code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Importacion general de librerias y de visualizacion (matplotlib y seaborn)
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
plt.style.use('default') # haciendo los graficos un poco mas bonitos en matplotlib
plt.rcParams['figure.figsize'] = (20, 10)
sns.set(style="whitegrid") # seteando tipo de grid en seaborn
# -
# ---------------------------------------------------------------------------------------
# ---------------------------------------------------------------------------------------
# # 2C 2017
# # Enunciado
#
# Tenemos un dataframe con la información de distintas playlists armadas por usuarios con el formato:
#
# ```
# (playlist, song_id, description)
# ```
#
# A su vez, contamos con un dataframe de canciones que contiene:
# ```
# (song_id, singer, year, lenght, genres)
# ```
#
# Se pide generar un programa en Pandas que indique para cada playlist cual es el cantante predominante (con mas canciones incluidas dentro de esa lista)
#
# +
## Reviews de usuarios de libros
m_playlist = [
[1, 1, "texto"],
[2, 1, "texto"],
[1, 2, "texto"],
[1, 3, "texto"],
[2, 3, "texto"],
[3, 1, "texto"],
[1, 4, "texto"],
[2, 5, "texto"],
[1, 4, "texto"],
[4, 3, "texto"],
[4, 1, "texto"],
[4, 2, "texto"],
[1, 3, "texto"],
[2, 9, "texto"],
[1, 4, "texto"],
[1, 7, "texto"],
[2, 1, "texto"],
[3, 9, "texto"],
[1, 3, "texto"],
[2, 5, "texto"],
[1, 3, "texto"],
[4, 2, "texto"],
[4, 8, "texto"],
[4, 7, "texto"],
]
songs_m = [
[1, "Singer 1", "1997", 3.14, "rock"],
[2, "Singer 1", "1997", 3.25, "rock"],
[3, "Singer 2", "1997", 2.50, "rock"],
[4, "Singer 2", "1997", 1.00, "rock"],
[5, "Singer 3", "1997", 4.00, "rock"],
[6, "Singer 4", "1997", 3.25, "rock"],
[7, "Singer 2", "1997", 2.50, "rock"],
[8, "Singer 2", "1997", 1.00, "rock"],
[9, "Singer 1", "1997", 3.14, "rock"],
[10, "Singer 1", "1997", 3.25, "rock"],
]
# -
playlists = pd.DataFrame(m_playlist, columns=list(("playlist", "song_id", "description")))
songs= pd.DataFrame(songs_m, columns=list(("song_id", "singer", "year", "lenght", "genres")))
playlists
songs
merged = pd.merge(playlists, songs, on="song_id", how="inner")
merged
groupped = merged.groupby(["playlist", "singer"]).agg({"song_id": "count"})
sorted = groupped.sort_values("song_id", ascending=False)
sorted.reset_index().drop_duplicates("playlist")
# ---------------------------------------------------------------------------------------
# ---------------------------------------------------------------------------------------
# # 1C 2017
# # Enunciado
#
# Un sitio de Ebooks tiene información sobre los reviews que los usuarios hacen de sus libros en un DataFrame con formato:
# ```
# (user_id, book_id, rating, timestamp)
# ```
#
# Por otro lado tenemos información en otro DataFrame que bajamos de GoodReads:
# ```
# (book_id, book_name, avg_rating)
# ```
#
# Podemos suponer que los Ids de los libros son compatibles. Se pide usar Python Pandas para:
#
# **a)** Obtener un DataFrame que indique el TOP5 de Ebooks en el sitio de Ebooks. (Para este punto se puede ignorar el segundo DataFrame)
#
# **b)** Obtener un DataFrame que indique qué libros tienen una diferencia de rating promedio mayor al 20% entre el sitio de Ebooks y GoodReads.
# +
## Reviews de usuarios de libros
m = [
[1, 1, 5, 1526245886],
[2, 1, 4, 1526245886],
[1, 2, 1, 1526245886],
[2, 2, 2, 1526245886],
[3, 1, 4, 1526245886],
[3, 2, 5, 1526245886],
[4, 1, 3, 1526245886],
[4, 2, 5, 1526245886],
[1, 3, 5, 1526245886],
[2, 6, 4, 1526245886],
[1, 3, 1, 1526245886],
[2, 3, 2, 1526245886],
[3, 6, 4, 1526245886],
[3, 6, 1, 1526245886],
[4, 6, 3, 1526245886],
[4, 3, 5, 1526245886],
]
m2 = [
[1, "Book 1", 4.00],
[2, "Book 2", 3.25],
[3, "Book 3", 2.50],
[4, "Book 4", 1.00],
[5, "Book 5", 4.00],
[6, "Book 6", 3.25],
[7, "Book 7", 2.50],
[8, "Book 8", 1.00],
]
# -
ebooks = pd.DataFrame(m, columns=list(("user_id", "book_id", "rating", "timestamp")))
goodReads = pd.DataFrame(m2, columns=list(("book_id", "book_name", "rating")))
ebooks
goodReads
groupped = ebooks.groupby(["book_id"]).agg({"rating": ["mean", "count"]})
groupped
groupped = groupped.reset_index()
groupped.columns = ["book_id", "mean", "count"]
groupped.sort_values(by="mean", ascending=False)
groupped.sort_values(by="mean", ascending=False).head(5)
merged = pd.merge(groupped, goodReads, on="book_id", how="inner")
merged
merged[["mean", "rating"]]
merged.apply(lambda x: (abs(x["rating"] - x["mean"])) / 5, axis="columns")
merged["diff"] = merged.apply(lambda x: (abs(x["rating"] - x["mean"])) / 5, axis="columns")
merged.loc[merged["diff"] > 0.1].sort_values("book_id", ascending=False)
merged.apply(lambda x: x[2])
# ---------------------------------------------------------------------------------------
# ---------------------------------------------------------------------------------------
# # 1C 2016
# # Enunciado
#
# Dados los archivos:
# - Notas (Padron, Codigo de Materia, Codigo de Curso, Nota)
# - Cursos (Codigo de Materia, Nombre de Materia, Codigo de Curso, - Profesor a Cargo)
#
# Hacer un programa en Pandas que liste para cada curso de cada materia, el promedio de notas de los alumnos que aprobaron la misma.
#
# El listado debe contener Codigo de Materia, Codigo de Curso, Profesor a cargo y Promedio de Notas, y debe estar ordenado por materia.
#
# Tener en cuenta solo los cursos que tengan al menos 100 alumnos aprobados.
#
#
# +
## Reviews de usuarios de libros
notas_m = [
[9000, 1, 100, 4],
[9001, 1, 100, 2],
[9002, 1, 101, 9],
[9004, 2, 200, 4],
[9005, 3, 300, 6],
[9006, 4, 401, 7],
[9007, 2, 203, 3],
[9008, 4, 401, 9],
[9009, 1, 101, 8],
[9010, 1, 100, 2],
[9011, 2, 203, 4],
[9012, 2, 202, 6],
[9013, 2, 205, 8],
[9014, 1, 100, 4],
[9015, 3, 301, 2],
[9016, 3, 100, 4],
[9017, 1, 100, 4],
[9018, 4, 400, 3],
[9019, 2, 204, 9],
[9020, 3, 303, 6],
]
cursos_m = [
[1, "Álgebra", 100, "Profesor 1"],
[1, "Álgebra", 101, "Profesor 2"],
[2, "Análisis Matemático", 200, "Profesor 3"],
[2, "Análisis Matemático", 201, "Profesor 4"],
[2, "Análisis Matemático", 202, "Profesor 5"],
[2, "Análisis Matemático", 203, "Profesor 7"],
[3, "Algoritmos I", 300, "Profesor 10"],
[3, "Algoritmos I", 301, "Profesor 10"],
[3, "Algoritmos I", 302, "Profesor 11"],
[4, "Inteligencia Artificial", 400, "Profesor 9"],
[4, "Inteligencia Artificial", 401, "Profesor 99"],
[2, "Análisis Matemático", 204, "Profesor 44"],
[2, "Análisis Matemático", 205, "Profesor 55"],
[3, "Algoritmos I", 303, "Profesor 4"],
]
# -
notas = pd.DataFrame(notas_m, columns=list(("padron", "codigo_materia", "codigo_curso", "nota")))
cursos = pd.DataFrame(cursos_m, columns=list(("codigo_materia", "nombre_materia", "codigo_curso","profesor")))
notas.shape
# Filtrar los alumnos que aprobaron
approved = notas.loc[notas["nota"] >= 4]
approved.shape
merged = pd.merge(cursos, approved, on=["codigo_materia", "codigo_curso"], how="inner")
merged
groupped = merged.groupby(["codigo_materia", "codigo_curso", "profesor"]).agg({"nota": ["count", "mean"]})
groupped
groupped = groupped.reset_index()
groupped
groupped.columns = ["codigo_materia", "codigo_curso", "profesor", "count", "mean"]
groupped
n = 1
filtered = groupped.loc[groupped["count"] >= n]
filtered
sorted = groupped.sort_values("codigo_materia")
sorted
| pandas/Ejercicios Parcial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="oIA8dahyEPo0"
import time
import toml
import numpy as np
import matplotlib.pyplot as plt
from ref_trajectory import plot_smooth_route
from ref_trajectory import generate_trajectory as traj
from astar import gen_astar_path, plot_astar
from route_directions import make_route,euclidean
from collision_dwa import simulate_unicycle,track,plot_final_path,plot_wvk
from environment_setup import grid_from_map,place_obstacles,plot_environment,densify_grid
# %matplotlib inline
config_params = toml.load("config.toml")['params']
locals().update(config_params)
# + colab={"base_uri": "https://localhost:8080/", "height": 670} id="4r_4TTsGWNsd" outputId="1d71108f-1a70-4b2a-fc9d-349eb56d35b4"
data = np.loadtxt("arena_2.map", skiprows=4, dtype='str')
data_header = np.loadtxt("arena_2.map", skiprows=1, max_rows = 2, dtype = 'str')
data_size = [int(data_header[0][1]), int(data_header[1][1])]
grid = grid_from_map(data,data_size)
start = (128,150)
goal= (174,123)
astar_path,extent_limits = gen_astar_path(grid,start,goal)
plot_astar(grid,start,goal,astar_path,extent_limits)
# + colab={"base_uri": "https://localhost:8080/", "height": 670} id="FHOOEFwLrDah" outputId="3c5bca3a-dcb2-49a8-81c6-895484b78b61"
grid_dense = densify_grid(grid)
plot_astar(grid_dense,start,goal,astar_path,extent_limits)
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="dAozuJivXxH2" outputId="1482b12e-48e4-457c-d82c-028f54681189"
pose = (*astar_path[0],0)
logs = []
path_index = 0
v, w = 0.0, 0.0
while path_index < len(astar_path)-1:
t0 = time.time()
local_ref_path = astar_path[path_index:path_index+pred_horizon]
# update path_index using current pose and local_ref_path
if euclidean(pose[:2],local_ref_path[-1][:2]) < goal_threshold*4*pred_horizon:
path_index = path_index + 1
local_ref_path = astar_path[path_index:path_index+pred_horizon]
# get next command
v, w = track(grid_dense,local_ref_path,pose,v,w)
#simulate vehicle for 1 step
# remember the function now returns a trajectory, not a single pose
pose = simulate_unicycle(pose,v,w)[-1]
#update logs
logs.append([*pose, v, w, local_ref_path[-1]])
t1 = time.time() #simplest way to time-profile your code
print(f"idx:{path_index}, v:{v:0.3f}, w:{w:0.3f}, current pose: {pose}, tracking point:{local_ref_path[-1][:2]}, time:{(t1-t0) * 1000:0.1f}ms")
# + id="HOryvzpK3kXJ" colab={"base_uri": "https://localhost:8080/", "height": 600} outputId="084b5f7d-9d15-4a22-f16f-090c383b0e56"
plot_final_path(grid_dense,astar_path,logs,extent_limits)
# + colab={"base_uri": "https://localhost:8080/", "height": 679} id="W_k__FW04WS3" outputId="b39951a3-6fae-4fa6-b406-c1dd55309619"
plot_wvk(logs)
# + id="dpz1AeRaNplC"
| project/submissions/akshathaj/tests/ipynb_files/arena_2_tracking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="9MiGmfrbNo9P"
# In this notebook we demonstrate Temporal IE using [duckling](https://github.com/FraBle/python-duckling), which is a python wrapper for wit.ai's [Duckling](https://github.com/facebookarchive/duckling_old) Clojure library.
# + colab={"base_uri": "https://localhost:8080/", "height": 90} colab_type="code" id="kt0BWbCFMLP0" outputId="a0669798-63d5-4871-e539-db70a3cac257"
#installing the package
# !pip install JPype1==0.7.4 #This is required as duckling is not compatible with recent versions of JPype.
# !pip install duckling==1.8.0
# + colab={} colab_type="code" id="taFQDgC7NUOt"
from duckling import DucklingWrapper
from pprint import pprint
# + colab={} colab_type="code" id="xDDNmWFQbFQF"
d = DucklingWrapper()
print(d.parse_time(u'Let\'s meet at 11:45am'))
# + [markdown] colab_type="text" id="M8ZX3J0vz5ro"
# Extracting time from text
#
# + colab={"base_uri": "https://localhost:8080/", "height": 454} colab_type="code" id="anWMHSQAbHi-" outputId="2c9cec94-3999-4a00-ff97-519196c167e4"
pprint(d.parse_time(u'Let\'s meet at 11:45am'))
pprint(d.parse_time(u'You owe me twenty bucks, please call me today'))
# + [markdown] colab_type="text" id="mTiNsnoG0BaK"
# Extracting temperature from text
# + colab={"base_uri": "https://localhost:8080/", "height": 381} colab_type="code" id="_ab9_gzGbab2" outputId="c73db449-9527-46b3-f80b-f43887f54bdf"
pprint(d.parse_temperature(u'Let\'s change the temperatur from thirty two celsius to 65 degrees'))
pprint(d.parse_temperature(u"It's getting hotter day by day, yesterday it was thirty-five degrees celcius today its 37 degrees "))
# + [markdown] colab_type="text" id="MIhGr8sD0EaK"
# Extracting timezone from text
# + colab={"base_uri": "https://localhost:8080/", "height": 199} colab_type="code" id="n80DX6jLbncT" outputId="bc33984a-e691-4527-aabd-9728ce934c82"
pprint(d.parse_timezone(u"Let's meet at 10pm IST"))
pprint(d.parse_timezone(u"Let's meet at 22:00 EST"))
# + [markdown] colab_type="text" id="namYN2uV03r-"
# Extracting number from text
# + colab={"base_uri": "https://localhost:8080/", "height": 108} colab_type="code" id="TPUGDEUocLou" outputId="02803174-7854-4036-ee90-3fbe68ec334f"
d.parse_number(u"Hey i am a 20 year old student from Alaska")
# + [markdown] colab_type="text" id="O-5RYo3j07Rc"
# Extracting ordinals from text
# + colab={"base_uri": "https://localhost:8080/", "height": 199} colab_type="code" id="N-lovNRKcUPA" outputId="41ebd328-9a7e-4aeb-e613-94d52bfd4d72"
d.parse_ordinal(u"I came 2nd and u came 1st in a race")
# + [markdown] colab_type="text" id="dyNu2zVG0_lN"
# Extracting currency and value from text
# + colab={"base_uri": "https://localhost:8080/", "height": 108} colab_type="code" id="RvEkCiBxcta4" outputId="79adfbbf-9690-4bf5-c899-25915f1b0f57"
d.parse_money(u"This meal costs 3$")
# + [markdown] colab_type="text" id="TtzDTHoy1GG2"
# Extracting email ids from text
# + colab={"base_uri": "https://localhost:8080/", "height": 108} colab_type="code" id="HJG0t9Iyc1rI" outputId="61465e7f-a4c8-4890-f70a-a8cc1c149763"
d.parse_email(u"my email is <EMAIL>")
# + [markdown] colab_type="text" id="ixVmrKBw1NPx"
# Extracting the durations in a text
# + colab={"base_uri": "https://localhost:8080/", "height": 454} colab_type="code" id="DxmmN78bdHqK" outputId="0f226807-215a-4a55-ad7e-2cd680bb4184"
d.parse_duration(u"I have been working on this project for 4 hrs every month for almost 2 years.")
# + [markdown] colab_type="text" id="ktDtLHCY1y_f"
# Extracting urls from text
# + colab={"base_uri": "https://localhost:8080/", "height": 199} colab_type="code" id="HEBKT4BT11sb" outputId="88e74b0d-5c58-4be8-972f-205deaede893"
d.parse_url(u"The official website for the book Practical NLP is http://www.practicalnlp.ai/")
# + [markdown] colab_type="text" id="k0Ox-vvV2K7I"
# Extracting phone numbers from text
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="gSRr-ae52Ryd" outputId="fa229777-ad1f-4f0f-93f4-86e299221a00"
d.parse_phone_number(u"Enter text here")#didnt demo this due to privacy reasons
# + [markdown] colab_type="text" id="xCOEyk_u1Rq-"
# Generally, a good idea would be to make a pipeline of all of these functions or which ever you require according to your use case.
# + colab={} colab_type="code" id="K_AV5damdQMK"
# -
| Ch5/08_Duckling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# This notebook preprocess the data extracted from the chess database.
#
# To run this notebook with all the 170 million of positions from the chess database is required at least 8GB of RAM (if you use a local machine, for some reason, I can't run it on google colab).
#
# I used a laptop with a SSD NVMe, Intel i7-9750h and 24GB RAM DDR4@2666Mhz
#
#
total_ram = 170e6*64/1024/1024/1024
print("If all data were loaded, it would take at least {:.1f} GB of RAM".format(total_ram))
# +
# #!pip install chesslab --upgrade
# -
from chesslab_.preprocessing import preprocess
download=False
#https://drive.google.com/file/d/1XwH0reHwaOA0Tpt0ihJkP_XW99EUhlp9/view?usp=sharing
if download:
from chesslab.utils import download_7z
path='./'
file_id = '1XwH0reHwaOA0Tpt0ihJkP_XW99EUhlp9'
download_7z(file_id,path)
else:
path='D:/database/ccrl/'
# +
block_size=1000000
blocks=170
path_files= path
start_name= 'chess'
min_elo= 2500
data_name= 'ccrl_states_elo2'
labels_name= 'ccrl_results_elo2'
elo_filter= 1 #1 = mean, 2 = min
nb_game_filter= 10 #0 no aplica el filtro
delete_duplicate=True
delete_draws= True
delete_both_winners = True
delete_eaten=True
undersampling=False
preprocess(
block_size= block_size,
blocks= blocks,
path= path_files,
start_name= start_name,
min_elo= min_elo,
data_name= data_name,
labels_name= labels_name,
elo_filter= elo_filter,
nb_game_filter= nb_game_filter,
delete_eaten=delete_eaten,
delete_duplicate=delete_duplicate,
delete_draws= delete_draws,
delete_both_winners = delete_both_winners,
undersampling=undersampling)
# -
| examples/preprocess/Preprocess.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import numpy as np
import pickle
import seaborn as sns
from scipy.stats import norm, skewnorm
from pwass.spline import SplineBasis, MonotoneQuadraticSplineBasis
from pwass.distributions import Distribution
from pwass.dimsensionality_reduction.geodesic_pca import GeodesicPCA
from pwass.dimsensionality_reduction.nested_pca import NestedPCA
from pwass.dimsensionality_reduction.projected_pca import ProjectedPCA
from pwass.dimsensionality_reduction.simplicial_pca import SimplicialPCA
np.random.seed(20200712)
# +
nbasis = 20
zero_one_grid = np.linspace(0, 1, 100)
wbasis = MonotoneQuadraticSplineBasis(nbasis, zero_one_grid)
# -
def simulate_data(ndata):
base_means = np.array([-3, 3])
out = []
for i in range(ndata):
xgrid = np.linspace(-10, 10, 1000)
m = np.random.choice(base_means) + np.random.normal(scale=0.2)
std = np.random.uniform(0.5, 2.0)
dens = norm.pdf(xgrid, m, std)
curr = Distribution()
curr.init_from_pdf(xgrid, dens)
out.append(curr)
return out
# +
data = simulate_data(100)
xgrid = data[0].pdf_grid
sbasis = SplineBasis(deg=3, xgrid=xgrid, nbasis=nbasis)
for d in data:
plt.plot(d.pdf_grid, d.pdf_eval)
# d.compute_inv_cdf(wbasis)
# plt.savefig("scenario1_data.pdf")
# -
spca = SimplicialPCA(nbasis)
spca.fit(data, 5)
# +
f = data[0]
spca.k = 3
fig, axes = plt.subplots(nrows=1, ncols=2)
reduced = spca.transform([f])
rec = spca.pt_from_proj(reduced) + spca.bary
rec_pdf = spca.get_pdf(rec)
axes[0].plot(f.pdf_grid, f.pdf_eval)
axes[0].plot(f.pdf_grid, rec_pdf[0, :])
spca.k = 10
reduced = spca.transform([f])
rec = spca.pt_from_proj(reduced) + spca.bary
rec_pdf = spca.get_pdf(rec)
axes[1].plot(spca.pdf_grid, f.pdf_eval)
axes[1].plot(spca.pdf_grid, rec_pdf[0, :])
# +
from scipy.interpolate import UnivariateSpline
def invcdf_to_pdf(zero_one_grid, invcdf_eval, s=0.1):
kept = np.unique(invcdf_eval, return_index=True)[1]
new_grid = np.linspace(np.min(invcdf_eval), np.max(invcdf_eval), 100)
cdf = UnivariateSpline(x=invcdf_eval[kept], y=zero_one_grid[kept], s=s)
der = cdf.derivative()(new_grid)
return new_grid, der
def plot_wpc(pca, ind, pos_lambdas, neg_lambdas, pos_palette, neg_palette, ax, smooth_val):
for j, lam in enumerate(pos_lambdas):
proj = pca.bary + pca.project(lam * pca.eig_vecs[:, ind])
grid, pdf = invcdf_to_pdf(
zero_one_grid,
wbasis.eval_spline(proj), smooth_val)
ax.plot(grid, pdf, color=pos_palette[j])
for j, lam in enumerate(neg_lambdas):
proj = pca.bary + pca.project(lam * pca.eig_vecs[:, ind])
grid, pdf = invcdf_to_pdf(
zero_one_grid,
wbasis.eval_spline(proj), smooth_val)
ax.plot(grid, pdf, color=neg_palette[j])
def plot_spc(pca, ind, pos_lambdas, neg_lambdas, pos_palette, neg_palette, ax):
for j, lam in enumerate(pos_lambdas):
proj = pca.bary + lam * pca.eig_vecs[:, ind]
ax.plot(pca.spline_basis.xgrid, pca.get_pdf(proj), color=pos_palette[j])
for j, lam in enumerate(neg_lambdas):
proj = pca.bary + lam * pca.eig_vecs[:, ind]
ax.plot(pca.spline_basis.xgrid, pca.get_pdf(proj), color=neg_palette[j])
# +
fig, axes = plt.subplots(nrows=2, ncols=4, figsize=(12, 6))
pos_lambdas = np.linspace(0.01, 100, 10)
neg_lambdas = np.linspace(-0.01, -100, 10)
pos_palette = sns.light_palette("red", n_colors=len(pos_lambdas))
neg_palette = sns.light_palette("navy", n_colors=len(neg_lambdas))
plot_spc(spca, 0, pos_lambdas, neg_lambdas, pos_palette, neg_palette, axes[0][0])
pos_lambdas = np.linspace(0.01, 100, 10)
neg_lambdas = np.linspace(-0.01, -100, 10)
plot_spc(spca, 1, pos_lambdas, neg_lambdas, pos_palette, neg_palette, axes[1][0])
# for i in range(2):
# axes[0][i].set_xlim(-5, 5)
wpca = ProjectedPCA(20)
wpca.fit(data, 2)
pos_lambdas = np.linspace(0.01, 1, 10)
neg_lambdas = np.linspace(-0.01, -1, 10)
plot_wpc(wpca, 0, pos_lambdas, neg_lambdas, pos_palette, neg_palette, axes[0][1], 1.0)
plot_wpc(wpca, 1, pos_lambdas, neg_lambdas, pos_palette, neg_palette, axes[1][1], 10.0)
gpca = GeodesicPCA(20)
gpca.fit(data, 2)
plot_wpc(gpca, 0, pos_lambdas, neg_lambdas, pos_palette, neg_palette, axes[0][2], 3.5)
plot_wpc(gpca, 1, pos_lambdas, neg_lambdas, pos_palette, neg_palette, axes[1][2], 10.0)
npca = NestedPCA(20)
npca.fit(data, 2)
plot_wpc(npca, 0, pos_lambdas, neg_lambdas, pos_palette, neg_palette, axes[0][3], 1.0)
plot_wpc(npca, 1, pos_lambdas, neg_lambdas, pos_palette, neg_palette, axes[1][3], 10.0)
axes[0][0].set_ylabel("1st PD")
axes[1][0].set_ylabel("2nd PD")
axes[0][0].set_title("SIMPLICIAL")
axes[0][1].set_title("WASS - PROJECTED")
axes[0][2].set_title("WASS - GLOBAL")
axes[0][3].set_title("WASS - NESTED")
for i in range(2):
for j in range(4):
axes[i][j].set_ylim(ymin=0.0)
plt.tight_layout()
# plt.savefig("scenario1_pca_comparison.pdf")
# -
# # comparison of geodesics
# +
xgrid = np.linspace(-10, 10, 200)
p1un = norm.pdf(xgrid, -4, 0.8)
p2un = norm.pdf(xgrid, 2, 3)
plt.plot(xgrid, p1un)
plt.plot(xgrid, p2un)
# +
from scipy.integrate import simps
from scipy.interpolate import UnivariateSpline, PchipInterpolator
def inv_clr(f_eval, grid):
out = np.exp(f_eval)
den = simps(out, grid)
return out / den
def clr(f_eval, grid):
log_f = np.log(f_eval)
out = log_f - simps(log_f / (grid[-1] - grid[0]), grid)
return out
def w_dist(f1, f2):
qgrid1 = np.cumsum(f1.f_eval) * (f1.grid[1] - f1.grid[0])
qeval1 = f1.grid
keep = np.where(np.diff(qgrid1) > 1e-5)
quant1 = PchipInterpolator(qgrid1[keep], qeval1[keep])
qgrid2 = np.cumsum(f2.f_eval) * (f2.grid[1] - f2.grid[0])
qeval2 = f2.grid
keep = np.where(np.diff(qgrid2) > 1e-5)
quant2 = PchipInterpolator(qgrid2[keep], qeval2[keep])
er = np.sqrt(simps( (quant1(zero_one_grid) - quant2(zero_one_grid))**2, zero_one_grid ))
return er
# +
import ot
from sklearn.metrics import pairwise_distances
p1 /= np.sum(p1un)
p2 /= np.sum(p2un)
A = np.vstack((p1, p2)).T
n_distributions = A.shape[1]
# loss matrix + normalization
M = pairwise_distances(xgrid.reshape(-1, 1))
M /= M.max()
wgrid = np.concatenate(
[np.linspace(0, 0.45, 5), np.linspace(0.45, 0.55, 90), np.linspace(0.55, 1.0, 5)])
wass_geod = np.empty((len(wgrid), len(xgrid)))
simp_geod = np.empty((len(wgrid), len(xgrid)))
wass_dists = np.zeros_like(wgrid)
simp_dists = np.zeros_like(wgrid)
p1clr = clr(p1un, xgrid)
p2clr = clr(p2un, xgrid)
dwass = ot.emd2(p1, p2, M)
dsimp = simps( (p2clr - p1clr)**2, xgrid)
for i, w in enumerate(wgrid):
print("\r{0} / {1}".format(i + 1, len(wgrid)), end=" ", flush=True)
reg = 1e-3
wass_geod[i, :] = ot.bregman.barycenter(A, M, reg, np.array([w, 1-w]))
wass_dists[i] = ot.sinkhorn2(p1, wass_geod[i, :], M, reg)
curr_simp = w * p1clr + (1-w) * p2clr
simp_geod[i, :] = inv_clr(curr_simp, xgrid)
simp_dists[i] = simps( (curr_simp - p1clr)**2, xgrid)
# -
plt.plot(wgrid, wass_dists)
# plt.plot(wgrid, simp_dists)
dwass = ot.sinkhorn2(p1, p2, M, reg)
plt.plot(xgrid, p1)
plt.plot(xgrid, p2)
idx = np.where(wass_dists > dwass * 0.25)[0][-1]
print(idx)
plt.plot(xgrid, wass_geod[idx, :])
idx = np.where(wass_dists > dwass * 0.5)[0][-1]
print(idx)
plt.plot(xgrid, wass_geod[idx, :])
idx = np.where(wass_dists > dwass * 0.75)[0][-1]
print(idx)
plt.plot(xgrid, wass_geod[idx, :])
plt.plot(xgrid, p1un)
plt.plot(xgrid, p2un)
idx = np.where(simp_dists > dsimp * 0.25)[0][-1]
print(idx)
plt.plot(xgrid, simp_geod[idx, :])
idx = np.where(simp_dists > dsimp * 0.5)[0][-1]
print(idx)
plt.plot(xgrid, simp_geod[idx, :])
idx = np.where(simp_dists > dsimp * 0.75)[0][-1]
print(idx)
plt.plot(xgrid, simp_geod[idx, :])
plt.plot(xgrid, p1)
plt.plot(xgrid, wass_geod[55, :])
# # Analysis of simulations
# +
import pickle
with open("bernstein_sim_res.pickle", "rb") as fp:
res = pickle.load(fp)
# +
basis_range = [5, 10, 15, 25, 50]
ncomp_range = [2, 5, 10]
fig, axes = plt.subplots(nrows=1, ncols=3, figsize=(10, 4))
sim_idx = 0
res["w_errors"][res["w_errors"] > 1] = np.nan
res["w_errors"] = np.ma.array(res["w_errors"], mask=np.isnan(res["w_errors"]))
res["s_errors"][res["s_errors"] > 1] = np.nan
for i in range(3):
mean = np.mean(res["w_errors"][:, i, :], axis=0)
std = np.std(res["w_errors"][:, i, :], axis=0)
axes[i].plot(basis_range, mean, label="projected")
axes[i].fill_between(basis_range, mean + std, mean - std, alpha=0.3)
mean = np.mean(res["s_errors"][:, i, :], axis=0)
std = np.std(res["s_errors"][:, i, :], axis=0)
axes[i].plot(basis_range, mean, label="simplicial")
axes[i].fill_between(basis_range, mean + std, mean - std, alpha=0.3)
for i in range(3):
# axes[i].set_title("# Components: {0}".format(ncomp_range[i]))
axes[i].set_ylim(0.001, 0.1)
axes[i].set_xlim(0, 55)
axes[i].set_xticks(np.arange(5, 51, 5))
axes[2].legend()
plt.tight_layout()
# plt.savefig("bernstein_simulation.pdf")
# -
res["w_errors"][:, :, -1]
np.std(res["w_errors"][:, 1, :], axis=0)
# +
xgrid = np.linspace(0, 1, 1000)
from scipy.stats import norm, gamma, beta, dirichlet
def simulate_data(ndata):
L = 500
beta_dens = np.zeros((L, len(xgrid)))
for j in range(L):
beta_dens[j, :] = beta.pdf(xgrid, j + 1, L - j)
out = []
for i in range(ndata):
ws = dirichlet.rvs(np.ones(L) * 0.01)[0]
curr = np.sum(beta_dens * ws[:, np.newaxis], axis=0)
out.append(GlobalTfunction(xgrid, curr))
return out
data = simulate_data(100)
for d in data:
plt.plot(d.grid, d.f_eval)
plt.savefig("bernstein_data.pdf")
# +
import pickle
with open("dpm_sim_res.pickle", "rb") as fp:
res = pickle.load(fp)
# +
dim_range = np.arange(2, 10 + 1, 2)
mean = np.mean(res["p_errors"], axis=0)
std = np.std(res["p_errors"], axis=0)
plt.plot(dim_range, mean, label="projected")
plt.fill_between(dim_range, mean + std, mean - std, alpha=0.3)
mean = np.mean(res["s_errors"], axis=0)
std = np.std(res["s_errors"], axis=0)
plt.plot(dim_range, mean, label="simplicial")
plt.fill_between(dim_range, mean + std, mean - std, alpha=0.3)
mean = np.mean(res["n_errors"], axis=0)
std = np.std(res["n_errors"], axis=0)
plt.plot(dim_range, mean, label="nested")
plt.fill_between(dim_range, mean + std, mean - std, alpha=0.3)
mean = np.mean(res["g_errors"], axis=0)
std = np.std(res["g_errors"], axis=0)
plt.plot(dim_range, mean, label="global")
plt.fill_between(dim_range, mean + std, mean - std, alpha=0.3)
plt.yscale("log")
plt.legend()
plt.savefig("dpm_simulation.pdf")
plt.show()
# +
xgrid = np.linspace(-10, 10, 1000)
from scipy.integrate import simps
def simulate_data(ndata):
# approximate a DP by truncation
gamma = 50
L = 500
out = []
for i in range(ndata):
weights = np.random.dirichlet(np.ones(L) / L, 1)
atoms = np.empty((L, 2))
atoms[:, 0] = np.random.normal(loc=0.0, scale=2.0, size=L)
atoms[:, 1] = np.random.uniform(0.5, 2.0, size=L)
dens_ = norm.pdf(xgrid.reshape(-1, 1), atoms[:, 0], atoms[:, 1])
dens = np.sum(dens_ * weights, axis=1)
dens += 1e-5
totmass = simps(dens, xgrid)
dens /= totmass
out.append(GlobalTfunction(xgrid, dens))
return out
data = simulate_data(100)
for d in data:
plt.plot(d.grid, d.f_eval)
plt.savefig("dpm_data.pdf")
| Comparison vs Simplicial PCA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
## global imports
import pandas as pd
import numpy as np
import jgraph
from datetime import datetime
# +
# read the data and harmonise = > takes a while to run
file_path= './data/'
covid_df = pd.read_csv(file_path + '/covid19-US.csv')
state_abbreviations_df = pd.read_csv(file_path + '/state_abbreviations.csv' )
abbreviations_dict = dict(zip(state_abbreviations_df.State, state_abbreviations_df.Code))
col_list = ['FL_DATE', 'ORIGIN_CITY_NAME', 'DEST_CITY_NAME', 'FLIGHTS']
flight_jan19 = pd.read_csv(file_path + '/flights-jan19.csv', usecols=col_list)
flight_jan20 = pd.read_csv(file_path + '/flights-jan20.csv', usecols=col_list)
flight_feb19 = pd.read_csv(file_path + '/flights-feb19.csv', usecols=col_list)
flight_mar19 = pd.read_csv(file_path + '/flights-mar19.csv', usecols=col_list)
flight_apr19 = pd.read_csv(file_path + '/flights-apr19.csv', usecols=col_list)
## harmonise dataframes
# convert to datetime makes things slightly faster
covid_df['date']= pd.to_datetime(covid_df.date)
covid_df['day'] = covid_df.date.apply(lambda x: x.day)
covid_df['month'] = covid_df.date.apply(lambda x: x.month)
# name origin as that is how we will agregate
covid_df['origin_state'] = covid_df.state.apply(lambda x: abbreviations_dict[x] if x in abbreviations_dict.keys() else 'other')
covid_df['year'] = 2020
## this is what takes a lot of time
flight_df = flight_jan20
flight_df = flight_df.append([flight_jan19, flight_feb19, flight_mar19, flight_apr19], ignore_index = True)
## might not need that but could be useful later ..takes a while
flight_df['FL_DATE']= pd.to_datetime(flight_df.FL_DATE)
flight_df['day'] = flight_df.FL_DATE.apply(lambda x: x.day)
flight_df['month'] = flight_df.FL_DATE.apply(lambda x: x.month)
flight_df['year'] = flight_df.FL_DATE.apply(lambda x: x.year)
flight_df['origin_state'] = flight_df.ORIGIN_CITY_NAME.apply(lambda x: x.strip().split()[-1])
flight_df['dest_state'] = flight_df.DEST_CITY_NAME.apply(lambda x: x.strip().split()[-1])
flight_df = flight_df.drop(['ORIGIN_CITY_NAME', 'DEST_CITY_NAME', 'FL_DATE'], axis =1)
covid_df = covid_df.drop(['date', 'county', 'fips', 'state'], axis = 1)
# -
## aggregate and sum
flight_df = flight_df.groupby(['day', 'month', 'year', 'origin_state',
'dest_state'])['FLIGHTS'].sum().reset_index().fillna(0)
flight_mar19
flight_df.loc[flight_df.month==3]
print('FLIGHT shape:', flight_df.shape)
print('COVID shape:', covid_df.shape)
covid_df.head()
# +
import networkx as nx
## create death and case graphs for a certain date
# choose date of interest
day = 1
month = 4
year = 2019
interesting_flight_data = flight_df.loc[(flight_df.day == day) &
(flight_df.month == month) &
(flight_df.year == year)]
interesting_covid_data = covid_df.loc[(covid_df.day == day) &
(covid_df.month == month), ['cases', 'deaths', 'origin_state']]
interesting_covid_data = interesting_covid_data.groupby(['origin_state'])['deaths',
'cases'].sum().reset_index().fillna(0)
print('Flight-data-of-interest shape:', interesting_flight_data.shape)
print('Covid-data-of-interest shape:', interesting_covid_data.shape)
# create death graph
edge_weights = [i / np.max(interesting_flight_data.FLIGHTS) for i in interesting_flight_data.FLIGHTS]
nodes = list(abbreviations_dict.values()) + ['other']
death_graph = nx.DiGraph()
death_graph.add_nodes_from(nodes)
deaths_dict = dict(zip(interesting_covid_data.origin_state, interesting_covid_data.deaths))
cases_dict = dict(zip(interesting_covid_data.origin_state, interesting_covid_data.cases))
for node in nodes:
## look up for weight of node else 0
weight = 0
if node in deaths_dict.keys():
weight = deaths_dict[node]
death_graph.add_node(node, weight=weight)
for origin, dest, weight in zip(interesting_flight_data.origin_state,
interesting_flight_data.dest_state, edge_weights):
death_graph.add_edge(origin, dest, weight = weight)
# create case graph
edge_weights = [i / np.max(interesting_flight_data.FLIGHTS) for i in interesting_flight_data.FLIGHTS]
nodes = list(abbreviations_dict.values()) + ['other']
case_graph = nx.DiGraph()
case_graph.add_nodes_from(nodes)
cases_dict = dict(zip(interesting_covid_data.origin_state, interesting_covid_data.cases))
for node in nodes:
## look up for weight of node else 0
weight = 0
if node in cases_dict.keys():
weight = cases_dict[node]
case_graph.add_node(node, weight=weight)
for origin, dest, weight in zip(interesting_flight_data.origin_state,
interesting_flight_data.dest_state, edge_weights):
case_graph.add_edge(origin, dest, weight = weight)
# -
interesting_covid_data.head()
# +
## This is to print the graph --need to do something with
# import matplotlib.pyplot as plt
# pos = nx.drawing.layout.circular_layout(death_graph)
# elarge = [(u, v) for (u, v, d) in death_graph.edges(data=True) if d['weight'] > 0.5]
# esmall = [(u, v) for (u, v, d) in death_graph.edges(data=True) if d['weight'] <= 0.5]
# #pos = nx.spring_layout(death_graph) # positions for all nodes
# # nodesG.nodes(data='weight'
# weights = [w for w in list(nx.get_node_attributes(death_graph,'weight').values())]* 10000
# nx.draw_networkx_nodes(death_graph.nodes, pos, node_size = weights)
# # edges
# nx.draw_networkx_edges(death_graph, pos, edgelist=elarge,
# width=6)
# nx.draw_networkx_edges(death_graph, pos, edgelist=esmall,
# width=6, alpha=0.5, edge_color='b', style='dashed')
# # labels
# nx.draw_networkx_labels(death_graph, pos, font_size=10, font_family='sans-serif')
# plt.axis('off')
# plt.show()
| Project/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('..')
# # Convert ID Sequences to Masked Sequences (idseqs_to_mask)
# +
from torch_tweaks import idseqs_to_mask
idseqs = [[1, 1, 0, 0, 2, 2, 3], [1, 3, 2, 1, 0, 0, 2]]
masks = idseqs_to_mask(
idseqs, n_seqlen=6, ignore=[1],
dtype=int, dense=False)
for i, mask in enumerate(masks):
print(f"Mask {i}:")
print(mask.to_dense())
# -
| examples/help1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Distribuciones Estadisticas
# ### Distribución Uniforme
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
a=1
b=100
n=200
data = np.random.uniform(a,b,n)<
# %matplotlib inline
plt.hist(data)
# ## Distribución Normal
data = np.random.randn(1000)
# %matplotlib inline
x=range(1,1001)
plt.plot(x,data) #Conjuntos de valores que se han obtenido con randn
plt.hist(data)
plt.plot(x,sorted(data))
mu = 5.5
sd= 2.5
data= 5.5 + 2.5*np.random.randn(1000) #Para usar una desviacion standar
#Segun los datos usados
plt.hist(data)
# ## Simulación de Monte Carlo
# Generamos dos numeros aleatorios entre 0 y 1
# Calcularemos x * x + y*y
# * Si el valor es inferior a 1 estamos dentro del circulo
# * Si es mayor a 1 estamos fuera del circulo
# Calculamos el numero total de veces que estan dentro del circulo y dividimos entre el numero de intentos obtenidos
def pi_montecarlo(n, n_exp):
pi_avg=0
pi_value_list=[]
n=1000
for i in range(n_exp):
value=0
x = np.random.uniform(0,1,n).tolist()
y=np.random.uniform(0,1,n).tolist()
for j in range(n):
z=np.sqrt(x[j]*x[j] + y[j]*y[j])
if z<=1:
value+=1
float_value= float(value)
pi_value= (float_value * 4 )/n
pi_value_list.append(pi_value)
pi_avg+=pi_value
pi=pi_avg/n_exp
fig=plt.plot(pi_value_list)
return(pi,fig)
pi_montecarlo(10,10)
# ## Dummy Data Sets
n=100000
data = pd.DataFrame(
{
"A": np.random.randn(n),
"B": 1.5 +2.5* np.random.randn(n),
"C": np.random.uniform(5,32,n)
}
)
data.describe()
plt.hist(data.A)
plt.hist(data.B)
plt.hist(data.C)
data= pd.read_csv("../datasets/customer-churn-model/Customer Churn Model.txt")
data.head()
colum_names= data.columns.values.tolist()
a=len(colum_names)
new_data= pd.DataFrame(
{
"Column Name": colum_names,
"A": np.random.randn(a),
"B": np.random.uniform(0,1,a)
}
)
new_data
| 2. Operando Datos/.ipynb_checkpoints/2. Distribuciones Estadisticas-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import itertools
import numpy as np
import pandas as pd
# read sample data
sample_df = pd.read_csv("../../data/sample_data.csv")
sample_df.head(2)
# +
def create_new_categories(x):
try:
categories = x.split('/')
if len(categories) == 3:
return pd.Series({'c1':categories[0].lower().strip(),
'c2':categories[1].lower().strip(),
'c3':categories[2].lower().strip()})
return pd.Series({'c1': None, 'c2':None, 'c3': None})
except:
return pd.Series({'c1': None, 'c2':None, 'c3': None})
# +
new_df = pd.concat([sample_df[['train_id', 'name', 'item_condition_id', 'brand_name','price', 'shipping', 'item_description']],
sample_df.category_name.apply(create_new_categories)], axis=1)
# -
new_df.dropna(inplace=True)
new_df.shape
new_df.head(3)
new_df.c1.value_counts()
print(len(new_df.c2.unique()))
new_df.c2.value_counts()
print(len(new_df.c3.unique()))
new_df.c3.value_counts()
| experiment/data_preprocess/CategoricalNameExtraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="657b2d49"
# # Challenge 08: k-Nearest Neighbors
#
# **_This challenge is brought to you by [@MaldoAlberto](https://github.com/MaldoAlberto) as a community contribution. Thank you!_**
#
# The k-Nearest Neighbors (k-NN) algorithm is one of the best known classification algorithms and an example of [supervised learning](https://en.wikipedia.org/wiki/Supervised_learning). The objective of a classification algorithm is to identify correctly which class a data point belongs to. The training data used to train this algorithm includes labels (e.g. "1", "2", "dog", "cat", etc.) already. The k in k-NN is the number of nearest neighbors that we are going to compare a data point to in order to choose which class it belongs, therefore this classifier will consider the test set that we will call states $|a_i>$ whose labels are determined by finding the minimum distance to the states of the training set $|b_j>$, whose labels are known beforehand [[1]](https://arxiv.org/pdf/2003.09187.pdf).
#
# 
#
# Figure 1. Example of a k-NN with k=3. (image obtained from [here](https://github.com/artifabrian/dynamic-knn-gpu))
# + [markdown] id="fI7bt50CjF9M"
# To find the distance between two instances, in classical computation one of these is the Euclidean distance, $\lvert a-b \rvert = \sqrt{\sum_{i=1}^N (a_i-b_i)^2}$. In the case of quantum computation, its equivalence is achieved from the inner product (or dot product) which is defined as $\lvert a-b \rvert = \lvert a \rvert \lvert b \rvert -a \cdot b$, where |a> and |b> are two quantum states. To implement the inner product corresponding to the Euclidean distance, it is necessary to use a quantum subroutine known as the SWAP test [[2]](https://arxiv.org/pdf/1401.2142.pdf), which is a quantum algorithm that can be used to estimate the fidelity of two pure states $|\phi>$ and $|\psi>$, i.e., $F=\lvert<\psi|\phi>\rvert ^2$. Figure (2) shows the quantum circuit to perform such an operation:
#
# 
#
# The measurement of the first qubit in the quantum circuit will indicate the following probability:
#
# $P(q_0 = 0) =\frac{1}{2}+\frac{1}{2}\lvert<\psi|\phi>\rvert^2 $
#
# If the two states are orthogonal, then the probability will be $\frac{1}{2}$ and if they match, then the probability will be $1$.
#
# -
# ### The Challenge
#
# **Task 1**
# Design a quantum circuit to perform k-NN classification using only two classes of your preference (for example classes "3", "7") from the MNIST dataset. Test your classifier using the following values of k: 1,3,5,7. With this being a binary classification, use odd values to break a tie to which class it belongs (similar to the [Pigeonhole principle](https://en.wikipedia.org/wiki/Pigeonhole_principle)). Achieve classification with at least 85% accuracy.
#
# How to obtain the MNIST dataset:
# https://www.tensorflow.org/tutorials/quickstart/beginner
# https://scikit-learn.org/stable/auto_examples/linear_model/plot_sparse_logistic_regression_mnist.html
# https://csc.lsu.edu/~saikat/n-mnist/
#
# **Task 2**
# Try solving the problem with a decreased number of qubits for holding the image information (using a classical [[3]](https://arxiv.org/pdf/1908.02626.pdf),[[4]](https://arxiv.org/pdf/1404.1100.pdf),[[5]](https://www.mdpi.com/2504-3900/54/1/40/pdf) preprocessing step) or using the idea of the [March 2021 Challenge](https://github.com/qosf/monthly-challenges/tree/main/challenge-2021.03-mar/challenge-2021.03-mar.ipynb).
#
# **Task 3 (Bonus)**
# Increase the challenge to include a third class, such as using classes "0", "7" and "8" with an accuracy of at least 85%. Having more than two classes I considered this [post](https://towardsdatascience.com/a-simple-introduction-to-k-nearest-neighbors-algorithm-b3519ed98e) to identify the value of k that would be the best for the classification. What about a fourth class? How high can you go?
# + [markdown] id="bcb2f283"
# ### References
#
# [1] Afham, & <NAME> & Goyal, Sandeep. (2020). Quantum k-nearest neighbor machine learning algorithm.
#
# [2] <NAME> & <NAME> & <NAME>. (2015). Quantum algorithms for nearest-neighbor methods for supervised and unsupervised learning. Quantum Information and Computation. 15. 318-358.
#
# ### Resources
#
# [3] Rudolph, Marco & <NAME> & <NAME>. (2019). Structuring Autoencoders.
#
# [4] <NAME>. (2014). A Tutorial on Principal Component Analysis. Educational. 51.
#
# [5] <NAME>. Feature Selection in Big Image Datasets. Proceedings 2020, 54, 40. https://doi.org/10.3390/proceedings2020054040
#
# [6] <NAME> & Xue, Xiling & Liu, Heng & Tan, Jianing & <NAME>. (2017). Quantum Algorithm for K-Nearest Neighbors Classification Based on the Metric of Hamming Distance. International Journal of Theoretical Physics. 56. 10.1007/s10773-017-3514-4.
#
# [7] <NAME> & <NAME>. (2008). Quantum Random Access Memory. Physical review letters. 100. 160501. 10.1103/PhysRevLett.100.160501.
#
# [8] <NAME> & <NAME>. (2018). MNIST Dataset Classification Utilizing k-NN Classifier with Modified Sliding Window Metric.
#
# [9] <NAME> & <NAME>. (2020). Quantum Computing: Start your journey with Qiskit! | Open Source for You (December 2020). 10.13140/RG.2.2.32232.08961
# -
| challenge-2021.06-jun/challenge-2021.06-jun.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Test Jupyter with python notebook
#
# I installed python2 and python3 with MacPorts.
# I even selected python36 before install juptyer with pip36.
# Still, only Python 2 notebook available. Sad.
#
# I figured it out.
x=[1,2,3]
x
import sys
sys.version_info
sys.version
| jupyter_python2_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + jupyter={"source_hidden": true}
import pandas as pd
import numpy as np
import os
import time
from matplotlib import pyplot as plt
from sklearn import datasets
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import auc
from sklearn.metrics import plot_roc_curve
from sklearn.model_selection import StratifiedShuffleSplit
from sklearn.preprocessing import Normalizer
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
from joblib import dump, load
from sklearn.neural_network import MLPClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import classification_report
import json
# -
# # Code for visualise the default number
# + jupyter={"source_hidden": true}
def visualise_default():
year_list = ['2004', '2005','2006','2007','2008','2009','2010','2011','2012','2013','2014','2015','2016','2017','2018']
data_ = []
default = []
total = []
for year_ in year_list:
print("Loading data...")
data_path = 'data_flag/{year}_flag.csv'.format(year=year_)
cols = pd.read_csv(data_path).columns
data = pd.read_csv(data_path, usecols = cols[1:])
y = np.asarray(data['default_flag'].astype(int))
u_ele, ct_ele = np.unique(y, return_counts=True)
default.append(ct_ele[1])
total.append(np.size(y))
data_.append(default)
data_.append(total)
columns = ['%d' % x for x in np.arange(2004,2019)]
rows = ['default','number of accounts']
fig, ax = plt.subplots(figsize=(30, 20))
values = np.arange(0, 2500, 500)
colors = plt.cm.BuPu(np.linspace(0, 0.5, len(rows)))
n_rows = len(data_)
index = np.arange(len(columns)) + 0.6
bar_width = 0.4
# Initialize the vertical-offset for the stacked bar chart.
y_offset = np.zeros(len(columns))
# Plot bars and create text labels for the table
for row in range(n_rows):
bar = plt.bar(index, data_[row], bar_width, bottom=y_offset, color=colors[row])
y_offset = y_offset + data_[row]
cell_text.append(['%d' % x for x in y_offset])
colors = colors[::1]
# Add a table at the bottom of the axes
the_table = plt.table(cellText=data_,
rowLabels=rows,
rowColours=colors,
colLabels=columns,
loc='bottom')
the_table.set_fontsize(25)
the_table.scale(1.0, 2.0) # may help
# Adjust layout to make room for the table:
values = np.arange(0, 2000000, 500000)
plt.ylabel("Person",fontsize = 20)
plt.yticks(values, ['%d' % val for val in values], fontsize = 20)
plt.xticks([])
plt.title('Default counts by years',fontsize = 20)
plt.show()
fig.savefig('./Defaults.jpg')
# -
# # Load Data
#
# + jupyter={"source_hidden": true}
def load_data(data_path, data_path_time):
time_start=time.time()
#Load data
data_list = []
for fname in sorted(os.listdir(data_path)):
subject_data_path = os.path.join(data_path, fname)
print(subject_data_path)
if not os.path.isfile(subject_data_path): continue
data_list.append(
pd.read_csv(
subject_data_path,
sep='|',
header=None,
names = [
'CREDIT_SCORE',
'FIRST_PAYMENT_DATE',
'FIRST_TIME_HOMEBUYER_FLAG',
'4','5','6',
'NUMBER_OF_UNITS',
'OCCUPANCY_STATUS',
'9',
'ORIGINAL_DTI_RATIO',
'ORIGINAL_UPB',
'ORIGINAL_LTV',
'ORIGINAL_INTEREST_RATE',
'CHANNEL',
'15',
'PRODUCT_TYPE',
'PROPERTY_STATE',
'PROPERTY_TYPE',
'19',
'LOAN_SQ_NUMBER',
'LOAN_PURPOSE',
'ORIGINAL_LOAN_TERM',
'NUMBER_OF_BORROWERS',
'24','25','26'#,'27'#data from every year may have different column number
#2004-2007: 27 2008: 26 2009: 27
],
usecols=[
'CREDIT_SCORE',
'FIRST_TIME_HOMEBUYER_FLAG',
'NUMBER_OF_UNITS',
'OCCUPANCY_STATUS',
'ORIGINAL_DTI_RATIO',
'ORIGINAL_UPB',
'ORIGINAL_LTV',
'ORIGINAL_INTEREST_RATE',
'CHANNEL',
'PROPERTY_TYPE',
'LOAN_SQ_NUMBER',
'LOAN_PURPOSE',
'ORIGINAL_LOAN_TERM',
'NUMBER_OF_BORROWERS'
],
dtype={'CREDIT_SCORE':np.float_,
'FIRST_TIME_HOMEBUYER_FLAG':np.str,
'NUMBER_OF_UNITS':np.int_,
'OCCUPANCY_STATUS':np.str,
'ORIGINAL_DTI_RATIO':np.float_,
'ORIGINAL_UPB':np.float_,
'ORIGINAL_LTV':np.float_,
'ORIGINAL_INTEREST_RATE':np.float_,
'CHANNEL':np.str,
'PROPERTY_TYPE':np.str,
'LOAN_SQ_NUMBER':np.str,
'LOAN_PURPOSE':np.str,
'ORIGINAL_LOAN_TERM':np.int_,
'NUMBER_OF_BORROWERS':np.int_},
low_memory=False
)
)
data = pd.concat(data_list)
#Load data with time
data_p_list=[]
for fname in sorted(os.listdir(data_path_time)):
subject_data_path = os.path.join(data_path_time, fname)
print(subject_data_path)
if not os.path.isfile(subject_data_path): continue
data_p_list.append(
pd.read_csv(subject_data_path,
sep='|',
header=None,
usecols=[0,3,4],
dtype={'0':np.str, '3':np.str, '4':np.int_}
)
)
#data_p = pd.concat(data_p_list)
#Calculate default
default_list=[]
for data_p in data_p_list:
data_p[3] = data_p[3].astype(str)
clean_index = data_p.iloc[:,1].str.isdigit()
data_p_cleaned = data_p[clean_index].copy()
data_p_cleaned[3] = data_p_cleaned[3].astype(int)
data_less_than_48 = data_p_cleaned[data_p_cleaned[4] < 48]
default_list.append(data_less_than_48[data_less_than_48[3] > 2])
data_default = pd.concat(default_list)
default_index = data['LOAN_SQ_NUMBER'].isin(data_default[0].tolist())
data['default_flag']=default_index
data.drop(columns=['LOAN_SQ_NUMBER'], inplace=True)
#data.to_csv('data/historical_data_withflag.csv',index=False)
#Imputation
CREDIT_SCORE = data['CREDIT_SCORE']
OIR = data['ORIGINAL_DTI_RATIO']
LTV = data['ORIGINAL_LTV']
CREDIT_clean = CREDIT_SCORE[CREDIT_SCORE != 9999]
OIR_clean = OIR[OIR != 999]
LTV_clean = LTV[LTV != 999]
data['CREDIT_SCORE'] = data['CREDIT_SCORE'].apply(lambda x : CREDIT_clean.mean() if x == 9999 else x)
data['ORIGINAL_DTI_RATIO'] = data['ORIGINAL_DTI_RATIO'].apply(lambda x : OIR_clean.mean() if x == 999 else x)
data['ORIGINAL_LTV'] = data['ORIGINAL_LTV'].apply(lambda x : LTV_clean.mean() if x == 999 else x)
#Timer stop
time_end=time.time()
print('Finished loading, time cost:',time_end-time_start,'s')
return data
# -
# # Standardize
# + jupyter={"source_hidden": true}
def data_standardize():
years = ['2004', '2005','2006','2007','2008','2009','2010','2011','2012','2013','2014','2015']
cols = pd.read_csv('data_flag/2004_flag.csv').columns
data = pd.DataFrame(columns = cols[1:].append(pd.Index(['YEAR'])))
for year_ in years:
data_path = 'data_flag/{year}_flag.csv'.format(year=year_)
cols = pd.read_csv(data_path).columns
data_this_year = pd.read_csv(data_path, usecols = cols[1:])
data_this_year['YEAR'] = year_
data = data.append(data_this_year, ignore_index=True)
year_list = data['YEAR']
data.drop(columns=['YEAR'])
data['FIRST_TIME_HOMEBUYER_FLAG'] = data['FIRST_TIME_HOMEBUYER_FLAG'].apply(lambda x : np.NaN if x == '9' else x)
data['NUMBER_OF_UNITS'] = data['NUMBER_OF_UNITS'].apply(lambda x : np.NaN if x == 99 else x)
data['CHANNEL'] = data['CHANNEL'].apply(lambda x : np.NaN if x == 'T' else x)
data['PROPERTY_TYPE'] = data['PROPERTY_TYPE'].apply(lambda x : np.NaN if x == '99' else x)
data['NUMBER_OF_BORROWERS'] = data['NUMBER_OF_BORROWERS'].apply(lambda x : np.NaN if x == 99 else x)
output_array = np.asarray(data['default_flag'].astype(int))
input_values = np.c_[
data[['CREDIT_SCORE',#0
'ORIGINAL_DTI_RATIO',#1
'ORIGINAL_UPB',#2
'ORIGINAL_LTV',#3
'ORIGINAL_LOAN_TERM',#4
'ORIGINAL_INTEREST_RATE'#5
]]
]
scaler = preprocessing.MinMaxScaler()
scaler.fit(input_values)
input_values = scaler.transform(input_values)
input_dummies = np.c_[
np.asarray(pd.get_dummies(data['FIRST_TIME_HOMEBUYER_FLAG'])), # N,Y,9 str remove 9
np.asarray(pd.get_dummies(data['NUMBER_OF_UNITS'])), # 1 2 3 4 99 int remove 99
np.asarray(pd.get_dummies(data['OCCUPANCY_STATUS'])), # P S I str
np.asarray(pd.get_dummies(data['CHANNEL'])), # T R C B str remove T
np.asarray(pd.get_dummies(data['PROPERTY_TYPE'])), # SF PU CO MH CP 99 str remove 99
np.asarray(pd.get_dummies(data['LOAN_PURPOSE'])), # P N C str
np.asarray(pd.get_dummies(data['NUMBER_OF_BORROWERS'])) # 2 1 99 int remove 99
]
input_array = np.c_[
input_values,
input_dummies
]
data_stand = pd.DataFrame(input_array)
data_stand['YEAR'] = year_list
output = pd.DataFrame(output_array)
output['YEAR'] = year_list
for year_ in years:
print('Preparing {year} ...'.format(year=year_))
data_path = 'data_train/{year}/'.format(year=year_)
data_this_year = data_stand.loc[data_stand['YEAR'] == year_]
data_this_year = data_this_year.drop(columns=['YEAR'])
output_this_year = output.loc[output['YEAR'] == year_]
output_this_year = output_this_year.drop(columns=['YEAR'])
folder = os.getcwd() + '/data_train/{year}/'.format(year=year_)
if not os.path.exists(folder):
os.makedirs(folder)
X_path = data_path + 'input.csv'
y_path = data_path + 'output.csv'
data_this_year.to_csv(X_path.format(year_))
output_this_year.to_csv(y_path.format(year_))
print('{year} done!'.format(year=year_))
# -
# # Logistic Regression
# + jupyter={"source_hidden": true}
def train_log_N_S(data, fig, ax, penalty):
time_start=time.time()
#Get dummy value
data['FIRST_TIME_HOMEBUYER_FLAG'] = data['FIRST_TIME_HOMEBUYER_FLAG'].apply(lambda x : np.NaN if x == '9' else x)
data['NUMBER_OF_UNITS'] = data['NUMBER_OF_UNITS'].apply(lambda x : np.NaN if x == 99 else x)
data['CHANNEL'] = data['CHANNEL'].apply(lambda x : np.NaN if x == 'T' else x)
data['PROPERTY_TYPE'] = data['PROPERTY_TYPE'].apply(lambda x : np.NaN if x == '99' else x)
data['NUMBER_OF_BORROWERS'] = data['NUMBER_OF_BORROWERS'].apply(lambda x : np.NaN if x == 99 else x)
output_array = np.asarray(data['default_flag'].astype(int))
input_array = np.c_[
data[['CREDIT_SCORE',#0
'ORIGINAL_DTI_RATIO',#1
'ORIGINAL_UPB',#2
'ORIGINAL_LTV',#3
'ORIGINAL_LOAN_TERM',#4
'ORIGINAL_INTEREST_RATE'#5
]],
np.asarray(pd.get_dummies(data['FIRST_TIME_HOMEBUYER_FLAG'])), # N,Y,9 str remove 9
np.asarray(pd.get_dummies(data['NUMBER_OF_UNITS'])), # 1 2 3 4 99 int remove 99
np.asarray(pd.get_dummies(data['OCCUPANCY_STATUS'])), # P S I str
np.asarray(pd.get_dummies(data['CHANNEL'])), # T R C B str remove T
np.asarray(pd.get_dummies(data['PROPERTY_TYPE'])), # SF PU CO MH CP 99 str remove 99
np.asarray(pd.get_dummies(data['LOAN_PURPOSE'])), # P N C str
np.asarray(pd.get_dummies(data['NUMBER_OF_BORROWERS'])) # 2 1 99 int remove 99
]
X = input_array
y = output_array
X_train, X_test, y_train, y_test = train_test_split(
X, y,
test_size=0.3,
random_state=13
)
#Normalise
# scaler = preprocessing.StandardScaler().fit(X_train)
# X_train = scaler.transform(X_train)
# X_test = scaler.transform(X_test)
# min_max_scaler = preprocessing.MinMaxScaler()
# X_train = min_max_scaler.fit_transform(X_train)
# X_test = min_max_scaler.transform(X_test)
if penalty == 1:
classifier = LogisticRegression(
tol= 1e-6,
C=0.05,
max_iter = 500,
n_jobs = -1
)
else:
classifier = LogisticRegression(
penalty = 'none',
tol= 1e-6,
#C=0.05,
#class_weight = 'balanced',
#class_weight = {0:0.01, 1:0.99},
#solver='sag',
max_iter=500,
n_jobs = -1
)
classifier.fit(X_train,y_train)
# viz = plot_roc_curve(
# classifier,
# X_test,
# y_test,
# name='Test ROC'.format(0),
# alpha=0.5, lw=1, ax=ax
# )
# viz_train = plot_roc_curve(
# classifier,
# X_train,
# y_train,
# name='Train ROC'.format(1),
# alpha=0.5, lw=1, ax=ax
# )
# ax.plot([0, 1], [0, 1], linestyle='--', lw=2, color='r',
# label='Chance', alpha=.8)
# ax.set(xlim=[-0.05, 1.05], ylim=[-0.05, 1.05],
# title="Receiver operating characteristic example")
# ax.legend(loc="lower right")
time_end=time.time()
print('Training done, time cost:',time_end-time_start,'s')
#print(classifier.predict_proba(X_test)[:, 1])
return classifier
# + jupyter={"source_hidden": true}
def train_log_Y_S(X, y, fig, ax, penalty):
time_start=time.time()
X_train, X_test, y_train, y_test = train_test_split(
X, y,
test_size=0.3,
random_state=13
)
if penalty == 1:
classifier = LogisticRegression(
tol= 1e-5,
C=0.05,
max_iter=500,
n_jobs = -1
)
else:
classifier = LogisticRegression(
penalty = 'none',
tol= 1e-5,
#class_weight = 'balanced',
#class_weight = {0:0.01, 1:0.99},
#solver='sag',
max_iter=500,
n_jobs = -1
)
classifier.fit(X_train,y_train)
# viz = plot_roc_curve(
# classifier,
# X_test,
# y_test,
# name='Test ROC'.format(0),
# alpha=0.5, lw=1, ax=ax
# )
# viz_train = plot_roc_curve(
# classifier,
# X_train,
# y_train,
# name='Train ROC'.format(1),
# alpha=0.5, lw=1, ax=ax
# )
time_end=time.time()
print('Training done, time cost:',time_end-time_start,'s')
return classifier
# + jupyter={"source_hidden": true}
def train_log_diff(X, y, c):
time_start=time.time()
X_train, X_test, y_train, y_test = train_test_split(
X, y,
test_size=0.3,
random_state=13
)
classifier = LogisticRegression(
penalty = 'l2',
tol= 1e-5,
C=c,
#class_weight = 'balanced',
#class_weight = {0:0.01, 1:0.99},
#solver='sag',
max_iter=1500,
n_jobs = -1
)
classifier.fit(X_train,y_train)
time_end=time.time()
print('Training done, time cost:',time_end-time_start,'s')
return classifier
# + jupyter={"source_hidden": true}
def train_log_cross_validation(data):
#Get dummy value
output_array = np.asarray(data['default_flag'].astype(int))
input_array = np.c_[
data[['CREDIT_SCORE',#0
'ORIGINAL_DTI_RATIO',#1
'ORIGINAL_UPB',#2
'ORIGINAL_LTV',#3
'ORIGINAL_LOAN_TERM',#4
'ORIGINAL_INTEREST_RATE'#5
]],
np.asarray(pd.get_dummies(data['FIRST_TIME_HOMEBUYER_FLAG'])),
np.asarray(pd.get_dummies(data['NUMBER_OF_UNITS'])),
np.asarray(pd.get_dummies(data['OCCUPANCY_STATUS'])),
np.asarray(pd.get_dummies(data['CHANNEL'])),
np.asarray(pd.get_dummies(data['PROPERTY_TYPE'])),
np.asarray(pd.get_dummies(data['LOAN_PURPOSE'])),
np.asarray(pd.get_dummies(data['NUMBER_OF_BORROWERS']))
]
#Normalise
min_max_scaler = preprocessing.MinMaxScaler()
input_array_N = min_max_scaler.fit_transform(input_array)
X = input_array_N
y = output_array
#devide Flods for cv
cv = StratifiedKFold(n_splits=6)
#define classifier
classifier = LogisticRegression(
solver='saga',
max_iter=1500
)
#tpr lists and auc value list
tprs = []
aucs = []
#For ploting, prepare 500 points from 0-1
mean_fpr = np.linspace(0, 1, 500)
#Loop training for every fold
for i, (train, test) in enumerate(cv.split(X, y)):
classifier.fit(X[train], y[train])
# put the curve in ax through 'ax = ax'
viz = plot_roc_curve(classifier, X[test], y[test],
name='ROC fold {}'.format(i),
alpha=0.3, lw=1, ax=ax)
#Plot every point (500) form 0-1, similiar to bin
interp_tpr = np.interp(mean_fpr, viz.fpr, viz.tpr)
interp_tpr[0] = 0.0
#Buff the result
tprs.append(interp_tpr)
aucs.append(viz.roc_auc)
#Plot chance
ax.plot([0, 1], [0, 1], linestyle='--', lw=2, color='r',
label='Chance', alpha=.8)
#mean value for each colomn
mean_tpr = np.mean(tprs, axis=0)
mean_tpr[-1] = 1.0
mean_auc = auc(mean_fpr, mean_tpr)
std_auc = np.std(aucs)
#Plot mean
ax.plot(mean_fpr, mean_tpr, color='b',
label=r'Mean ROC (AUC = %0.2f $\pm$ %0.2f)' % (mean_auc, std_auc),
lw=2, alpha=.8)
#Plot standard tpr (Doesn't know the point for this step yet)
std_tpr = np.std(tprs, axis=0)
tprs_upper = np.minimum(mean_tpr + std_tpr, 1)
tprs_lower = np.maximum(mean_tpr - std_tpr, 0)
ax.fill_between(mean_fpr, tprs_lower, tprs_upper, color='grey', alpha=.2,
label=r'$\pm$ 1 std. dev.')
#Configure the diagram
ax.set(xlim=[-0.05, 1.05], ylim=[-0.05, 1.05],
title="Receiver operating characteristic example")
ax.legend(loc="lower right")
# -
# # NN
# + jupyter={"source_hidden": true}
# def train_NN_cv(data, year):
# time_start=time.time()
# #prepare config list
# conf_list = [0.0001, 0.001, 0.01, 0.1]
# #conf_list = [(2,2,),(3,3,),(5,5,),(10,10,)] #dont forget to change name!!!!!!!
# #conf_list = [(2,),(5,),(10,),(15,)]
# conf_name = "alpha"
# para = dict()
# #Get dummy value
# data['FIRST_TIME_HOMEBUYER_FLAG'] = data['FIRST_TIME_HOMEBUYER_FLAG'].apply(lambda x : np.NaN if x == '9' else x)
# data['NUMBER_OF_UNITS'] = data['NUMBER_OF_UNITS'].apply(lambda x : np.NaN if x == 99 else x)
# data['CHANNEL'] = data['CHANNEL'].apply(lambda x : np.NaN if x == 'T' else x)
# data['PROPERTY_TYPE'] = data['PROPERTY_TYPE'].apply(lambda x : np.NaN if x == '99' else x)
# data['NUMBER_OF_BORROWERS'] = data['NUMBER_OF_BORROWERS'].apply(lambda x : np.NaN if x == 99 else x)
# output_array = np.asarray(data['default_flag'].astype(int))
# input_array = np.c_[
# data[['CREDIT_SCORE',#0
# 'ORIGINAL_DTI_RATIO',#1
# 'ORIGINAL_UPB',#2
# 'ORIGINAL_LTV',#3
# 'ORIGINAL_LOAN_TERM',#4
# 'ORIGINAL_INTEREST_RATE'#5
# ]]
# ]
# scaler = preprocessing.StandardScaler().fit(input_array)
# input_array_N = scaler.transform(input_array)
# #input_array_N = preprocessing.normalize(input_array, norm='l2')
# X = np.c_[
# input_array_N,
# np.asarray(pd.get_dummies(data['FIRST_TIME_HOMEBUYER_FLAG'])), # N,Y,9 str remove 9
# np.asarray(pd.get_dummies(data['NUMBER_OF_UNITS'])), # 1 2 3 4 99 int remove 99
# np.asarray(pd.get_dummies(data['OCCUPANCY_STATUS'])), # P S I str
# np.asarray(pd.get_dummies(data['CHANNEL'])), # T R C B str remove T
# np.asarray(pd.get_dummies(data['PROPERTY_TYPE'])), # SF PU CO MH CP 99 str remove 99
# np.asarray(pd.get_dummies(data['LOAN_PURPOSE'])), # P N C str
# np.asarray(pd.get_dummies(data['NUMBER_OF_BORROWERS'])) # 2 1 99 int remove 99
# ]
# y = output_array
# #devide Flods for cv
# cv = StratifiedShuffleSplit(n_splits=5, test_size=0.2, random_state=10)
# #tpr lists and auc value list
# auc_all_config = []
# mean_aucs = []
# #For ploting, prepare 500 points from 0-1
# mean_fpr = np.linspace(0, 1, 500)
# #Loop training for every fold
# for conf in conf_list:
# print("Training "+str(conf)+"...")
# tprs = []
# aucs = []
# fig, ax = plt.subplots(figsize=(15, 8))
# for i, (train, test) in enumerate(cv.split(X, y)):
# ##############################################################################################################################
# classifier = MLPClassifier(
# hidden_layer_sizes = (10,),
# alpha = conf,
# learning_rate_init = 0.001,
# tol = 1e-05,
# verbose = True,
# n_iter_no_change = 10
# )
# para = classifier.get_params(deep=True)
# ##############################################################################################################################
# print("Training fold "+str(i)+"...")
# classifier.fit(X[train], y[train])
# # put the curve in ax through 'ax = ax'
# viz = plot_roc_curve(classifier, X[test], y[test],
# name='ROC fold {}'.format(i),
# alpha=0.3, lw=1, ax=ax)
# #Plot every point (500) form 0-1, similiar to bin
# interp_tpr = np.interp(mean_fpr, viz.fpr, viz.tpr)
# interp_tpr[0] = 0.0
# #Buff the result
# tprs.append(interp_tpr)
# aucs.append(viz.roc_auc)
# #store the auc value for this conf
# std = np.std(aucs)
# aucs.append(std)
# auc_all_config.append(aucs)
# #Plot chance
# ax.plot([0, 1], [0, 1], linestyle='--', lw=2, color='r',
# label='Chance', alpha=.8)
# #mean value for each colomn
# mean_tpr = np.mean(tprs, axis=0)
# mean_tpr[-1] = 1.0
# mean_auc = auc(mean_fpr, mean_tpr)
# #store all mean_auc together
# mean_aucs.append(mean_auc)
# std_auc = np.std(aucs)
# #Plot mean
# ax.plot(mean_fpr, mean_tpr, color='b',
# label=r'Mean ROC (AUC = %0.2f $\pm$ %0.2f)' % (mean_auc, std_auc),
# lw=2, alpha=.8)
# #Configure the diagram
# ax.set(xlim=[-0.05, 1.05], ylim=[-0.05, 1.05],
# title="Receiver operating characteristic example")
# ax.legend(loc="lower right")
# file_name = str(conf)+conf_name+str(year)+'.jpg'
# folder = os.getcwd() + '/NN_cv/'+year+'/'+conf_name
# if not os.path.exists(folder):
# os.makedirs(folder)
# fig.savefig('./NN_cv/'+year+'/'+conf_name+'/'+file_name)
# txt_head = conf_name+'/'.join(str(x) for x in conf_list)
# file_name_all = './NN_cv/'+year+'/'+conf_name+'/'+str(year)+'all.txt'
# file_name_mean = './NN_cv/'+year+'/'+conf_name+'/'+str(year)+'mean.txt'
# file_name_para = './NN_cv/'+year+'/'+conf_name+'/'+'other_para.json'
# folder = os.getcwd() + '/NN_cv/'+year+'/'+conf_name
# if not os.path.exists(folder):
# os.makedirs(folder)
# np.savetxt(file_name_all,auc_all_config,fmt='%.7f',delimiter=',', header=txt_head)
# np.savetxt(file_name_mean,mean_aucs,fmt='%.7f',delimiter=',', header=txt_head)
# with open(file_name_para, 'w') as fp:
# json.dump(para, fp, indent=4)
# + jupyter={"source_hidden": true}
def train_NN_cv(X, y):
# Split the dataset in two equal parts
X_train, X_test, y_train, y_test = train_test_split(
X, y,
test_size=0.3,
stratify = y,
random_state=13
)
# Set the parameters by cross-validation
tuned_parameters = [{
'hidden_layer_sizes': [(5,),(10,),(15,),(5,5)],
'activation': ['logistic', 'tanh', 'relu'],
'alpha': [0.1, 0.01, 0.001],
'learning_rate_init':[0.01, 0.001, 0.0001]
}]
print("# Tuning hyper-parameters for AUC")
clf = GridSearchCV(
MLPClassifier(),
tuned_parameters,
scoring='roc_auc',
n_jobs=7,
verbose=1
)
clf.fit(X_train, y_train)
print("Best parameters set found on development set:")
print()
print(clf.best_params_)
print()
print("Grid scores on development set:")
print()
means = clf.cv_results_['mean_test_score']
stds = clf.cv_results_['std_test_score']
for mean, std, params in zip(means, stds, clf.cv_results_['params']):
print("%0.3f (+/-%0.03f) for %r"
% (mean, std * 2, params))
print()
print("Detailed classification report:")
print()
print("The model is trained on the full development set.")
print("The scores are computed on the full evaluation set.")
print()
y_true, y_pred = y_test, clf.predict(X_test)
print(classification_report(y_true, y_pred))
print()
# + jupyter={"source_hidden": true}
def train_NN_diff(X, y, conf):
# train test Random state 13
time_start=time.time()
#prepare config list
X_train, X_test, y_train, y_test = train_test_split(
X, y,
test_size=0.3,
stratify = y,
random_state = 13
)
classifier = MLPClassifier(
hidden_layer_sizes = conf,
alpha = 0.001,
learning_rate_init = 0.001,
tol = 1e-05,
verbose = True,
n_iter_no_change = 10
)
classifier.fit(X_train,y_train)
time_end=time.time()
print('Training done, time cost:',time_end-time_start,'s')
return classifier
# + jupyter={"source_hidden": true}
def train_NN(X, y, year):
# {'activation': 'tanh', 'alpha': 0.001, 'hidden_layer_sizes': (5,), 'learning_rate_init': 0.001}
# train test Random state 13
time_start=time.time()
#prepare config list
para = dict()
X_train, X_test, y_train, y_test = train_test_split(
X, y,
test_size=0.3,
stratify = y,
random_state = 13
)
fig, ax = plt.subplots(figsize=(15, 8))
classifier = MLPClassifier(
hidden_layer_sizes = (5,),
activation = 'tanh',
alpha = 0.001,
learning_rate_init = 0.001,
tol = 1e-04,
verbose = True
)
para = classifier.get_params(deep=True)
folder = os.getcwd() + '/NN/'
if not os.path.exists(folder):
os.makedirs(folder)
file_name_para = folder+year+'para.json'
with open(file_name_para, 'w') as fp:
json.dump(para, fp, indent=4)
classifier.fit(X_train,y_train)
viz = plot_roc_curve(
classifier,
X_test,
y_test,
name='Test ROC'.format(0),
alpha=0.5, lw=1, ax=ax
)
viz_train = plot_roc_curve(
classifier,
X_train,
y_train,
name='Train ROC'.format(1),
alpha=0.5, lw=1, ax=ax
)
ax.plot([0, 1], [0, 1], linestyle='--', lw=2, color='r',
label='Chance', alpha=.8)
ax.set(xlim=[-0.05, 1.05], ylim=[-0.05, 1.05],
title="Receiver operating characteristic example")
ax.legend(loc="lower right")
time_end=time.time()
print('Training done, time cost:',time_end-time_start,'s')
plt.show()
fig.savefig(folder+'AUC_{year}_NN.jpg'.format(year=year_))
fig_loss, ax_loss = plt.subplots(figsize=(15, 8))
loss_curve = classifier.loss_curve_
ax_loss.plot(np.arange(1, len(loss_curve)+1), loss_curve )
plt.show()
fig_loss.savefig(folder+'train_curve_{year}_NN.jpg'.format(year=year_))
return classifier
# -
# # Random Forest
# + jupyter={"source_hidden": true}
def train_random_forest_cv(X, y):
# Split the dataset in two equal parts
X_train, X_test, y_train, y_test = train_test_split(
X, y,
test_size=0.3,
stratify = y,
random_state=13
)
# Set the parameters by cross-validation
tuned_parameters = [{
'n_estimators': [800, 1200],
'max_depth': [10, 12, 14],
'min_samples_split': [2, 4]
}]
print("# Tuning hyper-parameters for AUC")
clf = GridSearchCV(
RandomForestClassifier(),
tuned_parameters,
scoring='roc_auc',
n_jobs=7,
verbose=3
)
clf.fit(X_train, y_train)
print("Best parameters set found on development set:")
print()
print(clf.best_params_)
print()
print("Grid scores on development set:")
print()
means = clf.cv_results_['mean_test_score']
stds = clf.cv_results_['std_test_score']
for mean, std, params in zip(means, stds, clf.cv_results_['params']):
print("%0.3f (+/-%0.03f) for %r"
% (mean, std * 2, params))
print()
print("Detailed classification report:")
print()
print("The model is trained on the full development set.")
print("The scores are computed on the full evaluation set.")
print()
y_true, y_pred = y_test, clf.predict(X_test)
print(classification_report(y_true, y_pred))
print()
# + jupyter={"source_hidden": true}
#{'max_depth': 10, 'min_samples_split': 2, 'n_estimators': 800}
def train_random_forest(X, y):
# Split the dataset in two equal parts
X_train, X_test, y_train, y_test = train_test_split(
X, y,
test_size=0.3,
stratify = y,
random_state=13
)
print("# Tuning hyper-parameters for AUC")
clf = RandomForestClassifier(
n_estimators = 800,
max_depth = 10,
min_samples_split = 2,
n_jobs=6,
verbose = 2
)
clf.fit(X_train, y_train)
return clf
# -
#{'max_depth': 10, 'min_samples_split': 2, 'n_estimators': 800}
def train_RF_diff(X, y, conf):
# Split the dataset in two equal parts
X_train, X_test, y_train, y_test = train_test_split(
X, y,
test_size=0.3,
stratify = y,
random_state=13
)
clf = RandomForestClassifier(
n_estimators = conf,
max_depth = 10,
min_samples_split = 2,
n_jobs=7,
verbose = 2
)
clf.fit(X_train, y_train)
return clf
# # Main Function
if __name__ == '__main__':
year_list = ['2004','2005','2006','2007','2008','2009','2010','2011','2012','2013','2014','2015']
year_list_2004 = ['2007','2008','2009','2010']
c_list = [0.05, 0.5, 1.0, 5.0, 10.0]
conf_list = [5, 50, 100, 800]
#model_save_path_NS_NP = './LR_experiments/Penalty_CV/{C}/models/{year}.joblib'
LR_model_save_path_NS_NP = './LR_experiments/N_Penalty/N_Standardized/models/{year}.joblib'
LR_model_save_path_NS_YP = './LR_experiments/Penalty/N_Standardized/models/{year}.joblib'
LR_model_save_path_YS_NP = './LR_experiments/N_Penalty/Standardized/models/{year}.joblib'
LR_model_save_path_YS_YP = './LR_experiments/Penalty/Standardized/models/{year}.joblib'
NN_model_save_path = './NN_models_0.01/models/{year}.joblib'
RF_model_save_path = './RF_models/models/{year}.joblib'
NN_Alpha_model_save_path = './NN_experiments/deep/{a}/models/{year}.joblib'
LR_C_model_save_path = './LR_experiments/Penalty_CV/{C}/models/{year}.joblib'
RF_Exp_model_save_path = './RF_experiments/Tree/{conf}/models/{year}.joblib'
X_path = 'data_train/{year}/input.csv'
y_path = 'data_train/{year}/output.csv'
# Command
mode_list = ["train_NN_cv"]
for mode in mode_list:
if mode == "data_process":
for year_ in year_list:
data_path = 'data/{year}/data'.format(year=year_)
data_path_time = 'data/{year}/data_time'.format(year=year_)
data = load_data(data_path, data_path_time)
data.to_csv('data_flag/{}_flag.csv'.format(year_))
elif mode == "train_NN_cv":
for year_ in year_list_2004:
print("Start year {}...".format(year_))
cols = pd.read_csv(X_path.format(year=year_)).columns
X = pd.read_csv(X_path.format(year=year_), usecols = cols[1:])
cols = pd.read_csv(y_path.format(year=year_)).columns
y = pd.read_csv(y_path.format(year=year_), usecols = cols[1:])
X = X.to_numpy()
y = np.asarray(y['0'].astype(int))
train_NN_cv(X, y)
elif mode =="train_LR_N_S": # Train LR without standardize
for year_ in year_list:
penalty_list = [0, 1]
for penalty in penalty_list:
print("Loading data...")
data_path = 'data_flag/{year}_flag.csv'.format(year=year_)
cols = pd.read_csv(data_path).columns
data = pd.read_csv(data_path, usecols = cols[1:])
fig, ax = plt.subplots(figsize=(15, 8))
print(str(year_)+" training start...")
model = train_log_N_S(data, fig, ax, penalty)
if penalty == 1:
folder = os.getcwd() + '/LR_experiments/Penalty/N_Standardized/models/'
if not os.path.exists(folder):
os.makedirs(folder)
dump(model, LR_model_save_path_NS_YP.format(year=year_))
else:
folder = os.getcwd() + '/LR_experiments/N_Penalty/N_Standardized/models/'
if not os.path.exists(folder):
os.makedirs(folder)
dump(model, LR_model_save_path_NS_NP.format(year=year_))
elif mode =="train_LR_S": # Train LR using data after standardizing
for year_ in year_list:
print("Loading data...")
cols = pd.read_csv(X_path.format(year=year_)).columns
X = pd.read_csv(X_path.format(year=year_), usecols = cols[1:])
cols = pd.read_csv(y_path.format(year=year_)).columns
y = pd.read_csv(y_path.format(year=year_), usecols = cols[1:])
X = X.to_numpy()
y = np.asarray(y['0'].astype(int))
fig, ax = plt.subplots(figsize=(15, 8))
print(str(year_)+" training start...")
penalty_list = [0, 1]
for penalty in penalty_list:
if penalty == 0:
model = train_log_Y_S(X, y, fig, ax, penalty)
folder = os.getcwd() + '/LR_experiments/N_Penalty/Standardized/models/'
if not os.path.exists(folder):
os.makedirs(folder)
dump(model, LR_model_save_path_YS_NP.format(year=year_))
else:
model = train_log_Y_S(X, y, fig, ax, penalty)
folder = os.getcwd() + '/LR_experiments/Penalty/Standardized/models/'
if not os.path.exists(folder):
os.makedirs(folder)
dump(model, LR_model_save_path_YS_YP.format(year=year_))
elif mode =="train_NN":
for year_ in year_list:
print("Loading data...")
cols = pd.read_csv(X_path.format(year=year_)).columns
X = pd.read_csv(X_path.format(year=year_), usecols = cols[1:])
cols = pd.read_csv(y_path.format(year=year_)).columns
y = pd.read_csv(y_path.format(year=year_), usecols = cols[1:])
X = X.to_numpy()
y = np.asarray(y['0'].astype(int))
print(str(year_)+" training start...")
model = train_NN(X, y, year_)
dump(model, NN_model_save_path.format(year=year_))
elif mode =="train_RF":
for year_ in year_list:
print("{} Loading data...".format(year_))
cols = pd.read_csv(X_path.format(year=year_)).columns
X = pd.read_csv(X_path.format(year=year_), usecols = cols[1:])
cols = pd.read_csv(y_path.format(year=year_)).columns
y = pd.read_csv(y_path.format(year=year_), usecols = cols[1:])
X = X.to_numpy()
y = np.asarray(y['0'].astype(int))
model = train_random_forest(X, y)
dump(model, RF_model_save_path.format(year=year_))
elif mode =="train_RF_cv":
for year_ in year_list_2004:
print("Loading data...")
cols = pd.read_csv(X_path.format(year=year_)).columns
X = pd.read_csv(X_path.format(year=year_), usecols = cols[1:])
cols = pd.read_csv(y_path.format(year=year_)).columns
y = pd.read_csv(y_path.format(year=year_), usecols = cols[1:])
X = X.to_numpy()
y = np.asarray(y['0'].astype(int))
train_random_forest_cv(X, y)
elif mode =="train_LR_C":
for year_ in year_list:
for c in c_list:
print("C="+str(c)+"Loading data...")
cols = pd.read_csv(X_path.format(year=year_)).columns
X = pd.read_csv(X_path.format(year=year_), usecols = cols[1:])
cols = pd.read_csv(y_path.format(year=year_)).columns
y = pd.read_csv(y_path.format(year=year_), usecols = cols[1:])
X = X.to_numpy()
y = np.asarray(y['0'].astype(int))
print(str(year_)+" training start...")
model = train_log_diff(X, y, c)
folder = os.getcwd() + '/LR_experiments/Penalty_CV/{}/models/'.format(c)
if not os.path.exists(folder):
os.makedirs(folder)
dump(model, LR_C_model_save_path.format(C=c, year=year_))
elif mode =="train_NN_alpha":
for year_ in year_list:
for conf in conf_list:
print("Conf = "+str(conf)+" Loading data...")
cols = pd.read_csv(X_path.format(year=year_)).columns
X = pd.read_csv(X_path.format(year=year_), usecols = cols[1:])
cols = pd.read_csv(y_path.format(year=year_)).columns
y = pd.read_csv(y_path.format(year=year_), usecols = cols[1:])
X = X.to_numpy()
y = np.asarray(y['0'].astype(int))
print(str(year_)+" training start...")
model = train_NN_diff(X, y, conf)
folder = os.getcwd() + '/NN_experiments/deep/{}/models/'.format(conf)
if not os.path.exists(folder):
os.makedirs(folder)
dump(model, NN_Alpha_model_save_path.format(a=conf, year=year_))
elif mode =="train_RF_conf":
for year_ in year_list:
for conf in conf_list:
print("Conf = "+str(conf)+" Loading data...")
cols = pd.read_csv(X_path.format(year=year_)).columns
X = pd.read_csv(X_path.format(year=year_), usecols = cols[1:])
cols = pd.read_csv(y_path.format(year=year_)).columns
y = pd.read_csv(y_path.format(year=year_), usecols = cols[1:])
X = X.to_numpy()
y = np.asarray(y['0'].astype(int))
print(str(year_)+" training start...")
model = train_RF_diff(X, y, conf)
folder = os.getcwd() + '/RF_experiments/Tree/{}/models/'.format(conf)
if not os.path.exists(folder):
os.makedirs(folder)
dump(model, RF_Exp_model_save_path.format(conf=conf, year=year_))
elif mode =="Standard":
data_standardize()
| Pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercises - Chapter 2
#
# ## Solutions
# ## Exercise 2.1
#
# Repeating my advice from the previous chapter, whenever you learn a new feature,
# you should try it out in interactive mode and make errors on purpose to see what goes wrong.
#
# - We have seen that `n = 42` is legal. What about `42 = n?`
42 = n # error
# - How about `x = y = 1?`
x = y = 1 # valid
# - In some languages every statement ends with a semi-colon, ;. What happens if you put a semi-colon at the end of a Python statement?
n = 42; # valid
# - What if you put a period at the end of a statement?
n = '42'. # error
# - In math notation you can multiply $x$ and $y$ like this: $xy$. What happens if you try that in Python?
x = 2
y = 5
result = xy # xy is interpreted as a new variable
# ***
# ## Exercise 2.2
#
# Practice using the Python interpreter as a calculator:
#
# 1. The volume of a sphere with radius $r$ is $\frac{4}{3}\pi r^3$. What is the volume of a sphere with radius 5?
# +
import math
radius = 5
volume = 4/3 * math.pi * radius**3
volume
# -
# 2. Suppose the cover price of a book is \\$24.95, but bookstores get a 40\% discount. Shipping costs \$3 for the first copy and 75 cents for each additional copy. What is the total wholesale cost for 60 copies?
# +
book_price = 24.95
discount = 40
shipping_first_copy = 3
shipping_additional_copy = 0.75
copies = 60
wholesale = copies * book_price * discount / 100 + shipping_first_copy + (copies - 1) * shipping_additional_copy
wholesale
# -
# 3. If I leave my house at 6:52 am and run 1 mile at an easy pace (8:15 per mile), then 3 miles at tempo (7:12 per mile) and 1 mile at easy pace again, what time do I get home for breakfast?
# +
import datetime
start_time = datetime.datetime(year=2020, month=1, day=1, hour=6, minute=34, second=0)
easy_pace = datetime.timedelta(minutes=8, seconds=15)
tempo = datetime.timedelta(minutes=7, seconds=12)
end_time = start_time + easy_pace + 3 * tempo + easy_pace
end_time.strftime("%H:%M:%S")
| Exercises/Solutions/Think-Python_Exercises_Chapter-2-Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text"
# This is a companion notebook for the book [Deep Learning with Python, Second Edition](https://www.manning.com/books/deep-learning-with-python-second-edition?a_aid=keras&a_bid=76564dff). For readability, it only contains runnable code blocks and section titles, and omits everything else in the book: text paragraphs, figures, and pseudocode.
#
# **If you want to be able to follow what's going on, I recommend reading the notebook side by side with your copy of the book.**
#
# This notebook was generated for TensorFlow 2.6.
# + [markdown] colab_type="text"
# # The mathematical building blocks of neural networks
# + [markdown] colab_type="text"
# ## A first look at a neural network
# + [markdown] colab_type="text"
# **Loading the MNIST dataset in Keras**
# + colab_type="code"
from tensorflow.keras.datasets import mnist
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
# + colab_type="code"
train_images.shape
# + colab_type="code"
len(train_labels)
# + colab_type="code"
train_labels
# + colab_type="code"
test_images.shape
# + colab_type="code"
len(test_labels)
# + colab_type="code"
test_labels
# + [markdown] colab_type="text"
# **The network architecture**
# + colab_type="code"
from tensorflow import keras
from tensorflow.keras import layers
model = keras.Sequential([
layers.Dense(512, activation="relu"),
layers.Dense(10, activation="softmax")
])
# + [markdown] colab_type="text"
# **The compilation step**
# + colab_type="code"
model.compile(optimizer="rmsprop",
loss="sparse_categorical_crossentropy",
metrics=["accuracy"])
# + [markdown] colab_type="text"
# **Preparing the image data**
# + colab_type="code"
train_images = train_images.reshape((60000, 28 * 28))
train_images = train_images.astype("float32") / 255
test_images = test_images.reshape((10000, 28 * 28))
test_images = test_images.astype("float32") / 255
# + [markdown] colab_type="text"
# **"Fitting" the model**
# + colab_type="code"
model.fit(train_images, train_labels, epochs=5, batch_size=128)
# + [markdown] colab_type="text"
# **Using the model to make predictions**
# + colab_type="code"
test_digits = test_images[0:10]
predictions = model.predict(test_digits)
predictions[0]
# + colab_type="code"
predictions[0].argmax()
# + colab_type="code"
predictions[0][7]
# + colab_type="code"
test_labels[0]
# + [markdown] colab_type="text"
# **Evaluating the model on new data**
# + colab_type="code"
test_loss, test_acc = model.evaluate(test_images, test_labels)
print(f"test_acc: {test_acc}")
# + [markdown] colab_type="text"
# ## Data representations for neural networks
# + [markdown] colab_type="text"
# ### Scalars (rank-0 tensors)
# + colab_type="code"
import numpy as np
x = np.array(12)
x
# + colab_type="code"
x.ndim
# + [markdown] colab_type="text"
# ### Vectors (rank-1 tensors)
# + colab_type="code"
x = np.array([12, 3, 6, 14, 7])
x
# + colab_type="code"
x.ndim
# + [markdown] colab_type="text"
# ### Matrices (rank-2 tensors)
# + colab_type="code"
x = np.array([[5, 78, 2, 34, 0],
[6, 79, 3, 35, 1],
[7, 80, 4, 36, 2]])
x.ndim
# + [markdown] colab_type="text"
# ### Rank-3 tensors and higher-rank tensors
# + colab_type="code"
x = np.array([[[5, 78, 2, 34, 0],
[6, 79, 3, 35, 1],
[7, 80, 4, 36, 2]],
[[5, 78, 2, 34, 0],
[6, 79, 3, 35, 1],
[7, 80, 4, 36, 2]],
[[5, 78, 2, 34, 0],
[6, 79, 3, 35, 1],
[7, 80, 4, 36, 2]]])
x.ndim
# + [markdown] colab_type="text"
# ### Key attributes
# + colab_type="code"
from tensorflow.keras.datasets import mnist
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
# + colab_type="code"
train_images.ndim
# + colab_type="code"
train_images.shape
# + colab_type="code"
train_images.dtype
# + [markdown] colab_type="text"
# **Displaying the fourth digit**
# + colab_type="code"
import matplotlib.pyplot as plt
digit = train_images[4]
plt.imshow(digit, cmap=plt.cm.binary)
plt.show()
# + colab_type="code"
train_labels[4]
# + [markdown] colab_type="text"
# ### Manipulating tensors in NumPy
# + colab_type="code"
my_slice = train_images[10:100]
my_slice.shape
# + colab_type="code"
my_slice = train_images[10:100, :, :]
my_slice.shape
# + colab_type="code"
my_slice = train_images[10:100, 0:28, 0:28]
my_slice.shape
# + colab_type="code"
my_slice = train_images[:, 14:, 14:]
# + colab_type="code"
my_slice = train_images[:, 7:-7, 7:-7]
# + [markdown] colab_type="text"
# ### The notion of data batches
# + colab_type="code"
batch = train_images[:128]
# + colab_type="code"
batch = train_images[128:256]
# + colab_type="code"
n = 3
batch = train_images[128 * n:128 * (n + 1)]
# + [markdown] colab_type="text"
# ### Real-world examples of data tensors
# + [markdown] colab_type="text"
# ### Vector data
# + [markdown] colab_type="text"
# ### Timeseries data or sequence data
# + [markdown] colab_type="text"
# ### Image data
# + [markdown] colab_type="text"
# ### Video data
# + [markdown] colab_type="text"
# ## The gears of neural networks: tensor operations
# + [markdown] colab_type="text"
# ### Element-wise operations
# + colab_type="code"
def naive_relu(x):
assert len(x.shape) == 2
x = x.copy()
for i in range(x.shape[0]):
for j in range(x.shape[1]):
x[i, j] = max(x[i, j], 0)
return x
# + colab_type="code"
def naive_add(x, y):
assert len(x.shape) == 2
assert x.shape == y.shape
x = x.copy()
for i in range(x.shape[0]):
for j in range(x.shape[1]):
x[i, j] += y[i, j]
return x
# + colab_type="code"
import time
x = np.random.random((20, 100))
y = np.random.random((20, 100))
t0 = time.time()
for _ in range(1000):
z = x + y
z = np.maximum(z, 0.)
print("Took: {0:.2f} s".format(time.time() - t0))
# + colab_type="code"
t0 = time.time()
for _ in range(1000):
z = naive_add(x, y)
z = naive_relu(z)
print("Took: {0:.2f} s".format(time.time() - t0))
# + [markdown] colab_type="text"
# ### Broadcasting
# + colab_type="code"
import numpy as np
X = np.random.random((32, 10))
y = np.random.random((10,))
# + colab_type="code"
y = np.expand_dims(y, axis=0)
# + colab_type="code"
Y = np.concatenate([y] * 32, axis=0)
# + colab_type="code"
def naive_add_matrix_and_vector(x, y):
assert len(x.shape) == 2
assert len(y.shape) == 1
assert x.shape[1] == y.shape[0]
x = x.copy()
for i in range(x.shape[0]):
for j in range(x.shape[1]):
x[i, j] += y[j]
return x
# + colab_type="code"
import numpy as np
x = np.random.random((64, 3, 32, 10))
y = np.random.random((32, 10))
z = np.maximum(x, y)
# + [markdown] colab_type="text"
# ### Tensor product
# + colab_type="code"
x = np.random.random((32,))
y = np.random.random((32,))
z = np.dot(x, y)
# + colab_type="code"
def naive_vector_dot(x, y):
assert len(x.shape) == 1
assert len(y.shape) == 1
assert x.shape[0] == y.shape[0]
z = 0.
for i in range(x.shape[0]):
z += x[i] * y[i]
return z
# + colab_type="code"
def naive_matrix_vector_dot(x, y):
assert len(x.shape) == 2
assert len(y.shape) == 1
assert x.shape[1] == y.shape[0]
z = np.zeros(x.shape[0])
for i in range(x.shape[0]):
for j in range(x.shape[1]):
z[i] += x[i, j] * y[j]
return z
# + colab_type="code"
def naive_matrix_vector_dot(x, y):
z = np.zeros(x.shape[0])
for i in range(x.shape[0]):
z[i] = naive_vector_dot(x[i, :], y)
return z
# + colab_type="code"
def naive_matrix_dot(x, y):
assert len(x.shape) == 2
assert len(y.shape) == 2
assert x.shape[1] == y.shape[0]
z = np.zeros((x.shape[0], y.shape[1]))
for i in range(x.shape[0]):
for j in range(y.shape[1]):
row_x = x[i, :]
column_y = y[:, j]
z[i, j] = naive_vector_dot(row_x, column_y)
return z
# + [markdown] colab_type="text"
# ### Tensor reshaping
# + colab_type="code"
train_images = train_images.reshape((60000, 28 * 28))
# + colab_type="code"
x = np.array([[0., 1.],
[2., 3.],
[4., 5.]])
x.shape
# + colab_type="code"
x = x.reshape((6, 1))
x
# + colab_type="code"
x = np.zeros((300, 20))
x = np.transpose(x)
x.shape
# + [markdown] colab_type="text"
# ### Geometric interpretation of tensor operations
# + [markdown] colab_type="text"
# ### A geometric interpretation of deep learning
# + [markdown] colab_type="text"
# ## The engine of neural networks: gradient-based optimization
# + [markdown] colab_type="text"
# ### What's a derivative?
# + [markdown] colab_type="text"
# ### Derivative of a tensor operation: the gradient
# + [markdown] colab_type="text"
# ### Stochastic gradient descent
# + [markdown] colab_type="text"
# ### Chaining derivatives: the Backpropagation algorithm
# + [markdown] colab_type="text"
# #### The chain rule
# + [markdown] colab_type="text"
# #### Automatic differentiation with computation graphs
# + [markdown] colab_type="text"
# #### The Gradient Tape in TensorFlow
# + colab_type="code"
import tensorflow as tf
x = tf.Variable(0.)
with tf.GradientTape() as tape:
y = 2 * x + 3
grad_of_y_wrt_x = tape.gradient(y, x)
# + colab_type="code"
x = tf.Variable(tf.random.uniform((2, 2)))
with tf.GradientTape() as tape:
y = 2 * x + 3
grad_of_y_wrt_x = tape.gradient(y, x)
# + colab_type="code"
W = tf.Variable(tf.random.uniform((2, 2)))
b = tf.Variable(tf.zeros((2,)))
x = tf.random.uniform((2, 2))
with tf.GradientTape() as tape:
y = tf.matmul(x, W) + b
grad_of_y_wrt_W_and_b = tape.gradient(y, [W, b])
# + [markdown] colab_type="text"
# ## Looking back at our first example
# + colab_type="code"
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
train_images = train_images.reshape((60000, 28 * 28))
train_images = train_images.astype("float32") / 255
test_images = test_images.reshape((10000, 28 * 28))
test_images = test_images.astype("float32") / 255
# + colab_type="code"
model = models.Sequential([
layers.Dense(512, activation="relu"),
layers.Dense(10, activation="softmax")
])
# + colab_type="code"
model.compile(optimizer="rmsprop",
loss="sparse_categorical_crossentropy",
metrics=["accuracy"])
# + colab_type="code"
model.fit(train_images, train_labels, epochs=5, batch_size=128)
# + [markdown] colab_type="text"
# ### Reimplementing our first example from scratch in TensorFlow
# + [markdown] colab_type="text"
# #### A simple Dense class
# + colab_type="code"
import tensorflow as tf
class NaiveDense:
def __init__(self, input_size, output_size, activation):
self.activation = activation
w_shape = (input_size, output_size)
w_initial_value = tf.random.uniform(w_shape, minval=0, maxval=1e-1)
self.W = tf.Variable(w_initial_value)
b_shape = (output_size,)
b_initial_value = tf.zeros(b_shape)
self.b = tf.Variable(b_initial_value)
def __call__(self, inputs):
return self.activation(tf.matmul(inputs, self.W) + self.b)
@property
def weights(self):
return [self.W, self.b]
# + [markdown] colab_type="text"
# #### A simple Sequential class
# + colab_type="code"
class NaiveSequential:
def __init__(self, layers):
self.layers = layers
def __call__(self, inputs):
x = inputs
for layer in self.layers:
x = layer(x)
return x
@property
def weights(self):
weights = []
for layer in self.layers:
weights += layer.weights
return weights
# + colab_type="code"
model = NaiveSequential([
NaiveDense(input_size=28 * 28, output_size=512, activation=tf.nn.relu),
NaiveDense(input_size=512, output_size=10, activation=tf.nn.softmax)
])
assert len(model.weights) == 4
# + [markdown] colab_type="text"
# #### A batch generator
# + colab_type="code"
class BatchGenerator:
def __init__(self, images, labels, batch_size=128):
self.index = 0
self.images = images
self.labels = labels
self.batch_size = batch_size
def next(self):
images = self.images[self.index : self.index + self.batch_size]
labels = self.labels[self.index : self.index + self.batch_size]
self.index += self.batch_size
return images, labels
# + [markdown] colab_type="text"
# ### Running one training step
# + colab_type="code"
def one_training_step(model, images_batch, labels_batch):
with tf.GradientTape() as tape:
predictions = model(images_batch)
per_sample_losses = tf.keras.losses.sparse_categorical_crossentropy(
labels_batch, predictions)
average_loss = tf.reduce_mean(per_sample_losses)
gradients = tape.gradient(average_loss, model.weights)
update_weights(gradients, model.weights)
return average_loss
# + colab_type="code"
learning_rate = 1e-3
def update_weights(gradients, weights):
for g, w in zip(gradients, model.weights):
w.assign_sub(g * learning_rate)
# + colab_type="code"
from tensorflow.keras import optimizers
optimizer = optimizers.SGD(learning_rate=1e-3)
def update_weights(gradients, weights):
optimizer.apply_gradients(zip(gradients, weights))
# + [markdown] colab_type="text"
# ### The full training loop
# + colab_type="code"
def fit(model, images, labels, epochs, batch_size=128):
for epoch_counter in range(epochs):
print(f"Epoch {epoch_counter}")
batch_generator = BatchGenerator(images, labels)
for batch_counter in range(len(images) // batch_size):
images_batch, labels_batch = batch_generator.next()
loss = one_training_step(model, images_batch, labels_batch)
if batch_counter % 100 == 0:
print(f"loss at batch {batch_counter}: {loss:.2f}")
# + colab_type="code"
from tensorflow.keras.datasets import mnist
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
train_images = train_images.reshape((60000, 28 * 28))
train_images = train_images.astype("float32") / 255
test_images = test_images.reshape((10000, 28 * 28))
test_images = test_images.astype("float32") / 255
fit(model, train_images, train_labels, epochs=10, batch_size=128)
# + [markdown] colab_type="text"
# ### Evaluating the model
# + colab_type="code"
predictions = model(test_images)
predictions = predictions.numpy()
predicted_labels = np.argmax(predictions, axis=1)
matches = predicted_labels == test_labels
print(f"accuracy: {matches.mean():.2f}")
# + [markdown] colab_type="text"
# ## Chapter summary
| chapter02_mathematical-building-blocks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 情感分类
#
# 停用词
#
# 加载原始文本,只考虑'1', '2', '3', '4', '-'五类,'x'不确定的暂时不考虑。
# +
import os
import json
import torch
from collections import Counter
from tqdm import tqdm_notebook as tqdm
import numpy as np
from thulac import thulac
thu = thulac(user_dict='data/emo-words.txt', seg_only=True)
from myclf import *
from sklearn.neural_network import MLPClassifier
# +
def load_stopword():
"""
加载停用词集合
"""
return set(json.load(open('data/stopword-zh.json')))
# stop_word = load_stopword()
def load_label_sentence():
"""
加载原始文本
"""
sentences = []
labels = []
for line in tqdm(open("data/labelled_split/labels_text.txt")):
label = line.split('\t')[0]
s = line.split('\t')[1]
# 1234:四种情绪,-:没有情绪,x:不确定
if label in ['1', '2', '3', '4', '-']:
if label == '-':
labels.append('0')
else:
labels.append(label)
sentences.append(s)
print(Counter(labels))
return labels, sentences
labels, sentences = load_label_sentence()
for y, s in zip(labels, sentences):
print(y + ' ' + ' '.join(sentences), file=open('data/labelled/{}.txt'.format(y), 'a'))
# -
# 信息增益来计算特征词
#
# ## one-hot表示法
# +
def get_word_freq():
"""
统计高频词汇
"""
stopwords = load_stopword()
words_freq = {}
words_ci = {} # 出现某个词,是某类的概率,此问题有五类
class_num = 5
labels_num = [0] * class_num
labels, sentences = load_label_sentence()
for y, s in zip(labels, sentences):
# 统计每个类别的数量
labels_num[int(y)] += 1
# 分词
for w in thu.cut(s):
w = w[0]
# 停用词等过滤
if w == '' or w in stopwords or w.isdigit():
continue
elif w in words_freq:
words_freq[w] += 1
words_ci[w][int(y)] += 1
else:
words_freq[w] = 1
words_ci[w] = [0] * class_num
words_ci[w][int(y)] += 1
# 数量转概率
num2pro = lambda nums: [num / sum(nums) for num in nums]
# 每类上的概率
v_ci = num2pro(labels_num)
word_gain = {}
for w in words_ci.keys():
word_ci = words_ci[w]
v_ci_t = num2pro(word_ci) # 句子出现t是Ci类的概率
non_word_ci = [labels_num[i] - word_ci[i] for i in range(class_num)] # 不是t时候的各类数量
v_ci_non_t = num2pro(non_word_ci) # 句子不出现t是Ci的概率
pr_t = words_freq[w] / sum(labels_num) # 存在t的概率
Gt = Info_gain_of_term(v_ci, v_ci_t, v_ci_non_t, pr_t)
word_gain[w] = Gt
word_gain = sorted(word_gain.items(), key=lambda d: d[1], reverse=True)
with open('data/word_gain_freq.txt', 'w') as f:
for w, gain in word_gain:
if words_freq[w] >= 5:
print(w, gain, words_freq[w], sep='\t', file=f)
def Info_gain_of_term(v_ci, v_ci_t, v_ci_non_t, pr_t):
"""
计算信息增益,需要每类的概率,句子出现t是Ci类的概率,不出现t是Ci的概率,存在t的概率
"""
def info_entropy(p):
if p == 0:
return 0
else:
return -p * np.log(p)
gain = 0
for i in range(len(v_ci)):
gain = gain + (info_entropy(v_ci[i]) - pr_t * info_entropy(v_ci_t[i]) - (1 - pr_t) * info_entropy(v_ci_non_t[i]))
return gain
# +
def load_word_list(first=2400):
word_list = []
for i, line in enumerate(open('data/word_gain_freq.txt')):
if i >= first:
break
try:
w, gain, freq = line.strip().split('\t')
except ValueError:
print('读取词向量出错:行 {}'.format(i))
word_list.append(w)
print('词向量大小', len(word_list))
return word_list
def make_features_onehot(features_file_name):
word_list = load_word_list()
print('---- 我的词表 ----')
i = 0
with open(features_file_name, 'w') as f:
for y, s in zip(labels, sentences):
i += 1
if not i % 1000:
print('行 ->', i)
vec = np.zeros(len(word_list))
for w in thu.cut(s):
w = w[0]
# print(w)
try:
_i = word_list.index(w)
vec[_i] = 1
except ValueError:
pass
f.write(y + '\t' + ','.join(['{:.1f}'.format(num) for num in list(vec)]) + '\n')
print('总行数:', i)
# -
# 引入ACL2018词向量(财经方面)
#
# 因为该文件是按出现次数排序,那么考虑“掐头去尾”
#
# 停用词要不要去?也是要考虑的,停用词有时候也起到作用。
# +
def load_word_vec():
"""
加载ACL2018词向量
"""
word_vec = {}
print('加载词向量中 ...')
for i, line in enumerate(open('data/sgns.financial.word')):
if i <= 10:
continue
if i > 150000:
break
words = line.strip().split(' ')
word = words[0]
word_vec[word] = np.array([float(num) for num in words[1:]])
# except UnicodeDecodeError:
# print("编码问题,行 {}".format(i))
print('加载词完成!一共 {}个词'.format(len(word_vec)))
return word_vec
def make_features_ACLwv(features_file_name):
word_vec = load_word_vec()
i = 0
# 建立训练文件:ACL的wv
print('---- ACL wv ----')
with open(features_file_name, 'w') as f:
for y, s in zip(labels, sentences):
i += 1
if not i % 1000:
print('行 -> {}'.format(i))
count = 0
vec = np.zeros(300)
for w in thu.cut(s): # 对分词结果进行处理
w = w[0]
# if w in stop_word:
# continue
if w in word_vec:
vec += word_vec[w]
count += 1
if count != 0:
vec = vec / count
# if count > 0:
f.write(y + '\t' + ','.join(['{:.6f}'.format(num) for num in list(vec)]) + '\n')
print('总行数:', i)
# +
from gensim.models import Word2Vec
def make_features_mywv(features_file_name):
mywv_model = Word2Vec.load("model/guba_word2vec.model")
i = 0
# 建立训练文件: 我的wv
print('---- 我的wv ----')
with open(features_file_name, 'w') as f:
for y, s in zip(labels, sentences):
i += 1
if not i % 1000:
print('行 -> {}'.format(i))
count = 0
vec = np.zeros(300)
for w in thu.cut(s): # 对分词结果进行处理
w = w[0]
if w in mywv_model.wv:
vec += mywv_model.wv[w]
count += 1
if count != 0:
vec = vec / count
# if count > 0:
f.write(y + '\t' + ','.join(['{:.6f}'.format(num) for num in list(vec)]) + '\n')
print('总行数:', i)
# -
get_word_freq() # 词分析
make_features_onehot('data/train/onehot.txt')
make_features_ACLwv('data/train/ACLwv.txt')
make_features_mywv('data/train/mywv.txt')
ig_word_list = load_word_list()
ACL_word_vec = load_word_vec()
mywv_model = Word2Vec.load("model/guba_word2vec.model")
# +
def my_w2v(w):
# 三合一
vec = np.zeros(3000)
if w in ig_word_list:
vec[ig_word_list.index(w)] = 1
if w in ACL_word_vec:
vec[2400: 2700] = ACL_word_vec[w]
if w in mywv_model.wv:
vec[2700:] = mywv_model.wv[w]
return vec
# ' '.join([str(i) for i in list(my_w2v('涨停'))])
# -
# ## ~ ⬆️准备训练数据 ⬇️开始训练
#
# 机器学习算法包括:KNN、LR、随机森林、决策树、GBDT、SVM
# +
def load_train_data(in_name, num=5):
"""
加载训练数据
"""
X = []
y = []
for line in open(in_name):
lab, vec = line.strip().split('\t')
if num == 0: # 是否有情绪
if lab != '0':
lab = '1'
elif num == 2: # 正负情绪
if lab == '2':
lab = '1'
elif lab == '0':
continue
else:
lab = '0'
elif num == 4: # 四种情绪
if lab == '0':
continue
x = np.array([float(v) for v in vec.split(',')])
X.append(x)
y.append(int(lab))
X = np.array(X)
y = np.array(y)
return X, y
def stack_X_y(X1, y1, X2, y2, out_name=0):
print(X1.shape, y1.shape, X2.shape, y2.shape)
if len(y1) != len(y2):
print('两列表长度不同,不同合并。')
return -1
_len = len(X1)
X = []
for i in range(_len):
xi= np.hstack([X1[i], X2[i]])
X.append(xi)
X = np.array(X)
y = np.array(y1)
if out_name != 0:
with open(out_name, 'w') as f:
for xi, yi in zip(X, y):
f.write(str(yi) + '\t' + ','.join(['{:.6f}'.format(num) for num in list(xi)]) + '\n')
print('合并数据完成。')
return X, y
# +
def train():
"""
调参
"""
# 合并数据
# X1, y1 = load_train_data('data/train/onehot.txt')
# X2, y2 = load_train_data('data/train/ACLwv.txt')
# X1, y1 = stack_X_y(X1, y1, X2, y2)
# X3, y3 = load_train_data('data/train/mywv.txt')
# X, y = stack_X_y(X1, y1, X3, y3, out_name='data/train/all-180920.txt')
X, y = load_train_data('data/train/all-180920.txt', num=4)
# from sklearn.datasets import dump_svmlight_file
# dump_svmlight_file(X, y, 'data/train/svmlib-180920-C4.txt')
# print('saving svm format ...')
print(X.shape, y.shape)
# 划分数据集
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=21)
# 初始化分类器
test_classifiers = ['LR', 'GBDT']
classifiers = {
'NB':naive_bayes_classifier,
'KNN':knn_classifier,
'LR':logistic_regression_classifier,
'RF':random_forest_classifier,
'DT':decision_tree_classifier,
'SVM':svm_classifier,
'SVMCV':svm_cross_validation,
'GBDT':gradient_boosting_classifier
}
## NN
print('****** NN ******')
clf = MLPClassifier((128, 128), solver='adam', alpha=0.1)
clf.fit(X_train, y_train)
evaluate(clf, X, y, X_test, y_test)
for classifier in test_classifiers:
print('******************* {} ********************'.format(classifier))
clf = classifiers[classifier](X_train, y_train)
evaluate(clf, X, y, X_test, y_test)
# SVC
original_params = {}
for i, setting in enumerate([{'C':0.125}, {'C': 0.25}, {'C':0.5}, {'C':1.0}]):
print('******************* SVC-{} ********************'.format(i))
print(setting)
params = dict(original_params)
params.update(setting)
clf = LinearSVC(**params)
clf.fit(X_train, y_train)
evaluate(clf, X, y, X_test, y_test)
# GBDT
# original_params = {'n_estimators': 1000, 'max_leaf_nodes': 4, 'max_depth': 3, 'random_state': 23,
# 'min_samples_split': 5}
# for i, setting in enumerate([{'learning_rate': 1.0, 'subsample': 1.0},
# {'learning_rate': 0.1, 'subsample': 1.0},
# {'learning_rate': 1.0, 'subsample': 0.5},
# {'learning_rate': 0.1, 'subsample': 0.5},
# {'learning_rate': 0.1, 'max_features': 2}]):
# print('******************* GBDT-{} ********************'.format(i))
# print(setting)
# params = dict(original_params)
# params.update(setting)
# clf = GradientBoostingClassifier(**params)
# clf.fit(X_train, y_train)
# evaluate(clf, X, y, X_test, y_test)
def evaluate(clf, X, y, X_test, y_test):
# CV
print('accuracy of CV:', cross_val_score(clf, X, y, cv=5).mean())
# 模型评估
y_pred = []
for i in range(len(X_test)):
y_hat = clf.predict(X_test[i].reshape(1, -1))
y_pred.append(y_hat[0])
print(classification_report(y_test, y_pred))
def train_model():
X, y = load_train_data('data/train/train_data_ACL-20180712.txt')
clf = LogisticRegression(penalty='l2')
print(X.shape, y.shape)
clf.fit(X, y)
# 保存模型
joblib.dump(clf, "emo-LR-v1.model")
train()
# -
| emotion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import packages
# from __future__ import print_function
import voila
import watermark
import pandas as pd
import numpy as np
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
import plotly.express as px
import plotly.graph_objects as go
# -
# Original Data from https://github.com/CSSEGISandData/COVID-19
# Copyright by: "COVID-19 Data Repository by the Center for Systems Science and Engineering (CSSE) at Johns Hopkins University"
glbconfirmed_df = pd.read_csv("https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_confirmed_global.csv")
glbrecovered_df = pd.read_csv("https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_recovered_global.csv")
glbdeath_df = pd.read_csv("https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_deaths_global.csv")
country_df = pd.read_csv("https://raw.githubusercontent.com/CSSEGISandData/COVID-19/web-data/data/cases_country.csv")
# Data cleaning (renaming columns to lower-case)
country_df.columns = map(str.lower, country_df.columns)
glbrecovered_df.columns = map(str.lower, glbrecovered_df.columns)
glbdeath_df.columns = map(str.lower, glbdeath_df.columns)
glbconfirmed_df.columns = map(str.lower, glbconfirmed_df.columns)
# Before changing column names save country_df as scatter_df for scatter plot
scatter_df = country_df.sort_values('confirmed', ascending=False)
# Change of 'province/state' to 'state' and 'country/region' to 'country'
glbconfirmed_df = glbconfirmed_df.rename(columns = {'province/state': 'state', 'country/region': 'country'})
glbrecovered_df = glbrecovered_df.rename(columns = {'province/state': 'state', 'country/region': 'country'})
glbdeath_df = glbdeath_df.rename(columns = {'province/state': 'state', 'country/region': 'country'})
country_df = country_df.rename(columns = {'country_region': 'country'})
# Change automatic float-datatype to int
country_df["confirmed"] = country_df["confirmed"].astype('Int32')
country_df["deaths"] = country_df["deaths"].astype('Int64')
country_df["active"] = country_df["active"].astype('Int64')
country_df["recovered"] = country_df["recovered"].astype('Int64')
# Set sorted_country_df for list-display
sorted_country_df = country_df.sort_values('confirmed', ascending=False)
sorted_country_df.drop(columns=['lat', 'long_', 'people_tested', 'people_hospitalized'], inplace = True)
# Define the 'higlight_col'-function to colorize certain columns
def highlight_col(x):
p = 'background-color: grey'
r = 'background-color: brown'
g = 'background-color: darkgreen'
temp_df = pd.DataFrame('', index = x.index, columns = x.columns)
temp_df.iloc[:, 2] = p
temp_df.iloc[:, 3] = r
temp_df.iloc[:, 4] = g
return temp_df
# Define the 'plot_cases_for_country'-function for graph-display
# Choose an input Country, iterate through glb_confirmed_df and glbdeath_df
# to display the countries total confirmed and death incidents using plotly.graphic_objects
def plot_cases_for_country(country):
labels = ["confirmed cases", "deaths"]
colors = ['#4C78A4', '#D62728']
mode_size = [6, 8]
line_size = [5, 4]
df_list = [glbconfirmed_df, glbdeath_df]
fig = go.Figure(layout=go.Layout(title='<span style="font-size: 24px;">Timeline</span>'))
if country in np.asarray(glbconfirmed_df['country']) or str.casefold(country) == 'world':
for i, df in enumerate(df_list):
if country == 'World' or country == 'world':
x_data = np.array(list(df.iloc[:, 5:].columns))
y_data = np.sum(np.asarray(df.iloc[:, 5:]), axis = 0)
else:
x_data = np.array(list(df.iloc[:, 5:].columns))
y_data = np.sum(np.asarray(df[df['country'] == country].iloc[:, 5:]), axis = 0)
fig.add_trace(go.Scatter(x = x_data, y = y_data, mode = 'lines',
name = labels[i],
line = dict(color = colors[i], width = line_size[i]),
connectgaps = True,
text = "Total" + str(labels[i]) + ": " + str(y_data[-1])
))
else:
print("Please check the extended list below for correct spelling.")
if country != 'World':
display(sorted_country_df.loc[sorted_country_df["country"] == country].style.apply(highlight_col, axis = None))
fig.show()
# # Covid-19 Dashboard
# This Dashboard offers an overview of the worldwide development of Covid-19.
# The data was distributed by John Hopkins University. For further information please visit: https://coronavirus.jhu.edu/about
# Graph-Display: Using interact to dynamically choose a country for its confirmed cases and deaths
interact(plot_cases_for_country, country = 'World');
# +
# scatter_df has different column names! plotting did not accept changed name-datatypes - is not displayed by voila
plt_scat = px.scatter(scatter_df.head(15), x='country_region', y='confirmed', size='confirmed', color='country_region', hover_name="country_region", size_max = 60)
plt_scat.update_layout(title_text = "Comparison of Confirmed Cases", title_font_size= 24, height=550, width=1000)
f = go.FigureWidget(data = plt_scat)
display(f)
# -
# ## Table of Worldwide Cases
# A list of all countries:
sorted_country_df.style.apply(highlight_col, axis = None)
#
# Please feel free to contact me for further enquiries or feedback. Link to the GitHub repository: https://github.com/JayVeezy1/CS50_final_project
#
#
# The dashboard was inspired by the following tutorial: https://www.youtube.com/watch?v=FngV4VdYrkA&list=LL&index=2&ab_channel=DataSciencewithHarshit
# This data set is licensed under the Creative Commons Attribution 4.0 International (CC BY 4.0) by the Johns Hopkins University on behalf of its Center for Systems Science in Engineering. Copyright Johns Hopkins University 2020.
# The correctness of this data can not by guaranteed.
| notebooks/covid19_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,py
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:generic_expression] *
# language: python
# name: conda-env-generic_expression-py
# ---
# # Process _P. aeruginosa_ multiplier model
# +
# %load_ext autoreload
# %autoreload 2
import os
import pandas as pd
import pickle
import rpy2.robjects as ro
from rpy2.robjects import pandas2ri
from rpy2.robjects.conversion import localconverter
# -
readRDS = ro.r["readRDS"]
# CHANGE LOCATION TO LOCAL WHEN COMMIT
multiplier_full_model = readRDS("Pa_compendium_PLIER_model.RDS")
# # Format multiplier Z data
#
# The Z data matrix contains the contribution (i.e. weight) per gene to each latent variable
multiplier_model_Z_matrix = multiplier_full_model.rx2("Z")
with localconverter(ro.default_converter + pandas2ri.converter):
multiplier_model_Z_matrix_values = ro.conversion.rpy2py(multiplier_model_Z_matrix)
# +
column_header = [f"LV{i}" for i in range(1, 73)]
multiplier_model_Z_matrix_df = pd.DataFrame(
data=multiplier_model_Z_matrix_values,
index=multiplier_model_Z_matrix.rownames,
columns=column_header,
)
# -
print(multiplier_model_Z_matrix_df.shape)
multiplier_model_Z_matrix_df.head()
# Save
multiplier_model_Z_matrix_df.to_csv("multiplier_Pa_model_z.tsv", sep="\t")
# # Format multiplier summary data
#
# This summary data matrix contains statistics about each LV - which pathways it was associated with and its significance score. This information is saved in the MultiPLIER model: https://github.com/greenelab/multi-plier/blob/7f4745847b45edf8fef3a49893843d9d40c258cf/23-explore_AAV_recount_LVs.Rmd
multiplier_model_matrix = multiplier_full_model.rx2("summary")
with localconverter(ro.default_converter + pandas2ri.converter):
multiplier_model_matrix_values = ro.conversion.rpy2py(multiplier_model_matrix)
multiplier_model_matrix_df = pd.DataFrame(
data=multiplier_model_matrix_values,
index=multiplier_model_matrix.rownames,
columns=multiplier_model_matrix.colnames,
)
multiplier_model_matrix_df.head()
# Save
multiplier_model_matrix_df.to_csv("multiplier_Pa_model_summary.tsv", sep="\t")
| LV_analysis/0_process_Pa_multiplier_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from multiprocessing import Pool, cpu_count
import copy
import cv2
# -
dfAll = pd.read_pickle("DataStudyEvaluation/AllData.pkl")
df = dfAll[(dfAll.Actual_Data == True) & (dfAll.Is_Pause == False)]
df.head()
df.userID.unique()
# +
# %%time
def is_max(df):
df_temp = df.copy(deep=True)
max_version = df_temp.RepetitionID.max()
df_temp["IsMax"] = np.where(df_temp.RepetitionID == max_version, True, False)
df_temp["MaxRepetition"] = [max_version] * len(df_temp)
return df_temp
df_grp = df.groupby([df.userID, df.TaskID, df.VersionID])
pool = Pool(cpu_count() - 1)
result_lst = pool.map(is_max, [grp for name, grp in df_grp])
df = pd.concat(result_lst)
pool.close()
# -
df.Image = df.Image.apply(lambda x: x.reshape(27, 15))
df.Image = df.Image.apply(lambda x: x.clip(min=0, max=255))
df.Image = df.Image.apply(lambda x: x.astype(np.uint8))
df["ImageSum"] = df.Image.apply(lambda x: np.sum(x))
df.to_pickle("DataStudyEvaluation/dfFiltered.pkl")
print("recorded actual: %s, used data: %s" % (len(dfAll), len(df)))
df = pd.read_pickle("DataStudyEvaluation/dfFiltered.pkl")
df.head()
# +
#Label if knuckle or finger
def f(row):
if row['TaskID'] < 17:
#val = "Knuckle"
val = 0
elif row['TaskID'] >= 17:
#val = "Finger"
val = 1
return val
df['InputMethod'] = df.apply(f, axis=1)
def f(row):
if row['TaskID'] < 17:
val = "Knuckle"
elif row['TaskID'] >= 17:
val = "Finger"
return val
df['Input'] = df.apply(f, axis=1)
# -
#Svens new Blob detection
def detect_blobs(image, task):
#image = e.Image
large = np.ones((29,17), dtype=np.uint8)
large[1:28,1:16] = np.copy(image)
temp, thresh = cv2.threshold(cv2.bitwise_not(large), 200, 255, cv2.THRESH_BINARY)
contours, hierarchy = cv2.findContours(thresh, cv2.RETR_TREE, cv2.CHAIN_APPROX_SIMPLE)
contours = [a for a in contours if cv2.contourArea(a) > 8 and cv2.contourArea(a) < 255]
lstBlob = []
lstMin = []
lstMax = []
count = 0
contours.sort(key=lambda a: cv2.contourArea(a))
if len(contours) > 0:
# if two finger or knuckle
cont_count = 2 if task in [1, 6, 7, 18, 23, 24] and len(contours) > 1 else 1
for i in range(1, cont_count + 1):
max_contour = contours[-1 * i]
xmax, ymax = np.max(max_contour.reshape(len(max_contour),2), axis=0)
xmin, ymin = np.min(max_contour.reshape(len(max_contour),2), axis=0)
#croped_im = np.zeros((27,15))
blob = large[max(ymin - 1, 0):min(ymax + 1, large.shape[0]),max(xmin - 1, 0):min(xmax + 1, large.shape[1])]
#croped_im[0:blob.shape[0],0:blob.shape[1]] = blob
#return (1, [croped_im])
lstBlob.append(blob)
lstMin.append(xmax-xmin)
lstMax.append(ymax-ymin)
count = count + 1
return (count, lstBlob, lstMin, lstMax)
else:
return (0, [np.zeros((29, 19))], 0, 0)
# %%time
pool = Pool(os.cpu_count()-1)
temp_blobs = pool.starmap(detect_blobs, zip(df.Image, df.TaskID))
pool.close()
df["BlobCount"] = [a[0] for a in temp_blobs]
df["BlobImages"] = [a[1] for a in temp_blobs]
df["BlobW"] = [a[2] for a in temp_blobs]
df["BlobH"] = [a[3] for a in temp_blobs]
df.BlobCount.value_counts()
# +
dfX = df[(df.BlobCount == 1)].copy(deep=True)
dfX.BlobImages = dfX.BlobImages.apply(lambda x : x[0])
dfX.BlobW = dfX.BlobW.apply(lambda x : x[0])
dfX.BlobH = dfX.BlobH.apply(lambda x : x[0])
dfY = df[(df.BlobCount == 2)].copy(deep=True)
dfY.BlobImages = dfY.BlobImages.apply(lambda x : x[0])
dfY.BlobW = dfY.BlobW.apply(lambda x : x[0])
dfY.BlobH = dfY.BlobH.apply(lambda x : x[0])
dfZ = df[(df.BlobCount == 2)].copy(deep=True)
dfZ.BlobImages = dfZ.BlobImages.apply(lambda x : x[1])
dfZ.BlobW = dfZ.BlobW.apply(lambda x : x[1])
dfZ.BlobH = dfZ.BlobH.apply(lambda x : x[1])
df = dfX.append([dfY, dfZ])
# -
print("Sample Size not Argumented:", len(df))
df["BlobArea"] = df["BlobW"] * df["BlobH"]
df.BlobArea.describe().round(1)
df.groupby("Input").BlobArea.describe().round(1)
df["BlobSum"] = df.BlobImages.apply(lambda x: np.sum(x))
df.BlobSum.describe()
df.BlobSum.hist()
#Small / Blobs where the pixels are only a "little" hit
dfX = df[df.BlobSum <= 255]
len(dfX)
print("Sample Size argumented:", len(df))
def pasteToEmpty (blob):
croped_im = np.zeros((27,15))
croped_im[0:blob.shape[0],0:blob.shape[1]] = blob
return croped_im
df["Blobs"] = df.BlobImages.apply(lambda x: pasteToEmpty(x))
df.to_pickle("DataStudyEvaluation/df_statistics.pkl")
df[["userID", "TaskID", "Blobs", "InputMethod"]].to_pickle("DataStudyEvaluation/df_blobs_area.pkl")
# # display blobs
plt.clf()
plt.figure(figsize=(6, 6))
ax = plt.gca()
data_point = 100
data = df.Blobs.iloc[data_point]
print(df.iloc[data_point])
plt.imshow(data, cmap='gray', vmin=0, vmax=255)
# Loop over data dimensions and create text annotations.
for i in range(0, data.shape[0]):
for j in range(0, data.shape[1]):
text = ax.text(j, i, int(data[i, j]),
ha="center", va="center", color="cyan", fontsize=1)
plt.show()
| python/Step_33_CNN_PreprocessData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:mu2e_utils]
# language: python
# name: conda-env-mu2e_utils-py
# ---
# # Working Through "Asymptotic formulae for likelihood-based tests of new physics" as Laid Out by Cowan, Cranmer, Gross, and Vitells
# - <NAME>
# - 02-01-2021
# - Paper: https://arxiv.org/abs/1007.1727
# # Imports
# +
# import time
# from copy import deepcopy
import numpy as np
# import pandas as pd
from scipy.stats import norm, poisson, chi2, ncx2
# from tqdm.notebook import tqdm
# from joblib import Parallel, delayed
# import multiprocessing
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
# %matplotlib inline
#from matplotlib.ticker import FormatStrFormatter, StrMethodFormatter, FuncFormatter, MultipleLocator
#import matplotlib.colors as mcolors
from plot_config import config_plots
# -
config_plots()
plt.rcParams.update({'font.size': 18.0})
plt.rc('text', usetex=True)
# # Scratch
# + [markdown] heading_collapsed=true
# ## Section 2: Formalism of a search as a statistical test
# + [markdown] heading_collapsed=true hidden=true
# ### Using median instead of expectation
# - Paragraph 5 states that sensitivity to discovery given some signal process $H_1$ can be characterized by expectation value, under assumption of $H_1$, of the value of $Z$ obtained from a test of $H_0$, but that Eqn. 1 (which uses the $p$ value of a particular measurement) does not hold as "$Z$ obtained from Eqn. 1 using expectation of $p$-value". The stated reason is $p$ and $Z$ have a non-linear relationship, but that Eq. 1 will hold for **median** $Z$ and $p$, since the relationship is monotonic.
# - Verify these statements.
# + hidden=true
def Z(p):
return norm.ppf(1-p, loc=0, scale=1)
# + hidden=true
# check 5 sigma and p=0.05
Z(2.87e-7), Z(0.05)
# + hidden=true
ps = np.linspace(1e-8, 1, 1000)
Zs = Z(ps)
# + hidden=true
Zs
# + hidden=true
plt.plot(ps, Zs)
# + [markdown] hidden=true
# - Suppose expected number of events under $H_1$ is 100 and expected number of events under $H_0$ is 50
# - Let's see what we get for sensitivity both by using "expectation" and "median"
# + hidden=true
N = 100000 # number of experiments
mu1 = 100
mu0 = 75
# + hidden=true
#ns = np.random.normal(loc=0, scale=1, size=N)
ns = np.random.poisson(lam=mu1, size=N)
# + hidden=true
ns
# + hidden=true
p_vals = poisson.sf(k=ns, mu=mu0)
# + hidden=true
np.mean(p_vals)
# + hidden=true
Z_vals = Z(p_vals)
# + hidden=true
Z_vals
# + hidden=true
np.sum(np.isinf(Z_vals))
# + hidden=true
np.mean(Z_vals)
# + hidden=true
Z(np.mean(p_vals))
# + hidden=true
plt.hist(Z_vals, bins=100, histtype='step');
# + hidden=true
plt.hist(p_vals, bins=100, histtype='step');
# + hidden=true
np.median(p_vals), np.median(Z_vals)
# + hidden=true
Z(np.median(p_vals))
# + [markdown] hidden=true
# - My test agrees with the paper's statement. Would like to think more and make sure mathematical reason is clear.
# + [markdown] heading_collapsed=true
# ## Section 3:
# + [markdown] hidden=true
# #### Checking Approximated Distribution
# + hidden=true
def t_mu(mu, mu_hat, sigma):
return (mu - mu_hat)**2 / sigma**2
# + hidden=true
lam = (mu-mu_prime)**2 / sigma**2
def f_t_mu(tmu, mu, muprime, sigma):
lam = (mu-muprime)**2/sigma**2
f = 1/(2*tmu**(1/2)) * 1 / (2*np.pi)**(1/2) *\
(np.exp(-1/2 * (tmu**(1/2) + lam**(1/2))**2) + np.exp(-1/2 * (tmu**(1/2) - lam**(1/2))**2))
return f
# def f_t_mu(tmu, mu, muprime, sigma):
# lam = (mu-muprime)**2/sigma**2
# f = 1/(2*tmu**(1/2)) * 1 / (2*np.pi)**(1/2) *\
# (np.exp(-1/2 * (tmu**(1/2) - lam**(1/2))**2))
# return f
# + hidden=true
# generate mu_hat, mean mu_prime, std sigma
N = 1000000
mu_prime = 3.
sigma= 1.
mu_hats = np.random.normal(loc=mu_prime, scale=sigma, size=N)
mus = 10 # 5
#mus = np.random.uniform(low=0, high=10, size=N)
# + hidden=true
t_mus = t_mu(mus, mu_hats, sigma)
# + hidden=true
# suppose mu fixed
#mu = 5
tmus = np.linspace(1e-2, 80, 81)
#mus_ = tmus**(1/2)*sigma + mu
fs = f_t_mu(tmus, mus, mu_prime, sigma)
# + hidden=true
#fs
# + hidden=true
# + hidden=true
# + hidden=true
# + hidden=true
print(f'mean: {np.mean(mu_hats):0.2f}, std: {np.std(mu_hats):0.2f}')
plt.hist(mu_hats, bins=100, histtype='step');
plt.xlabel(r'$\hat{\mu}$');
# + hidden=true
print(f'mean: {np.mean(t_mus):0.2f}, std: {np.std(t_mus):0.2f}')
plt.hist(t_mus, bins=100, histtype='step');
plt.xlabel(r'$\hat{\mu}$');
# + hidden=true
print(f'mean: {np.mean(t_mus):0.2f}, std: {np.std(t_mus):0.2f}')
plt.hist(t_mus, bins=100, histtype='step', density=1);
plt.plot(tmus, fs, 'r--')
plt.xlabel(r'$t_\mu$')
plt.ylabel(r'$f(t_\mu)$');
# + [markdown] hidden=true
# - This looks to be true for a few test cases.
# - Still not sure why there are two exponential terms instead of one.
# + hidden=true
# using scipy for non-central chi2, 1 DOF
fs2 = ncx2.pdf(tmus, df=1, nc=(mus-mu_prime)**2/sigma**2)
# + hidden=true
tmus
# + hidden=true
fs
# + hidden=true
fs2
# + hidden=true
all(np.isclose(fs, fs2))
# + hidden=true
mu_hats
# + hidden=true
muhs = np.linspace(-20, 20, 401)
tmus_ = (mus-muhs)**2/sigma**2
# + hidden=true
plt.plot(muhs, tmus_)
# + hidden=true
mus
# + [markdown] hidden=true
# - Note: going from $t_\mu$ to $f(t_\mu)$ is done by using theorems surrounding functions of continuous random variables where the function is not monotone (e.g. https://www.probabilitycourse.com/chapter4/4_1_3_functions_continuous_var.php)
# -
# # Examples
# ## Counting Experiment
# ### Asymptotic Approx.
# +
# likelihood is product of two poissonion variables
# drop factorials -- they cancel in likelihood ratio
def L(mu, b, s, n, m, tau):
return (mu*s+b)**n * np.exp(-(mu*s+b)) * (tau*b)**m * np.exp(-(tau*b))
# estimator functions
def muhat(n, m, tau, s):
return (n-m/tau)/s
def bhat(m, tau):
return m/tau
def bhat2(n, m, tau, s, mu):
return (n+m - (1+tau)*mu*s) / (2*(1+tau)) + (((n+m-(1+tau)*mu*s)**2 + 4*(1+tau)*m*mu*s)/(4*(1+tau)**2))**(1/2)
# approximation for PDF
# does not handle q0 = 0 --> delta
def f_q0(q0, muprime, sigma):
return 1 / (2 *(2*np.pi*q0)**(1/2)) * np.exp(-1/2 * (q0**(1/2) - muprime/sigma)**2)
# -
# find sigma from Asimov dataset
def q0_asimov(muprime, s, b, tau):
nA = muprime * s + b
mA = tau * b
bh2 = bhat2(nA, mA, tau, s, 0) # assuming mu = 0 for q0
bh = bhat(mA, tau)
muh = muhat(nA, mA, tau, s)
# calculate appropriate likelihood ratio
q0A = -2 * np.log(L(0, bh2, s, nA, mA, tau)/L(muh, bh, s, nA, mA, tau))
return q0A
q0_asimov(1, 10, 10, 1)
def sigma_q0A(q0A, muprime):
return muprime/q0A**(1/2)
sigma = sigma_q0A(q0_asimov(1, 10, 10, 1), 1)
sigma
#q0s_asym = np.linspace(1e-2, 40, 4000)
q0s_asym = np.linspace(1e-1, 40, 400)
fq0_0 = f_q0(q0s_asym, 0, 1) # sigma=1 is a kludge
fq0_1 = f_q0(q0s_asym, 1, sigma)
## DIGITIZED FIG 3a
qs = np.array([0.1982,1.9273,3.5484,5.4463,7.2058,9.964,12.3071,14.7368,
18.1953, 21.7447, 24.879, 28.2986, 30.1144])
fs = np.array([.387, .05402, .017712, .005567, .002001, .0004389,
.00012064, 3.232e-5, 5.221e-6, 8.141e-7, 1.58e-7,
2.6318e-8, 1.0516e-8])
# +
fig, ax = plt.subplots()
ax.xaxis.set_major_locator(ticker.MultipleLocator(base=5.))
ax.xaxis.set_minor_locator(ticker.MultipleLocator(base=1.))
ax.xaxis.set_ticks_position('both')
ax.set_yscale('log')
ax.yaxis.set_major_locator(ticker.LogLocator(base=10., numticks=12))
ax.yaxis.set_minor_locator(ticker.LogLocator(base=10.,subs=(0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9), numticks=12))
ax.yaxis.set_ticks_position('both')
ax.plot(q0s_asym, fq0_0, 'k--', label=r'$f(q_0|0)_{\mathrm{asymp.}}$ [Cole]')
ax.scatter(qs, fs, s=10, c='red', label=r'$f(q_0|0)_{\mathrm{asymp.}}$ [paper]')
ax.set_xlim([0,40])
ax.set_ylim([1e-8,10])
ax.set_xlabel(r'$q_0$')
ax.set_ylabel(r'$f(q_0|0)$')
ax.legend()
# -
# - Looks good.
# ### Monte Carlo
# +
# # likelihood is product of two poissonion variables
# # drop factorials -- they cancel in likelihood ratio
# def L(mu, b, s, n, m, tau):
# return (mu*s+b)**n * np.exp(-(mu*s+b)) * (tau*b)**m * np.exp(-(tau*b))
# # estimator functions
# def muhat(n, m, tau, s):
# return (n-m/tau)/s
# def bhat(m, tau):
# return m/tau
# def bhat2(n, m, tau, s, mu):
# return (n+m - (1+tau)*mu*s) / (2*(1+tau)) + (((n+m-(1+tau)*mu*s)**2 + 4*(1+tau)*m*mu*s)/(4*(1+tau)**2))**(1/2)
# -
def gen_q0s(mu, s, b, tau, N):
# generate n and m bin values for N experiments
ns = np.random.poisson(lam=mu*s+b, size=N)
ms = np.random.poisson(lam=tau*b, size=N)
# calculate ML estimators
bh2s = bhat2(ns, ms, tau, s, 0) # assuming mu = 0 for q0
bhs = bhat(ms, tau)
muhs = muhat(ns, ms, tau, s)
# calculate appropriate likelihood ratio
q0s = np.zeros(N)
#q0s[muhs<0] = 0
c = muhs>=0
q0s[c] = -2 * np.log(L(0, bh2s[c], s, ns[c], ms[c], tau)/L(muhs[c], bhs[c], s, ns[c], ms[c], tau))
# calculate appropriate likelihood ratio
#q0s = -2 * np.log(L(0, bh2s, s, ns, ms, tau)/L(muhs, bhs, s, ns, ms, tau))
return q0s
# - Figure 3a
N = 10000000
mu=0; s=10; tau=1
#bs = [0.5, 1, 2, 5, 20]
bs = [2, 5, 20]
q0s_list = []
for b in bs:
q0s_list.append(gen_q0s(mu, s, b, tau, N))
# +
fig, ax = plt.subplots()
ax.xaxis.set_major_locator(ticker.MultipleLocator(base=5.))
ax.xaxis.set_minor_locator(ticker.MultipleLocator(base=1.))
ax.xaxis.set_ticks_position('both')
ax.set_yscale('log')
ax.yaxis.set_major_locator(ticker.LogLocator(base=10., numticks=12))
ax.yaxis.set_minor_locator(ticker.LogLocator(base=10.,subs=(0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9), numticks=12))
ax.yaxis.set_ticks_position('both')
ax.plot(q0s_asym, fq0_0, 'k--', label=r'$f(q_0|0)_{\mathrm{asymp.}}$')
for b, q in zip(bs, q0s_list):
ax.hist(q, bins=np.linspace(0,40,23), linewidth=2, histtype="step", density=1, label=f'b={b}')
ax.set_xlim([0,40])
ax.set_ylim([1e-8,10])
ax.set_xlabel(r'$q_0$')
ax.set_ylabel(r'$f(q_0|0)$')
ax.legend()
# -
# - Figure 3b
N = 10000000
s=10; b=10; tau=1 # paper
#s=7; b=0.5; tau=1 # Mu2e
q0s_0 = gen_q0s(0, s, b, tau, N)
q0s_1 = gen_q0s(1, s, b, tau, N)
# +
fig, ax = plt.subplots()
ax.xaxis.set_major_locator(ticker.MultipleLocator(base=5.))
ax.xaxis.set_minor_locator(ticker.MultipleLocator(base=1.))
ax.xaxis.set_ticks_position('both')
ax.set_yscale('log')
ax.yaxis.set_major_locator(ticker.LogLocator(base=10., numticks=12))
ax.yaxis.set_minor_locator(ticker.LogLocator(base=10.,subs=(0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9), numticks=12))
ax.yaxis.set_ticks_position('both')
ax.plot(q0s_asym, fq0_0, 'k--', label=r'$f(q_0|0)_{\mathrm{asymp.}}$')
ax.plot(q0s_asym, fq0_1, 'r--', label=r'$f(q_0|1)_{\mathrm{asymp.}}$')
ax.hist(q0s_0, bins=np.linspace(0,40,21), linewidth=2, histtype="step", density=1, label=r'$f(q_0|0)$ [MC]')
ax.hist(q0s_1, bins=np.linspace(0,40,21), linewidth=2, histtype="step", density=1, label=r'$f(q_0|1)$ [MC]')
# ax.hist(q0s_0, bins=np.linspace(0,40,41), linewidth=2, histtype="step", density=False, label=r'$f(q_0|0)$ [MC]')
# ax.hist(q0s_1, bins=np.linspace(0,40,41), linewidth=2, histtype="step", density=False, label=r'$f(q_0|1)$ [MC]')
ax.set_xlim([0,40])
ax.set_ylim([1e-8,10])
ax.set_xlabel(r'$q_0$')
ax.set_ylabel(r'$f(q_0|n)$')
ax.legend()
# +
fig, ax = plt.subplots()
ax.xaxis.set_major_locator(ticker.MultipleLocator(base=5.))
ax.xaxis.set_minor_locator(ticker.MultipleLocator(base=1.))
ax.xaxis.set_ticks_position('both')
ax.set_yscale('log')
ax.yaxis.set_major_locator(ticker.LogLocator(base=10., numticks=12))
ax.yaxis.set_minor_locator(ticker.LogLocator(base=10.,subs=(0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9), numticks=12))
ax.yaxis.set_ticks_position('both')
ax.plot(q0s_asym, fq0_0, 'k--', label=r'$f(q_0|0)_{\mathrm{asymp.}}$')
ax.plot(q0s_asym, fq0_1, 'r--', label=r'$f(q_0|1)_{\mathrm{asymp.}}$')
ax.hist(np.concatenate([q0s_0[q0s_0<25],40*np.ones(10000000)]), bins=np.linspace(0,40,41), linewidth=2, histtype="step", density=1, label=r'$f(q_0|0)$ [MC]')
ax.hist(q0s_1[q0s_1<25], bins=np.linspace(0,25,26), linewidth=2, histtype="step", density=1, label=r'$f(q_0|1)$ [MC]')
# ax.hist(q0s_0, bins=np.linspace(0,40,41), linewidth=2, histtype="step", density=False, label=r'$f(q_0|0)$ [MC]')
# ax.hist(q0s_1, bins=np.linspace(0,40,41), linewidth=2, histtype="step", density=False, label=r'$f(q_0|1)$ [MC]')
ax.set_xlim([0,25])
#ax.set_xlim([0,40])
#ax.set_ylim([1e-8,10])
ax.set_xlabel(r'$q_0$')
ax.set_ylabel(r'$f(q_0|n)$')
ax.legend()
# -
def F(q0, muprime, sigma):
return norm.cdf(q0**(1/2) - muprime/sigma)
F(25, 0, 1)
F(25, 1, sigma)
1-norm.cdf(1/sigma)
norm
N = 1000000
s=10; b=10; tau=1
q0s_0 = gen_q0s(0, s, b, tau, N)
q0s_1 = gen_q0s(1, s, b, tau, N)
plt.hist(q0s, bins=np.linspace(0,40,25), histtype="step", density=1, label='')
plt.hist(q0s, bins=np.linspace(0,40,25), histtype="step", density=1)
plt.xlabel(r'$q_0$')
N = 1000000
mu = 1; s=10; b=10; tau=1
# generate n and m bin values for N experiments
ns = np.random.poisson(lam=mu*s+b, size=N)
ms = np.random.poisson(lam=tau*b, size=N)
# likelihood is product of two poissonion variables
# drop factorials -- they cancel in likelihood ratio
def L(mu, b, s, n, m, tau):
return (mu*s+b)**n * np.exp(-(mu*s+b)) * (tau*b)**m * np.exp(-(tau*b))
# +
# estimator functions
def muhat(n, m, tau, s):
return (n-m/tau)/s
def bhat(m, tau):
return m/tau
def bhat2(n, m, tau, s, mu):
return (n+m - (1+tau)*mu*s) / (2*(1+tau)) + (((n+m-(1+tau)*mu*s)**2 + 4*(1+tau)*m*mu*s)/(4*(1+tau)**2))**(1/2)
# -
# calculate ML estimators
bh2s = bhat2(ns, ms, tau, s, 0)
bhs = ms/tau
muhs = (ns-ms/tau)/s
# calculate discovery test statistics
q0s = -2 * np.log(L(0, bh2s, s, ns, ms, tau)/L(muhs, bhs, s, ns, ms, tau))
plt.hist(q0s, bins=np.linspace(0,40,25), histtype="step", density=1);
plt.xlabel(r'$q_0$')
# +
#####
# -
Is = np.linspace(0, 200, 2001)
Is
Bs = 1e-3 * Is
Bs
plt.plot(Is, Bs)
cutoff = 65
for I in Is:
if I < cutoff:
B = a + b * I
else:
B = c + d * I
Bs = np.zeros_like(Is)
Bs
Bs[Is < cutoff] = a + b * Is[Is < cutoff]
Bs[Is >= cutoff] = c + d * Is[Is >= cutoff]
B1 = a + b * Is[Is < cutoff]
B2 = c + d * Is[Is >= cutoff]
B = np.concatenate([B1, B2])
alpha = 0.5
ys = np.cos(Is**alpha)
plt.plot(Is, ys)
| stats/notebooks/Cowan_Cranmer_Gross_Vitells_Asymptotic_Formulae_Reconstruction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import modules
import psycopg2
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
import matplotlib.pyplot as plt
warnings.simplefilter('ignore')
pd.options.display.max_columns = 300
sns.set_style('darkgrid')
# Connect to PostgreSQL
conn = psycopg2.connect(database='usaspending', user='team', password='<PASSWORD>', host='dopelytics.site', port='5432')
# ### Initialize data set
# +
# Select data elements
sql_cols = ('federal_action_obligation, '
#'total_dollars_obligated, '
'base_and_exercised_options_value, '
'base_and_all_options_value, '
#'awarding_sub_agency_name, '
'awarding_sub_agency_code, '
#'awarding_office_name, '
'awarding_office_code, '
#'funding_sub_agency_name, '
'funding_sub_agency_code, '
#'funding_office_name, ' too many NaN
'primary_place_of_performance_state_code, '
'award_or_idv_flag, '
#'award_type, '
'award_type_code, '
#'type_of_contract_pricing, '
'type_of_contract_pricing_code, '
#'dod_claimant_program_description, '
'dod_claimant_program_code, '
'type_of_set_aside_code, '
#'multi_year_contract, ' too many NaN
#'dod_acquisition_program_description, ' too many NaN
#'subcontracting_plan, ' too many NaN
#'contract_bundling, '
'contract_bundling_code, '
#'evaluated_preference, ' too many NaN
#'national_interest_action, '
'national_interest_action_code, '
#'cost_or_pricing_data, ' too many NaN
#'gfe_gfp, '
'gfe_gfp_code, '
#'contract_financing, '
'contract_financing_code, '
'portfolio_group, '
#'product_or_service_code_description, '
'product_or_service_code, '
#'naics_bucket_title, ' too many NaN
#'naics_description'
'naics_code'
)
# Create dataframe
sql_tbl_name = 'consolidated_data2'
df = pd.read_sql_query('SELECT ' + sql_cols + ' FROM ' + sql_tbl_name, con=conn)
print('Shape of initial df:', df.shape)
# Drop rows with NaN values
df = df.dropna()
print('Shape with no NaN values:', df.shape)
# +
# Create two columns for set-aside (0/1) and contract value
def set_aside(c):
if c['type_of_set_aside_code'] == 'NONE':
return 0
else:
return 1
def contract_value(c):
if c['base_and_exercised_options_value'] > 0:
return c['base_and_exercised_options_value']
elif c['base_and_all_options_value'] > 0:
return c['base_and_all_options_value']
elif c['federal_action_obligation'] > 0:
return c['federal_action_obligation']
else:
return 0
df['set_aside'] = df.apply(set_aside, axis=1)
df['contract_value'] = df.apply(contract_value, axis=1)
# Drop columns that are no longer needed
df = df.drop(['type_of_set_aside_code','base_and_exercised_options_value','base_and_all_options_value',
'federal_action_obligation'], axis=1)
# -
# ## Feature Selection
# #### Initialize Model
# +
# Create feature and target dataframes
X_int = df.drop(['set_aside'], axis = 1)
y = df['set_aside']
# One hot encoding for features
X_int = pd.get_dummies(X_int)
print('Shape of OHE feature df:', X_int.shape)
# -
# Import Random Forest Classifier modules
from sklearn.model_selection import train_test_split, cross_val_score
from yellowbrick.classifier import ClassificationReport
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import confusion_matrix, recall_score, precision_score, classification_report
# Fit initial model
X_train, X_test, y_train, y_test = train_test_split(X_int, y, test_size=0.20, random_state=42)
model = RandomForestClassifier(n_estimators=17, n_jobs=-1, random_state=0)
model.fit(X_train, y_train)
print('Model Accuracy: {:.2%}'.format(model.score(X_test, y_test)))
# #### Find Important Features
# +
# Calcultae feature importance
feature_importances = pd.DataFrame(model.feature_importances_,
index = X_train.columns,
columns=['importance']).sort_values('importance', ascending=False)
# Sort descending important features...
# Calculate cumalative percentage of total importance...
# Only keep features accounting for top 80% of feature importance
feature_importances['cumpercent'] = feature_importances['importance'].cumsum()/feature_importances['importance'].sum()*100
relevant_features = feature_importances[feature_importances.cumpercent < 80]
print('Shape of relevant features:', relevant_features.shape)
# Create list of relevant features to create new dataframe with only relevant features
list_relevant_features = list(relevant_features.index)
X = X_int[list_relevant_features]
print('Shape of initialized feature dataframe X with only relevant features:', X.shape)
# -
# Test accuracy of initialized dataframe
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20, random_state=42)
model = RandomForestClassifier(n_estimators=17, n_jobs=-1, random_state=0)
model.fit(X_train, y_train)
print('Model Accuracy: {:.2%}'.format(model.score(X_test, y_test)))
predictions = model.predict(X_test)
# ## Perform Random Forest Classification
# #### Using only relevant features dataframe
classes = ['None', 'Set Aside']
visualizer = ClassificationReport(model, classes=classes, support=True)
visualizer.score(X_test, y_test)
visualizer.show()
model_score_f1 = cross_val_score(estimator=model, X=X, y=y, scoring='f1', cv=12)
model_score_precision = cross_val_score(estimator=model, X=X, y=y, scoring='precision', cv=12)
model_score_recall = cross_val_score(estimator=model, X=X, y=y, scoring='recall', cv=12)
print("Accuracy : ", round(model_score_f1.mean(),3))
print('Standard Deviation : ',round(model_score_f1.std(),3))
print('Precision : ', round(model_score_precision.mean(),3))
print('Recall : ', round(model_score_recall.mean(),3))
print('Confusion Matrix')
print(confusion_matrix(y_test, predictions))
print(classification_report(y_test, predictions))
import pickle
#Save Trained Model
filename = 'RandomForest_SetAside_None_Model.save'
pickle.dump(model, open(filename, 'wb'))
# ## Perform Second Model - Predict Type of Set-Aside
df1 = pd.read_sql_query('SELECT ' + sql_cols + ' FROM ' + sql_tbl_name, con=conn)
print('Shape of initial df:', df1.shape)
# Drop all instances where type_of_set_aside_code = NONE
none_set_asides = df1[df1['type_of_set_aside_code'] == 'NONE'].index
df1 = df1.drop(none_set_asides, axis=0)
print('Shape of dataframe WITH set-asides:', df1.shape)
# +
# Create column for contract value
def contract_value(c):
if c['base_and_exercised_options_value'] > 0:
return c['base_and_exercised_options_value']
elif c['base_and_all_options_value'] > 0:
return c['base_and_all_options_value']
elif c['federal_action_obligation'] > 0:
return c['federal_action_obligation']
else:
return 0
df1['contract_value'] = df1.apply(contract_value, axis=1)
# Assign numerics to set-aside codes
df1['set_aside_number'] = df1['type_of_set_aside_code'].map({'SBA':1, '8AN':2, '8A':3, 'SDVOSBC':4,'HZC':5, 'WOSB':6,
'SBP':7, 'EDWOSB':7, 'SDVOSBS':7, 'HZS':7, 'WOSBSS':7,
'EDWOSBSS':7, 'ISBEE':7, 'HS3':7, 'IEE':7})
# Drop columns that are no longer needed
df1 = df1.drop(['type_of_set_aside_code','base_and_exercised_options_value','base_and_all_options_value',
'federal_action_obligation'], axis=1)
df1 = df1.dropna()
print('Shape of dataframe WITH set-asides with no NaN values:', df1.shape)
# +
X1 = df1.drop(['set_aside_number'], axis=1).copy()
print('Shape of originial X1 dataframe:', X1.shape)
# One hot encoding
X1 = pd.get_dummies(X1)
# Create a list of relevant features in X1 based on the list of previous relevant features from feature selection
# Note numpy is taking only relevant features from the first feature selection that are also in X1
cols = list(X1.columns)
updated_list_relevant_features = np.asarray(list_relevant_features)[np.in1d(list_relevant_features, cols)].tolist()
# Updated dummy table with only relevant features
X1 = X1[updated_list_relevant_features]
print('Shape of X1 dummy dataframe:', X1.shape)
# -
y1 = df1['set_aside_number'].copy()
df1.head()
y1.value_counts()
X1_train, X1_test, y1_train, y1_test = train_test_split(X1, y1, test_size=0.20, random_state=42)
model1 = RandomForestClassifier(n_estimators=17)
model1.fit(X1_train, y1_train)
X1_train.shape
classes1 = ['SBA', '8AN', '8A', 'SDVOSBC','HZC', 'WOSB', 'OTHER SET ASIDE']
visualizer = ClassificationReport(model1, classes=classes1, support=True)
visualizer.score(X1_test, y1_test)
visualizer.show()
predictions_all_set_aside = model1.predict(X1_test)
print(classification_report(y1_test, predictions_all_set_aside))
X2 = pd.DataFrame(data=X1, columns=X1.columns)
X2.head()
X2['set_aside_number'] = y1
X2.head()
# +
# Next I am testing the accuracy of the model on each specific set aside. Because we have an unbalanced data set
# it seems that the model is great for predicting set asides in general, however it is also skewed to better
# predict certain categories compared to others.
# Create a dictionary object to capture set aside code and it's score
class scores(dict):
# __init__ function
def __init__(self):
self = dict()
# Function to add key:value
def add(self, key, value):
self[key] = value
scores = scores()
percent = ''
set_aside_codes = X2['set_aside_number'].unique()
print(set_aside_codes)
# Loop through each set aside, test it, and append to the dictionary
for set_aside in set_aside_codes:
dataPoint = X2.loc[X2['set_aside_number'] == set_aside]
XPoint = dataPoint.drop(['set_aside_number'],axis=1)
yPoint = dataPoint['set_aside_number']
percent = model1.score(XPoint, yPoint)
percent = round(percent, 4)
scores.add(set_aside, percent)
# -
# Sort the dictionary by score
import operator
sortedScores = sorted(scores.items(), key=operator.itemgetter(1))
# Print scores
for score in reversed(sortedScores):
print("{:<8} {:.2%}".format(score[0], score[1]))
model_score_all_set_aside_f1 = cross_val_score(estimator=model1, X=X1, y=y1, scoring='f1_weighted', cv=12)
model_score_all_set_aside_precision = cross_val_score(estimator=model1, X=X1, y=y1, scoring='precision_weighted', cv=12)
model_score_all_set_aside_recall = cross_val_score(estimator=model1, X=X1, y=y1, scoring='recall_weighted', cv=12)
print("Accuracy : ", round(model_score_all_set_aside_f1.mean(),2))
print('Standard Deviation : ',round(model_score_all_set_aside_f1.std(),3))
print('Precision : ', round(model_score_all_set_aside_precision.mean(),3))
print('Recall : ', round(model_score_all_set_aside_recall.mean(),3))
print("")
print('Confusion Matrix')
print(confusion_matrix(y1_test, predictions_all_set_aside))
#Save Trained Model
filename = 'RandomForest_All_Set_Aside_Model.save'
pickle.dump(model, open(filename, 'wb'))
| Random Forest/DOPE_Random_Forest_Classification_Model_-Final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JavierMedel/Data-Structures-Algorithms/blob/master/First_Non_Repeating_Character.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Bozo4vSDmLCp" colab_type="text"
# # Fist Non Repeation Character
# + id="VD1JZbqOglxd" colab_type="code" colab={}
# find the first character that is repeated in a String
# + id="P7p0aonIf7T8" colab_type="code" colab={}
array = ['a','a','b','c','c','f','g']
# + id="d-5Wf01RgCWq" colab_type="code" colab={}
dir_ = {}
# + id="j-sYcPO1gCt1" colab_type="code" colab={}
first_element = array[0]
for i in range(0, len(array)):
if array[i] in dir_:
dir_[array[i]] = dir_[array[i]] + 1
else:
dir_[array[i]] = 1
# + id="aGflpu_8gFwN" colab_type="code" outputId="a2a2efbf-1067-4096-bf1c-77571e7a829b" colab={"base_uri": "https://localhost:8080/", "height": 52}
for key in dir_:
if dir_[key] == 1:
print(key)
print(dir_[key])
break
# + id="vMuo2ufLgHgs" colab_type="code" outputId="31debd68-e4fc-4729-daed-ff25f96ad242" colab={"base_uri": "https://localhost:8080/", "height": 34}
dir_
# + [markdown] id="ZNOjE2byksF4" colab_type="text"
# # Squares of a Sorted Array
# + [markdown] id="Sfxq6nfPk5ud" colab_type="text"
# Given an array of integers A sorted in non-decreasing order, return an array of the squares of each number, also in sorted non-decreasing order.
#
# Example 1:
# Input: [-4,-1,0,3,10]
# Output: [0,1,9,16,100]
#
# Example 2:
# Input: [-7,-3,2,3,11]
# Output: [4,9,9,49,121]
# + id="SlDieNyJk9c4" colab_type="code" colab={}
def squaresMarge(arr1):
return sorted([x*x for x in arr1])
# + id="hdz3XesLgcJf" colab_type="code" outputId="4d94590e-a8b6-42f5-9921-644bf0dad8f1" colab={"base_uri": "https://localhost:8080/", "height": 34}
squaresMarge([-4,-1,0,3,10])
# + id="EVuF6LLjgksX" colab_type="code" colab={}
def squareMarge2(arr):
l = 0
r = len(arr)-1
result = [None] * len(arr)
idx = len(arr)-1
while l <= r:
if abs(arr[l]) <= abs(arr[r]):
result[idx] = arr[r] ** 2
r -=1
else:
result[idx] = arr[l] ** 2
l +=1
idx -= 1
return result
# + id="5GD0eZ4VhC27" colab_type="code" outputId="f8ff29a5-9af2-4caa-bce3-426b838ecad6" colab={"base_uri": "https://localhost:8080/", "height": 34}
squareMarge2([-4,-1,0,3,10])
# + id="JZ6SNZothEXu" colab_type="code" colab={}
# Find longest Subarray BySum
# + id="Ijo8mIvf_aXY" colab_type="code" colab={}
arr = [1,2,3,4,5,6,7,8]
n = 15
# + id="60HiGHjXJUNP" colab_type="code" colab={}
def longestSubarray(arr, n):
l = 0
r = 1
sum_rlt = 0
idxs = [0,0]
while (l < len(arr) - 1) & (r < len(arr)):
print(l,r)
sum_rlt = sum(arr[l:r])
if sum_rlt == n:
print('idx:' , l , '-' , r)
print(arr[l:r])
if (r - l) >= (idxs[1] - idxs[0]):
idxs = [l, r - 1]
r += 1
elif sum_rlt < n:
r += 1
else:
l += 1
return idxs
# + id="P6KL-HB6JVL0" colab_type="code" outputId="b7031896-fde6-40fb-82ab-4a75b10dd120" colab={"base_uri": "https://localhost:8080/", "height": 294}
longestSum = longestSubarray(arr, n)
# + id="8aI_VOMmJvjv" colab_type="code" outputId="6f1c6eb0-d6b0-4639-86bb-9cb1bde0a70c" colab={"base_uri": "https://localhost:8080/", "height": 34}
arr
# + id="albXxWC5YtcN" colab_type="code" outputId="12ac2cbf-6a85-439d-a141-faacc785bb97" colab={"base_uri": "https://localhost:8080/", "height": 34}
longestSum
# + id="-yRdiiNYj_jb" colab_type="code" colab={}
| First_Non_Repeating_Character.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 5.11 残差网络(ResNet)
#
# 让我们先思考一个问题:对神经网络模型添加新的层,充分训练后的模型是否只可能更有效地降低训练误差?理论上,原模型解的空间只是新模型解的空间的子空间。也就是说,如果我们能将新添加的层训练成恒等映射$f(x) = x$,新模型和原模型将同样有效。由于新模型可能得出更优的解来拟合训练数据集,因此添加层似乎更容易降低训练误差。然而在实践中,添加过多的层后训练误差往往不降反升。即使利用批量归一化带来的数值稳定性使训练深层模型更加容易,该问题仍然存在。针对这一问题,何恺明等人提出了残差网络(ResNet) [1]。它在2015年的ImageNet图像识别挑战赛夺魁,并深刻影响了后来的深度神经网络的设计。
#
#
# ## 5.11.1残差块
#
# 让我们聚焦于神经网络局部。如图5.9所示,设输入为$\boldsymbol{x}$。假设我们希望学出的理想映射为$f(\boldsymbol{x})$,从而作为图5.9上方激活函数的输入。左图虚线框中的部分需要直接拟合出该映射$f(\boldsymbol{x})$,而右图虚线框中的部分则需要拟合出有关恒等映射的残差映射$f(\boldsymbol{x})-\boldsymbol{x}$。残差映射在实际中往往更容易优化。以本节开头提到的恒等映射作为我们希望学出的理想映射$f(\boldsymbol{x})$。我们只需将图5.9中右图虚线框内上方的加权运算(如仿射)的权重和偏差参数学成0,那么$f(\boldsymbol{x})$即为恒等映射。实际中,当理想映射$f(\boldsymbol{x})$极接近于恒等映射时,残差映射也易于捕捉恒等映射的细微波动。图5.9右图也是ResNet的基础块,即残差块(residual block)。在残差块中,输入可通过跨层的数据线路更快地向前传播。
#
# 
#
# ResNet沿用了VGG全$3\times 3$卷积层的设计。残差块里首先有2个有相同输出通道数的$3\times 3$卷积层。每个卷积层后接一个批量归一化层和ReLU激活函数。然后我们将输入跳过这两个卷积运算后直接加在最后的ReLU激活函数前。这样的设计要求两个卷积层的输出与输入形状一样,从而可以相加。如果想改变通道数,就需要引入一个额外的$1\times 1$卷积层来将输入变换成需要的形状后再做相加运算。
#
# 残差块的实现如下。它可以设定输出通道数、是否使用额外的$1\times 1$卷积层来修改通道数以及卷积层的步幅。
import tensorflow as tf
from tensorflow.keras import layers,activations
class Residual(tf.keras.Model):
def __init__(self, num_channels, use_1x1conv=False, strides=1, **kwargs):
super(Residual, self).__init__(**kwargs)
self.conv1 = layers.Conv2D(num_channels,
padding='same',
kernel_size=3,
strides=strides)
self.conv2 = layers.Conv2D(num_channels, kernel_size=3,padding='same')
if use_1x1conv:
self.conv3 = layers.Conv2D(num_channels,
kernel_size=1,
strides=strides)
else:
self.conv3 = None
self.bn1 = layers.BatchNormalization()
self.bn2 = layers.BatchNormalization()
def call(self, X):
Y = activations.relu(self.bn1(self.conv1(X)))
Y = self.bn2(self.conv2(Y))
if self.conv3:
X = self.conv3(X)
return activations.relu(Y + X)
# 下面我们来查看输入和输出形状一致的情况。
blk = Residual(3)
#tensorflow input shpe (n_images, x_shape, y_shape, channels).
#mxnet.gluon.nn.conv_layers (batch_size, in_channels, height, width)
X = tf.random.uniform((4, 6, 6 , 3))
blk(X).shape
# 我们也可以在增加输出通道数的同时减半输出的高和宽。
blk = Residual(6, use_1x1conv=True, strides=2)
blk(X).shape
# ## 5.11.2 ResNet模型
#
# ResNet的前两层跟之前介绍的GoogLeNet中的一样:在输出通道数为64、步幅为2的$7\times 7$卷积层后接步幅为2的$3\times 3$的最大池化层。不同之处在于ResNet每个卷积层后增加的批量归一化层。
net = tf.keras.models.Sequential(
[layers.Conv2D(64, kernel_size=7, strides=2, padding='same'),
layers.BatchNormalization(), layers.Activation('relu'),
layers.MaxPool2D(pool_size=3, strides=2, padding='same')])
# 一个模块的通道数同输入通道数一致。由于之前已经使用了步幅为2的最大池化层,所以无须减小高和宽。之后的每个模块在第一个残差块里将上一个模块的通道数翻倍,并将高和宽减半。
#
# 下面我们来实现这个模块。注意,这里对第一个模块做了特别处理。
class ResnetBlock(tf.keras.layers.Layer):
def __init__(self,num_channels, num_residuals, first_block=False,**kwargs):
super(ResnetBlock, self).__init__(**kwargs)
self.listLayers=[]
for i in range(num_residuals):
if i == 0 and not first_block:
self.listLayers.append(Residual(num_channels, use_1x1conv=True, strides=2))
else:
self.listLayers.append(Residual(num_channels))
def call(self, X):
for layer in self.listLayers.layers:
X = layer(X)
return X
# 接着我们为ResNet加入所有残差块。这里每个模块使用两个残差块。
# +
class ResNet(tf.keras.Model):
def __init__(self,num_blocks,**kwargs):
super(ResNet, self).__init__(**kwargs)
self.conv=layers.Conv2D(64, kernel_size=7, strides=2, padding='same')
self.bn=layers.BatchNormalization()
self.relu=layers.Activation('relu')
self.mp=layers.MaxPool2D(pool_size=3, strides=2, padding='same')
self.resnet_block1=ResnetBlock(64,num_blocks[0], first_block=True)
self.resnet_block2=ResnetBlock(128,num_blocks[1])
self.resnet_block3=ResnetBlock(256,num_blocks[2])
self.resnet_block4=ResnetBlock(512,num_blocks[3])
self.gap=layers.GlobalAvgPool2D()
self.fc=layers.Dense(units=10,activation=tf.keras.activations.softmax)
def call(self, x):
x=self.conv(x)
x=self.bn(x)
x=self.relu(x)
x=self.mp(x)
x=self.resnet_block1(x)
x=self.resnet_block2(x)
x=self.resnet_block3(x)
x=self.resnet_block4(x)
x=self.gap(x)
x=self.fc(x)
return x
mynet=ResNet([2,2,2,2])
# -
# 最后,与GoogLeNet一样,加入全局平均池化层后接上全连接层输出。
# 这里每个模块里有4个卷积层(不计算 1×1卷积层),加上最开始的卷积层和最后的全连接层,共计18层。这个模型通常也被称为ResNet-18。通过配置不同的通道数和模块里的残差块数可以得到不同的ResNet模型,例如更深的含152层的ResNet-152。虽然ResNet的主体架构跟GoogLeNet的类似,但ResNet结构更简单,修改也更方便。这些因素都导致了ResNet迅速被广泛使用。
# 在训练ResNet之前,我们来观察一下输入形状在ResNet不同模块之间的变化。
X = tf.random.uniform(shape=(1, 224, 224 , 1))
for layer in mynet.layers:
X = layer(X)
print(layer.name, 'output shape:\t', X.shape)
# ## 5.11.3 获取数据和训练模型
#
# 下面我们在Fashion-MNIST数据集上训练ResNet。
# +
(x_train, y_train), (x_test, y_test) = tf.keras.datasets.fashion_mnist.load_data()
x_train = x_train.reshape((60000, 28, 28, 1)).astype('float32') / 255
x_test = x_test.reshape((10000, 28, 28, 1)).astype('float32') / 255
mynet.compile(loss='sparse_categorical_crossentropy',
optimizer=tf.keras.optimizers.Adam(),
metrics=['accuracy'])
history = mynet.fit(x_train, y_train,
batch_size=64,
epochs=5,
validation_split=0.2)
test_scores = mynet.evaluate(x_test, y_test, verbose=2)
# -
# ## 小结
#
# * 残差块通过跨层的数据通道从而能够训练出有效的深度神经网络。
# * ResNet深刻影响了后来的深度神经网络的设计。
#
#
# ## 练习
#
# * 参考ResNet论文的表1来实现不同版本的ResNet [1]。
# * 对于比较深的网络, ResNet论文中介绍了一个“瓶颈”架构来降低模型复杂度。尝试实现它 [1]。
# * 在ResNet的后续版本里,作者将残差块里的“卷积、批量归一化和激活”结构改成了“批量归一化、激活和卷积”,实现这个改进([2],图1)。
#
#
#
# ## 参考文献
#
# [1] <NAME>., <NAME>., <NAME>., & <NAME>. (2016). Deep residual learning for image recognition. In Proceedings of the IEEE conference on computer vision and pattern recognition (pp. 770-778).
#
# [2] <NAME>., <NAME>., <NAME>., & <NAME>. (2016, October). Identity mappings in deep residual networks. In European Conference on Computer Vision (pp. 630-645). Springer, Cham.
#
#
| code/chapter05_CNN/5.11_resnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="AMuplKoptNZG"
# # Lemmatizing Words Using Wordnet
# + id="2r14xyT1tRWd"
import nltk
from nltk.stem import *
import pandas as pd
# + colab={"base_uri": "https://localhost:8080/"} id="UzpyR-dRvQj6" outputId="de8ac96b-4276-445e-85dd-f8777b4fe223"
nltk.download('punkt')
# + colab={"base_uri": "https://localhost:8080/"} id="jv-tOrXPtp9P" outputId="66bea216-bdeb-446e-8eff-e92e4964d096"
nltk.download('wordnet')
# + [markdown] id="lodevmyitmPx"
# # Lemmatizing Words
# + colab={"base_uri": "https://localhost:8080/"} id="IC0bATELtcOm" outputId="d7c1250c-3cea-4711-aee3-5dc83697b1ae"
from nltk.stem import WordNetLemmatizer
wnl = WordNetLemmatizer()
print(wnl.lemmatize('definitions'))
# + [markdown] id="T3yYOcVFt1jd"
# # Lemmatizing words by specifying parts-of-speech
# + colab={"base_uri": "https://localhost:8080/"} id="pqLiNL4atn4k" outputId="2175429c-9383-49e6-a66d-15cfa9ba975f"
print('Adjective: ', wnl.lemmatize('running', pos='a'))
print('Adverb: ', wnl.lemmatize('running', pos='r'))
print('Noun: ', wnl.lemmatize('running', pos='n'))
print('Verb: ', wnl.lemmatize('running', pos='v'))
# + id="onOqC_Sut5JO"
input_tokens = ['dictionaries', 'dictionary',
'hushed', 'hush', 'hushing',
'functional', 'functionally',
'lying', 'lied', 'lies',
'flawed', 'flaws', 'flawless',
'friendship', 'friendships', 'friendly', 'friendless',
'definitions', 'definition', 'definitely',
'the', 'these', 'those',
'motivational', 'motivate', 'motivating']
# + id="SbvX3HrRuHdG"
ss = SnowballStemmer('english')
ss_stemmed_tokens = []
for token in input_tokens:
ss_stemmed_tokens.append(ss.stem(token))
# + id="5tAyt6WMuIzA"
wnl_lemmatized_tokens = []
for token in input_tokens:
wnl_lemmatized_tokens.append(wnl.lemmatize(token, pos='v'))
# + colab={"base_uri": "https://localhost:8080/", "height": 855} id="2oAtibVOuKfc" outputId="9d310514-8730-4dee-c2ae-683434a64d2b"
stems_lemmas_df = pd.DataFrame({
'words': input_tokens,
'Snowball Stemmer': ss_stemmed_tokens,
'WordNet Lemmatizer': wnl_lemmatized_tokens
})
stems_lemmas_df
# + colab={"base_uri": "https://localhost:8080/"} id="xs-Z28oWuMIO" outputId="dc275819-6443-4bec-f82e-4b13d1cd34bb"
from nltk.tokenize import word_tokenize
with open('DLdata.txt', 'r') as f:
file_contents = f.read()
print(file_contents)
# + id="-FRxqOVwuRPc"
word_tokens = word_tokenize(file_contents)
# + id="qkjWb_d4vFoT"
wnl = WordNetLemmatizer()
lemmatized_words = []
for word in word_tokens:
lemmatized_words.append(wnl.lemmatize(word, pos="v"))
# + colab={"base_uri": "https://localhost:8080/", "height": 239} id="QUwJPwr9vVjJ" outputId="d133b4dc-97fc-4771-c5c6-daeee23337ed"
" ".join(lemmatized_words)
# + [markdown] id="7ICnOjGEvbu4"
# # Stopwords
# + colab={"base_uri": "https://localhost:8080/"} id="nKru5dqnvY8R" outputId="5ebae5eb-749e-4673-eb20-71ae05744fdf"
from nltk import word_tokenize
from nltk.corpus import stopwords
import nltk
nltk.download('stopwords')
# + colab={"base_uri": "https://localhost:8080/"} id="jzDXpNYtvn_g" outputId="78681007-ed01-4ac9-e01d-d27b5608542e"
print(stopwords.fileids())
# + colab={"base_uri": "https://localhost:8080/"} id="AVaBSxykwUeQ" outputId="14eaa22e-3397-4f7d-eb48-8c06633ae17e"
print("English: ",stopwords.words('english'))
print("Arabic: ",stopwords.words('arabic'))
# + colab={"base_uri": "https://localhost:8080/", "height": 52} id="pDlpfIZYwQue" outputId="80720121-e72c-400d-aeef-3de6eeaf6882"
text_array = ["A bird in hand is worth two in the bush.",
"Good things come to those who wait.",
"These watches cost $1500! ",
"There are other fish in the sea.",
"The ball is in your court.",
"Mr. Smith Goes to Washington ",
"<NAME>.D."]
text = " ".join(text_array)
text
# + colab={"base_uri": "https://localhost:8080/"} id="BkZm8RQWwtGu" outputId="76671252-4873-4201-ab4e-3c807417b616"
word_tokens = word_tokenize(text)
word_tokens
# + colab={"base_uri": "https://localhost:8080/"} id="kIupMi94wvYQ" outputId="0c061f3f-106b-4d56-81eb-839d4da81f79"
stop_words = set(stopwords.words('english'))
filtered_words = []
for word in word_tokens:
if word not in stop_words:
filtered_words.append(word)
print(filtered_words)
# + id="S_wnXbbjwyWx"
with open("DLdata.txt", "w") as f:
for word in filtered_words:
f.write(word)
f.write(' ')
# + colab={"base_uri": "https://localhost:8080/"} id="AM6-s8u1w8NY" outputId="fc375a61-9607-4713-e833-2c3e8e86d22e"
with open("DLdata.txt", "r") as f:
file_contents = f.read()
print(file_contents)
# + colab={"base_uri": "https://localhost:8080/"} id="bZk5qum_w_z_" outputId="2f69e0f8-0646-4c4c-9dfb-550a5affbf14"
from sklearn.feature_extraction.text import CountVectorizer
count_vectorizer = CountVectorizer()
count_vectorizer.fit([file_contents])
# + colab={"base_uri": "https://localhost:8080/"} id="VcsGnZP1xCbC" outputId="d5d32a79-4e38-463a-df7a-16837380cbfe"
transformed_vector = count_vectorizer.transform(text_array)
transformed_vector.shape
# + id="BrRMBvXNxEIO"
feature_names_nltk = count_vectorizer.get_feature_names()
# + colab={"base_uri": "https://localhost:8080/"} id="4bHLJZwcxFaE" outputId="63869962-1eba-41b1-9d43-c3d8bbbc1325"
count_vectorizer.vocabulary_
# + colab={"base_uri": "https://localhost:8080/"} id="3xPrPNG1xHfJ" outputId="c9335e5a-f1cf-4711-8b78-b756cc0b97df"
transformed_vector.toarray()
# + colab={"base_uri": "https://localhost:8080/"} id="CeonNdPPxK7x" outputId="24dc99c5-69d9-4ef1-8110-d98d4846fc42"
count_vectorizer.inverse_transform(transformed_vector)
# + [markdown] id="YxfRUhhLxNvX"
# # Removing Stpwords Using sklearn
# + colab={"base_uri": "https://localhost:8080/"} id="0-XQyePXxMsa" outputId="58d9235c-c82f-4b0e-a6ea-9dc272b14e31"
count_vectorizer = CountVectorizer(stop_words='english')
transformed_vector = count_vectorizer.fit_transform(text_array)
transformed_vector.shape
# + id="iGjqOb-CxTbE"
feature_names_sklearn = count_vectorizer.get_feature_names()
# + colab={"base_uri": "https://localhost:8080/"} id="bfOu-dh8xVw7" outputId="bbb92dde-23bc-4cd9-af9c-b9cd820d3619"
transformed_vector.toarray()
# + colab={"base_uri": "https://localhost:8080/"} id="TR6bx6YMxXV2" outputId="f473eae0-511e-4b90-aea5-f26271eae434"
count_vectorizer.inverse_transform(transformed_vector)
# + [markdown] id="vhk5eAtAxcoe"
# # Set Difference of Both
#
# + id="BBeWV-YFxZAs"
def set_diff(first, second):
second = set(second)
return [item for item in first if item not in second]
# + colab={"base_uri": "https://localhost:8080/"} id="i4ACOu9uxa73" outputId="a9b1c196-2b21-4687-8b83-a0f700bc2c6a"
set_diff(feature_names_sklearn, feature_names_nltk)
# + colab={"base_uri": "https://localhost:8080/"} id="KwjTG4AXxir7" outputId="12465d78-8258-415f-e2a6-2d313551a960"
set_diff(feature_names_nltk, feature_names_sklearn)
# + [markdown] id="H9N-7bl3xmk2"
# # Filtering words based on frequency
# + colab={"base_uri": "https://localhost:8080/"} id="syqplXmfxj8v" outputId="3e465bce-e8c5-4b44-df6a-38a0807b4278"
from sklearn.datasets import fetch_20newsgroups
newsgroups = fetch_20newsgroups(subset='train')
# + colab={"base_uri": "https://localhost:8080/"} id="z5ztVt7axoa9" outputId="40d3256a-141d-4eae-b0b2-9305fbbf589c"
newsgroups.keys()
# + colab={"base_uri": "https://localhost:8080/"} id="pILvs8DDxrY9" outputId="93524777-94a2-4fb0-c5bc-1ee1d7d30669"
print(newsgroups.data[0])
# + colab={"base_uri": "https://localhost:8080/"} id="24_e2jYSxtqo" outputId="1a3b306b-b5f5-4cb4-f70f-8090b7a737c7"
newsgroups.target_names
# + colab={"base_uri": "https://localhost:8080/"} id="ss963Q5cxva8" outputId="f981a852-32a7-4b62-91b3-3e4cfaed69a5"
count_vectorizer = CountVectorizer()
transformed_vector = count_vectorizer.fit_transform(newsgroups.data)
transformed_vector.shape
| lemmatization and remove stopwords on text data using NLTK.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Finite State Machine Generator
#
# This notebook will show how to use the Finite State Machine (FSM) Generator to generate a state machine. The FSM we will build is a Gray code counter. The counter has three state bits and can count up or down through eight states. The counter outputs are Gray coded, meaning that there is only a single-bit transition between the output vector of any state and its next states.
# ### Step 1: Download the `logictools` overlay
# +
from pynq.overlays.logictools import LogicToolsOverlay
from pynq.lib.logictools import FSMGenerator
logictools_olay = LogicToolsOverlay('logictools.bit')
# -
# ### Step 2: Specify the FSM
fsm_spec = {'inputs': [('reset','D0'), ('direction','D1')],
'outputs': [('bit2','D3'), ('bit1','D4'), ('bit0','D5')],
'states': ['S0', 'S1', 'S2', 'S3', 'S4', 'S5', 'S6', 'S7'],
'transitions': [['01', 'S0', 'S1', '000'],
['00', 'S0', 'S7', '000'],
['01', 'S1', 'S2', '001'],
['00', 'S1', 'S0', '001'],
['01', 'S2', 'S3', '011'],
['00', 'S2', 'S1', '011'],
['01', 'S3', 'S4', '010'],
['00', 'S3', 'S2', '010'],
['01', 'S4', 'S5', '110'],
['00', 'S4', 'S3', '110'],
['01', 'S5', 'S6', '111'],
['00', 'S5', 'S4', '111'],
['01', 'S6', 'S7', '101'],
['00', 'S6', 'S5', '101'],
['01', 'S7', 'S0', '100'],
['00', 'S7', 'S6', '100'],
['1-', '*', 'S0', '']]}
# __Notes on the FSM specification format__
#
# 
# ### Step 3: Instantiate the FSM generator object
fsm_generator = logictools_olay.fsm_generator
# __Setup to use trace analyzer__
# In this notebook, the trace analyzer is used to check if the inputs and outputs of the FSM.
#
# Users can choose whether to use the trace analyzer by calling the `trace()` method.
fsm_generator.trace()
# ### Step 5: Setup the FSM generator
# The FSM generator will work at the default frequency of 10MHz. This can be modified using a `frequency` argument in the `setup()` method.
fsm_generator.setup(fsm_spec)
# __Display the FSM state diagram__
# This method should only be called after the generator has been properly set up.
fsm_generator.show_state_diagram()
# __Set up the FSM inputs on the PYNQ board__
# * Check that the reset and direction inputs are correctly wired on the PYNQ board, as shown below:
# * Connect D0 to GND
# * Connect D1 to 3.3V
#
# 
# __Notes:__
#
# * The 3-bit Gray code counter is an up-down, wrap-around counter that will count from states 000 to 100 in either ascending or descending order
#
#
# * The reset input is connected to pin D0 of the Arduino connector
# * Connect the reset input to GND for normal operation
# * When the reset input is set to logic 1 (3.3V), the counter resets to state 000
#
#
# * The direction input is connected to pin D1 of the Arduino connector
# * When the direction is set to logic 0, the counter counts down
# * Conversely, when the direction input is set to logic 1, the counter counts up
# ### Step 6: Run and display waveform
#
# The ` run()` method will execute all the samples, `show_waveform()` method is used to display the waveforms
fsm_generator.run()
fsm_generator.show_waveform()
# #### Verify the trace output against the expected Gray code count sequence
#
# | State | FSM output bits: bit2, bit1, bit0 |
# |:-----:|:----------------------------------------:|
# | s0 | 000 |
# | s1 | 001 |
# | s2 | 011 |
# | s3 | 010 |
# | s4 | 110 |
# | s5 | 111 |
# | s6 | 101 |
# | s7 | 100 |
#
#
#
# ### Step 7: Stop the FSM generator
# Calling `stop()` will clear the logic values on output pins; however, the waveform will be recorded locally in the FSM instance.
fsm_generator.stop()
| Session_3/4_fsm_generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="2quaM0o5G404"
# <h1><center>Universidade Federal de Minas Gerais</center></h1>
# <h3><center>Departamento de Ciência da Computação</center></h3>
# <h3><center>Introdução a Banco de Dados - Trabalho Prático #1</center></h3>
#
# + [markdown] id="__T58tCjoXSx"
# # 1. Introdução
# + [markdown] id="r9gaXCSG1-6y"
# ### 1.1. Contexto
# + [markdown] id="8glNz6nlG405"
# Nesse trabalho prático utilizaremos o banco de dados representado pelo schema abaixo:
#
# 
# + [markdown] id="O-XWIzUOyW_b" pycharm={"name": "#%% md\n"}
# ### 1.2. Entrega
#
# + [markdown] id="AvrMbGsEyjUu" pycharm={"name": "#%% md\n"}
#
#
# 1. Você deve completar esse notebook, executar todas as células e submetê-lo no **Minha UFMG**.
# 2. Você pode entregar até o dia 29/07/2021 às 23:59.
# 3. O trabalho é individual e vale 15 pontos.
#
#
#
#
#
# + [markdown] id="HJwmiueqok-8"
# # 2. Preparação do Ambiente
# + id="Wv-_wG6VG408"
# importando os packages necessários
import io
import sqlite3
import pandas as pd
import matplotlib.pyplot as plt
# + id="wFSKBvDfG41C" pycharm={"name": "#%%\n"} colab={"base_uri": "https://localhost:8080/"} outputId="d25ba8a1-8d3f-4bc1-9bfe-b2a479958245"
# download do dump do banco de dados:
# !wget https://raw.githubusercontent.com/claudiovaliense/ibd/master/data/despesas_publicas_tp1.sql
# + [markdown] id="FTqbHHOw2iEE" pycharm={"name": "#%% md\n"}
# ### 2.1. Criação da conexão com o banco
# * No momento da conexão, é informado o nome do arquivo onde será montado o banco de dados. No nosso caso, este arquivo não precisa ser pré-existente, porque iremos carregar as informações de um dump;
# * Se este comando for executado com um arquivo que já existe e possui as mesmas tabelas que serão criadas, ocorrerá um erro no momento da execução do script mais à frente.
# + id="wuRZ8VP3G41G" pycharm={"name": "#%%\n"}
conn = sqlite3.connect('despesas.db')
cursor = conn.cursor()
# + [markdown] id="Us7-HZg-G41N" pycharm={"name": "#%% md\n"}
# ### 2.2. Execução do script para inicializar e popular o banco de dados
#
# * Criada a conexão, iremos utilizar o arquivo .sql disponibilizado para recuperar o schema e os dados do banco
# + id="3tthWfmTG41P" pycharm={"name": "#%%\n"} colab={"base_uri": "https://localhost:8080/"} outputId="8f45966f-18fa-46c4-a9c4-150eb90ed2fd"
f = io.open('despesas_publicas_tp1.sql', 'r', encoding='ISO-8859-1')
sql = f.read()
cursor.executescript(sql)
# + [markdown] id="bq5lfEMRG41T"
# ### 2.3. Realizando consultas no banco de dados
#
# * Criada a conexão e realizada a carga dos dados no banco, pode ser executada qualquer consulta SQL;
# * O SQLite3 permite que o retorno dos dados seja feito em vários formatos, entretanto, utilizaremos o retorno em um dataframe do Pandas.
#
#
#
# **CONSULTA EXEMPLO:** todos os órgãos ordenados pela quantidade de orgãos subordinados de forma decrescente.
# + id="WcR2nrjLG41T" pycharm={"name": "#%%\n"} colab={"base_uri": "https://localhost:8080/", "height": 489} outputId="529a815a-36e3-4e3d-8e40-a1292ba091a0"
df = pd.read_sql_query("SELECT * FROM ORGAO ORDER BY QTD_ORGAOS_SUBORDINADOS DESC", conn)
df
# + [markdown] id="8n1eSJAcppwc" pycharm={"name": "#%% md\n"}
# #3. Consultas
# + [markdown] id="sf2LzUSWs4nM"
# Expresse cada consulta abaixo utilizando SQL:
#
# + [markdown] id="eM71aOGsG41X" pycharm={"name": "#%% md\n"}
# **1.** Retorne a quantidade de palavras em cada registro da coluna nome_orgao da tabela orgão.
# + id="x6bTFKHHG41j" pycharm={"name": "#%%\n"}
# + [markdown] id="5631TeDlG41m" pycharm={"name": "#%% md\n"}
#
#
# ```
# # Isto está formatado como código
# ```
#
# **2.** Dado a coluna nome_orgao, retorne a primeira posição que a string 'da' aparece da tabela orgão
# + id="XBFBGjjTG41n" pycharm={"name": "#%%\n"}
# + [markdown] id="0Hk-EcnpG41q" pycharm={"name": "#%% md\n"}
# **3.** Liste todos os nomes dos órgãos e o valor das despesas totais no ano de 2018 para os mesmos, ordenando de forma decrescente pelo valor.
# + id="LIEa2HJRG41r" pycharm={"name": "#%%\n"}
# + [markdown] id="XbWL79JAG41y" pycharm={"name": "#%% md\n"}
# **4.** Liste o mês e ano, modalidade, programa de governo e valor de todas as despesas do órgão Ministério da Fazenda no primeiro semestre de 2018.
#
# > Bloco com recuo
#
#
# + id="SNDpxc_3G41z" pycharm={"name": "#%%\n"}
# + [markdown] id="Yo9Iln_yG412" pycharm={"name": "#%% md\n"}
# **5.** Liste o código e nome dos programas de governo que possuíram os 3 maiores valores de despesas totais na função de educação durante o ano de 2018.
# + id="2C3s7o8sG413" pycharm={"name": "#%%\n"}
# + [markdown] id="4TpNWfhxG416" pycharm={"name": "#%% md\n"}
# **6.** Transforme os caracteres em minúsculo e remova todos os termos 'da' na coluna nome_orgao da tabela orgão.
# + id="eZ25oY25G417" pycharm={"name": "#%%\n"}
# + [markdown] id="5vg_FHhhG42C" pycharm={"name": "#%% md\n"}
# **7.** Liste os órgãos governamentais e a quantidade de programas de governo distintos envolvidos nas despesas públicas desses órgãos. Nesta consulta, desconsidere valores de despesa negativos ou iguais a zero no período.
# + id="8m2VR3rYG42C" pycharm={"name": "#%%\n"}
# + [markdown] id="4aTLYKVlG42F" pycharm={"name": "#%% md\n"}
# **8.** Liste os nomes das subfunções das despesas públicas envolvidas para cada programa de governo. Sua consulta deve retornar o nome do programa de governo, a subfunção da despesa, a quantidade de entradas de despesas públicas e o valor total dessas despesas.
# + id="SOJHEEkFG42G" pycharm={"name": "#%%\n"}
# + [markdown] id="OD7TxifCG42K" pycharm={"name": "#%% md\n"}
# **9.** Liste todas as despesas públicas (código da despesa, mês e ano da despesa, modalidade, valor) em que o órgão responsável é aquele que possui o maior número de órgãos subordinados.
# + id="qIdkOCpSG42K" pycharm={"name": "#%%\n"}
# + [markdown] id="k11q5Eh0G42Q" pycharm={"name": "#%% md\n"}
# **10.** Liste todas as despesas públicas (código da despesa, mês e ano da despesa, modalidade, valor) em que o órgão responsável é aquele que possui o maior valor total em despesas durante o ano de 2018.
# + id="W313teDOG42R" pycharm={"name": "#%%\n"}
# + [markdown] id="bDN01TwoG42U" pycharm={"name": "#%% md\n"}
# **11.** Liste os códigos e nomes dos programas de governo relacionados ao órgão (ou órgãos, caso haja empate) que teve menos registros de despesas públicas durante o ano.
# + id="r16cTfiGG42V" pycharm={"name": "#%%\n"}
# + [markdown] id="x18gb-yqG42Y" pycharm={"name": "#%% md\n"}
# **12.** Álgebra relacional: projeção do NOME_FUNCAO e NOME_SUBFUNCAO da junção natural das tabelas FUNCAO, DETALHAMENTO_FUNCAO e SUBFUNCAO
# + id="73jm4FOHG42Z" pycharm={"name": "#%%\n"}
# + [markdown] id="Mwv-emqQG42c" pycharm={"name": "#%% md\n"}
# **13.** Álgebra relacional: projeção de MODALIDADE_DESPESA, VALOR_DESPESA e NOME_PROGRAMA_GOVERNO da junção natural das tabelas PROGRAMA_GOVERNO e DESPESA_PUBLICA onde a MODALIDADE_DESPESA = 'Reserva de Contingência'
# + id="JG2E4JajG42d" pycharm={"name": "#%%\n"}
# + [markdown] id="TVqX_v22G42i" pycharm={"name": "#%% md\n"}
# **14.** Álgebra relacional: projeção de VALOR_DESPESA e NOME_ORGAO da junção natural das tabelas DESPESA_PUBLICA e ORGAO onde o NOME_ORGAO = 'Ministério da Fazenda'
# + id="0ylNfNJoG42j" pycharm={"name": "#%%\n"}
# + [markdown] id="ikjln39psbK6"
# **15.** Álgebra relacional: projeção de NOME_PROGRAMA_GOVERNO e VALOR_DESPESA da junção natural das tabelas DESPESA_PUBLICA, PROGRAMA_GOVERNO e ORGAO onde o NOME_ORGAO = 'Ministério da Saúde'
# + pycharm={"name": "#%%\n"} id="jetjhyCK1Lyj"
# + [markdown] id="clX3ruU9G42q"
# # 4. Fechamento da conexão com o banco
#
# * Após serem realizadas todas as consultas necessárias, é aconselhavel encerrar formalmente a conexão com o banco de dados
# + id="my0mPqYVG42r"
conn.close()
| ibd_tp_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/otwn/Japan-Geospatial-Data-Analysis/blob/master/Protected_Areas_in_Japan.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="b1Xi_EZTMSAh" colab_type="text"
# # Protected Areas in Japan
#
#
# + id="ZhwEx7yaNBFv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 260} outputId="b79404b8-4f21-46d6-b1df-047e60e34941"
# !pip install geopandas shapely
# + id="_FmdLtYBNJNP" colab_type="code" colab={}
import pandas as pd
import geopandas as gpd
import shapely
# + [markdown] id="p1T-BusNMWqt" colab_type="text"
# # Data
# 1. Protected Areas in Japan
# * UNEP-WCMC (2020). Protected Area Profile for Japan from the World Database of Protected Areas, August 2020. Available at: www.protectedplanet.net
# * https://hub.arcgis.com/datasets/b50f002de336438895df1a9b03efd423?geometry=-168.047%2C-88.664%2C168.047%2C88.173&layer=0
# * filter ISO3=="JPN"
# * Shapefile zip
#
#
# 2. Japan Prefecture
# * https://hub.arcgis.com/datasets/esri::%E5%B9%B3%E6%88%90-27-%E5%B9%B4%E5%9B%BD%E5%8B%A2%E8%AA%BF%E6%9F%BB-%E9%83%BD%E9%81%93%E5%BA%9C%E7%9C%8C%E7%95%8C-japan-prefecture-boundaries-ecm?layer=0
#
#
#
#
# CSV file downloaded from https://www.protectedplanet.net/country/JP
# Located in ./dataset
# + id="Ior2uwfrMNVZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 569} outputId="aba90880-d753-4f5a-f7ef-08bc7922df5f"
pts_url = "https://gis.unep-wcmc.org/arcgis/rest/services/wdpa/public/MapServer/0/query?outFields=*&f=geojson&where=ISO3%20%3D%20'JPN'"
protected_areas_pts_japan = gpd.read_file(pts_url)
protected_areas_pts_japan.head()
# + id="w5BMHpDOTi6S" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 569} outputId="94f7fbf5-0c11-424b-80ae-214d442e6981"
# Platform: ArcGIS Hub https://hub.arcgis.com/datasets
# Dataset: WDPA_poly_Aug2020
# Config: using a filter at the column: ISO3
# Parameter: ISO3=="JPN"
poly_url = "https://gis.unep-wcmc.org/arcgis/rest/services/wdpa/public/MapServer/1/query?outFields=*&f=geojson&where=ISO3%20%3D%20'JPN'"
protected_areas_poly_japan = gpd.read_file(poly_url)
protected_areas_poly_japan.head()
# + [markdown] id="_P9BvAceOSee" colab_type="text"
# # Plot the data (Japan)
# + id="XG0FKnqbOPGI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 202} outputId="44f8b2a5-1a1c-4462-8852-0d4c60cfc9c3"
url = "https://services.arcgis.com/P3ePLMYs2RVChkJx/arcgis/rest/services/JPN_Boundaries_ECM/FeatureServer/0/query?outFields=*&where=1%3D1&f=geojson"
japan = gpd.read_file(url)
japan.head()
# + id="hNXNV5o3bN-9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 202} outputId="2b8a3363-a54b-417f-c253-d1d3857de7fc"
# plotting the surrounding areas
# gpd has own world map
world_filepath = gpd.datasets.get_path('naturalearth_lowres')
world = gpd.read_file(world_filepath)
# east_asia = world.loc[world.continent=="Asia"].copy()
east_asia = world.loc[world.name.isin(["Japan", "South Korea", "North Korea", "China", "Taiwan"])].copy()
east_asia.head()
# + id="or104NXTNgs5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 881} outputId="3bebc613-9c49-4895-c4bd-d1c88c9ffda9"
# ax = east_asia.plot(figsize=(10,10), color="whitesmoke", linestyle=":", edgecolor="black")
# japan.plot(color="lightgray", ax=ax)
ax = japan.plot(figsize=(15,15), color="whitesmoke", linestyle=":", edgecolor="black")
protected_areas_pts_japan.plot(markersize=2, ax=ax)
protected_areas_poly_japan.plot(color="lightgreen", ax=ax)
# + id="sS136CiCPA6n" colab_type="code" colab={}
| Japan/Protected_Areas_in_Japan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.sys.path.append(os.path.dirname(os.path.abspath('.')))
# # 数据准备
import numpy as np
X = np.array([[1, 2], [3, 4], [1, 2], [3, 4]])
y = np.array([1, 2, 3, 4])
# # k-fold
# 迭代方式实现KFold很简单,不断地更改测试集的位置即可,剩余的位置就构成训练集,如下所示:
# +
K = 3 # fold数
test_size = len(X)//K # 测试集尺寸
idx_arr = np.arange(len(X)) # 生成索引数组
for epoch in range(K):
test_idx = idx_arr[epoch*test_size:(epoch+1)*test_size]
train_idx = np.append(idx_arr[0:epoch*test_size],
idx_arr[(epoch+1)*test_size:]) # 利用切片不会越界的性质
print(train_idx, test_idx)
# -
# 生成器方式:
# +
K = 3
test_size = len(X)//K
idx_arr = np.arange(len(X))
gen = ((np.append(idx_arr[0:epoch*test_size], idx_arr[(epoch+1)*test_size:]),
idx_arr[epoch*test_size:(epoch+1)*test_size]) for epoch in range(K))
for train, test in gen:
print(train, test)
# -
# # Stratified k-fold
# 当数据目标值为类别型数据时,分层抽样是最佳选择,在每个类别下都分别做k-fold。
X = np.ones(10)
y = np.array([0, 0, 0, 0, 1, 1, 1, 1, 1, 1])
# +
K = 2
n_samples = len(y)
for epoch in range(K):
# 空索引,等待扩展
train_idx = np.array(list()).astype(int)
test_idx = np.array(list()).astype(int)
# 遍历y所有可能的取值并扩展索引
for y_val in np.unique(y):
idxs = np.arange(n_samples)[y == y_val] # 当前类别下的全部索引
test_size = len(idxs)//K
train_idx = np.append(train_idx, np.append(
idxs[0:epoch*test_size], idxs[(epoch+1)*test_size:]))
test_idx = np.append(
test_idx, idxs[epoch*test_size:(epoch+1)*test_size])
print(train_idx, test_idx)
# -
| model_selection/KFold.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import cufflinks as cf
# %matplotlib inline
from plotly.offline import download_plotlyjs,init_notebook_mode,plot,iplot
init_notebook_mode(connected=True)
cf.go_offline()
df = pd.DataFrame(np.random.randn(100,4),columns='A B C D'.split())
df.head()
df2 = pd.DataFrame({'Category':"A B C".split(),'Values':[32,43,50]})
df2
df.iplot() # looucura
df.iplot(kind='scatter',x='A',y='B',mode='markers')
df2.iplot(kind='bar',x='Category')
df3 = pd.DataFrame({'x':[1,2,3,4,5],'y':[10,20,30,20,10],'z':[500,400,300,200,100]})
df3.iplot(kind='surface',colorscale='rdylbu')
df.iplot('hist')
| Plotly_visualization/.ipynb_checkpoints/PlotLy and cufflink-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
from ggplot import *
import pandas as pd
df = pd.melt(meat, id_vars="date")
ggplot(diamonds, aes(x='carat', y='price', color='cut')) + \
geom_point() + \
facet_grid("clarity", "color")
ggplot(diamonds, aes(x='carat', y='price')) + \
stat_smooth(method='loess') + \
facet_grid("clarity", "cut")
| docs/examples/Faceting Basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import xarray
import netCDF4 as nc
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
feature_fn = '/beegfs/DATA/pritchard/srasp/preprcessed_data/full_physics_essentials_train_test_features.nc'
target_fn = '/beegfs/DATA/pritchard/srasp/preprcessed_data/full_physics_essentials_train_test_targets.nc'
norm_fn = '/beegfs/DATA/pritchard/srasp/preprcessed_data/full_physics_essentials_train_test_norm.nc'
aqua_fn = '/beegfs/DATA/pritchard/srasp/Aquaplanet_enhance05/AndKua_aqua_SPCAM3.0_enhance05.cam2.h1.0000-01-01-00000.nc'
features = nc.Dataset(feature_fn)
norm = nc.Dataset(norm_fn)
features
norm
features['feature_names'][:]
unnorm_features = features['features'][:] * norm['feature_stds'][:] + norm['feature_means'][:]
unnorm_features.shape
aqua = nc.Dataset(aqua_fn); aqua
# sample, lev --> [time, lat, lon]
unnorm_features = unnorm_features.reshape(-1, 64, 128, 92)
ctrl_TBP = aqua['TAP'][:] - aqua['TPHYSTND'][:] * 1800.
ctrl_TBP.shape
ctrl_TBP = ctrl_TBP[1:]
unnorm_features.shape
unnorm_features[0, 0, 0, :5]
ctrl_TBP[0, :5, 0, 0]
# Check! Computed BPs correctly
unnorm_features[0, 0, 0, -1] # SOLIN
aqua['SOLIN'][1, 0, 0]
unnorm_features[0, 0, 0, -2] # PS
aqua['PS'][0, 0, 0]
# Check! Took the correct time steps for PS
norm['target_names'][:]
# Now check the targets.
targets = nc.Dataset(target_fn); targets
targets['target_names'][:]
plt.plot(np.mean(targets['targets'][:], axis=0))
# Now check the pure crm inputs file
feature_fn2 = '/beegfs/DATA/pritchard/srasp/preprcessed_data/full_physics_essentials_train_test2_features.nc'
target_fn2 = '/beegfs/DATA/pritchard/srasp/preprcessed_data/full_physics_essentials_train_test2_targets.nc'
norm_fn2 = '/beegfs/DATA/pritchard/srasp/preprcessed_data/full_physics_essentials_train_test2_norm.nc'
features2 = nc.Dataset(feature_fn2)
norm2 = nc.Dataset(norm_fn2)
features2['feature_names'][:]
unnorm_features2 = features2['features'][:] * norm2['feature_stds'][:] + norm2['feature_means'][:]
unnorm_features2.shape
# sample, lev --> [time, lat, lon]
unnorm_features2 = unnorm_features2.reshape(-1, 64, 128, 152)
# T_C = TAP[t-1] - DTV[t-1] * dt
ctrl_T_C = aqua['TAP'][:-1] - aqua['DTV'][:-1] * 1800.
unnorm_features2[0, 0, 0, :5]
ctrl_T_C[0, :5, 0, 0]
# adiab = (TBP - TC)/dt
ctrl_T_C.shape, ctrl_TBP.shape
ctrl_adiab = (ctrl_TBP - ctrl_T_C) / 1800.
features2['feature_names'][90]
unnorm_features2[0, 0, 0, 90:95]
ctrl_adiab[0, :4, 0, 0]
# Check, I think I did everything correct
| notebooks/dev/old_notebooks/test_preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] nbsphinx="hidden"
# This notebook is part of the $\omega radlib$ documentation: https://docs.wradlib.org.
#
# Copyright (c) $\omega radlib$ developers.
# Distributed under the MIT License. See LICENSE.txt for more info.
# -
# # xarray IRIS backend
#
# In this example, we read IRIS (sigmet) data files using the wradlib `iris` xarray backend.
import glob
import gzip
import io
import wradlib as wrl
import warnings
warnings.filterwarnings('ignore')
import matplotlib.pyplot as pl
import numpy as np
import xarray as xr
try:
get_ipython().magic("matplotlib inline")
except:
pl.ion()
# ## Load IRIS Volume Data
fpath = 'sigmet/SUR210819000227.RAWKPJV'
f = wrl.util.get_wradlib_data_file(fpath)
vol = wrl.io.open_iris_dataset(f, reindex_angle=False)
# ### Inspect RadarVolume
display(vol)
# ### Inspect root group
#
# The `sweep` dimension contains the number of scans in this radar volume. Further the dataset consists of variables (location coordinates, time_coverage) and attributes (Conventions, metadata).
vol.root
# ### Inspect sweep group(s)
#
# The sweep-groups can be accessed via their respective keys. The dimensions consist of `range` and `time` with added coordinates `azimuth`, `elevation`, `range` and `time`. There will be variables like radar moments (DBZH etc.) and sweep-dependend metadata (like `fixed_angle`, `sweep_mode` etc.).
display(vol[0])
# ### Goereferencing
swp = vol[0].copy().pipe(wrl.georef.georeference_dataset)
# ### Plotting
swp.DBZH.plot.pcolormesh(x='x', y='y')
pl.gca().set_aspect('equal')
fig = pl.figure(figsize=(10,10))
swp.DBZH.wradlib.plot_ppi(proj='cg', fig=fig)
# +
import cartopy
import cartopy.crs as ccrs
import cartopy.feature as cfeature
map_trans = ccrs.AzimuthalEquidistant(central_latitude=swp.latitude.values,
central_longitude=swp.longitude.values)
# -
map_proj = ccrs.AzimuthalEquidistant(central_latitude=swp.latitude.values,
central_longitude=swp.longitude.values)
pm = swp.DBZH.wradlib.plot_ppi(proj=map_proj)
ax = pl.gca()
ax.gridlines(crs=map_proj)
print(ax)
map_proj = ccrs.Mercator(central_longitude=swp.longitude.values)
fig = pl.figure(figsize=(10,8))
ax = fig.add_subplot(111, projection=map_proj)
pm = swp.DBZH.wradlib.plot_ppi(ax=ax)
ax.gridlines(draw_labels=True)
# +
import cartopy.feature as cfeature
def plot_borders(ax):
borders = cfeature.NaturalEarthFeature(category='physical',
name='coastline',
scale='10m',
facecolor='none')
ax.add_feature(borders, edgecolor='black', lw=2, zorder=4)
map_proj = ccrs.Mercator(central_longitude=swp.longitude.values)
fig = pl.figure(figsize=(10,8))
ax = fig.add_subplot(111, projection=map_proj)
DBZH = swp.DBZH
pm = DBZH.where(DBZH > 0).wradlib.plot_ppi(ax=ax)
plot_borders(ax)
ax.gridlines(draw_labels=True)
# +
import matplotlib.path as mpath
theta = np.linspace(0, 2*np.pi, 100)
center, radius = [0.5, 0.5], 0.5
verts = np.vstack([np.sin(theta), np.cos(theta)]).T
circle = mpath.Path(verts * radius + center)
map_proj = ccrs.AzimuthalEquidistant(central_latitude=swp.latitude.values,
central_longitude=swp.longitude.values,
)
fig = pl.figure(figsize=(10,8))
ax = fig.add_subplot(111, projection=map_proj)
ax.set_boundary(circle, transform=ax.transAxes)
pm = swp.DBZH.wradlib.plot_ppi(proj=map_proj, ax=ax)
ax = pl.gca()
ax.gridlines(crs=map_proj)
# -
fig = pl.figure(figsize=(10, 8))
proj=ccrs.AzimuthalEquidistant(central_latitude=swp.latitude.values,
central_longitude=swp.longitude.values)
ax = fig.add_subplot(111, projection=proj)
pm = swp.DBZH.wradlib.plot_ppi(ax=ax)
ax.gridlines()
swp.DBZH.wradlib.plot_ppi()
# ### Inspect radar moments
#
# The DataArrays can be accessed by key or by attribute. Each DataArray has dimensions and coordinates of it's parent dataset.
display(swp.DBZH)
# ### Create simple plot
#
# Using xarray features a simple plot can be created like this. Note the `sortby('rtime')` method, which sorts the radials by time.
swp.DBZH.sortby('rtime').plot(x="range", y="rtime", add_labels=False)
fig = pl.figure(figsize=(5,5))
pm = swp.DBZH.wradlib.plot_ppi(proj={'latmin': 3e3}, fig=fig)
# ### Mask some values
dbzh = swp['DBZH'].where(swp['DBZH'] >= 0)
dbzh.plot(x="x", y="y")
vol[0]
# ### Export to ODIM and CfRadial2
#
# Need to remove DB_XHDR since it can't be represented as ODIM/CfRadial2 moment.
vol[0] = vol[0].drop("DB_XHDR", errors="ignore")
vol[0].DBZH.sortby("rtime").plot(y="rtime")
vol.to_odim('iris_as_odim.h5')
vol.to_cfradial2('iris_as_cfradial2.nc')
# ### Import again
vola = wrl.io.open_odim_dataset('iris_as_odim.h5', reindex_angle=False, keep_elevation=True)
display(vola.root)
display(vola[0])
vola[0].DBZH.sortby("rtime").plot(y="rtime")
volb = wrl.io.open_cfradial2_dataset('iris_as_cfradial2.nc')
display(volb.root)
display(volb[0])
volb[0].DBZH.sortby("rtime").plot(y="rtime")
# ### Check equality
#
# We have to drop the time variable when checking equality since IRIS has millisecond resolution.
xr.testing.assert_allclose(vol.root.drop("time"), vola.root.drop("time"))
xr.testing.assert_allclose(vol[0].drop(["rtime", "time", "DB_HCLASS2"]), vola[0].drop(["rtime", "time"]))
xr.testing.assert_allclose(vol.root.drop("time"), volb.root.drop("time"))
xr.testing.assert_allclose(vol[0].drop("time"), volb[0].drop("time"))
xr.testing.assert_allclose(vola.root, volb.root)
xr.testing.assert_allclose(vola[0].drop("rtime"), volb[0].drop(["rtime", "DB_HCLASS2"]))
# ## More Iris loading mechanisms
# ### Use `xr.open_dataset` to retrieve explicit group
swp = xr.open_dataset(f, engine="iris", group=1, backend_kwargs=dict(reindex_angle=False))
display(swp)
| notebooks/fileio/wradlib_iris_backend.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.020212, "end_time": "2021-06-04T13:07:04.990938", "exception": false, "start_time": "2021-06-04T13:07:04.970726", "status": "completed"} tags=[]
# # Loops
#
# Loops are a way to repeatedly execute some code. Here's an example:
# + papermill={"duration": 0.03233, "end_time": "2021-06-04T13:07:05.042553", "exception": false, "start_time": "2021-06-04T13:07:05.010223", "status": "completed"} tags=[]
planets = ['Mercury', 'Venus', 'Earth', 'Mars', 'Jupiter', 'Saturn', 'Uranus', 'Neptune']
for planet in planets:
print(planet, end=' ') # print all on same line
# + [markdown] papermill={"duration": 0.019769, "end_time": "2021-06-04T13:07:05.082722", "exception": false, "start_time": "2021-06-04T13:07:05.062953", "status": "completed"} tags=[]
# The ``for`` loop specifies
# - the variable name to use (in this case, `planet`)
# - the set of values to loop over (in this case, `planets`)
#
# You use the word "``in``" to link them together.
#
# The object to the right of the "``in``" can be any object that supports iteration. Basically, if it can be thought of as a group of things, you can probably loop over it. In addition to lists, we can iterate over the elements of a tuple:
# + papermill={"duration": 0.033067, "end_time": "2021-06-04T13:07:05.135664", "exception": false, "start_time": "2021-06-04T13:07:05.102597", "status": "completed"} tags=[]
multiplicands = (2, 2, 2, 3, 3, 5)
product = 1
for mult in multiplicands:
product = product * mult
product
# + [markdown] papermill={"duration": 0.02135, "end_time": "2021-06-04T13:07:05.176223", "exception": false, "start_time": "2021-06-04T13:07:05.154873", "status": "completed"} tags=[]
# You can even loop through each character in a string:
# + papermill={"duration": 0.030704, "end_time": "2021-06-04T13:07:05.226782", "exception": false, "start_time": "2021-06-04T13:07:05.196078", "status": "completed"} tags=[]
s = 'steganograpHy is the practicE of conceaLing a file, message, image, or video within another fiLe, message, image, Or video.'
msg = ''
# print all the uppercase letters in s, one at a time
for char in s:
if char.isupper():
print(char, end='')
# + [markdown] papermill={"duration": 0.019434, "end_time": "2021-06-04T13:07:05.266644", "exception": false, "start_time": "2021-06-04T13:07:05.247210", "status": "completed"} tags=[]
# ### range()
#
# `range()` is a function that returns a sequence of numbers. It turns out to be very useful for writing loops.
#
# For example, if we want to repeat some action 5 times:
# + papermill={"duration": 0.031506, "end_time": "2021-06-04T13:07:05.319179", "exception": false, "start_time": "2021-06-04T13:07:05.287673", "status": "completed"} tags=[]
for i in range(5):
print("Doing important work. i =", i)
# + [markdown] papermill={"duration": 0.020551, "end_time": "2021-06-04T13:07:05.360531", "exception": false, "start_time": "2021-06-04T13:07:05.339980", "status": "completed"} tags=[]
# ## ``while`` loops
# The other type of loop in Python is a ``while`` loop, which iterates until some condition is met:
# + papermill={"duration": 0.0302, "end_time": "2021-06-04T13:07:05.411822", "exception": false, "start_time": "2021-06-04T13:07:05.381622", "status": "completed"} tags=[]
i = 0
while i < 10:
print(i, end=' ')
i += 1 # increase the value of i by 1
# + [markdown] papermill={"duration": 0.020397, "end_time": "2021-06-04T13:07:05.453144", "exception": false, "start_time": "2021-06-04T13:07:05.432747", "status": "completed"} tags=[]
# The argument of the ``while`` loop is evaluated as a boolean statement, and the loop is executed until the statement evaluates to False.
# + [markdown] papermill={"duration": 0.020523, "end_time": "2021-06-04T13:07:05.494524", "exception": false, "start_time": "2021-06-04T13:07:05.474001", "status": "completed"} tags=[]
# # List comprehensions
#
# List comprehensions are one of Python's most beloved and unique features. The easiest way to understand them is probably to just look at a few examples:
# + papermill={"duration": 0.030198, "end_time": "2021-06-04T13:07:05.545427", "exception": false, "start_time": "2021-06-04T13:07:05.515229", "status": "completed"} tags=[]
squares = [n**2 for n in range(10)]
squares
# + [markdown] papermill={"duration": 0.021028, "end_time": "2021-06-04T13:07:05.587657", "exception": false, "start_time": "2021-06-04T13:07:05.566629", "status": "completed"} tags=[]
# Here's how we would do the same thing without a list comprehension:
# + papermill={"duration": 0.031941, "end_time": "2021-06-04T13:07:05.640808", "exception": false, "start_time": "2021-06-04T13:07:05.608867", "status": "completed"} tags=[]
squares = []
for n in range(10):
squares.append(n**2)
squares
# + [markdown] papermill={"duration": 0.022868, "end_time": "2021-06-04T13:07:05.685761", "exception": false, "start_time": "2021-06-04T13:07:05.662893", "status": "completed"} tags=[]
# We can also add an `if` condition:
# + papermill={"duration": 0.030666, "end_time": "2021-06-04T13:07:05.738203", "exception": false, "start_time": "2021-06-04T13:07:05.707537", "status": "completed"} tags=[]
short_planets = [planet for planet in planets if len(planet) < 6]
short_planets
# + [markdown] papermill={"duration": 0.022689, "end_time": "2021-06-04T13:07:05.782735", "exception": false, "start_time": "2021-06-04T13:07:05.760046", "status": "completed"} tags=[]
# (If you're familiar with SQL, you might think of this as being like a "WHERE" clause)
#
# Here's an example of filtering with an `if` condition *and* applying some transformation to the loop variable:
# + papermill={"duration": 0.031425, "end_time": "2021-06-04T13:07:05.836336", "exception": false, "start_time": "2021-06-04T13:07:05.804911", "status": "completed"} tags=[]
# str.upper() returns an all-caps version of a string
loud_short_planets = [planet.upper() + '!' for planet in planets if len(planet) < 6]
loud_short_planets
# + [markdown] papermill={"duration": 0.022408, "end_time": "2021-06-04T13:07:05.881162", "exception": false, "start_time": "2021-06-04T13:07:05.858754", "status": "completed"} tags=[]
# People usually write these on a single line, but you might find the structure clearer when it's split up over 3 lines:
# + papermill={"duration": 0.032407, "end_time": "2021-06-04T13:07:05.936154", "exception": false, "start_time": "2021-06-04T13:07:05.903747", "status": "completed"} tags=[]
[
planet.upper() + '!'
for planet in planets
if len(planet) < 6
]
# + [markdown] papermill={"duration": 0.022777, "end_time": "2021-06-04T13:07:05.982812", "exception": false, "start_time": "2021-06-04T13:07:05.960035", "status": "completed"} tags=[]
# (Continuing the SQL analogy, you could think of these three lines as SELECT, FROM, and WHERE)
#
# The expression on the left doesn't technically have to involve the loop variable (though it'd be pretty unusual for it not to). What do you think the expression below will evaluate to? Press the 'output' button to check.
# + _kg_hide-output=true papermill={"duration": 0.03521, "end_time": "2021-06-04T13:07:06.041675", "exception": false, "start_time": "2021-06-04T13:07:06.006465", "status": "completed"} tags=[]
[32 for planet in planets]
# + [markdown] papermill={"duration": 0.022902, "end_time": "2021-06-04T13:07:06.087944", "exception": false, "start_time": "2021-06-04T13:07:06.065042", "status": "completed"} tags=[]
# List comprehensions combined with functions like `min`, `max`, and `sum` can lead to impressive one-line solutions for problems that would otherwise require several lines of code.
#
# For example, compare the following two cells of code that do the same thing.
#
# + papermill={"duration": 0.031786, "end_time": "2021-06-04T13:07:06.142796", "exception": false, "start_time": "2021-06-04T13:07:06.111010", "status": "completed"} tags=[]
def count_negatives(nums):
"""Return the number of negative numbers in the given list.
>>> count_negatives([5, -1, -2, 0, 3])
2
"""
n_negative = 0
for num in nums:
if num < 0:
n_negative = n_negative + 1
return n_negative
# + [markdown] papermill={"duration": 0.023078, "end_time": "2021-06-04T13:07:06.189266", "exception": false, "start_time": "2021-06-04T13:07:06.166188", "status": "completed"} tags=[]
# Here's a solution using a list comprehension:
# + papermill={"duration": 0.036906, "end_time": "2021-06-04T13:07:06.250610", "exception": false, "start_time": "2021-06-04T13:07:06.213704", "status": "completed"} tags=[]
def count_negatives(nums):
return len([num for num in nums if num < 0])
# + [markdown] papermill={"duration": 0.023068, "end_time": "2021-06-04T13:07:06.297650", "exception": false, "start_time": "2021-06-04T13:07:06.274582", "status": "completed"} tags=[]
# Much better, right?
#
# Well if all we care about is minimizing the length of our code, this third solution is better still!
# + papermill={"duration": 0.031353, "end_time": "2021-06-04T13:07:06.352923", "exception": false, "start_time": "2021-06-04T13:07:06.321570", "status": "completed"} tags=[]
def count_negatives(nums):
# Reminder: in the "booleans and conditionals" exercises, we learned about a quirk of
# Python where it calculates something like True + True + False + True to be equal to 3.
return sum([num < 0 for num in nums])
# + [markdown] papermill={"duration": 0.02343, "end_time": "2021-06-04T13:07:06.400260", "exception": false, "start_time": "2021-06-04T13:07:06.376830", "status": "completed"} tags=[]
# Which of these solutions is the "best" is entirely subjective. Solving a problem with less code is always nice, but it's worth keeping in mind the following lines from [The Zen of Python](https://en.wikipedia.org/wiki/Zen_of_Python):
#
# > Readability counts.
# > Explicit is better than implicit.
#
# So, use these tools to make compact readable programs. But when you have to choose, favor code that is easy for others to understand.
| Tutorials/5-loops-and-list-comprehensions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Scaffold hopping by holistic molecular representation in drug design
#
# #### <NAME>,* <NAME>
#
# ETH Zurich, Department of Chemistry and Applied Biosciences, RETHINK, Vladimir-Prelog-Weg 4, 8093, Zurich, Switzerland. <div>
# *Corresponding author: <EMAIL>
#
# ## Table of Contents
# 1. [Preliminary steps](#preliminary)
# 2. [Molecule import and pre-treatment](#import)
# 3. [WHALES descriptors](#whales) <div>
# a. [*WHALES calculation*](#calculation) <div>
# b. [*Descriptor scaling*](#scaling)
# 4. [Virtual screening](#vs)<div>
# a. [*Similarity calculation*](#similarity)<div>
# b. [*Identification of top hits*](#tophits)<div>
# 5. [Scaffold analysis](#scaffold)<div>
# a. [*Template scaffold*](#templatescaffold)<div>
# b. [*Library scaffold diversity*](#libraryscaffold)<div>
# c. [*Scaffold diversity of the top hits*](#tophitsscaffold)<div>
# 6. [How to cite](#cite)
# # Import of necessary packages <a name="preliminary"></a>
#
# Here you will find all of the necessary packages that will be used in this Jupyter Notebook. No worries, they should all be installed already in your virtual environment!
# chemistry toolkits
from rdkit import Chem
from rdkit.Chem import AllChem
from rdkit.Chem import Draw # for molecule depiction
from rdkit.Chem.Draw import IPythonConsole
from rdkit.Chem.Scaffolds import MurckoScaffold
# WHALES-related code
import ChemTools as tools # for molecule pretreatment
import do_whales # importing WHALES descriptors
from ChemTools import prepare_mol_from_sdf # to pretreat the virtual screening library
# for data analysis and plotting
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.metrics.pairwise import euclidean_distances
import numpy
# to delete before uploading
# %load_ext autoreload
# %autoreload
# %reload_ext ChemTools
# %reload_ext do_whales
# # Molecule import and pre-treatment <a name="import"></a>
#
# ## Template molecule
# Import and visualization of galantamine, starting from SMILES
# import template from SMILES
template = Chem.MolFromSmiles('C[C@@H]1CC[C@@H]2[C@@H]1[C@@H]([C@@]3(C[C@H]([C@]2(O3)C)OC(=O)CO)C(C)C)OC(=O)/C=C/c4ccccc4') # imports from SMILES
# add 2D coordinates for visualization
err = AllChem.Compute2DCoords(template)
template
# Prepare the molecule, by checking for errors and optimizing the 3D conformation
mol, err = tools.prepare_mol(template)
mol
# Inspect the partial charges that will be used for WHALES calculation
tools.do_map(template,lab_atom=True)
writer = Chem.rdmolfiles.SDWriter('out.sdf')
writer.write(mol)
# ## Compound library
# Import and prepare library
# set the path to the sdf file for input and output
input_filename = '../data/library.sdf'
vs_library_2D = Chem.SDMolSupplier(input_filename) # conserved for 2D representations
vs_library = prepare_mol_from_sdf(input_filename) # computes 3D geometry from a specified sdf file
# Visually inspect the database after geometry optimization
#Visualize the first 6 molecules of the database after geometry optimization
number_mol = 6 # number of molecules to inspect
Draw.MolsToGridImage(vs_library[:number_mol],molsPerRow=3,subImgSize=(100,100),legends=[x.GetProp("_Name") for x in vs_library[:number_mol]])
# # WHALES descriptors <a name="whales"></a>
#
# ## Descriptors calculation <a name="calculation"></a>
# Compute WHALES descriptors for the template molecule
# compute descriptors as an array
whales_template, lab = do_whales.whales_from_mol(template)
# convert the arrays into a pandas dataframe
df_whales_template = pd.DataFrame(whales_template.reshape(-1, len(whales_template)),index=['template'],columns=lab)
df_whales_template
# Compute WHALES for the virtual screening library
whales_library = []
for mol in vs_library: # runs over the library and updates WHALES
whales_temp, lab = do_whales.whales_from_mol(mol)
whales_library.append(whales_temp)
# convert the arrays into a pandas dataframe
df_whales_library = pd.DataFrame(whales_library,columns=lab)
df_whales_library.head() # library preview
# ## Descriptors scaling <a name="scaling"></a>
# ### Data visualization
# Visualizing the raw descriptor values for the virtual screening library using a boxplot
sns.set(rc={'figure.figsize':(16,8.27)}) # sets the size of the boxplot
sns.boxplot(data=df_whales_library,linewidth=2)
# ### Autoscaling
# Virtual library scaling
aver = df_whales_library.mean()
sdv = df_whales_library.std()
df_whales_library_scaled = (df_whales_library - aver)/sdv
df_whales_library_scaled.to_csv('WHALES_after.csv')
# Template scaling
df_whales_template_scaled = (df_whales_template - aver)/sdv
df_whales_template_scaled
# ### Data visualization
# Visualizing the descriptor values for the virtual screening library after autoscaling
#
sns.set(rc={'figure.figsize':(16,8.27)})
sns.boxplot(data=df_whales_library_scaled,linewidth=2)
# # Virtual screening <a name="vs"></a>
#
# ## Similarity calculation <a name="similarity"></a>
# compute Euclidean distance
D = euclidean_distances(df_whales_template_scaled,df_whales_library_scaled)
# ## Identification of top hits <a name="tophits"></a>
# Selection of the 10 closest compounds based on the computed distance on WHALES
# sorting based on distance
sort_index = numpy.argsort(D) # index for sorting according to D
D_neig = D[:,sort_index] # sorted distance
# selection of the 10 closest compounds
k = 10 # number of compounds to choose
neighbor_ID = sort_index[:,0:k]
neighbor_ID
# +
# display of the 10 closest compounds
hits = []
smiles_hits = []
for j in numpy.nditer(neighbor_ID):
hits.append(vs_library_2D[int(j)])
smiles_hits.append(Chem.MolToSmiles(mol))
#Visualize the first 10 molecules of the database after geometry optimization
number_mol = 10 # number of molecules to inspect
Draw.MolsToGridImage(hits[:number_mol+1],molsPerRow=5,subImgSize=(300,300),legends=[x.GetProp("_Name") for x in hits[:number_mol+1]])
# -
# # Scaffold analysis <a name="scaffold"></a>
#
# ## Template scaffold <a name="templatescaffold"></a>
core = MurckoScaffold.GetScaffoldForMol(template)
core
# ## Scaffold diversity of the virtual screening library
# Compute the Bemis-Murcko scaffolds for the virtual screening library
scaffold_vs = [] # generates an empty supplier to contain the computed scaffold
for mol in vs_library_2D:
scaffold_vs.append(MurckoScaffold.GetScaffoldForMol(mol))
# Preview of the scaffolds (4 for display)
# 4 scaffolds are displayed
k = 4
Draw.MolsToGridImage(scaffold_vs[:k],molsPerRow=2,subImgSize=(200,200),legends=[x.GetProp("_Name") for x in scaffold_vs[:k]])
# Most frequently recurring scaffolds in the virtual screening library
freq_scaffolds_library = tools.frequent_scaffolds(vs_library_2D) # contains the list of the (unique) scaffolds, sorted by number
# Display the four most occurring scaffolds
k = 4
Draw.MolsToGridImage(freq_scaffolds_library[:k],molsPerRow=2,subImgSize=(200,200),legends=[x.GetProp("_Name") for x in freq_scaffolds_library[:k]])
# Computes the relative scaffold diversity of the library
SD_rel = len(freq_scaffolds_library)/len(vs_library)*100
print(SD_rel)
# ## Top hits scaffolds
# +
scaffold_hits = []
for mol in hits:
scaffold_hits.append(MurckoScaffold.GetScaffoldForMol(mol))
Draw.MolsToGridImage(scaffold_hits[:10],molsPerRow=2,subImgSize=(200,200),legends=[x.GetProp("_Name") for x in scaffold_hits[:10]])
# -
# Computes the frequent scaffolds and the scaffold diversity
freq_scaffolds_hits = tools.frequent_scaffolds(hits) # contains the list of the (unique) scaffolds, sorted by number
k = len(freq_scaffolds_hits) # display all scaffolds
Draw.MolsToGridImage(freq_scaffolds_hits[:k],molsPerRow=2,subImgSize=(200,200),legends=[x.GetProp("_Name") for x in freq_scaffolds_hits[:k]])
SD_rel = len(freq_scaffolds_hits)/len(hits)*100
print(SD_rel)
| src/python/scaffold_hopping_whales/code/virtual_screening_pipeline.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: julia-1.5
# kernelspec:
# argv:
# - C:\Users\nicho\AppData\Local\Programs\Julia 1.5.3\bin\julia.exe
# - -i
# - --startup-file=yes
# - --color=yes
# - --project=@.
# - C:\Users\nicho\.julia\packages\IJulia\a1SNk\src\kernel.jl
# - '{connection_file}'
# display_name: Julia 1.5.3
# env: {}
# interrupt_mode: message
# language: julia
# name: julia-1.5
# ---
# + [markdown] nteract={"transient": {"deleting": false}}
# ## Computing a ϕ(ρz) curve using a Monte Carlo
# This notebook compares a simple model of the ionization cross-section with the more sophisticated model of Bote & Salvat by modeling a ϕ(ρz) curve using Monte Carlo simulation.
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
using NeXLCore, StaticArrays
using StatsBase
# + [markdown] nteract={"transient": {"deleting": false}}
# Use the standard crude model of the ionization cross-section used for most ϕ(ρz) models.
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
m(sh) = 0.86 + 0.12*exp(-(z(element(sh))/5)^2)
sigma(e, sh) = e>energy(sh) ? log(e/energy(sh))/ (((e/energy(sh))^m(sh))*energy(sh)^2) : 0.0
# + [markdown] nteract={"transient": {"deleting": false}}
# Also consider, the Bote 2009 model for the ionization cross-section.
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
sigma2(e,sh) = ionizationcrosssection(sh, e)
# + [markdown] nteract={"transient": {"deleting": false}}
# Construct a simple accumlator to accumulate the relative number of ionizations per unit depth.
# + [markdown] nteract={"transient": {"deleting": false}}
# Define the model parameters. Let's consider the O K-L3 transition in Albite at 10 keV.
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
using NeXLMatrixCorrection
# Sample material, beam energy, characteristic X-ray
mat, e0, cxr = material("NaAlSi3O8",1.0), 15.0e3, n"O K-L3"
# Shell and take-off angle
sh, toa = inner(cxr), deg2rad(40.0)
# reduced mass absorption coefficient
xi = χ(mat, cxr, toa)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
randbetween(a::T,b::T) where {T<:Real} = a+(b-a)*rand(T)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
width = 1.0e-5 # Accumulator bin width
rz=collect(0.0:width:60*width)
prz, prza = Histogram(rz,Float64), Histogram(rz,Float64)
prz2, prza2 = Histogram(rz,Float64), Histogram(rz,Float64)
traj = 10000
# Normalize relative to a thin film of width
norm, norm2 = sigma(e0, sh)*traj*width, sigma2(e0,sh)*traj*width
for i in 1:traj
# trajectory(...) excutes one full electron trajectory. It stops at each scattering
# point and runs the `do` code.
trajectory(gun(Electron, e0, 1.0e-6), bulk(mat), minE=energy(inner(cxr))) do part, reg
# Pick a randomized depth within the trajectory segment
z = randbetween(part.current[3], part.previous[3])
v = sigma(part.energy, sh)*NeXLCore.pathlength(part)/norm
push!(prz, z, v)
push!(prza, z, v*exp(-xi*z))
v2 = sigma2(part.energy, sh)*NeXLCore.pathlength(part)/norm2
push!(prz2, z, v2)
push!(prza2, z, v2*exp(-xi*z))
end
end
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
using Gadfly
set_default_plot_size(8inch,6inch)
mca = matrixcorrection(XPhi, mat, sh, e0)
plot(
layer(x=rz[1:end-1], y=[ ϕ(mca, ρz) for ρz in rz[1:end-1] ], Geom.line,Theme( default_color="blue")),
layer(x=rz[1:end-1], y=[ ϕabs(mca, ρz, cxr, toa) for ρz in rz[1:end-1] ], Geom.line,Theme( default_color="lightblue")),
layer(x=rz[1:end-1], y=prz.weights, Geom.line,Theme( default_color="red")),
layer(x=rz[1:end-1], y=prza.weights, Geom.line,Theme( default_color="pink")),
layer(x=rz[1:end-1], y=prz2.weights, Geom.line,Theme( default_color="green")),
layer(x=rz[1:end-1], y=prza2.weights, Geom.line,Theme( default_color="lightgreen")),
Guide.manual_color_key("Model",["XPhi","XPhi(absorbed)", "Simple","Simple(absorbed)","Bote2009","Bote2009(absorbed)"],["blue","lightblue", "red","pink","green","lightgreen",]), Guide.xlabel("ρz"), Guide.ylabel("ϕ(ρz)"))
# + [markdown] nteract={"transient": {"deleting": false}}
# The A-term in the ZAF expression is just the ratio of the absorbed over the generated.
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
sum(prza.weights) / sum(prz.weights), sum(prza2.weights) / sum(prz2.weights), A(mca, cxr,toa)
| jupyter/phirhoz_mc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Project: Train a Quadcopter How to Fly
#
# Design an agent to fly a quadcopter, and then train it using a reinforcement learning algorithm of your choice!
#
# Try to apply the techniques you have learnt, but also feel free to come up with innovative ideas and test them.
# ## Instructions
#
# Take a look at the files in the directory to better understand the structure of the project.
#
# - `task.py`: Define your task (environment) in this file.
# - `agents/`: Folder containing reinforcement learning agents.
# - `policy_search.py`: A sample agent has been provided here.
# - `agent.py`: Develop your agent here.
# - `physics_sim.py`: This file contains the simulator for the quadcopter. **DO NOT MODIFY THIS FILE**.
#
# For this project, you will define your own task in `task.py`. Although we have provided a example task to get you started, you are encouraged to change it. Later in this notebook, you will learn more about how to amend this file.
#
# You will also design a reinforcement learning agent in `agent.py` to complete your chosen task.
#
# You are welcome to create any additional files to help you to organize your code. For instance, you may find it useful to define a `model.py` file defining any needed neural network architectures.
#
# ## Controlling the Quadcopter
#
# We provide a sample agent in the code cell below to show you how to use the sim to control the quadcopter. This agent is even simpler than the sample agent that you'll examine (in `agents/policy_search.py`) later in this notebook!
#
# The agent controls the quadcopter by setting the revolutions per second on each of its four rotors. The provided agent in the `Basic_Agent` class below always selects a random action for each of the four rotors. These four speeds are returned by the `act` method as a list of four floating-point numbers.
#
# For this project, the agent that you will implement in `agents/agent.py` will have a far more intelligent method for selecting actions!
# +
import random
class Basic_Agent():
def __init__(self, task):
self.task = task
def act(self):
new_thrust = random.gauss(450., 25.)
return [new_thrust + random.gauss(0., 1.) for x in range(4)]
# -
# Run the code cell below to have the agent select actions to control the quadcopter.
#
# Feel free to change the provided values of `runtime`, `init_pose`, `init_velocities`, and `init_angle_velocities` below to change the starting conditions of the quadcopter.
#
# The `labels` list below annotates statistics that are saved while running the simulation. All of this information is saved in a text file `data.txt` and stored in the dictionary `results`.
# +
# %load_ext autoreload
# %autoreload 2
import csv
import numpy as np
from task import Task
from agents.agent import DDPG
# Modify the values below to give the quadcopter a different starting position.
runtime = 5. # time limit of the episode
init_pose = np.array([0., 0., 10., 0., 0., 0.]) # initial pose
init_velocities = np.array([0., 0., 0.]) # initial velocities
init_angle_velocities = np.array([0., 0., 0.]) # initial angle velocities
file_output = 'data.txt' # file name for saved results
# Setup
task = Task(init_pose, init_velocities, init_angle_velocities, runtime)
agent = Basic_Agent(task)
done = False
labels = ['time', 'x', 'y', 'z', 'phi', 'theta', 'psi', 'x_velocity',
'y_velocity', 'z_velocity', 'phi_velocity', 'theta_velocity',
'psi_velocity', 'rotor_speed1', 'rotor_speed2', 'rotor_speed3', 'rotor_speed4']
results = {x : [] for x in labels}
# Run the simulation, and save the results.
with open(file_output, 'w') as csvfile:
writer = csv.writer(csvfile)
writer.writerow(labels)
while True:
rotor_speeds = agent.act()
_, _, done = task.step(rotor_speeds)
to_write = [task.sim.time] + list(task.sim.pose) + list(task.sim.v) + list(task.sim.angular_v) + list(rotor_speeds)
for ii in range(len(labels)):
results[labels[ii]].append(to_write[ii])
writer.writerow(to_write)
if done:
break
# +
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# #%matplotlib notebook # Change to %matplotlib inline if dont work
# %matplotlib inline
fig = plt.figure(figsize = (14,8))
ax = fig.add_subplot(111, projection='3d')
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_zlabel('Z')
ax.scatter(results['x'], results['y'], results['z'])
# -
# Run the code cell below to visualize how the position of the quadcopter evolved during the simulation.
# +
import matplotlib.pyplot as plt
# %matplotlib inline
plt.plot(results['time'], results['x'], label='x')
plt.plot(results['time'], results['y'], label='y')
plt.plot(results['time'], results['z'], label='z')
plt.legend()
_ = plt.ylim()
# -
# The next code cell visualizes the velocity of the quadcopter.
plt.plot(results['time'], results['x_velocity'], label='x_hat')
plt.plot(results['time'], results['y_velocity'], label='y_hat')
plt.plot(results['time'], results['z_velocity'], label='z_hat')
plt.legend()
_ = plt.ylim()
# Next, you can plot the Euler angles (the rotation of the quadcopter over the $x$-, $y$-, and $z$-axes),
plt.plot(results['time'], results['phi'], label='phi')
plt.plot(results['time'], results['theta'], label='theta')
plt.plot(results['time'], results['psi'], label='psi')
plt.legend()
_ = plt.ylim()
# before plotting the velocities (in radians per second) corresponding to each of the Euler angles.
plt.plot(results['time'], results['phi_velocity'], label='phi_velocity')
plt.plot(results['time'], results['theta_velocity'], label='theta_velocity')
plt.plot(results['time'], results['psi_velocity'], label='psi_velocity')
plt.legend()
_ = plt.ylim()
# Finally, you can use the code cell below to print the agent's choice of actions.
plt.plot(results['time'], results['rotor_speed1'], label='Rotor 1 revolutions / second')
plt.plot(results['time'], results['rotor_speed2'], label='Rotor 2 revolutions / second')
plt.plot(results['time'], results['rotor_speed3'], label='Rotor 3 revolutions / second')
plt.plot(results['time'], results['rotor_speed4'], label='Rotor 4 revolutions / second')
plt.legend()
_ = plt.ylim()
# When specifying a task, you will derive the environment state from the simulator. Run the code cell below to print the values of the following variables at the end of the simulation:
# - `task.sim.pose` (the position of the quadcopter in ($x,y,z$) dimensions and the Euler angles),
# - `task.sim.v` (the velocity of the quadcopter in ($x,y,z$) dimensions), and
# - `task.sim.angular_v` (radians/second for each of the three Euler angles).
# the pose, velocity, and angular velocity of the quadcopter at the end of the episode
print(task.sim.pose)
print(task.sim.v)
print(task.sim.angular_v)
# In the sample task in `task.py`, we use the 6-dimensional pose of the quadcopter to construct the state of the environment at each timestep. However, when amending the task for your purposes, you are welcome to expand the size of the state vector by including the velocity information. You can use any combination of the pose, velocity, and angular velocity - feel free to tinker here, and construct the state to suit your task.
#
# ## The Task
#
# A sample task has been provided for you in `task.py`. Open this file in a new window now.
#
# The `__init__()` method is used to initialize several variables that are needed to specify the task.
# - The simulator is initialized as an instance of the `PhysicsSim` class (from `physics_sim.py`).
# - Inspired by the methodology in the original DDPG paper, we make use of action repeats. For each timestep of the agent, we step the simulation `action_repeats` timesteps. If you are not familiar with action repeats, please read the **Results** section in [the DDPG paper](https://arxiv.org/abs/1509.02971).
# - We set the number of elements in the state vector. For the sample task, we only work with the 6-dimensional pose information. To set the size of the state (`state_size`), we must take action repeats into account.
# - The environment will always have a 4-dimensional action space, with one entry for each rotor (`action_size=4`). You can set the minimum (`action_low`) and maximum (`action_high`) values of each entry here.
# - The sample task in this provided file is for the agent to reach a target position. We specify that target position as a variable.
#
# The `reset()` method resets the simulator. The agent should call this method every time the episode ends. You can see an example of this in the code cell below.
#
# The `step()` method is perhaps the most important. It accepts the agent's choice of action `rotor_speeds`, which is used to prepare the next state to pass on to the agent. Then, the reward is computed from `get_reward()`. The episode is considered done if the time limit has been exceeded, or the quadcopter has travelled outside of the bounds of the simulation.
#
# In the next section, you will learn how to test the performance of an agent on this task.
# ## The Agent
#
# The sample agent given in `agents/policy_search.py` uses a very simplistic linear policy to directly compute the action vector as a dot product of the state vector and a matrix of weights. Then, it randomly perturbs the parameters by adding some Gaussian noise, to produce a different policy. Based on the average reward obtained in each episode (`score`), it keeps track of the best set of parameters found so far, how the score is changing, and accordingly tweaks a scaling factor to widen or tighten the noise.
#
# Run the code cell below to see how the agent performs on the sample task.
# +
import sys
import pandas as pd
from agents.policy_search import PolicySearch_Agent
from task import Task
num_episodes = 1000
target_pos = np.array([0., 0., 10.])
task = Task(target_pos=target_pos)
agent = PolicySearch_Agent(task)
for i_episode in range(1, num_episodes+1):
state = agent.reset_episode() # start a new episode
while True:
action = agent.act(state)
next_state, reward, done = task.step(action)
agent.step(reward, done)
state = next_state
if done:
print("\rEpisode = {:4d}, score = {:7.3f} (best = {:7.3f}), noise_scale = {}".format(
i_episode, agent.score, agent.best_score, agent.noise_scale), end="") # [debug]
break
sys.stdout.flush()
# -
# This agent should perform very poorly on this task. And that's where you come in!
# ## Define the Task, Design the Agent, and Train Your Agent!
#
# Amend `task.py` to specify a task of your choosing. If you're unsure what kind of task to specify, you may like to teach your quadcopter to takeoff, hover in place, land softly, or reach a target pose.
#
# After specifying your task, use the sample agent in `agents/policy_search.py` as a template to define your own agent in `agents/agent.py`. You can borrow whatever you need from the sample agent, including ideas on how you might modularize your code (using helper methods like `act()`, `learn()`, `reset_episode()`, etc.).
#
# Note that it is **highly unlikely** that the first agent and task that you specify will learn well. You will likely have to tweak various hyperparameters and the reward function for your task until you arrive at reasonably good behavior.
#
# As you develop your agent, it's important to keep an eye on how it's performing. Use the code above as inspiration to build in a mechanism to log/save the total rewards obtained in each episode to file. If the episode rewards are gradually increasing, this is an indication that your agent is learning.
# +
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
# Make sure to change from notebook to inline after your tests
# %matplotlib notebook
import time
class AnimatedPlot():
def __init__(self):
"""Initialize parameters"""
self.X, self.Y, self.Z = [], [], []
self.fig = plt.figure(figsize = (14,8))
self.ax = self.fig.add_subplot(111, projection='3d')
def plot(self, task, i_episode=None):
pose = task.sim.pose[:3]
self.X.append(pose[0])
self.Y.append(pose[1])
self.Z.append(pose[2])
self.ax.clear()
if i_episode:
plt.title("Episode {}".format(i_episode))
if len(self.X)>1:
self.ax.scatter(self.X[:-1], self.Y[:-1], self.Z[:-1], c='k', alpha=0.3)
if task.sim.done and task.sim.runtime > task.sim.time:
# Colision
self.ax.scatter(pose[0], pose[1], pose[2], c='r', marker='*', linewidths=5)
else:
self.ax.scatter(pose[0], pose[1], pose[2], c='k', marker='s', linewidths=5)
self.fig.canvas.draw()
time.sleep(0.5)
# +
## TODO: Train your agent here.
import sys
import pandas as pd
import numpy as np
from agents.policy_search import PolicySearch_Agent
from agents.agent import DDPG
from task import Task
#from agents.agent import AnimatedPlot
num_episodes = 10
target_pos = np.array([0., 0., 20.])
task = Task(target_pos=target_pos)
agent = DDPG(task)
myplot = AnimatedPlot()
for i_episode in range(1, num_episodes+1):
state = agent.reset_episode() # start a new episode
score = []
while True:
action = agent.act(state)
next_state, reward, done = task.step(action)
agent.step(action, reward, next_state, done)
state = next_state
score += reward
myplot.plot(task)
if done:
print("The reward after {} episode is {}".format(i_episode, agent.total_reward))
break
# +
import sys
import pandas as pd
import numpy as np
from agents.policy_search import PolicySearch_Agent
from agents.agent import DDPG
from task import Task
#from agents.agent import AnimatedPlot
num_episodes = 1000
target_pos = np.array([0., 0., 20.])
task = Task(target_pos=target_pos)
agent = DDPG(task)
#myplot = AnimatedPlot()
for i_episode in range(1, num_episodes+1):
state = agent.reset_episode() # start a new episode
score = []
while True:
action = agent.act(state)
next_state, reward, done = task.step(action)
agent.step(action, reward, next_state, done)
state = next_state
score += reward
#myplot.plot(task)
if done:
print("The reward after {} episode is {}".format(i_episode, agent.total_reward))
break
# -
# ## Plot the Rewards
#
# Once you are satisfied with your performance, plot the episode rewards, either from a single run, or averaged over multiple runs.
# +
import numpy as np
import pandas as pd
import seaborn as sns
# %matplotlib inline
def map_function(reward_function, x, y, target_pos):
R = pd.DataFrame(np.zeros([len(x), len(y)]), index=y, columns=x)
for xx in x:
for yy in y:
R[xx][yy] = reward_function([xx, yy], target_pos)
return R
reward_function = lambda pose, target_pos: 0. + 10 -.3*(np.tanh(abs(pose - target_pos))**2).sum()
x_range = np.round(np.arange(-10.0,10,0.1), 2)
z_range = np.round(np.arange(20,0,-0.1), 2)
target_pos = np.array([0, 10])
R = map_function(reward_function, x_range, z_range, target_pos)
ax = sns.heatmap(R)
ax.set_xlabel("Position X-axis")
ax.set_ylabel("Position Z-axis")
plt.show()
# -
import matplotlib.pyplot as plt
# %matplotlib inline
plt.plot(np.array(agent.rewards))
plt.xlabel('Episodes')
plt.ylabel('Score')
plt.title("Quadcopter Takeoff")
plt.show()
np.mean(agent.rewards[-50:])
# ## Reflections
#
# **Question 1**: Describe the task that you specified in `task.py`. How did you design the reward function?
#
# **Answer**:
#
# The task I specified in is takeoff, specifically the quadcopter has an initial position of x,y,z : (0, 0, 0) and a target position of x,y,z : (0, 0, 10). it means initally the z:0 which means it is in ground it will take off from z:0 to z:10 i.e 10 units above.
#
# The reward Function Contains following points:
#
# --> Calculated the coordinate distance from the target position and added a penalty of porduct based on how far it ended up from the target position.
#
# --> The reward function should be normalized between -1 and 1 for this np.tanh() is used. This avoids instability in training due to exploding gradients.
#
# --> Added a bonus of 5 points for it successfully taking off and not crashing
# **Question 2**: Discuss your agent briefly, using the following questions as a guide:
#
# - What learning algorithm(s) did you try? What worked best for you?
# - What was your final choice of hyperparameters (such as $\alpha$, $\gamma$, $\epsilon$, etc.)?
# - What neural network architecture did you use (if any)? Specify layers, sizes, activation functions, etc.
#
# **Answer**:
#
# * Deep Deterministic Policy Gradient (DDPG) algorithm is the best one among other algorithm as quadcopter control problem has a continuous action space which is ideally handle by DDPG actor-Critic alogrithum.
# * The whole combination of buffer function, actor, crtic, DDPG and Noise function worked best for me.
#
# Following are the hyperparameters which is used.
#
# * gamma = 0.99
# * tau = 0.01
# * exploration_mu = 0
# * exploration_theta = 0.15
# * exploration_sigma = 0.2
#
# Neural network architecture which i used are:
# * Dense layer seprated by batch normalization to increase stability
# * "Relu" activation Function for all layer and final layer "Sigmoid" function for output.
#
# **Question 3**: Using the episode rewards plot, discuss how the agent learned over time.
#
# - Was it an easy task to learn or hard?
# - Was there a gradual learning curve, or an aha moment?
# - How good was the final performance of the agent? (e.g. mean rewards over the last 10 episodes)
#
# **Answer**:
#
# * Using the episode rewards plot initially the agent was learning quite well with good reward score.
# * It was a gradual learning curve in which no of episode are given high as we can determine better rewards and teach quadcopter more efficiently.
# * the final performance of the agent are quite good with mean of 1174 for last 50 episodes.
# **Question 4**: Briefly summarize your experience working on this project. You can use the following prompts for ideas.
#
# - What was the hardest part of the project? (e.g. getting started, plotting, specifying the task, etc.)
# - Did you find anything interesting in how the quadcopter or your agent behaved?
#
# **Answer**:
#
# The Project was definitely the hardest project i worked on any nanodegree so far as it contains the mixture of maths, coding phycis, deep nural network and many more. bascilly it was really hard get started with the project and understand the key things which is expected by the project and work on that. DDPG algorithm given in the project gave me quite good understanding on how to work on this project.
#
# The intresting thing which i find in the project was declaring the reward function as with slight change in the function causing significant change in the score and the animated simulation of my quadcopter gave me a better understanding how this going to work in practical.
#
# with the only one task defined (i.e. takeoff) understand few possiblities of quadcopter but will surely want to try other possiblities also like hover, landing and many more.
| Train a quadcopter to fly/Quadcopter_Project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] nbsphinx="hidden"
# This notebook is part of the $\omega radlib$ documentation: https://docs.wradlib.org.
#
# Copyright (c) $\omega radlib$ developers.
# Distributed under the MIT License. See LICENSE.txt for more info.
# -
# # xarray CfRadial2 backend
#
# In this example, we read CfRadial2 data files using the xarray `cfradial2` backend.
import wradlib as wrl
import warnings
warnings.filterwarnings('ignore')
import matplotlib.pyplot as pl
import numpy as np
import xarray as xr
try:
get_ipython().magic("matplotlib inline")
except:
pl.ion()
# ## Load CfRadial1 Volume Data
fpath = 'netcdf/cfrad.20080604_002217_000_SPOL_v36_SUR_cfradial2.nc'
f = wrl.util.get_wradlib_data_file(fpath)
vol = wrl.io.open_cfradial2_dataset(f)
# ### Inspect RadarVolume
display(vol)
# ### Inspect root group
#
# The `sweep` dimension contains the number of scans in this radar volume. Further the dataset consists of variables (location coordinates, time_coverage) and attributes (Conventions, metadata).
vol.root
# ### Inspect sweep group(s)
#
# The sweep-groups can be accessed via their respective keys. The dimensions consist of `range` and `time` with added coordinates `azimuth`, `elevation`, `range` and `time`. There will be variables like radar moments (DBZH etc.) and sweep-dependend metadata (like `fixed_angle`, `sweep_mode` etc.).
display(vol[0])
# ### Goereferencing
swp = vol[0].copy().pipe(wrl.georef.georeference_dataset)
# ### Plotting
swp.DBZ.plot.pcolormesh(x='x', y='y')
pl.gca().set_aspect('equal')
fig = pl.figure(figsize=(10,10))
swp.DBZ.wradlib.plot_ppi(proj='cg', fig=fig)
# +
import cartopy
import cartopy.crs as ccrs
import cartopy.feature as cfeature
map_trans = ccrs.AzimuthalEquidistant(central_latitude=swp.latitude.values,
central_longitude=swp.longitude.values)
# -
map_proj = ccrs.AzimuthalEquidistant(central_latitude=swp.latitude.values,
central_longitude=swp.longitude.values)
pm = swp.DBZ.wradlib.plot_ppi(proj=map_proj)
ax = pl.gca()
ax.gridlines(crs=map_proj)
print(ax)
map_proj = ccrs.Mercator(central_longitude=swp.longitude.values)
fig = pl.figure(figsize=(10,8))
ax = fig.add_subplot(111, projection=map_proj)
pm = swp.DBZ.wradlib.plot_ppi(ax=ax)
ax.gridlines(draw_labels=True)
# +
import cartopy.feature as cfeature
def plot_borders(ax):
borders = cfeature.NaturalEarthFeature(category='physical',
name='coastline',
scale='10m',
facecolor='none')
ax.add_feature(borders, edgecolor='black', lw=2, zorder=4)
map_proj = ccrs.Mercator(central_longitude=swp.longitude.values)
fig = pl.figure(figsize=(10,8))
ax = fig.add_subplot(111, projection=map_proj)
DBZ = swp.DBZ
pm = DBZ.where(DBZ > 0).wradlib.plot_ppi(ax=ax)
plot_borders(ax)
ax.gridlines(draw_labels=True)
# +
import matplotlib.path as mpath
theta = np.linspace(0, 2*np.pi, 100)
center, radius = [0.5, 0.5], 0.5
verts = np.vstack([np.sin(theta), np.cos(theta)]).T
circle = mpath.Path(verts * radius + center)
map_proj = ccrs.AzimuthalEquidistant(central_latitude=swp.latitude.values,
central_longitude=swp.longitude.values,
)
fig = pl.figure(figsize=(10,8))
ax = fig.add_subplot(111, projection=map_proj)
ax.set_boundary(circle, transform=ax.transAxes)
pm = swp.DBZ.wradlib.plot_ppi(proj=map_proj, ax=ax)
ax = pl.gca()
ax.gridlines(crs=map_proj)
# -
fig = pl.figure(figsize=(10, 8))
proj=ccrs.AzimuthalEquidistant(central_latitude=swp.latitude.values,
central_longitude=swp.longitude.values)
ax = fig.add_subplot(111, projection=proj)
pm = swp.DBZ.wradlib.plot_ppi(ax=ax)
ax.gridlines()
swp.DBZ.wradlib.plot_ppi()
# ### Inspect radar moments
#
# The DataArrays can be accessed by key or by attribute. Each DataArray has dimensions and coordinates of it's parent dataset. There are attributes connected which are defined by Cf/Radial standard.
display(swp.DBZ)
# ### Create simple plot
#
# Using xarray features a simple plot can be created like this. Note the `sortby('time')` method, which sorts the radials by time.
swp.DBZ.sortby('rtime').plot(x="range", y="rtime", add_labels=False)
fig = pl.figure(figsize=(5,5))
pm = swp.DBZ.wradlib.plot_ppi(proj={'latmin': 33e3}, fig=fig)
# ### Mask some values
swp['DBZ'] = swp['DBZ'].where(swp['DBZ'] >= 0)
swp['DBZ'].plot()
# ### Export to ODIM and CfRadial2
vol.to_odim('cfradial2_as_odim.h5')
vol.to_cfradial2('cfradial2_as_cfradial2.nc')
# ### Import again
vola = wrl.io.open_odim_dataset('cfradial2_as_odim.h5')
volb = wrl.io.open_cfradial2_dataset('cfradial2_as_cfradial2.nc')
# ### Check equality
#
# Some variables need to be dropped, since they are not exported to the other standards or differ slightly (eg. re-indexed ray times).
drop = set(vol[0]) ^ set(vola[0]) | set({"elevation", "rtime"})
xr.testing.assert_allclose(vol.root, vola.root)
xr.testing.assert_allclose(vol[0].drop_vars(drop), vola[0].drop_vars(drop, errors="ignore"))
xr.testing.assert_allclose(vol.root, volb.root)
xr.testing.assert_equal(vol[0], volb[0])
xr.testing.assert_allclose(vola.root, volb.root)
xr.testing.assert_allclose(vola[0].drop_vars(drop, errors="ignore"), volb[0].drop_vars(drop, errors="ignore"))
# ## More CfRadial2 loading mechanisms
# ### Use `xr.open_dataset` to retrieve explicit group
#
swp = xr.open_dataset(f, engine="cfradial2", group="sweep_8")
display(swp)
| notebooks/fileio/wradlib_cfradial2_backend.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 과거 주식 가격 데이터 확보를 위해 지금까지 삽질해 본 것들
#
# 요즘은 집에서 컴퓨터에 설치된 HTS 를 통해 주식을 거래하는 것을 넘어서 스마트폰으로 언제든지 어디서든 주식 가격을 확인하고 거래하는게 가능한 세상이다.
# 또한 각종 증권사들 뿐만 아니라 다른 다양한 곳에서, 예를 들어 트레이딩뷰 같은 곳에서 제공하는 여러 기본 보조지표들을 포함하는 강력한 차트 툴 처럼,
# 주식 가격 분석에 활용 가능한 여러 기능들을 탑재한 프로그램들을 제공하고 있어 누구나 쉽게 기본적인 가격 데이터 분석을 시작해 볼 수 있다.
#
# 하지만 그렇게 제공되는 기능들로는 만족하지 못하고, 대신 자신이 직접 나름대로의 방법으로 데이터를 분석해보고 싶은 사람들은,
# 아마도 가장 첫번째 난관으로 **주식 가격 데이터를 어떻게 확보해야할지** 부터 고민하게 되지 않을까 생각한다.
# 그리고 이 문서는 그렇게 나와 비슷하게 이러한 데이터 확보의 난관에 봉착한 사람들에게 내 일련의 삽질 경험을 공유함으로써 약간이나마 도움이 되었으면 하는 마음에서 적어보려고 한다.
# + [markdown] tags=[]
# ## 1. 우리에게 친숙한 포털의 금융 주제 페이지
#
# 여기저기 찾다보면 비교적 가장 먼저, 그러고 쉽게 먼저 접할 수 있는 데이터 소스중 하나지 않을까 싶다.
#
# * [네이버 금융 > 시세 탭 > 일별 시세](https://finance.naver.com/item/sise.nhn?code=035420)
# * [다음 금융 > 현재가 탭 > 일자별 주가](https://finance.daum.net/quotes/A035420#current/quote)
#
# 접근성 측면에서는 좋은 데이터 소스이지만 아쉬운 점들이 몇가지 있다.
#
# * 데이터 다운로드 기능은 따로 제공하지 않고 있다. 따라서 데이터를 다운로드 받기 위해서는 별도의 웹 크롤러를 사용해야 하는 수도 있다.
# * 직접 구현하는 경우 개발이 필요한 번거로움이 있다. 만약에 웹사이트의 구조가 바뀌는 경우.. 일일이 그에 맞춰 크롤러를 수정해주어야 한다.
# * [다른 능력자 분들께서 이미 구현해주신 코드](https://github.com/FinanceData/FinanceDataReader)가 있어서 그냥 가져다가 사용할 수 있으면 한결 다행이다.
# * 사실 엄밀하게 하자면 위의 라이브러리는 HTML 을 파싱하는 방식이 아니라 아예 XML 형태로 결과를 제공하는 내부 API 를 직접 사용하는것으로 보인다.
# * 서비스 제공자가 우리에게 순순히 크롤링을 당해줄 것인지는 또 다른 문제이다. 대량의 데이터를 다운로드 받고자 하는 경우 차단당할 리스크는 더욱 커진다.
# * 제공하는 과거 데이터는 일자별 데이터 뿐이다. 만약에 분봉 이하의 더 작은 타임프레임의 데이터가 필요한 경우 이를 얻을 수 없다.
# * 데이터를 관리하는 주체마다 데이터의 처리방식이나 보관정책이 조금씩 다를 수 있다. 사용 목적에 따라서 이를 먼저 파악하고 알맞는 데이터 소스를 선택해 다운로드 받아야 할 필요가 있지만 딱히 이에 대해 일일이 자세하게 설명하거나 알려주지는 않는다.
# * 제공하는 가격정보가 수정주가인가? 아니면 수정되지 않은 원래의 가격인가? 수정주가라면 수정주가를 산정하는 기준이나 수식은 어떻게 되어있는가?
# * 다음에서 제공하고 있는 NAVER(035420) 의 2003.01.02 종가는 49,800원 이지만 네이버에서 제공하고 있는 NAVER(035420) 의 2003.01.02 종가는 24,900원 이다. 과연 무엇이 더 정확한 혹은 사용목적에 적합한 가격인가?
# * 제공 주체가 가능한 한 가장 오래된 과거의 데이터까지 저장 및 제공하고 있는가?
# * 네이버는 NAVER(035420) 의 주가를 가장 오래된 2002.10.29 일자까지의 정보를 제공하고 있지만 다음에서는 확인 가능한 가장 오래된 날짜가 2003.01.02 이다.
# -
# 위의 내용중에 크롤러를 직접 구현해보는건 생각만 해도 머리가 아파서 따로 시도해보지 않았다.
#
# 참고로 위에서 언급된 [다른 능력자 분들께서 이미 구현해주신 코드](https://github.com/FinanceData/FinanceDataReader)는 아래처럼 사용이 가능한 정도로만 확인했다. 또한 해당 라이브러리는 내부적으로 네이버측의 주식 가격정보 API 를 활용하고 있는 것을 확인할 수 있었다.
import FinanceDataReader as fdr
code = '035420'
fdr_data = fdr.DataReader(code)
fdr_data
# ## 2. 아니 외국 사이트들은 주가를 다운로드 받을 수 있던데?
#
# 주로 해외 자료를 찾다보면 종종 나오는 웹사이트 기반 데이터 소스들로 아래 정도가 있던 걸로 기억한다.
#
# * [Yahoo Finance > Historical Data](https://finance.yahoo.com/quote/035420.KS/history?p=035420.KS)
# * [Investing.com > General > Historical Data](https://www.investing.com/equities/nhn-corp-historical-data)
#
# 앞선 국내 포털의 금융 주제 웹페이지와 구별되는 특징 중 장점이라고 볼 수 있는게 몇가지 있다.
#
# * 데이터를 다운로드 할 수 있는 버튼을 직접적으로 제공한다.
# * 일별 주가 뿐만 아니라 주단위, 월단위 형태의 주가 데이터 또한 얻을 수 있다.
#
# 하지만 역시나 한계점은 있다.
#
# * 역시나 제공하는 과거 데이터는 일자별 데이터 뿐이다. 만약에 분봉 이하의 더 작은 타임프레임의 데이터가 필요한 경우 이를 얻을 수 없다.
# * 역시나 데이터를 관리하는 주체마다 데이터의 처리방식이나 보관정책이 조금씩 다를 수 있다.
# * Yahoo Finance 에서는 일반 종가와 수정 종가를 둘 다 제공하고 있지만 역시 해당 값을 어떻게 계산하는지는 알 수 없다.
# * Investing.com 에서 제공하는 NAVER(035420) 의 2011.09.09 기준 종가는 254,340 지만 Yahoo Finance 에서는 50867.44141(Close) 혹은 49895.66797(AdjClose) 이다.
# * 참고로 네이버와 다음에서 확인되는 NAVER(035420) 의 2011.09.09 (일반) 종가는 207,500 이다.
# * Investing.com 의 경우 기간을 최대로 설정하더라도 한번에 가져올 수 있는 데이터의 개수에는 제한이 있어 보인다.
# * 다운로드를 자동화 하는 것은 또 다른 이슈다.
# * 다운로드 링크가 일반적인 하이퍼링크로 구현된 경우 해당 API 주소를 역으로 분석해 직접 활용할 수 있겠다.
# * Yahoo Finance 가 이에 해당되어 보인다.
# * Investing.com 은 다운로드 버튼이 자바스크립트 기반으로 동작한다.
# * 리버스 엔지니어링이 상대적으로 어렵게 된다.
# * 최악의 경우 selenium 과 같은 headless web browser 를 활용한 자동화가 필요할 수 있다.
# * Investing.com 의 경우 데이터를 다운로드 받기 위해서는 로그인이 필요하다. 자동화 하는데 있어서 또 하나의 걸림목이다.
# 비교적 쉬워보이는 Yahoo Finance 출처에 대한 데이터 다운로더 툴을 간단하게 개발해봤는데 다음과 같았다.
# +
import io
import requests
import pandas as pd
class YahooFinanceKrxHistoricalDailyPriceDataDownloader:
def __init__(self):
self._headers = {
'User-Agent': 'Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/87.0.4280.141 Safari/537.36',
}
def download(self, symbol, start_date=None, end_date=None):
if start_date is None:
start_date = pd.Timestamp(1990, 1, 1)
elif not isinstance(start_date, pd.Timestamp):
start_date = pd.Timestamp(start_date)
if end_date is None:
end_date = pd.Timestamp.now().normalize() + pd.Timedelta(1, unit='day')
elif not isinstance(end_date, pd.Timestamp):
end_date = pd.Timestamp(end_date)
url = 'https://query1.finance.yahoo.com/v7/finance/download/%s.KS' % symbol.upper()
params = {
'period1': int(start_date.timestamp()),
'period2': int(end_date.timestamp()),
'interval': '1d',
'events': 'history',
'includeAdjustedClose': 'true',
}
response = requests.get(url, params=params, headers=self._headers)
df = pd.read_csv(io.BytesIO(response.content), parse_dates=['Date'], index_col='Date')
return df
# -
yahoo_downloader = YahooFinanceKrxHistoricalDailyPriceDataDownloader()
yahoo_data = yahoo_downloader.download(code)
yahoo_data
# 좀 더 찾다보니 역시나 이미 다른 능력자들이 구현해놓은게 있었다. 역시나 머리가 나쁘면 혼자 바퀴를 발명하는 삽질을 하게 된다.
import yfinance as yf
stock = yf.Ticker('%s.KS' % code)
yf_data = stock.history(period='max')
yf_data
# 값을 앞선 결과와 비교해보니 기본적으로 수정주가의 형태로 가져오는 것으로 보인다.
# ## 3. 일반적인 웹 사이트들은 못써먹겠다. 공식 데이터 관리처에서 데이터를 받아올 수 있을까?
#
# 우리나라의 주식 거래 데이터는 당연하게도 [한국거래소(KRX)](http://krx.co.kr/main/main.jsp) 가 관리하고 있다. 이건 약간 TMI 지만 사실 실제 내부적으로 거래소의 전산관련 업무는 [코스콤](https://www.koscom.co.kr/portal/main.do)이라는 회사에서 맡아서 관리하고 있긴 하다.
# 어찌 되었든 여기서 어떻게 데이터를 받아올수만 있다면 데이터의 신뢰성과 관련해서는 걱정할 필요가 없어 보인다.
#
# ### KRX 데이터 상품
#
# 여기저기 찾다보니 거래소에서 데이터를 상품으로 팔고 있다는걸 먼저 알아냈다. 가격이 그리 부담스럽지 않다면 데이터를 구매하는게 가장 바람직하지 않을까?
#
# 아래 페이지에서 데이터 구입에 대한 안내를 하고 있다.
#
# * [KRX 데이터 구입 안내](https://data.krx.co.kr/contents/MDC/DATA/datasale/index.cmd?viewNm=MDCDATA001)
#
# 판매 데이터의 가격을 확인해봤다. 위의 페이지에 첨부된 문서에서 가장 일반적인 *주식 > 일별매매정보 > 유가/코스닥 > 전체항목* 을 먼저 확인해봤다. 가격은 **1년치 30만원**...
# 단순하게 2000년 부터 시작하는 것으로 가정하고 20년 정도를 구매한다 하면 600만원이다.
# 만약에 이 데이터를 구매하면 내가 600만원 이상을 벌 수 있을까? 나는 도저히 본전도 못 채울 것 같아서 바로 포기했다.
#
# 혹시나 해서 데이터 구입 안내 메뉴 바로 밑의 페이지에서 좀 더 구체적인 견적도 내어봤는데 결과는 아래와 같다.
#
# * 선택형 데이터 > 주식 > 일별 시세정보
# * 대상선택: 유가증권
# * 기간: 1996.01 ~ 2021.02
# * 항목선택: 기본옵션 (OHLCV 포함)
# * 예상가격: 3,775,000원
#
# 만약에 해당 가격이 부담되지 않는 사람이 있다면 (그 사람은 더이상 주식투자를 하지 않아도 되지 않나 싶지만) 도전해보고 알려주길 바란다.
#
# ### KRX 정보데이터 시스템
#
# 좀 더 찾다보니 어느정도 기본적인 통계 정보들은 해당 사이트상에서 확인이 가능하도록 제공중인 것을 알게 되었다.
#
# 구체적으로 아래 페이지에서 개별종목의 일별 시세추이를 확인할 수 있다. 무려 다운로드도 가능했다!
#
# * [KRX 정보데이터 시스템 > 통계 > 기본 통계 > 주식 > 종목시세 > 개별종목 시세추이](http://data.krx.co.kr/contents/MDC/MDI/mdiLoader/index.cmd?menuId=MDC0201020103)
#
# 관련해서 [다른 능력자 분들께서 이미 구현해놓으신 코드](https://github.com/FinanceData/FinanceDataReader/blob/b28bfb5f802f7ad5ad8012a59c32553225e40040/krx/data.py)를 참고하면서, 추가적으로 약간의 리버스 엔지니어링을 거쳐 아래와 같은 툴을 개발할 수 있었다.
# +
import requests
import pandas as pd
class KrxHistoricalDailyPriceDataDownloader:
def __init__(self):
self._headers = {
'User-Agent': 'Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/87.0.4280.141 Safari/537.36',
}
self._stocks = None
self._stocks_delisted = None
self._bld = 'dbms/MDC/STAT/standard/MDCSTAT01701'
def get_stocks(self):
data = {
'mktsel': 'ALL',
'typeNo': '0',
'searchText': '',
'bld': 'dbms/comm/finder/finder_stkisu',
}
url = 'http://data.krx.co.kr/comm/bldAttendant/getJsonData.cmd'
response = requests.post(url, data, headers=self._headers)
df = pd.json_normalize(response.json()['block1'])
df = df.set_index('short_code')
return df
def get_stocks_delisted(self):
data = {
'mktsel': 'ALL',
'searchText': '',
'bld': 'dbms/comm/finder/finder_listdelisu',
}
url = 'http://data.krx.co.kr/comm/bldAttendant/getJsonData.cmd'
response = requests.post(url, data, headers=self._headers)
df = pd.json_normalize(response.json()['block1'])
df = df.set_index('short_code')
return df
@property
def stocks(self):
if self._stocks is None:
self._stocks = self.get_stocks()
return self._stocks
@property
def stocks_delisted(self):
if self._stocks_delisted is None:
self._stocks_delisted = self.get_stocks_delisted()
return self._stocks_delisted
def get_full_code(self, symbol):
if symbol in self.stocks.index:
return self.stocks.loc[symbol]['full_code']
if symbol in self.stocks_delisted.index:
return self.stocks_delisted.loc[symbol]['full_code']
raise ValueError('No full_code found for given symbol %s' % symbol)
def download(self, symbol, start_date=None, end_date=None):
if start_date is None:
start_date = pd.Timestamp(1980, 1, 1)
if end_date is None:
end_date = pd.Timestamp.now().normalize() + pd.Timedelta(1, unit='day')
full_code = self.get_full_code(symbol)
url = 'http://data.krx.co.kr/comm/bldAttendant/getJsonData.cmd'
data = {
'bld': self._bld,
'isuCd': full_code,
'isuCd2': '',
'strtDd': start_date.strftime("%Y%m%d"),
'endDd': end_date.strftime("%Y%m%d"),
'share': '1',
'money': '1',
'csvxls_isNo': 'false',
}
response = requests.post(url, data, headers=self._headers)
df = pd.json_normalize(response.json()['output'])
if df.shape[0] == 0:
return None
column_names = {
'TRD_DD':'Date',
'ISU_CD':'Code',
'ISU_NM':'Name',
'MKT_NM':'Market',
'SECUGRP_NM':'SecuGroup',
'TDD_CLSPRC':'Close',
'FLUC_TP_CD':'UpDown',
'CMPPRVDD_PRC':'Change',
'FLUC_RT':'ChangeRate',
'TDD_OPNPRC':'Open',
'TDD_HGPRC':'High',
'TDD_LWPRC':'Low',
'ACC_TRDVOL':'Volume',
'ACC_TRDVAL':'Amount',
'MKTCAP':'MarCap',
'CMPPREVDD_PRC': 'Change',
'LIST_SHRS': 'Shares',
}
df = df.rename(columns=column_names)
df['Date'] = pd.to_datetime(df['Date'])
df['ChangeRate'] = pd.to_numeric(df['ChangeRate'].str.replace(',', ''))
int_columns = [
'Close',
'UpDown',
'Change',
'Open',
'High',
'Low',
'Volume',
'Amount',
'MarCap',
'Shares'
]
for col in int_columns:
if col in df.columns:
df[col] = pd.to_numeric(df[col].str.replace(',', ''), errors='coerce')
df.set_index('Date', inplace=True)
return df
# -
krx_downloader = KrxHistoricalDailyPriceDataDownloader()
krx_data = krx_downloader.download(code)
krx_data
# 해당 출처의 데이터를 사용하면 아래와 같은 장점이 있다고 생각한다.
#
# * 가장 공식적인 데이터이다.
# * 그렇기 때문에 데이터의 정확도와 관련해서 가장 신뢰할 수 있는 출처이지 않을까 싶다.
#
# 다만 아래는 조금은 아쉬운 부분이다.
#
# * 역시나 제공하는 과거 데이터는 일자별 데이터 뿐이다. 만약에 분봉 이하의 더 작은 타임프레임의 데이터가 필요한 경우 별도로 돈을 주고 데이터를 사야 한다.
# * 위에서 제공중인 페이지에서는 수정주가는 따로 제공되지 않는다. 필요하다면 별도로 계산을 해줘야 한다.
#
# 필자는 혹시나 매번 각 종목에 대한 전체 일자에 대한 데이터를 받아오는 것이 거래소의 서버에 부담이 되지 않을까 우려해,
# 아래처럼 로컬에 데이터를 저장하고 정말 필요한 경우에만 최신 데이터를 요청해 덧붙여 나가는 식으로 추가적으로 구현해 사용 중이다.
# 수정주가가 아닌 액면 그대로의 주가 데이터를 다루다 보니 과거의 데이터는 별도 오류로 인한 수정이 필요하지 않는 이상 갱신할 필요가 없어서 가능한 동작이다.
# +
import pandas as pd
from sqlalchemy import create_engine, inspect
from sqlalchemy import Table, MetaData
from exchange_calendars import get_calendar
from tqdm import tqdm
class KrxHistoricalDailyPriceDataLoader:
def __init__(self, filename):
self._downloader = KrxHistoricalDailyPriceDataDownloader()
self._engine = create_engine('sqlite:///' + filename)
self._inspector = inspect(self._engine)
self._calendar = get_calendar('XKRX')
def load_naive(self, symbol):
data = pd.read_sql_table(symbol, self._engine, index_col='Date', parse_dates=['Date'])
return data
def load_if_exists(self, symbol):
if self._inspector.has_table(symbol):
data = self.load_naive(symbol)
if data.shape[0] > 0:
return data
def load_or_download(self, symbol, start_date=None, end_date=None, save=True):
if end_date is None:
now = pd.Timestamp.now(self._calendar.tz).floor("T")
end_date = self._calendar.previous_close(now).astimezone(self._calendar.tz).normalize()
if self._inspector.has_table(symbol):
data = self.load_naive(symbol)
data = data.sort_index()
if data.shape[0] > 0:
start_date = data.index.max().tz_localize(self._calendar.tz) + self._calendar.day
if start_date < end_date:
recent_data = self._downloader.download(symbol, start_date, end_date)
if recent_data is not None and recent_data.shape[0] > 0:
data = data.combine_first(recent_data)[data.columns]
data = data.convert_dtypes(convert_floating=False)
data = data.sort_index()
if save:
data.to_sql(symbol, self._engine, if_exists='replace')
return data
else:
Table(symbol, MetaData()).drop(self._engine)
if not self._inspector.has_table(symbol):
if start_date is None:
start_date = pd.Timestamp(1980, 1, 1)
data = self._downloader.download(symbol, start_date, end_date)
if data is not None and data.shape[0] > 0:
data = data.convert_dtypes(convert_floating=False)
data = data.sort_index()
if save:
data.to_sql(symbol, self._engine, if_exists='replace')
return data
def load(self, symbol):
return self.load_or_download(symbol)
def load_all(self, include_delisted=False, progress_bar=False):
symbols_with_delisted = {}
result = {}
symbols = self._downloader.stocks.index.tolist()
for symbol in symbols:
symbols_with_delisted.setdefault(symbol, False)
if include_delisted:
symbols = self._downloader.stocks_delisted.index.tolist()
for symbol in symbols:
symbols_with_delisted.setdefault(symbol, True)
now = pd.Timestamp.now(self._calendar.tz)
end_date = self._calendar.previous_close(now).normalize()
disable = not progress_bar
for symbol, _delisted in tqdm(symbols_with_delisted.items(), disable=disable):
data = self.load_or_download(symbol, end_date=end_date)
if data is not None:
result[symbol] = data
return result
# -
krx_loader = KrxHistoricalDailyPriceDataLoader('data.sqlite3')
krx_data = krx_loader.load(code)
krx_data
# ## 4. 사실 당연하게도 HTS 에서 데이터를 얻을 수 있다.
#
# 키움증권의 영웅문 HTS 기준으로 아래처럼 하면 단일 종목의 과거 가격데이터를 엑셀파일로 저장할 수 있다.
#
# 1. 키움증권 영웅문 HTS 실행 및 접속
# 2. 키움종합차트 창 생성
# 3. 원하는 종목코드를 입력해 차트 불러오기
# 4. 필요하다면 우측상단의 톱니버튼을 클릭해 수정주가 적용여부 설정 (기본값은 수정주가 적용)
# 5. 우측상단의 연속조회 버튼을 최대한 많이 클릭해 최대한 많은 과거 데이터를 불러오기
# 6. 차트화면 우클릭 후 데이타표 저장 기능 클릭
# 7. 저장할 항목 설정 창에서 원하는 항목 체크 후 확인 버튼 클릭
# 8. 최종적으로 엑셀파일을 원하는 경로에 원하는 이름으로 저장
# 
# 증권사를 통해서 데이터를 받게 되면 아래와 같은 이점을 기대할 수 있다.
#
# * 증권사는 주식 가격데이터의 관리에 관해서는 전문가이다. 충분히 신뢰할 수 있다고 생각한다.
# * 수정주가 적용과 관련된 부분도 가장 합리적으로 수정주가를 계산해서 제공하고 있을 것으로 기대할 수 있다.
# * 일봉 데이터 뿐만 아니라 다른 다양한 타임프레임의 데이터들도 모두 확인 가능하다.
#
# 다만 위와 같은 방식은 아래와 같은 단점이 떠오른다.
#
# * 저걸 단순하게 있는 그대로 자동화 하기 위해서는 거의 자동 매크로와 같은 프로그램이 필요할 듯 하다.
#
# 여기서 자연스럽게 다음 단계로 고려되는 것이 각 증권사에서 제공하는 OpenAPI 와 같은 기능들을 직접 사용하는 것이 되겠다.
# ## 5. 증권사 API 를 활용해서 데이터를 가져와보자.
#
# 맨 처음 시도해볼 증권사 API 를 고를 때, 참고할 자료들이 가장 풍부해 보이는 키움증권의 OpenAPI+ 를 먼저 사용해보기로 마음먹고 개발을 시작했다.
# 이후 이것저것 삽질하면서 개발을 하다 보니 결론적으로는 `koapy` 를 개발하게 되었다.
# 개발과정에서의 여러 삽질들은 여기서 설명하자면 너무 길어질듯해 나중에 기회가 되면 이야기를 풀어봤으면 한다.
#
# 여기서는 `koapy` 를 사용해서 과거 일봉 데이터를 가져와 다른 데이터들과 한번 비교해본다.
# ### 키움증권의 OpenAPI+ 활용하기
#
# 먼저 현재 환경이 32Bit 환경이 맞는지 확인을 하고 진행할 것이다.
# +
import platform
assert platform.architecture()[0] == '32bit'
# -
# 로그인 처리와 관련해서는 이미 자동 로그인 설정이 되어있는 것이 좋지만,
# 그게 아니라면 이후 진행하면서 팝업되는 로그인 창에 수동으로 인증정보를 입력해 로그인을 진행한다.
#
# 이제 아래처럼 `koapy` 를 사용해서 과거 일봉 데이터를 가져올 수 있다.
from koapy import KiwoomOpenApiPlusEntrypoint
kiwoom_entrypoint = KiwoomOpenApiPlusEntrypoint()
kiwoom_entrypoint.EnsureConnected()
kiwoom_data = kiwoom_entrypoint.GetDailyStockDataAsDataFrame(code)
kiwoom_data
# 위의 주가는 수정주가가 적용되지 않은 결과이다. 이것을 앞서 KRX 에서 확인했던 데이터와 비교해보자.
# +
import numpy as np
kiwoom_data_close = kiwoom_data.sort_values('일자')['현재가'].astype(int).to_numpy()
krx_data_close = krx_data.sort_index()['Close'].to_numpy()
# -
np.sum(kiwoom_data_close - krx_data_close)
# 차이가 없는 것이 확인된다. 긍정적인 결과이다.
# 이번에는 수정주가를 확인해보자.
kiwoom_data_adjusted = kiwoom_entrypoint.GetDailyStockDataAsDataFrame(code, adjusted_price=True)
kiwoom_data_adjusted
# 맨 처음의 `FinancialDataReader` 가 네이버 금융의 데이터를 가져오고 있고, 해당 데이터는 수정주가가 적용된 데이터이다. 해당 데이터와 값을 비교해보자
# +
import numpy as np
kiwoom_data_adjusted_close = kiwoom_data_adjusted.sort_values('일자')['현재가'].astype(int).to_numpy()
fdr_data_adjusted_close = fdr_data.sort_index()['Close'].to_numpy()
# -
np.sum(kiwoom_data_adjusted_close - fdr_data_adjusted_close)
np.mean(kiwoom_data_adjusted_close - fdr_data_adjusted_close)
# 꽤나 차이가 발생하는 것을 확인할 수 있으며, 이것은 각 데이터 출처에서 수정주가를 계산하는 방식이 다른 것으로 인한 차이이지 않을까 추측해볼 수 있다.
# 좀 더 시각적으로 확인해보자면 아래처럼 해볼 수 있다.
# +
import matplotlib.pyplot as plt
plt.plot(fdr_data.index, kiwoom_data_adjusted_close)
plt.plot(fdr_data.index, fdr_data_adjusted_close)
plt.legend(['kiwoom', 'fdr'])
plt.show()
# -
# 대략 2013년과 2014년 사이에 뭔 일이 있었는지 급격한 가격변화가 있었고, 해당 시점에서 부터 두 데이터의 가격 차이가 발생하는 것으로 보인다.
#
# 사실 해당 시점은 현재 NAVER 의 전신인 당시의 NHN 이 2013.08.01 을 기점으로 NAVER 와 NHN엔터테인먼트 (현재의 NHN) 으로 기업분할을 한 시점이다.
# 여기서 유추해볼 수 있는건 네이버와 키움증권이 해당 이벤트를 전후해서 수정주가 처리를 다른 방식으로 하고 있다는 것이다.
# 어느 방식이 더 적절한지에 대해서는 당장은 알 수 없기 때문에 제삼자의 케이스를 추가로 확인해보는게 좋겠다.
# 내가 알고 있는 것 처럼 `FinancialDataReader` 의 데이터가 수정주가가 아닐 수도 있을 것 같아서 원래 키움증권의 데이터와도 비교해봤다.
# +
import matplotlib.pyplot as plt
plt.plot(fdr_data.index, kiwoom_data_close)
plt.plot(fdr_data.index, fdr_data_adjusted_close)
plt.legend(['kiwoom', 'fdr'])
plt.show()
# -
# 꽤나 차이가 크다. 이상하게 2007년부터 2009년 정도까지는 가격이 또 겹치는 구간이 있는거 같은데 뭘까.. 내가 보기에는 원래 수정주가가 최근부터 가장 오래된 시점까지 쭉 이어져서 수정되어야 하지만 여기서는 2009년에 한번 뚝 끊고 다시 거기서부터 수정주가를 적용한듯한 느낌도 든다.
# ### 대신증권의 Cybos Plus 활용해보기
#
# 여기서는 대신증권의 Cybos Plus 를 활용해 수정주가를 가져와 추가적으로 확인해보자.
# 대신증권 Cybos Plus 의 경우도 마찬가지로 32Bit 환경에서만 사용이 가능하다.
# +
import platform
assert platform.architecture()[0] == '32bit'
# -
# 앞의 키움증권 케이스와는 다르게 여기서는 자동 로그인과 같은 설정은 따로 없다.
# 대신에 Cybos Plus 프로그램을 미리 실행 및 로그인까지 진행해두는 것이 필요하다.
# 미리 실행시켜두지 않았다면 지금 실행하자.
# 이제 아래처럼 `koapy` 를 사용해서 과거 일봉 데이터를 가져올 수 있다.
from koapy import CybosPlusEntrypoint
cybos_entrypoint = CybosPlusEntrypoint()
cybos_entrypoint.EnsureConnected()
cybos_data_adjusted = cybos_entrypoint.GetDailyStockDataAsDataFrame(code, adjusted_price=True)
cybos_data_adjusted
# 앞서 시각적으로 확인해 봤던 결과에서 위의 케이스를 추가해보자.
cybos_data_adjusted_close = cybos_data_adjusted.sort_values('날짜')['종가'].astype(int).to_numpy()
# +
import matplotlib.pyplot as plt
plt.plot(fdr_data.index, kiwoom_data_adjusted_close)
plt.plot(fdr_data.index, fdr_data_adjusted_close)
plt.plot(fdr_data.index, cybos_data_adjusted_close)
plt.legend(['kiwoom', 'fdr', 'cybos'])
plt.show()
# -
# 키움증권의 OpenAPI+ 와 대신증권의 CybosPlus 사이에서도 데이터 값의 차이가 발생하는 것을 확인할 수 있었다.
# 대신에 그 둘의 차이가 나머지 `FinancialDataReader` 케이스와의 차이보다 확연하게 적은 것을 봤을 때 `FinancialDataReader` 의 데이터가 약간은 표준에서 동떨어진 데이터가 아닐지 의심해볼 수 있겠다.
# ### 번외1. 만약에 앞에서 확인했던 Yahoo Financial 의 데이터까지 그려보면 어떻게 될까?
yahoo_data
yahoo_data_adjusted_close = yahoo_data.sort_index()['Adj Close'].reindex(fdr_data.index).to_numpy()
# +
import matplotlib.pyplot as plt
plt.plot(fdr_data.index, kiwoom_data_adjusted_close)
plt.plot(fdr_data.index, fdr_data_adjusted_close)
plt.plot(fdr_data.index, cybos_data_adjusted_close)
plt.plot(fdr_data.index, yahoo_data_adjusted_close)
plt.legend(['kiwoom', 'fdr', 'cybos', 'yahoo'])
plt.show()
# -
# 무엇이 가장 올바른 수정주가일지는 아직 잘 모르겠지만, 개인적으로는 야후의 데이터도 신뢰가 간다.
# 야후의 데이터가 키움증권과 대신증권의 데이터의 중간즘에 있어서 더욱 그렇게 보이는 걸지도 모르겠다.
# 위아래의 둘도 사실은 별다른 이슈 없이 실전에 충분히 활용 가능하지 않을까 하는 생각은 든다.
# ### 번외2. 만약에 KRX 데이터를 임의로 수정주가 처리해 같이 그려보면 어떻게 될까?
#
# KRX 로 부터 받은 데이터를 어떻게 활용해볼 수 없을까 고민하다가 아래처럼 생각을 해보고 관련 아이디어를 적용해봤다.
#
# * 해당 데이터에서 Change 컬럼을 통해 당일의 주가 변동량을 확인할 수 있다.
# * 당일 종가해서 당일 Change 값을 빼면 당일의 기준가를 역산해 확인할 수 있다.
# * 위의 방식으로 역산한 기준가와 전일의 종가 사이에 값의 차이가 발생하는 경우 기준가가 알 수 없는 외부 영향으로 인해 인위적으로 변동이 된 것으로 생각할 수 있다.
# * 일반적인 상황에서는 전일의 종가가 자연스럽게 당일의 기준가가 된다.
# * 역산한 기준가와 전일 종가 사이의 비율을 통해 수정주가 비율을 추측해볼 수 있겠다고 생각이 들었다.
#
# 다만 해당 방식을 나이브하게 여러 종목에 적용해보다가 아래와 같은 문제가 있어서 약간은 무식하게 대응한게 있는데, 구체적으로 아래와 같다.
#
# * KRX 데이터에서는 특정 종목이 과거에 상장폐지 후 다시 상장하는 경우 그 모든 가격 데이터가 하나의 시계열로 저장되는 경우가 있다.
# * 위와 같은 경우에 상장폐지와 재 상장 사이를 위와 같은 방식으로 보정하는건 의미가 없다. 오히려 가격을 왜곡시킬 수 있다.
# * 데이터상으로 확인해 봤을때 상장폐지 이력이 없는 종목들에서는 가장 긴 연휴가 11일 정도로 확인되었다. 따라서 그보다 긴 공백이 발견되면 상장폐지가 한번 되었던 종목으로 판단하고 구분해 처리한다.
#
# 이외에 또 다른 문제가 있을지는 아직까지는 잘 모르겠다.
# +
import datetime
import numpy as np
def get_adjust_ratios(data):
data = data.sort_index(ascending=False)
adjust_ratios = []
last_close = data['Close'].iloc[0]
last_adjust_ratio = 1.0
adjust_ratios.append(last_adjust_ratio)
eleven_days = datetime.timedelta(days=11)
for i in range(data.shape[0] - 1):
if data.index[i] - data.index[i+1] > eleven_days:
last_close = data['Close'].iloc[i+1]
last_adjust_ratio = 1.0
else:
last_close = last_close - data['Change'].iloc[i] * last_adjust_ratio
last_adjust_ratio = last_close / data['Close'].iloc[i+1]
adjust_ratios.append(last_adjust_ratio)
adjust_ratios = np.array(adjust_ratios)
return adjust_ratios
def adjust_prices(data):
data = data.sort_index(ascending=False)
adjust_ratios = get_adjust_ratios(data)
data['Adj Open'] = data['Open'] * adjust_ratios
data['Adj High'] = data['High'] * adjust_ratios
data['Adj Low'] = data['Low'] * adjust_ratios
data['Adj Close'] = data['Close'] * adjust_ratios
data['Adj Volume'] = data['Volume'] / adjust_ratios
return data
# -
krx_data_adjusted = adjust_prices(krx_data)
krx_data_adjusted
krx_data_adjusted_close = krx_data_adjusted.sort_index()['Adj Close'].reindex(fdr_data.index).to_numpy()
# +
import matplotlib.pyplot as plt
plt.plot(fdr_data.index, kiwoom_data_adjusted_close)
plt.plot(fdr_data.index, fdr_data_adjusted_close)
plt.plot(fdr_data.index, cybos_data_adjusted_close)
plt.plot(fdr_data.index, yahoo_data_adjusted_close)
plt.plot(fdr_data.index, krx_data_adjusted_close)
plt.legend(['kiwoom', 'fdr', 'cybos', 'yahoo', 'krx'])
plt.show()
# -
# 놀랍게도 수정한 KRX 데이터가 앞서 대신증권 Cybos Plus 를 활용해 받은 수정주가 데이터를 거의 가려버리는 형국으로 보인다.
# 아마도 대신증권에서는 수정주가 계산시 내가 위에서 생각해낸 아이디어와 거의 비슷한 방향으로 접근하고 있는건 아닐지 조심스레 추측해본다.
# ## 6. 그래서 과거 주식 가격 데이터를 지속적으로 업데이트 하는데에 있어서 가장 좋은 방법은 뭘까?
#
# 일단 당장 자신있게 내릴 수 있는 결론 하나는 다음과 같다.
#
# > 아래의 조건에 만족하는 경우 KRX 의 데이터를 활용한다.
# > * 수정주가는 필요없다.
# > * 일봉보다 작은 타임프레임의 가격 데이터는 필요없다.
#
# 그 다음으로 아래도 생각해볼 수 있겠다.
#
# > 아래의 조건에 만족하는 경우 Yahoo Finance 의 데이터를 활용한다.
# > * 일봉보다 작은 타임프레임의 가격 데이터는 필요없지만 수정주가는 필요하다.
# > * 증권사 API 를 사용하는건 너무 번거로운 것 같다.
#
# 당장 Yahoo Finance 를 사용하지 말아야 할 이유는 떠오르지 않지만, 만약에 Yahoo Finance 는 싫은데 일봉 단위 수정주가가 필요한 경우, 차선책으로 KRX 데이터로부터 직접 수정주가를 생성하는 방식도 어느정도 유효하지 않을까 싶은 생각이다. 이 경우 자신이 직접 수정주가를 처리하는 것에 대한 리스크는 어느정도 감수해야 한다. (개인적으로 그리 크게 리스크가 있을 것 같지는 않다는 생각이다.)
#
# > 아래의 조건에 만족하는 경우 KRX 의 데이터에 기반해 직접 수정주가 처리를 해서 활용한다.
# > * 일봉보다 작은 타임프레임의 가격 데이터는 필요없지만 수정주가는 필요하다.
# > * 증권사 API 를 사용하는건 너무 번거로운 것 같다.
# > * 수정주가 처리방식을 내가 직접 관리하고 싶다.
# > * 왠지 Yahoo Finance 는 쓰기 싫다.
#
# 이외에 일봉보다 작은 타임프레임의 영역으로 들어가게 되면 결국 증권사 API 의 사용은 필수가 된다. 수정주가의 경우는 제공 안하는 증권사를 찾는게 더 어렵지 않을까 싶다.
#
# 여기서 고려했던 두 증권사 API 말고도 다른 다양한 API 가 있겠지만, 먼저 고려했던 둘 중에 하나를 고르자면, 적어도 주식 가격데이터 확보의 관점에서는, 개인적으로 대신증권의 Cybos Plus 를 권하고 싶다. 구체적인 이유는 다음과 같다.
#
# 1. Cybos Plus 가 속도 측면에서 압도적으로 빠르다.
# * 모든 증권사 API 는 과도한 요청을 제한하기 위해 시간당 API 요청 횟수의 제한을 두는데 상대적으로 Cybos Plus 가 훨씬 느슨하게 제한을 하고 있다.
# * 키움증권의 OpenAPI+ 의 경우 표면적으로 1초당 5회라고 이야기하고 있지만 내부적으로는 길게 봤을때 1시간당 1000회로, 거의 4초당 1회 꼴로 제한을 두고 있다.
# * 그러면서 1회 호출마다 확보 가능한 레코드 수는 600건 정도에 불과하다.
# * 일봉 데이터 기준 단순 속도를 계산해보면 **150일/초** 정도가 된다.
# * 반면에 대신증권의 Cybos Plus 는 단순하게 15초에 60회, 나눠보면 1초에 4회 정도까지 호출이 가능하다.
# * Cybos Plus 는 요청시에 필요한 컬럼들만 가져오도록 설정이 가능하다.
# * Cybos Plus 는 1회 요청에서 결과 테이블 내 전체 셀 개수의 총량을 제한하는 식으로 트래픽을 조절하고 있다.
# * 위의 두가지 성격으로 인해 1회 호출시에 확보 가능한 레코드 수는 요청하는 컬럼수에 반비례하게 된다.
# * 따라서 정말 필요한 컬럼들로만 제한해 호출하는 경우 속도는 더욱 빨라질 수 있다.
# * 일반적인 컬럼들로 설정했다고 가정했을때 1회 호출마다 대략 2000건의 레코드를 받을 수 있다.
# * 일봉 데이터 기준 단순 속도를 계산해보면 **8000일/초** 정도가 된다. 대략 키움증권 OpenAPI+ 의 **50배**가 되는 속도이다.
# 2. Cybos Plus 가 지원하는 확인 가능한 과거 데이터의 기간도 더 길다.
# * 더 과거의 데이터를 확인할 수 있다는 의미다.
# * 만약에 최대한 많은 데이터를 확보하려는 입장이라면 유의미한 차이로 볼 수 있다.
# 3. 요청시에 필요한 컬럼들만 가져오도록 설정 가능한 부분이 유용하게 사용될 수 있다. 예를 들어 아래와 같은 구성이 가능하다.
# 1. 일별 타임프레임하에 수정주가비율 관련 정보만 요청해 주기적으로 업데이트 한다.
# * 수정주가가 하루 단위보다 더 잘게 나뉘어서 적용되지는 않기 때문에 일별 타임프레임으로 저장하면 된다.
# * 해당 정보는 매번 전체 업데이트를 거쳐도 된다. 필요한 컬럼만 요청하기 때문에 비교적 빠르게 값을 갱신하는 것이 가능하다.
# * 주기적으로 업데이트 처리를 하는게 가능하다면 효율성 측면에서 아래처럼 처리할 수도 있어 보인다.
# 1. 업데이트시 현재 로컬에 가지고 있는 데이터 기준으로 가장 최근의 영업일과 중복되는/겹쳐지는 결과가 되도록 기간을 설정해 API 를 통해 수정주가비율을 요청한다.
# * 매일 업데이트 하는 경우 기간을 최근 2영업일로 설정해 요청하면 된다.
# 2. 겹치게 되는 영업일의 수정주가비율에서 차이가 발생하게 되면 전체 기간에 대한 업데이트를 진행한다.
# * 만약에 차이가 없으면 겹치는 날짜를 제외한 나머지 신규 데이터만 추가한다.
# * 차이가 발생하는 경우의 기존 과거기간 전체에 대한 업데이트는 아래와 같은 방식으로도 가능하다.
# 1. 겹치는 영업일에서 기존/신규 수정주가비율 사이의 값의 차이를 비율 형태로 확인한다.
# 2. 기존에 가지고 있던 과거 수정주가비율 전체를 해당 비율만큼 동일하게 조정해준다.
# * 겹치게 되는 영업일에서 기존 로컬의 수정주가비율이 요청을 통해 받은 신규 수정주가비율 만큼 되도록 조정해주는 것으로 보면 된다.
# 2. 다른 필요한 주식 가격 데이터들은 모두 수정주가를 적용하지 않은 버전으로 가져와 로컬에 저장한다.
# * 이 경우 매번 전체 데이터를 업데이트 할 필요 없이 누락된 최신 데이터만 추가로 요청해 덧붙이는 식으로 업데이트가 가능하다.
# 3. 이후 수정주가가 필요할때마다 아래와 같이 처리해 사용한다.
# 1. 원본주가 데이터를 먼저 로드한다.
# 2. 일별 주가수정비율 데이터도 같이 로드한다.
# 3. 날짜 기준으로 위의 두 데이터를 조인 후 주가수정비율에 맞게 각 가격의 값을 조정한다.
#
# 증권사 API 선택방식과 관련해서 정리하자면 아래처럼 될 수 있겠다.
#
# > 아래의 조건에 만족하는 경우 대신증권 Cybos Plus 의 데이터를 활용한다.
# > * 분봉 이하를 포함한 다양한 타임프레임의 가격 데이터가 필요하다.
# > * 데이터 업데이트를 효율적으로 하고 싶다.
# > * 더 많은 기간의 데이터를 확보하는 것이 중요하다.
#
# > 아래의 조건에 만족하는 경우 키움증권 OpenAPI+ 의 데이터를 활용한다.
# > * 분봉 이하를 포함한 다양한 타임프레임의 가격 데이터가 필요하다.
# > * 업데이트 속도나 효율성은 크게 중요하지 않다.
# > * 확보 가능한 과거 데이터의 기간도 크게 중요하지 않다.
# > * 여러 증권사 API 를 사용하는게 번거로울 것 같다. 나는 원래 쓰던 키움증권만 쓰련다.
#
# 각 증권사마다 수정주가 계산을 다르게 처리하는 것과 관련해서는, 필요하다면 각기 증권사에 문의해본 뒤에 자신에게 맞는 방식을 골라서 사용하면 되지 않을지 하는 생각이다.
# ### Cybos Plus 를 활용한 수정주가 별도 관리 방식의 구체적인 예시
#
# 위의 Cybos Plus 의 장점을 설명하는 내용중 마지막 3번에서 설명했던 방식을 좀 더 구체적인 예시를 들어보자면 아래와 같다.
adjustment_ratios = cybos_entrypoint.GetDailyAdjustmentRatioAsDataFrame(code)
adjustment_ratios
minutes_data = cybos_entrypoint.GetMinuteStockDataAsDataFrame(code, 15)
minutes_data
adjusted_minutes_data = pd.merge(minutes_data, adjustment_ratios, on='날짜', how='left')
adjusted_minutes_data
price_multiplier = adjusted_minutes_data['수정주가비율'] / 100
price_multiplier
adjusted_minutes_data['수정시가'] = adjusted_minutes_data['시가'] * price_multiplier
adjusted_minutes_data['수정고가'] = adjusted_minutes_data['고가'] * price_multiplier
adjusted_minutes_data['수정저가'] = adjusted_minutes_data['저가'] * price_multiplier
adjusted_minutes_data['수정종가'] = adjusted_minutes_data['종가'] * price_multiplier
adjusted_minutes_data['수정거래량'] = adjusted_minutes_data['거래량'] / price_multiplier
adjusted_minutes_data
# ### 데이터 소스별 지원 기간 비교
#
# 위에서 검토해본 데이터 소스들에서 제공 가능한 가장 오래된 데이터의 범위가 어디까지인지 비교해본다.
code = '005930'
fdr_data = fdr.DataReader(code)
yahoo_data = yahoo_downloader.download(code)
krx_data = krx_downloader.download(code)
kiwoom_data = kiwoom_entrypoint.GetDailyStockDataAsDataFrame(code)
cybos_data = cybos_entrypoint.GetDailyStockDataAsDataFrame(code)
data = [
fdr_data.index.to_series().describe(datetime_is_numeric=True),
yahoo_data.index.to_series().describe(datetime_is_numeric=True),
krx_data.index.to_series().describe(datetime_is_numeric=True),
pd.to_datetime(kiwoom_data['일자'], format='%Y%m%d').describe(datetime_is_numeric=True),
pd.to_datetime(cybos_data['날짜'], format='%Y%m%d').describe(datetime_is_numeric=True),
]
keys = ['fdr', 'yahoo', 'krx', 'kiwoom', 'cybos']
result = pd.concat(data, axis=1, keys=keys).T
result
result['count'].plot.bar()
plt.show()
# 대신증권의 Cybos Plus 가 가장 많은 데이터를 제공하는 것을 확인할 수 있다.
| docs/source/notebooks_ipynb/getting-historical-stock-price-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
# # CI/CD for TFX pipelines
# ## Learning Objectives
#
# 1. Develop a CI/CD workflow with Cloud Build to build and deploy TFX pipeline code.
# 2. Integrate with Github to automatically trigger pipeline deployment with source code repository changes.
# In this lab, you will walk through authoring a Cloud Build CI/CD workflow that automatically builds and deploys the same TFX pipeline from `lab-02.ipynb`. You will also integrate your workflow with GitHub by setting up a trigger that starts the workflow when a new tag is applied to the GitHub repo hosting the pipeline's code.
#
#
# ## Setup
# +
import yaml
# Set `PATH` to include the directory containing TFX CLI.
# PATH=%env PATH
# %env PATH=/home/jupyter/.local/bin:{PATH}
# -
# !python -c "import tfx; print('TFX version: {}'.format(tfx.__version__))"
# **Note**: this lab was built and tested with the following package versions:
#
# `TFX version: 0.25.0`
# (Optional) If the TFX version above does not match the lab tested defaults, run the command below:
# %pip install --upgrade --user tfx==0.25.0
# **Note**: you may need to restart the kernel to pick up the correct package versions.
# ## Understanding the Cloud Build workflow
# Review the `cloudbuild.yaml` file to understand how the CI/CD workflow is implemented and how environment specific settings are abstracted using **Cloud Build** variables.
#
# The **Cloud Build** CI/CD workflow automates the steps you walked through manually during `lab-02`:
# 1. Builds the custom TFX image to be used as a runtime execution environment for TFX components and as the AI Platform Training training container.
# 1. Compiles the pipeline and uploads the pipeline to the KFP environment
# 1. Pushes the custom TFX image to your project's **Container Registry**
#
# The **Cloud Build** workflow configuration uses both standard and custom [Cloud Build builders](https://cloud.google.com/cloud-build/docs/cloud-builders). The custom builder encapsulates **TFX CLI**.
#
# ## Configuring environment settings
#
# Navigate to [AI Platform Pipelines](https://console.cloud.google.com/ai-platform/pipelines/clusters) page in the Google Cloud Console.
#
# ### Create or select an existing Kubernetes cluster (GKE) and deploy AI Platform
#
# Make sure to select `"Allow access to the following Cloud APIs https://www.googleapis.com/auth/cloud-platform"` to allow for programmatic access to your pipeline by the Kubeflow SDK for the rest of the lab. Also, provide an `App instance name` such as "tfx" or "mlops". Note you may have already deployed an AI Pipelines instance during the Setup for the lab series. If so, you can proceed using that instance below in the next step.
#
# Validate the deployment of your AI Platform Pipelines instance in the console before proceeding.
# ### Configure environment settings
#
# Update the below constants with the settings reflecting your lab environment.
#
# - `GCP_REGION` - the compute region for AI Platform Training and Prediction
# - `ARTIFACT_STORE` - the GCS bucket created during installation of AI Platform Pipelines. The bucket name starts with the `kubeflowpipelines-` prefix.
# Use the following command to identify the GCS bucket for metadata and pipeline storage.
# !gsutil ls
# * `CUSTOM_SERVICE_ACCOUNT` - In the gcp console Click on the Navigation Menu and navigate to `IAM & Admin`, then to `Service Accounts` and use the service account starting with prefix - 'tfx-tuner-caip-service-account'. This enables CloudTuner and the Google Cloud AI Platform extensions Tuner component to work together and allows for distributed and parallel tuning backed by AI Platform Vizier's hyperparameter search algorithm. Please see the lab setup `README` for setup instructions.
# - `ENDPOINT` - set the `ENDPOINT` constant to the endpoint to your AI Platform Pipelines instance. The endpoint to the AI Platform Pipelines instance can be found on the [AI Platform Pipelines](https://console.cloud.google.com/ai-platform/pipelines/clusters) page in the Google Cloud Console. Open the *SETTINGS* for your instance and use the value of the `host` variable in the *Connect to this Kubeflow Pipelines instance from a Python client via Kubeflow Pipelines SKD* section of the *SETTINGS* window. The format is `'....[region].pipelines.googleusercontent.com'`.
# +
#TODO: Set your environment resource settings here for GCP_REGION, ARTIFACT_STORE_URI, ENDPOINT, and CUSTOM_SERVICE_ACCOUNT.
GCP_REGION = ''
ARTIFACT_STORE_URI = ''
CUSTOM_SERVICE_ACCOUNT = ''
ENDPOINT = ''
PROJECT_ID = !(gcloud config get-value core/project)
PROJECT_ID = PROJECT_ID[0]
# -
# ## Creating the TFX CLI builder
# ### Review the Dockerfile for the TFX CLI builder
# !cat tfx-cli/Dockerfile
# !cat tfx-cli/requirements.txt
# ### Build the image and push it to your project's Container Registry
# **Hint**: Review the [Cloud Build](https://cloud.google.com/cloud-build/docs/running-builds/start-build-manually#gcloud) gcloud command line reference for builds submit. Your image should follow the format `gcr.io/[PROJECT_ID]/[IMAGE_NAME]:latest`. Note the source code for the tfx-cli is in the directory `./tfx-cli`.
IMAGE_NAME='tfx-cli'
TAG='latest'
IMAGE_URI='gcr.io/{}/{}:{}'.format(PROJECT_ID, IMAGE_NAME, TAG)
# TODO: Your gcloud command here to build tfx-cli and submit to Container Registry.
# ## Exercise: manually trigger CI/CD pipeline run with Cloud Build
#
# You can manually trigger **Cloud Build** runs using the `gcloud builds submit` command.
# +
PIPELINE_NAME='tfx_covertype_continuous_training'
MODEL_NAME='tfx_covertype_classifier'
DATA_ROOT_URI='gs://workshop-datasets/covertype/small'
TAG_NAME='test'
TFX_IMAGE_NAME='lab-03-tfx-image'
PIPELINE_FOLDER='pipeline'
PIPELINE_DSL='runner.py'
RUNTIME_VERSION='2.3'
PYTHON_VERSION='3.7'
USE_KFP_SA='False'
ENABLE_TUNING='True'
SUBSTITUTIONS="""
_GCP_REGION={},\
_ARTIFACT_STORE_URI={},\
_CUSTOM_SERVICE_ACCOUNT={},\
_ENDPOINT={},\
_PIPELINE_NAME={},\
_MODEL_NAME={},\
_DATA_ROOT_URI={},\
_TFX_IMAGE_NAME={},\
TAG_NAME={},\
_PIPELINE_FOLDER={},\
_PIPELINE_DSL={},\
_RUNTIME_VERSION={},\
_PYTHON_VERSION={},\
_USE_KFP_SA={},\
_ENABLE_TUNING={},
""".format(GCP_REGION,
ARTIFACT_STORE_URI,
CUSTOM_SERVICE_ACCOUNT,
ENDPOINT,
PIPELINE_NAME,
MODEL_NAME,
DATA_ROOT_URI,
TFX_IMAGE_NAME,
TAG_NAME,
PIPELINE_FOLDER,
PIPELINE_DSL,
RUNTIME_VERSION,
PYTHON_VERSION,
USE_KFP_SA,
ENABLE_TUNING
).strip()
# -
# Hint: you can manually trigger **Cloud Build** runs using the `gcloud builds submit` command. See the [documentation](https://cloud.google.com/sdk/gcloud/reference/builds/submit) for pass the `cloudbuild.yaml` file and SUBSTITIONS as arguments.
# TODO: write gcloud builds submit command to trigger manual pipeline run.
# ## Exercise: Setting up GitHub integration
# In this exercise you integrate your CI/CD workflow with **GitHub**, using [Cloud Build GitHub App](https://github.com/marketplace/google-cloud-build).
# You will set up a trigger that starts the CI/CD workflow when a new tag is applied to the **GitHub** repo managing the pipeline source code. You will use a fork of this repo as your source GitHub repository.
# ### Create a fork of this repo
# #### [Follow the GitHub documentation](https://help.github.com/en/github/getting-started-with-github/fork-a-repo) to fork this repo
# #### Create a Cloud Build trigger
#
# Connect the fork you created in the previous step to your Google Cloud project and create a trigger following the steps in the [Creating GitHub app trigger](https://cloud.google.com/cloud-build/docs/create-github-app-triggers) article. Use the following values on the **Edit trigger** form:
#
# |Field|Value|
# |-----|-----|
# |Name|[YOUR TRIGGER NAME]|
# |Description|[YOUR TRIGGER DESCRIPTION]|
# |Event| Tag|
# |Source| [YOUR FORK]|
# |Tag (regex)|.\*|
# |Build Configuration|Cloud Build configuration file (yaml or json)|
# |Cloud Build configuration file location|/ workshops/tfx-caip-tf23/lab-03-tfx-cicd/labs/cloudbuild.yaml|
#
#
# Use the following values for the substitution variables:
#
# |Variable|Value|
# |--------|-----|
# |_GCP_REGION|[YOUR GCP_REGION]|
# |_CUSTOM_SERVICE_ACCOUNT|[YOUR CUSTOM_SERVICE_ACCOUNT]|
# |_ENDPOINT|[Your inverting proxy host pipeline ENDPOINT]|
# |_TFX_IMAGE_NAME|lab-03-tfx-image|
# |_PIPELINE_NAME|tfx_covertype_continuous_training|
# |_MODEL_NAME|tfx_covertype_classifier|
# |_DATA_ROOT_URI|gs://workshop-datasets/covertype/small|
# |_PIPELINE_FOLDER|workshops/tfx-caip-tf23/lab-03-tfx-cicd/labs/pipeline|
# |_PIPELINE_DSL|runner.py|
# |_PYTHON_VERSION|3.7|
# |_RUNTIME_VERSION|2.3|
# |_USE_KFP_SA|False|
# |_ENABLE_TUNING|True|
# #### Trigger the build
#
# To start an automated build [create a new release of the repo in GitHub](https://help.github.com/en/github/administering-a-repository/creating-releases). Alternatively, you can start the build by applying a tag using `git`.
# ```
# git tag [TAG NAME]
# git push origin --tags
# ```
#
# #### Verify triggered build in Cloud Build dashboard
#
# After you see the pipeline finish building on the Cloud Build dashboard, return to [AI Platform Pipelines](https://console.cloud.google.com/ai-platform/pipelines/clusters) in the console. Click `OPEN PIPELINES DASHBOARD` and view the newly deployed pipeline. Creating a release tag on GitHub will create a pipeline with the name `tfx_covertype_continuous_training-[TAG NAME]` while doing so from GitHub will create a pipeline with the name `tfx_covertype_continuous_training_github-[TAG NAME]`.
# ## Next Steps
# In this lab, you walked through authoring a Cloud Build CI/CD workflow that automatically builds and deploys a TFX pipeline. You also integrated your TFX workflow with GitHub by setting up a Cloud Build trigger. In the next lab, you will walk through inspection of TFX metadata and pipeline artifacts created during TFX pipeline runs.
# # License
# <font size=-1>Licensed under the Apache License, Version 2.0 (the \"License\");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at [https://www.apache.org/licenses/LICENSE-2.0](https://www.apache.org/licenses/LICENSE-2.0)
#
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an \"AS IS\" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.</font>
| workshops/tfx-caip-tf23/lab-03-tfx-cicd/labs/lab-03.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_tensorflow_p36)
# language: python
# name: conda_tensorflow_p36
# ---
# ## MVP Siamese LSTM Net
#
# This is a baseline siamese LSTM net. The purpose is to build out the architecture, and see if the net can get as good as validation score as the classifiers.
#
# Ideas Implemented:
# * Remove stop words
# * Add BatchNormalization
# +
# data manipulation
import utils
import pandas as pd
import numpy as np
import logging
# Keras
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.models import Sequential, Model
from keras.layers import Dense, Flatten, LSTM, Conv1D, MaxPooling1D, Dropout, Activation, Input, Add, concatenate, BatchNormalization
from keras.layers.embeddings import Embedding
from keras.utils.vis_utils import model_to_dot
from keras.callbacks import ModelCheckpoint, TensorBoard, Callback, EarlyStopping
from keras.models import load_model
from sklearn import metrics
from sklearn.model_selection import train_test_split
# plotting
from IPython.display import SVG
# -
X_train = utils.load('X_train')
y_train = utils.load('y_train')
model_name = 'lstm_LEMMA_SW_dropout_20_lstm_layer_DO_20'
# ## Tokenize and Encode vocabulary
#
# 1. Limit the vocab to 20,000 words.
# 2. Clean questions only and do not lemmatize.
# 3. Limit the question length to 100 tokens.
# +
vocabulary_size = 20000
max_q_len = 100
X_train_stack = utils.apply_lemma(
utils.clean_questions(
utils.stack_questions(X_train),
excl_num=False),
incl_stop_words=False)
tokenizer = Tokenizer(num_words= vocabulary_size)
tokenizer.fit_on_texts(X_train_stack)
sequences = tokenizer.texts_to_sequences(X_train_stack)
data = pad_sequences(sequences, maxlen=max_q_len)
print(data.shape)
data[:,0].sum()
# -
# ## Embedding Matrix
#
# 1. Calculates the embedding matrix utilizing spaCy `en_core_web_lg` word vectors.
# * https://spacy.io/models/en#en_core_web_lg
# * GloVe vectors trained on Common Crawl
try:
embedding_matrix = utils.load('embedding_matrix_lemma_sw')
except:
# create a weight matrix for words in training docs
embedding_matrix = np.zeros((vocabulary_size, 300))
for word, index in tokenizer.word_index.items():
# print(word, index, end='\r')
if index > vocabulary_size - 1:
break
else:
embedding_vector = utils.nlp(word).vector
if embedding_vector is not None:
embedding_matrix[index] = embedding_vector
# break
utils.save(embedding_matrix, 'embedding_matrix_lemma_sw')
# ## Define the batch to pass into the network
#
# Create arrays to split the stacked data into question 1 set and question 2 set for each pair.
# +
# cooncatenate the two questions
odd_idx = [i for i in range(data.shape[0]) if i % 2 == 1]
even_idx = [i for i in range(data.shape[0]) if i % 2 == 0]
data_1 = data[odd_idx]
data_2 = data[odd_idx]
# split the data set into a validation set
data_train, data_val, label_train, label_val = train_test_split(np.hstack([data_1, data_2]),
y_train,
stratify=y_train,
test_size = 0.33,
random_state=42)
# split the concatenation back into 2 data sets for the siamese network
data_1_train = data_train[:, :max_q_len]
data_2_train = data_train[:, max_q_len:]
data_1_val = data_val[:, :max_q_len]
data_2_val = data_val[:, max_q_len:]
print(f'Train major class: {len(label_train[label_train == 0]) / len(label_train):.2}')
print(f'Val major class: {len(label_val[label_val == 0]) / len(label_val):.2}')
# -
# ## Build out legs of the siamese network
#
# The architecure is the following,
#
# 0. Input - (100,) word tensor
# 1. Embedding Layer - outputs (300,) **not trainable**
# 2. LSTM - default outputs (300,)
# 3. Concatenate the two nets outputs (600,)
# 4. BatchNormalization
# 5. Dropout - 20%
# 6. Dense - outputs (100,), activation `tanh` -- somewhat random decision
# 7. BatchNormalization
# 8. Dropout - 20%
# 9. Dense - outputs (1,), activation `sigmoid`
# +
# Creating word embedding layer
embedding_layer = Embedding(vocabulary_size, 300, input_length=100,
weights=[embedding_matrix], trainable=False)
# Creating LSTM Encoder
# Bidirectional(LSTM(self.number_lstm_units, dropout=self.rate_drop_lstm, recurrent_dropout=self.rate_drop_lstm))
lstm_layer = LSTM(300, dropout=0.2, recurrent_dropout=0.2)
# Creating LSTM Encoder layer for First Sentence
sequence_1_input = Input(shape=(100,), dtype='int32')
embedded_sequences_1 = embedding_layer(sequence_1_input)
x1 = lstm_layer(embedded_sequences_1)
# Creating LSTM Encoder layer for Second Sentence
sequence_2_input = Input(shape=(100,), dtype='int32')
embedded_sequences_2 = embedding_layer(sequence_2_input)
x2 = lstm_layer(embedded_sequences_2)
# +
# Merging two LSTM encodes vectors from sentences to
# pass it to dense layer applying dropout and batch normalisation
merged = concatenate([x1, x2])
merged = BatchNormalization()(merged)
merged = Dropout(.2)(merged)
merged = Dense(100)(merged) # feed forward
merged = BatchNormalization()(merged)
merged = Dropout(0.2)(merged)
preds = Dense(1, activation='sigmoid')(merged)
model = Model(inputs=[sequence_1_input, sequence_2_input], outputs=preds)
model.compile(loss='binary_crossentropy', optimizer='nadam', metrics=['acc'])
# SVG(model_to_dot(model, show_shapes=True).create(prog='dot', format='svg'))
model.summary()
# +
# Callbacks
file_path = '../data/keras_models/' + model_name + '_{epoch:02d}-{val_loss:.2f}.hdf5'
model_checkpoint = ModelCheckpoint(filepath=file_path, save_best_only=True)
tensorboard = TensorBoard(log_dir='../data/tensorboard')
early_stopping = EarlyStopping(monitor='val_loss',
min_delta=0,
patience=3,
verbose=1,
mode='auto',
restore_best_weights=True)
# calc_auc = IntervalEvaluation(([data_1_val, data_2_val], label_val), interval=1)
# -
model.fit([data_1_train, data_2_train], label_train,
validation_data=([data_1_val, data_2_val], label_val),
epochs=200, batch_size=128, shuffle=True,
callbacks=[model_checkpoint, tensorboard, early_stopping])
# ## Results
# +
model = load_model('../data/keras_models/lstm_LEMMA_SW_dropout_20_lstm_layer_DO_20_08-0.52.hdf5')
y_prob = model.predict([data_1_val, data_2_val], batch_size=128, verbose=1)
# +
results_df = utils.load('results')
results_df = results_df.drop(index=model_name, errors='ignore')
results_df = results_df.append(utils.log_keras_scores(label_val, y_prob, model_name))
results_df.sort_values('avg_auc', ascending=False)
# -
utils.save(results_df, 'results')
# ### Next Steps
#
# Similar to the best NN model yet. Let's build upon this model.
#
# * Dense layer with sigmoid activation
# * Dense layer with same dimension as LSTM
| py_files/58. DL - Lemma_SW_Dropout_20_LSTM_Dropout_20.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 3. Building Prediction Model
# ### Tensorflow
import pandas as pd
import numpy as np
import tensorflow as tf
from sklearn.preprocessing import MinMaxScaler
X_train = np.load('X_train.npy')
X_test = np.load('X_test.npy')
y_train = np.load('y_train.npy')
y_test = np.load('y_test.npy')
X_test_input = np.load('X_test_input.npy')
display('shape of X :', np.shape(X_test),'shape of y :', np.shape(y_test))
y_test = np.reshape(y_test, (-1, 1))
y_train = np.reshape(y_train, (-1, 1))
np.nan_to_num(X_train, copy=False)
np.nan_to_num(X_test, copy=False)
# +
scaler = MinMaxScaler()
scaler.fit(X_train)
X_train = X_train[~np.isnan(X_train).any(axis=1)]
X_train = scaler.transform(X_train)
X_test = X_test[~np.isnan(X_test).any(axis=1)]
X_test = scaler.transform(X_test)
# -
tf.reset_default_graph()
# +
learning_rate = 0.01
X = tf.placeholder(tf.float32, [None, 78], name = 'x-input')
Y = tf.placeholder(tf.float32, [None, 1], name = 'y-input')
keep_prob = tf.placeholder(tf.float32)
with tf.name_scope("layer1") as scope:
W1 = tf.get_variable("weight1", shape=[78, 50],
initializer=tf.contrib.layers.xavier_initializer())
b1 = tf.Variable(tf.random_normal([50]), name='bias1')
layer1 = tf.nn.relu(tf.matmul(X, W1) + b1)
w1_hist = tf.summary.histogram("weights1", W1)
b1_hist = tf.summary.histogram("biases1", b1)
layer1_hist = tf.summary.histogram("layer1", layer1)
tf.add_to_collection('vars', W1)
layer1 = tf.nn.dropout(layer1, keep_prob=keep_prob)
with tf.name_scope("layer2") as scope:
W2 = tf.get_variable("weight2", shape = [50, 25],
initializer=tf.contrib.layers.xavier_initializer())
b2 = tf.Variable(tf.random_normal([25]), name='bias2')
layer2 = tf.nn.relu(tf.matmul(layer1, W2) + b2)
tf.add_to_collection('vars', W2)
w2_hist = tf.summary.histogram("weights2", W2)
b2_hist = tf.summary.histogram("biases2", b2)
layer2_hist = tf.summary.histogram("layer2", layer2)
layer2 = tf.nn.dropout(layer2, keep_prob=keep_prob)
with tf.name_scope("layer3") as scope:
W3 = tf.get_variable("weight3", shape = [25, 10],
initializer=tf.contrib.layers.xavier_initializer())
b3 = tf.Variable(tf.random_normal([10]), name='bias3')
layer3 = tf.nn.relu(tf.matmul(layer2, W3) + b3)
tf.add_to_collection('vars', W3)
w3_hist = tf.summary.histogram("weights3", W3)
b3_hist = tf.summary.histogram("biases3", b3)
layer3_hist = tf.summary.histogram("layer3", layer3)
layer3 = tf.nn.dropout(layer3, keep_prob=keep_prob)
with tf.name_scope("layer4") as scope:
W4 = tf.get_variable("weight4", shape = [10, 1],
initializer=tf.contrib.layers.xavier_initializer())
b4 = tf.Variable(tf.random_normal([1]), name='bias4')
hypothesis = tf.matmul(layer3, W4) + b4
tf.add_to_collection('vars', W4)
w4_hist = tf.summary.histogram("weights4", W4)
b4_hist = tf.summary.histogram("biases4", b4)
hypothesis_hist = tf.summary.histogram("hypothesis", hypothesis)
# cost/loss function
with tf.name_scope("cost") as scope:
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=hypothesis, labels=Y))
cost_summ = tf.summary.scalar("cost", cost)
with tf.name_scope("train") as scope:
train = tf.train.AdamOptimizer(learning_rate=learning_rate).minimize(cost)
# Launch graph
with tf.Session() as sess:
# tensorboard --logdir=./logs/xor_logs
merged_summary = tf.summary.merge_all()
writer = tf.summary.FileWriter("./logs/recommendation")
writer.add_graph(sess.graph) # Show the graph
# Initialize TensorFlow variables
sess.run(tf.global_variables_initializer())
for step in range(10001):
summary, _ = sess.run([merged_summary, train], feed_dict={X: X_train, Y: y_train, keep_prob: 0.7})
writer.add_summary(summary, global_step=global_step)
global_step = 1
global_step +=10
if step % 100 == 0:
print(step, sess.run(cost, feed_dict={
X: X_train, Y: y_train}), sess.run([W1, W2, W3, W4]))
print('Learning Finished!')
correct_prediction = tf.equal(tf.argmax(hypothesis, 1), tf.argmax(Y, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
print('Accuracy:', sess.run(accuracy, feed_dict={
X: X_test, Y: y_test, keep_prob: 1.0}))
# -
# #### Tensorboard
tensorboard --logdir=./logs
# #### Save Trained Tensorflow Model
saver = tf.train.Saver()
sess = tf.Session()
sess.run(tf.global_variables_initializer())
saver.save(sess, 'trained_model')
# #### Run Trained Tensorflow Model
sess = tf.Session()
new_saver = tf.train.import_meta_graph('trained_model.meta')
new_saver.restore(sess, tf.train.latest_checkpoint('./'))
all_vars = tf.get_collection('vars')
for v in all_vars:
v_ = sess.run(v)
print(v_)
# #### Dealing with Large Data
# +
filename_queue = tf.train.string_input_producer([...])
reader = tf.TextLineReader()
_, line = reader.read(filename_queue)
line = tf.decode_csv(line, record_defaults=default)
label_batch, feature_batch = tf.train.shuffle_batch([label, feature], batch_size=batch_size, capacity=512, min_after_dequeue=256, num_threads=8)
| Music_recommendation/3. Building Prediction Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=false editable=false heading_collapsed=true id="7P66a3NStetI" run_control={"frozen": true}
# import matplotlib.pyplot as plt
# # ^^^ pyforest auto-imports - don't write above this line
# # CE-40717: Machine Learning
# + deletable=false editable=false heading_collapsed=true hidden=true run_control={"frozen": true}
# ## HW4-MultiLayer Perceptron (MLP)
# + [markdown] hidden=true id="ZsX3itNwTc8T"
# The following lines of code will load the [MNIST](http://yann.lecun.com/exdb/mnist/) data and turn them
# into numpy arrays, you can print their shape if you like.
# You can also transform the data as you wish, including seperating
# the training data for cross validation.
#
# If you have the data (on google drive or locally) change the root
# address accordingly, if you don't, set download=True but you might encounter
# some problems downloading the data.
# + hidden=true id="eRDGolwttJJr"
# <NAME> (99210259)
import torchvision.datasets as ds
from sklearn.utils import shuffle
import numpy as np
import pandas as pd
data_train = np.array(ds.MNIST(root="./data", train=True, download=True).data)
target_train = np.array(ds.MNIST(root="./data", train=True, download=True).targets)
data_test = np.array(ds.MNIST(root="./data", train=False, download=True).data)
target_test = np.array(ds.MNIST(root="./data", train=False, download=True).targets)
#data_train, target_train = shuffle(data_train, target_train)
#### Transform the data! ####
data_train =data_train / 255
data_test = data_test / 255
target_train=pd.get_dummies(target_train).values
target_test = pd.get_dummies(target_test).values
data_train=data_train.reshape((-1,28*28))
data_test = data_test.reshape((-1,28*28))
# -
from IPython.core.debugger import set_trace
# + deletable=false editable=false heading_collapsed=true hidden=true id="OQ0i1tVuT3bb" run_control={"frozen": true}
# ### Part1:
# Complete the functions of the MLP class to create
# a MultiLayer Perceptron
# +
def sigmoid(x, derivative=False):
if derivative:
return (np.exp(-x))/((np.exp(-x)+1)**2)
return 1/(1 + np.exp(-x))
def softmax(x):
exps = np.exp(x - x.max())
return exps / np.sum(exps, axis=0)
def ReLU(x,derivative=False):
if derivative:
return x>0
return x * (x>0)
def safe_ln(x, minval=0.0000000001):
return np.log(x.clip(min=minval))
def calculate_accuracy(prediction,real):
predictedNumber = np.argmax(prediction,axis=0)
realNumber = np.argmax(real,axis=0)
return np.mean(predictedNumber == realNumber)
# + hidden=true id="86AdE8SntShx"
class MLP:
def __init__(self, in_dimensions, hidden_dimensions, out_dimensions):
self.w1 = np.random.normal(size=(hidden_dimensions, in_dimensions)) / np.sqrt(hidden_dimensions)
self.b1 = np.random.normal(size=(hidden_dimensions,1)) / np.sqrt(hidden_dimensions)
self.w2 = np.random.normal(size=(out_dimensions,hidden_dimensions)) /np.sqrt(out_dimensions)
self.b2 = np.random.normal(size=(out_dimensions,1)) /np.sqrt(out_dimensions)
def compute_loss(self,Y):
Y_hat = self.a2
L_sum = np.sum(np.multiply(Y, np.log(Y_hat+1e-10)))
m = Y.shape[1]
L = -(1/m) * L_sum
return L
def forward(self, x):
# perform a forward pass of the network and return the result
# remember to retain the value of each node (i.e. self.h1_forward)
# in order to use in backpropagation
# Use whatever activation function you wish for the first layer
# and softmax activation for the output layer
self.a0 = x.T
self.z1 = self.w1 @ self.a0 + self.b1
self.a1 = ReLU(self.z1,derivative=False)
self.z2 = self.w2 @ self.a1 + self.b2
self.a2 = softmax(self.z2)
return self.a2
def backward(self, y_target,batch_size):
# perform backpropagation on the loss value and compute the gradient
# w.r.t. every element of the network and retain them (i.e. self.w1_backward)
dZ2 = self.a2 - y_target
self.w2_backward = (1./batch_size) * dZ2 @ self.a1.T
self.b2_backward = (1./batch_size) * np.sum(dZ2)
dA1 = self.w2.T @ dZ2
dZ1 = dA1 * ReLU(self.z1,derivative=True)
self.w1_backward = (1./batch_size) * dZ1 @ self.a0.T
self.b1_backward = (1./batch_size) * np.sum(dZ1)
def step(self, lr, lam):
# simply update all the weights using the gradinets computed in backward
# and the given learning rate with SGD
# don't forget to use regularization
self.w2 = self.w2 - (lr * self.w2_backward - self.w2*lam*lr)
self.b2 = self.b2 - (lr * self.b2_backward - self.b2*lam*lr)
self.w1 = self.w1 - (lr * self.w1_backward - self.w1*lam*lr)
self.b1 = self.b1 - (lr * self.b1_backward - self.b1*lam*lr)
# + deletable=false editable=false heading_collapsed=true hidden=true id="EeMLiOlMUC2D" run_control={"frozen": true}
# ### Part2:
# Make instances of your network and train them **using l2 regularization and choose the lambda using k-fold cross validation
# (set the candidate lambda as you wish)**.
#
# You may choose the hyperparameters (i.e. num of epochs, learning rate etc.)
# as you wish.
#
# Then train a final model on all the training data with the chosen lambda.
#
# + hidden=true id="0Ojg9CSL4vei" tags=["outputPrepend"]
n_epochs =200 # number of epochs
lr =0.05 # learning rate
k = 4 # number of folds
in_dim =28*28 # MNIST has 28*28 images
hidden_dim = 64 # number of hidden dimensions for the hidden layer
out_dim = 10 # MNIST has 10 classes
fold_len = int(data_train.shape[0]/k)
lambdas = [1e-1,1e-2,1e-3,1e-4]
best_lambda = lambdas[-1]
best_acc = 0
for l in lambdas:
acc = 0 # accuracy for current lambda
loss = 0 # loss for current lambda
for j in range(k):
mlp = MLP(in_dim,hidden_dim,out_dim)
separated=slice(j*fold_len,(j+1)*fold_len)
fold_train_set = np.delete(data_train,separated,axis=0) # the training data for the current fold
fold_train_target =np.delete(target_train,separated,axis=0) # the training targets for the current fold
val_set =data_train[separated,:] # the validation data for the current fold
val_target =target_train[separated,:] # the validation targets for the current fold
for i in range(n_epochs):
# train the model on the data with the curent lambda
mlp.forward(fold_train_set)
#cost = mlp.compute_loss(fold_train_target.T)
mlp.backward(fold_train_target.T,fold_train_target.shape[0])
mlp.step(lr,l)
prediction=np.argmax(mlp.forward(val_set),axis=0)
labels = np.argmax(val_target.T,axis=0)
# test the model on the current validation data
fold_acc = np.sum(prediction == labels) / prediction.shape[0] # current fold accuracy
fold_loss = mlp.compute_loss(val_target.T) # current fold loss
print('fold no:' ,j,'fold acc: ',fold_acc,'fold_loss: ', fold_loss)
acc =acc+ fold_acc
loss = loss + fold_loss
acc =100* acc / k
loss = loss / k
print("Lambda:", l)
print("Loss: %.4f Accuracy: %.4f" % (loss, acc))
print()
if acc > best_acc:
best_acc = acc
best_lambda = l
print("Best lambda is",best_lambda, "with %.4f accuracy" % best_acc)
# + deletable=false editable=false heading_collapsed=true hidden=true id="6cDg4S27xD5Y" run_control={"frozen": true}
# ### Part3:
# Train a final model using the best lambda on all the training data
# + hidden=true id="fE1mC1BkxMdt"
n_epochs = 300
lr = 0.05
accuracies = []
losses =[]
model = MLP(in_dim,hidden_dim,out_dim)
for i in range(n_epochs):
#### training code here ####
prediction=np.argmax(model.forward(data_train),axis=0)
model.backward(target_train.T,target_train.shape[0])
model.step(lr,best_lambda)
loss=model.compute_loss(target_train.T)
accuracy = np.sum(prediction == np.argmax(target_train.T,axis=0)) / target_train.shape[0]
losses.append(loss)
accuracies.append(accuracy)
if (i % 20==0) or (i == n_epochs-1):
print('Epoch ',i, 'Loss: ' ,loss,'Accuracy:' ,accuracy)
# + deletable=false editable=false heading_collapsed=true hidden=true run_control={"frozen": true}
# ### Part4:
# + [markdown] hidden=true id="6X8hFKXQUeml"
# Plot the training loss value and accuracy (mean over all batches each epoch if you're using mini-batches) over epochs
# for the final model that is trained on all the training data
# + hidden=true id="_LpeQU225eGi"
X = np.arange(0,300)
loss_array = np.array(losses)
plt.plot(X,loss_array, label = 'plot of loss decay!')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# -
X = np.arange(0,300)
accuracy_array = np.array(accuracies)
plt.plot(X,accuracy_array,label='Accuracy increse over epochs')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.show()
# + [markdown] hidden=true id="WIaBwatyUqmC"
# Use your network on the test set and report the accuracy, you must get at least 70% accuracy on the test set.
# + hidden=true id="NmQiBh4C5ULJ"
calculate_accuracy(model.forward(data_test),target_test.T)
# + [markdown] hidden=true id="zyp8Wgx_nV-A"
# Below you can add code cells and improve on the network structure as you see fit (it still must be an MLP), train and test your network and explain why it works better.
#
# -
class MyMultiLayerPerceptron:
def __init__(self,in_dim,hidden_dim_1,hidden_dim_2,out_dim):
self.w1 = np.random.normal(size=(hidden_dim_1,in_dim)) * np.sqrt(1/hidden_dim_1)
self.b1 = np.random.normal(size=(hidden_dim_1,1)) * np.sqrt(1/hidden_dim_1)
self.w2 = np.random.normal(size=(hidden_dim_2,hidden_dim_1)) * np.sqrt(1/hidden_dim_2)
self.b2 = np.random.normal(size=(hidden_dim_2,1))* np.sqrt(1/hidden_dim_2)
self.w3 = np.random.normal(size=(out_dim,hidden_dim_2)) * np.sqrt(1/out_dim)
self.b3 = np.random.normal(size=(out_dim,1)) * np.sqrt(1/out_dim)
def forward(self,x):
self.a0 = x.T
self.z1 = self.w1 @ self.a0 + self.b1
self.a1 = ReLU(self.z1)
self.z2 = self.w2 @ self.a1 + self.b2
self.a2 =ReLU(self.z2)
self.z3 = self.w3 @ self.a2 + self.b3
self.a3 = softmax(self.z3)
self.prediction = self.a3
return self.prediction
def backward(self,Y_target):
batch_size = Y_target.shape[1]
dZ3 = self.a3 - Y_target
self.w3_backward = (1./batch_size) * dZ3 @ self.a2.T
self.b3_backward = (1./batch_size) * np.sum(dZ3)
dA2 = self.w3.T @ dZ3
dZ2 = dA2 * ReLU(self.z2,derivative=True)
self.w2_backward = (1./batch_size) * dZ2 @ self.a1.T
self.b2_backward = (1./batch_size) * np.sum(dZ2)
dA1 = self.w2.T @ dZ2
dZ1 = dA1 * ReLU(self.z1,derivative=True)
self.w1_backward = (1./batch_size) * dZ1 @ self.a0.T
self.b1_backward = (1./batch_size) * np.sum(dZ1)
def step(self,lr,lam):
self.w3 = self.w3 - (lr * self.w3_backward - self.w3*lam*lr)
self.b3 = self.b3 - (lr * self.b3_backward - self.b3*lam*lr)
self.w2 = self.w2 - (lr * self.w2_backward - self.w2*lam*lr)
self.b2 = self.b2 - (lr * self.b2_backward - self.b2*lam*lr)
self.w1 = self.w1 - (lr * self.w1_backward - self.w1*lam*lr)
self.b1 = self.b1 - (lr * self.b1_backward - self.b1*lam*lr)
def compute_loss(self,Y):
Y_hat = self.prediction
L_sum = np.sum(np.multiply(Y, np.log(Y_hat+1e-10)))
m = Y.shape[1]
L = -(1/m) * L_sum
return L
# +
model = MyMultiLayerPerceptron(28*28,128,64,10)
learning_rate = 0.6
regularization = 1e-3
epochs = 500
for i in range(epochs):
model.forward(data_train)
model.backward(target_train.T)
model.step(learning_rate,regularization)
if (i% 10 == 0) or (i == epochs-1):
loss=model.compute_loss(target_train.T)
accuracy=calculate_accuracy(model.prediction,target_train.T)
print(f'Epoch: {i},Loss: {loss}, Accuracy: {accuracy}')
# -
calculate_accuracy(model.forward(data_test),target_test.T)
# #### Here I simply added a hidden layer to neural network and I've set it's size to 128 and I've initialized the network differently for better results. Neural Network layer and other paramaters are chosen by trial and error.
| HW4/Practical/ML_HW4_AmirPourmand.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lesson 1 - Introduction to Pandas
# ## Lesson Video:
#hide_input
from IPython.lib.display import YouTubeVideo
from datetime import timedelta
start = int(timedelta(minutes=0, seconds=0).total_seconds())
YouTubeVideo('liTHAhdl1cQ', start=start)
#hide
#Run once per session
# !pip install fastai wwf -q --upgrade
#hide_input
from wwf.utils import state_versions
state_versions(['fastai','pandas'])
# ## Intro to Tabular and Pandas
#
# Before we begin doing tabular modeling, let's learn about the `Pandas` library
# Pandas? Like the bear? No!
#
# `pandas` is a library we can use for reading and analyzing any bit of Tabular data. We'll work out of the newly released 1.0 version
import pandas as pd
# We do this by creating what are called `DataFrames`. These can come from a variety of forms. Most commonly:
#
# * `read_csv`
# * `read_excel`
# * `read_feather`
# * `read_html`
# * `read_json`
# * `read_pickle`
# * `read_sql`
# * `read_table`
# We'll focus on the `read_csv` function. We'll use the Titanic dataset for today's tutorial
# ## Titanic
# !git clone https://github.com/pcsanwald/kaggle-titanic.git
# Let's use the `Pathlib` library to read our data
from fastai.basic_data import pathlib
path = pathlib.Path('kaggle-titanic')
# And look at what we grabbed
path.ls()
# We can see that we have a `train.csv` and `test.csv` file. We'll work out of the `train` file today. Let's make our `DataFrame`
df = pd.read_csv(path/'train.csv')
# And now for some common functionalities:
# ## Pandas Functionalities:
# ### Head and Tail
#
# `.head()` and `.tail()` will show the first and last few rows of a `DataFrame`. You can pass in `n` rows to look at (the default is 5)
df.head(n=4)
df.tail(n=3)
# ## Selecting
#
# We can select a variety of ways in pandas: row, value, even by a column:
# ### Row:
# There are two different ways we can select rows, `loc` and `iloc`. Each work a little differently
# ### `loc`
# `loc` is used to get rows (or columns) with a particular **label** from an index. IE:
df.loc[:3]
# Here we grabbed rows 0 through 3
# ### `iloc`
# `iloc` is used when we want to get rows (or columns) from a *position* on an index
df.iloc[:3]
# Whereas here we grabbed the first three rows
# ### Column
#
# We can pass in either a string, an index, or multiple columns to select:
df['survived'].head()
# To select multiple columns, pass in a double array of your names
df[['sex', 'age', 'survived']].head()
# And to pass as an index, first do the number of rows followed by the column index
df.iloc[:,0]
df.iloc[:,0:3]
# ### Value
#
# We can select based on a value a few different ways, most involving a boolean argument:
# +
# df.loc[df['column_name'] == some_value]
# -
df.loc[df['sex'] == 'female'].head()
# Or even a series of values:
df.loc[(df['sex'] == 'female') & (df['survived'] == 0)].head()
# Now that we have the basic selects done, let's go into some more complex ideas
# ## Plotting
#
# We can plot out any data we want from our `DataFrames`, like so:
df['pclass'].iloc[:5].plot(kind='bar', title='pclass')
df['fare'].plot(title='fare')
# We can also plot multiple columns:
df[['survived', 'pclass']][:20].plot(kind='bar')
# ## Dealing with multiple `DataFrames`
#
# Sometimes, we have multiple dataframes of data. How do we combine them?
df1 = df[['sibsp', 'parch', 'ticket', 'fare', 'cabin', 'embarked', 'name']].iloc[:50]
df2 = df[df.columns[:6]].iloc[:50]
# ### Merge
#
# `merge` allows you to do standard database operations on a `DataFrame` or `Series`. Here is the doc line, let's break it down:
# ```python
# pd.merge(left, right, how='inner', on=None, left_on=None, right_on=None,
# left_index=False, right_index=False, sort=True,
# suffixes=('_x', '_y'), copy=True, indicator=False,
# validate=None)
# ```
# * `left` and `right` should be two `DataFrame` or `Series` objects.
# * `on` is a **column** or **index** name to join on and must be found in **both** dataframes.
# * `left_index` and `right_index` uses the respective input's index to use as a key instead
# * `how`: Either `left`, `right`, `outer`, or `inner`.
df1.head()
df2.head()
# Let's practice merging on `name`
merge = pd.merge(df1, df2, on='name')
merge.head()
# ### Appending
#
# We can tag one dataframe onto another:
df_top = df.iloc[:5]
df_bottom = df.iloc[5:10]
len(df_top), len(df_bottom)
df_top.append(df_bottom)
# ## Dropping
#
# You can drop a column or row, and by default `pandas` will return the dropped table. If you choose to do `inplace`, it will directly modify the dataframe. You can also pass in an `axis` parameter. By default it is `1`, which will drop a row. `0` drops a column
df_top.drop(0)
df_top.drop('survived', axis=1)
df_top.head()
# ## GroupBy
#
# GroupBy can be used to split our data into groups if they fall into some criteria.
# ```python
# df_top.groupby(by=None, axis=0, level=None, as_index: bool=True, sort: bool=True, group_keys: bool=True, squeeze: bool=False, observed: bool=False)
# ```
# Parameters:
# * `by`: mapping, function, string or some iterable
# * `axis`: default is 0
# * `level`: If the axis is heirarchical, group by levels or a particular level
# * `group_keys`: Add group keys to index to identify pieces
# For an example, let's group by `survived`:
surv = df.groupby('survived')
# If we call `first`, we'll see the first entry in each group:
surv.first()
# To grab the group, we can call `get_group` and pass in either one of our classes:
surv.get_group(1)
# We can also group by multiple columns. Let's combine both `survived` and `sex`
surs = df.groupby(['survived', 'sex'])
surs.first()
# Now we can further analyze and split our data based on whatever queries we want to use!
# ## Map
#
# `map` can be used to map values from two different `series` (column) that share a same column. Basically we can repeat our adjust values:
type(df['survived'])
df['survived'].head()
# We can use a function or a dictionary:
df['survived'].map({0:'dead', 1:'survived'}).head()
df['survived'].map('I am a {}'.format).head()
# ## .apply
# Apply let's you pass in a function to apply to every value in a series. It takes in a function, `convert_dtype`, and some argumetns
# Let's make a basic one that returns if someone survived:
def lived(num):
if num == 0:
return "Died"
else:
return "Lived"
new = df['survived'].apply(lived)
new.head()
# We can also use a lambda:
new = df['age'].apply(lambda age: age + 5)
print(f'{df["age"].head()}\n{new.head()}')
| nbs/course2020/tabular/01_Pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="VSgzfxo5DmJy"
# ### Question 13
# A transformer having 145 turns in its primary winding has a 60 Hz input voltage of $V_{in} = 75 VAC$. It is desired that $V_{out}$ should be $120 VAC$. How many turns should the secondary winding have?
#
# __Given__
# - $N_p=145~turns~~~~~~~~~~~~~~~~$ The primary number of turns
# - $\epsilon_p=75~V~~~~~~~~~~~~~~~~~~~~~~~~~~~~$The primary emf of the transformer
# - $\epsilon_s=120V~~~~~~~~~~~~~~~~~~~~~~~~~~$ The secondary emf of the transformer
#
# __Formula__
# - $\frac{N_p}{N_s}=\frac{\epsilon_p}{\epsilon_s}~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~$ Topic 11.2
#
# __Solution__
# - We have the primary turns of the transformer, the emfs in and out for the transformer. To calculate the number of secondary turn is just need to subtitute the values and calculate.
# - $\frac{N_p}{N_s}=\frac{\epsilon_p}{\epsilon_s}=\frac{75V}{120V}=\frac{145turns}{N_s}$
# - $N_s=\frac{120V}{75V}\times145~turns$
#
# __Answer__
#
# The number of turn should be $62.1~turns$
# + id="QGEk3L3LDZy2" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5959fb87-25c1-4d8e-81e7-2d8074ace2fb"
N_p = 145 # The primary number of turns
epsilon_p = 75
epsilon_s = 120
N_s = epsilon_s*epsilon_p/N_p
print('The number of turn should be {:.1f} turns'.format(N_s))
a = 3 # voltage in volt
b = 7 # current in ampere
R = a / b # resistance in Ohm
print(“The resistance is {:.7f} Ohm.”.format(R))
| formative_ipynb-solution/11 Electromagnetic induction/11_2_13.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# [View in Colaboratory](https://colab.research.google.com/github/zhangbingxyx/deep-learning-coursera/blob/master/zhangbing_exp_1.ipynb)
# + id="ZHU3lb3q_M64" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 53} outputId="5261541a-de7e-4cd9-b9ed-a7de8c60f7bc"
import tensorflow as tf
input1 = tf.ones((2, 3))
input2 = tf.reshape(tf.range(1, 7, dtype=tf.float32), (2, 3))
output = input1 + input2
with tf.Session():
result = output.eval()
result
# + [markdown] id="080mezDk_U7_" colab_type="text"
#
| zhangbing_exp_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Paul-mwaura/ML-Hackathons/blob/main/Netflix.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="_71vSqORLwTq"
import os
import numpy as np # linear algebra
import seaborn as sns
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
import tensorflow as tf
from tensorflow import keras
from keras.models import Model
from keras.layers import Input, Reshape, Dot
from keras.layers.embeddings import Embedding
from keras.layers import Concatenate, Dense, Dropout
from keras.optimizers import Adam
from keras.regularizers import l2
# + colab={"base_uri": "https://localhost:8080/"} id="ELc2rrqoMw4r" outputId="f2a0862b-8407-407c-8ae2-79a85c26225b"
# DataFrame to store all imported data
if not os.path.isfile('data.csv'):
data = open('data.csv', mode='w')
files = ['/content/combined_data_1.txt',
'/content/combined_data_2.txt',
'/content/combined_data_3.txt',
'/content/combined_data_4.txt']
# Remove the line with movie_id: and add a new column of movie_id
# Combine all data files into a csv file
for file in files:
print("Opening file: {}".format(file))
with open(file) as f:
for line in f:
line = line.strip()
if line.endswith(':'):
movie_id = line.replace(':', '')
else:
data.write(movie_id + ',' + line)
data.write('\n')
data.close()
# + colab={"base_uri": "https://localhost:8080/", "height": 414} id="lpg12YdTYFP_" outputId="55649304-c2a3-4f74-d11c-18e643707a99"
# Read all data into a pd dataframe
df = pd.read_csv('data.csv', names=['movie_id',
'user_id',
'rating',
'date'])
df
# + colab={"base_uri": "https://localhost:8080/"} id="AMCydJ1qS4-e" outputId="93c5e761-282f-4f24-ac93-b90226e09d6b"
df.dropna(inplace=True)
df.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="sSLOwMPwVBbf" outputId="ad787e0b-6b3d-47ea-de08-691addf9c959"
df.duplicated().sum()
# + id="pKaI4B5-Vp7F" colab={"base_uri": "https://localhost:8080/", "height": 414} outputId="53817314-aee8-42da-b549-862d328d4cd1"
#no need to use the df_movie
#continue df
#pre-process stage
lite_rating_df = pd.DataFrame()
group = df.groupby('user_id')['rating'].count()
top_users = group.sort_values(ascending=False)[:20000]
group = df.groupby('movie_id')['rating'].count()
top_movies = group.sort_values(ascending=False)[:4000]
lite_rating_df = df.join(top_users, rsuffix='_r', how='inner', on='user_id')
lite_rating_df = lite_rating_df.join(top_movies, rsuffix='_r', how='inner', on='movie_id')
# Re-name the users and movies for uniform name from 0..2000 and 10000
user_enc = LabelEncoder()
lite_rating_df['user'] = user_enc.fit_transform(lite_rating_df['user_id'].values)
movie_enc = LabelEncoder()
lite_rating_df['movie'] = movie_enc.fit_transform(lite_rating_df['movie_id'].values)
n_movies = lite_rating_df['movie'].nunique()
n_users = lite_rating_df['user'].nunique()
# print(n_movies, n_users)
lite_rating_df
# + colab={"base_uri": "https://localhost:8080/"} id="w0vh6zxSblkN" outputId="aa25ee2c-1974-4545-acd2-4de5c3d36356"
top_users[:5]
# + colab={"base_uri": "https://localhost:8080/"} id="dEfI14ycbxEi" outputId="7bb06cac-e6f8-4b70-fb65-b450933ce24c"
top_movies[:5]
# + colab={"base_uri": "https://localhost:8080/"} id="YRAeMDnfcRVA" outputId="45632936-c390-4335-bd64-36a9829a1412"
lite_rating_df.isnull().sum()#dataset no null values
# + id="7HCJQWkAcm0J" colab={"base_uri": "https://localhost:8080/"} outputId="187c27a1-dce6-433e-e1dc-ff5acd7328dc"
X = lite_rating_df[['user', 'movie']].values
y = lite_rating_df['rating'].values
# Split train and test data (for test model performance at last)
X_training, X_test, y_training, y_test = train_test_split(X, y, test_size=0.1)
# Split train and validation data (to monitor model performance in training)
X_train, X_val, y_train, y_val = train_test_split(X_training, y_training, test_size=0.1)
# Set the embedding dimension d of Matrix factorization
e_dimension = 50
X_train_array = [X_train[:, 0], X_train[:, 1]]
X_val_array = [X_val[:, 0], X_val[:, 1]]
X_test_array = [X_test[:, 0], X_test[:, 1]]
print(f"X Train: {X_train.shape}\nX Test: {X_test.shape}\n")
print(f"Y Train: {Y_train.shape}\nY Test: {Y_test.shape}\n")
# + id="8agW02v0e4CM" colab={"base_uri": "https://localhost:8080/"} outputId="a7a582cf-1416-4646-fd86-90b71f97e384"
# Build user and movie embedding matrix
user = Input(shape=(1,))
u = Embedding(n_users, e_dimension, embeddings_initializer='he_normal',
embeddings_regularizer=l2(1e-6))(user)
u = Reshape((e_dimension,))(u)
movie = Input(shape=(1,))
m = Embedding(n_movies, e_dimension, embeddings_initializer='he_normal',
embeddings_regularizer=l2(1e-6))(movie)
m = Reshape((e_dimension,))(m)
x = Dot(axes=1)([u, m])
# Build last deep learning layers
x = Dense(128, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(1)(x)
model = Model(inputs=[user, movie], outputs=x)
model.compile(loss='mean_squared_error',
optimizer=Adam(lr=0.001),
metrics=[tf.keras.metrics.Accuracy()]
)
# Set up for early stop if the validation loss stop improving for more than 1 epoch
callbacks_list = [keras.callbacks.EarlyStopping(monitor='val_loss',
patience=1,
),
# Saves the weights after every epoch
keras.callbacks.ModelCheckpoint(
filepath='Model_1',
monitor='val_loss',
save_best_only=True,
)]
# Print model info summary
model.summary()
history = model.fit(x=X_train_array, y=y_train, batch_size=64, epochs=5,
verbose=1,
callbacks=callbacks_list,
validation_data=(X_val_array, y_val)
)
# Save the model (we should make a good habit of always saving our models after training)
model.save("Model_1")
# + id="m8QDPjkhhrI0" colab={"base_uri": "https://localhost:8080/", "height": 294} outputId="d6c07287-8873-4ba0-dcf4-c439d1e9d52f"
# Visualize the training and validation loss
history_dict = history.history
loss_values = history_dict['loss']
val_loss_values = history_dict['val_loss']
epochs = range(1, len(loss_values) + 1)
plt.plot(epochs, loss_values, 'ro', label='Training loss')
plt.plot(epochs, val_loss_values, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# + id="l8lFoz_Dsc6L" colab={"base_uri": "https://localhost:8080/"} outputId="9463b1c8-806a-4ae7-ca80-4da45f273884"
m = tf.keras.metrics.RootMeanSquaredError()
m.update_state(model.predict(X_test_array), y_test)
m.result().numpy()
# + [markdown] id="yzciCoqND9dM"
# In the test result, we can see that our model's RMSE is 0.87932473, which is quite good
| Netflix.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Global Validation ###
#
# This notebook combines several validation notebooks: `global_validation_tasmax_v2.ipynb` and `global_validation_dtr_v2.ipynb` along with `check_aiqpd_downscaled_data.ipynb` to create a "master" global validation notebook. It also borrows validation code from the ERA-5 workflow, `validate_era5_hourlyORdaily_files.ipynb`. It is intended to be run with `papermill`.
#
# ### Data Sources ###
#
# Coarse Resolution:
# - CMIP6
# - Bias corrected data
# - ERA-5
#
# Fine Resolution:
# - Bias corrected data
# - Downscaled data
# - ERA-5 (fine resolution)
# - ERA-5 (coarse resolution resampled to fine resolution)
#
# ### Types of Validation ###
#
# Basic:
# - maxes, means, mins
# - CMIP6, bias corrected and downscaled
# - historical (1995-2014), 2020-2040, 2040-2060, 2060-2080, 2080-2100
# - differences between historical and future time periods for bias corrected and downscaled
# - differences between bias corrected and downscaled data
#
# Variable-specific:
# - GMST
# - days over 95 (TO-DO)
# - max # of consecutive dry days, highest precip amt over 5-day rolling window (TO-DO)
# +
# %matplotlib inline
import xarray as xr
import numpy as np
import matplotlib.pyplot as plt
from cartopy import config
import cartopy.crs as ccrs
import cartopy.feature as cfeature
import os
import gcsfs
from matplotlib import cm
import warnings
from validation import *
# -
# ### Set Validation Parameters ###
# + tags=["parameters"]
# variable options: 'tasmax', 'tasmin', 'dtr', 'pr'
variable = 'tasmax'
# ssp options: 'ssp126', 'ssp245', 'ssp370', 'ssp585'
ssp = 'ssp370'
# data output types for running validation
cmip6 = True
bias_corrected = True
downscaled = True
# projection time period options: '2020_2040', '2040_2060', '2060_2080', '2080_2100'
projection_time_period = '2080_2100'
# validation plot options
basic_diagnostics = True
# options: 'mean', 'max', 'min'
basic_diag_type = 'mean'
gmst = True
difference_plots = True
# options: 'downscaled_minus_biascorrected' , 'change_from_historical'
diff_type = 'downscaled_minus_biascorrected'
# contains the gcs URLs to zarr locations for each specified dataset
data_dict = {'coarse': {'cmip6': {'historical': 'scratch/biascorrectdownscale-bk6n8/biascorrectdownscale-bk6n8-858077599/out.zarr',
ssp: 'scratch/biascorrectdownscale-bk6n8/biascorrectdownscale-bk6n8-269778292/out.zarr'},
'bias_corrected': {'historical': 'az://biascorrected-stage/CMIP/NOAA-GFDL/GFDL-ESM4/historical/r1i1p1f1/day/tasmax/gr1/v20210920214427.zarr',
ssp: 'az://biascorrected-stage/ScenarioMIP/NOAA-GFDL/GFDL-ESM4/ssp370/r1i1p1f1/day/tasmax/gr1/v20210920214427.zarr'},
'ERA-5':'az://scratch/biascorrectdownscale-bk6n8/biascorrectdownscale-bk6n8-131793962/out.zarr'},
'fine': {'bias_corrected': {'historical': 'az://scratch/biascorrectdownscale-bk6n8/biascorrectdownscale-bk6n8-1362934973/regridded.zarr',
ssp: 'az://scratch/biascorrectdownscale-bk6n8/biascorrectdownscale-bk6n8-377595554/regridded.zarr'},
'downscaled': {'historical': 'az://downscaled-stage/CMIP/NOAA-GFDL/GFDL-ESM4/historical/r1i1p1f1/day/tasmax/gr1/v20210920214427.zarr',
ssp: 'az://downscaled-stage/ScenarioMIP/NOAA-GFDL/GFDL-ESM4/ssp370/r1i1p1f1/day/tasmax/gr1/v20210920214427.zarr'},
'ERA-5_fine': 'az://scratch/biascorrectdownscale-bk6n8/biascorrectdownscale-bk6n8-491178896/rechunked.zarr',
'ERA-5_coarse': 'az://scratch/biascorrectdownscale-bk6n8/biascorrectdownscale-bk6n8-1213790070/rechunked.zarr'}}
# -
# we only plot gmst if validation variable is tasmax
if variable != 'tasmax':
gmst = False
warnings.warn("gmst plotting option changed to False since validation variable is not tasmax")
# + [markdown] tags=[]
# ### other data inputs ###
# -
units = {'tasmax': 'K', 'tasmin': 'K', 'dtr': 'K', 'pr': 'mm'}
years = {'hist': {'start_yr': '1995', 'end_yr': '2014'},
'2020_2040': {'start_yr': '2020', 'end_yr': '2040'},
'2040_2060': {'start_yr': '2040', 'end_yr': '2060'},
'2060_2080': {'start_yr': '2060', 'end_yr': '2080'},
'2080_2100': {'start_yr': '2080', 'end_yr': '2100'}}
years_test = {'hist': {'start_yr': '1995', 'end_yr': '2014'},
'2020_2040': {'start_yr': '2020', 'end_yr': '2040'},
'2040_2060': {'start_yr': '2040', 'end_yr': '2060'}}
# ### Validation ###
# ### basic diagnostic plots: means, maxes, mins ###
if cmip6 and basic_diagnostics:
plot_diagnostic_climo_periods(read_gcs_zarr(data_dict['coarse']['cmip6']['historical']),
read_gcs_zarr(data_dict['coarse']['cmip6'][ssp]),
ssp, years, variable, basic_diag_type, 'cmip6',
units, vmin=280, vmax=320)
if bias_corrected and basic_diagnostics:
plot_diagnostic_climo_periods(read_gcs_zarr(data_dict['coarse']['bias_corrected']['historical']),
read_gcs_zarr(data_dict['coarse']['bias_corrected'][ssp]),
ssp, years, variable, basic_diag_type, 'bias_corrected',
units, vmin=280, vmax=320)
if downscaled and basic_diagnostics:
plot_diagnostic_climo_periods(read_gcs_zarr(data_dict['coarse']['downscaled']['historical']),
read_gcs_zarr(data_dict['coarse']['downscaled'][ssp]),
ssp, years, variable, basic_diag_type, 'downscaled',
units, vmin=280, vmax=320)
# ### GMST ###
if gmst:
plot_gmst_diagnostic(read_gcs_zarr(data_dict['coarse']['cmip6']['historical']),
read_gcs_zarr(data_dict['coarse']['cmip6'][ssp]),
read_gcs_zarr(data_dict['coarse']['bias_corrected']['historical']),
read_gcs_zarr(data_dict['coarse']['bias_corrected'][ssp]),
variable=variable, ssp=ssp, ds_hist_downscaled=None, ds_fut_downscaled=None)
# ### Difference plots: bias corrected and downscaled OR historical/future (bias corrected and downscaled data outputs) ###
if bias_corrected and difference_plots:
plot_bias_correction_downscale_differences(read_gcs_zarr(data_dict['fine']['bias_corrected']['historical']),
read_gcs_zarr(data_dict['fine']['downscaled']['historical']),
read_gcs_zarr(data_dict['fine']['bias_corrected'][ssp]),
read_gcs_zarr(data_dict['fine']['downscaled'][ssp]),
diff_type, 'bias_corrected', variable,
ssp=ssp, time_period=projection_time_period)
if downscaled and difference_plots:
plot_bias_correction_downscale_differences(read_gcs_zarr(data_dict['fine']['bias_corrected']['historical']),
read_gcs_zarr(data_dict['fine']['downscaled']['historical']),
read_gcs_zarr(data_dict['fine']['bias_corrected'][ssp]),
read_gcs_zarr(data_dict['fine']['downscaled'][ssp]),
diff_type, 'downscaled', variable,
ssp=ssp, time_period=projection_time_period)
# ### TO-DO: Days over 95 degrees F/extreme precip metrics ###
| notebooks/downscaling_pipeline/global_validation_papermill.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Directly specyfing a list
# Thanks to the 'order' parameter of the violinplot function, you can set the order in which you expect the distributions to appear on the figure.
# +
# libraries & dataset
import seaborn as sns
import matplotlib.pyplot as plt
# set a grey background (use sns.set_theme() if seaborn version 0.11.0 or above)
sns.set(style="darkgrid")
df = sns.load_dataset('iris')
# specifying the group list as 'order' parameter and plotting
sns.violinplot(x='species', y='sepal_length', data=df, order=[ "versicolor", "virginica", "setosa"])
plt.show()
# -
# ## Ordering by decreasing median
# Instead of creating the list 'by hand' with various categories, you can also use the power of pandas operations (groupby, median or mean) in order to create a ranked list, which we stored in 'my_order' variable in the following example.
# +
# libraries & dataset
import seaborn as sns
import matplotlib.pyplot as plt
# set a grey background (use sns.set_theme() if seaborn version 0.11.0 or above)
sns.set(style="darkgrid")
df = sns.load_dataset('iris')
# Using pandas methods and slicing to determine the order by decreasing median
my_order = df.groupby(by=["species"])["sepal_length"].median().iloc[::-1].index
# Specifying the 'order' parameter with my_order and plotting
sns.violinplot(x='species', y='sepal_length', data=df, order=my_order)
plt.show()
| src/notebooks/55-control-order-of-groups-in-violinplot-seaborn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
import pandas as pd
from sklearn import preprocessing
from sklearn import tree
from sklearn import preprocessing
from sklearn.metrics import confusion_matrix,accuracy_score
from sklearn.metrics import precision_recall_fscore_support as score
from sklearn.ensemble import RandomForestClassifier
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
# +
df = pd.read_csv("../../Datasets/flight_weather.csv", low_memory=False)
df = df[df['Temp'] != '9999']
# -
print df.shape
df['FLIGHT_DATE']
# +
df['DAY_OF_WEEK'].astype('category')
# -
df.shape
## Converting variables to string, then to factors
#from datetime import datetime
df['SCHEDULED_DEPARTURE_HOURS'] = df['SCHEDULED_DEPARTURE'].apply(lambda x: str(x)[11:13])
df['SCHEDULED_DEPARTURE_HOURS'].astype('category')
print df['SCHEDULED_DEPARTURE_HOURS']
df['SCHEDULED_ARRIVAL']
# +
df['SCHEDULED_ARRIVAL']
df['SCHEDULED_ARRIVAL_Hours'] = df['SCHEDULED_ARRIVAL'].apply(lambda x: str(x)[:2])
df['SCHEDULED_ARRIVAL_Hours'].astype('category')
print df['SCHEDULED_ARRIVAL_Hours']
# -
df['DAY_OF_WEEK'].astype('category')
df.head()
print df.apply(lambda x:len(x.unique()))
# +
#### Selecting the required variables for ML algorithm.
df = df.loc[:, ['DAY_OF_WEEK', 'AIRLINE', 'ORIGIN_AIRPORT', 'DESTINATION_AIRPORT','DEPARTURE_DELAY', 'ELAPSED_TIME', 'AIR_TIME', 'DISTANCE', 'ARRIVAL_DELAY', 'DAY_TYPE', 'DEP_DELAY_BIN', 'SCHEDULED_DEPARTURE_HOURS', 'SCHEDULED_ARRIVAL_Hours', 'Temp', 'Conditions', 'Visibility', 'Wind_speed' ]]
print df
#df = df.drop([''])
print list(df)
df = df.dropna(axis = 0, how = 'any')
# +
### Encoding AIRLINE
le = preprocessing.LabelEncoder()
le.fit(df.iloc[:,1])
col_2_transformed = le.transform(df.iloc[:,1])
#print col_2_transformed
df.iloc[:,1] = col_2_transformed
le1 = preprocessing.LabelEncoder()
df['AIRLINE'].astype('category')
print df.head
# +
### Encoding ORIGIN_AIRPORT
le1 = preprocessing.LabelEncoder()
le1.fit(df.iloc[:,2])
col_3_transformed = le1.transform(df.iloc[:,2])
#print col_2_transformed
df.iloc[:,2] = col_3_transformed
df['ORIGIN_AIRPORT'].astype('category')
print df.head
# +
### Encoding DESTINATION_AIRPORT
le2 = preprocessing.LabelEncoder()
le2.fit(df.iloc[:,3])
col_4_transformed = le2.transform(df.iloc[:,3])
#print col_2_transformed
df.iloc[:,3] = col_4_transformed
df['DESTINATION_AIRPORT'].astype('category')
print df.head
# -
# +
# Encoding SCHEDULED_DEPARTURE_HOURS
### Encoding DESTINATION_AIRPORT
le4 = preprocessing.LabelEncoder()
le4.fit(df.iloc[:,11])
col_5_transformed = le4.transform(df.iloc[:,11])
#print col_2_transformed
df.iloc[:,11] = col_5_transformed
df['SCHEDULED_DEPARTURE_HOURS'].astype('category')
print df.head
# +
### Encoding SCHEDULED_ARRIVAL_Hours
le5 = preprocessing.LabelEncoder()
le5.fit(df.iloc[:,12])
col_6_transformed = le5.transform(df.iloc[:,12])
#print col_2_transformed
df.iloc[:,12] = col_6_transformed
df['SCHEDULED_ARRIVAL_Hours'].astype('category')
print df.head
# +
# Converting the new variables as categorical
df['Temp'] = df['Temp'].apply(lambda x: foo(x))
df['Wind_speed'] = df['Wind_speed'].apply(lambda x: foo(x))
#df['Visibility'] = df['Visibility'].apply(lambda x: foo(x))
df['Conditions'].astype('category')
df['Visibility'].astype('category')
# +
### Encoding Condition
def foo(x):
if x == "Calm" or x =='\n -\n' or x == 'NaN':
return 0
else:
return float(x)
le6 = preprocessing.LabelEncoder()
le6.fit(df.iloc[:,14])
col_7_transformed = le6.transform(df.iloc[:,14])
#print col_2_transformed
df.iloc[:,14] = col_7_transformed
df['Temp'] = df['Temp'].apply(lambda x: foo(x))
df['Visibility'] = df['Visibility'].apply(lambda x : foo(x))
df['Conditions'].astype('category')
df['Wind_speed'] = df['Wind_speed'].apply(lambda x : foo(x))
print df.head
# -
df['Temp'] = df['Temp'].apply(lambda x: foo(x))
# +
# Converting the new variables as categorical
df['Temp'] = df['Temp'].apply(lambda x: foo(x))
df['Wind_speed'] = df['Wind_speed'].apply(lambda x: foo(x))
#df['Visibility'] = df['Visibility'].apply(lambda x: foo(x))
df['Conditions'].astype('category')
df['Visibility'].astype('category')
# -
df.head()
# +
# Selecting the needed variables from the dataframe for training and testing set.
tr_data = df.iloc[:,[0,1,2,3,5,6,7,8,9,11,12,13,14,15,16]]
tr_target = df.iloc[:,10]
print tr_data.head
print tr_target.head
# +
#### Need to do upsampling / downsampling.
# -
# +
### Creating the training and test for validation
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(tr_data, tr_target, test_size = 0.25, random_state = 0, stratify=tr_target)
print X_train.head()
# +
#X_train['Temp'] = X_train['Temp'].apply(lambda x: foo(x))
# +
#X_train['Wind_speed'] = X_train['Wind_speed'].apply(lambda x: foo(x))
# +
#y_train.head()
# +
#df['Visibility'] = X_train['Visibility'].apply(lambda x: foo(x))
# +
#df['Visibility'] = X_train['Visibility'].apply(lambda x: foo(x))
# +
###Decision tree classifier ## "entropy"
clf = tree.DecisionTreeClassifier(criterion='entropy', min_samples_split=5)
clf = clf.fit(X_train, y_train)
#Predicting the results
y_pred = clf.predict(X_test)
#making the confusion matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
print cm
print "Accuracy score"
accuracy_score(y_test, y_pred)
#tree.export_graphviz(clf, out_file='decision_tree.dat',
# feature_names=list(tr_data)[0:15],
# filled=True)
# +
###Decision tree classifier ## "gini"
clf = tree.DecisionTreeClassifier(criterion='gini', min_samples_split=5)
clf = clf.fit(X_train, y_train)
#Predicting the results
y_pred = clf.predict(X_test)
#making the confusion matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
print cm
print "Accuracy score"
accuracy_score(y_test, y_pred)
#tree.export_graphviz(clf, out_file='decision_tree.dat',
# feature_names=list(tr_data)[0:10],# filled=True)
# -
X_train
# +
##### . Logistic regression and SGD
##### . Standardising the continuous variables
sc = StandardScaler()
sc.fit(X_train.iloc[:,[4]])
X_train.iloc[:,4] = sc.transform(X_train.iloc[:,4])
sc.fit(X_train.iloc[:,[5]])
X_train.iloc[:,5] = sc.transform(X_train.iloc[:,5])
sc.fit(X_train.iloc[:,[6]])
X_train.iloc[:,6] = sc.transform(X_train.iloc[:,6])
#sc.fit(X_train.iloc[:,[13]])
#X_train.iloc[:,13] = sc.transform(X_train.iloc[:,13])
#sc.fit(X_train.iloc[:,[16]])
#X_train.iloc[:,16] = sc.transform(X_train.iloc[:,16])
sc.fit(X_test.iloc[:,[4]])
X_test.iloc[:,4] = sc.transform(X_test.iloc[:,4])
sc.fit(X_test.iloc[:,[5]])
X_test.iloc[:,5] = sc.transform(X_test.iloc[:,5])
sc.fit(X_test.iloc[:,[6]])
X_test.iloc[:,6] = sc.transform(X_test.iloc[:,6])
#sc.fit(X_test.iloc[:,[13]])
#X_test.iloc[:,13] = sc.transform(X_test.iloc[:,13])
#sc.fit(X_test.iloc[:,[16]])
#X_test.iloc[:,16] = sc.transform(X_test.iloc[:,16])
X_train
# -
# +
### Random Forest
# -
rf = RandomForestClassifier(n_jobs=2, random_state=0)
y_pred_rf = clf.fit(X_train,y_train)
# +
###Random forest ## "gini"
rf = RandomForestClassifier(n_jobs=2, random_state=0)
rf = clf.fit(X_train, y_train)
#Predicting the results
y_pred = rf.predict(X_test)
#making the confusion matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
print cm
print "Accuracy score"
accuracy_score(y_test, y_pred)
# -
# +
### Logistic regressio and sgd
###Logistic Regression
lr = LogisticRegression( random_state=0)
lr.fit(X_train, y_train)
y_pred = lr.predict(X_test)
#making the confusion matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
print cm
print "Accuracy score"
accuracy_score(y_test, y_pred)
# +
###Stochastic Gradient Descent
from sklearn import linear_model
clf1 = linear_model.SGDClassifier()
clf1.fit(X_train, y_train)
y_pred = clf1.predict(X_test)
#making the confusion matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
print cm
# -
print "Accuracy score"
accuracy_score(y_test, y_pred)
### naive Bayes
X_train_cat = X_train.iloc[:, [0,1,2,3, ]]
X_test_cat = X_test.iloc[:, [0,1,2,3]]
# +
from sklearn.naive_bayes import MultinomialNB
clf = MultinomialNB()
clf.fit(X_train_cat, y_train)
# +
# Continuous var in X_train
X_train_cont = X_train.iloc[:, [5,6,8]]
X_test_cont = X_test.iloc[:, [5,6,8]]
from sklearn.naive_bayes import GaussianNB
clf1 = GaussianNB()
clf1.fit(X_train_cont, y_train)
# -
c = np.multiply(clf.predict_proba(X_test_cat)[1:2000],clf1.predict_proba(X_test_cont)[1:2000])
# +
#Final predicted value of delay vs Non- delay
fin = np.zeros(2000)
#print fin
for i in range(0,len(clf.predict_proba(X_test_cat)[0:1999])):
if c[i][0] > c[i][1]:
fin[i] = 0
else:
fin[i] = 1
fin
y_test[0:10]
# -
pd.crosstab(y_test[0:1999], fin[0:1999])
| IndividualContribution/Mithil/Decision tree_multi_class.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assigning ambiguous counts
#
# +
import matplotlib
import numpy as np
import pandas as pd
import scanpy as sc
import anndata
import scvelo as scv
import scipy
import json
import os
with open('../../configs/config.json') as f:
input_paths = json.load(f)
top_dir = input_paths['top_dir']
frydir = os.path.sep.join([top_dir, "results", "alevin_fry", "mouse_pancreas", "fry_knee_quant_usa_cr-like"])
e2n_path = os.path.sep.join([top_dir, "refs", "refdata-cellranger-mm10-2.1.0", "geneid_to_name.txt"])
os.makedirs("anndata", exist_ok= True)
# +
verbose = True
meta_info = json.load(open(os.path.sep.join([frydir, "meta_info.json"])))
ng = meta_info['num_genes']
usa_mode = meta_info['usa_mode']
if usa_mode:
if verbose:
print("processing input in USA mode, will return A+S as the spliced count, and U as the unspliced count")
else:
print("please follow previous steps to generate the ount matrix in the USA mode")
assert(False)
af_raw = sc.read_mtx(os.path.sep.join([frydir, "alevin", "quants_mat.mtx"]))
ng = int(ng/3)
e2n = dict([ l.rstrip().split() for l in open(e2n_path).readlines()])
var_names = [ l.rstrip() for l in open(os.path.sep.join([frydir, "alevin", "quants_mat_cols.txt"])).readlines()][:ng]
var_names = [e2n[e] for e in var_names]
obs_names = [ l.rstrip() for l in open(os.path.sep.join([frydir, "alevin", "quants_mat_rows.txt"])).readlines() ]
example_adata = scv.datasets.pancreas()
spliced = af_raw[:,range(0,ng)]
spliced.obs_names = obs_names
spliced.var_names = var_names
spliced.var_names_make_unique()
spliced = spliced[example_adata.obs_names, example_adata.var_names]
unspliced = af_raw[:,range(ng, 2*ng)]
unspliced.obs_names = obs_names
unspliced.var_names = var_names
unspliced.var_names_make_unique()
unspliced = unspliced[example_adata.obs_names, example_adata.var_names]
ambiguous = af_raw[:,range(2*ng,3*ng)]
ambiguous.obs_names = obs_names
ambiguous.var_names = var_names
ambiguous.var_names_make_unique()
ambiguous = ambiguous[example_adata.obs_names, example_adata.var_names]
spliced = pd.DataFrame.sparse.from_spmatrix(spliced.X, columns=spliced.var_names, index=spliced.obs_names).sparse.to_dense()
unspliced = pd.DataFrame.sparse.from_spmatrix(unspliced.X,columns=unspliced.var_names, index=unspliced.obs_names).sparse.to_dense()
ambiguous = pd.DataFrame.sparse.from_spmatrix(ambiguous.X,columns=ambiguous.var_names, index=ambiguous.obs_names).sparse.to_dense()
del(af_raw)
# -
spliced.sum().sum() / (spliced.sum().sum()+unspliced.sum().sum()+ambiguous.sum().sum())
unspliced.sum().sum() / (spliced.sum().sum()+unspliced.sum().sum()+ambiguous.sum().sum())
ambiguous.sum().sum() / (spliced.sum().sum()+unspliced.sum().sum()+ambiguous.sum().sum())
# ## A discard
# +
# create AnnData using spliced and unspliced count matrix
adata = anndata.AnnData(X = spliced,
layers = dict(spliced = spliced,
unspliced = unspliced))
adata.obs = example_adata.obs
adata.obsm['X_umap'] = example_adata.obsm['X_umap']
adata.write('anndata/pancreas_usa_trimmed_A_discard.h5ad', compression='gzip')
del(adata)
# -
# ## A to S:U
# +
s_ratio = spliced/(spliced+unspliced)
s_ratio = s_ratio.fillna(0.5)
new_spliced = spliced + s_ratio * ambiguous
new_unspliced = unspliced + (1-s_ratio)* ambiguous
adata = anndata.AnnData(X = new_spliced,
layers = dict(spliced = new_spliced,
unspliced = new_unspliced))
adata.obs = example_adata.obs
adata.write('anndata/pancreas_usa_trimmed_A_S2U.h5ad', compression='gzip')
del(s_ratio, new_spliced, new_unspliced, adata)
# -
# ## A to S+A:U
# +
s_ratio = (spliced+ambiguous)/(spliced+ambiguous+unspliced)
s_ratio = s_ratio.fillna(0.5)
new_spliced = spliced + s_ratio * ambiguous
new_unspliced = unspliced + (1-s_ratio)* ambiguous
adata = anndata.AnnData(X = new_spliced,
layers = dict(spliced = new_spliced,
unspliced = new_unspliced))
adata.obs = example_adata.obs
adata.obsm['X_umap'] = example_adata.obsm['X_umap']
adata.write('anndata/pancreas_usa_trimmed_A_S+A2U.h5ad', compression='gzip')
del(s_ratio, new_spliced, new_unspliced, adata)
# -
# ## A to S:U+A
# +
s_ratio = (spliced)/(spliced+ambiguous+unspliced)
s_ratio = s_ratio.fillna(0.5)
new_spliced = spliced + s_ratio * ambiguous
new_unspliced = unspliced + (1-s_ratio)* ambiguous
adata = anndata.AnnData(X = new_spliced,
layers = dict(spliced = new_spliced,
unspliced = new_unspliced))
adata.obs = example_adata.obs
adata.obsm['X_umap'] = example_adata.obsm['X_umap']
adata.write('anndata/pancreas_usa_trimmed_A_S2U+A.h5ad', compression='gzip')
del(s_ratio, new_spliced, new_unspliced, adata)
# -
# ## A to S
# +
new_spliced = spliced + ambiguous
adata = anndata.AnnData(X = new_spliced,
layers = dict(spliced = new_spliced,
unspliced = unspliced))
adata.obs = example_adata.obs
adata.obsm['X_umap'] = example_adata.obsm['X_umap']
adata.write('anndata/pancreas_usa_trimmed_A_S.h5ad', compression='gzip')
del(new_spliced, adata)
# -
# ## A to U
# +
new_unspliced = unspliced + ambiguous
adata = anndata.AnnData(X = spliced,
layers = dict(spliced = spliced,
unspliced = new_unspliced))
adata.obs = example_adata.obs
adata.obsm['X_umap'] = example_adata.obsm['X_umap']
adata.write('anndata/pancreas_usa_trimmed_A_U.h5ad', compression='gzip')
del(new_unspliced, adata)
# -
# ## A uniform
# +
s_ratio = 0.5
new_spliced = spliced + s_ratio * ambiguous
new_unspliced = unspliced + (1-s_ratio)* ambiguous
adata = anndata.AnnData(X = new_spliced,
layers = dict(spliced = new_spliced,
unspliced = new_unspliced))
subset_adata.obs = example_adata.obs
subset_adata.obsm['X_umap'] = example_adata.obsm['X_umap']
subset_adata.write('anndata/pancreas_usa_trimmed_A_unif.h5ad', compression='gzip')
del(s_ratio, new_spliced, new_unspliced, adata, subset_adata)
# -
# # Running scVelo
# ## discard A
#
# +
adata = scv.read("anndata/pancreas_usa_trimmed_A_discard.h5ad")
# get embeddings
sc.tl.pca(adata)
sc.pp.neighbors(adata)
sc.tl.tsne(adata)
# sc.tl.umap(adata, n_components = 2)
adata.obsm['X_umap'] = example_adata.obsm['X_umap']
# housekeeping
matplotlib.use('AGG')
scv.settings.set_figure_params('scvelo')
# get the proportion of spliced and unspliced count
scv.utils.show_proportions(adata)
# filter cells and genes, then normalize expression values
scv.pp.filter_and_normalize(adata, min_shared_counts=20, n_top_genes=2000,enforce=True)
# scVelo pipeline
scv.pp.moments(adata, n_pcs=30, n_neighbors=30)
scv.tl.recover_dynamics(adata, n_jobs = 11)
scv.tl.velocity(adata, mode = 'dynamical')
scv.tl.velocity_graph(adata)
scv.pl.velocity_embedding_stream(adata, basis='umap', save="umap_pancreas_usa_A_discard.png")
scv.pl.velocity_embedding_stream(adata, basis='tsne', save="tsne_pancreas_usa_A_discard.png")
scv.tl.latent_time(adata)
scv.pl.scatter(adata, color='latent_time', color_map='gnuplot', size=80, save = "latent_time_pancreas_usa_A_discard.png")
# -
# ## A to S
# +
adata = scv.read("anndata/pancreas_usa_trimmed_A_S.h5ad")
# get embeddings
sc.tl.pca(adata)
sc.pp.neighbors(adata)
sc.tl.tsne(adata)
# sc.tl.umap(adata, n_components = 2)
adata.obsm['X_umap'] = example_adata.obsm['X_umap']
# housekeeping
matplotlib.use('AGG')
scv.settings.set_figure_params('scvelo')
# get the proportion of spliced and unspliced count
scv.utils.show_proportions(adata)
# filter cells and genes, then normalize expression values
scv.pp.filter_and_normalize(adata, min_shared_counts=20, n_top_genes=2000,enforce=True)
# scVelo pipeline
scv.pp.moments(adata, n_pcs=30, n_neighbors=30)
scv.tl.recover_dynamics(adata, n_jobs = 11)
scv.tl.velocity(adata, mode = 'dynamical')
scv.tl.velocity_graph(adata)
scv.pl.velocity_embedding_stream(adata, basis='umap', save="umap_pancreas_usa_A_S.png")
scv.pl.velocity_embedding_stream(adata, basis='tsne', save="tsne_pancreas_usa_A_S.png")
# scv.pl.velocity_embedding(adata, basis='umap', save="test.png")
scv.tl.latent_time(adata)
scv.pl.scatter(adata, color='latent_time', color_map='gnuplot', size=80, save = "latent_time_pancreas_usa_A_S.png")
# -
# ## A to U
# +
adata = scv.read("anndata/pancreas_usa_trimmed_A_U.h5ad")
# get embeddings
sc.tl.pca(adata)
sc.pp.neighbors(adata)
sc.tl.tsne(adata)
# sc.tl.umap(adata, n_components = 2)
adata.obsm['X_umap'] = example_adata.obsm['X_umap']
# housekeeping
matplotlib.use('AGG')
scv.settings.set_figure_params('scvelo')
# get the proportion of spliced and unspliced count
scv.utils.show_proportions(adata)
# filter cells and genes, then normalize expression values
scv.pp.filter_and_normalize(adata, min_shared_counts=20, n_top_genes=2000,enforce=True)
# scVelo pipeline
scv.pp.moments(adata, n_pcs=30, n_neighbors=30)
scv.tl.recover_dynamics(adata, n_jobs = 11)
scv.tl.velocity(adata, mode = 'dynamical')
scv.tl.velocity_graph(adata)
scv.pl.velocity_embedding_stream(adata, basis='umap', save="umap_pancreas_usa_A_U.png")
scv.pl.velocity_embedding_stream(adata, basis='tsne', save="tsne_pancreas_usa_A_U.png")
# scv.pl.velocity_embedding(adata, basis='umap', save="test.png")
scv.tl.latent_time(adata)
scv.pl.scatter(adata, color='latent_time', color_map='gnuplot', size=80, save = "latent_time_pancreas_usa_A_U.png")
# -
# ## A to S:U
#
# +
adata = scv.read("anndata/pancreas_usa_trimmed_A_S2U.h5ad")
# get embeddings
sc.tl.pca(adata)
sc.pp.neighbors(adata)
sc.tl.tsne(adata)
# sc.tl.umap(adata, n_components = 2)
adata.obsm['X_umap'] = example_adata.obsm['X_umap']
# housekeeping
matplotlib.use('AGG')
scv.settings.set_figure_params('scvelo')
# get the proportion of spliced and unspliced count
scv.utils.show_proportions(adata)
# filter cells and genes, then normalize expression values
scv.pp.filter_and_normalize(adata, min_shared_counts=20, n_top_genes=2000,enforce=True)
# scVelo pipeline
scv.pp.moments(adata, n_pcs=30, n_neighbors=30)
scv.tl.recover_dynamics(adata, n_jobs = 11)
scv.tl.velocity(adata, mode = 'dynamical')
scv.tl.velocity_graph(adata)
scv.pl.velocity_embedding_stream(adata, basis='umap', save="umap_pancreas_usa_A_S2U.png")
scv.pl.velocity_embedding_stream(adata, basis='tsne', save="tsne_pancreas_usa_A_S2U.png")
scv.tl.latent_time(adata)
scv.pl.scatter(adata, color='latent_time', color_map='gnuplot', size=80, save = "latent_time_pancreas_usa_A_S2U.png")
# -
# ## A to S+A:U
#
# +
adata = scv.read("anndata/pancreas_usa_trimmed_A_S+A2U.h5ad")
# get embeddings
sc.tl.pca(adata)
sc.pp.neighbors(adata)
sc.tl.tsne(adata)
# sc.tl.umap(adata, n_components = 2)
adata.obsm['X_umap'] = example_adata.obsm['X_umap']
# housekeeping
matplotlib.use('AGG')
scv.settings.set_figure_params('scvelo')
# get the proportion of spliced and unspliced count
scv.utils.show_proportions(adata)
# filter cells and genes, then normalize expression values
scv.pp.filter_and_normalize(adata, min_shared_counts=20, n_top_genes=2000,enforce=True)
# scVelo pipeline
scv.pp.moments(adata, n_pcs=30, n_neighbors=30)
scv.tl.recover_dynamics(adata, n_jobs = 11)
scv.tl.velocity(adata, mode = 'dynamical')
scv.tl.velocity_graph(adata)
scv.pl.velocity_embedding_stream(adata, basis='umap', save="umap_pancreas_usa_A_S+A2U.png")
scv.pl.velocity_embedding_stream(adata, basis='tsne', save="tsne_pancreas_usa_A_S+A2U.png")
scv.tl.latent_time(adata)
scv.pl.scatter(adata, color='latent_time', color_map='gnuplot', size=80, save = "latent_time_pancreas_usa_A_S+A2U.png")
# -
# ## A to S:U+A
#
# +
adata = scv.read("anndata/pancreas_usa_trimmed_A_S2U+A.h5ad")
# get embeddings
sc.tl.pca(adata)
sc.pp.neighbors(adata)
sc.tl.tsne(adata)
# sc.tl.umap(adata, n_components = 2)
adata.obsm['X_umap'] = example_adata.obsm['X_umap']
# housekeeping
matplotlib.use('AGG')
scv.settings.set_figure_params('scvelo')
# get the proportion of spliced and unspliced count
scv.utils.show_proportions(adata)
# filter cells and genes, then normalize expression values
scv.pp.filter_and_normalize(adata, min_shared_counts=20, n_top_genes=2000,enforce=True)
# scVelo pipeline
scv.pp.moments(adata, n_pcs=30, n_neighbors=30)
scv.tl.recover_dynamics(adata, n_jobs = 11)
scv.tl.velocity(adata, mode = 'dynamical')
scv.tl.velocity_graph(adata)
scv.pl.velocity_embedding_stream(adata, basis='umap', save="umap_pancreas_usa_A_S2U+A.png")
scv.pl.velocity_embedding_stream(adata, basis='tsne', save="tsne_pancreas_usa_A_S2U+A.png")
scv.tl.latent_time(adata)
scv.pl.scatter(adata, color='latent_time', color_map='gnuplot', size=80, save = "latent_time_pancreas_usa_A_S2U+A.png")
# -
# ## A to uniform
#
# +
adata = scv.read("anndata/pancreas_usa_trimmed_A_unif.h5ad")
# get embeddings
sc.tl.pca(adata)
sc.pp.neighbors(adata)
sc.tl.tsne(adata)
# sc.tl.umap(adata, n_components = 2)
adata.obsm['X_umap'] = example_adata.obsm['X_umap']
# housekeeping
matplotlib.use('AGG')
scv.settings.set_figure_params('scvelo')
# get the proportion of spliced and unspliced count
scv.utils.show_proportions(adata)
# filter cells and genes, then normalize expression values
scv.pp.filter_and_normalize(adata, min_shared_counts=20, n_top_genes=2000,enforce=True)
# scVelo pipeline
scv.pp.moments(adata, n_pcs=30, n_neighbors=30)
scv.tl.recover_dynamics(adata, n_jobs = 11)
scv.tl.velocity(adata, mode = 'dynamical')
scv.tl.velocity_graph(adata)
scv.pl.velocity_embedding_stream(adata, basis='umap', save="umap_pancreas_usa_A_unif.png")
scv.pl.velocity_embedding_stream(adata, basis='tsne', save="tsne_pancreas_usa_A_unif.png")
scv.tl.latent_time(adata)
scv.pl.scatter(adata, color='latent_time', color_map='gnuplot', size=80, save = "latent_time_pancreas_usa_A_unif.png")
# -
# ## A to S
# +
adata = scv.read("anndata/pancreas_usa_trimmed_A_S.h5ad")
del adata.obs
# get embeddings
sc.tl.pca(adata)
sc.pp.neighbors(adata)
sc.tl.tsne(adata)
sc.tl.umap(adata, n_components = 2)
# adata.obsm['X_umap'] = example_adata.obsm['X_umap']
# housekeeping
matplotlib.use('AGG')
scv.settings.set_figure_params('scvelo')
# get the proportion of spliced and unspliced count
scv.utils.show_proportions(adata)
# filter cells and genes, then normalize expression values
scv.pp.filter_and_normalize(adata, min_shared_counts=20, n_top_genes=2000,enforce=True)
# scVelo pipeline
scv.pp.moments(adata, n_pcs=30, n_neighbors=30)
scv.tl.recover_dynamics(adata, n_jobs = 11)
scv.tl.velocity(adata, mode = 'dynamical')
scv.tl.velocity_graph(adata)
scv.pl.velocity_embedding_stream(adata, basis='umap', save="umap_pancreas_usa_A_S_self_embedding.png")
scv.pl.velocity_embedding_stream(adata, basis='tsne', save="tsne_pancreas_usa_A_S_self_embedding.png")
# scv.pl.velocity_embedding(adata, basis='umap', save="test.png")
scv.tl.latent_time(adata)
scv.pl.scatter(adata, color='latent_time', color_map='gnuplot', size=80, save = "latent_time_pancreas_usa_A_S_self_embedding.png")
# -
# # example dataset
# +
example_adata = scv.datasets.pancreas()
# housekeeping
matplotlib.use('AGG')
scv.settings.set_figure_params('scvelo')
# get the proportion of spliced and unspliced count
scv.utils.show_proportions(example_adata)
# filter cells and genes, then normalize expression values
scv.pp.filter_and_normalize(example_adata, min_shared_counts=20, n_top_genes=2000,enforce=True)
# scVelo pipeline
scv.pp.moments(example_adata, n_pcs=30, n_neighbors=30)
scv.tl.recover_dynamics(example_adata, n_jobs = 5)
scv.tl.velocity(example_adata, mode = 'dynamical')
scv.tl.velocity_graph(example_adata)
scv.pl.velocity_embedding_stream(example_adata, basis='umap', save="umap_pancreas_scveloExample.png")
scv.tl.latent_time(example_adata)
scv.pl.scatter(example_adata, color='latent_time', color_map='gnuplot', size=80, save = "latent_time_pancreas_scveloExample.png")
| analysis_scripts/mouse_pancreas_velocity/mouse_pancreas_af_velocity_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
os.environ['CUDA_VISIBLE_DEVICES'] = ''
# -
from tqdm import tqdm
import random
# +
import malaya
model = malaya.dependency.transformer(model = 'xlnet', quantized = True)
pos = malaya.pos.transformer(model = 'xlnet', quantized = True)
# -
t = 'Cerpen itu telah saya karang.'
tokenizer = malaya.preprocessing.TOKENIZER().tokenize
sastrawi = malaya.stem.sastrawi()
graph, tagging, indexing = model.predict(' '.join(tokenizer(t)))
graph.to_graphvis()
pos.predict(' '.join(tokenizer(t)))
malaya.stack.voting_stack([pos] * 3, ' '.join(tokenizer('Buku itu dibaca oleh Ahmad.')))
malaya.stack.voting_stack([pos] * 3, ' '.join(tokenizer('Surat itu dihantar oleh abang semalam.')))
r = malaya.stack.voting_stack([pos] * 3, ' '.join(tokenizer(t)))
list(zip(*r))[1]
# +
import copy
import re
from malaya.text.regex import _expressions
def reset_t(tokens):
t = []
for i in range(len(tokens)):
t.append([tokens[i], 2])
return t
def augment_12_0(t, row):
text, tokens, tokens_lower, tagging = row
for i in range(len(tokens) - 2):
if tagging[i] == 'ADV' \
and tagging[i + 1] in ['PRON', 'NOUN'] \
and tagging[i + 2] in ['VERB', 'NOUN'] \
and tokens_lower[i] in ['telah', 'mesti']:
v = f'di{tokens[i + 2]}'
n = f'oleh {tokens[i + 1]}'
t[i][1] = 12
t[i + 1][0] = v
t[i + 1][1] = 12
t[i + 2][0] = n
t[i + 2][1] = 12
def augment_12_1(t, row):
text, tokens, tokens_lower, tagging = row
for i in range(len(tokens) - 1):
if tagging[i] == 'PRON' and tagging[i + 1] == 'VERB' and sastrawi.stem(tokens[i + 1]) == tokens[i + 1]:
v = f'men{tokens[i + 1]}'
if sastrawi.stem(v) == v:
v = f'mem{tokens[i + 1]}'
t[i][1] = 12
t[i + 1][0] = v
t[i + 1][1] = 12
def augment_12_2(t, row):
text, tokens, tokens_lower, tagging = row
for i in range(len(tokens) - 2):
if tagging[i] == 'VERB' \
and tagging[i + 1] in ['ADP'] \
and tagging[i + 2] in ['PRON', 'NOUN'] \
and tokens_lower[i + 1] in ['oleh']:
v = sastrawi.stem(tokens[i])
t[i][0] = tokens[i + 2]
t[i][1] = 12
t[i + 1][0] = v
t[i + 1][1] = 12
t[i + 2][0] = ''
t[i + 2][1] = 12
# -
results = []
for text in tqdm(['Cerpen itu telah saya karang.',
'Latihan itu mesti kau buat.',
'Kereta itu saya beli daripada Ali.',
'Surat itu dihantar oleh abang semalam.']):
tokens = tokenizer(text)
t = reset_t(tokens)
t_ = copy.deepcopy(t)
tokens_lower = tokenizer(text.lower())
tagging = malaya.stack.voting_stack([pos] * 3, ' '.join(tokens))
tagging = list(zip(*tagging))[1]
r = (t, tokens, tokens_lower, tagging)
augment_12_0(t_, r)
a = list(zip(*t_))[1]
if 12 not in a:
augment_12_1(t_, r)
a = list(zip(*t_))[1]
if 12 not in a:
augment_12_2(t_, r)
results.append((t, t_))
results
| session/tatabahasa/kesalahan-tatabahasa-12.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Who are the members of one of my _projects_?
# ### Overview
# There are a number of API calls related to projects. Here we focus on _getting the members_ of a single project.
#
# ### Prerequisites
# 1. You need to be a member (or owner) of _at least one_ project.
# 2. You need your _authentication token_ and the API needs to know about it. See <a href="Setup_API_environment.ipynb">**Setup_API_environment.ipynb**</a> for details.
# 3. You understand how to <a href="projects_listAll.ipynb" target="_blank">list</a> projects you are a member of (we will just use that call directly here).
#
# ## Imports
# We import the _Api_ class from the official sevenbridges-python bindings below.
# + deletable=true editable=true
import sevenbridges as sbg
# + [markdown] deletable=true editable=true
# ## Initialize the object
# The `Api` object needs to know your **auth\_token** and the correct path. Here we assume you are using the credentials file in your home directory. For other options see <a href="Setup_API_environment.ipynb">Setup_API_environment.ipynb</a>
# + deletable=true editable=true
# [USER INPUT] specify credentials file profile {cgc, sbg, default}
prof = 'default'
config_file = sbg.Config(profile=prof)
api = sbg.Api(config=config_file)
# + [markdown] deletable=true editable=true
# ## List some projects & get members of one of them
# We start by listing all of your projects, then get more information on the first one. A **detail**-call for projects returns the following *attributes*:
# * **description** The user specified project description
# * **id** _Unique_ identifier for the project, generated based on Project Name
# * **name** Name of project specified by the user, maybe _non-unique_
# * **href** Address<sup>1</sup> of the project.
# * **tags** List of tags
# * **created_on** Project creation time
# * **modified_on** Project modification time
# * **created_by** User that created the project
# * **root_folder** ID of the root folder for that project
# * **billing_group** ID of the billing group for the project
# * **settings** Dictionary with project settings for storage and task execution
#
# <sup>1</sup> This is the address where, by using API you can get this resource
# + deletable=true editable=true jupyter={"outputs_hidden": false}
# [USER INPUT] project index
p_index = 0
existing_projects = list(api.projects.query(limit=100).all())
project_members = existing_projects[p_index].get_members()
print('The selected project (%s) has %i members:' % \
(existing_projects[p_index].name, len(project_members)))
for member in project_members:
if member.permissions['admin']:
print('\t User %s is a project ADMIN' % (member.username))
else:
print('\t User %s is a project MEMBER' % (member.username))
# + [markdown] deletable=true editable=true
# ## Additional Information
# Detailed documentation of this particular REST architectural style request is available [here](http://docs.sevenbridges.com/docs/list-members-of-a-project)
| Recipes/SBPLAT/projects_membersOne.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import spacy
import time, nltk
nlp=spacy.load('en') # this will use nltk data folder
data="hello I am feeling good thank you"
p_data=nlp(data)
for i in p_data:
print(i,"lemma --->", i.lemma_)
time.sleep(2)
print("pos of word is : " , i.pos_)
print(type(i.pos_))
| NLP_begins.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="rcgwsDOj0ccE" colab_type="code" outputId="81b01c83-1c79-4394-d6e0-701588b32c53" executionInfo={"status": "ok", "timestamp": 1581803335172, "user_tz": 0, "elapsed": 7080, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}} colab={"base_uri": "https://localhost:8080/", "height": 283}
# !pip install eli5
# + id="_APHJPG10hqZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 172} outputId="497c5819-ad0c-482c-c45f-c51be4d333c4" executionInfo={"status": "ok", "timestamp": 1581803337671, "user_tz": 0, "elapsed": 9562, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
import pandas as pd
import numpy as np
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_absolute_error
from sklearn.model_selection import cross_val_score
import eli5
from eli5.sklearn import PermutationImportance
from ast import literal_eval
from tqdm import tqdm_notebook
# + id="0l8H7ki61Nvv" colab_type="code" outputId="98418166-6f05-4289-e25e-470ded6cd562" executionInfo={"status": "ok", "timestamp": 1581803337673, "user_tz": 0, "elapsed": 9552, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd /content/
# + id="c4p1PuC91b6Q" colab_type="code" outputId="6072a0c6-e678-46cd-939a-63ce03432366" executionInfo={"status": "ok", "timestamp": 1581803337674, "user_tz": 0, "elapsed": 9539, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd drive/
# + id="Rg4ijd_m1e3_" colab_type="code" outputId="092af505-d23d-402c-b308-c3dddb68912f" executionInfo={"status": "ok", "timestamp": 1581803337675, "user_tz": 0, "elapsed": 9526, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd My Drive
# + id="DVIfFmZO1lfw" colab_type="code" outputId="3278d9b0-d818-4e95-981f-96ad5902d03e" executionInfo={"status": "ok", "timestamp": 1581803337675, "user_tz": 0, "elapsed": 9487, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd Colab Notebooks
# + id="f9jfseKq1rfn" colab_type="code" outputId="f47062db-4c1c-435c-f5f1-49bbfb168677" executionInfo={"status": "ok", "timestamp": 1581803337676, "user_tz": 0, "elapsed": 9474, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd dw_matrix/
# + id="lNqKba8XVJPn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f4c13ca9-c973-49d5-9f2e-ad7a04e579ae" executionInfo={"status": "ok", "timestamp": 1581803340761, "user_tz": 0, "elapsed": 12545, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
# ls data
# + id="IAZFGt4z1yqX" colab_type="code" colab={}
df = pd.read_csv('data/men_shoes.csv', low_memory=False)
# + id="i-e2TKCGVuuk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 230} outputId="9af14433-7b45-4df9-a7e8-7061fb0bfa25" executionInfo={"status": "ok", "timestamp": 1581803342540, "user_tz": 0, "elapsed": 14257, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
df.columns
# + id="HiXeBhuNVqSf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="228778db-6f2e-4a31-9d46-a74ed94ba7cf" executionInfo={"status": "ok", "timestamp": 1581803342541, "user_tz": 0, "elapsed": 14243, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
mean_price = np.mean( df['prices_amountmin'])
mean_price
# + id="nr1RuL2fWAuR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b98303e4-849a-4d85-cb27-7ed736ff9979" executionInfo={"status": "ok", "timestamp": 1581803342542, "user_tz": 0, "elapsed": 14230, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
y_true =df['prices_amountmin']
y_pred = [mean_price] * y_true.shape[0]
mean_absolute_error(y_true, y_pred)
# + id="CQr7i6aXWN5n" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 230} outputId="7db91485-4990-456b-bcd6-ccc693537c57" executionInfo={"status": "ok", "timestamp": 1581803342543, "user_tz": 0, "elapsed": 14221, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
df.brand.value_counts()
# + id="JIcUA5VKWSTb" colab_type="code" colab={}
df ['brand_cat']= df ['brand'].factorize()[0]
# + id="SbE0jrO_Wici" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b2f534d5-a77f-4998-f211-6f3897bf1b10" executionInfo={"status": "ok", "timestamp": 1581803342544, "user_tz": 0, "elapsed": 14203, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
feats = ['brand_cat']
X = df[feats].values
y = df['prices_amountmin'].values
model= DecisionTreeRegressor(max_depth=5)
scores=cross_val_score(model, X, y, scoring='neg_mean_absolute_error')
np.mean(scores), np.std(scores)
# + id="FLiphbgM2BDn" colab_type="code" colab={}
def run_model(feats, model = DecisionTreeRegressor(max_depth=5)):
X= df[ feats ].values
y = df['prices_amountmin'].values
scores = cross_val_score(model, X, y, scoring='neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
# + id="yRy3Sny8WQbS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2bc8facc-1d85-412f-a3cc-398a02560963" executionInfo={"status": "ok", "timestamp": 1581803343132, "user_tz": 0, "elapsed": 14772, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
df['brand_cat'] = df['brand'].map(lambda x:str(x).lower()).factorize()[0]
run_model(['brand_cat'])
# + id="wQdbkeGzGYIo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d09681f4-4264-4793-bec0-7784ac72f2bf" executionInfo={"status": "ok", "timestamp": 1581803345949, "user_tz": 0, "elapsed": 16935, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
model = RandomForestRegressor(max_depth=5, n_estimators=100, random_state=0)
run_model(['brand_cat'], model),
# + id="VCrPAvkcTdMJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 143} outputId="cda3e6f4-3731-4c41-9306-2d1739e13018" executionInfo={"status": "ok", "timestamp": 1581803345950, "user_tz": 0, "elapsed": 16188, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
df.features.head().values
# + id="EXXcy3o4YbBg" colab_type="code" colab={}
def parse_features(x):
output_dict = {}
if str(x) == 'nan': return output_dict
features = literal_eval(x.replace('\\"', '"'))
for item in features:
key = item['key'].lower().strip()
value = item['value'][0].lower().strip()
output_dict[key] = value
return output_dict
df['features_parsed'] = df['features'].map(parse_features)
# + id="n0QIlDoieW5V" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="45a9cd74-4eae-4be7-b33b-d6f254af03f7" executionInfo={"status": "ok", "timestamp": 1581803347575, "user_tz": 0, "elapsed": 16463, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
keys = set()
df['features_parsed'].map( lambda x: keys.update(x.keys()) )
len(keys)
# + id="N5OyWfI2fLqZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 66, "referenced_widgets": ["f6093f614a5d443aa73aafc1c4c6e674", "3cccdd9158db4584802de8a78dca72a2", "c53a03c93a414278965295eaccfdf79b", "f1fa5fe9d2624bc68251812968786ea9", "ba7d9712e86e48a58f2aa076f98df6a7", "<KEY>", "77e44896ee9f48e2818a85350d07bd92", "40394800ab954b6790c85bed36b23b71"]} outputId="9731e912-2bff-4c18-8726-9f2b10ca073e" executionInfo={"status": "ok", "timestamp": 1581803351226, "user_tz": 0, "elapsed": 18047, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
def get_name_feat(key):
return 'feat_' + key
for key in tqdm_notebook(keys):
df[get_name_feat(key)] = df.features_parsed.map(lambda feats: feats[key] if key in feats else np.nan)
# + id="eR4kMMZfyyqt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 141} outputId="e4563ae9-53ed-4dd5-9ced-88973a9a69e9" executionInfo={"status": "ok", "timestamp": 1581803351227, "user_tz": 0, "elapsed": 16547, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
df.columns
# + id="anZLvbMM7eFR" colab_type="code" colab={}
keys_stat = {}
for key in keys:
keys_stat[key]= df[ False== df[get_name_feat(key)].isnull() ].shape[0] / df.shape[0] * 100
# + id="JX4rzmFC8DNf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="ab1c5432-1f01-4472-c381-eae803f8191e" executionInfo={"status": "ok", "timestamp": 1581803353633, "user_tz": 0, "elapsed": 16941, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
{k:v for k,v in keys_stat.items() if v > 35}
# + id="0NdVg0-F8H41" colab_type="code" colab={}
df['feat_brand_cat'] = df['feat_brand'].factorize()[0]
df['feat_color_cat'] = df['feat_color'].factorize()[0]
df['feat_gender_cat'] = df['feat_gender'].factorize()[0]
df['feat_manufacturer part number_cat'] = df['feat_manufacturer part number'].factorize()[0]
df['feat_material_cat'] = df['feat_material'].factorize()[0]
df['feat_sport_cat'] = df['feat_sport'].factorize()[0]
df['feat_style_cat'] = df['feat_style'].factorize()[0]
for key in keys:
df[get_name_feat(key) + '_cat'] = df[get_name_feat(key)].factorize()[0]
# + id="bn4nIFOI_9wq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="284a124b-e619-4ed0-beeb-04724add886f" executionInfo={"status": "ok", "timestamp": 1581803385012, "user_tz": 0, "elapsed": 904, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
run_model(['feat_brand_cat'])
# + id="tdicAJWwADvI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="cd97908e-3d95-4b4b-80ee-43299e60932d" executionInfo={"status": "ok", "timestamp": 1581803389764, "user_tz": 0, "elapsed": 473, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
df['brand'] = df['brand'].map(lambda x: str(x).lower())
df[ df.brand == df.feat_brand ].shape
# + id="vPcWhEyUF2qZ" colab_type="code" colab={}
feats = ['']
# + id="kG3gRPOSHP-q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7aaa71d8-4c07-424c-edaf-28d0e9f8b94f" executionInfo={"status": "ok", "timestamp": 1581803396080, "user_tz": 0, "elapsed": 3544, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
model = RandomForestRegressor(max_depth= 5, n_estimators = 100)
run_model(['brand_cat'], model)
# + id="x872m9HFL92Z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="3c1d73cb-410e-40f2-86d9-e5abec7174f6" executionInfo={"status": "ok", "timestamp": 1581803400066, "user_tz": 0, "elapsed": 448, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
feats_cat = [x for x in df.columns if 'cat' in x ]
feats_cat
# + id="lyFhtnlIC3mY" colab_type="code" colab={}
feats = ['brand_cat','feat_brand_cat','feat_gender_cat','feat_material_cat', 'feat_sport_cat']
#feats += feats_cat
#feats = list(set(feats))
model = RandomForestRegressor(max_depth= 5, n_estimators = 100)
result= run_model(feats , model)
# + id="ns779q5eD0Mr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="62b70587-f4d2-46e2-cf13-07d413a88406" executionInfo={"status": "ok", "timestamp": 1581803646576, "user_tz": 0, "elapsed": 3019, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
X= df [feats].values
y =df['prices_amountmin'].values
m = RandomForestRegressor(max_depth= 5, n_estimators = 100, random_state= 0)
m.fit(X,y)
print(result)
perm =PermutationImportance(m, random_state= 1).fit(X, y);
eli5.show_weights(perm, feature_names = feats)
# + id="F__xUSfcFAb_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 230} outputId="fb47f2e2-1545-4b35-9792-7f094ff333ec" executionInfo={"status": "ok", "timestamp": 1581803412502, "user_tz": 0, "elapsed": 665, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
df['brand'].value_counts()
# + id="2zzXuMfTG-rm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="58596ec3-8e52-4bcc-9def-f7d015795278" executionInfo={"status": "ok", "timestamp": 1581792104211, "user_tz": 0, "elapsed": 485, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
df [ df['brand'] == 'nike'].features_parsed.sample(5).values
# + id="5xIpikhKHWL1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 301} outputId="9605fef0-5682-406f-91c0-302005e341a4" executionInfo={"status": "ok", "timestamp": 1581803464797, "user_tz": 0, "elapsed": 471, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
df['weight'].unique()
# + id="cACwCLHn1WX6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 194} outputId="995f4216-a96f-4ed6-ff21-d8358951f2ca" executionInfo={"status": "ok", "timestamp": 1581804292235, "user_tz": 0, "elapsed": 4650, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12267226697134176235"}}
# !git add matrix_one/day5.ipynb
# !git commit -m "day5"
# !git push
# + id="Z_KmYonO29sd" colab_type="code" colab={}
# !git config --global user.email "<EMAIL>"
# !git config --global user.name "jaro19853"
# + id="bpYkVLdH3OfT" colab_type="code" colab={}
| matrix_one/day5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 8: Lists
x = 10
x = 20
print(x)
# # a collection is varaible that can hold many values
x = [10, 20]
print(x)
# list of numbers
numbers = [1, 6, 12]
numbers
numbers = [1, 1.16, 12.5, 0.6, 7]
numbers
students = ['Ahmed', 'Ali', 'Muhammad']
students
alist = ['Ahmed', 20, 'Data Science']
alist
# nested lists
alist = ['Ahmed', [20, 17], 'Data Science', ['python', 'data preprocessing', 'ML']]
alist
# +
# create empty list
# -
list2 = list()
list2
list2 = []
list2
alist
# # access list by indeces
alist[0]
alist[1]
alist[1][0]
# # loop over list
#
# 1. using iteration variable
# 2. using index
# using iteration variable
for element in alist:
print(element)
# using index
for i in range(len(alist)):
print(alist[i])
friends = ['Majdy', 'Khitam', 'Goerge']
friends
for friend in friends:
print('hello', friend)
for friend in friends:
print('Happy new year', friend)
friends
# # list is mutable
friends[0] = 'Majdi'
friends
# # string is imputable list
s1 = 'banana'
s1
s1[0]
s1[0] = 'B' # string is imputable list
# # string length
len(s1)
len(friends)
alist
len(alist)
# # list operation
a = [1, 2, 3]
b = [4, 5, 6]
a + b # concat
a - b
t = [9, 41, 12, 3, 74, 15]
t
t[3]
t[3:]
t[:5]
t[:]
t[3:5]
t[-1]
t[:-2]
# # list functions
print(dir(t))
t
t.append(8)
t
t.clear()
t
t.append(1)
t
t.append(7)
t
t = [9, 41, 12, 3, 74, 15, 8]
list1 = [9, 41, 12, 3, 74, 15, 8]
list2 = list1
list1
list2
list1[0] += 1
list1
list2
list3 = list1.copy()
list3
list1
list1[0] += 1
list1
list3
list3
list3.extend(0)
list3.extend([0, 9]) # list3 + [0, 9] # concat
list3
list3.index(15)
list3.index(7)
list3
t = [3, 41, 12, 3, 74, 15, 8, 12, 3]
t.count(10)
t.count(3)
t.count(12)
t
t.insert(4, 79) # insert(pos, element)
t
v = t.pop() # remove last value and return it
v
t
t.remove(3)
t
t.remove(3)
t
t.remove(3)
t
t
t.reverse()
t
t.sort()
t
t.sort(reverse=True)
t
t2 = [12, 'b', 'c', 4]
t2.sort()
t2
t2 = ['a', 'b', 'c', 'f']
t2.sort()
t2
t2.sort(reverse=True)
t2
x = 10
x
x = 10
print(x)
x = 10
x
print('hello')
# # is somehting in the list using in operator
t
74 in t
7 in t
t
# # built-in functions len, max, min, sum
max(t)
min(t)
sum(t)
sum(t)/len(t)
a = list()
a
a = []
a
b = list([1, 2, 3])
b
b = [1, 2, 3]
b
# # write a program to take numbers from the user untill he/she entered done, the file the average
while True:
line = input('please enter a number: ')
if line == 'done':
break
number = int(line)
print('done!')
total = 0
count = 0
while True:
line = input('please enter a number: ')
if line == 'done':
break
number = int(line)
total += number
count += 1
print('average =', total/count)
print('done!')
# +
num_list = []
while True:
line = input('please enter a number: ')
if line == 'done':
break
number = int(line)
num_list.append(number)
print('average =', sum(num_list)/len(num_list))
print('done!')
# -
# # Write a program to sum the prices of products and provide disount
# +
num_list = []
while True:
line = input('please enter a number: ')
if line == 'done':
break
number = int(line)
num_list.append(number)
discount = float(input('please enter discount ratio: '))
print('Your total is', sum(num_list))
print('Your total after discount is ', sum(num_list) - sum(num_list) * discount)
print('done!')
# -
| notebooks/ch8_lists.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# To enable plotting graphs in Jupyter notebook
# %matplotlib inline
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn import metrics
from sklearn.model_selection import train_test_split
# +
# Since it is a data file with no header, we will supply the column names which have been obtained from the above URL
# Create a python list of column names called "names"
colnames = ['preg', 'plas', 'pres', 'skin', 'test', 'mass', 'pedi', 'age', 'class']
#Load the file from local directory using pd.read_csv which is a special form of read_table
#while reading the data, supply the "colnames" list
pima_df = pd.read_csv("pima-indians-diabetes.csv",skiprows=2, names= colnames)
# -
pima_df.head(5)
# +
# Let us check whether any of the columns has any value other than numeric i.e. data is not corrupted such as a "?" instead of
# a number.
# we use np.isreal a numpy function which checks each column for each row and returns a bool array,
# where True if input element is real.
# applymap is pandas dataframe function that applies the np.isreal function columnwise
# Following line selects those rows which have some non-numeric value in any of the columns hence the ~ symbol
pima_df[~pima_df.applymap(np.isreal).all(1)]
# -
#Lets analysze the distribution of the various attributes
pima_df.describe().transpose()
# +
# Let us look at the target column which is 'class' to understand how the data is distributed amongst the various values
pima_df.groupby(["class"]).count()
# Most are not diabetic. The ratio is almost 1:2 in favor or class 0. The model's ability to predict class 0 will
# be better than predicting class 1.
# -
pima_df
# +
# Pairplot using s
sns.pairplot(pima_df, hue='class')
# +
#data for all the attributes are skewed, especially for the variable "test"
#The mean for test is 80(rounded) while the median is 30.5 which clearly indicates an extreme long tail on the right
# +
# Attributes which look normally distributed (plas, pres, skin, and mass).
# Some of the attributes look like they may have an exponential distribution (preg, test, pedi, age).
# Age should probably have a normal distribution, the constraints on the data collection may have skewed the distribution.
# There is no obvious relationship between age and onset of diabetes.
# There is no obvious relationship between pedi function and onset of diabetes.
# +
from scipy.stats import zscore
numeric_cols = pima_df.drop('class', axis=1)
# Copy the 'mpg' column alone into the y dataframe. This is the dependent variable
class_values = pd.DataFrame(pima_df[['class']])
numeric_cols = numeric_cols.apply(zscore)
pima_df_z = numeric_cols.join(class_values) # Recreating mpg_df by combining numerical columns with car names
pima_df_z.head()
# +
import matplotlib.pylab as plt
pima_df_z.boxplot(by = 'class', layout=(3,4), figsize=(15, 20))
#print(pima_df.boxplot('preg'))
# -
pima_df_z.hist('age')
# +
pima_df_z["log_age"] = np.log(pima_df_z['age'])
pima_df_z["log_test"] = np.log(pima_df_z["test"])
pima_df_z["log_preg"] = np.log(pima_df_z["preg"])
pima_df_z.hist('log_age')
# -
pima_df_z.hist("log_test")
pima_df_z.hist("log_preg")
plt.scatter(pima_df_z['log_test'] , pima_df_z["class"])
# #Conclusion -
#
# #1. Too many outliers on each dimensions indicated by long tails with gaps
# #2. data no normally distributed n the dimensions.
# #3. log2 transformation of exponential distributed data results near normal distribution
#
# #4. None of the given attributes are able to distinguishe the two classes. The distribution of the two classes eclipse each
# #other on all dimensions
# #5. Expect poor performance
#
| CourseContent/03-Intro.to.Python.and.Basic.Statistics/Week4/Basic_statistics_Pima.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Developing an AI application
#
# Going forward, AI algorithms will be incorporated into more and more everyday applications. For example, you might want to include an image classifier in a smart phone app. To do this, you'd use a deep learning model trained on hundreds of thousands of images as part of the overall application architecture. A large part of software development in the future will be using these types of models as common parts of applications.
#
# In this project, you'll train an image classifier to recognize different species of flowers. You can imagine using something like this in a phone app that tells you the name of the flower your camera is looking at. In practice you'd train this classifier, then export it for use in your application. We'll be using [this dataset](http://www.robots.ox.ac.uk/~vgg/data/flowers/102/index.html) of 102 flower categories, you can see a few examples below.
#
# <img src='assets/Flowers.png' width=500px>
#
# The project is broken down into multiple steps:
#
# * Load and preprocess the image dataset
# * Train the image classifier on your dataset
# * Use the trained classifier to predict image content
#
# We'll lead you through each part which you'll implement in Python.
#
# When you've completed this project, you'll have an application that can be trained on any set of labeled images. Here your network will be learning about flowers and end up as a command line application. But, what you do with your new skills depends on your imagination and effort in building a dataset. For example, imagine an app where you take a picture of a car, it tells you what the make and model is, then looks up information about it. Go build your own dataset and make something new.
#
# First up is importing the packages you'll need. It's good practice to keep all the imports at the beginning of your code. As you work through this notebook and find you need to import a package, make sure to add the import up here.
# Imports here
import torch
import json
from torchvision import datasets, transforms, models
import matplotlib.pyplot as plt
import numpy as np
from torch import nn, optim
from collections import OrderedDict
from PIL import Image
# +
# All the values used globally here.
# Device used to run/test the network.
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# Data directories.
data_dir = 'flowers'
train_dir = data_dir + '/train'
valid_dir = data_dir + '/valid'
test_dir = data_dir + '/test'
# Data transformations.
batch_size = 64
network_means = [0.485, 0.456, 0.406]
network_std_dev = [0.229, 0.224, 0.225]
center_crop_size = 224
image_resize_size = 255
# Training hyperparameters.
learning_rate = 0.0008
epochs = 10
# Model saving location.
saved_model_path = 'models/image_classifier_model.pt'
# -
# ## Load the data
#
# Here you'll use `torchvision` to load the data ([documentation](http://pytorch.org/docs/0.3.0/torchvision/index.html)). The data should be included alongside this notebook, otherwise you can [download it here](https://s3.amazonaws.com/content.udacity-data.com/nd089/flower_data.tar.gz). The dataset is split into three parts, training, validation, and testing. For the training, you'll want to apply transformations such as random scaling, cropping, and flipping. This will help the network generalize leading to better performance. You'll also need to make sure the input data is resized to 224x224 pixels as required by the pre-trained networks.
#
# The validation and testing sets are used to measure the model's performance on data it hasn't seen yet. For this you don't want any scaling or rotation transformations, but you'll need to resize then crop the images to the appropriate size.
#
# The pre-trained networks you'll use were trained on the ImageNet dataset where each color channel was normalized separately. For all three sets you'll need to normalize the means and standard deviations of the images to what the network expects. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`, calculated from the ImageNet images. These values will shift each color channel to be centered at 0 and range from -1 to 1.
#
# Moving the imshow utility function here because I am using it to verify if the data is loading correctly or not.
def imshow(image, ax=None, title=None, is_np=False):
"""Imshow for Tensor."""
if ax is None:
fig, ax = plt.subplots()
# PyTorch tensors assume the color channel is the first dimension
# but matplotlib assumes is the third dimension
if not is_np:
image = image.numpy()
image = image.transpose((1, 2, 0))
# Undo preprocessing
mean = np.array(network_means)
std = np.array(network_std_dev)
image = std * image + mean
# Image needs to be clipped between 0 and 1 or it looks like noise when displayed
image = np.clip(image, 0, 1)
ax.imshow(image)
ax.set_title(title)
return ax
# +
# TODO: Define your transforms for the training, validation, and testing sets
train_transform = transforms.Compose(
[
transforms.RandomRotation(30),
transforms.RandomResizedCrop(center_crop_size),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize(network_means, network_std_dev)
]
)
test_transform = transforms.Compose(
[
transforms.Resize(image_resize_size),
transforms.CenterCrop(center_crop_size),
transforms.ToTensor(),
transforms.Normalize(network_means, network_std_dev)
]
)
# TODO: Load the datasets with ImageFolder
train_dataset = datasets.ImageFolder(train_dir, transform=train_transform)
test_dataset = datasets.ImageFolder(test_dir, transform=test_transform)
validation_dataset = datasets.ImageFolder(valid_dir, transform=test_transform)
# TODO: Using the image datasets and the trainforms, define the dataloaders
train_loader = torch.utils.data.DataLoader(train_dataset, batch_size=batch_size, shuffle=True, drop_last=True)
test_loader = torch.utils.data.DataLoader(test_dataset, batch_size=batch_size, shuffle=True, drop_last=True)
validation_loader = torch.utils.data.DataLoader(validation_dataset, batch_size=batch_size, shuffle=True, drop_last=True)
print(f"Training dataset size: {len(train_dataset)}")
print(f"Test dataset size: {len(test_dataset)}")
print(f"Validation dataset size: {len(validation_dataset)}")
# +
# Test if the data is loaded properly.
im1, _ = next(iter(train_loader))
im2, _ = next(iter(test_loader))
im3, _ = next(iter(validation_loader))
print(f"Shape of the image: {im1.shape}")
imshow(im1[0], title='Train Image')
imshow(im2[0], title='Test Image')
imshow(im3[0], title='Validation Image')
# -
# ### Label mapping
#
# You'll also need to load in a mapping from category label to category name. You can find this in the file `cat_to_name.json`. It's a JSON object which you can read in with the [`json` module](https://docs.python.org/2/library/json.html). This will give you a dictionary mapping the integer encoded categories to the actual names of the flowers.
# +
with open('cat_to_name.json', 'r') as f:
cat_to_name = json.load(f)
# number of classes we have
num_classes = len(cat_to_name)
# -
# # Building and training the classifier
#
# Now that the data is ready, it's time to build and train the classifier. As usual, you should use one of the pretrained models from `torchvision.models` to get the image features. Build and train a new feed-forward classifier using those features.
#
# We're going to leave this part up to you. If you want to talk through it with someone, chat with your fellow students! You can also ask questions on the forums or join the instructors in office hours.
#
# Refer to [the rubric](https://review.udacity.com/#!/rubrics/1663/view) for guidance on successfully completing this section. Things you'll need to do:
#
# * Load a [pre-trained network](http://pytorch.org/docs/master/torchvision/models.html) (If you need a starting point, the VGG networks work great and are straightforward to use)
# * Define a new, untrained feed-forward network as a classifier, using ReLU activations and dropout
# * Train the classifier layers using backpropagation using the pre-trained network to get the features
# * Track the loss and accuracy on the validation set to determine the best hyperparameters
#
# We've left a cell open for you below, but use as many as you need. Our advice is to break the problem up into smaller parts you can run separately. Check that each part is doing what you expect, then move on to the next. You'll likely find that as you work through each part, you'll need to go back and modify your previous code. This is totally normal!
#
# When training make sure you're updating only the weights of the feed-forward network. You should be able to get the validation accuracy above 70% if you build everything right. Make sure to try different hyperparameters (learning rate, units in the classifier, epochs, etc) to find the best model. Save those hyperparameters to use as default values in the next part of the project.
# +
# TODO: Build and train your network
# +
model = models.densenet121(pretrained=True)
for param in model.parameters():
param.requires_grad = False
# +
# Replace the old classifier
classifier = nn.Sequential(OrderedDict([
('fc1', nn.Linear(1024, 512)),
('relu1', nn.ReLU()),
('dropout1', nn.Dropout(p=0.2)),
('fc2', nn.Linear(512, 256)),
('relu2', nn.ReLU()),
('dropout2', nn.Dropout(p=0.2)),
('fc3', nn.Linear(256, 102)),
('output', nn.LogSoftmax(dim=1))
]))
model.classifier = classifier
# +
# %%capture
# Move the model to available device. Preferably GPU otherwise fallback to CPU.
model.to(device)
# +
criterion = nn.NLLLoss()
optimizer = optim.Adam(model.classifier.parameters(), lr=learning_rate)
# +
previous_accuracy = -1
# Train the model.
for epoch in range(epochs):
train_loss = 0
for images, labels in train_loader:
images, labels = images.to(device), labels.to(device)
outputs = model.forward(images)
optimizer.zero_grad()
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
train_loss += loss.item()
else:
model.eval()
with torch.no_grad():
validation_loss = 0
current_accuracy = 0
for validation_images, validation_labels in validation_loader:
validation_images, validation_labels = validation_images.to(device), validation_labels.to(device)
validation_outputs = model.forward(validation_images)
validation_loss = criterion(validation_outputs, validation_labels)
validation_loss += validation_loss.item()
validation_ps = torch.exp(validation_outputs)
validation_top_p, validation_top_class = validation_ps.topk(1, dim=1)
validation_equal_result = validation_top_class == validation_labels.view(*validation_top_class.shape)
current_accuracy += torch.mean(validation_equal_result.type(torch.FloatTensor))
calc_current_accuracy = current_accuracy / len(validation_loader)
if previous_accuracy != -1 and calc_current_accuracy < previous_accuracy:
print(f"\n\nCurrent accuracy: {calc_current_accuracy:.3f} < Previous accuracy: {previous_accuracy:.3f}. Stopping early for the best model.")
# We are breaking out prematurely. Let's make sure we return the model to one state.
model.train()
break
else:
calc_train_loss = train_loss / len(train_loader)
calc_validation_loss = validation_loss / len(validation_loader)
print(f"Epoch {epoch + 1}/{epochs} | "
f"Train loss: {calc_train_loss:.3f} | "
f"Validation loss: {calc_validation_loss:.3f} | "
f"Validation accuracy: {calc_current_accuracy:.3f}")
previous_accuracy = calc_current_accuracy
model.train()
print(f"\n\nModel training complete with the final accuracy of: {previous_accuracy * 100:.2f}%.")
previous_accuracy = -1
# -
# ## Testing your network
#
# It's good practice to test your trained network on test data, images the network has never seen either in training or validation. This will give you a good estimate for the model's performance on completely new images. Run the test images through the network and measure the accuracy, the same way you did validation. You should be able to reach around 70% accuracy on the test set if the model has been trained well.
# +
# TODO: Do validation on the test set
test_loss = 0
test_accuracy = 0
model.eval()
with torch.no_grad():
for test_images, test_labels in test_loader:
test_images, test_labels = test_images.to(device), test_labels.to(device)
test_outputs = model.forward(test_images)
test_loss = criterion(test_outputs, test_labels)
test_loss += test_loss.item()
test_ps = torch.exp(test_outputs)
test_top_p, test_top_class = test_ps.topk(1, dim=1)
test_equal_result = test_top_class == test_labels.view(*test_top_class.shape)
test_accuracy += torch.mean(test_equal_result.type(torch.FloatTensor))
model.train()
calc_test_accuracy = test_accuracy / len(test_loader)
calc_test_loss = test_loss / len(test_loader)
print(f"Test loss: {calc_test_loss:.3f} | "
f"Test accuracy: {calc_test_accuracy:.3f}")
# -
# ## Save the checkpoint
#
# Now that your network is trained, save the model so you can load it later for making predictions. You probably want to save other things such as the mapping of classes to indices which you get from one of the image datasets: `image_datasets['train'].class_to_idx`. You can attach this to the model as an attribute which makes inference easier later on.
#
# ```model.class_to_idx = image_datasets['train'].class_to_idx```
#
# Remember that you'll want to completely rebuild the model later so you can use it for inference. Make sure to include any information you need in the checkpoint. If you want to load the model and keep training, you'll want to save the number of epochs as well as the optimizer state, `optimizer.state_dict`. You'll likely want to use this trained model in the next part of the project, so best to save it now.
# +
# TODO: Save the checkpoint
model.eval()
model.class_to_idx = train_dataset.class_to_idx
model.optimizer_state_dict = optimizer.state_dict
torch.save(model, saved_model_path)
# -
# ## Loading the checkpoint
#
# At this point it's good to write a function that can load a checkpoint and rebuild the model. That way you can come back to this project and keep working on it without having to retrain the network.
# +
# %%capture
# TODO: Write a function that loads a checkpoint and rebuilds the model
loaded_model = torch.load(saved_model_path)
loaded_model.to(device)
# -
# # Inference for classification
#
# Now you'll write a function to use a trained network for inference. That is, you'll pass an image into the network and predict the class of the flower in the image. Write a function called `predict` that takes an image and a model, then returns the top $K$ most likely classes along with the probabilities. It should look like
#
# ```python
# probs, classes = predict(image_path, model)
# print(probs)
# print(classes)
# > [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]
# > ['70', '3', '45', '62', '55']
# ```
#
# First you'll need to handle processing the input image such that it can be used in your network.
#
# ## Image Preprocessing
#
# You'll want to use `PIL` to load the image ([documentation](https://pillow.readthedocs.io/en/latest/reference/Image.html)). It's best to write a function that preprocesses the image so it can be used as input for the model. This function should process the images in the same manner used for training.
#
# First, resize the images where the shortest side is 256 pixels, keeping the aspect ratio. This can be done with the [`thumbnail`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) or [`resize`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) methods. Then you'll need to crop out the center 224x224 portion of the image.
#
# Color channels of images are typically encoded as integers 0-255, but the model expected floats 0-1. You'll need to convert the values. It's easiest with a Numpy array, which you can get from a PIL image like so `np_image = np.array(pil_image)`.
#
# As before, the network expects the images to be normalized in a specific way. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`. You'll want to subtract the means from each color channel, then divide by the standard deviation.
#
# And finally, PyTorch expects the color channel to be the first dimension but it's the third dimension in the PIL image and Numpy array. You can reorder dimensions using [`ndarray.transpose`](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.ndarray.transpose.html). The color channel needs to be first and retain the order of the other two dimensions.
def process_image(image_path):
''' Scales, crops, and normalizes a PIL image for a PyTorch model,
returns an Numpy array
'''
# TODO: Process a PIL image for use in a PyTorch model
im = Image.open(image_path)
im = im.resize((image_resize_size, image_resize_size))
im = im.crop(
(
(image_resize_size - center_crop_size) // 2,
(image_resize_size - center_crop_size) // 2,
(image_resize_size + center_crop_size) // 2,
(image_resize_size + center_crop_size) // 2
)
)
# Imshow expects values between 0 and 1
np_image = np.array(im) / 255
np_image = (np_image - network_means) / network_std_dev
# pytorch expects color channel to be the first.
np_image = np_image.transpose((2, 0, 1))
return np_image
# +
# TODO: Show the image processed above.
np_image_data = process_image('flowers/test/1/image_06743.jpg')
imshow(np_image_data, is_np=True)
# -
# ## Class Prediction
#
# Once you can get images in the correct format, it's time to write a function for making predictions with your model. A common practice is to predict the top 5 or so (usually called top-$K$) most probable classes. You'll want to calculate the class probabilities then find the $K$ largest values.
#
# To get the top $K$ largest values in a tensor use [`x.topk(k)`](http://pytorch.org/docs/master/torch.html#torch.topk). This method returns both the highest `k` probabilities and the indices of those probabilities corresponding to the classes. You need to convert from these indices to the actual class labels using `class_to_idx` which hopefully you added to the model or from an `ImageFolder` you used to load the data ([see here](#Save-the-checkpoint)). Make sure to invert the dictionary so you get a mapping from index to class as well.
#
# Again, this method should take a path to an image and a model checkpoint, then return the probabilities and classes.
#
# ```python
# probs, classes = predict(image_path, model)
# print(probs)
# print(classes)
# > [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]
# > ['70', '3', '45', '62', '55']
# ```
def predict(image_path, model, topk=5):
''' Predict the class (or classes) of an image using a trained deep learning model.
'''
with torch.no_grad():
test_image = process_image(image_path)
test_image = torch.from_numpy(test_image).float().unsqueeze_(0)
test_image = test_image.to(device)
test_output = model.forward(test_image)
test_p = torch.exp(test_output)
return test_p.topk(5, dim=1)
# ## Sanity Checking
#
# Now that you can use a trained model for predictions, check to make sure it makes sense. Even if the testing accuracy is high, it's always good to check that there aren't obvious bugs. Use `matplotlib` to plot the probabilities for the top 5 classes as a bar graph, along with the input image. It should look like this:
#
# <img src='assets/inference_example.png' width=300px>
#
# You can convert from the class integer encoding to actual flower names with the `cat_to_name.json` file (should have been loaded earlier in the notebook). To show a PyTorch tensor as an image, use the `imshow` function defined above.
# +
class_to_idx = loaded_model.class_to_idx
idx_to_class = dict((v, k) for k, v in class_to_idx.items())
# +
# TODO: Display an image along with the top 5 classes
sanity_image_path = 'flowers/test/89/image_00708.jpg'
sanity_image_class = cat_to_name[sanity_image_path.split('/')[2]]
top_prob, top_classes = predict(sanity_image_path, loaded_model)
top_prob = top_prob[0].tolist()
top_class_list = [ cat_to_name[idx_to_class[top_class]] for top_class in top_classes[0].tolist() ]
# +
imshow(process_image(sanity_image_path), is_np=True, title=sanity_image_class)
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
ax.bar(top_class_list, top_prob)
plt.show()
# -
| Image Classifier Project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import csv
from sklearn import preprocessing
import numpy as np
# %matplotlib notebook
from mpl_toolkits.mplot3d import axes3d
import matplotlib.pyplot as plt
# #%matplotlib inline
datContent = [i.strip().split() for i in open("./doughs.dat").readlines()]
y=np.array(datContent[1:])
labels = y[:,7].astype(np.float32)
y = y[:,1:7].astype(np.float32)
labels
class pca:
def __init__(self,k,scaling = False, ratio = False):
self.k = k
self.scaling = scaling
self.ratio = ratio
def EV(self,y):
if self.scaling:
scaler = preprocessing.StandardScaler().fit(y)
y=scaler.transform(y)
else:
y = y - y.mean(axis=0)
self.y = y
s = (y.shape[0]-1)*np.cov(y.T)
l, v = np.linalg.eig(s)
self.v = v[:self.k]
if self.ratio:
print(l[:self.k]/np.sum(l))
return l[:self.k],v[:self.k]
def pdata(self):
self.pdata1 = np.dot(self.y,np.array(self.v).T)
return self.pdata1
def mse(self):
self.pdata1 = np.dot(self.y,np.array(self.v).T)
e = (self.y - np.dot(self.pdata1,self.v))**2
e = e.mean()
return e
def Scatter3D(self,labels):
l = np.ones(labels.shape[0])
for i in range(labels.shape[0]):
if labels[i] >= 5:
l[i]=0
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
for i in range(self.pdata1.shape[0]):
if l[i]==1:
m='^'
c='b'
else:
m='o'
c='r'
ax.scatter(self.pdata1[i,0], self.pdata1[i,1], self.pdata1[i,2], marker=m, color=c)
ax.set_xlabel('First PC')
ax.set_ylabel('Second PC')
ax.set_zlabel('Third PC')
def Scatter2D(self,labels):
l = np.ones(labels.shape[0])
for i in range(labels.shape[0]):
if labels[i] >= 5:
l[i]=0
fig = plt.figure()
fig.tight_layout()
ax = fig.add_subplot(131)
plt.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0.5, hspace=0.4)
for i in range(self.pdata1.shape[0]):
if l[i]==1:
m='^'
c='b'
else:
m='o'
c='r'
ax.scatter(self.pdata1[i,0],self.pdata1[i,1], marker=m, color=c)
plt.xlabel('First PC')
plt.ylabel('Second PC')
ax1 = fig.add_subplot(132)
for i in range(self.pdata1.shape[0]):
if l[i]==1:
m='^'
c='b'
else:
m='o'
c='r'
ax1.scatter(self.pdata1[i,0], self.pdata1[i,2], marker=m, color=c)
plt.xlabel('First PC')
plt.ylabel('Third PC')
ax2 = fig.add_subplot(133)
for i in range(self.pdata1.shape[0]):
if l[i]==1:
m='^'
c='b'
else:
m='o'
c='r'
ax2.scatter(self.pdata1[i,1], self.pdata1[i,2],marker=m, color=c)
plt.xlabel('Second PC')
plt.ylabel('Third PC')
p = pca(3)
p.__init__(3,scaling=False,ratio=True)
p.EV(y)
p.mse()
p.Scatter3D(labels)
p.Scatter2D(labels)
| 3-c.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
import time
from get_params import get_params
from Bow_grafic import grafic
from eval_rankings import eval_rankings
import matplotlib.pyplot as plt
from eval_rankings import *
params = get_params()
# +
params['descriptor_size'] = 1024
mean=grafic(params,'train')
meant=grafic(params,'val')
llista1=[mean]
llista2=[meant]
params['descriptor_size'] = 256
mean=grafic(params,'train')
meant=grafic(params,'val')
llista1.append(mean)
llista2.append(meant)
size=[2561024]
plt.plot(llista1,size)
# +
ap_list, dict_ = eval_rankings(params)
print "Precisió mitjana:",np.mean(ap_list)
for id in dict_.keys():
if not id == 'desconegut':
# We divide by 10 because it's the number of images per class in the validation set.
print id+":", dict_[id]/10
# -
query_id='apryzkbgzo'
single_eval(params,query_id)
query_id='adpcqmgnet'
single_eval(params,query_id)
| Notebooks/Informe5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from compute_filters import *
from matplotlib import pyplot as plt
# create phantom
sl = tomopy.misc.phantom.shepp2d(size=256)[0]
plt.imshow(sl)
# +
# create data using ASTRA
N = sl.shape[0]
N_theta = 32
angles = np.linspace(0,np.pi, N_theta, False)
vol_geom = astra.create_vol_geom(N,N)
proj_geom = astra.create_proj_geom('parallel', 1.0, N, angles)
proj_id = astra.create_projector('strip', proj_geom, vol_geom)
sino_id, sino = astra.create_sino(sl, proj_id)
# add noise to sino
sino = astra.add_noise_to_sino(sino,2**10)
# pad sinogram
sino = np.pad(sino, ((0,), (128,)), 'constant')
plt.imshow(sino, aspect='auto')
# +
# reconstruct this sinogram with various software packages and the Shepp-Logan filter
recos = Reconstructions(sino, N, angles, filter='shepp-logan')
reco_strip = recos.astra_fbp()
reco_line = recos.astra_fbp(proj_type='line')
reco_linear = recos.astra_fbp(proj_type='linear')
# reco_cuda = recos.astra_fbp_cuda()
reco_iradon = recos.skimage_iradon()
reco_tomopy_gr = recos.tomopy_gridrec()
# +
# compute implementation-adapted filters for each implementation and reconstruct using these filters
filters = ComputedFilters(sino, angles, N, exp_binning=True, large_bins=4)
strip = filters.filter_fbp('astra-strip')
line = filters.filter_fbp('astra-line')
linear = filters.filter_fbp('astra-linear')
# cuda = filters.filter_fbp('astra-cuda')
iradon = filters.filter_fbp('iradon')
tomopy_gr = filters.filter_fbp('tomopy-gridrec')
tomopy_gr[0] = 0.0 # this is required because the 0th component is zeroed out in tomopy
reco_filter_strip = fbp_with_filter(sino, angles, N, 'astra-strip', strip, circle=True)
reco_filter_line = fbp_with_filter(sino, angles, N, 'astra-line', line, circle=True)
reco_filter_linear = fbp_with_filter(sino, angles, N, 'astra-linear', linear, circle=True)
# reco_filter_cuda = fbp_with_filter(sino, angles, N, 'astra-cuda', cuda, circle=True)
reco_filter_iradon = fbp_with_filter(sino, angles, N, 'iradon', iradon, circle=True)
reco_filter_tomopy_gr = gridrec_with_filter(sino, angles, N, 'tomopy-gridrec', tomopy_gr, circle=True)
# +
# compute standard deviation between sets of reconstructions and display these as heat maps
## standard reconstructions
std_reco = np.std([reco_strip.flatten(),
reco_line.flatten(),
reco_linear.flatten(),
# reco_cuda.flatten(),
reco_iradon.flatten(),
reco_tomopy_gr.flatten()], axis=0).reshape([N,N])
## implementaion-adapted reconstructions
std_reco_filter = np.std([reco_filter_strip.flatten(),
reco_filter_line.flatten(),
reco_filter_linear.flatten(),
# reco_filter_cuda.flatten(),
reco_filter_iradon.flatten(),
reco_filter_tomopy_gr.flatten()], axis=0).reshape([N,N])
_min = 0.0
_max = np.max(std_reco)
plt.figure(figsize=(15,5))
plt.subplot(121)
plt.imshow(std_reco, vmin=_min, vmax=_max)
plt.axis('off')
plt.colorbar()
plt.title('Standard filter recons')
plt.subplot(122)
plt.imshow(std_reco_filter, vmin=_min, vmax=_max)
plt.axis('off')
plt.colorbar()
plt.title('Impl-adapted filter recons')
plt.tight_layout()
# -
| example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# Lambda School Data Science
#
# *Unit 2, Sprint 3, Module 1*
#
# ---
#
# Lambda School Data Science, Unit 2: Predictive Modeling
#
# # Define ML problems
#
# - Choose a **target** to predict, and check its **distribution**
# - Choose what data to hold out for your **test set**
# - Choose an appropriate **evaluation metric**
# - Avoid **leakage** of information from test to train or from target to features
# ### Setup
#
# Run the code cell below. You can work locally (follow the [local setup instructions](https://lambdaschool.github.io/ds/unit2/local/)) or on Colab.
#
# +
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# -
# ## Today we're focused on the "first arrow" ...
#
# #### <NAME>, [Becoming a Data Scientist, PyData DC 2016 Talk](https://www.becomingadatascientist.com/2016/10/11/pydata-dc-2016-talk/)
#
# 
#
# ## For these kinds of questions ...
#
# #### <NAME>, [Five Questions Data Science Answers](https://brohrer.github.io/five_questions_data_science_answers.html)
#
# > - Is this A or B? (or C or ...)
# > - How Much / How Many?
#
# ## Doing steps 1—3 in this workflow ...
#
# #### <NAME>, [Deep Learning with Python](https://github.com/fchollet/deep-learning-with-python-notebooks/blob/master/README.md), Chapter 4: Fundamentals of machine learning, "A universal workflow of machine learning"
#
# > **1. Define the problem at hand and the data on which you’ll train.** Collect this data, or annotate it with labels if need be.
#
# > **2. Choose how you’ll measure success on your problem.** Which metrics will you monitor on your validation data?
#
# > **3. Determine your evaluation protocol:** hold-out validation? K-fold validation? Which portion of the data should you use for validation?
#
# > **4. Develop a first model that does better than a basic baseline:** a model with statistical power.
#
# > **5. Develop a model that overfits.** The universal tension in machine learning is between optimization and generalization; the ideal model is one that stands right at the border between underfitting and overfitting; between undercapacity and overcapacity. To figure out where this border lies, first you must cross it.
#
# > **6. Regularize your model and tune its hyperparameters, based on performance on the validation data.** Repeatedly modify your model, train it, evaluate on your validation data (not the test data, at this point), modify it again, and repeat, until the model is as good as it can get.
#
# > **Iterate on feature engineering: add new features, or remove features that don’t seem to be informative.** Once you’ve developed a satisfactory model configuration, you can train your final production model on all the available data (training and validation) and evaluate it one last time on the test set.
#
# ## It's not easy!
#
# This opinionated blog post explains:
#
# #### [Data Science Is Not Taught At Universities - And Here Is Why](https://www.linkedin.com/pulse/data-science-taught-universities-here-why-maciej-wasiak/)
#
# > The tables they use in machine learning research already have the target information clearly defined. Here comes the famous IRIS dataset, then the Wisconsin Breast Cancer, there is even Credit Risk or Telco Churn data and they all have the **Target** column there ...
#
# > The problem is that in real life the **Target** flag is NEVER there.
#
# > For churn modelling you may have many churn types on the system and need to distil the few that need modelling. And hey - when a subscriber moves from Postpaid contract to Prepaid – is this a churn or not? (‘Yes’ – says the Postpaid Base Manager, ‘No’ says the CEO ). You have to make the call ...
#
# > Your source will be a database with tens or hundreds of **tables**, millions of records, usually after 3 painful migrations with gaps in history, columns without descriptions ...
#
# > Flooded by **leaks from the future**, ...a dozen of other traps ... And you need to disarm all of them, because even one left behind may result in a completely useless model.
#
# > These are the skills employers are looking for.
#
# # Choose a target to predict, and check its distribution
#
# ## Regression, Binary Classification, or Multi-Class Classification? It's up to you!
#
#
# #### You can convert problems from regression to classification
#
# 1. UCI, [Adult Census Income dataset](https://archive.ics.uci.edu/ml/datasets/adult)
#
# 2. DS5 student <NAME>, [Bitcoin Price Prediction app](https://dry-shore-97069.herokuapp.com/about):
#
# > We also cared a lot more about the direction of returns instead of magnitude of returns. A trade placed based on the prediction that the price to go up tomorrow will be fine if the magnitude is off but will be unprofitable if the direction is wrong. ... Yesterday's return is unsurprising a great predictor for today's return, but has a poor directional accuracy.
#
#
# #### You can convert problems from classification to regression
#
# <NAME>, [What questions can machine learning answer](https://brohrer.github.io/five_questions_data_science_answers.html)
#
# > Sometimes questions that look like multi-value classification questions are actually better suited to regression. For instance, “Which news story is the most interesting to this reader?” appears to ask for a category—a single item from the list of news stories. However, you can reformulate it to “How interesting is each story on this list to this reader?” and give each article a numerical score. Then it is a simple thing to identify the highest-scoring article. Questions of this type often occur as rankings or comparisons.
#
# > “Which van in my fleet needs servicing the most?” can be rephrased as “How badly does each van in my fleet need servicing?”
# “Which 5% of my customers will leave my business for a competitor in the next year?” can be rephrased as “How likely is each of my customers to leave my business for a competitor in the next year?”
#
# > Binary classification problems can also be reformulated as regression. (In fact, under the hood some algorithms reformulate every binary classification as regression.) This is especially helpful when an example can belong part A and part B, or have a chance of going either way. When an answer can be partly yes and no, probably on but possibly off, then regression can reflect that. Questions of this type often begin “How likely…” or “What fraction…”
#
# > How likely is this user to click on my ad? What fraction of pulls on this slot machine result in payout? How likely is this employee to be an insider security threat? What fraction of today’s flights will depart on time?
#
# We'll see examples of this, using predicted probabilities instead of discrete predictions, with Tanzania Waterpumps and Lending Club data.
#
#
# #### You can convert multi-class classification to binary classification
#
# By omitting or combining some classes. We'll also see examples of this, with Tanzania Waterpumps and Lending Club data.
#
# ## Classification problems with imbalanced classes
#
# [Learning from Imbalanced Classes](https://www.svds.com/tbt-learning-imbalanced-classes/) gives "a rough outline of useful approaches" :
#
# - Do nothing. Sometimes you get lucky and nothing needs to be done. You can train on the so-called natural (or stratified) distribution and sometimes it works without need for modification.
# - Balance the training set in some way:
# - Oversample the minority class.
# - Undersample the majority class.
# - Synthesize new minority classes.
# - Throw away minority examples and switch to an anomaly detection framework.
# - At the algorithm level, or after it:
# - Adjust the class weight (misclassification costs).
# - Adjust the decision threshold.
# - Modify an existing algorithm to be more sensitive to rare classes.
# - Construct an entirely new algorithm to perform well on imbalanced data.
#
# #### Technical implementations:
#
# - "Adjust the class weight (misclassification costs)" — many scikit-learn classifiers have a `class_balance` parameter
# - "Adjust the decision threshold" — we did this last week. You can lean more about it in a great blog post, [Visualizing Machine Learning Thresholds to Make Better Business Decisions](https://blog.insightdatascience.com/visualizing-machine-learning-thresholds-to-make-better-business-decisions-4ab07f823415).
# - "Oversample the minority class, undersample the majority class, or synthesize new minority classes" — the [imbalance-learn](https://github.com/scikit-learn-contrib/imbalanced-learn) library can be used for this.
#
# ## Regression problems with right-skewed targets
#
# #### <NAME> & <NAME>, [The Mechanics of Machine Learning, Chapter 5.5](https://mlbook.explained.ai/prep.html#logtarget)
#
# > Transforming the target variable (using the mathematical log function) into a tighter, more uniform space makes life easier for any model.
#
# > The only problem is that, while easy to execute, understanding why taking the log of the target variable works and how it affects the training/testing process is intellectually challenging. You can skip this section for now, if you like, but just remember that this technique exists and check back here if needed in the future.
#
# > Optimally, the distribution of prices would be a narrow “bell curve” distribution without a tail. This would make predictions based upon average prices more accurate. We need a mathematical operation that transforms the widely-distributed target prices into a new space. The “price in dollars space” has a long right tail because of outliers and we want to squeeze that space into a new space that is normally distributed (“bell curved”). More specifically, we need to shrink large values a lot and smaller values a little. That magic operation is called the logarithm or log for short.
#
# > To make actual predictions, we have to take the exp of model predictions to get prices in dollars instead of log dollars.
#
# # Choose what data to hold out for your test set
#
# #### <NAME>, [How (and why) to create a good validation set](https://www.fast.ai/2017/11/13/validation-sets/)
# # Choose an appropriate evaluation metric
#
# https://scikit-learn.org/stable/modules/model_evaluation.html#common-cases-predefined-values
#
# Do not only use accuracy score when classes are imbalanced.
#
# Consider custom evaluation metrics:
#
# - [Attacking discrimination with smarter machine learning](https://research.google.com/bigpicture/attacking-discrimination-in-ml/), by Google Research, with interactive visualizations. _"A threshold classifier essentially makes a yes/no decision, putting things in one category or another. We look at how these classifiers work, ways they can potentially be unfair, and how you might turn an unfair classifier into a fairer one. As an illustrative example, we focus on loan granting scenarios where a bank may grant or deny a loan based on a single, automatically computed number such as a credit score."_
# - [How Shopify Capital Uses Quantile Regression To Help Merchants Succeed](https://engineering.shopify.com/blogs/engineering/how-shopify-uses-machine-learning-to-help-our-merchants-grow-their-business)
# - [Machine Learning Meets Economics](http://blog.mldb.ai/blog/posts/2016/01/ml-meets-economics/)
# - [Maximizing Scarce Maintenance Resources with Data: Applying predictive modeling, precision at k, and clustering to optimize impact](https://towardsdatascience.com/maximizing-scarce-maintenance-resources-with-data-8f3491133050), by Lambda DS3 student <NAME>. His blog post extends the Tanzania Waterpumps scenario, far beyond what's in the lecture notebook.
# - [Notebook about how to calculate expected value from a confusion matrix by treating it as a cost-benefit matrix](https://github.com/podopie/DAT18NYC/blob/master/classes/13-expected_value_cost_benefit_analysis.ipynb)
#
# # Avoid leakage of information from test to train or from target to features
# [<NAME> recommends,](https://www.quora.com/What-are-some-best-practices-for-training-machine-learning-models/answer/Xavier-Amatriain)
#
# "Make sure your training features do not contain data from the “future” (aka time traveling). While this might be easy and obvious in some cases, it can get tricky. ... If your test metric becomes really good all of the sudden, ask yourself what you might be doing wrong. Chances are you are time travelling or overfitting in some way."
#
# # 1. Bank Marketing 🏦
#
# http://archive.ics.uci.edu/ml/datasets/Bank+Marketing
#
# ## Choose a target to predict, and check its distribution
#
# +
BANK_PATH = DATA_PATH + 'bank-marketing/bank-additional-full.csv'
BANK_PATH
import pandas as pd
df = pd.read_csv(BANK_PATH, sep=';')
# -
df.shape
df.head()
# +
y = df['y'] == 'yes'
X = df.drop(columns='y')
# Check its distribution
y.describe()
# -
y.value_counts()
y.value_counts(normalize=True)
# ## Choose what data to hold out for your test set
#
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=42)
# -
y_train.value_counts(normalize=True)
y_test.value_counts(normalize=True)
# ## Choose an appropriate evaluation metric
#
# Majority Class baseline
y_pred = [False] * len(y_test)
from sklearn.metrics import accuracy_score
accuracy_score(y_test, y_pred)
from sklearn.metrics import recall_score
recall_score(y_test, y_pred)
from sklearn.metrics import precision_score
precision_score(y_test, y_pred)
from sklearn.metrics import roc_auc_score
roc_auc_score(y_test, y_pred)
# ## Avoid leakage of information from test to train or from target to features
#
#
df['nonzero_duration'] = df['duration'] > 0
df['nonzero_duration'].value_counts()
# +
import category_encoders as ce
import pandas as pd
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline
from sklearn.tree import DecisionTreeClassifier
pipe = make_pipeline(
ce.OneHotEncoder(use_cat_names=True),
DecisionTreeClassifier(max_depth=2)
)
scores = cross_val_score(pipe, X_train, y_train, cv=5, scoring='roc_auc', n_jobs=-1)
print('Validation ROC AUC:', scores)
# +
import graphviz
from sklearn.tree import export_graphviz
pipe.fit(X_train, y_train)
tree = pipe.named_steps['decisiontreeclassifier']
encoder = pipe.named_steps['onehotencoder']
feature_names = encoder.transform(X_train).columns
feature_names[tree.feature_importances_ > 0]
dot_data = export_graphviz(tree, out_file=None,
feature_names=feature_names, class_names=['False', 'True'],
filled=True, impurity=False, proportion=True)
graphviz.Source(dot_data)
# -
# ### When the `duration` feature is dropped, then the ROC AUC score drops from ~0.85 to ~0.75
X_train = X_train.drop(columns='duration')
scores = cross_val_score(pipe, X_train, y_train, cv=5, scoring='roc_auc')
print('Validation ROC AUC:', scores)
# ### ROC AUC
#
# [Wikipedia explains,](https://en.wikipedia.org/wiki/Receiver_operating_characteristic) "A receiver operating characteristic curve, or ROC curve, is a graphical plot that illustrates the diagnostic ability of a binary classifier system as its discrimination threshold is varied. The ROC curve is created by plotting the true positive rate (TPR) against the false positive rate (FPR) at various threshold settings."
#
# ROC AUC is the area under the ROC curve. [It can be interpreted](https://stats.stackexchange.com/questions/132777/what-does-auc-stand-for-and-what-is-it) as "the expectation that a uniformly drawn random positive is ranked before a uniformly drawn random negative."
#
# ROC AUC measures how well a classifier ranks predicted probabilities. It ranges from 0 to 1. A naive majority class baseline will have an ROC AUC score of 0.5.
#
# +
import matplotlib.pyplot as plt
from sklearn.metrics import roc_auc_score, roc_curve
from sklearn.model_selection import cross_val_predict
y_pred_proba = cross_val_predict(pipe, X_train, y_train, cv=5, n_jobs=-1,
method='predict_proba')[:, 1]
fpr, tpr, thresholds = roc_curve(y_train, y_pred_proba)
plt.plot(fpr, tpr)
plt.title('ROC curve')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
print('Area under the Receiver Operating Characteristic curve:',
roc_auc_score(y_train, y_pred_proba))
# -
pd.DataFrame({'False Positive Rate': fpr,
'True Positive Rate': tpr,
'Threshold': thresholds})
# # 2. Caterpillar tube pricing problem 🚜
#
# #### [Description](https://www.kaggle.com/c/caterpillar-tube-pricing/overview/description)
#
# > Like snowflakes, it's difficult to find two tubes in Caterpillar's diverse catalogue of machinery that are exactly alike. Tubes can vary across a number of dimensions, including base materials, number of bends, bend radius, bolt patterns, and end types.
#
# > Currently, Caterpillar relies on a variety of suppliers to manufacture these tube assemblies, each having their own unique pricing model. This competition provides detailed tube, component, and annual volume datasets, and challenges you to predict the price a supplier will quote for a given tube assembly.
#
# #### [Data Description](https://www.kaggle.com/c/caterpillar-tube-pricing/data)
#
# > The dataset is comprised of a large number of relational tables that describe the physical properties of tube assemblies. You are challenged to combine the characteristics of each tube assembly with supplier pricing dynamics in order to forecast a quote price for each tube. The quote price is labeled as cost in the data.
#
#
#
# ## Choose a target to predict, and check its distribution
#
df = pd.read_csv('https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/caterpillar-tube-pricing/train_set.csv')
df.head()
df['cost'].describe()
#VERY SKEWED
import seaborn as sns
sns.distplot(df['cost']);
# https://docs.scipy.org/doc/numpy/reference/routines.math.html#exponents-and-logarithms
import numpy as np
df['cost_transformed'] = np.log1p(df['cost'])
#np.expm1 to convert back
df['cost_transformed'].describe()
sns.distplot(df['cost_transformed']);
# ## Choose what data to hold out for your test set
#
# ## Choose an appropriate evaluation metric
#
# ## Avoid leakage of information from test to train or from target to features
#
#
# # 3. Lending Club 🏦
#
# ### Background
#
# [According to Wikipedia,](https://en.wikipedia.org/wiki/Lending_Club)
#
# > Lending Club is the world's largest peer-to-peer lending platform. Lending Club enables borrowers to create unsecured personal loans between \$1,000 and \$40,000. The standard loan period is three years. Investors can search and browse the loan listings on Lending Club website and select loans that they want to invest in based on the information supplied about the borrower, amount of loan, loan grade, and loan purpose. Investors make money from interest. Lending Club makes money by charging borrowers an origination fee and investors a service fee.
#
# [Lending Club says,](https://www.lendingclub.com/) "Our mission is to transform the banking system to make credit more affordable and investing more rewarding." You can view their [loan statistics and visualizations](https://www.lendingclub.com/info/demand-and-credit-profile.action).
#
# Lending Club's [Investor Education Center](https://www.lendingclub.com/investing/investor-education) can help you grow your domain expertise. The article about [Benefits of diversification](https://www.lendingclub.com/investing/investor-education/benefits-of-diversification) explains,
#
# > With the investment minimum of \$1,000, you can get up to 40 Notes at \$25 each.
#
# 
#
# ### Data sources
# - [Current loans](https://www.lendingclub.com/browse/browse.action)
# - [Data Dictionary & Historical loans](https://www.lendingclub.com/info/download-data.action) (17 zip files, 450 MB total)
#
# ### What questions could we ask with this data?
#
#
#
# ## Choose a target to predict, and check its distribution
#
# ## Choose what data to hold out for your test set
#
# ## Choose an appropriate evaluation metric
#
# ## Avoid leakage of information from test to train or from target to features
#
#
#
| module1/My Notes lesson_applied_modeling_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Describing Data by Batch
# +
import os
import pathlib
import numpy as np
import pandas as pd
import plotnine as gg
from pycytominer.cyto_utils import infer_cp_features
from scripts.processing_utils import load_data
# +
def get_count_per_batch(df, batch_name):
result = (
df
.Metadata_Plate
.value_counts()
.reset_index()
.rename({
"index": "Metadata_Plate",
"Metadata_Plate": "profile_count"
}, axis="columns")
.assign(batch=batch_name)
)
return result
def count_treatments_per_plate(df, batch_name):
group_cols = ["Metadata_clone_number", "Metadata_treatment", "Metadata_Plate"]
result = (
df
.groupby(group_cols)
["Metadata_Well"]
.count()
.reset_index()
.rename({
"Metadata_Well": "profile_count",
group_cols[0]: "Metadata_clone"
}, axis="columns")
.assign(batch=batch_name)
)
return result
def process_counts(batch_name, profile_dir="profiles"):
df = load_data(
batch=batch_name,
plates="all",
profile_dir=profile_dir,
suffix="normalized_feature_selected.csv.gz",
combine_dfs=True,
harmonize_cols=True,
add_cell_count=False,
)
batch_count = get_count_per_batch(df, batch_name)
treatment_count = count_treatments_per_plate(df, batch_name)
return df, batch_count, treatment_count
# +
profile_dir = pathlib.Path("../0.generate-profiles/profiles")
batches = sorted([x for x in os.listdir(profile_dir) if x != ".DS_Store"])
batches
# -
profile_dir
batch_data = {}
profile_count_list = list()
for batch in batches:
print("Now processing... {}".format(batch))
df, batch_count, treatment_count = process_counts(batch, profile_dir=profile_dir)
batch_data[batch] = {
"dataframe": df,
"metafeatures": infer_cp_features(df, metadata=True),
"batch_count": batch_count,
"treatment_count": treatment_count
}
profile_count_list.append(
treatment_count.loc[:, ["Metadata_clone", "Metadata_treatment", "profile_count"]]
)
# +
sample_count_df = (
pd.DataFrame(
pd.concat(profile_count_list, axis="rows")
.fillna("DMSO")
.reset_index(drop=True)
.groupby(["Metadata_clone", "Metadata_treatment"])
["profile_count"]
.sum()
)
.sort_values("profile_count", ascending=False)
.reset_index()
)
sample_treatment_count_df = (
sample_count_df
.pivot_table(
values="profile_count",
index="Metadata_clone",
columns="Metadata_treatment",
aggfunc=lambda x: x.sum()
)
.fillna(0)
.astype(int)
)
sample_treatment_count_df.to_csv(
pathlib.Path("results/sample_summary_profile_counts.tsv"), sep="\t", index=True
)
sample_treatment_count_df
# +
plot_ready_df = (
sample_treatment_count_df
.reset_index()
.melt(
id_vars="Metadata_clone",
value_vars=sample_count_df.Metadata_treatment.unique(),
value_name="profile_count"
)
)
clone_order = (
plot_ready_df
.groupby("Metadata_clone")
.sum()
.reset_index()
.sort_values(by="profile_count")
.Metadata_clone
)
plot_ready_df.Metadata_clone = pd.Categorical(
plot_ready_df.Metadata_clone,
categories=clone_order
)
plot_ready_df.head()
# +
total_count = plot_ready_df.profile_count.sum()
total_label = "Total Profile Count: {}".format(total_count)
treatment_count_gg = (
gg.ggplot(plot_ready_df, gg.aes(y="profile_count", x="Metadata_clone")) +
gg.geom_bar(gg.aes(fill="Metadata_treatment"), position="stack", stat="identity") +
gg.coord_flip() +
gg.theme_bw() +
gg.theme(axis_text_y=gg.element_text(size=5)) +
gg.ylab("Profile Count") +
gg.xlab("Clone") +
gg.ggtitle(total_label)
)
output_figure = pathlib.Path("figures", "treatment_count.png")
treatment_count_gg.save(output_figure, height=4, width=5.5, dpi=400, verbose=False)
treatment_count_gg
# -
# How many unique clones
len(sample_treatment_count_df.index.unique())
# +
all_profile_counts = []
for key, value in batch_data.items():
all_profile_counts.append(batch_data[key]["batch_count"])
profile_counts_df = pd.concat(all_profile_counts, axis="rows").reset_index(drop=True)
profile_counts_df
# +
all_treatment_counts = []
for key, value in batch_data.items():
all_treatment_counts.append(batch_data[key]["treatment_count"])
treatment_counts_df = pd.concat(all_treatment_counts, axis="rows", sort=True).reset_index(drop=True)
treatment_counts_df.head()
# +
clone_counts_df = (
treatment_counts_df
.groupby(["Metadata_clone", "Metadata_treatment"])
["profile_count"]
.sum()
.reset_index()
.sort_values(by=["Metadata_clone", "Metadata_treatment"])
)
output_file = pathlib.Path("tables/clone_counts_bortezomib.csv")
clone_counts_df.to_csv(output_file, sep=',', index=False)
clone_counts_df
# -
# ## Visualize Counts
# +
total_count = profile_counts_df.profile_count.sum()
total_label = "Total Profile Count: {}".format(total_count)
profile_counts_df.Metadata_Plate = profile_counts_df.Metadata_Plate.astype(str)
batch_count_gg = (
gg.ggplot(profile_counts_df, gg.aes(y="profile_count", x="batch")) +
gg.geom_bar(gg.aes(fill="Metadata_Plate"), stat="identity") +
gg.coord_flip() +
gg.theme_bw() +
gg.ylab("Profile Count") +
gg.xlab("Batch") +
gg.ggtitle(total_label)
)
output_figure = pathlib.Path("figures/batch_count.png")
batch_count_gg.save(output_figure, height=4, width=6.5, dpi=400, verbose=False)
batch_count_gg
# -
# ## Output Metadata Counts for Each Batch
#
# For quick description
suspect_batches = [
"2019_06_25_Batch3", # Too confluent, not even DMSO control
"2019_11_11_Batch4", # Too confluent
"2019_11_19_Batch5", # Too confluent
]
# +
non_suspect_counts = treatment_counts_df.loc[~treatment_counts_df.batch.isin(suspect_batches), :]
treatment_counts_df.Metadata_clone = pd.Categorical(
treatment_counts_df.Metadata_clone,
categories=clone_order
)
total_count = non_suspect_counts.profile_count.sum()
total_label = "Total Usable Profile Count: {}".format(total_count)
treatment_count_by_batch_gg = (
gg.ggplot(treatment_counts_df, gg.aes(y="profile_count", x="Metadata_clone")) +
gg.geom_bar(gg.aes(fill="Metadata_treatment"), position="stack", stat="identity") +
gg.coord_flip() +
gg.facet_wrap("~batch") +
gg.theme_bw() +
gg.theme(
axis_text_y=gg.element_text(size=3.5),
axis_text_x=gg.element_text(size=6),
strip_text=gg.element_text(size=6, color="black"),
strip_background=gg.element_rect(colour="black", fill="#fdfff4")
) +
gg.ylab("Profile Count") +
gg.xlab("Clones") +
gg.ggtitle(total_label)
)
output_figure = pathlib.Path("figures/treatment_count_by_batch.png")
treatment_count_by_batch_gg.save(output_figure, height=8, width=5.5, dpi=400, verbose=False)
treatment_count_by_batch_gg
# -
batch1_40x_df = treatment_counts_df.query("batch == '2019_02_15_Batch1_40X'").dropna(axis="columns")
batch1_40x_df
batch1_20x_df = treatment_counts_df.query("batch == '2019_02_15_Batch1_20X'").dropna(axis="columns")
batch1_20x_df
batch2_df = treatment_counts_df.query("batch == '2019_03_20_Batch2'").dropna(axis="columns")
batch2_df
batch3_df = treatment_counts_df.query("batch == '2019_06_25_Batch3'").dropna(axis="columns")
batch3_df
batch4_df = treatment_counts_df.query("batch == '2019_11_11_Batch4'").dropna(axis="columns")
batch4_df
batch5_df = treatment_counts_df.query("batch == '2019_11_19_Batch5'").dropna(axis="columns")
batch5_df
batch6_df = treatment_counts_df.query("batch == '2019_11_20_Batch6'").dropna(axis="columns")
batch6_df
batch7_df = treatment_counts_df.query("batch == '2019_11_22_Batch7'").dropna(axis="columns")
batch7_df
batch8_df = treatment_counts_df.query("batch == '2020_07_02_Batch8'").dropna(axis="columns")
batch8_df
batch9_df = treatment_counts_df.query("batch == '2020_08_24_Batch9'").dropna(axis="columns")
batch9_df
batch10_df = treatment_counts_df.query("batch == '2020_09_08_Batch10'").dropna(axis="columns")
batch10_df
batch11_df = treatment_counts_df.query("batch == '2021_02_08_Batch11'").dropna(axis="columns")
batch11_df
batch12_df = treatment_counts_df.query("batch == '2021_03_03_Batch12'").dropna(axis="columns")
batch12_df
batch13_df = treatment_counts_df.query("batch == '2021_03_03_Batch13'").dropna(axis="columns")
batch13_df
batch14_df = treatment_counts_df.query("batch == '2021_03_03_Batch14'").dropna(axis="columns")
batch14_df
batch15_df = treatment_counts_df.query("batch == '2021_03_03_Batch15'").dropna(axis="columns")
batch15_df
batch16_df = treatment_counts_df.query("batch == '2021_03_05_Batch16'").dropna(axis="columns")
batch16_df
batch17_df = treatment_counts_df.query("batch == '2021_03_05_Batch17'").dropna(axis="columns")
batch17_df
batch18_df = treatment_counts_df.query("batch == '2021_03_12_Batch18'").dropna(axis="columns")
batch18_df
batch19_df = treatment_counts_df.query("batch == '2021_03_12_Batch19'").dropna(axis="columns")
batch19_df
batch20_df = treatment_counts_df.query("batch == '2021_06_25_Batch20'").dropna(axis="columns")
batch20_df
batch21_df = treatment_counts_df.query("batch == '2021_06_25_Batch21'").dropna(axis="columns")
batch21_df
batch22_df = treatment_counts_df.query("batch == '2021_07_21_Batch22'").dropna(axis="columns")
batch22_df
batch23_df = treatment_counts_df.query("batch == '2021_07_21_Batch23'").dropna(axis="columns")
batch23_df
batch24_df = treatment_counts_df.query("batch == '2021_08_02_Batch24'").dropna(axis="columns")
batch24_df
batch25_df = treatment_counts_df.query("batch == '2021_08_02_Batch25'").dropna(axis="columns")
batch25_df
batch26_df = treatment_counts_df.query("batch == '2021_08_03_Batch26'").dropna(axis="columns")
batch26_df
batch27_df = treatment_counts_df.query("batch == '2021_08_03_Batch27'").dropna(axis="columns")
batch27_df
| 2.describe-data/0.describe-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import scipy.misc
import scipy.io
import numpy as np
import tensorflow as tf
import warnings
from tensorflow.python.framework import ops
ops.reset_default_graph()
warnings.filterwarnings("ignore")
sess = tf.Session()
original_image_file = 'temp/book_cover.jpg'
style_image_file = 'temp/starry_night.jpg'
vgg_path = 'temp/imagenet-vgg-verydeep-19.mat'
original_image_weight = 5.0
style_image_weight = 500.0
regularization_weight = 100
learning_rate = 0.001
generations = 5000
output_generations = 250
beta1 = 0.9
beta2 = 0.999
original_image = scipy.misc.imread(original_image_file)
style_image = scipy.misc.imread(style_image_file)
target_shape = original_image.shape
style_image = scipy.misc.imresize(style_image, target_shape[1] / style_image.shape[1])
vgg_layers = ['conv1_1', 'relu1_1',
'conv1_2', 'relu1_2', 'pool1',
'conv2_1', 'relu2_1',
'conv2_2', 'relu2_2', 'pool2',
'conv3_1', 'relu3_1',
'conv3_2', 'relu3_2',
'conv3_3', 'relu3_3',
'conv3_4', 'relu3_4', 'pool3',
'conv4_1', 'relu4_1',
'conv4_2', 'relu4_2',
'conv4_3', 'relu4_3',
'conv4_4', 'relu4_4', 'pool4',
'conv5_1', 'relu5_1',
'conv5_2', 'relu5_2',
'conv5_3', 'relu5_3',
'conv5_4', 'relu5_4']
def extract_net_info(path_to_params):
vgg_data = scipy.io.loadmat(path_to_params)
normalization_matrix = vgg_data['normalization'][0][0][0]
mat_mean = np.mean(normalization_matrix, axis=(0,1))
network_weights = vgg_data['layers'][0]
return(mat_mean, network_weights)
def vgg_network(network_weights, init_image):
network = {}
image = init_image
for i, layer in enumerate(vgg_layers):
if layer[0] == 'c':
weights, bias = network_weights[i][0][0][0][0]
weights = np.transpose(weights, (1, 0, 2, 3))
bias = bias.reshape(-1)
conv_layer = tf.nn.conv2d(image, tf.constant(weights), (1, 1, 1, 1), 'SAME')
image = tf.nn.bias_add(conv_layer, bias)
elif layer[0] == 'r':
image = tf.nn.relu(image)
else:
image = tf.nn.max_pool(image, (1, 2, 2, 1), (1, 2, 2, 1), 'SAME')
network[layer] = image
return(network)
original_layer = 'relu4_2'
style_layers = ['relu1_1', 'relu2_1', 'relu3_1', 'relu4_1', 'relu5_1']
normalization_mean, network_weights = extract_net_info(vgg_path)
shape = (1,) + original_image.shape
style_shape = (1,) + style_image.shape
original_features = {}
style_features = {}
image = tf.placeholder('float', shape=shape)
vgg_net = vgg_network(network_weights, image)
original_minus_mean = original_image - normalization_mean
original_norm = np.array([original_minus_mean])
original_features[original_layer] = sess.run(vgg_net[original_layer], feed_dict={image: original_norm})
# +
image = tf.placeholder('float', shape=style_shape)
vgg_net = vgg_network(network_weights, image)
style_minus_mean = style_image - normalization_mean
style_norm = np.array([style_minus_mean])
for layer in style_layers:
layer_output = sess.run(vgg_net[layer], feed_dict={image: style_norm})
layer_output = np.reshape(layer_output, (-1, layer_output.shape[3]))
style_gram_matrix = np.matmul(layer_output.T, layer_output) / layer_output.size
style_features[layer] = style_gram_matrix
# -
initial = tf.random_normal(shape) * 0.256
image = tf.Variable(initial)
vgg_net = vgg_network(network_weights, image)
original_loss = original_image_weight * (2 * tf.nn.l2_loss(vgg_net[original_layer] - original_features[original_layer]) /
original_features[original_layer].size)
style_loss = 0
style_losses = []
for style_layer in style_layers:
layer = vgg_net[style_layer]
feats, height, width, channels = [x.value for x in layer.get_shape()]
size = height * width * channels
features = tf.reshape(layer, (-1, channels))
style_gram_matrix = tf.matmul(tf.transpose(features), features) / size
style_expected = style_features[style_layer]
style_losses.append(2 * tf.nn.l2_loss(style_gram_matrix - style_expected) / style_expected.size)
style_loss += style_image_weight * tf.reduce_sum(style_losses)
total_var_x = sess.run(tf.reduce_prod(image[:,1:,:,:].get_shape()))
total_var_y = sess.run(tf.reduce_prod(image[:,:,1:,:].get_shape()))
first_term = regularization_weight * 2
second_term_numerator = tf.nn.l2_loss(image[:,1:,:,:] - image[:,:shape[1]-1,:,:])
second_term = second_term_numerator / total_var_y
third_term = (tf.nn.l2_loss(image[:,:,1:,:] - image[:,:,:shape[2]-1,:]) / total_var_x)
total_variation_loss = first_term * (second_term + third_term)
loss = original_loss + style_loss + total_variation_loss
style_layer = 'relu2_1'
layer = vgg_net[style_layer]
feats, height, width, channels = [x.value for x in layer.get_shape()]
size = height * width * channels
features = tf.reshape(layer, (-1, channels))
style_gram_matrix = tf.matmul(tf.transpose(features), features) / size
style_expected = style_features[style_layer]
style_losses.append(2 * tf.nn.l2_loss(style_gram_matrix - style_expected) / style_expected.size)
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
train_step = optimizer.minimize(loss)
sess.run(tf.global_variables_initializer())
for i in range(generations):
sess.run(train_step)
if (i+1) % output_generations == 0:
print('Generation {} out of {}, loss: {}'.format(i + 1, generations,sess.run(loss)))
image_eval = sess.run(image)
best_image_add_mean = image_eval.reshape(shape[1:]) + normalization_mean
output_file = 'temp_output_{}.jpg'.format(i)
scipy.misc.imsave(output_file, best_image_add_mean)
image_eval = sess.run(image)
best_image_add_mean = image_eval.reshape(shape[1:]) + normalization_mean
output_file = 'final_output.jpg'
scipy.misc.imsave(output_file, best_image_add_mean)
| Section03/Applying Stylenet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="26xb3SKeUfXh"
# ## Instalação das dependências
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="U_W-gG5kUfXh" outputId="adcac35e-702a-4ae6-80ed-7f128853dd10"
# !pip install torch
# !pip install espnet==0.9.3
# -
# ## ESPNet
#
# Os modelos deste trabalho foram treinados utilizando o [ESPNet](https://github.com/espnet/espnet)
# .
#
# ESPnet é um conjunto de ferramentas para processamento de fala fim-a-fim (E2E) focado principalmente em reconhecimento de fala e síntese de fala. O framework do ESPNet utilizado neste trabalho é o Pytorch.
# + [markdown] id="Mv23KLr_UfXh"
# ## Imports
# + colab={"base_uri": "https://localhost:8080/"} id="yhbp95DXUfXh" outputId="c24d2b92-bc0a-4dc2-9b66-d4e9ba43fbec"
import os
import io
import copy
import yaml
import torch
import shutil
import zipfile
import librosa
import numpy as np
from pathlib import Path
from os.path import abspath, join, exists, dirname, isfile, isdir
from typing import Tuple, Any, Dict, Union, TypeVar
#Importa os módulos do ESPNet necessários
from espnet2.tasks.asr import ASRTask
#ASRTask é a classe principal de uma tarefa de ASR (Automatic Speech Recognition)
#Carrega as informações para carregar um modelo a partir de um arquivo yaml
#o mesmo utilizado no treinamento
from espnet.nets.beam_search import BeamSearch
#Beam Search é o algoritmo de decodificação proposto pelos autores da ferramenta
#Chamado de joint decoding, para cada frame a hipotese é balanceada entre uma rede
#com CTC loss e uma rede de atenção. A justificativa é que a rede CTC acelera o aprendizado
#do alinhamento da rede de atenção, principalmente Transformers, naturalmente difíceis de se treinar
from espnet.nets.scorers.ctc import CTCPrefixScorer
from espnet.nets.scorers.length_bonus import LengthBonus
#Inicialização e configuração do algoritmo de BeamSearch.
from espnet2.text.build_tokenizer import build_tokenizer
from espnet2.text.token_id_converter import TokenIDConverter
#Métodos úteis para conversão de tokens usados pelo treinamento pelos caracteres ortográficos
from espnet2.main_funcs.calculate_all_attentions import calculate_all_attentions
#Método que executa um passo forward na rede de atenção e salva os valores das matrizes
#de atenção para os labels de saída. Útil para plotar o alinhamento dos modelos de atenção
from espnet2.torch_utils.device_funcs import to_device
#Útil
# %matplotlib inline
import librosa.display
import ipywidgets as widgets
import matplotlib.pyplot as plt
from IPython.display import Image, display, Audio
from google_drive_downloader import GoogleDriveDownloader as gdd
# + [markdown] id="cgIOcyGPUfXg"
# # Modelos
# + id="CnlVEAnVUfXh"
MODEL_TYPES = ["rnn", "convrnn", "transformer"]
model_type_d = {"rnn":"rnn", "convrnn":"vggrnn", "transformer":"transformer"}
TOKEN_TYPES = ["char", "subword"]
token_type_d = {'char': 'char', 'subword': 'bpe'}
MODELS_URLS = {
"rnn_char" : "1c_J3MEEPQXhaYSYTMy-Pp6Wm7g4F3ppo",
"vggrnn_char" : "12SAYVc8LMDEg9Hm5vcVnwIw_H1xqH8Dh",
"transformer_char" : "1Sm_LZkna8RMCxWBCdoZwdPHecxJ5X24F",
"rnn_bpe" : "<KEY>
"vggrnn_bpe" : "<KEY>",
"transformer_bpe" : "<KEY>"
}
# + id="-_wR7dw-UfXh"
def download_model(model_type, token_type):
fname = f"asr_train_commonvoice_{model_type}_raw_{token_type}_valid.acc.ave.zip"
if not os.path.exists(fname):
print(f"Downloading {fname}")
gdd.download_file_from_google_drive(
file_id=MODELS_URLS[f"{model_type}_{token_type}"],
dest_path=f"./{fname}"
)
else:
print(f"Model file {fname} exists")
return fname
# + id="c3W0JXHFUfXh"
class Result(object):
def __init__(self) -> None:
self.text = None
self.tokens_txt = None
self.tokens_int = None
self.ctc_posteriors = None
self.attention_weights = None
self.encoded_vector = None
self.audio_samples = None
self.mel_features = None
class ASR(object):
def __init__(
self,
zip_model_file: Union[Path, str],
) -> None:
self.zip_model_file = abspath(zip_model_file)
self.device = 'cpu'
self.model = None
self.beam_search = None
self.tokenizer = None
self.converter = None
self.global_cmvn = None
self.extract_zip_model_file(self.zip_model_file)
def extract_zip_model_file(self, zip_model_file: str) -> Dict[str, Any]:
"""Extrai os dados de um zip contendo o arquivo com o estado do modelo e configurações
Args:
zip_model_file (str): ZipFile do modelo gerado dos scripts de treinamento
Raises:
ValueError: Se o arquivo não for correto
FileNotFoundError: Se o arquivo zip não contiver os arquivos necessários
Returns:
Dict[str, Any]: Dicionário do arquivo .yaml utilizado durante o treinamento para carregar o modelo corretamente
"""
print("Unzipping model")
if not zipfile.is_zipfile(zip_model_file):
raise ValueError(f"File {zip_model_file} is not a zipfile")
else:
zipfile.ZipFile(zip_model_file).extractall(dirname(zip_model_file))
check = ['exp', 'meta.yaml']
if not all([x for x in check]):
raise FileNotFoundError
print("Load yaml file")
with open('meta.yaml') as f:
meta = yaml.load(f, Loader=yaml.FullLoader)
model_stats_file = meta['files']['asr_model_file']
asr_model_config_file = meta['yaml_files']['asr_train_config']
self.model_config = {}
with open(asr_model_config_file) as f:
self.model_config = yaml.load(f, Loader=yaml.FullLoader)
try:
self.global_cmvn = self.model_config['normalize_conf']['stats_file']
except KeyError:
self.global_cmvn = None
print(f'Loading model config from {asr_model_config_file}')
print(f'Loading model state from {model_stats_file}')
#Build Model
print('Building model')
self.model, _ = ASRTask.build_model_from_file(
asr_model_config_file, model_stats_file, self.device
)
self.model.to(dtype=getattr(torch, 'float32')).eval()
#print("Loading extra modules")
self.build_beam_search()
self.build_tokenizer()
def build_beam_search(self, ctc_weight: float = 0.4, beam_size: int = 1):
"""Constroi o objeto de decodificação beam_search.
Esse objeto faz a decodificação do vetor de embeddings da saída da parte encoder
do modelo passando pelos decoders da rede que são o módulo CTC e Transformer ou RNN.
Como:
Loss = (1-λ)*DecoderLoss + λ*CTCLoss
Se ctc_weight=1 apenas o módulo CTC será usado na decodificação
Args:
ctc_weight (float, optional): Peso dado ao módulo CTC da rede. Defaults to 0.4.
beam_size (int, optional): Tamanho do feixe de busca durante a codificação. Defaults to 1.
"""
scorers = {}
ctc = CTCPrefixScorer(ctc=self.model.ctc, eos=self.model.eos)
token_list = self.model.token_list
scorers.update(
decoder=self.model.decoder,
ctc=ctc,
length_bonus=LengthBonus(len(token_list)),
)
#Variáveis com os pesos para cada parte da decodificação
#lm referente à modelos de linguagem não são utilizados aqui mas são necessários no objeto
weights = dict(
decoder=1.0 - ctc_weight,
ctc=ctc_weight,
lm=1.0,
length_bonus=0.0,
)
#Cria o objeto beam_search
self.beam_search = BeamSearch(
beam_size=beam_size,
weights=weights,
scorers=scorers,
sos=self.model.sos,
eos=self.model.eos,
vocab_size=len(token_list),
token_list=token_list,
pre_beam_score_key=None if ctc_weight == 1.0 else "full",
)
self.beam_search.to(device=self.device, dtype=getattr(torch, 'float32')).eval()
for scorer in scorers.values():
if isinstance(scorer, torch.nn.Module):
scorer.to(device=self.device, dtype=getattr(torch, 'float32')).eval()
def build_tokenizer(self):
"""Cria um objeto tokenizer para conversão dos tokens inteiros para o dicionário
de caracteres correspondente.
Caso o modelo possua um modelo BPE de tokenização, ele é utilizado. Se não, apenas a lista
de caracteres no arquivo de configuração é usada.
"""
token_type = self.model_config['token_type']
if token_type == 'bpe':
bpemodel = self.model_config['bpemodel']
self.tokenizer = build_tokenizer(token_type=token_type, bpemodel=bpemodel)
else:
self.tokenizer = build_tokenizer(token_type=token_type)
self.converter = TokenIDConverter(token_list=self.model.token_list)
def get_layers(self) -> Dict[str, Dict[str, torch.Size]]:
"""Retorna as camadas nomeadas e os respectivos shapes para todos os módulos da rede.
Os módulos são:
Encoder: RNN, VGGRNN, TransformerEncoder
Decoder: RNN, TransformerDecoder
CTC
Returns:
Dict[str, Dict[str, torch.Size]]: Dicionário de cada módulo com seus respectivos layers e shape
"""
r = {}
r['frontend'] = {x: self.model.frontend.state_dict()[x].shape
for x in self.model.frontend.state_dict().keys()}
r['specaug'] = {x: self.model.specaug.state_dict()[x].shape
for x in self.model.specaug.state_dict().keys()}
r['normalize'] = {x: self.model.normalize.state_dict()[x].shape
for x in self.model.normalize.state_dict().keys()}
r['encoder'] = {x: self.model.encoder.state_dict()[x].shape
for x in self.model.encoder.state_dict().keys()}
r['decoder'] = {x: self.model.decoder.state_dict()[x].shape
for x in self.model.decoder.state_dict().keys()}
r['ctc'] = {x: self.model.ctc.state_dict()[x].shape
for x in self.model.ctc.state_dict().keys()}
return r
def frontend(self, audiofile: Union[Path, str, bytes], normalize: bool = True) -> Tuple[torch.Tensor, torch.Tensor]:
"""Executa o frontend do modelo, transformando as amostras de áudio em parâmetros log mel spectrogram
Args:
audiofile (Union[Path, str]): arquivo de áudio
Returns:
Tuple[torch.Tensor, torch.Tensor]: Parâmetros, Tamanho do vetor de parâmetros
"""
if isinstance(audiofile, str):
audio_samples, rate = librosa.load(audiofile, sr=16000)
elif isinstance(audiofile, bytes):
audio_samples, rate = librosa.core.load(io.BytesIO(audiofile), sr=16000)
else:
raise ValueError("Failed to load audio file")
if isinstance(audio_samples, np.ndarray):
audio_samples = torch.tensor(audio_samples)
audio_samples = audio_samples.unsqueeze(0).to(getattr(torch, 'float32'))
lengths = audio_samples.new_full([1], dtype=torch.long, fill_value=audio_samples.size(1))
features, features_length = self.model.frontend(audio_samples, lengths)
if normalize:
features, features_length = self.model.normalize(features, features_length)
return features, features_length
def specaug(self, features: torch.Tensor, features_length: torch.Tensor) -> Tuple[torch.Tensor, torch.Tensor]:
"""Executa o módulo specaug, da parte de 'data augmentation'.
Útil para visualização apenas.
Não é utilizado na inferência, apenas no treinamento.
Args:
features (torch.Tensor): Parâmetros
features_length (torch.Tensor): tamanho do vetor de parâmetros
Returns:
Tuple[torch.Tensor, torch.Tensor]: Parâmetros com máscaras temporais, em frequência e distoção. Tamanho dos vetores
"""
return self.model.specaug(features, features_length)
def __del__(self) -> None:
"""Remove os arquivos temporários
"""
for f in ['exp', 'meta.yaml']:
print(f"Removing {f}")
ff = join(dirname(self.zip_model_file), f)
if exists(ff):
if isdir(ff):
shutil.rmtree(ff)
elif isfile(ff):
os.remove(ff)
else:
raise ValueError("Error ao remover arquivos temporários")
@torch.no_grad()
def recognize(self, audiofile: Union[Path, str, bytes]) -> Result:
result = Result()
if isinstance(audiofile, str):
audio_samples, rate = librosa.load(audiofile, sr=16000)
elif isinstance(audiofile, bytes):
audio_samples, rate = librosa.core.load(io.BytesIO(audiofile), sr=16000)
else:
raise ValueError("Failed to load audio file")
result.audio_samples = copy.deepcopy(audio_samples)
#a entrada do modelo é torch.tensor
if isinstance(audio_samples, np.ndarray):
audio_samples = torch.tensor(audio_samples)
audio_samples = audio_samples.unsqueeze(0).to(getattr(torch, 'float32'))
lengths = audio_samples.new_full([1], dtype=torch.long, fill_value=audio_samples.size(1))
batch = {"speech": audio_samples, "speech_lengths": lengths}
batch = to_device(batch, device=self.device)
#model encoder
enc, _ = self.model.encode(**batch)
#model decoder
nbest_hyps = self.beam_search(x=enc[0])
#Apenas a melhor hipótese
best_hyps = nbest_hyps[0]
#Conversão de tokenids do treinamento para texto
token_int = best_hyps.yseq[1:-1].tolist()
token_int = list(filter(lambda x: x != 0, token_int))
token = self.converter.ids2tokens(token_int)
text = self.tokenizer.tokens2text(token)
#Preenche o objeto result
result.text = text
result.encoded_vector = enc[0] #[0] remove dimensão de batch
#calcula todas as matrizes de atenção
#
text_tensor = torch.Tensor(token_int).unsqueeze(0).to(getattr(torch, 'long'))
batch["text"] = text_tensor
batch["text_lengths"] = text_tensor.new_full([1], dtype=torch.long, fill_value=text_tensor.size(1))
result.attention_weights = calculate_all_attentions(self.model, batch)
result.tokens_txt = token
#CTC posteriors
logp = self.model.ctc.log_softmax(enc.unsqueeze(0))[0]
result.ctc_posteriors = logp.exp_().numpy()
result.tokens_int = best_hyps.yseq
result.mel_features, _ = self.frontend(audiofile, normalize=False)
return result
def __call__(self, input: Union[Path, str, bytes]) -> Result:
return self.recognize(input)
asr = None
# + [markdown] id="u1wjPAJxUfXh"
# ## Escolha um tipo de modelo
# + colab={"base_uri": "https://localhost:8080/", "height": 113, "referenced_widgets": ["d2f91f61e4a34a9d916b7661da8889f8", "2dada558847e4553a2946838e5409f77", "25e36958528742ccae66e70e1b50941a"]} id="gtNmySVkUfXh" outputId="242f2a3b-ea10-4ddd-8027-c40feeb32041"
w1 = widgets.RadioButtons(
options=MODEL_TYPES,
value='rnn',
description='Model type:',
disabled=False
)
display(w1)
# + [markdown] id="Xd1u0q7eUfXh"
# ## Escolha um formato dos símbolos $Y$
#
# + colab={"base_uri": "https://localhost:8080/", "height": 81, "referenced_widgets": ["dbd51b9d0b594fd0a32e1ea5b84cf184", "47d3206e537f4774bb2e2f8bd4d98511", "d2ac50ce3ba24d33ba8b75344d878f27"]} id="BKbaQqQhUfXh" outputId="d4fc3b0b-4cbc-42be-bcc3-0b3cbdb5ca97"
w2 = widgets.RadioButtons(
options=TOKEN_TYPES,
value='char',
description='Token type:',
disabled=False
)
display(w2)
# + [markdown] id="nJt8xfujUfXh"
# ## Faça o upload de um arquivo de áudio para inferência (wav, 16kHz)
#
# + colab={"base_uri": "https://localhost:8080/", "height": 49, "referenced_widgets": ["7853e8c6f34044169cb115e70a0d5924", "98a4267b1e4f47059908c207f1165c84", "73c53552b4e542fd80739ca2aef4fc8b"]} id="GRQ5fnx6UfXh" outputId="93d71cc2-88a4-489c-d12c-406db685a9a8"
uploader = widgets.FileUpload()
display(uploader)
# + [markdown] id="ogJgJn5FUfXh"
# ## Execute a célula abaixo para carregar o modelo selecionado e realizar a inferência do áudio
#
# É possível alterar os modelos nos diálogos acima e executar apenas a célula abaixo para fazer uma nova inferência em um novo modelo. Para visualizar as figuras para o novo modelo, da mesma forma, basta executar as células abaixo.
# + colab={"base_uri": "https://localhost:8080/"} id="J1c4lBkQUfXh" outputId="a3890ea5-ab19-448f-d507-d50220876a7d"
if isinstance(asr, ASR):
del asr
asr = None
model_type = model_type_d[w1.value]
token_type = token_type_d[w2.value]
try:
audio_file = uploader.value[list(uploader.value.keys())[0]]['content']
except IndexError:
raise ValueError("Faça o upload de um arquivo de áudio")
asr_tag = "asr_train_commonvoice_"+model_type+"_raw_"+token_type
#Faz o downlooad do modelo caso o arquivo ainda não exista
model_file = download_model(model_type, token_type)
#Unzip do arquivo do modelo no diretório local e carregamento do objeto da classe ASR
#A classe ASR encapsula um modelo ESPNet em asr.model
asr = ASR(model_file)
#O método recognize recebe um arquivo de áudio ou um buffer de memória e fornece a melhor hipotese do modelo
results = asr.recognize(audio_file)
print(f"\nHipotese: {results.text}")
# -
# ### Arquitetura da rede completa
Image(f'imagens/{model_type}.png')
# ### Arquitetura da rede CTC
Image(f'imagens/ctc.png')
# ### Acurácia de treinamento e validação
# + colab={"base_uri": "https://localhost:8080/", "height": 497} id="exa3xX2ZG4R0" outputId="8038cc5c-06a3-4032-e99e-418bff703386"
Image(f'exp/{asr_tag}/images/acc.png')
# -
# ### Função custo do modelo de atenção
# + colab={"base_uri": "https://localhost:8080/", "height": 497} id="uQeIA2CWGy0r" outputId="373fc500-582f-45d2-a588-550653a77b1a"
Image(f'exp/{asr_tag}/images/loss_att.png')
# -
# ### Função custo do modelo CTC
# + colab={"base_uri": "https://localhost:8080/", "height": 497} id="qFV8vC5fG1Fq" outputId="6b9bcb41-357d-48bc-b1dd-7a2275cadbb6"
Image(f'exp/{asr_tag}/images/loss_ctc.png')
# -
# ### Função custo total
#
# A função custo total obedece o esquema de MTL (MultiTask Learning) equilibrando as funções custo anteriores através de um hiperparâmetro $\lambda$.
#
# Modelos de atenção com redes recorrentes(RNN e ConvRNN) treinaram com $\lambda=0.5$ dando um peso igual para a rede CTC e para a rede de atenção. O modelo transformer treinou com $\lambda=0.3$, dando um peso 0.3 para a função custo CTC e $0.7$ para o modelo transformer.
#
# A equação final da função custo é:
#
# $L_{total} = \lambda L_{ctc} + (1-\lambda)L_{att}$
# + colab={"base_uri": "https://localhost:8080/", "height": 497} id="4YoLDldHG7Na" outputId="d11378d8-3fe7-408d-d739-f82c1f11126e"
Image(f'exp/{asr_tag}/images/loss.png')
# -
# ### Learning rate durante o treinamento do modelo
# + colab={"base_uri": "https://localhost:8080/", "height": 497} id="ZGb2ndTEG9CS" outputId="84201344-8e74-4809-ac83-ea473a88b1e0"
Image(f'exp/{asr_tag}/images/lr_0.png')
# + [markdown] id="o2vLTyCGUfXh"
# ## Descrição do modelo
#
# Os módulos do Pytorch usados no modelo, com os respectivos argumentos, são descritos abaixo
# + colab={"base_uri": "https://localhost:8080/"} id="ORBh4EGKUfXh" outputId="bf79e5a5-49f8-40df-9704-290267d6d152"
asr.model
# + [markdown] id="Gd4P7GROUfXh"
# ## Camadas do modelo
# + colab={"base_uri": "https://localhost:8080/"} id="aw6cX70rUfXi" outputId="7d9f5476-a943-4ad3-836b-5fa3df688fb5"
asr.get_layers()
# + [markdown] id="klsQjDzjUfXi"
# ## Inferência
# + colab={"base_uri": "https://localhost:8080/", "height": 249} id="q3pG2b-2UfXi" outputId="ea43ea67-4b42-4a8b-a9c0-3d6a9b3bd5ce"
#Audio
plt.figure(figsize=(10, 2))
librosa.display.waveplot(results.audio_samples, sr=16000)
print(f"Hypotese: {results.text}")
Audio(audio_file)
# + [markdown] id="AmarGl15UfXi"
# ## Parâmetros mel-fbank (dim=80)
#
# (frontend): DefaultFrontend(
#
# (stft): Stft(n_fft=512, win_length=512, hop_length=128, center=True, normalized=False, onesided=True)
#
# (frontend): Frontend()
#
# (logmel): LogMel(sr=16000, n_fft=512, n_mels=80, fmin=0, fmax=8000.0, htk=False)
#
# )
# + colab={"base_uri": "https://localhost:8080/", "height": 273} id="b4hYzpCSUfXi" outputId="b3db00af-281b-4cb3-ab12-e40bc240a0a9"
fig, axs = plt.subplots(2, 1)
axs[0].plot(results.audio_samples)
axs[0].set_xlim(xmin=0, xmax=len(results.audio_samples))
axs[0].set_title("Amostras de áudio e LogMel Espectrograma")
axs[1].matshow(results.mel_features[0].T)
fig.tight_layout()
# + [markdown] id="FiPx-IBxUfXi"
# ## Specaug
#
# Data augmentation utilizada durante o treinamento, aqui mostrada apenas para visualização.
# Máscaras no tempo e na frequência, além de time warping são aplicadas aleatoriamente. É possível executar a célula abaixo várias vezes para visualizar diferentes resultados.
#
# (specaug): [SpecAug](https://arxiv.org/pdf/1904.08779)(
#
# (time_warp): TimeWarp(window=5, mode=bicubic)
#
# (freq_mask): MaskAlongAxis(mask_width_range=[0, 30], num_mask=2, axis=freq)
#
# (time_mask): MaskAlongAxis(mask_width_range=[0, 40], num_mask=2, axis=time)
#
# )
# + id="t0zhiOTAUfXi" outputId="52736ad2-67c2-4edc-f2b0-490a05c34f6d"
specaug_features,_ = asr.specaug(results.mel_features, results.mel_features.shape[1:])
plt.matshow(specaug_features[0].T)
plt.title("Data Augmentation")
#specaug_features = ASR.model.specaug(results.mel_features, results.mel_features.shape)
# + [markdown] id="ohMdD4PIUfXi"
# ## Resultado da camada Encoder
#
# Diferentes redes de encoder fazem diferentes tipos de transformação dos atributos de entrada $\mathbf{X}$. O vetor enc tem diferentes dimensões em cada um deles.
# + colab={"base_uri": "https://localhost:8080/", "height": 252} id="6wK6WnPpUfXi" outputId="0349a6df-7408-4ab6-8037-4fd2e295245c"
enc = results.encoded_vector
im = plt.matshow(enc.detach().numpy())
plt.colorbar(im)
# + [markdown] id="-jPXlabRUfXi"
# ## Visualização das matrizes de atenção
#
# Cada modelo de atenção possui diferentes subcamadas de atenção, utilize o menu abaixo para selecionar uma delas em cada modelo. O modelo transformer, baseado em Multihea-Attention é treinado com 8 heads mas aqui mostra apenas a primeira delas. Contém as 12 camadas do encoder e as 6 camadas do decoder.
# + colab={"base_uri": "https://localhost:8080/", "height": 475, "referenced_widgets": ["2671f930a01e476db4e9224b30fad763", "2dee31a6ad0f4720a48fe5237e205333", "e7fbf806b5284c6183a16aaf192d060a", "3838a2146ccc4645943232549370d221", "44d0a3d511c24d7c8b6b3bdfda2f81bf", "a3360cd5fd824b6da7031de86f1d1109", "01fe550043524922b37dd9f947757550"]} id="1mscnh3lUfXi" outputId="6f1ebc09-4ff4-4141-932c-f958a3c9e7b0"
# %pylab inline
from ipywidgets import interact, interactive, Layout
def f(x):
attn = results.attention_weights[x][0]
if len(attn.shape) == 3:
attn = attn[0]
fig, ax = plt.subplots(1, figsize=(10, 5))
ax.set_title(f"Attention {x}")
im = ax.matshow(attn, aspect="auto")
fig.colorbar(im)
if not 'self_attn' in x or not 'encoder.encoders' in x:
ax.set_yticks(range(len(results.tokens_txt)))
ax.set_yticklabels(results.tokens_txt)
fig.tight_layout()
int_widget = interactive(f, x=list(results.attention_weights.keys()))
#int_widget.children[0].layout = Layout(width='100px')
display(int_widget)
# + [markdown] id="qQ3pQa4YUfXi"
# ## Visualização dos CTC Posteriors
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="rTJDJKkIUfXi" outputId="0f00791b-2c29-4b69-b405-7fa0676743e5"
prob = results.ctc_posteriors[0]
fig, ax = plt.subplots(1, figsize=(10, 7))
ax.set_title("CTC posterior")
vs = set(results.tokens_int.tolist())
vs.add(0)
for n, i in enumerate(vs):
v = asr.converter.ids2tokens([i])[0]
ax.plot(prob[:, i], label=v, linestyle="-" if len(v) == 1 else "--")
ax.legend(loc='center left', bbox_to_anchor=(1, 0.5))
ax.set_xlim(0, len(results.encoded_vector)-1)
fig.tight_layout()
| egs2/ptbr/asr1/relatorio/.ipynb_checkpoints/Projeto_ML_ASR-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Random forests: finding optimal criterion ('gini' or 'entropy')
"""Random forests: finding optimal criterion ('gini' or 'entropy')
"""
# import libraries
import pandas as pd
import numpy as np
from sklearn.ensemble import RandomForestClassifier
from sklearn import preprocessing
from sklearn.model_selection import TimeSeriesSplit
# import data
df = pd.read_csv('data/SCADA_downtime_merged.csv', skip_blank_lines=True)
# list of turbines to plot
list1 = list(df['turbine_id'].unique())
# sort turbines in ascending order
list1 = sorted(list1, key=int)
# list of categories
list2 = list(df['TurbineCategory_id'].unique())
# remove NaN from list
list2 = [g for g in list2 if g >= 0]
# sort categories in ascending order
list2 = sorted(list2, key=int)
# categories to remove
list2 = [m for m in list2 if m not in (1, 12, 13, 14, 15, 17, 21, 22)]
# empty list to hold optimal n values for all turbines
num = []
# empty list to hold minimum error readings for all turbines
err = []
# filter only data for turbine x
for x in list1:
dfx = df[(df['turbine_id'] == x)].copy()
# copying fault to new column (mins) (fault when turbine category id is y)
for y in list2:
def f(c):
if c['TurbineCategory_id'] == y:
return 0
else:
return 1
dfx['mins'] = dfx.apply(f, axis=1)
# sort values by timestamp in descending order
dfx = dfx.sort_values(by='timestamp', ascending=False)
# reset index
dfx.reset_index(drop=True, inplace=True)
# assigning value to first cell if it's not 0 with a large number
if dfx.loc[0, 'mins'] == 0:
dfx.set_value(0, 'mins', 0)
else:
# to allow the following loop to work
dfx.set_value(0, 'mins', 999999999)
# using previous value's row to evaluate time
for i, e in enumerate(dfx['mins']):
if e == 1:
dfx.at[i, 'mins'] = dfx.at[i - 1, 'mins'] + 10
# sort in ascending order
dfx = dfx.sort_values(by='timestamp')
# reset index
dfx.reset_index(drop=True, inplace=True)
# convert to hours, then round to nearest hour
dfx['hours'] = dfx['mins'].astype(np.int64)
dfx['hours'] = dfx['hours']/60
# round to integer
dfx['hours'] = round(dfx['hours']).astype(np.int64)
# > 48 hours - label as normal (999)
def f1(c):
if c['hours'] > 48:
return 999
else:
return c['hours']
dfx['hours'] = dfx.apply(f1, axis=1)
# filter out curtailment - curtailed when turbine is pitching outside
# 0deg <= normal <= 3.5deg
def f2(c):
if 0 <= c['pitch'] <= 3.5 or c['hours'] != 999 or (
(c['pitch'] > 3.5 or c['pitch'] < 0) and (
c['ap_av'] <= (.1 * dfx['ap_av'].max()) or
c['ap_av'] >= (.9 * dfx['ap_av'].max()))):
return 'normal'
else:
return 'curtailed'
dfx['curtailment'] = dfx.apply(f2, axis=1)
# filter unusual readings, i.e., for normal operation, power <= 0 in
# operating wind speeds, power > 100 before cut-in, runtime < 600 and
# other downtime categories
def f3(c):
if c['hours'] == 999 and ((
3 < c['ws_av'] < 25 and (
c['ap_av'] <= 0 or c['runtime'] < 600 or
c['EnvironmentalCategory_id'] > 1 or
c['GridCategory_id'] > 1 or
c['InfrastructureCategory_id'] > 1 or
c['AvailabilityCategory_id'] == 2 or
12 <= c['TurbineCategory_id'] <= 15 or
21 <= c['TurbineCategory_id'] <= 22)) or
(c['ws_av'] < 3 and c['ap_av'] > 100)):
return 'unusual'
else:
return 'normal'
dfx['unusual'] = dfx.apply(f3, axis=1)
# round to 6 hour intervals to reduce number of classes
def f4(c):
if 1 <= c['hours'] <= 6:
return 6
elif 7 <= c['hours'] <= 12:
return 12
elif 13 <= c['hours'] <= 18:
return 18
elif 19 <= c['hours'] <= 24:
return 24
elif 25 <= c['hours'] <= 30:
return 30
elif 31 <= c['hours'] <= 36:
return 36
elif 37 <= c['hours'] <= 42:
return 42
elif 43 <= c['hours'] <= 48:
return 48
else:
return c['hours']
dfx['hours6'] = dfx.apply(f4, axis=1)
# change label for unusual and curtailed data (9999), if originally
# labelled as normal
def f5(c):
if c['unusual'] == 'unusual' or c['curtailment'] == 'curtailed':
return 9999
else:
return c['hours6']
# apply to new column specific to each fault
dfx['hours_%s' % y] = dfx.apply(f5, axis=1)
# drop unnecessary columns
dfx = dfx.drop('hours6', axis=1)
dfx = dfx.drop('hours', axis=1)
dfx = dfx.drop('mins', axis=1)
dfx = dfx.drop('curtailment', axis=1)
dfx = dfx.drop('unusual', axis=1)
features = [
'ap_av', 'ws_av', 'wd_av', 'pitch', 'ap_max', 'ap_dev',
'reactive_power', 'rs_av', 'gen_sp', 'nac_pos']
# separate features from classes for classification
classes = [col for col in dfx.columns if 'hours' in col]
# list of columns to copy into new df
list3 = features + classes + ['timestamp']
df2 = dfx[list3].copy()
# drop NaNs
df2 = df2.dropna()
X = df2[features]
# normalise features to values b/w 0 and 1
X = preprocessing.normalize(X)
Y = df2[classes]
# convert from pd dataframe to np array
Y = Y.as_matrix()
criterion = ['gini', 'entropy']
scores = []
# cross validation using time series split
tscv = TimeSeriesSplit(n_splits=5)
# looping for each value of c and defining random forest classifier
for c in criterion:
rf = RandomForestClassifier(criterion=c, n_jobs=-1)
# empty list to hold score for each cross validation fold
p1 = []
# looping for each cross validation fold
for train_index, test_index in tscv.split(X):
# split train and test sets
X_train, X_test = X[train_index], X[test_index]
Y_train, Y_test = Y[train_index], Y[test_index]
# fit to classifier and predict
rf1 = rf.fit(X_train, Y_train)
pred = rf1.predict(X_test)
# accuracy score
p2 = np.sum(np.equal(Y_test, pred))/Y_test.size
# add to list
p1.append(p2)
# average score across all cross validation folds
p = sum(p1)/len(p1)
scores.append(p)
# changing to misclassification error
MSE = [1 - x for x in scores]
# determining best n
optimal = criterion[MSE.index(min(MSE))]
num.append(optimal)
err.append(min(MSE))
d = pd.DataFrame(num, columns=['criterion'])
d['error'] = err
d['turbine'] = list1
d
| jupyter-notebooks/optimisation/random-forest-criterion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Python
# ## data types,
# ## basic math function
#
# 1.
# 2.
# 3.
#
#
# ## basic math
1 + 2
1 == 3
1 != 3
8 < 10
10 < 1000
10 >= 5
'Simon' == 'simon'
| UdemyPandas/2017-07-notebooks/01 python basic math.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Emekaborisama/Plot-and-identify-fraud-and-normal-transaction-from-a-credit-card-dataset/blob/master/Plot_and_identify_fraud_and_normal_transaction_from_a_credit_card_dataset.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="YW9mZFrR4JV5" colab_type="code" colab={}
#import packages and dependies
# + id="5_Qwlkhu4JWU" colab_type="code" colab={}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# + id="a9Ph_5Bt4JWt" colab_type="code" colab={}
#read our dataset via pd.read_csv
# + id="h-9HbNQk4JW8" colab_type="code" colab={}
dataset = pd.read_csv('creditcard.csv') #get dataset from Kaggle Creditcard Competition
# + id="m15hxlJ14JXN" colab_type="code" colab={} outputId="0ffb1e9e-c479-478f-b581-867f543b8609"
dataset
# + id="YurddGXN4JX6" colab_type="code" colab={} outputId="a3b84010-567f-495c-8012-d2d2c036cff9"
dataset.head(n=5)
# + id="Ou8V5iyY4JYU" colab_type="code" colab={} outputId="9947bc09-83b2-4679-a056-a6f7655c3bc7"
dataset.shape
# + id="ME3uAqDu4JYs" colab_type="code" colab={} outputId="378b885c-3de7-4379-a1e4-cca58f902b71"
dataset.isnull().values.any() #to check for empty or null row or column
# + id="x1IPegig4JZD" colab_type="code" colab={}
#check the amount of Fraud and Normal transaction
# + id="beqK9XrS4JZU" colab_type="code" colab={} outputId="8c274a80-918f-4d1b-8d4d-e55f1c13357c"
pd.value_counts(dataset['Class'], sort = True) #class comparison 0=normal 1=fraud
# + id="YbX7Q-Zc4JZv" colab_type="code" colab={}
#Assign a varible to it
# + id="VP69ZqMX4JaC" colab_type="code" colab={}
data = pd.value_counts(dataset['Class'],sort = True)
# + id="AkzC0C2Z4JaT" colab_type="code" colab={} outputId="197c8761-e0bc-42f3-dba2-c0d227edd1a0"
data
# + id="DBSi15Xc4Jal" colab_type="code" colab={}
#Plot a bar chart to represent our data
# + id="9SdTjWWx4Ja0" colab_type="code" colab={} outputId="c600e4e7-ca2e-4a8b-e01e-7cd966c5017b"
data.plot(kind = 'bar', rot=0)
plt.xticks(range(2),)
plt.title("Frequency by observation number")
plt.xlabel("Class")
plt.ylabel("Number of Observation");
# + id="CZWOiqH44JbW" colab_type="code" colab={}
normal_df = dataset[dataset.Class == 0] #save normal_df as normal transaction
# + id="1KQEQ9TP4Jbl" colab_type="code" colab={}
fraud_df = dataset[dataset.Class == 1] #do the same for frauds
# + id="v3ES3o0f4JcC" colab_type="code" colab={} outputId="16358141-0f13-42a9-a119-a16c7a5c184f"
normal_df.shape
# + id="3eGp23Wf4Jcl" colab_type="code" colab={} outputId="d84ef717-87b8-44be-8f3f-69a997ec580b"
fraud_df.shape
# + id="IzIro7CI4Jc2" colab_type="code" colab={}
#get the stats of our normal transaction
# + id="mvLVccc24JdE" colab_type="code" colab={} outputId="152efedb-b51a-4a7e-cb1b-9683e904f1b3"
normal_df.Amount.describe()
# + id="ZotqifhD4JdZ" colab_type="code" colab={}
#get the stats of our fraud transaction
# + id="lZ3V4ujY4Jdv" colab_type="code" colab={} outputId="9e2adf31-0bae-45dc-c42b-e1c59ce52975"
fraud_df.Amount.describe()
# + id="1-ejHkzT4JeK" colab_type="code" colab={}
#lets plot a scatter plot to see the amount of fraud transaction by hours
# + id="w3cUl2iv4JeX" colab_type="code" colab={} outputId="aa538497-48df-42f4-d015-12ba5ca9b620"
data = plt.scatter((normal_df.Time/(60*60)), normal_df.Amount, alpha=0.6, label='Normal')
plt.scatter((fraud_df.Time/(60*60)), fraud_df.Amount, alpha=0.9, label='Fraud')
plt.title("Amount of transaction by hour")
plt.xlabel("Transaction time as measured from first transaction in the dataset (hours)")
plt.ylabel('Amount (USD)')
plt.legend(loc='upper right')
plt.show()
# + id="PoBiFi4j4Jex" colab_type="code" colab={}
#where 0= normal and 1=fraud
# + id="GxW9QmA64JfB" colab_type="code" colab={}
# + id="za1PXzfi4JfN" colab_type="code" colab={}
#so we sucessfully ploted a Bar chart representing the amount of fraud and normal transaction
| Plot_and_identify_fraud_and_normal_transaction_from_a_credit_card_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Caracterização de Dispositivos Eletrônicos via Multímetros
# ## < <NAME>, <NAME>, Sicr<NAME> Santos >
# ### Disciplina DQF10441 Física Experimental II - Semestre 2021/1 EARTE
# ### [DQF - CCENS](http://alegre.ufes.br/ccens/departamento-de-quimica-e-fisica) - [UFES/Alegre](http://alegre.ufes.br/)
# #### Experimento realizado em 13 e 20/08/2021 - Relatório entregue em < 31/08/2021 >
# #### Professor : [<NAME> Jr.](https://github.com/rcolistete)
# + [markdown] slideshow={"slide_type": "slide"}
# # 1 - Objetivos
# + [markdown] slideshow={"slide_type": "fragment"}
# < Identifição da razão de se realizar o experimento.
#
# Parcialmente baseado em "Objetivos" do roteiro.
#
# Por exemplo, nesse experimento (fiquem à vontade em alterar) : >
# + [markdown] slideshow={"slide_type": "fragment"}
# Esse experimento visa inicialmente praticar circuito elétrico de uma malha com resistor, com fonte de tensão elétrica, protoboard e resistor. Para tanto é necessário o domínio da teoria envolvida de circuitos elétricos, seus componentes e suas grandezas. Bem como domínio da prática de medições das grandezas elétricas utilizando multímetros nos modos voltímetro, amperímetro e ohmímetro, sendo calculadas as incertezas de cada medida.
# + [markdown] slideshow={"slide_type": "fragment"}
# A segunda parte do experimento visa praticar circuitos eletrônicos de uma malha com resistor, diodo e LED (diodo emissor de luz). Para tanto é necessário o domínio da teoria envolvida de circuitos eletrônicos, seus componentes e suas grandezas. Bem como domínio da prática de medições das grandezas elétricas utilizando multímetros nos modos voltímetro e amperímetro para obter a curva $V \times I$ de caracterização de cada componente : resistor, diodo e LED. Sendo que as incertezas das medidas são calculadas e propagadas via software.
# + [markdown] slideshow={"slide_type": "slide"}
# # 2 - Teoria
# + [markdown] slideshow={"slide_type": "fragment"}
# < Resumo teórico, geralmente é advindo de um texto maior encontrado em livros de Física (que vem ser citados).
#
# Parcialmente baseada em "Teoria" e "Equipamento" do roteiro.
#
# Teoria resumida, tipicamente com 1 a 2 parágrafos, sobre certos tópicos e ítens. >
# + [markdown] slideshow={"slide_type": "fragment"}
# < Teoria de grandezas elétricas : tensão elétrica, corrente elétrica e resistência elétrica (e lei de Ohm).
#
# Segundo [HRW], ... >
# + [markdown] slideshow={"slide_type": "fragment"}
# < Teoria de circuitos elétricos de uma malha. Com imagem de um exemplo. >
# + [markdown] slideshow={"slide_type": "fragment"}
# < Teoria de condutores, isolantes e semicondutores elétricos. >
# + [markdown] slideshow={"slide_type": "subslide"}
# < Teoria de resistor (ôhmico) e código de cores. Com imagem/foto de exemplo. Com imagem do símbolo de resistor adotado em diagrama esquemático. >
# + [markdown] slideshow={"slide_type": "fragment"}
# < Teoria de diodo. Com imagem/foto de exemplo. Com imagem do símbolo de diodo adotado em diagrama esquemático. >
# + [markdown] slideshow={"slide_type": "fragment"}
# < Teoria de LED (diodo emissor de luz) e tipos/cores disponíveis. Com imagem/foto de exemplo. Com imagem do símbolo de LED adotado em diagrama esquemático. >
# + [markdown] slideshow={"slide_type": "fragment"}
# < Teoria de fonte de tensão elétrica. Com imagem/foto de exemplo. Com imagem do símbolo de fonte de tensão elétrica adotado em diagrama esquemático. >
# + [markdown] slideshow={"slide_type": "fragment"}
# < Teoria de princípio de funcionamento de voltímetro, amperímetro e ohmímetro. Com imagem/foto de exemplo. >
# + [markdown] slideshow={"slide_type": "slide"}
# # 3 - Procedimento Experimental
# + [markdown] slideshow={"slide_type": "fragment"}
# < Descrição precisa do que foi usado e como foi usado no experimento.
#
# Parcialmente baseado em "Equipamento" e "Procedimento Experimental" do roteiro.
#
# Nesta seção são descritos os procedimentos empregados para efetuar as medidas e são descritas as montagens experimentais utilizadas. Diagramas esquemáticos das experiências são bastante úteis pois facilitam a visualização. Este procedimento não é uma cópia do roteiro do experimento, podendo o relatório ser mais resumido que o roteiro.
#
# Por exemplo, nesse experimento (fiquem à vontade em alterar) : >
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 3.1 - Medindo Tensão, Corrente e Resistência Elétricas com Multímetro
# + [markdown] slideshow={"slide_type": "fragment"}
# Essa parte do experimento consiste de um circuito elétrico de uma malha com resistor, com fonte de tensão elétrica de 3,3/5V, protoboard, fios e resistor.
#
# Bem como 2 multímetros com pontas-de-prova e garras jacaré, sendo usados nos modos voltímetro, amperímetro e ohmímetro.
# + [markdown] slideshow={"slide_type": "fragment"}
# A montagem é feita sob orientação do professor, conectando a fonte de tensão 3,3/5V no protoboard, o resistor indicado pelo professor no protoboard, um multímetro em modo voltímetro conectado em paralelo ao resistor, outro multímetro em modo amperímetro em série ao resistor (entre o terminal positivo da fonte de tensão e um dos terminais do resistor).
#
# Todas as conexões devem ser feitas com a fonte de tensão elétrica desligada e os multímetros desligados. Ssempre deve ser desconectado um dos cabos do multímetro antes de ligar ou mudar a escala, conectando depois.
# + [markdown] slideshow={"slide_type": "subslide"}
# <center> Figura N - Diagrama esquemático do circuito elétrico com resistor. </center>
# + [markdown] slideshow={"slide_type": "fragment"}
# São feitas medições em todas as escalas possíveis de tensão elétrica contínua (DC) do multímetro, bem como medições em todas as escalas possíveis de corrente elétrica contínua (DC) do multímetro.
# + [markdown] slideshow={"slide_type": "fragment"}
# Ao final a resistência elétrica do resistor é medida via multímetro no modo ohmímetro, com o resistor desconectado do protoboard, novamente usando todas as escalas possíveis do multímetro.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 3.2 - Caracterização de Resistor, Diodo e LED com Multímetro
# + [markdown] slideshow={"slide_type": "fragment"}
# Essa segunda parte do experimento consiste de um circuito eletrônico de uma malha com resistor, com fonte de tensão elétrica regulável, protoboard, fios, 1-2 resistores, diodo e LED.
#
# Bem como 2 multímetros com pontas-de-prova e garras jacaré, sendo usados nos modos voltímetro e amperímetro.
# + [markdown] slideshow={"slide_type": "fragment"}
# A montagem e medições são feitas pelo professor, pois a fonte de tensão regulável é equipamento mais complexo e caro e por isso não está presente no kit de material fornecido aos grupos.
# + [markdown] slideshow={"slide_type": "fragment"}
# A montagem, se feita pelos alunos, é sob orientação do professor. Todas as conexões devem ser feitas com a fonte de tensão elétrica desligada e os multímetros desligados. Sempre deve ser desconectado um dos cabos do multímetro antes de ligar ou mudar a escala, conectando depois.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### 3.2.1 - Caracterização $V \times I$ de Resistor $R_C$
# + [markdown] slideshow={"slide_type": "fragment"}
# A montagem é feita conectando a fonte de tensão regulável no protoboard, os resistores indicados pelo professor em série no protoboard, um multímetro em modo voltímetro conectado em paralelo ao resistor carga, outro multímetro em modo amperímetro em série ao resistor carga (entre o resistor limitador e resistor carga).
# + [markdown] slideshow={"slide_type": "fragment"}
# <center> Figura N - Diagrama esquemático do circuito elétrico com 2 resistores em série. </center>
# + [markdown] slideshow={"slide_type": "fragment"}
# São feitas medições de tensão e corrente elétrica sobre o resistor carga $R_C$, variando a fonte de tensão regulável de $0\,V$ até $V_{Smax}$ em 10 passos (variando 10%), depois é invertida a polaridade e repetido o procedimento, com 21 valores ao todo.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### 3.2.2 - Caracterização $V \times I$ de Diodo $D$
# + [markdown] slideshow={"slide_type": "fragment"}
# É substituída a carga $R_C$ pelo diodo $D$ indicado (com catodo, indicado pela faixa, conectado no terminal negativo da fonte de tensão regulável).
# + [markdown] slideshow={"slide_type": "fragment"}
# <center> Figura N - Diagrama esquemático do circuito elétrico com 1 resistor e 1 diodo em série. </center>
# + [markdown] slideshow={"slide_type": "fragment"}
# São feitas medições de tensão e corrente elétrica sobre o diodo $D$, variando a fonte de tensão regulável de $0\,V$ até $V_{Smax}$ em 10 passos (variando 10%), depois é invertida a polaridade e repetido o procedimento porém variando 50%, com 13 valores ao todo.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### 3.2.3 - Caracterização $V \times I$ de LED
# + [markdown] slideshow={"slide_type": "fragment"}
# É substituída a carga $R_C$ pelo LED indicado (com catodo, indicado pelo terminal mais curto, conectado no terminal negativo da fonte de tensão regulável).
# + [markdown] slideshow={"slide_type": "fragment"}
# <center> Figura N - Diagrama esquemático do circuito elétrico com 1 resistor e 1 LED em série. </center>
# + [markdown] slideshow={"slide_type": "fragment"}
# São feitas medições de tensão e corrente elétrica sobre o LED, variando a fonte de tensão regulável de $0\,V$ até $V_{Smax}$ em 10 passos (variando 10%), depois é invertida a polaridade e repetido o procedimento porém variando 50%, com 13 valores ao todo.
# + [markdown] slideshow={"slide_type": "slide"}
# # 4 - Resultados e Discussão
# + [markdown] slideshow={"slide_type": "fragment"}
# < Apresentação dos dados medidos e dos principais resultados do experimento, com incertezas, o número correto de algarismos significativos e em notação científica. É nessa seção que se discute mais amplamente esses resultados apresentados.
#
# Parcialmente baseado em "Procedimento Experimental / Medidas" e "Análises" do roteiro.
#
# Esta seção é o coração do relatório. Nela são apresentados os dados obtidos em forma de fotos, tabelas, gráficos e diagramas. Lembre-se que quando o volume de dados é elevado os gráficos devem ter preferência sobre as tabelas.
#
# Lembre-se que toda medida experimental apresenta uma incerteza e portanto as contas efetuadas devem levar estas em consideração via propagação de incertezas.
#
# Por exemplo, nesse experimento (fiquem à vontade em alterar) : >
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 4.1 - Medindo Tensão, Corrente e Resistência Elétricas com Multímetro
# + [markdown] slideshow={"slide_type": "fragment"}
# 1) Calculando a corrente elétrica $I_R$ teórica para o valor do resistor $R$ escolhido e a tensão $V_S = 3,3\,V$ ou $5,0\,V$ (vide indicaçao do professor) nominal da fonte ("source") de tensão :
#
# $R = \underline{\ \ \ \ \ \ \ \ \ }\,k\Omega$
#
# $V_S = \underline{\ \ \ \ \ \ \ \ \ }\,V$
#
# $I_R = \underline{\ \ \ \ \ \ \ \ \ }\,mA$
#
# Vide 7.1.1 de Anexos.
# + [markdown] slideshow={"slide_type": "fragment"}
# 2) Após consulta do manual do multímetro, a escolha da escala de tensão elétrica contínua (DC) mais adequada do multímetro em modo voltímetro, i. e., a menor escala capaz de ler a tensão $V_S$ :
#
# Escala $= \underline{\ \ \ \ \ \ \ \ \ }\,V$
# + [markdown] slideshow={"slide_type": "fragment"}
# 3) Após consulta do manual do multímetro, a escolha da escala de corrente elétrica contínua (DC) mais adequada do multímetro em modo amperímetro, i. e., a menor escala capaz de ler a corrente $I_R$ :
#
# Escala $= \underline{\ \ \ \ \ \ \ \ \ }\, mA$
# + [markdown] slideshow={"slide_type": "subslide"}
# <center> Figura N - Foto do circuito elétrico com resistor. </center>
# + [markdown] slideshow={"slide_type": "fragment"}
# | n | Leitura da tela | Escala (V) | Resolução (V) | Precisão ±(p%+nD) | $V_R$ (V) com incerteza |
# |---------|-------------------|-------------|---------------|---------------------|--------------------------|
# | 1 | | | | | |
# | 2 | | | | | |
# | 3 | | | | | |
# | 4 | | | | | |
# | 5 | | | | | |
#
# <center>Tabela 1 : Medidas de $V_R$ variando escala do voltímetro.</center>
#
# Vide 7.1.1 de Anexos.
# + [markdown] slideshow={"slide_type": "fragment"}
# Na tabela 1 :
# - as medições de $V_R$ são compatíveis com o valor nominal da fonte de tensão $V_S$, no ítem (1) de "Medidas" ?
# - qual a escala com menor incerteza na medição de $V_R$ ?
# - é a mesma escala apontada no ítem (2) de "Medidas" ?
#
# ...
# + [markdown] slideshow={"slide_type": "subslide"}
# | n | Leitura da tela | Escala (A) | Resolução (A) | Precisão ±(p%+nD) | $I_R$ (mA) com incerteza |
# |---------|-------------------|--------------|----------------|---------------------|-----------------------------|
# | 1 | | | | | |
# | 2 | | | | | |
# | 3 | | | | | |
# | 4 | | | | | |
# | 5 | | | | | |
#
# <center>Tabela 2 : Medidas de $I_R$ variando escala do amperímetro.</center>
#
# Vide 7.1.1 de Anexos.
# + [markdown] slideshow={"slide_type": "fragment"}
# Na tabela 2 :
# - as medições de $I_R$ são compatíveis com o valor teórico calculado da corrente elétrica $I_R$, no ítem (1) de "Medidas" ?
# - qual a escala com menor incerteza na medição de $I_R$ ?
# - é a mesma escala apontada no ítem (3) de "Medidas" ?
#
# ...
# + [markdown] slideshow={"slide_type": "subslide"}
# 9) Após consulta do manual do multímetro, a escolha da escala de resistência elétrica mais adequada do multímetro em modo ohmímetro, i. e., a menor escala capaz de ler a resistência elétrica $R$ :
#
# Escala $= \underline{\ \ \ \ \ \ \ \ \ }\, \Omega$
# + [markdown] slideshow={"slide_type": "fragment"}
# | n | Leitura da tela | Escala ($\Omega$) | Resolução ($\Omega$) | Precisão ±(p%+nD) | $R$ ($k\Omega$) com incerteza |
# |---------|-------------------|--------------|----------------|---------------------|-----------------------------|
# | 1 | | | | | |
# | 2 | | | | | |
# | 3 | | | | | |
# | 4 | | | | | |
# | 5 | | | | | |
#
# <center>Tabela 3 : Medidas de $R$ variando escala do ohmímetro.</center>
#
# Vide 7.1.1 de Anexos.
# + [markdown] slideshow={"slide_type": "subslide"}
# Na tabela 3 :
# - as medições de $R$ são compatíveis com o valor nominal do resistor $R$ obtido via código de cores, no ítem (1) de "Medidas" ?
# - qual a escala com menor incerteza na medição de $R$ ?
# - é a mesma escala apontada no ítem (9) de "Medidas" ou não ?
# - calcule **uma vez** $R$ a partir de $V_R$ e $I_R$ medidos nas escalas mais adequadas, ítens (2)-(3) de "Medidas", usando propagação de incertezas (manualmente e via propagação de incerteza usando [módulo Python Uncertainties](https://pythonhosted.org/uncertainties/), comparando as incertezas resultantes). Compare com a medição de $R$ na melhor escala e com o valor nominal do resistor $R$.
#
# ...
#
# Vide 7.1.2 de Anexos.
#
# ...
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 4.2 - Caracterização de Resistor, Diodo e LED com Multímetro
# + [markdown] slideshow={"slide_type": "fragment"}
# As medições são feitas pelo professor, pois a fonte de tensão regulável é equipamento mais complexo e caro e por isso não está presente no kit de material fornecido aos grupos.
# + [markdown] slideshow={"slide_type": "fragment"}
# ### 4.2.1 - Caracterização $V \times I$ de Resistor $R_C$
# + [markdown] slideshow={"slide_type": "fragment"}
# 1) Calculando a corrente elétrica $I_{max}$ teórica máxima para os valores dos resistores $R_L$ e $R_C$ escolhidos e a tensão $V_{Smax}$ máxima nominal da fonte ("source") de tensão regulável :
#
# $R_L = \underline{\ \ \ \ \ \ \ \ \ }\,k\Omega$
#
# $R_C = \underline{\ \ \ \ \ \ \ \ \ }\,k\Omega$
#
# $V_{Smax} = \underline{\ \ \ \ \ \ \ \ \ }\,V$
#
# $I_{max} = \underline{\ \ \ \ \ \ \ \ \ }\,mA$
#
# Vide 7.2.1.1 de Anexos.
# + [markdown] slideshow={"slide_type": "subslide"}
# 2) Após consideração do divisor de tensão com $R_L$ e $R_C$ e consulta do manual do multímetro, a escolha da escala de tensão elétrica contínua (DC) mais adequada do multímetro em modo voltímetro, i. e., a menor escala capaz de ler a tensão $V_{R_Cmax}$ :
#
# Escala $= \underline{\ \ \ \ \ \ \ \ \ }\,V$
#
# Vide 7.2.1.1 de Anexos.
# + [markdown] slideshow={"slide_type": "fragment"}
# 3) Após consulta do manual do multímetro, a escolha da escala de corrente elétrica contínua (DC) mais adequada do multímetro em modo amperímetro, i. e., a menor escala capaz de ler a corrente $I_{max}$ :
#
# Escala $= \underline{\ \ \ \ \ \ \ \ \ }\, mA$
# + [markdown] slideshow={"slide_type": "fragment"}
# <center> Figura N - Foto do circuito elétrico com 2 resistores em série. </center>
# + [markdown] slideshow={"slide_type": "subslide"}
# | n | $V_S\,(V)$ | Leitura da tela | Escala (V) | Resolução (V) | Precisão ±(p%+nD) | $V_{R_C}$ (V) com incerteza |
# |----|------------|-------------------|-------------|---------------|---------------------|----------------------------------|
# | 1 | | | | | | |
# | 2 | | | | | | |
# | 3 | | | | | | |
# | 4 | | | | | | |
# | 5 | | | | | | |
# | 6 | | | | | | |
# | 7 | | | | | | |
# | 8 | | | | | | |
# | 9 | | | | | | |
# | 10 | | | | | | |
# | 11 | | | | | | |
# | 12 | | | | | | |
# | 13 | | | | | | |
# | 14 | | | | | | |
# | 15 | | | | | | |
# | 16 | | | | | | |
# | 17 | | | | | | |
# | 18 | | | | | | |
# | 19 | | | | | | |
# | 20 | | | | | | |
# | 21 | | | | | | |
#
# <center>Tabela 4 : Medidas de $V_{R_C}$ variando $V_S$, cujos dados são repassados para cada grupo.</center>
#
# Vide 7.2.1.1 de Anexos.
# + [markdown] slideshow={"slide_type": "subslide"}
# | n | $V_S\,(V)$ | Leitura da tela | Escala (A) | Resolução (A) | Precisão ±(p%+nD) | $I_{R_C}$ (V) com incerteza |
# |----|------------|-------------------|-------------|---------------|---------------------|----------------------------------|
# | 1 | | | | | | |
# | 2 | | | | | | |
# | 3 | | | | | | |
# | 4 | | | | | | |
# | 5 | | | | | | |
# | 6 | | | | | | |
# | 7 | | | | | | |
# | 8 | | | | | | |
# | 9 | | | | | | |
# | 10 | | | | | | |
# | 11 | | | | | | |
# | 12 | | | | | | |
# | 13 | | | | | | |
# | 14 | | | | | | |
# | 15 | | | | | | |
# | 16 | | | | | | |
# | 17 | | | | | | |
# | 18 | | | | | | |
# | 19 | | | | | | |
# | 20 | | | | | | |
# | 21 | | | | | | |
#
# <center>Tabela 5 : Medidas de $I_{R_C}$ variando $V_S$, cujos dados são repassados para cada grupo.</center>
#
# Vide 7.2.1.1 de Anexos.
# + [markdown] slideshow={"slide_type": "subslide"}
# Gráfico de $V_{R_C} \times I_{R_C}$ (ou seja, $I_{R_C}$ no eixo horizontal), usando SciDAVis (ou AlphaPlot, ou Python), com barras de incerteza e cálculo de ajuste de curva :
# + [markdown] slideshow={"slide_type": "fragment"}
# <center> Figura N - gráfico de $V_{R_C} \times I_{R_C}$. </center>
# + [markdown] slideshow={"slide_type": "subslide"}
# A curva é linear ou não-linear ?
#
# R.:
#
# Obtenha o coeficiente angular do ajuste linear e compare com o valor nominal de $R_C$.
#
# R.:
#
# Vide 7.2.2.1 de Anexos.
#
# Ou seja, analisando graficamente, o resistor $R_C$ segue a lei de Ohm ?
#
# R.:
# + [markdown] slideshow={"slide_type": "fragment"}
# Ao inverter a polaridade da fonte de tensão $V_S$, o que ocorre com as medidas elétricas do resistor $R_C$ ?
#
# R.:
#
# Então resistor tem polaridade, só conduzindo em um sentido ?
#
# R.:
# + [markdown] slideshow={"slide_type": "fragment"}
# A partir dos valores de $V_{R_C}$ da tabela 4 e $I_{R_C}$ da tabela 5, é calculada a potência $P_{R_C} = V_{R_C} I_{R_C}$ com propagação de incertezas usando módulo [Python Uncertainties](https://pythonhosted.org/uncertainties/) com [NumPy](https://pythonhosted.org/uncertainties/numpy_guide.html), criando a tabela 10 :
# + [markdown] slideshow={"slide_type": "subslide"}
# | n | $V_{R_C}\,(V)$ com incerteza | $I_{R_C}\,(V)$ com incerteza | $P_{R_C}\,(V)$ com incerteza |
# |:--:|:----------------------------:|:-----------------------------:|:----------------------------:|
# | 1 | | | |
# | 2 | | | |
# | 3 | | | |
# | 4 | | | |
# | 5 | | | |
# | 6 | | | |
# | 7 | | | |
# | 8 | | | |
# | 9 | | | |
# | 10 | | | |
# | 11 | | | |
# | 12 | | | |
# | 13 | | | |
# | 14 | | | |
# | 15 | | | |
# | 16 | | | |
# | 17 | | | |
# | 18 | | | |
# | 19 | | | |
# | 20 | | | |
# | 21 | | | |
#
# <center>Tabela 10 : $P_{R_C}$ variando $V_S$.</center>
#
# Vide 7.2.2.1 de Anexos.
# + [markdown] slideshow={"slide_type": "subslide"}
# Gráfico de $P_{R_C} \times I_{R_C}$ (ou seja, $I_{R_C}$ no eixo horizontal), usando SciDAVis (ou AlphaPlot, ou Python), com barras de incerteza :
# + [markdown] slideshow={"slide_type": "fragment"}
# <center> Figura N - gráfico de $P_{R_C} \times I_{R_C}$. </center>
# + [markdown] slideshow={"slide_type": "subslide"}
# A curva é linear ou não-linear, de que tipo ?
#
# R.:
#
# Compare com o comportamento previsto teoricamente de $P_{R_C}$ em termos de $R_C$ e $I_{R_C}$.
#
# R.:
# + [markdown] slideshow={"slide_type": "fragment"}
# ... Discussão ...
# + [markdown] slideshow={"slide_type": "subslide"}
# ### 4.2.2 - Caracterização $V \times I$ de Diodo $D$
# + [markdown] slideshow={"slide_type": "fragment"}
# 1) Calculando a corrente elétrica $I_{max}$ teórica máxima considerando a queda de tensão $V_{Dmax}$, os valores do resistor $R_L$ escolhido e a tensão $V_{Smax}$ máxima nominal da fonte ("source") de tensão regulável :
#
# $R_L = \underline{\ \ \ \ \ \ \ \ \ }\,k\Omega$
#
# $V_{Dmax} = \underline{\ \ \ \ \ \ \ \ \ }\,V$
#
# $V_{Smax} = \underline{\ \ \ \ \ \ \ \ \ }\,V$
#
# $I_{max} = \underline{\ \ \ \ \ \ \ \ \ }\,mA$
#
# Vide 7.2.1.2 de Anexos.
# + [markdown] slideshow={"slide_type": "fragment"}
# 2) Após consulta do manual do multímetro, a escolha da escala de tensão elétrica contínua (DC) mais adequada do multímetro em modo voltímetro, i. e., a menor escala capaz de ler a tensão $V_{Dmax}$ :
#
# Escala $= \underline{\ \ \ \ \ \ \ \ \ }\,V$
# + [markdown] slideshow={"slide_type": "subslide"}
# 3) Após consideração da queda de tensão $V_{Dmax}$ e consultando o manual do multímetro, a escolha da escala de corrente elétrica contínua (DC) mais adequada do multímetro em modo amperímetro, i. e., a menor escala capaz de ler a corrente $I_{max}$ :
#
# Escala $= \underline{\ \ \ \ \ \ \ \ \ }\, mA$
# + [markdown] slideshow={"slide_type": "fragment"}
# <center> Figura N - Foto do circuito elétrico com 1 resistor e 1 diodo em série. </center>
# + [markdown] slideshow={"slide_type": "subslide"}
# | n | $V_S\,(V)$ | Leitura da tela | Escala (V) | Resolução (V) | Precisão ±(p%+nD) | $V_{D}$ (V) com incerteza |
# |----|------------|-------------------|-------------|---------------|---------------------|----------------------------------|
# | 1 | | | | | | |
# | 2 | | | | | | |
# | 3 | | | | | | |
# | 4 | | | | | | |
# | 5 | | | | | | |
# | 6 | | | | | | |
# | 7 | | | | | | |
# | 8 | | | | | | |
# | 9 | | | | | | |
# | 10 | | | | | | |
# | 11 | | | | | | |
# | 12 | | | | | | |
# | 13 | | | | | | |
#
# <center>Tabela 6 : Medidas de $V_{D}$ variando $V_S$, cujos dados são repassados para cada grupo.</center>
#
# Vide 7.2.1.2 de Anexos.
# + [markdown] slideshow={"slide_type": "subslide"}
# | n | $V_S\,(V)$ | Leitura da tela | Escala (A) | Resolução (A) | Precisão ±(p%+nD) | $I_{D}$ (V) com incerteza |
# |----|------------|-------------------|-------------|---------------|---------------------|----------------------------------|
# | 1 | | | | | | |
# | 2 | | | | | | |
# | 3 | | | | | | |
# | 4 | | | | | | |
# | 5 | | | | | | |
# | 6 | | | | | | |
# | 7 | | | | | | |
# | 8 | | | | | | |
# | 9 | | | | | | |
# | 10 | | | | | | |
# | 11 | | | | | | |
# | 12 | | | | | | |
# | 13 | | | | | | |
#
# <center>Tabela 7 : Medidas de $I_{D}$ variando $V_S$, cujos dados são repassados para cada grupo.</center>
#
# Vide 7.2.1.2 de Anexos.
# + [markdown] slideshow={"slide_type": "subslide"}
# Gráfico de $V_{D} \times I_{D}$ (ou seja, $I_{D}$ no eixo horizontal) usando só as medições com $V_S$ não-negativas, usando SciDAVis (ou AlphaPlot, ou Python), com barras de incerteza :
# + [markdown] slideshow={"slide_type": "fragment"}
# <center> Figura N - gráfico de $V_{D} \times I_{D}$. </center>
# + [markdown] slideshow={"slide_type": "subslide"}
# A curva é linear ou não-linear ?
#
# R.:
#
# Compare a tensão direta $V_{Dmax}$ máxima nominal com o valor de tensão tendendo a linha horizontal no gráfico.
#
# R.:
# + [markdown] slideshow={"slide_type": "fragment"}
# Ao inverter a polaridade da fonte de tensão $V_S$, o que ocorre com as medidas elétricas do diodo ?
#
# R.:
#
# Então diodo tem polaridade, só conduzindo em um sentido ?
#
# R.:
# + [markdown] slideshow={"slide_type": "subslide"}
# Usando software de simulação de circuitos eletrônicos para o circuito com fonte de tensão regulável $V_S$, resistor limitador $R_L$ e diodo $D$, variando $V_S$ e medindo $V_{D}$ e $I_{D}$ sem incertezas via software.
#
# Obtendo assim outro gráfico de $V_{D} \times I_{D}$ (ou seja, $I_{D}$ no eixo horizontal) análogo, porém sem barras de incerteza. Compare com o gráfico feito a partir de medidas :
# + [markdown] slideshow={"slide_type": "fragment"}
# <center> Figura N - gráfico de $V_{D} \times I_{D}$ via simulação. </center>
# + [markdown] slideshow={"slide_type": "fragment"}
# ... Discussão ...
# + [markdown] slideshow={"slide_type": "subslide"}
# ### 4.2.3 - Caracterização $V \times I$ de LED
# + [markdown] slideshow={"slide_type": "fragment"}
# 1) Calculando a corrente elétrica $I_{max}$ teórica máxima considerando a queda de tensão $V_{LEDmax}$, os valores do resistor $R_L$ escolhido e a tensão $V_{Smax}$ máxima nominal da fonte ("source") de tensão regulável :
#
# $R_L = \underline{\ \ \ \ \ \ \ \ \ }\,k\Omega$
#
# $V_{LEDmax} = \underline{\ \ \ \ \ \ \ \ \ }\,V$
#
# $V_{Smax} = \underline{\ \ \ \ \ \ \ \ \ }\,V$
#
# $I_{max} = \underline{\ \ \ \ \ \ \ \ \ }\,mA$
#
# Vide 7.2.1.3 de Anexos.
# + [markdown] slideshow={"slide_type": "fragment"}
# 2) Após consulta do manual do multímetro, a escolha da escala de tensão elétrica contínua (DC) mais adequada do multímetro em modo voltímetro, i. e., a menor escala capaz de ler a tensão $V_{LEDmax}$ :
#
# Escala $= \underline{\ \ \ \ \ \ \ \ \ }\,V$
# + [markdown] slideshow={"slide_type": "subslide"}
# 3) Após consideração da queda de tensão $V_{LEDmax}$ e consultando o manual do multímetro, a escolha da escala de corrente elétrica contínua (DC) mais adequada do multímetro em modo amperímetro, i. e., a menor escala capaz de ler a corrente $I_{max}$ :
#
# Escala $= \underline{\ \ \ \ \ \ \ \ \ }\, mA$
# + [markdown] slideshow={"slide_type": "fragment"}
# <center> Figura N - Foto do circuito elétrico com 1 resistor e 1 LED em série. </center>
# + [markdown] slideshow={"slide_type": "subslide"}
# | n | $V_S\,(V)$ | Leitura da tela | Escala (V) | Resolução (V) | Precisão ±(p%+nD) | $V_{LED}$ (V) com incerteza |
# |----|------------|-------------------|-------------|---------------|---------------------|----------------------------------|
# | 1 | | | | | | |
# | 2 | | | | | | |
# | 3 | | | | | | |
# | 4 | | | | | | |
# | 5 | | | | | | |
# | 6 | | | | | | |
# | 7 | | | | | | |
# | 8 | | | | | | |
# | 9 | | | | | | |
# | 10 | | | | | | |
# | 11 | | | | | | |
# | 12 | | | | | | |
# | 13 | | | | | | |
#
# <center> Tabela 8 : Medidas de $V_{LED}$ variando $V_S$, cujos dados serão repassados para cada grupo. </center>
#
# Vide 7.2.1.3 de Anexos.
# + [markdown] slideshow={"slide_type": "subslide"}
# | n | $V_S\,(V)$ | Leitura da tela | Escala (A) | Resolução (A) | Precisão ±(p%+nD) | $I_{LED}$ (V) com incerteza |
# |----|------------|-------------------|-------------|---------------|---------------------|----------------------------------|
# | 1 | | | | | | |
# | 2 | | | | | | |
# | 3 | | | | | | |
# | 4 | | | | | | |
# | 5 | | | | | | |
# | 6 | | | | | | |
# | 7 | | | | | | |
# | 8 | | | | | | |
# | 9 | | | | | | |
# | 10 | | | | | | |
# | 11 | | | | | | |
# | 12 | | | | | | |
# | 13 | | | | | | |
#
# <center> Tabela 9 : Medidas de $I_{LED}$ variando $V_S$, cujos dados são repassados para cada grupo. </center>
#
# Vide 7.2.1.3 de Anexos.
# + [markdown] slideshow={"slide_type": "subslide"}
# Gráfico de $V_{LED} \times I_{LED}$ (ou seja, $I_{LED}$ no eixo horizontal) usando só as medições com $V_S$ não-negativas, usando SciDAVis (ou AlphaPlot, ou Python), com barras de incerteza.
# + [markdown] slideshow={"slide_type": "fragment"}
# <center> Figura N - gráfico de $V_{LED} \times I_{LED}$. </center>
# + [markdown] slideshow={"slide_type": "subslide"}
# A curva é linear ou não-linear ?
#
# R.:
#
# Compare a tensão direta $V_{LEDmax}$ máxima nominal com o valor de tensão tendendo a linha horizontal no gráfico.
#
# R.:
# + [markdown] slideshow={"slide_type": "fragment"}
# Ao inverter a polaridade da fonte de tensão $V_S$, o que ocorre com as medidas elétricas do LED ?
#
# R.:
#
# Então LED tem polaridade, só conduzindo em um sentido ?
#
# R.:
# + [markdown] slideshow={"slide_type": "fragment"}
# ... Discussão ...
# + [markdown] slideshow={"slide_type": "slide"}
# # 5 - Conclusão
# + [markdown] slideshow={"slide_type": "fragment"}
# <A conclusão deve abordar brevemente o experimento efetuado, os resultados obtidos e a que conclusões estes resultados levam. Em alguns casos se discute possíveis rumos desta investigação.
#
# Apresentar com clareza se o objetivo foi cumprido ou dar uma explicação satisfatória para o não cumprimento do objetivo.
#
# Comentários do tipo: “O experimento foi muito proveitoso....” “Foi muito legal ter feito isso...” e outro similares não tem sentido científico e portanto não devem ser utilizados.>
# + [markdown] slideshow={"slide_type": "fragment"}
# ...
# + [markdown] slideshow={"slide_type": "slide"}
# # 6 - Referências Bibliográficas
# + [markdown] slideshow={"slide_type": "fragment"}
# < Devem constar nessa seção todas as referências bibliográfias, seguindo norma da ABNT, utilizadas no relatório, como :
# - livros impressos;
# - livros eletrônicos;
# - sítios com material de referência;
# - softwares;
# - etc. >
# + [markdown] slideshow={"slide_type": "fragment"}
# [HRW] <NAME>.; <NAME>.; <NAME>.; Fundamentos de Física: Eletromagnetismo. Vol. 3,
# 10a ed, editora LTC, 2016. ISBN: 8521630379;
# + [markdown] slideshow={"slide_type": "fragment"}
# ...
# + [markdown] slideshow={"slide_type": "slide"}
# # 7 - Anexos
# + [markdown] slideshow={"slide_type": "fragment"}
# < Devem constar nessa seção todos os cálculos intermediários feitos no experimento e relatório, como :
# - cálculo de incerteza das medições;
# - cálculos de propagação de incertezas;
# - relatório de ajuste de curvas gerado por certo software (SciDAVis, etc);
# - cálculos diversos.
#
# Os cálculos podem ser organizados em subseções, subsubseçõe, etc. >
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 7.1 - Medindo Tensão, Corrente e Resistência Elétricas com Multímetro
# + [markdown] slideshow={"slide_type": "fragment"}
# ### 7.1.1 - Medidas
# + [markdown] slideshow={"slide_type": "fragment"}
# $R$ indicado pelo professor via código de cores : cor1 cor2 cor3 cor4.
# Consultando tabela de código de cores [ResCores], o valor é $R = (N k\,\Omega \pm p\%)$.
# + [markdown] slideshow={"slide_type": "fragment"}
# Corrente elétrica teórica, sem incertezas :
#
# $$I_R =\,...$$
# + [markdown] slideshow={"slide_type": "subslide"}
# Cálculos de incertezas das medições da Tabela 1 :
#
# ...
# + [markdown] slideshow={"slide_type": "subslide"}
# Cálculos de incertezas das medições da Tabela 2 :
#
# ...
# + [markdown] slideshow={"slide_type": "subslide"}
# Cálculos de incertezas das medições da Tabela 3 :
#
# ...
# + [markdown] slideshow={"slide_type": "subslide"}
# ### 7.1.2 - Análises
# + [markdown] slideshow={"slide_type": "fragment"}
# Análise da Tabela 3 :
#
# $R$ a partir de $V_R$ e $I_R$ usando propagação de incertezas manualmente :
#
# ...
#
# $R$ via propagação de incerteza usando [módulo Python Uncertainties](https://pythonhosted.org/uncertainties/) :
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 7.2 - Caracterização de Resistor, Diodo e LED com Multímetro
# + [markdown] slideshow={"slide_type": "fragment"}
# ### 7.2.1 - Medidas
# + [markdown] slideshow={"slide_type": "fragment"}
# #### 7.2.1.1 - Caracterização $V \times I$ de Resistor $R_C$
# + [markdown] slideshow={"slide_type": "fragment"}
# $R_L$ indicado pelo professor via código de cores : cor1 cor2 cor3 cor4.
# Consultando tabela de código de cores [ResCores], o valor é $R_L = (N k\,\Omega \pm p\%)$.
# + [markdown] slideshow={"slide_type": "fragment"}
# $R_C$ indicado pelo professor via código de cores : cor1 cor2 cor3 cor4.
# Consultando tabela de código de cores [ResCores], o valor é $R_L = (N k\,\Omega \pm p\%)$.
# + [markdown] slideshow={"slide_type": "subslide"}
# Cálculo de $I_{max}$ teórica, em termos de $V_{Smax}$, $R_L$ e $R_C$, sem incertezas :
#
# $$I_{max} = \,...$$
# + [markdown] slideshow={"slide_type": "fragment"}
# Cálculo de $V_{R_Cmax}$ teórica, em termos de $V_{Smax}$, $R_L$ e $R_C$, sem incertezas :
#
# $$V_{R_Cmax} =\,...$$
# + [markdown] slideshow={"slide_type": "subslide"}
# Cálculos de incertezas das medições da Tabela 4 :
#
# ...
# + [markdown] slideshow={"slide_type": "subslide"}
# Cálculos de incertezas das medições da Tabela 5 :
#
# ...
# + [markdown] slideshow={"slide_type": "subslide"}
# #### 7.2.1.2 - Caracterização $V \times I$ de Diodo $D$
# + [markdown] slideshow={"slide_type": "fragment"}
# Cálculo de $I_{max}$ teórica, em termos de $V_{Smax}$, $R_L$ e $V_{Dmax}$, sem incertezas :
#
# $$I_{max} = \,...$$
# + [markdown] slideshow={"slide_type": "subslide"}
# Cálculos de incertezas das medições da Tabela 6 :
#
# ...
# + [markdown] slideshow={"slide_type": "subslide"}
# Cálculos de incertezas das medições da Tabela 7 :
#
# ...
# + [markdown] slideshow={"slide_type": "subslide"}
# #### 7.2.1.3 - Caracterização $V \times I$ de LED
# + [markdown] slideshow={"slide_type": "fragment"}
# Cálculo de $I_{max}$ teórica, em termos de $V_{Smax}$, $R_L$ e $V_{LEDmax}$, sem incertezas :
#
# $$I_{max} = \,...$$
# + [markdown] slideshow={"slide_type": "subslide"}
# Cálculos de incertezas das medições da Tabela 8 :
#
# ...
# + [markdown] slideshow={"slide_type": "subslide"}
# Cálculos de incertezas das medições da Tabela 9 :
#
# ...
# + [markdown] slideshow={"slide_type": "subslide"}
# ### 7.2.2 - Análises
# + [markdown] slideshow={"slide_type": "fragment"}
# #### 7.2.2.1 - Caracterização $V \times I$ de Resistor $R_C$
# + [markdown] slideshow={"slide_type": "fragment"}
# Relatório de ajuste de curvas do gráfico de $V_{R_C} \times I_{R_C}$, gerado por certo software (SciDAVis, etc) :
#
#
# ...
# + [markdown] slideshow={"slide_type": "subslide"}
# Cálculo de propagação de incertezas da tabela 10, usando módulo [Python Uncertainties](https://pythonhosted.org/uncertainties/) com [NumPy](https://pythonhosted.org/uncertainties/numpy_guide.html) :
#
# ...
| Experimentos/1_Caracterizacao_Dispositivos_Eletronicos_via_Multimetros/Modelo_Relatorio_Caracterizacao_Dispositivos_Eletronicos_via_Multimetros.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pylab as plt
import warnings
warnings.simplefilter('ignore')
with open('Все идет по плану.txt', 'r', encoding='utf-8') as file:
file = file.read()
# +
from nltk import word_tokenize
from collections import Counter
from wordcloud import WordCloud, STOPWORDS
import pymorphy2
morph = pymorphy2.MorphAnalyzer()
a = ord('а')
russian = ''.join([chr(i) for i in range(a, a+32)])
words = word_tokenize(file)
russian_words = []
for word in words:
flag = True
for char in word:
if char not in russian:
flag = False
break
if flag:
russian_words.append(word)
russian_words = [word for word in russian_words
if ('NOUN' in morph.parse(word)[0].tag) or
('ADJF' in morph.parse(word)[0].tag)]
cloud = Counter(russian_words)
stopwords = set(STOPWORDS)
cloud = ' '.join(cloud)
wordcloud = WordCloud(width = 1000, height = 1000,
stopwords = stopwords,
min_font_size = 8,background_color='white'
).generate(cloud)
plt.figure(figsize = (14, 14))
plt.imshow(wordcloud)
pass
# +
import random
import importlib as imp
from algorithm import naive
from algorithm import rabin_karp
from algorithm import boyer_moore_horspool
from algorithm import knuth_morris_pratt
from utils import tools
tools = imp.reload(tools)
rabin_karp = imp.reload(rabin_karp)
naive = imp.reload(naive)
boyer_moore_horspool = imp.reload(boyer_moore_horspool)
knuth_morris_pratt = imp.reload(knuth_morris_pratt)
len_candidate = 3
len_reference = 150
dictionary = ['a', 'b', 'c', 'd']
reference = tools.gen_random_string(dictionary, len_reference)
candidate = tools.gen_random_string(dictionary, len_candidate)
print('Reference: {}'.format(reference))
print('Candidate: {}'.format(candidate))
alg = knuth_morris_pratt.KnuthMorrisPratt(reference)
# alg = boyer_moore_horspool.BoyerMooreHorspool(reference)
# alg = rabin_karp.RabinKarp(reference)
# alg = naive.BruteForce(reference)
alg.set_candidate(candidate)
offset_lst = alg.search(multiple_search=True)
print('Indexes: {}'.format(offset_lst))
reference = list(reference)
i = 0
for offset in offset_lst:
ofst = offset + i
reference.insert(ofst, '_')
for i_char in range(len_candidate):
reference[ofst + i_char + 1] = reference[ofst + i_char + 1].upper()
reference.insert(ofst + len_candidate + 1, '_')
i += 2
reference = ''.join(reference)
print('Result: {}'.format(reference))
# +
# %%time
from utils import tools
tools = imp.reload(tools)
algorithms = [naive.BruteForce,
rabin_karp.RabinKarp,
boyer_moore_horspool.BoyerMooreHorspool,
knuth_morris_pratt.KnuthMorrisPratt
]
set_params = [{},
{'d': 27, 'q': 1247},
{},
{}
]
reference_len = np.arange(1000, 50000, 5000).tolist()
dictionary = ['a', 'b', 'c', 'd', 'l', 'r']
n_observations = 3
coef = 0.42
info_df = tools.generate_stat( algorithms=algorithms,
set_params=set_params,
gen_string=tools.gen_random_string,
dictionary=dictionary,
reference_len=reference_len,
candidate_len=[round(rlen * coef) for rlen in reference_len],
n_observations=n_observations,
multiple_search=False)
# -
stat_df = info_df.groupby(['algorithm', 'reference_len']).agg({'execution':['mean','std']})
stat_df.columns = ['execution mean', 'execution std']
stat_df = stat_df.reset_index()
tools = imp.reload(tools)
tools.get_plots(stat_df, figsize=(15, 8))
import os
PATH_TO_BENCHMARKS = './benchmarks/'
all_files = os.listdir(PATH_TO_BENCHMARKS)
# +
names = ['bad_w', 'bad_t', 'good_w', 'good_t']
info_file = {key:[] for key in names}
for n in names:
for l in all_files:
if (l.startswith(n)):
info_file[n].append(l)
# -
info_file
# +
tools = imp.reload(tools)
algorithms = [naive.BruteForce,
rabin_karp.RabinKarp,
boyer_moore_horspool.BoyerMooreHorspool,
knuth_morris_pratt.KnuthMorrisPratt]
set_params = [{}, {'d': 59, 'q': 13}, {},{}]
info_df_good_benchmarks = tools.generate_stat_for_benchmarks( algorithms=algorithms,
set_params=set_params,
files_w = info_file['good_w'],
files_t = info_file['good_t'],
path_to_benchmarks = PATH_TO_BENCHMARKS,
n_observations=5,
multiple_search=False
)
# -
stat_df_benchmarks = info_df_good_benchmarks.groupby(['filename', 'algorithm']).agg({'execution':['mean','std'],
'n_operations': 'mean',
'reference_len': 'last'})
stat_df_benchmarks.columns = [ 'execution mean', 'execution std', 'operations', 'reference_len']
print(np.around(stat_df_benchmarks[[ 'execution mean', 'execution std', 'operations']], 4).reset_index().to_markdown())
tools = imp.reload(tools)
tools.get_plots(stat_df_benchmarks.reset_index(), figsize=(15, 8))
# +
tools = imp.reload(tools)
algorithms = [naive.BruteForce,
rabin_karp.RabinKarp,
boyer_moore_horspool.BoyerMooreHorspool,
knuth_morris_pratt.KnuthMorrisPratt
]
set_params = [ {},
{'d': 2117, 'q': 12},
{},
{}
]
info_df_bad_benchmarks = tools.generate_stat_for_benchmarks(algorithms=algorithms,
set_params=set_params,
files_w = info_file['bad_w'],
files_t = info_file['bad_t'],
path_to_benchmarks = PATH_TO_BENCHMARKS,
n_observations=5,
multiple_search=False
)
# -
stat_df_benchmarks = info_df_bad_benchmarks.groupby(['filename', 'algorithm']).agg({'execution':['mean','std'],
'n_operations': 'mean',
'reference_len': 'last'})
stat_df_benchmarks.columns = [ 'execution mean', 'execution std', 'operations', 'reference_len']
print(np.around(stat_df_benchmarks[[ 'execution mean', 'execution std', 'operations']], 4).reset_index().to_markdown())
tools = imp.reload(tools)
tools.get_plots(stat_df_benchmarks.reset_index(), figsize=(15, 8))
# ---
#
# # Лабораторная работа №1. Поиск подстроки в строке
#
# ---
# <NAME><br>
# <NAME><br>
# 17ПМИ<br>
#
# ---
#
# ### *`goodXXX.txt`* benchmarks<br>
# *Количество наблюдений на каждый алгоритм - 5*<br>
# *время представлено в секунда с точность до **4х** знаков после запятой*
#
# | | Файл | Алгоритм | Время (среднее), сек. | Время (отклонение), сек. | Операций |
# |---:|:-------------|:---------------------|-----------------:|----------------:|-------------:|
# | 0 | good_t_1.txt | Boyer-Moore-Horspool | 0.0002 | 0.0004 | 88 |
# | 1 | good_t_1.txt | Brute Force | 0.0002 | 0.0004 | 34 |
# | 2 | good_t_1.txt | Knuth-Morris-Pratt | 0.0007 | 0.0007 | 1233 |
# | 3 | good_t_1.txt | Rabin-Karp | 0.0012 | 0.0004 | 674 |
# | 4 | good_t_2.txt | Boyer-Moore-Horspool | 0.0002 | 0.0004 | 124 |
# | 5 | good_t_2.txt | Brute Force | 0.0002 | 0.0004 | 85 |
# | 6 | good_t_2.txt | Knuth-Morris-Pratt | 0.0006 | 0.0005 | 1390 |
# | 7 | good_t_2.txt | Rabin-Karp | 0.001 | 0.0007 | 733 |
# | 8 | good_t_3.txt | Boyer-Moore-Horspool | 0.0004 | 0.0005 | 487 |
# | 9 | good_t_3.txt | Brute Force | 0.0004 | 0.0005 | 445 |
# | 10 | good_t_3.txt | Knuth-Morris-Pratt | 0.0025 | 0.0015 | 4025 |
# | 11 | good_t_3.txt | Rabin-Karp | 0.0035 | 0.0019 | 2169 |
# | 12 | good_t_4.txt | Boyer-Moore-Horspool | 0.0006 | 0.0005 | 551 |
# | 13 | good_t_4.txt | Brute Force | 0.0038 | 0.0021 | 92 |
# | 14 | good_t_4.txt | Knuth-Morris-Pratt | 0.0084 | 0.0006 | 19228 |
# | 15 | good_t_4.txt | Rabin-Karp | 0.0165 | 0.002 | 10318 |
#
#
# 
#
# ---
#
# ### *`badXXX.txt`* benchmarks<br>
# *Количество наблюдений на каждый алгоритм - 5*<br>
# *время представлено в секунда с точность до **4х** знаков после запятой*
#
#
#
# | | Файл | Алгоритм | Время (среднее), сек. | Время (отклонение), сек. | Операций |
# |---:|:------------|:---------------------|-----------------:|----------------:|-------------:|
# | 0 | bad_t_1.txt | Boyer-Moore-Horspool | 0 | 0 | 11 |
# | 1 | bad_t_1.txt | Brute Force | 0 | 0 | 10 |
# | 2 | bad_t_1.txt | Knuth-Morris-Pratt | 0 | 0 | 20 |
# | 3 | bad_t_1.txt | Rabin-Karp | 0 | 0 | 11 |
# | 4 | bad_t_2.txt | Boyer-Moore-Horspool | 0 | 0 | 101 |
# | 5 | bad_t_2.txt | Brute Force | 0.0003 | 0.0004 | 820 |
# | 6 | bad_t_2.txt | Knuth-Morris-Pratt | 0.0002 | 0.0004 | 208 |
# | 7 | bad_t_2.txt | Rabin-Karp | 0.0002 | 0.0004 | 101 |
# | 8 | bad_t_3.txt | Boyer-Moore-Horspool | 0.0008 | 0.0008 | 1001 |
# | 9 | bad_t_3.txt | Brute Force | 0.0435 | 0.0142 | 89200 |
# | 10 | bad_t_3.txt | Knuth-Morris-Pratt | 0.0008 | 0.0004 | 2098 |
# | 11 | bad_t_3.txt | Rabin-Karp | 0.0016 | 0.0005 | 1001 |
# | 12 | bad_t_4.txt | Boyer-Moore-Horspool | 0.0033 | 0.001 | 5001 |
# | 13 | bad_t_4.txt | Brute Force | 1.8613 | 0.255 | 3997000 |
# | 14 | bad_t_4.txt | Knuth-Morris-Pratt | 0.0058 | 0.0018 | 10998 |
# | 15 | bad_t_4.txt | Rabin-Karp | 0.0067 | 0.0013 | 5001 |
#
#
# 
#
# ---
| StringSearching.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Vienna 4x22 note aligned ground truth data
#
# The Vienna 4x22 is a small dataset of note aligned scores and performances. If you're interested, you can find the original data here (MIDI performances and audio): https://repo.mdw.ac.at/projects/IWK/the_vienna_4x22_piano_corpus/data/index.html
#
# On moodle you find a zipped version of .match files of this data. Recall that .match files contain both a score (mostly) and performance as well as a notewise alignment. You can load them like so:
import partitura as pt
import os
path_to_dummy_match = os.path.join("..","introduction","example_data","Chopin_op10_no3_p01.match")
performedpart, alignment_groundtruth, part = pt.load_match(path_to_dummy_match, create_part=True)
alignment_groundtruth
# 1. Load the test/train data as above
# 2. use performedpart (e.g. performaedpart.note_array) and pert (e.g. via part.note_array) to generate a notewise alignment
# 3. store the predicted alignment in the same format as the alignment ground truth (a list of dictionaries with the keys "label", "score_id", and "performance_id" where the label can be "match", "insertion" or "deletion and the IDs are the corresponding note IDs from the performedpart and part)
# 4. compare your solution to the ground truth using the helper function we wrote. Ideally your solutions reaches precision 1.0, recall, 1.0 and f score 1.0
from compare_alignments import compare_alignments_
compare_alignments_(alignment_groundtruth, alignment_groundtruth, "match")
| challenge/Vienna4x22_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import os
import imp
#By <NAME>
# -
lumAPI = imp.load_source("lumAPI","C:\\Program Files\\Lumerical\\v212\\api\\python\\lumapi.py")
fdtd = lumAPI.FDTD()
# +
# Xsize = 240e-9
Ysize = 0.6e-6
Zsize = 0.36e-6
Xcenter=0
Ycenter=0
Zcenter=0
n_start=1e-6 #size of first layer
n_end=1e-6 #size of final layer
n_start_index=1
n_end_index=1
n1=4
n2=1.67
m=1 #Odd multiples 1,3,5...
WL0=1550e-9
d1=m*((WL0/(4*n1)))
d2=m*((WL0/(4*n2)))
N=10
fdtdSpan=(N*(d1+d2))+(n_end)+(n_start)
WaveL_Start = 0.5e-6
WaveL_Stop = 2.5e-6
# +
fdtd.addfdtd()
fdtd.set("x",Xcenter)
fdtd.set("x span",fdtdSpan)
fdtd.set("y",Xcenter)
fdtd.set("y span",Ysize)
fdtd.set("z",Zcenter)
fdtd.set("y min bc", "Periodic")
fdtd.set("dimension", "2D")
fdtd.addrect()
fdtd.set("name","n_start")
fdtd.set("index",n_start_index)
fdtd.set("x",-((fdtdSpan)-n_start)/2)
fdtd.set("x span",n_start)
fdtd.set("y",0)
fdtd.set("y span",Ysize)
fdtd.set("z",0)
fdtd.set("z span",Zsize)
fdtd.addrect()
fdtd.set("name","n_end")
fdtd.set("index",n_end_index)
fdtd.set("x",((fdtdSpan)-n_start)/2)
fdtd.set("x span",n_end)
fdtd.set("y",0)
fdtd.set("y span",Ysize)
fdtd.set("z",0)
fdtd.set("z span",Zsize)
fdtd.addplane()
fdtd.set("name","Plane Wave")
fdtd.set("injection axis","x")
fdtd.set("direction","forward")
fdtd.set("x",-((fdtdSpan)-n_start)/2)
fdtd.set("y",Ycenter)
fdtd.set("y span",Ysize)
fdtd.set("z",Zcenter)
fdtd.set("z span",Zsize)
fdtd.set("wavelength start",WaveL_Start)
fdtd.set("wavelength stop",WaveL_Stop)
fdtd.addpower()
fdtd.set("name","R_mointor")
fdtd.set("monitor type",3)
fdtd.set("x",(-((((fdtdSpan)-n_start)/2))-0.3e-6))
fdtd.set("y",Ycenter)
fdtd.set("y span",Ysize)
fdtd.set("z",Zcenter)
fdtd.addpower()
fdtd.set("name","T_mointor")
fdtd.set("monitor type",3)
fdtd.set("x",(((((fdtdSpan)-n_start)/2))+0.3e-6))
fdtd.set("y",Ycenter)
fdtd.set("y span",Ysize)
fdtd.set("z",Zcenter)
fdtd.addmovie()
fdtd.set("name","Moive")
fdtd.set("x",Xcenter)
fdtd.set("x span",fdtdSpan)
fdtd.set("y",Ycenter)
fdtd.set("y span",Ysize)
fdtd.setglobalmonitor("frequency points", 1001)
fdtd.setglobalmonitor("use wavelength spacing", 1)
# +
fdtd.select("group");
fdtd.delete();
xSpanN1=0
xSpanN2=0
xN1i=0
xN2i=0
# xSpanN1=0.232e-6
# xSpanN2=0.09688e-6
xN1i = -(fdtdSpan/2-1e-6-(d1/2))
xN2i = ((d1/2)+(d2/2))+xN1i
for _ in range(N):
fdtd.addrect()
fdtd.set("name","n1")
fdtd.set("index",n1)
fdtd.set("x",xN1i)
fdtd.set("x span",d1)
fdtd.set("y",0)
fdtd.set("y span",0.6e-6)
fdtd.set("z",0)
fdtd.set("z span", 0.36e-6)
fdtd.addtogroup("group")
fdtd.addrect()
fdtd.set("name","n2")
fdtd.set("index",n2)
fdtd.set("x",xN2i)
fdtd.set("x span",d2)
fdtd.set("y",0)
fdtd.set("y span",0.6e-6)
fdtd.set("z",0)
fdtd.set("z span", 0.36e-6)
fdtd.addtogroup("group")
xN1i = ((d1/2)+(d2/2))+xN2i
xSpanN1=0.232e-6
xSpanN2=0.09688e-6
xN2i = ((d1/2)+(d2/2))+xN1i
# +
fdtd.save("CAD_Report1_task1.fsp")
fdtd.run()
T = fdtd.getresult("T_mointor","T")
Lam = T["lambda"]
Lam = Lam[:,0]
Trans = T["T"]
R = fdtd.getresult("R_mointor","T")
Refl = R["T"]
plt.figure(figsize = (22,14))
plt.plot(Lam,Trans, "-b", label="Transmittance")
plt.plot(Lam,abs(Refl), "-r", label="Reflectance")
plt.legend(['Transmittance','Reflectance'])
plt.title('Reflectance & Transmittance')
plt.xlabel('Lambda')
plt.ylabel('Reflectance & Transmittance')
plt.grid(True)
plt.show()
fdtd.switchtolayout()
| Bragg_Reflector_FDTD_API_Python.ipynb |
# !pip install imutils
from tensorflow.keras.applications.mobilenet_v2 import preprocess_input
from tensorflow.keras.preprocessing.image import img_to_array
from tensorflow.keras.models import load_model
import matplotlib.pyplot as plt
from imutils import paths
import numpy as np
import cv2
import os
# from os.path import dirname, join,__file__
prototxtPath = 'deploy.prototxt.txt'
weightsPath= 'res10_300x300_ssd_iter_140000.caffemodel'
prototxtPath
net=cv2.dnn.readNet(weightsPath,prototxtPath)
net
## Load model
model=load_model(r'custom_4370_32_100_v2.h5')
# +
dataset = "test"
imagePaths=list(paths.list_images(dataset))
data=[]
for i in imagePaths:
data.append(i)
# -
image=cv2.imread("image (1).png")
image.shape
(h,w)=image.shape[:2]
blob=cv2.dnn.blobFromImage(image,1.0,(300,300),(104.0,177.0,123.0))
blob.shape
net.setInput(blob)
detections = net.forward()
detections
#loop over the detections
det =[]
a=0
for i in range(0,detections.shape[2]):
confidence=detections[0,0,i,2]
print("CONF: ",confidence)
if confidence>0.3:
box=detections[0,0,i,3:7]*np.array([w,h,w,h])
(startX,startY,endX,endY)=box.astype('int')
(startX,startY)=(max(0,startX),max(0,startY))
(endX,endY)=(min(w-1,endX), min(h-1,endY))
face=image[startY:endY, startX:endX]
face=cv2.cvtColor(face,cv2.COLOR_BGR2RGB)
plt.imshow(face)
face=cv2.resize(face,(96,96))
det.append(face)
face=img_to_array(face)
face=preprocess_input(face)
face=np.expand_dims(face,axis=0)
(withoutMask,mask)=model.predict(face)[0]
print("MASK: ",mask)
print("Without: ",withoutMask)
#determine the class label and color we will use to draw the bounding box and text
label='Mask' if mask>withoutMask else 'No Mask'
color=(0,255,0) if label=='Mask' else (255,0,0)
#include the probability in the label
label="{}: {:.2f}%".format(label,max(mask,withoutMask)*100)
#display the label and bounding boxes
cv2.putText(image,label,(startX,startY-10),cv2.FONT_HERSHEY_SIMPLEX,0.45,color,2)
cv2.rectangle(image,(startX,startY),(endX,endY),color,2)
a=a+1
# plt.imshow(det[1])
plt.imshow(image)
| Model_Image_Testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mario-i-caicedo-ai/Ondas-y-Optica/blob/main/cuadernos/Modos_Normales_Cuerda_Vibrante.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="nYhJFSHjMqNp"
# # Cuerda Vibrante de Extremos Fijos
#
# ### Visualización de los Modos Normales.
#
#
# Prof. <NAME>
# + [markdown] id="B0fALW-3n3_j"
# * El estudio de la dinámica de la cuerda cargada nos enseña que un tal sistema con $N$ masas, posee exactamente $N$ modos normales de oscilación
#
# * Es posible hacer un paso al límite
#
# $$N\rightarrow\infty$$
#
# * Este límite es conocido como límite del contínuo.
#
# * En el ímite del contínuo el índice $n$ que marca a cada partícula pasa a ser una coordenada ($x$), y las posiciones ($\psi_n(t)$) de las masas con respecto a sus posiciones de equilibrio se convierten en una función de dos variables $u(x,t)$ que representa las oscilaciones transversales de una cuerda vibrante con sus extremos ($x=0,L$) fijos.
#
# * El sistema de ecuaciones dinámicas para las masas se transforma en la ecuación de ondas unidimensional,
#
# $$
# \frac{\partial^{2}u(x,t)}{\partial{}x^{2}}-\frac{1}{v^2}\frac{\partial^{2}u(x,t)}{\partial{}x^{2}}=0
# $$
# donde, debido a las condiciones en los extremos,
# $$
# u(0,t)=0\,,\qquad{}u(L,t)=0
# $$
#
# * En esta situación, los $N$ modos normales de las $N$ masas se transforman en una cadena infinita de modos normales.
#
#
# + [markdown] id="09ZLKDJsJ3xG"
# * Si nos limitamos al caso en que una de las condiciones iniciales es
# $$\frac{\partial{}u(x,t=0)}{\partial{}t}=0$$
#
# Los modos normales adoptan la forma:
#
# $$u_p(x,t)=sen(k_0x)cos(p\omega_0t)\,\quad{}p=1,2,3,\dots$$
#
# $$k_0=\pi/L\,,\quad{}\omega_0=vk_0$$
#
# NOTA: El p-ésimo modo posee $p-1$ puntos que permanecen inmóviles (nodos).
# + [markdown] id="j86_6qHWPAC3"
# **Ejercicio**
#
# Utilice este cuaderno como base para visualizar los modos de la cuerda con extremos libres
# + [markdown] id="cYhG72hPN1gA"
# Bibliotecas
# + [markdown] id="RNy4LcmaqDy7"
# ------------------------------------------------------
# + id="-wSKAbJUH_ZQ"
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# + [markdown] id="enMTuvdPN7Nx"
# Definición de los modos
# + id="0cWSBQxSqMaY"
L=2.
v=.04
# + colab={"base_uri": "https://localhost:8080/"} id="R7j0widCVtow" outputId="1a0175fd-1880-4667-f18b-e0277c8f9530"
print('Cada modo normal está indexado por un entero (n)\n')
print('El número p-ésimo modo tiene p-1 ceros (nodos) modos\n')
p=float(input('Introduzca un entero para determinar el modo '))
# + id="DJsMqN8FIWGT"
def u(x,t):
u=np.zeros(1000)
u = 1.*np.sin(p*(np.pi/L)*x)*np.cos(p*(v*np.pi/L)*t)
return u
# + [markdown] id="foapvpINJoCG"
# * Creando la figura y los ejes
# + colab={"base_uri": "https://localhost:8080/", "height": 338} id="zInEVbvnujrt" outputId="9a34b94d-02f0-4105-d77d-834d2aa5720d"
fig = plt.figure(figsize=(12,5))
ax1 = plt.subplot(1,1,1)
ax1.set_xlim(( 0, 2))
ax1.set_ylim((-2, 2))
ax1.set_xlabel('x')
ax1.set_ylabel('u(x,t)')
txt_title = ax1.set_title('')
line1, = ax1.plot([], [], 'b', lw=2)
ax1.legend(['u(x,t)']);
# + [markdown] id="A00lgchCLgs7"
# * Animación
# + id="87QfQn9RG4CI"
def init():
line1.set_data([], [])
return line1,
# + id="-VRilK49uzgB"
def drawframe(n):
x = np.linspace(0, 2, 1000)
y1 = u(x,n)
line1.set_data(x, y1)
txt_title.set_text('Frame = {0:4d}'.format(n))
return line1,
# + id="RlvAQwmPu6ft"
from matplotlib import animation
# blit=True re-draws only the parts that have changed.
#anim = animation.FuncAnimation(fig, drawframe, frames=100, interval=20, blit=True)
anim = animation.FuncAnimation(fig, drawframe, frames=100, interval=20, blit=True)
# + colab={"base_uri": "https://localhost:8080/"} id="kHDZr1rFu_5h" outputId="64822d27-6200-4171-b743-bbc07e904f48"
from IPython.display import HTML
HTML(anim.to_html5_video())
| cuadernos/Modos_Normales_Cuerda_Vibrante.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Random Forest from scratch!
# %load_ext autoreload
# %autoreload 2
# +
# %matplotlib inline
from fastai.imports import *
from fastai.structured import *
from sklearn.ensemble import RandomForestRegressor, RandomForestClassifier
from IPython.display import display
from sklearn import metrics
# -
# ## Load in our data from last lesson
# +
PATH = "data/bulldozers/"
df_raw = pd.read_feather('tmp/bulldozers-raw')
df_trn, y_trn, nas = proc_df(df_raw, 'SalePrice')
# -
def split_vals(a,n): return a[:n], a[n:]
n_valid = 12000
n_trn = len(df_trn)-n_valid
X_train, X_valid = split_vals(df_trn, n_trn)
y_train, y_valid = split_vals(y_trn, n_trn)
raw_train, raw_valid = split_vals(df_raw, n_trn)
x_sub = X_train[['YearMade', 'MachineHoursCurrentMeter']]
# ## Basic data structures
class TreeEnsemble():
def __init__(self, x, y, n_trees, sample_sz, min_leaf=5):
np.random.seed(42)
self.x,self.y,self.sample_sz,self.min_leaf = x,y,sample_sz,min_leaf
self.trees = [self.create_tree() for i in range(n_trees)]
def create_tree(self):
rnd_idxs = np.random.permutation(len(self.y))[:self.sample_sz]
return DecisionTree(self.x.iloc[rnd_idxs], self.y[rnd_idxs], min_leaf=self.min_leaf)
def predict(self, x):
return np.mean([t.predict(x) for t in self.trees], axis=0)
class DecisionTree():
def __init__(self, x, y, idxs=None, min_leaf=5):
self.x,self.y,self.idxs,self.min_leaf = x,y,idxs,min_leaf
m = TreeEnsemble(X_train, y_train, n_trees=10, sample_sz=1000, min_leaf=3)
m.trees[0]
class DecisionTree():
def __init__(self, x, y, idxs=None, min_leaf=5):
if idxs is None: idxs=np.arange(len(y))
self.x,self.y,self.idxs,self.min_leaf = x,y,idxs,min_leaf
self.n,self.c = len(idxs), x.shape[1]
self.val = np.mean(y[idxs])
self.score = float('inf')
self.find_varsplit()
# This just does one decision; we'll make it recursive later
def find_varsplit(self):
for i in range(self.c): self.find_better_split(i)
# We'll write this later!
def find_better_split(self, var_idx): pass
@property
def split_name(self): return self.x.columns[self.var_idx]
@property
def split_col(self): return self.x.values[self.idxs,self.var_idx]
@property
def is_leaf(self): return self.score == float('inf')
def __repr__(self):
s = f'n: {self.n}; val:{self.val}'
if not self.is_leaf:
s += f'; score:{self.score}; split:{self.split}; var:{self.split_name}'
return s
m = TreeEnsemble(X_train, y_train, n_trees=10, sample_sz=1000, min_leaf=3)
m.trees[0]
m.trees[0].idxs
# ## Single branch
# ### Find best split given variable
ens = TreeEnsemble(x_sub, y_train, 1, 1000)
tree = ens.trees[0]
x_samp,y_samp = tree.x, tree.y
x_samp.columns
tree
m = RandomForestRegressor(n_estimators=1, max_depth=1, bootstrap=False)
m.fit(x_samp, y_samp)
draw_tree(m.estimators_[0], x_samp, precision=2)
def find_better_split(self, var_idx):
x,y = self.x.values[self.idxs,var_idx], self.y[self.idxs]
for i in range(1,self.n-1):
lhs = x<=x[i]
rhs = x>x[i]
if rhs.sum()==0: continue
lhs_std = y[lhs].std()
rhs_std = y[rhs].std()
curr_score = lhs_std*lhs.sum() + rhs_std*rhs.sum()
if curr_score<self.score:
self.var_idx,self.score,self.split = var_idx,curr_score,x[i]
# %timeit find_better_split(tree,1)
tree
find_better_split(tree,0); tree
# ### Speeding things up
tree = TreeEnsemble(x_sub, y_train, 1, 1000).trees[0]
# +
def std_agg(cnt, s1, s2): return math.sqrt((s2/cnt) - (s1/cnt)**2)
def find_better_split_foo(self, var_idx):
x,y = self.x.values[self.idxs,var_idx], self.y[self.idxs]
sort_idx = np.argsort(x)
sort_y,sort_x = y[sort_idx], x[sort_idx]
rhs_cnt,rhs_sum,rhs_sum2 = self.n, sort_y.sum(), (sort_y**2).sum()
lhs_cnt,lhs_sum,lhs_sum2 = 0,0.,0.
for i in range(0,self.n-self.min_leaf-1):
xi,yi = sort_x[i],sort_y[i]
lhs_cnt += 1; rhs_cnt -= 1
lhs_sum += yi; rhs_sum -= yi
lhs_sum2 += yi**2; rhs_sum2 -= yi**2
if i<self.min_leaf or xi==sort_x[i+1]:
continue
lhs_std = std_agg(lhs_cnt, lhs_sum, lhs_sum2)
rhs_std = std_agg(rhs_cnt, rhs_sum, rhs_sum2)
curr_score = lhs_std*lhs_cnt + rhs_std*rhs_cnt
if curr_score<self.score:
self.var_idx,self.score,self.split = var_idx,curr_score,xi
# -
# %timeit find_better_split_foo(tree,1)
tree
find_better_split_foo(tree,0); tree
DecisionTree.find_better_split = find_better_split
tree = TreeEnsemble(x_sub, y_train, 1, 1000).trees[0]; tree
# ## Full single tree
m = RandomForestRegressor(n_estimators=1, max_depth=2, bootstrap=False)
m.fit(x_samp, y_samp)
draw_tree(m.estimators_[0], x_samp, precision=2)
def find_varsplit(self):
for i in range(self.c): self.find_better_split(i)
if self.is_leaf: return
x = self.split_col
lhs = np.nonzero(x<=self.split)[0]
rhs = np.nonzero(x>self.split)[0]
self.lhs = DecisionTree(self.x, self.y, self.idxs[lhs])
self.rhs = DecisionTree(self.x, self.y, self.idxs[rhs])
DecisionTree.find_varsplit = find_varsplit
tree = TreeEnsemble(x_sub, y_train, 1, 1000).trees[0]; tree
tree.lhs
tree.rhs
tree.lhs.lhs
tree.lhs.rhs
# ## Predictions
cols = ['MachineID', 'YearMade', 'MachineHoursCurrentMeter', 'ProductSize', 'Enclosure',
'Coupler_System', 'saleYear']
# %time tree = TreeEnsemble(X_train[cols], y_train, 1, 1000).trees[0]
x_samp,y_samp = tree.x, tree.y
m = RandomForestRegressor(n_estimators=1, max_depth=3, bootstrap=False)
m.fit(x_samp, y_samp)
draw_tree(m.estimators_[0], x_samp, precision=2, ratio=0.9, size=7)
def predict(self, x): return np.array([self.predict_row(xi) for xi in x])
DecisionTree.predict = predict
if something:
x= do1()
else:
x= do2()
x = do1() if something else do2()
x = something ? do1() : do2()
# +
def predict_row(self, xi):
if self.is_leaf: return self.val
t = self.lhs if xi[self.var_idx]<=self.split else self.rhs
return t.predict_row(xi)
DecisionTree.predict_row = predict_row
# -
# %time preds = tree.predict(X_valid[cols].values)
plt.scatter(preds, y_valid, alpha=0.05)
metrics.r2_score(preds, y_valid)
m = RandomForestRegressor(n_estimators=1, min_samples_leaf=5, bootstrap=False)
# %time m.fit(x_samp, y_samp)
preds = m.predict(X_valid[cols].values)
plt.scatter(preds, y_valid, alpha=0.05)
metrics.r2_score(preds, y_valid)
# # Putting it together
# +
class TreeEnsemble():
def __init__(self, x, y, n_trees, sample_sz, min_leaf=5):
np.random.seed(42)
self.x,self.y,self.sample_sz,self.min_leaf = x,y,sample_sz,min_leaf
self.trees = [self.create_tree() for i in range(n_trees)]
def create_tree(self):
idxs = np.random.permutation(len(self.y))[:self.sample_sz]
return DecisionTree(self.x.iloc[idxs], self.y[idxs],
idxs=np.array(range(self.sample_sz)), min_leaf=self.min_leaf)
def predict(self, x):
return np.mean([t.predict(x) for t in self.trees], axis=0)
def std_agg(cnt, s1, s2): return math.sqrt((s2/cnt) - (s1/cnt)**2)
# -
class DecisionTree():
def __init__(self, x, y, idxs, min_leaf=5):
self.x,self.y,self.idxs,self.min_leaf = x,y,idxs,min_leaf
self.n,self.c = len(idxs), x.shape[1]
self.val = np.mean(y[idxs])
self.score = float('inf')
self.find_varsplit()
def find_varsplit(self):
for i in range(self.c): self.find_better_split(i)
if self.score == float('inf'): return
x = self.split_col
lhs = np.nonzero(x<=self.split)[0]
rhs = np.nonzero(x>self.split)[0]
self.lhs = DecisionTree(self.x, self.y, self.idxs[lhs])
self.rhs = DecisionTree(self.x, self.y, self.idxs[rhs])
def find_better_split(self, var_idx):
x,y = self.x.values[self.idxs,var_idx], self.y[self.idxs]
sort_idx = np.argsort(x)
sort_y,sort_x = y[sort_idx], x[sort_idx]
rhs_cnt,rhs_sum,rhs_sum2 = self.n, sort_y.sum(), (sort_y**2).sum()
lhs_cnt,lhs_sum,lhs_sum2 = 0,0.,0.
for i in range(0,self.n-self.min_leaf-1):
xi,yi = sort_x[i],sort_y[i]
lhs_cnt += 1; rhs_cnt -= 1
lhs_sum += yi; rhs_sum -= yi
lhs_sum2 += yi**2; rhs_sum2 -= yi**2
if i<self.min_leaf or xi==sort_x[i+1]:
continue
lhs_std = std_agg(lhs_cnt, lhs_sum, lhs_sum2)
rhs_std = std_agg(rhs_cnt, rhs_sum, rhs_sum2)
curr_score = lhs_std*lhs_cnt + rhs_std*rhs_cnt
if curr_score<self.score:
self.var_idx,self.score,self.split = var_idx,curr_score,xi
@property
def split_name(self): return self.x.columns[self.var_idx]
@property
def split_col(self): return self.x.values[self.idxs,self.var_idx]
@property
def is_leaf(self): return self.score == float('inf')
def __repr__(self):
s = f'n: {self.n}; val:{self.val}'
if not self.is_leaf:
s += f'; score:{self.score}; split:{self.split}; var:{self.split_name}'
return s
def predict(self, x):
return np.array([self.predict_row(xi) for xi in x])
def predict_row(self, xi):
if self.is_leaf: return self.val
t = self.lhs if xi[self.var_idx]<=self.split else self.rhs
return t.predict_row(xi)
ens = TreeEnsemble(X_train[cols], y_train, 5, 1000)
preds = ens.predict(X_valid[cols].values)
plt.scatter(y_valid, preds, alpha=0.1, s=6);
metrics.r2_score(y_valid, preds)
# %load_ext Cython
def fib1(n):
a, b = 0, 1
while b < n:
a, b = b, a + b
# + language="cython"
# def fib2(n):
# a, b = 0, 1
# while b < n:
# a, b = b, a + b
# + language="cython"
# def fib3(int n):
# cdef int b = 1
# cdef int a = 0
# cdef int t = 0
# while b < n:
# t = a
# a = b
# b = a + b
# -
# %timeit fib1(50)
# %timeit fib2(50)
# %timeit fib3(50)
| courses/ml1/lesson3-rf_foundations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.2
# language: julia
# name: julia-1.0
# ---
# # Interpolation
# # Load Packages
# +
using Dates, Interpolations
include("printmat.jl")
# +
using Plots
backend = "gr" #"gr" (default), "pyplot"
if backend == "pyplot"
pyplot(size=(600,400))
default(fmt = :svg)
else
gr(size=(480,320))
default(fmt = :svg)
end
# -
# # Interpolation of y = f(x)
#
# Interpolations are particularly useful when we repeatedly want to evaluate a function $f(x)$, or when we only know $f(x)$ for a grid of $x$ values but we have strong beliefs that the function is pretty smooth. We then do something like this:
#
# 1. Calculate $f(x)$ values for a grid of $x$. This creates a "look-up" table.
# 2. Replace the expensive calculations of $f(x_i)$ by interpolating $f(x_i)$ from the "look-up" table.
#
# This notebook uses the package Interpolations (see https://github.com/JuliaMath/Interpolations.jl). As an alternative, you may consider the (more traditional) Dierckx.jl package.
# ## Some Values to Be Interpolated
#
# As a simple illustration, we interpolate the sine function. (In practice, the interpolation technique is typically applied to more complicated functions.)
# +
xGrid = range(-pi,stop=pi,length=101) #uniformly spaced grid
yGrid = sin.(xGrid) #y values at xGrid
p1 = plot(xGrid,yGrid,color=:red,linewidth=2,legend=nothing)
title!("the sin function")
xlabel!("x")
ylabel!("y")
# -
# ## Interpolate
#
# The next cell calls on `CubicSplineInterpolation()` to create the "look-up" table (more precisely, create an interpolation object).
#
# To use a cublic spline it is required that the $x_i$ grid is *uniformly spaced* (for instance, 0.1,0.2,...). The case of a non-uniformly spaced $x$ grid is discussed later.
#
# The option `extrapolation_bc=...` determines how extrapolation beyond the range of the $x_i$ grid is done.
#
# The second cell interpolates and extrapolates $y$ at some specific $x$ values.
# +
itp = CubicSplineInterpolation(xGrid,yGrid,extrapolation_bc=Flat())
println()
# +
x = [0.25,0.75] #to interpolate the y values at
y_interpolated = itp(x)
println("x, interpolated y values and true y values")
printmat([x y_interpolated sin.(x)])
x2 = [1.25,pi+0.1,pi+0.5] #to extrapolate the y values at
y_extrapolated = itp(x2) #"extrapolation" can be done
#inside and outside the range xGrid
println("x2 and extrapolated values")
printmat([x2 y_extrapolated])
# -
# ## Plotting the Results
p1 = plot(xGrid,yGrid,color=:red,linewidth=2,label="sin function")
scatter!(x,y_interpolated,color=:magenta,markersize=5,marker=:square,label="interpolated")
scatter!(x2,y_extrapolated,color=:blue,markersize=8,label="extrapolated")
title!("the sin function")
xlabel!("x")
ylabel!("y")
# # Interpolation of y=f(x) for General Vectors
#
# That is, when we cannot guarantee that $y_i=f(x_i)$ is from uniformly spaced $x_i$ values. This is useful, for instance, when we have empirical data on $(x_i,y_i)$.
#
# The approach works similar to before, except that the `CubicSplineInterpolation` must be replaced by `LinearInterpolation`.
# +
xGrid2 = deleteat!(collect(xGrid),55:60) #non-uniformly spaced grid
yGrid2 = sin.(xGrid2)
p1 = scatter(xGrid2,yGrid2,color=:red,legend=false)
title!("y")
xlabel!("x")
# +
itp2 = LinearInterpolation(xGrid2,yGrid2,extrapolation_bc=Flat())
y_interpolated = itp2(x)
println("x, interpolated y values and true y values")
printmat([x y_interpolated sin.(x)])
y_extrapolated = itp2(x2)
println("x2 and extrapolated values")
printmat([x2 y_extrapolated])
# -
| Tutorial_23_Interpolation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3-azureml
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# We will start up trying to implement an end-to-end process with similar algorithm and preprocessing as described in the original article (https://docs.microsoft.com/en-us/security/engineering/identifying-security-bug-reports) and see where that gets us first.
#
# If you are new to Jupyter Notebooks, find instructions on use here: https://jupyter-notebook.readthedocs.io/en/stable/
#
# Short version: To run the contents of a cell, press ctrl+enter :)
#
# The first thing we need to do is connect to the Azure Machine Learning Workspace :
# +
from azureml.core import Workspace, Dataset
ws=Workspace.from_config()
# -
# The dataset has been uploaded and registered in the workspace already, so we just need to get it from there
# +
#import dataset from ws:
dataset = Dataset.get_by_name(ws, name='SecBugDataset')
df = dataset.to_pandas_dataframe()
# this is the dataset we want to use:
df.head()
# -
# We'll use the result from one of the labelers, found in the column L2.
# +
# these are the bugs labeled as security bugs by labeler 2
sb = df[df['L2'] == 'Integrity/Security']
print(sb)
# +
# how many bugs are labeled as security bugs?
print(len(sb))
# -
# First, lets run through locally what we want to deploy to run in the cloud:
#
#
# First of all, there are a couple of libraries we need to import. Internal AML libraries to work with datasets and to handle training runs that will be logged to our experiment, and libraries from the machine learning framework "Scikit Learn" which contains functionality for using a classifier algorithm to train on our dataset and output a model.
#
# +
import os
import math
import string
import numpy as np
from azureml.core import Dataset, Run
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import roc_auc_score, accuracy_score
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.externals import joblib
# -
# We could experiment with different labelers or even combine the results from the labelers as in the original article this dataset was retrieved from, but for now we will use the results from labeler 2, so we create a new column "Label", with the contents from the column with labels from labeler 2:
#
df['Label'] = [1 if x =='Integrity/Security' else 0 for x in df['L2']]
# The field we want to use to predict if it is a security bug or not is
# the "summary" field. It is a short text, and the text must be translated into a
# representation the machine learning alogrithm understand. For this we use the tf-idf vectorization algorithm:
#
# +
#do the vectorization - tf-idf
vectorizer = TfidfVectorizer(min_df=2)
tfidf = vectorizer.fit_transform(df['summary'])
tfidf = tfidf.toarray()
# -
# Now the vectorizer has built a vector that represents the summary field,
# built by using the number of times a word is present in a text, weighted by
# how many texts the word occurs in overall, in all the texts.
# We got a matrix with all our texts along the y-axis and the words along the x-axis:
print(tfidf.shape)
words = vectorizer.get_feature_names()
print(len(words))
print(words[10:20])
# Lets create a column in our dataframe with the vectors representing the summary text:
# +
df['summary_vec'] = list(tfidf)
print(df.head())
# -
# What we want to do now is take X - all the texts (the summary column)
# in their vector representation - and y - the column we are using to
# predict them (The label column) - and split them into one training set and one test set.
# We'll use the first portion to train a classifier algorithm, and the
# second portion to test the classifier afterwards, to see how well it performed:
# +
# split the dataset into test and train
x = df['summary_vec'].tolist()
y = df['Label'].tolist()
X_train, X_test, y_train, y_test = train_test_split(x, y, test_size=0.2,stratify=y, random_state=66)
# -
# Now we want to create and train the model:
model = LogisticRegression(random_state=0, solver='lbfgs', multi_class='ovr')
model.fit(X=X_train, y=y_train)
# Lets do the predictions on the test set, data that the classifier wasn't trained on:
y_pred = model.predict(X=X_test)
# After predicting let's use some common measures for performance and test how well our model perform::
# +
auc_weighted = roc_auc_score(y_test, y_pred,average="weighted")
accuracy = accuracy_score(y_test, y_pred)
print(auc_weighted)
print(accuracy)
# -
# We dont want to just run this locally, we would like to run this in a compute cluster in the cloud, and we want to be able to track metrics on how this training performed and so on, and make it available to others in our team
# +
# create Experiment, my container for Runs
from azureml.core import Experiment
experiment = Experiment(workspace=ws, name="SecurityBugClassification")
# +
# create compute resource that I will be using for training my classifier
# If a cluster by that name already exist, use it
from azureml.core.compute import AmlCompute
from azureml.core.compute import ComputeTarget
import os
# choose a name for your cluster
compute_name = os.environ.get('AML_COMPUTE_CLUSTER_NAME', 'cpu-cluster')
# I'll construct a cluster of nodes 0-1 because
# I'll be working with scikit-learn and there's no scaling out to sev
# nodes but I want the cluster to shut down when not in use
compute_min_nodes = os.environ.get('AML_COMPUTE_CLUSTER_MIN_NODES', 0)
compute_max_nodes = os.environ.get('AML_COMPUTE_CLUSTER_MAX_NODES', 1)
# This example uses CPU VM. For using GPU VM, set SKU to STANDARD_NC6
vm_size = os.environ.get('AML_COMPUTE_CLUSTER_SKU', 'STANDARD_D2_V2')
if compute_name in ws.compute_targets:
compute_target = ws.compute_targets[compute_name]
if compute_target and type(compute_target) is AmlCompute:
print('found compute target. just use it. ' + compute_name)
else:
print('creating a new compute target...')
provisioning_config = AmlCompute.provisioning_configuration(vm_size=vm_size,
min_nodes=compute_min_nodes,
max_nodes=compute_max_nodes)
# create the cluster
compute_target = ComputeTarget.create(ws, compute_name, provisioning_config)
# can poll for a minimum number of nodes and for a specific timeout.
# if no min node count is provided it will use the scale settings for the cluster
compute_target.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)
# For a more detailed view of current AmlCompute status, use get_status()
print(compute_target.get_status().serialize())
# -
# Now we want to submit a job to run on the remote training cluster we have created. To do that we need to:
#
# * Create a training script
# * Create an estimator object
# * Submit the job
#
# We will put the files that will be copied to the remote cluster nodes for execution in the folder "train-dataset":
script_folder = os.path.join(os.getcwd(), 'train-dataset')
# The directory must contain a file with the training script you want to run. For better visibiilty into what the script does, we'll create the file here and add it to the directory we just created:
# +
# %%writefile $script_folder/train.py
import os
import math
import string
import numpy as np
from azureml.core import Dataset, Run
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import roc_auc_score, accuracy_score
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.externals import joblib
run = Run.get_context()
# get input dataset by name
dataset = run.input_datasets['SecBugDataset']
df = dataset.to_pandas_dataframe()
# create column used as target
df['Label'] = [1 if x =='Integrity/Security' else 0 for x in df['L2']]
# do the vectorization - tf-idf
vectorizer = TfidfVectorizer(min_df=2)
tfidf = vectorizer.fit_transform(df['summary'])
tfidf = tfidf.toarray()
# create our feature column
df['summary_vec'] = list(tfidf)
#dividing X,y into train and test data
X_train, X_test, y_train, y_test = train_test_split(df['summary_vec'].tolist(), df['Label'].tolist(), test_size=0.2, random_state=66)
# create our classifier & train it
model = LogisticRegression(random_state=0, solver='lbfgs', multi_class='ovr')
model.fit(X=X_train, y=y_train)
# make predictions to see how well it does
y_pred = model.predict(X=X_test)
# measure it with to different metrics
auc_weighted = roc_auc_score(y_test, y_pred,average="weighted")
accuracy = accuracy_score(y_test, y_pred)
# log the metrics we want to track and measure on to the Run
run.log("AUC_Weighted", auc_weighted)
run.log("Accuracy", accuracy)
model_file_name = 'LogRegModel.pkl'
# The training script saves the model into a directory named ‘outputs’. Files saved in the
# outputs folder are automatically uploaded into experiment record. Anything written in this
# directory is automatically uploaded into the workspace.
os.makedirs('./outputs', exist_ok=True)
with open(model_file_name, 'wb') as file:
joblib.dump(value=model, filename='outputs/' + model_file_name)
# -
# In our training script we log important metrics to the current run, as well as saving the model created into a directory called 'outputs' that will be uploaded to the workspace and available through our run object when the training (run) is completed. Now, we need to create an estimator object that contains the run configuration:
# +
from azureml.train.sklearn import SKLearn
est = SKLearn(source_directory=script_folder,
entry_script='train.py',
inputs=[dataset.as_named_input('SecBugDataset')],
#environment_definition=env,
pip_packages=['azureml-dataprep[pandas]'],
compute_target=compute_target)
# -
# ... and we submit this to the Experiment it belongs to:
run = experiment.submit(config=est)
run
run.wait_for_completion(show_output=True)
# This is the contents of the output directory after the run:
print(run.get_file_names())
# Lets also register our model to the workspace so that we can retrieve it later for testing and deployment:
# register model
model = run.register_model(model_name='LogRegModel.pkl', model_path='outputs/LogRegModel.pkl')
print(model.name, model.id, model.version, sep='\t')
# Now, just running this model once, with no validation, no parameter tuning or testing out other algorithms to see if they perform better is not something we would to in reality - but for now, lets pretend we're satisfied and wants others to be able to use our model in a real world scenario. Then we need to deploy our model to a web service running in a container so that it can be consumed from other applications.
#
# For that we need:
# * A scoring script to show how to use the model
# * An environment file to show what packages need to be installed
# * A configuration file to build the ACI
# * The model we trained before
#
# Again, we will be creating the scoring script inline for visibility, called score.py. It is used by the web service call to show how to use the model.
#
# You must include two required functions into the scoring script:
# * The `init()` function, which typically loads the model into a global object. This function is run only once when the Docker container is started.
#
# * The `run(input_data)` function uses the model to predict a value based on the input data. Inputs and outputs to the run typically use JSON for serialization and de-serialization, but other formats are supported.
deploy_folder = os.path.join(os.getcwd(), 'deploy-model')
# +
# %%writefile $deploy_folder/score.py
import os
import pickle
import json
import numpy as np
from sklearn.externals import joblib
from sklearn.linear_model import LogisticRegression
from azureml.core.model import Model
from azureml.core import model
def init():
global model
# AZUREML_MODEL_DIR is an environment variable created during deployment.
# It is the path to the model folder (./azureml-models/$MODEL_NAME/$VERSION)
# For multiple models, it points to the folder containing all deployed models (./azureml-models)
model_path = os.path.join(os.getenv('AZUREML_MODEL_DIR'), 'LogRegModel.pkl')
# deserialize the model file back into a sklearn model
model = joblib.load(model_path)
# note you can pass in multiple rows for scoring
def run(raw_data):
try:
data = json.loads(raw_data)['data']
data = np.array(data)
result = model.predict([data])
# you can return any data type as long as it is JSON-serializable
return result.tolist()
except Exception as e:
result = str(e)
return result
# -
# Next, create an environment file, called myenv.yml, that specifies all of the script's package dependencies. This file is used to ensure that all of those dependencies are installed in the Docker image. This model needs `scikit-learn` and `azureml-sdk`.
# +
from azureml.core.conda_dependencies import CondaDependencies
myenv = CondaDependencies()
myenv.add_pip_package("scikit-learn==0.20.1")
myenv.add_pip_package("azureml-defaults")
myenv.add_pip_package('azureml-dataprep[pandas]')
with open("./deploy-model/myenv.yml","w") as f:
f.write(myenv.serialize_to_string())
# -
# Create a deployment configuration file and specify the number of CPUs and gigabyte of RAM needed for your ACI container.
# +
from azureml.core.webservice import AciWebservice
aciconfig = AciWebservice.deploy_configuration(cpu_cores=1,
memory_gb=1,
tags={"data": "SecBugDataset", "method" : "sklearn"},
description='Predict Security Bugs with sklearn')
# -
# Configure the image and deploy. The following code goes through these steps:
#
# 1. Create environment object containing dependencies needed by the model using the environment file (`myenv.yml`)
# 1. Create inference configuration necessary to deploy the model as a web service using:
# * The scoring file (`score.py`)
# * environment object created in previous step
# 1. Deploy the model to the ACI container.
# 1. Get the web service HTTP endpoint.
# +
# %%time
from azureml.core.webservice import Webservice
from azureml.core.model import InferenceConfig, Model
from azureml.core.environment import Environment
scorefile = os.path.join(os.getcwd(), 'deploy-model','score.py')
myenvfile = os.path.join(os.getcwd(), 'deploy-model','myenv.yml')
myenv = Environment.from_conda_specification(name="myenv", file_path=myenvfile)
inference_config = InferenceConfig(entry_script=scorefile, environment=myenv)
service = Model.deploy(workspace=ws,
name='secbug-sklearn-logreg-svc-5',
models=[model],
inference_config=inference_config,
deployment_config=aciconfig)
service.wait_for_deployment(show_output=True)
# -
# Get the scoring web service's HTTP endpoint, which accepts REST client calls. This endpoint can be shared with anyone who wants to test the web service or integrate it into an application:
print(service.scoring_uri)
# We have come this far and are able to test our model locally in this notebook, but there are several things that we must deal with still. First, we would need to be able to call the scoring script with text input, not the vector. To accomplish this, we'd need to do some additional work - and also, it might seem like with the high accuracy score the model did well. But really, with a dataset where the amount of elements in the class we are trying to predict are so few, compared to the rest non-security items, high accuracy is not a good way to measure. The AUC is a better measure for this kind of skewed classes, and we got an auc score of 0.5, which isn't very good. It means the model cant really discriminate well between the two classes. Read more about that here: https://towardsdatascience.com/understanding-auc-roc-curve-68b2303cc9c5 For now, we will move over to another notebook and another approach: Automated Machine Learning.
| AISec Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
df = pd.read_csv("HR_comma_sep.csv")
# df.head()
# df.shape
# df.info()
df.describe()
# df.salary = df.salary.replace(["low", "medium", "high"], [0,1,2])
import numpy as np
df = pd.get_dummies(df)
df.head()
import seaborn as sn
cor = df.corr()
# cor = np.abs(cor)
# print(cor)
sn.heatmap(cor)
tm = df.groupby("left").mean()
# print(df.columns)
# tm
# df.groupby("left")["time_spend_company"].describe()
df.groupby("left")["average_montly_hours"].describe()
# tm
v1 = tm.iloc[0,:]
v2 = tm.iloc[1,:]
abs((v1-v2)/(v1+v2))
df = pd.read_csv("HR_comma_sep.csv")
# pd.crosstab(df.salary, df.left).plot(kind='bar')
pd.crosstab(df.Department, df.left, normalize=True, margins=True) #requested by abhijit #not kaam ka like him
# pd.crosstab(df.Department, df.left, normalize=0, margins=True)
# +
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from xgboost import XGBClassifier
from sklearn.metrics import accuracy_score
import numpy as np
import pandas as pd
df = pd.read_csv("HR_comma_sep.csv")
df.drop(['last_evaluation', 'number_project'], axis='columns', inplace=True)
df["salary"].replace(["low", "medium", "high"], [0,1,2], inplace=True)
df = pd.get_dummies(df)
from sklearn.model_selection import train_test_split
#model = LogisticRegression()
model = RandomForestClassifier()
#model = XGBClassifier()
#from sklearn.svm import SVC
#model = SVC()
y = df["left"]
df.drop("left", axis="columns", inplace=True)
X = df
X["average_montly_hours"] = pd.cut(X["average_montly_hours"], bins=[0,150,200,1000], labels=[2,1,3])
X["time_spend_company"] = pd.cut(X["time_spend_company"], bins=[0,2,5, 1000], labels=[3,2,1])
X["average_montly_hours"] = X["average_montly_hours"].astype('int64')
X["time_spend_company"] = X["time_spend_company"].astype('int64')
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=13)
model.fit(X_train, y_train)
print(model.score(X_test,y_test))
from sklearn.metrics import confusion_matrix
yp = model.predict(X_test)
print("left", sum(yp!=0))
print("not left", sum(yp==0))
accuracy_score(y_test, yp)
#cm = confusion_matrix(y_test, yp)
# import seaborn as sn
# sn.heatmap(cm , annot=True)
cm
# -
| HR_DATA_analysis/HR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pylab as plt
import matplotlib as mpl
import mpl_toolkits.axes_grid1 as axgrid
from mpl_toolkits.axes_grid1 import make_axes_locatable
plt.style.use('dark_background')
plt.rcParams['mathtext.fontset'] = 'stix'
plt.rcParams['font.family'] = 'STIXGeneral'
plt.rcParams['font.size'] = 22
"""
How to plot several lines on a
standard plot with a color gradient.
"""
#--------------------------#
# Data Stuff #
#--------------------------#
x = np.linspace(0, 2*np.pi, 500)
y = np.cos(2*x)
#--------------------------#
# Colormap Stuff #
#--------------------------#
#note a colormap has RGB values mapped to a value between 0 and 1
cmap = plt.cm.inferno #choose cmap
n = 200 #number of colormap sample points to choose from
colors = cmap(np.linspace(0,1,n)) # sampling n different colors from the colormap
#--------------------------#
# Plotting Stuff #
#--------------------------#
fig, ax = plt.subplots(1,1,figsize=(12,8)) # initialize a figure and axis object
# ax.set_facecolor('#430085')
# fig.patch.set_facecolor('#430085')
for i in range(n): #iterate through each line you want to plot with a different color
ax.plot(x, i*y, color=colors[i])
ax.set_title("TITLE")
ax.set_xlabel("X - LABEL")
ax.set_ylabel("Y - LABEL")
#--------------------------#
# Colorbar Stuff #
#--------------------------#
norm_scaling = mpl.colors.Normalize(vmin=0, vmax=1) #set the max and min y value for your cmap
divider = axgrid.make_axes_locatable(ax)
cax = divider.append_axes("right", size='5%', pad=0.05)
cbar = plt.colorbar(mpl.cm.ScalarMappable(norm=norm_scaling, cmap=cmap), cax=cax)
# cbar = fig.colorbar(mpl.cm.ScalarMappable(norm=norm_scaling, cmap=cmap), ax=ax) #create color bar object
cbar.set_label("TESTING",rotation=270,labelpad=30) #give cbar a label and rotate it
fig.tight_layout() #tidy up the figure
plt.show()
# -
# ## Load into Cycler
#
# If you want to get some hexidecimal values to load into a cycler in a .mplstyle file, just use this:
cmap = plt.cm.prism #choose cmap
n = 10 #number of colormap sample points to choose from
colors = cmap(np.linspace(0,1,n)) # sampling n different colors from the colormap
print([mpl.colors.to_hex(colors[i])[1:] for i in range(n)])
| Useful_Tools/lines_with_gradient_cmap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JiaminJIAN/20MA573/blob/master/src/Importance_sampling_on_digital_put.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="5kehCwAVrYNq" colab_type="text"
# ## **Importance sampling**
#
# ### **Exercise:**
#
# Asset price under $\mathbb{Q}$ follows
#
# $$S_{t} = S_{0} \exp \{\mu t + \sigma W_{t} \}.$$
#
# Consider Digital put with its payoff
#
# $$h(S_{T}) = I (S_{T} < S_{0} e^{-b}) .$$
#
# We want to find the forward price:
#
# $$v = \mathbb{E}^{\mathbb{Q}} [h(S_{T})].$$
#
# Parameters are given as
#
# $$r = 0.03, \sigma = 0.2, \mu = r - \frac{1}{2} \sigma^{2} = 0.01, T = 1, b = 0.39.$$
#
# - Prove that the exact price is $0.02275$.
# - Use OMC find the price.
# - Use $IS(\alpha)$ find the price.
# - Can you show your approach is optimal?
# - Prove or demonstrate IS is more efficient to OMC.
#
# + id="tXxkrCtqrXLF" colab_type="code" colab={}
# + [markdown] id="F7YBRs1jrZWn" colab_type="text"
# ## **Solution:**
#
# (1) Prove that the exact price is $0.02275$.
#
# Solution:
#
# \begin{equation}
# \begin{aligned}
# v &= \mathbb{E}^{\mathbb{Q}} [h(S_{T})] \\
# &= \mathbb{E}^{\mathbb{Q}} [I (S_{T} < S_{0} e^{-b})] \\
# &= \mathbb{Q} (S_{T} < S_{0} e^{-b})\\
# &= \mathbb{Q} (S_{0} \exp \{(r - \frac{1}{2} \sigma^{2}) T + \sigma W_{T}\} < S_{0} e^{-b}) \\
# &= \mathbb{Q} (Z < - \frac{b + (r - \frac{1}{2} \sigma^{2}) T}{\sigma \sqrt{T}}) \\
# &= \mathbb{Q} (Z < - 2) \\
# &= \Phi(-2), \\
# \end{aligned}
# \end{equation}
# where $Z \sim \mathcal N (0, 1)$ under probability measure $\mathbb{Q}$ and $\Phi(\cdot)$ is the culmulative distribution function of standard normal variable. Then we know the exact price of this forward is $\Phi(-2)$.
# + id="GX9evyplxCwa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0ad5ac7f-07ae-46a9-e4fc-6974968bdc95"
import numpy as np
import scipy.stats as ss
v = ss.norm.cdf(-2)
print("The exact value of this forward is:", v)
# + id="HnK-lQA61qmg" colab_type="code" colab={}
# + [markdown] id="NhRvanaixcUb" colab_type="text"
# (2) Use OMC find the price.
#
# Solution:
#
# Since we get $v= \Phi(-2) $ in (1), then we have the estimator
#
# $$\hat{v} \approx \frac{1}{n} \sum_{i = 1}^{n} I(X_{i} < -2),$$
#
# where $X_{i} \sim \mathcal{N}(0, 1)$.
# + id="p7Uxv8QZz7jP" colab_type="code" colab={}
## Use monte carlo method to get the forward price
def monte_carlo_v(N):
sum_v = 0
for i in range(N):
x = np.random.normal()
if x < -2:
sum_v += 1
return sum_v/N
# + id="KxRmNmWV0zEj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b77e67ee-e5a8-40cc-a20f-1252e285ea8c"
## Test the monte carlo method
v_hat = monte_carlo_v(10000)
print("The value of this forward by the monte carlo method is:", v_hat)
# + id="4D2FIVB81pfI" colab_type="code" colab={}
# + [markdown] id="qF8lbtWB1pxl" colab_type="text"
# (3) Use $IS(\alpha)$ find the price.
#
# Solution:
#
# For the importance sampling , we suppose $\phi_{\alpha}(\cdot)$ is the probability distribution function of the Gaussian random variable with mean $-\alpha$ and variance $1$. Then we have
#
# \begin{equation}
# \begin{aligned}
# v &= \Phi(-2) \\
# &= \int_{- \infty}^{-2} \phi_{0}(x) \, d x \\
# &= \int_{- \infty}^{-2} \frac{\phi_{0}(x)}{\phi_{\alpha}(x)} \phi_{\alpha}(x) \, d x \\
# &= \int_{- \infty}^{-2} e^{\frac{1}{2} \alpha^{2} + \alpha x} \phi_{\alpha} (x) \, d x \\
# &= \mathbb{E} [I(Y < - 2) e^{\frac{1}{2} \alpha^{2} + \alpha Y} | Y \sim \phi_{\alpha}] \\
# &\approx \frac{1}{n} \sum_{i = 1}^{n} [I(Y_{i} < - 2) e^{\frac{1}{2} \alpha^{2} + \alpha Y_{i}}]. \\
# \end{aligned}
# \end{equation}
# + id="w6PZq6sZ4vU-" colab_type="code" colab={}
## Use importance sampling method to get the forward price
def importance_sampling_v(N, alpha):
sum_v = 0
for i in range(N):
y = np.random.normal(- alpha, 1)
if y < -2:
sum_v += np.exp(0.5 * alpha ** 2 + alpha * y)
return sum_v / N
# + id="MO_FFaKK6KA9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="28f7520f-0203-4d50-cced-b657ba1e1c71"
## Test the importance sampling method
v_hat = importance_sampling_v(10000, 4)
print("The value of this forward by the importance sampling method is:", v_hat)
# + [markdown] id="NF1Ac12vJ7X8" colab_type="text"
# We can use the importance sampling method to generate a sequence of estimators, and then we can calculate the sample variance of this sequence. And the $\alpha$ which is correspondent to the minimum variance is the desired one. But this method may not very exactly.
# + id="P7nB4l0S72zV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5010477b-a66f-4966-8185-f4d046da830a"
estimate_list = []
variance_list = []
for i in range(10):
for j in range(100):
estimate_list.append(importance_sampling_v(1000, i))
variance_list.append(np.var(estimate_list))
alpha_optimal1 = variance_list.index(min(variance_list))
print("The optimal choice of alpha is", alpha_optimal1)
# + id="e9hfjV4s_PCu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="bd9c55e6-cc4e-4082-849f-29a333e96e9a"
v_hat = importance_sampling_v(10000, alpha_optimal1)
print("The value of this forward by the importance sampling method is:", v_hat)
# + id="x2-MajnuA3xD" colab_type="code" colab={}
# + [markdown] id="nzeuw5S-A-vl" colab_type="text"
# (4) Can you show your approach is optimal?
#
# Solution:
#
# By the result we get from (3), we know that when $\alpha = 3$, the variance of the extimators by the importance sampling method is the minimum one. We can also calculate the varaince of the estimators. As
#
# $$\hat{v} = \frac{1}{n} \sum_{i = 1}^{n} [I(Y_{i} < - 2) e^{\frac{1}{2} \alpha^{2} + \alpha Y_{i}}]$$
#
# by the importance sampling method. We know that
#
# \begin{equation}
# \begin{aligned}
# MSE(\hat{v}) &= var(\hat{v}) \\
# &= \frac{1}{n} var(I(Y_{i} < - 2) e^{\frac{1}{2} \alpha^{2} + \alpha Y_{i}}) \\
# &= \frac{1}{n} \{ \mathbb{E} [I(Y_{i} < - 2) e^{ \alpha^{2} + 2 \alpha Y_{i}}] - \Phi^{2}(-2) \}. \\
# \end{aligned}
# \end{equation}
#
# As we have
#
# \begin{equation}
# \begin{aligned}
# \mathbb{E} [I(Y_{i} < - 2) e^{ \alpha^{2} + 2 \alpha Y_{i}}] &= \int_{- \infty}^{-2} e^{\alpha^{2}+ 2 \alpha y} \frac{1}{\sqrt{2 \pi}} e^{- \frac{(y + \alpha)^{2}}{2}} \, d y \\
# &= \int_{- \infty}^{-2} \frac{1}{\sqrt{2 \pi}} e^{- \frac{y^{2} - \alpha y - \alpha^{2}}{2}} \, d y \\
# &= \int_{- \infty}^{-2} \frac{1}{\sqrt{2 \pi}} e^{- \frac{(y - \alpha)^{2}}{2}} e^{\alpha^{2}} \, d y \\
# &= e^{\alpha^{2}} \Phi(-2-\alpha),
# \end{aligned}
# \end{equation}
#
# then we can get
#
# $$MSE(\hat{v}) = \frac{1}{n} \{e^{\alpha^{2}} \Phi(-2-\alpha) - \Phi^{2}(-2)\}.$$
#
# Thus we know that the desired $\alpha$ satisfies the equation:
#
# $$2 \alpha \Phi(- 2 - \alpha) = \phi(-2 - \alpha).$$
# + id="AD2kdGv7F2IA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c2d34c77-3db2-43e9-893f-b157d18eeff0"
## get the optimal alpha
mse_list = []
for i in range(10):
mse_list.append(np.exp(i**2) * ss.norm.cdf(-2 - i) - ss.norm.cdf(-2) ** 2)
alpha_optimal2 = mse_list.index(min(mse_list))
print("The optimal choice of alpha is", alpha_optimal2)
# + id="XY9acdfHLJ-f" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="15d1cdec-d773-4db1-a0a3-41d7680a579b"
v_hat = importance_sampling_v(10000, alpha_optimal2)
print("The value of this forward by the importance sampling method is:", v_hat)
# + id="DdMLgcUMLTkg" colab_type="code" colab={}
# + [markdown] id="fukqqZZg7ucv" colab_type="text"
# (5) Prove or demonstrate IS is more efficient to OMC.
#
# + [markdown] id="_kyaDtMpLVMa" colab_type="text"
# Solution:
#
# By the Monte-Carlo method, we have
#
# $$\hat{v_{1}} \approx \frac{1}{n} \sum_{i = 1}^{n} I(X_{i} < -2),$$
#
# then the variance of this estimator is
#
# \begin{equation}
# \begin{aligned}
# MSE(\hat{v_{1}}) &= var(\hat{v_{1}}) \\
# &= \frac{1}{n} var(I(X_{i} < - 2)) \\
# &= \frac{1}{n} \{ \mathbb{E} [I(X_{i} < - 2)] - (\mathbb{E} [I(X_{i} < - 2)])^{2} \} \\
# &= \frac{1}{n} \{\Phi(-2) - (\Phi(-2))^{2}\}.
# \end{aligned}
# \end{equation}
#
# For importance sampling method, we have proved that
#
# $$MSE(\hat{v_{2}}) = \frac{1}{n} \{e^{\alpha^{2}} \Phi(-2-\alpha) - \Phi^{2}(-2)\},$$
#
# hence we have
#
# $$MSE(\hat{v_{1}}) - MSE(\hat{v_{2}}) = \frac{\Phi(-2) - e^{\alpha^{2}} \Phi(-2-\alpha)}{n}.$$
#
# When $\alpha = 2$, the difference of MSE between these two method is
#
# $$\frac{\Phi(-2) - e^{4} \Phi(-4)}{n}$$
#
#
# + id="_uEqaHNpLUOs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="997fcce0-0e74-41fe-9921-349387075c63"
## the difference of MSE between these two method
distance = ss.norm.cdf(-2) - np.exp(4) * ss.norm.cdf(-4)
print("The difference of MSE between these two method is:", distance)
# + [markdown] id="70gMDIJTQZBw" colab_type="text"
# Since $\frac{\Phi(-2) - e^{4} \Phi(-4)}{n} > 0$, we know that importance sampling method is more efficient to ordinary Monte-Carlo method.
| src/Importance_sampling_on_digital_put.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# <h1 style="text-align:center;">Resolução numérica de Equações Diferenciais Não Homogêneas</h1>
# <h2>Introdução</h2>
#
# <p style="text-align: justify; text-indent:4em">Diversos fenômenos da ciência, engenharia, economia e da natureza (de modo geral), podem ser descritos/modelados por equações diferenciais. "Em síntese, uma
# equação diferencial é uma equação que envolve uma função desconhecida (incógnita) e suas derivadas"[1]. Em muitos casos, a resolução analítica das equações é extremamente custosa ou inviável. Assim, os métodos numéricos fornecem aproximações para a solução dessas equações diferenciais.</p>
# <p style="text-align: justify; text-indent:4em">Uma Equação Diferencial Ordinária (EDO) é a ED cuja a função incógnita desconhecida depende de uma única variável independente. Dessa maneira, apenas derivadas ordinárias são encontradas nessas equações[2]. Uma EDO pode ser classificada em relação a sua ordem. Por exemplo, se uma EDO tiver como derivada de maior grau uma derivado segunda, ela é dita como uma EDO de ordem 2. Essas equações possuem uma solução geral e também soluções particulares, que dependem das "condições iniciais". Para uma EDO de primeira ordem, apenas uma condição inicial é necessária. Analogamente, uma EDO de terceira ordem requer três condições iniciais para que uma solução particular possa ser encontrada. A imagem abaixo mostra como uma mesmo EDO pode possuir várias soluções particulares que dependem justamente das condições iniciais.</p>
# <center><img src="img/solucoes_edo.png"></center>
# <center style="margin-top:6px; font-size:12px; font-weight:bold; margin-top:-20px">Soluções particulares de uma ODE</center>
# <h2>Métodos Numéricos</h2>
# <p style="text-align: justify; text-indent:4em">Os métodos desenvolvidos nesse trabalho se baseiam na Série de Taylor. Em linhas gerais, é uma série que exprime uma função analítica f em termos de um somatório de potências, de modo que as primeiros valores da série possuem uma contribuição maior, ou seja, medida que o somatório contínua, os novos termos passam a contribuir menos na aproximação da série em um dado ponto. A equação abaixo representa a série de Taylor em torno do ponto <i>a</i>.</p>
#
# \begin{align*}
# f(x) = \sum_{i=0}^N \frac {f^{(n)}(a)}{n!}\ (x-a)^{n}\
# \end{align*}
# <center style="margin-top:6px; font-size:13px; font-weight:bold">Série de Taylor</center>
# <h3>Método de Euler</h3>
#
# <p style="text-align: justify; text-indent:4em">O método de Euler é descrito pela equação abaixo. Ele é conhecido como método da tangente e pode ser obtido como parte do polinômio de Taylor de
# ordem 1. Ele é o método explícito mais básico de resolução de EDO's, tendo sido desenvolvido por <NAME>. Como pode-se ver pela equação abaixo, esse método depende apenas de um valor precedente, por isso ele é classificado como um método de passo simples. </p>
# <br>
# \begin{align*}
# y_{n+1} = y_n + hf(x_n, y_n)
# \end{align*}
# <center style="margin-top:6px; font-size:13px; font-weight:bold">Método de Euler</center>
#
# <p style="text-align: justify; text-indent:4em"> Por ser bastante simples, ele é interessante para aproximações iniciais, porém ele não é eficiente, pois para conseguir aproximações "boas", o intervalo entre os valores do domínio deve ser "muito" pequeno, o que gera um maior esforço computacional. A célula abaixo contém a implementação desse método. Percebe-se que ela possui duas funções, uma que "monta" as listas com os valores da aproximação e a outra que retorna a predição - a ser somada com a imagem do elemento anterior do domínio - de acordo com a edo, o valor precedente do domínio e da imagem e o intervalo h entre os valores do domínio.
# +
# Euler = Range-Kutta de primeira ordem
def predicao_euler(f: object, x: float, y: float, h: float):
"""
funcao que retorna a variacao em y - delta y - em um intervalo x ate x+h
:param f: equcao diferencial de primeira ordem da funcao ex: y' = 2xy
:param x: valor do dominio
:param y: valor da imagem
:param h: 'distancia' entre os elementos do dominio
"""
return h*f(x, y)
def calcular_por_euler(f: object, x0: float, y0: float, h: float, n: int):
"""
Funcao que retorn duas lista: uma com seu dominio e outra com com sua imagem, atraves do metodo
de euler
:param f: equcao diferencial de primeira ordem da funcao ex: y' = 2xy
:param x0: valor inicial do dominio
:param y0: imagem do valor do valor inicial do dominio
:param h: 'distancia' entre os elementos do dominio
:param n: numero de total de elementos em cada conjunto - imagem e dominio
"""
imagem = [0]*n
dominio = [0]*n
imagem[0] = y0
dominio[0] = x0
aux = x0
for i in range(n-1):
imagem[i+1] = imagem[i] + predicao_euler(f, dominio[i], imagem[i], h)
aux += h
dominio[i+1] = aux
return dominio, imagem
# -
# <h3>Métodos de Runge-Kutta</h3>
# <p style="text-align: justify; text-indent:4em"> Os métodos de Runge-Kutta foram desenvolvidos por volta de 1900 por dois matemáticos Alemães, <NAME> e <NAME>. Eles são classificados como métodos de passo simples, porém implicitos iterativos. Nesse trabalho, foram implementandos os métodos de Runge-Kutta (RK) de segunda (RK2), terceira (RK3) e quarta ordem (RK4). O RK2 também é conhecido como Método de Euler Melhorado. Normalmente, quando se fala apenas método de Runge-Kutta, fica subentendido que trata-se do RK4. Abaixo, seguem as expressões desses métodos, bem como suas implementações.</p>
# <br>
#
# <h4>Método de Euler Melhorado / Runge-Kutta de 2ª Ordem</h4>
# <br>
# \begin{align*}
# f(x) = y_{n} + h\frac{ f(x_{n},y_{n}) + f(x_{n+1},y^*_{n+1}) } {2}\
# \end{align*}
# <center style="margin-top:6px; font-size:13px; font-weight:bold">Método de Runge-Kutta de Ordem 2</center>
#
# <p style="text-align: justify; text-indent:4em"> Onde tem-se que:</p>
#
# \begin{align*}
# y^*_{n+1} = y_n + hf(x_n, y_n)
# \end{align*}
# <center style="margin-top:6px; font-size:13px; font-weight:bold">Fator de correção de Runge-Kutta de Ordem 2</center>
# <br>
# +
# Método de euler melhorado = Runge-Kutta de ordem 2
def predicao_rk2(f: object, x: float, y: float, h: float):
"""
funcao que retorna a variacao em y - delta y - em um intervalo x ate x+h atraves
do metodo de euler melhorado
:param f: equcao diferencial de primeira ordem da funcao ex: y' = 2xy
:param x: valor do dominio
:param y: valor da imagem
:param h: 'distancia' entre os elementos do dominio
"""
y_pos = y + h*f(x,y)
return h*(( f(x, y) + f(x + h, y_pos)) /2)
def calcular_por_rk2(f: object, x0: float, y0: float, h: float, n: int):
"""
Funcao que retorn duas lista: uma com seu dominio e outra com com sua imagem, atraves do metodo
de euler melhorado, o mesmo que Runge Kutta de ordem 2
:param f: equcao diferencial de primeira ordem da funcao ex: y' = 2xy
:param x0: valor inicial do dominio
:param y0: imagem do valor do valor inicial do dominio
:param h: 'distancia' entre os elementos do dominio
:param n: numero de total de elementos em cada conjunto - imagem e dominio
"""
#Setup inicial
imagem = [0]*n
dominio = [0]*n
imagem[0] = y0
dominio[0] = x0
aux = x0
for i in range(n-1):
aux += h
dominio[i+1] = aux
imagem[i+1] = imagem[i] + predicao_rk2(f, dominio[i], imagem[i], h)
return dominio, imagem
# -
# <h4>Método Runge-Kutta de 3ª ordem</h4>
# <br>
# \begin{align*}
# \left(
# \begin{array}{c}
# y_{n+1} = y_n + \frac{h}{6}(k_1 + 4k_2 + k_3)\\
# k_1 = f(x_n, y_n)\\
# k_2 = f(x_n + \frac{h}{2}, y_n + \frac{h}{2}k_1)\\
# k_3 = f(x_n + h, y_n + 2hk_2 - hk_1)\\
# \end{array}
# \right)
# \end{align*}
# <center style="margin-top:6px; font-size:13px; font-weight:bold">Método de Runge-Kutta de Ordem 3</center>
# <br>
# +
# Método de range-kutta 3ª ordem
def predicao_rk3(f: object, x: float, y: float, h: float):
"""
funcao que retorna a variacao em y - delta y - em um intervalo x ate x+h atraves
do metodo Runge Kutta 3
:param f: equcao diferencial de primeira ordem da funcao ex: y' = 2xy
:param x: valor do dominio
:param y: valor da imagem
:param h: 'distancia' entre os elementos do dominio
"""
k1 = f(x, y)
k2 = f(x+(h/2), (y+(h*k1/2)))
k3 = f(x+h, y+2*h*k2 - h*k1)
return h/6*(k1 + 4*k2 + k3)
def calcular_por_rk3(f: object, x0: float, y0: float, h: float, n: int):
"""
Funcao que retorn duas lista: uma com seu dominio e outra com com sua imagem, atraves do
metodo Runge Kutta de ordem 3
:param f: equcao diferencial de primeira ordem da funcao ex: y' = 2xy
:param x0: valor inicial do dominio
:param y0: imagem do valor do valor inicial do dominio
:param h: 'distancia' entre os elementos do dominio
:param n: numero de total de elementos em cada conjunto - imagem e dominio
"""
#Setup inicial
imagem = [0]*n
dominio = [0]*n
imagem[0] = y0
dominio[0] = x0
aux = x0
for i in range(n-1):
aux += h
dominio[i+1] = aux
imagem[i+1] = imagem[i] + predicao_rk3(f, dominio[i], imagem[i], h)
return dominio, imagem
# -
# <h4>Método Runge-Kutta de 4ª ordem</h4>
#
# <br>
# \begin{align*}
# \left(
# \begin{array}{c}
# y_{n+1} = y_n + \frac{1}{6}(k_1 + 2k_2 + 2k_3 + k_4)\\
# k_1 = hf(x_n, y_n)\\
# k_2 = hf(x_n + \frac{h}{2}, y_n + \frac{k_1}{2})\\
# k_3 = hf(x_n + \frac{h}{2}, y_n + \frac{k_2}{2})\\
# k_4 = hf(x_n + h, y_n + k_3)\\
# \end{array}
# \right)
# \end{align*}
# <center style="margin-top:6px; font-size:13px; font-weight:bold">Método de Runge-Kutta de Ordem 4</center>
# <br>
# +
# Método de range-kutta 4ª ordem
def predicao_rk4(f: object, x: float, y: float, h: float):
"""
funcao que retorna a variacao em y - delta y - em um intervalo x ate x+h atraves
do metodo Runge Kutta 4
:param f: equcao diferencial de primeira ordem da funcao ex: y' = 2xy
:param x: valor do dominio
:param y: valor da imagem
:param h: 'distancia' entre os elementos do dominio
"""
k1 = h*f(x, y)
k2 = h*f(x+(h/2), y+(k1/2))
k3 = h*f(x+(h/2), y+(k2/2))
k4 = h*f(x+h, y+k3)
return 1/6*(k1 + 2*k2 + 2*k3 + k4)
def calcular_por_rk4(f: object, x0: float, y0: float, h: float, n: int):
"""
Funcao que retorn duas lista: uma com seu dominio e outra com com sua imagem, atraves do
metodo Runge Kutta de ordem 4
:param f: equcao diferencial de primeira ordem da funcao ex: y' = 2xy
:param x0: valor inicial do dominio
:param y0: imagem do valor do valor inicial do dominio
:param h: 'distancia' entre os elementos do dominio
:param n: numero de total de elementos em cada conjunto - imagem e dominio
"""
#Setup inicial
imagem = [0]*n
dominio = [0]*n
imagem[0] = y0
dominio[0] = x0
aux = x0
for i in range(n-1):
aux += h
dominio[i+1] = aux
imagem[i+1] = imagem[i] + predicao_rk4(f, dominio[i], imagem[i], h)
return dominio, imagem
# -
# <h3>Métodos de Passos Múltiplos</h3>
#
# <p style="text-align: justify; text-indent:4em">Conceitualmente, um método numérico começa a partir de um ponto inicial e, em seguida, leva um pequeno passo para a frente no tempo para encontrar o próximo ponto da solução. O processo continua com os passos subsequentes para mapear a solução. Métodos de uma etapa (como o método de Euler) referem-se a apenas um ponto anterior e sua derivada a determinar o valor atual. Métodos como os Runge-Kutta dão alguns passos intermediários (por exemplo, um meio-passo) para obter um método de ordem superior, mas, em seguida, descartam todas as informações anteriores antes de tomar uma segunda etapa. Métodos de várias etapas tentam ganhar eficiência, mantendo e usando as informações a partir das etapas anteriores, em vez de descartá-las. Consequentemente, os métodos de várias etapas referem-se a vários pontos anteriores e valores derivados. No caso de métodos de várias etapas lineares, uma combinação linear dos pontos anteriores e os valores derivados são utilizados. [3] </p>
#
# <h4>Método de Adam-Bashforth</h4>
#
# \begin{align*}
# y_{n+1} = y_n + \frac{h}{24}[55f(x_n, y_n) - 59f(x_{n-1}, y_{n-1} + 37f(x_{n-2}, y_{n-2} - 9f(x_{n-3}, y_{n-3} )]
# \end{align*}
#
# <br>
# +
# Método Adams-Bashforth de quarta ordem
def predicao_ab4(f: object, xn: float, yn: float, condicoes_anteriores: list, h: float):
"""
funcao que retorna a variacao em y - delta y - em um intervalo x ate x+h atraves
do metodo Adams Bashford de ordem 4
:param f: equcao diferencial de primeira ordem da funcao ex: y' = 2xy
:param x: valor do dominio
:param y: valor da imagem
:param condicoes_anteriores: ex:[[xn_-3, xn_-2, xn_-1], [yn_-3, yn_-2, yn_-1] ]
:param h: 'distancia' entre os elementos do dominio
"""
x_anteriores = condicoes_anteriores[0]
y_anteriores = condicoes_anteriores[1]
k1 = f(xn,yn)
k2 = f(x_anteriores[2], y_anteriores[2])
k3 = f(x_anteriores[1], y_anteriores[1])
k4 = f(x_anteriores[0], y_anteriores[0])
return (h/24)*(55*k1 - 59*k2 + 37*k3 - 9*k4)
def calcular_por_ab4(f: object, x0: float, y0: float, h: float, n: int):
"""
Funcao que retorn duas lista: uma com seu dominio e outra com com sua imagem, atraves do
metodo Adams Bashford de ordem 4
:param f: equcao diferencial de primeira ordem da funcao ex: y' = 2xy
:param x0: valor inicial do dominio
:param y0: imagem do valor do valor inicial do dominio
:param h: 'distancia' entre os elementos do dominio
:param n: numero de total de elementos em cada conjunto - imagem e dominio
"""
#Setup inicial
imagem = [0]*n
dominio = [0]*n
condicoes_iniciais = calcular_por_rk4(f, x0, y0, h, 4)
dominio[0:4] = condicoes_iniciais[0]
imagem[0:4] = condicoes_iniciais[1]
condicoes_anteriores = [[],[]]
aux = dominio[3]
for i in range(3, n-1):
aux += h
dominio[i+1] = aux
condicoes_anteriores[0] = dominio[i-3:i]
condicoes_anteriores[1] = imagem[i-3:i]
imagem[i+1] = imagem[i] + predicao_ab4(f, dominio[i], imagem[i], condicoes_anteriores, h)
return dominio, imagem
# -
# <h4>Método de Adam-Bashfort-Moulton</h4>
#
# <br>
# \begin{align*}
# \left(
# \begin{array}{c}
# y_{n+1} = y_n + \frac{h}{24}[9f(x_{n+1}, p_{n+1}) + 19f(x_{n}, y_{n} - 5f(x_{n-1}, y_{n-1} + f(x_{n-2}, y_{n-2} )]\\
# p_{n+1} = y_n + \frac{h}{24}[55f(x_n, y_n) - 59f(x_{n-1}, y_{n-1} + 37f(x_{n-2}, y_{n-2} - 9f(x_{n-3}, y_{n-3} )] \\
# \end{array}
# \right)
# \end{align*}
#
# <br>
# +
# Método Adams-Bashforth-Moulton de quarta ordem
def predicao_abm4(f: object, xn: float, yn: float, condicoes: list, h: float):
"""
funcao que retorna a variacao em y - delta y - em um intervalo x ate x+h atraves
do metodo Adams Bashford Moulton de ordem 4
:param f: equcao diferencial de primeira ordem da funcao ex: y' = 2xy
:param xn: valor do dominio
:param yn: valor da imagem
:param pn: valor predito por ab4
:param condicoes: ex:[[xn_-3, xn_-2, xn_-1], [yn_-3, yn_-2, yn_-1] ]
:param h: 'distancia' entre os elementos do dominio
"""
x_anteriores = condicoes[0]
y_anteriores = condicoes[1]
p_pos = yn + predicao_ab4(f, xn, yn, condicoes, h)
k_pos = f(xn+h, p_pos)
k_n = f(xn,yn)
k_ant1 = f(x_anteriores[2], y_anteriores[2])
k_ant2 = f(x_anteriores[1], y_anteriores[1])
return (h/24)*(9*k_pos + 19*k_n - 5*k_ant1 + k_ant2)
def calcular_por_abm4(f: object, x0: float, y0: float, h: float, n: int):
"""
Funcao que retorn duas lista: uma com seu dominio e outra com com sua imagem, atraves do
metodo Adams Bashford Moulton de ordem 4
:param f: equcao diferencial de primeira ordem da funcao ex: y' = 2xy
:param x0: valor inicial do dominio
:param y0: imagem do valor do valor inicial do dominio
:param h: 'distancia' entre os elementos do dominio
:param n: numero de total de elementos em cada conjunto - imagem e dominio
"""
#Setup inicial
imagem = [0]*n
dominio = [0]*n
condicoes_iniciais = calcular_por_rk4(f, x0, y0, h, 4)
dominio[0:4] = condicoes_iniciais[0]
imagem[0:4] = condicoes_iniciais[1]
condicoes_anteriores = [[],[]]
aux = dominio[3]
for i in range(3, n-1):
aux += h
dominio[i+1] = aux
condicoes_anteriores[0] = dominio[i-3:i]
condicoes_anteriores[1] = imagem[i-3:i]
imagem[i+1] = imagem[i] + predicao_abm4(f, dominio[i], imagem[i], condicoes_anteriores, h)
return dominio, imagem
# -
# <p style="text-align: justify; text-indent:4em"> Esta célula tem como objetivo implementar funções que realizem alguns procedimentos matemáticos. Uma dessas funções implementadas é a de calcular precisão com base no erro relativo entre os elementos de um sinal original e de um aproximado, baseada na Equação 2.</p>
# <br>
#
# \begin{align*}
# erro = \left |\frac{Original - Aproximado}{Original} \right |
# \end{align*}
# <center style="margin-top:6px;">Equação - Erro Reltivo</center>
#
# <br>
#
# <p style="text-align: justify; text-indent:4em">Outra função implementada é a de calcular o desvio padrão, que assim como a função de calcular precisão, recebe dois conjuntos de valores, retornando o valor do desvio baseado na Equação 3.</p>
#
# <br>
#
# \begin{align*}
# f(x) = \frac{\sqrt{ \sum_{i=0}^N (x_{i}-\bar x)^2 }} {n}\
# \end{align*}
# <center style="margin-top:6px;">Equação - Desvio Padrão</center>
# +
def calcular_precisao(original, aproximado):
erro = 0.0
qtd_numeros = len(original)
for i in range(qtd_numeros):
if original[i] == 0.0:
original[i] = 0.00000000000000000000001
erro += abs( ((original[i] - aproximado[i]))/original[i] )/qtd_numeros
return (1.0-erro)*100
def calcular_erro_absoluto(original, aproximado):
erro = 0.0
qtd_numeros = len(original)
for i in range(qtd_numeros):
erro += abs(original[i] - aproximado[i])
return erro
def calcular_media(lista):
total = 0.0
for num in lista:
total += num
return total/len(lista)
def calcular_desvio_padrao(lista):
desvio = 0.0
n = len(lista)
media = calcular_media(lista)
for num in lista:
desvio += (abs(num - media)**2)/n
return desvio**(1/2)
# -
# <p style="text-align: justify; text-indent:4em">As funções que seguem abaixo representam as equação diferencial e sua solução exata utilizadas para análise dos algoritmos desenvolvidos. A função <span style="font-style:italic'"> edoC</span>, por exemplo, representa a seguinte equação diferencial:</p>
#
# \begin{align*}
# y^{'} = x^2 - e^{-2x}
# \end{align*}
# <center style="margin-top:6px; font-size:13px; font-weight:bold">EDO 1</center>
#
#
# <br>
#
# <p style="text-align: justify; text-indent:4em"> Que possui como solução exata:</p>
# \begin{align*}
# y(x) = \frac{x^3}{3} + 2e^{-2x}
# \end{align*}
# <center style="margin-top:6px; font-size:13px; font-weight:bold">Solução exata da EDO 1</center>
#
# +
import numpy as np
e = 2.71828182846
import math
def edoA(x, y):
#y' = 0.2xy
return 0.2 * x*y
def equacaoA_exata(x):
# y = e^(0.1(x²-1))
return e**(0.1*((x**2)-1))
def edoB(x, y):
#y' = 2xy
return 2*x*y
def equacaoB_exata(x):
return np.exp((x**2)-1)
def edoC(x, y):
return (x**2) - (2*np.exp(-2*x))
def equacaoC_exata(x):
return (x**3)/3 + np.exp(-2*x)
# -
# <h2>Resultados</h2>
#
# <p style="text-align: justify; text-indent:4em">Utilizando a implementação desses métodos de resolução numérica de equações diferencias, pode-se comparar suas respostas com as respostas da solução exata de cada função apresentada acima. Abaixo, seguem os gráficos dos resultados obtidos para vários "steps" entre os valores do domínio, bem como tabelas que mostram a precisão e erro absoluto dos métodos para esses "steps"</p>
# +
from IPython.core.display import HTML
import matplotlib.pyplot as plt
import numpy as np
x0 = 0
y0 = 1
xf = 1.0
edo = edoC
splits = [0.005, 0.05, 0.1]
html = """"""
for h in splits:
n = int(round(np.floor((xf - x0)/h)))+1
# Solucao 'exata'
exata = []
for i in range(n):
x = x0 + h*i
y = equacaoC_exata(x)
exata.append(y)
# Utilizacao dos metodos de resolucao de edoh
t_euler, euler = calcular_por_euler(edo, x0, y0, h, n)
t_rk2, rk2 = calcular_por_rk2(edo, x0, y0, h, n)
t_rk3, rk3 = calcular_por_rk3(edo, x0, y0, h, n)
t_rk4, rk4 = calcular_por_rk4(edo, x0, y0, h, n)
t_ab4, ab4 = calcular_por_ab4(edo, x0, y0, h, n)
t_abm4, abm4 = calcular_por_abm4(edo, x0, y0, h, n)
# Plotagem dos resultados
plt.title('Análise dos resultados com step %.3f' %(h))
plt.xlabel("x")
plt.ylabel("y = f(x)")
legenda_exata, = plt.plot(t_euler, exata, label="Original")
legenda_euler, = plt.plot(t_euler, euler, 'o', label="Euler")
legenda_euler_melhorado, = plt.plot(t_rk2, rk2, 'o', label="RK2")
legenda_rk3, = plt.plot(t_rk3, rk3, 'o', label="RK3")
legenda_rk4, = plt.plot(t_rk4, rk4, 'o', label="RK4")
legenda_ab4, = plt.plot(t_ab4, ab4, 'o', label="AB4")
legenda_abm4, = plt.plot(t_abm4, abm4, 'o', label="ABM4")
plt.legend(handles=[legenda_exata, legenda_euler, legenda_euler_melhorado,
legenda_rk3, legenda_rk4, legenda_ab4, legenda_abm4])
plt.show()
html = html + """
<table style="border:none">
<tr style="border:none">
<th style="padding:20px; border:none; background-color:#f7f7f7;" colspan="3">
<center> h = %.3f </center> """ %h + """
</th>
</tr>
<tr style="border:none">
<th style="padding:20px; border:none; background-color:#f7f7f7;">Método</th>
<th style="padding:20px; border:none; background-color:#f7f7f7;">Precisão (porcentagem)</th>
<th style="padding:20px; border:none; background-color:#f7f7f7;"> Erro absoluto </th>
</tr>
<tr style="border:none;">
<td style="padding:20px; border:none;">Euler</td>
<td style="padding:20px; border:none; color:red;"> %f """ %calcular_precisao(exata, euler) + """ </td>
<td style="padding:20px; border:none; color:red;"> %f """ %calcular_erro_absoluto(exata, euler) + """ </td>
</tr>
<tr style="border:none;">
<td style="padding:20px; border:none;">Runge-Kutta 2</td>
<td style="padding:20px; border:none;"> %f """ %calcular_precisao(exata, rk2) + """ </td>
<td style="padding:20px; border:none;"> %f """ %calcular_erro_absoluto(exata, rk2) + """ </td>
</tr>
<tr style="border:none;">
<td style="padding:20px; border:none;">Runge-Kutta 3</td>
<td style="padding:20px; border:none; color:green;"> %f """ %calcular_precisao(exata, rk3) + """ </td>
<td style="padding:20px; border:none; color:green;"> %f """ %calcular_erro_absoluto(exata, rk3) + """ </td>
</tr>
<tr style="border:none;">
<td style="padding:20px; border:none;">Runge-Kutta 4</td>
<td style="padding:20px; border:none; color:green;"> %f """ %calcular_precisao(exata, rk4) + """ </td>
<td style="padding:20px; border:none; color:green;"> %f """ %calcular_erro_absoluto(exata, rk4) + """ </td>
</tr>
<tr style="border:none;">
<td style="padding:20px; border:none;">Adams-Bashforth 4</td>
<td style="padding:20px; border:none;"> %f """ %calcular_precisao(exata, ab4) + """ </td>
<td style="padding:20px; border:none;"> %f """ %calcular_erro_absoluto(exata, ab4) + """ </td>
</tr>
<tr style="border:none;">
<td style="padding:20px; border:none; color:blue;"><strong>Adams-Bashforth-Moulton 4</strong></td>
<td style="padding:20px; border:none; color:blue;">
<strong> %f """ %calcular_precisao(exata, abm4) + """</strong>
</td>
<td style="padding:20px; border:none; color:blue;">
<strong> %f """ %calcular_erro_absoluto(exata, abm4) + """</strong>
</td>
</tr>
"""
HTML(html)
# -
# <h3>Comparativo com a função ODE do Scipy</h3>
#
# <p style="text-align: justify; text-indent:4em">Para mensurar os resultados obtidos através da implementação dos métodos propostos, pode-se comparar os resultados do método de Runge-Kutta 4, que obteve os melhores resultados para a ODE analisada nesse trabalho, com a função ODE do Scipy.</p>
# +
import time
import numpy as np
from scipy import integrate
from matplotlib.pylab import *
from IPython.core.display import HTML
x0 = 0
y0 = 1
xf = 1.0
h = 0.05
edo = edoC
n = int(round(np.floor((xf - x0)/h)))+1
# Solucao 'exata'
exata = []
for i in range(n):
x = x0 + h*i
y = equacaoC_exata(x)
exata.append(y)
tempo_scipy = time.time()
# Solucao Scipy
r = integrate.ode(edoC).set_integrator('vode', method='bdf')
r.set_initial_value([y0], x0)
dominio_scipy = np.zeros((n, 1))
sol_scipy = np.zeros((n, 1))
dominio_scipy[0] = x0
sol_scipy[0] = y0
k = 1
while r.successful() and k < n:
r.integrate(r.t + h)
dominio_scipy[k] = r.t
sol_scipy[k] = r.y[0]
k += 1
tempo_scipy = time.time() - tempo_scipy
tempo_rk4_implementado = time.time()
t, rk4 = calcular_por_rk4(edo, x0, y0, h, n)
tempo_rk4_implementado = time.time() - tempo_rk4_implementado
legenda_exata, = plt.plot(t, exata, label="Solução Exata", linewidth=8.0)
legenda_rk4, = plt.plot(t, rk4, '--', label="RK4", color='black')
legenda_sol_scipy, = plt.plot(t, sol_scipy, 'o', color="y", label="Scipy-ODE")
plt.title("Comparativo com a função ODE do scipy")
plt.legend(handles=[legenda_exata, legenda_rk4, legenda_sol_scipy])
grid('on')
xlabel('x')
ylabel('y')
plt.show()
html = """
<table style="border:none">
<tr style="border:none">
<th style="padding:20px; border:none; background-color:#f7f7f7;" colspan="4">
<center> h = %.3f </center> """ %h + """
</th>
</tr>
<tr style="border:none">
<th style="padding:20px; border:none; background-color:#f7f7f7;">Método</th>
<th style="padding:20px; border:none; background-color:#f7f7f7;">Precisão (porcentagem)</th>
<th style="padding:20px; border:none; background-color:#f7f7f7;">Erro absoluto</th>
<th style="padding:20px; border:none; background-color:#f7f7f7;">Tempo Gasto (segundos)</th>
</tr>
<tr style="border:none;">
<td style="padding:20px; border:none;">Scipy-ODE</td>
<td style="padding:20px; border:none;">%s</td>
""" %str(calcular_precisao(exata, sol_scipy)[0]) + """
<td style="padding:20px; border:none;">%s</td>
""" %str(calcular_erro_absoluto(exata, sol_scipy)[0]) + """
<td style="padding:20px; border:none"> %f </td>""" %tempo_scipy + """
<\tr>
<tr style="border:none;">
<td style="padding:20px; border:none;">RK4 Implementado</td>
<td style="padding:20px; border:none;">%s</td> """ %str(calcular_precisao(exata, rk4)) + """
<td style="padding:20px; border:none;">%s</td> """ %str(calcular_erro_absoluto(exata, rk4)) + """
<td style="padding:20px; border:none;"> %f </td>""" %tempo_rk4_implementado + """
</tr>
<tr style="border:none">
<th style="padding:20px; border:none; background-color:#f7f7f7;">Resultados</th>
<th style="padding:20px; border:none; background-color:#f7f7f7; color:green;">
%f """ %(calcular_precisao(exata, rk4)-calcular_precisao(exata, sol_scipy)[0]) + """ mais eficiente
</th>
<th style="padding:20px; border:none; background-color:#f7f7f7; color:green;">
%f """ %(calcular_erro_absoluto(exata, sol_scipy)[0]-calcular_erro_absoluto(exata, rk4)) + """ menor </th>
<th style="padding:20px; border:none; background-color:#f7f7f7; color:green;">
%f """ %(tempo_scipy-tempo_rk4_implementado) + """ mais rápido
</th>
</tr>
</table>"""
HTML(html)
# -
# <p style="text-align: justify; text-indent:4em">Assim, constatou-se que para a ODE analisada nesse trabalho, o método de Runge-Kutta 4 implementado obteve um desempenho melhor do que a da função ODE do Scipy (com os parâmetros defaults) para o step 0.5 no intervalo 0-3. Alcançando uma maior precisão, com um menor erro absoluto, também sendo mais eficiente em relação ao tempo.</p>
# <h2>Equações Diferenciais Ordinárias de Ordem N</h2>
#
# <p style="text-align: justify; text-indent:4em">Para a resolução de equações diferenciais ordinárias de ordem maior do que 1, pode-se transformar essa EDO em um sistema de equações diferenciais de ordem 1. Nessa seção, utilizou-se a EDO abaixo. </p>
#
# \begin{align*}
# \left(
# \begin{array}{c}
# y^{''} = y + e^{x}\\
# y(0) = 1 \\
# y^{'}(0) = 0
# \end{array}
# \right)
# \end{align*}
# <p style="text-align: center"> <strong>EDO de ordem 2 - EDO2 A</strong> </p>
#
# <p style="text-align: justify; text-indent:4em"> Pode-se utilizar variáveis auxiliares que representem as derivadas, da seguinte maneira: </p>
#
# \begin{align*}
# \left(
# \begin{array}{c}
# y_1 = y \\
# y_2 = y^{'}\\
# \end{array}
# \right)
# \end{align*}
#
# <p style="text-align: justify; text-indent:4em"> Assim, a EDO de ordem 2 pode ser escrita através do seguinte sistema de EDO's de ordem 1: </p>
#
# \begin{align*}
# \left(
# \begin{array}{c}
# y_1^{'} = y_2 = f1 \\
# y_2{'} = y_1 + e^{x} = f2\\
# y_1(0) = 1 \\
# y_2(0) = 0
# \end{array}
# \right)
# \end{align*}
#
# <p style="text-align: justify; text-indent:4em"> Analogamente, pode-se fazer o mesmo processo para uma outra EDO de ordem 2, como a mostrada abaixo: </p>
#
# \begin{align*}
# \left(
# \begin{array}{c}
# y^{''} = x + 1 \\
# y(0) = 1 \\
# y^{'}(0) = 0
# \end{array}
# \right)
# \end{align*}
# <p style="text-align: center"> <strong>EDO de ordem 2 - EDO2 B</strong> </p>
#
# <p style="text-align: justify; text-indent:4em"> De tal maneira que tem-se: </p>
#
# \begin{align*}
# \left(
# \begin{array}{c}
# y_1 = y \\
# y_2 = y^{'}
# \end{array}
# \right)
# \end{align*}
#
# <p style="text-align: justify; text-indent:4em"> E, finalmente: </p>
#
# \begin{align*}
# \left(
# \begin{array}{c}
# y_1^{'} = y_2 = f1\\
# y_2{'} = x + y_1 =f2\\
# y_1(0) = 1 \\
# y_2(0) = 0
# \end{array}
# \right)
# \end{align*}
#
# <p style="text-align: justify; text-indent:4em">A eficácia da resolução desse método numérico pode ser comparado com a solução analítica a seguir: </p>
#
#
# \begin{align*}
# y(x) = x^2 + 3x - 3
# \end{align*}
#
# <p style="text-align: justify; text-indent:4em">Assim, essas duas equações ordinárias de ordem 2, foram expressas em sistemas de EDO's de ordem 1. As duas esquações que compõem cada sistema estão representadas na célula abaixo, bem como as soluções exatas utilizadas para a verificação dos métodos de predição.</p>
# +
import numpy as np
# edo2_f1 f1 do sistem de equacaoes diferenciais A: y = y_2
def edo2A_f1(x, ydot):
return ydot
# f2 do sistem de equacaoes diferenciais A: y'_2 = y + e**x
def edo2A_f2(x, y):
return y + np.exp(x)
# Solucao exata da EDO A obtida analiticamente
def edo2A_exata(x):
return (0.25*np.exp(x)) + (0.75 * np.exp(-x)) + (0.5*x*np.exp(x))
# f2 f1 do sistem de equacaoes diferenciais A: y = y_2
def edo2B_f1(x, ydot):
return ydot
# f2 do sistem de equacaoes diferenciais A: y'_2 = x +y
def edo2B_f2(x, y):
return x + y
# Solucao exata da EDO B obtida analiticamente
def edo2B_exata(x):
return np.exp(x) - x
# -
# <p style="text-align: justify; text-indent:4em">Para a resolução desses sistemas de equações diferenciais de ordem 1 que descrevem EDO's de ordem 2, desenvolveu-se uma função que recebe como parâmetro o sistema (as duas equações), bem como o tipo da aproximação (Euler, RK2, Rk3, RK4) e assim reaproveitou-se as aproximações já desenvolvidas na seção anterior.</p>
# +
def aproximacao_edo2(tipo: str, f: list, x_0: float, y_0: float, ydot_0: float, h: float, n: int):
imagem = [0]*n
dominio = [0]*n
ydot = [0]*n
dominio[0] = x_0
imagem[0] = y_0
ydot[0] = ydot_0
if tipo == "euler":
predicao = predicao_euler
elif tipo == "rk2":
predicao = predicao_rk2
elif tipo == "rk3":
predicao = predicao_rk3
elif tipo == "rk4":
predicao = predicao_rk4
aux = dominio[0]
for i in range(n-1):
aux += h
dominio[i+1] = aux
imagem[i+1] = imagem[i] + predicao(f[0], dominio[i], ydot[i], h)
ydot[i+1] = ydot[i] + predicao(f[1], dominio[i], imagem[i], h)
return dominio, imagem, ydot
# teste
sistema = [edo2A_f1, edo2A_f2]
aproximacao_edo2(tipo="rk4", f=sistema, x_0=0, y_0=1, ydot_0=0, h=0.1, n=4)
# -
# <h2>Resultados</h2>
# +
import matplotlib.pyplot as plt
sistema = [edo2A_f1, edo2A_f2]
x0 = 0
y0 = 1
xf = 1
ydot0 = 0
splits = [0.005, 0.05, 0.01]
html = """"""
for split in splits:
n = int(round(np.floor((xf - x0)/split)))+1
x_euler, y_euler, ydot_euler = aproximacao_edo2(tipo="euler", f=sistema,
x_0=x0, y_0=y0, ydot_0=ydot0, h=split, n=n)
x_rk2, y_rk2, ydot_rk2 = aproximacao_edo2(tipo="rk2", f=sistema,
x_0=x0, y_0=y0, ydot_0=ydot0, h=split, n=n)
x_rk3, y_rk3, ydot_rk3 = aproximacao_edo2(tipo="rk2", f=sistema,
x_0=x0, y_0=y0, ydot_0=ydot0, h=split, n=n)
x_rk4, y_rk4, ydot_rk4 = aproximacao_edo2(tipo="rk4", f=sistema,
x_0=x0, y_0=y0, ydot_0=ydot0, h=split, n=n)
# Solucao 'exata'
t = []
exata_2ordem = []
for i in range(n):
x = x0 + split*i
y = edo2A_exata(x)
exata_2ordem.append(y)
t.append(x)
# Plotagem dos resultados
plt.title('Análise dos resultados com step %.3f' %(split))
plt.xlabel("x")
plt.ylabel("y = f(x)")
legenda_exata02, = plt.plot(t, exata_2ordem, label="Exata")
legenda_euler, = plt.plot(x_euler, y_euler, 'o', label="Euler")
legenda_rk2, = plt.plot(x_rk2, y_rk2, 'o', label="RK2")
legenda_rk3, = plt.plot(x_rk2, y_rk3, 'o', label="RK3")
legenda_rk4, = plt.plot(x_rk2, y_rk4, 'o', label="RK4")
#
plt.legend(handles=[legenda_exata02, legenda_euler, legenda_rk2, legenda_rk3, legenda_rk4])
plt.show()
html += """
<table style="border:none">
<tr style="border:none">
<th style="padding:20px; border:none; background-color:#f7f7f7;" colspan="3">
<center> split = %.3f </center> """ %h + """
</th>
</tr>
<tr style="border:none">
<th style="padding:20px; border:none; background-color:#f7f7f7;">Método</th>
<th style="padding:20px; border:none; background-color:#f7f7f7;">Precisão (porcentagem)</th>
<th style="padding:20px; border:none; background-color:#f7f7f7;">Erro absoluto</th>
</tr>
<tr style="border:none;">
<td style="padding:20px; border:none;">Euler</td>
<td style="padding:20px; border:none; color:red;"> %f """ %calcular_precisao(exata_2ordem, y_euler) + """ </td>
<td style="padding:20px; border:none; color:red;"> %f """ %calcular_erro_absoluto(exata_2ordem, y_euler) + """ </td>
</tr>
<tr style="border:none;">
<td style="padding:20px; border:none;">RK2</td>
<td style="padding:20px; border:none;"> %f """ %calcular_precisao(exata_2ordem, y_rk2) + """ </td>
<td style="padding:20px; border:none;"> %f """ %calcular_erro_absoluto(exata_2ordem, y_rk2) + """ </td>
</tr>
<tr style="border:none;">
<td style="padding:20px; border:none;">RK3</td>
<td style="padding:20px; border:none;"> %f """ %calcular_precisao(exata_2ordem, y_rk3) + """ </td>
<td style="padding:20px; border:none;"> %f """ %calcular_erro_absoluto(exata_2ordem, y_rk3) + """ </td>
</tr>
<tr style="border:none;">
<td style="padding:20px; border:none; color:green; font-weight:bold;">RK4</td>
<td style="padding:20px; border:none; color:green; font-weight:bold"> %f
""" %calcular_precisao(exata_2ordem, y_rk4) + """ </td>
<td style="padding:20px; border:none; color:green; font-weight:bold"> %f
""" %calcular_erro_absoluto(exata_2ordem, y_rk4) + """ </td>
</tr>
"""
HTML(html)
# -
# <h2>Conclusão</h2>
#
# <p style="text-align: justify; text-indent:4em">Através do presente trabalho, concluiu-se que os métodos numéricos podem fornecer aproximações significativamente próximas das funções incógnitas de Equações Diferenciais. A eficiência e eficácia dessas aproximações estão instrinsecamente relacionadas com o "step" entre os valores do domínio. Além disso, comparou-se os resultados obtidos com o método de Runge-Kutta 4 no intervalo [0,1] com o step 0.5 na EDO 1 com os resultados oriundos da função ODE da biblioteca scipy - com parâmetros defaults(método de Adams). Nessa comparação, o método implementado foi mais eficiente (aproximação com menor tempo gasto), obteve uma maior precisão e um menor erro absoluto.</p>
# <h2>Referências</h2>
#
# <p>[1] https://www.ppgia.pucpr.br/~jamhour/Download/pub/MatComp/7.%20EquacoesDiferencaisOrdinarias.pdf </p>
# <p>[2] http://www.mat.ufmg.br/~espec/Monografias_Noturna/Monografia_KarineNayara.pdf</p>
# <p>[3] https://pt.wikipedia.org/wiki/M%C3%A9todo_de_passo_m%C3%BAltiplo</p>
# <p>[4] http://members.tripod.com/tiago_pinho/trabalho2/metodos.pdf</p>
#
#
#
| EDOH.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''unit2'': conda)'
# language: python
# name: python37664bitunit2conda5feb989ccf2443a0815999853992303c
# ---
# +
import pandas as pd
df = pd.read_csv('/Users/robertbuckley/repos/DS-Unit-2-Applied-Modeling/module2-wrangle-ml-datasets/PGA_Data_Historical.csv')
# -
df.head()
stat_list = df[df['Season'] == 2010].Statistic.unique().tolist()
stat_list
stat_list = ['Driving Distance',
'Driving Accuracy Percentage',
'Greens in Regulation Percentage',
'Putting Average',
'Scoring Average (Actual)',
'Money Leaders',
'Sand Save Percentage',
'Putts Per Round',
'Scoring Average',
'All-Around Ranking',
'Scrambling',
'Putts made Distance',
'Money per Event Leaders',
'World Money List',
'Ball Striking',
'Longest Drives',
'GIR Percentage from Fairway',
'Total Money (Official and Unofficial)',
'GIR Percentage from Other than Fairway',
'Average Distance of Putts made',
'Last 15 Events - Power',
'Last 15 Events - Accuracy',
'Last 15 Events - Short Game',
'Last 15 Events - Putting',
'Last 15 Events - Scoring',
'Going for the Green - Birdie or Better',
'Short Game Rating',
'Putting Rating',
'Fairway Bunker Tendency']
df = df[df['Statistic'].isin(stat_list)]
df.isnull().sum()
df.shape
df.drop(columns='Statistic')
df['Name_Year'] = df['Player Name'] + df['Season'].astype('str')
df.head()
df.isnull().sum()
df.shape
df = df.pivot(index='Name_Year', columns='Variable', values='Value')
df
df.isnull().sum()
df = df.dropna(subset=['Driving Accuracy Percentage - (%)'])
column_list = df.columns.tolist()
column_list
df['Driving Accuracy Percentage - (%)'].isnull().sum()
col_list = ['Average Distance of Putts made - (AVG DIST.)',
'Average Distance of Putts made - (AVG)', 'Ball Striking - (GIR RANK)',
'Ball Striking - (TOTAL DRV RANK)', 'Driving Accuracy Percentage - (%)',
'Driving Distance - (AVG.)', 'GIR Percentage from Fairway - (%)',
'GIR Percentage from Other than Fairway - (%)',
'Going for the Green - Birdie or Better - (%)',
'Greens in Regulation Percentage - (%)',
'Putting Average - (AVG)',
'Putting Average - (BIRDIE CONVERSION)',
'Putting Average - (GIR PUTTS)', 'Putting Average - (GIR RANK)',
'Putts Per Round - (AVG)',
'Putts made Distance - (DISTANCE IN INCHES)',
'Sand Save Percentage - (%)',
'Short Game Rating - (RATING)',
'Scrambling - (%)',
'Total Money (Official and Unofficial) - (MONEY)']
df = df[col_list]
df.info()
df['Year'] = df.index.str[-4:]
df['Name_Year'] = df.index.str[:-4]
df.info()
df.columns
df = df.apply(lambda x: x.str.replace(',', ''))
df = df.apply(lambda x: x.str.replace("'",''))
df = df.apply(lambda x: x.str.replace("$",''))
df = df.apply(lambda x: x.str.replace('"',''))
df = df.apply(lambda x: x.str.replace(' ','.'))
df = df.apply(lambda x: x.str.replace('T',''))
df = df.drop(columns='Name_Year')
df.isnull().sum()
df.columns
# +
col = ['Average Distance of Putts made - (AVG DIST.)',
'Average Distance of Putts made - (AVG)', 'Ball Striking - (GIR RANK)',
'Ball Striking - (TOTAL DRV RANK)', 'Driving Accuracy Percentage - (%)',
'Driving Distance - (AVG.)', 'GIR Percentage from Fairway - (%)',
'GIR Percentage from Other than Fairway - (%)',
'Going for the Green - Birdie or Better - (%)',
'Greens in Regulation Percentage - (%)', 'Putting Average - (AVG)',
'Putting Average - (BIRDIE CONVERSION)',
'Putting Average - (GIR PUTTS)', 'Putting Average - (GIR RANK)',
'Putts Per Round - (AVG)', 'Putts made Distance - (DISTANCE IN INCHES)',
'Sand Save Percentage - (%)',
'Scrambling - (%)',
'Short Game Rating - (RATING)',
'Total Money (Official and Unofficial) - (MONEY)', 'Year']
df = df[col].astype(float)
# -
test = df[df['Year'] == 2018]
train = df[df['Year'] <= 2017]
train['Total Money (Official and Unofficial) - (MONEY)'].mean()
guess = train['Total Money (Official and Unofficial) - (MONEY)'].mean()
errors = guess - train['Total Money (Official and Unofficial) - (MONEY)']
mean_absolute_error = errors.abs().mean()
print(f'Guessing the average would give me an average error: {mean_absolute_error:,.2f}')
# +
from sklearn.model_selection import train_test_split
train, val = train_test_split(train, stratify=train['Year'], random_state=42)
train.shape, val.shape
# -
target = 'Total Money (Official and Unofficial) - (MONEY)'
X_train = train.drop(columns='Total Money (Official and Unofficial) - (MONEY)')
y_train = train[target]
X_val = val.drop(columns='Total Money (Official and Unofficial) - (MONEY)')
y_val = val[target]
X_test = test.drop(columns='Total Money (Official and Unofficial) - (MONEY)')
y_test = test[target]
X_train.shape, X_val.shape, X_test.shape
from sklearn.linear_model import LinearRegression
model_linear = LinearRegression()
model_linear.fit(X_train, y_train)
y_pred = model_linear.predict(X_val)
errors = y_pred - y_val
mae = errors.abs().mean()
print(f'Train Error: ${mae:,.0f}')
# +
from xgboost import XGBRegressor
model = XGBRegressor(max_depth= 20, random_state=42, n_jobs=-1)
model.fit(X_train, y_train)
# -
model.score(X_val, y_val)
# +
import numpy as np
from scipy.stats import randint, uniform
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import RandomizedSearchCV
param_distributions = {
'n_estimators': randint(50, 500),
'max_depth': [5, 10, 15, 20, None],
'max_features': uniform(0, 1),
}
search = RandomizedSearchCV(
RandomForestRegressor(random_state=42),
param_distributions=param_distributions,
n_iter=5,
cv=2,
scoring='neg_mean_absolute_error',
verbose=10,
return_train_score=True,
n_jobs=-1,
random_state=42
)
search.fit(X_train, y_train);
# -
print('Best hyperparameters', search.best_params_)
print('Cross-validation MAE', -search.best_score_)
model = search.best_estimator_
# +
from xgboost import XGBRegressor
model = XGBRegressor(max_depth= 14, random_state=42, n_jobs=-1)
model.fit(X_train, y_train)
# +
from sklearn.metrics import mean_absolute_error
y_pred = model.predict(X_val)
mae = mean_absolute_error(y_val, y_pred)
print(f'Train Error: ${mae:,.0f}')
# -
y_pred = model.predict(X_test)
mae = mean_absolute_error(y_test, y_pred)
print(f'Test Error: ${mae:,.0f}')
# +
import shap
row = X_val.iloc[[0]]
explainer = shap.TreeExplainer(model)
shap_values = explainer.shap_values(row)
shap.initjs()
shap.force_plot(
base_value=explainer.expected_value,
shap_values=shap_values,
features=row
)
# +
import eli5
from eli5.sklearn import PermutationImportance
permuter = PermutationImportance(
model,
scoring='neg_mean_absolute_error',
n_iter=5,
random_state=42
)
permuter.fit(X_val, y_val)
# +
feature_names = X_val.columns.tolist()
eli5.show_weights(
permuter,
top=None,
feature_names=feature_names
)
# -
import matplotlib.pyplot as plt
plt.rcParams['figure.dpi'] = 92
# +
from pdpbox.pdp import pdp_isolate, pdp_plot
feature= 'Putting Average - (BIRDIE CONVERSION)'
isolated = pdp_isolate(
model=model,
dataset=X_val,
model_features=X_val.columns,
feature=feature
)
# -
pdp_plot(isolated, feature_name=feature, plot_lines=False);
train['Putting Average - (BIRDIE CONVERSION)'].describe()
# +
feature= 'Driving Distance - (AVG.)'
isolated = pdp_isolate(
model=model,
dataset=X_val,
model_features=X_val.columns,
feature=feature
)
# -
pdp_plot(isolated, feature_name=feature, plot_lines=False);
train['Driving Distance - (AVG.)'].describe()
# +
feature= 'Driving Accuracy Percentage - (%)'
isolated = pdp_isolate(
model=model,
dataset=X_val,
model_features=X_val.columns,
feature=feature
)
# -
pdp_plot(isolated, feature_name=feature, plot_lines=False);
train['Driving Accuracy Percentage - (%)'].describe()
# +
from pdpbox.pdp import pdp_interact, pdp_interact_plot
features= ['Putting Average - (BIRDIE CONVERSION)', 'Ball Striking - (GIR RANK)']
interaction = pdp_interact(
model=model,
dataset=X_val,
model_features=X_val.columns,
features=features
)
# -
pdp_interact_plot(interaction, plot_type='grid', feature_names=features);
train['Putting Average - (BIRDIE CONVERSION)']
train['Putting Average - (GIR PUTTS)']
| module2-wrangle-ml-datasets/Unit_2_BW.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#pip install openai
# -
import openai
import wget
import pathlib
import pdfplumber
import numpy as np
import pandas as pd
import json
# +
# def read_pubmed(path: str) -> pd.DataFrame:
# with open(path, 'r') as json_file:
# json_list = list(json_file)
# # read jsonl
# result = [json.loads(json_str) for json_str in json_list]
# return pd.DataFrame(result)
# df_train = read_pubmed('../csci-544-project/data/pubmed-dataset/train.txt')
# df_test = read_pubmed('../csci-544-project/data/pubmed-dataset/test.txt')
# df_val = read_pubmed('../csci-544-project/data/pubmed-dataset/val.txt')
# +
#import data after extraction result
import pickle
with open('../csci-544-project/data/extraction-pubmed.pkl', 'rb') as f:
data = pickle.load(f)
final_df = pd.DataFrame(data)
# -
final_df.head(12)
from torch.utils.data import Dataset, DataLoader
from datasets import load_metric
metric = load_metric("rouge")
# full data, The time of processing all 119924 paper is 250 days :), so run first 1000 paper
# Pubmed after extraction first 1000 paper
import os
import openai
from tqdm import tqdm
openai.organization = "org-nsNh9fJWAg5XDKfN84jwabqj"
openai.api_key = "<KEY>"
engine_list = openai.Engine.list()
#openai.api_key = os.getenv("OPENAI_API_KEY")
output = []
#final_output = []
for i in tqdm(range(1000)):
try:
beginning_tag = "Original:\n"
ending_tag = "\n````\nPolished Sentence:"
text = beginning_tag + str(final_df["predictions_joined"].iloc[i])+ ending_tag
response = openai.Completion.create(
engine="curie",
prompt=text,
temperature=0.5,
max_tokens=500,
top_p=1,
frequency_penalty=0.2,
presence_penalty=0,
stop=["````"]
)
output.append(response["choices"][0]['text'])
except:# if the token exceed the maximum tokens number in GPT-3 model,then just use first 1/60 text
beginning_tag = "Original:\n"
ending_tag = "\n````\nPolished Sentence:"
a = final_df["predictions_joined"].iloc[i][:int(len(final_df["predictions_joined"].iloc[i])/3.5)]
text = beginning_tag + str(a)+ ending_tag
response = openai.Completion.create(
engine="curie",
prompt=text,
temperature=0.5,
max_tokens=500,
top_p=1,
frequency_penalty=0.2,
presence_penalty=0,
stop=["````"]
)
output.append(response["choices"][0]['text'])
#final_output.append(output)
pred_df = pd.DataFrame()
pred_df["summary"] = output
pred_df.to_csv("pubmed_after_extraction_summary_1000.csv",sep = ' ',index = False,header = False)
#after extraction text
fake_preds = output
fake_labels = final_df['references_joined'].head(1000)
metric.compute(predictions=fake_preds, references=fake_labels)
report_df = pd.DataFrame(columns = ['data_resource','data_num','extraction_rough_1','extraction_rough_2','extraction_rough_L'])
# +
pubmed = {'data_resource':'pubmed', 'data_num':1000,'extraction_rough_1':28.43,'extraction_rough_2':8.66,'extraction_rough_L':16.30}
report_df = report_df.append(pubmed, ignore_index = True)
# -
report_df
| notebooks/GPT-3_pubmed.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="KKN-9w0bLG5U"
# # Evaluating the Robustness of Off-Policy Evaluation
# + [markdown] id="8fZr3WnuLFqI"
# ## Setup
# + id="aIsmJfbS739X"
# !pip install -q obp
# + id="Pe-0qKN9BsYv"
# !pip install matplotlib==3.1.1
# + id="klvJDHIcCPEz"
# !pip install -U pandas
# + [markdown] id="Omda8tuDqjkd"
# ## Imports
# + id="O8GhX4Mr7jp0" executionInfo={"status": "ok", "timestamp": 1633531025565, "user_tz": -330, "elapsed": 1790, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
from abc import ABCMeta, abstractmethod
from typing import Union
from scipy.stats import loguniform
from inspect import isclass
from pathlib import Path
from typing import Optional, List, Tuple, Union, Dict
from dataclasses import dataclass
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from tqdm import tqdm
from sklearn.base import BaseEstimator, clone
from sklearn.model_selection._search import BaseSearchCV
from sklearn.metrics import (
roc_auc_score,
log_loss,
mean_squared_error as calc_mse,
mean_absolute_error as calc_mae,
)
from obp.ope import (
RegressionModel,
OffPolicyEvaluation,
BaseOffPolicyEstimator,
)
from obp.types import BanditFeedback
import numpy as np
from sklearn.datasets import load_digits
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier as RandomForest
from sklearn.model_selection import RandomizedSearchCV
import obp
from obp.dataset import (
SyntheticBanditDataset,
logistic_reward_function,
linear_behavior_policy
)
from obp.policy import IPWLearner
from obp.ope import (
DirectMethod,
DoublyRobust,
DoublyRobustWithShrinkage,
InverseProbabilityWeighting,
)
from obp.dataset import MultiClassToBanditReduction
# + id="RdPPTf39GUny"
import warnings
from sklearn.exceptions import ConvergenceWarning
warnings.filterwarnings(action='ignore', category=ConvergenceWarning)
# + [markdown] id="aKsAy3S87vB9"
# ## Utils
# + id="ZZNJjEpH7wGb" executionInfo={"status": "ok", "timestamp": 1633531025566, "user_tz": -330, "elapsed": 6, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
def _choose_uniform(
s: int,
lower: Union[int, float],
upper: Union[int, float],
type_: type,
) -> Union[int, float]:
np.random.seed(seed=s)
assert lower <= upper, "`upper` must be larger than or equal to `lower`"
assert type_ in [int, float], f"`type_` must be int or float but {type_} is given"
if lower == upper:
return lower
if type_ == int:
return np.random.randint(lower, upper, dtype=type_)
else: # type_ == float:
return np.random.uniform(lower, upper)
def _choose_log_uniform(
s: int,
lower: Union[int, float],
upper: Union[int, float],
type_: type,
) -> Union[int, float]:
assert (
lower > 0
), f"`lower` must be greater than 0 when drawing from log uniform distribution but {lower} is given"
assert lower <= upper, "`upper` must be larger than or equal to `lower`"
assert type_ in [int, float], f"`type_` must be int or float but {type_} is given"
if lower == upper:
return lower
if type_ == int:
return int(loguniform.rvs(lower, upper, random_state=s))
else: # type_ == float:
return loguniform.rvs(lower, upper, random_state=s)
# + [markdown] id="G4WkQNyr7kJb"
# ## OPE Evaluators
# + id="An3QvViM7osQ" executionInfo={"status": "ok", "timestamp": 1633531025567, "user_tz": -330, "elapsed": 6, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
class BaseOPEEvaluator(metaclass=ABCMeta):
@abstractmethod
def estimate_policy_value(self) -> None:
"""Estimate policy values."""
raise NotImplementedError
@abstractmethod
def calculate_squared_error(self) -> None:
"""Calculate squared errors."""
raise NotImplementedError
@abstractmethod
def visualize_cdf(self) -> None:
"""Create graph of cumulative distribution function of an estimator."""
raise NotImplementedError
@abstractmethod
def visualize_cdf_aggregate(self) -> None:
"""Create graph of cumulative distribution function of all estimators."""
raise NotImplementedError
@abstractmethod
def save_policy_value(self) -> None:
"""Save estimate policy values to csv file."""
raise NotImplementedError
@abstractmethod
def save_squared_error(self) -> None:
"""Save squared errors to csv file."""
raise NotImplementedError
@abstractmethod
def calculate_au_cdf_score(self) -> None:
"""Calculate AU-CDF score."""
raise NotImplementedError
@abstractmethod
def calculate_cvar_score(self) -> None:
"""Calculate CVaR score."""
raise NotImplementedError
# + id="bF-Fjzis8G5V" executionInfo={"status": "ok", "timestamp": 1633531029291, "user_tz": -330, "elapsed": 3730, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
COLORS = [
"lightcoral",
"plum",
"lightgreen",
"lightskyblue",
"lightsalmon",
"orange",
"forestgreen",
"royalblue",
"gold",
"blueviolet",
"fuchsia",
"lightpink",
"firebrick",
"peru",
"darkkhaki",
"darkolivegreen",
"navy",
"deeppink",
"black",
"silver",
]
LINESTYLES = [
"solid",
(0, (1, 0.6)),
(0, (1, 1.2)),
(0, (1, 1.8)),
(0, (1, 2.4)),
(0, (1, 3)),
]
@dataclass
class InterpretableOPEEvaluator(BaseOPEEvaluator):
"""Class to carry out Interpretable OPE Evaluation.
Parameters
----------
random_states: np.ndarray
list of integers representing random states
length of random_states corresponds to the number of runs
bandit_feedbacks: List[BanditFeedback]
list of bandit feedbacks
evaluation_policies: List[Tuple[float, np.ndarray]]
list of tuples representing evaluation policies
first entry in tuple represents the ground truth policy value
second entry in tuple represents action distribution of evaluation policy
ope_estimators: List[BaseOffPolicyEstimator]
list of ope estimators from obp.ope
ope_estimator_hyperparams: dict
dictionary storing hyperparameters for ope estimators
must be in the following format
ope_estimator_hyperparams = dict(
[OffPolicyEstimator].estimator_name = dict(
[parameter_name] = dict(
"lower":
"upper":
"log":
"type":
)
),
)
regression_models: Optional[List[Union[BaseEstimator, BaseSearchCV]]]
list of regression models to be used in off policy evaluation
each element must either be of type BaseEstimator or BaseSearchCV
regression_model_hyperparams: dict
dictionary storing hyperparameters for regression models
must be in the following format
regression_model_hyperparams = dict(
[model_name] = dict(
[parameter_name] = dict(
"lower":
"upper":
"log":
"type":
)
),
)
pscore_estimators: Optional[List[Union[BaseEstimator, BaseSearchCV]]]
list of classification models to be used in estimating propensity scores of behavior policy
each element must either be of type BaseEstimator or BaseSearchCV
pscore_estimator_hyperparams: dict
dictionary storing hyperparameters for pscore estimators
must be in the following format
pscore_estimator_hyperparams = dict(
[model_name] = dict(
[parameter_name] = dict(
"lower":
"upper":
"log":
"type":
)
),
)
"""
random_states: np.ndarray
ope_estimators: List[BaseOffPolicyEstimator]
bandit_feedbacks: List[BanditFeedback]
evaluation_policies: List[Tuple[float, np.ndarray]]
ope_estimator_hyperparams: Optional[dict] = None
regression_models: Optional[List[Union[BaseEstimator, BaseSearchCV]]] = None
regression_model_hyperparams: Optional[dict] = None
pscore_estimators: Optional[List[Union[BaseEstimator, BaseSearchCV]]] = None
pscore_estimator_hyperparams: Optional[dict] = None
def __post_init__(self) -> None:
self.estimator_names = [est.estimator_name for est in self.ope_estimators]
self.policy_value = None
for i in np.arange(len(self.bandit_feedbacks)):
if self.bandit_feedbacks[i]["position"] is None:
self.bandit_feedbacks[i]["position"] = np.zeros_like(
self.bandit_feedbacks[i]["action"],
dtype=int,
)
if self.reward_type == "binary":
self.reg_model_metric_names = ["auc", "rel_ce"]
else:
self.reg_model_metric_names = ["rel_mse", "rel_mae"]
if not self.ope_estimator_hyperparams:
self.ope_estimator_hyperparams = {
estimator_name: dict() for estimator_name in self.estimator_names
}
if not self.regression_model_hyperparams:
self.regression_model_hyperparams = {
regression_model: dict() for regression_model in self.regression_models
}
if self.pscore_estimators and not self.pscore_estimator_hyperparams:
self.pscore_estimator_hyperparams = {
pscore_estimator: dict() for pscore_estimator in self.pscore_estimators
}
@property
def n_runs(self) -> int:
"""Number of iterations."""
return self.random_states.shape[0]
@property
def n_rounds(self) -> np.ndarray:
"""Number of observations in each given bandit_feedback in self.bandit_feedbacks"""
return np.asarray(
[bandit_feedback["n_rounds"] for bandit_feedback in self.bandit_feedbacks]
)
@property
def n_actions(self) -> np.ndarray:
"""Number of actions in each given bandit_feedback in self.bandit_feedbacks"""
return np.asarray(
[bandit_feedback["n_actions"] for bandit_feedback in self.bandit_feedbacks]
)
@property
def reward_type(self) -> np.ndarray:
"""Whether the reward is binary or continuous"""
if np.unique(self.bandit_feedbacks[0]["reward"]).shape[0] == 2:
return "binary"
else:
return "continuous"
@property
def len_list(self) -> np.ndarray:
"""Number of positions in each given bandit_feedback in self.bandit_feedbacks"""
return np.asarray(
[
int(bandit_feedback["position"].max() + 1)
for bandit_feedback in self.bandit_feedbacks
]
)
def estimate_policy_value(
self,
n_folds_: Union[int, Optional[dict]] = 2,
sample_size: Optional[int] = None,
) -> dict:
"""Estimates the policy values using selected ope estimators under a range of environments."""
# initialize dictionaries to store results
self.policy_value = {est: np.zeros(self.n_runs) for est in self.estimator_names}
self.squared_error = {
est: np.zeros(self.n_runs) for est in self.estimator_names
}
self.reg_model_metrics = {
metric: np.zeros(self.n_runs) for metric in self.reg_model_metric_names
}
for i, s in enumerate(tqdm(self.random_states)):
np.random.seed(seed=s)
# randomly select bandit_feedback
self.bandit_feedback = self._choose_bandit_feedback(s)
if self.pscore_estimators is not None:
# randomly choose pscore estimator
pscore_estimator = np.random.choice(self.pscore_estimators)
# randomly choose hyperparameters of pscore estimator
if isinstance(pscore_estimator, BaseEstimator):
classifier = pscore_estimator
setattr(classifier, "random_state", s)
elif isclass(pscore_estimator) and issubclass(
pscore_estimator, BaseEstimator
):
pscore_estimator_hyperparam = (
self._choose_pscore_estimator_hyperparam(s, pscore_estimator)
)
classifier = clone(pscore_estimator(**pscore_estimator_hyperparam))
else:
raise ValueError(
f"pscore_estimator must be BaseEstimator or BaseSearchCV, but {type(pscore_estimator)} is given."
)
# fit classifier
classifier.fit(
self.bandit_feedback["context"], self.bandit_feedback["action"]
)
estimated_pscore = classifier.predict_proba(
self.bandit_feedback["context"]
)
# replace pscore in bootstrap bandit feedback with estimated pscore
self.bandit_feedback["pscore"] = estimated_pscore[
np.arange(self.bandit_feedback["n_rounds"]),
self.bandit_feedback["action"],
]
# randomly sample from selected bandit_feedback
bootstrap_bandit_feedback = self._sample_bootstrap_bandit_feedback(
s, sample_size
)
# randomly choose hyperparameters of ope estimators
self._choose_ope_estimator_hyperparam(s)
# randomly choose regression model
regression_model = self._choose_regression_model(s)
# randomly choose hyperparameters of regression models
if isinstance(regression_model, BaseEstimator):
setattr(regression_model, "random_state", s)
elif isclass(regression_model) and issubclass(
regression_model, BaseEstimator
):
regression_model_hyperparam = self._choose_regression_model_hyperparam(
s, regression_model
)
regression_model = regression_model(**regression_model_hyperparam)
else:
raise ValueError(
f"regression_model must be BaseEstimator or BaseSearchCV, but {type(regression_model)} is given."
)
# randomly choose evaluation policy
ground_truth, bootstrap_action_dist = self._choose_evaluation_policy(s)
# randomly choose number of folds
if isinstance(n_folds_, dict):
n_folds = _choose_uniform(
s,
n_folds_["lower"],
n_folds_["upper"],
n_folds_["type"],
)
else:
n_folds = n_folds_
# estimate policy value using each ope estimator under setting s
(
policy_value_s,
estimated_rewards_by_reg_model_s,
) = self._estimate_policy_value_s(
s,
bootstrap_bandit_feedback,
regression_model,
bootstrap_action_dist,
n_folds,
)
# calculate squared error for each ope estimator
squared_error_s = self._calculate_squared_error_s(
policy_value_s,
ground_truth,
)
# evaluate the performance of reg_model
r_pred = estimated_rewards_by_reg_model_s[
np.arange(bootstrap_bandit_feedback["n_rounds"]),
bootstrap_bandit_feedback["action"],
bootstrap_bandit_feedback["position"],
]
reg_model_metrics = self._calculate_rec_model_performance_s(
r_true=bootstrap_bandit_feedback["reward"],
r_pred=r_pred,
)
# store results
for est in self.estimator_names:
self.policy_value[est][i] = policy_value_s[est]
self.squared_error[est][i] = squared_error_s[est]
for j, metric in enumerate(self.reg_model_metric_names):
self.reg_model_metrics[metric][i] = reg_model_metrics[j].mean()
return self.policy_value
def calculate_squared_error(self) -> dict:
"""Calculates the squared errors using selected ope estimators under a range of environments."""
if not self.policy_value:
_ = self.estimate_policy_value()
return self.squared_error
def calculate_variance(self, scale: bool = False, std: bool = True) -> dict:
"""Calculates the variance of squared errors."""
if not self.policy_value:
_ = self.estimate_policy_value()
if std:
self.variance = {
key: np.sqrt(np.var(val)) for key, val in self.squared_error.items()
}
else:
self.variance = {
key: np.var(val) for key, val in self.squared_error.items()
}
variance = self.variance.copy()
if scale:
c = min(variance.values())
for est in self.estimator_names:
if type(variance[est]) != str:
variance[est] = variance[est] / c
return variance
def calculate_mean(self, scale: bool = False, root: bool = False) -> dict:
"""Calculates the mean of squared errors."""
if not self.policy_value:
_ = self.estimate_policy_value()
if root: # root mean squared error
self.mean = {
key: np.sqrt(np.mean(val)) for key, val in self.squared_error.items()
}
else: # mean squared error
self.mean = {key: np.mean(val) for key, val in self.squared_error.items()}
mean = self.mean.copy()
if scale:
c = min(mean.values())
for est in self.estimator_names:
if type(mean[est]) != str:
mean[est] = mean[est] / c
return mean
def save_policy_value(
self,
file_dir: str = "results",
file_name: str = "ieoe_policy_value.csv",
) -> None:
"""Save policy_value to csv file."""
path = Path(file_dir)
path.mkdir(exist_ok=True, parents=True)
ieoe_policy_value_df = pd.DataFrame(self.policy_value, self.random_states)
ieoe_policy_value_df.to_csv(f"{file_dir}/{file_name}")
def save_squared_error(
self,
file_dir: str = "results",
file_name: str = "ieoe_squared_error.csv",
) -> None:
"""Save squared_error to csv file."""
path = Path(file_dir)
path.mkdir(exist_ok=True, parents=True)
ieoe_squared_error_df = pd.DataFrame(self.squared_error, self.random_states)
ieoe_squared_error_df.to_csv(f"{file_dir}/{file_name}")
def save_variance(
self,
file_dir: str = "results",
file_name: str = "ieoe_variance.csv",
) -> None:
"""Save squared_error to csv file."""
path = Path(file_dir)
path.mkdir(exist_ok=True, parents=True)
ieoe_variance_df = pd.DataFrame(self.variance.values(), self.variance.keys())
ieoe_variance_df.to_csv(f"{file_dir}/{file_name}")
def visualize_cdf(
self,
fig_dir: str = "figures",
fig_name: str = "cdf.png",
font_size: int = 12,
fig_width: float = 8,
fig_height: float = 6,
kde: Optional[bool] = False,
) -> None:
"""Create a cdf graph for each ope estimator."""
path = Path(fig_dir)
path.mkdir(exist_ok=True, parents=True)
for est in self.estimator_names:
plt.clf()
plt.style.use("ggplot")
plt.rcParams.update({"font.size": font_size})
_, ax = plt.subplots(figsize=(fig_width, fig_height))
if kde:
sns.kdeplot(
x=self.squared_error[est],
kernel="gaussian",
cumulative=True,
ax=ax,
label=est,
linewidth=3.0,
bw_method=0.05,
)
else:
sns.ecdfplot(
self.squared_error[est],
ax=ax,
label=est,
linewidth=3.0,
)
plt.legend()
plt.title(f"{est}: Cumulative distribution of squared error")
plt.xlabel("Squared error")
plt.ylabel("Cumulative probability")
plt.xlim(0, None)
plt.ylim(0, 1.1)
plt.savefig(f"{fig_dir}/{est}_{fig_name}", dpi=100)
plt.show()
def visualize_cdf_aggregate(
self,
fig_dir: str = "figures",
fig_name: str = "cdf.png",
font_size: int = 12,
fig_width: float = 8,
fig_height: float = 6,
xmax: Optional[float] = None,
kde: Optional[bool] = False,
linestyles: Optional[bool] = False,
) -> None:
"""Create a graph containing the cdf of all ope estimators."""
path = Path(fig_dir)
path.mkdir(exist_ok=True, parents=True)
plt.clf()
plt.style.use("ggplot")
plt.rcParams.update({"font.size": font_size})
_, ax = plt.subplots(figsize=(fig_width, fig_height))
for i, est in enumerate(self.estimator_names):
if i < len(COLORS):
color = COLORS[i]
else:
color = np.random.rand(
3,
)
if linestyles:
linestyle = LINESTYLES[i % len(LINESTYLES)]
else:
linestyle = "solid"
if kde:
sns.kdeplot(
x=self.squared_error[est],
kernel="gaussian",
cumulative=True,
ax=ax,
label=est,
linewidth=3.0,
bw_method=0.05,
alpha=0.7,
c=color,
linestyle=linestyle,
)
else:
sns.ecdfplot(
self.squared_error[est],
ax=ax,
label=est,
linewidth=3.0,
alpha=0.7,
c=color,
linestyle=linestyle,
)
plt.legend(loc="lower right")
plt.title("Cumulative distribution of squared error")
plt.xlabel("Squared error")
plt.ylabel("Cumulative probability")
plt.xlim(0, xmax)
plt.ylim(0, 1.1)
plt.savefig(f"{fig_dir}/{fig_name}", dpi=100)
plt.show()
def visualize_squared_error_density(
self,
fig_dir: str = "figures",
fig_name: str = "squared_error_density_estimation.png",
font_size: int = 12,
fig_width: float = 8,
fig_height: float = 6,
) -> None:
"""Create a graph based on kernel density estimation of squared error for each ope estimator."""
path = Path(fig_dir)
path.mkdir(exist_ok=True, parents=True)
for est in self.estimator_names:
plt.clf()
plt.style.use("ggplot")
plt.rcParams.update({"font.size": font_size})
_, ax = plt.subplots(figsize=(fig_width, fig_height))
sns.kdeplot(
self.squared_error[est],
ax=ax,
label=est,
linewidth=3.0,
)
plt.legend()
plt.title(f"{est}: Graph of estimated density of squared error")
plt.xlabel(
"Squared error",
)
plt.savefig(f"{fig_dir}/{est}_{fig_name}", dpi=100)
plt.show()
def calculate_au_cdf_score(
self,
threshold: float,
scale: bool = False,
) -> dict:
"""Calculate AU-CDF score."""
au_cdf_score = {est: None for est in self.estimator_names}
for est in self.estimator_names:
au_cdf_score[est] = np.mean(
np.clip(threshold - self.squared_error[est], 0, None)
)
if scale:
c = max(au_cdf_score.values())
for est in self.estimator_names:
au_cdf_score[est] = au_cdf_score[est] / c
return au_cdf_score
def calculate_cvar_score(
self,
alpha: float,
scale: bool = False,
) -> dict:
"""Calculate CVaR score."""
cvar_score = {est: None for est in self.estimator_names}
for est in self.estimator_names:
threshold = np.percentile(self.squared_error[est], alpha)
bool_ = self.squared_error[est] >= threshold
if any(bool_):
cvar_score[est] = np.sum(self.squared_error[est] * bool_) / np.sum(
bool_
)
else:
cvar_score[
est
] = f"the largest squared error is less than the threshold value {threshold}"
if scale:
c = min(cvar_score.values())
for est in self.estimator_names:
if type(cvar_score[est]) != str:
cvar_score[est] = cvar_score[est] / c
return cvar_score
def set_ope_estimator_hyperparam_space(
self,
ope_estimator_name: str,
param_name: str,
lower: Union[int, float],
upper: Union[int, float],
log: Optional[bool] = False,
type_: Optional[type] = int,
) -> None:
"""Specify sampling method of hyperparameter of ope estimator."""
assert type_ in [
int,
float,
], f"`type_` must be int or float but {type_} is given"
dic = {
"lower": lower,
"upper": upper,
"log": log,
"type": type_,
}
self.ope_estimator_hyperparams[ope_estimator_name][param_name] = dic
def set_regression_model_hyperparam_space(
self,
regression_model: Union[BaseEstimator, BaseSearchCV],
param_name: str,
lower: Union[int, float],
upper: Union[int, float],
log: Optional[bool] = False,
type_: Optional[type] = int,
) -> None:
"""Specify sampling method of hyperparameter of regression model."""
assert type_ in [
int,
float,
], f"`type_` must be int or float but {type_} is given"
dic = {
"lower": lower,
"upper": upper,
"log": log,
"type": type_,
}
self.regression_model_hyperparams[regression_model][param_name] = dic
def _choose_bandit_feedback(
self,
s: int,
) -> BanditFeedback:
"""Randomly select bandit_feedback."""
np.random.seed(seed=s)
idx = np.random.choice(len(self.bandit_feedbacks))
return self.bandit_feedbacks[idx]
def _sample_bootstrap_bandit_feedback(
self, s: int, sample_size: Optional[int]
) -> BanditFeedback:
"""Randomly sample bootstrap data from bandit_feedback."""
bootstrap_bandit_feedback = self.bandit_feedback.copy()
np.random.seed(seed=s)
if sample_size is None:
sample_size = self.bandit_feedback["n_rounds"]
self.bootstrap_idx = np.random.choice(
np.arange(sample_size), size=sample_size, replace=True
)
for key_ in self.bandit_feedback.keys():
# if the size of a certain key_ is not equal to n_rounds,
# we should not resample that certain key_
# e.g. we want to resample action and reward, but not n_rounds
if (
not isinstance(self.bandit_feedback[key_], np.ndarray)
or len(self.bandit_feedback[key_]) != self.bandit_feedback["n_rounds"]
):
continue
bootstrap_bandit_feedback[key_] = bootstrap_bandit_feedback[key_][
self.bootstrap_idx
]
bootstrap_bandit_feedback["n_rounds"] = sample_size
return bootstrap_bandit_feedback
def _choose_ope_estimator_hyperparam(
self,
s: int,
) -> None:
"""Randomly choose hyperparameters for ope estimators."""
for i, est in enumerate(self.ope_estimators):
hyperparam = self.ope_estimator_hyperparams.get(est.estimator_name, None)
if not hyperparam:
continue
for p in hyperparam:
if hyperparam[p].get("log", False):
val = _choose_log_uniform(
s,
hyperparam[p]["lower"],
hyperparam[p]["upper"],
hyperparam[p].get("type", int),
)
else:
val = _choose_uniform(
s,
hyperparam[p]["lower"],
hyperparam[p]["upper"],
hyperparam[p].get("type", int),
)
setattr(est, p, val)
self.ope_estimators[i] = est
def _choose_regression_model(
self,
s: int,
) -> Union[BaseEstimator, BaseSearchCV]:
"""Randomly choose regression model."""
idx = np.random.choice(len(self.regression_models))
return self.regression_models[idx]
def _choose_regression_model_hyperparam(
self,
s: int,
regression_model: Union[BaseEstimator, BaseSearchCV],
) -> dict:
"""Randomly choose hyperparameters for regression model."""
hyperparam = dict(
random_state=s,
)
hyperparam_set = self.regression_model_hyperparams.get(regression_model, None)
if not hyperparam_set:
return hyperparam
for p in hyperparam_set:
if hyperparam_set[p].get("log", False):
val = _choose_log_uniform(
s,
hyperparam_set[p]["lower"],
hyperparam_set[p]["upper"],
hyperparam_set[p].get("type", int),
)
else:
val = _choose_uniform(
s,
hyperparam_set[p]["lower"],
hyperparam_set[p]["upper"],
hyperparam_set[p].get("type", int),
)
hyperparam[p] = val
return hyperparam
def _choose_pscore_estimator_hyperparam(
self,
s: int,
pscore_estimator: Union[BaseEstimator, BaseSearchCV],
) -> dict:
"""Randomly choose hyperparameters for pscore estimator."""
hyperparam = dict(
random_state=s,
)
hyperparam_set = self.pscore_estimator_hyperparams.get(pscore_estimator, None)
if not hyperparam_set:
return hyperparam
for p in hyperparam_set:
if hyperparam_set[p].get("log", False):
val = _choose_log_uniform(
s,
hyperparam_set[p]["lower"],
hyperparam_set[p]["upper"],
hyperparam_set[p].get("type", int),
)
else:
val = _choose_uniform(
s,
hyperparam_set[p]["lower"],
hyperparam_set[p]["upper"],
hyperparam_set[p].get("type", int),
)
hyperparam[p] = val
return hyperparam
def _choose_evaluation_policy(
self,
s: int,
) -> Tuple[float, np.ndarray]:
"""Randomly choose evaluation policy and resample using bootstrap."""
np.random.seed(seed=s)
idx = np.random.choice(len(self.evaluation_policies))
ground_truth, action_dist = self.evaluation_policies[idx]
action_dist = action_dist[self.bootstrap_idx]
return ground_truth, action_dist
def _estimate_policy_value_s(
self,
s: int,
bootstrap_bandit_feedback: BanditFeedback,
_regression_model: Union[BaseEstimator, BaseSearchCV],
bootstrap_action_dist: np.ndarray,
n_folds: int,
) -> Tuple[dict, np.ndarray]:
"""Estimates the policy values using selected ope estimators under a particular environments."""
# prepare regression model for ope
regression_model = RegressionModel(
n_actions=self.bandit_feedback["n_actions"],
len_list=int(self.bandit_feedback["position"].max() + 1),
base_model=_regression_model,
fitting_method="normal",
)
estimated_reward_by_reg_model = regression_model.fit_predict(
context=bootstrap_bandit_feedback["context"],
action=bootstrap_bandit_feedback["action"],
reward=bootstrap_bandit_feedback["reward"],
position=bootstrap_bandit_feedback["position"],
pscore=bootstrap_bandit_feedback["pscore"],
action_dist=bootstrap_action_dist,
n_folds=n_folds,
random_state=int(s),
)
# estimate policy value using ope
ope = OffPolicyEvaluation(
bandit_feedback=bootstrap_bandit_feedback,
ope_estimators=self.ope_estimators,
)
estimated_policy_value = ope.estimate_policy_values(
action_dist=bootstrap_action_dist,
estimated_rewards_by_reg_model=estimated_reward_by_reg_model,
)
return estimated_policy_value, estimated_reward_by_reg_model
def _calculate_squared_error_s(
self,
policy_value: dict,
ground_truth: float,
) -> dict:
"""Calculate squared error."""
squared_error = {
est: np.square(policy_value[est] - ground_truth)
for est in self.estimator_names
}
return squared_error
def _calculate_rec_model_performance_s(
self,
r_true: np.ndarray,
r_pred: np.ndarray,
) -> Tuple[float, float]:
"""Calculate performance of reg model."""
r_naive_pred = np.ones_like(r_true) * r_true.mean()
if self.reward_type == "binary":
auc = roc_auc_score(r_true, r_pred)
ce = log_loss(r_true, r_pred)
ce_naive = log_loss(r_true, r_naive_pred)
rel_ce = 1 - (ce / ce_naive)
return auc, rel_ce
elif self.reward_type == "continuous":
mse = calc_mse(r_true, r_pred)
mse_naive = calc_mse(r_true, r_naive_pred)
rel_mse = 1 - (mse / mse_naive)
mae = calc_mae(r_true, r_pred)
mae_naive = calc_mae(r_true, r_naive_pred)
rel_mae = 1 - (mae / mse_naive)
return rel_mse, rel_mae
def load_squared_error(
self,
file_dir: str,
file_name: str,
) -> None:
df = pd.read_csv(f"{file_dir}/{file_name}")
self.squared_error = {est: None for est in self.estimator_names}
for est in self.estimator_names:
self.squared_error[est] = df[est].values
# + [markdown] id="ZkHWycdN8L3j"
# ## Example 1 - Synthetic dataset
# + [markdown] id="-2MW7AG28mcW"
# This section demonstrates an example of conducting Interpretable Evaluation for Off-Policy Evaluation (IEOE). We use synthetic logged bandit feedback data generated using [`obp`](https://github.com/st-tech/zr-obp) and evaluate the performance of Direct Method (DM), Doubly Robust (DR), Doubly Robust with Shrinkage (DRos), and Inverse Probability Weighting (IPW).
#
# Our example contains the following three major steps:
#
# 1. Data Preparation
# 2. Setting Hyperparameter Spaces for Off-Policy Evaluation
# 3. Interpretable Evaluation for Off-Policy Evaluation
# + [markdown] id="8LIsXGoo8mcc"
# ### Data Preparation
#
# In order to conduct IEOE using `pyieoe`, we need to prepare logged bandit feedback data, action distributions of evaluation policies, and ground truth policy values of evaluation policies. Because `pyieoe` is built with the intention of being used with `obp`, these inputs must follow the conventions in `obp`. Specifically, logged bandit feedback data must be of type `BanditFeedback`, action distributions must be of type `np.ndarray`, and ground truth policy values must be of type `float` (or `int`).
#
# In this example, we generate synthetic logged bandit feedback data and perform off-policy learning to obtain two sets of evaluation policies along with their action distributions and ground truth policy values using `obp`. For a detailed explanation of this process, please refer to the [official obp example](https://github.com/st-tech/zr-obp/blob/master/examples/quickstart/quickstart_synthetic.ipynb).
# + id="JqPDekwo8mcd" executionInfo={"status": "ok", "timestamp": 1633531029293, "user_tz": -330, "elapsed": 11, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
# generate a synthetic bandit dataset with 10 actions
# we use `logistic function` as the reward function and `linear_behavior_policy` as the behavior policy.
# one can define their own reward function and behavior policy such as nonlinear ones.
dataset = SyntheticBanditDataset(
n_actions=10,
dim_context=5,
reward_type="binary", # "binary" or "continuous"
reward_function=logistic_reward_function,
behavior_policy_function=linear_behavior_policy,
random_state=12345
)
# obtain training and test sets of synthetic logged bandit feedback
n_rounds_train, n_rounds_test = 10000, 10000
bandit_feedback_train = dataset.obtain_batch_bandit_feedback(n_rounds=n_rounds_train)
bandit_feedback_test = dataset.obtain_batch_bandit_feedback(n_rounds=n_rounds_test)
# define IPWLearner with Logistic Regression as its base ML model
evaluation_policy_a = IPWLearner(
n_actions=dataset.n_actions,
len_list=dataset.len_list,
base_classifier=LogisticRegression(C=100, random_state=12345)
)
# train IPWLearner on the training set of the synthetic logged bandit feedback
evaluation_policy_a.fit(
context=bandit_feedback_train["context"],
action=bandit_feedback_train["action"],
reward=bandit_feedback_train["reward"],
pscore=bandit_feedback_train["pscore"]
)
# obtains action choice probabilities for the test set of the synthetic logged bandit feedback
action_dist_a = evaluation_policy_a.predict_proba(
context=bandit_feedback_test["context"],
tau=0.1 # temperature hyperparameter
)
# define IPWLearner with Random Forest as its base ML model
evaluation_policy_b = IPWLearner(
n_actions=dataset.n_actions,
len_list=dataset.len_list,
base_classifier=RandomForest(n_estimators=30, min_samples_leaf=10, random_state=12345)
)
# train IPWLearner on the training set of the synthetic logged bandit feedback
evaluation_policy_b.fit(
context=bandit_feedback_train["context"],
action=bandit_feedback_train["action"],
reward=bandit_feedback_train["reward"],
pscore=bandit_feedback_train["pscore"]
)
# obtains action choice probabilities for the test set of the synthetic logged bandit feedback
action_dist_b = evaluation_policy_b.predict_proba(
context=bandit_feedback_test["context"],
tau=0.1 # temperature hyperparameter
)
# obtain ground truth policy value for each action choice probabilities
expected_rewards = bandit_feedback_test["expected_reward"]
ground_truth_a = np.average(expected_rewards, weights=action_dist_a[:, :, 0], axis=1).mean()
ground_truth_b = np.average(expected_rewards, weights=action_dist_b[:, :, 0], axis=1).mean()
# + [markdown] id="q9ekJZa38mch"
# ### Setting Hyperparameter Spaces for Off-Policy Evaluation
#
# An integral aspect of IEOE is the different sources of variance. The main sources of variance are evaluation policies, random states, hyperparameters of OPE estimators, and hyperparameters of regression models.
#
# In this step, we define the spaces from which the hyperparameters of OPE estimators / regression models are chosen. (The evaluation policy space is defined in the previous step, and the random state space will be defined in the next step.)
# + id="39YU1CiI8mci" executionInfo={"status": "ok", "timestamp": 1633531029293, "user_tz": -330, "elapsed": 8, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
# set hyperparameter space for ope estimators
# set hyperparameter space for the doubly robust with shrinkage estimator
# with the following code, lambda_ will be chosen from a logarithm uniform distribution over the interval [0.001, 1000]
lambda_ = {
"lower": 1e-3,
"upper": 1e3,
"log": True,
"type": float
}
dros_param = {"lambda_": lambda_}
# + id="MghLFZFN8mck" executionInfo={"status": "ok", "timestamp": 1633531029294, "user_tz": -330, "elapsed": 8, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
# set hyperparameter space for regression models
# set hyperparameter space for logistic regression
# with the following code, C will be chosen from a logarithm uniform distribution over the interval [0.001, 100]
C = {
"lower": 1e-3,
"upper": 1e2,
"log": True,
"type": float
}
# create a dictionary mapping hyperparamter names to hyperparamter spaces
logistic_regression_param = {"C": C}
# set hyperparameter space for the random forest classifier
# with the following code, n_estimators will be chosen from a logarithm uniform distribution over the interval [50, 100]
# the chosen value will be of type int
n_estimators = {
"lower": 5e1,
"upper": 1e2,
"log": True,
"type": int
}
# with the following code, max_depth will be chosen from a uniform distribution over the interval [2, 10]
# the chosen value will be of type int
max_depth = {
"lower": 2,
"upper": 10,
"log": False,
"type": int
}
# with the following code, min_samples_split will be chosen from a uniform distribution over the interval [2, 10]
# the chosen value will be of type int
min_samples_split = {
"lower": 2,
"upper": 10,
"log": False,
"type": int
}
# create a dictionary mapping hyperparamter names to hyperparamter spaces
random_forest_param = {
"n_estimators": n_estimators,
"max_depth": max_depth,
"min_samples_split": min_samples_split
}
# + [markdown] id="as9sJoQ18mck"
# ### Interpretable Evaluation for Off-Policy Evaluation
#
# With the above steps completed, we can finally conduct IEOE by utilizing the `InterpretableOPEEvaluator` class.
#
# Here is a brief description for each parameter that can be passed into `InterpretableOPEEvaluator`:
#
# - `random_states`: a list of integers representing the random_state used when performing OPE; corresponds to the number of iterations
# - `bandit_feedback`: a list of logged bandit feedback data
# - `evaluation_policies`: a list of tuples representing (ground truth policy value, action distribution)
# - `ope_estimators`: a list of OPE ope_estimators
# - `ope_estimator_hyperparams`: a dictionary mapping OPE estimator names to OPE estimator hyperparameter spaces defined in step 2
# - `regression_models`: a list of regression_models
# - `regression_model_hyperparams`: a dictionary mapping regression models to regression model hyperparameter spaces defined in step 2
# + id="9zS8aK0j8mcl" executionInfo={"status": "ok", "timestamp": 1633531030670, "user_tz": -330, "elapsed": 6, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
# initializing class
evaluator = InterpretableOPEEvaluator(
random_states=np.arange(1000),
bandit_feedbacks=[bandit_feedback_test],
evaluation_policies=[
(ground_truth_a, action_dist_a),
(ground_truth_b, action_dist_b)
],
ope_estimators=[
DirectMethod(),
DoublyRobust(),
DoublyRobustWithShrinkage(),
InverseProbabilityWeighting(),
],
ope_estimator_hyperparams={
DoublyRobustWithShrinkage.estimator_name: dros_param,
},
regression_models=[
LogisticRegression,
RandomForest
],
regression_model_hyperparams={
LogisticRegression: logistic_regression_param,
RandomForest: random_forest_param
}
)
# + [markdown] id="MJi8Sk2y8mcm"
# We can set the hyperparameters of OPE estimators / regression models after initializing `InterpretableOPEEvaluator` as well. Below is an example:
# + id="aNK67ojt8mcm" executionInfo={"status": "ok", "timestamp": 1633531031323, "user_tz": -330, "elapsed": 3, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
# re-set hyperparameter space for the doubly robust with shrinkage estimator
# with the following code, lambda_ will now be chosen from a logarithm uniform distribution over the interval [0.001, 100]
evaluator.set_ope_estimator_hyperparam_space(
DoublyRobustWithShrinkage.estimator_name,
param_name="lambda_",
lower=1e-3,
upper=1e2,
log=True,
type_=float,
)
# re-set hyperparameter space for logistic regression
# with the following code, C will now be chosen from a logarithm uniform distribution over the interval [0.001, 100]
evaluator.set_regression_model_hyperparam_space(
LogisticRegression,
param_name="C",
lower=1e-2,
upper=1e2,
log=True,
type_=float,
)
# + [markdown] id="3E7M-Sp88mcn"
# Once we have initialized `InterpretableOPEEvaluator`, we can call implemented methods to perform IEOE.
# + id="sinQVE1H8mcn" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1633531875551, "user_tz": -330, "elapsed": 843510, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="d63812df-ac84-41b4-ea3a-20f395165641"
# estimate policy values
# we obtain a dictionary mapping ope estimator names to np.ndarray storing the estimated policy value for each iteration
policy_value = evaluator.estimate_policy_value()
# + id="_xap440_8mco" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1633531875553, "user_tz": -330, "elapsed": 108, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="4b6176b4-5caf-45d9-b990-8fdb0c1cfa42"
print("dm:", policy_value["dm"][:3])
print("dr:", policy_value["dr"][:3])
print("dr-os:", policy_value["dr-os"][:3])
print("ipw:", policy_value["ipw"][:3])
# + id="N8_NPhB68mco" executionInfo={"status": "ok", "timestamp": 1633531875553, "user_tz": -330, "elapsed": 63, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
# compute squared errors
# we obtain a dictionary mapping ope estimator names to np.ndarray storing the calculated squared error for each iteration
squared_error = evaluator.calculate_squared_error()
# + id="WcqDkW4f8mcp" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1633531875554, "user_tz": -330, "elapsed": 62, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="8c10f4c7-f998-486f-eee4-6c6e203241fa"
print("dm:", squared_error["dm"][:3])
print("dr:", squared_error["dr"][:3])
print("dr-os:", squared_error["dr-os"][:3])
print("ipw:", squared_error["ipw"][:3])
# + id="7JrCAlGP8mcq" colab={"base_uri": "https://localhost:8080/", "height": 432} executionInfo={"status": "ok", "timestamp": 1633531875555, "user_tz": -330, "elapsed": 51, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="b2029a47-41ba-4813-caac-d1bf542e0ac7"
# visualize cdf of squared errors for all ope estimators
evaluator.visualize_cdf_aggregate()
# + id="Y4whQskM8mcr" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1633531875556, "user_tz": -330, "elapsed": 44, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="d50c50ec-01e2-45e3-b804-06147bc4cfd9"
# compute the au-cdf score (area under cdf of squared error over interval [0, thershold]), higher score is better
# we obtain a dictionary mapping ope estimator names to cvar scores
au_cdf = evaluator.calculate_au_cdf_score(threshold=0.0004)
au_cdf
# + id="mrJDfPla8mcs" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1633531876454, "user_tz": -330, "elapsed": 928, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="6f0236fa-36d2-4ab0-9883-5b173458a247"
# by activating the `scale` option,
# we obtain the au_cdf scores where the highest score is scaled to 1
au_cdf_scaled = evaluator.calculate_au_cdf_score(threshold=0.0004, scale=True)
au_cdf_scaled
# + id="QEvejCKy8mcs" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1633531876456, "user_tz": -330, "elapsed": 39, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="320351c3-8e28-465e-bbaf-dfaf818be8b2"
# compute the cvar score (expected value of squared error above probability alpha), lower score is better
# we obtain a dictionary mapping ope estimator names to cvar scores
cvar = evaluator.calculate_cvar_score(alpha=90)
cvar
# + id="G4NXJqVA8mct" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1633531876457, "user_tz": -330, "elapsed": 27, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="053bb41d-9725-49e5-f9bb-39dcce530fb5"
# by activating the `scale` option,
# we obtain the cvar scores where the lowest score is scaled to 1
cvar_scaled = evaluator.calculate_cvar_score(alpha=90, scale=True)
cvar_scaled
# + [markdown] id="skeoq1N28mct"
# ## Example 2 - Synthetic P-Score
# + [markdown] id="e1gRO7rYEY_L"
# A quickstart guide of pyIEOE using synthetic logged bandit feedback data and using estimated propensity scores of the behavior policy instead of the ground truth values.
# + [markdown] id="AmhBtw3nDu6d"
# This section demonstrates an example of conducting Interpretable Evaluation for Off-Policy Evaluation (IEOE). We use synthetic logged bandit feedback data generated using [`obp`](https://github.com/st-tech/zr-obp) and evaluate the performance of Direct Method (DM), Doubly Robust (DR), Doubly Robust with Shrinkage (DRos), and Inverse Probability Weighting (IPW).
#
# Our example contains the following three major steps:
#
# 1. Data Preparation
# 2. Setting Hyperparameter Spaces for Off-Policy Evaluation
# 3. Interpretable Evaluation for Off-Policy Evaluation
# + [markdown] id="4z-SwDXnDu6q"
# ### Data Preparation
#
# In order to conduct IEOE using `pyieoe`, we need to prepare logged bandit feedback data, action distributions of evaluation policies, and ground truth policy values of evaluation policies. Because `pyieoe` is built with the intention of being used with `obp`, these inputs must follow the conventions in `obp`. Specifically, logged bandit feedback data must be of type `BanditFeedback`, action distributions must be of type `np.ndarray`, and ground truth policy values must be of type `float` (or `int`).
#
# In this example, we generate synthetic logged bandit feedback data and perform off-policy learning to obtain two sets of evaluation policies along with their action distributions and ground truth policy values using `obp`. For a detailed explanation of this process, please refer to the [official obp example](https://github.com/st-tech/zr-obp/blob/master/examples/quickstart/quickstart_synthetic.ipynb).
# + id="jx284y0CDu6t" executionInfo={"status": "ok", "timestamp": 1633531907767, "user_tz": -330, "elapsed": 1147, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
# generate a synthetic bandit dataset with 10 actions
# we use `logistic function` as the reward function and `linear_behavior_policy` as the behavior policy.
# one can define their own reward function and behavior policy such as nonlinear ones.
dataset = SyntheticBanditDataset(
n_actions=10,
dim_context=5,
reward_type="binary", # "binary" or "continuous"
reward_function=logistic_reward_function,
behavior_policy_function=linear_behavior_policy,
random_state=12345
)
# obtain training and test sets of synthetic logged bandit feedback
n_rounds_train, n_rounds_test = 10000, 10000
bandit_feedback_train = dataset.obtain_batch_bandit_feedback(n_rounds=n_rounds_train)
bandit_feedback_test = dataset.obtain_batch_bandit_feedback(n_rounds=n_rounds_test)
# define IPWLearner with Logistic Regression as its base ML model
evaluation_policy_a = IPWLearner(
n_actions=dataset.n_actions,
len_list=dataset.len_list,
base_classifier=LogisticRegression(C=100, random_state=12345)
)
# train IPWLearner on the training set of the synthetic logged bandit feedback
evaluation_policy_a.fit(
context=bandit_feedback_train["context"],
action=bandit_feedback_train["action"],
reward=bandit_feedback_train["reward"],
pscore=bandit_feedback_train["pscore"]
)
# obtains action choice probabilities for the test set of the synthetic logged bandit feedback
action_dist_a = evaluation_policy_a.predict_proba(
context=bandit_feedback_test["context"],
tau=0.1 # temperature hyperparameter
)
# define IPWLearner with Random Forest as its base ML model
evaluation_policy_b = IPWLearner(
n_actions=dataset.n_actions,
len_list=dataset.len_list,
base_classifier=RandomForest(n_estimators=30, min_samples_leaf=10, random_state=12345)
)
# train IPWLearner on the training set of the synthetic logged bandit feedback
evaluation_policy_b.fit(
context=bandit_feedback_train["context"],
action=bandit_feedback_train["action"],
reward=bandit_feedback_train["reward"],
pscore=bandit_feedback_train["pscore"]
)
# obtains action choice probabilities for the test set of the synthetic logged bandit feedback
action_dist_b = evaluation_policy_b.predict_proba(
context=bandit_feedback_test["context"],
tau=0.1 # temperature hyperparameter
)
# obtain ground truth policy value for each action choice probabilities
expected_rewards = bandit_feedback_test["expected_reward"]
ground_truth_a = np.average(expected_rewards, weights=action_dist_a[:, :, 0], axis=1).mean()
ground_truth_b = np.average(expected_rewards, weights=action_dist_b[:, :, 0], axis=1).mean()
# + [markdown] id="oKC-lDodDu60"
# ### Setting Hyperparameter Spaces for Off-Policy Evaluation
#
# An integral aspect of IEOE is the different sources of variance. The main sources of variance are evaluation policies, random states, hyperparameters of OPE estimators, and hyperparameters of regression models.
#
# In this step, we define the spaces from which the hyperparameters of OPE estimators / regression models are chosen. (The evaluation policy space is defined in the previous step, and the random state space will be defined in the next step.)
# + id="c2jGLE-3Du64" executionInfo={"status": "ok", "timestamp": 1633531909694, "user_tz": -330, "elapsed": 6, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
# set hyperparameter space for ope estimators
# set hyperparameter space for the doubly robust with shrinkage estimator
# with the following code, lambda_ will be chosen from a logarithm uniform distribution over the interval [0.001, 1000]
lambda_ = {
"lower": 1e-3,
"upper": 1e3,
"log": True,
"type": float
}
dros_param = {"lambda_": lambda_}
# + id="siofnHwHDu66" executionInfo={"status": "ok", "timestamp": 1633531911713, "user_tz": -330, "elapsed": 414, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
# set hyperparameter space for regression models
# set hyperparameter space for logistic regression
# with the following code, C will be chosen from a logarithm uniform distribution over the interval [0.001, 100]
C = {
"lower": 1e-3,
"upper": 1e2,
"log": True,
"type": float
}
# create a dictionary mapping hyperparamter names to hyperparamter spaces
logistic_regression_param = {"C": C}
# set hyperparameter space for the random forest classifier
# with the following code, n_estimators will be chosen from a logarithm uniform distribution over the interval [50, 100]
# the chosen value will be of type int
n_estimators = {
"lower": 5e1,
"upper": 1e2,
"log": True,
"type": int
}
# with the following code, max_depth will be chosen from a uniform distribution over the interval [2, 10]
# the chosen value will be of type int
max_depth = {
"lower": 2,
"upper": 10,
"log": False,
"type": int
}
# with the following code, min_samples_split will be chosen from a uniform distribution over the interval [2, 10]
# the chosen value will be of type int
min_samples_split = {
"lower": 2,
"upper": 10,
"log": False,
"type": int
}
# create a dictionary mapping hyperparamter names to hyperparamter spaces
random_forest_param = {
"n_estimators": n_estimators,
"max_depth": max_depth,
"min_samples_split": min_samples_split
}
# + [markdown] id="AqoaVgxHDu68"
# ### Interpretable Evaluation for Off-Policy Evaluation
#
# With the above steps completed, we can finally conduct IEOE by utilizing the `InterpretableOPEEvaluator` class.
#
# Here is a brief description for each parameter that can be passed into `InterpretableOPEEvaluator`:
#
# - `random_states`: a list of integers representing the random_state used when performing OPE; corresponds to the number of iterations
# - `bandit_feedback`: a list of logged bandit feedback data
# - `evaluation_policies`: a list of tuples representing (ground truth policy value, action distribution)
# - `ope_estimators`: a list of OPE ope_estimators
# - `ope_estimator_hyperparams`: a dictionary mapping OPE estimator names to OPE estimator hyperparameter spaces defined in step 2
# - `regression_models`: a list of regression_models
# - `regression_model_hyperparams`: a dictionary mapping regression models to regression model hyperparameter spaces defined in step 2
# + id="blW97KoKDu6-" executionInfo={"status": "ok", "timestamp": 1633531914852, "user_tz": -330, "elapsed": 442, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
# initializing class
evaluator = InterpretableOPEEvaluator(
random_states=np.arange(1000),
bandit_feedbacks=[bandit_feedback_test],
evaluation_policies=[
(ground_truth_a, action_dist_a),
(ground_truth_b, action_dist_b)
],
ope_estimators=[
DirectMethod(),
DoublyRobust(),
DoublyRobustWithShrinkage(),
InverseProbabilityWeighting(),
],
ope_estimator_hyperparams={
DoublyRobustWithShrinkage.estimator_name: dros_param,
},
regression_models=[
LogisticRegression,
RandomForest
],
regression_model_hyperparams={
LogisticRegression: logistic_regression_param,
RandomForest: random_forest_param
},
pscore_estimators=[
LogisticRegression,
RandomForest
],
pscore_estimator_hyperparams={
LogisticRegression: logistic_regression_param,
RandomForest: random_forest_param
}
)
# + [markdown] id="i6cfcKBvDu7A"
# We can set the hyperparameters of OPE estimators / regression models after initializing `InterpretableOPEEvaluator` as well. Below is an example:
# + id="nk6WSm2BDu7B" executionInfo={"status": "ok", "timestamp": 1633531917054, "user_tz": -330, "elapsed": 454, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
# re-set hyperparameter space for the doubly robust with shrinkage estimator
# with the following code, lambda_ will now be chosen from a logarithm uniform distribution over the interval [0.001, 100]
evaluator.set_ope_estimator_hyperparam_space(
DoublyRobustWithShrinkage.estimator_name,
param_name="lambda_",
lower=1e-3,
upper=1e2,
log=True,
type_=float,
)
# re-set hyperparameter space for logistic regression
# with the following code, C will now be chosen from a logarithm uniform distribution over the interval [0.001, 100]
evaluator.set_regression_model_hyperparam_space(
LogisticRegression,
param_name="C",
lower=1e-2,
upper=1e2,
log=True,
type_=float,
)
# + [markdown] id="Hk4SExejDu7C"
# Once we have initialized `InterpretableOPEEvaluator`, we can call implemented methods to perform IEOE.
# + id="P7iKs11nDu7D" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1633533386212, "user_tz": -330, "elapsed": 1467183, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="b653b918-70fc-49a7-8f4d-aefaa4fb5797"
# estimate policy values
# we obtain a dictionary mapping ope estimator names to np.ndarray storing the estimated policy value for each iteration
policy_value = evaluator.estimate_policy_value()
# + id="7Z_welpvDu7F" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1633533386213, "user_tz": -330, "elapsed": 32, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="19682ffa-e010-4e08-e0e1-9ab069d65abf"
print("dm:", policy_value["dm"][:3])
print("dr:", policy_value["dr"][:3])
print("dr-os:", policy_value["dr-os"][:3])
print("ipw:", policy_value["ipw"][:3])
# + id="0ujRjvG5Du7G" executionInfo={"status": "ok", "timestamp": 1633533403052, "user_tz": -330, "elapsed": 1379, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}}
# compute squared errors
# we obtain a dictionary mapping ope estimator names to np.ndarray storing the calculated squared error for each iteration
squared_error = evaluator.calculate_squared_error()
# + id="vP_vo9hcDu7H" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1633533405765, "user_tz": -330, "elapsed": 21, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="2edba37d-678c-4f5c-b168-cf943f4e3441"
print("dm:", squared_error["dm"][:3])
print("dr:", squared_error["dr"][:3])
print("dr-os:", squared_error["dr-os"][:3])
print("ipw:", squared_error["ipw"][:3])
# + id="SZbxYSCjDu7H" colab={"base_uri": "https://localhost:8080/", "height": 432} executionInfo={"status": "ok", "timestamp": 1633533409081, "user_tz": -330, "elapsed": 1090, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="5c514cab-442c-4b37-d22d-27c3a6eb1363"
# visualize cdf of squared errors for all ope estimators
evaluator.visualize_cdf_aggregate(xmax=0.002)
# + id="36naCb1EDu7I" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1633533412696, "user_tz": -330, "elapsed": 620, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="de522526-3696-4abc-d458-11e226b41693"
# compute the au-cdf score (area under cdf of squared error over interval [0, thershold]), higher score is better
# we obtain a dictionary mapping ope estimator names to cvar scores
au_cdf = evaluator.calculate_au_cdf_score(threshold=0.0004)
au_cdf
# + id="09pr3_UuDu7J" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1633533414676, "user_tz": -330, "elapsed": 21, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="91bbe2ff-ef6f-4bc9-c6eb-46c03c075feb"
# by activating the `scale` option,
# we obtain the au_cdf scores where the highest score is scaled to 1
au_cdf_scaled = evaluator.calculate_au_cdf_score(threshold=0.0004, scale=True)
au_cdf_scaled
# + id="787ebELZDu7J" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1633533416444, "user_tz": -330, "elapsed": 17, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="42e97017-91d9-4e6c-f109-e09f7578215a"
# compute the cvar score (expected value of squared error above probability alpha), lower score is better
# we obtain a dictionary mapping ope estimator names to cvar scores
cvar = evaluator.calculate_cvar_score(alpha=90)
cvar
# + id="b2GuPiBiDu7K" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1633533418752, "user_tz": -330, "elapsed": 18, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13037694610922482904"}} outputId="aca49a97-6dce-4b3c-f618-6da009d69d9e"
# by activating the `scale` option,
# we obtain the cvar scores where the lowest score is scaled to 1
cvar_scaled = evaluator.calculate_cvar_score(alpha=90, scale=True)
cvar_scaled
# + [markdown] id="M91PB3GDDu7O"
# ## Example 3 - Synthetic RSCV
# + [markdown] id="f35iP_S8FB0C"
# A quickstart guide of pyIEOE using synthetic logged bandit feedback data and using RandomizedSearchCV for regression models and pscore estimators.
# + [markdown] id="uwJs5p_NE8Kh"
# ### Data Preparation
#
# In order to conduct IEOE using `pyieoe`, we need to prepare logged bandit feedback data, action distributions of evaluation policies, and ground truth policy values of evaluation policies. Because `pyieoe` is built with the intention of being used with `obp`, these inputs must follow the conventions in `obp`. Specifically, logged bandit feedback data must be of type `BanditFeedback`, action distributions must be of type `np.ndarray`, and ground truth policy values must be of type `float` (or `int`).
#
# In this example, we generate synthetic logged bandit feedback data and perform off-policy learning to obtain two sets of evaluation policies along with their action distributions and ground truth policy values using `obp`. For a detailed explanation of this process, please refer to the [official obp example](https://github.com/st-tech/zr-obp/blob/master/examples/quickstart/quickstart_synthetic.ipynb).
# + id="hiKvoK-cE8Ki"
# generate a synthetic bandit dataset with 10 actions
# we use `logistic function` as the reward function and `linear_behavior_policy` as the behavior policy.
# one can define their own reward function and behavior policy such as nonlinear ones.
dataset = SyntheticBanditDataset(
n_actions=10,
dim_context=5,
reward_type="binary", # "binary" or "continuous"
reward_function=logistic_reward_function,
behavior_policy_function=linear_behavior_policy,
random_state=12345
)
# obtain training and test sets of synthetic logged bandit feedback
n_rounds_train, n_rounds_test = 10000, 10000
bandit_feedback_train = dataset.obtain_batch_bandit_feedback(n_rounds=n_rounds_train)
bandit_feedback_test = dataset.obtain_batch_bandit_feedback(n_rounds=n_rounds_test)
# define IPWLearner with Logistic Regression as its base ML model
evaluation_policy_a = IPWLearner(
n_actions=dataset.n_actions,
len_list=dataset.len_list,
base_classifier=LogisticRegression(C=100, random_state=12345)
)
# train IPWLearner on the training set of the synthetic logged bandit feedback
evaluation_policy_a.fit(
context=bandit_feedback_train["context"],
action=bandit_feedback_train["action"],
reward=bandit_feedback_train["reward"],
pscore=bandit_feedback_train["pscore"]
)
# obtains action choice probabilities for the test set of the synthetic logged bandit feedback
action_dist_a = evaluation_policy_a.predict_proba(
context=bandit_feedback_test["context"],
tau=0.1 # temperature hyperparameter
)
# define IPWLearner with Random Forest as its base ML model
evaluation_policy_b = IPWLearner(
n_actions=dataset.n_actions,
len_list=dataset.len_list,
base_classifier=RandomForest(n_estimators=30, min_samples_leaf=10, random_state=12345)
)
# train IPWLearner on the training set of the synthetic logged bandit feedback
evaluation_policy_b.fit(
context=bandit_feedback_train["context"],
action=bandit_feedback_train["action"],
reward=bandit_feedback_train["reward"],
pscore=bandit_feedback_train["pscore"]
)
# obtains action choice probabilities for the test set of the synthetic logged bandit feedback
action_dist_b = evaluation_policy_b.predict_proba(
context=bandit_feedback_test["context"],
tau=0.1 # temperature hyperparameter
)
# obtain ground truth policy value for each action choice probabilities
expected_rewards = bandit_feedback_test["expected_reward"]
ground_truth_a = np.average(expected_rewards, weights=action_dist_a[:, :, 0], axis=1).mean()
ground_truth_b = np.average(expected_rewards, weights=action_dist_b[:, :, 0], axis=1).mean()
# + [markdown] id="Gz7FkzuFE8Kl"
# ### Setting Hyperparameter Spaces for Off-Policy Evaluation
#
# An integral aspect of IEOE is the different sources of variance. The main sources of variance are evaluation policies, random states, hyperparameters of OPE estimators, and hyperparameters of regression models.
#
# In this step, we define the spaces from which the hyperparameters of OPE estimators / regression models are chosen. (The evaluation policy space is defined in the previous step, and the random state space will be defined in the next step.)
# + id="NVH2dZzYE8Km"
# set hyperparameter space for ope estimators
# set hyperparameter space for the doubly robust with shrinkage estimator
# with the following code, lambda_ will be chosen from a logarithm uniform distribution over the interval [0.001, 1000]
lambda_ = {
"lower": 1e-3,
"upper": 1e3,
"log": True,
"type": float
}
dros_param = {"lambda_": lambda_}
# + id="auB7GcSYE8Kn"
# set hyperparameter space for logistic regression using RandomizedSearchCV
from sklearn.utils.fixes import loguniform
logistic = LogisticRegression()
distributions = {
"C": loguniform(1e-2, 1e2)
}
clf_logistic = RandomizedSearchCV(logistic, distributions, random_state=0, n_iter=5)
# + id="H20quziGE8Ko"
# set hyperparameter space for random forest classifier using RandomizedSearchCV
from scipy.stats import randint
randforest = RandomForest()
distributions = {
# n_estimators will be chosen from a uniform distribution over the interval [50, 100)
"n_estimators": randint(5e1, 1e2),
# max_depth will be chosen from a uniform distribution over the interval [2, 10)
"max_depth": randint(2, 10),
# min_samples_split will be chosen from a uniform distribution over the interval [2, 10)
"min_samples_split": randint(2, 10)
}
clf_randforest = RandomizedSearchCV(randforest, distributions, random_state=0, n_iter=5)
# + [markdown] id="YhIFJTDdE8Ko"
# ### Interpretable Evaluation for Off-Policy Evaluation
#
# With the above steps completed, we can finally conduct IEOE by utilizing the `InterpretableOPEEvaluator` class.
#
# Here is a brief description for each parameter that can be passed into `InterpretableOPEEvaluator`:
#
# - `random_states`: a list of integers representing the random_state used when performing OPE; corresponds to the number of iterations
# - `bandit_feedback`: a list of logged bandit feedback data
# - `evaluation_policies`: a list of tuples representing (ground truth policy value, action distribution)
# - `ope_estimators`: a list of OPE ope_estimators
# - `ope_estimator_hyperparams`: a dictionary mapping OPE estimator names to OPE estimator hyperparameter spaces defined in step 2
# - `regression_models`: a list of regression_models
# - `regression_model_hyperparams`: a dictionary mapping regression models to regression model hyperparameter spaces defined in step 2
# + id="favcoQnZE8Kp"
# initializing class
evaluator = InterpretableOPEEvaluator(
random_states=np.arange(100),
bandit_feedbacks=[bandit_feedback_test],
evaluation_policies=[
(ground_truth_a, action_dist_a),
(ground_truth_b, action_dist_b)
],
ope_estimators=[
DirectMethod(),
DoublyRobust(),
DoublyRobustWithShrinkage(),
InverseProbabilityWeighting(),
],
ope_estimator_hyperparams={
DoublyRobustWithShrinkage.estimator_name: dros_param,
},
regression_models=[
clf_logistic,
clf_randforest
],
pscore_estimators=[
clf_logistic,
clf_randforest
]
)
# + [markdown] id="aZ_pxjj-E8Kp"
# Once we have initialized `InterpretableOPEEvaluator`, we can call implemented methods to perform IEOE.
# + id="-WMc3JMAE8Kq" outputId="6e0d2868-c3a5-4db0-bff3-e8123a46934a"
# estimate policy values
# we obtain a dictionary mapping ope estimator names to np.ndarray storing the estimated policy value for each iteration
policy_value = evaluator.estimate_policy_value()
# + id="NV5s_KQNE8Kr" outputId="d9479f4e-67de-4ebe-ed30-52246595e4be"
print("dm:", policy_value["dm"][:3])
print("dr:", policy_value["dr"][:3])
print("dr-os:", policy_value["dr-os"][:3])
print("ipw:", policy_value["ipw"][:3])
# + id="2YpdxF85E8Kr"
# compute squared errors
# we obtain a dictionary mapping ope estimator names to np.ndarray storing the calculated squared error for each iteration
squared_error = evaluator.calculate_squared_error()
# + id="dZbd-eVbE8Ks" outputId="46f571d5-d382-45aa-c011-62a6c2f59251"
print("dm:", squared_error["dm"][:3])
print("dr:", squared_error["dr"][:3])
print("dr-os:", squared_error["dr-os"][:3])
print("ipw:", squared_error["ipw"][:3])
# + id="TAzYFpRRE8Ks" outputId="b1679b55-9f65-45f8-b927-00eaece7a139"
# visualize cdf of squared errors for all ope estimators
evaluator.visualize_cdf_aggregate(xmax=0.002)
# + id="NxYS2lquE8Kt" outputId="af7834ca-acf9-434b-9f44-48468e567960"
# compute the au-cdf score (area under cdf of squared error over interval [0, thershold]), higher score is better
# we obtain a dictionary mapping ope estimator names to cvar scores
au_cdf = evaluator.calculate_au_cdf_score(threshold=0.0004)
au_cdf
# + id="pGJMvH_vE8Ku" outputId="274f7992-a883-4030-eab3-ae0cead1e172"
# by activating the `scale` option,
# we obtain the au_cdf scores where the highest score is scaled to 1
au_cdf_scaled = evaluator.calculate_au_cdf_score(threshold=0.0004, scale=True)
au_cdf_scaled
# + id="064QHL4xE8Kv" outputId="e00e4aef-bc51-4790-896f-6407535169b9"
# compute the cvar score (expected value of squared error above probability alpha), lower score is better
# we obtain a dictionary mapping ope estimator names to cvar scores
cvar = evaluator.calculate_cvar_score(alpha=90)
cvar
# + id="vAAvr2xbE8Kv" outputId="f65d2728-b8db-4053-fcef-82f30c711d9f"
# by activating the `scale` option,
# we obtain the cvar scores where the lowest score is scaled to 1
cvar_scaled = evaluator.calculate_cvar_score(alpha=90, scale=True)
cvar_scaled
# + [markdown] id="vfgNKQaTF3f0"
# ## Example 4 - Multiclass dataset
# + [markdown] id="prn-5aqKF58Z"
# A quickstart guide of pyIEOE using multiclass classification data as logged bandit feedback data.
# + [markdown] id="v_FGRdjYGiJn"
# This section demonstrates an example of conducting Interpretable Evaluation for Off-Policy Evaluation (IEOE). We use logged bandit feedback data generated by modifying multiclass classification data using [`obp`](https://github.com/st-tech/zr-obp) and evaluate the performance of Direct Method (DM), Doubly Robust (DR), Doubly Robust with Shrinkage (DRos), and Inverse Probability Weighting (IPW).
#
# Our example contains the following three major steps:
#
# 1. Data Preparation
# 2. Setting Hyperparameter Spaces for Off-Policy Evaluation
# 3. Interpretable Evaluation for Off-Policy Evaluation
# + [markdown] id="PYJ2uM8bGe5L"
# ### Data Preparation
#
# In order to conduct IEOE using `pyieoe`, we need to prepare logged bandit feedback data, action distributions of evaluation policies, and ground truth policy values of evaluation policies. Because `pyieoe` is built with the intention of being used with `obp`, these inputs must follow the conventions in `obp`. Specifically, logged bandit feedback data must be of type `BanditFeedback`, action distributions must be of type `np.ndarray`, and ground truth policy values must be of type `float` (or `int`).
#
# In this example, we generate logged bandit feedback data by modifying multiclass classification data and obtain two sets of evaluation policies along with their action distributions and ground truth policy values using `obp`. For a detailed explanation of this process, please refer to the [official docs](https://zr-obp.readthedocs.io/en/latest/_autosummary/obp.dataset.multiclass.html#module-obp.dataset.multiclass).
# + id="BlxiQlSqGe5M"
# load raw digits data
X, y = load_digits(return_X_y=True)
# convert the raw classification data into the logged bandit dataset
dataset = MultiClassToBanditReduction(
X=X,
y=y,
base_classifier_b=LogisticRegression(random_state=12345),
alpha_b=0.8,
dataset_name="digits"
)
# split the original data into the training and evaluation sets
dataset.split_train_eval(eval_size=0.7, random_state=12345)
# obtain logged bandit feedback generated by the behavior policy
bandit_feedback = dataset.obtain_batch_bandit_feedback(random_state=12345)
# obtain action choice probabilities by an evaluation policy and its ground-truth policy value
action_dist_a = dataset.obtain_action_dist_by_eval_policy(
base_classifier_e=LogisticRegression(C=100, random_state=12345),
alpha_e=0.9
)
ground_truth_a = dataset.calc_ground_truth_policy_value(action_dist=action_dist_a)
action_dist_b = dataset.obtain_action_dist_by_eval_policy(
base_classifier_e=RandomForest(n_estimators=100, min_samples_split=5, random_state=12345),
alpha_e=0.9
)
ground_truth_b = dataset.calc_ground_truth_policy_value(action_dist=action_dist_b)
# + [markdown] id="D1WCrmkSGe5M"
# ### Setting Hyperparameter Spaces for Off-Policy Evaluation
#
# An integral aspect of IEOE is the different sources of variance. The main sources of variance are evaluation policies, random states, hyperparameters of OPE estimators, and hyperparameters of regression models.
#
# In this step, we define the spaces from which the hyperparameters of OPE estimators / regression models are chosen. (The evaluation policy space is defined in the previous step, and the random state space will be defined in the next step.)
# + id="1qeIQd79Ge5N"
# set hyperparameter space for ope estimators
# set hyperparameter space for the doubly robust with shrinkage estimator
# with the following code, lambda_ will be chosen from a logarithm uniform distribution over the interval [0.001, 1000]
lambda_ = {
"lower": 1e-3,
"upper": 1e3,
"log": True,
"type": float
}
dros_param = {"lambda_": lambda_}
# + id="LOpaUfGYGe5N"
# set hyperparameter space for regression models
# set hyperparameter space for logistic regression
# with the following code, C will be chosen from a logarithm uniform distribution over the interval [0.001, 100]
C = {
"lower": 1e-3,
"upper": 1e2,
"log": True,
"type": float
}
# with the following code, max_iter will be fixed at 10000 and of type int
max_iter = {
"lower": 1e4,
"upper": 1e4,
"log": False,
"type": int
}
# create a dictionary mapping hyperparamter names to hyperparamter spaces
logistic_regression_param = {
"C": C,
"max_iter": max_iter
}
# set hyperparameter space for random forest classifier
# with the following code, n_estimators will be chosen from a logarithm uniform distribution over the interval [50, 100]
# the chosen value will be of type int
n_estimators = {
"lower": 5e1,
"upper": 1e2,
"log": True,
"type": int
}
# with the following code, max_depth will be chosen from a uniform distribution over the interval [2, 10]
# the chosen value will be of type int
max_depth = {
"lower": 2,
"upper": 10,
"log": False,
"type": int
}
# with the following code, min_samples_split will be chosen from a uniform distribution over the interval [2, 10]
# the chosen value will be of type int
min_samples_split = {
"lower": 2,
"upper": 10,
"log": False,
"type": int
}
# create a dictionary mapping hyperparamter names to hyperparamter spaces
random_forest_param = {
"n_estimators": n_estimators,
"max_depth": max_depth,
"min_samples_split": min_samples_split
}
# + [markdown] id="jLOs3fUrGe5O"
# ### Interpretable Evaluation for Off-Policy Evaluation
#
# With the above steps completed, we can finally conduct IEOE by utilizing the `InterpretableOPEEvaluator` class.
#
# Here is a brief description for each parameter that can be passed into `InterpretableOPEEvaluator`:
#
# - `random_states`: a list of integers representing the random_state used when performing OPE; corresponds to the number of iterations
# - `bandit_feedback`: a list of logged bandit feedback data
# - `evaluation_policies`: a list of tuples representing (ground truth policy value, action distribution)
# - `ope_estimators`: a list of OPE ope_estimators
# - `ope_estimator_hyperparams`: a dictionary mapping OPE estimator names to OPE estimator hyperparameter spaces defined in step 2
# - `regression_models`: a list of regression regression_models
# - `regression_model_hyperparams`: a dictionary mapping regression models to regression model hyperparameter spaces defined in step 2
# + id="RpAqLCmUGe5R"
# initializing class
evaluator = InterpretableOPEEvaluator(
random_states=np.arange(1000),
bandit_feedbacks=[bandit_feedback],
evaluation_policies=[
(ground_truth_a, action_dist_a),
(ground_truth_b, action_dist_b)
],
ope_estimators=[
DirectMethod(),
DoublyRobust(),
DoublyRobustWithShrinkage(),
InverseProbabilityWeighting(),
],
ope_estimator_hyperparams={
DoublyRobustWithShrinkage.estimator_name: dros_param,
},
regression_models=[
LogisticRegression,
RandomForest
],
regression_model_hyperparams={
LogisticRegression: logistic_regression_param,
RandomForest: random_forest_param
}
)
# + [markdown] id="28uy9Gh6Ge5R"
# We can set the hyperparameters of OPE estimators / regression models after initializing `InterpretableOPEEvaluator` as well. Below is an example:
# + id="PTyLRuKKGe5S"
# re-set hyperparameter space for doubly robust with shrinkage estimator
# with the following code, lambda_ will now be chosen from a logarithm uniform distribution over the interval [0.001, 100]
evaluator.set_ope_estimator_hyperparam_space(
DoublyRobustWithShrinkage.estimator_name,
param_name="lambda_",
lower=1e-3,
upper=1e2,
log=True,
type_=float,
)
# re-set hyperparameter space for logistic regression
# with the following code, C will now be chosen from a logarithm uniform distribution over the interval [0.001, 100]
evaluator.set_regression_model_hyperparam_space(
LogisticRegression,
param_name="C",
lower=1e-2,
upper=1e2,
log=True,
type_=float,
)
# + [markdown] id="UEnmiLzmGe5S"
# Once we have initialized `InterpretableOPEEvaluator`, we can call implemented methods to perform IEOE.
# + id="XzZGsJyJGe5T" outputId="4359a6d9-05ab-4985-eddb-cf230d39d7d4"
# estimate policy values
# we obtain a dictionary mapping ope estimator names to np.ndarray storing the estimated policy value for each iteration
policy_value = evaluator.estimate_policy_value()
# + id="05QPkmrRGe5T" outputId="518cf6f5-ca32-42c5-f9b8-e8eac321614f"
print("dm:", policy_value["dm"][:3])
print("dr:", policy_value["dr"][:3])
print("dr-os:", policy_value["dr-os"][:3])
print("ipw:", policy_value["ipw"][:3])
# + id="LR8geYTkGe5U"
# compute squared errors
# we obtain a dictionary mapping ope estimator names to np.ndarray storing the calculated squared error for each iteration
squared_error = evaluator.calculate_squared_error()
# + id="fWQUMjXlGe5U" outputId="b67a1f8c-2077-4bed-9b9d-cd3413d18fc8"
print("dm:", squared_error["dm"][:3])
print("dr:", squared_error["dr"][:3])
print("dr-os:", squared_error["dr-os"][:3])
print("ipw:", squared_error["ipw"][:3])
# + id="vFxyRkNDGe5V" outputId="a55e6f4c-58ca-4c0e-ec2a-606075da3257"
# visualize cdf of squared errors for all ope estimators
evaluator.visualize_cdf_aggregate()
# + id="KzfGNZ-sGe5V" outputId="1a8f4342-c6fe-4758-fc3c-859bf51d99a0"
# compute the au-cdf score (area under cdf of squared error over interval [0, thershold]), higher score is better
# we obtain a dictionary mapping ope estimator names to cvar scores
au_cdf = evaluator.calculate_au_cdf_score(threshold=0.004)
au_cdf
# + id="kZgtGa7wGe5W" outputId="9a3b80f6-bc89-44c4-f669-1a19fbdeed05"
# by activating the `scale` option,
# we obtain au_cdf scores where the highest score is scaled to 1
au_cdf_scaled = evaluator.calculate_au_cdf_score(threshold=0.004, scale=True)
au_cdf_scaled
# + id="wrNEepNHGe5X" outputId="6b5755f8-d508-4a90-f334-0dc8f5ad28e1"
# compute the cvar score (expected value of squared error above probability alpha), lower score is better
# we obtain a dictionary mapping ope estimator names to cvar scores
cvar = evaluator.calculate_cvar_score(alpha=90)
cvar
# + id="JnFZf7znGe5c" outputId="813a83cc-5b7f-447c-d846-ccfa9be56539"
# by activating the `scale` option,
# we obtain cvar scores where the lowest score is scaled to 1
cvar_scaled = evaluator.calculate_cvar_score(alpha=90, scale=True)
cvar_scaled
# + [markdown] id="-XznTeoAG_qy"
# ## Example 5 - Multiclass P-Score
# + [markdown] id="9WghWS-xJdEy"
# A quickstart guide of pyIEOE using multiclass classification data and using estimated propensity scores of the behavior policy instead of the ground truth values.
# + [markdown] id="bRJuPPCDG6uA"
# This notebook demonstrates an example of conducting Interpretable Evaluation for Off-Policy Evaluation (IEOE). We use logged bandit feedback data generated by modifying multiclass classification data using [`obp`](https://github.com/st-tech/zr-obp) and evaluate the performance of Direct Method (DM), Doubly Robust (DR), Doubly Robust with Shrinkage (DRos), and Inverse Probability Weighting (IPW).
#
# Our example contains the following three major steps:
#
# 1. Data Preparation
# 2. Setting Hyperparameter Spaces for Off-Policy Evaluation
# 3. Interpretable Evaluation for Off-Policy Evaluation
# + [markdown] id="UDuL-UTxG6uL"
# ### Data Preparation
#
# In order to conduct IEOE using `pyieoe`, we need to prepare logged bandit feedback data, action distributions of evaluation policies, and ground truth policy values of evaluation policies. Because `pyieoe` is built with the intention of being used with `obp`, these inputs must follow the conventions in `obp`. Specifically, logged bandit feedback data must be of type `BanditFeedback`, action distributions must be of type `np.ndarray`, and ground truth policy values must be of type `float` (or `int`).
#
# In this example, we generate logged bandit feedback data by modifying multiclass classification data and obtain two sets of evaluation policies along with their action distributions and ground truth policy values using `obp`. For a detailed explanation of this process, please refer to the [official docs](https://zr-obp.readthedocs.io/en/latest/_autosummary/obp.dataset.multiclass.html#module-obp.dataset.multiclass).
# + id="EgLP5CSjG6uM"
# load raw digits data
X, y = load_digits(return_X_y=True)
# convert the raw classification data into the logged bandit dataset
dataset = MultiClassToBanditReduction(
X=X,
y=y,
base_classifier_b=LogisticRegression(random_state=12345),
alpha_b=0.8,
dataset_name="digits"
)
# split the original data into the training and evaluation sets
dataset.split_train_eval(eval_size=0.7, random_state=12345)
# obtain logged bandit feedback generated by the behavior policy
bandit_feedback = dataset.obtain_batch_bandit_feedback(random_state=12345)
# obtain action choice probabilities by an evaluation policy and its ground-truth policy value
action_dist_a = dataset.obtain_action_dist_by_eval_policy(
base_classifier_e=LogisticRegression(C=100, random_state=12345, max_iter=10000),
alpha_e=0.9
)
ground_truth_a = dataset.calc_ground_truth_policy_value(action_dist=action_dist_a)
action_dist_b = dataset.obtain_action_dist_by_eval_policy(
base_classifier_e=RandomForest(n_estimators=100, min_samples_split=5, random_state=12345),
alpha_e=0.9
)
ground_truth_b = dataset.calc_ground_truth_policy_value(action_dist=action_dist_b)
# + [markdown] id="xBPAdys_G6uP"
# ### Setting Hyperparameter Spaces for Off-Policy Evaluation
#
# An integral aspect of IEOE is the different sources of variance. The main sources of variance are evaluation policies, random states, hyperparameters of OPE estimators, and hyperparameters of regression models.
#
# In this step, we define the spaces from which the hyperparameters of OPE estimators / regression models are chosen. (The evaluation policy space is defined in the previous step, and the random state space will be defined in the next step.)
# + id="nvJYaTFVG6uR"
# set hyperparameter space for ope estimators
# set hyperparameter space for the doubly robust with shrinkage estimator
# with the following code, lambda_ will be chosen from a logarithm uniform distribution over the interval [0.001, 1000]
lambda_ = {
"lower": 1e-3,
"upper": 1e3,
"log": True,
"type": float
}
dros_param = {"lambda_": lambda_}
# + id="Tf--fTxVG6uU"
# set hyperparameter space for regression models
# set hyperparameter space for logistic regression
# with the following code, C will be chosen from a logarithm uniform distribution over the interval [0.001, 100]
# the chosen value will be of type float
C = {
"lower": 1e-3,
"upper": 1e2,
"log": True,
"type": float
}
# with the following code, max_iter will be fixed at 10000 and of type int
max_iter = {
"lower": 1e4,
"upper": 1e4,
"log": False,
"type": int
}
# create a dictionary mapping hyperparamter names to hyperparamter spaces
logistic_regression_param = {
"C": C,
"max_iter": max_iter
}
# set hyperparameter space for random forest classifier
# with the following code, n_estimators will be chosen from a logarithm uniform distribution over the interval [50, 100]
# the chosen value will be of type int
n_estimators = {
"lower": 5e1,
"upper": 1e2,
"log": True,
"type": int
}
# with the following code, max_depth will be chosen from a uniform distribution over the interval [2, 10]
# the chosen value will be of type int
max_depth = {
"lower": 2,
"upper": 10,
"log": False,
"type": int
}
# with the following code, min_samples_split will be chosen from a uniform distribution over the interval [2, 10]
# the chosen value will be of type int
min_samples_split = {
"lower": 2,
"upper": 10,
"log": False,
"type": int
}
# create a dictionary mapping hyperparamter names to hyperparamter spaces
random_forest_param = {
"n_estimators": n_estimators,
"max_depth": max_depth,
"min_samples_split": min_samples_split
}
# + [markdown] id="huLEWqzjG6uW"
# ### Interpretable Evaluation for Off-Policy Evaluation
#
# With the above steps completed, we can finally conduct IEOE by utilizing the `InterpretableOPEEvaluator` class.
#
# Here is a brief description for each parameter that can be passed into `InterpretableOPEEvaluator`:
#
# - `random_states`: a list of integers representing the random_state used when performing OPE; corresponds to the number of iterations
# - `bandit_feedback`: a list of logged bandit feedback data
# - `evaluation_policies`: a list of tuples representing (ground truth policy value, action distribution)
# - `ope_estimators`: a list of OPE ope_estimators
# - `ope_estimator_hyperparams`: a dictionary mapping OPE estimator names to OPE estimator hyperparameter spaces defined in step 2
# - `regression_models`: a list of regression regression_models
# - `regression_model_hyperparams`: a dictionary mapping regression models to regression model hyperparameter spaces defined in step 2
# + id="A8WeqUEWG6uX"
# initializing class
evaluator = InterpretableOPEEvaluator(
random_states=np.arange(1000),
bandit_feedbacks=[bandit_feedback],
evaluation_policies=[
(ground_truth_a, action_dist_a),
(ground_truth_b, action_dist_b)
],
ope_estimators=[
DirectMethod(),
DoublyRobust(),
DoublyRobustWithShrinkage(),
InverseProbabilityWeighting(),
],
ope_estimator_hyperparams={
DoublyRobustWithShrinkage.estimator_name: dros_param,
},
regression_models=[
LogisticRegression,
RandomForest
],
regression_model_hyperparams={
LogisticRegression: logistic_regression_param,
RandomForest: random_forest_param
},
pscore_estimators=[
LogisticRegression,
RandomForest
],
pscore_estimator_hyperparams={
LogisticRegression: logistic_regression_param,
RandomForest: random_forest_param
}
)
# + [markdown] id="EBeRcYo-G6uX"
# We can set the hyperparameters of OPE estimators / regression models after initializing `InterpretableOPEEvaluator` as well. Below is an example:
# + id="bHC2WIxbG6uY"
# re-set hyperparameter space for doubly robust with shrinkage estimator
# with the following code, lambda_ will now be chosen from a logarithm uniform distribution over the interval [0.001, 100]
evaluator.set_ope_estimator_hyperparam_space(
DoublyRobustWithShrinkage.estimator_name,
param_name="lambda_",
lower=1e-3,
upper=1e2,
log=True,
type_=float,
)
# re-set hyperparameter space for logistic regression
# with the following code, C will now be chosen from a logarithm uniform distribution over the interval [0.001, 100]
evaluator.set_regression_model_hyperparam_space(
LogisticRegression,
param_name="C",
lower=1e-2,
upper=1e2,
log=True,
type_=float,
)
# + [markdown] id="GGwQNCVyG6uY"
# Once we have initialized `InterpretableOPEEvaluator`, we can call implemented methods to perform IEOE.
# + id="uNzEIUMWG6uZ" outputId="8c914282-e8c9-4fe3-e627-7b02f154ab29"
# estimate policy values
# we obtain a dictionary mapping ope estimator names to np.ndarray storing the estimated policy value for each iteration
policy_value = evaluator.estimate_policy_value()
# + id="LwUCCvvwG6uZ" outputId="486f4818-1c81-4fc4-e82c-18585cecc42d"
print("dm:", policy_value["dm"][:3])
print("dr:", policy_value["dr"][:3])
print("dr-os:", policy_value["dr-os"][:3])
print("ipw:", policy_value["ipw"][:3])
# + id="0NPL_UYUG6ua"
# compute squared errors
# we obtain a dictionary mapping ope estimator names to np.ndarray storing the calculated squared error for each iteration
squared_error = evaluator.calculate_squared_error()
# + id="-ophfTl6G6ua" outputId="751cb388-ce5b-4d5b-d7ba-c814a8e5669f"
print("dm:", squared_error["dm"][:3])
print("dr:", squared_error["dr"][:3])
print("dr-os:", squared_error["dr-os"][:3])
print("ipw:", squared_error["ipw"][:3])
# + id="pROmONbVG6ua" outputId="c826906d-328c-4f06-8ac7-9188aff7a8d3"
# visualize cdf of squared errors for all ope estimators
evaluator.visualize_cdf_aggregate(xmax=0.04)
# + id="nhNcA5sWG6ug" outputId="dfd454dd-caee-4436-bcc6-bb9a4ad45c07"
# compute the au-cdf score (area under cdf of squared error over interval [0, thershold]), higher score is better
# we obtain a dictionary mapping ope estimator names to cvar scores
au_cdf = evaluator.calculate_au_cdf_score(threshold=0.004)
au_cdf
# + id="zIC_frEzG6uj" outputId="69cadae3-361c-444c-f257-ba98fa7519aa"
# by activating the `scale` option,
# we obtain au_cdf scores where the highest score is scaled to 1
au_cdf_scaled = evaluator.calculate_au_cdf_score(threshold=0.004, scale=True)
au_cdf_scaled
# + id="4ysQqt_OG6up" outputId="299a7247-eb15-417e-8b31-5788a890868f"
# compute the cvar score (expected value of squared error above probability alpha), lower score is better
# we obtain a dictionary mapping ope estimator names to cvar scores
cvar = evaluator.calculate_cvar_score(alpha=90)
cvar
# + id="MD-2jhGtG6ut" outputId="1fb3cdc7-f7e8-464d-f113-abf117b472cd"
# by activating the `scale` option,
# we obtain cvar scores where the lowest score is scaled to 1
cvar_scaled = evaluator.calculate_cvar_score(alpha=90, scale=True)
cvar_scaled
# + [markdown] id="Cl59UFs_G6uu"
# ## Example 6 - Multiclass RSCV
# + [markdown] id="7qGIsmOAJmG_"
# A quickstart guide of pyIEOE using multiclass classification data and using RandomizedSearchCV for regression models and pscore estimators.
# + [markdown] id="wY466oehJqXZ"
# This section demonstrates an example of conducting Interpretable Evaluation for Off-Policy Evaluation (IEOE). We use logged bandit feedback data generated by modifying multiclass classification data using [`obp`](https://github.com/st-tech/zr-obp) and evaluate the performance of Direct Method (DM), Doubly Robust (DR), Doubly Robust with Shrinkage (DRos), and Inverse Probability Weighting (IPW).
#
# Our example contains the following three major steps:
#
# 1. Data Preparation
# 2. Setting Hyperparameter Spaces for Off-Policy Evaluation
# 3. Interpretable Evaluation for Off-Policy Evaluation
# + [markdown] id="Uqy99AyZJqXf"
# ### Data Preparation
#
# In order to conduct IEOE using `pyieoe`, we need to prepare logged bandit feedback data, action distributions of evaluation policies, and ground truth policy values of evaluation policies. Because `pyieoe` is built with the intention of being used with `obp`, these inputs must follow the conventions in `obp`. Specifically, logged bandit feedback data must be of type `BanditFeedback`, action distributions must be of type `np.ndarray`, and ground truth policy values must be of type `float` (or `int`).
#
# In this example, we generate logged bandit feedback data by modifying multiclass classification data and obtain two sets of evaluation policies along with their action distributions and ground truth policy values using `obp`. For a detailed explanation of this process, please refer to the [official docs](https://zr-obp.readthedocs.io/en/latest/_autosummary/obp.dataset.multiclass.html#module-obp.dataset.multiclass).
# + id="n1EjJoJ_JqXg"
# load raw digits data
X, y = load_digits(return_X_y=True)
# convert the raw classification data into the logged bandit dataset
dataset = MultiClassToBanditReduction(
X=X,
y=y,
base_classifier_b=LogisticRegression(random_state=12345),
alpha_b=0.8,
dataset_name="digits"
)
# split the original data into the training and evaluation sets
dataset.split_train_eval(eval_size=0.7, random_state=12345)
# obtain logged bandit feedback generated by the behavior policy
bandit_feedback = dataset.obtain_batch_bandit_feedback(random_state=12345)
# obtain action choice probabilities by an evaluation policy and its ground-truth policy value
action_dist_a = dataset.obtain_action_dist_by_eval_policy(
base_classifier_e=LogisticRegression(C=100, random_state=12345, max_iter=10000),
alpha_e=0.9
)
ground_truth_a = dataset.calc_ground_truth_policy_value(action_dist=action_dist_a)
action_dist_b = dataset.obtain_action_dist_by_eval_policy(
base_classifier_e=RandomForest(n_estimators=100, min_samples_split=5, random_state=12345),
alpha_e=0.9
)
ground_truth_b = dataset.calc_ground_truth_policy_value(action_dist=action_dist_b)
# + [markdown] id="xDmLkCTnJqXi"
# ### Setting Hyperparameter Spaces for Off-Policy Evaluation
#
# An integral aspect of IEOE is the different sources of variance. The main sources of variance are evaluation policies, random states, hyperparameters of OPE estimators, and hyperparameters of regression models.
#
# In this step, we define the spaces from which the hyperparameters of OPE estimators / regression models are chosen. (The evaluation policy space is defined in the previous step, and the random state space will be defined in the next step.)
# + id="FG_hpxWcJqXj"
# set hyperparameter space for ope estimators
# set hyperparameter space for the doubly robust with shrinkage estimator
# with the following code, lambda_ will be chosen from a logarithm uniform distribution over the interval [0.001, 1000]
lambda_ = {
"lower": 1e-3,
"upper": 1e3,
"log": True,
"type": float
}
dros_param = {"lambda_": lambda_}
# + id="fy3DSPK9JqXk"
# set hyperparameter space for logistic regression using RandomizedSearchCV
from sklearn.utils.fixes import loguniform
logistic = LogisticRegression()
distributions = {
"C": loguniform(1e-2, 1e2)
}
clf_logistic = RandomizedSearchCV(logistic, distributions, random_state=0, n_iter=5)
# + id="z6_SJ6vrJqXl"
# set hyperparameter space for random forest classifier using RandomizedSearchCV
from scipy.stats import randint
randforest = RandomForest()
distributions = {
# n_estimators will be chosen from a uniform distribution over the interval [50, 100)
"n_estimators": randint(5e1, 1e2),
# max_depth will be chosen from a uniform distribution over the interval [2, 10)
"max_depth": randint(2, 10),
# min_samples_split will be chosen from a uniform distribution over the interval [2, 10)
"min_samples_split": randint(2, 10)
}
clf_randforest = RandomizedSearchCV(randforest, distributions, random_state=0, n_iter=5)
# + [markdown] id="7T7Rg-jXJqXl"
# ### Interpretable Evaluation for Off-Policy Evaluation
#
# With the above steps completed, we can finally conduct IEOE by utilizing the `InterpretableOPEEvaluator` class.
#
# Here is a brief description for each parameter that can be passed into `InterpretableOPEEvaluator`:
#
# - `random_states`: a list of integers representing the random_state used when performing OPE; corresponds to the number of iterations
# - `bandit_feedback`: a list of logged bandit feedback data
# - `evaluation_policies`: a list of tuples representing (ground truth policy value, action distribution)
# - `ope_estimators`: a list of OPE ope_estimators
# - `ope_estimator_hyperparams`: a dictionary mapping OPE estimator names to OPE estimator hyperparameter spaces defined in step 2
# - `regression_models`: a list of regression regression_models
# - `regression_model_hyperparams`: a dictionary mapping regression models to regression model hyperparameter spaces defined in step 2
# + id="1xSx2ZwvJqXm"
# initializing class
evaluator = InterpretableOPEEvaluator(
random_states=np.arange(100),
bandit_feedbacks=[bandit_feedback],
evaluation_policies=[
(ground_truth_a, action_dist_a),
(ground_truth_b, action_dist_b)
],
ope_estimators=[
DirectMethod(),
DoublyRobust(),
DoublyRobustWithShrinkage(),
InverseProbabilityWeighting(),
],
ope_estimator_hyperparams={
DoublyRobustWithShrinkage.estimator_name: dros_param,
},
regression_models=[
clf_logistic,
clf_randforest
],
pscore_estimators=[
clf_logistic,
clf_randforest
]
)
# + [markdown] id="sv_kCRYGJqXm"
# Once we have initialized `InterpretableOPEEvaluator`, we can call implemented methods to perform IEOE.
# + id="jGujF_feJqXn" outputId="642095bc-6cdc-473f-b143-3613441a320e"
# estimate policy values
# we obtain a dictionary mapping ope estimator names to np.ndarray storing the estimated policy value for each iteration
policy_value = evaluator.estimate_policy_value()
# + id="yfJ6E95TJqXn" outputId="ada81cd1-9fc5-407f-c03b-0f14b8b159d6"
print("dm:", policy_value["dm"][:3])
print("dr:", policy_value["dr"][:3])
print("dr-os:", policy_value["dr-os"][:3])
print("ipw:", policy_value["ipw"][:3])
# + id="KKQtHYxCJqXo"
# compute squared errors
# we obtain a dictionary mapping ope estimator names to np.ndarray storing the calculated squared error for each iteration
squared_error = evaluator.calculate_squared_error()
# + id="Ni5mcDu7JqXo" outputId="91e336ff-5915-4842-d6f0-c1255b36d016"
print("dm:", squared_error["dm"][:3])
print("dr:", squared_error["dr"][:3])
print("dr-os:", squared_error["dr-os"][:3])
print("ipw:", squared_error["ipw"][:3])
# + id="2aXY2t2FJqXp" outputId="69ab5fea-137b-44b6-a162-b7f147cf52c7"
# visualize cdf of squared errors for all ope estimators
evaluator.visualize_cdf_aggregate(xmax=0.04)
# + id="l_JfY0PVJqXq" outputId="68a06243-7692-494f-b723-92447c860e4d"
# compute the au-cdf score (area under cdf of squared error over interval [0, thershold]), higher score is better
# we obtain a dictionary mapping ope estimator names to cvar scores
au_cdf = evaluator.calculate_au_cdf_score(threshold=0.004)
au_cdf
# + id="HMMlyuvZJqXq" outputId="17fcbef6-98ca-4194-c67f-36533024a6a5"
# by activating the `scale` option,
# we obtain au_cdf scores where the highest score is scaled to 1
au_cdf_scaled = evaluator.calculate_au_cdf_score(threshold=0.004, scale=True)
au_cdf_scaled
# + id="Hbzzn13kJqXr" outputId="6a61c824-45ab-4cc2-e6a8-5116080942ee"
# compute the cvar score (expected value of squared error above probability alpha), lower score is better
# we obtain a dictionary mapping ope estimator names to cvar scores
cvar = evaluator.calculate_cvar_score(alpha=90)
cvar
# + id="ZvO-fTAUJqXr" outputId="f7093127-78a5-4f96-db68-2a08d8503e67"
# by activating the `scale` option,
# we obtain cvar scores where the lowest score is scaled to 1
cvar_scaled = evaluator.calculate_cvar_score(alpha=90, scale=True)
cvar_scaled
# + id="wgF3P3ZWJqXs"
| docs/T902666_Evaluating_the_Robustness_of_Off_Policy_Evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#All the needed libraries for the project
import pandas as pd
import numpy as np
import sys
from pandas import DataFrame
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.preprocessing import OneHotEncoder
from sklearn.model_selection import train_test_split
import random
scaled_ble= pd.read_csv('datasets/Scaled_BLE_RSSI.csv')
distance = pd.read_csv('datasets/iBeacon_Distance.csv')
X = scaled_ble.values
y = distance.values
X = np.array(X)
X = np.reshape(X, (-1, ))
X.shape
y = np.array(y)
y = np.reshape(y, (-1, ))
y.shape[0]
X = np.reshape(X, (-1,1 ))
y = np.reshape(y, (-1,1 ))
X[20,0] == 0.0
# +
index = []
for i in range(0,X.shape[0]):
if X[i,0] == 0.0:
index.append(i)
X = np.delete(X, index)
y = np.delete(y, index)
X.shape
# -
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
print( scaler.fit(X.reshape(-1, 1)))
Xs = scaler.transform(X.reshape(-1, 1))
from sklearn.preprocessing import quantile_transform
Xs = quantile_transform(X.reshape(-1, 1), n_quantiles=5)
plt.scatter(X,y)
plt.ylabel('Distance')
plt.xlabel('Scaled RSSI Values')
plt.show()
y1
# +
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeRegressor
X_train, X_test, y_train, y_test = train_test_split(Xs, y,
test_size=0.15,
random_state=4)
# -
regr_rf = DecisionTreeRegressor(max_depth=7,criterion='mse')
regr_rf.fit(X_train.reshape((-1,1)), y_train.reshape((-1,)))
regr_rf.predict(0.9996)
regr_rf.score(X_test.reshape((-1,1)), y_test)
| Data_testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # JIT Engine: Scalar + Scalar
#
# This example will go over how to compile MLIR code to a function callable from Python.
#
# The example MLIR code we'll use here performs scalar addition.
#
# Let’s first import some necessary modules and generate an instance of our JIT engine.
# +
import mlir_graphblas
import numpy as np
engine = mlir_graphblas.MlirJitEngine()
# -
# Here's some MLIR code to add two 32-bit floating point numbers.
mlir_text = r"""
func @scalar_add_f32(%a: f32, %b: f32) -> f32 {
%ans = arith.addf %a, %b : f32
return %ans : f32
}
"""
# Let's say we wanted to optimize our code with the following [MLIR passes](https://mlir.llvm.org/docs/Passes/):
passes = [
"--linalg-bufferize",
"--func-bufferize",
"--tensor-bufferize",
"--tensor-constant-bufferize",
"--finalizing-bufferize",
"--convert-linalg-to-loops",
"--convert-scf-to-std",
"--convert-arith-to-llvm",
"--convert-math-to-llvm",
"--convert-std-to-llvm",
]
# We can compile the MLIR code using our JIT engine.
engine.add(mlir_text, passes)
# The returned value above is a list of the names of all functions compiled in the given MLIR code.
#
# We can access the compiled Python callables in two ways:
func_1 = engine['scalar_add_f32']
func_2 = engine.scalar_add_f32
# They both point to the same function:
func_1 is func_2
# We can call our function in Python:
scalar_add_f32 = engine.scalar_add_f32
scalar_add_f32(100.0, 200.0)
# Let's try creating a function to add two 8-bit integers.
mlir_text = r"""
func @scalar_add_i8(%a: i8, %b: i8) -> i8 {
%ans = arith.addi %a, %b : i8
return %ans : i8
}
"""
engine.add(mlir_text, passes)
scalar_add_i8 = engine.scalar_add_i8
# Let's verify that it works.
scalar_add_i8(30, 40)
# What happens if we give invalid inputs, e.g. integers too large to fit into 8-bits?
scalar_add_i8(9999, 9999)
# We get an exception! There's some input and output type checking that takes place in compiled callables, so there's some safety provided by the JIT Engine.
| docs/tools/engine/scalar_plus_scalar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 8.1
# language: ''
# name: sagemath
# ---
# En este evaluación, aprenderemos a analizar gráficas de funciones encontrando puntos destacados tales como
# * Ceros (o raíces)
# * Puntos críticos
# * Puntos de inflexión
# ## Raíces
#
# Los ceros o raíces de una función $f:I\to\mathbb{R}$ son aquellos puntos $r\in I$ tales que $f(r)=0$.
#
# ### Ejemplo
# 1. Determina las raíces de $f(x)=xe^x$.
# 2. Bosqueja la gráfica marcando las intersecciones con los ejes.
# 3. Determina los intervalos en que la función es positiva y en los que es negativa.
# #### Solución
# Primero resolvemos la ecuación utilizando el método `solve`
f(x) = x*exp(x)
ecuacion_raices = f(x)==0
raices = solve(ecuacion_raices,x)
show(raices)
# En el bloque anterior determinamos que la única raíz es $x=0$, por lo que la única intersección con el eje $x$ es $(0,0)$. Por pura coincidencia, resulta que es también la intersección con el eje $y$.
# trazamos la gráfica de f
grafica = plot(f)
# añadimos (+=) la intersección con los ejes
## tamaño = 10, color = rojo
grafica += point((0,0), size=100, color="red")
# mostramos la gráfica
show(grafica)
# Ahora determinaremos cuando la función es positiva ($f(x)>0$) y cuando es negativa ($f(x)<0$).
# determinamos el intervalo I={x|f(x)>0}
f_positiva = solve(f(x)>0, x)
print(f_positiva)
# determinamos el intervalo I={x|f(x)<0}
f_negativa = solve(f(x)<0, x)
print(f_negativa)
# Concluímos que la función es positiva en el intervalo $x>0$, mientras que es negativa en $x<0$.
# ## Puntos críticos
#
# Los puntos críticos de una función diferenciable $f:I\to\mathbb{R}$ son aquellos puntos $r\in I$ tales que $f'(r)=0$.
#
# ### Ejemplo
# 1. Determina los puntos críticos de $f(x)=xe^x$.
# 2. Marca estos puntos en la gráfica anterior.
# 3. Determina los intervalos en que la función es creciente y en los que es decreciente.
# #### Solución
#
# Calcularemos primero la derivada de la función; plantearemos la ecuación de sus puntos críticos y resolveremos.
# calculamos la derivada fx de la función f
fx(x) = f.diff(x)
# planteamos la ecuación del punto crítico
ecuacion_pto_critico = fx(x)==0
# resolvemos
ptos_criticos = solve(ecuacion_pto_critico, x)
show(ptos_criticos)
# Del inciso anterior, concluímos que el único punto crítico es
| CALCULO 100 EVALUACION CONTINUA .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reading and writing data
# ## Read datasets into chunks
# There are two main options for loading an `xarray.Dataset` into Xarray-Beam. You can either [create the dataset](data-model.ipynb) from scratch or use the {py:class}`~xarray_beam.DatasetToChunks` transform starting at the root of a Beam pipeline:
# +
# hidden imports & helper functions
# + tags=["hide-input"]
import textwrap
import apache_beam as beam
import xarray_beam as xbeam
import xarray
def summarize_dataset(dataset):
return f'<xarray.Dataset data_vars={list(dataset.data_vars)} dims={dict(dataset.sizes)}>'
def print_summary(key, chunk):
print(f'{key}\n with {summarize_dataset(chunk)}')
# -
ds = xarray.tutorial.load_dataset('air_temperature')
with beam.Pipeline() as p:
p | xbeam.DatasetToChunks(ds, chunks={'time': 1000}) | beam.MapTuple(print_summary)
# Importantly, xarray datasets fed into `DatasetToChunks` **can be lazy**, with data not already loaded eagerly into NumPy arrays. When you feed lazy datasets into `DatasetToChunks`, each individual chunk will be indexed and evaluated separately on Beam workers.
#
# This pattern allows for leveraging Xarray's builtin dataset loaders (e.g., `open_dataset()` and `open_zarr()`) for feeding arbitrarily large datasets into Xarray-Beam.
#
# For best performance, set `chunks=None` when opening datasets and then _explicitly_ provide chunks in `DatasetToChunks`:
# +
# write data into the distributed Zarr format
ds.chunk({'time': 1000}).to_zarr('example-data.zarr', mode='w')
# load it with zarr
on_disk = xarray.open_zarr('example-data.zarr', chunks=None)
with beam.Pipeline() as p:
p | xbeam.DatasetToChunks(on_disk, chunks={'time': 1000}) | beam.MapTuple(print_summary)
# -
# `chunks=None` tells Xarray to use its builtin lazy indexing machinery, instead of using Dask. This is advantageous because datasets using Xarray's lazy indexing are serialized much more compactly (via [pickle](https://docs.python.org/3/library/pickle.html)) when passed into Beam transforms.
# Alternatively, you can pass in lazy datasets [using dask](http://xarray.pydata.org/en/stable/user-guide/dask.html). In this case, you don't need to explicitly supply `chunks` to `DatasetToChunks`:
# +
on_disk = xarray.open_zarr('example-data.zarr', chunks={'time': 1000})
with beam.Pipeline() as p:
p | xbeam.DatasetToChunks(on_disk) | beam.MapTuple(print_summary)
# -
# Dask's lazy evaluation system is much more general than Xarray's lazy indexing, so as long as resulting dataset can be independently evaluated in each chunk this can be a very convenient way to setup computation for Xarray-Beam.
#
# Unfortunately, it doesn't scale as well. In particular, the overhead of pickling large Dask graphs for passing to Beam workers can be prohibitive for large (typically multiple TB) datasets with millions of chunks. However, a current major effort in Dask on [high level graphs](https://blog.dask.org/2021/07/07/high-level-graphs) should improve this in the near future.
# ```{note}
# We are still figuring out the optimal APIs to facilitate opening data and building lazy datasets in Xarray-Beam. E.g., see [this issue](https://github.com/google/xarray-beam/issues/26) for discussion of a higher level `ZarrToChunks` transform embedding these best practices.
# ```
# ## Writing data to Zarr
# [Zarr](https://zarr.readthedocs.io/) is the preferred file format for reading and writing data with Xarray-Beam, due to its excellent scalability and support inside Xarray.
#
# {py:class}`~xarray_beam.ChunksToZarr` is Xarray-Beam's API for saving chunks into a Zarr store.
#
# You can get started just using it directly:
with beam.Pipeline() as p:
p | xbeam.DatasetToChunks(on_disk) | xbeam.ChunksToZarr('example-data-v2.zarr')
# By default, `ChunksToZarr` needs to evaluate and combine the entire distributed dataset in order to determine overall Zarr metadata (e.g., array names, shapes, dtypes and attributes). This is fine for relatively small datasets, but can entail significant additional communication and storage costs for large datasets.
#
# The optional `template` argument allows for prespecifying structure of the full on disk dataset in the form of another lazy `xarray.Dataset`. Like the lazy datasets fed into DatasetToChunks, lazy templates can built-up using either Xarray's lazy indexing or lazy operations with Dask, but the data _values_ in a `template` will never be written to disk -- only the metadata structure is used.
#
# One recommended pattern is to use a lazy Dask dataset consisting of a single value to build up the desired template, e.g.,
ds = xarray.open_zarr('example-data.zarr', chunks=None)
template = xarray.zeros_like(ds.chunk()) # a single virtual chunk of all zeros
# Xarray operations like indexing and expand dimensions (see {py:meth}`xarray.Dataset.expand_dims`) are entirely lazy on this dataset, which makes it relatively straightforward to build up a Dataset with the required variables and dimensions, e.g., as used in the [ERA5 climatology example](https://github.com/google/xarray-beam/blob/main/examples/era5_climatology.py).
#
# Note that if supply a `template`, you will also typically need to specify the `chunks` argument in order to ensure that the data ends up appropriately chunked in the Zarr store.
# ```{warning}
# Xarray-Beam does not use locks when writing data to Zarr. If multiple Beam chunks correspond to the same Zarr chunk, your will almost certainly end up with corrupted data due to concurrent writes. To avoid such issues, ensure your data is [chunked appropriately](rechunking.ipynb) before exporting to Zarr.
# ```
| docs/read-write.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pyross.tsi as pyrosstsi
import numpy as np
import matplotlib.pyplot as plt
# +
T = 15 # Longest infectious duration
Td = 5 # Doubling time in linear growth regime
Tf = 150 # Duration of simulation
tsi = np.array([0, 3, 5, 10, T]) # Time since infection (days)
beta = np.array([0, 0.5, 1, .5, 0]) # Mean infectiousness
M = 2 # Number of age groups to model
Ni = 10**6*np.ones(M) # Number of people in each age group
Np = sum(Ni) # Total population size
#how many 'stages' to resolve in time since infection?
Nk = 10
#define a time-dependent contact matrix.
#variations on timescale less than T/Nk may not be resolved.
def contactMatrix(t):
if t > 15 and t < 100:
return 1*np.array([[4, 1],[1, 2]])
else:
return np.array([[4, 1],[1, 2]]);
subclasses = ['Recovered', 'Hospitalized', 'Mortality']
pR = 0.99*np.ones(M); #probability of eventually recovering for each age class
pH = 0.05*np.ones(M); #probability of needing hospitalization for each age class
pD = 1-pR; #probability of death for each age class
#prepare for a linear interpolating function evaluated at times:
tsi_sc = np.array([0, 3., 6., 9., 12, T])
phiR = np.array([0, 0, 0.5, 3, 2, 0])#rate of transferring to 'recovered' (arbitrary units)
phiH_in = np.array([0, 0, 1, 1, 0, 0])#rate that people enter hospital (arbitrary units)
phiH_out = np.array([0, 0, 0, 1, 1, 0])#rate that people exit hospital (arbitrary units)
phiD = np.array([0, 0, 0, 1, 1, .5])#times at which a person dies (arbitrary units)
#combine hospital in/out to a single function for net change in hospitalized cases
phiH = np.add(-phiH_out/np.trapz(phiH_out,tsi_sc),phiH_in/np.trapz(phiH_in,tsi_sc))
#normalize all to one -- can then be rescaled by approprate pR, pH, pD, etc. at a later time
phiR, phiD = phiR/np.trapz(phiR,tsi_sc), phiD/np.trapz(phiD,tsi_sc)
#group them all together for later processing
phi_alpha, p_alpha = np.array([phiR, phiH, phiD]), np.array([pR, pH, pD])
# +
parameters = {'M':M, 'Ni':Ni, 'Nc':len(subclasses), 'Nk':Nk, 'Tf':Tf, 'Tc':(T/2), 'T':T, 'Td':Td,
'tsi':tsi,'beta':beta,'tsi_sc':tsi_sc, 'phi_alpha':phi_alpha, 'p_alpha':p_alpha,
'contactMatrix':contactMatrix}
model = pyrosstsi.deterministic.Simulator(parameters)
IC = model.get_IC()
data = model.simulate(IC)
# -
# ## Plot the results using default 'Predictor Corrector':
#
# This is preferred integrator for most cases, but it does not support adaptive timestepping.
# +
#unpack and rescale simulation output
t = data['t']; S_t = data['S_t']; I_t = data['I_t']; Ic_t = data['Ic_t']
plt.figure(figsize=(12, 4)); plt.subplot(121)
plt.plot(t,np.sum(S_t,0), color="#348ABD", lw=2, label = 'Susceptible') #all susceptible
plt.plot(t,np.sum(I_t,0), color="#A60628", lw=2, label = 'Infected') #all Infected
plt.plot(t,np.sum(Ic_t[0,:,:],0), color='green', lw=2, label = 'Recovered') #all Recovered
plt.xlabel('time (days)'); plt.xlim(0,Tf);
plt.ylabel('Fraction of compartment value'); plt.legend()
plt.subplot(122)
for i in (1 + np.arange(len(subclasses)-1)):
plt.plot(t,np.sum(Ic_t[i,:,:],0), lw=2, label = subclasses[i])
plt.legend(); plt.xlabel('time (days)'); plt.xlabel('time (days)'); plt.xlim(0,Tf);
# -
# ## Repeat same simulation using Galerkin Discretization and default integrator (odeint)
#
# This integrator supports adaptive timestepping but it is not recommended for time-dependent contact matrices or non-smooth dynamic more generally.
# +
parameters['NL'] = 5
model = pyrosstsi.deterministic.Simulator(parameters,'Galerkin')
IC = model.get_IC()
data = model.simulate(IC)#,10**-3,10**-2)# <- error tolerance options
#unpack and rescale simulation output
t = data['t']; S_t = data['S_t']; I_t = data['I_t']; Ic_t = data['Ic_t']
plt.figure(figsize=(12, 4)); plt.subplot(121)
plt.plot(t,np.sum(S_t,0), color="#348ABD", lw=2, label = 'Susceptible') #all susceptible
plt.plot(t,np.sum(I_t,0), color="#A60628", lw=2, label = 'Infected') #all Infected
plt.plot(t,np.sum(Ic_t[0,:,:],0), color='green', lw=2, label = 'Recovered') #all Recovered
plt.xlabel('time (days)'); plt.xlim(0,Tf);
plt.ylabel('Fraction of compartment value'); plt.legend()
plt.subplot(122)
for i in (1 + np.arange(len(subclasses)-1)):
plt.plot(t,np.sum(Ic_t[i,:,:],0), lw=2, label = subclasses[i])
plt.legend(); plt.xlabel('time (days)'); plt.xlabel('time (days)'); plt.xlim(0,Tf);
# -
# ## Repeat the same using Galerkin discretization and <NAME> integrator
#
# This integrator supports adaptive time-stepping and is preferable to 'odeint' whenever the contact matrix is time-dependent. Still not recommended for non-smooth dynamics (e.g. lockdown). When the contact matrix is time-dependent and piecewise smooth, consider using the Hybrid method (see example notebook on the subject).
# +
parameters['NL'] = 5
model = pyrosstsi.deterministic.Simulator(parameters,'Galerkin','<NAME>')
IC = model.get_IC()
data = model.simulate(IC)#,10**-1,10**-1)# <- error tolerance options
#unpack and rescale simulation output
t = data['t']; S_t = data['S_t']; I_t = data['I_t']; Ic_t = data['Ic_t']
plt.figure(figsize=(12, 4)); plt.subplot(121)
plt.plot(t,np.sum(S_t,0), color="#348ABD", lw=2, label = 'Susceptible') #all susceptible
plt.plot(t,np.sum(I_t,0), color="#A60628", lw=2, label = 'Infected') #all Infected
plt.plot(t,np.sum(Ic_t[0,:,:],0), color='green', lw=2, label = 'Recovered') #all Recovered
plt.xlabel('time (days)'); plt.xlim(0,Tf);
plt.ylabel('Fraction of compartment value'); plt.legend()
plt.subplot(122)
for i in (1 + np.arange(len(subclasses)-1)):
plt.plot(t,np.sum(Ic_t[i,:,:],0), lw=2, label = subclasses[i])
plt.legend(); plt.xlabel('time (days)'); plt.xlabel('time (days)'); plt.xlim(0,Tf);
| examples/tsi/ex01.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:percent
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %% [markdown]
# # Assignment 1: Neural Machine Translation
#
# Welcome to the first assignment of Course 4. Here, you will build an English-to-German neural machine translation (NMT) model using Long Short-Term Memory (LSTM) networks with attention. Machine translation is an important task in natural language processing and could be useful not only for translating one language to another but also for word sense disambiguation (e.g. determining whether the word "bank" refers to the financial bank, or the land alongside a river). Implementing this using just a Recurrent Neural Network (RNN) with LSTMs can work for short to medium length sentences but can result in vanishing gradients for very long sequences. To solve this, you will be adding an attention mechanism to allow the decoder to access all relevant parts of the input sentence regardless of its length. By completing this assignment, you will:
#
# - learn how to preprocess your training and evaluation data
# - implement an encoder-decoder system with attention
# - understand how attention works
# - build the NMT model from scratch using Trax
# - generate translations using greedy and Minimum Bayes Risk (MBR) decoding
# ## Outline
# - [Part 1: Data Preparation](#1)
# - [1.1 Importing the Data](#1.1)
# - [1.2 Tokenization and Formatting](#1.2)
# - [1.3 tokenize & detokenize helper functions](#1.3)
# - [1.4 Bucketing](#1.4)
# - [1.5 Exploring the data](#1.5)
# - [Part 2: Neural Machine Translation with Attention](#2)
# - [2.1 Attention Overview](#2.1)
# - [2.2 Helper functions](#2.2)
# - [Exercise 01](#ex01)
# - [Exercise 02](#ex02)
# - [Exercise 03](#ex03)
# - [2.3 Implementation Overview](#2.3)
# - [Exercise 04](#ex04)
# - [Part 3: Training](#3)
# - [3.1 TrainTask](#3.1)
# - [Exercise 05](#ex05)
# - [3.2 EvalTask](#3.2)
# - [3.3 Loop](#3.3)
# - [Part 4: Testing](#4)
# - [4.1 Decoding](#4.1)
# - [Exercise 06](#ex06)
# - [Exercise 07](#ex07)
# - [4.2 Minimum Bayes-Risk Decoding](#4.2)
# - [Exercise 08](#ex08)
# - [Exercise 09](#ex09)
# - [Exercise 10](#ex10)
#
# %% [markdown]
# <a name="1"></a>
# # Part 1: Data Preparation
#
# %% [markdown]
# <a name="1.1"></a>
# ## 1.1 Importing the Data
#
# We will first start by importing the packages we will use in this assignment. As in the previous course of this specialization, we will use the [Trax](https://github.com/google/trax) library created and maintained by the [Google Brain team](https://research.google/teams/brain/) to do most of the heavy lifting. It provides submodules to fetch and process the datasets, as well as build and train the model.
# %%
from termcolor import colored
import random
import numpy as np
import trax
from trax import layers as tl
from trax.fastmath import numpy as fastnp
from trax.supervised import training
# !pip list | grep trax
# %% [markdown]
# Next, we will import the dataset we will use to train the model. To meet the storage constraints in this lab environment, we will just use a small dataset from [Opus](http://opus.nlpl.eu/), a growing collection of translated texts from the web. Particularly, we will get an English to German translation subset specified as `opus/medical` which has medical related texts. If storage is not an issue, you can opt to get a larger corpus such as the English to German translation dataset from [ParaCrawl](https://paracrawl.eu/), a large multi-lingual translation dataset created by the European Union. Both of these datasets are available via [Tensorflow Datasets (TFDS)](https://www.tensorflow.org/datasets)
# and you can browse through the other available datasets [here](https://www.tensorflow.org/datasets/catalog/overview). We have downloaded the data for you in the `data/` directory of your workspace. As you'll see below, you can easily access this dataset from TFDS with `trax.data.TFDS`. The result is a python generator function yielding tuples. Use the `keys` argument to select what appears at which position in the tuple. For example, `keys=('en', 'de')` below will return pairs as (English sentence, German sentence).
# %%
# Get generator function for the training set
# This will download the train dataset if no data_dir is specified.
train_stream_fn = trax.data.TFDS('opus/medical',
data_dir='./data/',
keys=('en', 'de'),
eval_holdout_size=0.01, # 1% for eval
train=True)
# Get generator function for the eval set
eval_stream_fn = trax.data.TFDS('opus/medical',
data_dir='./data/',
keys=('en', 'de'),
eval_holdout_size=0.01, # 1% for eval
train=False)
# %% [markdown]
# Notice that TFDS returns a generator *function*, not a generator. This is because in Python, you cannot reset generators so you cannot go back to a previously yielded value. During deep learning training, you use Stochastic Gradient Descent and don't actually need to go back -- but it is sometimes good to be able to do that, and that's where the functions come in. It is actually very common to use generator functions in Python -- e.g., `zip` is a generator function. You can read more about [Python generators](https://book.pythontips.com/en/latest/generators.html) to understand why we use them. Let's print a a sample pair from our train and eval data. Notice that the raw ouput is represented in bytes (denoted by the `b'` prefix) and these will be converted to strings internally in the next steps.
# %%
train_stream = train_stream_fn()
print(colored('train data (en, de) tuple:', 'red'), next(train_stream))
print()
eval_stream = eval_stream_fn()
print(colored('eval data (en, de) tuple:', 'red'), next(eval_stream))
# %% [markdown]
# <a name="1.2"></a>
# ## 1.2 Tokenization and Formatting
#
# Now that we have imported our corpus, we will be preprocessing the sentences into a format that our model can accept. This will be composed of several steps:
#
# **Tokenizing the sentences using subword representations:** As you've learned in the earlier courses of this specialization, we want to represent each sentence as an array of integers instead of strings. For our application, we will use *subword* representations to tokenize our sentences. This is a common technique to avoid out-of-vocabulary words by allowing parts of words to be represented separately. For example, instead of having separate entries in your vocabulary for --"fear", "fearless", "fearsome", "some", and "less"--, you can simply store --"fear", "some", and "less"-- then allow your tokenizer to combine these subwords when needed. This allows it to be more flexible so you won't have to save uncommon words explicitly in your vocabulary (e.g. *stylebender*, *nonce*, etc). Tokenizing is done with the `trax.data.Tokenize()` command and we have provided you the combined subword vocabulary for English and German (i.e. `ende_32k.subword`) saved in the `data` directory. Feel free to open this file to see how the subwords look like.
# %%
# global variables that state the filename and directory of the vocabulary file
VOCAB_FILE = 'ende_32k.subword'
VOCAB_DIR = 'data/'
# Tokenize the dataset.
tokenized_train_stream = trax.data.Tokenize(vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR)(train_stream)
tokenized_eval_stream = trax.data.Tokenize(vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR)(eval_stream)
# %% [markdown]
# **Append an end-of-sentence token to each sentence:** We will assign a token (i.e. in this case `1`) to mark the end of a sentence. This will be useful in inference/prediction so we'll know that the model has completed the translation.
# %%
# Append EOS at the end of each sentence.
# Integer assigned as end-of-sentence (EOS)
EOS = 1
# generator helper function to append EOS to each sentence
def append_eos(stream):
for (inputs, targets) in stream:
inputs_with_eos = list(inputs) + [EOS]
targets_with_eos = list(targets) + [EOS]
yield np.array(inputs_with_eos), np.array(targets_with_eos)
# append EOS to the train data
tokenized_train_stream = append_eos(tokenized_train_stream)
# append EOS to the eval data
tokenized_eval_stream = append_eos(tokenized_eval_stream)
# %% [markdown]
# **Filter long sentences:** We will place a limit on the number of tokens per sentence to ensure we won't run out of memory. This is done with the `trax.data.FilterByLength()` method and you can see its syntax below.
# %%
# Filter too long sentences to not run out of memory.
# length_keys=[0, 1] means we filter both English and German sentences, so
# both much be not longer that 256 tokens for training / 512 for eval.
filtered_train_stream = trax.data.FilterByLength(
max_length=256, length_keys=[0, 1])(tokenized_train_stream)
filtered_eval_stream = trax.data.FilterByLength(
max_length=512, length_keys=[0, 1])(tokenized_eval_stream)
# print a sample input-target pair of tokenized sentences
train_input, train_target = next(filtered_train_stream)
print(colored(f'Single tokenized example input:', 'red' ), train_input)
print(colored(f'Single tokenized example target:', 'red'), train_target)
# %% [markdown]
# <a name="1.3"></a>
# ## 1.3 tokenize & detokenize helper functions
#
# Given any data set, you have to be able to map words to their indices, and indices to their words. The inputs and outputs to your trax models are usually tensors of numbers where each number corresponds to a word. If you were to process your data manually, you would have to make use of the following:
#
# - <span style='color:blue'> word2Ind: </span> a dictionary mapping the word to its index.
# - <span style='color:blue'> ind2Word:</span> a dictionary mapping the index to its word.
# - <span style='color:blue'> word2Count:</span> a dictionary mapping the word to the number of times it appears.
# - <span style='color:blue'> num_words:</span> total number of words that have appeared.
#
# Since you have already implemented these in previous assignments of the specialization, we will provide you with helper functions that will do this for you. Run the cell below to get the following functions:
#
# - <span style='color:blue'> tokenize(): </span> converts a text sentence to its corresponding token list (i.e. list of indices). Also converts words to subwords (parts of words).
# - <span style='color:blue'> detokenize(): </span> converts a token list to its corresponding sentence (i.e. string).
# %%
# Setup helper functions for tokenizing and detokenizing sentences
def tokenize(input_str, vocab_file=None, vocab_dir=None):
"""Encodes a string to an array of integers
Args:
input_str (str): human-readable string to encode
vocab_file (str): filename of the vocabulary text file
vocab_dir (str): path to the vocabulary file
Returns:
numpy.ndarray: tokenized version of the input string
"""
# Set the encoding of the "end of sentence" as 1
EOS = 1
# Use the trax.data.tokenize method. It takes streams and returns streams,
# we get around it by making a 1-element stream with `iter`.
inputs = next(trax.data.tokenize(iter([input_str]),
vocab_file=vocab_file, vocab_dir=vocab_dir))
# Mark the end of the sentence with EOS
inputs = list(inputs) + [EOS]
# Adding the batch dimension to the front of the shape
batch_inputs = np.reshape(np.array(inputs), [1, -1])
return batch_inputs
def detokenize(integers, vocab_file=None, vocab_dir=None):
"""Decodes an array of integers to a human readable string
Args:
integers (numpy.ndarray): array of integers to decode
vocab_file (str): filename of the vocabulary text file
vocab_dir (str): path to the vocabulary file
Returns:
str: the decoded sentence.
"""
# Remove the dimensions of size 1
integers = list(np.squeeze(integers))
# Set the encoding of the "end of sentence" as 1
EOS = 1
# Remove the EOS to decode only the original tokens
if EOS in integers:
integers = integers[:integers.index(EOS)]
return trax.data.detokenize(integers, vocab_file=vocab_file, vocab_dir=vocab_dir)
# %% [markdown]
# Let's see how we might use these functions:
# %%
# As declared earlier:
# VOCAB_FILE = 'ende_32k.subword'
# VOCAB_DIR = 'data/'
# Detokenize an input-target pair of tokenized sentences
print(colored(f'Single detokenized example input:', 'red'), detokenize(train_input, vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR))
print(colored(f'Single detokenized example target:', 'red'), detokenize(train_target, vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR))
print()
# Tokenize and detokenize a word that is not explicitly saved in the vocabulary file.
# See how it combines the subwords -- 'hell' and 'o'-- to form the word 'hello'.
print(colored(f"tokenize('hello'): ", 'green'), tokenize('hello', vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR))
print(colored(f"detokenize([17332, 140, 1]): ", 'green'), detokenize([17332, 140, 1], vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR))
# %% [markdown]
# <a name="1.4"></a>
# ## 1.4 Bucketing
#
# Bucketing the tokenized sentences is an important technique used to speed up training in NLP.
# Here is a
# [nice article describing it in detail](https://medium.com/@rashmi.margani/how-to-speed-up-the-training-of-the-sequence-model-using-bucketing-techniques-9e302b0fd976)
# but the gist is very simple. Our inputs have variable lengths and you want to make these the same when batching groups of sentences together. One way to do that is to pad each sentence to the length of the longest sentence in the dataset. This might lead to some wasted computation though. For example, if there are multiple short sentences with just two tokens, do we want to pad these when the longest sentence is composed of a 100 tokens? Instead of padding with 0s to the maximum length of a sentence each time, we can group our tokenized sentences by length and bucket, as on this image (from the article above):
#
# 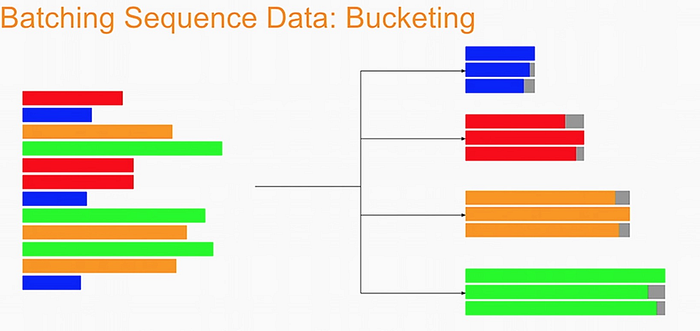
#
# We batch the sentences with similar length together (e.g. the blue sentences in the image above) and only add minimal padding to make them have equal length (usually up to the nearest power of two). This allows to waste less computation when processing padded sequences.
# In Trax, it is implemented in the [bucket_by_length](https://github.com/google/trax/blob/5fb8aa8c5cb86dabb2338938c745996d5d87d996/trax/supervised/inputs.py#L378) function.
# %%
# Bucketing to create streams of batches.
# Buckets are defined in terms of boundaries and batch sizes.
# Batch_sizes[i] determines the batch size for items with length < boundaries[i]
# So below, we'll take a batch of 256 sentences of length < 8, 128 if length is
# between 8 and 16, and so on -- and only 2 if length is over 512.
boundaries = [8, 16, 32, 64, 128, 256, 512]
batch_sizes = [256, 128, 64, 32, 16, 8, 4, 2]
# Create the generators.
train_batch_stream = trax.data.BucketByLength(
boundaries, batch_sizes,
length_keys=[0, 1] # As before: count inputs and targets to length.
)(filtered_train_stream)
eval_batch_stream = trax.data.BucketByLength(
boundaries, batch_sizes,
length_keys=[0, 1] # As before: count inputs and targets to length.
)(filtered_eval_stream)
# Add masking for the padding (0s).
train_batch_stream = trax.data.AddLossWeights(id_to_mask=0)(train_batch_stream)
eval_batch_stream = trax.data.AddLossWeights(id_to_mask=0)(eval_batch_stream)
# %% [markdown]
# <a name="1.5"></a>
# ## 1.5 Exploring the data
#
# We will now be displaying some of our data. You will see that the functions defined above (i.e. `tokenize()` and `detokenize()`) do the same things you have been doing again and again throughout the specialization. We gave these so you can focus more on building the model from scratch. Let us first get the data generator and get one batch of the data.
# %%
input_batch, target_batch, mask_batch = next(train_batch_stream)
# let's see the data type of a batch
print("input_batch data type: ", type(input_batch))
print("target_batch data type: ", type(target_batch))
# let's see the shape of this particular batch (batch length, sentence length)
print("input_batch shape: ", input_batch.shape)
print("target_batch shape: ", target_batch.shape)
# %% [markdown]
# The `input_batch` and `target_batch` are Numpy arrays consisting of tokenized English sentences and German sentences respectively. These tokens will later be used to produce embedding vectors for each word in the sentence (so the embedding for a sentence will be a matrix). The number of sentences in each batch is usually a power of 2 for optimal computer memory usage.
#
# We can now visually inspect some of the data. You can run the cell below several times to shuffle through the sentences. Just to note, while this is a standard data set that is used widely, it does have some known wrong translations. With that, let's pick a random sentence and print its tokenized representation.
# %%
# pick a random index less than the batch size.
index = random.randrange(len(input_batch))
# use the index to grab an entry from the input and target batch
print(colored('THIS IS THE ENGLISH SENTENCE: \n', 'red'), detokenize(input_batch[index], vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR), '\n')
print(colored('THIS IS THE TOKENIZED VERSION OF THE ENGLISH SENTENCE: \n ', 'red'), input_batch[index], '\n')
print(colored('THIS IS THE GERMAN TRANSLATION: \n', 'red'), detokenize(target_batch[index], vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR), '\n')
print(colored('THIS IS THE TOKENIZED VERSION OF THE GERMAN TRANSLATION: \n', 'red'), target_batch[index], '\n')
# %% [markdown]
# <a name="2"></a>
# # Part 2: Neural Machine Translation with Attention
#
# Now that you have the data generators and have handled the preprocessing, it is time for you to build the model. You will be implementing a neural machine translation model from scratch with attention.
#
# %% [markdown]
# <a name="2.1"></a>
# ## 2.1 Attention Overview
#
# The model we will be building uses an encoder-decoder architecture. This Recurrent Neural Network (RNN) will take in a tokenized version of a sentence in its encoder, then passes it on to the decoder for translation. As mentioned in the lectures, just using a a regular sequence-to-sequence model with LSTMs will work effectively for short to medium sentences but will start to degrade for longer ones. You can picture it like the figure below where all of the context of the input sentence is compressed into one vector that is passed into the decoder block. You can see how this will be an issue for very long sentences (e.g. 100 tokens or more) because the context of the first parts of the input will have very little effect on the final vector passed to the decoder.
#
# <img src='plain_rnn.png'>
#
# Adding an attention layer to this model avoids this problem by giving the decoder access to all parts of the input sentence. To illustrate, let's just use a 4-word input sentence as shown below. Remember that a hidden state is produced at each timestep of the encoder (represented by the orange rectangles). These are all passed to the attention layer and each are given a score given the current activation (i.e. hidden state) of the decoder. For instance, let's consider the figure below where the first prediction "Wie" is already made. To produce the next prediction, the attention layer will first receive all the encoder hidden states (i.e. orange rectangles) as well as the decoder hidden state when producing the word "Wie" (i.e. first green rectangle). Given these information, it will score each of the encoder hidden states to know which one the decoder should focus on to produce the next word. The result of the model training might have learned that it should align to the second encoder hidden state and subsequently assigns a high probability to the word "geht". If we are using greedy decoding, we will output the said word as the next symbol, then restart the process to produce the next word until we reach an end-of-sentence prediction.
#
# <img src='attention_overview.png'>
#
#
# There are different ways to implement attention and the one we'll use for this assignment is the Scaled Dot Product Attention which has the form:
#
# $$Attention(Q, K, V) = softmax(\frac{QK^T}{\sqrt{d_k}})V$$
#
# You will dive deeper into this equation in the next week but for now, you can think of it as computing scores using queries (Q) and keys (K), followed by a multiplication of values (V) to get a context vector at a particular timestep of the decoder. This context vector is fed to the decoder RNN to get a set of probabilities for the next predicted word. The division by square root of the keys dimensionality ($\sqrt{d_k}$) is for improving model performance and you'll also learn more about it next week. For our machine translation application, the encoder activations (i.e. encoder hidden states) will be the keys and values, while the decoder activations (i.e. decoder hidden states) will be the queries.
#
# You will see in the upcoming sections that this complex architecture and mechanism can be implemented with just a few lines of code. Let's get started!
# %% [markdown]
# <a name="2.2"></a>
# ## 2.2 Helper functions
#
# We will first implement a few functions that we will use later on. These will be for the input encoder, pre-attention decoder, and preparation of the queries, keys, values, and mask.
#
# ### 2.2.1 Input encoder
#
# The input encoder runs on the input tokens, creates its embeddings, and feeds it to an LSTM network. This outputs the activations that will be the keys and values for attention. It is a [Serial](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.combinators.Serial) network which uses:
#
# - [tl.Embedding](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.core.Embedding): Converts each token to its vector representation. In this case, it is the the size of the vocabulary by the dimension of the model: `tl.Embedding(vocab_size, d_model)`. `vocab_size` is the number of entries in the given vocabulary. `d_model` is the number of elements in the word embedding.
#
# - [tl.LSTM](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.rnn.LSTM): LSTM layer of size `d_model`. We want to be able to configure how many encoder layers we have so remember to create LSTM layers equal to the number of the `n_encoder_layers` parameter.
#
# <img src = "input_encoder.png">
#
# <a name="ex01"></a>
# ### Exercise 01
#
# **Instructions:** Implement the `input_encoder_fn` function.
# %%
# UNQ_C1
# GRADED FUNCTION
def input_encoder_fn(input_vocab_size, d_model, n_encoder_layers):
""" Input encoder runs on the input sentence and creates
activations that will be the keys and values for attention.
Args:
input_vocab_size: int: vocab size of the input
d_model: int: depth of embedding (n_units in the LSTM cell)
n_encoder_layers: int: number of LSTM layers in the encoder
Returns:
tl.Serial: The input encoder
"""
# create a serial network
input_encoder = tl.Serial(
### START CODE HERE (REPLACE INSTANCES OF `None` WITH YOUR CODE) ###
# create an embedding layer to convert tokens to vectors
tl.Embedding(input_vocab_size, d_model),
# feed the embeddings to the LSTM layers. It is a stack of n_encoder_layers LSTM layers
[tl.LSTM(d_model) for _ in range(n_encoder_layers)],
### END CODE HERE ###
)
return input_encoder
# %% [markdown]
# *Note: To make this notebook more neat, we moved the unit tests to a separate file called `w1_unittest.py`. Feel free to open it from your workspace if needed. Just click `File` on the upper left corner of this page then `Open` to see your Jupyter workspace directory. From there, you can see `w1_unittest.py` and you can open it in another tab or download to see the unit tests. We have placed comments in that file to indicate which functions are testing which part of the assignment (e.g. `test_input_encoder_fn()` has the unit tests for UNQ_C1).*
# %%
# BEGIN UNIT TEST
import w1_unittest
w1_unittest.test_input_encoder_fn(input_encoder_fn)
# END UNIT TEST
# %% [markdown]
# ### 2.2.2 Pre-attention decoder
#
# The pre-attention decoder runs on the targets and creates activations that are used as queries in attention. This is a Serial network which is composed of the following:
#
# - [tl.ShiftRight](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.attention.ShiftRight): This pads a token to the beginning of your target tokens (e.g. `[8, 34, 12]` shifted right is `[0, 8, 34, 12]`). This will act like a start-of-sentence token that will be the first input to the decoder. During training, this shift also allows the target tokens to be passed as input to do teacher forcing.
#
# - [tl.Embedding](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.core.Embedding): Like in the previous function, this converts each token to its vector representation. In this case, it is the the size of the vocabulary by the dimension of the model: `tl.Embedding(vocab_size, d_model)`. `vocab_size` is the number of entries in the given vocabulary. `d_model` is the number of elements in the word embedding.
#
# - [tl.LSTM](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.rnn.LSTM): LSTM layer of size `d_model`.
#
# <img src = "pre_attention_decoder.png">
#
# <a name="ex02"></a>
# ### Exercise 02
#
# **Instructions:** Implement the `pre_attention_decoder_fn` function.
#
# %%
# UNQ_C2
# GRADED FUNCTION
def pre_attention_decoder_fn(mode, target_vocab_size, d_model):
""" Pre-attention decoder runs on the targets and creates
activations that are used as queries in attention.
Args:
mode: str: 'train' or 'eval'
target_vocab_size: int: vocab size of the target
d_model: int: depth of embedding (n_units in the LSTM cell)
Returns:
tl.Serial: The pre-attention decoder
"""
# create a serial network
pre_attention_decoder = tl.Serial(
### START CODE HERE (REPLACE INSTANCES OF `None` WITH YOUR CODE) ###
# shift right to insert start-of-sentence token and implement
# teacher forcing during training
tl.ShiftRight(mode=mode),
# run an embedding layer to convert tokens to vectors
tl.Embedding(target_vocab_size, d_model),
# feed to an LSTM layer
tl.LSTM(d_model)
### END CODE HERE ###
)
return pre_attention_decoder
# %%
# BEGIN UNIT TEST
w1_unittest.test_pre_attention_decoder_fn(pre_attention_decoder_fn)
# END UNIT TEST
# %% [markdown]
# ### 2.2.3 Preparing the attention input
#
# This function will prepare the inputs to the attention layer. We want to take in the encoder and pre-attention decoder activations and assign it to the queries, keys, and values. In addition, another output here will be the mask to distinguish real tokens from padding tokens. This mask will be used internally by Trax when computing the softmax so padding tokens will not have an effect on the computated probabilities. From the data preparation steps in Section 1 of this assignment, you should know which tokens in the input correspond to padding.
#
# We have filled the last two lines in composing the mask for you because it includes a concept that will be discussed further next week. This is related to *multiheaded attention* which you can think of right now as computing the attention multiple times to improve the model's predictions. It is required to consider this additional axis in the output so we've included it already but you don't need to analyze it just yet. What's important now is for you to know which should be the queries, keys, and values, as well as to initialize the mask.
#
# <a name="ex03"></a>
# ### Exercise 03
#
# **Instructions:** Implement the `prepare_attention_input` function
#
# %%
# UNQ_C3
# GRADED FUNCTION
def prepare_attention_input(encoder_activations, decoder_activations, inputs):
"""Prepare queries, keys, values and mask for attention.
Args:
encoder_activations fastnp.array(batch_size, padded_input_length, d_model): output from the input encoder
decoder_activations fastnp.array(batch_size, padded_input_length, d_model): output from the pre-attention decoder
inputs fastnp.array(batch_size, padded_input_length): padded input tokens
Returns:
queries, keys, values and mask for attention.
"""
### START CODE HERE (REPLACE INSTANCES OF `None` WITH YOUR CODE) ###
# set the keys and values to the encoder activations
keys = encoder_activations
values = encoder_activations
# set the queries to the decoder activations
queries = decoder_activations
# generate the mask to distinguish real tokens from padding
# hint: inputs is 1 for real tokens and 0 where they are padding
mask = (inputs != 0)
### END CODE HERE ###
# add axes to the mask for attention heads and decoder length.
mask = fastnp.reshape(mask, (mask.shape[0], 1, 1, mask.shape[1]))
# broadcast so mask shape is [batch size, attention heads, decoder-len, encoder-len].
# note: for this assignment, attention heads is set to 1.
mask = mask + fastnp.zeros((1, 1, decoder_activations.shape[1], 1))
return queries, keys, values, mask
# %%
# BEGIN UNIT TEST
w1_unittest.test_prepare_attention_input(prepare_attention_input)
# END UNIT TEST
# %% [markdown]
# <a name="2.3"></a>
# ## 2.3 Implementation Overview
#
# We are now ready to implement our sequence-to-sequence model with attention. This will be a Serial network and is illustrated in the diagram below. It shows the layers you'll be using in Trax and you'll see that each step can be implemented quite easily with one line commands. We've placed several links to the documentation for each relevant layer in the discussion after the figure below.
#
# <img src = "NMTModel.png">
# %% [markdown]
# <a name="ex04"></a>
# ### Exercise 04
# **Instructions:** Implement the `NMTAttn` function below to define your machine translation model which uses attention. We have left hyperlinks below pointing to the Trax documentation of the relevant layers. Remember to consult it to get tips on what parameters to pass.
#
# **Step 0:** Prepare the input encoder and pre-attention decoder branches. You have already defined this earlier as helper functions so it's just a matter of calling those functions and assigning it to variables.
#
# **Step 1:** Create a Serial network. This will stack the layers in the next steps one after the other. Like the earlier exercises, you can use [tl.Serial](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.combinators.Serial).
#
# **Step 2:** Make a copy of the input and target tokens. As you see in the diagram above, the input and target tokens will be fed into different layers of the model. You can use [tl.Select](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.combinators.Select) layer to create copies of these tokens. Arrange them as `[input tokens, target tokens, input tokens, target tokens]`.
#
# **Step 3:** Create a parallel branch to feed the input tokens to the `input_encoder` and the target tokens to the `pre_attention_decoder`. You can use [tl.Parallel](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.combinators.Parallel) to create these sublayers in parallel. Remember to pass the variables you defined in Step 0 as parameters to this layer.
#
# **Step 4:** Next, call the `prepare_attention_input` function to convert the encoder and pre-attention decoder activations to a format that the attention layer will accept. You can use [tl.Fn](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.base.Fn) to call this function. Note: Pass the `prepare_attention_input` function as the `f` parameter in `tl.Fn` without any arguments or parenthesis.
#
# **Step 5:** We will now feed the (queries, keys, values, and mask) to the [tl.AttentionQKV](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.attention.AttentionQKV) layer. This computes the scaled dot product attention and outputs the attention weights and mask. Take note that although it is a one liner, this layer is actually composed of a deep network made up of several branches. We'll show the implementation taken [here](https://github.com/google/trax/blob/master/trax/layers/attention.py#L61) to see the different layers used.
#
# ```python
# def AttentionQKV(d_feature, n_heads=1, dropout=0.0, mode='train'):
# """Returns a layer that maps (q, k, v, mask) to (activations, mask).
#
# See `Attention` above for further context/details.
#
# Args:
# d_feature: Depth/dimensionality of feature embedding.
# n_heads: Number of attention heads.
# dropout: Probababilistic rate for internal dropout applied to attention
# activations (based on query-key pairs) before dotting them with values.
# mode: Either 'train' or 'eval'.
# """
# return cb.Serial(
# cb.Parallel(
# core.Dense(d_feature),
# core.Dense(d_feature),
# core.Dense(d_feature),
# ),
# PureAttention( # pylint: disable=no-value-for-parameter
# n_heads=n_heads, dropout=dropout, mode=mode),
# core.Dense(d_feature),
# )
# ```
#
# Having deep layers pose the risk of vanishing gradients during training and we would want to mitigate that. To improve the ability of the network to learn, we can insert a [tl.Residual](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.combinators.Residual) layer to add the output of AttentionQKV with the `queries` input. You can do this in trax by simply nesting the `AttentionQKV` layer inside the `Residual` layer. The library will take care of branching and adding for you.
#
# **Step 6:** We will not need the mask for the model we're building so we can safely drop it. At this point in the network, the signal stack currently has `[attention activations, mask, target tokens]` and you can use [tl.Select](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.combinators.Select) to output just `[attention activations, target tokens]`.
#
# **Step 7:** We can now feed the attention weighted output to the LSTM decoder. We can stack multiple [tl.LSTM](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.rnn.LSTM) layers to improve the output so remember to append LSTMs equal to the number defined by `n_decoder_layers` parameter to the model.
#
# **Step 8:** We want to determine the probabilities of each subword in the vocabulary and you can set this up easily with a [tl.Dense](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.core.Dense) layer by making its size equal to the size of our vocabulary.
#
# **Step 9:** Normalize the output to log probabilities by passing the activations in Step 8 to a [tl.LogSoftmax](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.core.LogSoftmax) layer.
# %%
# UNQ_C4
# GRADED FUNCTION
def NMTAttn(input_vocab_size=33300,
target_vocab_size=33300,
d_model=1024,
n_encoder_layers=2,
n_decoder_layers=2,
n_attention_heads=4,
attention_dropout=0.0,
mode='train'):
"""Returns an LSTM sequence-to-sequence model with attention.
The input to the model is a pair (input tokens, target tokens), e.g.,
an English sentence (tokenized) and its translation into German (tokenized).
Args:
input_vocab_size: int: vocab size of the input
target_vocab_size: int: vocab size of the target
d_model: int: depth of embedding (n_units in the LSTM cell)
n_encoder_layers: int: number of LSTM layers in the encoder
n_decoder_layers: int: number of LSTM layers in the decoder after attention
n_attention_heads: int: number of attention heads
attention_dropout: float, dropout for the attention layer
mode: str: 'train', 'eval' or 'predict', predict mode is for fast inference
Returns:
A LSTM sequence-to-sequence model with attention.
"""
### START CODE HERE (REPLACE INSTANCES OF `None` WITH YOUR CODE) ###
# Step 0: call the helper function to create layers for the input encoder
input_encoder = input_encoder_fn(input_vocab_size, d_model, n_encoder_layers)
# Step 0: call the helper function to create layers for the pre-attention decoder
pre_attention_decoder = pre_attention_decoder_fn(mode, target_vocab_size, d_model)
# Step 1: create a serial network
model = tl.Serial(
# Step 2: copy input tokens and target tokens as they will be needed later.
tl.Select([0, 1, 0, 1]),
# Step 3: run input encoder on the input and pre-attention decoder the target.
tl.Parallel(input_encoder, pre_attention_decoder),
# Step 4: prepare queries, keys, values and mask for attention.
tl.Fn('PrepareAttentionInput', prepare_attention_input, n_out=4),
# Step 5: run the AttentionQKV layer
# nest it inside a Residual layer to add to the pre-attention decoder activations(i.e. queries)
tl.Residual(tl.AttentionQKV(d_model, n_heads=n_attention_heads, dropout=attention_dropout, mode=mode)),
# Step 6: drop attention mask (i.e. index = None
tl.Select([0, 2]),
# Step 7: run the rest of the RNN decoder
[tl.LSTM(d_model) for _ in range(n_decoder_layers)],
# Step 8: prepare output by making it the right size
tl.Dense(target_vocab_size),
# Step 9: Log-softmax for output
tl.LogSoftmax()
)
### END CODE HERE
return model
# %%
# BEGIN UNIT TEST
w1_unittest.test_NMTAttn(NMTAttn)
# END UNIT TEST
# %%
# print your model
model = NMTAttn()
print(model)
# %% [markdown]
# **Expected Output:**
#
# ```
# Serial_in2_out2[
# Select[0,1,0,1]_in2_out4
# Parallel_in2_out2[
# Serial[
# Embedding_33300_1024
# LSTM_1024
# LSTM_1024
# ]
# Serial[
# ShiftRight(1)
# Embedding_33300_1024
# LSTM_1024
# ]
# ]
# PrepareAttentionInput_in3_out4
# Serial_in4_out2[
# Branch_in4_out3[
# None
# Serial_in4_out2[
# Parallel_in3_out3[
# Dense_1024
# Dense_1024
# Dense_1024
# ]
# PureAttention_in4_out2
# Dense_1024
# ]
# ]
# Add_in2
# ]
# Select[0,2]_in3_out2
# LSTM_1024
# LSTM_1024
# Dense_33300
# LogSoftmax
# ]
# ```
# %% [markdown]
# <a name="3"></a>
# # Part 3: Training
#
# We will now be training our model in this section. Doing supervised training in Trax is pretty straightforward (short example [here](https://trax-ml.readthedocs.io/en/latest/notebooks/trax_intro.html#Supervised-training)). We will be instantiating three classes for this: `TrainTask`, `EvalTask`, and `Loop`. Let's take a closer look at each of these in the sections below.
#
# %% [markdown]
# <a name="3.1"></a>
# ## 3.1 TrainTask
#
# The [TrainTask](https://trax-ml.readthedocs.io/en/latest/trax.supervised.html#trax.supervised.training.TrainTask) class allows us to define the labeled data to use for training and the feedback mechanisms to compute the loss and update the weights.
#
# <a name="ex05"></a>
# ### Exercise 05
#
# **Instructions:** Instantiate a train task.
# %%
# UNQ_C5
# GRADED
train_task = training.TrainTask(
### START CODE HERE (REPLACE INSTANCES OF `None` WITH YOUR CODE) ###
# use the train batch stream as labeled data
labeled_data= train_batch_stream,
# use the cross entropy loss
loss_layer= tl.CrossEntropyLoss(),
# use the Adam optimizer with learning rate of 0.01
optimizer= trax.optimizers.Adam(.01),
# use the `trax.lr.warmup_and_rsqrt_decay` as the learning rate schedule
# have 1000 warmup steps with a max value of 0.01
lr_schedule= trax.lr.warmup_and_rsqrt_decay(1000, .01),
# have a checkpoint every 10 steps
n_steps_per_checkpoint= 10,
### END CODE HERE ###
)
# %%
# BEGIN UNIT TEST
w1_unittest.test_train_task(train_task)
# END UNIT TEST
# %% [markdown]
# <a name="3.2"></a>
# ## 3.2 EvalTask
#
# The [EvalTask](https://trax-ml.readthedocs.io/en/latest/trax.supervised.html#trax.supervised.training.EvalTask) on the other hand allows us to see how the model is doing while training. For our application, we want it to report the cross entropy loss and accuracy.
# %%
eval_task = training.EvalTask(
## use the eval batch stream as labeled data
labeled_data=eval_batch_stream,
## use the cross entropy loss and accuracy as metrics
metrics=[tl.CrossEntropyLoss(), tl.Accuracy()],
)
# %% [markdown]
# <a name="3.3"></a>
# ## 3.3 Loop
#
# The [Loop](https://trax-ml.readthedocs.io/en/latest/trax.supervised.html#trax.supervised.training.Loop) class defines the model we will train as well as the train and eval tasks to execute. Its `run()` method allows us to execute the training for a specified number of steps.
# %%
# define the output directory
output_dir = 'output_dir/'
# remove old model if it exists. restarts training.
# !rm -f ~/output_dir/model.pkl.gz
# define the training loop
training_loop = training.Loop(NMTAttn(mode='train'),
train_task,
eval_tasks=[eval_task],
output_dir=output_dir)
# %%
# NOTE: Execute the training loop. This will take around 8 minutes to complete.
training_loop.run(10)
# %% [markdown]
# <a name="4"></a>
# # Part 4: Testing
#
# We will now be using the model you just trained to translate English sentences to German. We will implement this with two functions: The first allows you to identify the next symbol (i.e. output token). The second one takes care of combining the entire translated string.
#
# We will start by first loading in a pre-trained copy of the model you just coded. Please run the cell below to do just that.
# %%
# instantiate the model we built in eval mode
model = NMTAttn(mode='eval')
# initialize weights from a pre-trained model
model.init_from_file("model.pkl.gz", weights_only=True)
model = tl.Accelerate(model)
# %% [markdown]
# <a name="4.1"></a>
# ## 4.1 Decoding
#
# As discussed in the lectures, there are several ways to get the next token when translating a sentence. For instance, we can just get the most probable token at each step (i.e. greedy decoding) or get a sample from a distribution. We can generalize the implementation of these two approaches by using the `tl.logsoftmax_sample()` method. Let's briefly look at its implementation:
#
# ```python
# def logsoftmax_sample(log_probs, temperature=1.0): # pylint: disable=invalid-name
# """Returns a sample from a log-softmax output, with temperature.
#
# Args:
# log_probs: Logarithms of probabilities (often coming from LogSofmax)
# temperature: For scaling before sampling (1.0 = default, 0.0 = pick argmax)
# """
# # This is equivalent to sampling from a softmax with temperature.
# u = np.random.uniform(low=1e-6, high=1.0 - 1e-6, size=log_probs.shape)
# g = -np.log(-np.log(u))
# return np.argmax(log_probs + g * temperature, axis=-1)
# ```
#
# The key things to take away here are: 1. it gets random samples with the same shape as your input (i.e. `log_probs`), and 2. the amount of "noise" added to the input by these random samples is scaled by a `temperature` setting. You'll notice that setting it to `0` will just make the return statement equal to getting the argmax of `log_probs`. This will come in handy later.
#
# <a name="ex06"></a>
# ### Exercise 06
#
# **Instructions:** Implement the `next_symbol()` function that takes in the `input_tokens` and the `cur_output_tokens`, then return the index of the next word. You can click below for hints in completing this exercise.
#
# <details>
# <summary>
# <font size="3" color="darkgreen"><b>Click Here for Hints</b></font>
# </summary>
# <p>
# <ul>
# <li>To get the next power of two, you can compute <i>2^log_2(token_length + 1)</i> . We add 1 to avoid <i>log(0).</i></li>
# <li>You can use <i>np.ceil()</i> to get the ceiling of a float.</li>
# <li><i>np.log2()</i> will get the logarithm base 2 of a value</li>
# <li><i>int()</i> will cast a value into an integer type</li>
# <li>From the model diagram in part 2, you know that it takes two inputs. You can feed these with this syntax to get the model outputs: <i>model((input1, input2))</i>. It's up to you to determine which variables below to substitute for input1 and input2. Remember also from the diagram that the output has two elements: [log probabilities, target tokens]. You won't need the target tokens so we assigned it to _ below for you. </li>
# <li> The log probabilities output will have the shape: (batch size, decoder length, vocab size). It will contain log probabilities for each token in the <i>cur_output_tokens</i> plus 1 for the start symbol introduced by the ShiftRight in the preattention decoder. For example, if cur_output_tokens is [1, 2, 5], the model will output an array of log probabilities each for tokens 0 (start symbol), 1, 2, and 5. To generate the next symbol, you just want to get the log probabilities associated with the last token (i.e. token 5 at index 3). You can slice the model output at [0, 3, :] to get this. It will be up to you to generalize this for any length of cur_output_tokens </li>
# </ul>
#
# %%
# UNQ_C6
# GRADED FUNCTION
def next_symbol(NMTAttn, input_tokens, cur_output_tokens, temperature):
"""Returns the index of the next token.
Args:
NMTAttn (tl.Serial): An LSTM sequence-to-sequence model with attention.
input_tokens (np.ndarray 1 x n_tokens): tokenized representation of the input sentence
cur_output_tokens (list): tokenized representation of previously translated words
temperature (float): parameter for sampling ranging from 0.0 to 1.0.
0.0: same as argmax, always pick the most probable token
1.0: sampling from the distribution (can sometimes say random things)
Returns:
int: index of the next token in the translated sentence
float: log probability of the next symbol
"""
### START CODE HERE (REPLACE INSTANCES OF `None` WITH YOUR CODE) ###
# set the length of the current output tokens
token_length = len(cur_output_tokens)
# calculate next power of 2 for padding length
padded_length = 2**int(np.ceil(np.log2(token_length + 1)))
# pad cur_output_tokens up to the padded_length
padded = cur_output_tokens + [0] * (padded_length - token_length)
# model expects the output to have an axis for the batch size in front so
# convert `padded` list to a numpy array with shape (None, <padded_length>) where
# None is a placeholder for the batch size
padded_with_batch = np.expand_dims(padded, axis=0)
# get the model prediction (remember to use the `NMAttn` argument defined above)
output, _ = NMTAttn((input_tokens, padded_with_batch))
# get log probabilities from the last token output
log_probs = output[0, token_length, :]
# get the next symbol by getting a logsoftmax sample (*hint: cast to an int)
symbol = int(tl.logsoftmax_sample(log_probs, temperature))
### END CODE HERE ###
return symbol, float(log_probs[symbol])
# %%
# BEGIN UNIT TEST
w1_unittest.test_next_symbol(next_symbol, model)
# END UNIT TEST
# %% [markdown]
# Now you will implement the `sampling_decode()` function. This will call the `next_symbol()` function above several times until the next output is the end-of-sentence token (i.e. `EOS`). It takes in an input string and returns the translated version of that string.
#
# <a name="ex07"></a>
# ### Exercise 07
#
# **Instructions**: Implement the `sampling_decode()` function.
# %%
# UNQ_C7
# GRADED FUNCTION
def sampling_decode(input_sentence, NMTAttn = None, temperature=0.0, vocab_file=None, vocab_dir=None):
"""Returns the translated sentence.
Args:
input_sentence (str): sentence to translate.
NMTAttn (tl.Serial): An LSTM sequence-to-sequence model with attention.
temperature (float): parameter for sampling ranging from 0.0 to 1.0.
0.0: same as argmax, always pick the most probable token
1.0: sampling from the distribution (can sometimes say random things)
vocab_file (str): filename of the vocabulary
vocab_dir (str): path to the vocabulary file
Returns:
tuple: (list, str, float)
list of int: tokenized version of the translated sentence
float: log probability of the translated sentence
str: the translated sentence
"""
### START CODE HERE (REPLACE INSTANCES OF `None` WITH YOUR CODE) ###
# encode the input sentence
input_tokens = tokenize(input_sentence,vocab_file,vocab_dir)
# initialize the list of output tokens
cur_output_tokens = []
# initialize an integer that represents the current output index
cur_output = 0
# Set the encoding of the "end of sentence" as 1
EOS = 1
# check that the current output is not the end of sentence token
while cur_output != EOS:
# update the current output token by getting the index of the next word (hint: use next_symbol)
cur_output, log_prob = next_symbol(NMTAttn, input_tokens, cur_output_tokens, temperature)
# append the current output token to the list of output tokens
cur_output_tokens.append(cur_output)
# detokenize the output tokens
sentence = detokenize(cur_output_tokens, vocab_file, vocab_dir)
### END CODE HERE ###
return cur_output_tokens, log_prob, sentence
# %%
# Test the function above. Try varying the temperature setting with values from 0 to 1.
# Run it several times with each setting and see how often the output changes.
sampling_decode("I love languages.", model, temperature=0.0, vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR)
# %%
# BEGIN UNIT TEST
w1_unittest.test_sampling_decode(sampling_decode, model)
# END UNIT TEST
# %% [markdown]
# We have set a default value of `0` to the temperature setting in our implementation of `sampling_decode()` above. As you may have noticed in the `logsoftmax_sample()` method, this setting will ultimately result in greedy decoding. As mentioned in the lectures, this algorithm generates the translation by getting the most probable word at each step. It gets the argmax of the output array of your model and then returns that index. See the testing function and sample inputs below. You'll notice that the output will remain the same each time you run it.
# %%
def greedy_decode_test(sentence, NMTAttn=None, vocab_file=None, vocab_dir=None):
"""Prints the input and output of our NMTAttn model using greedy decode
Args:
sentence (str): a custom string.
NMTAttn (tl.Serial): An LSTM sequence-to-sequence model with attention.
vocab_file (str): filename of the vocabulary
vocab_dir (str): path to the vocabulary file
Returns:
str: the translated sentence
"""
_,_, translated_sentence = sampling_decode(sentence, NMTAttn, vocab_file=vocab_file, vocab_dir=vocab_dir)
print("English: ", sentence)
print("German: ", translated_sentence)
return translated_sentence
# %%
# put a custom string here
your_sentence = 'I love languages.'
greedy_decode_test(your_sentence, model, vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR);
# %%
greedy_decode_test('You are almost done with the assignment!', model, vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR);
# %% [markdown]
# <a name="4.2"></a>
# ## 4.2 Minimum Bayes-Risk Decoding
#
# As mentioned in the lectures, getting the most probable token at each step may not necessarily produce the best results. Another approach is to do Minimum Bayes Risk Decoding or MBR. The general steps to implement this are:
#
# 1. take several random samples
# 2. score each sample against all other samples
# 3. select the one with the highest score
#
# You will be building helper functions for these steps in the following sections.
# %% [markdown]
# <a name='4.2.1'></a>
# ### 4.2.1 Generating samples
#
# First, let's build a function to generate several samples. You can use the `sampling_decode()` function you developed earlier to do this easily. We want to record the token list and log probability for each sample as these will be needed in the next step.
# %%
def generate_samples(sentence, n_samples, NMTAttn=None, temperature=0.6, vocab_file=None, vocab_dir=None):
"""Generates samples using sampling_decode()
Args:
sentence (str): sentence to translate.
n_samples (int): number of samples to generate
NMTAttn (tl.Serial): An LSTM sequence-to-sequence model with attention.
temperature (float): parameter for sampling ranging from 0.0 to 1.0.
0.0: same as argmax, always pick the most probable token
1.0: sampling from the distribution (can sometimes say random things)
vocab_file (str): filename of the vocabulary
vocab_dir (str): path to the vocabulary file
Returns:
tuple: (list, list)
list of lists: token list per sample
list of floats: log probability per sample
"""
# define lists to contain samples and probabilities
samples, log_probs = [], []
# run a for loop to generate n samples
for _ in range(n_samples):
# get a sample using the sampling_decode() function
sample, logp, _ = sampling_decode(sentence, NMTAttn, temperature, vocab_file=vocab_file, vocab_dir=vocab_dir)
# append the token list to the samples list
samples.append(sample)
# append the log probability to the log_probs list
log_probs.append(logp)
return samples, log_probs
# %%
# generate 4 samples with the default temperature (0.6)
generate_samples('I love languages.', 4, model, vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR)
# %% [markdown]
# ### 4.2.2 Comparing overlaps
#
# Let us now build our functions to compare a sample against another. There are several metrics available as shown in the lectures and you can try experimenting with any one of these. For this assignment, we will be calculating scores for unigram overlaps. One of the more simple metrics is the [Jaccard similarity](https://en.wikipedia.org/wiki/Jaccard_index) which gets the intersection over union of two sets. We've already implemented it below for your perusal.
# %%
def jaccard_similarity(candidate, reference):
"""Returns the Jaccard similarity between two token lists
Args:
candidate (list of int): tokenized version of the candidate translation
reference (list of int): tokenized version of the reference translation
Returns:
float: overlap between the two token lists
"""
# convert the lists to a set to get the unique tokens
can_unigram_set, ref_unigram_set = set(candidate), set(reference)
# get the set of tokens common to both candidate and reference
joint_elems = can_unigram_set.intersection(ref_unigram_set)
# get the set of all tokens found in either candidate or reference
all_elems = can_unigram_set.union(ref_unigram_set)
# divide the number of joint elements by the number of all elements
overlap = len(joint_elems) / len(all_elems)
return overlap
# %%
# let's try using the function. remember the result here and compare with the next function below.
jaccard_similarity([1, 2, 3], [1, 2, 3, 4])
# %% [markdown]
# One of the more commonly used metrics in machine translation is the ROUGE score. For unigrams, this is called ROUGE-1 and as shown in class, you can output the scores for both precision and recall when comparing two samples. To get the final score, you will want to compute the F1-score as given by:
#
# $$score = 2* \frac{(precision * recall)}{(precision + recall)}$$
#
# <a name="ex08"></a>
# ### Exercise 08
#
# **Instructions**: Implement the `rouge1_similarity()` function.
# %%
# UNQ_C8
# GRADED FUNCTION
# for making a frequency table easily
from collections import Counter
def rouge1_similarity(system, reference):
"""Returns the ROUGE-1 score between two token lists
Args:
system (list of int): tokenized version of the system translation
reference (list of int): tokenized version of the reference translation
Returns:
float: overlap between the two token lists
"""
### START CODE HERE (REPLACE INSTANCES OF `None` WITH YOUR CODE) ###
# make a frequency table of the system tokens (hint: use the Counter class)
sys_counter = Counter(system)
# make a frequency table of the reference tokens (hint: use the Counter class)
ref_counter = Counter(reference)
# initialize overlap to 0
overlap = 0
# run a for loop over the sys_counter object (can be treated as a dictionary)
for token in sys_counter:
# lookup the value of the token in the sys_counter dictionary (hint: use the get() method)
token_count_sys = sys_counter.get(token,0)
# lookup the value of the token in the ref_counter dictionary (hint: use the get() method)
token_count_ref = ref_counter.get(token,0)
# update the overlap by getting the smaller number between the two token counts above
overlap += min(token_count_sys, token_count_ref)
# get the precision (i.e. number of overlapping tokens / number of system tokens)
precision = overlap / sum(sys_counter.values())
# get the recall (i.e. number of overlapping tokens / number of reference tokens)
recall = overlap / sum(ref_counter.values())
if precision + recall != 0:
# compute the f1-score
rouge1_score = 2 * ((precision * recall)/(precision + recall))
else:
rouge1_score = 0
### END CODE HERE ###
return rouge1_score
# %%
# notice that this produces a different value from the jaccard similarity earlier
rouge1_similarity([1, 2, 3], [1, 2, 3, 4])
# %%
# BEGIN UNIT TEST
w1_unittest.test_rouge1_similarity(rouge1_similarity)
# END UNIT TEST
# %% [markdown]
# ### 4.2.3 Overall score
#
# We will now build a function to generate the overall score for a particular sample. As mentioned earlier, we need to compare each sample with all other samples. For instance, if we generated 30 sentences, we will need to compare sentence 1 to sentences 2 to 30. Then, we compare sentence 2 to sentences 1 and 3 to 30, and so forth. At each step, we get the average score of all comparisons to get the overall score for a particular sample. To illustrate, these will be the steps to generate the scores of a 4-sample list.
#
# 1. Get similarity score between sample 1 and sample 2
# 2. Get similarity score between sample 1 and sample 3
# 3. Get similarity score between sample 1 and sample 4
# 4. Get average score of the first 3 steps. This will be the overall score of sample 1.
# 5. Iterate and repeat until samples 1 to 4 have overall scores.
#
# We will be storing the results in a dictionary for easy lookups.
#
# <a name="ex09"></a>
# ### Exercise 09
#
# **Instructions**: Implement the `average_overlap()` function.
# %%
# UNQ_C9
# GRADED FUNCTION
def average_overlap(similarity_fn, samples, *ignore_params):
"""Returns the arithmetic mean of each candidate sentence in the samples
Args:
similarity_fn (function): similarity function used to compute the overlap
samples (list of lists): tokenized version of the translated sentences
*ignore_params: additional parameters will be ignored
Returns:
dict: scores of each sample
key: index of the sample
value: score of the sample
"""
# initialize dictionary
scores = {}
# run a for loop for each sample
for index_candidate, candidate in enumerate(samples):
### START CODE HERE (REPLACE INSTANCES OF `None` WITH YOUR CODE) ###
# initialize overlap to 0.0
overlap = 0.0
# run a for loop for each sample
for index_sample, sample in enumerate(samples):
# skip if the candidate index is the same as the sample index
if index_candidate == index_sample:
continue
# get the overlap between candidate and sample using the similarity function
sample_overlap = similarity_fn(candidate,sample)
# add the sample overlap to the total overlap
overlap += sample_overlap
# get the score for the candidate by computing the average
score = overlap/index_sample
# save the score in the dictionary. use index as the key.
scores[index_candidate] = score
### END CODE HERE ###
return scores
# %%
average_overlap(jaccard_similarity, [[1, 2, 3], [1, 2, 4], [1, 2, 4, 5]], [0.4, 0.2, 0.5])
# %%
# BEGIN UNIT TEST
w1_unittest.test_average_overlap(average_overlap)
# END UNIT TEST
# %% [markdown]
# In practice, it is also common to see the weighted mean being used to calculate the overall score instead of just the arithmetic mean. We have implemented it below and you can use it in your experiements to see which one will give better results.
# %%
def weighted_avg_overlap(similarity_fn, samples, log_probs):
"""Returns the weighted mean of each candidate sentence in the samples
Args:
samples (list of lists): tokenized version of the translated sentences
log_probs (list of float): log probability of the translated sentences
Returns:
dict: scores of each sample
key: index of the sample
value: score of the sample
"""
# initialize dictionary
scores = {}
# run a for loop for each sample
for index_candidate, candidate in enumerate(samples):
# initialize overlap and weighted sum
overlap, weight_sum = 0.0, 0.0
# run a for loop for each sample
for index_sample, (sample, logp) in enumerate(zip(samples, log_probs)):
# skip if the candidate index is the same as the sample index
if index_candidate == index_sample:
continue
# convert log probability to linear scale
sample_p = float(np.exp(logp))
# update the weighted sum
weight_sum += sample_p
# get the unigram overlap between candidate and sample
sample_overlap = similarity_fn(candidate, sample)
# update the overlap
overlap += sample_p * sample_overlap
# get the score for the candidate
score = overlap / weight_sum
# save the score in the dictionary. use index as the key.
scores[index_candidate] = score
return scores
# %%
weighted_avg_overlap(jaccard_similarity, [[1, 2, 3], [1, 2, 4], [1, 2, 4, 5]], [0.4, 0.2, 0.5])
# %% [markdown]
# ### 4.2.4 Putting it all together
#
# We will now put everything together and develop the `mbr_decode()` function. Please use the helper functions you just developed to complete this. You will want to generate samples, get the score for each sample, get the highest score among all samples, then detokenize this sample to get the translated sentence.
#
# <a name="ex10"></a>
# ### Exercise 10
#
# **Instructions**: Implement the `mbr_overlap()` function.
# %%
# UNQ_C10
# GRADED FUNCTION
def mbr_decode(sentence, n_samples, score_fn, similarity_fn, NMTAttn=None, temperature=0.6, vocab_file=None, vocab_dir=None):
"""Returns the translated sentence using Minimum Bayes Risk decoding
Args:
sentence (str): sentence to translate.
n_samples (int): number of samples to generate
score_fn (function): function that generates the score for each sample
similarity_fn (function): function used to compute the overlap between a pair of samples
NMTAttn (tl.Serial): An LSTM sequence-to-sequence model with attention.
temperature (float): parameter for sampling ranging from 0.0 to 1.0.
0.0: same as argmax, always pick the most probable token
1.0: sampling from the distribution (can sometimes say random things)
vocab_file (str): filename of the vocabulary
vocab_dir (str): path to the vocabulary file
Returns:
str: the translated sentence
"""
### START CODE HERE (REPLACE INSTANCES OF `None` WITH YOUR CODE) ###
# generate samples
samples, log_probs = generate_samples(sentence, n_samples, NMTAttn, temperature, vocab_file, vocab_dir)
# use the scoring function to get a dictionary of scores
# pass in the relevant parameters as shown in the function definition of
# the mean methods you developed earlier
scores = score_fn(similarity_fn, samples, log_probs )
# find the key with the highest score
max_index = max(scores, key=scores.get)
# detokenize the token list associated with the max_index
translated_sentence = detokenize(samples[max_index], vocab_file, vocab_dir)
### END CODE HERE ###
return (translated_sentence, max_index, scores)
# %%
TEMPERATURE = 1.0
# put a custom string here
your_sentence = 'She speaks English and German.'
# %%
mbr_decode(your_sentence, 4, weighted_avg_overlap, jaccard_similarity, model, TEMPERATURE, vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR)[0]
# %%
mbr_decode('Congratulations!', 4, average_overlap, rouge1_similarity, model, TEMPERATURE, vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR)[0]
# %%
mbr_decode('You have completed the assignment!', 4, average_overlap, rouge1_similarity, model, TEMPERATURE, vocab_file=VOCAB_FILE, vocab_dir=VOCAB_DIR)[0]
# %% [markdown]
# **This unit test take a while to run. Please be patient**
# %%
# BEGIN UNIT TEST
w1_unittest.test_mbr_decode(mbr_decode, model)
# END UNIT TEST
# %% [markdown]
# #### Congratulations! Next week, you'll dive deeper into attention models and study the Transformer architecture. You will build another network but without the recurrent part. It will show that attention is all you need! It should be fun!
| Course4/Week1/C4W1Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re
p_text = ' 新浪财经讯 12月6日,由中国(深圳)综合开发研究院和国家发展和改革委员会国际合作中心共同举办的“2019协同创新高端论坛”在北京召开。会议重点研讨如何构建新型城乡关系,促进乡村振兴和农业农村现代化,以推动中国经济社会的高质量与和谐发展。'
print(p_text.startswith('\u3000\u3000新浪财经讯 '))
#p_text = '\u3000\u3000' + p_text[len('\u3000\u3000新浪财经讯 '):]
p_text = re.sub('^\u3000\u3000新浪.{0,6}讯 ', '\u3000\u3000', p_text)
p_text
from lxml.html import builder as E
from lxml import etree
import lxml.html
import logging
html = E.HTML(
E.HEAD(
E.META(content='text/html', charset='utf-8'),
E.LINK(rel='stylesheet', href='../css/style.css', type='text/css'),
E.TITLE(E.CLASS('title'), f'实时新闻摘要')
)
)
body = etree.SubElement(html, 'body')
print(f'html: {lxml.html.tostring(html, pretty_print=True, encoding="utf-8").decode("utf-8")}')
| jupyter/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.style as style
import numpy as np
import pandas as pd
import plotly.express as px
import seaborn as sns
from scipy import stats
import warnings
warnings.filterwarnings("ignore")
data = pd.read_csv('./train-data.csv')
test = pd.read_csv('./test-data.csv')
# +
from lightgbm import LGBMClassifier
from sklearn import metrics
from sklearn import model_selection
from sklearn import preprocessing
from sklearn.datasets import make_classification
from sklearn.ensemble import ExtraTreesRegressor
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import RandomForestRegressor
from sklearn.linear_model import LogisticRegression
from sklearn.linear_model import Ridge
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report, confusion_matrix
from sklearn.metrics import mean_squared_error
from sklearn.metrics import mean_squared_error, mean_absolute_error
from sklearn.metrics import plot_confusion_matrix
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import GaussianNB
from sklearn.neighbors import KNeighborsClassifier
from sklearn.neighbors import KNeighborsRegressor
from sklearn.preprocessing import StandardScaler, Normalizer
from sklearn.tree import DecisionTreeClassifier
from sklearn.tree import DecisionTreeRegressor
from xgboost import XGBClassifier
from xgboost.sklearn import XGBRegressor
from sklearn.ensemble import GradientBoostingRegressor,AdaBoostRegressor,BaggingRegressor, RandomForestRegressor
from sklearn.neural_network import MLPRegressor
import xgboost as xgb
# -
data.isnull().sum()
test.isnull().sum()
# +
data = data.drop('New_Price', axis=1)
data = data.drop('Unnamed: 0', axis=1)
test = test.drop('New_Price', axis=1)
test = test.drop('Unnamed: 0', axis=1)
# -
data = data.dropna(how='any')
test = test.dropna(how='any')
data.shape
test.shape
# +
listtrain = data['Name']
listtest = test['Name']
# prints the missing in listrain
print("Missing values in first list:", (set(listtest).difference(listtrain)))
# -
data['Cars'] = data['Name'].str.split(" ").str[0] + ' ' +data['Name'].str.split(" ").str[1]
test['Cars'] = test['Name'].str.split(" ").str[0] + ' ' +test['Name'].str.split(" ").str[1]
set(test['Cars']).issubset(set(data['Cars']))
# +
listtrain = data['Cars']
listtest = test['Cars']
# prints the missing and additional elements in list1
print("Missing values in first list:", (set(listtest).difference(listtrain)))
# -
test.drop(test[test['Cars'].isin(['Toyota Land', 'Hindustan Motors', 'Fiat Abarth', 'Nissan 370Z',
'Isuzu MU', 'Bentley Flying', 'OpelCorsa 1.4Gsi'])].index, inplace = True)
test.shape
# +
listtrain = data['Cars']
listtest = test['Cars']
# prints the missing and additional elements in list1
print("Missing values in first list:", (set(listtest).difference(listtrain)))
# -
test.drop(test[test['Cars'].isin(['Toyota Land', 'Hindustan Motors', 'Fiat Abarth', 'Nissan 370Z',
'Isuzu MU', 'B<NAME>', 'OpelCorsa 1.4Gsi'])].index, inplace = True)
test.shape
# +
listtrain = data['Cars']
listtest = test['Cars']
# prints the missing and additional elements in list1
print("Missing values in first list:", (set(listtest).difference(listtrain)))
# -
data.head(3)
# +
data['Mileage'] = data['Mileage'].str.replace(' kmpl','')
data['Mileage'] = data['Mileage'].str.replace(' km/kg','')
data['Engine'] = data['Engine'].str.replace(' CC','')
data['Power'] = data['Power'].str.replace('null bhp','112')
data['Power'] = data['Power'].str.replace(' bhp','')
test['Mileage'] = test['Mileage'].str.replace(' kmpl','')
test['Mileage'] = test['Mileage'].str.replace(' km/kg','')
test['Engine'] = test['Engine'].str.replace(' CC','')
test['Power'] = test['Power'].str.replace('null bhp','112')
test['Power'] = test['Power'].str.replace(' bhp','')
# -
data.isnull().sum()
test.isnull().sum()
data.dtypes
# +
data['Mileage'] = data['Mileage'].astype(float)
data['Mileage'] = data['Mileage'].astype(float)
data['Engine'] = data['Engine'].astype(float)
data['Power'] = data['Power'].astype(float)
test['Mileage'] = test['Mileage'].astype(float)
test['Mileage'] = test['Mileage'].astype(float)
test['Engine'] = test['Engine'].astype(float)
test['Power'] = test['Power'].astype(float)
# -
data.describe()
# +
feature = ['Cars', 'Location', 'Year', 'Kilometers_Driven', 'Fuel_Type', 'Transmission',
'Owner_Type', 'Mileage', 'Engine', 'Power', 'Seats','Price']
data = pd.DataFrame(data, columns=feature)
feature1 = ['Cars', 'Location', 'Year', 'Kilometers_Driven', 'Fuel_Type', 'Transmission',
'Owner_Type', 'Mileage', 'Engine', 'Power', 'Seats']
test = pd.DataFrame(test, columns=feature1)
# -
data.head(3)
# +
sns.distplot(data['Price'])
print("Skewness: %f" % data['Price'].skew())
print("Kurtosis: %f" % data['Price'].kurt())
# -
#applying log transformation
data['Price'] = np.log(data['Price'])
#transformed histogram and normal probability plot
#sns.distplot(data['Price']);
sns.distplot(data['Price'], fit=None);
fig = plt.figure()
res = stats.probplot(data['Price'], plot=plt)
# RELATIONSHIP OF PRICE WITH OTHER PARAMETER
# Find most important features relative to target Price
print("Find most important features relative to Price-target")
corr = data.corr()
corr.sort_values(["Price"], ascending = False, inplace = True)
print(corr.Price)
px.treemap(data.groupby(by='Fuel_Type').sum().reset_index(), path=['Fuel_Type'], labels='Fuel_Type',
values='Price', title='Price vs Fuel_type')
yprop = 'Price'
xprop = 'Power'
h= 'Fuel_Type'
px.scatter(data, x=xprop, y=yprop, color=h, marginal_y="violin", marginal_x="box", trendline="ols", template="simple_white")
plt.figure(figsize=(15,10))
xprop = 'Year'
yprop = 'Price'
sns.boxplot(data=data, x=xprop, y=yprop, hue='Transmission')
plt.xlabel('{} range'.format(xprop), size=14)
plt.ylabel('Number of {}'.format(yprop), size=14)
plt.title('Boxplot of {}'.format(yprop), size=20)
plt.show()
yprop = 'Price'
xprop = 'Year'
h= 'Owner_Type'
px.scatter(data, x=xprop, y=yprop, color=h, marginal_y="violin", marginal_x="box", trendline="ols", template="simple_white")
#fig.update_layout(xaxis_range=[0,5e5])
plt.figure(figsize=(15,10))
xprop = 'Year'
yprop = 'Price'
sns.boxplot(data=data, x=xprop, y=yprop, hue='Fuel_Type')
plt.xlabel('{} range'.format(xprop), size=14)
plt.ylabel('Number of {}'.format(yprop), size=14)
plt.title('Boxplot of {}'.format(yprop), size=20)
plt.show()
fig = px.box(data, x='Fuel_Type',y='Price', color='Transmission', notched=True)
fig.update_layout(legend=dict(orientation="h",yanchor="bottom",y=1.02,xanchor="right",x=1))
fig.show()
px.violin(data, y='Price', x='Seats', color=None, box=True, points="all", hover_data=data.columns)
import plotly.graph_objects as go
fig = go.Figure(data=[go.Pie(labels=data['Fuel_Type'], values=data['Price'], hole=.3)])
fig.update_layout(legend=dict(orientation="h", yanchor="bottom",y=1.02,xanchor="right",x=1))
fig.show()
# IV. MODEL DESCRIPTION
# +
import copy
df_train=copy.deepcopy(data)
df_test=copy.deepcopy(test)
cols=np.array(data.columns[data.dtypes != object])
for i in df_train.columns:
if i not in cols:
df_train[i]=df_train[i].map(str)
df_test[i]=df_test[i].map(str)
df_train.drop(columns=cols,inplace=True)
df_test.drop(columns=np.delete(cols,len(cols)-1),inplace=True)
# +
from sklearn.preprocessing import LabelEncoder
from collections import defaultdict
# build dictionary function
cols=np.array(data.columns[data.dtypes != object])
d = defaultdict(LabelEncoder)
# only for categorical columns apply dictionary by calling fit_transform
df_train = df_train.apply(lambda x: d[x.name].fit_transform(x))
df_test = df_test.apply(lambda x: d[x.name].transform(x))
df_train[cols] = data[cols]
df_test[np.delete(cols,len(cols)-1)]=test[np.delete(cols,len(cols)-1)]
# -
df_test.head(2)
df_train.head(2)
# B. Training and Testing
# +
ftrain = ['Cars', 'Location', 'Year', 'Kilometers_Driven', 'Fuel_Type', 'Transmission',
'Owner_Type', 'Mileage', 'Engine', 'Power', 'Seats','Price']
def Definedata():
# define dataset
data2 = df_train[ftrain]
X = data2.drop(columns=['Price']).values
y0 = data2['Price'].values
lab_enc = preprocessing.LabelEncoder()
y = lab_enc.fit_transform(y0)
return X, y
# +
def Models(models):
model = models
X, y = Definedata()
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.33, random_state = 25)
model.fit(X_train,y_train)
y_pred = model.predict(X_test)
y_total = model.predict(X)
print("\t\tError Table")
print('Mean Absolute Error : ', metrics.mean_absolute_error(y_test, y_pred))
print('Mean Squared Error : ', metrics.mean_squared_error(y_test, y_pred))
print('Root Mean Squared Error : ', np.sqrt(metrics.mean_squared_error(y_test, y_pred)))
print('Accuracy on Traing set : ', model.score(X_train,y_train))
print('Accuracy on Testing set : ', model.score(X_test,y_test))
return y_total, y
def Featureimportances(models):
model = models
model.fit(X_train,y_train)
importances = model.feature_importances_
features = df_test.columns[:9]
imp = pd.DataFrame({'Features': ftest, 'Importance': importances})
imp['Sum Importance'] = imp['Importance'].cumsum()
imp = imp.sort_values(by = 'Importance')
return imp
def Graph_prediction(n, y_actual, y_predicted):
y = y_actual
y_total = y_predicted
number = n
aa=[x for x in range(number)]
plt.figure(figsize=(25,10))
plt.plot(aa, y[:number], marker='.', label="actual")
plt.plot(aa, y_total[:number], 'b', label="prediction")
plt.xlabel('Price prediction of first {} used cars'.format(number), size=15)
plt.legend(fontsize=15)
plt.show()
# -
style.use('ggplot')
sns.set_style('whitegrid')
plt.subplots(figsize = (12,7))
## Plotting heatmap. # Generate a mask for the upper triangle (taken from seaborn example gallery)
mask = np.zeros_like(df_train.corr(), dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
sns.heatmap(df_train.corr(), cmap=sns.diverging_palette(20, 220, n=200), annot=True, mask=mask, center = 0, );
plt.title("Heatmap of all the Features of Train data set", fontsize = 25);
# C. Models comparison
Acc = pd.DataFrame(index=None, columns=['model','Root Mean Squared Error','Accuracy on Traing set','Accuracy on Testing set'])
# +
X, y = Definedata()
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.33, random_state = 25)
regressors = [['DecisionTreeRegressor',DecisionTreeRegressor()],
['XGBRegressor', XGBRegressor()],
['RandomForestRegressor', RandomForestRegressor()],
['MLPRegressor',MLPRegressor()],
['AdaBoostRegressor',AdaBoostRegressor()],
['ExtraTreesRegressor',ExtraTreesRegressor()]]
for mod in regressors:
name = mod[0]
model = mod[1]
model.fit(X_train,y_train)
y_pred = model.predict(X_test)
RMSE = np.sqrt(metrics.mean_squared_error(y_test, y_pred))
ATrS = model.score(X_train,y_train)
ATeS = model.score(X_test,y_test)
Acc = Acc.append(pd.Series({'model':name, 'Root Mean Squared Error': RMSE,'Accuracy on Traing set':ATrS,'Accuracy on Testing set':ATeS}),ignore_index=True )
# -
Acc.sort_values(by='Accuracy on Testing set')
y_predicted, y_actual = Models(RandomForestRegressor(n_estimators=10000,min_samples_split=2,min_samples_leaf=1,max_features='sqrt',max_depth=25))
Graph_prediction(150, y_actual, y_predicted)
y_predicted, y_actual = Models(GradientBoostingRegressor(random_state=21, n_estimators=3000))
Graph_prediction(150, y_actual, y_predicted)
compare = pd.DataFrame({'Prediction': y_predicted, 'Test Data' : y_actual, 'Abs error': abs(y_actual - y_predicted), 'AAD%': abs(y_actual - y_predicted)/y_actual*100})
compare.head(10)
# +
model = GradientBoostingRegressor(random_state=21, n_estimators=5000)
feature1 = ['Cars', 'Location', 'Year', 'Kilometers_Driven', 'Fuel_Type', 'Transmission',
'Owner_Type', 'Mileage', 'Engine', 'Power', 'Seats']
X0 = df_test[feature1]
X, y = Definedata()
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.25, random_state = 25)
model.fit(X_train,y_train)
y_predicted = model.predict(X0)
submission = pd.DataFrame({'Car_id':test.index,'Price':y_predicted})
submission.head(10)
# +
#Convert DataFrame to a csv file that can be uploaded
#This is saved in the same directory as your notebook
filename = 'submission.csv'
submission.to_csv(filename,index=True)
print('Saved file: ' + filename)
# -
| Used_car_prediction/car_predict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Author : <NAME>
# github link : https://github.com/amirshnll/Online-Shoppers-Purchasing-Intention/
# dataset link : http://archive.ics.uci.edu/ml/datasets/Online+Shoppers+Purchasing+Intention+Dataset
# email : <EMAIL>
# +
import csv
def transformDataMTT(trainingFile, features):
transformData=[]
labels = []
blank=""
# Now we are finally ready to read the csv file
with open(trainingFile,'r') as csvfile:
lineReader = csv.reader(csvfile,delimiter=',')
lineNum=1
# lineNum will help us keep track of which row we are in
for row in lineReader:
if lineNum==1:
header = row
else:
allFeatures=list(row)
featureVector = [allFeatures[header.index(feature)] for feature in features]
if blank not in featureVector:
transformData.append(featureVector)
labels.append(int(row[1]))
lineNum=lineNum+1
return transformData,labels
# return both our list of feature vectors and the list of labels
# -
# Let's take this for a spin now
trainingFile="O_S_I_train.csv"
features=["Administrative","Informational","ProductRelated","ProductRelated_Duration","BounceRates","ExitRates","PageValues","SpecialDay","Month","OperatingSystems","Browser","Region","TrafficType","VisitorType","Weekend","Revenue"]
trainingData=transformDataMTT(trainingFile,features)
# We are now ready to train our Decision Tree classifier
from sklearn import tree
import numpy as np
clf=tree.DecisionTreeClassifier(max_leaf_nodes=20)
X=np.array(trainingData[0])
y=np.array(trainingData[1])
clf=clf.fit(X,y)
import graphviz
with open("MTTTEST.dot","w") as f:
f = tree.export_graphviz(clf,
feature_names=features,out_file=f)
clf.feature_importances_
def transformTestDataMTT(testFile,features):
transformData=[]
ids=[]
blank=""
with open(testFile,"r") as csvfile:
lineReader = csv.reader(csvfile,delimiter=',')
lineNum=1
for row in lineReader:
if lineNum==1:
header=row
else:
allFeatures=list(row)
featureVector = [allFeatures[header.index(feature)] for feature in features]
#featureVector=list(map(lambda x:0 if x=="" else x, featureVector))
transformData.append(featureVector)
ids.append(row[0])
lineNum=lineNum+1
return transformData,ids
def MTTTest(classifier,resultFile,transformDataFunction=transformTestDataMTT):
testFile="O_S_I_test.csv"
testData=transformDataFunction(testFile,features)
result=classifier.predict(testData[0])
with open(resultFile,"w") as mf:
ids=testData[1]
lineWriter=csv.writer(mf,delimiter=',')
lineWriter.writerow(["ShopperId","Revenue"])
for rowNum in range(len(ids)):
try:
lineWriter.writerow([ids[rowNum],result[rowNum]])
except Exception as e:
print (e)
# Let's take this for a spin!
resultFile="result.csv"
MTTTest(clf,resultFile)
| .ipynb_checkpoints/Untitled4-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=["remove_cell"]
# # Transpiling Quantum Circuits
# -
# In this chapter we will investigate how quantum circuits are transformed when run on quantum devices. That we need to modify the circuits at all is a consequence of the limitations of current quantum computing hardware. Namely, the limited connectivity inherent in most quantum hardware, restricted gate sets, as well as environmental noise and gate errors, all conspire to limit the effective computational power on today's quantum devices. Fortunately, quantum circuit rewriting tool chains have been developed that directly address these issues, and return heavily optimized circuits mapped to targeted quantum devices. Here we will explore the IBM Qiskit 'transpiler' circuit rewriting framework.
import numpy as np
from qiskit import *
from qiskit.tools.jupyter import *
from qiskit.providers.ibmq import least_busy
# %matplotlib inline
# %config InlineBackend.figure_format = 'svg' # Makes the images look nice
# + tags=["uses-hardware"]
IBMQ.load_account()
# -
# ## Core Steps in Circuit Rewriting
# As we will see, rewriting quantum circuits to match hardware constraints and optimize for performance can be far from trivial. The flow of logic in the rewriting tool chain need not be linear, and can often have iterative sub-loops, conditional branches, and other complex behaviors. That being said, the basic building blocks follow the structure given below.
# 
#
# Our goal in this section is to see what each of these "passes" does at a high-level, and then begin exploring their usage on a set of common circuits.
# ### Unrolling to Basis Gates
# When writing a quantum circuit you are free to use any quantum gate (unitary operator) that you like, along with a collection of non-gate operations such as qubit measurements and reset operations. However, when running a circuit on a real quantum device one no longer has this flexibility. Due to limitations in, for example, the physical interactions between qubits, difficulty in implementing multi-qubit gates, control electronics etc, a quantum computing device can only natively support a handful of quantum gates and non-gate operations. In the present case of IBM Q devices, the native gate set can be found by querying the devices themselves:
# + tags=["uses-hardware"]
provider = IBMQ.get_provider(group='open')
provider.backends(simulator=False)
# + tags=["uses-hardware"]
backend = least_busy(provider.backends(filters=lambda x: x.configuration().n_qubits >= 5 and not x.configuration().simulator and x.status().operational==True))
backend.configuration().basis_gates
# -
# We see that the our device supports five native gates: three single-qubit gates (`u1`, `u2`, `u3`, and `id`) and one two-qubit entangling gate `cx`. In addition, the device supports qubit measurements (otherwise we can not read out an answer!). Although we have queried only a single device, all IBM Q devices support this gate set.
#
# The `u*` gates represent arbitrary single-qubit rotations of one, two, and three angles. The `u1` gates are single-parameter rotations that represent generalized phase gates of the form
#
# $$
# U_{1}(\lambda) = \begin{bmatrix}
# 1 & 0 \\
# 0 & e^{i\lambda}
# \end{bmatrix}
# $$
#
# This set includes common gates such as $Z$, $T$, $T^{\dagger}$, $S$, and $S^{\dagger}$. It turns out that these gates do not actually need to be performed on hardware, but instead, can be implemented in software as "virtual gates". These virtual gates are called "frame changes" and take zero time, and have no associated error; they are free gates on hardware.
#
# Two-angle rotations, $U_{2}(\phi,\lambda)$, are actually two frame changes with a single $X_{\pi/2}$ gate in between them, and can be used to synthesize gates like the Hadamard ($U_{2}(0,\pi)$) gate. As the only actual gate performed is the $X_{\pi/2}$ gate, the error and gate time associated with any $U_{2}$ gate is the same as an $X_{\pi/2}$ gate. Similarly, $U_{3}(\theta,\phi,\lambda)$ gates are formed from three frame changes with two $X_{\pi/2}$ gates in between them. The errors and gate times are twice those of a single $X_{\pi/2}$. The identity gate, $id$, is straightforward, and is a placeholder gate with a fixed time-interval.
#
# The only entangling gate supported by the IBM Q devices is the CNOT gate (`cx`) that, in the computational basis, can be written as:
#
# $$
# \mathrm{CNOT}(0,1) = \begin{bmatrix}
# 1 & 0 & 0 & 0 \\
# 0 & 0 & 0 & 1 \\
# 0 & 0 & 1 & 0 \\
# 0 & 1 & 0 & 0
# \end{bmatrix}
# $$,
#
# where we see that the matrix form follows from the specific bit-ordering convention used in Qiskit.
# Every quantum circuit run on a IBM Q device must be expressed using only these basis gates. For example, suppose one wants to run a simple phase estimation circuit:
# + tags=["uses-hardware"]
qr = QuantumRegister(2, 'q')
cr = ClassicalRegister(1, 'c')
qc = QuantumCircuit(qr, cr)
qc.h(qr[0])
qc.x(qr[1])
qc.cu1(np.pi/4, qr[0], qr[1])
qc.h(qr[0])
qc.measure(qr[0], cr[0])
qc.draw(output='mpl')
# -
# We have $H$, $X$, and controlled-$U_{1}$ gates, all of which are not in our devices basis gate set, and must be expanded. We will see that this expansion is taken care of for you, but for now let us just rewrite the circuit in the basis gate set:
# + tags=["uses-hardware"]
qr = QuantumRegister(2, 'q')
cr = ClassicalRegister(1, 'c')
qc_basis = QuantumCircuit(qr, cr)
# Hadamard in U2 format
qc_basis.u2(0, np.pi, qr[0])
# X gate in U3 format
qc_basis.u3(np.pi, 0, np.pi, qr[1])
# Decomposition for controlled-U1 with lambda=pi/4
qc_basis.u1(np.pi/8, qr[0])
qc_basis.cx(qr[0], qr[1])
qc_basis.u1(-np.pi/8, qr[1])
qc_basis.cx(qr[0], qr[1])
qc_basis.u1(np.pi/8, qr[1])
# Hadamard in U2 format
qc_basis.u2(0, np.pi, qr[0])
qc_basis.measure(qr[0], cr[0])
qc_basis.draw(output='mpl')
# -
# A few things to highlight. One, the circuit has gotten longer with respect to the initial one. This can be verified by checking the depth of the circuits:
# + tags=["uses-hardware"]
print(qc.depth(), ',', qc_basis.depth())
# -
# Second, although we had a single controlled gate, the fact that it was not in the basis set means that, when expanded, it requires more than a single `cx` gate to implement. All said, unrolling to the basis set of gates leads to an increase in the length of a quantum circuit and the number of gates. Both of these increases lead to more errors from the environment and gate errors, respectively, and further circuit rewriting steps must try to mitigate this effect through circuit optimizations.
# Finally, we will look at the particularly important example of a Toffoli, or controlled-controlled-not gate:
# + tags=["uses-hardware"]
qr = QuantumRegister(3, 'q')
qc = QuantumCircuit(qr)
qc.ccx(qr[0], qr[1], qr[2])
qc.draw(output='mpl')
# -
# As a three-qubit gate, it should already be clear that this is not in the basis set of our devices. We have already seen that controlled gates not in the basis set are typically decomposed into multiple CNOT gates. This is doubly true for controlled gates with more than two qubits, where multiple CNOT gates are needed to implement the entangling across the multiple qubits. In our basis set, the Toffoli gate can be written as:
# + tags=["uses-hardware"]
qr = QuantumRegister(3, 'q')
qc_basis = QuantumCircuit(qr)
qc_basis.u2(0,np.pi, qr[2])
qc_basis.cx(qr[1], qr[2])
qc_basis.u1(-np.pi/4, qr[2])
qc_basis.cx(qr[0], qr[2])
qc_basis.u1(np.pi/4, qr[2])
qc_basis.cx(qr[1], qr[2])
qc_basis.u1(np.pi/4, qr[1])
qc_basis.u1(-np.pi/4, qr[2])
qc_basis.cx(qr[0], qr[2])
qc_basis.cx(qr[0], qr[1])
qc_basis.u1(np.pi/4, qr[2])
qc_basis.u1(np.pi/4, qr[0])
qc_basis.u1(-np.pi/4, qr[1])
qc_basis.u2(0,np.pi, qr[2])
qc_basis.cx(qr[0], qr[1])
qc_basis.draw(output='mpl')
# -
# Therefore, for every Toffoli gate in a quantum circuit, the IBM Q hardware must execute six CNOT gates, and a handful of single-qubit gates. From this example, it should be clear that any algorithm that makes use of multiple Toffoli gates will end up as a circuit with large depth and with therefore be appreciably affected by noise and gate errors.
# ### Initial Layout
# + tags=["uses-hardware"]
qr = QuantumRegister(5, 'q')
cr = ClassicalRegister(5, 'c')
qc = QuantumCircuit(qr, cr)
qc.h(qr[0])
qc.cx(qr[0], qr[4])
qc.cx(qr[4], qr[3])
qc.cx(qr[3], qr[1])
qc.cx(qr[1], qr[2])
qc.draw(output='mpl')
# + tags=["uses-hardware"]
from qiskit.visualization.gate_map import plot_gate_map
plot_gate_map(backend, plot_directed=True)
# -
import qiskit
qiskit.__qiskit_version__
| content/ch-quantum-hardware/transpiling-quantum-circuits.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 업무자동화를 위한 파이썬 입문 CAMP #10-01-01
#
#
# > 주제 : 네이버에서 실시간 검색어 순위를 크롤링해서 출력하기
# ## Prerequisite
#
#
# ```shell
# $ pip install requests
# $ pip install beautifulsoup4
# ```
# +
import requests # http request 를 보내고 받기 위해서 사용됩니다.
from bs4 import BeautifulSoup # html 을 parsing 하기 위해서 사용됩니다.
# 즉, 저희는 크게 2가지 파이썬 패키지 ( 라이브러리 )를 사용하게 되는데요,
#
# 1. requests => html request 를 보내서, html response ( html, css, javascript )
# 를 받는다.
# 2. bs4 ( beautifulsoup4 ) => requests 를 통해서 받은 response 에서 우리가 필요한
# 데이터를 파싱한다.
#
# 알아두셔야할 점 => 우리가 흔히 크롤링 이라고 부르는 부분은 이렇게 항상 [1. Crawling] + [2. Parsing]이
# 합쳐진 거라고 생각하시면 좋을 것 같습니다.
# +
response = requests.get("http://www.naver.com/")
assert response.status_code is 200
# 위의 `assert response.status_code is 200`의 의미는 네이버 서버를 통해서
# response의 status_code 가 200인지 확인해보는 과정입니다. ( 테스트 )
#
# 각각의 status_code는 고유의 의미를 가지고 있는데, 많이 사용되는 것만 간단하게 소개를 드립니다:
# 200 => OK ( 성공 )
# 404 => NOT FOUND, 이 경우에는 제대로 크롤링을 하기가 힘들겠죠?
#
# 각각의 Status Code 들에 대해서는 https://www.w3.org/Protocols/HTTP/HTRESP.html 에서 추가적으로 살펴보실 수 있습니다.
# -
dom = BeautifulSoup(response.content, "html.parser")
# ```html
# ...
# <ol id="realrank">
# <li>패스트캠퍼스</li>
# <li>업무자동화</li>
# <li>업무자동화</li>
# ...
# </ol>
# ...
# ```
# +
# 일반적으로 bs4와 같은 parser에서 html element 를 선택하기 위한
# 다양한 방법이 있지만, 가장 선호되는 방식은 "css select" 방식입니다.
#
# id => "#"
# class => "."
# 예를 들어, 위의 html 코드에서 살펴보면,
# "realrank"라는 "id"를 가진 "ol" 태그 > 안에 들어있는 "li" 태그를 선택하려면,
# "ol#realrank li"로 선택을 할 수 있습니다.
ranking_elements = dom.select("ol#realrank li")
# -
for ranking_element in ranking_elements[:-1]:
ranking_title_element = ranking_element.select("a")[0]
print(ranking_title_element.attrs.get('title'))
# > 수고하셨습니다. 감사합니다. - 안수찬 올림.
| legacy/01-01-naver-search-ranking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sehejjain/cautious-guacamole/blob/master/Cuckoo_Search_Algorithm.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="qVhkrhPfDImy"
import random
import numpy as np
from numpy.random.mtrand import randint
from scipy.stats import multivariate_normal
import scipy.stats
import matplotlib.pyplot as plt
# + [markdown] id="w0B1eWEFDuk6"
# Class for Implementing Cuckoo Search Algorithm
# + id="ts-ate8xDMTe"
class cuckoo:
def __init__(self, populationSize, probability, iterations):
self.nestNumb = populationSize
self.Pa = probability
self.iterNumb = iterations
self.nests = []
def fitness(self, x): #x = (x,y)
mean = np.array([0,0])
cov = np.array([[0.1,0 ], [0,0.1]])
ans = multivariate_normal.pdf(x, mean, cov)
return pow((pow((x[0] - 3), 2) +pow((x[1] - 4), 2)), 2)
def generate(self):
x =random.uniform(-10,10)
y =random.uniform(-10,10)
return [x,y]
def LevyFlight(self, x):
beta = 1.5 # Lambda (between 1 and 3)
alpha = (random.uniform(-1,1)) #between -1 and 1
x += scipy.stats.levy_stable.pdf(x, alpha, beta)
return x
def abandonWorst(self, nest):
for i in range(len(nest)-int(self.Pa*len(nest)),len(nest)): # Pa of worst solutions
buf = self.generate() #generate new nests
val = self.fitness(buf) #evaluate new nests
nest[i] = [buf,val] #swap
return nest
def run(self):
# generate random nests
for i in range(self.nestNumb):
buf = self.generate()
self.nests.append((buf, self.fitness(buf)))
#start iteration
for step in range(self.iterNumb):
if (step%50 == 0):
print("iteration\t",step)
i = randint(0,self.nestNumb) #chose random nest
cuckoo = self.LevyFlight(self.nests[i][0]) #get random cuckoo and make him levy's flight
Fcuckoo = self.fitness(cuckoo) #evaluate cuckoo
jnest = randint(0,len(self.nests)) #nest chosen by cuckoo
if(Fcuckoo > self.nests[jnest][1]):
self.nests[jnest] = [cuckoo, Fcuckoo] #replace new solution
self.nests.sort(key=lambda val: val[1], reverse=True) #best solutions at start of list
self.nests = self.abandonWorst(self.nests)
self.nests.sort(key=lambda val: val[1], reverse=True)
return self.nests
# + id="-dKpbq_qDQWH"
def meanPoint(x,val):
mean=0
for i in range(len(x)):
mean= x[i]*val[i]
mean = mean/sum(val)
return mean
# + [markdown] id="oMXnfe74EBoS"
# #Initializing Parameters
# * n = Number of Nests
# * Pa = Probablility of Abandoning the nests
# * iterNumb = The number of Iterations
# + id="Lv3zIQVxDUeH"
n = 1000
Pa = 0.25
iterNumb = 1000
# + colab={"base_uri": "https://localhost:8080/"} id="ZYQxdVbMDZZk" outputId="a9e92026-fce8-418f-8a99-f4f7e4c921c2"
CS = cuckoo(n, Pa, iterNumb)
nests = CS.run()
# + [markdown] id="tzGqRrbnENnR"
# It can be seen that after all the iterations, the nests converge to the most optimal point, in the matlab plot below.
# + colab={"base_uri": "https://localhost:8080/", "height": 319} id="G09bKDdRDctz" outputId="40c6dbd3-fe07-43ea-c25a-1d1b59f6d852"
nestsPoints, nestsVal = zip(*nests)
print("vector of nests: ",nests)
x,y = zip(*nestsPoints)
print("Best point: ", nestsPoints[0], " which value is: ", nestsVal[0])
plt.plot(x,y, 'bo')
plt.show()
# + id="j4cuid3KDpFJ"
| Cuckoo_Search_Algorithm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## AI for Medicine Course 1 Week 1 lecture exercises
# <a name="densenet"></a>
# # Densenet
#
# In this week's assignment, you'll be using a pre-trained Densenet model for image classification.
#
# Densenet is a convolutional network where each layer is connected to all other layers that are deeper in the network
# - The first layer is connected to the 2nd, 3rd, 4th etc.
# - The second layer is connected to the 3rd, 4th, 5th etc.
#
# Like this:
#
# <img src="densenet.png" alt="U-net Image" width="400" align="middle"/>
#
# For a detailed explanation of Densenet, check out the source of the image above, a paper by <NAME> et al. 2018 called [Densely Connected Convolutional Networks](https://arxiv.org/pdf/1608.06993.pdf).
#
# The cells below are set up to provide an exploration of the Keras densenet implementation that you'll be using in the assignment. Run these cells to gain some insight into the network architecture.
# Import Densenet from Keras
from keras.applications.densenet import DenseNet121
from keras.layers import Dense, GlobalAveragePooling2D
from keras.models import Model
from keras import backend as K
# For your work in the assignment, you'll be loading a set of pre-trained weights to reduce training time.
# Create the base pre-trained model
base_model = DenseNet121(weights='./nih/densenet.hdf5', include_top=False);
# View a summary of the model
# Print the model summary
base_model.summary()
# +
# Print out the first five layers
layers_l = base_model.layers
print("First 5 layers")
layers_l[0:5]
# -
# Print out the last five layers
print("Last 5 layers")
layers_l[-6:-1]
# Get the convolutional layers and print the first 5
conv2D_layers = [layer for layer in base_model.layers
if str(type(layer)).find('Conv2D') > -1]
print("The first five conv2D layers")
conv2D_layers[0:5]
# Print out the total number of convolutional layers
print(f"There are {len(conv2D_layers)} convolutional layers")
# Print the number of channels in the input
print("The input has 3 channels")
base_model.input
# Print the number of output channels
print("The output has 1024 channels")
x = base_model.output
x
# Add a global spatial average pooling layer
x_pool = GlobalAveragePooling2D()(x)
x_pool
# Define a set of five class labels to use as an example
labels = ['Emphysema',
'Hernia',
'Mass',
'Pneumonia',
'Edema']
n_classes = len(labels)
print(f"In this example, you want your model to identify {n_classes} classes")
# Add a logistic layer the same size as the number of classes you're trying to predict
predictions = Dense(n_classes, activation="sigmoid")(x_pool)
print(f"Predictions have {n_classes} units, one for each class")
predictions
# Create an updated model
model = Model(inputs=base_model.input, outputs=predictions)
# Compile the model
model.compile(optimizer='adam',
loss='categorical_crossentropy')
# (You'll customize the loss function in the assignment!)
# #### This has been a brief exploration of the Densenet architecture you'll use in this week's graded assignment!
| AI for Medical Diagnosis/Week 1/C1W1_L3_Densenet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # INTRODUCTION
#
#
# > Ever heard of the terminologies like used car, a pre-owned vehicle or a second hand car? <br>
# >
# > Used cars are sold through a varirty of outlets, including franchise, independent car dealers and rental car companies.Have you ever wondered how their prices are evaluated for sale?
# >
# > You cannot take a wild guess at this because the Indian used car market was valued at USD 24.24 billion in 2019.
# > Let's build a model that helps the vendors evaluate the used cars.
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
#Importing the necessary libraries
import numpy as np
import pandas as pd
import seaborn as sb
import matplotlib.pyplot as plt
from sklearn.preprocessing import *
from sklearn.model_selection import *
from sklearn.ensemble import *
from sklearn.feature_selection import *
from sklearn.metrics import *
from sklearn.tree import *
from sklearn.svm import *
from sklearn.neighbors import *
from sklearn.linear_model import *
import xgboost as xgb
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
#To avoid unnecessary warning
import warnings
with warnings.catch_warnings():
warnings.filterwarnings("ignore")
# -
# # DATA GATHERING
# + _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0"
train = pd.read_csv('../input/used-cars-price-prediction/train-data.csv',index_col= 0)
train = train.reindex(np.random.permutation(train.index))
print("TRAIN SHAPE: ",train.shape)
train.info()
train.head()
# -
test = pd.read_csv('../input/used-cars-price-prediction/test-data.csv',index_col= 0)
print(test.shape)
print(test.info())
test.head()
# # DATA CLEANING AND ASSESSMENT
# Ensure to apply the changes to both the test and train sets inorder to maintain uniformity.
# ##### Quality issue 1: Missing values
# We can see that there are missing values in the following columns:
# * Engine
# * Power
# * Seats
# * New_Price
#percentage of missing values
percent_missing = train.isnull().sum() * 100 / len(train)
print(percent_missing)
percent_missing = test.isnull().sum() * 100 / len(test)
print(percent_missing)
# +
#dropping the "New_Price" column that has 86.3% of missing values
train.drop(columns =['New_Price'],axis =1, inplace = True)
test.drop(columns =['New_Price'],axis =1, inplace = True)
# +
#Mileage attribute has the least percentage of missing values. Let's fill them up manually.
train[train['Mileage'].isnull()]
# +
#Thanks to Google!
train.loc[4904, 'Mileage'] = '23.91 kmpl'
train.loc[4446, 'Mileage'] = '140 kmpl'
# +
#Now,let's drop the rest of the rows with missing values
train.dropna(how ='any',inplace = True)
test.dropna(how ='any',inplace = True)
# -
#CHECKING IF ALL THE MISSING VALUES ARE TAKEN CARE OF
train.info()
test.info()
# ##### Quality issue 2: Erroneous values and datatypes
# Some of our attributes' datatypes could be changed inorder to make the modelling process easier. Those attributes are
# * Mileage
# * Engine
# * Power
# * Year
# +
#Mileage - Before we change the datatype, we must extract the actual mileage in numbers without the "Kmpl"
train['Mileage']= train['Mileage'].str[:-5]
train['Mileage']=train['Mileage'].astype(float);
test['Mileage']= test['Mileage'].str[:-5]
test['Mileage']=test['Mileage'].astype(float);
# +
#Engine - Before we change the datatype, we must extract the actual engine cc in numbers without the "CC" string
train['Engine'] = train['Engine'].str.strip('CC')
train['Engine']= train['Engine'].astype(float);
test['Engine'] = test['Engine'].str.strip('CC')
test['Engine']= test['Engine'].astype(float);
# +
train['Power'] = train['Power'].fillna(value = "null")
train["Power"]= train["Power"].replace("null", "NaN")
train['Power'] = train['Power'].str.strip('bhp ')
train['Power'] = train['Power'].astype(float)
train.dropna(how ='any',inplace = True)
# +
test['Power'] = test['Power'].fillna(value = "null")
test["Power"]= test["Power"].replace("null", "NaN")
test['Power'] = test['Power'].str.strip('bhp ')
test['Power'] = test['Power'].astype(float)
test.dropna(how ='any',inplace = True)
# +
#Year
train['Year'] = train['Year'].astype(str)
test['Year'] = test['Year'].astype(str)
# -
#CHECKING
train.info()
test.info()
train.to_csv('trainfinal.csv')
test.to_csv('testfinal.csv')
# # Feature engineering
x = pd.read_csv('trainfinal.csv')
print(x.shape)
x.head()
#dropping the unnamed:0 column
x.drop(columns=['Unnamed: 0'],axis=1,inplace = True)
# **The name column has a diverse number of values.
# Let's break it down and extract the brand name of the car.**
#
x["breakdown"] = x.Name.str.split(" ")
x["breakdown"].head()
# +
#Lets store the brand name in our new column
brand_list=[]
for i in range(len(x)):
a = x.breakdown[i][0]
brand_list.append(a)
x['Brand'] = brand_list
# -
# We don't need these columns now
x.drop(columns=['Name','breakdown'],axis=1,inplace=True)
#Lets analyse the new attribute
x['Brand'].unique()
# We can see that "Isuzu" carries some duplicated values.
# <br>
# Let's sort that out
duplic = {'ISUZU': 'Isuzu'}
x.replace({"Brand": duplic},inplace = True)
# +
#CHECKING
x['Brand'].value_counts()
#Sorted!
# +
#Lets encode our categorical values
labelencoder = LabelEncoder()
label_array=[]
label_array = ['Location','Year','Fuel_Type','Transmission','Owner_Type','Brand']
for ele in label_array:
x[ele] = labelencoder.fit_transform(x[ele])
# -
#CHECKING
x.head()
# +
#feature selection
X_fs = x[['Location', 'Year', 'Kilometers_Driven', 'Fuel_Type', 'Transmission', 'Owner_Type',
'Mileage', 'Engine', 'Power', 'Seats', 'Brand']]
y_fs = x['Price']
y_fs = y_fs*100
y_fs = y_fs.astype(int)
bestfeatures = SelectKBest(score_func=chi2, k=5)
fit = bestfeatures.fit(X_fs,y_fs)
dfscores = pd.DataFrame(fit.scores_)
dfcolumns = pd.DataFrame(X_fs.columns)
featureScores = pd.concat([dfcolumns,dfscores],axis=1)
featureScores.columns = ['Specs','Score'] #naming the dataframe columns
fea = pd.DataFrame(featureScores.nlargest(10,'Score'))
print(featureScores.nlargest(10,'Score')) #print 10 best features
# -
# Therefore, the most important features are "Kilometers driven" followed by "Engine", "Power", "Brand".
# +
selection= ExtraTreesRegressor()
selection.fit(X_fs,y_fs)
plt.figure(figsize = (12,8))
feat_importances = pd.Series(selection.feature_importances_, index=X_fs.columns)
feat_importances.nlargest(20).plot(kind='barh')
plt.show()
# -
# This plot also validates our selection of important features.
# # MODEL BUILDING
#Preparing Training set
X = np.array(x.drop(['Price'],axis = 1))
Y = x.Price.values
#splitting into train and test set
X_train,X_test,Y_train,Y_test = train_test_split(X,Y,test_size=0.20,random_state=25)
# +
#selecting best models
model_selc = [LinearRegression(),
DecisionTreeRegressor(),
RandomForestRegressor(n_estimators=10),
KNeighborsRegressor(),
GradientBoostingRegressor()]
kfold = RepeatedKFold(n_splits=5, n_repeats=10, random_state= None)
cv_results = []
cv_results_mean =[]
for ele in model_selc:
cross_results = cross_val_score(ele, X_train, Y_train, cv=kfold, scoring ='r2')
cv_results.append(cross_results)
cv_results_mean.append(cross_results.mean())
print("\n MODEL: ",ele,"\nMEAN R2:",cross_results.mean() )
# -
#Let's try xgboost now
my_xgb = xgb.XGBRegressor(objective='reg:linear',learning_rate = 0.1, n_estimators = 100,verbosity = 0,silent=True)
xgb_results = cross_val_score(my_xgb, X_train, Y_train, cv=kfold, scoring ='r2')
print("\n MODEL: XGBOOST","\nMEAN R2:",xgb_results.mean() )
# AND THERE WE HAVE A WINNER TO GO THROUGH GRIDSEARCH FOR HYPER-PARAMTER TUNING!!
# ### Tuning HyperParameters for xgboost
# ##### GRID SEARCH
#
# Grid Search can be thought of as an exhaustive and computationally expensive method for selecting a model.<br>
#
# **For example** : <br>
#
# Searching 20 different parameter values for each of 4 parameters will require 160,000 trials of cross-validation. This equates to 1,600,000 model fits and 1,600,000 predictions if 10-fold cross validation is used
# +
#We use GridSearch for fine tuning Hyper Parameters
from sklearn.model_selection import *
n_estimator_val = np.arange(100,400,100).astype(int)
max_depth_val = [2,3,4]
grid_params = { 'loss' : ['ls'] ,
'learning_rate' : [0.1],
'n_jobs': [-1],
'n_estimators' : n_estimator_val,
'max_depth' : max_depth_val
}
# -
gs = GridSearchCV(xgb.XGBRegressor(silent= True),grid_params,verbose=1,cv=5,n_jobs =-1)
gs_results = gs.fit(X_train,Y_train)
#To Display the Best Score
gs_results.best_score_
#To Display the Best Estimator
gs_results.best_estimator_
#To Display the Best Parameters
gs_results.best_params_
# We could settle here but let's try randomized search cross validation.
# ##### RANDOMIZED SEARCH
# Randomized Search sets up a grid of hyperparameter values and selects random combinations to train the model and score.
# This allows you to explicitly control the number of parameter combinations that are attempted.
# The number of search iterations is set based on time or resources.
#
#
# While it’s possible that RandomizedSearchCV will not find as accurate of a result as GridSearchCV, it surprisingly picks the best result more often than not and in a fraction of the time it takes GridSearchCV would have taken. Given the same resources, Randomized Search can even outperform Grid Search.
XGBRegressor(base_score=0.5, booster='gbtree', colsample_bylevel=1,
colsample_bynode=1, colsample_bytree=0.8, gamma=0.5,
importance_type='gain', learning_rate=0.1, max_delta_step=0,
max_depth=4, min_child_weight=10, missing=None, n_estimators=1900,
n_jobs=-1, nthread=None, objective='reg:linear', random_state=0,
reg_alpha=0, reg_lambda=1, scale_pos_weight=1, seed=None,
silent=None, subsample=1.0, verbosity=1)
# +
folds = 3
param_comb = 10
skf = StratifiedKFold(n_splits=folds, shuffle = True, random_state = 1001)
params = { 'n_jobs': [-1],
'n_estimators' : n_estimator_val,
'learning_rate' : [0.1],
'min_child_weight': [9],
'gamma': [0.5],
'subsample': [0.6],
'colsample_bytree': [0.8, 1.0],
'max_depth': [3, 4]
}
xgb_regrsv = xgb.XGBRegressor()
# -
random_search = RandomizedSearchCV(xgb_regrsv, params, n_iter=param_comb, scoring='r2',
n_jobs=-1, cv=5 )
random_search.fit(X_train, Y_train);
random_search.best_score_
random_search.best_estimator_
# After some more time of trying various hyperparameters and tuning,
xgb_tuned = xgb.XGBRegressor(base_score=0.5, booster='gbtree', colsample_bylevel=1,
colsample_bynode=1, colsample_bytree=0.8, gamma=0.5,
importance_type='gain', learning_rate=0.1, max_delta_step=0,
max_depth=4, min_child_weight=9, missing=None, n_estimators=300,
n_jobs=-1, nthread=None, objective='reg:linear', random_state=0,
reg_alpha=0, reg_lambda=1, scale_pos_weight=1, seed=None,
silent=True, subsample=1.0, verbosity=0)
# +
xgb_tuned.fit(X_train,Y_train)
y_pred =xgb_tuned.predict(X_test)
print("Training set accuracy: ",xgb_tuned.score(X_train,Y_train))
print("Test set accuracy : ",xgb_tuned.score(X_test,Y_test))
# -
# Now, lets check how various metrics have evaluated our model on the test set
# +
print("\t\tError Table")
print('Mean Absolute Error : ', mean_absolute_error(Y_test, y_pred))
print('Mean Squared Error : ', mean_squared_error(Y_test, y_pred))
print('Root Mean Squared Error : ', np.sqrt(mean_squared_error(Y_test, y_pred)))
print('R Squared Error : ', r2_score(Y_test, y_pred))
# -
# # CONCLUSION
#
# This model is ready to be deployed in a pipeline. <br>
# We have accomplished the task of building a good model for our used cars' prediction purposes.
#
# We will be adding the exploratory data analysis of the same dataset soon. <br>
#
#
#
# ##### Please do let us know if you have any constructive feedbacks to help us improve our work.
# ##### If you guys could learn something from our notebook, do upvote and support us! **
# This brings us to the end of our first kaggle notebook.
#
| used-car-prices-prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %autosave 20
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import math
import cmath
cmath.sqrt(-1)
fh = open('diagram.csv')
text = fh.read()
fh.close()
print(text)
with open('diagram.csv') as fh:
text = fh.read()
print(fh.closed)
# +
x = []
y = []
with open('diagram.csv', encoding='utf-8') as fh:
for row in fh:
row = row.strip()
a, b = row.split(',')
x.append(float(a))
y.append(float(b))
x = np.array(x)
y = np.array(y)
print(x)
print(y)
plt.plot(x, y, 'x')
with open('diagram.tsv', 'w') as fh:
for a, b in zip(x * 7, y):
fh.write('{}\t{}\n'.format(a, b))
# -
with open('cat.jpg', 'rb') as fh:
data = fh.read(16)
print(data)
print(type(data))
# +
from PIL import Image
img = Image.open('cat.jpg')
cat = np.array(img)
cat.dtype
# -
table = np.genfromtxt('diagram.csv',
delimiter=',',
names=('d', 'v'))
print(table[0])
print(table['d'])
records = np.rec.array(table)
records.d
table = np.genfromtxt('freddi.dat', names=True,
skip_footer=100)
# +
import gzip
import os
import shutil
with gzip.open('V404Cyg.txt.gz') as fh:
for _ in range(10):
print(fh.readline())
# -
table = np.genfromtxt('V404Cyg.txt.gz', names=True,
usecols=(0, 1, 2,),
missing_values=b'', filling_values=0,)
table[:10]
# +
def magn_converter(s):
if s.startswith(b'<'):
x = float(s[1:]) + 900
return x
return float(s)
table = np.genfromtxt('V404Cyg.txt.gz', names=True,
usecols=(0, 1, 2,),
dtype=(float, float, float),
missing_values=b'', filling_values=0,
converters={
1: magn_converter
})
table[:10]
is_upper_limit = table['Magnitude'] > 500
good_data = table[np.logical_not(is_upper_limit)]
plt.plot(good_data['JD'], good_data['Magnitude'], 'x')
# -
df = pd.DataFrame(table)
df.JD
df['JD']
df.columns
# df[0]
df.loc[2]
df = pd.read_table('V404Cyg.txt.gz', low_memory=False)
print(df.dtypes)
print(type(df.Magnitude[0]))
# +
# import json
# .yaml, .yml, .ini
# +
def magn_converter(s):
if s.startswith('<'):
s = s[1:]
return float(s)
df['m'] = df.Magnitude.map(magn_converter)
df['is_upper_limit'] = df.Magnitude.map(
lambda s: s.startswith('<')
)
df['m']
# -
upper_limits = df[df.is_upper_limit]
| misc/jupyter_notebooks/18.10.17/read_files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
def get_data(filename):
data = pd.read_csv(filename)
start = data['From'].tolist()
end = data['To'].tolist()
duration = data['Time'].tolist()
network = dict()
for i in range(0,len(start)):
task = f'{start[i]} -> {end[i]}'
network[task]= dict()
network[task]['name'] = task
network[task]['duration'] = duration[i]
network[task]['dependencies'] = get_dependencies(start[i], start, end)
network[task]['neighbors'] = get_neighbors(end[i], start, end)
network[task]['ES'] = 0
network[task]['EF'] = 0
network[task]['LS'] = 0
network[task]['LF'] = 0
return start, end, duration, network
def print_network(start, end, duration):
print("Network:")
for i in range(0,len(start)):
print(f"Task: {start[i]} -> {end[i]}\tDuration: {duration[i]}")
def check_feasability(start, end):
if len(set(start)-set(end)):
return True
if len(set(end)-set(start)):
return True
return False
def get_dependencies(task_name, start, end):
dependency_index = [j for j, val in enumerate(end) if val == task_name]
if len(dependency_index):
dependencies = [f"{start[j]} -> {end[j]}" for j in dependency_index]
return dependencies
return []
def get_neighbors(task_name, start, end):
neighbor_index = [j for j, val in enumerate(start) if val == task_name]
if len(neighbor_index):
neighbors = [f"{start[j]} -> {end[j]}" for j in neighbor_index]
return neighbors
return []
def get_attribute(network, dependencies, attribute, cond="min"):
attributes = []
for dependency in dependencies:
attributes.append(network[dependency][attribute])
if cond == "min":
return min(attributes), dependencies[attributes.index(min(attributes))]
elif cond == "max":
return max(attributes), dependencies[attributes.index(max(attributes))]
else:
print("Wrong condition passed")
return -1
def forward_pass(network):
for task in network:
dependencies = network[task]['dependencies']
if len(dependencies) > 0:
# if not first task
# calculate ES = maximum EF value from immediate predecessors
network[task]['ES'], pos = get_attribute(network,dependencies,"EF","max")
network[task]['EF'] = network[task]['ES'] + network[task]['duration']
else:
#if first task
network[task]['ES'] = 0
network[task]['EF'] = (network[task]['duration'])
return network
def backward_pass(network):
back_network = [i for i in network.keys()]
back_network.reverse()
max_EF = 0
last_task = back_network[-1]
# Find the end node
for task in back_network:
if network[task]['EF'] > max_EF:
max_EF = network[task]['EF']
last_task = task
for task in back_network:
if len(network[task]["neighbors"])>0:
# if not last task
# calculate LF = minimum LS value from immediate successors
network[task]['LF'], pos = get_attribute(network,network[task]["neighbors"],"LS","min")
network[task]['LS'] = network[task]['LF'] - network[task]['duration']
else:
# if last task
network[task]['LF']=network[last_task]['EF']
network[task]['LS']=network[task]['LF'] - network[task]['duration']
return network
def print_network_table(network):
print('task name\tduration\tES\t\tEF\t\tLS\t\tLF')
for task in network:
print(f"{network[task]['name']}\t\t{network[task]['duration']}\t\t{network[task]['ES']}\t\t{network[task]['EF']}\t\t{network[task]['LS']}\t\t{network[task]['LF']}")
def critical_nodes(network):
nodes = []
for task in network:
if(network[task]['ES'] == network[task]['LS'] and network[task]['EF'] == network[task]['LF']):
nodes.append(task)
print(f"\nCritical Nodes: {set(nodes)}")
filename = 'critical_path_problem.csv'
start, end, duration, network = get_data(filename)
if check_feasability(start, end):
network = forward_pass(network)
network = backward_pass(network)
print_network_table(network)
critical_nodes(network)
else:
print("The provided network is incorrect!")
| 6/critical path.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="vgXbvCBnYDzt" colab_type="text"
# ## Outliers
#
# An outlier is an observation that lies outside the overall pattern of a distribution __[Moore and McCabe, 1999]__.
#
# - Outliers can either be treated special completely ignored
#
# - E.g., Fraudulant transactions are outliers, but since we want to avoid them, they must be paid special attention
#
# - If we think that the outliers are errors, we should remove them
#
#
# ## Which of the ML models care about Outliers?
#
# Affected models:
#
# - AdaBoost
# - Linear models
# - Linear regression
# - Neural Networks (if the number is high)
# - Logistic regression
# - KMeans
# - Heirarchical Clustering
# - PCA
#
# Unaffected models:
#
# - Decision trees
# - Naive bayes
# - SVMs
# - Random forest
# - Gradient boosted trees
# - K-Nearest Neighbors
#
#
# ### Identification
#
# - Extreme Value Analysis
# - IQR = 75th quantile - 25th quantile
# - Upper boundary = 75th quantile + (IQR * 1.5)
# - Lower boundary = 25th quantile - (IQR * 1.5)
# - Upper Extreme boundary = 75th quantile + (IQR * 3)
# - Lower Extreme boundary = 25th quantile - (IQR * 3)
#
# + id="42hbGwCeDd8-" colab_type="code" outputId="2c0bb18e-10e4-472c-d3e4-6bca63e2e0fc" colab={"base_uri": "https://localhost:8080/", "height": 72}
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
# + id="Ds9gl_oFEATI" colab_type="code" outputId="d86d96c7-aea8-44cf-fa45-91d3ef0cce5e" colab={"base_uri": "https://localhost:8080/", "height": 125}
from google.colab import drive
drive.mount('/content/gdrive')
data = pd.read_csv("gdrive/My Drive/Colab Notebooks/FeatureEngineering/train_date.csv")
# + id="FIDvE3p-s872" colab_type="code" outputId="c0914ce4-abdc-424c-f9fd-1b1c6018feaa" colab={"base_uri": "https://localhost:8080/", "height": 459}
data.head()
# + id="liUrG1RoYD0J" colab_type="code" outputId="ab6d0a19-a65e-439b-b776-3a58c11cda7e" colab={"base_uri": "https://localhost:8080/", "height": 369}
# Outliers according to the quantiles + 1.5 IQR
sns.catplot(x="Survived", y="Fare", kind="box", data=data)
sns.despine(left=False, right=False, top=False)
# + id="tuPMVeeMYD0P" colab_type="code" outputId="635894bf-6420-4b29-c876-41c02bbe51fa" colab={"base_uri": "https://localhost:8080/", "height": 175}
data['Fare'].describe()
# + id="Xsmlqi-mYD0l" colab_type="code" outputId="57b41350-ca2e-4f43-f851-9e32dd0e4bee" colab={"base_uri": "https://localhost:8080/", "height": 87}
# Get outliers
IQR = data['Fare'].quantile(0.75) - data['Fare'].quantile(0.25)
ub = data['Fare'].quantile(0.75) + (IQR * 3)
lb = data['Fare'].quantile(0.25) - (IQR * 3)
data[(data['Fare']>ub) | (data['Fare']<lb)].groupby('Survived')['Fare'].count()
# + id="hb9soWOMYD0x" colab_type="code" outputId="308daf8f-1b50-46ad-f54e-40a8d8697ae3" colab={"base_uri": "https://localhost:8080/", "height": 312}
# First let's plot the histogram to get an idea of the distribution
fig = data.Age.hist(bins=50, color='green')
fig.set_title('Distribution')
fig.set_xlabel('X')
fig.set_ylabel('#')
# + id="tKCDmb61voRC" colab_type="code" outputId="f813823a-85ba-48c1-9d94-c02cea81651b" colab={"base_uri": "https://localhost:8080/", "height": 87}
# Get outliers
IQR = data['Age'].quantile(0.75) - data['Age'].quantile(0.25)
ub = data['Age'].mean() + data['Age'].std()
lb = data['Age'].mean() - data['Age'].std()
data[(data['Age']>ub) | (data['Age']<lb)].groupby('Survived')['Age'].count()
# + id="Sd1QAUsvYD1J" colab_type="code" colab={}
from sklearn.metrics import roc_auc_score
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
# + id="7d9Ze6uwYD1M" colab_type="code" colab={}
data = data.drop(['Date'], axis=1)
# + id="6613mx4Z6B6C" colab_type="code" outputId="574fdade-77bd-4f18-b511-cb9309726fb9" colab={"base_uri": "https://localhost:8080/", "height": 70}
data.columns
# + id="vVDsyp3yYD1O" colab_type="code" outputId="d2be6910-24fc-46b7-f30a-c48d692463c1" colab={"base_uri": "https://localhost:8080/", "height": 70}
data[['Fare','Age']].isnull().mean()
# + id="TuUwgyv-YD1R" colab_type="code" outputId="c2693be0-d077-4806-d8b0-c15a99707ab0" colab={"base_uri": "https://localhost:8080/", "height": 34}
X_train, X_test, y_train, y_test = train_test_split(data[['Age', 'Fare']].fillna(0),
data['Survived'],
test_size=0.2)
X_train.shape, X_test.shape
# + id="VN_SePVTYD1T" colab_type="code" colab={}
# We will cap the values of outliers
data_processed = data.copy()
_temp = np.ceil(data['Age'].mean() + data['Age'].std())
data_processed.loc[data_processed.Age >= _temp, 'Age'] = _temp
IQR = data['Fare'].quantile(0.75) - data['Fare'].quantile(0.25)
_temp = np.ceil(data['Fare'].quantile(0.75) + (IQR * 3))
data_processed.loc[data_processed.Fare > _temp, 'Fare'] = _temp
X_train_processed, X_test_processed, y_train_processed, y_test_processed = train_test_split(
data_processed[['Age', 'Fare']].fillna(0),
data_processed['Survived'],
test_size=0.2)
# + id="kDCwav-bw1Ni" colab_type="code" outputId="68644e2d-9471-4e62-cdc9-807aa88d7750" colab={"base_uri": "https://localhost:8080/", "height": 122}
from sklearn.covariance import EllipticEnvelope
df_outliers = data.copy()
df_outliers = df_outliers.fillna(0)
column_name = 'Fare'
obj = EllipticEnvelope()
_temp = obj.fit_predict(df_outliers[[column_name]])
print(np.unique(_temp, return_counts=True))
central = df_outliers[_temp==1][column_name].mean()
max_val = df_outliers[_temp==1][column_name].max()
min_val = df_outliers[_temp==1][column_name].min()
df_outliers.loc[_temp==-1,[column_name]] = df_outliers.loc[_temp==-1,[column_name]].apply(lambda x: [max_val if y > central else y for y in x])
df_outliers.loc[_temp==-1,[column_name]] = df_outliers.loc[_temp==-1,[column_name]].apply(lambda x: [min_val if y < central else y for y in x])
print(data.shape)
print(df_outliers.shape)
column_name = 'Age'
obj = EllipticEnvelope()
_temp = obj.fit_predict(df_outliers[[column_name]])
print(np.unique(_temp, return_counts=True))
central = df_outliers[_temp==1][column_name].mean()
max_val = df_outliers[_temp==1][column_name].max()
min_val = df_outliers[_temp==1][column_name].min()
df_outliers.loc[_temp==-1,[column_name]] = df_outliers.loc[_temp==-1,[column_name]].apply(lambda x: [max_val if y > central else y for y in x])
df_outliers.loc[_temp==-1,[column_name]] = df_outliers.loc[_temp==-1,[column_name]].apply(lambda x: [min_val if y < central else y for y in x])
print(data.shape)
print(df_outliers.shape)
X_train_outliers, X_test_outliers, y_train_outliers, y_test_outliers = train_test_split(
df_outliers[['Age', 'Fare']],
df_outliers['Survived'],
test_size=0.2)
# + id="SUjEsyuX8FDq" colab_type="code" outputId="476832b2-f680-41b5-ac06-9e2625cdcca6" colab={"base_uri": "https://localhost:8080/", "height": 122}
print(X_train.shape)
print(X_train_processed.shape)
print(X_train_outliers.shape)
print(X_test.shape)
print(X_test_processed.shape)
print(X_test_outliers.shape)
# + id="tsRG3K5l8t_J" colab_type="code" outputId="42c98f3c-55e7-4531-815e-9f4e7b5ac504" colab={"base_uri": "https://localhost:8080/", "height": 70}
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
print(accuracy_score(y_test, y_pred))
classifier.fit(X_train_processed,y_train_processed)
y_pred_processed = classifier.predict(X_test_processed)
y_pred_processed = np.round(y_pred_processed).flatten()
print(accuracy_score(y_test_processed, y_pred_processed))
classifier.fit(X_train_outliers,y_train_outliers)
y_pred_outliers = classifier.predict(X_test_outliers)
y_pred_outliers = np.round(y_pred_outliers).flatten()
print(accuracy_score(y_test_outliers, y_pred_outliers))
# + id="2HgLWOz-9Uda" colab_type="code" outputId="35aff728-1c14-480e-f814-d539858acc92" colab={"base_uri": "https://localhost:8080/", "height": 70}
from sklearn.linear_model import RidgeClassifierCV
classifier = RidgeClassifierCV()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
print(accuracy_score(y_test, y_pred))
classifier.fit(X_train_processed,y_train_processed)
y_pred_processed = classifier.predict(X_test_processed)
y_pred_processed = np.round(y_pred_processed).flatten()
print(accuracy_score(y_test_processed, y_pred_processed))
classifier.fit(X_train_outliers,y_train_outliers)
y_pred_outliers = classifier.predict(X_test_outliers)
y_pred_outliers = np.round(y_pred_outliers).flatten()
print(accuracy_score(y_test_outliers, y_pred_outliers))
# + id="gS9bIfAB9eKW" colab_type="code" outputId="bb9c0ed0-ca1b-42a8-9105-29a452f3a0b7" colab={"base_uri": "https://localhost:8080/", "height": 70}
from sklearn.linear_model import RidgeClassifierCV
classifier = RidgeClassifierCV()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
print(accuracy_score(y_test, y_pred))
classifier.fit(X_train_processed,y_train_processed)
y_pred_processed = classifier.predict(X_test_processed)
y_pred_processed = np.round(y_pred_processed).flatten()
print(accuracy_score(y_test_processed, y_pred_processed))
classifier.fit(X_train_outliers,y_train_outliers)
y_pred_outliers = classifier.predict(X_test_outliers)
y_pred_outliers = np.round(y_pred_outliers).flatten()
print(accuracy_score(y_test_outliers, y_pred_outliers))
# + id="bZLwAZRI9vl3" colab_type="code" outputId="a11e8048-241a-42c6-9ad7-ce3027ccf5b6" colab={"base_uri": "https://localhost:8080/", "height": 70}
from sklearn.svm import SVC
classifier = SVC()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
print(accuracy_score(y_test, y_pred))
classifier.fit(X_train_processed,y_train_processed)
y_pred_processed = classifier.predict(X_test_processed)
y_pred_processed = np.round(y_pred_processed).flatten()
print(accuracy_score(y_test_processed, y_pred_processed))
classifier.fit(X_train_outliers,y_train_outliers)
y_pred_outliers = classifier.predict(X_test_outliers)
y_pred_outliers = np.round(y_pred_outliers).flatten()
print(accuracy_score(y_test_outliers, y_pred_outliers))
# + id="RO5oc4UJ9ykm" colab_type="code" outputId="6b043181-970b-4867-fa7d-7f9da177dc43" colab={"base_uri": "https://localhost:8080/", "height": 70}
from sklearn.neural_network import MLPClassifier
classifier = MLPClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
print(accuracy_score(y_test, y_pred))
classifier.fit(X_train_processed,y_train_processed)
y_pred_processed = classifier.predict(X_test_processed)
y_pred_processed = np.round(y_pred_processed).flatten()
print(accuracy_score(y_test_processed, y_pred_processed))
classifier.fit(X_train_outliers,y_train_outliers)
y_pred_outliers = classifier.predict(X_test_outliers)
y_pred_outliers = np.round(y_pred_outliers).flatten()
print(accuracy_score(y_test_outliers, y_pred_outliers))
# + id="RFKVucj790aA" colab_type="code" outputId="70da3fb2-c3d8-44ff-b841-f1dc4dec277c" colab={"base_uri": "https://localhost:8080/", "height": 195}
from sklearn.svm import LinearSVC
classifier = LinearSVC()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
print(accuracy_score(y_test, y_pred))
classifier.fit(X_train_processed,y_train_processed)
y_pred_processed = classifier.predict(X_test_processed)
y_pred_processed = np.round(y_pred_processed).flatten()
print(accuracy_score(y_test_processed, y_pred_processed))
classifier.fit(X_train_outliers,y_train_outliers)
y_pred_outliers = classifier.predict(X_test_outliers)
y_pred_outliers = np.round(y_pred_outliers).flatten()
print(accuracy_score(y_test_outliers, y_pred_outliers))
# + id="wgb5-ZI19274" colab_type="code" outputId="518d2659-9abf-4355-8aaa-5381d27ce336" colab={"base_uri": "https://localhost:8080/", "height": 70}
from sklearn.ensemble import RandomForestClassifier
classifier = RandomForestClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
print(accuracy_score(y_test, y_pred))
classifier.fit(X_train_processed,y_train_processed)
y_pred_processed = classifier.predict(X_test_processed)
y_pred_processed = np.round(y_pred_processed).flatten()
print(accuracy_score(y_test_processed, y_pred_processed))
classifier.fit(X_train_outliers,y_train_outliers)
y_pred_outliers = classifier.predict(X_test_outliers)
y_pred_outliers = np.round(y_pred_outliers).flatten()
print(accuracy_score(y_test_outliers, y_pred_outliers))
# + id="hrF5V_7_94hA" colab_type="code" outputId="a849520d-f53e-4173-c278-e560c7a7cf62" colab={"base_uri": "https://localhost:8080/", "height": 70}
from sklearn.tree import DecisionTreeClassifier
classifier = DecisionTreeClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
print(accuracy_score(y_test, y_pred))
classifier.fit(X_train_processed,y_train_processed)
y_pred_processed = classifier.predict(X_test_processed)
y_pred_processed = np.round(y_pred_processed).flatten()
print(accuracy_score(y_test_processed, y_pred_processed))
classifier.fit(X_train_outliers,y_train_outliers)
y_pred_outliers = classifier.predict(X_test_outliers)
y_pred_outliers = np.round(y_pred_outliers).flatten()
print(accuracy_score(y_test_outliers, y_pred_outliers))
# + id="0M4ySoHj97e5" colab_type="code" outputId="9a227b51-1244-423f-a224-8ecb4dcebcb1" colab={"base_uri": "https://localhost:8080/", "height": 70}
from sklearn.ensemble import GradientBoostingClassifier
classifier = GradientBoostingClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
print(accuracy_score(y_test, y_pred))
classifier.fit(X_train_processed,y_train_processed)
y_pred_processed = classifier.predict(X_test_processed)
y_pred_processed = np.round(y_pred_processed).flatten()
print(accuracy_score(y_test_processed, y_pred_processed))
classifier.fit(X_train_outliers,y_train_outliers)
y_pred_outliers = classifier.predict(X_test_outliers)
y_pred_outliers = np.round(y_pred_outliers).flatten()
print(accuracy_score(y_test_outliers, y_pred_outliers))
# + id="UpVGb0-p991g" colab_type="code" outputId="e903f4c5-6608-4d54-d5f9-47c9a77bd102" colab={"base_uri": "https://localhost:8080/", "height": 70}
from sklearn.linear_model import SGDClassifier
classifier = SGDClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
print(accuracy_score(y_test, y_pred))
classifier.fit(X_train_processed,y_train_processed)
y_pred_processed = classifier.predict(X_test_processed)
y_pred_processed = np.round(y_pred_processed).flatten()
print(accuracy_score(y_test_processed, y_pred_processed))
classifier.fit(X_train_outliers,y_train_outliers)
y_pred_outliers = classifier.predict(X_test_outliers)
y_pred_outliers = np.round(y_pred_outliers).flatten()
print(accuracy_score(y_test_outliers, y_pred_outliers))
# + id="nrl2BNRA9_mm" colab_type="code" outputId="68d759da-c112-464a-d156-674cba80f9fe" colab={"base_uri": "https://localhost:8080/", "height": 70}
from sklearn.linear_model import Perceptron
classifier = Perceptron()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
print(accuracy_score(y_test, y_pred))
classifier.fit(X_train_processed,y_train_processed)
y_pred_processed = classifier.predict(X_test_processed)
y_pred_processed = np.round(y_pred_processed).flatten()
print(accuracy_score(y_test_processed, y_pred_processed))
classifier.fit(X_train_outliers,y_train_outliers)
y_pred_outliers = classifier.predict(X_test_outliers)
y_pred_outliers = np.round(y_pred_outliers).flatten()
print(accuracy_score(y_test_outliers, y_pred_outliers))
# + id="L6SQhWSk-Btx" colab_type="code" outputId="f0000805-94ad-4711-e07a-2dc5497a9cd1" colab={"base_uri": "https://localhost:8080/", "height": 70}
from sklearn.naive_bayes import GaussianNB
classifier = GaussianNB()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
print(accuracy_score(y_test, y_pred))
classifier.fit(X_train_processed,y_train_processed)
y_pred_processed = classifier.predict(X_test_processed)
y_pred_processed = np.round(y_pred_processed).flatten()
print(accuracy_score(y_test_processed, y_pred_processed))
classifier.fit(X_train_outliers,y_train_outliers)
y_pred_outliers = classifier.predict(X_test_outliers)
y_pred_outliers = np.round(y_pred_outliers).flatten()
print(accuracy_score(y_test_outliers, y_pred_outliers))
# + id="Ty-k4q1L-Ew-" colab_type="code" outputId="5e272ee1-6f99-4571-e86d-3739ed28b55a" colab={"base_uri": "https://localhost:8080/", "height": 70}
from sklearn.neighbors import KNeighborsClassifier
classifier = KNeighborsClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
print(accuracy_score(y_test, y_pred))
classifier.fit(X_train_processed,y_train_processed)
y_pred_processed = classifier.predict(X_test_processed)
y_pred_processed = np.round(y_pred_processed).flatten()
print(accuracy_score(y_test_processed, y_pred_processed))
classifier.fit(X_train_outliers,y_train_outliers)
y_pred_outliers = classifier.predict(X_test_outliers)
y_pred_outliers = np.round(y_pred_outliers).flatten()
print(accuracy_score(y_test_outliers, y_pred_outliers))
# + id="D_-cjmP7NIpN" colab_type="code" colab={}
| FeatureEngineering_DataScience/Demo158_Outliers_Detection_EllipticEnvelope.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dask
# - [Dask 공식 문서](http://dask.pydata.org/en/latest/)
# - numpy, pandas, sklearn이랑 통합 가능
# - Dask is a flexible parallel computing library for analytic computing.
#
# - Dask is composed of two components:
#
# - **Dynamic task scheduling** optimized for computation. This is similar to Airflow, Luigi, Celery, or Make, but optimized for interactive computational workloads.
# - **“Big Data” collections** like parallel arrays, dataframes, and lists that extend common interfaces like NumPy, Pandas, or Python iterators to larger-than-memory or distributed environments. These parallel collections run on top of the dynamic task schedulers.
# 
# pip3 install dask로 하면 dask 베이직만 설치됨.
#
# ## Installation
# * pip install dask[complete]: Install everything
# * pip install dask[array]: Install dask and numpy
# * pip install dask[bag]: Install dask and cloudpickle
# * pip install dask[dataframe]: Install dask, numpy, and pandas
# * pip install dask: Install only dask, which depends only on the standard
# library. This is appropriate if you only want the task schedulers.
#
# We do this so that users of the lightweight core dask scheduler aren’t required
# to download the more exotic dependencies of the collections (numpy, pandas,
# etc.).
import dask
import pandas as pd
df = pd.read_csv('./user_log_2018_01_01.csv')
df
import dask.dataframe as dd
dask_df = dd.read_csv('./user_log_2018_01_01.csv')
dask_df
dir(dask_df)
dask_df["0"]
dask_df.index
len(dask_df.index)
dask_df.info
# ## Setup
# - 2 families of task scheduler
# - 1) Single machine scheduler : basic feature, default, does not scale
# - 2) Distributed scheduler : sophisticated, more feature, a bit more effort to set up
dask_df.head()
dask_df["user_id"].sum().compute()
dask_df["event_cnt"].sum().compute()
dask_df[dask_df["event_cnt"]>1].sum().compute()
from dask.distributed import Client
client2 = Client()
# client = Client(process=False)
dask_df["event_cnt"].sum().compute(scheduler='client2')
# - Single Machine
# - Default Scheduler : no-setup, local threads or process for larger than memory processing
# - Dask.distributed : newer system on a single machine. advanced features
# - Distributed computing
# - Manual SEtup : dask-scheduler and dask-worker
# - SSH
# - High Performance Computers
# - Kuberneters
# - Python API
# - Docker
# - Cloud
# with
with dask.config.set(scheduler='threads'):
x.compute()
y.compute()
# global setting
dask.config.set(scheduler='threads')
# ## LocalCluster
from dask.distributed import Client, LocalCluster
cluster = LocalCluster()
# class distributed.deploy.local.LocalCluster(n_workers=None, threads_per_worker=None, processes=True, loop=None, start=None, ip=None, scheduler_port=0, silence_logs=30, diagnostics_port=8787, services={}, worker_services={}, service_kwargs=None, asynchronous=False, **worker_kwargs)
client = Client(cluster)
cluster
client
# +
### Add a new worker to the cluster
# -
w = cluster.start_worker(ncores=2)
cluster.stop_worker(w)
# ## Command line
# dask-worker tcp://172.16.17.32:8786
# ## SSH
# ```
# pip3 install paramiko
# ```
# - dask-ssh 192.168.0.1 192.168.0.2 192.168.0.3 192.168.0.4
# - dask-ssh 192.168.0.{1,2,3,4}
# - dask-ssh --hostfile hostfile.txt
| python/dask.ipynb |