code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="6ZQslRT0Fqy-" colab_type="text"
# ## FARM: Use your own dataset
#
# In Tutorial 1 you already learned about the major building blocks.
# In this tutorial, you will see how to use FARM with your own dataset.
# + id="usDHxZCBFxgU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="1303e268-39b2-4fa5-a5bf-999965638dc7"
# !git clone https://github.com/deepset-ai/FARM.git
import os
os.chdir("FARM")
# !pip install -r requirements.txt
# !pip install --editable .
# + id="x2N7NwpuFqzD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="e2327856-63cf-42b6-f1a6-c930b2f00bd0"
# Let's start by adjust the working directory so that it is the root of the repository
# This should be run just once.
import os
os.chdir('../')
print("Current working directory is {}".format(os.getcwd()))
# + [markdown] id="ESbe1stqFqzT" colab_type="text"
# # 1) How a Processor works
# + [markdown] id="KlrvOVB4FqzV" colab_type="text"
# ### Architecture
# The Processor converts a <b>raw input (e.g File) into a Pytorch dataset</b>.
# For using an own dataset we need to adjust this Processor.
#
# <img src="https://raw.githubusercontent.com/deepset-ai/FARM/master/docs/img/data_silo_no_bg.jpg" width="400" height="400" align="left"/>
# <br/><br/>
# <br/><br/>
# <br/><br/>
# <br/><br/>
# <br/><br/>
# <br/><br/>
# <br/><br/>
#
#
# ### Main Conversion Stages
# 1. Read from file / raw input
# 2. Create samples
# 3. Featurize samples
# 4. Create PyTorch Dataset
#
# ### Functions to implement
# 1. file\_to_dicts()
# 2. \_dict_to_samples()
# 3. \_sample_to_features()
# + [markdown] id="11IJzSOtFqzW" colab_type="text"
# ## Example: TextClassificationProcessor
# + id="PjwdbEnOFqzZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} outputId="ef59991f-f22e-4821-dd43-659341d841a4"
from farm.data_handler.processor import *
from farm.data_handler.samples import Sample
from farm.modeling.tokenization import Tokenizer, tokenize_with_metadata
import os
class TextClassificationProcessor(Processor):
"""
Used to handle the text classification datasets that come in tabular format (CSV, TSV, etc.)
"""
def __init__(
self,
tokenizer,
max_seq_len,
data_dir,
label_list=None,
metric=None,
train_filename="train.tsv",
dev_filename=None,
test_filename="test.tsv",
dev_split=0.1,
delimiter="\t",
quote_char="'",
skiprows=None,
label_column_name="label",
multilabel=False,
header=0,
proxies=None,
max_samples=None,
**kwargs
):
#TODO If an arg is misspelt, e.g. metrics, it will be swallowed silently by kwargs
# Custom processor attributes
self.delimiter = delimiter
self.quote_char = quote_char
self.skiprows = skiprows
self.header = header
self.max_samples = max_samples
super(TextClassificationProcessor, self).__init__(
tokenizer=tokenizer,
max_seq_len=max_seq_len,
train_filename=train_filename,
dev_filename=dev_filename,
test_filename=test_filename,
dev_split=dev_split,
data_dir=data_dir,
tasks={},
proxies=proxies,
)
if metric and label_list:
if multilabel:
task_type = "multilabel_classification"
else:
task_type = "classification"
self.add_task(name="text_classification",
metric=metric,
label_list=label_list,
label_column_name=label_column_name,
task_type=task_type)
else:
logger.info("Initialized processor without tasks. Supply `metric` and `label_list` to the constructor for "
"using the default task or add a custom task later via processor.add_task()")
def file_to_dicts(self, file: str) -> [dict]:
column_mapping = {task["label_column_name"]: task["label_name"] for task in self.tasks.values()}
dicts = read_tsv(
filename=file,
delimiter=self.delimiter,
skiprows=self.skiprows,
quotechar=self.quote_char,
rename_columns=column_mapping,
header=self.header,
proxies=self.proxies,
max_samples=self.max_samples
)
return dicts
def _dict_to_samples(self, dictionary: dict, **kwargs) -> [Sample]:
# this tokenization also stores offsets and a start_of_word mask
text = dictionary["text"]
tokenized = tokenize_with_metadata(text, self.tokenizer)
if len(tokenized["tokens"]) == 0:
logger.warning(f"The following text could not be tokenized, likely because it contains a character that the tokenizer does not recognize: {text}")
return []
# truncate tokens, offsets and start_of_word to max_seq_len that can be handled by the model
for seq_name in tokenized.keys():
tokenized[seq_name], _, _ = truncate_sequences(seq_a=tokenized[seq_name], seq_b=None, tokenizer=self.tokenizer,
max_seq_len=self.max_seq_len)
return [Sample(id=None, clear_text=dictionary, tokenized=tokenized)]
def _sample_to_features(self, sample) -> dict:
features = sample_to_features_text(
sample=sample,
tasks=self.tasks,
max_seq_len=self.max_seq_len,
tokenizer=self.tokenizer,
)
return features
# Helper
def read_tsv(filename, rename_columns, quotechar='"', delimiter="\t", skiprows=None, header=0, proxies=None, max_samples=None):
"""Reads a tab separated value file. Tries to download the data if filename is not found"""
# get remote dataset if needed
if not (os.path.exists(filename)):
logger.info(f" Couldn't find {filename} locally. Trying to download ...")
_download_extract_downstream_data(filename)
# read file into df
df = pd.read_csv(
filename,
sep=delimiter,
encoding="utf-8",
quotechar=quotechar,
dtype=str,
skiprows=skiprows,
header=header
)
# let's rename our target columns to the default names FARM expects:
# "text": contains the text
# "text_classification_label": contains a label for text classification
columns = ["text"] + list(rename_columns.keys())
df = df[columns]
for source_column, label_name in rename_columns.items():
df[label_name] = df[source_column]
df.drop(columns=[source_column], inplace=True)
if "unused" in df.columns:
df.drop(columns=["unused"], inplace=True)
raw_dict = df.to_dict(orient="records")
return raw_dict
# + id="Zjsd5TFDFqzl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="b5fb3077-f0ae-499e-e9f6-9bedb0d6fa7f"
# The default format is:
# - tab separated
# - column "text"
# - column "label"
import pandas as pd
df = pd.DataFrame({"text": ["The concerts supercaliphractisch was great!", "I hate people ignoring climate change."],
"label": ["positive","negative"]
})
print(df)
df.to_csv("train.tsv", sep="\t")
# + id="F8msjxryFqzv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 203, "referenced_widgets": ["0fedf11acab6486d8c734b79290d6875", "daced229491c441ea69ffe85c36867a5", "c39407be900946029a6103d448fb8fb2", "c990a7513c1a4a3398eaef7699fcda7b", "a421ec39b3b944e6bd444ef6019a5808", "29c9b4c052f5482097b678a2b74bb626", "7dcf072091ac4753a9d2a8f41d5ecc82", "2be8f3cd6a81443a8d74f08a98c2b23b"]} outputId="80aed84a-7183-48b4-e932-fc61b049ab0c"
tokenizer = Tokenizer.load(
pretrained_model_name_or_path="bert-base-uncased")
processor = TextClassificationProcessor(data_dir = "",
tokenizer=tokenizer,
max_seq_len=64,
label_list=["positive","negative"],
label_column_name="label",
metric="acc",
)
# + id="euqZZc7iFqz3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="c0f90f65-49d8-4a71-e2f8-ff9ee61b65b8"
# 1. One File -> Dictionarie(s) with "raw data"
dicts = processor.file_to_dicts(file="train.tsv")
print(dicts)
# + id="QVyOl5evFqz9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 517} outputId="14ec7a69-1170-48aa-997a-1176ff2651e3"
# 2. One Dictionary -> Sample(s)
# (Sample = "clear text" model input + meta information)
samples = processor._dict_to_samples(dictionary=dicts[0])
# print each attribute of sample
print(samples[0].clear_text)
print(samples[0].tokenized)
print(samples[0].features)
print("----------------------------------\n\n\n")
# or in a nicer, formatted style
print(samples[0])
# + id="-sOWhXIHFq0C" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="96b296a4-69fc-48fd-8912-346defc20aae"
# 3. One Sample -> Features
# (Features = "vectorized" model input)
features = processor._sample_to_features(samples[0])
print(features[0])
# + [markdown] id="03hkuHsoFq0J" colab_type="text"
# # 2) Hands-On: Adjust it to your dataset
# + [markdown] id="wCoBmkpOFq0K" colab_type="text"
# ## Task 1: Use an existing Processor
#
# This works if you have:
# - standard tasks
# - common file formats
#
# **Example: Text classification on CSV with multiple columns**
#
# Dataset: GermEval18 (Hatespeech detection)
# Format: TSV
# Columns: `text coarse_label fine_label`
# + id="fCN7UkbGFq0L" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 236} outputId="1f86d316-0522-4a56-b195-a8a82f22034e"
# Download dataset
from farm.data_handler import utils
utils._download_extract_downstream_data("germeval18/train.tsv")
# !head -n 10 germeval18/train.tsv
# TODO: Initialize a processor for the above file by passing the right arguments
processor = TextClassificationProcessor(tokenizer=tokenizer,
max_seq_len=128,
data_dir="germeval18",
train_filename="train.tsv",
label_list=["OTHER","OFFENSE"],
metric="acc",
label_column_name="coarse_label"
)
# + id="aBX3eyjFFq0R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="acea2635-dd88-4382-ba7d-8d98a696a3e2"
# test it
dicts = processor.file_to_dicts(file="germeval18/train.tsv")
print(dicts[0])
assert dicts[0] == {'text': '@corinnamilborn <NAME>, wir würden dich gerne als Moderatorin für uns gewinnen! Wärst du begeisterbar?', 'text_classification_label': 'OTHER'}
# + [markdown] id="pnIO1whZFq0X" colab_type="text"
# ## Task 2: Build your own Processor
# This works best for:
# - custom input files
# - special preprocessing steps
# - advanced multitask learning
#
# **Example: Text classification with JSON as input file**
#
# Dataset: [100k Yelp reviews](https://s3.eu-central-1.amazonaws.com/deepset.ai-farm-downstream/yelp_reviews_100k.json) ( [full dataset](https://https://www.yelp.com/dataset/download), [documentation](https://https://www.yelp.com/dataset/documentation/main))
#
# Format:
#
# ```
# {
# ...
# // integer, star rating
# "stars": 4,
#
# // string, the review itself
# "text": "Great place to hang out after work: the prices are decent, and the ambience is fun. It's a bit loud, but very lively. The staff is friendly, and the food is good. They have a good selection of drinks.",
# ...
# }
# ```
# + id="gViE3EnNFq0Z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 302} outputId="91b6531b-18bc-4df6-9df5-031878905cf2"
# Download dataset
# !wget https://s3.eu-central-1.amazonaws.com/deepset.ai-farm-downstream/yelp_reviews_100k.json
# !head -5 yelp_reviews_100k.json
# + id="BHtzq11EFq0d" colab_type="code" colab={}
import pandas as pd
# TODO: Create a new Processor class and overwrite the function that reads from the file
# The dicts created should look like this to comply with the default TextClassificationProcessor.
#{'text': 'Total bill for this horrible service? ...',
# 'text_classification_label': '4'}
class CustomTextClassificationProcessor(TextClassificationProcessor):
# we need to overwrite this function from the parent class
def file_to_dicts(self, file: str) -> [dict]:
# read into df
df = pd.read_json(file, lines=True)
# rename
df["text_classification_label"] = df["stars"].astype(str)
# drop unused
columns = ["text_classification_label","text"]
df = df[columns]
# convert to dicts
dicts = df.to_dict(orient="records")
return dicts
# + id="ovkucFdrFq0k" colab_type="code" colab={}
processor = CustomTextClassificationProcessor(tokenizer=tokenizer,
max_seq_len=128,
data_dir="",
label_list=["1","2","3","4","5"],
metric="acc",
)
# + id="Kfy9RsbWFq0o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="3883887a-d931-497d-8ab9-e1024e039207"
# test it
dicts = processor.file_to_dicts(file="yelp_reviews_100k.json")
print(dicts[0])
assert dicts[0] == {'text_classification_label': '1', 'text': 'Total bill for this horrible service? Over $8Gs. These crooks actually had the nerve to charge us $69 for 3 pills. I checked online the pills can be had for 19 cents EACH! Avoid Hospital ERs at all costs.'}
# + id="ga1r_ZgbFq0t" colab_type="code" colab={}
| tutorials/2_Build_a_processor_for_your_own_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/shamoji101/TensorFlow2.0Gym/blob/master/TensorFlow20_test.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="e0nJc-0UktwV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="6e3aa93b-8f61-476a-e24c-c6b2610e2a21"
# %tensorflow_version 2.x
# + id="hamXgYXAlJ84" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="641f0809-5234-4e77-c2bd-46c81f527a74"
import tensorflow as tf
tf.__version__
# + id="8bCvj5fZlNwc" colab_type="code" colab={}
| TensorFlow20_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ma_modells
# language: python
# name: ma_modells
# ---
# # Auswertung des Fine-Tunings pro Anomalie-Art
# +
import arrow
import numpy as np
import os
import glob
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import torch
import joblib
from sklearn.preprocessing import MinMaxScaler
from torch.utils.data import TensorDataset
from sklearn.metrics import confusion_matrix
from models.SimpleAutoEncoder import SimpleAutoEncoder
from utils.evalUtils import calc_cm_metrics
from utils.evalUtils import print_confusion_matrix
# -
from matplotlib import rc
rc('text', usetex=True)
# %run -i ./scripts/setConfigs.py
os.getcwd()
# %run -i ./scripts/EvalPreperations.py
type(drifted_anormal_torch_tensor)
# ## Read Experimet Results from Phase I
os.chdir(os.path.join(exp_data_path, 'experiment'))
extension = 'csv'
result = glob.glob('*.{}'.format(extension))
result[0]
df_tVP1_m1 = pd.read_csv(result[0])
for file in result[1:]:
df = pd.read_csv(file)
df_tVP1_m1 = df_tVP1_m1.append(df)
df_tVP1_m1.reset_index(inplace=True)
len(df_tVP1_m1)
if 'Unnamed: 0' in df_tVP1_m1.columns:
print('Drop unnamed shit col...')
df_tVP1_m1.drop('Unnamed: 0', axis=1,inplace=True)
df_tVP1_m1.head(1)
df_results = pd.DataFrame()
df_results['ano_labels'] = s_ano_labels_drifted_ano
df_results.reset_index(inplace=True, drop=True)
# ## Let every model predict on $X_{drifted,ano}$
for i, row in df_tVP1_m1.iterrows():
print('Current Iteration: {} of {}'.format(i+1, len(df_tVP1_m1)))
model = None
model = SimpleAutoEncoder(num_inputs=17, val_lambda=42)
logreg = None
model_fn = row['model_fn']
logreg_fn = row['logreg_fn']
logreg = joblib.load(logreg_fn)
model.load_state_dict(torch.load(model_fn))
losses = []
for val in drifted_anormal_torch_tensor:
loss = model.calc_reconstruction_error(val)
losses.append(loss.item())
s_losses_anormal = pd.Series(losses)
X = s_losses_anormal.to_numpy()
X = X.reshape(-1, 1)
predictions = []
for val in X:
val = val.reshape(1,-1)
pred = logreg.predict(val)
predictions.append(pred[0])
col_name = 'preds_model_{}'.format(i)
df_results[col_name] = predictions
file_name = '{}_predictions_models_phase_1_on_x_drifted_ano.csv'.format(arrow.now().format('YYYYMMDD'))
full_fn = '/home/torge/dev/masterthesis_code/02_Experimente/03_Experimente/exp_data/evaluation/{}'.format(file_name)
df_results.to_csv(full_fn, sep=';', index=False)
len(df_results)
len(df_results.columns)
df_results.head()
df_results['binary_label'] = [1 if x > 0 else 0 for x in df_results['ano_labels']]
# +
phase_1_general_kpis_per_model = {
'acc': [],
'prec': [],
'spec': [],
'sen': []
}
for col in df_results.columns:
if col != 'ano_labels' and col != 'binary_label':
cm = confusion_matrix(df_results['binary_label'], df_results[col])
tn, fp, fn, tp = cm.ravel()
accuracy, precision, specifity, sensitivity, f1_score = calc_cm_metrics(tp, tn, fp, fn)
phase_1_general_kpis_per_model['acc'].append(accuracy)
phase_1_general_kpis_per_model['prec'].append(precision)
phase_1_general_kpis_per_model['spec'].append(specifity)
phase_1_general_kpis_per_model['sen'].append(sensitivity)
# -
df_phase_1_general_kpis_per_model = pd.DataFrame(phase_1_general_kpis_per_model)
df_phase_1_general_kpis_per_model.head()
df_phase_1_general_kpis_per_model.describe()
df_results_anos_0 = df_results[df_results['ano_labels'] == 0.0]
df_results_anos_1 = df_results[df_results['ano_labels'] == 1.0]
df_results_anos_2 = df_results[df_results['ano_labels'] == 2.0]
df_results_anos_3 = df_results[df_results['ano_labels'] == 3.0]
print('Anzahl Samples in Klasse 0: {}'.format(len(df_results_anos_0)))
print('Anzahl Samples in Klasse 1: {}'.format(len(df_results_anos_1)))
print('Anzahl Samples in Klasse 2: {}'.format(len(df_results_anos_2)))
print('Anzahl Samples in Klasse 3: {}'.format(len(df_results_anos_3)))
# ## Create KPIs for every model per Anomaly class
kpis_per_model_anos_0 = {
'acc': [],
'prec': [],
'spec': [],
'sen': []
}
kpis_per_model_anos_1 = {
'acc': [],
'prec': [],
'spec': [],
'sen': []
}
kpis_per_model_anos_2 = {
'acc': [],
'prec': [],
'spec': [],
'sen': []
}
kpis_per_model_anos_3 = {
'acc': [],
'prec': [],
'spec': [],
'sen': []
}
label_ano_class_0 = np.zeros(len(df_results_anos_0))
label_ano_class_1 = np.ones(len(df_results_anos_1))
label_ano_class_2 = np.ones(len(df_results_anos_2))
label_ano_class_3 = np.ones(len(df_results_anos_3))
for col in df_results_anos_0.columns:
if col != 'ano_labels':
cm = confusion_matrix(label_ano_class_0, df_results_anos_0[col])
tn, fp, fn, tp = cm.ravel()
accuracy, precision, specifity, sensitivity, f1_score = calc_cm_metrics(tp, tn, fp, fn)
kpis_per_model_anos_0['acc'].append(accuracy)
kpis_per_model_anos_0['prec'].append(precision)
kpis_per_model_anos_0['spec'].append(specifity)
kpis_per_model_anos_0['sen'].append(sensitivity)
df_kpis_per_model_anos_0 = pd.DataFrame(kpis_per_model_anos_0)
df_kpis_per_model_anos_0.head()
df_kpis_per_model_anos_0.describe()
print(1.428468e-14)
len(df_kpis_per_model_anos_0)
for col in df_results_anos_1.columns:
if col != 'ano_labels':
cm = confusion_matrix(label_ano_class_1, df_results_anos_1[col])
tn, fp, fn, tp = cm.ravel()
accuracy, precision, specifity, sensitivity, f1_score = calc_cm_metrics(tp, tn, fp, fn)
kpis_per_model_anos_1['acc'].append(accuracy)
kpis_per_model_anos_1['prec'].append(precision)
kpis_per_model_anos_1['spec'].append(specifity)
kpis_per_model_anos_1['sen'].append(sensitivity)
df_kpis_per_model_anos_1 = pd.DataFrame(kpis_per_model_anos_1)
df_kpis_per_model_anos_1.head()
df_kpis_per_model_anos_1.describe()
for col in df_results_anos_2.columns:
if col != 'ano_labels':
cm = confusion_matrix(label_ano_class_2, df_results_anos_2[col])
tn, fp, fn, tp = cm.ravel()
accuracy, precision, specifity, sensitivity, f1_score = calc_cm_metrics(tp, tn, fp, fn)
kpis_per_model_anos_2['acc'].append(accuracy)
kpis_per_model_anos_2['prec'].append(precision)
kpis_per_model_anos_2['spec'].append(specifity)
kpis_per_model_anos_2['sen'].append(sensitivity)
df_kpis_per_model_anos_2 = pd.DataFrame(kpis_per_model_anos_2)
df_kpis_per_model_anos_2.head()
df_kpis_per_model_anos_2.describe()
for col in df_results_anos_3.columns:
if col != 'ano_labels':
cm = confusion_matrix(label_ano_class_3, df_results_anos_3[col])
tn, fp, fn, tp = cm.ravel()
accuracy, precision, specifity, sensitivity, f1_score = calc_cm_metrics(tp, tn, fp, fn)
kpis_per_model_anos_3['acc'].append(accuracy)
kpis_per_model_anos_3['prec'].append(precision)
kpis_per_model_anos_3['spec'].append(specifity)
kpis_per_model_anos_3['sen'].append(sensitivity)
df_kpis_per_model_anos_3 = pd.DataFrame(kpis_per_model_anos_3)
df_kpis_per_model_anos_3.head()
df_kpis_per_model_anos_3.describe()
arrow.now()
# ## Results from Phase II
os.getcwd()
os.chdir(os.path.join(exp_data_path, 'experiment', 'fine_tuning'))
extension = 'csv'
result = glob.glob('*.{}'.format(extension))
# +
real_results = []
for res in result:
if 'tVPII_M2' not in res:
real_results.append(res)
# -
df_tVP2_m1 = pd.read_csv(real_results[0], sep=';')
df_tVP2_m1.head()
for file in real_results[1:]:
df = pd.read_csv(file, sep=';')
df_tVP2_m1 = df_tVP2_m1.append(df)
df_tVP2_m1.reset_index(drop=True, inplace=True)
df_tVP2_m1.head(2)
len(df_tVP2_m1)
df_results_phase_2 = pd.DataFrame()
df_results_phase_2['ano_labels'] = s_ano_labels_drifted_ano
df_results_phase_2.reset_index(inplace=True, drop=True)
# +
print('Started: {}'.format(arrow.now().format('HH:mm:ss')))
for i, row in df_tVP2_m1.iterrows():
print('Current Iteration: {} of {}'.format(i+1, len(df_tVP2_m1)))
model = None
model = SimpleAutoEncoder(num_inputs=17, val_lambda=42)
logreg = None
model_fn = row['model_fn']
logreg_fn = row['logreg_fn']
logreg = joblib.load(logreg_fn)
model.load_state_dict(torch.load(model_fn))
losses = []
for val in drifted_anormal_torch_tensor:
loss = model.calc_reconstruction_error(val)
losses.append(loss.item())
s_losses_anormal = pd.Series(losses)
X = s_losses_anormal.to_numpy()
X = X.reshape(-1, 1)
predictions = []
for val in X:
val = val.reshape(1,-1)
pred = logreg.predict(val)
predictions.append(pred[0])
col_name = 'preds_model_{}'.format(i)
df_results_phase_2[col_name] = predictions
print('Ended: {}'.format(arrow.now().format('HH:mm:ss')))
# -
file_name = '{}_predictions_models_phase_2_on_x_drifted_ano.csv'.format(arrow.now().format('YYYYMMDD'))
full_fn = '/home/torge/dev/masterthesis_code/02_Experimente/03_Experimente/exp_data/evaluation/{}'.format(file_name)
df_results_phase_2.to_csv(full_fn, sep=';', index=False)
len(df_results_phase_2)
len(df_results_phase_2.columns)
print(df_results_phase_2['binary_label'].sum())
print(len(df_results_phase_2))
df_results_phase_2.head()
df_results_phase_2['binary_label'] = [1 if x > 0 else 0 for x in df_results_phase_2['ano_labels']]
phase_2_general_kpis_per_model = {
'acc': [],
'prec': [],
'spec': [],
'sen': []
}
for col in df_results_phase_2.columns:
if col != 'ano_labels' and col != 'binary_label':
cm = confusion_matrix(df_results_phase_2['binary_label'], df_results_phase_2[col])
tn, fp, fn, tp = cm.ravel()
accuracy, precision, specifity, sensitivity, f1_score = calc_cm_metrics(tp, tn, fp, fn)
phase_2_general_kpis_per_model['acc'].append(accuracy)
phase_2_general_kpis_per_model['prec'].append(precision)
phase_2_general_kpis_per_model['spec'].append(specifity)
phase_2_general_kpis_per_model['sen'].append(sensitivity)
df_phase_2_general_kpis_per_model = pd.DataFrame(phase_2_general_kpis_per_model)
df_phase_2_general_kpis_per_model.describe()
# ## Analyse pro Anomalie Art für Phase II
df_results_phase_2_anos_0 = df_results_phase_2[df_results_phase_2['ano_labels'] == 0.0]
df_results_phase_2_anos_1 = df_results_phase_2[df_results_phase_2['ano_labels'] == 1.0]
df_results_phase_2_anos_2 = df_results_phase_2[df_results_phase_2['ano_labels'] == 2.0]
df_results_phase_2_anos_3 = df_results_phase_2[df_results_phase_2['ano_labels'] == 3.0]
print('Anzahl Samples in Klasse 0: {}'.format(len(df_results_phase_2_anos_0)))
print('Anzahl Samples in Klasse 1: {}'.format(len(df_results_phase_2_anos_1)))
print('Anzahl Samples in Klasse 2: {}'.format(len(df_results_phase_2_anos_2)))
print('Anzahl Samples in Klasse 3: {}'.format(len(df_results_phase_2_anos_3)))
phase_2_kpis_per_model_anos_0 = {
'acc': [],
'prec': [],
'spec': [],
'sen': []
}
phase_2_kpis_per_model_anos_1 = {
'acc': [],
'prec': [],
'spec': [],
'sen': []
}
phase_2_kpis_per_model_anos_2 = {
'acc': [],
'prec': [],
'spec': [],
'sen': []
}
phase_2_kpis_per_model_anos_3 = {
'acc': [],
'prec': [],
'spec': [],
'sen': []
}
phase_2_label_ano_class_0 = np.zeros(len(df_results_phase_2_anos_0))
phase_2_label_ano_class_1 = np.ones(len(df_results_phase_2_anos_1))
phase_2_label_ano_class_2 = np.ones(len(df_results_phase_2_anos_2))
phase_2_label_ano_class_3 = np.ones(len(df_results_phase_2_anos_3))
# ### Kennzahlen Anomaly Klasse 0 nach Phase II Modell 1
for col in df_results_phase_2_anos_0.columns:
if col != 'ano_labels' and col != 'binary_label':
cm = confusion_matrix(phase_2_label_ano_class_0, df_results_phase_2_anos_0[col])
try:
tn, fp, fn, tp = cm.ravel()
except:
print('Cant ravel CM : {}'.format(col))
if df_results_phase_2_anos_0[col].all() == 1:
tp = 0
tn = 0
fp = cm[0][0]
fn = 0
accuracy, precision, specifity, sensitivity, f1_score = calc_cm_metrics(tp, tn, fp, fn)
phase_2_kpis_per_model_anos_0['acc'].append(accuracy)
phase_2_kpis_per_model_anos_0['prec'].append(precision)
phase_2_kpis_per_model_anos_0['spec'].append(specifity)
phase_2_kpis_per_model_anos_0['sen'].append(sensitivity)
df_phase_2_kpis_per_model_anos_0 = pd.DataFrame(phase_2_kpis_per_model_anos_0)
df_phase_2_kpis_per_model_anos_0.head()
df_phase_2_kpis_per_model_anos_0.describe()
# ### Kennzahlen Anomaly Klasse 1 nach Phase II Modell 1
for col in df_results_phase_2_anos_1.columns:
if col != 'ano_labels' and col != 'binary_label':
cm = confusion_matrix(phase_2_label_ano_class_1, df_results_phase_2_anos_1[col])
try:
tn, fp, fn, tp = cm.ravel()
except:
print('Cant ravel CM : {}'.format(col))
if df_results_phase_2_anos_1[col].all() == 1:
tp = cm[0][0]
tn = 0
fp = 0
fn = 0
accuracy, precision, specifity, sensitivity, f1_score = calc_cm_metrics(tp, tn, fp, fn)
phase_2_kpis_per_model_anos_1['acc'].append(accuracy)
phase_2_kpis_per_model_anos_1['prec'].append(precision)
phase_2_kpis_per_model_anos_1['spec'].append(specifity)
phase_2_kpis_per_model_anos_1['sen'].append(sensitivity)
df_phase_2_kpis_per_model_anos_1 = pd.DataFrame(phase_2_kpis_per_model_anos_1)
df_phase_2_kpis_per_model_anos_1.head()
df_phase_2_kpis_per_model_anos_1.describe()
# ### Kennzahlen Anomaly Klasse 2 nach Phase II Modell 1
for col in df_results_phase_2_anos_2.columns:
if col != 'ano_labels' and col != 'binary_label':
cm = confusion_matrix(phase_2_label_ano_class_2, df_results_phase_2_anos_2[col])
try:
tn, fp, fn, tp = cm.ravel()
except:
print('Cant ravel CM : {}'.format(col))
if df_results_phase_2_anos_2[col].all() == 1:
tp = cm[0][0]
tn = 0
fp = 0
fn = 0
accuracy, precision, specifity, sensitivity, f1_score = calc_cm_metrics(tp, tn, fp, fn)
phase_2_kpis_per_model_anos_2['acc'].append(accuracy)
phase_2_kpis_per_model_anos_2['prec'].append(precision)
phase_2_kpis_per_model_anos_2['spec'].append(specifity)
phase_2_kpis_per_model_anos_2['sen'].append(sensitivity)
df_phase_2_kpis_per_model_anos_2 = pd.DataFrame(phase_2_kpis_per_model_anos_2)
df_phase_2_kpis_per_model_anos_2.head()
df_phase_2_kpis_per_model_anos_2.describe()
# ### Kennzahlen Anomaly Klasse 3 nach Phase II Modell 1
for col in df_results_phase_2_anos_3.columns:
if col != 'ano_labels' and col !='binary_label':
cm = confusion_matrix(phase_2_label_ano_class_3, df_results_phase_2_anos_3[col])
try:
tn, fp, fn, tp = cm.ravel()
except:
print('Cant ravel CM : {}'.format(col))
if df_results_phase_2_anos_3[col].all() == 1:
tp = cm[0][0]
tn = 0
fp = 0
fn = 0
accuracy, precision, specifity, sensitivity, f1_score = calc_cm_metrics(tp, tn, fp, fn)
phase_2_kpis_per_model_anos_3['acc'].append(accuracy)
phase_2_kpis_per_model_anos_3['prec'].append(precision)
phase_2_kpis_per_model_anos_3['spec'].append(specifity)
phase_2_kpis_per_model_anos_3['sen'].append(sensitivity)
df_phase_2_kpis_per_model_anos_3 = pd.DataFrame(phase_2_kpis_per_model_anos_3)
df_phase_2_kpis_per_model_anos_3.head()
df_phase_2_kpis_per_model_anos_3.describe()
# # Veränderung der Modellperformance pro Concept Drift Event Art
df_results_phase_1_concept_drift = df_results.copy()
df_results_phase_1_concept_drift.head(1)
s_drift_labls_copy = s_drift_labels_drifted_ano.copy()
s_drift_labls_copy.reset_index(drop=True, inplace=True)
df_results_phase_1_concept_drift['cd_label'] = s_drift_labls_copy
df_res_phase_1_cd_0 = df_results_phase_1_concept_drift[df_results_phase_1_concept_drift['cd_label'] == 0.0]
df_res_phase_1_cd_1 = df_results_phase_1_concept_drift[df_results_phase_1_concept_drift['cd_label'] == 1.0]
df_res_phase_1_cd_2 = df_results_phase_1_concept_drift[df_results_phase_1_concept_drift['cd_label'] == 2.0]
df_res_phase_1_cd_3 = df_results_phase_1_concept_drift[df_results_phase_1_concept_drift['cd_label'] == 3.0]
print('Anzahl CD Event 0: {}'.format(len(df_res_phase_1_cd_0)))
print('Anzahl CD Event 1: {}'.format(len(df_res_phase_1_cd_1)))
print('Anzahl CD Event 2: {}'.format(len(df_res_phase_1_cd_2)))
print('Anzahl CD Event 3: {}'.format(len(df_res_phase_1_cd_3)))
lab_res_phase_1_cd_0 = df_res_phase_1_cd_0['binary_label']
lab_res_phase_1_cd_1 = df_res_phase_1_cd_1['binary_label']
lab_res_phase_1_cd_2 = df_res_phase_1_cd_2['binary_label']
lab_res_phase_1_cd_3 = df_res_phase_1_cd_3['binary_label']
# ## Concept Drivt Event 0
# +
kpis_phase_1_cd_0 = {
'acc': [],
'prec': [],
'spec': [],
'sen': []
}
for col in df_res_phase_1_cd_0.columns:
if col != 'ano_labels' and col !='binary_label' and col !='cd_label':
cm = confusion_matrix(lab_res_phase_1_cd_0, df_res_phase_1_cd_0[col])
try:
tn, fp, fn, tp = cm.ravel()
except:
print('Cant ravel CM : {}'.format(col))
#if df_results_phase_2_anos_3[col].all() == 1:
# tp = cm[0][0]
# tn = 0
# fp = 0
#fn = 0
accuracy, precision, specifity, sensitivity, f1_score = calc_cm_metrics(tp, tn, fp, fn)
kpis_phase_1_cd_0['acc'].append(accuracy)
kpis_phase_1_cd_0['prec'].append(precision)
kpis_phase_1_cd_0['spec'].append(specifity)
kpis_phase_1_cd_0['sen'].append(sensitivity)
# -
df_kpis_phase_1_cd_0 = pd.DataFrame(kpis_phase_1_cd_0)
df_kpis_phase_1_cd_0.describe()
# ## Concept Drift Event 1
# +
kpis_phase_1_cd_1 = {
'acc': [],
'prec': [],
'spec': [],
'sen': []
}
for col in df_res_phase_1_cd_0.columns:
if col != 'ano_labels' and col !='binary_label' and col !='cd_label':
cm = confusion_matrix(lab_res_phase_1_cd_1, df_res_phase_1_cd_1[col])
try:
tn, fp, fn, tp = cm.ravel()
except:
print('Cant ravel CM : {}'.format(col))
#if df_results_phase_2_anos_3[col].all() == 1:
# tp = cm[0][0]
# tn = 0
# fp = 0
#fn = 0
accuracy, precision, specifity, sensitivity, f1_score = calc_cm_metrics(tp, tn, fp, fn)
kpis_phase_1_cd_1['acc'].append(accuracy)
kpis_phase_1_cd_1['prec'].append(precision)
kpis_phase_1_cd_1['spec'].append(specifity)
kpis_phase_1_cd_1['sen'].append(sensitivity)
# -
df_kpis_phase_1_cd_1 = pd.DataFrame(kpis_phase_1_cd_1)
df_kpis_phase_1_cd_1.describe()
# ## Concept Drift Event 2
# +
kpis_phase_1_cd_2 = {
'acc': [],
'prec': [],
'spec': [],
'sen': []
}
for col in df_res_phase_1_cd_0.columns:
if col != 'ano_labels' and col !='binary_label' and col !='cd_label':
cm = confusion_matrix(lab_res_phase_1_cd_2, df_res_phase_1_cd_2[col])
try:
tn, fp, fn, tp = cm.ravel()
except:
print('Cant ravel CM : {}'.format(col))
#if df_results_phase_2_anos_3[col].all() == 1:
# tp = cm[0][0]
# tn = 0
# fp = 0
#fn = 0
accuracy, precision, specifity, sensitivity, f1_score = calc_cm_metrics(tp, tn, fp, fn)
kpis_phase_1_cd_2['acc'].append(accuracy)
kpis_phase_1_cd_2['prec'].append(precision)
kpis_phase_1_cd_2['spec'].append(specifity)
kpis_phase_1_cd_2['sen'].append(sensitivity)
# -
df_kpis_phase_1_cd_2 = pd.DataFrame(kpis_phase_1_cd_2)
df_kpis_phase_1_cd_2.describe()
# ## Concept Drift Event 3
# +
kpis_phase_1_cd_3 = {
'acc': [],
'prec': [],
'spec': [],
'sen': []
}
for col in df_res_phase_1_cd_0.columns:
if col != 'ano_labels' and col !='binary_label' and col !='cd_label':
cm = confusion_matrix(lab_res_phase_1_cd_3, df_res_phase_1_cd_3[col])
try:
tn, fp, fn, tp = cm.ravel()
except:
print('Cant ravel CM : {}'.format(col))
#if df_results_phase_2_anos_3[col].all() == 1:
# tp = cm[0][0]
# tn = 0
# fp = 0
#fn = 0
accuracy, precision, specifity, sensitivity, f1_score = calc_cm_metrics(tp, tn, fp, fn)
kpis_phase_1_cd_3['acc'].append(accuracy)
kpis_phase_1_cd_3['prec'].append(precision)
kpis_phase_1_cd_3['spec'].append(specifity)
kpis_phase_1_cd_3['sen'].append(sensitivity)
# -
df_kpis_phase_1_cd_3 = pd.DataFrame(kpis_phase_1_cd_3)
df_kpis_phase_1_cd_3.describe()
# ## Auswertung Kennzahlen pro Concept Drift Event Art nach Phase II
df_results_phase_2_concept_drift = df_results_phase_2.copy()
df_results_phase_2_concept_drift['cd_label'] = s_drift_labls_copy
df_results_phase_2_concept_drift.head(1)
df_res_phase_2_cd_0 = df_results_phase_2_concept_drift[df_results_phase_2_concept_drift['cd_label'] == 0.0]
df_res_phase_2_cd_1 = df_results_phase_2_concept_drift[df_results_phase_2_concept_drift['cd_label'] == 1.0]
df_res_phase_2_cd_2 = df_results_phase_2_concept_drift[df_results_phase_2_concept_drift['cd_label'] == 2.0]
df_res_phase_2_cd_3 = df_results_phase_2_concept_drift[df_results_phase_2_concept_drift['cd_label'] == 3.0]
print('Anzahl CD Event 0: {}'.format(len(df_res_phase_2_cd_0)))
print('Anzahl CD Event 1: {}'.format(len(df_res_phase_2_cd_1)))
print('Anzahl CD Event 2: {}'.format(len(df_res_phase_2_cd_2)))
print('Anzahl CD Event 3: {}'.format(len(df_res_phase_2_cd_3)))
lab_res_phase_2_cd_0 = df_res_phase_2_cd_0['binary_label']
lab_res_phase_2_cd_1 = df_res_phase_2_cd_1['binary_label']
lab_res_phase_2_cd_2 = df_res_phase_2_cd_2['binary_label']
lab_res_phase_2_cd_3 = df_res_phase_2_cd_3['binary_label']
# ### Concept Drift Event Art 0
# +
kpis_phase_2_cd_0 = {
'acc': [],
'prec': [],
'spec': [],
'sen': []
}
for col in df_res_phase_2_cd_0.columns:
if col != 'ano_labels' and col !='binary_label' and col !='cd_label':
cm = confusion_matrix(lab_res_phase_2_cd_0, df_res_phase_2_cd_0[col])
try:
tn, fp, fn, tp = cm.ravel()
except:
print('Cant ravel CM : {}'.format(col))
#if df_results_phase_2_anos_3[col].all() == 1:
# tp = cm[0][0]
# tn = 0
# fp = 0
#fn = 0
accuracy, precision, specifity, sensitivity, f1_score = calc_cm_metrics(tp, tn, fp, fn)
kpis_phase_2_cd_0['acc'].append(accuracy)
kpis_phase_2_cd_0['prec'].append(precision)
kpis_phase_2_cd_0['spec'].append(specifity)
kpis_phase_2_cd_0['sen'].append(sensitivity)
# -
df_kpis_phase_2_cd_0 = pd.DataFrame(kpis_phase_2_cd_0)
df_kpis_phase_2_cd_0.describe()
#
# ### Concept Drift Event Art 1
#
# kpis_phase_2_cd_1 = {
# 'acc': [],
# 'prec': [],
# 'spec': [],
# 'sen': []
# }
#
# for col in df_res_phase_2_cd_1.columns:
# if col != 'ano_labels' and col !='binary_label' and col !='cd_label':
# cm = confusion_matrix(lab_res_phase_2_cd_1, df_res_phase_2_cd_1[col])
# try:
# tn, fp, fn, tp = cm.ravel()
# except:
# print('Cant ravel CM : {}'.format(col))
# #if df_results_phase_2_anos_3[col].all() == 1:
# # tp = cm[0][0]
# # tn = 0
# # fp = 0
# #fn = 0
#
# accuracy, precision, specifity, sensitivity, f1_score = calc_cm_metrics(tp, tn, fp, fn)
#
# kpis_phase_2_cd_1['acc'].append(accuracy)
# kpis_phase_2_cd_1['prec'].append(precision)
# kpis_phase_2_cd_1['spec'].append(specifity)
# kpis_phase_2_cd_1['sen'].append(sensitivity)
df_kpis_phase_2_cd_1 = pd.DataFrame(kpis_phase_2_cd_1)
df_kpis_phase_2_cd_1.describe()
# ### Concept Drift Event Art 2
# +
kpis_phase_2_cd_2 = {
'acc': [],
'prec': [],
'spec': [],
'sen': []
}
for col in df_res_phase_2_cd_2.columns:
if col != 'ano_labels' and col !='binary_label' and col !='cd_label':
cm = confusion_matrix(lab_res_phase_2_cd_2, df_res_phase_2_cd_2[col])
try:
tn, fp, fn, tp = cm.ravel()
except:
print('Cant ravel CM : {}'.format(col))
#if df_results_phase_2_anos_3[col].all() == 1:
# tp = cm[0][0]
# tn = 0
# fp = 0
#fn = 0
accuracy, precision, specifity, sensitivity, f1_score = calc_cm_metrics(tp, tn, fp, fn)
kpis_phase_2_cd_2['acc'].append(accuracy)
kpis_phase_2_cd_2['prec'].append(precision)
kpis_phase_2_cd_2['spec'].append(specifity)
kpis_phase_2_cd_2['sen'].append(sensitivity)
# -
df_kpis_phase_2_cd_2 = pd.DataFrame(kpis_phase_2_cd_2)
df_kpis_phase_2_cd_2.describe()
# ### Concept Drift Event Art 3
# +
kpis_phase_2_cd_3 = {
'acc': [],
'prec': [],
'spec': [],
'sen': []
}
for col in df_res_phase_2_cd_3.columns:
if col != 'ano_labels' and col !='binary_label' and col !='cd_label':
cm = confusion_matrix(lab_res_phase_2_cd_3, df_res_phase_2_cd_3[col])
try:
tn, fp, fn, tp = cm.ravel()
except:
print('Cant ravel CM : {}'.format(col))
#if df_results_phase_2_anos_3[col].all() == 1:
# tp = cm[0][0]
# tn = 0
# fp = 0
#fn = 0
accuracy, precision, specifity, sensitivity, f1_score = calc_cm_metrics(tp, tn, fp, fn)
kpis_phase_2_cd_3['acc'].append(accuracy)
kpis_phase_2_cd_3['prec'].append(precision)
kpis_phase_2_cd_3['spec'].append(specifity)
kpis_phase_2_cd_3['sen'].append(sensitivity)
# -
df_kpis_phase_2_cd_3 = pd.DataFrame(kpis_phase_2_cd_3)
df_kpis_phase_2_cd_3.describe()
| 03_Experimente/04_B_Interpretation_per_Anomalie_Art_and_Concept_Drift.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Steps:
# - find the changes
# - find the cause of the changes (may include more data searching)
# +
#imports
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
pd.set_option("max_rows", 5)
# -
# ### Load data
# +
df = pd.read_table("Churn_UsersProfile.txt", delimiter='|',encoding ='latin1')
print(df.columns)
df["log_Average_BusUsers_per_Day"] = np.log10(df["Average_BusUsers_per_Day"])
df
# -
# ### Encode Labels with sklearn with for loop
# +
from sklearn.preprocessing import LabelEncoder
df2 = df
for i in range(len(df.columns)-1):
#print(i)
enc = LabelEncoder()
enc.fit(df.iloc[:,i]) # get a LabelEncoder encoding for the labels (returns?)
#print(enc.classes_)
newcol_name = df.columns[i]
newcol_name += '_enc'
df2[newcol_name] = enc.transform(df.iloc[:,i])
df
# -
#plt.plot(df["Region_of_Origin"], df["Average_BusUsers_per_Day"], '.')
ax = df.boxplot(column='log_Average_BusUsers_per_Day', by='Region_of_Origin', grid=True)
ax = df.boxplot(column='log_Average_BusUsers_per_Day', by='District_of_Origin', grid=True)
ax = df.boxplot(column='log_Average_BusUsers_per_Day', by='AgeClassDescription', grid=True)
ax = df.boxplot(column='log_Average_BusUsers_per_Day', by='GenderDescription', grid=True)
ax = df.boxplot(column='log_Average_BusUsers_per_Day', by='Period', grid=True)
#plot a histogram of period
| notebooks/stage1/Data visualisation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# -
#export
from exp.nb_10c import *
# ## Imagenet(te) training
path = datasets.untar_data(datasets.URLs.IMAGENETTE_160)
# +
size = 128
tfms = [make_rgb, RandomResizedCrop(size, scale=(0.35,1)), np_to_float, PilRandomFlip()]
bs = 64
il = ImageList.from_files(path, tfms=tfms)
sd = SplitData.split_by_func(il, partial(grandparent_splitter, valid_name='val'))
ll = label_by_func(sd, parent_labeler, proc_y=CategoryProcessor())
ll.valid.x.tfms = [make_rgb, CenterCrop(size), np_to_float]
data = ll.to_databunch(bs, c_in=3, c_out=10, num_workers=8)
# -
data.train_ds
# ## XResNet
import numpy as np
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
a = np.arange(12).reshape(2,3,2)
a
a = torch.tensor(a)
# a.size(0)
a.view(a.size(0), -1)
# +
#export
def noop(x): return x
class Flatten(nn.Module):
def forward(self, x): return x.view(x.size(0), -1)
def conv(ni, nf, ks=3, stride=1, bias=False):
return nn.Conv2d(ni, nf, kernel_size=ks, stride=stride, padding=ks//2, bias=bias)
# +
#export
act_fn = nn.ReLU(inplace=True)
def init_cnn(m):
if getattr(m, 'bias', None) is not None: nn.init.constant_(m.bias, 0)
if isinstance(m, (nn.Conv2d,nn.Linear)): nn.init.kaiming_normal_(m.weight)
for l in m.children(): init_cnn(l)
def conv_layer(ni, nf, ks=3, stride=1, zero_bn=False, act=True):
bn = nn.BatchNorm2d(nf)
nn.init.constant_(bn.weight, 0. if zero_bn else 1.)
layers = [conv(ni, nf, ks, stride=stride), bn]
if act: layers.append(act_fn)
return nn.Sequential(*layers)
# -
#export
class ResBlock(nn.Module):
def __init__(self, expansion, ni, nh, stride=1):
super().__init__()
nf,ni = nh*expansion,ni*expansion
layers = [conv_layer(ni, nh, 3, stride=stride),
conv_layer(nh, nf, 3, zero_bn=True, act=False)
] if expansion == 1 else [
conv_layer(ni, nh, 1),
conv_layer(nh, nh, 3, stride=stride),
conv_layer(nh, nf, 1, zero_bn=True, act=False)
]
self.convs = nn.Sequential(*layers)
self.idconv = noop if ni==nf else conv_layer(ni, nf, 1, act=False)
self.pool = noop if stride==1 else nn.AvgPool2d(2, ceil_mode=True)
def forward(self, x): return act_fn(self.convs(x) + self.idconv(self.pool(x)))
#export
class XResNet(nn.Sequential):
@classmethod
def create(cls, expansion, layers, c_in=3, c_out=1000):
nfs = [c_in, (c_in+1)*8, 64, 64]
stem = [conv_layer(nfs[i], nfs[i+1], stride=2 if i==0 else 1)
for i in range(3)]
nfs = [64//expansion,64,128,256,512]
res_layers = [cls._make_layer(expansion, nfs[i], nfs[i+1],
n_blocks=l, stride=1 if i==0 else 2)
for i,l in enumerate(layers)]
res = cls(
*stem,
nn.MaxPool2d(kernel_size=3, stride=2, padding=1),
*res_layers,
nn.AdaptiveAvgPool2d(1), Flatten(),
nn.Linear(nfs[-1]*expansion, c_out),
)
init_cnn(res)
return res
@staticmethod
def _make_layer(expansion, ni, nf, n_blocks, stride):
return nn.Sequential(
*[ResBlock(expansion, ni if i==0 else nf, nf, stride if i==0 else 1)
for i in range(n_blocks)])
#export
def xresnet18 (**kwargs): return XResNet.create(1, [2, 2, 2, 2], **kwargs)
def xresnet34 (**kwargs): return XResNet.create(1, [3, 4, 6, 3], **kwargs)
def xresnet50 (**kwargs): return XResNet.create(4, [3, 4, 6, 3], **kwargs)
def xresnet101(**kwargs): return XResNet.create(4, [3, 4, 23, 3], **kwargs)
def xresnet152(**kwargs): return XResNet.create(4, [3, 8, 36, 3], **kwargs)
# ## Train
cbfs = [partial(AvgStatsCallback,accuracy), ProgressCallback, CudaCallback,
partial(BatchTransformXCallback, norm_imagenette),
# partial(MixUp, alpha=0.2)
]
loss_func = LabelSmoothingCrossEntropy()
arch = partial(xresnet18, c_out=10)
opt_func = adam_opt(mom=0.9, mom_sqr=0.99, eps=1e-6, wd=1e-2)
#export
def get_batch(dl, learn):
learn.xb,learn.yb = next(iter(dl))
learn.do_begin_fit(0)
learn('begin_batch')
learn('after_fit')
return learn.xb,learn.yb
# We need to replace the old `model_summary` since it used to take a `Runner`.
# export
def model_summary(model, data, find_all=False, print_mod=False):
xb,yb = get_batch(data.valid_dl, learn)
mods = find_modules(model, is_lin_layer) if find_all else model.children()
f = lambda hook,mod,inp,out: print(f"====\n{mod}\n" if print_mod else "", out.shape)
with Hooks(mods, f) as hooks: learn.model(xb)
learn = Learner(arch(), data, loss_func, lr=1, cb_funcs=cbfs, opt_func=opt_func)
learn.model = learn.model.cuda()
model_summary(learn.model, data, print_mod=False)
arch = partial(xresnet34, c_out=10)
learn = Learner(arch(), data, loss_func, lr=1, cb_funcs=cbfs, opt_func=opt_func)
learn.fit(1, cbs=[LR_Find(), Recorder()])
learn.recorder.plot(3)
#export
def create_phases(phases):
phases = listify(phases)
return phases + [1-sum(phases)]
print(create_phases(0.3))
print(create_phases([0.3,0.2]))
# +
# # cos_1cycle_anneal??
# combine_scheds??
# -
lr = 1e-2
pct_start = 0.5
phases = create_phases(pct_start)
print(phases)
sched_lr = combine_scheds(phases, cos_1cycle_anneal(lr/10., lr, lr/1e5))
sched_mom = combine_scheds(phases, cos_1cycle_anneal(0.95, 0.85, 0.95))
sched_lr
ParamScheduler('lr', sched_lr)
ParamScheduler('mom', sched_mom)
cbsched = [
ParamScheduler('lr', sched_lr),
ParamScheduler('mom', sched_mom)]
learn = Learner(arch(), data, loss_func, lr=lr, cb_funcs=cbfs, opt_func=opt_func)
learn.model
learn.data
# +
# ParamScheduler??
# -
cbsched
learn.fit(5, cbs=cbsched)
# ## cnn_learner
#export
def cnn_learner(arch, data, loss_func, opt_func, c_in=None, c_out=None,
lr=1e-2, cuda=True, norm=None, progress=True, mixup=0, xtra_cb=None, **kwargs):
cbfs = [partial(AvgStatsCallback,accuracy)]+listify(xtra_cb)
if progress: cbfs.append(ProgressCallback)
if cuda: cbfs.append(CudaCallback)
if norm: cbfs.append(partial(BatchTransformXCallback, norm))
if mixup: cbfs.append(partial(MixUp, mixup))
arch_args = {}
if not c_in : c_in = data.c_in
if not c_out: c_out = data.c_out
if c_in: arch_args['c_in' ]=c_in
if c_out: arch_args['c_out']=c_out
return Learner(arch(**arch_args), data, loss_func, opt_func=opt_func, lr=lr, cb_funcs=cbfs, **kwargs)
learn = cnn_learner(xresnet34, data, loss_func, opt_func, norm=norm_imagenette)
learn.fit(5, cbsched)
# ## Imagenet
# You can see all this put together in the fastai [imagenet training script](https://github.com/fastai/fastai/blob/master/examples/train_imagenet.py). It's the same as what we've seen so far, except it also handles multi-GPU training. So how well does this work?
#
# We trained for 60 epochs, and got an error of 5.9%, compared to the official PyTorch resnet which gets 7.5% error in 90 epochs! Our xresnet 50 training even surpasses standard resnet 152, which trains for 50% more epochs and has 3x as many layers.
# ## Export
# !./notebook2script.py 11_train_imagenette.ipynb
| nbs/dl2/11_train_imagenette_sz_20191016.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="MqZGoP3StCih" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 154} outputId="819eb654-3827-480c-caaf-1f3d05d97667"
#remove " > /dev/null 2>&1" to see what is going on under the hood
# !pip install gym pyvirtualdisplay > /dev/null 2>&1
# !apt-get install -y xvfb python-opengl ffmpeg > /dev/null 2>&1
# !apt-get update > /dev/null 2>&1
# !apt-get install cmake > /dev/null 2>&1
# !pip install --upgrade setuptools 2>&1
# !pip install ez_setup > /dev/null 2>&1
# !pip install gym[all] > /dev/null 2>&1
# !pip install Box2D
# !pip install box2d
# + id="BwyeVcUWu7fg" colab_type="code" colab={}
import gym
from gym import logger as gymlogger
from gym.wrappers import Monitor
gymlogger.set_level(40) #error only
import math
import random
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
import glob
import io
import base64
from IPython.display import HTML
from IPython import display as ipythondisplay
from collections import namedtuple
from itertools import count
from PIL import Image
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import torchvision.transforms as T
# + id="V73JzLT3tZHa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="978ba78b-e56a-427f-b800-986686a7c495"
is_ipython = 'inline' in matplotlib.get_backend()
# if is_python: from IPython import display
from pyvirtualdisplay import Display
display = Display(visible=0, size=(1400, 900))
display.start()
# + id="Ache6SLOtb_0" colab_type="code" colab={}
"""
Utility functions to enable video recording of gym environment and displaying it
To enable video, just do "env = wrap_env(env)""
"""
def show_video():
mp4list = glob.glob('video/*.mp4')
if len(mp4list) > 0:
mp4 = mp4list[0]
video = io.open(mp4, 'r+b').read()
encoded = base64.b64encode(video)
ipythondisplay.display(HTML(data='''<video alt="test" autoplay
loop controls style="height: 400px;">
<source src="data:video/mp4;base64,{0}" type="video/mp4" />
</video>'''.format(encoded.decode('ascii'))))
else:
print("Could not find video")
def wrap_env(env):
env = Monitor(env, './video', force=True)
return env
# + [markdown] id="innirRJ-N0CY" colab_type="text"
# The “Critic” estimates the value function. This could be the action-value (the Q value) or state-value (the V value).
#
# The “Actor” updates the policy distribution in the direction suggested by the Critic (such as with policy gradients).
# + id="xWd_HrN9t6rR" colab_type="code" colab={}
class ActorCriticNetwork(nn.Module):
def __init__(self,alpha,input_dims,fc1_dims,fc2_dims,n_actions):
super(ActorCriticNetwork,self).__init__()
self.alpha = alpha
self.input_dims = input_dims
self.fc1_dims = fc1_dims
self.fc2_dims = fc2_dims
self.n_actions = n_actions
self.fc1 = nn.Linear(*self.input_dims,self.fc1_dims)
self.fc2 = nn.Linear(self.fc1_dims,self.fc2_dims)
self.pi = nn.Linear(self.fc2_dims,n_actions)
self.v = nn.Linear(self.fc2_dims,1)
self.optimizer = optim.Adam(self.parameters(),lr=self.alpha)
self.device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
self.to(self.device)
def forward(self, observation):
state = torch.Tensor(observation).to(self.device)
x = F.relu(self.fc1(state))
x = F.relu(self.fc2(x))
pi = self.pi(x)
v = self.v(x)
return pi,v
# + [markdown] id="y-Tmg0OpOJDw" colab_type="text"
# Here the agent chooses an action on the basis of the evaluation that the network makes when presented the current states of the environment.
#
# Thr Critic evaluates the actor's choice of action by evaluating the value function.
# + id="5YkJozhBteny" colab_type="code" colab={}
class NewAgent(object):
def __init__(self,alpha,input_dims,gamma=0.99,layer1_size=256,layer2_size=256,n_actions=4):
self.gamma = gamma
self.actor_critic = ActorCriticNetwork(alpha,input_dims,layer1_size,layer2_size,n_actions=n_actions)
self.log_probs = None
def choose_action(self,observation):
policy,_ = self.actor_critic.forward(observation)
policy = F.softmax(policy)
action_probs = torch.distributions.Categorical(policy)
action = action_probs.sample()
self.log_probs = action_probs.log_prob(action)
return action.item()
def learn(self, state, reward, state_, done):
self.actor_critic.optimizer.zero_grad()
_,critic_value = self.actor_critic.forward(state)
_,critic_value_ = self.actor_critic.forward(state_)
reward = torch.tensor(reward, dtype=torch.float).to(self.actor_critic.device)
delta = reward + self.gamma*critic_value_*(1-int(done)) - critic_value
actor_loss = -self.log_probs*delta #gradient ascent so it is -ve
critic_loss = delta**2
(actor_loss+critic_loss).backward()
self.actor_critic.optimizer.step()
# + id="u4G_3m8y5RZz" colab_type="code" colab={}
def plot(values,moving_avg_period):
plt.figure(2)
plt.clf()
plt.title('Training..')
plt.xlabel('Episode')
plt.ylabel('Duration')
plt.plot(values)
moving_avg = get_moving_average(moving_avg_period,values)
plt.plot(moving_avg)
plt.pause(0.001)
print("Episode",len(values),"\n",moving_avg_period,"episode moving avg:",moving_avg[-1])
if is_ipython: ipythondisplay.clear_output(wait=True)
def get_moving_average(period,values):
values = torch.tensor(values,dtype=torch.float)
if len(values)>=period:
moving_avg = values.unfold(dimension=0, size=period,step=1).mean(dim=1).flatten(start_dim=0)
moving_avg = torch.cat((torch.zeros(period-1),moving_avg))
return moving_avg.numpy()
else:
moving_avg = torch.zeros(len(values))
return moving_avg.numpy()
# + id="lQR6F1ests8r" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="8153674b-5f0b-4d54-fdf5-95b77bf7c2a8"
agent = NewAgent(alpha=0.00001, input_dims=[8], gamma=0.99, n_actions=4,layer1_size=2048,layer2_size=512)
env = wrap_env(gym.make('LunarLander-v2').unwrapped)
score_history = []
num_episodes = 2000
for i in range(num_episodes):
done = False
observation = env.reset()
score = 0
while not done:
action = agent.choose_action(observation)
observation_,reward,done,info = env.step(action)
agent.learn(observation,reward,observation_,done)
observation = observation_
score+=reward
score_history.append(score)
# print("episode",i,"score %.2f" %score)
plot(score_history,100)
env.close()
show_video()
# + id="tNeDTnOf9jOL" colab_type="code" colab={}
| ActorCriticLunarLander.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def merge(a, p, q, r):
n1 = q - p + 1
n2 = r - q
L, R = [], []
for i in range(0, n1):
L.append(a[i + p])
for j in range(0, n2):
R.append(a[j + q + 1])
i, j = 0, 0
for k in range(p, r):
if i <= len(L) and (j >= len(R) or L[i] <= R[j]):
a[k] = L[i]
i += 1
else:
a[k] = R[j]
j += 1
# +
import math
def mergeSort(a, p, r):
if p < r:
mid = math.floor((p + r)/2)
mergeSort(a, p, mid)
mergeSort(a, mid + 1, r)
merge(a, p, mid, r)
# -
a = [3, 4, 1, 2]
mergeSort(a, 0, 3)
print(a)
a = [3, 4, 1, 2]
merge(a, 0, 1, 3)
print(a)
| python/divideAndConquer/Merge Sort practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# this makes sure that I can recover the PCAed vgg features.
import os
from itertools import product
import numpy as np
import h5py
from tang_jcompneuro.io import load_split_dataset, load_split_dataset_idx, load_split_dataset_pretrained_pca_params
from tang_jcompneuro import dir_dictionary
from tang_jcompneuro.cnn_pretrained import blob_corresponding_info
blob_names = list(blob_corresponding_info['vgg19'].keys())
# +
# dataset = load_split_dataset('MkA_Shape', 'all', True, slice(None), last_val=False, suffix='vgg19/legacy/conv3_1')
# +
# len(dataset)
# +
# data_idx = load_split_dataset_idx('MkA_Shape', 'all', True, last_val=False)
# +
# len(data_idx)
# +
# print(dataset[0].shape)
# print(dataset[1].shape)
# print(dataset[2].shape)
# print(dataset[3].shape)
# print(dataset[4].shape)
# print(dataset[5].shape)
# +
# print(data_idx[0].shape)
# print(data_idx[1].shape)
# print(data_idx[2].shape)
# +
# load original VGG features
# -
def load_original_features(layer_name):
file_in = os.path.join(dir_dictionary['features'], 'cnn_feature_extraction.hdf5')
layer_idx = blob_names.index(layer_name)
key_to_extract = '/'.join(['vgg19', 'legacy', str(layer_idx)])
with h5py.File(file_in, 'r') as f:
feature_this = f[f'Shape_9500/{key_to_extract}'][...].astype(np.float64, copy=False)
print(feature_this.shape)
return feature_this
# +
# load_original_features('conv3_1')
# -
# +
# pca_mean.shape
# +
# pca_components.shape
# -
def check_one_layer(layer_name, seed):
suffix=f'vgg19/legacy/{layer_name}'
dataset = load_split_dataset('MkA_Shape', 'all', True, slice(None), last_val=False, suffix=suffix, seed=seed)
dataset = (dataset[0], dataset[2], dataset[4])
data_idx = load_split_dataset_idx('MkA_Shape', 'all', True, last_val=False, seed=seed)
assert len(data_idx) == len(dataset) == 3
pca_mean, pca_components = load_split_dataset_pretrained_pca_params('MkA_Shape', 'all', suffix=suffix, seed=seed)
features_original = load_original_features(layer_name)
features_original = features_original.reshape(features_original.shape[0], -1)
# ok. let's check
for (dataset_out_this, dataset_idx_this) in zip(dataset, data_idx):
print(dataset_idx_this.shape)
assert (dataset_out_this.shape[0],) == dataset_idx_this.shape
dataset_out_this_debug = (features_original[dataset_idx_this] - pca_mean)@pca_components.T
assert dataset_out_this_debug.shape == dataset_out_this.shape
assert np.allclose(dataset_out_this_debug, dataset_out_this)
print(np.array_equal(dataset_out_this_debug, dataset_out_this))
def check_all():
for layer_name, seed in product(['conv2_1', 'conv3_1', 'conv4_1'], range(2)):
print(layer_name, seed)
check_one_layer(layer_name, seed)
check_all()
| results_ipynb/debug/cnn_pretrained/check_pca.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import firebase_admin
from firebase_admin import db
cred = firebase_admin.credentials.Certificate('/Users/sam/Downloads/flightlog-235e4-firebase-adminsdk-uvrwi-d8510b3147.json')
flightLog = firebase_admin.initialize_app(cred,{'databaseURL':'https://flightlog-235e4.firebaseio.com/'})
# +
import csv
with open("data/ParaglidingLog.csv") as csvfile:
flights = list(csv.DictReader(csvfile, delimiter=',', quotechar='"'))
from datetime import datetime, date
epoch = datetime.utcfromtimestamp(0)
def unix_time_millis(dt):
return (dt - epoch).total_seconds() * 1000.0
flights = [{}]+flights
for flight in flights:
if '' in flight.keys():
# print flight['date'],flight['start_time'],flight['end_time']
# print flight
#flight['id']=flight['flight']
flight['flight']=int(flight['flight'])
try:
d = datetime.strptime(flight['date'],'%m/%d/%y')
except:
d = datetime.strptime(flight['date'],'%Y-%m-%d')
flight['date'] = d.isoformat()[:10]
if flight['start_time']:
try:
launchTime = datetime.strptime(flight['date']+' '+flight['start_time'],'%m/%d/%y %I:%M %p')
except:
launchTime = datetime.strptime(flight['date']+' '+flight['start_time'],'%Y-%m-%d %I:%M %p')
flight['launch_time_iso'] = launchTime.isoformat()
# flight['launch_time_epoch'] = unix_time_millis(launchTime)
else:
flight['launch_time_iso'] = None
# flight['launch_time_epoch'] = None
if flight['end_time']:
try:
landingTime = datetime.strptime(flight['date']+' '+flight['end_time'],'%m/%d/%y %I:%M %p')
except:
landingTime = datetime.strptime(flight['date']+' '+flight['end_time'],'%Y-%m-%d %I:%M %p')
flight['landing_time_iso'] = landingTime.isoformat()
# flight['landing_time_epoch'] = unix_time_millis(landingTime)
else:
flight['landing_time_iso'] = None
# flight['landing_time_epoch'] = None
flight['duration_total_minutes']=int(flight['duration_hours'] or 0)*60+int(flight['duration_minutes'] or 0)
flight['launch_altitude']=int(flight['launch_altitude'] or 0) or None
flight['landing_altitude']=int(flight['landing_altitude'] or 0) or None
flight['max_altitude']=int(flight['max_altitude'] or 0) or None
flight['altitude_gain']=int(flight['altitude_gain'] or 0) or None
flight['vertical_drop']=int(flight['vertical_drop'] or 0) or None
flight['xc_miles']=float(flight['xc_miles'] or 0) or None
flight['total_airtime']=float(flight['total_airtime'])
flight['updated_epoch'] = int(unix_time_millis(datetime.utcnow()))
del flight['']
del flight['start_time']
del flight['end_time']
del flight['duration_hours']
del flight['duration_minutes']
# print flight['date'],flight['launch_time_iso'],flight['landing_time_iso'],'\n'
print flight
# -
myFlights = db.reference('/ALdWWkKJGGZZDJSvtdHpgKFxdLt2/flight_log')
# +
#myFlights.set(flights)
# +
#myFlights.set([])
# -
myFlights.set([])
for flight in flights:
newFlight = myFlights.push()
newFlight.set(flight)
totalMins=0
lastTotal=0
for flight in flights[21:]:
totalMins+=flight['duration_total_minutes']
flight['total_airtime_computed']=totalMins
if abs(flight['duration_total_minutes'] -(flight['total_airtime']*60-lastTotal)) > .5:
print flight['flight'], flight['duration_total_minutes'], flight['total_airtime'], flight['total_airtime']*60, flight['total_airtime_computed']
lastTotal = flight['total_airtime']*60
# if flight['landing_time_epoch']:
# flight['duration_total_minutes_computed']=int((flight['landing_time_epoch']-flight['launch_time_epoch'])/60000)
# if flight['duration_total_minutes'] != flight['duration_total_minutes_computed']:
# print flight['flight'],"check time", flight['launch_time_iso'], flight['landing_time_iso'], flight['duration_total_minutes'], flight['duration_total_minutes_computed'], flight['total_airtime'], flight['total_airtime_computed']/60.
# if flight['total_airtime'] != flight['total_airtime_computed']/60.:
# print flight['flight'], flight['total_airtime'], flight['total_airtime']*60, flight['total_airtime_computed']
| DatabaseImport.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Django Shell-Plus
# language: python
# name: django_extensions
# ---
# # <NAME> (Okay)
# +
import ipywidgets as widgets
from IPython.display import display
import esper.identity_clusters
from esper.identity_clusters import identity_clustering_workflow,_manual_recluster,visualization_workflow
shootings = [
('<NAME>', 'Chattanooga', 'Jul 16, 2015'),
('<NAME>', 'Umpqua Community College', 'Oct 1, 2015'),
('<NAME>', 'Colorado Springs - Planned Parenthood', 'Nov 27, 2015'),
('<NAME>', 'San Bernardino', 'Dec 2, 2015'),
('<NAME>', 'San Bernardino', 'Dec 2, 2015'),
('<NAME>', 'Charleston Shurch', 'Jun 17, 2015'),
('<NAME>', 'Orlando Nightclub', 'Jun 12, 2016'),
('<NAME>', 'Dallas Police', 'Jul 7-8, 2016'),
('<NAME>', 'Baton Rouge Police', 'Jul 17, 2016'),
('<NAME>', 'Ft. Lauderdale Airport', 'Jan 6, 2017'),
('<NAME>', 'Lincoln County', 'May 28, 2017'),
('<NAME>', 'Las Vegas', 'Oct 1, 2017'),
('<NAME>', 'San Antonio Church', 'Nov 5, 2017')
]
orm_set = { x.name for x in Identity.objects.filter(name__in=[s[0].lower() for s in shootings]) }
for s in shootings:
assert s[0].lower() in orm_set, '{} is not in the database'.format(s)
identity_clustering_workflow('<NAME>','Nov 5, 2017', True)
# -
| app/notebooks/michaela_notebooks/Analysis of Shooters and Victims/Devin Patrick Kelley (Shooter).ipynb |
# +
"""
Checking Imported Data II - How Big is the Data?
"""
import numpy as np
import pandas as pd
wine_reviews = pd.read_csv('../../winemag-data-130k.csv')
# Print the number of rows and columns in wine_reviews.
print(wine_reviews.shape) # (rows, columns) is (129971, 13)
# Print the number of elements in wine_reviews.
print(wine_reviews.size) # 1689623
| pset_pandas1_wine_reviews/check_imported_data/solutions/nb/p2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] Collapsed="false"
# <img src='./img/EU-Copernicus-EUM_3Logos.png' alt='Logo EU Copernicus EUMETSAT' align='right' width='50%'></img>
# <br>
# + [markdown] Collapsed="false"
# <br>
# + [markdown] Collapsed="false"
# <a href="./index_ltpy.ipynb"><< Index</a><span style="float:right;"><a href="./12_ltpy_WEkEO_harmonized_data_access_api.ipynb">12 - WEkEO Harmonized Data Access API >></a></span>
# + [markdown] Collapsed="false"
# # 1.1 Atmospheric composition data - Overview and data access
# + [markdown] Collapsed="false"
# This module gives an overview of the following atmospheric composition data services:
# * [EUMETSAT AC SAF - The EUMETSAT Satellite Application Facility on Atmospheric Composition Monitoring](#ac_saf)
# * [Copernicus Sentinel-5 Precursor (Sentinel-5P)](#sentinel_5p)
# * [Copernicus Sentinel-3](#sentinel3)
# * [Copernicus Atmosphere Monitoring Service (CAMS)](#cams)
# + [markdown] Collapsed="false"
# <br>
# + [markdown] Collapsed="false"
# ## <a id="ac_saf"></a>EUMETSAT AC SAF - The EUMETSAT Satellite Application Facility on Atmospheric Composition Monitoring
# -
# <span style=float:left><img src='./img/ac_saf_logo.png' alt='Logo EU Copernicus EUMETSAT' align='left' width='90%'></img></span>
#
# The [EUMETSAT Satellite Application Facility on Atmospheric Composition Monitoring (EUMETSAT AC SAF)](http://acsaf.org/) is one of eight EUMETSAT Satellite Application Facilities (SAFs). <br>
#
# SAFs generate and disseminate operational EUMETSAT products and services and are an integral part of the distributed EUMETSAT Application Ground Segment.
#
# AC SAF processes data on ozone, other trace gases, aerosols and ultraviolet data, obtained from satellite instrumentation.
# + [markdown] Collapsed="false"
# <br>
# + [markdown] Collapsed="false"
# ### Available AC SAF products
# + [markdown] Collapsed="false"
# AC-SAF offers three different product types: <br>
#
# |<font size='+0.2'><center>[Near real-time products](#nrt)</center></font> | <font size='+0.2'><center>[Offline products](#offline)</center></font> | <font size='+0.2'><center>[Data records](#records)</center></font> |
# |-----|-----|------|
# <img src='./img/nrt_no2_example.png' alt='Near-real time product - NO2' align='middle' width='60%'></img>|<img src='./img/offline_ozone_example.png' alt='Logo EU Copernicus EUMETSAT' align='middle' width='60%'></img>|<img src='./img/ac_saf_level3.png' alt='Logo EU Copernicus EUMETSAT' align='middle' width='100%'></img>|
#
# <br>
# Near real-time and offline products are often refered as Level 2 data. Data records are refered as Level 3 data.
#
# AC SAF products are sensed from two instruments onboard the Metop satellites:
# * [Global Ozone Monitoring Experiment-2 (GOME-2) instrument](https://acsaf.org/gome-2.html) <br>
# GOME-2 can measure a range of atmospheric trace constituents, with the emphasis on global ozone distributions. Furthermore, cloud properties and intensities of ultraviolet radiation are retrieved. These data are crucial for monitoring the atmospheric composition and the detection of pollutants. <br>
#
# * [Infrared Atmospheric Sounding Interferometer (IASI) instrument](https://acsaf.org/iasi.html)
#
# The [Metop satellites](https://acsaf.org/metop.html) is a series of three satellites that were launched in October 2006 (Metop-A), September 2012 (Metop-B) and November 2018 (Metop-C) respectively.
#
# All AC SAF products are disseminated under the [AC SAF Data policy](https://acsaf.org/data_policy.html).
# + [markdown] Collapsed="false"
# <br>
# + [markdown] Collapsed="false"
# #### <a id="nrt"></a>Near-real time (NRT) products
# + [markdown] Collapsed="false"
# NRT products are Level 2 products and are available to users in 3 hours from sensing at the latest and available for the past two months. NRT products are disseminated in HDF5 format.
#
# | <img width=100>NRT Product type name</img> | Description | Unit | <img width=80>Satellite</img> | Instrument |
# | ---- | ----- | ----- | ---- | -----|
# | Total Ozone (O<sub>3</sub>) column | NRT total ozone column product provided information about vertical column densities of ozone in the atmosphere | Dobson Units (DU) | Metop-A<br>Metop-B | GOME-2 |
# | Total and tropospheric NO<sub>2</sub> columns | NRT total and tropospheric NO2 column products provide information about vertical column densities of nitrogen dioxide in the atmosphere. | molecules/cm2 | Metop-A<br>Metop-B | GOME-2 |
# | Total SO<sub>2</sub> column | NRT total SO2 column product provides information about vertical column densities of the sulfur dioxide in the atmosphere. | Dobson Units (DU) | Metop-A<br>Metop-B | GOME-2
# | Total HCHO column | NRT HCHO column product provides information about vertical column densities of formaldehyde in the atmosphere. | molecules/cm2 | Metop-A<br>Metop-B | GOME-2 |
# | High-resolution vertical ozone profile | NRT high-resolution vertical ozone profile product provides an ozone profile from the GOME-2 nadir scanning mode. | Partial ozone columns in Dobson Units in 40 layers from the surface up to 0.001 hPa| Metop-A<br>Metop-B | GOME-2 |
# | Global tropospheric ozone column | The global tropospheric ozone column product provides information about vertical column densities of ozone in the troposphere, <br>from the surface to the tropopause and from the surface to 500 hPa (∼5km). | Dobson Units (DU) | Metop-A<br>Metop-B | GOME-2 |
# + [markdown] Collapsed="false"
# <br>
# + [markdown] Collapsed="false"
# #### <a id="offline"></a>Offline products
# + [markdown] Collapsed="false"
# Offline products are Level 2 products and are available to users in 15 days of sensing. Typical delay is 2-3 days. Offline products are disseminated in HDF5 format.
#
# | Offline Product type name | Description | Unit | <img width=80>Satellite</img> | Instrument | <img width=150px>Time period</img> |
# | ---- | ----- | ----- | ---- | -----|----|
# | Total Ozone (O<sub>3</sub>) column | Offline total ozone column product provided information about vertical column densities of ozone in the atmosphere | Dobson Units (DU) | Metop-A<br>Metop-B | GOME-2 | 1 Jan 2008 - almost NRT<br>13 Dec 2012 - almost NRT |
# | Total and tropospheric NO<sub>2</sub> columns | Offline total and tropospheric NO2 column products provide information about vertical column densities of nitrogen dioxide in the atmosphere. | molecules/cm2 | Metop-A<br>Metop-B | GOME-2 | 1 Jan 2008 - almost NRT<br>13 Dec 2012 - almost NRT |
# | Total SO<sub>2</sub> column | Offline total SO2 column product provides information about vertical column densities of the sulfur dioxide in the atmosphere. | Dobson Units (DU) | Metop-A<br>Metop-B | GOME-2 | 1 Jan 2008 - almost NRT<br>13 Dec 2012 - almost NRT |
# | Total HCHO column | Offline HCHO column product provides information about vertical column densities of formaldehyde in the atmosphere. | molecules/cm2 | Metop-A<br>Metop-B | GOME-2 | 1 Jan 2008 - almost NRT<br>13 Dec 2012 - almost NRT |
# | High-resolution vertical ozone profile | Offline high-resolution vertical ozone profile product provides an ozone profile from the GOME-2 nadir scanning mode. | Partial ozone columns in Dobson Units in 40 layers from the surface up to 0.001 hPa| Metop-A<br>Metop-B | GOME-2 | 1 Jan 2008 - almost NRT<br>13 Dec 2012 - almost NRT |
# | Global tropospheric ozone column | The offline global tropospheric ozone column product provides information about vertical column densities of ozone in the troposphere, from the surface to the tropopause and and from the surface to 500 hPa (∼5km). | Dobson Units (DU) | Metop-A<br>Metop-B | GOME-2 | 1 Jan 2008 - almost NRT<br>13 Dec 2012 - almost NRT |
# + [markdown] Collapsed="false"
# <br>
# + [markdown] Collapsed="false"
# #### <a id="records"></a>Data records
# + [markdown] Collapsed="false"
# Data records are reprocessed, gridded Level 3 data. Data records are monthly aggregated products, regridded on a regular latitude-longitude grid. Data records are disseminated in NetCDF format.
#
# | Data record name | Description | Unit | <img width=80>Satellite</img> | Instrument | <img width=150>Time period</img> |
# | ---- | ----- | ----- | ---- | -----|----|
# | Reprocessed **tropospheric O<sub>3</sub>** column data record for the Tropics | Tropospheric ozone column data record for the Tropics provides long-term information <br>about vertical densities of ozone in the atmosphere for the tropics. | Dobson Units (DU) | Metop-A<br>Metop-B | GOME-2 | Jan 2007- Dec 2018<br>Jan 2013- Jun 2019 |
# | Reprocessed **total column and tropospheric NO<sub>2</sub>** data record | Total and tropospheric NO2 column data record provides long-term information about vertical column densities of nitrogen dioxide in the atmosphere. | molecules/cm2 | Metop-A<br>Metop-B | GOME-2 | Jan 2007 - Nov 2017<br>Jan 2013 - Nov 2017 |
# | Reprocessed **total H<sub>2</sub>O column** data record | Total H2O column data record provides long-term information about vertical column densities of water vapour in the atmosphere. | kg/m2 | Metop-A<br>Metop-B | GOME-2 | Jan 2007 - Nov 2017<br>Jan 2013 - Nov 2017 |
# + [markdown] Collapsed="false"
# <br>
# + [markdown] Collapsed="false"
# ### <a id="ac_saf_access"></a>How to access AC SAF products
# + [markdown] Collapsed="false"
# AC SAF products can be accessed via different dissemination channels. There are channels where Level 2 and Level 3 are available for download. Other sources allow to browse through images and maps of the data. This is useful to see for what dates e.g. Level 2 data were sensed.
# + [markdown] Collapsed="false"
# #### DLR ftp server
# + [markdown] Collapsed="false"
# All near-real time, offline and reprocessed total column data are available at [DLR's ATMOS FTP-server](https://atmos.eoc.dlr.de/products/). Accessing data is a two step process:
# 1. [Register](https://acsaf.org/registration_form.html) as a user of AC SAF products
# 2. [Log in](https://atmos.eoc.dlr.de/products/)( (with the user name and password that is emailed to you after registration)
#
# Once logged in, you find data folders for GOME-2 products from Metop-A in the directory *'gome2a/'* and GOME-2 products from Metop-B in the directory: *'gome2b/'* respectively. In each GOME-2 directory, you find the following sub-directories: <br>
# * **`near_real_time/`**,
# * **`offline/`**, and
# * **`level3/`**.
#
# <br>
#
#
# <div style='text-align:center;'>
# <figure><img src='./img/dlr_ftp_directory.png' width='50%'/>
# <figcaption><i>Example of the directory structure of DLR's ATMOS FTP-server</i></figcaption>
# </figure>
# </div>
#
# + [markdown] Collapsed="false"
# <br>
# + [markdown] Collapsed="false"
# #### EUMETSAT Data Centre
# + [markdown] Collapsed="false"
# The EUMETSAT Data Centre provides a long-term archive of data and generated products from EUMETSAT, which can be ordered online. Ordering data is a two step process:
# 1. [Create an account](https://eoportal.eumetsat.int/userMgmt/register.faces) at the EUMETSAT Earth Observation Portal
# 2. [Log in](https://eoportal.eumetsat.int/userMgmt/login.faces) (with the user name and password that is emailed to you after registration)
#
# Once succesfully logged in, go to (1) Data Centre. You will be re-directed to (2) the User Services Client. Type in *'GOME'* as search term and you can get a list of all available GOME-2 products.
# + [markdown] Collapsed="false"
#
#
# <div style='text-align:center;'>
# <figure><img src='./img/eumetsat_data_centre.png' width='50%' />
# <figcaption><i>Example of the directory structure of EUMETSAT's Data Centre</i></figcaption>
# </figure>
# </div>
# + [markdown] Collapsed="false"
# <br>
# + [markdown] Collapsed="false"
# #### Web-based services
# + [markdown] Collapsed="false"
# There are two web-based services, [DLR's ATMOS webserver](https://atmos.eoc.dlr.de/app/missions/gome2) and the [TEMIS service by KNMI](http://temis.nl/index.php) that offer access to GOME-2/MetOp browse products. These services are helpful to see the availability of data for specific days, especially for AC SAF Level-2 parameters.
# + [markdown] Collapsed="false"
# <br>
# + [markdown] Collapsed="false"
# | <font size='+0.2'>[DLR's ATMOS webserver](https://atmos.eoc.dlr.de/app/missions/gome2)</font> | <font size='+0.2'>[TEMIS - Tropospheric Emission Monitoring Internet Service](http://temis.nl/index.php)</font> |
# | - | - |
# | <br>ATMOS (Atmospheric ParameTers Measured by in-Orbit Spectrosocopy is a webserver operated by DLR's Remote Sensing Technology Institute (IMF). The webserver provides access to browse products from GOME-2/Metop Products, both in NRT and offline mode. <br><br> | <br>TEMIS is a web-based service to browse and download atmospheric satellite data products maintained by KNMI. The data products consist mainly of tropospheric trace gases and aerosol concentrations, but also UV products, cloud information and surface albedo climatologies are provided. <br><br> |
# | <center><img src='./img/atmos_service.png' width='70%'></img></center> | <center><img src='./img/temis_service.png' width='70%'></img></center> |
# + [markdown] Collapsed="false"
# <br>
# + [markdown] Collapsed="false"
# ## <a id="sentinel_5p"></a>Copernicus Sentinel-5 Precursor (Sentinel-5P)
# + [markdown] Collapsed="false"
# [Sentinel-5 Precursor (Sentinel-5P)](https://sentinels.copernicus.eu/web/sentinel/missions/sentinel-5p) is the first Copernicus mission dedicated to monitoring our atmosphere. The satellite carries the state-of-the-art TROPOMI instrument to map a multitude of trace gases.
#
# Sentinel-5P was developed to reduce data gaps between the ENVISAT satellite - in particular the Sciamachy instrument - and the launch of Sentinel-5, and to complement GOME-2 on MetOp. In the future, both the geostationary Sentinel-4 and polar-orbiting Sentinel-5 missions will monitor the composition of the atmosphere for Copernicus Atmosphere Services. Both missions will be carried on meteorological satellites operated by [EUMETSAT](https://eumetsat.int).
# + [markdown] Collapsed="false"
# ### Available data products and trace gas information
# + [markdown] Collapsed="false"
# <span style=float:right><img src='./img/sentinel_5p_data_products.jpg' alt='Sentinel-5p data prodcuts' align='right' width='90%'></img></span>
# Data products from Sentinel-5P’s Tropomi instrument are distributed to users at two levels:
# * `Level-1B`: provides geo-located and radiometrically corrected top of the atmosphere Earth radiances in all spectral bands, as well as solar irradiances.
# * `Level-2`: provides atmospheric geophysical parameters.
#
# `Level-2` products are disseminated within three hours after sensing. This `near-real-time`(NRT) services disseminates the following products:
# * `Ozone`
# * `Sulphur dioxide`
# * `Nitrogen dioxide`
# * `Formaldehyde`
# * `Carbon monoxide`
# * `Vertical profiles of ozone` and
# * `Cloud / Aerosol distributions`
#
# `Level-1B` products are disseminated within 12 hours after sensing.
#
# `Methane`, `tropospheric ozone` and `corrected total nitrogen dioxide columns` are available withing 5 days after sensing.
#
#
# + [markdown] Collapsed="false"
# <br>
# + [markdown] Collapsed="false"
# ### <a id="sentinel5p_access"></a>How to access Sentinel-5P data
# + [markdown] Collapsed="false"
# Sentinel-5P data can be accessed via different dissemination channels. The data is accessible via the `Copernicus Open Access Hub` and `EUMETSAT's EUMETCast`.
# + [markdown] Collapsed="false"
# #### Copernicus Open Access Hub
# + [markdown] Collapsed="false"
# Sentinel-5P data is available for browsing and downloading via the [Copernicus Open Access Hub](https://scihub.copernicus.eu/). The Copernicus Open Access Hub provides complete, free and open access to Sentinel-1, Sentinel-2, Sentinel-3 and Sentinel-5P data.
# + [markdown] Collapsed="false"
#
# <div style='text-align:center;'>
# <figure><img src='./img/open_access_hub.png' alt='Sentinel-5p data products' align='middle' width='50%'/>
# <figcaption><i>Interface of the Copernicus Open Access Hub and the Sentinel-5P Pre-Operations Data Hub</i></figcaption>
# </figure>
# </div>
# + [markdown] Collapsed="false"
# #### EUMETSAT's EUMETCast
# + [markdown] Collapsed="false"
# Since August 2019, Sentinel-5p `Level 1B` and `Level 2` are as well available on EUMETSAT's EUMETCast:
# * **Level 1B** products will be distributed on EUMETCast Terrestrial
# * **Level 2** products are distributed on EUMETCast Europe, High Volume Service Transponder 2 (HVS-2)
#
# Sentinel-5P data on EUMETCast can be accessed via [EUMETSAT's Earth Observation Portal (EOP)](https://eoportal.eumetsat.int/userMgmt/login.faces).
# -
# #### TEMIS
# [TEMIS - Tropospheric Emission Monitoring Internet Service](http://temis.nl/airpollution/no2.html) provides access to selected Sentinel-5P parameters, e.g. `NO`<sub>2</sub>.
# + [markdown] Collapsed="false"
# <br>
# -
# ## <a id='sentinel3'></a>Copernicus Sentinel-3 - Ocean and Land Colour (OLCI)
# <span style=float:right><img src='./img/sentinel3.png' alt='Sentinel-5p data prodcuts' align='right' width='90%'></img></span>
# The Sentinel-3 is the Copernicus mission to monitor and measure sea surface topography, sea and land surface temperature and ocean and land surface.
#
# The Sentinel-3 mission carries five different instruments aboard the satellites: and offers four differnt data product types:
# - [Ocean and Land Colour Instrument (OLCI)](https://sentinel.esa.int/web/sentinel/missions/sentinel-3/data-products/olci)
# - [Sea and Land Surface Temperature Radiometer (SLSTR)](https://sentinel.esa.int/web/sentinel/missions/sentinel-3/data-products/slstr)
# - [Synergy](https://sentinel.esa.int/web/sentinel/missions/sentinel-3/data-products/synergy), and
# - [Altimetry](https://sentinel.esa.int/web/sentinel/missions/sentinel-3/data-products/altimetry).
#
# The Sentinel-3 OLCI mission supports maritime monitoring, land mapping and monitoring, atmospheric monitoring and climate change monitoring.
# ### Available OLCI data products
# OLCI product types are divided in three main categories:
#
# - #### Level-1B products
# Two different Level-1B products can be obtained:
# - OL_1_EFR - output during EO processing mode for Full Resolution
# - OL_1_ERR -output during EO processing mode for Reduced Resolution
#
# The Level-1B products in EO processing mode contain calibrated, ortho-geolocated and spatially re-sampled Top Of Atmosphere (TOA) radiances for [21 OLCI spectral bands](https://sentinel.esa.int/web/sentinel/user-guides/sentinel-3-olci/resolutions/radiometric). In Full Resolution products (i.e. at native instrument spatial resolution), these parameters are provided for each re-gridded pixel on the product image and for each removed pixel. In Reduced Resolution products (i.e. at a resolution four times coarser), the parameters are only provided on the product grid.
#
# - #### Level-2 Land products and Water products
# The level-2 land product provides land and atmospheric geophysical parameters. The Level-2 water product provides water and atmospheric geophysical parameters. All products are computed for full and reduced resolution:
# - OL_2_LFR - Land Full Resolution
# - OL_2_LRR - Land Reduced Resolution
#
#
# There are two timeframes for the delivery of the products:
# - **Near-Real-Time (NRT)**: delivered to the users less than three hours after acquisition of the data by the sensor
# - **Non-Time Critical (NTC)**: delivered no later than one month after acquisition or from long-term archives. Typically, the product is available within 24 or 48 hours.
#
# The data is disseminated in .zip archive containing free-standing `NetCDF4` product files.
#
# ### How to access Sentinel-3 data
# Sentinel-3 data can be accessed via different dissemination channels. The data is accessible via the `Copernicus Open Access Hub` and `WEkEO's Harmonized Data Access API`.
# #### Copernicus Open Access Hub
# Sentinel-3 data is available for browsing and downloading via the Copernicus Open Access Hub. The Copernicus Open Access Hub provides complete, free and open access to Sentinel-1, Sentinel-2, Sentinel-3 and Sentinel-5P data. See the an example of the Copernicus Open Access Hub interface [here](#sentinel5p_access).
# #### WEkEO's Harmonized Data Access API
# <span style=float:left><img src='./img/wekeo_logo2.png' alt='Logo WEkEO' align='center' width='90%'></img></span>
# [WEkEO](https://www.wekeo.eu/) is the EU Copernicus DIAS (Data and Information Access Service) reference service for environmental data, virtual processing environments and skilled user support.
#
# WEkEO offers access to a variety of data, including different parameters sensored from Sentinel-1, Sentinel-2 and Sentinel-3. It further offers access to climate reanalysis and seasonal forecast data.
#
# The [Harmonized Data Access (HDA) API](https://www.wekeo.eu/documentation/using_jupyter_notebooks), a REST interface, allows users to subset and download datasets from WEkEO.
#
# Please see [here](./12_ltpy_WEkEO_harmonized_access_api.ipynb) a practical example how you can retrieve Sentinel-3 data from WEkEO using the Harmonized Data Access API.
# <br>
# <br>
# ## <a id="cams"></a>Copernicus Atmosphere Monitoring Service (CAMS)
# <span style=float:left><img src='./img/cams_logo_2.png' alt='Copernicus Atmosphere Monitoring Service' align='left' width='95%'></img></span>
#
# [The Copernicus Atmosphere Monitoring Service (CAMS)](https://atmosphere.copernicus.eu/) provides consistent and quality-controlled information related to `air pollution and health`, `solar energy`, `greenhouse gases` and `climate forcing`, everywhere in the world.
#
# CAMS is one of six services that form [Copernicus, the European Union's Earth observation programme](https://www.copernicus.eu/en).
#
# CAMS is implemented by the [European Centre for Medium-Range Weather Forecasts (ECMWF)](http://ecmwf.int/) on behalf of the European Commission. ECMWF is an independent intergovernmental organisation supported by 34 states. It is both a research institute and a 24/7 operational service, producing and disseminating numerical weather predictions to its member states.
# <br>
# ### Available data products
# CAMS offers four different data product types:
#
# |<font size='+0.2'><center>[CAMS Global <br>Reanalysis](#cams_reanalysis)</center></font></img> | <font size='+0.2'><center>[CAMS Global Analyses <br>and Forecasts](#cams_an_fc)</center></font> | <img width=30><font size='+0.2'><center>[CAMS Global Fire Assimilation System (GFAS)](#cams_gfas)</center></font></img> | <img width=30><font size='+0.2'><center>[CAMS Greenhouse Gases Flux Inversions](#cams_greenhouse_flux)</center></font></img> |
# |-----|-----|------|------|
# <img src='./img/cams_reanalysis.png' alt='CAMS reanalysis' align='middle' width='100%'></img>|<img src='./img/cams_forecast.png' alt='CAMS Forecast' align='middle' width='100%'></img>|<img src='./img/cams_gfas.png' alt='CAMS GFAS' align='middle' width='100%'></img>|<img src='./img/cams_greenhouse_fluxes.png' alt='CAMS greenhous flux inversions' align='middle' width='100%'></img>|
#
# #### <a id="cams_reanalysis"></a>CAMS Global Reanalysis
# CAMS reanalysis data set provides consistent information on aerosols and reactive gases from 2003 to 2017. CAMS global reanalysis dataset has a global horizontal resolution of approximately 80 km and a refined temporal resolution of 3 hours. CAMS reanalysis are available in GRIB and NetCDF format.
#
# | Parameter family | Time period | <img width=80>Spatial resolution</img> | Temporal resolution |
# | ---- | ----- | ----- | -----|
# | [CAMS global reanalysis of total aerosol optical depth<br> at multiple wavelengths](https://atmosphere.copernicus.eu/catalogue#/product/urn:x-wmo:md:int.ecmwf::copernicus:cams:prod:rean:black-carbon-aod_dust-aod_organic-carbon-aod_sea-salt-aod_sulphate-aod_total-aod_warning_multiple_species:pid469) | 2003-2017 | ~80km | 3-hourly |
# | [CAMS global reanalysis of aerosol concentrations](https://atmosphere.copernicus.eu/catalogue#/product/urn:x-wmo:md:int.ecmwf::copernicus:cams:prod:rean:black-carbon-concentration_dust-concentration_organic-carbon-concentration_pm1_pm10_pm2.5_sea-salt-concentration_sulfates-concentration_warning_multiple_species:pid467) | 2003-2017 | ~80km | 3-hourly |
# | [CAMS global reanalysis chemical species](https://atmosphere.copernicus.eu/catalogue#/product/urn:x-wmo:md:int.ecmwf::copernicus:cams:prod:rean:ald2_c10h16_c2h4_c2h5oh_c2h6_c2o3_c3h6_c3h8_c5h8_ch3coch3_ch3cocho_ch3o2_ch3oh_ch3ooh_ch4_co_dms_h2o2_hcho_hcooh_hno3_ho2_ho2no2_mcooh_msa_n2o5_nh2_nh3_nh4_no_no2_no3_no3_a_nox_o3_oh_ole_onit_pan_par_pb_rooh_ror_ra_so2_so4_warning_multiple_species:pid468) | 2003-2017 | ~80km | 3-hourly |
# #### <a id="cams_an_fc"></a>CAMS Global analyses and forecasts
# CAMS daily global analyses and forecast data set provides daily global forecasts of atmospheric composition parameters up to five days in advance. CAMS analyses and forecast data are available in GRIB and NetCDF format.
#
# The forecast consists of 56 reactive trace gases in the troposphere, stratospheric ozone and five different types of aersol (desert dust, sea salt, organic matter, black carbon and sulphate).
#
# | Parameter family | Time period | <img width=80>Spatial resolution</img> | Forecast step |
# | ---- | ----- | ----- | -----|
# | CAMS global forecasts of aerosol optical depths | Jul 2012- 5 days in advance | ~40km | 3-hour |
# | CAMS global forecasts of aerosols | Jul 2012 - 5 days in advance | ~40km | 3-hour |
# | CAMS global forecasts of chemical species | Jul 2012- 5 days in advance | ~40km | 3-hour |
# | CAMS global forecasts of greenhouse gases | Jul 2012- 5 days in advance | ~9km | 3-hour |
# #### <a id="cams_gfas"></a>CAMS Global Fire Assimiliation System (GFAS)
# CAMS GFAS assimilated fire radiative power (FRP) observations from satellite-based sensors to produce daily estimates of wildfire and biomass burning emissions. The GFAS output includes spatially gridded Fire Radiative Power (FRP), dry matter burnt and biomass burning emissions for a large set of chemical, greenhouse gas and aerosol species. CAMS GFAS data are available in GRIB and NetCDF data.
#
# A full list of CAMS GFAS parameters can be found in the [CAMS Global Fire Assimilation System (GFAS) data documentation](https://atmosphere.copernicus.eu/sites/default/files/2018-05/CAMS%20%20Global%20Fire%20Assimilation%20System%20%28GFAS%29%20data%20documentation.pdf).
#
# | Parameter family | Time period | <img width=80>Spatial resolution</img> | Temporal resolution |
# | ---- | ----- | ----- | ---- |
# | CAMS GFAS analysis surface parameters | Jan 2003 - present | ~11km | daily |
# | CAMS GFAS gridded satellite parameters | Jan 2003 - present | ~11km | daily |
# #### <a id="cams_greenhouse_flux"></a>CAMS Greenhouse Gases Flux Inversions
# CAMS Greenhouse Gases Flux Inversion reanalysis describes the variations, in space and in time, of the surface sources and sinks (fluxes) of the three major greenhouse gases that are directly affected by human activities: `carbon dioxide (CO2)`, `methane (CH4)` and `nitrous oxide (N2O)`. CAMS Greenhouse Gases Flux data is available in GRIB and NetCDF format.
#
# | Parameter | Time period | <img width=80>Spatial resolution</img> | Frequency | Quantity |
# | ---- | ----- | ----- | ---- | -----|
# | Carbon Dioxide | Jan 1979 - Dec 2018 | ??? | 3 hourly<br>Monthly average | Concentration<br>Surface flux<br> Total colum |
# | Methane | Jan 1990 - Dec 2017 | ??? | 6-hourly<br>Daily average<br>Monthly average | Concentration<br>Surface flux<br>Total column
# | Nitrous Oxide | Jan 1995 - Dec 2017 | ???| 3-hourly<br>Monthly average | Concentration<br>Surface flux |
# <br>
# ### <a id="cams_access"></a>How to access CAMS data
# CAMS data can be accessed in two different ways: `ECMWF data archive` and `CAMS data catalogue of data visualizations`. A more detailed description of the different data access platforms can be found [here](https://confluence.ecmwf.int/display/CKB/Access+to+CAMS+global+forecast+data).
# #### ECMWF data archive
# ECMWF's data archive is called Meteorological and Archival Retrieval System (MARS) and provides access to ECMWF Public Datasets. The following CAMS data can be accessed through the ECMWF MARS archive: `CAMS reanalysis`, `CAMS GFAS data` (older than one day), and `CAMS global analyses and forecasts` (older than five days).
#
# The archive can be accessed in two ways:
# * via the [web interface](https://apps.ecmwf.int/datasets/) and
# * via the [ECMWF Web API](https://confluence.ecmwf.int/display/WEBAPI/Access+ECMWF+Public+Datasets).
#
# Subsequently, an example is shown how a MARS request can be executed within Python and data in either GRIB or netCDF can be downloaded on-demand.
# #### 1. Register for an ECMWF user account
# - Self-register at https://apps.ecmwf.int/registration/
# - Login at https://apps.ecmwf.int/auth/login
# #### 2. Install the `ecmwfapi` python library
# `pip install ecmwf-api-client`
# #### 3. Retrieve your API key
# You can retrieve your API key at https://api.ecmwf.int/v1/key/. Add the `url`, `key` and `email` information, when you define the `ECMWFDataServer` (see below).
#
# #### 3. Execute a MARS request and download data as `netCDF` file
# Below, you see the principle of a `data retrieval` request. You can use the web interface to browse through the datasets. At the end, there is the option to let generate the `data retrieval` request for the API.
#
# Additionally, you can have a look [here](./cams_ecmwfapi_example_requests.ipynb) at some example requests for different CAMS parameters.
#
# **NOTE**: per default, ECMWF data is stored on a grid with longitudes going from 0 to 360 degrees. It can be reprojected to a regular geographic latitude-longitude grid, by setting the keyword argument `area` and `grid`. Per default, data is retrieved in `GRIB`. If you wish to retrieve the data in `netCDF`, you have to specify it by using the keyword argument `format`.
#
# The example requests `Organic Matter Aerosol Optical Depth at 550 nm` forecast data for 3 June 2019 in `NetCDF`.
# +
# #!/usr/bin/env python
from ecmwfapi import ECMWFDataServer
server = ECMWFDataServer(url="https://api.ecmwf.int/v1", key="<KEY>", email="<KEY>")
# Retrieve data in NetCDF format
server.retrieve({
"class": "mc",
"dataset": "cams_nrealtime",
"date": "2019-06-03/to/2019-06-03",
"expver": "0001",
"levtype": "sfc",
"param": "210.210",
"step": "3",
"stream": "oper",
"time": "00:00:00",
"type": "fc",
"format": "netcdf",
"area": "90/-180/-90/180",
"grid": "0.4/0.4",
"target": "test.nc"
})
# -
# #### CAMS data catalogue of data visualizations
# CAMS provides an extensive [catalogue of data visualizations](https://atmosphere.copernicus.eu/data) in the form of maps and charts. Products are updated daily and are available for selected parameters of `CAMS daily analyses and forecasts`.
# <hr>
# + [markdown] Collapsed="false"
# ## Further information
# + [markdown] Collapsed="false"
# * [EUMETSAT AC SAF - The EUMETSAT Application Facility on Atmospheric Composition Monitoring](https://acsaf.org/index.html)
# * [AC SAF Data policy](https://acsaf.org/data_policy.html)
# * [AC SAF Algorithm Theoretical Basis Documents (atbds)](https://acsaf.org/atbds.html)
#
#
# * [DLR's ATMOS webserver](https://atmos.eoc.dlr.de/app/missions/gome2)
# * [TEMIS - Tropospheric Emission Monitoring Internet Service](http://temis.nl/index.php)
#
#
# * [Copernicus Open Access Hub](https://scihub.copernicus.eu/)
# * [EUMETSAT Earth Observation Portal](https://eoportal.eumetsat.int/userMgmt/login.faces)
# * [Sentinel-5P Mission information](https://sentinels.copernicus.eu/web/sentinel/missions/sentinel-5p)
#
#
# * [Sentinel-3 Mission information](https://sentinel.esa.int/web/sentinel/missions/sentinel-3)
# * [Sentinel-3 OLCI User Guide](https://sentinel.esa.int/web/sentinel/user-guides/sentinel-3-olci)
# * [WEkEO](https://www.wekeo.eu/)
#
#
# * [Copernicus Atmosphere Monitoring Service](https://atmosphere.copernicus.eu/)
# * [ECMWF Web Interface](https://apps.ecmwf.int/datasets/)
# * [ECMWF Web API](https://confluence.ecmwf.int/display/WEBAPI/Access+ECMWF+Public+Datasets)
# * [CAMS catalogue of data visualizations](https://atmosphere.copernicus.eu/data)
# * [CAMS Service Product Portfolio](https://atmosphere.copernicus.eu/sites/default/files/2018-12/CAMS%20Service%20Product%20Portfolio%20-%20July%202018.pdf)
# -
# <br>
# <a href="./index_ltpy.ipynb"><< Index</a><span style="float:right;"><a href="./12_ltpy_WEkEO_harmonized_data_access_api.ipynb">12 - WEkEO Harmonized Data Access API >></a></span>
# + [markdown] Collapsed="false"
# <hr>
# + [markdown] Collapsed="false"
# <p style="text-align:left;">This project is licensed under the <a href="./LICENSE">MIT License</a> <span style="float:right;"><a href="https://gitlab.eumetsat.int/eumetlab/atmosphere/atmosphere">View on GitLab</a> | <a href="https://training.eumetsat.int/">EUMETSAT Training</a> | <a href=mailto:<EMAIL>>Contact</a></span></p>
| 11_ltpy_atmospheric_composition_overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Weekly Assignment 6
#
# ### <NAME> and <NAME>
#
# ### Q. 1
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import LogisticRegression
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error
from sklearn.cluster import KMeans
from sklearn.decomposition import PCA
from sklearn.svm import SVC
# Import the dataset “iowa_housing.csv”. The dataset is about housing prices in Ames, Iowa
# You are going to predict sale price.
df = pd.read_csv("../dataFiles/iowa_housing.csv")
df.head()
# (a) Show the head and shape of the dataframe. Drop “Id” in the dataset.
df_drop = df.drop(["Id"], axis=1)
df_drop.head()
df_drop.shape
# (b) Use the dataset to:
# (i) create a correlation matrix (df.corr()), and then draw the heatmap on the correlation matrix.
corr = df_drop.corr()
corr.head()
sns.heatmap(corr)
# (ii) Find the features that the absolute value of correlation with “SalePrice” > 0.5.
# Then, draw the heatmap of correlation on those features.
features_abs = corr[abs((corr.SalePrice)>=.5)].SalePrice.keys()
abs_corr = corr.loc[features_abs,features_abs]
sns.heatmap(abs_corr)
# (iii) create pair plots on the following features
sns.pairplot(df_drop, vars=['SalePrice', 'OverallQual', 'GrLivArea', 'GarageCars', 'TotalBsmtSF', 'FullBath', 'YearBuilt'])
# (iv) Find the most important feature relative to the “SalePrice” based on absolute value of correlation coefficient.
corr.sort_values(["SalePrice"], ascending = False, inplace = True)
corr.SalePrice.head()
# (v) create a distribution plot on “SalePrice”
sns.distplot(df_drop['SalePrice'], kde =True)
# (vi) For all the numerical features except “SalePrice”,
# replace all the missing values of numerical features with the median value of each feature.
temp=pd.DataFrame()
temp['SalePrice'] = df['SalePrice']
num_cols = df.select_dtypes(exclude = ['object']).columns
num_cols = num_cols.drop('SalePrice')
med_replace = df[num_cols]
med_replace = med_replace.fillna(med_replace.median())
med_replace
# (vii) Create dummies for all categorical features.
# The final shape of dataset should be (1460, 246) (set drop_first = True)
dummies_cols = df.select_dtypes(include = ['object']).columns
new_housing = df[dummies_cols]
new_housing = pd.get_dummies(new_housing, drop_first = True)
new_housing.head()
newdf_housing = pd.concat([temp, med_replace, new_housing], axis =1)
newdf_housing.shape
newdf_housing['SalePrice']
# +
# (c) Then, do the modelling:
# -
# (i) Check for any missing values again.
newdf_housing.isnull().any()
# (ii) create y = “SalePrice”. Drop y from the dataframe.
y = "SalePrice"
house_temp = newdf_housing.drop([y], axis =1)
house_temp.head()
# +
# (iii) Split the dataset into train and test.
X_train, X_test, y_train, y_test = train_test_split(house_temp,newdf_housing, test_size = 0.3, random_state = 42)
print("X_train : " + str(X_train.shape))
print("X_test : " + str(X_test.shape))
print("y_train : " + str(y_train.shape))
print("y_test : " + str(y_test.shape))
# -
# (iv) Train a linear regression model.
model = LinearRegression()
model.fit(X_train, y_train)
# (v) predict “SalePrice” with the test data.
y_pred = model.predict(X_test)
plt.scatter(y_test,y_pred)
plt.xlabel('Y Test (True Values)')
plt.ylabel('Predicted Values')
plt.title('Predicted vs. Actual Values ')
# (vi) find the root mean squared error of the model on the test data.
from sklearn.metrics import mean_squared_error
mse = mean_squared_error(y_test, y_pred)
rmse = np.sqrt(mse)
print("Root Mean Squared Error = ", rmse)
print("The range of temperature = ",y_test.min(), y_test.max())
# ### Q2
# (a) Read in the dataset 'online_shoppers_intention.csv'
df_shoppers = pd.read_csv('../dataFiles/online_shoppers_intention.csv')
df_shoppers.head()
# (b) Count how many shoppers buy, i.e. “Revenue”==True.
# Count how many shoppers browse in the weekends, i.e. “Weekends”==True. Create the following plot.
fig, ax =plt.subplots(1,2)
df_shoppers['Revenue'].value_counts().plot.bar(ax=ax[0])
df_shoppers['Weekend'].value_counts().plot.bar(ax=ax[1])
# (c) Create the following plot, which shows the proportions of various kinds of visitors.
# (first, use value counts, and then use .plot.pie(autopct = '%.2f%%')
df_shoppers['VisitorType'].value_counts().plot.pie(autopct = '%.2f%%')
plt.legend()
# (d) Check the month with most shoppers visiting the online shopping sites. Create the following bar plot.
months = ['Jan', 'Feb', 'Mar', 'Apr','May','Jun', 'Jul', 'Aug','Sep', 'Oct', 'Nov', 'Dec']
new_df = df_shoppers['Month'].value_counts()
new_df = new_df.reindex(months)
new_df.plot.bar()
# (e) For all the numerical variables, fill the missing values with the median of that numerical variable.
num_features = df_shoppers.select_dtypes(exclude = ["object"]).columns
num_shoppers = df_shoppers[num_features]
num_shoppers = num_shoppers.fillna(num_shoppers.median())
num_shoppers
# +
# (f) Create dummies for categorical variables.
cate_vari = df_shoppers.select_dtypes(include = ["object"]).columns
shop_cat = df_shoppers[cate_vari]
shop_cat = pd.get_dummies(shop_cat,drop_first = True)
shop_cat.head()
# +
# (g) Use KMeans to group customers into 3 clusters. That is unsupervised learning. Create the following plot.
kmeans = KMeans(n_clusters=3)
kmeans.fit(num_shoppers)
y_kmeans = kmeans.predict(num_shoppers)
plt.figure(figsize=(10,5))
plt.title('Product related Duration vs Bounce rates', fontsize = 20)
plt.grid()
plt.xlabel('Product related Duration')
plt.ylabel('Bounce rates')
plt.scatter(df_shoppers["ProductRelated_Duration"], df_shoppers["BounceRates"], c=y_kmeans, s=50, cmap='viridis')
# -
# (h) Set y = “Revenue” and X is the dataframe without “Revenue”.
numerical_cols = df_shoppers.select_dtypes(exclude = ["number"]).columns
shop_nums = df_shoppers[numerical_cols]
df_dummies = pd.get_dummies(shop_nums, drop_first=True)
y = df_dummies["Revenue"]
X = df_dummies.drop("Revenue", axis=1)
df_dummies.isnull().any()
# (i) Split the dataset into train and test sets.
# Train a support vector machine classifier model. Use kernel = “rbf” and class_weight='balanced'.
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
model = SVC(kernel='rbf', class_weight='balanced')
model.fit(X_train, y_train)
# (j) Predict which online shopper will do a purchase. Find the accuracy score.
y_pred = model.predict(X_test)
from sklearn.metrics import accuracy_score
accuracy_score(y_test,y_pred)
# ### Q3
# (a) Load the dataset.
from sklearn.datasets import fetch_olivetti_faces
face= fetch_olivetti_faces()
faces = fetch_olivetti_faces().images
# (b) Show the first 10 images
fig, ax = plt.subplots(1, 10, figsize=(64, 64))
for i, axi in enumerate(ax.flat):
axi.imshow(faces[i], cmap=plt.cm.bone)
axi.set(xticks=[], yticks=[])
# +
# (c) Size of each image is 64x64. Use PCA to reduce it into 90 features.
# For the first row, show the first 10 original images.
# Then for the second row, show the first 10 images with reduced number of features.
x = face.data
y = face.target
pca = PCA(n_components = 90)
pca.fit(x)
transformed_data = pca.fit_transform(x)
x_approx = pca.inverse_transform(transformed_data)
x_approx_img = x_approx.reshape(400,64,64)
fig, ax = plt.subplots(1, 10, figsize=(64, 64))
for i, axi in enumerate(ax.flat):
axi.imshow(faces[i], cmap=plt.cm.bone)
axi.set(xticks=[], yticks=[])
fig, ax = plt.subplots(1, 10, figsize=(64, 64))
for i, axi in enumerate(ax.flat):
axi.imshow(x_approx_img[i] , cmap = plt.cm.bone)
axi.set(xticks=[], yticks=[])
# +
# (d) Using images of reduced features to conduct the machine learning task.
# (i) Split the dataset into train and test.
X_train, X_test, y_train, y_test = train_test_split(face.data,face.target, random_state=0)
# -
pca = PCA(n_components=90)
pca.fit(X_train)
# +
# (ii) Train a Random Forest Classifier model. Set n_estimators=100. Predict the target of the images in test set.
model = RandomForestClassifier(n_estimators=100)
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
# -
# (iii) Find the accuracy score.
accuracy_score(y_test,y_pred)
# (iv) Create a confusion matrix, and put it in a heatmap. (Your heatmap may look different due to the randomization in random forest classifier)
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
sns.heatmap(cm)
| assignments/.ipynb_checkpoints/assignment-6-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Calculate historical risk values** using *risk_calculation.py*
# + jupyter={"source_hidden": true}
# Calculate risk function - same as risk_calculation.py, just this time average_brightness is stored as a variable
import numpy as np
import pandas as pd
import ast
import time
import datetime
import statistics
from tqdm import tqdm
# function
def calculateDays(date1, date2):
day1 = time.strptime(date1, "%Y-%m-%d")
day1 = datetime.datetime(day1[0], day1[1], day1[2])
day2 = time.strptime(date2, "%Y-%m-%d")
day2 = datetime.datetime(day2[0], day2[1], day2[2])
#today = datetime.datetime.today()
interval = day1 - day2
return interval.days
# read the cleaned and procesed data
c_df = pd.read_csv('data/combined_dataframes.csv')
# Convert the contents of the pandas array from strings looking like lists to actual lists
brightness_MODIS = c_df.loc[:,'bright_ti4'].apply(ast.literal_eval)
brightness_VIIRS = c_df.loc[:,'bright_ti5'].apply(ast.literal_eval)
instrument = c_df.loc[:,'instrument'].apply(ast.literal_eval)
# Initialise the risk vector
risk = np.zeros(len(c_df.latitude))
for i,list in enumerate(tqdm(iterable = brightness_MODIS, desc = "Insert brightness_MODIS")):
risk[i] += statistics.mean(list)
for i,list in enumerate(tqdm(iterable = brightness_VIIRS, desc = "Insert brightness_VIIRS")):
risk[i] += statistics.mean(list)
# Calculate the average of each of the brightnesses
for i,list in enumerate(tqdm(iterable = risk, desc = "Calculate the average")):
risk[i] = risk[i] / len(instrument[i]) # divide by the number of instruments i.e. mean of 1 or mean of 2
average_brightness = risk.copy()
timeRange = np.zeros(len(c_df.latitude))
timeData = c_df["acq_date"].apply(ast.literal_eval)
for i, value in enumerate(tqdm(iterable = timeData, desc = "Calculate Time Range")):
# if only one day, the result will be the difference between that and the date today
if len(value) == 1:
timeRange[i] = abs(calculateDays("2020-02-15",timeData[i][0]))
# if more than one day, the result will be the difference between the start day and the end day
elif len(value) > 1:
# start day
date1 = timeData[i][0]
# end day
date2 = timeData[i][-1]
timeRange[i] = abs(calculateDays(date2,date1))
# divided by the time range
for i,list in enumerate(tqdm(iterable = risk, desc = "Generate the final Risk")):
risk[i] = risk[i] / timeRange[i]
# -
# **Import required packages**
# + jupyter={"source_hidden": true}
import numpy as np
import pandas as pd
import time
import pandas as pd
import ast
import matplotlib.pyplot as plt
from sklearn.datasets import fetch_california_housing
from sklearn.model_selection import train_test_split
from visualisation import generate_map
# -
# **Define the imput dataset to the Machine Learning Code from the historical risk values and cleaned data**
dataset = pd.concat([c_df.latitude, c_df.longitude,
pd.DataFrame(timeRange, columns=['timeRange']),
pd.DataFrame(average_brightness, columns=['avg_brightness']),
pd.DataFrame(risk, columns=['risk'])],
axis = 1)
dataset.tail()
dataset.info()
# +
import matplotlib.pyplot as plt
plt.style.use('seaborn')
dataset.hist(bins=30, figsize=(20,15))
# -
dataset.corr()
features = dataset[['latitude', 'longitude', 'timeRange', 'avg_brightness']]
labels = dataset['risk']
mean_values = features.describe().iloc[1, :]
# Set random seed to ensure reproducible runs
RSEED = 50
# **Machine Learning Approach 1**
# Split the dataset into train and test sets: 30% of the data is used for training
X_train, X_test, y_train, y_test = train_test_split(features, labels, test_size = 0.30, random_state = RSEED)
X_test.plot(kind='scatter', x='latitude', y='longitude', alpha=0.1, s=20)
X_train.plot(kind='scatter', x='latitude', y='longitude', alpha=0.1, s=20)
X_train.plot(kind='scatter', x='latitude', y='longitude', alpha=0.2, s=20*X_train['avg_brightness'])
corr_matrix = dataset.corr()
corr_matrix['risk'].sort_values(ascending=False)
# +
from pandas.plotting import scatter_matrix
attributes = ['avg_brightness', 'timeRange']
scatter_matrix(X_train[attributes], figsize=(15,10))
# +
# train our dataset on the train data and analyse our Mean Absolute Error on the test data.
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_squared_error
clf = RandomForestRegressor(verbose=True) # enable a progress bar
clf = RandomForestRegressor(n_estimators = 100, max_depth = 50)
clf.fit(X_train, y_train)
print("Mean Absolute Error: {}".format(mean_squared_error(y_test, clf.predict(X_test))))
# -
# Train the model on the complete dataset and then simply use the post endpoint to get the risk
endpoint_classifier = RandomForestRegressor(n_estimators = 100, max_depth = 50)
endpoint_classifier.fit(fetched_data.data, fetched_data.target)
# **Machine Learning Approach 2: random forrest**
# This takes the idea of a single decision tree, and creates an ensemble model out of hundreds or thousands of trees to reduce the variance. Each tree is trained on a random set of the observations, and for each split of a node, only a subset of the features are used for making a split. When making predictions, the random forest averages the predictions for each of the individual decision trees for each data point in order to arrive at a final classification.
labels = np.array(labels)
labels
# Tutorial: https://github.com/WillKoehrsen/Machine-Learning-Projects/blob/master/Random%20Forest%20Tutorial.ipynb
train, test, train_labels, test_labels = X_train, X_test, y_train, y_test
# +
from sklearn import preprocessing
from sklearn import utils
lab_enc = preprocessing.LabelEncoder()
encoded = lab_enc.fit_transform(labels)
# Create the model with 100 trees
model = RandomForestClassifier(n_estimators=100,
random_state=RSEED,
max_features = 'sqrt',
n_jobs=-1, verbose = 1)
# Fit on training data
model.fit(features, encoded)
# +
# We can see how many nodes there are for each tree on average and the maximum depth of each tree. There were 100 trees in the forest.
n_nodes = []
max_depths = []
for ind_tree in model.estimators_:
n_nodes.append(ind_tree.tree_.node_count)
max_depths.append(ind_tree.tree_.max_depth)
print(f'Average number of nodes {int(np.mean(n_nodes))}')
print(f'Average maximum depth {int(np.mean(max_depths))}')
# -
# +
#import all modules
import urllib.request
from bs4 import BeautifulSoup
import plotly.graph_objects as go
import pandas as pd
import numpy as np
import plotly.express as px
# Part 1, get a list of city for visualization ploting
# get lat and long for list of australia cities
city_df = pd.read_html('https://www.latlong.net/category/cities-14-15.html')
city_df = city_df[0]
# split the city, state and country
s = city_df['Place Name'].str.split(", ", n = 2, expand = True)
city_df["City"]= s[0]
city_df["State"]= s[1]
city_df["Country"]= s[2]
pd.options.mode.chained_assignment = None
# cleaning
city_df['City'][8] = 'Chessnok'
city_df['State'][8] = 'NSW'
city_df['Country'][8] = 'Australia'
city_df['City'][71] = 'Greenvale'
city_df['State'][71] = 'Victoria'
city_df['Country'][71] = 'Australia'
city_df['City'][83] = 'Gladstone'
city_df['State'][83] = 'QLD'
city_df['Country'][83] = 'Australia'
city_df['City'][80] = 'Gladstone'
city_df['State'][80] = 'QLD'
city_df['Country'][80] = 'Australia'
city_df['State'] = city_df['State'].str.replace('Queensland', 'QLD')
city_df['State'] = city_df['State'].str.replace('Tasmania', 'TAS')
city_df['State'] = city_df['State'].str.replace('Victoria', 'VIC')
city_df['State'] = city_df['State'].str.replace('Canberra', 'ACT')
city_df['State'] = city_df['State'].str.replace('Northern Territory', 'NT')
# Part 2, summarize historical numbers by state
#Open the url to be scraped
url = "https://en.wikipedia.org/wiki/List_of_major_bushfires_in_Australia"
page = urllib.request.urlopen(url)
#Convert page to a beautifulsoup object
soup = BeautifulSoup(page, "lxml")
#Need to find the table
fire_table = soup.find('table', class_='wikitable sortable')
#Set up individual lists for each of the columns
Date = []
States = []
HA = []
Acres = []
Fatalities = []
Homes = []
#go through each row and append each cell to respective list
for row in fire_table.find_all('tr'):
cells = row.find_all('td')
if len(cells) == 10:
Date.append(cells[0].find(text=True).strip("\n"))
States.append(cells[2].find(text=True).strip("\n"))
HA.append(cells[3].find(text=True).strip("\n"))
Acres.append(cells[4].find(text=True).strip("\n"))
Fatalities.append(cells[5].find(text=True).strip("approx. \n"))
Homes.append(cells[6].find(text=True).strip("approx. \n"))
#Convert all relevant scraped cells into a DataFrame
fire_df = pd.DataFrame(Date, columns=["Date"])
fire_df["States"] = States
fire_df["HA"] = HA
fire_df["Fatalities"] = Fatalities
fire_df["Homes"] = Homes
#Need to do some extra cleaning on the dataframe
fire_df = fire_df.replace(to_replace = "Nil", value = "0")
# cleaning
fire_df['HA'] = fire_df['HA'].str.replace(',', '')
fire_df['Fatalities'] = fire_df['Fatalities'].str.replace(',', '')
fire_df['Homes'] = fire_df['Homes'].str.replace(',', '')
fire_df['Homes'] = fire_df['Homes'].str.replace(',', '')
fire_df['HA'][7] = 160000
fire_df['Fatalities'][4] = 20
fire_df['Homes'][19] = 0
fire_df['Year'] = fire_df['Date'].str[-4:]
fire_df['Year'][197] = 2020
# transform data type to numeric
fire_df['HA'] = pd.to_numeric(fire_df['HA'], errors='coerce')
fire_df['Fatalities'] = pd.to_numeric(fire_df['Fatalities'], errors='coerce')
fire_df['Homes'] = pd.to_numeric(fire_df['Homes'], errors='coerce')
fire_df['Year'] = pd.to_numeric(fire_df['Year'], errors='coerce')
# pivot table to get summary by state
df1=pd.pivot_table(fire_df, index=['States'],values=['HA','Fatalities','Homes'],aggfunc=np.sum)
df2=fire_df.groupby('States').Date.nunique()
wiki_df = pd.concat([df1,df2],axis=1)
wiki_df= wiki_df.rename(columns={'Date': 'FireCount'})
wiki_df['State_ab']=('NA', 'ACT', 'NW', 'NSW', 'NT', 'SA', 'TAS', 'VIC', 'WA')
# left join two dataframes
combine_df = pd.merge(left=city_df,right=wiki_df, how='left', left_on='State', right_on='State_ab')
# plot on map
fig = px.scatter_mapbox(combine_df, lat="Latitude", lon="Longitude", hover_name="City", hover_data=["Fatalities", "Homes"],
color_discrete_sequence=["fuchsia"], zoom=3, height=300)
fig.update_layout(mapbox_style="open-street-map")
fig.update_layout(margin={"r":0,"t":0,"l":0,"b":0})
fig.show()
# -
fire_df
fire_df
| risk_forecast_ML.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/nnuncert/nnuncert/blob/master/notebooks/PNN-E_toy.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="5-sSRaon3d1d"
# # Git + Repo Installs
# + id="viDL06MYoQhB"
# !git clone https://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@github.com/nnuncert/nnuncert
# + id="2wqtWNj43fI9"
# %cd nnuncert
# !pip install -r requirements.txt
# + [markdown] id="76P-Kug5kfGl"
# # Imports
# + id="9LhGoMW9k3RN"
# # %cd nnuncert
# + id="8o_PzKDikhz6"
# general imports
import numpy as np
import numexpr as ne
import tensorflow as tf
import matplotlib.pyplot as plt
# thesis code
import nnuncert
from nnuncert.models import make_model, type2name
from nnuncert.app.toy import make_toy_data, make_toy_plot, gen_2d_gaussian_samples, input2grid, contour_plot_2d
from nnuncert.utils.traintest import TrainTestSplit
# + [markdown] id="nU1noww7k7OU"
# # Toy 1D
# + [markdown] id="SOV-7prllB8R"
# ## Make data
# Generate input features on (-4, 4) uniformly and calculate noisy targets with true function 'x**3' and additive homoscedastic noise N(0, 3).
# + id="rAppye78lFK4"
# set seed for reproducibility and make random number generator
seed = 21
rng = np.random.default_rng(seed)
# define function that generates true relationship, possible to use any kind of expression such as "sin(x)", "exp(x)", "3*x**2 - 8x + 14", ...
def reg_func(x):
reg_func = "x**3"
return ne.evaluate(reg_func)
# generate input data (x) uniformly from -4 to 4
x = rng.random((20, ))*8 - 4
# alternatively: setup (multiple) clusters and sample uniformly in between
# ppc = 20 # points per cluster
# clusters = [[-4, 4]] # list of cluster bounds (left, right)
# x = np.array([rng.choice(np.linspace(x1, x2, 1000), ppc)
# for (x1, x2) in clusters]).ravel()
# generate responses with function
noise_std = 3
data_1d = make_toy_data(x, reg_func, noise_std, seed=rng)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="tyEoygManN88" outputId="3336472e-42f6-4601-f119-e95b113e3d5c"
# have a look at the data
data_1d.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="X3GMKlA1nUOD" outputId="69e8584a-3190-43b6-9f13-4a840d840f30"
# plot ground truth
fig, ax = plt.subplots(figsize=(8, 4))
minx, maxx = [-6, 6]
x0 = np.linspace(minx, maxx, 80)
ax.scatter(data_1d["x1"], data_1d["y"])
ax.plot(x0, reg_func(x0), "--", color="black")
# + [markdown] id="x7uGJcxJlCxN"
# ## Fit model
# Fit a model to all training samples that were generated.
# Get predictitve mean, variance for 100 inputs in (-6, 6), evenly spaced.
# + id="JmtmfghLnq4F"
class TrainTestSplitToy1D(TrainTestSplit):
def __init__(self, df, train_id = None, test_id = None, test_ratio=0.1, norm_x=False, rng=None):
non_norm = []
if norm_x is False:
non_norm = ["x1"]
super(TrainTestSplitToy1D, self).__init__(df, "y", non_norm=non_norm,train_id=train_id, test_id=test_id, ratio=test_ratio, rng=rng)
# standardize features x
toy_1d = TrainTestSplitToy1D(data_1d, test_ratio=0, norm_x=True, rng=rng)
input_shape = toy_1d.x_train.shape[1]
# + id="DMx4EMS3oDvq"
# handle general settings
arch = [[50, "relu", 0]] # list of hidden layer description (size, act. func, dropout rate)
epochs = 40
verbose = 0
learning_rate = 0.01
# make model (5 ensemble members)
model = make_model("PNN-E", input_shape, arch, N=5)
model.compile(optimizer=tf.keras.optimizers.Adam(learning_rate=learning_rate), metrics=["mae", "mse"])
# fit model to all samples (test_ratio in TrainTestSplitToy1D was set to 0)
model.fit(toy_1d.x_train, toy_1d.y_train, epochs=epochs, verbose=verbose)
# get prediction on scaled test input features
xlim = (-6, 6)
x0 = np.linspace(*xlim, 100)
pred = model.make_prediction(toy_1d.scale_x(x0))
# + [markdown] id="qdgNVRH4qA5o"
# ## Plot
# Plot mean prediction and predictive uncertainty (2 standard deviations).
# + colab={"base_uri": "https://localhost:8080/", "height": 324} id="yPvdbu4GpJe5" outputId="dc0f035a-e526-4ce5-a438-2e25bf3515ca"
# setup some colors and setup limits for y axis
colors = ["mediumblue", "tab:blue", "#b3cde0"]
ylim = (-150, 150)
fig, ax = plt.subplots(figsize=(8, 5))
plt.setp(ax, xlim=xlim, ylim=ylim)
make_toy_plot(pred, toy_1d.x_train_us, toy_1d.y_train_us, x0=x0, reg_func=reg_func, std_devs=2, colors=colors, ax=ax)
# + [markdown] id="CJsa74J9qt-h"
# # Toy 2D
# + [markdown] id="ndxOEKb5qxqy"
# ## Make data
# Create input data clusters located at (-1, -1) and (1, 1) and generate noise response with standard deviation = 0.01 (very low noise).
# + id="mHYM10Bpq82d"
# set seed for reproducibility and make random number generator
seed = 21
rng = np.random.default_rng(seed)
# define function that generates true relationship
def reg_func_2d(x):
x1, x2 = x.T
reg_func = "0.8*x1 + 0.8*x2"
return ne.evaluate(reg_func)
# generate input data (x), defined by cluster centers muh and noise in the data
muh = [[-1, -1], [1, 1]]
x = gen_2d_gaussian_samples(muh, var=0.02, ppc=50, seed=rng)
# make y data
toy2d_noise_std = 0.1
data2d = make_toy_data(x, reg_func_2d, toy2d_noise_std, seed=rng)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="u1wEb6BRrqNs" outputId="545a3d94-0843-43ee-c793-f04456cb162f"
# show data
data2d.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="3ybeOq45rjcw" outputId="fd49ea72-c10f-401f-f7ff-0f4d6e674f48"
# plot noisy data in 3D
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.scatter(data2d.x1, data2d.x2, data2d.y, c="red")
# + [markdown] id="xtD2oVC_qys1"
# ## Fit model
# Fit a model to all training samples that were generated.
# Get predictitve mean, variance for inputs on a [-2, 2] x [-2, 2] grid.
# + id="cvpJQZg_rzt5"
class TrainTestSplitToy2D(TrainTestSplit):
def __init__(self, df, train_id = None, test_id = None, test_ratio=0.1, norm_x=False, rng=None):
non_norm = ["x1", "x2"]
if norm_x is True:
non_norm = []
super(TrainTestSplitToy2D, self).__init__(df, "y", non_norm=non_norm, train_id=train_id, test_id=test_id, ratio=test_ratio, rng=rng)
# standardize features x
toy_2d = TrainTestSplitToy2D(data2d, test_ratio=0, norm_x=True, rng=rng)
input_shape = toy_2d.x_train.shape[1]
# + id="dyT24IymsBps"
# handle general settings
arch = [[50, "relu", 0]] # list of hidden layer description (size, act. func, dropout rate)
epochs = 40
verbose = 0
learning_rate = 0.01
# make model (5 ensemble members)
model = make_model("PNN-E", input_shape, arch, N=5)
model.compile(optimizer=tf.keras.optimizers.Adam(learning_rate=learning_rate), metrics=["mae", "mse"])
# fit model to all samples (test_ratio in TrainTestSplitToy1D was set to 0)
model.fit(toy_2d.x_train, toy_2d.y_train, epochs=epochs, verbose=verbose)
# generate scaled test features
ppa = 100 # points per axis -> 100*100=10,000 test points
grid_in, x1, x2 = input2grid([-2, 2], [-2, 2], ppa)
x_test = toy_2d.scale_x(grid_in)
# get prediction of model for test features
pred = model.make_prediction(x_test)
# + [markdown] id="PyBO6DNZq0Dm"
# ## Plot
# Contour plot of predictive uncertainty (std. deviation) for input features.
# + colab={"base_uri": "https://localhost:8080/", "height": 488} id="6XEMIKTms9Lc" outputId="f19624e2-bd1a-4e47-f4d5-30f42811cd64"
# init plot and define colormap
fig, ax = plt.subplots(figsize=(8, 8))
cmap = plt.get_cmap("viridis")
# retrieve predictive standard deviation in proper form
std = pred.std_total.reshape(ppa, ppa)
# plot std as contour to ax
contour_plot_2d(x1, x2, std, x_train=(data2d[["x1", "x2"]].values, "white"), levels=20, cmap=cmap, ax=ax, fig_=fig, make_colbar=True)
| notebooks/PNN-E_toy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pangeo
# language: python
# name: pangeo
# ---
# <center><h1><strong>CLEANING MODEL FORECAST</strong></h1></center>
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
plt.style.use('ggplot')
import os
# ## **CLEANING MODEL FORECAST**
fili = '../DATASETS/forecast_air_quality_2020_report.xls'
df_model = pd.read_excel(fili)
df_model = df_model.rename(columns={'ZONA':'ZONE','FECHA_PRONOSTICO':'date','ESTADO_CALIDAD_AIRE':'MODEL'})
df_model = df_model[['ZONE','date','MODEL']]
df_model['ZONE'] = df_model['ZONE'].str.replace('Callao','Lima Oeste').str.replace('Lima','').str.upper()
df_model['date'] = pd.to_datetime(df_model['date'])
df_model.set_index('date',inplace=True)
# ## **CLEANING OBS**
# OBS
df_obs = pd.read_excel('../DATASETS/indice_carlos octubre.xlsx', skiprows=1, usecols = "A:D")
df_obs['date'] = pd.to_datetime(df_obs['date'], format='%m/%d/%Y')
df_obs = df_obs.melt(id_vars='date', var_name='ZONE',value_name='OBS')
df_obs = df_obs.apply(lambda x: x.replace({'SJL':'ESTE','CRB':'NORTE','CDM':'CENTRO'}) if x.name=='ZONE' else x)
df_obs['OBS'] = df_obs['OBS'].apply(lambda x: 'buena' if (x>0 and x<=12) else 'moderado' if (x>12 and x<=35.4) else \
'insalubre_grupo_sensible' if (x>35.4 and x<=55.4) else 'insalubre' if (x>55.4 and x<=150.4) else \
'muy_insalubre' if (x>150.4) else np.nan)
df_obs.set_index('date', inplace=True)
# ## **CONCATENING**
df_merged = pd.concat([df_model,df_obs])
df_merged['eval'] = df_merged.apply(lambda x: x['MODEL']==x['OBS'], axis=1)
df_merged['eval'] = df_merged['eval'].apply(lambda x: 'Acierto' if x==True else 'Desacierto')
df_merged.dropna(how='all', axis=0)
fig, ax = plt.subplots(figsize=(10,6), dpi=80)
sns.countplot(df_merged['eval'],ax=ax, hue=df_merged['ZONE'],palette=sns.color_palette("husl"))
ax.set_ylabel('Conteo de casos')
ax.set_xlabel('')
ax.set_title('PM2.5 evaluación pronostico')
| NOTEBOOKS/Validations_Cross.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Reading data for red and white wine
# \[_In case you’re unable to see the atoti visualizations in GitHub, try viewing the notebook in [nbviewer](https://nbviewer.org/github/atoti/notebooks/blob/master/notebooks/wine-analytics/main.ipynb)._]
#
# In this notebook, we are using the wine quality datasets from https://archive.ics.uci.edu/ml/datasets/Wine+Quality.
# <div style="text-align: center;" ><a href="https://www.atoti.io/?utm_source=gallery&utm_content=wineAnalytics" target="_blank"><img src="https://data.atoti.io/notebooks/banners/discover.png" alt="Try atoti"></a></div>
import atoti as tt
import numpy as np
import pandas as pd
import seaborn as sns
wine_red = pd.read_csv(
"https://data.atoti.io/notebooks/wine-analytics/winequality-red.csv", sep=";"
)
wine_red.head()
wine_white = pd.read_csv(
"https://data.atoti.io/notebooks/wine-analytics/winequality-white.csv", sep=";"
)
wine_white.head()
# ### Merging dataset for loading into atoti
#
# We can combine the 2 dataset by adding the category for the wine.
# +
wine_red["category"] = "Red"
wine_white["category"] = "White"
wines = pd.concat([wine_red, wine_white], axis=0, ignore_index=True)
wines.index.set_names("wine index", inplace=True)
wines.head()
# -
# We group the wines into 3 broad ratings-good, average and poor-based on the quality.
wines["rating"] = "Good"
wines.loc[wines["quality"] < 7, "rating"] = "Average"
wines.loc[wines["quality"] < 5, "rating"] = "Poor"
wines
# ### Wine analysis by correlated features
corr_red = wine_red.corr()
ax_red = sns.heatmap(
corr_red,
vmin=-1,
vmax=1,
center=0,
cmap=sns.diverging_palette(20, 220, n=200),
square=True,
)
ax_red.set_xticklabels(
ax_red.get_xticklabels(), rotation=45, horizontalalignment="right"
);
# Let's look at the features that are correlated to the quality.
# The correlation value is not very high, so just for demonstrative purpose, we will take the threshold value of 0.25 to select a few key features that are most correlated.
# We will not consider the interaction between the features.
corr_red[(corr_red["quality"] > 0.25) | (corr_red["quality"] < -0.25)]["quality"]
corr_white = wine_white.corr()
corr_white[(corr_white["quality"] > 0.25) | (corr_white["quality"] < -0.25)]["quality"]
wine_variables = wines.reset_index()
wine_variables.head()
wine_variables_melt = pd.melt(
wine_variables,
id_vars=["wine index"],
value_vars=[
"fixed acidity",
"volatile acidity",
"citric acid",
"residual sugar",
"chlorides",
"free sulfur dioxide",
"total sulfur dioxide",
"density",
"pH",
"sulphates",
"alcohol",
],
)
wine_variables_melt.head()
# ### Getting started with atoti
session = tt.create_session(config={"user_content_storage": "./content"})
wines_table = session.read_pandas(
wine_variables_melt, table_name="Wine quality", keys=["wine index", "variable"]
)
cube = session.create_cube(wines_table, "Wines")
cube.schema
# ### Joining additional tables to the cube
wines_detail = session.read_pandas(
wine_variables[["wine index", "category", "rating", "quality"]],
table_name="Wine detail",
keys=["wine index"],
)
wines_table.join(wines_detail)
cube.schema
# + atoti={"height": 434, "widget": {"mapping": {"horizontalSubplots": [], "splitBy": ["[Wine detail].[rating].[rating]", "ALL_MEASURES"], "values": ["[Measures].[value.MEAN]"], "verticalSubplots": ["[Wine detail].[category].[category]"], "xAxis": ["[Wine quality].[variable].[variable]"]}, "query": {"mdx": "SELECT NON EMPTY Hierarchize(Descendants({[Wine quality].[variable].[AllMember]}, 1, SELF_AND_BEFORE)) ON ROWS, NON EMPTY Crossjoin(Hierarchize(Descendants({[Wine detail].[rating].[AllMember]}, 1, SELF_AND_BEFORE)), {[Measures].[value.MEAN]}, Hierarchize(Descendants({[Wine detail].[category].[AllMember]}, 1, SELF_AND_BEFORE))) ON COLUMNS FROM [Wines] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "plotly-clustered-column-chart"}} tags=[]
session.visualize("Wine characteristics comparison")
# -
# ### Creating hierarchy from numerical columns
h, m = cube.hierarchies, cube.measures
h["value"] = [wines_table["value"]]
h
# ### Creating simple measure from referenced table
m["quality"] = tt.agg.mean(wines_detail["quality"])
# + atoti={"widget": {"filters": ["[Wine quality].[variable].[AllMember].[volatile acidity]"], "mapping": {"horizontalSubplots": [], "splitBy": ["[Wine detail].[category].[category]", "[Wine quality].[variable].[variable]", "ALL_MEASURES"], "values": ["[Measures].[quality]"], "verticalSubplots": [], "xAxis": ["[Wine quality].[value].[value]"]}, "query": {"mdx": "SELECT NON EMPTY Crossjoin(Hierarchize(Descendants({[Wine detail].[category].[AllMember]}, 1, SELF_AND_BEFORE)), Hierarchize(Descendants({[Wine quality].[variable].[AllMember]}, 1, SELF_AND_BEFORE)), {[Measures].[quality]}) ON COLUMNS, NON EMPTY Hierarchize(Descendants({[Wine quality].[value].[AllMember]}, 1, SELF_AND_BEFORE)) ON ROWS FROM [Wines] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "plotly-line-chart"}} tags=[]
session.visualize("Features vs Wine quality")
# -
session.link(path="/#/dashboard/29f")
# <div style="text-align: center;" ><a href="https://www.atoti.io/?utm_source=gallery&utm_content=wineAnalytics" target="_blank" rel="noopener noreferrer"><img src="https://data.atoti.io/notebooks/banners/discover-try.png" alt="Try atoti"></a></div>
| notebooks/wine-analytics/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import cv2
img = cv2.imread('images/g001.jpg', 1) # second param 1 -> 3 channels or 0 -> just one channel
type(img)
img.shape
img.dtype
img.size
| img_size.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="dquM-bfUIW-J"
import os
import numpy as np
import math
import matplotlib.pyplot as plt
import json
import pandas as pd
from IPython.display import display
from tqdm import tqdm, tqdm_notebook, trange
import sentencepiece as spm
import wget
import torch
import torch.nn as nn
import torch.nn.functional as F
# + colab={"base_uri": "https://localhost:8080/", "height": 136} colab_type="code" id="8pjohaiGId95" outputId="7e5d8ddc-d2f3-4059-b53b-d1d1304aa746"
data_dir = "../data"
for f in os.listdir(data_dir):
print(f)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="u69gqsv_IkrL" outputId="3609e01b-9c36-4009-9e0d-d5d9ac0a7e7e"
# vocab loading
vocab_file = f"{data_dir}/kowiki.model"
vocab = spm.SentencePieceProcessor()
vocab.load(vocab_file)
# + colab={} colab_type="code" id="IHGknKGNI9SU"
""" configuration json을 읽어들이는 class """
class Config(dict):
__getattr__ = dict.__getitem__
__setattr__ = dict.__setitem__
@classmethod
def load(cls, file):
with open(file, 'r') as f:
config = json.loads(f.read())
return Config(config)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="x3UQU0e8JM0b" outputId="202d1af5-c2c5-421f-bb0c-fc6ba16320eb"
config = Config({
"n_enc_vocab": len(vocab),
"n_dec_vocab": len(vocab),
"n_enc_seq": 256,
"n_dec_seq": 256,
"n_layer": 6,
"d_hidn": 256,
"i_pad": 0,
"d_ff": 1024,
"n_head": 4,
"d_head": 64,
"dropout": 0.1,
"layer_norm_epsilon": 1e-12
})
print(config)
# -
def get_sinusoid_encoding_table(n_seq, d_hidn):
def cal_angle(position, i_hidn):
return position / np.power(100000, 2 * (i_hidn // 2) / d_hidn)
def get_posi_angle_vec(position):
return [cal_angle(position, i_hidn) for i_hidn in range(d_hidn)]
sinusoid_table = np.array([get_posi_angle_vec(i+seq) for i_seq in range(n_seq)])
sinusoid_table[:, 0::2] = np.sin(sinusoid_table[:, 0::2])
sinusoid_table[:, 1::2] = np.cos(sinusoid_table[:, 1::2])
return sinusoid_table
# ### size() 함수를 쓰기 위해서 torch로 q와 k를 만들어준다.
# +
lines = [
"겨울은 추워요.",
"감기 조심하세요."
]
inputs = []
for line in lines:
pieces = vocab.encode_as_pieces(line)
ids = vocab.encode_as_ids(line)
inputs.append(torch.tensor(ids))
print(pieces)
# 입력 길이가 다르므로 입력 최대 길이에 맟춰 padding(0)을 추가 해 줌
k = torch.nn.utils.rnn.pad_sequence(inputs, batch_first=True, padding_value=0)
k.size()
# +
lines = [
"겨울은 추워요.",
"감기 조심하세요."
]
inputs = []
for line in lines:
pieces = vocab.encode_as_pieces(line)
ids = vocab.encode_as_ids(line)
inputs.append(torch.tensor(ids))
print(pieces)
# 입력 길이가 다르므로 입력 최대 길이에 맟춰 padding(0)을 추가 해 줌
q = torch.nn.utils.rnn.pad_sequence(inputs, batch_first=True, padding_value=0)
q.size()
# -
def get_attn_pad_mask(seq_q, seq_k, i_pad):
batch_size, len_q = seq_q.size()
batch_size, len_k = seq_k.size()
pad_attn_mask = seq_k.data.eq(i_pad).unsqueeze(1).expand(batch_size, len_q, len_k)
return pad_attn_mask
#torch.Size([2, 8]), torch.Size([2, 8])
get_attn_pad_mask(q, k , 1)
#torch.Size([2, 9]), torch.Size([2, 8])
get_attn_pad_mask(q, k , 2)
# 이 함수를 통해서 위와 같이 크기를 맞줘줘서 나중에 내적을 쉽게 할 수 있다.
# + [markdown] colab_type="text" id="IJbvIXsNjloN"
# ### Decoder Mask를 구하기
# -
# -----------------------
#
# unsqueeze(-1) : 가로 새로 바구기
# ```
# torch.unsqueeze(x, 0)
# tensor([[ 1, 2, 3, 4]])
# torch.unsqueeze(x, 1)
# tensor([[ 1],
# [ 2],
# [ 3],
# [ 4]])
# ```
# -----------
#
# torch.ones_like : 가로안 만큼 1 만들기
#
#
# ------------------
#
# triu(diagonal=1) : 뭉쳐 있는 배열들을 보기 쉽게 아래로 내려주기
# ```
# np.triu([[1,2,3],[4,5,6],[7,8,9],[10,11,12]], -1)
# array([[ 1, 2, 3],
# [ 4, 5, 6],
# [ 0, 8, 9],
# [ 0, 0, 12]])
# ```
# --------------------
#
# torch.triu(a, diagonal=1) # 아래는 다 0으로 조지기
# ```
# tensor([[ 0.0000, 0.5207, 2.0049],
# [ 0.0000, 0.0000, 0.6602],
# [ 0.0000, 0.0000, 0.0000]])
# ```
def get_dttn_decoder_mask(seq):
# 빈 마스크를 만들어주고 (크기만큼만)
subsequent_mask = torch.ones_like(seq).unsqueeze(-1).expand(seq.size(0), seq.size(1), seq.size(1))
# 행렬의 위쪽 삼각형 부분(2-D)
subsequent_mask = subsequent_mask.triu(diagonal=1)
return subsequent_mask
# + [markdown] colab_type="text" id="B7rndBaMj16n"
# ###### ScaledDotProductAttention
# 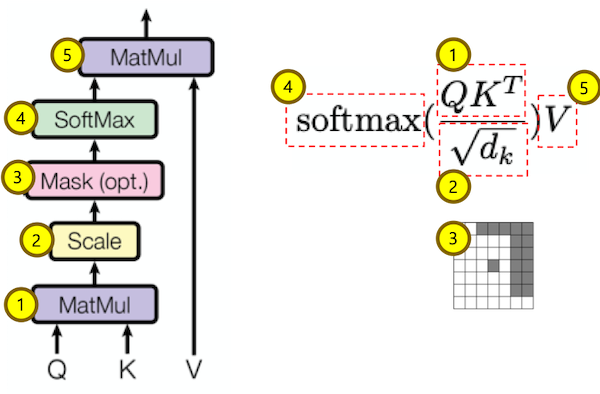
# -
q, q.transpose(-1, -2)
# +
# masked_fill_ 역할은 위치에 있는 것들 제거하는거
# -
# ```
# In [3]: mask = torch.tensor([[[1, 0]], [[0, 1]]], dtype=torch.uint8)
#
# In [4]: mask.shape
# Out[4]: torch.Size([2, 1, 2])
#
# In [5]: x = torch.randn(2, 2)
#
# In [6]: x, mask = torch.broadcast_tensors(x, mask)
#
# In [7]: mask.shape
# Out[7]: torch.Size([2, 2, 2])
#
# In [8]: x.is_contiguous()
# Out[8]: False
#
# In [9]: x.masked_fill(mask, 0) # This should be the correct behavior
# Out[9]:
# tensor([[[ 0.0000, 0.9482],
# [ 0.0000, 0.0719]],
#
# [[ 0.6267, 0.0000],
# [-0.6296, 0.0000]]])
# ```
class ScaledDotProductAttention(nn.Module):
def __init__(self, config):
super().__init__()
self.config = config
self.dropout = nn.Dropout(config.dropout)
self.scale = 1 / (self.config.d_head ** 0.5)
def forward(self, Q, K, V, attn_mask):
scores = torch.matmul(Q, K.transpose(-1, -2)).mul_(self.scale)
scores.masked_fill_(attn_mask, -1e9)
attn_prob = nn.Softmax(dim=-1)(scores)
attn_prob = self.dropout(attn_prob)
context = torch.matmul(attn_prob, V)
return context, attn_prob
# + [markdown] colab_type="text" id="zJMLVMjkj7_1"
# ###### MultiHeadAttention
# 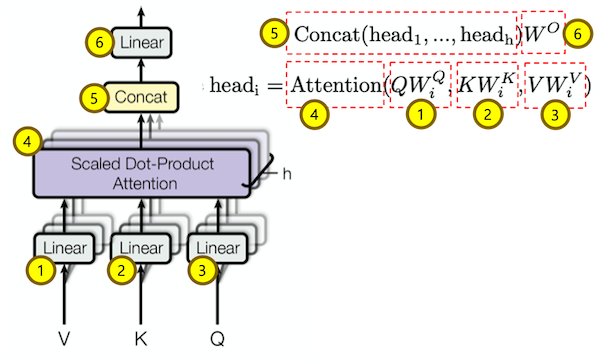
# -
# self.config.n_head * self.config.d_head 만큼 들어오고
#
#
# self.config.d_hidn 만큼 멀티해드 어텐션 저 위에 h가 이것이여
class MultiHeadAttention(nn.Module):
def __init__(self, config):
super().__init__()
self.config = config
self.W_Q = nn.Linear(self.config.d_hidn, self.config.n_head * self.config.d_head)
self.W_K = nn.Linear(self.config.d_hidn, self.config.n_head * self.config.d_head)
self.W_V = nn.Linear(self.config.d_hidn, self.config.n_head * self.config.d_head)
self.scaled_dot_attn = ScaledDotProductAttention(self.config)
self.linear = nn.Linear(self.config.n_head * self.config.d_head, self.config.d_hidn)
self.dropout = nn.Dropout(config.dropout)
def forward(self, Q, K, V, attn_mask):
batch_size = Q.size(0)
# (bs, n_head, n_q_seq, d_head) / view는 크기를 조절하는 거 일단 batch는 고정하고 seq는 자유롭게 r그리고 n_head 고정
# 마무리로 늘려주기 까지
q_s = self.W_Q(Q).view(batch_size, -1, self.config.n_head, self.config.d_head).transpose(1,2)
# (bs, n_head, n_k_seq, d_head)
k_s = self.W_K(K).view(batch_size, -1, self.config.n_head, self.config.d_head).transpose(1,2)
# (bs, n_head, n_v_seq, d_head)
v_s = self.W_V(V).view(batch_size, -1, self.config.n_head, self.config.d_head).transpose(1,2)
# (bs, n_head, n_q_seq, n_k_seq)
# unsqueeze(1) 살려두는 것들은 1로 한 줄로 만드는건가
attn_mask = attn_mask.unsqueeze(1).repeat(1, self.config.n_head, 1, 1)
# (bs, n_head, n_q_seq, d_head), (bs, n_head, n_q_seq, n_k_seq)
context, attn_prob = self.scaled_dot_attn(q_s, k_s, v_s, attn_mask)
# (bs, n_head, n_q_seq, h_head * d_head)
context = context.transpose(1, 2).contiguous().view(batch_size, -1, self.config.n_head * self.config.d_head)
# (bs, n_head, n_q_seq, e_embd)
output = self.linear(context)
output = self.dropout(output)
# (bs, n_q_seq, d_hidn), (bs, n_head, n_q_seq, n_k_seq)
return output, attn_prob
#
# sizes (torch.Size or python:int...) – The number of times to repeat this tensor along each dimension
#
# Example:
# ```
# >>> x = torch.tensor([1, 2, 3])
# >>> x.repeat(4, 2)
# tensor([[ 1, 2, 3, 1, 2, 3],
# [ 1, 2, 3, 1, 2, 3],
# [ 1, 2, 3, 1, 2, 3],
# [ 1, 2, 3, 1, 2, 3]])
# >>> x.repeat(4, 2, 1).size()
# torch.Size([4, 2, 3])
# ```
# + [markdown] colab_type="text" id="df1dKmZSkEiV"
# ###### PoswiseFeedForwardNet
#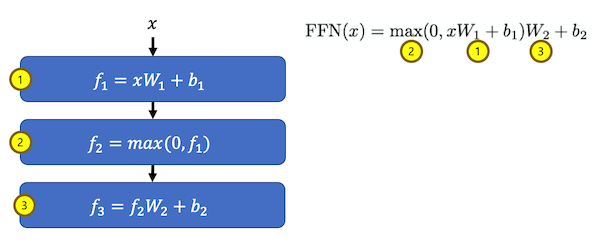 
# -
class PoswiseFeedForwardNet(nn.Module):
def __init__(self, config):
super().__init__()
self.config = config
self.conv1 = nn.Conv1d(in_channels=self.config.d_hidn, out_channels=self.config.d_ff, kernel_size=1)
self.conv2 = nn.Conv1d(in_channels=self.config.d_ff, out_channels=self.config.d_hidn, kernel_size=1)
self.active = F.gelu
self.dropout = nn.Dropout(config.dropout)
def forward(self, inputs):
# (bs, d_ff, n_seq)
output = self.active(self.conv1(inputs.transpose(1, 2)))
# (bs, n_seq, d_hidn)
output = self.conv2(output).transpose(1, 2)
output = self.dropout(output)
# (bs, n_seq, d_hidn)
return output
# + [markdown] colab_type="text" id="KQaP14UDkgpV"
# #### 7. Encoder
# 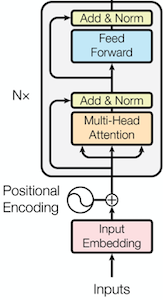
# + [markdown] colab_type="text" id="4HxrA0mdkoFu"
# ###### EncoderLayer
# -
class EncoderLayer(nn.Module):
def __init__(self, config):
super().__init__()
self.config = config
self.self_attn = MultiHeadAttention(self.config)
self.layer_norm1 = nn.LayerNorm(self.config.d_hidn, eps=self.config.layer_norm_epsilon)
self.pos_ffn = PoswiseFeedForwardNet(self.config)
self.layer_norm2 = nn.LayerNorm(self.config.d_hidn, eps=self.config.layer_norm_epsilon)
def forward(self, inputs, attn_mask):
# (bs, n_enc_seq, d_hidn), (bs, n_head, n_enc_seq, n_enc_seq)
att_outputs, attn_prob = self.self_attn(inputs, inputs, inputs, attn_mask)
att_outputs = self.layer_norm1(inputs + att_outputs)
# (bs, n_enc_seq, d_hidn)
ffn_outputs = self.pos_ffn(att_outputs)
ffn_outputs = self.layer_norm2(ffn_outputs + att_outputs)
# (bs, n_enc_seq, d_hidn), (bs, n_head, n_enc_seq, n_enc_seq)
return ffn_outputs, attn_prob
# + [markdown] colab_type="text" id="2OqU2L7IlDO_"
# ###### Encoder
# -
class Encoder(nn.Module):
def __init__(self, config):
super().__init__()
self.config = config
self.enc_emb = nn.Embedding(self.config.n_enc_vocab, self.config.d_hidn)
# 멀티헤드 어텐션으로 위치 찾아주기
sinusoid_table = torch.FloatTensor(get_sinusoid_encoding_table(self.config.n_enc_seq + 1, self.config.d_hidn))
# 포지션 임베딩 때리고
self.pos_emb = nn.Embedding.from_pretrained(sinusoid_table, freeze = True)
# 레이어 수만큼 반복
self.layers = nn.ModuleList([EncoderLayer(self.config) for _ in range(self.config.n_layer)])
def forward(self, inputs):
positions = torch.arange(inputs.size(1), device=inputs.device, dtype=inputs.dtype).expand(inputs.size(0), inputs.size(1)).contiguous() + 1
pos_mask = inputs.eq(self.config.i_pad)
positions.masked_fill_(pos_mask, 0)
# (bs, n_enc_seq, d_hidn)
outputs = self.enc_emb(inputs) + self.pos_emb(positions)
# (bs, n_enc_seq, n_enc_seq)
attn_mask = get_attn_pad_mask(inputs, inputs, self.config.i_pad)
attn_probs = []
for layer in self.layers:
# (bs, n_enc_seq, d_hidn), (bs, n_head, n_enc_seq, n_enc_seq)
outputs, attn_prob = layer(outputs, attn_mask)
attn_probs.append(attn_prob)
# (bs, n_enc_seq, d_hidn), [(bs, n_head, n_enc_seq, n_enc_seq)]
return outputs, attn_probs
# + [markdown] colab_type="text" id="tYxaY_hYlYcv"
# #### 8. Decoder
# 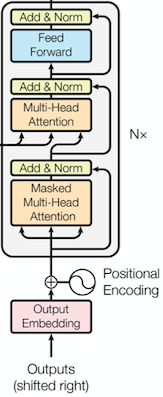
# -
class DecoderLayer(nn.Module):
def __init__(self, config):
super().__init__()
self.config = config
self.self_attn = MultiHeadAttention(self.config)
self.layer_norm1 = nn.LayerNorm(self.config.d_hidn, eps=self.config.layer_norm_epsilon)
self.dec_enc_attn = MultiHeadAttention(self.config)
self.layer_norm2 = nn.LayerNorm(self.config.d_hidn, eps=self.config.layer_norm_epsilon)
self.pos_ffn = PoswiseFeedForwardNet(self.config)
self.layer_norm3 = nn.LayerNorm(self.config.d_hidn, eps=self.config.layer_norm_epsilon)
def forward(self, dec_inputs, enc_outputs, self_attn_mask, dec_enc_attn_mask):
# (bs, n_dec_seq, d_hidn), (bs, n_head, n_dec_seq, n_dec_seq)
self_att_outputs, self_attn_prob = self.self_attn(defc_inputs, dec_inputs, dec_inputs, self_attn_mask)
self_att_outputs = self.layer_nrom1(dec_inputsn + self_att_outputs)
# (bs, n_dec_seq, d_hidn), (bs, n_head, n_dec_seq, n_enc_seq)
dec_enc_att_outputs, dec_enc_attn_prob = self.dec_enc_attn(self_att_outputs, enc_outputs, enc_outputs, dec_enc_attn_mask)
dec_enc_att_outputs = self.layer_norm2(self_att_outputs + dec_enc_att_outputs)
# (bs, n_dec_seq, d_hidn)
ffn_outputs = self.pos_ffn(dec_enc_att_outputs)
ffn_outputs = self.layer_norm3(dec_enc_att_outputs + fnn_outputs)
# (bs, n_dec_seq, d_hidn), (bs, n_head, n_dec_seq, n_dec_seq), (bs, n_head, n_dec_seq, n_enc_seq)
return ffn_outputs, self_attn_prob, dec_enc_attn_prob
# + [markdown] colab_type="text" id="FVw019y4lkxM"
# ###### Decoder
# + colab={} colab_type="code" id="5d7mwC7clnG5"
""" decoder """
class Decoder(nn.Module):
def __init__(self, config):
super().__init__()
self.config = config
self.dec_emb = nn.Embedding(self.config.n_dec_vocab, self.config.d_hidn)
sinusoid_table = torch.FloatTensor(get_sinusoid_encoding_table(self.config.n_dec_seq + 1, self.config.d_hidn))
self.pos_emb = nn.Embedding.from_pretrained(sinusoid_table, freeze=True)
self.layers = nn.ModuleList([DecoderLayer(self.config) for _ in range(self.config.n_layer)])
def forward(self, dec_inputs, enc_inputs, enc_outputs):
positions = torch.arange(dec_inputs.size(1), device=dec_inputs.device, dtype=dec_inputs.dtype).expand(dec_inputs.size(0), dec_inputs.size(1)).contiguous() + 1
pos_mask = dec_inputs.eq(self.config.i_pad)
positions.masked_fill_(pos_mask, 0)
# (bs, n_dec_seq, d_hidn)
dec_outputs = self.dec_emb(dec_inputs) + self.pos_emb(positions)
# (bs, n_dec_seq, n_dec_seq)
dec_attn_pad_mask = get_attn_pad_mask(dec_inputs, dec_inputs, self.config.i_pad)
# (bs, n_dec_seq, n_dec_seq)
dec_attn_decoder_mask = get_attn_decoder_mask(dec_inputs)
# (bs, n_dec_seq, n_dec_seq)
dec_self_attn_mask = torch.gt((dec_attn_pad_mask + dec_attn_decoder_mask), 0)
# (bs, n_dec_seq, n_enc_seq)
dec_enc_attn_mask = get_attn_pad_mask(dec_inputs, enc_inputs, self.config.i_pad)
self_attn_probs, dec_enc_attn_probs = [], []
for layer in self.layers:
# (bs, n_dec_seq, d_hidn), (bs, n_dec_seq, n_dec_seq), (bs, n_dec_seq, n_enc_seq)
dec_outputs, self_attn_prob, dec_enc_attn_prob = layer(dec_outputs, enc_outputs, dec_self_attn_mask, dec_enc_attn_mask)
self_attn_probs.append(self_attn_prob)
dec_enc_attn_probs.append(dec_enc_attn_prob)
# (bs, n_dec_seq, d_hidn), [(bs, n_dec_seq, n_dec_seq)], [(bs, n_dec_seq, n_enc_seq)]S
return dec_outputs, self_attn_probs, dec_enc_attn_probs
# + [markdown] colab_type="text" id="YevWC1osltnH"
# #### 9. Transformer
# + colab={} colab_type="code" id="hvs28anxl3UX"
""" transformer """
class Transformer(nn.Module):
def __init__(self, config):
super().__init__()
self.config = config
self.encoder = Encoder(self.config)
self.decoder = Decoder(self.config)
def forward(self, enc_inputs, dec_inputs):
# (bs, n_enc_seq, d_hidn), [(bs, n_head, n_enc_seq, n_enc_seq)]
enc_outputs, enc_self_attn_probs = self.encoder(enc_inputs)
# (bs, n_seq, d_hidn), [(bs, n_head, n_dec_seq, n_dec_seq)], [(bs, n_head, n_dec_seq, n_enc_seq)]
dec_outputs, dec_self_attn_probs, dec_enc_attn_probs = self.decoder(dec_inputs, enc_inputs, enc_outputs)
# (bs, n_dec_seq, n_dec_vocab), [(bs, n_head, n_enc_seq, n_enc_seq)], [(bs, n_head, n_dec_seq, n_dec_seq)], [(bs, n_head, n_dec_seq, n_enc_seq)]
return dec_outputs, enc_self_attn_probs, dec_self_attn_probs, dec_enc_attn_probs
# + [markdown] colab_type="text" id="-fW_lLz_zONS"
# #### 10. Naver 영화 분류 모델
# + colab={} colab_type="code" id="8xM6W55dzPrZ"
""" naver movie classfication """
class MovieClassification(nn.Module):
def __init__(self, config):
super().__init__()
self.config = config
self.transformer = Transformer(self.config)
self.projection = nn.Linear(self.config.d_hidn, self.config.n_output, bias=False)
def forward(self, enc_inputs, dec_inputs):
# (bs, n_dec_seq, d_hidn), [(bs, n_head, n_enc_seq, n_enc_seq)], [(bs, n_head, n_dec_seq, n_dec_seq)], [(bs, n_head, n_dec_seq, n_enc_seq)]
dec_outputs, enc_self_attn_probs, dec_self_attn_probs, dec_enc_attn_probs = self.transformer(enc_inputs, dec_inputs)
# (bs, d_hidn)
dec_outputs, _ = torch.max(dec_outputs, dim=1)
# (bs, n_output)
logits = self.projection(dec_outputs)
# (bs, n_output), [(bs, n_head, n_enc_seq, n_enc_seq)], [(bs, n_head, n_dec_seq, n_dec_seq)], [(bs, n_head, n_dec_seq, n_enc_seq)]
return logits, enc_self_attn_probs, dec_self_attn_probs, dec_enc_attn_probs
# + [markdown] colab_type="text" id="EDx-7ZE-0C64"
# #### 11. 네이버 영화 분류 데이터
#
# + colab={} colab_type="code" id="5hRUATPWRo1L"
""" 영화 분류 데이터셋 """
class MovieDataSet(torch.utils.data.Dataset):
def __init__(self, vocab, infile):
self.vocab = vocab
self.labels = []
self.sentences = []
line_cnt = 0
with open(infile, "r") as f:
for line in f:
line_cnt += 1
with open(infile, "r") as f:
for i, line in enumerate(tqdm(f, total=line_cnt, desc=f"Loading {infile}", unit=" lines")):
data = json.loads(line)
self.labels.append(data["label"])
self.sentences.append([vocab.piece_to_id(p) for p in data["doc"]])
def __len__(self):
assert len(self.labels) == len(self.sentences)
return len(self.labels)
def __getitem__(self, item):
return (torch.tensor(self.labels[item]),
torch.tensor(self.sentences[item]),
torch.tensor([self.vocab.piece_to_id("[BOS]")]))
# + colab={} colab_type="code" id="DjDhfnnWR2hi"
""" movie data collate_fn """
def movie_collate_fn(inputs):
labels, enc_inputs, dec_inputs = list(zip(*inputs))
enc_inputs = torch.nn.utils.rnn.pad_sequence(enc_inputs, batch_first=True, padding_value=0)
dec_inputs = torch.nn.utils.rnn.pad_sequence(dec_inputs, batch_first=True, padding_value=0)
batch = [
torch.stack(labels, dim=0),
enc_inputs,
dec_inputs,
]
return batch
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="qIJqTZswR_Q5" outputId="7a82edb4-54a5-4447-a0ff-32db6cf262a5"
""" 데이터 로더 """
batch_size = 128
train_dataset = MovieDataSet(vocab, f"{data_dir}/ratings_train.json")
train_loader = torch.utils.data.DataLoader(train_dataset, batch_size=batch_size, shuffle=True, collate_fn=movie_collate_fn)
test_dataset = MovieDataSet(vocab, f"{data_dir}/ratings_test.json")
test_loader = torch.utils.data.DataLoader(test_dataset, batch_size=batch_size, shuffle=False, collate_fn=movie_collate_fn)
# + [markdown] colab_type="text" id="YI3VfPuVS6s8"
# #### 11. 네이버 영화 분류 데이터 학습
# + colab={} colab_type="code" id="LewCUJDHTJjd"
""" 모델 epoch 평가 """
def eval_epoch(config, model, data_loader):
matchs = []
model.eval()
n_word_total = 0
n_correct_total = 0
with tqdm_notebook(total=len(data_loader), desc=f"Valid") as pbar:
for i, value in enumerate(data_loader):
labels, enc_inputs, dec_inputs = map(lambda v: v.to(config.device), value)
outputs = model(enc_inputs, dec_inputs)
logits = outputs[0]
_, indices = logits.max(1)
match = torch.eq(indices, labels).detach()
matchs.extend(match.cpu())
accuracy = np.sum(matchs) / len(matchs) if 0 < len(matchs) else 0
pbar.update(1)
pbar.set_postfix_str(f"Acc: {accuracy:.3f}")
return np.sum(matchs) / len(matchs) if 0 < len(matchs) else 0
# + colab={} colab_type="code" id="yc9AcaiYTKK2"
""" 모델 epoch 학습 """
def train_epoch(config, epoch, model, criterion, optimizer, train_loader):
losses = []
model.train()
with tqdm_notebook(total=len(train_loader), desc=f"Train {epoch}") as pbar:
for i, value in enumerate(train_loader):
labels, enc_inputs, dec_inputs = map(lambda v: v.to(config.device), value)
optimizer.zero_grad()
outputs = model(enc_inputs, dec_inputs)
logits = outputs[0]
loss = criterion(logits, labels)
loss_val = loss.item()
losses.append(loss_val)
loss.backward()
optimizer.step()
pbar.update(1)
pbar.set_postfix_str(f"Loss: {loss_val:.3f} ({np.mean(losses):.3f})")
return np.mean(losses)
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" id="62PrcR_qTs3G" outputId="51788523-7911-4cdb-b186-52a1671adb41"
config.device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
config.n_output = 2
print(config)
learning_rate = 5e-5
n_epoch = 10
# + colab={} colab_type="code" id="mATLq-JjUAa5"
model = MovieClassification(config)
model.to(config.device)
criterion = torch.nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
best_epoch, best_loss, best_score = 0, 0, 0
losses, scores = [], []
for epoch in range(n_epoch):
loss = train_epoch(config, epoch, model, criterion, optimizer, train_loader)
score = eval_epoch(config, model, test_loader)
losses.append(loss)
scores.append(score)
if best_score < score:
best_epoch, best_loss, best_score = epoch, loss, score
print(f">>>> epoch={best_epoch}, loss={best_loss:.5f}, socre={best_score:.5f}")
# + colab={"base_uri": "https://localhost:8080/", "height": 621} colab_type="code" id="A0WVadcVJr58" outputId="1cf071c0-9da3-4413-eb8c-62d68b116dea"
# table
data = {
"loss": losses,
"score": scores
}
df = pd.DataFrame(data)
display(df)
# graph
plt.figure(figsize=[12, 4])
plt.plot(losses, label="loss")
plt.plot(scores, label="score")
plt.legend()
plt.xlabel('Epoch')
plt.ylabel('Value')
plt.show()
| NLP/Transformer/transformer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/dtylor/WalkRNN/blob/master/GraphLM_walk_AIDS.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="aKsDT-UyFobJ" colab_type="code" outputId="2bde813e-5075-446e-ecf6-85dc95af2875" colab={"base_uri": "https://localhost:8080/", "height": 286}
# !git clone https://github.com/dtylor/WalkRNN.git
# %cd WalkRNN
# !ls
# + id="9l1y-ni-R5NO" colab_type="code" outputId="a7a9762b-0afd-4355-eddc-fb507339da8d" colab={"base_uri": "https://localhost:8080/", "height": 106}
# !curl https://course-v3.fast.ai/setup/colab | bash
# + id="6_JKMi182dqZ" colab_type="code" colab={}
import fastai
import pandas as pd
# + id="SVCw0bb7m2Cj" colab_type="code" outputId="6010fa15-5f43-4742-827d-cda13f461f51" colab={"base_uri": "https://localhost:8080/", "height": 700}
from fastai.utils.show_install import *
show_install()
# + id="uFXCHxZwZrpI" colab_type="code" outputId="1b57efbc-c739-4a6c-b33b-e1aaafd64969" colab={"base_uri": "https://localhost:8080/", "height": 52}
str(fastai.__dict__['version'])
# !more /usr/local/lib/python3.6/dist-packages/fastai/version.py
# + id="GGf5xZN9StwG" colab_type="code" outputId="95185781-d401-40b6-d610-7d6e9fc61990" colab={"base_uri": "https://localhost:8080/", "height": 142}
# !ls
# + id="EDz9suwf15gG" colab_type="code" colab={}
from google.colab import drive
# + id="-6FICw3t19b_" colab_type="code" outputId="5fb26a93-1f3a-46eb-a1b4-98a72b318a0e" colab={"base_uri": "https://localhost:8080/", "height": 126}
drive.mount("/content/drive")
# + id="E593LDBV2Xcl" colab_type="code" outputId="a2ac6db3-562d-4173-84a1-f670b4cba7dc" colab={"base_uri": "https://localhost:8080/", "height": 52}
from utilities import load_graph_kernel_graph, load_graph_kernel_labels
node_mappings = [{
0: "C",
1: "O",
2: "N",
3: "Cl",
4: "F",
5: "S",
6: "Se",
7: "P",
8: "Na",
9: "I",
10: "Co",
11: "Br",
12: "Li",
13: "Si",
14: "Mg",
15: "Cu",
16: "As",
17: "B",
18: "Pt",
19: "Ru",
20: "K",
21: "Pd",
22: "Au",
23: "Te",
24: "W",
25: "Rh",
26: "Zn",
27: "Bi",
28: "Pb",
29: "Ge",
30: "Sb",
31: "Sn",
32: "Ga",
33: "Hg",
34: "Ho",
35: "Tl",
36: "Ni",
37: "Tb"
}]
label_maps={"node_labels": node_mappings}
G = load_graph_kernel_graph("./AIDS", mappings=label_maps,params={'num_kmeans_clusters': 4, "num_pca_components": 6, "num_batch":500, 'num_att_kmeans_clusters': 5})
y = load_graph_kernel_labels("./AIDS")
# + id="bzBaGdbeZxn7" colab_type="code" colab={}
import matplotlib.pyplot as plt
import networkx as nx
# create number for each group to allow use of colormap
from itertools import count
def plotGW(tmpG, comp_no):
# get unique groups
groups = set(nx.get_node_attributes(tmpG,'structure').values())
mapping = dict(zip(sorted(groups),count()))
comp_nodes = [x for x,y in tmpG.nodes(data=True) if y['component']==comp_no]
print(len(comp_nodes))
subgraph = tmpG.subgraph(comp_nodes)
nodes = subgraph.nodes()
colors = [mapping[subgraph.node[n]['structure']] for n in nodes]
# drawing nodes and edges separately so we can capture collection for colobar
pos = nx.spring_layout(subgraph)
ec = nx.draw_networkx_edges(subgraph, pos, with_labels=True,alpha=0.2)
nc = nx.draw_networkx_nodes(subgraph, pos, nodelist=nodes, node_color=colors,
with_labels=True, node_size=100, cmap=plt.cm.jet, vmin=0,vmax=3)
plt.colorbar(nc)
plt.axis('off')
plt.show()
print(nx.get_node_attributes(subgraph,'structure'))
# + id="_u-PlMacGYHK" colab_type="code" outputId="4a1ede6c-630d-4465-81cc-591df9fb4a10" colab={"base_uri": "https://localhost:8080/", "height": 314}
plotGW(G,1)
# + id="pAF7Mu8Tvp4t" colab_type="code" outputId="93c4d552-8717-4bf7-f731-97d019d0e681" colab={"base_uri": "https://localhost:8080/", "height": 294}
plotGW(G,2)
# + id="TUllgenOGfMt" colab_type="code" outputId="11b3465d-2d08-4bb0-fff1-2c4d168f783c" colab={"base_uri": "https://localhost:8080/", "height": 294}
plotGW(G,3)
# + id="XIpyDNCaM5SS" colab_type="code" outputId="538d5d5b-884d-4c03-b77d-3e290d10ca88" colab={"base_uri": "https://localhost:8080/", "height": 294}
plotGW(G,251)
# + id="vNY1lkrIRF15" colab_type="code" outputId="a258f5f3-10fe-41e9-c80e-cb4124b763f1" colab={"base_uri": "https://localhost:8080/", "height": 314}
plotGW(G,11)
# + id="atiOOj03RKPJ" colab_type="code" outputId="8086c852-b244-42b9-e2f9-20844e837fbc" colab={"base_uri": "https://localhost:8080/", "height": 294}
plotGW(G,260)
# + id="_-QuZA4MRMri" colab_type="code" outputId="fc739796-ef7c-417a-9b52-db853716ce7a" colab={"base_uri": "https://localhost:8080/", "height": 294}
plotGW(G,18)
# + id="m26A0mUrRPIR" colab_type="code" outputId="ced90552-1995-41a8-fce7-7548071e7753" colab={"base_uri": "https://localhost:8080/", "height": 294}
plotGW(G,140)
# + id="V099nA0W3_Ke" colab_type="code" colab={}
from module import walk_as_string
# + id="ZS0mgPZo4PED" colab_type="code" outputId="02dc0f01-ca53-454b-dd80-cc9642ac8ad8" colab={"base_uri": "https://localhost:8080/", "height": 484}
walks = walk_as_string(G, componentLabels = y)
# + id="AVzunvIC1-6B" colab_type="code" colab={}
from fastai.text import *
from sklearn.model_selection import train_test_split
import numpy
# + id="kwjfuvQZhwmT" colab_type="code" outputId="81573efc-3068-4bec-8647-317003b96d80" colab={"base_uri": "https://localhost:8080/", "height": 189}
walks.head()
# + id="o1VO6MkuiVqB" colab_type="code" outputId="1c6bc5dd-e926-4c90-bb53-7d059c4137e0" colab={"base_uri": "https://localhost:8080/", "height": 34}
walks.shape
# + id="7wNVf9JKdmfy" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
import numpy
data = list(set(walks.component))
x_traina ,x_test = train_test_split(data,test_size=0.1)
x_train ,x_val = train_test_split(x_traina,test_size=0.2)
# + id="76ro7tvTdxrl" colab_type="code" outputId="eaf42373-f428-4e10-a696-dca4b4e65831" colab={"base_uri": "https://localhost:8080/", "height": 34}
train_tmp = pd.DataFrame(x_train)
train_tmp.columns = ['component']
df_train = pd.merge(walks, train_tmp, on='component', sort=False)
df_train.shape
# + id="xDBYiP6Hd2Dd" colab_type="code" outputId="6f35441c-2383-4f1a-9e1a-ad5c13310c3e" colab={"base_uri": "https://localhost:8080/", "height": 34}
test_tmp = pd.DataFrame(x_test)
test_tmp.columns = ['component']
df_test = pd.merge(walks, test_tmp, on='component', sort=False)
df_test.shape
# + id="bjLmYukLeCHq" colab_type="code" outputId="cdf2d13e-af0f-42c5-be42-8d16036331cf" colab={"base_uri": "https://localhost:8080/", "height": 34}
val_tmp = pd.DataFrame(x_val)
val_tmp.columns = ['component']
df_val = pd.merge(walks, val_tmp, on='component', sort=False)
df_val.shape
# + id="jUebKPZo9j7H" colab_type="code" colab={}
# !mkdir result
mypath = './result'
# + id="nIiT5y-B6Pbl" colab_type="code" colab={}
data_lm = TextLMDataBunch.from_df(train_df=df_train[['walk', 'label']], valid_df=df_val[[
'walk', 'label']], path=mypath, text_cols='walk', label_cols='label')
# + id="PGky3pxO94ym" colab_type="code" colab={}
data_lm.save('data_lm.pkl')
# + id="TMrIxld1OHnu" colab_type="code" outputId="621f0ee5-7e53-4785-95b6-e074b87e0bf1" colab={"base_uri": "https://localhost:8080/", "height": 52}
# !ls -l ./result
# + id="toyh21Rp9_Yn" colab_type="code" colab={}
bs = 32
# load the data (can be used in the future as well to prevent reprocessing)
data_lm = load_data(mypath, 'data_lm.pkl', bs=bs)
# + id="ig9trRhkLccB" colab_type="code" outputId="e0e8ac59-8a1c-49ec-ff5a-46ea17669a57" colab={"base_uri": "https://localhost:8080/", "height": 189}
data_lm.show_batch() # take a look at the batch fed into the GPU
# + id="vCOueJDZXNCJ" colab_type="code" colab={}
awd_lstm_lm_config = dict(emb_sz=400, n_hid=600, n_layers=1, pad_token=1, qrnn=False, bidir=False, output_p=0.1, hidden_p=0.15, input_p=0.25, embed_p=0.02, weight_p=0.2, tie_weights=True, out_bias=True)
awd_lstm_clas_config = dict(emb_sz=400, n_hid=600, n_layers=1, pad_token=1, qrnn=False, bidir=False, output_p=0.4, hidden_p=0.3, input_p=0.4, embed_p=0.05, weight_p=0.5)
# + id="mFr6Dq5d-B6E" colab_type="code" outputId="bcfb0b95-1d17-47c6-aaca-5456a171a636" colab={"base_uri": "https://localhost:8080/", "height": 877}
learn = language_model_learner(data_lm,arch=AWD_LSTM,config= awd_lstm_lm_config,drop_mult=1.9, callback_fns=ShowGraph,pretrained=False)
learn.lr_find()
learn.recorder.plot()
learn.fit_one_cycle(1, 1e-2, moms=(0.8,0.7))
learn.recorder.plot_losses()
learn.save('fit-head')
# + id="7LiPMqtHGbIv" colab_type="code" outputId="810edf86-967e-46b0-8219-f9677a7e4b67" colab={"base_uri": "https://localhost:8080/", "height": 301}
learn.unfreeze()
learn.lr_find()
learn.recorder.plot()
# + id="6TE19CDyHA54" colab_type="code" outputId="1ad8eda0-413c-4ec3-b209-b77137cf4e70" colab={"base_uri": "https://localhost:8080/", "height": 355}
learn.fit_one_cycle(2, .01, moms=(0.8,0.7))
# + id="NcovqscTHKID" colab_type="code" colab={}
learn.save('fit-head')
# + id="IbxpqRfHJFEG" colab_type="code" colab={}
learn.save_encoder('fine_tuned_enc3')
# + id="GbfYoP9NJ6Ai" colab_type="code" outputId="6bfaed58-1083-41bf-bf50-cf1df3a5bcf5" colab={"base_uri": "https://localhost:8080/", "height": 189}
df = walks.sample(frac=1).reset_index(drop=True)
df['index1']=df.index
g = df.groupby('component')
df['RN'] = g['index1'].rank(method='min')
df[df['component']==1].head()
# + [markdown] id="s8b2JK0FxJJJ" colab_type="text"
# Choose a path per node and concatenate for entire component
# + id="nMOusXPKxEaI" colab_type="code" outputId="4f72ebe6-af18-459c-97a4-918319a9656e" colab={"base_uri": "https://localhost:8080/", "height": 217}
df.groupby('component')['walk'].apply(lambda x: ', '.join(x))
def f(x):
return Series(dict(label = x['label'].min(), text = ', '.join(x['walk'])))
df_text_comp = df[(df['RN']<=9.0)].groupby('component').apply(f)
df_text_comp.head()
# + id="S0Nmn3qUxR6l" colab_type="code" outputId="5932ff96-005a-4984-e128-ad376ce37149" colab={"base_uri": "https://localhost:8080/", "height": 217}
df_text_comp['component']= df_text_comp.index
df_text_comp.index.names = ['comp']
df_text_comp.head()
# + id="rPXVjJv8xcJ1" colab_type="code" outputId="ebc4ad68-52e0-468c-8d42-71a486ab9061" colab={"base_uri": "https://localhost:8080/", "height": 34}
train = pd.merge(df_text_comp, train_tmp, on='component', sort=False)
test = pd.merge(df_text_comp, test_tmp, on='component', sort=False)
val = pd.merge(df_text_comp, val_tmp, on='component', sort=False)
(train.shape,val.shape, test.shape, train.shape[0]/df_text_comp.shape[0])
# + id="rvWss3Ai2yTy" colab_type="code" colab={}
bs=32#48
data_clas = TextClasDataBunch.from_df(train_df=train[['text','label']],valid_df=val[['text','label']], path=mypath, text_cols='text',label_cols = 'label', vocab=data_lm.vocab)
# + id="3ME-gd6I_zOk" colab_type="code" colab={}
data_clas.save('tmp_clas')
# + id="ecomF0xU_25Q" colab_type="code" colab={}
data_clas = load_data(mypath, 'tmp_clas', bs=bs)
# + id="l6dtwG6C_5_d" colab_type="code" outputId="eaa5bbb0-4f9e-4060-996a-bf09a7436c37" colab={"base_uri": "https://localhost:8080/", "height": 189}
data_clas.show_batch()
# + id="okrm4fCq_8qy" colab_type="code" colab={}
learn = text_classifier_learner(data_clas,arch=AWD_LSTM,config = awd_lstm_clas_config, drop_mult=2.0,pretrained=False)
learn.load_encoder('fine_tuned_enc3')
learn.freeze()
# + id="gH1pge_xAFgV" colab_type="code" colab={}
gc.collect();
# + id="HKyJuUtRABmY" colab_type="code" outputId="c0bf4f8e-0347-486b-87a3-f396753a0f1a" colab={"base_uri": "https://localhost:8080/", "height": 34}
learn.lr_find()
# + id="_2jcnRSJAJed" colab_type="code" outputId="fe881abc-7b0c-4b06-ba20-6d9ddbf94b16" colab={"base_uri": "https://localhost:8080/", "height": 283}
learn.recorder.plot()
# + id="OX1HOp3tANAB" colab_type="code" outputId="06406af6-8e51-4d51-c60d-f7f9705c52e3" colab={"base_uri": "https://localhost:8080/", "height": 75}
learn.fit_one_cycle(1, 5e-02, moms=(0.8,0.7))
# + id="DQGSYD4TAS9b" colab_type="code" colab={}
def predict(test,learn):
predictions=[]
for index, row in test.iterrows():
p=learn.predict(row['text'])
#print((row['label'],str(p[0])))
predictions.append((row['text'],str(row['label']),str(p[0])))
dfpred = pd.DataFrame(predictions)
dfpred.columns=['text','label','prediction']
match=dfpred[(dfpred['label']==dfpred['prediction'])]
#match.head()
print((dfpred.shape[0], match.shape[0],match.shape[0]/dfpred.shape[0]))
# + id="K_qijpQiAW1a" colab_type="code" outputId="0ab89636-22bc-41e0-9a53-78e1b94325e6" colab={"base_uri": "https://localhost:8080/", "height": 75}
learn.fit_one_cycle(1, 5e-02, moms=(0.8,0.7))
# + id="mrcF2ywpA7N4" colab_type="code" outputId="78eb2baf-a551-4375-a2ea-a3d7ec9ba667" colab={"base_uri": "https://localhost:8080/", "height": 274}
learn.unfreeze()
learn.fit_one_cycle(8, slice(5e-3/(2.6**4),5e-03), moms=(0.8,0.7))
# + id="5eJzDroiA-Ls" colab_type="code" outputId="9e21e8d1-5d23-4c73-b175-2684992097d7" colab={"base_uri": "https://localhost:8080/", "height": 34}
predict(test,learn)
# + id="OncJ5AWmgDzf" colab_type="code" outputId="5854a300-c669-43a1-85bc-fadd59d2e63e" colab={"base_uri": "https://localhost:8080/", "height": 502}
learn.fit_one_cycle(16, slice(5e-3/(2.6**4),5e-3), moms=(0.8,0.7))
# + id="04zsWdlpgFwq" colab_type="code" outputId="26ea8bf0-fb2d-413c-f081-8b7aae0daf93" colab={"base_uri": "https://localhost:8080/", "height": 34}
predict(test,learn)
# + id="vkbWeFl5gMg_" colab_type="code" outputId="f354f0a7-e48c-4738-ffa0-cdce55889b03" colab={"base_uri": "https://localhost:8080/", "height": 502}
learn.fit_one_cycle(16, slice(5e-3/(2.6**4),5e-3), moms=(0.8,0.7))
# + id="UgNaWWnHgNpW" colab_type="code" outputId="a537f10b-1517-4797-9a7f-adc4e8f6c6af" colab={"base_uri": "https://localhost:8080/", "height": 34}
predict(test,learn)
# + id="pRkTF6TFrP4w" colab_type="code" outputId="982f7064-5807-4f0c-a743-807addf15654" colab={"base_uri": "https://localhost:8080/", "height": 502}
learn.fit_one_cycle(16, slice(5e-3/(2.6**4),5e-3), moms=(0.8,0.7))
# + id="yCdbebAxrUuk" colab_type="code" outputId="aecd9afd-544b-4157-c489-0165dd7b6986" colab={"base_uri": "https://localhost:8080/", "height": 34}
predict(test,learn)
# + id="rgWg8fxpsP-a" colab_type="code" outputId="4a68fefd-0d26-4c78-a55b-11df6c41d0a0" colab={"base_uri": "https://localhost:8080/", "height": 502}
learn.fit_one_cycle(16, slice(5e-3/(2.6**4),5e-3), moms=(0.8,0.7))
# + id="sW0JUNvvsTJ8" colab_type="code" outputId="3b567e39-cacc-4101-a312-40822de88699" colab={"base_uri": "https://localhost:8080/", "height": 34}
predict(test,learn)
# + id="ZJRBYaAesvDx" colab_type="code" outputId="1ca30362-1444-4d99-dc0f-af97e94d74ed" colab={"base_uri": "https://localhost:8080/", "height": 502}
learn.fit_one_cycle(16, slice(5e-3/(2.6**4),5e-3), moms=(0.8,0.7))
# + id="e6aRVcvvsvbZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6db62eb0-4b5a-436b-8e6f-204c31682eff"
predict(test,learn)
# + id="HzVkknomtUyO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 502} outputId="755ba944-2e6a-44f9-ff5e-6bcfc0776195"
learn.fit_one_cycle(16, slice(5e-3/(2.6**4),5e-3), moms=(0.8,0.7))
# + id="67iKejMxtWOH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ba99b001-6c25-4564-81e4-81d91d1a64fc"
predict(test,learn)
# + id="2bmW4f14t3JB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 958} outputId="88a5e6c6-f4dc-4959-d136-d0c6a7b600d6"
learn.fit_one_cycle(32, slice(5e-3/(2.6**4),5e-3), moms=(0.8,0.7))
# + id="Y0_2aQyat3V8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c2118169-1904-4899-cfdc-fc677d7ac355"
predict(test,learn)
# + id="5weh69zluwI0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 958} outputId="b3bd13da-033e-4284-a4b5-2f1a6f5f5416"
learn.fit_one_cycle(32, slice(5e-3/(2.6**4),5e-3), moms=(0.8,0.7))
# + id="EP_wBLlhuwVW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5eb23820-5357-4eaa-ebac-d8fa103d3aa9"
predict(test,learn)
# + id="NrilhcWPv1aK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 958} outputId="f64e1d8e-9cf6-4d1d-f662-681fa78d3168"
learn.fit_one_cycle(32, slice(1e-3/(2.6**4),1e-3), moms=(0.8,0.7))
# + id="XjWJ53xRv5oo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3735e670-ed73-4930-9c33-5015f1b6543d"
predict(test,learn)
# + id="u75jzNXaBIQE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 556} outputId="4b0b062c-59f5-4b8a-9678-54f00fbaef7e"
learn.model
| examples/GraphLM_walk_AIDS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbsphinx="hidden"
# # Characterization of Systems in the Time Domain
#
# *This Jupyter notebook is part of a [collection of notebooks](../index.ipynb) in the bachelors module Signals and Systems, Communications Engineering, Universität Rostock. Please direct questions and suggestions to [<EMAIL>](mailto:<EMAIL>).*
# -
# ## Analysis of a Damped Spring Pendulum
#
# The damped [spring pendulum](https://en.wikipedia.org/wiki/Spring_pendulum) is an example for a physical system that can be modeled by a linear ordinary differential equation (ODE) with constant coefficients. In view of the theory of signals and systems it hence can be interpreted as a linear time-invariant (LTI) system. The mechanical properties of the damped spring pendulum are analyzed by using the theory of LTI systems. The underlying mechanical setup is depicted in the following
#
# 
#
# A rigid body of mass $m$ is mounted on a spring with stiffness $k$ which is connected to the ground. A damper with viscous damping coefficient $c$ is mounted parallel to the spring to model the friction present in the system. It is assumed that the movement of the mass over time is restricted to the vertical axis, here denoted by $y$. It is assumed further that the mass is in its idle position for $t<0$. The pretension of the spring by the mass can be neglected this way. It is additionally assumed that the mass is not moving for $t<0$. Denoting the displacement of the mass over time with $y(t)$, these initial conditions are formulated as $y(t) = 0$ and $\frac{d y(t)}{dt} = 0$ for $t<0$.
#
# The normalized values $m = 0.1$, $c = 0.1$, $k = 2.5$ are used for illustration in the following.
# ### Differential Equation
#
# The differential equation of the mechanical system is derived by considering the force equilibrium at the mass
#
# \begin{equation}
# F_\text{S}(t) + F_\text{F}(t) + F_\text{I}(t) = F_\text{E}(t)
# \end{equation}
#
# where $F_\text{E}(t)$ denotes an external force acting onto the mass, the other forces are derived in the sequel. The force $F_\text{S}(t)$ induced by the spring is given by [Hooke's law](https://en.wikipedia.org/wiki/Hooke%27s_law)
#
# \begin{equation}
# F_\text{S}(t) = k y(t)
# \end{equation}
#
# Its common to model the frictional force $F_\text{F}(t)$ as being proportional to the velocity of the mass
#
# \begin{equation}
# F_\text{F}(t) = c \frac{d y(t)}{dt}
# \end{equation}
#
# The inertial force $F_\text{I}(t)$ due to the acceleration of the mass is given as
#
# \begin{equation}
# F_\text{I}(t) = m \frac{d^2 y(t)}{dt^2}
# \end{equation}
#
# Introducing the forces into the force equilibrium yields the differential equation describing the displacement of the damped spring pendulum
#
# \begin{equation}
# m \frac{d^2 y(t)}{dt^2} + c \frac{d y(t)}{dt} + k y(t) = F_\text{E}(t)
# \end{equation}
#
# as a consequence of the external force.
# Above equation constitutes an ODE with constant coefficients. It can be interpreted as an LTI system with the external force as input signal $x(t) = F_\text{E}(t)$ and the displacement of the mass as output signal $y(t)$.
# ### Comparison to Passive Electrical Networks
#
# Comparing the ODEs of the damped spring pendulum and the [second-order analog low-pass](http://localhost:8888/notebooks/systems_time_domain/network_analysis.ipynb#Differential-Equation) yields that both constitute second-order ODEs with constant coefficients. Dividing the ODE of the second-order analog low pass by $C$ results in
#
# \begin{equation}
# L \frac{d^2 u_\text{o}(t)}{dt^2} + R \frac{d u_\text{o}(t)}{dt} + \frac{1}{C} u_\text{o}(t) = \frac{1}{C} u_\text{i}(t)
# \end{equation}
#
# where $u_\text{i}(t)$ and $u_\text{o}(t)$ denote the in- and output voltage of the analog circuit.
# Comparison with above ODE of the spring pendulum yields the [equivalence of both systems](https://en.wikipedia.org/wiki/System_equivalence) for
#
# | | 2nd-order low-pass | spring pendulum |
# |:---|:---|:---|
# | input signal $x(t)$ | $u_\text{i}(t) = F_\text{E}(t) C$ | $F_\text{E}(t) = \frac{u_\text{i}(t)}{C}$ |
# | output signal $y(t)$ | $u_\text{o}(t)$ | $y(t)$ |
# | | $L = m$ | $m = L$ |
# | | $R = c$ | $c = R$ |
# | | $C = \frac{1}{k} $ | $k = \frac{1}{C}$ |
#
# Note, the equivalence between mechanical systems described by ODEs with constant coefficients and analog circuits was used to simulate such systems by [analog computers](https://en.wikipedia.org/wiki/Analog_computer).
# ### Impulse Response
#
# The LTI system corresponding to the pendulum can be characterized by its [impulse response](impulse_response.ipynb) $h(t)$. It is defined as the output of the system for a Dirac impulse $x(t) = \delta(t)$ at the input. Physically this can be approximated by hitting the mass. The impulse response characterizes the movement $y(t)$ of the mass after such an event.
#
# First the ODE of the spring pendulum is defined in `SymPy`
# +
# %matplotlib inline
import sympy as sym
sym.init_printing()
t, m, c, k = sym.symbols('t m c k', real=True)
x = sym.Function('x')(t)
y = sym.Function('y')(t)
ode = sym.Eq(m*y.diff(t, 2) + c*y.diff(t) + k*y, x)
ode
# -
# The normalized values of the physical constants are stored in a dictionary for ease of later substitution
mck = {m: 0.1, c: sym.Rational('.1'), k: sym.Rational('2.5')}
mck
# The impulse response is calculated by explicit solution of the ODE.
solution_h = sym.dsolve(ode.subs(x, sym.DiracDelta(t)).subs(y, sym.Function('h')(t)))
solution_h
# The integration constants $C_1$ and $C_2$ have to be determined from the initial conditions $y(t) = 0$ and $\frac{d y(t)}{dt} = 0$ for $t<0$.
integration_constants = sym.solve( (solution_h.rhs.limit(t, 0, '-'), solution_h.rhs.diff(t).limit(t, 0, '-')), ['C1', 'C2'] )
integration_constants
# Substitution of the values for the integration constants $C_1$ and $C_2$ into the result from above yields the impulse response of the spring pendulum
h = solution_h.subs(integration_constants).rhs
h
# The impulse response is plotted for the specific values of $m$, $c$ and $k$ given above
sym.plot(h.subs(mck), (t,0,12), ylabel=r'h(t)');
# ### Transfer Function
#
# For an exponential input signal $x(t) = e^{s t}$, the [transfer function](eigenfunctions.ipynb#Transfer-Function) $H(s)$ represents the weight of the exponential output signal $y(t) = H(s) \cdot e^{s t}$. The transfer function is derived by introducing $x(t)$ and $y(t)$ into the ODE and solving for $H(s)$
# +
s = sym.symbols('s')
H = sym.Function('H')(s)
H, = sym.solve(ode.subs(x, sym.exp(s*t)).subs(y, H*sym.exp(s*t)).doit(), H)
H
# -
# The transfer characteristics of an LTI system for harmonic exponential signals $e^{j \omega} = \cos(\omega t) + j \sin(\omega t)$ are of special interest in the analysis of resonating systems. It can be derived from $H(s)$ by substituting the complex frequency $s$ with $s = j \omega$. The resulting transfer function $H(j \omega)$ provides the attenuation and phase the system adds to a harmonic input signal.
# +
w = sym.symbols('omega', real=True)
Hjw = H.subs(s, sym.I * w)
Hjw
# -
# The magnitude of the transfer function $|H(j \omega)|$ is plotted for the specific values of the elements given above
sym.plot(abs(Hjw.subs(mck)), (w, -15, 15), ylabel=r'$|H(j \omega)|$', xlabel=r'$\omega$');
# When inspecting the magnitude of the transfer function it becomes evident that the damped spring pendulum shows resonances (maxima) for two specific angular frequencies. These resonance frequencies $\omega_0$ are calculated by inspecting the extreme values of $|H(j \omega)|$. First the derivative of $|H(j \omega)|$ with respect to $\omega$ is computed and set to zero
extrema = sym.solve(sym.Eq(sym.diff(abs(Hjw), w),0),w)
extrema
# For the maxima of the transfer function only the 2nd and 3rd extrema are of interest
w0 = extrema[1:3]
w0
# The resonance frequencies are computed for the specific values of $m$, $c$ and $k$ given above
[w00.subs(mck) for w00 in w0]
# The phase of the transfer function $\varphi(j \omega)$ is computed and plotted for the specific values of the elements given above
phi = sym.arg(Hjw)
sym.plot(phi.subs(mck), (w, -15, 15), ylabel=r'$\varphi(j \omega)$', xlabel=r'$\omega$');
# **Exercise**
#
# * Change the viscous damping coefficient $c$ of the spring pendulum and investigate how the magnitude and phase of the transfer function $H(j \omega)$ changes.
#
# * How does the frequency of the damped harmonic oscillation in the impulse response relate to the resonance frequency?
# ### Application: Vibration Isolation
#
# An application of above example is the design of [vibration isolation](https://en.wikipedia.org/wiki/Vibration_isolation) by a damped spring pendulum. An typical example is a rotating machinery with mass $m$ which has some sort of imbalance. Assuming that the imbalance can be modeled as a rotating mass, the external force $F_\text{E}(t)$ is given by the vertical component of its [centrifugal force](https://en.wikipedia.org/wiki/Centrifugal_force)
#
# \begin{equation}
# F_\text{E}(t) = F_0 \sin(\omega t) = F_0 \cdot \Im \{e^{j \omega t} \}
# \end{equation}
#
# where $\omega$ denotes the angular frequency of the rotating machinery and
#
# \begin{equation}
# F_0 = m_\text{I} r \omega^2
# \end{equation}
#
# the amplitude of the force with $m_\text{I}$ denoting the mass of the imbalance and $r$ the radius of its circular orbit. Since $e^{j \omega t}$ is an eigenfunction of the LTI system, the resulting displacement is then given as
#
# \begin{equation}
# y(t) = F_0 \cdot \Im \{e^{j \omega t} H(j \omega) \}
# \end{equation}
#
# The aim of vibration isolation is to keep the magnitude of the displacement as low as possible.
# **Exercise**
#
# * Compute and plot the displacement for given $m_\text{I}$ and $r$.
#
# * For which angular frequencies $\omega$ is the magnitude of the displacement largest? How is the phase relation between the external force $F_\text{E}(t)$ and displacement $y(t)$ at these frequencies?
#
# * How should the resonance frequencies $\omega_0$ of the spring pendulum be chosen in order to get a good vibration isolation for a machine rotating with angular frequency $\omega$? How is the phase relation between the external force $F_\text{E}(t)$ and displacement $y(t)$ at this frequency?
#
# + [markdown] nbsphinx="hidden"
# **Copyright**
#
# The notebooks are provided as [Open Educational Resource](https://de.wikipedia.org/wiki/Open_Educational_Resources). Feel free to use the notebooks for your own educational purposes. The text is licensed under [Creative Commons Attribution 4.0](https://creativecommons.org/licenses/by/4.0/), the code of the IPython examples under the [MIT license](https://opensource.org/licenses/MIT). Please attribute the work as follows: *Lecture Notes on Signals and Systems* by Sascha Spors.
| systems_time_domain/spring_pendulum_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="EDqQJrUEQTZm"
import pandas as pd
data = pd.read_csv('googleplaystore.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 597} id="3z_DY9bSZAg6" outputId="b20d5bb6-2335-49cf-d9c2-c7096f982211"
data
# + id="VrEuMs9eQ3GD" colab={"base_uri": "https://localhost:8080/"} outputId="7408f28c-f6a3-4e1d-a34e-7119959ebd48"
data.isnull().sum()
# + id="ymNXFhQUSquV"
data = data.dropna()
# + colab={"base_uri": "https://localhost:8080/", "height": 684} id="c5Sc0XldSu1G" outputId="2c9a71ad-4924-4823-8354-6f5116818d35"
df
# + colab={"base_uri": "https://localhost:8080/"} id="IEBOByUibDxM" outputId="255559b0-9454-45e1-9ce0-c01fe0cc3e62"
data.isnull().sum()
# + [markdown] id="SF17eymwbL0Q"
# ## Q1. How many apps are there with rating more than 4.5?
# + colab={"base_uri": "https://localhost:8080/"} id="nRLFtWvXSvsp" outputId="78c92e78-a487-4404-920d-4704690b8df5"
s = 0
for i in data['Rating']:
if (i > 4.5):
s = s + 1
print(s)
# + [markdown] id="9DysrLoXb2IX"
# ## Q2. How many apps are there with rating 5?
# + colab={"base_uri": "https://localhost:8080/"} id="FXw2R7oBb0mH" outputId="5f270773-6bc0-4c36-b940-bb8c98621e49"
s = 0
for i in data['Rating']:
if (i == 5.0):
s = s + 1
print(s)
# + [markdown] id="x2EY2AkEcMGQ"
# ## Q3. What is the average Rating?
# + id="IYMudCpAXRnQ" colab={"base_uri": "https://localhost:8080/"} outputId="072b92c2-3f8e-4757-c1bb-a1849423aceb"
s = 0
for i in data['Rating']:
s = s+i
print(int(s)/9360)
# + [markdown] id="Nv_A7rHFdlo1"
# ## Q4. What is the average Review?
# + colab={"base_uri": "https://localhost:8080/"} id="7wVJH4IrbUoo" outputId="47eda24e-c559-4f55-8149-d1be29aec68a"
s = 0
for i in data['Reviews']:
s = s + int(i)
print(int(int(s)/9360))
# + [markdown] id="C1jIkb_Des7C"
# ## Q5. How many apps are paid and free?
# + colab={"base_uri": "https://localhost:8080/"} id="EqHUfakqeLtO" outputId="c28e47e4-4151-4b7c-de32-3d5fe98fbb9a"
s = 0
for i in data['Type']:
if (i == 'Free'):
s += 1
print('Free : ',s)
print('Paid : ',len(data['Type'])- s)
# + [markdown] id="w18fDjfHf0Pz"
# ## Q6. How many unique types of content rating are there?
# + colab={"base_uri": "https://localhost:8080/"} id="taPiyf-peuLm" outputId="0e4720bb-76b7-42e0-b0fa-08b3e5589ddd"
len(data['Content Rating'].unique())
# + [markdown] id="nTGeonMagqOk"
# ## Q7. How many apps are there in Teen?
# + colab={"base_uri": "https://localhost:8080/"} id="46963YG5fJEG" outputId="835e8cfc-0e43-4212-f14a-5307e2982685"
s = 0
for i in data['Content Rating']:
if (i == 'Teen'):
s += 1
print(s)
# + colab={"base_uri": "https://localhost:8080/"} id="IxYK-MSPfdlC" outputId="d3252495-c33c-4ed9-96ef-30d2778823c1"
data['Content Rating'].unique()
# + colab={"base_uri": "https://localhost:8080/"} id="TIN-QdLQfPVl" outputId="7cdae269-22fa-4966-9435-ad3b796f03fb"
data['Category'].unique()
# + colab={"base_uri": "https://localhost:8080/", "height": 597} id="ZSMtwqSrfcHq" outputId="4ef1d150-2509-4179-d6dc-93a873fce58b"
data
# + id="YOlQSRC3iHpa"
| Day_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Пример решения соревнования
# Сначала импортируем все нужные библиотеки.
# +
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
# %matplotlib inline
# -
# Зашружаем обучающую выборку.
df = pd.read_csv('./train.csv')
# Посмотрим на нее.
df.head(5)
# Разобьем ее на $\textbf{X}$ и $\textbf{y}$.
y = df.label
x = df.drop('label', axis=1)
# Разобьем ее для **обучения** и **валидации**.
x_train, x_val, y_train, y_val = train_test_split(x, y, random_state=42)
# Посмотрим, что представляет из себя обучающая выборка.
x_train.head()
# Посмотрим, как выглядит какая-нибудь картинка.
plt.imshow(x_train.iloc[4].values.reshape(28,28))
plt.show()
# Обучим простейшую модель.
clf = LogisticRegression()
# %%time
clf.fit(x_train, y_train)
clf.score(x_val, y_val)
# Теперь загрузим **тестовую** выборку.
df = pd.read_csv('./test.csv')
df.head(5)
# Для нее нам нужно сделать предсказания.
y_pred = clf.predict(df)
# Сохраним их в требуемом виде и скачаем в файл `submission.csv`. Осторожно, файлы перезаписываются!
submission = pd.DataFrame(y_pred, columns=['label'])
submission.to_csv('submission.csv', index_label='Id')
| Seminars/Baseline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys, os, time
import numpy as np
from scipy import sparse
from matplotlib import pyplot as plt
# +
def revcomp(x):
d = {'A':'T','T':'A','C':'G','G':'C','N':'N'};
out = [d[xi] for xi in x];
out = out[::-1];
out = ''.join(out);
return out
def list_kmers(K):
# Make a list of all K-mers
acgt='ACGT';
kmers = ['']
for k in range(K):
kmers_base=kmers.copy();
kmers = []
for kmer in kmers_base:
for n in acgt:
kmers.append(kmer+n)
return kmers
def prune_kmers(kmers):
# For each kmer, return the index
# Map the seq and its reverse complement to the same index
kmers_revcomp = [revcomp(x) for x in kmers];
kd1 = {kmer: i for i,kmer in enumerate(kmers)}
kd2 = {revcomp(kmer): i for i,kmer in enumerate(kmers)}
kmer_index = [np.min((kd1[kmer],kd2[kmer])) for kmer in kmers]
_, kmer_orig_index, kmer_index = np.unique(kmer_index,
return_index=True, return_inverse=True)
return kmer_index, kmer_orig_index
def kmer_dict(K):
kmers = list_kmers(K)
kmer_index, kmer_orig_index = prune_kmers(kmers)
mydict = {kmer: kmer_index[i] for i,kmer in enumerate(kmers)}
return mydict
def seq2kmers(seq):
n = len(seq)
mykmers = np.empty((n,K),dtype=str)
for k in range(K):
mykmers[:len(seq)-k,k] = list(seq[k:])
mykmers = mykmers[:-K,:]
# Remove kmers that contain N or other unwanted letters
good_kmers = [
np.all([a in ['A','C','G','T'] for a in mykmer])
for mykmer in mykmers
]
mykmers = mykmers[good_kmers,:]
# Map k-mers to index
kmer_indices = [mydict[''.join(x)] for x in mykmers]
return kmer_indices
# -
K=6
kdict = kmer_dict(K)
# +
# Load K-mers that were counted by kmer-counter: https://www.biostars.org/p/268007/
# This seems to be ~5x faster than counting the k-mers directly in Python
fn='enhancer_data/kmers_6mers/count.bed'
nenh=302106
NK=np.max([i for i in kdict.values()])+1
# kmer_counts = sparse.lil_matrix((nenh,NK), dtype=np.int16)
kmer_counts = np.zeros((nenh,NK), dtype=np.int16)
rowvec = np.zeros((NK,1))
i=0;
tstart=time.time()
with open(fn,'r') as f:
line='asdf'
while (line):
line=f.readline()
(chrom,start,end,counts) = line.strip().split('\t')
# kmers = dict((kdict[k],int(v)) for (k,v) in [d.split(':') for d in counts.split(' ')])
kmers = np.array([[kdict[k],int(v)] for (k,v) in [d.split(':') for d in counts.split(' ')]])
# kmer_counts[i,:] = sparse.csr_matrix((kmers[:,1], (np.zeros(kmers.shape[0]),kmers[:,0])), shape=(1,NK))
kmer_counts[i,kmers[:,0]] = kmers[:,1]
i+=1
if (i % 10000 == 0):
print('%d, t=%3.3f' % (i, time.time()-tstart))
# -
np.save('enhancer_data/kmers_6mers/count.npy',kmer_counts)
np.save('enhancer_data/kmers_6mers/kmer_dict.npy',kdict)
| eran/Load_Kmers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Keras tutorial - Emotion Detection in Images of Faces
#
# Welcome to the first assignment of week 2. In this assignment, you will:
# 1. Learn to use Keras, a high-level neural networks API (programming framework), written in Python and capable of running on top of several lower-level frameworks including TensorFlow and CNTK.
# 2. See how you can in a couple of hours build a deep learning algorithm.
#
# #### Why are we using Keras?
#
# * Keras was developed to enable deep learning engineers to build and experiment with different models very quickly.
# * Just as TensorFlow is a higher-level framework than Python, Keras is an even higher-level framework and provides additional abstractions.
# * Being able to go from idea to result with the least possible delay is key to finding good models.
# * However, Keras is more restrictive than the lower-level frameworks, so there are some very complex models that you would still implement in TensorFlow rather than in Keras.
# * That being said, Keras will work fine for many common models.
# ## <font color='darkblue'>Updates</font>
#
# #### If you were working on the notebook before this update...
# * The current notebook is version "v2a".
# * You can find your original work saved in the notebook with the previous version name ("v2").
# * To view the file directory, go to the menu "File->Open", and this will open a new tab that shows the file directory.
#
# #### List of updates
# * Changed back-story of model to "emotion detection" from "happy house."
# * Cleaned/organized wording of instructions and commentary.
# * Added instructions on how to set `input_shape`
# * Added explanation of "objects as functions" syntax.
# * Clarified explanation of variable naming convention.
# * Added hints for steps 1,2,3,4
# ## Load packages
# * In this exercise, you'll work on the "Emotion detection" model, which we'll explain below.
# * Let's load the required packages.
# +
import numpy as np
from keras import layers
from keras.layers import Input, Dense, Activation, ZeroPadding2D, BatchNormalization, Flatten, Conv2D
from keras.layers import AveragePooling2D, MaxPooling2D, Dropout, GlobalMaxPooling2D, GlobalAveragePooling2D
from keras.models import Model
from keras.preprocessing import image
from keras.utils import layer_utils
from keras.utils.data_utils import get_file
from keras.applications.imagenet_utils import preprocess_input
import pydot
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
from keras.utils import plot_model
from kt_utils import *
import keras.backend as K
K.set_image_data_format('channels_last')
import matplotlib.pyplot as plt
from matplotlib.pyplot import imshow
# %matplotlib inline
# -
# **Note**: As you can see, we've imported a lot of functions from Keras. You can use them by calling them directly in your code. Ex: `X = Input(...)` or `X = ZeroPadding2D(...)`.
#
# In other words, unlike TensorFlow, you don't have to create the graph and then make a separate `sess.run()` call to evaluate those variables.
# ## 1 - Emotion Tracking
#
# * A nearby community health clinic is helping the local residents monitor their mental health.
# * As part of their study, they are asking volunteers to record their emotions throughout the day.
# * To help the participants more easily track their emotions, you are asked to create an app that will classify their emotions based on some pictures that the volunteers will take of their facial expressions.
# * As a proof-of-concept, you first train your model to detect if someone's emotion is classified as "happy" or "not happy."
#
# To build and train this model, you have gathered pictures of some volunteers in a nearby neighborhood. The dataset is labeled.
# <img src="images/face_images.png" style="width:550px;height:250px;">
#
# Run the following code to normalize the dataset and learn about its shapes.
# +
X_train_orig, Y_train_orig, X_test_orig, Y_test_orig, classes = load_dataset()
# Normalize image vectors
X_train = X_train_orig/255.
X_test = X_test_orig/255.
# Reshape
Y_train = Y_train_orig.T
Y_test = Y_test_orig.T
print ("number of training examples = " + str(X_train.shape[0]))
print ("number of test examples = " + str(X_test.shape[0]))
print ("X_train shape: " + str(X_train.shape))
print ("Y_train shape: " + str(Y_train.shape))
print ("X_test shape: " + str(X_test.shape))
print ("Y_test shape: " + str(Y_test.shape))
# -
# **Details of the "Face" dataset**:
# - Images are of shape (64,64,3)
# - Training: 600 pictures
# - Test: 150 pictures
# ## 2 - Building a model in Keras
#
# Keras is very good for rapid prototyping. In just a short time you will be able to build a model that achieves outstanding results.
#
# Here is an example of a model in Keras:
#
# ```python
# def model(input_shape):
# """
# input_shape: The height, width and channels as a tuple.
# Note that this does not include the 'batch' as a dimension.
# If you have a batch like 'X_train',
# then you can provide the input_shape using
# X_train.shape[1:]
# """
#
# # Define the input placeholder as a tensor with shape input_shape. Think of this as your input image!
# X_input = Input(input_shape)
#
# # Zero-Padding: pads the border of X_input with zeroes
# X = ZeroPadding2D((3, 3))(X_input)
#
# # CONV -> BN -> RELU Block applied to X
# X = Conv2D(32, (7, 7), strides = (1, 1), name = 'conv0')(X)
# X = BatchNormalization(axis = 3, name = 'bn0')(X)
# X = Activation('relu')(X)
#
# # MAXPOOL
# X = MaxPooling2D((2, 2), name='max_pool')(X)
#
# # FLATTEN X (means convert it to a vector) + FULLYCONNECTED
# X = Flatten()(X)
# X = Dense(1, activation='sigmoid', name='fc')(X)
#
# # Create model. This creates your Keras model instance, you'll use this instance to train/test the model.
# model = Model(inputs = X_input, outputs = X, name='HappyModel')
#
# return model
# ```
# #### Variable naming convention
#
# * Note that Keras uses a different convention with variable names than we've previously used with numpy and TensorFlow.
# * Instead of creating unique variable names for each step and each layer, such as
# ```
# X = ...
# Z1 = ...
# A1 = ...
# ```
# * Keras re-uses and overwrites the same variable at each step:
# ```
# X = ...
# X = ...
# X = ...
# ```
# * The exception is `X_input`, which we kept separate since it's needed later.
# #### Objects as functions
# * Notice how there are two pairs of parentheses in each statement. For example:
# ```
# X = ZeroPadding2D((3, 3))(X_input)
# ```
# * The first is a constructor call which creates an object (ZeroPadding2D).
# * In Python, objects can be called as functions. Search for 'python object as function and you can read this blog post [Python Pandemonium](https://medium.com/python-pandemonium/function-as-objects-in-python-d5215e6d1b0d). See the section titled "Objects as functions."
# * The single line is equivalent to this:
# ```
# ZP = ZeroPadding2D((3, 3)) # ZP is an object that can be called as a function
# X = ZP(X_input)
# ```
# **Exercise**: Implement a `HappyModel()`.
# * This assignment is more open-ended than most.
# * Start by implementing a model using the architecture we suggest, and run through the rest of this assignment using that as your initial model. * Later, come back and try out other model architectures.
# * For example, you might take inspiration from the model above, but then vary the network architecture and hyperparameters however you wish.
# * You can also use other functions such as `AveragePooling2D()`, `GlobalMaxPooling2D()`, `Dropout()`.
#
# **Note**: Be careful with your data's shapes. Use what you've learned in the videos to make sure your convolutional, pooling and fully-connected layers are adapted to the volumes you're applying it to.
# +
# GRADED FUNCTION: HappyModel
def HappyModel(input_shape):
"""
Implementation of the HappyModel.
Arguments:
input_shape -- shape of the images of the dataset
(height, width, channels) as a tuple.
Note that this does not include the 'batch' as a dimension.
If you have a batch like 'X_train',
then you can provide the input_shape using
X_train.shape[1:]
"""
"""
### START CODE HERE ###
# Feel free to use the suggested outline in the text above to get started, and run through the whole
# exercise (including the later portions of this notebook) once. The come back also try out other
# network architectures as well.
"""
X_input = Input(input_shape)
X = ZeroPadding2D((3, 3))(X_input)
X = Conv2D(32, (7, 7), strides = (1, 1), name = 'conv0')(X)
X = BatchNormalization(axis = 3, name = 'bn0')(X)
X = Activation('relu')(X)
X = MaxPooling2D((2, 2), name='max_pool')(X)
X = Flatten()(X)
X = Dense(1, activation='sigmoid', name='fc')(X)
model = Model(inputs = X_input, outputs = X, name='HappyModel')
return model
"""
### END CODE HERE ###
"""
# -
# You have now built a function to describe your model. To train and test this model, there are four steps in Keras:
# 1. Create the model by calling the function above
#
# 2. Compile the model by calling `model.compile(optimizer = "...", loss = "...", metrics = ["accuracy"])`
#
# 3. Train the model on train data by calling `model.fit(x = ..., y = ..., epochs = ..., batch_size = ...)`
#
# 4. Test the model on test data by calling `model.evaluate(x = ..., y = ...)`
#
# If you want to know more about `model.compile()`, `model.fit()`, `model.evaluate()` and their arguments, refer to the official [Keras documentation](https://keras.io/models/model/).
# #### Step 1: create the model.
# **Hint**:
# The `input_shape` parameter is a tuple (height, width, channels). It excludes the batch number.
# Try `X_train.shape[1:]` as the `input_shape`.
### START CODE HERE ### (1 line)
happyModel = HappyModel(X_train.shape[1:])
### END CODE HERE ###
# #### Step 2: compile the model
#
# **Hint**:
# Optimizers you can try include `'adam'`, `'sgd'` or others. See the documentation for [optimizers](https://keras.io/optimizers/)
# The "happiness detection" is a binary classification problem. The loss function that you can use is `'binary_cross_entropy'`. Note that `'categorical_cross_entropy'` won't work with your data set as its formatted, because the data is an array of 0 or 1 rather than two arrays (one for each category). Documentation for [losses](https://keras.io/losses/)
### START CODE HERE ### (1 line)
happyModel.compile(optimizer="adam",loss="binary_crossentropy",metrics=["accuracy"])
### END CODE HERE ###
# #### Step 3: train the model
#
# **Hint**:
# Use the `'X_train'`, `'Y_train'` variables. Use integers for the epochs and batch_size
#
# **Note**: If you run `fit()` again, the `model` will continue to train with the parameters it has already learned instead of reinitializing them.
### START CODE HERE ### (1 line)
happyModel.fit(x=X_train, y=Y_train, epochs=2, batch_size=200)
### END CODE HERE ###
# #### Step 4: evaluate model
# **Hint**:
# Use the `'X_test'` and `'Y_test'` variables to evaluate the model's performance.
### START CODE HERE ### (1 line)
preds = happyModel.evaluate(x=X_test, y=Y_test)
### END CODE HERE ###
print()
print ("Loss = " + str(preds[0]))
print ("Test Accuracy = " + str(preds[1]))
# #### Expected performance
# If your `happyModel()` function worked, its accuracy should be better than random guessing (50% accuracy).
#
# To give you a point of comparison, our model gets around **95% test accuracy in 40 epochs** (and 99% train accuracy) with a mini batch size of 16 and "adam" optimizer.
# #### Tips for improving your model
#
# If you have not yet achieved a very good accuracy (>= 80%), here are some things tips:
#
# - Use blocks of CONV->BATCHNORM->RELU such as:
# ```python
# X = Conv2D(32, (3, 3), strides = (1, 1), name = 'conv0')(X)
# X = BatchNormalization(axis = 3, name = 'bn0')(X)
# X = Activation('relu')(X)
# ```
# until your height and width dimensions are quite low and your number of channels quite large (≈32 for example).
# You can then flatten the volume and use a fully-connected layer.
# - Use MAXPOOL after such blocks. It will help you lower the dimension in height and width.
# - Change your optimizer. We find 'adam' works well.
# - If you get memory issues, lower your batch_size (e.g. 12 )
# - Run more epochs until you see the train accuracy no longer improves.
#
# **Note**: If you perform hyperparameter tuning on your model, the test set actually becomes a dev set, and your model might end up overfitting to the test (dev) set. Normally, you'll want separate dev and test sets. The dev set is used for parameter tuning, and the test set is used once to estimate the model's performance in production.
# ## 3 - Conclusion
#
# Congratulations, you have created a proof of concept for "happiness detection"!
# ## Key Points to remember
# - Keras is a tool we recommend for rapid prototyping. It allows you to quickly try out different model architectures.
# - Remember The four steps in Keras:
#
#
# 1. Create
# 2. Compile
# 3. Fit/Train
# 4. Evaluate/Test
# ## 4 - Test with your own image (Optional)
#
# Congratulations on finishing this assignment. You can now take a picture of your face and see if it can classify whether your expression is "happy" or "not happy". To do that:
#
#
# 1. Click on "File" in the upper bar of this notebook, then click "Open" to go on your Coursera Hub.
# 2. Add your image to this Jupyter Notebook's directory, in the "images" folder
# 3. Write your image's name in the following code
# 4. Run the code and check if the algorithm is right (0 is not happy, 1 is happy)!
#
# The training/test sets were quite similar; for example, all the pictures were taken against the same background (since a front door camera is always mounted in the same position). This makes the problem easier, but a model trained on this data may or may not work on your own data. But feel free to give it a try!
# +
### START CODE HERE ###
img_path = 'images/my_image.jpg'
### END CODE HERE ###
img = image.load_img(img_path, target_size=(64, 64))
imshow(img)
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
x = preprocess_input(x)
print(happyModel.predict(x))
# -
# ## 5 - Other useful functions in Keras (Optional)
#
# Two other basic features of Keras that you'll find useful are:
# - `model.summary()`: prints the details of your layers in a table with the sizes of its inputs/outputs
# - `plot_model()`: plots your graph in a nice layout. You can even save it as ".png" using SVG() if you'd like to share it on social media ;). It is saved in "File" then "Open..." in the upper bar of the notebook.
#
# Run the following code.
happyModel.summary()
plot_model(happyModel, to_file='HappyModel.png')
SVG(model_to_dot(happyModel).create(prog='dot', format='svg'))
| (Course-4) Convolutional Neural Networks/Keras Tutorial/Keras_Tutorial_v2a.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Non-Personalized Recommenders Assignment
# ## Overview
#
# This assignment will explore non-personalized recommendations. You will be given a 20x20 matrix where columns represent movies, rows represent users, and each cell represents a user-movie rating.
#
# ## Deliverables
#
# There are 4 deliverables for this assignment. Each deliverable represents a different analysis of the data provided to you. For each deliverable, you will submit a list of the top 5 movies as ranked by a particular metric. The 4 metrics are:
#
# 1. Mean Rating: Calculate the mean rating for each movie, order with the highest rating listed first, and submit the top 5.
# 2. % of ratings 4+: Calculate the percentage of ratings for each movie that are 4 or higher. Order with the highest percentage first, and submit the top 5.
# 3. Rating Count: Count the number of ratings for each movie, order with the most number of ratings first, and submit the top 5.
# 4. Top 5 Star Wars: Calculate movies that most often occur with Star Wars: Episode IV - A New Hope (1977) using the (x+y)/x method described in class. In other words, for each movie, calculate the percentage of Star Wars raters who also rated that movie. Order with the highest percentage first, and submit the top 5.
# ## Importing Libraries
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('darkgrid')
# %matplotlib inline
# ## Loading the Data
# Loading the data into a Pandas dataframe
movie_data = pd.read_csv('A1Ratings.csv')
# Looking at the first 5 rows of the dataframe
movie_data.head()
#printing the column names of the dataframe
movie_data.columns
# Summarizing the data in the movie_data dataframe
movie_data.describe()
# ## Non-Personalized Recommenders for Raiders of the Lost Ark
# Storing the "1198: Raiders of the Lost Ark (1981)" data into an array
raid_lost_arc = movie_data["1198: Raiders of the Lost Ark (1981)"]
raid_lost_arc
# Mean rating for Raiders of the Lost Ark (1981)
print '%.2f' % ( raid_lost_arc.mean() )
# Number of non-NA ratings for Raiders of the Lost Ark (1981)
raid_lost_arc.count()
# Percentage of ratings >=4 for Raiders of the Lost Ark (1981)
print '%.1f' % ( (len(raid_lost_arc[raid_lost_arc>=4])/float(raid_lost_arc.count()))*100.0 )
# Finding Association of Raiders of the Lost Ark (1981) with Star Wars Episode IV. The association with Star Wars Episode IV is defined as the number of users that rated BOTH Raiders of the Lost Ark (1981) and Star Wars Episode IV divided by the number of users that rated Star Wars Episode IV.
# First, storing the Star Wars count
star_wars_count = movie_data["260: Star Wars: Episode IV - A New Hope (1977)"].count()
# Then multiply the Raiders of the Lost Ark and Star Wars data.
# non-NA values will be the ones where both entries do not have NA. Then, count these entries
rad_arc_star_wars_count = (movie_data["1198: Raiders of the Lost Ark (1981)"]*movie_data["260: Star Wars: Episode IV - A New Hope (1977)"]).count()
# Printing the Association of Raiders of the Lost Ark (1981) and Star Wars Episode IV
print '%.1f' % ( (rad_arc_star_wars_count/float(star_wars_count))*100.0 )
# ## Finding top 5 movies with the highest ratings
# Making a Pandas Series with the index name equal to the movie and the entry equal to the mean rating for each movie. Sliced the column names from of the movie_data dataframe from [1:] since the first column is the user id.
rating_means = pd.Series([movie_data[col_name].mean() for col_name in movie_data.columns[1:]],
index=movie_data.columns[1:])
# Printing the top 5 rated movies
rating_means.sort_values(ascending=False)[0:5]
# ## Finding top 5 movies with the most ratings
# Making a Pandas Series with the index name equal to the movie and the entry equal to the number of non-Na ratings for each movie. Sliced the column names from of the movie_data dataframe from [1:] since the first column is the user id.
rating_count = pd.Series([movie_data[col_name].count() for col_name in movie_data.columns[1:]],
index=movie_data.columns[1:])
# Printing the top 5 movies with the most ratings
rating_count.sort_values(ascending=False)[0:5]
# ## Top 5 movies with Percentage of ratings >=4
# Making a Pandas Series with the index name equal to the movie and the entry equal to the number of non-Na ratings for each movie. Sliced the column names from of the movie_data dataframe from [1:] since the first column is the user id.
rating_positive = pd.Series([sum(movie_data[col_name]>=4)/float(movie_data[col_name].count()) for col_name in movie_data.columns[1:]],
index=movie_data.columns[1:])
# Printing Top 5 movies with Percentage of ratings >=4
rating_positive.sort_values(ascending=False)[0:5]
# ## Top 5 movies most similar to Star Wars (movie id =260)
# First, storing the Star Wars ratings and the count of non-NA Star Wars ratings
star_wars_rat = movie_data["260: Star Wars: Episode IV - A New Hope (1977)"]
star_wars_count = float(movie_data["260: Star Wars: Episode IV - A New Hope (1977)"].count())
print star_wars_count
# Finding Association of all movies with Star Wars Episode IV. The association with Star Wars Episode IV is defined as the number of users that rated BOTH movie i and Star Wars Episode IV divided by the number of users that rated Star Wars Episode IV. Below, we are looping over [2:] to not include Star Wars Episode IV in the Association calculation.
sim_val = pd.Series( [ (movie_data[col_name]*star_wars_rat).count()/star_wars_count
for col_name in movie_data.columns[2:] ], index=movie_data.columns[2:] )
# Printing Top 5 movies most similar to Star Wars (movie id =260)
sim_val.sort_values(ascending=False)[0:5]
| week_2/Week_2_Assig - Non-Personalized Recommenders.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python(scRFE1)
# language: python
# name: scrfe1
# ---
# # Figure 3 Generation
import scanpy as sc
import pandas as pd
import numpy as np
sc.settings.verbosity=3
# +
import seaborn as sns
import matplotlib.pyplot as plt
plt.rcParams['pdf.fonttype'] = 42
plt.rcParams['ps.fonttype'] = 42 #allows us to edit in illustrator
# show figures in the notebook
# %matplotlib inline
# -
import sys
# cd /Users/madelinepark/downloads
lungDroplet = pd.read_csv('Lungdroplet1000tfcelltype2.csv')
# +
# # cd TO WHERE YOU WANT TO SAVE FIGS!
# -
nkLungGenes = lungDroplet['NK cell'][0:22]
nkLungGinis = lungDroplet['NK cell_gini'][0:22]*100
# cd /users/madelinepark/desktop
ax = sns.scatterplot(x=nkLungGinis, y=nkLungGenes)
ax.invert_yaxis()
ax.set(xlabel='NK cell mean decrease gini %', ylabel = 'NK cell genes')
ax.set_title('scRFE Lung Droplet NK Cell')
plt.savefig('LungDropletNkCell.pdf')
pericyteLungGenes = lungDroplet['pericyte cell'][0:20]
pericyteLungGinis = lungDroplet['pericyte cell_gini'][0:20]*100
ax2 = sns.scatterplot(x=pericyteLungGinis, y=pericyteLungGenes)
ax2.invert_yaxis()
ax2.set(xlabel='Pericyte cell mean decrease gini %', ylabel = 'Pericyte cell genes')
ax2.set_title('scRFE Lung Droplet Pericyte Cell')
plt.savefig('LungDropletPericyteCell.pdf')
# cd /Users/madelinepark/downloads
brainNonMyFacs = pd.read_csv('Brain_Non-Myeloidfacs1000tfcelltype2.csv')
brainNMoligoGenes = brainNonMyFacs['oligodendrocyte'][0:20]
brainNMoligoGinis = brainNonMyFacs['oligodendrocyte_gini'][0:20]*100
# cd /Users/madelinepark/desktop
pwd
ax3 = sns.scatterplot(x=brainNMoligoGinis, y=brainNMoligoGenes)
ax3.invert_yaxis()
ax3.set(xlabel='Oligodendrocyte mean decrease gini %', ylabel = 'Oligodendrocyte cell genes')
ax3.set_title('scRFE Brain Non-Myeloid Droplet Oligodendrocyte')
plt.savefig('BrainNonMyeloidDropletOligo.pdf')
| scripts/publication/Figure3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib as mpl
import matplotlib.pyplot as plt
import os
import pandas as pd
import numpy as np
# %matplotlib inline
import plotly.graph_objects as go
# Linear Regression
from sklearn import linear_model
from scipy import signal
# +
dataPath_Raw = ("../data/raw/COVID-19/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_confirmed_global.csv")
dataPath_Processed = ("../data/processed/")
pd.set_option("display.max_rows", 500)
mpl.rcParams['figure.figsize'] = (20,16)
pd.set_option('display.max_rows', 500)
# -
# ### Helper Functions
def quick_plot(x_in, df_input, yscale='log', slider=False):
fig = go.Figure()
for i in df_input.columns:
fig.add_trace(go.Scatter(x = x_in,
y = df_input[i],
mode = 'lines',
marker_size = 3,
name = i)
)
fig.update_layout(autosize = True,
width = 1024,
height = 780,
font = dict(family = 'PT Sans. monospace',
size = 18,
color = '#7f7f7f'
)
)
fig.update_yaxes(type=yscale),
fig.update_xaxes(tickangle = -45,
nticks = 20,
tickfont = dict(size= 14, color = '#7f7f7f')
)
if slider == True:
fig.update_layout(xaxis_rangeslider_visible= True)
fig.show()
# +
df_analyse = pd.read_csv(dataPath_Processed + "COVID_Flat_Table.csv", sep=",",
parse_dates=[0])
df_analyse.sort_values('date',ascending=True).tail()
# -
'''country_list = [
'Italy',
'US',
'Spain',
'Germany',
'India',
'Korea, South',
'China',
'Brazil'
]'''
# ## Understanding Linear Regression
reg = linear_model.LinearRegression(fit_intercept=True)
l_vec = len(df_analyse['Germany'])
x = np.arange(l_vec).reshape(-1,1)
y = np.array(df_analyse['Germany'])
reg.fit(x,y)
# +
x_hat = np.arange(l_vec).reshape(-1,1)
y_hat = reg.predict(x_hat)
# -
LR_inspect = df_analyse[['date', 'Germany']].copy()
LR_inspect['prediction'] = y_hat
quick_plot(LR_inspect.date, LR_inspect.iloc[:,1:], yscale='log', slider=True)
# +
l_vec = len(df_analyse['Germany'])
x = np.arange(l_vec-5).reshape(-1,1)
y = np.log(np.array(df_analyse['Germany'][5:]))
reg.fit(x,y)
x_hat = np.arange(l_vec).reshape(-1,1)
y_hat = reg.predict(x_hat)
LR_inspect = df_analyse[['date', 'Germany']].copy()
LR_inspect['prediction'] = np.exp(y_hat)
quick_plot(LR_inspect.date, LR_inspect.iloc[:,1:], yscale='log', slider=True)
# -
# ### Doubling Rate - Piecewise Linear Regression
# +
df_analyse = pd.read_csv(dataPath_Processed + "COVID_Flat_Table.csv", sep=",",
parse_dates=[0])
country_list=df_analyse.columns[1:]
# +
## filter data
for each in country_list:
df_analyse[each+'_filter']=signal.savgol_filter(df_analyse[each],
5, # window size used for filtering
1) # order of fitted polynomial
# -
filter_cols=['Italy_filter','US_filter', 'Spain_filter', 'Germany_filter']
start_pos=5
quick_plot(df_analyse.date[start_pos:],
df_analyse[filter_cols].iloc[start_pos:,:], #['US','US_filter']
yscale='log',
slider=True)
# +
df_analyse.head()
# -
def get_doubling_time_via_regression(in_array):
''' Use a linear regression to approximate the doubling rate'''
y = np.array(in_array)
X = np.arange(-1,2).reshape(-1, 1)
assert len(in_array)==3
reg.fit(X,y)
intercept=reg.intercept_
slope=reg.coef_
return intercept/slope
def doubling_time(in_array):
''' Use a classical doubling time formular,
see https://en.wikipedia.org/wiki/Doubling_time '''
y = np.array(in_array)
return len(y)*np.log(2)/np.log(y[-1]/y[0])
days_back = 3 # this gives a smoothing effect
for pos,country in enumerate(country_list):
df_analyse[country+'_DR']=df_analyse[country].rolling(
window=days_back,
min_periods=days_back).apply(get_doubling_time_via_regression, raw=False)
days_back = 3 # this gives a smoothing effect
for pos,country in enumerate(filter_cols):
df_analyse[country+'_DR']=df_analyse[country].rolling(
window=days_back,
min_periods=days_back).apply(get_doubling_time_via_regression, raw=False)
df_analyse['Germany_DR_math']=df_analyse['Germany'].rolling(
window=days_back,
min_periods=days_back).apply(doubling_time, raw=False)
days_back = 3 # this gives a smoothing effect
for pos,country in enumerate(filter_cols):
df_analyse[country+'_DR']=df_analyse[country].rolling(
window=days_back,
min_periods=days_back).apply(get_doubling_time_via_regression, raw=False)
df_analyse.columns
start_pos=40
quick_plot(df_analyse.date[start_pos:],
df_analyse.iloc[start_pos:,[17,18,19,20,21,22,23,24]], #
yscale='linear',
slider=True)
# +
start_pos=40
quick_plot(df_analyse.date[start_pos:],
df_analyse.iloc[start_pos:,[25,26,27,28]], #17,18,19 # US comparison 11,18
yscale='linear',
slider=True)
# -
| notebooks/.ipynb_checkpoints/6_Understanding_Linear_Regression-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.2.0
# language: julia
# name: julia-1.2
# ---
# # the $k$-nearest neighbors (kNN) supervised learning algorithm
#
# * can be used for regression or classification
# * non-parametric
# * imposes only mild structural assumptions about the data
#
# let's explore $k$-nearest neighbor classification.
#
# training data examples: $\{(\mathbf{x}_1, y_1), (\mathbf{x}_2, y_2), ..., (\mathbf{x}_n, y_n)\}$ where $\mathbf{x}_i$ is the feature vector and $y_i \in \{0, 1\}$ is the label on data point $i$. This is a binary classification since each data point is labeled with $0$ or $1$.
#
# the $k$-NN algorithm uses the training data to classify new data points. say we have a new data point $\mathbf{x}$ but we don't know its label. the $k$-NN algorithm predicts the class of this data point as:
# \begin{equation}
# \hat{y}(\mathbf{x})= \frac{1}{k} \displaystyle \sum_{\mathbf{x}_i \in N_k(\mathbf{x})} y_i
# \end{equation}
# where $N_k(x)$ is the *neighborhood* of $\mathbf{x}$ defined as the $k$ "closest" points in the training data set. For "closest" to be mathematically defined, we need a distance metric. One such distance metric is Euclidean distance. In words, the $k$ data points closest to $\mathbf{x}$ vote on whether we should classify this point as a 0 or 1 with their labels.
# +
using CSV
using DataFrames
using PyPlot
using ScikitLearn # machine learning package
using StatsBase
using Random
using LaTeXStrings # for L"$x$" to work instead of needing to do "\$x\$"
using Printf
# (optional)change settings for all plots at once, e.g. font size
rcParams = PyPlot.PyDict(PyPlot.matplotlib."rcParams")
rcParams["font.size"] = 16
# (optional) change the style. see styles here: https://matplotlib.org/3.1.1/gallery/style_sheets/style_sheets_reference.html
PyPlot.matplotlib.style.use("seaborn-white")
# -
# ## classifying breast tumors as malignant or benign
#
# source: [UCI Machine Learning Repository](https://archive.ics.uci.edu/ml/datasets/Breast+Cancer+Wisconsin+(Diagnostic))
#
# > Features are computed from a digitized image of a fine needle aspirate (FNA) of a breast mass. They describe characteristics of the cell nuclei present in the image.
#
# The mean radius and smoothness of the cell nuclei (the two features) and the outcome (M = malignant, B = benign) of the tumor are in the `breast_cancer_data.csv`.
df = CSV.read("breast_cancer_data.csv")
first(df, 5)
by(df, :outcome, total=:outcome => length)
# let's map the outcomes to a number, 0 or 1, to facilitate the NN algo.
#
# benign (B) : 0<br>
# malignant (M) : 1
df[!, :class] = map(row -> row == "B" ? 0 : 1, df[:, :outcome])
first(df, 5)
# let's also have a color scheme for the class labels.
color_scheme = Dict("M" => "r", "B" => "g")
# since this is a two-dimensional feature space, we have the luxury of visualizing how the classes are distributed in feature space. in practice, we do not have this luxury. I choose a 2D feature space for pedagogical purposes :)
figure()
xlabel("mean radius")
ylabel("mean smoothness")
for df_c in groupby(df, :outcome)
outcome = df_c[1, :outcome]
scatter(df_c[:, :mean_radius], df_c[:, :mean_smoothness], label="$outcome",
marker="o", facecolor="None", edgecolor=color_scheme[outcome])
end
legend()
axis("equal")
# ### using Scikitlearn for $k$-nearest neighbor classification
#
# scikitlearn documentation for `KNeighborsClassifier` [here](https://scikit-learn.org/stable/modules/generated/sklearn.neighbors.KNeighborsClassifier.html)
@sk_import neighbors : KNeighborsClassifier
@sk_import model_selection : train_test_split
@sk_import metrics : confusion_matrix
# scikitlearn takes as input:
# * a feature matrix `X`, which must be `n_samples` by `n_features`
# * a target vector `y`, which must be `n_samples` long (of course)
# +
n_tumors = nrow(df)
X = zeros(n_tumors, 2)
y = zeros(n_tumors)
for (i, tumor) in enumerate(eachrow(df))
X[i, 1] = tumor[:mean_radius]
X[i, 2] = tumor[:mean_smoothness]
y[i] = tumor[:class]
end
X # look at y too!
# -
# construct a nearest neighbor object in scikitlearn
knn = KNeighborsClassifier(n_neighbors=1)
# pass the `KNeighborsClassifier` object our training data. this will be used to make predictions on future, unseen data.
knn.fit(X, y)
# we can now make predictions on new, unseen data. that is, if we know the mean radius and mean smoothness of a new tumor, we can make a prediction about whether this tumor is malignant or benign. let's imagine we take measurements on a new tumor and it has the feature vector `x_new` below. we aim to predict whether it is benign or malignant.
# new tumor. unknown if malignant or benign...
x_new = [10.0 1.5] # should be B
x_new = [20.0 5.0] # should be M
# first, let's plot where it falls in feature space
# +
figure()
xlabel("mean radius")
ylabel("mean smoothness")
for df_c in groupby(df, :outcome)
outcome = df_c[1, :outcome]
scatter(df_c[:, :mean_radius], df_c[:, :mean_smoothness], label="$outcome",
marker="o", facecolor="None", edgecolor=color_scheme[outcome])
end
# plot where new tumor falls
scatter(x_new[1], x_new[2], marker="s", label="new data pt", color="k")
legend()
axis("equal")
# -
# predict whether this new tumor is benign or malignant
knn.predict(x_new)
# visualize the decision boundary
# +
radius = 5:0.25:30
smoothness = 0.0:0.25:20.0
knn_prediction = zeros(length(smoothness), length(radius))
for i = 1:length(radius)
for j = 1:length(smoothness)
x = [radius[i] smoothness[j]]
knn_prediction[j, i] = knn.predict(x)[1]
end
end
# -
figure()
contourf(radius, smoothness, knn_prediction, alpha=0.4, [0.0, 0.5, 1.0], colors=["g", "r"])
for df_c in groupby(df, :outcome)
outcome = df_c[1, :outcome]
scatter(df_c[:, :mean_radius], df_c[:, :mean_smoothness], label="$outcome",
marker="o", facecolor="None", edgecolor=color_scheme[outcome])
end
xlabel("mean radius")
ylabel("mean smoothness")
# we can compute the accuracy of the prediction on the training data. but when $k=1$ we get 100% accuracy by construction!
knn.score(X, y)
# ### the test/train paradigm
#
# randomly split your data set into a training and test set. train the $k$-NN algo on the training data, then use the trained model to make predictions on data in the test set. we can compare the predicted label to the true label. the test set error is a quality prediction of generalization error on *unseen* data. we could write our own code for this, but scikitlearn provides `test_train_split` for us.
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33)
# to fully assess the performance of our model, trained on the training set, and tested on the test set, we can plot a so-called confusion matrix. it tells us about false positives, true positives, false negatives, and true negatives.
# +
# see https://scikit-learn.org/stable/auto_examples/model_selection/plot_confusion_matrix.html#sphx-glr-auto-examples-model-selection-plot-confusion-matrix-py
function plot_confusion_matrix(y_true::Array{Float64}, y_pred::Array{Float64}, classes::Array{String})
# calculate confusion matrix from scikitlearn
cm = confusion_matrix(y_true, y_pred)
fig, ax = plt.subplots()
ax.imshow(cm, interpolation="nearest", cmap=plt.cm.Greens)
ax.set(
xlim=[-0.5, 1.5],
ylim=[-0.5, 1.5],
xticks=[0, 1],
yticks=[0, 1],
xticklabels=classes,
yticklabels=classes,
xlabel="predicted label",
ylabel="true label"
)
for i = 1:2
for j = 1:2
ax.text(j-1, i-1, @sprintf("%d", cm[i, j]), ha="center", va="center")
end
end
tight_layout()
end
k = 3
knn = KNeighborsClassifier(n_neighbors=k)
knn.fit(X_train, y_train)
y_pred = knn.predict(X_test)
plot_confusion_matrix(y_test, y_pred, ["benign", "malignant"])
# -
# to find the best $k$, we can scan over all $k$, train the model in the training set, then test the model on the test set (unseen data) to see how it performs. we take the "best" $k$ as the one that yields the lowest test set error, as we expect it to have the lowest generalization error on unseen data.
# +
ks = 1:20
test_accuracy = zeros(length(ks))
train_accuracy = zeros(length(ks))
for (i, k) in enumerate(ks)
knn = KNeighborsClassifier(n_neighbors=k)
knn.fit(X_train, y_train)
test_accuracy[i] = knn.score(X_test, y_test)
train_accuracy[i] = knn.score(X_train, y_train)
end
figure()
plot(ks, train_accuracy, marker="o", label="train")
plot(ks, test_accuracy, marker="o", label="test")
legend()
xlabel(L"$k$")
ylabel("accuracy")
# -
# the optimal $k$ is around 8. So we should train our $k$-NN algo with $k=8$, then deploy this model, since it should result in the best accuracy on unseen data.
# +
knn = KNeighborsClassifier(n_neighbors=8)
knn.fit(X, y) # *this* is the model we deploy
# look at new decision boundary
for i = 1:length(radius)
for j = 1:length(smoothness)
x = [radius[i] smoothness[j]]
knn_prediction[j, i] = knn.predict(x)[1]
end
end
figure()
contourf(radius, smoothness, knn_prediction, alpha=0.4, [0.0, 0.5, 1.0], colors=["g", "r"])
for df_c in groupby(df, :outcome)
outcome = df_c[1, :outcome]
scatter(df_c[:, :mean_radius], df_c[:, :mean_smoothness], label="$outcome",
marker="o", facecolor="None", edgecolor=color_scheme[outcome])
end
xlabel("mean radius")
ylabel("mean smoothness")
| CHE599-IntroDataScience/lectures/knn/kNN_algo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pyodbc
import pandas as pd
import config as cfg
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn.metrics import accuracy_score
# %matplotlib inline
import seaborn as sns
import matplotlib.pyplot as plt
cnxn = pyodbc.connect( 'DRIVER={ODBC Driver 13 for SQL Server};SERVER=' + cfg.mssql['server'] + ';DATABASE='
+ cfg.mssql['database'] + ';UID=' + cfg.mssql['username'] + ';PWD=' + cfg.mssql['password'] )
query = "SELECT * FROM BankSampleView;"
data = pd.read_sql(query, cnxn, index_col='BankID')
data.head()
data.columns
data.shape
# ## Data Prep
data2 = pd.DataFrame(index=data.index.copy())
data2 = data[['Name', 'City', 'Deposit', 'ClosestPSDistance',
'MeanPSDistance', 'PSCount', 'Take', 'PDistance', 'Officers1000',
'FFLCount', 'AvgRating', 'Target', 'Population', 'CrimeRate1000' ]].copy()
data2['CrimeRate1000'].mean()
data2['Population'].mean()
data2.isnull().sum()
values = {'CrimeRate1000': data2['CrimeRate1000'].mean(), 'Population': data2['Population'].mean(), 'AvgRating' : data2['AvgRating'].mean()}
data2.fillna(value=values, inplace=True)
#data2.dropna(axis=0, inplace=True)
data2.shape
# Map target class 1 - 5 (70% success rate and up) to 1 and the rest to 0
data2['target_bin'] = data2.Target.map({ 0:0, 1:1, 2:1, 3:1, 4:1, 5:1 })
data2.target_bin[data2.target_bin != 0].sum() #total target
# ## Correlation
plt.figure(figsize = (15,10))
sns.heatmap(data2.corr(), annot=True)
#Distance, P(Caught by Distance), Target correlation
g = sns.pairplot(data2[['ClosestPSDistance', 'FFLCount', 'Officers1000', 'Take', 'Population', 'AvgRating', 'CrimeRate1000', 'target_bin']], diag_kind="hist", kind="reg")
for ax in g.axes.flat:
plt.setp(ax.get_xticklabels(), rotation=45)
# ## RandomForest with GridSearch for best Features, Max Depth and Min Leaf
from sklearn.model_selection import GridSearchCV
from sklearn.ensemble import RandomForestClassifier
feature_cols = ['ClosestPSDistance',
'MeanPSDistance', 'PSCount', 'Take', 'PDistance', 'Officers1000',
'FFLCount', 'AvgRating', 'Population', 'CrimeRate1000']
X = data2[feature_cols]
y = data2['target_bin']
# +
#Use RandomForest
rf = RandomForestClassifier(random_state=1)
max_depth_range = range(1, 21)
param_grid = dict(max_depth=list(max_depth_range))
grid = GridSearchCV(rf, param_grid, cv=10, scoring='roc_auc')
grid.fit(X, y)
grid_mean_scores = grid.cv_results_['mean_test_score']
# plot the results
plt.plot(max_depth_range, grid_mean_scores)
plt.xlabel('Value of max_depth')
plt.ylabel('Cross-Validated AUC')
print (grid.best_score_)
print (grid.best_params_)
print (grid.best_estimator_)
# -
pd.DataFrame({'feature':feature_cols, 'importance':grid.best_estimator_.feature_importances_}).sort_values(by=['importance'])
# ### Remove Features
# +
# Remove features: MeanPSDistance, PSCount
# Use the rest
feature_cols = ['ClosestPSDistance', 'Take', 'PDistance', 'Officers1000',
'FFLCount', 'AvgRating', 'Population', 'CrimeRate1000']
X = data2[feature_cols]
y = data2['target_bin']
rf = RandomForestClassifier(random_state=1)
max_depth_range = range(1, 21)
param_grid = dict(max_depth=list(max_depth_range))
grid = GridSearchCV(rf, param_grid, cv=10, scoring='roc_auc')
grid.fit(X, y)
grid_mean_scores = grid.cv_results_['mean_test_score']
# plot the results
plt.plot(max_depth_range, grid_mean_scores)
plt.xlabel('Value of max_depth')
plt.ylabel('Cross-Validated AUC')
print (grid.best_score_)
# -
# ### Best Depth and Min Leaf
# Use Grid Search Cross Validation for best depth AND min leaf
max_depth_range = range(1, 21)
leaf_range = range(1, 11)
param_grid = dict(max_depth=list(max_depth_range), min_samples_leaf=list(leaf_range))
grid = GridSearchCV(rf, param_grid, cv=10, scoring='roc_auc')
grid.fit(X, y)
print (grid.best_score_)
print (grid.best_params_)
print (grid.best_estimator_)
# ## Use the max_depth and min_samples_leaf for the model
rf2 = RandomForestClassifier(random_state=1, max_depth=8, min_samples_leaf=2)
rf2.fit(X,y)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=1)
#Check the model with prediction
y_pred = rf2.predict(X_test)
metrics.accuracy_score(y_test, y_pred)
from sklearn.tree import export_graphviz
idx = 0
for tree in rf2.estimators_:
export_graphviz(tree, out_file='tree_banks_' + str(idx) + '.dot', feature_names=feature_cols)
idx = idx + 1
# ! dot -Tpng tree_banks_1.dot -o tree_banks_1.png
# +
from IPython.display import Image
from IPython.display import display
display(Image('tree_banks_1.png'))
# -
# ## Save Model to Pickle
try:
import cPickle as pickle
except ImportError:
import pickle
## Save the RandomForestClassifier model
out_s = open('targetbanks_randomforestclassifier.pkl', 'wb')
pickle.dump(rf2, out_s)
out_s.close()
| Modeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
from tensorflow_tts.inference import AutoProcessor
from IPython.display import Audio
processor = AutoProcessor.from_pretrained(pretrained_path="./kss_mapper.json")
tacotron2_model = 'tacotron2_korean_saved_model'
mb_melgan_model = 'mb_melgan_korean_saved_model'
mb_melgan = tf.saved_model.load(mb_melgan_model)
tacotron2 = tf.saved_model.load(tacotron2_model)
input_text = "신은 우리의 수학 문제에는 관심이 없다. 신은 다만 경험적으로 통합할 뿐이다."
input_ids = processor.text_to_sequence(input_text)
_, mel_outputs, stop_token_prediction, alignment_history = tacotron2.inference(
tf.expand_dims(tf.convert_to_tensor(input_ids, dtype=tf.int32), 0),
tf.convert_to_tensor([len(input_ids)], tf.int32),
tf.convert_to_tensor([0], dtype=tf.int32)
)
audios = mb_melgan.inference(mel_outputs)
Audio(data=audios[0, :, 0], rate=22050)
| notebooks/Tacotron2_Tensorflow_Inference_code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="SRcy_kIPDo6U"
# # CTR (Item ranking) models in Tensorflow on ML-1m
# + [markdown] id="0brc2-VLDo6W"
# ## **Step 1 - Setup the environment**
# + [markdown] id="rS5pc1KVDo6Y"
# ### **1.1 Install libraries**
# + id="iKc5Z6OOGqVD"
# !pip install tensorflow==2.5.0
# + id="zh3IrkWHGywI"
# !pip install -q -U git+https://github.com/RecoHut-Projects/recohut.git -b v0.0.5
# + [markdown] id="th2G0p2IDo6f"
# ### **1.2 Download datasets**
# + colab={"base_uri": "https://localhost:8080/"} id="lxCe3QHbGrkm" executionInfo={"status": "ok", "timestamp": 1640008563294, "user_tz": -330, "elapsed": 363332, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="a5c1b915-5c10-4a3d-bbce-8687be6dca08"
# !pip install -q -U kaggle
# !pip install --upgrade --force-reinstall --no-deps kaggle
# !mkdir ~/.kaggle
# !cp /content/drive/MyDrive/kaggle.json ~/.kaggle/
# !chmod 600 ~/.kaggle/kaggle.json
# !kaggle datasets download -d mrkmakr/criteo-dataset
# !unzip criteo-dataset.zip
# + [markdown] id="fIs1ygR3Do6i"
# ### **1.3 Import libraries**
# + id="VjtUOFBEG-Tp" executionInfo={"status": "ok", "timestamp": 1640009732310, "user_tz": -330, "elapsed": 4181, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
import os
import pandas as pd
import numpy as np
from sklearn.preprocessing import LabelEncoder, KBinsDiscretizer
from sklearn.model_selection import train_test_split
import tensorflow as tf
from tensorflow.keras.losses import binary_crossentropy
from tensorflow.keras.callbacks import EarlyStopping
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.metrics import AUC
# + id="wj-hW3UXHZLv" executionInfo={"status": "ok", "timestamp": 1640014506365, "user_tz": -330, "elapsed": 607, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
from recohut.transforms.datasets.criteo import create_criteo_dataset
from recohut.models.tf.fm import FM, FFM, NFM, AFM, DeepFM, xDeepFM
from recohut.models.tf.widedeep import WideDeep
from recohut.models.tf.deepcross import DeepCross
from recohut.models.tf.pnn import PNN
from recohut.models.tf.dcn import DCN
# + [markdown] id="6nhAeMCWDo6n"
# ### **1.4 Set params**
# + id="OJBMn-x6HLah" executionInfo={"status": "ok", "timestamp": 1640010334317, "user_tz": -330, "elapsed": 720, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '2'
os.environ['CUDA_VISIBLE_DEVICES'] = '0'
# + id="dWWoB0bnHFLD" executionInfo={"status": "ok", "timestamp": 1640014905006, "user_tz": -330, "elapsed": 704, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
class Args:
def __init__(self, model='fm'):
self.file = 'dac/train.txt'
self.read_part = True
self.sample_num = 10000
self.test_size = 0.2
self.k = self.embed_dim = 8
self.learning_rate = 0.001
self.batch_size = 4096
self.dnn_dropout = 0.5
self.dropout = 0.5
self.epochs = 2
self.att_vector = 8
self.mode = 'att' # 'max', 'avg'
self.activation = 'relu'
self.embed_reg = 1e-5
self.hidden_units = [256, 128, 64]
self.cin_size = [128, 128]
if model=='ffm':
self.k = 10
self.batch_size = 1024
# + [markdown] id="pn39JAHGDo6t"
# ## **Step 2 - Training & Evaluation**
# + [markdown] id="Y1VgAkXTHitA"
# ### **2.1 FM**
# + id="zVQBZwHKHoD9" executionInfo={"status": "ok", "timestamp": 1640010337793, "user_tz": -330, "elapsed": 5, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
args = Args(model='fm')
# + colab={"base_uri": "https://localhost:8080/"} id="CpnvJ3_UHkAi" executionInfo={"status": "ok", "timestamp": 1640010353121, "user_tz": -330, "elapsed": 13440, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="bd530a40-0984-44bc-b18c-7a2b189105e1"
# ========================== Create dataset =======================
feature_columns, train, test = create_criteo_dataset(file=args.file,
read_part=args.read_part,
sample_num=args.sample_num,
test_size=args.test_size)
train_X, train_y = train
test_X, test_y = test
# ============================Build Model==========================
mirrored_strategy = tf.distribute.MirroredStrategy()
with mirrored_strategy.scope():
model = FM(feature_columns=feature_columns, k=args.k)
model.summary()
# ============================Compile============================
model.compile(loss=binary_crossentropy, optimizer=Adam(learning_rate=args.learning_rate),
metrics=[AUC()])
# ============================model checkpoint======================
# check_path = '../save/fm_weights.epoch_{epoch:04d}.val_loss_{val_loss:.4f}.ckpt'
# checkpoint = tf.keras.callbacks.ModelCheckpoint(check_path, save_weights_only=True,
# verbose=1, period=5)
# ==============================Fit==============================
model.fit(
train_X,
train_y,
epochs=args.epochs,
callbacks=[EarlyStopping(monitor='val_loss', patience=2, restore_best_weights=True)], # checkpoint
batch_size=args.batch_size,
validation_split=0.1
)
# ===========================Test==============================
print('test AUC: %f' % model.evaluate(test_X, test_y, batch_size=args.batch_size)[1])
# + [markdown] id="XoJhQzq6O9Gl"
# ### **2.2 FFM**
# + id="0093ztfHa3Rh" executionInfo={"status": "ok", "timestamp": 1640013578579, "user_tz": -330, "elapsed": 11, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
args = Args(model='ffm')
# + colab={"base_uri": "https://localhost:8080/"} id="b8Ohm6ZzbT1C" executionInfo={"status": "ok", "timestamp": 1640013791161, "user_tz": -330, "elapsed": 156848, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="39f642f5-c3ff-4a85-f2a6-cee9a487e5fc"
# ========================== Create dataset =======================
feature_columns, train, test = create_criteo_dataset(file=args.file,
read_part=args.read_part,
sample_num=args.sample_num,
test_size=args.test_size)
train_X, train_y = train
test_X, test_y = test
# ============================Build Model==========================
model = FFM(feature_columns=feature_columns, k=args.k)
model.summary()
# ============================model checkpoint======================
# check_path = '../save/fm_weights.epoch_{epoch:04d}.val_loss_{val_loss:.4f}.ckpt'
# checkpoint = tf.keras.callbacks.ModelCheckpoint(check_path, save_weights_only=True,
# verbose=1, period=5)
# ============================Compile============================
model.compile(loss=binary_crossentropy, optimizer=Adam(learning_rate=args.learning_rate),
metrics=[AUC()])
# ==============================Fit==============================
model.fit(
train_X,
train_y,
epochs=args.epochs,
callbacks=[EarlyStopping(monitor='val_loss', patience=2, restore_best_weights=True)], # checkpoint
batch_size=args.batch_size,
validation_split=0.1
)
# ===========================Test==============================
print('test AUC: %f' % model.evaluate(test_X, test_y, batch_size=args.batch_size)[1])
# + [markdown] id="ovN5AJw6bhgk"
# ### **2.3 Wide & Deep**
# + id="JW_Ws-vlbrXt" executionInfo={"status": "ok", "timestamp": 1640013799473, "user_tz": -330, "elapsed": 610, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
args = Args(model='widedeep')
# + colab={"base_uri": "https://localhost:8080/"} id="xy-c1JRIcJmY" executionInfo={"status": "ok", "timestamp": 1640013884019, "user_tz": -330, "elapsed": 10308, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="6bcfddbd-ee86-42d2-8a22-136b1f85d3ad"
# ========================== Create dataset =======================
feature_columns, train, test = create_criteo_dataset(file=args.file,
embed_dim=args.embed_dim,
read_part=args.read_part,
sample_num=args.sample_num,
test_size=args.test_size)
train_X, train_y = train
test_X, test_y = test
# ============================Build Model==========================
mirrored_strategy = tf.distribute.MirroredStrategy()
with mirrored_strategy.scope():
model = WideDeep(feature_columns, hidden_units=args.hidden_units, dnn_dropout=args.dnn_dropout)
model.summary()
# ============================Compile============================
model.compile(loss=binary_crossentropy, optimizer=Adam(learning_rate=args.learning_rate),
metrics=[AUC()])
# ============================model checkpoint======================
# check_path = '../save/wide_deep_weights.epoch_{epoch:04d}.val_loss_{val_loss:.4f}.ckpt'
# checkpoint = tf.keras.callbacks.ModelCheckpoint(check_path, save_weights_only=True,
# verbose=1, period=5)
# ==============================Fit==============================
model.fit(
train_X,
train_y,
epochs=args.epochs,
callbacks=[EarlyStopping(monitor='val_loss', patience=2, restore_best_weights=True)], # checkpoint
batch_size=args.batch_size,
validation_split=0.1
)
# ===========================Test==============================
print('test AUC: %f' % model.evaluate(test_X, test_y, batch_size=args.batch_size)[1])
# + [markdown] id="OgBpmMgJcb80"
# ### **2.4 Deep Crossing**
# + id="z7cNC2C2ckS1" executionInfo={"status": "ok", "timestamp": 1640013945511, "user_tz": -330, "elapsed": 607, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
args = Args(model='deepcross')
# + colab={"base_uri": "https://localhost:8080/"} id="z7Zk6wNhcs-A" executionInfo={"status": "ok", "timestamp": 1640014056055, "user_tz": -330, "elapsed": 11232, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="ed5f5085-cae7-4e84-bdaa-46c9e65897c2"
# ========================== Create dataset =======================
feature_columns, train, test = create_criteo_dataset(file=args.file,
embed_dim=args.embed_dim,
read_part=args.read_part,
sample_num=args.sample_num,
test_size=args.test_size)
train_X, train_y = train
test_X, test_y = test
# ============================Build Model==========================
mirrored_strategy = tf.distribute.MirroredStrategy()
with mirrored_strategy.scope():
model = DeepCross(feature_columns, args.hidden_units)
model.summary()
# =========================Compile============================
model.compile(loss=binary_crossentropy, optimizer=Adam(learning_rate=args.learning_rate),
metrics=[AUC()])
# ============================model checkpoint======================
# check_path = 'save/deep_crossing_weights.epoch_{epoch:04d}.val_loss_{val_loss:.4f}.ckpt'
# checkpoint = tf.keras.callbacks.ModelCheckpoint(check_path, save_weights_only=True,
# verbose=1, period=5)
# ===========================Fit==============================
model.fit(
train_X,
train_y,
epochs=args.epochs,
callbacks=[EarlyStopping(monitor='val_loss', patience=2, restore_best_weights=True)], # checkpoint
batch_size=args.batch_size,
validation_split=0.1
)
# ===========================Test==============================
print('test AUC: %f' % model.evaluate(test_X, test_y, batch_size=args.batch_size)[1])
# + [markdown] id="YcPlkxq0dFo3"
# ### **2.5 PNN**
# + id="kNNxWtrWdP2b" executionInfo={"status": "ok", "timestamp": 1640014096024, "user_tz": -330, "elapsed": 793, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
args = Args(model='pnn')
# + colab={"base_uri": "https://localhost:8080/"} id="_pSQuGUsdSBo" executionInfo={"status": "ok", "timestamp": 1640014283370, "user_tz": -330, "elapsed": 53689, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="2dfad7b5-4ff7-42ee-dea4-7521e132f4b7"
# ========================== Create dataset =======================
feature_columns, train, test = create_criteo_dataset(file=args.file,
embed_dim=args.embed_dim,
read_part=args.read_part,
sample_num=args.sample_num,
test_size=args.test_size)
train_X, train_y = train
test_X, test_y = test
# ============================Build Model==========================
mirrored_strategy = tf.distribute.MirroredStrategy()
with mirrored_strategy.scope():
model = PNN(feature_columns, args.hidden_units, args.dnn_dropout)
model.summary()
# =========================Compile============================
model.compile(loss=binary_crossentropy, optimizer=Adam(learning_rate=args.learning_rate),
metrics=[AUC()])
# ============================model checkpoint======================
# check_path = 'save/pnn_weights.epoch_{epoch:04d}.val_loss_{val_loss:.4f}.ckpt'
# checkpoint = tf.keras.callbacks.ModelCheckpoint(check_path, save_weights_only=True,
# verbose=1, period=5)
# ===========================Fit==============================
model.fit(
train_X,
train_y,
epochs=args.epochs,
callbacks=[EarlyStopping(monitor='val_loss', patience=2, restore_best_weights=True)], # checkpoint
batch_size=args.batch_size,
validation_split=0.1
)
# ===========================Test==============================
print('test AUC: %f' % model.evaluate(test_X, test_y, batch_size=args.batch_size)[1])
# + [markdown] id="5QGMmWeZdy20"
# ### **2.6 DCN**
# + id="sWHzDu7Qd7Eo" executionInfo={"status": "ok", "timestamp": 1640014285372, "user_tz": -330, "elapsed": 8, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
args = Args(model='dcn')
# + colab={"base_uri": "https://localhost:8080/"} id="udqJU5Ahd-jq" executionInfo={"status": "ok", "timestamp": 1640014358793, "user_tz": -330, "elapsed": 10320, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="ef38e55b-1a5e-400f-8935-425586e3cecb"
# ========================== Create dataset =======================
feature_columns, train, test = create_criteo_dataset(file=args.file,
embed_dim=args.embed_dim,
read_part=args.read_part,
sample_num=args.sample_num,
test_size=args.test_size)
train_X, train_y = train
test_X, test_y = test
# ============================Build Model==========================
mirrored_strategy = tf.distribute.MirroredStrategy()
with mirrored_strategy.scope():
model = DCN(feature_columns, args.hidden_units, dnn_dropout=args.dnn_dropout)
model.summary()
# =========================Compile============================
model.compile(loss=binary_crossentropy, optimizer=Adam(learning_rate=args.learning_rate),
metrics=[AUC()])
# ============================model checkpoint======================
# check_path = 'save/dcn_weights.epoch_{epoch:04d}.val_loss_{val_loss:.4f}.ckpt'
# checkpoint = tf.keras.callbacks.ModelCheckpoint(check_path, save_weights_only=True,
# verbose=1, period=5)
# ===========================Fit==============================
model.fit(
train_X,
train_y,
epochs=args.epochs,
callbacks=[EarlyStopping(monitor='val_loss', patience=2, restore_best_weights=True)], # checkpoint
batch_size=args.batch_size,
validation_split=0.1
)
# ===========================Test==============================
print('test AUC: %f' % model.evaluate(test_X, test_y, batch_size=args.batch_size)[1])
# + [markdown] id="0sIomQLKeP0E"
# ### **2.7 NFM**
# + id="r8mXoX2BejRx" executionInfo={"status": "ok", "timestamp": 1640014513790, "user_tz": -330, "elapsed": 538, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
args = Args(model='nfm')
# + colab={"base_uri": "https://localhost:8080/"} id="DFAQtguRemF8" executionInfo={"status": "ok", "timestamp": 1640014528582, "user_tz": -330, "elapsed": 14045, "user": {"displayName": "<NAME>wal", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="3cc29459-0a17-47fa-b188-bb2fb0b490ec"
# ========================== Create dataset =======================
feature_columns, train, test = create_criteo_dataset(file=args.file,
embed_dim=args.embed_dim,
read_part=args.read_part,
sample_num=args.sample_num,
test_size=args.test_size)
train_X, train_y = train
test_X, test_y = test
# ============================Build Model==========================
mirrored_strategy = tf.distribute.MirroredStrategy()
with mirrored_strategy.scope():
model = NFM(feature_columns, args.hidden_units, dnn_dropout=args.dnn_dropout)
model.summary()
# =========================Compile============================
model.compile(loss=binary_crossentropy, optimizer=Adam(learning_rate=args.learning_rate),
metrics=[AUC()])
# ============================model checkpoint======================
# check_path = 'save/nfm_weights.epoch_{epoch:04d}.val_loss_{val_loss:.4f}.ckpt'
# checkpoint = tf.keras.callbacks.ModelCheckpoint(check_path, save_weights_only=True,
# verbose=1, period=5)
# ===========================Fit==============================
model.fit(
train_X,
train_y,
epochs=args.epochs,
callbacks=[EarlyStopping(monitor='val_loss', patience=2, restore_best_weights=True)], # checkpoint
batch_size=args.batch_size,
validation_split=0.1
)
# ===========================Test==============================
print('test AUC: %f' % model.evaluate(test_X, test_y, batch_size=args.batch_size)[1])
# + [markdown] id="EfR2ObfXeZWX"
# ### **2.8 AFM**
# + id="BIEsGwIxe6LA" executionInfo={"status": "ok", "timestamp": 1640014721216, "user_tz": -330, "elapsed": 448, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
args = Args(model='afm')
# + colab={"base_uri": "https://localhost:8080/"} id="oDrJt6mce78a" executionInfo={"status": "ok", "timestamp": 1640014736598, "user_tz": -330, "elapsed": 14600, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="fb7478f2-5526-4068-fc14-58a8c91d704d"
# ========================== Create dataset =======================
feature_columns, train, test = create_criteo_dataset(file=args.file,
embed_dim=args.embed_dim,
read_part=args.read_part,
sample_num=args.sample_num,
test_size=args.test_size)
train_X, train_y = train
test_X, test_y = test
# ============================Build Model==========================
mirrored_strategy = tf.distribute.MirroredStrategy()
with mirrored_strategy.scope():
model = AFM(feature_columns, args.mode, args.att_vector, args.activation, args.dropout, args.embed_reg)
model.summary()
# =========================Compile============================
model.compile(loss=binary_crossentropy, optimizer=Adam(learning_rate=args.learning_rate),
metrics=[AUC()])
# ============================model checkpoint======================
# check_path = 'save/afm_weights.epoch_{epoch:04d}.val_loss_{val_loss:.4f}.ckpt'
# checkpoint = tf.keras.callbacks.ModelCheckpoint(check_path, save_weights_only=True,
# verbose=1, period=5)
# ===========================Fit==============================
model.fit(
train_X,
train_y,
epochs=args.epochs,
callbacks=[EarlyStopping(monitor='val_loss', patience=2, restore_best_weights=True)], # checkpoint,
batch_size=args.batch_size,
validation_split=0.1
)
# ===========================Test==============================
print('test AUC: %f' % model.evaluate(test_X, test_y, batch_size=args.batch_size)[1])
# + [markdown] id="8fu7UVaZecN2"
# ### **2.9 DeepFM**
# + id="nAKsDMERf1jX"
args = Args(model='deepfm')
# + colab={"base_uri": "https://localhost:8080/"} id="so06_YSrf4I_" executionInfo={"status": "ok", "timestamp": 1640014858211, "user_tz": -330, "elapsed": 12392, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="0bcf7f70-1ae8-4971-b5a5-0d838423398a"
# ========================== Create dataset =======================
feature_columns, train, test = create_criteo_dataset(file=args.file,
embed_dim=args.embed_dim,
read_part=args.read_part,
sample_num=args.sample_num,
test_size=args.test_size)
train_X, train_y = train
test_X, test_y = test
# ============================Build Model==========================
mirrored_strategy = tf.distribute.MirroredStrategy()
with mirrored_strategy.scope():
model = DeepFM(feature_columns, hidden_units=args.hidden_units, dnn_dropout=args.dnn_dropout)
model.summary()
# ============================Compile============================
model.compile(loss=binary_crossentropy, optimizer=Adam(learning_rate=args.learning_rate),
metrics=[AUC()])
# ============================model checkpoint======================
# check_path = '../save/deepfm_weights.epoch_{epoch:04d}.val_loss_{val_loss:.4f}.ckpt'
# checkpoint = tf.keras.callbacks.ModelCheckpoint(check_path, save_weights_only=True,
# verbose=1, period=5)
# ==============================Fit==============================
model.fit(
train_X,
train_y,
epochs=args.epochs,
callbacks=[EarlyStopping(monitor='val_loss', patience=2, restore_best_weights=True)], # checkpoint,
batch_size=args.batch_size,
validation_split=0.1
)
# ===========================Test==============================
print('test AUC: %f' % model.evaluate(test_X, test_y, batch_size=args.batch_size)[1])
# + [markdown] id="5FW58X-UecQd"
# ### **2.10 xDeepFM**
# + id="ldF-rahyefh1" executionInfo={"status": "ok", "timestamp": 1640014915603, "user_tz": -330, "elapsed": 529, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
args = Args(model='xdeepfm')
# + colab={"base_uri": "https://localhost:8080/"} id="ZHFgD3rlgaJ6" executionInfo={"status": "ok", "timestamp": 1640015004067, "user_tz": -330, "elapsed": 44685, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="b9269666-743e-4c34-dfd5-9f75d5f85cb6"
# ========================== Create dataset =======================
feature_columns, train, test = create_criteo_dataset(file=args.file,
embed_dim=args.embed_dim,
read_part=args.read_part,
sample_num=args.sample_num,
test_size=args.test_size)
train_X, train_y = train
test_X, test_y = test
# ============================Build Model==========================
mirrored_strategy = tf.distribute.MirroredStrategy()
with mirrored_strategy.scope():
model = xDeepFM(feature_columns, args.hidden_units, args.cin_size)
model.summary()
# =========================Compile============================
model.compile(loss=binary_crossentropy, optimizer=Adam(learning_rate=args.learning_rate),
metrics=[AUC()])
# ============================model checkpoint======================
# check_path = 'save/xdeepfm_weights.epoch_{epoch:04d}.val_loss_{val_loss:.4f}.ckpt'
# checkpoint = tf.keras.callbacks.ModelCheckpoint(check_path, save_weights_only=True,
# verbose=1, period=5)
# ===========================Fit==============================
model.fit(
train_X,
train_y,
epochs=args.epochs,
callbacks=[EarlyStopping(monitor='val_loss', patience=2, restore_best_weights=True)], # checkpoint
batch_size=args.batch_size,
validation_split=0.1
)
# ===========================Test==============================
print('test AUC: %f' % model.evaluate(test_X, test_y, batch_size=args.batch_size)[1])
# + [markdown] id="NCFKoIs5Do6z"
# ## **Closure**
# + [markdown] id="sSKqb6t5Do60"
# For more details, you can refer to https://github.com/RecoHut-Stanzas/S021355.
# + [markdown] id="0R4uWucpDo61"
# <a href="https://github.com/RecoHut-Stanzas/S021355/blob/main/reports/S021355.ipynb" alt="S021355_Report"> <img src="https://img.shields.io/static/v1?label=report&message=active&color=green" /></a> <a href="https://github.com/RecoHut-Stanzas/S021355" alt="S021355"> <img src="https://img.shields.io/static/v1?label=code&message=github&color=blue" /></a>
# + colab={"base_uri": "https://localhost:8080/"} id="zuVA3NzxglAX" executionInfo={"status": "ok", "timestamp": 1640015143767, "user_tz": -330, "elapsed": 3846, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="427dd15e-62bb-4c39-f000-b94c68110ce2"
# !pip install -q watermark
# %reload_ext watermark
# %watermark -a "Sparsh A." -m -iv -u -t -d
# + [markdown] id="JOsqyvAkDo62"
# ---
# + [markdown] id="LLugelemDo63"
# **END**
| _docs/nbs/T844978-CTR-Item-ranking-models-in-Tensorflow-on-ML-1m.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/krishnaaxo/Drug_Discovery_AI/blob/main/EDA.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="c2Cbi02oLC3b" outputId="dfa29443-3a42-4e9e-8836-bf8d7c4e8307"
pip install rdkit-pypi
# + [markdown] id="QmxXXFa4wTNG"
# ## **Load bioactivity data**
# + colab={"base_uri": "https://localhost:8080/"} id="4jXqIPHbaq-b" outputId="738e21ac-9fa6-435d-d5a9-3380af51e30a"
# ! wget https://raw.githubusercontent.com/krishnaaxo/Drug_Discovery_AI/main/acetylcholinesterase_03_bioactivity_data_curated.csv
# + id="Fpu5C7HlwV9s"
import pandas as pd
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="GCcE8J5XwjtB" outputId="50a85f49-0238-41de-e92c-fa9bea79ab7f"
df = pd.read_csv('acetylcholinesterase_03_bioactivity_data_curated.csv')
df
# + id="7AMm19NW0bJR"
df_no_smiles = df.drop(columns='canonical_smiles')
# + id="aymiQsfdr5sY"
smiles = []
for i in df.canonical_smiles.tolist():
cpd = str(i).split('.')
cpd_longest = max(cpd, key = len)
smiles.append(cpd_longest)
smiles = pd.Series(smiles, name = 'canonical_smiles')
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="l7a8qW_U85ZK" outputId="1b7728df-aa42-491d-fd8f-ebbd9e60a28e"
df_clean_smiles = pd.concat([df_no_smiles,smiles], axis=1)
df_clean_smiles
# + [markdown] id="YzN_S4Quro5S"
# ## **Calculate Lipinski descriptors**
# <NAME>, a scientist at Pfizer, came up with a set of rule-of-thumb for evaluating the **druglikeness** of compounds. Such druglikeness is based on the Absorption, Distribution, Metabolism and Excretion (ADME) that is also known as the pharmacokinetic profile. Lipinski analyzed all orally active FDA-approved drugs in the formulation of what is to be known as the **Rule-of-Five** or **Lipinski's Rule**.
#
# The Lipinski's Rule stated the following:
# * Molecular weight < 500 Dalton
# * Octanol-water partition coefficient (LogP) < 5
# * Hydrogen bond donors < 5
# * Hydrogen bond acceptors < 10
# + [markdown] id="9qn_eQcnxY7C"
# ### **Import libraries**
# + id="CgBjIdT-rnRU"
import numpy as np
from rdkit import Chem
from rdkit.Chem import Descriptors, Lipinski
# + [markdown] id="JsgTV-ByxdMa"
# ### **Calculate descriptors**
# + id="bCXEY7a9ugO_"
# Inspired by: https://codeocean.com/explore/capsules?query=tag:data-curation
def lipinski(smiles, verbose=False):
moldata= []
for elem in smiles:
mol=Chem.MolFromSmiles(elem)
moldata.append(mol)
baseData= np.arange(1,1)
i=0
for mol in moldata:
desc_MolWt = Descriptors.MolWt(mol)
desc_MolLogP = Descriptors.MolLogP(mol)
desc_NumHDonors = Lipinski.NumHDonors(mol)
desc_NumHAcceptors = Lipinski.NumHAcceptors(mol)
row = np.array([desc_MolWt,
desc_MolLogP,
desc_NumHDonors,
desc_NumHAcceptors])
if(i==0):
baseData=row
else:
baseData=np.vstack([baseData, row])
i=i+1
columnNames=["MW","LogP","NumHDonors","NumHAcceptors"]
descriptors = pd.DataFrame(data=baseData,columns=columnNames)
return descriptors
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="ThFIFw8IukMY" outputId="2caabf72-812e-4af7-b678-d73689ec01ae"
df_lipinski = lipinski(df_clean_smiles.canonical_smiles)
df_lipinski
# + [markdown] id="gUMlPfFrxicj"
# ### **Combine DataFrames**
#
# Let's take a look at the 2 DataFrames that will be combined.
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="DaezyM5vwp9n" outputId="9424be5c-a670-44d2-c785-10da1f438c22"
df_lipinski
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="9-ChzM8_wuq_" outputId="77b425ad-3c88-4482-eb93-3fb1f7de4d5f"
df
# + [markdown] id="eET6iZ1Aw3oe"
# Now, let's combine the 2 DataFrame
# + id="L9nUZC0Ww3gp"
df_combined = pd.concat([df,df_lipinski], axis=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="FRBfBP3QxFJp" outputId="1ee758ad-b8ff-474b-b51b-cd8dea9a2bb2"
df_combined
# + [markdown] id="e0MLOedB6j96"
# ### **Convert IC50 to pIC50**
# To allow **IC50** data to be more uniformly distributed, we will convert **IC50** to the negative logarithmic scale which is essentially **-log10(IC50)**.
#
# This custom function pIC50() will accept a DataFrame as input and will:
# * Take the IC50 values from the ``standard_value`` column and converts it from nM to M by multiplying the value by 10$^{-9}$
# * Take the molar value and apply -log10
# * Delete the ``standard_value`` column and create a new ``pIC50`` column
# + id="UXMuFQoQ4pZF"
# https://github.com/chaninlab/estrogen-receptor-alpha-qsar/blob/master/02_ER_alpha_RO5.ipynb
import numpy as np
def pIC50(input):
pIC50 = []
for i in input['standard_value_norm']:
molar = i*(10**-9) # Converts nM to M
pIC50.append(-np.log10(molar))
input['pIC50'] = pIC50
x = input.drop('standard_value_norm', 1)
return x
# + [markdown] id="WU5Fh1h2OaJJ"
# Point to note: Values greater than 100,000,000 will be fixed at 100,000,000 otherwise the negative logarithmic value will become negative.
# + colab={"base_uri": "https://localhost:8080/"} id="QuUTFUpcR1wU" outputId="8f11022a-ecce-4938-d6a8-f96fbd4f27ce"
df_combined.standard_value.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="QyiJ0to5N6Z_" outputId="05c1b1ab-2c49-48e2-bbcd-a5e14c15799b"
-np.log10( (10**-9)* 100000000 )
# + colab={"base_uri": "https://localhost:8080/"} id="9S1aJkOYOP6K" outputId="51626bdf-2b68-4c8e-ebd2-124383cde2d4"
-np.log10( (10**-9)* 10000000000 )
# + id="iktHDDwtPDwl"
def norm_value(input):
norm = []
for i in input['standard_value']:
if i > 100000000:
i = 100000000
norm.append(i)
input['standard_value_norm'] = norm
x = input.drop('standard_value', 1)
return x
# + [markdown] id="EkrTs7RfPsrH"
# We will first apply the norm_value() function so that the values in the standard_value column is normalized.
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="EX2Mj2-ZP1Rj" outputId="c6325d4c-b3ee-4c95-de01-197c85aadca8"
df_norm = norm_value(df_combined)
df_norm
# + colab={"base_uri": "https://localhost:8080/"} id="hb1eKrIjRiH9" outputId="66c8fcce-020a-4664-c0ee-09b0c9db89b6"
df_norm.standard_value_norm.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="UDKZzmK57YnS" outputId="1d5ad0f5-e41a-4032-a52a-05ed4c4f0edf"
df_final = pIC50(df_norm)
df_final
# + colab={"base_uri": "https://localhost:8080/"} id="BoqY53udSTYC" outputId="a820fb83-2bc4-4212-f403-b5fef88b61a7"
df_final.pIC50.describe()
# + [markdown] id="TTB-xkOcgwRX"
# Let's write this to CSV file.
# + id="sbZfHUSOgykw"
df_final.to_csv('acetylcholinesterase_04_bioactivity_data_3class_pIC50.csv')
# + [markdown] id="05vHBWvqaQtb"
# ### **Removing the 'intermediate' bioactivity class**
# Here, we will be removing the ``intermediate`` class from our data set.
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="HmrndhDW3c7Z" outputId="78f27d48-31cb-4f2f-c08b-4b3e759a7424"
df_2class = df_final[df_final['class'] != 'intermediate']
df_2class
# + [markdown] id="GJGRNXXigd1o"
# Let's write this to CSV file.
# + id="XIiYdeL5ghCo"
df_2class.to_csv('acetylcholinesterase_05_bioactivity_data_2class_pIC50.csv')
# + [markdown] id="p9vA4-hQQ8sA"
# ---
# + [markdown] id="x0vqbQWfxsZu"
# ## **Exploratory Data Analysis (Chemical Space Analysis) via Lipinski descriptors**
# + [markdown] id="18heJagiyHoF"
# ### **Import library**
# + id="0Egq_rNsxtIj"
import seaborn as sns
sns.set(style='ticks')
import matplotlib.pyplot as plt
# + [markdown] id="NiarmFbOdG3H"
# ### **Frequency plot of the 2 bioactivity classes**
# + colab={"base_uri": "https://localhost:8080/", "height": 367} id="w2Ia0iycdMO2" outputId="a6aa38e4-33e8-4e27-9e7b-7e3430d152d3"
plt.figure(figsize=(5.5, 5.5))
sns.countplot(x='class', data=df_2class, edgecolor='black')
plt.xlabel('Bioactivity class', fontsize=14, fontweight='bold')
plt.ylabel('Frequency', fontsize=14, fontweight='bold')
plt.savefig('plot_bioactivity_class.pdf')
# + [markdown] id="wB68NKVG0j68"
# ### **Scatter plot of MW versus LogP**
#
# It can be seen that the 2 bioactivity classes are spanning similar chemical spaces as evident by the scatter plot of MW vs LogP.
# + colab={"base_uri": "https://localhost:8080/", "height": 367} id="F79BNwjF0nub" outputId="98023d1d-9d8a-4895-b5d9-0f17389b34bf"
plt.figure(figsize=(5.5, 5.5))
sns.scatterplot(x='MW', y='LogP', data=df_2class, hue='class', size='pIC50', edgecolor='black', alpha=0.7)
plt.xlabel('MW', fontsize=14, fontweight='bold')
plt.ylabel('LogP', fontsize=14, fontweight='bold')
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0)
plt.savefig('plot_MW_vs_LogP.pdf')
# + [markdown] id="oLAfyRwHyJfX"
# ### **Box plots**
# + [markdown] id="1n1uIAivyOkY"
# #### **pIC50 value**
# + colab={"base_uri": "https://localhost:8080/", "height": 367} id="EpPviw0hxue6" outputId="0d4183ff-84ed-487f-e029-c070b198f7cc"
plt.figure(figsize=(5.5, 5.5))
sns.boxplot(x = 'class', y = 'pIC50', data = df_2class)
plt.xlabel('Bioactivity class', fontsize=14, fontweight='bold')
plt.ylabel('pIC50 value', fontsize=14, fontweight='bold')
plt.savefig('plot_ic50.pdf')
# + [markdown] id="PsOqKyysCZCv"
# **Statistical analysis | Mann-Whitney U Test**
# + id="LPdV1vDnWBsh"
def mannwhitney(descriptor, verbose=False):
# https://machinelearningmastery.com/nonparametric-statistical-significance-tests-in-python/
from numpy.random import seed
from numpy.random import randn
from scipy.stats import mannwhitneyu
# seed the random number generator
seed(1)
# actives and inactives
selection = [descriptor, 'class']
df = df_2class[selection]
active = df[df['class'] == 'active']
active = active[descriptor]
selection = [descriptor, 'class']
df = df_2class[selection]
inactive = df[df['class'] == 'inactive']
inactive = inactive[descriptor]
# compare samples
stat, p = mannwhitneyu(active, inactive)
#print('Statistics=%.3f, p=%.3f' % (stat, p))
# interpret
alpha = 0.05
if p > alpha:
interpretation = 'Same distribution (fail to reject H0)'
else:
interpretation = 'Different distribution (reject H0)'
results = pd.DataFrame({'Descriptor':descriptor,
'Statistics':stat,
'p':p,
'alpha':alpha,
'Interpretation':interpretation}, index=[0])
filename = 'mannwhitneyu_' + descriptor + '.csv'
results.to_csv(filename)
return results
# + colab={"base_uri": "https://localhost:8080/", "height": 80} id="HZmUgOmdYVm5" outputId="f030ac46-2312-43f6-c622-98cb69126535"
mannwhitney('pIC50')
# + [markdown] id="o2UlCwPmyTBq"
# #### **MW**
# + colab={"base_uri": "https://localhost:8080/", "height": 370} id="ZNlEEsDEx3m6" outputId="1b6bb9b9-257c-4d63-b871-3adfa8b5fa98"
plt.figure(figsize=(5.5, 5.5))
sns.boxplot(x = 'class', y = 'MW', data = df_2class)
plt.xlabel('Bioactivity class', fontsize=14, fontweight='bold')
plt.ylabel('MW', fontsize=14, fontweight='bold')
plt.savefig('plot_MW.pdf')
# + colab={"base_uri": "https://localhost:8080/", "height": 80} id="wRl2FvgHYqaG" outputId="553e9100-849b-4d88-d905-e71b70c31eda"
mannwhitney('MW')
# + [markdown] id="z5hyBhGqyc6J"
# #### **LogP**
# + colab={"base_uri": "https://localhost:8080/", "height": 367} id="liEtkpI4yX9t" outputId="a7359701-1f1c-4f05-bfc6-1e6275567a5a"
plt.figure(figsize=(5.5, 5.5))
sns.boxplot(x = 'class', y = 'LogP', data = df_2class)
plt.xlabel('Bioactivity class', fontsize=14, fontweight='bold')
plt.ylabel('LogP', fontsize=14, fontweight='bold')
plt.savefig('plot_LogP.pdf')
# + [markdown] id="2KgV5v_oFLXh"
# **Statistical analysis | Mann-Whitney U Test**
# + colab={"base_uri": "https://localhost:8080/", "height": 80} id="B61UsGMIFLuE" outputId="29a46218-3917-42ce-9a2e-ddf069052bf7"
mannwhitney('LogP')
# + [markdown] id="4db7LZLRym2k"
# #### **NumHDonors**
# + colab={"base_uri": "https://localhost:8080/", "height": 367} id="iru1JPM1yg5A" outputId="55f53443-ad54-4251-a02c-9d331428ddda"
plt.figure(figsize=(5.5, 5.5))
sns.boxplot(x = 'class', y = 'NumHDonors', data = df_2class)
plt.xlabel('Bioactivity class', fontsize=14, fontweight='bold')
plt.ylabel('NumHDonors', fontsize=14, fontweight='bold')
plt.savefig('plot_NumHDonors.pdf')
# + [markdown] id="vM5vZWanFe3c"
# **Statistical analysis | Mann-Whitney U Test**
# + colab={"base_uri": "https://localhost:8080/", "height": 80} id="mS-rOqq7Fd1E" outputId="86b66ccc-9d6b-4e3b-8c1f-5db7459b9c64"
mannwhitney('NumHDonors')
# + [markdown] id="yOYQ3QiSyu7-"
# #### **NumHAcceptors**
# + colab={"base_uri": "https://localhost:8080/", "height": 367} id="yCw6tgNCyxHf" outputId="ea23d2a8-815a-44fa-e9a9-e5b5685264b2"
plt.figure(figsize=(5.5, 5.5))
sns.boxplot(x = 'class', y = 'NumHAcceptors', data = df_2class)
plt.xlabel('Bioactivity class', fontsize=14, fontweight='bold')
plt.ylabel('NumHAcceptors', fontsize=14, fontweight='bold')
plt.savefig('plot_NumHAcceptors.pdf')
# + colab={"base_uri": "https://localhost:8080/", "height": 80} id="NEQoDZctFtGG" outputId="5d333b1d-e80b-4fc1-c86d-13609c65144d"
mannwhitney('NumHAcceptors')
# + [markdown] id="p4QjdHVjKYum"
# #### **Interpretation of Statistical Results**
# + [markdown] id="mSoOBIqfLbDs"
# ##### **Box Plots**
#
# ###### **pIC50 values**
#
# Taking a look at pIC50 values, the **actives** and **inactives** displayed ***statistically significant difference***, which is to be expected since threshold values (``IC50 < 1,000 nM = Actives while IC50 > 10,000 nM = Inactives``, corresponding to ``pIC50 > 6 = Actives and pIC50 < 5 = Inactives``) were used to define actives and inactives.
#
# ###### **Lipinski's descriptors**
#
# All of the 4 Lipinski's descriptors exhibited ***statistically significant difference*** between the **actives** and **inactives**.
# + [markdown] id="U-rK8l0wWnKK"
# ## **Zip files**
# + colab={"base_uri": "https://localhost:8080/"} id="GW1ZSsfJWqbM" outputId="caa0da00-de46-416d-d646-511ef6766526"
# ! zip -r results.zip . -i *.csv *.pdf
# + id="LMWOG2UIXEg-"
| EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="c-ijxti0SiIn"
# Import necessary libraries
import numpy as np
import pandas as pd
import os , sys ,cv2
import json
import time
import csv
import keras.applications
from keras.applications.mobilenet_v2 import MobileNetV2
from keras.preprocessing import image
from keras.applications.resnet50 import ResNet50
from keras.applications.resnet50 import preprocess_input
from keras.preprocessing import image
from keras.preprocessing.image import ImageDataGenerator, array_to_img, img_to_array, load_img
from keras.models import Model
from urllib.request import urlopen
from PIL import Image
from sklearn.preprocessing import StandardScaler
from sklearn import svm
import shutil
# -
# Create new folder to store the snapshots from train videos
try:
os.mkdir("./output_dir")
except FileExistsError:
pass
# Create new folder to store the snapshots from test videos
try:
os.mkdir("./test_output_dir")
except FileExistsError:
pass
# +
# Get the snapshots of the training videos for 1/3rd frame of the total length
pathOut = "./output_dir/"
listing = os.listdir("./train_video/")
count = 0
counter = 1
for vid in listing:
vid1 = "./train_video/"+vid
cap = cv2.VideoCapture(vid1)
frame = int(cap.get(cv2.CAP_PROP_FRAME_COUNT) / 3) # Divide by 3 to get correct frame
count = 0
counter += 1
success = True
while success:
success,image = cap.read()
print('read a new frame:',success)
if count == frame:
cv2.imwrite(pathOut + str(vid) + ".jpg",image)
print (success)
count+=1
# +
# Get the snapshots of the test videos for 1/3rd frame of the total length
pathOut = "./test_output_dir/"
listing = os.listdir("./test_video/")
count = 0
counter = 1
for vid in listing:
vid1 = "./test_video/"+vid
cap = cv2.VideoCapture(vid1)
frame = int(cap.get(cv2.CAP_PROP_FRAME_COUNT) / 3) # Divide by 3 to get proper frame
count = 0
counter += 1
success = True
while success:
success,image = cap.read()
print('read a new frame:',success)
if count == frame:
cv2.imwrite(pathOut + str(vid) + ".jpg",image)
print (success)
count+=1
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="0Hrd6Qr2Seou" outputId="88cff61c-9a62-444c-dd3e-f53f36931ed4"
# Read the labels for the training images after appending .jpg to the image filename
df_train = pd.read_csv('tarin_tag.txt',header=None, sep=',')
df_train.columns = ['filename','label']
df_train['filename'] = df_train['filename'] + '.jpg'
df_train.head(5)
# + colab={} colab_type="code" id="d_kubYYPUMdr"
filename = df_train.filename.values
label = df_train.label.values
# + colab={} colab_type="code" id="DbaPe8ykeqJv"
# Function to get the image size in the image directory
def file_size_train(file_name):
stats=os.stat("output_dir/"+str(file_name))
return stats.st_size
# + colab={} colab_type="code" id="8Sq1JBOkUYd_"
# Function to call the keras pre-trained model to get feature weights of the train images
def resnet_model_train():
n=0
model = ResNet50(weights='imagenet', include_top=False, pooling='avg')
images_path = "./output_dir"
# Looping over every image present in the files list
for img_path in filename:
if(file_size_train(img_path)!=0):
print(str(img_path))
n+=1
print(n)
# load the image and resize it
img = image.load_img("./output_dir/"+str(img_path), target_size=(224, 224))
# extract features from each image
x_image = image.img_to_array(img)
x_image = np.expand_dims(x_image, axis=0) # increase dimensions of x to make it suitable for further feature extraction
x_image = preprocess_input(x_image)
x_features = model.predict(x_image) # extract image features from model
x_features = np.array(x_features) # convert features list to numpy array
x_flatten= x_features.flatten() # flatten out the features in x
train_features.append(x_flatten) # this list contains the final features of the train images
# + colab={} colab_type="code" id="KzoY9-_eU2_W"
# Function to get the image size in the image directory
def file_size_test(file_name):
stats=os.stat("test_output_dir/"+str(file_name))
return stats.st_size
# + colab={} colab_type="code" id="kcxP_HbuU6eC"
# Function to call the keras pre-trained model to get feature weights of the test images
def resnet_model_test():
n=0
model = ResNet50(weights='imagenet', include_top=False, pooling='avg')
images_path = "./test_output_dir"
for f in os.listdir(images_path):
test_files.append(f)
# Looping over every image present in the files list
for img_path in test_files:
if(file_size_test(img_path)!=0):
print(str(img_path))
n+=1
print(n)
# load the image and resize it
img = image.load_img("./test_output_dir/"+str(img_path), target_size=(224, 224))
# extract features from each image
x_image = image.img_to_array(img)
x_image = np.expand_dims(x_image, axis=0) # increase dimensions of x to make it suitable for further feature extraction
x_image = preprocess_input(x_image)
x_features = model.predict(x_image) # extract image features from model
x_features = np.array(x_features) # convert features list to numpy array
x_flatten= x_features.flatten() # flatten out the features in x
test_features.append(x_flatten) # this list contains the final features of the test images
# + colab={"base_uri": "https://localhost:8080/", "height": 70159} colab_type="code" id="3lKtpdFRY4nb" outputId="e012a1a3-0f05-4de9-b3c9-a267142ff40e"
# Get features for every image extracted from train videos
train_features=[]
resnet_model_train()
# + colab={} colab_type="code" id="Rkwlp-8FjYf_"
# Scale all features with standard scaler
ss = StandardScaler()
train_features = ss.fit_transform(train_features)
# + colab={"base_uri": "https://localhost:8080/", "height": 30481} colab_type="code" id="dNFPEcpqjuCO" outputId="9b6adb4e-1a2d-4381-d38a-20fd1bae84a9"
# Get features for every image extracted from test videos
test_features=[]
test_files = []
resnet_model_test()
test_features = ss.transform(test_features) # scale the test video frames
# + colab={} colab_type="code" id="2rbOTkCrjwRE"
#Train the model with SVC on extracted features and provided labels
model = svm.SVC()
model.fit(train_features,label)
pred = model.predict(test_features) # predict the labels for test videos
# + colab={} colab_type="code" id="xXu96WuOMdRy"
# Strip all .jpg from the end of the filename
test_files1 = [s.strip('.jpg') for s in test_files]
# + colab={} colab_type="code" id="3txyvpgtupK1"
# Write the output in CSV format
label_output = pd.DataFrame(pred, columns=['label'])
testfile_output = pd.DataFrame(test_files1, columns=['file_name'])
# + colab={} colab_type="code" id="Dqe3PO2cNCeH"
result = pd.concat([testfile_output, label_output], axis=1, ignore_index = True)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="85tAy6-ANmjy" outputId="3e5e7df9-b3cb-4c0e-948d-f9f41d1832b3"
result.columns=['file_name','label']
result.head(5)
# + colab={} colab_type="code" id="nEIBzv5bNjMU"
result.to_csv('output.csv', header=True, index=False) # save output in csv
| videoclassification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multitask GP Regression
# Source: https://docs.gpytorch.ai/en/latest/examples/03_Multitask_Exact_GPs/Multitask_GP_Regression.html
# +
import math
import torch
import gpytorch as gpt
from matplotlib import pyplot as plt
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# -
# ### Set up training data
# +
train_x = torch.linspace(0, 1, 108)
train_y = torch.stack([
torch.sin(train_x * (2 * math.pi)) + torch.randn(train_x.size()) * 0.2,
torch.cos(train_x * (2 * math.pi)) + torch.randn(train_x.size()) * 0.2,
], -1)
# -
train_x.shape, train_y.shape
fig, ax = plt.subplots(1,1, figsize=(4,3))
ax.scatter(train_x.numpy(), train_y.numpy()[:,0])
ax.scatter(train_x.numpy(), train_y.numpy()[:,1])
plt.legend(["sin", "cos"])
# ### Define a multitask model
# +
class MultitaskGPModel(gpt.models.ExactGP):
def __init__(self, train_x, train_y, likelihood):
super(MultitaskGPModel, self).__init__(train_x, train_y, likelihood)
self.mean_module = gpt.means.MultitaskMean(
gpt.means.ConstantMean(), num_tasks=2
)
self.covar_module = gpt.kernels.MultitaskKernel(
gpt.kernels.RBFKernel(), num_tasks=2, rank=1
)
def forward(self, x):
mean_x = self.mean_module(x)
covar_x = self.covar_module(x)
return gpt.distributions.MultitaskMultivariateNormal(mean_x, covar_x)
likelihood = gpt.likelihoods.MultitaskGaussianLikelihood(num_tasks=2)
model = MultitaskGPModel(train_x, train_y, likelihood)
# -
# ### Train the model
# +
training_iters = 50
model.train()
likelihood.train()
optimizer = torch.optim.Adam(model.parameters(), lr=0.1)
# Loss for the marginal log likelihood
mll = gpt.mlls.ExactMarginalLogLikelihood(likelihood, model)
for i in range(training_iters):
optimizer.zero_grad()
output = model(train_x)
loss = -mll(output, train_y)
loss.backward()
print('Iter %d/%d - Loss: %.3f' % (i + 1, training_iters, loss.item()))
optimizer.step()
# -
# ### Make predictions with the model
# +
# Set into eval mode
model.eval()
likelihood.eval()
# Make predictions
with torch.no_grad(), gpt.settings.fast_pred_var():
test_x = torch.linspace(0, 1, 51)
predictions = likelihood(model(test_x))
mean = predictions.mean
lower, upper = predictions.confidence_region()
# This contains predictions for both tasks, flattened out
# The first half of the predictions is for the first task
# The second half is for the second task
# +
# Initialize plots
f, (y1_ax, y2_ax) = plt.subplots(1, 2, figsize=(8, 3))
# Plot training data as black stars
y1_ax.plot(train_x.detach().numpy(), train_y[:, 0].detach().numpy(), 'k*')
# Predictive mean as blue line
y1_ax.plot(test_x.numpy(), mean[:, 0].numpy(), 'b')
# Shade in confidence
y1_ax.fill_between(test_x.numpy(), lower[:, 0].numpy(), upper[:, 0].numpy(), alpha=0.5)
y1_ax.set_ylim([-3, 3])
y1_ax.legend(['Observed Data', 'Mean', 'Confidence'])
y1_ax.set_title('Observed Values (Likelihood)')
# Plot training data as black stars
y2_ax.plot(train_x.detach().numpy(), train_y[:, 1].detach().numpy(), 'k*')
# Predictive mean as blue line
y2_ax.plot(test_x.numpy(), mean[:, 1].numpy(), 'b')
# Shade in confidence
y2_ax.fill_between(test_x.numpy(), lower[:, 1].numpy(), upper[:, 1].numpy(), alpha=0.5)
y2_ax.set_ylim([-3, 3])
y2_ax.legend(['Observed Data', 'Mean', 'Confidence'])
y2_ax.set_title('Observed Values (Likelihood)')
# -
| experiments/05_multitask_gp_regression_gpytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (pynamical)
# language: python
# name: pynamical
# ---
# # Pynamical: demo of the Cubic Map, Singer Map, and Mandelbrot Map
#
# **Citation info**: <NAME>. 2016. "[Visual Analysis of Nonlinear Dynamical Systems: Chaos, Fractals, Self-Similarity and the Limits of Prediction](http://geoffboeing.com/publications/nonlinear-chaos-fractals-prediction/)." *Systems*, 4 (4), 37. doi:10.3390/systems4040037.
#
# Pynamical documentation: http://pynamical.readthedocs.org
#
# Pynamical can simulate, visualize, and explore any discrete dynamical system. This notebook demonstrates how to do this with two additional models that come included in the Pynamical module (the *Cubic Map* and the *Singer Map*) as well as with one additional model defined in this notebook (the *Mandelbrot Map*). Any other models can be created and easily plugged-in, as demonstrated below.
from numba import jit
from pynamical import simulate, cubic_map, singer_map, bifurcation_plot, phase_diagram, phase_diagram_3d
# %matplotlib inline
# ## Cubic map
#
# First we visualize the behavior of the *Cubic Map*. To do this, our code is nearly identical to the previous [examples using the Logistic Map](pynamical-demo-logistic-model.ipynb), with the exception that we are now passing `model=cubic_map` into our `simulate` function.
pops = simulate(model=cubic_map, num_gens=100, rate_min=1, rate_max=4, num_rates=1000, num_discard=100)
bifurcation_plot(pops, title='Cubic Map Bifurcation Diagram', xmin=1, xmax=4, save=False)
pops = simulate(model=cubic_map, num_gens=1000, rate_min=3.99, num_rates=1, num_discard=100)
phase_diagram(pops, xmin=-1, xmax=1, ymin=-1, ymax=1, save=False, title='Cubic Map 2D Phase Diagram')
pops = simulate(model=cubic_map, num_gens=1000, rate_min=3.99, num_rates=1, num_discard=100)
phase_diagram_3d(pops, xmin=-1, xmax=1, ymin=-1, ymax=1, zmin=-1, zmax=1, save=False, title='Cubic Map 3D Phase Diagram')
pops = simulate(model=cubic_map, num_gens=3000, rate_min=3.5, num_rates=30, num_discard=100)
phase_diagram_3d(pops, xmin=-1, xmax=1, ymin=-1, ymax=1, zmin=-1, zmax=1, save=False, alpha=0.2, color='viridis',
azim=330, title='Cubic Map 3D Phase Diagram, r = 3.5 to 4.0')
# ## Singer Map
#
# Next we visualize the behavior of the *Singer Map*. We pass `model=singer_map` into our `simulate` function.
pops = simulate(model=singer_map, num_gens=100, rate_min=0.9, rate_max=1.08, num_rates=1000, num_discard=100)
bifurcation_plot(pops, title='Singer Map Bifurcation Diagram', xmin=0.9, xmax=1.08, save=False)
pops = simulate(model=singer_map, num_gens=1000, rate_min=1.07, num_rates=1, num_discard=100)
phase_diagram(pops, title='Singer Map 2D Phase Diagram, r = 1.07', save=False)
pops = simulate(model=singer_map, num_gens=1000, rate_min=1.07, num_rates=1, num_discard=100)
phase_diagram_3d(pops, title='Singer Map 3D Phase Diagram, r = 1.07', save=False)
pops = simulate(model=singer_map, num_gens=3000, rate_min=1.04, rate_max=1.07, num_rates=30, num_discard=100)
phase_diagram_3d(pops, save=False, alpha=0.2, color='viridis', title='Singer Map 3D Phase Diagram, r = 1.04 to 1.07')
# ## ...Or define your own model
#
# Here we define the equation for the *Mandelbrot Map* and then visualize its behavior (use jit to compile it for fast simulation).
@jit(nopython=True)
def mandelbrot_map(pop, rate):
return pop ** 2 + rate
pops = simulate(model=mandelbrot_map, num_gens=100, rate_min=-2, rate_max=0, num_rates=1000, num_discard=100)
bifurcation_plot(pops, title='Mandelbrot Map Bifurcation Diagram', xmin=-2, xmax=0, ymin=-2, ymax=2, save=False)
pops = simulate(model=mandelbrot_map, num_gens=1000, rate_min=-1.99, num_rates=1, num_discard=100)
phase_diagram(pops, title='Mandelbrot Map 2D Phase Diagram', xmin=-2, xmax=2, ymin=-2, ymax=2, save=False)
pops = simulate(model=mandelbrot_map, num_gens=1000, rate_min=-1.99, num_rates=1, num_discard=100)
phase_diagram_3d(pops, title='Mandelbrot Map 3D Phase Diagram', xmin=-2, xmax=2, ymin=-2, ymax=2, zmin=-2, zmax=2, save=False)
pops = simulate(model=mandelbrot_map, num_gens=3000, rate_min=-1.99, rate_max=-1.5, num_rates=30, num_discard=100)
phase_diagram_3d(pops, title='Mandelbrot Map 3D Phase Diagram', alpha=0.2, color='viridis', color_reverse=True,
xmin=-2, xmax=2, ymin=-2, ymax=2, zmin=-2, zmax=2, save=False)
# ## For more info:
# - [Read the journal article](http://geoffboeing.com/publications/nonlinear-chaos-fractals-prediction/)
# - [Pynamical documentation](http://pynamical.readthedocs.org)
# - [Chaos Theory and the Logistic Map](http://geoffboeing.com/2015/03/chaos-theory-logistic-map/)
# - [Visualizing Chaos and Randomness with Phase Diagrams](http://geoffboeing.com/2015/04/visualizing-chaos-and-randomness/)
# - [Animated 3D Plots in Python](http://geoffboeing.com/2015/04/animated-3d-plots-python/)
| examples/pynamical-demo-other-models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="HUMzkjeKUwj9" colab_type="text"
# Import necessory modules
# + id="KYiL9HRbkBqX" colab_type="code" outputId="1d9ee4b0-2c98-4018-d5af-9bbe4ee6243e" colab={"base_uri": "https://localhost:8080/", "height": 87}
import numpy as np
import pandas as pd
import os
import keras
import cv2
from keras.layers import Conv2D, MaxPool2D, Flatten, Dense, Dropout, BatchNormalization
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential
from keras import regularizers
from sklearn.model_selection import train_test_split
from keras.utils import to_categorical
from keras.optimizers import SGD
from keras.preprocessing.image import ImageDataGenerator
from keras.models import load_model
from keras.callbacks import ModelCheckpoint
import matplotlib.pyplot as plt
import seaborn as sns
# + [markdown] id="bK006_gVU7hn" colab_type="text"
# Following code snippit is for colab users. Ignore it if you are not working on colab
#
# + id="2O9q90s_kOaD" colab_type="code" outputId="62b28f61-80ef-42b5-d3f4-889d84d49225" colab={"base_uri": "https://localhost:8080/", "height": 121}
from google.colab import drive
drive.mount('/content/drive')
# + id="uPf8eSXBkBqh" colab_type="code" outputId="cc60d5d3-3841-4ea1-fb7f-ee7d69455107" colab={"base_uri": "https://localhost:8080/", "height": 118}
train_dir = "/content/drive/My Drive/Virtual_Dermatologist/training_images"
# !ls "/content/drive/My Drive/Virtual_Dermatologist/training_images"
# + [markdown] id="YzDHb5pqVHLE" colab_type="text"
# `load_unique()` loads all the lables
# + id="A4I6GhO6kBql" colab_type="code" outputId="74eacf5e-417c-4faf-e8fe-65b91b5ecf3b" colab={"base_uri": "https://localhost:8080/", "height": 54}
def load_unique():
size_img = 256,256
images_for_plot = []
labels_for_plot = []
for folder in os.listdir(train_dir):
for file in os.listdir(train_dir + '/' + folder):
filepath = train_dir + '/' + folder + '/' + file
image = cv2.imread(filepath)
final_img = cv2.resize(image, size_img)
final_img = cv2.cvtColor(final_img, cv2.COLOR_BGR2RGB)
images_for_plot.append(final_img)
labels_for_plot.append(folder)
break
return images_for_plot, labels_for_plot
images_for_plot, labels_for_plot = load_unique()
print("unique_labels = ", labels_for_plot)
# + [markdown] id="0Vn4BBUiVVdc" colab_type="text"
# `plot_images()` is defined to visualise the images
# + id="TLRVS-x5kBqo" colab_type="code" outputId="27d9da5f-c6fd-4a72-dbcb-2fa1ceda3db9" colab={"base_uri": "https://localhost:8080/", "height": 319}
fig = plt.figure(figsize = (5,5))
def plot_images(fig, image, label, row, col, index):
fig.add_subplot(row, col, index)
plt.axis('off')
plt.imshow(image)
plt.title(label)
return
image_index = 0
row = 2
col = 1
for i in range(0,(row*col)):
plot_images(fig, images_for_plot[image_index], labels_for_plot[image_index], row, col, i+1)
image_index = image_index + 1
plt.show()
# + [markdown] id="gjRbu-JnVhJh" colab_type="text"
# `load_data()` loads the images and labels and also split it for testing purpose
# + id="B_IUKcBUkBqr" colab_type="code" colab={}
labels_dict = {'Acne-and-Rosacea':0,'Bullous-Disease':1,'Cellulitis-Impetigo-and-other-Bacterial-Infections':2
,'Eczema':3, 'Nail-Fungus':4, 'Melanoma-Skin-Cancer-Nevi-and-Moles':5}
def load_data():
images = []
labels = []
size = 256,256
print("LOADING DATA FROM : ",end = "")
for folder in os.listdir(train_dir):
print(folder, end = ' | ')
for image in os.listdir(train_dir + "/" + folder):
temp_img = cv2.imread(train_dir + '/' + folder + '/' + image)
temp_img = cv2.resize(temp_img, size)
images.append(temp_img)
if folder == 'Acne-and-Rosacea-Photos':
labels.append(labels_dict['Acne-and-Rosacea'])
elif folder == 'Bullous-Disease-Photos':
labels.append(labels_dict['Bullous-Disease'])
elif folder == 'Cellulitis-Impetigo-and-other-Bacterial-Infections':
labels.append(labels_dict['Cellulitis-Impetigo-and-other-Bacterial-Infections'])
elif folder == 'Eczema-Photos':
labels.append(labels_dict['Eczema'])
elif folder == 'Melanoma-Skin-Cancer-Nevi-and-Moles':
labels.append(labels_dict['Melanoma-Skin-Cancer-Nevi-and-Moles'])
elif folder == 'Seborrheic-Keratoses-and-other-Benign-Tumors':
labels.append(labels_dict['Seborrheic-Keratoses-and-other-Benign-Tumors'])
elif folder == 'Nail-Fungus-and-other-Nail-Disease':
labels.append(labels_dict['Nail-Fungus'])
images = np.array(images)
#images = images.astype('float32')/255.0
templabels = labels
labels = keras.utils.to_categorical(labels)
X_train, X_test, Y_train, Y_test = train_test_split(images, labels, test_size = 0.000000001)
#print()
#print('Loaded', len(X_train),'images for training,','Train data shape =',X_train.shape)
#print('Loaded', len(X_test),'images for testing','Test data shape =',X_test.shape)
return X_train, Y_train, X_test, Y_test, templabels
# + id="KJ1u8bLakBqv" colab_type="code" outputId="c9f26fdc-e984-4b50-c331-824ffe47630d" colab={"base_uri": "https://localhost:8080/", "height": 87}
X_train, Y_train, X_test, Y_test, labels = load_data()
# + [markdown] id="FXQNz_KMWHyM" colab_type="text"
# Following code snippit normalise the image
# + id="_aRQf-9yuX7L" colab_type="code" colab={}
#x_train = np.asarray(X_train.tolist())
#x_test = np.asarray(X_test.tolist())
x_train_mean = np.mean(X_train)
x_train_std = np.std(X_train)
x_test_mean = np.mean(X_test)
x_test_std = np.std(X_test)
x_train = (X_train - x_train_mean)/x_train_std
x_test = (X_test - x_test_mean)/x_test_std
# + id="gdlhszVXnZmL" colab_type="code" colab={}
x_train = (X_train)/255
x_test = (X_test)/255
# + id="CaWLigU1kBqx" colab_type="code" outputId="ec8e67b4-e631-41bf-d028-534a1d5775ba" colab={"base_uri": "https://localhost:8080/", "height": 462}
labelsdf = pd.DataFrame()
labelsdf["labels"] = labels_dict
labelsdf["labels"].value_counts().plot(kind = "bar", figsize = (7,3))
# + [markdown] id="Dsqp3SgZWmZD" colab_type="text"
# Following code snippit contains the CNN architecture
# + id="mQsPoXs_h3Kb" colab_type="code" colab={}
from keras.optimizers import Adam
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPool2D, BatchNormalization, LeakyReLU, Activation
from keras.layers.advanced_activations import LeakyReLU
def create_model():
input_shape = (256, 256, 3)
num_classes = 6
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3),activation='relu',padding = 'Same',input_shape=input_shape))
model.add(Conv2D(32,kernel_size=(3, 3), activation='relu',padding = 'Same',))
model.add(MaxPool2D(pool_size = (2, 2)))
model.add(Dropout(0.25))
model.add(Conv2D(64, (3, 3), activation='relu',padding = 'Same'))
model.add(Conv2D(64, (3, 3), activation='relu',padding = 'Same'))
model.add(MaxPool2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
print("MODEL CREATED")
model.summary()
return model
# + id="wRPCqDkixFg_" colab_type="code" colab={}
datagen = ImageDataGenerator(
featurewise_center=False, # set input mean to 0 over the dataset
samplewise_center=False, # set each sample mean to 0
featurewise_std_normalization=False, # divide inputs by std of the dataset
samplewise_std_normalization=False, # divide each input by its std
zca_whitening=False, # apply ZCA whitening
rotation_range=10, # randomly rotate images in the range (degrees, 0 to 180)
zoom_range = 0.1, # Randomly zoom image
width_shift_range=0.1, # randomly shift images horizontally (fraction of total width)
height_shift_range=0.1, # randomly shift images vertically (fraction of total height)
horizontal_flip=False, # randomly flip images
vertical_flip=False) # randomly flip images
datagen.fit(x_train)
from keras.callbacks import EarlyStopping
def fit_model():
checkpoint = ModelCheckpoint(
'/content/drive/My Drive/Virtual_Dermatologist/Checkpoints' + '/trial-weights-improvement-{epoch:02d}-{loss:.4f}-bigger.hdf5',
monitor='loss',
verbose=0,
save_best_only=True,
mode='min')
#callbacks = [EarlyStopping(monitor='val_acc', patience=4, verbose=1, mode='max'), checkpoint]
model_hist = model.fit(datagen.flow(x_train,Y_train, batch_size=32), epochs = 200, validation_data = (x_test, Y_test), verbose=1)
return model_hist
# + id="JYa2lsVgkBq2" colab_type="code" colab={}
from keras.callbacks import EarlyStopping
def fit_model():
#callbacks = [EarlyStopping(monitor='val_acc', patience=4, verbose=1, mode='max')]
model_hist = model.fit(x_train, Y_train, batch_size = 32, epochs = 100, validation_split = 0.1, verbose=1)
return model_hist
# + id="6Vli0TqQkBq4" colab_type="code" outputId="f7513348-abe8-4b04-ca14-b56b5b0bf901" colab={"base_uri": "https://localhost:8080/", "height": 1000}
model = create_model()
curr_model_hist = fit_model()
# + [markdown] id="xFudAl-ZWvZM" colab_type="text"
# Following code snippit saves the .h5 file of model
# + id="lFLSgNkGJgG6" colab_type="code" outputId="844af1dd-6969-481c-9b48-4fe107ea4c90" colab={"base_uri": "https://localhost:8080/", "height": 178}
model.save('/content/drive/My Drive/Virtual_Dermatologist/Checkpoints/complete_trained_model_200epoc.h5')
model.save('/content/drive/My Drive/Virtual_Dermatologist/Checkpoints/complete_trained_model_200epoc.model')
# + [markdown] id="YGB_8GduW9M7" colab_type="text"
# Test accuracy is predicted as follows
# + id="eC-y9lyEkBq6" colab_type="code" outputId="9342ffe4-9478-41fb-8cfe-c68a58dbe502" colab={"base_uri": "https://localhost:8080/", "height": 84}
evaluate_metrics = model.evaluate(x_test, Y_test)
print("\nEvaluation Accuracy = ", "{:.2f}%".format(evaluate_metrics[1]*100),"\nEvaluation loss = " ,"{:.6f}".format(evaluate_metrics[0]))
# + id="5RIqUQzOkBq-" colab_type="code" outputId="61b91978-30e6-4b53-a5bf-5a0e01c54c2c" colab={"base_uri": "https://localhost:8080/", "height": 573}
def plot_accuracy(y):
if(y == True):
plt.plot(curr_model_hist.history['acc'])
plt.plot(curr_model_hist.history['val_acc'])
plt.legend(['train', 'test'], loc='lower right')
plt.title('accuracy plot - train vs test')
plt.xlabel('epoch')
plt.ylabel('accuracy')
plt.show()
else:
pass
return
def plot_loss(y):
if(y == True):
plt.plot(curr_model_hist.history['loss'])
plt.plot(curr_model_hist.history['val_loss'])
plt.legend(['training loss', 'validation loss'], loc = 'upper right')
plt.title('loss plot - training vs vaidation')
plt.xlabel('epoch')
plt.ylabel('loss')
plt.show()
else:
pass
return
plot_accuracy(True)
plot_loss(True)
# + id="kMwGkt_yQs7d" colab_type="code" colab={}
from keras.utils import to_categorical
from numpy import argmax
model = load_model('/content/drive/My Drive/Virtual_Dermatologist/Checkpoints/complete_trained_model_200epoc.model')
y_pred = np.round( model.predict_classes(x_test), 0)
# + id="qtym98_b5Nli" colab_type="code" outputId="f60fdacb-5c2e-4f72-bc40-a1287996a152" colab={"base_uri": "https://localhost:8080/", "height": 104}
from sklearn.preprocessing import OneHotEncoder
onehot_encoder = OneHotEncoder(sparse=False)
y_pred1 = y_pred.reshape(len(y_pred), 1)
onehot_encoded = onehot_encoder.fit_transform(y_pred1)
# + id="Eit_Bhj-4mW9" colab_type="code" outputId="4c655e49-7463-41a3-d6b2-a13191cca57c" colab={"base_uri": "https://localhost:8080/", "height": 134}
onehot_encoded
# + id="p_l0H9B-2W3U" colab_type="code" outputId="83d1fa5e-df65-4dcd-e4c2-61bc4f8afad2" colab={"base_uri": "https://localhost:8080/", "height": 34}
from sklearn.metrics import roc_auc_score
print("ROC AUC Score: ",roc_auc_score(Y_test, onehot_encoded))
# + id="Wg4JkFFuslP-" colab_type="code" outputId="bb280ce6-42c8-416b-f4c7-b09da11980c6" colab={"base_uri": "https://localhost:8080/", "height": 235}
from sklearn.metrics import f1_score
from sklearn import metrics
print(metrics.classification_report(Y_test.argmax(axis = 1), y_pred, digits=3))
# + id="mtEPdMUesPVR" colab_type="code" outputId="24d70c6d-5526-4eeb-ec49-eeb50caac404" colab={"base_uri": "https://localhost:8080/", "height": 118}
from sklearn.metrics import confusion_matrix
matrix = confusion_matrix(Y_test.argmax(axis=1), y_pred)
matrix
# + id="uZy3Fv80tMc7" colab_type="code" outputId="a7d3193e-93a3-4774-ab39-19fc27bda3db" colab={"base_uri": "https://localhost:8080/", "height": 218}
from sklearn.preprocessing import normalize
normed_matrix = normalize(matrix, axis=1, norm='l1')
normed_matrix
# + id="9SSLri2qMX_2" colab_type="code" outputId="25ceac89-b6d1-4445-8116-2606a520dcfe" colab={"base_uri": "https://localhost:8080/", "height": 271}
import seaborn as sns;
ax = sns.heatmap(normed_matrix, annot=True)
ax
# + id="Qyj66qWekBrB" colab_type="code" colab={}
predicted = model.predict(X_test)
pdt = []
for i in predicted:
if(i[0]>i[1]):
pdt.append([1.,0.])
else:
pdt.append([0.,1.])
pdt = np.array(pdt)
#pdt
# + id="BD4c8zvlkBrG" colab_type="code" outputId="c7b02cfe-35c6-4fc7-ea84-0c35b7aa8948" colab={"base_uri": "https://localhost:8080/", "height": 154}
loaded_model = load_model('/content/drive/My Drive/Virtual_Dermatologist/Checkpoints/complete_trained_model_200epoc.h5')
# + [markdown] id="6KSjBciUXH96" colab_type="text"
# Following code snippit can be used to test model in realtime or on custom image
# + id="trWq60d_kBrK" colab_type="code" outputId="eb6229fc-6eb9-4d84-bf56-7d91264fbd80" colab={"base_uri": "https://localhost:8080/", "height": 306}
images = []
size = 256,256
img_file = '/content/drive/My Drive/Virtual_Dermatologist/Aditya.jpg'
temp_img = cv2.imread(img_file)
temp_img = cv2.resize(temp_img, size)
images.append(temp_img)
temp_img_mean = np.mean(images)
temp_img_std = np.std(images)
x_temp_img = (images -temp_img_mean)/temp_img_std
i = loaded_model.predict(x_temp_img)
print(i)
pred_lable = i[0].index(max(i[0]))
if(pred_lable==0):
print("Acne")
elif(pred_lable==1):
print("Bullous Disease")
elif(pred_lable=2):
print("Cellulitus")
elif(pred_lable==3):
print("Eczema")
elif(pred_lable==4):
print("Melanoma")
elif(pred_lable==5):
print("<NAME>")
from IPython.display import Image
Image(filename=img_file,
width=256,
height=256,)
# + [markdown] id="LJ2YoDn7Xcpc" colab_type="text"
# Following code snippit converts the model to tflite file for further use in an android application
# + id="l6NcjGNR_FxM" colab_type="code" outputId="15841652-4bca-4a43-b881-ed4ec2bbf43d" colab={"base_uri": "https://localhost:8080/", "height": 188}
import tensorflow as tf
converter = tf.lite.TFLiteConverter.from_keras_model_file('/content/drive/My Drive/Virtual_Dermatologist/Checkpoints/complete_trained_model_200epoc.h5')
tflite_model = converter.convert()
open("/content/drive/My Drive/Virtual_Dermatologist/skin_tflite.tflite", "wb").write(tflite_model)
print("testf2.tflite generated")
| skin_disease_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# __@this notebook__ will guide you through a very simple case of generative adversarial networks.
#
# <img src="https://raw.githubusercontent.com/torch/torch.github.io/master/blog/_posts/images/model.png" width=320px height=240px>
#
# Like.. veeery simple. Generative adversarial networks that learn to convert 1d uniform noise distribution to a uniform 1d normal data distribution.
#
# Unlike the main notebooks (`adversarial_*.ipynb`), this one features a lot of useful visualizations that will help you both get acquainted with the behavior of two networks and debug common errors without having to wait hours of CPU time.
# +
import numpy as np
import torch, torch.nn as nn
import torch.nn.functional as F
def sample_noise(batch_size):
""" Uniform noise of shape [batch_size, 1] in range [0, 1]"""
return torch.rand(batch_size, 1)
def sample_real_data(batch_size):
""" Normal noise of shape [batch_size, 1], mu=5, std=1.5 """
return torch.randn(batch_size, 1) * 1.5 + 5
# +
# Generator converts 1d noise into 1d data
gen = nn.Sequential(nn.Linear(1, 16), nn.ELU(), nn.Linear(16, 1))
gen_opt = torch.optim.SGD(gen.parameters(), lr=1e-3)
# Discriminator converts 1d data into two logits (0th for real, 1st for fake).
# It is deliberately made stronger than generator to make sure disc is slightly "ahead in the game".
disc = nn.Sequential(nn.Linear(1, 64), nn.ELU(), nn.Linear(64, 2))
disc_opt = torch.optim.SGD(disc.parameters(), lr=1e-2)
# +
# we define 0-th output of discriminator as "is_fake" output and 1-st as "is_real"
IS_FAKE, IS_REAL = 0, 1
def train_disc(batch_size):
""" trains discriminator for one step """
# compute logp(real | x)
real_data = sample_real_data(batch_size)
logp_real_is_real = F.log_softmax(disc(real_data), dim=1)[:, IS_REAL]
# compute logp(fake | G(z)). We detach to avoid computing gradinents through G(z)
noise = <sample noise>
gen_data = <generate data given noise>
logp_gen_is_fake = <compute logp for 0th>
disc_loss = <compute loss>
# sgd step. We zero_grad first to clear any gradients left from generator training
disc_opt.zero_grad()
disc_loss.backward()
disc_opt.step()
return disc_loss.item()
# -
def train_gen(batch_size):
""" trains generator for one step """
# compute logp(fake | G(z)).
noise = <sample noise>
gen_data = <generate data given noise>
logp_gen_is_real = <compute logp gen is REAL>
gen_loss = <generator loss>
gen_opt.zero_grad()
gen_loss.backward()
gen_opt.step()
return gen_loss.item()
# +
import matplotlib.pyplot as plt
from IPython.display import clear_output
# %matplotlib inline
for i in range(100000):
for _ in range(5):
train_disc(128)
train_gen(128)
if i % 250 == 0:
clear_output(True)
plt.figure(figsize=[14, 6])
plt.subplot(1, 2, 1)
plt.title("Data distributions")
plt.hist(gen(sample_noise(1000)).data.numpy()[:, 0],
range=[0, 10], alpha=0.5, normed=True, label='gen')
plt.hist(sample_real_data(1000).data.numpy()[:,0],
range=[0, 10], alpha=0.5, normed=True, label='real')
x = np.linspace(0, 10, dtype='float32')
disc_preal = F.softmax(disc(torch.from_numpy(x[:, None])))[:, 1]
plt.plot(x, disc_preal.data.numpy(), label='disc P(real)')
plt.legend()
plt.subplot(1, 2, 2)
plt.title("Discriminator readout on real vs gen")
plt.hist(F.softmax(disc(gen(sample_noise(100))))[:, 1].data.numpy(),
range=[0, 1], alpha=0.5, label='D(G(z))')
plt.hist(F.softmax(disc(sample_real_data(100)))[:, 1].data.numpy(),
range=[0, 1], alpha=0.5, label='D(x)')
plt.legend()
plt.show()
# -
# __What to expect:__
# * __left:__ two distributions will start differently, but generator distribution should match real data _almost_ everywhere. The curve represents discriminator's opinion on all possible values of x. It should slowly get closer to 0.5 over areas where real data is dense.
# * __right:__ this chart shows how frequently does discriminator assign given probability to samples from real and generated data samples (shown in different colors). First several iterations will vary, but eventually they will both have nearly all probability mass around 0.5 as generator becomes better at it's job.
# * If instead it converges to two delta-functions around 0(gen) and 1(real) each, your discriminator has won. _Check generator loss function_. As a final measure, try decreasing discriminator learning rate. This can also happen if you replace mean over batch with sum or similar.
# * If it converges to 0.5 and stays there for several iterations but generator haven't learned to generate plausible data yet, generator is winning the game. _Double-check discriminator loss function_.
| homework03/simple_1d_gan_pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="2qmoeBUEBmvU"
# # Setup
# + [markdown] id="7F6PH06NCRGP"
# Mount Google Drive
# + colab={"base_uri": "https://localhost:8080/"} id="ZjHHUmz1Bf1N" executionInfo={"status": "ok", "timestamp": 1620171555402, "user_tz": 240, "elapsed": 87009, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjuu5ZxulIvlUF4kQ7Fx-JEkYMunqTV1k93wXJu=s64", "userId": "16538749842847312111"}} outputId="24936ff2-11cd-4f89-e03d-9fb1163a1d13"
from google.colab import drive
drive.mount('/content/gdrive/')
import sys
project_path = ('/content/gdrive/My Drive/DATA_690_NLP/Text_Summarization/')
sys.path.append(project_path)
# + [markdown] id="esrr0AdbCU4V"
# Import packages
# + id="bzu1KQmPCHa9" executionInfo={"status": "ok", "timestamp": 1620181410627, "user_tz": 240, "elapsed": 393, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjuu5ZxulIvlUF4kQ7Fx-JEkYMunqTV1k93wXJu=s64", "userId": "16538749842847312111"}}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from textblob import TextBlob
import seaborn as sns
sns.set()
% matplotlib inline
np.random.seed(42)
# + [markdown] id="fpHhwcibCWiG"
# Load articles
# + colab={"base_uri": "https://localhost:8080/", "height": 695} id="dnXgI29xCYvF" executionInfo={"status": "ok", "timestamp": 1620171559266, "user_tz": 240, "elapsed": 90856, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjuu5ZxulIvlUF4kQ7Fx-JEkYMunqTV1k93wXJu=s64", "userId": "16538749842847312111"}} outputId="9bddfe19-365e-4899-e716-8b43810df029"
articles_df = pd.read_csv(f'{project_path}Data/processed/Clean_Lemma_LongForm_Telehealth.csv')
articles_df.head()
# + [markdown] id="xwicWnldCZx-"
# Load clustering analysis
# + colab={"base_uri": "https://localhost:8080/", "height": 330} id="58UAdyLQCcEb" executionInfo={"status": "ok", "timestamp": 1620171559433, "user_tz": 240, "elapsed": 91017, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjuu5ZxulIvlUF4kQ7Fx-JEkYMunqTV1k93wXJu=s64", "userId": "16538749842847312111"}} outputId="ed0b39f5-68e5-4e71-cefc-f06c92ce3119"
clustering_df = pd.read_csv(f'{project_path}/Data/clustering_results.csv')
clustering_df.head()
# + [markdown] id="RDcb-0PCC08Y"
# # Adding polarity and subjectivity values
# + colab={"base_uri": "https://localhost:8080/", "height": 330} id="6s3C2AefDErf" executionInfo={"status": "ok", "timestamp": 1620171562644, "user_tz": 240, "elapsed": 94219, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjuu5ZxulIvlUF4kQ7Fx-JEkYMunqTV1k93wXJu=s64", "userId": "16538749842847312111"}} outputId="772480af-57d9-4fd2-e243-5fb4abe1a58e"
clustering_df['Polarity'] = articles_df['Clean_Content_Lemma_LongForm'].apply(lambda x: TextBlob(x).sentiment.polarity)
clustering_df['Subjectivity'] = articles_df['Clean_Content_Lemma_LongForm'].apply(lambda x: TextBlob(x).sentiment.subjectivity)
clustering_df.head()
# + [markdown] id="m3GgQv2Wu7ch"
# Save data
# + colab={"base_uri": "https://localhost:8080/", "height": 625} id="ZboOsnq_u6sf" executionInfo={"status": "ok", "timestamp": 1620171810814, "user_tz": 240, "elapsed": 337, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjuu5ZxulIvlUF4kQ7Fx-JEkYMunqTV1k93wXJu=s64", "userId": "16538749842847312111"}} outputId="67035f36-a152-44f8-b72d-7784a04e3010"
clustering_df['Journal Title'] = articles_df['Journal Title']
clustering_df['Article Title'] = articles_df['Article Title']
clustering_df['Abstract'] = articles_df['Abstract']
clustering_df['Keywords'] = articles_df['Keywords']
clustering_df['Content_Length'] = articles_df['Content_Length']
clustering_df.to_csv(f'{project_path}/Data/clustering_results.csv')
clustering_df.head()
# + [markdown] id="wRN39toKGrCP"
# # Analysis
# + [markdown] id="f89ukw8DGs_F"
# Comparing the variance in polarity by cleaned_pca_dbscan_class
# + colab={"base_uri": "https://localhost:8080/"} id="sSe5rc9jG6yU" executionInfo={"status": "ok", "timestamp": 1620173754006, "user_tz": 240, "elapsed": 254, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjuu5ZxulIvlUF4kQ7Fx-JEkYMunqTV1k93wXJu=s64", "userId": "16538749842847312111"}} outputId="5566863b-bfa6-419b-8c11-4b9a8a7dac83"
cleaned_pca_df = clustering_df[(clustering_df['cleaned_pca_dbscan_class'] != -1)&(clustering_df['cleaned_tsne_dbscan_class'] != -1)][['cleaned_pca_dbscan_class', 'cleaned_tsne_dbscan_class', 'Polarity', 'Subjectivity', 'Content_Length']]
print(cleaned_pca_df['cleaned_pca_dbscan_class'].value_counts())
print(cleaned_pca_df['cleaned_tsne_dbscan_class'].value_counts())
# + colab={"base_uri": "https://localhost:8080/", "height": 609} id="c6bpn4inIPdQ" executionInfo={"status": "ok", "timestamp": 1620181436049, "user_tz": 240, "elapsed": 725, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjuu5ZxulIvlUF4kQ7Fx-JEkYMunqTV1k93wXJu=s64", "userId": "16538749842847312111"}} outputId="1dc2676f-2b3d-4608-8249-d31da4d57873"
pca_class_var_df = cleaned_pca_df[['Polarity', 'Subjectivity']].var().to_frame().T
pca_class_var_df['Cluster'] = 'Overall'
for db_scan_class, count in cleaned_pca_df['cleaned_pca_dbscan_class'].value_counts().items():
cluster_df = cleaned_pca_df[cleaned_pca_df['cleaned_pca_dbscan_class'] == db_scan_class]
cluster_df = cluster_df[['Polarity', 'Subjectivity']].var().to_frame().T
cluster_df['Cluster'] = db_scan_class
pca_class_var_df = pd.concat([pca_class_var_df, cluster_df], ignore_index=True)
pca_class_var_df.set_index('Cluster')
print(cleaned_pca_df.value_counts('cleaned_pca_dbscan_class'))
a4_dims = (11.7, 8.27)
fig, ax = plt.subplots(figsize=a4_dims)
sns.lineplot(data=pca_class_var_df)
# + colab={"base_uri": "https://localhost:8080/", "height": 936} id="XaBplfoH3bb7" executionInfo={"status": "error", "timestamp": 1620181601302, "user_tz": 240, "elapsed": 659, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjuu5ZxulIvlUF4kQ7Fx-JEkYMunqTV1k93wXJu=s64", "userId": "16538749842847312111"}} outputId="6c28fd0a-079b-4de3-df01-ed4c3ec9e10e"
tsne_class_var_df = cleaned_pca_df[['Polarity', 'Subjectivity']].var().to_frame().T
tsne_class_var_df['Cluster'] = 'Overall'
for db_scan_class, count in cleaned_pca_df['cleaned_tsne_dbscan_class'].value_counts().items():
cluster_df = cleaned_pca_df[cleaned_pca_df['cleaned_tsne_dbscan_class'] == db_scan_class]
cluster_df = cluster_df[['Polarity', 'Subjectivity']].var().to_frame().T
cluster_df['Cluster'] = db_scan_class
tsne_class_var_df = pd.concat([tsne_class_var_df, cluster_df], ignore_index=True)
#tsne_class_var_df.set_index('Cluster')
print(cleaned_pca_df.value_counts('cleaned_tsne_dbscan_class'))
a4_dims = (11.7, 8.27)
fig, ax = plt.subplots(figsize=a4_dims)
sns.lineplot(data=tsne_class_var_df, x=['Cluster'])
| notebooks/Correlation_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Project 3 - Water Pump Clasificaiton
# # Setup
from __future__ import print_function
# +
import pandas as pd
import numpy as np
import seaborn as sns
from os import system
import matplotlib.pyplot as plt
# %matplotlib inline
from __future__ import division
pd.set_option('display.width',5000)
# +
import patsy
from sklearn import linear_model as lm
from sklearn.linear_model import LogisticRegression
from sklearn import cross_validation
from sklearn import metrics
from sklearn.metrics import confusion_matrix
# -
from sklearn.tree import DecisionTreeClassifier
from sklearn.svm import LinearSVC
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
# +
# system('say I am Done!')
# -
# # Data Import - Training Data
# This data is part of the Data Driven Competition
#
# https://www.drivendata.org/competitions/7/pump-it-up-data-mining-the-water-table/page/23/
#
# +
# 'Values' provided for each pump location - features
df_values = pd.read_csv('/Users/amycurneen/ds/metis/metisgh/Metis-Curneen/3 - Water Pumps/Data Downloads/Training set values.csv')
df_values.sample(1)
# +
# 'Labels' provided for each pump location - clasificaiton - what I am predicting
df_labels = pd.read_csv('/Users/amycurneen/ds/metis/metisgh/Metis-Curneen/3 - Water Pumps/Data Downloads/Training set labels.csv')
df_labels.sample(1)
# -
# # Data Import - Challenge Data
# +
# 'Values' provided for each competition pump location - features
df_test_values = pd.read_csv('/Users/amycurneen/ds/metis/metisgh/Metis-Curneen/3 - Water Pumps/Data Downloads/Test set values.csv')
df_test_values.sample(1)
# +
# 'Labels' I will provide for each pump location - clasificaiton
df_sub = pd.read_csv('/Users/amycurneen/ds/metis/metisgh/Metis-Curneen/3 - Water Pumps/Data Downloads/SubmissionFormat.csv')
df_sub = df_sub.drop('status_group', axis = 1)
df_sub.sample(1)
# -
# # Feature Analysis
# My goal is to predict the operating condition of a waterpoint for each record in the dataset. I was provided the following set of information about the waterpoints:
#
# * amount_tsh - Total static head (amount water available to waterpoint)
# * 98 unique
# * date_recorded - The date the row was entered
# * 365 unique
# * year - month - day
# * funder - Who funded the well
# * 1897 unique
# * look at top ones?
# * installer - Organization that installed the well
# * 2145 unique
# * DWE is main one - 10x closest other, 17k
# * wpt_name - Name of the waterpoint if there is one
# * 37400 unique
# * look at top ones?
# * num_private - (NO PROVIDED DESC)
# * 65 unique
# * USELESS FEATURE
# * population - Population around the well
# * 1049 unique
# * a lot are zero
# * public_meeting - True/False
# * 2 unique
# * recorded_by - Group entering this row of data
# * 1 unique
# * all the same - USELESS FEATURE
# * scheme_management - Who operates the waterpoint
# * 12 unique
# * scheme_name - Who operates the waterpoint
# * 2696 unique
# * USELESS FEATURE
# * permit - If the waterpoint is permitted
# * 2 unique
# * construction_year - Year the waterpoint was constructed
# * 55 unique
# * third are 0 - USELESS FEATURE
#
#
# * Geography
# * gps_height - Altitude of the well
# * numerical
# * longitude - GPS coordinate
# * numerical
# * latitude - GPS coordinate
# * numerical
# * basin - Geographic water basin
# * 9 unique
# * subvillage - Geographic location
# * 19287 unique
# * region - Geographic location
# * 21 unique
# * region_code - Geographic location (coded)
# * 27 unique
# * district_code - Geographic location (coded)
# * 20 unique
# * lga - Geographic location
# * 125 unique
# * ward - Geographic location
# * 2092 unique
#
#
# * Extraction
# * extraction_type - The kind of extraction the waterpoint uses
# * 18 unique
# * Most descriptive of extraction
# * extraction_type_group - The kind of extraction the waterpoint uses
# * 13 unique
# * Parent of extraction_type
# * extraction_type_class - The kind of extraction the waterpoint uses
# * 7 unique
# * Parent of extraction_type_group
#
#
# * Overhead
# * management - How the waterpoint is managed
# * 12 unique
# * management_group - How the waterpoint is managed
# * 5 unique
# * payment - What the water costs
# * 7 unique
# * same as payment type
# * payment_type - What the water costs
# * 7 unique
# * same as payment
#
#
# * Water
# * water_quality - The quality of the water
# * 3 unique
# * Subset of quality_group
# * quality_group - The quality of the water
# * 6 unique
# * Parent group of water_quality
# * quantity - The quantity of water
# * 5 unique
# * Same as quantity_group
# * quantity_group - The quantity of water
# * 5 unique
# * Same as quantity
# * source - The source of the water
# * 10 unique
# * source_type - The source of the water
# * 7 unique
# * Subset of source
# * source_class - The source of the water
# * 3 unique
# * Subset of source_type
# * waterpoint_type - The kind of waterpoint
# * 6 unique
# * Parent of waterpoint_type_group
# * waterpoint_type_group - The kind of waterpoint
# * 7 unique
# * Subset of waterpoint_type
myregions = list(df_values.region.unique())
myregions.sort()
df_values.region.value_counts()
payment = list(df_values.payment.unique())
payment.sort()
payment
# # Sorting features
total = list(df_values.columns)
useless = ['id','date_recorded','num_private','recorded_by','scheme_name','construction_year','subvillage','ward',
'payment_type','quantity_group','wpt_name']
subsets_to_go = ['quality_group','extraction_type_group','extraction_type','source','source_type',
'waterpoint_type_group','management']
numerical = ['amount_tsh','population','latitude','longitude','gps_height']
non_numerical = list(set(total) - set(useless) - set(subsets_to_go) - set(numerical))
my_features = numerical+non_numerical
# +
# get rid of features that dont show often
map_funder = df_values.funder.value_counts().to_dict()
for i in range(len(map_funder.keys())):
keys = list(map_funder.keys())
a = keys[i]
if map_funder[a] > 800:
map_funder[a] = a
else:
map_funder[a] = 'other'
# -
df_values['funder'].replace(map_funder, inplace=True)
# +
# get rid of features that dont show often
map_installer = df_values.installer.value_counts().to_dict()
for i in range(len(map_installer.keys())):
keys = list(map_installer.keys())
a = keys[i]
if map_installer[a] > 800:
map_installer[a] = a
else:
map_installer[a] = 'other'
# -
df_values['installer'].replace(map_installer, inplace=True)
# ## Nans
df_values = df_values.fillna('other')
# # Setup chalange data
funder_list = list(set(map_funder.values()))
funder_list.remove('other')
installer_list = list(set(map_installer.values()))
installer_list.remove('other')
for i in range(len(df_test_values.funder)):
if df_test_values.funder[i] not in funder_list:
df_test_values.at[i, 'funder'] = 'other'
for i in range(len(df_test_values.funder)):
if df_test_values.installer[i] not in funder_list:
df_test_values.at[i, 'installer'] = 'other'
df_test_values = df_test_values.fillna('other')
df_test = df_test_values[my_features]
# # Create selected feature DataFrame
# ## Randomize
# +
# randomize data
# df_lables, df_values - combine and shuffle this data
df = pd.merge(df_labels,df_values,how = 'left')
df = df.sample(frac=1).reset_index(drop=True)
# -
my_features.insert(0,'status_group')
df_features = df[my_features]
map_status_group = {'functional':0,'functional needs repair':1,'non functional':2}
# turn y into 3 class 0,1,2
df_features['status_group'].replace(map_status_group, inplace=True)
# ## Create X and Y
y = df_features.status_group
X = df_features.drop('status_group', axis=1)
X.info()
X['region_code'] = X['region_code'].astype(object)
X['district_code'] = X['district_code'].astype(object)
# ## Test train split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2,stratify=y)
X_train.head()
# # Catboost for limited features
from catboost import CatBoostRegressor, CatBoostClassifier, Pool
# +
categorical_features_indices = np.where((X_train.dtypes != np.float) & (X_train.dtypes != np.int))[0]
train_pool = Pool(X_train, y_train, cat_features=categorical_features_indices)
model=CatBoostClassifier(iterations=200, depth=10, learning_rate=0.3,loss_function='MultiClass')
model.fit(train_pool,eval_set=(X_test, y_test),plot=True)
# -
# +
# model2=CatBoostClassifier(depth=10, learning_rate=0.3,loss_function='Recall')
# model2.fit(train_pool,plot=True)
# -
feature_importances = model.get_feature_importance(train_pool)
feature_names = X_train.columns
for score, name in sorted(zip(feature_importances, feature_names), reverse=True):
print('{}: {}'.format(name, score))
model.save_model('catboost_model.dump')
# +
# model.load_model('catboost_model.dump')
# +
# model.load_model('3_catboost_model_it200_depth14_learn025.dump')
# +
from sklearn.metrics import confusion_matrix
y_predict = model.predict(X_test)
conf = confusion_matrix(y_test, y_predict)
# -
conf
print ('Functional:')
print ("Precision: %0.2f" %(conf[0, 0] / (conf[0, 0] + conf[1, 0] + conf[2, 0])))
print ("Recall: %0.2f"% (conf[0, 0] / (conf[0, 0] + conf[0, 1] + conf[0, 2])))
print ('Functional but needs repair:')
print ("Precision: %0.2f" %(conf[1, 0] / (conf[0, 0] + conf[1, 0] + conf[2, 0])))
print ("Recall: %0.2f"% (conf[0, 1] / (conf[0, 0] + conf[0, 1] + conf[0, 2])))
print ('Non functioning:')
print ("Precision: %0.2f" %(conf[2, 0] / (conf[0, 0] + conf[1, 0] + conf[2, 0])))
print ("Recall: %0.2f"% (conf[0, 2] / (conf[0, 0] + conf[0, 1] + conf[0, 2])))
print ("Accuracy: %0.2f" %((conf[0, 0] + conf[1, 1] + conf[2, 2]) / (len(y_test))))
# # Create decision list
answers = list(model.predict(df_test))
# +
answer_better = []
for i in range(len(answers)):
num = int(answers[i][0])
answer_better.append(num)
# -
df_sub['status_group'] = answer_better
inv_map_status_group = {v: k for k, v in map_status_group.items()}
# turn y into 3 class 0,1,2
df_sub['status_group'].replace(inv_map_status_group, inplace=True)
df_sub['status_group'].value_counts()
df_sub.to_csv('./Submissions/decisionTree5_14_215pm.csv',index=False)
# + [markdown] heading_collapsed=true
# # Appendix
# + [markdown] heading_collapsed=true hidden=true
# ## A
# + hidden=true
columns = list(feature_sel.columns)
feature_sel[columns[5:]] = feature_sel[columns[5:]].astype(int)
feature_sel[columns[0]] = feature_sel[columns[0]].astype(int)
# + hidden=true
feature_sel = feature_sel.sample(frac=1).reset_index(drop=True)
y = feature_sel.status_group
X = feature_sel.drop('status_group', axis=1)
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.84)
# + hidden=true
X_train.head(1)
# + [markdown] heading_collapsed=true hidden=true
# ## Adjust competition data
# + hidden=true
df_test_values = df_test_values[['gps_height', 'longitude', 'latitude', 'basin',
'extraction_type_class','payment','quantity','water_quality', 'source_class',
'waterpoint_type_group','population','public_meeting','scheme_management']]
# + hidden=true
# rearange columns
cols = df_test_values.columns.tolist()
cols = ['gps_height',
'longitude',
'latitude',
'population',
'basin',
'extraction_type_class',
'payment',
'quantity',
'water_quality',
'source_class',
'waterpoint_type_group',
'public_meeting',
'scheme_management']
df_test_values = df_test_values[cols]
# + hidden=true
a = [0] * len(df_test_values['gps_height'])
# + hidden=true
df_test_values = pd.get_dummies(df_test_values,columns=list(df_test_values.columns[4:]))
df_test_values.sample(5)
# + hidden=true
columns = list(feature_sel.columns)
feature_sel[columns[5:]] = feature_sel[columns[5:]].astype(int)
feature_sel[columns[0]] = feature_sel[columns[0]].astype(int)
# + hidden=true
df_test_values['scheme_management_None'] = a
df_test_values.head(10)
# + hidden=true
list(set(X_train.columns)-set(df_test_values.columns))
# + hidden=true
# + [markdown] heading_collapsed=true hidden=true
# ## B
# + hidden=true
from __future__ import print_function
import numpy as np
import pandas as pd
from IPython.display import Image
from sklearn import datasets
from sklearn.decomposition import PCA
from sklearn.metrics import accuracy_score, confusion_matrix
from sklearn.cross_validation import train_test_split
from sklearn import svm
from matplotlib import pyplot as plt
# %matplotlib inline
# + hidden=true
pd.Series(y_train).value_counts()
# + [markdown] hidden=true
# ## Principle Component Analysis
# + hidden=true
pca = PCA(n_components=2)
pca.fit(X_train)
# + hidden=true
pcafeatures_train = pca.transform(X_train)
# + hidden=true
from itertools import cycle
# + hidden=true
# def plot_PCA_2D(data, target, target_names):
# colors = cycle(['r','g','b'])
# target_ids = range(len(target_names))
# plt.figure()
# for i, c, label in zip(target_ids, colors, target_names):
# plt.scatter(data[target == i, 0], data[target == i, 1],
# c=c, label=label)
# plt.legend()
# + hidden=true
# plot_PCA_2D(pcafeatures_train, target=y_train, target_names=digits.target_names)
# + [markdown] hidden=true
# ## Fitting Linear and RBF SVM Models
# + hidden=true
# fit linear model
model_svm = svm.SVC(kernel='rbf',probability=False,cache_size=2000)
model_svm.fit(X_train, y_train)
# + hidden=true
# predict out of sample
y_pred = model_svm.predict(X_test)
# + hidden=true
# check accuracy
accuracy_score(y_test,y_pred)
# + hidden=true
# confusion matrix
confusion_matrix(y_test,y_pred)
# + hidden=true
# fit rbf model
# model_svm2 = svm.SVC(kernel='rbf', gamma = 0.001)
# model_svm2.fit(X_train, y_train)
# + hidden=true
# predict out of sample
y_pred2 = model_svm2.predict(X_test)
# + hidden=true
# check accuracy
accuracy_score(y_test,y_pred2)
# + hidden=true
# confusion matrix
confusion_matrix(y_test,y_pred2)
# + [markdown] heading_collapsed=true hidden=true
# ## C
# + [markdown] hidden=true
# Extra code from class to utilize
# + hidden=true
df.age=df.age.fillna(df.age.mean())
# + hidden=true
y,X=dmatrices('survived~ pclass +age+sibsp+parch+fare',data=df,return_type='dataframe')
# + hidden=true
# Generate a confusion matrix plot:
def plot_confusion_matrix(cm,title='Confusion matrix', cmap=plt.cm.Reds):
plt.imshow(cm, interpolation='nearest',cmap=cmap)
plt.title(title)
plt.colorbar()
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
#Could be a typical function for classifying:
def train_score(classifier,x,y):
xtrain, xtest, ytrain, ytest = cross_validation.train_test_split(x, y, test_size=0.2, random_state=1234)
ytrain=np.ravel(ytrain)
clf = classifier.fit(xtrain, ytrain)
# accuracy for test & train:
train_acc=clf.score(xtrain, ytrain)
test_acc=clf.score(xtest,ytest)
print("Training Data Accuracy: %0.2f" %(train_acc))
print("Test Data Accuracy: %0.2f" %(test_acc))
y_true = ytest
y_pred = clf.predict(xtest)
conf = confusion_matrix(y_true, y_pred)
print(conf)
print ('\n')
print ("Precision: %0.2f" %(conf[0, 0] / (conf[0, 0] + conf[1, 0])))
print ("Recall: %0.2f"% (conf[0, 0] / (conf[0, 0] + conf[0, 1])))
cm=confusion_matrix(y_true, y_pred, labels=None)
plt.figure()
plot_confusion_matrix(cm)
# + hidden=true
log_clf=LogisticRegression()
train_score(log_clf,X,y)
# + hidden=true
# What about ROC ?
from sklearn.metrics import roc_curve, auc
xtrain, xtest, ytrain, ytest = cross_validation.train_test_split(X, y, test_size=0.2, random_state=1234)
log = LogisticRegression()
log.fit(xtrain,np.ravel(ytrain))
y_score=log.predict_proba(xtest)[:,1]
fpr, tpr,_ = roc_curve(ytest, y_score)
roc_auc = auc(fpr, tpr)
plt.figure()
# Plotting our Baseline..
plt.plot([0,1],[0,1])
plt.plot(fpr,tpr)
plt.xlabel('FPR')
plt.ylabel('TPR')
# + hidden=true
tpr
# + [markdown] hidden=true
# #### Cost Benefit Example:
#
# We can also optimize our models based on specific costs associated with our classification errors; here we will use specific dollar amounts as weights.
#
# Let's say we were developing a classification model for Aircraft Delay prediction. For this example let's assume that a true positive would
# lead to a cost savings of 2160 dollars, a false negative would cost us 2900 dollars a false positive would cost 750 dollars.
#
# cb = np.array([[2160, -750.0], [-2900, 0]])
#
# Expected_Value = #TPs(2160) - #FNs(2900) -#FPs(750)
| 3_Water_Pumps/.ipynb_checkpoints/CatBoost-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
###Made by <NAME>
###To process Iolite baseline-subtracted NP-II files for BB
# -
# To Do:
#
# -Test out Error minimization
#
# -Determine if filtering is the correct way to remove negative measurements from dataset.
# +
import pandas as pd
import os
import re
import copy
import numpy as np
from scipy import stats
from scipy.stats import linregress
#Graphing stuff
from matplotlib import pyplot as plt
# %matplotlib inline
# # %pip install seaborn
# import seaborn as sns
# #%pip install PyPDF2
from PyPDF2 import PdfFileMerger, PdfFileReader
# #%pip install pdfkit
import pdfkit
#pd.set_option("display.precision", 8)
# +
#Functions for calculation and EXCEL export
def read_np2_timeseries(excel_file):
''' Excel input file is your baseline corrected time series export from Iolite for the NP-II.'''
df = pd.read_excel(excel_file, sheet_name = None)
keys = df.keys()
header_row = 0
new_dict = {}
for key in keys:
if '.' in key: #Kind of hard-coded right now, so if names get weird may need to change
df_test = df[key]
df_test.columns = df_test.iloc[header_row]
df_test = df_test.drop(header_row)
df_test = df_test.reset_index(drop=True)
# test1_new = df_test[['Absolute Time',
# 'Elapsed Time',
# 'm238_CPS',
# 'm232_CPS',
# 'm208_CPS',
# 'm207_CPS',
# 'm206_CPS',
# 'm204_CPS',
# 'm202_CPS',]]
new_string = key.split('time')[0].rstrip()
new_dict[new_string] = df_test #test1_new
return new_dict
def calc_CPS(np2_dict):
columns = ['Absolute Time',
'Elapsed Time',
'm238_CPS',
'm232_CPS',
'm208_CPS',
'm207_CPS',
'm206_CPS',
'm204_CPS',
'm202_CPS']
new_col = ['Absolute Time',
'Elapsed Time',
'238_CPS',
'232_CPS',
'208_CPS',
'207_CPS',
'206_CPS',
'204_CPS',
'202_CPS']
cut_col = ['238_CPS',
'232_CPS',
'208_CPS',
'207_CPS',
'206_CPS',
'204_CPS',
'202_CPS']
calc_dict = {}
for key in np2_dict:
#print(key)
test_df1 = np2_dict[key]
for col in columns:
test_df2 = test_df1.apply(lambda x: x * 62500000 if 'CPS' in x.name else x)
test_df2 = test_df2[['Absolute Time',
'Elapsed Time',
'm238_CPS',
'm232_CPS',
'm208_CPS',
'm207_CPS',
'm206_CPS',
'm204_CPS',
'm202_CPS',]]
test_df2.columns = new_col
test_df2 = test_df2[cut_col]
result = pd.concat([test_df1, test_df2], axis=1)
#Calculating OPZ
result['OPZ_238'] = result.apply(lambda x: x['m238'] - x['m238_CPS'], axis=1)
result['OPZ_232'] = result.apply(lambda x: x['m232'] - x['m232_CPS'], axis=1)
result['OPZ_208'] = result.apply(lambda x: x['m208'] - x['m208_CPS'], axis=1)
result['OPZ_207'] = result.apply(lambda x: x['m207'] - x['m207_CPS'], axis=1)
result['OPZ_206'] = result.apply(lambda x: x['m206'] - x['m206_CPS'], axis=1)
result['OPZ_204'] = result.apply(lambda x: x['m204'] - x['m204_CPS'], axis=1)
result['OPZ_202'] = result.apply(lambda x: x['m202'] - x['m202_CPS'], axis=1)
#Calculating Ratios
result['206/238'] = result.apply(lambda x: x['206_CPS']/x['238_CPS'], axis=1)
result['208/232'] = result.apply(lambda x: x['208_CPS']/x['232_CPS'], axis=1)
result['207/206'] = result.apply(lambda x: x['207_CPS']/x['206_CPS'], axis=1)
result['208/206'] = result.apply(lambda x: x['208_CPS']/x['206_CPS'], axis=1)
result['206/204'] = result.apply(lambda x: x['206_CPS']/x['204_CPS'], axis=1)
result['208/204'] = result.apply(lambda x: x['208_CPS']/x['204_CPS'], axis=1)
result['207/204'] = result.apply(lambda x: x['207_CPS']/x['204_CPS'], axis=1)
calc_dict[key] = result
return calc_dict
def ranked_minimization(sheet, ratio, reject_percentage = 20):
mytest = tester[sheet].copy(deep=True)
df_mean_before = mytest[ratio].mean()
df_1std_before = mytest[ratio].std()
df_count_before = mytest[ratio].count()
df_2se_perc_before = (2 * mytest[ratio].sem()) / df_mean_before * 100
dif_mean = ratio + '_dif_from_mean'
dif_1SD = ratio + '_dif_from_1SD'
mytest[dif_mean] = mytest.apply(lambda x: abs(x[ratio] - df_mean_before), axis=1)
mytest[dif_1SD] = mytest.apply(lambda x: x[dif_mean] - df_1std_before, axis=1)
mytest2 = mytest.sort_values(by = dif_1SD, ascending = False)
#mytest2.head()
ratios_to_reject = int(mytest[ratio].count() * reject_percentage / 100)
#print(ratios_to_reject)
after_rejection = mytest2[ratios_to_reject:]
df_mean_after = after_rejection[ratio].mean()
df_1std_after = after_rejection[ratio].std()
df_count_after = after_rejection[ratio].count()
df_2se_perc_after = (2 * after_rejection[ratio].sem()) / df_mean_after * 100
# print(df_mean_after)
# print(df_1std_after)
# print(df_2se_perc_after)
results_dict = {}
results_dict['avg_before'] = df_mean_before
results_dict['1sd_before'] = df_1std_before
results_dict['2se%_before'] = df_2se_perc_before
results_dict['avg_after'] = df_mean_after
results_dict['1sd_after'] = df_1std_after
results_dict['2se%_after'] = df_2se_perc_after
return results_dict
def statistics_NP2(calc_dict):
calc_list = ['238_CPS', '232_CPS',
'208_CPS', '207_CPS', '206_CPS', '204_CPS', '202_CPS', '206/238',
'208/232', '207/206', '208/206', '206/204','208/204','207/204' ]
mega_dict = {}
for sheet in calc_dict:
tester = calc_dict[sheet]
stats_dict = {}
for col in tester:
if col in calc_list:
#print(col)
if '/' in col:
key = col + '_before rejection'
else:
key = col + '_mean'
df_mean = tester[col].mean()
stats_dict[key] = df_mean
df_precision = (2 * tester[col].sem()) / df_mean * 100
stats_dict[col + '_se%'] = df_precision
if 'OPZ' in col:
stats_dict[col + '_mean'] = tester[col].mean()
stats_dict['Time (s)'] = tester['Elapsed Time'].max()
#new_string = sheet.replace('time series data', '')
new_string = sheet.split('time')[0].rstrip()
mega_dict[new_string] = stats_dict
df_1 = pd.DataFrame(mega_dict)
df_flip = pd.DataFrame.transpose(df_1)
return df_flip
def statistics_ranktest(calc_dict):
calc_list = ['238_CPS', '232_CPS',
'208_CPS', '207_CPS', '206_CPS', '204_CPS', '202_CPS', '206/238',
'208/232', '207/206', '208/206', '206/204','208/204','207/204' ]
mega_dict = {}
for sheet in calc_dict:
tester = calc_dict[sheet]
stats_dict = {}
for col in tester:
if col in calc_list:
#print(col)
if '/' in col:
key_bf = col + '_before rejection'
key_af = col + '_after rejection'
key_bf_se = col + '_before rejection 2se%'
key_af_se = col + '_after rejection 2se%'
ranked_dict = ranked_minimization(sheet, col)
stats_dict[key_bf] = ranked_dict['avg_before']
stats_dict[key_bf_se] = ranked_dict['2se%_before']
stats_dict[key_af] = ranked_dict['avg_after']
stats_dict[key_af_se] = ranked_dict['2se%_after']
else:
key = col + '_mean'
df_mean = tester[col].mean()
stats_dict[key] = df_mean
df_precision = (2 * tester[col].sem()) / df_mean * 100
stats_dict[col + '_se%'] = df_precision
if 'OPZ' in col:
stats_dict[col + '_mean'] = tester[col].mean()
stats_dict['Time (s)'] = tester['Elapsed Time'].max()
#new_string = sheet.replace('time series data', '')
new_string = sheet.split('time')[0].rstrip()
mega_dict[new_string] = stats_dict
df_1 = pd.DataFrame(mega_dict)
df_flip = pd.DataFrame.transpose(df_1)
return df_flip
def files_ranked_toEXCEL(calc_dict, excel_name):
stats = statistics_ranktest(calc_dict)
with pd.ExcelWriter(excel_name) as writer:
for sheet in calc_dict:
calc_dict[sheet].to_excel(writer, sheet_name = sheet, index = False)
stats.to_excel(writer, sheet_name = 'Statistics', index = True)
new_filename = str(excel_name.split('.')[0]) + '_statistics.xlsx'
with pd.ExcelWriter(new_filename) as writer:
stats.to_excel(writer, sheet_name = 'Statistics', index = True)
def files_process_toEXCEL(calc_dict, excel_name):
with pd.ExcelWriter(excel_name) as writer:
for sheet in calc_dict:
calc_dict[sheet].to_excel(writer, sheet_name = sheet, index = False)
statistics_NP2(calc_dict).to_excel(writer, sheet_name = 'Statistics', index = True)
def file_process_combine(filename):
calc_dict = calc_CPS(read_np2_timeseries(filename))
new_filename = str(filename.split('.')[0]) + '_processed.xlsx'
files_process_toEXCEL(calc_dict, new_filename)
# +
#Functons for graphing and report generation
def U_Pb_plots(calc_dict, sample, choice = True):
key_list = ['238_CPS', '232_CPS',
'208_CPS', '207_CPS', '206_CPS', '204_CPS', '202_CPS', '206/238',
'208/232', '207/206', '208/206', '206/204']
zet = calc_dict[sample]
new_string = sample.split('time')[0].rstrip()
y_list = []
for key in key_list:
y_list.append(zet[key])
x = zet['Elapsed Time']
fig, axs = plt.subplots(4, 3, sharex = True, figsize = (12, 12))
fig.suptitle(new_string, fontsize=24)
ax_list = [
axs[0, 0],
axs[0, 1],
axs[0, 2],
axs[1, 0],
axs[1, 1],
axs[1, 2],
axs[2, 0],
axs[2, 1],
axs[2, 2],
axs[3, 0],
axs[3, 1],
axs[3, 2]
]
axs[0, 0].plot(x, y_list[0])
axs[0, 1].plot(x, y_list[1])
axs[0, 2].plot(x, y_list[2])
axs[1, 0].plot(x, y_list[3])
axs[1, 1].plot(x, y_list[4])
axs[1, 2].plot(x, y_list[5])
axs[2, 0].plot(x, y_list[6])
axs[2, 1].plot(x, y_list[7])
axs[2, 2].plot(x, y_list[8])
axs[3, 0].plot(x, y_list[9])
axs[3, 0].set(xlabel = 'Time (s)')
axs[3, 1].plot(x, y_list[10])
axs[3, 1].set(xlabel = 'Time (s)')
axs[3, 2].plot(x, y_list[11])
axs[3, 2].set(xlabel = 'Time (s)')
for idx in range(len(ax_list)):
ax_list[idx].ticklabel_format(axis='y', style='sci', scilimits=(0,0))
ax_list[idx].set_title(key_list[idx])
y_mean = [np.mean(y_list[idx])]*len(x)
# Plot the average line
mean_line = ax_list[idx].plot(x,y_mean, label='Mean', linestyle='--', color = "black")
# Make a legend
legend = ax_list[idx].legend(loc='upper right')
MYDIR = ("Figures")
CHECK_FOLDER = os.path.isdir(MYDIR)
# If folder doesn't exist, then create it.
if not CHECK_FOLDER:
os.makedirs(MYDIR)
#print("created folder : ", MYDIR)
#new_string = sample.replace('time series data', '').rstrip()
filename = os.path.join(MYDIR, new_string + '.pdf')
plt.savefig(filename)
print('Plot for ', new_string, " is complete.")
if choice == False:
plt.close()
#else:
#plt.close()
def U_Pb_report(calc_dict, intro_filename, intro = False, output_name = 'U-Pb_output.pdf'):
MYDIR = ("Figures")
mergedObject = PdfFileMerger()
if intro:
mergedObject.append(PdfFileReader(intro_filename, 'rb'))
print(f'Succesfully incorporated {intro_filename} into PDF.')
pd.set_option('precision', 2)
stats = statistics_NP2(calc_dict)
stat_dict = {}
stat_dict['stat1'] = stats.iloc[:, 14:]
stat_dict['stat2'] = stats.iloc[:, :8]
stat_dict['stat3'] = stats.iloc[:, 8:14]
html_list = []
for key in stat_dict:
name = key + ".pdf"
stats_html = stat_dict[key].to_html()
pdfkit.from_string(stats_html, name)
mergedObject.append(PdfFileReader(name, 'rb'))
file_list = []
keys = calc_dict.keys()
for key in keys:
#print(key)
U_Pb_plots(calc_dict, key, False)
new_string = key.split('time')[0].rstrip()
filename = os.path.join(MYDIR, new_string + '.pdf')
mergedObject.append(PdfFileReader(filename, 'rb'))
if '.pdf' in output_name:
pass
else:
output_name = output_name + '.pdf'
#output_name = "U-Pb_output.pdf"
mergedObject.write(output_name)
print(f'PDF file named: {output_name} is complete.')
# +
#Currently deprecated. Need to update to reflect calc_CPS()
def calc_CPS2(np2_dict):
''' Eliminates negative values in 238, 232, 208, 207, 206, and 204'''
columns = ['Absolute Time',
'Elapsed Time',
'm238_CPS',
'm232_CPS',
'm208_CPS',
'm207_CPS',
'm206_CPS',
'm204_CPS',
'm202_CPS']
new_col = ['Absolute Time',
'Elapsed Time',
'238_CPS',
'232_CPS',
'208_CPS',
'207_CPS',
'206_CPS',
'204_CPS',
'202_CPS']
cut_col = ['238_CPS',
'232_CPS',
'208_CPS',
'207_CPS',
'206_CPS',
'204_CPS',
'202_CPS']
key_list = ['<KEY>',
'm232_CPS',
'm208_CPS',
'm207_CPS',
'm206_CPS',
'm204_CPS']
calc_dict = {}
for key in np2_dict:
#print(key)
test_df1 = np2_dict[key]
test_df_new = test_df1
for item in key_list:
filter_mass = test_df_new[item] > 0
test_df_new = test_df_new[filter_mass]
#print(test_df_new.shape)
test_df1 = test_df_new
for col in columns:
test_df2 = test_df1.apply(lambda x: x * 62500000 if 'CPS' in x.name else x)
test_df2.columns = new_col
test_df2 = test_df2[cut_col]
result = pd.concat([test_df1, test_df2], axis=1)
#Calculating ratios
result['206/238'] = result.apply(lambda x: x['206_CPS']/x['238_CPS'], axis=1)
result['208/232'] = result.apply(lambda x: x['208_CPS']/x['232_CPS'], axis=1)
result['207/206'] = result.apply(lambda x: x['207_CPS']/x['206_CPS'], axis=1)
result['208/206'] = result.apply(lambda x: x['208_CPS']/x['206_CPS'], axis=1)
result['206/204'] = result.apply(lambda x: x['206_CPS']/x['204_CPS'], axis=1)
calc_dict[key] = result
return calc_dict
def file_process_combine2(filename):
calc_dict = calc_CPS2(read_np2_timeseries(filename))
new_filename = str(filename.split('.')[0]) + '_processed.xlsx'
files_process_toEXCEL(calc_dict, new_filename)
# -
filename = '06May_ttnSS2_NP2_baseline_corrected.xlsx'
test_df = read_np2_timeseries(filename)
test_df.keys()
test_df['SRM NIST 610 1.1']
tester = calc_CPS(test_df)
tester.keys()
# +
# excel_name = '06May_ttnSS2_NP2_processed.xlsx'
# files_ranked_toEXCEL(tester, excel_name)
# -
tester['SRM NIST 610 1.1']
statistics_NP2(tester).keys()
statistics_ranktest(tester)
# +
#INPUT
reject_percentage = 20
sheet = 'SRM NIST 610 1.1'
ratio = '208/232'
#END INPUT
ranked_minimization(sheet, ratio, reject_percentage)
# +
#filename = '0416_glass_SS_NPII_baseline.xlsx'
#calc_dict = calc_CPS(read_np2_timeseries(filename))
# +
# filename = '06May_ttnSS2_NP2_baseline_corrected.xlsx'
# SS_dict = calc_CPS(read_np2_timeseries(filename))
# excel_name = str(filename.split('.')[0]) + '_processed.xlsx'
# files_process_toEXCEL(SS_dict, excel_name)
# U_Pb_report(SS_dict, 'SS2_6May.pdf', True, 'Splitstream_06May2021_results')
# +
# filename = '0416_ttn_SS_NP2_data1.xlsx'
# ttn_dict = calc_CPS(read_np2_timeseries(filename))
# U_Pb_report(ttn_dict, 'titaniteTE.pdf',True, 'Titanite_splitstream_results')
# +
#U_Pb_report(calc_dict, 'glassTE.pdf',True, 'Glass_splitstream_results')
| NP_test5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3-azureml
# kernelspec:
# display_name: Python 3.8 - AzureML
# language: python
# name: python38-azureml
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# # Part 1: Training Tensorflow 2.0 Model on Azure Machine Learning Service
#
# ## Overview of the part 1
# This notebook is Part 1 (Preparing Data and Model Training) of a four part workshop that demonstrates an end-to-end workflow using Tensorflow 2.0 on Azure Machine Learning service. The different components of the workshop are as follows:
#
# - Part 1: [Preparing Data and Model Training](https://github.com/microsoft/bert-stack-overflow/blob/master/1-Training/AzureServiceClassifier_Training.ipynb)
# - Part 2: [Inferencing and Deploying a Model](https://github.com/microsoft/bert-stack-overflow/blob/master/2-Inferencing/AzureServiceClassifier_Inferencing.ipynb)
# - Part 3: [Setting Up a Pipeline Using MLOps](https://github.com/microsoft/bert-stack-overflow/tree/master/3-ML-Ops)
# - Part 4: [Explaining Your Model Interpretability](https://github.com/microsoft/bert-stack-overflow/blob/master/4-Interpretibility/IBMEmployeeAttritionClassifier_Interpretability.ipynb)
#
# **This notebook will cover the following topics:**
#
# - Stackoverflow question tagging problem
# - Introduction to Transformer and BERT deep learning models
# - Introduction to Azure Machine Learning service
# - Preparing raw data for training using Apache Spark
# - Registering cleaned up training data as a Dataset
# - Debugging the model in Tensorflow 2.0 Eager Mode
# - Training the model on GPU cluster
# - Monitoring training progress with built-in Tensorboard dashboard
# - Automated search of best hyper-parameters of the model
# - Registering the trained model for future deployment
# ## Prerequisites
# This notebook is designed to be run in Azure ML Notebook VM. See [readme](https://github.com/microsoft/bert-stack-overflow/blob/master/README.md) file for instructions on how to create Notebook VM and open this notebook in it.
# ### Check Azure Machine Learning Python SDK version
#
# This tutorial requires version 1.27.0 or higher. Let's check the version of the SDK:
# +
import azureml.core
print("Azure Machine Learning Python SDK version:", azureml.core.VERSION)
# -
# ## Stackoverflow Question Tagging Problem
# In this workshop we will use powerful language understanding model to automatically route Stackoverflow questions to the appropriate support team on the example of Azure services.
#
# One of the key tasks to ensuring long term success of any Azure service is actively responding to related posts in online forums such as Stackoverflow. In order to keep track of these posts, Microsoft relies on the associated tags to direct questions to the appropriate support team. While Stackoverflow has different tags for each Azure service (azure-web-app-service, azure-virtual-machine-service, etc), people often use the generic **azure** tag. This makes it hard for specific teams to track down issues related to their product and as a result, many questions get left unanswered.
#
# **In order to solve this problem, we will build a model to classify posts on Stackoverflow with the appropriate Azure service tag.**
#
# We will be using a BERT (Bidirectional Encoder Representations from Transformers) model which was published by researchers at Google AI Reasearch. Unlike prior language representation models, BERT is designed to pre-train deep bidirectional representations from unlabeled text by jointly conditioning on both left and right context in all layers. As a result, the pre-trained BERT model can be fine-tuned with just one additional output layer to create state-of-the-art models for a wide range of natural language processing (NLP) tasks without substantial architecture modifications.
#
# ## Why use BERT model?
# [Introduction of BERT model](https://arxiv.org/pdf/1810.04805.pdf) changed the world of NLP. Many NLP problems that before relied on specialized models to achive state of the art performance are now solved with BERT better and with more generic approach.
#
# If we look at the leaderboards on such popular NLP problems as GLUE and SQUAD, most of the top models are based on BERT:
# * [GLUE Benchmark Leaderboard](https://gluebenchmark.com/leaderboard/)
# * [SQuAD Benchmark Leaderboard](https://rajpurkar.github.io/SQuAD-explorer/)
#
# Recently, Allen Institue for AI announced new language understanding system called Aristo [https://allenai.org/aristo/](https://allenai.org/aristo/). The system has been developed for 20 years, but it's performance was stuck at 60% on 8th grade science test. The result jumped to 90% once researchers adopted BERT as core language understanding component. With BERT Aristo now solves the test with A grade.
# ## Quick Overview of How BERT model works
#
# The foundation of BERT model is Transformer model, which was introduced in [Attention Is All You Need paper](https://arxiv.org/abs/1706.03762). Before that event the dominant way of processing language was Recurrent Neural Networks (RNNs). Let's start our overview with RNNs.
#
# ## RNNs
#
# RNNs were powerful way of processing language due to their ability to memorize its previous state and perform sophisticated inference based on that.
#
# <img src="https://miro.medium.com/max/400/1*L38xfe59H5tAgvuIjKoWPg.png" alt="Drawing" style="width: 100px;"/>
#
# _Taken from [1](https://towardsdatascience.com/transformers-141e32e69591)_
#
# Applied to language translation task, the processing dynamics looked like this.
#
# 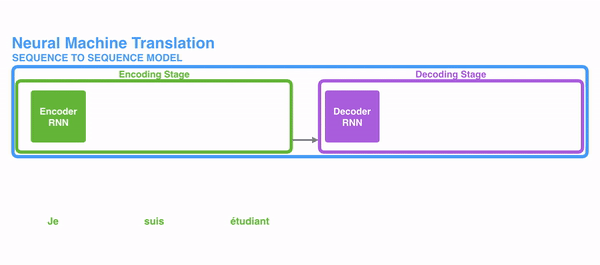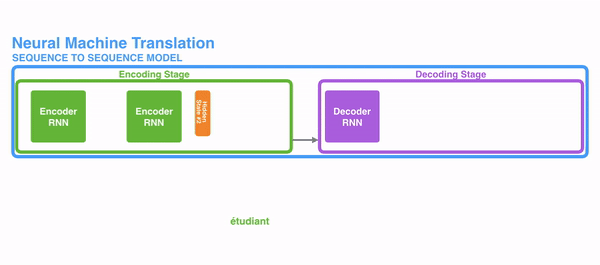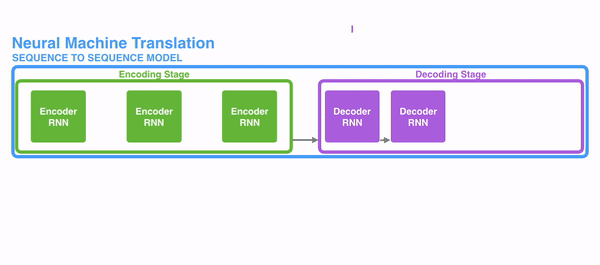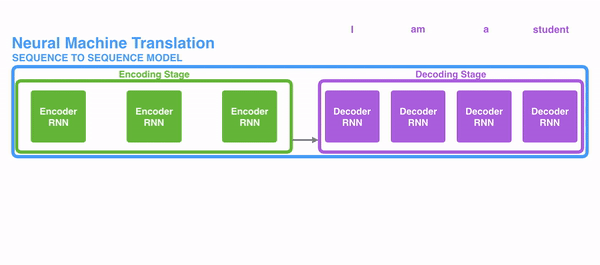
# _Taken from [2](https://jalammar.github.io/visualizing-neural-machine-translation-mechanics-of-seq2seq-models-with-attention/)_
#
# But RNNs suffered from 2 disadvantes:
# 1. Sequential computation put a limit on parallelization, which limited effectiveness of larger models.
# 2. Long term relationships between words were harder to detect.
# ## Transformers
#
# Transformers were designed to address these two limitations of RNNs.
#
# <img src="https://miro.medium.com/max/2436/1*V2435M1u0tiSOz4nRBfl4g.png" alt="Drawing" style="width: 500px;"/>
#
# _Taken from [3](http://jalammar.github.io/illustrated-transformer/)_
#
# In each Encoder layer Transformer performs Self-Attention operation which detects relationships between all word embeddings in one matrix multiplication operation.
#
# <img src="https://miro.medium.com/max/2176/1*fL8arkEFVKA3_A7VBgapKA.gif" alt="Drawing" style="width: 500px;"/>
#
# _Taken from [4](https://towardsdatascience.com/deconstructing-bert-part-2-visualizing-the-inner-workings-of-attention-60a16d86b5c1)_
#
# ## BERT Model
#
# BERT is a very large network with multiple layers of Transformers (12 for BERT-base, and 24 for BERT-large). The model is first pre-trained on large corpus of text data (WikiPedia + books) using un-superwised training (predicting masked words in a sentence). During pre-training the model absorbs significant level of language understanding.
#
# <img src="http://jalammar.github.io/images/bert-output-vector.png" alt="Drawing" style="width: 700px;"/>
#
# _Taken from [5](http://jalammar.github.io/illustrated-bert/)_
#
# Pre-trained network then can easily be fine-tuned to solve specific language task, like answering questions, or categorizing spam emails.
#
# <img src="http://jalammar.github.io/images/bert-classifier.png" alt="Drawing" style="width: 700px;"/>
#
# _Taken from [5](http://jalammar.github.io/illustrated-bert/)_
#
# The end-to-end training process of the stackoverflow question tagging model looks like this:
#
# 
#
# ## What is Azure Machine Learning Service?
# Azure Machine Learning service is a cloud service that you can use to develop and deploy machine learning models. Using Azure Machine Learning service, you can track your models as you build, train, deploy, and manage them, all at the broad scale that the cloud provides.
# 
#
#
# #### How can we use it for training machine learning models?
# Training machine learning models, particularly deep neural networks, is often a time- and compute-intensive task. Once you've finished writing your training script and running on a small subset of data on your local machine, you will likely want to scale up your workload.
#
# To facilitate training, the Azure Machine Learning Python SDK provides a high-level abstraction, the estimator class, which allows users to easily train their models in the Azure ecosystem. You can create and use an Estimator object to submit any training code you want to run on remote compute, whether it's a single-node run or distributed training across a GPU cluster.
# ## Connect To Workspace
#
# The [workspace](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.workspace(class)?view=azure-ml-py) is the top-level resource for Azure Machine Learning, providing a centralized place to work with all the artifacts you create when you use Azure Machine Learning. The workspace holds all your experiments, compute targets, models, datastores, etc.
#
# You can [open ml.azure.com](https://ml.azure.com) to access your workspace resources through a graphical user interface of **Azure Machine Learning studio**.
#
# 
#
# **You will be asked to login in the next step. Use your Microsoft AAD credentials.**
# +
from azureml.core import Workspace
ws = Workspace.from_config()
print('Workspace name: ' + ws.name,
'Azure region: ' + ws.location,
'Subscription id: ' + ws.subscription_id,
'Resource group: ' + ws.resource_group, sep = '\n')
# -
# ## Register Datastore
# A [Datastore](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.datastore.datastore?view=azure-ml-py) is used to store connection information to a central data storage. This allows you to access your storage without having to hard code this (potentially confidential) information into your scripts.
#
# In this tutorial, the data was been previously prepped and uploaded into a central [Blob Storage](https://azure.microsoft.com/en-us/services/storage/blobs/) container. We will register this container into our workspace as a datastore using a [shared access signature (SAS) token](https://docs.microsoft.com/en-us/azure/storage/common/storage-sas-overview).
#
#
# Here is files.
# 
# + gather={"logged": 1622006615321}
from azureml.core import Datastore, Dataset
datastore_name = 'mtcseattle'
container_name = 'azure-service-classifier'
account_name = 'mtcseattle'
sas_token = '?sv=2020-04-08&st=2021-05-26T04%3A39%3A46Z&se=2022-05-27T04%3A39%3A00Z&sr=c&sp=rl&sig=CTFMEu24bo2X06G%2B%2F2aKiiPZBzvlWHELe15rNFqULUk%3D'
datastore = Datastore.register_azure_blob_container(workspace=ws,
datastore_name=datastore_name,
container_name=container_name,
account_name=account_name,
sas_token=sas_token,
overwrite=True)
# -
# #### If the datastore has already been registered, then you (and other users in your workspace) can directly run this cell.
# + gather={"logged": 1622006619321}
from azureml.core import Datastore
datastore = Datastore.get(ws, 'mtcseattle')
datastore
# -
# ## Dataset
#
# Azure Machine Learning service supports first class notion of a Dataset. A [Dataset](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.dataset.dataset?view=azure-ml-py) is a resource for exploring, transforming and managing data in Azure Machine Learning. The following Dataset types are supported:
#
# * [TabularDataset](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.data.tabulardataset?view=azure-ml-py) represents data in a tabular format created by parsing the provided file or list of files.
#
# * [FileDataset](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.data.filedataset?view=azure-ml-py) references single or multiple files in datastores or from public URLs.
# ### Register Dataset using SDK
#
# We will register datasets using SDK. In this workshop we will register second type of Datasets using code - File Dataset. File Dataset allows specific folder in our datastore that contains our data files to be registered as a Dataset.
#
# There is a folder within our datastore called **data** that contains all our training and testing data. We will register this as a dataset.
# +
azure_dataset = Dataset.File.from_files(path=(datastore, 'data'))
azure_dataset = azure_dataset.register(workspace=ws,
name='Azure Services Dataset',
description='Dataset containing azure related posts on Stackoverflow',
create_new_version=True)
# -
# #### If the dataset has already been registered, then you (and other users in your workspace) can directly run this cell.
azure_dataset = Dataset.get_by_name(ws, 'Azure Services Dataset')
azure_dataset
# ## Perform Experiment using Azure Machine Learning for SDK
#
# Now that we have Compute Instance, dataset, and training script working locally, it is time to train a model running the script. We will start by creating an [experiment](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.experiment.experiment?view=azure-ml-py). An experiment is a grouping of many runs from a specified script. All runs in this tutorial will be performed under the same experiment.
# +
from azureml.core import Experiment
experiment_name = 'azure-service-classifier'
experiment = Experiment(ws, name=experiment_name)
# + gather={"logged": 1622006443581}
from azureml.core import Environment
from azureml.core.conda_dependencies import CondaDependencies
env = Environment.get(ws, name='AzureML-TensorFlow-2.0-GPU')
env.python.conda_dependencies.add_conda_package("pip")
env.python.conda_dependencies.add_pip_package("transformers==2.0.0")
env.python.conda_dependencies.add_pip_package("absl-py")
env.python.conda_dependencies.add_pip_package("azureml-dataprep")
env.python.conda_dependencies.add_pip_package("h5py<3.0.0")
env.python.conda_dependencies.add_pip_package("pandas")
env.name = "Bert_training"
env
# -
# ### Create ScriptRunConfig
#
# The RunConfiguration object encapsulates the information necessary to submit a training run in an experiment. Typically, you will not create a RunConfiguration object directly but get one from a method that returns it, such as the submit method of the Experiment class.
#
# RunConfiguration is a base environment configuration that is also used in other types of configuration steps that depend on what kind of run you are triggering. For example, when setting up a PythonScriptStep, you can access the step's RunConfiguration object and configure Conda dependencies or access the environment properties for the run.
# Let's go over how a Run is executed in Azure Machine Learning.
#
# 
# A quick description for each of the parameters we have just defined:
#
# - `source_directory`: This specifies the root directory of our source code.
# - `entry_script`: This specifies the training script to run. It should be relative to the source_directory.
# - `compute_target`: This specifies to compute target to run the job on. We will use the one created earlier.
# - `script_params`: This specifies the input parameters to the training script. Please note:
#
# 1) *azure_dataset.as_named_input('azureservicedata').as_mount()* mounts the dataset to the remote compute and provides the path to the dataset on our datastore.
#
# 2) All outputs from the training script must be outputted to an './outputs' directory as this is the only directory that will be saved to the run.
# #### Add Metrics Logging
#
# So we were able to clone a Tensorflow 2.0 project and run it without any changes. However, with larger scale projects we would want to log some metrics in order to make it easier to monitor the performance of our model.
#
# We can do this by adding a few lines of code into our training script:
#
# ```python
# # 1) Import SDK Run object
# from azureml.core.run import Run
#
# # 2) Get current service context
# run = Run.get_context()
#
# # 3) Log the metrics that we want
# run.log('val_accuracy', float(logs.get('val_accuracy')))
# run.log('accuracy', float(logs.get('accuracy')))
# ```
# We've created a *train_logging.py* script that includes logging metrics as shown above.
# %pycat train_logging.py
# + gather={"logged": 1622006507878}
from azureml.core import ScriptRun, ScriptRunConfig
scriptrun = ScriptRunConfig(source_directory='./',
script='train_logging.py',
arguments=['--data_dir', azure_dataset.as_named_input('azureservicedata').as_mount(),
'--max_seq_length', 128,
'--batch_size', 16,
'--learning_rate', 3e-5,
'--steps_per_epoch', 5, # to reduce time for workshop
'--num_epochs', 1, # to reduce time for workshop
'--export_dir','./outputs/exports'],
compute_target=locals,
environment=env)
run = experiment.submit(scriptrun)
# -
# Now if we view the current details of the run, you will notice that the metrics will be logged into graphs.
# + gather={"logged": 1622006698331}
from azureml.widgets import RunDetails
RunDetails(run).show()
# -
# ## Check the model performance
#
# Last training run produced model of decent accuracy. Let's test it out and see what it does. First, let's check what files our latest training run produced and download the model files.
#
# #### Download model files
run.get_file_names()
# +
modelpath = 'outputs/exports'
run.download_files(prefix=modelpath)
# # If you haven't finished training the model then just download pre-made model from datastore
# datastore.download('./outputs/exports',prefix='model')
# -
# ## Register Model
#
# A registered [model](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.model(class)?view=azure-ml-py) is a reference to the directory or file that make up your model. After registering a model, you and other people in your workspace can easily gain access to and deploy your model without having to run the training script again.
#
# We need to define the following parameters to register a model:
#
# - `model_name`: The name for your model. If the model name already exists in the workspace, it will create a new version for the model.
# - `model_path`: The path to where the model is stored. In our case, this was the *export_dir* defined in our estimators.
# - `description`: A description for the model.
#
# Let's register the best run from our hyperparameter tuning.
model = run.register_model(model_name='azure-service-classifier',
model_path=modelpath,
datasets=[('train, test, validation data', azure_dataset)],
description='BERT model for classifying azure services on stackoverflow posts.')
# ---
# ## Appendix
#
# __Followings are not for this workshop.__
# ## Distributed Training Across Multiple GPUs
#
# Distributed training allows us to train across multiple nodes if your cluster allows it. Azure Machine Learning service helps manage the infrastructure for training distributed jobs. All we have to do is add the following parameters to our estimator object in order to enable this:
#
# - `node_count`: The number of nodes to run this job across. Our cluster has a maximum node limit of 2, so we can set this number up to 2.
# - `process_count_per_node`: The number of processes to enable per node. The nodes in our cluster have 2 GPUs each. We will set this value to 2 which will allow us to distribute the load on both GPUs. Using multi-GPUs nodes is benefitial as communication channel bandwidth on local machine is higher.
# - `distributed_training`: The backend to use for our distributed job. We will be using an MPI (Message Passing Interface) backend which is used by Horovod framework.
#
# We use [Horovod](https://github.com/horovod/horovod), which is a framework that allows us to easily modifying our existing training script to be run across multiple nodes/GPUs. The distributed training script is saved as *train_horovod.py*.
#
# * **ACTION**: Explore _train_horovod.py_ using [Azure ML studio > Notebooks tab](images/azuremlstudio-notebooks-explore.png)
# %pycat train_horovod.py
# +
from azureml.core.compute import AmlCompute, ComputeTarget
cluster_name = 'train-gpu-nv6'
compute_config = AmlCompute.provisioning_configuration(vm_size='Standard_NV6',
idle_seconds_before_scaledown=6000,
min_nodes=0,
max_nodes=10)
compute_target = ComputeTarget.create(ws, cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
# -
# We can submit this run in the same way that we did with the others, but with the additional parameters.
# +
from azureml.core import ScriptRun, ScriptRunConfig
from azureml.core.runconfig import MpiConfiguration
scriptrun3 = ScriptRunConfig(source_directory='./',
script='train_horovod.py',
arguments=['--data_dir', azure_dataset.as_named_input('azureservicedata').as_mount(),
'--max_seq_length', 128,
'--batch_size', 32,
'--learning_rate', 3e-5,
'--steps_per_epoch', 150,
'--num_epochs', 3,
'--export_dir','./outputs/exports'],
compute_target=compute_target,
distributed_job_config=MpiConfiguration(process_count_per_node=1, node_count=1),
environment=env)
run3 = experiment.submit(scriptrun3)
# -
# Once again, we can view the current details of the run.
from azureml.widgets import RunDetails
RunDetails(run3).show()
# Once the run completes note the time it took. It should be around 5 minutes. As you can see, by moving to the cloud GPUs and using distibuted training we managed to reduce training time of our model from more than an hour to 5 minutes. This greatly improves speed of experimentation and innovation.
# ## Tune Hyperparameters Using Hyperdrive
#
# So far we have been putting in default hyperparameter values, but in practice we would need tune these values to optimize the performance. Azure Machine Learning service provides many methods for tuning hyperparameters using different strategies.
#
# The first step is to choose the parameter space that we want to search. We have a few choices to make here :
#
# - **Parameter Sampling Method**: This is how we select the combinations of parameters to sample. Azure Machine Learning service offers [RandomParameterSampling](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.hyperdrive.randomparametersampling?view=azure-ml-py), [GridParameterSampling](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.hyperdrive.gridparametersampling?view=azure-ml-py), and [BayesianParameterSampling](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.hyperdrive.bayesianparametersampling?view=azure-ml-py). We will use the `GridParameterSampling` method.
# - **Parameters To Search**: We will be searching for optimal combinations of `learning_rate` and `num_epochs`.
# - **Parameter Expressions**: This defines the [functions that can be used to describe a hyperparameter search space](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.hyperdrive.parameter_expressions?view=azure-ml-py), which can be discrete or continuous. We will be using a `discrete set of choices`.
#
# The following code allows us to define these options.
# +
from azureml.train.hyperdrive import GridParameterSampling
from azureml.train.hyperdrive.parameter_expressions import choice
param_sampling = GridParameterSampling( {
'--learning_rate': choice(3e-5, 3e-4),
'--num_epochs': choice(3, 4)
}
)
# -
# The next step is to a define how we want to measure our performance. We do so by specifying two classes:
#
# - **[PrimaryMetricGoal](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.hyperdrive.primarymetricgoal?view=azure-ml-py)**: We want to `MAXIMIZE` the `val_accuracy` that is logged in our training script.
# - **[BanditPolicy](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.hyperdrive.banditpolicy?view=azure-ml-py)**: A policy for early termination so that jobs which don't show promising results will stop automatically.
# +
from azureml.train.hyperdrive import BanditPolicy
from azureml.train.hyperdrive import PrimaryMetricGoal
primary_metric_name='val_accuracy'
primary_metric_goal=PrimaryMetricGoal.MAXIMIZE
early_termination_policy = BanditPolicy(slack_factor = 0.1, evaluation_interval=1, delay_evaluation=2)
# -
# We define an estimator as usual, but this time without the script parameters that we are planning to search.
# +
from azureml.core import ScriptRun, ScriptRunConfig
scriptrun4 = ScriptRunConfig(source_directory='./',
script='train_logging.py',
arguments=['--data_dir', azure_dataset.as_named_input('azureservicedata').as_mount(),
'--max_seq_length', 128,
'--batch_size', 32,
'--learning_rate', 3e-5,
'--steps_per_epoch', 150,
'--num_epochs', 3,
'--export_dir','./outputs/exports'],
compute_target=compute_target,
environment=env)
# -
# Finally, we add all our parameters in a [HyperDriveConfig](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.hyperdrive.hyperdriveconfig?view=azure-ml-py) class and submit it as a run.
# +
from azureml.train.hyperdrive import HyperDriveConfig
hyperdrive_run_config = HyperDriveConfig(run_config=scriptrun4,
hyperparameter_sampling=param_sampling,
policy=early_termination_policy,
primary_metric_name=primary_metric_name,
primary_metric_goal=PrimaryMetricGoal.MAXIMIZE,
max_total_runs=10,
max_concurrent_runs=2)
run4 = experiment.submit(hyperdrive_run_config)
# -
# When we view the details of our run this time, we will see information and metrics for every run in our hyperparameter tuning.
from azureml.widgets import RunDetails
RunDetails(run4).show()
# We can retrieve the best run based on our defined metric.
best_run = run4.get_best_run_by_primary_metric()
# We have registered the model with Dataset reference.
# * **ACTION**: Check dataset to model link in **Azure ML studio > Datasets tab > Azure Service Dataset**.
# In the [next tutorial](), we will perform inferencing on this model and deploy it to a web service.
| 1-Training/AzureServiceClassifier_Training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # `keras-unet-collection.models` user guide
#
# This user guide is proposed for `keras-unet-collection == 0.0.15`
import tensorflow as tf
from tensorflow import keras
print('TensorFlow {}; Keras {}'.format(tf.__version__, keras.__version__))
# ### Step 1: importing `models` from `keras_unet_collection`
from keras_unet_collection import models
# ### Step 2: defining your hyper-parameters
# Commonly used hyper-parameter options are listed as follows. Full details are available through the Python helper function:
#
# * `inpust_size`: a tuple or list that defines the shape of input tensors.
# * `models.resunet_a_2d` and other models that configured with `'PReLU'` support int only, others support NoneType.
#
#
# * `filter_num`: a list that defines the number of convolutional filters per down- and up-sampling blocks.
# * For `unet_2d`, `att_unet_2d`, `unet_plus_2d`, `r2_unet_2d`, depth $\ge$ 2 is expected.
# * For `resunet_a_2d` and `u2net_2d`, depth $\ge$ 3 is expected.
#
#
# * `n_labels`: number of output targets, e.g., `n_labels=2` for binary classification.
#
# * `activation`: the activation function of hidden layers. Available choices are `'ReLU'`, `'LeakyReLU'`, `'PReLU'`, `'ELU'`, `'GELU'`, `'Snake'`.
#
# * `output_activation`: the activation function of the output layer. Recommended choices are `'Sigmoid'`, `'Softmax'`, `None` (linear), `'Snake'`.
#
# * `batch_norm`: if specified as True, all convolutional layers will be configured as stacks of "Conv2D-BN-Activation".
#
# * `stack_num_down`: number of convolutional layers per downsampling level.
#
# * `stack_num_up`: number of convolutional layers (after concatenation) per upsampling level.
#
# * `pool`: the configuration of downsampling (encoding) blocks.
# * `pool=False`: downsampling with a convolutional layer (2-by-2 convolution kernels with 2 strides; optional batch normalization and activation).
# * `pool=True` or `pool='max'` downsampling with a max-pooling layer.
# * `pool='ave'` downsampling with a average-pooling layer.
#
#
# * `unpool`: the configuration of upsampling (decoding) blocks.
# * `unpool=False`: upsampling with a transpose convolutional layer (2-by-2 convolution kernels with 2 strides; optional batch normalization and activation).
# * `unpool=True` or `unpool='bilinear'` upsampling with bilinear interpolation.
# * `unpool='nearest'` upsampling with reflective padding.
#
#
# * `name`: user-specified prefix of the configured layer and model. Use `keras.models.Model.summary` to identify the exact name of each layer.
# ### Step 3: Configuring your model (examples are provided)
# **Example 1**: U-net for binary classification with:
#
# 1. Five down- and upsampliung levels (or four downsampling levels and one bottom level).
#
# 2. Two convolutional layers per downsampling level.
#
# 3. One convolutional layer (after concatenation) per upsamling level.
#
# 2. Gaussian Error Linear Unit (GELU) activcation, Softmax output activation, batch normalization.
#
# 3. Downsampling through Maxpooling.
#
# 4. Upsampling through reflective padding.
model = models.unet_2d((None, None, 3), [64, 128, 256, 512, 1024], n_labels=2,
stack_num_down=2, stack_num_up=1,
activation='GELU', output_activation='Softmax',
batch_norm=True, pool='max', unpool='nearest', name='unet')
# **Example 2**: Vnet (originally proposed for 3-d inputs, here modified for 2-d inputs) for binary classification with:
#
# 1. Input size of (256, 256, 1); PReLU does not support input tensor with shapes of NoneType
#
#
#
# 1. Five down- and upsampliung levels (or four downsampling levels and one bottom level).
#
# 1. Number of stacked convolutional layers of the residual path increase with downsampling levels from one to three (symmetrically, decrease with upsampling levels).
# * `res_num_ini=1`
# * `res_num_max=3`
#
#
# 2. PReLU activcation, Softmax output activation, batch normalization.
#
# 3. Downsampling through stride convolutional layers.
#
# 4. Upsampling through transpose convolutional layers.
model = models.vnet_2d((256, 256, 1), filter_num=[16, 32, 64, 128, 256], n_labels=2,
res_num_ini=1, res_num_max=3,
activation='PReLU', output_activation='Softmax',
batch_norm=True, pool=False, unpool=False, name='vnet')
# **Example 3**: attention-Unet for single target regression with:
#
# 1. Four down- and upsampling levels.
#
# 2. Two convolutional layers per downsampling level.
#
# 3. Two convolutional layers (after concatenation) per upsampling level.
#
# 2. ReLU activation, linear output activation (None), batch normalization.
#
# 3. Additive attention, ReLU attention activation.
#
# 4. Downsampling through stride convolutional layers.
#
# 5. Upsampling through bilinear interpolation.
#
model = models.att_unet_2d((None, None, 3), [64, 128, 256, 512], n_labels=1,
stack_num_down=2, stack_num_up=2,
activation='ReLU', atten_activation='ReLU', attention='add', output_activation=None,
batch_norm=True, pool=False, unpool='bilinear', name='attunet')
# **Example 4**: U-net++ for three-label classification with:
#
# 1. Four down- and upsampling levels.
#
# 2. Two convolutional layers per downsampling level.
#
# 3. Two convolutional layers (after concatenation) per upsampling level.
#
# 2. LeakyReLU activation, Softmax output activation, no batch normalization.
#
# 3. Downsampling through Maxpooling.
#
# 4. Upsampling through transpose convolutional layers.
#
# 5. Deep supervision.
model = models.unet_plus_2d((None, None, 3), [64, 128, 256, 512], n_labels=3,
stack_num_down=2, stack_num_up=2,
activation='LeakyReLU', output_activation='Softmax',
batch_norm=False, pool='max', unpool=False, deep_supervision=True, name='xnet')
# **Example 5**: UNet 3+ for binary classification with:
#
# 1. Four down- and upsampling levels.
#
# 2. Two convolutional layers per downsampling level.
#
# 3. One convolutional layers (after concatenation) per upsampling level.
#
# 2. ReLU activation, Sigmoid output activation, batch normalization.
#
# 3. Downsampling through Maxpooling.
#
# 4. Upsampling through transpose convolutional layers.
#
# 5. Deep supervision.
model = models.unet_3plus_2d((128, 128, 3), n_labels=2, filter_num_down=[64, 128, 256, 512],
filter_num_skip='auto', filter_num_aggregate='auto',
stack_num_down=2, stack_num_up=1, activation='ReLU', output_activation='Sigmoid',
batch_norm=True, pool='max', unpool=False, deep_supervision=True, name='unet3plus')
# * `filter_num_skip` and `filter_num_aggregate` can be specified explicitly:
model = models.unet_3plus_2d((128, 128, 3), n_labels=2, filter_num_down=[64, 128, 256, 512],
filter_num_skip=[64, 64, 64], filter_num_aggregate=256,
stack_num_down=2, stack_num_up=1, activation='ReLU', output_activation='Sigmoid',
batch_norm=True, pool='max', unpool=False, deep_supervision=True, name='unet3plus')
# **Example 6**: R2U-net for binary classification with:
#
# 1. Four down- and upsampling levels.
#
# 2. Two recurrent convolutional layers with two iterations per down- and upsampling level.
#
# 2. ReLU activation, Softmax output activation, no batch normalization.
#
# 3. Downsampling through Maxpooling.
#
# 4. Upsampling through reflective padding.
model = models.r2_unet_2d((None, None, 3), [64, 128, 256, 512], n_labels=2,
stack_num_down=2, stack_num_up=1, recur_num=2,
activation='ReLU', output_activation='Softmax',
batch_norm=True, pool='max', unpool='nearest', name='r2unet')
# **Example 7**: ResUnet-a for 16-label classification with:
#
# 1. input size of (128, 128, 3)
#
# 1. Six downsampling levels followed by an Atrous Spatial Pyramid Pooling (ASPP) layer with 256 filters.
#
# 1. Six upsampling levels followed by an ASPP layer with 128 filters.
#
# 2. dilation rates of {1, 3, 15, 31} for shallow layers, {1,3,15} for intermediate layers, and {1,} for deep layers.
#
# 3. ReLU activation, Sigmoid output activation, batch normalization.
#
# 4. Downsampling through stride convolutional layers.
#
# 4. Upsampling through reflective padding.
model = models.resunet_a_2d((128, 128, 3), [32, 64, 128, 256, 512, 1024],
dilation_num=[1, 3, 15, 31],
n_labels=16, aspp_num_down=256, aspp_num_up=128,
activation='ReLU', output_activation='Sigmoid',
batch_norm=True, pool=False, unpool='nearest', name='resunet')
# * `dilation_num` can be specified per down- and uplampling level:
model = models.resunet_a_2d((128, 128, 3), [32, 64, 128, 256, 512, 1024],
dilation_num=[[1, 3, 15, 31], [1, 3, 15, 31], [1, 3, 15], [1, 3, 15], [1,], [1,],],
n_labels=16, aspp_num_down=256, aspp_num_up=128,
activation='ReLU', output_activation='Sigmoid',
batch_norm=True, pool=False, unpool='nearest', name='resunet')
# **Example 8**: U^2-Net for binary classification with:
#
# 1. Six downsampling levels with the first four layers built with RSU, and the last two (one downsampling layer, one bottom layer) built with RSU-F4.
# * `filter_num_down=[64, 128, 256, 512]`
# * `filter_mid_num_down=[32, 32, 64, 128]`
# * `filter_4f_num=[512, 512]`
# * `filter_4f_mid_num=[256, 256]`
#
#
# 1. Six upsampling levels with the deepest layer built with RSU-F4, and the other four layers built with RSU.
# * `filter_num_up=[64, 64, 128, 256]`
# * `filter_mid_num_up=[16, 32, 64, 128]`
#
#
# 3. ReLU activation, Sigmoid output activation, batch normalization.
#
# 4. Deep supervision
#
# 5. Downsampling through stride convolutional layers.
#
# 6. Upsampling through transpose convolutional layers.
#
# *In the original work of U^2-Net, down- and upsampling were achieved through maxpooling (`pool=True` or `pool='max'`) and bilinear interpolation (`unpool=True` or unpool=`'bilinear'`).
u2net = models.u2net_2d((128, 128, 3), n_labels=2,
filter_num_down=[64, 128, 256, 512], filter_num_up=[64, 64, 128, 256],
filter_mid_num_down=[32, 32, 64, 128], filter_mid_num_up=[16, 32, 64, 128],
filter_4f_num=[512, 512], filter_4f_mid_num=[256, 256],
activation='ReLU', output_activation=None,
batch_norm=True, pool=False, unpool=False, deep_supervision=True, name='u2net')
# * `u2net_2d` supports automated determination of filter numbers per down- and upsampling level. Auto-mode may produce a slightly larger network.
u2net = models.u2net_2d((None, None, 3), n_labels=2,
filter_num_down=[64, 128, 256, 512],
activation='ReLU', output_activation='Sigmoid',
batch_norm=True, pool=False, unpool=False, deep_supervision=True, name='u2net')
| examples/user_guid_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/probml/pyprobml/blob/master/book1/supplements/colab_intro.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="24_OboyL7tqe"
# # Introduction to colab
#
# Colab is Google's version of Jupyter notebooks, but has the following advantages:
# - it runs in the cloud, not locally, so you can use it from a cheap laptop, such as a Chromebook.
# - The notebook is saved in your Google drive, so you can share your notebook with someone else and work on it collaboratively.
# - it has nearly all of the packages you need for doing ML pre-installed
# - it gives you free access to GPUs
# - it has a [file editor](https://colab.research.google.com/github/probml/pyprobml/blob/master/book1/supplements/colab_intro.ipynb#scrollTo=DdXlYCe1AlJa&line=7&uniqifier=1), so you can separate your code from the output of your code, as with other IDEs, such as [Jupyter lab](https://jupyterlab.readthedocs.io/en/stable/).
# - it has various other useful features, such as collapsible sections (cf. code folding), and ways to specify parameters to your functions via [various GUI widgets](https://colab.research.google.com/notebooks/forms.ipynb) for use by non-programmers. (You can automatically execute parameterized notebooks with different parameters using [papermill](https://papermill.readthedocs.io/en/latest/).)
#
# More details can be found in the [official introduction](https://colab.research.google.com/notebooks/intro.ipynb). Below we describe a few more tips and tricks, focusing on methods that I have found useful when developing the book. (More advanced tricks can be found in [this blog post](https://amitness.com/2020/06/google-colaboratory-tips/) and [this blog post](https://medium.com/@robertbracco1/configuring-google-colab-like-a-pro-d61c253f7573#4cf4).)
#
# + colab={"base_uri": "https://localhost:8080/"} id="ZjFsGQJ41k32" outputId="ba36c760-3d87-4b96-ea62-0fe44d308d95"
IS_COLAB = ('google.colab' in str(get_ipython()))
print(IS_COLAB)
# + id="B4KQOCig_xf1"
# Standard Python libraries
from __future__ import absolute_import, division, print_function, unicode_literals
import os
import time
import glob
from typing import Any, Callable, Dict, Iterator, Mapping, Optional, Sequence, Tuple
# + [markdown] id="8lAbDqny-vDq"
# # How to import and use standard libraries
# + [markdown] id="XHO2_uKXMbD4"
# Colab comes with most of the packages we need pre-installed.
# You can see them all using this command.
#
#
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="0C54AJx40vJq" outputId="accb047b-6ac3-4e3c-97c5-6f3e1ccf7823"
# !pip list -v
# + [markdown] id="U9PghW_NT1HY"
# To install a new package called 'foo', use the following (see [this page](https://colab.research.google.com/notebooks/snippets/importing_libraries.ipynb) for details):
#
# ```
# # # !pip install foo
# ```
#
#
# + [markdown] id="fOBdg02-_Jws"
# ## Numpy
# + id="AzP2LAtN_L1m" colab={"base_uri": "https://localhost:8080/"} outputId="775ab507-3a70-4816-d76e-e2d463aa571d"
import numpy as np
np.set_printoptions(precision=3)
A = np.random.randn(2,3)
print(A)
# + [markdown] id="76jPgsuk_1IP"
# ## Pandas
# + id="GimloDqo_4No" colab={"base_uri": "https://localhost:8080/", "height": 197} outputId="1ddca722-d652-4b44-c4e0-982c722dabeb"
import pandas as pd
pd.set_option('precision', 2) # 2 decimal places
pd.set_option('display.max_rows', 20)
pd.set_option('display.max_columns', 30)
pd.set_option('display.width', 100) # wide windows
url = 'https://archive.ics.uci.edu/ml/machine-learning-databases/auto-mpg/auto-mpg.data'
column_names = ['MPG','Cylinders','Displacement','Horsepower','Weight',
'Acceleration', 'Year', 'Origin', 'Name']
df = pd.read_csv(url, names=column_names, sep='\s+', na_values="?")
df.head()
# + [markdown] id="hUCC261x_7zZ"
# ## Sklearn
# + id="RCSwx_lE_7Jn" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="4036c988-bb39-4ad4-d9af-f45bc6330231"
import sklearn
from sklearn.datasets import load_iris
iris = load_iris()
# Extract numpy arrays
X = iris.data
y = iris.target
import matplotlib.pyplot as plt
plt.scatter(X[:,0], X[:,1])
# + [markdown] id="PJXF4csdBhsN"
# ## JAX
# + id="8JiSxcJJ79Bv" colab={"base_uri": "https://localhost:8080/"} outputId="97f3ebb7-93b8-4255-bccb-781d01ccb4cc"
# JAX (https://github.com/google/jax)
import jax
import jax.numpy as jnp
A = jnp.zeros((3,3))
# Check if JAX is using GPU
print("jax backend {}".format(jax.lib.xla_bridge.get_backend().platform))
# + [markdown] id="l99YLyorBdYE"
# ## Tensorflow
# + colab={"base_uri": "https://localhost:8080/"} id="StpReaSICLUm" outputId="5dfac104-8e5a-49cd-8400-eadb56650acd"
import tensorflow as tf
from tensorflow import keras
assert tf.__version__ >= "2.0"
print("tf version {}".format(tf.__version__))
print([d for d in tf.config.list_physical_devices()])
if not tf.config.list_physical_devices('GPU'):
print("No GPU was detected. DNNs can be very slow without a GPU.")
if IS_COLAB:
print("Go to Runtime > Change runtime and select a GPU hardware accelerator.")
# + [markdown] id="grUUK1GrBfIY"
# ## PyTorch
# + id="Oi4Zmzla73A_" colab={"base_uri": "https://localhost:8080/"} outputId="558ed73a-5dfe-491a-be1c-64c<PASSWORD>e48"
import torch
import torchvision
print("torch version {}".format(torch.__version__))
if torch.cuda.is_available():
print(torch.cuda.get_device_name(0))
else:
print("Torch cannot find GPU")
# + [markdown] id="M4ayVFuc0FD9"
# # Plotting
#
# Colab has excellent support for plotting. We give some examples below.
# + [markdown] id="ENCT0EqifCDO"
# ## Static plots
#
# Colab lets you make static plots using matplotlib, as shown below.
# Note that plots are displayed inline by default, so
# ```
# # # %matplotlib inline
# ```
# is not needed.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 267} id="j_k4tv4D1VaC" outputId="dda6cc16-8351-4aff-9c09-ab7ad5dc968f"
import matplotlib.pyplot as plt
import PIL
import imageio
import seaborn as sns;
seaborn.set()
seaborn.set_style("whitegrid")
plt.rcParams.update({'font.size': 48, 'legend.fontsize': 18})
plt.rc('xtick',labelsize=14)
plt.rc('ytick',labelsize=14)
from IPython.display import display
plt.figure()
plt.plot(range(10))
plt.savefig('myplot.png')
# + [markdown] id="iFwo8LA4fIh9"
# ## Interactive plots
#
# Colab also lets you create interactive plots using various javascript libraries - see [here](https://colab.research.google.com/notebooks/charts.ipynb#scrollTo=QSMmdrrVLZ-N) for details.
#
# Below we illustrate how to use the [bokeh library](https://docs.bokeh.org/en/latest/index.html) to create an interactive plot of a pandas time series, where if you mouse over the plot, it shows the corresponding (x,y) coordinates. (Another option is [plotly](https://plotly.com/graphing-libraries/).)
# + id="SiKRxZyo3Fa1"
import pandas as pd
import numpy as np
from bokeh.plotting import figure, show
from bokeh.io import output_notebook
from bokeh.models import ColumnDataSource, HoverTool
# Call once to configure Bokeh to display plots inline in the notebook.
output_notebook()
# + id="7_4CJ3mQhLNa" colab={"base_uri": "https://localhost:8080/", "height": 616} outputId="88eb0172-352f-46bd-8303-1baf898e0a79"
np.random.seed(0)
dates = pd.date_range(start='2018-04-24', end='2018-08-27')
N = len(dates)
vals = np.random.standard_t(1, size=N)
dd = pd.DataFrame({'vals': vals, 'dates': dates}, index=dates)
dd['days'] = dd.dates.dt.strftime("%Y-%m-%d")
source = ColumnDataSource(dd)
hover = HoverTool(tooltips=[("Date", "@days"),
("vals", "@vals")],
)
p = figure( x_axis_type="datetime")
p.line(x='dates', y='vals', source=source)
p.add_tools(hover)
show(p)
# + [markdown] id="do80zEzsil6O"
# When you mouse over a point, you should see something like the screenshot below.
#
#
# <img height="500" src="
# https://github.com/probml/pyprobml/blob/master/book1/intro/figures/bokeh-timeseries.png?raw=true">
#
# + [markdown] id="nHGH3o_R28G9"
# We can also make plots that can you pan and zoom into.
# + id="q1jy_cQl3AYc" colab={"base_uri": "https://localhost:8080/", "height": 616} outputId="ae8fc2f4-a7e5-48e9-f8f0-f9fd32331ded"
N = 4000
np.random.seed(0)
x = np.random.random(size=N) * 100
y = np.random.random(size=N) * 100
radii = np.random.random(size=N) * 1.5
colors = ["#%02x%02x%02x" % (r, g, 150) for r, g in zip(np.floor(50+2*x).astype(int), np.floor(30+2*y).astype(int))]
p = figure()
p.circle(x, y, radius=radii, fill_color=colors, fill_alpha=0.6, line_color=None)
show(p)
# + [markdown] id="qLh3fxl63IHW"
# ## Graphviz
#
# You can use graphviz to layout nodes of a graph and draw the structure.
# + colab={"base_uri": "https://localhost:8080/"} id="33v9WeNjfb2k" outputId="2567027c-8c11-4517-94b8-3ae984b1d328"
# !apt-get -y install python-pydot
# !apt-get -y install python-pydot-ng
# !apt-get -y install graphviz
# + colab={"base_uri": "https://localhost:8080/", "height": 498} id="ZXTgGVkaffjf" outputId="2037bb22-88c1-4013-cf8e-03c8ebe349d5"
from graphviz import Digraph
dot = Digraph(comment='Bayes net')
print(dot)
dot.node('C', 'Cloudy')
dot.node('R', 'Rain')
dot.node('S', 'Sprinkler')
dot.node('W', 'Wet grass')
dot.edge('C', 'R')
dot.edge('C', 'S')
dot.edge('R', 'W')
dot.edge('S', 'W')
print(dot.source)
dot.render('test-output/graph.jpg', view=True)
dot
# + [markdown] id="DaSY50JzpWnC"
# ## Progress bar
#
# + colab={"base_uri": "https://localhost:8080/"} id="CqcNHigZpYPO" outputId="b924fe64-137a-4342-e829-9d39d5fb52c8"
# ! pip install fastprogress
from fastprogress import master_bar, progress_bar
# + colab={"base_uri": "https://localhost:8080/", "height": 37} id="kN9YYBcBpfQy" outputId="a14d72a5-2b4c-44aa-9ccd-26a9d0a41dbf"
my_list = np.arange(0,100)
N = 1000
for item in progress_bar(my_list):
np.dot(np.ones((N,N)), np.ones((N,N)))
# + [markdown] id="lyWwltzKlAHS"
# # Filing system issues
#
# + [markdown] id="7aCgO-moU2WA"
# ## Accessing local files
#
# Clicking on the file folder icon on the left hand side of colab lets you browse local files. Right clicking on a filename lets you download it to your local machine. Double clicking on a file will open it in the file viewer/ editor, which appears on the right hand side.
#
# The result should look something like this:
#
# <img src="https://github.com/probml/pyprobml/blob/
# master/book1/intro/figures/colab-image-viewer.png?raw=true">
#
#
# You can also use standard unix commands to manipulate files, as we show below.
# + colab={"base_uri": "https://localhost:8080/"} id="dRh4BOIxHpEX" outputId="104c8f4f-9eff-4b9b-fc81-c084740e475b"
# !pwd
# + colab={"base_uri": "https://localhost:8080/"} id="T7i8bvaghwy7" outputId="bb0012d2-bc82-4d5d-dfe3-acf2ba3e77a7"
# !ls
# + colab={"base_uri": "https://localhost:8080/"} id="HDNijfMPjPsE" outputId="d2bb4d90-86fc-44b2-9900-6050507e082b"
# !echo 'foo bar' > foo.txt
# !cat foo.txt
# + [markdown] id="Ell60Ff4UE-P"
# You can open text files in the editor by clicking on their filename in the file browser, or programmatically as shown below.
# + colab={"base_uri": "https://localhost:8080/", "height": 16} id="CcxDCpyYjhfU" outputId="d521b728-16e6-43d7-f8e9-2c2d3c0b8eef"
from google.colab import files
files.view('foo.txt')
# + [markdown] id="PMet3XdcVF9O"
# If you make changes to a file containing code, the new version of the file will not be noticed unless you use the magic below.
# + colab={"base_uri": "https://localhost:8080/"} id="0ufY8AO1VEUh" outputId="13aeca9f-992b-4881-98d4-89508b0ad23a"
# %load_ext autoreload
# %autoreload 2
# + [markdown] id="6a6nkLsKWQpu"
# ## Syncing with Google drive
#
# Files that you generate in, or upload to, colab are ephemeral, since colab is a temporary environment with an idle timeout of 90 minutes and an absolute timeout of 12 hours (24 hours for Colab pro). To save any files permanently, you need to mount your google drive folder as we show below. (Executing this command will open a new window in your browser - you need cut and paste the password that is shown into the prompt box.)
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="cYZpcMiQkl15" outputId="f4cf4b76-100a-428f-8c63-14aad3207235"
from google.colab import drive
drive.mount('/content/gdrive')
# !pwd
# + id="wycWRaVxPh5P" colab={"base_uri": "https://localhost:8080/"} outputId="5571799c-360b-446f-9f70-375f7d14527b"
with open('/content/gdrive/MyDrive/foo.txt', 'w') as f:
f.write('Hello Google Drive!')
# !cat /content/gdrive/MyDrive/foo.txt
# + [markdown] id="CeGz5P_RUyzG"
# To ensure that local changes are detected by colab, use this piece of magic.
# + id="K6RR9dfoUyMG"
# %load_ext autoreload
# %autoreload 2
# + [markdown] id="bIqJtkFnlF8I"
# ## Downloading data
#
# You can use [wget](https://www.pair.com/support/kb/paircloud-downloading-files-with-wget/)
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="LjaBDSSQltez" outputId="b57a7fe9-3545-43ad-d9fc-<PASSWORD>"
# !pwd
# + id="3oypH8Vclu86"
# !rm timemachine.*
# + colab={"base_uri": "https://localhost:8080/"} id="H0dvvsUclgdu" outputId="d13d67f5-861d-4cab-a856-a719a4d2d503"
# #!wget https://github.com/probml/pyprobml/blob/master/data/timemachine.txt
# !wget https://raw.githubusercontent.com/probml/pyprobml/master/data/timemachine.txt
# + colab={"base_uri": "https://localhost:8080/"} id="kj3fMYuwlyNR" outputId="632cf687-d2d9-4e75-da37-07b096736521"
# !head timemachine.txt
# + [markdown] id="57S7dQXbPSh6"
# ## Viewing all your notebooks
#
# You can see the list of colab notebooks that you have saved as shown below.
# + id="cTJGGK29PYM4" colab={"base_uri": "https://localhost:8080/"} outputId="dbb73578-2dd8-4f3c-f270-4dfbdedb60ff"
import re, pathlib, shutil
from pathlib import PosixPath
# Get a list of all your Notebooks
notebooks = [x for x in pathlib.Path("/content/gdrive/MyDrive/Colab Notebooks").iterdir() if
re.search(r"\.ipynb", x.name, flags = re.I)]
print(notebooks[:2])
#n = PosixPath('/content/gdrive/MyDrive/Colab Notebooks/covid-open-data-paper.ipynb')
# + [markdown] id="buZsxpmUS37n"
# # Working with github
#
# You can open any jupyter notebook stored in github in a colab by replacing
# https://github.com/probml/.../intro.ipynb with https://colab.research.google.com/github/probml/.../intro.ipynb (see [this blog post](https://amitness.com/2020/06/google-colaboratory-tips/#6-open-notebooks-from-github).
#
# It is possible to download code (or data) from githib into a local directory on this virtual machine. It is also possible to upload local files back to github, although that is more complex. See details below.
# + [markdown] id="rVvGT6GUBg2Q"
# ## Cloning a repo from github
#
# You can clone a public github repo into your local colab VM, as we show below,
# using the repo for this book as an example.
# (To clone a private repo, you need to specify your password,
# as explained [here](https://medium.com/@robertbracco1/configuring-google-colab-like-a-pro-d61c253f7573#6b70). Alternatively you can use the ssh method we describe below.)
# + id="uVZWqzdW7_ZG" colab={"base_uri": "https://localhost:8080/"} outputId="69d00890-ba7e-4c24-9295-63580ac06e67"
# !rm -rf pyprobml # Remove any old local directory to ensure fresh install
# !git clone https://github.com/probml/pyprobml
# + id="sL0CLHTm7HSH" colab={"base_uri": "https://localhost:8080/"} outputId="bee2bf7f-a31d-4a97-fd63-ce65bc8629e0"
# !pwd
# + colab={"base_uri": "https://localhost:8080/"} id="XdC34HzKT8L8" outputId="7370c426-dc03-471a-f284-7905905faddf"
# !ls
# + [markdown] id="HuplkSqtkDuR"
# We can access data as shown below.
# + colab={"base_uri": "https://localhost:8080/"} id="_8m9VQ-3kFcq" outputId="4f078099-482d-4f09-c1b1-a950cd703d33"
datadir = 'pyprobml/data'
import re
fname = os.path.join(datadir, 'timemachine.txt')
with open(fname, 'r') as f:
lines = f.readlines()
sentences = [re.sub('[^A-Za-z]+', ' ', st).lower().split()
for st in lines]
for i in range(5):
words = sentences[i]
print(words)
# + [markdown] id="MNWWINngc5rn"
# We can run any script as shown below.
# (Note we first have to define the environment variable for where the figures will be stored.)
# + colab={"base_uri": "https://localhost:8080/", "height": 828} id="aYXkQP-DdApw" outputId="d04325f6-7aa0-428a-c56e-bfc8ac2c7ca6"
import os
os.environ['PYPROBML']='pyprobml'
# %run pyprobml/scripts/activation_fun_plot.py
# + id="qItn37RW7R3N" colab={"base_uri": "https://localhost:8080/"} outputId="fb8d5ba1-7a9d-48ce-e0d2-ef429aa453d2"
# !ls pyprobml/figures
# + [markdown] id="YTT5eJ_qUDFe"
# We can also import code, as we show below.
# + colab={"base_uri": "https://localhost:8080/"} id="0jUdrHWLd95C" outputId="a6ba93bc-118c-42e7-c894-34eeef80c95b"
# !ls
# + colab={"base_uri": "https://localhost:8080/"} id="-NkBCWPdePFj" outputId="b33684a4-cd62-46fb-ab15-ef650f12d5c2"
import pyprobml.scripts.pyprobml_utils as pml
pml.test()
# + [markdown] id="yaISmcnNmnS7"
# ## Pushing local files back to github
#
# You can easily save your entire colab notebook to github by choosing 'Save a copy in github' under the File menu in the top left. But if you want to save individual files (eg code that you edited in the colab file editor, or a bunch of images or data files you created), the process is more complex.
#
# There are two main methods. You can either specify your username and password every time, as explained [here](https://medium.com/@robertbracco1/configuring-google-colab-like-a-pro-d61c253f7573#6b70). Or you can authenticate via ssh. We explain the latter method here.
#
# You first need to do some setup to create SSH keys on your current colab VM (virtual machine), manually add the keys to your github account, and then copy the keys to your mounted google drive so you can reuse the same keys in the future. This only has to be done once.
#
# After setup, you can use the `git_ssh` function we define below to securely execute git commands. This works by copying your SSH keys from your google drive to the current colab VM, executing the git command, and then deleting the keys from the VM for safety.
#
# + [markdown] id="0gOzFmcKoUuO"
# ### Setup
#
# Follow these steps. (These instructions are text, not code, since they require user interaction.)
#
# ```
# # # !ssh-keygen -t rsa -b 4096
# # # !ssh-keyscan -t rsa github.com >> ~/.ssh/known_hosts
# # # !cat /root/.ssh/id_rsa.pub
# ```
# The cat command will display your public key in the colab window.
# Cut and paste this and manually add to your github account following [these instructions](https://github.com/settings/keys).
#
# Test it worked
# ```
# # # !ssh -T <EMAIL>@github.com
# ```
#
# Finally, save the generated keys to your Google drive
#
# ```
# from google.colab import drive
# drive.mount('/content/drive')
# # # !mkdir /content/drive/MyDrive/ssh/
# # # !cp -r ~/.ssh/* /content/drive/MyDrive/ssh/
# # # !ls /content/drive/MyDrive/ssh/
# ```
#
# + [markdown] id="oSiVyBG1xm44"
# ### Test previous setup
#
# Let us check that we can see our SSH keys in our mounted google drive.
# + colab={"base_uri": "https://localhost:8080/"} id="cCUxHiHAxcY2" outputId="0cab8327-3769-4bca-e831-a8c6cce09888"
from google.colab import drive
drive.mount('/content/drive')
# !ls /content/drive/MyDrive/ssh/
# + [markdown] id="C-lPchgDpD7t"
# ### Executing git commands from colab via SSH
#
# The following function lets you securely doing a git command via SSH.
# It copies the keys from your google drive to the local VM, excecutes the command, then removes the keys.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="ONJ0Ump4wEho" outputId="5a0ca00a-dca7-49c9-ffba-339c1a3b521a"
# !rm -rf probml_tools.py # remove any old copies of this file
# !wget https://raw.githubusercontent.com/probml/pyprobml/master/scripts/pyprobml_utils.py
# + [markdown] id="umvOwzMfvpmU"
# Below we show how to use this. We first clone the repo to this colab VM.
# + colab={"base_uri": "https://localhost:8080/"} id="IbX9-PDpwlO0" outputId="957ae2fd-cab7-4267-8f4d-9709d6554910"
from google.colab import drive
drive.mount('/content/drive') # must do this before running git_colab
import pyprobml_utils as pml
# !rm -rf pyprobml # remove any old copies of this directory
# #!git clone https://github.com/probml/pyprobml.git # clones using wrong credentials
pml.git_ssh("git clone https://github.com/probml/pyprobml.git") # clone using your credentials
# + [markdown] id="ft_pJJ4ZTLdl"
# Next we add a file and push it to github.
#
# + colab={"base_uri": "https://localhost:8080/"} id="3hfluZVYTSNd" outputId="5da089c4-2234-46ff-d175-ffab8c96c67f"
# !pwd
# !ls
# To add stuff to github, you must be inside the git directory
# %cd /content/pyprobml
# !echo 'this is a test' > scripts/foo.txt
pml.git_ssh("git add scripts; git commit -m 'push from colab'; git push")
# %cd /content
# + [markdown] id="DAqqAhzzTS7F"
# [Here](https://github.com/probml/pyprobml/blob/master/scripts/foo.txt) is a link to the file we just pushed.
#
# Finally we clean up our mess.
# + colab={"base_uri": "https://localhost:8080/"} id="r16zYiNTu_Mz" outputId="3a6e5fe4-2eb2-46da-f834-9d14fbdd6c2d"
# %cd /content/pyprobml
pml.git_ssh("git rm scripts/foo*.txt; git commit -m 'colab cleanup'; git push")
# %cd /content
# + [markdown] id="q-kRtmdm5d7X"
# # Software engineering tools
#
# [<NAME> has argued](https://docs.google.com/presentation/d/1n2RlMdmv1p25Xy5thJUhkKGvjtV-dkAIsUXP-AL4ffI/edit) that notebooks are bad for developing complex software, because they encourage creating monolithic notebooks instead of factoring out code into separate, well-tested files.
#
# [<NAME> has responded to Joel's critiques here](https://www.youtube.com/watch?v=9Q6sLbz37gk&feature=youtu.be). In particular, the FastAI organization has created [nbdev](https://github.com/fastai/nbdev) which has various tools that make notebooks more useful.
#
#
# + [markdown] id="PuSsmj_fZ106"
# ## Avoiding problems with global state
#
# One of the main drawbacks of colab is that all variables are globally visible, so you may accidently write a function that depends on the current state of the notebook, but which is not passed in as an argument. Such a function may fail if used in a different context.
#
# One solution to this is to put most of your code in files, and then have the notebook simply import the code and run it, like you would from the command line. Then you can always run the notebook from scratch, to ensure consistency.
#
# Another solution is to use the [localscope](https://localscope.readthedocs.io/en/latest/README.html) package can catch some of these errors.
#
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="Q0FmEeIgc0YI" outputId="0d71c4a8-a4fb-45bd-e137-c391e408c83f"
# !pip install localscope
# + id="9zfUiUB8d-jh"
from localscope import localscope
# + colab={"base_uri": "https://localhost:8080/"} id="wI5wXzUPdlOS" outputId="500dfc1f-1cbc-4fa6-a30e-02f32258fa31"
a = 'hello world'
def myfun():
print(a) # silently accesses global variable
myfun()
# + colab={"base_uri": "https://localhost:8080/", "height": 337} id="T3V1iV32czq8" outputId="b346cb7a-f252-41bc-992b-dd72e5aec825"
a = 'hello world'
@localscope
def myfun():
print(a)
myfun()
# + colab={"base_uri": "https://localhost:8080/", "height": 337} id="5t48_AMbAN8V" outputId="71fdaacf-4650-4a52-9fb5-da5873c33e73"
def myfun2():
return 42
@localscope
def myfun3():
return myfun2()
# + colab={"base_uri": "https://localhost:8080/"} id="DAqLZ8PdAquy" outputId="021e8282-a283-414c-ef1d-46521da89e49"
@localscope.mfc # allow for global methods, functions, classes
def myfun4():
return myfun2()
myfun4()
# + [markdown] id="iZaeVouoAhXP"
# ## Factoring out functionality into files stored on github
#
# The recommended workflow is to develop your code in the colab in the usual way, and when it is working, to factor out the core code into separate files. You can edit these files locally in the colab editor, and then push the code to github when ready (see details above). To run functions defined in a local file, just import them. For example, suppose we have created the file /content/pyprobml/scripts/fit_flax.py; we can use this idiom to run its test suite:
# ```
# import pyprobml.scripts.fit_flax as ff
# ff.test()
# ```
# If you make local edits, you want to be sure
# that you always import the latest version of the file (not a cached version). So you need to use this piece of colab magic first:
# ```
# # # %load_ext autoreload
# # # %autoreload 2
# ```
#
# + [markdown] id="DdXlYCe1AlJa"
# ## File editors
#
# Colab has a simple file editor, illustrated below for an example file.
# This lets you separate your code from the output of your code, as with other IDEs, such as [Jupyter lab](https://jupyterlab.readthedocs.io/en/stable/).
#
#
# <img src="https://github.com/probml/pyprobml/blob/
# master/book1/intro/figures/colab-file-editor.png?raw=true">
#
# + [markdown] id="LgLmSpaHBxLu"
# You can click on a class name when holding Ctrl and the source code will open in the file viewer. (h/t [Amit Choudhary's blog](https://amitness.com/2020/06/google-colaboratory-tips/).
#
# <img src="https://github.com/probml/pyprobml/blob/master/images/colab-goto-class.gif?raw=true">
# + [markdown] id="4qi2xKMbAnlj"
#
# ## VScode
# The default colab file editor is very primitive.
# See [this article](https://amitness.com/vscode-on-colab/) for how to run VScode
# from inside your Colab browser. Unfortunately this is a bit slow. It is also possible to run VScode locally on your laptop, and have it connect to colab via SSH, but this is more complex (see [this blog post](https://amitness.com/vscode-on-colab/) or [this medium post](https://medium.com/@robertbracco1/configuring-google-colab-like-a-pro-d61c253f7573#4cf4) for details).
#
#
# + [markdown] id="7KrRcbQ71ZyZ"
# # Hardware accelerators
#
# By default, Colab runs on a CPU, but you can select GPU or TPU for extra speed, as we show below. To get access to more powerful machines (with faster processors, more memory, and longer idle timeouts), you can subscript to [Colab Pro](https://colab.research.google.com/signup). At the time of writing (Jan 2021), the cost is $10/month (USD). This is a good deal if you use GPUs a lot.
#
# <img src="https://github.com/probml/pyprobml/blob/master/images/colab-pro-spec-2020.png?raw=true" height=300>
#
#
# + [markdown] id="Qpb9N-c-3RSf"
# ## CPUs
#
# To see what devices you have, use this command.
#
# + colab={"base_uri": "https://localhost:8080/"} id="SZYIR1Kk1ktp" outputId="04ab4999-74d4-47d6-f9cb-004ca13b590b"
from tensorflow.python.client import device_lib
device_lib.list_local_devices()
# + colab={"base_uri": "https://localhost:8080/"} id="hs_l1zUY3UuQ" outputId="99a7db7f-6325-452e-ca69-ab7439070d67"
# !cat /proc/version
# + colab={"base_uri": "https://localhost:8080/"} id="WHG13Iwa3Z5k" outputId="ebc74028-163e-48e2-a0c6-56667e4e0c0d"
from psutil import cpu_count
print('num cores', cpu_count())
# !cat /proc/cpuinfo
# + [markdown] id="PivaBI5za45p"
# ## Memory
#
# + colab={"base_uri": "https://localhost:8080/"} id="xg3C0Yrq3job" outputId="0b480191-8489-4b6c-fc26-c6f3b51aecc2"
from psutil import virtual_memory
ram_gb = virtual_memory().total / 1e9
print('RAM (GB)', ram_gb)
# + colab={"base_uri": "https://localhost:8080/"} id="W_I7m_hUbBpu" outputId="b688a7d0-1c53-4864-c79f-fde42cda4db6"
# !cat /proc/meminfo
# + [markdown] id="v0G2d13kIEz5"
# ## GPUs
#
# If you select the 'Runtime' menu at top left, and then select 'Change runtime type' and then select 'GPU', you can get free access to a GPU.
#
#
# <img src="https://github.com/probml/pyprobml/blob/
# master/book1/intro/figures/colab-change-runtime.png?raw=true" height=300>
# <img src="https://github.com/probml/pyprobml/blob/
# master/book1/intro/figures/colab-select-gpu.png?raw=true" height=200>
#
#
# + [markdown] id="MGVZB0esI0QG"
#
# To see what kind of GPU you are using, see below.
#
# + id="FikkXWQqBU9O" colab={"base_uri": "https://localhost:8080/"} outputId="655ec017-3552-4390-bd7e-21e48ebc4ea9"
# gpu_info = !nvidia-smi
gpu_info = '\n'.join(gpu_info)
print(gpu_info)
# + colab={"base_uri": "https://localhost:8080/"} id="GU6nII1F5S2S" outputId="d220a1a6-cc83-4718-c77e-2df38f13bf9c"
# !grep Model: /proc/driver/nvidia/gpus/*/information | awk '{$1="";print$0}'
# + [markdown] id="8hYLvG1KgM6K"
# # Using Julia in colab
#
# See [this colab](https://colab.research.google.com/github/ageron/julia_notebooks/blob/master/Julia_Colab_Notebook_Template.ipynb)
# + id="hzi1OpMaZlAc"
| book1/supplements/colab_intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Test ploting of categorical values
#
# <NAME> 20 feb 2019
# This notebook serves to test wether plotting of categorical values works as intended
#import sys
#sys.path.append('/home/sigurd/dev/ProcessOptimizer')
import numpy as np
from sklearn.datasets import load_breast_cancer
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import cross_val_score
from ProcessOptimizer.space import Integer, Categorical
from ProcessOptimizer import plots, gp_minimize
from ProcessOptimizer.plots import plot_objective
# For reproducibility
import numpy as np
np.random.seed(123)
import matplotlib.pyplot as plt
plt.set_cmap("viridis")
SPACE = [
Integer(1, 20, name='max_depth'),
Integer(2, 100, name='min_samples_split'),
Integer(5, 30, name='min_samples_leaf'),
Integer(1, 30, name='max_features'),
Categorical(list('abc'), name='dummy'),
Categorical(['gini', 'entropy'], name='criterion'),
Categorical(list('def'), name='dummy'),
]
def objective(params):
clf = DecisionTreeClassifier(**{dim.name: val for dim, val in zip(SPACE, params) if dim.name != 'dummy'})
return -np.mean(cross_val_score(clf, *load_breast_cancer(True)))
result = gp_minimize(objective, SPACE, n_calls=20)
# # plot_objective
#
# Plot objective now supports optional use of partial dependence as well as different methods of defining parameter values for dependency plots
# Here we see an example of using partial dependence. Even when setting n_points all the way down to 10 from the default of 40, this method is still very slow. This is because partial dependence calculates 250 extra predictions for each point on the plots.
_ = plot_objective(result,usepartialdependence = True, n_points = 10)
# Here we plot without partial dependence. We see that it is a lot faster. Also the values for the other parameters are set to the default "result" which is the parameter set of the best observed value so far. In the case of funny_func this is close to 0 for all parameters.
_ = plot_objective(result,usepartialdependence = False, n_points = 10)
# Here we try with setting the other parameters to something other than "result". First we try with "expected_minimum" which is the set of parameters that gives the miniumum value of the surogate function, using scipys minimum search method.
_ = plot_objective(result,usepartialdependence = False, n_points = 10,pars = 'expected_minimum')
# "expected_minimum_random" is a naive way of finding the minimum of the surogate by only using random sampling:
_ = plot_objective(result,usepartialdependence = False, n_points = 10,pars = 'expected_minimum_random')
# Lastly we can also define these parameters ourselfs by parsing a list as the pars argument:
_ = plot_objective(result,usepartialdependence = False, n_points = 10,pars = [15, 4, 7, 15, 'b', 'entropy', 'e'])
# We can also specify how many intial samples are used for the two different "expected_minimum" methods. We set it to a low value in the next examples to showcase how it affects the minimum for the two methods.
_ = plot_objective(result,usepartialdependence = False, n_points = 10,pars = 'expected_minimum_random',expected_minimum_samples = 10)
_ = plot_objective(result,usepartialdependence = False, n_points = 10,pars = 'expected_minimum',expected_minimum_samples = 1)
| plot_test/plot_objective_categorical_values.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using regularized logistic regression to classify email
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import utils
from sklearn import linear_model
#import sklearn.cross_validation
from sklearn import model_selection
#from sklearn.cross_validation import KFold
import scipy.io
import warnings
warnings.filterwarnings("ignore", category=FutureWarning)
# This is a bit of magic to make matplotlib figures appear inline in the notebook
# rather than in a new window.
# %matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# Some more magic so that the notebook will reload external python modules;
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
# +
# No modifications in this cell
# complete the functions in utils.py; then run the cell
Xtrain,Xtest,ytrain,ytest = utils.load_spam_data()
# Preprocess the data
Xtrain_std,mu,sigma = utils.std_features(Xtrain)
Xtrain_logt = utils.log_features(Xtrain)
Xtrain_bin = utils.bin_features(Xtrain)
Xtest_std = (Xtest - mu)/sigma
Xtest_logt = utils.log_features(Xtest)
Xtest_bin = utils.bin_features(Xtest)
# find good lambda by cross validation for these three sets
def run_dataset(X,ytrain,Xt,ytest,typea,penalty):
best_lambda = utils.select_lambda_crossval(X,ytrain,0.1,5.1,0.5,penalty)
print("best_lambda = %.3f" %best_lambda)
# train a classifier on best_lambda and run it
if penalty == "l2":
lreg = linear_model.LogisticRegression(penalty=penalty,C=1.0/best_lambda, solver='lbfgs',fit_intercept=True,max_iter=1000)
else:
lreg = linear_model.LogisticRegression(penalty=penalty,C=1.0/best_lambda, solver='liblinear',fit_intercept=True,max_iter=1000)
lreg.fit(X,ytrain)
print("Coefficients = %s" %lreg.intercept_,lreg.coef_)
predy = lreg.predict(Xt)
print("Accuracy on set aside test set for %s = %.4f" %(typea, np.mean(predy==ytest)))
print("L2 Penalty experiments -----------")
run_dataset(Xtrain_std,ytrain,Xtest_std,ytest,"std","l2")
run_dataset(Xtrain_logt,ytrain,Xtest_logt,ytest,"logt","l2")
run_dataset(Xtrain_bin,ytrain,Xtest_bin,ytest,"bin","l2")
print("L1 Penalty experiments -----------")
run_dataset(Xtrain_std,ytrain,Xtest_std,ytest,"std","l1")
run_dataset(Xtrain_logt,ytrain,Xtest_logt,ytest,"logt","l1")
run_dataset(Xtrain_bin,ytrain,Xtest_bin,ytest,"bin","l1")
# -
| hw2/logreg/logreg_spam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
df= pd.read_csv("textclass.csv")
df
# -
y=df["Label"]
x=df["Title"]
y
# +
###############################################################################################
# the words need to be encoded as integers or floating point values for use as input to a ML algorithm
#################################################################################################
# Create bag of words
from sklearn.feature_extraction.text import CountVectorizer
count = CountVectorizer()
"""CountVectorizer : vector is returned with a length of the entire vocabulary
and an integer count for the number of
times each word appeared in the document."""
count.fit(x)
print(count.vocabulary_)
#{'love': 6, 'india': 4, 'is': 5, 'best': 1, 'germany': 3, 'beats': 0, 'both': 2}
# +
bag_of_words = count.fit_transform(x)
print(bag_of_words)
# +
# Create feature matrix
X = bag_of_words.toarray()
print(X)
"""
#{'love': 6, 'india': 4, 'is': 5, 'best': 1, 'germany': 3, 'beats': 0, 'both': 2}
# 7 columns because we have 7 seperate words in our dataset and 3 rows because we have 3 rows in dataset
[[0 0 0 0 2 0 1]
[0 1 0 0 1 1 0]
[1 0 1 1 0 0 0]]
"""
# +
# Create multinomial naive Bayes object with prior probabilities of each class
from sklearn.naive_bayes import MultinomialNB
clf = MultinomialNB(class_prior=[0.35, 0.60])
# Train model
model = clf.fit(X, y)
#xtest=np.array(["India is my country. I want to go germany, India"])
xtest=np.array(["love germany"])
xtest=count.transform(xtest)
print(xtest)
xtest=xtest.toarray()
xtest
# +
pred=clf.predict(xtest)
# Create new observation
#new_observation = [[0, 0, 0, 1, 0, 1, 0]]
#new_observation = [[0, 0, 0, 1, 1, 1, 1]]
#new_observation = [[0, 0, 1, 1, 1, 1, 1]]
# Predict new observation's class
#pred = model.predict(new_observation)
print(pred)
| Lec 15 Naive Bayes Questions/Ma'am/multinomail.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lets make some simple examples with scala
#
# We'll make a very simple scala object compile it and use it in the python process
# !mkdir helloscala
# +
# %%file helloscala/hw.scala
object HelloScala
{
def sayHi(): String = "Hi! from scala"
def sum(x: Int, y: Int) = x + y
}
# -
# ---
#
# We'll compile things with sbt (the scala build tool)
#
# This makes us a jar that we can load with spark.
#
# ---
# + language="bash"
# cd helloscala
# sbt package
# + slideshow={"slide_type": "slide"}
import spylon
import spylon.spark as sp
c = sp.SparkConfiguration()
c._spark_home = "/path/to/spark-1.6.2-bin-hadoop2.6"
c.master = ["local[4]"]
# -
# Add the jar we built previously so that we can import our scala stuff
c.jars = ["./helloscala/target/scala-2.10/helloscala_2.10-0.1-SNAPSHOT.jar"]
# + slideshow={"slide_type": "fragment"}
(sc, sqlContext) = c.sql_context("MyApplicationName")
# -
# Lets load our helpers and import the Scala class we just wrote
# + slideshow={"slide_type": "fragment"}
from spylon.spark.utils import SparkJVMHelpers
helpers = SparkJVMHelpers(sc)
# -
Hi = helpers.import_scala_object("HelloScala")
print Hi.__doc__
Hi.sayHi()
Hi.sum(4, 6)
| examples/02_SpylonExample-WithJar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Inhaltsverzeichnis<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Hilfsfunktionen-definieren" data-toc-modified-id="Hilfsfunktionen-definieren-1"><span class="toc-item-num">1 </span>Hilfsfunktionen definieren</a></span></li><li><span><a href="#Experimmente:-Anzahl-an-Schichten" data-toc-modified-id="Experimmente:-Anzahl-an-Schichten-2"><span class="toc-item-num">2 </span>Experimmente: Anzahl an Schichten</a></span></li></ul></div>
# -
# # Hilfsfunktionen definieren
# +
import numpy as np
import matplotlib.pyplot as plt
from tensorflow.keras import Sequential
from tensorflow.keras.layers import Dense, Activation, Flatten
from tensorflow.keras.layers import Conv2D, MaxPooling2D, ZeroPadding2D
from tensorflow.keras import backend as K
def baue_cnn_model(shape_of_input, nr_output_neurons, nr_layers, kernel_side_len):
"""
Baut ein komplettes CNN auf
"""
# clear the last Keras session
# this will clear the underlying TensorFlow graph
K.clear_session()
model = Sequential()
for layer_nr in range(0, nr_layers):
nr_filter = 32
kernel_stride = 1
if layer_nr==0:
model.add( Conv2D(nr_filter,
kernel_size=(kernel_side_len,kernel_side_len),
strides=(kernel_stride,kernel_stride),
activation='relu',
input_shape=shape_of_input) )
else:
model.add( Conv2D(nr_filter,
kernel_size=(kernel_side_len,kernel_side_len),
strides=(kernel_stride,kernel_stride),
activation='relu') )
model.add( MaxPooling2D(pool_size=(2, 2), strides=(2, 2)) )
model.add(Flatten())
model.add(Dense(1024, activation="relu"))
model.add(Dense(nr_output_neurons, activation='softmax'))
model.compile(optimizer='adam', loss='categorical_crossentropy')
model.build()
model.summary()
return model
def trainiere_cnn(das_cnn, bildds, anz_train_schritte):
"""
Trainiert das angegebene CNN mit dem angegebenen
Bilddatensatz für die angegebene Anzahl von
Trainingsschritten
"""
from datetime import datetime
time_start = datetime.now()
height = input_shape[0]
width = input_shape[1]
nr_channels = input_shape[2]
X = np.zeros( (1,height,width,nr_channels) )
Y = np.zeros( (1,bildds.nr_classes) )
alle_fehler = []
for train_schritt in range(0,anz_train_schritte):
if train_schritt % 200 == 0:
print("Trainingsschritt:", train_schritt)
img, img_processed, class_id, class_name, teacher_vec = \
bildds.hole_irgendein_bild()
# Stecke das 3D Bild in das 4D Array
# denn Keras will als Input für die
# Trainingsmethod fit() ein 4D Array
X[0,:,:,:] = img_processed
# Der gewünschte Output ist dementsprechend
# ein 2D Array
Y[0,:] = teacher_vec
# Trainiere das Modell mit diesem Bild
history = das_cnn.fit(X,Y,verbose=0)
fehler_img = history.history["loss"][0]
alle_fehler.append( fehler_img )
plt.plot(alle_fehler)
plt.show()
time_stop = datetime.now()
print("Training zu Ende:")
print("\tTraining-Start: ", time_start)
print("\tTraining-Stop: ", time_stop)
print("\tTrainings-Dauer: ", time_stop - time_start)
def teste_cnn(das_cnn, bildds, zeige_bsp_klassifikationen=False):
"""
Testet das angegebene CNN mit dem angegebenen Bilddatensatz
und zeigt evtl. Beispielklassifikationen an
"""
korrekt = 0
print("Ich werde das CNN auf {} Tesbildern testen".
format(bildds.nr_images))
for test_bild_nr in range(0, bildds.nr_images):
if test_bild_nr % 100 == 0:
print("{} Bilder getestet".format(test_bild_nr))
# 1. hole nächstes Testbild
img, img_processed, class_id, class_name, teacher_vec = \
bildds.hole_bild_per_index(test_bild_nr)
# 2. 3D Bild in 4D Array umwandeln
X = img_processed.reshape((-1,
img_processed.shape[0],
img_processed.shape[1],
img_processed.shape[2] ))
# 3. Klassifiziere jetzt!
neuron_outputs = das_cnn.predict(X)
# 4. Bestimme prädizierte Klasse
predicted_class_nr = np.argmax(neuron_outputs.reshape(-1))
# 5. Prädiktion korrekt?
if predicted_class_nr == class_id:
korrekt +=1
# 6.
# Show image, predicted class and gt class?
if zeige_bsp_klassifikationen and np.random.randint(5)==0:
predicted_class_label = \
bildds.class_names[predicted_class_nr]
plt.title("Ist: {} vs. Prädiziert: {}\nOutputneurone: {}"
.format(class_name,
predicted_class_label,
neuron_outputs
)
)
plt.imshow( img )
plt.show()
# Berechne Korrektklassifikationsrate
korrekt_rate = float(korrekt) / float(bildds.nr_images)
print("Korrekt klassifiziert: {} of {} images:"
" --> rate: {:.2f}"
.format(korrekt,
bildds.nr_images,
korrekt_rate)
)
return korrekt_rate
# -
# # Experimmente: Anzahl an Schichten
# +
# Vorbereiten der Trainings- und Testdaten
from bilddatensatz import bilddatensatz
img_size_heute = (100,100)
input_shape = (img_size_heute[0], img_size_heute[1], 3)
train_folder = r"/home/juebrauer/link_to_vcd/07_datasets/08_bikes_vs_cars/train"
bd_train = bilddatensatz( train_folder, img_size_heute )
test_folder = r"/home/juebrauer/link_to_vcd/07_datasets/08_bikes_vs_cars/test"
bd_test = bilddatensatz( test_folder, img_size_heute )
x = []
y = []
kernel_side_len = 3
for nr_layers in range(1,6):
print("\n")
print("--------------------------------------------")
print(f"Experiment: CNN mit {nr_layers} Schicht(en)")
print("--------------------------------------------")
# 1. CNN Modell bauen
mein_cnn = baue_cnn_model(input_shape, bd_train.nr_classes, nr_layers, kernel_side_len)
# 2. CNN trainieren
NR_TRAIN_STEPS = 30000
trainiere_cnn(mein_cnn, bd_train, NR_TRAIN_STEPS)
# 3. CNN testen
korrekt_klassifikations_rate = teste_cnn(mein_cnn, bd_test)
print("Korrekt_klassifikations_rate :",korrekt_klassifikations_rate)
# 4. Speichere Ergebnisse
x.append( nr_layers )
y.append( korrekt_klassifikations_rate )
# 5. Plotte Ergebnisse so weit
plt.scatter(x,y)
plt.xlabel("Anzahl Schichten")
plt.ylabel("Korrekt-Klassifikationsrate")
plt.show()
# -
| linux/cnn_experimente_mit_hyperparametern/cnn_experimente_mit_hyperparametern.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
get_ipython().magic('matplotlib inline')
sns.set(style='white', font_scale=0.9)
flatui = ["#9b59b6", "#3498db", "#95a5a6", "#e74c3c", "#34495e", "#2ecc71"]
sns.color_palette(flatui)
np.set_printoptions(threshold=np.nan)
pd.set_option("display.max_columns",100)
# -
dataset = pd.read_csv('HW2_pokemon.csv')
dataset.head(10)
dataset.info()
dataset['Type 2'] = dataset['Type 2'].fillna('None')
dataset_num = dataset[['Total','HP','Attack','Defense','Sp. Atk','Sp. Def','Speed']]
from sklearn.preprocessing import StandardScaler
sc_X = StandardScaler()
dataset_num = sc_X.fit_transform(dataset_num)
# +
dataset_scaleddataset = dataset.copy()
dataset_scaled[['Total','HP','Attack','Defense','Sp. Atk','Sp. Def','Speed']] = dataset_num
# -
from sklearn.cluster import KMeans
wcss = []
for i in range(1, 41):
kmeans = KMeans(n_clusters = i, init = 'k-means++', random_state = 42)
kmeans.fit(dataset_num)
wcss.append(kmeans.inertia_)
plt.plot(range(1, 41), wcss)
plt.title('The Elbow Method')
plt.xlabel('Number of clusters')
plt.ylabel('WCSS')
plt.xticks(np.arange(1, 41, 1.0))
plt.grid(which='major', axis='x')
plt.show()
kmeans = KMeans(n_clusters = 5, init = 'k-means++', random_state = 42)
y_kmeans = kmeans.fit_predict(dataset_num)
dataset['y_kmeans'] = y_kmeans
dataset.head()
sns.violinplot(x='y_kmeans', y='Total', data=dataset)
plt.show()
sns.violinplot(x='y_kmeans', y='Attack', data=dataset)
plt.show()
sns.violinplot(x='y_kmeans', y='Defense', data=dataset)
plt.show()
dataset.sort_values('Defense', axis=0, ascending=False).head(10)
sns.violinplot(x='y_kmeans', y='Speed', data=dataset)
plt.show()
dataset.sort_values('Speed', axis=0, ascending=False).head(15)
sns.violinplot(x='y_kmeans', y='Sp. Atk', data=dataset)
plt.show()
sns.violinplot(x='y_kmeans', y='Sp. Def', data=dataset)
plt.show()
# +
datasetdataset..sort_valuessort_val ('Sp. Def', axis=0, ascending=False).head(10)
# +
# Clusters by Type
#Get counts by type and cluster
#We need to merge the two columns Type 1 and Type 2 together the type can appear in either column
data_pct_1 = dataset.groupby(['Type 1', 'y_kmeans'])['Name'].count().to_frame().reset_index()
data_pct_1.columns = ['Type', 'y_kmeans', 'count_1']
data_pct_2 = dataset.groupby(['Type 2', 'y_kmeans'])['Name'].count().to_frame().reset_index()
data_pct_2.columns = ['Type', 'y_kmeans', 'count_2']
data_pct = data_pct_1.merge(data_pct_2, how='outer',
left_on=['Type', 'y_kmeans'],
right_on=['Type', 'y_kmeans'])
data_pct.fillna(0, inplace=True)
data_pct['count'] = data_pct['count_1'] + data_pct['count_2']
#Get counts by type
data_pct_Total = data_pct.groupby(['Type']).sum()['count'].reset_index()
data_pct_Total.columns = ['Type', 'count_total']
#Merge two dataframes and create percentage column
data_pct = data_pct.merge(right=data_pct_Total,
how='inner',
left_on='Type',
right_on='Type')
data_pct['pct'] = data_pct['count'] / data_pct['count_total']
#Create Graph
sns.barplot(x='Type', y='pct', data=data_pct, estimator=sum, ci=None, color='#34495e', label='4')
sns.barplot(x='Type', y='pct', data=data_pct[data_pct['y_kmeans'] <= 3],
estimator=sum, ci=None, color='#e74c3c', label='3')
sns.barplot(x='Type', y='pct', data=data_pct[data_pct['y_kmeans'] <= 2],
estimator=sum, ci=None, color='#95a5a6', label='2')
sns.barplot(x='Type', y='pct', data=data_pct[data_pct['y_kmeans'] <= 1],
estimator=sum, ci=None, color='#3498db', label='1')
sns.barplot(x='Type', y='pct', data=data_pct[data_pct['y_kmeans'] == 0],
estimator=sum, ci=None, color='#9b59b6', label='0')
plt.legend(loc='upper right', bbox_to_anchor=(1.1, 1))
plt.xticks(rotation=90)
plt.ylabel('Percentage')
plt.tight_layout()
plt.show()
# +
#Clusters by Generation
#Get counts by generation and cluster
#We need to merge the two columns Type 1 and Type 2 together the type can appear in either column
data_pct = dataset.groupby(['Generation', 'y_kmeans'])['Name'].count().to_frame().reset_index()
data_pct.columns = ['Generation', 'y_kmeans', 'count']
#Get counts by type
data_pct_Total = data_pct.groupby(['Generation']).sum()['count'].reset_index()
data_pct_Total.columns = ['Generation', 'count_total']
#Merge two dataframes and create percentage column
data_pct = data_pct.merge(right=data_pct_Total,
how='inner',
left_on='Generation',
right_on='Generation')
data_pct['pct'] = data_pct['count'] / data_pct['count_total']
#Create Graph
sns.barplot(x='Generation', y='pct', data=data_pct, estimator=sum, ci=None, color='#34495e', label='4')
sns.barplot(x='Generation', y='pct', data=data_pct[data_pct['y_kmeans'] <= 3],
estimator=sum, ci=None, color='#e74c3c', label='3')
sns.barplot(x='Generation', y='pct', data=data_pct[data_pct['y_kmeans'] <= 2],
estimator=sum, ci=None, color='#95a5a6', label='2')
sns.barplot(x='Generation', y='pct', data=data_pct[data_pct['y_kmeans'] <= 1],
estimator=sum, ci=None, color='#3498db', label='1')
sns.barplot(x='Generation', y='pct', data=data_pct[data_pct['y_kmeans'] == 0],
estimator=sum, ci=None, color='#9b59b6', label='0')
plt.legend(loc='upper right', bbox_to_anchor=(1.1, 1))
plt.xticks(rotation=90)
plt.ylabel('Percentage')
plt.tight_layout()
plt.show()
# -
| homework2/main-2018_0531_1359.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow_probability as tfp
import tensorflow as tf
import collections
# +
tfd = tfp.distributions
try:
tf.compat.v1.enable_eager_execution()
except ValueError:
pass
import matplotlib.pyplot as plt
# -
normal_dist = tfd.Normal(loc=0,scale=1)
normal_dist.sample(10)
normal_dist.log_prob(0.)
data_normal = normal_dist.sample(100)
plt.scatter(range(len(data_normal)),data_normal,color="blue",alpha=0.4)
plt.title("Normal Distribution")
plt.show()
gamma_dist = tfd.Gamma(concentration=3.0,rate=2.0)
data_gamma = gamma_dist.sample(100)
plt.scatter(range(len(data_gamma)),data_gamma,color="blue")
plt.title("Gamma Distribution")
plt.show()
# +
#normal_dist.cdf(100.0),gamma_dist.cdf(value=100.0)
import numpy as np
np.std(data_gamma)
np.std(data_normal)
# -
#
# Here is an example typesetting mathematics in \LaTeX
# \begin{equation*}
# X(m,n) = \left\{\begin{array}{lr}
# x(n), & \text{for } 0\leq n\leq 1\\
# \frac{x(n-1)}{2}, & \text{for } 0\leq n\leq 1\\
# \log_2 \left\lceil n \right\rceil \qquad & \text{for } 0\leq n\leq 1
# \end{array}\right\} = xy
# \end{equation*}
#
#
| Python/tfp_intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <small><small><i>
# All of these python notebooks are available at [ https://github.com/milaan9/Python4DataScience ]
# </i></small></small>
# # Integer & Linear Programming
#
# ## An example
#
#
# Setting up data: cost matrix, demand, supply
F = range(2) # 2 factories
R = range(12) # 12 retailers
C = [[1,2,2,1,3,4,5,7,5,2,3,2],
[4,5,5,4,1,3,1,2,1,2,4,6]]
demand = [9,4,2,6,4,5,7,8,3,6,9,5]
print('Total demand =',sum(demand))
supply = [ 34, 45]
# Now we can define a linear program
# $$\min \sum_{i\in F}\sum_{j\in R} c_{ij} x_{ij}$$
# Subject To
# $$\sum_{r\in R} x_{fr} \le s_r\quad\forall f\in F$$
# $$\sum_{f\in F} x_{fr} = d_f\quad\forall r\in R$$
# $$x \ge 0$$
# +
from mymip.mycplex import Model
lp = Model()
# define double indexed variables and give them a meaningful names
x = [ [lp.variable('x%dto%d'%(i,j)) for j in R]
for i in F ]
lp.min( sum( C[i][j] * x[i][j] for i in F for j in R))
# constraints can be given names too:
lp.SubjectTo({"Supply%d"%f: sum(x[f][r] for r in R) <= supply[f] for f in F})
lp.SubjectTo(("Demand%02d"%r, sum(x[f][r] for f in F) == demand[r] ) for r in R)
for f in F:
for r in R: x[f][r] >= 0 # all variables non-negative
lp.param["SCRIND"]=1 # set parameter to show CPLEX output
lp.optimise()
print("The minimum cost is",lp.objective())
for r in R:
for f in F:
if x[f][r].x > 0: # amount is not zero
print("%.1f from F%d to R%02d"%(x[f][r].x,f,r))
# -
# To see how the solve sees this problem, try writing it out to file and printing the contents of the file:
lp.write("myfirst.lp")
print(open("myfirst.lp","r").read())
# ## Advanced Usage
# All of the raw CPLEX callable library functions as per the can be accessed if required using C interface (see [IBM's CPLEX documentation](http://www.ibm.com/support/knowledgecenter/SSSA5P_12.6.2/ilog.odms.cplex.help/refcallablelibrary/groups/homepagecallable.html)) can be accessed by using the `cplex` object.
#
# For example to identify a minimal conflict set leading to infeasiblity of the problem we can use the [CPXrefineconflict](http://www.ibm.com/support/knowledgecenter/SSSA5P_12.6.2/ilog.odms.cplex.help/refcallablelibrary/cpxapi/refineconflict.html) and write out the conflict set using [CPXclpwrite](http://www.ibm.com/support/knowledgecenter/en/SSSA5P_12.6.2/ilog.odms.cplex.help/refcallablelibrary/cpxapi/clpwrite.html)
from mymip.mycplex import cplex
lp.SubjectTo(x[0][0]+x[1][1] >= 50) # make it infeasible
lp.optimise() # error!
cplex.CPXrefineconflict(lp.Env,lp.LP,0,0) # 2 null pointers
cplex.CPXclpwrite(lp.Env,lp.LP,b"conflict.lp") # note binary (ascii) string
print("#"*10,"Conflict LP","#"*10)
print(open("conflict.lp","r").read())
| 09_Integer_&_Linear_Programming.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.offsetbox import AnchoredText
import mplhep as hep
import pickle
from coffea.util import load
from coffea.hist import plot
import re
lumi_weights = None
with open("filefetcher/aram_samples.scales.pkl", "rb") as fin:
lumi_weights = pickle.load(fin)
lumi_weights["data_mu"] = 1.0
lumi_weights["data_el"] = 1.0
output = load("hists_wpt_filefetcher_aram_sample.coffea")
from hist import Hist, Stack
data_mu = {name: h * lumi_weights[name] for name, h in output.items() if name.find("data") > -1 and name[-2:] == "mu"}
mc_mu = {name: h * lumi_weights[name] for name, h in output.items() if name.find("data") == -1 and name[-2:] == "mu"}
data_el = {name: h * lumi_weights[name] for name, h in output.items() if name.find("data") > -1 and name[-2:] == "el"}
mc_el = {name: h * lumi_weights[name] for name, h in output.items() if name.find("data") == -1 and name[-2:] == "el"}
# +
mts_mu = [h["cent", ...].project("charge", "mt") for _, h in mc_mu.items()]
mts_mu_data = sum(h["cent", :, :1, ...] for h in data_mu.values()).project("charge","mt")
mts_mu_scaled = Stack(*mts_mu)
mts_el = [h["cent", ...].project("mt") for _, h in mc_el.items()]
mts_el_scaled = Stack(*mts_el)
# -
print(data_mu["data_mu"]["cent", : , :, ...])
# +
plt.style.use([hep.style.ROOT, hep.style.firamath])
f, axs = plt.subplots(1, 2, sharex=True, sharey=True, figsize=(20,10))
for i, q in enumerate([-1, 1]):
hep.histplot([x[q, ...] for x in mts_mu_scaled], stack=True, label=[name for name, _ in mc_mu.items()], ax=axs[i])
hep.histplot(mts_mu_data[q, ...], None, yerr=True, histtype="errorbar", label="Data", color="black", ax=axs[i])
axs[i].legend(fontsize=12)
hep.cms.label(loc=0, data=True, llabel="Preliminary", ax=axs[i])
plt.show()
# -
analysis["wl0_mu", "cent", ...].project("mt") * lumi_weights["wl0_mu"]
# +
plt.style.use([hep.style.ROOT, {'font.size': 16}])
# plot options for data
data_err_opts = {
'linestyle': 'none',
'marker': '.',
'markersize': 10.,
'color': 'k',
'elinewidth': 1,
}
for i in range(0, int(round(len(output.keys())))-1, 2)[2:]:
fig, ((ax1, ax2),(rax1, rax2)) = plt.subplots(2, 2, figsize=(16, 7), gridspec_kw={"height_ratios": (3, 1)}, sharex=True)
fig.subplots_adjust(hspace=.07)
h1name = list(output.keys())[i]
h2name = list(output.keys())[i+1]
if any([h1name.startswith('cutflow'), h2name.startswith('cutflow')]): break
print(h1name, h2name)
h1 = output[h1name]
h2 = output[h2name]
# if True: normalize histo to 1
dense = False
for ax, rax, h in zip([ax1, ax2], [rax1, rax2], [h1, h2]):
# scale MC samples according to lumi (for now, dummy scale)
scales = {
'ttbar': 1.0,
'ttbarprime':1.0
}
h.scale(scales,axis='dataset')
notdata = re.compile('(?!Data)')
if hasattr(h, 'dim'):
plot.plot1d(h[notdata], ax=ax, legend_opts={'loc':1}, density=dense, stack=True, clear=False);
plot.plot1d(h["Data"], ax=ax, legend_opts={'loc':1}, density=dense, error_opts=data_err_opts, clear=False);
# now we build the ratio plot
plot.plotratio(
num=h["Data"].sum("dataset"),
denom=h[notdata].sum("dataset"),
ax=rax,
error_opts=data_err_opts,
denom_fill_opts={},
guide_opts={},
unc='num'
)
else:
continue
#break
for ax, rax, hname in zip([ax1, ax2], [rax1, rax2], [h1name, h2name]):
at = AnchoredText(r"$1\mu, 1e$"+"\n"+
"2+ jets"+"\n"+
r"$|\eta| < 2.5$",
loc=2, frameon=False)
ax.add_artist(at)
ax.set_ylim(0.001, None)
if hname.startswith("btag") or hname.startswith("DeepCSV_trackDecayLenVal"):
ax.semilogy()
rax.set_ylabel('Data/Pred.')
rax.set_ylim(0,2)
ax.set_xlabel(None)
hep.mpl_magic(ax1)
hep.mpl_magic(ax2)
# -
| plots_and_combine_inputs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
import torch.nn as nn
from torch.utils.data import Dataset
import pandas, numpy, random
import matplotlib.pyplot as plt
# +
# if CUDA is available, use GPU and set default tensor type to cuda
if torch.cuda.is_available():
torch.set_default_tensor_type(torch.cuda.FloatTensor)
print("using cuda:", torch.cuda.get_device_name(0))
pass
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
device
# +
max_length_smiles = 29
input_para = 200
class Token(nn.Embedding):
def __init__(self, vocab_size, embed_size=200, padding_idx=0):
super().__init__(vocab_size, embed_size, padding_idx)
token = Token(30)
def convertToASCII(item):
ascii = []
for char in item:
ascii.append(ord(char))
return ascii
def convertASCIIToString(items):
result = ""
for item in items:
#print("item", item)
result = result + chr(item)
return result
# +
# dataset class
class qm9Dataset(Dataset):
def __init__(self, csv_file):
self.data_df = pandas.read_csv(csv_file, header=None, skiprows=1, dtype='unicode')
#print(self.data_df)
pass
def __len__(self):
return len(self.data_df)
def __getitem__(self, index):
smiles = (self.data_df.iloc[index,0] + " " * max_length_smiles)[0:max_length_smiles]
smiles = torch.cuda.FloatTensor(convertToASCII(smiles)) / 128.0 #ascii max value
labels = []
labels.append(float(self.data_df.iloc[index,1]))
labels.append(float(self.data_df.iloc[index,2]))
labels.append(float(self.data_df.iloc[index,3]))
labels.append(float(self.data_df.iloc[index,4]))
labels.append(float(self.data_df.iloc[index,5]))
labels.append(float(self.data_df.iloc[index,6]))
labels.append(float(self.data_df.iloc[index,7]))
labels.append(float(self.data_df.iloc[index,8]))
labels.append(float(self.data_df.iloc[index,9]))
labels.append(float(self.data_df.iloc[index,10]))
labels.append(float(self.data_df.iloc[index,11]))
labels.append(float(self.data_df.iloc[index,11]))
labels.append(float(self.data_df.iloc[index,12]))
labels = torch.cuda.FloatTensor(labels)
target = torch.zeros((len(self.data_df))).to(device)
target[index] = 1.0
return smiles,labels,target
pass
# -
qm9dataset = qm9Dataset('dataset/qm9.csv')
qm9dataset[2]
# +
from torch.utils.data import DataLoader
data_loader = torch.utils.data.DataLoader(qm9dataset,batch_size=1,shuffle=False)
'''
max_length_smiles = 0
i = 0
for smiles,label,target in data_loader:
i += 1
if (i % 1000 == 0):
print(smiles)
max_length_smiles = max(max_length_smiles, len(smiles[0]))
#print("Max length smiles:", max_length_smiles)
#max_length_smiles = 29
'''
# +
def generate_random_smiles(size):
random_length = random.randint(0, size-1)
random_data = torch.randint(32, 128, (1,size))
random_data[0][random_length:size] = 32 # space
result = random_data.float()/128.0
return result
def generate_random_one_hot(size):
label_tensor = torch.zeros((size))
random_idx = random.randint(0,size)
label_tensor[random_idx] = 1.0
return label_tensor
def generate_random_seed(size):
random_data = torch.randn(size)
return random_data
class View(nn.Module):
def __init__(self, shape):
super().__init__()
self.shape = shape,
def forward(self, x):
return x.view(*self.shape)
generate_random_seed(max_length_smiles)
# +
# discriminator class
class Discriminator(nn.Module):
def __init__(self):
# initialise parent pytorch class
super().__init__()
# define neural network layers
self.model = nn.Sequential(
View(max_length_smiles),
#nn.Linear(30, 100),
#nn.LeakyReLU(),
#nn.LayerNorm(100),
#nn.Linear(100, 1),
#nn.Sigmoid()
nn.Linear(max_length_smiles, 200),
nn.Sigmoid(),
nn.Linear(200,1),
nn.Sigmoid()
)
# create loss function
self.loss_function = nn.BCELoss()
#nn.MSELoss()
#nn.BCELoss()
# create optimiser, simple stochastic gradient descent
self.optimiser = torch.optim.Adam(self.parameters(), lr=0.0001)
# counter and accumulator for progress
self.counter = 0;
self.progress = []
pass
def forward(self, inputs):
# combine seed and label
#inputs = torch.cat((image_tensor, label_tensor))
return self.model(inputs)
def train(self, inputs, targets):
#print("inputs:", inputs)
# calculate the output of the network
outputs = self.forward(inputs)
#print("outputs", outputs)
# calculate loss
loss = self.loss_function(outputs, targets)
# increase counter and accumulate error every 10
self.counter += 1;
if (self.counter % 10 == 0):
self.progress.append(loss.item())
pass
if (self.counter % 10000 == 0):
print("counter = ", self.counter)
pass
# zero gradients, perform a backward pass, update weights
self.optimiser.zero_grad()
loss.backward()
self.optimiser.step()
pass
def plot_progress(self):
df = pandas.DataFrame(self.progress, columns=['loss'])
df.plot(ylim=(0), figsize=(16,8), alpha=0.1, marker='.', grid=True, yticks=(0, 0.25, 0.5, 1.0, 5.0))
pass
pass
# +
# %%time
# test discriminator can separate real data from random noise
D = Discriminator()
D.to(device)
for smiles,label,target in data_loader:
# real
#print("smiles.shape", smiles.shape)
D.train(smiles[0], torch.cuda.FloatTensor([1.0]))
# fake
D.train(generate_random_seed(max_length_smiles), torch.cuda.FloatTensor([0.0]))
pass
# +
for i in range(4):
#random.randint(0,20000)
data_tensor,_,_ = qm9dataset[random.randint(0,130000)]
print( D.forward( data_tensor ).item() )
pass
for i in range(4):
print( D.forward(generate_random_seed(max_length_smiles)).item() )
pass
# -
D.plot_progress()
class Generator(nn.Module):
def __init__(self):
# initialise parent pytorch class
super().__init__()
# define neural network layers
self.model = nn.Sequential(
#nn.BatchNorm2d(input_para),
#nn.Upsample(scale_factor=2),
nn.Linear(input_para, 100),
#nn.LeakyReLU(0.02),
nn.ReLU(),
#nn.LayerNorm(200),
nn.Linear(100, max_length_smiles),
nn.Sigmoid()
)
# create optimiser, simple stochastic gradient descent
self.optimiser = torch.optim.Adam(self.parameters(), lr=0.001)
# counter and accumulator for progress
self.counter = 0;
self.progress = []
pass
def forward(self, inputs):
return self.model(inputs)
def train(self, D, inputs, targets):
# calculate the output of the network
g_output = self.forward(inputs)
# pass onto Discriminator
d_output = D.forward(g_output)
# calculate error
loss = D.loss_function(d_output, targets)
# increase counter and accumulate error every 10
self.counter += 1;
if (self.counter % 10 == 0):
self.progress.append(loss.item())
pass
# zero gradients, perform a backward pass, update weights
self.optimiser.zero_grad()
loss.backward()
self.optimiser.step()
pass
def plot_progress(self):
df = pandas.DataFrame(self.progress, columns=['loss'])
df.plot(ylim=(0), figsize=(16,8), alpha=0.1, marker='.', grid=True, yticks=(0, 0.25, 0.5, 1.0, 5.0))
pass
pass
# +
G = Generator()
G.to(device)
output = G.forward(generate_random_seed(input_para))
print("output", output)
# +
# %%time
D = Discriminator()
D.to(device)
G = Generator()
G.to(device)
epochs = 1
for epoch in range(epochs):
print ("epoch = ", epoch + 1)
i = 0
# train Discriminator and Generator
for smiles,label,target in data_loader:
# train discriminator on true
D.train(smiles[0], torch.cuda.FloatTensor([1.0]))
# train discriminator on false
# use detach() so gradients in G are not calculated
#D.train(G.forward(generate_random_smiles(input_para)).detach(), torch.cuda.FloatTensor([0.0]))
D.train(G.forward(generate_random_seed(input_para)).detach()[0], torch.cuda.FloatTensor([0.0]))
# train generator
#G.train(D, generate_random_seed(input_para), torch.cuda.FloatTensor([1.0]))
#G.train(D, generate_random_smiles(input_para), torch.cuda.FloatTensor([1.0]))
G.train(D, generate_random_seed(input_para), torch.cuda.FloatTensor([1.0]))
i +=1
pass
pass
import pickle
from datetime import datetime
now = datetime.now()
date_time = now.strftime("%Y_%m_%d_%H_%M")
G_filename = 'pre_train_model/GAN3_G_' + date_time + '.sav'
print('save model to file:', G_filename)
pickle.dump(G, open(G_filename, 'wb'))
D_filename = 'pre_train_model/GAN3_D_' + date_time + '.sav'
print('save model to file:', D_filename)
pickle.dump(D, open(D_filename, 'wb'))
G_model = pickle.load(open(G_filename, 'rb'))
D_model = pickle.load(open(D_filename, 'rb'))
output = G_model.forward(generate_random_smiles(input_para))
D_model.forward(output)
# -
D.plot_progress()
G.plot_progress()
for i in range(10):
output = G.forward(generate_random_seed(input_para))
smiles = output.detach().cpu().numpy() * 128
#print(convertASCIIToString(smiles))
print("smiles:", smiles.astype(int) )
pass
for i in range(10):
data_tensor,_,_ = qm9dataset[random.randint(0,130000)]
#print(D_model.forward(data_tensor).item())
print(D.forward( data_tensor ).item() )
for i in range(10):
output = G.forward(generate_random_seed(input_para))
print(D.forward( output ))
| archive/GAN3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from __future__ import division, print_function
import numpy as np
import matplotlib.pyplot as plt
from pymatgen.core import Element, Composition
# %matplotlib inline
# +
import csv
with open("ICSD/icsd-ternaries.csv", "r") as f:
csv_reader = csv.reader(f, dialect = csv.excel_tab)
data = [line for line in csv_reader]
formulas = [line[2] for line in data]
compositions = [Composition(f) for f in formulas]
# -
# ## Structure Types
#
# Structure types are assigned by hand by ICSD curators.
# How many ternaries have been assigned a structure type?
structure_types = [line[3] for line in data if line[3] is not '']
unique_structure_types = set(structure_types)
print("There are {} ICSD ternaries entries.".format(len(data)))
print("Structure types are assigned for {} entries.".format(len(structure_types)))
print("There are {} unique structure types.".format(len(unique_structure_types)))
# Filter for stoichiometric compounds only:
def is_stoichiometric(composition):
return np.all(np.mod(composition.values(), 1) == 0)
stoichiometric_compositions = [c for c in compositions if is_stoichiometric(c)]
print("Number of stoichiometric compositions: {}".format(len(stoichiometric_compositions)))
ternaries = set(c.formula for c in stoichiometric_compositions)
len(ternaries)
data_stoichiometric = [x for x in data if is_stoichiometric(Composition(x[2]))]
# +
from collections import Counter
struct_type_freq = Counter(x[3] for x in data_stoichiometric if x[3] is not '')
# -
plt.loglog(range(1, len(struct_type_freq)+1),
sorted(struct_type_freq.values(), reverse = True), 'o')
sorted(struct_type_freq.items(), key = lambda x: x[1], reverse = True)
len(set([x[2] for x in data if x[3] == 'Perovskite-GdFeO3']))
uniq_phases = set()
for row in data_stoichiometric:
spacegroup, formula, struct_type = row[1:4]
phase = (spacegroup, Composition(formula).formula, struct_type)
uniq_phases.add(phase)
uniq_struct_type_freq = Counter(x[2] for x in uniq_phases if x[2] is not '')
uniq_struct_type_freq_sorted = sorted(uniq_struc_type_freq.items(), key = lambda x: x[1], reverse = True)
plt.loglog(range(1, len(uniq_struct_type_freq_sorted)+1),
[x[1] for x in uniq_struct_type_freq_sorted], 'o')
uniq_struct_type_freq_sorted
for struct_type,freq in uniq_struct_type_freq_sorted[:10]:
print("{} : {}".format(struct_type, freq))
fffs = [p[1] for p in uniq_phases if p[2] == struct_type]
fmt = " ".join(["{:14}"]*5)
print(fmt.format(*fffs[0:5]))
print(fmt.format(*fffs[5:10]))
print(fmt.format(*fffs[10:15]))
print(fmt.format(*fffs[15:20]))
# ## Long Formulas
# What are the longest formulas?
for formula in sorted(formulas, key = lambda x: len(x), reverse = True)[:20]:
print(formula)
# Two key insights:
# 1. Just because there are three elements in the formula
# doesn't mean the compound is fundamentally a ternary.
# There are doped binaries which masquerade as ternaries.
# And there are doped ternaries which masquerade as quaternaries,
# or even quintenaries. Because I only asked for compositions
# with 3 elements, this data is missing.
# 2. ICSD has strategically placed parentheses in the formulas
# which give hints as to logical groupings. For example:
# (Ho1.3 Ti0.7) ((Ti0.64 Ho1.36) O6.67)
# is in fact in the pyrochlore family, A2B2O7.
# ## Intermetallics
#
# How many intermetallics does the ICSD database contain?
# +
def filter_in_set(compound, universe):
return all((e in universe) for e in Composition(compound))
transition_metals = [e for e in Element if e.is_transition_metal]
tm_ternaries = [c for c in formulas if filter_in_set(c, transition_metals)]
print("Number of intermetallics:", len(tm_ternaries))
# -
unique_tm_ternaries = set([Composition(c).formula for c in tm_ternaries])
print("Number of unique intermetallics:", len(unique_tm_ternaries))
unique_tm_ternaries
| notebooks/old_ICSD_Notebooks/Understanding ICSD data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## MNIST 手寫數字資料
# 
# 各數字墨水用量的平均、標準差
# ```
# digit: ink_mean, ink_std
# 0: 44.216828, 10.736191
# 1: 19.379654, 5.607750
# 2: 37.988658, 9.703046
# 3: 36.090187, 9.635850
# 4: 30.948226, 8.130984
# 5: 32.831095, 9.502106
# 6: 35.011953, 9.544340
# 7: 29.204563, 7.811263
# 8: 38.289775, 9.851285
# 9: 31.260435, 8.273293
# ```
# Q1(a). 用墨水量猜猜看以下分別是哪一個數字?
# ```
# index: ink
# 59677: 34.308673
# 35981: 25.968112
# 6865: 56.959184
# 26064: 22.743622
# 18741: 16.200255
# 22600: 37.936224
# 27950: 24.501276
# 16411: 36.030612
# 5475: 69.881378
# 32134: 28.904337
# ```
# Q1(b). 用每個數字附近的數字,來猜猜看它是哪個數字?
# ```
# ind: neighboring digits
# 28370: [2 1 1 2 6 1]
# 34420: [9 0 9 4 7 7]
# 11959: [6 1 1 1 1 1]
# 55532: [4 4 6 4 5 4]
# 54421: [0 5 7 4 7 4]
# 43773: [3 3 3 2 3 0]
# 25899: [1 1 1 1 1 1]
# 24254: [7 7 7 4 0 7]
# 2927: [8 0 8 5 3 6]
# 39562: [5 9 3 6 4 6]
# ```
#
#
#
#
#
#
#
#
# ## NSYSU-digits 手寫資料
# 
# 各數字墨水用量的平均、標準差
# ```
# digit: ink_mean, ink_std
# 0: 8.794989, 2.483019
# 1: 6.589010, 1.590713
# 2: 8.955215, 2.848100
# 3: 8.041135, 2.143122
# 4: 10.732587, 2.402498
# 5: 8.376941, 2.135341
# 6: 9.320737, 2.628407
# 7: 9.650199, 2.013895
# 8: 12.371447, 3.414354
# 9: 9.639391, 2.456494
# ```
# Q1(a). 用墨水量猜猜看以下分別是哪一個數字?
# ```
# index: ink
# 273: 12.673469
# 87: 14.271684
# 490: 13.397959
# 14: 13.533163
# 367: 4.100765
# 390: 8.561224
# 311: 5.316327
# 211: 9.485969
# 299: 8.251276
# 229: 10.153061
# ```
# Q1(b). 用每個數字附近的數字,來猜猜看它是哪個數字?
# ```
# ind: neighboring digits
# 53: [2 3 0 6 2]
# 483: [1 7 1 3 1]
# 60: [1 6 6 1 1]
# 257: [0 4 4 0 6]
# 358: [0 1 5 4 9]
# 230: [3 3 6 3 9]
# 287: [1 1 1 9 4]
# 280: [7 7 1 0 6]
# 84: [8 0 8 2 5]
# 264: [9 1 1 7 0]
# ```
#
#
#
#
#
#
#
#
#
# ## Answers
# A1(a). 有猜對嗎?
# ```
# index: ink <-- answer
# 59677: 34.308673 <-- 2: 37.988658, 9.703046
# 35981: 25.968112 <-- 8: 38.289775, 9.851285
# 6865: 56.959184 <-- 4: 30.948226, 8.130984
# 26064: 22.743622 <-- 9: 31.260435, 8.273293
# 18741: 16.200255 <-- 1: 19.379654, 5.607750
# 22600: 37.936224 <-- 6: 35.011953, 9.544340
# 27950: 24.501276 <-- 7: 29.204563, 7.811263
# 16411: 36.030612 <-- 3: 36.090187, 9.635850
# 5475: 69.881378 <-- 0: 44.216828, 10.736191
# 32134: 28.904337 <-- 5: 32.831095, 9.502106
# ```
# A1(b). 有猜對嗎?
# ```
# ind: neighboring digits <-- answer
# 28370: [2 1 1 2 6 1] <-- 2
# 34420: [9 0 9 4 7 7] <-- 9
# 11959: [6 1 1 1 1 1] <-- 6
# 55532: [4 4 6 4 5 4] <-- 4
# 54421: [0 5 7 4 7 4] <-- 0
# 43773: [3 3 3 2 3 0] <-- 3
# 25899: [1 1 1 1 1 1] <-- 1
# 24254: [7 7 7 4 0 7] <-- 7
# 2927: [8 0 8 5 3 6] <-- 8
# 39562: [5 9 3 6 4 6] <-- 5
# ```
# A2(a). 有猜對嗎?
# ```
# index: ink <-- answer
# 273: 12.673469 <-- 2: 8.955215, 2.848100
# 87: 14.271684 <-- 8: 12.371447, 3.414354
# 490: 13.397959 <-- 4: 10.732587, 2.402498
# 14: 13.533163 <-- 9: 9.639391, 2.456494
# 367: 4.100765 <-- 1: 6.589010, 1.590713
# 390: 8.561224 <-- 6: 9.320737, 2.628407
# 311: 5.316327 <-- 7: 9.650199, 2.013895
# 211: 9.485969 <-- 3: 8.041135, 2.143122
# 299: 8.251276 <-- 0: 8.794989, 2.483019
# 229: 10.153061 <-- 5: 8.376941, 2.135341
# ```
# A2(b). 有猜對嗎?
# ```
# ind: neighboring digits <-- answer
# 53: [2 3 0 6 2] <-- 2
# 483: [1 7 1 3 1] <-- 9
# 60: [1 6 6 1 1] <-- 6
# 257: [0 4 4 0 6] <-- 4
# 358: [0 1 5 4 9] <-- 0
# 230: [3 3 6 3 9] <-- 3
# 287: [1 1 1 9 4] <-- 1
# 280: [7 7 1 0 6] <-- 7
# 84: [8 0 8 2 5] <-- 8
# 264: [9 1 1 7 0] <-- 5
# ```
#
#
#
#
#
#
#
#
# ## Code
import os
import numpy as np
import matplotlib.pyplot as plt
# ### MNIST
# +
### load MNIST
import tensorflow as tf
from tensorflow.keras.datasets import mnist
(X_train, y_train), (X_test, y_test) = mnist.load_data()
X = X_train
y = y_train
N = X.shape[0]
# +
fig, axs = plt.subplots(2, 5, figsize=(10,4))
for k in range(10):
ax = axs[k // 5, k % 5]
ax.axis('off')
mask = (y == k)
mean_img = X[mask].mean(axis=0)
ax.imshow(mean_img, cmap='Greys')
# -
### ink density histogram
# fig = plt.figure(figsize=(12,4))
dwise = np.zeros((10,2), dtype=float)
print("digit: ink_mean, ink_std")
for k in range(10):
mask = (y == k)
wanted = X.reshape(N, -1)[mask].mean(axis=1)
dwise[k][0],dwise[k][1] = wanted.mean(),wanted.std()
print("%d: %f, %f"%(k, dwise[k][0], dwise[k][1]))
# plt.hist(wanted, bins=100, label='%s'%k)
# plt.legend()
# +
rng = np.random.RandomState(0)
choices = np.arange(10)
rng.shuffle(choices)
print(choices)
inds = np.zeros((10,), dtype=int)
for i in range(10):
mask = (y == choices[i])
inds[i] = rng.choice(np.arange(N)[mask])
print("index: ink")
for i in range(10):
ind = inds[i]
k = choices[i]
ink = X[ind].mean()
print("%d: %f"%(ind,ink))
print("index: ink <-- answer")
for i in range(10):
ind = inds[i]
k = choices[i]
ink = X[ind].mean()
print("%d: %f <-- %d: %f, %f"%(ind,ink,k,dwise[k][0],dwise[k][1]))
# +
rng = np.random.RandomState(1)
choices = np.arange(10)
rng.shuffle(choices)
print(choices)
inds = np.zeros((10,), dtype=int)
for i in range(10):
mask = (y == choices[i])
inds[i] = rng.choice(np.arange(N)[mask])
print("ind: neighboring digits")
for i in range(10):
ind = inds[i]
k = choices[i]
x = X[ind]
dist = ((x - X)**2).sum(axis=-1).sum(axis=-1)
nbr = y[dist.argsort()[:6]]
print("%d: %s"%(ind, nbr))
print("ind: neighboring digits <-- answer")
for i in range(10):
ind = inds[i]
k = choices[i]
x = X[ind]
dist = ((x - X)**2).sum(axis=-1).sum(axis=-1)
nbr = y[dist.argsort()[:6]]
print("%d: %s <-- %d"%(ind, nbr, k))
# -
# ### NSYSU-digits
# +
### load nsysu
import urllib
import numpy as np
base = r"https://github.com/SageLabTW/auto-grading/raw/master/nsysu-digits/"
for c in ['X', 'y']:
filename = "nsysu-digits-%s.csv"%c
if filename not in os.listdir('.'):
print(filename, 'not found --- will download')
urllib.request.urlretrieve(base + c + ".csv", filename)
Xsys = np.genfromtxt('nsysu-digits-X.csv', dtype=int, delimiter=',') ### flattened already
ysys = np.genfromtxt('nsysu-digits-y.csv', dtype=int, delimiter=',')
N = Xsys.shape[0]
X = Xsys.reshape(N,28,28)
y = ysys
# +
fig, axs = plt.subplots(2, 5, figsize=(10,4))
for k in range(10):
ax = axs[k // 5, k % 5]
ax.axis('off')
mask = (y == k)
mean_img = X[mask].mean(axis=0)
ax.imshow(mean_img, cmap='Greys')
# -
### ink density histogram
# fig = plt.figure(figsize=(12,4))
dwise = np.zeros((10,2), dtype=float)
print("digit: ink_mean, ink_std")
for k in range(10):
mask = (y == k)
wanted = X.reshape(N, -1)[mask].mean(axis=1)
dwise[k][0],dwise[k][1] = wanted.mean(),wanted.std()
print("%d: %f, %f"%(k, dwise[k][0], dwise[k][1]))
# plt.hist(wanted, bins=100, label='%s'%k)
# plt.legend()
# +
rng = np.random.RandomState(0)
choices = np.arange(10)
rng.shuffle(choices)
print(choices)
inds = np.zeros((10,), dtype=int)
for i in range(10):
mask = (y == choices[i])
inds[i] = rng.choice(np.arange(N)[mask])
print("index: ink")
for i in range(10):
ind = inds[i]
k = choices[i]
ink = X[ind].mean()
print("%d: %f"%(ind,ink))
print("index: ink <-- answer")
for i in range(10):
ind = inds[i]
k = choices[i]
ink = X[ind].mean()
print("%d: %f <-- %d: %f, %f"%(ind,ink,k,dwise[k][0],dwise[k][1]))
# +
rng = np.random.RandomState(1)
choices = np.arange(10)
rng.shuffle(choices)
print(choices)
inds = np.zeros((10,), dtype=int)
for i in range(10):
mask = (y == choices[i])
inds[i] = rng.choice(np.arange(N)[mask])
print("ind: neighboring digits")
for i in range(10):
ind = inds[i]
k = choices[i]
x = X[ind]
dist = ((x - X)**2).sum(axis=-1).sum(axis=-1)
nbr = y[dist.argsort()[1:6]]
print("%d: %s"%(ind, nbr))
print("ind: neighboring digits <-- answer")
for i in range(10):
ind = inds[i]
k = choices[i]
x = X[ind]
dist = ((x - X)**2).sum(axis=-1).sum(axis=-1)
nbr = y[dist.argsort()[1:6]]
print("%d: %s <-- %d"%(ind, nbr, k))
# + slideshow={"slide_type": "skip"}
| NSYSU-digits/guess-2021.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# # Automated Machine Learning
# _**Remote Execution using DSVM (Ubuntu)**_
#
# ## Contents
# 1. [Introduction](#Introduction)
# 1. [Setup](#Setup)
# 1. [Data](#Data)
# 1. [Train](#Train)
# 1. [Results](#Results)
# 1. [Test](#Test)
# ## Introduction
# In this example we use the scikit-learn's [digit dataset](http://scikit-learn.org/stable/datasets/index.html#optical-recognition-of-handwritten-digits-dataset) to showcase how you can use AutoML for a simple classification problem.
#
# Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.
#
# In this notebook you wiil learn how to:
# 1. Create an `Experiment` in an existing `Workspace`.
# 2. Attach an existing DSVM to a workspace.
# 3. Configure AutoML using `AutoMLConfig`.
# 4. Train the model using the DSVM.
# 5. Explore the results.
# 6. Test the best fitted model.
#
# In addition, this notebook showcases the following features:
# - **Parallel** executions for iterations
# - **Asynchronous** tracking of progress
# - **Cancellation** of individual iterations or the entire run
# - Retrieving models for any iteration or logged metric
# - Specifying AutoML settings as `**kwargs`
# ## Setup
#
# As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments.
# +
import logging
import os
import time
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
from sklearn import datasets
import azureml.core
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
from azureml.train.automl import AutoMLConfig
# +
ws = Workspace.from_config()
# Choose a name for the run history container in the workspace.
experiment_name = 'automl-remote-dsvm'
project_folder = './sample_projects/automl-remote-dsvm'
experiment = Experiment(ws, experiment_name)
output = {}
output['SDK version'] = azureml.core.VERSION
output['Subscription ID'] = ws.subscription_id
output['Workspace Name'] = ws.name
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Project Directory'] = project_folder
output['Experiment Name'] = experiment.name
pd.set_option('display.max_colwidth', -1)
outputDf = pd.DataFrame(data = output, index = [''])
outputDf.T
# -
# Opt-in diagnostics for better experience, quality, and security of future releases.
from azureml.telemetry import set_diagnostics_collection
set_diagnostics_collection(send_diagnostics = True)
# ### Create a Remote Linux DSVM
# **Note:** If creation fails with a message about Marketplace purchase eligibilty, start creation of a DSVM through the [Azure portal](https://portal.azure.com), and select "Want to create programmatically" to enable programmatic creation. Once you've enabled this setting, you can exit the portal without actually creating the DSVM, and creation of the DSVM through the notebook should work.
#
# +
from azureml.core.compute import DsvmCompute
dsvm_name = 'mydsvma'
try:
dsvm_compute = DsvmCompute(ws, dsvm_name)
print('Found an existing DSVM.')
except:
print('Creating a new DSVM.')
dsvm_config = DsvmCompute.provisioning_configuration(vm_size = "Standard_D2s_v3")
dsvm_compute = DsvmCompute.create(ws, name = dsvm_name, provisioning_configuration = dsvm_config)
dsvm_compute.wait_for_completion(show_output = True)
print("Waiting one minute for ssh to be accessible")
time.sleep(60) # Wait for ssh to be accessible
# +
from azureml.core.runconfig import RunConfiguration
from azureml.core.conda_dependencies import CondaDependencies
# create a new RunConfig object
conda_run_config = RunConfiguration(framework="python")
# Set compute target to the Linux DSVM
conda_run_config.target = dsvm_compute
cd = CondaDependencies.create(pip_packages=['azureml-sdk[automl]'], conda_packages=['numpy'])
conda_run_config.environment.python.conda_dependencies = cd
# -
# ## Data
# For remote executions you should author a `get_data.py` file containing a `get_data()` function. This file should be in the root directory of the project. You can encapsulate code to read data either from a blob storage or local disk in this file.
# In this example, the `get_data()` function returns data using scikit-learn's [load_digits](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_digits.html) method.
if not os.path.exists(project_folder):
os.makedirs(project_folder)
# +
# %%writefile $project_folder/get_data.py
from sklearn import datasets
from scipy import sparse
import numpy as np
def get_data():
digits = datasets.load_digits()
X_train = digits.data[100:,:]
y_train = digits.target[100:]
return { "X" : X_train, "y" : y_train }
# -
# ## Train
#
# You can specify `automl_settings` as `**kwargs` as well. Also note that you can use a `get_data()` function for local excutions too.
#
# **Note:** When using Remote DSVM, you can't pass Numpy arrays directly to the fit method.
#
# |Property|Description|
# |-|-|
# |**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics: <br><i>accuracy</i><br><i>AUC_weighted</i><br><i>average_precision_score_weighted</i><br><i>norm_macro_recall</i><br><i>precision_score_weighted</i>|
# |**iteration_timeout_minutes**|Time limit in minutes for each iteration.|
# |**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|
# |**n_cross_validations**|Number of cross validation splits.|
# |**max_concurrent_iterations**|Maximum number of iterations to execute in parallel. This should be less than the number of cores on the DSVM.|
# +
automl_settings = {
"iteration_timeout_minutes": 10,
"iterations": 20,
"n_cross_validations": 5,
"primary_metric": 'AUC_weighted',
"preprocess": False,
"max_concurrent_iterations": 2,
"verbosity": logging.INFO
}
automl_config = AutoMLConfig(task = 'classification',
debug_log = 'automl_errors.log',
path = project_folder,
run_configuration=conda_run_config,
data_script = project_folder + "/get_data.py",
**automl_settings
)
# -
# **Note:** The first run on a new DSVM may take several minutes to prepare the environment.
# Call the `submit` method on the experiment object and pass the run configuration. For remote runs the execution is asynchronous, so you will see the iterations get populated as they complete. You can interact with the widgets and models even when the experiment is running to retrieve the best model up to that point. Once you are satisfied with the model, you can cancel a particular iteration or the whole run.
#
# In this example, we specify `show_output = False` to suppress console output while the run is in progress.
remote_run = experiment.submit(automl_config, show_output = False)
remote_run
# ## Results
#
# #### Loading Executed Runs
# In case you need to load a previously executed run, enable the cell below and replace the `run_id` value.
# + active=""
# remote_run = AutoMLRun(experiment=experiment, run_id = 'AutoML_480d3ed6-fc94-44aa-8f4e-0b945db9d3ef')
# -
# #### Widget for Monitoring Runs
#
# The widget will first report a "loading" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.
#
# You can click on a pipeline to see run properties and output logs. Logs are also available on the DSVM under `/tmp/azureml_run/{iterationid}/azureml-logs`
#
# **Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details.
from azureml.widgets import RunDetails
RunDetails(remote_run).show()
# Wait until the run finishes.
remote_run.wait_for_completion(show_output = True)
#
# #### Retrieve All Child Runs
# You can also use SDK methods to fetch all the child runs and see individual metrics that we log.
# +
children = list(remote_run.get_children())
metricslist = {}
for run in children:
properties = run.get_properties()
metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}
metricslist[int(properties['iteration'])] = metrics
rundata = pd.DataFrame(metricslist).sort_index(1)
rundata
# -
# ### Cancelling Runs
#
# You can cancel ongoing remote runs using the `cancel` and `cancel_iteration` functions.
# +
# Cancel the ongoing experiment and stop scheduling new iterations.
# remote_run.cancel()
# Cancel iteration 1 and move onto iteration 2.
# remote_run.cancel_iteration(1)
# -
# ### Retrieve the Best Model
#
# Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*.
best_run, fitted_model = remote_run.get_output()
print(best_run)
print(fitted_model)
# #### Best Model Based on Any Other Metric
# Show the run and the model which has the smallest `log_loss` value:
lookup_metric = "log_loss"
best_run, fitted_model = remote_run.get_output(metric = lookup_metric)
print(best_run)
print(fitted_model)
# #### Model from a Specific Iteration
# Show the run and the model from the third iteration:
iteration = 3
third_run, third_model = remote_run.get_output(iteration = iteration)
print(third_run)
print(third_model)
# ## Test
#
# #### Load Test Data
digits = datasets.load_digits()
X_test = digits.data[:10, :]
y_test = digits.target[:10]
images = digits.images[:10]
# #### Test Our Best Fitted Model
# Randomly select digits and test.
for index in np.random.choice(len(y_test), 2, replace = False):
print(index)
predicted = fitted_model.predict(X_test[index:index + 1])[0]
label = y_test[index]
title = "Label value = %d Predicted value = %d " % (label, predicted)
fig = plt.figure(1, figsize=(3,3))
ax1 = fig.add_axes((0,0,.8,.8))
ax1.set_title(title)
plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')
plt.show()
| how-to-use-azureml/automated-machine-learning/remote-execution/auto-ml-remote-execution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import numpy as np
# +
# 问题:
# 1.无法调整大小
# 2.通道不同不能使用
img=cv2.imread("./images/lena.jpg")
imgHor=np.hstack((img,img))
imgVer=np.vstack((img,img))
cv2.imshow("Horizationtal",imgHor)
cv2.imshow("Vertical",imgVer)
cv2.waitKey(0)
# +
# 解决:
def stackImages(scale,imgArray):
rows = len(imgArray)
cols = len(imgArray[0])
rowsAvailable = isinstance(imgArray[0], list)
width = imgArray[0][0].shape[1]
height = imgArray[0][0].shape[0]
if rowsAvailable:
for x in range ( 0, rows):
for y in range(0, cols):
if imgArray[x][y].shape[:2] == imgArray[0][0].shape [:2]:
imgArray[x][y] = cv2.resize(imgArray[x][y], (0, 0), None, scale, scale)
else:
imgArray[x][y] = cv2.resize(imgArray[x][y], (imgArray[0][0].shape[1], imgArray[0][0].shape[0]), None, scale, scale)
if len(imgArray[x][y].shape) == 2: imgArray[x][y]= cv2.cvtColor( imgArray[x][y], cv2.COLOR_GRAY2BGR)
imageBlank = np.zeros((height, width, 3), np.uint8)
hor = [imageBlank]*rows
hor_con = [imageBlank]*rows
for x in range(0, rows):
hor[x] = np.hstack(imgArray[x])
ver = np.vstack(hor)
else:
for x in range(0, rows):
if imgArray[x].shape[:2] == imgArray[0].shape[:2]:
imgArray[x] = cv2.resize(imgArray[x], (0, 0), None, scale, scale)
else:
imgArray[x] = cv2.resize(imgArray[x], (imgArray[0].shape[1], imgArray[0].shape[0]), None,scale, scale)
if len(imgArray[x].shape) == 2: imgArray[x] = cv2.cvtColor(imgArray[x], cv2.COLOR_GRAY2BGR)
hor= np.hstack(imgArray)
ver = hor
return ver
# +
img=cv2.imread("./images/lena.jpg")
imgStack=stackImages(0.5,([img,img,img]))
cv2.imshow("Stack",imgStack)
cv2.waitKey(0)
# +
imgGray=cv2.cvtColor(img,cv2.COLOR_BGR2GRAY)
imgStack=stackImages(0.5,([img,imgGray,img],[img,img,img]))
cv2.imshow("Stack",imgStack)
cv2.waitKey(0)
# -
| CV/OpenCV/Learn_OpenCV_in_3_hours/tutorial/CHAPTER6.Joining Images.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Packages:
import pandas as pd
import matplotlib.pyplot as plt
import plotly.express as px
import plotly.graph_objects as go
from urllib.request import urlopen
import json
import datetime
import numpy as np
import warnings
from textblob import TextBlob
import re
from wordcloud import WordCloud
import scipy.stats as st
import scipy
from scipy import stats
import statsmodels.api as sm
df = pd.read_csv('../data/processed/branson_cleaned.csv')
df.head()
# +
fig, (ax1, ax2, ax3, ax4) = plt.subplots(1, 4, figsize = (20, 5))
sns.kdeplot(ax=ax1, x=df['Price_delta'], bw_adjust=1).set_title('Stock price change')
sns.kdeplot(ax=ax2, x=df['Polarity_delta'], bw_adjust=0.27).set_title('Sentiment change')
sns.kdeplot(ax=ax3, x=df['Favorite_scaled'], bw_adjust=0.27).set_title('No. of likes per tweet')
sns.kdeplot(ax=ax4, x=df['Retweet_scaled'], bw_adjust=0.27).set_title('No. of retweets')
plt.tight_layout()
# -
df.describe()
# +
#from scipy.stats import boxcox
#df['transformed_return'], lam = boxcox(df["Return"])
# -
numeric_df = df.filter(items=['Price_scaled','Price_delta', 'Polarity_delta', 'Favorite_scaled', 'Retweet_scaled'])
corr1 = numeric_df.corr()
corr2 = numeric_df.corr(method='spearman')
sns.heatmap(corr1, annot=True, cmap=sns.diverging_palette(140, 10, as_cmap=True), fmt='.2f', vmin=-1, vmax=1)
plt.show()
sns.heatmap(corr2, annot=True, cmap=sns.diverging_palette(140, 10, as_cmap=True), fmt='.2f', vmin=-1, vmax=1)
plt.show()
# +
print("H0: the changes in stock market prices are similar on average compared to the changes in CEO's Twitter sentiment")
print("H1: the changes in stock market prices are different on average compared to the changes in CEO's Twitter sentiment")
statistic = st.ttest_ind(df['Price_delta'], df['Polarity_delta'], axis=0, equal_var=False, alternative='two-sided', nan_policy='omit')[0]
pvalue = st.ttest_ind(df['Price_delta'], df['Polarity_delta'], axis=0, equal_var=False, alternative='two-sided', nan_policy='omit')[1]
print('statistic = ',statistic)
print('pvalue = ',pvalue)
if pvalue<= 0.05:
print('We reject the null hypothesis')
else:
print('We fail to reject the null hypothesis')
# -
from pandas.plotting import scatter_matrix
df_sm = df[['Price_scaled','Price_delta', 'Polarity_delta','Favorite_scaled', 'Retweet_scaled', 'Polarity_scaled']]
df_sm.dropna(inplace=True)
scatter_matrix(df_sm, figsize=(12,8));
scipy.stats.shapiro(df_sm['Price_delta'])
scipy.stats.shapiro(df_sm['Polarity_delta'])
# +
fig, (ax1, ax2) = plt.subplots(1, 2, figsize = (15, 5))
sns.regplot(data = df_sm, x='Price_delta', y='Polarity_delta', lowess=False, ax=ax1).set(title='Sentiment vs. Stock price')
sns.regplot(data = df_sm, x='Polarity_delta', y='Price_delta', lowess=False, ax=ax2).set(title='Stock Price vs. Sentiment')
plt.tight_layout()
# +
fig, (ax1, ax2) = plt.subplots(1, 2, figsize = (15, 5))
sns.regplot(data = df_sm, x='Price_delta', y='Retweet_scaled', lowess=False, ax=ax1). set(title='Stock Price vs. No. of retweets')
sns.regplot(data = df_sm, x='Price_delta', y='Favorite_scaled', lowess=False, ax=ax2). set(title='Stock Price vs. No. of likes')
plt.tight_layout()
# +
X = df_sm['Price_delta']
y = df_sm['Polarity_delta']
atm_const = sm.add_constant(X)
mod = sm.OLS(y, atm_const)
res = mod.fit()
print(res.summary())
# +
X = df_sm['Polarity_delta']
y = df_sm['Price_delta']
atm_const = sm.add_constant(X)
mod = sm.OLS(y, atm_const)
res = mod.fit()
print(res.summary())
# +
X = df_sm[['Favorite_scaled']]
y = df_sm['Price_delta']
atm_const = sm.add_constant(X)
mod = sm.OLS(y, atm_const)
res = mod.fit()
print(res.summary())
# +
X = df_sm[['Retweet_scaled']]
y = df_sm['Price_delta']
atm_const = sm.add_constant(X)
mod = sm.OLS(y, atm_const)
res = mod.fit()
print(res.summary())
# +
X = df_sm[['Retweet_scaled', 'Polarity_delta', 'Favorite_scaled']]
y = df_sm['Price_delta']
atm_const = sm.add_constant(X)
mod = sm.OLS(y, atm_const)
res = mod.fit()
print(res.summary())
# + tags=[]
X = df_sm[['Retweet_scaled', 'Polarity_delta', 'Favorite_scaled']]
y = df_sm['Price_delta']
atm_const = sm.add_constant(X)
mod = sm.OLS(y, atm_const)
res = mod.fit()
print(res.summary())
# + tags=[]
X = df_sm[['Polarity_delta', 'Favorite_scaled']]
y = df_sm['Price_delta']
atm_const = sm.add_constant(X)
mod = sm.OLS(y, atm_const)
res = mod.fit()
print(res.summary())
# -
| notebooks/data_analysis_branson.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
boston=np.loadtxt("/Users/anura_000/bostontrain.csv",delimiter=',')
test=np.loadtxt("/Users/anura_000/bostontest.csv",delimiter=',')
# +
X=boston[:,:13]
Y=boston[:,13]
# +
import xgboost as xgb
# Instantiate the XGBRegressor: xg_reg
xg_reg = xgb.XGBRegressor(objective="reg:linear", n_estimators=265, seed=123,max_depth=4)
# Fit the regressor to the training set
xg_reg.fit(X,Y)
# Predict the labels of the test set: preds
preds = xg_reg.predict(test)
np.savetxt("Michegin.csv",preds,fmt='%1.5f')
preds
# -
np.savetxt("Michegin.csv",preds,fmt='%1.5f')
preds
| Project Gradient Descent/Gradient Descent - Boston Dataset/Boston Project with xGboost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Network Data Science with NetworkX and Python
# ## Load graphs from Excel spreadsheet files
import pandas as pd
link = ("https://github.com/dnllvrvz/Social-Network-Dataset/"
"raw/master/Social%20Network%20Dataset.xlsx")
network_data = pd.read_excel(link, sheet_name=['Elements', 'Connections'])
elements_data = network_data['Elements'] # node list
connections_data = network_data['Connections'] # edge list
import networkx as nx
# +
edge_cols = ['Type', 'Weight', 'When']
graph = nx.convert_matrix.from_pandas_edgelist(connections_data,
source='From',
target='To',
edge_attr=edge_cols)
# -
from random import sample
sampled_edges = sample(graph.edges, 10)
graph.edges[sampled_edges[0]]
node_dict = elements_data.set_index('Label').to_dict(orient='index')
nx.set_node_attributes(graph, node_dict)
sampled_nodes = sample(graph.nodes, 10)
graph.nodes[sampled_nodes[0]]
| implementing-graph-theory-with-NetworkX-and-Python/Load graphs from Excel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Amazon, EDA
# ## Importar
# # %load basic
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
pd.set_option('display.max_columns',200)
pd.options.display.float_format = '{:.2f}'.format
liga = '/home/ef/Documents/Diplomado/Amazon/data/csv/'
import time
start = time.time()
import os
os.listdir(liga)
# +
df = pd.DataFrame()
#Consolidar tablas en una
for x in [f for f in os.listdir(liga) if f.endswith('.csv')]:
df = df.append(pd.read_csv(liga + x, low_memory = False), ignore_index = True)
df.sample(4)
# -
# ## Funciones
import math
#Tiempo en minutos y segundos
def time_exp(x):
print(str(int(math.floor(x/60))
) + " minutos con " + '{:.2f}'.format(60*(x/60 - math.floor(x/60))
) + " segundos")
def fechas(df, col, formato = "%m %d, %Y"):
df = df.copy()
df[col] = pd.to_datetime(df[col], format = formato)
df['Anio'] = df[col].dt.year
meses = ['ene','feb','mar','abr','may','jun',
'jul','ago','sep','oct','nov','dic']
meses = dict(zip(range(1,13),meses))
#Mes en formato MMM (español)
df['Mes'] = df[col].dt.month.replace(meses)
df['Sem'] = df[col].dt.isocalendar().week
diasem = ['lun','mar','mie','jue','vie','sab','dom']
diasem = dict(zip(range(7),diasem))
#Día de la semana en formato DDD
df['DiaSem'] = df[col].dt.dayofweek.replace(diasem)
display(df['Anio'].hist())
return df, meses, diasem
# +
'''
#Activar si es la primera vez que se ocupan stopwords o Lemmatizer
import nltk
nltk.download('stopwords')
nltk.download('wordnet')
'''
import re
import unicodedata
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer
lem = WordNetLemmatizer()
def clean_text(text,language = 'english',
add_stopwords = [],
pattern ='[^a-zA-Z]'):
cleaned_text = unicodedata.normalize('NFD', str(text)).encode('ascii', 'ignore')
cleaned_text = re.sub(pattern, ' ',
cleaned_text.decode('utf-8'),
flags = re.UNICODE)
cleaned_text = ' '.join([lem.lemmatize(word) for word in
cleaned_text.lower().split() if word not in
stopwords.words(language) + add_stopwords])
return cleaned_text
# +
from nltk import sent_tokenize
from nltk.sentiment.vader import SentimentIntensityAnalyzer
def sentiment(text):
sid = SentimentIntensityAnalyzer()
try:
list(map(sid.polarity_scores,sent_tokenize(text)))[0]
except:
sentences = sent_tokenize('a')
else:
sentences = sent_tokenize(text)
res = map(sid.polarity_scores, sentences)
return list(res)[0]
# -
# ## EDA
# ### Fechas
df, meses, diasem = fechas(df,'reviewTime')
# +
from datetime import datetime
df['unixReviewTime'] = [datetime.utcfromtimestamp(ts).strftime('%Y-%m-%d') for ts in df['unixReviewTime']]
df['unixReviewTime'] = pd.to_datetime(df['unixReviewTime'])
df['unixReviewTime'] -= df['reviewTime']
#No hay diferencia entre reviewTime y unixReviewTime
df['unixReviewTime'].value_counts(1)
# -
# ### Style
df['style'].value_counts(1, dropna = False)
# +
#Se intentará evaluar cada registro
aux = []
for x in df['style']:
try:
#Dado que es un string que representa un diccionario
eval(x)
except:
#En caso de encontrar nulos, crear un diccionario arbitrario
aux.append({'Vacio':'Sí'})
else:
aux.append(eval(x))
#Lista de diccionarios se convierte a DataFrame
aux = pd.DataFrame(aux)
aux.sample(4)
# -
#Son columnas poco pobladas, se procede a omitirlas, con excepción de la primera
for col in aux.columns:
display(aux[col].value_counts(1,dropna = False).reset_index())
df['style'] = aux.iloc[:,:1].fillna('No_especifica')
df['style'].value_counts(1, dropna = False)
# ### Images
eval(df[df['image'].notnull()]['image'].reset_index(drop = True)[22])[0]
#Se guarda el dataset con link de imágenes para trabajarlo en el módulo IV
image_dataset = df[df['image'].notnull()].copy()
image_dataset['image'] = list(map(lambda x:eval(x)[0],image_dataset['image']))
image_dataset.to_csv(liga + 'images/image_dataset.csv', index = False)
df = df[df['image'].isnull()].reset_index(drop = True).copy()
len(df)
# ### Review
df['reviewText'] += " " + df['summary']
# +
import concurrent.futures
texto = 'reviewText'
with concurrent.futures.ProcessPoolExecutor() as executor:
limpio = list(executor.map(clean_text, df[texto]))
df[f'{texto}_limpio'] = list(map(lambda x:str(x),limpio))
df[f'{texto}_long'] = df[texto].str.len()
df[f'{texto}_n_words'] = df[texto].str.split().str.len()
df[f'{texto}_relevant'] = (df[f'{texto}_limpio'].str.len()+1) / (df[texto].str.len()+1)
df[[texto,f'{texto}_limpio',f'{texto}_long',
f'{texto}_n_words',f'{texto}_relevant']].sample(7)
# -
df.to_csv(liga[:-4] + 'df.csv', index = False)
# +
import concurrent.futures
texto = 'reviewText_limpio'
with concurrent.futures.ProcessPoolExecutor() as executor:
sentim = pd.DataFrame(list(executor.map(sentiment,
df[texto])))
sentim.sample(4)
# -
sentim.to_csv(liga[:-4] + 'sentim.csv', index = False)
df = df.join(sentim)
df.drop(columns = ['reviewerID','asin','reviewerName',
'unixReviewTime','vote', 'image'],
inplace = True)
df.sample(4)
df.dropna().to_csv(liga[:-4] + 'df_sentim.csv', index = False)
# ## Fin
#Tiempo total para correr el notebook
end = time.time()
time_exp(end - start)
# +
#Tono para cuando termina el script
from IPython.lib.display import Audio
import numpy as np
framerate = 4410
play_time_seconds = 1
t = np.linspace(0, play_time_seconds, framerate*play_time_seconds)
audio_data = np.sin(5*np.pi*300*t) + np.sin(2*np.pi*240*t)
#La siguiente línea suena!
Audio(audio_data, rate=framerate, autoplay=True)
| notebooks/01_Ingenieria_variables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
from astropy.io import ascii
from astropy.io import fits
# %load_ext autoreload
# %autoreload 2
from matplotlib import rc
rc("font", **{"family": "serif", "serif": ["Times"]})
rc("text", usetex=True)
# Constants and definitions:
lmc_distance = 50e3
def Absolute_to_Apparent(AbsoluteMag, distance):
return AbsoluteMag + 5 * (np.log10(distance / 10))
# -
# ### Note!! To run this you'll need a series of CSVs and fits files. These are too large for github but can be found here:
#
# https://drive.google.com/drive/folders/1eM0v4pp5YE5u7wSn2JVbjbulShM0mblW
#
# #### Unless you need to rerun reduction and dust extinction you can also jump down a few cells.
# ## Read in and Prepare the Data:
# +
lmc = pd.read_csv("Data/LMC_combined_1_1.csv")
###########
## Reduce #
###########
print(f"Total LMC Sources: {lmc.shape[0]}")
# Drop Nans
lmc = lmc.dropna(
subset=[
"uvw2_mag",
"uvw2_mag_err",
"uvm2_mag",
"uvm2_mag_err",
"uvw1_mag",
"uvw1_mag_err",
"Umag",
"e_Umag",
"Bmag",
"e_Bmag",
"Vmag",
"e_Vmag",
]
)
print(f"After Dropping Nans LMC Sources: {lmc.shape[0]}")
# Saturated Flag
lmc = lmc[
(lmc.uvm2_saturated == 0) & (lmc.uvw1_saturated == 0) & (lmc.uvw2_saturated == 0)
]
print(f"Keeping Saturated Flag = 0 LMC Sources: {lmc.shape[0]}")
# SSS Flag
lmc = lmc[(lmc.uvm2_sss == 1.0) & (lmc.uvw1_sss == 1.0) & (lmc.uvw2_sss == 1.0)]
print(f"Keeping SSS Flag = 1 LMC Sources: {lmc.shape[0]}")
# Edge Flag
lmc = lmc[(lmc.uvm2_edge == 1.0) & (lmc.uvw1_edge == 1.0) & (lmc.uvw2_edge == 1.0)]
print(f"Keeping Edge Flag = 1 LMC Sources: {lmc.shape[0]}")
# Residual Frac Error
max_resid_error = 0.5
lmc = lmc[
(np.abs(lmc.uvm2_resid_frac) < max_resid_error)
& (np.abs(lmc.uvw1_resid_frac) < max_resid_error)
& (np.abs(lmc.uvw2_resid_frac) < max_resid_error)
]
print(f"Keeping Max Residaul Under {max_resid_error} LMC Sources: {lmc.shape[0]}")
# UVOT - Mag Error
max_mag_error = 0.25
lmc = lmc[
(lmc.uvm2_mag_err < max_mag_error)
& (lmc.uvw1_mag_err < max_mag_error)
& (lmc.uvw2_mag_err < max_mag_error)
]
print(f"Keeping Max UVOT Mag Error Under {max_mag_error} LMC Sources: {lmc.shape[0]}")
# Optical - Mag Error
max_emag = 0.25
lmc = lmc[
(lmc.e_Umag < max_mag_error)
& (lmc.e_Bmag < max_mag_error)
& (lmc.e_Vmag < max_mag_error)
& (lmc.e_Imag < max_mag_error)
]
print(f"Remaining LMC Sources: {lmc.shape[0]}")
# Drop sources base on colors:
lmc = lmc.drop(lmc[lmc["uvw1_mag"] - lmc["Umag"] < -3].index).reset_index(drop=True)
lmc = lmc.drop(lmc[lmc["uvw1_mag"] - lmc["Umag"] > 3].index).reset_index(drop=True)
lmc = lmc.drop(lmc[lmc["Umag"] - lmc["Bmag"] < -2].index).reset_index(drop=True)
lmc = lmc.drop(lmc[lmc["Umag"] - lmc["Bmag"] > 2].index).reset_index(drop=True)
lmc = lmc.drop(lmc[lmc["Bmag"] - lmc["Vmag"] < -1].index).reset_index(drop=True)
lmc = lmc.drop(lmc[lmc["Bmag"] - lmc["Vmag"] > 1].index).reset_index(drop=True)
lmc = lmc.drop(lmc[lmc["Vmag"] - lmc["Imag"] < -1].index).reset_index(drop=True)
lmc = lmc.drop(lmc[lmc["Vmag"] - lmc["Imag"] > 1].index).reset_index(drop=True)
print(f"Remaining LMC Sources: {lmc.shape[0]}")
lmc.to_csv("Data/LMC_Reduced.csv", index=False)
# +
###################
# Dust Correction #
###################
import Dust
lmc_corr = Dust.DustCorrection("Data/LMC_Reduced.csv")
# -
# ## ** Start here unless you need to rerun reduction or dust
# +
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
from astropy.io import ascii
from astropy.io import fits
# %load_ext autoreload
# %autoreload 2
from matplotlib import rc
rc("font", **{"family": "serif", "serif": ["Times"]})
rc("text", usetex=True)
# Constants and definitions:
lmc_distance = 50e3
def Absolute_to_Apparent(AbsoluteMag, distance):
return AbsoluteMag + 5 * (np.log10(distance / 10))
# +
###############
# M o d e l s #
###############
# These are all in absolute magnitude.
zams_lmc = ascii.read("Data/ZAMS_Z0.006_Vegamag.txt")
she_lmc = ascii.read("Data/stripped_stars_Z0.006_Vegamag.txt")
zams_lmc_uvm2 = Absolute_to_Apparent(zams_lmc["UVM2_spec"], lmc_distance)
zams_lmc_v = Absolute_to_Apparent(zams_lmc["V_spec"], lmc_distance)
zams_lmcAB = ascii.read("Data/ZAMS_Z0.006_ABmag.txt")
she_lmcAB = ascii.read("Data/stripped_stars_Z0.006_ABmag.txt")
zams_lmc_uvm2AB = Absolute_to_Apparent(zams_lmcAB["UVM2_spec"], lmc_distance)
zams_lmc_vAB = Absolute_to_Apparent(zams_lmcAB["V_spec"], lmc_distance)
she_lmc_uvm2AB = Absolute_to_Apparent(she_lmcAB["UVM2"], lmc_distance)
she_lmc_vAB = Absolute_to_Apparent(she_lmcAB["V"], lmc_distance)
# +
def get_colors(x, y, zams_uvm2, zams_v):
"""x is the uvm2-v color of data; y is uvm2"""
data_x = np.array(x)
data_y = np.array(y)
curve_x = np.array(zams_uvm2) - np.array(zams_v)
curve_y = np.array(zams_uvm2)
# Interpolate the ZAMS to the y (uvm2) value of the data
zams_color_ref = np.interp(data_y, np.flip(curve_y, 0), np.flip(curve_x, 0))
m_map = []
sizes = []
alphas = []
for i in range(len(x)):
if x[i] < zams_color_ref[i]:
# c = "#0099b5"
c = np.array([13, 193, 231]) / 255.0
s = 5
if x[i] >= zams_color_ref[i]:
# c = "#061728"
c = np.array([9, 7, 52]) / 255.0
s = 10
m_map.append(c)
sizes.append(s)
return m_map, sizes
def get_blue(x, y, zams_uvm2, zams_v):
"""x is the uvm2-v color of data; y is uvm2"""
data_x = np.array(x)
data_y = np.array(y)
curve_x = np.array(zams_uvm2) - np.array(zams_v)
curve_y = np.array(zams_uvm2)
# Interpolate the ZAMS to the y (uvm2) value of the data
zams_color_ref = np.interp(data_y, np.flip(curve_y, 0), np.flip(curve_x, 0))
x_sub = []
y_sub = []
for i in range(len(x)):
if x[i] < zams_color_ref[i]:
x_sub.append(x[i])
y_sub.append(y[i])
return x_sub, y_sub
def get_red(x, y, zams_uvm2, zams_v):
"""x is the uvm2-v color of data; y is uvm2"""
data_x = np.array(x)
data_y = np.array(y)
curve_x = np.array(zams_uvm2) - np.array(zams_v)
curve_y = np.array(zams_uvm2)
# Interpolate the ZAMS to the y (uvm2) value of the data
zams_color_ref = np.interp(data_y, np.flip(curve_y, 0), np.flip(curve_x, 0))
x_sub = []
y_sub = []
for i in range(len(x)):
if x[i] >= zams_color_ref[i]:
x_sub.append(x[i])
y_sub.append(y[i])
return x_sub, y_sub
import matplotlib.colors as colors
from scipy import stats
def truncate_colormap(cmap, minval=0.0, maxval=1.0, n=100):
new_cmap = colors.LinearSegmentedColormap.from_list(
"trunc({n},{a:.2f},{b:.2f})".format(n=cmap.name, a=minval, b=maxval),
cmap(np.linspace(minval, maxval, n)),
)
return new_cmap
# -
# ## Actual CMD PLOTS:
# +
#########
# L M C #
#########
# Read in Data:
l = pd.read_csv("Data/LMC_Reduced_DeRed.csv")
lmc_mean_extinction = 0.25 # 0.44
# Define Axes
l_uvm2 = l["uvm2_mag"] + 1.69 - 3.07 * lmc_mean_extinction
l_v = l["vmag"] + 0.02 - lmc_mean_extinction
l_uvm2_v = l_uvm2 - l_v
# Separate based on line:
lx_cand, ly_cand = get_blue(l_uvm2_v, l_uvm2, zams_lmc_uvm2AB, zams_lmc_vAB)
lx_ms, ly_ms = get_red(l_uvm2_v, l_uvm2, zams_lmc_uvm2AB, zams_lmc_vAB)
# Create Main sequence density (Top Cat styling):
lxy_ms = np.vstack([lx_ms, ly_ms])
lz_ms = stats.gaussian_kde(lxy_ms)(lxy_ms)
lidx_ms = lz_ms.argsort()
lx_ms, ly_ms, lz_ms = (
np.array(lx_ms)[lidx_ms],
np.array(ly_ms)[lidx_ms],
np.array(lz_ms)[lidx_ms],
)
# +
# Stellar Data:
iso_path = "Data/LMC_Isolated_2019b.csv"
# iso_path = 'LMC_Isolated_2019.csv'
iso = pd.read_csv(iso_path)
#iso = iso[iso["name"] != "lmc-obs45578-835"]
iso_uvm2 = iso["uvm2_mag"] + 1.69 - 3.07 * iso["Av"]
iso_v = iso["Vmag"] + 0.02 - iso["Av"]
iso_uvm2_v = iso_uvm2 - iso_v
wn_path = "Data/LMC_WN3O3_2019.csv"
wn = pd.read_csv(wn_path, comment=";")
wn_uvm2 = wn["uvm2_mag"] + 1.69 - 3.07 * 3.1 * wn["E(B-V)"]
wn_v = wn["Vmag"] + 0.02 - 3.1 * wn["E(B-V)"]
wn_uvm2_v = wn_uvm2 - wn_v
wr_path = "Data/LMC_WR_2019b.csv"
wr = pd.read_csv(wr_path, comment=";")
wr_uvm2 = wr["uvm2_mag"] + 1.69 - 3.07 * 3.1 * wr["E(B-V)"]
wr_v = wr["Vmag"] + 0.02 - 3.1 * wr["E(B-V)"]
wr_uvm2_v = wr_uvm2 - wr_v
# -
iso_uvm2.iloc[[1,5]]
iso
# +
##LABELS FOR STRIPPED STAR MASSES.
start = 10
progenitor_lmc_x = [
(she_lmc_uvm2AB - she_lmc_vAB)[i] for i in range(start, len(she_lmc["UVM2"]), 3)
]
progenitor_lmc_y = [she_lmc_uvm2AB[i] for i in range(start, len(she_lmc["UVM2"]), 3)]
progenitor_lmc_label = [
she_lmc["Mstrip"][i] for i in range(start, len(she_lmc["UVM2"]), 3)
]
progenitor_lmc_label = np.array(progenitor_lmc_label).astype(str)
# +
##Background/Colors:
labelsize = 26
ticksize = 20
textsize = 20
legendsize = 18
ypad = 15
titlesize = 40
progenitor_sep = -0.25
progenitorsize = 15
sky2 = np.array([84, 107, 171]) / 255.0 # color of stripped stars
blue2 = np.array([9, 7, 52]) / 255.0 # Color of other stars
blue1 = np.array([13, 193, 231]) / 255.0 # Background for UV excess (apply alpha=0.03)
violet2 = np.array([161, 25, 107]) / 255.0 # models
royalblue = "#0000ff"
# xcand "#0099b5"
purple1 = np.array([54, 27, 77]) / 255
purple2 = "#32e300" # np.array([164,162,197])/255
lightblue = np.array([88, 149, 169]) / 255
darkblue = np.array([51, 73, 110]) / 255
mediumblue = np.array([164, 162, 197]) / 255 # np.array([51,73,110])*2/255
# 244,250,252
# 88,149,169
# 51,73,110
# targetcolor = "#00f9ff"
# targetcolor = "#0900ff"
# targetcolor="#4287f5"
# targetcolor="#EB9605"
# targetcolor="#fd9937"
# targetcolor="#ccccff"
# targetcolor="#340089"
# targetcolor="#4682B4"
# targetcolor="#4B0082"
targetcolor = "#7575cf"
# +
# Set plot:
f, (axes, bx) = plt.subplots(
2, 1, figsize=(10, 15), gridspec_kw={"height_ratios": [3, 1]}
)
sns.set(style="whitegrid", font="serif")
# rc('font',**{'family':'serif','serif':['Times']})
# rc('text', usetex=True)
# All Data:
plot_all = True
if plot_all:
# More filtering:
index_l = np.where(np.array(lx_cand) > -1.7)[0]
axes.scatter(
lx_ms,
ly_ms,
c=lz_ms,
zorder=0,
s=10,
edgecolor="",
cmap=truncate_colormap(plt.get_cmap("Greys"), 0.55, 0.95),
label="",
)
axes.scatter(
np.array(lx_cand)[index_l],
np.array(ly_cand)[index_l],
color=sky2,
zorder=0,
s=5,
label="",
alpha=0.5,
)
# Shading
axes.fill_betweenx(
zams_lmc_uvm2AB,
np.repeat(-3, len(zams_lmc_uvm2AB)),
zams_lmc_uvm2AB - zams_lmc_vAB,
alpha=0.4,
color="#e8fafc",
)
# ZAMS:
axes.plot(
zams_lmc_uvm2AB - zams_lmc_vAB,
zams_lmc_uvm2AB,
linewidth=5,
color="darkgray",
zorder=0,
label="",
)
# Stripped Stars Model:
axes.plot(
she_lmc_uvm2AB - she_lmc_vAB,
she_lmc_uvm2AB,
linewidth=5,
ms=10,
marker="o",
linestyle="-",
label="Stripped Helium Star Models",
zorder=1,
color=purple1,
)
[
axes.text(
x + progenitor_sep,
y,
"%.1f" % float(string),
fontweight="bold",
fontsize=progenitorsize,
)
for x, y, string in zip(progenitor_lmc_x, progenitor_lmc_y, progenitor_lmc_label)
]
# Chandra Isolated Targets:
axes.scatter(
iso_uvm2_v.iloc[[0,2,3,4]],
iso_uvm2.iloc[[0,2,3,4]],
marker="o",
edgecolor="black",
color=targetcolor,
s=250,
label="Chandra Targets",
zorder=3,
)
# Chandra Isolated Targets with XMM Detection:
axes.scatter(
iso_uvm2_v.iloc[[1,5]],
iso_uvm2.iloc[[1,5]],
marker="*",
edgecolor="black",
color=targetcolor,
s=400,
label="XMM Detections",
zorder=4,
)
# WN3/O3:
axes.scatter(
wn_uvm2_v,
wn_uvm2,
marker="D",
color=mediumblue,
edgecolor="black",
s=150,
zorder=1,
)
# WR LMC:
axes.scatter(
wr_uvm2_v,
wr_uvm2,
marker="D",
color=mediumblue,
edgecolor="black",
s=150,
zorder=1,
label="Wolf Rayet Stars",
)
axes.text(
progenitor_lmc_x[-1] - 0.3,
progenitor_lmc_y[-1] - 0.15,
r"$M_{\mathrm{strip}} [M_{\odot}]$",
fontweight="bold",
fontsize=progenitorsize + 4,
)
# Legend and Clean up:
axes.text(
-1.65,
13.5,
"ZAMS",
fontsize=textsize,
zorder=1,
weight="bold",
rotation=-85,
color="darkgray",
)
legend = axes.legend(prop={"size": legendsize, "weight": "bold"})
legend.set_title("Observations in the LMC", prop={"size": textsize})
axes.xaxis.label.set_size(labelsize)
axes.set_xlabel("UVM2 - V [AB mag]")
axes.yaxis.label.set_size(labelsize)
axes.set_ylabel("UVM2 [AB mag]", labelpad=ypad)
axes.grid(False)
axes.set_xlim(-2.0, 2)
axes.set_ylim(19.0, 13)
axes.set_xticks([-2.0, -1.0, 0.0, 1.0, 2.0])
axes.set_xticklabels([-2.0, -1.0, 0.0, 1.0, 2.0], fontsize=ticksize)
axes.set_yticks([19, 18, 17, 16, 15, 14, 13, 12.9])
axes.set_yticklabels([19, 18, 17, 16, 15, 14, 13], fontsize=ticksize)
# Spectra
lmc4349 = ascii.read("Data/New_Spectra/lmc-obs45516-4349_smooth.txt")
lmc156 = ascii.read("Data/New_Spectra/lmc-obs45510-156_smooth.txt")
lmc2273 = ascii.read("Data/New_Spectra/lmc-obs45461-2273_smooth.txt")
lmc335 = ascii.read("Data/New_Spectra/lmc-obs45536-335_smooth.txt")
lmc206 = ascii.read("Data/New_Spectra/lmc-obs45446-206_smooth.txt")
spec = lmc335
spec2 = lmc2273
bx.plot(spec["col1"], spec["col2"] + 0.2, c=targetcolor)
bx.plot(spec2["col1"], spec2["col2"] - 0.2, c=targetcolor)
bx.set_xlim(3750, 5000)
bx.set_ylim(0.5, 1.4)
waves = [3797, 3889, 3969, 4101+2, 4201, 4339, 4542, 4686, 4860]
wavenames = [
"HI-3797",
r"HeII-3889",
r"HeII-3969",
r"HeII-4101",
"HeII-4201",
r"HeII-4339",
"HeII-4542",
"HeII-4686",
r"HeII-4860",
]
newwaves = [4057+.05,4604]
newwavenames = ["NIV-4057","NV-4604/20"
]
[bx.axvline(x=lam, c=mediumblue, ls="dashed", alpha=0.4) for lam in waves]
[bx.axvline(x=lam, c=mediumblue, ls="dashed", alpha=0.4) for lam in newwaves]
bx.axvline(x=4620, c=mediumblue, ls="dashed", alpha=0.4)
bx.axhline(y=1, c=mediumblue, alpha=0.1)
bx.xaxis.label.set_size(labelsize)
bx.yaxis.label.set_size(labelsize)
bx.patch.set_facecolor("#e8fafc")
bx.patch.set_alpha(0.4)
bx.set_xlabel(r"Wavelength [$\AA$] ")
bx.grid(False)
# bx.set_ylabel("Normalized Intensity")
bx.text(4600, 1.3, "lmc-obs45536-335", fontsize=ticksize)
bx.text(4600, 0.9, "lmc-obs45461-2273", fontsize=ticksize)
# bx.text(4900, 1.3, "M01", fontsize=ticksize)
# bx.text(4900, 0.95, "M04", fontsize=ticksize)
# Last adjustments
adjust = [0,0,0,0,0,0,0,0,0,0]
adjust[3] = 10
[bx.text(w+a, 1.41, n, rotation=27, fontsize=ticksize-5)
for w, n,a in zip(waves, wavenames,adjust)]
[bx.text(w, 1.41, n, rotation=27, fontsize=ticksize-5)
for w, n in zip(newwaves, newwavenames)]
bx.set_xticks([3800, 4000, 4200, 4400, 4600, 4800, 5000])
bx.set_xticklabels([3800, 4000, 4200, 4400, 4600,
4800, 5000], fontsize=ticksize)
bx.set_yticks([0.6, 0.8, 1., 1.2, 1.4])
bx.set_yticklabels([0.6, 0.8, 1., 1.2, 1.4], fontsize=ticksize)
# plt.suptitle("Observations in the Magellanic Clouds",size=titlesize,x=0.5,y=0.92,fontweight="bold")
plt.subplots_adjust(top=0.85, hspace=0.29)
plt.savefig("cmd.png", dpi=500)
plt.show()
# -
adjust[0]
# +
lmc4349 = ascii.read("Data/New_Spectra/lmc-obs45516-4349_smooth.txt")
lmc156 = ascii.read("Data/New_Spectra/lmc-obs45510-156_smooth.txt")
lmc2273 = ascii.read("Data/New_Spectra/lmc-obs45461-2273_smooth.txt")
lmc335 = ascii.read("Data/New_Spectra/lmc-obs45536-335_smooth.txt")
lmc206 = ascii.read("Data/New_Spectra/lmc-obs45446-206_smooth.txt")
spec = [lmc4349, lmc156, lmc2273, lmc335, lmc206]
filenames = ["4349", "156", "2273", "335", "206"]
f, axes = plt.subplots(5, 1, figsize=(10, 20))
for ax, fname, s in zip(axes, filenames, spec):
ax.set_title(fname)
ax.plot(s["col1"], s["col2"])
ax.set_xlim(3750, 5000)
ax.set_ylim(0.5, 1.4)
plt.savefig("spectra.png")
# -
| CMD/CMD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Soft Robot Simulations
#
# ## <NAME>
# ## 1 April 2016
# + [markdown] slideshow={"slide_type": "slide"}
# # Problem
#
# * Simulate deformable robots making lots of contact with the world
# * Control those robots to do something dynamic and interesting
# * e.g. juggling a ball using a soft arm
#
# + [markdown] slideshow={"slide_type": "slide"}
# # A Possible Approach
#
# * Simulate deformable robots and contact with the world in a way that is easy to differentiate
# * Prefer informative derivatives to high accuracy
# * Use nifty nonlinear MPC techniques to try to generate and track trajectories
# + [markdown] slideshow={"slide_type": "slide"}
# # Some Inspiration
#
# Neunert et al., *Fast Nonlinear Model Predictive Control for Unified Trajectory Optimization and Tracking*
# + slideshow={"slide_type": "skip"}
from IPython.display import YouTubeVideo
# -
YouTubeVideo("Y7-1CBqs4x4")
# + [markdown] slideshow={"slide_type": "slide"}
# # Simulation
#
# * Model a robot as a network of point masses with damped springs connecting them
# * Perform indirect collision checking between robots:
# * Compute a barrier that tightly surrounds the robot, represented as an implicit function
# * Check for collisions between the points of one robot and the barriers of another
#
#
| presentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/scottslusher/Alphabet-Soup/blob/main/inventory_projection_fb_prophet.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="D-DzilsrMojU" outputId="95b5522e-d1fd-44f7-8ad7-6d68b4658e2e"
# !pip install pystan
# !pip install fbprophet
# + id="6HIajlMdM8BX"
# Import the required libraries and dependencies
import pandas as pd
from pathlib import Path
from fbprophet import Prophet
import datetime as dt
from datetime import datetime
# %matplotlib inline
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": "OK"}}, "base_uri": "https://localhost:8080/", "height": 73} id="265_S1nbM-8U" outputId="3f231347-340c-4a26-9cba-7a63885b1aa8"
from google.colab import files
uploaded = files.upload()
# + colab={"base_uri": "https://localhost:8080/"} id="WohE9UgRYZyh" outputId="e8b9c2d2-e997-4e18-82c6-262b7fe8a216"
df_orig = pd.read_csv('pivetal_latex_gloves.csv', parse_dates=True, infer_datetime_format=True)
df_orig.dropna(inplace=True)
item_list = df_orig['Item#'].unique()
print(df_orig.head())
print(item_list[:5])
# + colab={"base_uri": "https://localhost:8080/", "height": 780} id="4mxNtDyGNA1E" outputId="27dd56e7-6843-414b-e7fb-207754f046da"
finalized_list = []
for i in range(len(item_list)):
try:
item_num = item_list[i]
df = df_orig.loc[df_orig['Item#'] == item_list[i]]
df = df[['InvoiceDate', 'Qty']].copy()
df['InvoiceDate'] = pd.to_datetime(df['InvoiceDate'])
df_grouped = df.groupby(df.InvoiceDate).sum()
df_grouped.reset_index(inplace=True)
df_grouped.sort_values('InvoiceDate', inplace=True)
df_grouped['t_qty'] = df_grouped['Qty'].cumsum()
df = df_grouped[['InvoiceDate', 't_qty']].copy()
df_sliced = df.loc[df['InvoiceDate'] < '2020-03-01']
df_sliced.columns = ['ds', 'y']
model = Prophet()
model.fit(df_sliced)
future_trends = model.make_future_dataframe(periods=173, freq='D')
forecast_trends = model.predict(future_trends)
forecast_trends = forecast_trends.set_index(["ds"])
forecast = forecast_trends[["yhat", "yhat_lower", "yhat_upper"]].iloc[-173:,:]
estimated_total = forecast['yhat'][-1] - forecast['yhat'][0]
data = {"Item #": item_num, "Estimated 8 Month Qty":estimated_total}
finalized_list.append(data)
except:
pass
final_df = pd.DataFrame(finalized_list)
final_df
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="dX3u_znBNKhU" outputId="5684679b-fb08-4071-fd48-6631f4adf0e6"
df = df[['InvoiceDate', 'Qty']].copy()
df['InvoiceDate'] = pd.to_datetime(df['InvoiceDate'])
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="88F7rtbeNTjk" outputId="197d4301-ce93-43ea-a24f-2ecd66d47732"
df_grouped = df.groupby(df.InvoiceDate).sum()
df_grouped.reset_index(inplace=True)
df_grouped.sort_values('InvoiceDate', inplace=True)
df_grouped['t_qty'] = df_grouped['Qty'].cumsum()
df = df_grouped[['InvoiceDate', 't_qty']].copy()
df.tail()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="RUb4lJElSwZu" outputId="49bf9abd-ea84-4879-83f1-d25e27311dd0"
df_sliced = df.loc[df['InvoiceDate'] < '2020-03-01']
df_sliced.tail()
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="zEU1wuPxNbET" outputId="ddcfcbdc-77eb-447f-daa6-157ac75609e1"
df_sliced.columns = ['ds', 'y']
df_sliced
# + colab={"base_uri": "https://localhost:8080/"} id="_IvF7i6fNhUE" outputId="b639ebaa-4725-47c8-f849-12849a5a21a7"
df_sliced.shape
# + id="Xw9VA1qUNqGT"
model = Prophet()
# + colab={"base_uri": "https://localhost:8080/"} id="S9H5hHc4NvjF" outputId="ccd68118-4691-436e-e883-e1f9ffb76671"
model.fit(df_sliced)
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="Ws6FNbzaNwwV" outputId="a9a007f6-f18f-449b-b074-3cdad27db272"
future_trends = model.make_future_dataframe(periods=365, freq='D')
future_trends
# + colab={"base_uri": "https://localhost:8080/", "height": 661} id="4_0BNLzdN6I0" outputId="03bd1511-023e-4380-c209-90b035af83bd"
forecast_trends = model.predict(future_trends)
forecast_trends
# + colab={"base_uri": "https://localhost:8080/", "height": 357} id="V5yY4C4QOAcz" outputId="612f5e35-3998-44f5-f6c0-4dd7a7570907"
forecast_trends.tail()
# + colab={"base_uri": "https://localhost:8080/", "height": 865} id="oJW5aKeAOFdE" outputId="010049a5-fb7b-4b59-e7a3-0da39348ddc7"
model.plot(forecast_trends)
# + colab={"base_uri": "https://localhost:8080/", "height": 657} id="c09Ksz7LOIKU" outputId="efb74520-a081-4176-f153-726094f65f0a"
# Use the plot_components function to visualize the forecast results.
figures = model.plot_components(forecast_trends)
# + colab={"base_uri": "https://localhost:8080/", "height": 388} id="W0ZEsv2tOQ2F" outputId="05996348-8af4-4a3b-cc08-c4b8747cddda"
# At this point, it's useful to set the `datetime` index of the forecast data.
forecast_trends = forecast_trends.set_index(["ds"])
forecast_trends.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 310} id="tcAUPzhXOYR1" outputId="c3f754d8-81be-448a-a251-8db67dae9fd1"
# From the `forecast_trends` DataFrame, use hvPlot to visualize
# the yhat, yhat_lower, and yhat_upper columns over the last 10 days (24*10 = 240)
forecast = forecast_trends[["yhat", "yhat_lower", "yhat_upper"]].iloc[-173:,:]
forecast.plot()
# + colab={"base_uri": "https://localhost:8080/"} id="VM55b-nqOZvV" outputId="d0a70b7a-392b-4373-ee0e-e5d6e87483e4"
forecast = forecast_trends[["yhat", "yhat_lower", "yhat_upper"]].iloc[-173:,:]
estimated_total = forecast['yhat'][-1] - forecast['yhat'][0]
estimated_total
# + id="K-2wrxkhUuOV"
| inventory_projection_fb_prophet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
from thermof.interpenetration import *
# -
# ### Path for the lammps structure file
data_dir = os.path.join(os.getcwd(), 'sample')
data_path = os.path.join(data_dir, 'lammps_single.data.sample')
# ### Create interpenetrated structure
interpenetrate_lampps_data(data_path, output_dir=data_dir)
# ### Convert interpenetrated structure to xyz format
new_data_path = os.path.join(data_dir, 'lammps_ipmof.data')
lammps_data2xyz(new_data_path, output_dir=data_dir)
| notebooks/interpenetrate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Pandas Series
import pandas as pd
df = pd.read_csv("../data/500_Person_Gender_Height_Weight_Index.csv")
# ## Exploring data
# examine first few rows
df.head()
# examine last few rows
df.tail()
# shape of data
df.shape
# column
df.columns
# sampling
df.sample(10)
# info
df.info()
# summary stats
df.describe()
# check missing values
df.isnull().sum()
df["Weight"]
type(df["Weight"])
df["Weight"].equals(df.Weight)
weight = df["Weight"]
weight.head(2)
weight.tail()
weight.dtype
weight.shape
len(weight)
weight.index
weight.info()
weight.to_frame().info()
# ## Analyzing Numerical Series
weight
weight.describe()
weight.count()
weight.size
len(weight)
weight.sum()
weight.sum(skipna = False)
weight.sum(skipna = True)
sum(weight)
weight.mean()
weight.median()
weight.std()
weight.min()
weight.max()
weight.unique()
len(weight.unique())
weight.nunique()
weight.nunique(dropna = False)
weight.nunique(dropna = True)
weight.value_counts()
weight.value_counts(sort = True)
weight.value_counts(sort = False)
weight.value_counts(dropna = True)
weight.value_counts(dropna = False)
weight.value_counts(ascending = False)
weight.value_counts(ascending = True)
weight.value_counts(sort = True, dropna = True, ascending = False, normalize = False)
weight.value_counts(sort = True, dropna = True, ascending = False, normalize = True)
weight.value_counts(sort = True, dropna = True, ascending = False, normalize = True) * 100
weight.value_counts(sort = True, dropna = False, ascending = False, normalize = True)
weight.value_counts(sort = True, dropna = True, ascending= False, normalize = False, bins = 5)
weight.value_counts(sort = True, dropna = True, ascending= False, normalize = True, bins = 10)
weight.value_counts(sort = True, dropna = True, ascending= False, normalize = True, bins = 10) * 100
# ## Analyzing non-numerical Series
import pandas as pd
df = pd.read_excel("../data/LungCapData.xls")
df.head()
df.shape
df.info()
df['Gender'].unique()
df['Smoke'].unique()
df['Caesarean'].unique()
smoker = df["Smoke"]
smoker.head()
smoker.tail(5)
type(smoker)
smoker.dtype
smoker.shape
smoker.describe()
smoker.size
smoker.count()
smoker.min()
smoker.unique()
len(smoker.unique())
smoker.nunique(dropna= False)
smoker.value_counts()
smoker.value_counts(sort = True, ascending=True)
smoker.value_counts(sort = True, ascending=False, normalize = True)
smoker.value_counts(sort = True, ascending=False, normalize = True) * 100
# ## Sorting and introduction to the inplace-parameter
import pandas as pd
dic = {1:10, 3:25, 2:6, 4:36, 5:2, 6:0, 7:None}
dic
ages = pd.Series(dic)
ages
ages.sort_index()
x = ages.sort_index(ascending=False)
x
ages
ages.sort_index(ascending = False, inplace= True)
ages
ages.sort_values(inplace=False)
ages
ages.sort_values(ascending=False, na_position="last", inplace= True)
ages
dic = {"Mon":10, "Tue":25, "Wed":6, "Thu": 36, "Fri": 2}
dic
s = pd.Series(dic)
s
s.sort_index(ascending=False)
# ## nlargest() and nsmallest()
import pandas as pd
df = pd.read_csv("../data/diabetes.csv")
df.head()
age = df.Age
age.head()
age.sort_values(ascending=False).head(3)
age.sort_values(ascending=True).iloc[:3]
age.nlargest(n = 3).index[0]
age.nsmallest(n = 3).index[0]
# ## idxmin() and idxmax()
df.head()
df.Age.idxmax()
df.Age.idxmin()
df.loc[630]
df.loc[df.Age.idxmin()]
dic = {"Mon":10,"Tue":25, "Wed":6, "Thu":36, "Fri":2, "Sat":0, "Sun":None}
dic
sales = pd.Series(dic)
sales
sales.sort_values(ascending=True).index[0]
sales.idxmin()
sales.sort_values(ascending=False).index[0]
sales.idxmax()
# ### Manipulating Series
import pandas as pd
df = pd.read_csv("../data/500_Person_Gender_Height_Weight_Index.csv")
df.head()
height = df["Height"]
height.head()
height.tail()
height.iloc[1] = 30
height.head()
| 04 - Data Analysis With Pandas/notebooks/04_WorkingWithPandasSeries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Activity 6.02
# +
# Import the libraries
import numpy as np
import pandas as pd
from tensorflow import random
# Load the Data
X = pd.read_csv("../data/aps_failure_training_feats.csv")
y = pd.read_csv("../data/aps_failure_training_target.csv")
# -
# Split the data into training and testing sets
from sklearn.model_selection import train_test_split
seed = 42
X_train, X_test, y_train, y_test= train_test_split(X, y, test_size=0.20, random_state=seed)
# +
# Initialize StandardScaler
from sklearn.preprocessing import StandardScaler
sc=StandardScaler()
# Transform the training data
X_train = sc.fit_transform(X_train)
X_train = pd.DataFrame(X_train,columns=X_test.columns)
# Transform the testing data
X_test=sc.transform(X_test)
X_test=pd.DataFrame(X_test,columns=X_train.columns)
# +
# Import the relevant Keras libraries
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Dropout
random.set_seed(seed)
np.random.seed(seed)
model = Sequential()
# Add the hidden dense layers and with dropout Layer
model.add(Dense(units=64, activation='relu', kernel_initializer='uniform', input_dim=X_train.shape[1]))
model.add(Dropout(rate=0.5))
model.add(Dense(units=32, activation='relu', kernel_initializer='uniform'))
model.add(Dropout(rate=0.4))
model.add(Dense(units=16, activation='relu', kernel_initializer='uniform'))
model.add(Dropout(rate=0.3))
model.add(Dense(units=8, activation='relu', kernel_initializer='uniform'))
model.add(Dropout(rate=0.2))
model.add(Dense(units=4, activation='relu', kernel_initializer='uniform'))
model.add(Dropout(rate=0.1))
# Add Output Dense Layer
model.add(Dense(units=1, activation='sigmoid', kernel_initializer='uniform'))
# Compile the Model
model.compile(optimizer='adam', loss='binary_crossentropy', metrics=['accuracy'])
# -
# Fit the Model
model.fit(X_train, y_train, epochs=100, batch_size=20, verbose=1, validation_split=0.2, shuffle=False)
y_pred = model.predict(X_test)
y_pred_prob = model.predict_proba(X_test)
# Calculate ROC curve
from sklearn.metrics import roc_curve
fpr, tpr, thresholds = roc_curve(y_test, y_pred_prob)
import matplotlib.pyplot as plt
plt.plot(fpr,tpr)
plt.title("ROC Curve for APS Failure")
plt.xlabel("False Positive rate (1-Specificity)")
plt.ylabel("True Positive rate (Sensitivity)")
plt.grid(True)
plt.show()
# Calculate AUC score
from sklearn.metrics import roc_auc_score
roc_auc_score(y_test, y_pred_prob)
| Chapter06/Activity6.02/Activity6_02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## File IO
#
# These notebooks are used to check certain examples in the course notes, hence they are mainly for the benefit of the course developers, rather than a teaching resource.
import numpy as np
import pandas as pd
# +
#Go to the home directory for the GitBook
# -
# cd /home/researcher/Python
# !head -3 "data/titanic.csv"
# !tail -3 "data/titanic.csv"
# ## Python's open function
#Python has a built-in function open() to open a file.
f = open("data/titanic.csv", mode = 'r')
print(type(f))
data = f.read() # read in all data until the end of file as a single string
print(type(data), len(data))
# we can use readline() method to read individual lines of a file. This method reads a file till the newline, including the newline character
f = open("data/titanic.csv", mode = 'r')
f.readline()
# The next time we run readline, it will start from where we left off
f.readline()
# As opposed to much text, where line breaks signify a geometric limitation, in the world of data, lines often have a a crirical significance (i.e an individual record).
#
# The following examples show how we could extract indivdual lines and store them in a list.
f = open("data/titanic.csv", mode = 'r')
#We can read a file line-by-line using a for loop.
lines = []
for line in f:
lines.append(f)
print(len(lines))
#Or use the readlines() method returns a list of remaining lines of the entire file.
f = open("data/titanic.csv", mode = 'r')
lines = f.readlines()
print(len(lines))
f = open("data/titanic.csv", mode = 'r')
for lineGuy in f:
print(line)
f.close()
# When we are done with operations to the file, we need to properly close it. Python has a garbage collector to clean up unreferenced objects. But we must not rely on it to close the file. Closing a file will free up the resources that were tied with the file and is done using the close() method.
f.close()
print(type(f))
# If an exception occurs when we are performing some operation with the file, the code exits without closing the file. he best way to do this is using the with statement. This ensures that the file is closed when the block inside with is exited.
with open("data/titanic.csv",encoding = 'utf-8') as f:
lines = f.readlines()
print(len(lines))
f.read()
# ## Numpy
# The last section showed how files can be read into Python as strings. What about when our data is numeric? In previous lessons we have looked at the case of importing .csv files with purely numeric data. In this case the numpy methods `np.loadtxt` or `np.genfromtxt` can easily turn our data in a numer numpy array. `np.genfromtxt` is a more robust version of loadtxt that can better handle missing data.
#
# If our csv file contains both characters and numbers, as in the current example, the situation is a little more challenging.
#
# First, let's try the naive approach:
data = np.genfromtxt("data/titanic.csv", delimiter=",")
# This failed because the column headers have a 11 rows while the data appears to have 12 rows...have a think about why that is.
#
# If we ignored the header we can succesfully import the data
data = np.genfromtxt("data/titanic.csv", delimiter=",", skip_header=1)
data.shape
data[0, :]
# Unfortunately this is still not that a great solution. Our strings have become nans and our array has 12 colums because it interpreted the comma inside the name field as a column delimiter. without resorting to even more complexity, it may be easiest at this stage to tell `np.genfromtxt` exactly which columns we want to use, thereby creating a purely numeric array:
# !head -2 "data/titanic.csv"
data = np.genfromtxt("data/titanic.csv", delimiter=",", skip_header=1, usecols=tuple((0,1,5,6,7,8,9)))
data[0,:]
# This is fine, and if you definately want to use the numpy package, it may be the easiest way to get just the numeric data out of a delimited plain txt file (i.e a csv ).
#
# But if your data consists of both number and characters, there is a more applicable package available in Python...
# ## Pandas
# For .read_csv, always use header=0 when you know row 0 is the header row
df = pd.read_csv("data/titanic.csv", header=0)
print(df.shape, type(df))
# Okay, that seemed to work pretty well. Pandas was even clever enough to interpret the comma in the names field as a data value, rather than a delimiter. Well done Pandas.
#
# In the course we don't teach Pandas, but given we're here lets have a quick look at some out of the box tools.
df.info()
# There's a lot of useful info there! You can see immediately we have 418 entries (rows), and for most of the variables we have complete values (418 are non-null). But not for Age, or fare, Cabin -- those have nulls somewhere.
df.describe()
# This is also very useful: pandas has taken all of the numerical columns and quickly calculated the mean, std, minimum and maximum value. Convenient! But also a word of caution: we know there are a lot of missing values in Age, for example. How did pandas deal with that? It must have left out any nulls from the calculation. So if we start quoting the "average age on the Titanic" we need to caveat how we derived that number.
# %matplotlib inline
import matplotlib.pyplot as plt
df['Age'].dropna().hist(bins=16, range=(0,80), alpha = .5)
plt.show()
# If you're intersted in looking at more examples with this data, checkout this kaggle page: https://www.kaggle.com/c/titanic/details/getting-started-with-python-ii
| examples/fileIO.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Select Democrat and Republican states for case study
# ## using numpy random numbr generator
# +
# United States of America Python Dictionary to translate States,
# Districts & Territories to Two-Letter codes and vice versa.
#
# https://gist.github.com/rogerallen/1583593
#
# Dedicated to the public domain. To the extent possible under law,
# <NAME> has waived all copyright and related or neighboring
# rights to this code.
us_state_abbrev = {
'Alabama': 'AL',
'Alaska': 'AK',
#'American Samoa': 'AS',
'Arizona': 'AZ',
'Arkansas': 'AR',
'California': 'CA',
'Colorado': 'CO',
'Connecticut': 'CT',
'Delaware': 'DE',
'District of Columbia': 'DC',
'Florida': 'FL',
'Georgia': 'GA',
#'Guam': 'GU',
'Hawaii': 'HI',
'Idaho': 'ID',
'Illinois': 'IL',
'Indiana': 'IN',
'Iowa': 'IA',
'Kansas': 'KS',
'Kentucky': 'KY',
'Louisiana': 'LA',
'Maine': 'ME',
'Maryland': 'MD',
'Massachusetts': 'MA',
'Michigan': 'MI',
'Minnesota': 'MN',
'Mississippi': 'MS',
'Missouri': 'MO',
'Montana': 'MT',
'Nebraska': 'NE',
'Nevada': 'NV',
'New Hampshire': 'NH',
'New Jersey': 'NJ',
'New Mexico': 'NM',
'New York': 'NY',
'North Carolina': 'NC',
'North Dakota': 'ND',
#'Northern Mariana Islands':'MP',
'Ohio': 'OH',
'Oklahoma': 'OK',
'Oregon': 'OR',
'Pennsylvania': 'PA',
#'Puerto Rico': 'PR',
'Rhode Island': 'RI',
'South Carolina': 'SC',
'South Dakota': 'SD',
'Tennessee': 'TN',
'Texas': 'TX',
'Utah': 'UT',
'Vermont': 'VT',
#'Virgin Islands': 'VI',
'Virginia': 'VA',
'Washington': 'WA',
'West Virginia': 'WV',
'Wisconsin': 'WI',
'Wyoming': 'WY'
}
print(us_state_abbrev)
# -
# ### from https://www.bbc.com/news/election/us2020/results
#
# Solid Democratic: CA, NY, IL, NJ, VA, WA, MA, MN, CO, MD, OR, CT, NM, ME, NH, HI, RI, DE, VT, DC
#
# Solid Republican: TN, IN, MO, SC, AL, LA, KY, OK, UT, KS, MS, AR, NE, ID, WV, MT, SD, ND, AK, WY
#
democrat = ['CA', 'NY', 'IL', 'NJ', 'VA', 'WA', 'MA', 'MN', 'CO', 'MD', 'OR', 'CT', 'NM', 'ME', 'NH', 'HI', 'RI', 'DE', 'VT', 'DC']
republican = ['TN', 'IN', 'MO', 'SC', 'AL', 'LA', 'KY', 'OK', 'UT', 'KS', 'MS', 'AR', 'NE', 'ID', 'WV', 'MT', 'SD', 'ND', 'AK', 'WY']
# +
import numpy as np
np.random.seed(100)
dem_state = np.random.randint(0,len(democrat)+1)
print("Democrat State: "+ democrat[dem_state])
# +
np.random.seed(100)
rep_state = np.random.randint(0,len(republican)+1)
print("Republican State: "+ republican[rep_state])
# -
| data_analysis/case_studies/select_states.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="rTKvoFXUk3FN"
# ## Startup codes
#
# + [markdown] id="6Ex-M-kukB_8"
# ### Imports
# + colab={"base_uri": "https://localhost:8080/"} id="kLG28lX8RIq9" outputId="57053ae8-d072-4f54-8d2d-1d379ba6339d"
# %load_ext autotime
import warnings
warnings.filterwarnings('always')
warnings.simplefilter('always')
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
warnings.simplefilter(action='ignore', category=DeprecationWarning)
warnings.filterwarnings("ignore", category=DeprecationWarning)
warnings.filterwarnings("ignore", message="numpy.dtype size changed")
warnings.filterwarnings("ignore", message="numpy.ufunc size changed")
import glob, os, sys
# import matplotlib.pylab as plt
# from matplotlib import pyplot
import numpy as np
from sklearn.model_selection import StratifiedKFold
import tensorflow as tf
print(tf.__version__)
import pandas as pd
pd.options.mode.chained_assignment = None
import csv
import xml.etree.ElementTree as ET
import pickle
import math
import matplotlib.pyplot as plt
# %config Completer.use_jedi = False
from importlib import reload
# -
# ### Utility methods
# + colab={"base_uri": "https://localhost:8080/"} id="NCFF6nHC9TzM" outputId="9d7df00c-9f5c-4bf8-b59d-afb5b6987d4b"
def save_obj(obj, name ):
with open('pickle/'+ name + '.pkl', 'wb') as f:
pickle.dump(obj, f, pickle.HIGHEST_PROTOCOL)
print("Saved object to a file: %s" % (str(f)))
def load_obj(name ):
with open('pickle/' + name + '.pkl', 'rb') as f:
return pickle.load(f)
def save_df(df, file_name):
df.to_csv(file_name, index=False, quotechar='"', quoting=csv.QUOTE_NONNUMERIC)
import re
def get_tags(tags_str):
return re.findall(r'<(.+?)>', tags_str)
def make_link(id, type):
'''
id = postid
type : 'q' for question
'a' for answer
'''
url = f'https://stackoverflow.com/{type}/{id}'
return f'=HYPERLINK("{url}", "{id}")'
def linkToId(x):
'''
Takes a excel styled link and retures the QuestionID
'''
return int(x.split('"')[::-1][1])
COLS = ["Id", "PostTypeId", "AcceptedAnswerId", "ParentId", "CreationDate", "DeletionDate", "Score", "ViewCount", "Body",
"OwnerUserId", "OwnerDisplayName", "LastEditorUserId", "LastEditorDisplayName", "LastEditDate", "LastActivityDate",
"Title", "Tags", "AnswerCount", "CommentCount", "FavoriteCount", "ClosedDate", "CommunityOwnedDate", "ContentLicense"]
ROOT_DIR = '/'
# + [markdown] id="HB9FAjcAmdcn"
# ## Dataset filtering
# + [markdown] id="4vYOmSeg6QAW"
# #### Bootstrap
# + colab={"base_uri": "https://localhost:8080/"} id="KDy97QW9W_VY" outputId="f62ac00b-2ef3-4d22-ddd1-d5e85eb377fa"
TAG_list_initial_file = os.path.join( ROOT_DIR, "Data", "tag_list_initial.csv")
TAG_list_final_file = os.path.join( ROOT_DIR, "Data", "tag_list_final.csv")
QUESTIONS_initial_tag_file = os.path.join(ROOT_DIR, "Data", "questions_initial_tag.csv")
QUESTIONS_final_tag_file = os.path.join(ROOT_DIR, "Data", "questions_final_tag.csv")
QUESTIONS_nss_final_tag_file = os.path.join(ROOT_DIR, "Data", "questions_nss_final_tag.csv")
ANSWERS_nss_final_tag_file = os.path.join(ROOT_DIR, "Data", "answers_nss_final_tag.csv")
TAG_manual_relevance_file = os.path.join( ROOT_DIR, "Data", "tag_manual_relevance.csv")
# + colab={"base_uri": "https://localhost:8080/"} id="G15QyRdxswNr" outputId="2cbe92cd-1020-4d9e-d91a-a80e6424fa8d"
Tag_list_initial = None
df_questions_initial_tag = None
Tag_stat_Q_initial = {}
Tag_stat_Q_all = {}
Significance_tag = {}
Relevance_tag = {}
Tag_stat_manual_relevance = []
DF_tag_stat_summary = None
Tag_list_relevent = []
Tag_list_final = []
DF_question_final_tag = None
DF_answer_final_tag = None
DF_tag_stat_summary = None
DF_question_nss_final_tag = None
DF_answer_nss_final_tag = None
# + [markdown] id="Rt4P94GE54Uj"
# ##### Load all variables
# + colab={"base_uri": "https://localhost:8080/"} id="TZtUkTvd5ipI" outputId="dc6a9096-0bee-47f8-bdc0-f43da22b122c"
Tag_list_initial = load_obj("Tag_list_initial")
df_questions_initial_tag = load_obj("df_questions_initial_tag")
Tag_stat_Q_initial = load_obj("Tag_stat_Q_initial")
Tag_stat_Q_all = load_obj("Tag_stat_Q_all")
Significance_tag = load_obj("Significance_tag")
Relevance_tag = load_obj("Relevance_tag")
Tag_stat_manual_relevance = load_obj("Tag_stat_manual_relevance")
Tag_list_final = load_obj("Tag_list_final")
DF_question_final_tag = load_obj("DF_question_final_tag")
DF_answer_final_tag = load_obj("DF_answer_final_tag")
DF_question_nss_final_tag = load_obj("DF_question_nss_final_tag")
DF_answer_nss_final_tag = load_obj("DF_answer_nss_final_tag")
Map_tag_qids = load_obj("Map_tag_qids")
# DF_answer_final_tag = load_obj("DF_answer_final_tag")
# DF_question_nss_final_tag = load_obj("DF_question_nss_final_tag")
# DF_answer_nss_final_tag = load_obj("DF_answer_nss_final_tag")
print(len(Tag_list_initial))
print(len(df_questions_initial_tag))
print(len(Tag_stat_Q_initial))
print(len(Tag_stat_Q_all))
print(len(Significance_tag))
print(len(Relevance_tag))
print(len(Tag_stat_manual_relevance))
print(len(Tag_list_final))
print(len(DF_question_final_tag))
print(len(DF_answer_final_tag))
print(len(DF_question_nss_final_tag))
print(len(DF_answer_nss_final_tag))
# print(len(DF_answer_final_tag))
# print(len(DF_question_nss_final_tag))
# print(len(DF_answer_nss_final_tag))
# + [markdown] id="nDKPfV56RjeP"
# ##### Helper methods
# + colab={"base_uri": "https://localhost:8080/"} id="xVz0ThylRnO3" outputId="21fc1650-622c-4ddf-acac-c2d532c977da"
def get_initial_taglist(input_file):
df_initial_tag_list = pd.read_csv(input_file)
df_initial_tag_list = df_initial_tag_list['Tag'].dropna()
initial_tag_list = df_initial_tag_list.tolist()
return initial_tag_list
import lxml.etree as ET
def get_questions_from_tags(POSTS_file, TAGS):
context = ET.iterparse(POSTS_file, events=("end",))
print("Going to extract questions")
df_posts = pd.DataFrame(columns = COLS)
total_questions = 0
count = 0
_, root = next(context)
for event, elem in context:
# if (count > 5):
# break
if elem.tag == "row":
tags = elem.attrib.get('Tags', '')
tags_list = get_tags(tags) # list of tags
for tag in TAGS:
if tag in tags_list:
dic = {}
for col in COLS:
dic[col] = elem.attrib.get(col, '')
# data.append(elem.attrib.get(col, ''))
df_posts = df_posts.append(pd.Series(dic), ignore_index = True)
continue
# progress
if total_questions % 1000000 == 0:
print('done', elem.attrib['Id'])
elem.clear()
root.clear()
total_questions += 1
# count += 1
df_posts.drop_duplicates('Id',inplace=True)
return df_posts
# + [markdown] id="cHGybFuhTdJq"
# ### Extract Initial analysis
# + [markdown] id="WGVnQm13Q2uu"
# #### Get Initial taglist
# + colab={"base_uri": "https://localhost:8080/"} id="J94yWPaSmxdy" outputId="3995667c-6ce6-470f-ac4a-7525438dac54"
Tag_list_initial = get_initial_taglist(TAG_list_initial_file)
print(len(Tag_list_initial))
print(Tag_list_initial)
# + colab={"base_uri": "https://localhost:8080/"} id="AMWdgSWp5xNV" outputId="b3ff3e31-5ceb-4cde-8a70-448b6fc02005"
save_obj(Tag_list_initial, "Tag_list_initial")
# + [markdown] id="E-yiiFR5RYWH"
# #### Extract Initial Qestions
# + id="W9jSGRMORXlT"
df_questions_initial_tag = get_questions_from_tags(POSTS_file, Tag_list_initial)
# + colab={"base_uri": "https://localhost:8080/"} id="En5bJuiXQr_r" outputId="741f149b-1a45-4daa-aefd-b6f812c5554f"
print(len(df_questions_initial_tag))
df_questions_initial_tag.drop_duplicates('Id', inplace=True)
print(len(df_questions_initial_tag))
# + colab={"base_uri": "https://localhost:8080/"} id="jBtacFhgWT6C" outputId="de6aa3f9-5a18-402f-cc37-3f3caf3a7b01"
print(len(df_questions_initial_tag))
output_file = os.path.join(ROOT_dir, "output", "questions_initial_tag.csv")
df_questions_initial_tag.to_csv(output_file, index=False, quotechar='"', quoting=csv.QUOTE_NONNUMERIC)
print("Check the output file and then move it to dataset folder")
save_obj(df_questions_initial_tag, "df_questions_initial_tag")
# + [markdown] id="2pgwBZIKWpJW"
# ### Significance and Relevance
# + [markdown] id="jXvOIBraXbOo"
# #### Tag stat selected question
# + colab={"base_uri": "https://localhost:8080/"} id="2asXK71SXz85" outputId="90484f75-3f86-4088-8ae5-50f6fd07f524"
def get_tags_stat(df_questions):
tags_stat = {}
for index, row in df_questions.iterrows():
tags = get_tags(row['Tags'])
for tag in tags:
tags_stat[tag] = tags_stat.get(tag, 0) + 1
return tags_stat
Tag_stat_Q_initial = get_tags_stat(df_questions_initial_tag)
print(len(Tag_stat_Q_initial))
# + colab={"base_uri": "https://localhost:8080/"} id="RgRKmKK7bplJ" outputId="aa02e3b3-e127-45fd-afef-d6af287dd111"
save_obj(Tag_stat_Q_initial, "Tag_stat_Q_initial")
# Tag_stat_Q_initial = load_obj("Tag_stat_Q_initial")
print(len(Tag_stat_Q_initial))
# + [markdown] id="aDvUNJLS0dFv"
# #### Tag stat all questions
# + id="gf72_O05efSq"
def tag_stat_all_questions(selected_tag_list):
tags_stat = {}
context = ET.iterparse(POSTS_file, events=("end",))
print("Going to extract all tags stat from questions")
# df_posts = pd.DataFrame(columns = COLS)
total_questions = 0
_, root = next(context)
for event, elem in context:
if elem.tag == "row":
post_tags = elem.attrib.get('Tags', '')
post_tags_list = get_tags(post_tags) # list of tags
for tag in post_tags_list:
if tag in selected_tag_list:
tags_stat[tag] = tags_stat.get(tag, 0) + 1
# progress
if total_questions % 1000000 == 0:
print('done', elem.attrib['Id'])
elem.clear()
root.clear()
total_questions += 1
# count += 1
# print(len(all_tags_in_Q_all))
return tags_stat
Tag_stat_Q_all = tag_stat_all_questions(Tag_stat_Q_initial)
# + colab={"base_uri": "https://localhost:8080/"} id="L1bClBHHuzmB" outputId="e99eeb94-6bb9-4895-abeb-9c31ded4af07"
# save_obj(Tag_stat_Q_all, "Tag_stat_Q_all")
Tag_stat_Q_all = load_obj("Tag_stat_Q_all")
print(len(Tag_stat_Q_all))
# Tag_stat_Q_all = load_obj("all_tags_in_Q_all_stat")
# save_obj(all_tags_in_Q_all, "all_tags_in_Q_all_stat")
# # all_tags_in_Q_all = load_obj("all_tags_in_Q_all")
# + [markdown] id="oMV3BbeCW1hL"
# #### Calculate Significance relevance
# + colab={"base_uri": "https://localhost:8080/"} id="Fqfc1Rl9W5Kn" outputId="96bc327c-a5e4-4f26-aca8-53450cf43d92"
for tag in Tag_stat_Q_initial:
if tag in Tag_list_initial:
continue
sig = Tag_stat_Q_initial[tag] * 1.0 / Tag_stat_Q_all[tag]
rel = Tag_stat_Q_initial[tag] * 1.0 / len(df_questions_initial_tag)
Significance_tag[tag] = sig
Relevance_tag[tag] = rel
# + id="Al6xv5OqXCxf"
for tag in Tag_stat_Q_initial:
if tag in Tag_list_initial:
continue
print("Tag: %s sig: %f rel: %f" % (tag, Significance_tag[tag], Relevance_tag[tag] ))
# + id="2CYtUzpV40gz"
save_obj(Significance_tag, "Significance_tag")
save_obj(Relevance_tag, "Relevance_tag")
# + [markdown] id="HdGeeD4r8HGD"
# #### Extract based on sig rel
# + colab={"base_uri": "https://localhost:8080/"} id="Cm1kW5kcZtyO" outputId="6299576c-f1d1-4ae5-812d-6a143451e55f"
def get_tag_with_higher_sig_and_rel(sig, rel, sig_map, rel_map):
assert len(sig_map) == len(rel_map)
results = []
for tag in sig_map:
if(sig_map[tag] >= sig and rel_map[tag] >= rel):
results.append(tag)
return results
def get_intersection_info(list_big, list_ref):
list_ref = set(list_ref)
count = 0
for item in list_big:
if item in list_ref:
count += 1
pct = round(count * 100.0 / len(list_big), 1)
return count, pct
# + colab={"base_uri": "https://localhost:8080/"} id="uDiWvTp6K23x" outputId="c9a4fa70-79c5-4f98-c816-4614a773af39"
tags_with_lowest_sig_rel = get_tag_with_higher_sig_and_rel(.05, .001, Significance_tag, Relevance_tag)
print(len(tags_with_lowest_sig_rel))
# + colab={"base_uri": "https://localhost:8080/"} id="edw2uvLT9ULm" outputId="98354cc4-e44e-461c-c56f-7b7079137141"
tmp_sigs = []
tmp_rel = []
tmp_posts = []
for tag in tags_with_lowest_sig_rel:
tmp_sigs.append(Significance_tag[tag])
tmp_rel.append(Relevance_tag[tag])
tmp_posts.append(Tag_stat_Q_all[tag])
df_tag_recommend = pd.DataFrame()
df_tag_recommend['sig'] = tmp_sigs
df_tag_recommend['tag'] = tags_with_lowest_sig_rel
df_tag_recommend['rel'] = tmp_rel
df_tag_recommend['posts'] = tmp_posts
output_file = os.path.join("tags", "tags_recommended_stat.csv")
df_tag_recommend.to_csv(output_file, index=False)
# + colab={"base_uri": "https://localhost:8080/"} id="fiBTFd2Ubu5v" outputId="a4acf3aa-e195-4220-ef28-6e41c83a0e83"
df_Tag_stat_manual_relevance = pd.read_csv(TAG_manual_relevance_file)
print(len(df_Tag_stat_manual_relevance))
df_Tag_stat_manual_relevance = df_Tag_stat_manual_relevance[df_Tag_stat_manual_relevance['isrelavent'] == True]
print(len(df_Tag_stat_manual_relevance))
Tag_stat_manual_relevance = df_Tag_stat_manual_relevance['tags'].tolist()
# print(Tag_stat_manual_relevance)
temp = []
for tag in Tag_stat_manual_relevance:
tag = tag[1:-1]
temp.append(tag)
Tag_stat_manual_relevance = temp
print(Tag_stat_manual_relevance)
# + colab={"base_uri": "https://localhost:8080/"} id="8mEywGqb0Ald" outputId="b30919b3-a7ef-44e7-bf72-edebdc2301aa"
save_obj(Tag_stat_manual_relevance, 'Tag_stat_manual_relevance')
# + colab={"base_uri": "https://localhost:8080/"} id="GcnGSOXrY4JX" outputId="bc4eeab8-d084-45ff-d8d3-2a220cd591b6"
# Uses global variable of Significance_tag and Relevance_tag map
def get_tag_stat_summary(significance_values, relavance_values, Tag_stat_manual_relevance):
sig_res = []
rel_res = []
tags_recommended = []
tags_recommended_count = []
tags_relevant = []
tags_relavant_pct = []
for rel in relavance_values:
for sig in significance_values:
reco_tags = get_tag_with_higher_sig_and_rel(sig, rel, Significance_tag, Relevance_tag)
rel_tags_count, rel_pct = get_intersection_info(reco_tags, Tag_stat_manual_relevance)
# df = pd.DataFrame()
# df['tags'] = selected_tags
# file_name = "Sig_%.3f_Rel_%.3f.csv" % (sig, rel)
# file_name = os.path.join("tags", file_name)
# # print(file_name)
# df.to_csv(file_name, index=False)
sig_res.append(sig)
rel_res.append(rel)
tags_recommended.append(reco_tags)
tags_recommended_count.append(len(reco_tags))
tags_relevant.append(rel_tags_count)
tags_relavant_pct.append(rel_pct)
# print("Sig: %f Rel: %f len_tags: %d" % (sig, rel, len(res)))
df = pd.DataFrame()
df["significance"] = sig_res
df['relevance'] = rel_res
df['tags_recommended'] = tags_recommended
df['tags_relevant'] = tags_relevant
df['tags_relavant_pct'] = tags_relavant_pct
df['tags_recommended_count'] = tags_recommended_count
return df
significance_values = [0.05, 0.10, 0.15, 0.20, 0.25, 0.30, 0.35]
relavance_values = [0.001, .005, .01, .015, .02, .025, .03]
DF_tag_stat_summary = get_tag_stat_summary(significance_values, relavance_values, Tag_stat_manual_relevance)
file_name = os.path.join("tags", "tags_stat_summary.csv")
DF_tag_stat_summary.to_csv(file_name, index=False)
save_obj(DF_tag_stat_summary, 'DF_tag_stat_summary')
# + [markdown] id="jK16NVqMTOWi"
# ### Extract final posts
# + [markdown] id="CMATt3v4IH_A"
# #### Final taglist
#
# + colab={"base_uri": "https://localhost:8080/"} id="UH-n28sndH4f" outputId="e1417af2-d15b-4df7-f5d4-69e1fbb64a8a"
print(Tag_list_relevent)
print(len(Tag_list_relevent))
print(len(Tag_list_initial))
tag_unique = set(Tag_list_relevent + Tag_list_initial)
print(len(tag_unique))
# + id="IF9UR3G0gkL3"
# + id="WwnjkTQaIK0l"
Tag_list_relevent = util.load_obj(ROOT_dir, "Tag_stat_manual_relevance")
Tag_list_initial = util.load_obj(ROOT_dir, "Tag_list_initial")
Tag_list_final = Tag_list_relevent + Tag_list_initial
print(len(Tag_list_final))
# save_obj(Tag_list_final, "Tag_list_final")
# + [markdown] id="fnWWzOR7TUzR"
# #### Final questions
# + colab={"base_uri": "https://localhost:8080/"} id="RSRFsFlanwVm" outputId="d4f906cd-b6c4-44a4-d78f-57727ab1ffec"
DF_question_final_tag = util.load_obj(ROOT_dir, "DF_question_final_tag")
print(len(DF_question_final_tag))
DF_question_final_tag = util.load_obj(ROOT_dir, "DF_question_final_tag")
# + id="jhF2OKUfJjcC"
DF_question_final_tag = get_questions_from_tags(POSTS_file, Tag_list_final)
print(len(DF_question_final_tag))
output_file = os.path.join("output", "questions_final_tag.csv")
DF_question_final_tag.to_csv(output_file, index=False)
save_obj(DF_question_final_tag, "DF_question_final_tag")
# + id="zkVgjOMJSVPF"
# print(len(DF_question_final_tag))
# DF_question_final_tag.drop_duplicates('Id', inplace=True)
# output_file = os.path.join("output", "questions_final_tag.csv")
# DF_question_final_tag.to_csv(output_file, index=False)
# save_obj(DF_question_final_tag, "DF_question_final_tag")
# print(len(DF_question_final_tag))
# + colab={"base_uri": "https://localhost:8080/"} id="MpaG59oKWdvg" outputId="4f7d935b-6d33-4745-917e-961c3d58f9ef"
# print(len(DF_question_final_tag))
# + colab={"base_uri": "https://localhost:8080/"} id="6XU_jmdHH725" outputId="c157fed9-6a5c-4947-9110-3bc22c4c4613"
DF_question_nss_final_tag = DF_question_final_tag[DF_question_final_tag['Score'].astype(int) >= 0]
print(len(DF_question_nss_final_tag))
save_df(DF_question_nss_final_tag, QUESTIONS_nss_final_tag_file)
save_obj(DF_question_nss_final_tag, "DF_question_nss_final_tag")
# + [markdown] id="1l5i4BGWTXyy"
# #### Final answers
# + id="BLsAaq--YwIn"
def get_answers_from_questions(question_ids):
question_ids = set(question_ids)
context = ET.iterparse(POSTS_file, events=("end",))
print("Going to extract answers")
df_answers = pd.DataFrame(columns = COLS)
total_answers = 0
count = 0
_, root = next(context)
for event, elem in context:
if elem.tag == "row":
post_type = int(elem.attrib.get('PostTypeId'))
if(post_type == 2):
parent_id = elem.attrib.get('ParentId')
if(parent_id in question_ids):
dic = {}
for col in COLS:
dic[col] = elem.attrib.get(col, '')
df_answers = df_answers.append(pd.Series(dic), ignore_index = True)
# progress
if (int(elem.attrib['Id']) % 1000000) == 0:
print('done id: %s and len of res: %d' % (elem.attrib['Id'], len(df_answers)))
total_answers += 1
elem.clear()
root.clear()
print("Total answers: %d" % (total_answers))
return df_answers
question_ids = DF_question_final_tag['Id'].tolist()
print(len(question_ids))
DF_answer_final_tag = get_answers_from_questions(question_ids)
print(len(DF_answer_final_tag))
output_file = os.path.join("output", "answers_final_tag.csv")
DF_answer_final_tag.to_csv(output_file, index=False)
save_obj(DF_answer_final_tag, "DF_answer_final_tag")
# + colab={"base_uri": "https://localhost:8080/"} id="K1pbtSpATEG1" outputId="c1001f24-263b-4fbf-c1d3-f017d2e178db"
print(len(DF_answer_final_tag))
DF_answer_final_tag.drop_duplicates('Id', inplace=True)
output_file = os.path.join("output", "answers_final_tag.csv")
DF_answer_final_tag.to_csv(output_file, index=False)
save_obj(DF_answer_final_tag, "DF_answer_final_tag")
print(len(DF_answer_final_tag))
# + colab={"base_uri": "https://localhost:8080/"} id="Ma4NLcXsIeob" outputId="038ab143-c69c-4635-cf89-22d1293dfa4f"
q_ids = DF_question_nss_final_tag["Id"].tolist()
print(len(q_ids))
print(len(DF_answer_final_tag))
DF_answer_nss_final_tag = DF_answer_final_tag[DF_answer_final_tag['ParentId'].isin(q_ids)]
print(len(DF_answer_nss_final_tag))
save_df(DF_answer_nss_final_tag, ANSWERS_nss_final_tag_file)
save_obj(DF_answer_nss_final_tag, "DF_answer_nss_final_tag")
# + [markdown] id="XqDLfiZUBqfW"
# ### Tag list to Questions
# + id="YLbKLwLaBt9b"
Map_tag_qids = {}
for tag in Tag_list_final:
Map_tag_qids[tag] = []
for index, row in All_Questions_df.iterrows():
tags = get_tags(row['Tags'])
for tag in tags:
if(tag in Tag_list_final):
Map_tag_qids[tag].append(int(row['Id']))
save_obj(Map_tag_qids, "Map_tag_qids")
total = 0
for tag in Map_tag_qids:
total += len(Map_tag_qids[tag])
print(tag, len(Map_tag_qids[tag]))
print(total)
# + id="YDzENhFHNtk8"
# + [markdown] id="QwiRrfGQNuEy"
# ### Generate final stats on posts.xml
# + colab={"base_uri": "https://localhost:8080/"} id="zZAKvaADBuSD" outputId="4229706e-e5b7-4eb5-ccde-7db74ffe400b"
def generate_final_stats(POSTS_file):
context = ET.iterparse(POSTS_file, events=("end",))
print("Going to extract questions")
total_posts = 0
total_Q = 0
total_A = 0
total_Q_with_acc = 0
all_post_ids = set()
_, root = next(context)
for event, elem in context:
if elem.tag == "row":
id = int(elem.attrib.get('Id'))
if id in all_post_ids:
continue
all_post_ids.add(id)
total_posts += 1
post_type = int(elem.attrib.get('PostTypeId'))
if(post_type == 1):
total_Q += 1
acc_id = (elem.attrib.get('AcceptedAnswerId'))
if(acc_id is not None and len(acc_id) > 4):
total_Q_with_acc += 1
else:
total_A += 1
# progress
if total_posts % 10000000 == 0:
print('done', elem.attrib['Id'])
# break
elem.clear()
root.clear()
return total_posts, total_Q, total_A, total_Q_with_acc
total_posts, total_Q, total_A, total_Q_with_acc = generate_final_stats(POSTS_file)
print(total_posts, total_Q, total_A, total_Q_with_acc)
# + [markdown] id="QldpYk17FErQ"
# ### Sanity testing
# + colab={"base_uri": "https://localhost:8080/"} id="vU9Lq0VJnUg3" outputId="cb0d5b77-d1da-4594-ee1c-7418451bbfe0"
df = DF_question_final_tag[DF_question_final_tag['Score'].astype(int) < 0]
print(len(df))
# + colab={"base_uri": "https://localhost:8080/"} id="zmY6vv-hRCpD" outputId="f699b2bf-13e0-4ca6-e141-097c4ec18430"
ids = df_questions_initial_tag['Id'].tolist()
print(len(ids))
print(len(set(ids)))
# + id="iUxRzsVpGo7B"
df = pd.read_csv("experiment/IoTPostInformation.csv")
print(len(df))
print(df.head())
ids = df['PostId'].tolist()
print(len(ids))
print(len(set(ids)))
# + colab={"base_uri": "https://localhost:8080/"} id="Zu25xJmDFJMs" outputId="64e41c30-21a5-4650-ba48-64c9ad23dc8e"
# assert len(question_ids) == len(set(question_ids))
question_ids = DF_question_final_tag['Id'].tolist()
print(len(question_ids))
print(len(set(question_ids)))
# + colab={"base_uri": "https://localhost:8080/"} id="YQJ5v1yEBMgz" outputId="33208b1d-d664-46fe-cf52-4dc5d5f15c3f"
total_answers = DF_question_final_tag['AnswerCount'].astype(int).sum()
accepted_answers = DF_question_final_tag[DF_question_final_tag['AcceptedAnswerId'].str.len() > 0]['AcceptedAnswerId'].tolist()
# print(accepted_answers)
print(len(accepted_answers))
print(total_answers)
print(len(DF_answer_final_tag))
# + colab={"base_uri": "https://localhost:8080/"} id="XJ6rT-RWEl6j" outputId="e5389e43-22fb-49d8-c636-722303dd6b32"
print(type(question_ids[0]))
# + [markdown] id="EpMofJ6REsHU"
# ## Topic Modelling
# + [markdown] id="pofweTsgR9Ux"
# ### Bootstrap
#
# + colab={"base_uri": "https://localhost:8080/"} id="EeepMhchj86Z" outputId="69bb4e78-bbef-48b8-9d4d-fe55a9c4163e"
import string
import gensim
import unicodedata
from gensim import corpora
from nltk.corpus import stopwords
from nltk.tokenize import sent_tokenize, word_tokenize
from nltk.stem import WordNetLemmatizer
from nltk.stem.porter import *
from gensim.models.wrappers import LdaMallet
from gensim.models import CoherenceModel
import pyLDAvis.gensim
import pyLDAvis
import json
import html
import warnings
from tqdm import tqdm
import matplotlib.pyplot as plt
warnings.filterwarnings('ignore')
# + colab={"base_uri": "https://localhost:8080/"} id="SCcOVdMQkHii" outputId="752568ce-53c1-42ae-fd24-cb5ec160054b"
stemmer = PorterStemmer()
pyLDAvis.enable_notebook()
# os.environ.update({'MALLET_HOME':'./Mallet'})
# mallet_path = './Mallet/bin/mallet'
os.environ.update({'MALLET_HOME': r"C:\Mallet"})
mallet_path = r'C:\Mallet\bin\mallet' # update this path
current_dir = os.getcwd()
coherence_scores = []
# ldamallet = gensim.models.wrappers.LdaMallet(mallet_path, corpus=corpus, num_topics=5, id2word=dictionary, alpha=5)
# + colab={"base_uri": "https://localhost:8080/"} id="v-BBCleKkyIv" outputId="fa2757d7-cb11-4971-c889-d0983fcbcd95"
def create_dir(parent_dir, dir_name):
temp = os.path.join(parent_dir,dir_name)
try:
os.mkdir(temp)
except OSError as error:
print(error)
return temp
# + [markdown] id="soO624mdkQMI"
# #### Variable initializations
# + colab={"base_uri": "https://localhost:8080/"} id="yLZmGSSmR_jV" outputId="382b595c-5f77-4119-de06-57d8e55d2469"
DF_question_nss_final_tag = load_obj("DF_question_nss_final_tag")
DF_answer_nss_final_tag = load_obj("DF_answer_nss_final_tag")
DF_accepted_answer_nss_final_tag = None
TM_data_file = os.path.join("output", "topicModeling", "TM_data.csv")
TM_data_processed_file = os.path.join("output", "topicModeling", "TM_data_processed.csv")
# + colab={"base_uri": "https://localhost:8080/"} id="mAOC83b-ow2m" outputId="72a96c04-8f40-41e0-c2bc-7063f5ac4975"
# + [markdown] id="Q4J7B_j0RxyL"
# ### Preprocess the posts
# + colab={"base_uri": "https://localhost:8080/"} id="TXUW-cskj480" outputId="b344621b-d288-4e51-949a-3887a8c53cd9"
accepted_answers_ids = DF_question_nss_final_tag[DF_question_nss_final_tag['AcceptedAnswerId'].str.len() > 0]['AcceptedAnswerId'].tolist()
print(len(accepted_answers_ids))
DF_accepted_answer_nss_final_tag = DF_answer_nss_final_tag[DF_answer_nss_final_tag['Id'].isin(accepted_answers_ids)]
print(len(DF_accepted_answer_nss_final_tag))
# + colab={"base_uri": "https://localhost:8080/"} id="iTikx0U3lDhm" outputId="294824d6-0d5b-4bc4-8e63-01270a5eedb9"
answers_df = DF_accepted_answer_nss_final_tag.copy()
answers_df = answers_df[['Id', 'Body']]
answers_df = answers_df.dropna()
answers_df.columns = ['Id', 'raw_data']
answers_df.insert(1, 'qa', 'a')
answers_df.Id = answers_df.Id.apply(np.int64)
print(answers_df.head())
print(len(answers_df))
# + colab={"base_uri": "https://localhost:8080/"} id="DWHlTKxmnwl9" outputId="d44fead5-7a77-4732-e0b2-e2667eb4daed"
questions_df = DF_question_nss_final_tag.copy()
questions_df = questions_df[['Id', 'Body', 'Title']]
questions_df['titlePlusQuestion'] = questions_df[['Title', 'Body']].apply(lambda x: ' '.join(x), axis=1)
questions_df = questions_df.drop(['Title', 'Body'], axis=1)
questions_df.columns = ['Id', 'raw_data']
questions_df.insert(1, 'qa', 'q')
questions_df.Id = questions_df.Id.apply(np.int64)
print(questions_df.head())
print(len(questions_df))
# + colab={"base_uri": "https://localhost:8080/"} id="4ytbj19xoMMp" outputId="16200a98-fe77-4d63-fe01-97f71631e8af"
df = pd.concat([questions_df,answers_df])
print(len(df))
# output_file = os.path.join("output", "topicModeling", "TM_data")
# df.to_csv(TM_data_file, index=False, quotechar='"', quoting=csv.QUOTE_NONNUMERIC)
save_df(df, TM_data_file)
# + [markdown] id="mYpEoYk6lNAD"
# #### Generate temporary TM file
# + [markdown] id="CCmgdOVmkbOb"
# #### Process stop words
# + colab={"base_uri": "https://localhost:8080/"} id="5ivdAYg3rEa9" outputId="20a96a94-3f6b-4625-94fb-738170aeed85"
import nltk
# nltk.download('stopwords')
nltk.download('wordnet')
# + colab={"base_uri": "https://localhost:8080/"} id="NZrAVkg8SNAk" outputId="f75c13a8-79b9-42eb-9402-95b9fdfa02d9"
# stop words set
STOP_WORDS_FILES = ['mallet_stop_words.txt']
stop_words_set = set()
stop_words = set(stopwords.words('english'))
for word in stop_words:
if('\'' in word):
stop_words_set.add(word.strip().replace('\'', ''))
stop_words_set.add(word)
for swfile in STOP_WORDS_FILES:
try:
with open(swfile, 'r') as f:
words = f.readlines()
for word in words:
if('\'' in word):
stop_words_set.add(word.strip().replace('\'', ''))
stop_words_set.add(word.strip())
except:
pass
# + colab={"base_uri": "https://localhost:8080/"} id="JdgTGaArkkld" outputId="1efdcd73-2b11-4536-932b-fb091e4ecfa6"
def preprocess_text(text):
# remove non ascii
text = unicodedata.normalize('NFKD', text).encode(
'ascii', 'ignore').decode('utf-8', 'ignore')
text = text.lower()
# unescaping
text = html.unescape(text)
text = re.sub(r'<code>(.|\n)*?</code>','', text) # removing <code>...</code>
text = re.sub(r'<a.*?</a>', '', text) # removing whole anchor tags
text = re.sub(r'(<.*?>)', '', text) # removing html markup
text = re.sub(r'[^\w\s]', '', text) # removing punctuation
text = re.sub(r'[\d]', '', text) # removing digits
# remove stopwords
tokenized = []
for word in text.split():
if word in stop_words_set:
continue
tokenized.append(word)
for i in range(len(tokenized)):
word = tokenized[i]
word = WordNetLemmatizer().lemmatize(word, pos='v')
tokenized[i] = stemmer.stem(word)
# tokenized[i] = word
return tokenized
# + colab={"base_uri": "https://localhost:8080/", "height": 224} id="5dMroUcWkpq-" outputId="d945e8d4-d317-480f-8513-225035657d6b"
df_raw = pd.read_csv(TM_data_file)
df_raw['preprocessed'] = ""
for i in range(len(df_raw)):
df_raw['preprocessed'].iloc[i] = preprocess_text(df_raw.raw_data.iloc[i])
save_df(df_raw, TM_data_processed_file)
# df.to_csv('preprocesseedData.csv')
df_raw.head()
# + colab={"base_uri": "https://localhost:8080/"} id="MAKwkD80K5AX" outputId="276d9ef6-7c5f-4fbd-c02e-083afb5aaabf"
save_obj(df_raw, "processed_df")
# + colab={"base_uri": "https://localhost:8080/"} id="9qCuLkwR-ZxK" outputId="a6cd6689-67a5-43d0-9a9a-2dd14dcb0bb8"
# save_obj(df_raw, "processed_df")
df_TM = load_obj("processed_df")
print(len(df_TM))
# + [markdown] id="_ZpFrX7jR1ME"
# ### Topic Modelling
# + [markdown] id="zo3BmJ8Wk6my"
# #### Optimal number of topics
# + id="AE5Os6Fe0W6l"
# df_TM = pd.read_csv(TM_data_processed_file)
# print(df.head)
# + colab={"base_uri": "https://localhost:8080/"} id="h02IClOcr3I0" outputId="1af5d776-1336-4c9f-8598-90957c32e5b8"
topic_range = []
print(topic_range)
for i in range(5, 71, 5):
if i not in topic_range:
topic_range.append(i)
topic_range.sort()
print(topic_range)
# + colab={"base_uri": "https://localhost:8080/", "height": 534} id="p7Y5S6_mSNko" outputId="9f1e79e5-50ef-488d-fc78-20ba5a8a1e38"
# finding out optimum topic number
coherence_file = os.path.join("output", "topicModeling", "Coherence_Scores_5_70_nns.csv")
image_file = "scores_5_70_nns.png"
data = df_TM['preprocessed']
print(data)
dictionary = gensim.corpora.Dictionary(data)
corpus = [dictionary.doc2bow(doc) for doc in data]
coherence_scores = []
for num_topics in tqdm(topic_range):
ldamallet = gensim.models.wrappers.LdaMallet(
mallet_path, corpus=corpus, num_topics=num_topics, id2word=dictionary, alpha=50/num_topics, iterations=1000)
coherenceModel = CoherenceModel(model=ldamallet, texts=data, dictionary=dictionary, coherence='c_v')
score = coherenceModel.get_coherence()
coherence_scores.append([num_topics,score])
# save scores as csv
ch_df = pd.DataFrame(coherence_scores,columns=['Num Topic','Score'])
save_df(ch_df, coherence_file)
# ch_df.to_csv('Coherence_Scores.csv')
# plot
plt.xlabel('Number of Topics')
plt.ylabel('Coherence Score')
x = []
y = []
for score in coherence_scores:
x.append(score[0])
y.append(score[1])
plt.plot(x,y,c='r')
plt.gca().set_aspect('auto')
plt.grid()
plt.savefig(image_file, dpi=500)
plt.show()
# + [markdown] id="82_pdJ28k9te"
# #### Generating topics
# + colab={"base_uri": "https://localhost:8080/"} id="JyKl6hRuunR0" outputId="83705d1e-015f-4ece-eb33-c3d5fe703732"
# + colab={"base_uri": "https://localhost:8080/"} id="Vld1AVuRlAsp" outputId="802f59f7-2388-4012-dda3-d39a2ccd2a6f"
res_dir = create_dir(current_dir, 'TM_Run2')
data = df_TM['preprocessed']
dictionary = gensim.corpora.Dictionary(data)
corpus = [dictionary.doc2bow(doc) for doc in data]
# create folder for topic number
NUM_TOPIS = [25, 45]
for num_topics in NUM_TOPIS:
topic_dir = create_dir(res_dir, f'{num_topics}_Topics')
if os.path.isfile(os.path.join(topic_dir, 'ldamallet.pkl')):
ldamallet = pickle.load(
open(os.path.join(topic_dir, 'ldamallet.pkl'), "rb"))
else:
ldamallet = gensim.models.wrappers.LdaMallet(
mallet_path, corpus=corpus, num_topics=num_topics, id2word=dictionary, alpha=50/num_topics, iterations=1000)
# save the model as pickle
pickle.dump(ldamallet, open(os.path.join(
topic_dir, 'ldamallet.pkl'), "wb"))
topics = [[(word, word_prob) for word, word_prob in ldamallet.show_topic(
n, topn=30)] for n in range(ldamallet.num_topics)]
# term-topic matrix
topics_df = pd.DataFrame([[f'{word} {round(word_prob,4)}' for word, word_prob in topic] for topic in topics], columns=[
f'Term {i}' for i in range(1, 31)], index=[f'Topic {n}' for n in range(ldamallet.num_topics)]).T
topics_df.to_csv(os.path.join(topic_dir, 'term x topic.csv'))
# topic words
topic_words_dir = create_dir(topic_dir, 'TopicWords')
for n in range(num_topics):
topic_words_df = pd.DataFrame(
[[word_prob, word]for word, word_prob in topics[n]], columns=['Prob', 'Word'])
topic_words_df.to_csv(os.path.join(topic_words_dir, f'{n}.csv'))
# post to dominant topic
corpus_topic_df = pd.DataFrame()
corpus_topic_df['link'] = df.Id
corpus_topic_df['qa'] = df.qa
for i in range(len(corpus_topic_df)):
corpus_topic_df.link.iloc[i] = make_link(df.Id.iloc[i],df.qa.iloc[i])
topic_model_results = ldamallet[corpus]
corpus_topics = [sorted(doc, key=lambda x: -x[1])[0]
for doc in topic_model_results]
corpus_topic_df['Dominant Topic'] = [item[0] for item in corpus_topics]
corpus_topic_df['Correlation'] = [item[1] for item in corpus_topics]
corpus_topic_df.to_csv(os.path.join(topic_dir, 'postToTopic.csv'))
topic_to_post_dir = create_dir(topic_dir, 'TopicToPost')
for i in range(num_topics):
temp = create_dir(topic_to_post_dir, str(i))
temp_q_df = corpus_topic_df.loc[corpus_topic_df['Dominant Topic'] == i]
temp_q_df = temp_q_df.loc[temp_q_df['qa'] == 'q']
temp_a_df = corpus_topic_df.loc[corpus_topic_df['Dominant Topic'] == i]
temp_a_df = temp_a_df.loc[temp_a_df['qa'] == 'a']
temp_q_df.drop(columns=['Dominant Topic','qa']).to_csv(
os.path.join(temp, 'Questions.csv'), index=False)
# temp_q_df.drop(columns=['Dominant Topic','qa']).to_excel(
# os.path.join(temp, 'Questions.xlsx'), index=False)
temp_a_df.drop(columns=['Dominant Topic','qa']).to_csv(
os.path.join(temp, 'Answers.csv'), index=False)
# temp_a_df.drop(columns=['Dominant Topic','qa']).to_excel(
# os.path.join(temp, 'Answers.xlsx'), index=False)
# post count under any topic
topic_post_cnt_df = corpus_topic_df.groupby('Dominant Topic').agg(
Document_Count=('Dominant Topic', np.size),
Percentage=('Dominant Topic', np.size)).reset_index()
topic_post_cnt_df['Percentage'] = topic_post_cnt_df['Percentage'].apply(
lambda x: round((x*100) / len(corpus), 2))
topic_post_cnt_df.to_csv(os.path.join(topic_dir, 'postPerTopic.csv'))
# pyLDAvis
vis = pyLDAvis.gensim.prepare(
gensim.models.wrappers.ldamallet.malletmodel2ldamodel(ldamallet), corpus, dictionary)
pyLDAvis.save_html(vis, os.path.join(topic_dir, f'pyLDAvis.html'))
# + [markdown] id="d766kEtq1QDF"
# #### Update topic outputs
# + colab={"base_uri": "https://localhost:8080/"} id="0GoP6-AVEwKe" outputId="e70e2ef3-c129-4069-e79c-e8fdae511a21"
print(res_dir)
def get_post_stat(df, qids):
res_score = []
res_view = []
for qid in qids:
q = df[df['Id'] == str(qid)].iloc[0]
res_score.append(q.Score)
res_view.append(q.ViewCount)
return res_score, res_view
def update_TM_reresult(file):
df = pd.read_csv(file)
links = df['link'].tolist()
# print(links[0])
# print(linkToId(links[0]))
ids = [linkToId(i) for i in links]
res_score, res_view = get_post_stat(DF_question_nss_final_tag, ids)
df['score'] = res_score
df['view_count'] = res_view
# print(df.head)
save_df(df, file)
# df.to_csv("updated.csv")
# return df
update_TM_reresult(file)
# + [markdown] id="uSsCmJ5sdnet"
# # Results
# + colab={"base_uri": "https://localhost:8080/"} id="lDVr2cY00Tyn" outputId="fb4ca13c-625e-4556-eed6-dded4cbdf4ab"
MAP_tm_id_quid = load_obj("MAP_tm_id_quid")
MAP_merged_tm_quid = load_obj("MAP_merged_tm_quid")
TMs = load_obj("TMs")
Map_cat_topic = load_obj("Map_cat_topic")
DF_question_nss_final_tag = load_obj("DF_question_nss_final_tag")
Map_topic_pop_diff = load_obj("Map_topic_pop_diff")
Map_cat_ques = load_obj("Map_cat_ques")
All_Questions_df = load_obj("DF_question_nss_final_tag")
# + [markdown] id="ieocUnhxwiIH"
# ## RQ Topic Modelling
# + colab={"base_uri": "https://localhost:8080/"} id="zowfEkulzyzz" outputId="73abe0ab-6e39-471f-91ac-72acdbdb8b98"
class TM:
def __init__(self):
self.name = ""
self.ids = []
self.low_cat = ""
self.mid_cat = ""
self.high_cat = ""
def __str__(self):
print("Name: %s, Ids: %s, Low: %s, Mid: %s, High: %s" % (self.name, self.ids, self.low_cat, self.mid_cat, self.high_cat))
# + colab={"base_uri": "https://localhost:8080/"} id="o5sskrKcwp8Y" outputId="e6d8d7b0-1166-4489-9eed-2822c5766367"
# https://www.surveysystem.com/sscalc.htm
High_cats = ["Customization", "Data Storage", "Platform Adoption", "Platform Maintenance", "Third-Party Integration" ]
Sample_size = {"Customization": 95 , "Data Storage": 95, "Platform Adoption": 94, "Platform Maintenance": 94, "Third-Party Integration": 93}
File_Cat_sample = os.path.join(ROOT_dir, "output", "Category_sample.csv")
Total_Questions = 26763
Total_Ansers = 11010
# + colab={"base_uri": "https://localhost:8080/"} id="im-RCIGSyYXN" outputId="83d45123-39c7-44dc-8a21-482374f55bbc"
MAP_tm_id_quid = load_obj("MAP_tm_id_quid")
MAP_merged_tm_quid = load_obj("MAP_merged_tm_quid")
MAP_tm_id_ansid = load_obj("MAP_tm_id_ansid")
MAP_merged_tm_ansid = load_obj("MAP_merged_tm_ansid")
TMs = load_obj("TMs")
Map_sample = load_obj("Map_sample")
Map_cat_ques = load_obj("Map_cat_ques")
Map_cat_topic = load_obj("Map_cat_topic")
DF_sample = load_obj("DF_sample")
# MAP_tm_id_quid = load_obj("MAP_tm_id_quid")
# MAP_tm_id_quid = load_obj("MAP_tm_id_quid")
# MAP_tm_id_quid = load_obj("MAP_tm_id_quid")
# MAP_tm_id_quid = load_obj("MAP_tm_id_quid")
# MAP_tm_id_quid = load_obj("MAP_tm_id_quid")
# MAP_tm_id_quid = load_obj("MAP_tm_id_quid")
# + colab={"base_uri": "https://localhost:8080/"} id="olIP8Zh97gBI" outputId="06057498-9183-49d0-8c86-b4ee431f5e13"
Map_cat_ques = {}
# + [markdown] id="wzjfy56wwlTM"
# #### TopicID vs QuestionID
# + [markdown] id="J5Q_kOSJEX_A"
# topics to question_ids
# + id="bClzTUO-DH8b"
df_tm_ques = df_tm[df_tm['qa'] == 'q']
print(len(df_tm_ques))
for topic_id in range(45):
MAP_tm_id_quid[topic_id] = []
ques_ids = df_tm_ques[df_tm_ques['Dominant Topic'] == topic_id]['link'].tolist()
for i in range(len(ques_ids)):
ques_ids[i] = linkToId(ques_ids[i])
MAP_tm_id_quid[topic_id] = ques_ids
# + colab={"base_uri": "https://localhost:8080/"} id="_Vbddk55HNEZ" outputId="13afb826-49a5-4c87-fdec-e506de43293a"
save_obj(MAP_tm_id_quid, "MAP_tm_id_quid")
# + [markdown] id="m11gMYMAEcPK"
# Topics to answers ids
# + colab={"base_uri": "https://localhost:8080/"} id="vjVyshG6Ee5k" outputId="a25c2940-236e-4144-dffd-fb4ff0f0c097"
df_tm_ans = df_tm[df_tm['qa'] == 'a']
print(len(df_tm_ans))
MAP_tm_id_ansid = {}
for topic_id in range(45):
MAP_tm_id_ansid[topic_id] = []
ans_ids = df_tm_ans[df_tm_ans['Dominant Topic'] == topic_id]['link'].tolist()
for i in range(len(ans_ids)):
ans_ids[i] = linkToId(ans_ids[i])
MAP_tm_id_ansid[topic_id] = ans_ids
save_obj(MAP_tm_id_ansid, "MAP_tm_id_ansid")
# + [markdown] id="f0s_eILLwq5w"
# #### Topic Categorization
# + [markdown] id="XZTKCIPnw1Ij"
# ##### Parse TM labeling file
# + colab={"base_uri": "https://localhost:8080/"} id="vtAEivX9KCbR" outputId="a80796ef-1d0f-4440-d589-1ebd2c553008"
input_file = os.path.join(ROOT_dir, "dataset", "TM_label.csv")
df_tm_label = pd.read_csv(input_file)
print(df_tm_label.columns)
df_tm_label = pd.read_csv(input_file).dropna( subset=['Id', "Topic_name"])
print(len(df_tm_label))
# print(df_tm_label)
# + colab={"base_uri": "https://localhost:8080/"} id="_QWEqIg3Mj0u" outputId="898fbfb7-eba0-4e9c-ae00-51a7ffbd73e3"
TMs = []
for index, row in df_tm_label.iterrows():
id = int(row["Id"])
topic_name = row["Topic_name"]
merged_id = row["Merged Topic"]
low_cat = row["Lower Cat"]
mid_cat = row["Middle Cat"]
high_cat = row["Higher Cat"]
if(math.isnan(merged_id)):
merged_id = -2
merged_id = int(merged_id)
# print(id)
# print(merged_id)
if(merged_id == -1):
# print(math.isnan(merged_id))
print("Id %d is merged" % (id))
continue
tm = TM()
tm.name = topic_name
tm.ids.append(id)
if(merged_id != -2):
tm.ids.append(merged_id)
tm.low_cat = low_cat
tm.mid_cat = mid_cat
tm.high_cat = high_cat
TMs.append(tm)
print("name: %s and ids: %s" % (tm.name, tm.ids))
# print(tm)
# break
save_obj(TMs, "TMs")
# + [markdown] id="_jxlt1lx6-4k"
# #### Map Questions to Categories and samples
# + colab={"base_uri": "https://localhost:8080/"} id="TWHMEGPTSy2w" outputId="16e053cf-fa09-4f9c-b91a-eb7daee0f297"
import random
print(len(TMs))
total = 0
Map_sample = {}
for high_cat in High_cats:
# print(high_cat)
ids = []
topics = 0
for tm in TMs:
if tm.high_cat == high_cat:
topics += 1
ids += tm.ids
print(ids)
questions = []
for id in ids:
total_questions += len(MAP_tm_id_quid[id])
questions += MAP_tm_id_quid[id]
total += len(questions)
sample = random.sample(questions, Sample_size[high_cat])
assert len(sample) == len(set(sample))
assert len(sample) == Sample_size[high_cat]
Map_sample[high_cat] = sample
Map_cat_ques[high_cat] = questions
print("High Cat: %s Topics: %d and topics_ids: %d total_ques: %d" % (high_cat, topics, len(ids), len(questions)))
print(sample)
print(total)
save_obj(Map_sample, "Map_sample")
save_obj(Map_cat_ques, "Map_cat_ques")
# + colab={"base_uri": "https://localhost:8080/"} id="cqzAsnr46fa_" outputId="55c5327d-85dd-48dd-92eb-40a6510e18ea"
for high_cat in High_cats:
print("%s Categories has %d questions" % (high_cat, len(Map_cat_ques[high_cat])))
# + [markdown] id="EaZwvdxAN6CE"
# #### Map cat to topic
# + colab={"base_uri": "https://localhost:8080/"} id="ILyDiKVBN-05" outputId="5393f26f-253a-4c6c-e3a7-bddfc044539f"
Map_cat_topic = {}
for high_cat in High_cats:
Map_cat_topic[high_cat] = []
for tm in TMs:
Map_cat_topic[tm.high_cat].append(tm)
for high_cat in High_cats:
print("%s => #%d" % (high_cat, len(Map_cat_topic[high_cat])))
save_obj(Map_cat_topic, "Map_cat_topic")
# print("%s Categories has %d questions" % (high_cat, len(Map_cat_ques[high_cat])))
# + [markdown] id="7_6x95hxxExM"
# #### Create labeling sample
# + colab={"base_uri": "https://localhost:8080/"} id="uUpig-xIrVdK" outputId="ff33c9f4-a61b-4530-f2e2-5084fc1d5e1a"
input_file = os.path.join(ROOT_dir, "dataset", "MSR_SDLC.csv")
df_msr_sdlc = pd.read_csv(input_file)
print(len(df_msr_sdlc))
df_msr_sdlc = df_msr_sdlc[['Id', "SDLC"]]
print(df_msr_sdlc)
# + colab={"base_uri": "https://localhost:8080/"} id="bYQbXOHAse1D" outputId="f08100de-61bc-4d4f-e366-3662c8856b5d"
# df_merged = pd.merge(DF_sample, df_msr_sdlc, on="Id", how="left")
# print(len(df_merged))
# save_df(df_merged, File_Cat_sample)
# + colab={"base_uri": "https://localhost:8080/"} id="kFFnmaVuhD2_" outputId="22c656bb-0da4-4a06-b881-8664c3e85687"
DF_question_nss_final_tag = load_obj("DF_question_nss_final_tag")
print(len(DF_question_nss_final_tag))
sample_ques_ids = []
for high_cat in High_cats:
print("High Cat: %s total_ques: %d" % (high_cat, len(Map_sample[high_cat])))
sample_ques_ids += Map_sample[high_cat]
print(len(sample_ques_ids))
DF_question_nss_final_tag['Id'] = DF_question_nss_final_tag['Id'].astype(int)
DF_sample = DF_question_nss_final_tag[DF_question_nss_final_tag['Id'].isin(sample_ques_ids)]
print(len(DF_sample))
df_merged = pd.merge(DF_sample, df_msr_sdlc, on="Id", how="left")
save_obj(df_merged, "DF_sample")
save_df(df_merged, File_Cat_sample)
# + id="jHPOAyFUl4YK"
print(sample_ques_ids)
DF_sample = DF_question_nss_final_tag[DF_question_nss_final_tag['Id'] == "12556"]
print(DF_sample)
# print(type(sample_ques_ids[0]))
# print(type(DF_question_nss_final_tag.Id))
# DF_question_nss_final_tag.Id
# + colab={"base_uri": "https://localhost:8080/"} id="IDrWVvw5lNO2" outputId="d8b540c9-1cf6-4d30-8602-38439c102fe7"
print(len(DF_question_nss_final_tag))
# + [markdown] id="3Ibgb8It7mvF"
# #### TMs statistics
# + id="-MilT2CK7l8k"
# Total_Questions = 26763
# print(len(TMs))
# for tm in TMs:
# print(tm.ids, tm.name)
# + colab={"base_uri": "https://localhost:8080/"} id="LesHIPM0gs7h" outputId="ba3e0b80-d6ff-4259-8ae2-c27a2a4d1ac9"
a = [1, 2, 3]
b = a
a = []
b[0] = 100
print(b, a)
# + [markdown] id="LrD2HbYuxJfh"
# Generate merged TM stat
# + id="sXaoHqZg8amR"
MAP_merged_tm_quid = {}
t = 0
for tm in TMs:
qids = []
for id in tm.ids:
qids += MAP_tm_id_quid[id]
t += len(qids)
pct = round(len(qids) * 100.0 / Total_Questions, 1)
print("%s %s %d %.1f" % (tm.ids, tm.name, len(qids), pct))
MAP_merged_tm_quid[tm.name] = qids
assert t == Total_Questions
save_obj(MAP_merged_tm_quid, "MAP_merged_tm_quid")
# print(tm.ids, tm.name)
# + [markdown] id="evWjQsuSF2oZ"
# Merged TM stat with answers
#
# + colab={"base_uri": "https://localhost:8080/"} id="MnWoPvU3F5-z" outputId="53dba10b-c0c8-4972-b52f-15f19eae1b18"
MAP_merged_tm_ansid = {}
t = 0
for tm in TMs:
ans_ids = []
for id in tm.ids:
ans_ids += MAP_tm_id_ansid[id]
t += len(ans_ids)
pct = round(len(ans_ids) * 100.0 / Total_Ansers, 1)
print("%s %s %d %.1f" % (tm.ids, tm.name, len(ans_ids), pct))
MAP_merged_tm_ansid[tm.name] = ans_ids
assert t == Total_Ansers
save_obj(MAP_merged_tm_ansid, "MAP_merged_tm_ansid")
# print(tm.ids, tm.name)
# + [markdown] id="k0Emiqtaf-Ue"
# Initialization code
# + [markdown] id="bOjX3eAp0ldD"
# ## RQ Topic Evolution
# + colab={"base_uri": "https://localhost:8080/"} id="1Ly9u5Is05_c" outputId="10d051fa-3f48-44b3-82eb-bac38352b9c6"
Tag_list_final = load_obj("Tag_list_final")
# + [markdown] id="CElL9LtW08sy"
# ### Absolute impact
# + [markdown] id="mH4qZ55VtqOO"
# #### 5 Topic categories
# + id="OzrDUJsL1D5z"
SAMPLING = '1MS'
DUMMY_DATE = pd.to_datetime('2008-8-1 18:18:37.777')
Map_cat_ques_df = {}
df_all_absolute = All_Questions_df[['Id', 'CreationDate']]
df_all_absolute.CreationDate = df_all_absolute.CreationDate.apply(pd.to_datetime)
df_all_absolute = df_all_absolute.resample(SAMPLING, on="CreationDate").count()[["Id"]]
df_all_absolute.columns = ['All']
# print(df_all.head())
for high_cat in High_cats:
# print("%s Categories has %d questions" % (high_cat, len(Map_cat_ques[high_cat])))
qids = Map_cat_ques[high_cat]
df_cat = All_Questions_df[All_Questions_df['Id'].astype(int).isin(qids)]
print("%s ==========> %d" % (high_cat, len(df_cat)))
# df_cat = Map_cat_ques_df['Customization']
df_cat.CreationDate = df_cat.CreationDate.apply(pd.to_datetime)
df_cat = df_cat.append(pd.Series(data=[DUMMY_DATE], index = ['CreationDate']), ignore_index=True)
df_cat = df_cat.resample(SAMPLING, on='CreationDate').count()[["Id"]]
df_cat.columns = [high_cat]
df_all_absolute = df_all_absolute.merge(df_cat, how='left', on='CreationDate', validate='one_to_one')
Map_cat_ques_df[high_cat] = df_cat
print(df_all_absolute.head())
# for high_cat in High_cats:
# print("%s Categories has %d questions" % (high_cat, len(Map_cat_ques_df[high_cat])))
# + colab={"base_uri": "https://localhost:8080/"} id="i2R2mCB7zdXu" outputId="9b2c7fac-8dde-468f-af82-2d217c557eae"
file_name = os.path.join(ROOT_dir, "Output", "category_evolution_monthly.csv")
# save_df(df_all_absolute, file_name)
df_all_absolute.to_csv(file_name, quotechar='"', quoting=csv.QUOTE_NONNUMERIC)
# + [markdown] id="GY5IckEm11os"
# Absolute topic impact
# + id="yC_9YX8H6yY2"
# + [markdown] id="zAGj_AY_6FqZ"
# #### All LCSD posts evolution
# + colab={"base_uri": "https://localhost:8080/", "height": 435} id="wpM9rHtn0PHe" outputId="3393e30c-9f1e-48a4-c352-099e36e60a5f"
file_name = os.path.join(ROOT_dir, "Output", "All_posts_absolute_impact.png")
df = df_all_absolute[['All']]
df.columns = ['LCSD Posts']
ax = df.plot(figsize=(10,6))
ax.xaxis.label.set_visible(False)
# plt.xlabel('Time')
plt.ylabel('# of Questions')
labels = [x.strftime("%b, %Y") for x in df_all_absolute.index]
ax.set_xticks(labels)
ax.set_xticklabels(labels, rotation=90)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.title.set_visible(False)
# plt.axis('off')
# plt.grid()
plt.savefig(file_name, dpi=1000, bbox_inches = "tight")
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 435} id="InzKuGQg16D9" outputId="d9e8864b-86a1-4364-8114-83e0c79af32c"
file_name = os.path.join(ROOT_dir, "Output", "Topic_absolute_impact.png")
ax = df_all_absolute.drop(columns='All').plot(figsize=(10,6))
ax.xaxis.label.set_visible(False)
# plt.xlabel('Time')
plt.ylabel('# of Questions')
labels = [x.strftime("%b, %Y") for x in df_all_absolute.index]
ax.set_xticks(labels)
ax.set_xticklabels(labels, rotation=90)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
# plt.grid()
plt.savefig(file_name, dpi=1000, bbox_inches = "tight")
plt.show()
# + [markdown] id="7SRLY5fntlFY"
# #### 40 topics
# + id="4X7gzH22aBgb"
# + id="mVHZ3dzfzZd-"
# for topic in MAP_merged_tm_quid:
# print(topic, len(MAP_merged_tm_quid[topic]))
for cat in Map_cat_topic:
print(cat, Map_cat_topic[cat])
for tm in Map_cat_topic[cat]:
print(tm.name, len(tm.ids))
# + colab={"base_uri": "https://localhost:8080/"} id="EcuGbTC_z2sl" outputId="c3b4209b-151c-4216-881e-0a0138ade6ac"
def plot_dataframe(df, file_name, xlablel="Time"):
ax = df.drop(columns='All').plot(figsize=(10,6))
ax.xaxis.label.set_visible(False)
plt.xlabel(xlablel)
plt.ylabel('# of Questions')
labels = [x.strftime("%b, %Y") for x in df.index]
ax.set_xticks(labels)
ax.set_xticklabels(labels, rotation=90)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
plt.savefig(file_name, dpi=1000, bbox_inches = "tight")
plt.show()
def get_initial_df():
df = All_Questions_df[['Id', 'CreationDate']]
df.CreationDate = df.CreationDate.apply(pd.to_datetime)
df = df.resample(SAMPLING, on="CreationDate").count()[["Id"]]
df.columns = ['All']
return df
# + colab={"base_uri": "https://localhost:8080/"} id="HVd1Oe_F9yiO" outputId="f63ef77d-81ea-4a0d-a238-b47b0a477a25"
pd.options.mode.chained_assignment = None
# + [markdown] id="5M1LBeDcZ2jP"
# ### Covid19
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="Br1tizxBtxNl" outputId="35df78eb-3f7c-4420-bad5-efcaffde6352"
SAMPLING = '3MS'
file_name = os.path.join(ROOT_dir, "Output", "topic_evolution.csv")
# fig_file_name = os.path.join(ROOT_dir, "Output", "Topics_absolute_impact.png")
for cat in Map_cat_topic:
# print(cat, Map_cat_topic[cat])
df = get_initial_df()
for tm in Map_cat_topic[cat]:
topic = tm.name
# for topic in MAP_merged_tm_quid:
# print("%s Categories has %d questions" % (high_cat, len(Map_cat_ques[high_cat])))
qids = MAP_merged_tm_quid[topic]
df_cat = All_Questions_df[All_Questions_df['Id'].astype(int).isin(qids)]
# print("%s ==========> %d" % (topic, len(df_cat)))
# df_cat = Map_cat_ques_df['Customization']
df_cat.CreationDate = df_cat.CreationDate.apply(pd.to_datetime)
df_cat = df_cat.append(pd.Series(data=[DUMMY_DATE], index = ['CreationDate']), ignore_index=True)
df_cat = df_cat.resample(SAMPLING, on='CreationDate').count()[["Id"]]
df_cat.columns = [topic]
df = df.merge(df_cat, how='left', on='CreationDate', validate='one_to_one')
Map_cat_ques_df[topic] = df_cat
# print(df.head())
# df.to_csv(file_name, quotechar='"', quoting=csv.QUOTE_NONNUMERIC)
fig_file_name = "%s_absolute_impact.png" % cat
fig_file_name = os.path.join(ROOT_dir, "Output", fig_file_name)
plot_dataframe(df, fig_file_name, cat)
# break
# + [markdown] id="Qa0EBBVCZ9Fn"
# #### Prepandemic vs post pandemic
# + colab={"base_uri": "https://localhost:8080/"} id="R7vxacs5aAd5" outputId="1386f9fc-a4c7-4703-81d8-3486911cf1ef"
df_pandemic = pd.DataFrame()
# pre_pandemic = []
# post_pandemic = []
for high_cat in High_cats:
# print("%s Categories has %d questions" % (high_cat, len(Map_cat_ques[high_cat])))
qids = Map_cat_ques[high_cat]
df_cat = All_Questions_df[All_Questions_df['Id'].astype(int).isin(qids)]
print("%s ==========> %d" % (high_cat, len(df_cat)))
# # df_cat = Map_cat_ques_df['Customization']
df_cat.CreationDate = df_cat.CreationDate.apply(pd.to_datetime)
df_pre_pandemic = len(df_cat[(df_cat['CreationDate'] > '2018-03-01') & (df_cat['CreationDate'] < '2020-2-29')]) / 24
df_post_pandemic = len(df_cat[(df_cat['CreationDate'] > '2020-03-01') & (df_cat['CreationDate'] < '2021-04-30')]) / 14
dic = {}
dic["Cat"] = high_cat
dic["Pre_pandemic"] = int(df_pre_pandemic)
dic["Post_pandemic"] = int(df_post_pandemic)
df_pandemic = df_pandemic.append(pd.Series(dic), ignore_index = True)
# last_month = len(df_cat[(df_cat['CreationDate'] > '2021-03-01') & (df_cat['CreationDate'] < '2026-3-31')])
# print(last_month)
# pre_pandemic.append(int(df_pre_pandemic))
# post_pandemic.append(int(df_post_pandemic))
# print(df_pre_pandemic)
# print(df_post_pandemic)
# break
print(df_pandemic)
# + colab={"base_uri": "https://localhost:8080/"} id="GwesXn7tpac9" outputId="93dc8a8b-358c-4c7c-85c1-316929232af8"
file_name = os.path.join(ROOT_dir, "output", "pandemic.csv")
df_pandemic.to_csv(file_name, index=False, quotechar='"', quoting=csv.QUOTE_NONNUMERIC)
# + [markdown] id="XFXjs3dPtuM9"
# #### Tags
# + colab={"base_uri": "https://localhost:8080/"} id="xnxktOWUck1W" outputId="c6a15933-08e5-4d67-ce59-9a698d9e02ff"
a = [1, 3]
b = [ 4, 5]
c = a + b
print(c)
# + id="VNQTTrhJAAC2"
for tag in sorted(Map_tag_qids, key = lambda topic: len(Map_tag_qids[topic]), reverse=True):
print(tag, len(Map_tag_qids[tag]))
# + colab={"base_uri": "https://localhost:8080/", "height": 728} id="FLiRiD5ntx4B" outputId="c7f3e2fc-23d9-49c2-a2df-6231c8158ffb"
SAMPLING = '3MS'
file_name = os.path.join(ROOT_dir, "Output", "tag_evolution.csv")
fig_file_name = os.path.join(ROOT_dir, "Output", "Tag_absolute_impact.png")
df = get_initial_df()
count = 0
for tag in sorted(Map_tag_qids, key = lambda topic: len(Map_tag_qids[topic]), reverse=True):
qids = Map_tag_qids[tag]
print(tag, len(qids))
df_cat = All_Questions_df[All_Questions_df['Id'].astype(int).isin(qids)]
# print("%s ==========> %d" % (topic, len(df_cat)))
# df_cat = Map_cat_ques_df['Customization']
df_cat.CreationDate = df_cat.CreationDate.apply(pd.to_datetime)
df_cat = df_cat.append(pd.Series(data=[DUMMY_DATE], index = ['CreationDate']), ignore_index=True)
df_cat = df_cat.resample(SAMPLING, on='CreationDate').count()[["Id"]]
df_cat.columns = [tag]
df = df.merge(df_cat, how='left', on='CreationDate', validate='one_to_one')
count += 1
if(count > 15):
break
# df.to_csv(file_name, quotechar='"', quoting=csv.QUOTE_NONNUMERIC)
plot_dataframe(df, fig_file_name)
# break
# + [markdown] id="DY05PqYTd1UH"
# ##### Platform wise
# + colab={"base_uri": "https://localhost:8080/"} id="s-JSI5fEX3HN" outputId="37e41691-4452-4e29-d7c3-35738e56b715"
PLATFORMS = {'Salesforce': ['salesforce', 'visualforce', 'salesforce-lightning', 'salesforce-service-cloud', 'lwc',
'salesforce-communities', 'salesforce-marketing-cloud', 'salesforce-chatter',
'salesforce-development', 'salesforce-ios-sdk'],
"Oracle Apex": ['apex-code', 'apex-code', 'apex', 'oracle-apex-5', 'oracle-apex-5.1', 'oracle-apex-19.1', 'oracle-apex-18.2', 'apex-trigger'],
"Lotus Software": ['lotus-domino', 'lotus-notes'],
'Filemaker': ['filemaker'],
'Microsoft PowerApps': ['powerapps', 'powerapps-canvas', 'powerapps-formula', 'powerapps-modeldriven', 'powerapps-selected-items', 'powerapps-collection'],
'Service Now': ['servicenow', 'servicenow-rest-api'],
'Tibco': ['tibco'],
'Zoho Creator': ['zoho'],
'OutSystems': ['outsystems'],
'Pega': ['pega'],
'Process Maker': ['processmaker'],
'Mendix': ['mendix']
}
Map_Platfoms_quids = {}
for platform in PLATFORMS:
Map_Platfoms_quids[platform] = []
for tag in PLATFORMS[platform]:
Map_Platfoms_quids[platform] = Map_Platfoms_quids[platform] + Map_tag_qids[tag]
# for platform in PLATFORMS:
# print(platform, len(Map_Platfoms_quids[platform]))
for platform in sorted(Map_Platfoms_quids, key = lambda platform: len(Map_Platfoms_quids[platform]), reverse=True):
print(platform, len(Map_Platfoms_quids[platform]))
save_obj(Map_Platfoms_quids, "Map_Platfoms_quids")
# + colab={"base_uri": "https://localhost:8080/", "height": 644} id="NriWtuSSd6uk" outputId="888620fe-a9b6-4440-c071-460ce3a47982"
SAMPLING = '3MS'
file_name = os.path.join(ROOT_dir, "Output", "platform_evolution.csv")
fig_file_name = os.path.join(ROOT_dir, "Output", "platform_absolute_impact.png")
df = get_initial_df()
count = 0
for platform in sorted(Map_Platfoms_quids, key = lambda topic: len(Map_Platfoms_quids[topic]), reverse=True):
qids = Map_Platfoms_quids[platform]
print(platform, len(qids))
df_cat = All_Questions_df[All_Questions_df['Id'].astype(int).isin(qids)]
# print("%s ==========> %d" % (topic, len(df_cat)))
# df_cat = Map_cat_ques_df['Customization']
df_cat.CreationDate = df_cat.CreationDate.apply(pd.to_datetime)
df_cat = df_cat.append(pd.Series(data=[DUMMY_DATE], index = ['CreationDate']), ignore_index=True)
df_cat = df_cat.resample(SAMPLING, on='CreationDate').count()[["Id"]]
df_cat.columns = [platform]
df = df.merge(df_cat, how='left', on='CreationDate', validate='one_to_one')
# df.to_csv(file_name, quotechar='"', quoting=csv.QUOTE_NONNUMERIC)
plot_dataframe(df, fig_file_name)
# break
# + [markdown] id="zS_I1Jyw0_Ul"
# ### Relative impact
# + colab={"base_uri": "https://localhost:8080/", "height": 527} id="7UFvCV4_4e7u" outputId="cddc665b-640d-453c-e4db-88b9630418dd"
df_all_relative = df_all_absolute.copy(deep=True)
for high_cat in High_cats:
df_all_relative[high_cat] = df_all_relative[high_cat] / df_all_relative['All']
# print(df_all_relative.head())
file_name = os.path.join(ROOT_dir, "Output", "Topic_relative_impact.png")
ax = df_all_relative.drop(columns='All').plot(figsize=(10,6))
ax.xaxis.label.set_visible(False)
# plt.xlabel('Time')
plt.ylabel('# of Questions')
y_labels = [ str(i)+"%" for i in range(0, 51, 10)]
labels = [x.strftime("%b, %Y") for x in df_all_relative.index]
print(type(labels[0]))
print(labels)
ax.set_xticks(labels)
ax.set_xticklabels(labels, rotation=90)
# ax.set_yticks(y_labels)
ax.set_yticklabels(y_labels)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
# plt.grid()
plt.savefig(file_name, dpi=1000, bbox_inches = "tight")
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="dqHn-Zet1EUZ" outputId="bf6801f0-cca2-4cec-d2cf-c0d19d2c6494"
print(len(All_Questions_df))
# + [markdown] id="M2xW-yemfnHD"
# ## RQ: Popularity difficulty
# + colab={"base_uri": "https://localhost:8080/"} id="JORuNebWnbCO" outputId="6b260d4a-37a1-436e-c06a-b0811e439069"
class Info:
def __init__(self):
self.view = 0.0
self.favorite = 0.0
self.score = 0.0
self.viewN = 0.0
self.favoriteN = 0.0
self.scoreN = 0.0
self.fusedP = 0.0
def __str__(self):
return ("View: %.2f, Favorite: %.2f, Score: %.2f, ViewN: %.2f, FavoriteN: %.2f, ScoreN: %.2f, FusedP: %.2f"
% ( self.view, self.favorite, self.score, self.viewN, self.favoriteN, self.scoreN, self.fusedP))
class InfoD:
def __init__(self):
self.pct_qwo_acc = 0.0
self.med_hours = 0.0
self.pct_qwo_accN = 0.0
self.med_hoursN = 0.0
self.fusedD = 0.0
def __str__(self):
return ("pct_qwo_acc: %.2f, med_hours: %.2f, pct_qwo_accN: %.2f, med_hours: %.2f, fusedD: %.2f"
% ( self.pct_qwo_acc, self.med_hours, self.pct_qwo_accN, self.med_hoursN, self.fusedD))
# + colab={"base_uri": "https://localhost:8080/"} id="VfSQggR155tD" outputId="2d4a946b-1238-437a-90e7-539a55d292b6"
a = Info()
a.view = 5
b = Info ()
b.view = 10
print(a)
# print(a.view, b.view)
# + [markdown] id="6fLLRIW4j_4c"
# ### Popularity
# + id="tQJdm6Qnj_Td"
View_mean_sum = 0
Favorite_mean_sum = 0
Score_mean_sum = 0
Map_topic_pop_diff = {}
for topic in MAP_merged_tm_quid:
quids = MAP_merged_tm_quid[topic]
# print(quids)
# print(type(quids[0]))
df = DF_question_nss_final_tag[DF_question_nss_final_tag['Id'].astype(int).isin(quids)]
# print(len(df))
info = Info()
info.view = round(pd.to_numeric(df['ViewCount']).fillna(0).astype(int).mean(), 1)
info.favorite = round(pd.to_numeric(df['FavoriteCount']).fillna(0).astype(int).mean(), 1)
info.score = round(pd.to_numeric(df['Score']).fillna(0).astype(int).mean(), 1)
# print(info)
View_mean_sum += info.view
Favorite_mean_sum += info.favorite
Score_mean_sum += info.score
Map_topic_pop_diff[topic] = info
assert len(quids) == len(df)
# break
print("view_sum: %.2f favorite_sum: %.2f score_sum: %.2f" % (View_mean_sum, Favorite_mean_sum, Score_mean_sum))
for topic in MAP_merged_tm_quid:
info = Map_topic_pop_diff[topic]
info.viewN = info.view * 40.0 / View_mean_sum
info.favoriteN = info.favorite * 40.0 / Favorite_mean_sum
info.scoreN = info.score * 40.0 / Score_mean_sum
info.fusedP = round((info.viewN + info.favoriteN + info.scoreN) / 3.0, 2)
# for topic in MAP_merged_tm_quid:
# print(Map_topic_pop_diff[topic])
for key in sorted(Map_topic_pop_diff, key = lambda topic: Map_topic_pop_diff[topic].fusedP, reverse=True):
print(Map_topic_pop_diff[key])
save_obj(Map_topic_pop_diff, "Map_topic_pop_diff")
# + [markdown] id="L8LboNkS1AQ_"
# #### Generate topic popularity table
# + colab={"base_uri": "https://localhost:8080/"} id="BJ64two8PQnS" outputId="873520fa-1f11-4837-88f7-0d82753aca40"
def get_high_cat(topic):
for tm in TMs:
if tm.name == topic:
return tm.high_cat
return None
# + id="isoREvUS0NqC"
DF_pop = pd.DataFrame()
topics = []
categories = []
fusedPs = []
scores = []
favorites = []
views = []
for key in sorted(Map_topic_pop_diff, key = lambda topic: Map_topic_pop_diff[topic].fusedP, reverse=True):
info = Map_topic_pop_diff[key]
topics.append(key)
categories.append(get_high_cat(key))
fusedPs.append(info.fusedP)
views.append(info.view)
favorites.append(info.favorite)
scores.append(info.score)
DF_pop['Topic'] = topics
DF_pop['Category'] = categories
DF_pop['FusedP'] = fusedPs
DF_pop['#View'] = views
DF_pop['#Favorite'] = favorites
DF_pop['#Score'] = scores
print(df_pop)
save_obj(DF_pop, "DF_pop")
file_name = os.path.join(ROOT_dir, "output", "Topic_popularity.csv")
save_df(DF_pop, file_name=file_name)
# + colab={"base_uri": "https://localhost:8080/"} id="lnEw9lqTf9AK" outputId="7cc9f4ae-5794-4d52-dbdf-4ed963f1a444"
TMs[0].name
# + [markdown] id="3EdAYm84uvrR"
# ### Difficulty
# + colab={"base_uri": "https://localhost:8080/"} id="Fn_1n2ffIsk9" outputId="62dc2075-baaa-4cb6-dfba-0dac0521ac7c"
Map_topic_diff = {}
# for topic in MAP_merged_tm_quid:
# Map_topic_diff[topic] = InfoD()
# + colab={"base_uri": "https://localhost:8080/"} id="CMvXsEG10DQs" outputId="1e986260-f308-4cfc-87d1-8f22c3c1db1e"
def get_acc_time(ques_id):
question = DF_question_nss_final_tag[DF_question_nss_final_tag.Id == str(ques_id)].iloc[0]
# print(type(question))
question_time = pd.to_datetime(question.CreationDate)
# print(type(question_time))
# print("question_time: %s" % (question_time) )
answer_id = question['AcceptedAnswerId']
answer = DF_answer_nss_final_tag[DF_answer_nss_final_tag.Id == str(answer_id)].iloc[0]
answer_time = pd.to_datetime(answer.CreationDate)
# print("answer_time: %s" % (answer_time) )
time_taken = (answer_time - question_time) / (pd.Timedelta(hours=1))
# print(time_taken)
return time_taken
# print(question_time, answer_time, time_taken)
print(get_acc_time(22940521))
# + [markdown] id="MF_scCwg1KCT"
# #### Generate topic difficulty table
# + [markdown] id="nheCP2iVIcct"
# Get questions ids with acc
# + colab={"base_uri": "https://localhost:8080/"} id="rIvRwhpPvhHq" outputId="41fe57c5-4b68-4152-8566-3bc8ede0c0d9"
df_ques_acc = DF_question_nss_final_tag[DF_question_nss_final_tag['AcceptedAnswerId'].astype(str).str.len() > 0]
print(len(df_ques_acc))
set_ques_acc_ids = set(df_ques_acc['Id'].tolist())
# DF_question_nss_final_tag.dtypes
# + id="Wr1A8ixoFjZq"
import statistics
pct_qwo_acc_sum = 0.0
med_hours_sum = 0.0
for topic in MAP_merged_tm_quid:
quids = MAP_merged_tm_quid[topic]
time_taken_list = []
acc_count = 0
for q in quids:
q = str(q)
# print(q)
# print(type(q))
if str(q) in set_ques_acc_ids:
acc_count += 1
time = get_acc_time(q)
time_taken_list.append(time)
# print(time)
# break
info = InfoD()
info.pct_qwo_acc = round(((len(quids) - acc_count)) * 100.0 / len(quids), 0)
info.med_hours = statistics.median(time_taken_list)
pct_qwo_acc_sum += info.pct_qwo_acc
med_hours_sum += info.med_hours
Map_topic_diff[topic] = info
# print(info)
# break
for topic in MAP_merged_tm_quid:
info = Map_topic_diff[topic]
info.pct_qwo_accN = info.pct_qwo_acc * 40.0 / pct_qwo_acc_sum
info.med_hoursN = info.med_hours * 40.0 / med_hours_sum
# info.scoreN = info.score * 40.0 / Score_mean_sum
info.fusedD = round((info.pct_qwo_accN + info.med_hoursN) / 2.0, 2)
for key in sorted(Map_topic_diff, key = lambda topic: Map_topic_diff[topic].fusedD, reverse=True):
print(Map_topic_diff[key])
save_obj(Map_topic_diff, "Map_topic_diff")
# + colab={"base_uri": "https://localhost:8080/"} id="OQ-KK3YUTPMb" outputId="7da1a4ae-f164-4a92-abdb-4510d5aa3500"
DF_diff = pd.DataFrame()
topics = []
categories = []
fusedDs = []
pct_qwo_acc_list = []
med_hours_list = []
for key in sorted(Map_topic_diff, key = lambda topic: Map_topic_diff[topic].fusedD, reverse=True):
print(Map_topic_diff[key])
info = Map_topic_diff[key]
topics.append(key)
categories.append(get_high_cat(key))
fusedDs.append(info.fusedD)
med_hours_list.append(info.med_hours)
pct_qwo_acc_list.append(info.pct_qwo_acc)
# favorites.append(info.favorite)
# scores.append(info.score)
DF_diff['Topic'] = topics
DF_diff['Category'] = categories
DF_diff['FusedD'] = fusedDs
DF_diff['Med Hrs. to Acc'] = med_hours_list
DF_diff['W/O Acc.'] = pct_qwo_acc_list
# print(DF_diff)
save_obj(DF_diff, "DF_diff")
file_name = os.path.join(ROOT_dir, "output", "Topic_difficulty.csv")
save_df(DF_diff, file_name=file_name)
# + [markdown] id="8VfhZIs_YlwN"
# ### Statistical significant
# + colab={"base_uri": "https://localhost:8080/"} id="g2vubJxwaPUi" outputId="fe352117-f4c7-4c37-d7d4-3eed5e19bf43"
import scipy.stats as stats
Map_pop = {'view': [], 'favorite': [], 'score': []}
Map_diff = {'pct_wo_acc': [], 'med_hours': []}
for topic in MAP_merged_tm_quid:
infoP = Map_topic_pop_diff[topic]
infoD = Map_topic_diff[topic]
Map_pop['view'].append(infoP.viewN)
Map_pop['favorite'].append(infoP.favoriteN)
Map_pop['score'].append(infoP.scoreN)
Map_diff['pct_wo_acc'].append(infoD.pct_qwo_accN)
Map_diff['med_hours'].append(infoD.med_hoursN)
for diff in Map_diff:
for pop in Map_pop:
X1 = Map_pop[pop]
X2 = Map_diff[diff]
tau, p_value = stats.kendalltau(X1, X2)
issig = False
if p_value <= 0.05:
issig = True
print ("%s => %s: %.2f/%.5f %s" % (pop, diff, tau, p_value, issig))
print("# " * 15)
for diff in Map_diff:
for pop in Map_pop:
X1 = Map_pop[pop]
X2 = Map_diff[diff]
tau, p_value = stats.kendalltau(X1, X2)
issig = False
if p_value <= 0.05:
issig = True
print ("%.2f/%.2f & " % ( tau, p_value), end="")
print()
# + [markdown] id="np_XyKZLout7"
# ### Bubble charts
# + id="25J_AsOopc4w"
topics = []
total_questions = []
fusedP = []
fusedD = []
for topic in MAP_merged_tm_quid:
topics.append(topic)
total_questions.append(len(MAP_merged_tm_quid[topic]))
fusedP.append(Map_topic_pop_diff[topic].fusedP)
fusedD.append(Map_topic_diff[topic].fusedD)
df_topic_bubble = pd.DataFrame()
df_topic_bubble['topics'] = topics
df_topic_bubble['total_questions'] = total_questions
df_topic_bubble['fusedP'] = fusedP
df_topic_bubble['fusedD'] = fusedD
print(df_topic_bubble)
file_name = os.path.join(ROOT_dir, "output", "topic_bubble.csv")
save_df(df_topic_bubble, file_name)
save_obj(df_topic_bubble, "df_topic_bubble")
# + colab={"base_uri": "https://localhost:8080/"} id="uh7dKGOFoyVi" outputId="df83ace7-d5bd-414b-dee8-8bd46c062137"
categories = []
total_questions = []
fusedP = []
fusedD = []
for cat in Map_cat_topic:
categories.append(cat)
q_total = 0
fusedP_total = 0
fusedD_total = 0
for tm in Map_cat_topic[cat]:
# print(tm.name)
q_total += len(Map_cat_ques[cat])
fusedP_total += Map_topic_pop_diff[tm.name].fusedP
fusedD_total += Map_topic_diff[tm.name].fusedD
total_questions.append(q_total)
fusedP.append(fusedP_total)
fusedD.append(fusedD_total)
df_cat_bubble = pd.DataFrame()
df_cat_bubble['Categories'] = categories
df_cat_bubble['total_questions'] = total_questions
df_cat_bubble['fusedP'] = fusedP
df_cat_bubble['fusedD'] = fusedD
print(df_cat_bubble)
file_name = os.path.join(ROOT_dir, "output", "category_bubble.csv")
save_df(df_cat_bubble, file_name)
save_obj(df_cat_bubble, "df_cat_bubble")
# + [markdown] id="Kqmlzekx2mhf"
# ### Table of topic stats
#
# + colab={"base_uri": "https://localhost:8080/"} id="RFCblRWX2qoV" outputId="8acfd242-0569-4942-c0f1-131c8ab7828d"
df_topic_stat = pd.DataFrame()
topics = []
num_ques = []
num_ans = []
for topic in MAP_merged_tm_quid:
topics.append(topic)
num_ques.append(len(MAP_merged_tm_quid[topic]))
num_ans.append(len(MAP_merged_tm_ansid[topic]))
# print(topic)
df_topic_stat['Topic'] = topics
df_topic_stat['#Question'] = num_ques
df_topic_stat['#Answer'] = num_ans
file_name = os.path.join(ROOT_dir, "output", "topic_stats.csv")
save_df(df_topic_stat, file_name)
save_obj(df_topic_stat, "df_topic_stat")
print(len(df_topic_stat))
# + [markdown] id="ndHXaOxaCIvj"
# ## RQ types of questions
#
# + colab={"base_uri": "https://localhost:8080/"} id="UruoDsA3CLs2" outputId="4a1a0f37-eff2-44c3-95dd-d0ef71ed7ee3"
File_annotation = os.path.join(ROOT_dir, "dataset", "RQ_annotation.csv")
# + [markdown] id="ZNEcl5qbCUF1"
# ### Labeling file
# + colab={"base_uri": "https://localhost:8080/"} id="8Zt-ylUsCXyS" outputId="4e066065-c326-4d49-d89a-ffb163da8bbc"
df_rq_annotation = pd.read_csv(File_annotation).fillna("")
print(len(df_rq_annotation))
# + [markdown] id="2LYCsl4R_m5c"
# ### Category => type of questions
# + colab={"base_uri": "https://localhost:8080/"} id="Nt-DS_bvCu3S" outputId="cbdab0a8-83bf-4a97-9631-1f587004a614"
def print_map(map_tmp):
print(map_tmp)
total = 0
for qtype in map_tmp:
total += map_tmp[qtype]
for qtype in map_tmp:
pct = round(map_tmp[qtype] * 100.0 / total, 1)
print("Type: %s pct: %.1f" % (qtype, pct))
for qtype in map_tmp:
pct = round(map_tmp[qtype] * 100.0 / total, 1)
print(" %.1f\%% & " % (pct), end="")
print()
Map_type_to_question = {'How': 0, "What": 0, "Why": 0, "Others": 0}
Map_cat_to_type = {}
for high_cat in High_cats:
Map_cat_to_type[high_cat] = 0
ques_ids = Map_sample[high_cat]
map_tmp = {'How': 0, "What": 0, "Why": 0, "Others": 0}
for quid in ques_ids:
label = df_rq_annotation[df_rq_annotation['Id'] == quid].iloc[0]
type1 = label.Type
type2 = label.Type2
# print(type1)
# print(type(type1))
Map_type_to_question[type1] = Map_type_to_question[type1] + 1
map_tmp[type1] = map_tmp[type1] + 1
# print(type2)
if(len(type2) > 2):
Map_type_to_question[type2] = Map_type_to_question[type2] + 1
map_tmp[type2] = map_tmp[type2] + 1
# print(label.Type)
print("Category: %s and %s" % (high_cat, map_tmp))
print_map(map_tmp)
print("#" * 10)
# break
# print(Map_type_to_question)
print_map(Map_type_to_question)
# total = 0
# for qtype in Map_type_to_question:
# total += Map_type_to_question[qtype]
# for qtype in Map_type_to_question:
# pct = round(Map_type_to_question[qtype] * 100.0 / total, 0)
# print("Type: %s pct: %.1f" % (qtype, pct))
# save_obj(Map_question_type, "Map_question_type")
# + [markdown] id="RjF3B-1srxyx"
# ## RQ SDLC
# + colab={"base_uri": "https://localhost:8080/"} id="xkUdnwDEsAzP" outputId="ed10fda7-e941-436e-ef98-ab3a3b0d9159"
File_annotation = os.path.join(ROOT_dir, "dataset", "RQ_annotation.csv")
SDLC_phases = ["Requirement Analysis & Planning", "Application Design", "Implementation", "Testing", "Deployment", "Maintenance"]
# + [markdown] id="6qLk3WJLsB3J"
# Read annotation file
# + colab={"base_uri": "https://localhost:8080/"} id="rpHCRL_DsDuY" outputId="2283476b-e8f9-4d0f-bdec-d01e8f836019"
df_rq_annotation = pd.read_csv(File_annotation).fillna("")
print(len(df_rq_annotation))
df_SDLC = df_rq_annotation[['Id', 'SDLC']]
df_SDLC_type = df_rq_annotation[['Id', 'SDLC', 'Type', 'Type2']]
# + [markdown] id="tW_b3n-ZsNii"
# ### SDLC stats
# + colab={"base_uri": "https://localhost:8080/"} id="VdS44jeFuFLa" outputId="3dba545f-c398-493c-bace-02941ed58137"
dist = df_SDLC['SDLC'].value_counts(normalize=True)
print(type(dist))
print(dist)
# + colab={"base_uri": "https://localhost:8080/"} id="oanZtvj9GP7P" outputId="b7bc2127-bdf6-4da3-b26f-83a93e6c850a"
dist = df_SDLC['SDLC'].value_counts()
print(type(dist))
print(dist)
# + [markdown] id="DBX0rRFAzMGH"
# ### SDLC => Type
# + colab={"base_uri": "https://localhost:8080/"} id="B_Fb2Rm4sPzO" outputId="0e8ac8d1-d6c4-4547-b97c-42a1dd28c20e"
def get_stat_SDLC_type(df, label):
# print(len(df))
total = len(df)
s1 = df['Type'].value_counts()
s2 = df['Type2'].dropna().value_counts()
how_count = s1['How']
if "How" in s2:
how_count += s2['How']
what_count = s1['What']
if "What" in s2:
what_count += s2['What']
why_count = s1['Why']
if "Why" in s2:
why_count += s2['Why']
other_count = s1['Others']
if "Others" in s2:
other_count += s2['Others']
# print(other_count)
total_pct = round(total * 100.0/471, 0)
how_pct = round(how_count * 100.0/total, 0)
what_pct = round(what_count * 100.0/total, 0)
why_pct = round(why_count * 100.0/total, 0)
other_pct = round(other_count * 100.0/total, 0)
print("%s(%.0f\\%%) & %.0f\\%% & %.0f\\%% & %.0f\\%% & %.0f\\%% \\\\" % (label, total_pct, how_pct, what_pct, why_pct, other_pct))
for phase in SDLC_phases:
df = df_SDLC_type[df_SDLC_type['SDLC'] == phase]
get_stat_SDLC_type(df, phase)
# print(type(df))
# print(len(df))
# print(df.sample)
# get_stat_SDLC_type(df, "Requirement")
# + [markdown] id="pUD_l3m__eHE"
# ### Category => SDLC
# + [markdown] id="QEiWhqBJULPB"
#
# + colab={"base_uri": "https://localhost:8080/"} id="uHw_Witr_h3M" outputId="8e93c559-48da-46f2-c423-fa618dc67bfa"
def get_stat_cat_sdlc(df, label):
# print(len(df), label)
counts = df['SDLC'].value_counts()
print("\\textbf{%s} & \\sixbars" % (label), end="")
for phase in SDLC_phases:
val = 0
if phase in counts:
val = counts[phase]
print("{%d}" % val, end="")
print(" \\\\", end="")
print()
for high_cat in High_cats:
Map_cat_to_type[high_cat] = 0
ques_ids = Map_sample[high_cat]
df = df_rq_annotation[df_rq_annotation["Id"].isin(ques_ids)]
# print(len(df))
get_stat_cat_sdlc(df, high_cat)
# break
# map_tmp = {'How': 0, "What": 0, "Why": 0, "Others": 0}
# for quid in ques_ids:
# label = df_rq_annotation[df_rq_annotation['Id'] == quid].iloc[0]
# type1 = label.Type
# type2 = label.Type2
# # print(type1)
# # print(type(type1))
# Map_type_to_question[type1] = Map_type_to_question[type1] + 1
# map_tmp[type1] = map_tmp[type1] + 1
# # print(type2)
# if(len(type2) > 2):
# Map_type_to_question[type2] = Map_type_to_question[type2] + 1
# map_tmp[type2] = map_tmp[type2] + 1
# # print(label.Type)
# print("Category: %s and %s" % (high_cat, map_tmp))
# print_map(map_tmp)
# print("#" * 10)
# -
# ## Graph: Significance va Relevance of final tags
# +
df = pd.read_csv('data/tags_stat.csv')
df.head()
fields = ['tags_relevant', 'tags_recommended_count']
colors = ['#4C72B0', '#55A868']
labels = ['Total relevant', 'Total recomended']
# figure and axis
fig, ax = plt.subplots(1, figsize=(19.20, 10.80))
# plot bars
left = len(df) * [0]
for idx, name in enumerate(fields):
plt.barh(df.index, df[name], left=left, color=colors[idx], height=0.50)
left = left + df[name]
# title, legend, labels
plt.legend(labels, loc='upper right', frameon=False, prop={'family': 'serif'})
# remove spines
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
# Add y-axis labels
ytick_labels = []
for _, df_entry in df.iterrows():
ytick_labels.append(
f"\u03bc = {df_entry.significance}, \u03BD = {df_entry.relevance}: {df_entry.tags_relavant_pct}%")
plt.yticks(np.arange(0, len(df.index)), ytick_labels)
# adjust limits and draw grid lines
plt.ylim(-0.5, ax.get_yticks()[-1] + 0.5)
ax.set_axisbelow(True)
plt.savefig('figures/significane_relevance.png', dpi=300)
# -
# ## Graph Genration
# ### Evolution of top 10 platforms over time
# +
def plot_dataframe(df, file_name, xlablel="Time"):
ax = df.drop(columns='All').plot(figsize=(10,6))
ax.xaxis.label.set_visible(False)
plt.xlabel(xlablel)
plt.ylabel('# of Questions')
labels = [x.strftime("%b, %Y") for x in df.index]
ax.set_xticks(labels)
ax.set_xticklabels(labels, rotation=90)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
plt.savefig(file_name, dpi=1000, bbox_inches = "tight")
plt.show()
def get_initial_df(sampling='3MS'):
df = all_q_df[['Id', 'CreationDate']]
df.CreationDate = df.CreationDate.apply(pd.to_datetime)
df = df.resample(sampling, on="CreationDate").count()[["Id"]]
df.columns = ['All']
return df
SAMPLING = '4MS'
DUMMY_DATE = pd.to_datetime('2008-8-1 18:18:37.777')
df = get_initial_df(SAMPLING)
count = 0
for platform_name in top_10_plats_to_qid:
qids = top_10_plats_to_qid[platform_name]
# print(platform_name, len(qids))
df_cat = all_q_df[all_q_df['Id'].astype(int).isin(qids)]
# print("%s ==========> %d" % (platform_name, len(df_cat)))
df_cat.CreationDate = df_cat.CreationDate.apply(pd.to_datetime)
df_cat = df_cat.append(pd.Series(data=[DUMMY_DATE], index = ['CreationDate']), ignore_index=True)
df_cat = df_cat.resample(SAMPLING, on='CreationDate').count()[["Id"]]
df_cat.columns = [platform_name]
df = df.merge(df_cat, how='left', on='CreationDate', validate='one_to_one')
# df.to_csv(file_name, quotechar='"', quoting=csv.QUOTE_NONNUMERIC)
plot_dataframe(df, 'top5platform_evolution')
# -
# ### Category Distribution in top 5 platforms
# +
cat_to_qid = load_obj('map_cat_to_ques')
top_5_plats_to_qid = {}
for platform_name in sorted(plat_to_qid_df, key=lambda k: len(plat_to_qid_df[k]), reverse=True)[:5]:
top_5_plats_to_qid[platform_name] = plat_to_qid_df[platform_name]
# for key in top_5_plats_to_qid:
# print(key, len(top_5_plats_to_qid[key]))
platforms = []
categories = []
questions = []
for platform_name in top_5_plats_to_qid:
for category_name in cat_to_qid:
intersection = list(set(top_5_plats_to_qid[platform_name]) & set(cat_to_qid[category_name]))
# print(platform_name, category_name, len(intersection))
platforms.append(platform_name)
categories.append(category_name)
questions.append(len(intersection))
# print()
plat_to_cat_df = pd.DataFrame()
plat_to_cat_df['Platform'] = platforms
plat_to_cat_df['Category'] = categories
plat_to_cat_df['Count'] = questions
file_name = os.path.join("output", "Plat_To_Cateogory_Distribution.csv")
save_df(plat_to_cat_df, file_name=file_name)
cat_to_qid = load_obj('map_cat_to_ques')
for category_name in cat_to_qid:
print(category_name, len(cat_to_qid[category_name]))
base_q_df = load_obj('questions')
base_q_df['Id'] = base_q_df['Id'].astype(int)
base_ans_df = load_obj('answers')
def get_acc_time(ques_id):
question = base_q_df[base_q_df.Id == ques_id].iloc[0]
# print(type(question))
question_time = pd.to_datetime(question.CreationDate)
# print(type(question_time))
# print("question_time: %s" % (question_time) )
answer_id = question['AcceptedAnswerId']
# print(answer_id)
answer = base_ans_df[base_ans_df.Id == str(answer_id)].iloc[0]
answer_time = pd.to_datetime(answer.CreationDate)
# print("answer_time: %s" % (answer_time) )
time_taken = (answer_time - question_time) / (pd.Timedelta(hours=1))
# print(time_taken)
return time_taken
# print(question_time, answer_time, time_taken)
print(get_acc_time(22940521))
df_ques_acc = base_q_df[base_q_df['AcceptedAnswerId'].astype(str).str.len() > 0]
# print(len(df_ques_acc))
set_ques_acc_ids = set(df_ques_acc['Id'].tolist())
# -
# ### Overall popularity and diffuculty
# +
all_q_df = load_obj('questions')
base_ans_df = load_obj('answers')
print("# Posts (questions):", len(all_q_df))
avg_view = round(pd.to_numeric(all_q_df['ViewCount']).dropna().astype(int).mean(), 1)
print("Avg View", avg_view)
avg_fav = round(pd.to_numeric(all_q_df['FavoriteCount']).dropna().astype(int).mean(), 1)
print("Avg Favorite (droped n/a)", avg_fav)
avg_fav = round(pd.to_numeric(all_q_df['FavoriteCount']).fillna(0).astype(int).mean(), 1)
print("Avg Favorite (filling n/a favorits with zero)", avg_fav)
avg_score = round(pd.to_numeric(all_q_df['Score']).dropna().astype(int).mean(), 1)
print("Avg Score", avg_score)
import statistics
time_taken_list = []
acc_count = 0
for _, el in all_q_df.iterrows():
if int(el['Id']) in set_ques_acc_ids:
acc_count += 1
time = get_acc_time(int(el['Id']))
time_taken_list.append(time)
# print(acc_count)
pct_qwo_acc = round(((len(all_q_df) - acc_count)) * 100.0 / len(all_q_df), 0)
print("% W/o Acct Ans", pct_qwo_acc)
med_hours = statistics.median(time_taken_list)
print("Med Hrs to Acc.", med_hours)
# -
# ### Comparison of Question Types Between Different Domains
rq_df = pd.read_csv('data/RQ_annotation.csv')
ss = '''
{'How': 271, 'What': 87, 'Why': 68, 'Others': 61}
Type: How pct: 55.6
Type: What pct: 17.9
Type: Why pct: 14.0
Type: Others pct: 12.5
'''
print(ss)
| Codes/LCSD_Challenges_EMSE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: gpu
# language: python
# name: gpu
# ---
# # Classification
#
# *Some of of the following materials borrow concepts or code snippets from
# materials written by our friends <NAME>, <NAME> and Quentin
# Batista. We thank them for their willingness to allow us to build on their
# ideas*
#
# **Prerequisites**
#
# - [Regression](regression.ipynb)
#
#
# **Outcomes**
# ## Introduction to Classification
#
# We now move from regression to the second main branch of machine learning:
# classification
#
# Recall that the regression problem was to construct a mapping from a set of
# feature variable to a continuous target
#
# Classification is similar to regression, but instead of predicting a continuous
# target, classification algorithms attempt to apply one (or more) of a discrete
# number of labels or classes to each observation
#
# Another way to put it is that for regression the targets are usually
# continuous-valued, while in classification the targets are categorical
#
# Common examples of classification problems are
#
# - Labeling emails as spam or not
# - Person identification in a photo
# - Speech recognition
# - Whether or not a country is or will be in a recession
#
#
# Classification can also be applied in settings where the target isn’t naturally
# categorical
#
# For example, suppose we want to predict if the unemployment rate for a state
# will be low ($ <3\% $), medium ($ \in [3\%, 5\%] $), or high ($ >5\% $)
# but don’t necessarily care about the actual number
#
# While most economic problems are posed in continuous terms, it may take a bit
# of creativity to determine the optimal way to categorize a target variable so
# classification algorithms can be applied
#
# Just as many problems can be posed either as classification or regression, many
# machine learning algorithms have variants that perform regression or
# classification tasks
#
# Throughout this lecture we will revisit some of the algorithms from
# [regression](regression.ipynb) lecture and discuss how they can be applied in
# classification settings
#
# As we have seen relatives of many of these algorithms, this lecture will be
# lighter on exposition and build up to an application at the end
# + hide-output=false
import datetime
import numpy as np
import pandas as pd
import seaborn as sns
# import pandas_datareader.data as web
from sklearn import (
linear_model, metrics, neural_network, pipeline, preprocessing, model_selection
)
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ## Warmup Example: Logistic Regression
#
# We have actually already come across a classification algorithm
#
# In the recidivism example we attempted to predict whether
# or not an individual would commit another crime using a combination of the
# assigned COMPAS score and the individual’s gender or race
#
# In that example we used a *logistic regression* model, which is a close
# relative of the linear regression model we saw in the [regression](regression.ipynb) section
#
# The logistic regression model for predicting the likelihood of recidivism using
# the `COMPAS` score as the single feature is written
#
# $$
# p(\text{recid}) = L(\beta_0 + \beta_1 \text{COMPAS} + \epsilon)
# $$
#
# where $ L $ is the *logistic function*: $ L(x) = \frac{1}{1 + e^{-x}} $
#
# To get some intuitions for this function, let’s plot it below
# + hide-output=false
x = np.linspace(-5, 5, 100)
y = 1/(1+np.exp(-x))
plt.plot(x, y)
# -
# Notice that for all values of $ x $ the value of the logistic function is
# always between 0 and 1
#
# This makes it perfectly suited for binary classification problems that need to
# output a probability of one of the two labels
#
# Let’s load up the recidivism data and fit the Logistic regression model
df.head()
# + hide-output=false
data_url = "https://raw.githubusercontent.com/propublica/compas-analysis"
data_url += "/master/compas-scores-two-years.csv"
df = pd.read_csv(data_url)
df.head()
X = df[["decile_score"]]
y = df["two_year_recid"]
X_train, X_test, y_train, y_test = model_selection.train_test_split(X, y, test_size=0.25)
logistic_model = linear_model.LogisticRegression(solver="lbfgs")
logistic_model.fit(X_train, y_train)
beta_0 = logistic_model.intercept_[0]
beta_1 = logistic_model.coef_[0][0]
print(f"Fit model: p(recid) = L({beta_0:.4f} + {beta_1:.4f} decile_score)")
# -
# We see from these coefficients that an increase in the `decile_score` leads
# to an increase in the predicted probability of recidivism
#
# Suppose we choose to classify any model output greater than 0.5 as at risk of
# recidivism
#
# Then, because of the positive coefficient on `decile_score` there is a
# threshold level of the COMPAS score above which all individuals will be labeled
# as high-risk
#
# <blockquote>
#
# **Check for understanding**
#
# Determine what the level of this cutoff value is. Recall that the COMPAS
# score takes on integer values between 1 and 10, inclusive.
#
# What happens to the cutoff level of the `decile_score` when you change
# the classification threshold from 0.5 to 0.7? What about 0.3? Remember this
# idea – we’ll come back to it soon
#
#
# </blockquote>
# ### Visualization: Decision Boundaries
#
# With just one feature that has a positive coefficient, the predictions of the
# model will always have this cutoff structure
#
# Let’s add a second feature the model, the age of the individual
# + hide-output=false
X = df[["decile_score", "age"]]
X_train, X_test, y_train, y_test = model_selection.train_test_split(
X, y, test_size=0.25, random_state=42
)
logistic_age_model = linear_model.LogisticRegression()
logistic_age_model.fit(X_train, y_train)
beta_0 = logistic_age_model.intercept_[0]
beta_1, beta_2 = logistic_age_model.coef_[0]
print(f"Fit model: p(recid) = L({beta_0:.4f} + {beta_1:.4f} decile_score + {beta_2:.4f} age)")
# -
# Here we that an increase in the `decile_score` still leads to an increase in
# the predicted probability of recidivism, while older individuals are slightly
# less likely to commit crime again
#
# We’ll build on an example from the [scikit-learn documentation](https://scikit-learn.org/stable/auto_examples/svm/plot_iris.html) to visualize the predictions of this model
# + hide-output=false
def plot_contours(ax, mod, xx, yy, **params):
"""
Plot the decision boundaries for a classifier with 2 features x and y.
Parameters
----------
ax: matplotlib axes object
mod: a classifier
xx: meshgrid ndarray
yy: meshgrid ndarray
params: dictionary of params to pass to contourf, optional
"""
Z = mod.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
out = ax.contourf(xx, yy, Z, **params)
return out
def fit_and_plot_decision_boundary(mod, X, y, **params):
# fit model
mod.fit(X, y)
# generate grids of first two columns of X
def gen_grid(xseries):
if xseries.nunique() < 50:
return sorted(xseries.unique())
else:
return np.linspace(xseries.min(), xseries.max(), 50)
x1, x2 = np.meshgrid(gen_grid(X.iloc[:, 0]), gen_grid(X.iloc[:, 1]))
# plot contours and scatter
fig, ax = plt.subplots()
plot_contours(ax, mod, x1, x2, **params)
x1_name, x2_name = list(X)[:2]
X.plot.scatter(x=x1_name, y=x2_name, color=y, ax=ax)
ax.set_xlabel(x1_name)
ax.set_ylabel(x2_name)
return ax
fit_and_plot_decision_boundary(
linear_model.LogisticRegression(),
X_train, y_train, cmap=plt.cm.Greys
)
# -
# In this plot we can see clearly the relationships we identified from the
# coefficients
#
# However we do see that the model is not perfect, there are some solid circles
# in the light section and some light circles in the solid section
#
# This is likely due to two things:
#
# 1. The model inside the logistic function is a linear regression – thus only a
# linear combination of the input features can be used for prediction
# 1. There is no way to draw a straight line (linear) that perfectly separates
# the true from false observations
#
#
# <blockquote>
#
# **Check for understanding**
#
# Experiment with different pairs of features to see which ones show the
# clearest decision boundaries
#
# Use the `fit_and_plot_decision_boundary` function above and feed it
# different `X` DataFrames
#
#
# </blockquote>
# ## Model Evaluation
#
# Before we get too far into additional classification algorithms, let’s take a
# step back and think about how to evaluate the performance of a classification
# model
# ### Accuracy
#
# Perhaps the most intuitive classification metric is *accuracy*, which is the
# fraction of correct predictions
#
# For a scikit-learn classifier this can be computed using the `score` method
# + hide-output=false
train_acc = logistic_age_model.score(X_train, y_train)
test_acc = logistic_age_model.score(X_test, y_test)
train_acc, test_acc
# -
# When we see that the testing accuracy is similar to or higher than the training
# accuracy (as we do here) that is a signal that the model might be underfitting
# and we should consider either using a more powerful model or adding additional
# features to the model
#
# In many contexts this would be an appropriate way to evaluate a model, but in
# other contexts this would be in sufficient
#
# For example, suppose we want to use a classification model to predict the
# likelihood of someone having a rare, but serious health condition
#
# If the condition is very rare, say it shows up in 0.01% of the population, then
# a model that always predicts false would have 99.99% accuracy but the false
# negatives would have potentially large consequences
# ### Precision and Recall
#
# In order to capture situations like that, data scientists often uses two other
# very common are two other very common metrics:
#
# - *Precision*: the number of true positives over the number of positive
# predictions. Precision tells us how often the model was correct when it
# predicted true
# - *Recall*: The number of true positives over the number of actual positives.
# Recall answer the question, “what fraction of the positives did we get
# correct?”
#
#
# In the classifying probability of rare health condition example, you may prefer
# a model with high recall (never misses an at risk patient) even if the
# precision is a bit low (sometimes you have false positives)
#
# On the other hand, if your algorithm was trying to filter spam out of an email
# inbox you may prefer a model with high precision so that when something is
# classified as spam it is very likely to actually be spam (i.e. non-spam
# messages don’t get sent to spam folder)
#
# In many settings, both precision and recall are equally important and a
# compound metric, known as the F1-score is used:
#
# $$
# F1 = 2 \frac{\text{precision} \cdot \text{recall}}{\text{precision} + \text{recall}}
# $$
#
# The F1 score is bounded between 0 and 1 and will only achieve a value of 1 if
# both precision and recall are exactly 1
#
# We can have scikit-learn produce a textual report with a precision and
#
# Scikit-learn
# + hide-output=false
report = metrics.classification_report(
y_train, logistic_age_model.predict(X_train),
target_names=["no recid", "recid"]
)
print(report)
# -
# ### ROC and AUC
#
# For classification algorithms there is a tradeoff between precision and recall
#
# Let’s illustrate this point in the context of the logistic regression model
#
# The output of a logistic regression is a probability of an event or label
#
# In order to get a definite prediction from the algorithm, the modeler would
# first select a threshold parameter $ p $ such that all model outputs above the
# threshold are given the label of true
#
# As this $ p $ increases the model must be relatively more confident before
# assigning a label of true
#
# In this case the model’s precision will increase (very confident when applying
# true label), but the recall will suffer (will apply false to some true cases
# that had a model output just below the raised threshold)
#
# Machine learning practitioners have adapted a way to help us visualize
# this tradeoff
#
# The visualization technique is known as the receiver operating characteristic
# – or more commonly used ROC – curve <sup>[1](#roc)</sup>
#
# To understand what this curve is, consider two extremes choices for $ p $:
#
# - When $ p=1 $ we will (almost surely) never predict any observation to
# have a label 1. In this case the false positive rate will be equal to 0 as
# will the true positive rate
# - When $ p=0 $ we will predict that all observations always have a label
# of 1. The false positive rate and true positive rates will be equal to 1
#
#
# The *ROC curve* traces out the relationship between the false positive rate (on
# the x axis) and the true positive rate (on the y axis) as the probability
# threshold \$p\$ is changed
#
# Below we define a function that uses scikit-learn to compute the true positive
# rate and false positive rates for us and then plot these rates against
# each-other
#
# <a id='roc'></a>
# **[1]** The name receiver operating characteristic comes from the
# context where ROC curves were first used. ROC curves were
# first used to describe radar receivers during World War II,
# where the goal was to classify a noisy radar signal as
# indicating an aircraft or not.
# + hide-output=false
def plot_roc(mod, X, y):
# predicted_probs is an N x 2 array, where N is number of observations
# and 2 is number of classes
predicted_probs = mod.predict_proba(X_test)
# keep the second column, for label=1
predicted_prob1 = predicted_probs[:, 1]
fpr, tpr, _ = metrics.roc_curve(y_test, predicted_prob1)
# Plot ROC curve
fig, ax = plt.subplots()
ax.plot([0, 1], [0, 1], "k--")
ax.plot(fpr, tpr)
ax.set_xlabel("False Positive Rate")
ax.set_ylabel("True Positive Rate")
ax.set_title("ROC Curve")
plot_roc(logistic_age_model, X_test, y_test)
# -
# We can use the ROC curve to determine the optimal value for the threshold
#
# Given that the output of our model for the recidivism application could
# potentially inform judicial decisions that impact the lives of individuals, we
# should be careful when considering a threshold value with low false
# positive rate vs high recall (low false negative rate)
#
# We may choose to err on the side of low false negative rate so that when the model
# predicts recidivism it is likely that recidivism will occur – in other words
# we would favor a high true positive rate even if the false positive rate is
# higher
#
# <blockquote>
#
# **Check for understanding**
#
# Use the `metrics.roc_curve` function to determine an appropriate value
# for the probability threshold, while keeping in mind our preference for
# high precision over high recall
#
# The third return value of `metrics.roc_curve` is an array of the
# probability thresholds (`p`) used to compute each false positive rate and
# true positive rate
#
# To do this problem, you may wish to do the following steps:
#
# - Come up with some objective function in terms of the `fpr` and `tpr`
# - Evaluate the objective function using the `fpr` and `tpr` variables returned by the `metrics.roc_curve` function
# - Use `np.argmin` to find the *index* of the smallest value of the objective function
# - Extract from the probability threshold values array the value at the argmin index
#
#
# *Hint*: If we cared about both precision and recall equally (we don’t here)
# we might choose as on objective function `(fpr - tpr)**2`. With this
# objective function we will find the value of the probability threshold
# that makes the false positive and true positive rates as close to equal as
# possible
#
#
# </blockquote>
# + hide-output=false
# your code here
# -
# <blockquote>
#
# </blockquote>
# The ROC curve can also be used to do hyper-parameter selection for the model’s
# parameters
#
# To see how, consider a model with an ROC curve that has a single point at (0, 1)
# – meaning the true positive rate is 1 and false positive rate is zero or
# that the model has 100% accuracy
#
# Notice that if we were to integrate to obtain the area under the ROC curve, we
# would get a value of 1 for the perfect model
#
# The area under any other ROC curve would be less than 1
#
# Thus, we could use the area under the curve (abbreviated AUC) as an objective
# metric in cross-validation
#
# Let’s see an example
# + hide-output=false
predicted_prob1 = logistic_age_model.predict_proba(X)[:, 1]
auc = metrics.roc_auc_score(y, predicted_prob1)
print(f"Initial AUC value is {auc:.4f}")
help(linear_model.LogisticRegression)
# -
# <blockquote>
#
# **Check for understanding**
#
# The `LogisticRegression` class with default arguments implements the
# regression including `l2` regularization (it penalizes coefficient
# vectors with an l2-norm)
#
# The strength of regularization is controlled by a parameter `C` that is
# passed to the `LogisticRegression` constructor
#
# Smaller values of `C` lead to stronger regularization
#
# For example `LogisticRegression(C=10)` would have weaker regularization
# than `LogisticRegression(C=0.5)`
#
# Your task here is to use the `model_selection.cross_val_score` method,
# with the `scoring` argument set to `roc_auc` in order to select an
# optimal level for the regularization parameter `C`
#
# Refer to the example in the recidivism lecture for how
# to use `model_selection.cross_val_score`
#
#
# </blockquote>
# + hide-output=false
# your code here
# -
# <blockquote>
#
# </blockquote>
# ## Neural Network Classifiers
#
# The final classifier we will visit today is a neural-network classifier, using
# the multi-layer perceptron network architecture
#
# Recall from the [regression](regression.ipynb) chapter that a multi-layer
# perceptron is made up of a series of nested linear regressions, separated by
# non-linear activation functions
#
# The number of neurons (size of weight matrices and bias vectors) in each layer
# are hyperparameters that can be chosen by modeler, but for regression the last
# layer had to have exactly one neuron which represented the single regression
# target
#
# In order to use the MLP for classification tasks we need to make three adjustments:
#
# 1. Construct a final layer with $ N $ neurons instead of 1, where $ N $ is the number of classes in the classification task
# 1. Apply a *softmax* function on the output of the network
# 1. Use the cross-entropy loss function instead of the MSE to optimize network weights and biases
#
#
# The softmax function applied to a vector $ x \in \mathbb{R}^N $ is computed as
#
# $$
# \sigma(x)_i = \frac{e^{x_i}}{\sum_{j=1}^{N} e^{x_j}}
# $$
#
# In words, the softmax function is computed by exponentiating all the values,
# then dividing by the sum of exponentiated values
#
# The output of the softmax function is a probability distribution (all
# non-negative and sum to 1), weighted by the relative value of the input values
#
# Finally, the cross entropy loss function for $ M $ observations $ y $, with associated softmax vectors $ z $ is
#
# $$
# -\frac{1}{M} \sum_{j=1}^M \sum_{i=1}^N 1_{y_j = i} log\left(z_{i,j}\right)
# $$
#
# where $ 1_{y_j = i} $ is an indicator variable taking on the value of 1 if
# the observed class was equal to $ i $ for the :math:`j`th observation, 0
# otherwise
#
# All the same tradeoffs we saw when we used the multi-layer perceptron for
# regression will apply for classification tasks
#
# This includes positives like automated-feature enginnering and theoretically unlimited flexibility
#
# It also includes potential negatives such as a risk of overfitting, high
# computational expenses compared to many classification algorithms, lack of
# interpretability
#
# For a more detailed discussion, review the [regression lecture](regression.ipynb)
#
# <blockquote>
#
# **Check for understanding**
#
# Use the `neural_network.MLPClassifier` class to use a multi-layer
# perceptron in our recidivism example
#
# Experiment with different inputs such as:
#
# - The features to include
# - The number of layers and number of neurons in each layer
# - The l2 regularization parameter `alpha`
# - The solver
#
#
# See if you can come up with a model that outperforms logistic regression
#
# Keep in mind other things like the degree of overfitting and time required
# to estimate the model parameters. How do these compare to logistic
# regression?
#
#
# </blockquote>
# + hide-output=false
# your code here
# -
# <blockquote>
#
# </blockquote>
# ### Aside: neural network toolboxes
#
# Thus far we have been using the routines in scikit-learn’s `neural_network` package
#
# These are great for learning and exploratory analysis like we have been doing,
# but are rarely used in production or real world settings
#
# The two main reasons are that the scikit-learn routines do not leverage modern
# hardware like GPUs (so performance is likely much slower than it could be) and
# it only provides an implementation of the most basic of deep neural networks
#
# If you were to use neural networks in mission-critical situations you would
# want to use modern neural network libraries such as Google’s [tensorflow](https://www.tensorflow.org/),
# Facebook’s [pytorch](https://pytorch.org/), the amazon supported [MXNet](https://mxnet.apache.org/), or
# [fastai](https://www.fast.ai/)
#
# Each of these toolkits has its own relative strengths and weaknesses, but we’ve
# seen tensorflow and pytorch used the most
#
# Thankfully, they all support Python as either the only or the primary point of
# access, so you will be well-prepared to start using them
# ## Application: predicting US recessions
#
# Let’s apply our new classification algorithm knowledge and try to use
# [leading indicators](https://www.investopedia.com/terms/l/leadingindicator.asp)
# to predict recessions in the US economy
#
# A leading indicator is a variable that tends to move or change before the rest
# of the economy
#
# Many different leading indicators have been proposed – we’ll use a few of them
#
# We won’t go into detail defending the choice of these variables as leading
# indicators, but will show a plot of each of the variables that will let us
# visually inspect the hypothesis
# ### Data prep
#
# Let’s first gather the data from FRED
# + hide-output=false
start = "1974-01-01"
end = datetime.date.today()
def pct_change_on_last_year(df):
"compute pct_change on previous year, assuming quarterly"
return (df - df.shift(4))/df.shift(4)
def get_indicators_from_fred(start=start, end=end):
"""
Fetch quarterly data on 6 leading indicators from time period start:end
"""
# yield curve, unemployment, change in inventory, new private housing permits
yc_unemp_inv_permit = (
web.DataReader(["T10Y2Y", "UNRATE", "CBIC1", "PERMIT"], "fred", start, end)
.resample("QS")
.mean()
)
# percent change in housing prices and retail sales
hpi_retail = (
web.DataReader(["USSTHPI", "SLRTTO01USQ661S"], "fred", start, end)
.resample("QS") # already quarterly, adjusting so index is same
.mean()
.pipe(pct_change_on_last_year)
.dropna()
)
indicators = (
yc_unemp_inv_permit
.join(hpi_retail)
.dropna()
.rename(columns=dict(
USSTHPI="pct_change_hpi",
T10Y2Y="yield_curve",
UNRATE="unemp",
CBIC1="inventory",
SLRTTO01USQ661S="retail_sales",
PERMIT="house_permits"
))
)
return indicators
indicators = get_indicators_from_fred()
indicators.head()
# -
# Now we also need data on recessions
# + hide-output=false
def get_recession_data():
recession = (
web.DataReader(["USRECQ"], "fred", start, end)
.rename(columns=dict(USRECQ="recession"))
["recession"]
)
# extract start and end date for each recession
start_dates = recession.loc[recession.diff() > 0].index.tolist()
if recession.iloc[0] > 0:
start_dates = [recession.index[0]] + start_dates
end_dates = recession.loc[recession.diff() < 0].index.tolist()
if len(start_dates) != len(end_dates):
raise ValueError("Need to have same number of start/end dates!")
return recession, start_dates, end_dates
recession, start_dates, end_dates = get_recession_data()
# -
# Now let’s take a look at the data we have:
# + hide-output=false
def add_recession_bands(ax):
for s, e in zip(start_dates, end_dates):
ax.axvspan(s, e, color="grey", alpha=0.2)
axs = indicators.plot(subplots=True, figsize=(8, 6), layout=(3, 2), legend=False)
for i, ax in enumerate(axs.flatten()):
add_recession_bands(ax)
ax.set_title(list(indicators)[i])
fig = axs[0, 0].get_figure()
fig.tight_layout();
# -
# For each of the variables we have chosen you can see that in periods leading up
# to a recession (noted by the grey bands in background), the leading indicator
# has a distinct move
#
# <blockquote>
#
# **Check for understanding**
#
# Let’s pause here to take a few minutes and digest
#
# If the task is to use these leading indicators to predict a recession,
# would high recall or high precision be more important for our model?
#
# Would your answer change if you worked at the federal reserve?
#
# What if you worked at a news company such as the Economist or the New York
# Times?
#
#
# </blockquote>
# ### How many leads?
#
# If the variables we have chosen truly are leading indicators, we should be able
# to use leading values of the variables to predict current or future recessions
#
# A natural question is how many leads should we include?
#
# Let’s explore that question by looking at many different sets of leads
# + hide-output=false
def make_train_data(indicators, rec, nlead=4):
return indicators.join(rec.shift(nlead)).dropna()
def fit_for_nlead(ind, rec, nlead, mod):
df = make_train_data(ind, rec, nlead)
X = df.drop(["recession"], axis=1).copy()
y = df["recession"].copy()
X_train, X_test, y_train, y_test = model_selection.train_test_split(X, y)
mod.fit(X_train, y_train)
cmat = metrics.confusion_matrix(y_test, mod.predict(X_test))
return cmat
mod = pipeline.make_pipeline(
preprocessing.StandardScaler(),
linear_model.LogisticRegression(solver="lbfgs")
)
cmats = dict()
for nlead in range(1, 11):
cmats[nlead] = np.zeros((2, 2))
print(f"starting for {nlead} leads")
for rep in range(200):
cmats[nlead] += fit_for_nlead(indicators, recession, nlead, mod)
cmats[nlead] = cmats[nlead] / 200
for k, v in cmats.items():
print(f"\n\nThe average confusion matrix for {k} lag(s) was:\n {v}")
# -
# From the averaged confusion matrices reported above we see that the model with
# only one leading period was the most accurate
#
# After that was the model with 4 leading quarters
#
# Depending on the application, we might favor a model with higher accuracy or
# one that gives us more time to prepare (the 4 quarter model)
#
# One reason why the 1 lead and 4 lead models did better than the models with
# another number of leads might be because different variables start to move a
# different number of periods before the recession hits
#
# The exercise below asks you to explore this idea
#
# <blockquote>
#
# **Check for understanding**
#
# Extend the logic from the previous example and allow for a different number
# of leading periods for each variable
#
# How would you find the “optimal” number of leads for each variable? How
# could you try to avoid overfitting?
#
# Use `make_train_data_varying_leads` function below to construct
#
#
# </blockquote>
# + hide-output=false
def make_train_data_varying_leads(indicators, rec, nlead):
"""
Apply per-indicator leads to each indicator and join with recession data
Parameters
----------
indicators: pd.DataFrame
A DataFrame with timestamps on index and leading indicators as columns
rec: pd.Series
A Series indicating if the US economy was in a recession each period
nlead: dict
A dictionary mapping a column name to a positive integer
specifying how many periods to shift each indicator. Any
indicator not given a key in this dictionary will not be
included in the output DataFrame
Returns
-------
df: pd.DataFrame
A DataFrame with the leads applied and merged with the recession
indicator
Example
-------
```
df = make_train_data_varying_leads(
indicators,
recession,
nlead=dict(yield_curve=3, unemp=4)
)
df.shape[1] # == 3 (yield_curve, unemp, recession))
```
"""
cols = []
for col in list(indicators):
if col in nlead:
cols.append(indicators[col].shift(-nlead[col]))
X = pd.concat(cols, axis=1)
return X.join(rec).dropna()
# your code here!
# -
# <blockquote>
#
# </blockquote>
# <blockquote>
#
# **Check for understanding**
#
# Experiment with different classifiers. Which ones perform better or worse?
#
# See if you can beat your neighbor at accuracy, precision, and/or recall
#
#
# </blockquote>
| 2019-06-03__Orientation/classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Massive Comparison of different cluster algorithms
# (adpated from sklearn)
# (c) <NAME>
import time
import warnings
import numpy as np
import matplotlib.pyplot as plt
from sklearn import cluster, datasets, mixture
from sklearn.neighbors import kneighbors_graph
from sklearn.preprocessing import StandardScaler
from itertools import cycle, islice
# Global rnd num seed: change here for quick and dirty
np.random.seed(0)
# Number of samples
n_samples = 150
# Generate sample point clouds
noisy_circles = datasets.make_circles(n_samples=n_samples, factor=.5, noise=.05)
noisy_moons = datasets.make_moons(n_samples=n_samples, noise=.05)
blobs = datasets.make_blobs(n_samples=n_samples, random_state=8)
# Anisotropicly distributed data
random_state = 170
X, y = datasets.make_blobs(n_samples=n_samples, random_state=random_state)
transformation = [[0.6, -0.6], [-0.4, 0.8]]
X_aniso = np.dot(X, transformation)
aniso = (X_aniso, y)
# blobs with varied variances
varied = datasets.make_blobs(n_samples=n_samples,
cluster_std=[1.0, 2.5, 0.5],
random_state=random_state)
# ============
# Set up cluster parameters
# ============
plt.figure(figsize=(9 * 2 + 3, 12.5))
plt.subplots_adjust(left=.02, right=.98, bottom=.001, top=.96, wspace=.05,
hspace=.01)
# Default paramters (used for blobs and no_structure later)
default_base = {'quantile': .3,
'eps': .3,
'damping': .9,
'preference': -200,
'n_neighbors': 10,
'n_clusters': 3,
'min_samples': 20,
'xi': 0.05,
'min_cluster_size': 0.1}
# Actual default parameters
datasets = [
(noisy_circles, {'damping': .77, 'preference': -240,
'quantile': .2, 'n_clusters': 2,
'min_samples': 20, 'xi': 0.25}),
(noisy_moons, {'damping': .75, 'preference': -220, 'n_clusters': 2}),
(varied, {'eps': .18, 'n_neighbors': 2,
'min_samples': 5, 'xi': 0.035, 'min_cluster_size': .2}),
(aniso, {'eps': .15, 'n_neighbors': 2,
'min_samples': 20, 'xi': 0.1, 'min_cluster_size': .2}),
(blobs, {})
]
# Subplot index
plot_num = 1
# Compute and plot everything
for i_dataset, (dataset, algo_params) in enumerate(datasets):
# Set parameters with dataset-specific values
params = default_base.copy()
params.update(algo_params)
# Set dataset (hidden index i_dataset)
X, y = dataset
# Normalize and fit
X = StandardScaler().fit_transform(X)
# BANDWITH method
# Estimate bandwidth for mean shift
bandwidth = cluster.estimate_bandwidth(X, quantile=params['quantile'])
# WARD method
# Connectivity matrix for structured Ward
connectivity = kneighbors_graph(
X, n_neighbors=params['n_neighbors'], include_self=False)
# make connectivity symmetric
connectivity = 0.5 * (connectivity + connectivity.T)
# ============
# Create cluster objects
# ============
#Mean Shift method
ms = cluster.MeanShift(bandwidth=bandwidth, bin_seeding=True)
#Mini Batch K-Means
two_means = cluster.MiniBatchKMeans(n_clusters=params['n_clusters'])
#Ward method (Hierachical clustering)
ward = cluster.AgglomerativeClustering(
n_clusters=params['n_clusters'], linkage='ward',
connectivity=connectivity)
#Spectal Clustering method
spectral = cluster.SpectralClustering(
n_clusters=params['n_clusters'], eigen_solver='arpack',
affinity="nearest_neighbors")
# DBSCAN
dbscan = cluster.DBSCAN(eps=params['eps'])
#Affinity propagation method
affinity_propagation = cluster.AffinityPropagation(
damping=params['damping'], preference=params['preference'])
#Average Linkage method (Agglomerative Clustering)
average_linkage = cluster.AgglomerativeClustering(
linkage="average", affinity="cityblock",
n_clusters=params['n_clusters'], connectivity=connectivity)
#Birch method (balanced iterative reducing and clustering using hierarchies)
#(Hierarchical clustering)
birch = cluster.Birch(n_clusters=params['n_clusters'])
#Gaussian Mixure (EM method with finite number of gaussian centers)
gmm = mixture.GaussianMixture(
n_components=params['n_clusters'], covariance_type='full')
#List of tupels with method name and pointer to it
clustering_algorithms = (
('MiniBatchKMeans', two_means),
('Affinity', affinity_propagation),
('MeanShift', ms),
('SpectralC', spectral),
('Ward', ward),
('AggloC', average_linkage),
('DBSCAN', dbscan),
('Birch', birch),
('GaussMixtureEM', gmm)
)
# Algorithm call loop
for name, algorithm in clustering_algorithms:
# For runtime calculation
t0 = time.time()
#XXX Catch warnings using WITCH, related to kneighbors_graph
#main thing: call algorithm.fit(X)
with warnings.catch_warnings():
warnings.filterwarnings(
"ignore",
message="the number of connected components of the " +
"connectivity matrix is [0-9]{1,2}" +
" > 1. Completing it to avoid stopping the tree early.",
category=UserWarning)
warnings.filterwarnings(
"ignore",
message="Graph is not fully connected, spectral embedding" +
" may not work as expected.",
category=UserWarning)
algorithm.fit(X)
# Runtime is t1-t0 now
t1 = time.time()
#XX
if hasattr(algorithm, 'labels_'):
y_pred = algorithm.labels_.astype(np.int)
else:
y_pred = algorithm.predict(X)
# Plot Lables and Plot Title
plt.subplot(len(datasets), len(clustering_algorithms), plot_num)
if i_dataset == 0:
plt.title(name, size=18)
# Define colors (dynamic, cyclic, as #predicted classes)
colors = np.array(list(islice(cycle(['r', 'g', 'b',
'#ffff00', '#00ffff', '#ff00ff',
'#999999', '#e41a1c', '#dede00']),
int(max(y_pred) + 1))))
# Add black color for outliers (if any)
colors = np.append(colors, ["#000000"])
plt.scatter(X[:, 0], X[:, 1], s=10, color=colors[y_pred],
alpha=0.3, marker='o', edgecolor='black')
plt.xlim(-2.5, 2.5)
plt.ylim(-2.5, 2.5)
plt.xticks(())
plt.yticks(())
#display runtime for output
plt.text(.99, .01, ('%.2fs' % (t1 - t0)).lstrip('0'),
transform=plt.gca().transAxes, size=15,
horizontalalignment='right')
#increase subplot index
plot_num += 1
# Conversion to high-resolution pdf (call this before plt.show!)
plt.savefig('aaaa_clustering_monster.pdf')
# Show entire plot figure AND save now
plt.show()
# -
| Codes/ClusterMonster.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
import cv2
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
# from subprocess import call
# call(["python", "C:\\Users\\z5130037\\Desktop\\Udacity\\Camera_calibaration\\CarND-Camera-Calibration-master\\CarND-Camera-Calibration-master\\camera_calibration.ipynb"])
# Read in the saved camera matrix and distortion coefficients
# These are the arrays you calculated using cv2.calibrateCamera()
# and this the adress of cv2.calibrateCamera()::\Users\z5130037\Desktop\Udacity\Camera_calibaration\CarND-Camera-Calibration-master\CarND-Camera-Calibration-master\calibration_wide
dist_pickle = pickle.load( open( "calibration_wide\wide_dist_pickle.p", "rb" ) )
#dist_pickle = pickle.load( open( "wide_dist_pickle.p", "rb" ) )
mtx = dist_pickle["mtx"]
dist = dist_pickle["dist"]
print(mtx)
print(dist)
# +
# Read in an image
#img = cv2.imread('test_image2.png')
#img = cv2.imread('calibration_wide\test_image.jpg') #this test image has some issue.
img = cv2.imread('calibration_wide\GOPR0035.jpg')
#img = cv2.imread('calibration_wide\test_image.jpg')
nx = 8 # the number of inside corners in x
ny = 6 # the number of inside corners in y
# +
#undist = cv2.undistort(img, mtx, dist, None, mtx)
#go through this link : https://adventuresandwhathaveyou.wordpress.com/category/coding-2/
# -
# MODIFY THIS FUNCTION TO GENERATE OUTPUT
# THAT LOOKS LIKE THE IMAGE ABOVE
def corners_unwarp(img, nx, ny, mtx, dist):
# Pass in your image into this function
# Write code to do the following steps
# 1) Undistort using mtx and dist
# Use the OpenCV undistort() function to remove distortion
undist = cv2.undistort(img, mtx, dist, None, mtx)
# 2) Convert to grayscale
# Convert undistorted image to grayscale
gray = cv2.cvtColor(undist, cv2.COLOR_BGR2GRAY)
# 3) Find the chessboard corners
# Search for corners in the grayscaled image
ret, corners = cv2.findChessboardCorners(gray, (nx, ny), None)
# 4) If corners found:
if ret == True:
# a) draw corners
# If we found corners, draw them! (just for fun)
cv2.drawChessboardCorners(undist, (nx, ny), corners, ret)
# Choose offset from image corners to plot detected corners
# This should be chosen to present the result at the proper aspect ratio
# My choice of 100 pixels is not exact, but close enough for our purpose here
offset = 100 # offset for dst points
# Grab the image shape
img_size = (gray.shape[1], gray.shape[0])
# b) define 4 source points src = np.float32([[,],[,],[,],[,]])
# For source points I'm grabbing the outer four detected corners
src = np.float32([corners[0], corners[nx-1], corners[-1], corners[-nx]])
#Note: you could pick any four of the detected corners
# as long as those four corners define a rectangle
#One especially smart way to do this would be to use four well-chosen
# corners that were automatically detected during the undistortion steps
#We recommend using the automatic detection of corners in your code
# c) define 4 destination points dst = np.float32([[,],[,],[,],[,]])
# For destination points, I'm arbitrarily choosing some points to be
# a nice fit for displaying our warped result
# again, not exact, but close enough for our purposes
dst = np.float32([[offset, offset], [img_size[0]-offset, offset],
[img_size[0]-offset, img_size[1]-offset],
[offset, img_size[1]-offset]])
# d) use cv2.getPerspectiveTransform() to get M, the transform matrix
# Given src and dst points, calculate the perspective transform matrix
M = cv2.getPerspectiveTransform(src, dst)
# e) use cv2.warpPerspective() to warp your image to a top-down view
# Warp the image using OpenCV warpPerspective()
warped = cv2.warpPerspective(undist, M, img_size)
#delete the next two lines
return warped, M
top_down, perspective_M = corners_unwarp(img, nx, ny, mtx, dist)
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(24, 9))
f.tight_layout()
ax1.imshow(img)
ax1.set_title('Original Image', fontsize=50)
ax2.imshow(top_down)
ax2.set_title('Undistorted and Warped Image', fontsize=50)
plt.subplots_adjust(left=0., right=1, top=0.9, bottom=0.)
| Undistort_and_Tranform.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/flower-go/DiplomaThesis/blob/master/Taggin_and_Lemmatization_Working_Example.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="L4hZQFhWKAAX"
# This is a Jupyter Notebook presenting the usage of pretrained tagging and lemmatization model presented as a part of my thesis. For running please run all cells in Helper code part (PLEASE RESTART RUNTIME after installing requirements). Then run the rest of cells. You can change the input in [this cell](https://colab.research.google.com/drive/14U0zNpnU9VUkvMH2GwzdcULV6Xp_qjhR#scrollTo=Ej_NzOSnNgk6&line=1&uniqifier=1). An output will be printed in the last cell. For more information, do not hesitate to contact me on: <EMAIL>
# + [markdown] id="A15adFmDBEaH"
# # Helper code
#
# + id="4s8Ke81RP9Vv" colab={"base_uri": "https://localhost:8080/"} outputId="ad22eb58-9ec5-459e-c86c-9a068d910f50"
#clone repo
# !git clone https://github.com/flower-go/DiplomaThesis.git
# + colab={"base_uri": "https://localhost:8080/"} id="ZmNEcwiXX42-" outputId="485ceb56-f819-4dde-cd2a-cd1618d920b5"
# !pip install ufal.morphodita
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="E4O9tV0UBesv" outputId="525f4c8b-a6a4-41e7-9498-b874bd58f055"
# !pip install -r "DiplomaThesis/requirements.txt"
# + colab={"base_uri": "https://localhost:8080/"} id="vHj1DmVIz-ke" outputId="d2814ed8-e13f-4c56-beaf-e08b9efc57ee"
# !mkdir /content/DiplomaThesis/code/morphodita-research/models
# !mkdir /content/DiplomaThesis/code/morphodita-research/models/tl_18
# !wget --no-check-certificate aic.ufal.mff.cuni.cz/~doubrap1/mappings.pickle
# !cp /content/mappings.pickle /content/DiplomaThesis/code/morphodita-research/models/tl_18/mappings.pickle
# !rm /content/mappings.pickle
# + colab={"base_uri": "https://localhost:8080/"} id="I9Ab2KZuK356" outputId="763adcaf-0b65-4457-a9e8-d565c4dd2814"
# !wget --no-check-certificate aic.ufal.mff.cuni.cz/~doubrap1/ch18.index
# !cp /content/ch18.index /content/DiplomaThesis/code/morphodita-research/models/ch18.index
# !rm /content/ch18.index
# !wget --no-check-certificate aic.ufal.mff.cuni.cz/~doubrap1/ch18.data-00000-of-00001
# !cp /content/ch18.data-00000-of-00001 /content/DiplomaThesis/code/morphodita-research/models/ch18.data-00000-of-00001
# !rm /content/ch18.data-00000-of-00001
# + colab={"base_uri": "https://localhost:8080/"} id="pa5hwEmfMC_q" outputId="8a8696ff-0d2a-400c-9f6e-5be77fd88940"
# !wget --no-check-certificate aic.ufal.mff.cuni.cz/~doubrap1/forms.vectors-w5-d300-ns5.16b.npz
# + [markdown] id="Wq685N8tZR8y"
# # Creating model and input data
# + id="_loWe77oLMVC"
import sys
sys.path.append("DiplomaThesis/code/morphodita-research")
import tensorflow as tf
import morpho_tagger_2 as mt
# + id="umC6UsT_YUbJ"
import ufal.morphodita
# + id="hLHQFnVt0BvS"
import os
os.chdir("DiplomaThesis/code/morphodita-research")
# + id="MCeEeu1PYaT0"
def tokenize_to_vertical(text):
tokenizer = ufal.morphodita.Tokenizer.newCzechTokenizer()
forms = ufal.morphodita.Forms()
f = open('vstup', 'w')
tokenizer.setText(text)
while tokenizer.nextSentence(forms, None):
for form in forms:
print(form, "_", "_", sep="\t", file=f)
print(file=f)
f.close()
# + [markdown] id="_LuJOilWZcsO"
# -------------------------------------------
# Insert input:
# + id="Ej_NzOSnNgk6"
vstup = "Dnes byl krásný den."
# + [markdown] id="j6g6swEwZYmm"
# ---------------------------------------
# + id="QoH_lGDcPlPm" colab={"base_uri": "https://localhost:8080/", "height": 698, "referenced_widgets": ["3300439a28e34b2e8ba6cd9786e2b7b0", "<KEY>", "e463bcf6fffa457ea0c9f01775b11923", "1538c8d411394847a35364101b24603e", "dd9e541df6034fbc9d8723924c65683f", "1ec923740bf04877bb6decae58cae1bf", "ee7ee701c1134be1ac93ef246cd99d12", "a224bc0a108142c99268cf554459fdd3"]} outputId="7fcebe50-3baa-4e21-deb8-2a8f6f75ad58"
tokenize_to_vertical(vstup)
mt.main(["./vstup", "--checkp", "ch18", "--embeddings", "/content/forms.vectors-w5-d300-ns5.16b.npz","--exp", "tl_18", "--warmup_decay", "c:1", "--label_smoothing", "0.03", "--bert_model", "ufal/robeczech-base", "--epochs", "60:1e-3", "--predict", '/content/DiplomaThesis/code/morphodita-research/models/ch18'])
# + id="uA9j6IwkZ0pH" colab={"base_uri": "https://localhost:8080/"} outputId="1231da70-4de9-406c-e616-8c817b8b7867"
# !cat tl_18_vystup
| Taggin_and_Lemmatization_Working_Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from urllib.request import urlopen, Request
from bs4 import BeautifulSoup
from nltk.sentiment.vader import SentimentIntensityAnalyzer
import pandas as pd
import matplotlib.pyplot as plt
import nltk
nltk.downloader.download('vader_lexicon')
# +
finviz_url = 'https://finviz.com/quote.ashx?t='
tickers = ['AMZN']
news_tables = {}
for ticker in tickers:
url = finviz_url + ticker
req = Request(url=url, headers={'user-agent': 'my-app'})
response = urlopen(req)
html = BeautifulSoup(response, features='html.parser')
news_table = html.find(id='news-table')
news_tables[ticker] = news_table
parsed_data = []
for ticker, news_table in news_tables.items():
for row in news_table.findAll('tr'):
title = row.a.text
date_data = row.td.text.split(' ')
if len(date_data) == 1:
time = date_data[0]
else:
date = date_data[0]
time = date_data[1]
parsed_data.append([ticker, date, time, title])
df = pd.DataFrame(parsed_data, columns=['ticker', 'date', 'time', 'title'])
df['date'] = pd.to_datetime(df.date).dt.date
vader = SentimentIntensityAnalyzer()
f = lambda title: vader.polarity_scores(title)['compound']
df['compound'] = df['title'].apply(f)
plt.figure(figsize=(800,680))
mean_df = df.groupby(['ticker', 'date']).mean().unstack()
mean_df = mean_df.xs('compound', axis="columns").transpose()
mean_df.plot(kind='bar')
plt.show()
# -
| Sentiment_analysis_tesla.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import pandas as pd
import sklearn
from ailib.ml.datasets.binary_classification_toy_problems import gen_2d_samples
from sklearn import datasets
# + [markdown] tags=["meta", "draft"]
# # Keras Toy 2D binary classification
# -
# ## Install Keras
#
# https://keras.io/#installation
#
# ### Install dependencies
#
# Install TensorFlow backend: https://www.tensorflow.org/install/
#
# ```
# pip install tensorflow
# ```
#
# Insall h5py (required if you plan on saving Keras models to disk): http://docs.h5py.org/en/latest/build.html#wheels
#
# ```
# pip install h5py
# ```
#
# Install pydot (used by visualization utilities to plot model graphs): https://github.com/pydot/pydot#installation
#
# ```
# pip install pydot
# ```
#
# ### Install Keras
#
# ```
# pip install keras
# ```
# ## Import packages and check versions
import tensorflow as tf
tf.__version__
import keras
keras.__version__
import h5py
h5py.__version__
import pydot
pydot.__version__
# ## Make the dataset
# +
df_train = gen_2d_samples(n_samples=200)
x_train = df_train[['x1', 'x2']].values
y_train = df_train.y.values
ax = df_train.loc[df_train.y == 0].plot.scatter(x='x1', y='x2', color="r")
df_train.loc[df_train.y == 1].plot.scatter(x='x1', y='x2', ax=ax);
# +
df_test = gen_2d_samples(n_samples=200)
x_test = df_test[['x1', 'x2']].values
y_test = df_test.y.values
ax = df_test.loc[df_test.y == 0].plot.scatter(x='x1', y='x2', color="r")
df_test.loc[df_test.y == 1].plot.scatter(x='x1', y='x2', ax=ax);
# -
# ## Make the classifier
# +
model = keras.models.Sequential()
model.add(keras.layers.Dense(units=2, activation='relu', input_dim=2))
model.add(keras.layers.Dense(units=1, activation='sigmoid'))
model.compile(loss='binary_crossentropy',
optimizer='rmsprop',
metrics=['accuracy'])
model.summary()
# -
print(model.get_config())
hist = model.fit(x_train, y_train, batch_size=100, epochs=10, verbose=False)
plt.plot(hist.history['loss']);
model.evaluate(x_test, y_test)
y_predicted = model.predict(x_test)
# +
predicted_data = np.concatenate([x_test, y_predicted], axis=1)
predicted_df = pd.DataFrame(predicted_data, columns=['x1', 'x2', 'y'])
ax = predicted_df.loc[predicted_df.y <= 0.5].plot.scatter(x='x1', y='x2', color="r")
predicted_df.loc[predicted_df.y > 0.5].plot.scatter(x='x1', y='x2', ax=ax);
# -
# ## Bonnus: plot the regressor
# +
from keras.utils import plot_model
plot_model(model, show_shapes=True, to_file="model.png")
| nb_dev_python/python_keras_toy_2d_binary_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Copyright 2021 NVIDIA Corporation. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# <img src="http://developer.download.nvidia.com/compute/machine-learning/frameworks/nvidia_logo.png" style="width: 90px; float: center;">
# # Triton server with FIL backend
# ## Overview
#
# This notebook shows the procedure to deploy a [XGBoost model](https://xgboost.readthedocs.io/en/latest/) in Triton Inference Server with Forest Inference Library (FIL) backend. The FIL backend allows forest models trained by several popular machine learning frameworks (including XGBoost, LightGBM, Scikit-Learn, and cuML) to be deployed in a Triton inference server using the RAPIDS Forest Inference LIbrary for fast GPU-based inference. Using this backend, forest models can be deployed seamlessly alongside deep learning models for fast, unified inference pipelines.
#
# ### Contents
# * [Train XGBoost model on dummy data](http://localhost:7001/notebooks/simple_xgboost_example.ipynb#Train-XGBoost-model)
# * [Export, load and deploy XGBoost model in Triton Inference Server](http://localhost:8888/notebooks/simple_xgboost_example.ipynb#Export,-load-and-deploy-XGBoost-model-in-Triton-Inference-Server)
# * [Determine throughput and latency using Perf Analyzer](http://localhost:7001/notebooks/simple_xgboost_example.ipynb#Determine-throughput-and-latency-with-Perf-Analyzer)
# * [Find best configuration using Model Analyzer](http://localhost:7001/notebooks/simple_xgboost_example.ipynb#Find-best-configuration-using-Model-Analyzer)
# * [Deploy model with best configuration](http://localhost:7001/notebooks/simple_xgboost_example.ipynb#Deploy-model-with-best-configuration)
# * [Triton Client](http://localhost:7001/notebooks/simple_xgboost_example.ipynb#Triton-Client)
# * [Conclusion](http://localhost:7001/notebooks/simple_xgboost_example.ipynb#Conclusion)
# ## Requirements
#
# * Nvidia GPU (Pascal+ Recommended GPUs: T4, V100 or A100)
# * [Latest NVIDIA driver](https://docs.nvidia.com/datacenter/tesla/tesla-installation-notes/index.html)
# * [Docker](https://docs.docker.com/get-docker/)
# * [The NVIDIA container toolkit](https://docs.nvidia.com/datacenter/cloud-native/container-toolkit/install-guide.html#docker)
# ## Setup
#
# To begin, check that the NVIDIA driver has been installed correctly. The `nvidia-smi` command should run and output information about the GPUs on your system:"
# !nvidia-smi
# ## Install XGBoost and Sklearn
#
# We'd need to install XGBoost and SKlearn using the following pip3 commands inside the container as follows:
# Install sklearn and XGBoost
# !pip3 install -U scikit-learn xgboost
# ## Train XGBoost model
#
# If you have a pre-trained xgboost model, save it as `xgboost.model` and skip this step. We'll train a XGBoost model on random data in this section
# +
# Import required libraries
import numpy
from numpy import loadtxt
from xgboost import XGBClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
import os
import signal
import subprocess
# +
# Generate dummy data to perform binary classification
seed = 7
features = 9 # number of sample features
samples = 10000 # number of samples
X = numpy.random.rand(samples, features).astype('float32')
Y = numpy.random.randint(2, size=samples)
test_size = 0.33
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size=test_size, random_state=seed)
# -
model = XGBClassifier()
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
predictions = [round(value) for value in y_pred]
accuracy = accuracy_score(y_test, predictions)
print("Test Accuracy: {:.2f}".format(accuracy * 100.0))
# ## Export, load and deploy XGBoost model in Triton Inference Server
#
# For deploying the trained XGBoost model in Triton Inference Server, follow the steps below:
#
# **1. Create a model repository and save xgboost model checkpoint:**
#
# We'll need to create a model repository that looks as follows:
#
# ```
# model_repository/
# `-- fil
# |-- 1
# | `-- xgboost.model
# `-- config.pbtxt
# ```
# +
# Create directory to save the model
![ ! -d "/model_repository" ] && mkdir -p /model_repository/fil/1
# Save your xgboost model as xgboost.model
# For more information on saving xgboost model check https://xgboost.readthedocs.io/en/latest/python/python_intro.html#training
# Model can also be dumped to json format
model.save_model('/model_repository/fil/1/xgboost.model')
# -
# **Note:**
# The FIL backend's testing infrastructure includes a script for generating example models, putting them in the correct directory layout, and generating an associated config file. This can be helpful both for providing a template for your own models and for testing your Triton deployment. Please check this [link](https://github.com/triton-inference-server/fil_backend/blob/main/Example_Models.md) for the sample script.
# **2. Create and save config.pbtxt**
#
# To deploy the model in Triton Inference Server, we need to create and save a protobuf config file called config.pbtxt under `model_repository/fil/` directory that contains information about the model and the deployment. Sample config file is available here: [link](https://github.com/triton-inference-server/fil_backend#configuration)
#
# Essentially, the following parameters need to be updated as per your configuration
#
# ```
# name: "fil" # Name of the model directory (fil in our case)
# backend: "fil" # Triton FIL backend for deploying forest models
# max_batch_size: 8192
# input [
# {
# name: "input__0"
# data_type: TYPE_FP32
# dims: [ 9 ] # Input feature dimensions, in our sample case it's 9
# }
# ]
# output [
# {
# name: "output__0"
# data_type: TYPE_FP32
# dims: [ 2 ] # Output 2 for binary classification model
# }
# ]
# instance_group [{ kind: KIND_GPU }]
# parameters [
# {
# key: "model_type"
# value: { string_value: "xgboost" }
# },
# {
# key: "predict_proba"
# value: { string_value: "false" }
# },
# {
# key: "output_class"
# value: { string_value: "true" }
# },
# {
# key: "threshold"
# value: { string_value: "0.5" }
# },
# {
# key: "algo"
# value: { string_value: "ALGO_AUTO" }
# },
# {
# key: "storage_type"
# value: { string_value: "AUTO" }
# },
# {
# key: "blocks_per_sm"
# value: { string_value: "0" }
# }
# ]
# ```
#
# Triton server looks for this configuration file before deploying XGBoost model for inference. It'll setup the server parameters as per the configuration passed within config.pbtxt. Store the above config at `/model_repository/fil/` directory as config.pbtxt as follows:
#
# For more information on sample configs, please refer this [link](https://github.com/triton-inference-server/server/blob/main/docs/model_configuration.md)
# + language="bash"
# # Writing config to file
# cat > /model_repository/fil/config.pbtxt <<EOL
# name: "fil" # Name of the model directory (fil in our case)
# backend: "fil" # Triton FIL backend for deploying forest models
# max_batch_size: 8192
# input [
# {
# name: "input__0"
# data_type: TYPE_FP32
# dims: [ 9 ] # Input feature dimensions, in our sample case it's 9
# }
# ]
# output [
# {
# name: "output__0"
# data_type: TYPE_FP32
# dims: [ 2 ] # Output 2 for binary classification model
# }
# ]
# instance_group [{ kind: KIND_GPU }]
# parameters [
# {
# key: "model_type"
# value: { string_value: "xgboost" }
# },
# {
# key: "predict_proba"
# value: { string_value: "false" }
# },
# {
# key: "output_class"
# value: { string_value: "true" }
# },
# {
# key: "threshold"
# value: { string_value: "0.5" }
# },
# {
# key: "algo"
# value: { string_value: "ALGO_AUTO" }
# },
# {
# key: "storage_type"
# value: { string_value: "AUTO" }
# },
# {
# key: "blocks_per_sm"
# value: { string_value: "0" }
# }
# ]
#
# EOL
# -
# The model repository should look like this:
#
# ```
# model_repository/
# `-- fil
# |-- 1
# | `-- xgboost.model
# `-- config.pbtxt
# ```
#
# **3. Deploy the model in Triton Inference Server**
#
# Finally, we can deploy the xgboost model in Triton Inference Server using the following command:
# +
# Run the Triton Inference Server in a Subprocess from Jupyter notebook
triton_process = subprocess.Popen(["tritonserver", "--model-repository=/model_repository"], stdout=subprocess.PIPE, preexec_fn=os.setsid)
# -
# The above command should load the model and print the log `successfully loaded 'fil' version 1`. Triton server listens on the following endpoints:
#
# ```
# Port 8000 -> HTTP Service
# Port 8001 -> GRPC Service
# Port 8002 -> Metrics
# ```
#
# We can test the status of the server connection by running the curl command: `curl -v <IP of machine>:8000/v2/health/ready` which should return `HTTP/1.1 200 OK`
#
# **NOTE:-** In our case the IP of machine on which Triton Server and this notebook are currently running is `localhost`
# !curl -v localhost:8000/v2/health/ready
# ## Determine throughput and latency with Perf Analyzer
#
# Once the model is deployed for inference in Triton, we can measure its inference performance using `perf_analyzer`. The perf_analyzer application generates inference requests to the deployed model and measures the throughput and latency of those requests. For more information on `perf_analyzer` utility, please refer this [link](https://github.com/triton-inference-server/server/blob/main/docs/perf_analyzer.md)
# Install nvidia-pyindex
# !pip3 install nvidia-pyindex
# Install Triton client
# !pip3 install tritonclient[http]
# Run Perf analyzer to simulate incoming inference requests to Triton server
# Stabilize p99 latency with threshold of 5 msec and concurrency of incoming request from 10 to 15 with batch size 1
# !perf_analyzer -m fil --percentile=99 --latency-threshold=5 --concurrency-range=10:15 --async -b 1
# Stopping Triton Server before proceeding further
os.killpg(os.getpgid(triton_process.pid), signal.SIGTERM) # Send the signal to all the process groups
# ## Find best configuration using Model Analyzer
#
# [Triton Model Analyzer](https://github.com/triton-inference-server/model_analyzer) is a tool to profile and evaluate the best deployment configuration that maximizes inference performance of your model when deployed in Triton Inference Server. Using this tool, you can find the appropriate batch size, instances of your model etc. based on the constraints specified like maximum latency budget, minimum throughput and maximum GPU utilization limit. Model Analyzer installation steps are available here: [link](https://github.com/triton-inference-server/model_analyzer/blob/main/docs/install.md)
# Install library
# !pip3 install triton-model-analyzer
# Create a config file specifying the profiling constrains as follows:
# * `perf_throughput` - Specify minimum desired throughput.
#
#
# * `perf_latency` - Specify maximum tolerable latency or latency budget.
#
#
# * `gpu_used_memory` - Specify maximum GPU memory used by model.
# + language="bash"
# # Writing constraints to file
# cat > model_analyzer_constraints.yaml <<EOL
# model_repository: /model_repository/
# triton_launch_mode: "local"
# latency_budget: 5
# run_config_search_max_concurrency: 64
# run_config_search_max_instance_count: 3
# run_config_search_max_preferred_batch_size: 8
# profile_models:
# fil
#
# EOL
# -
# Run model_analyzer profiler on XGBoost model
# !model-analyzer profile -f model_analyzer_constraints.yaml --override-output-model-repository
# The above command will perform a search across various config parameters on the `fil` XGBoost model. Complete execution of this cell might take a while (30-40 mins) as the model analyzer searches for the optimum configuration based on the given constraints. When finished, model analyzer stores all of the profiling measurements it has taken in a binary file in the checkpoint directory. Now we can generate and visualize results using `analyze` command as follows:
# Install wkhtmltopdf to generate pdf reports
# !apt-get update && apt-get install -y wkhtmltopdf
![ ! -d "analysis_results" ] && mkdir analysis_results
# !model-analyzer analyze --analysis-models fil -e analysis_results
# The detailed summary report from model analyzer can be found under `/notebook/analysis_results/reports/summaries/fil/result_summary.pdf`. It'll look something like this:
# 
# 
# ## Deploy model with best configuration
#
# Now we can deploy the model with configuration that gives best throughput and latency numbers as evaluated by the model analyzer. In our case, model configuration `fil_i10` gives the best configuration. We can now copy this config.pbtxt to model directory and re-deploy Triton Inference Server
# + language="bash"
# # Change the best model configuration as per required constraints
# export best_model='fil_i0'
# cp -r output_model_repository/$best_model/ /model_repository/
# mkdir -p /model_repository/$best_model/1 && cp /model_repository/fil/1/xgboost.model /model_repository/$best_model/1/
# +
# Run Triton Inference Server in background from Jupyter notebook
triton_process = subprocess.Popen(["tritonserver", "--model-repository=/model_repository"], stdout=subprocess.PIPE, preexec_fn=os.setsid)
# -
# Run the perf_analyzer with same parameters
# Change the model as per your configuration
# !perf_analyzer -m fil_i0 --percentile=99 --latency-threshold=5 --concurrency-range=10:15 --async -b 1
# Stopping Triton Server before proceeding further
os.killpg(os.getpgid(triton_process.pid), signal.SIGTERM) # Send the signal to all the process groups
# ## Triton Client
#
# After model profiling is done and the final model is selected as per required configuration and deployed in Triton, we can now test the inference by sending real inference request from Triton Client and checking the accuracy of responses. For more information on installation steps, please check [Triton Client Github](https://github.com/triton-inference-server/client)
# +
# Run Triton Inference Server in background from Jupyter notebook
triton_process = subprocess.Popen(["tritonserver", "--model-repository=/model_repository"], stdout=subprocess.PIPE, preexec_fn=os.setsid)
# -
# Check client library can be imported
import numpy
import tritonclient.http as triton_http
# +
# Set up HTTP client.
http_client = triton_http.InferenceServerClient(
url='localhost:8000',
verbose=False,
concurrency=1
)
# Set up Triton input and output objects for both HTTP and GRPC
triton_input_http = triton_http.InferInput(
'input__0',
(X_test.shape[0], X_test.shape[1]),
'FP32'
)
triton_input_http.set_data_from_numpy(X_test, binary_data=True)
triton_output_http = triton_http.InferRequestedOutput(
'output__0',
binary_data=True
)
# Submit inference requests
request_http = http_client.infer(
'fil',
model_version='1',
inputs=[triton_input_http],
outputs=[triton_output_http]
)
# Get results as numpy arrays
result_http = request_http.as_numpy('output__0')
# Check that we got the same accuracy as previously
predictions = [round(value) for value in result_http]
accuracy = accuracy_score(y_test, predictions)
print("Accuracy: {:.2f}".format(accuracy * 100.0))
# -
# The above test accuracy score of the model deployed in Triton using FIL backend matches with the one previously computed using XGBoost library's predict function.
# Stopping Triton Server before proceeding further
os.killpg(os.getpgid(triton_process.pid), signal.SIGTERM) # Send the signal to all the process groups
# # Conclusion
#
# Triton FIL backend can be used for deploying tree based models trained in frameworks like LightGBM, Scikit-Learn, and cuML for fast GPU-based inference. Essentially, tree based models can now be deployed with other deep learning based models in Triton Inference Server seamlessly. Moreover, Model Analyzer utility tool can be used to profile the models and get the best deployment configuration that satisfy the deployment constraints. The trained model can then be deployed using the best configuration in Triton and Triton Client can be used for sending inference requests.
| notebooks/simple_xgboost_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="lWSdA0nx5-Wr" colab_type="code" colab={}
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn import linear_model
from sklearn import metrics
import matplotlib.pyplot as plt
# %matplotlib inline
# + id="ZX7PUPVJjgZz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 126} outputId="bba70985-b8c4-44ba-98c8-41254b866b59"
from google.colab import drive
drive.mount('/content/drive')
# + id="8uTaaE2hjhRr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="2b40a9d0-d270-4c46-e796-7f9d4f2e1f67"
# %cd /content/drive/My Drive/Colab Notebooks/ML_and_NN_course/module 1
# + id="XC3EXsSolAUa" colab_type="code" colab={}
from utils import *
# + [markdown] id="1V_niaY25-XO" colab_type="text"
# ## Dataset
# + id="xmGqSkbt5-XQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 226} outputId="accb8cfc-9d64-4bcc-ea77-0255bb24c8fc"
df=pd.read_csv("kc_house_data.csv") #Load the dataset from csv file
df.head()
# + id="xOUx0bBE5-Xf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 320} outputId="5f07ab08-8f04-4aec-82c7-dfb3829eda1b"
df.describe() #Gives the information about the data
# + id="5BHkzuTn5-Xr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 106} outputId="181e36cd-bcb9-4027-d5ee-d5633210c89d"
df.columns #Columns in dataframe
# + id="i-bkmLMh5-X5" colab_type="code" colab={}
# Dropping the id and date columns
df = df.drop(['id', 'date'],axis=1)
# + id="-6gBcDmcmDpK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 383} outputId="4b03889b-0da2-42e8-970a-718825d03d9a"
df.head(10)
# + [markdown] id="x_p2--OO5-YH" colab_type="text"
# ### Visualizing data
# + [markdown] id="HQALYUK25-YI" colab_type="text"
# More on Markers: https://matplotlib.org/3.1.1/api/markers_api.html
# + id="mLP37ctS5-YL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="de19f082-1245-4ad1-8a2a-7a3eebb7d48c"
plt.scatter(df['sqft_living'],df['price'],alpha=0.4,marker='1')
plt.title("bedrooms v/s price")
plt.xlabel("Sq. ft")
plt.ylabel("Price")
plt.show()
# + id="t90npbxL5-YT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="2bcda134-0e97-4237-966b-35b5a8f8af4e"
plt.scatter(df['floors'],df['price'],alpha=0.4,marker='+')
plt.title("bedrooms v/s price")
#plt.xlabel("No. of bedrooms")
plt.xlabel('floors')
plt.ylabel("Price")
plt.show()
# + [markdown] id="RG0Vd7K95-Yd" colab_type="text"
# ## Prepare the dataset
# + id="eC5jD6IE5-Ye" colab_type="code" colab={}
# First extract the target variable which is our House prices
Y = df.price.values
# Drop price from the house dataframe and create a matrix out of the house data
df = df.drop(['price'], axis=1)
X=np.matrix(df)
# Store the column/feature names into a list "colnames"
colnames = df.columns
# + [markdown] id="_8OOhiHXBPL7" colab_type="text"
# > **Training data:** actual dataset that we use to train the model. The model sees and learns from this data.
#
#
# > **Validation data:** sample of data used to provide an unbiased evaluation of a model fit on the training dataset while tuning model hyperparameters
#
#
#
# + id="fbgIU_HD5-Yo" colab_type="code" colab={}
#Splitting the data into training and validation dataset
X_train,X_test,Y_train,Y_test = train_test_split(X,Y,train_size = 0.8,random_state=3)
# + [markdown] id="ZcuRrgxL5-Y0" colab_type="text"
# ## Training the model
# 
# + id="x5IomKGB5-Y2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="e76b4116-4ca1-4014-b255-7e07eaebc985"
# Using Linear Regression
lr = linear_model.LinearRegression(normalize=True) #Creates the Linear regression model
lr.fit(X_train,Y_train)
# + id="yinbMMPS5-ZC" colab_type="code" colab={}
# lr??
# + id="An9VjtZy5-ZO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 339} outputId="82598dab-bb29-42d1-c980-76930aad12f9"
ranking(np.abs(lr.coef_), colnames)
# + id="1mw3N3e45-ZY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 53} outputId="60e79603-6f60-4790-d97f-555996d05754"
predictions=lr.predict(X_test)
test_error=metrics.mean_absolute_error(Y_test,predictions)
print("Mean absolute error on Test set: ",test_error)
predictions=lr.predict(X_train)
training_error=metrics.mean_absolute_error(Y_train,predictions)
print("Mean absolute error on Training set: ",training_error)
# + [markdown] id="4eR2B5HU5-Zi" colab_type="text"
# ## Prediction
# + id="BVZn_dbJ5-Zj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 53} outputId="64e5d11f-291b-410d-bf5f-afc0ea0f845b"
index=4 #Change Index to see how it performs on other examples
check=X_train[index]
check=np.array(check).reshape(1,-1)
print("predicted Value:",lr.predict(check))
print("True value: ",Y_train[index])
# + id="zAjngcMZ5-Zq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="6bae591b-c6e8-4d6b-af17-3b2450eb7234"
Y.max(),Y.min(),Y.mean()
# + id="BldZ8SCt5-Zz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 106} outputId="6b25ee2e-afc1-42d5-e154-e667c9bd9ff8"
X_train[index]
# + id="4BZ7aLrp5-Z8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 356} outputId="26a7b528-d84b-4eac-ed3b-d7956f5cd912"
df.iloc[9]
# + [markdown] id="JZ47BiD35-aD" colab_type="text"
# ## Checking Prediction for house
# + id="s-K0jITU5-aM" colab_type="code" colab={}
# change these values according to your house
bedrooms = 3.0000
bathrooms = 2.0000
sqft_living = 1680.0000
sqft_lot = 8080.0000
floors = 1.0000
waterfront = 0.0000
view = 0.0000
condition = 3.0000
grade = 8.0000
sqft_above = 1680.0000
sqft_basement = 0.0000
yr_built = 1987.0000
yr_renovated = 0.0000
zipcode = 98074.0000
lat = 47.6168
long = -122.0450
sqft_living15 = 1800.0000
sqft_lot15 = 7503.0000
#Creating array from the values taken
my_house=np.array([bedrooms,bathrooms, sqft_living, sqft_lot ,floors ,waterfront ,view ,condition, grade,sqft_above,
sqft_basement, yr_built, yr_renovated,zipcode, lat, long ,sqft_living15 ,sqft_lot15 ])
# + id="kNRFKCsN5-aT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="bd6036b0-4cab-4eaa-b9a9-0864aa02b4fc"
print("predicted Value:",lr.predict(my_house.reshape(1,-1))) #Predicting the value
# + id="3jlkog-G5-aZ" colab_type="code" colab={}
| module 1/Linear_Regression_(module_1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## seaborn excercise
#
# these plots are only for numerical values:
#
# 1.dist plot ------> univariate
# 2.joint plot ------> bivariate
# 3.pair plot ------> multivariate
#
# Note----> hue can be used to view the ploting with reference to a categorical feature
#
# these plots are only for catrgorical values:
#
# 1.box plot ------> five line summary
# 2.violin plot ------> gives both the boxplot and kde
# 3.count plot ------> count of each feature
# 4.bar plot ------> same as countplot
#
from sklearn.datasets import load_iris
import pandas as pd
iris = load_iris()
dir(iris)
iris.feature_names
df = pd.DataFrame(iris.data)
df.columns = ['sepal length (cm)',
'sepal width (cm)',
'petal length (cm)',
'petal width (cm)']
df
df['target'] = iris.target
df
import seaborn as sns
df['target'] = df.target.apply(lambda x:iris.target_names[x])
df
df.corr()
sns.heatmap(df.corr(),cmap="YlGnBu",annot=True, fmt="f",linewidths=.1)
# +
#join plot -- univariate or bivariate
# -
sns.jointplot(x='sepal length (cm)', y='sepal width (cm)', data = df, kind = "hex")#kind : { "scatter" | "reg" | "resid" | "kde" | "hex" }
sns.jointplot(x='petal length (cm)', y='petal width (cm)', data = df, kind = "kde" )
# +
## pair plot used when we have features are more than two independent features
sns.pairplot(df,hue='target',kind='scatter',palette="husl")#kind : {scatter,reg}
# -
# +
### dist plot to check the distribution of a column feature
sns.distplot(df['sepal length (cm)'])
# +
sns.distplot(df['sepal width (cm)'],kde=False,bins=10)
# -
sns.distplot(df['sepal width (cm)'],kde=True,bins=10)
sns.distplot(df['petal length (cm)'],kde=True,bins=10)
sns.distplot(df['petal width (cm)'],kde=True,bins=10)
##showes the count of the catogorical feature
sns.countplot(x='target',data=df)
# +
## bar plot x and y both axis is given for plotting
sns.barplot(x='target',y='sepal length (cm)',data=df)
# -
sns.barplot(x='target',y='sepal width (cm)',data=df)
sns.barplot(x='target',y='petal length (cm)',data=df)
sns.barplot(x='target',y='petal width (cm)',data=df)
### boxplot gives the information into a five line summary
#sns.boxplot("Flower_name","target",data=df)
sns.boxplot(data=df,orient = "v")
### violin plot - to see both the data distribution in kernel density estimator and boxplot
sns.violinplot(data=df,orient="h")
| Seaborn_Visuals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.9 64-bit (''venv'': venv)'
# name: pythonjvsc74a57bd031cce29e8fb201be9f4f5076271a9ab310f410b2e79720eb45fe746a4bcc676b
# ---
import med_dataloader as med_dl
# +
import shutil
try:
shutil.rmtree("Test_Dataset_segmentation_TF")
except:
pass
# + tags=[]
num_classes = 3
med_dl.generate_dataset(data_dir=r"Test_Dataset_segmentation",
imgA_label="CT",
imgB_label="Labels",
input_size=192,
is_B_categorical=True,
num_classes=num_classes,
norm_boundsA=None,
norm_boundsB=None,
use_3D=True,
)
train_ds, valid_ds, test_ds = med_dl.get_dataset(data_dir=r"Test_Dataset_segmentation_TF",
percentages=[1,0,0],
batch_size=1,
train_augmentation=True,
random_crop_size=None,
random_rotate=False,
random_flip=False,
)
# +
import matplotlib.pyplot as plt
slice_index = 80
for batch in train_ds.take(3):
volume_batch, label_batch = batch
for volume, label in zip(volume_batch, label_batch):
plt.subplot(121)
plt.imshow(volume[:,:,slice_index,0], cmap="gray")
plt.subplot(122)
for i in range(num_classes):
plt.imshow(label[:,:,slice_index,i], cmap="gray", alpha=0.5)
plt.show()
# -
| examples/basic_usage_segmentation_3D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
import os
import pickle
from torch.utils.data import Dataset
from torchvision import transforms
from torch.utils.data import DataLoader
from tqdm import tqdm
from time import time
import torch.nn as nn
import torch.nn.parallel
import torch.utils.data
from torch.autograd import Variable
import numpy as np
import torch.nn.functional as F
import torch.nn as nn
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits import mplot3d
# +
# from open3d import *
# import numpy as np
# -
NUM_CLASSES = 21
block_points = 8091
root = '/media/ken/B60A03C60A03829B/data/scannet'
BATCH_SIZE = 8
NUM_POINT = 8091
# +
import numpy as np
def normalize_data(batch_data):
""" Normalize the batch data, use coordinates of the block centered at origin,
Input:
BxNxC array
Output:
BxNxC array
"""
B, N, C = batch_data.shape
normal_data = np.zeros((B, N, C))
for b in range(B):
pc = batch_data[b]
centroid = np.mean(pc, axis=0)
pc = pc - centroid
m = np.max(np.sqrt(np.sum(pc ** 2, axis=1)))
pc = pc / m
normal_data[b] = pc
return normal_data
def shuffle_data(data, labels):
""" Shuffle data and labels.
Input:
data: B,N,... numpy array
label: B,... numpy array
Return:
shuffled data, label and shuffle indices
"""
idx = np.arange(len(labels))
np.random.shuffle(idx)
return data[idx, ...], labels[idx], idx
def shuffle_points(batch_data):
""" Shuffle orders of points in each point cloud -- changes FPS behavior.
Use the same shuffling idx for the entire batch.
Input:
BxNxC array
Output:
BxNxC array
"""
idx = np.arange(batch_data.shape[1])
np.random.shuffle(idx)
return batch_data[:,idx,:]
def rotate_point_cloud(batch_data):
""" Randomly rotate the point clouds to augument the dataset
rotation is per shape based along up direction
Input:
BxNx3 array, original batch of point clouds
Return:
BxNx3 array, rotated batch of point clouds
"""
rotated_data = np.zeros(batch_data.shape, dtype=np.float32)
for k in range(batch_data.shape[0]):
rotation_angle = np.random.uniform() * 2 * np.pi
cosval = np.cos(rotation_angle)
sinval = np.sin(rotation_angle)
rotation_matrix = np.array([[cosval, 0, sinval],
[0, 1, 0],
[-sinval, 0, cosval]])
shape_pc = batch_data[k, ...]
rotated_data[k, ...] = np.dot(shape_pc.reshape((-1, 3)), rotation_matrix)
return rotated_data
def rotate_point_cloud_z(batch_data):
""" Randomly rotate the point clouds to augument the dataset
rotation is per shape based along up direction
Input:
BxNx3 array, original batch of point clouds
Return:
BxNx3 array, rotated batch of point clouds
"""
rotated_data = np.zeros(batch_data.shape, dtype=np.float32)
for k in range(batch_data.shape[0]):
rotation_angle = np.random.uniform() * 2 * np.pi
cosval = np.cos(rotation_angle)
sinval = np.sin(rotation_angle)
rotation_matrix = np.array([[cosval, sinval, 0],
[-sinval, cosval, 0],
[0, 0, 1]])
shape_pc = batch_data[k, ...]
rotated_data[k, ...] = np.dot(shape_pc.reshape((-1, 3)), rotation_matrix)
return rotated_data
def rotate_point_cloud_with_normal(batch_xyz_normal):
''' Randomly rotate XYZ, normal point cloud.
Input:
batch_xyz_normal: B,N,6, first three channels are XYZ, last 3 all normal
Output:
B,N,6, rotated XYZ, normal point cloud
'''
for k in range(batch_xyz_normal.shape[0]):
rotation_angle = np.random.uniform() * 2 * np.pi
cosval = np.cos(rotation_angle)
sinval = np.sin(rotation_angle)
rotation_matrix = np.array([[cosval, 0, sinval],
[0, 1, 0],
[-sinval, 0, cosval]])
shape_pc = batch_xyz_normal[k,:,0:3]
shape_normal = batch_xyz_normal[k,:,3:6]
batch_xyz_normal[k,:,0:3] = np.dot(shape_pc.reshape((-1, 3)), rotation_matrix)
batch_xyz_normal[k,:,3:6] = np.dot(shape_normal.reshape((-1, 3)), rotation_matrix)
return batch_xyz_normal
def rotate_perturbation_point_cloud_with_normal(batch_data, angle_sigma=0.06, angle_clip=0.18):
""" Randomly perturb the point clouds by small rotations
Input:
BxNx6 array, original batch of point clouds and point normals
Return:
BxNx3 array, rotated batch of point clouds
"""
rotated_data = np.zeros(batch_data.shape, dtype=np.float32)
for k in range(batch_data.shape[0]):
angles = np.clip(angle_sigma*np.random.randn(3), -angle_clip, angle_clip)
Rx = np.array([[1,0,0],
[0,np.cos(angles[0]),-np.sin(angles[0])],
[0,np.sin(angles[0]),np.cos(angles[0])]])
Ry = np.array([[np.cos(angles[1]),0,np.sin(angles[1])],
[0,1,0],
[-np.sin(angles[1]),0,np.cos(angles[1])]])
Rz = np.array([[np.cos(angles[2]),-np.sin(angles[2]),0],
[np.sin(angles[2]),np.cos(angles[2]),0],
[0,0,1]])
R = np.dot(Rz, np.dot(Ry,Rx))
shape_pc = batch_data[k,:,0:3]
shape_normal = batch_data[k,:,3:6]
rotated_data[k,:,0:3] = np.dot(shape_pc.reshape((-1, 3)), R)
rotated_data[k,:,3:6] = np.dot(shape_normal.reshape((-1, 3)), R)
return rotated_data
def rotate_point_cloud_by_angle(batch_data, rotation_angle):
""" Rotate the point cloud along up direction with certain angle.
Input:
BxNx3 array, original batch of point clouds
Return:
BxNx3 array, rotated batch of point clouds
"""
rotated_data = np.zeros(batch_data.shape, dtype=np.float32)
for k in range(batch_data.shape[0]):
#rotation_angle = np.random.uniform() * 2 * np.pi
cosval = np.cos(rotation_angle)
sinval = np.sin(rotation_angle)
rotation_matrix = np.array([[cosval, 0, sinval],
[0, 1, 0],
[-sinval, 0, cosval]])
shape_pc = batch_data[k,:,0:3]
rotated_data[k,:,0:3] = np.dot(shape_pc.reshape((-1, 3)), rotation_matrix)
return rotated_data
def rotate_point_cloud_by_angle_with_normal(batch_data, rotation_angle):
""" Rotate the point cloud along up direction with certain angle.
Input:
BxNx6 array, original batch of point clouds with normal
scalar, angle of rotation
Return:
BxNx6 array, rotated batch of point clouds iwth normal
"""
rotated_data = np.zeros(batch_data.shape, dtype=np.float32)
for k in range(batch_data.shape[0]):
#rotation_angle = np.random.uniform() * 2 * np.pi
cosval = np.cos(rotation_angle)
sinval = np.sin(rotation_angle)
rotation_matrix = np.array([[cosval, 0, sinval],
[0, 1, 0],
[-sinval, 0, cosval]])
shape_pc = batch_data[k,:,0:3]
shape_normal = batch_data[k,:,3:6]
rotated_data[k,:,0:3] = np.dot(shape_pc.reshape((-1, 3)), rotation_matrix)
rotated_data[k,:,3:6] = np.dot(shape_normal.reshape((-1,3)), rotation_matrix)
return rotated_data
def rotate_perturbation_point_cloud(batch_data, angle_sigma=0.06, angle_clip=0.18):
""" Randomly perturb the point clouds by small rotations
Input:
BxNx3 array, original batch of point clouds
Return:
BxNx3 array, rotated batch of point clouds
"""
rotated_data = np.zeros(batch_data.shape, dtype=np.float32)
for k in range(batch_data.shape[0]):
angles = np.clip(angle_sigma*np.random.randn(3), -angle_clip, angle_clip)
Rx = np.array([[1,0,0],
[0,np.cos(angles[0]),-np.sin(angles[0])],
[0,np.sin(angles[0]),np.cos(angles[0])]])
Ry = np.array([[np.cos(angles[1]),0,np.sin(angles[1])],
[0,1,0],
[-np.sin(angles[1]),0,np.cos(angles[1])]])
Rz = np.array([[np.cos(angles[2]),-np.sin(angles[2]),0],
[np.sin(angles[2]),np.cos(angles[2]),0],
[0,0,1]])
R = np.dot(Rz, np.dot(Ry,Rx))
shape_pc = batch_data[k, ...]
rotated_data[k, ...] = np.dot(shape_pc.reshape((-1, 3)), R)
return rotated_data
def jitter_point_cloud(batch_data, sigma=0.01, clip=0.05):
""" Randomly jitter points. jittering is per point.
Input:
BxNx3 array, original batch of point clouds
Return:
BxNx3 array, jittered batch of point clouds
"""
B, N, C = batch_data.shape
assert(clip > 0)
jittered_data = np.clip(sigma * np.random.randn(B, N, C), -1*clip, clip)
jittered_data += batch_data
return jittered_data
def shift_point_cloud(batch_data, shift_range=0.1):
""" Randomly shift point cloud. Shift is per point cloud.
Input:
BxNx3 array, original batch of point clouds
Return:
BxNx3 array, shifted batch of point clouds
"""
B, N, C = batch_data.shape
shifts = np.random.uniform(-shift_range, shift_range, (B,3))
for batch_index in range(B):
batch_data[batch_index,:,:] += shifts[batch_index,:]
return batch_data
def random_scale_point_cloud(batch_data, scale_low=0.8, scale_high=1.25):
""" Randomly scale the point cloud. Scale is per point cloud.
Input:
BxNx3 array, original batch of point clouds
Return:
BxNx3 array, scaled batch of point clouds
"""
B, N, C = batch_data.shape
scales = np.random.uniform(scale_low, scale_high, B)
for batch_index in range(B):
batch_data[batch_index,:,:] *= scales[batch_index]
return batch_data
def random_point_dropout(batch_pc, max_dropout_ratio=0.875):
''' batch_pc: BxNx3 '''
for b in range(batch_pc.shape[0]):
dropout_ratio = np.random.random()*max_dropout_ratio # 0~0.875
drop_idx = np.where(np.random.random((batch_pc.shape[1]))<=dropout_ratio)[0]
if len(drop_idx)>0:
batch_pc[b,drop_idx,:] = batch_pc[b,0,:] # set to the first point
return batch_pc
# -
# # Scannet dataloder
# +
class ScannetDataset():
def __init__(self, root, block_points=2048, split='train', with_rgb = True):
self.npoints = block_points
self.root = root
self.with_rgb = with_rgb
self.split = split
self.data_filename = os.path.join(self.root, 'scannet_%s_rgb21c_pointid.pickle'%(split))
with open(self.data_filename,'rb') as fp:
self.scene_points_list = pickle.load(fp)
self.semantic_labels_list = pickle.load(fp)
self.scene_points_id = pickle.load(fp)
self.scene_points_num = pickle.load(fp)
if split=='train':
labelweights = np.zeros(21)
for seg in self.semantic_labels_list:
tmp,_ = np.histogram(seg,range(22))
labelweights += tmp
labelweights = labelweights.astype(np.float32)
labelweights = labelweights/np.sum(labelweights)
self.labelweights = np.power(np.amax(labelweights[1:]) / labelweights, 1/3.0)
print(self.labelweights)
elif split=='val':
self.labelweights = np.ones(21)
def __getitem__(self, index):
if self.with_rgb:
point_set = self.scene_points_list[index]
else:
point_set = self.scene_points_list[index][:, 0:3]
semantic_seg = self.semantic_labels_list[index].astype(np.int32)
coordmax = np.max(point_set[:, 0:3],axis=0)
coordmin = np.min(point_set[:, 0:3],axis=0)
isvalid = False
for i in range(10):
curcenter = point_set[np.random.choice(len(semantic_seg),1)[0],0:3]
curmin = curcenter-[0.75,0.75,1.5]
curmax = curcenter+[0.75,0.75,1.5]
curmin[2] = coordmin[2]
curmax[2] = coordmax[2]
curchoice = np.sum((point_set[:, 0:3]>=(curmin-0.2))*(point_set[:, 0:3]<=(curmax+0.2)),axis=1)==3
cur_point_set = point_set[curchoice,0:3]
cur_point_full = point_set[curchoice,:]
cur_semantic_seg = semantic_seg[curchoice]
if len(cur_semantic_seg)==0:
continue
mask = np.sum((cur_point_set>=(curmin-0.01))*(cur_point_set<=(curmax+0.01)),axis=1)==3
vidx = np.ceil((cur_point_set[mask,:]-curmin)/(curmax-curmin)*[31.0,31.0,62.0])
vidx = np.unique(vidx[:,0]*31.0*62.0+vidx[:,1]*62.0+vidx[:,2])
isvalid = np.sum(cur_semantic_seg>0)/len(cur_semantic_seg)>=0.7 and len(vidx)/31.0/31.0/62.0>=0.02
if isvalid:
break
choice = np.random.choice(len(cur_semantic_seg), self.npoints, replace=True)
point_set = cur_point_full[choice,:]
semantic_seg = cur_semantic_seg[choice]
mask = mask[choice]
sample_weight = self.labelweights[semantic_seg]
sample_weight *= mask
return point_set, semantic_seg, sample_weight
def __len__(self):
return len(self.scene_points_list)
class ScannetDatasetWholeScene():
def __init__(self, root, block_points=8192, split='val', with_rgb = True):
self.npoints = block_points
self.root = root
self.with_rgb = with_rgb
self.split = split
self.data_filename = os.path.join(self.root, 'scannet_%s_rgb21c_pointid.pickle'%(split))
with open(self.data_filename,'rb') as fp:
self.scene_points_list = pickle.load(fp)
self.semantic_labels_list = pickle.load(fp)
self.scene_points_id = pickle.load(fp)
self.scene_points_num = pickle.load(fp)
if split=='train':
labelweights = np.zeros(21)
for seg in self.semantic_labels_list:
tmp,_ = np.histogram(seg,range(22))
labelweights += tmp
labelweights = labelweights.astype(np.float32)
labelweights = labelweights/np.sum(labelweights)
self.labelweights = 1/np.log(1.2+labelweights)
elif split=='val':
self.labelweights = np.ones(21)
def __getitem__(self, index):
if self.with_rgb:
point_set_ini = self.scene_points_list[index]
else:
point_set_ini = self.scene_points_list[index][:, 0:3]
semantic_seg_ini = self.semantic_labels_list[index].astype(np.int32)
coordmax = np.max(point_set_ini[:, 0:3],axis=0)
coordmin = np.min(point_set_ini[:, 0:3],axis=0)
nsubvolume_x = np.ceil((coordmax[0]-coordmin[0])/1.5).astype(np.int32)
nsubvolume_y = np.ceil((coordmax[1]-coordmin[1])/1.5).astype(np.int32)
point_sets = list()
semantic_segs = list()
sample_weights = list()
for i in range(nsubvolume_x):
for j in range(nsubvolume_y):
curmin = coordmin+[i*1.5,j*1.5,0]
curmax = coordmin+[(i+1)*1.5,(j+1)*1.5,coordmax[2]-coordmin[2]]
curchoice = np.sum((point_set_ini[:, 0:3]>=(curmin-0.2))*(point_set_ini[:, 0:3]<=(curmax+0.2)),axis=1)==3
cur_point_set = point_set_ini[curchoice,0:3]
cur_point_full = point_set_ini[curchoice,:]
cur_semantic_seg = semantic_seg_ini[curchoice]
if len(cur_semantic_seg)==0:
continue
mask = np.sum((cur_point_set>=(curmin-0.001))*(cur_point_set<=(curmax+0.001)),axis=1)==3
choice = np.random.choice(len(cur_semantic_seg), self.npoints, replace=True)
point_set = cur_point_full[choice,:] # Nx3/6
semantic_seg = cur_semantic_seg[choice] # N
mask = mask[choice]
if sum(mask)/float(len(mask))<0.01:
continue
sample_weight = self.labelweights[semantic_seg]
sample_weight *= mask # N
point_sets.append(np.expand_dims(point_set,0)) # 1xNx3
semantic_segs.append(np.expand_dims(semantic_seg,0)) # 1xN
sample_weights.append(np.expand_dims(sample_weight,0)) # 1xN
point_sets = np.concatenate(tuple(point_sets),axis=0)
semantic_segs = np.concatenate(tuple(semantic_segs),axis=0)
sample_weights = np.concatenate(tuple(sample_weights),axis=0)
return point_sets, semantic_segs, sample_weights
def __len__(self):
return len(self.scene_points_list)
# -
TRAIN_DATASET = ScannetDataset(root, split='val')
VAL_DATASET = ScannetDataset(root, split='val')
train_data_loader = DataLoader(VAL_DATASET, batch_size=2, shuffle=True, num_workers=4)
# +
# testDataLoader = DataLoader(VAL_DATASET, batch_size=2, shuffle=True, num_workers=4)
# -
# # visualize
for i, data in tqdm(enumerate(train_data_loader), total=len(train_data_loader), smoothing=0.9, disable=True):
points, target,w = data
points.shape
target.shape
# +
# def visualize_pointcloud(xyz, rgb=None):
# pcd = PointCloud()
# pcd.points = Vector3dVector(xyz) # XYZ points
# if rgb != None:
# pcd.colors = Vector3dVector(rgb/ 255.0) #open3d requires colors (RGB) to be in range[0,1]
# draw_geometries([pcd])
# return
# +
# augmentation data
# -
def rotate_point_cloud_with_r_group(batch_data):
""" Randomly perturb the point clouds by small rotations
Input:
BxNx3 array, original batch of point clouds
Return:
BxNx3 array, rotated batch of point clouds
"""
G4 = np.array([[[1., 0., 0.],
[0., 1., 0.],
[0., 0., 1.]],
[[1., 0., 0.],
[0., 0., -1.],
[0., 1., 0.]],
[[1., 0., 0.],
[0., -1., 0.],
[0., 0., -1.]],
[[1., 0., 0.],
[0., 0., 1.],
[0., -1., 0.]], ])
G1_grouped_xyz = torch.mm(batch_data, G4[1])
return G1_grouped_xyz
# +
# model_training
# +
def timeit(tag, t):
print("{}: {}s".format(tag, time() - t))
return time()
def pc_normalize(pc):
l = pc.shape[0]
centroid = np.mean(pc, axis=0)
pc = pc - centroid
m = np.max(np.sqrt(np.sum(pc**2, axis=1)))
pc = pc / m
return pc
def square_distance(src, dst):
"""
Calculate Euclid distance between each two points.
src^T * dst = xn * xm + yn * ym + zn * zm;
sum(src^2, dim=-1) = xn*xn + yn*yn + zn*zn;
sum(dst^2, dim=-1) = xm*xm + ym*ym + zm*zm;
dist = (xn - xm)^2 + (yn - ym)^2 + (zn - zm)^2
= sum(src**2,dim=-1)+sum(dst**2,dim=-1)-2*src^T*dst
Input:
src: source points, [B, N, C]
dst: target points, [B, M, C]
Output:
dist: per-point square distance, [B, N, M]
"""
B, N, _ = src.shape
_, M, _ = dst.shape
dist = -2 * torch.matmul(src, dst.permute(0, 2, 1))
dist += torch.sum(src ** 2, -1).view(B, N, 1)
dist += torch.sum(dst ** 2, -1).view(B, 1, M)
return dist
def index_points(points, idx):
"""
Input:
points: input points data, [B, N, C]
idx: sample index data, [B, S]
Return:
new_points:, indexed points data, [B, S, C]
"""
device = points.device
B = points.shape[0]
view_shape = list(idx.shape)
view_shape[1:] = [1] * (len(view_shape) - 1)
repeat_shape = list(idx.shape)
repeat_shape[0] = 1
batch_indices = torch.arange(B, dtype=torch.long).to(device).view(view_shape).repeat(repeat_shape)
new_points = points[batch_indices, idx, :]
return new_points
def farthest_point_sample(xyz, npoint):
"""
Input:
xyz: pointcloud data, [B, N, 3]
npoint: number of samples
Return:
centroids: sampled pointcloud index, [B, npoint]
"""
device = xyz.device
B, N, C = xyz.shape
centroids = torch.zeros(B, npoint, dtype=torch.long).to(device)
distance = torch.ones(B, N).to(device) * 1e10
farthest = torch.randint(0, N, (B,), dtype=torch.long).to(device)
batch_indices = torch.arange(B, dtype=torch.long).to(device)
for i in range(npoint):
centroids[:, i] = farthest
centroid = xyz[batch_indices, farthest, :].view(B, 1, 3)
dist = torch.sum((xyz - centroid) ** 2, -1)
mask = dist < distance
distance[mask] = dist[mask]
farthest = torch.max(distance, -1)[1]
return centroids
def query_ball_point(radius, nsample, xyz, new_xyz):
"""
Input:
radius: local region radius
nsample: max sample number in local region
xyz: all points, [B, N, 3]
new_xyz: query points, [B, S, 3]
Return:
group_idx: grouped points index, [B, S, nsample]
"""
device = xyz.device
B, N, C = xyz.shape
_, S, _ = new_xyz.shape
group_idx = torch.arange(N, dtype=torch.long).to(device).view(1, 1, N).repeat([B, S, 1])
sqrdists = square_distance(new_xyz, xyz)
group_idx[sqrdists > radius ** 2] = N
group_idx = group_idx.sort(dim=-1)[0][:, :, :nsample]
group_first = group_idx[:, :, 0].view(B, S, 1).repeat([1, 1, nsample])
mask = group_idx == N
group_idx[mask] = group_first[mask]
return group_idx
def sample_and_group(npoint, radius, nsample, xyz, points, returnfps=False):
"""
Input:
npoint:
radius:
nsample:
xyz: input points position data, [B, N, 3]
points: input points data, [B, N, D]
Return:
new_xyz: sampled points position data, [B, npoint, nsample, 3]
new_points: sampled points data, [B, npoint, nsample, 3+D]
"""
B, N, C = xyz.shape
S = npoint
fps_idx = farthest_point_sample(xyz, npoint) # [B, npoint, C]
torch.cuda.empty_cache()
new_xyz = index_points(xyz, fps_idx)
torch.cuda.empty_cache()
idx = query_ball_point(radius, nsample, xyz, new_xyz)
torch.cuda.empty_cache()
grouped_xyz = index_points(xyz, idx) # [B, npoint, nsample, C]
torch.cuda.empty_cache()
grouped_xyz_norm = grouped_xyz - new_xyz.view(B, S, 1, C)
torch.cuda.empty_cache()
if points is not None:
grouped_points = index_points(points, idx)
new_points = torch.cat([grouped_xyz_norm, grouped_points], dim=-1) # [B, npoint, nsample, C+D]
else:
new_points = grouped_xyz_norm
if returnfps:
return new_xyz, new_points, grouped_xyz, fps_idx
else:
return new_xyz, new_points
def sample_and_group_all(xyz, points):
"""
Input:
xyz: input points position data, [B, N, 3]
points: input points data, [B, N, D]
Return:
new_xyz: sampled points position data, [B, 1, 3]
new_points: sampled points data, [B, 1, N, 3+D]
"""
device = xyz.device
B, N, C = xyz.shape
new_xyz = torch.zeros(B, 1, C).to(device)
grouped_xyz = xyz.view(B, 1, N, C)
if points is not None:
new_points = torch.cat([grouped_xyz, points.view(B, 1, N, -1)], dim=-1)
else:
new_points = grouped_xyz
return new_xyz, new_points
# +
def square_distance_conv(src, dst):
"""
Calculate Euclid distance between each two points.
src^T * dst = xn * xm + yn * ym + zn * zm;
sum(src^2, dim=-1) = xn*xn + yn*yn + zn*zn;
sum(dst^2, dim=-1) = xm*xm + ym*ym + zm*zm;
dist = (xn - xm)^2 + (yn - ym)^2 + (zn - zm)^2
= sum(src**2,dim=-1)+sum(dst**2,dim=-1)-2*src^T*dst
Input:
src: source points, [B, N, C]
dst: target points, [B, M, C]
Output:
dist: per-point square distance, [B, N, M]
"""
B, N, _ = src.shape
_, M, _ = dst.shape
dist = -2 * torch.matmul(src, dst.permute(0, 2, 1))
dist += torch.sum(src ** 2, -1).view(B, N, 1)
dist += torch.sum(dst ** 2, -1).view(B, 1, M)
return dist
def index_points_conv(points, idx):
"""
Input:
points: input points data, [B, N, C]
idx: sample index data, [B, S]
Return:
new_points:, indexed points data, [B, S, C]
"""
device = points.device
B = points.shape[0]
view_shape = list(idx.shape)
view_shape[1:] = [1] * (len(view_shape) - 1)
repeat_shape = list(idx.shape)
repeat_shape[0] = 1
batch_indices = torch.arange(B, dtype=torch.long).to(device).view(view_shape).repeat(repeat_shape)
new_points = points[batch_indices, idx, :]
return new_points
def farthest_point_sample_conv(xyz, npoint):
"""
Input:
xyz: pointcloud data, [B, N, C]
npoint: number of samples
Return:
centroids: sampled pointcloud index, [B, npoint]
"""
#import ipdb; ipdb.set_trace()
device = xyz.device
B, N, C = xyz.shape
centroids = torch.zeros(B, npoint, dtype=torch.long).to(device)
distance = torch.ones(B, N).to(device) * 1e10
farthest = torch.randint(0, N, (B,), dtype=torch.long).to(device)
batch_indices = torch.arange(B, dtype=torch.long).to(device)
# -----------------------------------------------------------------------------------------
# Chien.dotruong 12-11-2020 Add
chien_centroids = torch.mean(xyz, dim=1)
chien_distance = torch.sum((xyz - chien_centroids.view(B, 1, C)) ** 2, dim=2)
farthest = torch.argmax(chien_distance.reshape(B,N),dim=1)
# -----------------------------------------------------------------------------------------
for i in range(npoint):
centroids[:, i] = farthest
centroid = xyz[batch_indices, farthest, :].view(B, 1, 3)
dist = torch.sum((xyz - centroid) ** 2, -1)
mask = dist < distance
distance[mask] = dist[mask]
farthest = torch.max(distance, -1)[1]
return centroids
def query_ball_point_conv(radius, nsample, xyz, new_xyz):
"""
Input:
radius: local region radius
nsample: max sample number in local region
xyz: all points, [B, N, C]
new_xyz: query points, [B, S, C]
Return:
group_idx: grouped points index, [B, S, nsample]
"""
device = xyz.device
B, N, C = xyz.shape
_, S, _ = new_xyz.shape
group_idx = torch.arange(N, dtype=torch.long).to(device).view(1, 1, N).repeat([B, S, 1])
sqrdists = square_distance_conv(new_xyz, xyz)
group_idx[sqrdists > radius ** 2] = N
group_idx = group_idx.sort(dim=-1)[0][:, :, :nsample]
group_first = group_idx[:, :, 0].view(B, S, 1).repeat([1, 1, nsample])
mask = group_idx == N
group_idx[mask] = group_first[mask]
return group_idx
def knn_point_conv(nsample, xyz, new_xyz):
"""
Input:
nsample: max sample number in local region
xyz: all points, [B, N, C]
new_xyz: query points, [B, S, C]
Return:
group_idx: grouped points index, [B, S, nsample]
"""
sqrdists = square_distance_conv(new_xyz, xyz)
_, group_idx = torch.topk(sqrdists, nsample, dim = -1, largest=False, sorted=False)
return group_idx
def sample_and_group_conv(npoint, nsample, xyz, points, density_scale = None):
"""
Input:
npoint:
nsample:
xyz: input points position data, [B, N, C]
points: input points data, [B, N, D]
Return:
new_xyz: sampled points position data, [B, 1, C]
new_points: sampled points data, [B, 1, N, C+D]
"""
B, N, C = xyz.shape
S = npoint
fps_idx = farthest_point_sample_conv(xyz, npoint) # [B, npoint, C]
new_xyz = index_points_conv(xyz, fps_idx)
idx = knn_point_conv(nsample, xyz, new_xyz)
grouped_xyz = index_points_conv(xyz, idx) # [B, npoint, nsample, C]
grouped_xyz_norm = grouped_xyz - new_xyz.view(B, S, 1, C)
if points is not None:
grouped_points = index_points_conv(points, idx)
# print(grouped_points.shape)
# print("Debug grouped points: {}".format(grouped_points[0,0,0,0:5]))
new_points = torch.cat([grouped_xyz_norm, grouped_points], dim=-1) # [B, npoint, nsample, C+D]
else:
new_points = grouped_xyz_norm
if density_scale is None:
return new_xyz, new_points, grouped_xyz_norm, idx
else:
grouped_density = index_points_conv(density_scale, idx)
return new_xyz, new_points, grouped_xyz_norm, idx, grouped_density
def sample_and_group_all_conv(xyz, points, density_scale = None):
"""
Input:
xyz: input points position data, [B, N, C]
points: input points data, [B, N, D]
Return:
new_xyz: sampled points position data, [B, 1, C]
new_points: sampled points data, [B, 1, N, C+D]
"""
device = xyz.device
B, N, C = xyz.shape
#new_xyz = torch.zeros(B, 1, C).to(device)
new_xyz = xyz.mean(dim = 1, keepdim = True)
grouped_xyz = xyz.view(B, 1, N, C) - new_xyz.view(B, 1, 1, C)
if points is not None:
new_points = torch.cat([grouped_xyz, points.view(B, 1, N, -1)], dim=-1)
else:
new_points = grouped_xyz
if density_scale is None:
return new_xyz, new_points, grouped_xyz
else:
grouped_density = density_scale.view(B, 1, N, 1)
return new_xyz, new_points, grouped_xyz, grouped_density
def group_conv(nsample, xyz, points):
"""
Input:
npoint:
nsample:
xyz: input points position data, [B, N, C]
points: input points data, [B, N, D]
Return:
new_xyz: sampled points position data, [B, 1, C]
new_points: sampled points data, [B, 1, N, C+D]
"""
B, N, C = xyz.shape
S = N
new_xyz = xyz
idx = knn_point_conv(nsample, xyz, new_xyz)
grouped_xyz = index_points_conv(xyz, idx) # [B, npoint, nsample, C]
grouped_xyz_norm = grouped_xyz - new_xyz.view(B, S, 1, C)
if points is not None:
grouped_points = index_points_conv(points, idx)
new_points = torch.cat([grouped_xyz_norm, grouped_points], dim=-1) # [B, npoint, nsample, C+D]
else:
new_points = grouped_xyz_norm
return new_points, grouped_xyz_norm
# def compute_density(xyz, bandwidth):
# '''
# xyz: input points position data, [B, N, C]
# '''
# #import ipdb; ipdb.set_trace()
# B, N, C = xyz.shape
# sqrdists = square_distance_conv(xyz, xyz)
# gaussion_density = torch.exp(- sqrdists / (2.0 * bandwidth * bandwidth)) / (2.5 * bandwidth)
# xyz_density = gaussion_density.mean(dim = -1)
# return xyz_density
# +
def compute_density(xyz, bandwidth):
'''
xyz: input points position data, [B, N, C]
'''
#import ipdb; ipdb.set_trace()
B, N, C = xyz.shape
sqrdists = square_distance(xyz, xyz)
gaussion_density = torch.exp(- sqrdists / (2.0 * bandwidth * bandwidth)) / (2.5 * bandwidth)
xyz_density = gaussion_density.mean(dim = -1)
return xyz_density
class DensityNet(nn.Module):
def __init__(self, hidden_unit = [16, 8]):
super(DensityNet, self).__init__()
self.mlp_convs = nn.ModuleList()
self.mlp_bns = nn.ModuleList()
self.mlp_convs.append(nn.Conv2d(1, hidden_unit[0], 1))
self.mlp_bns.append(nn.BatchNorm2d(hidden_unit[0]))
for i in range(1, len(hidden_unit)):
self.mlp_convs.append(nn.Conv2d(hidden_unit[i - 1], hidden_unit[i], 1))
self.mlp_bns.append(nn.BatchNorm2d(hidden_unit[i]))
self.mlp_convs.append(nn.Conv2d(hidden_unit[-1], 1, 1))
self.mlp_bns.append(nn.BatchNorm2d(1))
def forward(self, density_scale):
for i, conv in enumerate(self.mlp_convs):
bn = self.mlp_bns[i]
density_scale = bn(conv(density_scale))
if i == len(self.mlp_convs):
density_scale = F.sigmoid(density_scale)
else:
density_scale = F.relu(density_scale)
return density_scale
class WeightNet(nn.Module):
def __init__(self, in_channel, out_channel, hidden_unit = [8, 8]):
super(WeightNet, self).__init__()
self.mlp_convs = nn.ModuleList()
self.mlp_bns = nn.ModuleList()
if hidden_unit is None or len(hidden_unit) == 0:
self.mlp_convs.append(nn.Conv2d(in_channel, out_channel, 1))
self.mlp_bns.append(nn.BatchNorm2d(out_channel))
else:
self.mlp_convs.append(nn.Conv2d(in_channel, hidden_unit[0], 1))
self.mlp_bns.append(nn.BatchNorm2d(hidden_unit[0]))
for i in range(1, len(hidden_unit)):
self.mlp_convs.append(nn.Conv2d(hidden_unit[i - 1], hidden_unit[i], 1))
self.mlp_bns.append(nn.BatchNorm2d(hidden_unit[i]))
self.mlp_convs.append(nn.Conv2d(hidden_unit[-1], out_channel, 1))
self.mlp_bns.append(nn.BatchNorm2d(out_channel))
def forward(self, localized_xyz):
#xyz : BxCxKxN
weights = localized_xyz
for i, conv in enumerate(self.mlp_convs):
bn = self.mlp_bns[i]
weights = F.relu(bn(conv(weights)))
return weights
class PointNetSetAbstraction(nn.Module):
# npoint, nsample, in_channel, mlp, bandwidth, group_all
def __init__(self, npoint, nsample, in_channel, mlp, bandwidth,group_all):
super(PointNetSetAbstraction, self).__init__()
self.npoint = npoint
# self.radius = radius
self.bandwidth = bandwidth
self.nsample = nsample
self.mlp_convs = nn.ModuleList()
self.mlp_bns = nn.ModuleList()
last_channel = in_channel
for out_channel in mlp:
self.mlp_convs.append(nn.Conv2d(last_channel, out_channel, 1))
self.mlp_bns.append(nn.BatchNorm2d(out_channel))
last_channel = out_channel
self.group_all = group_all
self.weightnet = WeightNet(3, 16)
self.linear = nn.Linear(16 * mlp[-1], mlp[-1])
self.bn_linear = nn.BatchNorm1d(mlp[-1])
self.densitynet = DensityNet()
def forward(self, xyz, points):
"""
Input:
xyz: input points position data, [B, C, N]
points: input points data, [B, D, N]
Return:
new_xyz: sampled points position data, [B, C, S]
new_points_concat: sample points feature data, [B, D', S]
"""
print("SA")
B = xyz.shape[0]
N = xyz.shape[2]
xyz = xyz.permute(0, 2, 1)
if points is not None:
points = points.permute(0, 2, 1)
xyz_density = compute_density(xyz, self.bandwidth)
inverse_density = 1.0 / xyz_density
if self.group_all:
new_xyz, new_points, grouped_xyz_norm, grouped_density = sample_and_group_all_conv(xyz,
points,
inverse_density.view(B, N, 1))
else:
new_xyz, new_points, grouped_xyz_norm, _, grouped_density = sample_and_group_conv(self.npoint,
self.nsample,
xyz,
points,
inverse_density.view(B, N, 1))
new_points = new_points.permute(0, 3, 2, 1) # [B, C+D, nsample,npoint]
print("----new_points_1:", new_points.shape)
for i, conv in enumerate(self.mlp_convs):
bn = self.mlp_bns[i]
new_points = F.relu(bn(conv(new_points)))
print("----new_points_none_linear_1:", new_points.shape)
inverse_max_density = grouped_density.max(dim = 2, keepdim=True)[0]
density_scale = grouped_density / inverse_max_density
density_scale = self.densitynet(density_scale.permute(0, 3, 2, 1))
new_points = new_points * density_scale
grouped_xyz = grouped_xyz_norm.permute(0, 3, 2, 1)
weights = self.weightnet(grouped_xyz)
new_points = torch.matmul(input=new_points.permute(0, 3, 1, 2), other = weights.permute(0, 3, 2, 1)).view(B, self.npoint, -1)
new_points = self.linear(new_points)
print("----new_points_none_linear_2:",new_points.shape)
new_points = self.bn_linear(new_points.permute(0, 2, 1))
new_points = F.relu(new_points)
new_xyz = new_xyz.permute(0, 2, 1)
#add new noneline
print("----new_points_out:",new_points.shape)
return new_xyz, new_points
class PointNetFeaturePropagation(nn.Module):
def __init__(self, npoint, nsample, in_channel, out_put, mlp, bandwidth,group_all):
super(PointNetFeaturePropagation, self).__init__()
self.npoint = npoint
self.nsample = nsample
self.mlp_convs = nn.ModuleList()
self.mlp_bns = nn.ModuleList()
self.mlp_convs2d = nn.ModuleList()
self.mlp_bns2d = nn.ModuleList()
self.out_put = out_put
last_channel = in_channel
for out_channel in mlp:
self.mlp_convs2d.append(nn.Conv2d(last_channel, out_channel, 1))
self.mlp_bns2d.append(nn.BatchNorm2d(out_channel))
last_channel = out_channel
last_channel = out_put
for out_channel in mlp:
self.mlp_convs.append(nn.Conv2d(last_channel, out_channel, 1))
self.mlp_bns.append(nn.BatchNorm2d(out_channel))
last_channel = out_channel
self.bandwidth = bandwidth
self.group_all = group_all
self.weightnet = WeightNet(3, 16)
self.linear = nn.Linear(16 * mlp[-1], mlp[-1])
self.bn_linear = nn.BatchNorm1d(mlp[-1])
self.densitynet = DensityNet()
def forward(self, xyz1, xyz2, points1, points2):
"""
Input:
xyz1: input points position data, [B, C, N]
xyz2: sampled input points position data, [B, C, S]
points1: input points data, [B, D, N]
points2: input points data, [B, D, S]
Return:
new_points: upsampled points data, [B, D', N]
"""
print("FP")
xyz1 = xyz1.permute(0, 2, 1)
xyz2 = xyz2.permute(0, 2, 1)
points2 = points2.permute(0, 2, 1)
B, N, C = xyz1.shape
_, S, _ = xyz2.shape
if S == 1:
interpolated_points = points2.repeat(1, N, 1)
else:
dists = square_distance(xyz1, xyz2)
dists, idx = dists.sort(dim=-1)
dists, idx = dists[:, :, :3], idx[:, :, :3] # [B, N, 3]
dist_recip = 1.0 / (dists + 1e-8)
norm = torch.sum(dist_recip, dim=2, keepdim=True)
weight = dist_recip / norm
interpolated_points = torch.sum(index_points(points2, idx) * weight.view(B, N, 3, 1), dim=2)
print("----interpolated_points:", interpolated_points.shape)
#deconv
xyz_density = compute_density(xyz1, self.bandwidth)
inverse_density = 1.0 / xyz_density
if self.group_all:
new_xyz, new_points, grouped_xyz_norm, grouped_density = sample_and_group_all_conv(xyz1,
interpolated_points,
inverse_density.view(B, N, 1))
else:
new_xyz, new_points, grouped_xyz_norm, _, grouped_density = sample_and_group_conv(self.npoint,
self.nsample,
xyz1,
interpolated_points,
inverse_density.view(B, N, 1))
print("----new_points:", new_points.shape)
new_points = new_points.permute(0, 3, 2, 1) # [B, C+D, nsample,npoint]
print("----new_points_permute_1:", new_points.shape)
for i, conv in enumerate(self.mlp_convs2d):
# print("i:",i)
bn = self.mlp_bns2d[i]
new_points = F.relu(bn(conv(new_points)))
# print("-----new_points:",new_points.shape)
print("----new_points_none_linear:", new_points.shape)
########weight-net#############3
grouped_xyz = grouped_xyz_norm.permute(0, 3, 2, 1)
weights = self.weightnet(grouped_xyz)
print("----weights:", weights.shape)
################################
################################
if points1 is not None:
print("points1:", points1.shape)
points1 = points1.permute(0, 2, 1)
new_points = torch.cat([points1, interpolated_points], dim=-1)
else:
new_points = interpolated_points
new_points = new_points.permute(0, 2, 1)
print("new_points_permute_FP:",new_points.shape)
new_points.unsqueeze_(2)
new_points = new_points.permute(0, 1, 2, 3)
for i, conv in enumerate(self.mlp_convs):
bn = self.mlp_bns[i]
new_points = F.relu(bn(conv(new_points)))
new_points = torch.squeeze(new_points,dim=2)
print("new_points_FP:",new_points.shape)
return new_points
# -
for i, data in tqdm(enumerate(train_data_loader), total=len(train_data_loader), smoothing=0.9, disable=True):
points, target,w = data
points = points.transpose(2, 1)
points.shape
l0_points = points
l0_xyz = points[:,:3,:]
sa1 = PointNetSetAbstraction(1024, 32, 6 + 3, [32, 32, 64],0.2, False)
sa2 = PointNetSetAbstraction(256, 32, 64 + 3, [64, 64, 128],0.2, False)
sa3 = PointNetSetAbstraction(64, 32, 128 + 3, [128, 128, 256],0.2, False)
sa4 = PointNetSetAbstraction(16, 32, 256 + 3, [256, 256, 512],0.2, False)
fp4 = PointNetFeaturePropagation(512, 32, 515, 768, [512, 256], bandwidth =0.1,group_all=False)
fp3 = PointNetFeaturePropagation(256, 32, 259, 384, [256, 256], bandwidth =0.2,group_all=False)
fp2 = PointNetFeaturePropagation(256, 32, 259, 320, [256, 128], bandwidth =0.4,group_all=False)
fp1 = PointNetFeaturePropagation(128, 32, 131, 128, [128, 128, 128], bandwidth =0.6,group_all=False)
l1_xyz, l1_points = sa1(l0_xyz, l0_points)
l2_xyz, l2_points = sa2(l1_xyz, l1_points)
l3_xyz, l3_points = sa3(l2_xyz, l2_points)
l4_xyz, l4_points = sa4(l3_xyz, l3_points)
l3_points = fp4(l3_xyz, l4_xyz, l3_points, l4_points)
l2_points = fp3(l2_xyz, l3_xyz, l2_points, l3_points)
l1_points = fp2(l1_xyz, l2_xyz, l1_points, l2_points)
l0_points = fp1(l0_xyz, l1_xyz, None, l1_points)
l0_points.shape
num_classes=21
NUM_CLASSES=21
conv1 = nn.Conv1d(128, 128, 1)
bn1 = nn.BatchNorm1d(128)
drop1 = nn.Dropout(0.5)
conv2 = nn.Conv1d(128, num_classes, 1)
x = drop1(F.relu(bn1(conv1(l0_points))))
x = conv2(x)
x = F.log_softmax(x, dim=1)
x = x.permute(0, 2, 1)
# return x, l4_points
x.shape
seg_pred=x
trans_feat = l4_points
weights = torch.Tensor(TRAIN_DATASET.labelweights).cuda()
weights.shape
class get_loss(nn.Module):
def __init__(self):
super(get_loss, self).__init__()
def forward(self, pred, target, trans_feat, weight):
total_loss = F.nll_loss(pred, target, weight=weight)
return total_loss
criterion = get_loss().cuda()
# seg_pred, trans_feat = classifier(points)
seg_pred = seg_pred.contiguous().view(-1, NUM_CLASSES)
batch_label = target.view(-1, 1)[:, 0].cpu().data.numpy()
target = target.view(-1, 1)[:, 0]
seg_pred.shape
batch_label.shape
target.shape
loss = criterion(seg_pred, target, trans_feat, weights)
# +
class PointNet2(nn.Module):
def __init__(self, npoint, radius, nsample, in_channel, mlp, group_all):
super(PointNet2, self).__init__()
self.npoint = npoint
self.radius = radius
self.nsample = nsample
self.mlp_convs = nn.ModuleList()
self.mlp_bns = nn.ModuleList()
last_channel = in_channel
for out_channel in mlp:
self.mlp_convs.append(nn.Conv2d(last_channel, out_channel, 1))
self.mlp_bns.append(nn.BatchNorm2d(out_channel))
last_channel = out_channel
self.group_all = group_all
def forward(self, xyz, points):
"""
Input:
xyz: input points position data, [B, C, N]
points: input points data, [B, D, N]
Return:
new_xyz: sampled points position data, [B, C, S]
new_points_concat: sample points feature data, [B, D', S]
"""
print("points:",points.shape)
xyz = xyz.permute(0, 2, 1)
if points is not None:
points = points.permute(0, 2, 1)
if self.group_all:
new_xyz, new_points = sample_and_group_all(xyz, points)
else:
new_xyz, new_points = sample_and_group(self.npoint, self.radius, self.nsample, xyz, points)
print("new_points_grouping:",new_points.shape)
# new_xyz: sampled points position data, [B, npoint, C]
# new_points: sampled points data, [B, npoint, nsample, C+D]
new_points = new_points.permute(0, 3, 2, 1) # [B, C+D, nsample,npoint]
for i, conv in enumerate(self.mlp_convs):
bn = self.mlp_bns[i]
new_points = F.relu(bn(conv(new_points)))
new_points = torch.max(new_points, 2)[0]
new_xyz = new_xyz.permute(0, 2, 1)
print("new_points_out:",new_points.shape)
return new_xyz, new_points
class PointNet2FP(nn.Module):
def __init__(self, in_channel, mlp):
super(PointNet2FP, self).__init__()
self.mlp_convs = nn.ModuleList()
self.mlp_bns = nn.ModuleList()
last_channel = in_channel
for out_channel in mlp:
self.mlp_convs.append(nn.Conv1d(last_channel, out_channel, 1))
self.mlp_bns.append(nn.BatchNorm1d(out_channel))
last_channel = out_channel
def forward(self, xyz1, xyz2, points1, points2):
"""
Input:
xyz1: input points position data, [B, C, N]
xyz2: sampled input points position data, [B, C, S]
points1: input points data, [B, D, N]
points2: input points data, [B, D, S]
Return:
new_points: upsampled points data, [B, D', N]
"""
xyz1 = xyz1.permute(0, 2, 1)
xyz2 = xyz2.permute(0, 2, 1)
points2 = points2.permute(0, 2, 1)
B, N, C = xyz1.shape
_, S, _ = xyz2.shape
if S == 1:
interpolated_points = points2.repeat(1, N, 1)
else:
dists = square_distance(xyz1, xyz2)
dists, idx = dists.sort(dim=-1)
dists, idx = dists[:, :, :3], idx[:, :, :3] # [B, N, 3]
dist_recip = 1.0 / (dists + 1e-8)
norm = torch.sum(dist_recip, dim=2, keepdim=True)
weight = dist_recip / norm
interpolated_points = torch.sum(index_points(points2, idx) * weight.view(B, N, 3, 1), dim=2)
if points1 is not None:
points1 = points1.permute(0, 2, 1)
new_points = torch.cat([points1, interpolated_points], dim=-1)
else:
new_points = interpolated_points
print("new_points_FP:",new_points.shape)
new_points = new_points.permute(0, 2, 1)
for i, conv in enumerate(self.mlp_convs):
bn = self.mlp_bns[i]
new_points = F.relu(bn(conv(new_points)))
print("new_points_FP:",new_points.shape)
return new_points
# -
l0_points = points
l0_xyz = points[:,:3,:]
sa1 = PointNet2(1024, 0.1, 32, 6 + 3, [32, 32, 64], False)
sa2 = PointNet2(256, 0.2, 32, 64 + 3, [64, 64, 128], False)
sa3 = PointNet2(64, 0.4, 32, 128 + 3, [128, 128, 256], False)
sa4 = PointNet2(16, 0.8, 32, 256 + 3, [256, 256, 512], False)
fp4 = PointNet2FP(768, [256, 256])
fp3 = PointNet2FP(384, [256, 256])
fp2 = PointNet2FP(320, [256, 128])
fp1 = PointNet2FP(128, [128, 128, 128])
conv1 = nn.Conv1d(128, 128, 1)
bn1 = nn.BatchNorm1d(128)
drop1 = nn.Dropout(0.5)
conv2 = nn.Conv1d(128, num_classes, 1)
# +
l1_xyz, l1_points = sa1(l0_xyz, l0_points)
l2_xyz, l2_points = sa2(l1_xyz, l1_points)
l3_xyz, l3_points = sa3(l2_xyz, l2_points)
l4_xyz, l4_points = sa4(l3_xyz, l3_points)
l3_points = fp4(l3_xyz, l4_xyz, l3_points, l4_points)
l2_points = fp3(l2_xyz, l3_xyz, l2_points, l3_points)
l1_points = fp2(l1_xyz, l2_xyz, l1_points, l2_points)
l0_points = fp1(l0_xyz, l1_xyz, None, l1_points)
# -
>>> x = torch.zeros(2, 1, 2, 1, 2)
>>> x.size()
>>> y = torch.squeeze(x,dim=1)
y.shape
| pointconv_segmentation_pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Manim in Jupyter
# Working with manim in jupyter notebooks has several advantages:
#
# * code snippets and rendered outputs are close together
# * easy to iterate examples
# * easy to try different varieties of one scene in multiple cells
# * computation intensive code can be executed separately from the scenes
# * global Mobjects can be used in multiple scenes.
# ## Simple Example
# First, we need to import manim
from manim import *
# Now we build up our scene
# %%manim -v WARNING --progress_bar None -s -ql --disable_caching MyExample
class MyExample(Scene):
def construct(self):
m= ManimBanner()
self.add(m)
# Note, that I use the following parameters:
#
# * `-v WARNING` means that only warnings are shown in the log
# * `--progress_bar None` will not show the animation progress bar
# * `-s ` will only show the last frame
# * `-ql` renders in low quality
# * `--disable_caching` will disable the manim caching system
# * `MyExample` gives the scene name
#
# for rendering a video, just remove the -s flag. To lower the resolution, you can use -r 400,200 (pixel values in x and y direction).
# %%manim -v WARNING --progress_bar None -r 400,200 --disable_caching HelloManim
class HelloManim(Scene):
def construct(self):
self.camera.background_color = "#ece6e2"
banner_large = ManimBanner(dark_theme=False).scale(0.7)
self.play(banner_large.create())
self.play(banner_large.expand())
# Rendering with a transparent background can be done like this:
# %%manim -v ERROR --progress_bar None -r 400,200 -t --disable_caching HelloManim
class HelloManim(Scene):
def construct(self):
self.camera.background_color = "#ece6e2"
banner_large = ManimBanner(dark_theme=False).scale(1.5)
self.add(banner_large)
# We can define the parameters as a string `param` and call this string by the cell magic with `$param`
param = "-v WARNING -s -ql --disable_caching Example"
paramSMALL = "-v WARNING -r 400,200 -s --disable_caching Example"
# %%manim $param
class Example(Scene):
def construct(self):
m= ManimBanner()
self.add(m)
# ## Initializing Mobjects Outside the Class
# In some cases, it might be convenient to define mobjects outside the `Scene` class (e.g. for uncluttering or for speeding up the animation).
m = ManimBanner()
# %%manim $paramSMALL
class Example(Scene):
def construct(self):
m.scale(0.4 )
m.shift(1.5*UP)
self.add(m)
# Because the mobject is manipulated in the class, the next cell might show some unexpected scaling and shifting:
# %%manim $paramSMALL
class Example(Scene):
def construct(self):
m.scale(0.4)
m.shift(1.5*UP)
self.add(m)
# To aviod this, it is better to add only a copy of these mobjects to scenes, and keep the originals untouched:
m_reference = ManimBanner()
# %%manim $paramSMALL
class Example(Scene):
def construct(self):
m = m_reference.copy()
m.scale(0.4)
m.shift(2*UP)
self.add(m)
# %%manim $paramSMALL
class Example(Scene):
def construct(self):
m = m_reference.copy()
m.scale(0.4)
m.shift(2*UP)
self.add(m)
# ## Defining Global Mobjects
# When you have to build complex scenes, you might want to use parts of that scene for your next scene. That is possible with global variables, which can be accessed in any other scene.
# %%manim $paramSMALL
class Example(Scene):
def construct(self):
stars= VGroup()
for i in range(0,20):
s= Star(color= random_bright_color(), fill_opacity=1).scale(0.8)
stars.add(s)
stars.arrange_in_grid()
self.add(stars)
global favoritstar
favoritstar = stars[9]
# %%manim $paramSMALL
class Example(Scene):
def construct(self):
self.add(favoritstar)
# ## Pre-Execute Slow Code
# In this example, calculating a random walk for 500 particles and 100000 steps takes about 4 seconds.
# This step can be done before the actual scene construction, which takes about 0.2 seconds.
# Making aesthetic changes to the scene will then become easier.
# Note: The `%%time` command will print the execution time of the cells.
# %%time
np.random.seed(20)
steps = np.random.choice(a=[-1, 0, 1], size=(100000,1000))
stop = steps.cumsum(0)
end_points= stop[-1]/stop[-1].max()
end_pointsX = end_points[0:499]
end_pointsY = end_points[500:-1]
# %%time
# %%manim $param
class Example(Scene):
def construct(self):
radius= (end_pointsX*end_pointsX + end_pointsY * end_pointsY)**0.5
dots = VGroup()
for x,y,r in zip(end_pointsX, end_pointsY,radius):
c= interpolate_color(YELLOW, RED, r)
dots.add(Dot(color=c,point=[3*x,3*y,0]).scale(0.7))
self.add(dots)
# ## Installing Plugins
# plugins can be found at https://plugins.manim.community/
# !pip install manim-physics
# +
# %%manim -v WARNING --progress_bar None -qm --disable_caching Example
from manim_physics import *
class Example(SpaceScene):
def construct(self):
circle = Dot(radius=1).shift(1.5*LEFT+3*UP)
rect = Square(color=YELLOW, fill_opacity=1)
ground = Line([-4, -3.5, 0], [4, -3.5, 0])
wall1 = Line([-4, -3.5, 0], [-4, 3.5, 0])
wall2 = Line([4, -3.5, 0], [4, 3.5, 0])
walls = VGroup(ground, wall1, wall2)
self.add(walls)
self.add(rect, circle)
self.make_rigid_body(rect, circle)
self.make_static_body(walls)
self.wait(5)
| docs/source/ch1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Front matter
import os,datetime
import pandas as pd
import numpy as np
import scipy
from scipy import constants
import matplotlib
import matplotlib.pyplot as plt
from matplotlib.ticker import AutoMinorLocator
from matplotlib import gridspec
import re
# Seaborn, useful for graphics
import seaborn as sns
# Magic function to make matplotlib inline; other style specs must come AFTER
# %matplotlib inline
matplotlib.rc('xtick', labelsize=16)
matplotlib.rc('ytick', labelsize=16)
rc = {'lines.linewidth': 1,
'axes.labelsize': 20,
'axes.titlesize': 20,
'legend.fontsize': 26,
'xtick.direction': u'in',
'ytick.direction': u'in'}
sns.set_style('ticks', rc=rc)
# +
# Import data sets
studylist = []
labelchoice = dict()
phase = dict()
NRIXSdatapath = dict()
XRDdatapath = dict()
colorchoice = dict()
symbol = dict()
symbolsize = dict()
study = 'bccFe'
studylist.append(study)
labelchoice[study] = 'bcc Fe (this study)'
phase[study] = 'bcc'
NRIXSdatapath[study] = '../050_phox_Fe_man/Results/phox_valsFromPDOS.csv'
XRDdatapath[study] = '../010_XRDAnalysis/Results/XRD_results_bccFe.csv'
colorchoice[study] = 'Gray'
symbol[study] = '^'
symbolsize[study] = 9
study = 'bccFeNi'
studylist.append(study)
labelchoice[study] = 'bcc Fe$_{0.91}$Ni$_{0.09}$ (this study)'
phase[study] = 'bcc'
NRIXSdatapath[study] = '../060_phox_FeNi_man/Results/phox_valsFromPDOS.csv'
XRDdatapath[study] = '../010_XRDAnalysis/Results/XRD_results_bccFeNi.csv'
colorchoice[study] = 'DarkOrange'
symbol[study] = '^'
symbolsize[study] = 9
study = 'bccFeNiSi'
studylist.append(study)
labelchoice[study] = 'hcp Fe$_{0.80}$Ni$_{0.10}$Si$_{0.10}$ (this study)'
phase[study] = 'bcc'
NRIXSdatapath[study] = '../070_phox_FeNiSi_man/Results/phox_valsFromPDOS.csv'
XRDdatapath[study] = '../010_XRDAnalysis/Results/XRD_results_bccFeNiSi.csv'
colorchoice[study] = 'DeepSkyBlue'
symbol[study] = '^'
symbolsize[study] = 9
study = 'hcpFe'
studylist.append(study)
labelchoice[study] = 'hcp Fe (Murphy et al.)'
phase[study] = 'hcp'
NRIXSdatapath[study] = '../005_PubNRIXSVals/hcpFe_Murphy.csv'
XRDdatapath[study] = '../005_PubNRIXSVals/hcpFe_Murphy.csv'
colorchoice[study] = 'Gray'
symbol[study] = 'o'
symbolsize[study] = 8
study = 'hcpFeNi'
studylist.append(study)
labelchoice[study] = 'hcp Fe$_{0.91}$Ni$_{0.09}$ (this study)'
phase[study] = 'hcp'
NRIXSdatapath[study] = '../060_phox_FeNi_man/Results/phox_valsFromPDOS.csv'
XRDdatapath[study] = '../010_XRDAnalysis/Results/XRD_results_hcpFeNi.csv'
colorchoice[study] = 'DarkOrange'
symbol[study] = 'o'
symbolsize[study] = 8
study = 'hcpFeNiSi'
studylist.append(study)
labelchoice[study] = 'hcp Fe$_{0.80}$Ni$_{0.10}$Si$_{0.10}$ (this study)'
phase[study] = 'hcp'
NRIXSdatapath[study] = '../070_phox_FeNiSi_man/Results/phox_valsFromPDOS.csv'
XRDdatapath[study] = '../010_XRDAnalysis/Results/XRD_results_hcpFeNiSi.csv'
colorchoice[study] = 'DeepSkyBlue'
symbol[study] = 'o'
symbolsize[study] = 8
# +
precious_dfdict = dict()
for study in studylist:
print('Now importing '+study)
# Import XRD data
init_XRD_df = pd.read_csv(XRDdatapath[study])
# Change index name to match other data sets
if 'NRIXS exp' in init_XRD_df:
init_XRD_df = init_XRD_df.rename(columns={'NRIXS exp': 'Index'})
# We only need some of the info here
if phase[study] == 'bcc':
XRD_df = init_XRD_df[['Index','a','da','V','dV','rho','drho','P','dP']]
else:
XRD_df = init_XRD_df[['Index','a','da','c','dc','V','dV','rho','drho','P','dP']]
# Import NRIXS data
init_NRIXS_df = pd.read_csv(NRIXSdatapath[study])
# We're only going to look at some of the values here, b/c only some values are reported in Murphy et al.
NRIXS_df = init_NRIXS_df[['Index','fLM','dfLM','KE','dKE','MFC','dMFC','cvib','dcvib','Svib','dSvib','TLM','dTLM','IF','dIF','Fvib','dFvib']]
# Get rows in NRIXS df that also appear in XRD df. (So we only look at bcc or hcp data.)
NRIXS_df = NRIXS_df[NRIXS_df['Index'].isin(XRD_df['Index'].values)]
# Combine XRD df and NRIXS df into one df to rule them all
precious_df = pd.merge(XRD_df,NRIXS_df,on='Index')
precious_dfdict[study] = precious_df
# -
alphaval = 0.6
# +
# Lamb-Moessbauer factor figure
fig, (ax0) = plt.subplots(nrows = 1, ncols=1, sharex=True, figsize=(8, 8))
# Plot bcc Fe
#####################
study = 'bccFe'
df = precious_dfdict[study]
h0, = ax0.plot(df['P'], df['fLM'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['fLM'], xerr=df['dP'], yerr=df['dfLM'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot bcc FeNi
#####################
study = 'bccFeNi'
df = precious_dfdict[study]
h1, = ax0.plot(df['P'], df['fLM'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['fLM'], xerr=df['dP'], yerr=df['dfLM'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot bcc FeNiSi
#####################
study = 'bccFeNiSi'
df = precious_dfdict[study]
h2, = ax0.plot(df['P'], df['fLM'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['fLM'], xerr=df['dP'], yerr=df['dfLM'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot hcp Fe
#####################
study = 'hcpFe'
df = precious_dfdict[study]
h3, = ax0.plot(df['P'], df['fLM'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['fLM'], xerr=df['dP'], yerr=df['dfLM'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot hcp FeNi
#####################
study = 'hcpFeNi'
df = precious_dfdict[study]
h4, = ax0.plot(df['P'], df['fLM'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['fLM'], xerr=df['dP'], yerr=df['dfLM'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot hcp FeNiSi
#####################
study = 'hcpFeNiSi'
df = precious_dfdict[study]
h5, = ax0.plot(df['P'], df['fLM'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['fLM'], xerr=df['dP'], yerr=df['dfLM'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
hlabels = [h0,h1,h2,h3,h4,h5]
ax0.legend(fontsize=14,loc=4,handles = hlabels)
ax0.set_xlabel(r'Pressure (GPa)', fontsize=18)
ax0.set_ylabel(r'$f_{LM}$', fontsize=18)
plt.tight_layout()
fig = plt.gcf()
fig.savefig('hcpFeAlloy_fLM.pdf', format='pdf')
# +
# Mean force constant figure
fig, (ax0) = plt.subplots(nrows = 1, ncols=1, sharex=True, figsize=(8, 8))
# Plot bcc Fe
#####################
study = 'bccFe'
df = precious_dfdict[study]
h0, = ax0.plot(df['P'], df['MFC'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['MFC'], xerr=df['dP'], yerr=df['dMFC'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot bcc FeNi
#####################
study = 'bccFeNi'
df = precious_dfdict[study]
h1, = ax0.plot(df['P'], df['MFC'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['MFC'], xerr=df['dP'], yerr=df['dMFC'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot bcc FeNiSi
#####################
study = 'bccFeNiSi'
df = precious_dfdict[study]
h2, = ax0.plot(df['P'], df['MFC'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['MFC'], xerr=df['dP'], yerr=df['dMFC'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot hcp Fe
#####################
study = 'hcpFe'
df = precious_dfdict[study]
h3, = ax0.plot(df['P'], df['MFC'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['MFC'], xerr=df['dP'], yerr=df['dMFC'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot hcp FeNi
#####################
study = 'hcpFeNi'
df = precious_dfdict[study]
h4, = ax0.plot(df['P'], df['MFC'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['MFC'], xerr=df['dP'], yerr=df['dMFC'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot hcp FeNiSi
#####################
study = 'hcpFeNiSi'
df = precious_dfdict[study]
h5, = ax0.plot(df['P'], df['MFC'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['MFC'], xerr=df['dP'], yerr=df['dMFC'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
hlabels = [h0,h1,h2,h3,h4,h5]
ax0.legend(fontsize=14,loc=4,handles = hlabels)
ax0.set_xlabel(r'Pressure (GPa)', fontsize=18)
ax0.set_ylabel(r'$MFC$', fontsize=18)
plt.tight_layout()
fig = plt.gcf()
fig.savefig('hcpFeAlloy_MFC.pdf', format='pdf')
# +
# Vibrational entropy figure
fig, (ax0) = plt.subplots(nrows = 1, ncols=1, sharex=True, figsize=(8, 8))
# Plot bcc Fe
#####################
study = 'bccFe'
df = precious_dfdict[study]
h0, = ax0.plot(df['P'], df['Svib'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['Svib'], xerr=df['dP'], yerr=df['dSvib'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot bcc FeNi
#####################
study = 'bccFeNi'
df = precious_dfdict[study]
h1, = ax0.plot(df['P'], df['Svib'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['Svib'], xerr=df['dP'], yerr=df['dSvib'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot bcc FeNiSi
#####################
study = 'bccFeNiSi'
df = precious_dfdict[study]
h2, = ax0.plot(df['P'], df['Svib'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['Svib'], xerr=df['dP'], yerr=df['dSvib'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot hcp Fe
#####################
study = 'hcpFe'
df = precious_dfdict[study]
h3, = ax0.plot(df['P'], df['Svib'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['Svib'], xerr=df['dP'], yerr=df['dSvib'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot hcp FeNi
#####################
study = 'hcpFeNi'
df = precious_dfdict[study]
h4, = ax0.plot(df['P'], df['Svib'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['Svib'], xerr=df['dP'], yerr=df['dSvib'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot hcp FeNiSi
#####################
study = 'hcpFeNiSi'
df = precious_dfdict[study]
h5, = ax0.plot(df['P'], df['Svib'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['Svib'], xerr=df['dP'], yerr=df['dSvib'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
hlabels = [h0,h1,h2,h3,h4,h5]
ax0.legend(fontsize=14,loc=1,handles = hlabels)
ax0.set_xlabel(r'Pressure (GPa)', fontsize=18)
ax0.set_ylabel(r'$S_{vib}$', fontsize=18)
plt.tight_layout()
fig = plt.gcf()
fig.savefig('hcpFeAlloy_Svib.pdf', format='pdf')
# +
# Vibrational free energy figure
fig, (ax0) = plt.subplots(nrows = 1, ncols=1, sharex=True, figsize=(8, 8))
# Plot bcc Fe
#####################
study = 'bccFe'
df = precious_dfdict[study]
h0, = ax0.plot(df['P'], df['Fvib'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['Fvib'], xerr=df['dP'], yerr=df['dFvib'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot bcc FeNi
#####################
study = 'bccFeNi'
df = precious_dfdict[study]
h1, = ax0.plot(df['P'], df['Fvib'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['Fvib'], xerr=df['dP'], yerr=df['dFvib'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot bcc FeNiSi
#####################
study = 'bccFeNiSi'
df = precious_dfdict[study]
h2, = ax0.plot(df['P'], df['Fvib'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['Fvib'], xerr=df['dP'], yerr=df['dFvib'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot hcp Fe
#####################
study = 'hcpFe'
df = precious_dfdict[study]
h3, = ax0.plot(df['P'], df['Fvib'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['Fvib'], xerr=df['dP'], yerr=df['dFvib'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot hcp FeNi
#####################
study = 'hcpFeNi'
df = precious_dfdict[study]
h4, = ax0.plot(df['P'], df['Fvib'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['Fvib'], xerr=df['dP'], yerr=df['dFvib'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
# Plot hcp FeNiSi
#####################
study = 'hcpFeNiSi'
df = precious_dfdict[study]
h5, = ax0.plot(df['P'], df['Fvib'], marker=symbol[study], ms=symbolsize[study], color=colorchoice[study],
mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none')
ax0.errorbar(df['P'], df['Fvib'], xerr=df['dP'], yerr=df['dFvib'],
marker=symbol[study], ms=symbolsize[study], capsize=0, color=colorchoice[study], mfc=colorchoice[study],
mec=matplotlib.colors.colorConverter.to_rgba('White', alpha=.75),
label=labelchoice[study],markeredgewidth=0.5,ls='none',elinewidth=1)
hlabels = [h0,h1,h2,h3,h4,h5]
ax0.legend(fontsize=14,loc=2,handles = hlabels)
ax0.set_xlabel(r'Pressure (GPa)', fontsize=18)
ax0.set_ylabel(r'$F_{vib}$', fontsize=18)
plt.tight_layout()
fig = plt.gcf()
fig.savefig('hcpFeAlloy_Fvib.pdf', format='pdf')
# -
| 078_phoxPlots_FeAlloys/FeAlloy_phox_plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualizations of editing activity in en.wikipedia.org
#
# By [<NAME>](http://stuartgeiger.com), Berkeley Institute for Data Science
#
# (C) 2016, Released under [The MIT license](https://opensource.org/licenses/MIT).
#
# This data is collected and aggregated by <NAME>, which is [here](https://stats.wikimedia.org/EN/TablesWikipediaEN.htm) for the English Wikipedia. I have just copied that data from HTML tables into a CSV (which is not done here), then imported it into Pandas dataframes, and plotted it with matplotlib.
#
# ## Processing and cleaning data
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
from matplotlib.ticker import ScalarFormatter
import numpy as np
# %matplotlib inline
matplotlib.style.use('ggplot')
# Data by <NAME> at https://stats.wikimedia.org/EN/TablesWikipediaEN.htm
counts = pd.read_csv("edit_counts.tsv", sep="\t")
# +
# Data by <NAME>
# Random sample of 50000 users who have registered and made a userpage
# Then, how many edits did they make 1 to 2 years after their registration date
retention = pd.read_csv("retention.tsv", sep="\t")
retention.reg_date=pd.to_datetime(retention.reg_date,format="%Y%m%d%H%M%S")
# -
def survived(edits):
if edits > 0:
return 1
else:
return 0
retention['survival'] = retention.edits_1yr.apply(survived)
retention[0:10]
# Convert dates to datetimes
counts.date=pd.to_datetime(counts.date,infer_datetime_format=True)
# Peek at the dataset
counts[0:10]
# Some of the columns use 'k' for thousands and 'M' for millions, so we need to convert them.
# +
def units_convert(s):
"""
Convert cells with k and M to times 1,000 and 1,000,000 respectively
I got this solution from
http://stackoverflow.com/questions/14218728/converting-string-of-numbers-and-letters-to-int-float-in-pandas-dataframe
"""
powers = {'k': 1000, 'M': 10 ** 6}
if(s[-1] == 'k' or s[-1] == 'M'):
try:
power = s[-1]
return float(s[:-1]) * powers[power]
except TypeError:
return float(s)
else:
return float(s)
# -
# Apply this function to the columns that have 'k' or 'M' units, store them as new _float columns
counts['edits_float']=counts.edits.apply(units_convert)
counts['article_count_float']=counts['article count'].apply(units_convert)
# Make sure we've got data types figured out
counts.dtypes
# +
# Set date column as index
counts.set_index(['date'])
# Calculate some ratios
counts['highly_active_to_newcomer_ratio']=counts['>100 edits']/counts['new accts']
counts['active_to_newcomer_ratio']=counts['>5 edits']/counts['new accts']
counts['highly_active_to_active_ratio']=counts['>100 edits']/(counts['>5 edits']-counts['>100 edits'])
# -
# ## Graphs
# ### Editor retention
import datetime
def dt_to_yearmofirst(dt):
"""
Adding one year to the reg date, because we are looking at if people who registered were still editing 1 year later
"""
year= dt.year + 1
month= dt.month
return datetime.datetime(year=year,month=month,day=1)
# +
retention['reg_mo_first'] = retention.reg_date.apply(dt_to_yearmofirst)
retention[0:10]
# -
retention_group = retention.groupby(["reg_mo_first"])
monthly_averages = retention_group.aggregate({"survival":np.mean})
def add_year(dt):
year = dt.year + 1
month = dt.month
day = dt.day
return datetime.datetime(year, month, day)
# Filter because there is missing data before 2003 and counts after July 2014 aren't accurate
# +
monthly_avg = monthly_averages[monthly_averages.index>datetime.datetime(2004,1,1)]
monthly_avg= monthly_avg[monthly_avg.index<datetime.datetime(2014,9,1)]
# -
monthly_avg.plot()
# ### Number of editors
# +
matplotlib.style.use(['bmh'])
font = {'weight' : 'regular',
'size' : 16}
matplotlib.rc('font', **font)
ax1 = counts.plot(x='date',y=['>5 edits', '>100 edits'], figsize=(12,4),
label="Users making >5 edits in a month", color="r")
ax1.set_xlabel("Year")
ax1.set_ylabel("Number of users")
ax2 = counts.plot(x='date',y='>100 edits', figsize=(12,4),
label="Users making >100 edits in a month",color="g")
ax2.set_xlabel("Year")
ax2.set_ylabel("Number of editors")
ax3 = counts.plot(x='date',y='new accts', figsize=(12,4),
label="New users making >10 edits in a month",color="b")
ax3.set_xlabel("Year")
ax3.set_ylabel("Number of editors")
ax3.yaxis.set_major_formatter(ScalarFormatter())
# +
matplotlib.style.use(['bmh'])
font = {'weight' : 'regular',
'size' : 16}
matplotlib.rc('font', **font)
ax1 = counts.plot(x='date',y=['>5 edits', '>100 edits'], figsize=(12,7),
label="Users making >5 edits in a month", color="bg")
ax1.set_xlabel("Year")
ax1.set_ylabel("Number of users")
ax1.set_ylim(0,70000)
ax2 = ax1.twinx()
ax2 = monthly_avg.plot(ax=ax2,secondary_y=True,color="r")
ax1.set_xlim(372,519)
ax1.legend(loc='upper center', bbox_to_anchor=(0.35, 1.15),
ncol=2, fancybox=True, shadow=True)
ax2.legend(loc='upper center', bbox_to_anchor=(0.75, 1.15),
ncol=1, fancybox=True, shadow=True)
ax2.set_ylabel("1 year newcomer survival")
# +
matplotlib.style.use(['bmh'])
font = {'weight' : 'regular',
'size' : 16}
matplotlib.rc('font', **font)
ax1 = counts.plot(x='date',y=['>5 edits', '>100 edits'], figsize=(12,7),
label="Users making >5 edits in a month", color="bg")
ax1.set_xlabel("Year")
ax1.set_ylabel("Number of users making >x edits/month")
ax1.set_ylim(0,60000)
ax1.set_xlim(372,529)
ax1.legend(loc='upper center', bbox_to_anchor=(0.35, 1.15),
ncol=2, fancybox=True, shadow=True)
# +
ax1 = counts.plot(x='date',y=['>5 edits','>100 edits','new accts'], figsize=(12,4),
label="Users making >5 edits in a month",color=['r','g','b'])
ax1.set_xlabel("Year")
ax1.set_ylabel("Number of users")
ax1.yaxis.set_major_formatter(ScalarFormatter())
# +
ax1 = counts.plot(x='date',y=['>5 edits','>100 edits','new accts'], figsize=(12,4),
label="Users making >5 edits in a month",logy=True, color=['r','g','b'])
ax1.set_xlabel("Year")
ax1.set_ylabel("Number of users")
ax1.yaxis.set_major_formatter(ScalarFormatter())
# -
ax3 = counts.plot(x='date',y='highly_active_to_active_ratio', figsize=(12,4),
label="Highly active users to active users ratio",color="k")
ax3.set_xlabel("Year")
ax3.set_ylabel("Ratio")
# +
ax3 = counts.plot(x='date',y='highly_active_to_newcomer_ratio', figsize=(12,4),
label="Highly active users to newcomers ratio",color="k")
ax3.set_xlabel("Year")
ax3.set_ylabel("Ratio")
# +
ax3 = counts.plot(x='date',y='active_to_newcomer_ratio', figsize=(12,4),
label="Active users to newcomers ratio",color="k")
ax3.set_xlabel("Year")
ax3.set_ylabel("Ratio")
# +
ax3 = counts.plot(x='date',y='edits_float', figsize=(12,4),
label="Number of edits per month",color="k")
ax3.set_xlabel("Year")
ax3.set_ylabel("Number of editors")
# -
# +
ax3 = counts.plot(x='date',y='new per day', figsize=(12,4),
label="New articles written per day",color="k")
ax3.set_xlabel("Year")
ax3.set_ylabel("Number of articles")
# +
ax3 = counts.plot(x='date',y='article_count_float', figsize=(12,4),
label="Number of articles",color="k")
ax3.set_xlabel("Year")
ax3.set_ylabel("Number of articles")
# -
ax3 = counts.plot(x='date',y='article_count_float', figsize=(12,4),
label="Number of articles",color="k",logy=True)
ax3.set_xlabel("Year")
ax3.set_ylabel("Number of articles")
| retention/wiki_edit_counts_retention.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import seaborn as sns
from pandas.plotting import scatter_matrix
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
import math
"""ACTIVITY 1"""
# --- Load data ---
df = pd.read_csv('train.csv')
# --- LINTING 1 --- Drop unecessary data
df.drop('Name', 1, inplace=True)
df.drop('Cabin', 1, inplace=True)
df.drop('PassengerId', 1, inplace=True)
# --- LINTING 2 --- Replace nullish Age values
df['Age'].interpolate(inplace=True)
# --- LINTING 3 --- Drop leftovers wih null
df.dropna(inplace=True)
# --- LINTING 4 --- Replace literals with numeric values
df['Sex'], unique_sex_keys = pd.factorize(df['Sex'])
df['Ticket'], unique_ticket_keys = pd.factorize(df['Ticket'])
#df['Embarked'], unique_embarked_keys = pd.factorize(df['Embarked'])
df = pd.get_dummies(df, columns=['Embarked'])
# --- LINTING 5 --- Normalize values
df.loc[:, ~df.columns.isin(['Survived', 'Embarked'])] = StandardScaler().fit_transform(
df.loc[:, ~df.columns.isin(['Survived', 'Embarked'])]
)
# --- Head visualization ---
df.head()
df.describe()
df.info()
# +
""" ACTIVITY 2"""
def get_euclidian_distance(X_train_row, X_test_row):
for i, val in enumerate(X_test_row):
X_train_row[i] = (X_test_row[i] - X_train_row[i]) ** 2
return X_train_row
class MyKNeighborsClassifier():
def __init__(self, k=5):
self.k = k
def fit(self, X_train, y_train):
self.X_train = X_train.copy()
self.y_train = y_train.copy()
def predict(self, X_test):
prediction = pd.DataFrame(index=X_test.index.copy())
prediction['Survived'] = [None] * len(prediction)
for i, X_test_row in X_test.iterrows():
# Calculate euclidian
X_train_copy = X_train.copy()
y_train_copy = y_train.copy()
X_train_copy.apply(
lambda X_train_row: get_euclidian_distance(X_train_row, X_test_row),
axis=1
)
y_train_copy['Distance'] = (
X_train_copy['Pclass']
#+ X_train_copy['Sex']
+ X_train_copy['Age']
+ X_train_copy['SibSp']
+ X_train_copy['Parch']
+ X_train_copy['Ticket']
+ X_train_copy['Fare']
+ X_train_copy['Embarked_C']
+ X_train_copy['Embarked_Q']
+ X_train_copy['Embarked_S']
).apply(math.sqrt)
# Predict
aux = y_train_copy.nsmallest(
self.k,
['Distance'],
keep='first'
)
prediction['Survived'][i] = aux['Survived'].value_counts()[:1].index.tolist()[0]
return prediction
# Split dataset
X = df.iloc[:, 1:11] # Training data
y = pd.DataFrame(df.iloc[:, 0]) # Answer
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
my_classifier = MyKNeighborsClassifier()
my_classifier.fit(X_train, y_train)
y_pred = my_classifier.predict(X_test)
# Proximo bloco verifica o resultado!!!!
# +
""" ACTIVITY 2: Comparation """
y_pred_copy = y_pred.copy()
y_test_copy = y_test.copy()
y_pred_copy['Survived'] = pd.to_numeric(y_pred_copy['Survived'], downcast="float")
y_test_copy['Survived'] = pd.to_numeric(y_test_copy['Survived'], downcast="float")
wrong = len(y_pred_copy['Survived'].compare(y_test_copy['Survived']))
total = len(y_test)
success = (total - wrong) / total
print(success * 100)
# -
| machine-learning/kNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from scipy.io import loadmat
import pandas as pd
import numpy as np
from scipy.optimize import fmin_cg
import copy
import matplotlib.pyplot as plt
# %matplotlib inline
# -
loc = r'C:\Users\c0w00f8.WMSC\Documents\Coursera\1. Machine Learning\machine-learning-ex4\ex4\ex4data1.mat'
mat_data = loadmat(loc)
data_x = mat_data['X']
data_y = mat_data['y']
data_y.shape
# +
# change the label 10 back to 0
#data_y[data_y == 10] = 0
# here for this assignment, don't change 10 to 0!!! since the weights provided was based on label 10!!!
# if change to 0, the cost test result would be around 10.44
# if stay 10, it would be 0.287629, the correct reference provided by the writeup!
# -
# load provided weights
loc = r'C:\Users\c0w00f8.WMSC\Documents\Coursera\1. Machine Learning\machine-learning-ex4\ex4\ex4weights.mat'
mat = loadmat(loc)
theta1 = mat['Theta1']
theta2 = mat['Theta2']
theta1.shape
# +
#data_x.shape
#theta1.shape - 25 * 401
#theta2.shape - 10 * 26
# -
t1_f = theta1.flatten()
t2_f = theta2.flatten()
t = np.append(t1_f, t2_f)
t.shape
def sigmoid(x):
return 1 / (1 + np.exp(-x))
def mReform(x, k):
m = x.shape[0]
mat = np.zeros((m, 1))
for i in range(k):
label = (x == i+1).astype(int)
mat = np.hstack((mat, label))
return mat[:, 1:]
testy = mReform(data_y, 10)
testy
a1 = np.arange(9).reshape(3, 3)
b1 = np.array([
[3, 1, 2],
[9, 1, 5],
[7, 4, 6]
])
b1
b1 **2
a1
(- a1 - b1)
np.multiply(a1, b1)
np.multiply(a1, b1).sum()
np.log(b1)
np.multiply(np.log(b1), a1)
def nnCost(theta, hid_size, x, y, num_labels, lam):
# assume the network only have 1 input layer, 1 hidden layer, and 1 output layer
m = x.shape[0]
# reshape theta1 and theta2
inp_size = x.shape[1]
theta_cutoff = hid_size * (inp_size + 1)
theta1 = theta[:theta_cutoff].reshape((hid_size, inp_size + 1))
theta2 = theta[theta_cutoff:].reshape((num_labels, hid_size + 1))
#print ('theta1', type(theta1), theta1.shape)
#print ('theta2', type(theta2), theta2.shape)
# adjust x and insert x0 = 1
x = np.insert(x, 0, 1, axis = 1)
# x.shape - 5000 * 401
# compute hidden layer value
hidden = sigmoid(x.dot(theta1.T))
#print ('hidden layer', type(hidden), hidden.shape)
# hidden.shape - 5000 * 25
# add bias unit to hidden layer
hid = np.insert(hidden, 0, 1, axis = 1)
#print ('hid+bias unit', type(hid), hid.shape)
# hid.shape - 5000 * 26
pred = sigmoid(hid.dot(theta2.T))
#print ('pred', type(pred), pred.shape)
# pred.shape - 5000 * 10
#out = np.argmax(pred, axis = 1).reshape((m, 1))
#print ('out', type(out), out.shape)
#out = mReform(out, num_labels)
# reform y to matrix
y = mReform(y, num_labels)
# turn y into 5000 * 10 matrix
# compute cost
j = ( - np.multiply(y, np.log(pred)) - np.multiply(1 - y, np.log(1 - pred))).sum() / m
# perform the same when using for-loop
#cost = 0
#for i in range(pred.shape[0]):
# for j in range(pred.shape[1]):
# cost = cost - y[i, j] * np.log(pred[i, j]) - (1 - y[i, j]) * np.log(1 - pred[i, j])
return j
# +
# test nnCost
testC = nnCost(t, 25, data_x, data_y, 10, 1)
testC
# ~0.287629
#tt1, tt2 = nnCost(t, 25, data_x, data_y, 10, 1)
#if ((tt1 == theta1).all() == True) and ((tt2 == theta2).all() == True):
# print ("theta reshape test passed")
# -
def nnCostReg(theta, hid_size, x, y, num_labels, lam):
m = x.shape[0]
# reshape theta1 and theta2
inp_size = x.shape[1]
theta_cutoff = hid_size * (inp_size + 1)
theta1 = theta[:theta_cutoff].reshape((hid_size, inp_size + 1))
theta2 = theta[theta_cutoff:].reshape((num_labels, hid_size + 1))
#theta1.shape - 25 * 401
#theta2.shape - 10 * 26
# adjust x and insert x0 = 1
x = np.insert(x, 0, 1, axis = 1)
# x.shape - 5000 * 401
# compute hidden layer value
hidden = sigmoid(x.dot(theta1.T))
# hidden.shape - 5000 * 25
# add bias unit to hidden layer
hid = np.insert(hidden, 0, 1, axis = 1)
# hid.shape - 5000 * 26
pred = sigmoid(hid.dot(theta2.T))
# pred.shape - 5000 * 10
# reform y to matrix
y = mReform(y, num_labels)
# turn y into 5000 * 10 matrix
# compute cost
j = ( - np.multiply(y, np.log(pred)) - np.multiply(1 - y, np.log(1 - pred))).sum() / m
# Regularition
reg = ((theta1[:, 1:] ** 2).sum() + (theta2[:, 1:] ** 2).sum()) * lam / 2 / m
return j + reg
testR = nnCostReg(t, 25, data_x, data_y, 10, 1)
testR
# ~0.38377
def sigmdGrad(x):
return np.multiply(sigmoid(x), 1 - sigmoid(x))
def randInit(l_in, l_out):
epsilon = 0.12
sample = np.random.uniform(-epsilon, epsilon, l_out * (l_in + 1))
return sample.reshape(l_out, l_in + 1)
def backprop(theta, hid_size, x, y, num_labels, lam):
m = x.shape[0]
inp_size = x.shape[1]
theta_cutoff = hid_size * (inp_size + 1)
theta1 = theta[:theta_cutoff].reshape((hid_size, inp_size + 1))
theta2 = theta[theta_cutoff:].reshape((num_labels, hid_size + 1))
x = np.insert(x, 0, 1, axis = 1)
hidden = sigmoid(x.dot(theta1.T))
# a2 = hid
a2 = np.insert(hidden, 0, 1, axis = 1)
# a3 = pred
a3 = sigmoid(a2.dot(theta2.T))
y = mReform(y, num_labels)
error3 = a3 - y
#print ('error3', error3.shape)
#error3.shape - 5000 * 10
# theta1 - 25 * 401
# theta2 - 10 * 26
# add 1 column to z2
z2 = x.dot(theta1.T)
z2 = np.insert(z2, 0, 1, axis = 1)
# z2.shape - 5000 * 26
gz2 = sigmdGrad(z2)
# gz2.shape - 5000 * 26
error2 = np.multiply(error3.dot(theta2), gz2)
#print ('error2', error2.shape)
#error2.shape - 5000 * 26
delta2 = error3.T.dot(a2)
# print ('delta2', delta2.shape)
# delta2.shape - 10 * 26
# ignore error2[:,0]
delta1 = error2[:, 1:].T.dot(x)
# print ('delta1', delta1.shape)
# delta1.shape - 25 * 401
delta1 = (delta1 / m).flatten()
delta2 = (delta2 / m).flatten()
delta = np.append(delta1, delta2)
#print ('delta', delta.shape)
return delta
testB = backprop(t, 25, data_x, data_y, 10, 1)
#testB
def compGrad(theta, hid_size, x, y, num_labels, lam):
epsilon = 0.0001
grad = np.zeros(theta.shape)
for i in range(len(theta)):
theta_pos = copy.copy(theta)
theta_neg = copy.copy(theta)
theta_pos[i] += epsilon
theta_neg[i] -= epsilon
grad[i] = (nnCost(theta_pos, hid_size, x, y, num_labels, lam) - nnCost(theta_neg, hid_size, x, y, num_labels, lam)) /2/epsilon
return grad
def checkGrad(in_size, hid_size, n_labels, m, lam=1):
# prep data
theta1 = randInit(in_size, hid_size)
#print ("theta1", theta1.shape)
theta2 = randInit(hid_size, n_labels)
#print ("theta2", theta2.shape)
x = randInit(in_size - 1, m)
#print ("x", x.shape)
y = (np.random.randint(1, n_labels+1, m)).reshape(m, 1)
t1 = theta1.flatten()
t2 = theta2.flatten()
t = np.append(t1, t2)
gradNN = backprop(t, hid_size, x, y, n_labels, lam)
gradC = compGrad(t, hid_size, x, y, n_labels, lam)
diff = abs(gradNN - gradC)
return (diff <= 1e-9).all() == True
if checkGrad(3, 5, 3, 5) == True:
print ('Gradient check passed!')
# +
#yy = np.random.randint(1, 6, size=10)
#yy = yy.reshape(10, 1)
#yy = mReform(yy, 5)
#yy
# -
def backpropReg(theta, hid_size, x, y, num_labels, lam):
m = x.shape[0]
inp_size = x.shape[1]
theta_cutoff = hid_size * (inp_size + 1)
theta1 = theta[:theta_cutoff].reshape((hid_size, inp_size + 1))
theta2 = theta[theta_cutoff:].reshape((num_labels, hid_size + 1))
x = np.insert(x, 0, 1, axis = 1)
hidden = sigmoid(x.dot(theta1.T))
# a2 = hid
a2 = np.insert(hidden, 0, 1, axis = 1)
# a3 = pred
a3 = sigmoid(a2.dot(theta2.T))
y = mReform(y, num_labels)
error3 = a3 - y
#error3.shape - 5000 * 10
# theta1 - 25 * 401
# theta2 - 10 * 26
# add 1 column to z2
z2 = x.dot(theta1.T)
z2 = np.insert(z2, 0, 1, axis = 1)
# z2.shape - 5000 * 26
gz2 = sigmdGrad(z2)
# gz2.shape - 5000 * 26
error2 = np.multiply(error3.dot(theta2), gz2)
#error2.shape - 5000 * 26
delta2 = error3.T.dot(a2)
# delta2.shape - 10 * 26
# ignore error2[:,0]
delta1 = error2[:, 1:].T.dot(x)
# delta1.shape - 25 * 401
delta1 = delta1 / m
delta2 = delta2 / m
reg1 = theta1 * lam / m
reg1[:, 0] = 0
reg2 = theta2 * lam / m
reg2[:, 0] = 0
delta1 += reg1
delta2 += reg2
delta = np.append(delta1.flatten(), delta2.flatten())
#print ('delta', delta.shape)
return delta
def compGradReg(theta, hid_size, x, y, num_labels, lam):
epsilon = 0.0001
grad = np.zeros(theta.shape)
for i in range(len(theta)):
theta_pos = copy.copy(theta)
theta_neg = copy.copy(theta)
theta_pos[i] += epsilon
theta_neg[i] -= epsilon
grad[i] = (nnCostReg(theta_pos, hid_size, x, y, num_labels, lam) - nnCostReg(theta_neg, hid_size, x, y, num_labels, lam)) /2/epsilon
return grad
def checkGradReg(in_size, hid_size, n_labels, m, lam=1):
# prep data
theta1 = randInit(in_size, hid_size)
#print ("theta1", theta1.shape)
theta2 = randInit(hid_size, n_labels)
#print ("theta2", theta2.shape)
x = randInit(in_size - 1, m)
#print ("x", x.shape)
y = (np.random.randint(1, n_labels+1, m)).reshape(m, 1)
t1 = theta1.flatten()
t2 = theta2.flatten()
t = np.append(t1, t2)
gradNN = backpropReg(t, hid_size, x, y, n_labels, lam)
gradC = compGradReg(t, hid_size, x, y, n_labels, lam)
diff = abs(gradNN - gradC)
return (diff <= 1e-9).all() == True
if checkGradReg(3, 5, 3, 5) == True:
print ('Gradient check passed!')
else: print ('Go check again')
# +
# learn the model with fmin_cg
# initialize theta
theta1 = randInit(400, 25)
theta2 = randInit(25, 10)
theta0 = np.append(theta1.flatten(), theta2.flatten())
lam = 1
hid_size = 25
n_labels = 10
x = data_x
y = data_y
myargs = (hid_size, x, y, n_labels, lam)
# train the model
train = fmin_cg(nnCostReg, theta0, args = myargs, fprime = backpropReg, maxiter = 400)
#train = fmin_cg(nnCostReg, theta0, args = myargs, fprime = backpropReg)
# -
train
def predict(theta, hid_size, x, num_labels):
m = x.shape[0]
inp_size = x.shape[1]
theta_cutoff = hid_size * (inp_size + 1)
theta1 = theta[:theta_cutoff].reshape((hid_size, inp_size + 1))
theta2 = theta[theta_cutoff:].reshape((num_labels, hid_size + 1))
x = np.insert(x, 0, 1, axis = 1)
hidden = sigmoid(x.dot(theta1.T))
# a2 = hid
a2 = np.insert(hidden, 0, 1, axis = 1)
# a3 = pred
a3 = sigmoid(a2.dot(theta2.T))
pred = np.argmax(a3, axis = 1)
pred += 1
return pred
def accuracy(y, pred):
y = y.flatten()
m = len(y)
count = (y == pred).sum()
accuracy = count / m
return accuracy
pred = predict(train, 25, data_x, 10)
accu = accuracy(data_y, pred)
accu
# the accuracy is 99.42% when lambda is 1
# accu = 98.62% when lambda is 2
# accu = 97.66% when lambda is 3
# visualize hidden layer
theta1 = train[:10025].reshape(25, 401)
tt1 = theta1[:, 1:]
tt1.shape
def toMatrix(data, row, col):
mat = np.zeros(shape = (row, col))
m = 0
for j in range(col):
for i in range(row):
mat[i, j] = data[m]
m += 1
return mat
test2 = toMatrix(tt1[3, :], 20, 20)
imgplot = plt.imshow(test2, cmap='gray')
| hw4/ex4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # What kind of descriptors are we dealing with?
import pandas as pd
data = pd.read_excel("../data/Descriptors.xls")
data.head(5)
# how many descriptor classes are there?
def isNaN(num):
return num != num
descriptor_class = data["Descriptor Java Class"]
i = 0
for descriptor in descriptor_class:
if isNaN(descriptor):
descriptor_class[i] = descriptor_class[i-1]
i += 1
descriptor_counts = pd.DataFrame(descriptor_class.value_counts())
descriptor_counts.shape
sum(descriptor_counts.iloc[:,0])
# +
from collections import Counter
def countDistinct(descriptor_class):
# counter method gives dictionary of elements in list
# with their corresponding frequency.
# using keys() method of dictionary data structure
# we can count distinct values in array
return len(Counter(descriptor_class).keys())
Counter(descriptor_class)
| EDA-Notebooks/Workbook_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: astro_sprint
# language: python
# name: astro_sprint
# ---
import keras,os
from keras.models import Sequential
from keras.layers import Dense, Conv2D, MaxPool2D , Flatten
# from keras.preprocessing.image import ImageDataGenerator
import numpy as np
import imageio
import glob
import matplotlib.pyplot as plt
from keras.utils import to_categorical
from sklearn.model_selection import train_test_split
import tensorflow as tf
x1 = []
for filename in glob.glob('~/Documents/astroSprint/output_rgb_format_data/lsst_triplets/single_fake2/*.png'):
im = imageio.imread(filename)
x1.append(im)
x2 = []
for filename in glob.glob('~/Documents/astroSprint//output_rgb_format_data/lsst_triplets/two_fakes2/*.png'):
im = imageio.imread(filename)
x2.append(im)
y1=[]
for i in enumerate(x1):
y1.append(0)
y2=[]
for i in enumerate(x2):
y2.append(1)
x = []
x.extend(x1)
x.extend(x2)
# x = np.expand_dims(x, axis=1)
x = np.array(x)
y = []
y.extend(y1)
y.extend(y2)
# y = to_categorical(y, num_classes=2)
# y = np.array(y)
# #### Approach 1 - Training our own VGG16 model from scratch
X_train, X_test, y_train, y_test = train_test_split(x, y, random_state=42, test_size=0.1)
X_train.shape
# +
model = Sequential()
model.add(Conv2D(input_shape=(224,224,3),filters=64,kernel_size=(3,3),padding="same", activation="relu"))
model.add(Conv2D(filters=64,kernel_size=(3,3),padding="same", activation="relu"))
model.add(MaxPool2D(pool_size=(2,2),strides=(2,2)))
model.add(Conv2D(filters=128, kernel_size=(3,3), padding="same", activation="relu"))
model.add(Conv2D(filters=128, kernel_size=(3,3), padding="same", activation="relu"))
model.add(MaxPool2D(pool_size=(2,2),strides=(2,2)))
model.add(Conv2D(filters=256, kernel_size=(3,3), padding="same", activation="relu"))
model.add(Conv2D(filters=256, kernel_size=(3,3), padding="same", activation="relu"))
model.add(Conv2D(filters=256, kernel_size=(3,3), padding="same", activation="relu"))
model.add(MaxPool2D(pool_size=(2,2),strides=(2,2)))
model.add(Conv2D(filters=512, kernel_size=(3,3), padding="same", activation="relu"))
model.add(Conv2D(filters=512, kernel_size=(3,3), padding="same", activation="relu"))
model.add(Conv2D(filters=512, kernel_size=(3,3), padding="same", activation="relu"))
model.add(MaxPool2D(pool_size=(2,2),strides=(2,2)))
model.add(Conv2D(filters=512, kernel_size=(3,3), padding="same", activation="relu"))
model.add(Conv2D(filters=512, kernel_size=(3,3), padding="same", activation="relu"))
model.add(Conv2D(filters=512, kernel_size=(3,3), padding="same", activation="relu"))
model.add(MaxPool2D(pool_size=(2,2),strides=(2,2)))
model.add(Flatten())
model.add(Dense(units=4096,activation="relu"))
model.add(Dense(units=4096,activation="relu"))
model.add(Dense(units=2, activation="softmax"))
# -
from keras.optimizers import Adam
opt = Adam(lr=0.001)
model.compile(optimizer=opt, loss=keras.losses.categorical_crossentropy,
metrics=['accuracy',
tf.keras.metrics.AUC(num_thresholds=1200, curve='ROC', summation_method='interpolation')])
model.summary()
from keras.callbacks import ModelCheckpoint, EarlyStopping
checkpoint = ModelCheckpoint("vgg16_1.h5", monitor='val_accuracy', verbose=1, save_best_only=True, save_weights_only=False, mode='auto')
early = EarlyStopping(monitor='val_accuracy', min_delta=0, patience=5, verbose=1, mode='auto')
# hist = model.fit_generator(steps_per_epoch=10,generator=(X_train, y_train), validation_data= (X_test, y_test), validation_steps=10,epochs=50,callbacks=[checkpoint,early])
hist = model.fit(X_train, y_train, epochs=50, validation_data=(X_test, y_test), batch_size=32,callbacks=[checkpoint,early])
import matplotlib.pyplot as plt
plt.plot(hist.history["acc"])
plt.plot(hist.history['val_acc'])
plt.plot(hist.history['loss'])
plt.plot(hist.history['val_loss'])
plt.title("model accuracy")
plt.ylabel("Accuracy")
plt.xlabel("Epoch")
plt.legend(["Accuracy","Validation Accuracy","loss","Validation Loss"])
plt.savefig("trainvalid.png")
# #### Approach 2 - Using weights of pre-trained VGG16 model for classification
from keras.models import Model
from keras.applications import vgg16
vgg_model = vgg16.VGG16(weights='imagenet')
feat_extractor = Model(inputs=vgg_model.input, outputs=vgg_model.get_layer("fc2").output)
feat_1 =tf.keras.applications.VGG16()
feat_1.summary()
# +
# from keras.applications.imagenet_utils import preprocess_input
# img_arr=np.vstack(x)
# processed_imgs= preprocess_input(img_arr.copy())
img_features=feat_extractor.predict(x)
img_features.shape
# -
X_train2, X_test2, y_train, y_test = train_test_split(img_features, y, random_state=42, test_size=0.1)
# +
penalty = ['l2','l1']
solver = ['liblinear']
C = [1e-4,1e-3,1e-2,1e-1]
class_weight = ['balanced']
multi_class = ['ovr']
# Create hyperparameter options
hyperparameters = dict(C=C, penalty=penalty, class_weight=class_weight, solver=solver, multi_class=multi_class)
# -
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import GridSearchCV, cross_val_predict
# model = LogisticRegression()
logreg = GridSearchCV(LogisticRegression(), hyperparameters, scoring='roc_auc', cv=5)
logreg.fit(X_train2, y_train)
# +
from sklearn.metrics import confusion_matrix, roc_auc_score, classification_report
print(logreg.best_params_)
y_pred = cross_val_predict(logreg, X_test2, y_test, cv=5)
print('Best Penalty:', logreg.best_estimator_.get_params()['penalty'])
print('Best C:', logreg.best_estimator_.get_params()['C'])
print('Overall AUC:', roc_auc_score(y_test, y_pred))
print('Accuracy of logistic regression classifier on validate set (RFE): {:.2f}'.format(
logreg.score(X_test2, y_test)))
confusion_matrix = confusion_matrix(y_test, y_pred)
print(confusion_matrix)
# Compute precision, recall, F-measure and support
print(classification_report(y_test, y_pred))
specificity = confusion_matrix[0, 0] / (confusion_matrix[0, 0] + confusion_matrix[0, 1])
print('specificity : ', specificity)
sensitivity = confusion_matrix[1, 1] / (confusion_matrix[1, 0] + confusion_matrix[1, 1])
print('sensitivity : ', sensitivity)
print('fawadMetric : ', sensitivity * specificity)
# -
n_estimators = [5, 10, 15]
# max_features = [1]
class_weight = ['balanced_subsample']
criterion = ['gini']
min_samples_split = [2, 5]
min_samples_leaf = [2, 5, 10]
# max_depth = [None]
# max_leaf_nodes = [3, 7, None]
# min_impurity_decrease = [0.0]
# min_weight_fraction_leaf = [0]
n_jobs = [-1]
# oob_score = [True]
random_state = [888888]
param_grid = dict(n_estimators=n_estimators, criterion=criterion, min_samples_split=min_samples_split,
n_jobs=n_jobs, random_state=random_state, class_weight=class_weight,
min_samples_leaf=min_samples_leaf)
X_train2.shape[1]
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV, cross_val_predict
trained_model = RandomForestClassifier()
rfc = GridSearchCV(estimator=trained_model, param_grid=dict(param_grid, max_features=[X_train2.shape[1]]), scoring='roc_auc', cv=5)
rfc.fit(X_train2, y_train)
# +
from sklearn.metrics import confusion_matrix, roc_auc_score, classification_report
print(rfc.best_params_)
y_pred = cross_val_predict(rfc, X_test2, y_test, cv=5)
print('Overall AUC:', roc_auc_score(y_test, y_pred))
print('Accuracy of Random Forest classifier on validate set (RFE): {:.2f}'.format(
rfc.score(X_test2, y_test)))
confusion_matrix = confusion_matrix(y_test, y_pred)
print(confusion_matrix)
# Compute precision, recall, F-measure and support
print(classification_report(y_test, y_pred))
specificity = confusion_matrix[0, 0] / (confusion_matrix[0, 0] + confusion_matrix[0, 1])
print('specificity : ', specificity)
sensitivity = confusion_matrix[1, 1] / (confusion_matrix[1, 0] + confusion_matrix[1, 1])
print('sensitivity : ', sensitivity)
print('fawadMetric : ', sensitivity * specificity)
# -
| diffimageml/examples/lsst_vgg16.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/probml/pyprobml/blob/master/book1/supplements/logreg_pytorch.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="b520E1nCIBHc"
#
# # Logistic regression using PyTorch
#
# We show how to fit a logistic regression model using PyTorch. The log likelihood for this model is convex, so we can compute the globally optimal MLE. This makes it easy to compare to sklearn (and other implementations).
#
#
# + id="UeuOgABaIENZ"
import sklearn
import scipy
import scipy.optimize
import matplotlib.pyplot as plt
from mpl_toolkits import mplot3d
from mpl_toolkits.mplot3d import Axes3D
import seaborn as sns
import warnings
warnings.filterwarnings('ignore')
import itertools
import time
from functools import partial
import os
import numpy as np
from scipy.special import logsumexp
np.set_printoptions(precision=3)
np.set_printoptions(formatter={'float': lambda x: "{0:0.3f}".format(x)})
# + id="GPozRwDAKFb8" colab={"base_uri": "https://localhost:8080/"} outputId="2a8248be-61d4-43fb-9cce-2c36e47d3be5"
import torch
import torch.nn as nn
import torchvision
print("torch version {}".format(torch.__version__))
if torch.cuda.is_available():
print(torch.cuda.get_device_name(0))
print("current device {}".format(torch.cuda.current_device()))
else:
print("Torch cannot find GPU")
def set_seed(seed):
np.random.seed(seed)
torch.manual_seed(seed)
torch.cuda.manual_seed_all(seed)
use_cuda = torch.cuda.is_available()
device = torch.device("cuda:0" if use_cuda else "cpu")
#torch.backends.cudnn.benchmark = True
# + [markdown] id="kjP6xqkvbKxe"
# # Logistic regression using sklearn
#
# We fit binary logistic regresion on the Iris dataset.
# + colab={"base_uri": "https://localhost:8080/"} id="aSYkjaAO6n3A" outputId="e5c90ccb-b01b-4115-ac78-1524e72b63e1"
# Fit the model usign sklearn
import sklearn.datasets
from sklearn.model_selection import train_test_split
iris = sklearn.datasets.load_iris()
X = iris["data"]
y = (iris["target"] == 2).astype(np.int) # 1 if Iris-Virginica, else 0'
N, D = X.shape # 150, 4
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.33, random_state=42)
from sklearn.linear_model import LogisticRegression
# We set C to a large number to turn off regularization.
# We don't fit the bias term to simplify the comparison below.
log_reg = LogisticRegression(solver="lbfgs", C=1e5, fit_intercept=False)
log_reg.fit(X_train, y_train)
w_mle_sklearn = np.ravel(log_reg.coef_)
print(w_mle_sklearn)
# + [markdown] id="-pIgD7iRLUBt"
# # Automatic differentiation <a class="anchor" id="AD"></a>
#
#
# In this section, we illustrate how to use autograd to compute the gradient of the negative log likelihood for binary logistic regression. We first compute the gradient by hand, and then use PyTorch's autograd feature.
# (See also [the JAX optimization colab](https://colab.research.google.com/github/probml/pyprobml/blob/master/book1/supplements/opt.ipynb).)
#
# + [markdown] id="0p5y7b8NbyZp"
# ## Computing gradients by hand
#
# + id="iS5AB9NjLZ_i"
# Binary cross entropy
def BCE_with_logits(logits, targets):
N = logits.shape[0]
logits = logits.reshape(N,1)
logits_plus = np.hstack([np.zeros((N,1)), logits]) # e^0=1
logits_minus = np.hstack([np.zeros((N,1)), -logits])
logp1 = -logsumexp(logits_minus, axis=1)
logp0 = -logsumexp(logits_plus, axis=1)
logprobs = logp1 * targets + logp0 * (1-targets)
return -np.sum(logprobs)/N
# Compute using numpy
def sigmoid(x): return 0.5 * (np.tanh(x / 2.) + 1)
def predict_logit(weights, inputs):
return np.dot(inputs, weights) # Already vectorized
def predict_prob(weights, inputs):
return sigmoid(predict_logit(weights, inputs))
def NLL(weights, batch):
X, y = batch
logits = predict_logit(weights, X)
return BCE_with_logits(logits, y)
def NLL_grad(weights, batch):
X, y = batch
N = X.shape[0]
mu = predict_prob(weights, X)
g = np.sum(np.dot(np.diag(mu - y), X), axis=0)/N
return g
# + colab={"base_uri": "https://localhost:8080/"} id="f9mD8S18746_" outputId="25c03ff4-36f7-4e49-81c6-c56c9f369bd3"
w_np = w_mle_sklearn
y_pred = predict_prob(w_np, X_test)
loss_np = NLL(w_np, (X_test, y_test))
grad_np = NLL_grad(w_np, (X_test, y_test))
print("params {}".format(w_np))
#print("pred {}".format(y_pred))
print("loss {}".format(loss_np))
print("grad {}".format(grad_np))
# + [markdown] id="YeGQ7SJTNHMk"
# ## PyTorch code
# + [markdown] id="Is7yJlgsL4BT"
# To compute the gradient using torch, we proceed as follows.
#
# - declare all the variables that you want to take derivatives with respect to using the requires_grad=True argumnet
# - define the (scalar output) objective function you want to differentiate in terms of these variables, and evaluate it at a point. This will generate a computation graph and store all the tensors.
# - call objective.backward() to trigger backpropagation (chain rule) on this graph.
# - extract the gradients from each variable using variable.grad field. (These will be torch tensors.)
#
# See the example below.
# + id="Wl_SK0WUlvNl"
# data. By default, numpy uses double but torch uses float
X_train_t = torch.tensor(X_train, dtype=torch.float)
y_train_t = torch.tensor(y_train, dtype=torch.float)
X_test_t = torch.tensor(X_test, dtype=torch.float)
y_test_t = torch.tensor(y_test, dtype=torch.float)
# + id="0L5NxIaVLu64" colab={"base_uri": "https://localhost:8080/"} outputId="2215d4e6-76c5-4437-950f-cb0430708f3a"
# parameters
W = np.reshape(w_mle_sklearn, [D, 1]) # convert 1d vector to 2d matrix
w_torch = torch.tensor(W, requires_grad=True, dtype=torch.float)
#w_torch.requires_grad_()
# binary logistic regression in one line of Pytorch
def predict_t(w, X):
y_pred = torch.sigmoid(torch.matmul(X, w))[:,0]
return y_pred
# This returns Nx1 probabilities
y_pred = predict_t(w_torch, X_test_t)
# loss function is average NLL
criterion = torch.nn.BCELoss(reduction='mean')
loss_torch = criterion(y_pred, y_test_t)
print(loss_torch)
# Backprop
loss_torch.backward()
print(w_torch.grad)
# convert to numpy. We have to "detach" the gradient tracing feature
loss_torch = loss_torch.detach().numpy()
grad_torch = w_torch.grad[:,0].detach().numpy()
# + colab={"base_uri": "https://localhost:8080/"} id="CSKAJvrBNKQC" outputId="5e6fc814-03aa-4518-8263-3951a40d4ed3"
# Test
assert np.allclose(loss_np, loss_torch)
assert np.allclose(grad_np, grad_torch)
print("loss {}".format(loss_torch))
print("grad {}".format(grad_torch))
# + [markdown] id="DLWeq4d-6Upz"
# # Batch optimization using BFGS
#
# We will use BFGS from PyTorch for fitting a logistic regression model, and compare to sklearn.
# + colab={"base_uri": "https://localhost:8080/"} id="yiefA00AuXK4" outputId="7d19a74b-69c1-4aea-eeb2-a95a2f429553"
set_seed(0)
params = torch.randn((D,1), requires_grad=True)
optimizer = torch.optim.LBFGS([params], history_size=10)
def closure():
optimizer.zero_grad()
y_pred = predict_t(params, X_train_t)
loss = criterion(y_pred, y_train_t)
loss.backward()
return loss
max_iter = 10
for i in range(max_iter):
loss = optimizer.step(closure)
print(loss.item())
# + colab={"base_uri": "https://localhost:8080/"} id="gcsx3JCGuISp" outputId="1f33c971-164f-4555-bd74-efaecb4664b3"
print("parameters from sklearn {}".format(w_mle_sklearn))
print("parameters from torch {}".format(params[:,0]))
# + colab={"base_uri": "https://localhost:8080/"} id="LSt8z7m5uuvK" outputId="dba94f42-47db-43e9-bf9d-ed62ce3bcb5d"
p_pred_np = predict_prob(w_np, X_test)
p_pred_t = predict_t(params, X_test_t)
p_pred = p_pred_t.detach().numpy()
np.set_printoptions(formatter={'float': lambda x: "{0:0.3f}".format(x)})
print(p_pred_np)
print(p_pred)
# + [markdown] id="8TMzOBNtUaW6"
# # Stochastic optimization using SGD
# + [markdown] id="9byvNfJ9QpsH"
# ## DataLoader
#
# First we need a way to get minbatches of data.
# + colab={"base_uri": "https://localhost:8080/"} id="O_jliQydRXUB" outputId="3ddf9e6b-caaa-40ea-c67d-25e615e26fc5"
from torch.utils.data import DataLoader, TensorDataset
# To make things interesting, we pick a batchsize of B=33, which is not divisible by N=100
dataset = TensorDataset(X_train_t, y_train_t)
B = 33
dataloader = DataLoader(dataset, batch_size=B, shuffle=True)
print(X_train_t.shape)
print('{} examples divided into {} batches of size {}'.format(
len(dataloader.dataset), len(dataloader), dataloader.batch_size))
for i, batch in enumerate(dataloader):
X, y = batch
print(X.shape)
print(y.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="ui_gFE0wWSIS" outputId="ab3b8828-c4eb-4be0-a7cd-dac11770c7c9"
datastream = iter(dataloader)
for i in range(3):
X,y = next(datastream)
print(y)
# + [markdown] id="Wux6hg6JVe7O"
# ## Vanilla SGD training loop
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="bXBNl-vwVejO" outputId="6bbec107-9bfb-432d-f519-f34da6a95cb5"
set_seed(0)
params = torch.randn((D,1), requires_grad=True)
nepochs = 100
nbatches = len(dataloader)
criterion = torch.nn.BCELoss(reduction='mean')
learning_rate = 1e-1
loss_trace = []
for epoch in range(nepochs):
for b, batch in enumerate(dataloader):
X, y = batch
if params.grad is not None:
params.grad.zero_() # reset gradient to zero
y_pred = predict_t(params, X)
loss = criterion(y_pred, y)
#print(f'epoch {epoch}, batch {b}, loss: {loss.item()}')
loss_trace.append(loss)
loss.backward()
with torch.no_grad():
params -= learning_rate * params.grad
#print(f'end of epoch {epoch}, loss: {loss.item()}')
plt.figure()
plt.plot(loss_trace)
# + colab={"base_uri": "https://localhost:8080/"} id="TK-4_-N5o4sK" outputId="fd3a3ed5-ddac-4bd3-e4ad-618956245754"
# SGD does not converge to a value that is close to the batch solver...
print("parameters from sklearn {}".format(w_mle_sklearn))
print("parameters from torch {}".format(params[:,0]))
# + colab={"base_uri": "https://localhost:8080/"} id="0dBeBatDo_Xy" outputId="073ed389-f504-4bf9-ba07-d06a5da0d6a4"
# Predicted probabilities from SGD are very different to sklearn
# although the thresholded labels are similar
p_pred_np = predict_prob(w_np, X_test)
p_pred_t = predict_t(params, X_test_t)
p_pred = p_pred_t.detach().numpy()
print(p_pred_np)
print(p_pred)
# + colab={"base_uri": "https://localhost:8080/"} id="5e8Wugc1eLHT" outputId="e9c62c71-bbad-4e7a-fe81-5e4279f0a6d2"
y_pred_np = p_pred_np > 0.5
y_pred = p_pred > 0.5
print(y_pred_np)
print(y_pred)
print(np.sum(y_pred_np == y_pred)/len(y_pred))
# + [markdown] id="2AlO5fUmrMzI"
# ## Use Torch SGD optimizer
#
# Instead of writing our own optimizer, we can use a torch optimizer. This should give identical results.
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="YzC12T6mrOmb" outputId="8725516e-dbd3-4e20-efdc-7ecc76944d5d"
set_seed(0)
params = torch.randn((D,1), requires_grad=True)
nepochs = 100
nbatches = len(dataloader)
criterion = torch.nn.BCELoss(reduction='mean')
learning_rate = 1e-1
loss_trace = []
# optimizer has pointer to params, so can mutate its state
optimizer = torch.optim.SGD([params], lr=learning_rate)
for epoch in range(nepochs):
for b, batch in enumerate(dataloader):
X, y = batch
y_pred = predict_t(params, X)
loss = criterion(y_pred, y)
#print(f'epoch {epoch}, batch {b}, loss: {loss.item()}')
loss_trace.append(loss)
optimizer.zero_grad()
loss.backward()
optimizer.step()
#print(f'end of epoch {epoch}, loss: {loss.item()}')
plt.figure()
plt.plot(loss_trace)
# + id="IMxLRicCvW_Y" outputId="eb6e6e21-7533-4326-c4c7-3aafe46f2f2f" colab={"base_uri": "https://localhost:8080/"}
print("parameters from sklearn {}".format(w_mle_sklearn))
print("parameters from torch {}".format(params[:,0]))
p_pred_np = predict_prob(w_np, X_test)
p_pred_t = predict_t(params, X_test_t)
p_pred = p_pred_t.detach().numpy()
print('predictions from sklearn')
print(p_pred_np)
print('predictions from torch')
print(p_pred)
y_pred_np = p_pred_np > 0.5
y_pred = p_pred > 0.5
print('fraction of predicted labels that agree ', np.sum(y_pred_np == y_pred)/len(y_pred))
# + [markdown] id="Hr8WRZP6vtBT"
# ## Use momentum optimizer
#
# Adding momentum helps a lot, and gives results which are very similar to batch optimization.
# + id="3D4E4JGdvvcU" outputId="09114882-d6fd-493d-86cb-41d9d100be99" colab={"base_uri": "https://localhost:8080/", "height": 282}
set_seed(0)
params = torch.randn((D,1), requires_grad=True)
nepochs = 100
nbatches = len(dataloader)
criterion = torch.nn.BCELoss(reduction='mean')
learning_rate = 1e-1
loss_trace = []
# optimizer has pointer to params, so can mutate its state
optimizer = torch.optim.SGD([params], lr=learning_rate, momentum=0.9)
for epoch in range(nepochs):
for b, batch in enumerate(dataloader):
X, y = batch
y_pred = predict_t(params, X)
loss = criterion(y_pred, y)
#print(f'epoch {epoch}, batch {b}, loss: {loss.item()}')
loss_trace.append(loss)
optimizer.zero_grad()
loss.backward()
optimizer.step()
#print(f'end of epoch {epoch}, loss: {loss.item()}')
plt.figure()
plt.plot(loss_trace)
# + id="ym9Lz7tCv41V" outputId="23d5a414-f457-4c25-d28e-3b4319db53c5" colab={"base_uri": "https://localhost:8080/"}
print("parameters from sklearn {}".format(w_mle_sklearn))
print("parameters from torch {}".format(params[:,0]))
p_pred_np = predict_prob(w_np, X_test)
p_pred_t = predict_t(params, X_test_t)
p_pred = p_pred_t.detach().numpy()
print('predictions from sklearn')
print(p_pred_np)
print('predictions from torch')
print(p_pred)
y_pred_np = p_pred_np > 0.5
y_pred = p_pred > 0.5
print('fraction of predicted labels that agree ', np.sum(y_pred_np == y_pred)/len(y_pred))
# + [markdown] id="Jn1sZgoJ0d7s"
# # Modules
#
# We can define logistic regression as multilayer perceptron (MLP) with no hidden layers. This can be defined as a sequential neural network module. Modules hide the parameters inside each layer, which makes it easy to construct complex models, as we will see later on.
#
# + [markdown] id="DN7AA9V_lm9W"
# ## Sequential model
# + colab={"base_uri": "https://localhost:8080/"} id="fjF4RwWWe3-g" outputId="c4a911a2-9810-4929-f77d-e1e118733509"
# Make an MLP with no hidden layers
model = nn.Sequential(
nn.Linear(D, 1, bias=False),
nn.Sigmoid()
)
print(model)
print(model[0].weight)
print(model[0].bias)
# + colab={"base_uri": "https://localhost:8080/"} id="Oie5FZnThX1B" outputId="91bec64e-ff8b-4c79-c74b-a284f6b19b36"
# We set the parameters of the MLP by hand to match sklearn.
# Torch linear layer computes X*W' + b (see https://pytorch.org/docs/stable/generated/torch.nn.Linear.html)
# where X is N*Din, so W must be Dout*Din. Here Dout=1.
print(model[0].weight.shape)
print(w_np.shape)
w = np.reshape(w_np, [-1, 1]).transpose()
print(w.shape)
model[0].weight = nn.Parameter(torch.Tensor(w))
print(model[0].weight.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="simLA1V0fz4Y" outputId="4ed01d9d-9246-4ad4-e088-f30823fe07ab"
p_pred_np = predict_prob(w_np, X_test)
p_pred_t = model(X_test_t).detach().numpy()[:,0]
print(p_pred_np)
print(p_pred_t)
assert np.allclose(p_pred_np, p_pred_t)
# + colab={"base_uri": "https://localhost:8080/"} id="1K60WLEOl-_3" outputId="bee54e14-5b0f-459b-f794-09dcdfadb216"
# we can assign names to each layer in the sequence
from collections import OrderedDict
model = nn.Sequential(OrderedDict([
('linear_layer', nn.Linear(D, 1, bias=False)),
('output_activation', nn.Sigmoid())
])
)
print(model)
print(model.linear_layer)
print(model.linear_layer.weight)
print(model.output_activation)
# + colab={"base_uri": "https://localhost:8080/"} id="c-O4sR1zmpn2" outputId="6ca16746-d4b3-443c-aaa8-431bdf739810"
# some layers define adjustable parameters, which can be optimized.
# we can inspect them thus:
for name, param in model.named_parameters():
print(name, param.shape)
# + [markdown] id="MlirdZ6rlrE0"
# ## Subclass the Module class
#
# For more complex models (eg non-sequential), we can create our own subclass. We just need to define a 'forward' method that maps inputs to outputs, as we show below.
# + id="xp1y2uzD6xGD" colab={"base_uri": "https://localhost:8080/"} outputId="f68029fb-0bf9-44f0-a348-462acd329afa"
class Model(torch.nn.Module):
def __init__(self):
super(Model, self).__init__()
self.linear = torch.nn.Linear(D, 1, bias=False)
def forward(self, x):
y_pred = torch.sigmoid(self.linear(x))
return y_pred[:,0] # (N,1) -> (N)
set_seed(0)
model = Model()
w = np.reshape(w_np, [-1, 1]).transpose()
model.linear.weight = nn.Parameter(torch.Tensor(w))
p_pred_np = predict_prob(w_np, X_test)
p_pred_t = model(X_test_t) # calls model.__call__ which calls model.forward()
p_pred = p_pred_t.detach().numpy()
print(p_pred_np)
print(p_pred)
assert np.allclose(p_pred_np, p_pred)
# + [markdown] id="dZqfTc03JIV7"
# ## SGD on a module
#
# We can optimize the parameters of a module by passing a reference to them into the optimizer, as we show below.
# + colab={"base_uri": "https://localhost:8080/", "height": 435} id="1K-Suo6jHynP" outputId="0c7918c6-76fc-49f2-e1cb-5a717568a680"
nepochs = 100
nbatches = len(dataloader)
criterion = torch.nn.BCELoss(reduction='mean')
learning_rate = 1e-1
loss_trace = []
set_seed(0)
model = Model()
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
for epoch in range(nepochs):
for b, batch in enumerate(dataloader):
X, y = batch
y_pred = model(X) # predict/ forward function
loss = criterion(y_pred, y)
#print(f'epoch {epoch}, batch {b}, loss: {loss.item()}')
loss_trace.append(loss)
optimizer.zero_grad()
loss.backward()
optimizer.step()
#print(f'end of epoch {epoch}, loss: {loss.item()}')
plt.figure()
plt.plot(loss_trace)
y_pred_np = predict_prob(w_np, X_test)
y_pred_t = model(X_test_t)
y_pred = y_pred_t.detach().numpy()
print(y_pred_np)
print(y_pred)
# + [markdown] id="MGbegp5xJKSN"
# ## Batch optimization on a module
#
# SGD does not match the results of sklearn. However, this is not because of the way we defined the model, it's just because SGD is a bad optimizer. Here we show that BFGS gives exactly the same results as sklearn.
#
# + colab={"base_uri": "https://localhost:8080/"} id="5BN5X-1w62ST" outputId="a05136f2-326e-4675-da1a-8de71cffaeed"
set_seed(0)
model = Model()
optimizer = torch.optim.LBFGS(model.parameters(), history_size=10)
criterion = torch.nn.BCELoss(reduction='mean')
def closure():
optimizer.zero_grad()
y_pred = model(X_train_t)
loss = criterion(y_pred, y_train_t)
loss.backward()
return loss
max_iter = 10
loss_trace = []
for i in range(max_iter):
loss = optimizer.step(closure)
#print(loss)
y_pred_np = predict_prob(w_np, X_test)
y_pred_t = model(X_test_t)
y_pred = y_pred_t.detach().numpy()
print(y_pred_np)
print(y_pred)
# + [markdown] id="jQTxqUFg4L1W"
# # Multi-class logistic regression
#
# For binary classification problems, we can use a sigmoid as the final layer, to return probabilities. The corresponding loss is the binary cross entropy, [nn.BCELoss(pred_prob, true_label)](https://pytorch.org/docs/stable/generated/torch.nn.BCELoss.html), where pred_prob is of shape (B) with entries in [0,1], and true_label is of shape (B) with entries in 0 or 1. (Here B=batch size.) Alternatively the model can return the logit score, and use [nn.BCEWithLogitsLoss(pred_score, true_label)](https://pytorch.org/docs/stable/generated/torch.nn.BCEWithLogitsLoss.html).
#
# For multiclass classifiction, the final layer can return the log probabilities using LogSoftmax layer, combined with the negative log likelihood loss, [nn.NLLLoss(pred_log_probs, true_label)](https://pytorch.org/docs/stable/generated/torch.nn.NLLLoss.html), where pred_log_probs is of shape B*C matrix, and true_label is of shape B with entries in {0,1,..C-1}.
# (Note that the target labels are integers, not sparse one-hot vectors.)
# Alternatively, we can just return the vector of logit scores, and use [nn.CrossEntropyLoss(logits, true_label)](https://pytorch.org/docs/stable/generated/torch.nn.CrossEntropyLoss.html). The above two methods should give the same results.
#
# + id="j7g6aFCD7KI5"
# code me
| notebooks/logreg_pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
import keras
import numpy as np
np.random.seed(123)
# ## Preprocessing Data
def get_mnist_data():
from keras.datasets import mnist
from keras.utils import np_utils
(X_train, y_train), (X_test, y_test) = mnist.load_data()
X_train = X_train.reshape(X_train.shape[0], 1, X_train.shape[1], X_train.shape[2])
y_train = y_train.reshape(y_train.shape[0], 1)
X_test = X_test.reshape(X_test.shape[0], 1, X_test.shape[1], X_test.shape[2])
y_test = y_test.reshape(y_test.shape[0], 1)
y_train = np_utils.to_categorical(y_train)
y_test = np_utils.to_categorical(y_test)
return X_train, y_train, X_test, y_test
X_train, y_train, X_test, y_test = get_mnist_data()
X_train.shape, y_train.shape, X_test.shape, y_test.shape
X_mean = X_train.mean().astype(np.float32)
X_std = X_train.std().astype(np.float32)
def normalizer(x):
return (x - X_mean) / X_std
# ## Building the model
from keras.layers import Convolution2D, Dense, Flatten, Lambda, Dropout
from keras.models import Sequential
from keras.optimizers import Adam
# ### Linear Model
nb_epoch = 5
model = Sequential([Lambda(normalizer, input_shape=(1, 28, 28)), Flatten(), Dense(10, activation='softmax')])
model.compile(Adam(), loss='categorical_crossentropy', metrics=['accuracy'] )
model.fit(X_train, y_train,validation_data=(X_test, y_test), nb_epoch=nb_epoch)
# ### Is learning rate optimal?
#
# default in Adam is 0.001
model.optimizer.lr = 0.1
model.fit(X_train, y_train,validation_data=(X_test, y_test), nb_epoch=nb_epoch)
model.optimizer.lr = 0.0001
model.fit(X_train, y_train,validation_data=(X_test, y_test), nb_epoch=15)
# Looks like learning rate is already good enough at 0.001 (default value)
| mnist/Linear Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## TIS notebook
# Notebook settings and importing packages
# +
# To support both python 2 and python 3
from __future__ import division, print_function, unicode_literals
# Common imports
import numpy as np
import os
import pandas as pd
# To plot pretty figures
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# to make this notebook's output stable across runs
np.random.seed(42)
# -
# Read data from file 'tis_data.csv'
# (in the same directory that your python process is based)
datatis = pd.read_csv("raw_v2.csv")
# Preview the first 10 lines of the loaded data
datatis.head(10)
# Import train_test_split function
from sklearn.model_selection import train_test_split
x = datatis[['WI','DT','HC','RP','OS']]
y = datatis['Class'] # Labels
# Split dataset into training set and test set
# +
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.3) # 70% training and 30% test
daisplay(y_train)
daisplay(y_test)
# -
# ## Random Forest Model
# Train the model using the training sets and import confusion matrix
from sklearn.metrics import confusion_matrix
from sklearn.ensemble import RandomForestClassifier
# Create a Gaussian Classifier
forest_clf = RandomForestClassifier(bootstrap=True, class_weight=None, criterion='gini',
max_depth=None, max_features='auto', max_leaf_nodes=None,
min_impurity_decrease=0.0, min_impurity_split=None,
min_samples_leaf=1, min_samples_split=2,
min_weight_fraction_leaf=0.0, n_estimators=50, n_jobs=-1,
oob_score=False, random_state=None, verbose=0,
warm_start=False)
# Train the model using cross validation
# +
from sklearn.model_selection import cross_val_score, cross_val_predict
y_pred_cross = cross_val_predict(forest_clf, x_train, y_train, cv=10)
#cross_val_score(forest_clf, x_train, y_train, cv=10)
conf_mx_cross = confusion_matrix(y_train, y_pred_cross)
# -
# Import scikit-learn metrics module for accuracy calculation
# + [markdown] toc-hr-collapsed=false
# # Performance Measures
# -
# Accuracy, Percision, Recall, F1 Score and Confusion Matrix
from sklearn.metrics import recall_score, precision_score, f1_score, confusion_matrix, accuracy_score
print("Accuracy:",accuracy_score(y_test, y_pred_cross))
#print("Percision:",precision_score(y_test,y_pred_cross, average='micro'))
#print("Recall:",recall_score(y_test,y_pred_cross, average='micro'))
#print("F1 score:",f1_score(y_test,y_pred_cross, average='micro'))
#print("confusion Matrix:",confusion_matrix(y_test,y_pred_cross))
# Feature Importance
#feature_imp = pd.Series(forest_clf.feature_importances_,index=datatis.feature_names).sort_values(ascending=False)
#print(feature_imp)
print(forest_clf.feature_importances_)
# Plot confusion matrix
plt.matshow(conf_mx_cross, cmap=plt.cm.gray)
plt.show()
# + [markdown] toc-hr-collapsed=false
# ## MLPClassifier
# +
from sklearn.neural_network import MLPClassifier
MLP = MLPClassifier(activation= 'relu', random_state= 8, hidden_layer_sizes= (50),
learning_rate_init= 0.05, learning_rate= 'invscaling', momentum= 0.1, max_iter= 50, verbose= 0)
# activation= relu & logistic & tanh & identity, relu: y=0 (x<0) & y=x (x=>0), hidden_layer_sizes: number of neurons in each layer
# learning_rate_init: initial learning rate (<0.2), learning_rate: constant or decreasing learning rate
# verbose: see training process
scores = cross_val_score(MLP, X, Y, cv= 10, scoring= 'f1')
#scores.mean()
#y_pred=MLP.predict(X_test)
# -
# Import more libraries
import itertools
import numpy as np
import matplotlib.pyplot as plt
# +
from sklearn.metrics import confusion_matrix
class_names = datatis.target_names
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.tight_layout()
# -
# Compute confusion matrix
cnf_matrix = confusion_matrix(y_test, y_pred)
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=class_names,
title='Confusion matrix, without normalization')
# Plot normalized confusion matrix
# +
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=class_names, normalize=True,
title='Normalized confusion matrix')
plt.show()
| tis_prediction/tis_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_csv("InsideAirbnbDataUrls.csv")
df
url = "http://data.insideairbnb.com/united-states/ca/los-angeles/2020-06-10/data/listings.csv.gz"
url.split("/")
def generate_country_data(url):
return url.split("/")[3]
df["country"] = df["url"].apply(generate_country_data)
df
def generate_state_data(url):
return url.split("/")[4]
df["state"] = df["url"].apply(generate_state_data)
df
def generate_date_data(url):
return url.split("/")[6]
df["date"] = df["url"].apply(generate_date_data)
df
def generate_city_data(url):
return url.split("/")[5]
df["city"] = df["url"].apply(generate_city_data)
df
df = df[['country', 'state', 'city','date', 'url']]
df
df.to_csv("urls", index=False)
| airflow/utils/ConvertUrls.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from functools import partial
from multiprocessing import Pool, cpu_count
ids_tsl = np.load("../data/misc/glacier_IDs.npy", allow_pickle=True).tolist()
tsl_path = "../results/tsl_csv/"
meteo_path = "../data/FIT_forcing/meteo/"
def read_tsl(rgi_id, path=tsl_path):
tsl = pd.read_csv(f"{tsl_path}{rgi_id}.csv", index_col=0, parse_dates=True)
return tsl
def read_meteo(rgi_id, path=meteo_path):
meteo = pd.read_hdf(f"{meteo_path}{rgi_id}.h5")
return meteo
def create_features(dataframe, back_to=12):
# convert circular wind_dir_mean
# to two components of cos() and sin()
# source: https://stats.stackexchange.com/questions/336613/
# regression-using-circular-variable-hour-from-023-as-predictor
# copy for safety
df = dataframe.copy()
# create cos() and sin() components
df["wind_dir_mean_cos"] = np.cos(np.deg2rad(df["wind_dir_mean"]))
df["wind_dir_mean_sin"] = np.sin(np.deg2rad(df["wind_dir_mean"]))
# drop "wind_dir_mean"
df = df.drop(["wind_dir_mean"], axis=1)
# make shifts and rolling means
cols = df.columns
for col in cols:
for shift in range(1, back_to+1, 1):
df["{}-{}".format(col, shift)] = df[col].shift(shift).values
for rol in range(1, back_to+1, 1):
df["{}rol-{}".format(col, rol)] = df[col].rolling(window=rol).mean().values
# delete NaNs
df = df.dropna()
return df
def dataset_construction(freq, subset_jjas, rgi_id):
# get raw TSL measurements
tsl = read_tsl(rgi_id)
# resample to specific frequency
tsl_resample = tsl.resample(freq).mean()
# get raw ERA5-Land forcing
meteo = read_meteo(rgi_id)
# resample to specific frequency
meteo_resample = pd.DataFrame({'t2m_min': meteo['t2m_min'].resample(freq).min(),
't2m_max': meteo['t2m_max'].resample(freq).max(),
't2m_mean': meteo['t2m_mean'].resample(freq).mean(),
'tp': meteo['tp'].resample(freq).sum(),
'sf': meteo['sf'].resample(freq).sum(),
'ssrd': meteo['ssrd'].resample(freq).sum(),
'strd': meteo['strd_mean'].resample(freq).sum(),
'wind_max': meteo['wind_max'].resample(freq).max(),
'wind_mean': meteo['wind_mean'].resample(freq).mean(),
'wind_dir_mean': meteo['wind_dir_mean'].resample(freq).mean(),
'tcc': meteo['tcc'].resample(freq).mean()})
# enrich meteo features
if freq == "M":
meteo_enrich = create_features(meteo_resample, back_to=12)
elif freq == "W":
meteo_enrich = create_features(meteo_resample, back_to=48) #12 months back considering 4 weeks in each month
# merge datasets
dataset = pd.concat([tsl_resample, meteo_enrich], axis=1)
# drop NaNs
dataset = dataset.dropna()
if subset_jjas:
dataset = dataset[(dataset.index.month == 6) | (dataset.index.month == 7) |
(dataset.index.month == 8) | (dataset.index.month == 9)]
if freq == "M":
freq_prefix = "monthly"
elif freq == "W":
freq_prefix = "weekly"
if subset_jjas:
subset_prefix = "JJAS"
else:
subset_prefix = "full"
dataset.to_csv(f"../results/data4ml/{freq_prefix}_{subset_prefix}/{rgi_id}.csv", compression="gzip")
#print(rgi_id)
return rgi_id
def detect_no_tsl_data(rgi_id):
tsl = read_tsl(rgi_id)
try:
tsl = tsl.resample("M").mean()
except:
print(rgi_id)
return rgi_id
# +
# %%time
p = Pool(cpu_count())
no_tsl_data = list(p.imap(detect_no_tsl_data, ids_tsl))
p.close()
p.join()
# -
no_tsl_data = [i for i in no_tsl_data if i is not None]
len(no_tsl_data)
ids_tsl_valid = [i for i in ids_tsl if i not in no_tsl_data]
len(ids_tsl_valid)
# +
# monthly, full data
p = Pool(cpu_count())
freq="M"
subset_jjas = False
func = partial(dataset_construction, freq, subset_jjas)
saved = list(p.imap(func, ids_tsl_valid))
p.close()
p.join()
print(len(saved))
# +
# monthly, JJAS data
p = Pool(cpu_count())
freq="M"
subset_jjas = True
func = partial(dataset_construction, freq, subset_jjas)
saved = list(p.map(func, ids_tsl_valid))
p.close()
p.join()
print(len(saved))
# +
# weekly, full data
p = Pool(cpu_count())
freq="W"
subset_jjas = False
func = partial(dataset_construction, freq, subset_jjas)
saved = list(p.map(func, ids_tsl_valid))
p.close()
p.join()
print(len(saved))
# +
# weekly, JJAS data
p = Pool(cpu_count())
freq="W"
subset_jjas = True
func = partial(dataset_construction, freq, subset_jjas)
saved = list(p.map(func, ids_tsl_valid))
p.close()
p.join()
print(len(saved))
# -
| code/examples/17_TSL_ERA5_data_for_modelling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1-4.1 Intro Python
# ## Conditionals
# - **`if`, `else`, `pass`**
# - **Conditionals using Boolean String Methods**
# - Comparison operators
# - String comparisons
#
# -----
#
# ><font size="5" color="#00A0B2" face="verdana"> <B>Student will be able to</B></font>
# - **control code flow with `if`... `else` conditional logic**
# - **using Boolean string methods (`.isupper(), .isalpha(), startswith()...`)**
# - using comparison (`>, <, >=, <=, ==, !=`)
# - using Strings in comparisons
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
# ## Conditionals use `True` or `False`
# - **`if`**
# - **`else`**
# - **`pass`**
#
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/c53fdb30-b2b0-4183-9686-64b0e5b46dd2/Unit1_Section4.1-Conditionals.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/c53fdb30-b2b0-4183-9686-64b0e5b46dd2/Unit1_Section4.1-Conditonals.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
if True:
print("True means do something")
else:
print("Not True means do something else")
# +
hot_tea = True
if hot_tea:
print("enjoy some hot tea!")
else:
print("enjoy some tea, and perhaps try hot tea next time.")
# -
someone_i_know = False
if someone_i_know:
print("how have you been?")
else:
# use pass if there is no need to execute code
pass
# changed the value of someone_i_know
someone_i_know = True
if someone_i_know:
print("how have you been?")
else:
pass
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 1</B></font>
#
# ## Conditionals
# ### Using Boolean with `if, else`
#
# - **Give a weather report using `if, else`**
sunny_today = True
# [ ] test if it is sunny_today and give proper responses using if and else
if sunny_today:
print("It is sunny today!")
else:
print("It is not sunny today!")
sunny_today = False
# [ ] use code you created above and test sunny_today = False
if sunny_today:
print("It is sunny today!")
else:
print("It is not sunny today!")
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
# ## Conditionals: Boolean String test methods with `if`
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/caa56256-733a-4172-96f7-9ecfc12d49d0/Unit1_Section4.1-conditionals-bool.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/caa56256-733a-4172-96f7-9ecfc12d49d0/Unit1_Section4.1-conditonals-bool.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# ```python
# if student_name.isalpha():
# ```
# - **`.isalnum()`**
# - **`.istitle()`**
# - **`.isdigit()`**
# - **`.islower()`**
# - **`.startswith()`**
#
# ###
# <font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
# +
# review code and run cell
favorite_book = input("Enter the title of a favorite book: ")
if favorite_book.istitle():
print(favorite_book, "- nice capitalization in that title!")
else:
print(favorite_book, "- consider capitalization throughout for book titles.")
# +
# review code and run cell
a_number = input("enter a positive integer number: ")
if a_number.isdigit():
print(a_number, "is a positive integer")
else:
print(a_number, "is not a positive integer")
# another if
if a_number.isalpha():
print(a_number, "is more like a word")
else:
pass
# +
# review code and run cell
vehicle_type = input('"enter a type of vehicle that starts with "P": ')
if vehicle_type.upper().startswith("P"):
print(vehicle_type, 'starts with "P"')
else:
print(vehicle_type, 'does not start with "P"')
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 2: multi-part</B></font>
#
# ## Evaluating Boolean Conditionals
# ### create evaluations for `.islower()`
# - print output describing **if** each of the 2 strings is all lower or not
#
test_string_1 = "welcome"
test_string_2 = "I have $3"
# [ ] use if, else to test for islower() for the 2 strings
if test_string_1.islower():
print("String 1 is lowercase.")
else:
print("String 1 is not lowercase.")
# Test the second string
if test_string_2.islower():
print("String 2 is lowercase.")
else:
print("String 2 is not lowercase.")
# <font size="3" color="#B24C00" face="verdana"> <B>Task 2 continued.. </B></font>
# ### create a functions using `startswith('w')`
# - w_start_test() tests if starts with "w"
# **function should have a parameter for `test_string` and print the test result**
test_string_1 = "welcome"
test_string_2 = "I have $3"
test_string_3 = "With a function it's efficient to repeat code"
# [ ] create a function w_start_test() use if & else to test with startswith('w')
def w_start_test(string):
if string.startswith('w'):
print("String starts with a 'w'")
else:
print("String does not start with a 'w'")
# [ ] Test the 3 string variables provided by calling w_start_test()
w_start_test(test_string_1)
w_start_test(test_string_2)
w_start_test(test_string_3)
# [Terms of use](http://go.microsoft.com/fwlink/?LinkID=206977) [Privacy & cookies](https://go.microsoft.com/fwlink/?LinkId=521839) © 2017 Microsoft
| Python Absolute Beginner/Module_3_1_Absolute_Beginner.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="jaT3om3nMpml" colab_type="text"
# # Textworld starting kit notebook
#
# Model: *LSTM-DQN*
#
# When running first:
# 1. Run the first 2 code cells(with pip installations)
# 2. Restart runtime
# 3. Continue with the next cells
#
# This is done, because there is a problem with dependencies of **textworld** and **colab**, requiring different versions of **prompt-toolkit**
# + [markdown] id="-QVq2c31UByC" colab_type="text"
# ## Todo
# ### RL:
# * [x] Prioritized Replay Memory
# * [x] [N-step DQN](https://www.groundai.com/project/understanding-multi-step-deep-reinforcement-learning-a-systematic-study-of-the-dqn-target/)
# * [x] [Fixed Q-targets](https://www.freecodecamp.org/news/improvements-in-deep-q-learning-dueling-double-dqn-prioritized-experience-replay-and-fixed-58b130cc5682/)
# * [x] [Double DQN](https://towardsdatascience.com/deep-double-q-learning-7fca410b193a)
# * [ ] Dueling DQN
# * [ ] Multiple inputs (description, inventory, quests, etc)
# * [x] Replay memory sample, when not having any alpha/beta priority samples, should take the whole sample of the opposite priority.
# * [ ] Noisy nets
# * [ ] DRQN ?
# * [ ] [Rainbow Paper](https://arxiv.org/pdf/1710.02298.pdf) ?
#
# ### NLP:
# * [x] Normalize tokens
# * [x] Add l2 regularization
# * [ ] Remove apostrophes from descriptions
#
# ### Game Env:
# * [x] Train with simple games in starting kit
# * [ ] Check game generation and complexity
# * [ ] Make many simple(easy) games, each of which teaches different skill.
# * [ ] Train first with simple games, and progressively train on more complex games.
#
# ### Debugging:
# * [x] Extended log for checking steps and scores on every epoch
# * [ ] Graphs with speed, reward, epsilon movement, loss function, episode length (use Tensorboard or similar)
# * [ ] Model comparisson
#
# ### Colab:
# * [x] Export model parameters to Drive
#
# + [markdown] id="oBFYrJbqjScT" colab_type="text"
# ## Setup
# + id="2-XUxa37Z1S5" colab_type="code" outputId="ac29c111-e70e-4af8-b951-74174d148502" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !pip install textworld
# + id="XPpv7-6bVOsP" colab_type="code" outputId="e7fa9a4c-e2e3-4237-e779-1cf4262f9c70" colab={"base_uri": "https://localhost:8080/", "height": 68}
# !pip install prompt-toolkit==1.0.16
# + id="Oqz6G9llwXOc" colab_type="code" outputId="98cf2593-c8f3-41b2-f634-9a5597e022a1" colab={"base_uri": "https://localhost:8080/", "height": 51}
# !pip install -U -q PyDrive
# + id="feZKkDlnQIZz" colab_type="code" outputId="62ce66f2-4c58-433c-a345-d80ad50869d4" colab={"base_uri": "https://localhost:8080/", "height": 476}
# !pip install pytorch_pretrained_bert
# + id="LuCM9FzGl2Zk" colab_type="code" outputId="7ffb839e-d507-4962-8e78-00af9e9247ca" colab={"base_uri": "https://localhost:8080/", "height": 34}
import os
import random
import logging
import yaml
import copy
import spacy
import numpy as np
import glob
from tqdm import tqdm
from typing import List, Dict, Any
from collections import namedtuple
import pandas as pd
import torch
import torch.nn.functional as F
import gym
import textworld.gym
from textworld import EnvInfos
import random
from pytorch_pretrained_bert import BertTokenizer, BertModel
torch.cuda.is_available()
# + [markdown] id="GWGX1rdFtIgW" colab_type="text"
# ## Generic functions
# + id="7zO94P3hjyH_" colab_type="code" colab={}
def to_np(x):
if isinstance(x, np.ndarray):
return x
return x.data.cpu().numpy()
def to_pt(np_matrix, enable_cuda=False, type='long'):
if type == 'long':
if enable_cuda:
return torch.autograd.Variable(torch.from_numpy(np_matrix).type(torch.LongTensor).cuda())
else:
return torch.autograd.Variable(torch.from_numpy(np_matrix).type(torch.LongTensor))
elif type == 'float':
if enable_cuda:
return torch.autograd.Variable(torch.from_numpy(np_matrix).type(torch.FloatTensor).cuda())
else:
return torch.autograd.Variable(torch.from_numpy(np_matrix).type(torch.FloatTensor))
def _words_to_ids(words, word2id):
ids = []
for word in words:
try:
ids.append(word2id[word])
except KeyError:
ids.append(1)
return ids
def preproc(s, str_type='None', tokenizer=None, lower_case=True):
if s is None:
return ["nothing"]
s = s.replace("\n", ' ')
if s.strip() == "":
return ["nothing"]
if str_type == 'feedback':
if "$$$$$$$" in s:
s = ""
if "-=" in s:
s = s.split("-=")[0]
s = s.strip()
if len(s) == 0:
return ["nothing"]
tokens = [t.text for t in tokenizer(s)]
# NORMALIZE WORDS
#tokens = [t.norm_ for t in tokenizer(s)]
if lower_case:
tokens = [t.lower() for t in tokens]
return tokens
def max_len(list_of_list):
return max(map(len, list_of_list))
def pad_sequences(sequences, maxlen=None, dtype='int32', value=0.):
'''
Partially borrowed from Keras
# Arguments
sequences: list of lists where each element is a sequence
maxlen: int, maximum length
dtype: type to cast the resulting sequence.
value: float, value to pad the sequences to the desired value.
# Returns
x: numpy array with dimensions (number_of_sequences, maxlen)
'''
lengths = [len(s) for s in sequences]
nb_samples = len(sequences)
if maxlen is None:
maxlen = np.max(lengths)
# take the sample shape from the first non empty sequence
# checking for consistency in the main loop below.
sample_shape = tuple()
for s in sequences:
if len(s) > 0:
sample_shape = np.asarray(s).shape[1:]
break
x = (np.ones((nb_samples, maxlen) + sample_shape) * value).astype(dtype)
for idx, s in enumerate(sequences):
if len(s) == 0:
continue # empty list was found
# pre truncating
trunc = s[-maxlen:]
# check `trunc` has expected shape
trunc = np.asarray(trunc, dtype=dtype)
if trunc.shape[1:] != sample_shape:
raise ValueError('Shape of sample %s of sequence at position %s is different from expected shape %s' %
(trunc.shape[1:], idx, sample_shape))
# post padding
x[idx, :len(trunc)] = trunc
return x
def freeze_layer(layer):
for param in layer.parameters():
param.requires_grad = False
def preproc_example(s):
s = s.replace('$', '')
s = s.replace('#', '')
s = s.replace('\n', ' ')
s = s.replace(' ', ' ')
s = s.replace('_', '')
s = s.replace('|', '')
s = s.replace('\\', '')
s = s.replace('/', '')
s = s.replace('-', '')
s = s.replace('=', '')
return s
def convert_examples_to_features(sequences, tokenizer):
"""Loads a data file into a list of `InputFeature`s."""
batch_tokens = []
batch_input_ids = []
batch_input_masks = []
for example in sequences:
_example = preproc_example(example)
# print(_example)
tokens = tokenizer.tokenize(_example)
batch_tokens.append(tokens)
del _example
del tokens
max_length = max([len(x) for x in batch_tokens])
# print('bert_max_seqence', max_length)
for tokens in batch_tokens:
input_ids = tokenizer.convert_tokens_to_ids(tokens)
# The mask has 1 for real tokens and 0 for padding tokens. Only real
# tokens are attended to.
input_mask = [1] * len(input_ids)
# Zero-pad up to the sequence length.
while len(input_ids) < max_length:
input_ids.append(0)
input_mask.append(0)
batch_input_ids.append(input_ids)
batch_input_masks.append(input_mask)
del input_ids
del input_mask
return batch_tokens, batch_input_ids, batch_input_masks
# + [markdown] id="6HTSoK93tM6V" colab_type="text"
# ## Layers
# + id="K2VSAkD3s7Ke" colab_type="code" colab={}
def masked_mean(x, m=None, dim=-1):
"""
mean pooling when there're paddings
input: tensor: batch x time x h
mask: batch x time
output: tensor: batch x h
"""
if m is None:
return torch.mean(x, dim=dim)
mask_sum = torch.sum(m, dim=-1) # batch
res = torch.sum(x, dim=1) # batch x h
res = res / (mask_sum.unsqueeze(-1) + 1e-6)
return res
class FastUniLSTM(torch.nn.Module):
"""
Adapted from https://github.com/facebookresearch/DrQA/
now supports: different rnn size for each layer
all zero rows in batch (from time distributed layer, by reshaping certain dimension)
"""
def __init__(self, ninp, nhids, bidir=False, dropout_between_rnn_layers=0.):
super(FastUniLSTM, self).__init__()
self.ninp = ninp
self.nhids = nhids
self.nlayers = len(self.nhids)
self.dropout_between_rnn_layers = dropout_between_rnn_layers
self.stack_rnns(bidir)
def stack_rnns(self, bidir):
rnns = [torch.nn.LSTM(self.ninp if i == 0 else self.nhids[i - 1],
self.nhids[i],
num_layers=1,
bidirectional=bidir) for i in range(self.nlayers)]
self.rnns = torch.nn.ModuleList(rnns)
def forward(self, x, mask):
def pad_(tensor, n):
if n > 0:
zero_pad = torch.autograd.Variable(torch.zeros((n,) + tensor.size()[1:]))
if x.is_cuda:
zero_pad = zero_pad.cuda()
tensor = torch.cat([tensor, zero_pad])
return tensor
"""
inputs: x: batch x time x inp
mask: batch x time
output: encoding: batch x time x hidden[-1]
"""
# Compute sorted sequence lengths
batch_size = x.size(0)
lengths = mask.data.eq(1).long().sum(1) # .squeeze()
_, idx_sort = torch.sort(lengths, dim=0, descending=True)
_, idx_unsort = torch.sort(idx_sort, dim=0)
lengths = list(lengths[idx_sort])
idx_sort = torch.autograd.Variable(idx_sort)
idx_unsort = torch.autograd.Variable(idx_unsort)
# Sort x
x = x.index_select(0, idx_sort)
# remove non-zero rows, and remember how many zeros
n_nonzero = np.count_nonzero(lengths)
n_zero = batch_size - n_nonzero
if n_zero != 0:
lengths = lengths[:n_nonzero]
x = x[:n_nonzero]
# Transpose batch and sequence dims
x = x.transpose(0, 1)
# Pack it up
rnn_input = torch.nn.utils.rnn.pack_padded_sequence(x, lengths)
# Encode all layers
outputs = [rnn_input]
for i in range(self.nlayers):
rnn_input = outputs[-1]
# dropout between rnn layers
if self.dropout_between_rnn_layers > 0:
dropout_input = F.dropout(rnn_input.data,
p=self.dropout_between_rnn_layers,
training=self.training)
rnn_input = torch.nn.utils.rnn.PackedSequence(dropout_input,
rnn_input.batch_sizes)
seq, last = self.rnns[i](rnn_input)
outputs.append(seq)
if i == self.nlayers - 1:
# last layer
last_state = last[0] # (num_layers * num_directions, batch, hidden_size)
last_state = last_state[0] # batch x hidden_size
# Unpack everything
for i, o in enumerate(outputs[1:], 1):
outputs[i] = torch.nn.utils.rnn.pad_packed_sequence(o)[0]
output = outputs[-1]
# Transpose and unsort
output = output.transpose(0, 1) # batch x time x enc
# re-padding
output = pad_(output, n_zero)
last_state = pad_(last_state, n_zero)
output = output.index_select(0, idx_unsort)
last_state = last_state.index_select(0, idx_unsort)
# Pad up to original batch sequence length
if output.size(1) != mask.size(1):
padding = torch.zeros(output.size(0),
mask.size(1) - output.size(1),
output.size(2)).type(output.data.type())
output = torch.cat([output, torch.autograd.Variable(padding)], 1)
output = output.contiguous() * mask.unsqueeze(-1)
return output, last_state, mask
# + [markdown] id="l0NRD6XDtRKa" colab_type="text"
# ## Model
# + id="vd4txzFctSFi" colab_type="code" colab={}
class LSTM_DQN(torch.nn.Module):
model_name = 'lstm_dqn'
def __init__(self, model_config, embedding_weights, word_vocab, generate_length=5, enable_cuda=False):
super(LSTM_DQN, self).__init__()
self.model_config = model_config
self.enable_cuda = enable_cuda
self.word_vocab_size = len(word_vocab)
self.id2word = word_vocab
self.generate_length = generate_length
self.read_config()
self._def_layers(embedding_weights)
self.init_weights()
self.print_parameters()
def print_parameters(self):
# print(self)
amount = 0
for p in self.parameters():
amount += np.prod(p.size())
print("Total number of parameters: {}".format(amount))
parameters = filter(lambda p: p.requires_grad, self.parameters())
amount = 0
for p in parameters:
amount += np.prod(p.size())
print("Number of trainable parameters: {}".format(amount))
def read_config(self):
# model config
self.freeze_embedding = self.model_config['freeze_embedding']
self.embedding_size = self.model_config['embedding_size']
self.encoder_rnn_hidden_size = self.model_config['encoder_rnn_hidden_size']
self.action_scorer_hidden_dim = self.model_config['action_scorer_hidden_dim']
self.dropout_between_rnn_layers = self.model_config['dropout_between_rnn_layers']
self.bidirectional_lstm = self.model_config['bidirectional_lstm']
def _def_layers(self, embedding_weights=None):
# word embeddings
self.word_embedding = Embedding(embedding_size=self.embedding_size,
vocab_size=self.word_vocab_size,
enable_cuda=self.enable_cuda)
if not(embedding_weights is None):
self.word_embedding.set_weights(embedding_weights)
print("Embedding imported!")
if self.freeze_embedding:
freeze_layer(self.word_embedding.embedding_layer)
print("Embedding freezed!")
# lstm encoder
self.encoder = FastUniLSTM(ninp=self.embedding_size,
nhids=self.encoder_rnn_hidden_size,
bidir=self.bidirectional_lstm,
dropout_between_rnn_layers=self.dropout_between_rnn_layers)
self.action_scorer_shared = torch.nn.Linear(self.encoder_rnn_hidden_size[-1], self.action_scorer_hidden_dim)
action_scorers = []
for _ in range(self.generate_length):
action_scorers.append(torch.nn.Linear(self.action_scorer_hidden_dim, self.word_vocab_size, bias=False))
self.action_scorers = torch.nn.ModuleList(action_scorers)
self.fake_recurrent_mask = None
def init_weights(self):
torch.nn.init.xavier_uniform_(self.action_scorer_shared.weight.data)
for i in range(len(self.action_scorers)):
torch.nn.init.xavier_uniform_(self.action_scorers[i].weight.data)
self.action_scorer_shared.bias.data.fill_(0)
def representation_generator(self, _input_words):
embeddings, mask = self.word_embedding.forward(_input_words) # batch x time x emb
encoding_sequence, _, _ = self.encoder.forward(embeddings, mask) # batch x time x h
mean_encoding = masked_mean(encoding_sequence, mask) # batch x h
return mean_encoding
def action_scorer(self, state_representation):
hidden = self.action_scorer_shared.forward(state_representation) # batch x hid
hidden = F.relu(hidden) # batch x hid
action_ranks = []
for i in range(len(self.action_scorers)):
action_ranks.append(self.action_scorers[i].forward(hidden)) # batch x n_vocab
return action_ranks
# + id="SoRlncyh_U5L" colab_type="code" colab={}
logger = logging.getLogger(__name__)
class Bert_DQN(torch.nn.Module):
model_name = 'bert_dqn'
def __init__(self, model_config, word_vocab, generate_length=5, enable_cuda=False):
super(Bert_DQN, self).__init__()
self.model_config = model_config
self.enable_cuda = enable_cuda
self.word_vocab_size = len(word_vocab)
self.id2word = word_vocab
self.generate_length = generate_length
self.read_config()
# print(enable_cuda)
self.device = torch.device("cuda" if enable_cuda else "cpu")
self.tokenizer = BertTokenizer.from_pretrained(self.bert_model, do_lower_case=True)
self._def_layers()
self.init_weights()
def print_parameters(self):
# print(self)
amount = 0
for p in self.parameters():
amount += np.prod(p.size())
print("total number of parameters: %s" % (amount))
parameters = filter(lambda p: p.requires_grad, self.parameters())
amount = 0
for p in parameters:
amount += np.prod(p.size())
print("number of trainable parameters: %s" % (amount))
def read_config(self):
# model config
# self.embedding_size = self.model_config['embedding_size']
# self.encoder_rnn_hidden_size = self.model_config['encoder_rnn_hidden_size']
# self.action_scorer_hidden_dim = self.model_config['action_scorer_hidden_dim']
# self.dropout_between_rnn_layers = self.model_config['dropout_between_rnn_layers']
self.bert_model = self.model_config['bert_model']
self.action_scorer_hidden_dim = self.model_config['action_scorer_hidden_dim']
self.train_bert = self.model_config['train_bert']
def _def_layers(self):
# word embeddings
# self.word_embedding = Embedding(embedding_size=self.embedding_size,
# vocab_size=self.word_vocab_size,
# enable_cuda=self.enable_cuda)
# # lstm encoder
# self.encoder = FastUniLSTM(ninp=self.embedding_size,
# nhids=self.encoder_rnn_hidden_size,
# dropout_between_rnn_layers=self.dropout_
self.encoder = BertModel.from_pretrained(self.bert_model).to(self.device)
if not self.train_bert:
freeze_layer(self.encoder)
# only for base models
# for large models is
bert_embeddings = 768
self.action_scorer_shared = torch.nn.Linear(bert_embeddings, self.action_scorer_hidden_dim)
action_scorers = []
for _ in range(self.generate_length):
action_scorers.append(torch.nn.Linear(self.action_scorer_hidden_dim, self.word_vocab_size, bias=False))
self.action_scorers = torch.nn.ModuleList(action_scorers)
self.fake_recurrent_mask = None
def init_weights(self):
torch.nn.init.xavier_uniform_(self.action_scorer_shared.weight.data)
for i in range(len(self.action_scorers)):
torch.nn.init.xavier_uniform_(self.action_scorers[i].weight.data)
self.action_scorer_shared.bias.data.fill_(0)
def representation_generator(self, ids, mask):
ids = ids.to(self.device)
mask = mask.to(self.device)
layers, _ = self.encoder(ids, attention_mask=mask)
# encoding_sequence = layers[self.layer_index]
# print('layer length: ', len(layers))
encoding_sequence = layers[-2].type(torch.FloatTensor)
encoding_sequence = encoding_sequence.to(self.device)
# print('encoding_sequence: ', type(encoding_sequence))
# print('encoding_sequence: ', encoding_sequence)
mask = mask.type(torch.FloatTensor).to(self.device)
# print('mask: ', type(mask))
# print('mask: ', mask)
# embeddings, mask = self.word_embedding.forward(_input_words) # batch x time x emb
# encoding_sequence, _, _ = self.encoder.forward(embeddings, mask) # batch x time x h
res_mean = masked_mean(encoding_sequence, mask) # batch x h
del layers
del encoding_sequence
del mask
return res_mean
def action_scorer(self, state_representation):
hidden = self.action_scorer_shared.forward(state_representation) # batch x hid
hidden = F.relu(hidden) # batch x hid
action_ranks = []
for i in range(len(self.action_scorers)):
action_ranks.append(self.action_scorers[i].forward(hidden)) # batch x n_vocab
del hidden
return action_ranks
# + [markdown] id="2vJ6KiC6nYA1" colab_type="text"
# Cache score
# + id="75-I610unU8W" colab_type="code" colab={}
class HistoryScoreCache(object):
def __init__(self, capacity=1):
self.capacity = capacity
self.reset()
def push(self, stuff):
"""stuff is float."""
if len(self.memory) < self.capacity:
self.memory.append(stuff)
else:
self.memory = self.memory[1:] + [stuff]
def get_avg(self):
return np.mean(np.array(self.memory))
def reset(self):
self.memory = []
def __len__(self):
return len(self.memory)
# + [markdown] id="F98HXiXKnjIH" colab_type="text"
# ## Memory
# + id="vJv3PcbJnXG2" colab_type="code" colab={}
# a snapshot of state to be stored in replay memory
Transition = namedtuple('Transition', ('bert_ids', 'bert_masks',
'word_indices',
'reward', 'mask', 'done',
'next_bert_ids', 'next_bert_masks',
'next_word_masks'))
# + id="tuUH9-QrnQxI" colab_type="code" colab={}
class PrioritizedReplayMemory(object):
def __init__(self, capacity=100000, priority_fraction=0.0):
# prioritized replay memory
self.priority_fraction = priority_fraction
self.alpha_capacity = int(capacity * priority_fraction)
self.beta_capacity = capacity - self.alpha_capacity
self.alpha_memory, self.beta_memory = [], []
self.alpha_position, self.beta_position = 0, 0
def push(self, is_prior, transition):
"""Saves a transition."""
if self.priority_fraction == 0.0:
is_prior = False
if is_prior:
if len(self.alpha_memory) < self.alpha_capacity:
self.alpha_memory.append(None)
self.alpha_memory[self.alpha_position] = transition
self.alpha_position = (self.alpha_position + 1) % self.alpha_capacity
else:
if len(self.beta_memory) < self.beta_capacity:
self.beta_memory.append(None)
self.beta_memory[self.beta_position] = transition
self.beta_position = (self.beta_position + 1) % self.beta_capacity
def sample(self, batch_size):
if self.priority_fraction == 0.0 or len(self.alpha_memory) == 0:
from_beta = min(batch_size, len(self.beta_memory))
res = random.sample(self.beta_memory, from_beta)
elif len(self.beta_memory) == 0:
from_alpha = min(batch_size, len(self.alpha_memory))
res = random.sample(self.alpha_memory, from_alpha)
else:
priority_batch = int(self.priority_fraction * batch_size)
from_alpha = min(priority_batch, len(self.alpha_memory))
from_beta = min(batch_size - priority_batch, len(self.beta_memory))
res = random.sample(self.alpha_memory, from_alpha) + random.sample(self.beta_memory, from_beta)
random.shuffle(res)
return res
def __len__(self):
return len(self.alpha_memory) + len(self.beta_memory)
# + [markdown] id="6qMQboCLcEu7" colab_type="text"
# ## Agent
# + colab_type="code" id="jF57tZRBzMa8" colab={}
class CustomAgent:
def __init__(self):
global embedding_weights
"""
Arguments:
word_vocab: List of words supported.
"""
self.mode = "train"
with open("./vocab.txt") as f:
self.word_vocab = f.read().split("\n")
with open("config.yaml") as reader:
self.config = yaml.safe_load(reader)
self.word2id = {}
self.last_wid = 0
for w in self.word_vocab:
self.word2id[w] = self.last_wid
self.last_wid+=1
self.EOS_id = self.word2id["</S>"]
self.batch_size = self.config['training']['batch_size']
self.max_nb_steps_per_episode = self.config['training']['max_nb_steps_per_episode']
self.nb_epochs = self.config['training']['nb_epochs']
# Set the random seed manually for reproducibility.
np.random.seed(self.config['general']['random_seed'])
torch.manual_seed(self.config['general']['random_seed'])
if torch.cuda.is_available():
if not self.config['general']['use_cuda']:
logging.warning("WARNING: CUDA device detected but 'use_cuda: false' found in config.yaml")
self.use_cuda = False
else:
torch.backends.cudnn.deterministic = True
torch.cuda.manual_seed(self.config['general']['random_seed'])
self.use_cuda = True
else:
self.use_cuda = False
print("Creating Q-Network")
self.model = Bert_DQN(model_config=self.config["model"],
word_vocab=self.word_vocab,
enable_cuda=self.use_cuda)
print("Creating Target Network")
self.target_model = Bert_DQN(model_config=self.config["model"],
word_vocab=self.word_vocab,
enable_cuda=self.use_cuda)
self.target_model.print_parameters()
self.update_target_model_count = 0
self.target_model_update_frequency = self.config['training']['target_model_update_frequency']
self.experiment_tag = self.config['checkpoint']['experiment_tag']
self.model_checkpoint_dir = self.config['checkpoint']['model_checkpoint_dir']
self.save_frequency = self.config['checkpoint']['save_frequency']
if self.config['checkpoint']['load_pretrained']:
self.load_pretrained_model(self.model_checkpoint_dir)
if self.use_cuda:
self.model.cuda()
self.target_model.cuda()
self.replay_batch_size = self.config['general']['replay_batch_size']
self.replay_memory = PrioritizedReplayMemory(self.config['general']['replay_memory_capacity'],
priority_fraction=self.config['general']['replay_memory_priority_fraction'])
self.wt_index = 0
# optimizer
parameters = filter(lambda p: p.requires_grad, self.model.parameters())
self.optimizer = torch.optim.Adam(parameters, lr=self.config['training']['optimizer']['learning_rate'])
# n-step
self.nsteps = self.config['general']['nsteps']
self.nstep_buffer = []
# epsilon greedy
self.epsilon_anneal_episodes = self.config['general']['epsilon_anneal_episodes']
self.epsilon_anneal_from = self.config['general']['epsilon_anneal_from']
self.epsilon_anneal_to = self.config['general']['epsilon_anneal_to']
self.epsilon = self.epsilon_anneal_from
self.update_per_k_game_steps = self.config['general']['update_per_k_game_steps']
self.clip_grad_norm = self.config['training']['optimizer']['clip_grad_norm']
self.nlp = spacy.load('en', disable=['ner', 'parser', 'tagger'])
self.preposition_map = {"take": "from",
"chop": "with",
"slice": "with",
"dice": "with",
"cook": "with",
"insert": "into",
"put": "on"}
self.single_word_verbs = set(["inventory", "look"])
self.discount_gamma = self.config['general']['discount_gamma']
self.current_episode = 0
self.current_step = 0
self._epsiode_has_started = False
self.history_avg_scores = HistoryScoreCache(capacity=1000)
self.best_avg_score_so_far = 0.0
self.loss = []
def train(self, imitate = False):
"""
Tell the agent that it's training phase.
"""
self.mode = "train"
self.imitate = imitate
self.wt_index = 0
# print(self.wt_index)
self.model.train()
def eval(self):
"""
Tell the agent that it's evaluation phase.
"""
self.mode = "eval"
self.model.eval()
def _start_episode(self, obs: List[str], infos: Dict[str, List[Any]]) -> None:
"""
Prepare the agent for the upcoming episode.
Arguments:
obs: Initial feedback for each game.
infos: Additional information for each game.
"""
self.init(obs, infos)
self._epsiode_has_started = True
def _end_episode(self, obs: List[str], scores: List[int], infos: Dict[str, List[Any]]) -> None:
"""
Tell the agent the episode has terminated.
Arguments:
obs: Previous command's feedback for each game.
score: The score obtained so far for each game.
infos: Additional information for each game.
"""
self.finish()
self._epsiode_has_started = False
def load_pretrained_model(self, load_from_dir):
"""
Load the pretrained model's last checkpoint from a directory
Arguments:
load_from_dir: Directory with save model parameters
"""
checkpoints_glob = glob.glob(os.path.join(load_from_dir, '*.pt'))
if len(checkpoints_glob) == 0:
print("No checkpoints to load from: " + load_from_dir)
return
load_from = max(checkpoints_glob)
print("loading model from %s\n" % (load_from))
try:
if self.use_cuda:
state_dict = torch.load(load_from)
else:
state_dict = torch.load(load_from, map_location='cpu')
self.model.load_state_dict(state_dict)
except:
print("Failed to load checkpoint...")
def select_additional_infos(self) -> EnvInfos:
"""
Returns what additional information should be made available at each game step.
Requested information will be included within the `infos` dictionary
passed to `CustomAgent.act()`. To request specific information, create a
:py:class:`textworld.EnvInfos <textworld.envs.wrappers.filter.EnvInfos>`
and set the appropriate attributes to `True`. The possible choices are:
* `description`: text description of the current room, i.e. output of the `look` command;
* `inventory`: text listing of the player's inventory, i.e. output of the `inventory` command;
* `max_score`: maximum reachable score of the game;
* `objective`: objective of the game described in text;
* `entities`: names of all entities in the game;
* `verbs`: verbs understood by the the game;
* `command_templates`: templates for commands understood by the the game;
* `admissible_commands`: all commands relevant to the current state;
In addition to the standard information, game specific information
can be requested by appending corresponding strings to the `extras`
attribute. For this competition, the possible extras are:
* `'recipe'`: description of the cookbook;
* `'walkthrough'`: one possible solution to the game (not guaranteed to be optimal);
Example:
Here is an example of how to request information and retrieve it.
>>> from textworld import EnvInfos
>>> request_infos = EnvInfos(description=True, inventory=True, extras=["recipe"])
...
>>> env = gym.make(env_id)
>>> ob, infos = env.reset()
>>> print(infos["description"])
>>> print(infos["inventory"])
>>> print(infos["extra.recipe"])
Notes:
The following information *won't* be available at test time:
* 'walkthrough'
"""
request_infos = EnvInfos()
request_infos.description = True
request_infos.inventory = True
request_infos.entities = True
request_infos.verbs = True
request_infos.extras = ["recipe", "walkthrough"]
return request_infos
def init(self, obs: List[str], infos: Dict[str, List[Any]]):
"""
Prepare the agent for the upcoming games.
Arguments:
obs: Previous command's feedback for each game.
infos: Additional information for each game.
"""
# reset agent, get vocabulary masks for verbs / adjectives / nouns
self.scores = []
self.dones = []
self.prev_actions = ["" for _ in range(len(obs))]
# get word masks
# print(infos['verbs'])
batch_size = len(infos["verbs"])
# print("VERBS SIZE:")
# print(batch_size)
verbs_word_list = infos["verbs"]
noun_word_list, adj_word_list = [], []
for entities in infos["entities"]:
tmp_nouns, tmp_adjs = [], []
for name in entities:
split = name.split()
tmp_nouns.append(split[-1])
if len(split) > 1:
tmp_adjs.append(" ".join(split[:-1]))
noun_word_list.append(list(set(tmp_nouns)))
adj_word_list.append(list(set(tmp_adjs)))
verb_mask = np.zeros((batch_size, len(self.word_vocab)), dtype="float32")
noun_mask = np.zeros((batch_size, len(self.word_vocab)), dtype="float32")
adj_mask = np.zeros((batch_size, len(self.word_vocab)), dtype="float32")
for i in range(batch_size):
for w in verbs_word_list[i]:
if w in self.word2id:
verb_mask[i][self.word2id[w]] = 1.0
for w in noun_word_list[i]:
if w in self.word2id:
noun_mask[i][self.word2id[w]] = 1.0
for w in adj_word_list[i]:
if w in self.word2id:
adj_mask[i][self.word2id[w]] = 1.0
# else:
# self.word2id[w] = self.last_wid
# self.last_wid += 1
# adj_mask[i][self.word2id[w]] = 1.0
# self.word_vocab.append(w)
second_noun_mask = copy.copy(noun_mask)
second_adj_mask = copy.copy(adj_mask)
second_noun_mask[:, self.EOS_id] = 1.0
adj_mask[:, self.EOS_id] = 1.0
second_adj_mask[:, self.EOS_id] = 1.0
self.word_masks_np = [verb_mask, adj_mask, noun_mask, second_adj_mask, second_noun_mask]
self.cache_bert_ids = None
self.cache_bert_mask = None
self.cache_chosen_indices = None
self.current_step = 0
def append_to_replay(self, is_prior, transition):
self.nstep_buffer.append((is_prior, transition))
if len(self.nstep_buffer) < self.nsteps:
return
R = sum([self.nstep_buffer[i][1].reward * (self.discount_gamma**i) for i in range(self.nsteps)])
prior, transition = self.nstep_buffer.pop(0)
self.replay_memory.push(prior, transition._replace(reward=R))
def get_game_step_info(self, obs: List[str], infos: Dict[str, List[Any]]):
"""
Get all the available information, and concat them together to be tensor for
a neural model. we use post padding here, all information are tokenized here.
Arguments:
obs: Previous command's feedback for each game.
infos: Additional information for each game.
"""
sep = ' [SEP] '
description_text_list = [_d + sep + _i + sep + _q + sep + _f + sep + _pa for (_d, _i, _q, _f, _pa)
in zip(infos['description'], infos['inventory'], infos['extra.recipe'], obs, self.prev_actions)]
_, bert_ids, bert_masks = convert_examples_to_features(description_text_list, self.model.tokenizer)
del description_text_list
return bert_ids, bert_masks
def word_ids_to_commands(self, verb, adj, noun, adj_2, noun_2):
"""
Turn the 5 indices into actual command strings.
Arguments:
verb: Index of the guessing verb in vocabulary
adj: Index of the guessing adjective in vocabulary
noun: Index of the guessing noun in vocabulary
adj_2: Index of the second guessing adjective in vocabulary
noun_2: Index of the second guessing noun in vocabulary
"""
# turns 5 indices into actual command strings
if self.word_vocab[verb] in self.single_word_verbs:
return self.word_vocab[verb]
if adj == self.EOS_id:
res = self.word_vocab[verb] + " " + self.word_vocab[noun]
else:
res = self.word_vocab[verb] + " " + self.word_vocab[adj] + " " + self.word_vocab[noun]
if self.word_vocab[verb] not in self.preposition_map:
return res
if noun_2 == self.EOS_id:
return res
prep = self.preposition_map[self.word_vocab[verb]]
if adj_2 == self.EOS_id:
res = res + " " + prep + " " + self.word_vocab[noun_2]
else:
res = res + " " + prep + " " + self.word_vocab[adj_2] + " " + self.word_vocab[noun_2]
return res
def get_wordid_from_vocab(self, word):
if word in self.word2id.keys():
return self.word2id[word]
else:
return self.EOS_id
def command_to_word_ids(self, cmd, batch_size):
verb_id=self.EOS_id
first_adj=self.EOS_id
first_noun=self.EOS_id
second_adj=self.EOS_id
second_noun=self.EOS_id
# print('cmd_to_ids')
# print(cmd.split())
ids = _words_to_ids(cmd.split(), self.word2id)
# print(ids)
for ind, i in enumerate(ids):
if self.word_masks_np[0][0][i]==1.0:
verb = ind
verb_id = i
nouns=[]
for ind, i in enumerate(ids):
if self.word_masks_np[2][0][i]==1.0:
nouns.append((ind,i))
if len(nouns) > 0:
if nouns[0][0] != verb - 1:
adj_ids = ids[verb + 1: nouns[0][0]]
adj=''
adj= ' '.join([self.word_vocab[x] for x in adj_ids])
# print(adj)
first_adj=self.get_wordid_from_vocab(adj)
# print(nouns)
first_noun=nouns[0][1]
if len(nouns) > 1:
if nouns[1][0] != nouns[0][0] - 1:
adj_ids = ids[nouns[0][0]: nouns[1][0]]
adj= ' '.join([self.word_vocab[x] for x in adj_ids])
second_adj=self.get_wordid_from_vocab(adj)
second_noun=nouns[1][1]
list_ids = [verb_id, first_adj, first_noun, second_adj, second_noun]
return [to_pt(np.array([[x]]*batch_size), self.use_cuda) for x in list_ids]
def get_chosen_strings(self, chosen_indices):
"""
Turns list of word indices into actual command strings.
Arguments:
chosen_indices: Word indices chosen by model.
"""
chosen_indices_np = [to_np(item)[:, 0] for item in chosen_indices]
res_str = []
batch_size = chosen_indices_np[0].shape[0]
for i in range(batch_size):
verb, adj, noun, adj_2, noun_2 = chosen_indices_np[0][i],\
chosen_indices_np[1][i],\
chosen_indices_np[2][i],\
chosen_indices_np[3][i],\
chosen_indices_np[4][i]
res_str.append(self.word_ids_to_commands(verb, adj, noun, adj_2, noun_2))
del verb
del adj
del noun
del adj_2
del noun_2
del chosen_indices_np
return res_str
def choose_random_command(self, word_ranks, word_masks_np):
"""
Generate a command randomly, for epsilon greedy.
Arguments:
word_ranks: Q values for each word by model.action_scorer.
word_masks_np: Vocabulary masks for words depending on their type (verb, adj, noun).
"""
batch_size = word_ranks[0].size(0)
word_ranks_np = [to_np(item) for item in word_ranks] # list of batch x n_vocab
word_ranks_np = [r * m for r, m in zip(word_ranks_np, word_masks_np)] # list of batch x n_vocab
word_indices = []
for i in range(len(word_ranks_np)):
indices = []
for j in range(batch_size):
msk = word_masks_np[i][j] # vocab
indices.append(np.random.choice(len(msk), p=msk / np.sum(msk, -1)))
del msk
# print('random: ', indices)
word_indices.append(np.array(indices))
del indices
# word_indices: list of batch
word_qvalues = [[] for _ in word_masks_np]
for i in range(batch_size):
for j in range(len(word_qvalues)):
word_qvalues[j].append(word_ranks[j][i][word_indices[j][i]])
word_qvalues = [torch.stack(item) for item in word_qvalues]
word_indices = [to_pt(item, self.use_cuda) for item in word_indices]
word_indices = [item.unsqueeze(-1) for item in word_indices] # list of batch x 1
del word_ranks_np
return word_qvalues, word_indices
def choose_maxQ_command(self, word_ranks, word_masks_np):
"""
Generate a command by maximum q values, for epsilon greedy.
Arguments:
word_ranks: Q values for each word by model.action_scorer.
word_masks_np: Vocabulary masks for words depending on their type (verb, adj, noun).
"""
batch_size = word_ranks[0].size(0)
word_ranks_np = [to_np(item) for item in word_ranks] # list of batch x n_vocab
word_ranks_np = [r - np.min(r) for r in word_ranks_np] # minus the min value, so that all values are non-negative
word_ranks_np = [r * m for r, m in zip(word_ranks_np, word_masks_np)] # list of batch x n_vocab
word_indices = [np.argmax(item, -1) for item in word_ranks_np] # list of batch
word_qvalues = [[] for _ in word_masks_np]
for i in range(batch_size):
for j in range(len(word_qvalues)):
word_qvalues[j].append(word_ranks[j][i][word_indices[j][i]])
word_qvalues = [torch.stack(item) for item in word_qvalues]
word_indices = [to_pt(item, self.use_cuda) for item in word_indices]
word_indices = [item.unsqueeze(-1) for item in word_indices] # list of batch x 1
del word_ranks_np
return word_qvalues, word_indices
def get_ranks(self, model, bert_ids, bert_masks):
"""
Given input description tensor, call model forward, to get Q values of words.
Arguments:
input_description: Input tensors, which include all the information chosen in
select_additional_infos() concatenated together.
"""
bert_ids = torch.tensor([x for x in bert_ids], dtype=torch.long)
bert_masks = torch.tensor([x for x in bert_masks], dtype=torch.long)
state_representation = model.representation_generator(bert_ids, bert_masks)
del bert_ids
del bert_masks
word_ranks = model.action_scorer(state_representation) # each element in list has batch x n_vocab size
del state_representation
return word_ranks
def act_eval(self, obs: List[str], scores: List[int], dones: List[bool], infos: Dict[str, List[Any]]) -> List[str]:
"""
Acts upon the current list of observations, during evaluation.
One text command must be returned for each observation.
Arguments:
obs: Previous command's feedback for each game.
score: The score obtained so far for each game (at previous step).
done: Whether a game is finished (at previous step).
infos: Additional information for each game.
Returns:
Text commands to be performed (one per observation).
Notes:
Commands returned for games marked as `done` have no effect.
The states for finished games are simply copy over until all
games are done, in which case `CustomAgent.finish()` is called
instead.
"""
if self.current_step > 0:
# append scores / dones from previous step into memory
self.scores.append(scores)
self.dones.append(dones)
if all(dones):
self._end_episode(obs, scores, infos)
return # Nothing to return.
bert_ids, bert_masks = self.get_game_step_info(obs, infos)
word_ranks = self.get_ranks(self.model, bert_ids, bert_masks) # list of batch x vocab
del bert_ids
del bert_masks
_, word_indices_maxq = self.choose_maxQ_command(word_ranks, self.word_masks_np)
chosen_indices = word_indices_maxq
chosen_indices = [item.detach() for item in chosen_indices]
chosen_strings = self.get_chosen_strings(chosen_indices)
self.prev_actions = chosen_strings
self.current_step += 1
del word_indices_max_q
return chosen_strings
def act(self, obs: List[str], scores: List[int], dones: List[bool], infos: Dict[str, List[Any]]) -> List[str]:
"""
Acts upon the current list of observations.
One text command must be returned for each observation.
Arguments:
obs: Previous command's feedback for each game.
score: The score obtained so far for each game (at previous step).
done: Whether a game is finished (at previous step).
infos: Additional information for each game.
Returns:
Text commands to be performed (one per observation).
Notes:
Commands returned for games marked as `done` have no effect.
The states for finished games are simply copy over until all
games are done, in which case `CustomAgent.finish()` is called
instead.
"""
if not self._epsiode_has_started:
self._start_episode(obs, infos)
if self.mode == "eval":
return self.act_eval(obs, scores, dones, infos)
if self.current_step > 0:
# append scores / dones from previous step into memory
self.scores.append(scores)
self.dones.append(dones)
# compute previous step's rewards and masks
rewards_np, rewards, mask_np, mask = self.compute_reward()
bert_ids, bert_masks = self.get_game_step_info(obs, infos)
# generate commands for one game step, epsilon greedy is applied, i.e.,
# there is epsilon of chance to generate random commands
if self.imitate:
print('imitate')
correct_cmd=infos['extra.walkthrough'][0][self.wt_index]
print(correct_cmd)
if self.wt_index != len(infos['extra.walkthrough'][0]) - 1:
self.wt_index+=1
chosen_indices = self.command_to_word_ids(correct_cmd, len(bert_ids))
else:
word_ranks = self.get_ranks(self.model, bert_ids, bert_masks) # list of batch x vocab
_, word_indices_maxq = self.choose_maxQ_command(word_ranks, self.word_masks_np)
_, word_indices_random = self.choose_random_command(word_ranks, self.word_masks_np)
# random number for epsilon greedyupdate
rand_num = np.random.uniform(low=0.0, high=1.0, size=(len(bert_ids), 1))
less_than_epsilon = (rand_num < self.epsilon).astype("float32") # batch
greater_than_epsilon = 1.0 - less_than_epsilon
less_than_epsilon = to_pt(less_than_epsilon, self.use_cuda, type='float')
greater_than_epsilon = to_pt(greater_than_epsilon, self.use_cuda, type='float')
less_than_epsilon, greater_than_epsilon = less_than_epsilon.long(), greater_than_epsilon.long()
# print('Random_step: ',less_than_epsilon.tolist())
chosen_indices = [
less_than_epsilon * idx_random + greater_than_epsilon * idx_maxq
for idx_random, idx_maxq in zip(word_indices_random, word_indices_maxq)
]
chosen_indices = [item.detach() for item in chosen_indices]
chosen_strings = self.get_chosen_strings(chosen_indices)
# print(chosen_strings)
self.prev_actions = chosen_strings
# push info from previous game step into replay memory
if self.current_step > 0:
for b in range(len(obs)):
if mask_np[b] == 0:
continue
is_prior = rewards_np[b] > 0.0
t = Transition(self.cache_bert_ids[b],
self.cache_bert_masks[b],
[ item[b] for item in self.cache_chosen_indices],
rewards[b],
mask[b],
dones[b],
bert_ids[b],
bert_masks[b],
[item[b] for item in self.word_masks_np])
# print("ACT: {}".format(t.observation_id_list))
self.append_to_replay(is_prior, t)
# cache new info in current game step into caches
self.cache_bert_ids = bert_ids
self.cache_bert_masks = bert_masks
self.cache_chosen_indices = chosen_indices
# update neural model by replaying snapshots in replay memory
if self.current_step > 0 and self.current_step % self.update_per_k_game_steps == 0:
loss = self.update()
if loss is not None:
self.loss.append(to_np(loss).mean())
# Backpropagate
self.optimizer.zero_grad()
loss.backward(retain_graph=True)
# `clip_grad_norm` helps prevent the exploding gradient problem in RNNs / LSTMs.
torch.nn.utils.clip_grad_norm_(self.model.parameters(), self.clip_grad_norm)
self.optimizer.step() # apply gradients
self.current_step += 1
if all(dones):
self._end_episode(obs, scores, infos)
return # Nothing to return.
return chosen_strings
def compute_reward(self):
"""
Compute rewards by agent. Note this is different from what the training/evaluation
scripts do. Agent keeps track of scores and other game information for training purpose.
"""
# mask = 1 if game is not finished or just finished at current step
if len(self.dones) == 1:
# it's not possible to finish a game at 0th step
mask = [1.0 for _ in self.dones[-1]]
else:
assert len(self.dones) > 1
mask = [1.0 if not self.dones[-2][i] else 0.0 for i in range(len(self.dones[-1]))]
mask = np.array(mask, dtype='float32')
mask_pt = to_pt(mask, self.use_cuda, type='float')
# rewards returned by game engine are always accumulated value the
# agent have recieved. so the reward it gets in the current game step
# is the new value minus values at previous step.
rewards = np.array(self.scores[-1], dtype='float32') # batch
if len(self.scores) > 1:
prev_rewards = np.array(self.scores[-2], dtype='float32')
rewards = rewards - prev_rewards
rewards_pt = to_pt(rewards, self.use_cuda, type='float')
return rewards, rewards_pt, mask, mask_pt
def update_target_model(self):
self.update_target_model_count = (self.update_target_model_count + 1) % self.target_model_update_frequency
if self.update_target_model_count == 0:
self.target_model.load_state_dict(self.model.state_dict())
def update(self):
"""
Update neural model in agent. In this example we follow algorithm
of updating model in dqn with replay memory.
"""
if len(self.replay_memory) < self.replay_batch_size:
return None
self.update_target_model()
transitions = self.replay_memory.sample(self.replay_batch_size)
batch = Transition(*zip(*transitions))
del transitions
bert_ids = pad_sequences(batch.bert_ids, maxlen=max_len(batch.bert_ids)).astype('int32')
bert_masks = pad_sequences(batch.bert_masks, maxlen=max_len(batch.bert_masks)).astype('int32')
next_bert_ids = pad_sequences(batch.next_bert_ids, maxlen=max_len(batch.next_bert_ids)).astype('int32')
next_bert_masks = pad_sequences(batch.next_bert_masks, maxlen=max_len(batch.next_bert_masks)).astype('int32')
chosen_indices = list(list(zip(*batch.word_indices)))
chosen_indices = [torch.stack(item, 0) for item in chosen_indices] # list of batch x 1
word_ranks = self.get_ranks(self.model, bert_ids, bert_masks) # list of batch x vocab
del bert_ids
del bert_masks
word_qvalues = [w_rank.gather(1, idx).squeeze(-1) for w_rank, idx in zip(word_ranks, chosen_indices)] # list of batch
del chosen_indices
del word_ranks
q_value = torch.mean(torch.stack(word_qvalues, -1), -1) # batch
del word_qvalues
# Action selection, using q-network
next_word_ranks = self.get_ranks(self.model, next_bert_ids, next_bert_masks) # batch x n_verb, batch x n_noun, batch x n_second_noun
next_word_masks = list(list(zip(*batch.next_word_masks)))
next_word_masks = [np.stack(item, 0) for item in next_word_masks]
_, next_word_indexes = self.choose_maxQ_command(next_word_ranks, next_word_masks)
del next_word_masks
del next_word_ranks
# Action evaluation, using target network
eval_next_word_ranks = self.get_ranks(self.target_model, next_bert_ids, next_bert_masks)
next_word_qvalues = [
rank.gather(1, idx.detach()).squeeze(-1)
for rank, idx in zip(eval_next_word_ranks, next_word_indexes)
]
del next_word_indexes
del eval_next_word_ranks
del next_bert_ids
del next_bert_masks
next_q_value = torch.mean(torch.stack(next_word_qvalues, -1), -1) # batch
next_q_value = next_q_value.detach()
rewards = torch.stack(batch.reward) # batch
not_done = 1.0 - np.array(batch.done, dtype='float32') # batch
not_done = to_pt(not_done, self.use_cuda, type='float')
# NB: Should not_done be used?
rewards = rewards + not_done * next_q_value * (self.discount_gamma**self.nsteps) # batch
#rewards = rewards + next_q_value * (self.discount_gamma**self.nsteps) # batch
mask = torch.stack(batch.mask) # batch
loss = F.smooth_l1_loss(q_value * mask, rewards * mask)
del q_value
del mask
del rewards
del batch
return loss
def finish(self) -> None:
"""
All games in the batch are finished. One can choose to save checkpoints,
evaluate on validation set, or do parameter annealing here.
"""
# Game has finished (either win, lose, or exhausted all the given steps).
self.final_rewards = np.array(self.scores[-1], dtype='float32') # batch
dones = []
for d in self.dones:
d = np.array([float(dd) for dd in d], dtype='float32')
dones.append(d)
dones = np.array(dones)
step_used = 1.0 - dones
self.step_used_before_done = np.sum(step_used, 0) # batch
self.history_avg_scores.push(np.mean(self.final_rewards))
# save checkpoint
if self.mode == "train" and self.current_episode % self.save_frequency == 0:
avg_score = self.history_avg_scores.get_avg()
if avg_score > self.best_avg_score_so_far:
self.best_avg_score_so_far = avg_score
save_to = os.path.join(self.model_checkpoint_dir, self.experiment_tag + "_episode_" + str(self.current_episode) + ".pt")
if not os.path.isdir(self.model_checkpoint_dir):
os.mkdir(self.model_checkpoint_dir)
torch.save(self.model.state_dict(), save_to)
print("\n========= saved checkpoint =========")
self.current_episode += 1
# annealing
if self.current_episode < self.epsilon_anneal_episodes:
self.epsilon -= (self.epsilon_anneal_from - self.epsilon_anneal_to) / float(self.epsilon_anneal_episodes)
def get_mean_loss(self):
mean_loss = 0.
if len(self.loss) != 0:
mean_loss = sum(self.loss) / len(self.loss)
self.loss = []
return mean_loss
# + [markdown] id="bHuITVpD0nAN" colab_type="text"
# ## Configs and environments
# + [markdown] id="N9jFzEnr1zt9" colab_type="text"
# ### Vocab
# Upload vocab.txt file`
# + id="Xfc78DzULeIf" colab_type="code" outputId="edea043b-468a-4125-b7ce-baeee5e39b59" colab={"base_uri": "https://localhost:8080/", "height": 34}
from google.colab import files
if not os.path.isfile('./vocab.txt'):
uploaded = files.upload()
# Upload vocab.txt
for fn in uploaded.keys():
print('User uploaded file "{name}" with length {length} bytes'.format(name=fn, length=len(uploaded[fn])))
else:
print("Vocab already uploaded!")
# + id="Vhn6sd_gLvC-" colab_type="code" outputId="890bfe76-77a1-465f-de15-d6bfa70fad19" colab={"base_uri": "https://localhost:8080/", "height": 187}
# !head vocab.txt
# + [markdown] id="YWAhh8mL2WCf" colab_type="text"
# ### Configuration
# + id="ufN2JE2f1eTO" colab_type="code" colab={}
with open('./config.yaml', 'w') as config:
config.write("""
general:
discount_gamma: 0.7
random_seed: 42
use_cuda: True # disable this when running on machine without cuda
# replay memory
replay_memory_capacity: 1000000 # adjust this depending on your RAM size
replay_memory_priority_fraction: 0.5
update_per_k_game_steps: 8
replay_batch_size: 32
nsteps: 3
# epsilon greedy
epsilon_anneal_episodes: 50 # -1 if not annealing
epsilon_anneal_from: 0.2
epsilon_anneal_to: 0.2
checkpoint:
experiment_tag: 'starting-kit'
model_checkpoint_dir: '/gdrive/My Drive/Masters/TextWorld/models'
load_pretrained: False # during test, enable this so that the agent load your pretrained model
#pretrained_experiment_dir: 'starting-kit'
save_frequency: 100
training:
batch_size: 16 # Parallel games played at once
nb_epochs: 100
max_nb_steps_per_episode: 100 # after this many steps, a game is terminated
target_model_update_frequency: 16 # update target model after that number of backprops
optimizer:
step_rule: 'adam' # adam
learning_rate: 0.001
clip_grad_norm: 5
model:
embedding_size: 50
freeze_embedding: False
encoder_rnn_hidden_size: [192]
bidirectional_lstm: False
action_scorer_hidden_dim: 63
dropout_between_rnn_layers: 0.0
bert_model: 'bert-base-uncased'
train_bert: False
""")
# + [markdown] id="Je184JvpCfos" colab_type="text"
# ### Mount drive to load games
#
# Notebook takes sample games from google drive(requires authentication).
#
# To train the agent with games, upload archive with them in google drive and fix the path to the archive inside drive below.
#
#
# + id="XugC3fAz9hpB" colab_type="code" outputId="7e3431db-569e-4cb6-897f-e976a902366c" colab={"base_uri": "https://localhost:8080/", "height": 34}
from google.colab import drive
drive.mount('/gdrive')
# + id="sIGWqhW4Z7NB" colab_type="code" colab={}
home_dir = '/gdrive/My Drive/Masters/TextWorld/'
path_to_sample_games = home_dir + 'sample_games'
# + [markdown] id="924KmPsyvmwF" colab_type="text"
# ## Train
# + colab_type="code" id="2HzAOxlCPDNI" colab={}
# List of additional information available during evaluation.
AVAILABLE_INFORMATION = EnvInfos(
description=True, inventory=True,
max_score=True, objective=True, entities=True, verbs=True,
command_templates=True, admissible_commands=True,
has_won=True, has_lost=True,
extras=["recipe"]
)
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
def _validate_requested_infos(infos: EnvInfos):
msg = "The following information cannot be requested: {}"
for key in infos.basics:
if not getattr(AVAILABLE_INFORMATION, key):
raise ValueError(msg.format(key))
for key in infos.extras:
if key not in AVAILABLE_INFORMATION.extras:
raise ValueError(msg.format(key))
def get_index(game_no, stats):
return "{}_{}".format(game_no, stats)
def print_epoch_stats(epoch_no, stats):
print("\n\nEpoch: {:3d}".format(epoch_no))
steps, scores, loss = stats["steps"], stats["scores"], stats["loss"]
games_cnt, parallel_cnt = len(steps), len(steps[0])
columns = [ get_index(col, st) for col in range(games_cnt) for st in ['st', 'sc']]
stats_df = pd.DataFrame(index=list(range(parallel_cnt)) + ["avr", "loss"], columns=columns)
for col in range(games_cnt):
for row in range(parallel_cnt):
stats_df[get_index(col, 'st')][row] = steps[col][row]
stats_df[get_index(col, 'sc')][row] = scores[col][row]
stats_df[get_index(col, 'sc')]['avr'] = stats_df[get_index(col, 'sc')].mean()
stats_df[get_index(col, 'st')]['avr'] = stats_df[get_index(col, 'st')].mean()
stats_df[get_index(col, 'sc')]['loss'] = "{:.5f}".format(loss[col])
print(stats_df)
def train(game_files):
print("Agent starting...")
agent = CustomAgent()
print("Agent started")
requested_infos = agent.select_additional_infos()
# _validate_requested_infos(requested_infos)
env_id = textworld.gym.register_games(game_files, requested_infos,
max_episode_steps=agent.max_nb_steps_per_episode,
name="training")
env_id = textworld.gym.make_batch(env_id, batch_size=agent.batch_size, parallel=True)
print("Making {} parallel environments to train on them\n".format(agent.batch_size))
env = gym.make(env_id)
max_score = -1
game_range = range(len(game_files))
for epoch_no in range(1, agent.nb_epochs + 1):
stats = {
"scores": [],
"steps": [],
"loss": [],
}
for game_no in tqdm(game_range):
obs, infos = env.reset()
imitate = random.random() > 0.7
agent.train(imitate)
scores = [0] * len(obs)
dones = [False] * len(obs)
steps = [0] * len(obs)
while not all(dones):
# Increase step counts.
steps = [step + int(not done) for step, done in zip(steps, dones)]
commands = agent.act(obs, scores, dones, infos)
obs, scores, dones, infos = env.step(commands)
# Let the agent knows the game is done.
agent.act(obs, scores, dones, infos)
stats["scores"].append(scores)
stats["steps"].append(steps)
stats["loss"].append( agent.get_mean_loss())
print_epoch_stats(epoch_no, stats)
#torch.save(agent.model, './agent_model.pt')
return
# + id="gCw4zpIiqyGK" colab_type="code" outputId="bb9b0d07-41b5-49d4-90d3-d2812a8e55f1" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# %%time
game_dir = path_to_sample_games
games = []
if os.path.isdir(game_dir):
games += glob.glob(os.path.join(game_dir, "*.ulx"))
print("{} games found for training.".format(len(games)))
if len(games) != 0:
train(games)
# + [markdown] id="k8hIulvHzNce" colab_type="text"
# ## Save models
# + id="nrIFg_ec1uvV" colab_type="code" colab={}
# !ls -lh '/gdrive/My Drive/saved_models'
# + id="ScjbitVE95E1" colab_type="code" colab={}
| experiments/bert_double_dqn_textworld_imitation_learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="7uctu-ZzRZOv"
# # Task 1: Try the algo on Same Whether dataset - LabelEncoding of features: and Train test Division 95%-5%
# + [markdown] id="De7AICwsdTMR"
# **Aim: Implement Decsion Tree classifier**
#
#
# - Implement Decision Tree classifier using scikit learn library
# - Test the classifier for Weather dataset
# + [markdown] id="alhwH449dTMd"
# Step 1: Import necessary libraries.
# + id="1CNH0GcydTMk" executionInfo={"status": "ok", "timestamp": 1630589190174, "user_tz": -330, "elapsed": 1807, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitS8CqtpOcM8tH8itmtSfpDm13rQsGdSUSpu-5KQ=s64", "userId": "07832619671270321867"}}
from sklearn import preprocessing
from sklearn.tree import DecisionTreeClassifier
# + [markdown] id="jw3TNFuOdTNH"
# Step 2: Prepare dataset.
# + id="8FYtofzfdTNM" executionInfo={"status": "ok", "timestamp": 1630589195868, "user_tz": -330, "elapsed": 502, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitS8CqtpOcM8tH8itmtSfpDm13rQsGdSUSpu-5KQ=s64", "userId": "07832619671270321867"}}
#Predictor variables
Outlook = ['Rainy', 'Rainy', 'Overcast', 'Sunny', 'Sunny', 'Sunny', 'Overcast',
'Rainy', 'Rainy', 'Sunny', 'Rainy','Overcast', 'Overcast', 'Sunny']
Temperature = ['Hot', 'Hot', 'Hot', 'Mild', 'Cool', 'Cool', 'Cool',
'Mild', 'Cool', 'Mild', 'Mild', 'Mild', 'Hot', 'Mild']
Humidity = ['High', 'High', 'High', 'High', 'Normal', 'Normal', 'Normal',
'High', 'Normal', 'Normal', 'Normal', 'High', 'Normal', 'High']
Wind = ['False', 'True', 'False', 'False', 'False', 'True', 'True',
'False', 'False', 'False', 'True', 'True', 'False', 'True']
#Class Label:
Play = ['No', 'No', 'Yes', 'Yes', 'Yes', 'No', 'Yes', 'No',
'Yes', 'Yes', 'Yes', 'Yes', 'Yes', 'No']
# + [markdown] id="8FncMW4IdTNk"
# Step 3: Digitize the data set using encoding
# + colab={"base_uri": "https://localhost:8080/"} id="Awu5kIQxdTNo" executionInfo={"status": "ok", "timestamp": 1630589205734, "user_tz": -330, "elapsed": 514, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitS8CqtpOcM8tH8itmtSfpDm13rQsGdSUSpu-5KQ=s64", "userId": "07832619671270321867"}} outputId="cd14fde5-bc7e-495a-ecb5-50ea5a05bd47"
#creating labelEncoder
le = preprocessing.LabelEncoder()
# Converting string labels into numbers.
Outlook_encoded = le.fit_transform(Outlook)
Outlook_name_mapping = dict(zip(le.classes_, le.transform(le.classes_)))
print("Outllok mapping:",Outlook_name_mapping)
Temperature_encoded = le.fit_transform(Temperature)
Temperature_name_mapping = dict(zip(le.classes_, le.transform(le.classes_)))
print("Temperature mapping:",Temperature_name_mapping)
Humidity_encoded = le.fit_transform(Humidity)
Humidity_name_mapping = dict(zip(le.classes_, le.transform(le.classes_)))
print("Humidity mapping:",Humidity_name_mapping)
Wind_encoded = le.fit_transform(Wind)
Wind_name_mapping = dict(zip(le.classes_, le.transform(le.classes_)))
print("Wind mapping:",Wind_name_mapping)
Play_encoded = le.fit_transform(Play)
Play_name_mapping = dict(zip(le.classes_, le.transform(le.classes_)))
print("Play mapping:",Play_name_mapping)
print("\n\n")
print("Weather:" ,Outlook_encoded)
print("Temerature:" ,Temperature_encoded)
print("Humidity:" ,Humidity_encoded)
print("Wind:" ,Wind_encoded)
print("Play:" ,Play_encoded)
# + [markdown] id="ftSYL4UFdTN8"
# Step 4: Merge different features to prepare dataset
# + colab={"base_uri": "https://localhost:8080/"} id="RUCHRYb3dTOC" executionInfo={"status": "ok", "timestamp": 1630589210278, "user_tz": -330, "elapsed": 914, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitS8CqtpOcM8tH8itmtSfpDm13rQsGdSUSpu-5KQ=s64", "userId": "07832619671270321867"}} outputId="6cc3f01d-608f-47d9-9be1-f2da75cebd66"
#Combinig weather and Weather, Temerature, Humidity, Wind, Play into single listof tuples
features=tuple(zip(Outlook_encoded,Temperature_encoded,Humidity_encoded,Wind_encoded))
print("Features:",features)
# + [markdown] id="NMgpif_GdTOW"
# Step 5: Train ’Create and Train DecisionTreeClassifier’
# + id="XiW1S39HRZO3" executionInfo={"status": "ok", "timestamp": 1630589225436, "user_tz": -330, "elapsed": 431, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitS8CqtpOcM8tH8itmtSfpDm13rQsGdSUSpu-5KQ=s64", "userId": "07832619671270321867"}}
#import the necessary module
from sklearn.model_selection import train_test_split
#split data set into train and test sets
data_train, data_test, target_train, target_test = train_test_split(features,Play_encoded, test_size = 0.05, random_state = 130)
# + id="jDnsCMnGdTOa" executionInfo={"status": "ok", "timestamp": 1630589228155, "user_tz": -330, "elapsed": 3, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitS8CqtpOcM8tH8itmtSfpDm13rQsGdSUSpu-5KQ=s64", "userId": "07832619671270321867"}}
#Create a Decision Tree Classifier (using Entropy)
# Train the model using the training sets
import numpy as np
DecTr = DecisionTreeClassifier()
#Train the model using the training sets
DecTr=DecTr.fit(data_train, target_train)
# + [markdown] id="ZmgZ1-obdTOw"
# Step 6: Predict Output for new data
# + colab={"base_uri": "https://localhost:8080/"} id="lD0pzkWtdTOy" executionInfo={"status": "ok", "timestamp": 1630589232927, "user_tz": -330, "elapsed": 628, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitS8CqtpOcM8tH8itmtSfpDm13rQsGdSUSpu-5KQ=s64", "userId": "07832619671270321867"}} outputId="e9f86f02-12b1-478f-db9f-b3f15966be1a"
#Predict Output
#Predict the response for test dataset
target_pred = DecTr.predict(data_test)
print(data_test)
print(target_pred)
# + colab={"base_uri": "https://localhost:8080/"} id="77tJyE-XRZO6" executionInfo={"status": "ok", "timestamp": 1630589236023, "user_tz": -330, "elapsed": 4, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitS8CqtpOcM8tH8itmtSfpDm13rQsGdSUSpu-5KQ=s64", "userId": "07832619671270321867"}} outputId="438cf9a1-f8ce-4345-d74b-e8e5ce934538"
#Import confusion_matrix from scikit-learn metrics module for confusion_matrix
from sklearn.metrics import confusion_matrix
confusion_matrix(target_test, target_pred)
# + colab={"base_uri": "https://localhost:8080/"} id="n-xks5stRZO6" executionInfo={"status": "ok", "timestamp": 1630589239998, "user_tz": -330, "elapsed": 416, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitS8CqtpOcM8tH8itmtSfpDm13rQsGdSUSpu-5KQ=s64", "userId": "07832619671270321867"}} outputId="8f68f78f-f5ec-408f-c695-6f21bebb884c"
#Import scikit-learn metrics module for accuracy calculation
from sklearn import metrics
# Model Accuracy, how often is the classifier correct?
print("Accuracy:",metrics.accuracy_score(target_test, target_pred))
# + colab={"base_uri": "https://localhost:8080/"} id="GrNYi6RRRZO7" executionInfo={"status": "ok", "timestamp": 1630589243674, "user_tz": -330, "elapsed": 447, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitS8CqtpOcM8tH8itmtSfpDm13rQsGdSUSpu-5KQ=s64", "userId": "07832619671270321867"}} outputId="d8dc8f74-79f3-489b-fb0f-681470d895af"
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
precision = precision_score(target_test, target_pred)
recall = recall_score(target_test, target_pred)
print('precision: {}'.format(precision))
print('recall: {}'.format(recall))
# + colab={"base_uri": "https://localhost:8080/", "height": 485} id="FZ9srf8QRZO8" executionInfo={"status": "ok", "timestamp": 1630589248569, "user_tz": -330, "elapsed": 1193, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitS8CqtpOcM8tH8itmtSfpDm13rQsGdSUSpu-5KQ=s64", "userId": "07832619671270321867"}} outputId="6d0cca8e-fdb8-4d3d-fe58-fac7b10543ff"
from sklearn import tree
tree.plot_tree(DecTr)
# + colab={"base_uri": "https://localhost:8080/"} id="Z1MJJBAIRZO8" executionInfo={"status": "ok", "timestamp": 1630589276193, "user_tz": -330, "elapsed": 441, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitS8CqtpOcM8tH8itmtSfpDm13rQsGdSUSpu-5KQ=s64", "userId": "07832619671270321867"}} outputId="b04d3e42-caf4-4051-a9f6-fec56fab69f1"
# Q.1
target_pred1 = DecTr.predict([(1,2,1,0)])
print(target_pred1)
# + colab={"base_uri": "https://localhost:8080/"} id="SbV0dJEKRZO9" executionInfo={"status": "ok", "timestamp": 1630589295791, "user_tz": -330, "elapsed": 786, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitS8CqtpOcM8tH8itmtSfpDm13rQsGdSUSpu-5KQ=s64", "userId": "07832619671270321867"}} outputId="a84c8d68-5528-4065-ad0e-047b7467c0bb"
# Q.2
target_pred2 = DecTr.predict([(2,0,0,1)])
print(target_pred2)
| Lab-04/130_04_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Alternative method for querying Landsat 8 P/R from lat/lon
#
# The idea: each unique P/R combination has a footprint that forms a polygon. So we can use the point-in-polygon algorithm to see what P/Rs have the point of interest covered.
import pandas as pd
import geopandas as gpd
import numpy as np
from shapely.geometry import Point
from shapely.geometry import Polygon
from shapely.geometry import MultiPolygon
# ls8 = pd.read_csv('./LS8.csv')
ls8 = pd.read_excel('./LS8_cornerPts.xlsx')
# +
# ls8_s = ls8[:10]
ls8_s = ls8
ls8_s
# -
# Polygons that run across the antimeridian need a special attention.... use MultiPolygon.
# +
def check_crossing(lon_list):
"""
Checks if the antimeridian is crossed.
"""
import itertools
return any(abs(pair[0] - pair[1]) > 180.0 for pair in itertools.combinations(lon_list, 2))
# return abs(lon2 - lon1) > 180.0
# if any(abs(x) > 180.0 for x in [lon1, lon2, lon3, lon4]):
# raise ValueError("longitudes must be in degrees [-180.0, 180.0]")
# +
# for record in ls8_s:
# print(record.lat_UR)
# for xy in zip(ls8_s.lon_UL, ls8_s.lat_UL):
# print(xy)
geometry_2 = []
for index, row in ls8_s.iterrows():
lon_list = [row.lon_UL, row.lon_UR, row.lon_LR, row.lon_LL]
if check_crossing(lon_list):
set1 = [x % 360.0 for x in lon_list]
set2 = [x % -360.0 for x in lon_list]
poly1 = Polygon([(set1[0], row.lat_UL), (set1[1], row.lat_UR), (set1[2], row.lat_LR), (set1[3], row.lat_LL)])
poly2 = Polygon([(set2[0], row.lat_UL), (set2[1], row.lat_UR), (set2[2], row.lat_LR), (set2[3], row.lat_LL)])
feature_geometry = MultiPolygon([poly1, poly2])
else:
feature_geometry = Polygon([(row.lon_UL, row.lat_UL), (row.lon_UR, row.lat_UR), (row.lon_LR, row.lat_LR), (row.lon_LL, row.lat_LL)])
geometry_2.append(feature_geometry)
# geometry = [Point(xy) for xy in zip(ls8_s.lon_UL, ls8_s.lat_UL)]
# geometry_2 = Polygon(zip(record )) for record in ls8_s
ls8_s_gdf = gpd.GeoDataFrame(ls8_s, geometry=geometry_2)
ls8_s_gdf = ls8_s_gdf.drop(['lat_UL', 'lon_UL', 'lat_UR', 'lon_UR', 'lat_LL', 'lon_LL', 'lat_LR', 'lon_LR'], axis=1)
ls8_s_gdf
# geometry
# zip(lon_point_list, lat_point_list))
# -
# +
def lquery_alt(point_geometry, polygon_data):
# points_geometry: a shapely.geometry.Point object or a 2-element list showing [lon, lat]
# return: print P/R
import geopandas as gpd
from shapely.geometry import Point
# from shapely.geometry import mapping
# shapefile = gpd.read_file(shp_filename)
# poly_geometries = [shapefile.loc[i]['geometry'] for i in range(len(shapefile))]
# poly_geometries = polygon_data['geometry']
if type(point_geometry) is Point:
pt = point_geometry
else:
pt = Point(point_geometry)
# if type(points_geometry) is gpd.GeoDataFrame:
# pt_gs = gpd.GeoSeries(points_geometry.geometry)
# elif type(points_geometry) is gpd.GeoSeries:
# pt_gs = points_geometry
# else:
# pt_geometries = [Point(xy) for xy in zip(points_geometry[:, 0], points_geometry[:, 1])]
# pt_gs = gpd.GeoSeries(pt_geometries)
# idx = []
for idx, row in polygon_data.iterrows():
if pt.within(row.geometry):
print(row.path, row.row)
# print(pt.within(row.geometry))
# print(pt_gs.within(row.geometry))
# if pt_gs.within(row.geometry):
# print(row.path, row.row)
# if idx is None:
# idx = pt_gs.within(single_poly)
# else:
# tmp = pt_gs.within(single_poly)
# idx = np.logical_or(idx, tmp)
# return idx
# +
# %%timeit
lquery_alt(Point([np.random.sample()*180, np.random.sample()*88]), ls8_s_gdf)
# for ind, row in ls8_s_gdf.iterrows():
# print(ind, row.geometry)
# -
lquery_alt(Point([179.8242, 71.0027]), ls8_s_gdf)
# !aws s3 ls s3://landsat-pds/c1/L8/094/010/ --no-sign-request
# !aws s3 ls s3://landsat-pds/c1/L8/094/010/LC08_L1TP_094010_20200924_20201005_01_T1/ --no-sign-request
# ## Can we process these cloud images directly without downlonding them first?
#
# Quick answer: Yes with Rasterio, but No with Gdal.
#
# This is not good news if we want to do feature tracking on the cloud because all of the packages that I know use gdal to handle raster images.
# +
import rasterio
from rasterio.plot import show
dataset = rasterio.open('https://landsat-pds.s3.amazonaws.com/c1/L8/094/010/LC08_L1TP_094010_20200924_20201005_01_T1/LC08_L1TP_094010_20200924_20201005_01_T1_B8.TIF')
show(dataset)
# +
print(type(dataset))
rio_array = dataset.read(1)
print(rio_array.shape)
rio_array[10000:10100, 10000:10100]
# dataset.shape
# +
from osgeo import gdal
gdal.GetDriverByName('DODS').Deregister()
gd_dataset = gdal.Open('https://landsat-pds.s3.amazonaws.com/c1/L8/094/010/LC08_L1TP_094010_20200924_20201005_01_T1/LC08_L1TP_094010_20200924_20201005_01_T1_B8.TIF')
band = gd_dataset.GetRasterBand(1)
# -
band
# gdal can't deal URL directly!
# +
# from usgs import api
# def submit_where_query():
# # USGS uses numerical codes to identify queryable fields
# # To see which fields are queryable for a specific dataset,
# # send off a request to dataset-fields
# fields = api.dataset_fields('LANDSAT_8', 'EE')
# for field in fields:
# print(field)
# # WRS Path happens to have the field id 10036
# where = {
# 10036: '043'
# }
# scenes = api.search('LANDSAT_8', 'EE', where=where, start_date='2015-04-01', end_date='2015-05-01', max_results=10, extended=True)
# for scene in scenes:
# print(scene)
# submit_where_query()
# +
# def points_in_polygon(points_geometry, polygon_data):
# # points_geometry: N-by-2 np array or GeoDataFram or GeoSeries defining the geometry of points
# # shp_filename: (multi-)polygon shapefile name
# # Both datasets should have the SAME CRS!
# # return: np mask array showing where the targeted points are.
# import geopandas as gpd
# from shapely.geometry import Point
# # from shapely.geometry import mapping
# # shapefile = gpd.read_file(shp_filename)
# # poly_geometries = [shapefile.loc[i]['geometry'] for i in range(len(shapefile))]
# poly_geometries = polygon_data['geometry']
# if type(points_geometry) is gpd.GeoDataFrame:
# pt_gs = gpd.GeoSeries(points_geometry.geometry)
# elif type(points_geometry) is gpd.GeoSeries:
# pt_gs = points_geometry
# else:
# pt_geometries = [Point(xy) for xy in zip(points_geometry[:, 0], points_geometry[:, 1])]
# pt_gs = gpd.GeoSeries(pt_geometries)
# idx = None
# for single_poly in poly_geometries:
# if idx is None:
# idx = pt_gs.within(single_poly)
# else:
# tmp = pt_gs.within(single_poly)
# idx = np.logical_or(idx, tmp)
# return idx
# +
# shp_filename = '/media/whyj/Data/Document/Projects/Atacama/flood_region_broad_wgs84.shp'
# shapefile = gpd.read_file(shp_filename)
# poly_geometries = [shapefile.loc[i]['geometry'] for i in range(len(shapefile))]
# for single_poly in poly_geometries:
# print(single_poly)
# type(single_poly)
# -
# ## Test with USGS API
# +
import landsatxplore.api
# Initialize a new API instance and get an access key
api = landsatxplore.api.API("username", "********")
# +
# Request
scenes = api.search(
dataset='LANDSAT_8_C1',
latitude=71.0027,
longitude=179.8242,
start_date='2020-01-01',
end_date='2021-01-01',
max_cloud_cover=100)
print('{} scenes found.'.format(len(scenes)))
for scene in scenes:
print(scene['acquisitionDate'])
print(scene)
# api.logout()
# -
# OK -- you have to download first and unzip the data for sure.
| examples/Alt_Landsat8_Fetch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import importlib.util
try:
import cirq
except ImportError:
print("installing cirq...")
# !pip install --quiet cirq
print("installed cirq.")
try:
import quimb
except ImportError:
print("installing cirq[contrib]...")
# !pip install --quiet cirq[contrib]
print("installed cirq[contrib].")
# -
# # Contract a Grid Circuit
# Shallow circuits on a planar grid with low-weight observables permit easy contraction.
# ### Imports
# +
import numpy as np
import networkx as nx
import cirq
import quimb
import quimb.tensor as qtn
from cirq.contrib.svg import SVGCircuit
import cirq.contrib.quimb as ccq
# +
# %matplotlib inline
from matplotlib import pyplot as plt
import seaborn as sns
sns.set_style('ticks')
plt.rc('axes', labelsize=16, titlesize=16)
plt.rc('xtick', labelsize=14)
plt.rc('ytick', labelsize=14)
plt.rc('legend', fontsize=14, title_fontsize=16)
# +
# theme colors
QBLUE = '#1967d2'
QRED = '#ea4335ff'
QGOLD = '#fbbc05ff'
QGREEN = '#34a853ff'
QGOLD2 = '#ffca28'
QBLUE2 = '#1e88e5'
# -
# ## Make an example circuit topology
# We'll use entangling gates according to this topology and compute the value of an observable on the red nodes.
# +
width = 3
height = 4
graph = nx.grid_2d_graph(width, height)
rs = np.random.RandomState(52)
nx.set_edge_attributes(graph, name='weight',
values={e: np.round(rs.uniform(), 2) for e in graph.edges})
zz_inds = ((width//2, (height//2-1)), (width//2, (height//2)))
nx.draw_networkx(graph,
pos={n:n for n in graph.nodes},
node_color=[QRED if node in zz_inds else QBLUE for node in graph.nodes])
# -
# ### Circuit
qubits = [cirq.GridQubit(*n) for n in graph]
circuit = cirq.Circuit(
cirq.H.on_each(qubits),
ccq.get_grid_moments(graph),
cirq.Moment([cirq.rx(0.456).on_each(qubits)]),
)
SVGCircuit(circuit)
# ### Observable
ZZ = cirq.Z(cirq.GridQubit(*zz_inds[0])) * cirq.Z(cirq.GridQubit(*zz_inds[1]))
ZZ
# ### The contraction
# The value of the observable is $\langle 0 | U^\dagger (ZZ) U |0 \rangle$.
tot_c = ccq.circuit_for_expectation_value(circuit, ZZ)
SVGCircuit(tot_c)
# ## We can simplify the circuit
# By cancelling the "forwards" and "backwards" part of the circuit that are outside of the light-cone of the observable, we can reduce the number of gates to consider --- and sometimes the number of qubits involved at all. To see this in action, run the following cell and then keep re-running the following cell to watch gates disappear from the circuit.
compressed_c = tot_c.copy()
print(len(list(compressed_c.all_operations())), len(compressed_c.all_qubits()))
# **(try re-running the following cell to watch the circuit get smaller)**
# +
ccq.MergeNQubitGates(n_qubits=2).optimize_circuit(compressed_c)
ccq.MergeNQubitGates(n_qubits=1).optimize_circuit(compressed_c)
cirq.DropNegligible(tolerance=1e-6).optimize_circuit(compressed_c)
cirq.DropEmptyMoments().optimize_circuit(compressed_c)
print(len(list(compressed_c.all_operations())), len(compressed_c.all_qubits()))
SVGCircuit(compressed_c)
# -
# ### Utility function to fully-simplify
#
# We provide this utility function to fully simplify a circuit.
ccq.simplify_expectation_value_circuit(tot_c)
SVGCircuit(tot_c)
# simplification might eliminate qubits entirely for large graphs and
# shallow `p`, so re-get the current qubits.
qubits = sorted(tot_c.all_qubits())
print(len(qubits))
# ## Turn it into a Tensor Netowork
#
# We explicitly "cap" the tensor network with `<0..0|` bras so the entire thing contracts to the expectation value $\langle 0 | U^\dagger (ZZ) U |0 \rangle$.
# +
tensors, qubit_frontier, fix = ccq.circuit_to_tensors(
circuit=tot_c, qubits=qubits)
end_bras = [
qtn.Tensor(
data=quimb.up().squeeze(),
inds=(f'i{qubit_frontier[q]}_q{q}',),
tags={'Q0', 'bra0'}) for q in qubits
]
tn = qtn.TensorNetwork(tensors + end_bras)
tn.graph(color=['Q0', 'Q1', 'Q2'])
plt.show()
# -
# ### `rank_simplify` effectively folds together 1- and 2-qubit gates
#
# In practice, using this is faster than running the circuit optimizer to remove gates that cancel themselves, but please benchmark for your particular use case.
tn.rank_simplify(inplace=True)
tn.graph(color=['Q0', 'Q1', 'Q2'])
# ### The tensor contraction path tells us how expensive this will be
path_info = tn.contract(get='path-info')
path_info.opt_cost / int(3e9) # assuming 3gflop, in seconds
path_info.largest_intermediate * 128 / 8 / 1024 / 1024 / 1024 # gb
# ### Do the contraction
zz = tn.contract(inplace=True)
zz = np.real_if_close(zz)
print(zz)
# ## Big Circuit
# +
width = 8
height = 8
graph = nx.grid_2d_graph(width, height)
rs = np.random.RandomState(52)
nx.set_edge_attributes(graph, name='weight',
values={e: np.round(rs.uniform(), 2) for e in graph.edges})
zz_inds = ((width//2, (height//2-1)), (width//2, (height//2)))
nx.draw_networkx(graph,
pos={n:n for n in graph.nodes},
node_color=[QRED if node in zz_inds else QBLUE for node in graph.nodes])
# -
qubits = [cirq.GridQubit(*n) for n in graph]
circuit = cirq.Circuit(
cirq.H.on_each(qubits),
ccq.get_grid_moments(graph),
cirq.Moment([cirq.rx(0.456).on_each(qubits)]),
)
ZZ = cirq.Z(cirq.GridQubit(*zz_inds[0])) * cirq.Z(cirq.GridQubit(*zz_inds[1]))
ZZ
ccq.tensor_expectation_value(circuit, ZZ)
| cirq/contrib/quimb/Contract-a-Grid-Circuit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="Y28BJkCOo05i"
# + [markdown] id="l-lvJgGVCT_d"
# ## Question - 1:
#
# #### Create a numpy array starting from 2 till 50 with a stepsize of 3.
# + id="0MtdHExjo3xj" outputId="fb8f7d34-0642-433e-a118-9f9fe4c5a9d7" colab={"base_uri": "https://localhost:8080/"}
# importing numpy with alias name
import numpy as np
# creating a array using numpy arange
arr = np.arange(2, 50, 3)
# printing array
print(arr)
# + id="iXsUyD56nf1d"
# + [markdown] id="Iuu9BcU6o7CH"
# ## Question - 2
#
# #### Accept two lists of 5 elements each from the user. Convert them to numpy arrays. Concatenate these arrays and print it. Also sort these arrays and print it.
# + id="ZbVJG91lsw2x" outputId="3a01351a-b05e-45a9-ce6d-91e91ebe1dbd" colab={"base_uri": "https://localhost:8080/"}
# importing numpy with alias name
import numpy as np
# creating a 2 empty list:
lst_1 = []
lst_2 = []
# getting values for lst_1 from user:
for i in range(5):
x=int(input("\nEnter the value for lst_1 : "))
lst_1.append(x)
# getting values for lst_2 from user:
for i in range(5):
x=int(input("\nEnter the value for lst_2 : "))
lst_2.append(x)
# converting list into array:
arr_1 = np.array(lst_1)
arr_2 = np.array(lst_2)
# concatenating 2 array:
arr = np.concatenate((arr_1, arr_2))
print("\nArray value for concatenated arr :",arr)
# sorting up concatenated array
sortArr = np.sort(arr)
print("\nSorted Array for concatenated arr :",sortArr)
# + id="95S3hI5ze8Va"
# + [markdown] id="LXS3gu0wgcJb"
# ## Question - 3:
#
# #### Write a code snippet to find the dimensions of a ndarray and its size.
# + [markdown] id="PAoJSV4D9qbn"
# #### snippet for finding dimension:
#
# ArrayName.ndim
#
# #### snippet for finding size:
#
# ArrayName.size
#
# + id="A-mc68nxghPe" outputId="114272d9-efc5-41d3-ee1b-ba0307e468ba" colab={"base_uri": "https://localhost:8080/"}
# importing numpy
import numpy as np
# creating a 2d array
arr = np.array([[1,2,3],[1,3,5]])
# Printing array dimensions (axes)
print("\nDimensions in array : ", arr.ndim)
# Printing size of array
print("\nSize of array : ", arr.size)
# + id="-5C5Io-OkV0s"
# + [markdown] id="SUeCpqiIkK-b"
# ## Question - 4:
#
# #### How to convert a 1D array into a 2D array? Demonstrate with the help of a code snippet
# ##### Hint: np.newaxis, np.expand_dims
# + [markdown] id="5XQbYiU7G425"
# ### newaxis snippet:
#
# ArrayName[numpy.newaxis, :] // row major
#
# ArrayName[:, numpy.newaxis] // column major
# + id="9y52Rwh9hC5L" outputId="789a4f54-7ced-44e3-9b26-97613131cdf0" colab={"base_uri": "https://localhost:8080/"}
# importing numpy
import numpy as np
# creating a 1d array
arr = np.arange(10)
print("\nShape of Array :",arr.shape)
# converting using newaxis in row wise
row_arr = arr[np.newaxis, :]
print("\nShape of 2d Array in row major :",row_arr.shape)
# converting using newaxis in column wise
col_arr = arr[:, np.newaxis]
print("\nShape of 2d Array in column major :",col_arr.shape)
# + id="o07qezARjORu"
# + [markdown] id="861Ag-19KT6W"
# ### expand_dims snippet:
#
# numpy.expand_dims(ArrayName, axis = 0) // row major
#
# numpy.expand_dims(ArrayName, axis = 1) // column major
# + id="KP4hiF-9lapO" outputId="3a12bef6-3f2c-4706-ab03-7a8f51955525" colab={"base_uri": "https://localhost:8080/"}
# importing numpy
import numpy as np
# creating a 1d array
arr = np.arange(10)
print("\nShape of Array :",arr.shape)
# converting using newaxis in row wise
row_arr = np.expand_dims(arr, axis=0)
print("\nShape of 2d Array in row major :",row_arr.shape)
# converting using newaxis in column wise
col_arr = np.expand_dims(arr, axis=1)
print("\nShape of 2d Array in column major :",col_arr.shape)
# + id="6WSK3fF1K4rc"
# + [markdown] id="WrQzooVRLcox"
# ## Question - 5:
#
# #### Consider two square numpy arrays. Stack them vertically and horizontally.
#
# ##### Hint: Use vstack(), hstack()
# + id="yIvVbspoLkL3" outputId="027cec44-195d-456a-d2ad-d87524e54bb0" colab={"base_uri": "https://localhost:8080/"}
# importing numpy
import numpy as np
# creating a array with square values
arr_1 = np.square([1, 2, 3, 4, 5])
arr_2 = np.square([6, 7, 8, 9, 10])
# stacking and printing the value
print("\nHorizontal Append :", np.hstack((arr_1, arr_2)))
print("\nVertical Append :", np.vstack((arr_1, arr_2)))
# + id="1XcixDsRLi7Y"
# + [markdown] id="RRj09BcYM22m"
# ## Question - 6:
#
# ##### How to get unique items and counts of unique items?
#
# unique method in numpy is used to get the unique values from the list and return count attribute of unique method return the count of unique value from the list.
# + id="D_L6jirvOIzZ" outputId="9d02d835-74ec-4a12-c8e9-55b20cc935ea" colab={"base_uri": "https://localhost:8080/"}
# importing numpy
import numpy as np
# creating a array
arr_1 = np.array([1, 4, 9, 16, 25, 1, 4, 2, 3, 2, 5, 6, 5, 7, 8, 10])
# getting unique charater and its count
unique, counts = np.unique(arr_1, return_counts=True)
# making a
arr = np.asarray((unique, counts)).T
print(arr)
# + id="r1Ci2n2LOguj"
| Day 3/Day_3_Assignment.ipynb |