code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Pearson Correlation
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from pylab import rcParams
import seaborn as sb
import scipy
from scipy.stats.stats import pearsonr
# -
# %matplotlib inline
rcParams['figure.figsize'] = 5, 4
sb.set_style('whitegrid')
address = '../datasets/staandelamp_realistic.json'
data = pd.read_json(address)
data.head()
sb.pairplot(data[0:20])
name = data['name']
state = data['state']
time = data['time']
pearsonr_coefficient, p_value = pearsonr(time, state)
print('Pr %0.3f' % pearsonr_coefficient)
corr = data.corr()
corr
sb.heatmap(corr)
subset = data[0:100]
# +
plt.plot(subset['time'], subset['name'], 'o', color='black');
# +
colors = subset['state']
plt.scatter(subset['time'], subset['name'], c=colors, s=10)
plt.colorbar();
# -
data.hist()
plt.show()
names = ['name', 'state', 'time']
data.plot(kind='density', subplots=True, layout=(3,3), sharex=False)
plt.show()
data.plot(kind='box', subplots=True, layout=(3,3), sharex=False, sharey=False)
plt.show()
from pandas.plotting import scatter_matrix
scatter_matrix(data)
plt.show()
sb.lmplot(x='name', y='time', fit_reg=False, data=subset);
data.groupby(['name']).corr()
data['time'].values
| old/linear_correlations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# # Tracing Executions
#
# In this chapter, we show how to _observe program state during an execution_ – a prerequisite for logging and interactive debugging. Thanks to the power of Python, we can do this in a few lines of code.
# + slideshow={"slide_type": "skip"}
from bookutils import YouTubeVideo
YouTubeVideo("UYAvCl-5NGY")
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# **Prerequisites**
#
# * You should have read the [Introduction to Debugging](Intro_Debugging.ipynb).
# * Knowing a bit of _Python_ is helpful for understanding the code examples in the book.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
import bookutils
# + slideshow={"slide_type": "skip"}
from bookutils import quiz
# + slideshow={"slide_type": "skip"}
import Intro_Debugging
# + [markdown] slideshow={"slide_type": "skip"}
# ## Synopsis
# <!-- Automatically generated. Do not edit. -->
#
# To [use the code provided in this chapter](Importing.ipynb), write
#
# ```python
# >>> from debuggingbook.Tracer import <identifier>
# ```
#
# and then make use of the following features.
#
#
# This chapter provides a `Tracer` class that allows to log events during program execution. The advanced subclass `EventTracer` allows to restrict logs to specific conditions. Logs are shown only while the given `condition` holds:
#
# ```python
# >>> with EventTracer(condition='line == 223 or len(out) >= 6'):
# >>> remove_html_markup('<b>foo</b>bar')
# ...
# # s = '<b>foo</b>bar', function = 'remove_html_markup', line = 243, tag = False, quote = False, out = 'foobar', c = 'r'
# 243 for c in s:
# # line = 255
# 255 return out
# remove_html_markup() returns 'foobar'
#
# ```
# It also allows to restrict logs to specific events. Log entries are shown only if one of the given `events` changes its value:
#
# ```python
# >>> with EventTracer(events=["c == '/'"]):
# >>> remove_html_markup('<b>foo</b>bar')
# ...
# Calling remove_html_markup(s = '<b>foo</b>bar', function = 'remove_html_markup', line = 238)
# ...
# # line = 244, tag = False, quote = False, out = '', c = '<'
# 244 assert tag or not quote
# ...
# # tag = True, out = 'foo', c = '/'
# 244 assert tag or not quote
# ...
# # c = 'b'
# 244 assert tag or not quote
#
# ```
# `Tracer` and `EventTracer` classes allow for subclassing and further customization.
#
# 
#
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Tracing Python Programs
#
# How do debugging tools access the state of a program during execution? For _interpreted_ languages such as Python, this is a simple task. If a language is interpreted, it is typically fairly easy to control execution and to inspect state – since this is what the interpreter is doing already anyway. Debuggers are then implemented on top of _hooks_ that allow to interrupt execution and access program state.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# Python makes such a hook available in the function `sys.settrace()`. You invoke it with a *tracing function* that will be called at every line executed, as in
#
# ```python
# sys.settrace(traceit)
# ```
#
# Such a tracing function is convenient, as it simply traces _everything_. In contrast to an interactive debugger, where you have to select which aspect of the execution you're interested in, you can just print out a long trace into an *execution log*, to examine it later.
#
# This tracing function takes the format
# + slideshow={"slide_type": "skip"}
from types import FrameType, TracebackType
# + slideshow={"slide_type": "subslide"}
# ignore
from typing import Any, Optional, Callable, Dict, List, Type, TextIO, cast
# + slideshow={"slide_type": "fragment"}
def traceit(frame: FrameType, event: str, arg: Any) -> Optional[Callable]:
...
# + [markdown] slideshow={"slide_type": "fragment"}
# Here, `event` is a string telling what has happened in the program – for instance,
#
# * `'line'` – a new line is executed
# * `'call'` – a function just has been called
# * `'return'` – a function returns
# + [markdown] slideshow={"slide_type": "subslide"}
# The `frame` argument holds the current execution frame – that is, the function and its local variables:
#
# * `frame.f_lineno` – the current line
# * `frame.f_locals` – the current variables (as a Python dictionary)
# * `frame.f_code` – the current code (as a Code object), with attributes such as
# * `frame.f_code.co_name` – the name of the current function
# + [markdown] slideshow={"slide_type": "fragment"}
# We can thus get a *trace* of the program by simply printing out these values:
# + slideshow={"slide_type": "fragment"}
def traceit(frame: FrameType, event: str, arg: Any) -> Optional[Callable]: # type: ignore
print(event, frame.f_lineno, frame.f_code.co_name, frame.f_locals)
# + [markdown] slideshow={"slide_type": "subslide"}
# The return value of the trace function is the function to be executed at the next event – typically, this is the function itself:
# + slideshow={"slide_type": "fragment"}
def traceit(frame: FrameType, event: str, arg: Any) -> Optional[Callable]: # type: ignore
print(event, frame.f_lineno, frame.f_code.co_name, frame.f_locals)
return traceit
# + [markdown] slideshow={"slide_type": "fragment"}
# Let us try this out on the `remove_html_markup()` function introduced in the [Introduction to Debugging](Intro_Debugging.ipynb):
# + slideshow={"slide_type": "skip"}
from Intro_Debugging import remove_html_markup
# + slideshow={"slide_type": "skip"}
import inspect
# + slideshow={"slide_type": "skip"}
from bookutils import print_content
# + slideshow={"slide_type": "subslide"}
content, start_line_number = inspect.getsourcelines(remove_html_markup)
print_content(content="".join(content).strip(), filename='.py', start_line_number=start_line_number)
# + [markdown] slideshow={"slide_type": "subslide"}
# We define a variant `remove_html_markup_traced()` which turns on tracing, invokes `remove_html_markup()`, and turns tracing off again.
# + slideshow={"slide_type": "skip"}
import sys
# + slideshow={"slide_type": "fragment"}
def remove_html_markup_traced(s): # type: ignore
sys.settrace(traceit)
ret = remove_html_markup(s)
sys.settrace(None)
return ret
# + [markdown] slideshow={"slide_type": "fragment"}
# Here is what we get when we run `remove_html_markup_traced()`:
# * We first get a `call` event (showing the call of `remove_html_markup()`)
# * We then get various `line` events (for each line of `remove_html_markup()`)
# * In the end, we get a `return` event (showing the return from `remove_html_markup()`)
# + slideshow={"slide_type": "subslide"}
remove_html_markup_traced('xyz')
# + [markdown] slideshow={"slide_type": "subslide"}
# During the execution, we also see all local _variables_. As `remove_html_markup()` is called at the very beginning, the parameter `s` holds the argument `"xyz"`. As more local variables are being assigned, these show up in our dictionary of local variables.
# + [markdown] slideshow={"slide_type": "fragment"}
# We see how the variable `c` takes one character of the input string at a time; the `out` variable accumulates them. and the `tag` and `quote` flags stay unchanged throughout the execution.
# + [markdown] slideshow={"slide_type": "fragment"}
# An interesting aspect is that we can actually _access_ all these local variables as regular Python objects. We can, for instance, separately access the value of `c` by looking up `frame.f_locals['c']`:
# + slideshow={"slide_type": "subslide"}
def traceit(frame: FrameType, event: str, arg: Any) -> Optional[Callable]: # type: ignore
if 'c' in frame.f_locals:
value_of_c = frame.f_locals['c']
print(f"{frame.f_lineno:} c = {repr(value_of_c)}")
else:
print(f"{frame.f_lineno:} c is undefined")
return traceit
# + [markdown] slideshow={"slide_type": "fragment"}
# This allows us to specifically monitor individual variables:
# + slideshow={"slide_type": "subslide"}
remove_html_markup_traced('xyz')
# + [markdown] slideshow={"slide_type": "subslide"}
# This tracing capability is tremendously powerful – actually, it is one of the reasons this book uses Python all over the place. In most other languages, inspecting the program state during execution is much more complex than the handful of lines we have needed so far.
# + [markdown] slideshow={"slide_type": "fragment"}
# To learn more about `sys.settrace()`, spend a moment to look up [its documentation in the Python reference](https://docs.python.org/3/library/sys.html).
# + slideshow={"slide_type": "fragment"}
# ignore
import math
# + slideshow={"slide_type": "subslide"}
quiz("What happens if the tracing function returns `None` while tracing function `f()`?"
" (You can also try this out yourself.)",
[
'Tracing stops for all functions;'
' the tracing function is no longer called',
'Tracing stops for `f()`: the tracing function is called when `f()` returns',
'Tracing stops for `f()` the rest of the execution: the tracing function'
' is no longer called for calls to `f()`',
'Nothing changes'
], "int(math.log(7.38905609893065))", globals())
# + [markdown] slideshow={"slide_type": "fragment"}
# Indeed, as listed in the documentation: if `sys.settrace()` returns `None`, then tracing stops for the current scope; tracing will resume when the current function returns. This can be helpful for momentarily disable (expensive) tracing.
# + [markdown] slideshow={"slide_type": "slide"}
# ## A Tracer Class
# + [markdown] slideshow={"slide_type": "fragment"}
# Let us refine our tracing function a bit. First, it would be nice if one could actually _customize_ tracing just as needed. To this end, we introduce a `Tracer` class that does all the formatting for us, and which can be _subclassed_ to allow for different output formats.
# + [markdown] slideshow={"slide_type": "fragment"}
# The `traceit()` method in `Tracer` is the same as above, and again is set up via `sys.settrace()`. It uses a `log()` method after the Python `print()` function.
# + [markdown] slideshow={"slide_type": "subslide"}
# The typical usage of `Tracer`, however, is as follows:
#
# ```python
# with Tracer():
# # Code to be traced
# ...
#
# # Code no longer traced
# ...
# ```
#
# When the `with` statement is encountered, the `__enter__()` method is called, which starts tracing. When the `with` block ends, the `__exit__()` method is called, and tracing is turned off. We take special care that the internal `__exit__()` method is not part of the trace, and that any other tracing function that was active before is being restored.
# + [markdown] slideshow={"slide_type": "subslide"}
# We build `Tracer` on top of a class named `StackInspector`, whose `our_frame()` and `is_internal_error()` methods us with providing better diagnostics in case of error.
# + slideshow={"slide_type": "skip"}
from StackInspector import StackInspector
# + slideshow={"slide_type": "subslide"}
class Tracer(StackInspector):
"""A class for tracing a piece of code. Use as `with Tracer(): block()`"""
def __init__(self, *, file: TextIO = sys.stdout) -> None:
"""Trace a block of code, sending logs to `file` (default: stdout)"""
self.original_trace_function: Optional[Callable] = None
self.file = file
def traceit(self, frame: FrameType, event: str, arg: Any) -> None:
"""Tracing function. To be overridden in subclasses."""
self.log(event, frame.f_lineno, frame.f_code.co_name, frame.f_locals)
def _traceit(self, frame: FrameType, event: str, arg: Any) -> Optional[Callable]:
"""Internal tracing function."""
if self.our_frame(frame):
# Do not trace our own methods
pass
else:
self.traceit(frame, event, arg)
return self._traceit
def log(self, *objects: Any,
sep: str = ' ', end: str = '\n',
flush: bool = True) -> None:
"""
Like `print()`, but always sending to `file` given at initialization,
and flushing by default.
"""
print(*objects, sep=sep, end=end, file=self.file, flush=flush)
def __enter__(self) -> Any:
"""Called at begin of `with` block. Turn tracing on."""
self.original_trace_function = sys.gettrace()
sys.settrace(self._traceit)
# This extra line also enables tracing for the current block
# inspect.currentframe().f_back.f_trace = self._traceit
return self
def __exit__(self, exc_tp: Type, exc_value: BaseException,
exc_traceback: TracebackType) -> Optional[bool]:
"""
Called at end of `with` block. Turn tracing off.
Return `None` if ok, not `None` if internal error.
"""
sys.settrace(self.original_trace_function)
# Note: we must return a non-True value here,
# such that we re-raise all exceptions
if self.is_internal_error(exc_tp, exc_value, exc_traceback):
return False # internal error
else:
return None # all ok
# + [markdown] slideshow={"slide_type": "subslide"}
# Here's how we use the `Tracer` class. You see that everything works as before, except that it is nicer to use:
# + slideshow={"slide_type": "subslide"}
with Tracer():
remove_html_markup("abc")
# + [markdown] slideshow={"slide_type": "slide"}
# ## Accessing Source Code
#
# We can now go and _extend_ the class with additional features. It would be nice if it could actually display the source code of the function being tracked, such that we know where we are. In Python, the function `inspect.getsource()` returns the source code of a function or module. Looking up
#
# ```python
# module = inspect.getmodule(frame.f_code)
# ```
#
# gives us the current module, and
#
# ```python
# inspect.getsource(module)
# ```
#
# gives us its source code. All we then have to do is to retrieve the current line.
# + [markdown] slideshow={"slide_type": "subslide"}
# To implement our extended `traceit()` method, we use a bit of a hack. The Python language requires us to define an entire class with all methods as a single, continuous unit; however, we would like to introduce one method after another. To avoid this problem, we use a special hack: Whenever we want to introduce a new method to some class `C`, we use the construct
#
# ```python
# class C(C):
# def new_method(self, args):
# pass
# ```
#
# This seems to define `C` as a subclass of itself, which would make no sense – but actually, it introduces a new `C` class as a subclass of the _old_ `C` class, and then shadowing the old `C` definition. What this gets us is a `C` class with `new_method()` as a method, which is just what we want. (`C` objects defined earlier will retain the earlier `C` definition, though, and thus must be rebuilt.)
# + [markdown] slideshow={"slide_type": "subslide"}
# Using this hack, we can now redefine the `traceit()` method. Our new tracer shows the current line as it is executed.
# + slideshow={"slide_type": "skip"}
import inspect
# + slideshow={"slide_type": "fragment"}
class Tracer(Tracer):
def traceit(self, frame: FrameType, event: str, arg: Any) -> None:
"""Tracing function; called at every line. To be overloaded in subclasses."""
if event == 'line':
module = inspect.getmodule(frame.f_code)
if module is None:
source = inspect.getsource(frame.f_code)
else:
source = inspect.getsource(module)
current_line = source.split('\n')[frame.f_lineno - 1]
self.log(frame.f_lineno, current_line)
# + slideshow={"slide_type": "subslide"}
with Tracer():
remove_html_markup("abc")
# + [markdown] slideshow={"slide_type": "slide"}
# ## Tracing Calls and Returns
#
# Next, we'd like to report calling and returning from functions. For the `return` event, `arg` holds the value being returned.
# + slideshow={"slide_type": "subslide"}
class Tracer(Tracer):
def traceit(self, frame: FrameType, event: str, arg: Any) -> None:
"""Tracing function. To be overridden in subclasses."""
if event == 'call':
self.log(f"Calling {frame.f_code.co_name}()")
if event == 'line':
module = inspect.getmodule(frame.f_code)
if module:
source = inspect.getsource(module)
if source:
current_line = source.split('\n')[frame.f_lineno - 1]
self.log(frame.f_lineno, current_line)
if event == 'return':
self.log(f"{frame.f_code.co_name}() returns {repr(arg)}")
# + slideshow={"slide_type": "subslide"}
with Tracer():
remove_html_markup("abc")
# + [markdown] slideshow={"slide_type": "slide"}
# ## Tracing Variable Changes
#
# Finally, we'd again like to report variables – but only those that have changed. To this end, we save a copy of the last reported variables in the class, reporting only the changed values.
# + slideshow={"slide_type": "subslide"}
class Tracer(Tracer):
def __init__(self, file: TextIO = sys.stdout) -> None:
"""
Create a new tracer.
If `file` is given, output to `file` instead of stdout.
"""
self.last_vars: Dict[str, Any] = {}
super().__init__(file=file)
def changed_vars(self, new_vars: Dict[str, Any]) -> Dict[str, Any]:
"""Track changed variables, based on `new_vars` observed."""
changed = {}
for var_name, var_value in new_vars.items():
if (var_name not in self.last_vars or
self.last_vars[var_name] != var_value):
changed[var_name] = var_value
self.last_vars = new_vars.copy()
return changed
# + [markdown] slideshow={"slide_type": "subslide"}
# Here's how this works: If variable `a` is set to 10 (and we didn't have it so far), it is marked as changed:
# + slideshow={"slide_type": "fragment"}
tracer = Tracer()
# + slideshow={"slide_type": "fragment"}
tracer.changed_vars({'a': 10})
# + [markdown] slideshow={"slide_type": "fragment"}
# If another variable `b` is added, and only `b` is changed, then only `b` is marked as changed:
# + slideshow={"slide_type": "fragment"}
tracer.changed_vars({'a': 10, 'b': 25})
# + [markdown] slideshow={"slide_type": "fragment"}
# If both variables keep their values, nothing changes:
# + slideshow={"slide_type": "fragment"}
tracer.changed_vars({'a': 10, 'b': 25})
# + [markdown] slideshow={"slide_type": "fragment"}
# But if new variables come along, they are listed again.
# + slideshow={"slide_type": "subslide"}
changes = tracer.changed_vars({'c': 10, 'd': 25})
changes
# + [markdown] slideshow={"slide_type": "fragment"}
# The following expression creates a comma-separated list of variables and values:
# + slideshow={"slide_type": "fragment"}
", ".join([var + " = " + repr(changes[var]) for var in changes])
# + [markdown] slideshow={"slide_type": "fragment"}
# We can now put all of this together in our tracing function, reporting any variable changes as we see them. Note how we exploit the fact that in a call, all variables have a "new" value; and when we return from a function, we explicitly delete the "last" variables.
# + slideshow={"slide_type": "subslide"}
class Tracer(Tracer):
def print_debugger_status(self, frame: FrameType, event: str, arg: Any) -> None:
"""Show current source line and changed vars"""
changes = self.changed_vars(frame.f_locals)
changes_s = ", ".join([var + " = " + repr(changes[var])
for var in changes])
if event == 'call':
self.log("Calling " + frame.f_code.co_name + '(' + changes_s + ')')
elif changes:
self.log(' ' * 40, '#', changes_s)
if event == 'line':
try:
module = inspect.getmodule(frame.f_code)
if module is None:
source = inspect.getsource(frame.f_code)
else:
source = inspect.getsource(module)
current_line = source.split('\n')[frame.f_lineno - 1]
except OSError as err:
self.log(f"{err.__class__.__name__}: {err}")
current_line = ""
self.log(repr(frame.f_lineno) + ' ' + current_line)
if event == 'return':
self.log(frame.f_code.co_name + '()' + " returns " + repr(arg))
self.last_vars = {} # Delete 'last' variables
def traceit(self, frame: FrameType, event: str, arg: Any) -> None:
"""Tracing function; called at every line. To be overloaded in subclasses."""
self.print_debugger_status(frame, event, arg)
# + [markdown] slideshow={"slide_type": "subslide"}
# Here's the resulting trace of `remove_html_markup()` for a more complex input. You can see that the tracing output allows us to see which lines are executed as well as the variables whose value changes.
# + slideshow={"slide_type": "subslide"}
with Tracer():
remove_html_markup('<b>x</b>')
# + [markdown] slideshow={"slide_type": "subslide"}
# As you see, even a simple function can create a long execution log. Hence, we will now explore how to focus tracing on particular _events_.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Conditional Tracing
#
# A log such as the above can very quickly become very messy – notably if executions take a long time, or if data structures become very complex. If one of our local variables were a list with 1,000 entries for instance, and were changed with each line, we'd be printing out the entire list with 1,000 entries for each step.
#
# We could still load the log into, say, a text editor or a database and then search for specific values, but this is still cumbersome – and expensive. A better alternative, however, is to have our tracer only log while specific _conditions_ hold.
# + [markdown] slideshow={"slide_type": "subslide"}
# To this end, we introduce a class `ConditionalTracer`, which gets a _conditional expression_ to be checked during executions. Only if this condition holds do we list the current status. With
#
# ```python
# with ConditionalTracer(condition='c == "z"'):
# remove_html_markup(...)
# ```
#
# we would obtain only the lines executed while `c` gets a value of `'z'`, and with
#
# ```python
# with ConditionalTracer(condition='quote'):
# remove_html_markup(...)
# ```
#
# we would obtain only the lines executed while `quote` is True. If we have multiple conditions, we can combine them into one using `and`, `or`, or `not`.
# + [markdown] slideshow={"slide_type": "subslide"}
# Our `ConditionalTracer` class stores the condition in its `condition` attribute:
# + slideshow={"slide_type": "fragment"}
class ConditionalTracer(Tracer):
def __init__(self, *, condition: Optional[str] = None, file: TextIO = sys.stdout) -> None:
"""Constructor. Trace all events for which `condition` (a Python expr) holds."""
if condition is None:
condition = 'False'
self.condition: str = condition
self.last_report: Optional[bool] = None
super().__init__(file=file)
# + [markdown] slideshow={"slide_type": "subslide"}
# Its `traceit()` function _evaluates_ `condition` and reports the current line only if it holds. To this end, it uses the Python `eval()` function which evaluates the condition in the context of the local variables of the program under test. If the condition gets set, we print out three dots to indicate the elapsed time.
# + slideshow={"slide_type": "fragment"}
class ConditionalTracer(ConditionalTracer):
def eval_in_context(self, expr: str, frame: FrameType) -> Optional[bool]:
try:
cond = eval(expr, None, frame.f_locals)
except NameError: # (yet) undefined variable
cond = None
return cond
# + [markdown] slideshow={"slide_type": "fragment"}
# The `do_report()` function returns True if the status is to be reported:
# + slideshow={"slide_type": "subslide"}
class ConditionalTracer(ConditionalTracer):
def do_report(self, frame: FrameType, event: str, arg: Any) -> Optional[bool]:
return self.eval_in_context(self.condition, frame)
# + [markdown] slideshow={"slide_type": "fragment"}
# We put everything together in our `traceit()` function:
# + slideshow={"slide_type": "fragment"}
class ConditionalTracer(ConditionalTracer):
def traceit(self, frame: FrameType, event: str, arg: Any) -> None:
report = self.do_report(frame, event, arg)
if report != self.last_report:
if report:
self.log("...")
self.last_report = report
if report:
self.print_debugger_status(frame, event, arg)
# + [markdown] slideshow={"slide_type": "subslide"}
# Here's an example. We see that `quote` is set only while the three characters `b`, `a`, and `r` are processed (as should be).
# + slideshow={"slide_type": "subslide"}
with ConditionalTracer(condition='quote'):
remove_html_markup('<b title="bar">"foo"</b>')
# + slideshow={"slide_type": "subslide"}
quiz("What happens if the condition contains a syntax error?",
[
"The tracer stops, raising an exception",
"The tracer continues as if the condition were `True`",
"The tracer continues as if the condition were `False`",
], '393 % 7')
# + [markdown] slideshow={"slide_type": "fragment"}
# Here's the answer, illustrated in two examples. For syntax errors, we indeed get an exception:
# + slideshow={"slide_type": "skip"}
from ExpectError import ExpectError
# + slideshow={"slide_type": "subslide"}
with ExpectError(SyntaxError):
with ConditionalTracer(condition='2 +'):
remove_html_markup('<b title="bar">"foo"</b>')
# + [markdown] slideshow={"slide_type": "subslide"}
# If a variable is undefined, though, the condition evaluates to False:
# + slideshow={"slide_type": "fragment"}
with ExpectError():
with ConditionalTracer(condition='undefined_variable'):
remove_html_markup('<b title="bar">"foo"</b>')
# + [markdown] slideshow={"slide_type": "fragment"}
# We can also have the log focus on _particular code locations_ only. To this end, we add the pseudo-variables `function` and `line` to our evaluation context, which can be used within our condition to refer to the current function name or line. Then, we invoke the original `eval_cond()` as above.
# + slideshow={"slide_type": "fragment"}
class ConditionalTracer(ConditionalTracer):
def eval_in_context(self, expr: str, frame: FrameType) -> Any:
frame.f_locals['function'] = frame.f_code.co_name
frame.f_locals['line'] = frame.f_lineno
return super().eval_in_context(expr, frame)
# + [markdown] slideshow={"slide_type": "subslide"}
# Again, here is an example. We focus on the parts of the function where the `out` variable is being set:
# + slideshow={"slide_type": "subslide"}
with ConditionalTracer(condition="function == 'remove_html_markup' and line >= 237"):
remove_html_markup('xyz')
# + [markdown] slideshow={"slide_type": "subslide"}
# Using `line` and `function` in conditions is equivalent to conventional _breakpoints_ in interactive debuggers. We will reencounter them in the next chapter.
# + slideshow={"slide_type": "fragment"}
quiz("If the program under test contains a variable named `line`, "
"which `line` does the condition refer to?",
[
"`line` as in the debugger",
"`line` as in the program"
], '(326 * 27 == 8888) + 1')
# + [markdown] slideshow={"slide_type": "slide"}
# ## Watching Events
#
# As an alternative to conditional logging, we may also be interested to exactly trace when a variable not only _has_ a particular value, but also when it _changes_ its value.
#
# To this end, we set up an `EventTracer` class that _watches_ when some event takes place. It takes a list of expressions ("events") and evaluates them for each line; if any event changes its value, we log the status.
# + [markdown] slideshow={"slide_type": "subslide"}
# With
#
# ```python
# with EventTracer(events=['tag', 'quote']):
# remove_html_markup(...)
# ```
#
# for instance, we would get a listing of all lines where `tag` or `quote` change their value; and with
#
# ```python
# with EventTracer(events=['function']):
# remove_html_markup(...)
# ```
#
# we would obtain a listing of all lines where the current function changes.
# + [markdown] slideshow={"slide_type": "subslide"}
# Our `EventTracer` class stores the list of events in its `events` attribute:
# + slideshow={"slide_type": "fragment"}
class EventTracer(ConditionalTracer):
"""Log when a given event expression changes its value"""
def __init__(self, *, condition: Optional[str] = None,
events: List[str] = [], file: TextIO = sys.stdout) -> None:
"""Constructor. `events` is a list of expressions to watch."""
self.events = events
self.last_event_values: Dict[str, Any] = {}
super().__init__(file=file, condition=condition)
# + [markdown] slideshow={"slide_type": "fragment"}
# Its `events_changed()` function _evaluates_ the individual events and checks if they change.
# + slideshow={"slide_type": "subslide"}
class EventTracer(EventTracer):
def events_changed(self, events: List[str], frame: FrameType) -> bool:
"""Return True if any of the observed `events` has changed"""
change = False
for event in events:
value = self.eval_in_context(event, frame)
if (event not in self.last_event_values or
value != self.last_event_values[event]):
self.last_event_values[event] = value
change = True
return change
# + [markdown] slideshow={"slide_type": "fragment"}
# We hook this into `do_report()`, the method that determines whether a line should be shown.
# + slideshow={"slide_type": "subslide"}
class EventTracer(EventTracer):
def do_report(self, frame: FrameType, event: str, arg: Any) -> bool:
"""Return True if a line should be shown"""
return (self.eval_in_context(self.condition, frame) or
self.events_changed(self.events, frame))
# + [markdown] slideshow={"slide_type": "fragment"}
# This allows us to track, for instance, how `quote` and `tag` change their values over time.
# + slideshow={"slide_type": "subslide"}
with EventTracer(events=['quote', 'tag']):
remove_html_markup('<b title="bar">"foo"</b>')
# + [markdown] slideshow={"slide_type": "subslide"}
# Continuously monitoring variable values at execution time is equivalent to the concept of *watchpoints* in interactive debuggers.
# + [markdown] slideshow={"slide_type": "fragment"}
# With this, we have all we need for observing what happens during execution: We can explore the entire state, and we can evaluate conditions and events we are interested in. In the next chapter, we will see how to turn these capabilities into an interactive debugger, where we can query all these things interactively.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Efficient Tracing
#
# While our framework is very flexible (and can still be extended further), it also is _slow_, since we have to evaluate all conditions and events for every single line of the program. Just how slow are things? We can easily measure this.
# + slideshow={"slide_type": "skip"}
from Timer import Timer
# + slideshow={"slide_type": "fragment"}
runs = 1000
# + [markdown] slideshow={"slide_type": "fragment"}
# Here's the untraced execution time in seconds:
# + slideshow={"slide_type": "fragment"}
with Timer() as t:
for i in range(runs):
remove_html_markup('<b title="bar">"foo"</b>')
untraced_execution_time = t.elapsed_time()
untraced_execution_time
# + [markdown] slideshow={"slide_type": "fragment"}
# And here's the _traced_ execution time:
# + slideshow={"slide_type": "subslide"}
with Timer() as t:
for i in range(runs):
with EventTracer():
remove_html_markup('<b title="bar">"foo"</b>')
traced_execution_time = t.elapsed_time()
traced_execution_time
# + [markdown] slideshow={"slide_type": "fragment"}
# We see that the _traced_ execution time is several hundred times slower:
# + slideshow={"slide_type": "fragment"}
traced_execution_time / untraced_execution_time
# + [markdown] slideshow={"slide_type": "fragment"}
# We can still speed up our implementation somewhat, but still will get nowhere near the untraced execution time.
# + [markdown] slideshow={"slide_type": "subslide"}
# There is a trick, though, that allows us to execute programs at full speed while being traced. Rather than _dynamically_ checking at run time whether a condition is met, we can also _statically_ inject appropriate code into the program under test. This way, the non-traced code is executed at normal speed.
# + [markdown] slideshow={"slide_type": "fragment"}
# There is a downside, though: This only works if the condition to be checked is limited to specific _locations_ – because it is precisely these locations where we insert our tracing code. With this limitation, though, _static_ tracing can speed up things significantly.
# + [markdown] slideshow={"slide_type": "fragment"}
# How does static code injection work? The trick involves _rewriting_ the program code to insert special _debugging statements_ at the given locations. This way, we do not need to use the tracing function at all.
# + [markdown] slideshow={"slide_type": "subslide"}
# The following `insert_tracer()` function demonstrates this. It takes a function as well as a list of _breakpoint_ lines where to insert tracing statements. At each given line, it injects the code
# + slideshow={"slide_type": "fragment"}
TRACER_CODE = \
"TRACER.print_debugger_status(inspect.currentframe(), 'line', None); "
# + [markdown] slideshow={"slide_type": "fragment"}
# into the function definition, which calls into this tracer:
# + slideshow={"slide_type": "fragment"}
TRACER = Tracer()
# + [markdown] slideshow={"slide_type": "fragment"}
# `insert_tracer()` then _compiles_ the resulting code into a new "traced" function, which it then returns.
# + slideshow={"slide_type": "subslide"}
def insert_tracer(function: Callable, breakpoints: List[int] = []) -> Callable:
"""Return a variant of `function` with tracing code `TRACER_CODE` inserted
at each line given by `breakpoints`."""
source_lines, starting_line_number = inspect.getsourcelines(function)
breakpoints.sort(reverse=True)
for given_line in breakpoints:
# Set new source line
relative_line = given_line - starting_line_number + 1
inject_line = source_lines[relative_line - 1]
indent = len(inject_line) - len(inject_line.lstrip())
source_lines[relative_line - 1] = ' ' * indent + TRACER_CODE + inject_line.lstrip()
# Rename function
new_function_name = function.__name__ + "_traced"
source_lines[0] = source_lines[0].replace(function.__name__, new_function_name)
new_def = "".join(source_lines)
# For debugging
print_content(new_def, '.py', start_line_number=starting_line_number)
# We keep original source and filename to ease debugging
prefix = '\n' * starting_line_number # Get line number right
new_function_code = compile(prefix + new_def, function.__code__.co_filename, 'exec')
exec(new_function_code)
new_function = eval(new_function_name)
return new_function
# + [markdown] slideshow={"slide_type": "subslide"}
# Here's an example: inserting two breakpoints in (relative) Lines 7 and 18 of `remove_html_markup()` results in the following (rewritten) definition of `remove_html_markup_traced()`:
# + slideshow={"slide_type": "fragment"}
_, remove_html_markup_starting_line_number = inspect.getsourcelines(remove_html_markup)
breakpoints = [(remove_html_markup_starting_line_number - 1) + 7,
(remove_html_markup_starting_line_number - 1) + 18]
# + slideshow={"slide_type": "subslide"}
remove_html_markup_traced = insert_tracer(remove_html_markup, breakpoints)
# + [markdown] slideshow={"slide_type": "subslide"}
# If we execute the statically instrumented `remove_html_markup_traced()`, we obtain the same output as when using a dynamic tracer. Note that the source code listed shows the original code; the injected calls into `TRACER` do not show up.
# + slideshow={"slide_type": "subslide"}
with Timer() as t:
remove_html_markup_traced('<b title="bar">"foo"</b>')
static_tracer_execution_time = t.elapsed_time()
# + [markdown] slideshow={"slide_type": "subslide"}
# How fast is the static tracer compared with the dynamic tracer? This is the execution time of the above code:
# + slideshow={"slide_type": "fragment"}
static_tracer_execution_time
# + [markdown] slideshow={"slide_type": "fragment"}
# Compare this with the equivalent dynamic tracer:
# + slideshow={"slide_type": "subslide"}
line7 = (remove_html_markup_starting_line_number - 1) + 7
line18 = (remove_html_markup_starting_line_number - 1) + 18
with Timer() as t:
with EventTracer(condition=f'line == {line7} or line == {line18}'):
remove_html_markup('<b title="bar">"foo"</b>')
dynamic_tracer_execution_time = t.elapsed_time()
dynamic_tracer_execution_time
# + slideshow={"slide_type": "subslide"}
dynamic_tracer_execution_time / static_tracer_execution_time
# + [markdown] slideshow={"slide_type": "fragment"}
# We see that the static tracker is several times faster – an advantage that will only increase further as more non-traced code is executed. If our code looks like this:
# + slideshow={"slide_type": "fragment"}
def some_extreme_function(s: str) -> None:
... # Long-running function
remove_html_markup(s)
# + [markdown] slideshow={"slide_type": "fragment"}
# and we then execute it with
# + slideshow={"slide_type": "subslide"}
with EventTracer(condition=f"function=='remove_html_markup' and line == {line18}"):
some_extreme_function("foo")
# + [markdown] slideshow={"slide_type": "fragment"}
# we will spend quite some time.
# + slideshow={"slide_type": "subslide"}
quiz("In the above example, "
"where is the `EventTracer.traceit()` function called?",
[
"When `some_extreme_function()` returns",
"For each line of `some_extreme_function()`",
"When `remove_html_markup()` returns",
"For each line of `remove_html_markup()`"
], "[ord(c) - 100 for c in 'efgh']")
# + [markdown] slideshow={"slide_type": "fragment"}
# Indeed: Stepping line by line through some function can be pretty expensive, as every call, line, and return of `some_extreme_function()` and `remove_html_markup()` is tracked.
# + [markdown] slideshow={"slide_type": "subslide"}
# On the other hand, the static tracker is limited to conditions that refer to a _specific location in the code._ If we want to check whether some variable changes, for instance, we have to perform a (nontrivial) static analysis of the code to determine possible locations for a change. If a variable is changed indirectly through references or pointers (a common risk in system-level languages like C), there is no alternative to actually watching its value after each instruction.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Tracing Binary Executables
#
# Debuggers that act on binary code (say, code compiled from C) operate in a similar way as our "static" tracer: They take a location in the binary code and replace its instruction with a _break instruction_ that interrupts execution, returning control to the debugger. The debugger then replaces the break instruction with the original instruction before resuming execution.
#
# If the code cannot be altered (for instance, because it is in read-only memory), however, then debuggers resort to the "dynamic" tracing method, executing one instruction at a time and checking the program counter for its current value after each step.
#
# To provide a minimum of efficient support, some processor architectures, such as x86, provide *hardware breakpoints*. Programmers (or more precisely, debugging tools) can define a set of specific values for the program counter to watch, and if the program counter reaches one of these values, execution is interrupted to return to the debugger. Likewise, *hardware watchpoints* will check specific memory locations at run time for changes and given values. There are also hardware watchpoints that break when a specific memory location is read from. Both hardware watchpoints and hardware breakpoints allow a limited tracking of stopping conditions while still maintaining original execution speed – and the best debugging tools will use a mix of static tracing, dynamic tracing, and hardware tracing.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Synopsis
# + [markdown] slideshow={"slide_type": "fragment"}
# This chapter provides a `Tracer` class that allows to log events during program execution. The advanced subclass `EventTracer` allows to restrict logs to specific conditions. Logs are shown only while the given `condition` holds:
# + slideshow={"slide_type": "fragment"}
with EventTracer(condition='line == 223 or len(out) >= 6'):
remove_html_markup('<b>foo</b>bar')
# + [markdown] slideshow={"slide_type": "subslide"}
# It also allows to restrict logs to specific events. Log entries are shown only if one of the given `events` changes its value:
# + slideshow={"slide_type": "subslide"}
with EventTracer(events=["c == '/'"]):
remove_html_markup('<b>foo</b>bar')
# + [markdown] slideshow={"slide_type": "subslide"}
# `Tracer` and `EventTracer` classes allow for subclassing and further customization.
# + slideshow={"slide_type": "fragment"}
# ignore
from ClassDiagram import display_class_hierarchy
# + slideshow={"slide_type": "subslide"}
# ignore
display_class_hierarchy(EventTracer,
public_methods=[
Tracer.__init__,
Tracer.__enter__,
Tracer.__exit__,
Tracer.changed_vars,
Tracer.print_debugger_status,
ConditionalTracer.__init__,
EventTracer.__init__,
],
project='debuggingbook')
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Lessons Learned
#
# * Interpreted languages can provide _debugging hooks_ that allow to dynamically control program execution and access program state.
# * Tracing can be limited to specific conditions and events:
# * A _breakpoint_ is a condition referring to a particular location in the code.
# * A _watchpoint_ is an event referring to a particular state change.
# * Compiled languages allow to _instrument_ code at compile time, injecting code that allows to hand over control to a tracing or debugging tool.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Next Steps
#
# In the next chapter, we will see how to
#
# * [leverage our tracing infrastructure for interactive debugging](Debugger.ipynb)
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Background
#
# Debugging interfaces like Python `sys.settrace()` are common in all programming languages that provide support for interactive debugging, providing support for executing programs step by step and inspecting program state along the way.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Low-Level Debugging Interfaces
#
# The first set of interfaces considered takes place at a _low level_, allowing access to _machine level_ features. On Linux and other UNIX-like systems, the [ptrace()](https://en.wikipedia.org/wiki/Ptrace) system call provides a means by which one process (the 'tracer') may observe and control the execution of another process (the 'tracee'), and examine and change the tracee's memory and registers.
#
# `ptrace()` is a low-level interface, which allows to step over individual machine instructions and to read raw memory. In order to map instructions back to original statements and translate memory contents to variable values, compilers can include *debugging information* in the produced binaries, which debuggers then read out during a debugging session.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### High-Level Debugging Interfaces
#
# The second set of interfaces allows to access the program's internals using the concepts of the program – i.e. variables and code locations, as Python does. The [Java Debug Interface](https://docs.oracle.com/javase/8/docs/jdk/api/jpda/jdi/) (JDI) is a _high-level interface_ for implementing a debugger (or tracer) on top of Java. [This introduction to JDI](https://www.baeldung.com/java-debug-interface) shows how to build a debugger using this interface in a few steps.
#
# For JavaScript, Mozilla's [Debugger API](https://developer.mozilla.org/en-US/docs/Tools/Debugger-API) and Google's [chrome.debugger API](https://developer.chrome.com/docs/extensions/reference/debugger/) similarly allow to trace and inspect program execution.
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Exercises
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# ### Exercise 1: Exception Handling
#
# So far, we have only seen execution of lines in individual functions. But if a function raises an exception, we also may want to catch and report this. Right now, an exception is being raised right through our tracer, interrupting the trace.
# + slideshow={"slide_type": "fragment"}
def fail() -> float:
return 2 / 0
# + slideshow={"slide_type": "subslide"}
with Tracer():
try:
fail()
except Exception:
pass
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"} solution2="hidden" solution2_first=true
# Extend the `Tracer` class (or the `EventTracer` subclasses) such that exceptions (event type `'exception'`) are properly traced, too, say as
#
# ```
# fail() raises ZeroDivisionError: division by zero
# ```
#
# See the Python documentation for `sys.settrace()`.
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** Simply extend `print_debugger_status()`:
# + cell_style="split" slideshow={"slide_type": "skip"} solution2="hidden"
class Tracer(Tracer):
def print_debugger_status(self, frame: FrameType, event: str, arg: Any) -> None:
if event == 'exception':
exception, value, tb = arg
self.log(f"{frame.f_code.co_name}() "
f"raises {exception.__name__}: {value}")
else:
super().print_debugger_status(frame, event, arg)
# + slideshow={"slide_type": "skip"} solution2="hidden"
with Tracer():
try:
fail()
except Exception:
pass
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# ### Exercise 2: Syntax-Based Instrumentation
#
# Adding instrumentation to source code is a complicated business, notably because it is not always easy to determine where and how to instrument. If a Python line starts with
#
# ```python
# if condition:
# ```
#
# where should one insert code to instrument it?
# + [markdown] slideshow={"slide_type": "subslide"}
# A much more elegant way to instrument code is to add instrumentation _after the code has already been parsed_. Python code, like most other code, is first _parsed_ into an intermediate tree-like structure (called an *abstract syntax tree*, or *AST*). This AST can then be inspected and manipulated, before a second step compiles it into low-level instruction sequences to be executed.
# + [markdown] slideshow={"slide_type": "fragment"}
# Let us start with an example. Here is an AST resulting from parsing a very simple piece of code:
# + slideshow={"slide_type": "fragment"}
def foo(): # type: ignore
ret = 2 * 2
return ret
# + slideshow={"slide_type": "subslide"}
source = inspect.getsource(foo)
print_content(source, '.py')
# + slideshow={"slide_type": "skip"}
import ast
# + slideshow={"slide_type": "skip"}
from bookutils import show_ast
# + slideshow={"slide_type": "fragment"}
tree = ast.parse(source)
# + slideshow={"slide_type": "fragment"}
show_ast(tree)
# + [markdown] slideshow={"slide_type": "fragment"}
# You see that the function `foo()` has a `FunctionDef` node with four children: The function name (`"foo"`), its arguments (`arguments`; currently empty), followed by the statements that make the function body – `Assign` for the assignment, `Return` for the `return` statement.
# + [markdown] slideshow={"slide_type": "subslide"}
# We obtain and manipulate the AST through the Python `ast` module. The [official Python `ast` reference](http://docs.python.org/3/library/ast) is complete, but a bit brief; the documentation ["Green Tree Snakes - the missing Python AST docs"](https://greentreesnakes.readthedocs.io/en/latest/) provides an excellent introduction.
# + [markdown] slideshow={"slide_type": "fragment"}
# To instrument the above code, we need to insert a new statement as a child to `FunctionDef` node.
# + slideshow={"slide_type": "skip"}
from ast import NodeTransformer, FunctionDef, fix_missing_locations, AST, Module
# + [markdown] slideshow={"slide_type": "fragment"}
# Here's the code we want to inject:
# + slideshow={"slide_type": "fragment"}
subtree_to_be_injected: AST = ast.parse("print('entering function')")
# + slideshow={"slide_type": "fragment"}
show_ast(subtree_to_be_injected)
# + [markdown] slideshow={"slide_type": "fragment"}
# The root of an `ast.parse()` tree actually is a `Module` node; we go directly to its child, which is the `Expr` node we want to inject.
# + slideshow={"slide_type": "fragment"}
subtree_to_be_injected = cast(Module, subtree_to_be_injected).body[0]
# + [markdown] slideshow={"slide_type": "subslide"}
# To inject the code, we use the `NodeTransformer` class as described in the Python `ast` documentation. We vist all function definitions (`FunctionDef`) and replace them with a new function definition in which the `body` gets an additional child – namely our subtree to be injected.
# + slideshow={"slide_type": "fragment"}
class InjectPass(NodeTransformer):
def visit_FunctionDef(self, node: FunctionDef) -> AST:
return FunctionDef(
name=node.name,
args=node.args,
body=[subtree_to_be_injected] + node.body,
decorator_list=node.decorator_list,
returns=node.returns
)
# + slideshow={"slide_type": "fragment"}
new_tree = fix_missing_locations(InjectPass().visit(tree))
# + [markdown] slideshow={"slide_type": "subslide"}
# This is what our new tree looks like:
# + slideshow={"slide_type": "fragment"}
show_ast(new_tree)
# + [markdown] slideshow={"slide_type": "fragment"}
# This is what the tree looks like when converted back to source code:
# + slideshow={"slide_type": "fragment"}
new_source = ast.unparse(new_tree)
print_content(new_source, '.py')
# + [markdown] slideshow={"slide_type": "fragment"}
# We can now compile the new source into a function:
# + slideshow={"slide_type": "fragment"}
exec(new_source)
# + [markdown] slideshow={"slide_type": "fragment"}
# ... and happily invoke our instrumented function.
# + slideshow={"slide_type": "subslide"}
foo()
# + [markdown] slideshow={"slide_type": "fragment"}
# Your task is to implement a function `insert_tracer_ast(function, breakpoints)` that works like `insert_tracer()`, above, except that it uses this AST-based mechanism to inject debugging code into the given function.
| docs/notebooks/Tracer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ___
# <a href='https://cafe.naver.com/jmhonglab'><img src='https://lh3.googleusercontent.com/lY3ySXooSmwsq5r-mRi7uiypbo0Vez6pmNoQxMFhl9fmZJkRHu5lO2vo7se_0YOzgmDyJif9fi4_z0o3ZFdwd8NVSWG6Ea80uWaf3pOHpR4GHGDV7kaFeuHR3yAjIJjDgfXMxsvw=w2400'/></a>
# ___
# <center><em>Content Copyright by HongLab, Inc.</em></center>
# # 주피터 노트북으로 파이썬을 사용해봅시다!
#
# 주피터 노트북에서는 주로 두 가지 종류의 셀(Cell)들을 사용하게 됩니다.
#
# 1. 마크다운(Markdown) 셀에서는 [마크다운 언어](https://ko.wikipedia.org/wiki/%EB%A7%88%ED%81%AC%EB%8B%A4%EC%9A%B4)를 이용하여 서식이 있는 문서를 편리하게 작성할 수 있습니다.
# 1. 코드(Code) 셀에서는 파이썬 언어를 편리하게 사용할 수 있습니다.
#
# 처음에는 낯설게 느껴질 수도 있지만 곧 익숙해질거에요.
# ### 파이썬을 계산기처럼 사용해봅시다.
#
# 간단한 산술 연산을 해봅시다
#
# >`1+1`
1 + 1
# ### 내가 만들고 싶은 것을 출력해봅시다.
#
# > print('여기에 내가 만들고 싶은 것을 적어보세요')
| Chapter1/ex1_helloworld.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import decimal
import numpy as np
xmin = -10
xmax = 10
dx = 0.1
xlist = np.around(np.arange(xmin, xmax, dx), decimals=4)
ylist = 1 / xlist
plt.plot(xlist, ylist)
plt.show()
| 2021 Осенний семестр/Практическое задание 4_5/КонтрольныйВопрос2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Calculate projection effects in redmapper
# Here, given the runpos.fit files recieved from Eli, I can calculate projection effects.
#
# So far, Eli has ran his runpos code on SDSS DR8 and des y1a1. This notebook should be able to handle either.
#import everything
import fitsio, sys, os
import numpy as np
from scipy.optimize import minimize
import matplotlib.pyplot as plt
# %matplotlib inline
#Get the data
datapath = "data_files/dr8_run_0.4_runpos.fit"
data, header = fitsio.read(datapath, header=True)
lambda_true = data['LAMBDA_CHISQ']
z_true = data['Z_LAMBDA']
lambda_arrays = data['LAMBDA_CHISQS']
z_domain = np.loadtxt("data_files/redshift_list.txt")
print lambda_true.shape, z_true.shape
print z_domain.shape #This is how many redshifts we have sampled
print lambda_arrays.shape
# Now we define a comparison function, as well as a model for $\lambda(z)$.
# +
#Lambda model function
def lambda_model(z, sigmaz, z_peak, lambda_peak):
return lambda_peak * np.exp(-0.5*(z_peak - z)**2/sigmaz**2)
#The comparison function
def comparison(params, z_domain, lambda_array):
z_peak, sigmaz, lambda_peak = params #Pull out the parameters
#Some priors
if any(params < 0.0): return np.inf
if z_peak > 4.0: return np.inf #Way too high redshift
if lambda_peak > 1000: return np.inf #Way too big
if sigmaz < 0.005: return np.inf #Avoids numerical issues
lam_model = lambda_model(z_domain, sigmaz, z_peak, lambda_peak)
X = (lambda_array - lam_model)**2/lam_model**2
indices = (lambda_array > max(lambda_array)*0.5) * (z_domain < 0.32) #THE SECOND PART IS SDSS SPECIFIC
return sum(X[indices])
# -
# Try this procedure on a single cluster
x0 = [z_true[0], 0.03, lambda_true[0]]
result = minimize(comparison, x0=x0, args=(z_domain, lambda_arrays[0]), method='Nelder-Mead')
zbest, szbest, lambest = result['x']
print result
# Write a function that can create a comparison plot and test it on the cluster test.
# +
def make_plot(z_peak, sigmaz, lambda_peak, z_true, lambda_true, z_domain, lambda_array, save=False, index=None):
plt.plot(z_domain, lambda_array)
plt.scatter(z_true, lambda_true, marker='^')
plt.axvline(z_true, c='r')
plt.plot(z_domain, lambda_model(z_domain, sigmaz, z_peak, lambda_peak))
plt.ylabel("Richness", fontsize=24)
plt.xlabel("Redshift", fontsize=24)
if index is not None:
plt.title("Cluster %d"%index)
plt.ylim(-10, max(plt.gca().get_ylim()))
return
make_plot(zbest, szbest, lambest, z_true[0], lambda_true[0], z_domain, lambda_arrays[0], index=0)
# -
# Great. Let's proceed with every other cluster and save the outputs.
N = len(lambda_arrays)
sz_all = np.zeros((N)) #sigmaz for all clusters
zp_all = np.zeros((N)) #z_peak for all clusters
lp_all = np.zeros((N)) #lambda_peak for all clusters
for i in xrange(0,N):
x0 = [z_true[i], 0.03, lambda_true[i]]
result = minimize(comparison, x0=x0, args=(z_domain, lambda_arrays[i]), method='Nelder-Mead')
zp_all[i], sz_all[i], lp_all[i] = result['x']
if i%500==0: print "At cluster %d"%i
outpath = "output_files/"
dataset = 'dr8_0.4_'
np.savetxt(outpath+dataset+"zpeak.txt",zp_all)
np.savetxt(outpath+dataset+"sigmaz.txt",sz_all)
np.savetxt(outpath+dataset+"lambdapeak.txt",lp_all)
print "Saved successfully."
# Make the scatter plot of the resulting $\sigma_z(z)$ points.
#First split by richnesses
colors = ['g','r','b']
edges = [20, 30, 60, 9999]
for i in range(len(edges) - 1):
lo = edges[i]
hi = edges[i+1]
indices = (lambda_true >= lo) * (lambda_true < hi)
plt.scatter(z_true[indices], sz_all[indices], c=colors[i], label=r'$\lambda\in(%d,%d)$'%(lo,hi), alpha=0.2)
plt.xlabel('Redshift', fontsize=24)
plt.ylabel(r'$\sigma_z$', fontsize=24)
plt.legend(loc='upper left')
plt.ylim(0,.2)
# Looks like there is a trend, but it is difficult to tell without inspecting everything individually whether there are significant outliers.
| RM_Projection_Effects.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib widget
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
plt.style.use(['dark_background'])
# +
WEIGHTS_AND = (1, 1)
BIAS_AND = -1.25
WEIGHTS_OR = (1, 1)
BIAS_OR = -0.75
WEIGHTS_NOT = (0, -1)
BIAS_NOT = 0
def linear_combination(x, y, bias, weights):
return weights[0] * x + weights[1] * y + bias
def f(x, bias, weights):
return -((weights[0] * x + bias) / weights[1]) if weights[1] != 0 else weights[0] * x + bias
inputs = [(0, 0), (0, 1), (1, 0), (1, 1)]
boundary = np.linspace(1.5, -0.5, 2)
# AND
fig, axs = plt.subplots()
fig.suptitle('AND classifier')
for x, y in inputs:
axs.plot(x, y, 'bo') if linear_combination(x, y, BIAS_AND, WEIGHTS_AND) >= 0 else axs.plot(x, y, 'ro')
axs.plot(boundary, f(boundary, BIAS_AND, WEIGHTS_AND))
# OR
fig, axs = plt.subplots()
fig.suptitle('OR classifier')
for x, y in inputs:
axs.plot(x, y, 'bo') if linear_combination(x, y, BIAS_OR, WEIGHTS_OR) >= 0 else axs.plot(x, y, 'ro')
axs.plot(boundary, f(boundary, BIAS_OR, WEIGHTS_OR))
# NOT
fig, axs = plt.subplots()
fig.suptitle('NOT classifier')
for x, y in inputs:
# print(str(x) + ' ' + str(y) + ' ' + str(linear_combination(x, BIAS_NOT, WEIGHTS_NOT) < y))
axs.plot(x, y, 'bo') if linear_combination(x, y, BIAS_NOT, WEIGHTS_NOT) >= 0 else axs.plot(x, y, 'ro')
axs.plot(boundary, f(boundary, BIAS_NOT, WEIGHTS_NOT))
# -
| Machine Learning/PyTorch Scholarship/Lesson2 - Neural Networks Intro/and_or_not_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''examples'': conda)'
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import tensorflow as tf
from utils.plotting import plot_dataset
from tensorflow.keras import layers
from sklearn.model_selection import train_test_split
# +
# Load Dataset
df = pd.read_csv('data/ex.csv')
dataset = df.copy()
X = dataset.values
x_cords = dataset['x'].values
y_cords = dataset['y'].values
# -
plot_dataset(x_cords, y_cords, 'Full Dataset')
# ### Split the data into train and test
#
# Now split the dataset into a training set and a test set.
# Use the test set in the final evaluation of the model.
#
# ### Split features from labels
# Separate the target value, the "label", from the features. This label is the value that you will train the model to predict.
# +
# Train Test Split
x_train, x_test, y_train, y_test = train_test_split(x_cords, y_cords, test_size=0.20, random_state=np.random.seed(6))
X = np.stack((x_test, y_test), axis=1)
# -
# ### Linear regression
#
# Before building a DNN model, start with a linear regression.
# One Variable
#
# Start with a single-variable linear regression, to predict `y` from `x`.
#
# Training a model with `tf.keras` typically starts by defining the model architecture.
#
# In this case use a `keras.Sequential` model. This model represents a sequence of steps. In this case there are two steps:
#
# - Normalize the input `x`.
# - Apply a linear transformation $(y = mx+b)$ to produce 1 output using `layers.Dense`.
#
# The number of inputs can either be set by the `input_shape` argument, or automatically when the model is run for the first time.
#
# First create the horsepower `Normalization` layer:
# +
# Build the sequential model
model = tf.keras.Sequential([
layers.Dense(1, input_dim=1)
])
model.summary()
# -
# This model will predict `y` from `x`.
#
# Run the untrained model on the first 10 `x` values. The output won't be good, but you'll see that it has the expected shape, (10,1):
#
#
#
model.predict(x_cords[:10])
print(model.predict(x_cords[:10]))
# Once the model is built, configure the training procedure using the `Model.compile()` method. The most important arguments to compile are the `loss` and the `optimizer` since these define what will be optimized (`mean_absolute_error`) and how (using the `optimizers.Adam`).
model.compile(
optimizer=tf.optimizers.Adam(lr=1e-3),
loss='logcosh')
# Once the training is configured, use `Model.fit()` to execute the training:
# %%time
history = model.fit(
x_train, y_train,
epochs=100,
validation_split=0.2,
verbose=0)
# Visualize the model's training progress using the stats stored in the history object.
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
hist.tail()
# +
def plot_loss(history):
plt.plot(history.history['loss'], label='loss')
plt.plot(history.history['val_loss'], label='val_loss')
plt.ylim([0, 1])
plt.xlabel('Epoch')
plt.ylabel('Error [Y]')
plt.legend()
plt.grid(True)
plot_loss(history)
# +
test_results = model.evaluate(
x_test, y_test, verbose=0)
x = tf.linspace(-4.0, 4.0, 9)
y = model.predict(x)
def plot_model(x, y):
plt.scatter(x_train, y_train, label='Data')
plt.plot(x, y, color='k', label='Predictions')
plt.xlabel('x')
plt.ylabel('y')
plt.legend()
plot_model(x,y)
print(f"Loss: {test_results}")
# -
| regression/src/linear_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] id="JsZlBCxbro4e" colab_type="text"
# <small><i>June 2016 - This notebook was created by [<NAME>](http://www.maia.ub.es/~oriol). Source and [license](./LICENSE.txt) info are in the folder.</i></small>
# + [markdown] id="cwth3APAro4g" colab_type="text"
# # Unsupervised learning
# + [markdown] id="-01LWpsNro4h" colab_type="text"
# # + Autoencoders
# # + Pretraining
# # + Manifold learning
# # + Sparse coding
# + [markdown] id="Iew9RTaLro4i" colab_type="text"
# # Autoencoders
# + id="B6zIMNiB9s-A" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 122} outputId="e8ab45de-1947-4b54-c778-0c67fc14ee5b" executionInfo={"status": "ok", "timestamp": 1528923878417, "user_tz": -120, "elapsed": 2771, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
# !pip install tqdm
# + id="Wv6_JvzKro4j" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="fb1b34c8-576a-47e6-9ecc-653b7a40b89f" executionInfo={"status": "ok", "timestamp": 1528902374398, "user_tz": -120, "elapsed": 7741, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
# %reset
# %matplotlib inline
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
tf.reset_default_graph()
# + id="sDV_N0zaro4p" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 17} outputId="39ca0caa-f504-442a-efdf-9d08c963b1eb" executionInfo={"status": "ok", "timestamp": 1528902383023, "user_tz": -120, "elapsed": 623, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
# Network Parameters
n_input = 64
n_hidden_1 = 64 # 1st layer num features
n_hidden_2 = 32 # 2nd layer num features
n_hidden_3 = 10 # 3rd layer num features
n_hidden_4 = 10 # 4th layer num features
n_hidden_5 = 32 # 5th layer num features
n_hidden_6 = 64 # 6th layer num features
# Parameters
learning_rate = 1e-3
# + id="TiLbexbqro4r" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 17} outputId="c41812c8-1202-41e5-b6b1-20e1a87cbe22" executionInfo={"status": "ok", "timestamp": 1528902383804, "user_tz": -120, "elapsed": 642, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
# Graph inputs
x = tf.placeholder("float", [None, n_input], name='x')
y = tf.placeholder("float", [None, n_input], name='y')
keep_prob = tf.placeholder(tf.float32) #Input parameter: dropout probability
# Store layers weight & bias
c = 0.1
weights = {
'h1': tf.Variable(c*tf.random_normal([n_input, n_hidden_1]), name='W1'),
'h2': tf.Variable(c*tf.random_normal([n_hidden_1, n_hidden_2]), name='W2'),
'h3': tf.Variable(c*tf.random_normal([n_hidden_2, n_hidden_3]), name='W3'),
'h4': tf.Variable(c*tf.random_normal([n_hidden_3, n_hidden_4]), name='W4'),
'h5': tf.Variable(c*tf.random_normal([n_hidden_4, n_hidden_5]), name='W5'),
'h6': tf.Variable(c*tf.random_normal([n_hidden_5, n_hidden_6]), name='W6'),
'out': tf.Variable(c*tf.random_normal([n_hidden_6, n_input]))
}
biases = {
'b1': tf.Variable(c*tf.random_normal([n_hidden_1]), name='b1'),
'b2': tf.Variable(c*tf.random_normal([n_hidden_2]), name='b2'),
'b3': tf.Variable(c*tf.random_normal([n_hidden_3]), name='b3'),
'b4': tf.Variable(c*tf.random_normal([n_hidden_4]), name='b4'),
'b5': tf.Variable(c*tf.random_normal([n_hidden_5]), name='b5'),
'b6': tf.Variable(c*tf.random_normal([n_hidden_6]), name='b6'),
'out': tf.Variable(c*tf.random_normal([n_input]))
}
pre_layer_drop = tf.nn.dropout(x, keep_prob)
layer_1 = tf.nn.relu(tf.add(tf.matmul(pre_layer_drop, weights['h1']), biases['b1']))
layer_2 = tf.nn.relu(tf.add(tf.matmul(layer_1, weights['h2']), biases['b2']))
layer_3 = tf.nn.relu(tf.add(tf.matmul(layer_2, weights['h3']), biases['b3']))
layer_4 = tf.nn.relu(tf.add(tf.matmul(layer_3, weights['h4']), biases['b4']))
layer_5 = tf.nn.relu(tf.add(tf.matmul(layer_4, weights['h5']), biases['b5']))
layer_6 = tf.nn.relu(tf.add(tf.matmul(layer_5, weights['h6']), biases['b6']))
output = tf.add(tf.matmul(layer_6, weights['out']), biases['out'])
# + id="XPxUxuWcro4t" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 17} outputId="920440f8-3923-4e2d-c6bd-bb22de963998" executionInfo={"status": "ok", "timestamp": 1528902385319, "user_tz": -120, "elapsed": 632, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
#Reconstruction loss
loss = tf.reduce_mean(tf.pow(output - y, 2))
# + id="U2dFlafNro4w" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 17} outputId="f76bde76-83c6-4140-8157-7a143c1dfaf0" executionInfo={"status": "ok", "timestamp": 1528902386874, "user_tz": -120, "elapsed": 971, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
#Optimizer
opt = tf.train.AdamOptimizer(learning_rate).minimize(loss)
# + id="swivB1wmro4y" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 17} outputId="0e0f3bd4-79ce-446b-f87d-0377e88779f6" executionInfo={"status": "ok", "timestamp": 1528902401729, "user_tz": -120, "elapsed": 612, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
init = tf.global_variables_initializer()
# + id="hOh_78g4ro40" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 17} outputId="ab2bfedd-80cb-432b-82ac-55bc6088d352" executionInfo={"status": "ok", "timestamp": 1528902404793, "user_tz": -120, "elapsed": 658, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
# Load data
from sklearn.datasets import load_digits
data = load_digits()
idx = np.random.permutation(data.data.shape[0])
idx_train = idx[:-100]
idx_test = idx[-100:]
train = data.data[idx_train,:]
test = data.data[idx_test,:]
# + id="HgySTTXZro42" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 51} outputId="ae70c50a-85df-4698-e27d-cc0ea28bebf4" executionInfo={"status": "ok", "timestamp": 1528902525432, "user_tz": -120, "elapsed": 96655, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
sess = tf.Session()
sess.run(init)
import tqdm
training_epochs = 100000
display_step = 50
# Training cycle
cost = []
batch_size=16
for epoch in tqdm.tqdm(xrange(training_epochs)):
xs = train[np.random.randint(0,train.shape[0],batch_size),:]/16.
sess.run(opt, feed_dict={x: xs, y: xs, keep_prob: 0.8})
if epoch % display_step == 1:
cost.append(sess.run(loss, feed_dict={x: xs, y: xs, keep_prob: 1.}))
print "Optimization Finished!"
# + id="Uqrtk2JCro46" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 840} outputId="c328eb0f-166a-4e9f-e069-ac64d6381d90" executionInfo={"status": "ok", "timestamp": 1528902530345, "user_tz": -120, "elapsed": 1986, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
import matplotlib.pyplot as plt
plt.plot(cost[10:])
plt.show()
N = 8
xs = test[np.random.randint(0,test.shape[0],N),:]/16.
rec = sess.run(output, feed_dict={x: xs, keep_prob: 1.})
# %matplotlib inline
import matplotlib.pyplot as plt
f, axarr = plt.subplots(N, 2)
for i in xrange(N):
axarr[i,0].imshow(xs[i].reshape((8,8)),cmap='gray', interpolation='nearest')
axarr[i,1].imshow(rec[i].reshape((8,8)),cmap='gray', interpolation='nearest')
f.set_size_inches(10,10)
# + id="NTOG7O-oEmMh" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 374} outputId="8d44c41b-0546-4bff-d6ce-9ad92f004a65" executionInfo={"status": "ok", "timestamp": 1528927286536, "user_tz": -120, "elapsed": 1622, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
# !ls -l
# + [markdown] id="hCF8jSK2ro49" colab_type="text"
# # Check reconstruction
# + id="PABO5hNRro49" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 249} outputId="15c1c3d4-3727-4ea3-fe5b-829a29384404" executionInfo={"status": "error", "timestamp": 1528927710000, "user_tz": -120, "elapsed": 1496, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
import tensorflow as tf
import numpy as np
tf.reset_default_graph()
# Network Parameters
n_input = 64
n_hidden_1 = 64 # 1st layer num features
n_hidden_2 = 32 # 2nd layer num features
n_hidden_3 = 10 # 3nd layer num features
n_hidden_4 = 10 # 4nd layer num features
n_hidden_5 = 32 # 4nd layer num features
n_hidden_6 = 64 # 1st layer num features
# Parameters
learning_rate = 1e-3
# tf Graph input
x = tf.placeholder("float", [None, n_input], name='x')
y = tf.placeholder("float", [None, n_input], name='y')
# Store layers weight & bias
c = 0.1
weights = {
'h1': tf.Variable(c*tf.random_normal([n_input, n_hidden_1]), name='W1'),
'h2': tf.Variable(c*tf.random_normal([n_hidden_1, n_hidden_2]), name='W2'),
'h3': tf.Variable(c*tf.random_normal([n_hidden_2, n_hidden_3]), name='W3'),
'h4': tf.Variable(c*tf.random_normal([n_hidden_3, n_hidden_4]), name='W4'),
'h5': tf.Variable(c*tf.random_normal([n_hidden_4, n_hidden_5]), name='W5'),
'h6': tf.Variable(c*tf.random_normal([n_hidden_5, n_hidden_6]), name='W6'),
'out': tf.Variable(c*tf.random_normal([n_hidden_6, n_input]))
}
biases = {
'b1': tf.Variable(c*tf.random_normal([n_hidden_1]), name='b1'),
'b2': tf.Variable(c*tf.random_normal([n_hidden_2]), name='b2'),
'b3': tf.Variable(c*tf.random_normal([n_hidden_3]), name='b3'),
'b4': tf.Variable(c*tf.random_normal([n_hidden_4]), name='b4'),
'b5': tf.Variable(c*tf.random_normal([n_hidden_5]), name='b5'),
'b6': tf.Variable(c*tf.random_normal([n_hidden_6]), name='b6'),
'out': tf.Variable(c*tf.random_normal([n_input]))
}
keep_prob = tf.placeholder(tf.float32)
pre_layer_drop = tf.nn.dropout(x, keep_prob)
layer_1 = tf.nn.relu(tf.add(tf.matmul(pre_layer_drop, weights['h1']), biases['b1']))
layer_2 = tf.nn.relu(tf.add(tf.matmul(layer_1, weights['h2']), biases['b2']))
layer_3 = tf.nn.relu(tf.add(tf.matmul(layer_2, weights['h3']), biases['b3']))
layer_4 = tf.nn.relu(tf.add(tf.matmul(layer_3, weights['h4']), biases['b4']))
layer_5 = tf.nn.relu(tf.add(tf.matmul(layer_4, weights['h5']), biases['b5']))
layer_6 = tf.nn.relu(tf.add(tf.matmul(layer_5, weights['h6']), biases['b6']))
output = tf.add(tf.matmul(layer_6, weights['out']), biases['out'])
loss = tf.reduce_mean(tf.pow(output - y, 2))
opt = tf.train.AdamOptimizer(learning_rate).minimize(loss)
init = tf.global_variables_initializer()
# Add ops to save and restore all the variables.
saver = tf.train.Saver()
import tqdm
training_epochs = 100000
display_step = 50
from sklearn.datasets import load_digits
data = load_digits()
import pickle
from google.colab import files
print "Upload file ... digits_idx_part1_autoencoder.pkl"
uploaded = files.upload()
import io
idx_train,idx_test = pickle.load(io.StringIO(uploaded['digits_idx_part1_autoencoder.pkl'].decode('utf-8')))
train = data.data[idx_train,:]
test = data.data[idx_test,:]
sess = tf.Session()
sess.run(init)
print "Upload file ... autoencoder_digits_part1_autoencoder.ckpt"
uploaded = files.upload()
saver.restore(sess, "autoencoder_digits_part1_autoencoder.ckpt")
print("Model restored.")
N = 8
xs = test[np.random.randint(0,test.shape[0],N),:]/16.
rec = sess.run(output, feed_dict={x: xs, keep_prob: 1.})
# %matplotlib inline
import matplotlib.pyplot as plt
f, axarr = plt.subplots(N, 2)
for i in xrange(N):
axarr[i,0].imshow(xs[i].reshape((8,8)),cmap='gray', interpolation='nearest')
axarr[i,1].imshow(rec[i].reshape((8,8)),cmap='gray', interpolation='nearest')
f.set_size_inches(10,10)
# + [markdown] id="RD7dB16fro5A" colab_type="text"
# # Learning representations and pretraining
# + id="mXe8ZVtSro5B" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "Ly8gQ29weXJpZ2h0IDIwMTcgR<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 86} outputId="ee2f238a-02d1-47e0-f7d8-debcb2a2e293" executionInfo={"status": "ok", "timestamp": 1528928073893, "user_tz": -120, "elapsed": 11419, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
import tensorflow as tf
import numpy as np
tf.reset_default_graph()
# Network Parameters
n_input = 64
n_hidden_1 = 64 # 1st layer num features
n_hidden_2 = 32 # 2nd layer num features
n_hidden_3 = 10 # 3rd layer num features
n_hidden_4 = 10 # 4th layer num features
n_hidden_5 = 32 # 5th layer num features
n_hidden_6 = 64 # 6th layer num features
# Parameters
learning_rate = 1e-3
# tf Graph input
x = tf.placeholder("float", [None, n_input], name='x')
y = tf.placeholder("float", [None, n_input], name='y')
# Store layers weight & bias
c = 0.1
weights = {
'h1': tf.Variable(c*tf.random_normal([n_input, n_hidden_1]), name='W1'),
'h2': tf.Variable(c*tf.random_normal([n_hidden_1, n_hidden_2]), name='W2'),
'h3': tf.Variable(c*tf.random_normal([n_hidden_2, n_hidden_3]), name='W3'),
'h4': tf.Variable(c*tf.random_normal([n_hidden_3, n_hidden_4]), name='W4'),
'h5': tf.Variable(c*tf.random_normal([n_hidden_4, n_hidden_5]), name='W5'),
'h6': tf.Variable(c*tf.random_normal([n_hidden_5, n_hidden_6]), name='W6'),
'out': tf.Variable(c*tf.random_normal([n_hidden_6, n_input]))
}
biases = {
'b1': tf.Variable(c*tf.random_normal([n_hidden_1]), name='b1'),
'b2': tf.Variable(c*tf.random_normal([n_hidden_2]), name='b2'),
'b3': tf.Variable(c*tf.random_normal([n_hidden_3]), name='b3'),
'b4': tf.Variable(c*tf.random_normal([n_hidden_4]), name='b4'),
'b5': tf.Variable(c*tf.random_normal([n_hidden_5]), name='b5'),
'b6': tf.Variable(c*tf.random_normal([n_hidden_6]), name='b6'),
'out': tf.Variable(c*tf.random_normal([n_input]))
}
keep_prob = tf.placeholder(tf.float32)
pre_layer_drop = tf.nn.dropout(x, keep_prob)
layer_1 = tf.nn.relu(tf.add(tf.matmul(pre_layer_drop, weights['h1']), biases['b1']))
layer_2 = tf.nn.relu(tf.add(tf.matmul(layer_1, weights['h2']), biases['b2']))
layer_3 = tf.nn.relu(tf.add(tf.matmul(layer_2, weights['h3']), biases['b3']))
layer_4 = tf.nn.relu(tf.add(tf.matmul(layer_3, weights['h4']), biases['b4']))
layer_5 = tf.nn.relu(tf.add(tf.matmul(layer_4, weights['h5']), biases['b5']))
layer_6 = tf.nn.relu(tf.add(tf.matmul(layer_5, weights['h6']), biases['b6']))
output = tf.add(tf.matmul(layer_6, weights['out']), biases['out'])
loss = tf.reduce_mean(tf.pow(output - y, 2))
opt = tf.train.AdamOptimizer(learning_rate).minimize(loss)
init = tf.initialize_all_variables()
# Add ops to save and restore all the variables.
saver = tf.train.Saver()
import tqdm
training_epochs = 100000
display_step = 50
from sklearn.datasets import load_digits
data = load_digits()
from google.colab import files
print "Upload file ... digits_idx_part1_autoencoder.pkl"
uploaded = files.upload()
import io
idx_train,idx_test = pickle.load(io.StringIO(uploaded['digits_idx_part1_autoencoder.pkl'].decode('utf-8')))
train = data.data[idx_train,:]/16.
test = data.data[idx_test,:]/16.
train_y = data.target[idx_train]
test_y = data.target[idx_test]
# + id="I_wkby5Dro5E" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 51} outputId="2cddf9ed-6d45-4fc9-fabd-560102cff50e" executionInfo={"status": "ok", "timestamp": 1528928084137, "user_tz": -120, "elapsed": 559, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
sess = tf.Session()
sess.run(init)
saver.restore(sess, "autoencoder_digits_part1_autoencoder.ckpt")
print("Model restored.")
code = sess.run(layer_3, feed_dict={x: train, keep_prob: 1.})
# + id="e3BjwcJOro5G" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="4b19418b-ec9f-4fa8-d11d-0575d3489d85" executionInfo={"status": "ok", "timestamp": 1528928089565, "user_tz": -120, "elapsed": 2449, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
code.shape
# + id="l7U1JYpAro5J" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 119} outputId="b54a9d97-ab58-4596-b5dd-9922959b363b" executionInfo={"status": "ok", "timestamp": 1528928092743, "user_tz": -120, "elapsed": 686, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
from sklearn.ensemble import RandomForestClassifier
clf = RandomForestClassifier(3,random_state=0)
clf.fit(code,train_y)
clf2 = RandomForestClassifier(3,random_state=0)
clf2.fit(train,train_y)
# + id="B3-1SVx9ro5M" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 51} outputId="b8525b74-0030-46b9-99f3-d93f4cabfbbf" executionInfo={"status": "ok", "timestamp": 1528928095813, "user_tz": -120, "elapsed": 867, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
code = sess.run(layer_3, feed_dict={x: test, keep_prob: 1.})
print clf.score(code,test_y)
print clf2.score(test,test_y)
# + [markdown] id="LkuZNl9Oro5P" colab_type="text"
# # Sparse coding
# + id="X_bEpwiuro5R" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 189} outputId="30c1a3e5-efc3-480e-9fb5-402356cd7acc" executionInfo={"status": "ok", "timestamp": 1528928266418, "user_tz": -120, "elapsed": 33234, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
import tensorflow as tf
import numpy as np
tf.reset_default_graph()
# Network Parameters
n_input = 64
n_hidden_1 = 64 # 1st layer num features
n_hidden_2 = 128 # 2nd layer num features
n_hidden_3 = 256 # 3nd layer num features
n_hidden_4 = 256 # 4nd layer num features
n_hidden_5 = 128 # 4nd layer num features
n_hidden_6 = 64 # 1st layer num features
# Parameters
learning_rate = 1e-3
# tf Graph input
x = tf.placeholder("float", [None, n_input], name='x')
y = tf.placeholder("float", [None, n_input], name='y')
lmd = tf.placeholder("float",[], name = 'lambda')
# Store layers weight & bias
c = 0.1
weights = {
'h1': tf.Variable(c*tf.random_normal([n_input, n_hidden_1]), name='W1'),
'h2': tf.Variable(c*tf.random_normal([n_hidden_1, n_hidden_2]), name='W2'),
'h3': tf.Variable(c*tf.random_normal([n_hidden_2, n_hidden_3]), name='W3'),
'h4': tf.Variable(c*tf.random_normal([n_hidden_3, n_hidden_4]), name='W4'),
'h5': tf.Variable(c*tf.random_normal([n_hidden_4, n_hidden_5]), name='W5'),
'h6': tf.Variable(c*tf.random_normal([n_hidden_5, n_hidden_6]), name='W6'),
'out': tf.Variable(c*tf.random_normal([n_hidden_6, n_input]))
}
biases = {
'b1': tf.Variable(c*tf.random_normal([n_hidden_1]), name='b1'),
'b2': tf.Variable(c*tf.random_normal([n_hidden_2]), name='b2'),
'b3': tf.Variable(c*tf.random_normal([n_hidden_3]), name='b3'),
'b4': tf.Variable(c*tf.random_normal([n_hidden_4]), name='b4'),
'b5': tf.Variable(c*tf.random_normal([n_hidden_5]), name='b5'),
'b6': tf.Variable(c*tf.random_normal([n_hidden_6]), name='b6'),
'out': tf.Variable(c*tf.random_normal([n_input]))
}
keep_prob = tf.placeholder(tf.float32)
pre_layer_drop = tf.nn.dropout(x, keep_prob)
layer_1 = tf.nn.relu(tf.add(tf.matmul(pre_layer_drop, weights['h1']), biases['b1']))
layer_2 = tf.nn.relu(tf.add(tf.matmul(layer_1, weights['h2']), biases['b2']))
layer_3 = tf.nn.relu(tf.add(tf.matmul(layer_2, weights['h3']), biases['b3']))
layer_4 = tf.nn.relu(tf.add(tf.matmul(layer_3, weights['h4']), biases['b4']))
layer_5 = tf.nn.relu(tf.add(tf.matmul(layer_4, weights['h5']), biases['b5']))
layer_6 = tf.nn.relu(tf.add(tf.matmul(layer_5, weights['h6']), biases['b6']))
output = tf.add(tf.matmul(layer_6, weights['out']), biases['out'])
loss = tf.reduce_mean(tf.pow(output - y, 2))
loss2 = tf.reduce_mean(tf.abs(layer_3))
floss = loss + lmd * loss2
opt = tf.train.AdamOptimizer(learning_rate).minimize(floss)
init = tf.initialize_all_variables()
# Add ops to save and restore all the variables.
saver = tf.train.Saver()
import tqdm
training_epochs = 100000
display_step = 50
from sklearn.datasets import load_digits
data = load_digits()
idx = np.random.permutation(data.data.shape[0])
idx_train = idx[:-100]
idx_test = idx[-100:]
from google.colab import files
print "Upload file ... digits_idx_part2_sparse.pkl"
uploaded = files.upload()
import io
idx_train,idx_test = pickle.load(io.StringIO(uploaded['digits_idx_part2_sparse.pkl'].decode('utf-8')))
train = data.data[idx_train,:]
test = data.data[idx_test,:]
sess = tf.Session()
sess.run(init)
print "Upload file ... autoencoder_digits_part2_sparse.ckpt"
uploaded = files.upload()
saver.restore(sess, "autoencoder_digits_part2_sparse.ckpt")
print("Model restored.")
# + id="qVLvekN6ro5T" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 17} outputId="f87ca8c8-50ec-4e67-f4f2-11c5e16dc06d" executionInfo={"status": "ok", "timestamp": 1528928271533, "user_tz": -120, "elapsed": 840, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
code = sess.run(layer_3, feed_dict={x: train, keep_prob: 1.})
# + id="aUCb0_h8ro5V" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 901} outputId="472eac25-fd39-473d-f8ef-7af34eb21a29" executionInfo={"status": "ok", "timestamp": 1528928273386, "user_tz": -120, "elapsed": 832, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
code[1]
# + id="kOKexY95ro5Y" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 17} outputId="af18f872-e2d4-42be-e8ff-bd537418301b" executionInfo={"status": "ok", "timestamp": 1528928275665, "user_tz": -120, "elapsed": 585, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
non_zero=[]
for i in xrange(code.shape[0]):
non_zero.append(len(np.where(code[i]>1e-6)[0]))
# + id="d4M3h-_Uro5Z" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 316} outputId="5ed0ac37-6b92-4cd5-b11e-b6662fc3d099" executionInfo={"status": "ok", "timestamp": 1528928278703, "user_tz": -120, "elapsed": 763, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
# Histogram of coded values
plt.hist(non_zero)
# + id="lwNY3oXFro5c" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 17} outputId="2e181835-46a7-4c9c-9fcc-762bc223d393" executionInfo={"status": "ok", "timestamp": 1528928282444, "user_tz": -120, "elapsed": 564, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
code = sess.run(layer_3, feed_dict={x: test, keep_prob: 1.})
# + id="7c81DSyPro5d" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 238} outputId="a43acb5e-cb8f-4020-818b-1c148cc2f3c4" executionInfo={"status": "ok", "timestamp": 1528928285069, "user_tz": -120, "elapsed": 892, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
code
# + [markdown] id="GCK8WW_Uro5g" colab_type="text"
# ## From compressed to restored
# + id="LhbAUT6Rro5g" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "Ly8gQ29weXJpZ2h0IDIwMTcgR29vZ2xlIExMQwovLwovLyBMaWNlbnNlZCB1bmRlciB0aGUgQXBhY2hlIExpY2Vuc2UsIFZlcnNpb24gMi4wICh0aGUgIkxpY2Vuc2UiKTsKLy8geW91IG1heSBub3QgdXNlIHRoaXMgZmlsZSBleGNlcHQgaW4gY29tcGxpYW5jZSB3aXRoIHRoZSBMaWNlbnNlLgovLyBZb3UgbWF5IG9idGFpbiBhIGNvcHkgb2YgdGhlIExpY2Vuc2UgYXQKLy8KLy8gICAgICBodHRwOi8vd3d3LmFwYWNoZS5vcmcvbGljZW5zZXMvTElDRU5TRS0yLjAKLy8KLy8gVW5sZXNzIHJlcXVpcmVkIGJ5IGFwcGxpY2FibGUgbGF3IG9yIGFncmVlZCB0byBpbiB3cml0aW5nLCBzb2Z0d2FyZQovLyBkaXN0cmlidXRlZCB1bmRlciB0aGUgTGljZW5zZSBpcyBkaXN0cmlidXRlZCBvbiBhbiAiQVMgSVMiIEJBU0lTLAovLyBXSVRIT1VUIFdBUlJBTlRJRVMgT1IgQ09ORElUSU9OUyBPRiBBTlkgS0lORCwgZWl0aGVyIGV4cHJlc3Mgb3IgaW1wbGllZC4KLy8gU2VlIHRoZSBMaWNlbnNlIGZvciB0aGUgc3BlY2lmaWMgbGFuZ3VhZ2UgZ292ZXJuaW5nIHBlcm1pc3Npb25zIGFuZAovLyBsaW1pdGF0aW9ucyB1bmRlciB0aGUgTGljZW5zZS4KCi8qKgogKiBAZmlsZW92ZXJ2aWV3IEhlbHBlcnMgZm9yIGdvb2dsZS5jb2xhYiBQeXRob24gbW9kdWxlLgogKi8KKGZ1bmN0aW9uKHNjb3BlKSB7CmZ1bmN0aW9uIHNwYW4odGV4dCwgc3R5bGVBdHRyaWJ1dGVzID0ge30pIHsKICBjb25zdCBlbGVtZW50ID0gZG9jdW1lbnQuY3JlYXRlRWxlbWVudCgnc3BhbicpOwogIGVsZW1lbnQudGV4dENvbnRlbnQgPSB0ZXh0OwogIGZvciAoY29uc3Qga2V5IG9mIE9iamVjdC5rZXlzKHN0eWxlQXR0cmlidXRlcykpIHsKICAgIGVsZW1lbnQuc3R5bGVba2V5XSA9IHN0eWxlQXR0cmlidXRlc1trZXldOwogIH0KICByZXR1cm4gZWxlbWVudDsKfQoKLy8gTWF4IG51bWJlciBvZiBieXRlcyB3aGljaCB3aWxsIGJlIHVwbG9hZGVkIGF0IGEgdGltZS4KY29uc3QgTUFYX1BBWUxPQURfU0laRSA9IDEwMCAqIDEwMjQ7Ci8vIE1heCBhbW91bnQgb2YgdGltZSB0byBibG9jayB3YWl0aW5nIGZvciB0aGUgdXNlci4KY29uc3QgRklMRV9DSEFOR0VfVElNRU9VVF9NUyA9IDMwICogMTAwMDsKCmZ1bmN0aW9uIF91cGxvYWRGaWxlcyhpbnB1dElkLCBvdXRwdXRJZCkgewogIGNvbnN0IHN0ZXBzID0gdXBsb2FkRmlsZXNTdGVwKGlucHV0SWQsIG91dHB1dElkKTsKICBjb25zdCBvdXRwdXRFbGVtZW50ID0gZG9jdW1lbnQuZ2V0RWxlbWVudEJ5SWQob3V0cHV0SWQpOwogIC8vIENhY2hlIHN0ZXBzIG9uIHRoZSBvdXRwdXRFbGVtZW50IHRvIG1ha2UgaXQgYXZhaWxhYmxlIGZvciB0aGUgbmV4dCBjYWxsCiAgLy8gdG8gdXBsb2FkRmlsZXNDb250aW51ZSBmcm9tIFB5dGhvbi4KICBvdXRwdXRFbGVtZW50LnN0ZXBzID0gc3RlcHM7CgogIHJldHVybiBfdXBsb2FkRmlsZXNDb250aW51ZShvdXRwdXRJZCk7Cn0KCi8vIFRoaXMgaXMgcm91Z2hseSBhbiBhc3luYyBnZW5lcmF0b3IgKG5vdCBzdXBwb3J0ZWQgaW4gdGhlIGJyb3dzZXIgeWV0KSwKLy8gd2hlcmUgdGhlcmUgYXJlIG11bHRpcGxlIGFzeW5jaHJvbm91cyBzdGVwcyBhbmQgdGhlIFB5dGhvbiBzaWRlIGlzIGdvaW5nCi8vIHRvIHBvbGwgZm9yIGNvbXBsZXRpb24gb2YgZWFjaCBzdGVwLgovLyBUaGlzIHVzZXMgYSBQcm9taXNlIHRvIGJsb2NrIHRoZSBweXRob24gc2lkZSBvbiBjb21wbGV0aW9uIG9mIGVhY2ggc3RlcCwKLy8gdGhlbiBwYXNzZXMgdGhlIHJlc3VsdCBvZiB0aGUgcHJldmlvdXMgc3RlcCBhcyB0aGUgaW5wdXQgdG8gdGhlIG5leHQgc3RlcC4KZnVuY3Rpb24gX3VwbG9hZEZpbGVzQ29udGludWUob3V0cHV0SWQpIHsKICBjb25zdCBvdXRwdXRFbGVtZW50ID0gZG9jdW1lbnQuZ2V0RWxlbWVudEJ5SWQob3V0cHV0SWQpOwogIGNvbnN0IHN0ZXBzID0gb3V0cHV0RWxlbWVudC5zdGVwczsKCiAgY29uc3QgbmV4dCA9IHN0ZXBzLm5leHQob3V0cHV0RWxlbWVudC5sYXN0UHJvbWlzZVZhbHVlKTsKICByZXR1cm4gUHJvbWlzZS5yZXNvbHZlKG5leHQudmFsdWUucHJvbWlzZSkudGhlbigodmFsdWUpID0+IHsKICAgIC8vIENhY2hlIHRoZSBsYXN0IHByb21pc2UgdmFsdWUgdG8gbWFrZSBpdCBhdmFpbGFibGUgdG8gdGhlIG5leHQKICAgIC8vIHN0ZXAgb2YgdGhlIGdlbmVyYXRvci4KICAgIG91dHB1dEVsZW1lbnQubGFzdFByb21pc2VWYWx1ZSA9IHZhbHVlOwogICAgcmV0dXJuIG5leHQudmFsdWUucmVzcG9uc2U7CiAgfSk7Cn0KCi8qKgogKiBHZW5lcmF0b3IgZnVuY3Rpb24gd2hpY2ggaXMgY2FsbGVkIGJldHdlZW4gZWFjaCBhc3luYyBzdGVwIG9mIHRoZSB1cGxvYWQKICogcHJvY2Vzcy4KICogQHBhcmFtIHtzdHJpbmd9IGlucHV0SWQgRWxlbWVudCBJRCBvZiB0aGUgaW5wdXQgZmlsZSBwaWNrZXIgZWxlbWVudC4KICogQHBhcmFtIHtzdHJpbmd9IG91dHB1dElkIEVsZW1lbnQgSUQgb2YgdGhlIG91dHB1dCBkaXNwbGF5LgogKiBAcmV0dXJuIHshSXRlcmFibGU8IU9iamVjdD59IEl0ZXJhYmxlIG9mIG5leHQgc3RlcHMuCiAqLwpmdW5jdGlvbiogdXBsb2FkRmlsZXNTdGVwKGlucHV0SWQsIG91dHB1dElkKSB7CiAgY29uc3QgaW5wdXRFbGVtZW50ID0gZG9jdW1lbnQuZ2V0RWxlbWVudEJ5SWQoaW5wdXRJZCk7CiAgaW5wdXRFbGVtZW50LmRpc2FibGVkID0gZmFsc2U7CgogIGNvbnN0IG91dHB1dEVsZW1lbnQgPSBkb2N1bWVudC5nZXRFbGVtZW50QnlJZChvdXRwdXRJZCk7CiAgb3V0cHV0RWxlbWVudC5pbm5lckhUTUwgPSAnJzsKCiAgY29uc3QgcGlja2VkUHJvbWlzZSA9IG5ldyBQcm9taXNlKChyZXNvbHZlKSA9PiB7CiAgICBpbnB1dEVsZW1lbnQuYWRkRXZlbnRMaXN0ZW5lcignY2hhbmdlJywgKGUpID0+IHsKICAgICAgcmVzb2x2ZShlLnRhcmdldC5maWxlcyk7CiAgICB9KTsKICB9KTsKCiAgY29uc3QgY2FuY2VsID0gZG9jdW1lbnQuY3JlYXRlRWxlbWVudCgnYnV0dG9uJyk7CiAgaW5wdXRFbGVtZW50LnBhcmVudEVsZW1lbnQuYXBwZW5kQ2hpbGQoY2FuY2VsKTsKICBjYW5jZWwudGV4dENvbnRlbnQgPSAnQ2FuY2VsIHVwbG9hZCc7CiAgY29uc3QgY2FuY2VsUHJvbWlzZSA9IG5ldyBQcm9taXNlKChyZXNvbHZlKSA9PiB7CiAgICBjYW5jZWwub25jbGljayA9ICgpID0+IHsKICAgICAgcmVzb2x2ZShudWxsKTsKICAgIH07CiAgfSk7CgogIC8vIENhbmNlbCB1cGxvYWQgaWYgdXNlciBoYXNuJ3QgcGlja2VkIGFueXRoaW5nIGluIHRpbWVvdXQuCiAgY29uc3QgdGltZW91dFByb21pc2UgPSBuZXcgUHJvbWlzZSgocmVzb2x2ZSkgPT4gewogICAgc2V0VGltZW91dCgoKSA9PiB7CiAgICAgIHJlc29sdmUobnVsbCk7CiAgICB9LCBGSUxFX0NIQU5HRV9USU1FT1VUX01TKTsKICB9KTsKCiAgLy8gV2FpdCBmb3IgdGhlIHVzZXIgdG8gcGljayB0aGUgZmlsZXMuCiAgY29uc3QgZmlsZXMgPSB5aWVsZCB7CiAgICBwcm9taXNlOiBQcm9taXNlLnJhY2UoW3BpY2tlZFByb21pc2UsIHRpbWVvdXRQcm9taXNlLCBjYW5jZWxQcm9taXNlXSksCiAgICByZXNwb25zZTogewogICAgICBhY3Rpb246ICdzdGFydGluZycsCiAgICB9CiAgfTsKCiAgaWYgKCFmaWxlcykgewogICAgcmV0dXJuIHsKICAgICAgcmVzcG9uc2U6IHsKICAgICAgICBhY3Rpb246ICdjb21wbGV0ZScsCiAgICAgIH0KICAgIH07CiAgfQoKICBjYW5jZWwucmVtb3ZlKCk7CgogIC8vIERpc2FibGUgdGhlIGlucHV0IGVsZW1lbnQgc2luY2UgZnVydGhlciBwaWNrcyBhcmUgbm90IGFsbG93ZWQuCiAgaW5wdXRFbGVtZW50LmRpc2FibGVkID0gdHJ1ZTsKCiAgZm9yIChjb25zdCBmaWxlIG9mIGZpbGVzKSB7CiAgICBjb25zdCBsaSA9IGRvY3VtZW50LmNyZWF0ZUVsZW1lbnQoJ2xpJyk7CiAgICBsaS5hcHBlbmQoc3BhbihmaWxlLm5hbWUsIHtmb250V2VpZ2h0OiAnYm9sZCd9KSk7CiAgICBsaS5hcHBlbmQoc3BhbigKICAgICAgICBgKCR7ZmlsZS50eXBlIHx8ICduL2EnfSkgLSAke2ZpbGUuc2l6ZX0gYnl0ZXMsIGAgKwogICAgICAgIGBsYXN0IG1vZGlmaWVkOiAkewogICAgICAgICAgICBmaWxlLmxhc3RNb2RpZmllZERhdGUgPyBmaWxlLmxhc3RNb2RpZmllZERhdGUudG9Mb2NhbGVEYXRlU3RyaW5nKCkgOgogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAnbi9hJ30gLSBgKSk7CiAgICBjb25zdCBwZXJjZW50ID0gc3BhbignMCUgZG9uZScpOwogICAgbGkuYXBwZW5kQ2hpbGQocGVyY2VudCk7CgogICAgb3V0cHV0RWxlbWVudC5hcHBlbmRDaGlsZChsaSk7CgogICAgY29uc3QgZmlsZURhdGFQcm9taXNlID0gbmV3IFByb21pc2UoKHJlc29sdmUpID0+IHsKICAgICAgY29uc3QgcmVhZGVyID0gbmV3IEZpbGVSZWFkZXIoKTsKICAgICAgcmVhZGVyLm9ubG9hZCA9IChlKSA9PiB7CiAgICAgICAgcmVzb2x2ZShlLnRhcmdldC5yZXN1bHQpOwogICAgICB9OwogICAgICByZWFkZXIucmVhZEFzQXJyYXlCdWZmZXIoZmlsZSk7CiAgICB9KTsKICAgIC8vIFdhaXQgZm9yIHRoZSBkYXRhIHRvIGJlIHJlYWR5LgogICAgbGV0IGZpbGVEYXRhID0geWllbGQgewogICAgICBwcm9taXNlOiBmaWxlRGF0YVByb21pc2UsCiAgICAgIHJlc3BvbnNlOiB7CiAgICAgICAgYWN0aW9uOiAnY29udGludWUnLAogICAgICB9CiAgICB9OwoKICAgIC8vIFVzZSBhIGNodW5rZWQgc2VuZGluZyB0byBhdm9pZCBtZXNzYWdlIHNpemUgbGltaXRzLiBTZWUgYi82MjExNTY2MC4KICAgIGxldCBwb3NpdGlvbiA9IDA7CiAgICB3aGlsZSAocG9zaXRpb24gPCBmaWxlRGF0YS5ieXRlTGVuZ3RoKSB7CiAgICAgIGNvbnN0IGxlbmd0aCA9IE1hdGgubWluKGZpbGVEYXRhLmJ5dGVMZW5ndGggLSBwb3NpdGlvbiwgTUFYX1BBWUxPQURfU0laRSk7CiAgICAgIGNvbnN0IGNodW5rID0gbmV3IFVpbnQ4QXJyYXkoZmlsZURhdGEsIHBvc2l0aW9uLCBsZW5ndGgpOwogICAgICBwb3NpdGlvbiArPSBsZW5ndGg7CgogICAgICBjb25zdCBiYXNlNjQgPSBidG9hKFN0cmluZy5mcm9tQ2hhckNvZGUuYXBwbHkobnVsbCwgY2h1bmspKTsKICAgICAgeWllbGQgewogICAgICAgIHJlc3BvbnNlOiB7CiAgICAgICAgICBhY3Rpb246ICdhcHBlbmQnLAogICAgICAgICAgZmlsZTogZmlsZS5uYW1lLAogICAgICAgICAgZGF0YTogYmFzZTY0LAogICAgICAgIH0sCiAgICAgIH07CiAgICAgIHBlcmNlbnQudGV4dENvbnRlbnQgPQogICAgICAgICAgYCR7TWF0aC5yb3VuZCgocG9zaXRpb24gLyBmaWxlRGF0YS5ieXRlTGVuZ3RoKSAqIDEwMCl9JSBkb25lYDsKICAgIH0KICB9CgogIC8vIEFsbCBkb25lLgogIHlpZWxkIHsKICAgIHJlc3BvbnNlOiB7CiAgICAgIGFjdGlvbjogJ2NvbXBsZXRlJywKICAgIH0KICB9Owp9CgpzY29wZS5nb29nbGUgPSBzY29wZS5nb29nbGUgfHwge307CnNjb3BlLmdvb2dsZS5jb2xhYiA9IHNjb3BlLmdvb2dsZS5jb2xhYiB8fCB7fTsKc2NvcGUuZ29vZ2xlLmNvbGFiLl9maWxlcyA9IHsKICBfdXBsb2FkRmlsZXMsCiAgX3VwbG9hZEZpbGVzQ29udGludWUsCn07Cn0pKHNlbGYpOwo=", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 120} outputId="2df71cb6-0c55-4b9a-ae3b-81309641f691" executionInfo={"status": "ok", "timestamp": 1528928379363, "user_tz": -120, "elapsed": 8603, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
import tensorflow as tf
import numpy as np
tf.reset_default_graph()
# Network Parameters
n_input = 64
n_hidden_1 = 64 # 1st layer num features
n_hidden_2 = 128 # 2nd layer num features
n_hidden_3 = 256 # 3nd layer num features
n_hidden_4 = 256 # 4nd layer num features
n_hidden_5 = 128 # 4nd layer num features
n_hidden_6 = 64 # 1st layer num features
# Parameters
learning_rate = 1e-3
# tf Graph input
x = tf.placeholder("float", [None, n_input], name='x')
y = tf.placeholder("float", [None, n_input], name='y')
lmd = tf.placeholder("float",[], name = 'lambda')
# Store layers weight & bias
c = 0.1
weights = {
'h1': tf.Variable(c*tf.random_normal([n_input, n_hidden_1]), name='W1'),
'h2': tf.Variable(c*tf.random_normal([n_hidden_1, n_hidden_2]), name='W2'),
'h3': tf.Variable(c*tf.random_normal([n_hidden_2, n_hidden_3]), name='W3'),
'h4': tf.Variable(c*tf.random_normal([n_hidden_3, n_hidden_4]), name='W4'),
'h5': tf.Variable(c*tf.random_normal([n_hidden_4, n_hidden_5]), name='W5'),
'h6': tf.Variable(c*tf.random_normal([n_hidden_5, n_hidden_6]), name='W6'),
'out': tf.Variable(c*tf.random_normal([n_hidden_6, n_input]))
}
biases = {
'b1': tf.Variable(c*tf.random_normal([n_hidden_1]), name='b1'),
'b2': tf.Variable(c*tf.random_normal([n_hidden_2]), name='b2'),
'b3': tf.Variable(c*tf.random_normal([n_hidden_3]), name='b3'),
'b4': tf.Variable(c*tf.random_normal([n_hidden_4]), name='b4'),
'b5': tf.Variable(c*tf.random_normal([n_hidden_5]), name='b5'),
'b6': tf.Variable(c*tf.random_normal([n_hidden_6]), name='b6'),
'out': tf.Variable(c*tf.random_normal([n_input]))
}
keep_prob = tf.placeholder(tf.float32)
pre_layer_drop = tf.nn.dropout(x, keep_prob)
layer_1 = tf.nn.relu(tf.add(tf.matmul(pre_layer_drop, weights['h1']), biases['b1']))
layer_2 = tf.nn.relu(tf.add(tf.matmul(layer_1, weights['h2']), biases['b2']))
layer_3 = tf.nn.relu(tf.add(tf.matmul(layer_2, weights['h3']), biases['b3']))
layer_4 = tf.nn.relu(tf.add(tf.matmul(layer_3, weights['h4']), biases['b4']))
layer_5 = tf.nn.relu(tf.add(tf.matmul(layer_4, weights['h5']), biases['b5']))
layer_6 = tf.nn.relu(tf.add(tf.matmul(layer_5, weights['h6']), biases['b6']))
output = tf.add(tf.matmul(layer_6, weights['out']), biases['out'])
loss = tf.reduce_mean(tf.pow(output - y, 2))
loss2 = tf.reduce_mean(tf.abs(layer_3))
floss = loss + lmd * loss2
opt = tf.train.AdamOptimizer(learning_rate).minimize(floss)
#################################### Decoder
x_code = tf.placeholder("float", [None, n_hidden_4], name='x')
dec_layer_1 = tf.nn.relu(tf.add(tf.matmul(x_code, weights['h4']), biases['b4']))
dec_layer_2 = tf.nn.relu(tf.add(tf.matmul(dec_layer_1, weights['h5']), biases['b5']))
dec_layer_3 = tf.nn.relu(tf.add(tf.matmul(dec_layer_2, weights['h6']), biases['b6']))
dec_output = tf.add(tf.matmul(dec_layer_3, weights['out']), biases['out'])
#############################################
init = tf.initialize_all_variables()
# Add ops to save and restore all the variables.
saver = tf.train.Saver()
from sklearn.datasets import load_digits
data = load_digits()
idx = np.random.permutation(data.data.shape[0])
idx_train = idx[:-100]
idx_test = idx[-100:]
import pickle
from google.colab import files
print "Upload file ... digits_idx_part2_sparse.pkl"
uploaded = files.upload()
import io
idx_train,idx_test = pickle.load(io.StringIO(uploaded['digits_idx_part2_sparse.pkl'].decode('utf-8')))
train = data.data[idx_train,:]/16.
test = data.data[idx_test,:]/16.
sess = tf.Session()
sess.run(init)
saver.restore(sess, "autoencoder_digits_part2_sparse.ckpt")
print("Model restored.")
# + id="3-t1EnkIro5j" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 17} outputId="2eb11c4c-f817-459d-b4d3-749090929d8b" executionInfo={"status": "ok", "timestamp": 1528928383661, "user_tz": -120, "elapsed": 805, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
code = sess.run(layer_3, feed_dict={x: test, keep_prob: 1.})
# + id="kgM84jH6ro5n" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 17} outputId="3196637c-5735-4836-962b-3421f395c674" executionInfo={"status": "ok", "timestamp": 1528928385506, "user_tz": -120, "elapsed": 842, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
res = sess.run(dec_output, feed_dict={x_code: code})
# + id="R2Qm4zJ9ro5q" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 4977} outputId="3d34468d-cbd6-4f9e-961d-26c06910b205" executionInfo={"status": "ok", "timestamp": 1528928388927, "user_tz": -120, "elapsed": 2583, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
for i in xrange(10):
plt.figure()
plt.imshow(test[i].reshape((8,8)),cmap='gray',interpolation='nearest')
plt.figure()
plt.imshow(res[i].reshape((8,8)),cmap='gray',interpolation='nearest')
# + [markdown] id="Tw7y9syqro5t" colab_type="text"
# # Manifold learning
# + id="wTpV_Fqcro5u" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 689} outputId="222bd1b5-5883-4d0c-c276-2f576a4bbf09" executionInfo={"status": "ok", "timestamp": 1528928597697, "user_tz": -120, "elapsed": 16815, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
# %matplotlib inline
import tensorflow as tf
import numpy as np
tf.reset_default_graph()
# Network Parameters
n_input = 64
n_hidden_1 = 32 # 1st layer num features
n_hidden_2 = 16 # 2nd layer num features
n_hidden_3 = 2 # 3nd layer num features
n_hidden_4 = 2 # 4nd layer num features
n_hidden_5 = 16 # 4nd layer num features
n_hidden_6 = 32 # 1st layer num features
# Parameters
lr = 1e-3
lr_decay_step = 100000
lr_decay_factor = 0.5
training_epochs = 50000
display_step = 100
batch_size=16
# tf Graph input
x = tf.placeholder("float", [None, n_input], name='x')
y = tf.placeholder("float", [None, n_input], name='y')
learning_rate = tf.placeholder(tf.float32, [], name='learning_rate')
# Store layers weight & bias
c = 0.1
weights = {
'h1': tf.Variable(c*tf.random_normal([n_input, n_hidden_1]), name='W1'),
'h2': tf.Variable(c*tf.random_normal([n_hidden_1, n_hidden_2]), name='W2'),
'h3': tf.Variable(c*tf.random_normal([n_hidden_2, n_hidden_3]), name='W3'),
'h4': tf.Variable(c*tf.random_normal([n_hidden_3, n_hidden_4]), name='W4'),
'h5': tf.Variable(c*tf.random_normal([n_hidden_4, n_hidden_5]), name='W5'),
'h6': tf.Variable(c*tf.random_normal([n_hidden_5, n_hidden_6]), name='W6'),
'out': tf.Variable(c*tf.random_normal([n_hidden_6, n_input]))
}
biases = {
'b1': tf.Variable(c*tf.random_normal([n_hidden_1]), name='b1'),
'b2': tf.Variable(c*tf.random_normal([n_hidden_2]), name='b2'),
'b3': tf.Variable(c*tf.random_normal([n_hidden_3]), name='b3'),
'b4': tf.Variable(c*tf.random_normal([n_hidden_4]), name='b4'),
'b5': tf.Variable(c*tf.random_normal([n_hidden_5]), name='b5'),
'b6': tf.Variable(c*tf.random_normal([n_hidden_6]), name='b6'),
'out': tf.Variable(c*tf.random_normal([n_input]))
}
keep_prob = tf.placeholder(tf.float32)
pre_layer_drop = tf.nn.dropout(x, keep_prob)
layer_1 = tf.nn.relu(tf.add(tf.matmul(pre_layer_drop, weights['h1']), biases['b1']))
layer_2 = tf.nn.relu(tf.add(tf.matmul(layer_1, weights['h2']), biases['b2']))
layer_3 = tf.nn.relu(tf.add(tf.matmul(layer_2, weights['h3']), biases['b3']))
layer_4 = tf.nn.relu(tf.add(tf.matmul(layer_3, weights['h4']), biases['b4']))
layer_5 = tf.nn.relu(tf.add(tf.matmul(layer_4, weights['h5']), biases['b5']))
layer_6 = tf.nn.relu(tf.add(tf.matmul(layer_5, weights['h6']), biases['b6']))
output = tf.add(tf.matmul(layer_6, weights['out']), biases['out'])
loss = tf.reduce_mean(tf.pow(output - y, 2))
opt = tf.train.AdamOptimizer(learning_rate).minimize(loss)
init = tf.initialize_all_variables()
# Add ops to save and restore all the variables.
saver = tf.train.Saver()
from sklearn.datasets import load_digits
data = load_digits()
idx = np.random.permutation(data.data.shape[0])
idx_train = idx[:-100]
idx_test = idx[-100:]
import pickle
from google.colab import files
print "Upload file ... digits_idx_part3_viz.pkl"
uploaded = files.upload()
import io
idx_train,idx_test = pickle.load(io.StringIO(uploaded['digits_idx_part3_viz.pkl'].decode('utf-8')))
train = data.data[idx_train,:]/16.
test = data.data[idx_test,:]/16.
sess = tf.Session()
sess.run(init)
print "Upload file ... autoencoder_digits_part3_viz.ckpt"
uploaded = files.upload()
saver.restore(sess, "autoencoder_digits_part3_viz.ckpt")
print("Model restored.")
import matplotlib.pyplot as plt
xs = test[np.random.randint(0,test.shape[0],1),:]
rec = sess.run(output, feed_dict={x: xs, keep_prob: 1.})
plt.imshow(xs.reshape((8, 8)),interpolation='nearest',cmap='gray')
plt.colorbar()
plt.show()
plt.imshow(rec.reshape((8, 8)),interpolation='nearest',cmap='gray')
plt.colorbar()
plt.show()
# + id="wgWqlFlUro5x" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 17} outputId="25e2f435-9adc-458d-b6ef-713fd2116eeb" executionInfo={"status": "ok", "timestamp": 1528928608340, "user_tz": -120, "elapsed": 665, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
xs = train
representation = sess.run(layer_3, feed_dict={x: xs, keep_prob: 1.})
# + id="KlqU4KN-ro5z" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="d1502828-303d-4c83-ba65-1edd96a88eba" executionInfo={"status": "ok", "timestamp": 1528928610256, "user_tz": -120, "elapsed": 521, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
representation.shape
# + id="jpEtTE-Gro53" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 17} outputId="52fda744-969d-4993-da4b-1aed432c801a" executionInfo={"status": "ok", "timestamp": 1528928633530, "user_tz": -120, "elapsed": 21870, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
from scipy.spatial import distance
N=10
vx = np.linspace(np.min(representation[:,0]),np.max(representation[:,0]),N)
vy = np.linspace(np.min(representation[:,1]),np.max(representation[:,1]),N)
def is_visited(x,l):
for item in l:
if np.abs(x-item)<1e-10:
return True
return False
visited=[]
idx_mat=np.zeros((N,N))
for i in xrange(N):
for j in xrange(N):
d = distance.cdist(np.array([vx[i],vy[j]])[np.newaxis,:], representation)
idx_sort = np.argsort(d)[0]
idx_not_visited=[tmp for tmp in idx_sort if not(is_visited(tmp,visited))]
if len(idx_not_visited)>0:
idx_mat[i,j] = idx_not_visited[0]
visited.append(idx_not_visited[0])
# + id="MIXtktc8ro55" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 593} outputId="341c54db-5dd3-4777-eee1-31631b1100c1" executionInfo={"status": "ok", "timestamp": 1528928720658, "user_tz": -120, "elapsed": 6919, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
# %matplotlib inline
import matplotlib.pyplot as plt
f, axarr = plt.subplots(N, N)
for i in xrange(N):
for j in xrange(N):
axarr[i,j].imshow(xs[int(idx_mat[i,j])].reshape((8,8)),cmap='gray', interpolation='nearest')
f.set_size_inches(10,10)
# + id="2rtK0Kicro58" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
| 6_Unsupervised_final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: newtq37
# language: python
# name: newtq37
# ---
# # Unitary ansatz entering the VQE
#
# The final energy output of a VQE calculation will crucially depend on the ansatz/form of the parameterized unitary $\hat U(\boldsymbol{\theta})$ employed in state preparation. Here we review two popular approaches, the unitary coupled cluster and qubit coupled cluster methodologies, and benchmark them for energy calculations of small molecules.
import numpy as np
import tequila as tq
from utility import *
threshold = 1e-6 #Cutoff for UCC MP2 amplitudes and QCC ranking gradients
# ## Unitary Coupled Cluster (UCC)
#
# The UCC ansatz is obtained by 'unitarizing' the traditional coupled cluster ansatz,
# $$ e^{\hat T} \rightarrow e^{\hat T - \hat T^\dagger} \equiv \hat U_{\text{UCC}}$$
#
#
# Due to non-commutativity of terms in $\hat T - \hat T^\dagger$, the UCC ansatz does not have a straightforward decomposition in terms of circuit primitives implementable on the quantum computer. Therefore, to obtain a form which can be compiled, we employ the Trotter approximation. The accuracy of the circuit ansatz relative to the exact UCC operator will be dependent on how many Trotter steps are employed. The number of Trotter steps is commonly set to its minimal value of one to avoid excessive circuit depth.
trotter_steps = 1
# ### H2 in STO-3G basis
#
# Below is a sample VQE simulation using the UCCSD ansatz compiled using a single trotter step for H$_2$ in minimal basis at $R=2.5$ (Angstrom). For comparison, we can run FCI to obtain the true ground state energy.
#
# +
xyz_data = get_molecular_data('h2', geometry=2.5, xyz_format=True)
basis='sto-3g'
h2 = tq.quantumchemistry.Molecule(geometry=xyz_data, basis_set=basis)
print('Number of spin-orbitals (qubits): {} \n'.format(2*h2.n_orbitals))
E_FCI = h2.compute_energy(method='fci')
print('FCI energy: {}'.format(E_FCI))
# -
# The UCCSD VQE optimization is of the form
# $$E = \min_{\boldsymbol{\theta}} \langle \text{HF} | \hat U_{\text{UCC}}^\dagger(\boldsymbol{\theta}) \hat H \hat U_{\text{UCC}} (\boldsymbol{\theta}) | \text{HF} \rangle $$
# To expedite the optimization process, we can set the initial guess of the amplitudes to zero, i.e. the optimization will begin at the Hartree Fock state. This heuristic is best suited for when Hartree Fock is believed to be in qualitative agreement with the true ground state. To further alleviate quantum resources, we can estimate the amplitudes using classical electronic structure methods (here, MP2 perturbation theory), and only include the unitaries with non-zero estimated amplitudes.
# +
H = h2.make_hamiltonian()
print("\nHamiltonian has {} terms\n".format(len(H)))
U_UCCSD = h2.make_uccsd_ansatz(initial_amplitudes='MP2',threshold=threshold, trotter_steps=trotter_steps)
E = tq.ExpectationValue(H=H, U=U_UCCSD)
print('\nNumber of UCCSD amplitudes: {} \n'.format(len(E.extract_variables())))
print('\nStarting optimization:\n')
result = tq.minimize(objective=E, method="BFGS", initial_values={k:0.0 for k in E.extract_variables()}, tol=1e-6)
print('\nObtained UCCSD energy: {}'.format(result.energy))
# -
# We see that the converged UCCSD energy is in exact agreement with the FCI energy, as expected for a $2$-electron system.
#
# ### H2O in 6-31G basis
#
# Now let us try a larger problem of H$_2$O in 6-31G basis. However, we will restrict the active space. The unrestricted problem leads to a $14$-qubit Hamiltonian, and $34$ UCCSD amplitudes to optimize even after neglecting the zero MP2 amplitudes. Therefore, we will remove some orbital degrees of freedom which are less important in accurately describing the electronic structure. By freezing all orbitals other than $0b_1$, $1b_1$, $2a_1$, and $3a_1$, we reduce the problem to an $8$-qubit Hamiltonian with $8$ UCCSD variational amplitudes.
# +
xyz_data = get_molecular_data('h2o', geometry=1, xyz_format=True)
basis = '6-31g'
active = {'B1':[0,1], 'A1':[2,3]}
h2o = tq.quantumchemistry.Molecule(geometry=xyz_data, basis_set = basis, active_orbitals = active)
print('Number of spin-orbitals (qubits): {} \n'.format(2*h2o.n_orbitals))
E_FCI = h2o.compute_energy(method='fci')
print('FCI energy: {}'.format(E_FCI))
# -
# We will then run the UCCSD VQE simulation (***warning: tq.minimize will take several minutes - 1 hour + to converge for a VQE instance of this size.*** Smaller active spaces can be employed to lower VQE simulation runtimes).
# +
H = h2o.make_hamiltonian()
print("\nHamiltonian has {} terms\n".format(len(H)))
U_UCCSD = h2o.make_uccsd_ansatz(initial_amplitudes='MP2',threshold=threshold, trotter_steps=trotter_steps)
E = tq.ExpectationValue(H=H, U=U_UCCSD)
print('\nNumber of UCCSD amplitudes: {} \n'.format(len(E.extract_variables())))
print('\nStarting optimization:\n')
result = tq.minimize(objective=E, method="BFGS", initial_values={k:0.0 for k in E.extract_variables()}, tol=1e-4)
print('\nObtained UCCSD energy: {}'.format(result.energy))
# -
# We obtain a UCCSD energy within chemical accuracy (< 1.6 miliHartree from FCI energy) for the H$_2$O molecule near equilibrium geometry. As the bond distance approaches the dissociation limit, the energy deviation from FCI is typically expected to increase as electronic correlations increase during covalent bond-breaking. Furthermore, as HF becomes energetically more distant from the FCI wavefunction, the initial guess of all amplitudes being zero may lead to a local minimum. One can then instead initialize the amplitudes using random guesses, and repeat for $n$ samples to attempt to find the global minimum. To facilitate this, one can run the commented lines below. This will execute $n$ instances of tq.minimize for uniformly random initial parameters in interval $[0, 2\pi]$ and return the lowest optimization result.
# +
#n=10
#result = minimize_E_random_guesses(objective=E, method='BFGS', tol=1e-4, n=n)
# -
# ## Qubit Coupled Cluster (QCC)
#
# In contrast to UCC, the QCC methodology makes no direct reference to fermionic algebra and seeks to construct an efficient ansatz directly in qubit-space by finding multi-qubit Pauli strings (entanglers) which lower energy. This is done through an energy-lowering heuristic employing the energy gradient with respect to a Pauli strings variational amplitude. As opposed to UCCSD, the circuit depth and number of parameter is chosen to meet hardware limitations, i.e. one must choose how many exponentiated Pauli strings will be entering the QCC ansatz.
#
# ### H2 in STO-3G basis
#
# Below we perform the entangler screening protocol for H2 in minimal basis, and obtain one grouping of entanglers with non-zero energy gradient. We then select one of them to be used in the QCC VQE simulation.
# +
xyz_data = get_molecular_data('h2', geometry=2.5, xyz_format=True)
basis='sto-3g'
h2 = tq.quantumchemistry.Molecule(geometry=xyz_data, basis_set='sto-3g')
hf_reference = hf_occ(2*h2.n_orbitals, h2.n_electrons)
H = h2.make_hamiltonian()
print("\nHamiltonian has {} terms\n".format(len(H)))
#Define number of entanglers to enter ansatz
n_ents = 1
#Rank entanglers using energy gradient criterion
ranked_entangler_groupings = generate_QCC_gradient_groupings(H.to_openfermion(),
2*h2.n_orbitals,
hf_reference,
cutoff=threshold)
print('Grouping gradient magnitudes (Grouping : Gradient magnitude):')
for i in range(len(ranked_entangler_groupings)):
print('{} : {}'.format(i+1,ranked_entangler_groupings[i][1]))
entanglers = get_QCC_entanglers(ranked_entangler_groupings, n_ents, 2*h2.n_orbitals)
print('\nSelected entanglers:')
for ent in entanglers:
print(ent)
# -
# Once the QCC ranking procedure has been ran, we can simulate the QCC VQE optimization with the generated entanglers. The VQE optimization for the QCC ansatz is of the form
# $$E = \min_{\boldsymbol{\Omega}, \boldsymbol{\tau}} \langle \boldsymbol{\Omega} | U_{\text{ENT}}^\dagger (\boldsymbol{\tau}) \hat H U_{\text{ENT}} (\boldsymbol{\tau}) | \boldsymbol{\Omega} \rangle $$
# where $\boldsymbol{\Omega}$ denote collective Euler angles parameterizing single-qubit rotations, and $\boldsymbol{\tau}$ are entangler amplitudes.
# +
#Mean-field part of U (Omega):
U_MF = construct_QMF_ansatz(n_qubits = 2*h2.n_orbitals)
#Entangling part of U:
U_ENT = construct_QCC_ansatz(entanglers)
U_QCC = U_MF + U_ENT
E = tq.ExpectationValue(H=H, U=U_QCC)
initial_vals = init_qcc_params(hf_reference, E.extract_variables())
#Minimize wrt the entangler amplitude and MF angles:
result = tq.minimize(objective=E, method="BFGS", initial_values=initial_vals, tol=1.e-6)
print('\nObtained QCC energy ({} entanglers): {}'.format(len(entanglers), result.energy))
# -
# We see that the QCC energy converged to the FCI energy with only a single entangler!
#
# ### H2O in 6-31G basis
#
# Let us move on to the problem of H$_2$O in 6-31G basis using the same active space as the UCCSD example.
# +
xyz_data = get_molecular_data('h2o', geometry=1, xyz_format=True)
basis = '6-31g'
active = {'B1':[0,1], 'A1':[2,3]}
h2o = tq.quantumchemistry.Molecule(geometry=xyz_data, basis_set = basis, active_orbitals = active)
hf_reference = hf_occ(2*h2o.n_orbitals, h2o.n_electrons)
H = h2o.make_hamiltonian()
print("\nHamiltonian has {} terms\n".format(len(H)))
#Define number of entanglers to enter ansatz
n_ents = 6
#Rank entanglers using energy gradient criterion
ranked_entangler_groupings = generate_QCC_gradient_groupings(H.to_openfermion(),
2*h2o.n_orbitals,
hf_reference,
cutoff=threshold)
print('Grouping gradient magnitudes (Grouping : Gradient magnitude):')
for i in range(len(ranked_entangler_groupings)):
print('{} : {}'.format(i+1,ranked_entangler_groupings[i][1]))
entanglers = get_QCC_entanglers(ranked_entangler_groupings, n_ents, 2*h2o.n_orbitals)
print('\nSelected entanglers:')
for ent in entanglers:
print(ent)
# -
# With the 6 chosen entanglers, we can then run the QCC VQE simulation.
# +
#Mean-field part of U (Omega):
U_MF = construct_QMF_ansatz(n_qubits = 2*h2o.n_orbitals)
#Entangling part of U:
U_ENT = construct_QCC_ansatz(entanglers)
U_QCC = U_MF + U_ENT
E = tq.ExpectationValue(H=H, U=U_QCC)
initial_vals = init_qcc_params(hf_reference, E.extract_variables())
#Minimize wrt the entangler amplitude and MF angles:
result = tq.minimize(objective=E, method="BFGS", initial_values=initial_vals, tol=1.e-4)
print('\nObtained QCC energy ({} entanglers): {}'.format(len(entanglers), result.energy))
# -
# We obtain chemical accuracy for water near equilibrium geometry with only 6 entanglers. The obtained energy is not as accurate as that of UCCSD for this problem, however the QCC optimization may be performed at a fraction of the UCCSD circuit depth. One can also increase the number of entanglers entering the QCC ansatz to increase accuracy. As a final check, one can always run $n$ VQE trials with random initial guesses to test if the optimization fell into a local minimum. ***(Warning: Completing n=10 trials may take a few minutes for this VQE instance).***
# +
n = 10
result = minimize_E_random_guesses(objective=E, method='BFGS', tol=1e-4, n=n)
print('\nObtained QCC energy ({} entanglers): {}'.format(len(entanglers), result))
| Project_2_VQE_Molecules/S3_Unitary_Ansatz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ===================
# ## Credit Scoring
# Tanggal 28 Agustus 2018
# Analisis Data
# =================
# import librari
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# import kedua
# import library
from random import sample
from xgboost import XGBClassifier
import seaborn as sn
import numpy as np
import pandas
from pandas.plotting import scatter_matrix
import matplotlib.pyplot as plt
from sklearn import model_selection
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
from sklearn.ensemble import VotingClassifier,BaggingClassifier,RandomForestClassifier, AdaBoostClassifier,GradientBoostingClassifier
from sklearn.model_selection import train_test_split, cross_val_score, KFold, GridSearchCV
import seaborn as sns
import numpy as np
import pandas as pd
from sklearn.neighbors import KNeighborsClassifier
from sklearn.naive_bayes import GaussianNB
# display semua
pd.set_option('display.max_rows', None)
pd.set_option('display.max_columns', None)
# import data
train=pd.read_csv('npl_train.csv')
test=pd.read_csv('npl_test.csv')
# menampilakan data semua
train.head(4)
train.columns
# ## Data Ku
# data yang akan saya analisis
train=train[['rasio_pembayaran','persentasi_overlimit', 'rasio_pembayaran_3bulan','rasio_pembayaran_6bulan', 'skor_delikuensi','jumlah_tahun_sejak_pembukaan_kredit', 'total_pemakaian','flag_kredit_macet']]
# summary data
train.describe()
# ukuran data train
train.shape
# Variabel Target
# banyaknya target
train.flag_kredit_macet.value_counts()
# proporsi target
train.flag_kredit_macet.value_counts(normalize=True)
# visualisasi banyak target
plt.figure(figsize=(8,2))
plt.subplot(121)
train.flag_kredit_macet.value_counts().plot.bar(rot=0)
plt.subplot(122)
train.flag_kredit_macet.value_counts(normalize=True).plot.bar(rot=0)
# Missing value
# missing
def missing_data(data):
total = data.isnull().sum().sort_values(ascending = False)
percent = (data.isnull().sum()/data.isnull().count()*100).sort_values(ascending = False)
return pd.concat([total, percent], axis=1, keys=['Total', 'Percent'])
missing_data(train)
train.head()
# ## Analisis Awal
# rasio_pembayaran
# +
# Distribusi data
plt.figure(figsize=(18,5))
plt.subplot(121)
sns.distplot(train['rasio_pembayaran'], kde=True, color = 'grey' )
plt.title('rasio_pembayaran')
# Cek Outlier
plt.subplot(122)
sns.boxplot(x='rasio_pembayaran',data=train)
plt.title('rasio_pembayaran')
# -
# persentasi_overlimit
# +
# Distribusi data
plt.figure(figsize=(18,5))
plt.subplot(121)
sns.distplot(train['persentasi_overlimit'], kde=True, color = 'grey' )
plt.title('persentasi_overlimit')
# Cek Outlier
plt.subplot(122)
sns.boxplot(x='persentasi_overlimit',data=train)
plt.title('persentasi_overlimit')
# -
# rasio_pembayaran_3bulan
# +
# Distribusi data
plt.figure(figsize=(18,5))
plt.subplot(121)
sns.distplot(train['rasio_pembayaran_3bulan'], kde=True, color = 'grey' )
plt.title('rasio_pembayaran_3bulan')
# Cek Outlier
plt.subplot(122)
sns.boxplot(x='rasio_pembayaran_3bulan',data=train)
plt.title('rasio_pembayaran_3bulan')
# -
# rasio_pembayaran_6bulan
# +
# Distribusi data
plt.figure(figsize=(18,5))
plt.subplot(121)
sns.distplot(train['rasio_pembayaran_6bulan'], kde=True, color = 'grey' )
plt.title('rasio_pembayaran_6bulan')
# Cek Outlier
plt.subplot(122)
sns.boxplot(x='rasio_pembayaran_6bulan',data=train)
plt.title('rasio_pembayaran_6bulan')
# -
# skor_delikuensi
train.skor_delikuensi.value_counts()
# +
# Distribusi data
plt.figure(figsize=(18,5))
plt.subplot(121)
sns.distplot(train['skor_delikuensi'], kde=True, color = 'grey' )
plt.title('skor_delikuensi')
# Cek Outlier
plt.subplot(122)
sns.boxplot(x='skor_delikuensi',data=train)
plt.title('skor_delikuensi')
# -
# jumlah_tahun_sejak_pembukaan_kredit
# +
# Distribusi data
plt.figure(figsize=(18,5))
plt.subplot(121)
sns.distplot(train['jumlah_tahun_sejak_pembukaan_kredit'], kde=True, color = 'grey' )
plt.title('jumlah_tahun_sejak_pembukaan_kredit')
# Cek Outlier
plt.subplot(122)
sns.boxplot(x='jumlah_tahun_sejak_pembukaan_kredit',data=train)
plt.title('jumlah_tahun_sejak_pembukaan_kredit')
# -
# total_pemakain == Total pemakaian kartu kredit (tunai dan retail) pada bulan terakhir
# +
# Distribusi data
plt.figure(figsize=(18,5))
plt.subplot(121)
sns.distplot(train['total_pemakaian'], kde=True, color = 'grey' )
plt.title('Total pemakaian')
# Cek Outlier
plt.subplot(122)
sns.boxplot(x='total_pemakaian',data=train)
plt.title('Total Pemakaian')
# -
# ## Target vs Prediktor
# Prediktor
train.head()
train.flag_kredit_macet.value_counts().plot.bar(rot=0)
# rasio_pembayaran
plt.figure(figsize=(18,5))
train.groupby('flag_kredit_macet').rasio_pembayaran.plot.density(title='rasio_pembayaran',legend=True)
# persentasi_overlimit
plt.figure(figsize=(18,5))
train.groupby('flag_kredit_macet').persentasi_overlimit.plot.density(title='persentasi_overlimit',legend=True)
# rasio_pembayaran_3bulan
plt.figure(figsize=(18,5))
train.groupby('flag_kredit_macet').rasio_pembayaran_3bulan.plot.density(title='rasio_pembayaran_3bulan',legend=True)
# rasio_pembayaran_6bulan
plt.figure(figsize=(18,5))
train.groupby('flag_kredit_macet').rasio_pembayaran_6bulan.plot.density(title='rasio_pembayaran_6bulan',legend=True)
plt.figure(figsize=(15,5))
train.groupby('flag_kredit_macet').skor_delikuensi.plot.density(title='skor_delikuensi',legend=True)
plt.figure(figsize=(15,5))
train.groupby('flag_kredit_macet').jumlah_tahun_sejak_pembukaan_kredit.plot.density(title='jumlah_tahun_sejak_pembukaan_kredit',legend=True)
plt.figure(figsize=(15,5))
train.groupby('flag_kredit_macet').total_pemakaian.plot.density(title='total_pemakaian',legend=True)
# skor delikuensi
train.skor_delikuensi.value_counts()
# ## Cek Korelasi
# +
# hitung corelasi
# cor= train.corr(method='pearson')
# print(cor)
# plot the heatmap
plt.figure(figsize=(15,7))
sns.heatmap(cor, xticklabels=cor.columns,yticklabels=cor.columns,annot=True)
# -
# ## X vs X
train.head()
# +
# sns.lmplot(x="sisa_tagihan_tidak_terbayar",y="rasio_pembayaran", data=train)
# plt.figure(figsize=(15,10))
train.plot(kind='scatter',x='rasio_pembayaran',y='rasio_pembayaran_3bulan')
# plt.subplot(222)
# train.plot(kind='scatter',x='rasio_pembayaran',y='rasio_pembayaran_6bulan')
# plt.tight_layout()
# plt.show()
# -
train.plot(kind='scatter',x='rasio_pembayaran',y='rasio_pembayaran_6bulan')
train.plot(kind='scatter',x='rasio_pembayaran',y='jumlah_tahun_sejak_pembukaan_kredit')
train.plot(kind='scatter',x='rasio_pembayaran_3bulan',y='rasio_pembayaran_6bulan')
train.plot(kind='scatter',y='total_pemakaian',x='jumlah_tahun_sejak_pembukaan_kredit')
| Deskripsi data masing2/Data Understanding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # NameX Payments
# We need to load in these libraries into our notebook in order to query, load, manipulate and view the data
# + pycharm={"is_executing": false, "name": "#%%\n"}
import os
from datetime import tzinfo, timedelta, datetime, timezone
import psycopg2
import pandas as pd
import matplotlib
import time
# from datetime import datetime, timedelta
from dateutil.relativedelta import relativedelta
from IPython.core.display import HTML
# %load_ext sql
# %config SqlMagic.displaylimit = 5
# -
# %run ./utility.ipynb
# This will create the connection to the database and prep the jupyter magic for SQL
# +
sbc_pay = 'postgresql://postgres:postgres@docker.for.mac.localhost:6666/pay-db';
# %sql $sbc_pay
# +
namex = 'postgresql://postgres:postgres@docker.<EMAIL>:4444/namex';
# %sql $namex
# -
# Simplest query to run to ensure our libraries are loaded and our DB connection is working
# + pycharm={"is_executing": false, "name": "#%%\n"} tags=[] magic_args="$sbc_pay" language="sql"
# select now() AT TIME ZONE 'PST' as current_date
# + pycharm={"is_executing": false, "name": "#%%\n"} tags=[] magic_args="$namex" language="sql"
# select now() AT TIME ZONE 'PST' as current_date
# + magic_args="$namex rs_today <<" language="sql"
# select (now() AT TIME ZONE 'PST')::date as today
# -
# Set the number of days we want the report to be run over.
# +
inlude_last_number_of_days=2
report_start_date=rs_today[0].today - timedelta(days=inlude_last_number_of_days)
# inlude_last_number_of_hours=24
# report_start_date=rs_today[0].today - timedelta(hours=inlude_last_number_of_hours)
# report_start_date = '2020-12-01'
report_start_date
# -
# ## get all payments
# + pycharm={"is_executing": false, "name": "#%%\n"} tags=[] magic_args="$sbc_pay paid <<" language="sql"
# SELECT i.business_identifier,
# i.id invoice_id,
# i.created_on,
# ir.invoice_number,
# i.invoice_status_code invoice_status,
# p.payment_status_code pay_status,
# i.total,
# i.paid,
# r.receipt_number
# FROM invoices i
# LEFT OUTER JOIN invoice_references ir
# ON ir.invoice_id = i.id
# LEFT OUTER JOIN payments p
# ON p.invoice_number = ir.invoice_number
# LEFT OUTER JOIN receipts r
# ON r.invoice_id = i.id
# WHERE
# created_on >= :report_start_date
# and i.invoice_status_code = 'PAID'
# and i.business_identifier like 'NR%'
# and i.paid <> 101.5
# ORDER BY invoice_id ASC;
# -
paid_frame = paid.DataFrame()
paid_frame['nr_num']=paid_frame['business_identifier']
paid_frame
# ## get all duplicate names
# + magic_args="$namex name_requests <<" language="sql"
# select distinct
# r.id, r.nr_num, r.priority_cd as pri, r.state_cd as nr_state,r.submitted_date,r.source,
# n.name,
# a.first_name||' '||a.last_name as name, a.phone_number, a.email_address
# from requests r, names n, applicants a
# where r.id = n.nr_id
# and r.id = a.nr_id
# and r.submitted_date::date >= :report_start_date
# and
# n.choice=1
# and
# n.name in (
#
# select
# n.name
# from requests r, names n
# where r.id = n.nr_id
# and
# r.submitted_date::date >= :report_start_date
# -- and r.state_cd in ('DRAFT','HOLD','PENDING_PAYMENT','CANCELLED')
# -- and r.state_cd in ('DRAFT','HOLD','PENDING_PAYMENT')
# --and n.choice=1
# group by n.name
# having count(n.name) > 1
# )
# order by n.name
# ;
# -
nr_frame = name_requests.DataFrame()
nr_frame
# ## Merge the Duplicate Names with Payment information
result = pd.merge(nr_frame, paid_frame, how='left', on=['nr_num'])
result=result.drop(['id','business_identifier','created_on','invoice_number','total','receipt_number'], axis=1)
result
# +
time_stamp = time.time()
now = datetime.utcfromtimestamp(time_stamp).replace(tzinfo=timezone.utc)
local_now = now.astimezone(Pacific)
local_now.strftime("%Y.%m.%d.%H")
# -
# ## Write the results to a CSV file
with open('nr_duplicates_with_pay_info.'+local_now.strftime("%Y.%m.%d.%H")+'.csv', 'a') as f:
f.write('\n\n Name Requests\n')
result.to_csv(f, sep=',', encoding='utf-8', index=False)
| support/ops/entities/names-duplicates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
theText = "No food is good food. Ha. I am on a diet and the food is awful and lame. How bad I am!"
positiveWords=['awesome','good','nice','super','fun','delightful']
negativeWords=['awful','lame','horrible','bad']
len(positiveWords)
type(positiveWords)
from nltk.tokenize import sent_tokenize
# +
# This function takes a text and tokenizes it into a list of sentences
def tokenSentences(textIn):
theSentences = sent_tokenize(theText)
return theSentences
# Test how many sentences we get
len(tokenSentences(theText))
# +
import re
def tokenizer(txt2Token): # Function for tokenizing
theTokens = re.findall(r'\b\w[\w-]*\b', txt2Token.lower())
return theTokens
# We test if the function works with the first sentnece from the list above
tokensOfSent = tokenizer(tokenSentences(theText)[0])
print(tokensOfSent)
# +
# Function that counts how many target words are in a list of tokens
def countSentimentalTokens(listOfTokens,listOfTargetWords):
numTargetWords = 0
matchedWords = []
for token in listOfTokens: # Goes through the tokens in the list
if token in listOfTargetWords: # For each one it checks if it is in the target list
numTargetWords += 1
matchedWords.append(token)
return numTargetWords, matchedWords # Note that we are returning a tuple (2 values)
theTuple = countSentimentalTokens(tokensOfSent,positiveWords)
print(str(theTuple[0]) + " " + str(theTuple[1]))
# +
def calculatePercent(listOfTokens,positiveList,negativeList):
numWords = len(listOfTokens) # How many words total
# We call the function to count the tokens from the positive list in the sentence
positiveMatches = countSentimentalTokens(listOfTokens,positiveList)
percntPos = positiveMatches[0] / numWords # We divide by the total number of words for percentage
# We call the function to count the tokens from the negative list in the sentence
negativeMatches = countSentimentalTokens(listOfTokens,negativeList)
percntNeg = negativeMatches[0] / numWords # We divide by the total number of words for percentage
return percntPos, percntNeg # We return the percentage of positive and negative words
# We test the function on the first sentence
results = calculatePercent(tokensOfSent,positiveWords,negativeWords)
print("Positive: " + "{:.0%}".format(results[0]) + " Negative: " + "{:.0%}".format(results[1]))
# -
len(tokensOfSent)
# +
def calculateSentiment(percntPos,percntNeg):
sentiment = percntPos - percntNeg # Subtract the percentage of negative words from positive words
return sentiment
# Test what we get
calculateSentiment(results[0],results[1])
# +
def processText(textIn,posMatchWords,negMatchWords):
listOfSentences = tokenSentences(textIn) # Tokenize the text
listOfSentiments = []
for sentence in listOfSentences: # Process sentence by sentence
sentTokens = tokenizer(sentence) # Tokenize the sentences
percentages = calculatePercent(sentTokens,posMatchWords,negMatchWords) # Calculates percents
theSentiment = calculateSentiment(percentages[0],percentages[1]) # Calculates sentiment
listOfSentiments.append(theSentiment) # Appends sentiment to list
return listOfSentiments # Return the final list
# Test the function
theFinalList = processText(theText,positiveWords,negativeWords)
theFinalList
# -
type(theText)
type(positiveWords)
Text="What a good idea."
processText(Text,positiveWords,negativeWords)
from textblob import TextBlob
text = "Textblob is amazingly simple to use. What great fun!"
testimonial = TextBlob(text) # any string (such as our sonnets)
testimonial.sentiment.polarity
from textblob import TextBlob
blob = TextBlob(theText)
for sentence in blob.sentences:
print(sentence.sentiment.polarity)
| Sentiment exam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: az-ml-realtime-score
# language: python
# name: az-ml-realtime-score
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# # Data Preparation
# In this notebook, we use a subset of [Stack Exchange network](https://archive.org/details/stackexchange) question data
# which includes original questions tagged as 'JavaScript', their duplicate questions and their answers. Here, we
# provide the steps to prepare the data to use in model development for training a model that will match a new
# question with an existing original question.
# +
import os
import pandas as pd
from azure_utils.utilities import read_csv_gz, clean_text, round_sample_strat, random_merge
from notebooks import directory
# -
# Below, we define some parameters that will be used in the data cleaning as well as train and test set preparation.
# The size of the test set
test_size = 0.21
# The minimum length of clean text
min_text = 150
# The minimum number of duplicates per question
min_dupes = 12
# The maximum number of duplicate matches
match = 20
# The output files path
outputs_path = directory + "/data_folder"
# ## Data cleaning
# Next, we download the questions, duplicate questions and answers and load the datasets into pandas dataframes using
# the helper functions.
# URLs to original questions, duplicate questions, and answers.
data_url = "https://bostondata.blob.core.windows.net/stackoverflow/{}"
questions_url = data_url.format("orig-q.tsv.gz")
dupes_url = data_url.format("dup-q.tsv.gz")
answers_url = data_url.format("ans.tsv.gz")
# Load datasets.
questions = read_csv_gz(questions_url, names=('Id', 'AnswerId', 'Text0', 'CreationDate'))
dupes = read_csv_gz(dupes_url, names=('Id', 'AnswerId', 'Text0', 'CreationDate'))
answers = read_csv_gz(answers_url, names=('Id', 'Text0'))
# Let's now check the dataframes. Notice that questions and duplicates have "AnswerID" column that would help match
# ith the index of answers dataframe.
questions.head()
dupes.head()
answers.head()
# Let's check the first original question's text.
print(questions.iloc[0, 1])
# Let's now check the duplicates for that question.
print(dupes[dupes.AnswerId == questions.iloc[0, 0]])
# Below is the answer to the original question.
print(answers.at[questions.iloc[0, 0], 'Text0'])
# Next, we use the helper functions to clean questions, duplicates and answers from unwanted text such as code, html
# tags and links. Notice that we add a new column 'Text' to each dataframe for clean text in lowercase.
# Clean up all text, and keep only data with some clean text.
for df in (questions, dupes, answers):
df["Text"] = df.Text0.apply(clean_text).str.lower()
questions = questions[questions.Text.str.len() > 0]
answers = answers[answers.Text.str.len() > 0]
dupes = dupes[dupes.Text.str.len() > 0]
# Let's compare the first original question and cleaned version as an example.
# Original question.
print(questions.iloc[0, 1])
# After cleaning.
print(questions.iloc[0, 3])
# It turns out that some duplicate questions were also in original questions. Also, some original questions and some
# duplicate questions were duplicated in the datasets. In the following, we remove them from the dataframes.
# First, remove dupes that are questions, then remove duplicated questions and dupes.
dupes = dupes[~dupes.index.isin(questions.index)]
questions = questions[~questions.index.duplicated(keep='first')]
dupes = dupes[~dupes.index.duplicated(keep='first')]
# We also make sure we keep questions with answers and duplicates.
# Keep only questions with answers and dupes, answers to questions, and dupes of questions.
questions = questions[
questions.AnswerId.isin(answers.index) & questions.AnswerId.isin(dupes.AnswerId)
]
answers = answers[answers.index.isin(questions.AnswerId)]
dupes = dupes[dupes.AnswerId.isin(questions.AnswerId)]
# Verify data integrity.
assert questions.AnswerId.isin(answers.index).all()
assert answers.index.isin(questions.AnswerId).all()
assert questions.AnswerId.isin(dupes.AnswerId).all()
assert dupes.AnswerId.isin(questions.AnswerId).all()
# Below are some statistics on the data. Notice that some questions have very low number of duplicates while others may
# have a large number.
# Report on the data.
print("Text statistics:")
print(
pd.DataFrame(
[
questions.Text.str.len().describe().rename("questions"),
answers.Text.str.len().describe().rename("answers"),
dupes.Text.str.len().describe().rename("dupes"),
]
)
)
print("\nDuplication statistics:")
print(pd.DataFrame([dupes.AnswerId.value_counts().describe().rename("duplications")]))
print(
"\nLargest class: {:.2%}".format(
dupes.AnswerId.value_counts().max() / dupes.shape[0]
)
)
# Now, we reset all indexes to use them as columns in the rest of the steps.
# Reset each dataframe's index.
questions.reset_index(inplace=True)
answers.reset_index(inplace=True)
dupes.reset_index(inplace=True)
# We filter the questions and duplicates to have at least min_text number of characters.
# Apply the minimum text length to questions and dupes.
questions = questions[questions.Text.str.len() >= min_text]
dupes = dupes[dupes.Text.str.len() >= min_text]
# Keep only questions with dupes, and dupes of questions.
label_column = "AnswerId"
questions = questions[questions[label_column].isin(dupes[label_column])]
dupes = dupes[dupes[label_column].isin(questions[label_column])]
# Here, we remove questions and their duplicates that are less than min_dupes parameter.
# Restrict the questions to those with a minimum number of dupes.
answerid_count = dupes.groupby(label_column)[label_column].count()
answerid_min = answerid_count.index[answerid_count >= min_dupes]
questions = questions[questions[label_column].isin(answerid_min)]
dupes = dupes[dupes[label_column].isin(answerid_min)]
# Verify data integrity.
assert questions[label_column].isin(dupes[label_column]).all()
assert dupes[label_column].isin(questions[label_column]).all()
# Here are some statistics on the resulting dataset.
# Report on the data.
print("Restrictions: min_text={}, min_dupes={}".format(min_text, min_dupes))
print("Restricted text statistics:")
print(
pd.DataFrame(
[
questions.Text.str.len().describe().rename("questions"),
dupes.Text.str.len().describe().rename("dupes"),
]
)
)
print("\nRestricted duplication statistics:")
print(
pd.DataFrame([dupes[label_column].value_counts().describe().rename("duplications")])
)
print(
"\nRestricted largest class: {:.2%}".format(
dupes[label_column].value_counts().max() / dupes.shape[0]
)
)
# ## Prepare train and test sets
# In this part, we prepare train and test sets. For training a binary classification model, we will need to construct
# match and non-match pairs from duplicates and their questions. Finding matching pairs can be accomplished by joining
# each duplicate with its question. However, non-match examples need to be constructed randomly.
# As a first step, to make sure we train and test the performance of the model on each question, we will need to have
# examples of match and non-match pairs for each question both in train and test sets. In order to achieve that,
# we split the duplicates in a stratified manner into train and test sets making sure at least 1 or more duplicates per
# question is in the test set depending on test_size parameter and number of duplicates per each question.
# Split dupes into train and test ensuring at least one of each label class is in test.
dupes_test = round_sample_strat(dupes, dupes[label_column], frac=test_size)
dupes_train = dupes[~dupes.Id.isin(dupes_test.Id)]
assert dupes_test[label_column].unique().shape[0] == dupes[label_column].unique().shape[0]
# The relevant columns for text pairs data.
balanced_pairs_columns = ['Id_x', 'AnswerId_x', 'Text_x', 'Id_y', 'Text_y', 'AnswerId_y', 'Label', 'n']
# Next, we pair each training duplicate in train set with its matching question and N-1 random questions using the
# helper function.
# Use AnswerId to pair each training dupe with its matching question and also with N-1 questions not its match.
balanced_pairs_train = random_merge(dupes_train, questions, N=match)
# Labeling is done such that matching pairs are labeled as 1 and non-match pairs are labeled as 0.
# Label records by matching AnswerIds.
balanced_pairs_train["Label"] = (
balanced_pairs_train.AnswerId_x == balanced_pairs_train.AnswerId_y
).astype(int)
# Keep only the relevant data.
balanced_pairs_train = balanced_pairs_train[balanced_pairs_columns]
balanced_pairs_train.head()
# Sort the data by dupe ID and Label.
balanced_pairs_train.sort_values(by=['Id_x', 'Label'], ascending=[True, False], inplace=True)
# In testing set, we match each duplicate with all the original questions and label them same way as training set.
# Use AnswerId to pair each testing dupe with all questions.
balanced_pairs_test = random_merge(dupes_test, questions, N=questions.shape[0])
# Label records by matching AnswerIds.
balanced_pairs_test["Label"] = (
balanced_pairs_test.AnswerId_x == balanced_pairs_test.AnswerId_y
).astype(int)
# Keep only the relevant data.
balanced_pairs_test = balanced_pairs_test[balanced_pairs_columns]
balanced_pairs_test.head()
# Sort the data by dupe ID and Label.
balanced_pairs_test.sort_values(
by=["Id_x", "Label"], ascending=[True, False], inplace=True
)
# Finally, we report the final train and test sets and save as text files to be used by modeling.
# Report on the datasets.
print(
"balanced_pairs_train: {:,} rows with {:.2%} matches".format(
balanced_pairs_train.shape[0], balanced_pairs_train.Label.mean()
)
)
print(
"balanced_pairs_test: {:,} rows with {:.2%} matches".format(
balanced_pairs_test.shape[0], balanced_pairs_test.Label.mean()
)
)
# +
os.makedirs(outputs_path, exist_ok=True)
# Save the data.
balanced_pairs_train_path = os.path.join(outputs_path, "balanced_pairs_train.tsv")
print(
"Writing {:,} to {}".format(
balanced_pairs_train.shape[0], balanced_pairs_train_path
)
)
balanced_pairs_train.to_csv(
balanced_pairs_train_path, sep="\t", header=True, index=False
)
balanced_pairs_test_path = os.path.join(outputs_path, "balanced_pairs_test.tsv")
print(
"Writing {:,} to {}".format(balanced_pairs_test.shape[0], balanced_pairs_test_path)
)
balanced_pairs_test.to_csv(balanced_pairs_test_path, sep="\t", header=True, index=False)
# Save original questions to be used for scoring later.
questions_path = os.path.join(outputs_path, "questions.tsv")
print("Writing {:,} to {}".format(questions.shape[0], questions_path))
questions.to_csv(questions_path, sep="\t", header=True, index=False)
# Save the test duplicate questions to be used with the scoring function.
dupes_test_path = os.path.join(outputs_path, "dupes_test.tsv")
print("Writing {:,} to {}".format(dupes_test.shape[0], dupes_test_path))
dupes_test.to_csv(dupes_test_path, sep="\t", header=True, index=False)
# -
# We can now move on to [train on local](02_TrainOnLocal.ipynb) notebook to train our model using Azure Machine
# Learning.
| notebooks/01_DataPrep.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
## Importing the necesssary Libraries
# +
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
import psycopg2 as pg2
import datetime as dt
# package used for converting the data into datetime format
from sklearn.preprocessing import LabelEncoder
from sklearn.feature_selection import RFE, f_regression
from sklearn.linear_model import (LinearRegression, Ridge, Lasso, RandomizedLasso)
from sklearn.preprocessing import MinMaxScaler
from sklearn.ensemble import RandomForestRegressor
import warnings
warnings.filterwarnings("ignore")
# +
##Establish connection to the postgres database
# -
conn= pg2.connect('dbname = Amazon user= postgres password = <PASSWORD> host= 127.0.0.1')
cur=conn.cursor()
df_raw = pd.read_sql_query('select * from public."keepa"', conn)
#Check the dimension of the raw data to see if its properly imported
print('Starting size of our Dataset ')
df_raw.shape
# Print out count of each datatype in the dataframe
df_raw.dtypes.value_counts()
# ### Price Aggregator
# Price column is divided into three different coumns depending on availability and condition of books carried by Amazon.
# According to keepa, prise is set based on availability of information in the following order. The aggregator function adds
# a new column called 'price' to the dataset and assigns the value that appers first from the following list and finally
# deletes the three price columns.
# * amazon_Price
# * marketplace_new
# * marketplace_used_price
# +
def PriceAggregator(original_df):
df=original_df
# create a copy of the three columns to choose amazon price from
df_copy=df[['amazon_price','marketplace_new_price','marketplace_used_price']]
# Replace missing price denoted by -1 to Null in all three price columns
for item in df_copy:
df_copy[item].replace('-1',np.nan, inplace=True)
# Add a new column to store the aggregated price with default value of 'amazon_price'
df.insert(79,'price',df_copy['amazon_price'].astype('float'))
#Loop throgh all three columns to assign non-null value to the newly created price column.
#Keep amazon_price as is if not null, otherwise assign marketplace_new_price as the new price.
#Where both 'amazon_price' and 'marketplace_new_price' are null, price will be set to
#'marketplace_used_price' regardless of its value.
for i in range(df['price'].size):
if pd.isnull(df['price'][i]):
if pd.isnull(df_copy['marketplace_new_price'][i]):
if pd.isnull(df_copy['marketplace_used_price'][i]):
pass
else:
df['price'][i]=df_copy['marketplace_used_price'][i]
else:
df['price'][i]=df_copy['marketplace_new_price'][i]
else:
pass
#Delete records where price record is missing since there is no value to cross check
#the accuracy of the model in the test set.
df.dropna(subset=['price'], axis=0, inplace=True)
#Reset index after dropping rows with missing price
df.reset_index(drop= True, inplace=True)
#Delete old price columns after assigning aggregated price to a brand new column
df.drop(['amazon_price','marketplace_new_price','marketplace_used_price'], axis=1 , inplace=True)
#Return the a dataframe with a new price column added to the original dataframe
return df
# -
df=PriceAggregator(df_raw)
df.shape
# ## Delete duplicate records if there are any
df.shape
df.drop_duplicates(inplace = True)
df.shape
# +
## categorical valued features
# -
cat=['author', 'binding','brand','categoryTree_0','categoryTree_1','categoryTree_2','categoryTree_3','categoryTree_4',
'color','edition','features','format','genre','label','languages_0', 'manufacturer','productGroup','publisher','studio',
'title','type']
df[cat].head(5)
# ## Replace every missing value with Null Values for further analysis.
df.replace('', np.NaN, inplace=True)
#df.fillna(np.NaN)
df.isna().sum().sort_values(ascending=False).to_frame(name='Count of Null Values')
# We can delete those columns that contain Null values for the majority of the record set since those features are not common
# behaviour to our instances.
Null_features=['coupon','offers','liveOffersOrder','promotions','buyBoxSellerIdHistory','features','upcList','variations',
'hazardousMaterialType','genre','platform','variationCSV','parentAsin','department','size','model','color'
,'partNumber','mpn','brand','edition','format']
df[Null_features].isna().sum()
#
# We can delete these features without losing any useful information from our data since more than 50% of the records in the above list contain null values.
#
df.drop(Null_features, axis=1, inplace=True)
df.shape
# For the remaining null values in our data where the total count is relatively small, we will replace them by a statistically representative values like mean or mode.
#
# * Mode for categorical values where there is a clear majority or filled with 'Unknown'
# * Mean value is used for numerical columns
with_Nulls=df.loc[:, df.isna().sum()!=0].columns.tolist()
df[with_Nulls].isna().sum().sort_values(ascending=False)
# +
## For our records sets mainly comprised of string or categorival data
# -
Nulls2Unknown=['categoryTree_4','categoryTree_3','categoryTree_2','author','studio','publisher','manufacturer',
'label']
df[with_Nulls].head(3)
for item in with_Nulls:
print (f'{item}\t\t{df[item].value_counts().max()}')
# Given that our data contains 100,000 records we can clearly see the high mode value for some of the features to replace the nu
Nulls2Mode=['languages_0','categoryTree_0','categoryTree_1']
mode = df.filter(['languages_0','categoryTree_0','categoryTree_1']).mode()
df[Nulls2Mode]=df[Nulls2Mode].fillna(df.mode().iloc[0])
#
#
#
#
#
# For the following features since there is no one single category with a high frequency(Mode) in the group, we are filling the missing(Null) values with 'Unknown'.
#
NullswithNoMode=df.loc[:, df.isna().sum()!=0].columns.tolist()
for item in NullswithNoMode:
print(item)
print(df[item].value_counts().nlargest(3))
df[NullswithNoMode]=df[NullswithNoMode].fillna('Unknown')
# Check if there are still missing or null values in the dataset
df[df.loc[:, df.isna().sum()!=0].columns].isna().sum()
# We have entirely replaced the null and missing values in the dataset by statistically representative values.
# ## Data Type Conversion
df.dtypes.value_counts()
# Lets group all those features that are in string (object) format and convert them to numeric
# +
#df[strings] = df[strings].apply(pd.to_numeric, errors='coerce', axis=1)
# -
df.dtypes.value_counts()
#Convert columns that contain numerical values to numeric data type using pandsas to_numeric
numeric=['availabilityAmazon',
'ean','hasReviews', 'isEligibleForSuperSaverShipping', 'isEligibleForTradeIn',
'isRedirectASIN', 'isSNS', 'lastPriceChange','lastRatingUpdate', 'lastUpdate', 'listedSince',
'newPriceIsMAP', 'numberOfItems','numberOfPages', 'offersSuccessful', 'packageHeight',
'packageLength', 'packageQuantity', 'packageWeight', 'packageWidth',
'publicationDate', 'releaseDate', 'rootCategory','stats_atIntervalStart', 'stats_avg', 'stats_avg30', 'stats_avg90',
'stats_avg180', 'stats_current', 'stats_outOfStockPercentage30',
'stats_outOfStockPercentage90', 'stats_outOfStockPercentageInInterval',
'trackingSince','sales_rank', 'price']
#cols = ['productType','rootCategory','stats_atIntervalStart','availabilityAmazon','hasReviews','isRedirectASIN','isSNS','isEligibleForTradeIn','isEligibleForSuperSaverShipping', 'ean','hasReviews', 'availabilityAmazon','isEligibleForTradeIn','lastPriceChange','lastRatingUpdate','lastUpdate','lastRatingUpdate','lastUpdate','listedSince',"newPriceIsMAP", "numberOfItems", "numberOfPages","packageHeight", "packageLength","packageQuantity", "packageWeight", "packageWidth",'stats_avg', 'stats_avg30', 'stats_avg90', 'stats_avg180', 'stats_current',"stats_outOfStockPercentage30", "stats_outOfStockPercentage90","stats_outOfStockPercentageInInterval","trackingSince",'upc','price','amazon_price', 'marketplace_new_price', 'marketplace_used_price', 'sales_rank']
df[numeric] = df[numeric].apply(pd.to_numeric, errors='coerce', axis=1)
df.dtypes.value_counts()
strings=df.loc[:, df.dtypes == np.object].columns.tolist()
print('\n'+ 'Sample of the dataset with only categorical information'+'\n')
df[strings].head(3)
# We can delete asin, ean and imageCSV columns since the information contained in them is not characteristic discription of books.
df.drop(['asin','imagesCSV','ean', 'upc'], axis=1, inplace=True)
#upc might break code watch for it
df.shape
df.dtypes.value_counts()
df.loc[:, df.dtypes == np.object].columns
# The language_0 column contains aggregated information separated by comma, we are going to split it into 2 parts;
df['languages_0'].head(5)
new = df['languages_0'].str.split(",", n = 1, expand = True)
df['language_1']=new[0]
df['language_2']=new[1]
# +
# reduced categories froom 9 to 6 grouping related categories together
#df['language_1'].value_counts().to_frame()
# +
#group English, english and Middle English to one categry
df['language_1'].replace(('English', 'english','Middle English'),'English', inplace = True)
#grouping Spanish,Portuguese and Latin under "Spanish"
df['language_1'].replace(('Spanish', 'Portuguese','Latin'),'Spanish', inplace = True)
#grouping Chinese, mandarin Chinese and simplified chinese to Chinese
df['language_1'].replace(('Simplified Chinese', 'Mandarin Chinese','Chinese'),'Chinese', inplace = True)
#grouping Arabic,Hebrew and Turkish under Middle Eastern
df['language_1'].replace(('Arabic', 'Hebrew','Turkish'),'Middle Eastern', inplace = True)
# group languages with single entry record in to one group called 'Others'
df['language_1'].replace(('Hindi', 'Scots','Filipino','Malay','Dutch','Greek','Korean','Romanian','Czech'),'Others', inplace = True)
#grouping Danish and Norwegian into one group of 'Scandinavian'
df['language_1'].replace(('Danish', 'Norwegian'),'Scandinavian', inplace=True)
# -
#replaced ('published','Published,Dolby Digital 1.0','Published,DTS-HD 5.1') by Published
df['language_2'].replace(('published','Published,Dolby Digital 1.0','Published,DTS-HD 5.1'),'Published', inplace=True)
df[['language_1','language_2']].head(5)
#Since we have copied the information into new columns we can delete the languages_0 column
df.drop(['languages_0'], axis=1 , inplace=True)
df.columns
df.shape
df.binding.value_counts()
# The binding column contains 73 differnt categories that are mostly related and some of them contain very small elements, we will aggregate closely related categories to reduce the dimension of our variables to avoid curse of dimentioanlity
#
df.binding.nunique()
dict={'Unknown':['Printed Access Code', 'Unknown','Health and Beauty', 'Lawn & Patio', 'Workbook', 'Kitchen', 'Automotive', 'Jewelry'],
'spiral':[ 'Spiral-bound', 'Staple Bound', 'Ring-bound', 'Plastic Comb', 'Loose Leaf', 'Thread Bound'],
'magazines':[ 'Journal', 'Single Issue Magazine', 'Print Magazine'],
'audios':[ 'Audible Audiobook', 'Audio CD', 'DVD', 'Album', 'MP3 CD', 'Audio CD Library Binding'],
'digital_prints':[ 'CD-ROM', 'Blu-ray', 'DVD-ROM', 'Kindle Edition', 'Video Game', 'Sheet music', 'Software Download',
'Personal Computers', 'Electronics', 'Game', 'Wireless Phone Accessory'],
'hardcovers':['Hardcover', 'Hardcover-spiral', 'Turtleback', 'Roughcut'],
'others':[ 'Cards', 'Pamphlet', 'Calendar', 'Map', 'Stationery', 'Accessory', 'Misc. Supplies', 'Office Product', 'Poster',
'Wall Chart', 'Bookmark', 'JP Oversized'],
'paperbacks':[ 'Paperback', 'Perfect Paperback', 'Mass Market Paperback', 'Flexibound', 'Print on Demand (Paperback)',
'Comic', 'Puzzle', 'Paperback Bunko'],
'leather_bonded':[ 'Bonded Leather', 'Leather Bound', 'Imitation Leather', 'Vinyl Bound'],
'board_book':[ 'Board book', 'Baby Product', 'Toy', 'Rag Book', 'Card Book', 'Bath Book', 'Pocket Book'],
'schoolLibrary_binding':[ 'School & Library Binding', 'Library Binding', 'Textbook Binding']}
for key,val in dict.items():
df.binding.replace(val,key, inplace=True)
df.binding.value_counts()
df.head()
#catTree_under10.categoryTree_2.values= 'Other'
def groupUnder10(x):
cond = df[x].value_counts()
threshold = 10
df[x] = np.where(df[x].isin(cond.index[cond > threshold ]), df[x], 'Others')
return('All the different categories that contain less than 10 items in the %s column are renamed to Others \n inorder to avoid curse of dimensionality' %x)
df[['categoryTree_1','categoryTree_2','categoryTree_3','categoryTree_4']].nunique()
groupUnder10('categoryTree_2')
#group under 10 counts in to one for categoryTree_3 column
groupUnder10('categoryTree_3')
groupUnder10('categoryTree_4')
df[['categoryTree_0','categoryTree_1','categoryTree_2','categoryTree_3','categoryTree_4']].nunique()
# +
## Some features are duplicated within the dataset, lets delete those duplicated columns
# +
## Delete duplicated features
duplicates=df[['label', 'manufacturer', 'publisher', 'studio']]
# -
df['label'].equals(df['manufacturer'])
df['label'].equals(duplicates['publisher'])
df['label'].equals(duplicates['studio'])
# +
#df[df.duplicated(['label', 'manufacturer', 'publisher', 'studio'])]
# -
duplicates.describe(include='all')
df.duplicated(subset=['label', 'manufacturer', 'publisher', 'studio'],keep='first').value_counts()
# Since the above 4 columns contain duplicated informartion in 89493 out of 99600 total records we can keep one of those and drop the reamining ones without losing useful information.
# Keep publisher and drop the rest
df.drop(['label', 'manufacturer','studio'], axis =1, inplace=True)
df.shape
df.describe(include='all').transpose()
# ## Encoding categorical columns
cat_cols=['author','language_1','language_2','binding','categoryTree_0', 'categoryTree_1', 'categoryTree_2', 'categoryTree_3',
'categoryTree_4','productGroup','publisher','title','type','language_1','language_2']
df[cat_cols].head()
# +
#might not be necessary
df['author']=df['author'].astype(str)
df['language_2']=df['language_2'].astype(str)
df['categoryTree_1']=df['categoryTree_1'].astype(str)
df['categoryTree_2']=df['categoryTree_2'].astype(str)
df['categoryTree_3']=df['categoryTree_3'].astype(str)
df['categoryTree_4']=df['categoryTree_4'].astype(str)
# -
#
# ## Outlier detection and transformation
# Before we decide whether to use standard deviation or interquntile range to identify outliers, lets plot the data points using a distribution plot.
def distWithBox(data):
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
sns.set(style="ticks")
x = df[data]
f, (ax_box, ax_hist) = plt.subplots(2, sharex=True,
gridspec_kw={"height_ratios": (.15, .85)})
sns.boxplot(x, ax=ax_box)
sns.distplot(x, ax=ax_hist)
ax_box.set(yticks=[])
sns.despine(ax=ax_hist)
sns.despine(ax=ax_box, left=True)
## Distribution and box plot of the raw data with outliers
distWithBox('price')
# We can see from the graph that the distribution is not normal so we have to use interquantile range to cutoff outliers
from numpy import percentile
data=df['price']
q25, q75 = percentile(data, 25), percentile(data, 75)
iqr = q75 - q25
print('Percentiles: 25th=%.3f, 75th=%.3f, IQR=%.3f' % (q25, q75, iqr))
# calculate the outlier cutoff
cut_off = iqr * 1.5
lower, upper = q25 - cut_off, q75 + cut_off
# identify outliers
outliers = [x for x in data if x < lower or x > upper]
print('Identified outliers: %d' % len(outliers))
outliers_removed = [x for x in data if x >= lower and x <= upper]
print('Non-outlier observations: %d' % len(outliers_removed))
outliers=[]
data_1=df['price']
for item in data_1:
if item <lower or item>upper:
outliers.append(item)
x=df['price']
outlier_indices=list(data_1.index[(x<lower) | (x> upper)])
len(outlier_indices)
df.drop(axis=0,index=outlier_indices, inplace=True)
df.shape
## lets plot distribution with and box plot to see the change after we trim down the outliers
distWithBox('price')
# ### Label Encoding
df[cat_cols]= df[cat_cols].apply(LabelEncoder().fit_transform)
# ### Feature Selection
#
# VarianceThreshold is a simple baseline approach to feature selection. It removes all features whose variance doesn’t meet some threshold. By default, it removes all zero-variance features, i.e. features that have the same value in all samples.
#
# so we can select using the threshold .8 * (1 - .8):
df_X=df.loc[:, df.columns != 'price']
df_y=df['price']
# +
from sklearn.feature_selection import VarianceThreshold
print('%s Number of features before VarianceThreshhold'%len(df_X.columns))
selector=VarianceThreshold(threshold=(.8*(1-.8)))
FeaturesTransformed=selector.fit_transform(df_X)
## print the support and shape of the transformed features
print(selector.get_support())
# -
data=df_X[df_X.columns[selector.get_support(indices=True)]]
cols=data.columns
df_reduced=pd.DataFrame(FeaturesTransformed, columns=cols)
df_reduced.shape
df_X=df_reduced
# +
from sklearn.ensemble import RandomForestRegressor
from yellowbrick.features.importances import FeatureImportances
fig = plt.figure(figsize=(20,20))
ax = fig.add_subplot()
viz = FeatureImportances(RandomForestRegressor(), ax=ax)
viz.fit(df_X, df_y)
viz.poof()
# -
feature_importances = pd.DataFrame(viz.feature_importances_,
index=df_X.columns,
columns=['importance']).sort_values('importance', ascending=False)
## important features for Random Forest Regression
importants=feature_importances.index[feature_importances.importance!=0]
importants
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import train_test_split as split
X=df[['language_1', 'sales_rank', 'type', 'trackingSince', 'title',
'stats_outOfStockPercentageInInterval', 'stats_outOfStockPercentage90',
'stats_outOfStockPercentage30', 'stats_current', 'stats_avg180',
'stats_avg90', 'stats_avg30', 'stats_avg', 'stats_atIntervalStart',
'rootCategory', 'releaseDate', 'publisher', 'publicationDate',
'productGroup', 'packageWidth', 'packageWeight', 'packageQuantity',
'packageLength', 'packageHeight', 'numberOfPages', 'numberOfItems',
'listedSince', 'lastUpdate', 'lastRatingUpdate', 'lastPriceChange',
'isEligibleForSuperSaverShipping', 'categoryTree_4', 'categoryTree_3',
'categoryTree_2', 'binding', 'author', 'productType']]
Y=df['price']
model=RandomForestRegressor()
X_train, X_test, Y_train, Y_test= split(X,Y,test_size=0.25, random_state=42)
model.fit(X_train,Y_train)
# +
from sklearn.ensemble import GradientBoostingRegressor
from yellowbrick.features.importances import FeatureImportances
fig = plt.figure(figsize=(20,20))
ax = fig.add_subplot()
viz = FeatureImportances(GradientBoostingRegressor(), ax=ax)
viz.fit(df_X, df_y)
viz.poof()
# -
feature_importances = pd.DataFrame(viz.feature_importances_,
index=df_X.columns,
columns=['importance']).sort_values('importance', ascending=False)
## important features for gradient boosting regression
importants=feature_importances.index[feature_importances.importance!=0]
importants
# +
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.model_selection import train_test_split as split
X=df[['language_1', 'sales_rank', 'type', 'trackingSince', 'title',
'stats_outOfStockPercentageInInterval', 'stats_outOfStockPercentage90',
'stats_outOfStockPercentage30', 'stats_current', 'stats_avg180',
'stats_avg90', 'stats_avg30', 'stats_avg', 'stats_atIntervalStart',
'rootCategory', 'releaseDate', 'publisher', 'publicationDate',
'productGroup', 'packageWidth', 'packageWeight', 'packageQuantity',
'packageLength', 'packageHeight', 'numberOfPages', 'numberOfItems',
'listedSince', 'lastUpdate', 'lastRatingUpdate', 'lastPriceChange',
'isEligibleForSuperSaverShipping', 'categoryTree_4']]
Y=df['price']
model=GradientBoostingRegressor()
X_train, X_test, Y_train, Y_test= split(X,Y,test_size=0.25, random_state=42)
model.fit(X_train,Y_train)
# -
# +
# %matplotlib inline
from yellowbrick.classifier import ClassificationReport
from yellowbrick.classifier import ClassPredictionError
from yellowbrick.regressor import ResidualsPlot
from sklearn.svm import SVR
from sklearn.neural_network import MLPRegressor
from sklearn.neighbors import KNeighborsRegressor
from sklearn.linear_model import BayesianRidge, LinearRegression
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.tree import DecisionTreeClassifier
from sklearn.linear_model import Lasso, LinearRegression
regressors = {
"support vector machine": SVR(),
"Gradient Boost": GradientBoostingRegressor(),
"Random Forest": RandomForestRegressor(),
"lasso linear model": Lasso(),
"linear regression": LinearRegression()
}
for _, regressor in regressors.items():
visualizer = ResidualsPlot(regressor)
visualizer.fit(X_train, Y_train)
visualizer.score(X_test, Y_test)
visualizer.poof()
# +
from sklearn.model_selection import StratifiedKFold
from sklearn.naive_bayes import MultinomialNB
from yellowbrick.model_selection import CVScores
# Create a cross-validation strategy
cv = StratifiedKFold(n_splits=12, random_state=42)
x_val = X.values
y_val = Y.values
# Instantiate the classification model and visualizer
model = MultinomialNB()
visualizer = CVScores(
model, cv=cv, scoring='f1_weighted', size=(1080, 720)
)
visualizer.fit(x_val, y_val)
visualizer.poof()
# -
| codeworks/capstone.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="aenHhwSWhMGu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 393.0} outputId="7296d0cd-fd1d-4ff7-9312-57d52b05e64d"
'''
code by <NAME>(<NAME>) @graykode
Reference : https://github.com/prakashpandey9/Text-Classification-Pytorch/blob/master/models/LSTM_Attn.py
'''
import tensorflow as tf
import matplotlib.pyplot as plt
import numpy as np
tf.reset_default_graph()
# Bi-LSTM(Attention) Parameters
embedding_dim = 2
n_hidden = 5 # number of hidden units in one cell
n_step = 3 # all sentence is consist of 3 words
n_class = 2 # 0 or 1
# 3 words sentences (=sequence_length is 3)
sentences = ["i love you", "he loves me", "she likes baseball", "i hate you", "sorry for that", "this is awful"]
labels = [1, 1, 1, 0, 0, 0] # 1 is good, 0 is not good.
word_list = " ".join(sentences).split()
word_list = list(set(word_list))
word_dict = {w: i for i, w in enumerate(word_list)}
vocab_size = len(word_dict)
input_batch = []
for sen in sentences:
input_batch.append(np.asarray([word_dict[n] for n in sen.split()]))
target_batch = []
for out in labels:
target_batch.append(np.eye(n_class)[out]) # ONE-HOT : To using Tensor Softmax Loss function
# LSTM Model
X = tf.placeholder(tf.int32, [None, n_step])
Y = tf.placeholder(tf.int32, [None, n_class])
out = tf.Variable(tf.random_normal([n_hidden * 2, n_class]))
embedding = tf.Variable(tf.random_uniform([vocab_size, embedding_dim]))
input = tf.nn.embedding_lookup(embedding, X) # [batch_size, len_seq, embedding_dim]
lstm_fw_cell = tf.nn.rnn_cell.LSTMCell(n_hidden)
lstm_bw_cell = tf.nn.rnn_cell.LSTMCell(n_hidden)
# output : [batch_size, len_seq, n_hidden], states : [batch_size, n_hidden]
output, final_state = tf.nn.bidirectional_dynamic_rnn(lstm_fw_cell,lstm_bw_cell, input, dtype=tf.float32)
# Attention
output = tf.concat([output[0], output[1]], 2) # output[0] : lstm_fw, output[1] : lstm_bw
final_hidden_state = tf.concat([final_state[1][0], final_state[1][1]], 1) # final_hidden_state : [batch_size, n_hidden * num_directions(=2)]
final_hidden_state = tf.expand_dims(final_hidden_state, 2) # final_hidden_state : [batch_size, n_hidden * num_directions(=2), 1]
attn_weights = tf.squeeze(tf.matmul(output, final_hidden_state), 2) # attn_weights : [batch_size, n_step]
soft_attn_weights = tf.nn.softmax(attn_weights, 1)
context = tf.matmul(tf.transpose(output, [0, 2, 1]), tf.expand_dims(soft_attn_weights, 2)) # context : [batch_size, n_hidden * num_directions(=2), 1]
context = tf.squeeze(context, 2) # [batch_size, n_hidden * num_directions(=2)]
model = tf.matmul(context, out)
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits_v2(logits=model, labels=Y))
optimizer = tf.train.AdamOptimizer(0.001).minimize(cost)
# Model-Predict
hypothesis = tf.nn.softmax(model)
predictions = tf.argmax(hypothesis, 1)
# Training
with tf.Session() as sess:
init = tf.global_variables_initializer()
sess.run(init)
for epoch in range(5000):
_, loss, attention = sess.run([optimizer, cost, soft_attn_weights], feed_dict={X: input_batch, Y: target_batch})
if (epoch + 1)%1000 == 0:
print('Epoch:', '%06d' % (epoch + 1), 'cost =', '{:.6f}'.format(loss))
# Test
test_text = 'sorry hate you'
tests = [np.asarray([word_dict[n] for n in test_text.split()])]
predict = sess.run([predictions], feed_dict={X: tests})
result = predict[0][0]
if result == 0:
print(test_text,"is Bad Mean...")
else:
print(test_text,"is Good Mean!!")
fig = plt.figure(figsize=(6, 3)) # [batch_size, n_step]
ax = fig.add_subplot(1, 1, 1)
ax.matshow(attention, cmap='viridis')
ax.set_xticklabels([''] + ['first_word', 'second_word', 'third_word'], fontdict={'fontsize': 14}, rotation=90)
ax.set_yticklabels([''] + ['batch_1', 'batch_2', 'batch_3', 'batch_4', 'batch_5', 'batch_6'], fontdict={'fontsize': 14})
plt.show()
| 4-3.Bi-LSTM(Attention)/Bi_LSTM(Attention)_Tensor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stark effect calculations
# Demonstrate the calculation of the Stark effect for the lowest few states of hydrogen, and the resulting UV/VIS spectrum
from sympy import * # use sympy math library
init_printing()
n, l, m = symbols('n, l, m', integer=True)
n_max = 3 # highest value of n to use in the calculation
from sympy.physics.hydrogen import *
r, th, ph = symbols('r, th, ph')
n, l, m = symbols('n,l,m')
integrate(Psi_nlm(3,2,0,r,ph,th)*r*cos(th)*Psi_nlm(3,1,0,r,ph,th)*r**2*sin(th),(ph,0,2*pi),(th,0,pi),(r,0,oo))
integrate(Psi_nlm(3,1,0,r,ph,th)*r*cos(th)*Psi_nlm(3,0,0,r,ph,th)*r**2*sin(th),(ph,0,2*pi),(th,0,pi),(r,0,oo))
# Using the rule $Y_{\ell,m}^* = -Y_{\ell,-m} $:
integrate(-Psi_nlm(3,2,-1,r,ph,th)*r*cos(th)*Psi_nlm(3,1,1,r,ph,th)*r**2*sin(th),(ph,0,2*pi),(th,0,pi),(r,0,oo))
integrate(-Psi_nlm(3,2,1,r,ph,th)*r*cos(th)*Psi_nlm(3,1,-1,r,ph,th)*r**2*sin(th),(ph,0,2*pi),(th,0,pi),(r,0,oo))
| StarkEffect.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Python Worksheet
# 1)
x = 'hello world'
print(x)
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
x = 5
x = (x**3)-25
print(x)
y = np.linspace(0,10,11)
print(y[0], y[1], y[-1], y[1:5], y[5:])
mask = (y<3)
print(mask)
print(y[mask])
print(y[y<3])
a = np.array([0,1,2,3])
b = np.array([0,15,30,45])
print(2*a + a*b)
x = 14.35244
print("%d"%(x))
print("%.3f"%(x))
print("%.4e"%(x))
print("%05d"%(x))
def myfunc1(x):
return(x**2)
myfunc1(2.7)
# 2)
def linfit(xdata, ydata, yerror):
def u_ni(n):
i = 0
u_ni = []
while(i<=len(xdata)-1):
a = ((xdata[i])**n)/((yerror[i])**2)
u_ni.append(a)
i += 1
return(u_ni)
def w_ni(n):
i = 0
w_ni = []
while(i<=len(ydata)-1):
b = ((ydata[i])*((xdata[i])**n))/((yerror[i])**2)
w_ni.append(b)
i += 1
return(w_ni)
def u_nsum(n):
u = u_ni(n)
t = np.sum(u)
return(t)
def w_nsum(n):
w = w_ni(n)
c = np.sum(w)
return(c)
d = (u_nsum(0))*(u_nsum(2)) - (u_nsum(1))**2
slope = ((u_nsum(0))*(w_nsum(1)) - (u_nsum(1))*(w_nsum(0)))/d
intercept = ((u_nsum(2))*(w_nsum(0)) - (u_nsum(1))*(w_nsum(1)))/d
sigma_slope = (u_nsum(0))/d
sigma_intercept = (u_nsum(2))/d
return([slope, intercept, sigma_slope, sigma_intercept])
# +
x = np.array([0.,1.,2.,3.,4.,5.])
y = np.array([-8.0,-12.8,-17.7,-23.3,-27.6,-31.7])
yerr = np.array([0.2,-0.5,-0.9,-1.6,-2.0,-2.5])
d = linfit(x,y,yerr)
def fit(t):
return d[0]*t + d[1]
t = np.linspace(0,8)
print("Results")
print("slope = %.3f +/- %.3f"%(d[0], d[1]))
print("intercept = %.3f +/- %.3f"%(d[2], d[3]))
plt.title('Linear Regression')
plt.xlabel('x axis')
plt.ylabel('y axis')
plt.grid()
plt.plot(t,fit(t), label = 'Fit line ($y = %.5sx + %.5s$)'%(d[0],d[2]))
plt.errorbar(x,y,yerr,ls='none',marker='.',c='k', label = 'y error')
plt.plot(x,y, marker = 'o', linestyle = 'none', label = 'Data points')
plt.legend()
plt.savefig('linear_regression_wksh.pdf')
# -
# 3)
# +
def z(f,R,L,C):
return ( (1/R) + (1/(np.pi*(f)*L*2j)) + (np.pi*(f)*C*2j) )**(-1)
rang = np.linspace(1,20000,1000)
complex_z = z(rang,2000,.005,300e-9)
mag_z = np.abs(complex_z)
arg_z = np.angle(complex_z)
complex_z2 = z(rang,4000,.005,300e-9)
mag_z2 = np.abs(complex_z2)
arg_z2 = np.angle(complex_z2)
plt.figure(1)
plt.plot(rang,mag_z,'r', label = '2000 K-Ohms')
plt.plot(rang,mag_z2,'b', label = '4000 K-Ohms')
plt.title('Magnitude vs Frequency')
plt.xlabel('Frequency')
plt.ylabel('Magnitude')
plt.legend()
plt.figure(2)
plt.plot(rang,arg_z,'r', label = '2000 K-Ohms')
plt.plot(rang,arg_z2,'b', label = '4000 K-Ohms')
plt.title('Phase vs Frequency')
plt.xlabel('Frequency')
plt.ylabel('Phase')
plt.legend()
plt.savefig('Impedence.pdf')
# -
# 4)
# +
lower = 0
upper = 100
p = []
print("Prime numbers between",lower,"and",upper,"are:")
for num in range(lower,upper + 1):
# prime numbers are greater than 1
if num > 1:
for i in range(2,num):
if (num % i) == 0:
break
else:
print(num)
# -
| Python Worksheet v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#hide
# %load_ext autoreload
# %autoreload 2
# +
# default_exp seasonal
# -
# # Seasonal Components
#
# > This module contains functions to define the seasonal components in a DGLM. These are *harmonic* seasonal components, meaning they are defined by sine and cosine functions with a specific period. For example, when working with daily time series we often use a weekly seasonal effect of period 7 or an annual seasonal effect of period 365.
#hide
#exporti
import numpy as np
from pybats.forecast import forecast_aR, forecast_R_cov
# ## Seasonal Components for a DGLM
#export
def seascomp(period, harmComponents):
p = len(harmComponents)
n = 2*p
F = np.zeros([n, 1])
F[0:n:2] = 1
G = np.zeros([n, n])
for j in range(p):
c = np.cos(2*np.pi*harmComponents[j]/period)
s = np.sin(2*np.pi*harmComponents[j]/period)
idx = 2*j
G[idx:(idx+2), idx:(idx+2)] = np.array([[c, s],[-s, c]])
return [F, G]
# This function is called from `dglm.__init__` to define the seasonal components.
#exporti
def createFourierToSeasonalL(period, harmComponents, Fseas, Gseas):
p = len(harmComponents)
L = np.zeros([period, 2*p])
L[0,:] = Fseas.reshape(-1)
for i in range(1, period):
L[i,:] = L[i-1,:] @ Gseas
return L
#export
def fourierToSeasonal(mod, comp=0):
phi = mod.L[comp] @ mod.m[mod.iseas[comp]]
var = mod.L[comp] @ mod.C[np.ix_(mod.iseas[comp], mod.iseas[comp])] @ mod.L[comp].T
return phi, var
# This function transforms the seasonal component of a model from fourier form into more interpretable seasonal components. For example, if `seasPeriods = [7]`, then this would return a vector of length $7$, with each of the seasonal effects.
#
# A simple use case is given below. For a more detailed use of this function, see the following [example](https://github.com/lavinei/pybats_nbdev/blob/master/examples/Poisson_DGLM_In_Depth_Example.ipynb).
# +
import numpy as np
import pandas as pd
from pybats.analysis import analysis
from pybats.shared import load_sales_example2
data = load_sales_example2()
prior_length = 21 # Number of days of data used to set prior
mod = analysis(data.Sales.values, data[['Price', 'Promotion']].values, k=1,
family='poisson',
seasPeriods=[7], seasHarmComponents=[[1,2,3]],
prior_length=prior_length, dates=data.index,
ret = ['model'])
# +
seas_mean, seas_cov = fourierToSeasonal(mod)
days = ['Monday', 'Tuesday', 'Wednesday', 'Thursday', 'Friday', 'Saturday', 'Sunday']
lastday = data.index[-1]
days = [*days[lastday.isoweekday()-1:], *days[:lastday.isoweekday()-1]]
seas_eff = pd.DataFrame({'Day':days,
'Effect Mean':np.exp(seas_mean.reshape(-1))})
seas_eff
# -
#exporti
def fourierToSeasonalFxnl(L, m, C, iseas):
phi = L @ m[iseas]
var = L @ C[np.ix_(iseas, iseas)] @ L.T
return phi, var
#exporti
def get_seasonal_effect_fxnl(L, m, C, iseas):
phi, var = fourierToSeasonalFxnl(L, m, C, iseas)
return phi[0], var[0, 0]
#exporti
def sample_seasonal_effect_fxnl(L, m, C, iseas, delVar, n, nsamps):
phi_samps = np.zeros([nsamps])
phi, var = fourierToSeasonalFxnl(L, m, C, iseas)
phi_samps[:] = phi[0] + np.sqrt(var[0,0])*np.random.standard_t(delVar*n, size = [nsamps])
return phi_samps
#exporti
def forecast_weekly_seasonal_factor(mod, k, sample = False, nsamps = 1):
a, R = forecast_aR(mod, k)
idx = np.where(np.array(mod.seasPeriods) == 7)[0][0]
if sample:
return sample_seasonal_effect_fxnl(mod.L[idx], a, R, mod.iseas[idx], mod.delVar, mod.n, nsamps)
else:
return get_seasonal_effect_fxnl(mod.L[idx], a, R, mod.iseas[idx])
#exporti
def forecast_path_weekly_seasonal_factor(mod, k, today, period):
phi_mu = [np.zeros([period]) for h in range(k)]
phi_sigma = [np.zeros([period, period]) for h in range(k)]
phi_psi = [np.zeros([period, period, h]) for h in range(1, k)]
idx = np.where(np.array(mod.seasPeriods) == 7)[0][0]
L = mod.L[idx]
iseas = mod.iseas[idx]
for h in range(k):
# Get the marginal a, R
a, R = forecast_aR(mod, h + 1)
m, v = get_seasonal_effect_fxnl(L, a, R, iseas)
day = (today + h) % period
phi_mu[h][day] = m
phi_sigma[h][day, day] = v
# phi_mu[h], phi_sigma[h] = get_latent_factor_fxnl_old((today + h) % period, mod.L, a, R, mod.iseas, mod.seasPeriods[0])
# Find covariances with previous latent factor values
for j in range(h):
# Covariance matrix between the state vector at times j, i, i > j
day_j = (today + j) % period
cov_jh = forecast_R_cov(mod, j, h)[np.ix_(iseas, iseas)]
phi_psi[h-1][day, day_j, j] = phi_psi[h-1][day_j, day, j] = (L @ cov_jh @ L.T)[day, day_j]
# cov_ij = (np.linalg.matrix_power(mod.G, h-j) @ Rlist[j])[np.ix_(mod.iseas, mod.iseas)]
return phi_mu, phi_sigma, phi_psi
#hide
from nbdev.export import notebook2script
notebook2script()
| nbs/04_seasonal.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import os
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from utils.dependencies import *
# %matplotlib inline
warnings.filterwarnings("ignore")
# +
DATA_INTER_PATH = os.path.join('..', 'data', 'interim')
DATA_PROCE_PATH = os.path.join('..', 'data', 'processed')
IMG_PATH = os.path.join('..','reports', 'figures')
DATA_INTER_NAME_2 = 'general_2.csv'
DATA_PROCE_NAME = 'processed.csv'
DATE_CHURN = pd.to_datetime('2018-08-01 00:00:00+00:00')
# +
df_2 = pd.read_csv(os.path.join(DATA_INTER_PATH, DATA_INTER_NAME_2))
df_2['register_date'] = pd.to_datetime(df_2['register_date'])
df_2['last_purchase_date'] = pd.to_datetime(df_2['last_purchase_date'])
df_2.head(2)
# -
df_3 = df_2\
.drop('discount', axis=1)\
.groupby(
by=[
'customer_code',
'group_code',
'segment_code',
'is_churn'
], as_index=False)\
.agg(
{
'register_date': 'max',
'total_price': 'sum',
'item_total_price': 'sum',
'last_purchase_date': 'max'
}
)
(df_3['last_purchase_date'] == df_3['register_date']).unique()
df_3.is_churn.value_counts()
df_3['discount'] = df_3.apply(lambda x: discount(x.item_total_price, x.total_price), axis=1)
df_3.discount.value_counts()
df_3['qnt_days'] = DATE_CHURN - df_3.last_purchase_date
df_3['qnt_days'] = df_3.apply(lambda x: x['qnt_days'].days, axis=1)
df_3 = df_3[
[
'customer_code', 'group_code', 'segment_code',
'total_price', 'item_total_price', 'discount',
'last_purchase_date', 'qnt_days', 'is_churn'
]
]
df_3.head(1)
# +
plt.figure(figsize=(20,6))
corrmat = df_3.corr(method='spearman')
sns.heatmap([corrmat['is_churn']], xticklabels = corrmat.index,
annot=True, fmt='.2f', annot_kws={'size': 14},
cbar=False, center=0)
plt.title('Correlação pelo método de Spearman')
plt.savefig(os.path.join(IMG_PATH,'corr-spearman.png'), format='png')
plt.tight_layout()
plt.show()
# -
df_3.to_csv(os.path.join(DATA_PROCE_PATH, DATA_PROCE_NAME), index=False)
# ## Considerações
#
# 1. Todas as features foram mantidas.
# 2. A variável **discount** representa se o cliente teve algum tipo de desconto até **01 de Ago de 2018**;
# 3. A variável **qnt_days** represemta quantos dias desde a sua ultima compra até **01 de Ago de 2018**.
| notebooks/03-FEATURE_SELECTION.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Adapted from https://github.com/pytorch/examples/blob/master/mnist/main.py
import os
os.chdir('/home/rcgonzal/DSC180Malware/m2v-adversarial-hindroid/')
from __future__ import print_function
from scipy import sparse
import pandas as pd
import numpy as np
import argparse
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torchvision import datasets, transforms
from torch.optim.lr_scheduler import StepLR
from torch.utils.data import Dataset, DataLoader
from sklearn.metrics import classification_report, confusion_matrix
from src.imbalanced_dataset_sampler.imbalanced import ImbalancedDatasetSampler
# -
with open('data/out/all-apps/hindroid-train-set/A_test.npz', 'rb') as file:
print(sparse.load_npz(file).shape)
pd.read_csv('data/out/train-set/method_map.csv').shape
# +
class HindroidDataset(Dataset):
def __init__(self, features_path, labels_path, label_col='m2vDroid', transform=None):
'''
Creates a dataset from the A matrix representation of apps and their associated labels.
Parameters:
-------------------
features_path: Path to A matrix in sparse format.
labels_path: Path to labels in csv format.
label_col: Default 'm2vDroid'. Useful for specifying which kernel to use for HinDroid.
'''
self.features = sparse.load_npz(os.path.join(features_path))
self.feature_width = self.features.shape[1]
features_folder = os.path.split(features_path)[0]
self.features_idx = list(pd.read_csv(
os.path.join(features_folder, 'predictions.csv'),
usecols=['app'],
squeeze=True
))
self.transform = transform
try:
self.labels = pd.read_csv(
labels_path,
usecols=['app', label_col],
index_col = 'app',
squeeze=True
)
self.labels = self.labels[self.features_idx].values # align labels with features index
except (KeyError, ValueError) as e:
print(e)
print('Seems like you may be trying to use a different model. This class is setup for m2vDroid by default.')
print('For HinDroid you must specify `label_col` as either AAT, ABAT, APAT, ABPBTAT, or APBPTAT.')
assert (self.features.shape[0] == self.labels.size), 'Length mismatch between features and labels.'
def __len__(self):
return self.features.shape[0]
def __getitem__(self, idx):
if torch.is_tensor(idx):
idx = idx.tolist()
features = self.features[idx]
features = features.todense().astype('float').A
labels = self.labels[idx]
# if self.transform:
# features = self.transform(features)
# labels = self.transform(labels)
# sample = {'features': features, 'labels': labels}
return features, labels
def get_labels(self, idx):
return self.labels[idx]
def hindroid_custom_get_label(dataset, idx):
return dataset.get_labels(idx)
# -
class HindroidSubstitute(nn.Module):
def __init__(self, n_features):
super(HindroidSubstitute, self).__init__()
self.layer_1 = nn.Linear(n_features, 64, bias=False)
# Linear - how to freeze layer ^
# biases = false
self.layer_2 = nn.Linear(64, 64, bias=False)
self.layer_3 = nn.Linear(64, 64, bias=False)
self.layer_4 = nn.Linear(64, 2, bias=False)
def forward(self, x):
if not torch.is_tensor(x):
x = torch.from_numpy(x)
x = x.view(x.shape[0], -1)
x = F.relu(self.layer_1(x))
x = F.relu(self.layer_2(x))
x = F.relu(self.layer_3(x))
x = self.layer_4(x)
return x # logits
# +
# Test model
# model = HindroidSubstitute(dataset.feature_width).double()
# model(dataset[-100:][0])
# +
# F.nll_loss?
# +
def train(model, device, train_loader, optimizer, epoch, weight=None):
model.train()
for batch_idx, (data, target) in enumerate(train_loader):
data, target = data.to(device), target.to(device)
optimizer.zero_grad()
output = F.log_softmax(model(data), dim=1)
loss = F.nll_loss(output, target, weight=weight) # do we use different loss?
loss.backward()
optimizer.step()
# logging
log_interval = 100
if batch_idx % log_interval == 0:
print('Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}'.format(
epoch, batch_idx * len(data), len(train_loader.dataset),
100. * batch_idx / len(train_loader), loss.item()))
# if batch_idx % args.log_interval == 0:
# print('Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}'.format(
# epoch, batch_idx * len(data), len(train_loader.dataset),
# 100. * batch_idx / len(train_loader), loss.item()))
# if args.dry_run:
# break
def test(model, device, test_loader, weight=None):
model.eval()
test_loss = 0
correct = 0
with torch.no_grad():
for data, target in test_loader:
data, target = data.to(device), target.to(device)
output = model(data)
# print(output)
output = F.log_softmax(output, dim=1)
# print(output)
loss = F.nll_loss(output, target, weight=weight, reduction='sum').item() # sum up batch loss
# print('loss: ', loss)
test_loss += loss
# print(output.argmax(dim=1, keepdim=True))
pred = output.argmax(dim=1, keepdim=True) # get the index of the max log-probability
# print(target.view_as(pred))
correct += pred.eq(target.view_as(pred)).sum().item()
test_loss /= len(test_loader.dataset)
print('\nTest set: Average loss: {:.4f}, Accuracy: {}/{} ({:.0f}%)\n'.format(
test_loss, correct, len(test_loader.dataset),
100. * correct / len(test_loader.dataset)))
# +
# model = HindroidSubstitute(dataset.feature_width).double()
# weights = torch.Tensor([dataset.labels.mean() / (1-dataset.labels.mean()), 1]).double()
# train_loader = torch.utils.data.DataLoader(dataset, batch_size=10)
# test_loader = torch.utils.data.DataLoader(dataset, batch_size=10)
# test(model, 'cpu', test_loader, weights)
# -
# # Train
# +
use_cuda = torch.cuda.is_available()
# use_cuda = False
# torch.manual_seed(args.seed)
device = torch.device("cuda" if use_cuda else "cpu")
# train_kwargs = {'batch_size': args.batch_size}
# test_kwargs = {'batch_size': args.test_batch_size}
if use_cuda:
cuda_kwargs = {'num_workers': 1,
# 'shuffle': True,
'pin_memory': True}
# train_kwargs.update(cuda_kwargs)
# test_kwargs.update(cuda_kwargs)
# load data (will need to be adapted as well)
# 1) load A_test
# 2) load labels
# 3) Perform train-test-split
dataset = HindroidDataset(
'data/out/all-apps/hindroid-train-set/A_test.npz',
'data/out/all-apps/hindroid-train-set/predictions.csv',
'ABAT',
)
# weights = torch.Tensor([dataset.labels.mean() / (1-dataset.labels.mean()), 1]).to(device).double()
weights = None
train_loader = torch.utils.data.DataLoader(
dataset,
batch_size=5,
shuffle = False,
sampler = ImbalancedDatasetSampler(dataset, callback_get_label = hindroid_custom_get_label),
**cuda_kwargs)
test_loader = torch.utils.data.DataLoader(
dataset,
batch_size=5,
shuffle = False,
sampler = ImbalancedDatasetSampler(dataset, callback_get_label = hindroid_custom_get_label),
**cuda_kwargs)
# train_loader = torch.utils.data.DataLoader(dataset, batch_size=10)
# test_loader = torch.utils.data.DataLoader(dataset, batch_size=10)
model = HindroidSubstitute(dataset.feature_width).double().to(device)
optimizer = optim.Adadelta(model.parameters(), lr=0.005)
# -
scheduler = StepLR(optimizer, step_size=1)
for epoch in range(1, 20 + 1):
train(model, device, train_loader, optimizer, epoch, weights)
test(model, device, test_loader, weights)
scheduler.step()
# +
test_loader = torch.utils.data.DataLoader(
dataset,
batch_size=10,
**cuda_kwargs)
model.eval()
all_preds = []
with torch.no_grad():
for data, target in test_loader:
data, target = data.to(device), target.to(device)
output = model(data)
output = F.log_softmax(output, dim=1)
pred = output.argmax(dim=1, keepdim=True) # get the index of the max log-probability
all_preds.extend(pred.tolist())
print(classification_report(dataset.labels, all_preds))
print(confusion_matrix(dataset.labels, all_preds))
# -
with open(os.path.join(outfolder, 'NN_sub.pkl'), 'wb') as file:
torch.save(model, file)
# +
outfolder = 'data/out/all-apps/hindroid-train-set-ABAT/'
with open(os.path.join(outfolder, 'NN_sub.pkl'), 'rb') as file:
model = torch.load(file).to(device)
batch_size = 10
test_loader = torch.utils.data.DataLoader(
dataset,
batch_size=batch_size,
shuffle = True,
# sampler = ImbalancedDatasetSampler(dataset, callback_get_label = hindroid_custom_get_label),
**cuda_kwargs)
# +
from src.attack.cw import to_tanh_space, from_tanh_space, L2Adversary
# %load_ext autoreload
# %autoreload 2
# +
cw_attack = L2Adversary(
targeted=False, confidence=0.0, c_range=(1e-1, 1e10),
search_steps=3, max_steps=1000, abort_early=True,
box=(0., 1.), optimizer_lr=1e-2, init_rand=True
)
advxs = None
all_inputs = []
all_advx = []
for data, target in test_loader:
if len(all_advx) * batch_size >= 500:
break
print("OG Labels: ", target)
target = F.softmax(model(data.to(device)), dim=1)
print("Model pred: ", target)
target = torch.argmax(target, dim=1)
print(target)
data = data.view(batch_size, -1)
with open(os.path.join(outfolder, 'input.npz'), 'wb') as file:
all_inputs.append(sparse.csr_matrix(data))
sparse.save_npz(file, sparse.csr_matrix(data))
data, target = data.to(device), target.to(device)
advxs = cw_attack(model, data, target, to_numpy=False)
sparse_advxs = sparse.csr_matrix(torch.round(advxs), dtype='i1')
all_advx.append(sparse_advxs)
with open(os.path.join(outfolder, 'advxs.npz'), 'wb') as file:
sparse.save_npz(file, sparse_advxs)
all_inputs = sparse.vstack(all_inputs)
all_advx = sparse.vstack(all_advx)
# +
# attack_folder = os.path.join('data', 'out', 'all-apps', 'hind')
# os.makedirs(attack_folder, exist_ok=True)
with open(os.path.join(outfolder, 'inputs.npz'), 'wb') as file:
sparse.save_npz(file, sparse.csr_matrix(all_inputs))
with open(os.path.join(outfolder, 'advxs.npz'), 'wb') as file:
sparse.save_npz(file, sparse.csr_matrix(all_advx))
# -
with open('advxs.npz', 'wb') as file:
sparse.save_npz(file, sparse.csr_matrix(advxs))
torch.min(torch.abs(advxs - 0.5))
torch.argmax(F.softmax(model(torch.round(advxs.double().to(device))), dim=1), dim=1)
mat = sparse.csr_matrix(torch.round(advxs))
density = mat.getnnz() / np.prod(mat.shape)
density
mat.data.nbytes + mat.indptr.nbytes + mat.indices.nbytes
2535703 * 1e-5
with open('advxs.npz', 'wb') as file:
sparse.save_npz(file, sparse.csr_matrix(advxs))
to_tanh_space(torch.Tensor(np.arange(-1,1,.001)).double(), [0., 1.])
pd.Series(to_tanh_space(torch.Tensor(np.arange(-1,1,.001)).double(), [0., 1.]).tolist(),
index=np.arange(-1,1,.001)).plot()
inputs = torch.Tensor(np.random.randint(0,2, (3,4)))
inputs
to_tanh_space(inputs, [0., 1.])
torch.clamp(to_tanh_space(inputs, [0., 1.]), min=-1e4)
from_tanh_space(torch.clamp(to_tanh_space(inputs, [0., 1.]), min=-1e4), [0., 1.])
torch.clamp(inputs + from_tanh_space(pert_tanh*1000, [0., 1.]), max=1.0)
pert_tanh = torch.zeros((3,4)) # type: torch.FloatTensor
nn.init.normal_(pert_tanh, mean=0, std=1)
pert_tanh
from_tanh_space(pert_tanh*1e4, [0., 1.])
torch.clamp(inputs + from_tanh_space(pert_tanh*1e4, [0., 1.]), max=1.0)
pd.Series(from_tanh_space(
(to_tanh_space(torch.Tensor(np.arange(-1,1,.001)).double(), [-1., 1.])*1000), [-1., 1]).tolist(),
index=to_tanh_space(torch.Tensor(np.arange(-1,1,.001)).double(), [-1., 1.]).tolist()).plot()
pd.Series(from_tanh_space(torch.Tensor(np.arange(-1000, 1000, 1)), box=[0,1]).tolist(),
index=np.arange(-1000, 1000, 1)).plot()
from_tanh_space(to_tanh_space(torch.Tensor([0,.999]).double(), [-1., 1.]), [-1., 1])
# +
# def main():
# # Training settings
# parser = argparse.ArgumentParser(description='PyTorch MNIST Example')
# parser.add_argument('--batch-size', type=int, default=64, metavar='N',
# help='input batch size for training (default: 64)')
# parser.add_argument('--test-batch-size', type=int, default=1000, metavar='N',
# help='input batch size for testing (default: 1000)')
# parser.add_argument('--epochs', type=int, default=14, metavar='N',
# help='number of epochs to train (default: 14)')
# parser.add_argument('--lr', type=float, default=1.0, metavar='LR',
# help='learning rate (default: 1.0)')
# parser.add_argument('--gamma', type=float, default=0.7, metavar='M',
# help='Learning rate step gamma (default: 0.7)')
# parser.add_argument('--no-cuda', action='store_true', default=False,
# help='disables CUDA training')
# parser.add_argument('--dry-run', action='store_true', default=False,
# help='quickly check a single pass')
# parser.add_argument('--seed', type=int, default=1, metavar='S',
# help='random seed (default: 1)')
# parser.add_argument('--log-interval', type=int, default=10, metavar='N',
# help='how many batches to wait before logging training status')
# parser.add_argument('--save-model', action='store_true', default=False,
# help='For Saving the current Model')
# args = parser.parse_args()
use_cuda = not args.no_cuda and torch.cuda.is_available()
torch.manual_seed(args.seed)
device = torch.device("cuda" if use_cuda else "cpu")
train_kwargs = {'batch_size': args.batch_size}
test_kwargs = {'batch_size': args.test_batch_size}
if use_cuda:
cuda_kwargs = {'num_workers': 1,
'pin_memory': True,
'shuffle': True}
train_kwargs.update(cuda_kwargs)
test_kwargs.update(cuda_kwargs)
# load data (will need to be adapted as well)
# 1) load A_test
# 2) load labels
# 3) Perform train-test-split
# transform=transforms.Compose([
# transforms.ToTensor(),
# transforms.Normalize((0.1307,), (0.3081,))
# ])
# dataset1 = datasets.MNIST('../data', train=True, download=True,
# transform=transform)
# dataset2 = datasets.MNIST('../data', train=False,
# transform=transform)
dataset = HindroidDataset(
'data/out/all-apps/hindroid-train-half/A_test.npz',
'data/out/all-apps/hindroid-train-half/predictions.csv',
'AAT'
)
train_loader = torch.utils.data.DataLoader(dataset,**train_kwargs)
test_loader = torch.utils.data.DataLoader(dataset, **test_kwargs)
model = HindroidSubstitute().to(device)
optimizer = optim.Adadelta(model.parameters(), lr=args.lr)
scheduler = StepLR(optimizer, step_size=1, gamma=args.gamma)
for epoch in range(1, args.epochs + 1):
train(args, model, device, train_loader, optimizer, epoch)
test(model, device, test_loader)
scheduler.step()
if args.save_model:
torch.save(model.state_dict(), "mnist_cnn.pt")
# if __name__ == '__main__':
# main()
| notebooks/Attack.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
tf.compat.v1.enable_eager_execution()
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import os
import sys
sys.path.insert(0, 'G:\\My Drive\\Colab Notebooks\\tensorflow-wavelets\src')
from tensorflow_wavelets.utils.cast import *
from tensorflow_wavelets.Layers.DWT import *
from sklearn.metrics import accuracy_score, precision_score, recall_score
from sklearn.model_selection import train_test_split
from tensorflow.keras import layers, losses
from tensorflow.keras.datasets import mnist
from tensorflow.keras.models import Model
# +
(x_train, _), (x_test, _) = mnist.load_data()
x_train = x_train.astype('float32') / 255.
x_test = x_test.astype('float32') / 255.
x_train = np.expand_dims(x_train, axis=-1)
x_test = np.expand_dims(x_test, axis=-1)
print (x_train.shape)
print (x_test.shape)
# +
latent_dim = 64
input_shape=(28, 28, 1)
class Autoencoder(Model):
def __init__(self, latent_dim):
super(Autoencoder, self).__init__()
self.latent_dim = latent_dim
self.encoder = tf.keras.Sequential([
DWT(name="db2", concat=1),
layers.Flatten(),
layers.Dense(latent_dim, activation='sigmoid'),
])
self.decoder = tf.keras.Sequential([
layers.Dense(900, activation='sigmoid'),
layers.Reshape((30, 30, 1)),
IDWT(name="db2", splited=0),
])
def call(self, x):
encoded = self.encoder(x)
decoded = self.decoder(encoded)
return decoded
autoencoder = Autoencoder(latent_dim)
autoencoder.compile(optimizer='adam', loss=losses.MeanSquaredError())
# -
autoencoder.fit(x_train, x_train,
epochs=10,
shuffle=True,
validation_data=(x_test, x_test))
encoded_imgs = autoencoder.encoder(x_test).numpy()
decoded_imgs = autoencoder.decoder(encoded_imgs).numpy()
n = 10
plt.figure(figsize=(20, 4))
for i in range(n):
# display original
ax = plt.subplot(2, n, i + 1)
plt.imshow(x_test[i])
plt.title("original")
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
# display reconstruction
ax = plt.subplot(2, n, i + 1 + n)
plt.imshow(decoded_imgs[i])
plt.title("reconstructed")
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
# +
model = keras.Sequential()
model.add(keras.Input(shape=input_shape))
model.add(DWT(name="db2", concat=0))
model.add(keras.layers.Flatten())
model.add(keras.layers.Dense(64, activation="sigmoid"))
model.add(keras.layers.Dense(786, activation="sigmoid"))
model.add(keras.layers.Reshape((30, 30, 1)))
model.add(IDWT(name="db2", splited=0))
model.summary()
# -
| Development/AutoEncoders_compare/AutoEncoder_DWT_db2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import skimage
from skimage import data,io,img_as_float
from skimage.morphology import convex_hull_image
import matplotlib
from matplotlib import pyplot as plt
from skimage.util import invert
horse_image=skimage.io.imread(r'C:\Users\user\AppData\Local\Programs\Python\Python37\Lib\site-packages\skimage\data\cell.png')
horse_inverted=invert(horse_image)
fig,axes=plt.subplots(1,2,figsize=(12,8),sharex=True,sharey=True)
ax=axes.ravel()
ax[0].set_title('Original')
ax[0].imshow(horse_image, cmap='gray')
ax[1].imshow(horse_inverted,cmap='gray',interpolation='nearest')
ax[1].set_title('inverted')
fig.tight_layout()
plt.show()
# -
| Convex Hull.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#read file
import pandas as pd
filepath = "pollution_us_2000_2016.csv"
us_pollution_data = pd.read_csv(filepath)
us_pollution_data.head()
# -
us_pollution_data.columns
# # Create a Year Column:
# +
#Create Year column. which we will use to group the data by year.
Date_data = []
#run a loop on the 'Date Local' Column
for i in us_pollution_data['Date Local']:
#Split each row (which is date in string format) into a list
date = i.split("-")
#The first element of the list will be the year.
year= date[0]
#Append the year to a list of dates
Date_data.append(year)
#Create a year column using Date data
us_pollution_data['Year'] = Date_data
#clean data by dropping rows with null values.
us_pollution_data_clean = us_pollution_data.dropna(how='any')
# -
# # Select Relevant Columns:
#Create a collection of columns of interest
Chem_Columns_data = us_pollution_data_clean[['State', 'Year', 'SO2 AQI', 'CO AQI', 'O3 AQI', 'NO2 AQI']]
#reindex data to set indices starting from 0
Chem_Columns_data = Chem_Columns_data.reset_index()
#drop the index column
Chem_Columns_data = Chem_Columns_data.drop(columns=['index'])
Chem_Columns_data.head()
# # Group by Year and State
Chem_by_ST_Year = Chem_Columns_data.groupby(['Year', 'State']).mean()
Chem_by_ST_Year.head()
# # Create a State Average Column
# +
#Create a column for the average of the mean measurements
Mean_us_pol = []
for index, row in Chem_by_ST_Year.iterrows():
avg_pol = (row[0] + row[1] + row[2] + row[3])/4
Mean_us_pol.append(avg_pol)
Chem_by_ST_Year['State Average'] = Mean_us_pol
Chem_by_ST_Year.head()
# -
# # Create a Year's Average for Each Pollutant AQI
Chem_by_Year = Chem_Columns_data.groupby(['Year']).mean()
Chem_by_Year.head()
# # Save CSVs
#save DataFrame as Csv for easy access to Dataframe data
Chem_by_ST_Year.to_csv("../DataFrames/Chem_mean_by_State.csv", header=True)
#save second DataFrame as Csv for easy access to Dataframe data
Chem_by_Year.to_csv("../DataFrames/Chem_mean_by_Year.csv", header=True)
# # Create a Final Pandas DataFrame With State and City Information
AQI_data = us_pollution_data_clean[['State Code', 'County Code', 'State', 'County', 'City','Year','NO2 AQI', 'O3 AQI', 'SO2 AQI', 'CO AQI']]
AQI_data.head()
AQI_data.to_csv("../DataFrames/AQI_data.csv", header=True)
| World_Heatmaps_US_Pollution_Trend/US poll dataframes/Final US Pollution Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Analysis of Cosine-Similarity Model
#
# In this notebook, a parsimonious version of K-Nearest Neighbors (dubbed Cosine-Similarity Model) is proposed that results in a slightly higher accuracy than standard K-Nearest Neighbor models, along with a 2 times (or greater) classification speedup. The models's speed and accuracy is a reminder that building in-house models is often the best option when solving specific problems, as opposed to using pre-built models.
#
# ## Introduction
#
# #### Standard K-Nearest Neighbors
# K-Nearest Neighbor (K-NN) algorithms are simple supervised classification algorithms that have the capability of making accurate predictions on complex problems. K-NN is a lazy learner, so it requires no training and can thus get right to classification, making it a much faster algorithm than other classification models such as SVM, regression, multi-layer perceptron, etc.. K-NN is also non-parametric, so it makes to assumptions about the data. Because the algorithm requires no training, data can be added or taken away seamlessly, without making any major adjustments.
#
# Given a point $p$ to classify, a K-NN model will "compare" the passed point with each point $x_i$ the model has available to it, using some distance metric (most commonly Euclidean distance). This process will generate the unordered set $D$ that holds the distances (similarities) between $p$ and every other point in the dataset, $x_i$, giving $d_i$. Next, the algorithm pulls the $k$ lowest distances (greatest similarities) from $D$, and uses some voting technique to classify $p$ as being a member of some class $C$. See [this](https://importq.wordpress.com/2017/11/24/mnist-analysis-using-knn/) tutorial for a more thorough description of K-NN.
#
# #### The Cosine-Similarity Model
# The Cosine-Similarity Model works in the same general way as most K-NN classifiers. The primary difference with the Cosine-Similarity Classifier is in its name: it uses cosine-similarity as a distance metric instead of standard Euclidean or Manhatten distance (which are far more common then cosine similarity). Cosine-similarity is given by
#
# $$
# similarity=cos(\theta) = \frac{\vec{a} \cdot \vec{b}}{||\vec{a}|| ||\vec{b}||}
# $$
#
# where $\vec{a}$ and $\vec{b}$ are vectors whos similarity is returned. Cosine similarity is often not a perfect distance metric, as it doesn't work on negative data, and violates the triangle inequality. However for certain problems (as shown below) it is a solid choice.
#
# After testing the Cosine-Similarity Model on the MNIST data set, it is found that the Cosine-Similarity classifier is both faster and just as, if not more accurate than go-to K-NN models from the Scikit-Learn library. In the analysis below, I will build out the Cosine-Similarity Model, and run it on the MNIST data set. I will then test a go-to K-NN model from Scikit-Learn on the MNIST dataset, finally comparing both the accuracy and classification time of the two models in a variety of situations. All tests were run on a Intel Core 3570K CPU (no GPU this time :( ).
#
# ## Analysis
#
# Start with required imports
# +
import numpy as np
import heapq
from collections import Counter
from sklearn.metrics.pairwise import cosine_similarity
from sklearn import datasets, model_selection
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import classification_report, confusion_matrix
mnist = datasets.fetch_mldata('MNIST original')
data, target = mnist.data, mnist.target
# -
data.shape, target.shape
# Make different data sets out of the full MNIST data set for testing the Cosine-Similarity Model and Scikit-Learn K-NN under different situations.
# +
# make an array of indices to use for making the data sets, in random order, the same length of the MNIST dataset
indx = np.random.choice(len(target), 70000, replace=False)
# method for building datasets to test with
def mk_dataset(size):
"""makes a dataset of size "size", and returns that datasets images and targets
This is used to make the dataset that will be stored by a model and used in
experimenting with different stored dataset sizes
"""
train_img = [data[i] for i in indx[:size]]
train_img = np.array(train_img)
train_target = [target[i] for i in indx[:size]]
train_target = np.array(train_target)
return train_img, train_target
################################################
# TEST SETS
# will be keeping test set the same for different stored data sets
# test_data/test_target: the smaller dataset used to test model accuracy for the data sets
test_img = [data[i] for i in indx[60000:70000]]
test_img = np.array(test_img)
test_target = [target[i] for i in indx[60000:70000]]
test_target = np.array(test_target)
# Single random test image, this is just used for testing the speed at which each model can
# classify a single point
t_img = test_img[563]
t_img = np.array([t_img])
t_target = test_target[563]
t_target = np.array([t_target])
t_target
test_img.shape, test_target.shape, t_target
# -
# make different datasets
sixty_x, sixty_y = mk_dataset(60000)
sixty_x.shape, sixty_y.shape
fifty_x, fifty_y = mk_dataset(50000)
fifty_x.shape, fifty_y.shape
fourty_x, fourty_y = mk_dataset(40000)
fourty_x.shape, fourty_y.shape
thirty_x, thirty_y = mk_dataset(30000)
thirty_x.shape, thirty_y.shape
twenty_x, twenty_y = mk_dataset(20000)
twenty_x.shape, twenty_y.shape
ten_x, ten_y = mk_dataset(10000)
ten_x.shape, ten_y.shape
one_x, one_y = mk_dataset(1000)
one_x.shape, one_y.shape
# Great. Now we have 7 data sets to test on the classifier, ranging from size 1,000 to size 60,000. We also have a testing data set of size 10,000 to calculate accuracy and speed of the classifiers, as well as a smaller testing dataset of size 1 used for computing sinlge point classification speed.
#
# Now we build the Cosine-Similarity Classifier. The method only takes the `test_target` argument to calculate prediction accuracy, it is not actually needed for classification.
def cos_knn(k, test_data, test_target, stored_data, stored_target):
"""k: number of neighbors to use for voting
test_data: a set of unobserved images to classify
test_target: the labels for the test_data (for calculating accuracy)
stored_data: the images already observed and available to the model
stored_target: labels for stored_data
"""
# find similarity for every point in test_data between every other point in stored_data
cosim = cosine_similarity(test_data, stored_data)
# get indices of images in stored_data that are most similar to any given test_data point
top = [(heapq.nlargest((k+1), range(len(i)), i.take)) for i in cosim]
# convert indices to numbers
top = [[stored_target[j] for j in i[:k]] for i in top]
# vote, and return prediction for every image in test_data
pred = [max(set(i), key=i.count) for i in top]
pred = np.array(pred)
# print table giving classifier accuracy using test_target
print(classification_report(test_target, pred))
# That is all, the developed Cosine-Similarity Model is very straight forward.
#
# For the Scikit-Learn K-NN model, all we have to worry about is the value for the `n_neighbors` argument (number of neighbors to use for classification), the `weights` argument for `KNeighborsClassifier()` (how to vote), which we will just leave at its default value of `uniform`, as that is the same method used in the Cosine-Similarity Model. Finally, we have the `algorithm` argument for `KNeighborsClassifier()`, which we will also leave at its default value of `auto`, as it will find the optimal algorithm to use for the given data. (Note, I ran a few tests with the `KNeighborsClassifier()` `metric` argument set to `cosine_similarity`. I did not notice a difference so I will just be leaving `metric` at its default value).
def skl_knn(k, test_data, test_target, stored_data, stored_target):
"""k: number of neighbors to use in classication
test_data: the data/targets used to test the classifier
stored_data: the data/targets used to classify the test_data
"""
classifier = KNeighborsClassifier(n_neighbors=k)
classifier.fit(stored_data, stored_target)
y_pred = classifier.predict(test_data)
print(classification_report(test_target, y_pred))
# ### Testing
# Now that we have the models setup, we will test how each model performs on each of the seven data sets.
#
# For each data set/model pair, we will be measuring classification accuracy and speed of `test_data` classification. For the Scikit-Learn model, a $k$ value of 5 will be used, and for the Cosine-Similarity model a $k$ value of 3 will be used, as those were the values found to be optimal (no techniques such as cross validation were used here for finding $k$, just performance on a few example tests).
#
# The tests on each data set alternate between the Cosine-Similarity Model (`cos_knn`) and the Scikit-Learn K-NN model (`skl_knn`).
#
# **NOTE**: immidiately below I show the Cosine-Similarity Model run with a $k$ size of 5, to show that the major difference in classification speed between the Cosine-Similarity Model and the Scikit-Learn model does not have very much to do with the different sizes of $k$ (in the case below, using a larger $k$ value on the Cosine similarity model actually resulted in a faster classification time than when the model ran with a $k$ value of 3. The classification time with a larger $k$ can vary, and is usually about 10-30 seconds slower than running with a $k$ value of 3).
# %%time
cos_knn(5, test_img, test_target, sixty_x, sixty_y)
# %%time
cos_knn(3, test_img, test_target, sixty_x, sixty_y)
# %%time
skl_knn(5, test_img, test_target, sixty_x, sixty_y)
# %%time
cos_knn(3, test_img, test_target, fifty_x, fifty_y)
# %%time
skl_knn(5, test_img, test_target, fifty_x, fifty_y)
# %%time
cos_knn(3, test_img, test_target, fourty_x, fourty_y)
# %%time
skl_knn(5, test_img, test_target, fourty_x, fourty_y)
# %%time
cos_knn(3, test_img, test_target, thirty_x, thirty_y)
# %%time
skl_knn(5, test_img, test_target, thirty_x, thirty_y)
# %%time
cos_knn(3, test_img, test_target, twenty_x, twenty_y)
# %%time
skl_knn(5, test_img, test_target, twenty_x, twenty_y)
# %%time
cos_knn(3, test_img, test_target, ten_x, ten_y)
# %%time
skl_knn(5, test_img, test_target, ten_x, ten_y)
# %%time
cos_knn(3, test_img, test_target, one_x, one_y)
# %%time
skl_knn(5, test_img, test_target, one_x, one_y)
# The test results show the following:
# * The Cosine-Similarity Model either matched the Scikit-Learn K-NN accuracy wise, or beat it by 1%-2%.
# * As far as classification speed goes, the Cosine-Similarity Model tends to be between 1.5-2 times faster than the Scikit-Learn K-NN.
# * Strangely, the Cosine-Similarity Classifier tends to underperform when classifying the digit 9. This could be on account of the fact that 4 and 9 are so similar, and thus tend to muddle the similarity metric for classification. It is possible that the cosine similarity metric performs better on data with higher variance, where as other similarity metrics perform better on data with lower variance.
#
# Below, the Cosine-Similarity Model and Scikit-Learn K-NN are tested in the time taken to classify a single image.
# %%time
cos_knn(3, t_img, t_target, sixty_x, sixty_y)
# %%time
skl_knn(5, t_img, t_target, sixty_x, sixty_y)
# As shown by the test results, the Cosine-Similarity Model is significantly faster (about 45 times faster in this example) at classifying single points than its corresponding Scikit-Learn classifier for larger stored data sets.
#
#
# ## Conclusion
# Although I cannot speak as to why the Cosine-Similarity Model is generally more accurate than the Scikit-Learn K-NN model, it is likely faster because the Scikit-Learn model is far more complex. The Scikit-Learn model takes many more arguments, adding complexity to the classification task, so it takes more time.
#
# The main lesson learned from this project is, depending on the problem being solved, it may be best to make an algorithm in-house instead of using pre-built algorithms, as these pre-built algorithms are meant to work well across a wide variety of situations, and are not optimized for just one problem.
#
#
# ### Room For Improvement
# Below are some points that can be used in the improvement of the Cosine-Similarity Model.
#
# The current Cosine-Similarity Model uses a brute force approach to classifying input data, where each point to be classified has its distance computed with each other point in the model's stored data set. This current brute force approach scales as $O(DN^2)$ for $D$ dimensions and $N$ samples. To address this massive computational inefficiency, one of the many tree based data structures could be implemented into the model, greatly reducing the number of distance calculations made for each data point, furthur reducing classification complexity to $O[DN\text{log}(N)]$.
#
# In the current Cosine-Similarity Model, each of the $k$ points used for voting are equally weighted. While the optimal voting approach used for classification varies from problem to problem, it would be worth experimenting with other weighted voting methods. Using cross validation to find the optimal $k$/weight "setting" for each problem would be worth the time.
#
#
# ### Lessons Learned & Moving Forward
# Below is a list of the primary takeaways from the project, and possible future applications of the model.
# * One of the greatest characteristics about K-NN is its simplicity. The Cosine-Similarity Model, an incredibly simple model, out performs some standard convolutional neural networks accuracy wise. This demonstrates the beauty of simple, parsimonious models, that seem to have been largely overlooked lately with the incredibly performance of deep learning.
# * Cosine similarity is often not used as it tends to violate the triangle inequality and does not work on negative data. However, as this analysis proves, cosine similarity holds its own in certain problems, as it is generally both accurate and efficient.
# * The results from this model are by no means state of the art, however they do show that proprietary algorithms often perform quite well on specific problems.
# * I doubt the Cosine-Similarity Model will generalize well to more complex image classification tasks. However, using techniques such as max pooling in more complex image classification tasks may help improve the model's accuracy.
# * In future projects, I would like to incorporate weighted voting, tree data structures, and cross-validation into the development of the model.
#
#
# #### References
# * [Scikit-Learn Documentation](http://scikit-learn.org/stable/modules/neighbors.html)
| Cosine-Similarity Model Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Analysis and Machine Learning Applications for Physicists
# *Material for a* [*University of Illinois*](http://illinois.edu) *course offered by the* [*Physics Department*](https://physics.illinois.edu). *This content is maintained on* [*GitHub*](https://github.com/illinois-mla) *and is distributed under a* [*BSD3 license*](https://opensource.org/licenses/BSD-3-Clause).
#
# [Table of contents](Contents.ipynb)
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns; sns.set()
import numpy as np
import pandas as pd
import warnings
warnings.filterwarnings('ignore')
from mls import plot_KL, plot_ELBO
# ## Variational Inference
# Most Bayesian inference problems cannot be solved exactly, so require an approximate method. The MCMC method is one such method, invented in the 1950s. **Variational inference (VI)** is an alternative approximate method, invented in the 1990s:
# - MCMC: provides an approximate description of the exact posterior distribution (using sampling).
# - VI: provides an exact description of an approximate posterior distribution (using optimization).
#
# The underlying assumptions and numerical algorithms involved (sampling and optimization) are fundamentally different, leading to different tradeoffs between these methods.
#
# The essence of VI is to first define a family of PDFs that balance two competing criteria:
# - convenient for calculations, and
# - flexible enough to approximately match some unknown target PDF.
#
# We then select the family member that is "closest" to the target. In a Bayesian context, our target PDF is a posterior distribution, but VI is a more general technique for finding approximate PDFs.
# ### Kullback-Leibler Divergence
# Variational inference relies on a concept of "closeness" between two PDFs, which we call $q(\theta)$ and $p(\theta)$. Note that we are talking about "separation" in an abstract function space, rather than a coordinate space. Just as with coordinate separation, there are many possible valid definitions, e.g.
#
# $$ \Large
# (\sum_i (x_i - y_i)^2)^{1/2} \quad, \quad
# \sum_i |x_i - y_i| \quad, \quad
# \max_i\, |x_i - y_i| \quad, \ldots
# $$
#
# VI uses the [Kullback Leibler (KL) divergence](https://en.wikipedia.org/wiki/Kullback%E2%80%93Leibler_divergence) to measure the "closeness" of PDFs $q(\theta)$ and $p(\theta)$:
#
# $$ \Large
# \boxed{
# \text{KL}( q \parallel p ) \equiv \int d\theta\, q(\theta)\, \log\frac{q(\theta)}{p(\theta)}} \; .
# $$
#
# Since $q$ is a PDF, KL divergence can also be written as a difference of expectation values over $q$:
#
# $$ \Large
# \text{KL}( q \parallel p ) = \langle \log q(\theta)\rangle_q - \langle \log p(\theta)\rangle_q \; .
# $$
# + [markdown] solution2="hidden" solution2_first=true
# **EXERCISE:**
# - Is KL divergence symmetric, $\text{KL}(q\parallel p) = \text{KL}(p\parallel q)$?
# - What is the value of $\text{KL}(q\parallel p)$ when $p = q$?
# - What happens to the integrand when either $q(\theta)$ or $p(\theta)$ approaches zero?
# - What bounds, if any, can you place on the value of $\text{KL}(q\parallel p)$ given that $p$ and $q$ are PDFs?
# + [markdown] solution2="hidden"
# KL divergence is not symmetric since exchanging $q$ and $p$ in the integrand changes its value. This makes KL divergence an unusual measure of separation and means that it is not a true
# [metric](https://en.wikipedia.org/wiki/Metric_%28mathematics%29).
#
# When $p=q$, the log zeros the integrand (except possibly where $q$ has a singularity), resulting in a KL divergence of zero. This is what we would expect for a useful measure of separation.
#
# When $q \rightarrow 0$ the combination $q \log q \rightarrow 0$. When $p(\theta)\rightarrow 0$, the log term diverges $\log(1/p)\rightarrow +\infty$. As a result, the KL integrand blows up wherever $\theta$ is very unlikely according to $p$ doesn't care when $\theta$ is very unlikely according to $q$.
#
# A PDF is always $\ge 0$ but not bounded from above, so the KL divergence is not bounded from above. However, nothing prevents $q(\theta) < p(\theta)$, so the integrand can be negative (due to the log) even with $p, q \ge 0$.
#
# It turns out that the KL divergence is always $\ge 0$ but this is not obvious. The proof relies on the [log sum inequality](https://en.wikipedia.org/wiki/Log_sum_inequality), which in turns relies on [Jensen's inequality](https://en.wikipedia.org/wiki/Jensen's_inequality) which we met earlier.
#
# The key insight is that the KL divergence is [convex](https://en.wikipedia.org/wiki/Convex_function) in $q$:
#
# $$ \Large
# \text{KL}\big(\lambda q_1 + (1-\lambda) q_2\parallel p\big) \le
# \lambda\,\text{KL}(q_1\parallel p) + (1-\lambda)\,\text{KL}(q_2\parallel p) \; .
# $$
#
# since, for any value of $\theta$, $p$ and $q$ are just numbers and the integrand
#
# $$ \Large
# f(q) = q \log q/p
# $$
#
# is a convex function of $q$:
# + solution2="hidden"
def plot_convex():
q = np.linspace(0, 2.5, 100)[1:]
f = lambda q, p: q * np.log(q / p)
qlo, qhi = 0.5, 2
for p in (0.5, 1, 2):
plt.plot(q, f(q, p), label='$p={:.1f}$'.format(p))
plt.plot([qlo, qhi], [f(qlo,p), f(qhi,p)], 'k:')
plt.legend()
plt.xlabel('$q$')
plt.ylabel('$f(q) = q\, log (q/p)$')
plot_convex()
# + [markdown] solution2="hidden"
# ---
# -
# We will use the MLS `plot_KL` function to explore some examples:
help(plot_KL)
# For our first example, we chose a family $q$ that includes the target $p$. This is generally not feasible but nicely demonstrates our earlier claim that KL$\ge 0$ with KL$=0$ when $q=p$.
#
# In this example, we explore the family of PDFs $q(\theta; s)$ by varying the scale factor $s$. More generally, the family can be explored with any (multidimensional) parameterization that is convenient for calculations. We need a parameterization of the family $q$ in order to use standard optimization algorithms to find the minimum KL divergence.
plot_KL(q='norm', q_scale_range=[0.75, 1.3], p='norm', p_scale=1.0, theta_range=[-5, +5])
# Note how (in the bottom left panel) the KL integrand has positive and negative regions: the net area is always positive, however, since KL$\ge 0$.
#
# For our next example, we consider the more realistic case where the family $q$ does not include the target $p$ so we have to settle for the "closest" approximation, according to the KL divergence:
plot_KL(q='norm', q_scale_range=[0.8, 1.2], p='laplace', p_scale=0.8, theta_range=[-5, +5])
# Notice how the "closest" $q$ now has KL$>0$. It also avoids regions where $p \simeq 0$, since that would blow up the KL integrand.
#
# The example above showed that a Gaussian PDF with $s \simeq 1$ gives the best match a Laplacian PDF with $s = 0.8$. Now, turn this around and find the closest Laplacian $q$ to a Gaussian $p$ with $s = 1$:
plot_KL(q='laplace', q_scale_range=[0.6, 0.9], p='norm', p_scale=1.0, theta_range=[-5, +5])
# The answer is a Laplacian with $s\simeq 0.74$, rather than $0.8$, so a round trip does not end up back where we started! However, this shouldn't surprise us because the KL divergence is not symmetric in $q$ and $p$.
# Note that the KL divergences between Gaussian and Laplacian distributions in the examples above can all be calculated analytically, which is useful for testing but not generally true. The analytic results are summarized below, for reference:
#
# $$
# \begin{aligned}
# \text{KL}_{qp}[G(s_1)\parallel G(s_2)] &=
# \frac{1}{2} r^2 - \log r - \frac{1}{2} \quad& \quad
# \text{KL}_{qp}[G(s_1)\parallel L(s_2)] &=
# \sqrt{\frac{2}{\pi}} r - \log\left( \sqrt{\frac{\pi}{2}} r\right) - \frac{1}{2} \\
# \text{KL}_{qp}[L(s_1)\parallel G(s_2)] &=
# r^2 - \log\left(\sqrt{\frac{2}{\pi}}r\right) - 1 \quad& \quad
# \text{KL}_{qp}[L(s_1)\parallel L(s_2)] &=
# r - \log r - 1 \; ,
# \end{aligned}
# $$
#
# where $r \equiv s_1/s_2$ is the ratio of scale parameters. With $s_2$ fixed, the corresponding minimum KL divergences are:
#
# $$
# \begin{aligned}
# \min \text{KL}_{qp}[G(s_1=s_2)\parallel G(s_2)] &= 0 \quad& \quad
# \min \text{KL}_{qp}[G(s_1=(\pi/2)s_2)\parallel L(s_2)] &=
# \sqrt{\frac{2}{\pi}} - \frac{1}{2}\log\frac{\pi}{2} - \frac{1}{2} \\
# \min \text{KL}_{qp}[L(s_1=s_2/\sqrt{2})\parallel G(s_2)] &=
# \frac{1}{2}\log\frac{\pi}{2} \quad& \quad
# \min \text{KL}_{qp}[L(s_1=s_2)\parallel L(s_2)] &= 0 \; .
# \end{aligned}
# $$
# ### Evidence Lower Bound
# The KL divergence is a generic method to find the parameterized PDF $q(x,s)$ that "best" approximates some target PDF $p(x)$. For Bayesian inference, the $p$ we care about is the posterior:
#
# $$ \Large
# p(\theta) = P(\theta\mid D) = \frac{P(D\mid \theta)\, P(\theta)}{P(D)} \; ,
# $$
#
# where:
# - $\theta$ are the model parameters and $D$ represents the observed data.
# - $P(D\mid \theta)$ is the likelihood of the data assuming parameters $\theta$.
# - $P(\theta)$ is our prior on the parameters.
# - $P(D)$ is the "evidence".
#
# Since we generally cannot calculate the evidence $P(D)$, a useful inference method should not require that we know its value.
#
# The **variational Bayesian inference** method has three steps:
# - Define a family of PDFs $q(\theta; s)$ that approximate the true posterior $P(\theta\mid D)$.
# - Use optimization to find the value $s^\ast$ that, according to the KL divergence, best approximates the true posterior.
# - Use $q(\theta; s=s^\ast)$ as an approximation of the true posterior for calculating expectation values, etc.
#
# The main tradeoff is in picking the approximate PDF family $q$. A more flexible choice will generally do a better job of approximating the true posterior, but also require more difficult calculations.
#
# Plugging the posterior into the KL definition, we can rewrite:
#
# $$ \Large
# \begin{aligned}
# \text{KL}(q\parallel p) &= \int d\theta\, q(\theta) \log\left[
# \frac{P(D)\, q(\theta)}{P(D\mid \theta)\, P(\theta)}
# \right] \\
# &= \int d\theta\, q(\theta) \left[\log P(D) +
# \log\frac{q(\theta)}{P(\theta)} - \log P(D\mid\theta) \right] \\
# &= \log P(D) + \text{KL}(q\parallel P) - \int d\theta\, q(\theta) \log P(D\mid\theta) \; .
# \end{aligned}
# $$
#
# The three terms on the right-hand side are:
# - The log of the evidence $P(D)$.
# - The KL divergence of $q(\theta)$ with respect to the prior $P(\theta)$.
# - The $q$-weighted log-likelihood of the data.
# + [markdown] solution2="hidden" solution2_first=true
# **DISCUSS:** Describe the $q(\theta)$ that would minimize the contribution of each term to their sum (assuming a fixed dataset $D$).
# + [markdown] solution2="hidden"
# Solution:
# - The log of the evidence is a constant offset in the sum, independent of $q$.
# - The KL divergence term is minimized when $q(\theta) = P(\theta)$, i.e., it drives $q$ to look like the prior.
# - The log-likelihood term is minimized when $q(\theta)$ prefers parameters $\theta$ that explain the data.
#
# The competition between the last two terms is exactly what we need for a useful learning rule that balances prior knowledge with the information gained from new data.
#
# ---
# -
# We can rewrite the expression above in terms of the log-evidence as:
#
# $$ \Large
# \log P(D) = \int d\theta\, q(\theta) \log P(D\mid\theta) - \text{KL}(q\parallel P) + \text{KL}(q\parallel p) \; .
# $$
#
# Since the last term is $\ge 0$ (since any KL$\ge 0$), we find:
#
# $$ \Large
# \log P(D) \ge \int d\theta\, q(\theta) \log P(D\mid\theta) - \text{KL}(q\parallel P) \; ,
# $$
#
# and call this right-hand side the **evidence lower bound (ELBO)**:
#
# $$ \Large
# \text{ELBO}(q) \equiv \int d\theta\, q(\theta) \log P(D\mid\theta) - \text{KL}(q\parallel P) \; .
# $$
#
# Substituting above, we find that
#
# $$ \Large
# \text{KL}(q\parallel p) = \log P(D) - \text{ELBO}(q) \; ,
# $$
#
# so that the ELBO contains all of the $q$ dependence of the KL divergence of $q$ with respect to $p$. The crucial insights are that:
# - Minimizing $-\text{ELBO}(q)$ with respect to $q$ is equivalent to minimizing $\text{KL}(q\parallel p)$.
# - $\text{ELBO}(q)$ is much easier to calculate since it does not depend on the evidence $P(D)$.
#
# Note that, as with the KL divergence, the ELBO can be evaluated in terms of expectation values,
#
# $$ \Large
# \text{ELBO}(q) = \langle \log P(D\mid\theta)\rangle_q + \langle \log P(\theta)\rangle_q - \langle \log q\rangle_q \; .
# $$
#
# The practical significance of this fact is that we can estimate the ELBO using averages of known quantities calculated with (finite) samples drawn from $q$, which effectively uses Monte Carlo integration with [importance sampling](https://en.wikipedia.org/wiki/Importance_sampling).
# We will use the MLS `plot_ELBO` function to explore some examples:
help(plot_ELBO)
# The example below specifies a [Laplacian PDF](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.laplace.html) for observing $x$ with an unknown offset parameter $\theta$,
#
# $$ \Large
# P(x\mid \theta) = \frac{1}{2}\, e^{-|x - \theta|} \; .
# $$
#
# The resulting likelihood is then:
#
# $$ \Large
# P(D\mid\theta) = \prod_i P(x_i\mid\theta) \; .
# $$
#
# Our prior knowledge of $\theta$ is specified by a unit Gaussian,
#
# $$ \Large
# P(\theta) = (2\pi)^{-1/2}\, e^{-\theta^2/2} \; .
# $$
#
# The resulting posterior PDF
#
# $$ \Large
# P(\theta\mid D) = \frac{P(D\mid\theta)\, P(\theta)}{P(D)}
# $$
#
# is no longer a simple distribution since it depends on the random data $D$ and reflects its statistical fluctuations. However, as shown below, it is roughly Gaussian, so we use a family $q$ of Gaussians to approximate it.
plot_ELBO(q='norm', q_scale_range=[0.05, 0.15], likelihood='laplace', prior='norm',
theta_range=[-0.6, +0.6], n_data=100)
# + [markdown] solution2="hidden" solution2_first=true
# **DISCUSS:**
# - Is the offset between the KL divergence and -ELBO significant on the scale of variations shown in the upper-right panel?
# - Is the posterior dominated by the prior or the new data in this example?
# - How do you expect these plots to change when doubling the number of data samples? (think about it before trying it).
# + [markdown] solution2="hidden"
# The offset equals $\log P(D) \simeq -145$ so is very significant compared with the variations $\simeq 0.3$ shown in the upper-right panel. However, since the offset is constant (with respect to $s$) it does not affect the location of the minimum.
#
# Referring to the top-left panel, the posterior has a standard deviation $s\simeq 0.1$ but but prior is much wider with $s = 1$, so the posterior is dominated by the new data.
#
# Doubling the number of data samples will make the data even more informative, leading to a narrower posterior. The best-fitting $q$ will therefore also be narrow, leading to a minimum KL divergence (upper-right panel) at a lower value of $s$.
#
# Re-run the example above with `n_data` changed from 100 to 200 to confirm these predictions:
# + solution2="hidden"
plot_ELBO(q='norm', q_scale_range=[0.05, 0.15], likelihood='laplace', prior='norm',
theta_range=[-0.6, +0.6], n_data=200)
# -
# ## Practical Calculations with VI
# MCMC with Metroplis-Hastings updates can be used as a black box for an arbitrary inference problem that only requires that you can calculate your likelihood $P(D\mid \theta)$ and prior $P(\theta)$ for arbitrary parameter values $\theta$.
#
# VI, on the other hand, generally requires more work to setup for a particular problem, but is then often more computationally efficient since it replaces sampling with optimization. A necessary step in any VI inference is to select an approximating family $q$, and this generally requires knowledge of the particular problem and some judgment on how to tradeoff calculational convenience against approximation error.
#
# Once you selected a family $q(\theta; s)$ that is explored by some $s$, you need to be able to:
# - evaluate the KL divergence of $q(s)$ with respect to $p$ for any $s$, and
# - find the value of $s$ that minimizes the KL divergence.
#
# There are standard numerical optimization methods for the second step, which perform best when you can evaluate derivatives of $q(s)$ with respect to $s$. The first step either requires an analytic integral over $\theta$ or a sufficiently accurate numerical approximation to the KL integral. A [recent development](https://arxiv.org/abs/1401.0118) is to use the expectation form of the KL divergence,
#
# $$ \Large
# \text{KL}( q \parallel p ) = \langle \log q(\theta)\rangle_q - \langle \log p(\theta)\rangle_q \; ,
# $$
#
# to replace the integral with averages. This approach changes our requirements on the family $q$ from being able to do integrals involving $q$ to being able to sample from $q$, which is generally much easier. Although this method is known as "Black Box Variational Inference", it still lacks the turn-key convenience of MCMC with MH updates.
#
# The examples above used a single parameter $\theta$, to simplify plotting and allow straightforward numerical integration. More interesting problems generally have many parameters, which makes picking a suitable family $q$ much harder. A common approach, known as the **mean field approximation**, is to assume that the posterior can be factored:
#
# $$ \Large
# P(\theta_1, \theta_2, \ldots\mid D) = p_1(\theta_1)\, p_2(\theta_2) \ldots
# $$
#
# This is certainly not true in general, but does break a difficult multidimensional optimization problem into a sequence of simpler 1D optimization problems, so is sometimes necessary. Note that this approach is not able to capture any correlations between $\theta_i$ and $\theta_j$, so is not a good choice when correlations are expected to be important.
| notebooks/Variational.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from numpy import array
import pandas as pd
from fox_toolbox.utils import xml_parser
from tsr import csv_parser, tsr
import matplotlib.pyplot as plt
# %load_ext autoreload
# %autoreload 2
# -
# ### Read IRSM FORM
# +
irsmform = xml_parser.get_files('irsmform xml', folder = 'linear TSR logs')
irsmout = xml_parser.get_files('out xml', folder = 'linear TSR logs')
csv = xml_parser.get_files('CMS 10y csv', folder = 'linear TSR logs')
replic_basket = csv_parser.parse_csv(csv)
cal_basket = list(xml_parser.get_calib_basket(irsmform))
settings = xml_parser.get_model_settings(irsmform)
# +
main_curve, sprds = xml_parser.get_rate_curves(irsmform)
dsc_curve = main_curve
try:
estim_curve = sprds[0]
except TypeError:
estim_curve = main_curve
n = settings.SpotIterations
# -
dsc_adj_cms_flows = []
for swo in cal_basket:
pmnt_date = swo.payment_dates[0]
mr = xml_parser.get_tsr_params(irsmform).meanRevTSRSwapRate(swo.start_date)
adj_cms_flow = tsr.cmsflow(swo, dsc_curve, estim_curve, n, mr, pmnt_date).adjCMSrate
dsc_adj_cms_flows.append(cmsflow * dsc_curve(pmnt_date))
adj_cms_flows = array(adj_cms_flows)
adj_cms_flows
sum(adj_cms_flows)
swo.strike
swo.strike
n = 100_000_000
1639019.212092/ (swo.strike * n)
1652490/1639019 - 1
_30 * 365
| notebooks/gold standard/.ipynb_checkpoints/3. recon price-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # `fitgrid` EEG modeling
#
# 1. Read or construct a 2-D pandas.DataFrame() of time-stamped epochs with time series observations in rows and variables in columns.
# +
import pandas as pd
import fitgrid as fg
from fitgrid import sample_data, DATA_DIR
# fitgrid built-in sample data download
p3_f = "sub000p3.ms1500.epochs.feather"
sample_data.get_file(p3_f)
# read as a pandas.DataFrame
p3_df = pd.read_feather(DATA_DIR / p3_f)
p3_df["time_ms"] = p3_df["match_time"]
# EEG quality control (specific to these data)
good_epoch_ids = p3_df.query("time_ms == 0 and log_flags == 0")["epoch_id"]
p3_df = p3_df.query("epoch_id in @good_epoch_ids")
# select EEG channels ("left hand side") and predictor variables ("right hand side")
columns_of_interest = [
"epoch_id",
"time_ms", # index columns
"MiPf",
"MiCe",
"MiPa",
"MiOc", # EEG
"stim",
"tone", # predictors
]
p3_df = p3_df[columns_of_interest].query("stim in ['standard', 'target']")
p3_df
# -
# 2. Load the epochs data into fitgrid for modeling
p3_df.set_index(['epoch_id', 'time_ms'], inplace=True)
p3_epochs_fg = fg.epochs_from_dataframe(
p3_df,
epoch_id='epoch_id',
time='time_ms',
channels=['MiPf', 'MiCe', 'MiPa', 'MiOc'],
)
# 3. Fit a model (formula) to the observations at each timepoint and channel.
lm_grid = fg.lm(
p3_epochs_fg,
RHS="1 + stim",
LHS=["MiPf", "MiCe", "MiPa", "MiOc"]
)
# 4. The `FitGrid[time, channel]` object is a container for the model fits.
lm_grid
# Slice it like a dataframe by times and/or channelsl
lm_grid[-200:600, ["MiCe", "MiPa"]]
# Access attributes by name like a single fit.
# The results come back as a :py:class:`pandas.DataFrame` or another
# FitGrid[time, channel].
# estimated predictor coefficients (betas)
lm_grid.params
# coefficient standard errors
lm_grid.bse
# model log likelihood.
lm_grid.llf
# Plot results with a fitgrid built-in.
f, axs = lm_grid.plot_betas()
# Or make your own with pandas, matplotlib, seaborn, etc..
# +
from matplotlib import pyplot as plt
# Slice a time range and compute means with pandas
p3_effect = lm_grid.params.loc[
pd.IndexSlice[250:400, "stim[T.target]"], :
].mean()
ax = p3_effect.plot.bar()
ax.set_title("Treatment effect: Mean amplitude 250 to 400 ms")
_ = ax.set(ylabel="$\mu$V")
# -
# 5. Compare grid summaries for models sets and pairs.
# +
from fitgrid.utils import summary as fgs
p3_summaries = fgs.summarize(
p3_epochs_fg,
modeler="lm",
RHS=["1 + stim + tone", "1 + stim", "1 + tone", "1"],
LHS=["MiPf", "MiCe", "MiPa", "MiOc"],
)
p3_summaries
# -
# Compare models on Akiake's information criterion (AIC) as
# the difference between the model's AIC and the
# lowest in the set. Larger AIC differences indicate relatively
# less support for the model in comparison with the alternative(s).
fig, axs = fgs.plot_AICmin_deltas(p3_summaries)
for axi in axs:
axi[0].set(ylim=(0, 30))
axs[-1][0].set(xlabel="Time (ms)", ylabel="$\mathsf{AIC - AIC_{min}}$")
fig.tight_layout()
| notebooks/Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
initial_value = 'hello'
for c in initial_value:
print(ord(c))
ord('a')
ord('z')
chr(97)
def caesar_shift(message, shift=0):
if shift > 25:
raise AssertionError("Shift cannot be more than 25")
min_ord, max_ord = ord('a'), ord('z')
message = message.lower()
result = []
for c in message:
ord_c = ord(c)
if ord_c < min_ord or ord_c > max_ord:
result.append(c)
continue
shifted_c = ord_c + shift
if shifted_c > max_ord:
shifted_c = (shifted_c - max_ord) + (min_ord - 1)
result.append(chr(shifted_c))
return ''.join(result)
caesar_shift('abc', 0)
caesar_shift('abc', 1)
caesar_shift('xyz', 1)
caesar_shift('ab cd', 5)
ord(' ')
caesar_shift('A quick brown fox jump over a lazy dog.', 6)
| Python/Problems/CaesarCipher.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## A first neural network in MXNet
# ### General imports
% matplotlib inline
import pickle
import matplotlib.pyplot as plt
import numpy as np
import random
import time
# ### First step is to load data from file and plot a few examples
train_x = pickle.load(open("MNIST_train_x.pkl", 'rb'))
train_y = pickle.load(open("MNIST_train_y.pkl", 'rb'))
test_x = pickle.load(open("MNIST_test_x.pkl", 'rb'))
test_y = pickle.load(open("MNIST_test_y.pkl", 'rb'))
print("Data stats")
print(type(train_x))
print(train_x.shape)
print(type(train_y))
print(train_y.shape)
print(type(test_x))
print(test_x.shape)
print(type(test_y))
print(test_y.shape)
# ### Visualization function
# - It plots a grid of 2 x 4 examples, with the real and predicted values
def plotExamples(data, labels, model_predict):
plt.figure(figsize=(8,5))
for i in range(8):
sub = 241 + i
ax = plt.subplot(sub)
index = np.random.randint(0, data.shape[0])
ax.set_title("num: " + str(np.argmax(labels[index])) + "," + str(np.argmax(model_predict[index])))
im = np.reshape(data[index], (28, 28))
plt.imshow(im, cmap="gray")
plt.show()
'''
Currently we are just duplicating the correct labels
When we have a model we can plot both the correct and predicted labels
'''
plotExamples(train_x, train_y, train_y)
# ### Building a first MXNet model
# - Simple 3 layer feedforward network
# - One input, one hidden, and one output layer
# - Six steps:
# 1. Preparing the data
# 2. Build model
# 3. Initialize parameters
# 4. Prepare loss and optimizer
# 5. Prepare training and evaluation code loops
# 6. Train the model
# - Please note, this section on how to set up an MXNet model closely tracks the tutorial available as part of the Gluon documentation, available [here](http://thestraightdope.mxnet.io/P03-C02-mlp-gluon.html)
# ### MXNet imports
import mxnet as mx
from mxnet import nd, autograd
from mxnet import gluon
# ### 1. Preparing the data
# - We will use MXNet's NDArray to store our data. This is the core data structure to hold data for computation in MXNet.
# - NDArray is similar to numpy.ndarray but also supports execution on different types of hardware (CPU, GPU for example) and can easily by parallelized
# - We can also use it to build dataIterators, a useful structure which makes it easy to organize your data into batches to feed into a model for training and evaluation
# - For more information about NDArrays see [this tutorial](http://mxnet.io/tutorials/basic/ndarray.html)
''' NDArrays can be created from numpy arrays '''
train_x_mx = mx.nd.array(train_x)
train_y_mx = mx.nd.array(train_y)
test_x_mx = mx.nd.array(test_x)
test_y_mx = mx.nd.array(test_y)
'''NDArrays have a shape, data type and context (processor that the computation will be executed on)'''
print("NDArray attributes")
print(train_x_mx.shape)
print(train_x_mx.dtype)
print(train_x_mx.context)
print()
'''The two different representations have different types'''
print("Numpy data type")
print(type(train_x_mx))
print("NDArray data type")
print(type(train_x_mx.asnumpy()))
print()
'''
Each type can be printed directly
Also note that NDArrays have similar indexing operators to numpy
'''
print("Numpy format")
print(train_x[0,150:160])
print("NDArray format")
print(train_x_mx[0,150:160])
''' Or convert from an NDArray back to numpy '''
print("NDArray converted to numpy")
print(train_x_mx[0,150:160].asnumpy())
print()
''' Printing statistics of the data in the NDArray format '''
print("Training data")
print(train_x_mx.shape)
print(train_x_mx.dtype)
print("Training labels")
print(train_y_mx.shape)
print(train_y_mx.dtype)
print("Test data")
print(test_x_mx.shape)
print(test_x_mx.dtype)
print("Test labels")
print(test_y_mx.shape)
print(test_y_mx.dtype)
''' It is easy to change data types as needed '''
test_y_mx = test_y_mx.astype('int32')
print(test_y_mx.dtype)
test_y_mx = test_y_mx.astype('float32')
print(test_y_mx.dtype)
'''
Create an iterator for the training and test data
This packages the data and labels into individual batches to be fed into the network
'''
batch_size = 32
train_data = mx.io.NDArrayIter(train_x_mx, train_y_mx, batch_size, shuffle=True, \
last_batch_handle='pad', data_name='train_data', label_name='train_label')
test_data = mx.io.NDArrayIter(test_x_mx, test_y_mx, batch_size, shuffle=True, \
last_batch_handle='pad', data_name='test_data', label_name='test_label')
''' Iterators have a number of useful descriptive attributes '''
print(train_data)
print(train_data.provide_data)
print(train_data.provide_label)
print(test_data)
print(test_data.provide_data)
print(test_data.provide_label)
# ### 2. Build model
'''First step is to initialize your model'''
net = mx.gluon.nn.Sequential()
'''Then, define your model architecture'''
with net.name_scope():
'''
Add a fully connected hidden layer with 128 nodes and specify the activation function
The input layer is implicit
'''
net.add(mx.gluon.nn.Dense(128, activation="sigmoid"))
'''Add an output layer with 10 output nodes and specify the activation function'''
net.add(mx.gluon.nn.Dense(10, activation="sigmoid"))
# ### 3. Initialize parameters
# +
'''
Lists information about the parameters in the network
Placeholder of 0 for the dimension of the input data.
In this example it will be 784 and is inferred when data is fed through the network
'''
print(net.collect_params())
'''
All parameters need a context - the device on which they carry out the computations
In this example computation is carried out on cpu 0
'''
ctx = mx.cpu()
print("Context for calculations: {}".format(ctx))
'''
Initialize all of the network's weights and biases to
Assign all of the networks parameters to the context - cpu 0
'''
net.collect_params().initialize(mx.init.Xavier(magnitude=2.24), ctx=ctx)
# -
# ### 4. Prepare the loss and optimizer
'''Next, define your loss function and optimizer through MXNet's Trainer class
1. Loss function - how your model defines the error between
the correct output and its prediction
2. Optimizer (part of the Trainer) - how your model learns.
'''
loss = gluon.loss.L2Loss()
trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': .01})
# ### 5. Prepare training and evaluation code
'''
The dataIterator structure packages data nicely and has a number of interesting features
The code below demonstrates some of the features of a single batch of data
'''
train_data.reset()
for i, batch in enumerate(train_data):
print("Batch object type: {}".format(type(batch)))
print(batch)
print()
print("Batch.data is a list of length 1 containing the data for this batch")
print("Batch data type {}".format(type(batch.data)))
print("Batch data list length: {}".format(len(batch.data)))
print("Batch data example values")
print(batch.data[0][0,150:160])
print()
print("Batch.label is a list of length 1 containing the label for this batch")
print("Batch label type {}".format(type(batch.label)))
print("Batch label list length: {}".format(len(batch.label)))
print("Batch label example values")
print(batch.label[0][:5])
print()
data = batch.data[0]
print("Data context: {}, shape: {}, data type: {}".format(data.context, data.shape, data.dtype))
label = batch.label[0]
print("Label context: {}, shape: {}, data type: {}".format(label.context, label.shape, label.dtype))
break
# +
''' Next step is to define a performance measure'''
''' The typical measure for this problem is overall accuracy
- the percentage of examples for which the model predicted the correct class
For more challenging classification tasks, the classic ImageNet challenge for example,
which contains images from 1000 classes it is common to look at a top k accuracy,
the percentage of examples for which the model predicted the correct class
in the the top k results
This also demonstrates MXNet's CompositeEvalMetric,
which allows you to evaluate models using multiple metrics
'''
''' Helper function creating the composite metric'''
def create_metrics():
m_acc = mx.metric.Accuracy()
m_acc_topk = mx.metric.TopKAccuracy(top_k=3, name='top_3_accuracy')
metrics = mx.metric.CompositeEvalMetric()
metrics.add(m_acc)
metrics.add(m_acc_topk)
return metrics
''' Helper function converting one hot labels to a single number'''
def convert_from_one_hot(label):
label_np = label.asnumpy()
label_np = label_np.argmax(axis=1)
label_np = np.expand_dims(label_np, axis=1)
label_np = mx.nd.array(label_np)
return label_np
''' This helper function takes some data and a network and calculates the performance of that
network using the metrics defined above
'''
def evaluate_accuracy(data_iterator, net, metrics):
data_iterator.reset()
''' Iterate through the data'''
for i, batch in enumerate(data_iterator):
with autograd.record():
data = batch.data[0]
label_one_hot = batch.label[0]
'''
Labels need to have a single value per example.
So we need to convert from a one hot encoding.
e.g. from [0, 0, 0, 1, 0, 0, 0, 0, 0, 0] --> [3]
'''
label = convert_from_one_hot(label_one_hot)
''' Forward pass '''
output = net(data)
metrics.update([label], [output])
'''
Metrics.get() returns a tuple containing two lists.
The first list contains the names of the evaluation metrics
The second contains a list of the values
e.g. (['accuracy', top_3_accuracy'],[0.75, 1.0])
'''
return metrics.get()[1]
# -
# ### 6. Train the model
def train_model(net, train_data, test_data, metrics, loss, trainer, batch_limit=0, num_epochs=10, verbose=1):
'''
net: neural network
train_data: training data
test_data: test data
metrics: instance of mx.metric class defining evaluation metrics for the network
loss: instance of gluon.loss class defining the loss function - the measure of error
trainer: instance of gluon.Trainer class defining an optimizer over parameters
batch_limit: number of batches to limit training to.
Can be useful to limit data for quick experiments
A zero value indicates no limit
num_epochs: number of iterations of the data to train on
verbose: Three values - 1, 2, or 3
Controls how much information to print
1 = least verbose, 3 = most verbose
'''
''' Lists to hold training statistics '''
losses = []
train_accuracies = []
test_accuracies = []
for e in range(num_epochs):
''' Keep track of how long an epoch takes '''
start = time.time()
epoch_loss = []
train_data.reset()
num_batches = 0
'''Iterate through the training data'''
for i, batch in enumerate(train_data):
data = batch.data[0]
label_one_hot = batch.label[0]
'''
Labels for the SoftmaxCrossEntropy loss need to have a single value per example.
So we need to convert from a one hot encoding.
e.g. from [0, 0, 0, 1, 0, 0, 0, 0, 0, 0] --> [3]
'''
label = convert_from_one_hot(label_one_hot)
with autograd.record():
'''Forward pass'''
output = net(data)
'''Calculate loss for this batch'''
try:
'''format for mse loss'''
b_loss = loss(output, label_one_hot)
except:
'''format for softmax cross entropy'''
b_loss = loss(output, label)
'''Calculate gradient of the weights and biases with respect to the loss'''
b_loss.backward()
'''Update all of the weights and biases'''
trainer.step(batch_size=data.shape[0])
'''Keep track of average loss per batch'''
loss_avg = mx.nd.mean(b_loss).asnumpy()[0]
epoch_loss.append(loss_avg)
num_batches += 1
if batch_limit > 0 and i >= batch_limit - 1:
break
'''Select 10 random epochs to print'''
sample_losses = random.sample(epoch_loss, 10)
average_loss = sum(epoch_loss) / (len(epoch_loss) * 1.0)
test_accuracy = evaluate_accuracy(test_data, net, metrics)
train_accuracy = evaluate_accuracy(train_data, net, metrics)
'''Store results'''
losses.append(average_loss)
train_accuracies.append(train_accuracy)
test_accuracies.append(test_accuracy)
end = time.time()
'''Print stats depending on verbosity'''
print("Epoch {} Loss: {:.5f} Time: {:.1f}s Num batches: {}".format(e, average_loss, end - start, num_batches))
if verbose == 3:
print("Sample batch losses: {}".format(sample_losses))
if verbose == 2 or verbose == 3:
print("Train_acc {:.5f}, Train_top_3_accuracy {:.5f}".format(train_accuracy[0], train_accuracy[1]))
print("Test_acc {:.5f}, Test_top_3_accuracy {:.5f}".format(test_accuracy[0], test_accuracy[1]))
if verbose > 3 or verbose < 1:
print("Not a valid verbose option. Valid options 1, 2, 3")
print()
return (losses, train_accuracies, test_accuracies)
batch_limit = 625 # Approximately 1/3 of the data
epochs = 10
metrics = create_metrics()
''' Note the loss, net, and trainer were created in parts 2 and 4'''
results = train_model(net, train_data, test_data, metrics, loss, trainer, \
batch_limit=batch_limit, num_epochs=epochs, verbose=2)
# ### Quadratic cost (mean squared error) vs. categorical crossentropy loss functions
# - Categorical cross-entropy significantly speeds up training
# - Softmax output layers are the most appropriate for the MNIST problem since each image can only belong to one class and the softmax functions over a set of values converts the values to a proability distribution. The result in this example is a probability distribution across the 10 classes.
# - As the value of one output node increases, the value of one or more other output nodes must decrease
# - This is consistent with our intuition that as we become more confident and image belongs to one class, we reduce our confidence that an image belongs to other classes
# - In MXNet the softmax operation is combined with the categorical cross-entropy loss function
# #### Softmax output layer, categorical cross-entropy
# +
net = mx.gluon.nn.Sequential()
with net.name_scope():
net.add(mx.gluon.nn.Dense(128, activation="sigmoid"))
net.add(mx.gluon.nn.Dense(10))
ctx = mx.cpu()
net.collect_params().initialize(mx.init.Xavier(magnitude=2.24), ctx=ctx)
loss = gluon.loss.SoftmaxCrossEntropyLoss()
trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': .01})
batch_limit = 625
epochs = 10
metrics = create_metrics()
results = train_model(net, train_data, test_data, metrics, loss, trainer, \
batch_limit=batch_limit, num_epochs=epochs, verbose=2)
# -
# ## Rectified Linear Unit (ReLU) vs. Sigmoid
# ### Observations
# - ReLU needs a low learning rate for the network to learn anything
# - May perform worse than a sigmoid hidden layer for shallow networks
# +
''' Learning rate of 0.01 results in poor learning'''
net = mx.gluon.nn.Sequential()
with net.name_scope():
net.add(mx.gluon.nn.Dense(128, activation="relu"))
net.add(mx.gluon.nn.Dense(10))
ctx = mx.cpu()
net.collect_params().initialize(mx.init.Xavier(magnitude=2.24), ctx=ctx)
loss = gluon.loss.SoftmaxCrossEntropyLoss()
trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': .01})
batch_limit = 625
epochs = 5
metrics = create_metrics()
results = train_model(net, train_data, test_data, metrics, loss, trainer, \
batch_limit=batch_limit, num_epochs=epochs, verbose=2)
# +
''' Learning rate of 0.001 is a significant improvement'''
net = mx.gluon.nn.Sequential()
with net.name_scope():
net.add(mx.gluon.nn.Dense(128, activation="relu"))
net.add(mx.gluon.nn.Dense(10))
ctx = mx.cpu()
net.collect_params().initialize(mx.init.Xavier(magnitude=2.24), ctx=ctx)
loss = gluon.loss.SoftmaxCrossEntropyLoss()
trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': .001})
batch_limit = 625
epochs = 10
metrics = create_metrics()
results = train_model(net, train_data, test_data, metrics, loss, trainer, \
batch_limit=batch_limit, num_epochs=epochs, verbose=2)
# -
# ### Relu really comes into its own for deep networks
# - Deeper networks tend to perform better than shallow networks for complex tasks
# - But they are hard to train. Relu's make it easier for deep networks to learn because their gradients don't saturate for postive inputs
# +
net = mx.gluon.nn.Sequential()
with net.name_scope():
net.add(mx.gluon.nn.Dense(512, activation="relu"))
net.add(mx.gluon.nn.Dense(256, activation="relu"))
net.add(mx.gluon.nn.Dense(128, activation="relu"))
net.add(mx.gluon.nn.Dense(64, activation="relu"))
net.add(mx.gluon.nn.Dense(10))
ctx = mx.cpu()
net.collect_params().initialize(mx.init.Xavier(magnitude=2.24), ctx=ctx)
loss = gluon.loss.SoftmaxCrossEntropyLoss()
trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': .001})
batch_limit = 625
epochs = 10
metrics = create_metrics()
results = train_model(net, train_data, test_data, metrics, loss, trainer, \
batch_limit=batch_limit, num_epochs=epochs, verbose=2)
# +
''' Same network as above but with sigmoid units - performance is much worse'''
net = mx.gluon.nn.Sequential()
with net.name_scope():
net.add(mx.gluon.nn.Dense(512, activation="sigmoid"))
net.add(mx.gluon.nn.Dense(256, activation="sigmoid"))
net.add(mx.gluon.nn.Dense(128, activation="sigmoid"))
net.add(mx.gluon.nn.Dense(64, activation="sigmoid"))
net.add(mx.gluon.nn.Dense(10))
ctx = mx.cpu()
net.collect_params().initialize(mx.init.Xavier(magnitude=2.24), ctx=ctx)
loss = gluon.loss.SoftmaxCrossEntropyLoss()
trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': .01})
batch_limit = 625
epochs = 10
metrics = create_metrics()
results = train_model(net, train_data, test_data, metrics, loss, trainer, \
batch_limit=batch_limit, num_epochs=epochs, verbose=2)
# -
# ### Putting everything together and training on all the data
# +
net = mx.gluon.nn.Sequential()
with net.name_scope():
net.add(mx.gluon.nn.Dense(512, activation="relu"))
net.add(mx.gluon.nn.Dense(256, activation="relu"))
net.add(mx.gluon.nn.Dense(128, activation="relu"))
net.add(mx.gluon.nn.Dense(64, activation="relu"))
net.add(mx.gluon.nn.Dense(10))
ctx = mx.cpu()
net.collect_params().initialize(mx.init.Xavier(magnitude=2.24), ctx=ctx)
loss = gluon.loss.SoftmaxCrossEntropyLoss()
trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': .001})
batch_limit = 0 # No limit, use all the data
epochs = 10
metrics = create_metrics()
results = train_model(net, train_data, test_data, metrics, loss, trainer, \
batch_limit=batch_limit, num_epochs=epochs, verbose=2)
# -
# ### Evaluation helper functions
# +
def accuracy(test_data, net):
predictions = []
labels = []
alldata = []
test_data.reset()
''' Collect model predictions '''
for i, batch in enumerate(test_data):
with autograd.record():
data = batch.data[0]
label_one_hot = batch.label[0]
output = net(data)
predictions.append(output.asnumpy())
labels.append(label_one_hot.asnumpy())
''' Selecting only the first 10,000 values since the last batch was padded'''
predictions = np.vstack(predictions)[:10000]
labels = np.vstack(labels)[:10000]
''' Calculate accuracy '''
num_correct = np.argmax(predictions, axis=1)==np.argmax(labels, axis=1)
accuracy = 1.0 * np.sum(num_correct) / predictions.shape[0]
print("Accuracy on data is: {}%".format(accuracy * 100))
def get_correct_and_incorrect(net, test_data):
predictions = []
labels = []
alldata = []
test_data.reset()
''' Collect model predictions '''
for i, batch in enumerate(test_data):
with autograd.record():
data = batch.data[0]
label_one_hot = batch.label[0]
output = net(data)
predictions.append(output.asnumpy())
labels.append(label_one_hot.asnumpy())
alldata.append(data.asnumpy())
''' Selecting only the first 10,000 values since the last batch was padded'''
predictions = np.vstack(predictions)[:10000]
labels = np.vstack(labels)[:10000]
alldata = np.vstack(alldata)[:10000]
''' Separate the data into correct and incorrect subsets'''
correct_indices = np.equal(np.argmax(predictions, axis=1),np.argmax(labels, axis=1))
test_x_correct = alldata[correct_indices]
test_y_correct = labels[correct_indices]
predict_test_y_correct = predictions[correct_indices]
incorrect_indices = np.not_equal(np.argmax(predictions, axis=1), np.argmax(labels, axis=1))
test_x_incorrect = alldata[incorrect_indices]
test_y_incorrect = labels[incorrect_indices]
predict_test_y_incorrect = predictions[incorrect_indices]
return test_x_correct, test_y_correct, test_x_incorrect, test_y_incorrect, predict_test_y_correct, predict_test_y_incorrect
# -
# ### Final model performance
test_accuracy = evaluate_accuracy(test_data, net, metrics)
print("Test_acc {:.5f}, Test_top_3_accuracy {:.5f}".format(test_accuracy[0], test_accuracy[1]))
accuracy(train_data, net)
accuracy(test_data, net)
test_x_correct, test_y_correct, test_x_incorrect, test_y_incorrect, predict_test_y_correct, predict_test_y_incorrect = \
get_correct_and_incorrect(net, test_data)
print(test_x_correct.shape)
plotExamples(test_x_correct, test_y_correct, predict_test_y_correct)
print(test_x_incorrect.shape)
plotExamples(test_x_incorrect, test_y_incorrect, predict_test_y_incorrect)
| MXNet_Basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from multiprocessing import shared_memory
import sys
import matplotlib
import numpy as np
from tqdm import tqdm
import xarray as xr
climate_indices_home_path = "/home/james/git/climate_indices"
if climate_indices_home_path not in sys.path:
sys.path.append(climate_indices_home_path)
from climate_indices import compute, indices, utils
# %matplotlib inline
# -
# Open the precipitation dataset as an xarray Dataset object.
ds_prcp = xr.open_dataset("/home/james/data/nclimgrid/nclimgrid_lowres_prcp.nc")
# Get the precipitation data and reshape the array to have the time dimension as the inner-most axis:
da_prcp = ds_prcp['prcp'].transpose('lat', 'lon', 'time')
# Create a NumPy array backed by shared memory and copy the precipitation data into it:
# create a shared memory array for precipitation, copy the precipitation data
# into it, then replace the original underlying array with the shared memory
shm_prcp = shared_memory.SharedMemory(create=True, size=da_prcp.data.nbytes)
shared_prcp = np.ndarray(da_prcp.shape, dtype=da_prcp.dtype, buffer=shm_prcp.buf)
shared_prcp[:,:,:] = da_prcp[:,:,:]
da_prcp.data = shared_prcp
shm_name_prcp = shm_prcp.name
shm_name_prcp
initial_year = int(da_prcp['time'][0].dt.year)
calibration_year_initial = 1900
calibration_year_final = 2000
period_times = 12
total_lats = da_prcp.shape[0]
total_lons = da_prcp.shape[1]
fitting_shape = (total_lats, total_lons, period_times)
scales = [1] # , 2, 3, 6, 9, 12, 24]
periodicity = compute.Periodicity.monthly
# Define a function that can be used to compute the gamma fitting parameters for a particular month scale:
def compute_gammas(
da_precip: xr.DataArray,
scale: int,
calibration_year_initial,
calibration_year_final,
periodicity: compute.Periodicity,
) -> (xr.DataArray, xr.DataArray):
initial_year = int(da_precip['time'][0].dt.year)
if periodicity == compute.Periodicity.monthly:
period_times = 12
gamma_time_coord = "month"
elif periodicity == compute.Periodicity.daily:
period_times = 366
gamma_time_coord = "day"
gamma_coords={"lat": ds.lat, "lon": ds.lon, gamma_time_coord: range(period_times)}
total_lats = da_precip.shape[0]
total_lons = da_precip.shape[1]
fitting_shape = (total_lats, total_lons, period_times)
# create a shared memory array for precipitation, copy the precipitation data
# into it, then replace the original underlying array with the shared memory
shm_prcp = shared_memory.SharedMemory(create=True, size=da_prcp.data.nbytes)
shared_prcp = np.ndarray(da_precip.shape, dtype=da_precip.dtype, buffer=shm_prcp.buf)
shared_prcp[:,:,:] = da_precip[:,:,:]
shm_name_prcp = shm_prcp.name
# create a shared memory array for the gamma distribution alpha and beta fitting parameters
shm_alpha = shared_memory.SharedMemory(create=True) # , size=da_prcp.data.nbytes)
shm_name_alpha = shm_alpha.name
alphas = np.full(shape=fitting_shape, fill_value=np.NaN, buffer=shm_alpha.buf)
shm_beta = shared_memory.SharedMemory(create=True) # , size=da_prcp.data.nbytes)
shm_name_beta = shm_beta.name
betas = np.full(shape=fitting_shape, fill_value=np.NaN, buffer=shm_beta.buf)
# loop over the grid cells and add a new arguments element for each
resize_arguments = {
"file_id": file_id,
"image_ext": image_ext,
"annotation_format": annotation_format,
"annotation_ext": annotation_ext,
"input_images_dir": input_images_dir,
"input_annotations_dir": input_annotations_dir,
"output_images_dir": output_images_dir,
"output_annotations_dir": output_annotations_dir,
"new_width": new_width,
"new_height": new_height,
}
resize_arguments_list.append(resize_arguments)
arguments_list = []
for lat_index in range(total_lats):
for lon_index in range(total_lons):
arguments = {
'lat_index': lat_index,
'lon_index': lon_index,
'shm_name_prcp': shm_name_prcp,
'shm_name_alpha': shm_name_alpha,
'shm_name_beta': shm_name_beta,
'scale': scale,
'calibration_year_initial': calibration_year_initial,
'calibration_year_final': calibration_year_final,
'periodicity': periodicity,
}
arguments_list.append(arguments)
# use a ProcessPoolExecutor to download the images in parallel
with concurrent.futures.ProcessPoolExecutor() as executor:
_logger.info("Computing gamma fitting parameters")
# use the executor to map the download function to the iterable of arguments
list(tqdm(executor.map(compute_gammas_gridcell, arguments_list), total=len(arguments_list)))
alpha_attrs = {
'description': 'shape parameter of the gamma distribution (also referred to as the concentration) ' + \
f'computed from the {scale}-month scaled precipitation values',
}
da_alpha = xr.DataArray(
data=alphas,
coords=gamma_coords,
dims=tuple(gamma_coords.keys()),
name=f"alpha_{str(scale).zfill(2)}",
attrs=alpha_attrs,
)
beta_attrs = {
'description': '1 / scale of the distribution (also referred to as the rate) ' + \
f'computed from the {scale}-month scaled precipitation values',
}
da_beta = xr.DataArray(
data=betas,
coords=gamma_coords,
dims=tuple(gamma_coords.keys()),
name=f"beta_{str(scale).zfill(2)}",
attrs=beta_attrs,
)
return da_alpha, da_beta
def compute_gammas_gridcell(
arguments: Dict,
):
initial_year = int(da_precip['time'][0].dt.year)
if periodicity == compute.Periodicity.monthly:
period_times = 12
gamma_time_coord = "month"
elif periodicity == compute.Periodicity.daily:
period_times = 366
gamma_time_coord = "day"
gamma_coords={"lat": ds.lat, "lon": ds.lon, gamma_time_coord: range(period_times)}
total_lats = da_precip.shape[0]
total_lons = da_precip.shape[1]
fitting_shape = (total_lats, total_lons, period_times)
# create a shared memory array for precipitation, copy the precipitation data
# into it, then replace the original underlying array with the shared memory
shm_prcp = shared_memory.SharedMemory(create=True, size=da_prcp.data.nbytes)
shared_prcp = np.ndarray(da_precip.shape, dtype=da_precip.dtype, buffer=shm_prcp.buf)
shared_prcp[:,:,:] = da_precip[:,:,:]
shm_name_prcp = shm_prcp.name
shm_alpha = shared_memory.SharedMemory(create=True) # , size=da_prcp.data.nbytes)
shm_name_alpha = shm_alpha.name
alphas = np.full(shape=fitting_shape, fill_value=np.NaN, buffer=shm_alpha.buf)
shm_beta = shared_memory.SharedMemory(create=True) # , size=da_prcp.data.nbytes)
shm_name_beta = shm_beta.name
betas = np.full(shape=fitting_shape, fill_value=np.NaN, buffer=shm_beta.buf)
# loop over the grid cells and add a new arguments element for each
resize_arguments = {
"file_id": file_id,
"image_ext": image_ext,
"annotation_format": annotation_format,
"annotation_ext": annotation_ext,
"input_images_dir": input_images_dir,
"input_annotations_dir": input_annotations_dir,
"output_images_dir": output_images_dir,
"output_annotations_dir": output_annotations_dir,
"new_width": new_width,
"new_height": new_height,
}
resize_arguments_list.append(resize_arguments)
arguments_list = []
for lat_index in range(total_lats):
for lon_index in range(total_lons):
arguments = {
'lat_index': lat_index,
'lon_index': lon_index,
'shm_name_prcp': shm_name_prcp,
'shm_name_alpha': shm_name_alpha,
'shm_name_beta': shm_name_beta,
'scale': scale,
'calibration_year_initial': calibration_year_initial,
'calibration_year_final': calibration_year_final,
'periodicity': periodicity,
}
# get the precipitation values for the lat/lon grid cell
values = da_precip[lat_index, lon_index]
# skip over this grid cell if all NaN values
if (np.ma.is_masked(values) and values.mask.all()) or np.all(np.isnan(values)):
continue
# convolve to scale
scaled_values = \
compute.scale_values(
values,
scale=scale,
periodicity=periodicity,
)
# compute the fitting parameters on the scaled data
alphas[lat_index, lon_index], betas[lat_index, lon_index] = \
compute.gamma_parameters(
scaled_values,
data_start_year=initial_year,
calibration_start_year=calibration_year_initial,
calibration_end_year=calibration_year_final,
periodicity=periodicity,
)
# use a ProcessPoolExecutor to download the images in parallel
with concurrent.futures.ProcessPoolExecutor() as executor:
_logger.info("Resizing files")
# use the executor to map the download function to the iterable of arguments
list(tqdm(executor.map(resize_image, resize_arguments_list),
total=len(resize_arguments_list)))
alpha_attrs = {
'description': 'shape parameter of the gamma distribution (also referred to as the concentration) ' + \
f'computed from the {scale}-month scaled precipitation values',
}
da_alpha = xr.DataArray(
data=alphas,
coords=gamma_coords,
dims=tuple(gamma_coords.keys()),
name=f"alpha_{str(scale).zfill(2)}",
attrs=alpha_attrs,
)
beta_attrs = {
'description': '1 / scale of the distribution (also referred to as the rate) ' + \
f'computed from the {scale}-month scaled precipitation values',
}
da_beta = xr.DataArray(
data=betas,
coords=gamma_coords,
dims=tuple(gamma_coords.keys()),
name=f"beta_{str(scale).zfill(2)}",
attrs=beta_attrs,
)
return da_alpha, da_beta
# Define a function that can be used to compute the SPI for a particular month scale:
def compute_spi_gamma(
da_precip: xr.DataArray,
da_alpha: xr.DataArray,
da_beta: xr.DataArray,
scale: int,
periodicity: compute.Periodicity,
) -> xr.DataArray:
initial_year = int(da_precip['time'][0].dt.year)
total_lats = da_precip.shape[0]
total_lons = da_precip.shape[1]
spi = np.full(shape=da_precip.shape, fill_value=np.NaN)
for lat_index in range(total_lats):
for lon_index in range(total_lons):
# get the values for the lat/lon grid cell
values = da_precip[lat_index, lon_index]
# skip over this grid cell if all NaN values
if (np.ma.is_masked(values) and values.mask.all()) or np.all(np.isnan(values)):
continue
gamma_parameters = {
"alphas": da_alpha[lat_index, lon_index],
"betas": da_beta[lat_index, lon_index],
}
# compute the SPI
spi[lat_index, lon_index] = \
indices.spi(
values,
scale=scale,
distribution=indices.Distribution.gamma,
data_start_year=initial_year,
calibration_year_initial=calibration_year_initial,
calibration_year_final=calibration_year_final,
periodicity=compute.Periodicity.monthly,
fitting_params=gamma_parameters,
)
# build a DataArray for this scale's SPI
da_spi = xr.DataArray(
data=spi,
coords=da_precip.coords,
dims=("lat", "lon", "time"),
name=f"spi_gamma_{str(scale).zfill(2)}",
)
da_spi.attrs = {
'description': f'SPI ({scale}-{periodicity} gamma) computed from monthly precipitation ' + \
f'data for the period {da_precip.time[0]} through {da_precip.time[-1]} using a ' + \
f'calibration period from {calibration_year_initial} through {calibration_year_final}',
'valid_min': -3.09,
'valid_max': 3.09,
'long_name': f'{scale}-{periodicity} SPI(gamma)',
'calibration_year_initial': calibration_year_initial,
'calibration_year_final': calibration_year_final,
}
return da_spi
# Copy the attributes from the precipitation dataset that will be applicable to the corresponding gamma fitting parameters and SPI datasets:
attrs_to_copy = [
'Conventions',
'ncei_template_version',
'naming_authority',
'standard_name_vocabulary',
'institution',
'geospatial_lat_min',
'geospatial_lat_max',
'geospatial_lon_min',
'geospatial_lon_max',
'geospatial_lat_units',
'geospatial_lon_units',
]
global_attrs = {key: value for (key, value) in ds.attrs.items() if key in attrs_to_copy}
# Compute the gamma fitting parameters for all scales and add these into a Dataset that we'll write to NetCDF:
# +
# %%time
if periodicity == compute.Periodicity.monthly:
period_times = 12
gamma_time_coord = "month"
elif periodicity == compute.Periodicity.daily:
period_times = 366
gamma_time_coord = "day"
ds_gamma = xr.Dataset(
coords={"lat": ds.lat, "lon": ds.lon, gamma_time_coord: range(period_times)},
attrs=global_attrs,
)
for scale in scales:
var_name_alpha = f"alpha_{str(scale).zfill(2)}"
var_name_beta = f"beta_{str(scale).zfill(2)}"
da_alpha, da_beta = compute_gammas(
da_prcp,
scale,
calibration_year_initial,
calibration_year_final,
periodicity,
)
ds_gamma[f"alpha_{str(scale).zfill(2)}"] = da_alpha
ds_gamma[f"beta_{str(scale).zfill(2)}"] = da_beta
netcdf_gamma = '/home/james/data/nclimgrid/nclimgrid_lowres_gamma.nc'
ds_gamma.to_netcdf(netcdf_gamma)
# -
# Compute the SPI using the pre-computed gamma fitting parameters for all scales and add these into a SPI(gamma) Dataset that we'll write to NetCDF:
# +
# %%time
ds_spi = xr.Dataset(
coords=ds.coords,
attrs=global_attrs,
)
for scale in scales:
var_name_alpha = f"alpha_{str(scale).zfill(2)}"
var_name_beta = f"beta_{str(scale).zfill(2)}"
da_spi = compute_spi_gamma(
da_prcp: xr.DataArray,
ds_gamma[f'alpha_{str(scale).zfill(2)}'],
ds_gamma[f'beta_{str(scale).zfill(2)}'],
scale,
periodicity,
)
ds_spi[f"spi_gamma_{str(scale).zfill(2)}"] = da_spi
netcdf_spi = '/home/james/data/nclimgrid/nclimgrid_lowres_spi_gamma.nc'
ds_spi.to_netcdf(netcdf_spi)
# -
# Plot a time step to validate that the SPI values look reasonable:
ds_spi["spi_gamma_03"].isel(time=500).plot()
| notebooks/muitprocess_spi_nclimgrid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: fundl-dev
# language: python
# name: fundl-dev
# ---
# +
"""Actual moleculenet task."""
from fundl.datasets import make_graph_counting_dataset
from fundl.utils import pad_graph
import numpy as onp
import networkx as nx
import jax.numpy as np
import panel as pn
import param
import holoviews as hv
import hvplot
import hvplot.streamz
# hv.extension("bokeh")
# -
from streamz.dataframe import DataFrame
from streamz import Stream
# +
# pn.extension()
# +
from chemgraph import atom_graph, bond_graph
import janitor.chemistry
# Gs = make_graph_counting_dataset(n_graphs=1000)
import pandas as pd
df = (
pd.read_csv("bace.csv")
.rename_column("mol", "structure")
.smiles2mol("structure", "mol")
.join_apply(lambda x: atom_graph(x["mol"]), "atom_graph")
.join_apply(lambda x: bond_graph(x["mol"]), "bond_graph")
.join_apply(lambda x: len(x["atom_graph"]), "atom_graph_size")
.join_apply(lambda x: len(x["bond_graph"]), "bond_graph_size")
.shuffle()
)
# -
def generate_adjacency_and_feature_matrices(Gs):
Fs = []
As = []
for G in Gs:
Fs.append(np.vstack([d["features"] for n, d in G.nodes(data=True)]))
As.append(onp.asarray(nx.adjacency_matrix(G).todense()))
return As, Fs
# +
aGs = df["atom_graph"].tolist()
bGs = df["bond_graph"].tolist()
print("Generating atom graph feature matrices and adjacency matrices...")
aAs, aFs = generate_adjacency_and_feature_matrices(aGs)
# -
print("Generating bond graph feature matrices and adjacency matrices...")
bAs, bFs = generate_adjacency_and_feature_matrices(bGs)
largest_agraph_size = max([len(G) for G in aGs])
largest_bgraph_size = max([len(G) for G in bGs])
largest_agraph_size, largest_bgraph_size
# +
print("Preparing outputs...")
y = df["pIC50"].values.reshape(-1, 1)
y_train = y[0:800, :]
y_test = y[800:, :]
# y_train = onp.ma.masked_array(y_train, value=7.09691)
# +
def pad_and_stack_graphs(Fs, As, largest_graph_size):
for i, (F, A) in enumerate(zip(Fs, As)):
F, A = pad_graph(F, A, largest_graph_size)
Fs[i] = F
As[i] = A
Fs = np.stack(Fs).astype(np.float32)
As = np.stack(As).astype(np.float32)
return np.array(Fs), np.array(As)
def simple_split(Fs, As):
Fs_train, Fs_test = Fs[0:800, :], Fs[800:]
As_train, As_test = As[0:800, :], As[800:]
return Fs_train, Fs_test, As_train, As_test
# +
# print(aFs.shape), print(aAs.shape)
# print(bFs.shape), print(bAs.shape)
# +
print("Padding atom graphs to correct size...")
aFs, aAs = pad_and_stack_graphs(aFs, aAs, largest_agraph_size)
bFs, bAs = pad_and_stack_graphs(bFs, bAs, largest_bgraph_size)
aFs_train, aFs_test, aAs_train, aAs_test = simple_split(aFs, aAs)
bFs_train, bFs_test, bAs_train, bAs_test = simple_split(bFs, bAs)
def train_data():
return aFs_train, aAs_train, bFs_train, bAs_train, y_train
def test_data():
return aFs_test, aAs_test, bFs_test, bAs_test, y_test
# +
from fundl.weights import add_dense_params
params = dict()
params = add_dense_params(params, name="agraph1", input_dim=9, output_dim=20)
params = add_dense_params(params, name="agraph2", input_dim=20, output_dim=10)
params = add_dense_params(params, name="bgraph1", input_dim=4, output_dim=20)
params = add_dense_params(params, name="bgraph2", input_dim=20, output_dim=10)
params = add_dense_params(params, name="dense1", input_dim=20, output_dim=1)
# +
from fundl.layers.graph import mpnn, gather
from fundl.layers import dense
# from fundl.activations import relu
arr = np.array([-1, 1, 0])
def relu(x, alpha=0.1):
return x * (x > 0) + alpha * x * (x <= 0)
from fundl.losses import _mse_loss
from jax import grad
def mseloss(p, model, aFs, aAs, bFs, bAs, y):
yhat = model(p, aFs, aAs, bFs, bAs)
return np.mean(_mse_loss(y, yhat))
def model(params, aFs, aAs, bFs, bAs):
aFs = mpnn(params["agraph1"], aAs, aFs, nonlin=relu)
aFs = mpnn(params["agraph2"], aAs, aFs, nonlin=relu)
bFs = mpnn(params["bgraph1"], bAs, bFs, nonlin=relu)
bFs = mpnn(params["bgraph2"], bAs, bFs, nonlin=relu)
aOut = gather(aFs)
bOut = gather(bFs)
out = np.concatenate([aOut, bOut], axis=1)
out = dense(params["dense1"], out, nonlin=relu)
return out
dloss = grad(mseloss)
# +
# As a sanity check, calculate mseloss
out = model(params, aFs, aAs, bFs, bAs)
out.sum()
# +
from jax.experimental.optimizers import adam
init, update, get_params = adam(step_size=0.001)
state = init(params)
# Make sure things fit over 10 epochs
for i in range(10):
g = dloss(params, model, aFs_train, aAs_train, bFs_train, bAs_train, y_train)
state = update(i, g, state)
params = get_params(state)
l_test = mseloss(params, model, aFs_test, aAs_test, bFs_test, bAs_test, y_test)
l_train = mseloss(params, model, aFs_train, aAs_train, bFs_train, bAs_train, y_train)
sum_loss = l_test + l_train + abs(l_test - l_train)
print(sum_loss)
# -
def fit(n_epochs, model, params, train_data, test_data):
init, update, get_params = adam(step_size=0.001)
aFs_train, aAs_train, bFs_train, bAs_train, y_train = train_data()
aFs_test, aAs_test, bFs_test, bAs_test, y_test = test_data()
state = init(params)
# Make sure things fit over 10 epochs
for i in range(n_epochs):
g = dloss(params, model, aFs_train, aAs_train, bFs_train, bAs_train, y_train)
state = update(i, g, state)
params = get_params(state)
l_test = mseloss(params, model, aFs_test, aAs_test, bFs_test, bAs_test, y_test)
l_train = mseloss(params, model, aFs_train, aAs_train, bFs_train, bAs_train, y_train)
if i % 10 == 0:
print(l_train, l_test)
return l_train, l_test, params
def objective(trial):
n_epochs = trial.suggest_int("n_epochs", 30, 1000)
loss_train, loss_test, params = fit(n_epochs, model, params, train_data, test_data)
return abs(loss_train - loss_test) + loss_train + loss_test
# +
import optuna
study = optuna.create_study()
study.optimize(objective, n_trials=20)
# -
study.best_params
loss_train, loss_test, params = fit(study.best_params["n_epochs"], model, params, train_data, test_data)
loss_train, loss_test
preds_train = model(params, aFs_train, aAs_train, bFs_train, bAs_train)
preds_test = model(params, aFs_test, aAs_test, bFs_test, bAs_test)
# hv.Scatter((preds_train, y_train)) * hv.Scatter((preds_test, y_test)) * hv.Curve(data=[(0, 0), (20, 20)])
y_train.squeeze().shape
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
# +
import matplotlib.pyplot as plt
plt.scatter(y_train.squeeze(), preds_train.squeeze())
plt.scatter(y_test.squeeze(), preds_test.squeeze())
# -
| examples/moleculenet/graph-live-training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import csv
import tensorflow as tf
import numpy as np
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
import torch
import torchvision
import matplotlib.pyplot as plt
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
n_epochs = 3
batch_size_train = 64
batch_size_test = 1000
learning_rate = 0.01
momentum = 0.5
log_interval = 20
torch.backends.cudnn.enabled = True
# +
vocab_size = 1000
embedding_dim = 16
max_length = 120
trunc_type='post'
padding_type='post'
oov_tok = "<OOV>"
training_portion = .8
#Stopwords list from https://github.com/Yoast/YoastSEO.js/blob/develop/src/config/stopwords.js
stopwords = [ "a", "about", "above", "after", "again", "against",
"all", "am", "an", "and", "any", "are", "as", "at",
"be", "because", "been", "before", "being", "below",
"between", "both", "but", "by", "could", "did", "do",
"does", "doing", "down", "during", "each", "few", "for",
"from", "further", "had", "has", "have", "having", "he",
"he'd", "he'll", "he's", "her", "here", "here's", "hers",
"herself", "him", "himself", "his", "how", "how's", "i",
"i'd", "i'll", "i'm", "i've", "if", "in", "into", "is", "it",
"it's", "its", "itself", "let's", "me", "more", "most", "my",
"myself", "nor", "of", "on", "once", "only", "or", "other",
"ought", "our", "ours", "ourselves", "out", "over", "own",
"same", "she", "she'd", "she'll", "she's", "should", "so",
"some", "such", "than", "that", "that's", "the", "their",
"theirs", "them", "themselves", "then", "there", "there's",
"these", "they", "they'd", "they'll", "they're", "they've",
"this", "those", "through", "to", "too", "under", "until",
"up", "very", "was", "we", "we'd", "we'll", "we're", "we've",
"were", "what", "what's", "when", "when's", "where", "where's",
"which", "while", "who", "who's", "whom", "why", "why's", "with",
"would", "you", "you'd", "you'll", "you're", "you've", "your",
"yours", "yourself", "yourselves" ]
sentences = []; labels = []
with open('D:/44754/Documents/Data/bbc-text2.csv','r') as csvfile:
reader = csv.reader(csvfile,delimiter=',')
next(reader)
for row in reader:
labels.append(row[0])
sentence = row[1]
for word in stopwords:
token = " "+word+" "
sentence = sentence.replace(token," ")
sentences.append(sentence)
# -
train_size = int(len(sentences) * training_portion)
train_sentences = sentences[:train_size]
train_labels = labels[:train_size]
validation_sentences = sentences[train_size:]
validation_labels = labels[train_size:]
tokenizer = Tokenizer(num_words = vocab_size, oov_token=oov_tok)
tokenizer.fit_on_texts(train_sentences)
word_index = tokenizer.word_index
train_sequences = tokenizer.texts_to_sequences(train_sentences)
train_padded = pad_sequences(train_sequences, padding=padding_type, maxlen=max_length)
validation_sequences = tokenizer.texts_to_sequences(validation_sentences)
validation_padded = pad_sequences(validation_sequences, padding=padding_type, maxlen=max_length)
label_tokenizer = Tokenizer()
label_tokenizer.fit_on_texts(labels)
training_label_seq = np.array(label_tokenizer.texts_to_sequences(train_labels))
validation_label_seq = np.array(label_tokenizer.texts_to_sequences(validation_labels))
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.embedding = nn.Embedding(vocab_size,embedding_dim)
self.fc1 = nn.Linear(16, 24)
self.fc2 = nn.Linear(24, 6)
def forward(self, x):
x = self.embedding(x)
x = torch.mean(x,1)
x = F.relu(self.fc1(x))
x = self.fc2(x)
return F.log_softmax(x)
network = Net()
optimizer = optim.SGD(network.parameters(), lr=learning_rate,
momentum=momentum)
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
validation_padded.shape
tmp = np.asarray(train_padded[:3],dtype=np.int64)
tmp = torch.from_numpy(tmp)
a = network(tmp)
train_loader = torch.utils.data.DataLoader(
torchvision.datasets.MNIST('/files/', train=True, download=True,
transform=torchvision.transforms.Compose([torchvision.transforms.ToTensor(),
torchvision.transforms.Normalize((0.1307,), (0.3081,))])),
batch_size=batch_size_train, shuffle=True)
for data,target in train_loader:
print(target.shape)
train_losses = []
train_counter = []
test_losses = []
def train(epoch):
network.train()
for i in range(89):
optimizer.zero_grad()
tmpdata = torch.from_numpy(np.asarray(
train_padded[i*20:(i+1)*20],dtype=np.int64))
output = network(tmpdata)
tmptarget = torch.from_numpy(np.asarray(
training_label_seq[i*20:(i+1)*20].squeeze(),dtype=np.int64))
loss = F.nll_loss(output, tmptarget)
loss.backward()
optimizer.step()
if i % log_interval == 0:
print('Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}'.format(
epoch, i * 20, 1780,
100. *i / 89, loss.item()))
def test():
network.eval()
test_loss = 0
correct = 0
with torch.no_grad():
for i in range(89):
tmpdata = torch.from_numpy(np.asarray(
train_padded[i*5:(i+1)*5],dtype=np.int64))
tmptarget = torch.from_numpy(np.asarray(
training_label_seq[i*5:(i+1)*5].squeeze(),dtype=np.int64))
output = network(tmpdata)
test_loss += F.nll_loss(output,tmptarget, size_average=False).item()
pred = output.data.max(1, keepdim=True)[1]
correct += pred.eq(tmptarget.data.view_as(pred)).sum()
test_loss /= 89
test_losses.append(test_loss)
print('\nTest set: Avg. loss: {:.4f}, Accuracy: {}/{} ({:.0f}%)\n'.format(
test_loss, correct, 445,
100. * correct / 445))
n_epochs = 30
test()
for epoch in range(1, n_epochs + 1):
train(epoch)
test()
| NLP_Keras2TFPyTorch/PyTorchnlp2.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Cvičení 1 - Stručný úvod do R, Kombinatorika
# ## <NAME>, <NAME>, <NAME>
# Nejprve si projdeme některé základy Rka a naimplementujeme si některé kombinatorické funkce.
#
# # Krátký úvod do R
# jednoduché početní operace
2+4
5/2
# POZOR na závorky! Pro počítání se používají pouze kulaté!
# Hranaté a složené mají v R jinou funkci!
(((10+2)*(340-33))-2)/3
# kombinační číslo, faktoriály
choose(10,2)
factorial(4)
# datové typy -> numeric, character, logical, (complex)
# funkcí class zjišťujeme typ objektu
a=2+3
class(a)
b="pismenko"
class(b)
c=(1>3)
class(c)
d=3+1i
class(d)
# ## datové struktury v R
#
# - vector (rozumíme sloupcový vektor)
# - factor (speciální případ vektoru)
# - matrix (matice s rozměry n x m)
# - data.frame (datový rámec s rozměry n x p)
# - list (seznam)
# definování vektoru (sloupcový=column)
a = c(3,4,6,7)
a <- c(3,4,6,7)
a[2]
# další možnosti
rep(1,4) # vytvoří vektor se čtyřmi jedničkami
seq(1,10,2) #posloupnost od 1 do 10 s krokem 2
1:10 #posloupnost od 1 do 10 s krokem 1
b=c("A","B","C","D")
b
class(b)
#předefinování objektu na jiný typ - např. as.vector, as.matrix, as.factor,...
b=as.factor(b)
b
#práce s vektory - slučování podle sloupců/řádků
cbind(a,b)
rbind(a,b)
c(a,b)
## definování matice
A=matrix(c(3,4,6,7,3,2),nrow=2,ncol=3)
B=matrix(c(3,4,6,7,3,2),nrow=2,ncol=3,byrow=TRUE)
C=matrix(c(3,4,6,7,3,2),nrow=3,ncol=2)
B
B[1,3]
A[1,]
A[,2:3]
#diagonální matice
diag(4)
diag(4,2)
## operace s maticemi - pozor na maticové násobení -> %*%
A+B
A-B
A*B
A%*%C
# <hr>
#
# # Kombinatorika
#
# ## Variace
#
# V(n,k) - variace bez opakování, první argument bude celkový počet entit, druhý argument velikost výběru
# funkce se vytváří příkazem fucntion, je to objekt jehož jméno je dáno až proměnnou
# do které tento objekt přiřadím
variace = function(n,k) # zde zadávám počet parametrů a jejich jména
{ # celé tělo funkce je uzavřeno mezi závorkami {...}
citatel = factorial(n) # faktoriál v originálním Rku existuje tak jej použijeme
jmenovatel = factorial(n-k)
return(citatel/jmenovatel) # to co funkce vrátí se dává do příkazu return(...)
}
# V*(n,k) - variace s opakováním
variace_opak = function(n,k)
{
return(n^k)
}
# ## Permutace
#
# P(n)=V(n,n) - permutace
permutace = function(n)
{
return(variace(n,n))
}
# P*(n1,n2,n3,....,nk) - permutace s opakováním, vstup bude vektor s jednotlivými počty unikátních entit
permutace_opak = function(vec_n) # vec_n je vektro počtů hodnot př.: vec_n = c(2,2,2,4,3)
{
n = sum(vec_n) # spočteme kolik máme hodnot celkem
res_temp=factorial(n) #jejich faktoriál = hodnota v čitateli
# jednoduchý cyklus začíná příkazem for, pak v závorkách následuje název iterátoru a z
# jakého seznamu bude brán
for(pocet in vec_n) # pocet je iterátor a postupně bude nabývat hodnot z vektoru vec_n
{
# postupně dělíme faktoriálem každého počtu unikátních entit
res_temp=res_temp/factorial(pocet)
}
return(res_temp)
}
# ## Kombinace
#
# C(n,k) - kombinace
kombinace = function(n,k)
{
return(choose(n,k)) # funkce for kombinace už existuje v Rku a jmenuje se choose
}
# C*(n,k) - kombinace s opakováním
kombinace_opak = function(n,k)
{
return(choose(n+k-1,k)) # použijeme známý vzorec
}
# # Úlohy na cvičení
#
# ## Příklad 1.
#
# Které heslo je bezpečnější?
# * Heslo o délce osm znaků složené pouze z číslic.
# * Heslo o délce pět znaků složené pouze z písmen anglické abecedy.
# heslo 1
h1 = variace_opak(n = 10, k = 8)
# heslo 2
h2 = variace_opak(n = 26, k = 5)
h1/h2
# ## Příklad 2.
#
# Jak dlouho by trvalo vyřešení problému obchodního cestujícího pro n = 10 měst hrubou silou, jestliže vyhodnocení délky každé z možných cest trvá 1 µs?
n = 10
pocet = permutace(n-1)/2
cas = pocet/1000000
cas
# ## Příklad 3.
#
# Jak rozdělit kořist mezi 2 loupežníky, aby dostali oba věci ve stejné hodnotě (případně co nejbližší hodnotě). Tj. lze rozdělit N zadaných čísel do dvou skupin tak, aby součet čísel v obou skupinách byl stejný?
#
# **Kolik možností by bylo třeba vyzkoušet, pokud bychom úlohu řešili hrubou silou?**
N = 10
L = 4
variace_opak(n = L, k = N)
# ## Příklad 4.
#
# Kolik anagramů slova "AUTO" můžeme vytvořit?
#
# Kolik anagramů slova "AUTOMOBILKA" můžeme vytvořit? Kolik z nich začína na "K"?
# +
permutace(4)
vec = c(2,1,1,2,1,1,1,1,1)
sum(vec)
permutace_opak(vec)
vec = c(2,1,1,2,1,1,1,1)
sum(vec)
permutace_opak(vec)
# -
# ## Příklad 5.
#
# V obchodě mají 6 druhů barevných hrníčků.
# - Kolika různými způsoby můžeme koupit 4 různě-barevné hrníčky?
# - Kolika různými možnostmi můžeme nakoupit 5 hrníčků (pokud nám nevadí více od stejné barvy)?
# - Jak se situace změní, pokud budou mít od každého pouze 4 kusy (a nám nevadí více stejné barvy)?
kombinace(6,4)
kombinace_opak(6,5)
kombinace_opak(6,5) - 6
# ## Příklad 6. (sbírka kap. 1, př. 7,8)
#
# Z urny se třemi koulemi, dvěma červenými a jednou bílou, budou současně vybrány dvě koule.
# Student a učitel uzavřou sázku. Pokud budou obě koule stejné barvy, vyhraje student. Pokud
# budou mít koule různou barvu, vyhraje učitel.
# - Je hra férová? Jaké jsou pravděpodobnosti výhry učitele a studenta?
# - Jakou kouli je třeba přidat, aby hra byla férová?
kombinace(3,2)
kombinace(4,2)
# ## Příklad 7.
#
# V balíčku je 5 různých párů ponožek (levá a pravá ponožka je vždy stejná).
# - Kolik různých dvojic ponožek lze vybrat?
# - Kolika různými způsoby se mohu obout? (tj. záleží na tom co je na které noze)
kombinace_opak(n = 5,k = 2)
variace_opak(n=5,k=2)
kombinace_opak(n = 5,k = 2)*2 - 5
# ## Příklad 8.
#
#
# Mám 12 závaží o hmotnostech 1,2,...,12 kg.
# - Kolika způsoby je mohu rozdělit na 2 hromádky?
# - Kolika způsoby je mohu rozdělit na 3 hromádky?
# - Kolika způsoby je mohu rozdělit na 3 hromádky, má-li na všech být stejný počet závaží?
# - Kolika způsoby je mohu rozdělit na 3 hromádky o stejném počtu závaží, pokud hmotnost žádné z nich nesmí překročit 40 kg?
variace_opak(2,12)
variace_opak(3,12)
(variace_opak(3,12)-3)/permutace(3)+1
(variace_opak(3,12)-(variace_opak(2,12)-2)*3-3)/permutace(3)
kombinace(12,4)*kombinace(8,4)/permutace(3)
permutace(12)/(permutace(4)*permutace(4)*permutace(4)*permutace(3))
kombinace(12,4)*kombinace(8,4)/permutace(3)-kombinace(8,4)
# ## Příklad 9.
#
# Mám 20 semínek od každého ze tří druhů zeleniny (mrkev, ředkvička, celer). Bohužel se pomíchala.
# - Do truhlíku zasadím 5 náhodných semínek. Jaká je pravděpodobnost, že mezi nimi budou alespoň tři ředkvičky?
# - Do truhlíku zasadím 5 náhodných semínek. Jaká je pravděpodobnost, že mezi nimi bude více mrkví než celerů?
(kombinace(20,3)*kombinace(40,2)+kombinace(20,4)*kombinace(40,1)+kombinace(20,5))/kombinace(60,5)
| CV1/cv1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib as mpl
from matplotlib import pyplot as plt
import h5py
import tensorflow as tf
print("Num GPUs Available: ", len(tf.config.list_physical_devices('GPU')))
# %load_ext autoreload
# %autoreload 2
import importlib
# ## Set Training Label
label = 'NewSim_TypeSelfAttention'
# ## Find all files
from glob import glob
# files_loc = "/sdf/home/r/rafaeltl/home/rafaeltl/Muon/Feb15/si-mu-lator/batch_slac/out_files/March24_bkgr_2375000/"
files_loc = "/gpfs/slac/atlas/fs1/u/rafaeltl/Muon/Apr2022/si-mu-lator/batch_slac/out_files/bkgr_1/"
# files_bkg = glob(files_loc+'*Muon*bkgr*70000.0*.h5')
files_bkg = glob(files_loc+'*NoMuon*.h5')
files_sig = glob(files_loc+'*WithMuon*.h5')
all_files = files_bkg+files_sig
# ## Open files
import dataprep
importlib.reload(dataprep)
# + jupyter={"outputs_hidden": true} tags=[]
data, dmat, Y, Y_mu, Y_hit, sig_keys = dataprep.make_data_matrix(all_files, max_files=500, sort_by='z')
# -
Y_mu.shape
sig_keys
dmat[:,:,sig_keys.index('time')] = dmat[:,:,sig_keys.index('time')].astype(int)
dmat[:,:,sig_keys.index('time')] = dmat[:,:,sig_keys.index('time')].astype(float)
dmat[:,:,sig_keys.index('z')]
dmat[:,:,sig_keys.index('is_muon')]
# + tags=[]
fig, axs = plt.subplots(4, 4, figsize=(20,20))
axs = axs.flatten()
for ivar in range(dmat.shape[2]):
valid_hits_mu = dmat[Y_mu == 1,:,sig_keys.index('is_muon')].flatten() > -90
valid_hits_nomu = dmat[Y_mu == 0,:,sig_keys.index('is_muon')].flatten() > -90
this_var_mu = dmat[Y_mu == 1,:,ivar].flatten()[valid_hits_mu]
this_var_nomu = dmat[Y_mu == 0,:,ivar].flatten()[valid_hits_nomu]
this_max = np.max([*this_var_mu, *this_var_nomu])
this_min = np.min([*this_var_mu, *this_var_nomu])
axs[ivar].hist( this_var_nomu, histtype='step', range=(this_min, this_max), bins=50, label = 'No muon Evts' )
axs[ivar].hist( this_var_mu, histtype='step', range=(this_min, this_max), bins=50, label = 'Muon Evts' )
axs[ivar].set_xlabel(sig_keys[ivar])
if ivar == dmat.shape[2] - 1:
axs[ivar].legend()
plt.show()
# +
vars_of_interest = np.zeros(dmat.shape[2], dtype=bool)
training_vars = [ 'z', 'ptilt', 'time', 'projX_at_rightend_x', 'projX_at_middle_x' ]
for tv in training_vars:
vars_of_interest[sig_keys.index(tv)] = 1
# -
X = dmat[:,:,vars_of_interest]
X.shape
Y_mu.sum()
# ## Define network
import sys
sys.path.insert(0, '../')
import models
# +
lambs = [1]
mymods = []
for ll in lambs:
# mymodel = models.muon_nn_type2( (X.shape[1],X.shape[2]), ll)
# mymodel = models.muon_nn_selfatt( (X.shape[1],X.shape[2]), ll)
# mymodel = models.muon_nn_type0( (X.shape[1],X.shape[2]))
mymodel = models.muon_nn_selfatt_muonly( (X.shape[1],X.shape[2]))
# mymodel = models.recurrent_model( (X.shape[1],X.shape[2]))
mymods.append(mymodel)
# -
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(X, Y[:,0], test_size=0.33, random_state=42)
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
# + tags=[]
histories = []
for mod,ll in zip(mymods, lambs):
history = mod.fit( X_train, Y_train,
callbacks = [
EarlyStopping(monitor='val_loss', patience=1000, verbose=1),
ModelCheckpoint(f'weights/{label}_ll_{ll}.h5', monitor='val_loss', verbose=True, save_best_only=True) ],
epochs=3000,
validation_split = 0.1,
batch_size=2**14,
verbose=1
)
mod.load_weights(f'weights/{label}_ll_{ll}.h5')
histories.append(history)
# -
for history,ll in zip(histories,lambs):
plt.Figure()
for kk in history.history.keys():
plt.plot(history.history[kk], label=kk)
plt.legend()
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.title(f'Lambda = {ll}')
plt.savefig(f'plots/{label}_loss_ll_{ll}.pdf')
plt.show()
Y_test.shape
from sklearn.metrics import roc_curve
for mod,ll in zip(mymods,lambs):
Y_pred = mod.predict(X_test, verbose=1).flatten()
plt.Figure()
plt.hist(Y_pred[Y_test==0], histtype='step', bins=50, range=(0,1))
plt.hist(Y_pred[Y_test==1], histtype='step', bins=50, range=(0,1))
plt.yscale('log')
plt.title(f'Lambda = {ll}')
plt.savefig(f'plots/{label}_muon_pred_ll_{ll}.pdf')
plt.show()
plt.show()
# +
fig, axs = plt.subplots(1, 2, figsize=(16, 8) )
axs = axs.flatten()
coli = 3
icol = 0
for mod,ll in zip(mymods,lambs):
Y_pred = mod.predict(X_test, verbose=1)
# Y_pred_hits = Y_pred[:,1:]
# Y_pred_mu = Y_pred[:,0]
Y_pred_mu = Y_pred
# Y_pred_hits_f_mu = Y_pred_hits[Y_test_mu==1].flatten()
# Y_pred_hits_f_nomu = Y_pred_hits[Y_test_mu==0].flatten()
# fpr_hits, tpr_hits, _ = roc_curve(Y_test_hits_f_mu[Y_test_hits_f_mu>-90], Y_pred_hits_f_mu[Y_test_hits_f_mu>-90])
# axs[0].semilogy(tpr_hits, 1./fpr_hits, color=f'C{coli+icol}', label=f'lambda = {ll}')
fpr_mus, tpr_mus, _ = roc_curve(Y_test, Y_pred_mu)
axs[1].semilogy(tpr_mus, 1./fpr_mus, color=f'C{coli+icol}', label=f'lambda = {ll}')
icol+=1
axs[0].set_ylabel('Background hits rejection')
axs[0].set_xlabel('Signal hits efficiency')
axs[0].legend()
axs[0].set_xlim(-0.01, 1.01)
axs[0].set_ylim(0.5, 1e6)
axs[1].set_ylabel('Rejection of events with no muons')
axs[1].set_xlabel('Efficiency of events with muons')
axs[1].set_xlim(0.8,1.01)
axs[1].set_ylim(0.5, 1e5)
axs[1].legend()
plt.savefig(f'plots/{label}_ROCs.pdf', transparent=True)
plt.show()
# -
| nn/notebooks/Simple-Mu-Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/reneholt/buffer-overflow-stack/blob/main/Buffer_overflow_stack.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="vxMoHg9zQBf0"
# # What are buffer overflow attacks and how are they thwarted?
# + [markdown] id="YBDFy-ktQGcg"
# **Example – Encoding hexadecimal characters as byte values**
#
# For software developers interested in a recent buffer overflow discovered in 2021, we offer the following code in C, which is a simplified and rewritten version of a vulnerability in the ZTE MF971R LTE router tracked as CVE‑2021‑21748:
# + colab={"base_uri": "https://localhost:8080/"} id="SyxE3_e1O0Vp" outputId="fad7d507-d3e6-4e00-930b-a02088901188"
# %%writefile EncodeHex.c
#include <stdio.h>
#include <string.h>
void encodeHexAsByteValues(const char *hexString, char *byteValues) {
signed int offset = 0;
int hexValues = 0;
int byte = 0;
while (offset < strlen(hexString)) {
strncpy((char *) &hexValues, &hexString[2 * offset], 2u);
sscanf((const char *) &hexValues, "%02X", &byte);
byteValues[offset++] = byte; // The return address can be overwritten opening a path for the
// insertion of exploit code
}
}
int main(void) {
const char* hexString = "0123456789ABCDEF01234";
char byteValues[4];
encodeHexAsByteValues(hexString, byteValues); // There is no size check to ensure that
// hexString is not too long for byteValues
// before calling the function
return 0;
}
# + colab={"base_uri": "https://localhost:8080/", "height": 346} id="9JWTMKN_PF9d" outputId="3391ee28-5c44-414c-d525-cb9af76b8f6a"
# %%shell
gcc EncodeHex.c -o EncodeHex
./EncodeHex
# + [markdown] id="2eXAkzxoQTYx"
# The program above demonstrates a function that encodes a string consisting of hexadecimal-compatible characters into a form with half the memory requirement. Two characters can stand in as actual byte values (in hexadecimal), so that the characters ‘0’ and ‘1’, represented with the byte values 30 and 31, respectively, can be represented literally as the byte value 01. This functionality was used as part of the ZTE router’s handling of passwords.
#
# As noted in the comments of the code, the hexString, having a size of 21 characters, is too large for the byteValues buffer, which only has a size of 4 characters (even though it can accept up to 8 characters in encoded form), and there is no check to ensure that the encodeHexAsByteValues function won’t lead to a buffer overflow.
| Buffer_overflow_stack.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Of1XjJLzOeCR"
# Cho một list các số nguyên n phần tử lst được nhập từ bàn phím. Bạn hãy viết chương trình hiển thị ra màn hình số nhỏ nhất trong list vừa nhập.
# + colab={"base_uri": "https://localhost:8080/"} id="ohsanXuON_k8" outputId="fbcd3d3f-b148-44cd-a594-4c5fd4de4860"
n = int(input())
lst = []
for i in range(n):
lst.append(int(input()))
print(min(lst))
# + [markdown] id="CXJwqSBUOg4M"
# Cho một list các số nguyên n phần tử lst được nhập vào từ bàn phím, bạn hãy viết chương trình tính tổng các phần tử trong list vừa nhập.
# + colab={"base_uri": "https://localhost:8080/"} id="vnhwOKa_OkGF" outputId="726de501-e25e-42ed-cc87-b620b03b63e4"
n = int(input())
lst = []
for i in range(n):
lst.append(int(input()))
print(sum(lst))
# + [markdown] id="H2o9U8nRPHVs"
# Cho một list các số nguyên n phần tử lst được nhập vào từ bàn phím, bạn hãy viết chương trình sắp xếp các phần tử trong list theo thứ tự tăng dần và hiển thị list đó ra màn hình.
# + id="1Mu3hDxXPIVF"
n = int(input())
lst = []
for i in range(n):
lst.append(int(input()))
lst.sort()
print(lst)
# + [markdown] id="zMXaoCxAPMS4"
# Cho một list các số nguyên n phần tử lst được nhập vào từ bàn phím, bạn hãy viết chương trình hiển thị ra màn hình một list chứa các số số lẻ trong list vừa nhập.
# + id="a0Fa2fGzPM4Y"
n = int(input())
lst = []
odd = []
for i in range(n):
lst.append(int(input()))
for i in lst:
if i % 2 != 0:
odd.append(i)
print(odd)
# + [markdown] id="qrhK7EKCPTTk"
# Cho một list các số nguyên n phần tử lst được nhập vào từ bàn phím, bạn hãy viết chương trình hiển thị ra màn hình một list chứa các số chia hết cho 5 trong list vừa nhập, nếu list không có số nào chia hết cho 5 thì hiển thị ra màn hình [0].
# + id="gVz5HWHKPUKz"
n = int(input())
lst = []
five = []
for i in range(n):
lst.append(int(input()))
for i in lst:
if i % 5 == 0:
five.append(i)
if len(five) == 0:
five = [0]
print(five)
| lab7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-1:081325390199:image/datascience-1.0
# ---
# #### Imports
import boto3
import json
# #### Essentials
# ENDPOINT_NAME = 'ENTER DEPLOYMENT ENDPOINT HERE'
ENDPOINT_NAME = 'jumpstart-dft-news-sentiment-classification'
CONTENT_TYPE = 'application/x-text'
sagemaker = boto3.client('runtime.sagemaker')
# #### Invoke SageMaker Endpoint for Prediction
label_map = {0: 'negative', 1: 'positive', 2: 'neutral'}
def predict(encoded_text):
response = sagemaker.invoke_endpoint(EndpointName=ENDPOINT_NAME,
ContentType=CONTENT_TYPE,
Body=encoded_text)
prediction = json.loads(response['Body'].read())['predictions'][0]
class_index = prediction.index(max(prediction))
return label_map[class_index]
text = 'Apple plans to develop in stages an area of no less than 100,000 sq. meters in order to host companies working in information technologies and telecommunications'
text
prediction = predict(text)
print(f'Predicted sentiment = {prediction}')
| finserv/language/news-sentiment-classification/make-prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <div style="text-align:center"><u>REPORT:</u> Technical interview proposed by Jolimoi</div>
# ## <div style="text-align:center;"><u>DATA:</u> Title, artist, date, and medium of every artwork in the MoMA collection</div>
# ### Some Context:
# According to [kaggle](https://www.kaggle.com/momanyc/museum-collection), the Museum of Modern Art (MoMA) acquired its first artworks in 1929, the year it was established. Today, the Museum’s evolving collection contains almost 200,000 works from around the world spanning the last 150 years. The collection includes an ever-expanding range of visual expression, including painting, sculpture, printmaking, drawing, photography, architecture, design, film, and media and performance art.
# Note that the data source was last updated 3 years ago.
# ### Which artist in this data set lived the longest?
# <strong><NAME></strong>, a Russian cartoonist, born on September 28, <em>1900</em> and died on October 1, <em>2008</em>, is the oldest artist in the dataset at <strong>108</strong> years of age.
# 
# ### Who are the top 10 artists by the number of artworks?
import files.utils as utils
utils.topTenArtists()
# ### Which artist has created the most artwork by total surface area?
# We have focused on 2D artworks to avoid misclassification.
utils.topTenArtistsByArea()
# <strong><NAME></strong> is clearly the one who produced the most artwork with a total of <strong>735.05 m²</strong> of 2D art.
# ### Did any artists have artwork acquired during their lifetime?
# Yes, <strong>2593</strong> artists had artwork(s) acquired during their lifetime. They are distributed this way:
utils.lifeTimeAquirement()
# ### Please review the quality of the data, and present any issues
# Fisrt, we can notice a <em>lack of data types consistency</em> between artists and artworks tables.
# Ex: Dates are not in the same type too. `Acquisition Date` and `Date` are strings and `Birth Year` is a float, just as `Death Year`. As we've seen earlier, there is also a format difference between those 4 columns, worse still, inside the `Acquisition Date` and the `Date` column.
# To fix this issue, we can reduce each date at its `yyyy` year format and delete the other values and then unify the data type.
# Then there is way too many sources of information about dimensions of the artworks. This looks messy and ambiguous because of the 4 columns concerning cartesian dimensional data.
# We also have 2 information inputs about dimensions of circular objects when we only need one. The `Radius` would have been a <em>better indicator</em> (standard metric) than the `Diameter` and the `Circunference`.
# On top of that, the columns' type is `float64` and 4 out of 6 are <em>almost empty</em>, that is <em>space consumming</em>.
# We also can find an `artworks.Dimensions` input which is quite useless here.
# Giving instead, the previously computed `Area` informations is a solution to avoid loosing too much data and reduce the size our data set from the fact that it's an agglomeration of the other columns.
# `Area` is expressed in <em>cm²</em>.
# `artworks.weight`, `artworks.duration` are empty, we removed them.
# `artworks.'Object Number'` is redundant with `artworks.'Acquisition date'` and not really clear, we removed it.
# `Medium`, `Department`, `Classification` are highly correlated, for that reason we can remove `Medium` and `Department`.
# Then we merged the resulting tables on `Artist ID` to get this table :
utils.header()
# The biggest issue of this database is <em>data consistency</em>. The feeling after beeing through this data exploration is the following : it does not seem to be any validation or verification about the format the data should have. To boost the quality of the data it should have at least one validation script on the server-side before inserting the data in the database. Sometimes even the data type is not very accurate (ex: dates should have a dateTime type at least).
# The solution is to set up a server side process to validate every transaction with the database. It is also possible to set up a correction process to try to 'restore' a value if it comes with a wrong format.
# The other issue is data redundancy. As we seen previously, some inputs reprensent of the same thing, that leads to unecessary strorage consumption.
# ### Please group the artworks into as many clusters as you feel is appropriate, using attributes from both the artist and artworks tables, and assign each artwork to this new cluster
# We can clearly identify 3 distinct "average" individuals which represent 3 major types of artworks within the database.
utils.clusters()
# ### Code's Notebook
# If you want to know the details, please have a look the [Code](./moma_study.ipynb).
| notebook/Report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Post Mortem - Take 1
#
# Version 1 of the analysis notebook.
#
# Project goals were to see how much data there was for each CAN ID
class CanMsg(object):
"""Generic class to represent can data."""
def __init__(self, canid, data):
self.canid=canid
self.data=data
def __repr__(self):
return f"CAN<{self.canid}, {self.data}>"
# Open the log file for analysis and return unique data messages for each CAN ID.
#
#
# `candump` creates a text file as such:
#
# ```
# (1596811036.445425) can2 2A8#020000000080A0CC
# (1596811036.447770) can2 311#7ACC095041
# ```
def parse_log(log_file):
# Create a list to hold all of the CanMsg objects.
messages = list()
""" Read the log_file. """
with open(log_file, "r") as f:
for can_msg in f.readlines():
can_id, can_data = can_msg.strip().split(" ")[-1].split("#")
cm = CanMsg(can_id, can_data)
messages.append(cm)
# Create a dict to hold unique data
unique_datas = dict()
# For each unique CAN ID.
for canid in set([msg.canid for msg in messages]):
# Get the unique messages on for each ID
id_datas = set([msg.data for msg in messages if msg.canid==canid])
# Add it to the dict using the canid as the key.
unique_datas[canid] = id_datas
# Return it.
return unique_datas
data = parse_log("final_log")
# Split the data messages into high and low frequency groups with the number of times the ID was seen:
thresh=16
print("High Frequency Message IDs:")
for key, items in data.items():
l = len(items)
if l>thresh:
print(f" {key}: {len(items)}")
print("Low Frequency Message IDs:")
for key, items in data.items():
l = len(items)
if l<=thresh:
print(f" {key}: {len(items)}")
# Pick out one ID and see which bits change and by how much:
data["3A6"]
# # TODO:
#
# - Analyze each of the can messages taking guesses at datatypes.
# - Look at the dbc files to see if any of those message ID's
# - Sleep.
#
# Thanks to everyone at the vCHV team! I haven't had this much fun since HvZ CTF.
# *4 months later*
#
# Documented documentation and writeup.
| 01_PostMortem_Local.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import itertools
import numpy as np
np.random.seed(42)
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import train_test_split
from sklearn.datasets import load_breast_cancer
from sklearn.model_selection import validation_curve
# +
dataset = load_breast_cancer()
x = dataset.data
y = dataset.target
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.3)
# +
param_range = range(1, 30)
train_scores, test_scores = validation_curve(
KNeighborsClassifier(),
x_train,
y_train,
param_name="n_neighbors",
param_range=param_range,
cv=10,
scoring="accuracy",
n_jobs=-1
)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
# +
plt.title("Validation Curve with KNN")
plt.xlabel("n_neighbors")
plt.ylabel("Score")
plt.ylim(0.8, 1.1)
lw = 2
plt.plot(
param_range,
train_scores_mean,
label="Training score",
color="darkorange",
lw=lw
)
plt.fill_between(
param_range,
train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std,
alpha=0.2,
color="darkorange",
lw=lw
)
plt.plot(
param_range,
test_scores_mean,
label="Cross-validation score",
color="navy",
lw=lw)
plt.fill_between(
param_range,
test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std,
alpha=0.2,
color="navy",
lw=lw
)
plt.legend(loc="best")
plt.show()
| Chapter8_MetricsAndEvaluation/ValidationCurve/ValidationCurve.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introducción a Series Temporales
# importación de paquetes
import pandas as pd
import numpy as np
# loading data...
rawCsvData = pd.read_csv("../../DataSets/Index2018.csv")
# Observaciones y variables
rawCsvData.shape
# Visualización de la data
rawCsvData.head()
# Hacemos una copia de la data para hacer algunas transformaciones
dfComp = rawCsvData.copy()
# Veamos si se copio correctamente
dfComp.head(5)
# #### Spx(500): Bolsa de Extados Unidos
# #### Dax(): Bolsa Alemana
# #### ftse(): Bolsa de Londres
# #### nikkei(): Bolsa de Japón
#
# ##### Precios de cierre en cada uno de esos días
# Veamos si nuestra lista de data set tiene datos no disponibles
dfComp.isna().sum()
# +
# si me interesa ver na`s en una columna en particular
dfComp.spx.isna().sum()
# -
# Una descripción general de los datos
dfComp.describe()
# #### Obs: Diferencias de magnitud de cada serie de tiempo
| Temas/1.0.Introduction/Carga de data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Recommendation Systems
#
# In this module, we review a few of the most popular algorithms for recommendation systems.
#
# ## System 1: Population Averages
#
# The simplest method is to take the average of all available data on a particular item, and then rank them based on that average.
#
# Pros:
#
# * Incredibly easy to implement (no data science required really)
#
# Cons:
#
# * All of them. If you have no data or little data, either doesn't work or wildly biased from the initial reviews.
# * Treats all people as having the same preferences.
# * All reviews are treated equally
#
#
# ### Improvements
#
# There are a few corrections that one can make which address the first con.
#
# #### Correcting for Small Numbers of Reviews
#
# Take the new rating to be the: $$\text{average rating} - \text{Constant}/\sqrt{\text{# of ratings}}$$
#
# Where the constant is strategically chosen based on the possibly range of values for the average. If it's between 0 and 1, try a constant of 0.5. If it's between 0 and 5, then try a constant of 2.5.
#
# The behavior of this correction is for more conservative reviews. If we flip the correction to be positive instead of negative, we would then have a system which learned on higher rating and encouraged more exploration from our users.
| 4. Recommendation Systems.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 信号処理シンポジウム分析
# モジュール
import os
import pandas as pd
from sipsymp import SipSymp
from sipwdc import SipWordCloud
# 対象年度の設定
syear = 2008
eyear = 2017
# 期間内の論文タイトル
path = './siprep{0}_{1}.csv'.format(syear,eyear)
if os.path.exists(path):
print('{0} file exists.'.format(path))
df = pd.read_csv(path,index_col=0)
else:
print('{0} file doesn\'t exist.'.format(path))
term = range(syear,eyear)
df = SipSymp.titlesDuring(term)
# CSVへの書き出し
df.to_csv(path)
# SIPワードクラウドオブジェクトの生成
swc = SipWordCloud()
# 2008-2012のタイトルワードクラウド
df2008_2012 = df.loc[df.loc[:,'Year']<2013].reset_index(drop=True)
swc.generate(df2008_2012,'sipwdc2008_2012.png')
# 2013-2017のタイトルワードクラウド
df2013_2017 = df.loc[df.loc[:,'Year']>2012].reset_index(drop=True)
swc.generate(df2013_2017,'sipwdc2013_2017.png')
| siprep.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
#
# ### ODPi Egeria Hands-On Lab
# # Welcome to the Configuring Egeria Servers Lab
# ## Introduction
#
# ODPi Egeria is an open source project that provides open standards and implementation libraries to connect tools, catalogs and platforms together so they can share information about data and technology. This information is called metadata.
#
# In this hands-on lab you will learn how to configure the metadata servers used by [Coco Pharmaceuticals](https://opengovernance.odpi.org/coco-pharmaceuticals/).
# ## The scenario
#
# <img src="https://raw.githubusercontent.com/odpi/data-governance/master/docs/coco-pharmaceuticals/personas/gary-geeke.png" style="float:left">
#
# Coco Pharmaceuticals is going through a major business transformation that requires them to drastically reduce their cycle times, collaborate laterally across the different parts of the business and react quickly to the changing needs of their customers. (See [this link](https://opengovernance.odpi.org/coco-pharmaceuticals/) for the background to this transformation).
#
# Part of the changes needed to the IT systems that support the business is the roll out of a distributed metadata and governance capability that is provided by ODPi Egeria.
#
# [<NAME>](https://opengovernance.odpi.org/coco-pharmaceuticals/personas/gary-geeke.html) is the IT Infrastructure leader at Coco Pharmaceuticals.
#
# In this hands-on lab Gary is configuring the metadata servers ready for deployment. Gary's userId is `garygeeke`.
# +
import requests
adminUserId = "garygeeke"
# -
# He needs to define the metadata servers for Coco Pharmaceuticals.
organizationName = "Coco Pharmaceuticals"
# ## Metadata management landscape
#
# Gary has decided to deploy a separate metadata server for each part of the organization that owns assets.
# You can think of a metadata server supporting a community of users. The metadata repositories are as follows:
#
# * cocoMDS1 - Data Lake Operations - used to manage the data in the data lake.
# * cocoMDS2 - Governance - used by all of the governance teams to operate the governance programs.
# * cocoMDS3 - Research - used by the research teams who are developing new treatments.
# * cocoMDS4 - Data Lake Users - used by general business users and the executive team to access data from the data lake.
# * cocoMDS5 - Business Systems - used to record information about the operational business systems such as procurements, sales, human resources and finance.
# * cocoMDS6 - Manufacturing - used by the warehouse, manufacturing and distribution teams.
# * cocoMDSx - Development - used by the development teams building new IT capablity.
# * cocoEDGE*i* - Manufacturing sensors edge node servers (many of them).
#
# These metadata servers will each be configured in later parts of this hands-on lab, but first there are decisons to be made about the platform that the servers will run on and how they will be connected together.
# ### Open Metadata and Governance (OMAG) Server Platforms
#
# The Coco Pharmaceuticals' metadata servers along with the governance servers that interact with them must be hosted on at least one OMAG Server Platform.
#
# The OMAG Server Platform is a single executable (application) that can be started from the command line or a script or as part of a pre-built container environment such as `docker-compose` or `kubernetes`.
#
# If you are running this notebook as part of an Egeria hands on lab then the server platforms you need are already started. Run the following command to check that the admin platform is running.
#
# %run common/environment-check.ipynb
# ----
# If one of the platforms is not running, follow [this link to set up and run the platform](https://egeria.odpi.org/open-metadata-resources/open-metadata-labs/). Once the platforms are running you are ready to proceed.
#
# ----
# Most of the metadata servers are pretty stable and can share an OMAG Server Platform.
# The data lake however requires a lot of active governance so Gary chooses to put all of the metadata and governance servers for the data lake on to their own platform. The development team requested that their infrastructure is completely separate from the operational systems,
# so they are given their own server platform. Finally each of the edge servers will run an OMAG Server Platform to support
# their metadata server.
#
# Figure 1 shows which metadata servers will sit in each platform.
#
# 
# > **Figure 1:** Coco Pharmaceuticals' OMAG Server Platforms
#
#
# The sensor edge node servers used to monitor the warehouse operation and manufacturing process each have their own platform and are not yet included in this notebook.
# ### Open Metadata Repository Cohorts
#
# Metadata servers communicate via open metadata repository cohorts. A server can become a member of none, one or many cohorts. Once it has joined a cohort it can exchange metadata with the other members of that cohort. So the cohorts define scopes of sharing.
#
# Gary decides to begin with three open metadata repository cohorts:
#
# * **cocoCohort** - The production cohort contains all of the metadata servers that are used to run, coordinate and govern the business.
# * **devCohort** - The development cohort where the development teams are building and testing new capablity. Much of their metadata describes the software components under construction and the governance of the software development lifecycle.
# * **iotCohort** - The IoT cohort used to manage the sensors and robots in the manufacturing systems. The metadata produced by the sensors and robots is only of interest to the manufactuing and governance team.
#
# Figure 2 shows which metadata servers belong to each cohort.
#
# 
# > **Figure 2:** Membership of Coco Pharmaceuticals' cohorts
#
# Below are the names of the three cohorts.
cocoCohort = "cocoCohort"
devCohort = "devCohort"
iotCohort = "iotCohort"
# At the heart of each cohort is an event topic. By default, ODPi Egeria uses [Apache Kafka](https://kafka.apache.org/) topics.
# The metadata servers will need to be configured with the host name and port where Kafka is running.
# The command below pulls the value from an environment variable called `eventBusURLroot` with a default value of
# `localhost:9092`. It is used in all of the server configuration documents to connect it to Kafka.
# +
eventBusURLroot = os.environ.get('eventBusURLroot', 'localhost:9092')
jsonContentHeader = {'content-type':'application/json'}
eventBusBody = {
"producer": {
"bootstrap.servers": eventBusURLroot
},
"consumer":{
"bootstrap.servers": eventBusURLroot
}
}
# -
# ## Access services
#
# [The Open Metadata Access Services (OMAS)](https://egeria.odpi.org/open-metadata-implementation/access-services/) provide domain-specific services for data tools, engines and platforms to integrate with open metadata.
#
# The table below shows which access services are needed by each server.
#
#
# | Access Service | cocoMDS1 | cocoMDS2 | cocoMDS3 | cocoMDS4 | cocoMDS5 | cocoMDS6 | cocoMDSx | cocoEDGE*i* |
# | :------------------- | :------: | :------: | :------: | :------: | :------: | :------: | :------: | :---------: |
# | asset-catalog | Yes | Yes | Yes | Yes | No | Yes | Yes | No |
# | asset-consumer | Yes | Yes | Yes | Yes | No | Yes | Yes | No |
# | asset-owner | Yes | Yes | Yes | No | No | Yes | Yes | No |
# | community-profile | Yes | Yes | Yes | Yes | No | Yes | Yes | No |
# | glossary-view | Yes | Yes | Yes | Yes | No | Yes | Yes | No |
# | ------------------- | ------ | ------ | ------ | ------ | ------ | ------ | ------ | ---------- |
# | data-science | No | No | Yes | Yes | No | Yes | Yes | No |
# | ------------------- | ------ | ------ | ------ | ------ | ------ | ------ | ------ | ---------- |
# | subject-area | No | Yes | Yes | Yes | No | Yes | Yes | No |
# | ------------------- | ------ | ------ | ------ | ------ | ------ | ------ | ------ | ---------- |
# | governance-program | No | Yes | No | No | No | No | No | No |
# | data-privacy | No | Yes | No | No | No | No | No | No |
# | security-officer | No | Yes | No | No | No | No | No | No |
# | asset-lineage | No | Yes | No | No | No | No | No | No |
# | -------------------- | ------ | ------ | ------ | ------ | ------ | ------ | ------ | ---------- |
# | discovery-engine | Yes | No | Yes | No | No | Yes | Yes | No |
# | stewardship-action | Yes | No | Yes | No | No | Yes | Yes | No |
# | -------------------- | ------ | ------ | ------ | ------ | ------ | ------ | ------ | ---------- |
# | data-engine | Yes | No | No | No | No | Yes | No | Yes |
# | data-manager | Yes | No | No | No | No | Yes | No | Yes |
# | governance-engine | Yes | No | No | No | No | Yes | No | No |
# | information-view | Yes | No | No | No | No | Yes | No | No |
# | -------------------- | ------ | ------ | ------ | ------ | ------ | ------ | ------ | ---------- |
# | it-infrastructure | No | Yes | No | No | No | Yes | Yes | No |
# | project-management | No | Yes | Yes | No | No | Yes | Yes | No |
# | -------------------- | ------ | ------ | ------ | ------ | ------ | ------ | ------ | ---------- |
# | software-developer | No | No | No | No | No | No | Yes | No |
# | devops | No | No | No | No | No | No | Yes | No |
# | digital-architecture | No | No | No | No | No | No | Yes | No |
# | design-model | No | No | No | No | No | No | Yes | No |
# | -------------------- | ------ | ------ | ------ | ------ | ------ | ------ | ------ | ---------- |
#
#
# ## ODPi Egeria Server Configuration Overview
#
# Open metadata servers are configured using REST API calls to an OMAG Server Platform. Each call either defines a default value or configures a service that must run within the server when it is started.
#
# As each configuration call is made, the OMAG Server Platform builds up a [configuration document](https://egeria.odpi.org/open-metadata-implementation/admin-services/docs/concepts/configuration-document.html) with the values passed. When the configuration is finished, the configuration document will have all of the information needed to start the server.
#
# The configuration document will then be deployed with the OMAG Server Platform that is to host the server. When a request is made to this OMAG Server Platform to start the server, it reads the configuration document and initializes the server with the appropriate services.
#
# ## Configuration Set Up
#
# A server can be configured by any OMAG Server Platform - it does not have to be the same platform where the server will run. For this hands on lab we will use the development team's OMAG Server Platform to create the servers' configuration documents and then deploy them to the platforms where they will run.
adminPlatformURL = devPlatformURL
# The URLs for the configuration REST APIs have a common structure and begin with the following root:
adminCommandURLRoot = adminPlatformURL + '/open-metadata/admin-services/users/' + adminUserId + '/servers/'
# Many of Coco Pharmaceuticals' metadata servers need a local repository to store metadata about the data and processing occuring in the data lake.
#
# Egeria includes two types of repositories natively. One is an **in-memory repository** that stores metadata in hash maps. It is useful for demos and testing because a restart of the server results in an empty metadata repository. However, if you need metadata to persist from one run of the server to the next, you should use the **local graph repository**.
#
# The choice of local repository is made by specifying the local repository mode. The variables below show the two options. The `metadataRepositoryType` identfies which one is going to be used in the configuration.
# +
inMemoryRepositoryOption = "in-memory-repository"
graphRepositoryOption = "local-graph-repository"
metadataRepositoryType = inMemoryRepositoryOption
# -
# Egeria supports instance based security. These checks can be customized through an **Open Metadata Security Verifier**. Coco Pharaceuticals have written their own connector to support the specific rules of their industry. The Connection definition below tells a server how to load this connector. It needs to be included in each metadata server's configuration document.
serverSecurityConnectionBody = {
"class": "Connection",
"connectorType": {
"class": "ConnectorType",
"connectorProviderClassName": "org.odpi.openmetadata.metadatasecurity.samples.CocoPharmaServerSecurityProvider"
}
}
# Finally, to ensure that a caller can not request too much metadata in a single request, it is possible to set a maximum page size for requests that return a list of items. The maximum page size puts a limit on the number of items that can be requested. The variable below defines the value that will be added to the configuration document for each server.
maxPageSize = '100'
# ## Configuring cocoMDS1 - Data Lake Operations metadata server
#
# This section configures the `cocoMDS1` server. The server name is passed on every configuration call to identify which configuration document to update with the new configuration. The configuration document is created automatically on first use.
# +
mdrServerName = "cocoMDS1"
mdrServerUserId = "cocoMDS1npa"
mdrServerPassword = "<PASSWORD>"
mdrServerPlatform = dataLakePlatformURL
metadataCollectionName = "Data Lake Catalog"
print("Configuring " + mdrServerName + "...")
configurePlatformURL(adminPlatformURL, adminUserId, mdrServerName, mdrServerPlatform)
configureMaxPageSize(adminPlatformURL, adminUserId, mdrServerName, maxPageSize)
clearServerType(adminPlatformURL, adminUserId, mdrServerName)
configureOwningOrganization(adminPlatformURL, adminUserId, mdrServerName, organizationName)
configureUserId(adminPlatformURL, adminUserId, mdrServerName, mdrServerUserId)
configurePassword(adminPlatformURL, adminUserId, mdrServerName, mdrServerPassword)
configureSecurityConnection(adminPlatformURL, adminUserId, mdrServerName, serverSecurityConnectionBody)
configureEventBus(adminPlatformURL, adminUserId, mdrServerName, eventBusBody)
configureMetadataRepository(adminPlatformURL, adminUserId, mdrServerName, metadataRepositoryType)
configureDescriptiveName(adminPlatformURL, adminUserId, mdrServerName, metadataCollectionName)
configureCohortMembership(adminPlatformURL, adminUserId, mdrServerName, cocoCohort)
print("\nConfiguring " + mdrServerName + " Access Services (OMAS)...")
accessServiceOptions = {
"SupportedZones": ["quarantine", "clinical-trials", "research", "data-lake", "trash-can"]
}
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'asset-catalog', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'asset-consumer', accessServiceOptions)
accessServiceOptions["DefaultZones"] = [ "quarantine" ]
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'asset-owner', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'community-profile', {"KarmaPointPlateau":"500"})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'glossary-view', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'discovery-engine', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'stewardship-action', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'data-engine', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'data-manager', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'governance-engine', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'information-view', accessServiceOptions)
print("\nDone.")
# -
# ----
#
# ## Configuring cocoMDS2 - Governance metadata server
#
# This section configures the `cocoMDS2` server. This server is configured in a similar way to cocoMDS1 except that is has different Open Metadata Access Services (OMASs) enabled and it joins all of the cohorts.
#
# The code below covers the basic set up of the server properties, security, event bus and local repository.
# +
mdrServerName = "cocoMDS2"
mdrServerUserId = "cocoMDS2npa"
mdrServerPassword = "<PASSWORD>"
mdrServerPlatform = corePlatformURL
metadataCollectionName = "Governance Catalog"
print("Configuring " + mdrServerName + "...")
configurePlatformURL(adminPlatformURL, adminUserId, mdrServerName, mdrServerPlatform)
configureMaxPageSize(adminPlatformURL, adminUserId, mdrServerName, maxPageSize)
clearServerType(adminPlatformURL, adminUserId, mdrServerName)
configureOwningOrganization(adminPlatformURL, adminUserId, mdrServerName, organizationName)
configureUserId(adminPlatformURL, adminUserId, mdrServerName, mdrServerUserId)
configurePassword(adminPlatformURL, adminUserId, mdrServerName, mdrServerPassword)
configureSecurityConnection(adminPlatformURL, adminUserId, mdrServerName, serverSecurityConnectionBody)
configureEventBus(adminPlatformURL, adminUserId, mdrServerName, eventBusBody)
configureMetadataRepository(adminPlatformURL, adminUserId, mdrServerName, metadataRepositoryType)
configureDescriptiveName(adminPlatformURL, adminUserId, mdrServerName, metadataCollectionName)
# Note: cohort membership is configured for all of the cohorts here
configureCohortMembership(adminPlatformURL, adminUserId, mdrServerName, cocoCohort)
configureCohortMembership(adminPlatformURL, adminUserId, mdrServerName, devCohort)
configureCohortMembership(adminPlatformURL, adminUserId, mdrServerName, iotCohort)
print("\nConfiguring " + mdrServerName + " Access Services (OMAS)...")
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'asset-catalog', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'asset-consumer', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'asset-owner', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'community-profile', {"KarmaPointPlateau":"500"})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'glossary-view', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'subject-area', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'governance-program', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'data-privacy', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'security-officer', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'asset-lineage', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'it-infrastructure', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'project-management', {})
print("\nDone.")
# -
# ----
#
# ## Configuring cocoMDS3 - Research
#
# Server cocoMDS3 is used by the research teams who are developing new treatments.
# These teams are working with their own assets as well as assets coming from the data lake.
# They have their own discovery server to automate analysis of their data.
# They are also creating new data science models that they run by hand, or deploy to the discovery server.
# +
mdrServerName = "cocoMDS3"
mdrServerUserId = "cocoMDS3npa"
mdrServerPassword = "<PASSWORD>"
mdrServerPlatform = corePlatformURL
metadataCollectionName = "Research Catalog"
print("Configuring " + mdrServerName + "...")
configurePlatformURL(adminPlatformURL, adminUserId, mdrServerName, mdrServerPlatform)
configureMaxPageSize(adminPlatformURL, adminUserId, mdrServerName, maxPageSize)
clearServerType(adminPlatformURL, adminUserId, mdrServerName)
configureOwningOrganization(adminPlatformURL, adminUserId, mdrServerName, organizationName)
configureUserId(adminPlatformURL, adminUserId, mdrServerName, mdrServerUserId)
configurePassword(adminPlatformURL, adminUserId, mdrServerName, mdrServerPassword)
configureSecurityConnection(adminPlatformURL, adminUserId, mdrServerName, serverSecurityConnectionBody)
configureEventBus(adminPlatformURL, adminUserId, mdrServerName, eventBusBody)
configureMetadataRepository(adminPlatformURL, adminUserId, mdrServerName, metadataRepositoryType)
configureDescriptiveName(adminPlatformURL, adminUserId, mdrServerName, metadataCollectionName)
configureCohortMembership(adminPlatformURL, adminUserId, mdrServerName, cocoCohort)
print("\nConfiguring " + mdrServerName + " Access Services (OMAS)...")
accessServiceOptions = {
"SupportedZones": ["personal-files", "clinical-trials", "research", "data-lake", "trash-can"]
}
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'asset-catalog', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'asset-consumer', accessServiceOptions)
accessServiceOptions["DefaultZones"] = [ "personal-files" ]
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'asset-owner', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'community-profile', {"KarmaPointPlateau":"500"})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'glossary-view', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'data-science', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'subject-area', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'discovery-engine', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'stewardship-action', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'project-management', accessServiceOptions)
print("\nDone.")
# -
# ----
# ## Configuring cocoMDS4 - Data Lake Users
#
# Server cocoMDS4 used by general business users and the executive team to access data from the data lake.
# It does not have a repository of its own. Instead it issues federated queries to the other repositories in the `cocoCohort`.
# +
mdrServerName = "cocoMDS4"
mdrServerUserId = "cocoMDS4npa"
mdrServerPassword = "<PASSWORD>"
mdrServerPlatform = dataLakePlatformURL
metadataCollectionName = "Data Lake Catalog"
print("Configuring " + mdrServerName + "...")
configurePlatformURL(adminPlatformURL, adminUserId, mdrServerName, mdrServerPlatform)
configureMaxPageSize(adminPlatformURL, adminUserId, mdrServerName, maxPageSize)
clearServerType(adminPlatformURL, adminUserId, mdrServerName)
configureOwningOrganization(adminPlatformURL, adminUserId, mdrServerName, organizationName)
configureUserId(adminPlatformURL, adminUserId, mdrServerName, mdrServerUserId)
configurePassword(adminPlatformURL, adminUserId, mdrServerName, mdrServerPassword)
configureSecurityConnection(adminPlatformURL, adminUserId, mdrServerName, serverSecurityConnectionBody)
configureEventBus(adminPlatformURL, adminUserId, mdrServerName, eventBusBody)
# Note: no metadata repository or collection configuration here
configureCohortMembership(adminPlatformURL, adminUserId, mdrServerName, cocoCohort)
print("\nConfiguring " + mdrServerName + " Access Services (OMAS)...")
accessServiceOptions = {
"SupportedZones": [ "data-lake" ]
}
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'asset-catalog', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'asset-consumer', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'community-profile', {"KarmaPointPlateau":"500"})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'glossary-view', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'data-science', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'subject-area', {})
print("\nDone.")
# -
# ----
#
# ## Configuring cocoMDS5 - Business Systems
#
# Server cocoMDS5 is a repository proxy to an ETL tool called `iisCore01`. This ETL tool is well established in Coco Pharmaceuticals and has a built-in metadata repository that contains information about their operational business systems such as procurement, sales, human resources and finance.
#
# This ETL tool has its own user interface and services so the OMASs are not enabled.
# +
mdrServerName = "cocoMDS5"
mdrServerUserId = "cocoMDS5npa"
mdrServerPassword = "<PASSWORD>"
mdrServerPlatform = corePlatformURL
metadataCollectionName = "Business Systems Catalog"
print("Configuring " + mdrServerName + "...")
configurePlatformURL(adminPlatformURL, adminUserId, mdrServerName, mdrServerPlatform)
configureMaxPageSize(adminPlatformURL, adminUserId, mdrServerName, maxPageSize)
clearServerType(adminPlatformURL, adminUserId, mdrServerName)
configureOwningOrganization(adminPlatformURL, adminUserId, mdrServerName, organizationName)
configureUserId(adminPlatformURL, adminUserId, mdrServerName, mdrServerUserId)
configurePassword(adminPlatformURL, adminUserId, mdrServerName, mdrServerPassword)
configureSecurityConnection(adminPlatformURL, adminUserId, mdrServerName, serverSecurityConnectionBody)
configureEventBus(adminPlatformURL, adminUserId, mdrServerName, eventBusBody)
configureRepositoryProxyDetails(adminPlatformURL, adminUserId, mdrServerName, "org.odpi.openmetadata.adapters.repositoryservices.readonly.repositoryconnector.ReadOnlyOMRSRepositoryConnectorProvider")
configureDescriptiveName(adminPlatformURL, adminUserId, mdrServerName, metadataCollectionName)
configureCohortMembership(adminPlatformURL, adminUserId, mdrServerName, cocoCohort)
# Note: no access service configuration here
# Still need to add startup Archive
print("\nDone.")
# -
# ----
#
# ## Configuring cocoMDS6 - Manufacturing
#
# Server cocoMDS6 is the repository server used by the warehouse, manufacturing and distribution teams. It supports the systems for this part of the organization and acts as a hub for monitoring the IoT environment.
# +
mdrServerName = "cocoMDS6"
mdrServerUserId = "cocoMDS6npa"
mdrServerPassword = "<PASSWORD>"
mdrServerPlatform = corePlatformURL
metadataCollectionName = "Manufacturing Catalog"
print("Configuring " + mdrServerName + "...")
configurePlatformURL(adminPlatformURL, adminUserId, mdrServerName, mdrServerPlatform)
configureMaxPageSize(adminPlatformURL, adminUserId, mdrServerName, maxPageSize)
clearServerType(adminPlatformURL, adminUserId, mdrServerName)
configureOwningOrganization(adminPlatformURL, adminUserId, mdrServerName, organizationName)
configureUserId(adminPlatformURL, adminUserId, mdrServerName, mdrServerUserId)
configurePassword(adminPlatformURL, adminUserId, mdrServerName, mdrServerPassword)
configureSecurityConnection(adminPlatformURL, adminUserId, mdrServerName, serverSecurityConnectionBody)
configureEventBus(adminPlatformURL, adminUserId, mdrServerName, eventBusBody)
configureMetadataRepository(adminPlatformURL, adminUserId, mdrServerName, metadataRepositoryType)
configureDescriptiveName(adminPlatformURL, adminUserId, mdrServerName, metadataCollectionName)
configureCohortMembership(adminPlatformURL, adminUserId, mdrServerName, cocoCohort)
configureCohortMembership(adminPlatformURL, adminUserId, mdrServerName, iotCohort)
print("\nConfiguring " + mdrServerName + " Access Services (OMAS)...")
accessServiceOptions = {
"SupportedZones": [ "manufacturing" ],
"DefaultZones" : [ "manufacturing"]
}
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'asset-catalog', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'asset-consumer', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'asset-owner', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'community-profile', {"KarmaPointPlateau":"500"})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'glossary-view', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'data-science', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'subject-area', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'discovery-engine', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'stewardship-action', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'data-engine', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'data-manager', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'governance-engine', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'information-view', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'it-infrastructure', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'project-management', accessServiceOptions)
print("\nDone.")
# -
# ----
#
# ## Configuring cocoMDSx - Development
#
# Server cocoMDSx is used by the development teams building new IT capablity. It will hold all of the software component assets and servers used for development and devOps. The development teams have their own OMAG Server Platform and cohort called 'devCohort'.
# +
mdrServerName = "cocoMDSx"
mdrServerUserId = "cocoMDSxnpa"
mdrServerPassword = "<PASSWORD>"
mdrServerPlatform = devPlatformURL
metadataCollectionName = "Development Catalog"
print("Configuring " + mdrServerName + "...")
configurePlatformURL(adminPlatformURL, adminUserId, mdrServerName, mdrServerPlatform)
configureMaxPageSize(adminPlatformURL, adminUserId, mdrServerName, maxPageSize)
clearServerType(adminPlatformURL, adminUserId, mdrServerName)
configureOwningOrganization(adminPlatformURL, adminUserId, mdrServerName, organizationName)
configureUserId(adminPlatformURL, adminUserId, mdrServerName, mdrServerUserId)
configurePassword(adminPlatformURL, adminUserId, mdrServerName, mdrServerPassword)
configureSecurityConnection(adminPlatformURL, adminUserId, mdrServerName, serverSecurityConnectionBody)
configureEventBus(adminPlatformURL, adminUserId, mdrServerName, eventBusBody)
configureMetadataRepository(adminPlatformURL, adminUserId, mdrServerName, metadataRepositoryType)
configureDescriptiveName(adminPlatformURL, adminUserId, mdrServerName, metadataCollectionName)
configureCohortMembership(adminPlatformURL, adminUserId, mdrServerName, devCohort)
print("\nConfiguring " + mdrServerName + " Access Services (OMAS)...")
accessServiceOptions = {
"SupportedZones": [ "sdlc" ],
"DefaultZones": [ "sdlc" ]
}
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'asset-catalog', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'asset-consumer', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'asset-owner', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'community-profile', {"KarmaPointPlateau":"500"})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'glossary-view', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'data-science', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'subject-area', {})
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'discovery-engine', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'stewardship-action', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'it-infrastructure', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'project-management', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'software-developer', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'devops', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'digital-architecture', accessServiceOptions)
configureAccessService(adminPlatformURL, adminUserId, mdrServerName, 'design-model', accessServiceOptions)
print("\nDone.")
# -
# ----
# ## Configuring the Discovery Server
#
# The discovery server is a special kind of governance server that hosts one or more discovery engines. A discovery engine is a set of discovery services that analyzes the content of an organization's assets. For example, a discovery service may open up a data set and assess the quality of the data inside.
#
# The result of a discovery service's analysis is stored in a metadata server as a discovery analysis report that is chained off of the asset's definition. This report can be retrieved either through the discovery server's API or through the metadata server's APIs, specifically the Discovery Engine OMAS and the Asset Owner OMAS.
#
# The behavior of the discovery server, its discovery engines and the discovery services within are defined in the [Open Discovery Framework (ODF)](https://egeria.odpi.org/open-metadata-implementation/frameworks/open-discovery-framework/). This framework enables new implementations of discovery services to be deployed to the discovery engines.
#
# Typically a discovery server is deployed close to where the data is stored because it can generate a lot of network
# traffic when it is retrieving all of the content of an asset.
#
# Coco Pharmaceuticals runs one discovery server for its data lake. It is called `findItDL01` and it runs on the This discovery server has three discovery engines running on it:
#
# * **AssetDiscovery** - extracts metadata about different types of assets on request.
# * **AssetDeduplicator** - detects and reports on asset definitions in the open metadata repositories that seem to be
# duplicate descriptions of the same physical asset.
# * **AssetQuality** - assesses the quality of the content of assets on request.
#
# The commands below configure the discovery server. The definitions of the discovery engines and their services
# are retrieved from the `cocoMDS1` metadata server through its Discovery Engine OMAS.
#
# +
discoServerName = "findItDL01"
discoServerPlatform = dataLakePlatformURL
discoServerUserId = "findItDL01npa"
discoServerPassword = "<PASSWORD>"
mdrServerName = "cocoMDS1"
mdrServerPlatform = dataLakePlatformURL
print("Configuring " + discoServerName + "...")
configurePlatformURL(adminPlatformURL, adminUserId, discoServerName, discoServerPlatform)
configureMaxPageSize(adminPlatformURL, adminUserId, discoServerName, maxPageSize)
clearServerType(adminPlatformURL, adminUserId, mdrServerName)
configureOwningOrganization(adminPlatformURL, adminUserId, discoServerName, organizationName)
configureUserId(adminPlatformURL, adminUserId, discoServerName, discoServerUserId)
configurePassword(adminPlatformURL, adminUserId, discoServerName, discoServerPassword)
configureSecurityConnection(adminPlatformURL, adminUserId, discoServerName, serverSecurityConnectionBody)
configureEventBus(adminPlatformURL, adminUserId, discoServerName, eventBusBody)
configureDefaultAuditLog(adminPlatformURL, adminUserId, discoServerName)
print("\nConfiguring " + discoServerName + " as a discovery server ...")
discoveryEngines = ["AssetDiscovery", "AssetDeduplicator", "AssetQuality"]
configureDiscoveryEngineServices(adminPlatformURL, adminUserId, discoServerName, mdrServerName, mdrServerPlatform, discoveryEngines)
# -
# ----
# # Configuring the View Server & services
# A new UI known as 'presentation server' allows coco's employees to understand more
# about their metadata environment
#
# This is an initial version of an example to configure the view services
# provided as an early sample in this release - this area is in development
#
# The configuration is likely to change, and this section of the notebook consolidated with our common functions
#
# A new UI is deployed in the k8s and docker-compose environments (nodeport 30091 in k8s, 18091 in compose )
#
# The tenant (coco in this case) must be explicitly provided in the URL, as must navigation to the login page
# For example in compose go to https://localhost:18091/coco/login
#
# Further docs will be added in future releases. Please use odpi.slack.com to get further help
# +
# Common functions
def configureGovernanceSolutionViewService(adminPlatformURL, adminUserId, viewServerName, viewService, remotePlatformURL,remoteServerName):
adminCommandURLRoot = adminPlatformURL + '/open-metadata/admin-services/users/' + adminUserId + '/servers/'
print (" ... configuring the " + viewService + " Governance Solution View Service for this server...")
url = adminCommandURLRoot + viewServerName + '/view-services/' + viewService
jsonContentHeader = {'content-type':'application/json'}
viewBody = {
"class": "ViewServiceConfig",
"omagserverPlatformRootURL": remotePlatformURL,
"omagserverName" : remoteServerName
}
postAndPrintResult(url, json=viewBody, headers=jsonContentHeader)
def configureIntegrationViewService(adminPlatformURL, adminUserId, viewServerName, viewService, configBody):
adminCommandURLRoot = adminPlatformURL + '/open-metadata/admin-services/users/' + adminUserId + '/servers/'
print (" ... configuring the " + viewService + " Integration View Service for this server...")
url = adminCommandURLRoot + viewServerName + '/view-services/' + viewService
jsonContentHeader = {'content-type':'application/json'}
postAndPrintResult(url, json=configBody, headers=jsonContentHeader)
# A view server supports the presentation server UI (a node based app). Here we run it on the datalake platform
viewServerName = "cocoView1"
viewServerUserId = "cocoView1npa"
viewServerPassword = "<PASSWORD>"
viewServerPlatform = dataLakePlatformURL
viewServerType = "View Server"
# Configuration is similar to most servers
print("Configuring " + viewServerName + "...")
configurePlatformURL(adminPlatformURL, adminUserId, viewServerName, viewServerPlatform)
configureMaxPageSize(adminPlatformURL, adminUserId, mdrServerName, maxPageSize)
clearServerType(adminPlatformURL, adminUserId, viewServerName)
configureServerType(adminPlatformURL,adminUserId,viewServerName,viewServerType)
configureOwningOrganization(adminPlatformURL, adminUserId, viewServerName, organizationName)
configureUserId(adminPlatformURL, adminUserId, viewServerName, viewServerUserId)
configurePassword(adminPlatformURL, adminUserId, viewServerName, viewServerPassword)
configureSecurityConnection(adminPlatformURL, adminUserId, viewServerName, serverSecurityConnectionBody)
configureEventBus(adminPlatformURL, adminUserId, viewServerName, eventBusBody)
configureDefaultAuditLog(adminPlatformURL, adminUserId, viewServerName)
# The governance solution view services currently only consist of glossary author
print ("Configuring the Governance Solution View Services")
remotePlatformURL=dataLakePlatformURL
remoteServerName="cocoMDS4"
viewService="glossary-author"
configureGovernanceSolutionViewService(adminPlatformURL, adminUserId, viewServerName, viewService, remotePlatformURL,remoteServerName)
print ("Configuring the Integration View Services")
# repository explorer integration view service
viewService="rex"
rexConfigBody = {
"class":"IntegrationViewServiceConfig",
"viewServiceAdminClass":"org.odpi.openmetadata.viewservices.rex.admin.RexViewAdmin",
"viewServiceFullName":"Repository Explorer",
"viewServiceOperationalStatus":"ENABLED",
"omagserverPlatformRootURL": "UNUSED",
"omagserverName" : "UNUSED",
"resourceEndpoints" : [
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Platform",
"description" : "Core Platform",
"platformName" : "Core",
"platformRootURL" : corePlatformURL
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Platform",
"description" : "DataLake Platform",
"platformName" : "DataLake",
"platformRootURL" : dataLakePlatformURL
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Platform",
"description" : "Development Platform",
"platformName" : "Development",
"platformRootURL" : devPlatformURL
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDS1",
"description" : "Data Lake Operations",
"platformName" : "DataLake",
"serverName" : "cocoMDS1"
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDS2",
"description" : "Governance",
"platformName" : "Core",
"serverName" : "cocoMDS2"
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDS3",
"description" : "Research",
"platformName" : "Core",
"serverName" : "cocoMDS3"
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDS5",
"description" : "Business Systems",
"platformName" : "Core",
"serverName" : "cocoMDS5"
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDS6",
"description" : "Manufacturing",
"platformName" : "Core",
"serverName" : "cocoMDS6"
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDSx",
"description" : "Development",
"platformName" : "Development",
"serverName" : "cocoMDSx"
},
]
}
configureIntegrationViewService(adminPlatformURL, adminUserId, viewServerName, viewService, rexConfigBody)
# type-explorer has endpoints
viewService="tex"
texConfigBody = {
"class":"IntegrationViewServiceConfig",
"viewServiceAdminClass":"org.odpi.openmetadata.viewservices.tex.admin.TexViewAdmin",
"viewServiceFullName":"Type Explorer",
"viewServiceOperationalStatus":"ENABLED",
"omagserverPlatformRootURL": "UNUSED",
"omagserverName" : "UNUSED",
"resourceEndpoints" : [
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Platform",
"description" : "Core Platform",
"platformName" : "Core",
"platformRootURL" : corePlatformURL
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Platform",
"description" : "DataLake Platform",
"platformName" : "DataLake",
"platformRootURL" : dataLakePlatformURL
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Platform",
"description" : "Development Platform",
"platformName" : "Development",
"platformRootURL" : devPlatformURL
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDS1",
"description" : "Data Lake Operations",
"platformName" : "DataLake",
"serverName" : "cocoMDS1"
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDS2",
"description" : "Governance",
"platformName" : "Core",
"serverName" : "cocoMDS2"
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDS3",
"description" : "Research",
"platformName" : "Core",
"serverName" : "cocoMDS3"
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDS5",
"description" : "Business Systems",
"platformName" : "Core",
"serverName" : "cocoMDS5"
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDS6",
"description" : "Manufacturing",
"platformName" : "Core",
"serverName" : "cocoMDS6"
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDSx",
"description" : "Development",
"platformName" : "Development",
"serverName" : "cocoMDSx"
},
]
}
configureIntegrationViewService(adminPlatformURL, adminUserId, viewServerName, viewService, texConfigBody)
# Dino provides insight into the operational environment of egeria - this config body allows coco's platforms & servers to be accessed
viewService="dino"
DinoConfigBody = {
"class":"IntegrationViewServiceConfig",
"viewServiceAdminClass":"org.odpi.openmetadata.viewservices.dino.admin.DinoViewAdmin",
"viewServiceFullName":"Dino",
"viewServiceOperationalStatus":"ENABLED",
"omagserverPlatformRootURL": "UNUSED",
"omagserverName" : "UNUSED",
"resourceEndpoints" : [
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Platform",
"description" : "Core Platform",
"platformName" : "Core",
"platformRootURL" : corePlatformURL
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Platform",
"description" : "DataLake Platform",
"platformName" : "DataLake",
"platformRootURL" : dataLakePlatformURL
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Platform",
"description" : "Development Platform",
"platformName" : "Development",
"platformRootURL" : devPlatformURL
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDS1",
"description" : "Data Lake Operations",
"platformName" : "DataLake",
"serverName" : "cocoMDS1"
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDS2",
"description" : "Governance",
"platformName" : "Core",
"serverName" : "cocoMDS2"
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDS3",
"description" : "Research",
"platformName" : "Core",
"serverName" : "cocoMDS3"
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDS4",
"description" : "Data Lake Users",
"platformName" : "DataLake",
"serverName" : "cocoMDS4"
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDS5",
"description" : "Business Systems",
"platformName" : "Core",
"serverName" : "cocoMDS5"
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDS6",
"description" : "Manufacturing",
"platformName" : "Core",
"serverName" : "cocoMDS6"
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoMDSx",
"description" : "Development",
"platformName" : "Development",
"serverName" : "cocoMDSx"
},
{
"class" : "ResourceEndpointConfig",
"resourceCategory" : "Server",
"serverInstanceName" : "cocoView1",
"description" : "View Server",
"platformName" : "DataLake",
"serverName" : "cocoView1"
},
]
}
configureIntegrationViewService(adminPlatformURL, adminUserId, viewServerName, viewService, DinoConfigBody)
# -
# # Deploying server configuration
# The commands that have been issued so far have created a configuration document for each server.
# These configuration documents are currently local to the Development OMAG Server Platform where the
# adminstration commands were issued (figure 3).
#
# 
# > **Figure 3:** Creating configuration documents using administration commands
#
# If servers are to be started on the other server platforms then their configuration documents
# need to be deployed (copied) to these platforms (figure 4).
#
# 
# > **Figure 4:** Deploying configuration documents
#
# However, before deploying the configuration documents, the receiving OMAG Server Platforms
# need to be running.
#
# The code below checks the Core and Data Lake OMAG Server Platforms are running.
# +
print("\nChecking OMAG Server Platform availability...")
checkServerPlatform("Data Lake Platform", dataLakePlatformURL)
checkServerPlatform("Core Platform", corePlatformURL)
checkServerPlatform("Dev Platform", devPlatformURL)
print ("\nDone.")
# -
# ----
# Make sure the each of the platforms is running.
#
# ----
# The commands below deploy the server configuration documents to the server platforms where the
# servers will run.
# +
print("\nDeploying server configuration documents to appropriate platforms...")
deployServerToPlatform(adminPlatformURL, adminUserId, "cocoMDS1", dataLakePlatformURL)
deployServerToPlatform(adminPlatformURL, adminUserId, "cocoMDS2", corePlatformURL)
deployServerToPlatform(adminPlatformURL, adminUserId, "cocoMDS3", corePlatformURL)
deployServerToPlatform(adminPlatformURL, adminUserId, "cocoMDS4", dataLakePlatformURL)
deployServerToPlatform(adminPlatformURL, adminUserId, "cocoMDS5", corePlatformURL)
deployServerToPlatform(adminPlatformURL, adminUserId, "cocoMDS6", corePlatformURL)
deployServerToPlatform(adminPlatformURL, adminUserId, "cocoMDSx", devPlatformURL)
deployServerToPlatform(adminPlatformURL, adminUserId, "findItDL01", dataLakePlatformURL)
deployServerToPlatform(adminPlatformURL, adminUserId, "cocoView1", dataLakePlatformURL)
print("\nDone.")
# -
# ----
| open-metadata-resources/open-metadata-labs/egeria-server-config.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="P_9IavYxfqKF"
# # 12. Machine learning techniques
# + id="W1qnFjBIftmC" outputId="42e40d7b-451d-4573-d1ff-d544a4e34df2" colab={"base_uri": "https://localhost:8080/"}
# !git clone https://github.com/s7s/machine_learning_1.git
# %cd machine_learning_1/ML_in_practice
# + id="m3nP5yDNfqKI"
import random as rd
rd.seed(0)
# + id="bAbJWz2MfqKH"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# + [markdown] id="E8GM6HeSfqKI"
# ## 12.1 Loading and exploring the dataset
#
# First, we use pandas to load the dataset from a csv file.
# + id="FFuOhKGrfqKJ" outputId="96eb755e-f26e-4ec3-bc63-c7e315ce483e" colab={"base_uri": "https://localhost:8080/", "height": 417}
# use pabdas to read './titanic.csv'
raw_data = pd.read_csv('./titanic.csv',sep=',')
raw_data
# + [markdown] id="P8cSfkeYfqKL"
# Next, we can explore the dataset.
# + id="B9A_Cb6afqKL" outputId="d62c81e0-604e-486d-cf03-5943d5fe350e" colab={"base_uri": "https://localhost:8080/"}
# Use pandas to examine the length of the dataset
raw_data.shape
# + id="SwGiRZ6FfqKM" outputId="d5513ebd-adfb-4d47-f88e-e95021acc7a0" colab={"base_uri": "https://localhost:8080/"}
# Use pandas to examine the columns in the dataset
raw_data.columns
# + id="-pwL3iZNfqKN" outputId="50813215-3286-45b7-8dc8-02d0650d56c1" colab={"base_uri": "https://localhost:8080/"}
# Use pandas to examine "survived" column (labels)
raw_data['Survived']
# + id="eT6BD83VfqKO" outputId="1fa83b3a-4343-4e0a-e515-e002d836f917" colab={"base_uri": "https://localhost:8080/", "height": 417}
# Use pandas to exanine more than one column at the same time []
raw_data[["Name", "Age"]]
# + id="87L5P5_GfqKN" outputId="fe48299a-fe2f-410b-9cfb-b35b6df7efb8" colab={"base_uri": "https://localhost:8080/"}
# Use pandas to check how many passengers survived
sum(raw_data['Survived'])
# + [markdown] id="IKYLTXINfqKO"
# ## 12.2. Cleaning up the data
#
# Now, let's look at how many columns have missing data
# + id="9azVLEVtfqKO" outputId="dd3705e2-7d01-4400-e9e4-77a96bfb8279" colab={"base_uri": "https://localhost:8080/"}
# use pandas to check missing data (NA(not available) values) for all the columns
raw_data.isnull().sum()
# + [markdown] id="uXFTGpN5fqKP"
# The Cabin column is missing too many values to be useful. Let's drop it altogether.
# + id="A49SL-0NfqKQ"
# Use pandas to drop "Cabin" column
clean_data = raw_data.drop(columns='Cabin',axis=1)
# + id="2227vwAofqKQ" outputId="f593d25b-d212-4ee6-d5fd-b9a0dec4d651" colab={"base_uri": "https://localhost:8080/", "height": 417}
clean_data
# + [markdown] id="wDdfjznYfqKQ"
# Other columns such as Age or Embarked are missing some values, but they can still be useful.
#
# For the age column, let's fill in the missing values with the median of all ages.
#
# For the Embarked column, let's make a new category called 'U', for Unknown port of embarkment.
# + id="VesqtzGsfqKR" outputId="b44b7ee1-bbd8-4489-dcf0-db26e29ea723" colab={"base_uri": "https://localhost:8080/"}
# get the median of age column using pandas
median_age = raw_data['Age'].median()
median_age
# + id="MVksNV43fqKR"
# use pandas to fill the na values in age column with the median age
clean_data["Age"] =raw_data["Age"].fillna(median_age)
# + id="cf_Z2CzvfqKR"
# use pandas to fill the na values in embarked column with 'U'
clean_data["Embarked"] = raw_data['Embarked'].fillna('U')
# + id="wAL-CsGyfqKR" outputId="81d7afc5-66d5-4caf-a7a8-9b42fcea8a4e" colab={"base_uri": "https://localhost:8080/"}
clean_data.isna().sum()
# + id="Ks6LMctufqKS" outputId="eff0b913-444c-4e6a-ec23-52660464fa73" colab={"base_uri": "https://localhost:8080/", "height": 202}
# view 10 rows of the clean dataset
clean_data.head()
# + [markdown] id="FUfU0glCfqKS"
# ### 12.2.3 Saving our data for the future
# + id="WtAY8WYTfqKS"
# save the clean dataset to './clean_titanic_data.csv'
clean_data.to_csv('./clean_titanic_data1.csv',sep=',',index=None)
# + [markdown] id="9Y_4pk_RfqKS"
# ## 12.3 Manipulating the features
#
# - One-hot encoding
# - Binning
# - Feature selection
#
# ### 12.3.1 One-hot encoding
# + id="a8pojgwefqKS" outputId="059fd061-211b-453a-a1d7-ef2294d445fc" colab={"base_uri": "https://localhost:8080/", "height": 415}
preprocessed_data = pd.read_csv('clean_titanic_data1.csv')
preprocessed_data
# + id="ZivvKrlcfqKS"
# Use pandas method .get_dummies() to get the one hot encoding of “embarked”, “pclass” and “gender”
preprocessed_data=pd.get_dummies(preprocessed_data, columns = ['Embarked', 'Pclass', 'Sex'])
# Use pandas method .drop() to remove the old columns and method .concat() to add the new columns
# + id="NwEwCHPFfqKT" outputId="defb60d9-9964-4511-ea93-50f5f37c4af7" colab={"base_uri": "https://localhost:8080/"}
preprocessed_data.columns
# + [markdown] id="cV-AbnysfqKU"
# ### 12.3.2 Binning
# + id="gThz2CLlfqKU" outputId="6f391183-45a3-413a-b4d7-ae2c09f32be5" colab={"base_uri": "https://localhost:8080/", "height": 554}
bins = [0, 10, 20, 30, 40, 50, 60, 70, 80]
# Use .cut() method to make bins from the age column
categorized_age =pd.cut(x=preprocessed_data['Age'], bins=[0, 10, 20, 30, 40, 50, 60, 70, 80])
preprocessed_data['Categorized_age'] = categorized_age
preprocessed_data = preprocessed_data.drop(["Age"], axis=1)
preprocessed_data
# + id="nBtwwk_ofqKU" outputId="2ea32f94-01a5-4be8-9724-b126a09299fc" colab={"base_uri": "https://localhost:8080/", "height": 937}
# Use pandas method .get_dummies() to get the one hot encoding of “Categorized_age”
# Use pandas method .drop() to remove the old column and method .concat() to add the new columns
preprocessed_data=pd.get_dummies(preprocessed_data, columns = ['Categorized_age'])
preprocessed_data
# + [markdown] id="M8251vH3fqKV"
# ### 12.3.4 Feature selection
# + id="Z_SErhJ9fqKV" outputId="d7b27c83-39ae-4b80-98dd-2a8690b8546b" colab={"base_uri": "https://localhost:8080/", "height": 239}
# drop these columns['Name', 'Ticket', 'PassengerId']
preprocessed_data=preprocessed_data.drop(columns=['Name', 'Ticket', 'PassengerId'],axis=1)
preprocessed_data.head()
# + [markdown] id="mw-TKDt1fqKV"
# ### 12.3.5 Saving for future use
# + id="MzlAdRmffqKW"
preprocessed_data.to_csv('./preprocessed_titanic_data.csv', index=None)
# + [markdown] id="bS3XGQrXfqKW"
# # 12.4 Training models
# + id="bnmsC1G7fqKW" outputId="0bfb5a51-ae84-46df-ff77-9389281a9c06" colab={"base_uri": "https://localhost:8080/", "height": 239}
data = pd.read_csv('./preprocessed_titanic_data.csv')
data.head()
# + [markdown] id="akXchKUDfqKW"
# ### 12.4.1 Features-labels split and train-validation split
# + id="_lzo2QNufqKX"
# drop ["Survived"] column and save that to features
features = data.drop(columns=['Survived'])
# save the ["Survived"] column to labels
labels = data['Survived']
# + id="sf5D_149oK5F" outputId="34b31bf9-e6df-46da-cbff-25585c930521" colab={"base_uri": "https://localhost:8080/", "height": 452}
features
# + id="U-gIWUBhfqKY"
from sklearn.model_selection import train_test_split
# + id="n-gnQzTvooqW"
# + id="p3CZd8UGfqKY"
# split data by 60% train ; use random_state=100
features_train, features_validation_test, labels_train, labels_validation_test = train_test_split(features,labels,test_size=0.40,random_state=100)
# + id="oZXTP5lgfqKY"
# split test data by 50% validation and 50% test ; use random_state=100
features_validation, features_test, labels_validation, labels_test = train_test_split(features_validation_test,labels_validation_test,test_size=0.50,random_state=100)
# + id="oCRpVXr0fqKY" outputId="55e7358b-2a97-4327-8a73-42b6627caadd" colab={"base_uri": "https://localhost:8080/"}
print(len(features_train))
print(len(features_validation))
print(len(features_test))
print(len(labels_train))
print(len(labels_validation))
print(len(labels_test))
# + [markdown] id="5dJMAfTUfqKZ"
# ### 12.4.2 Training different models on our dataset
#
# We'll train six models:
# - Logistic regression (perceptron)
# - Decision tree
# - Support vector machine (SVM)
# - RandomForestClassifier
# - GradientBoostingClassifier
# - AdaBoostClassifier
# + id="_gP26kwwfqKZ" outputId="b617531e-b1b9-4a8b-d1a4-1c7336813ba2" colab={"base_uri": "https://localhost:8080/"}
# Train logistic regression model
from sklearn.linear_model import LogisticRegression
lr_model = LogisticRegression().fit(features_train,labels_train)
# + id="DThp9iwIfqKZ"
# Train decision tree model ; don't use any hyperparameter
from sklearn.tree import DecisionTreeClassifier
dt_model = DecisionTreeClassifier().fit(features_train,labels_train)
# + id="8yMO82c9fqKa"
# Train SVM model ; don't use any hyperparameter
from sklearn.svm import SVC
svm_model = SVC().fit(features_train,labels_train)
# + id="yhIfxavqfqKa"
# Train random forest model ; don't use any hyperparameter
from sklearn.ensemble import RandomForestClassifier
rf_model = RandomForestClassifier().fit(features_train,labels_train)
# + id="EW7wDPHIfqKa"
# Train gradient boosting model ; don't use any hyperparameter
from sklearn.ensemble import GradientBoostingClassifier
gb_model = GradientBoostingClassifier().fit(features_train,labels_train)
# + id="YPmyHC-ZfqKa"
# Train Adaboost model ; don't use any hyperparameter
from sklearn.ensemble import AdaBoostClassifier
ab_model = AdaBoostClassifier().fit(features_train,labels_train)
# + [markdown] id="6jF0KbS_fqKb"
# ### 12.4.3 Evaluating the models
#
# #### Accuracy
# + id="hjeMJk4KfqKb" outputId="44cd2b76-fe2e-45a9-e98a-b824ad692439" colab={"base_uri": "https://localhost:8080/"}
# print accuracy of each model on validation data
print("Scores of the models")
print("Logistic regression:",lr_model.score(features_validation,labels_validation) )
print("Decision tree:", dt_model.score(features_validation,labels_validation))
print("SVM:", svm_model.score(features_validation,labels_validation))
print("Random forest:",rf_model.score(features_validation,labels_validation) )
print("Gradient boosting:",gb_model.score(features_validation,labels_validation) )
print("AdaBoost:",ab_model.score(features_validation,labels_validation) )
# + [markdown] id="gCUK9NJXfqKb"
# #### F1-score
# + id="QU9uZMJkfqKb" outputId="5b5c811a-1379-4492-8445-a5ce7161fe69" colab={"base_uri": "https://localhost:8080/"}
# print F1-score of each model on validation data
from sklearn.metrics import f1_score
print("F1-scores of the models:")
lr_predicted_labels = lr_model.predict(features_validation)
print("Logistic regression:", f1_score(labels_validation, lr_predicted_labels))
dt_predicted_labels = dt_model.predict(features_validation)
print("Decision Tree:", f1_score(labels_validation, dt_predicted_labels))
svm_predicted_labels = svm_model.predict(features_validation)
print("Support Vector Machine:", f1_score(labels_validation, svm_predicted_labels))
rf_predicted_labels = rf_model.predict(features_validation)
print("Random Forest:", f1_score(labels_validation, rf_predicted_labels))
gb_predicted_labels = gb_model.predict(features_validation)
print("Gradient boosting:", f1_score(labels_validation, gb_predicted_labels))
ab_predicted_labels = ab_model.predict(features_validation)
print("AdaBoost:", f1_score(labels_validation, ab_predicted_labels))
# + [markdown] id="cN_q429mfqKb"
# ### 12.4.4 Testing the model
#
# Finding the accuracy and the F1-score of the model in the testing set.
# + id="_nfxWhppfqKb" outputId="613ed4c5-73f4-4797-b8b5-ea84c42549cb" colab={"base_uri": "https://localhost:8080/"}
# print accuracy of gradient boost model on testing data
gb_model.score(features_test, labels_test)
# + id="oW0DEPGxfqKc" outputId="32899407-d68b-4129-dafb-01c2fb1583ec" colab={"base_uri": "https://localhost:8080/"}
# print F1-score of gradient boost model on testing data
gb_predicted_test_labels = gb_model.predict(features_test)
f1_score(labels_test, gb_predicted_test_labels)
# + [markdown] id="r1XGztd2fqKc"
# # 12.5 Grid search
# + id="gAK1vV5WfqKc"
from sklearn.model_selection import GridSearchCV
# + id="XrcPjLM1fqKc" outputId="cf6676be-a0b5-446b-9785-ad80df69b072" colab={"base_uri": "https://localhost:8080/"}
svm_parameters = {'kernel': ['rbf'],
'C': [0.01, 0.1, 1 , 10, 100],
'gamma': [0.01, 0.1, 1, 10, 100]
}
# use gridsearch to find the best hyperparameters
svm = SVC()
svm_gs = GridSearchCV(estimator = svm,
param_grid = svm_parameters)
svm_gs.fit(features_train, labels_train)
svm_winner = svm_gs.best_estimator_
svm_winner
svm_winner.score(features_test, labels_test)
# + id="jmtr0bLVfqKd" outputId="f8b9a0d6-11ed-455b-f169-f1e766c08938" colab={"base_uri": "https://localhost:8080/"}
svm_winner
# + [markdown] id="VDqUNXY1fqKd"
# # 12.6 Cross validation
# + id="kqi1xhi0fqKd" outputId="58451138-ff41-4c0c-fa29-2d11349f5f20" colab={"base_uri": "https://localhost:8080/"}
# print the k-fold cross validation output
svm_gs.cv_results_
# + id="aQEacljQyT80"
| ML_in_practice/End_to_end_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: eunil_py38
# language: python
# name: python3
# ---
import os
os.environ["CUDA_VISIBLE_DEVICES"]="1"
# # 사용 패키지
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import cv2
from tqdm import tqdm
from glob import glob
import os
import json
import torch
from torch import nn
from torchvision import models
from torch.utils.data import Dataset
from sklearn.metrics import f1_score
# # 데이터 살펴보기
# +
sample = 10123
sample_csv = pd.read_csv(f'sample_data/{sample}/{sample}.csv')
sample_image = cv2.imread(f'sample_data/{sample}/{sample}.jpg')
sample_json = json.load(open(f'sample_data/{sample}/{sample}.json', 'r'))
# -
# csv
sample_csv
# image
plt.imshow(cv2.cvtColor(sample_image, cv2.COLOR_BGR2RGB))
plt.show()
# json
sample_json
# +
# visualize bbox
plt.figure(figsize=(7,7))
points = sample_json['annotations']['bbox'][0]
part_points = sample_json['annotations']['part']
img = cv2.cvtColor(sample_image, cv2.COLOR_BGR2RGB)
cv2.rectangle(
img,
(int(points['x']), int(points['y'])),
(int((points['x']+points['w'])), int((points['y']+points['h']))),
(0, 255, 0),
2
)
for part_point in part_points:
point = part_point
cv2.rectangle(
img,
(int(point['x']), int(point['y'])),
(int((point['x']+point['w'])), int((point['y']+point['h']))),
(255, 0, 0),
1
)
plt.imshow(img)
plt.show()
# -
# # 데이터 로드
#
# ## 환경 데이터 통계량 계산 for MinMax Scaling
# +
# 분석에 사용할 feature 선택
csv_features = ['내부 온도 1 평균', '내부 온도 1 최고', '내부 온도 1 최저', '내부 습도 1 평균', '내부 습도 1 최고',
'내부 습도 1 최저', '내부 이슬점 평균', '내부 이슬점 최고', '내부 이슬점 최저']
csv_files = sorted(glob('sample_data/*/*.csv'))
temp_csv = pd.read_csv(csv_files[0])[csv_features]
max_arr, min_arr = temp_csv.max().to_numpy(), temp_csv.min().to_numpy()
# feature 별 최대값, 최솟값 계산
for csv in tqdm(csv_files[1:]):
temp_csv = pd.read_csv(csv)[csv_features]
temp_max, temp_min = temp_csv.max().to_numpy(), temp_csv.min().to_numpy()
max_arr = np.max([max_arr,temp_max], axis=0)
min_arr = np.min([min_arr,temp_min], axis=0)
# feature 별 최대값, 최솟값 dictionary 생성
csv_feature_dict = {csv_features[i]:[min_arr[i], max_arr[i]] for i in range(len(csv_features))}
csv_feature_dict
# -
# ## CustomDataset 제작
# +
# 제공된 sample data는 파프리카와 시설포도 2종류의 작물만 존재
label_description = {
'3_00_0': '파프리카_정상',
'3_a9_1': '파프리카흰가루병_초기',
'3_a9_2': '파프리카흰가루병_중기',
'3_a9_3': '파프리카흰가루병_말기',
'3_a10_1': '파프리카잘록병_초기',
'3_a10_2': '파프리카잘록병_중기',
'3_a10_3': '파프리카잘록병_말기',
'3_b3_1': '칼슘결핍_초기',
'3_b3_2': '칼슘결핍_중기',
'3_b3_3': '칼슘결핍_말기',
'3_b6_1': '다량원소결핍 (N)_초기',
'3_b6_2': '다량원소결핍 (N)_중기',
'3_b6_3': '다량원소결핍 (N)_말기',
'3_b7_1': '다량원소결핍 (P)_초기',
'3_b7_2': '다량원소결핍 (P)_중기',
'3_b7_3': '다량원소결핍 (P)_말기',
'3_b8_1': '다량원소결핍 (K)_초기',
'3_b8_2': '다량원소결핍 (K)_중기',
'3_b8_3': '다량원소결핍 (K)_말기',
'6_00_0': '시설포도_정상',
'6_a11_1': '시설포도탄저병_초기',
'6_a11_2': '시설포도탄저병_중기',
'6_a11_3': '시설포도탄저병_말기',
'6_a12_1': '시설포도노균병_초기',
'6_a12_2': '시설포도노균병_중기',
'6_a12_3': '시설포도노균병_말기',
'6_b4_1': '일소피해_초기',
'6_b4_2': '일소피해_중기',
'6_b4_3': '일소피해_말기',
'6_b5_1': '축과병_초기',
'6_b5_2': '축과병_중기',
'6_b5_3': '축과병_말기',
}
label_encoder = {key:idx for idx, key in enumerate(label_description)}
label_decoder = {val:key for key, val in label_encoder.items()}
# -
class CustomDataset(Dataset):
def __init__(self, files, labels=None, mode='train'):
self.mode = mode
self.files = files
self.csv_feature_dict = csv_feature_dict
self.csv_feature_check = [0]*len(self.files)
self.csv_features = [None]*len(self.files)
self.max_len = -1 * 24*6
self.label_encoder = label_encoder
def __len__(self):
return len(self.files)
def __getitem__(self, i):
file = self.files[i]
file_name = file.split('/')[-1]
json_path = f'{file}/{file_name}.json'
image_path = f'{file}/{file_name}.jpg'
if self.csv_feature_check[i] == 0:
csv_path = f'{file}/{file_name}.csv'
df = pd.read_csv(csv_path)
# MinMax scaling
for col in self.csv_feature_dict.keys():
df[col] = df[col] - self.csv_feature_dict[col][0]
df[col] = df[col] / (self.csv_feature_dict[col][1]-self.csv_feature_dict[col][0])
# transpose to sequential data
csv_feature = df[self.csv_feature_dict.keys()].to_numpy()[self.max_len:].T
self.csv_features[i] = csv_feature
self.csv_feature_check[i] = 1
else:
csv_feature = self.csv_features[i]
img = cv2.imread(image_path)
img = cv2.resize(img, dsize=(256, 256), interpolation=cv2.INTER_AREA)
img = img.astype(np.float32)/255
img = np.transpose(img, (2,0,1))
if self.mode == 'train':
with open(json_path, 'r') as f:
json_file = json.load(f)
crop = json_file['annotations']['crop']
disease = json_file['annotations']['disease']
risk = json_file['annotations']['risk']
label = f'{crop}_{disease}_{risk}'
return {
'img' : torch.tensor(img, dtype=torch.float32),
'csv_feature' : torch.tensor(csv_feature, dtype=torch.float32),
'label' : torch.tensor(self.label_encoder[label], dtype=torch.long)
}
else:
return {
'img' : torch.tensor(img, dtype=torch.float32),
'csv_feature' : torch.tensor(csv_feature, dtype=torch.float32)
}
# # 하이퍼파라미터 및 변수
device = torch.device("cuda:0")
batch_size = 256
class_n = len(label_encoder)
learning_rate = 1e-4
embedding_dim = 512
num_features = len(csv_feature_dict)
max_len = 24*6
dropout_rate = 0.1
epochs = 30
vision_pretrain = True
save_path = 'best_model.pt'
# # 데이터셋 구성
# +
data_files = glob('sample_data/*')
train = data_files[:250]
val = data_files[250:]
# +
train_dataset = CustomDataset(train)
val_dataset = CustomDataset(val)
train_dataloader = torch.utils.data.DataLoader(train_dataset, batch_size=batch_size, num_workers=16, shuffle=True)
val_dataloader = torch.utils.data.DataLoader(val_dataset, batch_size=batch_size, num_workers=16, shuffle=False)
# -
# # 모델
#
# ## 이미지 분류 모델 : Resnet50
class CNN_Encoder(nn.Module):
def __init__(self, class_n, rate=0.1):
super(CNN_Encoder, self).__init__()
self.model = models.resnet50(pretrained=True)
def forward(self, inputs):
output = self.model(inputs)
return output
# ## 시계열 모델 : LSTM
class RNN_Decoder(nn.Module):
def __init__(self, max_len, embedding_dim, num_features, class_n, rate):
super(RNN_Decoder, self).__init__()
self.lstm = nn.LSTM(max_len, embedding_dim)
self.rnn_fc = nn.Linear(num_features*embedding_dim, 1000)
self.final_layer = nn.Linear(1000 + 1000, class_n)
self.dropout = nn.Dropout(rate)
def forward(self, enc_out, dec_inp):
hidden, _ = self.lstm(dec_inp)
hidden = hidden.view(hidden.size(0), -1)
hidden = self.rnn_fc(hidden)
concat = torch.cat([enc_out, hidden], dim=1) # enc_out + hidden
fc_input = concat
output = self.dropout((self.final_layer(fc_input)))
return output
# ## 앙상블
class CNN2RNN(nn.Module):
def __init__(self, max_len, embedding_dim, num_features, class_n, rate):
super(CNN2RNN, self).__init__()
self.cnn = CNN_Encoder(embedding_dim, rate)
self.rnn = RNN_Decoder(max_len, embedding_dim, num_features, class_n, rate)
def forward(self, img, seq):
cnn_output = self.cnn(img)
output = self.rnn(cnn_output, seq)
return output
model = CNN2RNN(max_len=max_len, embedding_dim=embedding_dim, num_features=num_features, class_n=class_n, rate=dropout_rate)
model = model.to(device)
# # 학습
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
criterion = nn.CrossEntropyLoss()
# +
def accuracy_function(real, pred):
real = real.cpu()
pred = torch.argmax(pred, dim=1).cpu()
score = f1_score(real, pred, average='macro')
return score
def train_step(batch_item, training):
img = batch_item['img'].to(device)
csv_feature = batch_item['csv_feature'].to(device)
label = batch_item['label'].to(device)
if training is True:
model.train()
optimizer.zero_grad()
with torch.cuda.amp.autocast():
output = model(img, csv_feature)
loss = criterion(output, label)
loss.backward()
optimizer.step()
score = accuracy_function(label, output)
return loss, score
else:
model.eval()
with torch.no_grad():
output = model(img, csv_feature)
loss = criterion(output, label)
score = accuracy_function(label, output)
return loss, score
# +
loss_plot, val_loss_plot = [], []
metric_plot, val_metric_plot = [], []
for epoch in range(epochs):
total_loss, total_val_loss = 0, 0
total_acc, total_val_acc = 0, 0
tqdm_dataset = tqdm(enumerate(train_dataloader))
training = True
for batch, batch_item in tqdm_dataset:
batch_loss, batch_acc = train_step(batch_item, training)
total_loss += batch_loss
total_acc += batch_acc
tqdm_dataset.set_postfix({
'Epoch': epoch + 1,
'Loss': '{:06f}'.format(batch_loss.item()),
'Mean Loss' : '{:06f}'.format(total_loss/(batch+1)),
'Mean F-1' : '{:06f}'.format(total_acc/(batch+1))
})
loss_plot.append(total_loss/(batch+1))
metric_plot.append(total_acc/(batch+1))
tqdm_dataset = tqdm(enumerate(val_dataloader))
training = False
for batch, batch_item in tqdm_dataset:
batch_loss, batch_acc = train_step(batch_item, training)
total_val_loss += batch_loss
total_val_acc += batch_acc
tqdm_dataset.set_postfix({
'Epoch': epoch + 1,
'Val Loss': '{:06f}'.format(batch_loss.item()),
'Mean Val Loss' : '{:06f}'.format(total_val_loss/(batch+1)),
'Mean Val F-1' : '{:06f}'.format(total_val_acc/(batch+1))
})
val_loss_plot.append(total_val_loss/(batch+1))
val_metric_plot.append(total_val_acc/(batch+1))
if np.max(val_metric_plot) == val_metric_plot[-1]:
torch.save(model, save_path)
# -
# # 학습 결과
plt.figure(figsize=(10,7))
plt.plot(loss_plot, label='train_loss')
plt.plot(val_loss_plot, label='val_loss')
plt.xlabel('epoch')
plt.ylabel('loss')
plt.title("Loss", fontsize=25)
plt.legend()
plt.show()
# # 추론
# +
def predict(dataset):
model.eval()
tqdm_dataset = tqdm(enumerate(dataset))
training = False
results = []
answer = []
for batch, batch_item in tqdm_dataset:
img = batch_item['img'].to(device)
seq = batch_item['csv_feature'].to(device)
with torch.no_grad():
output = model(img, seq)
output = torch.tensor(torch.argmax(output, axis=-1), dtype=torch.int32).cpu().numpy()
results.extend(output)
answer.extend(batch_item['label'])
return results, answer
preds, answer = predict(val_dataloader)
# -
# ## 추론 결과 시각화
# +
answer = np.array([label_description[label_decoder[int(val)]] for val in answer])
preds = np.array([label_description[label_decoder[int(val)]] for val in preds])
new_crosstab = pd.crosstab(answer, preds, rownames=['answer'], colnames=['preds'])
new_crosstab
# -
| base code/[Sample_Baseline] Resnet50 + LSTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/NoCodeProgram/CodingTest/blob/main/hashMap/contigArray.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Cz7Wsu-UlNRx"
# Title : Contiguous Array
#
# Chapter : Hash Map
#
# Link : [YouTube](https://youtu.be/yDwH1QwiaWQ)
#
# ChapterLink : [PlayList](https://www.youtube.com/playlist?list=PLDV-cCQnUlIYjwJ_b-f8Z0OQlIqKpMkDr)
#
# 문제: 1과 0으로 이루어진 array에서 1과 0의 갯수가 같은 subarray의 최대길이는 몇 인가?
# + id="zviLSTsnlJgn"
from typing import List
def findMaxLength(nums: List[int]) -> int:
for idx in range(len(nums)):
if nums[idx] == 0 :
nums[idx] = -1
cml_sums = []
tmp_sum = 0
for num in nums:
tmp_sum += num
cml_sums.append(tmp_sum)
table = {}
max_length = 0
table[0] = [-1]
for idx, cml_sum in enumerate(cml_sums):
if cml_sum not in table:
table[cml_sum] = [idx]
else:
table[cml_sum].append(idx)
indices = table[cml_sum]
first_idx = indices[0]
last_idx = indices[-1]
length = last_idx - first_idx
max_length = max(max_length,length)
return max_length
# + colab={"base_uri": "https://localhost:8080/"} id="xNx6K85Xlijw" outputId="10719982-450d-4db1-e595-77655cbb9f36"
findMaxLength(nums=[1,0,1,1,1,0,0,1,1])
| hashMap/contigArray.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 ('base')
# language: python
# name: python3
# ---
# +
import xpress as xp
xp.controls.outputlog = 0 # turn off output log
import numpy as np
import pylab as pl
# +
# Define weight and values of the knapsack items
n_items = 8
n_bags = 2
max_bag_weight = 15
weights = np.array([1, 7, 4, 3, 5, 6, 2, 7])
ratings = np.array([2, 9, 3, 8, 10, 6, 2, 10])
availabilities = np.array([4, 2, 3, 2, 1, 1, 3, 1])
# -
knapsack = xp.problem()
x = np.array([[xp.var(vartype=xp.integer, lb=0) for i in range(n_items)] for j in range(n_bags)], dtype=xp.npvar)
knapsack.addVariable(x)
# Weight constraint
knapsack.addConstraint(xp.Dot(x, weights) <= max_bag_weight)
# Availability constraint
knapsack.addConstraint(xp.Dot(np.ones(n_bags), x) <= availabilities)
knapsack.setObjective(xp.Sum(x*ratings), sense=xp.maximize)
knapsack.solve()
knapsack.getProbStatusString()
pl.plot(knapsack.getSolution())
# Print knapsack solution
knapsack.getSolution(), knapsack.getObjVal()
| Exercises/Labs/knapsack.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # COUNT EVENS
#
# Return the number of even integers in the given array.
#
# ## Examples:
# `count_evens([2, 1, 2, 3, 4]) → 3`
# `count_evens([2, 2, 0]) → 3`
# `count_evens([1, 3, 5]) → 0`
# + inputHidden=false outputHidden=false
count_evens = lambda numbers: len([number for number in numbers if number % 2 == 0])
# + inputHidden=false outputHidden=false
count_evens([2, 1, 2, 3, 4])
# -
count_evens([2, 2, 0])
count_evens([1, 3, 5])
| mk026-count_evens.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: atac_env
# language: python
# name: atac_env
# ---
# ## ATAC + MULTIOME GERM CELLS (april 2021)
#
# ### Build anndata from cellatac output
# Load libraries
import pandas as pd
import scanpy as sc
import numpy as np
import scipy.sparse
# Define variables
outdir = "/nfs/team292/vl6/my_MULTIOME_dir/germcells_apr2021/"
experiment_prefix = 'germcells_'
cellatac_outdir = '/lustre/scratch117/cellgen/cellgeni/TIC-atacseq/tic-1005/germcells-work/results200k-b-germ/'
# +
input={'cnts': cellatac_outdir + 'peak_matrix/peaks_bc_matrix.mmtx.gz',
"bcs": cellatac_outdir + 'peak_matrix/bc.txt',
"peaks": cellatac_outdir + 'peak_matrix/peaks.txt',
'clusters': cellatac_outdir + 'qc/seurat-clades.tsv'}
output = {
"cnt_mmtx":outdir + experiment_prefix + '_ATAC_cisTopic.mmtx',
"cnt_peaks":outdir + experiment_prefix + '_ATAC_cisTopic.peaks.tsv',
"cnt_cells":outdir + experiment_prefix + '_ATAC_cisTopic.cells.tsv',
"h5ad":outdir + experiment_prefix + '_ATAC_raw.h5ad',
}
# -
# #### Make anndata object
# +
adata = sc.read_mtx(input["cnts"]).T
bc = pd.read_table(input["bcs"], header=None)
feat = pd.read_table(input["peaks"], header=None)
adata.obs_names = bc[0]
adata.var_names = feat[0]
# -
adata
# #### Load peak annotations (done with R script by Emma)
peak_anno_df = pd.read_csv(outdir + "ATACpeaks_annotation.csv", index_col=0)
peak_anno_df.index = peak_anno_df["peak_id"]
peak_anno_df.drop("peak_id",1, inplace=True)
adata.var = pd.concat([adata.var, peak_anno_df], 1)
# #### Save binary data to layers
adata.layers["binary_raw"] = adata.X
adata.layers["binary_raw"][adata.layers["binary_raw"] > 1] = 1
# #### Peak filtering
adata.var.hist(column = 'peak_width', bins = 200, grid = False, figsize = (25,6), color = '#870052')
var_qc = sc.pp.calculate_qc_metrics(adata, layer = "binary_raw")[1]
adata.var = pd.concat([adata.var, var_qc], 1)
adata
adata.var.head()
adata.var.hist(column = 'total_counts', bins = 200, grid = False, figsize = (25,6), color = '#870052')
thirty_percent = len(adata.obs_names) / 100 * 30
point_two_percent = len(adata.obs_names) / 100 * 0.2
print("30% : {}".format(thirty_percent))
print("0.2% : {}".format(point_two_percent))
# Accessible in at least k cells
adata = adata[:,adata.var.total_counts > point_two_percent]
adata = adata[:,adata.var.total_counts < thirty_percent]
adata
# Remove peaks in ENCODE blacklist
adata = adata[:, adata.var.ENCODE_blacklist == 0]
adata
# +
# Filter by width (remove peaks at the lowest end, closest to min peak width in MACS2)
adata = adata[:, adata.var.peak_width > 210]
adata = adata[:, adata.var.peak_width < 1500]
adata
# -
adata.var['annotation'].value_counts()
### Filter peaks that are not accessible in at least 4% of cells from a coarse cluster
min_frac=0.04
# Load cluster information from cellatac outputs
clusters = pd.read_table(input["clusters"], header = None, index_col = 0)
clusters.columns = ["cellatac_clusters"]
adata.obs = clusters.loc[adata.obs_names]
adata
adata.obs.head()
# +
n_clusters = len(np.unique(adata.obs[["cellatac_clusters"]]))
clus_mat = np.empty([adata.n_obs, n_clusters])
for cl in np.unique(adata.obs[["cellatac_clusters"]]):
clus_mat[np.where(adata.obs['cellatac_clusters']==cl)[0],cl] = 1
clus_mat = scipy.sparse.csr_matrix(clus_mat)
clus_mat[clus_mat != 1 ] = 0
# -
cl_peak_mat = np.dot(clus_mat.T, adata.layers["binary_raw"])
cl_peak_frac = cl_peak_mat/clus_mat.sum(0).T
cl_peak_frac.max(0).shape
bool_matrix = cl_peak_frac.max(0) > min_frac
bool_matrix.shape
bool_vector = np.squeeze(np.asarray(bool_matrix))
bool_vector.shape
adata = adata[:, bool_vector]
adata
# #### Filter low quality cells
sc.pp.calculate_qc_metrics(adata, layer = "binary_raw", inplace = True)
adata.var.hist(column = 'log1p_total_counts', bins = 200, grid = False, figsize = (25,6), color = '#870052')
adata.obs.hist(column = 'log1p_total_counts', bins = 200, grid = False, figsize = (25,6), color = '#870052')
adata = adata[adata.obs.log1p_total_counts >= 5.5]
adata
## Write output anndata
adata.write_h5ad(output["h5ad"])
# #### End of notebook
| scATACseq_germcells_0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # 14.2 Tidal period and the moon's revolution
#
# Tides are caused by the gravitational pull of the moon on the earth's water, stretching it into an eliptical shape.
#
# 
#
# So the side of the earth facing the moon, as well as the opposite side, will experience high tide. The sides perpendicular to the direction of the moon will experience a low tide.
#
# As the earth rotates on its axis, each spot on the earth will experience two high tides every rotation.
#
# How long will this take?
# + [markdown] slideshow={"slide_type": "slide"}
# It seems reasonable to argue that there should be two high tides every 24 hours. However this is missing two important factors. The first of these is that the moon revolves around the earth (in the same direction as the earth's rotation) and so during 24 hours the moon will have moved through a portion of its revolution.
#
# Looking at this from above the north pole, we could define and angle, $\theta$, to be the "extra" amount that the earth would need to rotate in order for the same spot on the surface to be under the moon.
#
# 
# + [markdown] slideshow={"slide_type": "slide"}
# We know the rotational period of the earth is 24 hours and the orbital period of the moon, $T_M$, is 29 days, 12 hours, 44 minutes, and 2.8 seconds.
# + slideshow={"slide_type": "subslide"}
T_M = 29*24 + 12 + 44/60 + 2.8/(60*60)
print(T_M, 'hours for one orbit of the moon')
# + [markdown] slideshow={"slide_type": "subslide"}
# So $T_E = 24$ hours and $T_M = 708.734$ hours.
#
# This means that the rotational speed of the earth is $\omega_E = \frac{1}{24}$ cycles per hour and the orbital speed of the moon is $\omega_M = \frac{1}{708.734}$ cycles per hour.
#
# Use the equation $\omega = \frac{\theta}{t}$ to find the time that it would take for a spot on the earth to line up with the moon again.
#
# As a hint, we know that the time for the earth to rotate to that position must be the same as the time for the moon to revolve to that position. This means we can say $t_E = t_M = T_E + \frac{\theta}{\omega_E} = \frac{\theta}{\omega_M}$. Find $t_E$ or $t_M$.
# + slideshow={"slide_type": "slide"}
T_E = 24
T_M = 29*24 + 12 + 44/60 + 2.8/(60*60)
omega_E = 1/T_E
omega_M = 1/T_M
# fill in your equation below then run this cell
t_M =
print(t_M)
| 14-The-period-of-the-tides/14.2-Tidal-period-and-moon-revolution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # VGG16 model for Keras
# - [Very Deep Convolutional Networks for Large-Scale Image Recognition](https://arxiv.org/abs/1409.1556)
# +
from __future__ import print_function
from __future__ import absolute_import
import warnings
from keras.models import Model
from keras.layers import Flatten
from keras.layers import Dense
from keras.layers import Input
from keras.layers import Conv2D
from keras.layers import MaxPooling2D
from keras.layers import GlobalAveragePooling2D
from keras.layers import GlobalMaxPooling2D
from keras.engine.topology import get_source_inputs
from keras.utils import layer_utils
from keras.utils.data_utils import get_file
from keras import backend as K
from keras.preprocessing.image import load_img
from keras.preprocessing import image
from keras.applications.imagenet_utils import decode_predictions
from keras.applications.imagenet_utils import preprocess_input
from keras.applications.imagenet_utils import _obtain_input_shape
import numpy as np
# -
WEIGHTS_PATH = 'https://github.com/fchollet/deep-learning-models/releases/download/v0.1/vgg16_weights_tf_dim_ordering_tf_kernels.h5'
WEIGHTS_PATH_NO_TOP = 'https://github.com/fchollet/deep-learning-models/releases/download/v0.1/vgg16_weights_tf_dim_ordering_tf_kernels_notop.h5'
# ## *Let's have some coding fun!*
#
#
# In the below VGG16 function, you will need to fill in Neural Network Architecture Design from the Diagram in the above VGG-16 paper.
def VGG16(include_top=True, weights='imagenet',
input_tensor=None, input_shape=None,
pooling=None,
classes=1000):
"""Instantiates the VGG16 architecture.
Optionally loads weights pre-trained
on ImageNet. Note that when using TensorFlow,
for best performance you should set
`image_data_format="channels_last"` in your Keras config
at ~/.keras/keras.json.
The model and the weights are compatible with both
TensorFlow and Theano. The data format
convention used by the model is the one
specified in your Keras config file.
# Arguments
include_top: whether to include the 3 fully-connected
layers at the top of the network.
weights: one of `None` (random initialization)
or "imagenet" (pre-training on ImageNet).
input_tensor: optional Keras tensor (i.e. output of `layers.Input()`)
to use as image input for the model.
input_shape: optional shape tuple, only to be specified
if `include_top` is False (otherwise the input shape
has to be `(224, 224, 3)` (with `channels_last` data format)
or `(3, 224, 224)` (with `channels_first` data format).
It should have exactly 3 inputs channels,
and width and height should be no smaller than 48.
E.g. `(200, 200, 3)` would be one valid value.
pooling: Optional pooling mode for feature extraction
when `include_top` is `False`.
- `None` means that the output of the model will be
the 4D tensor output of the
last convolutional layer.
- `avg` means that global average pooling
will be applied to the output of the
last convolutional layer, and thus
the output of the model will be a 2D tensor.
- `max` means that global max pooling will
be applied.
classes: optional number of classes to classify images
into, only to be specified if `include_top` is True, and
if no `weights` argument is specified.
# Returns
A Keras model instance.
# Raises
ValueError: in case of invalid argument for `weights`,
or invalid input shape.
"""
if weights not in {'imagenet', None}:
raise ValueError('The `weights` argument should be either '
'`None` (random initialization) or `imagenet` '
'(pre-training on ImageNet).')
if weights == 'imagenet' and include_top and classes != 1000:
raise ValueError('If using `weights` as imagenet with `include_top`'
' as true, `classes` should be 1000')
# Determine proper input shape
input_shape = _obtain_input_shape(input_shape,
default_size=224,
min_size=48,
data_format=K.image_data_format(),
include_top=include_top)
if input_tensor is None:
img_input = Input(shape=input_shape)
else:
if not K.is_keras_tensor(input_tensor):
img_input = Input(tensor=input_tensor, shape=input_shape)
else:
img_input = input_tensor
# Block 1
x = Conv2D(64, (3, 3), activation='relu', padding='same', name='block1_conv1')(img_input)
x = Conv2D(64, (3, 3), activation='relu', padding='same', name='block1_conv2')(x)
x = MaxPooling2D((2, 2), strides=(2, 2), name='block1_pool')(x)
# Block 2
# TODO: Fill in from the Diagram in the above VGG-16 paper
# Block 3
# TODO: Fill in from the Diagram in the above VGG-16 paper
# Block 4
# TODO: Fill in from the Diagram in the above VGG-16 paper
# Block 5
# TODO: Fill in from the Diagram in the above VGG-16 paper
if include_top:
# Classification block
x = Flatten(name='flatten')(x)
x = Dense(4096, activation='relu', name='fc1')(x)
x = Dense(4096, activation='relu', name='fc2')(x)
x = Dense(classes, activation='softmax', name='predictions')(x)
else:
if pooling == 'avg':
x = GlobalAveragePooling2D()(x)
elif pooling == 'max':
x = GlobalMaxPooling2D()(x)
# Ensure that the model takes into account
# any potential predecessors of `input_tensor`.
if input_tensor is not None:
inputs = get_source_inputs(input_tensor)
else:
inputs = img_input
# Create model.
model = Model(inputs, x, name='vgg16')
# load weights
if weights == 'imagenet':
if include_top:
weights_path = get_file('vgg16_weights_tf_dim_ordering_tf_kernels.h5',
WEIGHTS_PATH,
cache_subdir='models')
else:
weights_path = get_file('vgg16_weights_tf_dim_ordering_tf_kernels_notop.h5',
WEIGHTS_PATH_NO_TOP,
cache_subdir='models')
model.load_weights(weights_path)
if K.backend() == 'theano':
layer_utils.convert_all_kernels_in_model(model)
if K.image_data_format() == 'channels_first':
if include_top:
maxpool = model.get_layer(name='block5_pool')
shape = maxpool.output_shape[1:]
dense = model.get_layer(name='fc1')
layer_utils.convert_dense_weights_data_format(dense, shape, 'channels_first')
if K.backend() == 'tensorflow':
warnings.warn('You are using the TensorFlow backend, yet you '
'are using the Theano '
'image data format convention '
'(`image_data_format="channels_first"`). '
'For best performance, set '
'`image_data_format="channels_last"` in '
'your Keras config '
'at ~/.keras/keras.json.')
return model
model = VGG16(include_top=True, weights='imagenet', input_tensor=None, input_shape=None)
# +
img_path = "img/WeRise.png"
img = image.load_img(img_path, target_size=(224, 224))
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
x = preprocess_input(x)
preds = model.predict(x)
# -
# This next command should generate an integer.
#
# You can look up the image category in the ```code/imagenet-classes.csv``` file.
out = model.predict(x)
print(np.argmax(out))
# # Try your own images!!!
| notebooks/03_VGG16.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### 1. What is a common reason for an ML model that works well in training but fails in production?
# ##### Ans: The ML dataset was improperly created
# #### 2. Personalized Algorithms are often built using which type of ML model?
# ##### Ans: Recommendation systems
# #### 3. What is a key lesson Google has learned with regards to reducing the chance of failure in production ML models?
# ##### Ans: Process batch data and streaming data the same way
| Coursera/How Google does Machine Learning/Week-1/Quiz/Module-1-Quiz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 20170702 실험
#
# 더 많은 Places 데이터셋에 대해서 학습한 DCGAN모델을 이용하여 피쳐를 만들고 다시 해본다..
# +
import os
import scipy.misc
import numpy as np
from sklearn.decomposition import PCA
from model import DCGAN
from utils import pp, visualize, to_json, show_all_variables
import tensorflow as tf
from glob import glob
import sys
flags = tf.app.flags
flags.DEFINE_integer("epoch", 25, "Epoch to train [25]")
flags.DEFINE_float("learning_rate", 0.0002, "Learning rate of for adam [0.0002]")
flags.DEFINE_float("beta1", 0.5, "Momentum term of adam [0.5]")
flags.DEFINE_integer("train_size", np.inf, "The size of train images [np.inf]")
flags.DEFINE_integer("batch_size", 64, "The size of batch images [64]")
flags.DEFINE_integer("input_height", 64, "The size of image to use (will be center cropped). [108]")
flags.DEFINE_integer("input_width", None,
"The size of image to use (will be center cropped). If None, same value as input_height [None]")
flags.DEFINE_integer("output_height", 64, "The size of the output images to produce [64]")
flags.DEFINE_integer("output_width", None,
"The size of the output images to produce. If None, same value as output_height [None]")
flags.DEFINE_string("dataset", "PatchofPlaces", "The name of dataset [celebA, mnist, lsun]")
flags.DEFINE_string("input_fname_pattern", "*/*.jpg", "Glob pattern of filename of input images [*]")
flags.DEFINE_string("checkpoint_dir", "checkpoint", "Directory name to save the checkpoints [checkpoint]")
flags.DEFINE_string("sample_dir", "samples", "Directory name to save the image samples [samples]")
flags.DEFINE_boolean("train", False, "True for training, False for testing [False]")
flags.DEFINE_boolean("crop", False, "True for training, False for testing [False]")
flags.DEFINE_boolean("visualize", False, "True for visualizing, False for nothing [False]")
FLAGS = flags.FLAGS
pp.pprint(flags.FLAGS.__flags)
if FLAGS.input_width is None:
FLAGS.input_width = FLAGS.input_height
if FLAGS.output_width is None:
FLAGS.output_width = FLAGS.output_height
if not os.path.exists(FLAGS.checkpoint_dir):
os.makedirs(FLAGS.checkpoint_dir)
if not os.path.exists(FLAGS.sample_dir):
os.makedirs(FLAGS.sample_dir)
# gpu_options = tf.GPUOptions(per_process_gpu_memory_fraction=0.333)
run_config = tf.ConfigProto()
run_config.gpu_options.allow_growth = True
sess = tf.Session(config=run_config)
dcgan = DCGAN(
sess,
input_width=FLAGS.input_width,
input_height=FLAGS.input_height,
output_width=FLAGS.output_width,
output_height=FLAGS.output_height,
batch_size=FLAGS.batch_size,
sample_num=FLAGS.batch_size,
dataset_name=FLAGS.dataset,
input_fname_pattern=FLAGS.input_fname_pattern,
crop=FLAGS.crop,
checkpoint_dir=FLAGS.checkpoint_dir,
sample_dir=FLAGS.sample_dir)
show_all_variables()
if not dcgan.load(FLAGS.checkpoint_dir)[0]:
raise Exception("[!] Train a model first, then run test mode")
# -
# ## Utility functions
# +
def layer_extraction(dcgan, file_names):
return dcgan.get_feature(FLAGS, file_names)
def maxpooling(disc):
kernel_stride_size = 4
maxpooling = [
tf.nn.max_pool(disc[i],ksize=[1,2**(4-i),2**(4-i),1],
strides=[1,2**(4-i),2**(4-i),1],padding='SAME')
for i in range(4)
]
# tf.global_variables_initializer().run()
maxpool_result = sess.run(maxpooling)
# for idx in range(4):
# print(idx, maxpool_result[idx].shape)
return maxpool_result
def flatten(disc):
flatten = [
tf.reshape(disc[i],[64, -1])
for i in range(4)
]
# tf.global_variables_initializer().run()
flatten_result = sess.run(flatten)
return flatten_result
def concat(disc):
concat = tf.concat(disc,1)
# tf.global_variables_initializer().run()
concat_result = sess.run(concat)
return concat_result
def feature_ext_GAN(file_names):
ret = layer_extraction(dcgan, file_names)
ret = maxpooling(ret)
ret = flatten(ret)
ret = concat(ret)
return ret
# -
# # Integration
# +
pca = PCA(n_components = 128)
patch_path ="/media/dongwonshin/Ubuntu Data/Datasets/FAB-MAP/Image Data/City Centre/patches/#300/"
data = sorted(glob("%s/%04d/*.jpg" % (patch_path, idx)))
output_filename = '/media/dongwonshin/Ubuntu Data/Datasets/FAB-MAP/Image Data/City Centre/descs/20170702/' + (name.split('/')[-2])+'.desc'
# patch_path ="/media/dongwonshin/Ubuntu Data/Datasets/Places365/Large_images/val_large/patches"
# data = glob("%s/Places365_val_%08d/*.jpg" % (patch_path, idx))
# output_filename = '/media/dongwonshin/Ubuntu Data/Datasets/Places365/Large_images/val_large/descs/20170702/' + (name.split('/')[-2])+'.desc'
for term in [24]:
print('%d ~ %d' % (100*term,100*(term+1)))
disc_list = []
batch_list = []
file_names = []
for idx in range(100*term,100*(term+1)):
file_names.append(data)
file_names=np.concatenate(file_names)
print('total:',len(file_names))
# print(file_names)
for idx in range(0, len(file_names)-64,64):
batch_files = file_names[idx: idx+64]
disc = feature_ext_GAN(batch_files)
disc_list.append(disc)
batch_list.append(batch_files)
sys.stdout.write('.')
final_disc_list = np.concatenate(disc_list)
final_batch_list = np.concatenate(batch_list)
X = np.array(final_disc_list)
pca.fit(X)
final_disc_list = pca.transform(X)
for idx, name in enumerate(final_batch_list):
with open(output_filename,'at') as fp:
for v in final_disc_list[idx]:
fp.write('%f ' % v)
fp.write('\n')
print('done.')
| 20170702 Experiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 3. 3-D example with 1 parameters
# The following example shows how to construct the kernel, automatically, from a symbolic expression defining the linear differential operator in **3D**.
#
# We consider the following operator, for an unknwon *u*
#
# $$
# \mathcal{L}^{\phi} u := \phi u + \partial_x u + \partial_{y} u + \partial_{zz} u
# $$
# +
# imports
from mlhiphy.calculus import dx, dy, dz
from mlhiphy.calculus import Constant
from mlhiphy.calculus import Unknown
from mlhiphy.kernels import compute_kernel, generic_kernel
from sympy import expand
from sympy import symbols
from sympy import exp
from sympy import Tuple
# +
x, x_i, x_j = symbols('x x_i x_j')
y, y_i, y_j = symbols('y y_i y_j')
z, z_i, z_j = symbols('z z_i z_j')
X = Tuple(x,y)
X_i = Tuple(x_i,y_i,z_i)
X_j = Tuple(x_j,y_j,z_j)
u = Unknown('u')
phi = Constant('phi')
theta_1 = Constant('theta_1')
theta_2 = Constant('theta_2')
theta_3 = Constant('theta_3')
expr = phi * u + dx(u) + dy(u) + dz(dz(u))
# -
kuu = generic_kernel(expr, u, (X_i, X_j))
# +
from IPython.display import Math
from sympy import latex
Math(latex(expand(kuu)))
# +
# RBF kernel
kuu = exp(- theta_1 * (x_i - x_j)**2 - theta_2 * (y_i - y_j)**2 - theta_3 * (z_i - z_j)**2)
kuf = compute_kernel(expr, kuu, X_i)
kfu = compute_kernel(expr, kuu, X_j)
kff = compute_kernel(expr, kuu, (X_i, X_j))
# -
Math(latex(expand(kuf)))
Math(latex(expand(kfu)))
Math(latex(expand(kff)))
from IPython.core.display import HTML
def css_styling():
styles = open("../styles/custom.css", "r").read()
return HTML(styles)
css_styling()
| autoker/03_example_3d.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:thor_py38]
# language: python
# name: conda-env-thor_py38-py
# ---
# ## ZTF
# In this notebook, we run THOR on two weeks of ZTF alerts. We select test orbits from the catalog of known objects (MPCORB.DAT).
#
# Data and results files for this notebook may be downloaded [here](https://dirac.astro.washington.edu/~moeyensj/projects/thor/paper1/).
# +
# %load_ext autoreload
# %autoreload 2
import glob
import os
import numpy as np
import pandas as pd
import sqlite3 as sql
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import seaborn as sns
sns.set(font_scale=1.2, context="paper", style="ticks")
sns.set_palette("viridis")
from astropy.time import Time
# %matplotlib inline
import plotly
plotly.offline.init_notebook_mode(connected=True)
# +
import thor
from thor import __version__
print("THOR version: {}".format(__version__))
# +
DATA_DIR = "/mnt/data/projects/thor/thor_data/ztf"
preprocessed_observations = pd.read_csv(
"/mnt/data/projects/thor/thor_data/ztf/preprocessed_observations.csv",
index_col=False,
dtype={
"obs_id" : str,
}
)
# -
preprocessed_observations.head(10)
# +
# Read orbits file (MPCORB in OORB format from 2018)
orbits = pd.read_csv(
"/mnt/data/projects/thor/thor_data/ztf/MPCORB_20181106_ZTF_keplerian.orb",
delim_whitespace=True,
skiprows=4,
names=["designation", "a_au", "e", "i_deg", "ascNode_deg", "argPeri_deg", "meanAnom_deg", "epoch_mjd_tt", "H", "G"],
low_memory=False
)
# -
simulated_ephemeris = pd.read_csv(
"/mnt/data/projects/thor/thor_data/ztf/MPCORB_20181106_ZTF.eph",
delim_whitespace=True,
header=0,
low_memory=False
)
simulated_ephemeris.rename(
columns={
"#Designation" : "designation",
"RA" : "RA_deg",
"Dec" : "Dec_deg",
"MJD_UTC/UT1" : "exp_mjd",
"r" : "r_au",
"HEclObj_X" : "HEclObj_X_au",
"HEclObj_Y" : "HEclObj_Y_au",
"HEclObj_Z" : "HEclObj_Z_au",
"HEclObj_dX/dt" : "HEclObj_dX/dt_au_p_day",
"HEclObj_dY/dt" : "HEclObj_dY/dt_au_p_day",
"HEclObj_dZ/dt" : "HEclObj_dZ/dt_au_p_day",
},
inplace=True
)
size = 15
ras = np.arange(0, 360 + size, size)
decs = np.arange(-90, 90 + size, size)
# +
from thor import findAverageOrbits
average_orbits_list = []
patch_number = 0
for ra_i, ra_f in zip(ras[:-1], ras[1:]):
for dec_i, dec_f in zip(decs[:-1], decs[1:]):
# See if there are any observations in the patch
observations_in_patch = preprocessed_observations[
(preprocessed_observations["RA_deg"] >= ra_i)
& (preprocessed_observations["RA_deg"] < ra_f)
& (preprocessed_observations["Dec_deg"] < dec_f)
& (preprocessed_observations["Dec_deg"] >= dec_i)
].copy()
if len(observations_in_patch) > 0:
# Find time of first set of observations in patch (need to propagate orbit to that time)
exp_mjd = observations_in_patch["mjd_utc"].min()
simulated_ephemeris_mask = (
(simulated_ephemeris["RA_deg"] >= ra_i)
& (simulated_ephemeris["RA_deg"] < ra_f)
& (simulated_ephemeris["Dec_deg"] < dec_f)
& (simulated_ephemeris["Dec_deg"] >= dec_i)
)
average_orbits_hun1_patch = findAverageOrbits(
simulated_ephemeris[simulated_ephemeris_mask],
orbits[(orbits["a_au"] < 2.06) & (orbits["a_au"] >= 1.7) & (orbits["e"] <= 0.1)],
element_type="keplerian",
d_values=[1.7, 2.06]
)
average_orbits_hun2_patch = findAverageOrbits(
simulated_ephemeris[simulated_ephemeris_mask],
orbits[(orbits["a_au"] < 2.06) & (orbits["a_au"] >= 1.7) & (orbits["e"] > 0.1) & (orbits["e"] <= 0.2)],
element_type="keplerian",
d_values=[1.7, 2.06]
)
average_orbits_hun3_patch = findAverageOrbits(
simulated_ephemeris[simulated_ephemeris_mask],
orbits[(orbits["a_au"] < 2.06) & (orbits["a_au"] >= 1.7) & (orbits["e"] > 0.2) & (orbits["e"] <= 0.4)],
element_type="keplerian",
d_values=[1.7, 2.06]
)
average_orbits_patch = findAverageOrbits(
simulated_ephemeris[simulated_ephemeris_mask],
orbits[(orbits["e"] < 0.5)].reset_index(drop=True),
element_type="keplerian",
d_values=[2.06, 2.5, 2.82, 2.95, 3.27, 5.0, 50.0],
)
average_orbits_patch = pd.concat(
[
average_orbits_hun1_patch,
average_orbits_hun2_patch,
average_orbits_hun3_patch,
average_orbits_patch
],
ignore_index=True
)
average_orbits_patch.loc[:, "exp_mjd_start"] = exp_mjd
average_orbits_patch.insert(0, "patch_number", patch_number)
average_orbits_list.append(average_orbits_patch)
patch_number += 1
average_orbits = pd.concat(average_orbits_list)
average_orbits.sort_values(by=["patch_number", "a_au"], inplace=True)
average_orbits["orbit_id"] = np.arange(1, len(average_orbits) + 1)
average_orbits.reset_index(inplace=True, drop=True)
orbits["i_rad"] = np.radians(orbits["i_deg"])
orbits["ascNode_rad"] = np.radians(orbits["ascNode_deg"])
orbits["argPeri_rad"] = np.radians(orbits["argPeri_deg"])
orbits["meanAnom_rad"] = np.radians(orbits["meanAnom_deg"])
average_orbits = average_orbits.drop(index=average_orbits[average_orbits["designation"].isna()].index)
average_orbits.reset_index(
inplace=True,
drop=True
)
# +
from astropy.time import Time
from thor.orbits import Orbits
average_orbits["epoch"] = Time(average_orbits["exp_mjd"].values, scale="utc", format="mjd").tdb.mjd
average_orbits.rename(
columns={
""
"HEclObj_X_au" : "x",
"HEclObj_Y_au" : "y",
"HEclObj_Z_au" : "z",
"HEclObj_dX/dt_au_p_day" : "vx",
"HEclObj_dY/dt_au_p_day" : "vy",
"HEclObj_dZ/dt_au_p_day" : "vz",
},
inplace=True
)
# -
average_orbits
test_orbits = Orbits.from_df(
average_orbits
)
orbit_file = "/mnt/data/projects/thor/thor_results/ztf/v1.1/test_orbits.csv"
if not os.path.exists(orbit_file):
test_orbits.to_csv(orbit_file)
# +
from thor.orbits import Orbits
# Patches in chunks of 5 were submitted to Hyak, we now combine each patch's recovered orbits
RUN_DIR = "/mnt/data/projects/thor/thor_results/ztf/v1.1/run4/"
patch_orbits = []
patch_orbit_members = []
contents = sorted(glob.glob(os.path.join(RUN_DIR, "patch_*")))
for c in contents:
if os.path.isdir(c):
patch_orbits_i = Orbits.from_csv(
os.path.join(c, "recovered_orbits.csv")
)
patch_orbit_members_i = pd.read_csv(
os.path.join(c, "recovered_orbit_members.csv"),
index_col=False,
dtype={
"obs_id" : str
}
)
patch_orbits.append(
patch_orbits_i.to_df(include_units=False)
)
patch_orbit_members.append(patch_orbit_members_i)
patch_orbits = pd.concat(
patch_orbits,
ignore_index=True
)
patch_orbit_members = pd.concat(
patch_orbit_members,
ignore_index=True
)
# +
from thor.utils import removeDuplicateLinkages
from thor.utils import removeDuplicateObservations
from thor.utils import sortLinkages
recovered_orbits, recovered_orbit_members = removeDuplicateLinkages(
patch_orbits,
patch_orbit_members
)
recovered_orbits, recovered_orbit_members = removeDuplicateObservations(
recovered_orbits,
recovered_orbit_members
)
recovered_orbits, recovered_orbit_members = sortLinkages(
recovered_orbits,
recovered_orbit_members,
preprocessed_observations
)
# +
from thor.orbits import differentialCorrection
recovered_orbits, recovered_orbit_members = differentialCorrection(
recovered_orbits,
recovered_orbit_members,
preprocessed_observations,
min_obs=5,
min_arc_length=1.0,
rchi2_threshold=10,
contamination_percentage=0.0,
delta=1e-8,
max_iter=10,
method="central",
fit_epoch=False,
threads=60,
backend="PYOORB",
)
# +
Orbits.from_df(recovered_orbits).to_csv(
os.path.join(RUN_DIR, "recovered_orbits.csv")
)
recovered_orbit_members.to_csv(
os.path.join(RUN_DIR, "recovered_orbit_members.csv"),
index=False
)
# -
| paper1/ztf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.6 64-bit ('normal')
# metadata:
# interpreter:
# hash: 304dd9ef76ad2d6bf7237ef8f6fde70bef676e0aff923029a3f8742854b86f8f
# name: python3
# ---
import shutil
import os
import pandas as pd
valset_path = 'resources/valset.csv'
data_dir = '../mimic3benchmark/scripts/data/in-hospital-mortality'
out_dir = 'result'
train_listfile_path = os.path.join(data_dir, 'train_listfile.csv')
train_listfile = pd.read_csv(train_listfile_path)
train_listfile = train_listfile.sample(frac=1.0)
length_of_all = len(train_listfile)
length_of_train = int(length_of_all * 0.8)
length_of_val = length_of_all - length_of_train
train_listfile_after_split = train_listfile[: length_of_train]
val_listfile = train_listfile[length_of_train-1: -1]
#print(f'length_of_all: {length_of_all}')
#print(f'train_listfile_after_split length : {len(train_listfile_after_split)}')
#print(f'val_listfile length: {len(val_listfile)}')
assert len(train_listfile_after_split) + len(val_listfile) == length_of_all
# +
listfile_outpath = os.path.join(out_dir, 'mortality')
train_listfile_after_split.to_csv(listfile_outpath + '/train_listfile.csv', index=False)
val_listfile.to_csv(listfile_outpath + '/val_listfile.csv', index=False)
| mimic3models/split_train_val.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: fets
# language: python
# name: fets
# ---
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from feature_engine.encoding import DecisionTreeEncoder
# +
# let's load the data set
data = pd.read_csv("credit_approval_uci.csv")
data.head()
# +
# Let's separate into training and testing sets
X_train, X_test, y_train, y_test = train_test_split(
data.drop(labels=["target"], axis=1), # predictors
data["target"], # target
test_size=0.3, # percentage of observations in test set
random_state=0, # seed to ensure reproducibility
)
X_train.shape, X_test.shape
# +
# Set up the decision encoder
tree_encoder = DecisionTreeEncoder(
encoding_method="arbitrary", # how to convert the strings to numbers
cv=3, # cross-validation
scoring="roc_auc", # the scoring metric to optimise during the grid search
param_grid=None, # defaults to optimizing the tree depth
regression=False,
random_state=10,
variables=None,
)
# +
# let's fit the encoder to the train set
tree_encoder.fit(X_train, y_train)
# -
tree_encoder.variables_
# +
# the pipeline used to encode the categorical
# features
tree_encoder.encoder_
# +
# let's transform the train and test sets
X_train_enc = tree_encoder.transform(X_train)
X_test_enc = tree_encoder.transform(X_test)
# +
# Inspect the encoded variables
X_train_enc[tree_encoder.variables_].head()
# +
# Inspect the encoded variables
X_test_enc[tree_encoder.variables_].head()
# -
| ch02-categorical-encoding/Recipe-10-Encoding-with-decision-trees.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: data-x
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Linear regression & Logistic regression
#
# some exercises on prediction using sklearn.
from IPython.display import display, Latex, Markdown
import seaborn as sns
import csv
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import zipfile
from pathlib import Path
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn import linear_model
from sklearn.metrics import confusion_matrix
from sklearn.metrics import precision_score
import re
from sklearn.model_selection import KFold
#from sklearn.cross_validation import KFold
from sklearn.model_selection import cross_val_score
from sklearn.preprocessing import MinMaxScaler
#
# ### Data:
# __Data Source__:
# Data file is named: __Energy.csv__
#
# The dataset was created by <NAME> ( Civil/Structural Engineer) and was processed by <NAME>, Oxford Centre for Industrial and Applied Mathematics, University of Oxford, UK).
#
# __Data Description__:
#
# The dataset contains eight attributes of a building (or features, denoted by X1...X8) and response being the heating load on the building, y1.
#
# * X1 Relative Compactness
# * X2 Surface Area
# * X3 Wall Area
# * X4 Roof Area
# * X5 Overall Height
# * X6 Orientation
# * X7 Glazing Area
# * X8 Glazing Area Distribution
# * y1 Heating Load
#
df = pd.read_csv('Energy.csv')
display(df.head())
print(df.isnull().sum())
(df.describe()).loc[["min", 'max', '25%', '50%', '75%']]
# __REGRESSION__:
#
# Using the data, we want to predict "Heating load". The output variable is continuous. Hence, we need to use a regression algorithm.
#
# +
#spt = np.random.rand(len(df)) < 0.8
#train = df[spt]
#test = df[~spt]
X = df[['X1','X2','X3','X4','X5','X6','X7','X8']].values
y = df[['Y1']].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state = 5)
regressor = LinearRegression()
regressor.fit(X_train, y_train)
print('intercept:', regressor.intercept_)
print('coefficient values:', regressor.coef_)
# +
#np.sqrt(metrics.mean_squared_error
def RMSE(pred, tar):
return np.sqrt(((pred - tar) ** 2).mean())
q1_3_a = RMSE(regressor.predict(X_train), y_train)
print(' training RMSE:', q1_3_a)
q1_3_b = RMSE(regressor.predict(X_test), y_test)
print(' test RMSE:', q1_3_b)
# -
# Let's see the effect of amount of data on the performance of prediction model. Use varying amounts of data (100,200,300,400,500,all) from the training data used previously to train different regression models.
# +
regressor.fit(X_train[:100], y_train[:100])
xtr1 = RMSE(regressor.predict(X_test), y_test)
x1 = RMSE(regressor.predict(X_train[:100]), y_train[:100])
print("for 100 data points: (Training) ", x1)
print('for 100 data points: (Test)', xtr1)
regressor.fit(X_train[:200], y_train[:200])
x2 = RMSE(regressor.predict(X_train[:200]), y_train[:200])
xtr2 = RMSE(regressor.predict(X_test), y_test)
print("for 200 data points: (Training) ", x2)
print('for 200 data points: (Test)', RMSE(regressor.predict(X_test), y_test))
regressor.fit(X_train[:300], y_train[:300])
x3 = RMSE(regressor.predict(X_train[:300]), y_train[:300])
xtr3 = RMSE(regressor.predict(X_test), y_test)
print("for 300 data points: (Training) ", x3)
print('for 300 data points: (Test)', RMSE(regressor.predict(X_test), y_test))
regressor.fit(X_train[:400], y_train[:400])
x4 = RMSE(regressor.predict(X_train[:400]), y_train[:400])
xtr4 = RMSE(regressor.predict(X_test), y_test)
print("for 400 data points: (Training) ", x4)
print('for 400 data points: (Test)', RMSE(regressor.predict(X_test), y_test))
regressor.fit(X_train[:500], y_train[:500])
x5 = RMSE(regressor.predict(X_train[:500]), y_train[:500])
xtr5 = RMSE(regressor.predict(X_test), y_test)
print("for 500 data points: (Training) ", x5)
print('for 500 data points: (Test)', RMSE(regressor.predict(X_test), y_test))
regressor.fit(X_train, y_train)
xall = RMSE(regressor.predict(X_train), y_train)
xtrall = RMSE(regressor.predict(X_test), y_test)
print("for all data points: (Training) ", xall)
print('for all data points: (Test)', RMSE(regressor.predict(X_test), y_test))
print()
print("the more data I have, my error increases")
value = [x1, x2, x3, x4, x5, xall]
value2 = [xtr1, xtr2, xtr3, xtr4, xtr5, xtrall,]
nums = [100, 200, 300, 400, 500, len(X_train)]
plt.plot(nums, value, label = 'train')
plt.plot(nums, value2, label = 'test')
plt.legend()
plt.show()
# -
#
# __CLASSIFICATION__:
# LABELS ARE DISCRETE VALUES.
#
# Here the model is trained to classify each instance into a set of predefined discrete classes. On inputting a feature vector into the model, the trained model is able to predict a class of that instance.
#
# Bucket the values of 'y1' i.e 'Heating Load' from the original dataset into 3 classes:
#
# 0: 'Low' ( < 14),
# 1: 'Medium' (14-28),
# 2: 'High' (>28)
#
df['class'] = pd.cut(df["Y1"], [0,14,28, np.inf], include_lowest = True, labels = ["Low", 'Medium', 'High'])
df.head()
X = df[['X1','X2','X3','X4','X5','X6','X7','X8']]
y = df['class']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state = 5)
LogisticRegressionModel = linear_model.LogisticRegression()
LogisticRegressionModel.fit(X_train, y_train)
# - Print the training and test accuracies
# - Print the confusion matrix
# - Print the precision and recall numbers for all the classes
#
training_accuracy=LogisticRegressionModel.score(X_train,y_train)
print ('Training Accuracy:',training_accuracy)
test_accuracy=LogisticRegressionModel.score(X_test,y_test)
print ('Test Accuracy:',test_accuracy)
print()
y_true = y_test
y_pred = LogisticRegressionModel.predict(X_test)
ConfusionMatrix=pd.DataFrame(confusion_matrix(y_true, y_pred),columns=['Predicted Low','Predicted Medium','Predicted High'],index=['Actual Low','Actual Medium','Actual High'])
print ('Confusion matrix of test data is: \n',ConfusionMatrix)
print()
print("Average precision for the 3 classes is - ", precision_score(y_true, y_pred, average = None))
print()
from sklearn.metrics import recall_score
print("Average recall for the 3 classes is - ", recall_score(y_true, y_pred, average = None))
# ##### K Fold Cross Validation
#
# In k-fold cross-validation, the shuffled training data is partitioned into k disjoint sets and the model is trained on k −1 sets and validated on the kth set. This process is repeated k times with each set chosen as the validation set once. The cross-validation accuracy is reported as the average accuracy of the k iterations
#
X = df[['X1','X2','X3','X4','X5','X6','X7','X8']]
y = df['class']
F = KFold(n_splits=7, random_state=5, shuffle = True)
LM = linear_model.LogisticRegression()
scores = cross_val_score(LM, X, y, cv = F)
print('Cross-validated scores:', scores)
np.mean(scores)
# One of the preprocessing steps in Data science is Feature Scaling i.e getting all our data on the same scale by setting same Min-Max of feature values.
# This makes training less sensitive to the scale of features .
# Scaling is important in algorithms that use distance functions as a part of classification. If we Scale features in the range [0,1] it is called unity based normalization.
#
# __Performing unity based normalization on the above dataset and train the model
from sklearn import preprocessing
min_max_scaler = preprocessing.MinMaxScaler()
y = df['class']
X = df[['X1','X2','X3','X4','X5','X6','X7','X8']]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state = 5)
X_train_minmax = min_max_scaler.fit_transform(X_train)
X_test_minmax = min_max_scaler.fit_transform(X_test)
LogisticRegressionModel = linear_model.LogisticRegression()
LogisticRegressionModel.fit(X_train_minmax, y_train)
training_accuracy=LogisticRegressionModel.score(X_train_minmax,y_train)
print ('Training Accuracy:',training_accuracy)
test_accuracy=LogisticRegressionModel.score(X_test_minmax,y_test)
print ('Test Accuracy:',test_accuracy)
print()
y_true = y_test
y_pred = LogisticRegressionModel.predict(X_test_minmax)
ConfusionMatrix=pd.DataFrame(confusion_matrix(y_true, y_pred),columns=['Predicted Low','Predicted Medium','Predicted High'],index=['Actual Low','Actual Medium','Actual High'])
print ('Confusion matrix of test data is: \n',ConfusionMatrix)
print()
print("Average precision for the 3 classes is - ", precision_score(y_true, y_pred, average = None))
print()
from sklearn.metrics import recall_score
print("Average recall for the 3 classes is - ", recall_score(y_true, y_pred, average = None))
print("It works significantly better as compared to q2.2 for the test accuracy but the training accuracy is very slightly worse. It was 0.8078175895765473 for Training Accuracy before and 0.7857142857142857 Test Accuracy before." )
"But if I change my random state to say a 100 I cansee my accuracies improve a lot. "
| Regression/Linear & Logistic regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/trista-paul/DS-Unit-2-Sprint-3-Advanced-Regression/blob/master/Survival_Analysis_Assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="gfECCxqbvpjB" colab_type="code" colab={}
#Lecture summary
#Probability density function
#Example: Hazard function
#"instantaneous likelihood of failure" ex.lifetime mortality U curve
#Cumulative Distribution Function
#CDF is the integral of the PDF
#it is the area under the curve of sigmoid
#y P event less than or equal to x
#example: Survival function
#Survival analysis is applied to retention; also known as time to event
#birth, death are general terms for the interval
#birth: person issued welfare
#death: person improoves income and is no longer eligible
#survival answers how long the interval they are on welfare
#'data censorship' occurs when subject doesn't trigger death event
#or literally dies
#logistic reg doesn't have a way to interpret censorship, and survival barely does
#defaulting end of period as death or dropping are both bad
# + [markdown] id="8tMyVxHRa3cQ" colab_type="text"
# # Assignment - Customer Churn
#
# Treselle Systems, a data consulting service, [analyzed customer churn data using logistic regression](http://www.treselle.com/blog/customer-churn-logistic-regression-with-r/). For simply modeling whether or not a customer left this can work, but if we want to model the actual tenure of a customer, survival analysis is more appropriate.
#
# The "tenure" feature represents the duration that a given customer has been with them, and "churn" represents whether or not that customer left (i.e. the "event", from a survival analysis perspective). So, any situation where churn is "no" means that a customer is still active, and so from a survival analysis perspective the observation is censored (we have their tenure up to now, but we don't know their *true* duration until event).
#
# Your assignment is to [use their data](https://github.com/treselle-systems/customer_churn_analysis) to fit a survival model, and answer the following questions:
#
# - What features best model customer churn?
# - What would you characterize as the "warning signs" that a customer may discontinue service?
# - What actions would you recommend to this business to try to improve their customer retention?
#
# Please create at least *3* plots or visualizations to support your findings, and in general write your summary/results targeting an "interested layperson" (e.g. your hypothetical business manager) as your audience.
#
# This means that, as is often the case in data science, there isn't a single objective right answer - your goal is to *support* your answer, whatever it is, with data and reasoning.
#
# Good luck!
# + id="piO805esv_YF" colab_type="code" outputId="94e6a47f-d7be-437e-ecbb-7e6c1028fda8" colab={"base_uri": "https://localhost:8080/", "height": 336}
# !pip install lifelines
# + id="xrkqTt0_wSdq" colab_type="code" colab={}
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import lifelines
cph = lifelines.CoxPHFitter()
# + id="_48G5Nkvv-SK" colab_type="code" outputId="dea9dac7-d464-4bb5-cac1-21bebabec3e9" colab={"base_uri": "https://localhost:8080/", "height": 299}
# Loading the data to get you started
df = pd.read_csv(
'https://raw.githubusercontent.com/treselle-systems/'
'customer_churn_analysis/master/WA_Fn-UseC_-Telco-Customer-Churn.csv')
pd.set_option('display.max_columns', None)
df.head()
# + id="kL-PnLZlLVas" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 215} outputId="7da8f500-04a0-4a7a-e917-5287e914f8b1"
#cell that does all cleaning operations at once
#key for classes below
#I erased the work because I redid the cleaning from a new angle Wednesday night
#and the notebook was huge and depressing
#basics - drop id and convert totalcharge to float
df = df.drop(columns = ['customerID'])
df['TotalCharges'] = pd.to_numeric(df['TotalCharges'], errors='coerce')
#convert special classes into numeric
Classes = ['gender', 'InternetService', 'Contract', 'PaymentMethod']
classes = df[Classes]
df = df.drop(columns=Classes)
for label, col in classes.iteritems():
n = 0
for types in col.unique():
col = col.replace({types:n})
n = n+1
classes[label] = col
#manually assign Internet Service so None is 0
classes['InternetService'] = classes['InternetService'].replace({2:0, 0:2})
#convert remaining objects, binaries, into 0 for No and 1 for Yes
objects = df.select_dtypes(include=object)
notobjects = df.select_dtypes(exclude=object)
objects = objects.replace({'Yes': str(1), 'No': str(0), 'No internet service':str(0), 'No phone service':str(0)})
for label, col in objects.iteritems():
col = pd.to_numeric(col, errors='coerce')
objects[label] = col
#concat objects, notobjects and classes
df = pd.concat([objects, notobjects], axis=1)
df = pd.concat([df, classes], axis=1)
df = df.dropna()
df.head()
# + id="DAotjdS7JWr6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 655} outputId="d4707e90-6cc9-46df-cb06-cee4b6c88640"
#convert special classes into numeric
Classes = ['gender', 'InternetService', 'Contract', 'PaymentMethod']
classes = df[Classes]
for label, col in classes.iteritems():
print(col.value_counts())
n = 0
for types in col.unique():
col = col.replace({types:n})
n = n+1
classes[label] = col
print(classes[label].value_counts())
# + id="5VGOVTRmLwsi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 185} outputId="c74e8295-edc1-44f5-ec87-385b8a27178f"
classes['InternetService'] = classes['InternetService'].replace({2:0, 0:2})
classes['InternetService'].value_counts()
# + id="ym_LNHpmwCST" colab_type="code" outputId="3d7bfaf3-906b-4006-8b37-20357f76bb42" colab={"base_uri": "https://localhost:8080/", "height": 437}
df.info()
# + id="97TdHRv-PRUh" colab_type="code" outputId="0bed6474-d7e4-4339-c8b5-c8d54c0d095c" colab={"base_uri": "https://localhost:8080/", "height": 304}
df.describe()
# + id="bqai_UnpaJeb" colab_type="code" outputId="810bbf34-0608-4932-a0b1-979b1a2688f8" colab={"base_uri": "https://localhost:8080/", "height": 538}
cph = lifelines.CoxPHFitter()
cph.fit(df, 'tenure', event_col='Churn')
cph.print_summary()
#something is happening with onlinebackup, partner and phoneservice.
#their likelihood to cancel increases in response to small changes in unit
#probably irrelevant (likelihood change is or near 1 w low coefs):
#totalcharges, gender, senior, monthlycharges
#low P club! (under 0.05)
#let's start improving the model by reducing features to these
#Partner, PhoneService, MultipleLines, OnlineSecurity, OnlineBackup,
#TechSupport, Contract
#PaperlessBilling, PaymentMethod, MonthlyCharges
#TotalCharges just seems to act unusual
# + id="S54c3_eAcB9F" colab_type="code" outputId="ce61f1ee-d1c1-46c0-d88c-7edd59c403c3" colab={"base_uri": "https://localhost:8080/", "height": 77}
LowPClub = ['tenure', 'Churn',
'Partner', 'PhoneService', 'MultipleLines', 'OnlineSecurity',
'OnlineBackup', 'TechSupport', 'Contract', 'PaperlessBilling',
'PaymentMethod', 'MonthlyCharges']
dfnew = df[LowPClub]
dfnew.head(1)
# + id="PhTU6CpgfyUF" colab_type="code" outputId="4d8c526f-db16-4fdb-84e8-9c5ca2586fbf" colab={"base_uri": "https://localhost:8080/", "height": 403}
cph.fit(dfnew, 'tenure', event_col='Churn')
cph.print_summary()
#like I thought MonthlyCharges wasn't really that relevant
#but had inflated P significance
# + id="G0rAqRYRhAxK" colab_type="code" outputId="0c89d12c-c631-4823-b2d5-c6c411e56566" colab={"base_uri": "https://localhost:8080/", "height": 386}
dfnew = dfnew.drop(columns=['MonthlyCharges', 'Contract'])
cph.fit(dfnew, 'tenure', event_col='Churn')
cph.print_summary()
#The exp(coef) of these features is closer to reality now
#PhoneService, OnlineBackup and Partner are largest changes
# + id="4Tn53SMmh8QK" colab_type="code" outputId="a401bda7-f947-4260-f85c-1851f12d9acb" colab={"base_uri": "https://localhost:8080/", "height": 411}
ax = cph.plot();
plt.xlim(-2, 2)
ax.tick_params(axis='both', labelcolor='black')
ax.text(x = -2, y=9.5, s='Monthly payments, not having a cloud and not sharing',
fontsize=18, fontweight='bold');
ax.text(x = -2, y=9, s='with a partner are the main factors why customers churn.',
fontsize=18, fontweight='bold');
ax.text(x = -2, y=8.5, s='Plot of cox proportional hazards for features with a P<0.01 significance',
fontsize=15, color='#424242');
#the confidence interval of feature coefficient
# + id="Q5k3ymKYQIX2" colab_type="code" outputId="4f3dd7f0-61e8-4fc1-db68-20e3b1c6684a" colab={"base_uri": "https://localhost:8080/", "height": 151}
# !pip install -U matplotlib
# + id="fe0oYap4lTBt" colab_type="code" outputId="f3e250a7-83d1-4d85-8291-9d97437c3fad" colab={"base_uri": "https://localhost:8080/", "height": 34}
time = dfnew.tenure.values
event = dfnew.Churn.values
kmf = lifelines.KaplanMeierFitter()
kmf.fit(time, event_observed=event)
# + id="k7LFGySWQKYv" colab_type="code" outputId="a7942f42-1bdf-4b5a-ea54-ec1d5e49982e" colab={"base_uri": "https://localhost:8080/", "height": 415}
plt.style.use('ggplot')
ax = kmf.survival_function_.plot()
plt.ylim(0.00, 1.00)
plt.xlim(0, 80)
ax.tick_params(axis='both', labelcolor='black')
ax.text(x = 0, y=1.13, s='Customers are still 60% likely to be subscribed after 72 months',
fontsize=18, fontweight='bold');
ax.text(x = 0, y=1.08, s='Kaplan-Meier survival curve of probability of not having churned',
fontsize=15, color='#424242');
ax.text(x = 0, y=1.03, s='against months subscribed',
fontsize=15, color='#424242');
# + id="S8FITHPFQpt9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 407} outputId="d53100f6-1db4-47f7-a33e-689a03cae9c4"
ax = cph.plot_covariate_groups(covariates='Partner', values=[0,1]);
ax.text(x = 0, y=1.13, s='Its 1:30 AM this is as nice as these will get',
fontsize=18, fontweight='bold');
# + id="ktIk4IBybibH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 398} outputId="2f7af391-1e35-4d83-8880-6c9f07d4e9c0"
ax = cph.plot_covariate_groups(covariates='Contract', values=np.arange(0,2,1));
ax.text(x = 0, y=1.13, s='Contract spread of survival by bin much larger than for bools',
fontsize=18, fontweight='bold');
ax.text(x = 0, y=1.08, s='Indicates a dramatic effect with little overlap between dead and survival groups',
fontsize=15, color='#424242');
#spread of survival by bin nearly identical to partner
# + id="Nk2lIZj2cGVn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 380} outputId="52a6cc75-faf9-494b-b4be-9d27a05a4a4b"
cph.plot_covariate_groups(covariates='OnlineBackup', values=[0,1])
#spread of survival by bin nearly identical to partner
# + [markdown] id="kjKxnx4RuzWD" colab_type="text"
# Most of the features I looked at are what's called 'binary' or 'boolean' in the programming world - they either have a yes or no answer. Their numeric symbolism is 1 and 0. Two of the most impactful features on churn (above 0.5 or below -0.5 on the Cox hazards graph), sharing with a partner or using the phone service, are boolean. Our negative placement of partner on the cox hazards graph means that we are more at risk of losing a customer if they are not sharing it with another adult. This makes intuitive sense and there isn't a lot that can be done about it other than making our online service show more Hot Singles In Your Area (don't do this). OnlineBackup has a nearly identical placement, but's its worth saying it was originally three conditions (cloud - 1, not having cloud but having internet service - 0, not having internet service - 0). Pure booleans like partner sharing are probably slightly more accurate models of churn than compressed ones like online backup use or pure categorical classes like contract type.
#
# By far the most dramatic feature either direction is the contract type, where I mapped 0 (and that strongly negative trend) to the **month-by-month payment**, 1 to payment per 1 year, and 2 to payment by 2 years. This also makes sense, as there are more opportunities to withdrawl than with annual plans.
#
# I advise you to look into how we incentivize customers, particularly existing monthly customers, to our annual plans and cloud services. This strategy could look like offering discounts on cloud frequently and marketing the pricing-per-month of the annual plans as clearly more cost-effective than the monthly or the competition.
| Survival_Analysis_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pylab inline
import jax.numpy as np
import jax
points = np.array([[i, np.sin(i / 3.0), np.cos(i / 2)] for i in range(0, 11)])
knots = np.array([0, 0, 0, 0, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 1.0, 1.0, 1.0, 1.0])
knots.shape
knots[6], knots[7]
points.shape
def deBoor(k, x, t, c, p):
"""
Evaluates S(x).
Args
----
k: index of knot interval that contains x
x: position
t: array of knot positions, needs to be padded as described above
c: array of control points
p: degree of B-spline
"""
d = [c[j + k - p] for j in range(0, p+1)]
for r in range(1, p+1):
for j in range(p, r-1, -1):
alpha = (x - t[j+k-p]) / (t[j+1+k-r] - t[j+k-p])
d[j] = (1.0 - alpha) * d[j-1] + alpha * d[j]
return d[p]
# Can we do a vectorized version of that?
def deBoorVectorized(x, t, c, p):
"""
Evaluates S(x).
Args
----
x: position
t: array of knot positions, needs to be padded as described above
c: array of control points
p: degree of B-spline
"""
k = np.digitize(x, t) -1
d = [c[j + k - p] for j in range(0, p+1)]
for r in range(1, p+1):
for j in range(p, r-1, -1):
alpha = (x - t[j+k-p]) / (t[j+1+k-r] - t[j+k-p])
d[j] = (1.0 - alpha) * d[j-1] + alpha * d[j]
return d[p]
deBoor(6, 0.45, knots, points[:,1], 3 )
x = np.linspace(0,1.,100, endpoint=False)
y = deBoorVectorized(x, knots, points[:,1], 3 )
plot(x, y)
y
x
| notebooks/DeBoor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/agemagician/CodeTrans/blob/main/prediction/multitask/fine-tuning/commit%20generation/small_model.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="c9eStCoLX0pZ"
# **<h3>Generate the commit for github code changes using codeTrans multitask finetuning model</h3>**
# <h4>You can make free prediction online through this
# <a href="https://huggingface.co/SEBIS/code_trans_t5_small_commit_generation_multitask_finetune">Link</a></h4> (When using the prediction online, you need to parse and tokenize the code first.)
# + [markdown] id="6YPrvwDIHdBe"
# **1. Load necessry libraries including huggingface transformers**
# + id="6FAVWAN1UOJ4" colab={"base_uri": "https://localhost:8080/"} outputId="6f3593c3-b21c-431f-c80f-957d0b8e057a"
# !pip install -q transformers sentencepiece
# + id="53TAO7mmUOyI"
from transformers import AutoTokenizer, AutoModelWithLMHead, SummarizationPipeline
# + [markdown] id="xq9v-guFWXHy"
# **2. Load the token classification pipeline and load it into the GPU if avilabile**
# + colab={"base_uri": "https://localhost:8080/", "height": 316, "referenced_widgets": ["17d7ac880f1b4204bad7cf6dfbc8146b", "98e879d8002b4237aa137e36af7dfffa", "3274e44ad1014402a53d11e4eaae7a7b", "d512b0dab64049aaa2dd824e9e224ca2", "9e78956e855843d2823a9c64d4c6c7fd", "37463a48920f4bbc8f7e53445b25e00a", "e38c21e10e144eb19102fb17758a8a1c", "590f49f12afd49d8ad579ec7cf8fffe3", "e22dc23ff5f24c15b7b8f701f353811d", "babe92f28d3e4083aa47fffc3b13ec03", "96a701545e2d45b2b00272040c39d0d7", "6e5013f7150d45c4bb1888df3fd70a5c", "a23b3fe205c94d0a89a6470d7657c15a", "a3c706fe7a35437183f85c057b8ecc41", "4b4d88dd993b4f23ab1b3d3f8f3b7b15", "ae5db7956d4045428c70c07aa0542396", "680491e134924675be82e4cfbb1a7d03", "<KEY>", "22c04615ae7a4418850f108ff21433a7", "<KEY>", "<KEY>", "b5a97444838d47968827a3bd10580067", "1b89134e6c124902a08b5ce353a4c63e", "<KEY>", "f75c3a5d64a8410b9bbb7250c4770cef", "<KEY>", "05a0d3e153a64557bb86ba3d701dc6c3", "<KEY>", "03ee927700354efa8d558fe0add5dcad", "<KEY>", "<KEY>", "<KEY>", "7f04ee84f37d48af8401509fe76f1ff0", "<KEY>", "<KEY>", "<KEY>", "a1fed4b0443a47ecaedd2e773a8fac8c", "<KEY>", "<KEY>", "<KEY>"]} id="5ybX8hZ3UcK2" outputId="a7ceed4e-0dc0-4bbf-e52d-32f056abe251"
pipeline = SummarizationPipeline(
model=AutoModelWithLMHead.from_pretrained("SEBIS/code_trans_t5_small_commit_generation_multitask_finetune"),
tokenizer=AutoTokenizer.from_pretrained("SEBIS/code_trans_t5_small_commit_generation_multitask_finetune"),
device=0
)
# + [markdown] id="hkynwKIcEvHh"
# **3 Give the code for summarization, parse and tokenize it**
# + id="nld-UUmII-2e"
code = "new file mode 100644\n index 000000000 . . 892fda21b\n Binary files/dev/null and b/src/plugins/gateway/lib/joscar.jar differ\n" #@param {type:"raw"}
# + id="hqACvTcjtwYK" colab={"base_uri": "https://localhost:8080/"} outputId="bf908c1e-14b3-4996-e733-ed1370270abf"
import nltk
nltk.download('punkt')
from nltk.tokenize import WordPunctTokenizer
tokenized_list = WordPunctTokenizer().tokenize(code)
tokenized_code = ' '.join(tokenized_list)
print("tokenized code: " + tokenized_code)
# + [markdown] id="sVBz9jHNW1PI"
# **4. Make Prediction**
# + colab={"base_uri": "https://localhost:8080/"} id="KAItQ9U9UwqW" outputId="4240465d-4dea-436a-aceb-396f54b43b59"
pipeline([tokenized_code])
| prediction/multitask/fine-tuning/commit generation/small_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1><b>HomeWork Weekly (LINKED LIST) 3-6</b></h1>
# ## Linked List
# adalah suatu struktur data linier, berbeda dengan list yang juga merupakan struktur data linier dan tipe data komposit, linked list dibentuk secara dinamik. Pada saat awal program dijalankan elemen linked list sebelum data, Elemen linked list (disebut node) dibentuk sambil jalan sesuai instruksi. Apabila setiap elemen dapat diakses secara langsung dengan menggunakan indeks, sebuah node linked list diakses dengan menggunakan pointer yang mengacu (menunjuk) ke node tersebut<br><p>
# <img src="images.png">
# <center>Contoh gambar Linked List<center><p>
# <b>Berikut implementasi dalam syntax :</b>
class Node:
def __init__(self, init_data):
self.data = init_data
self.next = None
def getData(self):
return self.data
def getNext(self):
return self.next
def setData(self, newData):
self.data = newData
def setNext(self, newNext):
self.next = newNext
# <p><p><b>Beberapa operasi list tak-berurut (unordered) adalah : </b>
# <ul><li>List() membuat suatu list baru yang kosong. Tidak memerlukan parameter dan
# mengembalikan suatu list kosong.</ul>
# <ul><li>add(item) menambahkan suatu item baru ke dalam list. Diperlukan item yang akan
# ditambahkan dan tidak mengembalikan apapun. Anggapan: item tersebut belum ada
# dalam list.</ul>
# <ul><li>remove(item) menghapus item dari dalam list. Diperlukan item dan akan mengubah
# list. Anggapan: item tersebut tersedia di dalam list.
# search(item) mencari item di dalam list. Perlu item dan mengembalikan suatu nilai
# boolean.</ul>
# <ul><li>isEmpty() menguji untuk melihat apakah list dalam keadaan kosong (empty). Tidak
# memerlukan parameter dan mengembalikan suatu nilai boolean.</ul>
# <ul><li>size() mengembalikan jumlah item di dalam list. Tidak memerlukan parametyer dan
# mengembalikan suatu integer.</ul>
# <ul><li>append(item) menambahkan item baru ke akhir (ujung) list dan menjadikannya
# sebagai item terakhir di dalam koleksi. Diperlukan item dan tidak mengembalikan
# apapun. Anggapan: item tersebut belum ada di dalam list.</ul>
# <ul><li>index(item) mengembalikan posisi dari item di dalam list. Diperlukan item dan
# mengembalikan index dari item tersebut. Anggapan: item terdapat di dalam list.
# insert(pos, item) menambahkan suatu item baru ke dalam list pada posisi pos.
# Diperlukan item dan tidak mengembalikan apapun. Anggapan: item belum terdapat di
# dalam list dan ada cukup item existing (telah ada di dalam list) sehingga dapat
# diketahui posisi pos.</ul>
# <ul><li>pop() menghapus dan mengembalikan item terakhir di dalam list. Tidak memerlukan
# parameter dan mengembalikan suatu item. Anggapan: list tidak kosong, setidaknya
# ada satu item.</ul>
# <ul><li>pop(pos) menghapus dan mengembalikan item pada posisi pos. Diperlukan posisi
# sebagai parameter dan mengembalikan suatu item. Anggapan: item ada dalam list.</ul><br>
# +
class LinkedList:
def __init__(self):
self.head = None
def isEmpty(self):
return self.head == None
def add(self, item):
temp = Node(item)
temp.setNext(self.head)
self.head = temp
def size(self):
current = self.head
count = 0
while current != None:
count +=1
current = current.getNext()
return count
def search(self, item):
current = self.head
found = False
hit = 0
while current != None and not found:
hit+=1
if current.getData() == item:
found = True
print ("Node Position : ", hit)
else:
current = current.getNext()
return found
rlist= LinkedList()
rlist.add(54)
# print(rlist.head)
rlist.add(34)
rlist.add(70)
print("___________")
print(rlist.search(54))
print("___________")
# print(rlist.search(54))
print(rlist.size())
# print(rlist.head.da)
print("------")
data = Node(93)
data1 = Node(23)
print(data.getData())
print(data1.getNext())
data.setNext(data1)
print(data.getNext().getData())
print(data.getNext().getNext())
# print(data.getNext().getData())
# -
# > Kelas Node dan UnorderedList (Linked List) yang mengimplementasikan suatu linked-list (list
# berkait atau bersambungan). Perlu ditentukan nilai data awal untuk node. Juga ada metode
# untuk mengakses dan mengubah datanya dan referensi berikutnya. Perhatikan konstruktor,
# metode add, search, remove dan bagaimana itu semua digunakan.
# <p><p><b>Implementasi add(item)</b>
def add(self, item):
temp = Node(item)
temp.setNext(self.head)
self.head = temp
# <p><p><center><img src="img_add.png"></center>
# <p><p><b>Implementasi Search(item)</b>
def search(self, item):
current = self.head
found = False
hit = 0
while current != None and not found:
hit+=1
if current.getData() == item:
found = True
print ("Node Position : ", hit)
else:
current = current.getNext()
return found
# <p><p><center><img src="img_search.png"></center><p>
| Minggu 4-6/Linked_List.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''tesi'': conda)'
# language: python
# name: python3
# ---
# # Testing DEMV on Drug Dataset
# - Sensitive group: white women (`race: 1, gender: 0`)
# - Positive label: never (0)
# +
import pandas as pd
import numpy as np
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
from fairlearn.reductions import ExponentiatedGradient, BoundedGroupLoss, ZeroOneLoss, GridSearch
import matplotlib.pyplot as plt
import seaborn as sns
from utils import *
from demv import DEMV
import warnings
warnings.filterwarnings('ignore')
sns.set_style('whitegrid')
# -
def prepare_data():
data = pd.read_csv('data2/drugs.csv')
data.drop(['yhat','a'], axis=1, inplace=True)
data.loc[data['gender']==0.48246,'gender']=1
data.loc[data['gender']==-0.48246,'gender']=0
data['y'].replace({
'never': 0,
'not last year': 1,
'last year': 2}, inplace=True)
data['race'].replace({
'non-white': 0,
'white': 1}, inplace=True)
string_cols = data.dtypes[data.dtypes == 'object'].index.values
data.drop(string_cols, axis=1, inplace=True)
return data
data = prepare_data()
data
label = 'y'
protected_group = {'race': 1, 'gender': 0}
positive_label = 0
sensitive_features = ['race', 'gender']
pipeline = Pipeline([
('scaler', StandardScaler()),
('classifier', LogisticRegression(n_jobs=-1))
])
# ## Biased classifier
model, lr_bias = cross_val(pipeline, data, label, protected_group, sensitive_features, positive_label)
print_metrics(lr_bias)
# ## DEMV application
demv = DEMV(round_level=1)
demv_data = data.copy()
model, lr_demv = cross_val(pipeline, demv_data, label, protected_group, sensitive_features, debiaser=demv, positive_label=positive_label)
print_metrics(lr_demv)
# ## DEMV Evaluation
demv.get_iters()
metrics = eval_demv(17, 89, data.copy(), pipeline, label, protected_group, sensitive_features, positive_label=positive_label)
# ## Blackbox Postprocessing
model, blackboxmetrics, pred = cross_val2(pipeline, data, label, protected_group, sensitive_features, positive_label=positive_label)
# ## Plot
df = prepareplots(metrics, 'drugs')
# df = pd.read_csv('ris/drugs_eval.csv')
points = preparepoints(blackboxmetrics, 85)
plot_metrics_curves(df, points, 'Drug')
unprivpergentage(data, protected_group, 89)
save_metrics('blackbox', 'drug', blackboxmetrics)
| drug.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# New DG Food Agro are a multinational exporter of various grains from India since nearly 130 years. But their main product of exporting since early 1980s has been Wheat. They export wheat to countries like America, Afghanistan, Australia etc. They started seeing varying exports of sales year on year for various countries. The reason that was theorized by them had a lot of natural causes like floods, country growth, population explosion etc. Now they need to decide which countries fall in the same range of export and which don’t. They also need to know which countries export is low and can be improved and which countries are performing very well across the years.
# The data provided right now is across 18 years. What they need is a repeatable solution which won’t get affected no matter how much data is added across time and that they should be able to explain the data across years in less number of variables.
# Objective: Our objective is to cluster the countries based on various sales data provided to us across years. We have to apply an unsupervised learning technique like K means or Hierarchical clustering so as to get the final solution. But before that we have to bring the exports (in tons) of all countries down to same scale across years. Plus, as this solution needs to be repeatable we will have to do PCA so as to get the principal components which explain max variance.
# Implementation:
#
# 1) Read the data file and check for any missing values
#
# 2) Change the headers to country and year accordingly.
#
# 3) Cleanse the data if required and remove null or blank values
#
# 4) After the EDA part is done, try to think which algorithm should be applied here.
#
# 5) As we need to make this across years we need to apply PCA first.
#
# 6) Apply PCA on the dataset and find the number of principal components which explain nearly all the variance.
#
# 7) Plot elbow chart or scree plot to find out optimal number of clusters.
#
# 8) Then try to apply K means, Hierarchical clustering and showcase the results.
#
# 9) You can either choose to group the countries based on years of data or using the principal components.
#
# 10) Then see which countries are consistent and which are largest importers of the good based on scale and position of cluster.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# 1. Read the data file and check for any missing values
dfdata = pd.read_csv('542_proj_dataset_2_v1.0\Project_Data_2.csv')
dfdata.head()
dfdata.shape
len(dfdata['Sales of Wheat in tons'].unique())
dfdata.info()
# 3. Cleanse the data if required and remove null or blank values
dfdata['1990'] = dfdata['1990'].str.replace(',', '').astype('int64')
dfdata['1991'] = dfdata['1991'].str.replace(',', '').astype('int64')
dfdata['1992'] = dfdata['1992'].str.replace(',', '').astype('int64')
dfdata['1993'] = dfdata['1993'].str.replace(',', '').astype('int64')
dfdata['1994'] = dfdata['1994'].str.replace(',', '').astype('int64')
dfdata['1995'] = dfdata['1995'].str.replace(',', '').astype('int64')
dfdata['2004'] = dfdata['2004'].str.replace(',', '').astype('int64')
dfdata['2005'] = dfdata['2005'].str.replace(',', '').astype('int64')
dfdata['2006'] = dfdata['2006'].str.replace(',', '').astype('int64')
dfdata['2007'] = dfdata['2007'].str.replace(',', '').astype('int64')
dfdata.info()
# 2. Change the headers to country and year accordingly.
dfdata.rename(columns = {"Sales of Wheat in tons": "Country"},
inplace = True)
dfdata.head()
# +
# 4. After the EDA part is done, try to think which algorithm should be applied here.
# 5. As we need to make this across years we need to apply PCA first.
# -
from sklearn.decomposition import PCA
pca = PCA(n_components=10)
# +
# 6. Apply PCA on the dataset and find the number of principal components which explain nearly all the variance.
# 7. Plot elbow chart or scree plot to find out optimal number of clusters.
# -
X = dfdata.drop('Country', axis =1)
pca.fit(X)
pca.explained_variance_ratio_*100
plt.plot(pca.explained_variance_ratio_*100) #elbow graph
dftransformed = pd.DataFrame(pca.transform(X))
dftransformed.head()
dftransformedRequired = dftransformed[[0,1]]
dftransformedRequired.head()
# +
# 8. Then try to apply K means, Hierarchical clustering and showcase the results.
# 9. You can either choose to group the countries based on years of data or using the principal components.
# 10. Then see which countries are consistent and which are largest importers of the good based on scale and position of cluster.
# -
from sklearn.cluster import KMeans
kmeans = KMeans(n_clusters=4)
kmeans.fit(dftransformedRequired)
kmeans.cluster_centers_
kmeans.labels_
dftransformedRequired['Class'] = kmeans.labels_
dftransformedRequired.head()
dftransformedRequired.rename(columns={0: "col_1",1:"col_2"}, inplace=True)
dftransformedRequired.head()
ax = sns.lmplot("col_1","col_2", hue='Class', data= dftransformedRequired, palette='bright')
plt.figure(figsize=(8,8))
sns.countplot(dftransformedRequired.Class, palette='bright')
dfdata['Class'] = dftransformedRequired['Class']
dfdata.sample(10)
list(dfdata[dfdata['Class']== 2]['Country']) #consistent countries
from scipy.cluster.hierarchy import dendrogram, linkage
from matplotlib import pyplot as plt
f1 = dftransformedRequired['col_1'].values
f2 = dftransformedRequired['col_2'].values
fb = f1[0:10]
fg = f2[0:10]
X = np.array(list(zip(fb, fg)))
Z = linkage(X, 'ward')
fig = plt.figure(figsize = (5,5))
dn = dendrogram(Z)
Z = linkage(X, 'single')
fig = plt.figure(figsize = (5,5))
dn = dendrogram(Z)
| Certification Project/Certification_Project_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Analyzing Probabilities
import pandas as pd
import numpy as np
df = pd.read_csv("Root_Insurance_data.csv")
probRF = pd.read_csv("classes with probability.csv")
df.head()
probRF.head()
# +
_index = [['Y', 1, 1, 'M'],
['Y', 2, 2, 'M'],
['Y', 3, 1, 'S'],
['Y', 1, 2, 'S'],
['Y', 2, 1, 'M'],
['Y', 3, 2, 'M'],
['Y', 1, 1, 'S'],
['Y', 2, 2, 'S'],
['Y', 3, 1, 'M'],
['Y', 1, 2, 'M'],
['Y', 2, 1, 'S'],
['Y', 3, 2, 'S'],
['N', 1, 1, 'M'],
['N', 2, 2, 'M'],
['N', 3, 1, 'S'],
['N', 1, 2, 'S'],
['N', 2, 1, 'M'],
['N', 3, 2, 'M'],
['N', 1, 1, 'S'],
['N', 2, 2, 'S'],
['N', 3, 1, 'M'],
['N', 1, 2, 'M'],
['N', 2, 1, 'S'],
['N', 3, 2, 'S'],
['unknown', 1, 1, 'M'],
['unknown', 2, 2, 'M'],
['unknown', 3, 1, 'S'],
['unknown', 1, 2, 'S'],
['unknown', 2, 1, 'M'],
['unknown', 3, 2, 'M'],
['unknown', 1, 1, 'S'],
['unknown', 2, 2, 'S'],
['unknown', 3, 1, 'M'],
['unknown', 1, 2, 'M'],
['unknown', 2, 1, 'S'],
['unknown', 3, 2, 'S']]
#click = number of clicks for each status
#sold = number of policies sold
#total = number in each category
click = [37, 21, 24, 77, 19, 18, 78, 16, 21, 76, 15, 11, 25, 33, 22, 79, 15, 21, 43, 32, 15, 74, 16, 13, 112, 94, 67, 128, 84, 73, 127, 110, 69, 105, 108, 0]
sold = [11, 8, 5, 24, 6, 2, 38, 3, 4, 24, 4, 4, 14, 15, 8, 32, 6, 11, 25, 15, 9, 31, 5, 3, 59, 26, 26, 55, 41, 32, 69, 42, 25, 57, 44, 0]
total = [274, 317, 267, 282, 284, 276, 290, 304, 294, 295, 266, 270, 300, 296, 286, 283, 263, 326, 271, 316, 265, 275, 272, 291, 294, 306, 256, 303, 268, 277, 280, 288, 287, 294, 284, 0]
#probsold = number sold/10000
#probSoldGivenClick = numbersold/number click
probsold=[]#probably sold
probSoldGivenClick = [] #prob sold given click
for i in range(0,len(click)):
if click[i] !=0:
probsold.append(sold[i]/10000)
probSoldGivenClick.append(sold[i]/click[i])
else:
probsold.append(0)
probSoldGivenClick.append(0)
# +
currentlyinsured, numcars, numdrivers, married = [], [],[],[]
for i in range(0, len(_index)):
currentlyinsured.append(_index[i][0])
numcars.append(_index[i][1])
numdrivers.append(_index[i][2])
married.append(_index[i][3])
index2 = pd.DataFrame({'Currently Insured':currentlyinsured,
'Number of Vehicles':numcars,
'Number of Drivers':numdrivers,
'Marital Status':married})
# -
index2
xdf = pd.DataFrame()
xdf[["insured","unknown"]] = pd.get_dummies(index2["Currently Insured"])[["Y","unknown"]]
xdf[["cars2","cars3"]] = pd.get_dummies(index2["Number of Vehicles"])[[2,3]]
xdf[["drivers2"]] = pd.get_dummies(index2["Number of Drivers"])[[1]]
xdf[["married"]] = pd.get_dummies(index2["Marital Status"])[["M"]]
# +
probforests = []
for i in range(0, 36):
probforests.append(0)
for i in range(0, len(_index)):
for j in range(0, len(_index)):
if xdf.loc[i]['insured'] == probRF.loc[j]['insured'] and xdf.loc[i]['unknown'] == probRF.loc[j]['unknown'] and xdf.loc[i]['cars2'] == probRF.loc[j]['cars2'] and xdf.loc[i]['cars3'] == probRF.loc[j]['cars3'] and xdf.loc[i]['drivers2'] == probRF.loc[j]['drivers2'] and xdf.loc[i]['married'] == probRF.loc[j]['married']:
probforests[i] = probRF['prob_to_buy'][j]
# -
xdf['naive prob'] = probSoldGivenClick
xdf['prob from random forests'] = probforests
xdf['% difference/naive'] = -(xdf['naive prob'] - xdf['prob from random forests'])/xdf['naive prob']
xdf['% difference/forests'] = (xdf['naive prob'] - xdf['prob from random forests'])/xdf['prob from random forests']
xdf.to_csv("probabilities.csv")
| Probabilities/comparing probabilities.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hospital Readmissions Data Analysis and Recommendations for Reduction
#
# ### Background
# In October 2012, the US government's Center for Medicare and Medicaid Services (CMS) began reducing Medicare payments for Inpatient Prospective Payment System hospitals with excess readmissions. Excess readmissions are measured by a ratio, by dividing a hospital’s number of “predicted” 30-day readmissions for heart attack, heart failure, and pneumonia by the number that would be “expected,” based on an average hospital with similar patients. A ratio greater than 1 indicates excess readmissions.
#
# ### Exercise Directions
#
# In this exercise, you will:
# + critique a preliminary analysis of readmissions data and recommendations (provided below) for reducing the readmissions rate
# + construct a statistically sound analysis and make recommendations of your own
#
# More instructions provided below. Include your work **in this notebook and submit to your Github account**.
#
# ### Resources
# + Data source: https://data.medicare.gov/Hospital-Compare/Hospital-Readmission-Reduction/9n3s-kdb3
# + More information: http://www.cms.gov/Medicare/medicare-fee-for-service-payment/acuteinpatientPPS/readmissions-reduction-program.html
# + Markdown syntax: http://nestacms.com/docs/creating-content/markdown-cheat-sheet
# ****
# +
# %matplotlib inline
import pandas as pd
import numpy as np
from scipy import stats
import matplotlib.pyplot as plt
import bokeh.plotting as bkp
from mpl_toolkits.axes_grid1 import make_axes_locatable
# -
# read in readmissions data provided
hospital_read_df = pd.read_csv('data/cms_hospital_readmissions.csv')
# ****
# ## Preliminary Analysis
# deal with missing and inconvenient portions of data
clean_hospital_read_df = hospital_read_df[hospital_read_df['Number of Discharges'] != 'Not Available']
clean_hospital_read_df.loc[:, 'Number of Discharges'] = clean_hospital_read_df['Number of Discharges'].astype(int)
clean_hospital_read_df = clean_hospital_read_df.sort_values('Number of Discharges')
# +
# generate a scatterplot for number of discharges vs. excess rate of readmissions
# lists work better with matplotlib scatterplot function
x = [a for a in clean_hospital_read_df['Number of Discharges'][81:-3]]
y = list(clean_hospital_read_df['Excess Readmission Ratio'][81:-3])
fig, ax = plt.subplots(figsize=(8,5))
ax.scatter(x, y,alpha=0.2)
ax.fill_between([0,350], 1.15, 2, facecolor='red', alpha = .15, interpolate=True)
ax.fill_between([800,2500], .5, .95, facecolor='green', alpha = .15, interpolate=True)
ax.set_xlim([0, max(x)])
ax.set_xlabel('Number of discharges', fontsize=12)
ax.set_ylabel('Excess rate of readmissions', fontsize=12)
ax.set_title('Scatterplot of number of discharges vs. excess rate of readmissions', fontsize=14)
ax.grid(True)
fig.tight_layout()
# -
# ****
#
# ## Preliminary Report
#
# Read the following results/report. While you are reading it, think about if the conclusions are correct, incorrect, misleading or unfounded. Think about what you would change or what additional analyses you would perform.
#
# **A. Initial observations based on the plot above**
# + Overall, rate of readmissions is trending down with increasing number of discharges
# + With lower number of discharges, there is a greater incidence of excess rate of readmissions (area shaded red)
# + With higher number of discharges, there is a greater incidence of lower rates of readmissions (area shaded green)
#
# **B. Statistics**
# + In hospitals/facilities with number of discharges < 100, mean excess readmission rate is 1.023 and 63% have excess readmission rate greater than 1
# + In hospitals/facilities with number of discharges > 1000, mean excess readmission rate is 0.978 and 44% have excess readmission rate greater than 1
#
# **C. Conclusions**
# + There is a significant correlation between hospital capacity (number of discharges) and readmission rates.
# + Smaller hospitals/facilities may be lacking necessary resources to ensure quality care and prevent complications that lead to readmissions.
#
# **D. Regulatory policy recommendations**
# + Hospitals/facilties with small capacity (< 300) should be required to demonstrate upgraded resource allocation for quality care to continue operation.
# + Directives and incentives should be provided for consolidation of hospitals and facilities to have a smaller number of them with higher capacity and number of discharges.
# ****
# ## Exercise
#
# Include your work on the following **in this notebook and submit to your Github account**.
#
# A. Do you agree with the above analysis and recommendations? Why or why not?
#
# B. Provide support for your arguments and your own recommendations with a statistically sound analysis:
#
# 1. Setup an appropriate hypothesis test.
# 2. Compute and report the observed significance value (or p-value).
# 3. Report statistical significance for $\alpha$ = .01.
# 4. Discuss statistical significance and practical significance. Do they differ here? How does this change your recommendation to the client?
# 5. Look at the scatterplot above.
# - What are the advantages and disadvantages of using this plot to convey information?
# - Construct another plot that conveys the same information in a more direct manner.
#
#
#
# You can compose in notebook cells using Markdown:
# + In the control panel at the top, choose Cell > Cell Type > Markdown
# + Markdown syntax: http://nestacms.com/docs/creating-content/markdown-cheat-sheet
# ****
# +
# Your turn
# -
# ### A. Do you agree with the above analysis and recommendations? Why or why not?
#
# **A. Initial observations based on the plot above**
# + Overall, rate of readmissions is trending down with increasing number of discharges
# + With lower number of discharges, there is a greater incidence of excess rate of readmissions (area shaded red)
# + With higher number of discharges, there is a greater incidence of lower rates of readmissions (area shaded green)
# ****
# *-> The first point is very hard to tell on the graph above, especially considering the handfull of outliers in the upper left area, shaded red. Ideally, a correlation and/or regression line should be used to confirm a trend.*
# *- For the second two points these are specific statements about only a subsection of the data. Better would be to categorize 'number of discharges' into low and high, for example, and test whether they are different (e.g. t test).*
#
# **B. Statistics**
# + In hospitals/facilities with number of discharges < 100, mean excess readmission rate is 1.023 and 63% have excess readmission rate greater than 1
# + In hospitals/facilities with number of discharges > 1000, mean excess readmission rate is 0.978 and 44% have excess readmission rate greater than 1
#
# ****
# *-> These are likely accurate statements, but don't seem to originate from statistical tests. If the given threshold values are important, then again, a t-test could be performed to check difference in proportion of excess readmission greater than 1.*
#
# **C. Conclusions**
# + There is a significant correlation between hospital capacity (number of discharges) and readmission rates.
# + Smaller hospitals/facilities may be lacking necessary resources to ensure quality care and prevent complications that lead to readmissions.
#
# ****
# *-> I see no evidence in the report that suggests significant correlation of any factors. A significance test is required. The interpretation of this 'result' is therefore misleading.*
#
# **D. Regulatory policy recommendations**
# + Hospitals/facilties with small capacity (< 300) should be required to demonstrate upgraded resource allocation for quality care to continue operation.
# + Directives and incentives should be provided for consolidation of hospitals and facilities to have a smaller number of them with higher capacity and number of discharges.
#
# ****
# *-> Again, these applications of the 'analysis' are very misleading since they are not based on any significance test.*
# ### B. Provide support for your arguments and your own recommendations with a statistically sound analysis:
#
# B1. Setup an appropriate hypothesis test.
#
# Based on the direction of the provided report, it seems the critical issue is whether the excess readmissions ratio is larger or smaller depending on the size of the hospital. Although we aren't given the size (e.g. number of beds) for each hospital, we are given number of discharges. This number is most likely well correlated with hospital size. The above analysis makes policy implications based on the theshold of 300 discharges, so I will use this same threshold in my own analysis.
#
# Given the data, an appropriate hypothesis test would be:
#
# **Null hypothesis:** Excess readmission ratio for hospitals with discharge rates less than 300 ("small") is the same as that for hospitals with discharge rates greater than 300 ("large").
#
# **Alternative hypothesis:** Excess readmission ratio for hospitals with discharge rates less than 300 ("small") is NOT the same as that for hospitals with discharge rates greater than 300 ("large").
clean_hospital_read_df.head()
#subset dataframe by threshold value 300
small_df = clean_hospital_read_df[clean_hospital_read_df['Number of Discharges'] < 300]
large_df = clean_hospital_read_df[clean_hospital_read_df['Number of Discharges'] > 300]
# +
#define series and means to be used in analysis for excess readmission ratio
small_ratio = small_df['Excess Readmission Ratio']
large_ratio = large_df['Excess Readmission Ratio']
small_ratio_mean = np.mean(small_df['Excess Readmission Ratio'])
large_ratio_mean = np.mean(large_df['Excess Readmission Ratio'])
#define series for discharge values
small_disch = small_df['Number of Discharges']
large_disch = large_df['Number of Discharges']
# -
# B2. Compute and report the observed significance value (or p-value).
#
# Define mean difference
mean_diff = small_ratio_mean - large_ratio_mean
print('Mean difference',mean_diff)
# Print sample size
print(str(large_ratio.shape[0]))
print(str(small_ratio.shape[0]))
SE = mean_diff / np.sqrt(small_ratio.var()/small_ratio.size + large_ratio.var()/large_ratio.size)
print("Standard error:", SE)
p_val = 2*(1-stats.norm.cdf(np.abs(SE)))
p_val
# This p-value is <0.05, and we can accept the alternative hypothesis that readmission rates are different between small and large hospitals.
# B3. Report statistical significance for $\alpha$ = .01.
#
# The p-value also passes the test when considering $\alpha$ = .01.
# B4. Discuss statistical significance and practical significance. Do they differ here? How does this change your recommendation to the client?
#
# Calculate relative difference between groups, percent difference
mean_diff_perc = (mean_diff/small_ratio_mean) * 100
print('Mean percent difference',mean_diff_perc)
# Calculate confidence interval
small_conf = stats.t.interval(0.95, len(small_ratio)-1, loc=small_ratio_mean, scale=stats.sem(small_ratio))
large_conf = stats.t.interval(0.95, len(large_ratio)-1, loc=large_ratio_mean, scale=stats.sem(large_ratio))
print("95% Confidence interval, small hospitals:",small_conf)
print("95% Confidence interval, large hospitals:",large_conf)
# - The mean difference between the two sizes of hospital was 0.014. The increase in readmission for smaller hospitals was about 1% greater than for larger hospitals.
# - While the confidence intervals for readmission rates for small and large hospitals are not overlapping, they are very close.
# - Because the difference in readmission rates is so small, it may NOT be worth spending time and money on addressing this in the manner suggested in the previous analysis, which went as far as suggesting hospital closures. This seems extreme when considering the minor differences.
# B5. Look at the scatterplot above.
# - What are the advantages and disadvantages of using this plot to convey information?
# - Construct another plot that conveys the same information in a more direct manner.
# Advantages:
# - clear labels (axis and chart title)
# - scatter plot style usually allows reader to see all data points
#
# Disadvantages:
# - shaded areas are misleading
# - plot is crowded, and so it is hard to gather information about the general trends (a trendline might help)
# - data are not segmented by categories of interest (e.g. small and large hospitals)
import seaborn as sns
fig, ax = plt.subplots(figsize=(10,10))
sns.boxplot(data=[small_df['Excess Readmission Ratio'],large_df['Excess Readmission Ratio']])
xmin,xmax=ax.get_xlim()
ymin,ymax=ax.get_ylim()
labels=['Small Hospitals','Large Hospitals']
plt.hlines(y=1.0,xmin=xmin,xmax=xmax,color='r')
ax.set_xlabel('Hospital Size',fontsize=20)
ax.set_ylabel('Readmission Rate',fontsize=20)
ax.set_xticklabels(labels)
ax.fill_between([xmin,xmax], 1,ymax, facecolor='orange', alpha = .15, interpolate=True)
ax.fill_between([xmin,xmax], ymin, 1, facecolor='blue', alpha = .15, interpolate=True)
fig, ax = plt.subplots(figsize=(10,10))
sns.regplot(large_df['Number of Discharges'], large_df['Excess Readmission Ratio'], scatter_kws={'alpha':0.15})
sns.regplot(small_df['Number of Discharges'], small_df['Excess Readmission Ratio'], scatter_kws={'alpha':0.15})
ax.set_xlabel('Hospital Size',fontsize=20)
ax.set_ylabel('Readmission Rate',fontsize=20)
# The box plot makes it easier to see the threshold for the ratio of readmissions (red line at 1), and you can visualize the spead of the data.
#
# The color coded scatter plot shows the distinction in hospital size for the raw data.
| sliderule_dsi_inferential_statistics_exercise_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/prashanth-acsq/Colab-Notebooks/blob/main/fgvc9/FGVC9_(R50%2C_Full%2C_NA256%2C_OCLR).ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ZfxSNLQluo8d"
# ### **Bring Data Into Notebook**
# + id="O0VYp5vnuP-w" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 180} outputId="c7017d97-de34-45c7-f239-3c2d94d84e53"
from time import time
start_time = time()
from IPython.display import clear_output
from google.colab import files
files.upload()
# !pip install -q kaggle
# !mkdir -p ~/.kaggle
# !cp kaggle.json ~/.kaggle/
# !ls ~/.kaggle
# !chmod 600 /root/.kaggle/kaggle.json
# !mkdir ~/.data
# !kaggle kernels output prashanthacsq/fgvc9-images-256 -p /.data
clear_output()
print(f"Time Taken to download data : {(time()-start_time)/60:.2f} minutes")
# + colab={"base_uri": "https://localhost:8080/"} id="oREGm3PFd7MZ" outputId="a9c03b6b-fb76-4641-8168-476474afbbd7"
# !ls /.data
# + [markdown] id="TOOFDqrjurKt"
# ### **Library Imports**
# + id="27P89OuzutMx"
import os
import re
import torch
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from torch import nn, optim
from torch.utils.data import Dataset
from torch.utils.data import DataLoader as DL
from torchvision import models, transforms
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import StratifiedKFold
# + [markdown] id="ayDKd9nGu_WM"
# ### **Utilities and Constants**
# + id="qCyu4W798eiY"
SEED = 42
DEVICE = torch.device("cuda" if torch.cuda.is_available() else "cpu")
TRANSFORM = transforms.Compose([transforms.ToTensor(),
transforms.Normalize([0.37250, 0.37591, 0.37537],
[0.20316, 0.21171, 0.21160]),
])
SAVE_PATH = "/content/saves"
if not os.path.exists(SAVE_PATH):
os.makedirs(SAVE_PATH)
# + id="949la0RZeo_Z"
def breaker(num: int=50, char: str="*") -> None:
print("\n" + num*char + "\n")
def show_graphs(L: list, A: list) -> None:
TL, VL, TA, VA = [], [], [], []
for i in range(len(L)):
TL.append(L[i]["train"])
VL.append(L[i]["valid"])
TA.append(A[i]["train"])
VA.append(A[i]["valid"])
x_Axis = np.arange(1, len(TL) + 1)
plt.figure(figsize=(8, 6))
plt.subplot(1, 2, 1)
plt.plot(x_Axis, TL, "r", label="Train")
plt.plot(x_Axis, VL, "b", label="Valid")
plt.legend()
plt.grid()
plt.title("Loss Graph")
plt.subplot(1, 2, 2)
plt.plot(x_Axis, TA, "r", label="Train")
plt.plot(x_Axis, VA, "b", label="Valid")
plt.legend()
plt.grid()
plt.title("Accuracy Graph")
plt.show()
# + [markdown] id="wUr6F2MRvC16"
# ### **Configuration**
# + id="A-R7w0tcu-1z"
class CFG(object):
def __init__(self,
seed: int = 42,
n_splits: int = 5,
batch_size: int = 16,
epochs: int = 25,
early_stopping: int = 5,
lr: float = 1e-4,
wd: float = 0.0,
max_lr: float = 1e-3,
pct_start: float = 0.2,
steps_per_epoch: int = 100,
div_factor: int = 1e3,
final_div_factor: float = 1e3,
):
self.seed = seed
self.n_splits = n_splits
self.batch_size = batch_size
self.epochs = epochs
self.early_stopping = early_stopping
self.lr = lr
self.wd = wd
self.max_lr = max_lr
self.pct_start = pct_start
self.steps_per_epoch = steps_per_epoch
self.div_factor = div_factor
self.final_div_factor = final_div_factor
cfg = CFG(seed=SEED)
# + [markdown] id="-7ods3fduuL1"
# ### **Dataset Template**
# + id="YJsiPr6MuwN1"
class DS(Dataset):
def __init__(self, images: np.ndarray, labels: np.ndarray = None, transform=None):
self.images = images
self.labels = labels
self.transform = transform
def __len__(self):
return self.images.shape[0]
def __getitem__(self, idx):
if self.labels is not None:
return self.transform(self.images[idx]), torch.LongTensor(self.labels[idx])
else:
return self.transform(self.images[idx])
# + [markdown] id="G0qqqUyNuxRh"
# ### **Model**
# + id="uQSUzCF3uwxD"
class MyResnet(nn.Module):
def __init__(self):
super(MyResnet, self).__init__()
self.model = models.resnet50(pretrained=True, progress=True)
self.model.fc = nn.Linear(in_features=self.model.fc.in_features, out_features=100)
def forward(self, x):
return nn.LogSoftmax(dim=1)(self.model(x))
# + [markdown] id="dkPYM6vJuzjT"
# ### **Fit and Predict Helper**
# + id="7UQ8SB6BuzPZ"
def fit(model=None,
optimizer=None,
scheduler=None,
epochs=None,
early_stopping_patience=None,
dataloaders=None,
fold=None,
verbose=False) -> tuple:
def get_accuracy(y_pred, y_true):
y_pred = torch.argmax(y_pred, dim=1)
return torch.count_nonzero(y_pred == y_true).item() / len(y_pred)
if verbose:
breaker()
if fold: print(f"Training Fold {fold}...")
else: print("Training ...")
breaker()
bestLoss, bestAccs = {"train" : np.inf, "valid" : np.inf}, {"train" : 0.0, "valid" : 0.0}
Losses, Accuracies, LRs = [], [], []
if fold: name = f"state_fold_{fold}.pt"
else: name = "state.pt"
start_time = time()
for e in range(epochs):
e_st = time()
epochLoss, epochAccs = {"train" : 0.0, "valid" : 0.0}, {"train" : 0.0, "valid" : 0.0}
for phase in ["train", "valid"]:
if phase == "train":
model.train()
else:
model.eval()
lossPerPass, accsPerPass = [], []
for X, y in dataloaders[phase]:
X, y = X.to(DEVICE), y.to(DEVICE).view(-1)
optimizer.zero_grad()
with torch.set_grad_enabled(phase == "train"):
output = model(X)
loss = torch.nn.NLLLoss()(output, y)
if phase == "train":
loss.backward()
optimizer.step()
if scheduler: scheduler.step()
lossPerPass.append(loss.item())
accsPerPass.append(get_accuracy(output, y))
epochLoss[phase] = np.mean(np.array(lossPerPass))
epochAccs[phase] = np.mean(np.array(accsPerPass))
LRs.append(scheduler.get_last_lr())
Losses.append(epochLoss)
Accuracies.append(epochAccs)
if early_stopping_patience:
if epochLoss["valid"] < bestLoss["valid"]:
bestLoss = epochLoss
BLE = e + 1
torch.save({"model_state_dict" : model.state_dict(),
"optim_state_dict" : optimizer.state_dict(),
"scheduler_state_dict" : scheduler.state_dict()},
os.path.join(SAVE_PATH, name))
early_stopping_step = 0
else:
early_stopping_step += 1
if early_stopping_step > early_stopping_patience:
print("\nEarly Stopping at Epoch {}".format(e + 1))
break
if epochLoss["valid"] < bestLoss["valid"]:
bestLoss = epochLoss
BLE = e + 1
torch.save({"model_state_dict" : model.state_dict(),
"optim_state_dict" : optimizer.state_dict(),
"scheduler_state_dict" : scheduler.state_dict()},
os.path.join(SAVE_PATH, name))
if epochAccs["valid"] > bestAccs["valid"]:
bestAccs = epochAccs
BAE = e + 1
if verbose:
print("Epoch: {} | Train Loss: {:.5f} | Valid Loss: {:.5f} |\
Train Accs: {:.5f} | Valid Accs: {:.5f} | Time: {:.2f} seconds".format(e+1,
epochLoss["train"], epochLoss["valid"],
epochAccs["train"], epochAccs["valid"],
time()-e_st))
if verbose:
breaker()
print(f"Best Validation Loss at Epoch {BLE}")
breaker()
print(f"Best Validation Accs at Epoch {BAE}")
breaker()
print("Time Taken [{} Epochs] : {:.2f} minutes".format(len(Losses), (time()-start_time)/60))
return Losses, Accuracies, LRs, BLE, BAE, name
# def predict(model=None, dataloader=None, path=None) -> np.ndarray:
# model.load_state_dict(torch.load(path, map_location=DEVICE)["model_state_dict"])
# model.to(DEVICE)
# model.eval()
# y_pred = torch.zeros(1, 1).to(DEVICE)
# for X in dataloader:
# X = X.to(DEVICE)
# with torch.no_grad():
# output = torch.argmax(torch.exp(model(X)), dim=1)
# y_pred = torch.cat((y_pred, output.view(-1, 1)), dim=0)
# return y_pred[1:].detach().cpu().numpy()
# + [markdown] id="ETCO3Q4Qu7TG"
# ### **Train**
# + id="cCc0FBiqdQw9"
images = np.load("/.data/images_256.npy")
labels = np.load("/.data/labels.npy")
# + id="HE-f1Djku8lc" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="ade2b685-697e-4d30-eb10-1e20dc11acc4"
for tr_idx, va_idx in StratifiedKFold(n_splits=cfg.n_splits, random_state=SEED, shuffle=True).split(images, labels):
break
tr_images, va_images, tr_labels, va_labels = images[tr_idx], images[va_idx], labels[tr_idx], labels[va_idx]
tr_data_setup = DS(tr_images, tr_labels.reshape(-1, 1), TRANSFORM)
va_data_setup = DS(va_images, va_labels.reshape(-1, 1), TRANSFORM)
dataloaders = {
"train" : DL(tr_data_setup, batch_size=cfg.batch_size, shuffle=True, generator=torch.manual_seed(cfg.seed)),
"valid" : DL(va_data_setup, batch_size=cfg.batch_size, shuffle=False),
}
cfg = CFG(epochs=50, steps_per_epoch=len(dataloaders["train"]))
torch.manual_seed(SEED)
model = MyResnet().to(DEVICE)
optimizer = optim.Adam([p for p in model.parameters() if p.requires_grad], lr=cfg.lr, weight_decay=cfg.wd)
scheduler = optim.lr_scheduler.OneCycleLR(optimizer=optimizer,
max_lr=cfg.max_lr,
epochs=cfg.epochs,
steps_per_epoch=cfg.steps_per_epoch,
pct_start=cfg.pct_start,
div_factor=cfg.div_factor,
final_div_factor=cfg.final_div_factor)
L, A, LRs, BLE, BAE, name = fit(model=model,
optimizer=optimizer,
scheduler=scheduler,
epochs=cfg.epochs,
early_stopping_patience=cfg.early_stopping,
dataloaders=dataloaders,
verbose=True)
breaker()
show_graphs(L, A)
# + id="a4vuWYiXDsiT"
# from google.colab import drive
# drive.mount("/content/gdrive")
# !cp "/content/saves/state.pt" "/content/gdrive/My Drive/Models/fgvc9/resnet50-state.pt"
| fgvc9/FGVC9_(R50,_Full,_NA256,_OCLR).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chatting Slang Converter
import re
import pandas as pd
import numpy as np
# + active=""
# N ={'2night': 'tonight',
# '2nite': 'tonight',
# '4mer': 'former',
# '7k': 'sick:-d laugher',
# }
# N
# -
a = {'a-town': 'atlanta,ga',
'a.o.e.': 'area of effect',
'a/n': 'authors note',
'a3': 'anytime, anywhere, anyplace',
'a7x': 'avenged sevenfold',
'aa': 'allahu akbar',
'aaa': 'anywhere, any place, any time',
'aaftddup': 'as a friend till death do us part',
'aanfctw': 'a**h**es are not f**king checking these words',
'aapl': 'apple computer inc.',
'aatk': 'asleep at the keyboard',
'ab': 'about',
'abb': 'anything but bush',
'abc': 'already been chewed',
'abot': 'about',
'acgad': "absolutely couldn't give a damn",
'ad': 'advertisement',
'adad': 'another day another doller',
'adidas': 'all day i dream about sex',
'adiem': 'all day i eat meat',
'afaik': 'as far as i know',
'afap': 'as far as possible',
'afk': 'away from keyboard',
'afkst': 'away from keyboard stop talking',
'afm': 'away from mouse',
'afol': 'adult fan of lego',
'afp': 'away from phone',
'afret': 'air force retired',
'ahic': 'ah i see',
'ahz': 'ahz',
'aiiegaf': 'as if i even give a f**k',
'aijcomkb': 'aw, i just came on my keyboard',
"ain't": 'is not',
'alaittrwl': 'at least add it to the rejected words list.',
'alwas': 'always',
'amazin': 'amazing',
'anof': 'enough',
'ansx': 'anal sex',
'ao': 'auto op irc',
'aom': 'age of mythology',
'aotm': 'artist of the month',
'aoyp': 'angels on your pillow',
'ap': 'already posted',
'apl': 'approved paid leave',
"aren't": 'are not',
'arent': 'are not',
'asap': 'as soon as possible',
'asl': 'age, sex, location',
'asn': 'age, sex, name',
'aswell': 'as well',
'aswgaff': 'ask someone who gives a flying f**k',
'atay': 'always thinking about you',
'atk': 'at the keyboard',
'atlw': 'against the law',
'atm': 'at the moment',
'atom': 'always think of me',
'att': 'all the time',
'attoml': 'at the top of my lungs',
'atuw': 'any time u want',
'aufkmls': 'are you f**king kidding me like seriously',
'aul': 'as you like',
'av': 'avatar',
'avi': 'avatar',
'avon': 'marijuana',
'aw': 'ass-wipe',
'awd': 'all will die',
'awlol': 'a whole lot of love',
'ax': 'ask',
'aycbqo': 'abs you can bounce quaters off of',
'aycbqoo': 'abs you can bounce quarters off of',
'aygs': 'are you going somewhere',
'aypudh': 'answer your phone you d**k head',
'ayor': 'at your own risk',
'aysos': 'are you stupid or something',
'aznpryde': 'asian pride'}
b = {'b-lo': 'buffalo',
'b/a': 'birth announcment',
'b/m': 'bar mitzvah',
'b4': 'before',
'b4n': 'bye for now',
'ba': 'bad ass',
'babesz': 'babe',
'babz': 'baby',
'bac': 'berries and cream!',
'bacap': 'be as cool as possible',
'bak': 'back at keyboard',
'bass': 'b***h ass',
'bat': 'big ass titties',
'bay': 'back at you',
'bb4l': 'bad boy for life',
'bbb': 'bye bye b***h',
'bbbjcim': 'bare back blow job c** in mouth',
'bbbjtc': 'bare back blow job to completition',
'bbc': 'big black c**k',
'bbiwc': 'be back in a while crocodile',
'bbl': 'be back later',
'bbmfic': 'big black mouther f**ker in charge',
'bbs': 'be back soon',
'bc': 'because',
'bcou': 'because of you',
'bd': 'birthday',
'bday': 'birthday',
'beeotch': 'b***h',
'beg': 'big evil grin',
'bep': 'black eyed peas',
'bfbfl': 'best f**k buddie for life',
'bfdb': 'big fat d****e bag',
'bfff': 'best friend for forever',
'bfffl': 'best friends for f**king life',
'bfflaanmw': 'best friends for life and always no matter what',
'bfflabnmw': 'best friends for life and beyond no matter what',
'bfflead': 'best friend for life, even after death',
'bffttandc': 'best friend for today tommorow and next day continusely',
'bffttwgb': 'best friends forever til the world goes bang',
'bffutsolfdwin': 'best friends forever until the statue of liberty falls down which is never',
'bffwgsaly': 'best freinds forever who go shopping a lot yo',
'bffwtalotfagsatt': 'best friends forever who talk a lot on the phone and go shopping all the time',
'bfhm': 'big fat hairy man',
'bfl': 'b***h for life',
'bfmn': 'butt f**k me now',
'bfn': 'bye for now',
'bfo': 'b***h f**k off',
'bfplz': 'be friend please',
'bftimsb': 'best friends till i meet someone better',
'bfwb': 'best female with benefits',
'bhimbgo': 'bloody hell i must be getting old',
'bhm': 'black hat man (from xkcd.com)',
'bhmf': 'blondes have more fun',
'bias': 'back in a sec',
'bidw': 'but i dont wanna!',
'biffles': 'best friends',
'bily': 'but i love you',
'bio': 'bring it on',
'bjb': 'bond james bond',
'bjih': "best joke i've heard",
'bk': 'back',
'blfe': 'b***h lovers forever',
'bllin': 'ballin',
'blos': 'brother looming over shoulder',
'blt': 'bacon lettuce tomato',
'blw': 'below',
'bm43n3': 'best mates for ever and ever',
'bmg': 'be my girlfreind',
'bmm': 'blows my mind',
'bmth': 'bring me the horizon',
'bmttfe': 'best mates til the f**king end',
'bmv': 'be my valentine',
'bmw': 'be my wife',
'bmwb': 'best male with benefits',
'bnc': 'bouncer irc',
'bner': 'beaner',
'bnkch': 'bank character',
'bob': 'back off b***h',
'boc': 'but of course',
'bok': 'back on keyboard',
'boma': 'best of mates always',
'boobs': 'breasts',
'booooodie': 'buddy',
'booziest': 'busiest',
'bot': 'robot',
'botd': 'bring out the dead',
'bout': 'about',
'bp': 'beer pong',
'bplz': 'b***h please',
'brb': 'be right back',
'brbigtgp': 'be right back, i got to go pee',
'brbinte': 'be right back i need to eat',
'brbmpawoms': 'be right back, my parents are watching over my shoulder.',
'brfhrn': 'be real f**king honest right now',
'brittneyspears': 'cold beers',
'brt': 'be right there',
'bs': 'bull shit',
'bsilf': 'bens sister id like to f**k',
'bsofd': 'blue screen of f**king death',
'bstl': 'balls stuck to leg',
'bsw': 'b***h s**t w***e',
'btdtgttsawio': 'been there, done that, got the t-shirt and wore it out',
'btdtgttswio': 'been there done that got the t-shirt wore it out',
'btee': 'booty',
'btfwi': 'by the f**king way idiot',
'btmc': 'b***h took my chips',
'bttpmu': 'be there to pick me up',
'btw': 'by the way',
'btwitiailwy': 'by the way i think i am in love with you',
'btwitiainlwy': 'by the way i think i am in love with you',
'bu': 'butt ugly',
'bubbiii': 'bye',
'buff': 'good looking',
'butters': 'ugly',
'butterz': 'ugly',
'butts3xs': 'butt sex',
'bw': 'buttwipe',
'bwb': 'b***h with boobs',
'bwboin': 'beware boss is near',
'bwlotf': 'bursting with laughing on the floor',
'bygts': 'baby you got to stop',
'byitf': 'bust you in the face',
'bylasc': 'beat you like a step child'}
c = {'c': 'see',
'c 2 c': 'cam to cam (webcams)',
'c&c': 'command and conquer',
"c'mon": 'come on',
'c-p': 'sleepy',
'c.y.a': 'cover your ass',
'c/b': 'comment back',
'c/t': "can't talk",
'c14n': 'canonicalization',
'c2': 'come to',
'c2c': 'care to chat?',
'c2tc': 'cut to the chase',
'c4ashg': 'care for a shag',
'c4y': 'cool for you',
'c@': 'cat',
'cam': 'camera',
"can't": 'cannot',
"can't've": 'cannot have',
'cancer stick': 'cigarette',
'cant': 'cannot',
'catwot': 'complete and total waste of time',
'cawk': 'c**k',
'cayc': 'call at your convenience',
'cb': 'come back',
'cba': "can't be arsed",
'cbb': "can't be bothered",
'cbf': 'cant be f**ked',
'cbfa': "can't be f**king arsed",
'cbfed': "can't be f**ked",
'cbi': "can't believe it",
'ccl': "couldn't care less",
'ccna': 'cisco certified network associate',
'cd9': 'code 9 (other people nearby)',
'celly': 'cell phone',
'cex': 'sex',
'cexy': 'sexy',
'cfas': 'care for a secret?',
'cfid': 'check for identification',
'cfm': 'come f**k me',
'cg': 'congratulations',
'cgad': "couldn't give a damn",
'cgaf': "couldn't give a f**k",
'cgf': 'cute guy friend',
'ch@': 'chat',
'champs': 'champions',
'char': 'character',
'cheezburger': 'cheeseburger',
'chik': 'chick',
'chilax': 'chill and relax in one word',
'chillax': 'chill and relax',
'chilliin': 'chilling in',
'chillin': 'relaxing',
'chk': 'check',
'chohw': 'come hell or high water',
'chr': 'character',
'chronic': 'marijuana',
'chswm': 'come have sex with me',
'chswmrn': 'come have sex with me right now',
'chu': 'you',
'chut': 'p***y',
'cid': 'consider it done',
'cig': 'cigarette',
'cigs': 'cigarettes',
'cihswu': 'can i have sex with you',
'cihyn': 'can i have your number',
'cilf': "child i'd like to f**k",
'cing': 'seeing',
'cis': 'computer information science',
'ciwwaf': 'cute is what we aim for',
'cless': 'clanless',
'clm': 'cool like me',
'clt': 'cool like that',
'cluebie': 'clueless newbie',
'cm': 'call me',
'cma': 'cover my ass',
'cmao': 'crying my ass off',
'cmar': 'cry me a river',
'cmb': 'comment me back',
'cmbo': 'combo',
'cmcp': 'call my cell phone',
'cmeo': 'crying my eyes out',
'cmh': 'call my house',
'cmiiw': "correct me if i'm wrong",
'cmitm': 'call me in the morning',
'cml': 'call me later',
'cml8r': 'call me later',
'cmliuw2': 'call me later if you want to',
'cmn': 'call me now',
'cmomc': 'call me on my cell',
'cmon': 'come on',
'cmplcdd': 'complicated',
'cmplte': 'complete',
'cmptr': 'computer',
'cms': 'content management system',
'cmt': 'comment',
'cmw': 'cutting my wrists',
'cn': 'can',
'cnc': 'command and conquer',
'cnt': "can't",
'cob': 'close of business',
'cod': 'call of duty',
'cod4': 'call of duty 4',
'cod5': 'call of duty 5',
'codbo': 'call of duty: black ops',
'codbo2': 'call of duty: black ops ii',
'code 29': 'moderator is watching',
'code 8': 'parents are watching',
'code 9': 'parents are watching',
'code9': 'other people near by',
'cof': 'crying on the floor',
'coiwta': 'come on i wont tell anyone',
'col': 'crying out loud',
"comin'": 'coming',
'comnt': 'comment',
'comp': 'computer',
'compy': 'computer',
'congrats': 'congratulations',
'contrib': 'contribution',
'contribs': 'contributions',
'convo': 'conversation',
'coo': 'cool',
'cood': 'could',
'copyvio': 'copyright violation',
'cos': 'because',
'cotf': 'crying on the floor',
'cotm': 'check out this myspace',
"could've": 'could have',
"couldn't": 'could not',
"couldn't've": 'could not have',
'cowboy choker': 'cigarette',
'coz': 'because',
'cp': 'child porn',
'cpl': 'cyber athlete professional league',
'cpm': 'cost per 1000 impressions',
'cptn': 'captain',
'cpu': 'computer',
'cpy': 'copy',
'cr': "can't remember",
'cr8': 'crate',
'crakalakin': 'happening',
'cray-cray': 'crazy',
'crazn': 'crazy asian',
'cre8or': 'creator',
'crm': 'customer relationship management',
'crp': 'crap',
'crs': "can't remember s**t",
'crunk': 'combination of crazy and drunk',
'crzy': 'crazy',
'cs': 'counter-strike',
'cs:s': 'counter-strike: source',
'csi': 'crime scene investigation',
'cskr': 'c**k sucker',
'csl': "can't stop laughing",
'css': 'cascading style sheets',
'ct': "can't talk",
'ctc': 'call the cell',
'ctf': 'capture the flag',
'ctfd': 'calm the f**k down',
'ctfo': 'chill the f**k out',
'ctfu': 'cracking the f**k up',
'ctm': 'chuckle to myself',
'ctn': "can't talk now",
'ctnbos': "can't talk now boss over shoulder",
'ctncl': "can't talk now call later",
'ctpc': 'cant talk parent(s) coming',
'ctpos': "can't talk parent over sholder",
'ctrl': 'control',
'ctrn': "can't talk right now",
'cts': 'change the subject',
'ctt': 'change the topic',
'cu': 'goodbye',
'cu2': 'see you too',
'cu2nit': 'see you tonight',
'cu46': 'see you for sex',
'cubi': 'can you believe it',
'cud': 'could',
'cuic': 'see you in class',
'cul': 'see you later',
'cul83r': 'see you later',
'cul8er': 'see you later',
'cul8r': 'see you later',
'cul8tr': 'see you later',
'culd': 'could',
'cunt': 'vagina',
'cuom': 'see you on monday',
'cuple': 'couple',
'curn': 'calling you right now',
'cut3': 'cute',
'cuwul': 'catch up with you later',
'cuz': 'because',
'cuzz': 'because',
'cvq': 'chucking very quietly',
'cw2cu': 'can`t wait to see you',
'cwd': 'comment when done',
'cwm': 'come with me',
'cwmaos': 'coffee with milk and one sugar',
'cwot': 'complete waste of time',
'cwtgypo': "can't wait to get your panties off",
'cwyl': 'chat with ya later',
'cy2': 'see you too',
'cya': 'goodbye',
'cyal': 'see you later',
'cyal8r': 'see you later',
'cyas': 'see you soon',
'cyb': 'cyber',
'cybl': 'call you back later',
'cybr': 'cyber',
'cybseckz': 'cyber sex',
'cye': 'close your eyes',
'cyff': 'change your font, f**ker',
'cyl': 'see you later',
'cyl,a': 'see ya later, alligator',
'cyl8': 'see you later',
'cyl8er': 'see you later',
'cylbd': 'catch ya later baby doll',
'cylor': 'check your local orhtodox rabbi',
'cym': 'check your mail',
'cyntott': 'see you next time on tech today',
'cyt': 'see you tomorrow',
'cyu': 'see you',
'c|n>k': 'coffee through nose into keyboard'}
d = {'d&c': 'divide and conquer',
'd&df': 'drug & disease free',
'd.t.f': 'down to f**k',
'd.w': "don't worry",
'd/c': 'disconnected',
'd/l': 'download',
'd/m': "doesn't matter",
'd/w': "don't worry",
'd00d': 'dude',
'd1ck': 'd**k',
'd2': '<NAME>',
'd2m': 'dead to me',
'd2t': 'drink to that',
'd8': 'date',
'da': 'the',
'da2': 'dragon age 2',
'dads': 'father is',
'dadt': "don't ask. don't tell.",
'dafs': 'do a f**king search',
'dafuq': 'what the f**k',
'dah': 'dumb as hell',
'daii': 'day',
'damhik': "don't ask me how i know",
'damhikijk': "don't ask me how i know - i just know",
'damhikt': "don't ask me how i know this",
'dass': 'dumb ass',
'dat': 'that',
'dats': "that's",
'dawg': 'friend',
'dayum': 'damn',
'dayumm': 'damn',
'daze': 'days',
'db': 'database',
'db4l': 'drinking buddy for life',
'dbab': "don't be a b***h",
'dbafwtt': "don't be a fool wrap the tool",
'dbag': 'd****ebag',
'dbeyr': "don't believe everything you read",
'dbg': "don't be gay",
'dbh': "don't be hating",
'dbi': "don't beg it",
'dbm': "don't bother me",
'dbz': 'dragonball z',
'dc': "don't care",
"dc'd": 'disconnected',
'dctnry': 'dictionary',
'dcw': 'doing class work',
'dd': "don't die",
'ddf': 'drug and disease free',
'ddg': 'drop dead gorgeous',
'ddl': 'direct download',
'ddos': 'distributed denial of service',
'ddr': 'dance dance revolution',
'ddt': "don't do that",
'ded': 'dead',
'deets': 'details',
'deez': 'these',
'def': 'definitely',
'defs': 'definetly',
'degmt': "don't even give me that",
'dem': 'them',
'demo': 'demonstration',
'dept': 'department',
'der': 'there',
'dernoe': "i don't know",
'detai': "don't even think about it",
'dewd': 'dude',
'dey': 'they',
'df': 'dumb f**k',
'dfc': "don't f**king care",
'dfo': 'dumb f**king operator',
'dftba': "don't forget to be awesome",
'dftc': 'down for the count',
'dfu': "don't f**k up",
'dfw': 'down for whatever',
'dfw/m': "don't f**k with me",
'dfwm': "don't f**k with me",
'dfwmt': "don't f**king waste my time",
'dg': "don't go",
'dga': "don't go anywhere",
'dgac': "don't give a crap",
'dgaf': "don't give a f**k",
'dgara': "don't give a rats ass",
'dgas': "don't give a s**t",
'dgms': "don't get me started",
'dgoai': "don't go on about it",
'dgt': "don't go there",
'dgu': "don't give up",
'dgypiab': "don't get your panties in a bunch",
'dh': 'dickhead',
'dhac': "don't have a clue",
'dhcp': 'dynamic host configuration protocol',
'dhly': 'does he like you',
'dhv': 'demonstration of higher value',
'diacf': 'die in a car fire',
'diaf': 'die in a fire',
'diah': 'die in a hole',
'dic': 'do i care',
'dick': 'penis',
"didn't": 'did not',
'didnt': 'did not',
'diez': 'dies',
'diff': 'difference',
'dih': 'd**k in hand',
'dikhed': 'dickhead',
'diku': 'do i know you',
'diky': 'do i know you',
'dil': 'daughter in law',
'dilf': "dad i'd like to f**k",
'dillic': 'do i look like i care',
'dillifc': 'do i look like i f**king care',
'dilligad': 'do i look like i give a damn',
'dilligaf': 'do i look like i give a f**k',
'dilligas': 'do i look like i give a s**t',
'din': "didn't",
"din't": "didn't",
'dirl': 'die in real life',
'dis': 'this',
'dit': 'details in thread',
'diy': 'do it yourself',
'dju': 'did you',
'dk': "don't know",
'dkdc': "don't know, don't care",
'dl': 'download',
'dlf': 'dropping like flies',
'dlibu': 'dont let it bother you',
'dln': "don't look now",
'dm': 'deathmatch',
'dmaf': 'do me a favor',
'dmba*': 'dumbass',
'dmi': "don't mention it",
'dmn': 'damn',
'dmu': "don't mess up",
'dmwm': "don't mess with me",
'dmy': "don't mess yourself",
'dn': "don't know",
'dnd': 'do not disturb',
'dndp': 'do not double post',
'dnimb': 'dancing naked in my bra',
'dnk': 'do not know',
'dno': "don't know",
'dnrta': 'did not read the article',
'dnrtfa': 'did not read the f**king article',
'dns': 'domain name system',
'dnt': "do not",
'dnw': 'do not want',
'doa': 'dead on arrival',
'dob': 'date of birth',
'dod': 'day of defeat',
"doesn't": 'does not',
'doesnt': 'does not',
'dogg': 'friend',
'doin': 'doing',
"doin'": 'doing',
'don': 'denial of normal',
"don't": 'do not',
'donbt': 'do not',
'doncha': "don't you",
'donno': "don't know",
'dont': "don't",
'dontcha': "don't you",
'dood': 'dude',
'doodz': 'dudes',
'dos': 'denial of service',
"dosen't": 'does not',
'dotc': 'dancing on the ceiling',
'doypov': 'depends on your point of view',
'dp': 'display picture',
'dpmo': "don't piss me off",
'dprsd': 'depressed',
'dqmot': "don't quote me on this",
'dqydj': "don't quit your day job",
'dr00d': 'druid',
'drc': "don't really care",
'drm': 'dream',
'drood': 'druid',
'dsided': 'decided',
'dsl' : 'digital satellite link',
'dsu': "don't screw up",
'dt': 'double team',
'dta': "don't trust anyone",
'dtb': "don't text back",
'dtf': 'down to f**k',
'dtg': 'days to go',
'dth': 'down to hang',
'dtl': 'damn the luck',
'dtp': "don't type please",
'dtrt': 'do the right thing',
'dts': "don't think so",
'dttm': "don't talk to me",
'dttml': "don't talk to me loser",
'dttpou': "don't tell the police on us",
'dttriaa': "don't tell the riaa",
'du2h': 'damn you to hell',
'ducy': 'do you see why',
'dugi': 'do you get it?',
'dugt': 'did you get that?',
'dui': 'driving under the influence',
'duk': 'did you know',
'dulm': 'do you like me',
'dum': 'dumb',
'dun': "don't",
'dunna': "i don't know",
'dunno': "i don't know",
'duno': "don't know",
'dupe': 'duplicate',
'dutma': "don't you text me again",
'dvda': 'double vaginal, double anal',
'dw': "don't worry",
'dwai': "don't worry about it",
'dwb': 'driving while black',
'dwbh': "don't worry, be happy",
'dwbi': "don't worry about it.",
'dwi': 'deal with it',
'dwioyot': 'deal with it on your own time',
'dwmt': "don't waste my time",
'dwn': 'down',
'dwt': "don't wanna talk",
'dwud': 'do what you do',
'dwy': "don't wet yourself",
'dy2h': 'damn you to hell',
'dya': 'do you',
'dyac': 'damn you auto correct',
'dycotfc': 'do you cyber on the first chat',
'dyec': "don't you ever care",
'dygtp': 'did you get the picture',
'dyk': 'did you know',
'dylh': 'do you like him',
'dylm': 'do you love me',
'dylos': 'do you like oral sex',
'dym': 'do you mind',
'dymm': 'do you miss me',
'dynk': 'do you not know',
'dynm': 'do you know me',
'dyt': "don't you think",
'dyth': 'damn you to hell',
'dyw': "don't you worry",
'dyw2gwm': 'do you want to go with me',
'dywtmusw': 'do you want to meet up some where'}
e = { 'e-ok': 'electronically ok',
'e.g.': 'example',
'e4u2s': 'easy for you to say',
'eabod': 'eat a bag of dicks',
'ead': 'eat a d**k',
'ebitda': 'earnings before interest, taxes, depreciation and amortization',
'ecf': 'error carried forward',
'edumacation': 'education',
'eedyat': 'idiot',
'eejit': 'idiot',
'ef': 'f**k',
'ef-ing': 'f**king',
'efct': 'effect',
'effed': 'f**ked',
'efffl': 'extra friendly friends for life',
'effin': 'f**king',
'effing': 'f**king',
'eg': 'evil grin',
'ehlp': 'help',
'eil': 'explode into laughter',
'el!t': 'elite',
'eleo': 'extremely low earth orbit',
'ello': 'hello',
'elo': 'hello',
'em': 'them',
'emm': 'email me',
'emo': 'emotional',
'emp': 'eat my p***y',
'enat': 'every now and then',
'enit': "isn't it",
'enof': 'enough',
'enuf': 'enough',
'enuff': 'enough',
'eob': 'end of business',
'eoc': 'end of conversation',
'eod': 'end of day',
'eof': 'end of file',
'eom': 'end of message',
'eos': 'end of story',
'eot': 'end of transmission',
'eotw': 'end of the world',
'epa': 'emergency parent alert',
'eq': 'everquest',
'eq2': 'everquest2',
'ere': 'here',
'errythang': 'everything',
'errythin': 'everything',
'esad': 'eat s**t and die',
'esadyffb': 'eat s**t and die you fat f**king b*****d',
'esbm': 'everyone sucks but me',
'esc': 'escape',
'esl': 'eat s**t loser',
'eta': 'estimated time of arrival',
'etla': 'extended three letter acronym',
'etmda': 'explain it to my dumb ass',
'etp': 'eager to please',
'eula': 'end user license agreement',
'ev1': 'everyone',
'eva': 'ever',
'evaa': 'ever',
'evar': 'ever',
'evercrack': 'everquest',
'every1': 'everyone',
'evn': 'even',
'evr': 'ever',
'evry': 'every',
'evry1': 'every one',
'evrytin': 'everything',
'ex-bf': 'ex-boy friend',
'ex-gf': 'ex-girl friend',
'exp': 'experience',
'ey': 'hey',
'eyez': 'eyes',
'ez': 'easy',
'ezi': 'easy'}
f = {'f u': 'f**k you',
'f#cking': 'f**king',
'f&e': 'forever and ever',
"f'n": 'f**king',
'f-ing': 'f**king',
'f.b.': 'facebook',
'f.m.l.': 'f**k my life',
'f.u.': 'f**k you',
'f/o': 'f**k off',
'f00k': 'f**k.',
'f2f': 'face to face',
'f2m': 'female to male',
'f2p': 'free to play',
'f2t': 'free to talk',
'f4c3': 'face',
'f4eaa': 'friends forever and always',
'f4f': 'female for female',
'f4m': 'female for male',
'f8': 'fate',
'f9': 'fine',
'f@': 'fat',
'fa-q': 'f**k you',
'faa': 'forever and always',
'fab': 'fabulous',
'faggit': 'faggot',
'fah': 'funny as hell',
'faic': 'for all i care',
'fam': 'family',
'fankle': 'area between foot and ankle',
'fao': 'for attention of',
'fap': 'masturbate',
'fapping': 'masterbating',
'faq': 'frequently asked question',
'farg': 'f**k',
'fashizzle': 'for sure',
'fav': 'favorite',
'fave': 'favorite',
'fawk': 'f**k',
'fb': 'facebook',
'fbgm': 'f**k b***hes get money',
'fbimcl': 'fall back in my chair laughing',
'fbk': 'facebook',
'fbtw': 'fine be that way',
'fc': 'fruit cake',
'fcbk': 'facebook',
'fcfs': 'first come first served',
'fck': 'f**k',
'fckd': 'f**ked',
'fckin': 'f**king',
'fcking': 'f**king',
'fckm3hdbayb': 'f**k me hard baby',
'fcku': 'f**k you',
'fcol': 'for crying out loud',
'fcuk': 'f**k',
'fe': 'fatal error',
'feat': 'featuring',
'feck': 'f**k',
'fer': 'for',
'ferr': 'for',
'ff': 'friendly fire',
'ffa': 'free for all',
'ffcl': 'falling from chair laughing',
'ffl': 'friend for life',
'ffr': 'for future reference',
'ffs': "for f**k's sake",
'fft': 'food for thought',
'ffxi': 'final fantasy 11',
'fg': 'f**king gay',
'fgi': 'f**king google it',
'fgs': "for god's sake",
'fgssu': 'for gods sake shut up',
'fgt': 'faggot',
'fhrihp': 'f**k her right in her p***y',
'fi': 'f**k it',
'fi9': 'fine',
'fibijar': "f**k it buddy, i'm just a reserve",
'fifo': 'first in, first out',
'fify': 'fixed it for you',
'figjam': "f**k i'm good, just ask me",
'figmo': 'f*ck it - got my orders',
'fiic': 'f**ked if i care',
'fiik': 'f**ked if i know',
"fillin'": 'filling',
'fimh': 'forever in my heart',
'finna': 'fitting to',
'fio': 'figure it out',
'fitb': 'fill in the blank',
'fiv': 'five',
'fk': 'f**k',
'fka': 'formerly known as',
'fkd': 'f**ked',
'fker': 'f**ker',
'fkin': 'f**king',
'fking': 'f**king',
'fkn': 'f**king',
'fku': 'f**k you',
'flamer': 'angry poster',
'flames': 'angry comments',
'flicks': 'pictures',
'floabt': 'for lack of a better term',
'fm': 'f**k me',
'fmah': 'f**k my ass hole',
'fmao': 'freezing my ass of',
'fmb': 'f**k me b***h',
'fmbb': 'f**k me baby',
'fmbo': 'f**k my brains out',
'fmfl': 'f**k my f**king life',
'fmflth': 'f**k my f**king life to hell',
'fmh': 'f**k me hard',
'fmhb': 'f**k me hard b***h',
'fmi': 'for my information',
'fmir': 'family member in room',
'fmita': 'f**k me in the ass',
'fml': 'f**k my life',
'fmltwia': 'f**k me like the w***e i am',
'fmn': 'f**k me now',
'fmnb': 'f**k me now b***h',
'fmnkml': 'f**k me now kiss me later',
'fmph': 'f**k my p***y hard',
'fmq': 'f**k me quick',
'fmr': 'f**k me runnig',
'fmsh': 'f**k me so hard',
'fmth': 'f**k me to hell',
'fmu': 'f**k me up',
'fmup': 'f**k me up',
'fmuta': 'f**k me up the ass',
'fmutp': 'f**k me up the p***y',
'fn': 'first name',
'fnar': 'for no apparent reason',
'fnci': 'fancy',
'fnny': 'funny',
'fnpr': 'for no particular reason',
'fny': 'funny',
'fo': 'f**k off',
'fo shizzle': 'for sure',
'fo sho': 'for sure',
'fo`': 'for',
'foa': 'f**k off a**h**e',
'foad': 'f**k off and die',
'foaf': 'friend of a friend',
'foah': 'f**k off a**h**e',
'fob': 'fresh off the boat',
'focl': 'falling off chair laughing.',
'fofl': 'fall on the floor laughing',
'foia': 'freedom of information act',
'fol': 'farting out loud',
'folo': 'follow',
'fomofo': 'f**k off mother f**ker',
'foms': 'fell off my seat',
'fone': 'phone',
'foo': 'fool',
'foobar': 'f**ked up beyond all recognition',
'foocl': 'falls out of chair laughing',
'fook': 'f**k',
'for sheeze': 'for sure',
'fos': 'full of s**t',
'foshizzle': 'for sure',
'fosho': 'for sure',
'foss': 'free, open source software',
'fotcl': 'fell off the chair laughing',
'fotm': 'flavour of the month',
'fouc': 'f**k off you c**t',
'fov': 'field of view',
'foyb': 'f**k off you b***h',
'fp': 'first post',
'fpmitap': 'federal pound me in the ass prison',
'fpos': 'f**king piece of s**t',
'fps': 'first person shooter',
'fr': 'for real',
'frag': 'kill',
'fragged': 'killed',
'fren': 'friend',
'frens': 'friends',
'frgt': 'forgot',
'fri.': 'friday',
'friggin': 'freaking',
'frk': 'freak',
'frm': 'from',
'frnd': 'friend',
'frnds': 'friends',
'fs': 'for sure',
'fse': 'funniest s**t ever',
'fsho': 'for sure',
'fsm': 'flying spaghetti monster',
'fsob': 'f**king son of a b***h',
'fsod': 'frosn screen of death',
'fsr': 'for some reason',
'fst': 'fast',
'ft': 'f**k that',
'ft2t': 'from time to time',
'fta': 'from the article',
'ftb': 'f**k that b***h',
'ftbfs': 'failed to build from source',
'ftf': 'face to face',
'ftfa': 'from the f**king article',
'ftfw': 'for the f**king win!',
'ftfy': 'fixed that for you',
'ftio': 'fun time is over',
'ftk': 'for the kill',
'ftl': 'for the lose',
'ftlog': 'for the love of god',
'ftlt': 'for the last time',
'ftmfw': 'for the mother f**king win',
'ftmp': 'for the most part',
'ftp': 'file transfer protocol',
'ftr': 'for the record',
'fts': 'f**k that s**t',
'fttp': 'for the time being',
'ftw': 'for the win!',
'fu': 'f**k you',
'fu2': 'f**k you too',
'fua': 'f**k you all',
'fuah': 'f**k you ass hole',
'fub': 'f**k you b***h',
'fubah': 'f**ked up beyond all hope',
'fubalm': 'f**ked up beyond all local maintenance',
'fubar': 'f**ked up beyond all recognition',
'fubb': 'f**ked up beyond belief',
'fubh': 'f**ked up beyond hope',
'fubohic': 'f**k you bend over here it comes',
'fubr': 'f**ked up beyond recognition',
'fucken': 'f**king',
'fucktard': 'f**king retard',
'fuctard': 'f**king retard',
'fud': 'fear, uncertainty and doubt',
'fudh': 'f**k you d**k head',
'fudie': 'f**k you and die',
'fugly': 'f**king ugly',
'fuh-q': 'f**k you',
'fuhget': 'forget',
'fuk': 'f**k',
'fukin': 'f**king',
'fukk': 'f**k',
'fukkin': 'f**king',
'fukn': 'f**king',
'fukr': 'f**ker',
'fulla': 'full of',
'fumfer': 'f**k you mother f**ker',
'funee': 'funny',
'funner': 'more fun',
'funy': 'funny',
'fuq': 'f**k you',
'fus': 'f**k yourself',
'fut': 'f**k you too',
'fuu': 'f**k you up',
'fux': 'f**k',
'fuxing': 'f**king',
'fuxor': 'f**ker',
'fuxored': 'f**ked',
'fvck': 'f**k',
'fwb': 'friends with benefits',
'fwd': 'forward',
'fwiw': "for what it's worth",
'fwm': 'fine with me',
'fwob': 'friends with occasional benefits',
'fwp': 'first world problems',
'fxe': 'foxy',
'fxp': 'file exchange protocol',
'fy': 'f**k you',
'fya': 'for your attention',
'fyad': 'f**k you and die',
'fyah': 'f**k you a**h**e',
'fyb': 'f**k you b***h',
'fyc': 'f**k your couch',
'fye': 'for your entertainment',
'fyeo': 'for your eyes only',
'fyf': 'f**k your face',
'fyfi': 'for your f**king information',
'fyi': 'for your information',
'fyk': 'for your knowledge',
'fyl': 'for your love',
'fym': 'f**k your mom',
'fyp': 'fixed your post',
'fyrb': 'f**k you right back',
'fytd': 'f**k you to death'}
g = { "g'nite": 'good night',
'g. o. a. t.': 'greatest of all time',
'g/f': 'girlfriend',
'g/g': 'got to go',
'g/o': 'get out',
'g0': 'go',
'g00g13': 'google',
'g1': 'good one',
'g2': 'go to',
'g2/-/': 'go to hell',
'g2bg': 'got to be going',
'g2bl8': 'going to be late',
'g2cu': 'glad to see you',
'g2e': 'got to eat',
'g2g': 'got to go',
'g2g2tb': 'got to go to the bathroom',
'g2g2w': 'got to go to work',
'g2g4aw': 'got to go for a while',
'g2gb': 'got to go bye',
'g2gb2wn': 'got to go back to work now',
'g2ge': 'got to go eat',
'g2gn': 'got to go now',
'g2gp': 'got to go pee',
'g2gpc': 'got 2 go parents coming',
'g2gpp': 'got to go pee pee',
'g2gs': 'got to go sorry',
'g2h': 'go to hell',
'g2hb': 'go to hell b***h',
'g2k': 'good to know',
'g2p': 'got to pee',
'g2t2s': 'got to talk to someone',
'g3y': 'gay',
'g4u': 'good for you',
'g4y': 'good for you',
'g8': 'gate',
'g9': 'good night',
'g@y': 'gay',
'ga': 'go ahead',
'gaalma': 'go away and leave me alone',
'gaf': 'good as f**k',
'gafi': 'get away from it',
'gafl': 'get a f**king life',
'gafm': 'get away from me',
'gagf': 'go and get f**ked',
'gagp': 'go and get pissed',
'gah': 'gay ass homo',
'gai': 'gay',
'gaj': 'get a job',
'gal': 'get a life',
'gamez': 'illegally obtained games',
'gangsta': 'gangster',
'gank': 'kill',
'gaoep': 'generally accepted office etiquette principles',
'gaw': 'grandparents are watching',
'gawd': 'god',
'gb': 'go back',
'gb2': 'go back to',
'gba': 'game boy advance',
'gbioua': 'go blow it out your ass',
'gbnf': 'gone but not forgotten',
'gbtw': 'go back to work',
'gbu': 'god bless you',
'gby': 'good bye',
'gcad': 'get cancer and die',
'gcf': 'google click fraud',
'gd': 'good',
'gd&r': 'grins, ducks, and runs',
'gd4u': 'good for you',
'gday': 'good day',
'gdby': 'good bye',
'gded': 'grounded',
'gdgd': 'good good',
'gdi': 'god damn it',
'gdiaf': 'go die in a fire',
'gdih': 'go die in hell',
'gdilf': "grandad i'd like to f**k",
'gdmfpos': 'god damn mother f**king piece of s**t',
'gdr': 'grinning, ducking, running',
'gemo': 'gay emo',
'getcha': 'get you',
'geto': 'ghetto',
'gewd': 'good',
'gey': 'gay',
'gf': 'girlfriend',
'gfad': 'go f**k a duck',
'gfadh': 'go f**k a dead horse',
'gfak': 'go fly a kite',
'gfam': 'go f**k a monkey',
'gfar2cu': 'go find a rock to crawl under',
'gfas': 'go f**k a sheep',
'gfd': 'god f**king damnit',
'gfe': 'girl friend experience',
'gfe2e': 'grinning from ear to ear',
'gfg': 'good f**king game',
'gfgi': 'go f**king google it',
'gfi': 'good f**king idea',
'gfj': 'good f**king job',
'gfl': 'grounded for life',
'gfm': 'god forgive me',
'gfo': 'go f**k off',
'gfu': 'go f**k yourself',
'gfurs': 'go f**k yourself',
'gfus': 'go f**k yourself',
'gfx': 'graphics',
'gfy': 'good for you',
'gfyd': 'go f**k your dad',
'gfym': 'go f**k your mom',
'gfys': 'go f**k yourself',
'gg': 'good game',
'gga': 'good game all',
'ggal': 'go get a life',
'ggf': 'go get f**ked',
'ggg': 'go, go, go!',
'ggi': 'go google it',
'ggnore': 'good game no rematch',
'ggp': 'gotta go pee',
'ggpaw': 'gotta go parents are watching',
'ggs': 'good games',
'ggwp': 'good game well-played',
'gh': 'good half',
'ghei': 'gay',
'ghey': 'gay',
'gigig': 'get it got it good',
'gigo': 'garbage in garbage out',
'gilf': "grandma i'd like to f**k",
'gim': 'google instant messanger',
'gimme': 'give me',
'gimmie': 'give me',
'gir': 'google it retard',
'gis': 'google image search',
'gitar': 'guitar',
'giv': 'give',
'giyf': 'google is your friend',
'gj': 'good job',
'gjial': 'go jump in a lake',
'gjp': 'good job partner',
'gjsu': 'god just shut up',
'gjt': 'good job team',
'gky': 'go kill yourself',
'gkys': 'go kill yourself',
'gl': 'good luck',
'gl hf': 'good luck, have fun',
'gl&hf': 'good luck and have fun',
'gla': 'good luck all',
'glbt': 'gay, lesbian, bisexual, transgenderd',
'glf': 'group looking for',
'glhf': 'good luck have fun',
'glln': 'got laid last night',
'glnhf': 'good luck and have fun',
'glty': 'good luck this year',
'glu': 'girl like us',
'glu2': 'good luck to you too',
'glux': 'good luck',
'glwt': 'good luck with that',
'gm': 'good morning',
'gma': 'grandma',
'gmab': 'give me a break',
'gmabj': 'give me a blowjob',
'gmafb': 'give me a f**king break',
'gmao': 'giggling my ass off',
'gmfao': 'giggling my f**king ass off',
'gmilf': "grandmother i'd like to f**k",
'gmod': 'global moderator',
'gmta': 'great minds think alike',
'gmtyt': 'good morning to you too',
'gmv': 'got my vote',
'gmybs': 'give me your best shot',
'gn': 'good night',
'gn8': 'good night',
'gna': 'going to',
'gnasd': 'good night and sweet dreams',
'gndn': 'goes nowhere,does nothing',
'gnfpwlbn': 'good news for people who love bad news',
'gng': 'going',
'gng2': 'going to',
'gngbng': 'gang bang',
'gnight': 'good night',
'gnite': 'good night',
'gnn': 'get naked now',
'gno': 'going to do',
'gnoc': 'get naked on cam',
'gnos': 'get naked on screen',
'gnr': "guns n' roses",
'gnrn': 'get naked right now',
'gnst': 'goodnight sleep tight',
'gnstdltbbb': "good night sleep tight don't let the bed bugs bite",
'goc': 'get on camera',
'goi': 'get over it',
'goia': 'get over it already',
'goin': 'going',
'gok': 'god only knows',
'gokid': 'got observers - keep it decent',
'gokw': 'god only knows why',
'gol': 'giggle out loud',
'gomb': 'get off my back',
'goml': 'get out of my life',
'gona': 'gonna',
'gonna': 'going to',
'good9': 'goodnite',
'goodmorning': 'good morning',
'gooh': 'get out of here!',
'goomh': 'get out of my head',
'gork': 'god only really knows',
'gosad': 'go suck a d**k',
'gotc': 'get on the computer',
'gotcha': 'got you',
'gotta': 'got to',
'gow': 'gears of war',
'goya': 'get off your ass',
'goyhh': 'get off your high horse',
'gp': 'good point',
'gpb': 'gotta pee bad',
'gpwm': 'good point well made',
'gpytfaht': 'gladly pay you tuesday for a hamburger today',
'gr8': 'great',
'gr8t': 'great',
'grad': 'graduation',
'grats': 'congratulations',
'gratz': 'congratulations',
'grfx': 'graphics',
'grillz': 'metal teeth',
'grl': 'girl',
'grmbl': 'grumble',
'grog': 'beer',
'grrl': 'girl',
'grtg': 'getting ready to go',
'grvy': 'groovy',
'gsad': 'go suck a d**k',
'gsave': 'global struggle against violent extremists',
'gsd': 'getting s**t done',
'gsfg': 'go search f**king google',
'gsi': 'go suck it',
'gsoh': 'good sense of humor',
'gsp': 'get some p***y',
'gsta': 'gangster',
'gt': 'get',
'gta': 'grand theft auto',
'gtas': 'go take a s**t',
'gtb': 'go to bed',
'gtf': 'get the f**k',
'gtfa': 'go the f**k away',
'gtfbtw': 'get the f**k back to work',
'gtfh': 'go to f**king hell',
'gtfo': 'get the f**k out',
'gtfoi': 'get the f**k over it',
'gtfon': 'get the f**k out noob',
'gtfooh': 'get the f**k out of here',
'gtfoomf': 'get the f**k out of my face',
'gtfu': 'grow the f**k up',
'gtfuotb': 'get the f**k up out this b***h',
'gtg': 'got to go',
'gtgb': 'got to go bye',
'gtgbb': 'got to go bye bye',
'gtgfn': 'got to go for now',
'gtglyb': 'got to go love you bye',
'gtgmmiloms': 'got to go my mum is looking over my shoulder',
'gtgn': 'got to go now',
'gtgp': 'got to go pee',
'gtgpp': 'got to go pee pee',
'gtgtb': 'got to go to bed',
'gtgtpirio': 'got to go the price is right is on',
'gtgtwn': 'got to go to work now',
'gth': 'go to hell',
'gtha': 'go the hell away',
'gthb': 'go to hell b***h',
'gthmf': 'go to hell mothaf**ka',
'gtho': 'get the hell out',
'gthu': 'grow the heck up',
'gthyfah': 'go to hell you f**king a**h**e',
'gtk': 'good to know',
'gtm': 'giggling to myself',
'gtn': 'getting',
'gtp': 'got to pee',
'gtr': 'got to run',
'gts': 'going to school',
'gtsy': 'good to see you',
'gttp': 'get to the point',
'gtty': 'good talking to you',
'gu': 'grow up',
'gu2i': 'get used to it',
'gud': 'good',
'gudd': 'good',
'gui': 'graphical user interface',
'gurl': 'girl',
'gurlz': 'girls',
'guru': 'expert',
'gw': 'good work',
'gweetin': 'greeting',
'gwijd': 'guess what i just did',
'gwm': 'gay white male',
'gwork': 'good work',
'gwrk': 'good work',
'gws': 'get well soon',
'gwytose': 'go waste your time on someone else',
'gy': 'gay',
'gyal': 'girl',
'gypo': 'get your penis out'}
h = {'h&k': 'hugs and kisses',
'h*r': 'homestar runner',
'h+k': 'hugs and kisses',
'h.o': 'hold on',
'h/e': 'however',
'h/mo': 'homo',
'h/o': 'hold on',
'h/u': 'hold up',
'h/w': 'homework',
'h2': 'halo 2',
'h2gtb': 'have to go to the bathroom',
'h2o': 'water',
'h2sys': 'hope to see you soon',
'h3y': 'hey',
'h4kz0r5': 'hackers',
'h4x': 'hacks',
'h4x0r': 'hacker',
'h4xor': 'hacker',
'h4xr': 'hacker',
'h4xrz': 'hackers',
'h4xx0rz': 'hacker',
'h4xxor': 'hacker',
'h8': 'hate',
'h80r': 'hater',
'h82sit': 'hate to say it',
'h83r': 'hater',
'h8ed': 'hated',
'h8er': 'hater',
'h8r': 'hater',
'h8red': 'hatred',
'h8s': 'hates',
'h8t': 'hate',
'h8t0r': 'hater',
'h8t3r': 'hater',
'h8te': 'hate',
'h8ter': 'hater',
'h8tr': 'hater',
'h8u': 'i hate you',
'h9': 'husband in room',
'h@x0r': 'hacker',
'habt': 'how about this',
"hadn't": 'had not',
"hadn't've": 'had not have',
'hafta': 'have to',
'hagd': 'have a good day',
'hagl': 'have a great life',
'hagn': 'have a good night',
'hago': 'have a good one',
'hags': 'have a great summer',
'hai': 'hello',
'hait': 'hate',
'hak': "here's a kiss",
'hakas': 'have a kick ass summer',
'hammrd': 'hammered',
'han': 'how about now',
"hasn't": 'has not',
'hau': 'how are you',
'hav': 'have',
"haven't": 'have not',
'havnt': "haven't",
'hawf': 'husband and wife forever',
'hawt': 'hot',
'hawtie': 'hottie',
'hax': 'hacks',
'hax0r': 'hacker',
'hax0red': 'hacked',
'hax0rz': 'hackers',
'haxer': 'hacker',
'haxor': 'hacker',
'haxoring': 'hacking',
'haxors': 'hackers',
'haxorz': 'hackers',
'haxxor': 'hacker',
'haxxzor': 'hacker',
'haxz0r': 'hacker',
'haxzor': 'hacker',
'hayd': 'how are you doing',
'hazzunt' : 'has not',
'hb': 'hurry back',
'hb4b': 'hoes before bros',
'hbd': 'happy birthday',
'hbic': 'head b***h in charge',
'hbii': 'how big is it',
'hbu': 'how about you',
'hbuf': 'how about your family',
'hby': 'how about you',
'hc': 'how come',
'hcbt1': 'he could be the one',
'hcib': 'how can it be',
'hcihy': 'how can i help you',
'hdop': 'help delete online predators',
'hdu': 'how dare you',
'hdydi': 'how do you do it',
'hdydt': 'how did you do that',
"he'd": 'he would',
"he'd've": 'he would have',
"he'll": 'he will',
"he'll've": 'he he will have',
"he's": 'he is',
'headin': 'heading',
'heh': 'haha',
'hella': 'very',
'hes': 'he is',
'hey': 'hi',
'heya': 'hey',
'heyt': 'hate',
'heyy': 'hello',
'heyya': 'hello',
'hf': 'have fun',
'hfn': 'hell f**king no',
'hfs': 'holy f**king s**t!',
'hfsbm': 'holy f**king s**t batman',
'hfwt': 'have fun with that',
'hg': 'hockeygod',
'hght': 'height',
'hhiad': 'holy hole in a doughnut',
'hhiadb': 'holy hole in a donut batman',
'hhok': 'ha ha only kidding',
'hhyb': 'how have you been',
'hi2u': 'hello',
'hi2u2': 'hello to you too',
'hiet': 'height',
'hiik': 'hell if i know',
'hijack': 'start an off topic discussion',
'hith': 'how in the hell',
'hiw': 'husband is watching',
'hiya': 'hello',
'hiybbprqag': "copying somebody else's search results",
'hj': 'hand job',
'hl': 'half-life',
'hl2': 'half-life 2',
'hla': 'hot lesbian action',
'hlb': 'horny little b*****d',
'hld': 'hold',
'hldn': 'hold on',
'hldon': 'hold on',
'hll': 'hell',
'hlm': 'he loves me',
'hlo': 'hello',
'hlp': 'help',
'hly': 'holy',
'hlysht': 'holy s**t',
'hmb': 'hold me back',
'hmewrk': 'homework',
'hmfic': 'head mother f**ker in charge',
'hml': 'hate my life',
'hmoj': 'holy mother of jesus',
'hmp': 'help me please',
'hmu': 'hit me up',
'hmul': 'hit me up later',
'hmus': 'hit me up sometime',
'hmw': 'homework',
'hmwk': 'homework',
'hmwrk': 'homework',
'hng': 'horny net geek',
'hngry': 'hungry',
'hnic': 'head n****r in charge',
'ho': 'hold on',
'hoas': 'hang on a second',
'hoay': 'how old are you',
'hodl': 'hold',
'hoh': 'head of household',
'hom': 'home',
'homey': 'friend',
'homie': 'good friend',
'homo': 'homosexual',
'hoopty': 'broke down automobile',
'hott': 'hot',
"how'd": 'how did',
"how'd'y": 'how do you',
"how'll": 'how will',
"how's": 'how is',
'howdey': 'hello',
'howz': 'hows',
'hpb': 'high ping b*****d',
'hpoa': 'hot piece of ass',
'hppy': 'happy',
'hpy': 'happy',
'hpybdy': 'happy birthday',
'hr': 'hour',
'hre': 'here',
'hrny': 'horny',
'hrs': 'hours',
'hru': 'how are you',
'hrud': 'how are you doing',
'hruf': 'how are you friend',
'hs': 'headshot',
'hsd': 'high school dropout',
'hsik': 'how should i know',
'hsr': 'homestar runner',
'hss': 'horse s**t and splinters',
'hswm': 'have sex with me',
'ht': 'heard through',
'htc': 'hit the cell',
'htf': 'how the f**k',
'htfu': 'hurry the f**k up',
'htg': 'have to go',
'hth': 'hope that helps',
'hthu': 'hurry the hell up',
'htr': 'hater',
'http': 'hyper text transfer protocol',
'hu': 'hey you',
'hubby': 'husband',
'hud': 'heads up display',
'huggle': 'hug and cuddle',
'hugz': 'hugs',
'hun': 'honey',
'hv': 'have',
'hve': 'have',
'hvnt': "haven't",
'hw': 'homework',
'hw/hw': 'help me with homework',
'hwg': 'here we go',
'hwga': 'here we go again',
'hwik': 'how would i know',
'hwk': 'homework',
'hwmbo': 'he who must be obeyed',
'hwmnbn': 'he who must not be named',
'hwms': 'hot wild monkey sex',
'hwu': "hey what's up",
'hwz': 'how is',
'hxc': 'hardcore',
'hy': 'hell yeah',
'hyb': 'how you been',
'hyg': 'here you go',
'hyk': 'how you know'}
i = {"i'd": 'i would',
"i'd've": 'i would have',
"i'll": 'i will',
"i'll've": 'i will have',
"i'm": 'i am',
"i've": 'i have',
'i <3 u': 'i love you',
'i c': 'i see',
'i8': 'alright',
'i8u': 'i hate you',
'i<3 u': 'i love you',
'i<3u': 'i love you',
'iab': 'i am bored',
'iafh': 'i am f**king hot',
'iafi': 'i am from india',
'iag': "it's all good",
'iah': 'i am horny',
'iai': 'i am interested',
'iajsn': 'i am just scraping noslang.com',
'ianabs': 'i am not a brain surgeon',
'ianacl': 'i am not a copyright lawyer',
'ianal': 'i am not a lawyer',
'ianalb': 'i am not a lawyer, but..',
'ianars': 'i am not a rocket scientist',
'ians': 'i am not sure',
'ianyl': 'i am not your lawyer',
'iap': 'i am pissed',
'iasb': 'i am so bored',
'iatb': 'i am the best',
'iateu': 'i hate you',
'iavb': 'i am very bored',
'iaw': 'in another window',
'iawtc': 'i agree with this comment',
'iawtp': 'i agree with this post',
'iawy': 'i agree with you',
'ib': "i'm back",
'ibbl': "i'll be back later",
'ibcd': 'idiot between chair & desk',
'ibs': 'internet b***h slap',
'ibt': "i'll be there",
'ibtl':'in before the lock',
'ibw':"i'll be waiting",
'ic': 'i see',
'icb': "i can't believe",
'icbi': "i can't believe it",
'icbiwoop': 'i chuckled, but it was out of pity.',
'icbt': "i can't believe that",
'icbyst': 'i cant believe you said that',
'iccl': 'i could care less',
'icgup': 'i can give you pleasure',
'icic': 'i see. i see',
'icp': 'insane clown posse',
'icq': "i can't rememer",
'icr': 'i seek you (also a chat program)',
'icsrg': 'i can still reach google',
'ictrn': "i can't talk right now",
'icty': "i can't tell you",
'icu': 'i see you',
'icudk': "in case you didn't know",
'icup': 'i see you pee',
'icw': 'i care why?',
'icwudt': 'i see what you did there',
'icwum': 'i see what you mean',
'icydci': "in csae you didn't know",
'icydk': "in case you didn't catch it",
'icydn': "in case you didn't know",
'icymi': 'in case you missed it',
'id10t': 'idiot',
'idac': "i don't actually care",
'idak': "i don't actually know",
'idbtwdsat': "i don't believe they would do such a thing",
'idby': "i don't believe you",
'idc': "i don't care",
'iddi': "i didn't do it",
'idec': "i don't even care",
'idek': "i don't even know",
'idfc': "i don't f**king care",
'idfk': "i don't f**king know",
'idfts': "i don't f**king think so",
'idgac': "i don't give a crap",
'idgad': "i don't give a damn",
'idgaf': "i don't give a f**k",
'idgaff': "i don't give a flying f**k",
'idgafs': "i don't give a f**king s**t",
'idgara': "i don't give a rat's ass",
'idgas': "i don't give a s**t",
'idgi': "i don't get it",
'idjit': 'idiot',
'idk': "i don't know",
'idkbibt': "i don't know but i've been told",
'idke': "i don't know ethier",
'idkh': "i don't know how",
'idkh2s': "i don't know how to spell",
'idkt': "i don't know that",
'idkw': "i don't know why",
'idkwiwdwu': "i don't know what i would do without you",
'idkwtfytb': "i don't know what to say",
'idkwts': "i dont know what the f**k you're talking about",
'idkwurta': "i don't know what you are talking about.",
'idkwym': "i don't know what you mean",
'idky': "i don't know you",
'idkyb': "i don't know why but",
'idkymb2': "i didn't know yoru mom blogs too",
'idl': "i don't like",
'idli': "i don't like it",
'idlu': "i don't like you",
'idly': "i don't like you",
'idlyitw': "i don't like you in that way",
'idm': "i don't mind",
'idmhw': 'im doing my homework',
'idn': "i don't know",
'idnk': "i don't know",
'idno': 'i do not know',
'idntk': 'i dont need to know',
'idnwths': 'i do not want to have sex',
'idonno': 'i do not know',
'idop': 'it depends on price',
'idot': 'idiot',
'idr': "i don't remember",
'idrc': "i don't really care",
'idrfk': "i don't really f**king know",
'idrgaf': "i don't really give a f**k",
'idrgaff': "i don't really give a flying f**k",
'idrk': "i don't really know",
'idrts': "i don't really think so",
'idsw': "i don't see why",
'idtis': "i don't think i should",
'idtkso': "i don't think so",
'idts': "i don't think so",
'idu': "i don't understand",
'idunno': 'i do not know',
'iduwym': "i don't understand what you mean.",
'idw2': "i don't want to",
'idw2n': "i don't want to know",
'idwk': "i don't wanna know",
'idwt': "i don't want to",
'idwtg': "i don't want to go",
'idyat': 'idiot',
'iebkac': 'issue exists between keyboard and chair',
'ietf': 'internet engineering task force',
'iff': 'if and only if',
'ifhu': 'i f**king hate you',
'ifhy': 'i f**king hate you',
'ifk': 'i f**king know',
'iflu': 'i f**king love you',
'ifthtb': 'i find that hard to belive',
'ifttt': 'if this then that',
'ifwis': 'i forgot what i said',
'ig': 'i guess',
'ig2g': 'i got to go',
'ig5oi': 'i got 5 on it',
'igahp': "i've got a huge penis",
'igalboc': "i've got a lovely bunch of cocnuts",
'igg': 'i gotta go',
'ight': 'alright',
'igkymfa': "i'm gonna kick your mother f**king ass",
'igs': 'i guess so',
'igt': 'i got this',
'igtg': "i've got to go",
'igtgt': 'i got to go tinkle',
'igtkya': 'im going to kick your ass',
'igu': 'i give up',
'igyb': 'i got your back',
'ih': 'it happens',
'ih2gp': 'i have to go pee',
'ih2p': "i'll have to pass",
'ih8': 'i hate',
'ih8evry1': 'i hate everyone',
'ih8mls': 'i hate my little sister',
'ih8p': 'i hate parents',
'ih8tu': 'i hate you',
'ih8u': 'i hate you',
'ih8usm': 'i hate you so much',
'ih8y': 'i hate you',
'ihac': 'i have a customer',
'ihat3u': 'i hate you',
'ihavent': 'i hope i spelled that right',
'ihistr': 'i have not',
'ihiwydt': 'i hate it when you do that',
'ihml': 'i hate my life',
'ihmp': 'i hate my parents',
'ihnc': 'i have no clue',
'ihnfc': 'i have no f**king clue',
'ihni': 'i have no idea',
'iht': 'i heard that',
'ihtfp': 'i hate this f**king place',
'ihtgttbwijd': 'i have to go to the bathroom, wait i just did.',
'ihtp': 'i have to poop',
'ihtsm': 'i hate this so much',
'ihtutbr': 'i have to use the bathroom',
'ihu': 'i hate you',
'ihurg': 'i hate your guts',
'ihusb': 'i hate you so bad',
'ihusfm': 'i hate you so f**king much',
'ihusm': 'i hate you so much',
'ihy': 'i hate you',
'ihya': 'i hate you all',
'ihysm': 'i hate you so much',
'ihysmrn': 'i hate you so much right now',
'ii': 'alright',
'iigh': 'i',
'iight': 'alright',
'iiok': 'is it okay',
'iirc': 'if i recall correctly',
'iistgtbtipi': 'if it sounds too good to be true it probably is',
'iit': 'is it tight',
'iitywimwybmad': 'if i tell you what it means will you buy me a drink',
'iitywybmad': 'if i tell you, will you buy me a drink?',
'iiuc': 'if i understand correctly',
'iiw2': 'is it web 2.0?',
'iiwii': 'it is what it is',
'ij': 'indide joke',
'ijaf': "it's just a fact",
'ijcomk': 'i just came on my keyboard',
'ijdk': "i just don't know",
'ijdl': 'i just died laughing',
'ijeomk': 'i just ejaculated on my keybord',
'ijf': 'i just farted',
'ijgl': 'i just got laid',
'ijit': 'idiot',
'ijk': "i'm just kidding",
'ijp': 'internet job posting',
'ijpmp': 'i just peed my pants',
'ijpms': 'i just pissed myself',
'ijr': 'i just remembered',
'ijs': "i'm just saying",
'ijsabomomcibstg': 'i just saved a bunch of money on my car insurance by switching to geico',
'ik': 'i know',
'iki': 'i know it',
'ikic': 'i know i can',
'ikm': 'i know man',
'ikr': 'i know really',
'ikt': 'i knew that',
'ikwud': 'i know what you did',
'ikwum': 'i know what you meant',
'ikwyl': 'i know where you live',
'ikwym': 'i know what you mean',
'ilbbaicnl': 'i like big butts and i can not lie',
'ilbcnu': "i'll be seeing you",
'ilcul8r': "i'll see you later",
'ilhsm': 'i love him/her so much',
'ili': 'i love it',
'ilk2fku': 'i would like to f**k you',
'ilml': 'i love my life',
'ilms': 'i love my self',
'ilotibinlirl': "i'm laughing on the internet, but i'm not laughing in real life",
'ilshipmp': 'i laughed so hard i peed my pants',
'iltf': 'i love to f**k',
'iltwummf': 'i love the way you make me feel',
'iltwymmf': 'i love the way you make me feel',
'ilu': 'i love you',
'ilu2': 'i love you too',
'iluaaf': 'i love you as a friend',
'ilul': 'i love you loads',
'ilulafklc': 'i love you like a fat kid loves cake.',
'ilum': 'i love you more',
'ilusfm': 'i love you so f**king much',
'ilusm': 'i love you so much',
'ilyt': 'i love you too',
'ilyttmab': 'i love you to the moon and back',
'iluvm': 'i love you very much',
'iluvu': 'i love you',
'iluvya': 'i love you',
'iluwamh': 'i love you with all my heart',
'ilvu': 'i love you',
'ily': 'i love you',
'ily2': 'i love you too',
'ily4e': 'i love you forever',
'ily4ev': 'i love you forever',
'ilyaas': 'i love you as a sister',
'ilyal': 'i like you a lot',
'ilyb': 'i love you b***h',
'ilybby': 'i love you baby',
'ilybtid': "i love you but then i don't",
'ilyf': "i'll love you forever",
'ilygsm': 'i love you guys so much',
'ilykthnxbai': 'i love you k thanks bye',
'ilyl': 'i love you loads',
'ilylab': 'i love you like a brother',
'ilylabf': 'i love you like a best friend',
'ilylafklc': 'i love you like a fat kid loves cake',
'ilylas': 'i love you like a sister',
'ilylc': 'i love you like crazy',
'ilym': 'i love you more',
'ilymtyk': 'i love you more than you know',
'ilymtylm': 'i love you more than you love me',
'ilys': 'i love you sexy',
'ilysb': 'i love you so bad',
'ilysfm': 'i love you so f**king much',
'ilysfmb': 'i love you so f**king much baby',
'ilysm': 'i love you so much',
'ilysmih': 'i love you so much it hurts',
'ilysmm': 'i love you so much more',
'ilysmydek': "i love you so much you don't even know",
'ilysvm': 'i love you so very much',
'ilyvm': 'i love you very much',
'ilywamh': 'i love you with all my heart',
'im': 'instant message',
"im'd": 'i am',
'im26c4u': 'i am too sexy for you',
'ima': 'i am a',
'imao': 'in my arrogant opinion',
'imb': 'i am back',
'imcdo': 'in my conceited dogmatic opinion',
'imed': 'instant messaged',
'imfao': 'in my f**king arrogant opinion',
'imfo': 'in my f**king opinion',
'imh': 'i am here',
'imhbco': 'in my humble but correct opinion',
'imhe': 'in my humble experience',
'imho': 'in my humble opinion',
'imm': 'instant message me',
'imma': "i'm going to",
'imnerho': 'in my not even remotely humble opinion',
'imnl': "i'm not laughing",
'imnshmfo': 'in my not so humble mother f**king opinion',
'imnsho': 'in my not so humble opinion',
'imo': 'in my opinion',
'imoo': 'in my own opinion',
'impo': 'in my personal opinion',
'impov': 'in my point of view',
'imr': 'in my room',
'imsb': 'i am so bored',
'imsry': 'i am sorry',
'imtaw': 'it may take a while',
'imts': 'i meant to say',
'imu': 'i miss you',
'imusm': 'i miss you so much',
'imvho': 'in my very humble opinion',
'imwtk': 'inquiring minds want to know',
'imy': 'i miss you',
'imy2': 'i miss you to',
'imya': 'i miss you already',
'imysfm': 'i miss you so f**king much',
'in2': 'into',
'inb4': 'in before',
'inbd': "it's no big deal",
'incld': 'include',
'incrse': 'increase',
'ind2p': 'i need to pee',
'indie': 'independent',
'inef': "it's not even funny",
'inet': 'internet',
'inh': 'i need help',
'inho': 'in my honest opinion',
'inhwh': 'i need homework help',
'init': "isn't it",
'inmp': "it's not my problem",
'innit': "isn't it",
'ino': 'i know',
'instagib': 'instant kill',
'instakill': 'instant kill',
'intarwebs': 'internet',
'intel': 'intelligence',
'interweb': 'internet',
'intpftpotm': 'i nominate this post for the post of the month',
'inttwmf': 'i am not typing this with my fingers',
'invu': 'i envy you',
'ioh': "i'm out of here",
'iois': 'indicators of interest',
'iokiya': "it's ok if you are",
'iomw': "i'm on my way",
'ionno': "i don't know",
'iono': "i don't know",
'iotd': 'image of the day',
'iou': 'i owe you',
'iow': 'in other words',
'ioya': "i'd own your ass",
'ioyk': 'if only you knew',
'ip': 'internet protocol',
'irc': 'internet relay chat',
'ircd': 'internet relay chat daemon',
'ircx': 'internet relay chat extension',
'irdc': "i really don't care",
'irdgaf': "i really don't give a f**k",
'irdk': "i really don't know",
'irgtgbtw': "i've really got to get back to work",
'irhtgttbr': 'i really have to go to the bathroom',
'irhy': 'i really hate you',
'irl': 'in real life',
'irly': 'i really love you',
'irt': 'in reply to',
'irtf': "i'll return the favor",
'is2g': 'i swear to god',
'isb': "i'm so bored.",
'isbya': 'im sorry but you asked',
'isd': 'internet slang dictionary',
'isdc': "i so don't care",
'ise': 'internal server error',
'isfly': 'i so f**king love you',
'isg': 'i speak geek',
'ishii': 'i see how it is',
'isianmtu': 'i swear i am not making this up',
'isj': 'inside joke',
"isn'": 'in search of',
"isn't": 'is in it',
'iso': 'is not',
'isp': 'internet service provider',
'iss': 'im so sorry',
'istg': 'i swear to god',
'istr': 'i seem to remember',
'istwfn': 'i stole this word from noslang.com',
'iswydt': 'i see what you did there',
"it'd": 'i totally agree',
"it'd've": 'it would',
"it'll": 'it would have',
"it'll've": 'it will',
"it's": 'it is',
'ita': 'it is',
'itb': 'in the butt',
'itc': 'in that case',
'itd': 'in the dark',
'ite': 'alright',
'itk': 'in the know',
'itn': 'i think not',
'itt': 'in this thread',
'ittif': 'i think that is fine',
'ityltk': "i thought you'd like to know",
'itys': 'i told you so',
'itz': "it's",
'itzk': "it's ok",
'iucmd': 'if you catch my drift',
'iukwim': 'if you know what i mean',
'iunno': "i don't know",
'iuno': 'i dunno',
'iv': 'i have',
'ive': 'i have',
'iw2f': 'i want to f**k',
'iw2fu': 'i want to f**k you',
'iw2mu': 'i want to meet you',
'iwaa': 'it was an accident',
'iwbrbl@r': 'i will be right back later',
'iwc': 'in which case',
'iwfusb': 'i wanna f**k you so bad',
'iwfy': 'i want to f**k you',
'iwfybo': 'i will f**k your brains out',
'iwg': 'it was good',
'iwhi': 'i would hit it',
'iwhswu': 'i want to have sex with you',
'iwjk': 'i was just kidding',
'iwk': "i wouldn't know",
'iwlu4e': 'i will love you for ever',
'iwmu': 'i will miss you',
'iwmy': 'i will miss you',
'iws': 'i want sex',
'iwsn': 'i want sex now',
'iwsul8r': 'i will see you later',
'iwtfu': 'i want to f**k you',
'iwtfy': 'i want to f**k you',
'iwthswy': 'i want to have sex with you',
'iwtk': 'i want to know',
'iwtly': 'i want to love you',
'iwu': 'i want you',
'iwuwh': 'i wish you were here',
'iwy': 'i want you',
'iwyb': 'i want your body',
'iwyn': 'i want you now',
'iwythmb': 'i want you to have my baby',
'iya': 'in your ass',
'iyam': 'if you ask me',
'iyc': 'if you can',
'iyd': 'in your dreams',
'iydhantsdsaaa': "if you don't have anything nice to say don't say anything at all",
'iydmma': "if you don't mind me asking",
'iyf': 'in your face',
'iyflg': "if you're feeling less generous",
'iygm': 'if you get me',
'iykwim': 'if you know what i mean',
'iym': 'i am your man',
'iyo': 'in your opinion',
'iyq': 'i like you',
'iyss': 'if you say so',
'iyswim': 'if you see what i mean',
'iywt': 'if you want to'}
j = {'j-c': 'just chilling',
'j/a': 'just asking',
'j/c': 'just curious',
'j/j': 'just joking',
'j/k': 'just kidding',
'j/o': 'jackoff',
'j/p': 'just playing',
'j/s': 'just saying',
'j/t': 'just talking',
'j/w': 'just wondering',
'j00': 'you',
'j00r': 'your',
'j2bs': 'just to be sure',
'j2c': 'just too cute',
'j2f': 'just too funny',
'j2luk': 'just to let you know',
'j2lyk': 'just to let you know',
'j4f': 'just for fun',
'j4g': 'just for grins',
'j4l': 'just for laughs',
'j4u': 'just for you',
'jalaudlm': "just as long as you don't leave me",
'jas': 'just a second',
'jb': 'jailbait',
'jbu': 'just between us',
'jc': 'just curious',
'jcam': 'just checking away message',
'jcath': 'just chilling at the house',
'jdfi': 'just f**king do it',
'jdi': 'just do it',
'jebus': 'jesus',
'jeomk': 'just ejaculated on my keyboard',
'jf': 'just fooling',
'jfc': 'jesus f**king christ',
'jfdi': 'just f**king do it!',
'jff': 'just for fun',
'jfg': 'just for giggles',
'jfgi': 'just f**king google it',
'jfi': 'just forget it',
'jfj': 'jump for joy',
'jfk': 'just f**king kidding',
'jfl': 'just for laughs',
'jflts': 'just felt like typing something',
'jfn': 'just for now',
'jfo': 'just f**k off',
'jfr': 'just for reference',
'jftr': 'just for the record',
'jfu': 'just for you',
'jfwy': 'just f**king with you',
'jg2h': 'just go to hell',
'jgiyn': 'just google it you noob',
'jgtfooh': 'just get the f**k out of here',
'jh': 'just hanging',
'jhm': 'just hold me',
'jho': 'just hanging out',
'jic': 'just in case',
'jit': 'just in time',
'jizz': 'semen',
'jj': 'just joking',
'jj/k': 'just joking',
'jja': 'just joking around',
'jk': 'just kidding',
'jka': 'just kidding around',
'jking': 'joking',
'jkl': 'just kidding loser',
'jklol': 'just kidding laughing out loud',
'jkn': 'joking',
'jks': 'jokes',
'jkz': 'jokes',
'jlma': 'just leave me alone',
'jlt': 'just like that',
'jm': 'just messing',
'jma': 'just messing around',
'jml': 'just my luck',
'jmo': 'just my opinion',
'jms': 'just making sure',
'jom': 'just one minuite',
'joo': 'you',
'jooc': 'just out of curiosity',
'jooce': 'juice',
'joor': 'your',
'jp': 'just playing',
'js': 'just saying',
'jsa': 'just stop already',
'jsing': 'just saying',
'jst': 'just',
'jsuk': 'just so you know',
'jsun': 'just so you know',
'jsut': 'just',
'jsyk': 'just so you know',
'jsyn': 'just so you know',
'jtay': 'just thinking about you',
'jtbs': 'just to be sure',
'jtc': 'join the club',
'jtfo': 'joke the f**k out',
'jtluk': 'just to let you know',
'jtlyk': 'just to let you know',
'jtoi': 'just thought of it',
'jtol': 'just thinking out loud',
'jttsiowctw': 'just testing to see if other websites copy this word',
'jtty': 'just to tell you',
'jtumltk': 'just thought you might like to know',
'jtwii': 'just the way it is.',
'jtwiw': 'just the way it was.',
'jtyltk': "just thought you'd like to know",
'jtysk': 'just thought you should know',
'jumping the couch': 'acting strange',
'jus': 'just',
'juss': 'just',
'juz': 'just',
'juzt': 'just',
'jw': 'just wondering',
'jw2k': 'just wanted to know',
'jwas': 'just wait a second',
'jwing': 'just wondering',
'jwtlyk': 'just wanted to let you know',
'jyfihp': 'jam your finger in her p***y'}
k = {'k': 'ok',
'k3wl': 'cool',
'ka': 'kick ass',
'kafn': 'kill all f**king noobs',
'kah': 'kisses and hugs',
'kaw': 'kick ass work',
'kay': 'okay',
'kb': 'kilobite',
'kcco': 'keep calm chive on',
'kek': 'laughing out loud',
'kewel': 'cool',
'kewl': 'cool',
'kfc': 'kentucky fried chicken',
'khitbash': 'kick her in the box and shove her',
'khuf': 'know how you feel',
'kia': 'killed in action',
'kib': 'okay, im back',
'kic': 'keep it clean',
'kicks': 'sneakers',
'kig': 'keep it going',
'kiled': 'killed',
'kinda': 'kind of',
'kir': 'kid in room',
'kis': 'keep it simple',
'kisa': 'knight in shining armor',
'kiss': 'keep it simple, stupid',
'kit': 'keep in touch',
'kitfo': 'knock it the f**k off',
'kitteh': 'kitten',
'kiu': 'keep it up',
'kiwf': 'kill it with fire',
'kk': 'ok',
'kkk': 'ku klux klan',
'kkthnxbye': 'okay thanks bye',
'kky': 'kinky',
'kl': 'cool',
'km': 'kiss me',
'kma': 'kiss my ass',
'kmag': 'kiss my ass goodbye',
'kmao': 'kick my ass off',
'kmb': 'kiss my butt',
'kmfa': 'kiss my f**king ass',
'kmhba': 'kiss my hairy big ass',
'kml': 'killing myself laughing',
'kmn': 'kill me now',
'kmp': 'kill me please',
'kmsl': 'killing myself laughing',
'kmswl': 'killing myself with laughter',
'kmu': 'kinda miss you',
'knackered': 'drunk',
'knewb': 'new player',
'knn': 'f**k your mother',
'kno': 'know',
'kno!': 'know',
'knw': 'know',
'ko': 'knock out',
'kol': 'kiss on lips',
'koo': 'cool',
'kool': 'cool',
'kos': 'kid over shoulder',
'kotc': 'kiss on the cheek',
'kotd': 'kicks of the day',
'kotl': 'kiss on the lips',
'kotor': 'knights of the old republic',
'kots': 'keep on talking s**t',
'kpc': 'keeping parents clueless',
'kq': 'keep quiet',
'ks': 'kill steal',
'kss': 'kiss',
'kssd': 'kissed',
'ksw': 'ok so what',
'kt': 'keep talking',
'ktc': 'kill the cat',
'ktfo': 'knocked the f**k out',
'kthanxbi': 'okay, thanks. bye.',
'kthnxbai': 'okay, thanks, bye',
'kthnxbye': 'okay, thanks, bye',
'kthx': 'ok, thank you',
'kthxbai': 'ok thanks bye!',
'kthxbi': 'ok, thank you, goodbye',
'kthxbye': 'ok, thank you, goodbye',
'kthxgb': 'ok thanks goodbye',
'kthxmn': 'ok thanks man',
'kthz': 'ok thanks',
'ktnx': 'okay and thanks',
'kuhl': 'cool',
'kul': 'cool',
'kute': 'cute',
'kutgw': 'keep up the good work',
'kuwl': 'cool',
'kwik': 'quick',
'kwim': 'know what i mean',
'kwis': "know what i'm saying?",
'kwit': 'quit',
'kwiz': 'quiz',
'kwl': 'cool',
'kwtsds': "kiss where the sun don't shine",
'kyag': 'kiss your ass goodbye',
'kyfag': 'kiss your f**king ass goodbye',
'kyfc': 'keep your fingers crossed',
'kyko': 'keep your knickers on',
'kyms': 'keep your mouth shut',
'kys': 'kill yourself'}
# +
l = { 'l0lz': 'laugh out loud',
'l2': 'learn to',
'l2m': 'listening to music',
'l2ms': 'laughing to myself',
'l2p': 'learn to play',
'l2r': 'learn to read',
'l337': 'elite',
'l33t': 'elite',
'l4m3rz': 'lamers',
'l8': 'late',
'l84skool': 'late for school',
'l8a': 'later',
'l8er': 'later',
'l8ers': 'later',
'l8r': 'see you later',
'l8rs': 'laters',
'l8rz': 'later',
'l8s': 'later',
'l8t': 'later',
'l8ta': 'later',
'l8ter': 'later',
'l8tr': 'later',
'l@u': 'laughing at you',
'laff': 'laugh',
'lafs': 'love at first sight',
'lak': 'love and kisses',
'lal': 'laughing a little',
'lalol': 'lots and lots of laughs',
'lam': 'leave a message',
'lamf': 'like a motherf**ker',
'lan': 'local area network',
'lappy': 'laptop',
'larp': 'live action role-play',
'lasb': 'lame ass stupid b***h',
'lat': 'laugh at that',
'lata': 'later',
'lates': 'later',
'latn': 'laugh at the newbs',
'latr': 'later',
'latwttb': 'laughing all the way to the bank',
'lau': 'laugh at you',
'lawd': 'lord',
'lawl': 'lauging out loud with a southern drawl',
"lawl'd": 'laughed out loud',
'lawled': 'laughed out loud',
'lawls': 'laughing out loud with a southern drawl',
'lawlz': 'laughing out loud with a southern drawl',
'lazer': 'laser',
'lazor': 'laser',
'lbh': "let's be honest",
'lbnolbiwfetalol': "laughing but not out loud because it wasn't funny enough to actually laugh out loud",
'lbnr': 'laughing but not really',
'lbo': 'laughing butt off',
'lbr': "little boy's room",
'lbvs': 'laughing but very serious',
'lcsnpc': 'low cost small notebook personal computer',
'ldr': 'long-distance relationship',
'lee7': 'elite',
'leet': 'elite',
'legit': 'legitimate',
'leik': 'like',
'leme': 'let me',
'lemme': 'let me',
'lesbo': 'lesbian',
'less than 3': 'love',
'less than three': 'love',
"let's": 'let us',
'lez': 'lesbian',
'lezbean': 'lesbian',
'lezbo': 'lesbian',
'lezzzie': 'lesbian',
'lf': 'looking for',
'lf1m': 'looking for one more',
'lf2m': 'looking for 2 more',
'lfg': 'looking for group',
'lfl': "let's f**k later",
'lfm': 'looking for mate',
'lfnar': 'laughing for no aparent reason',
'lfp': 'looking for p***y',
'lfr': 'laughing for real',
'lgb': 'lesbian/gay/bisexual',
'lgbnaf': 'lets get butt naked and f**k',
'lgbtq': 'lesbien, gay, bisexual, transgender and queer.',
'lgds': "let's go do something",
'lgf': 'little green footballs',
'lggd': "let's go get drunk",
'lgn': 'link goes nowhere',
'lgo': 'life goes on',
'lgot': "let's go out tonight",
'lgr': 'little girls room',
'lgs': "let's go shopping!",
'lgtm': 'looks good to me',
'lhao': 'laughing her ass off',
'lhs': 'lets have sex',
'lhsrn': "let's have sex right now",
'lia': 'life is awesome',
'liaoyb': "like it's any of your business",
'lic': 'like i care',
'liec': 'like i even care',
'liek': 'like',
'liekz': 'likes',
'lifo': 'last in first out',
'ligad': 'like i give a damn',
'ligaff': 'like i give a flying f**k',
'ligafs': 'like i give a flying s**t',
'ligas': 'like i give a s**t',
'lih': 'laugh in head',
'liita': 'love is in the air',
'lik': 'like',
'lil': 'little',
'lim': 'like it matters',
'limh': 'laugh in my head',
'liol': 'laugh insanely out loud',
'lirl': 'laughing in real life',
'liu': 'look it up',
'liv': 'live',
'liyf': 'laughing in your face',
'lj': 'live journal',
'lk': 'like',
'lke': 'like',
'llab': 'laughing like a b***h.',
'llap': 'live long and prosper',
'llc': 'laughing like crazy',
'llf': 'laugh like f**k',
'llh': 'laughing like hell',
'llol': 'literally laughing out loud',
'lltnt': 'live like theres no tomorrow',
'lm4aq': "let's meet for a quickie.",
'lma': 'leave me alone',
'lmamf': 'leave me alone mother f**ker',
'lmao': 'laughing my ass off',
'lmaol': 'laughing my ass out loud',
'lmaomtoaoa': 'laugh my ass off many times over and over again',
'lmaonade': 'laughing my ass off',
'lmaool': 'laughing my ass off out loud',
'lmaootf': 'laughing my ass off on the floor',
'lmaorof': 'laughing my ass off rolling on the floor',
'lmaorotf': 'laughing my ass off rolling on the floor',
'lmaowrotf': 'laughing my ass of while rolling on the floor',
'lmaowtntpm': 'laughing my ass off whilst trying not to piss myself',
'lmaoxh': 'laughing my ass off extremely hard',
'lmap': 'leave me alone please',
'lmb': 'lick my balls',
'lmbao': 'laughing my black ass off',
'lmbfwao': 'laughing my big fat white ass off',
'lmbo': 'laughing my butt off',
'lmcao': 'laughing my crazy ass off',
'lmclao': 'laughing my cute little ass off',
'lmd': 'lick my d**k',
'lmfao': 'laughing my f**king ass off',
'lmfbo': 'laugh my f**king butt off',
'lmffao': 'laughing my f**king fat ass off',
'lmffo': 'laughing my f**king face off',
'lmfho': 'laughing my f**king head off',
'lmfo': 'laughing my face off',
'lmfpo': 'laughing my f**king p***y off',
'lmfr': 'lets meet for real',
'lmfto': 'laughing my f**kin tits off',
'lmg': 'let me guess',
'lmgdao': 'laughing my god damn ass off',
'lmgtfy': 'let me google that for you',
'lmhao': 'laughing my hairy ass off',
'lmho': 'laughing my heiny off',
'lmip': 'lets meet in person',
'lmirl': "let's meet in real life",
'lmk': 'let me know',
'lmks': 'let me know soon',
'lmkwc': 'let me know when clear',
'lmkwut': 'let me know what you think',
'lml': 'love my life',
'lmmfao': 'laughing my mother f**king ass off',
'lmmfaos': 'laughing my mother f**king ass off silly',
'lmmfas': 'laugh my mother f**kin ass off',
'lmmffao': 'laughing my mother f**king fat ass off',
'lmo': 'leave me one',
'lmoao': 'laughing my other ass off',
'lmp': 'lick my p***y',
'lmpitw': 'let me put it this way',
'lmpo': 'laughing my panties off',
'lms': 'leave me some',
'lmsao': 'laughing my sexy ass off',
'lmso': 'laughing my socks off',
'lmtd': 'limited',
'lmtfa': 'leave me the f**k alone',
'lmto': 'laughing my tits off',
'lmtus': 'let me tell you something',
'lmty': 'laughing more than you',
'lmvo': 'laugh my vagina off',
'ln': 'last name',
'lnk': 'link',
'lobfl': 'laugh out bloody f**king loud',
'lobl': 'laugh out bloody loud',
'lof': 'laughing on floor',
'lofi': 'uncool',
'lofl': 'laugh out f**king loud',
'loflmao': 'laying on floor laughing my ass off',
'loi': 'laughing on the inside',
'lol': 'laughing out loud',
"lol'd": 'laughed out loud',
'lol2u': 'laugh out loud to you',
'lol@u': 'laugh out loud at you',
'lolarotf': 'laughing out loud and rolling on the floor',
'lolaw': 'laugh out loud at work',
'lolbs': 'laugh out loud but seriously',
'lolcano': 'laugh out loud',
'lolci': 'laughing out loud, crying inside',
'lolcity': 'the whole city laughs out loud',
'lold': 'laughed out loud',
'lolees': 'laugh out loud',
'lolerz': 'laugh out loud',
'lolf': 'lots of love forever',
'lolin': 'laughing out loud',
'lolio': 'laugh out loud i own',
'lollam': 'laughing out loud like a maniac',
'lollercaust': 'an extreme event of hilarity',
'lollercoaster': 'laugh out loud (a lot)',
'lollerskates': 'laughing out loud',
'lolm': 'laugh out loud man',
'loln': 'laught out loud... not',
'lolngs': 'laghing out loud never gonna stop',
'lolocost': 'laugh out loud',
'lolol': 'saying "lol" out loud',
'lololz': 'laugh out loud',
'lolpimp': 'laughing out loud peeing in my pants',
'lolq': 'laugh out loud quietly',
'lolrof': 'laughing out loud while rolling on the floor.',
'lolrotf': 'laughing out loud rolling on the floor',
'lols': 'laugh out loud',
'lolvq': 'laugh out loud very quietly',
'lolwtime': 'laughing out loud with tears in my eyes',
'lolz': 'laugh out loud',
'lomg': 'like oh my god',
'loml': 'love of my life',
'lomy': 'love of my life',
'loomm': 'laughing out of my mind',
'lorl': 'laugh out real loud',
'lorrl': 'laugh out really really loud',
'lotf': 'laughing on the floor',
'loti': 'laughing on the inside',
'loto': 'laughing on the outside',
'lotr': 'lord of the rings',
'lov': 'love',
'lovu': 'love you',
'loxen': 'laughing out loud',
'loxxen': 'laughing out loud',
'lozer': 'loser',
'lpb': 'low ping b*****d',
'lpiaw': 'large penis is always welcome',
'lpms': 'life pretty much sucks',
'lq': 'laughing quietly',
'lq2m': 'laughing quietly to myself',
'lqtm': 'laugh quietly to myself',
'lqtms': 'laughing quietly to myself',
'lqts': 'laughing quietly to self',
'lrfl': 'laughing really f**king loud',
'lrh': 'laughing really hard',
'lrqtms': 'laughing really quietly to myself',
'lrt': 'last retweet',
'lsfw': 'less safe for work',
'lshic': "laughing so hard i'm crying",
'lshid': 'laugh so hard i die',
'lshifabmh': 'laughing so hard i fell and broke my hip',
'lshipmp': 'laughing so hard i piss my pants',
'lshismp': 'laughed so hard i s**t my pants',
'lshiwms': 'laughing so hard i wet myself',
'lshmson': 'laughing so hard milk shot out nose',
'lshrn': 'laughing so hard right now',
'lsmih': 'laughing so much it hurts',
'lsr': 'loser',
'lsudi': 'lets see you do it',
'lt': 'long time',
'ltb': 'looking to buy',
'lthtt': 'laughing too hard to type',
'ltip': 'laughting until i puke',
'ltl': "let's talk later",
'ltm': 'listen to me',
'ltmq': 'laugh to myself quietly',
'ltms': 'laughing to my self',
'ltnc': 'long time no see',
'ltns': 'long time no see',
'ltnsoh': 'long time, no see or hear',
'ltnt': 'long time no talk',
'ltp': 'lay the pipe',
'ltr': 'later',
'lttpot': 'laughing to the point of tears',
'ltw': 'lead the way',
'ltywl': 'love the way you lie',
'lu2': 'love you too',
'lu2d': 'love you to death',
'lu4l': 'love you for life',
'lub': 'laugh under breath',
'luf': 'love',
'luff': 'love',
'lug': 'lesbian until graduation',
'luk': 'look',
'lukin': 'looking',
'lul': 'love you lots',
'lulab': 'love you like a brother',
'lulas': 'love you like a sister',
'lulz': 'laughing out loud.',
'lumumi': 'love you miss you mean it',
'lurker': "one who reads but doesn't reply",
'lurve': 'love',
'luser': 'user who is a loser',
'lusm': 'love you so much',
'luv': 'love',
'luver': 'lover',
'luvuvm': 'love you very much',
'luvv': 'love',
'luzar': 'loser',
'lv': 'love',
'lve': 'love',
'lvl': 'level',
'lvn': 'loving',
'lvr': 'lover',
'lvya': 'love you',
'lwih': 'look what i have',
'lwn': "last week's news",
'lwwy': "live while we're young",
'ly': 'love you',
'ly2': 'love you to',
'lya': 'love you always',
'lyaab': 'love you as a brother',
'lyaaf': 'love you as a friend',
'lyao': 'laugh your ass off',
'lybo': 'laugh your butt off',
'lyf': 'life',
'lyfao': 'laughing your f**king ass off',
'lyfe': 'life',
'lyk': 'like',
'lyk3': 'like',
'lyke': 'like',
'lyl': 'love you lots',
'lylab': 'love you like a brother',
'lylaba': 'love you like a brother always',
'lylad': 'love you like a dad',
'lylafklc': 'love you like a fat kid loves cake',
'lylal': 'love you lots and lots',
'lylam': 'love you like a mom',
'lylas': 'love you like a sister',
'lylasa': 'love you like a sister always',
'lylno': 'love you like no other',
'lyls': 'love you lots',
'lymi': 'love you mean it',
'lymywy': 'love you, miss you, want you',
'lysfm': 'love you so f**king much',
'lysm': 'love you so much',
'lyt': 'love you too',
'lyvm': 'love you very much',
'lzer': 'laser',
'lzr': 'loser'}
# -
m = {'m': 'am',
'm$': 'microsoft',
'm$wxp': 'microsoft windows xp',
'm&d': 'mom and dad',
"m'kay": 'okay',
'm.i.a': 'missing in action',
'm.o': 'makeout',
'm/b': 'maybe',
'm/f': 'male or female',
'm2': 'me too',
'm3': 'me',
'm473s': 'friends',
'm473z': 'friends',
'm4f': 'male for female',
'm4m': 'male for male',
'm8': 'friend',
'm84l': 'mate for life',
'm8s': 'mates',
'm8t': 'mate',
"m8t's": 'friends',
'm9': 'mine',
"ma'am": 'madam',
'mabby': 'maybe',
'mabe': 'maybe',
'maga': 'make america great again',
'mah': 'my',
'mai': 'my',
'mama': 'mother',
'mao': 'my ass off',
'marvy': 'marvelous',
'masterb8': 'masterbate',
'mastrb8': 'masturbate.',
'mayb': 'maybe',
"mayn't": 'may not',
'mayte': 'mate',
'mb': 'my bad',
'mbf': 'my best friend',
'mbfal': 'my best friend and lover',
'mbhsm': 'my boobs hurt so much',
'mbl8r': 'maybe later',
'mc': 'master of ceremonies',
'mcds': 'mcdonalds',
'mcm': 'man crush monday',
'mcs': 'my computer sucks',
'mcse': 'microsoft certified systems engineer',
'mdf': 'my dear friend',
'mdk': 'murder death kill',
'me2': 'me too',
'meatcurtain': "woman's private parts",
'meatspace': 'the real world',
'meeh': 'me',
'mego': 'my eyes glaze over',
'meh': 'whatever',
'messg': 'message',
'mf': 'motherf**ker',
'mf2f4sx': 'meet face to face for sex',
'mfa': 'mother f**king a**h**e',
'mfah': 'motherf**king a**h**e',
'mfao': 'my f**king ass off',
'mfb': 'mother f**king b***h',
'mfer': 'motherf**ker',
'mfg': 'merge from current',
'mfkr': 'motherf**ker',
'mflfs': 'married female looking for sex',
'mfr': 'motherf**ker',
'mfw': 'my face when',
'mgiwjsdchmw': "my girlfriend is watching jeff so don't call her my wife",
'mgmt': 'management',
'mhh': 'my head hurts',
'mhm': 'yes',
'mho': 'my humble opinion',
'mia': 'missing in action',
'mic': 'microphone',
"might've": 'might have',
"mightn't": 'might not',
"mightn't've": 'might not have',
'miid': 'my internet is down',
'milf': "mom i'd like to f**k",
'miltf': "mom i'd like to f**k",
'min': 'minute',
'mins': 'minutes',
'miq': 'make it quick',
'mir': 'mom in room',
'mirl': 'meet in real life',
'misc.': 'miscellaneous',
'miself': 'myself',
'missin': 'missing',
'mite': 'might',
'miw': 'mom is watching',
'miwnlf': 'mom i would not like to f**k.',
'mk': 'mmm....ok',
'mkay': 'ok',
'mlc': 'mid life crisis',
'mle': 'emily',
'mlg': 'major league gaming',
'mlia': 'my life is amazing',
'mlm': 'multi level marketer',
'mlod': 'mega laugh out loud of doom',
'mlp': 'my little pony',
'mmamp': 'meet me at my place',
'mmas': 'meet me after school',
'mmatc': 'meet me around the corner',
'mmatp': 'meet me at the park',
'mmbocmb': 'message me back or comment me back',
'mmd': 'make my day',
'mmiw': 'my mom is watching',
'mmk': 'umm, ok',
'mml': 'making me laugh',
'mml8r': 'meet me later',
'mmlfs': 'married man looking for sex',
'mmmkay': 'okay',
'mmo': 'massive multiplayer online',
'mmorpg': 'massively multiplayer online role playing game',
'mmt': 'meet me there',
'mmtyh': "my mom thinks you're hot",
'mmw': 'making me wet',
'mngmt': 'management',
'mngr': 'manager',
'mnm': 'eminem',
'mnt': 'more next time',
'mobo': 'motherboard',
'mof': 'matter of fact',
'mofo': 'mother f**ker',
'moh': 'medal of honor',
'mohaa': 'medal of honor allied assult',
'mol': 'more or less',
'mommies': 'mother',
'mommy': 'mother',
'mompl': 'moment please',
'moms': 'mother is',
'moobs': 'man boobs',
'mor': 'more',
'morf': 'male or female',
'moro': 'tomorrow',
'mos': 'mom over shoulder',
'moss': 'member of same sex',
'motarded': 'more retarded',
'motd': 'message of the day',
'motos': 'member of the opposite sex',
'mpaw': 'my parents are watching',
'mpbis': 'most popular boy in school',
'mpd': 'multiple personality disorder',
'mpgis': 'most popular girl in school',
'mph': 'miles per hour',
'mpih': 'my penis is hard',
'mpty': 'more power to you',
'mrau': 'message received and understood',
'msf': 'male seeking female',
'msg': 'message',
'msgs': 'messages',
'msh': 'me so horny',
'msibo': 'my side is busting open',
'msie': "microsoft's internet explorer",
'msm': 'main stream media',
'msmd': 'monkey see - monkey do',
'msngr': 'messenger',
'mssg': 'message',
'mstrb8r': 'masturbator',
'msv': 'microsoft vista',
'mtc': 'more to come',
'mte': 'my thoughts exactly',
'mtf': 'more to follow',
'mtfbwu': 'may the force be with you',
'mtfbwy': 'may the force be with you',
'mtg': 'meeting',
'mtherfker': 'mother f**ker',
'mthrfkr': 'mother f**ker',
'mtl': 'more than likely',
'mtr': 'matter',
'mtrfkr': 'motherf**ker',
'mty': 'empty',
'mu': 'miss you',
'mudda': 'mother',
'muh': 'my',
'mul': 'miss you lots',
'mum': 'mother',
"mum's": 'mother is',
'mums': 'mother is',
'musiq': 'music',
'musm': 'miss you so much',
"must've": 'must have',
"mustn't": 'must not',
"mustn't've": 'must not have',
'mutha': 'mother',
'muve': 'multi-user virtual environment',
'muvva': 'mother',
'muzik': 'music',
'mvp': 'most valuble player',
'mw2': 'modern warfare 2',
'mw3': 'modern warfare 3',
'mwah': 'kiss',
'mwf': 'married white female',
'mwm': 'married white man',
'mwsmirl': 'maybe we should meet in real life',
'myaly': 'miss you and love you',
'myers': 'many years',
'myfb': 'mind your f**king business',
'myke': 'man-dyke',
'myn': 'mine',
'myob': 'mind your own business',
'myodb': 'mind your own damn business',
'myofb': 'mind your own f**king business',
'mypl': 'my young padawan learner',
'mysm': 'miss you so much',
'myspce': 'myspace'}
n = {'n': 'and',
'n e': 'any',
'n/a': 'not applicable',
'n/a/s/l': 'name, age, sex location',
'n/c': 'no comment',
'n/m': 'nevermind',
'n/n': 'nickname',
'n/o': 'no offense',
'n/r': 'no replys',
'n/t': 'no text',
'n00b': 'newbie',
'n00bs': 'newbies',
'n00dz': 'nudes',
'n00s': 'news',
'n1': 'nice one',
'n199312': 'african american',
'n1994': 'african american',
'n2': 'into',
'n2b': 'not too bad',
'n2bb': 'nice to be back',
'n2bm': 'not to be mean',
'n2br': 'not to be rude',
'n2g': 'not too good',
'n2m': 'not too much',
'n2mh': 'not too much here',
'n2mhbu': 'not too much how about you?',
'n2mhjc': 'not too much here just chillin',
'n2mu': 'not too much, you?',
'n2n': 'need to know',
'n2p': 'need to pee',
'n64': 'nintendo 64',
'n8v': 'native',
'na': 'not applicable',
'na4w': 'not appropriate for work',
'naa': 'not at all',
'nade': 'grenade',
'nafc': 'not appropriate for children',
'nafkam': 'not away from keyboard any more',
'naft': 'not a f**king thing',
'nafta': 'north american free trade agreement',
'nah': 'no',
'namh': 'not at my house',
'nao': 'not as often',
'natch': 'naturally',
'natm': 'not at the minute',
'naw': 'no',
'naw-t': 'naughty',
'nawidt': 'never again will i do that',
'nawt': 'not',
'naww': 'no',
'nayl': 'in a while',
'nb': 'not bad',
'nb,p': 'nothing bad, parents',
'nba': 'national basketball association',
'nbd': 'no big deal',
'nbdy': 'nobody',
'nbf': 'never been f**ked',
'nc': 'not cool',
'ncaa': 'national collegiate athletic association',
'ncs': 'no crap sherlock',
'nd': 'and',
'ndit': 'no details in thread',
'ndn': 'indian',
'nds': 'nintendo ds',
'ne': 'any',
'ne1': 'anyone',
'neday': 'any day',
'nedn': 'any day now',
"needn't": 'need not',
"needn't've": 'need not have',
'nefing': 'anything',
'negl': 'not even going to lie',
'nei': 'not enough information',
'neida': 'any idea',
'nekkid': 'naked',
'nemore': 'anymore',
'nes': 'nintendo entertainment system',
'nethin': 'anything',
'nething': 'anything',
'neva': 'never',
'nevah': 'never',
'nevar': 'never',
'nevarz': 'never',
'nevm': 'never mind',
'nevr': 'never',
'newais': 'anyways',
'neway': 'anyway',
'neways': 'anyways',
'newayz': 'anyways',
'newb': 'someone who is new',
'newbie': 'new player',
'newez': 'anyways',
'nf': 'not funny',
'nfbsk': 'not for british school kids',
'nfc': 'no f**king clue',
'nfd': 'no f**king deal',
'nff': 'not f**king fair',
'nfg': 'no f**king good',
'nfi': 'no f**king idea',
'nfr': 'not for real',
'nfs': 'not for sale',
'nft': 'no further text',
'nfw': 'no f**king way',
'ng': 'nice game',
'ngaf': 'nobody gives a f**k',
'ngl': 'not gonna lie',
'nh': 'nice hand',
'nhatm': 'not here at the moment',
'ni': 'no idea',
'ni994': 'n***a',
'nib': 'new in box',
'nic': 'network interface card',
'nif': 'non internet friend',
'nifoc': 'naked in front of computer',
'nifok': 'naked in front of keyboard',
'nigysob': "now i've got you son of a b***h",
'nimby': 'not in my backyard',
'nin': 'no its not',
'nip': 'nothing in particular',
'nips': 'nipples',
'nite': 'night',
'nitfm': 'not in the f**king mood',
'nitm': 'not in the mood',
'niwdi': "no i won't do it",
'nizzle': 'n****r',
'nj': 'nice job',
'njoy': 'enjoy',
'njp': 'nice job partner',
'nk': 'no kidding',
'nkd': 'naked',
'nkt': 'never knew that',
'nld': 'nice lay down',
'nm': 'not much',
'nm u': 'not much, you',
'nmb': 'not my business',
'nmbr': 'number',
'nme': 'enemy',
'nmf': 'not my fault',
'nmfp': 'not my f**king problem',
'nmh': 'not much here',
'nmhau': 'nothing much how about you',
'nmhbu': 'nothing much how about you',
'nmhm': 'nothing much here, man',
'nmhu': 'nothing much here, you?',
'nmhwby': 'nothing much here what about you',
'nmjb': 'nothing much just bored',
'nmjc': "not much, just chillin'",
'nmjch': 'nothing much just chilling',
'nmjcu': 'nothing much, just chilling, you?',
'nmjdhw': 'nothing much just doing homework',
'nmjfa': 'nothing much, just f**king around',
'nmnhnlm': 'no money, no honey, nobody loves me',
'nmp': 'not my problem',
'nmu': 'nothing much, you',
'nmw': 'no matter what',
'nmwh': 'no matter what happens',
'nn': 'good night',
'nn2r': 'no need to respond',
'nnaa': 'no not at all',
'nnfaa': 'no need for an apology',
'nnr': 'no not really',
'nntr': 'no need to reply',
'nntst': 'no need to say thanks',
'no pro': 'no problem',
'no1': 'no one',
'noaa': 'national oceanic and atmospheric administration',
'noc': 'naked on camera',
'noe': 'know',
'noes': 'no',
'nofi': 'no flame intended',
'nolm': 'no one loves me',
'nomw': 'not on my watch',
'noob': 'someone who is new',
'noobie': 'new person',
'nooblet': 'new player',
'noobz0r': 'newbie',
'noodz': 'nude pictures',
'nookie': 'sex',
'nop': 'normal operating procedure',
'nope': 'no',
'norc': 'no one really cares',
'norwich': 'knickers off ready when i come home',
'nos': 'numbers',
'notin': 'nothing',
'noty': 'no thank you',
'noub': 'none of your business',
'nowai': 'no way',
'nowin': 'knowing',
'noyb': 'none of your business',
'noygdb': 'none of your god damn business',
'np': 'no problem',
'np4np': 'naked pic for naked pic',
'npa': 'not paying attention',
'npc': 'non-playable character',
'npe': 'nope',
'npgf': 'no problem girl friend',
'nph': 'no problem here',
'npi': 'no pun intended',
'npnt': 'no picture, no talk',
'nps': 'no problems',
'nq': 'thank you',
'nqa': 'no questions asked',
'nr': 'no reserve',
'nr4u': 'not right for you',
'nrg': 'energy',
'nrn': 'no response necessary',
'ns': 'nice',
'nsa': 'no strings attached',
'nsas': 'no strings attached sex',
'nsfmf': 'not so fast my friend',
'nsfu': 'no sex for you',
'nsfw': 'not safe for work',
'nss': 'no s**t sherlock',
'nst': 'no school today',
'nstaafl': 'no such thing as a free lunch',
'nt': 'nice try',
'ntb': 'not to bad',
'ntbn': 'no text-back needed',
'nthg': 'nothing',
'nthin': 'nothing',
'nthn': 'nothing',
'ntigaf': 'not that i give a f**k',
'ntk': 'need to know',
'ntkb': 'need to know basis',
'ntm': 'not to much',
'ntmk': 'not to my knowledge',
'ntmu': 'nice to meet you',
'ntmy': 'nice to meet you',
'ntn': 'nothing',
'ntrly': 'not really',
'nts': 'note to self',
'ntstt': 'not safe to talk',
'ntt': 'need to talk',
'ntta': 'nothing to talk about',
'nttawwt': 'not that there is anything wrong with that',
'nttiawwt': 'not that there is anything wrong with that.',
'ntty': 'nice talking to you',
'ntw': 'not to worry',
'ntxt': 'no text',
'nty': 'no thank you',
'nu': 'new',
'nub': 'inexperienced person',
'nuff': 'enough',
'nuffin': 'nothing',
'nufin': 'nothing',
'nutin': 'nothing',
'nuttin': 'nothing',
'nv': 'envy',
'nvm': 'never mind',
'nvmd': 'nevermind',
'nvmdt': 'never mind then',
'nvmt': 'nevermind that',
'nvr': 'never',
'nvrm': 'nevermind',
'nvrmnd': 'never mind',
'nw': 'no way',
'nwb': 'a new person',
'nwih': 'no way in hell',
'nwj': 'no way jose',
'nwo': 'new world order',
'nwrus': 'no way are your serious',
'nws': 'not work safe',
'nwtf': 'now what the f**k',
'nwy': 'no way',
'nxt': 'next',
'ny1': 'anyone',
'nyc': 'new york city',
'nyf': 'not your fault',
'nyp': 'not your problem',
'nywy': 'anyway'}
o = {'o': 'oh',
'o rly': 'oh really',
'o&o': 'over and out',
"o'": 'of',
"o'clock": 'of the clock',
'o.p.': 'original poster',
'o/y': 'oh yeah',
'o4b': 'open for business',
'oaoa': 'over and over again',
'oar': 'on a roll',
'oaw': 'on a website',
'obgjfioyo': 'old but good job finding it on your own',
'obj': 'object',
'obl': 'osama bin laden',
'obo': 'or best offer',
'obtw': 'oh, by the way',
'obv': 'obviously',
'obvi': 'obviously',
'oc': 'original character',
'occ': 'occupation',
'ocd': 'obsessive compulsive disorder',
'ocgg': 'oh crap, gotta go',
'od': 'over dose',
'oday': 'software illegally obtained before it was released',
'odg': 'oh dear god',
'odtaa': 'one damn thing after another',
'oe': 'or else',
'oed': 'oxford english dictionary',
'of10': 'often',
'ofc': 'of course',
'ofcol': 'oh for crying out loud',
'ofn': 'old f**king news',
'oftc': 'out for the count',
'oftn': 'often',
'oftpc': 'off topic',
'ofwg': 'old fat white guys',
'og': 'original gangster',
'ogw': 'oh guess what',
'oh noes': 'oh s**t!',
'oh noez': 'oh no!',
'ohic': 'oh i see',
'ohn': 'oh hell no',
'ohnoez': 'oh no',
'ohy': 'oh hell yeah',
'oibmpc': 'oops i broke my computer',
'oic': 'oh, i see',
'oicic': 'oh i see i see',
'oicu': 'oh, i see you!',
'oicwydt': 'oh, i see what you did there',
'oidia': 'oops i did it again',
'oink': 'oh i never knew',
'oiyd': 'only in your dreams',
'oj': 'orange juice',
'ojsu': 'oh, just shut up!',
'ok': 'ok',
'okey': 'ok',
'oll': 'online love',
'olpc': 'one laptop per child',
'omdg': 'oh my dear god',
'omdz': 'oh my days',
'omfd': 'oh my f**king days',
'omfg': 'oh my f**king god',
'omfgn': 'oh my f**king god noob',
'omfgsh': 'oh my f**king gosh',
'omfj': 'oh my f**king jesus',
'omfl': 'oh my f**king internet connection is slow',
'omfsm': 'oh my flying spaghetti monster',
'omfwtg': 'oh my f**k what the god?',
'omg': 'oh my god',
"omg's": "oh my god's",
'omgd': 'oh my gosh dude',
'omgf': 'oh my god...f**k!',
'omgg': 'oh my gosh girl',
'omgicfbi': "oh my god i can't f**king believe it",
'omgih': 'oh my god in heaven',
'omgihv2p': 'oh my god i have to pee',
'omginbd': "oh my god, it's no big deal",
'omgn': 'oh my goodness',
'omgny': 'oh my god no way',
'omgosh': 'oh my gosh',
'omgroflmao': 'oh my god roll on the floor laughing my ass off',
'omgsh': 'oh my gosh',
'omgty': 'oh my god thank you',
'omgukk': 'oh my god you killed kenny',
'omgwtf': 'on my god, what the f**k',
'omgwtfbbq': 'oh my god, what the f**k',
'omgwtfhax': 'oh my god what the f**k, hacks!',
'omgwtfit': 'oh my god, what the f**k is that',
'omgwtfnipples': 'on my god, what the f**k',
'omgyg2bk': 'oh my god you got to be kidding',
'omgykkyb': 'oh my god you killed kenny you b*****ds',
'omgz': 'oh my god',
'omgzors': 'oh my god',
'omhg': 'oh my hell god',
'omj': 'oh my jesus',
'oml': 'oh my lord',
'ommfg': 'oh my mother f**king god',
'omt': 'one more time',
'omw': 'on my way',
'omwh': 'on my way home',
'omwts': 'on my way to school',
'omy': 'oh my!',
'onoez': 'oh no',
'onoz': 'oh no',
'onud': "oh no you didn't",
'onyd': "oh no you didn't",
'oob': 'out of buisness',
'oobl': 'out of breath laughing',
'ooc': 'out of character',
'oohm': 'out of his/her mind',
'oom': 'out of mana',
'oomf': 'one of my followers',
'oomm': 'out of my mind',
'ootb': 'out of the blue',
'ootd': 'outfit of the day',
'oow': 'on our way',
'ooym': 'out of your mind',
'op': 'operator',
'orgy': 'orgasm',
'orlsx': 'oral sex',
'orly': 'oh really?',
'orpg': 'online role playing game',
'os': 'operating system',
'osbutctt': 'only sad b*****ds use this crappy text talk',
'osd': 'on screen display',
'osifgt': 'oh s**t i forgot',
'oslt': 'or something like that',
'osy': 'oh screw you',
'ot': 'off topic',
'otb': 'off the boat',
'otc': 'off the chain',
'otfcu': 'on the floor cracking up',
'otfl': 'on the floor laughing',
'otflmao': 'on the floor laughing my ass off',
'otflmfao': 'on the floor laughing my f**king ass off',
'otflol': 'on the floor laughing out loud',
'otfp': 'on the f**king phone',
'otft': 'over the f**king top',
'oti': 'on the internet',
'otl': 'out to lunch',
'otoh': 'on the other hand',
'otp': 'on the phone',
'ots': 'over the shoulder',
'ott': 'over the top',
'otw': 'on the way',
"oughtn't": 'ought not',
"oughtn't've": 'ought not have',
'outa': 'out of',
'ova': 'over',
'oways': 'oh wow are you serious',
'owned': 'made to look bad',
'ownt': 'made to look bad',
'ownz': 'owns',
'ownzer': 'one who makes others look bad',
'ownzorz': 'owned.',
'owt': 'out',
'oww': 'oops, wrong window',
'oyfe': 'open your f**king eyes',
'oyid': 'oh yes i did',
'oyo': 'on your own',
'oyr': 'oh yeah right'}
p = {'p-nis': 'penis',
'p.o.b.': 'parent over back',
'p.o.s': 'parent over shoulder',
'p.o.s.': 'parent over shoulder',
'p/oed': 'pissed off',
'p/w': 'password',
'p00p': 'poop',
'p0wn': 'make to look bad',
'p2p': 'peer to peer',
'p2w': 'pay to win',
'p33n': 'penis',
'p3n0r': 'penis',
'p3n15': 'penis',
'p3n1s': 'penis',
'p4p': 'pic for pic',
'p911': 'parent emergency (parent near)',
'p@w': 'parents are watching',
'p^s': 'parent over shoulder',
'pach': 'parents are coming home',
'pachs': 'parents are coming home soon',
'pae': 'pimpin aint easy',
'pag': 'parents are gone',
'pah': 'parents at home',
'panl': 'party all night long',
'parnts': 'parents',
'pas': 'parent at side',
'pasii': 'put a sock in it',
'patd': 'panic at the disco',
'paw': 'parents are watching',
'pb': 'peanut butter',
'pb&j': 'peanut butter and jelly',
'pbb': 'parent behind back',
'pbcakb': 'problem between chair and keyboard',
'pbj': 'peanut butter and jelly',
'pbjt': 'peanut butter jelly time',
'pbkc': 'problem between keyboard & chair',
'pbly': 'probably',
'pbm': 'parent behind me',
'pbp': 'please be patient',
'pcbd': 'page cannot be displayed',
'pce': 'peace',
'pcent': 'percent',
'pcm': 'please call me',
'pco': 'please come over',
'pcrs': 'parents can read slang',
'pda': 'public display of affection',
'pdg': 'pretty damn good',
'pdq': 'pretty damn quick',
'peanus': 'penis',
'pearoast': 'repost',
'pebcak': 'problem exists between chair and keyboard',
'pebkac': 'problem exists between keyboard and chair',
'pebmac': 'problem exist between monitor and chair',
'peep dis': "check out what i'm telling you",
'peeps': 'people',
'pen0r': 'penis',
'pen15': 'penis',
'penor': 'penis',
'peoples': 'people',
'perv': 'pervert',
'pewp': 'poop',
'pex': 'please explain?',
'pezzas': 'parents',
'pf': 'profile',
'pfa': 'please find attached',
'pfm': 'please forgive me',
'pfo': 'please f**k off',
'pfos': 'parental figure over sholder',
'pfy': 'pimply faced youth',
'pg': 'page',
'ph#': 'phone number',
'ph33r': 'fear',
'ph34r': 'fear',
'phag': 'f**',
'phail': 'fail',
'phat': 'pretty hot and tasty',
'phayl': 'fail',
'phear': 'fear',
'phlr': 'peace hugs love respect',
'phm': 'please help me',
'phq': 'f**k you',
'phreak': 'freak',
'phreaker': 'phone hacker',
'phuck': 'f**k',
'phucker': 'f**ker',
'phuk': 'f**k',
'phun': 'fun',
'phux': 'f**k',
'phuxor': 'f**k',
'piab': 'panties in a bunch',
'pic': 'picture',
'piccies': 'pictures',
'pics': 'pictures',
'pihb': 'pee in his/her butt',
'piihb': 'put it in her butt',
'piitb': 'put it in the butt',
'pima': 'pain in my ass',
'pimfa': 'pain in my f**king ass',
'pimha': 'pain in my hairy ass',
'pimpl': 'pissing in my pants laughing',
'pino': 'filipino',
'pir': 'parents in room',
'pirlos': 'parent in room looking over shoulder',
'pita': 'pain in the ass',
'pitfa': 'pain in the f**king ass',
'pitr': 'parent in the room',
'pitrtul': 'parents in the room text you later',
'piw': 'parent is watching',
'pix': 'pictures',
'pk': 'player kill',
'pkemon': 'pokemon',
'pker': 'player killer',
'pking': 'player killing',
'pl': 'parent looking',
'pl0x': 'please',
'pl8': 'plate',
'plac': 'parent looking at computer',
'plams': 'parents looking at my screen',
'plars': 'party like a rock star',
'platcs': 'parent looking at the computer screen',
"ple's": 'please',
'pleaz': 'please',
'pleez': 'please',
'pleeze': 'please',
'pleze': 'please',
'pliz': 'please',
'plma': 'please leave me alone',
'plmk': 'please let me know',
'plocks': 'please',
'plom': 'parents looking over me',
'plomb': 'parents looking over my back',
'ploms': 'parent looking over my shoulder',
'plos': 'parents looking over shoulder',
'plox': 'please',
'ploxxorz': 'please',
'pls': 'please',
'plse': 'please',
'plx': 'please/thanks',
'plywm': 'play with me',
'plz': 'please',
'plzkthx': 'please? ok, thank you',
'plzthx': 'please? thanks',
'pmfji': 'pardon me for jumping in',
'pmfsl': 'piss my f**king self laughing',
'pmg': 'oh my god',
'pmita': 'pound me in the ass',
'pmitap': 'pound me in the ass prison',
'pml': 'pissing myself laughing',
'pmo': 'pissing me off',
'pmp': 'pissing my pants',
'pmpl': 'piss my pants laughing',
'pmsfl': 'pissed myself f**king laughing',
'pmsl': 'piss my self laughing',
'pmt': 'pretty much this',
'pnbf': 'potential new boy friend',
'pnhlgd': "parents not home, let's get dirty",
'pns': 'penis',
'pnus': 'penis',
'po': 'piss off',
'po po': 'police',
"po'd": 'pissed off',
'pob': 'parent over back',
'poc': 'piece of crap',
'poed': 'pissed off',
'poets': "piss off early, tomorrow's saturday",
'poi': 'point of interest',
'poidnh': 'pics or it did not happen',
'pol': 'parent over looking',
'poms': 'parent over my shoulder',
'poo': 'poop',
'poontang': 'female genitalia',
'pooter': 'computer',
'popo': 'police',
'poq': 'piss off quick',
'pos': 'parent over shoulder',
'poscs': 'parents over sholder change subject',
'posmbri': 'parent over shoulder might be reading it',
'potc': 'pirates of the caribbean',
'pots': 'plain old telephone service',
'pov': 'point of view',
'pow': 'prisoner of war',
'pp': 'pee pee',
'ppl': 'people',
'ppls': 'people',
'pplz': 'people',
'ppor': 'post proof or recant',
'ppppppp': 'prior proper planning prevents piss poor performance',
'pr0': 'professional',
'pr0n': 'porn',
'pr0nz': 'porn',
'prblm': 'problem',
'prd': 'period',
'preggers': 'pregnant',
'prego': 'pregnant',
'prfct': 'perfect',
'prn': 'porn',
'prncpl': 'principal',
'prncss': 'princess',
'prnoscrn': 'porn on screen',
'pro': 'professional',
'prob': 'probably',
'probly': 'probably',
'probz': 'probably',
'prod': 'product',
'prolly': 'probably',
'prollz': 'probably',
'proly': 'probably',
'promos': 'promotions',
'pron': 'porn',
'proxie': 'proxy',
'prp': 'please reply',
'prsn': 'person',
'prt': 'party',
'prty': 'party',
'prv': 'private',
'prvrt': 'pervert',
'prw': 'parents are watching',
'ps1': 'play station 1',
'ps2': 'play station 2',
'ps3': 'play station 3',
'psa': 'public service announcement',
'psbms': 'parent standing by my side',
'psl': 'pumpkin spice latte',
'psn': 'playstation netwok',
'psos': 'parent standing over sholder',
'psp': 'playstation portable',
'pssy': 'p***y',
'pst': 'please send tell',
'pt33n': 'preteen',
'ptbb': 'pass the barf bag',
'ptfo': 'passed the f**k out',
'pthc': 'preteen hardcore',
'ptl': 'praise the lord',
'pto': 'personal time off',
'ptw': 'play to win',
'puh-leaze': 'please',
'purty': 'pretty',
'puter': 'computer',
'pvp': 'player versus player',
'pvt': 'pervert',
'pw': 'parent watching',
'pwb': 'p***y whipped b***h',
'pwcb': 'parents watching close by',
'pwd': 'password',
'pwn': 'made to look bad',
'pwn3d': 'owned',
'pwn3r': 'owner',
'pwnage': 'ownage',
'pwnd': 'owned',
'pwned': 'made to look bad',
'pwner': 'owner',
'pwnr': 'owner',
'pwnt': 'owned',
'pwnz': 'owns',
'pwnzor': 'owner',
'pwob': 'parent watching over back',
'pwoms': 'parent watching over my shoulder',
'pwor': 'power',
'pwos': 'parent was over sholder',
'pww': 'parents were watching',
'pxr': 'punk rocker',
'pydim': 'put your d**k in me',
'pyfco': 'put your freaking clothes on',
'pyt': 'pretty young thing',
'pz': 'peace',
'pzled': 'puzzled'}
q = {'q2c': 'quick to c**',
'q33r': 'queer',
'q4u': 'question for you',
'qed': "i've made my point",
'qfe': 'quoted for emphasis',
'qfmft': 'quoted for motherf**king truth',
'qft': 'quoted for truth',
'qft&gj': 'quoted for truth and great justice',
'ql': 'cool',
'qltm': 'quietly laughing to myself',
'qna': 'question and answer',
'qool': 'cool',
'qoolz': 'cool',
'qotd': 'quote of the day',
'qotsa': 'queens of the stone age',
'qoty': 'quote of the year',
'qpr': 'quite pathetic really',
'qpwd': 'quit posting while drunk',
'qq': 'crying eyes',
'qt': 'cutie',
'qt3.14': 'cutie pie',
'qte': 'cutie',
'qtpi': 'cutie pie'}
r = { 'r': 'are',
'r-tard': 'retard',
'r.i.p': 'rest in peace',
'r.i.p.': 'rest in peace',
'r0x0rz': 'rocks',
'r2f': 'ready to f**k',
'r8': 'rate',
'r8p': 'rape',
'r8pist': 'rapist',
'r8t': 'rate',
'ra2': 'red alert 2 (game)',
'ra3': 'red alert 3 (game)',
'raoflmao': 'rolling around on floor laughing my ass off',
'rawk': 'rock',
'rawks': 'rocks',
'rawr': 'roar',
'rb@u': 'right back at you',
'rbau': 'right back at you',
'rbay': 'right back at you',
'rbm': 'right behind me',
'rbtl': 'read between the lines',
'rbty': 'right back to you',
'rcks': 'rocks',
'rcsa': 'right click save as',
'rcvd': 'received',
'rdy': 'ready',
're': 'reply',
're/rehi': 'hello again',
'realy': 'really',
'reefer': 'marijuana',
'refl': 'rolling on the floor laughing',
'refusn': 'refusing',
'rehi': 'hello again',
'rele': 'really',
'rents': 'parents',
'rentz': 'parents',
'rep': 'to represent',
'reppin': 'representing',
'retrotextual': 'one who is using out of date words and abbreviations while texting.',
'rff': 'really f**king funny',
'rflmao': 'rolling on the floor laughing my ass off',
'rfn': 'right f**king now',
'rgr': 'roger',
'rhcp': 'red hot chilli peppers',
'rhgir': 'really hot guy in room',
'rhs': 'right hand side',
'ricl': 'rolling in chair laughing',
'rifk': 'rolling on the floor laughing',
'rihad': 'rot in hell and die',
'rino': 'republican in name only',
'rite': 'right',
'ritjive': 'non virgin',
'rjct': 'reject',
'rl': 'real life',
'rlbf': 'real life boy friend',
'rlf': 'real life friend',
'rlg': 'really loud giggle',
'rlgf': 'real life girl friend',
'rlly': 'really',
'rln': 'real life name',
'rly': 'really',
'rlz': 'rules',
'rlze': 'realize',
'rm': 'room',
'rmao': 'rolling my ass off',
'rme': 'rolling my eyes',
'rmr': 'remember',
'rmso': 'rock my socks off',
'rn': 'right now',
'rnt': "aren't",
'ro': 'rock out',
'rockr': 'rocker',
'rodger': 'affirmative',
'rofalol': 'roll on the floor and laugh out loud',
'rofc': 'rolling on floor crying',
'roffle': 'rolling on the floor laughing',
'roffle out loud': 'rolling on the floor laughing out loud',
'rofflecake': 'rolling on the floor laughing',
'rofflecopters': 'rolling on the floor with laughter',
'roffleol': 'rolling on the floor laughing out loud',
'roffles': 'rolling on floor laughing',
'rofflmfao': 'rolling on the floor laughing my f**king ass',
'rofl': 'rolling on the floor laughing',
'rofl&pmp': 'rolling on floor laughing and peeing my pants',
'roflao': 'rolling on the floor laughing my ass off',
'roflastc': 'rolling on floor laughing and scaring the cat',
'roflcopter': 'rolling on the floor laughing',
'roflcopters': 'rolling on the floor laughing, very funny.',
'roflkmd': 'rolling on the floor laughing kicking my dog',
'rofllh': 'rolling on the floor laughing like hell',
'roflmao': 'rolling on the floor laughing my ass off',
'roflmaoapimp': 'rolling on the floor laughing my ass off and peeing in my pants',
'roflmaool': 'rolling on the floor laughing my ass off out loud',
'roflmaopmp': 'rolling on the floor, laughing my ass off, pissing my pants',
'roflmaouts': 'rolling on floor laughing my f**king ass off unable to speak',
'roflmaowpimp': 'rolling on floor laughing my ass off while peeing in my pants',
'roflmbfao': 'rolling on floor laughing my big fat ass off',
'roflmbo': 'rolling on floor laughing my butt off',
'roflmfao': 'rolling on the floor laughing my f**king ass off',
'roflmfaopimp': 'rolling on the floor laughing my f**king ass off pissing in my pants',
'roflmfaopmp': 'rolling on flor laughing my f**king ass of peeing my pants',
'roflmgao': 'rolling on the floor laughing my gay ass off',
'roflmgdao': 'rolling on the floor laughing my god damn ass off',
'roflmgdmfao': 'roling on floor laughing my god damn mother f**king ass off',
'roflmgo': 'rolling on floor laughing my guts out',
'roflmho': 'rolling on the floor laughing my head off',
'roflmiaha': 'rolling on the floor laughing myself into a heart attack',
'roflmmfao': 'rolling on the floor laughing my mother f**king ass off',
'roflol': 'rolling on floor laughing out loud',
'roflolbag': 'rolling on the floor laughing out loud busting a gut',
'roflpimp': 'rolling on the floor laughing pissing in my pants',
'roflpmp': 'rolling on the floor laughing peeing my pants',
'roflwtime': 'rolling on the floor laughing with tears in my eyes',
'rofpml': 'rolling on the floor pissing myself laughing',
'rofwl': 'rolling on the floor while laughing',
'roger': 'affirmative',
'rogl': 'rolling on ground laughing',
'roglmfao': 'rolling on ground laughing my f**king ass off',
'roi': 'return on investment',
'roids': 'steroids',
'roj': 'affirmative',
'rol': 'rolling over laugihng',
'rolmao': 'rolling over laughing my ass off',
'rolmfao': 'rolling over laughing my f**king ass off',
'romalsh': 'rolling on my ass laughing so hard',
'rombl': 'rolled off my bed laughing',
'rong': 'wrong',
'roofles': 'rolling on the floor laughing',
'ror': 'raughing out roud',
'rotf': 'rolling on the floor',
'rotfalol': 'roll on the floor and laugh out loud',
'rotffl': 'roll on the f**king floor laughing',
'rotfflmao': 'rolling on the f**king floor laughing my ass off',
'rotfflmfao': 'rolling on the f**king floor laughing my f**king ass off',
'rotfl': 'rolling on the floor laughing',
'rotflaviab': 'rolling on the floor laughing and vomiting in a bucket',
'rotflmao': 'rolling on the floor laughing my ass off',
'rotflmaofaktd': 'rolling on the floor laughing my ass off farted and killed the dog',
'rotflmaool': 'rolling on the floor laughing my ass off out loud',
'rotflmaostc': 'rolling on the floor laughing my ass off scaring the cat',
'rotflmbo': 'rolling on the floor laughing my butt off',
'rotflmfao': 'rolling on the floor laughing my f**king ass off',
'rotflmfaopimp': 'rolling on the floor laughing my f**king ass off peeing in my pants',
'rotflmfaopmp': 'rolling on the floor laughing my ass off pissin my pants',
'rotflmfho': 'rolling on the floor laughing my f**king head off',
'rotflmho': 'rolling on the floor laughing my head off',
'rotflmmfao': 'rolling on the floor laughing my mother f**king ass off',
'rotflol': 'rolling on the floor laughing out loud',
'rotfpm': 'rolling on the floor pissing myself',
'rotfwlmao': 'rolling on the floor while laughing my ass off',
'rotg': 'rolling on the ground',
'rotgl': 'roll on the ground laughing',
'rotglmao': 'rolling on the ground laughing my ass off',
'rotw': 'rest of the world',
'rowyco': 'rock out with your c**k out',
'rox': 'rocks',
'roxor': 'rock',
'roxorz': 'rocks',
'roxxor': 'rock',
'rp': 'roleplay',
'rpg': 'role playing game',
'rpita': 'royal pain in the ass',
'rplbk': 'reply back',
'rpo': 'royally pissed off',
'rq': 'real quick',
'rr': 'rest room',
'rrb': 'restroom break',
'rsn': 'real soon now',
'rsp': 'respawn',
'rspct': 'respect',
'rsps': 'runescape private server',
'rt': 'retweet',
'rta': 'read the article',
'rtard': 'retard',
'rtbq': 'read the blinking question',
'rtf': 'return the favor',
'rtfa': 'read the f**king article',
'rtffp': 'read the f**king front page',
'rtfm': 'read the f**king manual',
'rtfmfm': 'read the f**king manual f**king moron',
'rtfmm': 'read the f**king manual moron',
'rtfms': 'read the f**king manual stupid',
'rtfp': 'read the f**king post',
'rtfq': 'read the f**king question',
'rtfs': 'read the f**king summary',
'rtfu': 'ready the f**k up',
'rtg': 'ready to go',
'rtl': 'report the loss',
'rtm': 'read the manual',
'rtr': 'read the rules',
'rtry': 'retry',
'rts': 'real-time strategy',
'ru': 'are you',
'ru18': 'are you 18',
'rua': 'are you alone',
'ruabog': 'are you a boy or girl',
'ruagoab': 'are you a girl or a boy',
'rubz2nt': 'are you busy tonight',
'rufkm': 'are you f**king kidding me',
'rugay': 'are you gay',
'rugta': 'are you going to answer',
'ruh': 'are you horny',
'ruk': 'are you ok?',
'rukm': 'are you kidding me',
'rul8': 'are you late',
'rumf': 'are you male or female',
'runnin': 'running',
'ruok': 'are you ok?',
'rur': 'are you ready',
'rut': 'are you there',
'ruwm': 'are you watching me',
'rwb': 'rich white b***h',
'rys': 'are you single',
'ryt': 'right',
'ryte': 'right'}
s = { "s'ok": 'yes ok',
"s'pose": 'suppose',
"s'up": 'what is up',
's.i.n.g.l.e': 'stay intoxicated nightly, get laid everyday.',
's.i.t.': 'stay in touch',
's.o.a.b.': 'son of a b***h',
's.o.b.': 'son of a b***h',
's.w.a.k.': 'sealed with a kiss',
's/b': 'should be',
's2a': 'sent to all',
's2bu': 'sucks to be you',
's2g': 'swear to god',
's2r': 'send to receive',
's2u': 'same to you',
's2us': 'speak to you soon',
's3x': 'sex.',
's4se': 'sight for sore eyes',
's8ter': 'skater',
'sab': 'slap a b***h',
'safm': 'stay away from me',
'sagn': 'spelling and grammar nazi',
'sah': 'sexy as hell',
'sahm': 'stay at home mom',
'sase': 'self addressed stamped envelope',
'sbc': 'sorry bout caps',
'sbcg4ap': 'strongbads cool game for attractive people',
'sbd': 'silent but deadly',
'sblai': 'stop babbaling like an idiot',
'sbrd': 'so bored',
'sbs': 'such bull s**t',
'sbt': 'sorry bout that',
'scnr': "sorry, i couldn't resist",
'scool': 'school',
'scrilla': 'money',
'scrt': 'secret',
'scurred': 'scared',
'sd': 'suck d**k',
'sde': 'software defined environment',
'sdf^': 'shut da f**k up',
'sdk': 'software development kit',
'sdlc': 'software development life cycle',
'sec': 'second',
'secks': 'sex',
'secksea': 'sexy',
'secksy': 'sexy',
'sed': 'said',
'see through your eyes': 'stye',
'seg': 's**t eatin grin',
'seks': 'sex',
'sellin': 'selling',
'seo': 'search engine optimization',
'serp': 'search engine results page',
'sexc': 'sexy',
'sexe': 'sexy',
'sexi': 'sexy',
'sexii': 'sexy',
'sexilicious': 'very sexy',
'sexx0rz': 'sex',
'sez': 'says',
'sfam': 'sister from another mother',
'sfao': 'sorry for any offense',
'sfe': 'safe',
'sfh': 'so f**king hot',
'sfipmp': 'so funny i peed my pants',
'sfm': 'so f**king much',
'sfr': 'so f**king random',
'sfs': 'so f**king stupid',
'sfsg': 'so far so good',
'sftbc': 'sorry for the broad cast',
'sfu': 'shut the f**k up',
'sfw': 'safe for work',
'sfwuz': 'safe for work until zoomed',
'sfy': 'speak for yourself',
'sfyl': 'sorry for your loss',
'sg': 'so good',
'sgb': 'straight/gay/bisexual',
'sgbadq': 'search google before asking dumb questions',
'sgi': 'still got it',
'sgtm': 'slightly gigling to myself',
'sh': 's**t happens',
"sha'n't": 'shall not',
'shag': 'f**k',
"shan't": 'shall not',
"shan't've": 'shall not have',
'shawty': 'girl',
'shd': 'should',
"she'd": 'she would',
"she'd've": 'she would have',
"she'll": 'she will',
"she'll've": 'she will have',
"she's": 'she is',
'shexi': 'sexy',
'shexy': 'sexy',
'shiat': 's**t',
'shiet': 's**t',
'shite': 's**t',
'shiz': 's**t',
'shizit': 's**t',
'shiznat': 's**t',
'shiznit': 's**t',
'shizz': 's**t',
'shizzle': 's**t',
'shld': 'should',
'shmexy': 'sexy',
'shmily': 'see how much i love you',
'sho': 'sure',
"sho'nuff": 'sure enough',
"shoulda" : "should have"
"should've": 'should have',
"shouldn't": 'should not',
"shouldn't've": 'should not have',
'showin': 'showing',
'shr': 's**t happens right',
'shrn': 'so hot right now',
'sht': 's**t',
'shtf': 's**t hits the fan',
'shud': 'should',
'shuddup': 'shut up',
'shup': 'shut up',
'shure': 'sure',
'shut^': 'shut up',
'shwr': 'shower',
'shyat': 's**t',
'shyt': 's**t',
'siao': 'school is almost over',
'sibir': 'sibling in room',
'sic': 'said in context',
'sicl': 'sitting in chair laughing',
'sif': 'as if',
"sifn't": 'as if not',
'sig': 'signature',
'siggy': 'signature',
'silf': "sister i'd like to f**k",
'simcl': 'sitting in my chair laughing',
'simclmao': 'sitting in my chair laughing my ass off',
'siol': 'shout it out loud',
'sis': 'sister',
'sista': 'sister',
'sitb': 'sex in the but',
'sitd': 'still in the dark',
'sitmf': 'say it to my face',
'siu': 'suck it up',
'siuya': 'shove it up your ass',
'sk': 'spawn kill',
'sk8': 'skate',
'sk8er': 'skater',
'sk8ing': 'skating',
'sk8r': 'skater',
'sk8ter': 'skater',
'sk8tr': 'skater',
'skb': 'should know better',
'sked': 'schedule',
'skeet': 'ejaculate',
'skewl': 'school',
'skhool': 'school',
'skillz': 'skills',
'skl': 'school',
'skool': 'school',
'skoul': 'school',
'sktr': 'skater',
'skwl': 'school',
'sl4n': 'so long for now',
'sl^t': 's**t',
'sleepin': 'sleeping',
'sleepn': 'sleeping',
'slf': 'sexy little f**k',
'slgb': 'straight/lesbian/gay/bisexual',
'slng': 'slang',
'slo': 'slow',
'slore': 's**tty w***e',
'slos': 'someone looking over shoulder',
'slp': 'sleep',
'slt': 'something like that',
'sm': 'social media',
'sm1': 'someone',
'sma': 'suck my ass',
'smb': 'see my blog',
'smbd': 'suck my big d**k',
'smbt': 'suck my big toe',
'smc': 'suck my c**k',
'smd': 'suck my d**k',
'smdb': 'suck my d**k b***h',
'smdvq': 'suck my d**k quickly',
'smeg': 'f**k',
'smexy': 'sexy',
'smf': 'stupid motherf**ker',
'smfd': 'suck my f**king d**k',
'smfpos': 'stupid mother f**king piece of s**t',
'smh': 'shaking my head',
'smhb': 'suck my hairy balls',
'smhid': 'shaking my head in disgust',
'smho': 'screaming my head off',
'smithwaws': 'smack me in the head with a wooden spoon',
'smofo': 'stupid mother f**ker',
'smst': 'somebody missed snack time',
'smt': 'suck my tits',
'smthin': 'something',
'smthng': 'something',
'smtm': 'sometime',
'smto': 'sticking my tongue out',
'smtoay': 'sticking my tongue out at you',
'sn': 'screen name',
'snafu': 'situation normal all f**ked up',
'snafubar': 'situation normal all f**ked up beyond any recognition',
'snes': 'super nintendo entertainment system',
'snew': "what's new",
'snf': 'so not fair',
'snl': 'saturday night live',
'snm': 'say no more',
'snog': 'kiss',
'snogged': 'kissed',
"so's": 'so as',
"so've": 'so have',
'soa': 'service oriented architecture',
'soab': 'son of a btch',
'soad': 'system of a down',
'soafb': 'son of a f**king b***h',
'sob': 'son of a b***h',
'sobs': 'same, old, boring s**t',
'soc': 'same old crap',
'soe': 'service oriented enterprise',
'sof': 'smile on face',
'sofas': 'stepping out for a smoke',
'sofs': 'same old f**king s**t',
'soi': 'service oriented infrastructure',
'sok': "it's ok",
'sokay': "it's okay",
'sol': 's**t outta luck',
"som'm": 'something',
'som1': 'someone',
'somadn': 'sitting on my ass doing nothing',
'some1': 'someone',
'soml': 'story of my life',
'soo': 'so',
'soobs': 'saggy boobs',
'sool': 's**t out of luck',
'sop': 'same old place',
'sorg': 'straight or gay',
'sorreh': 'sorry',
'sorta': 'sort of',
'sos': 'same old s**t',
'sosdd': 'same old s**t, different day',
'sosg': 'spouse over shoulder gone',
'sot': 'suck on this',
'sotc': 'stupid off topic crap',
'sotr': 'sex on the road',
'sow': 'statement of work',
'sowi': 'sorry',
'sowwy': 'sorry',
'soz': 'sorry',
'spesh': 'special',
'spk': 'speak',
'spk2ul8r': 'speak to you later',
'sploits': 'exploits',
'sploitz': 'exploits',
'spos': 'stupid peace of s**t',
'sprm': 'sperm',
'sqtm': 'snickering quietly to myself',
'srch': 'search',
'srly': 'seriously',
'sroucks': "that's cool, but it still sucks",
'srry': 'sorry',
'srs': 'serious',
'srsly': 'seriously',
'srvis': 'service',
'sry': 'sorry',
'srynd2g': 'sorry need to go',
'srzly': 'seriously',
'ss': 'screenshot',
'ss4l': 'smoking sista for life',
'ssdd': 'same s**t, different day',
'ssdp': 'same s**t different pile',
'ssia': 'subject says it all',
'ssl': 'secure sockets layer',
'ssob': 'stupid sons of b***hes',
'ssry': 'so sorry',
'sssd': 'same s**t same day',
'st': 'stop that',
'st1': 'stoned',
'st8': 'state',
'stats': 'your sex and age',
'stb': 'soon to be',
'stbm': 'sucks to be me',
'stbx': 'soon to be ex',
'stby': 'sucks to be you',
'std': 'sexually transmitted disease',
'steamloller': 'laughing. alot.',
'stfd': 'sit the f**k down',
'stff': 'stuff',
'stfm': 'search the f**king manual',
'stfng': 'search the f**king news group',
'stfu': 'shut the f**k up',
'stfua': 'shut the f**k up already',
'stfuah': 'shut the f**k up a**h**e',
'stfub': 'shut the f**k up b***h',
'stfuda': 'shut the f**k up dumb ass',
'stfugbtw': 'shut the f**k up and get back to work',
'stfun': 'shut the f**k up n****r',
'stfuogtfo': 'shut the f**k up or get the f**k out.',
'stfuppercut': 'shut the f**k up',
'stfuyb': 'shut the f**k up you b***h',
'stfuysoab': 'shut the f**k up you son of a b***h',
'stfw': 'search the f**king web',
'stg': 'swear to god',
'sth': 'something',
'sthing': 'something',
'sthu': 'shut the hell up',
'stm': 'smiling to myself',
'stoopid': 'stupid',
'stpd': 'stupid',
'str8': 'straight',
'str8up': 'straight up',
'sts': 'so to speak',
'stsp': 'same time same place',
'stt': 'same time tomorrow',
'stufu': 'stupid f**ker',
'stupd': 'stupid',
'stw': 'share the wealth',
'stys': 'speak to you soon',
'su': 'shut up',
'sua': 'see you at',
'suabq': 'shut up and be quiet',
'suagooml': 'shut up and get out of my life',
'suib': 'shut up im busy',
'suk': 'suck',
'suka': 'sucker',
'sukz': 'sucks',
'sul': 'see you later',
'sum1': 'someone',
'sumfin': 'something',
'summin': 'something',
'sumone': 'someone',
"sumthin'": 'something',
'sumtin': 'something',
'sup': "what's up",
'supa': 'super',
'supposably': 'supposedly',
'sus': 'see you soon',
'susfu': 'situation unchanged, still f**ked up',
'sut': 'see you tomorrow',
'sutuct': 'so you think you can type',
'sux': 'sucks',
'sux0rz': 'sucks',
'sux2bu': 'sucks to be you',
'suxor': 'sucks',
'suxors': 'sucks',
'suxorz': 'sucks',
'suxx': 'sucks',
'suxxor': 'sucks',
'suyah': 'shut up you ass hole!',
'svn': 'seven',
'svu': 'special victims unit',
'sw': 'so what',
'swafk': 'sealed with a friendly kiss',
'swak': 'sealed with a kiss',
'swakaah': 'sealed with a kiss and a hug',
'swalk': 'sealed with a loving kiss',
'swf': 'single white female',
'swm': 'single white male',
'swmbo': 'she who must be obeyed',
'swmt': 'stop wasting my time',
'swp': 'sorry wrong person',
'swswta': 'so what should we talk about',
'swsw2b': 'single when she wants to be',
'swt': 'sweet',
'swtf': 'seriously, what the f**k',
'sx': 'sex',
'sxc': 'sexy',
'sxcy': 'sexy',
'sxe': 'straight edge',
'sxi': 'sexy',
'sxs': 'sex',
'sxy': 'sexy',
'sya': 'see you again.',
'syatp': 'see you at the party',
'sydim': 'stick your d**k in me',
'sydlm': 'shut your dirty little mouth',
'sydm': 'shut your damn mouth',
'syfm': 'shut your f**king mouth',
'syiab': 'see you in a bit',
'syiaf': 'see you in a few',
'syl': 'see you later',
'syl8r': 'see you later',
'sym': 'shut your mouth',
'syoa': 'save your own ass',
'syotbf': 'see you on the battlefield',
'syrs': 'see ya real soon',
'sys': 'see you soon',
'sysop': 'system operator',
'syt': 'see you there',
'sytycd': 'so you think you can dance',
'syu': 'sex you up',
'sz': 'sorry'}
t = {'t#3': 'the',
't,ftfy': 'there, fixed that for you',
't.t.y.l': 'talk to you later',
't/a': 'try again',
't2b': 'time to blunt',
't2m': 'talk to me',
't2u': 'talking to you',
't2ul': 'talk to you later',
't2ul8r': 'talk to you later',
't3h': 'the',
't4a': 'thanks for asking',
't4m': 'transgender for male',
't8st': 'taste',
'ta': 'thanks again',
'taci': "that's a crappy idea",
'tafn': "that's all for now",
'taht': 'that',
'tai': 'think about it',
'taig': "that's all i got.",
'tal': 'thanks a lot',
'talkin': 'talking',
'tanq': 'thank you',
'tanstaafl': "there ain't no such thing as a free lunch",
'tard': 'retard',
'tarfu': 'things are really f**ked up',
'tat': 'that',
'tat2': 'tattoo',
'tau': 'thinking about you',
'taunch': 'te amo un chingo',
'taw': 'teachers are watching',
'tay': 'thinking about you',
'tb': 'text back',
'tb4u': 'too bad for you',
'tba': 'to be anounced',
'tbc': 'to be continued',
'tbd': 'to be decided',
'tbf': 'to be fair',
'tbfh': 'to be f**king honest',
'tbfu': 'too bad for you',
'tbh': 'to be honest',
'tbhimo': 'to be honest in my opinion',
'tbhwu': 'to be honest with you',
'tbnt': 'thanks but no thanks',
'tbp': 'the pirate bay',
'tbpfh': 'to be perfectly f**king honest',
'tbph': 'to be perfectly honest',
'tbqf': 'to be quite frank',
'tbqh': 'to be quite honest',
'tbss': 'too bad so sad',
'tbt': 'throwback thursday',
'tbtfh': 'to be totally freaking honest',
'tbvh': 'to be very honest',
'tbya': 'think before you act',
'tc': 'take care',
'tcfc': 'too close for comfort',
'tcfm': 'too cool for me',
'tcg': 'trading card game',
'tchbo': 'topic creater has been owned',
'tcial': 'the cake is a lie',
'tcoy': 'take care of yourself',
'tcp': 'transmission control protocol',
'tcp/ip': 'transmission control protocol/internet protocol',
'td2m': 'talk dirty to me',
'tddup': 'till death do us part',
'tdf': 'to die for',
'tdl': 'too damn lazy',
'tdtm': 'talk dirty to me',
'tdtml': 'talk dirty to me later',
'tdwdtg': 'the devil went down to georgia',
'te': 'team effort',
'teh': 'the',
'teotwawki': 'the end of the world as we know it',
'terd': 's**t',
'tf': 'The f**k',
'tf2': 'team fortress 2',
'tfa': 'the f**king article',
'tfb': 'time for bed',
'tfbundy': 'totaly f**ked but unfortunatly not dead yet',
'tfc': 'team fortress classic',
'tfd': 'total f**king disaster',
'tff': "that's f**king funny",
'tfft': 'thank f**k for that',
'tffw': 'too funny for words',
'tfh': 'thread from hell',
'tfic': 'tongue firmly in cheek',
'tfiik': 'the f**k if i know',
'tfl': 'thanks for looking',
'tfln': 'thanx for last night',
'tfm': 'too f**king much',
'tfs': 'thanks for sharing',
'tfta': 'thanks for the add',
'tfti': 'thanks for the information',
'tfu': "that's f**ked up",
'tfu2baw': 'time for you to buy a watch',
'tfw': 'that feeling when',
'tg': 'thank god',
'tgfe': 'together forever',
'tgfitw': 'the greatest fans in the world',
'tgft': 'thank god for that',
'tgfu': 'too good for you',
'tgfuap': 'thank god for unanswered prayers',
'tghig': 'thank god husband is gone',
'tgif': "thank god it's friday",
'tgiff': 'thank god its f**king friday',
'tgis': "thank god it's saturday",
'tgiwjo': 'thank god it was just once',
'tgsttttptct': 'thank god someone took the time to put this crap together',
'tgtbt': 'too good to be true',
'tgwig': 'thank god wife is gone',
'tgws': 'that goes without saying',
'th4nk5': 'thanks',
'th@>': 'that',
'th@' : 'though',
'tha': 'the',
'thankies': 'thank you',
'thankx': 'thank you',
'thanq': 'thank you',
'thanx': 'thank you',
'thanxx': 'thank you',
'thar': 'there',
"that'd": 'that would',
"that'd've": 'that would have',
"that's": 'that is',
'thatz': "that's",
"there'd": 'there would',
"there'd've": 'there would have',
"there's": 'there is',
"they'd": 'they would',
"they'd've": 'they would have',
"they'll": 'they will',
"they'll've": 'they will have',
"they're": 'they are',
"they've": 'they have',
'theyre': 'they are',
"this's": 'this is',
'thku': 'thank you',
'thn': 'then',
'thnk': 'think',
'thnx': 'thanks',
'tho': 'though',
'thot': 'the hoe of today',
'thr': 'there',
'thr4': 'therefore',
'thru': 'through',
'tht': 'that',
'thwdi': 'thats how we do it',
'thwy': 'the hell with you!',
'thx': 'thank you',
'thxx': 'thanks',
'thz': 'thank you',
'ti2o': 'that is too obious',
'tia': 'thanks in advance',
'tiafayh': 'thanks in advance for all your help',
'tiai': 'take it all in',
'tias': 'try it and see',
'tiatwtcc': 'this is a trap word to catch copiers',
'tif': 'this is fun',
'tif2m': 'this is f**king 2 much',
'tifs': 'this is funny s**t',
'tifu': 'that is f**ked up',
'tigger': 'tiger',
'tiic': 'the idiots in control',
'til': 'until',
'tilf': "teenager i'd like to f**k",
'tinf': 'this is not fair',
'tinla': 'this is not legal advice',
'tinstaafl': 'there is no such thing as a free lunch',
'tioli': 'take it or leave it',
'tis': 'is',
'tisc': 'that is so cool',
'tisfu': 'that is so f**ked up',
'tisg': 'this is so gay',
'tish': 'this',
'tisly': 'that is so last year',
'tisnf': 'that is so not fair',
'tiss': 'this is some s**t',
'tisw': 'that is so wrong',
'tiw': 'teacher is watching',
'tix': 'tickets',
'tjb': 'thats just boring',
'tk': 'team kill',
'tk2ul': 'talk to you later',
'tkd': 'tae kwon do',
'tker': 'team killer',
'tks': 'thanks',
'tku': 'thank you',
'tl': 'tough luck',
'tl,dr': "too long; didn't read",
'tl8r': 'talk later',
'tl:dr': "too long; didn't read",
'tl; dr': "to long; didn't read",
'tl;dr': "too long; didn't read",
'tla': 'three letter acronym',
'tlc': 'tender loving care',
'tld': 'told',
'tldnr': 'too long, did not read',
'tldr': "too long, didn't read.",
'tlgo': 'the list goes on',
'tliwwv': 'this link is worthless without video',
'tlk': 'talk',
'tlk2me': 'talk to me',
'tlk2ul8r': 'talk to you later',
'tlkin': 'talking',
'tlkn': 'talking',
'tltpr': 'too long to proof read.',
'tlyk': 'to let you know',
'tma': 'take my advice',
'tmaai': 'tell me all about it',
'tmai': 'tell me about it',
'tmbi': 'tell me about it',
'tmi': 'too much information',
'tmk': 'to my knowledge',
'tml': 'tell me later',
'tmmrw': 'tomorrow',
'tmnt': 'teenage mutant ninja turtles',
'tmo': 'take me out',
'tmoro': 'tomorrow',
'tmoz': 'tomorrow',
'tmp': 'text my phone',
'tmr': 'tomorrow',
'tmr@ia': 'the monkeys are at it again',
'tmrrw': 'tomorrow',
'tmrw': 'tomorrow',
'tmrz': 'tomorrow',
'tms': 'that makes sense',
'tmsaisti': "that's my story and i'm sticking to it.",
'tmsg': 'tell me something good',
'tmsidk': "tell me somthing i don't know",
'tmth': 'too much to handle',
'tmtmo': 'text me tomorrow',
'tmtoyh': 'too much time on your hands',
'tmtt': 'tell me the truth',
'tmw': 'too much work',
'tmwfi': 'take my word for it',
'tmz': 'tomorrow',
'tn1': 'trust no-one',
'tna': 'tits and ass',
'tnc': 'totally not cool',
'tnf': "that's not funny",
'tnlnsl': 'took nothing left nothing signed log',
'tnmp': 'totally not my problem',
'tnomb': "that's none of my business",
'tnx': 'thanks',
'tnxz': 'thanks',
"to've": 'to have',
'tob': 'teacher over back',
'tofy': 'thinking of you',
'togtfooh': 'tits or get the f**k out of here',
'toh': 'typing one handed',
'tok': "that's ok",
'tok2ul8r': "i'll talk to you later",
'tolol': 'thinking of laughing out loud',
'tomm': 'tommorow',
'tomoro': 'tommorrow',
'tomoz': 'tomorrow',
'tonite': 'tonight',
'tos': 'terms of service',
'totd': 'tip of the day',
'totes': 'totally',
'totl': 'total',
'totm': 'top of the morning',
'totp': 'talking on the phone',
'totpd': 'top of the page dance',
'tou': 'thinking of you',
'toya': 'thinking of you always',
'tp': 'toilet paper',
'tpb': 'the pirate bay',
'tpf': "that's pretty funny",
'tpiwwp': 'this post is worthless without pictures',
'tps': 'test procedure specification',
'tptb': 'the powers that be',
'tq': 'thank you',
'trani': 'transexual',
'tranny': 'transexual',
'trans': 'transfer',
'trble': 'trouble',
'trd': 'tired',
'tres': 'so',
'trnsl8': 'translate',
'trnsltr': 'translator',
'troll': 'person who diliberately stirs up trouble',
'tru': 'through',
'ts': 'talking s**t',
'tsc': "that's so cool",
'tsff': 'thats so f**kin funny',
'tsig': 'that site is gay',
'tsl': 'the single life',
'tsm': 'thanks so much',
'tsnf': "that's so not fair",
'tss': "that's so sweet",
'tstoac': 'too stupid to own a computer',
'tswc': 'tell someone who cares',
'tt4n': 'ta ta for now',
'ttbc': 'try to be cool',
'ttbomk': 'to the best o fmy knowledge',
'ttc': 'text the cell',
'ttf': "that's too funny",
'ttfaf': 'through the fire and flames',
'ttfn': 'ta ta for now',
'ttg': 'time to go',
'tthb': 'try to hurry back',
'ttihlic': 'try to imagine how little i care',
'ttiuwiop': 'this thread is useless without pics',
'ttiuwop': 'this thread is useless without pics',
'ttiuwp': 'this thread is useless without pictures',
'ttiwwop': 'this thread is worthless without pics',
'ttiwwp': 'this thread is worthless without pics',
'ttl': 'total',
'ttlly': 'totally',
'ttly': 'totally',
'ttm': 'talk to me',
'ttml': 'talk to me later',
'ttmn': 'talk to me now',
'ttms': 'talking to myself',
'ttr': 'time to run',
'ttrf': "that's the rules, f**ker",
'tts': 'text to speech',
'ttt': 'to the top',
'ttth': 'talk to the hand',
'tttt': 'to tell the truth',
'ttul': 'talk to you later',
'ttul8r': 'talk to you later',
'ttus': 'talk to you soon',
'ttut': 'talk to you tomorrow',
'ttutt': 'to tell you the truth',
'tty': 'talk to you',
'ttya': 'thanks to you all',
'ttyab': 'talk to you after breakfast',
'ttyad': 'talk to you after dinner',
'ttyal': 'talk to you after lunch',
'ttyas': 'talk to you at school',
'ttyiam': 'talk to you in a minute',
'ttyitm': 'talk to you in the morning',
'ttyl': 'talk to you later',
'ttyl2': 'talk to you later too',
'ttyl8r': 'talk to you later',
'ttylo': 'talk to you later on',
'ttylt': 'talk to you later today',
'ttyn': 'talk to you never',
'ttyna': 'talk to you never again',
'ttynl': 'talk to you never loser',
'ttynw': 'talk to you next week',
'ttyo': 'talk to you online',
'ttyob': 'tend to your own business',
'ttyotp': 'talk to you on the phone',
'ttyrs': 'talk to you really soon',
'ttys': 'talk to you soon',
'ttyt': 'talk to you tomorrow',
'ttytm': 'talk to you tomorrow',
'ttytt': 'to tell you the truth',
'ttyw': 'talk to you whenever',
'ttywl': 'talk to you way later',
'tu': 'thank you',
'tuff': 'tough',
'tuh': 'to',
'tul': 'text you later',
'tut': 'take your time',
'tuvm': 'thank you very much',
'tv': 'television',
'tvm': 'thanks very much',
'tw': 'teacher watching',
'twajs': 'that was a joke, son.',
'twat': 'vagina',
'twbc': 'that would be cool',
'twdah': 'that was dumb as hell',
'twf': 'that was funny',
'twfaf': 'thats what friends are for',
'twg': 'that was great',
'twi': 'texting while intoxicated',
'twis': "that's what i said",
'twit': 'tweet',
'twn': 'town',
'twoh': 'typing with one hand',
'tws2wa': 'that was so 2 weeks ago',
'twss': "that's what she said",
'twsy': 'that was so yeterday',
'twttr': 'twitter',
'twvsoy': 'that was very stupid of you',
'twyl': 'talk with you later',
'twys': 'talk with you soon',
'tx': 'thanks',
'txs': 'thanks',
'txt': 'text',
'txting': 'texting',
'txtms': 'text me soon',
'txtyl': 'text you later',
'ty': 'thank you',
'tyclos': 'turn your caps lock off, stupid',
'tyfi': 'thank you for invite',
'tyfn': 'thank you for nothing',
'tyfyc': 'thank you for your comment',
'tyfyt': 'thank you for your time',
'tyl': 'text you later',
'tym': 'time',
'tyme': 'time',
'typ': 'thank you partner',
'typo': 'typing mistake',
'tyred': 'tired',
'tys': 'told you so',
'tysfm': 'thank you so f**king much',
'tysm': 'thank you so much',
'tysvm': 'thank you so very much',
'tyt': 'take your time',
'tyto': 'take your top off',
'tyty': 'thank you thank you',
'tyvm': 'thank you very much',
'tyvvm': 'thank you very very much'}
u = {'u': 'you',
'u iz a 304': 'you is a hoe',
"u'd": 'you would',
"u'll": 'you will',
"u'r": "you're",
"u'v": 'you have',
"u've": "you've",
'u/l': 'upload',
'u/n': 'username',
'u2': 'you too',
'u2c': 'unable to contact',
'u2u': 'up to you',
'u4e': 'yours for ever',
'u4i': 'up for it',
'ua': 'user agreement',
'uaaaa': 'universal association against acronym abuse',
'uat': 'user acceptance testing',
'uayor': 'use at your own risk',
'ub3r': 'super',
'uctaodnt': "you can't teach an old dog new tricks",
'udc': "you don't care",
'udcbm': "you don't care about me",
'udek': "you don't even know",
'uds': 'you dumb s**t',
'udwk': "you don't want to know",
'udy': 'you done yet',
'ufab': 'ugly fat ass b***h',
'ufia': 'unsolicited finger in the anus',
'ufic': 'unsolicited finger in chili',
'ufmf': 'you funny mother f**ker',
'ufr': 'upon further review',
'ugba': 'you gay b***h ass',
'ugtr': 'you got that right',
'uhab': 'you have a blog',
'uhems': 'you hardly ever make sense',
'ui': 'user interface',
'ujds': 'u just did s**t',
'ukr': 'you know right',
'ukwim': 'you know what i mean',
'ul': 'unlucky',
'ulbom': 'you looked better on myspace',
'umfriend': 'sexual partner',
'un2bo': 'you need to back off',
'un4rtun8ly': 'unfortunately',
'unt': 'until next time',
'uom': 'you owe me',
'upcia': 'unsolicited pool cue in anus',
'upia': 'unsolicited pencil in anus',
'upmo': 'you piss me off',
'upos': 'you piece of s**t',
'upw': 'unidentified party wound',
'ur': 'your',
'ur2g': 'you are too good',
'ur6c': "you're sexy",
'ura': 'you are a',
'uradrk': "you're a dork",
'urafb': 'you are a f**king b***h',
'uraqt': 'you are a cutie',
'urcrzy': 'you are crazy',
'ure': 'you are',
'urg': 'you are gay',
'urht': "you're hot",
'url': 'uniform resource locator',
'url8': 'you are late',
'urms': 'you rock my socks',
'urmw': 'you are my world',
'urnc': 'you are not cool',
'urng2bt': 'you are not going to believe this',
'urs': 'yours',
'ursab': 'you are such a b***h',
'ursdf': 'you are so damn fine',
'ursg': 'you are so gay',
'ursh': 'you are so hot',
'urssb': 'you are so sexy baby',
'urstpid': 'you are stupid',
'urstu': 'you are stupid',
'urtb': 'you are the best',
'urtbitw': 'you are the best in the world!',
'urtrd': 'you retard',
'urtw': 'you are the worst',
'urw': 'you are weird',
'uryyfm': 'you are too wise for me',
'usa': 'united states of america',
'usck': 'you suck',
'usd': 'united states dollar',
'ussr': 'the union of soviet socialist republics',
'usuk': 'you suck',
'usux': 'you suck',
'ut': 'you there',
'uta': 'up the ass',
'utfs': 'use the f**king search',
'utfse': 'use the f**king search engine',
'utm': 'you tell me',
'uttm': 'you talking to me?',
'utube': 'youtube',
'utw': 'used to work',
'uty': "it's up to you",
'uve': "you've",
'uvgtbsm': 'you have got to be shiting me',
'uw': "you're welcome",
'uwc': 'you are welcome',
'uya': 'up your ass',
'uyab': 'up your ass b***h'}
v = {'v4g1n4': 'vagina',
'vag': 'vagina',
'vajayjay': 'vagina',
'vb': 'visual basic',
'vbeg': 'very big evil grin',
'vbg': 'very big grin',
'vet': 'vetenary doctor',
'vf': 'very funny',
'vfe': 'virgins 4 ever',
'vff': 'verry f**king funny',
'vfm': 'value for money',
'vgg': 'very good game',
'vgh': 'very good hand',
'vgl': 'very good looking',
'vgn': 'video game nerd',
'vid': 'video',
'vids': 'videos',
'vip': 'very important person',
'vleo': 'very low earth orbit',
'vlog': 'video log',
'vn': 'very nice',
'vnc': 'virtual network computing',
'vnh': 'very nice hand',
'voip': 'voice over ip',
'vrsty': 'varsity',
'vry': 'very',
'vs': 'versus',
'vwd': 'very well done',
'vweg': 'very wicked evil grin',
'vzit': 'visit',
'vzn': 'verizon'}
w = {"w'sup": "what is up",
'w.b.s.': 'write back soon',
'w.e': 'whatever',
'w.e.': 'whatever',
'w.o.w': 'world of warcraft',
'w.o.w.': 'world of warcraft',
'w/': 'with',
'w/b': 'write back',
'w/e': 'whatever',
'w/end': 'weekend',
'w/eva': 'whatever',
'w/o': 'with out',
'w/out': 'without',
'w/u': 'with you',
'w00t': 'woohoo',
'w012d': 'word',
'w2d': 'what to do',
'w2f': 'want to f**k',
'w2g': 'way to go',
'w2ho': 'want to hang out',
'w2m': 'want to meet',
'w33d': 'weed',
'w8': 'wait',
'w8am': 'wait a minute',
'w8ing': 'waiting',
'w8t4me': 'wait for me',
'w8ter': 'waiter',
'w911': 'wife in room',
'w\\e': 'whatever',
'wab': 'what a b***h',
'wad': 'without a doubt',
'wad ^': "what's up?",
'wadr': 'with all due respect',
'wadzup': "what's up?",
'waf': 'weird as f**k',
'wafda': 'what a f**king dumb ass',
'wafl': 'what a f**king loser',
'wafm': 'wait a f**king minute',
'wafn': 'what a f**ken noob',
'wai': 'what an idiot',
'waloc': 'what a load of crap',
'walstib': "what a long strange trip it's been",
'wam': 'wait a minute',
'wamh': 'with all my heart',
'wan2tlk': 'want to talk',
'wana': 'want to',
'wanafuk': 'wanna f**k',
'wanker': 'masturbater',
'wanking': 'masturbating',
'wanna': 'want to',
'wansta': 'wanna be ganster',
'warez': 'illegally obtained software',
'was^': "what's up",
"wasn't": 'was not',
'wassup': "what's up?",
'wasup': "what's up",
'wat': 'what',
"wat's^": 'whats up',
'watcha': 'what are you',
'watev': 'whatever',
'wateva': 'whatever',
'watevr': 'whatever',
'watevs': 'whatever',
'watp': 'we are the people',
'wats': 'whats',
'wats ^': 'whats up',
'wats^': "what's up?",
'watz ^': "what's up",
'wau': 'what about you',
'wau^2': 'what are you up to?',
'waud': 'what are you doing',
'waug': 'where are you going',
'wauw': 'what are you wearing',
'waw': 'what a w***e',
'waycb': 'when are you coming back',
'wayd': 'what are you doing',
'waygow': 'who are you going out with',
'wayh': 'why are you here',
'wayjdin': 'why are you just doing it now',
'wayn': 'where are you now',
'waysttm': 'why are you still talking to me',
'waysw': 'why are you so weird',
'wayt': 'what are you thinking?',
'wayta': 'what are you talking about',
'wayut': 'what are you up to',
'waz': 'what is',
'waz ^': "what's up",
'waz^': "what's up?",
'wazz': "what's",
'wazza': "what's up",
'wazzed': 'drunk',
'wazzup': "what's up",
'wb': 'welcome back',
'wbagnfarb': 'would be a good name for a rock band',
'wbb': 'will be back',
'wbbs': 'will be back soon',
'wbk': 'welcome back',
'wbp': 'welcome back partner',
'wbrb': 'will be right back',
'wbs': 'write back soon',
'wbu': 'what about you',
'wby': 'what about you',
'wc': 'who cares',
'wc3': 'warcraft iii',
'wcb': 'welcome back',
'wcm': 'women crush monday',
'wcutm': 'what can you tell me',
'wcw': 'webcam w***e',
'wd': 'well done',
'wdf': 'worth dying for',
'wdhlm': 'why doesnt he love me?',
'wdidn': 'what do i do now',
'wdim': 'what did i miss',
'wdk': "we don't know",
'wdtm': 'what does that mean',
'wduc': 'what do you care',
'wdum': 'what do you mean',
'wdus': 'what did you say',
'wdut': 'what do you think?',
'wdutom': 'what do you think of me',
'wduw': 'what do you want',
'wduwta': 'what do you wanna talk about',
'wduwtta': 'what do you want to talk about',
'wdwdn': 'what do we do now',
'wdwgw': 'where did we go wrong',
'wdya': 'why do you ask',
'wdydt': 'why do you do that',
'wdye': 'what do you expect',
'wdyl': 'who do you like',
'wdym': 'what do you mean',
'wdys': 'what did you say',
'wdyt': 'what do you think',
'wdytia': 'who do you think i am?',
'wdyw': 'what do you want',
'wdywd': 'what do you want to do?',
'wdywta': 'what do you wanna talk about',
'wdywtd': 'what do you want to do',
'wdywtdt': 'why do you want to do that?',
'wdywtta': 'what do you want to talk about',
"we'd": 'we would',
"we'd've": 'we would have',
"we'll": 'we will',
"we'll've": 'we will have',
"we're": 'we are',
"we've": 'we have',
'webby': 'webcam',
'weg': 'wicked evil grin',
'welc': 'welcome',
'wen': 'when',
"weren't": 'were not',
'werkz': 'works',
'wev': 'whatever',
'weve': 'what ever',
'wevr': 'whatever',
'wfh': 'working from home',
'wfhw': "what's for homework",
'wfm': 'works for me',
'wfyb': 'whatever floats your boat',
'wg': 'wicked gril',
'wgac': 'who gives a crap',
'wgaf': 'who gives a f**k',
'wgas': 'who gives a s**t',
'wgasa': 'who gives a s**t anyway',
'wgo': "what's going on",
'wgph2': 'want to go play halo 2?',
'wha': 'what?',
'whaddya': 'what do you',
'whaletail': 'thong',
"what'll": 'what will',
"what'll've": 'what will have',
"what're": 'what are',
"what's": 'what is',
"what've": 'what have',
'what^': "what's up?",
'whatcha': 'what are you',
'whatev': 'whatever',
'whatevs': 'whatever',
'whats ^': 'whats up',
"when's": 'when is',
"when've": 'when have',
'whenevs': 'whenever',
"where'd": 'where did',
"where's": 'where is',
"where've": 'where have',
'whevah': 'where ever',
'whever': 'whatever',
'whf': 'wanna have fun?',
'whit': 'with',
"who'll": 'who will',
"who'll've": 'who will have',
"who's": 'who is',
"who've": 'who have',
'whodi': 'friend',
'whr': 'where',
'whs': 'wanna have sex',
'wht': 'what',
'wht^': 'what up',
'whteva': 'what ever',
'whteve': 'whatever',
'whtever': 'whatever',
'whtevr': 'whatever',
'whtvr': 'whatever',
'whubu2': 'what have you been up to',
'whubut': 'what have you been up to',
'whut': 'what',
"why's": 'why is',
"why've": 'why have',
'whyb': 'where have you been',
'whyd': 'what have you done',
'wid': 'with',
'widout': 'without',
'wieu2': 'what is everyone up to',
'wif': 'with',
'wiid': 'what if i did',
'wilco': 'will comply',
"will've": 'will have',
'winnar': 'winner',
'wio': 'without',
'wip': 'work in progress',
'wit': 'with',
'witcha': 'with you',
'witfp': 'what is the f**king point',
'witu': 'with you',
'witw': 'what in the world',
'witwct': 'what is the world coming too',
'witwu': 'who is there with you',
'witwwyt': 'what in the world were you thinking',
'wiu': 'what is up?',
'wiuwu': 'what is up with you',
'wiv': 'with',
'wiw': 'wife is watching',
'wiwhu': 'wish i was holding you',
'wiwt': 'wish i was there',
'wiyp': 'what is your problem',
'wjwd': 'what jesus would do',
'wk': 'week',
'wkd': 'wicked',
'wkend': 'weekend',
'wl': 'will',
'wlc': 'welcome',
'wlcb': 'welcome back',
'wlcm': 'welcome',
'wld': 'would',
'wlkd': 'walked',
'wlos': 'wife looking over shoulder',
'wltm': 'would like to meet',
'wmao': 'working my ass off',
'wmd': 'weapons of mass destruction',
'wmgl': 'wish me good luck',
'wml': 'wish me luck',
'wmts': 'we must talk soon',
'wmyb': 'what makes you beautiful',
'wmytic': 'what makes you think i care',
'wn': 'when',
'wna': 'want to',
'wnkr': 'wanker',
'wnrn': 'why not right now',
'wnt': 'want',
'wntd': 'what not to do',
'woa': 'word of advice',
'woc': 'welcome on cam',
'wochit': 'watch it',
'woe': 'what on earth',
'woft': 'waste of f**king time',
'wogge': "what on god's green earth?",
'wogs': 'waste of good sperm',
'wolo': 'we only live once',
'wom': 'word of mouth',
'wombat': 'waste of money, brains, and time',
"won't": 'will not',
"won't've": 'will not have',
'woot': 'woohoo',
'workin': 'working',
'wos': 'waste of space',
'wot': 'what',
'wotevs': 'whatever',
'wotv': "what's on television?",
'wotw': 'word of the week',
"would've": 'would have',
"wouldn't": 'would not',
"wouldn't've": 'would not have',
'woum': "what's on your mind",
'wowzers': 'wow',
'woz': 'was',
'wp': 'wrong person',
'wpe': 'worst president ever (bush)',
'wrd': 'word',
'wrdo': 'weirdo',
'wrgad': 'who really gives a damn',
'wrgaf': 'who really gives a f**k?',
'wrk': 'work',
'wrm': 'which reminds me',
'wrng': 'wrong',
'wrt': 'with regard to',
'wrtg': 'writing',
'wrthls': 'worthless',
'wru': 'where are you',
'wrud': 'what are you doing',
'wruf': 'where are you from',
'wruu2': 'what are you up to',
'wryd': 'what are you doing',
'wsb': 'wanna cyber?',
'wsf': 'we should f**k',
'wshtf': 'when s**t hits the fan',
'wsi': 'why should i',
'wsibt': 'when should i be there',
'wsidi': 'why should i do it',
'wsop': 'world series of poker',
'wswta': 'what shall we talk about?',
'wt': 'what',
'wtaf': 'what the actual f**k',
'wtb': 'want to buy',
'wtbd': "what's the big deal",
'wtbh': 'what the bloody hell',
'wtc': 'what the crap',
'wtcf': 'what the crazy f**k',
'wtd': 'what the deuce',
'wtf': 'what the f**k',
'wtfaud': 'what the f**k are you doing?',
'wtfay': 'who the f**k are you',
'wtfayd': 'what the f**k are you doing',
'wtfayt': 'why the f**k are you talking',
'wtfayta': 'what the f**k are you talking about?',
'wtfb': 'what the f**k b***h',
'wtfbs': 'what the f**k bull s**t',
'wtfc': 'who the f**k cares',
'wtfdik': 'what the f**k do i know',
'wtfdtm': 'what the f**k does that mean',
'wtfdum': 'what the f**k do you mean',
'wtfduw': 'what the f**k do you want?',
'wtfdyw': 'what the f**k do you want',
'wtfe': 'what the f**k ever',
'wtfever': 'what the f**k ever',
'wtfg': 'what the f**king god',
'wtfh': 'what the f**king hell',
'wtfhb': 'what the f**king hell b***h',
'wtfhwt': 'what the f**king hell was that',
'wtfigo': 'what the f**k is going on',
'wtfigoh': 'what the f**k is going on here',
'wtfit': 'what the f**k is that',
'wtfits': 'what the f**k is this s**t',
'wtfiu': 'what the f**k is up',
'wtfiup': 'what the f**k is your problem',
'wtfiuwy': 'what the f**k is up with you',
'wtfiwwu': 'what the f**k is wrong with you',
'wtfiwwy': 'what the f**k is wrong with you',
'wtfiyp': 'what the f**k is your problem',
'wtfm': 'what the f**k, mate?',
'wtfmf': 'what the f**k mother f**ker',
'wtfo': 'what the f**k over',
'wtfru': 'what the f**k are you',
'wtfrud': 'what the f**k are you doing?',
'wtfrudng': 'what the f**k are you doing',
'wtfrudoin': 'what the f**k are you doing',
'wtfruo': 'what the f**k are you on?',
'wtfruttd': 'what the f**k are you trying to do',
'wtfs': 'what the f**king s**t?',
'wtfuah': 'what the f**k you a**h**e',
'wtful': 'what the f**k you loser',
'wtfwjd': 'what the f**k would jesus do',
'wtfwt': 'what the f**k was that',
'wtfwtd': 'what the f**k was that dude',
'wtfwtf': 'what the f**k was that for?',
'wtfwydt': 'why the f**k would you do that',
'wtfya': 'what the f**k you a**h**e',
'wtfyb': 'what the f**k you b***h',
'wtg': 'way to go',
'wtgds': 'way to go dumb s**t',
'wtgp': 'want to go private',
'wth': 'what the heck',
'wtharud': 'what the heck are you doing',
'wthau': 'who the hell are you',
'wthauwf': 'what the hell are you waiting for',
'wthay': 'who the hell are you',
'wthayd': 'what the heck are you doing',
'wthaydwmgf': 'what the hell are you doing with my girlfriend',
'wthdydt': 'why the hell did you do that',
'wthhyb': 'where the hell have you been?',
'wthigo': 'what the hell is going on',
'wthiwwu': 'what the hell is wrong with you',
'wtho': 'want to hang out?',
'wthru': 'who the heck are you',
'wthrud': 'what the hell are you doing?',
'wths': 'want to have sex',
'wthswm': 'want to have sex with me',
'wthwt': 'what the hell was that?',
'wthwut': 'what the hell were you thinking',
'wthyi': 'what the hell you idiot',
'wtii': 'what time is it',
'wtiiot': 'what time is it over there?',
'wtityb': 'whatever, tell it to your blog',
'wtly': 'welcome to last year',
'wtmf': 'what the mother f**k',
'wtmfh': 'what the mother f**king hell',
'wtmi': 'way too much information',
'wtmtr': "what's the matter",
'wtp': "where's the party",
'wtrud': 'what are you doing',
'wts': 'want to sell',
'wtt': 'want to trade',
'wttp': 'want to trade pictures?',
'wtv': 'whatever',
'wtva': 'whatever',
'wtvr': 'whatever',
'wtwm': 'what time are we meeting?',
'wtwr': 'well that was random',
'wu': "what's up?",
'wu2kilu': 'want you to know i love you',
'wub': 'love',
'wubmgf': 'will you be my girlfriend?',
'wubu2': 'what you been up to',
'wubut': 'what you been up too',
'wuciwug': 'what you see is what you get',
'wud': 'would',
'wudev': 'whatever',
'wudn': 'what you doing now',
'wuf': 'where are you from?',
'wufa': 'where you from again',
'wugowm': 'will you go out with me',
'wula': 'what you looking at?',
'wuld': 'would',
'wuny': 'wait until next year',
'wussup': 'what is up?',
'wut': 'what',
'wuta': 'what you talking about',
'wutb': 'what are you talking about',
'wutcha': 'what are you',
'wuteva': 'whatever',
'wutevr': 'what ever',
'wuts': 'what is',
'wutup': "what's up",
'wuu2': 'what you up to',
'wuu22m': 'what you up to tomorrow',
'wuut': 'what you up to',
'wuv': 'love',
'wuwh': 'wish you were here',
'wuwt': "what's up with that",
'wuwta': 'what do you want to talk about',
'wuwtab': 'what do you want to talk about',
'wuwtb': 'what do you want to talk about',
'wuwtta': 'what you want to talk about',
'wuwttb': 'what you want to talk about',
'wuwu': 'what up with you',
'wuz': 'was',
'wuza': "what's up",
'wuzup': "what's up",
'ww2': 'world war 2',
'wwc': 'who would care',
'wwcnd': 'what would <NAME> do',
'wwdhd': 'what would <NAME> do',
'wwe': 'world wrestling entertainment',
'wwgf': 'when we gonna f**k',
'wwhw': 'when where how why',
'wwikt': 'why would i know that',
'wwjd': 'what would jesus do?',
'wwt': 'what was that',
'wwtf': 'what was that for',
'wwudtm': 'what would you do to me',
'wwut': 'what were you thinking',
'www': 'world wide web',
'wwwu': 'whats wrong with you',
'wwwy': "what's wrong with you",
'wwy': 'where were you',
'wwycm': 'when will you call me',
'wwyd': 'what would you do?',
'wwyd2m': 'what would you do to me',
'wwygac': 'write when you get a chance',
'wwyt': 'what were you thinking',
'wy': 'why?',
'wyas': 'wow you are stupid',
'wyatb': 'wish you all the best',
'wyauimg': 'why you all up in my grill?',
'wyb': 'watch your back',
'wybts': 'were you born this sexy',
'wyc': 'will you come',
'wycm': 'will you call me',
'wyd': 'what are you doing',
'wyg': 'will you go',
'wygac': 'when you get a chance',
'wygam': 'when you get a minute',
'wygf': 'what you going for',
'wygowm': 'will you go out with me',
'wygwm': 'will you go with me',
'wyhi': 'would you hit it?',
'wyhswm': 'would you have sex with me',
'wylion': 'whether you like it or not',
'wyltk': "wouldn't you like to know",
'wylym': 'watch your language young man',
'wym': 'what you mean?',
'wyn': "what's your name",
'wyp': "what's your problem?",
'wypsu': 'will you please shut up',
'wys': "wow you're stupid",
'wysiayg': 'what you see is all you get',
'wysitwirl': 'what you see is totally worthless in real life',
'wysiwyg': 'what you see is what you get',
'wyut': 'what you up to',
'wyw': 'what you want',
'wywh': 'wish you were here',
'wywo': 'while you were out'}
x = {'x treme': 'extreme',
'xb36t': 'xbox 360',
'xbf': 'ex-boyfriend',
'xbl': 'xbox live',
'xcept': 'except',
'xcpt': 'except',
'xd': 'extreme droll',
'xellent': 'excellent',
'xfer': 'transfer',
'xgf': 'exgirlfriend',
'xing': 'crossing',
'xit': 'exit',
'xl': 'extra large',
'xlnt': 'excellent',
'xmas': 'christmas',
'xmpl': 'example',
'xoac': 'christ on a crutch',
'xor': 'hacker',
'xover': 'crossover',
'xox': 'hugs and kisses',
'xoxo': 'hugs and kisses',
'xp': 'experience',
'xpect': 'expect',
'xplaned': 'explained',
'xpt': 'except',
'xroads': 'crossroads',
'xs': 'excess',
'xtc': 'ecstasy',
'xtra': 'extra',
'xtreme': 'extreme',
'xyz': 'examine your zipper',
'xyzpdq': 'examine your zipper pretty darn quick'}
y = { 'y': 'why',
'y w': "you're welcome",
'y!a': 'yahoo answers',
"y'all": 'you all',
"y'all'd": 'you all would',
"y'all'd've": 'you all would have',
"y'all're": 'you all are',
"y'all've": 'you all have',
'y/n': 'yes or no',
'y/o': 'years old',
'y00': 'you',
'y2b': 'youtube',
'y2k': 'year 2000',
'ya': 'yeah',
'yaab': 'you are a b***h',
'yaaf': 'you are a f**',
'yaafm': 'you are a f**king moron',
'yaagf': 'you are a good friend',
'yaai': 'you are an idiot',
'yaf': "you're a f**",
'yafi': "you're a f**king idiot",
'yag': 'you are gay',
'yall': 'you all',
'yapa': 'yet another pointless acronym',
'yaqw': 'you are quite welcome',
'yarly': 'yeah really',
'yas': 'you are stupid',
'yasan': 'you are such a nerd',
'yasf': 'you are so funny',
'yasfg': 'you are so f**king gay',
'yasg': 'you are so gay',
'yasw': 'you are so weird',
'yatb': 'you are the best',
'yatwl': 'you are the weakest link',
'yaw': 'you are welcome',
'yayo': 'cocaine',
'ybbg': 'your brother by grace',
'ybs': "you'll be sorry",
'ybya': 'you bet your ass',
'ycliu': 'you could look it up',
'ycmtsu': "you can't make this s**t up",
'ycntu': 'why cant you?',
'yctwuw': 'you can think what you want',
'ydpos': 'you dumb piece of s**t',
'ydtm': "you're dead to me",
'ydufc': 'why do f**king care?',
'yduwtk': 'why do you want to know',
'ye': 'yes',
'yea': 'yes',
'yeh': 'yes',
'yep': 'yes',
'yer': "you're",
'yermom': 'your mother',
'yesh': 'yes',
'yessir': 'yes sir',
'yew': 'you',
'yfb': 'you f**king b*****d',
'yfg': "you're f**king gay",
'yfi': 'you f**king idiot',
'ygg': 'you go girl',
'ygm': 'you got mail',
'ygp': 'you got punked!',
'ygpm': "you got a private message",
'ygrr': 'you got rick rolled',
'ygtbfkm': "you got to be f**king kidding me",
'ygtbk': "you got to be kidding",
'ygtbkm': 'you got to be kidding me',
'ygtbsm': "you got to be shitting me",
'ygtsr': 'you got that s**t right',
'yh': 'yeah',
'yhbt': "you've been trolled",
'yhew': 'you',
'yhf': 'you have failed',
'yhgtbsm': 'you have got to be shitting me',
'yhl': 'you have lost',
'yhm': 'you have mail',
'yhni': 'you have no idea',
'yhpm': 'you have a private messge',
'yhtbt': 'you had to be there',
'yid': 'yes, i do',
'yim': 'yahoo instant messenger',
'yiwtgo': 'yes, i want to go private',
'yk': 'you kidding',
'yki': 'you know it',
'ykisa': 'your knight in shining armor',
'ykm': "you are killing me",
'ykn': 'you know nothing',
'ykw': 'you know what',
'ykwim': 'you know what i mean',
'ykwya': 'you know who you are',
'ykywywm': 'you know you wish you were me',
'ylb': 'you little b***h',
'ym': 'your mom',
'ymbg': 'you must be guessing',
'ymbkm': 'you must be kidding me',
'yme': 'why me',
'ymfp': 'your most favorite person',
'ymg2c': 'your mom goes to college',
'ymgtc': 'your mom goes to college',
'ymiaw': 'your mom is a w***e',
'ymislidi': 'you make it sound like i did it',
'ymmd': 'you made my day',
'ymmv': 'your mileage may vary',
'ymrasu': 'yes, my retarded ass signed up',
'yn': 'why not',
'yng': 'young',
'ynk': 'you never know',
'ynm': 'yes, no, maybe',
'ynt': 'why not',
'ynw': 'you know what',
'yo': 'year old',
"yo'": 'your',
'yodo': 'you only die once',
'yolo': 'you only live once',
'yolt': 'you only live twice',
'yomank': 'you owe me a new keyboard',
'yooh': 'you',
'yor': 'your',
"you'd": 'you would',
"you'd've": 'you would have',
"you'll": 'you will',
"you'll've": 'you will have',
"you're": 'you are',
"you've": 'you have',
'youngin': 'young person',
'yoy': 'why oh why',
'ypmf': 'you pissed me off',
'ypmo': 'you piss me off',
'ypom': 'your place or mine',
'yqw': "you are quite welcome",
'yr': 'year',
'yrbk': 'yearbook',
'yrbook': 'year book',
'yrms': 'you rock my socks',
'yrs': 'years',
'yrsaf': 'you are such a fool',
'yrsm': 'you really scare me',
'yrss': 'you are so sexy',
'yru': 'why are you?',
'yrubm': 'why are you bugging me?',
'yrusm': 'why are you so mean',
'ys': 'you suck',
'ysa': 'you suck ass',
'ysal': 'you suck at life',
'ysati': 'you suck at the internet',
'ysf': 'you stupid f**k',
'ysic': 'why should i care',
'ysitm': 'your shirt is too small',
'ysk': 'you should know',
'ysm': 'you scare me',
'ysoab': 'you son of a b***h',
'yss': 'you stupid s**t',
'ystrdy': 'yesterday',
'yswnt': 'why sleep when not tired?',
'yt': 'you there?',
'ytd': 'year to date',
'ytf': 'why the f**k',
'ytfwudt': 'why the f**k would you do that?',
'ythwudt': 'why the hell would you do that',
'ytis': "you think i am special?",
'ytm': 'you tell me',
'ytmnd': "you are the man now, dog!",
'yty': 'why thank you',
'yu': 'you',
'yua': 'you ugly ass',
'yuo': 'you',
'yup': 'yes',
'yur': 'your',
'yust': 'why you say that',
'yvfw': "you are very f**king welcome",
'yvw': "you are very welcome",
'yw': "you are welcome",
'ywapom': 'you want a piece of me?',
'ywia': "you are welcome in advance",
'ywic': 'why would i care',
'yws': 'you want sex',
'ywsyls': 'you win some you lose some',
'ywud': 'yo what is up dude',
'ywvm': "you are welcome very much",
'ywywm': 'you wish you were me',
'yysw': 'yeah, yeah, sure, whatever'}
z = {"z'omg": 'oh my god',
'z0mg': 'oh my god',
'zex': 'sex',
'zh': 'zero hour',
'zig': 'cigarette',
'zomfg': 'oh my f**king god',
'zomg': 'oh my god',
'zomgzorrz': 'oh my god',
'zoot': 'woohoo',
'zot': 'zero tolerance',
'zt': 'zoo tycoon',
'zup': "what is up?"}
# +
# mylis = sorted(z.items())
# print(len(mylis))
# mydict = {}
# for k, v in mylis:
# mydict[k.lower()] = v.lower()
# print(len(mydict))
# mydict
# -
slang = pd.DataFrame([ a, b, c, d, e, f, g, h, i, j, k, l, m, n, o, p, q, r, s, t, u, v, w, x, y, z], index = list('abcdefghijklmnopqrstuvwxyz'))
slang.head()
# +
# for i in CONTRACTION_MAP.keys():
# if i not in list(slang.columns):
# print(i)
# else:
# print("***************",i)
# -
slang.shape
slang_list = list(slang.columns)
# +
# for i in mytext1:
# for j in i.split():
# if j.lower() in slang_list:
# print(j , " ",slang[j.lower()].loc[j[0].lower()])
# else:
# continue
# print(i)
# print()
# +
# for text in mytext1:
# import re
# print(text)
# pattern = re.compile(r'(\w)(\1{2,})')
# text = re.sub(r'(\w)(\1{2,})', r"\1", text)
# print(text)
# +
# # re.sub(r"(\w)\1*", r'\1', 'sleeeeepy')
# for text in mytext2:
# pat = re.compile(r"(\w)\1*")
# res = pat.findall(text)
# print(''.join(res))
# +
# mylis = sorted(a.items())
# mydict = {}
# for k, v in mylis:
# mydict[k.lower()] = v.lower()
# mydict
# -
slang['luk'].loc['l']
# + active=""
# 'iaspfm' :'i am sorry please forgive me'
#
# Dang euphemism for damn.
# TMH Touch My Hand
#
# fml f**k my life
#
#
# p.s. postscript
#
# yg <NAME>
#
#
# SF San Franscisco
# BMFing 'bad m****r f*****g
#
# cuss 'hell
#
# LMFAO "laughing my fucking ass off"
#
#
#
#
#
# woulda "would have"
#
# deffo Definitely
#
# LMAO "laughing my ass off"
# Smh shaking my head
# l'm
#
# MCFLY
# mcfly
# gig
# l'm
# boi!
# NKOTB
# gn'r
# lmao
# jkin
# (dsl digital satellite link)
# F!
# (th@ though)
# # ok
# ACSM
# 'iaspfm': ''
#
# -
slang.to_csv('slang.csv')
# +
sounds = ['aaaaaah', 'aha', 'ahem', 'ahh', 'ahhh', 'alas', 'argh', 'augh', 'aw', 'aww', 'awww', 'awwwww', 'bah', 'bam', 'blah', 'boo', 'booh', 'boohoo', 'booo', 'boooo', 'brr', 'brrrr', 'doh', 'duh', 'eeeek', 'eek', 'eep', 'egh', 'eh', 'ehh', 'ehhh', 'ehhhh', 'eww', 'fuff', 'gah', 'gahhh', 'gee', 'grr', 'grrrr', 'hah', 'haha', 'hahaa', 'hahaha', 'hahahahaha', 'harumph', 'hay', 'heh', 'hm', 'hmm', 'hmmm', 'hmmmm', 'hmmmmm', 'hoooray', 'huh', 'humph', 'hun', 'hurrah', 'ick', 'jeez', 'meh', 'mhm', 'mm', 'muahaha', 'mwah', 'naaaaah', 'naaaah', 'naaah', 'naah', 'nah', 'nup', 'oh', 'ohh', 'ohhh', 'ohhhh', 'ohoh', 'ooh', 'oomph', 'oooh', 'oooops', 'ooops', 'oops', 'ouch', 'oww', 'owww', 'owwww', 'oy', 'pew', 'pff', 'phew', 'poo', 'pooo', 'poooo', 'pooooo', 'psst', 'sheesh', 'shh', 'shoo', 'ugh', 'uh', 'uhh', 'um', 'umm', 'umph', 'waaaaah', 'waah', 'wee', 'whoa', 'whoo', 'whooo', 'whoooo', 'whooooo', 'woof', 'wow', 'yahoo', 'yay', 'yeah', 'yeehaw', 'yikes', 'yippee', 'yoohoo', 'yuck', 'zing']
# -
print(sounds)
text = re.sub(r'\b([a-z]+)\s+\1{1,}\b', r"\1", " ".join(sounds))
print(text)
slang["'"]
# Checking words
mylis = "back from aot last lec.. alot of question marks on my head.. but thank god for all the tips.. should be able to pull tru".split()
mylis
new = []
for i in mylis:
if isinstance(i, str):
if i in slang_columns:
new.append(slang[i].loc[i[0]])
else:
new.append(i)
else:
continue
print(' '.join(new))
| Chatting_Slang_Converter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 9.2
# language: sage
# name: sagemath
# ---
# +
def transmissionProcess(N,initialN=0,lambda1=1,lambda2=0,lambda3=0,p_b = 0,plt = 0,z_end=0):
#initialisation of SIRSCN
C = N.copy()
z = 0 # infection event count
n = C.order() #total number of nodes
infectedNs = [initialN]
recoveredNs = []
n_i = [1]
n_s = [n-1]
n_r = [0] #put number of SIR in lists for easy plotting later on
C.set_vertex(initialN,'i')
lex = {e:[0,0] for e in range(n)} #lexicon. [#how many nodes infected,#times infected]
lex[initialN] = [0,1]
currentSOE = susceptibleOutEdges(C,infectedNs)
if p_b > 0:
for e in currentSOE:
if random() < p_b:
C.set_edge_label(e[0],e[1],'b')
currentSOE.remove(e)
# initialisation of Transmission Tree
pathsDict={} # dictionary of nodes -> paths from root in tree
LBT = LabelledBinaryTree
# individuals in tree are labelled by "n"+str(integer_label)
T = LBT([None,None],label="n"+str(initialN)).clone()
pathsDict["n"+str(initialN)]=[]
while (n_i[-1] > 0 and lambda2 > 0) or (n_s[-1] > 0 and lambda2 == 0):
z=z+1
p = random()
m = len(currentSOE)
p_inf = m*lambda1/(m*lambda1+n_i[-1]*lambda2+n_r[-1]*lambda3) #s -> i
p_rec = n_i[-1]*lambda2/(len(currentSOE)*lambda1+n_i[-1]*lambda2+n_r[-1]*lambda3) #i -> r
if 0 < p < p_inf: #infection event
next_edge = currentSOE.pop(randrange(0,m)) #find next edge
infectedNs.append(next_edge[1])
C.set_vertex(next_edge[1],'i')
currentSOE = [e for e in currentSOE if e[1] != next_edge[1]] #remove edges leading to this new infected node
new_SOE = susceptibleOutEdges(C,next_edge[1])
if p_b > 0:
for e in C.outgoing_edge_iterator(next_edge[1]):
if e[2] == 'b':
C.set_edge_label(e[0],e[1],None)
if random() < p_b:#*n_i[-1]/n:
C.set_edge_label(e[0],e[1],'b')
if e in new_SOE:
new_SOE.remove(e)
currentSOE = currentSOE+ new_SOE #add new susceptible edges
n_s.append(n_s[-1]-1)
n_i.append(n_i[-1]+1)
n_r.append(n_r[-1])
T=growTransmissionTree(T, pathsDict, z, "n"+str(next_edge[0]),"n"+str(next_edge[1]))
lex[next_edge[0]][0] += 1 #add stuff to lexicon
lex[next_edge[1]][1] += 1
elif p_inf < p < p_inf+p_rec: #recovery event
next_node = infectedNs.pop(randrange(0,len(infectedNs)))
currentSOE = [e for e in currentSOE if e[0] != next_node] #remove edges coming from this node
if lambda3 == -1: #skip recovery, go straight to susceptible
C.set_vertex(next_node,None)
currentSOE = currentSOE+infectedInEdges(C,next_node)
n_i.append(n_i[-1]-1)
n_r.append(n_r[-1])
n_s.append(n_s[-1]+1)
else:
recoveredNs.append(next_node)
C.set_vertex(next_node,'r')
n_i.append(n_i[-1]-1)
n_s.append(n_s[-1])
n_r.append(n_r[-1]+1)
else:
next_node = recoveredNs.pop(randrange(0,len(recoveredNs)))
C.set_vertex(next_node,None)
currentSOE = currentSOE+infectedInEdges(C,next_node)
n_i.append(n_i[-1])
n_r.append(n_r[-1]-1)
n_s.append(n_s[-1]+1)
#print(ascii_art(T))
#print "step z = ",z; print ascii_art(T); print "--------------------"
if (z_end != 0) and (z == z_end):
print(f"{z_end} events, stopping simulation.")
converge = 0
break
if n_i[-1] == 0:
converge = 1
if plt == 1:
iplotvalues = [(i,e) for i,e in enumerate(n_i)]
splotvalues = [(i,e) for i,e in enumerate(n_s)]
iplot = line(iplotvalues, color='red', legend_label='Infected', legend_color='red',axes_labels=['Event','Count'])
splot = line(splotvalues, color='green', legend_label='Susceptible', legend_color='green',axes_labels=['Event','Count'])
plotter = splot+iplot
if (lambda2 > 0) and (lambda3 != -1):
rplotvalues = [(i,e) for i,e in enumerate(n_r)]
rplot = line(rplotvalues, color='blue', legend_label='Recovered', legend_color='blue',axes_labels=['Event','Count'])
plotter = plotter + rplot
show(plotter)
#print("Parameters: lambda1 = "+str(lambda1)+", lambda2 = "+str(lambda2)+", lambda3 = "+str(lambda3)+", p_block = "+str(RR(p_b)))
return [[n_s,n_i,n_r,lex,z,converge],T.as_ordered_tree(with_leaves=False)]
def CountsDict(X):
'''convert a list X into a Dictionary of counts or frequencies'''
CD = {}
for x in X:
CD[x] = (CD[x] + 1) if (x in CD) else 1
return CD
def susceptibleOutEdges(C,vs):
'''return the the susceptible outedges of node v in vs in SICN C'''
SOE = [e for e in C.outgoing_edge_iterator(vs) if C.get_vertex(e[1])==None]
return SOE
def infectedInEdges(C,vs):
'''return the the infected inedges of node v in vs in SICN C'''
SOE = [e for e in C.incoming_edge_iterator(vs) if (C.get_vertex(e[0])=='i') and (e[2] !='b')]
return SOE
def growTransmissionTree(Ttree, pDict, z, infector, infectee):
'''grow the transmission tree Ttree and update pathsDict pDict by adding the
z-th infection event with infector -> infectee '''
LBT = LabelledBinaryTree
newSubTree = LBT([LBT([None,None], label=infector), LBT([None, None], label=infectee)], label=z).clone()
path2Infector = pDict[infector]
if z==1:
Ttree = newSubTree
else:
Ttree[tuple(path2Infector)] =newSubTree
#print ascii_art(Ttree)
pDict[infector]=path2Infector+[0]
pDict[infectee]=path2Infector+[1]
pDict[z]=path2Infector
return Ttree
def SIS(C,lambda1=1,lambda2=1,z_end=5000,plt=0):
return transmissionProcess(C,0,lambda1,lambda2,-1,0, plt,z_end)
def SIR(C,lambda1=1,lambda2=1,plt=0):
return transmissionProcess(C,0,lambda1,lambda2,0,0,plt,z_end=0)
def SIRS(C,lambda1=1,lambda2=1,lambda3=1,z_end = 3000,plt=0,p_b=0):
return transmissionProcess(C,0,lambda1,lambda2,lambda3,p_b,plt,z_end)
# +
#SIS simulations
#Small World
from sage.plot.density_plot import DensityPlot
n = 300
k0 = n/10
p = 0.4
k = graphs.RandomNewmanWattsStrogatz(n, k0,p).to_directed()
t = transmissionProcess(k,0,1,15,-1,0,1,15000)
l = t[0][3]
k2 = list(l.values())
node_lex_ic = {}
for i in range(n):
node_lex_ic[i] = k.out_degree(i)
node_lex2_ic = {}
for i,e in enumerate(node_lex_ic.values()):
node_lex2_ic[e] = (node_lex2_ic[e] + [i]) if (e in node_lex2_ic) else [i]
ic_lex = {}
for i,e in enumerate(node_lex2_ic):
ic_lex[e] = [l[y][0] for y in node_lex2_ic[e]]
plot_values_ic = []
for i in ic_lex:
for j in ic_lex[i]:
plot_values_ic.append((i,j))
plot_mean_ic = sorted([(e,mean(ic_lex[e])) for e in ic_lex])
pl = line(plot_mean_ic,color='red',axes_labels=["Neighbours","Infector frequency"])
pl2 = points(plot_values_ic,color='blue')
show(pl+pl2)
node_lex_ti = {}
for i in range(n):
node_lex_ti[i] = k.in_degree(i)
node_lex2_ti = {}
for i,e in enumerate(node_lex_ti.values()):
node_lex2_ti[e] = (node_lex2_ti[e] + [i]) if (e in node_lex2_ti) else [i]
ti_lex = {}
for i,e in enumerate(node_lex2_ti):
ti_lex[e] = [l[y][1] for y in node_lex2_ti[e]]
plot_values_ti = []
for i in ti_lex:
for j in ti_lex[i]:
plot_values_ti.append((i,j))
plot_mean_ti = sorted([(e,mean(ti_lex[e])) for e in ti_lex])
pl = line(plot_mean_ti,color='red',axes_labels=["Neighbours","Infectee frequency"])
pl2 = points(plot_values_ti,color='blue')
show(pl+pl2)
# -
#SIS simulations
#Star
n = 30
k = graphs.StarGraph(n).to_directed()
t = transmissionProcess(k,0,3,1,-1,0,1,5000)
l = t[0][3]
k = [l[i] for i in range(len(l)) if i != 0]
infected = [e[0] for e in k]
been_inf = [e[1] for e in k]
plotvalues1 = [(i,infected[i]) for i in range(len(infected))]
plotvalues2 = [(i,been_inf[i]) for i in range(len(been_inf))]
plot1 = line(plotvalues1, color = 'red',legend_label='Infector', legend_color='red',axes_labels=['Node label','Frequency'])
plot2 = line(plotvalues2, color = 'green',legend_label='Infectee', legend_color='green',axes_labels=['Node label','Frequency'])
show(plot1+plot2)
print('infected nodes, mean ',RR(mean(t[0][1])))
print('infected nodes, std ',RR(std(t[0][1])))
print('susceptible nodes, mean ',RR(mean(t[0][0])))
print('susceptible nodes, std ',RR(std(t[0][0])))
print('Infected per node, mean: ',RR(mean(infected)))
print('Infected per node, std: ',RR(std(infected)))
print('times been infected, mean: ',RR(mean(been_inf)))
print('times been infected, std: ',RR(std(been_inf)))
print('Star node: ',l[0])
print("z = "+str(t[0][4]))
#SIS simulations
#path
n = 30
k = graphs.PathGraph(n).to_directed()
t = SIS(k,3,1,15000,1)
l = t[0][3]
k = [l[i] for i in range(len(l))]
infected = [e[0] for e in k]
been_inf = [e[1] for e in k]
plotvalues1 = [(i,infected[i]) for i in range(len(infected))]
plotvalues2 = [(i,been_inf[i]) for i in range(len(been_inf))]
plot1 = line(plotvalues1, color = 'red',legend_label='Infector', legend_color='red',axes_labels=['Node label','Frequency'])
plot2 = line(plotvalues2, color = 'green',legend_label='Infectee', legend_color='green',axes_labels=['Node label','Frequency'])
show(plot1+plot2)
print('infected nodes, mean ',RR(mean(t[0][1])))
print('infected nodes, std ',RR(std(t[0][1])))
print('susceptible nodes, mean ',RR(mean(t[0][0])))
print('susceptible nodes, std ',RR(std(t[0][0])))
print('Infected per node, mean: ',RR(mean(infected)))
print('Infected per node, std: ',RR(std(infected)))
print('times been infected, mean: ',RR(mean(been_inf)))
print('times been infected, std: ',RR(std(been_inf)))
print("z = "+str(t[0][4]))
#SIS simulations
#complete
n = 30
k = graphs.CompleteGraph(n).to_directed()
t = transmissionProcess(k,0,1,5,-1,0,1,15000)
l = t[0][3]
k = [l[i] for i in range(len(l))]
infected = [e[0] for e in k]
been_inf = [e[1] for e in k]
plotvalues1 = [(i,infected[i]) for i in range(len(infected))]
plotvalues2 = [(i,been_inf[i]) for i in range(len(been_inf))]
plot1 = line(plotvalues1, color = 'red',legend_label='Infector', legend_color='red',axes_labels=['Node label','Frequency'])
plot2 = line(plotvalues2, color = 'green',legend_label='Infectee', legend_color='green',axes_labels=['Node label','Frequency'])
show(plot1+plot2)
print('infected nodes, mean ',RR(mean(t[0][1])))
print('infected nodes, std ',RR(std(t[0][1])))
print('susceptible nodes, mean ',RR(mean(t[0][0])))
print('susceptible nodes, std ',RR(std(t[0][0])))
print('Infected per node, mean: ',RR(mean(infected)))
print('Infected per node, std: ',RR(std(infected)))
print('times been infected, mean: ',RR(mean(been_inf)))
print('times been infected, std: ',RR(std(been_inf)))
print("z = "+str(t[0][4]))
#SIS simulations
#complete
n = 30
k = graphs.CompleteGraph(n).to_directed()
t = transmissionProcess(k,0,1,5,-1,0,1,40)
l = t[0][3]
k = [l[i] for i in range(len(l))]
infected = [e[0] for e in k]
been_inf = [e[1] for e in k]
plotvalues1 = [(i,e) for i,e in infected]
plotvalues2 = [(i,e) for i,e in been_inf]
plot1 = line(plotvalues1, color = 'red',legend_label='Infector', legend_color='red',axes_labels=['Node label','Frequency'])
plot2 = line(plotvalues2, color = 'green',legend_label='Infectee', legend_color='green',axes_labels=['Node label','Frequency'])
show(plot1+plot2)
print('infected nodes, mean ',RR(mean(t[0][1])))
print('infected nodes, std ',RR(std(t[0][1])))
print('susceptible nodes, mean ',RR(mean(t[0][0])))
print('susceptible nodes, std ',RR(std(t[0][0])))
print('Infected per node, mean: ',RR(mean(infected)))
print('Infected per node, std: ',RR(std(infected)))
print('times been infected, mean: ',RR(mean(been_inf)))
print('times been infected, std: ',RR(std(been_inf)))
print("z = "+str(t[0][4]))
# +
#SIR simulations
#complete
n = 30
m = 1000
lambda1 = 1
lambda2 = 6
plotvalues = []
lambdas = [5,10,15,20,25]
for i in lambdas:
gr = graphs.CompleteGraph(n).to_directed()
t = [transmissionProcess(gr,0,lambda1,i,0,0) for _ in range(m)]
#t = [SIR(gr,lambda1,i) for _ in range(m)]
l = [n-e[0][0][-1] for e in t]
k = CountsDict(l)
plotvalues.append(sorted([(e,k[e]) for e in k]))
print(i)
print(plotvalues[-1][0])
print(plotvalues[-1][1])
colorss = ['blue','green','orange','purple','yellow']
plot1 = line(plotvalues.pop(0), color = 'red',marker = 'o',markersize = 2,legend_label=f'lambda2 = {lambdas.pop(0)}',axes_labels=['Recovered nodes','Frequency'])
for i,e in enumerate(plotvalues):
plot1 += line(e, color = colorss[i],marker = 'o',markersize = 2,legend_label=f'lambda2 = {lambdas[i]}',axes_labels=['Recovered nodes','Frequency'])
show(plot1)
# +
#SIR simulations
#star
n = 30
m = 1000
lambda1 = 1
lambda2 = 6
plotvalues = []
lambdas = [1/2,1,5,10]
for i in lambdas:
gr = graphs.StarGraph(n).to_directed()
t = [transmissionProcess(gr,0,lambda1,i,0,0,0) for _ in range(m)]
l = [n-e[0][0][-1] for e in t]
k = CountsDict(l)
plotvalues.append(sorted([(e+1,k[e]) for e in k]))
print(i)
print(plotvalues[-1][0])
print(plotvalues[-1][1])
colorss = ['blue','green','orange']
plot1 = line(plotvalues.pop(0), color = 'red',marker = 'o',markersize = 2,legend_label=f'lambda2 = {lambdas.pop(0)}',axes_labels=['Recovered nodes','Frequency'])
for i,e in enumerate(plotvalues):
plot1 += line(e, color = colorss[i],marker = 'o',markersize = 2,legend_label=f'lambda2 = {lambdas[i]}',axes_labels=['Recovered nodes','Frequency'])
show(plot1)
# +
#SIR simulations
#path
n = 30
m = 1000
lambda1 = 1
lambda2 = 6
plotvalues = []
lambdas = [1/16,1/8,1/3,1]
for i in lambdas:
gr = graphs.PathGraph(n).to_directed()
t = [transmissionProcess(gr,0,lambda1,i,0,0,0) for _ in range(m)]
l = [n-e[0][0][-1] for e in t]
k = CountsDict(l)
plotvalues.append(sorted([(e,k[e]) for e in k]))
print(i)
print(plotvalues[-1][0])
print(plotvalues[-1][1])
colorss = ['blue','green','orange']
plot1 = line(plotvalues.pop(0), color = 'red',marker = 'o',markersize = 2,legend_label=f'lambda2 = {lambdas.pop(0)}',axes_labels=['Recovered nodes','Frequency'])
for i,e in enumerate(plotvalues):
plot1 += line(e, color = colorss[i],marker = 'o',markersize = 2,legend_label=f'lambda2 = {lambdas[i]}',axes_labels=['Recovered nodes','Frequency'])
show(plot1)
# +
#star probs
print(RR(0.5/(30+0.5)*1000))
print(RR(30/(30+1/2)*0.5/(29+1)*(1+0.5/(29.5))*1000))
print(RR(1/(30+1)*1000))
print(RR(30/(30+1)*1/(29+2)*(1+1/(30))*1000))
print(RR(5/(30+5)*1000))
print(RR(30/(30+5)*5/(29+10)*(1+5/(34))*1000))
print(RR(10/(30+10)*1000))
print(RR(30/(30+10)*10/(29+20)*(1+10/(39))*1000))
print('')
#complete probs
print(RR(5/(29+5)*1000))
print(RR(2*29/(29+5)*(5/(56+10)*5/(28+5))*1000))
print(RR(10/(29+10)*1000))
print(RR(2*29/(29+10)*(10/(56+20)*10/(28+10))*1000))
print(RR(15/(29+15)*1000))
print(RR(2*29/(29+15)*(15/(56+30)*15/(28+15))*1000))
print(RR(20/(29+20)*1000))
print(RR(2*29/(29+20)*(20/(56+40)*20/(28+20))*1000))
print(RR(25/(29+25)*1000))
print(RR(2*29/(29+25)*(25/(56+50)*25/(28+25))*1000))
print('')
#path probs
print(RR(1/(1+1)*1000))
print(RR(1/(1+1)*1/(1+2)*(1+1/(1+1))*1000))
print(RR(1/3/(1+1/3)*1000))
print(RR(1/(1+1/3)*1/3/(1+2/3)*(1+1/3/(1+1/3))*1000))
print(RR(1/8/(1+1/8)*1000))
print(RR(1/(1+1/8)*1/8/(1+2/8)*(1+1/8/(1+1/8))*1000))
print(RR(1/16/(1+1/16)*1000))
print(RR(1/(1+1/16)*1/16/(1+2/16)*(1+1/16/(1+1/16))*1000))
# -
#presentation sims
#SIRS simulations
#complete
n = 300
k = graphs.CompleteGraph(n).to_directed()
t = SIRS(k,1,5,0.01,plt=1) #kinda SIR
t = SIRS(k,1,50,10000,plt=1) #kinda SIS
t = SIRS(k,1,2,1,plt=1,p_b=0.9) #p block
#t = SIRS(k,1,1,2,plt=1,p_b=4,z_end=15000) #p block, remember to change p_b and n=300
# +
#SIS simulations
#Small World
import matplotlib.pyplot as plt
n = 300
k0 = n/10
p = 0.4
k = graphs.RandomNewmanWattsStrogatz(n, k0,p).to_directed()
t = transmissionProcess(k,0,1,15,-1,0,1,15000)
l = t[0][3]
k2 = list(l.values())
node_lex_ic = {}
for i in range(n):
node_lex_ic[i] = k.out_degree(i)
node_lex2_ic = {}
for i,e in enumerate(node_lex_ic.values()):
node_lex2_ic[e] = (node_lex2_ic[e] + [i]) if (e in node_lex2_ic) else [i]
ic_lex = {}
for i,e in enumerate(node_lex2_ic):
ic_lex[e] = [l[y][0] for y in node_lex2_ic[e]]
plot_values_ic = []
for i in ic_lex:
for j in ic_lex[i]:
plot_values_ic.append([i,j])
plot_mean_ic = sorted([(e,mean(ic_lex[e])) for e in ic_lex])
pl = line(plot_mean_ic,color='red',axes_labels=["Neighbours","Infector frequency"])
pl2 = plt.hist2d([e[0] for e in plot_values_ic],[e[1] for e in plot_values_ic],bins=(50,50),cmap=plt.cm.BuPu)
plt.show()
"""
node_lex_ti = {}
for i in range(n):
node_lex_ti[i] = k.in_degree(i)
node_lex2_ti = {}
for i,e in enumerate(node_lex_ti.values()):
node_lex2_ti[e] = (node_lex2_ti[e] + [i]) if (e in node_lex2_ti) else [i]
ti_lex = {}
for i,e in enumerate(node_lex2_ti):
ti_lex[e] = [l[y][1] for y in node_lex2_ti[e]]
plot_values_ti = []
for i in ti_lex:
for j in ti_lex[i]:
plot_values_ti.append((i,j))
plot_mean_ti = sorted([(e,mean(ti_lex[e])) for e in ti_lex])
pl = line(plot_mean_ti,color='red',axes_labels=["Neighbours","Infectee frequency"])
pl2 = points(plot_values_ti,color='blue')
show(pl+pl2)
"""
# -
pl2 = plt.hist2d([e[0] for e in plot_values_ic],[e[1] for e in plot_values_ic],bins=(100,100),cmap=plt.cm.BuPu)
plt.show()
DensityPlot?
| codes/finalTransmission.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "Using Python3's ‘venv’ with tox"
# > "Using the built-in Python module in favor of virtualenv to create your testing or project automation environments."
# - author: jhermann
# - toc: false
# - branch: master
# - badges: true
# - comments: true
# - published: true
# - categories: [python, testing]
# - image: images/copied_from_nb/img/python/tox-venv.png
# 
# `tox` is a generic virtualenv management and test command line tool, especially useful for multi-environment testing. It has a plugin architecture, with plenty of both built-in and 3rd party extensions.
#
# This post assumes you are already familiar with `tox` and have a working configuration for it. If not, check out [its documentation](https://tox.readthedocs.io/).
# In order to make `tox` use the built-in virtual environment `venv` of Python 3.3+, there is a plugin named [tox-venv](https://pypi.org/project/tox-venv/) that switches from using `virtualenv` to `venv` whenever it is available.
#
# Typically, `venv` is more robust when faced with ever-changing runtime environments and versions of related tooling (`pip`, `setuptools`, …).
# To enable that plugin, add this to your `tox.ini`:
#
# ```ini
# [tox]
# requires = tox-venv
# ```
#
# That merely triggers `tox` to check (on startup) that the plugin is installed. You still have to add it to your `dev-requirements.txt` or a similar file, so it gets installed together with `tox`. You can also install `tox` globally using `dephell jail install tox tox-venv` – see the [related post](https://jhermann.github.io/blog/python/deployment/2020/03/03/install_tools_with_dephell.html) in this blog for details.
#
# The end result is this (call `tox -v` to see those messages):
#
# ```
# py38 create: …/.tox/py38
# …/.tox$ /usr/bin/python3.8 -m venv py38 >…/log/py38-0.log
# ```
#
# And there you have it, no more virtualenv package needed. 🎉 🎊
| _notebooks/2020-03-21-tox_venv.ipynb |
;; -*- coding: utf-8 -*-
;; ---
;; jupyter:
;; jupytext:
;; text_representation:
;; extension: .scm
;; format_name: light
;; format_version: '1.5'
;; jupytext_version: 1.14.4
;; kernelspec:
;; display_name: Calysto Scheme 3
;; language: scheme
;; name: calysto_scheme
;; ---
;; ### 練習問題2.60
;; 上の例では、集合は重複のないリストとして表現するよう規定した。
;; ここで、重複を許す場合について考えてみよう。
;; その場合、例えば{1,2,3}という集合は(2 3 2 1 3 2 2)というリストとして表現することもできる。
;; この表現に対して演算を⾏う⼿続きelement-of-set?,adjoin-set,union-set,intersectionsetを設計せよ。
;; それぞれの効率は、重複なし表現に対する⼿続きでそれに対応するものと⽐べてどうだろうか。
;; 重複なしの表現よりもこの表現のほうが向いているような応⽤はあるだろうか。
;; +
; 変更なし
(define (element-of-set? x set)
(cond ((null? set) #f)
((equal? x (car set)) #t)
(else (element-of-set? x (cdr set))))
)
; 変更
(define (adjoin-set x set)
(cons x set)
)
; 変更なし
(define (intersection-set set1 set2)
(cond ((or (null? set1) (null? set2)) '())
((element-of-set? (car set1) set2) (cons (car set1) (intersection-set (cdr set1) set2)))
(else (intersection-set (cdr set1) set2)))
)
; 変更
(define (union-set set1 set2)
(cond ((and (null? set1) (null? set2)) '())
((null? set1) set2)
((null? set2) set1)
;(else (cons (car set1) (union-set (cdr set1) set2)))
;(else (union-set (cdr set1) (adjoin-set (car set1) set2)))
(else (append set1 set2))
)
)
;; +
; 動作確認
(define A (list 1 2 3 4))
(define B (list 2 4 5 6))
(define C (list 4 6 7))
(union-set A B)
;; -
(union-set (union-set A B) C)
(adjoin-set 3 B)
;; 重複なしの表現よりもこの表現のほうが向いているような応⽤はあるだろうか。
;;
;; →リストの要素数が増えるので、element-of-set?のステップ数は増大するので、element-of-set?を多用するシステムは不利と思われる。。
;; union-setはステップ数が減るはずなので、union-setを多用するシステムは有利と考えられる。
| exercises/2.60.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
from wikidata2df import wikidata2df
query = """
SELECT ?item ?itemLabel ?countryLabel
WHERE
{
?item p:P31 ?statement.
?statement ps:P31 wd:Q3241045.
?statement pq:P642 wd:Q84263196.
?statement pq:P3005 ?country.
VALUES ?countryTypes {wd:Q6256 wd:Q3624078 wd:Q1048835}.
?country wdt:P31 ?countryTypes.
SERVICE wikibase:label { bd:serviceParam wikibase:language "[AUTO_LANGUAGE],en". }
}
"""
local_outbreak_items = wikidata2df(query)
# -
local_outbreak_items
# +
manual_matches = pd.DataFrame(data={'countryLabel':['Kosovo', 'Liberia', 'State of Palestine'],
'itemLabel':["COVID-19 pandemic in Kosovo","2020 COVID-19 pandemic in Liberia","2020 COVID-19 pandemic in the State of Palestine"],
'item': ["Q87655119","Q87766242","Q87199320"]
})
# -
local_outbreak_items.append(manual_matches)
local_outbreak_items.append(manual_matches).drop_duplicates().to_csv("reference.csv", index=False)
| wikipedia_case_import/create_ref_table.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #이미지 증강기법을 적용한 결과 보기
import cv2
import matplotlib.pyplot as plt
import os
find_list=os.listdir('C:/Users/gptjd/OneDrive/Desktop/label')
print(find_list)
# +
class_label={'Chair':'0','Light bulb':'1','Window':'2','Person':'3',
'Door':'4','Picture frame':'5','Book':'6','Table':'7',
'Drawer':'8','Houseplant':'9'}
class_label={v:k for k,v in class_label.items()}
IMAGE_PATH='C:/Users/gptjd/OneDrive/Desktop/OIDv4_ToolKit/OID/Dataset/train/images/images'
LABEL_PATH='C:/Users/gptjd/OneDrive/Desktop/label'
for label_ in find_list:
label_path=os.path.join(LABEL_PATH,label_)
image_path=os.path.join(IMAGE_PATH,label_.replace('.txt','.jpg'))
image=cv2.imread(image_path)
f=open(label_path,'r')
list_box=[]
while 1:
line=f.readline()
if not line:
break
list_box.append(line.split())
f.close()
imW,imH=image.shape[:2]
f=open(LABEL_PATH+'/'+label_)
for box in list_box:
label=class_label.get(box[0])
x1=int(float(box[1]))
y1=int(float(box[2]))
x2=int(float(box[3]))
y2=int(float(box[4]))
cv2.rectangle(image,(x1,y1),(x2,y2),(255,0,0),thickness=3)
cv2.putText(image,label,(int((x1+x2)/2),int((y1+y2)/2)),cv2.FONT_HERSHEY_SIMPLEX, 1, (0,255), 2)
SAVE_PATH='C:/Users/gptjd/OneDrive/Desktop/OIDv4_ToolKit/OID/Dataset/train/images/check/'
SAVE_PATH=SAVE_PATH+label_.replace('.txt','.jpg')
cv2.imwrite(SAVE_PATH,image)
# -
| show_result.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## With dependencies, keys loaded and function defined, all we need is to call the function while give it the target_user (account) and page number
# ### What we will get is: 1) print out to notebook telling how many tweets pulled and what time period covers; 2) a named CSV file generated in your work directory
# ### Remember to modify this template and use your own Twitter API keys, because the information is not there. For each Twitter account, we can play around with the page number and see whether we can pull more tweets.
# +
# Dependencies
import tweepy
import numpy as np
import pandas as pd
from datetime import datetime
import time
from pprint import pprint
# +
# Import and Initialize Sentiment Analyzer
from vaderSentiment.vaderSentiment import SentimentIntensityAnalyzer
analyzer = SentimentIntensityAnalyzer()
# Twitter API Keys
from TwitterbotKeys import *
# Setup Tweepy API Authentication
auth = tweepy.OAuthHandler(consumer_key, consumer_secret)
auth.set_access_token(access_token, access_token_secret)
api = tweepy.API(auth, parser=tweepy.parsers.JSONParser())
# -
# define a function to pull data from twitter account and generate CSV file
def pull_tweets_to_csv(target_user, pages):
# part1: get as many tweets as we can
tweets_list = []
for x in range(pages):
response = api.user_timeline(target_user, page=x)
for tweet in response:
tweets_list.append(tweet)
print(f"We have {len(tweets_list)} tweets from account: {target_user}.")
start = datetime.strptime(tweets_list[0]['created_at'], "%a %b %d %H:%M:%S %z %Y")
end = datetime.strptime(tweets_list[len(tweets_list)-1]['created_at'], "%a %b %d %H:%M:%S %z %Y")
recent_date = f"{start.year}{start.month}{start.day}"
oldest_date = f"{end.year}{end.month}{end.day}"
print(f"Those tweets from {oldest_date} to {recent_date}.")
# part2: extract data from pulled tweets and generate lists for next
Acc_name = []
Tweet_date = []
Tweet_id = []
Text = []
Favor_count = []
Retweet_count = []
Lan = []
Acc_date = []
followers_count = []
Acc_location = []
for tweet in tweets_list:
Acc_name.append(tweet['user']['screen_name'])
Tweet_date.append(tweet['created_at'])
Tweet_id.append(tweet['id_str'])
Text.append(tweet['text'])
Favor_count.append(tweet['favorite_count'])
Retweet_count.append(tweet['retweet_count'])
Lan.append(tweet['lang'])
Acc_date.append(tweet['user']['created_at'])
followers_count.append(tweet['user']['followers_count'])
Acc_location.append(tweet['user']['location'])
# part3: generate data frame
df = pd.DataFrame({
"Account Name": Acc_name,
"Tweet Date": Tweet_date,
"Tweet ID": Tweet_id,
"Text": Text,
"Favorite Count": Favor_count,
"Retweet Count": Retweet_count,
"Language": Lan,
"Account Created Date": Acc_date,
"Followers Count": followers_count,
"Account Location": Acc_location
})
df=df[["Account Name", "Tweet Date", "Tweet ID", "Text", "Favorite Count", "Retweet Count", "Language",
"Account Created Date", "Followers Count", "Account Location"]]
# part4: sentiment analysis and add results to data frame
df["Compound"] = ""
df["Positive"] = ""
df["Negative"] = ""
df["Neutral"] = ""
for index, row in df.iterrows():
results = analyzer.polarity_scores(row["Text"])
df.set_value(index, 'Compound', results['compound'])
df.set_value(index, 'Positive', results['pos'])
df.set_value(index, 'Negative', results['neg'])
df.set_value(index, 'Neutral', results['neu'])
# part5: save data to CSV file
filename = f"{oldest_date}-{recent_date}-{target_user}.csv"
df.to_csv(filename)
target_user = "@Coinsquare"
pages = 50
pull_tweets_to_csv(target_user, pages)
| tweets_pull_template.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# +
import os
from dotenv import load_dotenv
import psycopg2
import pandas
load_dotenv() #> loads contents of the .env file into the script's environment
DB_NAME = os.getenv("DB_NAME")
DB_USER = os.getenv("DB_USER")
DB_PASSWORD = os.getenv("DB_PASSWORD")
DB_HOST = os.getenv("DB_HOST")
print(DB_NAME, DB_USER, DB_PASSWORD, DB_HOST)
connection = psycopg2.connect(dbname=DB_NAME, user=DB_USER, password=<PASSWORD>, host=DB_HOST)
print("CONNECTION:", connection)
cursor = connection.cursor()
print("CURSOR:", cursor)
# -
sql_whatever = """COPY titanic(survived,pclass,name,sex,age,siblings_spouses,parents_children,fare
)
FROM 'titanic.csv' with (format csv, header true, DELIMITER ',');"""
new_table =""" CREATE TABLE titanic2
(
survived INT,
pclass INT,
name VARCHAR(50),
sex VARCHAR(10),
age INT,
siblings_spouses INT,
parents_children INT,
fare FLOAT
);
"""
new_query = "CREATE TABLE titanic4 (Survived INT, Class INT, Name VARCHAR, Sex CHAR, Age FLOAT, Sibling_Spouse INT, Parent_Child INT, Fare FLOAT);"
cursor.execute(sql_whatever)
connection.commit()
cursor.execute(new_query)
connection.commit()
connection.close()
cursor.execute('SELECT * from titanic3;')
result = cursor.fetchall()
print("RESULT:", type(result))
print(result)
# +
import csv
cursor = connection.cursor()
print("CURSOR:", cursor)
with open('titanic.csv', 'r') as f:
reader = csv.reader(f)
next(reader) # Skip the header row.
for row in reader:
cursor.execute(new_query
)
conn.commit()
# +
import sqlite3
conn = sqlite3.connect('rpg_db.sqlite3')
curs = conn.cursor()
query = 'SELECT COUNT(*) FROM armory_item;'
curs.execute(query)
curs.execute(query).fetchall()
# +
import sqlite3
conn = sqlite3.connect('study_part1.sqlite3')
curs = conn.cursor()
students = "CREATE TABLE study_part1 (student VARCHAR,studied VARCHAR,grade INT,age INT,sex VARCHAR);"
query = students
curs.execute(query)
# -
fill = '''
INSERT INTO study_part1 (student, studied, grade, age, sex)
VALUES ('Lion-O', 'True', 85, 24, 'Male'),
('Cheetara', 'True', 95, 22, 'Female'),
('Mumm-Ra', 'False', 65, 153, 'Male'),
('Snarf', 'False', 70, 15, 'Male'),
('Panthro', 'True', 80, 30, 'Male');
'''
curs.execute(fill)
# +
gender ='''SELECT student
FROM study_part1
WHERE sex= 'Female'
'''
curs.execute(gender).fetchall()
# -
import sqlite3
conn = sqlite3.connect('demo_data.sqlite3')
curs = conn.cursor()
create = "CREATE TABLE demp(s VARCHAR,x INT, y INT);"
query = create
curs.execute(query)
fill = '''
INSERT INTO demp (s, x, y)
VALUES ('g', 3, 9),
('v', 5, 7),
('f', 8, 7)
;
'''
curs.execute(fill)
conn.commit()
count_rows = '''SELECT COUNT(*) FROM demp'''
curs.execute(count_rows)
print(conn.commit())
x_and_y='''
SELECT COUNT(*)
FROM demp
WHERE x>4 AND y>4
'''
curs.execute(x_and_y)
distincty = '''
SELECT COUNT (DISTINCT y) as d
FROM demp
'''
curs.execute(distincty)
| Challenge/scratchpad.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="pgJJLX1SG2qx" colab_type="text"
# # Matplotlib
# + [markdown] id="XXwIlyCUMknl" colab_type="text"
# ## Working with Multiple Figures
# + id="LP81RpO4VNyX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="4615060e-e185-4d26-f957-778b09eba302"
# installs wget for downloading files
# !pip install wget
# install pycroscopy for loading ibw files
# !pip install pycroscopy
# installs ncempy for loading dm3 files
# !pip install ncempy
# + id="GbMplRWUnguW" colab_type="code" colab={}
import wget
import pycroscopy as px
import h5py
import pyUSID as usid
import matplotlib.pyplot as plt
from ncempy.io import dm
from numpy import genfromtxt
import pandas as pd
import numpy as np
from matplotlib import image
# + id="c4mRM9TgduQJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 107} outputId="3e6f1a42-8b80-4f59-8686-8f5c3ef41672"
urls = ['https://github.com/jagar2/Summer_2020_MAT-395-495_Scientific-Data-Analysis-and-Computing/blob/master/lectures/4_Visualization/Data/gb_5_HAADF.dm3?raw=true',
'https://github.com/jagar2/Summer_2020_MAT-395-495_Scientific-Data-Analysis-and-Computing/blob/master/lectures/4_Visualization/Data/SP128_NSO_VPFM0001.ibw?raw=true',
'https://github.com/jagar2/Summer_2020_MAT-395-495_Scientific-Data-Analysis-and-Computing/blob/master/lectures/4_Visualization/Data/Data.xlsx?raw=true',
'https://github.com/jagar2/Summer_2020_MAT-395-495_Scientific-Data-Analysis-and-Computing/blob/master/lectures/4_Visualization/Data/Data.csv?raw=true']
# loops around the list of URLs
for url in urls:
print(url.split('/')[-1].split('?')[0]) # prints the string to save the file
wget.download(url, url.split('/')[-1].split('?')[0]) # downloads and saves the file
wget.download('https://www2.lehigh.edu/sites/www2/files/2020-02/Hulvat-4710.jpg','lehigh.jpg')
# + id="rUiPnQONYGWG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 541} outputId="0772fa3f-7a9a-4784-aaf5-b7b4e5b1517f"
# file path for ibw file
file_path = '/content/SP128_NSO_VPFM0001.ibw'
# instanciates the translator
translator = px.io.translators.IgorIBWTranslator()
# translates the file to h5
h5_path = translator.translate(file_path)
# + id="e8bJEryPtAXX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="02d2c954-d239-40d4-82ad-03e059793a2f"
# loads the dm3 file
im0 = dm.dmReader('gb_5_HAADF.dm3')
# views the content of the dm3 file
print(im0)
# + id="8DvQuJarYxRW" colab_type="code" colab={}
# reads the H5 file
h5_f = h5py.File(h5_path, 'r+')
# + id="nR8N5L4GZTij" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 467} outputId="ef9c8d5c-3fb1-4b05-95a6-d7626d73f3ab"
# prints the tree where the data is stored
usid.hdf_utils.print_tree(h5_f)
# + id="ltqedLPXZomE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="f36a3e89-0b17-4255-ef85-0da85da9565f"
# gets the main datasets
main_dset = usid.hdf_utils.get_all_main(h5_f)
# prints the information about the main datasets
print(main_dset)
# + id="rnC-1Euyap3H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="e188de83-87a3-48a8-c8d1-b5579f3d27e8"
# looks at the shape of the h5 file
np.array(h5_f['/Measurement_000/Channel_000/Raw_Data']).shape
# + id="dPuCSnkjg9gf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 143} outputId="af9fc5b9-0d4c-44ee-f360-b269114c04fb"
my_data = genfromtxt('Data.csv', delimiter=',',skip_header=1) # loads the csv file
# prints the csv file data
print(my_data)
# + id="eVhgJWf_hq_3" colab_type="code" colab={}
# loads the xlsx file into a pandas dataframe
data_xlsx = pd.read_excel('Data.xlsx')
# + id="YGpdgnnhhuff" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="75de786a-eef9-4dec-ae7b-f4c63ef771b9"
# prints the header of the dataframe
data_xlsx.head()
# + id="MlSlaYugjEwD" colab_type="code" colab={}
# loads the image file using matplotlib
image_file = image.imread('lehigh.jpg')
# + id="slhMsKyFjN9o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="765ca8da-7fd3-4845-cf0a-3f1ef4c5d20e"
# looks at the shape of the image file
image_file.shape
# + id="atzfqGMK1iZl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 313} outputId="8b629a3e-aebc-49f0-ff58-4cf33a26b336"
import matplotlib.pyplot as plt
from matplotlib.gridspec import GridSpec
import numpy as np
# builds the figure
fig = plt.figure(constrained_layout=True)
# uses gridspec to define the position of the axis
gs = GridSpec(3, 3, figure=fig)
# adds all the axis
ax1 = fig.add_subplot(gs[0, 0])
ax11 = fig.add_subplot(gs[0,1])
ax12 = fig.add_subplot(gs[0,2])
ax2 = fig.add_subplot(gs[1, :-1])
ax3 = fig.add_subplot(gs[1:, -1])
ax4 = fig.add_subplot(gs[-1, 0])
ax5 = fig.add_subplot(gs[-1, -2])
# creates a list for the 3 AFM images
ax_list = [ax1, ax11, ax12]
# iterates around each graph in a list plotting the i*2 channel
for i, ax in enumerate(ax_list):
# prints the name of the channel plotting
#print(f'/Measurement_000/Channel_00{i*2}/Raw_Data')
# plots the graph, fstring used for indexing. Reshape used to make 2d image
ax.imshow(np.array(h5_f[f'/Measurement_000/Channel_00{i*2}/Raw_Data']).reshape(1024,1024))
# plots the dm3 file
ax3.imshow(im0['data'])
# plots the data from the csv file
ax2.plot(my_data[:,0],my_data[:,1])
# plots the data from the xlsx file
ax2.plot(data_xlsx['X'],data_xlsx['cos(x)'])
# plots the image of Lehigh
ax4.imshow(image_file)
# plots the R color channel from the Lehigh image.
ax5.imshow(image_file[:,:,0])
# saves the figure at an image.
plt.savefig('fig.png', dpi = 300)
# saves the figure as a scaled vector graphic
plt.savefig('fig.svg', dpi = 300)
# + [markdown] id="E7d5KUN4wOUI" colab_type="text"
# ### Loading Data from a variety of sources
# + [markdown] colab_type="text" id="C3AVXktropQT"
# ## Parts of a Figure
# 
# + [markdown] colab_type="text" id="j_qxlM46opQY"
# # Graphical Integrity
# + [markdown] colab_type="text" id="8ayJKno-opQc"
# # Color scales
# + [markdown] id="QC9fwFQZZy_0" colab_type="text"
# # More details about perceptually correct colormaps
# + [markdown] colab_type="text" id="cI28x-YeopQl"
# # Choosing Colors for your Figures
# + [markdown] colab_type="text" id="U5aFUWM5opQo"
# # Types of Graphs
# + [markdown] id="ufMImc2LZy_9" colab_type="text"
# # Violin Plots
# * Like a box plot but provides a deeper understanding of data density
# * Good when you have large datasets
# + [markdown] id="x9eab9CXZzAA" colab_type="text"
# # 2D Density Plot
# * Used to compare 2D quantitative information
# * Good for small data sets
# * When the density of data is high (shouldn't use a scatter plot)
# + [markdown] id="s9A5MJ3eZzAC" colab_type="text"
# # Correlogram
# A correlogram or correlation matrix allows to analyse the relationship between each pair of numerical variables of a matrix.
# + [markdown] id="6p7tgK9nZzAF" colab_type="text"
# # Dendrogram
# * A dendrogram or tree diagram allows to illustrate the hierarchical organisation of several entities.
# + [markdown] id="LEWvfPzBZzAH" colab_type="text"
# # Graph Structures
# * Show interconnections between a set of entities.
# * Each entity is represented by a Node (or vertices).
# * Connection between nodes are represented through links (or edges).
# * Directed or undirected, weighted or unweighted.
# + [markdown] colab_type="text" id="Y9CDzbKIopQ6"
# # Guiding Principles
# + [markdown] colab_type="text" id="bTdkDQmyopRN"
# # WTF Graphs
# + [markdown] id="yfttWnj-TT0L" colab_type="text"
# # Assignment:
#
# Take a graph that you have previously used for a publication or assignment and recreate it based on concepts discussed in class. Please provide the original image and the updated image exported from python.
| lectures/4_Visualization/MAT_495_Pre_Lecture_2_Visual_Displays_of_Graphical_Information.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib notebook
import numpy as np
import math
import scipy
import copy
import os
import matplotlib.pyplot as plt
# llops, which provides backend-independent operations
import llops as yp
import llops.operators as ops
# coptic
from comptic import simulation, noise
# Project-specific imports
from htdeblur import blurkernel, analysis
yp.setDefaultBackend('numpy')
yp.setDefaultDatatype('float32')
# plt.style.use('publication')
# Figure output directory
figure_output_directory = os.path.expanduser('/Users/zfphil/Dropbox/Berkeley/My Talks/fom2019/')
# -
# ## Save forward Model Figures
# +
# Generater Blur kernel
vector, dnf = blurkernel.vector(pulse_count=12)
kernel = blurkernel.fromVector(vector, (64,64))
plt.figure(figsize=(4,4))
plt.imshow(kernel)
yp.savefig(figure_output_directory + '_kernel_crop.png')
# -
# ## Multi-Frame Figure
california = simulation.california(min_value=0, max_value=1, invert=False, shape=(320, 1000))
plt.figure()
plt.imshow(california)
plt.colorbar()
# +
object_shape = (600, 1200)
image_shape = (600, 600)
# Generate object
object_true = np.abs(yp.pad(california, object_shape, center=True))
# Define Crop ROIs
roi_list = (yp.Roi(start=(0,0), shape=image_shape, input_shape=object_shape),
yp.Roi(start=(0, object_shape[1] // 4), shape=image_shape, input_shape=object_shape),
yp.Roi(start=(0, object_shape[1] // 2), shape=image_shape, input_shape=object_shape))
# Generater Blur kernel
vector, dnf = blurkernel.vector(pulse_count=50)
kernel = blurkernel.fromVector(vector, object_shape)
# Define Operators
C = ops.Convolution(kernel)
G = ops.Segmentation(roi_list)
# Crop to measurements
measurement_list = ops.VecSplit(G * C * object_true,3)
plt.figure(figsize=(3,1.5))
plt.imshow(object_true)
plt.axis('off')
yp.savefig(figure_output_directory + 'system_ground_truth.png')
plt.figure(figsize=(1.5,1.5))
for index, measurement in enumerate(measurement_list):
plt.clf()
plt.imshow(measurement)
plt.axis('off')
yp.savefig(figure_output_directory + 'system_measurement_%d.png' % index)
plt.figure(figsize=(3,1.5))
plt.imshow(roi_list[1].mask)
plt.axis('off')
yp.savefig(figure_output_directory + 'mask.png')
vector, dnf = blurkernel.vector(pulse_count=15)
kernel_crop = blurkernel.fromVector(vector, (40,80))
plt.figure(figsize=(3,1.5))
plt.imshow(yp.circshift(kernel_crop, (0, -15)))
plt.axis('off')
yp.savefig(figure_output_directory + 'kernel_0.png')
plt.figure(figsize=(3,1.5))
plt.imshow(yp.circshift(kernel_crop, (0, 0)))
plt.axis('off')
yp.savefig(figure_output_directory + 'kernel_1.png')
plt.figure(figsize=(3,1.5))
plt.imshow(yp.circshift(kernel_crop, (0, 15)))
plt.axis('off')
yp.savefig(figure_output_directory + 'kernel_2.png')
# -
# ## Define Constants
plt.figure()
plt.imshow(kernel_crop)
# +
# Get system parameters
system_params = analysis.getDefaultSystemParams()
pulse_count = 25
# -
# ## Calculate SNR Improvement
t_strobe = 1e-5
t_coded = t_strobe * pulse_count
dnf_coded = analysis.getOptimalDnf(pulse_count*2)
system_params_illum_copy =copy.deepcopy(system_params)
system_params_illum_copy['camera_ad_conversion'] = 0.46
system_params_illum_copy['camera_quantum_efficency'] = 0.9
system_params_illum_copy['illuminance'] = 1000
system_params_illum_copy['readout_noise'] = 10
system_params_illum_copy['dark_current'] = 0.5
system_params_illum_copy['velocity_max'] = 2
snr_coded = analysis.exposureTimeToSnr(t_coded, dnf=dnf_coded, **system_params_illum_copy)
snr_strobe = analysis.exposureTimeToSnr(t_strobe, dnf=1, **system_params_illum_copy)
# print(snr_coded / snr_strobe)
print(snr_coded)
# +
illuminance_list = 10000, 2000, 10000
readout_noise_list = 10, 20, 40
frame_rate = 3
# Initialize lists
snr_strobed_list = []
snr_sns_list = []
snr_coded_list = []
snr_coded_raw_list = []
for illuminance, readout_noise in zip(illuminance_list, readout_noise_list):
# Define illuminance
system_params_illum = copy.deepcopy(system_params)
system_params_illum['illuminance'] = illuminance
system_params_illum['readout_noise'] = readout_noise
# Calculate Photon Pixel rate
photon_pixel_rate = analysis.illuminanceToPhotonPixelRate(**system_params)
# SNS
t_sns, dnf_sns = analysis.frameRateToExposure(frame_rate, photon_pixel_rate, 'stop_and_stare', pulse_count=pulse_count, **system_params_illum)
snr_sns = analysis.exposureTimeToSnr(t_sns, dnf=dnf_sns, **system_params_illum)
snr_sns_list.append(snr_sns)
# Strobed
t_strobe, dnf_strobe = analysis.frameRateToExposure(frame_rate, photon_pixel_rate, 'strobe', pulse_count=pulse_count, **system_params_illum)
snr_strobe = analysis.exposureTimeToSnr(t_strobe, dnf=dnf_strobe, **system_params_illum)
snr_strobed_list.append(snr_strobe)
# Coded
t_coded, dnf_coded = analysis.frameRateToExposure(frame_rate, photon_pixel_rate, 'code', pulse_count=pulse_count, **system_params_illum)
snr_coded_list.append(analysis.exposureTimeToSnr(t_coded, dnf=dnf_coded, **system_params_illum))
snr_coded_raw_list.append( analysis.exposureTimeToSnr(t_coded, dnf=1, **system_params_illum))
print(snr_strobed_list)
print(snr_coded_list)
print(snr_coded_list[1] / snr_strobed_list[1])
print(snr_coded_list[2] / snr_strobed_list[2])
# -
# ## Generate Blurry and Strobed Groups of Images as Examples
# https://www.ptgrey.com/support/downloads/10501
# +
# Generate object
object_true = np.abs(yp.pad(simulation.testObject('ucbseal', shape=(400,400)), (512,512), center=True, pad_value='edge')) * 100
# Generater Blur kernel
vector, dnf = blurkernel.vector(pulse_count=30)
kernel = blurkernel.fromVector(vector, object_true.shape)
# Define SNR function
snr = lambda x: noise.snr(x, signal_roi=yp.Roi(shape=(40, 40), start=(280, 30)), noise_roi=yp.Roi(shape=(10, 10), start=(40, 40)))
# Generate blurry object
object_blurry = yp.convolve(object_true, kernel)
# Define SNR to generate images from
measurement_list = []
for snr_strobed, snr_coded, snr_coded_raw in zip(snr_strobed_list, snr_coded_list, snr_coded_raw_list):
data_sublist = []
# Generate strobed data
data_sublist.append(noise.add(object_true, snr=snr_strobed))
# Generate coded measurement
data_sublist.append(noise.add(object_blurry, snr=snr_coded_raw))
# Deconvolve blurry measurement
data_sublist.append(yp.deconvolve(data_sublist[-1], kernel, reg=1e-3))
# Append to measurement list
measurement_list.append(data_sublist)
plt.figure(figsize=(12, 6))
index = 0
clim=(50,150)
cmap = 'gray'
plt.subplot(331)
plt.imshow(measurement_list[index][0], cmap=cmap)
plt.title('%g / %g' % (snr(measurement_list[index][0]), snr_strobed_list[index]))
plt.clim(clim)
plt.axis('off')
plt.subplot(332)
plt.imshow(measurement_list[index][1], cmap=cmap)
plt.title('%g / %g' % (snr(measurement_list[index][1]), snr_coded_raw_list[index]))
plt.clim(clim)
plt.axis('off')
plt.subplot(333)
plt.imshow(measurement_list[index][2], cmap=cmap)
plt.title('%g / %g' % (snr(measurement_list[index][2]), snr_coded_list[index]))
plt.clim(clim)
plt.axis('off')
index = 1
plt.subplot(334)
plt.imshow(measurement_list[index][0], cmap=cmap)
plt.title('%g / %g' % (snr(measurement_list[index][0]), snr_strobed_list[index]))
plt.clim(clim)
plt.axis('off')
plt.subplot(335)
plt.imshow(measurement_list[index][1], cmap=cmap)
plt.title('%g / %g' % (snr(measurement_list[index][1]), snr_coded_raw_list[index]))
plt.clim(clim)
plt.axis('off')
plt.subplot(336)
plt.imshow(measurement_list[index][2], cmap=cmap)
plt.title('%g / %g' % (snr(measurement_list[index][2]), snr_coded_list[index]))
plt.clim(clim)
plt.axis('off')
index = 2
plt.subplot(337)
plt.imshow(measurement_list[index][0], cmap=cmap)
plt.title('%g / %g' % (snr(measurement_list[index][0]), snr_strobed_list[index]))
plt.clim(clim)
plt.axis('off')
plt.subplot(338)
plt.imshow(measurement_list[index][1], cmap=cmap)
plt.title('%g / %g' % (snr(measurement_list[index][1]), snr_coded_raw_list[index]))
plt.clim(clim)
plt.axis('off')
plt.subplot(339)
plt.imshow(measurement_list[index][2], cmap=cmap)
plt.title('%g / %g' % (snr(measurement_list[index][2]), snr_coded_list[index]))
plt.clim(clim)
plt.axis('off')
plt.tight_layout()
# -
# ## Sweep System Pulse Count
# The parameter pulse_count is a scaler which represents the ratio of the amount of pulses used vs the amount of pulses which would saturate the camera
# +
# Set frame rate
frame_rate = 10
# Calculate Photon Pixel rate
photon_pixel_rate = analysis.illuminanceToPhotonPixelRate(**system_params)
# Stop and stare signal-to-noise
t_sns, dnf_sns = analysis.frameRateToExposure(frame_rate, photon_pixel_rate,'stop_and_stare', **system_params)
snr_sns = analysis.exposureTimeToSnr(t_sns, dnf=dnf_sns, **system_params)
counts_sns, noise_dependent, noise_independent = analysis.exposureTimeToNoiseComponents(t_sns, dnf=dnf_sns, **system_params)
noise_independent = 1 if noise_independent == 0 else noise_independent
print("SNS illumination at %d fps will have exposure %g seconds, %g counts, %g SNR (dnf = %g), and a noise ratio of %g" % (frame_rate, t_sns, counts_sns, snr_sns, dnf_sns, noise_dependent / noise_independent))
# Strobed signal-to-noise
t_strobe, dnf_strobe = analysis.frameRateToExposure(frame_rate, photon_pixel_rate,'strobe', **system_params)
snr_strobe = analysis.exposureTimeToSnr(t_strobe, dnf=dnf_strobe, **system_params)
counts_strobe, noise_dependent, noise_independent = analysis.exposureTimeToNoiseComponents(t_strobe, dnf=dnf_strobe, **system_params)
noise_independent = 1 if noise_independent == 0 else noise_independent
print("Strobed illumination at %d fps will have exposure %g seconds, %g counts, %g SNR (dnf = %g), and a noise ratio of %g" % (frame_rate, t_strobe, counts_strobe, snr_strobe, dnf_strobe, noise_dependent / noise_independent))
# Loop over illumination beta
for pulse_count in np.arange(1,1000,10):
t_coded, dnf_coded = analysis.frameRateToExposure(frame_rate, photon_pixel_rate,'code', pulse_count=pulse_count, **system_params)
snr_coded = analysis.exposureTimeToSnr(t_coded, dnf=dnf_coded, **system_params)
counts_coded, noise_dependent, noise_independent = analysis.exposureTimeToNoiseComponents(t_coded, dnf=dnf_coded, **system_params)
noise_independent = 1 if noise_independent == 0 else noise_independent
print("pulse_count=%g coded illumination at %d fps will have exposure %g seconds, %g counts, %g SNR (dnf = %g), and a noise ratio of %g" % (pulse_count, frame_rate, t_coded, counts_coded, snr_coded, dnf_coded, noise_dependent / noise_independent))
# -
# ## Plot SNR vs Frame Rate for 10 Lux
# +
# Define which frame rates to use
frame_rates = np.arange(0.1, 50, 1)
# Define number of pulses for coded
pulse_count = 30
# Define which illuminance to use
illuminance_list = [1000, 40000]
# Initialize lists
snr_strobe_list = []
snr_sns_list = []
snr_coded_list = [] # list of lists
# Loop over frame rates
for index, frame_rate in enumerate(frame_rates):
# Loop over illuminance
snr_sns_sublist, snr_strobed_sublist, snr_coded_sublist = [], [], []
for illuminance in illuminance_list:
# Define illuminance
system_params_illum = copy.deepcopy(system_params)
system_params_illum['illuminance'] = illuminance
# Calculate Photon Pixel rate
photon_pixel_rate = analysis.illuminanceToPhotonPixelRate(**system_params_illum)
# SNS
t_sns, dnf_sns = analysis.frameRateToExposure(frame_rate, photon_pixel_rate, 'stop_and_stare', pulse_count=pulse_count, **system_params_illum)
snr_sns = analysis.exposureTimeToSnr(t_sns, dnf=dnf_sns, **system_params_illum)
snr_sns_sublist.append(snr_sns)
# Strobed
t_strobe, dnf_strobe = analysis.frameRateToExposure(frame_rate, photon_pixel_rate, 'strobe', pulse_count=pulse_count, **system_params_illum)
snr_strobe = analysis.exposureTimeToSnr(t_strobe, dnf=dnf_strobe, **system_params_illum)
snr_strobed_sublist.append(snr_strobe)
# Coded
t_coded, dnf_coded = analysis.frameRateToExposure(frame_rate, photon_pixel_rate, 'code', pulse_count=pulse_count, **system_params_illum)
snr_coded = analysis.exposureTimeToSnr(t_coded, dnf=dnf_coded, **system_params_illum)
snr_coded_sublist.append(snr_coded)
# Append
snr_sns_list.append(snr_sns_sublist)
snr_strobe_list.append(snr_strobed_sublist)
snr_coded_list.append(snr_coded_sublist)
snr_sns_list_transpose = np.asarray(snr_sns_list).T.tolist()
snr_strobe_list_transpose = np.asarray(snr_strobe_list).T.tolist()
snr_coded_list_transpose = np.asarray(snr_coded_list).T.tolist()
# Perform plotting
lw = 3
plt.figure(figsize=(8, 8))
# Loop over illuminance
c = ['g', 'r', 'b', 'y']
for index, (illuminance, snr_sns, snr_strobe, snr_coded) in enumerate(zip(illuminance_list, snr_sns_list_transpose, snr_strobe_list_transpose, snr_coded_list_transpose)):
plt.semilogy(frame_rates, snr_coded, linewidth=lw, label='Coded (%d lux)' % illuminance, c=c[index])
# plt.semilogy(frame_rates, snr_sns, 'r-', linewidth=lw, label='Stop and Stare (%d lux)' % illuminance)
if index == 0:
plt.semilogy(frame_rates, snr_strobe, 'k-', linewidth=lw, label='Stobed (%d lux)' % illuminance)
else:
plt.semilogy(frame_rates, snr_strobe, 'k--', linewidth=lw, label='Stobed (%d lux)' % illuminance)
# Configure figure
plt.legend()
plt.xlabel('Frame Rate (Hz)', fontsize=28)
plt.ylabel('Imaging SNR', fontsize=28)
plt.ylim((1e-1, 1000))
plt.xlim((0, 50))
plt.grid('on', which='both')
plt.title('Illuminance and SNR Improvement')
# Set up ticks
ax = plt.gca()
for tick in ax.xaxis.get_major_ticks():
tick.label.set_fontsize(20)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(20)
# Save
plt.tight_layout()
# plt.savefig(os.path.join(figure_directory, 'snr_plot.png'))
# -
# ## Generate Example Images
# +
index = 100
snr_coded_0 = snr_coded_list_transpose[0][index]
snr_coded_1 = snr_coded_list_transpose[1][index]
snr_strobe_0 = snr_strobe_list_transpose[0][index]
snr_strobe_1 = snr_strobe_list_transpose[1][index]
# Generate object
object_true = np.abs(sim.ucb())
# Define SNR to generate images from
snr_list = [snr_coded_0, snr_strobe_0, snr_coded_1, snr_strobe_1]
noisy_object_list = []
for snr in snr_list:
noisy_object_list.append(noise.add(object_true, snr=snr))
# Show result
plt.figure(figsize=(2,6))
clim = [0.5, 1.5]
for index, (noisy, snr) in enumerate(zip(noisy_object_list, snr_list)):
plt.subplot(411 + index)
plt.imshow(np.abs(noisy))
# plt.title('SNR: %g' % snr)
plt.clim(clim)
plt.axis('off')
plt.savefig(os.path.join(figure_directory, 'snr_images.png'))
| notebooks/fig_fom_2019.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # WeatherPy
# ----
#
# #### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import time
# Import API key
import api_keys
# Incorporated citipy to determine city based on latitude and longitude
from citipy import citipy
# Output File (CSV)
output_data_file = "output_data/cities.csv"
# Range of latitudes and longitudes
lat_range = (-90, 90)
lng_range = (-180, 180)
# -
# ## Generate Cities List
# +
# List for holding lat_lngs and cities
lat_lngs = []
cities = []
# Create a set of random lat and lng combinations
lats = np.random.uniform(low=-90.000, high=90.000, size=1500)
lngs = np.random.uniform(low=-180.000, high=180.000, size=1500)
lat_lngs = zip(lats, lngs)
# Identify nearest city for each lat, lng combination
for lat_lng in lat_lngs:
city = citipy.nearest_city(lat_lng[0], lat_lng[1]).city_name
# Replace spaces with %20 to create url correctly
city = city.replace(" ", "%20")
# If the city is unique, then add it to a our cities list
if city not in cities:
cities.append(city)
# Print the city count to confirm sufficient count
len(cities)
# -
# ### Perform API Calls
# * Perform a weather check on each city using a series of successive API calls.
# * Include a print log of each city as it'sbeing processed (with the city number and city name).
#
# +
# OpenWeatherMap API Key
api_key = weather_api_key
# Starting URL for Weather Map API Call
url = "http://api.openweathermap.org/data/2.5/weather?units=Imperial&APPID=" + api_key
# +
# Create empty lists to append the API data into lists
city_name = []
cloudiness = []
country = []
date = []
humidity = []
lat = []
lng = []
max_temp = []
wind_speed = []
# Start the call counter
record = 1
# Log file print statement
print(f"Beginning Data Retrieval")
print(f"-------------------------------")
#Loop through the cities in the city list
for city in cities:
# Try statement to append calls where value is found
# Not all calls return data as OpenWeatherMap will not have have records in all the cities generated by CityPy module
try:
response = requests.get(f"{url}&q={city}").json()
city_name.append(response["name"])
cloudiness.append(response["clouds"]["all"])
country.append(response["sys"]["country"])
date.append(response["dt"])
humidity.append(response["main"]["humidity"])
max_temp.append(response["main"]["temp_max"])
lat.append(response["coord"]["lat"])
lng.append(response["coord"]["lon"])
wind_speed.append(response["wind"]["speed"])
city_record = response["name"]
print(f"Processing Record {record} | {city_record}")
print(f"{url}&q={city}")
# Increase counter by one
record= record + 1
# Wait a second in loop to not over exceed rate limit of API
time.sleep(1.01)
# If no record found "skip" to next call
except:
print("City not found. Skipping...")
continue
# +
# Create a dictonary with the lists generated
weatherpy_dict = {
"City": city_name,
"Cloudiness":cloudiness,
"Country":country,
"Date":date,
"Humidity": humidity,
"Lat":lat,
"Lng":lng,
"Max Temp": max_temp,
"Wind Speed":wind_speed
}
# Create a data frame from dictionary
weather_data = pd.DataFrame(weatherpy_dict)
# Display count of weather data values
weather_data.count()
# +
# Collect and parse the data into a Data Frame called
city= []
clouds= []
country= []
date= []
humidity= []
lat= []
lng= []
temp= []
wind= []
for data in city_data:
if data['cod']==200:
city.append(data['name'])
clouds.append(data['clouds']['all'])
country.append(data['sys']['country'])
humidity.append(data['main']['humidity'])
lat.append(data['coord']['lat'])
lng.append(data['coord']['lon'])
temp.append(data['main']['temp_max'])
wind.append(data['wind']['speed'])
weather_dict = {"City": city,
"Cloudiness": clouds,
"Country": country,
"Humidity": humidity,
"Lat": lat,
"Lng": lng,
"Temperature (°F)": temp,
"Wind Speed": wind}
weather_data_df = pd.DataFrame(weather_dict)
weather_data_df=weather_data_df[["City","Country","Lat","Lng","Temperature (°F)","Humidity","Wind Speed","Cloudiness"]]
weather_data_df.head()
# +
# Collect and parse the data into a Data Frame called
city= []
clouds= []
country= []
date= []
humidity= []
lat= []
lng= []
temp= []
wind= []
for data in city_data:
if data['cod']==200:
city.append(data['name'])
clouds.append(data['clouds']['all'])
country.append(data['sys']['country'])
humidity.append(data['main']['humidity'])
lat.append(data['coord']['lat'])
lng.append(data['coord']['lon'])
temp.append(data['main']['temp_max'])
wind.append(data['wind']['speed'])
weather_dict = {"City": city,
"Cloudiness": clouds,
"Country": country,
"Humidity": humidity,
"Lat": lat,
"Lng": lng,
"Temperature (°F)": temp,
"Wind Speed": wind}
weather_data_df = pd.DataFrame(weather_dict)
weather_data_df=weather_data_df[["City","Country","Lat","Lng","Temperature (°F)","Humidity","Wind Speed","Cloudiness"]]
weather_data_df.head()
# -
#Save weather data to a cities csv file
weather_data.to_csv("../output_data/cities.csv", index=False)
# ## Inspect the data and remove the cities where the humidity > 100%.
# ----
# Skip this step if there are no cities that have humidity > 100%.
# ## Plotting the Data
# * Use proper labeling of the plots using plot titles (including date of analysis) and axes labels.
# * Save the plotted figures as .pngs.
# ## Latitude vs. Temperature Plot
# +
#Latitude vs. Temperature Plot scatter plot
plt.scatter(weather_data["Lat"], weather_data["Max Temp"], color = "steelblue", edgecolor = "black")
plt.title("City Latitude vs. Max Temperature (06/18/20)")
plt.xlabel("Latitude")
plt.ylabel("Max Temperature (F)")
plt.grid(True)
plt.savefig("../Images/City Latitude vs Max Temperature.png")
# -
# ## Latitude vs. Humidity Plot
# +
#Latitude vs. Humidity Plot scatter plot
plt.scatter(weather_data["Lat"], weather_data["Humidity"], color = "steelblue", edgecolor = "black")
plt.title("City Latitude vs. Humidity (06/18/20)")
plt.xlabel("Latitude")
plt.ylabel("Humidity (%))")
plt.grid(True)
plt.savefig("../Images/City Latitude vs Humidity.png")
# -
# ## Latitude vs. Cloudiness Plot
# +
#Latitude vs. Cloudiness Plot scatter plot
plt.scatter(weather_data["Lat"], weather_data["Cloudiness"], color = "steelblue", edgecolor = "black")
plt.title("City Latitude vs. Humidity (06/18/20)")
plt.xlabel("Latitude")
plt.ylabel("Cloudines (%))")
plt.grid(True)
plt.savefig("../Images/City Latitude vs Cloudiness.png")
# -
# ## Latitude vs. Wind Speed Plot
# +
#Latitude vs. Wind Speed Plot scatter plot
plt.scatter(weather_data["Lat"], weather_data["Wind Speed"], color = "steelblue", edgecolor = "black")
plt.title("City Latitude vs. Wind Speed (06/18/20)")
plt.xlabel("Latitude")
plt.ylabel("Wind Speed (%))")
plt.grid(True)
plt.savefig("../Images/City Latitude vs Wind Speed.png")
| WeatherPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Export WALIS data for ShinyAppp
# This notebook contains the scripts to download the full WALIS database and prepare a CSV file for the R Shiny App hosted at: https://warmcoasts.shinyapps.io/WALIS_Visualization/
# ## Dependencies and packages
# This notebook calls various scripts that are included in the \scripts folder. The following is a list of the python libraries needed to run this notebook.
# +
#Main packages
import pandas as pd
import pandas.io.sql as psql
import geopandas
import pygeos
import numpy as np
import mysql.connector
from datetime import date
import xlsxwriter as writer
import math
from scipy import optimize
from scipy import stats
#Plots
import seaborn as sns
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1 import make_axes_locatable
#Jupyter data display
import tqdm
from tqdm.notebook import tqdm_notebook
from IPython.display import *
import ipywidgets as widgets
from ipywidgets import *
#Geographic
from shapely.geometry import Point
from shapely.geometry import box
import cartopy as ccrs
import cartopy.feature as cfeature
#System
import os
import glob
import shutil
#pandas options for debugging
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
#Set a date string for exported file names
date=date.today()
dt_string = date.strftime("_%d_%m_%Y")
# Ignore warnings
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
warnings.filterwarnings('ignore')
# -
# ## Import database
# Connect to the online MySQL database containing WALIS data and download data into a series of pandas data frames.
# + hide_input=false
## Connect to the WALIS database server
# %run -i scripts/connection.py
## Import data tables and show progress bar
with tqdm_notebook(total=len(SQLtables),desc='Importing tables from WALIS') as pbar:
for i in range(len(SQLtables)):
query = "SELECT * FROM {}".format(SQLtables[i])
walis_dict[i] = psql.read_sql(query, con=db)
query2 = "SHOW FULL COLUMNS FROM {}".format(SQLtables[i])
walis_cols[i] = psql.read_sql(query2, con=db)
pbar.update(1)
path = os.getcwd()
CHECK_FOLDER = os.path.isdir('Output')
Output = 'Output'
if not CHECK_FOLDER:
Output_path=os.path.join(path,Output)
os.mkdir(Output_path)
else:
Output_path=os.path.join(path,Output)
CHECK_FOLDER = os.path.isdir('Output/Shiny_input')
Shiny_app = 'Shiny_input'
if not CHECK_FOLDER:
Data_path=os.path.join(Output_path,Shiny_app)
os.mkdir(Data_path)
else:
Data_path=os.path.join(Output_path,Shiny_app)
# -
# The following scripts make connections between the data and produce the summary file, which will be processed in the next step.
# %run -i scripts/select_user.py
# %run -i scripts/multi_author_query.py
# %run -i scripts/substitutions.py
# %run -i scripts/make_summary.py
Summary.to_csv('Output/Shiny_input/Summary.csv',index = False,encoding='utf-8-sig')
# # Make data analysis
# This section takes the "Summary.csv" file and performs some basic data analysis on it.
#
# ## RSL percentiles
# Then, the script takes information on relative sea level values and calculates RSL percentiles in the following way.
# 1. If the RSL Indicator is a "Single Coral": the percentiles are obtained from a gamma function interpolated considering the upper limit of living range inserted in the database as, respectively, the 2.3 and 97.7 percentiles of the distribution.
# 2. If the RSL Indicator is a "Sea Level Indicator" or "Single Speleothem": the percentiles on paleo RSL are calculated from the gaussian distribution represented by the field "Paleo RSL (m)" and its associated uncertainty (1-sigma).
# 3. If the RSL Indicator is a "Terrestrial Limiting" or "Marine Limiting", the RSL percentiles are not calculated.
#
# ## Age percentiles
# The following script takes information on age values and calculates age percentiles according to the table below. The following modifications are done on the original data:
#
# - If a percentile goes below zero, it is set to zero.
# - If Lower age > Upper age, the two values are reversed.
# - If there is no age, the corresponding record is deleted.
#
# | Dating technique | Pre-selection | Lower age | Age (ka) 0.1 perc | Age (ka) 2.3 perc | Age (ka) 15.9 perc | Age (ka) 50 perc | Age (ka) 84.1 perc | Age (ka) 97.7 perc | Age (ka) 99.5 perc | Upper age |
# |-|-|-|-|-|-|-|-|-|-|-|
# | U-series / coral | Recalculated age used if available. If not, Reported age is used | NaN | Average age - 3 Sigma age | Average age - 2 Sigma age | Average age - 1 Sigma age | Average age | Average age + 1 Sigma age | Average age + 2 Sigma age | Average age + 3 Sigma age | NaN |
# | U-series / speleothem | Recalculated age used if available. If not, Reported age is used | NaN | Average age - 3 Sigma age | Average age - 2 Sigma age | Average age - 1 Sigma age | Average age | Average age + 1 Sigma age | Average age + 2 Sigma age | Average age + 3 Sigma age | NaN |
# | U-series / mollusks or algae | Upper and lower age derived from the MIS to which the sample is associated with | Lower age |<--|--|--| Uniform distribution |--|--|-->| Upper age |
# | AAR / Age reported | | NaN | Average age - 3 Sigma age | Average age - 2 Sigma age | Average age - 1 Sigma age | Average age | Average age + 1 Sigma age | Average age + 2 Sigma age | Average age + 3 Sigma age | NaN |
# | AAR / Only MIS reported | Upper and lower age derived from the MIS to which the sample is associated with | Lower age |<--|--|--| Uniform distribution |--|--|-->| Upper age |
# | ESR / Age reported | | NaN | Average age - 3 Sigma age | Average age - 2 Sigma age | Average age - 1 Sigma age | Average age | Average age + 1 Sigma age | Average age + 2 Sigma age | Average age + 3 Sigma age | NaN |
# | ESR / Only MIS reported | Upper and lower age derived from the MIS to which the sample is associated with | Lower age |<--|--|--| Uniform distribution |--|--|-->| Upper age |
# | Luminescence / Age reported | | NaN | Average age - 3 Sigma age | Average age - 2 Sigma age | Average age - 1 Sigma age | Average age | Average age + 1 Sigma age | Average age + 2 Sigma age | Average age + 3 Sigma age | NaN |
# | Luminescence / Only MIS reported | Upper and lower age derived from the MIS to which the sample is associated with | Lower age |<--|--|--| Uniform distribution |--|--|-->| Upper age |
# | Stratigraphic constraint / Age reported| Upper and lower age derived from the reported age | Lower age |<--|--|--| Uniform distribution |--|--|-->| Upper age |
# | Stratigraphic constraint / Only MIS reported| Upper and lower age derived from the MIS to which the sample is associated with | Lower age |<--|--|--| Uniform distribution |--|--|-->| Upper age |
# | Other age constraint / Age reported| Upper and lower age derived from the reported age | Lower age |<--|--|--| Uniform distribution |--|--|-->| Upper age |
# | Other age constraint / Only MIS reported| Upper and lower age derived from the MIS to which the sample is associated with | Lower age |<--|--|--| Uniform distribution |--|--|-->| Upper age |
# %run -i scripts/percentiles_from_summary.py
# # Suggested acknowledgments
# WALIS is the result of the work of several people, within different projects. For this reason, we kindly ask you to follow these simple rules to properly acknowledge those who worked on it:
#
# 1. Cite the original authors - Please maintain the original citations for each datapoint, to give proper credit to those who worked to collect the original data in the field or in the lab.
# 2. Acknowledge the database contributor - The name of each contributor is listed in all public datapoints. This is the data creator, who spent time to make sure the data is standardized and (as much as possible) free of errors.
# 3. Acknowledge the database structure and interface creators - The database template used in this study was developed by the ERC Starting Grant "WARMCOASTS" (ERC-StG-802414) and is a community effort under the PALSEA (PAGES / INQUA) working group.
#
# Example of acknowledgments: The data used in this study were *[extracted from / compiled in]* WALIS, a sea-level database interface developed by the ERC Starting Grant "WARMCOASTS" (ERC-StG-802414), in collaboration with PALSEA (PAGES / INQUA) working group. The database structure was designed by <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME> and <NAME>. The data points used in this study were contributed to WALIS by *[list names of contributors here]*.
| Code/.ipynb_checkpoints/Export_to_ShinyApp-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pytorch
# language: python
# name: bento_kernel_pytorch
# ---
# # torchrec Criteo Terabyte Tutorial
# ## Table of contents
# 1. Instantiating Criteo Terabyte dataset
# 2. Defining and applying batch data transformation function
# 3. Defining model
# 4. Training and evaluating model
# 5. Training and evaluating model on GPU
# +
from typing import Dict, List, Tuple, Union
import torch
from torchrec.datasets.criteo import criteo_terabyte
torch.set_printoptions(threshold=20)
# -
# ## 1. Instantiating Criteo Terabyte dataset
# Let's begin by instantiating a datapipe representing the Criteo 1TB Click Logs https://ailab.criteo.com/download-criteo-1tb-click-logs-dataset/ dataset (we'll refer to it here as the Criteo Terabyte dataset).
datapipe = criteo_terabyte(
("/home/jeffhwang/local/datasets/criteo/day_11.tsv",),
)
# By default, the datapipe returns each sample as a dictionary that maps each default feature name to a typecasted feature value (int for each of the label and 13 integer features, and str for each of the 26 categorical features).
next(iter(datapipe))
# We can adjust the format of each sample via input parameter `row_mapper`. For instance, if we'd prefer to work with lists of feature values, we can define and provide a function that maps a raw split TSV line to a list of typecasted values:
# +
from torchrec.datasets.utils import safe_cast
def row_to_list(row):
return [
safe_cast(val, int, 0) for val in row[:14]
] + [
safe_cast(val, str, "") for val in row[14:]
]
list_datapipe = criteo_terabyte(
("/home/jeffhwang/local/datasets/criteo/day_11.tsv",),
row_mapper=row_to_list,
)
next(iter(list_datapipe))
# -
# Or, if we'd prefer to operate directly on raw split TSV lines, we can pass `None`:
raw_datapipe = criteo_terabyte(
("/home/jeffhwang/local/datasets/criteo/day_11.tsv",),
row_mapper=None,
)
next(iter(raw_datapipe))
# Next, we move onto creating train and validation datapipes representing complementary subsets of the dataset and applying a sample limit, batching, and collation to each:
# +
from torchrec.datasets.utils import idx_split_train_val
datapipe = criteo_terabyte(
("/home/jeffhwang/local/datasets/criteo/day_11.tsv",),
)
train_datapipe, val_datapipe = idx_split_train_val(datapipe, 0.7)
train_datapipe = train_datapipe.limit(int(1e3)).batch(100).collate()
val_datapipe = val_datapipe.limit(int(1e3)).batch(100).collate()
# -
# ## 2. Defining and applying batch data transformation function
#
# At this point, each item that is read from `train_datapipe` and `val_datapipe` is a dictionary representing a batch of 100 Criteo Terabyte samples ("batch dictionary"). The dictionary maps each string feature name to 100 feature values, each corresponding to a sample in the batch.
#
# Each of the 13 feature names corresponding to integer-valued features ("int_0" through "int_12") maps to a shape-(100,) tensor of integers; each of the 26 feature names corresponding to categorical features ("cat_0" through "cat_25") maps to a length-100 list of hex strings.
batch = next(iter(train_datapipe))
print("int_0:", batch["int_0"])
print("cat_0:", batch["cat_0"])
# There are a few data transformations we'd like to apply to each batch dictionary to produce the data we want to feed into our model:
# - Normalize integer feature values, e.g. by applying a logarithmic function.
# - Map each categorical feature hex string value to an integer that can be used to index into an embedding table.
# - Separate integer features, categorical features, and labels into individual tensors reshaped appropriately.
#
# Towards accomplishing this, we define a function `_transform` that accepts a batch dictionary as an input, applies the aforementioned transformations, and returns a tuple of three tensors corresponding to integer features, categorical features, and labels:
# +
from torchrec.datasets.criteo import DEFAULT_CAT_NAMES, DEFAULT_INT_NAMES, DEFAULT_LABEL_NAME
NUM_EMBEDDINGS = int(1e5)
col_transforms = {
**{name: lambda x: torch.log(x + 2) for name in DEFAULT_INT_NAMES},
**{
name: lambda x: x.fmod(NUM_EMBEDDINGS - 1) + 1
for name in DEFAULT_CAT_NAMES
},
}
def _transform(
batch: Dict[str, List[Union[int, str]]]
) -> Tuple[torch.Tensor, torch.Tensor, torch.Tensor]:
int_x = torch.cat(
[
col_transforms[col_name](torch.tensor(batch[col_name]).unsqueeze(0).T)
for col_name in DEFAULT_INT_NAMES
if col_name in col_transforms
],
dim=1,
)
cat_x = torch.cat(
[
col_transforms[col_name](
torch.tensor([int(v, 16) if v else -1 for v in batch[col_name]])
.unsqueeze(0)
.T
)
for col_name in DEFAULT_CAT_NAMES
if col_name in col_transforms
],
dim=1,
)
y = torch.tensor(batch[DEFAULT_LABEL_NAME], dtype=torch.float32).unsqueeze(1)
return int_x, cat_x, y
# -
# Then, using `map`, we produce a new pair of train and validation datapipes that applies `_transform` to each batch dictionary of data:
train_datapipe = train_datapipe.map(_transform)
val_datapipe = val_datapipe.map(_transform)
next(iter(train_datapipe))
# Now we've got datapipes that produce data that we can train and evaluate a model on!
# ## 3. Defining model
# To utilize the integer (dense) and categorical (sparse) features present in the Criteo Terabyte dataset, we define `TestSparseNN`, which maps dense and sparse features to embeddings and interacts the embeddings to produce an output:
# +
from torchrec.fb.modules.mlp import LazyMLP
class TestSparseNN(torch.nn.Module):
def __init__(
self,
*,
hidden_layer_size,
output_dim,
sparse_input_size,
num_embeddings,
embedding_dim,
):
super(TestSparseNN, self).__init__()
self.dense_arch = LazyMLP([hidden_layer_size, embedding_dim])
self.embedding_layers = self._embedding_layers(
sparse_input_size, num_embeddings, embedding_dim
)
self.over_arch = LazyMLP([output_dim])
self.final = torch.nn.LazyLinear(1)
def _embedding_layers(self, sparse_input_size, num_embeddings, embedding_dim):
return torch.nn.ModuleList(
[
torch.nn.Embedding(num_embeddings, embedding_dim, padding_idx=0)
for _ in range(sparse_input_size)
]
)
def _interact(self, embeddings):
batch_size, embedding_dim = embeddings[0].shape
stacked_embeddings = torch.cat(embeddings, dim=1).view(
batch_size, -1, embedding_dim
)
interactions = torch.matmul(
stacked_embeddings, torch.transpose(stacked_embeddings, 1, 2)
)
_, embedding_count, _ = interactions.shape
rows, cols = torch.tril_indices(embedding_count, embedding_count)
return interactions[:, rows, cols]
def forward(self, dense_x, cat_x):
embedded_dense = self.dense_arch(dense_x)
embedded_sparse = [
embedding_layer(cat_x[:, idx])
for idx, embedding_layer in enumerate(self.embedding_layers)
]
interactions = self._interact([embedded_dense] + embedded_sparse)
return self.final(
self.over_arch(torch.cat([embedded_dense, interactions], dim=1))
)
# -
# ## 4. Training and evaluating model
# We can now train an instance of `TestSparseNN` on data supplied by `train_datapipe`
# +
model = TestSparseNN(
hidden_layer_size=20,
output_dim=10,
sparse_input_size=26,
num_embeddings=NUM_EMBEDDINGS,
embedding_dim=16,
)
# Initialize lazy modules.
int_x, cat_x, y = next(iter(train_datapipe))
model(int_x, cat_x)
loss_fn = torch.nn.BCEWithLogitsLoss()
optimizer = torch.optim.Adagrad(model.parameters(), lr=1e-2, weight_decay=1e-6)
for batch_num, (int_x, cat_x, y) in enumerate(train_datapipe):
res = model(int_x, cat_x)
loss = loss_fn(res, y)
optimizer.zero_grad()
loss.backward()
optimizer.step()
if batch_num % 1 == 0:
loss, current = loss.item(), batch_num * len(y)
print(f"loss: {loss:>7f} {current}")
# -
# , and evaluate the trained model on data supplied by `val_datapipe`
# +
import sklearn.metrics
y_true = []
y_pred = []
with torch.no_grad():
for int_x, cat_x, y in val_datapipe:
pred = model(int_x, cat_x)
y_pred.append(pred)
y_true.append(y)
auroc = sklearn.metrics.roc_auc_score(
torch.cat(y_true).view(-1),
torch.sigmoid(torch.cat(y_pred).view(-1)),
)
val_loss = loss_fn(
torch.cat(y_pred).view(-1),
torch.cat(y_true).view(-1),
)
print("Test results:")
print(f"AUROC: {auroc:>8f} Avg loss: {val_loss:>8f}")
# -
# ## 5. Training and evaluating model on GPU
#
# If we have access to a GPU device, we can leverage it as follows to accelerate model training and evaluation.
# +
assert(torch.cuda.is_available())
device = torch.device("cuda:0")
datapipe = criteo_terabyte(
("/home/jeffhwang/local/datasets/criteo/day_11.tsv",),
)
train_datapipe, val_datapipe = idx_split_train_val(datapipe, 70)
train_datapipe = train_datapipe.limit(int(1e6)).batch(1000).collate().map(_transform)
val_datapipe = val_datapipe.limit(int(1e5)).batch(1000).collate().map(_transform)
model.to(device)
int_x, cat_x, y = next(iter(train_datapipe))
int_x, cat_x, y = int_x.to(device), cat_x.to(device), y.to(device)
model(int_x, cat_x)
loss_fn = torch.nn.BCEWithLogitsLoss()
optimizer = torch.optim.Adagrad(model.parameters(), lr=1e-2, weight_decay=1e-6)
for batch_num, (int_x, cat_x, y) in enumerate(train_datapipe):
int_x, cat_x, y = int_x.to(device), cat_x.to(device), y.to(device)
res = model(int_x, cat_x)
loss = loss_fn(res, y)
optimizer.zero_grad()
loss.backward()
optimizer.step()
if batch_num % 10 == 0:
loss, current = loss.item(), batch_num * len(y)
print(f"loss: {loss:>7f} {current}")
y_true = []
y_pred = []
with torch.no_grad():
for int_x, cat_x, y in val_datapipe:
int_x, cat_x, y = int_x.to(device), cat_x.to(device), y.to(device)
pred = model(int_x, cat_x)
y_pred.append(pred)
y_true.append(y)
auroc = sklearn.metrics.roc_auc_score(
torch.cat(y_true).view(-1).cpu(),
torch.sigmoid(torch.cat(y_pred).view(-1)).cpu(),
)
val_loss = loss_fn(
torch.cat(y_pred).view(-1).cpu(),
torch.cat(y_true).view(-1).cpu(),
)
print("Test results:")
print(f"AUROC: {auroc:>8f} Avg loss: {val_loss:>8f}")
# -
| torchrec/examples/notebooks/criteo_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Observations and Insights
# Observations:
# 1. There was less variability in tumor volumes for drug regimens 'Capomulin' and 'Ramicane'
# 2. Both 'Capomulin' and 'Ramicane'have lower SEM showing that the standard error falls as the sample size increases.
# 2. Drug regimen 'Capomulin' shows a strong correlation between weight and tumor volume.
#
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import scipy.stats as st
# Study data files
mouse_metadata_path = "data/Mouse_metadata.csv"
study_results_path = "data/Study_results.csv"
# Read the mouse data and the study results
mouse_metadata = pd.read_csv(mouse_metadata_path)
study_results = pd.read_csv(study_results_path)
# Combine the data into a single dataset
mouse_study_results = pd.merge(mouse_metadata,study_results,on='Mouse ID')
# Display the data table for preview
mouse_study_results.head()
# -
# Checking the number of mice.
number_of_mice = len(mouse_study_results['Mouse ID'].unique())
print(f'Number of mice: {number_of_mice}')
# Getting the duplicate mice by ID number that shows up for Mouse ID and Timepoint.
grouped_df=mouse_study_results.groupby(['Mouse ID','Timepoint'])['Timepoint'].count()
grouped_df[grouped_df.values > 1]
# Optional: Get all the data for the duplicate mouse ID.
mouse_study_results[mouse_study_results['Mouse ID'] == 'g989']
# Create a clean DataFrame by dropping the duplicate mouse by its ID.
mouse_study_results = mouse_study_results[mouse_study_results['Mouse ID'] != 'g989']
# Checking the number of mice in the clean DataFrame.
number_of_mice = len(mouse_study_results['Mouse ID'].unique())
print(f'Number of mice after dropping duplicate mouse: {number_of_mice}')
# ## Summary Statistics
# +
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
# Use groupby and summary statistical methods to calculate the following properties of each drug regimen:
# mean, median, variance, standard deviation, and SEM of the tumor volume.
mean_tumor_value = mouse_study_results.groupby('Drug Regimen')['Tumor Volume (mm3)'].mean()
median_tumor_value = mouse_study_results.groupby('Drug Regimen')['Tumor Volume (mm3)'].median()
tumor_value_variance = mouse_study_results.groupby('Drug Regimen')['Tumor Volume (mm3)'].var()
tumor_vol_std = mouse_study_results.groupby('Drug Regimen')['Tumor Volume (mm3)'].std()
tumor_vol_std_error = mouse_study_results.groupby('Drug Regimen')['Tumor Volume (mm3)'].sem()
# Assemble the resulting series into a single summary dataframe.
summary_df = pd.DataFrame({'Mean Tumor Value': mean_tumor_value,
'Median Tumor Value': median_tumor_value,
'Tumor Value Variance': tumor_value_variance,
'Tumor Value Std. Dev.': tumor_vol_std,
'Tumor Value Std. Err.' :tumor_vol_std_error})
summary_df
# +
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
# Using the aggregation method, produce the same summary statistics in a single line
mouse_study_results.groupby(
['Drug Regimen']
).agg(
{
'Tumor Volume (mm3)':['mean','median','var','std','sem']
}
)
# -
# ## Bar and Pie Charts
# Generate a bar plot showing the total number of measurements taken on each drug regimen using pandas.
groupbydrugregimen = mouse_study_results.groupby(['Drug Regimen'])['Mouse ID'].count().sort_values(ascending=False)
# groupbydrugregimen
ax = groupbydrugregimen.plot(kind='bar',figsize=(10,6))
ax.set_xlabel("Drug Regimen")
ax.set_ylabel("Number of Unique Mice Tested");
# Generate a bar plot showing the total number of measurements taken on each drug regimen using pyplot.
y_vals = mouse_study_results.groupby(['Drug Regimen'])['Mouse ID'].count().sort_values(ascending=False)
plt.figure(figsize=(10,6))
plt.bar(y_vals.keys(),y_vals)
plt.xlabel('Drug Regimen')
plt.ylabel('Number of Unique Mice Tested')
plt.xticks(rotation=90)
plt.show()
# Generate a pie plot showing the distribution of female versus male mice using pandas
ax = mouse_study_results.groupby(['Sex'])['Mouse ID'].count().sort_values(ascending=False).plot(kind='pie',autopct='%1.1f%%')
ax.set_ylabel('Sex');
# Generate a pie plot showing the distribution of female versus male mice using pyplot
val = mouse_study_results.groupby(['Sex'])['Mouse ID'].count().sort_values(ascending=False)
plt.figure(figsize=(10,4))
plt.pie(val,labels=val.keys(),autopct='%1.1f%%')
plt.ylabel('Sex')
plt.show()
# ## Quartiles, Outliers and Boxplots
# +
# Calculate the final tumor volume of each mouse across four of the treatment regimens:
# Capomulin, Ramicane, Infubinol, and Ceftamin
reduced_mouse_study_results = mouse_study_results[mouse_study_results['Drug Regimen'].isin(['Capomulin','Ramicane','Infubinol','Ceftamin'])]
reduced_mouse_study_results = reduced_mouse_study_results[['Drug Regimen','Mouse ID','Timepoint','Tumor Volume (mm3)']]
# Start by getting the last (greatest) timepoint for each mouse
lasttimepoint = reduced_mouse_study_results.groupby('Mouse ID')['Timepoint'].max()
# Merge this group df with the original dataframe to get the tumor volume at the last timepoint
tumorvalueoflasttimepoint = pd.merge(reduced_mouse_study_results, lasttimepoint, on=['Mouse ID','Timepoint'])
tumorvalueoflasttimepoint = tumorvalueoflasttimepoint[['Drug Regimen', 'Timepoint','Tumor Volume (mm3)']]
# +
import numpy as np
# Put treatments into a list for for loop (and later for plot labels)
treatments = list(tumorvalueoflasttimepoint['Drug Regimen'].unique())
# Create empty list to fill with tumor vol data (for plotting)
tumorvol = []
outliers = []
# Calculate the IQR and quantitatively determine if there are any potential outliers.
for t in range(len(treatments)):
# Locate the rows which contain mice on each drug and get the tumor volumes
tv = tumorvalueoflasttimepoint[tumorvalueoflasttimepoint['Drug Regimen'] == treatments[t]]['Tumor Volume (mm3)']
tumorvol.append(tv)
q1 = np.quantile(tv,0.25)
q3 = np.quantile(tv,0.75)
iqr = q3 - q1
lower_bound = q1 -(1.5 * iqr)
upper_bound = q3 +(1.5 * iqr)
# print("\ntreatment: ", treatments[t], "\nq1: ",q1,"\nq3: ",q3,"\niqr: ",iqr,"\nlower: ",lower_bound,"\nupper: ",upper_bound)
# Determine outliers using upper and lower bounds
for index, value in tv.items():
if (value > upper_bound) or (value < lower_bound):
outliers.append(value)
print('Drug Regimen ', treatments[t], ' has an outlier: ', outliers)
# +
# Generate a box plot of the final tumor volume of each mouse across four regimens of interest
boxplot = tumorvalueoflasttimepoint.boxplot(by='Drug Regimen',column=['Tumor Volume (mm3)'])
boxplot.set_title("")
plt.suptitle("")
boxplot.set_ylabel("Final Tumor Value (mm3)")
boxplot.set_xlabel("")
plt.show()
# -
# ## Line and Scatter Plots
# Generate a line plot of tumor volume vs. time point for a mouse treated with Capomulin
capomulin_u364_mouse_study_results = mouse_study_results[(mouse_study_results['Drug Regimen']=="Capomulin") & (mouse_study_results['Mouse ID']=="u364")]
ax = capomulin_u364_mouse_study_results.plot.line(x='Timepoint', y='Tumor Volume (mm3)')
ax.set_ylabel('Tumor Volume (mm3)')
ax.set_title('Tumor Volume for Drug Regimen Capomulin for Mouse ID u364')
plt.show()
# +
# Generate a scatter plot of average tumor volume vs. mouse weight for the Capomulin regimen
capomulin_mouse_study_results = mouse_study_results[(mouse_study_results['Drug Regimen']=="Capomulin")]
avgtumorvol=capomulin_mouse_study_results.groupby(['Weight (g)','Mouse ID'])['Tumor Volume (mm3)'].mean()
avgtumorvol
for name, group in avgtumorvol.iteritems():
plt.scatter(x=name[0],y=group,color='blue')
plt.xlabel('Weight(g)')
plt.ylabel('Average Tumor Volume (mm3)')
# -
# ## Correlation and Regression
# +
# Calculate the correlation coefficient and linear regression model
# for mouse weight and average tumor volume for the Capomulin regimen
from scipy.stats import linregress
capomulin_mouse_study_results = mouse_study_results[(mouse_study_results['Drug Regimen']=="Capomulin")]
avgtumorvol=capomulin_mouse_study_results.groupby(['Weight (g)','Mouse ID'])['Tumor Volume (mm3)'].mean()
weight = []
tumorvol = []
for name, group in avgtumorvol.iteritems():
weight.append(name[0])
tumorvol.append(group)
plt.scatter(x=weight,y=tumorvol,color='blue')
plt.xlabel('Weight(g)')
plt.ylabel('Average Tumor Volume (mm3)')
# print(type(weight[0]))
#y = mx + b
(slope, intercept, rvalue, pvalue, stderr) = linregress(weight,tumorvol)
# print(type(slope))
y = []
for index in range(len(weight)):
y.append(weight[index] * slope + intercept)
plt.plot(weight,y,'r-')
ax = plt.title('The correlation between mouse weight and average tumor value is ' + str(round(rvalue,2)))
| Pymaceuticals/pymaceuticals_starter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Read data.
import os
# Folder containing all NIPS papers.
data_dir = 'nipstxt/'
# Folders containin individual NIPS papers.
yrs = ['00', '01', '02', '03', '04', '05', '06', '07', '08', '09', '10', '11', '12']
dirs = ['nips' + yr for yr in yrs]
# Read all texts into a list.
docs = []
for yr_dir in dirs:
files = os.listdir(data_dir + yr_dir)
for filen in files:
# Note: ignoring characters that cause encoding errors.
with open(data_dir + yr_dir + '/' + filen, 'r') as fid:
txt = fid.read()
docs.append(txt)
# +
text = docs
# Tokenize the documents.
from nltk.tokenize import RegexpTokenizer
# Split the documents into tokens.
tokenizer = RegexpTokenizer(r'\w+')
for idx in range(len(docs)):
docs[idx] = docs[idx].lower() # Convert to lowercase.
docs[idx] = tokenizer.tokenize(docs[idx]) # Split into words.
# Remove numbers, but not words that contain numbers.
docs = [[token for token in doc if not token.isnumeric()] for doc in docs]
# Remove words that are only one character.
docs = [[token for token in doc if len(token) > 1] for doc in docs]
# +
# Lemmatize the documents.
from nltk.stem.wordnet import WordNetLemmatizer
# Lemmatize all words in documents.
lemmatizer = WordNetLemmatizer()
docs = [[lemmatizer.lemmatize(token) for token in doc] for doc in docs]
# Compute bigrams.
from gensim.models import Phrases
# Add bigrams and trigrams to docs (only ones that appear 20 times or more).
bigram = Phrases(docs, min_count=20)
for idx in range(len(docs)):
for token in bigram[docs[idx]]:
if '_' in token:
# Token is a bigram, add to document.
docs[idx].append(token)
# Remove rare and common tokens.
from gensim.corpora import Dictionary
# Create a dictionary representation of the documents.
dictionary = Dictionary(docs)
# Filter out words that occur less than 20 documents, or more than 50% of the documents.
dictionary.filter_extremes(no_below=20, no_above=0.5)
# Vectorize data.
# Bag-of-words representation of the documents.
corpus = [dictionary.doc2bow(doc) for doc in docs]
print('Number of unique tokens: %d' % len(dictionary))
print('Number of documents: %d' % len(corpus))
# +
# Train LDA model.
from gensim.models import LdaModel
# Set training parameters.
num_topics = 10
chunksize = 2000
passes = 20
iterations = 400
eval_every = None # Don't evaluate model perplexity, takes too much time.
# Make a index to word dictionary.
temp = dictionary[0] # This is only to "load" the dictionary.
id2word = dictionary.id2token
# %time model = LdaModel(corpus=corpus, id2word=id2word, chunksize=chunksize, \
# alpha='auto', eta='auto', \
# iterations=iterations, num_topics=num_topics, \
# passes=passes, eval_every=eval_every)
# -
import pyLDAvis.gensim
pyLDAvis.enable_notebook()
pyLDAvis.gensim.prepare(model, corpus, dictionary)
# +
def format_topics_sentences(ldamodel=None, corpus=corpus, texts=data):
# Init output
sent_topics_df = pd.DataFrame()
# Get main topic in each document
for i, row_list in enumerate(ldamodel[corpus]):
row = row_list[0] if ldamodel.per_word_topics else row_list
# print(row)
row = sorted(row, key=lambda x: (x[1]), reverse=True)
# Get the Dominant topic, Perc Contribution and Keywords for each document
for j, (topic_num, prop_topic) in enumerate(row):
if j == 0: # => dominant topic
wp = ldamodel.show_topic(topic_num)
topic_keywords = ", ".join([word for word, prop in wp])
sent_topics_df = sent_topics_df.append(pd.Series([int(topic_num), round(prop_topic,4), topic_keywords]), ignore_index=True)
else:
break
sent_topics_df.columns = ['Dominant_Topic', 'Perc_Contribution', 'Topic_Keywords']
# Add original text to the end of the output
contents = pd.Series(texts)
sent_topics_df = pd.concat([sent_topics_df, contents], axis=1)
return(sent_topics_df)
df_topic_sents_keywords = format_topics_sentences(ldamodel=model, corpus=corpus, texts=text)
# Format
df_dominant_topic = df_topic_sents_keywords.reset_index()
df_dominant_topic.columns = ['Document_No', 'Dominant_Topic', 'Topic_Perc_Contrib', 'Keywords', 'Text']
df_dominant_topic.head(10)
# +
# 1. Wordcloud of Top N words in each topic
from matplotlib import pyplot as plt
from wordcloud import WordCloud, STOPWORDS
import matplotlib.colors as mcolors
cols = [color for name, color in mcolors.TABLEAU_COLORS.items()] # more colors: 'mcolors.XKCD_COLORS'
cloud = WordCloud(stopwords=stop_words,
background_color='white',
width=2500,
height=1800,
max_words=100,
colormap='tab10',
color_func=lambda *args, **kwargs: cols[i],
prefer_horizontal=1.0)
topics = model.show_topics(formatted=False)
fig, axes = plt.subplots(2, 2, figsize=(10,10), sharex=True, sharey=True)
for i, ax in enumerate(axes.flatten()):
fig.add_subplot(ax)
topic_words = dict(topics[i][1])
cloud.generate_from_frequencies(topic_words, max_font_size=300)
plt.gca().imshow(cloud)
plt.gca().set_title('Topic ' + str(i), fontdict=dict(size=16))
plt.gca().axis('off')
plt.subplots_adjust(wspace=0, hspace=0)
plt.axis('off')
plt.margins(x=0, y=0)
plt.tight_layout()
plt.show()
# +
from collections import Counter
topics = model.show_topics(formatted=False)
data_flat = [w for w_list in text for w in w_list]
counter = Counter(data_flat)
out = []
for i, topic in topics:
for word, weight in topic:
out.append([word, i , weight, counter[word]])
df = pd.DataFrame(out, columns=['word', 'topic_id', 'importance', 'word_count'])
# Plot Word Count and Weights of Topic Keywords
fig, axes = plt.subplots(2, 2, figsize=(8,8), sharey=True, dpi=160)
cols = [color for name, color in mcolors.TABLEAU_COLORS.items()]
for i, ax in enumerate(axes.flatten()):
ax.bar(x='word', height="word_count", data=df.loc[df.topic_id==i, :], color=cols[i], width=0.4, alpha=0.3, label='Word Count')
ax_twin = ax.twinx()
ax_twin.bar(x='word', height="importance", data=df.loc[df.topic_id==i, :], color=cols[i], width=0.2, label='Weights')
ax.set_ylabel('Word Count', color=cols[i])
ax_twin.set_ylim(0, 0.030); ax.set_ylim(0, 3500)
ax.set_title('Topic: ' + str(i), color=cols[i], fontsize=12)
ax.tick_params(axis='y', left=False)
ax.set_xticklabels(df.loc[df.topic_id==i, 'word'], rotation=30, horizontalalignment= 'right')
ax.legend(loc='upper left'); ax_twin.legend(loc='upper right')
fig.tight_layout(w_pad=2)
fig.suptitle('Word Count and Importance of Topic Keywords', fontsize=16, y=1.05)
plt.show()
# +
# Get topic weights and dominant topics ------------
from sklearn.manifold import TSNE
from bokeh.plotting import figure, output_file, show
from bokeh.models import Label
from bokeh.io import output_notebook
# Get topic weights
topic_weights = []
for i, row_list in enumerate(model[corpus]):
#print(row_list)
topic_weights.append([w for i, w in row_list])
# Array of topic weights
arr = pd.DataFrame(topic_weights).fillna(0).values
# Keep the well separated points (optional)
arr = arr[np.amax(arr, axis=1) > 0.35]
# Dominant topic number in each doc
topic_num = np.argmax(arr, axis=1)
# tSNE Dimension Reduction
tsne_model = TSNE(n_components=2, verbose=1, random_state=0, angle=.99, init='pca')
tsne_lda = tsne_model.fit_transform(arr)
# Plot the Topic Clusters using Bokeh
output_notebook()
n_topics = 4
mycolors = np.array([color for name, color in mcolors.TABLEAU_COLORS.items()])
plot = figure(title="t-SNE Clustering of {} LDA Topics".format(n_topics),
plot_width=900, plot_height=700)
plot.scatter(x=tsne_lda[:,0], y=tsne_lda[:,1], color=mycolors[topic_num])
show(plot)
# -
| topic_modelling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Working with Matrices
import numpy as np
# ## Matrices and liner combinations
#
# ### Post-multiplication with vector
#
# Matrix-vector multiplication is a linear combination of the columns of the matrix
#
# $$
# \begin{bmatrix}
# 1 & 2 \\
# 3 & 4 \\
# 5 & 6
# \end{bmatrix}
# \begin{bmatrix}
# 2 \\ 3
# \end{bmatrix} =
# 2 \begin{bmatrix}
# 1 \\ 3 \\ 5
# \end{bmatrix} +
# 3 \begin{bmatrix}
# 2 \\ 4 \\ 6
# \end{bmatrix} =
# \begin{bmatrix}
# 8 \\
# 18 \\
# 28
# \end{bmatrix}
# $$
#
# $$
# \begin{bmatrix}
# 1 & 2 \\
# 3 & 4 \\
# 5 & 6
# \end{bmatrix}
# \begin{bmatrix}
# 1 \\ 4
# \end{bmatrix} =
# 1 \begin{bmatrix}
# 1 \\ 3 \\ 5
# \end{bmatrix} +
# 4 \begin{bmatrix}
# 2 \\ 4 \\ 6
# \end{bmatrix} =
# \begin{bmatrix}
# 9 \\
# 19 \\
# 29
# \end{bmatrix}
# $$
#
# We can stack the columns horizontally to get matrix multiplication.
#
# $$
# \begin{bmatrix}
# 1 & 2 \\
# 3 & 4 \\
# 5 & 6
# \end{bmatrix}
# \begin{bmatrix}
# 2 & 1 \\ 3 & 4
# \end{bmatrix} =
# \begin{bmatrix}
# 8 & 9 \\
# 18 & 19 \\
# 28 & 29
# \end{bmatrix}
# $$
A = np.arange(1, 7).reshape((3,2))
x1 = np.array([2, 3]).reshape((2,1))
x2 = np.array([1,4]).reshape((2,1))
A @ x1
A @ x2
np.c_[x1, x2]
A @ np.c_[x1, x2]
# ### Pre-multiplication with vector
#
# Vector-matrix multiplication is a linear combination of the rows of the matrix
#
# $$
# \begin{bmatrix}
# 1 & 2 & 3
# \end{bmatrix}
# \begin{bmatrix}
# 1 & 2 \\
# 3 & 4 \\
# 5 & 6
# \end{bmatrix}=
# 1 \begin{bmatrix}
# 1 & 2
# \end{bmatrix} +
# 2 \begin{bmatrix}
# 3 & 4
# \end{bmatrix} +
# 3 \begin{bmatrix}
# 5 & 6
# \end{bmatrix} =
# \begin{bmatrix}
# 22 & 28
# \end{bmatrix}
# $$
#
# $$
# \begin{bmatrix}
# 4 & 5 & 6
# \end{bmatrix}
# \begin{bmatrix}
# 1 & 2 \\
# 3 & 4 \\
# 5 & 6
# \end{bmatrix}=
# 4 \begin{bmatrix}
# 1 & 2
# \end{bmatrix} +
# 5 \begin{bmatrix}
# 3 & 4
# \end{bmatrix} +
# 6 \begin{bmatrix}
# 5 & 6
# \end{bmatrix} =
# \begin{bmatrix}
# 49 & 64
# \end{bmatrix}
# $$
#
# We can stack the rows vertically to get matrix multiplication.
#
# $$
# \begin{bmatrix}
# 1 & 2 & 3 \\
# 4 & 5 & 4
# \end{bmatrix}
# \begin{bmatrix}
# 1 & 2 \\
# 3 & 4 \\
# 5 & 6
# \end{bmatrix} =
# \begin{bmatrix}
# 22 & 28 \\
# 49 & 64
# \end{bmatrix}
# $$
#
# Matrix-matrix multiplication can be seen as the horizontal stacking of column operations or as the vertical stacking of row operations.
y1 = np.array([1,2,3]).reshape((1,3))
y2 = np.array([4,5,6]).reshape((1,3))
y1 @ A
y2 @ A
np.r_[y1, y2]
np.r_[y1, y2] @ A
# ### Extract columns of a matrix by post-multiplication with standard unit column vector
A
e2 = np.array([0,1]).reshape((-1,1))
A @ e2
# ### Extract rows of a matrix by pre-multiplication with standard unit row vector
e2 = np.array([0,1,0]).reshape((-1, 1))
e2.T @ A
# ## Permutation matrices
#
# From the column extraction by post-multiplication with a standard unit column vector, we generalize to permutation matrices (identity matrix with permuted columns). Post-multiplication of a matrix $A$ with a permutation matrix $P$ rearranges the columns of $A$. To recover the original matrix, multiply with $P^T$ - i.e. $P^{-1} = P^T$ and the inverse of $P$ is its inverse, $P$ being our first example of an orthogonal matrix.
A = np.arange(1, 17).reshape((4,4))
A
I = np.eye(4, dtype='int')
I
A @ I
p = I[:, [2,1,3,0]]
p
A @ p
A @ p @ p.T
# ## Matrix partitioning
#
# We see above that matrix multiplication can be seen as separate operations on the row or column vectors. We can actually partition matrices into blocks (not just vectors) for matrix multiplication. Suppose we want to calculate $AB$, where
#
# \begin{align}
# A = \begin{bmatrix}
# 1 & 0 & 1 & 0 \\
# 0 & 1 & 0 & 1 \\
# 0 & 0 & 2 & 0 \\
# 0 & 0 & 0 & 3
# \end{bmatrix}&, & B = \begin{bmatrix}
# 1 & 2 & 3 & 4 \\
# 5 & 6 & 7 & 8 \\
# 0 & 0 & 1 & 0 \\
# 0 & 0 & 0 & 1
# \end{bmatrix}
# \end{align}
#
# We can consider (say) $A$ and $B$ as each being a $2 \times 2$ matrix where each element is a $2 \times 2$ sub-matrix (or block). This simplifies the computation since many blocks are the identity or null matrix.
#
# \begin{align}
# A = \begin{bmatrix}
# A_{11} & A_{12} \\
# A_{21} & A_{22}
# \end{bmatrix}&, & B = \begin{bmatrix}
# B_{11} & B_{12} \\
# B_{21} & B_{22}
# \end{bmatrix}
# \end{align}
#
# and
#
# $$
# AB = \begin{bmatrix}
# A_{11}B_{11} + A_{12}B_{21} & A_{11}B_{12} + A_{12}B_{22} \\
# A_{21}B_{11} + A_{22}B_{22} & A_{21}B_{12} + A_{22}B_{22}
# \end{bmatrix}
# $$
#
# In fact, we can see by inspection that the result will be
#
# $$
# AB = \begin{bmatrix}
# B_{11} & B_{12}+I_2 \\
# 0_2 & A_{22}
# \end{bmatrix} = \begin{bmatrix}
# 1 & 2 & 4 & 4 \\
# 5 & 6 & 7 & 9 \\
# 0 & 0 & 2 & 0 \\
# 0 & 0 & 0 & 3
# \end{bmatrix}
# $$
#
# In general, any sub-block structure consistent with matrix multiplication (more formally, $A$ and $B$ are *conformable* for multiplication) is fine. In particular, the blocks do not have to be square.
# +
a11 = np.eye(2)
a12 = np.eye(2)
a21 = np.zeros((2,2))
a22 = np.diag((2,3))
b11 = np.array([
[1,2],
[5,6]
])
b12 = np.array([
[3,4],
[7,8]
])
b21 = np.zeros((2,2))
b22 = np.eye(2)
# -
A = np.block([
[a11, a12],
[a21, a22]
]).astype('int')
A
B = np.block([
[b11, b12],
[b21, b22]
]).astype('int')
B
A @ B
np.block([
[a11@b11 + a12@b21, a11@b12 + a12@b22],
[a21@b11 + a22@b21, a21@b12 + a22@b22]
]).astype('int')
| notebooks/copies/lectures/T02A_Matrix_Multiplication.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# ### Rare event or Unbalanced data set
#
# Providing equal sample of positive and negative instances to the classification algorithm will result in optimal result. Dataset which are highly skewed towards one or more classes have proven to be a challenge.
#
# Resampling is a common practice of addressing this issue. Although there are many techniques within re-sampling, here we'll be learning the 3 most popular techniques.
#
# * Random under-sampling - Reduce majority class to match minority class count
# * Random over-sampling - Increase minority class by randomly picking samples within minority class till counts of both class match
# * Synthetic Minority Over Sampling Technique (SMOTE) - Increase minority class by introducing synthetic examples through connecting all k (default = 5) minority class nearest negihbors using feature space similarity (Euclidean distance)
#
# A number of techniques have been implemented in "imbalanced-learn" python package hosted at https://github.com/scikit-learn-contrib/imbalanced-learn. Follow the instructions in the link to install it, if not already installed.
from IPython.display import Image
Image(filename='../Chapter 4 Figures/Sampling.png', width=800)
# +
# Load libraries
import matplotlib.pyplot as plt
from sklearn.datasets import make_classification
from imblearn.under_sampling import RandomUnderSampler
from imblearn.over_sampling import RandomOverSampler
from imblearn.over_sampling import SMOTE
# +
# Generate the dataset with 2 features to keep it simple
X, y = make_classification(n_samples=5000, n_features=2, n_informative=2,
n_redundant=0, weights=[0.9, 0.1], random_state=2017)
print "Positive class: ", y.tolist().count(1)
print "Negative class: ", y.tolist().count(0)
# -
# ### Applying the sampling techniques
# +
# Apply the random under-sampling
rus = RandomUnderSampler()
X_RUS, y_RUS = rus.fit_sample(X, y)
# Apply the random over-sampling
ros = RandomOverSampler()
X_ROS, y_ROS = ros.fit_sample(X, y)
# Apply regular SMOTE
sm = SMOTE(kind='regular')
X_SMOTE, y_SMOTE = sm.fit_sample(X, y)
# -
# ### Plot the original vs re-sampled
# +
# Original vs resampled subplots
plt.figure(figsize=(10, 6))
plt.subplot(2,2,1)
plt.scatter(X[y==0,0], X[y==0,1], marker='o', color='blue')
plt.scatter(X[y==1,0], X[y==1,1], marker='+', color='red')
plt.xlabel('x1')
plt.ylabel('x2')
plt.title('Original: 1=%s and 0=%s' %(y.tolist().count(1), y.tolist().count(0)))
plt.subplot(2,2,2)
plt.scatter(X_RUS[y_RUS==0,0], X_RUS[y_RUS==0,1], marker='o', color='blue')
plt.scatter(X_RUS[y_RUS==1,0], X_RUS[y_RUS==1,1], marker='+', color='red')
plt.xlabel('x1')
plt.ylabel('y2')
plt.title('Random Under-sampling: 1=%s and 0=%s' %(y_RUS.tolist().count(1), y_RUS.tolist().count(0)))
plt.subplot(2,2,3)
plt.scatter(X_ROS[y_ROS==0,0], X_ROS[y_ROS==0,1], marker='o', color='blue')
plt.scatter(X_ROS[y_ROS==1,0], X_ROS[y_ROS==1,1], marker='+', color='red')
plt.xlabel('x1')
plt.ylabel('x2')
plt.title('Random over-sampling: 1=%s and 0=%s' %(y_ROS.tolist().count(1), y_ROS.tolist().count(0)))
plt.subplot(2,2,4)
plt.scatter(X_SMOTE[y_SMOTE==0,0], X_SMOTE[y_SMOTE==0,1], marker='o', color='blue')
plt.scatter(X_SMOTE[y_SMOTE==1,0], X_SMOTE[y_SMOTE==1,1], marker='+', color='red')
plt.xlabel('x1')
plt.ylabel('y2')
plt.title('SMOTE: 1=%s and 0=%s' %(y_SMOTE.tolist().count(1), y_SMOTE.tolist().count(0)))
plt.tight_layout()
plt.show()
# -
# ### Known disadvantages:
# * Random Under-Sampling rises opportunity for loss of information or concepts as we are reducing the majority class
# * Random Over-Sampling & SMOTE can lead to overfitting issue due to multiple related instances
#
# ### Which re-sampling technique is the best?
#
# Well, as always there is no one answer to this question! Let's try a quick classification model on 3 re-sampled data and compare the accuracy (we'll use AUC as this one of the best representation of model performance)
#
#
# +
from sklearn import tree
from sklearn import metrics
# split data into train and test
from sklearn.cross_validation import train_test_split
X_RUS_train, X_RUS_test, y_RUS_train, y_RUS_test = train_test_split(X_RUS, y_RUS, test_size=0.3, random_state=2017)
X_ROS_train, X_ROS_test, y_ROS_train, y_ROS_test = train_test_split(X_ROS, y_ROS, test_size=0.3, random_state=2017)
X_SMOTE_train, X_SMOTE_test, y_SMOTE_train, y_SMOTE_test = train_test_split(X_SMOTE, y_SMOTE, test_size=0.3, random_state=2017)
# build a decision tree classifier
clf = tree.DecisionTreeClassifier(random_state=2017)
clf_rus = clf.fit(X_RUS_train, y_RUS_train)
clf_ros = clf.fit(X_ROS_train, y_ROS_train)
clf_smote = clf.fit(X_SMOTE_train, y_SMOTE_train)
# evaluate model performance
print "\nRUS - Train AUC : ",metrics.roc_auc_score(y_RUS_train, clf.predict(X_RUS_train))
print "RUS - Test AUC : ",metrics.roc_auc_score(y_RUS_test, clf.predict(X_RUS_test))
print "ROS - Train AUC : ",metrics.roc_auc_score(y_ROS_train, clf.predict(X_ROS_train))
print "ROS - Test AUC : ",metrics.roc_auc_score(y_ROS_test, clf.predict(X_ROS_test))
print "\nSMOTE - Train AUC : ",metrics.roc_auc_score(y_SMOTE_train, clf.predict(X_SMOTE_train))
print "SMOTE - Test AUC : ",metrics.roc_auc_score(y_SMOTE_test, clf.predict(X_SMOTE_test))
# -
# ### Conclusion
#
# Here random over sampling is performing better on both train and test sets. As a best practice, in real world use cases it is recommended to look at other matrics (such as confusion matrix) and apply business context or domain knowledge to assess true performance of model.
| jupyter_notebooks/machine_learning/ebook_mastering_ml_in_6_steps/Chapter_4_Code/Code/Handling_Imbalanced_Dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Accessing data from a web API
# One of the best things about "web 2.0" is more formal, external facing API's to allow developers to build tools on top of data. These APIs provide a reliable connection to data over a reliable protocol returning a reliable response in an easy, machine-readable format. There are tons of APIs out there. Sometimes you will need to register for a developer key (see the St. Louis Fed's API [here](https://research.stlouisfed.org/docs/api/)) and other times the APIs are free (see Coindesk's API [here](https://www.coindesk.com/api)). ProgrammableWeb has a great [directory](https://www.programmableweb.com/apis/directory).
#
# ## With great power comes great responsibility. We should be responsible and polite internet citizens when hitting websites programmatically
#
# Below we will use Coindesk's API to get daily prices for bitcoin. Coindesk's API, like many others, returns a JSON (javascript object notation) object, which we can easily turn into a dataframe. Here we will need the `json` and `requests` libraries.
# +
import datetime
import pandas as pd
import json
import requests
coindeskURL = 'https://api.coindesk.com/v1/bpi/historical/close.json?'
# from API
start = datetime.date(2019, 1 ,1)
end = datetime.date(2019, 7, 1)
url = f'{coindeskURL}start={start:%Y-%m-%d}&end={end:%Y-%m-%d}'
print(f'Hitting this url:{url}')
result = requests.get(url)
result.content
# -
# Digging into this a little bit, there are a few critical components to accessing data. Not all APIs will return the same structure of data. Many times you can look at what is returned by just hitting the URL with your browser. Let's try that
#
# [https://api.coindesk.com/v1/bpi/historical/close.json?start=2019-01-01&end=2019-07-01](https://api.coindesk.com/v1/bpi/historical/close.json?start=2019-01-01&end=2019-07-01)
#
# In this case we get:
# ```
# {
# "bpi":
# {
# "2019-01-01":3869.47,
# "2019-01-02":3941.2167,
# ...
# },
# "disclaimer": "This data was produced from the CoinDesk Bitcoin Price Index.",
# "time":
# {
# "updated":"Jul 2, 2019 00:03:00 UTC",
# "updatedISO":"2019-07-02T00:03:00+00:00"
# }
# }
# ```
# `bpi`, `disclaimer`, and `time` are all data that we can reference. The data we're primarily interested in is `bpi`. Different APIs can break up data differently, we will see examples in other APIs later.
# Let's take a look at the json result in python.
jsondata = json.loads(result.content)
jsondata
# We can then wrap that up in to a pandas dataframe! Note: pandas does have a `read_json` helper as well, but sometimes it is not well suited given the structure of the json output.
# note the json gets read with the disclaimer and time outputs as well
data = pd.read_json(result.content)
data
data = pd.DataFrame({'Bitcoin Price Index': jsondata['bpi']})
data
# %matplotlib inline
data.plot()
| notebooks/4_webapi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.10.0 (''.venvpythonexcel'': venv)'
# language: python
# name: python3
# ---
import pandas as pd
from numpy import nan
# ## File einlesen und info
# File einlesen,
# ursprünglich von https://en.wikipedia.org/wiki/Ky%C5%8Diku_kanji
df = pd.read_excel("kyouiku_kanji.xlsx", sheet_name="kyouiku kanji", na_filter=True)
df.info()
# ## filter lines mit kanji
# https://kanokidotorg.github.io/2022/02/16/How-to-filter-out-the-NaN-values-in-a-pandas-dataframe/
#filter
ft = df["#"].notna()
df = df.loc[ft, ].set_index("#")
df = df.rename(columns={"New (Shinjitai)" : "shinjitai", "Old (Kyūjitai)" : "kyuujitai", "Radical" : "radical", "Strokes" : "strokes", "Grade" : "grade" , "Year added": "year", "English meaning" : "meaning", "Readings" : "readings"})
df.info()
## Filter getestefe
tf = (df["strokes"] <= 3) | (df["grade"] == "5")
df.loc[tf, :]
| svadilfari_examples/kyouiku_kanji_pdtest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/IdealisticINTJ/TLDR-Summarizer/blob/main/TextSummarizer.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="QzKkKAM3NFdf" colab={"base_uri": "https://localhost:8080/"} outputId="dc07f5e3-ba5f-48b8-c779-a5c92d8dca49"
#libraries
import bs4 as bs
import urllib.request as url
import re
import nltk
nltk.download('punkt')
nltk.download('stopwords')
from nltk.tokenize import sent_tokenize, word_tokenize
from nltk.corpus import stopwords
from nltk.probability import FreqDist
import heapq
from string import punctuation
# + id="C4M7CqyGNKIy"
scraped_data = url.urlopen('https://en.wikipedia.org/wiki/Tourette_syndrome')
article = scraped_data.read()
parsed_article = bs.BeautifulSoup(article,'lxml')
paragraphs = parsed_article.find_all('p')
article_text = ""
for p in paragraphs:
article_text += p.text
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="2qcwZrZMOdox" outputId="eee3b462-b9e2-42df-a505-d90d6efc7c2e"
article_text
# + id="oxIcAcGKOsJw"
# remove square brackets and extra spaces
article_text = re.sub(r'\[[0-9]*\]', ' ', article_text)
article_text = re.sub(r'\s+', ' ', article_text)
# + id="j2Dd-wQsOtgV"
# remove special characters and digits
formatted_article_text = re.sub('[^a-zA-Z]', ' ', article_text )
formatted_article_text = re.sub(r'\s+', ' ', formatted_article_text)
# + id="XOQgTMtvOxRo"
sentence_list = nltk.sent_tokenize(article_text)
# + id="LOv9LSJdO0qa"
stopwords = nltk.corpus.stopwords.words('english')
# + id="uNjCz6ZhO3C4"
word_frequencies = {}
for word in nltk.word_tokenize(formatted_article_text):
if word not in stopwords and word not in punctuation:
if word not in word_frequencies.keys():
word_frequencies[word] = 1
else:
word_frequencies[word] += 1
# + colab={"base_uri": "https://localhost:8080/"} id="7DibWmAOO95A" outputId="08bfabd8-8b46-4332-e61c-d5a772200f35"
word_frequencies
# + id="i7mgRu_RO3KN"
maximum_frequncy = max(word_frequencies.values())
for word in word_frequencies.keys():
word_frequencies[word] = (word_frequencies[word]/maximum_frequncy)
# + colab={"base_uri": "https://localhost:8080/", "height": 328} id="a7Dp7YGiPVj7" outputId="68d3d184-9727-4168-bc19-ff0248720551"
frequency_dist = nltk.FreqDist(word_frequencies)
frequency_dist.plot(30)
# + id="XY5fifDoPcfB"
sentence_scores = {}
for sent in sentence_list:
for word in nltk.word_tokenize(sent.lower()):
if word in word_frequencies.keys():
if len(sent.split(' ')) < 30:
if sent not in sentence_scores.keys():
sentence_scores[sent] = word_frequencies[word]
else:
sentence_scores[sent] += word_frequencies[word]
# + colab={"base_uri": "https://localhost:8080/"} id="YzfO3nXgPgi2" outputId="876d3ac5-d563-4368-f46d-3bfa7957ddd5"
sentence_scores
# + id="Iv1zyL5tPmkT"
summary_sentences = heapq.nlargest(7, sentence_scores, key=sentence_scores.get)
summary = ' '.join(summary_sentences)
# + colab={"base_uri": "https://localhost:8080/", "height": 174} id="_itNgxabPp0R" outputId="75eb853c-a0e1-4380-fa9d-601a376561d2"
summary
| TextSummarizer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/matteuscruz/computer_vision_models/blob/main/YoloR.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="15hxxPw2nHfx" outputId="c2809a46-6ebc-4420-d797-561f0da6890c"
# Mount Google Drive
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="4Z4O49qknXIc" outputId="f6087a3a-bcdd-4e31-e5df-f798f59dc41b"
# Clone YOLOR
# !git clone https://github.com/augmentedstartups/yolor
# %cd yolor
# + colab={"base_uri": "https://localhost:8080/"} id="A8BcdHTmpAlo" outputId="bb836326-16ce-4448-9158-e5c8d1ae7374"
# Install Requirements
# !pip install -qr requirements.txt
# + colab={"base_uri": "https://localhost:8080/"} id="yFXZuQeEpS_z" outputId="c7c4e6e3-6d9b-4e3a-d6d1-3045c9b1d041"
# Clone and Install CUDA
# !git clone https://github.com/JunnYu/mish-cuda
# %cd mish-cuda
# !python setup.py build install
# + id="9ormoR-UqfBW" colab={"base_uri": "https://localhost:8080/"} outputId="5f64542e-e5fe-4b56-94ba-35f323dd8454"
# Clone and Install PyTorch
# !git clone http://github.com/fbcotter/pytorch_wavelets
# %cd pytorch_wavelets
# !pip install .
# + colab={"base_uri": "https://localhost:8080/"} id="Y3lRhJ_o67RC" outputId="25f677c6-3fce-4755-c0a4-ea83207891ac"
# %cd ..
# + id="buGLdC6a7AN6" colab={"base_uri": "https://localhost:8080/"} outputId="d57c86e7-aaad-48e3-b467-46866ba90145"
# %cd ..
# + colab={"base_uri": "https://localhost:8080/"} id="R_jD986L6ODF" outputId="c59066ea-f388-46d8-bcaa-6137c901950a"
# !bash scripts/get_pretrain.sh
# + colab={"base_uri": "https://localhost:8080/"} id="Yn2QLwkE6zzX" outputId="d03573a6-8caa-478e-bd7c-1980a13cefd7"
# !python detect.py --source inference/images/horses.jpg --cfg cfg/yolor_p6.cfg --weights yolor_p6.pt --conf 0.25 --img-size 1280 --device 0 --output /content/drive/MyDrive/YoloR
# + colab={"base_uri": "https://localhost:8080/", "height": 529} id="hOiBCanw7-Ez" outputId="1abe27b5-3dce-4f9d-afd8-4420b7f3be2c"
import glob
from IPython.display import Image, display
for imageName in glob.glob('/content/drive/MyDrive/YoloR/*.jpg'):
display(Image(filename=imageName))
print('\n')
# + colab={"base_uri": "https://localhost:8080/"} id="DKEsyR_V-vHm" outputId="106fac59-0137-4ccc-fbf1-58c1f2ad7b3d"
# !python detect.py --source /content/drive/MyDrive/videos_originais/video05.mp4 --cfg cfg/yolor_p6.cfg --weights yolor_p6.pt --conf 0.25 --img-size 1280 --device 0 --output /content/drive/MyDrive/YoloR
# + id="4gOCCGBh2tcR"
| YoloR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TECHMIMO
#
#
# Autor: <NAME>
#
# Seguem alguns recados para ajudá-los e para contribuir com o curso:
#
# - Fiquem à vontade para me contatar pelo Linkdin, costumo responder por lá também: https://www.linkedin.com/in/rafael-pereira-da-silva-23890799/
#
# - Fiquem a vontade para compartilharem o certificado do curso no Linkedin. Eu costumo curtir e comentar para dar mais credibilidade
#
# - Vocês podem usar esses notebooks para resolver os exercícios e desafios
#
# - Não se esqueçam de avaliar o curso e dar feedback, eu costumo criar conteúdos baseado nas demandas de vocês
# - Se tiverem gostando do curso, recomendem aos amigos, pois isso também ajuda a impulsionar e a crescer a comunidade
# - Bons estudos e grande abraços!
#
# ## Sobre o Material
#
# Este material é parte do curso **Python para Engenheiros e Cientistas** e está sobre a licença MIT.
# Caso você tenha interesse em conhecer o curso, acesso o link abaixo.
#
#
# https://www.udemy.com/course/python-para-engenheiros-e-cientistas/?referralCode=FA25B2C024FF2D3F8232
#
#
# Algumas aulas, exercícios e desafios estarão sem códigos propositalmente para que o aluno possa preencher.
#
# # Seção 3 - Condionais e loops
#
# Palavras chave: *Control flow* ou *Control structures*
#
# Essa aula é inspirada em: https://docs.python.org/3/tutorial/controlflow.html
# ## 3.1 Condicional *if*
#
# ### Construções
# - **if** condição**:**
# - **elif** condição**:**
# - **else:**
#
# ### Operações booleanas
# Além dos comparadores, também podemos fazer operações booleanas. Elas são úteis em estruturas condicionais. Duas delas são: **or** e **and**.
#
#
# Acesse: https://docs.python.org/3/library/stdtypes.html#
# +
x = 5
if x > 0:
print('positivo')
elif x < 0:
print('negativo')
else:
print('nenhuma alternativa')
# +
x = 'Matemática'
if x == 'Matemática':
print(x)
# +
x = 5
if type(x) is not str:
print(x)
# +
x = -10
if x > 4 or x < 0:
print(x)
# -
# ## 3.2 Loop *for*
#
#
# - **for** variável **in** o_que_será_iterado**:**
#
# Há uma série de tipos que podem ser utilizado como iteradores, como:
# - range --> seus elementos são inteiros
# - list --> seus elementos são os que estão contidos na lista
# - enumerate --> seus elementos são tuplas que contem inteiros e valores
#
#
#
#
# +
lista = ['a',2,'b',4,5,5,'w']
for i in range(len(lista)):
print(i)
# +
lista = ['a',2,'b',4,5,5,'w']
for i in enumerate(lista):
print(i[1])
# -
# ## 3.3 Loop *while*
# - **while** condição**:**
#
# Enquanto a condição for verdadeira, o programa executará os comandos
#
# Uma noção sobre tipos booleanos é interessante para executar esse comando
#
# +
x = 10
while x > 5:
print(x)
#x = x - 1
x += 1
if x > 20:
break
# -
# ## 3.4 Exercício 1 -
# Crie um condicional que, dada uma variável, faça as operações:
# - Se for número positivo, retorna "Número positivo"
# - Se for número negativo, retorna "Número negativo"
# - Se for zero, retorna "zero"
# - Se não for número, retorna "Não é um número"
#
#
# +
# A ser resolvido pelo aluno (assista o vídeo com a solução)
# -
# ## 3.5 Exercício 2 -
# Faça um loop **for** para criar uma lista que intercale os valores das listas a seguir:
# - valores = [1,2,3,4,5]
# - letras = ['a','b','c','d','e']
#
# Dica: você pode utilizar o método .append() que adiciona um elemento ao final da lista.
# +
# A ser resolvido pelo aluno (assista o vídeo com a solução)
# -
# ## 3.6 Exercício 3 -
# Dado um número inteiro, faça um operador while que calcule o fatorial desse número. Dicas: utilize as operações ** *=** e também **-=**
# +
# A ser resolvido pelo aluno (assista o vídeo com a solução)
# -
# ## 3.7 Desafio 1 - Números primos
# Crie uma lista com os cinquenta primeiros números primos.
#
# Dica: pode ser utilizado o comando break e continue
# +
# A ser resolvido pelo aluno (assista o vídeo com a solução)
# -
# ## 3.8 Desafio 2 - Combinação
# Crie uma lista de todas as palavras formadas com as letras: ARARA
#
# Dica: o problema se trata de uma permutação com elementos repetidos.
#
#
# $$
# P^{3,2}_{5} = \frac{5!}{3!2!} = 10
# $$
#
#
# Anotações conforme: IEZZI, Gelson *et al*. Matemática: 1a série. São Paulo: Atual, 1981.
#
# Para escrever fórmulas no Markdown: http://luvxuan.top/posts/Markdown-math/
# +
# A ser resolvido pelo aluno
# -
| Notebooks/Secao-03-Condicionais_e_loops.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fine-tuning a Model on Your Own Data
#
# [](https://colab.research.google.com/github/deepset-ai/haystack/blob/master/tutorials/Tutorial2_Finetune_a_model_on_your_data.ipynb)
#
# For many use cases it is sufficient to just use one of the existing public models that were trained on SQuAD or other public QA datasets (e.g. Natural Questions).
# However, if you have domain-specific questions, fine-tuning your model on custom examples will very likely boost your performance.
# While this varies by domain, we saw that ~ 2000 examples can easily increase performance by +5-20%.
#
# This tutorial shows you how to fine-tune a pretrained model on your own dataset.
# ### Prepare environment
#
# #### Colab: Enable the GPU runtime
# Make sure you enable the GPU runtime to experience decent speed in this tutorial.
# **Runtime -> Change Runtime type -> Hardware accelerator -> GPU**
#
# <img src="https://raw.githubusercontent.com/deepset-ai/haystack/master/docs/_src/img/colab_gpu_runtime.jpg">
# + pycharm={"name": "#%%\n"}
# Make sure you have a GPU running
# !nvidia-smi
# +
# Install the latest release of Haystack in your own environment
# #! pip install farm-haystack
# Install the latest master of Haystack
# !pip install grpcio-tools==1.34.1
# !pip install git+https://github.com/deepset-ai/haystack.git
# If you run this notebook on Google Colab, you might need to
# restart the runtime after installing haystack.
# + pycharm={"name": "#%%\n"}
from haystack.nodes import FARMReader
# -
#
# ## Create Training Data
#
# There are two ways to generate training data
#
# 1. **Annotation**: You can use the [annotation tool](https://haystack.deepset.ai/guides/annotation) to label your data, i.e. highlighting answers to your questions in a document. The tool supports structuring your workflow with organizations, projects, and users. The labels can be exported in SQuAD format that is compatible for training with Haystack.
#
# 
#
# 2. **Feedback**: For production systems, you can collect training data from direct user feedback via Haystack's [REST API interface](https://github.com/deepset-ai/haystack#rest-api). This includes a customizable user feedback API for providing feedback on the answer returned by the API. The API provides a feedback export endpoint to obtain the feedback data for fine-tuning your model further.
#
#
# ## Fine-tune your model
#
# Once you have collected training data, you can fine-tune your base models.
# We initialize a reader as a base model and fine-tune it on our own custom dataset (should be in SQuAD-like format).
# We recommend using a base model that was trained on SQuAD or a similar QA dataset before to benefit from Transfer Learning effects.
#
# **Recommendation**: Run training on a GPU.
# If you are using Colab: Enable this in the menu "Runtime" > "Change Runtime type" > Select "GPU" in dropdown.
# Then change the `use_gpu` arguments below to `True`
# + pycharm={"name": "#%%\n"}
reader = FARMReader(model_name_or_path="distilbert-base-uncased-distilled-squad", use_gpu=True)
data_dir = "data/squad20"
# data_dir = "PATH/TO_YOUR/TRAIN_DATA"
reader.train(data_dir=data_dir, train_filename="dev-v2.0.json", use_gpu=True, n_epochs=1, save_dir="my_model")
# + pycharm={"name": "#%%\n"}
# Saving the model happens automatically at the end of training into the `save_dir` you specified
# However, you could also save a reader manually again via:
reader.save(directory="my_model")
# + pycharm={"name": "#%%\n"}
# If you want to load it at a later point, just do:
new_reader = FARMReader(model_name_or_path="my_model")
# + [markdown] pycharm={"name": "#%% md\n"}
# ## About us
#
# This [Haystack](https://github.com/deepset-ai/haystack/) notebook was made with love by [deepset](https://deepset.ai/) in Berlin, Germany
#
# We bring NLP to the industry via open source!
# Our focus: Industry specific language models & large scale QA systems.
#
# Some of our other work:
# - [German BERT](https://deepset.ai/german-bert)
# - [GermanQuAD and GermanDPR](https://deepset.ai/germanquad)
# - [FARM](https://github.com/deepset-ai/FARM)
#
# Get in touch:
# [Twitter](https://twitter.com/deepset_ai) | [LinkedIn](https://www.linkedin.com/company/deepset-ai/) | [Slack](https://haystack.deepset.ai/community/join) | [GitHub Discussions](https://github.com/deepset-ai/haystack/discussions) | [Website](https://deepset.ai)
#
# By the way: [we're hiring!](https://www.deepset.ai/jobs)
| tutorials/Tutorial2_Finetune_a_model_on_your_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:phd]
# language: python
# name: conda-env-phd-py
# ---
name = '2017-03-24-climate-model-output'
title = 'Two ways of preparing climate model output for analysis'
tags = 'numpy, iris'
author = '<NAME>'
# +
from nb_tools import connect_notebook_to_post
from IPython.core.display import HTML, Image
html = connect_notebook_to_post(name, title, tags, author)
# -
# Today one of the group members asked for help with reading climate model output and preparing it for data analysis.
# This notebook shows a couple of ways of doing that with the help of `numpy` and `iris` Python packages.
# Luckily, the model output is quite small and stored in a simple ASCII file. However, it has some properties that can be a hurdle for a programming novice.
# ### Download the data from UEA archive
# We start with downloading data from a given link.
URL = 'https://raw.githubusercontent.com/ueapy/ueapy.github.io/src/content/data/run1_U_60N_10hPa.dat'
# Instead of copy-pasting the contents manually, we are going to use Python's standard library and download the file, making this part of scientific analysis more [reproducible](https://www.youtube.com/playlist?list=PLYCpMb24GpOC704uO9svUrihl-HY1tTJJ).
from urllib.request import urlretrieve
# To organise data and code folders, we also import `os` module.
import os
# +
datadir = os.path.join(os.path.pardir, 'data') # directory is one level up
if not os.path.isdir(datadir):
# if the directory does not exist, create it
os.makedirs(datadir)
# File with data
fname = os.path.join(datadir, 'data.dat')
# -
# Now that we have a directory to store data, we can save the model output there.
urlretrieve(URL, fname)
# ### Read the data using `numpy`
# Since the data are purely numeric, we use `numpy` module.
import numpy as np
data = np.genfromtxt(fname)
data.shape
data
# For some reason the data are stored in 6 columns by 1500 rows, which in total is 9000 values.
# We know *a priori* that the file contains **75 years** of data written every **third** day, and the climate model's calendar is **360-day** calendar. Hence, we have 120 values per year:
data.shape[0] * data.shape[1] / 75
# Keeping data in $1500\times6$ array does not seem to be useful, so we make it 1-D:
data = data.flatten()
data.shape
# #### Wrap it up in a function
# To make the code above reusable, we create the following function to get data.
def get_model_data(url=URL, fname='climate_model_data.dat', force_download=False):
"""
Function to download climate model output from UEA server
Parameters
---------
url : string (optional)
web location of the data
fname : string (optional)
full path to save the data
force_download : bool (optional)
if True, force redownload of data
Returns
-------
data : numpy.ndarray
1-D array of data
"""
if not os.path.exists(fname) or force_download:
urlretrieve(URL, fname)
# print('Downloading...')
data = np.genfromtxt(fname)
return data.flatten()
data = get_model_data()
# ## 1. Plain NumPy
# ### Reshape the array to YEARS $\times$ DAYS
# Now we transform the array into a more useful shape.
NDAYS = 120 # the number of 3-day periods in a 360-day year
NYEARS = 75 # the total number of years
data_yd = data.reshape((NYEARS, NDAYS))
print(data_yd.shape)
# For example, this is a value of $u$-wind on 30 January of the last year:
data_yd[-1, 10]
# ### Select only winter months
# What if we want to extract only winter data? We can't use the first winter, because it's incomplete: it only has January and February. So the first *winter* period will comprise December data from the year 1:
data_yd[0, -10:]
# plus January and February data from the year 2:
data_yd[1, :20]
# To join them, we can use `numpy.concatentate()` function:
np.concatenate([data_yd[0, -10:], data_yd[1, :20]])
# And of course we can apply the same logic to the whole dataset:
data_djf = np.concatenate([data_yd[:-1, -10:], data_yd[1:, :20]], axis=1)
print(data_djf.shape)
# ### Selecting years by a certain criterion
# > How to find winters when at least 20 days of constant wind direction followed by its change?
# Here we are just applying [this](http://stackoverflow.com/a/24343375/5365232) answer on Stack Overflow to our problem.
for i, yr in enumerate(data_djf):
condition = yr > 0
lens_true = np.diff(np.where(np.concatenate(([condition[0]], condition[:-1] != condition[1:], [True])))[0])[::2]
if 20 <= lens_true.max() < 30:
print(i, lens_true.max())
# ## 2. What if you want to use labelled arrays?
# In the above example `numpy`'s capabilities were probably enough. But when you have more dimensions and data are more complex, it is mostly always better to use labelled arrays and all the great functionality offered by such libraries as `xarray` or `iris`.
# We show how `iris` library can be used with the same dataset. We chose `iris`, mostly because it can handle non-standard calendars, like `360-day` one.
# To create an appropriate time coordinate, we will use `iris` companion package - [`cf_units`](https://github.com/SciTools/cf_units).
import cf_units
import iris
DAYS_PER_YEAR = 360
t_unit = cf_units.Unit('days since 0001-01-01 00:00:00',
calendar='360_day')
t_coord = iris.coords.DimCoord(np.arange(0, DAYS_PER_YEAR * NYEARS, 3),
units=t_unit,
standard_name='time')
# Now we can attach the newly created time coordinate to the data themselves by creating an iris cube:
cube = iris.cube.Cube(data=data,
units='m/s',
dim_coords_and_dims=[(t_coord, 0)])
cube.rename('eastward_wind')
print(cube)
# #### Calculate seasonal means
# Source: http://scitools.org.uk/iris/docs/latest/userguide/cube_statistics.html
# Since we now have a labelled aray with appropriate metadata, we can use `iris` to make statistical analysis easier and make the code more readable.
import iris.coord_categorisation
iris.coord_categorisation.add_season(cube, 'time', name='clim_season')
iris.coord_categorisation.add_season_year(cube, 'time', name='season_year')
print(cube)
cube.coord('clim_season')
for season, year in zip(cube.coord('clim_season')[:100:10].points,
cube.coord('season_year')[:100:10].points):
print('{} {}'.format(season, year))
annual_seasonal_mean = cube.aggregated_by(['clim_season', 'season_year'],
iris.analysis.MEAN)
print(annual_seasonal_mean)
HTML(html)
| content/notebooks/2017-03-24-climate-model-output.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import random
random.random()
num = random.random() * 100
num
round(num)
int(num)
# +
print(40* "*")
print("Bem vindo ao jogo da adivinhação")
print(40* "*")
num_secreto = round(random.random() * 100)
chances = 4
rodada = 1
# +
print("Você tem", chances, "chances para acertar!")
for rodada in range(1, chances + 1):
print("Tentativa {} de {}".format(rodada, chances))
chute = int(input("Digite o seu número entre 1 e 100: "))
print("Você digitou", chute, end='.')
if chute < 1 or chute > 100:
print("\nEsse número não é válido")
continue
acertou = chute == num_secreto
maior = chute > num_secreto
menor = chute < num_secreto
if acertou:
print("\nVocê acertou! :)")
break
else:
if maior:
print("\nSeu chute foi maior do que o número. Você errou! :(")
elif menor:
print("\nSeu chute foi menor do que o número. Você errou! :(")
print(f'O número secreto era {num_secreto}')
print("Fim de Jogo")
# -
# ### função random.randrange()
# +
print(40* "*")
print("Bem vindo ao jogo da adivinhação")
print(40* "*")
num_secreto = round(random.randrange(1,101))
chances = 5
rodada = 1
print("Você tem", chances, "chances para acertar!")
for rodada in range(1, chances + 1):
print("Tentativa {} de {}".format(rodada, chances))
chute = int(input("Digite o seu número entre 1 e 100: "))
print("Você digitou", chute, end='.')
if chute < 1 or chute > 100:
print("\nEsse número não é válido")
continue
acertou = chute == num_secreto
maior = chute > num_secreto
menor = chute < num_secreto
if acertou:
print("\nVocê acertou! :)")
break
else:
if maior:
print("\nSeu chute foi maior do que o número. Você errou! :(")
elif menor:
print("\nSeu chute foi menor do que o número. Você errou! :(")
print(f'O número secreto era {num_secreto}!')
print("Fim de Jogo")
# +
## random.randrange(0,101,2) Vai do 0 ao 100 (último valor -1), de 2 em 2.
# -
| PY_ 01_Intro_parte_1/jogo_da_adivinhacao_pt5_random.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # Module 4 Required Coding Activity
# Introduction to Python (Unit 2) Fundamentals
#
# This is an activity based on code similar to the Jupyter Notebook **`Practice_MOD04_IntroPy.ipynb`** which you may have completed.
#
# | **Assignment Requirements** |
# |:-------------------------------|
# | **NOTE:** This program requires **`print`** output and code syntax used in module 4 such as variable assignment, **`while`**, **`open`** keywords, **`.split()`**, **`.readline()`**, **`.seek()`**, **`.write()`**, **`.close()`** methods |
#
# ## The Weather
# #### Create a program that:
# - imports and opens a file
# - appends additional data to a file
# - reads from the file to displays each city name and month average high temperature in Celsius
#
# **Output:** The output should resemble the following
# ```
# City of Beijing month ave: highest high is 30.9 Celsius
# City of Cairo month ave: highest high is 34.7 Celsius
# City of London month ave: highest high is 23.5 Celsius
# City of Nairobi month ave: highest high is 26.3 Celsius
# City of New York City month ave: highest high is 28.9 Celsius
# City of Sydney month ave: highest high is 26.5 Celsius
# City of Tokyo month ave: highest high is 30.8 Celsius
# City of Rio De Janeiro month ave: highest high is 30.0 Celsius
# ```
# all of the above text output is generated from the file
# the only strings are *hard coded*:
# - "is"
# - "of"
# - "Celsius"
#
#
#
# #### import the file into the Jupyter Notebook environment
# - use `!curl` to download https://raw.githubusercontent.com/MicrosoftLearning/intropython/master/world_temp_mean.csv as `mean_temp.txt`
#
#
# ```python
# # [ ] The Weather: import world_mean_team.csv as mean_temp.txt into the Jupyter notebook
#
#
# ```
#
# #### Add the weather for Rio
# 1. Open the file in append plus mode (**`'a+'`**)
# 2. Write a new line for Rio de Janeiro `"Rio de Janeiro,Brazil,30.0,18.0\n"`
#
#
# #### Grab the column headings
# 1. use .seek() to move the pointer to the beginning of the file
# 2. read the first line of text into a variable called: `headings`
# 3. convert `headings` to a list using **`.split(',')`** which splits on each comma
#
#
# ```python
# # [ ] The Weather: open file, read/print first line, convert line to list (splitting on comma)
#
#
# ```
#
# #### Read the remaining lines from the file using a while loop
# 1. assign remaining lines to a **`city_temp`** variable
# 2. convert the city_temp to a list using **`.split(',')`** for each **`.readline()`** in the loop
# 3. print each city & the highest monthly average temperature
# 4. close mean_temps
#
# >Tips & Hints:
# - print **`headings`** to determine indexes to use for the final output (what is in headings[0], [1], [2]..?)
# - the city_temp data follows the order of the headings (city_temp[0] is described by headings[0])
# - The output should look like: **`"month ave: highest high" for Beijing is 30.9 Celsius`**
# - convert `city_temp` to lists with `.split(',')`
#
#
# ```python
# # [ ] The Weather: use while loop to print city and highest monthly average temp in celsius
#
#
# ```
#
# ### Combine All Code into one cell, Copy and Submit on edX
#
# +
# [] create The Weather
# [] copy and paste in edX assignment page
# !curl https://raw.githubusercontent.com/MicrosoftLearning/intropython/master/world_temp_mean.csv -o mean_temp.txt
mean_temp_file = open('mean_temp.txt', 'a+')
mean_temp_file.write("Rio de Janeiro,Brazil,30.0,18.0\n")
mean_temp_file.seek(0)
headings = mean_temp_file.readline().strip()
headings_list = headings.split(',')
while mean_temp_file:
city_temp = mean_temp_file.readline().strip()
city_temp_list = city_temp.split(',')
if "" in city_temp_list:
break
else:
print(headings_list[2],"for",city_temp_list[0],"is",city_temp_list[2],"Celsius")
mean_temp_file.close()
# -
# Submit this by creating a python file (.py) and submitting it in D2L. Be sure to test that it works. Know that For this to work correctly in Python rather than Jupyter, you would need to switch to using import os rather than !curl. To convert !curl to run in the normal python interpreter try a method such as importing the os library and calling os.system(cmd) with your shell command in the cmd variable.
#
| Python Fundamentals/Module_4_Required_Code_Python_Fundamentals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="LsyWEfBiiV3P"
# # ML 101
#
# ## Comparing two classifiers
#
# The choice of a [statistical hypothesis test](https://machinelearningmastery.com/statistical-hypothesis-tests/) is a challenging open problem for interpreting machine learning results.
#
# Model Evaluation is the subsidiary part of the model development process. It is the phase that is decided whether the model performs better. Therefore, it is critical to consider the model outcomes according to every possible evaluation method. Applying different methods can provide different perspectives.
#
# One of the mistakes while evaluating the classification model is considering only the true cases. It means that looking for only how the model estimates actual cases correctly. Therefore, when the results are unsatisfactory, people try to apply different methods or different variations to get the result that makes them satisfied, without considering the main reason for that result. It shouldn’t be forgotten the accuracy also depends on the false predictions as much as it depends on the true predictions. Thus, false predictions also have to be taken into consideration before rendering a certain verdict. These are the predictions which we want to be as minimum as possible. The metrics called Recall and Precision slightly explain the performance of the positive classes (or negative) by considering the false cases too. But, what I try to say is, the false positives and the false negatives should be compared like they are compared for the true cases. This is where the McNemar test should be used for obtaining a probability of difference between the cases of false negative and false positives.
#
# McNemar’s test is applied to $2\times 2$ contingency tables to find whether row and column marginal frequencies are equal for paired samples. What row and column marginal frequencies mean for confusion matrices is the number of false predictions for both positive and negative classes. It uses the Chi-Square distribution to determine the probability of difference.
#
# 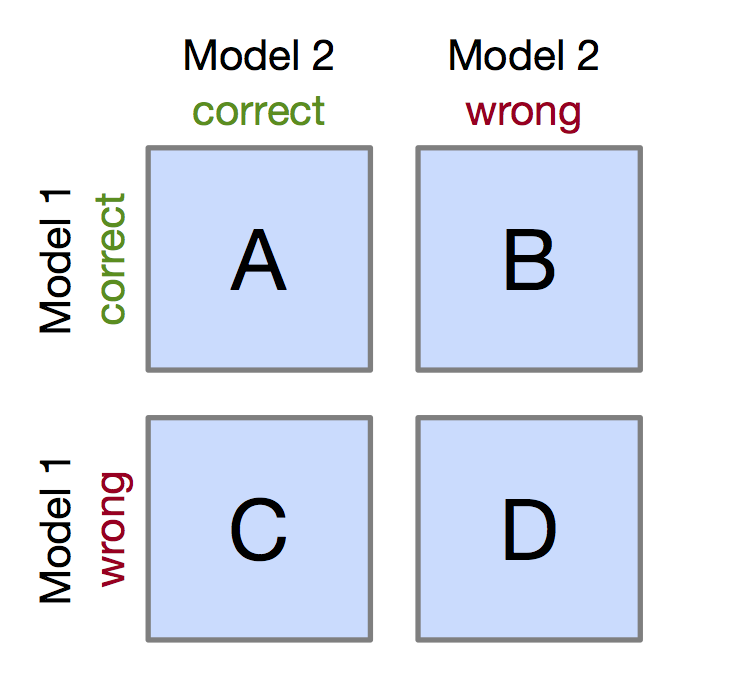
#
# + id="CDE8CTnLiThN" executionInfo={"status": "ok", "timestamp": 1643393949986, "user_tz": 0, "elapsed": 1216, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjyA-_qhOJOz0fgt8nLXmM1Vm0Hkd1Y6B77dsqd0do=s64", "userId": "09349029796619273737"}}
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import seaborn as sns
from mlxtend.evaluate import mcnemar_table
from mlxtend.evaluate import mcnemar
from sklearn import datasets
from sklearn.model_selection import train_test_split
from sklearn.metrics import matthews_corrcoef
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
# + id="p_Xd9Hsui4c8" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643393990352, "user_tz": 0, "elapsed": 244, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjyA-_qhOJOz0fgt8nLXmM1Vm0Hkd1Y6B77dsqd0do=s64", "userId": "09349029796619273737"}} outputId="a98723e5-f850-4360-8445-792ba8bb27d4"
iris = datasets.load_iris()
X = iris.data
y = iris.target
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42, stratify=y)
# Logistic Regression
clf = LogisticRegression().fit(X_train, y_train)
y_pred_lr = clf.predict(X_test)
m = matthews_corrcoef(y_test, y_pred_lr)
print(f'LR MCC {m}')
# Naive Bayes
clf = GaussianNB().fit(X_train, y_train)
y_pred_nb = clf.predict(X_test)
m = matthews_corrcoef(y_test, y_pred_nb)
print(f'NB MCC {m}')
# SVM
clf = SVC(probability=True, kernel='linear').fit(X_train,y_train)
y_pred_svm = clf.predict(X_test)
m = matthews_corrcoef(y_test, y_pred_svm)
print(f'SVM MCC {m}')
# + id="DGzbSjrqkUEB" colab={"base_uri": "https://localhost:8080/", "height": 283} executionInfo={"status": "ok", "timestamp": 1643394054991, "user_tz": 0, "elapsed": 611, "user": {"displayName": "M\u00e1<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjyA-_qhOJOz0fgt8nLXmM1Vm0Hkd1Y6B77dsqd0do=s64", "userId": "09349029796619273737"}} outputId="7f492ae9-c9cd-4a8c-97df-2f806b169df9"
tb = mcnemar_table(y_target=y_test, y_model1=y_pred_nb, y_model2=y_pred_svm)
sns.heatmap(tb, annot=True)
# + id="Nu6m4wkzkt1i" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643394087672, "user_tz": 0, "elapsed": 232, "user": {"displayName": "M\u00e1<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjyA-_qhOJOz0fgt8nLXmM1Vm0Hkd1Y6B77dsqd0do=s64", "userId": "09349029796619273737"}} outputId="8028a540-b6f6-4484-c04b-7734fb8466cd"
chi2, p = mcnemar(ary=tb, exact=True)
print('chi-squared:', chi2)
print('p-value:', p)
alpha = 0.05
if p < alpha:
print('The models are significantly different')
else:
print('The models are similar')
| classes/03 evaluation/03_evaluation_03.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Plot Comulative Distribution Of Sportive Behavior Over Time
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib notebook
from sensible_raw.loaders import loader
from world_viewer.cns_world import CNSWorld
from world_viewer.synthetic_world import SyntheticWorld
from world_viewer.glasses import Glasses
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm, PowerNorm
import math
import pandas as pd
import numpy as np
#import dask.dataframe as dd
import time
import seaborn as sns
# -
# load data and restict timeseries
# data from "PreprocessOpinions/FitnessAsBehavior.ipynb"
data = pd.read_pickle("data/op_fitness.pkl")
#data.reset_index(inplace=True)
opinion = "op_fitness"
data = data[data.time >= CNSWorld.CNS_TIME_BEGIN]
data = data[data.time <= CNSWorld.CNS_TIME_END]
data.head()
# +
# calc cummulative distribution function
def cdf_from_data(data, cdfx):
size_data = len(data)
y_values = []
for i in cdfx:
# all the values in data less than the ith value in x_values
temp = data[data <= i]
# fraction of that value with respect to the size of the x_values
value = temp.size / size_data
# pushing the value in the y_values
y_values.append(value)
# return both x and y values
return pd.DataFrame({'x':cdfx, 'cdf':y_values}).set_index("x")
cdfx = np.linspace(start=0,stop=4,num=400)
cdf = data.groupby("time")[opinion + "_abs"].apply(lambda d: cdf_from_data(d, cdfx))#
# +
# load cdf if previously calculated
#cdf = pd.read_pickle("tmp/cdf_fitness.pkl")
# +
# plot cdf as heatmap (fig.: 3.3)
fig, ax = plt.subplots(1,1)
num_ticks = 5
# the index of the position of yticks
yticks = np.linspace(0, len(cdfx)-1, num_ticks, dtype=np.int)
# the content of labels of these yticks
yticklabels = [round(cdfx[idx]) for idx in yticks]
cmap = sns.cubehelix_palette(60, hue=0.05, rot=0, light=0.9, dark=0, as_cmap=True)
ax = sns.heatmap(df2, cmap=cmap, xticklabels=80, yticklabels=yticklabels, vmin=0.4, vmax=1, cbar_kws={'label': 'cumulative distribution function'})#, norm=LogNorm(vmin=0.1, vmax=1))#, , cbar_kws={"ticks": cbar_ticks})
#ax.hlines([300], *ax.get_xlim(), linestyles="dashed")
ax.set_yticks(yticks)
ax.invert_yaxis()
plt.xticks(rotation=70)
plt.yticks(rotation=0)
plt.ylabel(r"$\bar b(t)$")
#ax.set_yscale('log')
#sns.heatmap(cdf.cdf, annot=False)
fig.savefig("test.png" , dpi=600, bbox_inches='tight')
# +
# plot cdf for singe timestep
fig, ax = plt.subplots(1,1)
ax.plot(cdf.loc["2014-02-09"].reset_index().x, 1-cdf.loc["2014-11-30","cdf"].values)
ax.set_yscale('log')
| CNS_CDF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/daffamilliano/Data-Mining/blob/main/Apriori.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="nxvqADBH55vo"
# **Menganalisa Pembelian Barang Menggunakan Algoritma Apriori**
#
#
# + [markdown] id="h-mHoMbP6jqW"
# Teman saya yang bernama Joko ingin membuat acara diskon besar-besaran pada toko online nya. Namun, Ia bingung dengan barang apa yang sering dibeli para pembelinya. Disini saya memberitahu kepada Joko. "Sepertinya masalah ini bisa dipecahkan dengan algoritma apriori deh" Ucap saya. Lalu Joko meminta saya untuk memecahkan masalahnya tersebut.
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="LHdNwCs0gaZ_" outputId="6aa3605b-c7a6-4e64-93d7-72a062dfbca6"
# Pertama kita load dataset transaksi barang tersebut
df = pd.read_csv('drive/MyDrive/Dataset/Apriori.csv')
df.head()
# + id="aJuTtVQ-nWIk" colab={"base_uri": "https://localhost:8080/"} outputId="d3d19b11-afd7-4013-847e-ec5a28ea77fe"
# Menginstall alat tempur untuk memecahkan masalah joko
# !pip install apyori
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from apyori import apriori
# + [markdown] id="DyF1ac2i50cC"
# Ketika kita sudah menginstall alat tempur, kita harus menghancurkan lawannya alias memecahkan masalahnya.
# + id="4rdFwMPug7ys"
# Membuat "data" untuk tabel yang hanya berisi Barang dari dataset sebelumnya
data=df.drop(['ID_Transaksi'],axis=1)
# + colab={"base_uri": "https://localhost:8080/"} id="ZXak3fpSquZc" outputId="19d38c1e-0454-462b-e1d4-ecd7030a5796"
# Lalu kita pastikan lagi hanya tinggal barang, sebelum nya dengan kolom 2 dan baris 11
data.shape
# + colab={"base_uri": "https://localhost:8080/"} id="zXkE3MsxhDD2" outputId="488de734-0e0a-47c1-9c79-652be9af89f0"
# Membuat list dalam list dari transaksi pembelian barang
records = []
for i in range(data.shape[0]):
records.append([str(data.values[i,j]).split(',') for j in range(data.shape[1])])
trx = [[] for trx in range(len(records))]
for i in range(len(records)):
for j in records[i][0]:
trx[i].append(j)
trx
# + id="judv0A72hFWf"
association_rules = apriori(trx)
association_results = association_rules
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="kifdj8JXrmYo" outputId="81e01d89-d219-4e14-efc6-9413035d2593"
# Menampilkan hasil asosiasi dari item
pd.set_option('max_colwidth', 1000)
Result=pd.DataFrame(columns=['Rule','Support','Confidence'])
for item in association_results:
pair = item[2]
for i in pair:
items = str([x for x in i[0]])
if i[3]!=1:
Result=Result.append({
'Rule':str([x for x in i[0]])+ " -> " +str([x for x in i[1]]),
'Support':str(round(item[1]*100,2))+'%',
'Confidence':str(round(i[2] *100,2))+'%'
},ignore_index=True)
Result
# + [markdown] id="EBUjML5U8lSC"
# dari daftar diatas kita dapat mencari nilai confidence nya paling besar yaitu :
#
# [' Asparagus'] -> [' Beans'] 27.27% 100.0%
#
# [' Broccoli'] -> [' Green Peppers'] 27.27% 100.0%
#
# [' Asparagus', 'Squash'] -> [' Beans'] 18.18% 100.0%
#
# ['Squash', ' Broccoli'] -> [' Green Peppers'] 18.18% 100.0%
#
# [' Green Peppers', 'Squash'] -> [' Broccoli'] 18.18% 100.0%
#
# [' Corn', 'Broccoli'] -> [' Green Peppers'] 18.18% 100.0%
#
# ['Broccoli', ' Tomatoes'] -> [' Green Peppers'] 18.18% 100.0%
#
# Kita dapat melihat bahwa ketika orang membeli Asparagus berarti mereka juga membeli Beans dan seterusnya.
#
# Masalah joko pun terselesaikan dan tidak lagi bingung untuk menjual barang yang ingin ia diskon.
| Apriori.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Load Data from CSVs
# +
import unicodecsv
def read_csv(filename):
with open(filename, 'rb') as f:
reader = unicodecsv.DictReader(f)
return list(reader)
# +
## variaveis que armazenam as funcoes para abrir as tabelas.
enrollments = read_csv('enrollments.csv')
daily_engagement = read_csv('daily_engagement.csv')
project_submissions = read_csv('project_submissions.csv')
# -
# testa a tabela e mostra as coluna de enrollments[0]
enrollments[0]
# testa a tabela e mostra as coluna de daily_engagement[0]
daily_engagement[0]
# testa a tabela e mostra as coluna de project_submissions[0]
project_submissions[0]
# ## Fixing Data Types
#
# - conversao de dados
# +
from datetime import datetime as dt
# Takes a date as a string, and returns a Python datetime object.
# If there is no date given, returns None
def parse_date(date):
if date == '':
return None
else:
return dt.strptime(date, '%Y-%m-%d')
# Takes a string which is either an empty string or represents an integer,
# and returns an int or None.
def parse_maybe_int(i):
if i == '':
return None
else:
return int(i)
# +
# Clean up the data types in the enrollments table
# percorre linha por linha da tabela ENROLLMENTS e tenta converter o dado usando as funcoes acima
for enrollment in enrollments:
enrollment['cancel_date'] = parse_date(enrollment['cancel_date'])
enrollment['days_to_cancel'] = parse_maybe_int(enrollment['days_to_cancel'])
enrollment['is_canceled'] = enrollment['is_canceled'] == 'True'
enrollment['is_udacity'] = enrollment['is_udacity'] == 'True'
enrollment['join_date'] = parse_date(enrollment['join_date'])
enrollments[0]
# +
# Clean up the data types in the engagement table
for engagement_record in daily_engagement:
engagement_record['lessons_completed'] = int(float(engagement_record['lessons_completed']))
engagement_record['num_courses_visited'] = int(float(engagement_record['num_courses_visited']))
engagement_record['projects_completed'] = int(float(engagement_record['projects_completed']))
engagement_record['total_minutes_visited'] = float(engagement_record['total_minutes_visited'])
engagement_record['utc_date'] = parse_date(engagement_record['utc_date'])
daily_engagement[0]
# +
# Clean up the data types in the submissions table
for submission in project_submissions:
submission['completion_date'] = parse_date(submission['completion_date'])
submission['creation_date'] = parse_date(submission['creation_date'])
project_submissions[0]
# -
# ## Investigating the Data
# - Remoção de elementos repetidos
# +
## Garanta que nao haja elementos repetidos
len(enrollments)
# utiliza a estrutura de dados set() para garantir que não haja alunos repetidos
unique_enrolled_students = set()
for enrollment in enrollments:
# armazena cada aluno da tabela num set()
unique_enrolled_students.add(enrollment['account_key'])
len(unique_enrolled_students)
print(f'Total enrollments = {len(enrollments)}')
print(f'Total enrollments set() = {len(unique_enrolled_students)}')
len(daily_engagement)
unique_engagement_students = set()
for engagement_record in daily_engagement:
unique_engagement_students.add(engagement_record['acct'])
len(unique_engagement_students)
print(f'Total daily_engagement = {len(daily_engagement)}')
print(f'Total daily_engagement set() = {len(unique_engagement_students)}')
len(project_submissions)
unique_project_submitters = set()
for submission in project_submissions:
unique_project_submitters.add(submission['account_key'])
len(unique_project_submitters)
print(f'Total project_submissions = {len(project_submissions)}')
print(f'Total project_submissions set() = {len(unique_project_submitters)}')
# -
# ## Relacionamento
# - PK de todas as tabelas
# +
## Rename the "acct" column in the daily_engagement table to "account_key".
for engagement in daily_engagement:
engagement['account_key'] = engagement['acct']
del[engagement['acct']]
daily_engagement[0]['account_key']
# -
# - LEFT, RIGHT, NATURAL JOIN
# - WHERE ()
#
# Depende do o quê esta se precisando.
#
# Neste caso foi solicitado para saber quais alunos inscritos (enrollments) mas NAO estao enganjados diariamente:
# `where student **not in** unique_engagement_students`
# TABLES: enrollments AND daily_engagement
for enrollment in enrollments:
# get student da tabela enrollments
student = enrollment['account_key']
# verifica se o student NAO esta matriculado
if student not in unique_engagement_students:
print (enrollment)
# ## Rename collumns
# Rename the column names by specifying a list
# ['label, 'sms_message'] to the 'names' argument of read_table().
df = pd.read_table('/home/brunocampos01/projetos/data_science/machine_learning/'
'supervised_learning/data_base/smsspamcollection/SMSSpamCollection',
names=['label', 'sms_message'])
# ## Convert String in number
# +
# Convert the values in the 'label'
dict = {'ham':0, 'spam':1}
# mapped collumn to dict
df ['label'] = df.label.map(dict)
# -
# ### Tokenizen words
# To handle this, we will be using sklearns
# [count vectorizer](http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html#sklearn.feature_extraction.text.CountVectorizer) method which does the following:
#
# * It tokenizes the string(separates the string into individual words) and gives an integer ID to each token.
# * It counts the occurrence of each of those tokens.
#
# ** Please Note: **
#
# * The CountVectorizer method automatically converts all tokenized words to their lower case form so that it does not treat words like 'He' and 'he' differently. It does this using the `lowercase` parameter which is by default set to `True`.
#
# * It also ignores all punctuation so that words followed by a punctuation mark (for example: 'hello!') are not treated differently than the same words not prefixed or suffixed by a punctuation mark (for example: 'hello'). It does this using the `token_pattern` parameter which has a default regular expression which selects tokens of 2 or more alphanumeric characters.
#
# * The third parameter to take note of is the `stop_words` parameter. Stop words refer to the most commonly used words in a language. They include words like 'am', 'an', 'and', 'the' etc. By setting this parameter value to `english`, CountVectorizer will automatically ignore all words(from our input text) that are found in the built in list of english stop words in scikit-learn. This is extremely helpful as stop words can skew our calculations when we are trying to find certain key words that are indicative of spam.
#
# We will dive into the application of each of these into our model in a later step, but for now it is important to be aware of such preprocessing techniques available to us when dealing with textual data.
# ### Data cleaning in words
# - lower()
# - punctuation (translate())
# - split()
#
#
# ### Count frequencies words
# Counter é uma subclasse de dict para contar e mapear objetos hashable
#
# `from collections import Counter`
#
# `for i in preprocessed_documents:
# frequency_counts = Counter(i)
# frequency_list.append(frequency_counts)`
#
#
#
# https://docs.python.org/2//library/collections.html
# ## Vectorize words
#
# Serve para depois converter os dados em uma matriz.
from sklearn.feature_extraction.text import CountVectorizer
count_vector = CountVectorizer()
print(count_vector)
| data_analysis_udacity/.ipynb_checkpoints/load_and_clean_data-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Blind Source Separations
#
#
# ## Introduction
# How can you determine how many people are talking in a room at a party? It's a [challenging problem](https://en.wikipedia.org/wiki/Source_separation) and the basic premise of the problems creeps up in a lot of different fields.
#
# In my own field, EEG is a perfect example of trying to "unmix" signals to find out what I actually measured. Scalp EEG is highly correlated, with a single brain source being "smeared" across multiple channels.
#
# Blind source separation is the problem of trying to split out independent processes that are generating data. Doing this without a priori information about the system/s generating the data is the "blind" part of this.
#
# A common example of this type of problem is trying to identify the number of people speaking in a noisy room with a certain number of microphones. Each microphone picks up each speaker, but to varying degrees. With information about where the microphones are, this problem is not so "blind". Without information about where the microphones are, this problem becomes "blind", but not insurmountable.
# + [markdown] deletable=true editable=true
# ## Generate our data
# We know the properties that our data needs:
#
# * Gaussian with noise
# * Multimodal
#
# Since we'll be dealing with timeseries in other notebooks, we'll focus our conversation around timeseries knowing that the principles are generalizable.
# + deletable=true editable=true
# %reset
import numpy as np
import matplotlib.pyplot as plt
import scipy.signal as sig
# + deletable=true editable=true
#We'll deal with 2D data
#Simple cross example
mean1 = [0,0]
cov1 = [[0,0.7],[-3.5,6]]
cov2 = [[0,-0.7],[3.5,6]]
mean2 = [4,0]
x,y = np.random.multivariate_normal(mean1,cov1,100).T
u,v = np.random.multivariate_normal(mean2,cov2,100).T
plt.plot(x,y,'x')
plt.plot(u,v,'x',color='r')
plt.axis('equal')
plt.show()
# + [markdown] deletable=true editable=true
# We made a dataset where two independent processes are observed. We know it's two independent processes because we *made them from scratch using two separate function calls to multivariate normal*. Of course, since it's all pseudorandom number generation, might have to make sure the seeds are different for each call, but I'm not sure that's how it's supposed to work.
#
# We have a dataset with two independent processes. We want to now study this and *find* these processes from data where we won't know where each datapoint is actually coming from.
#
# This set is actually very easy to see *visually* but let's do the process from the ground up. We start with linear approaches, move to ICA, then to gaussian processes.
# + [markdown] deletable=true editable=true
# ### Principle Component Analysis
# First we'll do a PCA on the aggregate dataset. This will give us two components: a component in the direction of maximal variance, and another one orthogonal to that
# + deletable=true editable=true
data = np.vstack((np.hstack((x,u)),np.hstack((y,v))))
plt.plot(data[0,:],data[1,:],'x')
plt.axis('equal')
plt.show()
# + deletable=true editable=true
from sklearn.decomposition import PCA as sklPCA
def do_PCA(data):
skl_PCA = sklPCA(n_components=2)
skl_Xform = skl_PCA.fit_transform(data.T)
plt.plot(skl_Xform[:,0],skl_Xform[:,1],'o')
plt.axis('equal')
plt.show()
pcs = skl_PCA.components_
plt.figure()
ax=plt.axes()
plt.plot(data[0,:],data[1,:],'x')
ax.arrow(0,0,5*pcs[0,0],5*pcs[1,0],color='r',head_width=0.5)
ax.arrow(0,0,2*pcs[0,1],2*pcs[1,1],color='g',head_width=0.5)
#plt.plot(pcs[0,:])
plt.axis('equal')
plt.show()
return pcs
orig_PCs = do_PCA(data)
# + [markdown] deletable=true editable=true
# So, according to PCA, we've got two components in our data. One going in the red direction, the other going in the green direction. We know this isn't true, we don't have a single linear system in 2D that maps inputs to outputs. So PCA completely misses the mark on this.
#
# A next step, if we didn't know what we knew about the source of our data, would be to look at the eigenvalues for each component. This would give us an idea of how much of the data variance is "explained" by that component. The red would be higher, by definition, but the green would account for a sizable portion.
#
# #### Outliers
# In some cases, we can actually see how sensitive PCA is to even a single outlier. Let's add a single point to the above dataset at (2,-6) and see how the principle components shift.
# + deletable=true editable=true
outlier_pt = np.array([[2,-6]]).T
data_outlier = np.hstack(([data,outlier_pt]))
outlier_PCs = do_PCA(data_outlier)
# -
# With just one outlier added to the mix, we have completely changed our principle components. This is why some scientists are, validly, weary of *"machine learning"* as a whole; certain techniques have to be very specifically applied.
# +
plt.figure()
ax=plt.axes()
ax.arrow(0,0,5*orig_PCs[0,0],5*orig_PCs[1,0],color='r',head_width=0.5)
ax.arrow(0,0,5*orig_PCs[0,1],5*orig_PCs[1,1],color='g',head_width=0.5)
ax.arrow(0,0,2*outlier_PCs[0,0],2*outlier_PCs[1,0],color='r',linestyle='dotted',head_width=0.5)
ax.arrow(0,0,2*outlier_PCs[0,1],2*outlier_PCs[1,1],color='g',linestyle='dotted',head_width=0.5)
#plt.plot(pcs[0,:])
plt.axis('equal')
plt.show()
# + [markdown] deletable=true editable=true
# However, calling PCA "machine learning" is like calling a tin can a boat; technically not wrong, but misses the point of a boat.
#
# Let's dive into some more advanced approaches.
# + [markdown] deletable=true editable=true
# ### Independent Component Analysis
# ICA should give us the two components themselves, though since one component is 2d symmetric, not sure what will happen there...
# +
from sklearn.decomposition import FastICA
def do_ICA(data):
rng = np.random.RandomState(42)
skl_ICA = FastICA(random_state = rng)
skl_Xform = ica.fit(data).transform(data)
plt.plot(skl_Xform[:,0],skl_Xform[:,1],'o')
plt.axis('equal')
plt.show()
pcs = skl_PCA.mixing_
plt.figure()
ax=plt.axes()
plt.plot(data[0,:],data[1,:],'x')
ax.arrow(0,0,5*pcs[0,0],5*pcs[1,0],color='r',head_width=0.5)
ax.arrow(0,0,2*pcs[0,1],2*pcs[1,1],color='g',head_width=0.5)
#plt.plot(pcs[0,:])
plt.axis('equal')
plt.show()
return pcs
_ = do_ICA(data.T)
# + [markdown] deletable=true editable=true
# ### Gaussian Mixture Models
# Gaussian mixture models take in a "number" of gaussians expected in the data, and then does a maximization of likelihood of seeing the data for the underlying model trying to be learned.
#
# GMM should give us the two gaussian! Let's just go for it
# + deletable=true editable=true
from sklearn import mixture
from matplotlib.colors import LogNorm
def GMM(data):
clf = mixture.GaussianMixture(n_components=2,covariance_type='full')
clf.fit(data.T)
xd = np.linspace(-20,20)
yd = np.linspace(-20,20)
Xd,Yd = np.meshgrid(xd,yd)
XX = np.array([Xd.ravel(),Yd.ravel()]).T
Z = -clf.score_samples(XX)
Z = Z.reshape(Xd.T.shape)
CS = plt.contour(Xd,Yd,Z,norm=LogNorm(vmin=1.0,vmax=1000),levels=np.logspace(0,2,20))
plt.scatter(data[0,:],data[1,:],.8)
plt.axis('equal')
plt.axis('tight')
plt.show()
print('Learned Gaussian Means ' + str(clf.means_) + '\n\n')
print('Learned Gaussian Covariances \n' + str(clf.covariances_))
GMM(data)
# + [markdown] deletable=true editable=true
# Cool! We see what we expect. What happens with the outlier dataset?
# + deletable=true editable=true
GMM(data_outlier)
# + [markdown] deletable=true editable=true
# Not much is different. Let's look at the learned Gaussians:
#
#
#
# In this case, it's a bit trivial, since we know, a priori, that two sources were used to generate the data. The GMM centered two gaussians accordingly to maximize the likelihood of the data being generated. What happens if we change the number of a priori components in the Gaussian Mixture?
# + deletable=true editable=true
clf = mixture.GaussianMixture(n_components=4,covariance_type='full')
clf.fit(data.T)
xd = np.linspace(-20,20)
yd = np.linspace(-20,20)
Xd,Yd = np.meshgrid(xd,yd)
XX = np.array([Xd.ravel(),Yd.ravel()]).T
Z = -clf.score_samples(XX)
Z = Z.reshape(Xd.T.shape)
CS = plt.contour(Xd,Yd,Z,norm=LogNorm(vmin=1.0,vmax=1000),levels=np.logspace(0,2,20))
plt.scatter(data[0,:],data[1,:],.8)
plt.axis('equal')
plt.axis('tight')
plt.show()
# + [markdown] deletable=true editable=true
# This is a key consideration of these types of methods; I say, a priori, there are 4 components. The algorithm will *find* four gaussians and mold them to maximize the likelihood of seeing the data. We know this to be *wrong*, but it's a different type of wrong. It's an "overfitting" type of wrong.
| BlindSourceSep.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using custom containers with AI Platform Training
#
# **Learning Objectives:**
# 1. Learn how to create a train and a validation split with Big Query
# 1. Learn how to wrap a machine learning model into a Docker container and train in on CAIP
# 1. Learn how to use the hyperparameter tunning engine on GCP to find the best hyperparameters
# 1. Learn how to deploy a trained machine learning model GCP as a rest API and query it.
#
# In this lab, you develop, package as a docker image, and run on **AI Platform Training** a training application that trains a multi-class classification model that predicts the type of forest cover from cartographic data. The [dataset](../../../datasets/covertype/README.md) used in the lab is based on **Covertype Data Set** from UCI Machine Learning Repository.
#
# The training code uses `scikit-learn` for data pre-processing and modeling. The code has been instrumented using the `hypertune` package so it can be used with **AI Platform** hyperparameter tuning.
#
# +
import json
import os
import numpy as np
import pandas as pd
import pickle
import uuid
import time
import tempfile
from googleapiclient import discovery
from googleapiclient import errors
from google.cloud import bigquery
from jinja2 import Template
from kfp.components import func_to_container_op
from typing import NamedTuple
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
from sklearn.linear_model import SGDClassifier
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler, OneHotEncoder
from sklearn.compose import ColumnTransformer
# -
# ## Configure environment settings
# Set location paths, connections strings, and other environment settings. Make sure to update `REGION`, and `ARTIFACT_STORE` with the settings reflecting your lab environment.
#
# - `REGION` - the compute region for AI Platform Training and Prediction
# - `ARTIFACT_STORE` - the GCS bucket created during installation of AI Platform Pipelines. The bucket name starts with the `hostedkfp-default-` prefix.
# !gsutil ls
# +
REGION = 'us-central1'
ARTIFACT_STORE = 'gs://qwiklabs-gcp-04-8722038efd75'
#ARTIFACT_STORE = 'gs://hostedkfp-default-l2iv13wnek'
PROJECT_ID = !(gcloud config get-value core/project)
PROJECT_ID = PROJECT_ID[0]
DATA_ROOT='{}/data'.format(ARTIFACT_STORE)
JOB_DIR_ROOT='{}/jobs'.format(ARTIFACT_STORE)
TRAINING_FILE_PATH='{}/{}/{}'.format(DATA_ROOT, 'training', 'dataset.csv')
VALIDATION_FILE_PATH='{}/{}/{}'.format(DATA_ROOT, 'validation', 'dataset.csv')
# -
# ## Explore the Covertype dataset
# %%bigquery
SELECT *
FROM `covertype_dataset.covertype`
# ## Create training and validation splits
#
# Use BigQuery to sample training and validation splits and save them to GCS storage
# ### Create a training split
# !bq query \
# -n 0 \
# --destination_table covertype_dataset.training \
# --replace \
# --use_legacy_sql=false \
# 'SELECT * \
# FROM `covertype_dataset.covertype` AS cover \
# WHERE \
# MOD(ABS(FARM_FINGERPRINT(TO_JSON_STRING(cover))), 10) IN (1, 2, 3, 4)'
# !bq extract \
# --destination_format CSV \
# covertype_dataset.training \
# $TRAINING_FILE_PATH
# ### Create a validation split
# ### Exercise
#
# In the first cell below, create
# a validation split that takes 10% of the data using the `bq` command and
# export this split into the BigQuery table `covertype_dataset.validation`.
#
# In the second cell, use the `bq` command to export that BigQuery validation table to GCS at `$VALIDATION_FILE_PATH`.
# !bq query \
# -n 0 \
# --destination_table covertype_dataset.validation \
# --replace \
# --use_legacy_sql=false \
# 'SELECT * \
# FROM `covertype_dataset.covertype` AS cover \
# WHERE \
# MOD(ABS(FARM_FINGERPRINT(TO_JSON_STRING(cover))), 10) IN (5)'
# !bq extract \
# --destination_format CSV \
# covertype_dataset.validation \
# $VALIDATION_FILE_PATH
df_train = pd.read_csv(TRAINING_FILE_PATH)
df_validation = pd.read_csv(VALIDATION_FILE_PATH)
print(df_train.shape)
print(df_validation.shape)
# ## Develop a training application
# ### Configure the `sklearn` training pipeline.
#
# The training pipeline preprocesses data by standardizing all numeric features using `sklearn.preprocessing.StandardScaler` and encoding all categorical features using `sklearn.preprocessing.OneHotEncoder`. It uses stochastic gradient descent linear classifier (`SGDClassifier`) for modeling.
# +
numeric_feature_indexes = slice(0, 10)
categorical_feature_indexes = slice(10, 12)
preprocessor = ColumnTransformer(
transformers=[
('num', StandardScaler(), numeric_feature_indexes),
('cat', OneHotEncoder(), categorical_feature_indexes)
])
pipeline = Pipeline([
('preprocessor', preprocessor),
('classifier', SGDClassifier(loss='log', tol=1e-3))
])
# -
# ### Convert all numeric features to `float64`
#
# To avoid warning messages from `StandardScaler` all numeric features are converted to `float64`.
# +
num_features_type_map = {feature: 'float64' for feature in df_train.columns[numeric_feature_indexes]}
df_train = df_train.astype(num_features_type_map)
df_validation = df_validation.astype(num_features_type_map)
# -
# ### Run the pipeline locally.
# +
X_train = df_train.drop('Cover_Type', axis=1)
y_train = df_train['Cover_Type']
X_validation = df_validation.drop('Cover_Type', axis=1)
y_validation = df_validation['Cover_Type']
pipeline.set_params(classifier__alpha=0.001, classifier__max_iter=200)
pipeline.fit(X_train, y_train)
# -
# ### Calculate the trained model's accuracy.
accuracy = pipeline.score(X_validation, y_validation)
print(accuracy)
# ### Prepare the hyperparameter tuning application.
# Since the training run on this dataset is computationally expensive you can benefit from running a distributed hyperparameter tuning job on AI Platform Training.
TRAINING_APP_FOLDER = 'training_app'
os.makedirs(TRAINING_APP_FOLDER, exist_ok=True)
# ### Write the tuning script.
#
# Notice the use of the `hypertune` package to report the `accuracy` optimization metric to AI Platform hyperparameter tuning service.
# ### Exercise
#
# Complete the code below to capture the metric that the hyper parameter tunning engine will use to optimize
# the hyper parameter.
# +
# %%writefile {TRAINING_APP_FOLDER}/train.py
# Copyright 2019 Google Inc. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import os
import subprocess
import sys
import fire
import pickle
import numpy as np
import pandas as pd
import hypertune
from sklearn.compose import ColumnTransformer
from sklearn.linear_model import SGDClassifier
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler, OneHotEncoder
def train_evaluate(job_dir, training_dataset_path,
validation_dataset_path, alpha, max_iter, hptune):
df_train = pd.read_csv(training_dataset_path)
df_validation = pd.read_csv(validation_dataset_path)
if not hptune:
df_train = pd.concat([df_train, df_validation])
numeric_feature_indexes = slice(0, 10)
categorical_feature_indexes = slice(10, 12)
preprocessor = ColumnTransformer(
transformers=[
('num', StandardScaler(), numeric_feature_indexes),
('cat', OneHotEncoder(), categorical_feature_indexes)
])
pipeline = Pipeline([
('preprocessor', preprocessor),
('classifier', SGDClassifier(loss='log',tol=1e-3))
])
num_features_type_map = {feature: 'float64' for feature
in df_train.columns[numeric_feature_indexes]}
df_train = df_train.astype(num_features_type_map)
df_validation = df_validation.astype(num_features_type_map)
print('Starting training: alpha={}, max_iter={}'.format(alpha, max_iter))
X_train = df_train.drop('Cover_Type', axis=1)
y_train = df_train['Cover_Type']
pipeline.set_params(classifier__alpha=alpha, classifier__max_iter=max_iter)
pipeline.fit(X_train, y_train)
if hptune:
# TODO: Score the model with the validation data and capture the result
# with the hypertune library
X_validation = df_validation.drop('Cover_Type', axis=1)
y_validation = df_validation['Cover_Type']
accuracy = pipeline.score(X_validation, y_validation)
print('Model accuracy: {}'.format(accuracy))
hpt = hypertune.HyperTune()
hpt.report_hyperparameter_tuning_metric(
hyperparameter_metric_tag='accuracy',
metric_value=accuracy)
# Save the model
if not hptune:
model_filename = 'model.pkl'
with open(model_filename, 'wb') as model_file:
pickle.dump(pipeline, model_file)
gcs_model_path = "{}/{}".format(job_dir, model_filename)
subprocess.check_call(['gsutil', 'cp', model_filename, gcs_model_path],
stderr=sys.stdout)
print("Saved model in: {}".format(gcs_model_path))
if __name__ == "__main__":
fire.Fire(train_evaluate)
# -
# ### Package the script into a docker image.
#
# Notice that we are installing specific versions of `scikit-learn` and `pandas` in the training image. This is done to make sure that the training runtime is aligned with the serving runtime. Later in the notebook you will deploy the model to AI Platform Prediction, using the 1.15 version of AI Platform Prediction runtime.
#
# Make sure to update the URI for the base image so that it points to your project's **Container Registry**.
# ### Exercise
#
# Complete the Dockerfile below so that it copies the 'train.py' file into the container
# at `/app` and runs it when the container is started.
# +
# %%writefile {TRAINING_APP_FOLDER}/Dockerfile
FROM gcr.io/deeplearning-platform-release/base-cpu
RUN pip install -U fire cloudml-hypertune scikit-learn==0.20.4 pandas==0.24.2
WORKDIR /app
COPY train.py .
ENTRYPOINT ["python", "train.py"]
# -
# ### Build the docker image.
#
# You use **Cloud Build** to build the image and push it your project's **Container Registry**. As you use the remote cloud service to build the image, you don't need a local installation of Docker.
IMAGE_NAME='trainer_image'
IMAGE_TAG='latest'
IMAGE_URI='gcr.io/{}/{}:{}'.format(PROJECT_ID, IMAGE_NAME, IMAGE_TAG)
# !gcloud builds submit --tag $IMAGE_URI $TRAINING_APP_FOLDER
# ## Submit an AI Platform hyperparameter tuning job
# ### Create the hyperparameter configuration file.
# Recall that the training code uses `SGDClassifier`. The training application has been designed to accept two hyperparameters that control `SGDClassifier`:
# - Max iterations
# - Alpha
#
# The below file configures AI Platform hypertuning to run up to 6 trials on up to three nodes and to choose from two discrete values of `max_iter` and the linear range betwee 0.00001 and 0.001 for `alpha`.
# ### Exercise
#
# Complete the `hptuning_config.yaml` file below so that the hyperparameter
# tunning engine try for parameter values
# * `max_iter` the two values 200 and 300
# * `alpha` a linear range of values between 0.00001 and 0.001
# +
# %%writefile {TRAINING_APP_FOLDER}/hptuning_config.yaml
# Copyright 2019 Google Inc. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
trainingInput:
hyperparameters:
goal: MAXIMIZE
maxTrials: 4
maxParallelTrials: 4
hyperparameterMetricTag: accuracy
enableTrialEarlyStopping: TRUE
params:
- parameterName: max_iter
type: DISCRETE
discreteValues: [
200,
500
]
- parameterName: alpha
type: DOUBLE
minValue: 0.00001
maxValue: 0.001
scaleType: UNIT_LINEAR_SCALE
# -
# ### Start the hyperparameter tuning job.
#
#
# ### Exercise
# Use the `gcloud` command to start the hyperparameter tuning job.
# +
JOB_NAME = "JOB_{}".format(time.strftime("%Y%m%d_%H%M%S"))
JOB_DIR = "{}/{}".format(JOB_DIR_ROOT, JOB_NAME)
SCALE_TIER = "BASIC"
# !gcloud ai-platform jobs submit training $JOB_NAME \
# --region=$REGION \
# --job-dir=$JOB_DIR \
# --master-image-uri=$IMAGE_URI \
# --scale-tier=$SCALE_TIER \
# --config $TRAINING_APP_FOLDER/hptuning_config.yaml \
# -- \
# --training_dataset_path=$TRAINING_FILE_PATH \
# --validation_dataset_path=$VALIDATION_FILE_PATH \
# --hptune
# -
# ### Monitor the job.
#
# You can monitor the job using GCP console or from within the notebook using `gcloud` commands.
# !gcloud ai-platform jobs describe $JOB_NAME
# !gcloud ai-platform jobs stream-logs $JOB_NAME
# ### Retrieve HP-tuning results.
# After the job completes you can review the results using GCP Console or programatically by calling the AI Platform Training REST end-point.
# +
ml = discovery.build('ml', 'v1')
job_id = 'projects/{}/jobs/{}'.format(PROJECT_ID, JOB_NAME)
request = ml.projects().jobs().get(name=job_id)
try:
response = request.execute()
except errors.HttpError as err:
print(err)
except:
print("Unexpected error")
response
# -
# The returned run results are sorted by a value of the optimization metric. The best run is the first item on the returned list.
response['trainingOutput']['trials'][0]
# ## Retrain the model with the best hyperparameters
#
# You can now retrain the model using the best hyperparameters and using combined training and validation splits as a training dataset.
# ### Configure and run the training job
alpha = response['trainingOutput']['trials'][0]['hyperparameters']['alpha']
max_iter = response['trainingOutput']['trials'][0]['hyperparameters']['max_iter']
# +
JOB_NAME = "JOB_{}".format(time.strftime("%Y%m%d_%H%M%S"))
JOB_DIR = "{}/{}".format(JOB_DIR_ROOT, JOB_NAME)
SCALE_TIER = "BASIC"
# !gcloud ai-platform jobs submit training $JOB_NAME \
# --region=$REGION \
# --job-dir=$JOB_DIR \
# --master-image-uri=$IMAGE_URI \
# --scale-tier=$SCALE_TIER \
# -- \
# --training_dataset_path=$TRAINING_FILE_PATH \
# --validation_dataset_path=$VALIDATION_FILE_PATH \
# --alpha=$alpha \
# --max_iter=$max_iter \
# --nohptune
# -
# !gcloud ai-platform jobs stream-logs $JOB_NAME
# ### Examine the training output
#
# The training script saved the trained model as the 'model.pkl' in the `JOB_DIR` folder on GCS.
# !gsutil ls $JOB_DIR
# ## Deploy the model to AI Platform Prediction
# ### Create a model resource
# ### Exercise
#
# Complete the `gcloud` command below to create a model with
# `model_name` in `$REGION` tagged with `labels`:
# +
model_name = 'forest_cover_classifier'
labels = "task=classifier,domain=forestry"
# !gcloud ai-platform models create $model_name \
# --regions=$REGION \
# --labels=$labels
# -
# ### Create a model version
# ### Exercise
# Complete the `gcloud` command below to create a version of the model:
# +
model_version = 'v01'
# !gcloud ai-platform versions create {model_version} \
# --model={model_name} \
# --origin=$JOB_DIR \
# --runtime-version=1.15 \
# --framework=scikit-learn \
# --python-version=3.7
# -
# ### Serve predictions
# #### Prepare the input file with JSON formated instances.
# +
input_file = 'serving_instances.json'
with open(input_file, 'w') as f:
for index, row in X_validation.head().iterrows():
f.write(json.dumps(list(row.values)))
f.write('\n')
# -
# !cat $input_file
# #### Invoke the model
# ### Exercise
#
# Using the `gcloud` command send the data in `$input_file` to
# your model deployed as a REST API:
# !gcloud ai-platform predict \
# --model $model_name \
# --version $model_version \
# --json-instances $input_file
# <font size=-1>Licensed under the Apache License, Version 2.0 (the \"License\");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at [https://www.apache.org/licenses/LICENSE-2.0](https://www.apache.org/licenses/LICENSE-2.0)
#
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an \"AS IS\" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.</font>
| notebooks/mlops/kubeflow_pipelines/walkthrough/labs/lab-01.ipynb |