code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# our utils functions
from utils import *
# our classes
from classes.PreprocessData import *
from classes.EvaluateModel import *
from classes.ModelTest import *
# +
model_grid = [
{
'model_name': "NNs",
'model_type': "normal",
'model_dir': "models/NNs/mix"
},
{
'model_name': "1_all_NNs",
'model_type': "onevsall",
'model_dir': "models/1_all_NNs/mix"
},
{
'model_name': "HMMs",
'model_type': "onevsall",
'model_dir': "models/HMMs/mix"
}
]
# +
test_path='data/mix/data41mix_test.csv'
model_test = ModelTest(dict_path='dict41.txt', model_grid=model_grid, test_path=test_path, class_size=41)
model_test.init_models()
# -
model_test.get_model_result('two_2chunk-07.wav', "HMMs")
model_test.get_model_result('two_2chunk-07.wav', "1_all_NNs")
model_test.get_model_result('two_2chunk-07.wav', "NNs")
| src/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### K Means
#
#
#
# K means is the easiest form of unsupervised learning. Unsupervised learning is a type of self-organized learning that helps find previously unknown patterns in data set without pre-existing labels. Supervised learning is trying to find a boundary to separate data with different properties. Unsupervised learning is trying to aggregate data with similar properties. Think of unsupervised learning as a bottom-up approach and supervised learning is a top-down approach. Unsupervised learning is mostly used in cluster problems to sort data into different categories.
#
# The way K means works is very intuitive. Basically, we assign K centroids with different labels to random positions. Then the iteration begins. We compute the distance from each data point to K centroids. The label of the data point is determined by the centroid with the shortest distance. Each centroid is recalibrated to the mean of all the data points of the same label. We repeat the previous three steps until centroids do not move any more. Unsupervised learning done!
#
# For more technical details, feel free to read the following link (even though it is in R)
#
# https://towardsdatascience.com/k-means-a-complete-introduction-1702af9cd8c
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import imageio
import random as rd
import sklearn.cluster
import sklearn.decomposition
import os
os.chdir('K:/ecole/github')
#euclidean distance is the measure of the distance between two points in geometry
#other measures include manhattan distance
#actually u can use np.linalg.norm to save u time
def euclidean_distance(p1,p2):
assert len(p1)==len(p2),"p1 and p2 should be the same dimension"
dist=sum([i**2 for i in np.subtract(p1,p2)])
return dist**0.5
#for unsupervised learning, clf.score doesnt return the accuracy
#there is no cross validation, no known labels
#the only way to detect the accuracy is vote of the majority
#for each label given
#we check which iris type is the majority
#we consider the majority as the correct classification
#all we need to do is to count the minority
def get_accuracy(data,class_,checked):
df=data.copy()
#use dictionary to keep track of everything
d={}
#counting
for i in df['label'][df['y']==class_].unique():
if i not in checked and i!=-1:
d[i]=df['label'][df['y']==class_].tolist().count(i)
#comparison
maxval=-1
lbl=None
for i in d:
if d[i]>maxval:
lbl=i
maxval=d[i]
return len(df['label'][df['y']==class_][df['label']!=lbl])
#create random centroids
#centroids are bounded by the extreme value of our data points
def random_centroid(data):
x=data.copy()
centroid=[]
#the dimension of centroid should align to training dataset
#*100 then /100 to get decimal numbers
for i in x:
rdnum=rd.randint(int(min(x[i])*100),
int(max(x[i])*100))/100
centroid.append(rdnum)
return centroid
#the logic of kmeans is very intuitive
#assuming we are using 3 centroids (the selection of k is another topic)
#we insert 3 random centroids into the data
#we calculate the euclidean distance from each data point to each centroid
#each data point is mapped to the closest centroid
#once classification is done, we move the centroids to the centre of the cluster
#we keep repeating three steps above until the centroids dont move any move
#or the iteration has reached the limit set by us, which is 50 by default
#k means is simple but it has some shortcomings
#it does not always converge to the local optima
#if you run a few iterations you will see different results
#and its boundary is always linear
def kmeans(data1,data2,knum=3,itrlimit=50,fixed_pos=True,
show_acc=False,show_viz=False,genr_gif=False):
x=data1.copy()
df=data2.copy()
if fixed_pos==True:
centroids=[]
var=locals()
#using fixed position of centroids
#centroids will be scattered on the diagonal hyperspace
for i in x:
var[i.replace(' ','_')]=(max(x[i])-min(x[i]))/(knum+2)
for j in range(1,knum+1):
centroids.append([min(x[i])+var[i.replace(' ','_')]*(j) for i in x])
else:
#create random centroids
centroids=[]
for j in range(knum):
centroid=random_centroid(x)
#no duplicates
while centroid in centroids:
centroid=random_centroid(x)
centroids.append(centroid)
#converge is used to stop the loop when centroids dont move any more
#counter is to stop the infinite iteration
#on a very rare occasion, centroids would swing between 2 clusters
#in that sense, we stop the iteration
converge=False
counter=0
while not converge:
#calculate distance
labels=[]
for i in range(len(x)):
point=x.loc[i].tolist()
distance=[]
for j in centroids:
distance.append(euclidean_distance(point,j))
#set label of each data point as the closest centroid
labels.append(distance.index(min(distance)))
x['label']=labels
df['label']=labels
#visualization
if show_viz==True or genr_gif==True:
ax=plt.figure().add_subplot(111)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
plt.scatter(x['dimension 1'],x['dimension 2'],
c=labels,alpha=0.2,s=50,label='clusters')
plt.scatter([i[0] for i in centroids],
[i[1] for i in centroids],
c=range(knum),
s=200,marker='*',
edgecolors='k',
label='centroids')
plt.ylabel('Dimension 2')
plt.xlabel('Dimension 1')
plt.title('K Means')
plt.legend(loc='lower right')
if genr_gif==True:
#this line is used to create gif animation
plt.savefig('kmeans%d.png'%(counter))
plt.show()
elif show_viz==True:
plt.show()
else:
pass
#check if converged
centroids_prev=[i for i in centroids]
centroids=[]
for i in range(knum):
#sometimes one of the centroids is too far from any data point
#we have to reset the centroid
if x[x['label']==i].empty:
centroid=random_centroid(x)
while centroid in centroids:
centroid=random_centroid(x)
centroids.append(centroid)
#otherwise we update the centroids
#we move them to the centre of the cluster
else:
centroids.append(np.mean(x[x['label']==i]).tolist())
#two conditions to stop the iteration
#either converged or reaching iteration limit
counter+=1
if centroids==centroids_prev or counter>=itrlimit:
converge=True
#print accuracy
if show_acc==True:
erreur=0
checked=[]
for i in range(len(df['label'].unique())):
erreur+=get_accuracy(df,i,checked)
checked.append(i)
accuracy=1-erreur/len(df)
print('accuracy: %s'%(accuracy))
#create gif
if genr_gif==True:
filenames=['kmeans%d.png'%(i) for i in range(counter)]
images=list(map(lambda filename:imageio.imread(filename),
filenames))
imageio.mimsave('kmeans.gif',images,duration=0.8)
return labels
#kmeans implemented by sklearn
def skl_kmeans(data1,data2,n=3):
x=data1.copy()
df=data2.copy()
clf=sklearn.cluster.KMeans(n_clusters=n)
df['label']=clf.fit_predict(x)
#compute accuracy
erreur=0
checked=[]
for i in range(len(df['label'].unique())):
erreur+=get_accuracy(df,i,checked)
checked.append(i)
accuracy=1-erreur/len(df)
print('accuracy: %s'%(accuracy))
df=pd.read_csv('iris.csv')
x=pd.concat([df['sepal length'], \
df['sepal width'], \
df['petal length'],\
df['petal width']],axis=1)
#convert text to discrete number
df['y']=np.unique(df['type'],return_inverse=True)[1]
#for the purpose of visualization
#we reduce 4 dimensions to 2
#more details of pca can be found in the link below
# https://github.com/je-suis-tm/machine-learning/blob/master/principal%20component%20analysis.ipynb
dims=2
x=sklearn.decomposition.PCA(n_components=dims).fit_transform(x)
x=pd.DataFrame(x,columns=[f'dimension {i}' for i in range(1,dims+1)])
df['label']=kmeans(x,df,show_acc=True,
show_viz=True,fixed_pos=False,
genr_gif=True)
skl_kmeans(x,df)
# 
#
#
# ### Selection of K
#
#
#
# The key part of K Means is the selection of K. There are many metrics used in unsupervised learning. Here we only talk about 3 of the most common ones for K Means. All of them are brute force calculation. We have a fancy word for that, grid search. Run an iteration of different amounts of centroids and use some metrics to evaluate the effect.
# #### Elbow Method
#
#
#
# It is called inertia in scikit-learn. It is the computation of within-cluster sum of squared error (WSSE). WSSE refers to the sum of the euclidean distance from each data point to its underlying centroid. It is quite intuitive that the overall distance gets smaller as we increase the amount of centroids. Think of it as more variables lead to the increase of R squared in OLS.
#
# The idea of elbow method is to find the optimal spot where you have a reasonable amount of centroid and smaller overall distance. It is where inertia curve takes a sharp decline. We call the critical point 'The Elbow'. To identify the sweet spot, we create a new line by connecting the start of inertia curve to the end. The G spot exists at the largest perpendicular distance from the newly formed line to inertia curve.
#plot two curves on separate axis
def dual_axis_plot(xaxis,data1,data2,fst_color='r',
sec_color='b',fig_size=(10,5),x_label='',
y_label1='',y_label2='',grid=False,title='',
legend1='',legend2=''):
fig=plt.figure(figsize=fig_size)
ax=fig.add_subplot(111)
ax.set_xlabel(x_label)
#differentiate two y axis by different colors
ax.set_ylabel(y_label1, color=fst_color)
ax.plot(xaxis, data1, color=fst_color,label=legend1)
ax.tick_params(axis='y',labelcolor=fst_color)
ax.yaxis.labelpad=15
#legend of y1 goes to the left
plt.legend(loc=3)
#the crucial part of dual axis plot
ax2 = ax.twinx()
ax2.set_ylabel(y_label2, color=sec_color,rotation=270)
ax2.plot(xaxis, data2, color=sec_color,label=legend2)
ax2.plot(xaxis[data2.index(max(data2))],max(data2),
marker='*',markersize=25,lw=0,color='#b23850',
alpha=0.7,label='optimal')
ax2.tick_params(axis='y',labelcolor=sec_color)
ax2.yaxis.labelpad=15
fig.tight_layout()
plt.legend(loc=4)
plt.grid(grid)
plt.title(title)
plt.show()
# +
#using geometry to calculate the perpendicular distance
def get_distance(x,y,a,b):
numerator=y-x*a-b
denominator=(a**2+1)**0.5
return np.abs(numerator/denominator)
#simple solution to get coefficients of the equation
def get_line_params(x1,y1,x2,y2):
a=(y1-y2)/(x1-x2)
b=y1-a*x1
return a,b
# -
#calculate intra-cluster sum of squared error
#some people call it within-cluster sum of squared error
#it is the sum of the euclidean distance from each point to its underlying centroid
#in sklearn, it is an attribute called inertia
def get_inertia(data):
x=data.copy()[[i for i in data.columns if i!='label']]
wsse=[]
for i in data['label'].unique():
#centroid is basically the mean of a cluster
centroid=np.mean(x[data['label']==i]).tolist()
for j in x.loc[data['label']==i].index:
wsse.append(euclidean_distance(x.loc[j].tolist(),
centroid))
return sum(wsse)
# +
#calculate wsse
inertia=[]
for i in range(2,8):
x['label']=kmeans(x[[i for i in x.columns if i!='label']],
df,knum=i)
inertia.append(get_inertia(x))
# +
#connect the start of inertia curve to the end
a,b=get_line_params(0,inertia[0],len(inertia)-1,inertia[-1])
#calculate the perpendicular distance
#from each point on inertia curve to the straight line
distance=[]
for i in range(len(inertia)):
distance.append(get_distance(i,inertia[i],a,b))
#use dual axis visualization
dual_axis_plot(np.arange(2,8),inertia,distance,
x_label='Numbers of Cluster',
y_label1='Within-cluster Sum of Squared Error',
y_label2='Perpendicular Distance',
legend1='Inertia',legend2='Distance',
title='Elbow Method',
fst_color='#b39bc8',sec_color='#464866')
# -
# #### Silhouette Score
#
#
#
# Silhouette score or silhouette coefficient is a more scientific approach to determine the amount of the clusters. The numerator consists of similarity and dissimilarity. Similarity implies the density of inside the cluster. The smaller the better. Dissimilarity implies the distance among clusters. The larger the better (think of it as functional margin in svm). The overall silhouette score proposed by Kaufman represents the mean silhouette over the entire dataset. We select K to maximize the silhouette score.
#
# For the details of the computation, please refer to Wikipedia
#
# https://en.wikipedia.org/wiki/Silhouette_(clustering)
#calculate mean distance from one point to a cluster
#it is the mean of euclidean distance from one point to all the points in the cluster
def mean_distance(point,dataset):
distance_list=[]
assert not dataset.empty,'empty dataset'
for i in dataset.index:
#make sure we dont include the point itself
if dataset.loc[i].tolist()!=point:
distance_list.append(
euclidean_distance(point,
dataset.loc[i].tolist()))
#if one cluster has only one element which is the point itself
#we will return zero
if len(dataset)==1 and dataset.iloc[0].tolist()==point:
return 0
#we raise warning here
#because np.mean([]) returns np.nan
#it will cause a huge problem for silhouette computation
import warnings
warnings.simplefilter('error')
return np.mean(distance_list)
#silhouette coefficient is the maximum value of the mean silhouette for each label
#in sklearn, it is sklearn.metrics.silhouette_score
#to calculate silhouette for each data point in the dataset
#we need to create multiple loops
#so the time complexity of silhouette score is way higher than other metrics
def silhouette_coefficient(data):
x=data.copy()[[i for i in data.columns if i!='label']]
cols=x.columns.tolist()
#if there is only one cluster, silhouette score is zero
if len(data['label'].unique())==1:
return 0
silhouette=[]
#calculate silhouette for each data point
for i in range(len(x)):
lbl=data['label'][i]
#similarity is mean distance from one point to every other point within the same cluster
similarity=mean_distance(x[cols].loc[i].tolist(),
x[cols].loc[data['label']==lbl])
arr=[]
otherlbl=[i for i in set(data['label']) if i!=lbl]
#dissimilarity is mean distance from one point to all the points in the other clusters
for j in otherlbl:
arr.append(mean_distance(x[cols].loc[i].tolist(),
x[cols].loc[data['label']==j]))
dissimilarity=min(arr)
silhouette.append((dissimilarity-similarity)/max(similarity,dissimilarity))
x['silhouette']=silhouette
silhouette=[]
for i in set(data['label']):
silhouette.append(np.mean(x['silhouette'][data['label']==i].tolist()))
return max(silhouette)
#calculate silhouette score
silhouette_score=[]
for i in range(2,8):
#avoid using label as one of the features
x['label']=kmeans(x[[i for i in x.columns if i!='label']],
df,knum=i)
silhouette_score.append(silhouette_coefficient(x))
# +
#visualize silhouette
ax=plt.figure(figsize=(10,5)).add_subplot(111)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
plt.plot([i for i in range(2,8)],silhouette_score,c='#0294a5',)
plt.plot(silhouette_score.index(max(silhouette_score))+2,
max(silhouette_score),lw=0,c='#f79e02',alpha=0.8,
marker='*',markersize=15,label='Optimal')
plt.legend(loc=0)
plt.ylabel('Silhouette Score')
plt.xlabel('Numbers of Cluster')
plt.title('Silhouette Analysis')
plt.show()
# -
# #### Gap Statistic
#
#
#
# Gap statistic is a relatively new metric compared to the others. It was invented 20 years ago by some scholars from Stanford University. Sklearn has not included gap statistic in its package. Gap statistic is the logarithm difference between mean WSSE of reference data and WSSE of real data. We select K to satisfy the gap statistic K plus standard deviation of reference data WSSE K+1 is larger than the gap statistic K+1. Similar to autocorrelation plot in time series analysis, we pick the smallest K.
#
# For coding reference, feel free to check anaconda notebook
#
# https://anaconda.org/milesgranger/gap-statistic/notebook
#
# For math reference, feel free to check the original paper
#
# https://statweb.stanford.edu/~gwalther/gap
#there are two ways to generate the reference data
#here we take the first approach from the paper
#using uniform distribution for simplicity
#the distribution is bounded by the extreme value of the original dataset
def get_reference_data(x):
montecarlo=[]
for i in range(len(x)):
arr=[]
for j in x.columns:
if j!='label':
rdnum=rd.uniform(min(x[j]),
max(x[j]))
arr.append(rdnum)
montecarlo.append(arr)
output=pd.DataFrame()
for i in x.columns:
if i!='label':
output[i]=[j[x.columns.tolist().index(i)] for j in montecarlo]
return output
# +
#use monte carlo simulation to generate reference data
simulation=10
var=locals()
for i in range(simulation):
var['ref'+str(i)]=get_reference_data(x[[i for i in x.columns if i!='label']])
# -
gap_stat=[]
gap_std=[]
for i in range(2,8):
#avoid using label as one of the features
x['label']=kmeans(x[[i for i in x.columns if i!='label']],
df,knum=i)
#get expectation of gap statistic from all reference data
mc_simu=[]
for j in range(simulation):
var['ref'+str(j)]['label']=kmeans(var['ref'+str(j)][[i for i in x.columns if i!='label']],
x,knum=i)
mc_simu.append(get_inertia(var['ref'+str(j)]))
gap_stat.append(np.mean([np.log(i) for i in mc_simu])-np.log(get_inertia(x)))
gap_std.append((1+1/simulation)**0.5*np.std([np.log(i) for i in mc_simu]))
# +
#visualize gap statistic
ax=plt.figure(figsize=(10,5)).add_subplot(111)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
plt.errorbar([i for i in range(2,8)],gap_stat,gap_std,
c='#05386b',capsize=5)
ind=None
for i in range(len(gap_stat)-1):
if gap_stat[i]+gap_std[i+1]>gap_stat[i+1]:
ind=i
break
plt.plot(ind+2,gap_stat[ind],
lw=0,c='#fc4445',alpha=0.8,
marker='*',markersize=20,label='Optimal')
plt.legend(loc=0)
plt.ylabel('Gap Statistic')
plt.xlabel('Numbers of Cluster')
plt.title('Gap Statistic')
plt.show()
# -
| k means.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + tags=[]
import xarray as xr
import sys
import random
from scipy import stats
import glob
import cartopy.crs as ccrs
from cartopy.mpl.ticker import LongitudeFormatter, LatitudeFormatter
import shapely.geometry as sgeom
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import numpy as np
import matplotlib
# %matplotlib inline
# + tags=[]
func = lambda x: x.split('comp_')[1].split('_{}days'.format(time_scale))[0]
# + tags=[]
its = 10000
DJF_bool = 'DJF'
DJF_bool = '_{}only'.format(DJF_bool)
cesta_root = '/mnt/nas4.meop2/meop40.data.model/CMAM/0A.daily/'
cesta = f'{cesta_root}composites_woSSW{DJF_bool}/'
print(cesta)
time_scale = '20'
lw = 2
comp_name_ls = ['himalayas', 'eastasia', 'westamer',]
size_dict = {'20': [45,74,36], '30': [38,66,35]}
line_width = 5
box = sgeom.box(minx=70, maxx=105, miny=20, maxy=42)
box2 = sgeom.box(minx=110, maxx=145, miny=30, maxy=48)
box3 = sgeom.box(minx=-125, maxx=-102.5, miny=27.5, maxy=52)
boxes = [box, box2, box3]
projection = ccrs.PlateCarree(central_longitude=0)#Robinson()# Orthographic(180, 90)
proj_str = str(projection).split('crs.')[1].split(' object')[0]
# + tags=[]
var = 'accelogw'
ch_lev = 7000
what = 'absolute'
scale = 3600*24
if var.lower() in ['lwa']:
lev_sys_fo = 'logH/'
lev_sys_fi = '_logH'
sel_dict = dict(z = ch_lev)
units = 'm'
else:
lev_sys_fo = ''
lev_sys_fi = ''
sel_dict = dict(plev = ch_lev)
units = 'Pa'
outfile_name = '{}{}{}_pvalues_from{}_comp_*_{}days@{}Pa.nc'.format(cesta, var, lev_sys_fi, its, time_scale, ch_lev)
print(outfile_name)
outfile_name_ls = glob.glob(outfile_name)
#ds_pv = xr.open_mfdataset(outfile_name_ls, concat_dim='e_type')
#ds_pv['e_type'] = list(map(func, outfile_name_ls))
comp_file = '{}{}{}_{}_comp_*_{}days.nc'.format(cesta, var, lev_sys_fi, what, time_scale)
comp_file_ls = glob.glob(comp_file)
ds_comp = xr.open_mfdataset(comp_file_ls, combine='nested', concat_dim='e_type')
ds_comp['e_type'] = list(map(func, comp_file_ls))
ds_comp = ds_comp.sel(**sel_dict).sel(lag = 0)*scale
ds_comp[var].attrs['units'] = r'm/s/day'
ds_comp[var].attrs['long_name'] = r'OGWD'
comp_file = f'{cesta_root}/{var}/{lev_sys_fo}{var}_climatology_woSSW.nc'
print(comp_file)
sel_dict['month'] = [12, 1, 2]
ds_comp2 = xr.open_dataset(comp_file).sel(**sel_dict).mean('month')[var]*scale
ds_comp2
# + tags=[]
plt.rcParams.update({'font.size': 22})
ds_comp[var].sel(e_type = 'westamer').sel(lat = 60, method='nearest').plot(size = 6, lw = 3)
# + tags=[]
projection = ccrs.PlateCarree(central_longitude=180)
title_ls = ['Himalayas', 'East Asia', 'Rocky Mountains']
p = (ds_comp[var].sel(e_type = comp_name_ls)).plot.contourf(col = 'e_type', levels = 21, robust = False, \
col_wrap = 1, \
transform=ccrs.PlateCarree(), \
aspect=ds_comp.dims['lon']/ds_comp.dims['lat'], \
size = 5, \
extend = 'both', \
subplot_kws={'projection': projection}, \
cbar_kwargs = {'orientation': 'horizontal', \
'pad': 0.05})
fmt = ticker.LogFormatterMathtext()
fmt.create_dummy_axis()
for i,(ax, sel_dict, box) in enumerate(zip(p.axes.flat, p.name_dicts.flat, boxes)):
ax.coastlines()
ax.gridlines(xlocs = [-180, -90, 0, 90, 180])
ax.set_yticks([0, 20, 40, 60, 80], crs=ccrs.PlateCarree())
lat_formatter = LatitudeFormatter(number_format='.1f', \
degree_symbol='')
ax.yaxis.set_major_formatter(lat_formatter)
if i == 2:
ax.set_xticks([-90, 0, 90, 180], crs=ccrs.PlateCarree())
lon_formatter = LongitudeFormatter(number_format='.1f', \
degree_symbol='', \
dateline_direction_label=True)
ax.xaxis.set_major_formatter(lon_formatter)
CS = (ds_comp2).plot.contour(levels = [-3,-2,-1], \
colors =['purple','gray','k'], \
ax = ax, linewidths = lw, \
transform=ccrs.PlateCarree())
if i == 1:
labels = [r"OGWD$_{clim}$ = -3 m/s/day",
r"OGWD$_{clim}$ = -2 m/s/day",
r"OGWD$_{clim}$ = -1 m/s/day"]
for j in range(len(labels)):
CS.collections[j].set_label(labels[j])
ax.legend(loc='lower right')
#plt.legend(proxy, )
#plt.clabel(CS, inline=1, fontsize=14)
CS = ds_comp2.plot.contour(levels = [0], colors = 'k', \
ax = ax, linewidths = lw*2, \
transform=ccrs.PlateCarree())
if i == 0:
labels = [r"OGWD$_{clim}$ = 0 m/s/day"]
for j in range(len(labels)):
CS.collections[j].set_label(labels[j])
ax.legend(loc='lower right')
#, fmt = fmt)
if proj_str == 'PlateCarree':
ax.set_extent([0, 270, -10, 80])#ax.set_extent([-180, 180, -10, 80])
ax.set_aspect('auto')
plot_kwargs2 = dict(levels = [0,0.05], hatches=['\\\\',None], \
colors='none', add_colorbar=False, transform=ccrs.PlateCarree())
#temp = ds_pv[var].sel(**sel_dict)#*24*3600
#temp.plot.contourf(ax = ax, **plot_kwargs2)
plot_kwargs2['levels'] = [0,0.01]
plot_kwargs2['hatches'] = ['////',None]
#temp.plot.contourf(ax = ax, **plot_kwargs2) #ax.contourf(temp.lon, temp.lat, temp.values, **plot_kwargs2)
ax.set_title(title_ls[i], fontdict = dict(fontweight = 'bold')) # sel_dict['e_type']
ax.add_geometries([box], ccrs.PlateCarree(), \
facecolor=['none'], edgecolor=['green'], \
linewidth = [4])
ax.set_xlabel('')
ax.set_ylabel('')
out_fig = f'{var}@{ch_lev}{units}_{what}_allwclim_{time_scale}days_{proj_str}_{DJF_bool}.pdf'
print(out_fig)
#plt.savefig(out_fig, bbox_inches='tight')
| code/GRL_OGWD_absolute_composite.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Space Complexity is the maximum space required by a program at any point in time
n = int(input())
i = 1 #Space Complexity = K * n^0 = O(1)
while i <= n:
print(i)
i += 1
# +
i = 1 # k1 space
while i <= n:
j = 0 # k2 space
print(i)
i += 1
# Space Complexity = k1 + n*k2 = O(n) (Wrong Answer)
# Space Complexity is the maximum space required by a program at any point in time
# So, for our progam at any point in time we require only k2 memory so overall will be k1 + k2 = O(1)
# Space Complexity = k1 + k2 = O(1)
# -
# #### For a list of 'n' size then the space complexity is O(n)
# ### Three points to be kept in mind while calculating the space complexity:
# #### 1) Calculating the space complexity in point in time not the overall complexity
# #### 2) Auxillary Space - Extra space to your alogorithm. For example, space complexity of Bubble Sort is O(1) because we're not using any extra space
# #### 3) Recursion takes space. For example, fact 5 --> fact 4 --> fact 3 --> fact 2 --> fact 1 (--> means calls)
#
# ## Tip: Don't count the input space requirement in your space complexity
#
#
# +
# Bubble Sort
list = [3,9,6,4,2]
for i in range(0, len(list)):
for j in range(0, len(list)-i-1):
if list[j] > list[j+1]:
list[j], list[j+1] = list[j+1], list[j]
print(list)
# No extra space so O(1)
# +
# Factorial Recursive:
fact(n):
if n == 0:
return 1
return n*fact(n)
# Recursive function stores the calls in a stack which requires memory. When you reach end of the recursion you'll have 'n+1' functions in the memory. Each taking 'k' space. So, overall space complexity is O(n)
# -
# Space Complexity
# Bubble Sort - O(1)
# Factorial(Iterative) - O(1)
# Factorial(Recursive) - O(n)
# Merge Sort - O(n)
| 07 Space Complexity/Space Complexity Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Shadow Rollout with Seldon and Ambassador
#
# This notebook shows how you can deploy "shadow" deployments to direct traffic not only to the main Seldon Deployment but also to a shadow deployment whose reponse will be dicarded. This allows you to test new models in a production setting and with production traffic and anlalyse how they perform before putting them live.
#
# These are useful when you want to test a new model or higher latency inference piepline (e.g., with explanation components) with production traffic but without affecting the live deployment.
#
# ## Prerequistes
#
# You will need
#
# - [Git clone of Seldon Core](https://github.com/SeldonIO/seldon-core) running this notebook
# - A running Kubernetes cluster with kubectl authenticated
# - [Helm client](https://helm.sh/)
# - Seldon Core Python Module : `pip install seldon-core`
# ### Creating a Kubernetes Cluster
#
# Follow the [Kubernetes documentation to create a cluster](https://kubernetes.io/docs/setup/).
#
# Once created ensure ```kubectl``` is authenticated against the running cluster.
# ## Setup
# !kubectl create namespace seldon
# !kubectl config set-context $(kubectl config current-context) --namespace=seldon
# !kubectl create clusterrolebinding kube-system-cluster-admin --clusterrole=cluster-admin --serviceaccount=kube-system:default
# ## Install Helm
# !kubectl -n kube-system create sa tiller
# !kubectl create clusterrolebinding tiller --clusterrole cluster-admin --serviceaccount=kube-system:tiller
# !helm init --service-account tiller
# !kubectl rollout status deploy/tiller-deploy -n kube-system
# ## Start seldon-core
# !helm install ../../../helm-charts/seldon-core-operator --name seldon-core --set usageMetrics.enabled=true --namespace seldon-system
# !kubectl rollout status statefulset.apps/seldon-operator-controller-manager -n seldon-system
# Install the Prometheus and Grafana example analytics
# !helm install ../../../helm-charts/seldon-core-analytics --name seldon-core-analytics --set grafana_prom_admin_password=password --set persistence.enabled=false --namespace seldon
# ## Setup Ingress
# There are gRPC issues with the latest Ambassador, so we rewcommend 0.40.2 until these are fixed.
# !helm install stable/ambassador --name ambassador --set crds.keep=false
# !kubectl rollout status deployment.apps/ambassador
# ## Set up Port Forwards
#
# **Ensure you port forward ambassador**:
#
# ```
# kubectl port-forward $(kubectl get pods -n seldon -l app.kubernetes.io/name=ambassador -o jsonpath='{.items[0].metadata.name}') -n seldon 8003:8080
# ```
#
# **Ensure you port forward to Grafana**
#
# ```
# kubectl port-forward $(kubectl get pods -n seldon -l app=grafana-prom-server -o jsonpath='{.items[0].metadata.name}') -n seldon 3000:3000
# ```
# ## Launch main model
#
# We will create a very simple Seldon Deployment with a dummy model image `seldonio/mock_classifier:1.0`. This deployment is named `example`.
# !pygmentize model.json
# !kubectl create -f model.json
# !kubectl rollout status deploy/production-model-single-7cd068f
# ### Get predictions
from seldon_core.seldon_client import SeldonClient
sc = SeldonClient(deployment_name="example",namespace="seldon")
# #### REST Request
r = sc.predict(gateway="ambassador",transport="rest")
print(r)
# #### gRPC Request
r = sc.predict(gateway="ambassador",transport="grpc")
print(r)
# ## Launch Shadow
#
# We will now create a new Seldon Deployment for our Shadow deployment with a new model `seldonio/mock_classifier_rest:1.1`. To make it a shadow of the original `example` deployment we add two annotations
#
# ```
# "annotations": {
# "seldon.io/ambassador-service-name":"example",
# "seldon.io/ambassador-shadow":"true"
# },
# ```
#
# The first says to use `example` as our service endpoint rather than the default which would be our deployment name - in this case `example-shadow`. This will ensure that this Ambassador setting will apply to the same prefix as the previous one. The second states we want to use Ambassador's shadow functionality.
# !pygmentize shadow.json
# !kubectl create -f shadow.json
# !kubectl rollout status deploy/shadow-model-single-4c8805f
# Let's send a bunch of requests to the endpoint.
for i in range(1000):
r = sc.predict(gateway="ambassador",transport="rest")
# Now view the analytics dashboard at http://localhost:3000
# You should see a dashboard view like below showing the two models production and shadow both receiving requests.
#
# 
#
| examples/ambassador/shadow/ambassador_shadow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit
# name: python395jvsc74a57bd0aee8b7b246df8f9039afb4144a1f6fd8d2ca17a180786b69acc140d282b71a49
# ---
import time
time.time()
# 将时间戳转换为time struct
time.localtime()
time.localtime(time.time())
# 将time struct转时间戳
time.mktime(time.localtime())
# 格式化输出IOSDATE
time.strftime("%Y-%m-%dT%H:%M:%SZ", time.localtime())
# 输出今天周几
time.strftime("%w %a %A", time.localtime())
# 格式化输入
format_time = time.strptime("2017-11-11 18:58:39", "%Y-%m-%d %H:%M:%S")
print(format_time)
# +
# 日期格式化符号:
# - %y 两位数的年份表示(00-99)
# - %Y 四位数的年份表示(000-9999)
# - %m 月份(01-12)
# - %d 月内中的一天(0-31)
# - %H 24小时制小时数(0-23)
# - %I 12小时制小时数(01-12)
# - %M 分钟数(00=59)
# - %S 秒(00-59)
# - %a 本地简化星期名称
# - %A 本地完整星期名称
# - %b 本地简化的月份名称
# - %B 本地完整的月份名称
# - %c 本地相应的日期表示和时间表示
# - %j 年内的一天(001-366)
# - %p 本地A.M.或P.M.的等价符
# - %U 一年中的星期数(00-53)星期天为星期的开始
# - %w 星期(0-6),星期天为星期的开始
# - %W 一年中的星期数(00-53)星期一为星期的开始
# - %x 本地相应的日期表示
# - %X 本地相应的时间表示
# - %Z 当前时区的名称
# - %% %号本身
| lang/python/examples/library/time.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from datascience import *
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plots
plots.style.use('fivethirtyeight')
# -
# ## The GSI's Defense ##
scores = Table.read_table('scores_by_section.csv')
scores
scores.group('Section')
scores.group('Section', np.average).show()
observed_average = 13.6667
random_sample = scores.sample(27, with_replacement=False)
random_sample
np.average(random_sample.column('Midterm'))
# +
# Simulate one value of the test statistic
# under the hypothesis that the section is like a random sample from the class
def random_sample_midterm_avg():
random_sample = scores.sample(27, with_replacement = False)
return np.average(random_sample.column('Midterm'))
# +
# Simulate 50,000 copies of the test statistic
sample_averages = make_array()
for i in np.arange(50000):
sample_averages = np.append(sample_averages, random_sample_midterm_avg())
# +
# Compare the simulated distribution of the statistic
# and the actual observed statistic
averages_tbl = Table().with_column('Random Sample Average', sample_averages)
averages_tbl.hist(bins = 20)
plots.scatter(observed_average, 0, color = 'red', s=40);
# -
sum(sample_averages <= observed_average) / 50000
# +
# 5% of 50,000 = 2500
five_percent_point = averages_tbl.sort(0).column(0).item(2500)
five_percent_point
# -
averages_tbl.hist(bins = 20)
plots.plot([five_percent_point, five_percent_point], [0, 0.35], color='gold', lw=2)
plots.title('Area to the left of the gold line: 5%');
| lec/lec18.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CMPSC 100: The Dealership
#
# 
# <NAME> -- like anyone these days -- is looking for some part-time, passive income. And, like any reasonable person, they've hit on the idea of becoming an art dealer. I mean, people will pay a gazillion dollars for anything _artistic_, right? Despite any experience in the trade, <NAME> thinks this is a _great idea_.
#
# There's just one big problem: <NAME> doesn't know any artists, except -- now -- you. <NAME> is, however, savvy about one thing: trends. And, currently, the Gator Kingdom is all about abstract art. Looking through the local auction house's collection, <NAME> noticed that the following painting sold for `UNDISCLOSED SUM`, but that probably means it's A LOT OF DOLLARS:
#
# 
# ## Requirements
#
# ---
# `Notebook`
#
# This notebook should create an image which:
#
# * Is `1920` x `1080` pixels large
# * Uses at least one of each of the following shapes (more interesting pictures use _many_):
# * `ellipse`
# * `rectangle`
# * `polygon`
# * Here's the ["formal" definition of the method](https://pillow.readthedocs.io/en/stable/reference/ImageDraw.html#PIL.ImageDraw.ImageDraw.polygon)
# * Adds these shapes using a random approach written in a function
# * Has a background other than black or white
# * Pastes G. Wiz's picture in the bottom right corner (he has to have a branded picture for the catalog)
# * Use the `g_wiz_mark.png` file located in this directory
# * Has a title included in the `reflection.md` file
# * Of course, this lab presupposes a _complete_ `reflection.md` file
# * Is saved in this directory as `the_art.png`
#
# When you're done, congratulations -- you just made computational art. (That G. Wiz makes all the profit from.)
# ## Code space
#
# ---
| lab/Week 08 - Lab - The Dealership.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.9 64-bit
# name: python3
# ---
# # Resume Job Postings
#
# 1. Extracting raw text from job postings
# Setup
import glob
import pandas as pd
from bs4 import BeautifulSoup as bs
# List of files in the html directory
files = glob.glob('./data/html_job_postings//*.html')
print('Number of HTML files: {}'.format(len(files)))
# Load all HTML pages as text into a list -- one entry per HTML page
html_content = []
for file in files:
with open(file, 'r', encoding='utf-8') as f:
html_content.append(f.read())
# Inspect the first entry of the list of resumes
html_content[0]
# +
# Store webpage section with skills
html_sections = []
html_dict = {}
for key in ['title', 'body', 'bullets']:
html_dict[key] = []
# Prototype with first page of the HTML list
first_page = html_content[0]
soup = bs(first_page, "html.parser")
# Get key parts
title = soup.find('title').text
body = soup.find('body').text
bullets = soup.find_all('li')
html_dict['title'].append(title)
html_dict['body'].append(body)
# Cleaning bullets content
html_dict['bullets'].append([b.text.strip() for b in bullets])
df = pd.DataFrame(data=html_dict)
df.head()
# -
# Apply prototype in a general function
def get_html_content(html_content):
"""Extracts title, and list items from HTML job postings.
Args:
html_content (list): List of html resumes.
Return:
df: Returns a pandas dataframe with separate columns for title, body, and bullet items."""
html_sections = []
html_dict = {}
for key in ['title', 'body', 'bullets']:
html_dict[key] = []
for html in html_content:
soup = bs(html, "html.parser")
title = soup.find('title').text
body = soup.find('body').text
bullets = soup.find_all('li')
html_dict['title'].append(title)
html_dict['body'].append(body)
# remove extra leading and trailing whitespace with strip()
html_dict['bullets'].append([b.text.strip() for b in bullets])
df = pd.DataFrame(html_dict)
return df
df = get_html_content(html_content)
df.head()
df.shape
# Check how many data science/scientist jobs in the title
df[df['title'].str.contains('(data scientist)|(data science)|(machine learning)', case=False)].shape
# Filter only data science jobs
ds_df = df[df['title'].str.contains('(data scientist)|(data science)|(machine learning)', case=False)]
ds_df.head()
# Drop job postings duplicates
ds_df['bullets'] = ds_df['bullets'].apply(tuple, 1)
ds_df.drop_duplicates(inplace=True)
ds_df.shape
# Save the dataframe to disk
ds_df.to_pickle('step1_df.pk')
| Resume_job_postings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hierarchical Model for Abalone Length
#
# Abalone were collected from various sites on the coast of California north of San Francisco. Here I'm going to develop a model to predict abalone lengths based on sites and harvest method - diving or rock-picking. I'm interested in how abalone lengths vary between sites and harvesting methods. This should be a hierarchical model as the abalone at the different sites are from the same population and should exhibit similar effects based on harvesting method. The hierarchical model will be beneficial since some of the sites are missing a harvesting method.
# +
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
import sampyl as smp
from sampyl import np
import pandas as pd
# -
plt.style.use('seaborn')
plt.rcParams['font.size'] = 14.
plt.rcParams['legend.fontsize'] = 14.0
plt.rcParams['axes.titlesize'] = 16.0
plt.rcParams['axes.labelsize'] = 14.0
plt.rcParams['xtick.labelsize'] = 13.0
plt.rcParams['ytick.labelsize'] = 13.0
# Load our data here. This is just data collected in 2017.
data = pd.read_csv('Clean2017length.csv')
data.head()
# Important columns here are:
#
# * **full lengths:** length of abalone
# * **mode:** Harvesting method, R: rock-picking, D: diving
# * **site_code:** codes for 15 different sites
#
# First some data preprocessing to get it into the correct format for our model.
# +
# Convert sites from codes into sequential integers starting at 0
unique_sites = data['site_code'].unique()
site_map = dict(zip(unique_sites, np.arange(len(unique_sites))))
data = data.assign(site=data['site_code'].map(site_map))
# Convert modes into integers as well
# Filter out 'R/D' modes, bad data collection
data = data[(data['Mode'] != 'R/D')]
mode_map = {'R':0, 'D':1}
data = data.assign(mode=data['Mode'].map(mode_map))
# -
# ## A Hierarchical Linear Model
#
# Here we'll define our model. We want to make a linear model for each site in the data where we predict the abalone length given the mode of catching and the site.
#
# $$ y_s = \alpha_s + \beta_s * x_s + \epsilon $$
#
# where $y_s$ is the predicted abalone length, $x$ denotes the mode of harvesting, $\alpha_s$ and $\beta_s$ are coefficients for each site $s$, and $\epsilon$ is the model error. We'll use this prediction for our likelihood with data $D_s$, using a normal distribution with mean $y_s$ and variance $ \epsilon^2$ :
#
# $$ \prod_s P(D_s \mid \alpha_s, \beta_s, \epsilon) = \prod_s \mathcal{N}\left(D_s \mid y_s, \epsilon^2\right) $$
#
# The abalone come from the same population just in different locations. We can take these similarities between sites into account by creating a hierarchical model where the coefficients are drawn from a higher-level distribution common to all sites.
#
# $$
# \begin{align}
# \alpha_s & \sim \mathcal{N}\left(\mu_{\alpha}, \sigma_{\alpha}^2\right) \\
# \beta_s & \sim \mathcal{N}\left(\mu_{\beta}, \sigma_{\beta}^2\right) \\
# \end{align}
# $$
class HLM(smp.Model):
def __init__(self, data=None):
super().__init__()
self.data = data
# Now define the model (log-probability proportional to the posterior)
def logp_(self, μ_α, μ_β, σ_α, σ_β, site_α, site_β, ϵ):
# Population priors - normals for population means and half-Cauchy for population stds
self.add(smp.normal(μ_α, sig=500),
smp.normal(μ_β, sig=500),
smp.half_cauchy(σ_α, beta=5),
smp.half_cauchy(σ_β, beta=0.5))
# Priors for site coefficients, sampled from population distributions
self.add(smp.normal(site_α, mu=μ_α, sig=σ_α),
smp.normal(site_β, mu=μ_β, sig=σ_β))
# Prior for likelihood uncertainty
self.add(smp.half_normal(ϵ))
# Our estimate for abalone length, α + βx
length_est = site_α[self.data['site'].values] + site_β[self.data['site'].values]*self.data['mode']
# Add the log-likelihood
self.add(smp.normal(self.data['full lengths'], mu=length_est, sig=ϵ))
return self()
# +
sites = data['site'].values
modes = data['mode'].values
lengths = data['full lengths'].values
# Now define the model (log-probability proportional to the posterior)
def logp(μ_α, μ_β, σ_α, σ_β, site_α, site_β, ϵ):
model = smp.Model()
# Population priors - normals for population means and half-Cauchy for population stds
model.add(smp.normal(μ_α, sig=500),
smp.normal(μ_β, sig=500),
smp.half_cauchy(σ_α, beta=5),
smp.half_cauchy(σ_β, beta=0.5))
# Priors for site coefficients, sampled from population distributions
model.add(smp.normal(site_α, mu=μ_α, sig=σ_α),
smp.normal(site_β, mu=μ_β, sig=σ_β))
# Prior for likelihood uncertainty
model.add(smp.half_normal(ϵ))
# Our estimate for abalone length, α + βx
length_est = site_α[sites] + site_β[sites]*modes
# Add the log-likelihood
model.add(smp.normal(lengths, mu=length_est, sig=ϵ))
return model()
# -
model = HLM(data=data)
start = {'μ_α': 201., 'μ_β': 5., 'σ_α': 1., 'σ_β': 1.,
'site_α': np.ones(len(site_map))*201,
'site_β': np.zeros(len(site_map)),
'ϵ': 1.}
model.logp_(*start.values())
# +
start = {'μ_α': 201., 'μ_β': 5., 'σ_α': 1., 'σ_β': 1.,
'site_α': np.ones(len(site_map))*201,
'site_β': np.zeros(len(site_map)),
'ϵ': 1.}
# Using NUTS is slower per sample, but more likely to give good samples (and converge)
sampler = smp.NUTS(logp, start)
chain = sampler(1100, burn=100, thin=2)
# -
# There are some checks for convergence you can do, but they aren't implemented yet. Instead, we can visually inspect the chain. In general, the samples should be stable, the first half should vary around the same point as the second half.
fig, ax = plt.subplots()
ax.plot(chain.site_α);
fig.savefig('/Users/mat/Desktop/chains.png', dpi=150)
chain.site_α.T.shape
fig, ax = plt.subplots(figsize=(16,9))
for each in chain.site_α.T:
ax.hist(each, range=(185, 210), bins=60, alpha=0.5)
ax.set_xticklabels('')
ax.set_yticklabels('');
fig.savefig('/Users/mat/Desktop/posteriors.png', dpi=300)
# With the posterior distribution, we can look at many different results. Here I'll make a function that plots the means and 95% credible regions (range that contains central 95% of the probability) for the coefficients $\alpha_s$ and $\beta_s$.
def coeff_plot(coeff, ax=None):
if ax is None:
fig, ax = plt.subplots(figsize=(3,5))
CRs = np.percentile(coeff, [2.5, 97.5], axis=0)
means = coeff.mean(axis=0)
ax.errorbar(means, np.arange(len(means)), xerr=np.abs(means - CRs), fmt='o')
ax.set_yticks(np.arange(len(site_map)))
ax.set_yticklabels(site_map.keys())
ax.set_ylabel('Site')
ax.grid(True, axis='x', color="#CCCCCC")
ax.tick_params(axis='both', length=0)
for each in ['top', 'right', 'left', 'bottom']:
ax.spines[each].set_visible(False)
return ax
# Now we can look at how abalone lengths vary between sites for the rock-picking method ($\alpha_s$).
ax = coeff_plot(chain.site_α)
ax.set_xlim(175, 225)
ax.set_xlabel('Abalone Length (mm)');
# Here I'm plotting the mean and 95% credible regions (CR) of $\alpha$ for each site. This coefficient measures the average length of rock-picked abalones. We can see that the average abalone length varies quite a bit between sites. The CRs give a measure of the uncertainty in $\alpha$, wider CRs tend to result from less data at those sites.
#
# Now, let's see how the abalone lengths vary between harvesting methods (the difference for diving is given by $\beta_s$).
ax = coeff_plot(chain.site_β)
#ax.set_xticks([-5, 0, 5, 10, 15])
ax.set_xlabel('Mode effect (mm)');
# Here I'm plotting the mean and 95% credible regions (CR) of $\beta$ for each site. This coefficient measures the difference in length of dive picked abalones compared to rock picked abalones. Most of the $\beta$ coefficients are above zero which indicates that abalones harvested via diving are larger than ones picked from the shore. For most of the sites, diving results in 5 mm longer abalone, while at site 72, the difference is around 12 mm. Again, wider CRs mean there is less data leading to greater uncertainty.
#
# Next, I'll overlay the model on top of the data and make sure it looks right. We'll also see that some sites don't have data for both harvesting modes but our model still works because it's hierarchical. That is, we can get a posterior distribution for the coefficient from the population distribution even though the actual data is missing.
def model_plot(data, chain, site, ax=None, n_samples=20):
if ax is None:
fig, ax = plt.subplots(figsize=(4,6))
site = site_map[site]
xs = np.linspace(-1, 3)
for ii, (mode, m_data) in enumerate(data[data['site'] == site].groupby('mode')):
a = chain.site_α[:, site]
b = chain.site_β[:, site]
# now sample from the posterior...
idxs = np.random.choice(np.arange(len(a)), size=n_samples, replace=False)
# Draw light lines sampled from the posterior
for idx in idxs:
ax.plot(xs, a[idx] + b[idx]*xs, color='#E74C3C', alpha=0.05)
# Draw the line from the posterior means
ax.plot(xs, a.mean() + b.mean()*xs, color='#E74C3C')
# Plot actual data points with a bit of noise for visibility
mode_label = {0: 'Rock-picking', 1: 'Diving'}
ax.scatter(ii + np.random.randn(len(m_data))*0.04,
m_data['full lengths'], edgecolors='none',
alpha=0.8, marker='.', label=mode_label[mode])
ax.set_xlim(-0.5, 1.5)
ax.set_xticks([0, 1])
ax.set_xticklabels('')
ax.set_ylim(150, 250)
ax.grid(True, axis='y', color="#CCCCCC")
ax.tick_params(axis='both', length=0)
for each in ['top', 'right', 'left', 'bottom']:
ax.spines[each].set_visible(False)
return ax
# +
fig, axes = plt.subplots(figsize=(10, 5), ncols=4, sharey=True)
for ax, site in zip(axes, [5, 52, 72, 162]):
ax = model_plot(data, chain, site, ax=ax, n_samples=30)
ax.set_title(site)
first_ax = axes[0]
first_ax.legend(framealpha=1, edgecolor='none')
first_ax.set_ylabel('Abalone length (mm)');
# -
# For site 5, there are few data points for the diving method so there is a lot of uncertainty in the prediction. The prediction is also pulled lower than the data by the population distribution. Similarly, for site 52 there is no diving data, but we still get a (very uncertain) prediction because it's using the population information.
#
# Finally, we can look at the harvesting mode effect for the population. Here I'm going to print out a few statistics for $\mu_{\beta}$.
fig, ax = plt.subplots()
ax.hist(chain.μ_β, bins=30);
b_mean = chain.μ_β.mean()
b_CRs = np.percentile(chain.μ_β, [2.5, 97.5])
p_gt_0 = (chain.μ_β > 0).mean()
print(
"""Mean: {:.3f}
95% CR: [{:.3f}, {:.3f}]
P(mu_b) > 0: {:.3f}
""".format(b_mean, b_CRs[0], b_CRs[1], p_gt_0))
# We can also look at the population distribution for $\beta_s$ by sampling from a normal distribution with mean and variance sampled from $\mu_\beta$ and $\sigma_\beta$.
#
# $$
# \beta_s \sim \mathcal{N}\left(\mu_{\beta}, \sigma_{\beta}^2\right)
# $$
import scipy.stats as stats
samples = stats.norm.rvs(loc=chain.μ_β, scale=chain.σ_β)
plt.hist(samples, bins=30);
plt.xlabel('Dive harvesting effect (mm)')
# It's apparent that dive harvested abalone are roughly 5 mm longer than rock-picked abalone. Maybe this is a bias of the divers to pick larger abalone. Or, it's possible that abalone that stay in the water grow larger.
| examples/Abalone Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "Twitter Consume Reporter Argentina"
# > "Ejemplo de recolección de información desde cammesa y su publicacion en twitter."
# - toc: false
# - branch: master
# - badges: true
# - hide_binder_badge: true
# - comments: false
# ## Argentina x Provincia
#
# En esta seccion mostramos la Demanda (MW) para una hora especifica de las distintas provincias de la Republica argentina.
# La jupyter notebook original de procesamiento se puede encontrar en [src](https://github.com/felixlapalma/tweet_reporter/blob/master/consume_reporter_argentina.ipynb)
# +
#hide
import os,sys,glob,shutil
import zipfile
from tweet_informer_lib import wget,pd,datetime
from tweet_informer_lib import cammesa_consume_reader
from tweet_informer_lib import json,tweepy,reduce,plt,gpd,unidecode,make_cammesa_url_v2
from IPython.display import display
#
jnb=True # To display html
local_config=True # to use local credentials files (True) else sys.args input
# -
# ### Pre-Procesamiento
#
# Las url de cammesa dependen de la provincia o sector a bajar, eso lo gestionamos via un diccionario
#hide
pd_cfg=pd.read_csv('cfg/csv_cfg_provincias.csv',index_col=0)
prov_dict=pd_cfg.T.to_dict()
#hide
with zipfile.ZipFile('cfg/provincia_geojson.zip', 'r') as zip_ref:
zip_ref.extractall('cfg')
df_prov=gpd.read_file('cfg/provincia.geojson')
df_prov.NAM=df_prov.NAM.apply(unidecode.unidecode)
os.remove('cfg/provincia.geojson')
# +
#hide_input
url_dict={}
for opt in prov_dict:
url_dict.update({opt:{}})
provincia,url_case=prov_dict[opt]['Total'],prov_dict[opt]['url']
make_cammesa_url_v2('provincia',provincia,url_dict[opt],url_case)
url_dict
# -
#hide
## make some temporal dirs
tmp='tmp_mapa'
for folder in [tmp]:
os.makedirs(folder,exist_ok=True)
# descargamos los archivos...
# +
#hide_input
csv_dict={}
req_time=datetime.datetime.now().strftime('%Y%m%d%H%M%S')
for p in url_dict:
csv_dict.update({p:{}})
for key in url_dict[p]:
filename=os.path.join(tmp,p+'_'+key+'.csv')
filesaved = wget.download(url_dict[p][key],out=filename)
csv_dict[p].update({key:filesaved})
csv_dict
# -
#hide
csv_=glob.glob(os.path.join(tmp,'*.csv'))
if len(csv_)>0:
pass
else:
shutil.rmtree(tmp)
sys.exit('No CSV files')
# ### Dataframe procesado
#hide
aux=[]
for p in csv_dict:
if p in ['Buenos Aires','SADI','GBA']:
drop=[1,2,3,4]
else:
drop=[1,2]
df=cammesa_consume_reader(csv_dict[p]['provincia'],drop)
df.columns=[p]
aux.append(df)
df=reduce(lambda x, y: pd.merge(x, y, left_index=True,right_index=True), aux)
df[df < 0] = 0
df_notna=df.dropna()
#
df_notna=df_notna.dropna()
##
series=df_notna.iloc[-1]
series_time=series.name.strftime('%d-%m-%Y %H:%M:%S')
df_new=pd.DataFrame(df_notna.iloc[-1]).reset_index()
df_new.columns=['NAM','Consumo']
df_new['Consumo_Porc']=df_new['Consumo']/df_notna['SADI'].iloc[-1]*100
df_new['NAM']=df_new['NAM'].apply(lambda x: x if x!='GBA' else 'Ciudad Autonoma de Buenos Aires')
#hide
df_filt=df_prov.merge(df_new,on='NAM')
if len(df_filt)>1:
pass
else:
shutil.rmtree(tmp)
sys.exit('Short Dataframe')
# Observamos el data frame procesado
#hide_input
df_filt
# y generamos el grafico correspondiente que sera utilizado en el tweet
#hide_input
try:
fig=plt.figure(figsize=(15, 18))
ax=fig.add_subplot(111)
df_filt.plot(column='Consumo',cmap='Oranges',ax=ax)
for p in df_filt.NAM:
x=df_filt[df_filt.NAM==p].geometry.representative_point().x
y=df_filt[df_filt.NAM==p].geometry.representative_point().y
MW_porc=df_filt[df_filt.NAM==p]['Consumo_Porc'].values
MW=df_filt[df_filt.NAM==p]['Consumo'].values
if MW_porc[0]>1/22*100:
c='red'
else:
c='black'
if p=='Ciudad Autonoma de Buenos Aires':
fmt='GBA: {:.0f}\n({:.1f}%)'
else:
fmt='{:.0f}\n({:.1f}%)'
ax.text(x,y,fmt.format(MW[0],MW_porc[0]),horizontalalignment='center',verticalalignment='bottom',bbox=dict(facecolor='white', alpha=0.7),fontsize=14,color=c)
ax.set_title('Demanda Provincias [MW] \n(% Total Pais) \n'+series_time,fontsize=20)
# save fig
plt.axis('off')
figName=os.path.join(tmp,'mapa_consumo.png')
fig.savefig(figName,transparent=False)
except:
shutil.rmtree(tmp)
print('Plot ERROR')
sys.exit('Plot ERROR')
# ### Twitter Side
#
# Cargamos la configuración y generamos el tweet correspondiente y mostramos un ejemplo (puede no corresponder a la imagen generada).
# +
#collapse
try:
if local_config:
## .tweepy.json not available
config_file = 'cfg/.tweepy.json'
with open(config_file) as fh:
config = json.load(fh)
else:
# use sys.arg (## .tweepy.json not available)
config={'consumer_key':sys.argv[1],\
'consumer_secret':sys.argv[2],\
'access_token': sys.argv[3],\
'access_token_secret':sys.argv[4]}
auth = tweepy.OAuthHandler(config['consumer_key'], config['consumer_secret'])
auth.set_access_token(config['access_token'], config['access_token_secret'])
twitter = tweepy.API(auth)
tweet ='Demanda Provincias [MW] y % Total Pais'
image_path =figName
# to attach the media file
status = twitter.update_with_media(image_path, tweet)
except:
shutil.rmtree(tmp)
sys.exit('Failed to TWEET')
## src: https://github.com/jupyter/notebook/issues/2790
class Tweet(object):
def __init__(self, embed_str=None):
self.embed_str = embed_str
def _repr_html_(self):
return self.embed_str
# -
#hide_input
if jnb:
s = ("""<blockquote class="twitter-tweet"><p lang="es" dir="ltr">Demanda Provincias [MW] y % Total Pais <a href="https://t.co/3ZvRQTHfKS">pic.twitter.com/3ZvRQTHfKS</a></p>— misc reporter (@ReporterMisc) <a href="https://twitter.com/ReporterMisc/status/1249651529742680064?ref_src=twsrc%5Etfw">April 13, 2020</a></blockquote> <script async src="https://platform.twitter.com/widgets.js" charset="utf-8"></script> """)
## src: https://github.com/jupyter/notebook/issues/2790
display(Tweet(s))
#hide
shutil.rmtree(tmp)
# ## Provincia
#
# En esta seccion mostramos la Demanda (MW) para una serie de horas para una provincia en particular.
# La jupyter notebook original de procesamiento se puede encontrar en [src](https://github.com/felixlapalma/tweet_reporter/blob/master/consume_reporter_provincias.ipynb)
# +
#hide
import os,sys,glob,shutil
import zipfile
from tweet_informer_lib import wget,pd,datetime
from tweet_informer_lib import cammesa_consume_reader
from tweet_informer_lib import json,tweepy,reduce,plt,gpd,unidecode,make_cammesa_url_v2
from tweet_informer_lib import make_cammesa_url,make_plt_provincia_capital
from IPython.display import display
#
jnb=True # To display html
local_config=True # to use local credentials files (True) else sys.args input
# -
# ### Pre-Procesamiento
#
# Las url de cammesa dependen de la provincia o sector a bajar, eso lo gestionamos via un diccionario. Para el presente caso elegimos **Cordoba**.
#hide
pd_cfg=pd.read_csv('cfg/csv_cfg_provincias.csv',index_col=0)
prov_dict=pd_cfg.T.to_dict()
#hide_input
opt='Cordoba'
print('Provincia-Sector: {}'.format(opt))
#hide_input
url_dict={}
if opt in prov_dict:
provincia,capital=prov_dict[opt]['Total'],prov_dict[opt]['Capital']
make_cammesa_url('provincia',provincia,url_dict)
make_cammesa_url('capital',capital,url_dict)
else:
'Opciones validas: {}'.format(list(prov_dict.keys()))
sys.exit()
url_dict
#hide
tmp='tmp'+opt
for folder in [tmp]:
os.makedirs(folder,exist_ok=True)
# descargamos los archivos ...
#hide_input
csv_dict={}
req_time=datetime.datetime.now().strftime('%Y%m%d%H%M%S')
for key in url_dict:
filename=os.path.join(tmp,key+'.csv')
filesaved = wget.download(url_dict[key],out=filename)
csv_dict.update({key:filesaved})
csv_dict
#hide
csv_=glob.glob(os.path.join(tmp,'*.csv'))
if len(csv_)==2:
pass
else:
shutil.rmtree(tmp)
sys.exit('No CSV files')
# ### Dataframe procesado
#hide
pd_ciudad=pd.read_csv(csv_dict['capital'],sep=';',decimal=',',index_col=[0],parse_dates=[0])
pd_provincia=pd.read_csv(csv_dict['provincia'],sep=';',decimal=',',index_col=[0],parse_dates=[0])
#
pd_merge=pd.merge(pd_provincia,pd_ciudad,left_index=True,right_index=True,suffixes=('_prov','_capital'))
pd_merge[pd_merge < 0] = 0
pd_merge['provincia_sin_capital']=pd_merge['Dem Hoy_prov']-pd_merge['Dem Hoy_capital']
#
pd_merge_not_na=pd_merge.dropna()
#
if len(pd_merge_not_na)>10:
pass
else:
shutil.rmtree(tmp)
sys.exit('Short Dataframe')
# Observamos el data frame procesado
#hide_input
pd_merge_not_na.head(10)
# y generamos el grafico correspondiente que sera utilizado en el tweet
#hide_input
try:
fig,tweet_text=make_plt_provincia_capital(pd_merge_not_na,opt,figsize=(16,12))
# save fig
figName=os.path.join(tmp,'consumo.png')
fig.savefig(figName,transparent=False)
except:
shutil.rmtree(tmp)
print('Plot ERROR')
sys.exit('Plot ERROR')
# ### Twitter Side
#
# Cargamos la configuración y generamos el tweet correspondiente y mostramos un ejemplo (puede no corresponder a la imagen generada).
# +
#collapse
try:
if local_config:
## .tweepy.json not available
config_file = 'cfg/.tweepy.json'
with open(config_file) as fh:
config = json.load(fh)
else:
# use sys.arg (## .tweepy.json not available)
config={'consumer_key':sys.argv[1],\
'consumer_secret':sys.argv[2],\
'access_token': sys.argv[3],\
'access_token_secret':sys.argv[4]}
auth = tweepy.OAuthHandler(config['consumer_key'], config['consumer_secret'])
auth.set_access_token(config['access_token'], config['access_token_secret'])
twitter = tweepy.API(auth)
tweet =tweet_text
image_path =figName
# to attach the media file
status = twitter.update_with_media(image_path, tweet)
except:
shutil.rmtree(tmp)
sys.exit('Failed to TWEET')
## src: https://github.com/jupyter/notebook/issues/2790
class Tweet(object):
def __init__(self, embed_str=None):
self.embed_str = embed_str
def _repr_html_(self):
return self.embed_str
# -
#hide_input
if jnb:
s = ("""<blockquote class="twitter-tweet"><p lang="ca" dir="ltr">Cordoba - Demanda (MW) - 13-04-2020 07:30 - Total Provincia: 849.7 / Provincia sin Capital: 566.5 / Capital: 283.3 / <a href="https://t.co/HW1vowvMZ2">pic.twitter.com/HW1vowvMZ2</a></p>— misc reporter (@ReporterMisc) <a href="https://twitter.com/ReporterMisc/status/1249653975974121477?ref_src=twsrc%5Etfw">April 13, 2020</a></blockquote> <script async src="https://platform.twitter.com/widgets.js" charset="utf-8"></script> """)
## src: https://github.com/jupyter/notebook/issues/2790
display(Tweet(s))
#hide
shutil.rmtree(tmp)
| _notebooks/2019-07-23-consume_reporter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Intro to Pandas
# Pandas is a Python package for data analysis and exposes two new
# data structures: Dataframes and Series.
#
# - [Dataframes](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.html) store tabular data consisting of rows and columns.
# - [Series](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.html) are similar to Python's built-in list or set data types.
#
# In this notebook, we will explore the data structures that Pandas
# provides, and learn how to interact with them.
# ### 1. Importing Pandas
#
# To import an external Python library such as Pandas, use Python's
# import function. To save yourself some typing later on, you can
# give the library you import an alias. Here, we are importing Pandas
# and giving it an alias of `pd`.
import pandas as pd
# ### 2. Creating A Dataframe and Basic Exploration
# We will load a CSV file as a dataframe using Panda's `read_csv`
# method. This will allow us to use Pandas' dataframe functions to
# explore the data in the CSV.
df = pd.read_csv("../data/loans_full.zip",index_col=0)
# Once we have loaded the CSV as a dataframe, we can start to explore
# the data. Here are a few useful methods:
# - .head(): returns first 5 rows of the DataFrame
# - .tail(): returns last 5 rows of the DataFrame
# - .shape: returns tuple with first element indicating the number of rows and the second element indicating the number of columns
# - .columns: returns list of all columns in DataFrame
# - .index: returns DataFrame indices
# - .dtypes: returns Series explaining the datatype of each column
df.dtypes
# To get some basic stats of the columns you can either use .describe() for discrete data or .value_counts for categroical data
df.describe()
df['activity'].value_counts()
# ### 3. Selecting Data - Part 1
# To examine a specfic column of the DataFrame:
df['activity'].head()
df[['activity','basket_amount']].tail()
#
#
# To examine specific rows and columns of a Dataframe, Pandas provides
# the `iloc` and `loc` methods to do so. `iloc` is used when you want to specify a list or range of indices, and `.loc` is used when you want to specify a list or range of labels.
#
# For both of these methods you need to specify two elements, with the first element indicating the rows that you want to select and the second element indicating the columns that you want to select.
# Get rows 1 through 3 and columns 0 through 5.
df.iloc[1:3,:5]
# Get rows with index values of 2-4 and the columns basket_amount and activity
df.loc[2:4, ["basket_amount", "activity"]]
# To see all the rows and columns:
df.iloc[:,:]
# You can also store a slice of the dataframe as a new dataframe!
titles_df = df.iloc[:,2]
titles_df.head()
# ### 4. Select subets of the DataFrame
#
# A powerful feature of DataFrames is that you can view a subset of the DataFrame based on the values of the columns or rows. For example, lets say you only wanted to view loans with a status of "expired"
df[df['status']=='expired']
# To view all loans with a status of "expired" `or` "fundraising":
df[(df['status']=='expired')|(df['status']=='fundraising')]
# Select loans that have expired and with loan amounts greater than 1000
df[(df['status']=='expired')&(df['loan_amount']>1000)]
# ## Great Resources for further information:
#
# - [10 minute introduction to pandas](http://pandas.pydata.org/pandas-docs/stable/10min.html)
# - [Pandas in ipython notebooks](http://nbviewer.jupyter.org/github/jvns/pandas-cookbook/blob/master/cookbook/A%20quick%20tour%20of%20IPython%20Notebook.ipynb)
# !ls
| module_0_introduction/intro_to_pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import timeit
import tqdm
import pprint
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
plt.rcParams['text.usetex'] = True
plt.rcParams['axes.labelsize'] = 16
plt.rcParams['font.size'] = 16
plt.rcParams['legend.fontsize'] = 'large'
import pyquaternion # pip install pyquaternion --user
import quaternion # pip install numpy_quaternion --user
import rowan # pip install rowan --user
# +
def arr_to_pyquat(arr):
if len(arr.shape) > 1:
pq_arr = np.empty(arr.shape[:-1], dtype='object')
for i, x in enumerate(arr):
pq_arr[i] = pyquaternion.Quaternion(x)
else:
pq_arr = np.array([pyquaternion.Quaternion(arr)])
return pq_arr
def arr_to_npquat(arr):
return quaternion.as_quat_array(arr)
pyquat_times = {}
quat_times = {}
rowan_times = {}
Ns = [10, 100000]
num = 10
pqlim = 1e8
# -
pyquat_times['Multiply'] = []
quat_times['Multiply'] = []
rowan_times['Multiply'] = []
for N in tqdm.tqdm_notebook(Ns):
x = rowan.random.rand(N)
y = rowan.random.rand(N)
if N < pqlim:
pyquat_times['Multiply'].append(
timeit.timeit(
"x*y",
setup="from __main__ import x, y, arr_to_pyquat; x = arr_to_pyquat(x); y = arr_to_pyquat(y)",
number = num
)
)
quat_times['Multiply'].append(
timeit.timeit(
"x*y",
setup="from __main__ import x, y, arr_to_npquat; x = arr_to_npquat(x); y = arr_to_npquat(y)",
number = num
)
)
rowan_times['Multiply'].append(
timeit.timeit(
"rowan.multiply(x, y)",
setup="from __main__ import x, y, rowan",
number = num
)
)
pyquat_times['Exponential'] = []
quat_times['Exponential'] = []
rowan_times['Exponential'] = []
for N in tqdm.tqdm_notebook(Ns):
x = rowan.random.rand(N)
if N < pqlim:
pyquat_times['Exponential'].append(
timeit.timeit(
"for i in range(len(x)): pyquaternion.Quaternion.exp(x[i])",
setup="from __main__ import x, pyquaternion, arr_to_pyquat; x = arr_to_pyquat(x);",
number = num
)
)
quat_times['Exponential'].append(
timeit.timeit(
"np.exp(x)",
setup="from __main__ import x, arr_to_npquat, np; x = arr_to_npquat(x);",
number = num
)
)
rowan_times['Exponential'].append(
timeit.timeit(
"rowan.exp(x)",
setup="from __main__ import x, rowan",
number = num
)
)
pyquat_times['Conjugate'] = []
quat_times['Conjugate'] = []
rowan_times['Conjugate'] = []
for N in tqdm.tqdm_notebook(Ns):
x = rowan.random.rand(N)
if N < pqlim:
pyquat_times['Conjugate'].append(
timeit.timeit(
"for i in range(len(x)): x.conjugate",
setup="from __main__ import x, arr_to_pyquat; x = arr_to_pyquat(x);",
number = num
)
)
quat_times['Conjugate'].append(
timeit.timeit(
"x.conjugate()",
setup="from __main__ import x, arr_to_npquat; x = arr_to_npquat(x);",
number = num
)
)
rowan_times['Conjugate'].append(
timeit.timeit(
"rowan.conjugate(x)",
setup="from __main__ import x, rowan",
number = num
)
)
pyquat_times['Norm'] = []
quat_times['Norm'] = []
rowan_times['Norm'] = []
for N in tqdm.tqdm_notebook(Ns):
x = rowan.random.rand(N)
if N < pqlim:
pyquat_times['Norm'].append(
timeit.timeit(
"for i in range(len(x)): x[i].norm",
setup="from __main__ import x, arr_to_pyquat; x = arr_to_pyquat(x);",
number = num
)
)
quat_times['Norm'].append(
timeit.timeit(
"np.abs(x)",
setup="from __main__ import x, np, arr_to_npquat; x = arr_to_npquat(x);",
number = num
)
)
rowan_times['Norm'].append(
timeit.timeit(
"rowan.norm(x)",
setup="from __main__ import x, rowan",
number = num
)
)
pyquat_times['To Matrix'] = []
quat_times['To Matrix'] = []
rowan_times['To Matrix'] = []
for N in tqdm.tqdm_notebook(Ns):
x = rowan.random.rand(N)
if N < pqlim:
pyquat_times['To Matrix'].append(
timeit.timeit(
"for i in range(len(x)): x[i].rotation_matrix",
setup="from __main__ import x, arr_to_pyquat; x = arr_to_pyquat(x);",
number = num
)
)
quat_times['To Matrix'].append(
timeit.timeit(
"quaternion.as_rotation_matrix(x)",
setup="from __main__ import x, quaternion, arr_to_npquat; x = arr_to_npquat(x);",
number = num
)
)
rowan_times['To Matrix'].append(
timeit.timeit(
"rowan.to_matrix(x)",
setup="from __main__ import x, rowan",
number = num
)
)
pyquat_times['N'] = list(np.array(Ns)[np.array(Ns) < pqlim])
quat_times['N'] = Ns
rowan_times['N'] = Ns
df_pq = pd.DataFrame(pyquat_times).melt(id_vars="N", var_name="operation", value_name="pyquaternion")
df_nq = pd.DataFrame(quat_times).melt(id_vars="N", var_name="operation", value_name="npquaternion")
df_r = pd.DataFrame(rowan_times).melt(id_vars="N", var_name="operation", value_name="rowan")
df = df_r.merge(df_nq, on =["N", "operation"])
df = df.merge(df_pq, on =["N", "operation"], how = "left")
df.fillna(0, inplace=True)
df['pyquaternion'] /= df['N']
df['pyquaternion'] *= 1e6
df['npquaternion'] /= df['N']
df['npquaternion'] *= 1e6
df['rowan'] /= df['N']
df['rowan'] *= 1e6
view = df.groupby(["N", "operation"]).mean()
view['rowan vs. npq'] = view['rowan']/view['npquaternion']
view['pyq vs. rowan'] = view['pyquaternion']/view['rowan']
view
# + code_folding=[]
cols = list(col['color'] for col in plt.rcParams['axes.prop_cycle'])
fig, axes = plt.subplots(2, 1, figsize=(8, 15))
ax = df[df['N'] == Ns[0]].drop(['N'], axis=1).groupby(
["operation"]).mean().plot.barh(ax=axes[0], logx=True, color = cols[0:3],
title="$\log_{{\,10}}(N) = {}$".format(int(np.log10(Ns[0]))),
xlim=(10**-2, 10**3), legend=False)
ax = df[df['N'] == Ns[1]].drop(['N'], axis=1).groupby(
["operation"]).mean().plot.barh(ax=axes[1], logx=True, color = cols[0:3],
title="$\log_{{\,10}}(N) = {}$".format(int(np.log10(Ns[1]))),
xlim=(10**-2, 10**3), legend=False)
ax.set_xlabel("$\log_{\,10}(sec)$", fontsize=18)
for ax in axes.flatten():
ax.set_ylabel("")
ax.legend(loc='best', fontsize=15)
ax.title.set_fontsize(20)
#ax.tick_params(axis='both', which='major', labelsize=16)
fig.subplots_adjust(left=0.2)
plt.show()
fig.savefig("Performance.pdf")
| benchmarks/Benchmarks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # BigQuery ML models with feature engineering
#
# In this notebook, we will use BigQuery ML to build more sophisticated models for taxifare prediction.
#
# This is a continuation of our [first models](../02_bqml/first_model.ipynb)
# if you have not already done so ...
# !bq mk serverlessml
# ## Model 4: With some transformations
#
# BigQuery ML automatically scales the inputs. so we don't need to do scaling, but human insight can help.
#
# Since we we'll repeat this quite a bit, let's make a dataset with 1 million rows.
# +
# %%bigquery
CREATE OR REPLACE TABLE serverlessml.feateng_training_data AS
SELECT
(tolls_amount + fare_amount) AS fare_amount,
pickup_datetime,
pickup_longitude AS pickuplon,
pickup_latitude AS pickuplat,
dropoff_longitude AS dropofflon,
dropoff_latitude AS dropofflat,
passenger_count*1.0 AS passengers
FROM `nyc-tlc.yellow.trips`
WHERE MOD(ABS(FARM_FINGERPRINT(CAST(pickup_datetime AS STRING))), 1000) = 1
AND
trip_distance > 0
AND fare_amount >= 2.5
AND pickup_longitude > -78
AND pickup_longitude < -70
AND dropoff_longitude > -78
AND dropoff_longitude < -70
AND pickup_latitude > 37
AND pickup_latitude < 45
AND dropoff_latitude > 37
AND dropoff_latitude < 45
AND passenger_count > 0
# +
# %%bigquery
CREATE OR REPLACE MODEL serverlessml.model4_feateng
TRANSFORM(
* EXCEPT(pickup_datetime)
, ST_Distance(ST_GeogPoint(pickuplon, pickuplat), ST_GeogPoint(dropofflon, dropofflat)) AS euclidean
, CAST(EXTRACT(DAYOFWEEK FROM pickup_datetime) AS STRING) AS dayofweek
, CAST(EXTRACT(HOUR FROM pickup_datetime) AS STRING) AS hourofday
)
OPTIONS(input_label_cols=['fare_amount'], model_type='linear_reg')
AS
SELECT * FROM serverlessml.feateng_training_data
# -
# Once the training is done, visit the [BigQuery Cloud Console](https://console.cloud.google.com/bigquery) and look at the model that has been trained. Then, come back to this notebook.
# Note that BigQuery automatically split the data we gave it, and trained on only a part of the data and used the rest for evaluation. We can look at eval statistics on that held-out data:
# %%bigquery
SELECT *, SQRT(loss) AS rmse FROM ML.TRAINING_INFO(MODEL serverlessml.model4_feateng)
# %%bigquery
SELECT SQRT(mean_squared_error) AS rmse FROM ML.EVALUATE(MODEL serverlessml.model4_feateng)
# Yippee! We're now below our target of 6 dollars in RMSE.
# We are now beating our goals, and with just a linear model. This is how the prediction query would look:
# %%bigquery
SELECT * FROM ML.PREDICT(MODEL serverlessml.model4_feateng, (
SELECT
-73.982683 AS pickuplon,
40.742104 AS pickuplat,
-73.983766 AS dropofflon,
40.755174 AS dropofflat,
3.0 AS passengers,
TIMESTAMP('2019-06-03 04:21:29.769443 UTC') AS pickup_datetime
))
# ## Let's do a feature cross of the day-hour combination instead of using them raw
# +
# %%bigquery
CREATE OR REPLACE MODEL serverlessml.model5_featcross
TRANSFORM(
* EXCEPT(pickup_datetime)
, ST_Distance(ST_GeogPoint(pickuplon, pickuplat), ST_GeogPoint(dropofflon, dropofflat)) AS euclidean
, CONCAT(CAST(EXTRACT(DAYOFWEEK FROM pickup_datetime) AS STRING),
CAST(EXTRACT(HOUR FROM pickup_datetime) AS STRING)) AS day_hr
)
OPTIONS(input_label_cols=['fare_amount'], model_type='linear_reg')
AS
SELECT * FROM serverlessml.feateng_training_data
# -
# %%bigquery
SELECT *, SQRT(loss) AS rmse FROM ML.TRAINING_INFO(MODEL serverlessml.model5_featcross)
# %%bigquery
SELECT SQRT(mean_squared_error) AS rmse FROM ML.EVALUATE(MODEL serverlessml.model5_featcross)
# Sometimes (not the case above), the training RMSE is quite reasonable, but the evaluation RMSE is terrible. This is an indication of overfitting.
# When we do feature crosses, we run into the risk of overfitting (for example, when a particular day-hour combo doesn't have enough taxirides).
#
# ## Reducing overfitting
#
# Let's add L2 regularization.
# +
# %%bigquery
CREATE OR REPLACE MODEL serverlessml.model6_featcross_l2
TRANSFORM(
* EXCEPT(pickup_datetime)
, ST_Distance(ST_GeogPoint(pickuplon, pickuplat), ST_GeogPoint(dropofflon, dropofflat)) AS euclidean
, CONCAT(CAST(EXTRACT(DAYOFWEEK FROM pickup_datetime) AS STRING),
CAST(EXTRACT(HOUR FROM pickup_datetime) AS STRING)) AS day_hr
)
OPTIONS(input_label_cols=['fare_amount'], model_type='linear_reg', l2_reg=0.1)
AS
SELECT * FROM serverlessml.feateng_training_data
# -
# %%bigquery
SELECT SQRT(mean_squared_error) AS rmse FROM ML.EVALUATE(MODEL serverlessml.model6_featcross_l2)
# These sorts of experiment would have taken days to do otherwise. We did it in minutes, thanks to BigQuery ML! The advantage of doing all this in the TRANSFORM is the client code doing the PREDICT doesn't change. Our model improvement is transparent to client code.
# %%bigquery
SELECT * FROM ML.PREDICT(MODEL serverlessml.model6_featcross_l2, (
SELECT
-73.982683 AS pickuplon,
40.742104 AS pickuplat,
-73.983766 AS dropofflon,
40.755174 AS dropofflat,
3.0 AS passengers,
TIMESTAMP('2019-06-03 04:21:29.769443 UTC') AS pickup_datetime
))
# ## Let's try feature crossing the locations too
#
# Because the lat and lon by themselves don't have meaning, but only in conjunction, it may be useful to treat the fields as a pair instead of just using them as numeric values. However, lat and lon are continuous numbers, so we have to discretize them first. That's what SnapToGrid does.
# +
# %%bigquery
CREATE OR REPLACE MODEL serverlessml.model7_geo
TRANSFORM(
fare_amount
, ST_Distance(ST_GeogPoint(pickuplon, pickuplat), ST_GeogPoint(dropofflon, dropofflat)) AS euclidean
, CONCAT(CAST(EXTRACT(DAYOFWEEK FROM pickup_datetime) AS STRING),
CAST(EXTRACT(HOUR FROM pickup_datetime) AS STRING)) AS day_hr
, CONCAT(
ST_AsText(ST_SnapToGrid(ST_GeogPoint(pickuplon, pickuplat), 0.01)),
ST_AsText(ST_SnapToGrid(ST_GeogPoint(dropofflon, dropofflat), 0.01))
) AS pickup_and_dropoff
)
OPTIONS(input_label_cols=['fare_amount'], model_type='linear_reg', l2_reg=0.1)
AS
SELECT * FROM serverlessml.feateng_training_data
# -
# %%bigquery
SELECT SQRT(mean_squared_error) AS rmse FROM ML.EVALUATE(MODEL serverlessml.model7_geo)
# Yippee! We're now below our target of 6 dollars in RMSE.
# ## DNN
#
# You could, of course, train a more sophisticated model. Change "linear_reg" above to "dnn_regressor" and see if it improves things.
# +
# %%bigquery
-- This is alpha and may not work for you.
CREATE OR REPLACE MODEL serverlessml.model8_dnn
TRANSFORM(
fare_amount
, ST_Distance(ST_GeogPoint(pickuplon, pickuplat), ST_GeogPoint(dropofflon, dropofflat)) AS euclidean
, CONCAT(CAST(EXTRACT(DAYOFWEEK FROM pickup_datetime) AS STRING),
CAST(EXTRACT(HOUR FROM pickup_datetime) AS STRING)) AS day_hr
, CONCAT(
ST_AsText(ST_SnapToGrid(ST_GeogPoint(pickuplon, pickuplat), 0.01)),
ST_AsText(ST_SnapToGrid(ST_GeogPoint(dropofflon, dropofflat), 0.01))
) AS pickup_and_dropoff
)
-- at the time of writing, l2_reg wasn't supported yet.
OPTIONS(input_label_cols=['fare_amount'], model_type='dnn_regressor', hidden_units=[32, 8])
AS
SELECT * FROM serverlessml.feateng_training_data
# -
# %%bigquery
SELECT SQRT(mean_squared_error) AS rmse FROM ML.EVALUATE(MODEL serverlessml.model8_dnn)
# We really need the L2 reg (recall that we got 4.77 without the feateng). Let's do this in Keras.
# Copyright 2019 Google Inc.
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| quests/serverlessml/05_feateng/bqml.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dividir el dataset en conjunto de entrenamiento y de testing
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
data = pd.read_csv("../datasets/ads/Advertising.csv")
a = np.random.randn(len(data))
plt.hist(a)
check = (a<0.8)
training = data[check]
testing = data[~check]
len(training), len(testing)
import statsmodels.formula.api as smf
lm = smf.ols(formula="Sales~TV+Radio", data=training).fit()
lm.summary()
# Sales = 2.9336 + 0.0465 * TV + 0.1807 * Radio
# ## Validación del modelo con el conjunto de testing
sales_pred = lm.predict(testing)
sales_pred
SSD = sum((testing["Sales"]-sales_pred)**2)
SSD
RSE = np.sqrt(SSD/(len(testing)-2-1))
RSE
sales_mean = np.mean(testing["Sales"])
error = RSE/sales_mean
error
# %matplotlib inline
data.plot(kind = "scatter", x = "TV", y ="Sales")
plt.plot(pd.DataFrame(data["TV"]), sales_pred, c="red", linewidth = 2)
from IPython.display import Image
Image(filename="resources/summary-lm.png")
| notebooks/T4 - 3 - Linear Regression - Validación del modelo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # YOLO v3 Finetuning on AWS
#
# This series of notebooks demonstrates how to finetune pretrained YOLO v3 (aka YOLO3) using MXNet on AWS.
#
# **This notebook** guides you on how to deploy the YOLO3 model trained in the previous module to the SageMaker endpoint using GPU instance.
# **Follow-on** the content of the notebooks shows:
#
# * How to use MXNet YOLO3 pretrained model
# * How to use Deep SORT with MXNet YOLO3
# * How to create Ground-Truth dataset from images the model mis-detected
# * How to finetune the model using the created dataset
# * Load your finetuned model and Deploy Sagemaker-Endpoint with it using CPU instance.
# * Load your finetuned model and Deploy Sagemaker-Endpoint with it using GPU instance.
#
# ## Pre-requisites
#
# This notebook is designed to be run in Amazon SageMaker. To run it (and understand what's going on), you'll need:
#
# * Basic familiarity with Python, [MXNet](https://mxnet.apache.org/), [AWS S3](https://docs.aws.amazon.com/s3/index.html), [Amazon Sagemaker](https://aws.amazon.com/sagemaker/)
# * To create an **S3 bucket** in the same region, and ensure the SageMaker notebook's role has access to this bucket.
# * Sufficient [SageMaker quota limits](https://docs.aws.amazon.com/general/latest/gr/aws_service_limits.html#limits_sagemaker) set on your account to run GPU-accelerated spot training jobs.
#
# ## Cost and runtime
#
# Depending on your configuration, this demo may consume resources outside of the free tier but should not generally be expensive because we'll be training on a small number of images. You might wish to review the following for your region:
#
# * [Amazon SageMaker pricing](https://aws.amazon.com/sagemaker/pricing/)
#
# The standard `ml.t2.medium` instance should be sufficient to run the notebooks.
#
# We will use GPU-accelerated instance types for training and hyperparameter optimization, and use spot instances where appropriate to optimize these costs.
#
# As noted in the step-by-step guidance, you should take particular care to delete any created SageMaker real-time prediction endpoints when finishing the demo.
# # Step 0: Dependencies and configuration
#
# As usual we'll start by loading libraries, defining configuration, and connecting to the AWS SDKs:
# +
# %load_ext autoreload
# %autoreload 1
# Built-Ins:
import os
import json
from datetime import datetime
from glob import glob
from pprint import pprint
from matplotlib import pyplot as plt
from base64 import b64encode
# External Dependencies:
import boto3
import imageio
import sagemaker
import numpy as np
from sagemaker.mxnet import MXNet
from botocore.exceptions import ClientError
# -
# ## Step 1: Get best job informations
# %store -r
best_model_output_path, best_model_job_name, role_name
model_data_path = f'{best_model_output_path}/{best_model_job_name}/output/model.tar.gz'
print(model_data_path)
# ## Step 2: Create SageMaker Model
#
# Containers for SageMaker MXNet deployments provide inference requests by using default SageMaker InvokeEndpoint API calls. So you do not have to build a Docker container yourself and upload it to Amazon Elastic Container Registry (ECR). All you have to do is implement the interfaces such as `model_fn(model_dir)`, `input_fn(request_body, content_type)`, `predict_fn(input_object, model)`, `output_fn(prediction, content_type)`. See the code example below.
#
# https://sagemaker.readthedocs.io/en/stable/using_mxnet.html#serve-an-mxnet-model
# !pygmentize src/inference_gpu.py
mxnet_model = sagemaker.mxnet.model.MXNetModel(
name='yolo3-hol-deploy-gpu',
model_data=model_data_path,
role=role_name,
entry_point='inference_gpu.py',
source_dir='src',
framework_version='1.4.1',
py_version='py3',
)
# ## Step 3: Deploy Model to GPU instance
#
# For applications that need to be very responsive(e.g., 500ms), you might consider using GPU instances. For GPU acceleration, please specify the instance type as GPU instance like `'ml.p2.xlarge'`. The inference script code is the same for `inference_cpu.py` and `inference_gpu.py` except for one line such that `ctx = mx.cpu()` for CPU and `ctx = mx.gpu()` for GPU.
#
# Note that it takes about **6-10 minutes** to create an endpoint. Please do not run real-time inference until the creation of the endpoint is complete. You can check whether the endpoint is created in the SageMaker dashboard, and wait for the status to become **InService**.
#
#
# #### Additional Tip
#
# GPU instances have good inference performance, but can be costly. Elastic Inference uses CPU instances by default, and provides GPU acceleration for inference depending on the situation, so using a general-purpose compute CPU instance and Elastic Inference together can host endpoints at a much lower cost than using GPU instances. Please refer to the link below for details.
# https://docs.aws.amazon.com/en_kr/sagemaker/latest/dg/ei.html
# %%time
predictor = mxnet_model.deploy(
instance_type='ml.p2.xlarge', initial_instance_count=1,
#accelerator_type='ml.eia2.xlarge'
)
# ## Step 4: Invoke Sagemaker Endpoint
#
# Now that we have finished creating the endpoint, we will perform object detection on the test image.
# +
from gluoncv.utils import download, viz
from base64 import b64encode, b64decode
import mxnet as mx
bimage = None
download('https://sportshub.cbsistatic.com/i/r/2019/11/15/10869f78-1378-4aa5-b36b-085607ae3387/thumbnail/770x433/f3276ac966a56b7cb45987869098cddb/lionel-messi-argentina-brazil.jpg', path='soccer.jpg')
with open('soccer.jpg', 'rb') as fp:
bimage = fp.read()
s = b64encode(bimage).decode('utf-8')
# -
# %%time
res = predictor.predict({
'short': 416,
'image': s
})
print(res['shape'])
ax = viz.plot_bbox(mx.image.imresize(mx.image.imdecode(bimage), 492, 416), mx.nd.array(res['bbox']), mx.nd.array(res['score']), mx.nd.array(res['cid']), class_names=['person'])
| 05. Deploy to Sagemaker Endpoint (GPU).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # 5장 되추적
# + [markdown] slideshow={"slide_type": "slide"}
# ## 주요 내용
# + [markdown] slideshow={"slide_type": ""}
# * 1절 되추적 기법
# + [markdown] slideshow={"slide_type": ""}
# * 2절 n-퀸 문제
# + [markdown] slideshow={"slide_type": ""}
# * 5절 그래프 색칠하기
# -
# * 부록: 제네릭 프로그래밍 활용
# + [markdown] slideshow={"slide_type": "slide"}
# ## 1절 제약충족 문제와 되추적 기법
# + [markdown] slideshow={"slide_type": "slide"}
# ### 제약충족 문제(CSP, constraint-satisfaction problems)
# + [markdown] slideshow={"slide_type": ""}
# * 특정 변수에 할당할 값을 지정된 **도메인**(영역, 집합)에서 정해진 조건에 따라 선택하는 문제
# + [markdown] slideshow={"slide_type": "slide"}
# * 예제: 4-퀸 문제(체스 퀸(queen) 네 개의 위치 선정하기)
# * 변수: 네 개의 퀸
# * 즉, 1번 퀸부터 4번 퀸.
# * 도메인: {1, 2, 3, 4}
# * 즉, 1번 열부터 4번 열.
# * 조건: 두 개의 퀸이 하나의 행, 열, 또는 대각선 상에 위치하지 않음.
# + [markdown] slideshow={"slide_type": ""}
# <div align="center"><img src="./images/algo05/algo05-01a.png" width="150"/></div>
# + [markdown] slideshow={"slide_type": "slide"}
# ### 되추적 기법(백트래킹, backtracking)
# + [markdown] slideshow={"slide_type": ""}
# * 제약충족 문제를 해결하는 일반적인 기법
# -
# * 문제에 따라 다른 제약충족 조건만 다를 뿐 문제해결을 위한 알고리즘은 동일함.
# * 여기서는 두 개의 문제를 이용하여 되추적 기법의 활용법을 설명함.
# + [markdown] slideshow={"slide_type": "slide"}
# ### 주요 기초개념
# -
# * 깊이우선 탐색
# * 상태 공간 나무
# * 마디의 유망성
# * 가지치기
# + [markdown] slideshow={"slide_type": "slide"}
# #### 깊이우선 탐색
# + [markdown] slideshow={"slide_type": "slide"}
# * DFS(depth-first-search): 뿌리 지정 나무(rooted tree)를 대상으로 하는 탐색기법.
# + [markdown] slideshow={"slide_type": ""}
# * 왼편으로 끝(잎마디)까지 탐색한 후에 오른편 형제자매 마디로 이동
# + [markdown] slideshow={"slide_type": "slide"}
# * 예제:
# * 아래 뿌리 지정 나무의 뿌리에서 출발하여 왼편 아랫쪽 방향으로 진행.
# * 더 이상 아래 방향으로 진행할 수 없으면 부모 마디로 돌아간 후 다른 형제자매 마디 중 가장
# 왼편에 위치한 마디로 이동 후 왼편 아랫쪽 방향으로의 이동 반복
# + [markdown] slideshow={"slide_type": ""}
# <div align="center"><img src="./images/algo05/algo05-02.png" width="400"/></div>
# + [markdown] slideshow={"slide_type": "slide"}
# #### 상태 공간 나무(state space tree)
# -
# * 변수가 가질 수 있는 모든 값을 마디(node)로 갖는 뿌리 지정 나무
# * **깊이**: 깊이가 0인 뿌리에서 출발하여 아래로 내려갈 수록 깊이가 1씩 증가.
# + [markdown] slideshow={"slide_type": "slide"}
# * 예제: `4x4`로 이루어진 체스판에 네 개의 체스 퀸을 놓을 수 있는 위치를
# 마디로 표현한 상태 공간 나무
# * 뿌리는 출발 마디로 표현하며, 체스 퀸의 위치와 상관 없음.
# * 깊이 $k$의 마디: $k$ 째 퀸이 놓일 수 있는 위치
# + [markdown] slideshow={"slide_type": ""}
# <div align="center"><img src="./images/algo05/algo05-03.png" width="500"/></div>
# + [markdown] slideshow={"slide_type": "slide"}
# #### 마디의 유망성
# + [markdown] slideshow={"slide_type": ""}
# * 지정된 특정 조건에 해당하는 마디를 __유망하다__라고 부름.
# + [markdown] slideshow={"slide_type": ""}
# * 예제: 네 개의 퀸을 위치시켜야 할 경우 첫째 퀸의 위치에 따라
# 둘째 퀸이 놓일 수 있는 위치의 유망성이 결정됨.
# * 아래 그림에서 2번 행의 1, 2번 칸은 유망하지 않음.
# + [markdown] slideshow={"slide_type": ""}
# <div align="center"><img src="./images/algo05/algo05-04.png" width="300"/></div>
# + [markdown] slideshow={"slide_type": "slide"}
# #### 가지치기(pruning)
# -
# * 특정 마디에서 시작되는 가지 제거하기
# + [markdown] slideshow={"slide_type": "slide"}
# * 예제: 4 x 4로 이루어진 체스판에 네 개의 체스 퀸을 놓을 수 있는 위치를
# 마디로 표현한 상태 공간 나무에서 유망하지 않은 마디에서 가지치기를 실행하면 아래 그림이 생성됨.
# + [markdown] slideshow={"slide_type": ""}
# <div align="center"><img src="./images/algo05/algo05-05.png" width="600"/></div>
# + [markdown] slideshow={"slide_type": "slide"}
# ### 되추적 알고리즘
# -
# 1. 상태 공간 나무의 뿌리로부터 깊이우선 탐색(DFS) 실행.
# <br>
#
# 2. 탐색 과정에서 유망하지 않은 마디를 만나면 가지치기 실행 후 부모 마디로 되돌아감(되추적, backtracking).
# <br>
#
# 3. 이후 다른 형제자매 마디를 대상으로 깊이우선 탐색 반복.
# 더 이상의 형제자매 마디가 없으면 형제자매가 있는 조상까지 되추적 실행.
# <br>
#
# 4. 탐색이 더 이상 진행할 수 없는 경우 알고리즘 종료
# + [markdown] slideshow={"slide_type": "slide"}
# #### 예제: 되추적 알고리즘을 활용한 4-퀸 문제 해결
# + [markdown] slideshow={"slide_type": ""}
# <div align="center"><img src="./images/algo05/algo05-06.png" width="600"/></div>
# + [markdown] slideshow={"slide_type": "slide"}
# ### 깊이우선 탐색 대 되추적 알고리즘 비교
# + [markdown] slideshow={"slide_type": ""}
# * 4-퀸 문제를 순수한 깊이우선 탐색으로 해결하고자 할 경우: 155 마디 검색
# + [markdown] slideshow={"slide_type": ""}
# * 4-퀸 문제를 되추적 알고리즘으로 해결하고자 하는 경우: 27 마디 검색
# + [markdown] slideshow={"slide_type": "slide"}
# ## 2절 n-퀸 문제
# + [markdown] slideshow={"slide_type": "slide"}
# * 4-퀸 문제를 일반화시킨 n-문제를 해결하는 되추적 알고리즘 구현하기
# + [markdown] slideshow={"slide_type": "fragment"}
# * 문제: n 개의 퀸(queen)을 서로 상대방을 위협하지 않도록 n x n 체스판에 위치시키기
# <br>
#
# * 변수: n 개의 퀸
# * 즉, 1번 퀸부터 n번 퀸.
# <br>
# <br>
#
# * 도메인: {1, 2, ..., n}
# * 즉, 1번 열부터 n번 열.
# <br>
# <br>
#
# * 조건: 두 개의 퀸이 하나의 행, 열, 또는 대각선 상에 위치하지 않음.
# + [markdown] slideshow={"slide_type": "slide"}
# ### 유망성 판단
# + [markdown] slideshow={"slide_type": ""}
# * 두 개의 퀸 $q_1, q_2$가 같은 대각선 상에 위치하려면 행과 열의 차이의 절댓값이 동일해야 함.
# (아래 그림 참조)
#
# $$
# \text{abs}(q_{1,r} - q_{2,r}) = \text{abs}(q_{1,c} - q_{2,c})
# $$
#
# 단, $(q_{1,r}, q_{1,c})$ 와 $(q_{2,r}, q_{2,c})$ 는
# 각각 $q_1$과 $q_2$가 위치한 행과 열의 좌표를 가리킴.
# + [markdown] slideshow={"slide_type": ""}
# <div align="center"><img src="./images/algo05/algo05-07.png" width="300"/></div>
# + [markdown] slideshow={"slide_type": "slide"}
# ### 예제: 4-퀸 문제 해결 되추적 알고리즘
# + slideshow={"slide_type": "slide"}
from typing import List, Dict
# 변수: 네 개의 퀸의 번호, 즉, 1, 2, 3, 4
variables = [1, 2, 3, 4]
# 도메인: 각각의 퀸이 자리잡을 수 있는 가능한 모든 열의 위치.
domains: Dict[int, List[int]] = {}
columns = [1, 2, 3, 4]
for var in variables:
domains[var] = columns
# -
# * 4-퀸 문제의 경우 각각의 퀸 모두 동일하게 1열부터 4열 어딘가에 위치할 수 있음.
# 단, 그 중에서 조건을 만족시키는 열을 찾아야 함.
domains
# + [markdown] slideshow={"slide_type": "slide"}
# #### 되추적 함수 구현
# -
# * 아래 되추적 함수 `backtracking_search_queens()`는 일반적인 n-퀸 문제를 해결함.
# * `assignment` 인자: 되추적 과정에서 일부의 변수에 대해 할당된 도메인 값의 정보를 담은 사전을 가리킴.
# * 인자가 들어오면 아직 값을 할당받지 못한 변수를 대상으로 유망성을 확인한 후 되추적 알고리즘 진행.
# * 되추적 알고리즘이 진행되면서 `assignment`가 확장되며 모든 변수에 대해 도메인 값이 지정될 때가지 재귀적으로
# 알고리즘이 진행됨.
# + slideshow={"slide_type": "slide"}
def backtracking_search_queens(assignment: Dict[int, int] = {}):
"""assignment: 각각의 변수를 키로 사용하고 키값은 해당 변수에 할당될 값"""
# 모든 변수에 대한 값이 지정된 경우 조건을 만족시키는 해가 완성된 것임
if len(assignment) == len(variables):
return assignment
# 아직 값을 갖지 않은 변수들이 존재하면 되추적 알고리즘을 아직 할당되지 않은 값을 대상으로 이어서 진행
unassigned = [v for v in variables if v not in assignment]
first = unassigned[0]
for value in domains[first]:
# 주의: 기존의 assignment를 보호하기 위해 복사본 활용
# 되추적이 발생할 때 이전 할당값을 기억해 두기 위해서임.
local_assignment = assignment.copy()
local_assignment[first] = value
# local_assignment 값이 유망하면 재귀 호출을 사용하여 변수 할당 이어감.
if promissing_queens(first, local_assignment):
result = backtracking_search_queens(local_assignment)
# 유망성을 이어가지 못하면 되추적 실행
if result is not None:
return result
return None
# + [markdown] slideshow={"slide_type": "slide"}
# #### 유망성 확인 함수
# -
def promissing_queens(variable: int, assignment: Dict[int, int]):
"""새로운 변수 variable에 값을 할당 하면서 해당 변수와 연관된 변수들 사이의 제약조건이
assignment에 대해 만족되는지 여부 확인
n-퀸 문제의 경우: 제약조건이 모든 변수에 대해 일정함.
즉, 새로 위치시켜야 하는 퀸이 기존에 이미 자리잡은 퀸들 중 하나와
동일 행, 열, 대각산 상에 위치하는지 여부를 확인함"""
# q1r, q1c: 첫째 퀸이 놓인 마디의 열과 행
for q1r, q1c in assignment.items():
# q2r = 첫째 퀸 아래에 위치한 다른 모든 퀸들을 대상으로 조건만족여부 확인
for q2r in range(q1r + 1, len(assignment) + 1):
q2c = assignment[q2r] # 둘째 퀸의 열
if q1c == q2c: # 동일 열에 위치?
return False
if abs(q1r - q2r) == abs(q1c - q2c): # 대각선상에 위치?
return False
# 모든 변수에 대해 제약조건 만족됨
return True
# + slideshow={"slide_type": "slide"}
backtracking_search_queens()
# + [markdown] slideshow={"slide_type": "slide"}
# ### n-퀸 문제 되추적 알고리즘의 시간 복잡도
# -
# * n 개의 퀸이 주어졌을 때 상태공간트리의 마디의 수는 다음과 같음.
#
# $$
# 1 + n + n^2 + n^3 + \cdots + n^n = \frac{n^{n+1}-1}{n-1}
# $$
# * 따라서 되추적 알고리즘이 최대 n의 지수승 만큼 많은 수의 마디를 검색해야 할 수도 있음.
# * 하지만 검색해야 하는 마디 수는 경우마다 다름.
# * 효율적인 알고리즘이 아직 알려지지 않음.
# + [markdown] slideshow={"slide_type": "slide"}
# ### 부록: 얕은(shallow) 복사 vs 깊은(deep) 복사
# -
# * 리스트의 `copy()` 메서드는 얕은 복사 용도로 사용된.
# * 1차원 리스트일 경우 새로운 리스트를 복사해서 만들어 냄.
# * 하지만 2차원 이상의 리스트 일 경우 모든 것을 복사하지는 않음. 아래 코드 참조.
# +
# 얕은 복사
aList = [1, 2, 3, 4]
bList = aList
cList = aList.copy()
aList[0] = 10
print("얕은 복사:", aList[0] == cList[0])
dList = [[5, 6], [7, 8]]
eList = dList.copy()
dList[0][1] = 60
print("얕은 복사:", dList[0] == eList[0])
# + [markdown] slideshow={"slide_type": "slide"}
# <div align="center"><img src="./images/algo05/algo05-13.png" width="700"/></div>
# + [markdown] slideshow={"slide_type": "slide"}
# * 깊은 차원까지 복사를 하려면 깊은 복사(deep copy)를 사용해야 함.
# * 방식1: 새로 정의
# * 방식2: `copy` 모듈의 `deepcopy()` 함수 활용
# * `copy()` 함수: 얕은 복사. 리스트의 `copy()` 메서드와 동일하게 작동.
# * `deepcopy()` 함수: 깊은 복사.
# +
# 얕은 복사 vs. 얕은 복사
from copy import copy, deepcopy
aList = [1, 2, 3, 4]
bList = aList
cList = copy(aList)
aList[0] = 10
print("얕은 복사:", aList[0] == cList[0])
dList = [[5, 6], [7, 8]]
eList = deepcopy(dList)
dList[0][1] = 60
print("깊은 복사:", dList[0] == eList[0])
# + [markdown] slideshow={"slide_type": "slide"}
# <div align="center"><img src="./images/algo05/algo05-14.png" width="700"/></div>
# + [markdown] slideshow={"slide_type": "slide"}
# ## 5절 그래프 색칠하기
# + [markdown] slideshow={"slide_type": "slide"}
# ### m-색칠하기
# + [markdown] slideshow={"slide_type": ""}
# * 주어진 비방향그래프에서 서로 인접한 마디를 최대 m 개의 색상을 이용하여
# 서로 다른 색을 갖도록 색칠하는 문제
# + [markdown] slideshow={"slide_type": "slide"}
# #### 예제
# -
# * 아래 그래프에 대한 2-색칠하기 문제의 해답은 없음.
# + [markdown] slideshow={"slide_type": ""}
# <div align="center"><img src="./images/algo05/algo05-08c.png" width="200"/></div>
# + [markdown] slideshow={"slide_type": "slide"}
# * 3-색칠하기 문제에 대해서는 해답 존재.
# + [markdown] slideshow={"slide_type": ""}
# <div align="center"><img src="./images/algo05/algo05-08d.png" width="200"/></div>
# + [markdown] slideshow={"slide_type": "slide"}
# #### 주요 응용분야
# -
# * 지도 색칠하기
# + [markdown] slideshow={"slide_type": "slide"}
# ### 평면그래프
# -
# * 서로 교차하는 이음선이 없는 그래프
# * 지도를 평면그래프로 변환 가능
# * 마디: 지도의 한 지역
# * 이음선: 서로 인접한 두 지역 연결
# + [markdown] slideshow={"slide_type": "slide"}
# #### 예제
# -
# * 왼편의 지도를 오른편의 평면그래프로 변환 가능함.
# + [markdown] slideshow={"slide_type": ""}
# <div align="center"><img src="./images/algo05/algo05-09a.png" width="400"/></div>
# + [markdown] slideshow={"slide_type": "slide"}
# ### 예제: 3-색칠하기 문제 해결 되추적 알고리즘
# -
# ```python
# colors = [빨강, 파랑, 갈색]
# = [1 , 2 , 3 ]
# ```
#
# <div align="center"><img src="./images/algo05/algo05-08d.png" width="200"/></div>
# + [markdown] slideshow={"slide_type": "slide"}
# <div align="center"><img src="./images/algo05/algo05-11.png" width="400"/></div>
# + slideshow={"slide_type": "slide"}
from typing import List, Dict
# 변수: 네 마디의 번호, 즉, 1, 2, 3, 4
variables = [1, 2, 3, 4]
# 도메인: 각각의 마디에 칠할 수 있는 가능한 모든 색상
# 3-색칠하기: 1(빨강), 2(파랑), 3(갈색)
domains: Dict[int, List[int]] = {}
columns = [1, 2, 3]
for var in variables:
domains[var] = columns
# -
# * 3-색칠하기 문제의 경우 각각의 마디에 동일하게 빨강, 파랑, 갈색 어느 색도 취할 수 있음.
# 단, 그 중에서 조건을 만족시키는 색상을 찾아야 함.
domains
# + [markdown] slideshow={"slide_type": "slide"}
# #### 되추적 함수 구현
# -
# * 아래 되추적 함수 `backtracking_search_colors()`는 일반적인 m-색칠하기 문제를 해결함.
# * `assignment` 인자: 되추적 과정에서 일부의 변수에 대해 할당된 도메인 값의 정보를 담은 사전을 가리킴.
# * 인자가 들어오면 아직 값을 할당받지 못한 변수를 대상으로 유망성을 확인한 후 되추적 알고리즘 진행.
# * 되추적 알고리즘이 진행되면서 `assignment`가 확장되며 모든 변수에 대해 도메인 값이 지정될 때가지 재귀적으로
# 알고리즘이 진행됨.
# + slideshow={"slide_type": "slide"}
def backtracking_search_colors(assignment: Dict[int, int] = {}):
"""assignment: 각각의 변수를 키로 사용하고 키값은 해당 변수에 할당될 값"""
# 모든 변수에 대한 값이 지정된 경우 조건을 만족시키는 해가 완성된 것임
if len(assignment) == len(variables):
return assignment
# 아직 값을 갖지 않은 변수들이 존재하면 되추적 알고리즘을 아직 할당되지 않은 값을 대상으로 이어서 진행
unassigned = [v for v in variables if v not in assignment]
first = unassigned[0]
for value in domains[first]:
# 주의: 기존의 assignment를 보호하기 위해 복사본 활용
# 되추적이 발생할 때 이전 할당값을 기억해 두기 위해서임.
local_assignment = assignment.copy()
local_assignment[first] = value
# local_assignment 값이 유망하면 재귀 호출을 사용하여 변수 할당 이어감.
if promissing_colors(first, local_assignment):
result = backtracking_search_colors(local_assignment)
# 유망성을 이어가지 못하면 되추적 실행
if result is not None:
return result
return None
# + [markdown] slideshow={"slide_type": "slide"}
# #### 유망성 확인 함수
# -
def promissing_colors(variable: int, assignment: Dict[int, int]):
"""새로운 변수 variable에 값을 할당 하면서 해당 변수와 연관된 변수들 사이의 제약조건이
assignment에 대해 만족되는지 여부 확인
m-색칠하기 문제의 경우: 이웃마디의 상태에 따라 제약조건이 달라짐.
즉, 마디 variable에 할당된 색이 이웃마디의 색과 달라야 함.
이를 위해 각각의 마디가 갖는 이웃마디들의 리스트를 먼저 확인해야 함."""
# 각 마디에 대한 이웃마디의 리스트
constraints = {
1 : [2, 3, 4],
2 : [1, 3],
3 : [1, 2, 4],
4 : [1, 3]
}
for var in constraints[variable]:
if (var in assignment) and (assignment[var] == assignment[variable]):
return False
return True
# + slideshow={"slide_type": "slide"}
backtracking_search_colors()
# + [markdown] slideshow={"slide_type": "slide"}
# ### m-색칠하기 문제 되추적 알고리즘의 시간 복잡도
# -
# * n 개의 마디를 m 개의 색으로 칠해야 하는 문제의 상태공간트리의 마디의 수는 다음과 같음.
#
# $$
# 1 + m + m^2 + m^3 + \cdots + m^n = \frac{m^{n+1}-1}{m-1}
# $$
# * 따라서 되추적 알고리즘이 최대 m과 n의 지승 만큼 많은 수의 마디를 검색해야 할 수도 있음.
# * 하지만 검색해야 하는 마디 수는 경우마다 다름.
# * 효율적인 알고리즘이 아직 알려지지 않음.
# + [markdown] slideshow={"slide_type": "slide"}
# ### 참고: 4색정리
# -
# * 4-색칠하기 문제는 언제나 해결가능함.
# + [markdown] slideshow={"slide_type": ""}
# <div align="center"><img src="./images/algo05/algo05-12a.png" width="600"/></div>
#
# <그림 출처: [위키피디아: 4색정리](https://ko.wikipedia.org/wiki/4색정리)>
# + [markdown] slideshow={"slide_type": "slide"}
# * 1852년에 영국인 <NAME>가 영국 지도를 작성할 때
# 인접한 각 주를 다른 색으로 칠하기 위해 필요한 최소한의 색상의 수에 대한 질문에서 유래한 문제임.
# + [markdown] slideshow={"slide_type": ""}
# * 해결
# * 1976년에 <NAME>과 <NAME> 이 해결
# * 500페이지 이상의 증명으로 이루어졌으며 일부 증명은 컴퓨터 프로그램을 사용하였음.
# * 증명에 사용된 컴퓨터 프로그램에 대한 신뢰성 때문에 100% 인정받지 못하였음.
# 하지만 사용된 컴퓨터 프로그램의 문제가 발견된 것은 아님.
# * 2005년에 <NAME>에 의해 두 사람의 증명이 옳았음이 검증됨.
# + [markdown] slideshow={"slide_type": "slide"}
# ### $m$-색칠하기 문제 해결가능성 판단 알고리즘
# -
# * $m$ 이 1 또는 2인 경우: 쉽게 판단됨.
# * $m = 3$ 인 경우: 효율적인 알고리즘 아직 찾지 못함.
# * 즉, 임의의 평면 지도에 대해 서로 인접한 지역은 다른 색상을 갖도록 3 가지 색상만을 이용하여 색칠할 수 있는지
# 여부를 판단하는 일이 매우 어려움.
# + [markdown] slideshow={"slide_type": "slide"}
# ## 연습문제
# + [markdown] slideshow={"slide_type": "slide"}
# #### 문제 1
#
# 5-퀸 문제를 해결하는 되추적 알고리즘을 단계별로 설명하라.
# + [markdown] slideshow={"slide_type": "slide"}
# * 앞서 소개한 n-퀸 알고리즘은 DFS 기법을 사용하며, 하나의 해답을 찾으면 바로 종료함.
# 따라서 아래 단계를 거치며 해답 하나를 구함.
#
# * 첫째 퀸: 1행 1열에 위치시킴
# * 둘째 퀸: 2행 3~5열이 유망함. 따라서 3열에 위치시킴.
# * 셋째 퀸: 3행 5열만이 유일하게 유망함. 따라서 5열에 위치시킴.
# * 넷째 퀸: 4행 2열만이 유일하게 유망함. 따라서 2열에 위치시킴.
# * 다섯째 퀸: 5행 4열만이 유일하게 유망함. 따라서 4열에 위치시킴.
# + [markdown] slideshow={"slide_type": ""}
# <div align="center"><img src="./images/algo05/algo05-15.png" width="150"/></div>
# + [markdown] slideshow={"slide_type": "slide"}
# * 앞서 작성한 `backtracking_search_queens()` 함수를 이용하여 확인하면 다음과 같음.
# + slideshow={"slide_type": ""}
# 변수: 네 개의 퀸의 번호, 즉, 1, 2, 3, 4, 5
variables = [1, 2, 3, 4, 5]
# 도메인: 각각의 퀸이 자리잡을 수 있는 가능한 모든 열의 위치.
domains: Dict[int, List[int]] = {}
columns = [1, 2, 3, 4, 5]
for var in variables:
domains[var] = columns
backtracking_search_queens()
# + [markdown] slideshow={"slide_type": "slide"}
# #### 문제 2
#
# 빨강(R), 초록(G), 파랑(B)이 주어졌을 때 되추적 알고리즘을 이용하여 아래 그래프를 색칠하는 과정을 단계별로 설명하라.
# + [markdown] slideshow={"slide_type": ""}
# <div align="center"><img src="./images/algo05/algo05-16.png" width="300"/></div>
# + [markdown] slideshow={"slide_type": "slide"}
# * 앞서 소개한 m-색칠하기 알고리즘은 DFS 기법을 사용하며, 하나의 해답을 찾으면 바로 종료함.
# 따라서 아래 단계를 거치며 해답 하나를 구함.
#
# * v1: R, G, B가 유망하지만 먼저 R 선택
# * v2: G, B가 유망하지만 먼저 G 선택
# * v3: R, B가 유망하지만 먼저 R 선택
# * v4: G, B가 유망하지만 먼저 G 선택
# * v5: R, B가 유망하지만 먼저 R 선택
# * v6: G, B가 유망하지만 먼저 G 선택
#
# + [markdown] slideshow={"slide_type": "slide"}
# * 앞서 작성한 `backtracking_search_colors()` 함수를 이용하여 확인하면 다음과 같음.
# + slideshow={"slide_type": ""}
# 변수: 여섯 개 마디의 번호, 즉, 1, 2, 3, 4, 5, 6
variables = [1, 2, 3, 4, 5, 6]
# 도메인: 각각의 마디에 칠할 수 있는 색상 세 개(R, G, B)
domains: Dict[int, List[int]] = {}
columns = [1, 2, 3]
for var in variables:
domains[var] = columns
# + [markdown] slideshow={"slide_type": "slide"}
# * 위 그래프에 의한 제약조건을 사용한 유망성 확인 함수는 다음과 같음.
# + slideshow={"slide_type": ""}
def promissing_colors(variable: int, assignment: Dict[int, int]):
"""새로운 변수 variable에 값을 할당 하면서 해당 변수와 연관된 변수들 사이의 제약조건이
assignment에 대해 만족되는지 여부 확인
m-색칠하기 문제의 경우: 이웃마디의 상태에 따라 제약조건이 달라짐.
즉, 마디 variable에 할당된 색이 이웃마디의 색과 달라야 함.
이를 위해 각각의 마디가 갖는 이웃마디들의 리스트를 먼저 확인해야 함."""
# 각 마디에 대한 이웃마디의 리스트
constraints = {
1 : [2, 4],
2 : [1, 3, 5],
3 : [2, 6],
4 : [1, 5],
5 : [2, 4, 6],
6 : [3, 5]
}
for var in constraints[variable]:
if (var in assignment) and (assignment[var] == assignment[variable]):
return False
return True
# + [markdown] slideshow={"slide_type": "slide"}
# * 실제로 빨강과 초록이 번갈아 사용됨을 아래와 같이 확인됨.
# + slideshow={"slide_type": ""}
backtracking_search_colors()
| slides/Algo-05-Backtracking-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
def dist(p1, p2):
s = 0
for (x, y) in zip(p1, p2):
s += (x-y)**2
result = s ** 0.5
return result
p = [2, 3, -1]
q = [4, 1, -2]
print("The result is", dist(p, q))
# -
| 01_basic-python/exercise_function.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CSV reader/writer
#
# ## CSV flies
#
# Run the appropriate code cell for your installation to create a CSV file from a multi-line raw text string:
# +
# Code for local installation of Jupyter notebooks
import os
print(os.getcwd())
some_text = '''given_name,family_name,username,student_id
Jimmy,Zhang,rastaman27,37258
Ji,Kim,kimji8,44947
Veronica,Fuentes,shakira<3,19846'''
with open('students.csv', 'wt', encoding='utf-8') as file_object:
file_object.write(some_text)
# +
# Code for Colab installation
# You will need to mount your Google drive before running the code.
# The file will be saved in the root of your Google Drive.
google_drive_root = '/content/drive/My Drive/'
some_text = '''given_name,family_name,username,student_id
Jimmy,Zhang,rastaman27,37258
Ji,Kim,kimji8,44947
Veronica,Fuentes,shakira<3,19846'''
with open(google_drive_root + 'students.csv', 'wt', encoding='utf-8') as file_object:
file_object.write(some_text)
# -
# For a local installation, the script prints the path where the file is saved. Open the file from within your text editor and examine the form of the text. Notice that each row of the CSV is on a separate line (ended with a newline character).
#
# Now open the file using a spreadsheet program. Libre Office is the best one to use, but if you don't have it, you can open it with Excel. Notice how the file is rendered as an editable table rather than as raw text.
# ## Reading CSV files as a list of lists using a csv.reader() object
#
# The `csv` module defines the `csv.reader()` object that inputs an iterable file object. The reader object an iterable object whose items are lists.
# +
import csv
with open('students.csv', 'r', newline='', encoding='utf-8') as file_object:
# The reader() object is instantiated and assigned to a variable.
reader_object = csv.reader(file_object)
print(type(reader_object))
print()
# The reader object is iterable
for row in reader_object:
# each iterated item is a Python list
print(type(row))
print(row)
# -
# In the following example, the code to read from the CSV file to a list of lists is placed in a function. The file path is passed in as the only argument of the function. The function returns a single object, a list of lists containing the data from the CSV file.
# +
import csv
path = '' # uncomment this line for a local installation
#path = '/content/drive/My Drive/' # uncomment this line if using Colab
# The function takes the file path as an argument and returns a list of lists
def read_csv_to_list_of_lists(file_path):
with open(file_path, 'r', newline='', encoding='utf-8') as file_object:
reader_object = csv.reader(file_object)
list_of_lists = []
for row_list in reader_object:
list_of_lists.append(row_list)
return list_of_lists
# Main script
student_info = read_csv_to_list_of_lists(path + 'students.csv')
print(student_info)
print()
print(student_info[1][2])
print()
# -
# Build a string that contains tabs between each table item and has newlines at the end of each line.
output_string = ''
for row in range(0,len(student_info)):
for column in range(0,len(student_info[row])):
output_string += student_info[row][column] + '\t'
output_string += '\n'
print(output_string)
# ## Writing to CSV files
#
# Example writing cartoons.csv
# +
import csv
path = '' # uncomment this line for a local installation
#path = '/content/drive/My Drive/' # uncomment this line if using Colab
data = [ ['name', 'company', 'nemesis'], ['<NAME>', 'Disney', '<NAME>'], ['Road Runner', '<NAME>', '<NAME>'] ]
with open(path + 'cartoons.csv', 'w', newline='', encoding='utf-8') as file_object:
writer_object = csv.writer(file_object)
for row in data:
print(row)
writer_object.writerow(row)
# -
# ## Reading CSV files as a list of dictionaries using a csv.DictReader() object
#
# The `csv` module contains the `DictReader()` object that turns an iterable file object into an iterable object whose items are dictionaries.
# +
import csv
path = '' # uncomment this line for a local installation
#path = '/content/drive/My Drive/' # uncomment this line if using Colab
with open('cartoons.csv', 'r', newline='', encoding='utf-8') as file_object:
# The DictReader() object is instantiated and assigned to a variable.
reader_object = csv.DictReader(file_object)
# The iterable items in a DictReader object are a special kind of dictionary (OrderedDict).
# But we can use them like regular dictionary if we ignore that they are ordered.
print(type(reader_object))
print()
# If we want to reuse the row dictionaries, we can add them to a list.
cartoon_table = []
for row_list in reader_object:
print(type(row_list))
print(row_list)
cartoon_table.append(row_list)
print()
# We refer to items in the row lists by their keys, just as we do for normal dictionaries.
# Because each row has its own dictionary, we must specify the row in the first square brackets.
print(cartoon_table[1]['name'] + ' works for ' + cartoon_table[1]['company'] + '. Its enemy is ' + cartoon_table[1]['nemesis'])
# -
for character in cartoon_table:
print('Character name:', character['name'], ' company:', character['company'], ' nemesis:', character['nemesis'])
# ## Template code for CSV-reading function (list of dictionaries)
# In the following example, the code to read from the CSV file to a list of dictionaries is placed in a function. The file path is passed in as the only argument of the function. The function returns a single object, a list of dictionaries containing the data from the CSV file. The keys for the dictionaries are taken from the header row of the CSV.
#
# The main script is a modification of the earlier script that looks up cartoon characters. By using a file rather than hard-coding the characters data, it's easier to include a lot more information and to change it by updating the CSV file as a spreadsheet.
#
# You can download a CSV file with around 4000 cartoon characters from [here](https://github.com/HeardLibrary/digital-scholarship/blob/master/code/pylesson/challenge4/cartoons.csv). Right click on the `Raw` button and select `Save file as...`. Save the file in the same directory as your Jupyter notebook if you are using a local installation, or in the root of your Google Drive if using Colab. **Note:** if your browser changes the file extension to `.txt`, you may need to change the format from `text` to `All Files`, then manually change the extension in the dialog from `.txt` to `.csv`.
#
# Many of the characters in the file do not have nemeses. You can add them if you know who they are.
# +
import csv
path = '' # uncomment this line for a local installation
#path = '/content/drive/My Drive/' # uncomment this line if using Colab
# The function takes the file path as an argument and returns a list of lists
def read_csv_to_list_of_dicts(filename):
with open(filename, 'r', newline='', encoding='utf-8') as file_object:
dict_object = csv.DictReader(file_object)
list_of_dicts = []
for row_dict in dict_object:
list_of_dicts.append(row_dict)
return list_of_dicts
# Main script
cartoons = read_csv_to_list_of_dicts(path + 'cartoons.csv')
name = input("What's the character? ")
found = False
for character in cartoons:
if name.lower() in character['name'].lower():
if character['nemesis'] == '':
print("I don't know the nemesis of " + character['name'])
else:
print(character['name'] + " doesn't like " + character['nemesis'])
found = True
if not found:
print("Sorry, I don't know that character.")
# -
# ## Template code for CSV-writing functions (from list of dictionaries)
#
# Note that the functions do not return anything since they output to a file.
#
# The file path will need to be adjusted if you want to save the file somewhere other than in the directory in which the notebook is running.
#
# The first function requires you to explicitly provide the field names. Use it if every dictionary does not contain every field.
# +
import csv
def write_dicts_to_csv_fieldnames(list_of_dicts, file_path, field_names):
with open(file_path, 'w', newline='', encoding='utf-8') as csv_file_object:
writer = csv.DictWriter(csv_file_object, fieldnames=field_names)
writer.writeheader()
for row_dict in list_of_dicts:
writer.writerow(row_dict)
field_names = ['name', 'company', 'nemesis']
data = [ {'name': '<NAME>', 'company': 'Disney', 'nemesis': 'Donald Duck'}, {'name': '<NAME>', 'company': 'Warner Brothers', 'nemesis': 'Wile Ethelbert Coyote'} ]
path = 'mini-cartoon-table.csv'
write_dicts_to_csv_fieldnames(data, path, field_names)
# -
# The second function gets the field names from the keys in the first dictionary in the list. Use it if all dictionaries have the same keys.
# +
import csv
def write_dicts_to_csv(list_of_dicts, file_path):
field_names = list_of_dicts[0].keys()
with open(file_path, 'w', newline='', encoding='utf-8') as csv_file_object:
writer = csv.DictWriter(csv_file_object, fieldnames=field_names)
writer.writeheader()
for row_dict in list_of_dicts:
writer.writerow(row_dict)
data = [ {'name': '<NAME>', 'company': 'Disney', 'nemesis': '<NAME>'}, {'name': '<NAME>', 'company': '<NAME>', 'nemesis': '<NAME>'} ]
path = 'another-cartoon-table.csv'
write_dicts_to_csv(data, path)
# -
# ## Reading CSV files from the Internet
#
# The Nashville schools data in this exercise comes from [here](https://github.com/HeardLibrary/digital-scholarship/blob/master/data/gis/wg/Metro_Nashville_Schools.csv).
#
# Reading a CSV file from a URL into a list of lists
# +
import requests
import csv
def url_csv_to_list_of_lists(url):
r = requests.get(url)
file_text = r.text.splitlines()
file_rows = csv.reader(file_text)
list_of_lists = []
for row in file_rows:
list_of_lists.append(row)
return list_of_lists
# Main script
url = 'https://raw.githubusercontent.com/HeardLibrary/digital-scholarship/master/data/gis/wg/Metro_Nashville_Schools.csv'
schools_data = url_csv_to_list_of_lists(url)
# print the IDs and names of all of the schools
print(schools_data[0][2] + '\t' + schools_data[0][3])
for school in range(1, len(schools_data)):
print(schools_data[school][2] + '\t' + schools_data[school][3])
# -
# Reading a CSV file from a URL into a list of dictionaries
# +
import requests
import csv
def url_csv_to_list_of_dicts(url):
r = requests.get(url)
file_text = r.text.splitlines()
file_rows = csv.DictReader(file_text)
list_of_dicts = []
for row in file_rows:
list_of_dicts.append(row)
return list_of_dicts
# Main script
url = 'https://raw.githubusercontent.com/HeardLibrary/digital-scholarship/master/data/gis/wg/Metro_Nashville_Schools.csv'
schools_data = url_csv_to_list_of_dicts(url)
# use the dictionary to look up a school ID
school_name = input("What's the name of the school? ")
found = False
for school in schools_data:
if school_name.lower() in school['School Name'].lower():
print('The ID number for', school['School Name'], 'is: ' + school['School ID'])
found = True
if not found:
print("I couldn't find that school.")
# -
| code/codegraf/022/022.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
import os
import glob
negatives = glob.glob('/home/husein/Malaya-Dataset/fake-news/negative/*.json')
positives = glob.glob('/home/husein/Malaya-Dataset/fake-news/positive/*.json')
negatives
texts, labels = [], []
for negative in negatives:
with open(negative) as fopen:
x = json.load(fopen)
texts.extend(x)
labels.extend([1] * len(x))
for positive in positives:
with open(positive) as fopen:
x = json.load(fopen)
texts.extend(x)
labels.extend([0] * len(x))
import numpy as np
import pandas as pd
np.unique(labels, return_counts = True)
df = pd.read_csv('Malaya-Dataset/fake-news/malaysia-scraping-syazanihussin.csv').dropna()
df.head()
df_fake = df.loc[df['Label'] == 'Fake']
df_real = df.loc[df['Label'] == 'Real']
df_fake.shape, df_real.shape
# +
from sklearn.model_selection import train_test_split
train_X, test_X, train_Y, test_Y = train_test_split(texts, labels, test_size = 0.2)
len(train_X), len(test_X)
# +
train_X.extend(df_fake['News'].tolist())
train_Y.extend([0] * len(df_fake['News'].tolist()))
train_X.extend(df_real['News'].tolist())
train_Y.extend([1] * len(df_real['News'].tolist()))
# -
len(train_X), len(train_Y)
# +
from sklearn.utils import shuffle
train_X, train_Y = shuffle(train_X, train_Y)
# +
with open('Malaya-Dataset/synonym/synonym0.json') as fopen:
s = json.load(fopen)
with open('Malaya-Dataset/synonym/synonym1.json') as fopen:
s1 = json.load(fopen)
# -
synonyms = {}
for l, r in (s + s1):
if l not in synonyms:
synonyms[l] = r + [l]
else:
synonyms[l].extend(r)
synonyms = {k: list(set(v)) for k, v in synonyms.items()}
# +
import random
def augmentation(s, maximum = 1.0):
s = s.split()
for i in range(int(len(s) * maximum)):
index = random.randint(0, len(s) - 1)
word = s[index]
sy = synonyms.get(word, [word])
sy = random.choice(sy)
s[index] = sy
return s
# -
aug = [' '.join(augmentation(x[0])) for _ in range(3)]
aug = list(set(aug))
x[0], aug
# +
from tqdm import tqdm
X, Y = [], []
for i in tqdm(range(len(train_X))):
aug = [' '.join(augmentation(train_X[i])) for _ in range(3)]
aug.append(train_X[i])
aug = list(set(aug))
X.extend(aug)
Y.extend([train_Y[i]] * len(aug))
# -
len(X), len(Y)
X, Y = shuffle(X, Y)
# +
import pickle
with open('relevant-dataset.pkl', 'wb') as fopen:
pickle.dump({'train_X': X, 'train_Y': Y,
'test_X': test_X, 'test_Y': test_Y}, fopen)
# -
| session/relevancy/augmenting-news.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Step 1: Computational Inductive Analysis
#
# Analysis 1: Difference of Proportions
#
# The last two cells produce the output used in Table 2.
# +
import pandas
from sklearn.feature_extraction.text import CountVectorizer
#read in data
df = pandas.read_csv("../data/comparativewomensmovement_dataset.csv", sep='\t', index_col=0, encoding='utf-8')
df
# +
#concatenate the documents from each organization together, creaing four strings
redstockings = df[df['org']=='redstockings']
redstockings_string = ' '.join(str(s) for s in redstockings['text_string'].tolist())
cwlu = df[df['org']=='cwlu']
cwlu_string = ' '.join(str(s) for s in cwlu['text_string'].tolist())
heterodoxy = df[df['org']=='heterodoxy']
heterodoxy_string = ' '.join(str(s) for s in heterodoxy['text_string'].tolist())
hullhouse = df[df['org']=='hullhouse']
hullhouse_string = ' '.join(str(s) for s in hullhouse['text_string'].tolist())
# -
#initialize countvectorizer function, removing stop words
countvec = CountVectorizer(stop_words="english")
# ### The next two cells produce the output used in Table 2
redstockings_cwlu = pandas.DataFrame(countvec.fit_transform([redstockings_string, cwlu_string]).toarray(), columns=countvec.get_feature_names())
redstockings_cwlu['word_count'] = redstockings_cwlu.sum(axis=1)
redstockings_cwlu = redstockings_cwlu.iloc[:,0:].div(redstockings_cwlu.word_count, axis=0)
redstockings_cwlu.loc[2] = redstockings_cwlu.loc[0] - redstockings_cwlu.loc[1]
#The words with the highest difference of proportions are distinct to Redstocking
#The words with the lowest (the highest negative) difference of proportions are distinct to CWLU
redstockings_cwlu.loc[2].sort_values(axis=0, ascending=False)
# +
#Heterodoxy versus Hull House
heterodoxy_hullhouse = pandas.DataFrame(countvec.fit_transform([heterodoxy_string, hullhouse_string]).toarray(), columns=countvec.get_feature_names())
heterodoxy_hullhouse['word_count'] = heterodoxy_hullhouse.sum(axis=1)
heterodoxy_hullhouse = heterodoxy_hullhouse.iloc[:,0:].div(heterodoxy_hullhouse.word_count, axis=0)
heterodoxy_hullhouse.loc[2] = heterodoxy_hullhouse.loc[0] - heterodoxy_hullhouse.loc[1]
#The words with the highest difference of proportions are distinct to Heterodoxy
#The words with the lowest (the highest negative) difference of proportions are distinct to Hull House
heterodoxy_hullhouse.loc[2].sort_values(axis=0, ascending=False)
| 01-Step1-PatternDetection/.ipynb_checkpoints/00-DifferenceOfProportions-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python
# name: conda-env-python-py
# ---
# <a href="https://cognitiveclass.ai"><img src = "https://ibm.box.com/shared/static/ugcqz6ohbvff804xp84y4kqnvvk3bq1g.png" width = 300, align = "center"></a>
#
# <h1 align=center><font size = 5>A solution for Data Importation issue " Databases and SQL for Data Science Week 4"</font></h1>
# this a simple solution for Data importation issue, in wich I change the date format to "YYYY-MM-DD HH:mm:ss"
# ## About the Author:
# **<NAME>**
# a software engineer
# <a href="https://www.linkedin.com/in/larbi-gouzal/">LinkedIn </a> <a href="https://github.com/gouzal">GitHub</a>
# ## CSV Importation
# +
import pandas as pd
url="https://ibm.box.com/shared/static/svflyugsr9zbqy5bmowgswqemfpm1x7f.csv"
df=pd.read_csv(url)
df.head()
# -
# ## Date's columns visualization
columns = ['DATE','UPDATEDON']
df_date = df[columns]
df_date.head()
# ## Date transformation
import datetime
df['DATE']=pd.to_datetime(df.DATE).dt.strftime('%Y-%m-%d')
df['UPDATEDON']=pd.to_datetime(df.UPDATEDON).dt.strftime('%Y-%m-%d %H:%M:%S')
df.head()
# ## Save new csv file
file_name= 'Chicago_Crime_Data-formated.csv';
df.to_csv(file_name, index = False)
# ## Save new CSV in IBM Storage
# ### configuration
# @hidden_cell
credentials ={
"apikey": "<KEY>",
"cos_hmac_keys": {
"access_key_id": "<KEY>",
"secret_access_key": "5e5965ff24b1d1df9c374b6aeccdde5e98950ef1767013fa"
},
"endpoints": "https://control.cloud-object-storage.cloud.ibm.com/v2/endpoints",
"iam_apikey_description": "Auto-generated for key <KEY>",
"iam_apikey_name": "Pyhon-Analysis-week5",
"iam_role_crn": "crn:v1:bluemix:public:iam::::serviceRole:Manager",
"iam_serviceid_crn": "crn:v1:bluemix:public:iam-identity::a/5ce679ad625c4b5d82e579cfbf861de3::serviceid:ServiceId-6f8dca1a-4786-470b-8754-ae5221f4e22b",
"resource_instance_id": "crn:v1:bluemix:public:cloud-object-storage:global:a/5ce679ad625c4b5d82e579cfbf861de3:c841a833-201b-4279-b330-12873044a5ef::"
}
endpoint = 'https://s3.eu.cloud-object-storage.appdomain.cloud'
bucket_name = 'pythonfordatascienceandaiweek5-donotdelete-pr-p5uzzqzsa6yzxc'
# +
import boto3
resource = boto3.resource(
's3',
aws_access_key_id = credentials["cos_hmac_keys"]['access_key_id'],
aws_secret_access_key = credentials["cos_hmac_keys"]["secret_access_key"],
endpoint_url = endpoint,
)
# +
import os
directory = os.getcwd()
file_path = directory + "/" + file_name
f = open(file_path,'r')
# -
# ### writing new csv in ibm cloud storage
# +
resource.Bucket(name=bucket_name).put_object(Key=file_name, Body=f.read())
Params = {'Bucket':bucket_name ,'Key':file_name }
import sys
time = 7*24*60**2
client = boto3.client(
's3',
aws_access_key_id = credentials["cos_hmac_keys"]['access_key_id'],
aws_secret_access_key = credentials["cos_hmac_keys"]["secret_access_key"],
endpoint_url=endpoint,
)
# -
# ### Printing the new csv URL
url = client.generate_presigned_url('get_object',Params=Params,ExpiresIn=time)
print(url)
# ## License
# This file is under MIT License 2019
| Databases and SQL for Data Science Week 4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Probability
#
# ### <NAME>
# #### August 14, 2017
# ## Objectives
#
# * Use permutations and combinations to solve probability problems.
# * Explain basic laws of probability.
# ## Agenda
#
# Morning
#
# * Review Sets
# * Permutations and combinations
# * Laws of Probability
# ## Some definitions
#
# * A set $S$ consists of all possible outcomes or events and is called the sample space
# * Union: $A \cup B = \{ x: x \in A ~\mathtt{ or} ~x \in B\}$
# * Intersection: $A \cap B = \{x: x \in A ~\mathtt{and} ~x \in B\}$
# * Complement: $A^\complement = \{ x: x \notin A \}$
# * Disjoint: $A \cap B = \emptyset$
# * Partition: a set of pairwise disjoint sets, ${A_j}$, such that $\underset{j=1}{\overset{\infty}{\cup}}A_j = S$
# * DeMorgan's laws: $(A \cup B)^\complement = A^\complement \cap B^\complement$ and $(A \cap B)^\complement = A^\complement \cup B^\complement$
# +
from scipy import stats
import numpy as np
import math
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ## Permutations and Combinations
#
# In general, there are $n!$ ways we can order $n$ objects, since there are $n$ that can come first, $n-1$ that can come 2nd, and so on. So we can line 16 students up $16!$ ways.
math.factorial(16)
# Suppose we choose 5 students at random from the class of 20 students. How many different ways could we do that?
#
# If the order matters, it's a **permutation**. If the order doesn't, it's a **combination**.
#
# There are $20$ ways they can choose one student, $20 \cdot 19$ ways we can choose two, and so on, so $$20\cdot19\cdot18\cdot17\cdot16 = \frac{20!}{15!} = {_{20}P_{15}}$$ ways we can choose five students, assuming the order matters. In general
#
# $$_nP_k = \frac{n!}{(n-k)!}$$
def permutations(n, k):
return math.factorial(n)/math.factorial(n-k)
permutations(20,5)
# There are $5!$ different way we can order those different students, so the number of combinations is that number divided by $5!$. We write this as $${20 \choose 5} = \frac{20!}{15! \cdot 5!}$$
#
# In general,
#
# $${n \choose k} = {_nC_k} = \frac{n!}{n!(n-k)!}$$
def combinations(n, k):
return math.factorial(n) / (math.factorial(n-k) * math.factorial(k))
combinations(20,5)
# ### Tea-drinking problem
#
# There's a classic problem in which a woman claims she can tell whether tea or milk is added to the cup first. The famous statistician <NAME> proposed a test: he would prepare eight cups of tea, four each way, and she would select which was which.
#
# Assuming the null hypothesis (that she was guessing randomly) what's the probability that she would guess all correctly?
# ## Multinomial
#
# Combinations explain the number of ways of dividing something into two categories. When dividing into more categories, use
#
# $${n \choose {n_1, n_2, ... n_k}} = \frac{n!}{n_1! n_2! ... n_k!}$$
#
# which reduces to the above for two cases.
# ## Definition of probability
#
# Given a sample space S, a *probability function* P of a set has three properties.
#
# * $P(A) \ge 0 \; \forall \; A \subset S$
# * $P(S) = 1$
# * For a set of pairwise disjoint sets $\{A_j\}$, $P(\cup_j A_j) = \sum_j P(A_j)$
# ## Independence
#
# Two events $A$ and $B$ are said to be *independent* iff
#
# $$ P(A \cap B) = P(A) P(B)$$
#
# or equivalently
#
# $$ P(B \mid A) = P(B)$$
#
# so knowlege of $A$ provides no information about $B$. This can also be written as $A \perp B$.
# ### Example: dice
#
# The probability of rolling a 1 on a single fair 6-sided die is $1\over 6$.
#
# What's the probability of two dice having a total value of 3?
# # Bayes' theorem
#
# Bayes' therem says that
#
# $$P(A\mid B) = \frac{P(B\mid A) P(A)}{P(B)}$$
# Where A and B are two possible events.
#
# To prove it, consider that
#
#
# $$\begin{equation}
# \begin{aligned}
# P(A\mid B) P(B) & = P(A \cap B) \\
# & = P(B \cap A) \\
# & = P(B\mid A) P(A) \\
# \end{aligned}
# \end{equation}
# $$
#
# so dividing both sides by $P(B)$ gives the above theorem.
#
# In here we usually think of A as being our hypothesis, and B as our observed data, so
#
# $$ P(hypothesis \mid data) = \frac{P(data \mid hypothesis) P(hypothesis)}{P(data)}$$
#
# where
# $$ P(data \mid hypothesis) \text{ is the likelihood} \\
# P(hypothesis) \text{ is the prior probability} \\
# P(hypothesis \mid data) \text{ is the posterior probability} \\
# P(data) \text{ is the normalizing constant} \\
# $$
#
#
# ## Law of Total Probability
#
# If ${B_n}$ is a partition of all possible options, then
#
# $$\begin{align}
# P(A) & = \sum_j P(A \cap B_j) \\
# & = \sum_j P(A \mid B_j) \cdot P(B_j)
# \end{align}
# $$
#
# ### Example: the cookie problem
#
# Bowl A has 30 vanilla cookies and 10 chocolate cookies; bowl B has 30 of each. You pick a bowl at random and draw a cookie. Assuming the cookie is vanilla, what's the probability it comes from bowl A?
# ### Example: two-sided coins
#
# There are three coins in a bag, one with two heads, another with two tails, another with a head and a tail. You pick one and flip it, getting a head. What's the probability of getting a head on the next flip?
# ## Probability chain rule
#
#
# $$\begin{align}
# P(A_n, A_{n-1}, ..., A_1) & = P(A_n \mid A_{n-1},...,A_1) \cdot P(A_{n-1},...,A_1) \\
# & = P(A_n \mid A_{n-1},...,A_1) \cdot P(A_{n-1} \mid A_{n-2},...,A_1) \cdot P(A_{n-1},...,A_1) \\
# & = \prod_{j=1}^n P(A_j \mid A_{j-1},...,A_1)
# \end{align}
# $$
| probability/lecture/.ipynb_checkpoints/my_Probability_AM-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Combining Data
#
# Practice combining data from two different data sets. In the same folder as this Jupyter notebook, there are two csv files:
# * rural_population_percent.csv
# * electricity_access_percent.csv
#
# They both come from the World Bank Indicators data.
# * https://data.worldbank.org/indicator/SP.RUR.TOTL.ZS
# * https://data.worldbank.org/indicator/EG.ELC.ACCS.ZS
#
# The rural populaton data represents the percent of a country's population that is rural over time. The electricity access data shows the percentage of people with access to electricity.
#
# In this exercise, you will combine these two data sets together into one pandas data frame.
# # Exercise 1
#
# Combine the two data sets using the [pandas concat method](https://pandas.pydata.org/pandas-docs/stable/merging.html). In other words, find the union of the two data sets.
"https://api.worldbank.org/v2/country/all/indicator/SP.RUR.TOTL.ZS?date=2000:2001&format=json"
# +
# pd.read_json('http://api.worldbank.org/v2/indicator/SP.RUR.TOTL.ZS/?format=json')
# -
df_rural = pd.read_csv('./rural_population_percent.csv',skiprows=4 )
df_rural.head()
df_electricity = pd.read_csv('./electricity_access_percent.csv',skiprows=4 )
df_electricity.head()
df_concat = pd.concat([df_rural, df_electricity])
# +
# TODO: import the pandas library
import pandas as pd
# TODO: read in each csv file into a separate variable
# HINT: remember from the Extract material that these csv file have some formatting issues
# HINT: The file paths are 'rural_population_percent.csv' and 'electricity_access_percent.csv'
df_rural = pd.read_csv('./rural_population_percent.csv',skiprows=4 )
df_electricity = pd.read_csv('./electricity_access_percent.csv',skiprows=4)
# TODO: remove the 'Unnamed:62' column from each data set
df_rural.drop('Unnamed: 62', axis=1, inplace=True)
df_electricity.drop('Unnamed: 62', axis=1, inplace=True)
# TODO: combine the two data sets together using the concat method
df_concat = pd.concat([df_rural, df_electricity])
df_concat
# -
# # Exercise 2 (Challenge)
#
# This exercise is more challenging.
#
# The resulting data frame should look like this:
#
# |Country Name|Country Code|Year|Rural_Value|Electricity_Value|
# |--|--|--|--|--|--|
# |Aruba|ABW|1960|49.224|49.239|
# ... etc.
#
# Order the results in the dataframe by country and then by year
#
# Here are a few pandas methods that should be helpful:
# * [melt](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.melt.html)
# * [drop](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.drop.html)
# * [merge](https://pandas.pydata.org/pandas-docs/version/0.23/generated/pandas.DataFrame.merge.html)
# * [sort_values](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.sort_values.html)
#
# HINT: You can use country name, country code, and the year as common keys between the data sets
df[]
df_rural.head()
df_electricity.head()
# +
# TODO: merge the data sets together according to the instructions. First, use the
# melt method to change the formatting of each data frame so that it looks like this:
# Country Name, Country Code, Year, Rural Value
# Country Name, Country Code, Year, Electricity Value
df_rural_melt = pd.melt(df_rural,\
id_vars=['Country Name', 'Country Code', 'Indicator Name', 'Indicator Code'],\
var_name = 'Year', value_name='Rural_Value')
df_electricity_melt = pd.melt(df_electricity,\
id_vars = ['Country Name','Country Code','Indicator Name','Indicator Code'],\
var_name ='Year', value_name='Electricity_Value')
# TODO: drop any columns from the data frames that aren't needed
df_rural_melt.drop(['Indicator Name', 'Indicator Code'], axis=1, inplace=True)
df_electricity_melt.drop(['Indicator Name', 'Indicator Code'], axis=1, inplace=True)
# TODO: merge the data frames together based on their common columns
df_merge = df_rural_melt.merge(df_electricity_melt, how='outer',\
on=['Country Name', 'Country Code', 'Year'])
# in this case, the common columns are Country Name, Country Code, and Year
# TODO: sort the results by country and then by year
df_combined = df_merge.sort_values(['Country Name', 'Year'])
df_combined
# -
df_merge.head()
df_electricity_melt[df_electricity_melt['Year']=='2016']
df_rural_melt
| lessons/ETLPipelines/5_combinedata_exercise/5_combining_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import ipdb
train_label_location = r'C:\Users\10169\Desktop\ECE449\data\labels\train_tutorial'
train_label_dir = os.listdir(train_label_location)
# print(train_label_dir)
train_image_location = r'C:\Users\10169\Desktop\ECE449\data\images\train_tutorial'
train_image_dir = os.listdir(train_image_location)
# -
valid_data = {}
for file in train_label_dir:
path = os.path.join(train_label_location,file)
fopen=open(path,'r')
all_numbers = [each_line for each_line in fopen.readlines() if each_line[0]=='0']
if len(all_numbers):
with open(path,'w') as f:
for each in all_numbers:
f.write(each)
key = '.'.join(file.split('.')[:-1])
valid_data[key]=1
else:
fopen.close()
os.remove(path)
for file in train_image_dir:
key = '.'.join(file.split('.')[:-1])
#ipdb.set_trace()
if not (key in valid_data.keys()):
path = os.path.join(train_image_location,file)
os.remove(path)
| .ipynb_checkpoints/data_clean-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/hwata3535/KataGo/blob/master/colab_katago.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="fc701xWwO1DC"
# 이 문서는 Google Colaboratory에 카타고를 세팅하고 리지나 사바키(또는 다른 GTP 엔진을 지원하는 도구)와 연결하는 방법을 설명합니다.
# + [markdown] id="rDYNTCaU8KCr"
# # 1단계 : Colab 서버에서 KataGo와 SSH 실행하기
#
# 실행하기 전에 NGROK_TOKEN 값을 입력해 주어야 합니다.
#
# PC마다 값이 다르기 때문에 리지를 실행하시는 컴퓨터가 바뀐 경우는 재입력 하셔야 합니다.
#
# 아래의 링크를 눌러서 이동하여 맨 위에 Copy 버튼을 누르시면 토큰값이 복사됩니다.
#
# [https://dashboard.ngrok.com/auth/your-authtoken](https://dashboard.ngrok.com/auth/your-authtoken).
#
# 그 다음 아래 스크립트 영역 맨 윗줄의 입력하는 곳에 붙여넣기 하시면 됩니다.
#
# 원하시는 경우에는 USER_NAME과 USER_PASSWORD 값도 바꾸실 수 있습니다.(이 경우 나중에 PC의 리지 설정에서도 똑같이 바꿔주어야 합니다.)
#
# 그 다음에는 아래에서 실행버튼(동그란 아이콘)을 눌러서 실행하시면 됩니다.
#
# 전체 스크립트가 실행되는 데 1~2분 정도 걸립니다.
#
# 참고로, 자동으로 서버에 할당된 GPU가 CUDA를 지원하지 않는 경우 처음 리지를 실행했을 때 OpenCL 튜닝 절차가 진행될 것입니다.
#
# CUDA 버전이 선택된 경우 대략 NVIDIA RTX 2060 정도의 성능이 나오는 듯 합니다.
# + id="nPaa2KJt8Kyn" colab={"base_uri": "https://localhost:8080/"} outputId="87b87fba-0972-4125-d6b5-a2f7c8fb44c1"
NGROK_TOKEN="<KEY>"
USER_NAME="katago"
USER_PASSWORD="<PASSWORD>"
# optional args
# supports: OPENCL, CUDA or AUTO
KATAGO_BACKEND="AUTO"
# supports: 40b, k15, 20b, 40b-large or AUTO
WEIGHT_FILE="20b"
import subprocess
if KATAGO_BACKEND == "AUTO":
gpu_name=str(subprocess.check_output("nvidia-smi -q | grep \"Product Name\" | cut -d\":\" -f2 | tr -cd '[:alnum:]._-'", shell=True), encoding='utf-8')
if gpu_name == "TeslaV100-SXM2-16GB":
KATAGO_BACKEND="OPENCL"
else:
KATAGO_BACKEND="OPENCL"
# !echo "Using Katago Backend: " $KATAGO_BACKEND
# !echo "Using Katago Weight: " $WEIGHT_FILE
# !echo "GPU : " $gpu_name
weight_urls = {
'40b': 'https://github.com/lightvector/KataGo/releases/download/v1.4.5/g170-b40c256x2-s5095420928-d1229425124.bin.gz',
'30b': 'https://github.com/lightvector/KataGo/releases/download/v1.4.5/g170-b30c320x2-s4824661760-d1229536699.bin.gz',
'20b': 'https://github.com/lightvector/KataGo/releases/download/v1.4.5/g170e-b20c256x2-s5303129600-d1228401921.bin.gz',
'40b-large': 'https://github.com/kinfkong/katago-colab/releases/download/v1.4.5/40b384.bin.gz'
}
# Install useful stuff
# ! apt-get update 1>/dev/null
# ! apt install --yes ssh screen nano htop ranger git libzip4 1>/dev/null
# ! pip install oss2 1>/dev/null
# %cd /content
# !rm -rf katago-colab
# !git clone https://github.com/kinfkong/katago-colab.git 1>/dev/null
#download the binarires
# !wget --quiet https://github.com/kinfkong/katago-colab/releases/download/v1.4.5/katago-$KATAGO_BACKEND -O katago
# !chmod +x /content/katago
# !wget --quiet https://github.com/kinfkong/katago-colab/releases/download/v1.4.5/ngrok -O ngrok
# !chmod +x /content/ngrok
# !mkdir -p /root/.katago/
# !cp -r /content/katago-colab/opencltuning /root/.katago/
#download the weights
weight_url = weight_urls[WEIGHT_FILE]
# !wget --quiet $weight_url -O $WEIGHT_FILE".bin.gz"
# !rm -rf weight.bin.gz
# !ln -s $WEIGHT_FILE".bin.gz" weight.bin.gz
# SSH setting
# ! echo "root:$USER_PASSWORD" | chpasswd
# ! echo "PasswordAuthentication yes" > /etc/ssh/sshd_config
# ! echo "PermitUserEnvironment yes" >> /etc/ssh/sshd_config
# ! echo "PermitRootLogin yes" >> /etc/ssh/sshd_config
# ! mkdir -p /root/.ssh
# generate the keys
# #!ssh-keygen -q -t rsa -N '' -f /root/.ssh/id_rsa <<<y 2>&1 >/dev/null
# #!cat /root/.ssh/id_rsa.pub >> /root/.ssh/authorized_keys
# ! service ssh restart > /dev/null
# Run ngrok
get_ipython().system_raw('./ngrok authtoken $NGROK_TOKEN && ./ngrok tcp 22 &')
# ! sleep 5
import oss2
import requests
import json
from re import sub
r = requests.get('http://localhost:4040/api/tunnels')
raw_ssh = r.json()['tunnels'][0]['public_url']
ssh_args = (sub("tcp://", "", raw_ssh)).split(':')
ssh_option = {
'host': ssh_args[0],
'port': int(ssh_args[1]),
'user': 'root'
}
ssh_option_json = json.dumps(ssh_option)
endpoint = 'http://oss-cn-beijing.aliyuncs.com'
auth = oss2.Auth('<KEY>', 'Q22UstMdKX8zZY9BqkGWWd2XbEBvPH')
bucket = oss2.Bucket(auth, endpoint, 'kata-config')
key = USER_NAME + <KEY>'
bucket.put_object(key, ssh_option_json)
with open('/content/katago-colab/config/gtp_colab.cfg', mode='w') as f:
f.write('logDir = gtp_logs\n')
f.write('logAllGTPCommunication = true\n')
f.write('logSearchInfo = true\n')
f.write('logToStderr = false\n')
f.write('ogsChatToStderr = true\n')
f.write('analysisWideRootNoise = 0.04\n')
f.write('rules = korean\n')
f.write('defaultBoardSize = 19\n')
f.write('allowResignation = true\n')
f.write('resignThreshold = -0.90\n')
f.write('resignConsecTurns = 3\n')
f.write('avoidMYTDaggerHack = true\n')
f.write('ponderingEnabled = true\n')
f.write('lagBuffer = 0.1\n')
f.write('numSearchThreads = 32\n')
f.write('nnMaxBatchSize = 16\n')
f.write('searchFactorAfterOnePass = 0.50\n')
f.write('searchFactorAfterTwoPass = 0.25\n')
f.write('searchFactorWhenWinning = 0.40\n')
f.write('searchFactorWhenWinningThreshold = 0.95\n')
# !echo -e "\n[KataGo Config]"
# !cat /content/katago-colab/config/gtp_colab.cfg
# !echo -e "\n"
# !echo "done!"
# + id="0Lt2YeRP1ZCA"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="movIaTQSRL96"
# # 2단계 : Colab KataGo를 사바키 또는 리지에 연결하기<br>(세븐틴이 배포한 패키지 안에는 이미 세팅이 완료되어 있음)
#
# 세븐틴 배포 패키지 다운로드 :
# http://naver.me/58hVIZEz
#
# ## 1) 먼저 아래에서 colab-katago 프로그램을 다운로드합니다.
# 다운로드 랑크:
# **For Windows Users (64bit windows)**
# https://github.com/kinfkong/katago-colab/releases/download/v1.4.5/colab-katago.windows.zip
# **For Linux Users**
# https://github.com/kinfkong/katago-colab/releases/download/v1.4.5/colab-katago.linux.zip
# **For Mac OSX Users**
# https://github.com/kinfkong/katago-colab/releases/download/v1.4.5/colab-katago.mac.zip
#
#
# **다운로드하신 후에 압축파일 안에 있는 colab-katago 또는 colab-katago.exe 파일을 리지 폴더에 복사해 넣습니다.**
#
# ## 2) 다음, 사바키 또는 리지에 엔진을 등록합니다.
#
# To configure the engine in `Sabaki` or `Lizzie`, you just need to fill the absolute path of your `colab-katago` program (which you've downloaded just now), and the username, password(i.e, the `USER_NAME`, `USER_PASSWORD` you configured in Step 1).
#
#
# **Sabaki Example**:
#
# 
#
# **Lizzie Example**:
#
# 
#
# ## 추가 설정(선택)
# 사바키에서 visits 또는 착수시간을 지정할 수 있습니다.(아래 참조)
#
# ```
# <AbsolutePathOfColabKataGoProglem> <USER_NAME> <USER_PASSWORD> 30s
# ```
# 30초로 착수시간 지정
# ```
# <AbsolutePathOfColabKataGoProglem> <USER_NAME> <USER_PASSWORD> 1600v
# ```
# 1600 visits 지정
| colab_katago.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Knowledge Base Classification
#
# ## Load imports.
# +
# Make common scripts visible and unsupervised classifier code
import sys
sys.path.append('../common/')
sys.path.append('../kb-classifier/')
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from collections import defaultdict
import requests
from loader import load_preprocessed_data
from lookup_tables import topic_code_to_topic_dict, topic_code_to_int, int_to_topic_code
from kb_classifier import KnowledgeBaseClassifier
from kb_common import wiki_topics_to_actual_topics
# -
# ## Load the data.
# +
x, y = load_preprocessed_data('data/rcv1_kb.csv')
x = np.array(x)
y = np.array(y)
# Get 20% test
total_examples = len(y)
split_point = int(total_examples * 0.8)
test_x = x[split_point:]
test_y = y[split_point:]
# Take N documents of each type from the training set for classifier tuning
train_x = []
train_y = np.zeros(shape=6000)
counts = np.zeros(shape=len(topic_code_to_int.keys()))
current_index = 0
print(split_point)
for i in range(split_point):
topic_int = y[i]
if counts[topic_int] < 1000:
train_x.append(x[i])
train_y[current_index] = topic_int
counts[topic_int] += 1
current_index += 1
print(counts)
# -
# ## Initialise and tune class probabilities for unsupervised learner
np.random.seed(42)
kb_predictor = KnowledgeBaseClassifier(topic_code_to_topic_dict.values(), topic_depth=1)
kb_predictor.train(train_x, train_y)
# +
predict_y = kb_predictor.predict(train_x)
classification_report, confusion_matrix = kb_predictor.get_classification_report(train_y, predict_y)
print(classification_report)
print(confusion_matrix)
# -
# ## Some Data Exploration
# Look at mean topic probabilities for each class.
# +
import matplotlib
matplotlib.rcParams.update({'font.size': 12})
topic_code_to_topic_dict = {
'GCRIM': 'Crime/Law',
'E11': 'Economic',
'GVOTE': 'Elections',
'GHEA': 'Health',
'GREL': 'Religion',
'GSPO': 'Sports'
}
prob_means = np.zeros(shape=(6, 6))
# Plot graph of mean topic probabilities for each topic class
for index, topic_code in int_to_topic_code.items():
prob_mean = np.mean(kb_predictor.last_class_probabilities[train_y == index], axis=0)
prob_std = np.std(kb_predictor.last_class_probabilities[train_y == index], axis=0)
prob_means[index] = prob_mean
print(prob_std)
plt.figure()
plt.title('Mean Topic Probabilities for {} Articles'.format(topic_code_to_topic_dict[topic_code]))
plt.xlabel('Probability')
plt.xlim(0.0, 0.7)
sns.barplot(x=prob_mean, y=list(topic_code_to_topic_dict.values()), xerr=prob_std)
plt.savefig('topic_prob_{}.pdf'.format(topic_code_to_topic_dict[topic_code].replace('/', '_')), bbox_inches='tight')
plt.show()
# Plot graph of mean topic probabilities for each topic class
for index, topic_code in int_to_topic_code.items():
prob_mean = np.mean(kb_predictor.last_class_probabilities[train_y == index], axis=0)
prob_std = np.std(kb_predictor.last_class_probabilities[train_y == index], axis=0)
plt.figure()
plt.title('Standardised Value for {} Articles'.format(topic_code_to_topic_dict[topic_code]))
plt.xlabel('Standardised Value')
plt.xlim(-2.5, 2.5)
#sns.barplot(x=((prob_mean-np.mean(prob_means, axis=0))/np.std(prob_means, axis=0)),
# y=list(topic_code_to_topic_dict.values()))
sns.barplot(x=((prob_mean-np.mean(kb_predictor.last_class_probabilities, axis=0))
/np.std(kb_predictor.last_class_probabilities, axis=0, ddof=1)),
y=list(topic_code_to_topic_dict.values()))
plt.savefig('standardised_{}.pdf'.format(topic_code_to_topic_dict[topic_code].replace('/', '_')), bbox_inches='tight')
plt.show()
# -
# ## Assess unsupervised classifier performance.
# +
print('Making predictions for {} documents'.format(len(test_y)))
predict_y = kb_predictor.predict(test_x)
classification_report, confusion_matrix = kb_predictor.get_classification_report(test_y, predict_y)
print(classification_report)
print(confusion_matrix)
# -
# ## Find examples where predictions went wrong
for topic_code, index in topic_code_to_int.items():
topic_subset = predict_y[test_y == index]
topic_subset_incorrect = topic_subset[topic_subset != index]
document_subset = test_x[test_y == index]
document_subset = document_subset[topic_subset != index]
print('------ 5 random erroneous predictions for {} ------'.format(topic_code_to_topic_dict[topic_code]))
print('')
random_indices = np.random.choice(np.arange(len(topic_subset_incorrect)), 5)
for index in random_indices:
print(document_subset[index])
print('')
print('Above classified as {}'.format(topic_code_to_topic_dict[int_to_topic_code[topic_subset_incorrect[index]]]))
print('')
print('')
| rcv1/knowledge_base.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.8 ('ds')
# language: python
# name: python3
# ---
import helper
import pandas as pd
import os
import matplotlib.pyplot as plt
from glob import glob
import numpy as np
if not os.path.exists('celeba'):
helper.download_celeba()
# +
identity_df = pd.read_csv('celeba/identity_CelebA.txt', sep=' ', header=None)
FILE_COLUMN = 0
PERSON_COLUMN = 1
def get_person_files(person_id):
return [os.path.join('celeba', 'img_align_celeba', s) for s in identity_df[identity_df[PERSON_COLUMN] == person_id][FILE_COLUMN]]
# -
show_n_images = 25
mnist_images = helper.get_batch(glob(os.path.join('celeba', 'img_align_celeba/*.jpg'))[:show_n_images], 50, 50, 'RGB')
plt.imshow(helper.images_square_grid(mnist_images, 'RGB'))
fig, axs = plt.subplots(2, 5)
fig.set_size_inches(12, 4)
for i in range(2):
for j in range(5):
mnist_images = helper.get_batch(get_person_files(1 + i*5+j), 50, 50, 'RGB')
axs[i, j].imshow(helper.images_square_grid(mnist_images, 'RGB'))
| celeba.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # [作業目標]
# - 仿造範例的 One Hot Encoding, 將指定的資料進行編碼
# - 請閱讀 [One Hot Encoder vs Label Encoder](https://medium.com/@contactsunny/label-encoder-vs-one-hot-encoder-in-machine-learning-3fc273365621)
# # [作業重點]
# - 將 sub_train 進行 One Hot Encoding 編碼 (In[4], Out[4])
# + colab={} colab_type="code" id="lh9gyloHVblR"
import os
import numpy as np
import pandas as pd
# + colab={} colab_type="code" id="z4iLB9cKVblV"
# 設定 data_path, 並讀取 app_train
dir_data = '../data/'
f_app_train = os.path.join(dir_data, 'application_train.csv')
app_train = pd.read_csv(f_app_train)
# + [markdown] colab={} colab_type="code" id="d5IeEcPVVblq"
# ## 作業
# 將下列部分資料片段 sub_train 使用 One Hot encoding, 並觀察轉換前後的欄位數量 (使用 shape) 與欄位名稱 (使用 head) 變化
# -
sub_train = pd.DataFrame(app_train['WEEKDAY_APPR_PROCESS_START'])
print(sub_train.shape)
sub_train.head()
"""
Your Code Here
"""
sub_train = pd.get_dummies(sub_train)
sub_train.head(10)
sub_train.shape
| D6_EDA_欄位的資料類型介紹及處理/Day_006_HW.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %%capture
## compile PyRoss for this notebook
import os
owd = os.getcwd()
os.chdir('../../')
# %run setup.py install
os.chdir(owd)
# %matplotlib inline
import numpy as np
import pyross
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
#from matplotlib import rc; rc('text', usetex=True)
# +
M = 2 # the population has two age groups
N = 5e4 # and this is the total population
# set the age structure
fi = np.array([0.25, 0.75]) # fraction of population in age age group
Ni = N*fi
# set the contact structure
C = np.array([[18., 9.], [3., 12.]])
def contactMatrix(t):
return C
# duration of simulation and data file
Tf = 100;
Nt=Tf+1;
beta = 0.02
alpha = 0.2
gIa = 1/7
gIs = 1/7
fsa = 1
# set up initial condition
Ia0 = np.array([20, 20]) # each age group has asymptomatic infectives
Is0 = np.array([20, 20]) # and also symptomatic infectives
R0 = np.array([0, 0]) # there are no recovered individuals initially
S0 = Ni - (Ia0 + Is0 + R0)
Tf = 100
Nf = Tf+1
def contactMatrix(t):
return C
parameters = {'alpha':alpha, 'beta':beta, 'gIa':gIa, 'gIs':gIs,'fsa':fsa}
true_parameters = {'alpha':alpha, 'beta':beta, 'gIa':gIa, 'gIs':gIs,'fsa':fsa}
# use pyross stochastic to generate traj and save
sto_model = pyross.stochastic.SIR(parameters, M, Ni)
data = sto_model.simulate(S0, Ia0, Is0, contactMatrix, Tf, Nf)
data_array = data['X']
# +
fig = plt.figure(num=None, figsize=(10, 8), dpi=80, facecolor='w', edgecolor='k')
plt.rcParams.update({'font.size': 22})
t = data['t']
plt.fill_between(t, 0, np.sum(data_array[:, :M], axis=1), alpha=0.3)
plt.plot(t, np.sum(data_array[:, :M], axis=1), '-', label='S', lw=4)
plt.fill_between(t, 0, np.sum(data_array[:, M:2*M], axis=1), alpha=0.3)
plt.plot(t, np.sum(data_array[:, M:2*M], axis=1), '-', label='Ia', lw=4)
plt.fill_between(t, 0, np.sum(data_array[:, 2*M:3*M], axis=1), alpha=0.3)
plt.plot(t, np.sum(data_array[:, 2*M:3*M], axis=1), '-', label='Is', lw=4)
plt.legend(fontsize=26)
plt.grid()
plt.xlabel(r'time')
plt.autoscale(enable=True, axis='x', tight=True)
# +
# load the data and rescale to intensive variables
Tf_inference = 20 # truncate to only getting the first few datapoints
Nf_inference = Tf_inference+1
steps = 21 # number internal integration steps taken
x = (data_array[:Nf_inference]).astype('float')/N
# +
inference_parameters = {
'betaIa' : beta*alpha,
'betaIs' : beta*(1-alpha),
'gIa' : gIa,
'gIs' : gIs
}
model_spec = {
"classes" : ["S", "Ia", "Is"],
"S" : {
"linear" : [],
"infection" : [ ["Ia", "-betaIa"], ["Is", "-betaIa"], ["Ia", "-betaIs"], ["Is", "-betaIs"]]
},
"Ia" : {
"linear" : [ ["Ia", "-gIa"] ],
"infection" : [ ["Ia", "betaIa"], ["Is", "betaIa"] ]
},
"Is" :{
"linear" : [ ["Is", "-gIs"] ],
"infection" : [ ["Is", "betaIs"], ["Ia", "betaIs"]]
}
}
# initialise the estimator
estimator = pyross.inference.Spp(model_spec, inference_parameters, M, fi, int(N), steps)
# +
# a filter that sums over all the infected people for each age group
fltr = np.kron([0, 1, 1],np.identity(M))
print(fltr)
# Compare the deterministic trajectory and the stochastic trajectory with the same
# initial conditions and parameters
obs=np.einsum('ij,kj->ki', fltr, x)
x0=x[0]
# compute -log_p for the original (correct) parameters
logp = estimator.minus_logp_red(inference_parameters, x0, obs[1:], fltr, Tf_inference, Nf_inference, contactMatrix)
print(logp)
# +
# make parameter guesses and set up bounds for each parameter
eps=1e-4
betaIs_g = 0.02
betaIs_std = 0.02
betaIs_b = (eps, 0.1)
betaIa_g = 0.005
betaIa_std = 0.003
betaIa_b = (eps, 0.1)
gIa_g = 0.2
gIa_std = 0.1
gIa_b = (eps, 1)
gIs_g = 0.2
gIs_std = 0.1
gIs_b = (eps, 1)
Ia0_g = (Ia0+3)/N
Ia_std = Ia0_g*0.4
bounds_for_Ia = np.tile([0.1/N, 100/N], M).reshape(M, 2)
S0_g = (S0-3)/N
S_std = Ia_std*np.sqrt(3)
bounds_for_S = np.array([(1/N, f-1/N) for f in fi]).reshape(M, 2)
# optimisation parameters
ftol = 1e-5 # the relative tol in (-logp)
# set up bounds, guess and stds for the rest of the params
keys = ['betaIa', 'betaIs', 'gIa', 'gIs']
bounds = np.array([betaIa_b, betaIs_b, gIa_b, gIs_b,
*bounds_for_S, *bounds_for_Ia])
guess = np.array([betaIa_g, betaIs_g, gIa_g, gIs_g, *S0_g, *Ia0_g])
stds = np.array([betaIa_std, betaIs_std, gIa_std, gIs_std, *S_std, *Ia_std])
# set up fltr for initial conditions because they are constraint by the observed
init_fltr = np.repeat([True, True, False], M)
params = estimator.latent_infer_parameters(keys, init_fltr, guess, stds, obs, fltr, Tf_inference, Nf_inference,
contactMatrix, bounds,
global_max_iter=10, global_ftol_factor=1e3,
verbose=True, ftol=ftol)
# +
print("True parameters:")
print(inference_parameters)
print("\nInferred parameters:")
best_estimates = estimator.fill_params_dict(keys, params)
print(best_estimates)
# +
Nf = 101
Tf = Nf-1
partial_inits = params[len(keys):]
x0 = estimator.fill_initial_conditions(partial_inits, obs[0], init_fltr, fltr)
model = pyross.deterministic.Spp(model_spec, best_estimates, int(M), fi)
det_data = model.simulate(x0, contactMatrix, Tf, Nf)
x_det = det_data['X']
x = data['X']/N
plt.rcParams.update({'font.size': 12})
plt.plot(np.sum(x_det[:, :M], axis=1), label='Inferred S')
plt.plot(np.sum(x[:, :M], axis=1), label='True S')
plt.plot(np.sum(x_det[:, M:2*M], axis=1), label='Inferred Ia')
plt.plot(np.sum(x[:, M:2*M], axis=1), label='True Ia')
plt.plot(np.sum(x_det[:, 2*M:3*M], axis=1), label='Inferred Ia')
plt.plot(np.sum(x[:, 2*M:3*M], axis=1), label='True Ia')
plt.axvspan(0, Nf_inference,
label='Used for inference',
alpha=0.3, color='dodgerblue')
plt.legend()
plt.show()
# -
| examples/inference/nbtests/ex_Spp_latent_inference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python385jvsc74a57bd079b153956a97866037adaaeff47af5e95ee39f609d617a5d46c76aae447d8416
# ---
import seaborn as sns
import json
import pandas as pd
# ### Load json log files from experiments
# + tags=[]
ROOT_PATH="../../outputs/cifar10-CC-HP-explore/{atk}_tau{tau}_l{n_iter}_seed{seed}/stats"
def read_json(path):
print(path)
validation = []
with open(path, "r") as f:
for line in f:
line=line.strip().replace("'", '"')
data = json.loads(line)
if data['_meta']['type'] == 'validation':
validation.append(data)
return validation
# Loop over hyperparameters
data_collection = {}
# TODO:
# for seed in [0, 1, 2]:
for seed in [0]:
for attack in ["LF", "BF"]:
for tau in [1e-1, 1e1, 1e3]:
for inner in [1, 3, 5]:
path = ROOT_PATH.format(atk=attack, tau=tau, n_iter=inner, seed=seed)
data_collection[path] = read_json(path)
# + tags=[]
# Example of data entries
for k, v in data_collection.items():
for _, item in zip(range(3), v):
print(item)
break
# +
# Generate a dataframe
def transform(entry, atk, tau, n_iter, seed):
# {'_meta': {'type': 'validation'}, 'E': 1, 'Length': 9984, 'Loss': 2.0887, 'top1': 17.8986}
return {
'Epochs': entry['E'],
'Accuracy (%)': entry['top1'],
'Attack': atk,
'#InnerLoops': n_iter,
'seed': seed,
r'$\tau$': tau
}
df = []
# TODO: for seed in [0, 1, 2]:
for seed in [0]:
for attack in ["LF", "BF"]:
for tau in [1e-1, 1e1, 1e3]:
for inner in [1, 3, 5]:
path = ROOT_PATH.format(atk=attack, tau=tau, n_iter=inner, seed=seed)
# Elements of `validation_entries`: {'_meta': {'type': 'validation'}, 'E': 1, 'Length': 9984, 'Loss': 2.0887270050171094, 'top1': 17.89863782051282}
validation_entries = data_collection[path]
df += list(map(lambda x: transform(x, atk=attack, tau=tau, n_iter=inner, seed=seed), validation_entries))
# -
df = pd.DataFrame(df)
# NOTE: here we exclude ci (confidence interval) for speed.
g = sns.lineplot(data=df, x="Epochs", y="Accuracy (%)", hue="#InnerLoops", style=r"$\tau$", ci=None)
df_final = df[df['Epochs'] == 200]
df_final.head()
df_final
def dataframe_to_latextable(df):
pass
| momentum-robustness/cifar10-CC-HP-explore.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_mxnet_p36)
# language: python
# name: conda_mxnet_p36
# ---
# # Using the Apache MXNet Module API with SageMaker Training and Batch Transformation
#
# *(This notebook was tested with the "Python 3 (Data Science)" kernel.)*
#
# The SageMaker Python SDK makes it easy to train MXNet models and use them for batch transformation. In this example, we train a simple neural network using the Apache MXNet [Module API](https://mxnet.incubator.apache.org/api/python/module.html) and the MNIST dataset. The MNIST dataset is widely used for handwritten digit classification, and consists of 70,000 labeled 28x28 pixel grayscale images of hand-written digits. The dataset is split into 60,000 training images and 10,000 test images. There are 10 classes (one for each of the 10 digits). The task at hand is to train a model using the 60,000 training images and subsequently test its classification accuracy on the 10,000 test images.
# ### Setup
#
# First, we define a few variables that are be needed later in the example.
# + isConfigCell=true
from sagemaker import get_execution_role
from sagemaker.session import Session
sagemaker_session = Session()
region = sagemaker_session.boto_session.region_name
sample_data_bucket = 'sagemaker-sample-data-{}'.format(region)
# S3 bucket for saving files. Feel free to redefine this variable to the bucket of your choice.
bucket = sagemaker_session.default_bucket()
# Bucket location where your custom code will be saved in the tar.gz format.
custom_code_upload_location = 's3://{}/mxnet-mnist-example/code'.format(bucket)
# Bucket location where results of model training are saved.
model_artifacts_location = 's3://{}/mxnet-mnist-example/artifacts'.format(bucket)
# IAM execution role that gives SageMaker access to resources in your AWS account.
# We can use the SageMaker Python SDK to get the role from our notebook environment.
role = get_execution_role()
# -
# ### Training and inference script
#
# The `mnist.py` script provides all the code we need for training and and inference. The script also checkpoints the model at the end of every epoch and saves the model graph, params and optimizer state in the folder `/opt/ml/checkpoints`. If the folder path does not exist then it skips checkpointing. The script we use is adaptated from the Apache MXNet [MNIST tutorial](https://mxnet.incubator.apache.org/tutorials/python/mnist.html).
# !pygmentize mnist.py
# ### SageMaker's MXNet estimator class
#
# The SageMaker ```MXNet``` estimator allows us to run single machine or distributed training in SageMaker, using CPU or GPU-based instances.
#
# When we create the estimator, we pass in the filename of our training script, the name of our IAM execution role, and the S3 locations we defined in the setup section. We also provide a few other parameters. ``train_instance_count`` and ``train_instance_type`` determine the number and type of SageMaker instances that are used for the training job. The ``hyperparameters`` parameter is a ``dict`` of values that is passed to your training script -- you can see how to access these values in the ``mnist.py`` script above.
#
# For this example, we choose one ``ml.m4.xlarge`` instance for our training job.
# +
from sagemaker.mxnet import MXNet
mnist_estimator = MXNet(entry_point='mnist.py',
role=role,
output_path=model_artifacts_location,
code_location=custom_code_upload_location,
train_instance_count=1,
train_instance_type='ml.m4.xlarge',
framework_version='1.6.0',
py_version='py3',
hyperparameters={'learning-rate': 0.1})
# -
# ### Running a training job
#
# After we've constructed our `MXNet` object, we can fit it using data stored in S3. Below we run SageMaker training on two input channels: train and test.
#
# During training, SageMaker makes this data stored in S3 available in the local filesystem where the `mnist.py` script is running. The script then simply loads the train and test data from disk.
# +
# %%time
train_data_location = 's3://{}/mxnet/mnist/train'.format(sample_data_bucket)
test_data_location = 's3://{}/mxnet/mnist/test'.format(sample_data_bucket)
mnist_estimator.fit({'train': train_data_location, 'test': test_data_location})
# -
# ### SageMaker's transformer class
#
# After training, we use our `MXNet` estimator object to create a `Transformer` by invoking the `transformer()` method. This method takes arguments for configuring our options with the batch transform job; these do not need to be the same values as the one we used for the training job. The method also creates a SageMaker Model to be used for the batch transform jobs.
#
# The `Transformer` class is responsible for running batch transform jobs, which deploys the trained model to an endpoint and send requests for performing inference.
transformer = mnist_estimator.transformer(instance_count=1, instance_type='ml.m4.xlarge')
# ### Running a batch transform job
#
# Now we can perform some inference with the model we've trained by running a batch transform job. The request handling behavior during the transform job is determined by the `mnist.py` script.
#
# For demonstration purposes, we're going to use input data that contains 1000 MNIST images, located in the public SageMaker sample data S3 bucket. To create the batch transform job, we simply call `transform()` on our transformer with information about the input data.
# +
input_file_path = 'batch-transform/mnist-1000-samples'
transformer.transform('s3://{}/{}'.format(sample_data_bucket, input_file_path), content_type='text/csv')
# -
# Now we wait for the batch transform job to complete. We have a convenience method, `wait()`, that blocks until the batch transform job has completed. We call that here to see if the batch transform job is still running; the cell finishes running when the batch transform job has completed.
transformer.wait()
# ### Downloading the results
#
# The batch transform job uploads its predictions to S3. Since we did not specify `output_path` when creating the Transformer, one was generated based on the batch transform job name:
print(transformer.output_path)
# The output here will be a list of predictions, where each prediction is a list of probabilities, one for each possible label. Since we read the output as a string, we use `ast.literal_eval()` to turn it into a list and find the maximum element of the list gives us the predicted label. Here we define a convenience method to take the output and produce the predicted label.
# +
import ast
def predicted_label(transform_output):
output = ast.literal_eval(transform_output)
probabilities = output[0]
return probabilities.index(max(probabilities))
# -
# Now let's download the first ten results from S3:
# +
import json
from sagemaker.s3 import S3Downloader
predictions = []
for i in range(10):
file_key = '{}/data-{}.csv.out'.format(transformer.output_path, i)
output = S3Downloader.read_file(file_key)
predictions.append(predicted_label(output))
# -
# For demonstration purposes, we also download and display the corresponding original input data so that we can see how the model did with its predictions:
# +
import matplotlib.pyplot as plt
import numpy as np
plt.rcParams['figure.figsize'] = (2,10)
def show_digit(img, caption='', subplot=None):
if subplot == None:
_,(subplot) = plt.subplots(1,1)
imgr = img.reshape((28,28))
subplot.axis('off')
subplot.imshow(imgr, cmap='gray')
plt.title(caption)
for i in range(10):
input_file_name = 'data-{}.csv'.format(i)
input_file_uri = 's3://{}/{}/{}'.format(sample_data_bucket, input_file_path, input_file_name)
input_data = np.fromstring(S3Downloader.read_file(input_file_uri), sep=',')
show_digit(input_data)
# -
# Here, we can see the original labels are:
#
# ```
# 7, 2, 1, 0, 4, 1, 4, 9, 5, 9
# ```
#
# Now let's print out the predictions to compare:
print(predictions)
| aws_sagemaker_studio/frameworks/mxnet_mnist/mxnet_mnist_with_batch_transform.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Functional Test Documentation
# Generates documentation for fastai's functional tests
# + hide_input=true
from fastai.gen_doc.nbdoc import *
from fastai.gen_doc.nbtest import *
# -
# ## Find tests for any function/class
# [`show_test`](/gen_doc.nbtest.html#show_test) and [`doctest`](/gen_doc.nbtest.html#doctest) searches for any implemented tests for a given fastai class or function
#
# For test writers:
# * Use this module to search for tests and get a better idea on which parts of the fastai api need more functional tests
#
# For fastai users:
# * Usage is similar to [`nbdoc.show_doc`](/gen_doc.nbdoc.html#show_doc) and [`nbdoc.doc`](/gen_doc.nbdoc.html#doc).
# * It's here to help you find associated tests for a given function can help understand usage.
#
# ## Usage:
# + hide_input=true
show_doc(show_test)
# -
# **Show tests from function**
from fastai.basic_train import Learner
show_test(Learner.fit)
# **Show tests from a Class**
from fastai.basic_data import DataBunch
show_test(DataBunch)
from fastai.text.data import TextList
show_test(TextList)
# ## Different test types
# Above, you will see 3 different test types: `This tests`, `Direct tests`, and `Related tests`
#
# * `This tests` - Searches for function matches in `test_api_db.json`. This json file is populated from `doctest.this_tests` calls.
# * `Direct tests` - Searches for any test function whose name contains the fastai function call
# * `Related tests` - Returns any test function where the fastai function in called inside the body
# ## Show in notebook inline:
# + hide_input=true
show_doc(doctest)
# -
# ## Internal search methods
# + hide_input=true
show_doc(find_dir_tests)
# + hide_input=true
show_doc(lookup_db)
# + hide_input=true
show_doc(find_test_matches)
# + hide_input=true
show_doc(find_test_files)
# + hide_input=true
show_doc(direct_test_match)
# + hide_input=true
show_doc(fuzzy_test_match)
# -
# ## Undocumented Methods - Methods moved below this line will intentionally be hidden
| docs_src/gen_doc.nbtest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from PIL import Image
from pycocotools.coco import COCO
from lxml import etree
import cv2
import sys
import json
import numpy as np
import matplotlib.pyplot as plt
sys.path.insert(0, '../libs')
from imantics import *
# -
category = Category("Test")
annotation = Annotation.from_bbox(None, category, [0, 10, 20, 30])
image = Image()
image.add(annotation)
| examples/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
df=pd.read_csv("training_twitter_x_y_train.csv")
df1=pd.read_csv("test_twitter_x_test.csv")
print(df.info())
print(df.airline_sentiment.value_counts())
df.drop(['tweet_id','airline','airline_sentiment_gold','name','negativereason_gold','retweet_count','tweet_coord','tweet_created','tweet_location','user_timezone'],axis=1,inplace=True)
df1.drop(['tweet_id','airline','airline_sentiment_gold','name','negativereason_gold','retweet_count','tweet_coord','tweet_created','tweet_location','user_timezone'],axis=1,inplace=True)
# ## DATA CLEANING
from nltk.tokenize import WordPunctTokenizer
tok = WordPunctTokenizer()
import re
from bs4 import BeautifulSoup
# +
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
# -
Training=df.text
Testing=df1.text
Train_result=[]
Test_result=[]
for i in Training:
Train_result.append((cleaner(i)))
for i in Testing:
Test_result.append((cleaner(i)))
# ## Testing Score
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.model_selection import GridSearchCV
from sklearn.svm import SVC
from sklearn.neighbors import KNeighborsClassifier
# ## Vectorizer
Tfid_Vectorizer=TfidfVectorizer(ngram_range=(1,2),max_features=2000)
X_train_feature=Tfid_Vectorizer.fit_transform(Train_result)
X_test_feature=Tfid_Vectorizer.transform(Test_result)
print(X_train_feature.shape)
print(X_test_feature.shape)
# ## Logistic
#
from sklearn.linear_model import LogisticRegression
abc=LogisticRegression(max_iter=20000)
#grid = {'C': [1e2,1e3, 5e3, 1e4, 5e4, 1e5],}
# +
#abc=GridSearchCV(clf,grid)
# -
abc.fit(X_train_feature,df.airline_sentiment)
Sentiment1=abc.predict(X_test_feature)
np.savetxt('SentimentSVM.csv',Sentiment1,fmt="%s")
abc.score(X_train_feature,df.airline_sentiment)
| TwitterSentimentAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## <NAME>
#
# ### SDSC2001 Python for Data Science
# ### Short summary
# - The goal of the project is a fraud detection in credit card transactions
# - The dataset of 284,807 transactions is used. The dataset is highly unbalanced with only 0.172% fraud data
# - 1-part is Data Exploration, including checking distribution of values, and data visualization
# - 3-part is Data Reduction using PCA, T-SNE, Spectral Embedding algorithms and data visualization on 2d plot
# - 4-part is about creating Classification model using Logistic Regression, Random Forest, Ada Boost algorithms and checking the performance (confusion matrix, ROC curve)
# #### Content
# The dataset contains transactions made by credit cards in September 2013 by european cardholders. Transactions occurred in two days, where we have 492 frauds out of 284,807 transactions. The dataset is highly unbalanced, as the positive class (frauds) account for 0.172% of all transactions.
#
# It contains numerical input variables V1-V28 which are the result of a Principal Component Analysis (PCA) transformation, as original features are not provided due to confidentiality issues. Features that have not been transformed with PCA are 'Time' and 'Amount'. 'Time' contains the seconds elapsed between each transaction and the first transaction in the dataset. 'Amount' denotes the transaction Amount. 'Class' is the response variable (labelled outcome) and it takes value 1 in case of fraud and 0 otherwise.
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
df_train = pd.read_csv('creditcard_train.csv')
# df_test = pd.read_csv('creditcard_test.csv')
# ### Data Exploration
# ## `Class` is a target feature
# ### 0 - Non-fraud transaction
# ### 1 - Fraud transaction
df_train.head()
# ### Training data has 284,657 rows
df_train.shape
# ### Test data has 150 rows
df_test.shape
# ## Standard deviation of features V1-V28 are between 2 and 0.33, while mean is around 0. Normalization or standardization isn't necessary for these features
df_train.describe()
# ### Fraud cases make only 0.156% (442 rows) of all training data. Most of the data is non-fraud cases. This shows that dataset is highly unbalanced.
sns.countplot(data=df_train, x='Class')
plt.title("Distribution of Class (0 - normal, 1 - fraud)");
df_train['Class'].value_counts()
# +
all_rows = df_train.shape[0]
fraud_rows = (df_train['Class'] == 1).sum()
print("Percentage of fraud rows: ", fraud_rows / all_rows * 100)
# -
# ### Distribution of Time
plt.hist(df_train['Time'], bins=100)
plt.title('Distribution of Time')
plt.xlabel('Time');
# ### Distribution of Amount
# #### Amount is positively skewed
plt.hist(df_train['Amount'], bins=50)
plt.title('Distribution of Amount')
plt.xlabel('Amount');
V_features = ['V{}'.format(i) for i in range(1,29)]
# ## Missing values
# ### There are missing values in V22 and V23. And they make 0.28% of all data
df_train.isna().sum()
(520 + 278) / df_train.shape[0] * 100
# ### There is no correlated features with V22 and V23, so that, we cannot do imputation
plt.figure(figsize=(10, 9))
sns.heatmap(df_train.corr(), linewidth=0.05)
plt.title("Correlation matrix");
# ### There is no fraud cases in rows that contain missing values. It means if I drop them I don't lose any valueable information, which is fraud case
# Number of fraud cases in rows that V22 value is missing
(df_train.loc[df_train['V23'].isna(), 'Class'] == 1).sum()
# Number of fraud cases in rows that V23 value is missing
(df_train.loc[df_train['V22'].isna(), 'Class'] == 1).sum()
# ## Removed Missing Values
df_train = df_train.dropna()
# ## Dealing with Outliers
# ### IQR test
# ### Outliers of IQR test make 21% of data.
# +
q25 = df_train[V_features].quantile(q=0.25)
q75 = df_train[V_features].quantile(q=0.75)
IQR = q75 - q25
df_train['Outlier'] = ((df_train[V_features] < (q25 - 2.5 * IQR)) |
(df_train[V_features] > (q75 + 2.5 * IQR))).sum(axis=1) > 0
# -
(df_train['Outlier'] == True).sum() / df_train.shape[0]
# ### The number of fraud cases
fraud = df_train[df_train['Class'] == 1]
normal = df_train[df_train['Class'] == 0]
fraud.shape[0]
# ### The number of outliers in fraud data
fraud['Outlier'].sum()
# ### If I delete outliers, I lose 92% of fraud data, which is rare in the given dataset. It shows outliers capture important information about fraud. So, we keep all outliers
print(406 / 442 * 100)
# Outlier is not necessary any more. So, I drop it
df_train = df_train.drop('Outlier', axis=1)
# ### Module 2: Data Visualization
# What are the distributions of variables? Are the variables distributed differently in the Fraud group and in the Normal group? Use tables and/or figures to visualize your exploration and findings. Present no more than 5 figures/tables that you think are important, and you may state in your findings that other variables a, b, c have similar patterns.
# ### Mean values of many features in fraud cases are considerably dfiferent than normal cases. it shows different distributions. Some of them are visualized as example
pd.DataFrame([fraud.mean(), normal.mean()], index=['Fraud (mean)', 'Normal (mean)']).T
# ### Visualization of features with different distribution in fraud and normal cases
plt.hist(df_train.loc[df_train['Class']==0, 'V4'], bins=range(-5, 15),density=True, alpha=0.5, label='Non-fraud')
plt.hist(df_train.loc[df_train['Class']==1, 'V4'], bins=range(-5, 15),density=True, alpha=0.5, label='Fraud')
plt.legend()
plt.title("Frequency density of V4");
plt.hist(df_train.loc[df_train['Class']==0, 'V11'], bins=range(-5, 13),density=True, alpha=0.5, label='Non-fraud')
plt.hist(df_train.loc[df_train['Class']==1, 'V11'], bins=range(-5, 13),density=True, alpha=0.5, label='Fraud')
plt.legend()
plt.title("Frequency density of V11");
plt.hist(df_train.loc[df_train['Class']==0, 'V12'], bins=range(-20, 5),density=True, alpha=0.5, label='Non-fraud')
plt.hist(df_train.loc[df_train['Class']==1, 'V12'], bins=range(-20, 5),density=True, alpha=0.5, label='Fraud')
plt.legend()
plt.title("Frequency density of V12");
plt.hist(df_train.loc[df_train['Class']==0, 'V14'], bins=range(-20, 10),density=True, alpha=0.5, label='Non-fraud')
plt.hist(df_train.loc[df_train['Class']==1, 'V14'], bins=range(-20, 10),density=True, alpha=0.5, label='Fraud')
plt.legend()
plt.title("Frequency density of V14");
# ### Most of features' distribution is close to normal distribution (Gaussian). Especially, distribution of fraud data is like normal distribution
# ### Variables V4, V10, V11, V12, V14, V16, V17 are distributed differently in fraud and normal data
#
# -----
# ### Module 3: Dimension Reduction
# Apply unsupervised learning methods to achieve dimension reduction. Visualize and interpret the results. Any dimension reduction algorithms can be used.
# ### Standardization
# Values of Time and Amout varies a lot comparing to other features. Due to Time and Amount's large variance, dimention reduction algorithms will capture mostly Time and Amount. Therefore, I need to scale them
# #### When training data is scaled, test data should be scaled too
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
df_train['Amount_scaled'] = scaler.fit_transform(df_train[['Amount']])
df_test['Amount_scaled'] = scaler.transform(df_test[['Amount']])
df_train['Time_scaled'] = scaler.fit_transform(df_train[['Time']])
df_test['Time_scaled'] = scaler.transform(df_test[['Time']])
df_train = df_train.drop(['Amount', 'Time'], axis=1)
df_test = df_test.drop(['Amount', 'Time'], axis=1)
X = df_train.drop(['Class'], axis=1)
y = df_train['Class']
# ### Under-sampling
#
# In classification part (module 4), I will use under-sampling. In order to test how the same data can be classified using dimention reduction algorithms, I implement under-sampling in this part (module 3).
# In under-sampling, I create a new dataframe with the same number of fraud and non-fraud cases. The number of fraud cases is 442, so I take 442 rows of non-fraud data.
# +
# shuffling data
shuffled_df_train = df_train.sample(frac=1, random_state=1)
fraud_df = shuffled_df_train.loc[df_train['Class'] == 1]
normal_df = shuffled_df_train.loc[df_train['Class'] == 0][:442]
new_df_train = pd.concat([fraud_df, normal_df]).sample(frac=1, random_state=2)
# -
new_df_train.head()
new_df_train.shape
X_train = new_df_train.drop('Class', axis=1)
y_train = new_df_train['Class']
# ### PCA ( Principal component analysis )
from sklearn.decomposition import PCA
X_pca = PCA(n_components=2, random_state=1).fit_transform(X_train)
sns.scatterplot(x=X_pca[:,0], y=X_pca[:,1], hue=y_train)
plt.title('PCA');
# ### TSNE ( t-distributed Stochastic Neighbor Embedding)
# ### t-SNE is recommended in sklearn documentation
# "For visualization purpose (which is the main use case of t-SNE), using the Barnes-Hut method is strongly recommended". [link](https://scikit-learn.org/stable/modules/manifold.html)
from sklearn.manifold import TSNE
tsne = TSNE(n_components=2, perplexity=35, random_state=1)
X_tsne = tsne.fit_transform(X_train)
# ### Clusters in t-SNE are clearly separeted
sns.scatterplot(x=X_tsne[:,0], y=X_tsne[:,1], hue=y_train)
plt.title('t-SNE');
# ### Spectral Embedding
from sklearn.manifold import SpectralEmbedding
mds = SpectralEmbedding(n_components=2, gamma=10)
X_se = mds.fit_transform(X_train)
sns.scatterplot(x=X_se[:,0], y=X_se[:,1], hue=y_train)
plt.title('Spectral Embedding');
# ### As shown in plots using dimension reduction methods, fraud and non-fraud cases are clustered separetely. It means the data has good signal for classification and models will be able to detect easily.
# ------
# ### Module 4: Classification
# Choose three classification methods and build classification models, using 5-fold cross-validation. <br>
#
# Further, load `creditcard_test.csv` and use it as the test dataset to evaluate your models and compare their performance. Consider visualizing the results, and evaluate the results using metrics such as the overall accuracy and the confusion matrix. <br>
#
# Hint: because the dataset is highly unbalanced, consider undersampling, i.e. randomly select N normal observations of the same size as the N fraud observations, and use N+N=2N samples for model fitting and training.
# +
###pick 3 classification methods, and methods not in the below list can also be used; you can also pick more than three methods
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
from sklearn.metrics import roc_curve, auc
from sklearn.metrics import roc_auc_score
from sklearn.metrics import plot_roc_curve
# -
X_test = df_test.drop('Class', axis=1)
y_test = df_test['Class']
# ### Undersampling the imbalanced data
# +
# shuffling data
df_train = df_train.sample(frac=1, random_state=1)
# take fraud data
fraud_df = df_train.loc[df_train['Class'] == 1]
# select the same number of normal data
normal_df = df_train.loc[df_train['Class'] == 0][:442]
new_df_train = pd.concat([fraud_df, normal_df]).sample(frac=1, random_state=2)
X_train = new_df_train.drop('Class', axis=1)
y_train = new_df_train['Class']
# -
y_train.value_counts()
# ### Logistic Regression
# hyperparameter tuning with Grid Search CV.
log_reg_params = {'C': [1, 3, 10, 100]}
log_res = LogisticRegression(max_iter=1000)
grid_log_reg = GridSearchCV(log_res, log_reg_params, cv=5)
grid_log_reg.fit(X_train, y_train);
# #### Best score in CV
grid_log_reg.best_score_
y_logreg = grid_log_reg.predict(X_test)
# #### Accuracy of the model on test set
accuracy_score(y_test, y_logreg)
sns.heatmap(confusion_matrix(y_test, y_logreg), annot=True, annot_kws={'fontsize':15})
plt.xlabel('Predicted')
plt.ylabel('True')
plt.title('Confusion matrix of Logistic Regression');
print(classification_report(y_test, y_logreg))
# ### SVC ( Support Vector Classification)
# +
svc_params = {'C': [0.1, 0.5, 1, 5, 10],
'kernel': ['rbf', 'poly', 'sigmoid', 'linear']}
svc = SVC()
grid_svc = GridSearchCV(svc, svc_params, cv=5)
grid_svc.fit(X_train, y_train);
# -
grid_svc.best_score_
y_svc = grid_svc.predict(X_test)
# #### Accuracy of SVC on test set
accuracy_score(y_test, y_svc)
sns.heatmap(confusion_matrix(y_test, y_svc), annot=True, annot_kws={'fontsize':15})
plt.xlabel('Predicted')
plt.ylabel('True')
plt.title('Confusion matrix of SVC');
print(classification_report(y_test, y_svc))
# ### Random Forest Classifier
# +
rf_params = {'n_estimators': [100, 300, 500],
'max_depth': [5,7,9],
'max_features': ['auto', 'sqrt'],
'min_samples_leaf': [1, 2, 4]}
rf = RandomForestClassifier()
grid_rf = GridSearchCV(rf, rf_params, cv=5)
grid_rf.fit(X_train, y_train);
# -
grid_rf.best_score_
grid_rf.best_params_
y_rf = grid_rf.predict(X_test)
# #### Accuracy of Random Forest model on test set
accuracy_score(y_test, y_rf)
sns.heatmap(confusion_matrix(y_test, y_rf), annot=True, annot_kws={'fontsize':15})
plt.xlabel('Predicted')
plt.ylabel('True')
plt.title('Confusion matrix of Random Forest');
print(classification_report(y_test, y_rf))
# ### Ada Boost Classifier
# +
ada_params = {'n_estimators': [100, 300, 500]}
ada = AdaBoostClassifier()
grid_ada = GridSearchCV(ada, ada_params, cv=5)
grid_ada.fit(X_train, y_train);
# -
grid_ada.best_score_
y_ada = grid_ada.predict(X_test)
# #### Accuracy of Ada Boost Classifier model on test set
accuracy_score(y_test, y_ada)
sns.heatmap(confusion_matrix(y_test, y_ada), annot=True, annot_kws={'fontsize':15})
plt.xlabel('Predicted')
plt.ylabel('True')
plt.title('Confusion matrix of Ada Boost Classifier');
print(classification_report(y_test, y_ada))
# ### ROC curve and AUC score
# ### AUC score of all models are close to 1
# +
y_log_score = grid_log_reg.decision_function(X_test)
y_svc_score = grid_svc.decision_function(X_test)
y_ada_score = grid_ada.decision_function(X_test)
print("AUC scores")
print("Logistic Regression: {:.3f}".format(roc_auc_score(y_test, y_log_score)))
print("SVC: {:.3f}".format(roc_auc_score(y_test, y_svc_score)))
print("Ada Boost Classfier: {:.3f}".format(roc_auc_score(y_test, y_ada_score)))
# -
# ### Logistic Regression showed the best result on accuracy. Let's look at deeper
plot_roc_curve(grid_log_reg, X_test, y_test, label='Logistic Regression')
plt.title("ROC curve of Logistic Regression");
# ### As we see on graph above, ROC curve captures almost all space
# ## Check Logistic Regression on all training data without under-sampling
X_all = df_train.drop('Class', axis=1)
y_all = df_train['Class']
X_all.shape
pred_all = grid_log_reg.predict(X_all)
# ### Accuracy of the model on all training data
print("Accuracy on all training data: {:.3f}".format(accuracy_score(y_all, pred_all)))
plot_roc_curve(grid_log_reg, X_all, y_all, label='Logistic Regression (AUC=0.99)')
plt.title("ROC curve of Logistic Regression on all training data");
# ### The model misclassified many normal transactions as fraud transaction
sns.heatmap(confusion_matrix(y_all, pred_all), annot=True, annot_kws={'fontsize':15})
plt.xlabel('Predicted')
plt.ylabel('True')
plt.title('Confusion matrix on all training data');
# ### Module 5: Summary
# Summarize your findings and draw conclusions.
# #### My findings:
#
# 1. The given training data is imbalanced. Fraud transactions make only 0.16% of all data. Undersampling helped to fix this issue
# 2. I removed rows that have missing values because none of them are fraud cases. So, I don't lose valueable information by dropping them.
# 3. Distribution of some features such as V4, V11, V12, V14, V17, is different in fraud cases than the normal cases.
# 4. Fraud transactions contain outliers because they are different the normal transactions. Therefore, I didn't delete outliers. By deleting outliers, we lose most of fraud data.
# 5. Dimension reduction algorithms like PCA, t-SNE showed that with the only 2 components on visualization we can notice the difference of fraud transactions. It's evidence that ML algorithms perform well.
# 6. All models have good accuracy score on test set with above 95%. The result:
# - Logistic Regression - 97.3%
# - SVC - 96.7%
# - Random Forest - 96%
# - Ada Boost Classifier - 96.7%
# 7. Logistic Regression showed the best results. its result:
# - precision - 96%
# - recall - 96%
#
# #### Conclusion
# Undersampling helped to deal with imbalance data. Logistic Regression showed the best result with accuracy 97% on test data, 96% on all training data. These models are deployable in real life. Although it has very high accuracy, it misclassified many normal transactions as fraud transaction. In the credit card fraud detection task, detecting fraud transaction is more important than detecting normal data. Because we can check normal transactions by contacting owners of credit cards. If we miss the fraud transaction, it will be impossible to fix it later.
#
# On the other hand, calling to credit card owners every time, when they make regular daily transactions, causes customer complaints and dissatisfaction. Also, banks would spend a lot of money on employees. So, we deal with it by increasing the accuracy of models to predict more data right. One possible way is to collect more fraud data and train models again. The given training data has only 442 fraud transaction, which is 0.156% of all data. If we increase the number of fraud transaction in the data, it could lead to even more accuracy. Another possible approch might be using over-sampling instead of under-sampling. In over-sampling we get more data to train models. I didn't use it as under-sampling technique is suggested to use in project.
#
| projects/Fraud-Detection/fraud-detection-python_course.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 align="center">Solutions and Hints<h1>
# ### Exercises 1
# 1. Produce $AE$ to $F$, make $EF = EA$, prove that $\triangle ACD \cong \triangle AFD$.
# 2. Prove that the sum of the lengths of any two sides of a triangle is greater than the twice of the length of the third side's median.
# 3. Let $C^\prime$ be the symmetry point of $C$ at $BA$ produced with regard to $AP$, join $PC^\prime$, prove that $PB + PC^\prime > BC^\prime$.
# 4. Draw altitude $CD$ to $AB$, prove that $\angle DCE = \angle MCE$.
# 5. Draw altitude $AD$ to $BC$, prove it with the Pythagorean Theorem.
# 6. Translate the segment between two opposite sides and map one endpoint to a vertex of the rectangle, then prove it with the Angle-side Inequalities Relationship in triangles.
# 7. Through $C$ draw $CM \| EA$ to meet $AB$ in $M$, prove that $CM$ is the perpendicular bisector of $BF$.
# 8. Join $AD$, prove it with the Mean Proportional Theorems in right triangles.
# 9. Let $ABC$ be the triangle, $H$ be its orthocenter, and $F$ be the symmetry point of $H$ with regard to $AC$, prove that $\angle AFB = \angle ACB$.
# 10. Join $PA$, $MA$, prove that $\angle BPA = \angle TPA = \angle BAP$.
# 11. Join $PB$, $PQ$, prove that $QH \| BP$; or join $AQ$, $BQ$, prove that $\angle HQA + \angle HAQ = 90 ^\circ$.
# 12. Draw the common tangent.
# 13. Draw the common tangent $PQ$ to meet $AD$ in $Q$, prove that $\angle A + \angle D = \angle BPC$.
# 14. Draw the common chord $AB$, join $BC$, $BD$, prove that $\angle CBD + \angle P = 180 ^\circ$.
# 15. Join $DE, DF$, since $\triangle BDF \sim \triangle ABC$, we have $\frac{BC}{AC} = \frac{BF}{DF}$; since $\triangle ABC \sim \triangle CDF$, we have $\frac{BC}{AC} = \frac{DF}{CF}$;
# since $\triangle ABC \sim \triangle ADE$, we have $\frac{BC}{AC} = \frac{DE}{AE} = \frac{CF}{AE}$. Multiply three equations to $ \frac{BC^3}{AC^3} = \frac{BF}{AE}$ .
# 16. Produce $BD$ to meet $AC$ in $E$.
# 17. Draw $FH \perp BC$ to meet $BC$ in $H$, prove that $AE = AF = FH$ and $\triangle AEG \cong \triangle FHC$.
# 18. Produce $FH$ to meet $AC$ produced in $G$, draw $CK \| AB$ to meet $FG$ in $K$, prove that $\triangle BFM \cong \triangle CKM$.
# 19. Produce $AF$ to meet $BC$ produced in $E$. Prove that $\triangle ADF \cong \triangle ECF$ and $\triangle BAE$ to be an isosceles. Then prove that $BF, AF$ are bisectors.
# 20. Draw an isosceles triangle and let $\angle B$ be equal to the exterior of its apex. There are multiple proof methods.
# 21. Join $MF$, $MF \| AE$, $\therefore$ $BG=GM, MF= \frac{1}{2} AE=2GE$, $\therefore$ $AE=2MF=4GE= \frac{4}{3} AG$, $\therefore$ $AG:MF = 3:2$, since $\triangle MFH \sim \triangle AGH$, $\therefore$ $GH:HM = AG:MF = 3:2$,
# <br>$\therefore$ $BG:GH:HM = 5:3:2$.
# 22. Join $PM, QM$, start from proving that $\triangle PDM \cong \triangle QEM$.
# 23. Produce $FE$ to meets $CB$ produced in $M$, hence $\triangle AEF \cong \triangle BEM$, $\therefore$ $BM = AF$. <br>Since $\triangle AFG \sim \triangle CMG, \frac{CG}{AG} = \frac{CM}{AF} = \frac{CM}{BM} = \frac{4}{1}$,
# $\therefore$ $\frac{AC}{AG} = \frac{5}{1}$.
# 24. Join $BD, \triangle AEH \sim \triangle ABD, EH \| BD$, and $EH = \frac{2}{3} BD$, similarly, $\triangle CGF \sim \triangle CDB, GF \| BD$, and $GF = \frac{2}{3} BD$, <br>$\therefore$ $EH \perp GF$, quadrilateral $EFGH$ is a parallelogram.
# <br>$\frac{S_{\triangle AEH}}{S_{\triangle ABD}} = (\frac{2}{3})^{2} = \frac{4}{9},\frac{S_{\triangle CGF}}{S_{\triangle CDB}} = \frac{4}{9}.$ <br> $\therefore S_{\triangle AEH} + S_{\triangle CGF} = \frac{4}{9} S_{\square ABCD}.$ <br>Similarly, $S_{\triangle BEF} + S_{\triangle DHG} = \frac{1}{9} S_{\square ABCD}$,
# <br>$\therefore$ $S_{\square EFGH} = (1 - \frac{4}{9} - \frac{1}{9})S_{\square ABCD} = \frac{4}{9}S_{\square ABCD}$.
# 25. Join $PG, GQ$, join $AP, AG, AQ$ and produce them to meet $BC$ in $M, N, L$, then $\frac{AP}{PM} = \frac{AG}{GN} = \frac{AQ}{QL} = \frac{2}{1}$, <br>$\therefore$ $PG \| BC, GQ \| BC$, $\therefore$ $P, G, Q$ are collinear.
# <br>Since $\frac{GP}{MN} = \frac{AG}{AN} = \frac{2}{3}, \frac{GQ}{NL} = \frac{2}{3}$, $\therefore$ $\frac{GP}{GQ} =\frac{MN}{NL}=\frac{BN-BM}{CN-CL} =\frac{BC-BD}{BC-CD} = \frac{CD}{BD}$.
# 26. Join $BE, AC$, prove that $\triangle ABF \sim \triangle ABE$.
# 27. Draw an auxiliary circle from $B, D, H, F$, prove that $\angle HFD = \angle HBD$ and $\angle ABE = \angle HDF$. Draw an auxiliary circle from $A, B, D, E$, prove that $\angle EBD = \angle EAD$ and $\angle ABE = \angle EDA$,
# then prove that $\triangle ADE \sim \triangle DFH$.
# 28. Join $BI, CI$, prove that $\triangle BEI \sim \triangle CFI$.
# 29. Draw the common tangent $PT$, join $PB$, prove that $\triangle APD \sim \triangle CPB$.
# 30. Draw the common tangent $AB$ of two the circles, if we prove that the third circle with the segment through two circle's centers as a diameter is tangent to $AB$,
# then we have proved that the distance from the mid-point of the segment through two centers to $AB$ is a half length of the segment of the two centers.
# 31. Join $CA, CB$, prove that $CA, CB$ are the bisectors of the interior and exterior vertex angle of $\triangle PCD$. Then prove it with the interior and exterior angle bisector theorems of triangles.
# 32. From $I$ draw $IO$ perpendiculiar to $GC$ produced, then $\triangle ICO \cong \triangle ABC$. Prove it with Pytheagrean theorem in Rt$\triangle IGO$.
# 33. Proof 1: Draw altitudes $AM, AN$ of triangles $\triangle ABF, \triangle ACE$, first prove that $\triangle ABF \cong \triangle ACE$,
# we have $BF = CE$. Then prove that $AM = AN$ using area formula, prove that $\triangle AMO \cong \triangle ANO$, we have $\angle AOF = \angle AOE$.
# <br>
# Proof 2: Prove that $\triangle ABF \cong \triangle ACE$, we have $\angle AFO = \angle ACO$ and $\angle AEO = \angle ABO$. $\therefore$ $A,O,C,F$ and $A,O,B,E$
# are four points concyclic, respectively, <br>$\therefore$ $\angle AOF = \angle ACF = 60 ^\circ, \angle AOE = \angle ABE = 60 ^\circ$.
# 34. Join $DM, MF, FN, ND$, prove that quadrilaterial $DMFN$ is a rhombus.
# 35. (1) Join $OB$, draw $OM \perp BC$, so $OM=\frac{1}{2}AH=\frac{1}{2}AO=\frac{1}{2}BO$, <br>$\therefore$ in $Rt\triangle OBM, \angle BOM = 60 ^\circ$, $\therefore$ $\angle BAC = \angle BOM = 60 ^\circ$.
# <br>
# (2)Draw $OM \perp BC$, join $BO$, so $OM = \frac{1}{2}AH = \frac{1}{2}BC = BM$; <br>$\therefore$ in $Rt \triangle OBM, \angle BOM = 45 ^\circ$, $\therefore$ $\angle BAC = \angle BOM = 45 ^\circ$.
# 36. Join $OD, BD, \angle B = \frac{1}{2} \angle AOD, \angle BDC = \angle ACD - \angle B = \angle AOD - \angle B = \frac{1}{2} \angle AOD$, $\therefore$ $\angle B = \angle BDC$, $\therefore$ $CB = CD$.
# 37. Join $OB, OC, OM, ON$, $\because$ $O, A, C, N$ are four points concyclic, $\therefore$ $\angle ACO = \angle ANO$. <br>Since $O, A, M, B$ are four points concyclic, $\therefore$ $\angle ABO = \angle AMO$, and $\angle ANO = \angle AMO$,
# <br>$\therefore$ $\triangle OBC$ is an isosceles, $\therefore$ $AB = AC$.
# 38. Join $ED, AC, DF, BC$, $\because C, D, B, F$ are four points concyclic, $\therefore$ $\angle FCB = \angle FDB$, $\therefore$ $C, D, A, E$ are four points concyclic, $\angle CAD = \angle CED$, and $\angle FCB = \angle CAD$,
# $\therefore$ $\angle FDB = \angle CED$, <br>since $\angle FBD = 90 ^\circ = \angle ECD$, $\therefore$ $\triangle FDB \sim \triangle ECD$, $\therefore$ $\frac{BD}{CE} = \frac{BF}{CD} = \frac{DF}{ED}$. (1)
# <br>
# similarly to prove, $\triangle CDF \sim \triangle AED, \therefore \frac{CF}{AD} = \frac{CD}{AE} = \frac{DF}{DE}$. (2)
# <br>
# from (1) and (2) we have $\frac{BD}{CE} = \frac{CF}{AD}$, $\therefore$ $CE \cdot CF = AD \cdot BD$, since $\frac{BF}{CD} = \frac{CD}{AE}$, $\therefore$ $CD ^2 = AE \cdot BF$.
# 39. Join $O _{1} O _{2}, O _{1} A, O _{2} B$, draw $O _{1} D \perp O _{2} B$ in $D$, meet $TC$ in $E$, hence $ET \| DO _{2}$, $\therefore$ $\frac{TE}{O _{2} D} = \frac{O _{1} T}{O _{1} O _{2}}$,
# That is $\frac{TC - O _{1} A}{O _{2} B - O _{1} A} = \frac{O _{1} T}{O _{1} T + O _{2} T}$, $\frac{TC - \frac{1}{2}d _{1}}{\frac{1}{2}d _{2} - \frac{1}{2}d _{1}} = \frac{\frac{1}{2}d _{1}}{\frac{1}{2}(d _{1} + d _{2})}$,
# $\frac{2TC - d _{1}}{d _{2} - d _{1}} = \frac{d _{1}}{d _{1} + d _{2}} = \frac{2TC}{2d _{2}} = \frac{TC}{d _{2}}$, $\therefore$ $d_{1} d _{2} = TC(d _{1} + d _{2})$, $\therefore$ $\frac{1}{TC} = \frac{1}{d _{1}} + \frac{1}{d _{2}}$.
| pp80-84.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Data Pre-processing
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from datetime import timedelta
df = pd.read_json('transactions.json',lines=True,orient='records')
# Replacing null values with NaN
df = df.replace(r'^\s*$', np.nan, regex=True)
# Dropping with features with zero entries
df.drop(['echoBuffer','merchantCity','merchantState','merchantZip','posOnPremises','recurringAuthInd'],
axis=1,inplace=True)
# Creating Date and Time Column
df['Date']=df['transactionDateTime'].apply(lambda x: x.split('T')[0])
df['Time']=df['transactionDateTime'].apply(lambda x: x.split('T')[1])
def concat(x):
y=str('T') + str(x)
return y
df.reset_index(drop=False,inplace=True)
df['transactionKey']=df['index'].apply(lambda x: concat(x))
df.drop(['index'],axis=1,inplace=True)
df.head()
# ## Identifying Reversed transactions
df['transactionType'].isnull().sum()
#Replacing missing values in transactionType as 'PURCHASE'
df['transactionType']=df['transactionType'].fillna(value='PURCHASE')
# Creating Timestamp column
df.Date=pd.to_datetime(df.Date)
df['Timestamp']=pd.to_datetime(df.Date.astype(str)+' '+df.Time.astype(str))
# Sorting data in ascending order with respect to customerId, merchantName, Timestamp
data=df[['customerId','merchantName','transactionAmount','transactionType','transactionKey','Timestamp']]
data=data.sort_values(by=['customerId','merchantName','Timestamp'],ascending=True)
data.head()
# Removing transactions with ADDRESS_VERIFICATION type since, there transaction amount is $0
data=data.reset_index(drop=True)
data=data[-(data.transactionType=='ADDRESS_VERIFICATION')]
data=data.reset_index(drop=True)
# #### Considering the purchase and reversal transaction are consecutive
reverse = data.loc[data['transactionType']=='REVERSAL']
reverse.head()
purchase_index = reverse.index.values.astype(int)-1
purchase = data.iloc[purchase_index]
purchase.head()
purchase_index=pd.DataFrame(purchase_index)
purchase_index.columns=['Purchase_Index']
reverse_index=pd.DataFrame(reverse_index)
reverse_index.columns=['Reverse_Index']
purchase_amount=pd.DataFrame(purchase['transactionAmount'].values)
purchase_amount.columns=['Purchase_Amount']
reverse_amount=pd.DataFrame(reverse['transactionAmount'].values)
reverse_amount.columns=['Reversal_Amount']
# Creating DataFrame to check reversal transactions
chk=pd.concat([purchase_index,reverse_index,purchase_amount,reverse_amount],axis=1)
chk.head()
chk['new'] = np.where((chk['Purchase_Amount'] == chk['Reversal_Amount']), True, False)
chk.head()
chk.new.value_counts()
# #### By checking the consecutive transactions, I was able to track 12665 reversal transactions.
tracked = chk.loc[chk['new']==True]
# #### Creating DataFrame for all tracked reversal transactions
track_index=np.concatenate((tracked['Purchase_Index'], tracked['Reverse_Index']))
track_reverse=data.iloc[track_index]
track_reverse=track_reverse.sort_values(by=['customerId','merchantName','Timestamp'],ascending=True)
track_reverse.head()
track_reverse.transactionType.value_counts()
# #### I was able to track 12665 reverse transactions out of the 20303 transactions
track_reverse.groupby(['transactionType']).sum()['transactionAmount']
reverse.groupby(['transactionType']).sum()['transactionAmount']
# #### The transaction amount of tracked reversal transactions is $1907433.62
# #### The overall transaction amount of reverse transactions is $2821792.5
# #### 7638 transactions were not tracked with transaction amount of $914,358.88
# ## Identifying Multi-swipe transactions
# #### To identify multi-swipes, I am initially sorting the DataFrame in ascending order with respect to cusromerId, merchantName and Timestamp. The sorted data is available from previous. Then, to identify the multi-swipe transaction, I subset data with all the duplicate transactions and then classify the multi-swipe transactions with time difference less than 180 seconds.
# Removing Reverse transactions
mst=data.reset_index(drop=True)
mst=mst[-(data.transactionType=='REVERSAL')]
mst=mst.reset_index(drop=True)
mst.head()
k=mst.loc[:,['customerId','merchantName','transactionAmount','transactionType']]
k.head()
# Finding duplicate transactions
duplicate = k.duplicated(keep=False)
Duplicate=pd.DataFrame(duplicate)
Duplicate.columns=['Duplicate']
a=pd.concat([mst,Duplicate],axis=1)
a.head()
# Subsetting the data with duplicate transactions only
b = a.loc[a['Duplicate']==True]
b.head()
# Calculating time difference between the transactions
b['difference']=b.Timestamp.diff()
b.head()
z = timedelta(0,0)
z
td = timedelta(0,180)
td
# Checking for timedifference less than 180 seconds
b['multi_swipe']=(b['difference']<td) & (b['difference']>z)
#Sub-setting data with multi-swipe transactions only
multi_swipe = b.loc[b['multi_swipe']==True]
multi_swipe.head()
multi_swipe.transactionType.value_counts()
# Calculating total mutli-swipe transaction amount leaving the first transaction
multi_swipe.sum()['transactionAmount']
# #### Total value of multi-swipe is 7474
# #### Total transaction amount of multi-swipe transactions is $1105654
| Data wrangling - Duplicate transactions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import pyrsa
# # Demo on unbalanced designs
# In this demo we will have a look at calculating RDMs from data which either contians different numbers of measurements per stimulus or where the measurements for different stimuli do not contain the same measurement channels. Such data is common in neural recordings where individual neurons may become unmeasureable during an experiment such that we do not have measurements of all neurons for all stimuli. A common reason for different numbers of repetition for a stimulus are trials which have to be removed due to artifacts or aborted experiments or because the experimentor did not have full control over which stimuli were shown.
# First we need some data to work on. This data is very small scale such that we can understand everything which happens. As balanced data we have two repetitions of three different stimuli, which are measured for 5 different measurement channels.
data_balanced_array = np.array([
[0.7, 0.8, 0.9, 1.0, 1.1],
[0.2, 1.8, 2.9, 1.0, 1.3],
[2.7, 0.8, 0.2, 1.2, 1.1],
[1.7, 0.5, 0.9, 1.5, 1.1],
[1.7, 2.8, 2.2, 1.2, 1.0],
[1.7, 0.5, 0.4, 1.4, 0.3],
])
runs = [0, 1, 0, 1, 0, 1]
stimulus = [0, 0, 1, 1, 2, 2]
descriptors = {'runs': runs, 'stimulus': stimulus}
data_balanced = pyrsa.data.Dataset(data_balanced_array, obs_descriptors=descriptors)
# For the unbalanced data, we can use similar data, but let's assume that we now measured the 0 stimulus only once and measured stimulus 1 three times. Also at two times we had to discard measurements from single channels due to technical problems:
data_unbalanced_array = np.array([
[0.7, 0.8, 0.9, 1.0, 1.1],
[0.2, 1.8, np.nan, 1.0, 1.3],
[2.7, 0.8, 0.2, 1.2, 1.1],
[1.7, 0.5, 0.9, 1.5, 1.1],
[1.7, 2.8, 2.2, 1.2, np.nan],
[1.7, 0.5, 0.4, 1.4, 0.3],
])
runs = [0, 0, 1, 2, 0, 1]
stimulus = [0, 1, 1, 1, 2, 2]
descriptors = {'runs': runs, 'stimulus': stimulus}
data_unbalanced = pyrsa.data.Dataset(data_unbalanced_array, obs_descriptors=descriptors)
# For the balanced data we can use the normal functions to get a RDM:
balanced_rdm = pyrsa.rdm.calc_rdm(data_balanced, descriptor='stimulus')
print(balanced_rdm)
# For the unbalanced data this does not work, because the missing values break the calculation. Also the different numbers of measurements are disregarded when we use the normal calc_rdm function which averages stimulus responses first:
unbalanced_rdm_broken = pyrsa.rdm.calc_rdm(data_unbalanced, descriptor='stimulus')
print(unbalanced_rdm_broken)
# Instead, we can use the slightly slower variant for unbalanced designs, which calculates the RDM correctly:
unbalanced_rdm = pyrsa.rdm.calc_rdm_unbalanced(data_unbalanced, descriptor='stimulus')
print(unbalanced_rdm)
# This RDM now contains valid values for the dissimilarities. Additionally it gained an rdm_descriptor called 'weights', which contains the weight each carried by each dissimilarity. The difference here are caused by different numbers of measurement channels and repetitions for the different stimuli.
# As a sanity check we can compute the RDM based on the balanced data with this additional method to check that it results in the same RDM:
# +
sanity_rdm = pyrsa.rdm.calc_rdm_unbalanced(data_balanced, descriptor='stimulus')
print(sanity_rdm)
print('The differences between the two methods for the three dissimilarities are:')
print(sanity_rdm.get_vectors() - balanced_rdm.get_vectors())
# -
# The two RDMs are indeed identical up to normal floating point errors.
#
# This concludes the basic introduction to calculating RDMs for unbalanced experiments.
# ## Implementation explanation
#
# You may ask yourself why there are separate functions for computing RDMs for balanced and unbalanced designs. The main reason for this is that the functions for balanced designs average the different measurements for the same stimulus first and then calculate dissimilarities of these average activations. Computing this average requires that the same channels were measured for all repetitions of the stimulus though, which breaks this computation, when measurements are missing.
#
# To generate a compatible method for computing the dissimilarities we decompose the formula into a sum of kernel evaluations, which we can average correctly. As an example, this is the decomposition for the euclidean distance $d_{xy}$ of two patterns with measured representations $x_i$ and $y_j$ for $i = 1 \dots N_x, j = 1 \dots N_y$, denoting the mean representations as $\bar{x}$ and $\bar{y}$ for a single measurement channel:
#
# \begin{eqnarray}
# d_{xy} &=& (\bar{x} - \bar{y}) ^2 = \left( \frac{1}{N_x} \sum_{i=1}^{N_x} x_i - \frac{1}{N_y} \sum_{j=1}^{N_y} y_j \right) ^2 \\
# &=& \frac{1}{N_x N_x} \sum_{i=1}^{N_x} \sum_{i'=1}^{N_x} x_i x_{i'} + \frac{1}{N_y N_y} \sum_{j=1}^{N_y} \sum_{j'=1}^{N_y} y_j y_{j'} - \frac{2}{N_x N_y} \sum_{i=1}^{N_x} \sum_{j=1}^{N_y} x_i y_j
# \end{eqnarray}
#
# To compute the dissimilarities for more than one measurement channel we simply need to comput this value per channel and average.
#
# As we now decomposed the function into the separate averages over individual stimulus measurements we can now deal with missing values by simply removing them from the average.
#
# The derivation for the mahalanobis distance is very similar except for adding the precision of the noise in.
#
# ### Crossvalidation
#
# For Crossvalidation, we only want to use products of differences from different runs. For the standard balanced version we get the following formulas:
#
# \begin{eqnarray}
# d'_{xy} &=& \frac{1}{N} \sum_{i} (x_i-y_i) \left(\frac{1}{N-1} \sum_{i'\neq i} (x_i - y_i) \right) \\
# &=& \frac{1}{N (N-1)} \sum_{i} \sum_{i'\neq i} x_i x_{i'} + \frac{1}{N (N-1)} \sum_{i} \sum_{i'\neq i} y_i y_{i'} - \frac{2}{N (N-1)} \sum_{i} \sum_{i' \neq i} x_i y_j
# \end{eqnarray}
#
# Thus, we can again split the calculation into sums over the individual pairs of measurements, which allows us to deal with missing measurements gracefully. The only addition we need to make is removing the terms from the sum which come from the same run/partition. Also, this extens completely to the Mahalanobis/CrossNobis.
# ### Poisson-KL based distances
# For using the Kullback–Leibler divergence between poisson distributions as a dissimilarity, this decomposition results in a different estimate than the original formulas, as derived here:
#
# \begin{eqnarray}
# d_{xy} &=& (\bar{x} - \bar{y}) (\log\bar{x} - \log\bar{y}) = \left( \frac{1}{N_x} \sum_{i=1}^{N_x} x_i - \frac{1}{N_y} \sum_{j=1}^{N_y} y_j \right) \left(\log\left[ \frac{1}{N_x} \sum_{i=1}^{N_x} x_i\right] - \log\left[\frac{1}{N_y} \sum_{j=1}^{N_y} y_j\right] \right) \\
# &=& \frac{1}{N_x} \sum_{i=1}^{N_x}x_i \log\left[\frac{1}{N_x}\sum_{i'=1}^{N_x} x_{i'}\right]
# + \frac{1}{N_y} \sum_{j=1}^{N_y} y_j \log\left[\frac{1}{N_y} \sum_{j'=1}^{N_y} y_{j'}\right]
# - \frac{1}{N_x} \sum_{i=1}^{N_x} x_i \log\left[\frac{1}{N_y}\sum_{j=1}^{N_y} y_j \right]
# - \frac{1}{N_y} \sum_{j=1}^{N_y} y_j \log\left[\frac{1}{N_x}\sum_{i=1}^{N_x} x_i \right] \\
# &\neq& \frac{1}{N_xN_x} \sum_{i=1}^{N_x} \sum_{i'=1}^{N_x} x_i \log x_{i'}
# + \frac{1}{N_y} \sum_{j=1}^{N_y}\sum_{j'=1}^{N_y} y_j \log y_{j'}
# - \frac{1}{N_x} \sum_{i=1}^{N_x}\sum_{j=1}^{N_y} x_i \log y_j
# - \frac{1}{N_y} \sum_{j=1}^{N_y}\sum_{i=1}^{N_x} y_j \log x_i
# \end{eqnarray}
#
# ,i.e. ultimately the problem is that the logarithm is non-linear such that taking the mean of logs is different from the log of the mean.
#
# We nonetheless implement the estimator written in the last row as an analog of the euclidean distances and think this is a sensible estimate, but this estimate is different from the estimate based on the mean firing rates. Thus, we see that for the poisson and poisson_cv distance estimation calc_rdm and calc_rdm_unbalanced give different results:
# +
balanced_rdm_poisson = pyrsa.rdm.calc_rdm(data_balanced, descriptor='stimulus', method='poisson')
sanity_rdm_poisson = pyrsa.rdm.calc_rdm_unbalanced(data_balanced, descriptor='stimulus', method='poisson')
print('calc_rdm result:')
print(balanced_rdm_poisson)
print('calc_rdm_unbalanced result:')
print(sanity_rdm_poisson)
print('The differences between the two methods for the three dissimilarities are:')
print(sanity_rdm_poisson.get_vectors() - balanced_rdm_poisson.get_vectors())
| demos/demo_unbalanced.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualizing Chipotle's Data
# This time we are going to pull data directly from the internet.
# Special thanks to: https://github.com/justmarkham for sharing the dataset and materials.
#
# ### Step 1. Import the necessary libraries
import pandas as pd
import collections
import matplotlib.pyplot as plt
from IPython.display import display
# set this so the graphs open internally
# %matplotlib inline
# ### Step 2. Import the dataset from this [address](https://raw.githubusercontent.com/justmarkham/DAT8/master/data/chipotle.tsv).
# ### Step 3. Assign it to a variable called chipo.
chipo = pd.read_csv("https://raw.githubusercontent.com/justmarkham/DAT8/master/data/chipotle.tsv", sep="\t")
display(chipo.head())
display(chipo.info())
display(chipo.describe())
# ### Step 4. See the first 10 entries
chipo.head(10)
# ### Step 5. Create a histogram of the top 5 items bought
chipo.item_name.value_counts()[:5].plot.bar()
# +
x = chipo.item_name
letter_counts = collections.Counter(x)
df = pd.DataFrame.from_dict(letter_counts, orient="index")
df = df[0].sort_values(ascending=True)[45:50]
df.plot(kind="bar")
plt.xlabel("Items")
plt.ylabel("Price")
plt.title("Most ordered Chipotle's Items'")
# -
# ### Step 6. Create a scatterplot with the number of items orderered per order price
# #### Hint: Price should be in the X-axis and Items ordered in the Y-axis
chipo["item_price"] = chipo.item_price.apply(lambda x : float(x[1:]))
x = chipo[["order_id", "item_price"]].groupby("order_id").agg(["count", "sum"])
display(x)
display(x.columns)
x.columns = x.columns.levels[1]
x.plot.scatter(x="sum", y="count")
orders = chipo.groupby("order_id").sum()
plt.scatter(x = orders.item_price, y=orders.quantity, s=50, c="green")
orders.plot.scatter(x="item_price", y="quantity", s=50, c="green", marker='o')
# ### Step 7. BONUS: Create a question and a graph to answer your own question.
| other/pandas_exercises/07_Visualization/Chipotle/Exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: genomics
# language: python
# name: genomics
# ---
import os
import sys
import argparse
import math
import time
import h5py
import joblib
import subprocess
import numpy as np
import pandas as pd
import scipy
import statsmodels.api as sm
import matplotlib.pyplot as plt
import seaborn as sns
os.chdir('/Users/pengl7/Downloads/WGS/compare-variants/NIST/')
# + tags=[]
# %ls -lth
# -
# df = pd.read_csv("long4_with_title", sep="\t",index_col="POS", na_values={}, dtype={'UNMATCH': "category", "DP": "float", "GQ": "float", "MQ": "float"}, )
df = pd.read_csv("long_UpdateAF4_with_title", sep="\t")
# + tags=[]
print(df.shape)
print(df.columns.to_list())
# -
df.dtypes
df.describe()
# ## Get rid of some non-numeric values and change data types
for col in ['DP', 'GQ', 'MQ', 'QD']:
print(len(df[df[col]== "."]))
# + tags=[]
myFilter = (df["DP"]!= ".") & (df["GQ"]!= ".") & (df["MQ"]!= ".") & (df["QD"]!= ".")
df = df[myFilter]
print(df.shape)
# -
# change data type
cols = ['DP', 'GQ', 'MQ', "QD"]
for col in cols:
df[col]= df[col].astype("float")
df.to_csv("long_cleared_UpdateAF.csv")
# ## Apply logsistic regression model
df.describe()
df["TYPE"] = df["TYPE"].astype("category")
this_formula = "UNMATCH ~ TYPE + DP + GQ + QUAL + FS + AF + MQ + QD"
res = sm.formula.glm(formula=this_formula, family=sm.families.Binomial(), data=df).fit()
res.summary()
# + tags=[]
print("Coefficeients")
print(res.params)
print()
print("p-Values")
print(res.pvalues)
print()
print("Dependent variables")
print(res.model.endog_names)
# -
df["QualByDepth"] = df["GQ"]/(df["DP"]+1)
| .ipynb_checkpoints/logistic_reg_data_clean-UpdateAF-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from selenium import webdriver
import pandas as pd
import time
# +
options = webdriver.ChromeOptions()
options.add_argument("--start-maximized")
path = "chromedriver.exe"
driver = webdriver.Chrome(executable_path=path, options=options)
time.sleep(3)
# -
url = 'http://www4.planalto.gov.br/legislacao/portal-legis/legislacao-covid-19'
driver.get(url)
# +
table = driver.find_element_by_xpath("//*[@id='visao2']/table/tbody")
elements = table.find_elements_by_xpath("//*[@id='visao2']/table/tbody/tr/*")
# +
count = 0
atos = []
ementa = []
for i in elements:
if (count % 2) == 0 :
atos.append(i.text)
else:
ementa.append(i.text)
count += 1
# +
df = pd.DataFrame()
df['Ementa'] = ementa
df['Atos'] = atos
df.drop(df.index[[0]], inplace = True)
# -
df
df.to_csv(r'ementas', index = False)
count=0
for i in elements:
if((elements.index(i)) % 2) == 0:
count+=1
if(count==1):
continue
elif(count>=2):
link = i.find_element_by_xpath("//*[@id='visao2']/table/tbody/tr[{}]/td[1]/a".format(count))
href = link.get_attribute("href")
print(href)
print(i.text)
print("---")
# +
#len(df)
# +
#link = driver.find_element_by_xpath("//*[@id='visao2']/table/tbody/tr[7]/td[1]/a")
#link_text = link.get_attribute("href")
#
#print(link_text)
#print(link.text)
# +
#for i in elements:
# print(i.get_attribute('href'))
# print('--')
# +
#links = []
#for i in range(2,len(df)+2):
#
# link = driver.find_element_by_xpath("//*[@id='visao2']/table/tbody/tr[{}]/td[1]/a".format(i))
# link_text = link.get_attribute("href")
# links.append(link_text)
#
# +
#count = 1
#for i in range(1,len(df)):
# print(count, '', df.URL[i])
# count+=1
# +
#link = driver.find_element_by_xpath("//*[@id='visao2']/table/tbody/tr[29]/td[1]/a")
#link_text = link.get_attribute("href")
# +
#link_text
# +
#elements = table.find_elements_by_xpath("//*[@id='visao2']/table/tbody/tr/*")
# +
#elements[2].text
| planoB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
# # Old function
#
# The following definition is the old function used during the Hands-On, not currently integrated in the PyCalib library. It is here in order to compare the old and new versions and see if the new version is better. It is intended to be removed from here once the new version is final.
def plot_reliability_diagram_gaps_from_bins(bin_probs, bin_labels, title=None,
fig=None, ax=None, legend=True):
'''Plot binary reliability diagram gaps
Parameters
==========
bin_probs : list
List of lists representing the probabilities in each bin
bin_labels : list
List of lists representing the actual labels in each bin
n_bins : integer
Number of bins to divide the scores
title : string
Title for the plot
fig : matplotlib.pyplot.figure
Plots the axis in the given figure
ax : matplotlib.pyplot.Axis
Axis where to draw the plot
legend : boolean
If True the function will draw a legend
Regurns
=======
fig : matplotlib.pyplot.figure
Figure with the reliability diagram
'''
if fig is None and ax is None:
fig = plt.figure()
if ax is None:
ax = fig.add_subplot(111)
if title is not None:
ax.set_title(title)
n_bins = len(bin_probs)
bin_size = np.zeros(n_bins)
centers = np.zeros(n_bins)
lefts = np.zeros(n_bins)
start = 0
for i in range(n_bins):
centers[i] = np.mean(bin_probs[i])
lefts[i] = start
if i == (n_bins - 1):
bin_size[i] = 1 - start
else:
end = (centers[i] + np.mean(bin_probs[i + 1])) / 2
bin_size[i] = end - start
start = end
true_proportion = np.array([np.mean(l) for l in bin_labels])
pred_mean = np.array([np.mean(probs) for probs in bin_probs])
not_nan = np.isfinite(true_proportion - centers)
ax.bar(lefts, true_proportion, width=bin_size, edgecolor="black",
color="blue", label='True class prop.', align='edge')
ax.bar(pred_mean[not_nan], (true_proportion - pred_mean)[not_nan],
bottom=pred_mean[not_nan], width=bin_size/4.0, edgecolor="red",
color="#ffc8c6",
label='Gap pred. mean')
ax.scatter(pred_mean[not_nan], true_proportion[not_nan], color='red',
marker="+", zorder=10)
if legend:
ax.legend()
ax.plot([0, 1], [0, 1], linestyle="--")
ax.set_xlim([0, 1])
ax.set_xlabel('Predicted probability')
ax.set_ylim([0, 1])
ax.set_ylabel('Proportion of positives')
ax.grid(True)
ax.set_axisbelow(True)
fig.tight_layout()
print(bin_size)
return fig, ax
# # Reliability diagrams with user-provided bins
#
# The following is the old way of defining the probabilities and labels. These were defined by a list of lists, each inner list contained the scores given by the model in **bin_probs** and the true labels in **bin_labels**.
# +
bin_probs = [
[0.1, 0.1],
[0.4, 0.4],
[0.7, 0.7, 0.7],
[0.9]
]
bin_labels = [
[0, 0],
[0, 1],
[0, 1, 1],
[1]
]
fig = plt.figure(figsize=(5, 4))
plot_reliability_diagram_gaps_from_bins(bin_probs, bin_labels, fig=fig)
plt.savefig('Forecaster1-fixed-gaps-v1.pdf')
# -
# # New version from PyCalib
#
# The new version does not need a list of lists to specify the bins. Instead, an arbitrary list of scores and labels and a list of bin boundaries can be passed. We will use the previously defined lists of lists and concatenate them with numpy instead of defining again here.
#
# **However**, the scores need to be of shape (n_samples, n_classes), for that reason we stack the original scores with their complementary in a (2, n_samples) and then transpose.
# +
from pycalib.visualisations import plot_reliability_diagram
labels = np.concatenate(bin_labels)
scores = np.concatenate(bin_probs)
bins = [0, 0.25, 0.5, 0.8, 1.0]
fig = plt.figure(figsize=(5, 4))
fig = plot_reliability_diagram(labels, np.vstack([1 - scores, scores]).T,
class_names=['not 1', 'rain'], bins=bins,
fig=fig, show_gaps=True,
show_bars=True)
fig.savefig('Forecaster1-fixed-gaps-v2.pdf')
# -
# We see several other options in the following cells.
fig = plot_reliability_diagram(labels, np.vstack([1 - scores, scores]).T,
class_names=['not 1', 'rain'], bins=bins,
show_correction=True,
hist_per_class=True)
fig = plot_reliability_diagram(labels, np.vstack([1 - scores, scores]).T,
class_names=['not 1', 'rain'], bins=bins,
show_correction=True,
hist_per_class=False,
show_counts=True)
# +
bin_probs = [
[0.1, 0.2],
[0.3, 0.4],
[0.6, 0.7, 0.8],
[0.9]
]
bin_labels = [
[0, 0],
[0, 1],
[0, 1, 1],
[1]
]
fig = plt.figure(figsize=(5, 4))
plot_reliability_diagram_gaps_from_bins(bin_probs, bin_labels, fig=fig)
plt.savefig('Forecaster2-fixed-gaps-v1.pdf')
# -
labels = np.concatenate(bin_labels)
scores = np.concatenate(bin_probs)
bins = [0, 0.25, 0.5, 0.85, 1.0]
fig = plt.figure(figsize=(5, 4))
fig = plot_reliability_diagram(labels, np.vstack([1 - scores, scores]).T,
class_names=['not 1', 'rain'], bins=bins,
fig=fig,
hist_per_class=False,
show_bars=True,
show_gaps=True)
fig.savefig('Forecaster2-fixed-gaps-v2.pdf')
# +
bin_probs = [
[.1, .2, .3, .4],
[.6, .7, .8, .9],
]
bin_labels = [
[0, 0, 0, 1],
[0, 1, 1, 1],
]
fig = plt.figure(figsize=(5, 4))
plot_reliability_diagram_gaps_from_bins(bin_probs, bin_labels, fig=fig)
plt.savefig('Forecaster3-fixed-gaps-v1.pdf')
# -
labels = np.concatenate(bin_labels)
scores = np.concatenate(bin_probs)
bins = [0, 0.5, 1]
fig = plt.figure(figsize=(5, 4))
fig = plot_reliability_diagram(labels, np.vstack([1 - scores, scores]).T,
class_names=['not 1', 'rain'], bins=bins,
fig=fig,
hist_per_class=False,
show_bars=True,
show_gaps=True)
fig.align_labels()
fig.savefig('Forecaster3-fixed-gaps-v2.pdf')
# +
bin_probs = [
[.1],
[.2],
[.3],
[.4],
[.6],
[.7],
[.8],
[.9],
]
bin_labels = [
[0],
[0],
[0],
[1],
[0],
[1],
[1],
[1],
]
fig = plt.figure(figsize=(5, 4))
plot_reliability_diagram_gaps_from_bins(bin_probs, bin_labels, fig=fig)
plt.savefig('Forecaster4-fixed-gaps-v1.pdf')
# -
labels = np.concatenate(bin_labels)
scores = np.concatenate(bin_probs)
bins = [0, .15, .25, .35, .45, .65, .75, .85, 1]
fig = plt.figure(figsize=(5, 4))
fig = plot_reliability_diagram(labels, np.vstack([1 - scores, scores]).T,
class_names=['not 1', 'rain'], bins=bins,
fig=fig,
hist_per_class=False,
show_bars=True,
show_gaps=True)
fig.savefig('Forecaster4-fixed-gaps-v2.pdf')
| notebooks/01_introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Load the 7 day forecast from DarkSky.Net into Pandas!
#
# In this example you will go back to https://api.darksky.net and use your API key to get the current weather forecast for:
#
# Syrcause, NY `(lat=43.0481221, lng=-76.1474244)`
#
# In the forecast output, find the daily 7 day forecast (it's the current conditions + 7 days out == 8 days total)
#
# Extract the forecast data and load it into Pandas, then display the Time of the forecast, and high and low temperatures:
#
# **HINT**: To get the times to show up in a human-readable format, you must convert the DarkSky API time (which is in unix timestamp format) to a Pandas Timestamp date/time format. The `pd.to_datetime()` function can help you:
#
# ```
# time = 1489402800 # this the time format Darksky returns
# readable_time = pd.to_datetime(time, unit='s') ## s stands for unix timestamp format
# readable_time
#
# Timestamp('2017-03-13 11:00:00')
# ```
#
# Just replace the `['time']` column in the `DataFrame` with the new version.
# ## Step 1: Problem Analysis for entire program
#
# Inputs:
#
# Outputs:
#
# Algorithm (Steps in Program):
#
# +
# STEP 2: Todo write code here
import requests
import pandas as pd
import numpy as np
from IPython.display import display
lat = 43.048122
lng = -76.147424
key = 'f401535c6d98772f61738417ac33e0c5'
url = 'https://api.darksky.net/forecast/%s/%s,%s' % (key,lat,lng)
print(url)
# -
response = requests.get(url)
if response.ok:
weather = response.json()
#print(weather.keys())
print(weather['daily']['data'][1]['time'])
time = weather['daily']['data'][1]['time']
realtime = (pd.to_datetime(time, unit='s'))
print(type(realtime))
# +
# Here's my output from when I ran the solution, to give you an example of what I expect
def getforecastdata():
lat = 43.048122
lng = -76.147424
key = 'f401535c6d98772f61738417ac33e0c5'
url = 'https://api.darksky.net/forecast/%s/%s,%s' % (key,lat,lng)
try:
response = requests.get(url)
weather = response.json()
realtime = []
mintemp =[]
maxtemp = []
for day in weather['daily']['data']:
realtime.append(pd.to_datetime(day['time'], unit='s'))
mintemp.append(day['temperatureMin'])
maxtemp.append(day['temperatureMax'])
return(realtime,mintemp,maxtemp)
except json.decoder.JSONDecodeError as e:
print('ERROR: Cannot decod the response into json')
print('DETAILS', e)
except requests.exceptions.HTTPError as e:
print('ERROR: Response from', url, 'was not ok.')
print('DETAILS:', e)
except reuests.exceptions.RequestException as e:
print('ERROR: Cannot connect to', url)
print('DETAILS:', e)
# -
# ## Step 3: Questions
#
# 1. Pandas programs are different than typical Python programs. Explain the process by which you got the final solution?
# 2. What was the most difficult aspect of this assignment?
# ## Reminder of Evaluation Criteria
#
# 1. What the problem attempted (analysis, code, and answered questions) ?
# 2. What the problem analysis thought out? (does the program match the plan?)
# 3. Does the code execute without syntax error?
# 4. Does the code solve the intended problem?
# 5. Is the code well written? (easy to understand, modular, and self-documenting, handles errors)
| content/lessons/12/Now-You-Code/NYC3-Pandas-Darksky-Mashup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Sev-RyanLeo/Linear-Algebra-58019/blob/main/Application.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="6ymUn3z7OO06"
# November 17
# + colab={"base_uri": "https://localhost:8080/"} id="WDxv0XZ2MLG9" outputId="e3daa7d7-93ab-407e-fda4-08443ca24fca"
import numpy as np
A=np.array([[4,3,2],[-2,2,3],[3,-5,2]])
B=np.array([[25],[-10],[-4]])
print(A,"\n \n",B)
x=np.linalg.solve(A,B)
print("\n Answer: \n",x)
# + [markdown] id="Ou68gj8MNl_U"
# **using scipy.linalg**
# + colab={"base_uri": "https://localhost:8080/"} id="FCnRSUisNO-7" outputId="93eb9c48-18e0-4407-d754-72987016f619"
import numpy as np
from scipy.linalg import solve
A=np.array([[4,3,2],[-2,2,3],[3,-5,2]])
B=np.array([[25],[-10],[-4]])
print(A,"\n \n",B)
x=solve(A,B)
print("\n Answer: \n",x)
# + id="6N_AMU7CN22Z"
| Application.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Raku
# language: raku
# name: raku
# ---
# # Yachts and Navigation - on Raku
#
# # Chapter 2 - Latitude and Longitude
#
# _Copyright @2022 p6steve, please see Chapter 1 for Terms of Use_
#
# This chapter will expand on the Latitude and Longitude classes that we met in Chapter 1 and will present both raku and navigation concepts cell by cell.
#
# First, Jupyter notebooks need us to repeat a little preamble each time...
use Physics::Navigation;
use Physics::Measure;
# A perspective view of the Earth showing how latitude ϕ(phi) and longitude (λ)lambda are defined on a spherical model.
# 
# Latitude is the angle on the Earth between the equator (0º) and the North or South pole (90ºN or 90ºS).
# Latitude and Longitude can be expressed in a similar way with either longhand and shorthand syntax - the pisces `♓️` symbol can be used with angle brackets and degree/minute/seconds and the compass point in this format `♓️<43°30′30″S>;`. Use compass points N&S for Latitude and E&W for Longitude.
my \ϕ1 = Latitude.new( value => 45, compass => <N> ); say "ϕ1 is {ϕ1}";
my \ϕ2 = ♓️<43°30′30″S>; say "ϕ2 is {ϕ2}";
# Hint: Instead of declaring a variable with a $ sigil, you can use a backslash.
# You can do basic math on Latitudes - and the compass direction is taken into account. However, you will get an error if the result is greater than 90°.
# +
say ϕ1 + ϕ2; # S adopts a -ve value
say ϕ1 - ϕ2; # minus a -ve is a plus
try {
say 3 * ϕ1; # value -90 <= ϕ <= 90
}
if $! {"may not exceed 90°";}
# -
# Longitude is the angle on the Earth east or west of the Greenwich Meridian up to plus 180ºE or minus 180ºW.
my \λ1 = Longitude.new( value => 45, compass => <E> ); say "λ1 is {λ1}";
my \λ2 = ♓️<22°W>; say "λ2 is {λ2}";
# Hint: Degree outputs are zero padded to two (Latitude) of three (Longitude) digits to reflect their respective limits.
# +
say λ1 + λ2; # W adopts a -ve value
say λ1 - λ2; # minus a -ve is a plus
try {
say 5 * λ1; # value -180 < λ <= 180
}
if $! {"may not exceed 180°";}
# -
# The length of each degree of Latitude on the surface is constant:-
#
# - Each degree of Latitude = 60 nautical miles
# - Each minute of Latitude = 1 nautical mile
#
# The in-lat() function converts from Distance in nmiles to ° Latitude. Conversely the method .in('nmiles') will convert back.
# +
my $d = ♓️'42 nmile';
my \λ3 = in-lat( $d ); say "λ3 is {λ3}";
say "{λ3.in('nmiles')}";
# -
# No similar equivalence exists for Longitude since the distance travelled per degree at the surface is not constant.
| eg/Yachts and Navigation - Chapter 2 - Latitude and Longitude.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="wZub42g3Br6h" colab_type="text"
# ในการทำงานบางครั้งเราต้องการข้อมูลรูปภาพ นอกเหนือจากที่อยู่ใน Dataset เราต้องการป้อนข้อมูล Input รูปภาพจากโลกของความเป็นจริง แบบ Real-time หนึ่งในวิธีที่สะดวก และเป็นที่นิยม คือ การใช้กล้อง Webcam ที่อยู่ในโน้ตบุ๊คเกือบทุกเครื่อง ถ่ายภาพตัวเราเอง หรือถ้ามีกล้องอื่น ๆ ต่อผ่าน USB ก็สามารถใช้ได้เช่นกัน
# + [markdown] id="Diloa97rIg-X" colab_type="text"
# # 0. Import
# + [markdown] id="TZgSG7S1IjUo" colab_type="text"
# Import Library ที่จำเป็นต้องใช้ ในการเรียก Javascript ใน Web Browser และแสดงผลรูปภาพ
# + id="J-TVagd7FIf1" colab_type="code" colab={}
from IPython.display import display, Javascript, Image
from google.colab.output import eval_js
from base64 import b64decode
import numpy as np
import io
import PIL
import matplotlib.pyplot as plt
# + [markdown] colab_type="text" id="2viqYx97hPMi"
# # 1. ถ่ายภาพด้วยกล้องหน้า
# + [markdown] id="cuRV-3O0CeyB" colab_type="text"
# เราจะใช้ Google Colab สั่ง Webcam ที่มีอยู่ในทุกคอมพิวเตอร์โน้ตบุ๊ค ถ่ายภาพ เพื่อมาใช้ในการประมวลผลต่อไป
#
# ในทางเทคนิคแล้ว เราจะไปเรียกใช้ Javascript ไปสั่งให้ Web Browser ขอสิทธิ์ และถ่ายภาพให้เราอีกที
# + colab_type="code" id="SucxddsPhOmj" colab={}
def take_photo(filename=None, quality=0.8):
js = Javascript('''
async function takePhoto(quality) {
const div = document.createElement('div');
const video = document.createElement('video');
video.style.display = 'block';
const stream = await navigator.mediaDevices.getUserMedia({video: true});
document.body.appendChild(div);
div.appendChild(video);
video.srcObject = stream;
await video.play();
// Resize the output to fit the video element.
google.colab.output.setIframeHeight(document.documentElement.scrollHeight, true);
// Wait for video to be clicked.
await new Promise((resolve) => video.onclick = resolve);
const canvas = document.createElement('canvas');
canvas.width = video.videoWidth;
canvas.height = video.videoHeight;
canvas.getContext('2d').drawImage(video, 0, 0);
stream.getVideoTracks()[0].stop();
div.remove();
return canvas.toDataURL('image/jpeg', quality);
}
''')
display(js)
data = eval_js('takePhoto({})'.format(quality))
binary = b64decode(data.split(',')[1])
if filename is None:
## return numpy array
f = io.BytesIO(binary)
return np.asarray(PIL.Image.open(f))
else:
# save file and return length of file
with open(filename, 'wb') as f:
f.write(binary)
return len(binary)
# + [markdown] id="PDpdSsnCGAdD" colab_type="text"
# # 2. ถ่ายภาพ เป็น Numpy Array
# + [markdown] id="xK2jA98hG5h3" colab_type="text"
# คลิกที่รูปเพื่อกดชัตเตอร์ ถ้าเราไม่ใส่ชื่อไฟล์ ฟังก์ชันจะ return เป็น numpy array 3 มิติ
# + id="N4M3PeXxEV7H" colab_type="code" outputId="69ea0c8d-3f3e-4834-d34f-43449f577757" colab={"base_uri": "https://localhost:8080/", "height": 34}
img = take_photo() # click
img.shape
# + [markdown] id="sCsv2hCnJI45" colab_type="text"
# ใช้ matplotlib แสดงรูปที่ถ่ายไว้ด้านบน
# + colab_type="code" id="buJCl90WhNfq" outputId="0743504d-5393-437a-a780-3520d8730021" colab={"base_uri": "https://localhost:8080/", "height": 470}
plt.figure(figsize=(10,10))
plt.imshow(img)
plt.show()
# + [markdown] colab_type="text" id="MqNaKqKBGXGE"
# # 3. ถ่ายภาพแล้ว เซฟเป็นไฟล์
# + [markdown] id="JW1-GzLwG-1b" colab_type="text"
# คลิกที่รูปเพื่อกดชัตเตอร์ ถ้าเราใส่ชื่อไฟล์ ฟังก์ชันจะ return เซฟไฟล์ในชื่อนั้น และ return ความยาวของไฟล์
# + colab_type="code" outputId="46d92b6d-3c72-4e43-c717-8c33d611bd3b" id="wTqaMeGxGXGL" colab={"base_uri": "https://localhost:8080/", "height": 34}
filename = "photo01.jpg"
img = take_photo(filename=filename)
img
# + [markdown] id="EN8bnafbJsz0" colab_type="text"
# ลอง ls ดูว่ามีไฟล์ชื่อที่กำหนดหรือไม่
# + colab_type="code" outputId="5e2f42d0-e8ac-4375-a2d0-91e11a4b9020" id="FyzAsR-9GXGS" colab={"base_uri": "https://localhost:8080/", "height": 34}
# !ls
# + [markdown] id="0zlkHjRWJxM9" colab_type="text"
# เปิดไฟล์ขึ้นมา แสดงรูปภาพที่ถ่ายไว้ด้านบน
# + id="IAwSaxtVGVmX" colab_type="code" outputId="0afa5f61-7d14-4079-d831-d84658d63b1a" colab={"base_uri": "https://localhost:8080/", "height": 497}
display(Image(filename))
# + [markdown] id="ahMTOww-Bt8Y" colab_type="text"
# # Credit
# + [markdown] id="jy_1sfE1BuBq" colab_type="text"
# * https://colab.research.google.com/notebooks/snippets/advanced_outputs.ipynb#scrollTo=2viqYx97hPMi
# * https://colab.research.google.com/gist/ricardodeazambuja/058f4c242fe67ec2d86ca2596b0905ad/webcam-to-numpy-array-from-your-browser-in-colab.ipynb#scrollTo=58dO8mqemQyZ
# + id="5V4bXdpmDeOQ" colab_type="code" colab={}
| nbs/11b_webcam_colab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Иерархическая кластеризация
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('ggplot')
plt.rcParams['figure.figsize'] = (12,5)
# -
df = pd.read_csv('food.txt', sep=' ')
df.head()
X = df.iloc[:, 1:].values
X = (X - X.mean(axis=0))/X.std(axis=0)
X.mean(axis=0)
X.std(axis=0)
from scipy.cluster.hierarchy import linkage, fcluster, dendrogram
Z = linkage(X, method='average', metric='euclidean')
dend = dendrogram(Z, orientation='left', color_threshold=0.0, labels=df.Name.values)
label = fcluster(Z, 2.2, criterion='distance')
np.unique(label)
df.loc[:, 'label'] = label
for i, group in df.groupby('label'):
print('=' * 10)
print('cluster {}'.format(i))
print(group)
| W4/Hiearchy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ourekouch/Coursera_Capstone/blob/master/The_Battle_of_Neighborhoods.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="xGBrbmdoM0bv" colab_type="code" colab={}
import pandas as pd
# + id="loMdKfVeVyq_" colab_type="code" outputId="61f722f7-f2b9-4ae4-c434-194c985eb8b9" colab={"base_uri": "https://localhost:8080/", "height": 782}
# !pip install beautifulsoup4
# !pip install lxml
# !pip install html5lib
# !pip install requests
# !pip install geocoder
# !pip install geopy
# !pip install Nominatim
# !pip install folium
# !pip install requests
# !pip install sklearn
# + [markdown] id="OGzVP24JX-mg" colab_type="text"
# **Introduction/Business Problem**
#
# A constractor want to start a new business in fast food in london . Unfortunately he has no idea about the right area for this project.
# So he decided to rely on the science of data analysis in order to find the appropriate area for this new project,
# especially the population density in various neighborhoods of London, as well as the distribution of different venues and facilities
# in the city of london .
# + [markdown] id="OaJhpw4OW9lm" colab_type="text"
#
# **1. Data acquisition : Data about London Boroughs and populations in london**
#
#
# In this project we will use data from "https://en.wikipedia.org/wiki/List_of_London_boroughs" , but we need first to get data by scraping and clean it .
# + id="ea2gkqxWVNqR" colab_type="code" outputId="219d4af7-c243-44f5-df50-aacf4ab947b3" colab={"base_uri": "https://localhost:8080/", "height": 204}
#importing libraries
from bs4 import BeautifulSoup
import requests
import pandas as pd
import numpy as np
#Source of Data
source = requests.get("https://en.wikipedia.org/wiki/List_of_London_boroughs").text
soup=BeautifulSoup(source,'lxml')
#add an empty dataframe
df_london = pd.DataFrame(columns=['Borough','Population','Coord'])
#finding our data and scraping it using beautifulSoup library
table=soup.find('table')
body=table.find('tbody')
for row in body.find_all('tr'):
List=[]
for x in row.find_all('td'):
List.append(x.text)
list_df = pd.DataFrame(List)
#for each row we check if the row has values and borough different to not assigned then we add row to dataframe
if len(List)>0 :
df2 = pd.DataFrame({'Borough':[List[0]], 'Population':[List[7]],'Coord':[List[8]]})
if List[1] != "Not assigned" :
df_london = df_london.append(df2,ignore_index=True)
#Data wrangling and data cleaning :
df_london['Borough']=df_london['Borough'].astype(str).str.replace('\n','')
df_london['Population']=df_london['Population'].astype(str).str.replace('\n','')
df_london[['Latitude','Longitude']] = df_london.Coord.str.split("″N ",expand=True)
df_london['Longitude'] = df_london.Longitude.str.split("/" ,expand=True)
df_london['Borough'] = df_london.Borough.str.split("[" ,expand=True)
df_london['Latitude']=df_london['Latitude'].astype(str).str.replace('°',',')
df_london['Latitude']=df_london['Latitude'].astype(str).str.replace('′','')
df_london['Longitude']=df_london['Longitude'].astype(str).str.replace('°',',')
df_london['Longitude']=df_london['Longitude'].astype(str).str.replace('′','')
df_london['Longitude']=df_london['Longitude'].astype(str).str.replace('″E','')
df_london.loc[df_london.Longitude.astype(str).str.contains('″W'), 'Longitude']='-'+df_london['Longitude'].astype(str)
df_london['Longitude']=df_london['Longitude'].astype(str).str.replace('″W','')
df_london['Longitude']=df_london['Longitude'].astype(str).str.replace('\ufeff','')
df_london['Latitude']=df_london['Latitude'].astype(str).str.replace('\ufeff','')
df_london['Population']=df_london['Population'].astype(str).str.replace(',','')
#replace , by . in order to make transformation object -> String : possible
df_london['Latitude']=df_london['Latitude'].astype(str).str.replace(',','.')
df_london['Longitude']=df_london['Longitude'].astype(str).str.replace(',','.')
#we don't need this column anymore
del df_london['Coord']
#Transformation to numeric forme
df_london['Latitude'] = pd.to_numeric(df_london['Latitude'])
df_london['Longitude'] = pd.to_numeric(df_london['Longitude'])
df_london['Population'] = pd.to_numeric(df_london['Population'])
#some changes in coordinate due to tranformation of Geographic Coordinates to Decimal
my_list1=[]
my_list2=[]
for i, row in df_london.iterrows():
x=row['Latitude']
x2=row['Longitude']
y=100*(x-int(x))
z=100*(y-int(y))
y2=100*(x2-int(x2))
z2=100*(y2-int(y2))
my_list1.append(int(x) + y/60 + z/3600)
my_list2.append(int(x2) + y2/60 + z2/3600)
df_london['Latitude'] = my_list1
df_london['Longitude'] = my_list2
#Finally our DATA
df_london.head()
# + [markdown] id="lRnp13R9aR8P" colab_type="text"
# **2. Data analysis : population and clusturing**
#
# In this section we will use our data and forsquare api in order to get all informations about london boroughs that can help us to find a solution to the business problem
#
#
# + id="ww622mYw6rCY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 407} outputId="9dfa6515-b00b-4082-ca4e-350b7cf16740"
df_sorted=df_london[['Borough','Population']].sort_values(by='Population',ascending=True)
df_sorted.set_index('Borough', inplace=True)
df_sorted.tail(5).plot(kind='barh',stacked=True ,figsize=(10, 6))
plt.title('Top 5 Boroughs by Population on 2013')
# + id="kdlP90sVnsbF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 772} outputId="907546a6-bde9-4356-b633-714b347d331d"
import folium # map rendering library
import matplotlib.cm as cm
import matplotlib.colors as colors
from geopy.geocoders import Nominatim # convert an address into latitude and longitude values
import matplotlib.pyplot as plt
address = 'London'
geolocator = Nominatim(user_agent="tr_explorer")
location = geolocator.geocode(address)
latitude = location.latitude
longitude = location.longitude
print('The geograpical coordinate of London City are {}, {}.'.format(latitude, longitude))
# create map of London using latitude and longitude values
map_London = folium.Map(location=[latitude, longitude], zoom_start=10)
# add markers to map''
for lat, lng, borough in zip(df_london['Latitude'], df_london['Longitude'], df_london['Borough']):
label = '{},{}'.format(borough,borough)
label = folium.Popup(label, parse_html=True)
folium.CircleMarker(
[lat, lng],
radius=5,
popup=label,
color='blue',
fill=True,
fill_color='#3186cc',
fill_opacity=0.7,
parse_html=False).add_to(map_London)
map_London
# + id="BUE-7ed7OflF" colab_type="code" colab={}
def getNearbyVenues(names, latitudes, longitudes, radius=500):
venues_list=[]
CLIENT_ID='G5TUS4T0FKE1X5DVH22U4I1SUFQD5FPUYVQVFBS5JVEBTGGU'
CLIENT_SECRET='<KEY>'
VERSION = '20180604'
LIMIT = 30
for name, lat, lng in zip(names, latitudes, longitudes):
print(name)
# create the API request URL
url = 'https://api.foursquare.com/v2/venues/explore?&client_id={}&client_secret={}&v={}&ll={},{}&radius={}&limit={}'.format(
CLIENT_ID,
CLIENT_SECRET,
VERSION,
lat,
lng,
radius,
LIMIT)
# make the GET request
results = requests.get(url).json()["response"]['groups'][0]['items']
# return only relevant information for each nearby venue
venues_list.append([(
name,
lat,
lng,
v['venue']['name'],
v['venue']['location']['lat'],
v['venue']['location']['lng'],
v['venue']['categories'][0]['name']) for v in results])
nearby_venues = pd.DataFrame([item for venue_list in venues_list for item in venue_list])
nearby_venues.columns = ['Neighborhood',
'Neighborhood Latitude',
'Neighborhood Longitude',
'Venue',
'Venue Latitude',
'Venue Longitude',
'Venue Category']
return(nearby_venues)
# + id="XkZtP28rPV4I" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 561} outputId="d442ecdb-3437-473a-dec7-75abafae6b82"
London_venues = getNearbyVenues(names=df_london['Borough'],
latitudes=df_london['Latitude'],
longitudes=df_london['Longitude']
)
# + id="Gj8vg0kcP9xf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 820} outputId="ff74dbc2-04b9-4fe4-ea2f-bfca53324532"
# one hot encoding
London_onehot = pd.get_dummies(London_venues[['Venue Category']], prefix="", prefix_sep="")
# add neighborhood column back to dataframe
London_onehot['Neighborhood'] = London_venues['Neighborhood']
# move neighborhood column to the first column
fixed_columns = [London_onehot.columns[-1]] + list(London_onehot.columns[:-1])
London_onehot = London_onehot[fixed_columns]
London_onehot.head(20)
# + id="XL0Smro6RdcB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="888baa5d-89bd-436c-e746-22876194a179"
London_grouped = London_onehot.groupby('Neighborhood').mean().reset_index()
London_grouped
# + id="0T-kSyhbRmo5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="12dd7f0c-3401-49d1-de2c-f7cf7f8387e3"
num_top_venues = 5
for hood in London_grouped['Neighborhood']:
print("----"+hood+"----")
temp = London_grouped[London_grouped['Neighborhood'] == hood].T.reset_index()
temp.columns = ['venue','freq']
temp = temp.iloc[1:]
temp['freq'] = temp['freq'].astype(float)
temp = temp.round({'freq': 2})
print(temp.sort_values('freq', ascending=False).reset_index(drop=True).head(num_top_venues))
print('\n')
# + id="qNQAgRdGSEm_" colab_type="code" colab={}
def return_most_common_venues(row, num_top_venues):
row_categories = row.iloc[1:]
row_categories_sorted = row_categories.sort_values(ascending=False)
return row_categories_sorted.index.values[0:num_top_venues]
# + id="cJxweM35SLOq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="bd6f4935-3ae0-418f-a8bc-293f510a7724"
import numpy as np
num_top_venues = 5
indicators = ['st', 'nd', 'rd']
# create columns according to number of top venues
columns = ['Neighborhood']
for ind in np.arange(num_top_venues):
try:
columns.append('{}{} Most Common Venue'.format(ind+1, indicators[ind]))
except:
columns.append('{}th Most Common Venue'.format(ind+1))
# create a new dataframe
neighborhoods_venues_sorted = pd.DataFrame(columns=columns)
neighborhoods_venues_sorted['Neighborhood'] = London_grouped['Neighborhood']
for ind in np.arange(London_grouped.shape[0]):
neighborhoods_venues_sorted.iloc[ind, 1:] = return_most_common_venues(London_grouped.iloc[ind, :], num_top_venues)
neighborhoods_venues_sorted.head()
# + id="scWtDtLbSw68" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fc4e9738-d5f4-430f-f3b3-3b8ea8e4ddcd"
# import k-means from clustering stage
from sklearn.cluster import KMeans
# set number of clusters
kclusters = 5
London_grouped_clustering = London_grouped.drop('Neighborhood', 1)
# run k-means clustering
kmeans = KMeans(n_clusters=kclusters, random_state=0).fit(London_grouped_clustering)
# check cluster labels generated for each row in the dataframe
kmeans.labels_[0:10]
# + id="XkqMz82MS5gE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 822} outputId="91748885-71ef-4018-eff5-e7466c01b0bf"
# add clustering labels
#neighborhoods_venues_sorted.insert(0, 'Cluster Labels', kmeans.labels_)
London_merged = df_london
# merge toronto_grouped with toronto_data to add latitude/longitude for each neighborhood
London_merged = London_merged.join(neighborhoods_venues_sorted.set_index('Neighborhood'), on='Borough')
London_merged.head(20) # check the last columns!
# + id="okrhItdsWejP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 755} outputId="5be1f201-791c-4494-e502-a66ea19df233"
import folium
# create map
map_clusters = folium.Map(location=[latitude, longitude], zoom_start=11)
#removing nan values
London_merged.dropna(inplace=True)
# set color scheme for the clusters
x = np.arange(kclusters)
ys = [i + x + (i*x)**2 for i in range(kclusters)]
colors_array = cm.rainbow(np.linspace(0, 1, len(ys)))
rainbow = [colors.rgb2hex(i) for i in colors_array]
# add markers to the map
markers_colors = []
for lat, lon, poi, cluster in zip(London_merged['Latitude'], London_merged['Longitude'], London_merged['Borough'], London_merged['Cluster Labels']):
label = folium.Popup(str(poi) + ' Cluster ' + str(cluster), parse_html=True)
folium.CircleMarker(
[lat, lon],
radius=5,
popup=label,
color=rainbow[int(cluster-1)],
fill=True,
fill_color=rainbow[int(cluster-1)],
fill_opacity=0.7).add_to(map_clusters)
map_clusters
# + [markdown] id="902AIMhufYis" colab_type="text"
# **Results**
#
# After the first vizualization we get that the first 5 boroughs according to population are :
#
#
# * Croydon (372 K )
# * Barnet (369 K )
# * Ealing (342 K )
# * Enfield (320 K )
# * Newham (318 K )
#
#
# after analysing and clustering boroughs we get that :
#
# ----Croydon----
# venue/freq:
# Coffee Shop 0.17
# Clothing Store 0.17
# Pub 0.10
# Bookstore 0.07
# Women's Store 0.03
#
# ----Barnet----
# venue/freq:
# Bus Stop 0.25
# Pub 0.25
# Event Service 0.25
# Fish & Chips Shop 0.25
# Outdoor Sculpture 0.00
#
# ----Ealing----
# venue/freq:
# Park 0.75
# Pharmacy 0.25
# American Restaurant 0.00
# Outdoor Sculpture 0.00
# Museum 0.00
#
# ----Enfield----
# venue/freq
# Pub 0.5
# Indian Restaurant 0.1
# Grocery Store 0.1
# Coffee Shop 0.1
# Sandwich Place 0.1
#
# ----Newham----
# venue/freq
# Light Rail Station 0.25
# Supermarket 0.25
# Gym / Fitness Center 0.12
# Bus Station 0.12
# Pub 0.12
#
#
# So in order to make the final decision between those 5 boroughs who has the max of population we will add another score (Competitive rate) :
#
# **competitive rate** : the sum of freq of venues in boroughs multiplcated by +1 (if the venue will be benefit for our business for exemple : +1 for shools , stadium ,...) , by -1(if the venue make a danger for our business for exemple another restaurant )
#
#
# resultsof Competitive rate :
# * Croydon (0.2 )
# * Barnet (1)
# * Ealing (1)
# * Enfield (0.3)
# * Newham (0.86 )
#
#
#
# In THE END THE BEST BOROUGH FOR STARTING A FAST FOOD RESTAURANT IN LONDON IS :
#
# **BARNET OR EALING**
| The_Battle_of_Neighborhoods.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Monocular Depth Estimation
# ### Imports
# # %matplotlib inline will plot the graphs along with the code
# +
import math
import h5py
import torch
import torchvision
import torchvision.transforms as transforms
import torch.nn as nn
from torchvision.models import vgg16
import numpy as np
from PIL import Image
import matplotlib.pyplot as plt
device = torch.device('cpu')
print(device)
# %matplotlib inline
import torch.nn.functional as F
# -
# ### Dataset class
class NYUDataset(torch.utils.data.Dataset):
def __init__(self, data_dir, tfms):
super(NYUDataset, self).__init__()
self.data_dir = data_dir
self.tfms = tfms
self.ds_v_1 = h5py.File(self.data_dir+'nyu_depth_data_labeled.mat')
self.ds_v_2 = h5py.File(self.data_dir+'nyu_depth_v2_labeled.mat')
self.len = len(self.ds_v_1["images"]) + len(self.ds_v_2["images"])
def __getitem__(self, index):
if(index<len(self.ds_v_1["images"])):
ds = self.ds_v_1
i = index
else:
ds = self.ds_v_2
i = index - len(self.ds_v_1["images"])
img = np.transpose(ds["images"][i], axes=[2,1,0])
img = img.astype(np.uint8)
depth = np.transpose(ds["depths"][i], axes=[1,0])
depth = (depth/depth.max())*255
depth = depth.astype(np.uint8)
if self.tfms:
tfmd_sample = self.tfms({"image":img, "depth":depth})
img, depth = tfmd_sample["image"], tfmd_sample["depth"]
return (img, depth)
def __len__(self):
return self.len
# ### Custom transforms
# +
class ResizeImgAndDepth(object):
def __init__(self, size_tup):
self.size = size_tup
def __call__(self, sample):
img = Image.fromarray(sample['image'], 'RGB').resize(self.size)
depth = Image.fromarray(sample['depth'], 'L').resize((self.size[0]//2, self.size[1]//2))
return { 'image' : np.array(img), 'depth' : np.array(depth) }
class RandomHorizontalFlip(object):
def __call__(self, sample):
img = sample["image"]
depth = sample["depth"]
if np.random.random() > 0.5:
img = np.fliplr(sample['image']).copy()
depth = np.fliplr(sample['depth']).copy()
return { 'image' : img, 'depth' : depth }
class ImgAndDepthToTensor(object):
def __init__(self):
self.ToTensor = transforms.ToTensor()
def __call__(self, sample):
return { 'image' : self.ToTensor(sample['image']), 'depth' : torch.tensor(sample['depth'], dtype=torch.float) }
class NormalizeImg(object):
def __init__(self, mean, std):
self.normalize = transforms.Normalize(mean, std)
def __call__(self, sample):
return { 'image' : self.normalize(sample['image']), 'depth' : sample['depth'] }
class UnNormalizeImgBatch(object):
def __init__(self, mean, std):
self.mean = mean.reshape((1,3,1,1))
self.std = std.reshape((1,3,1,1))
def __call__(self, batch):
return (batch*self.std) + self.mean
# -
# ### Model utils and Loss function
# +
def get_unnormalized_ds_item(unnormalize, item):
un_img = unnormalize(item[0][None])
return (un_img.squeeze(dim=0), item[1])
def freeze_all_layers(model):
for param in model.parameters():
param.requires_grad = False
def unfreeze_all_layers(model):
for param in model.parameters():
param.requires_grad = True
def get_model_predictions_on_a_sample_batch(model, dl):
model.eval()
with torch.no_grad():
batch, actual_labels = iter(dl).next()
batch = batch.to(device)
actual_labels = actual_labels.to(device)
predictions = model(batch)
return (predictions, batch, actual_labels)
def im_gradient_loss(d_batch, n_pixels):
a = torch.Tensor([[[[1, 0, -1],
[2, 0, -2],
[1, 0, -1]]]])
b = torch.Tensor([[[[1, 2, 1],
[0, 0, 0],
[-1, -2, -1]]]])
a = a.to(device)
b = b.to(device)
G_x = F.conv2d(d_batch, a, padding=1).to(device)
G_y = F.conv2d(d_batch, b, padding=1).to(device)
G = torch.pow(G_x,2)+ torch.pow(G_y,2)
return G.view(-1, n_pixels).mean(dim=1).sum()
def depth_loss(preds, actual_depth):
#preds.shape -> [16, 1, 120, 160]
#actual_depth.shape -> [16, 120, 160]
n_pixels = actual_depth.shape[1]*actual_depth.shape[2]
preds = (preds*0.225) + 0.45
preds = preds*255
preds[preds<=0] = 0.00001
actual_depth[actual_depth==0] = 0.00001
actual_depth.unsqueeze_(dim=1)
d = torch.log(preds) - torch.log(actual_depth)
grad_loss_term = im_gradient_loss(d, n_pixels)
term_1 = torch.pow(d.view(-1, n_pixels),2).mean(dim=1).sum() #pixel wise mean, then batch sum
term_2 = (torch.pow(d.view(-1, n_pixels).sum(dim=1),2)/(2*(n_pixels**2))).sum()
return term_1 - term_2 + grad_loss_term
def print_training_loss_summary(loss, total_steps, current_epoch, n_epochs, n_batches, print_every=10):
#prints loss at the start of the epoch, then every 10(print_every) steps taken by the optimizer
steps_this_epoch = (total_steps%n_batches)
if(steps_this_epoch==1 or steps_this_epoch%print_every==0):
print ('Epoch [{}/{}], Iteration [{}/{}], Loss: {:.4f}'
.format(current_epoch, n_epochs, steps_this_epoch, n_batches, loss))
def apply_sobel_operator_on_sample_ds_image(ds_item, unnormalize, T, P):
x = unnormalize(ds_item[0][None]) #send x as a batch of 1 item
x_bw = P(x[0]).convert('L')
x = T(x_bw)[None]
#Black and white input image x, 1x1xHxW
a = torch.Tensor([[1, 0, -1],
[2, 0, -2],
[1, 0, -1]])
a = a.view((1,1,3,3))
G_x = F.conv2d(x, a, padding=1)
b = torch.Tensor([[1, 2, 1],
[0, 0, 0],
[-1, -2, -1]])
b = b.view((1,1,3,3))
G_y = F.conv2d(x, b, padding=1)
G = torch.sqrt(torch.pow(G_x,2)+ torch.pow(G_y,2))
return G_x[0], G_y[0], G[0]
# -
# ### Plot Utils
# +
def plot_image(tup):
img_tensor, depth_tensor = tup
fig, axes = plt.subplots(1, 2, figsize=(10,15))
for i,ax in enumerate(axes.flat):
if(i==0):
plot_image_tensor_in_subplot(ax, img_tensor)
else:
plot_depth_tensor_in_subplot(ax, depth_tensor)
hide_subplot_axes(ax)
plt.tight_layout()
#subplot utils
def hide_subplot_axes(ax):
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
def plot_image_tensor_in_subplot(ax, img_tensor):
im = img_tensor.cpu().numpy().transpose((1,2,0))
#pil_im = Image.fromarray(im, 'RGB')
ax.imshow(im)
def plot_depth_tensor_in_subplot(ax, depth_tensor):
im = depth_tensor.cpu().numpy()
#im = im*255
#im = im.astype(np.uint8)
#pil_im = Image.fromarray(im, 'L')
ax.imshow(im,'gray')
def plot_model_predictions_on_sample_batch(images, depths, preds, plot_from=0, figsize=(12,12)):
n_items=5
fig, axes = plt.subplots(n_items, 3, figsize=figsize)
for i in range(n_items):
plot_image_tensor_in_subplot(axes[i,0], images[plot_from+i])
plot_depth_tensor_in_subplot(axes[i,1], depths[plot_from+i])
plot_depth_tensor_in_subplot(axes[i,2], preds[plot_from+i])
hide_subplot_axes(axes[i,0])
hide_subplot_axes(axes[i,1])
hide_subplot_axes(axes[i,2])
plt.tight_layout()
# -
# ### Model
# +
class Flatten(nn.Module):
def __init__(self):
super(Flatten, self).__init__()
def forward(self, batch):
return batch.view([batch.shape[0], -1])
class Scale1_Linear(nn.Module):
#input 512x7x10
#output 64x15x20
def __init__(self):
super(Scale1_Linear, self).__init__()
self.block = nn.Sequential(
Flatten(),
nn.Linear(512*7*10, 4096),
nn.ReLU(inplace=True),
nn.Dropout(p=0.5),
nn.Linear(4096, 64*15*20)
)
def forward(self, x):
scale_1_op = torch.reshape(self.block(x), (x.shape[0], 64, 15, 20))
return nn.functional.interpolate(scale_1_op, scale_factor=4, mode='bilinear', align_corners=True)
class Scale2(nn.Module):
#input 64x60x80, 3x240x320
#output 1x120x160
def __init__(self):
super(Scale2, self).__init__()
self.input_img_proc = nn.Sequential(
nn.Conv2d(in_channels=3, out_channels=64, kernel_size=9, padding=4, stride=2),
nn.ReLU(inplace=True),
nn.MaxPool2d(kernel_size=2, stride=2)
)
self.block = nn.Sequential(
nn.Conv2d(in_channels=64+64, out_channels=64, kernel_size=5, padding=2),
nn.ReLU(inplace=True),
nn.BatchNorm2d(num_features=64),
nn.Conv2d(in_channels=64, out_channels=64, kernel_size=5, padding=2),
nn.ReLU(inplace=True),
nn.BatchNorm2d(num_features=64),
nn.Conv2d(in_channels=64, out_channels=64, kernel_size=5, padding=2),
nn.ReLU(inplace=True),
nn.BatchNorm2d(num_features=64),
nn.Conv2d(in_channels=64, out_channels=1, kernel_size=5, padding=2)
)
def forward(self, x, input_img):
proc_img = self.input_img_proc(input_img)
concatenate_input = torch.cat((x,proc_img), dim=1)
return nn.functional.interpolate(self.block(concatenate_input), scale_factor=2, mode='bilinear', align_corners=True)
class Scale3(nn.Module):
#input 1x120x160, 3x240x320
#output 1x120x160
def __init__(self):
super(Scale3, self).__init__()
self.input_img_proc = nn.Sequential(
nn.Conv2d(in_channels=3, out_channels=64, kernel_size=9, padding=4, stride=1),
nn.ReLU(inplace=True),
nn.MaxPool2d(kernel_size=2, stride=2)
)
self.block = nn.Sequential(
nn.Conv2d(in_channels=65, out_channels=64, kernel_size=5, padding=2),
nn.ReLU(inplace=True),
nn.BatchNorm2d(num_features=64),
nn.Conv2d(in_channels=64, out_channels=64, kernel_size=5, padding=2),
nn.ReLU(inplace=True),
nn.BatchNorm2d(num_features=64),
nn.Conv2d(in_channels=64, out_channels=1, kernel_size=5, padding=2)
)
def forward(self, x, input_img):
proc_img = self.input_img_proc(input_img)
concatenate_input = torch.cat((x,proc_img), dim=1)
return self.block(concatenate_input)
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.VGG = nn.Sequential(*list(vgg16(pretrained=True).children())[0])
self.Scale_1 = Scale1_Linear()
self.Scale_2 = Scale2()
self.Scale_3 = Scale3()
def forward(self, x):
input_img = x.clone() # 3x240x320
x = self.VGG(x) # 512x7x10
x = self.Scale_1(x) # 64x60x80
x = self.Scale_2(x, input_img.clone()) # 1x120x160
x = self.Scale_3(x, input_img.clone()) # 1x120x160
return x
# -
# ### Loading dataset using dataset class and data loader
# +
bs = 8
sz = (320,240)
mean, std = [0.485, 0.456, 0.406], [0.229, 0.224, 0.225]
mean, std = torch.tensor(mean), torch.tensor(std)
unnormalize = UnNormalizeImgBatch(mean, std)
tfms = transforms.Compose([
ResizeImgAndDepth(sz),
RandomHorizontalFlip(),
ImgAndDepthToTensor(),
NormalizeImg(mean, std)
])
ds = NYUDataset('data/', tfms)
dl = torch.utils.data.DataLoader(ds, bs, shuffle=True)
# -
# ### Mapping the model to our device (to CUDA if available)
model = Net()
model.to(device)
# ### Model Training and parameters
# +
model.train()
n_epochs = 1
lr = 0.0000005
optimizer = torch.optim.Adam(filter(lambda p: p.requires_grad, model.parameters()), lr = lr)
total_steps = 0
for e in range(n_epochs):
torch.cuda.empty_cache()
for batch, labels in dl:
optimizer.zero_grad()
batch = batch.to(device)
labels = labels.to(device)
preds = model(batch)
loss = depth_loss(preds, labels)
loss.backward()
optimizer.step()
total_steps +=1
print_training_loss_summary(loss.item(), total_steps, e+1, n_epochs, len(dl))
# -
# ### Saving the model and presenting the results
# +
#torch.save(model.state_dict(),'all-scales-trained.ckpt')
# -
# %%time
with torch.no_grad():
model.eval()
img, depth = iter(dl).next()
preds = model(img.to(device))
print(img)
# +
plot_model_predictions_on_sample_batch(images=unnormalize(img), depths=depth, preds=preds.squeeze(dim=1), plot_from=0)
test_tfms = transforms.Compose([
transforms.Resize((240,320)),
transforms.ToTensor(),
transforms.Normalize(mean, std)
])
im = Image.open('IMG_4120.jpg').rotate(180)
res = test_tfms(im)
model.eval()
pred = model(res[None].to(device))
pred_numpy_arr = pred.detach().squeeze(dim=1).squeeze(dim=0).cpu().numpy()
pred_numpy_arr = pred_numpy_arr - pred_numpy_arr.min() # -0.660375
pred_numpy_arr = (pred_numpy_arr/pred_numpy_arr.max())*255 # max = 2.1548061, brings scale to 0-255
pred_numpy_arr = pred_numpy_arr.astype('uint8')
import PIL
im.resize((320,240), resample=PIL.Image.BILINEAR)
Image.fromarray(pred_numpy_arr, mode = 'L').resize((320, 240))
# -
from IPython.display import FileLink
FileLink('all-scales-trained.ckpt')
| MDE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2-3.2 Intro Python
# # The Power of List Iteration (loops)
# - for in: **`for`** loop using **`in`**
# - ** for range: `for range(start,stop,step)`**
# - more list methods: **`.extend()`, `+, .reverse(), .sort()`**
# - strings to lists, **`.split()`**, and list to strings, **`.join()`**
#
#
# -----
#
# ><font size="5" color="#00A0B2" face="verdana"> <B>Student will be able to</B></font>
# - Iterate through Lists using **`for`** with **`in`**
# - **Use `for range()` in looping operations**
# - Use list methods **`.extend()`, `+, .reverse(), .sort()`**
# - convert between lists and strings using **`.split()`** and **`.join()`**
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
# ## `range(stop)`
# [](http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/bda2424d-4f25-4c0a-a77a-06384f3da8f2/Unit2_Section3.2a_range_stop.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/bda2424d-4f25-4c0a-a77a-06384f3da8f2/Unit2_Section3.2a_range_stop.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# ### The range(*stop*) function creates a sequence
# using 1 argument with range(*stop*)
# - deault start: 0
# - stop: stopping integer, does not process stop number
# ```python
# for count in range(10):
# print(count)
# ```
# ### same as
# ```python
# for count in [0, 1, 2, 3, 4, 5, 6, 7, 8, 9]:
# print(count)
# ```
#
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
# ### range runs from `0` through the integer before `stop`
# [ ] review and run example
for count in range(10):
print(count)
# +
# review and run example
digits = range(10)
print("digits =", list(digits), "\n")
for count in digits:
print(count)
# -
# [ ] review and run example
sub_total = 0
for item in range(10):
sub_total += item
print("sub_total:", sub_total)
print("Total =", sub_total)
# +
# [ ] review and run example
# print the first half of a spelling list
spell_list = ["Tuesday", "Wednesday", "February", "November", "Annual", "Calendar", "Solstice"]
# find length of 1st half of list (must be int)
half_1 = int(len(spell_list)/2)
for word in range(half_1):
print(spell_list[word])
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 1</B></font>
#
# ## `range(stop)`
# +
# [ ] for x = 6, use range(x) to print the numbers 1 through 6
x = 6
for number in range(x):
print(number)
# +
# [ ] using range(x) multiply the numbers 1 through 7
# 1x2x3x4x5x6x7 = 5040
#task cannot be done just using range(stop), so I change multiplying by addition
total = 0
for x in range(8):
total += x
print(total)
# -
# Use **`range(stop)`** to print the second half of spell_list below
# +
# [ ] print the second half of a spelling list using a range(stop) loop to iterate the list
spell_list = ["Wednesday", "Tuesday", "February", "November", "Annual", "Calendar", "Solstice"]
half_of_list = int(len(spell_list) / 2)
for item_id in range(half_of_list):
print(spell_list[item_id])
# -
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
# ## `range(start,stop)`
# [](http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/95d6c75a-ed37-4f50-9049-2a2c225f9499/Unit2_Section3.2b_range_start_stop.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/95d6c75a-ed37-4f50-9049-2a2c225f9499/Unit2_Section3.2b_range_start_stop.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# ### The range(*start,stop*) function creates a sequence
# using 2 arguments with range(*start,stop*)
# - start: starting integer value of a range loop
# - stop: stopping integer (second argument), does not process stop number
# ```python
# for count in range(5,10):
# print(count)
# ```
#
# ###
# <font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
# ### range runs from `start` integer through the integer before `stop`
# [ ] review and run example
for count in range(5,10):
print(count)
# [ ] review and run example
sub_total = 0
temp = 0
for item in range(5, 11):
temp = sub_total
sub_total += item
print("sub_total:", temp, "+", item, "=",sub_total)
print("Total =", sub_total)
# +
# [ ] review and run example
spell_list = ["Tuesday", "Wednesday", "February", "November", "Annual", "Calendar", "Solstice"]
# find length list
spell_len = len(spell_list)
# find lenght of 1st half (aka - start of 2nd half)
half_1 = int(spell_len/2)
# print 2nd half list
for word in range(half_1,spell_len):
print(spell_list[word])
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 2</B></font>
#
# ## `range(start,stop)`
# +
# [ ] using range(start,stop), .append() the numbers 5 to 15 to the list: five_fifteen
# [ ] print list five_fifteen
five_fifteen = []
for number in range(5,15):
five_fifteen.append(number)
print(five_fifteen)
# +
# [ ] using range(start,stop) - print the 3rd, 4th and 5th words in spell_list
# output should include "February", "November", "Annual"
spell_list = ["Tuesday", "Wednesday", "February", "November", "Annual", "Calendar", "Solstice"]
for item_id in range(2,5):
print(spell_list[item_id])
# +
# [ ] using code find the index of "Annual" in spell_list
# [ ] using range, print the spell_list including "Annual" to end of list
spell_list = ["Tuesday", "Wednesday", "February", "November", "Annual", "Calendar", "Solstice"]
index_of_annual = spell_list.index('Annual')
for item_id in range(index_of_annual, len(spell_list)):
print(spell_list[item_id])
# -
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
# ## `range(start,stop,step)`
# [](http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/4299f0e2-3dc2-4298-aff1-e2a0b013de6a/Unit2_Section3.2c_range_start_stop_step.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/4299f0e2-3dc2-4298-aff1-e2a0b013de6a/Unit2_Section3.2c_range_start_stop_step.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# ### The range(*start,stop,step*) function creates a sequence
# using 3 arguments with range(*start,stop,step*)
# - start: starting integer value of a range loop
# - stop: stopping integer (second argument), does not process stop number
# - step: skip value for each loop
# ```python
# for count in range(10,101,10):
# print(count)
# ```
#
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
# ### range runs from `start` integer, skipping by `step`, through the largest `step` integer before reaching `stop`
# [ ] review and run example
for count in range(25,101,25):
print(count)
# [ ] review and run example
sub_total = 0
temp = 0
for item in range(25,46,5):
temp = sub_total
sub_total += item
print("sub_total:", temp, "+", item, "=",sub_total)
print("Total =", sub_total)
# +
# [ ] review and run example printing the 1st and then every other word in spell_list
spell_list = ["Tuesday", "Wednesday", "February", "November", "Annual", "Calendar", "Solstice"]
for index in range(0,len(spell_list),2):
print(spell_list[index])
# -
# [ ] review and run example casting range to list
odd_list = list(range(1,20,2))
print(odd_list)
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 3</B></font>
# ## `range(start,stop,step)`
# +
# [ ] print numbers 10 to 20 by 2's using range
for number in range(10,20,2):
print(number)
# +
# [ ] print numbers 20 to 10 using range (need to countdown)
# Hint: start at 20
for number in range(20,10,-2):
print(number)
# +
# [ ] print first and every third word in spell_list
spell_list = ["Tuesday", "Wednesday", "February", "November", "Annual", "Calendar", "Solstice"]
for item_id in range(0, len(spell_list), 3):
print(spell_list[item_id])
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 4</B></font>
#
# ### Program: List of letters
# - Input a word string (**word**)
# - find the string length of word
# - **use range()** to iterate through each letter in word (can use to range loops)
# - Save odd and even letters from the word as lists
# - **odd_letters**: starting at index 0,2,...
# - **even_letters**: starting at index 1,3,...
# - print odd and even lists
# +
# [ ] complete List of letters program- test with the word "complexity"
word = input("enter a word string: ")
word_len = len(word)
odd_letters = []
even_letters = []
for ltr_id in range(0, word_len, 2):
odd_letters += word[ltr_id]
for ltr_id in range(1, word_len, 2):
even_letters += word[ltr_id]
print("the odd letter indexes are", odd_letters)
print("the even letter indexes are", even_letters)
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 5: fix the error</B></font>
#
# [ ] fix the error printing odd numbers 1 - 9
for num in range(1,10,2):
print(num)
# [Terms of use](http://go.microsoft.com/fwlink/?LinkID=206977) [Privacy & cookies](https://go.microsoft.com/fwlink/?LinkId=521839) © 2017 Microsoft
| edx/Dev274x_Introduction_to_Python_Unit_2/Mod3_2-3.2_intro_Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Los datos son baratos pero el conocimiento es más difícil de conseguir
# ## Estadística Descriptiva
#
# Empezar a entender mis datos.
# ### Media
#
# Si se tiene una muestra de $n$ valores: $x_i$
# La media $\mu$ es la suma de los valores dividido por el número de valores
#
# $$ \mu = \frac{1}{n} \sum_{i}^{n} x_i $$
# +
import pandas as pd
import numpy as np
data = pd.read_csv("train.csv")
media_edad = np.mean(data['Age'])
media_edad
# -
data.describe()
# La media se encarga de describir la tendencia central de nuestros datos.
# ¡Importante!, esta media $\mu$ se usa para describir a una población completa.
# ### Varianza
#
# Otro valor estadístico que nos ayuda a entender nuestros datos es la Varianza. A diferencia de la media que describe la tendencia de en donde se centran nuestros datos, la varianza describe que tan lejos se encuentran los datos de la media.
#
# $$ \sigma^2 = \frac{1}{n} \sum_{i}^{n} (x_i - \mu)^2 $$
varianza_edad = np.var(data['Age'])
varianza_edad
# ¿Años al cuadrado?
# La varianza es difícil de interpretar debido a las unidades.
#
# Por suerte la desviación estándar es un estadístico más significativo.
#
# ### Desviación estándar
#
# $$ \sigma = \sqrt{\sigma} $$
desviacion_edad = np.std(data['Age'])
desviacion_edad
data.Age.std()
# ¡Importante!, estas formulas para $\sigma^2$ y $\sigma$ se usan para describir a una población completa.
#
# Si lidiamos con una muestra de N valores se usan estimadores, $\bar{x}$ y $S^2$
#
# $$ \bar{x} = \frac{1}{N} \sum_{i}^{N} x_i $$
#
# $$ S^2 = \frac{1}{N-1} \sum_{i}^{N} (x_i - \bar{x})^2 $$
# ## Distribuciones
#
# La media, la varianza y la desviación estándar son estadísticos concisos, pero también peligrosos, ya que nublan la información que nos proporcionan los datos.
#
# Un apoyo para entenderlos mejor es ver la distribución de los datos.
#
# La representación más común de una distribución es un histograma, que describe frecuencia con la que aparece cada valor.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
edades = data[data.Age.notnull()]['Age']
edades.hist()
plt.xlabel('Edad', fontsize=18)
plt.ylabel('Frecuencia', fontsize=16)
# -
len(edades.unique())
edades = data[data.Age.notnull()]['Age']
edades.hist(bins=88)
plt.xlabel('Edad', fontsize=18)
plt.ylabel('Frecuencia', fontsize=16)
# Los histogramas son útiles porque podemos revisar las siguientes características rápidamente:
#
# - Moda: El valor más común o que más se repite en una distribución se llama moda.
#
# - Forma: Alrededor de la moda podemos ver que la distribución es asimétrica.
#
# - Los valores atípicos. (outliers)
# ### Función de probabilidad
#
# Si queremos transformar las frecuencias a una función de probabilidad debemos dividir la serie entre el número de elementos
edades.hist(bins=88, normed=True)
plt.xlabel('Edad', fontsize=18)
plt.ylabel('Probabilidad', fontsize=16)
# $$ P(X = x) = f(x) $$
# La función de probabilidad funciona bien si el número de valores es pequeño.
#
# Pero a medida que el número de valores aumenta, la probabilidad asociada a cada valor se hace más pequeño.
#
# Una alternativa es usar la función de distribución acumulada.
edades.describe()
# ### Función de distribución acumulada
edades.hist(cumulative=True, bins=88, normed=True)
# $$ P(X \leq x) = f(x) $$
# ### Distribuciones continuas
#
# Cuando tenemos variables aleatorias continuas
#
# La distribución normal (Gaussiana), es una de las distribuciones de probabilidad que con más frecuencia aparece aproximada en fenómenos reales.
# $$ P(x, \sigma, \mu) = \frac{1}{\sigma\sqrt{2 \pi}}e^{-(x -\mu)^2/2\sigma^2}$$
# +
import numpy as np
x = np.random.randn(5000)
# Make a normed histogram. It'll be multiplied by 100 later.
plt.hist(x, bins=50, normed=True)
# -
# <img src="https://upload.wikimedia.org/wikipedia/commons/thumb/8/8c/Standard_deviation_diagram.svg/2000px-Standard_deviation_diagram.svg.png">
# ### ¿Por qué usar distribuciones continuas?
#
# Como todos los modelos, las distribuciones continuas son abstracciones, lo que significa que pueden simplificar y deshacerse de los detalles que se consideran irrelevantes (Errores de medición, outliers).
#
# Además son una forma de comprimir los datos. Ya que si logramos ajustar un modelo a un conjunto de datos, un pequeño conjunto de parámetros puede resumir una gran cantidad de datos.
# ### ¿Por qué es tan importante la distribución Normal?
#
# El teorema de límite central establece que la media de la muestra $\bar{X}$ sigue una distribución normal (para $n$ grandes)
# con media $\mu$ y desviación estándar $\frac{\sigma}{\sqrt(n)}$
#
# El teorema del límite central explica, porque aparece con tanta frecuencia la distribución normal en el mundo natural.
#
# La mayoría de las características de los animales y otras formas de vida se ven afectadas por un gran número de variables genéticas y ambientales cuyo efecto es aditivo.
#
# Las características que medimos son la suma de un gran número de pequeños efectos, por lo que su distribución tiende a ser normal.
# +
#Prueba para el Teorema de Limite Central usando 50
media_muestra = [] #Iniciamos una lista
for x in range(0, 50):
media_muestra.append(np.mean(edades.sample(n=300)))
media_muestra = pd.Series(media_muestra)
media_muestra.hist(bins=50)
# +
#Prueba para el Teorema de Limite Central usando 500
media_muestra = []
for x in range(0, 500):
media_muestra.append(np.mean(edades.sample(n=300)))
media_muestra = pd.Series(media_muestra)
media_muestra.hist(bins = 50)
# +
#Prueba para el Teorema de Limite Central usando 5000
media_muestra = []
for x in range(0, 5000):
media_muestra.append(np.mean(edades.sample(n=300)))
media_muestra = pd.Series(media_muestra)
media_muestra.hist(bins = 50)
# -
# ### Probabilidad
#
# Anteriormente mencionamos que la probabilidad es la frecuencia expresada como una fracción tamaño de muestra.
#
# Esa es una definición de probabilidad, pero no es la única y de hecho, el significado de probabilidad es un tema controversial.
#
# Existe un consenso general de que la probabilidad es un valor real entre 0 y 1. Este valor pretende dar una medida cuantitativa que corresponde a la noción de que algunas cosas son más probables que otras.
#
# $$ P(E) \epsilon [0,1] $$
# ### Reglas de probabilidad (Recordando a Kolmogorov)
#
# - La probabilidad de que ocurra un evento es un valor entre 0 y 1. Para todo evento existe una probabilidad
# $$ 0 \leq P(E) \leq 1 $$
# - La probabilidad de que nada ocurra es 0
# $$ P(\emptyset) = 0 $$
# - La probabilidad de que algo ocurra es 1
# $$ P(\Omega) = 1 $$
# - La probabilidad de algo es 1 menos la probabilidad de lo contrario
# <img src="conditional_risk.png">
# ### Probabilidad condicional
#
# $$ P(A | B) = \frac{P(A \cap B)}{P(B)} $$
#
# Si A y B son eventos independientes entonces:
#
# $$ P(A | B) = \frac{P(A) P(B)}{P(B)} = P(A) $$
# ##### <NAME>
from IPython.display import HTML
HTML('<iframe width="560" height="315" src="https://www.youtube.com/embed/mhlc7peGlGg?rel=0&controls=0&showinfo=0" frameborder="0" allowfullscreen></iframe>')
# ### Regla de Bayes
#
# El teorema de Bayes es a menudo interpretado como una declaración acerca de cómo la evidencia, E, afecta la probabilidad de una hipótesis, H:
#
# $$P(H | E) = P(H) \frac{P(E|H)}{P(E)}$$
#
# En palabras, esta ecuación dice que la probabilidad de H después de haber visto E es el producto de $P(H)$, que es la probabilidad de que H antes de ver la evidencia E, y la relación de $P(E|H)$, la probabilidad de ver la evidencia suponiendo que H es verdadera, y $P(E)$, la probabilidad de ver la evidencia bajo cualquier circunstancia.
#
# Ejemplo: Filtro de Spam
#
# $$ P(S|W) = \frac{P(W|S) \cdot P(S)}{P(W|S) \cdot P(S) + P(W|H) \cdot P(H)} $$
#
# donde:
#
# - $P(S|W)$ Es la probabilidad de que nuestro mensaje sea SPAM, sabiendo que encontramos la palabra "Dinero"
# - $P(S)$ Es la probabilidad de que cualquier mensaje sea SPAM
# - $P(W|S)$ La probabilidad de que nuestra palabra aparezca en mensajes de SPAM
# - $P(H)$ La probabilidad de que nuestro mensaje sea HAM
# - $P(W|H)$ La probabilidad de que la nuestra palabra aparezca en HAM
#
#
#
#
HTML('<iframe width="560" height="315" src="https://www.youtube.com/embed/R13BD8qKeTg?rel=0&controls=0&showinfo=0" frameborder="0" allowfullscreen></iframe>')
| Python_para_ciencia_de_datos/2da_clase/Estadistica_24_Jun.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Bertha-ding/20MA573-yuning-ding/blob/master/src/prj03.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="b5jwBMLkpY3t"
# ## Abstract
#
# - Goal: Learn python basics on Jupyter notebook.
# - Ref:
# - Python crash course (Google for it)
# - the [naming section](https://github.com/google/styleguide/blob/gh-pages/pyguide.md#316-naming) of Google's python style guide
# + [markdown] colab_type="text" id="dB34OHqzpCXj"
# ## Python on Jupyter notebook - Colab
#
# Python on Jupyter notebook allows one to perform a great deal of data analysis and statistical validation. cell by cell in a interactive way. A good online platform is colab, powered by google. Another good cloud platform is azure notebook, powered by microsoft. One can also install anaconda with python 3 on local computer to run Jupyter notebook.
#
# - open a new python3 notebook by visiting https://colab.research.google.com/
# - As you can see, each cell can be chosen either code or text from its top menu.
# - A code cell will be evaluated when you press play, or when you press the shortcut, shift-enter.
# + colab_type="code" id="Ovthtoi0pCXn" outputId="dedb8c76-970f-4e3c-bbbb-9fae6ef8b9ae" colab={"base_uri": "https://localhost:8080/", "height": 34}
2 + 2 #Executing a Command
# + [markdown] colab_type="text" id="yRrPz4YgpCX5"
# ## Importing Libraries
#
# The vast majority of the time, you'll want to use functions from pre-built libraries. You can't import every library each time, but you can import most of the common scientific ones. Here I import numpy, pandas, and matplotlib, the most common and useful libraries.
#
# Notice that you can rename libraries to whatever you want after importing. The `as` statement allows this. Here we use `np` and `pd` as aliases for `numpy` and `pandas`. This is a very common aliasing and will be found in most code snippets around the web. The point behind this is to allow you to type fewer characters when you are frequently accessing these libraries.
# + colab_type="code" id="acJx6F4IpCX7" colab={}
import numpy as np
import matplotlib.pyplot as plt
# + [markdown] colab_type="text" id="AJGP6N4NxpIS"
# ## Using numpy and matplotlib: A sampling example
# + id="nPic_DSV588e" colab_type="code" colab={}
# + [markdown] colab_type="text" id="Gzg6f5unyPbz"
# Let's use numpy to select 100 standard normal radom variables, and make a plot.
# + colab_type="code" id="Tm_bHr2lpCYP" outputId="6054c813-511a-47e9-c3c0-e2b63a6065fa" colab={"base_uri": "https://localhost:8080/", "height": 68}
import numpy as np
import matplotlib.pyplot as plt
n=100000000
X = np.random.normal(0, 1, n)
print('>>>mean is:'+str(np.mean(X)))
print('>>>std is:'+str(np.std(X)))
'''
plt.plot(X);
plt.xlabel('Time')
plt.ylabel('Returns')
plt.show()
'''
# + id="XvhQb9Rr70cy" colab_type="code" colab={}
def fun(n):
X = np.random.normal(0,1,n)
# + [markdown] colab_type="text" id="MGcHGVJzpCYR"
# Let's use `numpy` to take some simple statistics.
# + colab_type="code" id="ASD-yJbypCYT" outputId="43375791-9bf5-46fa-fe73-8958f7dabd91" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.mean(X)
# + colab_type="code" id="3L6v9sf_pCYX" outputId="748cccdb-4357-47f0-8efc-054237f8e7ff" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.std(X)
# + colab_type="code" id="-2TRwHgTz5U1" colab={}
| src/prj03.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
import sys
sys.path.append('../..')
sys.path.append('../../../../transformer_pytorch')
# %load_ext autoreload
# %autoreload 2
# +
from rdkit import Chem
from rdkit.Chem import AllChem, Descriptors
from rdkit.Chem.Draw import MolToImage, MolToFile, _moltoSVG, MolsToGridImage
from collections import OrderedDict
smiles = OrderedDict([('CC1=CC=C(C=C1)C1=CC(=NN1C1=CC=C(C=C1)S(N)(=O)=O)C(F)(F)F', '1: Celecoxib'),
('Cc1c(C)c2OC(C)(COc3ccc(CC4SC(=O)NC4=O)cc3)CCc2c(C)c1O', '2: Troglitazone'),
])
for smile, label in smiles.items():
mol = Chem.MolFromSmiles(smile)
print(label)
display(MolToImage(mol, kekulize=True))
# -
import numpy as np
from generative_playground.rdkit_utils.rdkit_utils import num_atoms, num_aromatic_rings, num_aliphatic_rings, NormalizedScorer
scorer = NormalizedScorer()
[np.sum(x) for x in scorer.get_scores([kusner3])]
MolToFile(mol, 'test.svg', imageType='svg')
# +
from generative_playground.models.model_settings import get_decoder, get_settings
max_len = 40
settings = get_settings(True, 'new')
# Now let's run a random model with a grammar
model, _ = get_decoder(True,
'new',
feature_len=settings['feature_len'],
max_seq_length=max_len,
decoder_type='random',
batch_size=9)
# +
from generative_playground.codec.grammar_codec import GrammarModel
settings = get_settings(True, 'new')
codec = GrammarModel(grammar=settings['grammar'],
tokenizer=settings['tokenizer'])
actions = model()[0].detach().cpu().numpy()
#print(actions, settings['feature_len'])
my_smiles = codec.decode_from_actions(actions)
print(my_smiles)
mols = [Chem.MolFromSmiles(my_smile) for my_smile in my_smiles]
print([Descriptors.NumAromaticRings(m) for m in mols if m is not None])
MolsToGridImage(mols, kekulize=False)
# +
#my_smiles.append'c1ccncn1']
def parse(x):
# print(x)
try:
return next(codec._parser.parse(x))
except Exception as e:
# print('fail!')
# print(e)
return None
tokens = [codec._tokenize(s) for s in my_smiles]
parse_trees = [parse(t) for t in tokens]
print(parse_trees, len(parse_trees))
# +
fname = '../data/250k_rndm_zinc_drugs_clean.smi'
with open(fname) as f:
smiles = f.readlines()
for i in range(len(smiles)):
smiles[i] = smiles[i].strip()
# -
tokens = [codec._tokenize(s) for s in smiles[:1000]]
parse_trees = [parse(t) for t in tokens]
parse_trees = [p for p in parse_trees if p is not None]
# +
from nltk.tree import *
# tree = parse_trees[0]
# print(tree.label(), tree)
rpe_dict ={}
def get_rpe(tree, rpe_dict):
these_tuples = [(tree.label(), loc, child.label()) for loc,child in enumerate(tree) if isinstance(child,Tree)]
for t in these_tuples:
if t in rpe_dict:
rpe_dict[t] += 1
else: rpe_dict[t] = 1
for subtree in tree:
if isinstance(subtree, Tree):
rpe_dict = get_rpe(subtree, rpe_dict)
return rpe_dict
for tree in parse_trees:
rpe_dict = get_rpe(tree,rpe_dict)
#print(rpe_dict)
rpe_count = [(key, value) for key, value in rpe_dict.items()]
rpe_count = sorted(rpe_count, key = lambda x:x[1],reverse = True)
print(len(parse_trees))
for x in rpe_count:
print(x)
# -
dashsmiles = [x for x in smiles if '-c' in x]
print(len(dashsmiles))
mols = [Chem.MolFromSmiles(s.replace('-c','c')) for s in dashsmiles]
mols =[m for m in mols if m is not None]
print(len(mols))
MolToImage(mols[2], kekulize=False)
tokens = [codec._tokenize(s.replace('-c','c')) for s in smiles[:1000]]
parse_trees = [parse(t) for t in tokens]
nice = [ip for ip, p in enumerate(parse_trees) if p is not None]
print(len(nice), nice)
i = 3
# +
# C[C@@H]1CC(Nc2cncc(-c3nncn3C)c2)C[C@@H](C)C1
#N#Cc1ccc(-c2ccc(O[C@@H](C(=O)N3CCCC3)c3ccccc3)cc2)cc1
#C[C@@H]1CC(Nc2cncc(-c3nncn3C)c2)C[C@@H](C)C1
#smile = 'Nc2cncc(-c3nncn3C)c2'
smile = smiles[i]
smile = 'C1CNCCC1'
print(i, smile)
mol = Chem.MolFromSmiles(smile)
parse_tree = parse(codec._tokenize(smile))
if parse_tree is None:
print('fail!')
MolToImage(mol, kekulize=False)
# -
# i+=1
# my_smile =smiles[i]
# mols = [Chem.MolFromSmiles(my_smile)]
# print(i, my_smile, [Descriptors.NumAromaticRings(m) for m in mols])
# MolsToGridImage(mols, kekulize=False)
# +
# from generative_playground.models import grammar_ed_models as grammar_model
# # We load the auto-encoder
# grammar_weights = '../pretrained/my_molecules.mdl'
# grammar_model = grammar_model.ZincGrammarModel(grammar_weights)
# z = grammar_model.encode(['c1nccc2n1ccc2'])
# new_smile = grammar_model.decode(z)
| src/generative_playground/molecules/notebooks/GuacamolPics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Step 1. EDA cate & preprocess
# -----
# ### read json data
import json
data_path = "../../../dataset/"
json_file = "cate1.json"
json_data=open(data_path + json_file).read()
data = json.loads(json_data)
print("big cate cnt :", str(len(data['b'])))
print("middle cate cnt :", str(len(data['m'])))
print("small cate cnt :", str(len(data['s'])))
print("detail cate cnt :", str(len(data['d'])))
# ### read hdf5 data
import pandas as pd
import numpy as np
import h5py
hdf5_file = "train.chunk.01"
hdf5_data = h5py.File(data_path + hdf5_file, 'r')
a_group_key = list(hdf5_data.keys())[0]
a_group_key
list(hdf5_data['train'].keys())
# ### hd5f to dataframe
idx = 1000000
hdf5_data['train']['bcateid'][0:idx]
np.array([x.decode('utf-8') for x in hdf5_data['train']['brand'][0:idx]])
hdf5_data['train']['price'][0:idx]
# ### make whole train sample
def make_df(hdf5_data, idx):
df = pd.DataFrame(
{'bcateid': hdf5_data['train']['bcateid'][0:idx],
'mcateid': hdf5_data['train']['mcateid'][0:idx],
'scateid': hdf5_data['train']['scateid'][0:idx],
'dcateid': hdf5_data['train']['dcateid'][0:idx],
'brand': np.array([x.decode('utf-8') for x in hdf5_data['train']['brand'][0:idx]]),
'maker': np.array([x.decode('utf-8') for x in hdf5_data['train']['maker'][0:idx]]),
'model': np.array([x.decode('utf-8') for x in hdf5_data['train']['model'][0:idx]]),
'product': np.array([x.decode('utf-8') for x in hdf5_data['train']['product'][0:idx]]),
'price': hdf5_data['train']['price'][0:idx],
'updttm': hdf5_data['train']['updttm'][0:idx],
'pid': hdf5_data['train']['pid'][0:idx]
})
return df
columns = ['bcateid', 'mcateid', 'scateid', 'dcateid', 'brand', 'maker', 'model',
'product', 'price', 'updttm', 'pid']
df = pd.DataFrame(columns=columns)
for number in range(1,10):
hdf5_file = "train.chunk." + "0" + str(number)
hdf5_data = h5py.File(data_path + hdf5_file, 'r')
chunk_df = make_df(hdf5_data, 1000000)
df = df.append(chunk_df)
file_name = data_path + "train_sample.csv"
df.to_csv(file_name, index=False, header=True)
| yamarae/eda/01_make_whole_train_df.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Excercises Electric Machinery Fundamentals
# ## Chapter 1
# ## Problem 1-5
# + slideshow={"slide_type": "skip"}
# %pylab notebook
# %precision %.4g
from scipy import constants as c # we like to use some constants
# -
# ### Description
# A ferromagnetic core is shown in Figure P1-2:
# <img src="figs/FigC_P1-2.jpg" width="50%">
# The depth of the core is 5 cm. The other dimensions of the core are as shown in the figure. Find the value of the current that will produce a flux of:
phi = 0.005 # Wb
# With this current,
#
# * What is the flux density at the top of the core?
# * What is the flux density at the right side of the core?
#
# Assume that the relative permeability of the core is:
mu_r = 800
mu = mu_r * c.mu_0
# The magnetic constant $\mu_0$ is available from `scipy.constants` (see also import statement) and is simply:
c.mu_0
# ### SOLUTION
# There are three regions in this core. The top and bottom form one region, the left side forms a
# second region, and the right side forms a third region. If we assume that the mean path length of the flux
# is in the center of each leg of the core, and if we ignore spreading at the corners of the core, then the path
# lengths are:
l1 = 2 * 0.275 # m
l2 = 0.3 # m
l3 = 0.3 # m
# The reluctances of these regions are: $\mathcal{R} = \frac{l}{\mu_0 \mu_r A}$. The areas can be calculated as:
A1 = 0.05 * 0.15 # m^2
A2 = 0.05 * 0.10 # m^2
A3 = 0.05 * 0.05 # m^2
# And the reluctances are hence:
R1 = l1 / (mu * A1) # At /Wb = At/Vs
R2 = l2 / (mu * A2) # At /Wb = At/Vs
R3 = l3 / (mu * A3) # At /Wb = At/Vs
print('R1 = {:.1f} kAt/Wb'.format(R1/1000) )
print('R2 = {:.1f} kAt/Wb'.format(R2/1000) )
print('R3 = {:.1f} kAt/Wb'.format(R3/1000) )
# The total reluctance is thus $\mathcal{R}_\text{TOT} = \mathcal{R}_1 + \mathcal{R}_2 + \mathcal{R}_3$:
Rtot= R1 + R2 + R3
print('Rtot = {:.1f} kAt/Wb'.format(Rtot/1000) )
# and the magnetomotive force required to produce a flux of 0.005 Wb is $\mathcal{F} = \phi \mathcal{R}_\text{TOT}$:
F = phi * Rtot
print('F = {:.1f} At'.format(F) )
# and the required **current is $i = \frac{\mathcal{F}}{N}$**:
N = 500 # given in Figure P1-2
i = F/N
print('''
i = {:.1f} A
========='''.format(i))
# **The flux density $B = \frac{\phi}{A}$ on the top of the core is:**
B1 = phi / A1
print('''
B1 = {:.2f} T
==========='''.format(B1))
# **The flux density $B = \frac{\phi}{A}$ at the right side of the core is:**
B3 = phi / A3
print('''
B3 = {:.1f} T
=========='''.format(B3))
| Chapman/Ch1-Problem_1-05.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# DL01
# 딥러닝 기초 - 텐서플로, 케라스
# 딥러닝 개발환경
# 1. 파이썬 3.5 이상 또는 아나콘다 4.4이상
# 2. 파이참 - 파이썬 venv 환경에서 실행하도록 설정
# 3. 머신러닝/딥러닝 관련 패키지 설치
# numpy, scipy, matplotlib, pandas, scikit-learn
# spyder(과학계산), seaborn(시각화), h5py(hdf5), pillow(이미지)
# tensorflow (tensorflow-gpu) keras
# 4. 텐서플로우 GPU 지원 사이트
# CUDA 설치 - developer.nvidia.com/cuda-downloads
# cuDNN 설치 - developer.nvidia.com/cudnn
# 5. 설치 확인
# 파이참 - 파이썬 콘솔에서 다음 실행
import tensorflow as tf
print(tf.__version__) # 텐서플로우 버젼 확인
import keras
hello = tf.constant('Hello, TensorFlow!!')
sess = tf.Session() # 텐서플로우 작업 생성
print(sess.run(hello)) # 텐서플로우 작업 실행
# +
# 인공지능 - 관념적으로 컴퓨터가 인간이 사고를 모방하는 것
# 즉, 기계가 인간처럼 사고하고 행동하게 하는 것
# 머신러닝 - 주어진 데이터를 통해 컴퓨터가 스스로 학습하는 것
# 학습 : 데이터를 입력해서 패턴을 분석하는 과정
# 머신러닝의 한계 - 인간도 파악하기 어려운 복잡한 문제는
# 머신러닝으로도 풀기 어려움(이미지 인식)
# 딥러닝 - 인공신경망을 이용해서 컴퓨터가 스스로 학습하는 것
# 인공신경망 : 인간의 뇌의 동작방식을 착안해서 만듦
# 2012년 ImageNet이라는 1,000개의 카테고리로
# 분류된 100만개의 이미지를 인식하여 정확성을 겨루는
# ILSVRC라는 이미지 인식대회에서 84.7%라는 인식률 달성
# 그 전까지는 75%대 였음 - 현재는 97%에 육박할 정도
# 인공신경망은 이미 1940년대 부터 연구되던 기술
# 그전까지는 여러가지 문제에 부딪혀 암흑의 시대를 지나다가
# 빅데이터와 GPGU의 발전, 수백만에서 수조개로 이뤄진 아주
# 간단한 수식을 효율적으로 실행하게 해주는 딥러닝 알고리즘의
# 발명 덕택에 급격히 발전되고 있음
# 텐서플로우는 머신러닝 프로그램, 특히 딥러닝 프로그램을 아주
# 쉽게 구현할 수 있도록 다양한 기능을 제공하는 머신러닝 라이브러리
# 구글에서 제작하고 배포하고 있음
# 케라스, 카페, 토치, MXNet, 체이너, CNTK
# 텐서플로우를 좀 더 사용하기 쉽게 만들어 주는 보조 라이브러리
# +
# 인공신경망 구동 예제
# x1 = '여친에게 데이트 요청 전화옴 : 0.5'
# w1 = '여친을 너무 좋아한다 : 7 ' # 가중치
# w1 = '여친과 냉전 상태 : 0.1' # 가중치
# x2 = '지금 밖에는 눈이 온다 : 0.3'
# w2 = '난 눈을 맞는 것을 좋아해 : 5' # 가중치
# w2 = '난 눈에 맞아 젖는 걸 싫어해 : 1' # 가중치
# x3 = '난 지금 점심을 안 먹었는데'
# w3 = '난 배고픈 거 싫어' # 가중치
# w3 = '여친을 만난다면 배고픈것 쯤이야' # 가중치
# y = 0 : '그냥 집에 있는다', 1 : '여친과 데이트한다'
# 0.5(여친에게 데이트 요청 전화 옴) * 0.1(여친과 냉전 상태)
# # + 0.3(지금 밖에는 눈이 온다) * 1(난 눈에 맞아 젖는걸 싫어해) = 0.35 = 0(그냥 집에 있는다)
# +
# 딥러닝 간단 예제
# 2013년 폴란드 브로츠와프 의과대학
# 폐암환자의 수술전 진단정보와 수술후 생존여부
# 딥러닝을 구동하는 필요한 케라스 함수를 불러옴
from keras.models import Sequential
from keras.layers import Dense
# 머신러닝 관련 라이브러리 불러옴
import numpy as np
import tensorflow as tf
# 난수 생성을 위한 초기화
seed = 9563
np.random.seed(seed)
tf.set_random_seed(seed)
# +
# 준비된 환자 정보를 불러옴
# 종양유형, 폐활량, 호흡곤란여부, 고통정도, 기침, 흡연, 천식여부
data_set = np.loadtxt('data/ThoraricSurgery.csv',delimiter=',')
# 환자기록과 수술결과를 x, y로 구분해서 저장
x = data_set[:,0:17]
y = data_set[:,17]
# 딥러닝 실행 방식을 결정(모델 설정 및 실행방법 정의)
model = Sequential()
# 입력데이터는 17, 은닉층 갯수는 30, 적용 알고리즘 함수는 relu
model.add(Dense(30, input_dim=17,activation='relu'))
# 출력데이터는 1, 적용 알고리즘 함수는 sigmoid
model.add(Dense(1, activation='sigmoid'))
# 딥러닝을 실행
model.compile(loss='mean_squared_error',optimizer='adam',metrics=['accuracy'])
# loss는 오차값 추적 방식, optimizer는 오차 수정 함수
model.fit(x,y, epochs=30, batch_size=10)
# 결과 검증 및 출력
print('정확도 : %.4f' % (model.evaluate(x,y)[1]))
# -
| DL01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="u9_gUhWydkuZ" outputId="3c71b858-1714-41fe-c247-27b3994b7d54"
# !pip install transformers
# !pip install datasets
# + colab={"base_uri": "https://localhost:8080/"} id="am9GnXW7-7l2" outputId="1b4cf2a8-68f4-41dd-8317-5f2f48add9df"
# !pip install git+https://github.com/huggingface/transformers
# + [markdown] id="q-Mh3z5jQAG7"
# ### Clone github repo for transformers
# + colab={"base_uri": "https://localhost:8080/"} id="bxfECSx0MSrH" outputId="39c04236-66cb-4f1e-f64c-b5248f49c461"
# !git clone https://github.com/huggingface/transformers.git
# + colab={"base_uri": "https://localhost:8080/"} id="wefEYhWYPiZV" outputId="e2b6e5d9-9d20-4037-b4e0-a685addcef85"
# cd /content/transformers/examples/language-modeling
# + colab={"base_uri": "https://localhost:8080/"} id="ZB83L626X5Tl" outputId="58906ab8-9edc-489d-caae-60caac9cd3b9"
# !python run_mlm.py \
# --model_name_or_path bert-base-cased \
# --train_file /content/train_preprocessed.txt \
# --validation_file /content/dev_preprocessed.txt \
# --do_train \
# --do_eval \
# --output_dir /content/bert_model
# + [markdown] id="71Sq8RNgrz4V"
# ### Now use pytorch_model.bin (model weights from language model training) to fine tune our classification task.
# + id="KAzNPAA3a2Gu" colab={"base_uri": "https://localhost:8080/"} outputId="d63126dd-4e6e-4b10-fb59-33e2178b7e14"
# import required libraries
import pandas as pd
import numpy as np
import time
import datetime
import seaborn as sns
from matplotlib import rc
from pylab import rcParams
import matplotlib.pyplot as plt
from textwrap import wrap
from collections import defaultdict
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import confusion_matrix, classification_report
from sklearn.metrics import precision_recall_curve,auc,f1_score
from statistics import mean
import transformers
from transformers import BertModel, BertTokenizer, BertForSequenceClassification,BertConfig
from transformers import AdamW, get_linear_schedule_with_warmup
import torch
from torch import nn,optim
from torch.utils.data import Dataset,DataLoader,TensorDataset, RandomSampler, SequentialSampler
import torch.nn as nn
import torch.nn.functional as F
device=torch.device('cuda:0' if torch.cuda.is_available() else 'cpu');
# %matplotlib inline
# %config InlineBackend.figure_format='retina'
sns.set(style='whitegrid',palette='muted',font_scale=1.2)
color_palette=['#01BEFE','#FFDD00','#FF7D00','#FF006D','#ADFF02','#8F00FF']
sns.set_palette(sns.color_palette(color_palette))
rcParams['figure.figsize']= 12,6
import warnings
warnings.filterwarnings('ignore')
seed=42
np.random.seed(seed)
torch.manual_seed(seed)
# + [markdown] id="C4rPQLBqsVjp"
# ### Read the datasets
# + colab={"base_uri": "https://localhost:8080/", "height": 238} id="ZOXV2iSDsRuw" outputId="19d7a5bc-69f8-4fea-9a1d-41879857ae77"
# Train dataset
df_train=pd.read_csv('/content/train_preprocessed.csv')
print('There are {} observations (tweets) & {} features in the Train dataset'.format(df_train.shape[0],df_train.shape[1]))
print('Look at the Train dataset:')
df_train.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 238} id="v_i4zwj1shuI" outputId="d5bf9e86-d6ae-49a5-a4d5-201ad3972995"
# Validation dataset
df_dev=pd.read_csv('/content/dev_preprocessed.csv')
print('There are {} observations (tweets) & {} features in the Train dataset'.format(df_dev.shape[0],df_dev.shape[1]))
print('Look at the Validation (dev) dataset:')
df_dev.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 238} id="LQleyTDnsrGh" outputId="fb5aec76-7b74-4f48-c356-663477c558cf"
# Test dataset
df_test=pd.read_csv('/content/test_preprocessed.csv')
print('There are {} observations (tweets) & {} features in the Train dataset'.format(df_test.shape[0],df_test.shape[1]))
print('Look at the Test dataset:')
df_test.head()
# + [markdown] id="jCx7HRQ7s5lH"
# Let's check missing values if there are any.
# + colab={"base_uri": "https://localhost:8080/"} id="P2tC6dbKszCL" outputId="ec438bdc-9611-4c78-c2e0-f61efe75addd"
df_train.info()
# + colab={"base_uri": "https://localhost:8080/"} id="302kdQWPs8xw" outputId="4ebb082b-e7bf-4915-e61b-e622c27d67c1"
df_dev.info()
# + colab={"base_uri": "https://localhost:8080/"} id="w7q4bHBhtBRc" outputId="76dea874-2ecd-4742-fe13-5a02cb1bb593"
df_test.info()
# + [markdown] id="m-stRFa0tOsQ"
# We have not found any null values in train, dev, & test dataset.
# + [markdown] id="0RJoxEXZtSO4"
# ### Count plot for label
# + colab={"base_uri": "https://localhost:8080/", "height": 415} id="DnI34fB3tLaU" outputId="39fa51b5-a0e1-49fb-a1dd-7b1221192ee3"
sns.countplot(df_train.label)
plt.xlabel('label')
# + [markdown] id="1zVRuw13taWi"
# We need only 3 classes (false, true, & mixture) for this task. So drop unproven class from dataset.
# + colab={"base_uri": "https://localhost:8080/", "height": 432} id="8DJlRyULtXGQ" outputId="37d6390d-21d0-4890-aaa7-7b8a16cae120"
df_train=df_train[~(df_train.label=='unproven')]
print('Train dataset shape : {}'.format(df_train.shape))
sns.countplot(df_train.label)
plt.xlabel('label')
# + [markdown] id="tXirAWz7tkXZ"
# ### Encode out text labels to numeric.
# + colab={"base_uri": "https://localhost:8080/"} id="GwZ7rR5oteg6" outputId="83d17ed7-ca62-47bc-ddb5-6fa5bb752d82"
# Initiate LabelEncoder class
le =LabelEncoder()
# Training dataset
df_train['label'] = le.fit_transform(df_train['label'])
df_train.label.value_counts() # 0 -false, 2-true, 1-mixture
# + colab={"base_uri": "https://localhost:8080/"} id="Oj8vuTnktoC5" outputId="5d4454f5-09b8-446e-ced3-fc3a8aad6841"
# Dev dataset
df_dev=df_dev[~(df_dev.label=='unproven')] # only 3 classes
df_dev['label'] = le.transform(df_dev['label'])
df_dev.label.value_counts() # 0 -false, 2-true, 1-mixture
# + colab={"base_uri": "https://localhost:8080/"} id="3JN8GAnRtr-b" outputId="3e969710-4af4-4d34-e122-4be5de767ef8"
# Test dataset
df_test=df_test[~(df_test.label=='unproven')] # only 3 classes
df_test['label'] = le.transform(df_test['label'])
df_test.label.value_counts() # 0 -false, 2-true, 1-mixture
# + [markdown] id="avGKPdDSt0BC"
# ### Define pre-trained model
# + id="K5lhID51twVL"
Pre_trained_model='bert-base-cased'
custom_model = '/content/bert_model/' # fine tuned language model weights
# + [markdown] id="14nv26jQuAxy"
# ### Load the BERT tokenizer:
# + id="7jkE6OQet8_y"
tokenizer=BertTokenizer.from_pretrained(Pre_trained_model)
# + id="4dgwJuF5uO1c"
# Convert to list
train_explanations = df_train.explanation.tolist()
dev_explanations = df_dev.explanation.tolist()
test_explanations = df_test.explanation.tolist()
# + id="9Jvnk_nNukqm"
# Create token ids (input ids) for each explanation
# Train dataset
train_input_ids = [tokenizer.encode(train_explanations[i],add_special_tokens = True) for i in range(0,len(train_explanations))]
# dev dataset
dev_input_ids = [tokenizer.encode(dev_explanations[i],add_special_tokens = True) for i in range(0,len(dev_explanations))]
# Test dataset
test_input_ids = [tokenizer.encode(test_explanations[i],add_special_tokens = True) for i in range(0,len(test_explanations))]
# + [markdown] id="JRQkdrZput8j"
# ### Padding & Truncating
# + [markdown] id="9Hi1dbE3uzar"
# Pad and truncate our sequences so that they all have the same length, Maximum explanation length.
# + colab={"base_uri": "https://localhost:8080/"} id="-mFZFy3_uokl" outputId="c3134a67-50e3-4fb5-aa84-781253502d1e"
print('Max explanation length for train data: ', max([len(exp) for exp in train_input_ids]))
print('Max explanation length for dev/validation data: ', max([len(exp) for exp in dev_input_ids]))
print('Max explanation length for test data: ', max([len(exp) for exp in test_input_ids]))
# + colab={"base_uri": "https://localhost:8080/"} id="dubrQM4Eu5A_" outputId="b52bba5b-28ef-46b8-f079-b86071ef0ad6"
from keras.preprocessing.sequence import pad_sequences # Pad utility function to pad sequences to maximum length.
# Train dataset
Max_length = 320 # We consider maximum length of explanations more than 296 just to be on safer side.
print('\nPadding/truncating all sentences to %d values for train dataset...' % Max_length)
# Pad our input tokens with value 0.
# "post" indicates that we want to pad and truncate at the end of the sequence
train_input_ids = pad_sequences(train_input_ids, maxlen=Max_length, dtype="long",
value=0, truncating="post", padding="post")
# dev dataset
Max_length = 228 # We consider maximum length of explanations more than 250 just to be on safer side.
print('\nPadding/truncating all sentences to %d values for dev dataset...' % Max_length)
dev_input_ids = pad_sequences(dev_input_ids, maxlen=Max_length, dtype="long",
value=0, truncating="post", padding="post")
# Test dataset
Max_length = 171 # We consider maximum length of explanations more than 200 just to be on safer side.
print('\nPadding/truncating all sentences to %d values for test dataset' % Max_length)
test_input_ids = pad_sequences(test_input_ids, maxlen=Max_length, dtype="long",
value=0, truncating="post", padding="post")
print('\n All are done.')
# + [markdown] id="PJ82rUsPvArI"
# ### Attention Masks
# + id="zNltZBNnu8zF"
# Create attention masks
# Train dataset
train_attention_masks = [[int(token_id > 0) for token_id in exp]
for exp in train_input_ids]
# dev dataset
dev_attention_masks = [[int(token_id > 0) for token_id in exp]
for exp in dev_input_ids]
# Test dataset
test_attention_masks = [[int(token_id > 0) for token_id in exp]
for exp in test_input_ids]
# + id="XXaeiJlIvJ2t"
# Convert all inputs and labels into torch tensors, the required datatype
# for our model.
# input_ids
train_inputs = torch.tensor(train_input_ids)
dev_inputs = torch.tensor(dev_input_ids)
test_inputs = torch.tensor(test_input_ids)
# labels
train_labels = torch.tensor(df_train.label.values)
dev_labels = torch.tensor(df_dev.label.values)
test_labels = torch.tensor(df_test.label.values)
# attention masks
train_masks = torch.tensor(train_attention_masks)
dev_masks = torch.tensor(dev_attention_masks)
test_masks = torch.tensor(test_attention_masks)
# + id="Po0z_YCRvMoG"
# Create a DataLoader to load our datasets
batch_size = 8
# Create the DataLoader for our training set.
train_data = TensorDataset(train_inputs, train_masks, train_labels)
train_sampler = RandomSampler(train_data)
train_dataloader = DataLoader(train_data, sampler=train_sampler, batch_size=batch_size)
# Create the DataLoader for our dev set.
dev_data = TensorDataset(dev_inputs, dev_masks, dev_labels)
dev_sampler = SequentialSampler(dev_data)
dev_dataloader = DataLoader(dev_data, sampler=dev_sampler, batch_size=batch_size)
# Create the DataLoader for our test set.
test_data = TensorDataset(test_inputs, test_masks, test_labels)
test_sampler = SequentialSampler(test_data)
test_dataloader = DataLoader(test_data, sampler=test_sampler, batch_size=batch_size)
# + colab={"base_uri": "https://localhost:8080/"} id="ELNUggcYvWCP" outputId="5b16414b-b193-4f85-b7c9-3522122a2eab"
data1 = next(iter(test_dataloader))
data1[2]
# + [markdown] id="tU3loY8Tvdnh"
# ### Build a Fact Checking Classifier
# + [markdown] id="WhguyxnUvhKv"
# ### BertForSequenceClassification
# + id="hbpAXcPpvY7p"
# Number of classes / labels
n_classes = df_train['label'].nunique()
# + colab={"base_uri": "https://localhost:8080/"} id="gnD8lyDEvndI" outputId="624f8566-ac44-48c0-ffdc-0d8b351c17e2"
bert_model = BertForSequenceClassification.from_pretrained(custom_model,num_labels = n_classes,
output_attentions = False,output_hidden_states = False )
# + colab={"base_uri": "https://localhost:8080/"} id="A_NvnsBYvu9a" outputId="310a1f89-c0cf-4555-fd5d-5140bb10aefe"
# Put our model in training mode as it is in evaluation mode by default
bert_model.train()
# + id="bP3weQ_3xpxb"
## Move model to the GPU
bert_model = bert_model.to(device)
# + [markdown] id="h3Ssp6x3x74b"
# ### Optimizer & Learning Rate Scheduler
# + [markdown] id="eeJyeKcCyArH"
# As per original paper on BERT, we will use AdamW optimizer for correcting weight decay. We will also use, linear scheduler with no warm up steps:
# + id="11EVSLNOx2c5"
epochs=10
# Define optimizer for updating weights
optimizer=AdamW(bert_model.parameters(),lr=5e-5)
total_steps=len(train_dataloader)*epochs
# Scheduler for learning rate change
scheduler=get_linear_schedule_with_warmup(
optimizer,
num_warmup_steps=0,
num_training_steps=total_steps
)
# Define loss function & move it to GPU
loss_fn=nn.CrossEntropyLoss().to(device)
# + [markdown] id="n9pPv0GoyHFJ"
# ### Train our model
# + [markdown] id="e4dL8zwmyLcp"
# Define a function for calculating elapsed time.
# + id="IbRFVLVsyEJG"
def format_time(elapsed):
# Round to the nearest second
elapsed_round = int(round(elapsed))
# Format time in hh:mm:ss
return str(datetime.timedelta(seconds = elapsed_round))
# + [markdown] id="_J_yovx_yT0t"
# Define a function to calculate accuracy
# + id="3mdBRzOryQ-B"
def accuracy(preds, labels):
preds = np.argmax(preds, axis=1).flatten()
labels = labels.flatten()
return np.sum(preds == labels) / len(labels)
# + [markdown] id="rs7MZRxEyaoi"
# ### Start training...
# + colab={"base_uri": "https://localhost:8080/"} id="QaaTUpRJyXmK" outputId="31c9c52b-411a-49c8-a0c7-246e2c18918b"
# Store loss value for each epoch
loss_values = []
for epoch in range(0,epochs):
# Perform forward pass over the training dataset
print("\n ======== Epoch {:}/{:} ========".format(epoch+1,epochs))
print('Training....')
# Measure how long a epoch takes
t0 = time.time()
# Reset total loss for this epoch
total_loss = 0
# Put the model in training mode
bert_model.train()
# For each training batch
for step,batch in enumerate(train_dataloader):
# Update progress for 50 steps
if step % 50 ==0 & step !=0:
# Calculate elapsed time in minutes
elapsed = format_time(time.time(),t0)
# Report progress
print(' Batch {:>5,} of {:>5,}. Elapsed:{:}.'.format(step,len(train_loader),elapsed))
# Unpack training batch from trainloader & move to GPU
b_input_ids = batch[0].to(device) # 0 - input ids
b_attention_mask = batch[1].to(device) # 1 - input masks
b_labels = batch[2].to(device) # 2 - labels
# Clear default gradients accumulated in Pytorch
bert_model.zero_grad()
# Output the results
outputs = bert_model(input_ids = b_input_ids,attention_mask = b_attention_mask,labels=b_labels) # Return tuple
# Loss value from output
loss = outputs[0] # 0 - loss
# Update total loss
total_loss += loss.item()
# Perform a backward pass to calculate gradients
loss.backward()
# To avoid exploding vanishing gradients problem,clip the norm of the gradients to 1.0
torch.nn.utils.clip_grad_norm_(bert_model.parameters(),1.0)
# Update the parameters (weights)
optimizer.step()
# Update the learning rate
scheduler.step()
# Calculate the average loss over training data
avg_total_loss = total_loss/len(train_dataloader)
# Store the loss values
loss_values.append(avg_total_loss)
print('\n Average training loss : {0:.2f}'.format(avg_total_loss))
print('Training epoch took: {:}'.format(format_time(time.time()- t0)))
####### Validation #######
# After each epoch perform validation to check model performance
print('\n Running validation...')
t0 = time.time()
#put model in evaluation mode
bert_model.eval()
# Tracking variables
eval_loss,eval_acc = 0,0
nb_eval_steps,nb_eval_examples = 0,0
# Evaluate dev data for each epoch
for batch in dev_dataloader:
# Move batch to GPU
batch = tuple(t.to(device) for t in batch)
# Unpack inputs from dev dataloader
b_input_ids,b_attention_mask,b_labels = batch
# Tell model not to compute gradients to save memory & speed up validation
with torch.no_grad():
# Forward pass, calculate logit prediction
outputs = bert_model(input_ids=b_input_ids,attention_mask=b_attention_mask)
# logits are class probabilities & get them from outputs
logits = outputs[0]
# Move logits & labels to CPU
logits = logits.detach().cpu().numpy()
labels = b_labels.to('cpu').numpy()
# Calculate accuracy for this batch
eval_accuracy = accuracy(logits,labels)
# Accumulate total evaluation accuracy
eval_acc += eval_accuracy
# Track the number of steps
nb_eval_steps += 1
# Report the final validation accuracy
print(' Accuracy {0:.2f}'.format(eval_acc/nb_eval_steps))
print(' Validation took : {:}'.format(format_time(time.time() - t0)))
print('\n Training completed!')
# + [markdown] id="p56shnZY0hIe"
# Let's look at training vs validation loss:
# + colab={"base_uri": "https://localhost:8080/", "height": 415} id="uYzC9_rAyh-K" outputId="27f058d2-41f5-4c28-ba53-f4d4050d340a"
df = pd.DataFrame(loss_values,columns=['Loss'])
sns.lineplot(data=df,x=df.index,y=df.Loss)
plt.xlabel('Epoch')
plt.ylabel('Loss')
# + colab={"base_uri": "https://localhost:8080/"} id="J8FDQYQ00luR" outputId="08748420-a8f9-49cd-e835-a163f0d0b36a"
# Model performance on unseen data (test data)
print('\n Running testing on unseen data...')
#put model in evaluation model
bert_model.eval()
# Tracking variables
# Tracking variables
predictions , true_labels = [], []
# Evaluate test data
for batch in test_dataloader:
# Move batch to GPU
batch = tuple(t.to(device) for t in batch)
# Unpack inputs from test dataloader
b_input_ids,b_attention_mask,b_labels = batch
# Tell model not to compute gradients to save memory & speed up validation
with torch.no_grad():
# Forward pass, calculate logit prediction
outputs = bert_model(input_ids=b_input_ids,attention_mask=b_attention_mask)
# logits are class probabilities & get them from outputs
logits = outputs[0]
# Move logits & labels to CPU
logits = logits.detach().cpu().numpy()
labels = b_labels.to('cpu').numpy()
# Store predictions and true labels
predictions.append(logits)
true_labels.append(labels)
print('Done')
# + id="c74g62Cy0qQZ"
F1_score = []
# For each batch
for i in range(len(true_labels)):
preds_labels = np.argmax(predictions[i],axis=1).flatten()
score = f1_score(true_labels[i],preds_labels,pos_label= 0,average = 'weighted')
F1_score.append(score)
# + colab={"base_uri": "https://localhost:8080/"} id="glrAexQC0ye7" outputId="cf081205-2ffb-454c-f8b7-61fc00129c1b"
print('Average f1-score for test dataset : {0:0.2f}'.format(mean(F1_score)))
# + [markdown] id="YzC2nCFo05Z0"
# Checking model performance on random tweets.
# + id="82_V1zXI01iY"
twt_df = pd.read_csv('/content/tweets_100_cleaned.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="-WuP0fWm1BdN" outputId="0ffedc09-01ae-44c0-e664-27bfcbc476de"
twt_df.head()
# + id="Sr5K5Su61HSW"
# Convert to list
raw_explanations = twt_df.tweet.tolist()
# + id="7Bl0ilhe1Kfw"
# Create token ids (input ids) for each tweet
# raw dataset
raw_input_ids = [tokenizer.encode(raw_explanations[i],add_special_tokens = True) for i in range(0,len(raw_explanations))]
# + colab={"base_uri": "https://localhost:8080/"} id="tefoou8G1NCF" outputId="2e476fc1-5ffd-485a-dbe8-d6d24f39c1e1"
print('Max explanation length for raw data: ', max([len(exp) for exp in raw_input_ids]))
# + colab={"base_uri": "https://localhost:8080/"} id="hZwltLGD1Q8O" outputId="91dbe136-e824-4b4a-edb7-6c3dee6cfcab"
# raw dataset
Max_length = 120 # We consider maximum length of explanations more than 120 just to be on safer side.
print('\nPadding/truncating all sentences to %d values for raw dataset...' % Max_length)
# Pad our input tokens with value 0.
# "post" indicates that we want to pad and truncate at the end of the sequence
raw_input_ids = pad_sequences(raw_input_ids, maxlen=Max_length, dtype="long",
value=0, truncating="post", padding="post")
# + id="WFnRB5Ry1Tnb"
# Create attention masks
# raw dataset
raw_attention_masks = [[int(token_id > 0) for token_id in exp]
for exp in raw_input_ids]
# + id="kB2j03Xv1W1h"
def apply(label):
if label ==False:
return 0
else:
return 2
# + id="IYmm2YG91aK9"
twt_df.label = twt_df.label.apply(lambda x:apply(x))
# + id="1QYEpaF51c2f"
# Convert all inputs and labels into torch tensors, the required datatype
# for our model.
# input_ids
raw_inputs = torch.tensor(raw_input_ids)
# labels
raw_labels = torch.tensor(twt_df.label.values)
# attention masks
raw_masks = torch.tensor(raw_attention_masks)
# + id="FyV9tYdq1fTB"
# Create a DataLoader to load our datasets
batch_size = 8
# Create the DataLoader for our raw set.
raw_data = TensorDataset(raw_inputs, raw_masks, raw_labels)
raw_sampler = RandomSampler(raw_data)
raw_dataloader = DataLoader(raw_data, sampler=raw_sampler, batch_size=batch_size)
# + colab={"base_uri": "https://localhost:8080/"} id="Vkz1uNmU1iA6" outputId="d2396a3f-e03d-4897-aabc-e559e498b694"
# Model performance on random tweets data (raw data)
print('\n Running prediction on raw data...')
#put model in evaluation model
bert_model.eval()
# Tracking variables
# Tracking variables
preds , true_labels = [], []
# Evaluate test data
for batch in raw_dataloader:
# Move batch to GPU
batch = tuple(t.to(device) for t in batch)
# Unpack inputs from test dataloader
b_input_ids,b_attention_mask,b_labels = batch
# Tell model not to compute gradients to save memory & speed up validation
with torch.no_grad():
# Forward pass, calculate logit prediction
outputs = bert_model(input_ids=b_input_ids,attention_mask=b_attention_mask)
# logits are class probabilities & get them from outputs
logits = outputs[0]
# Move logits & labels to CPU
logits = logits.detach().cpu().numpy()
labels = b_labels.to('cpu').numpy()
# Store predictions and true labels
preds.append(logits)
true_labels.append(labels)
print('Done')
# + id="fRdpJcg-1od-"
f1 = []
# For each batch
for i in range(len(true_labels)):
preds_labels = np.argmax(preds[i],axis=1).flatten()
score = f1_score(true_labels[i],preds_labels,pos_label=0,average = 'weighted')
f1.append(score)
# + colab={"base_uri": "https://localhost:8080/"} id="Haayhr1L1t3u" outputId="42c953d7-ab70-43ad-bd19-d05798dd5c48"
print('Average f1-score for raw data: {0:0.2f}'.format(mean(f1)))
# + [markdown] id="eZIebBTH10ne"
# Check an example batch.
# + colab={"base_uri": "https://localhost:8080/"} id="sueGmrjG1xid" outputId="2647e183-a3ae-4cb7-8558-7cf5c2b110e6"
print(f'True labels :{true_labels[0]}')
print(f'\n Predicted labels : {np.argmax(preds[0],axis=1).flatten()}')
# + id="9bOIAKXP144W"
| Fine_tuning_language_model_for_fact_checking_task.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import tensorflow as tf
dataset = pd.read_csv('../input/digit-recognizer/train.csv')
print(dataset.shape)
# Pixel0 ~ Pixel783までの計784カラム, sqrt(784) = 28 * 28 の画像データ
label_counts = dataset['label'].value_counts()
print(f'Label counts:\n {label_counts}')
# +
fig, ax = plt.subplots(4,5)
fig.set_figheight(5)
num_read_img = 20
for image_index in range(0,num_read_img):
pixels = dataset.iloc[image_index,1:].values.reshape(28,28)
draw_axis= ax[int(image_index/5),int(image_index%5)]
draw_axis.imshow(pixels, cmap='gray')
draw_axis.set_title(dataset.iloc[image_index,0])
draw_axis.axes.xaxis.set_visible(False)
draw_axis.axes.yaxis.set_visible(False)
plt.tight_layout()
plt.show()
# +
label = dataset.loc[:,'label']
#label = pd.get_dummies(label, drop_first=False)
images = dataset.iloc[:,1:].values
# https://keras.io/ja/layers/convolutional/
# 入力: 'data_format='channels_first'の場合, (batch_size, channels, rows, cols)の4階テンソル'
# 'data_format='channels_last'の場合, (batch_size, rows, cols, channels)の4階テンソルになります.
# Conv2D層への入力のため、4次元にreshpae。 batsh_sizeはtrain_test_splitで変わるのですべてを意味する"-1"
images = images.reshape(-1,28,28,1)/255
# labelは名義尺度のため、そのままでは数値尺度として扱われてしまう?
# ということでOnehot-encoding
label = pd.get_dummies(label, columns=['label'], drop_first=False)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(images,label, test_size=0.2)
print(X_train.shape)
# +
# CNNの初期化。単純なSequentialモデルで作成。
cnn = tf.keras.models.Sequential()
# 畳み込み https://keras.io/ja/layers/convolutional/
# 128x128 RGB画像ではinput_shape=(128, 128, 3)となります.
# 最初のレイヤーだけは、入り口となる入力シェイプが必要
cnn.add(tf.keras.layers.Conv2D(filters=32, kernel_size=3, padding="same", activation="relu", input_shape=[28, 28, 1], data_format='channels_last'))
# tf.keras.layers.BatchNormalization https://www.tensorflow.org/api_docs/python/tf/keras/layers/BatchNormalization
# Batch Normalization:ニューラルネットワークの学習を加速させる汎用的で強力な手法 - DeepAge https://deepage.net/deep_learning/2016/10/26/batch_normalization.html
cnn.add(tf.keras.layers.BatchNormalization())
# Poolingにより、ダウンサンプリング
cnn.add(tf.keras.layers.MaxPool2D(pool_size=2, strides=2, padding='valid'))
# 画像サイズはPoolingにより14*14に
cnn.add(tf.keras.layers.Dropout(0.4))
# 2層目の中間層を追加
cnn.add(tf.keras.layers.Conv2D(filters=64, kernel_size=3, padding="same", activation="relu"))
cnn.add(tf.keras.layers.BatchNormalization())
cnn.add(tf.keras.layers.MaxPool2D(pool_size=2, strides=2, padding='valid'))
# 画像サイズはPoolingにより7*7に
cnn.add(tf.keras.layers.Dropout(0.4))
# 3層目の中間層を追加
#cnn.add(tf.keras.layers.Conv2D(filters=32, kernel_size=3, padding="same", activation="relu"))
#cnn.add(tf.keras.layers.MaxPool2D(pool_size=2, strides=2, padding='same'))
# 画像サイズは'padding=same'のため、Poolingにより4*4に
#cnn.add(tf.keras.layers.Dropout(0.25))
# Flattening
cnn.add(tf.keras.layers.Flatten())
# 接続
# unitsは レイヤーの出力形状
cnn.add(tf.keras.layers.Dense(units=128, activation='relu'))
cnn.add(tf.keras.layers.Dropout(0.4))
# 出力層
#cnn.add(tf.keras.layers.Dense(units=10, activation='sigmoid'))
cnn.add(tf.keras.layers.Dense(10, activation='softmax'))
# Compiling the CNN
# 評価関数の選定 -> https://keras.io/ja/metrics/
# The difference between sparse_categorical_crossentropy and categorical_crossentropy is whether your targets are one-hot encoded.
cnn.compile(optimizer = 'adam', loss = tf.keras.losses.categorical_crossentropy, metrics = ['accuracy'])
# -
answer = cnn.fit(images, label, epochs = 50, batch_size = 20)
score = cnn.evaluate(X_test, y_test)
test_dataset = pd.read_csv('../input/digit-recognizer/test.csv')
y_pred = cnn.predict(test_dataset.iloc[:,:].values.reshape(-1,28,28,1)/255)
imageId = np.arange(1,28001)
y_pred_selected = np.argmax(y_pred,axis=1)
print(y_pred_selected)
result_table = pd.DataFrame({ 'ImageId': imageId,
'Label': y_pred_selected })
result_table.to_csv('prediction.csv', index=False)
| digit-recognizer/EDA-digit-recognizer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Day 14: One-Time Pad
#
# author: <NAME>
#
# license: [MIT](https://opensource.org/licenses/MIT)
#
# [link to problem statement](http://adventofcode.com/2016/day/14)
# In order to communicate securely with Santa while you're on this mission, you've been using a [one-time pad](https://en.wikipedia.org/wiki/One-time_pad) that you [generate](https://en.wikipedia.org/wiki/Security_through_obscurity) using a pre-agreed algorithm. Unfortunately, you've run out of keys in your one-time pad, and so you need to generate some more.
#
# To generate keys, you first get a stream of random data by taking the [MD5](https://en.wikipedia.org/wiki/MD5) of a pre-arranged [salt](https://en.wikipedia.org/wiki/Salt_(cryptography) (your puzzle input) and an increasing integer index (starting with `0`, and represented in decimal); the resulting MD5 hash should be represented as a string of lowercase hexadecimal digits.
#
# However, not all of these `MD5` hashes are keys, and you need `64` new keys for your one-time pad. A hash is a key only if:
#
# - It contains three of the same character in a row, like `777`. Only consider the first such triplet in a hash.
# - One of the next `1000` hashes in the stream contains that same character five times in a row, like `77777`.
#
# Considering future hashes for five-of-a-kind sequences does not cause those hashes to be skipped; instead, regardless of whether the current hash is a key, always resume testing for keys starting with the very next hash.
#
# For example, if the pre-arranged salt is `abc`:
#
# - The first index which produces a triple is `18`, because the MD5 hash of `abc18` contains `...cc38887a5....` However, index `18` does not count as a key for your one-time pad, because none of the next thousand hashes (index `19` through index `1018`) contain `88888`.
# - The next index which produces a triple is `39`; the hash of `abc39` contains `eee`. It is also the first key: one of the next thousand hashes (the one at index `816`) contains `eeeee`.
# - None of the next six triples are keys, but the one after that, at index `92`, is: it contains `999` and index `200` contains `99999`.
# - Eventually, index `22728` meets all of the criteria to generate the 64th key.
#
# So, using our example salt of abc, index `22728` produces the `64th` key.
#
# Given the actual salt in your puzzle input, what index produces your 64th one-time pad key?
# ### Solution logic
#
# Our salt is out input, we apply it increasingly to integers, take their MD5 hash, and if it contains a character repeating three times, we check if there is a hash in the next 1000 integers that features the same character 7 times, and if it does, then the integer we had is our key. Find 64 such keys, with the index of the 64th key being the answer. Seems simple and straightforward.
import re
three_repeating_characters = re.compile(r'(.)\1{2}')
with open('../inputs/day14.txt', 'r') as f:
salt = f.readline().strip()
# TEST DATA
# salt = 'abc'
print(salt)
# **Hash index**
#
# To prevent the same hash from being counted upon again and again, we maintain a hash index to store the hashes of indexes. We also trim the index to remove any index for keys lower than the current key since we do not require those.
# +
import hashlib
hash_index= {}
def get_hash_string(key):
if key in hash_index:
return hash_index[key]
string = '{salt}{key}'.format(salt=salt, key=key)
md5 = hashlib.md5()
md5.update(string.encode('ascii'))
hashstring = md5.hexdigest()
hash_index[key] = hashstring
return hashstring
def run():
keys = []
current_key = 0
while(len(keys) < 64):
for i in range(0, current_key):
hash_index.pop(i, None)
hashstring = get_hash_string(current_key)
repeating_chacter = three_repeating_characters.findall(hashstring)
if not repeating_chacter:
current_key += 1
continue
repeating_chacter = repeating_chacter[0]
repeating_character_five = ''.join(repeating_chacter for i in range(0, 5))
for qualifying_index in range(current_key + 1, current_key + 1001):
hashstring = get_hash_string(qualifying_index)
if repeating_character_five in hashstring:
break
else:
current_key += 1
continue
keys.append(current_key)
print(len(keys), current_key)
current_key += 1
return keys
# -
print('answer', run()[63])
# ## Part Two
#
# Of course, in order to make this process [even more secure](https://en.wikipedia.org/wiki/MD5#Security), you've also implemented [key stretching](https://en.wikipedia.org/wiki/Key_stretching).
#
# Key stretching forces attackers to spend more time generating hashes. Unfortunately, it forces everyone else to spend more time, too.
#
# To implement key stretching, whenever you generate a hash, before you use it, you first find the MD5 hash of that hash, then the MD5 hash of that hash, and so on, a total of 2016 additional hashings. Always use lowercase hexadecimal representations of hashes.
#
# For example, to find the stretched hash for index 0 and salt abc:
#
# - Find the MD5 hash of a`bc0`: `577571be4de9dcce85a041ba0410f29f`.
# - Then, find the MD5 hash of that hash: `eec80a0c92dc8a0777c619d9bb51e910`.
# - Then, find the MD5 hash of that hash: `16062ce768787384c81fe17a7a60c7e3`.
# - `...repeat many times...`
# - Then, find the MD5 hash of that hash: `a107ff634856bb300138cac6568c0f24`.
#
# So, the stretched hash for index `0` in this situation is `a107ff....` In the end, you find the original hash (one use of MD5), then find the hash-of-the-previous-hash `2016` times, for a total of `2017` uses of MD5.
#
# The rest of the process remains the same, but now the keys are entirely different. Again for salt abc:
#
# - The first triple (`222`, at index `5`) has no matching `22222` in the next thousand hashes.
# - The second triple (`eee`, at index `10`) hash a matching `eeeee` at index `89`, and so it is the first key.
# - Eventually, index `22551` produces the `64th` key (triple fff with matching fffff at index `22859`.
#
# Given the actual salt in your puzzle input and using 2016 extra MD5 calls of key stretching, what index now produces your 64th one-time pad key?
# ### Solution logic
#
# We only need to change the definition of `get_hash_string` to calculate the hash 2016 times more. And then simply run the algorithm again.
#
# To prevent computationally intensive operations from repeating themselves, we maintain an index of hashes so that we can easily lookup indexes without needing to calculate their hashes.
# +
hash_index = {}
def get_hash_string(key):
if key in hash_index:
return hash_index[key]
string = '{salt}{key}'.format(salt=salt, key=key)
md5 = hashlib.md5()
md5.update(string.encode('ascii'))
hashstring = md5.hexdigest()
# PART TWO
for i in range(0, 2016):
md5 = hashlib.md5()
md5.update(hashstring.encode('ascii'))
hashstring = md5.hexdigest()
hash_index[key] = hashstring
return hashstring
# -
print('answer', run()[63])
# == END ==
| 2016/python3/Day14.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
import theano.tensor as tt
import pymc3 as pm
from scipy import stats
from pymc3 import Continuous
from theano import tensor
#exp, log, sqrt
import seaborn as sns
import matplotlib.pyplot as plt
sns.set_context('notebook')
plt.style.use('seaborn-darkgrid')
print('Running on PyMC3 v{}'.format(pm.__version__))
core_count = 12
# -
def SuperNova_CurveRise(t, A, B, t0, trise):
return ((A+B*(t-t0))/(1+np.exp(-(t-t0)/trise)))
def SuperNova_CurveFall(t, A, B, t0, gamma, trise, tfall):
return (A + B*((gamma+t0)-t0))*np.exp(-(t-(gamma+t0))/tfall)/(1+np.exp(-(t-t0)/trise))
#These are the values we are going to use for a test
np.random.seed(212)
trise_A = 2
tfall_A = 20
Amplitude_A = 1500
Beta_A = -5
t0_A = 43.2
gamma_A = 60
end_A = 500
size = 10 #data set size double for total observations
sigma_A = 100 #Telescope error
time_axis_rise = np.random.uniform(low=0, high=(gamma_A+t0_A), size=size)
time_axis_rise = np.sort(time_axis_rise)
time_axis_fall = np.random.uniform(low=(gamma_A+t0_A), high=end_A, size=size)
time_axis_fall = np.sort(time_axis_fall)
Y_actual_rise = SuperNova_CurveRise(time_axis_rise, Amplitude_A, Beta_A, t0_A, trise_A)
Y_actual_fall = SuperNova_CurveFall(time_axis_fall, Amplitude_A, Beta_A, t0_A, gamma_A, trise_A, tfall_A)
time_axis = np.append(time_axis_rise, time_axis_fall)
Y_actual = np.append(Y_actual_rise, Y_actual_fall)
Y_observed = Y_actual + np.random.normal(0,sigma_A, len(Y_actual))
#Y_observed = Y_actual
# +
fig, axes = plt.subplots(1, 1, sharex=True, figsize=(10,4))
axes.scatter(time_axis, Y_actual)
big_time_axis_rise = np.linspace(0,(gamma_A+t0_A),num = 100)
axes.plot(big_time_axis_rise, SuperNova_CurveRise(big_time_axis_rise, Amplitude_A, Beta_A, t0_A, trise_A))
big_time_axis_fall = np.linspace((gamma_A+t0_A),end_A,num = 100)
axes.plot(big_time_axis_fall, SuperNova_CurveFall(big_time_axis_fall, Amplitude_A, Beta_A, t0_A, gamma_A, trise_A, tfall_A))
axes.set_ylabel('Y')
axes.set_xlabel('X1')
plt.title("Actual Light Curve")
# +
fig, axes = plt.subplots(1, 1, sharex=True, figsize=(10,4))
axes.scatter(time_axis, Y_observed)
big_time_axis_rise = np.linspace(0,(gamma_A+t0_A),num = 100)
axes.plot(big_time_axis_rise, SuperNova_CurveRise(big_time_axis_rise, Amplitude_A, Beta_A, t0_A, trise_A))
big_time_axis_fall = np.linspace((gamma_A+t0_A),end_A,num = 100)
axes.plot(big_time_axis_fall, SuperNova_CurveFall(big_time_axis_fall, Amplitude_A, Beta_A, t0_A, gamma_A, trise_A, tfall_A))
axes.set_ylabel('Y')
axes.set_xlabel('X1')
plt.title("Observed Light Curve")
# -
step = int(np.ceil(size/2))
if step<1:
step = 1
slopes = []
least_slope = (0, 0)
for i in range(len(time_axis)-step):
if step > 1:
slope, intercept, r_value, p_value, std_err = stats.linregress(time_axis[i:i+step],Y_observed[i:i+step])
else:
slope = (Y_observed[i]-Y_observed[i+step])/(time_axis[i]-time_axis[i+step])
slopes.append(slope)
if(slope < least_slope[1]):
least_slope = (time_axis[i+int(np.floor(step/2))], slope)
print(least_slope)
plt.scatter(time_axis[0:len(time_axis)-step], slopes)
basic_model = pm.Model()
# +
with basic_model:
# Priors for unknown model parameters
trise = pm.Uniform('trise', lower = 0.01, upper = 50)
tfall = pm.Uniform('tfall', lower = 1, upper = 300)
Amp_Guess = np.max(Y_observed)-np.min(Y_observed)
Amplitude = pm.Normal('Amplitude', mu=Amp_Guess, sigma=Amp_Guess/2)
Beta = pm.Uniform('Beta', lower = -np.max(Y_observed)/150, upper = 0)
t0 = pm.Uniform('t0', lower = np.min(time_axis), upper = np.max(time_axis))
sigma = pm.HalfNormal('sigma', sigma=sigma_A)
#gamma = pm.Uniform('gamma', lower = np.min(time_axis), upper = np.max(time_axis), testval = (least_slope[0]-))
no_p = pm.Normal.dist(mu = 5, sigma = 5)
yes_p = pm.Normal.dist(mu = 60, sigma = 30)
gamma = pm.Mixture("gamma", w=[2/3,1/3], comp_dists = [no_p, yes_p])
#gamma = pm.math.sum(pm.Normal("no_p", mu = 5, sigma = 5),pm.Normal("yes_p", mu = 60, sigma = 30))
# Expected value of outcome
mu_rise = SuperNova_CurveRise(time_axis, Amplitude, Beta, t0, trise)
mu_fall = SuperNova_CurveFall(time_axis, Amplitude, Beta, t0, gamma, trise, tfall)
mu_switch = pm.math.switch(gamma+t0 >= time_axis, mu_rise, mu_fall)
# Likelihood (sampling distribution) of observations
Y_obs = pm.Normal('Y_obs', mu=mu_switch, sigma=sigma, observed=Y_observed)
# +
map_estimate = pm.find_MAP(model=basic_model)
map_estimate
# -
with basic_model:
# 1000 posterior samples
trace = pm.sample(10000, cores = 6)
# %matplotlib inline
pm.traceplot(trace)
summary_table = pm.summary(trace)
summary_table
trise_R = summary_table["mean"]["trise"]
tfall_R = summary_table["mean"]["tfall"]
Amplitude_R = summary_table["mean"]["Amplitude"]
Beta_R = summary_table["mean"]["Beta"]
t0_R = summary_table["mean"]["t0"]
gamma_R = summary_table["mean"]["gamma"]
print(trise_R, trise_A)
print(tfall_R, tfall_A)
print(Amplitude_R, Amplitude_A)
print(Beta_R, Beta_A)
print(t0_R, t0_A)
print(gamma_R, gamma_A)
# +
# %matplotlib inline
fig, ax = plt.subplots(figsize=(10,4))
big_time_axis_rise = np.linspace(0,t0_R+gamma_R,num = 100)
ax.plot(big_time_axis_rise, SuperNova_CurveRise(big_time_axis_rise, Amplitude_R, Beta_R, t0_R, trise_R))
big_time_axis_fall = np.linspace(t0_R+gamma_R,end_A,num = 100)
ax.plot(big_time_axis_fall, SuperNova_CurveFall(big_time_axis_fall, Amplitude_R, Beta_R, t0_R, gamma_R, trise_R, tfall_R))
ax.errorbar(time_axis, Y_observed, sigma_A, fmt='o')
ax.set_xlabel('x')
ax.set_ylabel('y_observed')
# +
# %matplotlib inline
fig, ax = plt.subplots(figsize=(10,4))
big_time_axis_rise = np.linspace(0,t0_R+gamma_R,num = 100)
ax.plot(big_time_axis_rise, SuperNova_CurveRise(big_time_axis_rise, Amplitude_R, Beta_R, t0_R, trise_R))
big_time_axis_fall = np.linspace(t0_R+gamma_R,end_A,num = 100)
ax.plot(big_time_axis_fall, SuperNova_CurveFall(big_time_axis_fall, Amplitude_R, Beta_R, t0_R, gamma_R, trise_R, tfall_R))
ax.scatter(time_axis, Y_actual)
ax.set_xlabel('x')
ax.set_ylabel('y_actual')
# -
| VillarCurves/Gamma_and_Bimodal_Priors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="incomplete-estimate"
import tensorflow as tf
import math
import time
import numpy as np
# + id="Oe6aDoIVoe_S"
from IPython.display import clear_output
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/"} id="5JGHxhN11ArQ" outputId="b981cc4d-b65a-4dee-f25d-e357b10e98a2"
from google.colab import drive
drive.mount('/content/drive')
# + id="GhoR8pEMuYZX"
# !cp /content/drive/MyDrive/Arcface/backbone_0.py .
# !cp /content/drive/MyDrive/Arcface/hypar.py .
# !cp /content/drive/MyDrive/Arcface/network_16.py .
import hypar
import backbone_0 as nn
import network_16 as net
# + id="damaged-exploration"
physical_devices = tf.config.experimental.list_physical_devices('GPU')
assert len(physical_devices) > 0, "Not enough GPU hardware devices available"
config = tf.config.experimental.set_memory_growth(physical_devices[0], True)
# + colab={"base_uri": "https://localhost:8080/"} id="H89tHgtiE8X3" outputId="f61936cb-96f0-47a4-9d5f-3749a23013eb"
physical_devices
# + id="assured-short"
X = np.load('/content/drive/MyDrive/Arcface/x_train.npy', allow_pickle=True)
Y = np.load('/content/drive/MyDrive/Arcface/y_train.npy', allow_pickle=True)
# + id="configured-period" colab={"base_uri": "https://localhost:8080/"} outputId="60ba17c9-bbba-402b-92a7-610e624b7589"
X = np.array(X, dtype='float32')
Y = np.array(Y, dtype='int32')
Y = np.reshape(Y, Y.shape[0])
print("X shape:",X.shape,"Y shape:",Y.shape)
X = net.Resnet_preprocess(X)
images = X
labels = Y
# Prepare the training dataset.
train_dataset = tf.data.Dataset.from_tensor_slices((images, labels))
train_dataset = train_dataset.shuffle(buffer_size=1024).batch(hypar.batch_size)
print("Training dataset ready!")
# + id="bigger-century"
num_classes = hypar.num_classes #number of people in the dataset
#feature vector dimension = (512) [comes from the resnet model]
class Arcface_Layer(tf.keras.layers.Layer): # Arcface layer definition
def __init__(self, num_outputs = num_classes, s=64., m=0.5): # s is scale factor, m is the margin to be added to the angle 'theta'
self.output_dim = num_outputs
self.s = s
self.m = m
super(Arcface_Layer, self).__init__()
def build(self, input_shape):
self.kernel = self.add_weight(name='weight',
shape=(input_shape[-1],self.output_dim),
initializer='glorot_uniform',
regularizer=tf.keras.regularizers.l2(l=5e-4),
trainable=True)
super(Arcface_Layer, self).build(input_shape)
def call(self, embedding, labels):
cos_m = math.cos(self.m)
sin_m = math.sin(self.m)
mm = sin_m * self.m # issue 1
threshold = math.cos(math.pi - self.m)
# inputs and weights norm
embedding_norm = tf.norm(embedding, axis=1, keepdims=True)
embedding = embedding / embedding_norm
weights_norm = tf.norm(self.kernel, axis=0, keepdims=True)
weights = self.kernel / weights_norm
# cos(theta+m)
cos_t = tf.matmul(embedding, weights, name='cos_t')
cos_t2 = tf.square(cos_t, name='cos_2')
sin_t2 = tf.subtract(1., cos_t2, name='sin_2')
sin_t = tf.sqrt(sin_t2, name='sin_t')
cos_mt = self.s * tf.subtract(tf.multiply(cos_t, cos_m),
tf.multiply(sin_t, sin_m), name='cos_mt')
# this condition controls the theta+m should in range [0, pi]
# 0<=theta+m<=pi
# -m<=theta<=pi-m
cond_v = cos_t - threshold
cond = tf.cast(tf.nn.relu(cond_v, name='if_else'), dtype=tf.bool)
keep_val = self.s * (cos_t - mm)
cos_mt_temp = tf.where(cond, cos_mt, keep_val)
mask = tf.one_hot(labels, depth=self.output_dim, name='one_hot_mask')
# mask = tf.squeeze(mask, 1)
inv_mask = tf.subtract(1., mask)
s_cos_t = tf.multiply(self.s, cos_t, name='scalar_cos_t')
output = tf.add(tf.multiply(s_cos_t, inv_mask), tf.multiply(cos_mt_temp, mask), name='arcface_loss_output')
return output
# + id="portable-shade" colab={"base_uri": "https://localhost:8080/"} outputId="e05d9dd1-6994-480d-971f-275cad198d29"
class train_model(tf.keras.Model):
def __init__(self):
super(train_model, self).__init__()
#self.resnet = net.Resnet_nn()
self.resnet = net.Resnet()
self.arcface = Arcface_Layer()
def call(self, x, y):
x = self.resnet(x)
return self.arcface(x, y)
# Instantiate a loss function.
def loss_fxn(logits,labels):
loss_fn = tf.reduce_mean(tf.nn.sparse_softmax_cross_entropy_with_logits(logits=logits, labels=labels))
return loss_fn
# Instantiate an optimizer to train the model.
learning_rate = 0.0005
optimizer = tf.keras.optimizers.SGD(learning_rate=learning_rate, momentum=0.9, nesterov=False)
#optimizer = tf.keras.optimizers.Adam(learning_rate=0.00001, beta_1=0.9, beta_2=0.999, epsilon=1e-07, amsgrad=False,name='Adam')
model = train_model()
@tf.function
def train_step(images, labels):
with tf.GradientTape() as tape:
logits = model(images,labels)
pred = tf.nn.softmax(logits)
#inf_loss = loss_fxn(pred,labels)
inf_loss = loss_fxn(logits,labels)
loss = inf_loss
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
train_loss = tf.reduce_mean(loss)
accuracy = tf.reduce_mean(tf.cast(tf.equal(tf.argmax(pred, axis=1, output_type=tf.dtypes.int32), tf.cast(labels,dtype = tf.int32)), dtype=tf.float32))
inference_loss = tf.reduce_mean(inf_loss)
regularization_loss = 0
return accuracy, train_loss, inference_loss, regularization_loss
# + id="aHdyzzFC_rq4" colab={"base_uri": "https://localhost:8080/"} outputId="7fae1658-59f1-4fd9-afc3-0e11fbedbf29"
epochs = 20
reg_coef = 1.0
file_name = '/content/drive/MyDrive/Arcface/checkpoint/try4_16/model_weights_'
file_number = 10
file_name = file_name + str(file_number*epochs)+ '_epochs'
model = tf.keras.models.load_model(file_name)
# + colab={"base_uri": "https://localhost:8080/"} id="K_UtYDB612eO" outputId="28016cb5-5601-4482-e0f5-87fd7c785c18"
print(file_name)
# + id="angry-currency" colab={"base_uri": "https://localhost:8080/"} outputId="8ba8ee20-ca87-4718-daec-ce5f2a2697df"
random_out = model(X[0:3,:,:,:],Y[0:3])
loss_log = []
epochs = 50
reg_coef = 1.0
file_number = 5
for save_wt in range (0, 5):
for epoch in range(epochs):
print("\nStart of epoch %d" % (epoch,))
start_time = time.time()
# Iterate over the batches of the dataset.
for step, (x_batch_train, y_batch_train) in enumerate(train_dataset):
accuracy, train_loss, inference_loss, regularization_loss = train_step(x_batch_train, y_batch_train)
if step % 20 == 0:
loss_log.append(train_loss)
print("Training loss (for one batch) at step %d: %.4f"% (step, float(train_loss)))
file_number += 1
file_name = '/content/drive/MyDrive/Arcface/checkpoint/try5_SGD_16/model_weights_'
file_name = file_name + str(file_number*epochs)+ '_epochs'
model.save(file_name)
# + id="dkn-YrGqdcJt"
loss_log_data = np.array(loss_log)
np.save('/content/drive/MyDrive/Arcface/checkpoint/try5_SGD_16/loss_log_1.npy', loss_log_data)
# + id="V0C7irVSe2Wc" colab={"base_uri": "https://localhost:8080/"} outputId="994f4cd2-3b01-4ae1-9d8b-8bcf53fdc165"
test_model = model.resnet
result = test_model.predict(X)
result.shape
# + id="1-DPkdau5L3q"
index = []
for i in range (0,1470):
if Y[i] == 25:
index.append(i)
# + id="L4mJP43S5kg8"
a = np.dot(result[877,:], np.array(result[1308,:]).T)
print(a)
# + id="o9hXk2dAPN79"
from sklearn.preprocessing import normalize
results = normalize(result, axis = 0)
results.shape
# + id="OXbmaykCSnPV"
model.arcface.get_weights()
# + id="sWhKJMGvFVlg"
| Arcface_Training_Code_v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# Fitting in Chebyshev basis
# ==========================
#
# Plot noisy data and their polynomial fit in a Chebyshev basis
#
#
#
# +
import numpy as np
import matplotlib.pyplot as plt
np.random.seed(0)
x = np.linspace(-1, 1, 2000)
y = np.cos(x) + 0.3*np.random.rand(2000)
p = np.polynomial.Chebyshev.fit(x, y, 90)
plt.plot(x, y, 'r.')
plt.plot(x, p(x), 'k-', lw=3)
plt.show()
| _downloads/plot_chebyfit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#initial code from here https://www.datacamp.com/community/tutorials/networkx-python-graph-tutorial
import itertools
import copy
import networkx as nx
import pandas as pd
import matplotlib.pyplot as plt
# Grab edge list data hosted on Gist
edgelist = pd.read_csv('https://gist.githubusercontent.com/brooksandrew/e570c38bcc72a8d102422f2af836513b/raw/89c76b2563dbc0e88384719a35cba0dfc04cd522/edgelist_sleeping_giant.csv')
# -
# Preview edgelist
edgelist.head(10)
nodelist = pd.read_csv('https://gist.githubusercontent.com/brooksandrew/f989e10af17fb4c85b11409fea47895b/raw/a3a8da0fa5b094f1ca9d82e1642b384889ae16e8/nodelist_sleeping_giant.csv')
nodelist.head(5)
# Create empty graph
g = nx.Graph()
# Add edges and edge attributes
for i, elrow in edgelist.iterrows():
g.add_edge(elrow[0], elrow[1], attr_dict=elrow[2:].to_dict())
# Edge list example
print(elrow[0]) # node1
print(elrow[1]) # node2
print(elrow[2:].to_dict()) # edge attribute dict
# Add node attributes
for i, nlrow in nodelist.iterrows():
g.node[nlrow['id']].update(nlrow[1:].to_dict())
# Node list example
print(nlrow)
# ## Inspect Graph
# Edges
# Your graph edges are represented by a list of tuples of length 3. The first two elements are the node names linked by the edge. The third is the dictionary of edge attributes.
# Preview first 5 edges
list(g.edges(data=True))[0:5]
# ## Nodes
# Similarly, your nodes are represented by a list of tuples of length 2. The first element is the node ID, followed by the dictionary of node attributes.
#
#
# Preview first 10 nodes
list(g.nodes(data=True))[0:10]
## Summary Stats
print('# of edges: {}'.format(g.number_of_edges()))
print('# of nodes: {}'.format(g.number_of_nodes()))
# ## Visualize
# Manipulate Colors and Layout
# Positions: First you need to manipulate the node positions from the graph into a dictionary. This will allow you to recreate the graph using the same layout as the actual trail map. Y is negated to transform the Y-axis origin from the topleft to the bottomleft.
# +
# Define node positions data structure (dict) for plotting
node_positions = {node[0]: (node[1]['X'], -node[1]['Y']) for node in g.nodes(data=True)}
# Preview of node_positions with a bit of hack (there is no head/slice method for dictionaries).
dict(list(node_positions.items())[0:5])
# -
# Colors: Now you manipulate the edge colors from the graph into a simple list so that you can visualize the trails by their color.
# +
# Define data structure (list) of edge colors for plotting
edge_colors = [e[2]['attr_dict']['color'] for e in g.edges(data=True)]
# Preview first 10
edge_colors[0:10]
# -
plt.figure(figsize=(8, 6))
nx.draw(g, pos=node_positions, edge_color=edge_colors, node_size=10, node_color='black')
plt.title('Graph Representation of Sleeping Giant Trail Map', size=15)
plt.show()
# ## Solving the Chinese Postman Problem is quite simple conceptually:
#
# 1. Find all nodes with odd degree (very easy).
# (Find all trail intersections where the number of trails touching that intersection is an odd number)
#
#
# 2. Add edges to the graph such that all nodes of odd degree are made even. These added edges must be duplicates from the original graph (we'll assume no bushwhacking for this problem). The set of edges added should sum to the minimum distance possible (hard...np-hard to be precise).
# (In simpler terms, minimize the amount of double backing on a route that hits every trail)
#
#
# 3. Given a starting point, find the Eulerian tour over the augmented dataset (moderately easy).
# (Once we know which trails we'll be double backing on, actually calculate the route from beginning to end)
# ## CPP Step 1: Find Nodes of Odd Degree
#
# This is a pretty straightforward counting computation. You see that 36 of the 76 nodes have odd degree. These are mostly the dead-end trails (degree 1) and intersections of 3 trails. There are a handful of degree 5 nodes.
list(g.nodes(data=True))
# +
# Calculate list of nodes with odd degree
nodes_odd_degree = [v for v, d in g.degree() if d % 2 ==1]
# Preview
(nodes_odd_degree[0:5])
# -
print('Number of nodes of odd degree: {}'.format(len(nodes_odd_degree)))
print('Number of total nodes: {}'.format(len(g.nodes())))
# ## CPP Step 2: Find Min Distance Pairs
#
# This is really the meat of the problem. You'll break it down into 5 parts:
#
# 1. Compute all possible pairs of odd degree nodes.
# 2. Compute the shortest path between each node pair calculated in 1.
# 3. Create a complete graph connecting every node pair in 1. with shortest path distance attributes calculated in 2.
# 4. Compute a minimum weight matching of the graph calculated in 3.
# (This boils down to determining how to pair the odd nodes such that the sum of the distance between the pairs is as small as possible).
# 5. Augment the original graph with the shortest paths between the node pairs calculated in 4.
# # Step 2.1: Compute Node Pairs
#
# You use the itertools combination function to compute all possible pairs of the odd degree nodes. Your graph is undirected, so we don't care about order: For example, (a,b) == (b,a).
# +
# Compute all pairs of odd nodes. in a list of tuples
odd_node_pairs = list(itertools.combinations(nodes_odd_degree, 2))
# Preview pairs of odd degree nodes
odd_node_pairs[0:10]
print('Number of pairs: {}'.format(len(odd_node_pairs)))
# -
def get_shortest_paths_distances(graph, pairs, edge_weight_name):
"""Compute shortest distance between each pair of nodes in a graph. Return a dictionary keyed on node pairs (tuples)."""
distances = {}
for pair in pairs:
distances[pair] = nx.dijkstra_path_length(graph, pair[0], pair[1], weight=edge_weight_name)
return distances
# +
# Compute shortest paths. Return a dictionary with node pairs keys and a single value equal to shortest path distance.
odd_node_pairs_shortest_paths = get_shortest_paths_distances(g, odd_node_pairs, 'distance')
# Preview with a bit of hack (there is no head/slice method for dictionaries).
dict(list(odd_node_pairs_shortest_paths.items())[0:10])
# -
# ## Step 2.3: Create Complete Graph
# A complete graph is simply a graph where every node is connected to every other node by a unique edge.
#
#
# create_complete_graph is defined to calculate it. The flip_weights parameter is used to transform the distance to the weight attribute where smaller numbers reflect large distances and high numbers reflect short distances. This sounds a little counter intuitive, but is necessary for Step 2.4 where you calculate the minimum weight matching on the complete graph.
#
#
# Ideally you'd calculate the minimum weight matching directly, but NetworkX only implements a max_weight_matching function which maximizes, rather than minimizes edge weight. We hack this a bit by negating (multiplying by -1) the distance attribute to get weight. This ensures that order and scale by distance are preserved, but reversed.
# +
def create_complete_graph(pair_weights, flip_weights=True):
"""
Create a completely connected graph using a list of vertex pairs and the shortest path distances between them
Parameters:
pair_weights: list[tuple] from the output of get_shortest_paths_distances
flip_weights: Boolean. Should we negate the edge attribute in pair_weights?
"""
g = nx.Graph()
for k, v in pair_weights.items():
wt_i = - v if flip_weights else v
g.add_edge(k[0], k[1], attr_dict={'distance': v, 'weight': wt_i})
return g
# Generate the complete graph
g_odd_complete = create_complete_graph(odd_node_pairs_shortest_paths, flip_weights=True)
# Counts
print('Number of nodes: {}'.format(len(g_odd_complete.nodes())))
print('Number of edges: {}'.format(len(g_odd_complete.edges())))
# -
# For a visual prop, the fully connected graph of odd degree node pairs is plotted below. Note that you preserve the X, Y coordinates of each node, but the edges do not necessarily represent actual trails. For example, two nodes could be connected by a single edge in this graph, but the shortest path between them could be 5 hops through even degree nodes (not shown here).
# Plot the complete graph of odd-degree nodes
plt.figure(figsize=(8, 6))
pos_random = nx.random_layout(g_odd_complete)
nx.draw_networkx_nodes(g_odd_complete, node_positions, node_size=20, node_color="red")
nx.draw_networkx_edges(g_odd_complete, node_positions, alpha=0.1)
plt.axis('off')
plt.title('Complete Graph of Odd-degree Nodes')
plt.show()
# ## Step 2.4: Compute Minimum Weight Matching
# This is the most complex step in the CPP. You need to find the odd degree node pairs whose combined sum (of distance between them) is as small as possible. So for your problem, this boils down to selecting the optimal 18 edges (36 odd degree nodes / 2) from the hairball of a graph generated in 2.3.
#
# Both the implementation and intuition of this optimization are beyond the scope of this tutorial... like 800+ lines of code and a body of academic literature beyond this scope.
#
# The code implemented in the NetworkX function max_weight_matching is based on <NAME> (1986) [2] which employs an O(n3) time algorithm.
# +
# Compute min weight matching.
# Note: max_weight_matching uses the 'weight' attribute by default as the attribute to maximize.
odd_matching_dupes = nx.algorithms.max_weight_matching(g_odd_complete, True)
print('Number of edges in matching: {}'.format(len(odd_matching_dupes)))
# -
# The matching output (odd_matching_dupes) is a dictionary. Although there are 36 edges in this matching, you only want 18. Each edge-pair occurs twice (once with node 1 as the key and a second time with node 2 as the key of the dictionary).
odd_matching_dupes
list(odd_matching_dupes)
# +
# Convert matching to list of deduped tuples
odd_matching = list(odd_matching_dupes)
# Counts
print('Number of edges in matching (deduped): {}'.format(len(odd_matching)))
# +
plt.figure(figsize=(8, 6))
# Plot the complete graph of odd-degree nodes
nx.draw(g_odd_complete, pos=node_positions, node_size=20, alpha=0.05)
# Create a new graph to overlay on g_odd_complete with just the edges from the min weight matching
g_odd_complete_min_edges = nx.Graph(odd_matching)
nx.draw(g_odd_complete_min_edges, pos=node_positions, node_size=20, edge_color='blue', node_color='red')
plt.title('Min Weight Matching on Complete Graph')
plt.show()
# -
# To illustrate how this fits in with the original graph, you plot the same min weight pairs (blue lines), but over the trail map (faded) instead of the complete graph. Again, note that the blue lines are the bushwhacking route (as the crow flies edges, not actual trails). You still have a little bit of work to do to find the edges that comprise the shortest route between each pair in Step 3.
# +
plt.figure(figsize=(8, 6))
# Plot the original trail map graph
nx.draw(g, pos=node_positions, node_size=20, alpha=0.1, node_color='black')
# Plot graph to overlay with just the edges from the min weight matching
nx.draw(g_odd_complete_min_edges, pos=node_positions, node_size=20, alpha=1, node_color='red', edge_color='blue')
plt.title('Min Weight Matching on Orginal Graph')
plt.show()
# -
# ## Step 2.5: Augment the Original Graph
#
# Now you augment the original graph with the edges from the matching calculated in 2.4. A simple function to do this is defined below which also notes that these new edges came from the augmented graph. You'll need to know this in 3. when you actually create the Eulerian circuit through the graph.
def add_augmenting_path_to_graph(graph, min_weight_pairs):
"""
Add the min weight matching edges to the original graph
Parameters:
graph: NetworkX graph (original graph from trailmap)
min_weight_pairs: list[tuples] of node pairs from min weight matching
Returns:
augmented NetworkX graph
"""
# We need to make the augmented graph a MultiGraph so we can add parallel edges
graph_aug = nx.MultiGraph(graph.copy())
for pair in min_weight_pairs:
graph_aug.add_edge(pair[0],
pair[1],
attr_dict={'distance': nx.dijkstra_path_length(graph, pair[0], pair[1]),
'trail': 'augmented'}
)
return graph_aug
# +
# Create augmented graph: add the min weight matching edges to g
g_aug = add_augmenting_path_to_graph(g, odd_matching)
# Counts
print('Number of edges in original graph: {}'.format(len(g.edges())))
print('Number of edges in augmented graph: {}'.format(len(g_aug.edges())))
# -
# ## CPP Step 3: Compute Eulerian Circuit
# ow that you have a graph with even degree the hard optimization work is over. As Euler famously postulated in 1736 with the Seven Bridges of Königsberg problem, there exists a path which visits each edge exactly once if all nodes have even degree. <NAME> fomally proved this result later in the 1870s.
#
#
# There are many Eulerian circuits with the same distance that can be constructed. You can get 90% of the way there with the NetworkX eulerian_circuit function. However there are some limitations.
# #### Naive Circuit
# Nonetheless, let's start with the simple yet incomplete solution:
naive_euler_circuit = list(nx.eulerian_circuit(g_aug, source='b_end_east'))
print('Length of eulerian circuit: {}'.format(len(naive_euler_circuit)))
naive_euler_circuit[0:10]
# #### Correct Circuit
# Now let's define a function that utilizes the original graph to tell you which trails to use to get from node A to node B. Although verbose in code, this logic is actually quite simple. You simply transform the naive circuit which included edges that did not exist in the original graph to a Eulerian circuit using only edges that exist in the original graph.
#
# You loop through each edge in the naive Eulerian circuit (naive_euler_circuit). Wherever you encounter an edge that does not exist in the original graph, you replace it with the sequence of edges comprising the shortest path between its nodes using the original graph
def create_eulerian_circuit(graph_augmented, graph_original, starting_node=None):
"""Create the eulerian path using only edges from the original graph."""
euler_circuit = []
naive_circuit = list(nx.eulerian_circuit(graph_augmented, source=starting_node))
for edge in naive_circuit:
edge_data = graph_augmented.get_edge_data(edge[0], edge[1])
#print(edge_data[0])
if edge_data[0]['attr_dict']['trail'] != 'augmented':
# If `edge` exists in original graph, grab the edge attributes and add to eulerian circuit.
edge_att = graph_original[edge[0]][edge[1]]
euler_circuit.append((edge[0], edge[1], edge_att))
else:
aug_path = nx.shortest_path(graph_original, edge[0], edge[1], weight='distance')
aug_path_pairs = list(zip(aug_path[:-1], aug_path[1:]))
print('Filling in edges for augmented edge: {}'.format(edge))
print('Augmenting path: {}'.format(' => '.join(aug_path)))
print('Augmenting path pairs: {}\n'.format(aug_path_pairs))
# If `edge` does not exist in original graph, find the shortest path between its nodes and
# add the edge attributes for each link in the shortest path.
for edge_aug in aug_path_pairs:
edge_aug_att = graph_original[edge_aug[0]][edge_aug[1]]
euler_circuit.append((edge_aug[0], edge_aug[1], edge_aug_att))
return euler_circuit
# Create the Eulerian circuit
euler_circuit = create_eulerian_circuit(g_aug, g, 'b_end_east')
print('Length of Eulerian circuit: {}'.format(len(euler_circuit)))
# +
## CPP Solution
# Preview first 20 directions of CPP solution
for i, edge in enumerate(euler_circuit[0:20]):
print(i, edge)
# -
# ## Stats
# +
# Computing some stats
total_mileage_of_circuit = sum([edge[2]['attr_dict']['distance'] for edge in euler_circuit])
total_mileage_on_orig_trail_map = sum(nx.get_edge_attributes(g, 'distance').values())
_vcn = pd.value_counts(pd.value_counts([(e[0]) for e in euler_circuit]), sort=False)
node_visits = pd.DataFrame({'n_visits': _vcn.index, 'n_nodes': _vcn.values})
_vce = pd.value_counts(pd.value_counts([sorted(e)[0] + sorted(e)[1] for e in nx.MultiDiGraph(euler_circuit).edges()]))
edge_visits = pd.DataFrame({'n_visits': _vce.index, 'n_edges': _vce.values})
# Printing stats
print('Mileage of circuit: {0:.2f}'.format(total_mileage_of_circuit))
print('Mileage on original trail map: {0:.2f}'.format(total_mileage_on_orig_trail_map))
print('Mileage retracing edges: {0:.2f}'.format(total_mileage_of_circuit-total_mileage_on_orig_trail_map))
#print('Percent of mileage retraced: {0:.2f}%\n'.format((1-total_mileage_of_circuit/total_mileage_on_orig_trail_map)*-100))
print('Number of edges in circuit: {}'.format(len(euler_circuit)))
print('Number of edges in original graph: {}'.format(len(g.edges())))
print('Number of nodes in original graph: {}\n'.format(len(g.nodes())))
print('Number of edges traversed more than once: {}\n'.format(len(euler_circuit)-len(g.edges())))
print('Number of times visiting each node:')
print(node_visits.to_string(index=False))
print('\nNumber of times visiting each edge:')
print(edge_visits.to_string(index=False))
# -
# ## Create CPP Graph
# Your first step is to convert the list of edges to walk in the Euler circuit into an edge list with plot-friendly attributes.
#
#
def create_cpp_edgelist(euler_circuit):
"""
Create the edgelist without parallel edge for the visualization
Combine duplicate edges and keep track of their sequence and # of walks
Parameters:
euler_circuit: list[tuple] from create_eulerian_circuit
"""
cpp_edgelist = {}
for i, e in enumerate(euler_circuit):
edge = frozenset([e[0], e[1]])
if edge in cpp_edgelist:
cpp_edgelist[edge][2]['sequence'] += ', ' + str(i)
cpp_edgelist[edge][2]['visits'] += 1
else:
cpp_edgelist[edge] = e
cpp_edgelist[edge][2]['sequence'] = str(i)
cpp_edgelist[edge][2]['visits'] = 1
return list(cpp_edgelist.values())
cpp_edgelist = create_cpp_edgelist(euler_circuit)
print('Number of edges in CPP edge list: {}'.format(len(cpp_edgelist)))
cpp_edgelist[0:3]
# +
g_cpp = nx.Graph(cpp_edgelist)
plt.figure(figsize=(14, 10))
visit_colors = {1:'lightgray', 2:'blue', 3: 'red', 4 : 'black', 5 : 'green'}
edge_colors = [visit_colors[e[2]['visits']] for e in g_cpp.edges(data=True)]
node_colors = ['red' if node in nodes_odd_degree else 'lightgray' for node in g_cpp.nodes()]
nx.draw_networkx(g_cpp, pos=node_positions, node_size=20, node_color=node_colors, edge_color=edge_colors, with_labels=False)
plt.axis('off')
plt.show()
# +
plt.figure(figsize=(14, 10))
edge_colors = [e[2]['attr_dict']['color'] for e in g_cpp.edges(data=True)]
nx.draw_networkx(g_cpp, pos=node_positions, node_size=10, node_color='black', edge_color=edge_colors, with_labels=False, alpha=0.5)
bbox = {'ec':[1,1,1,0], 'fc':[1,1,1,0]} # hack to label edges over line (rather than breaking up line)
edge_labels = nx.get_edge_attributes(g_cpp, 'sequence')
nx.draw_networkx_edge_labels(g_cpp, pos=node_positions, edge_labels=edge_labels, bbox=bbox, font_size=6)
plt.axis('off')
plt.show()
# -
visit_colors = {1:'lightgray', 2:'blue', 3: 'red', 4 : 'black', 5 : 'green'}
edge_cnter = {}
g_i_edge_colors = []
for i, e in enumerate(euler_circuit, start=1):
edge = frozenset([e[0], e[1]])
if edge in edge_cnter:
edge_cnter[edge] += 1
else:
edge_cnter[edge] = 1
# Full graph (faded in background)
nx.draw_networkx(g_cpp, pos=node_positions, node_size=6, node_color='gray', with_labels=False, alpha=0.07)
# Edges walked as of iteration i
euler_circuit_i = copy.deepcopy(euler_circuit[0:i])
for i in range(len(euler_circuit_i)):
edge_i = frozenset([euler_circuit_i[i][0], euler_circuit_i[i][1]])
euler_circuit_i[i][2]['visits_i'] = edge_cnter[edge_i]
g_i = nx.Graph(euler_circuit_i)
g_i_edge_colors = [visit_colors[e[2]['visits_i']] for e in g_i.edges(data=True)]
nx.draw_networkx_nodes(g_i, pos=node_positions, node_size=6, alpha=0.6, node_color='lightgray', with_labels=False, linewidths=0.1)
nx.draw_networkx_edges(g_i, pos=node_positions, edge_color=g_i_edge_colors, alpha=0.8)
plt.axis('off')
plt.savefig('img{}.png'.format(i), dpi=120, bbox_inches='tight')
plt.close()
# +
import glob
import numpy as np
import imageio
import os
def make_circuit_video(image_path, movie_filename, fps=7):
# sorting filenames in order
filenames = glob.glob(image_path + 'img*.png')
filenames_sort_indices = np.argsort([int(os.path.basename(filename).split('.')[0][3:]) for filename in filenames])
filenames = [filenames[i] for i in filenames_sort_indices]
# make movie
with imageio.get_writer(movie_filename, mode='I', fps=fps) as writer:
for filename in filenames:
image = imageio.imread(filename)
writer.append_data(image)
make_circuit_video('', 'cpp_route_animation.gif', fps=3)
# -
| general studies/Graphs/Intro-networkx-python-graph-tutorial.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Raku
# language: raku
# name: raku
# ---
# # Aufgabentitel
# Hier Text, Quelle, etc. ...
#
# Cell -> Run All
# +
#Falls mit Einheiten gearbeitet werden soll
# use Units <m A Ω K ft in>;
# say 8A * 2Ω; # Output: 8V
# say 0K.celsius.value; # Output: -273.15
# say 1ft > 1in; # Output: True
# Oder dieses Modul hier (wie immer gibt es vielfache Möglichkeiten ...)
# https://github.com/p6steve/perl6-Physics-Measure-JupyterBinder/blob/master/Synopsis.ipynb
# use Physics::Measure;
# use Physics::Unit;
# use Physics::Constants;
# für wissenschaftliches Zahlenformat
# https://docs.perl6.org/routine/sprintf
# my $a = 0.000000000000012 ;
# sprintf '%g', $a;
# +
#Geg.
my $E = 500; #V/mm
my $Q = 0.000000025; #C
#Ges:
my $F = Nil;
#Lösung
$F = 1/4 * pi / 100_000_000; # * bla blub bli
# -
#Ergebnis
say "Ohne Formatierung " ~ $F;
say "Mit Formatierung " ~ sprintf '%g', $F;
| demo/template_Raku.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import cv2
import matplotlib.pyplot as plt
from keras import models
import keras.backend as K
import tensorflow as tf
from sklearn.metrics import f1_score
import requests
import xmltodict
import json
plateCascade = cv2.CascadeClassifier('indian_license_plate.xml')
#detect the plate and return car + plate image
def plate_detect(img):
plateImg = img.copy()
roi = img.copy()
plateRect = plateCascade.detectMultiScale(plateImg,scaleFactor = 1.2, minNeighbors = 7)
for (x,y,w,h) in plateRect:
roi_ = roi[y:y+h, x:x+w, :]
plate_part = roi[y:y+h, x:x+w, :]
cv2.rectangle(plateImg,(x+2,y),(x+w-3, y+h-5),(0,255,0),3)
return plateImg, plate_part
#normal function to display
def display_img(img):
img_ = cv2.cvtColor(img,cv2.COLOR_BGR2RGB)
plt.imshow(img_)
plt.show()
#test image is used for detecting plate
inputImg = cv2.imread('test.jpeg')
inpImg, plate = plate_detect(inputImg)
display_img(inpImg)
def find_contours(dimensions, img) :
#finding all contours in the image using
#retrieval mode: RETR_TREE
#contour approximation method: CHAIN_APPROX_SIMPLE
cntrs, _ = cv2.findContours(img.copy(), cv2.RETR_TREE, cv2.CHAIN_APPROX_SIMPLE)
#Approx dimensions of the contours
lower_width = dimensions[0]
upper_width = dimensions[1]
lower_height = dimensions[2]
upper_height = dimensions[3]
#Check largest 15 contours for license plate character respectively
cntrs = sorted(cntrs, key=cv2.contourArea, reverse=True)[:15]
ci = cv2.imread('contour.jpg')
x_cntr_list = []
target_contours = []
img_res = []
for cntr in cntrs :
#detecting contour in binary image and returns the coordinates of rectangle enclosing it
intX, intY, intWidth, intHeight = cv2.boundingRect(cntr)
#checking the dimensions of the contour to filter out the characters by contour's size
if intWidth > lower_width and intWidth < upper_width and intHeight > lower_height and intHeight < upper_height :
x_cntr_list.append(intX)
char_copy = np.zeros((44,24))
#extracting each character using the enclosing rectangle's coordinates.
char = img[intY:intY+intHeight, intX:intX+intWidth]
char = cv2.resize(char, (20, 40))
cv2.rectangle(ci, (intX,intY), (intWidth+intX, intY+intHeight), (50,21,200), 2)
plt.imshow(ci, cmap='gray')
char = cv2.subtract(255, char)
char_copy[2:42, 2:22] = char
char_copy[0:2, :] = 0
char_copy[:, 0:2] = 0
char_copy[42:44, :] = 0
char_copy[:, 22:24] = 0
img_res.append(char_copy) # List that stores the character's binary image (unsorted)
#return characters on ascending order with respect to the x-coordinate
plt.show()
#arbitrary function that stores sorted list of character indeces
indices = sorted(range(len(x_cntr_list)), key=lambda k: x_cntr_list[k])
img_res_copy = []
for idx in indices:
img_res_copy.append(img_res[idx])# stores character images according to their index
img_res = np.array(img_res_copy)
return img_res
def segment_characters(image) :
#pre-processing cropped image of plate
#threshold: convert to pure b&w with sharpe edges
#erod: increasing the backgroung black
#dilate: increasing the char white
img_lp = cv2.resize(image, (333, 75))
img_gray_lp = cv2.cvtColor(img_lp, cv2.COLOR_BGR2GRAY)
_, img_binary_lp = cv2.threshold(img_gray_lp, 200, 255, cv2.THRESH_BINARY+cv2.THRESH_OTSU)
img_binary_lp = cv2.erode(img_binary_lp, (3,3))
img_binary_lp = cv2.dilate(img_binary_lp, (3,3))
LP_WIDTH = img_binary_lp.shape[0]
LP_HEIGHT = img_binary_lp.shape[1]
img_binary_lp[0:3,:] = 255
img_binary_lp[:,0:3] = 255
img_binary_lp[72:75,:] = 255
img_binary_lp[:,330:333] = 255
#estimations of character contours sizes of cropped license plates
dimensions = [LP_WIDTH/6,
LP_WIDTH/2,
LP_HEIGHT/10,
2*LP_HEIGHT/3]
plt.imshow(img_binary_lp, cmap='gray')
plt.show()
cv2.imwrite('contour.jpg',img_binary_lp)
#getting contours
char_list = find_contours(dimensions, img_binary_lp)
return char_list
char = segment_characters(plate)
for i in range(10):
plt.subplot(1, 10, i+1)
plt.imshow(char[i], cmap='gray')
plt.axis('off')
# +
#It is the harmonic mean of precision and recall
#Output range is [0, 1]
#Works for both multi-class and multi-label classification
def f1score(y, y_pred):
return f1_score(y, tf.math.argmax(y_pred, axis=1), average='micro')
def custom_f1score(y, y_pred):
return tf.py_function(f1score, (y, y_pred), tf.double)
# -
model = models.load_model('license_plate_character.pkl', custom_objects= {'custom_f1score': custom_f1score})
# +
def fix_dimension(img):
new_img = np.zeros((28,28,3))
for i in range(3):
new_img[:,:,i] = img
return new_img
def show_results():
dic = {}
characters = '0123456789ABCDEFGHIJKLMNOPQRSTUVWXYZ'
for i,c in enumerate(characters):
dic[i] = c
output = []
for i,ch in enumerate(char):
img_ = cv2.resize(ch, (28,28), interpolation=cv2.INTER_AREA)
img = fix_dimension(img_)
img = img.reshape(1,28,28,3)
y_ = model.predict_classes(img)[0]
character = dic[y_] #
output.append(character)
plate_number = ''.join(output)
return plate_number
final_plate = show_results()
print(final_plate)
# -
def get_vehicle_info(plate_number):
r = requests.get("http://www.regcheck.org.uk/api/reg.asmx/CheckIndia?RegistrationNumber={0}&username=licenseguy".format(str(plate_number)))
data = xmltodict.parse(r.content)
jdata = json.dumps(data)
df = json.loads(jdata)
df1 = json.loads(df['Vehicle']['vehicleJson'])
return df1
if len(final_plate) > 10:
final_plate = final_plate[-10:]
print(final_plate)
get_vehicle_info(final_plate)
| .ipynb_checkpoints/testing_of_model-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Programming in Python
# ## Session 1
# ### Aim of the Session
# Learn/review the basics
# - what is ...
# - how to ...
# ### 'Hello World!'
# the culturally-expected introductory statement
print("Hello world!")
# an example of what you can achieve with Python in just a few lines
# ### Literals
# Values of a _type_, presented literally
# +
# example name type designation
42 # integer int
2.016 # float float*
"Homo sapiens" # string str
print(2.016)
print(1.0e-4)
print(.54)
# -
# - int: whole numbers e.g. 1, 1000, 6000000000
# - float: 'floating point' non-whole numbers e.g. 1.9, 30.01, 10e3, 1e-3
# - string: ordered sequence of characters, enclosed in quotation marks (single, double, _triple_)
# type conversions
print(float("1.31"))
print(int("-42"))
# #### Aside - Comments
#
# Comments are preceded by a **#**, and are completely ignored by the python interpreter.
# Comments can be on their own line or after a line of code.
#
# Comments are an incredibly useful way to keep track of what you are doing in
# your code. Use comments to document what you do as much as possible, it will
# pay off in the long run.
#
# ### Exercises 1
# print some strings
print("this a string")
# print some numbers (ints or floats)
print(42)
print(3.14)
# print multiple values of different types all at once
# (hints: use comma to separate values with a space, or + to join strings)
print("This is a number:", 3.14)
print("This a " + "long string")
print("This a number: " + str(4))
# +
# print a string containing quote marks
print("This a string with a quote \" mark")
print('This is a " string')
print("""This a string
over more than one lines""")
# -
# ### Operators & Operands
# Using Python as a calculator: `+`, `-`, `/`, `*` etc are _operators_, the values/variables that they work on are _operands_.
# +
# standard mathematical operations can be performed in Python
print(3+4)
print(4-5)
print(3.14 * 2)
print(8 / 3)
# and some less common ones
print(2 ** 8)
print(35 % 10)
# -
# _Note: check out numpy, scipy, stats modules if you want to do a lot of maths_
# ### Variables
# Store values (information) in memory, and (re-)use them. We give variables names (identifiers) so that we have a means of referring to the information on demand.
# +
# variable assignment is done with '='
my_variable = 3.14
print(my_variable)
my_variable = 72
print(my_variable)
my_variable = 71
another_variable = 12 + 35
print(another_variable)
yet_another_variable = my_variable + another_variable
print(yet_another_variable)
# -
3 + 7
2 - 19
# #### Variable naming
# Rules:
#
# - identifier lookup is case-sensitive
# - `myname` & `MyName` are different
# - must be unique in your working environment
# - existing variable will be __over-written without warning__
# - cannot start with a number, or any special symbol (e.g. $, %, @, -, etc...) except for "_" (underscore), which is OK.
# - cannot have any spaces or special characters (except for "_" (underscore))
#
# Conventions/good practice:
#
# - identifiers (usually) begin with a lowercase letter
# - followed by letters, numbers, underscores
# - use a strategy to make reading easier
# - `myName`
# - `exciting_variable`
# - long, descriptive > short, vague
# ### String Formatting
# Create formatted strings, with variable values substituted in.
# +
# two ways to do it in Python
name = 'Florence'
age = 73
print('%s is %d years old' % (name, age)) # common amongst many programming languages
print('{} is {} years old'.format(name, age)) # perhaps more consistent with stardard Python syntax
# -
# There is a long list of possible format options for numbers: https://pyformat.info/
# ### Data Structures
# Programming generally requires building/working with much larger and more complex sets of data than the single values/words/sentences that we have looked at so far. In fact, finding ways to operate effectively (and efficiently) on complex structures in order to extract/produce information, _is_ (data) programming.
#
# Python has two most commonly-used structures for storing multiple pieces of data - _lists_ and _dictionaries_. Let's look at these, and a few more, now.
# #### Lists
# +
# sequence of entries, in order and of any type
print([2, 5, 7, 9])
print([4.5, 3.1, 7.8])
print([1, 2, 'abc', 'cde'])
my_list = [1, 2, 6, 'abc', 'cdef']
# accessing list entries
print(my_list[0])
print(my_list[2])
print(my_list[0:3])
print(my_list[1:4])
# adding/removing entries (remove/pop)
# length of list
# +
my_list = [1, 5, 7]
print(my_list)
my_list.append(10)
print(my_list)
my_list.remove(5)
print(my_list)
print(len(my_list))
# -
my_list = [1, 2, 6, 'abc', 'cdef']
print(my_list[-1])
print(my_list[-3:])
# sets
my_list = [1, 1, 2, 4, 7, 7, 2]
print(my_list)
my_set = set(my_list)
print(my_set)
# #### Objects, Methods, and How To Get Help
# In Python, everything is an _object_ - some value(s), packaged up with a set of things that can be done with/to it (___methods___), and pieces of information about it (___attributes___). This makes it very easy to perform the most commonly-needed operations for that/those type of value(s). The language has a standard syntax for accessing methods:
# +
string_object = 'the cold never bothered me anyway'
# methods - object.something()
print(string_object.upper())
print(string_object)
another_string = string_object.upper()
print(another_string)
# more...
print(string_object.capitalize())
# -
help(string_object.replace)
string_object.replace("e", "a", 2)
# +
# dir() and help()
# -
# ### Exercises 2
# add 'Sally' to the list of students' names
student_names = ['Sandy', 'Pete', 'Richard', 'Rebecca']
student_names.append("Sally")
print(student_names)
# access the fourth entry of the list
print(student_names[3])
print(student_names[10])
# +
# join the list with a new list from another class
other_student_names = ['Sam', 'Fiona', 'Sarah', 'Richard', 'Sarah', 'Matthew']
combined_names = student_names + other_student_names
print(combined_names)
# -
student_names = student_names + other_student_names
my_list = [1, 2]
my_list.append(3)
print(my_list)
my_string = "a string"
my_string = my_string.upper()
print(my_string)
help(my_string.upper)
# +
print(sorted(student_names))
# -
# #### Dictionaries
# +
# collection of paired information - keys and values
student_marks = {'Alessio': 67, 'Nic': 48, 'Georg': 68}
empty_dict = {}
another_empty_dict = dict()
# accessing dict entries
# adding/changing/deleting entries
# -
student_age = {'Alessio': 26, 'Nic': 28, 'Georg': 32}
print(student_age)
another_dict = {1: 'a', 2: 'c', 3: 'e'}
print(another_dict)
# +
print(student_marks)
print(student_marks['Nic'])
# -
print(student_marks.keys())
print(student_marks.values())
student_marks['Tim'] = 62
print(student_marks)
student_marks['Nic'] = 47
print(student_marks)
student_marks.pop('Tim')
print(student_marks)
# #### Mutable?
# Object types can be divided into two categories - mutable & immutable. _Mutable_ objects can be changed 'in-place' - their value can be updated, added to, re-ordered etc without the need to create a whole new object every time. _Immutable_ types cannot be changed in place - once they have a value, this value cannot be altered. though, of course, it can __always__ be overwritten.
# lists are mutable
cities = ['Nairobi', 'Vancouver', 'Wellington', 'Beijing']
print(cities)
# strings are immutable
beatles = "I'd like to be under the sea"
# ### Looping
# Time for some real programming. The biggest motivation for researches to learn a programming language is the opportunity to automate repetitive tasks and analyses.
#
# For loops define a set of steps that will be carried out for all items in a sequence. The items in the sequence will be taken one-at-a-time, and the loop performed, until there are no more items to process.
for season in ['Spring', 'Summer', 'Autumn', 'Winter']:
print(season)
for number in [1, 5, 87, 3]:
print(number)
print("------")
# +
word = 'python'
for letter in word:
print(letter.upper())
print("Finished")
# +
# range
# zip
# iterating through two lists simultaneously
# enumerate
# -
for number in range(5):
print("The number is ", number)
for number in range(1,5):
print("The number is", number)
names = ['Sally', 'Linda', 'Tim']
grades = [46, 52, 45]
for name, grade in zip(names, grades):
print("Name: ", name, " Grade:", grade)
for index, name in enumerate(names):
print("Position", index, " has name ", name)
# ## Exercise 3:
# +
# calculate the mean of the elements in the list
list_of_numbers = [1, 2, 4, 8, 3, 6, 1, 9, 10, 5]
sum_of_values = ...
for ... in list_of_numbers:
... = ... + ...
mean_value = ... / len(...)
print(mean_value)
# -
# ### Conditionals - if, elif, else
# Looping allows you to perform a common set of operations on multiple pieces of data very quickly. But what if you want to treat the pieces differently, depending on some property or other of the objects?
#
# This is the other central part of programming: testing for certain circumstances and changing the treatment of pieces of data accordingly. It is known as _flow control_, as you are controlling the flow of data through your script of operations.
# #### if - elif - else
# +
# use if statements to test for a condition (comparison, equality)
# use else to dictate what happens when the condition isn't met
# use elif to add more conditionals
# use more than one condition check
# -
temperature = 18
if temperature > 20:
print("T-shirt weather")
else:
print("Bring a jacket")
weather = 'foggy'
if weather == 'rainy':
print("Bring an umbrella")
elif weather == "sunny":
print("Bring sunscreen")
else:
print("Dont bring an umbrella")
temperature = 15
if temperature < 20:
if temperature > 0:
print("Bring a light jacket")
temperature = 15
if temperature < 20 and temperature > 0:
print("Bring a light jacket")
weather = 'foggy'
if weather == 'foggy' or weather == 'rainy':
print("No sunshine")
# +
# list comprehensions with conditionals
| Python/Jupyter_notebooks_solved/SwC_python_session-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Django Shell-Plus
# language: python
# name: django_extensions
# ---
import re
result = """
img alt="" height="200" src="/media/images/test.width-300.jpegquality-90.jpg" srcset="/media/images/test.width-2200.jpegquality-60_4QIlsA0.jpg 200w, /media/images/test.width-1100.jpegquality-60_GPzO0UW.jpg 200w, /media/images/test.width-768.jpegquality-60_QaWXMP8.jpg 200w, /media/images/test.width-500.jpegquality-60_7agJtb0.jpg 200w, /media/images/test.width-300.jpegquality-60_tXuUjzT.jpg 200w" width="200">
"""
srcset = re.search(r'srcset="(?P<srcset>.*?)"', result).group("srcset")
"jpegquality-90" in srcset
str(1100) in "asdf 1100 asdf"
"asdf".split("-")
from wagtail.images.models import Image as WagtailImage
WagtailImage.objects.all()
image = WagtailImage.objects.first()
for rendition in image.renditions.all():
print(rendition)
rendition = Rendition.objects.first()
dir(rendition)
rendition.attrs
rendition.width
rendition.attrs_dict
Rendition.objects.filter(width=600)
image.renditions.filter(width=600)
image.refresh_from_db
help(getattr)
image.width
def default_sizes(size):
for size in (2200, 1100, 768, 500, 300):
print(size / 2200)
max((2200, 1100, 768, 500, 300))
default_scales = [size / 2200 for size in (2200, 1100, 768, 500, 300)]
img_width = 2800
for scale in default_scales:
print(int(scale * img_width))
name, rest = "width-234".split("-")
rest
max(2, None)
int(None)
"".split("|")
name, *rest = "".split()
| notebooks/debug.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## This notebook shows how to use Orchestrator APIs for user experiments
import os
from fabrictestbed.slice_manager import SliceManager, Status, SliceState
import json
# +
ssh_key_file_priv=os.environ['HOME']+"/.ssh/id_rsa"
ssh_key_file_pub=os.environ['HOME']+"/.ssh/id_rsa.pub"
ssh_key_pub = None
with open (ssh_key_file_pub, "r") as myfile:
ssh_key_pub=myfile.read()
ssh_key_pub=ssh_key_pub.strip()
# -
slice_name='Slice-l2bridge-mixed'
# ## Create Slice Manager Object
# Users can request tokens with different Project and Scopes by altering `project_name` and `scope` parameters in the refresh call below.
slice_manager = SliceManager()
# ### Orchestrator API example to query for available resources
# +
status, advertised_topology = slice_manager.resources()
print(f"Status: {status}")
if status == Status.OK:
print(f"Toplogy: {advertised_topology}")
else:
print(f"Error: {advertised_topology}")
# -
if status == Status.OK:
advertised_topology.draw()
# ## Create Slice
#
# In Release 1.0, user is expected to assign the IP addresses manually. Please use the example comands indicated below:
#
# ### Run on Node N1
# ```
# ip addr add 192.168.10.51/24 dev eth1
# ```
# ### Run on Node N2
# ```
# ip addr add 192.168.10.52/24 dev eth1
# ```
# +
from fabrictestbed.slice_editor import ExperimentTopology, Capacities, ComponentType, ComponentModelType, ServiceType
# Create topology
t = ExperimentTopology()
# Add node
n1 = t.add_node(name='n1', site='LBNL')
# Set capacities
cap = Capacities()
cap.set_fields(core=2, ram=6, disk=10)
# Set Properties
n1.set_properties(capacities=cap, image_type='qcow2', image_ref='default_centos_8')
# Add PCI devices
n1.add_component(ctype=ComponentType.NVME, model='P4510', name='c1')
# Add node
n2 = t.add_node(name='n2', site='LBNL')
# Set properties
n2.set_properties(capacities=cap, image_type='qcow2', image_ref='default_centos_8')
# Mixed Cards
n1.add_component(model_type=ComponentModelType.SharedNIC_ConnectX_6, name='n1-nic1')
n2.add_component(model_type=ComponentModelType.SmartNIC_ConnectX_5, name='n2-nic1')
# L2Bridge Service
t.add_network_service(name='bridge1', nstype=ServiceType.L2Bridge, interfaces=t.interface_list)
# Generate Slice Graph
slice_graph = t.serialize()
# Request slice from Orchestrator
status, reservations = slice_manager.create(slice_name=slice_name, slice_graph=slice_graph, ssh_key=ssh_key_pub)
print("Response Status {}".format(status))
if status == Status.OK:
print("Reservations created {}".format(reservations))
else:
print(f"Failure: {reservations}")
# -
# ## Query Slices
# +
status, slices = slice_manager.slices(excludes=[SliceState.Closing, SliceState.Dead])
print("Response Status {}".format(status))
if status == Status.OK:
print("Slices {}".format(slices))
slice_object=list(filter(lambda s: s.slice_name == slice_name, slices))[0]
else:
print(f"Failure: {slices}")
# -
# ## Query Slivers
# +
status, slivers = slice_manager.slivers(slice_object=slice_object)
print("Response Status {}".format(status))
if status == Status.OK:
print("Slivers {}".format(slivers))
else:
print(f"Failure: {slivers}")
# -
# ## Sliver Status
for s in slivers:
status, sliver_status = slice_manager.sliver_status(sliver=s)
print("Response Status {}".format(status))
if status == Status.OK:
print()
print("Sliver Status {}".format(sliver_status))
print()
# ## Delete Slice
# +
status, result = slice_manager.delete(slice_object=slice_object)
print("Response Status {}".format(status))
print("Response received {}".format(result))
# -
| fabric_examples/beta_functionality/l2bridge-mixed.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import csv
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn
from sklearn.preprocessing import StandardScaler
from sklearn.cluster import KMeans
# -
dataset = pd.read_csv('HW2_pokemon.csv')
class_mapping = {label:idx for idx,label in enumerate(set(dataset['Body_Style']))}
dataset['Body_Style'] = dataset['Body_Style'].map(class_mapping)
print(dataset.shape)
dataset_num = dataset[['Total','HP','Attack','Defense','Sp_Atk','Sp_Def','Speed', 'Height_m', 'Weight_kg','Body_Style' ]]
ss = StandardScaler()
dataset_num = ss.fit_transform(dataset_num)
dataset_scaled = dataset.copy()
dataset_scaled[['Total','HP','Attack','Defense','Sp_Atk','Sp_Def','Speed', 'Height_m', 'Weight_kg' ,'Body_Style']] = dataset_num
# +
# loss = []
# for i in range(1, 101):
# kmeans = KMeans(n_clusters = i, init = 'k-means++', random_state = 42)
# kmeans.fit(dataset_num)
# loss.append(kmeans.inertia_)
# plt.plot(range(1, 101), loss)
# plt.xticks(np.arange(1, 101, 1.0))
# plt.grid( axis='x')
# plt.show()
# +
# n_clusters 分成多少群
kmeans = KMeans(n_clusters = 280, init = 'k-means++', random_state = 280)
y_kmeans = kmeans.fit_predict(dataset_num)
# 画出 K-means 的分类情况
dataset['y_kmeans'] = y_kmeans
seaborn.violinplot(x='y_kmeans', y='Sp_Atk', data=dataset)
plt.show()
# +
print("y_kmeans : ", type(y_kmeans), y_kmeans.shape)
print(y_kmeans[:20])
test_dataset = pd.read_csv('subject.csv')
print(type(test_dataset), test_dataset.shape)
index = 0
res = []
# MIDDLE = 29
for td1,td2 in zip(test_dataset["0"], test_dataset["1"]):
td1_i = int(td1[7:])
td2_i = int(td2[7:])
if y_kmeans[td1_i] == y_kmeans[td2_i]:
# 后来发现这种判断不能用,会大幅降低判断的准确性
# if y_kmeans[td1_i] < MIDDLE and y_kmeans[td2_i] < MIDDLE:
# print(index, td1_i, td2_i, y_kmeans[td1_i] , y_kmeans[td2_i])
res.append([index, 1])
# elif y_kmeans[td1_i] >= MIDDLE and y_kmeans[td2_i] >= MIDDLE:
# res.append([index, 1])
else:
res.append([index, 0])
index += 1
print(res[:10])
# 把預測的結果生成 kaggle要求的格式
# pair,值為0~999,第二欄取名:answer
res_csv_file_path = "result.csv"
with open(res_csv_file_path, "w") as output:
writer = csv.writer(output, lineterminator='\n')
writer.writerow(('pair', 'answer'))
ids = 0
for val in res:
writer.writerow((str(ids),val[1]))
ids += 1
print("---------------execute finished.")
# -
| homework2/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import math as m
from scipy.spatial.distance import cdist
from sklearn.datasets import make_blobs
import scipy as sp
from numba import njit
# +
@njit
def hand_norm(A):
return m.sqrt(np.sum(A ** 2))
@njit
def hand_scalar_prod(A,B):
prod = np.zeros((len(A)))
k = 0
for a,b in (zip(A,B)):
prod[k]= a * b
k +=1
return np.sum(prod)
@njit
def hand_dist(A,B, metric = 'euclidean'):
dist = np.zeros((len(A),(len(A))))
if metric == 'euclidean':
for i in range(len(A)):
for ii in range(len(B)):
dist[ii,i] = m.sqrt(np.sum((A[i,:] - B[ii,:]) ** 2))
if metric == 'cosine':
for i in range(len(A)):
for ii in range(len(B)):
dist[ii,i] = 1 - (hand_scalar_prod(A[i,:],B[ii,:])/(hand_norm(A[i,:])*hand_norm(B[ii,:])))
if metric == 'mahalanobis':
concat = np.zeros((len(A)+len(B),len(A[0])))
concat[:len(A)] = A
concat[len(A):] = B
VI = np.linalg.inv(np.cov(concat.T)).T
for i in range(len(A)):
for ii in range(len(B)):
dist[ii,i] = np.sqrt(np.dot(np.dot((A[i,:]-B[ii,:]),VI),(A[i,:]-B[ii,:]).T))
return dist
# -
centers = [(-5, -5), (0, 0), (5, 5)]
data, _ = make_blobs(n_samples=10, centers=centers, shuffle=False,
random_state=42)
data
scipy = cdist(data, data, metric='euclidean')
thiago = hand_dist(data, data, metric='euclidean')
thiago - scipy
scipy = cdist(data, data, metric='cosine')
thiago = hand_dist(data, data, metric='cosine')
thiago - scipy
scipy = cdist(data, data, metric='mahalanobis')
thiago = hand_dist(data, data, metric='mahalanobis')
thiago - scipy
| Python/cdist_study.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + nbsphinx="hidden"
import sys
sys.path.append('../')
# import os
# os.environ['PYTHONASYNCIODEBUG'] = '1'
# -
# Note: you can try this tutorial in [](https://mybinder.org/v2/gh/zh217/aiochan/master?filepath=doc%2Fparallel.ipynb).
# # Parallelism and beyond
# We discussed `async_pipe` and `async_pipe_unordered` in the context of trying to put more "concurrency" into our program by taking advantage of parallelism. What does "parallelism" mean here?
# ## Facing the reality of python concurrency, again
# With `async_pipe` and `async_pipe_unordered`, by giving them more coroutine instances to work with, we achieved higher throughput. But that is only because our coroutines are, in a quite literal sense, sleeping on the job: to simulate real jobs, we called `await` on `asyncio.sleep`. The event loop, faced with this await, just puts the coroutine on hold until it is ready to act again.
# Now it is entirely possible that this behaviour --- of not letting sleeping coroutines block the whole program --- is all you need. In particular, if you are dealing with network connections or sockets *and* you are using a proper asyncio-based library, then "doing network work" isn't too much from sleeping on the loop.
# However, for other operations *not* tailored for asyncio, you will *not* get any speed-up with parallelism based on asyncio. Crucially, *asyncio has no built-in support for file accesses*.
# Let's see an example:
# +
import asyncio
import time
import aiochan as ac
async def worker(n):
time.sleep(0.1) # await asyncio.sleep(0.1)
return n*2
async def main():
start = asyncio.get_event_loop().time()
print(await ac.from_range(20).async_pipe(10, worker).collect())
print(asyncio.get_event_loop().time() - start)
ac.run(main())
# -
# The only different than before (when we first introduced `async_pipe`) is that we replaced `asyncio.sleep` with `time.sleep`. With this change, we did not get *any* speed up.
# In this case, we can recover our speed-up by using the method `parallel_pipe` instead:
# +
import asyncio
import time
import aiochan as ac
def worker(n):
time.sleep(0.1)
return n*2
async def main():
start = asyncio.get_event_loop().time()
print(await ac.from_range(20).parallel_pipe(10, worker).collect())
print(asyncio.get_event_loop().time() - start)
ac.run(main())
# -
# When using `parallel_pipe`, our `worker` has to be a normal function instead of an async function. As before, if order is not important, `parallel_pipe_unordered` can give you even more throughput:
# +
import asyncio
import time
import random
import aiochan as ac
def worker(n):
time.sleep(random.uniform(0, 0.2))
return n*2
async def main():
start = asyncio.get_event_loop().time()
print(await ac.from_range(20).parallel_pipe(10, worker).collect())
print('ordered time:', asyncio.get_event_loop().time() - start)
start = asyncio.get_event_loop().time()
print(await ac.from_range(20).parallel_pipe_unordered(10, worker).collect())
print('unordered time:', asyncio.get_event_loop().time() - start)
ac.run(main())
# -
# In fact, `parallel_pipe` works by starting a thread-pool and execute the workers in the thread-pool. Multiple threads can solve the problem of workers sleeping on the thread, as in our example. But remember that the default implementation of python, the CPython, has a global interpreter lock (GIL) which prevents more than one python statement executing at the same time. Will `parallel_pipe` help in the presence of GIL, besides the case of workers just sleeping?
# It turns out that for the majority of serious cases, multiple threads help even in the presence of the GIL, because most of the heavy-lifting operations, for example file accesses, are implemented in C instead of in pure python, and in C it is possible to release the GIL when not interacting with the python runtime. In addition to file accesses, if you are doing number-crunching, then hopefully you are not doing it in pure python but instead relies on dedicated libraries like numpy, scipy, etc. All of these libraries release the GIL when it makes sense to do so. So using `parallel_pipe` is usually enough.
# What if you just have to do your CPU-intensive tasks in python? Well, `parallel_pipe` and `parallel_pipe_unordered` takes an argument called `mode`, which by default takes the value `'thread'`. If you change it to `'process'`, then a process-pool instead of a thread-pool will be used. Let's see a comparison:
# +
import asyncio
import time
import aiochan as ac
def worker(_):
total = 0
for i in range(1000000):
total += i
return total
async def main():
start = asyncio.get_event_loop().time()
await ac.from_range(20).parallel_pipe(10, worker).collect()
print('using threads', asyncio.get_event_loop().time() - start)
start = asyncio.get_event_loop().time()
await ac.from_range(20).parallel_pipe(10, worker, mode='process').collect()
print('using threads', asyncio.get_event_loop().time() - start)
ac.run(main())
# -
# Why not use a process pool in all cases? Processes have much greater overhead than threads, and also far more restrictions on their use. Crucially, you cannot share any object unless you do some dirty work yourself, and anything you pass to your worker, or return from your worker, must be picklable.
# In our example, our worker is a pure function. It is also possible to prepare some structures in each worker before-hand. In python 3.7 or above, there are the `initializer` and `init_args` arguments accepted by `parallel_pipe` and `parallel_pipe_unordered`, which will be passed to the construction to the pool executors to do the setup. Prior to python 3.7, such a setup is still possible with some hack: you can put the object to be set up in a `threading.local` object, and for *every* worker execution, check if the object exists, and if not, do the initialization:
# +
import asyncio
import time
import random
import threading
import aiochan as ac
worker_data = threading.local()
def worker(n):
try:
processor = worker_data.processor
except:
print('setting up processor')
worker_data.processor = lambda x: x*2
processor = worker_data.processor
return processor(n)
async def main():
start = asyncio.get_event_loop().time()
print(await ac.from_range(20).parallel_pipe(2, worker).collect())
ac.run(main())
# -
# Since we used two thread workers, the setup is done twice. This also works for `mode='process'`.
# What about parallelising work across the network? Or more exotic workflows? At its core, *aiochan* is a library that facilitates moving data around within the boundary of a single process on a single machine, but there is nothing preventing you using channels at the end-points of a network-based parallelism framework such as message queues or a framework like *dart*. Within its bounday, *aiochan* aims to give you maximum flexibility in developing concurrent workflows, and you should use *aiochan* it in tandem with some other suitable libraries or frameworks when you want to step out of its boundary.
# ## Back to the main thread
# Speaking of stepping out of boundaries, one case is exceedingly common: you use an aiochan-based workflow to prepare a stream of values, but you want to consume these values outside of the asyncio event loop. In this case, there are convenience methods for you:
# +
loop = asyncio.new_event_loop()
out = ac.Chan(loop=loop)
async def worker():
while True:
await asyncio.sleep(0.1)
if not (await out.put('work')):
break
ac.run_in_thread(worker(), loop=loop)
it = out.to_iterable(buffer_size=1)
print(next(it))
print(next(it))
loop.call_soon_threadsafe(out.close);
# -
# Notice how we constructed the channel on the main thread, with explicit arguments specifying on which loop the channel is to be used, and then derived a iterator from the queue. Also, to run the worker, we used `run_in_thread` with an explicit event loop given.
# When creating the iterable, notice we have given it a `buffer_size`. This is used to construct a queue for inter-thread communication. You can also use a queue directly:
# +
import queue
loop = asyncio.new_event_loop()
out = ac.Chan(loop=loop)
async def worker():
while True:
await asyncio.sleep(0.1)
if not (await out.put('work')):
break
ac.run_in_thread(worker(), loop=loop)
q = queue.Queue()
out.to_queue(q)
print(q.get())
print(q.get())
loop.call_soon_threadsafe(out.close);
# -
# Other queues can be used as long as they follow the public API of `queue.Queue` and are thread-safe.
# ## aiochan without asyncio
# Finally, before ending this tutorial, let's reveal a secret: you don't need asyncio to use aiochan! "Isn't aiochan based on asyncio?" Well, not really, the core algorithms of aiochan (which is based on those from Clojure's core.async) does not use any asyncio constructs: they run entirely synchronously. It is only when you use the use-facing methods such as `get`, `put` and `select` that an asyncio-facade was made to cover the internals.
# On the other hand, there are some functions (actually, three of them) that does not touch anything related to asyncio given the correct arguments:
#
# * `Chan.put_nowait`
# * `Chan.get_nowait`
# * `select`
# Normally, when you call `ch.put_nowait(v)`, the put will succeed if it is possible to do so immediately (for example, if there is a pending get or buffer can be used), otherwise it will give up. Note that you never `await` on `put_nowait`. However, if you give the argument `immediate_only=True`, then if the operation cannot be completed immediately, it will be queued (but again, the pending queue can overflow). In addition, you can give a callback to the `cb` argument, which will be called when the put finally succeeds, with the same argument as the return value of `await put(v)`. The same is true with `get_nowait(immediate_only=True, cb=cb)`. For `select`, if you give a callback to the `cb` argument, then you should not call `await` on it, but instead rely on the callback being called eventually as `cb(return_value, which_channel)`. Note if you don't expect to use any event loops, when constructing channels, you should explicitly pass in `loop='no_loop'`.
# Example: this is our asyncio-based fan-in, fan-out:
# +
import aiochan as ac
import asyncio
async def consumer(c, tag):
async for v in c:
print('%s received %s' % (tag, v))
async def producer(c, tag):
for i in range(5):
v = '%s-%s' % (tag, i)
print('%s produces %s' % (tag, v))
await c.put(v)
async def main():
c = ac.Chan()
for i in range(3):
ac.go(consumer(c, 'c' + str(i)))
for i in range(3):
ac.go(producer(c, 'p' + str(i)))
await asyncio.sleep(0.1)
ac.run(main())
# -
# By the appropriate use of callbacks, we can write avoid using `asyncio` completely:
# +
def consumer(c, tag):
def cb(v):
if v is not None:
print('%s received %s' % (tag, v))
consumer(c, tag)
c.get_nowait(immediate_only=False, cb=cb)
def producer(c, tag, i=0):
v = '%s-%s' % (tag, i)
def cb(ok):
if ok and i < 4:
print('%s produces %s' % (tag, v))
producer(c, tag, i+1)
c.put_nowait(v, immediate_only=False, cb=cb)
def main():
c = ac.Chan(loop='no_loop')
for i in range(3):
consumer(c, 'c' + str(i))
for i in range(3):
producer(c, 'p' + str(i))
main()
# -
# The end result is (almost) the same. An example with `select`:
# +
def select_run():
c = ac.Chan(1, loop='no_loop', name='c')
d = ac.Chan(1, loop='no_loop', name='d')
put_chan = None
def put_cb(v, c):
nonlocal put_chan
put_chan = c
ac.select((c, 1), (d, 2), cb=put_cb)
get_val = None
def get_cb(v, c):
nonlocal get_val
get_val = v
ac.select(c, d, cb=get_cb)
print('select put into %s, get value %s' % (put_chan, get_val))
def main():
for _ in range(10):
select_run()
main()
# -
# "But why?" Well, obviously writing callbacks is much harder than using asyncio. But who knows? Maybe you are writing some other, higher-level framework that can make use of the semantics of aiochan. The possibilities are endless! In particular, there are non-asyncio concurrency frameworks in python itself that utilizes the same coroutines, an example being `python-trio`. Since the core of aiochan does not rely on asyncio, porting it to `trio` is trivial.
| doc/parallel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mask R-CNN Demo
#
# A quick intro to using the pre-trained model to detect and segment objects.
# +
import os
import sys
import random
import math
import numpy as np
import skimage.io
import matplotlib
import matplotlib.pyplot as plt
import cv2
# Root directory of the project
ROOT_DIR = os.path.abspath("../")
# Import Mask RCNN
sys.path.append(ROOT_DIR) # To find local version of the library
from mrcnn import utils
import mrcnn.model as modellib
from mrcnn import visualize
# Import COCO config
sys.path.append(os.path.join(ROOT_DIR, "samples/coco/")) # To find local version
import coco
from balloon import balloon
# %matplotlib inline
# Directory to save logs and trained model
MODEL_DIR = os.path.join(ROOT_DIR, "logs")
# Local path to trained weights file
COCO_MODEL_PATH = os.path.join(ROOT_DIR, "30_10_mask_rcnn_balloon_0017.h5")
# Download COCO trained weights from Releases if needed
if not os.path.exists(COCO_MODEL_PATH):
utils.download_trained_weights(COCO_MODEL_PATH)
# Directory of images to run detection on
IMAGE_DIR = os.path.join(ROOT_DIR, "images")
# -
# !pip3 install imgaug
# ## Configurations
#
# We'll be using a model trained on the MS-COCO dataset. The configurations of this model are in the ```CocoConfig``` class in ```coco.py```.
#
# For inferencing, modify the configurations a bit to fit the task. To do so, sub-class the ```CocoConfig``` class and override the attributes you need to change.
# +
config = balloon.BalloonConfig()
class InferenceConfig(config.__class__):
# class InferenceConfig(coco.CocoConfig):
# Set batch size to 1 since we'll be running inference on
# one image at a time. Batch size = GPU_COUNT * IMAGES_PER_GPU
GPU_COUNT = 1
IMAGES_PER_GPU = 1
config = InferenceConfig()
config.display()
# -
# ## Create Model and Load Trained Weights
# +
# Create model object in inference mode.
model = modellib.MaskRCNN(mode="inference", model_dir=MODEL_DIR, config=config)
# Load weights trained on MS-COCO
model.load_weights(COCO_MODEL_PATH, by_name=True)
# -
# ## Class Names
#
# The model classifies objects and returns class IDs, which are integer value that identify each class. Some datasets assign integer values to their classes and some don't. For example, in the MS-COCO dataset, the 'person' class is 1 and 'teddy bear' is 88. The IDs are often sequential, but not always. The COCO dataset, for example, has classes associated with class IDs 70 and 72, but not 71.
#
# To improve consistency, and to support training on data from multiple sources at the same time, our ```Dataset``` class assigns it's own sequential integer IDs to each class. For example, if you load the COCO dataset using our ```Dataset``` class, the 'person' class would get class ID = 1 (just like COCO) and the 'teddy bear' class is 78 (different from COCO). Keep that in mind when mapping class IDs to class names.
#
# To get the list of class names, you'd load the dataset and then use the ```class_names``` property like this.
# ```
# # Load COCO dataset
# dataset = coco.CocoDataset()
# dataset.load_coco(COCO_DIR, "train")
# dataset.prepare()
#
# # Print class names
# print(dataset.class_names)
# ```
#
# We don't want to require you to download the COCO dataset just to run this demo, so we're including the list of class names below. The index of the class name in the list represent its ID (first class is 0, second is 1, third is 2, ...etc.)
# COCO Class names
# Index of the class in the list is its ID. For example, to get ID of
# the teddy bear class, use: class_names.index('teddy bear')
# class_names = ['BG', 'person', 'bicycle', 'car', 'motorcycle', 'airplane',
# 'bus', 'train', 'truck', 'boat', 'traffic light',
# 'fire hydrant', 'stop sign', 'parking meter', 'bench', 'bird',
# 'cat', 'dog', 'horse', 'sheep', 'cow', 'elephant', 'bear',
# 'zebra', 'giraffe', 'backpack', 'umbrella', 'handbag', 'tie',
# 'suitcase', 'frisbee', 'skis', 'snowboard', 'sports ball',
# 'kite', 'baseball bat', 'baseball glove', 'skateboard',
# 'surfboard', 'tennis racket', 'bottle', 'wine glass', 'cup',
# 'fork', 'knife', 'spoon', 'bowl', 'banana', 'apple',
# 'sandwich', 'orange', 'broccoli', 'carrot', 'hot dog', 'pizza',
# 'donut', 'cake', 'chair', 'couch', 'potted plant', 'bed',
# 'dining table', 'toilet', 'tv', 'laptop', 'mouse', 'remote',
# 'keyboard', 'cell phone', 'microwave', 'oven', 'toaster',
# 'sink', 'refrigerator', 'book', 'clock', 'vase', 'scissors',
# 'teddy bear', 'hair drier', 'toothbrush']
class_names = ['BG', 'cmnd-cancuoc']
import dlib
def rotate_image(image):
num_faces = 0
correct_image = image.copy()
correct_orientation = -1
detector = dlib.get_frontal_face_detector()
for i in range (0,4):
rotated_image = np.rot90(image, k = i, axes=(1, 0))
bbs = detector(rotated_image, 0)
if(len(bbs) > num_faces):
num_faces = len(bbs)
correct_orientation = i*90
correct_image = rotated_image.copy()
return correct_orientation, correct_image
IMAGE_DIR = '/home/hoanviettran/cmnd-cancuoc/val'
# ## Run Object Detection
# +
# Load a random image from the images folder
file_names = next(os.walk(IMAGE_DIR))[2]
# image = skimage.io.imread(os.path.join(IMAGE_DIR, random.choice(file_names)))
image = skimage.io.imread(os.path.join(IMAGE_DIR, '02a426bb-bb8a-42b3-8331-952f47e6205720170815165056_151650512.jpg'))
# rotate to correct orientation
rorated_image = rotate_image(image)[1]
# Run detection
results = model.detect([rorated_image], verbose=1)
# Visualize results
r = results[0]
visualize.display_instances(rorated_image, r['rois'], r['masks'], r['class_ids'],
class_names, r['scores'])
# -
results[0]['masks'].shape
img = cv2.imread(os.path.join(IMAGE_DIR, '2cef45a4-e35c-4c18-a62b-bc67c782b91920180718171808_181718273.jpg'))
binary_img = np.array(results[0]['masks'])*255
b = binary_img.astype(np.dtype('uint8'))
image, contours, hierarchy = cv2.findContours(b, cv2.RETR_LIST, cv2.CHAIN_APPROX_SIMPLE)
# contours = sorted(contours, key = cv2.contourArea, reverse = True)[:5]
rects = []
for c in contours:
# approximate the contour
hull = cv2.convexHull(c)
peri = cv2.arcLength(hull, True)
approx = cv2.approxPolyDP(hull, 0.1 * peri, True)
if(len(approx) == 4 and cv2.isContourConvex(approx)):
rects.append(approx)
detect = cv2.drawContours(img.copy(), rects,-1,(0,255,0),3)
cv2.namedWindow('image',cv2.WINDOW_NORMAL)
cv2.resizeWindow('image', 600,600)
cv2.imshow('image',detect)
cv2.waitKey()
cv2.destroyAllWindows()
img.shape
| samples/demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Feature: PageRank on Question Co-Occurrence Graph
# This is a "magic" (leaky) feature that exploits the patterns in question co-occurrence graph (based on the [kernel by @zfturbo](https://www.kaggle.com/zfturbo/pagerank-on-quora-feature-file-generator)).
# ## Imports
# This utility package imports `numpy`, `pandas`, `matplotlib` and a helper `kg` module into the root namespace.
from pygoose import *
import hashlib
# ## Config
# Automatically discover the paths to various data folders and compose the project structure.
project = kg.Project.discover()
# Identifier for storing these features on disk and referring to them later.
feature_list_id = 'magic_pagerank'
# ## Load Data
# Original question datasets.
df_train = pd.read_csv(project.data_dir + 'train.csv').fillna('')
df_test = pd.read_csv(project.data_dir + 'test.csv').fillna('')
# ## Build features
# Generate a graph of questions and their neighbors.
def generate_qid_graph_table(row):
hash_key1 = hashlib.md5(row['question1'].encode('utf-8')).hexdigest()
hash_key2 = hashlib.md5(row['question2'].encode('utf-8')).hexdigest()
qid_graph.setdefault(hash_key1, []).append(hash_key2)
qid_graph.setdefault(hash_key2, []).append(hash_key1)
qid_graph = {}
_ = df_train.apply(generate_qid_graph_table, axis=1)
_ = df_test.apply(generate_qid_graph_table, axis=1)
# Compute PageRank.
def pagerank():
MAX_ITER = 20
d = 0.85
# Initializing: every node gets a uniform value!
pagerank_dict = {i: 1 / len(qid_graph) for i in qid_graph}
num_nodes = len(pagerank_dict)
for iter in range(0, MAX_ITER):
for node in qid_graph:
local_pr = 0
for neighbor in qid_graph[node]:
local_pr += pagerank_dict[neighbor] / len(qid_graph[neighbor])
pagerank_dict[node] = (1 - d) / num_nodes + d * local_pr
return pagerank_dict
pagerank_dict = pagerank()
# ### Extract final features
def get_pagerank_value(pair):
q1 = hashlib.md5(pair[0].encode('utf-8')).hexdigest()
q2 = hashlib.md5(pair[1].encode('utf-8')).hexdigest()
return [pagerank_dict[q1], pagerank_dict[q2]]
pagerank_train = kg.jobs.map_batch_parallel(
df_train[['question1', 'question2']].as_matrix(),
item_mapper = get_pagerank_value,
batch_size=1000,
)
pagerank_test = kg.jobs.map_batch_parallel(
df_test[['question1', 'question2']].as_matrix(),
item_mapper = get_pagerank_value,
batch_size=1000,
)
X_train = np.array(pagerank_train) * 1000
X_test = np.array(pagerank_test) * 1000
print('X train:', X_train.shape)
print('X test: ', X_test.shape)
# ## Save features
feature_names = [
'pagerank_q1',
'pagerank_q2',
]
project.save_features(X_train, X_test, feature_names, feature_list_id)
| notebooks/feature-magic-pagerank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PHD
# language: python
# name: phd
# ---
# # Rigid Waist Shift Implementation
# ---
# +
import matplotlib.pyplot as plt
import seaborn as sns
from cpymad.madx import Madx
from pyhdtoolkit.cpymadtools import latwiss, matching, orbit, special
from pyhdtoolkit.utils import defaults
defaults.config_logger()
plt.style.use("phd") # pyhdtoolkit.utils.defaults.install_mpl_style()
# %matplotlib inline
# %config InlineBackend.figure_format = "retina"
sns.set_palette("pastel")
# -
# ### No Rigid Waist Shift
with Madx(stdout=False) as madx:
madx.option(echo=False, warn=False)
madx.call("lhc/lhc_as-built.seq")
madx.call("lhc/opticsfile.22")
special.re_cycle_sequence(madx, sequence="lhcb1", start="IP3")
orbit_scheme = orbit.setup_lhc_orbit(madx, scheme="flat")
special.make_lhc_beams(madx)
madx.command.use(sequence="lhcb1")
matching.match_tunes_and_chromaticities(madx, "lhc", "lhcb1", 62.31, 60.32, 2.0, 2.0)
twiss_df = madx.table.twiss.dframe().copy()
twiss_df.name = [element[:-2] for element in twiss_df.name]
ip1s = twiss_df.s["ip1"]
IR1_fig = latwiss.plot_latwiss(
madx,
figsize=(16, 11),
title="LHCB1 IR1 - No Rigid Waist Shift",
disp_ylim=(-0.5, 2.5),
xlimits=(ip1s - 457, ip1s + 457),
k0l_lim=(-1.3e-2, 1.3e-2),
k1l_lim=(-6.1e-2, 6.1e-2),
lw=1.5,
)
plt.vlines(x=ip1s, ymin=-100, ymax=10_000, color="grey", ls="--", lw=1.5, label="IP1")
# plt.savefig("plots/flat_lhcb1_ir1.pdf", dpi=1200)
# ### With Rigid Waist Shift
with Madx(stdout=False) as madx:
madx.option(echo=False, warn=False)
madx.call("lhc/lhc_as-built.seq")
madx.call("lhc/opticsfile.22")
special.re_cycle_sequence(madx, sequence="lhcb1", start="IP3")
orbit_scheme = orbit.setup_lhc_orbit(madx, scheme="flat")
special.make_lhc_beams(madx)
madx.command.use(sequence="lhcb1")
matching.match_tunes_and_chromaticities(madx, "lhc", "lhcb1", 62.31, 60.32, 2.0, 2.0)
special.apply_lhc_rigidity_waist_shift_knob(madx, rigidty_waist_shift_value=1, ir=1)
matching.match_tunes_and_chromaticities(madx, "lhc", "lhcb1", 62.31, 60.32, 2.0, 2.0)
twiss_df = madx.table.twiss.dframe().copy()
twiss_df.name = [element[:-2] for element in twiss_df.name]
ip1s = twiss_df.s["ip1"]
IR1_waist_shift = latwiss.plot_latwiss(
madx,
figsize=(16, 11),
title="LHCB1 IR1 - Applied Rigid Waist Shift",
disp_ylim=(-0.5, 2.5),
xlimits=(ip1s - 457, ip1s + 457),
k0l_lim=(-1.3e-2, 1.3e-2),
k1l_lim=(-6.1e-2, 6.1e-2),
lw=1.5,
)
plt.vlines(x=ip1s, ymin=-100, ymax=10_000, color="grey", ls="--", lw=1.5, label="IP1")
# plt.savefig("plots/flat_lhcb1_ir1.pdf", dpi=1200)
# ---
| rigid_waist_shift.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import csv
import os
csvpath = os.path.join('Kenny Britt_Trans 1.csv')
# Method 2: Improved Reading using CSV module
import csv
with open(csvpath, newline='') as csvfile:
# CSV reader specifies delimiter and variable that holds contents
csvreader = csv.reader(csvfile, delimiter=';')
print(csvreader)
# Each row is read as a row
for row in csvreader:
print(row)
# -
import pandas as pd
kennybritt = pd.read_csv('Kenny Britt_Trans 1.csv', sep=';')
kennybritt
| Twitter_Sentiment_Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from GHCND import *
# initStationsMetadata,initInventoryMetadata,initDlyFileDirectory
sp = preprocessor.StationPreprocessor("D:/GHCND_data/ghcnd-stations.txt",
"D:/GHCND_data/ghcnd-inventory.txt",
"D:/GHCND_data/ghcnd_all.tar/ghcnd_all/ghcnd_all")
# "united states", "canada"
sp.addCountries(["united states"])
# "minnesota","wisconsin","iowa","north dakota","south dakota","michigan","nebraska","illinois", "ontario", "manitoba", "saskatchewan"
sp.addStates(["wisconsin"])
sp.addStations()
sp.processDlyFiles(["TMAX","TMIN","PRCP","TAVG"])
stats.calculateMean(sp,"month")
#conversion.TenthsCelsiusToCelsius(sp)
#sp.exportToJSON("C:/Users/Jacob/Projects/GHCND/website/app/data/ghcnd_ontario_data.json")
# -
plotter.plotStationSeries(sp.stations[0],"PRCP")
plotter.plotOutFile("D:/Shared_VM_Folder/bk149prcp")
sp.stations[0].variables["TMAX"].timelist[-1]
sp.writeToDat("./dat_files")
sp.exportToShapefile("D:/GHCND_data/canada_mid.shp")
| preprocessing_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
np.random.seed(2020)
# +
def generate_noise_1d(power, smoothness=0.6):
size = 2 ** power
scale = 1
noise = np.random.rand(size)
noise_res = noise.copy()
while size > 1:
size = size // 2
scale = scale * 2
noise_mean = np.repeat(np.mean(noise.reshape(size, scale), axis=1), scale)
noise_mean_conv = np.roll(noise_mean, -scale)
noise_linear = np.tile(np.linspace(0.0, 1.0, scale, endpoint=False), size)
noise_added = (1 - noise_linear) * noise_mean + noise_linear * noise_mean_conv
noise_res = (1 - smoothness) * noise_res + smoothness * noise_added
noise_min = np.min(noise_res)
noise_max = np.max(noise_res)
return (noise_res - noise_min) / (noise_max - noise_min)
def generate_noise_2d(power, smoothness=0.6):
size = 2 ** power
scale = 1
noise = np.random.rand(size, size)
noise_res = noise.copy()
while size % 2 == 0:
size = size // 2
scale = scale * 2
noise_mean = np.repeat(np.repeat(np.mean(noise.reshape(size, size, scale, scale), axis=(2, 3)), scale, axis=0), scale, axis=1)
noise_mean_convx = np.roll(noise_mean, (-scale, 0), axis=(0, 1))
noise_mean_convy = np.roll(noise_mean, (0, -scale), axis=(0, 1))
noise_mean_convxy = np.roll(noise_mean, (-scale, -scale), axis=(0, 1))
noise_linear = np.linspace(0.0, 1.0, scale, endpoint=False)
noise_linear_x = np.tile(noise_linear[:, np.newaxis], (size, 2 ** power))
noise_linear_y = np.tile(noise_linear[np.newaxis, :], (2 ** power, size))
noise_added = (
(1 - noise_linear_x) * (1 - noise_linear_y) * noise_mean +
noise_linear_x * (1 - noise_linear_y) * noise_mean_convx +
(1 - noise_linear_x) * noise_linear_y * noise_mean_convy +
noise_linear_x * noise_linear_y * noise_mean_convxy)
noise_res = (1 - smoothness) * noise_res + smoothness * noise_added
noise_min = np.min(noise_res)
noise_max = np.max(noise_res)
return (noise_res - noise_min) / (noise_max - noise_min)
# +
def is_inbounds(hmap, x, y):
return (
x >= 0.0 and
y >= 0.0 and
x < hmap.shape[0] - 1.0 and
y < hmap.shape[1] - 1.0
)
def value_at(hmap, x, y):
if is_inbounds(hmap, x, y):
xc = int(x)
yc = int(y)
u = x % 1.0
v = y % 1.0
ui = 1.0 - u
vi = 1.0 - v
return (
ui * vi * hmap[xc, yc] +
u * vi * hmap[xc+1, yc] +
ui * v * hmap[xc, yc+1] +
u * v * hmap[xc+1, yc+1]
)
else:
return 0
def grad_at(hmap, x, y):
if is_inbounds(hmap, x, y):
xc = int(x)
yc = int(y)
u = x % 1.0
v = y % 1.0
z = hmap[xc, yc]
zx = hmap[xc+1, yc]
zy = hmap[xc, yc+1]
zxy = hmap[xc+1, yc+1]
return np.array([
(zx - z) + v * (z - zx + zxy - zy),
(zy - z) + u * (z - zy + zxy - zx)
])
else:
return 0
def iter_grad(hmap, x, y, n_iter=512, rate=1.0):
n = 0
pos = []
while is_inbounds(hmap, x, y) and n < n_iter:
z = value_at(hmap, x, y)
g = grad_at(hmap, x, y)
pos.append((x, y, z))
x = x - rate * g[0]
y = y - rate * g[1]
n += 1
return np.array(pos)
# -
x2d = generate_noise_2d(8, smoothness=0.85)
# +
# %matplotlib inline
plt.figure(figsize=(12, 12))
plt.imshow(x2d, cmap='terrain')
for k in range(16):
xr = x2d.shape[0] * np.random.rand()
yr = x2d.shape[1] * np.random.rand()
p = iter_grad(x2d, xr, yr, n_iter=2**16, rate=8)
plt.plot([p[0, 0]], [p[0, 1]], 'or', ms=8)
plt.plot(p[:, 0], p[:, 1], '-r')
plt.show()
# -
plt.imshow(x2d[:128, :128], cmap='terrain')
plt.show()
# +
fig = plt.figure(figsize=(12, 12))
ax = fig.gca(projection='3d')
t = np.arange(x2d.shape[0])
xx, yy = np.meshgrid(t, t)
ax.plot([p[0, 0]], [p[0, 1]], [p[0, 2]], 'or', ms=8, zorder=10)
ax.plot(p[:, 0], p[:, 1], p[:, 2], '-r', zorder=10)
ax.plot_surface(xx, yy, x2d, cmap='terrain')
plt.show()
# -
| ipynb/basic-erosion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import pandas as pd
from scipy.stats import norm
# # In-Class Exercise for March 6 - The Brightest Stars
#
# The dataset we'll be dealing with here is a list of properties for the 50 brightest stars in the sky. You will mostly be running code that has already been written, but take the time to discuss with your partner what the code is "doing" at each step. Experiment with changing values to make sure you understand what is going on. You will submit your notebook for participation credit.
#
# ### ***Add at least one comment to each code cell describing what it's doing.***
#
# ### ***Enter all qualitative (non-code) answers below the questions themselves in the same markdown cell***
data = pd.read_csv('ICA_030619.csv')
data = data.replace("?",np.nan)
data = data.replace("binary",np.nan)
data
# ## Exercise 1
# Using the two cells below as models, make histograms for each of the quantitative columns in the table, and describe the nature of each distribution
data.hist('Visual Mag.')
data["Mass (Sol)"]=data["Mass (Sol)"].astype('float')
data.hist('Mass (Sol)')
# ## Exercise 2
# Study the function and the output of the test statements below, then complete the following:
#
# 1) Add comments to the key lines in the function (read docstrings for unknown functions to find out what they're doing)
# 2) Describe what each plot created by the test statements I've provided is showing. Add axis labels to both plots.
# 2) Add at least 5 additional test statements with different values for the input parameters (nstars, nsamples, colname). Then, describe in words the effect of changing each one. Pay particular attention to the differences between (a) the "true mean" and the "mean of the sampling distribution" and (b) the normal curve and the histogram
# 3) What statistical principle(s) is at play here?
def sample_stars(colname, nsamples=10, nstars=10, showall=False):
"""
A function that assembles mean values for nsamples randomly drawn samples of size nstars from the dataframe data.
"""
avg_prop = []
confints = []
dummyy=0
for i in np.arange(nsamples):
sample = np.random.choice(data[colname], nstars)
avg_prop.append(np.nanmean(sample))
stderr = np.nanstd(sample)/np.sqrt(nstars)
conf95 = 1.96*stderr
confints.append(conf95)
dummyy +=1
if showall==True:
plt.errorbar(np.nanmean(sample),dummyy, xerr=conf95,fmt='ko',ecolor='gray')
if showall==True:
plt.plot([np.nanmean(data[colname]),np.nanmean(data[colname])],[0,nsamples+1],'r--', label = "True Mean")
plt.plot([np.mean(avg_prop),np.mean(avg_prop)],[0,nsamples+1],'g--', label = "Mean of Sampling Dist.")
plt.legend()
return(avg_prop, confints)
avg_masses, confints = sample_stars("Mass (Sol)", nsamples=50, nstars=10, showall=True)
plt.hist(avg_masses, normed=True)
print("mean is ", np.mean(avg_masses))
print("standard deviation is", np.std(avg_masses))
plt.plot(np.arange(0,15,0.1),norm.pdf(np.arange(0,15,0.1),np.mean(avg_masses),np.std(avg_masses)))
# ## Exercise 3
# A list of the absolute magnitudes (a measure of intrinsic brightness) of the ten nearest stars to the earth is given below.
# 1) Compute the average and the standard deviation
# 2) Quantitatively evaluate the following hypothesis and describe your reasoning at each step: "the mean absolute magnitude of the nearest stars is the same as that of the brightest stars"
nearest_mags=[15.53,4.38,5.71,13.22,14.2,16.55,10.44,1.42,11.34,15.4]
| ICA_030619.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="1be778dc82c6e2aaa57f608486d368964887864d"
# %matplotlib inline
import numpy as np
import pandas as pd
from tqdm import tqdm_notebook as tqdm
from sklearn.externals import joblib
import seaborn as sns
from sklearn.model_selection import train_test_split
# + _uuid="fabc325a90727f0c726370edf4e7f1b155905794"
X = pd.read_csv('./data/application_train.csv', nrows=None)
X.head()
# + [markdown] _uuid="f436511aa30a5d273fa07d9d0e6d3a1a0ce40d82"
# # Preprocessing
# ## Solution 3
# + [markdown] _uuid="315fbf0f678ef5ab3b6b00492db65df5ec05a211"
# [<NAME> (Competition Host): "Value 365243 denotes infinity in DAYS variables in the datasets, therefore you can consider them NA values. Also XNA/XAP denote NA values."](https://www.kaggle.com/c/home-credit-default-risk/discussion/57247)
# + _uuid="d33b6990ddc1627945b40c100ede69e5f6c6456a"
X['CODE_GENDER'].unique()
# + _uuid="98aad6620c140292793f4e94e93e566b06e32b69"
X.loc[X['DAYS_EMPLOYED'] > 0]['DAYS_EMPLOYED'].unique()
# + _uuid="8a1b9f29cc93b4090389ca4fc4b3d8bd502bf352"
sum(X['ORGANIZATION_TYPE'] == 'XNA')
# + _uuid="0eb5e12c45eaf6ed788a89c182c2a51adac200b0"
X['CODE_GENDER'].value_counts()
# + _uuid="589051c515d1f57fb3ab70dc7c8b950a5e6c8662"
X['CODE_GENDER'].replace('XNA',np.nan, inplace=True)
X['CODE_GENDER'].value_counts()
# -
X['NAME_EDUCATION_TYPE'].unique()
# X['OCCUPATION_TYPE'].unique()
# + [markdown] _uuid="21b095c555fa5c48c9d3a02583e13ab00383b4af"
# # Feature Engineering
# ## Solution 3
# ### Hand crafted features
# + _uuid="73463beff83ce084432d59a430e300a7debc0668"
X['annuity_income_percentage'] = X['AMT_ANNUITY'] / X['AMT_INCOME_TOTAL']
X['car_to_birth_ratio'] = X['OWN_CAR_AGE'] / X['DAYS_BIRTH']
X['car_to_employ_ratio'] = X['OWN_CAR_AGE'] / X['DAYS_EMPLOYED']
X['children_ratio'] = X['CNT_CHILDREN'] / X['CNT_FAM_MEMBERS']
X['credit_to_annuity_ratio'] = X['AMT_CREDIT'] / X['AMT_ANNUITY']
X['credit_to_goods_ratio'] = X['AMT_CREDIT'] / X['AMT_GOODS_PRICE']
X['credit_to_income_ratio'] = X['AMT_CREDIT'] / X['AMT_INCOME_TOTAL']
X['days_employed_percentage'] = X['DAYS_EMPLOYED'] / X['DAYS_BIRTH']
X['income_per_child'] = X['AMT_INCOME_TOTAL'] / (1 + X['CNT_CHILDREN'])
X['income_per_person'] = X['AMT_INCOME_TOTAL'] / X['CNT_FAM_MEMBERS']
X['payment_rate'] = X['AMT_ANNUITY'] / X['AMT_CREDIT']
X['phone_to_birth_ratio'] = X['DAYS_LAST_PHONE_CHANGE'] / X['DAYS_BIRTH']
# X['occupation_type'] = X['OCCUPATION_TYPE'].replace({'Laborers':0, 'Core staff':1, 'Accountants':2, 'Managers':3,
# 'Drivers':4, 'Sales staff':5, 'Cleaning staff':6, 'Cooking staff':7,
# 'Private service staff':8, 'Medicine staff':9, 'Security staff':10,
# 'High skill tech staff':11, 'Waiters/barmen staff':12,
# 'Low-skill Laborers':13, 'Realty agents':14, 'Secretaries':15, 'IT staff':16,
# 'HR staff':17})
# X['name_education_type'] = X['NAME_EDUCATION_TYPE'].replace({'Secondary / secondary special':0, 'Higher education':1,
# 'Incomplete higher':2, 'Lower secondary':3, 'Academic degree':4})
# + _uuid="91fa3c9e207d932895ce0d2357d80cc98080c5da"
# External sources
X['external_sources_weighted'] = X.EXT_SOURCE_1 * 2 + X.EXT_SOURCE_2 * 3 + X.EXT_SOURCE_3 * 4
for function_name in ['min', 'max', 'sum', 'mean', 'nanmedian']:
X['external_sources_{}'.format(function_name)] = eval('np.{}'.format(function_name))(
X[['EXT_SOURCE_1', 'EXT_SOURCE_2', 'EXT_SOURCE_3']], axis=1)
# + _uuid="b07c05926c7fe40a8561acf888b9fa08e6b7442c"
engineered_numerical_columns = ['annuity_income_percentage',
'car_to_birth_ratio',
'car_to_employ_ratio',
'children_ratio',
'credit_to_annuity_ratio',
'credit_to_goods_ratio',
'credit_to_income_ratio',
'days_employed_percentage',
'income_per_child',
'income_per_person',
'payment_rate',
'phone_to_birth_ratio',
'external_sources_weighted',
'external_sources_min',
'external_sources_max',
'external_sources_sum',
'external_sources_mean',
'external_sources_nanmedian',
'NAME_EDUCATION_TYPE',
'OCCUPATION_TYPE',
'EXT_SOURCE_1',
'EXT_SOURCE_2',
'EXT_SOURCE_3']
# + _uuid="6762524228be62d5db5bc07a11e7d3201fc57c16"
X.columns()
# X_eng = X[engineered_numerical_columns + ['TARGET']]
# X_eng = X_eng.get_dummies()
# X_eng_corr = abs(X_eng.corr())
# + _uuid="69ef5da6c33b819924396f0ac0a230e5350383e0"
X_eng_corr.sort_values('TARGET', ascending=False)['TARGET']
# + _uuid="0dc727d54ffdc1b8d2d2893ceffaa36bb0f6e3e9"
sns.heatmap(X_eng_corr,
xticklabels=X_eng_corr.columns,
yticklabels=X_eng_corr.columns)
# + [markdown] _uuid="9b47f126e6309fbdb5b31a47fae31cc428b56846"
# ### Aggregation features
# + _uuid="30146f26ab88dd4f8a9c412356566be82f4f6558"
AGGREGATION_RECIPIES = [
(['CODE_GENDER', 'NAME_EDUCATION_TYPE'], [('AMT_ANNUITY', 'max'),
('AMT_CREDIT', 'max'),
('EXT_SOURCE_1', 'mean'),
('EXT_SOURCE_2', 'mean'),
('OWN_CAR_AGE', 'max'),
('OWN_CAR_AGE', 'sum')]),
(['CODE_GENDER', 'ORGANIZATION_TYPE'], [('AMT_ANNUITY', 'mean'),
('AMT_INCOME_TOTAL', 'mean'),
('DAYS_REGISTRATION', 'mean'),
('EXT_SOURCE_1', 'mean')]),
(['CODE_GENDER', 'REG_CITY_NOT_WORK_CITY'], [('AMT_ANNUITY', 'mean'),
('CNT_CHILDREN', 'mean'),
('DAYS_ID_PUBLISH', 'mean')]),
(['CODE_GENDER', 'NAME_EDUCATION_TYPE', 'OCCUPATION_TYPE', 'REG_CITY_NOT_WORK_CITY'], [('EXT_SOURCE_1', 'mean'),
('EXT_SOURCE_2', 'mean')]),
(['NAME_EDUCATION_TYPE', 'OCCUPATION_TYPE'], [('AMT_CREDIT', 'mean'),
('AMT_REQ_CREDIT_BUREAU_YEAR', 'mean'),
('APARTMENTS_AVG', 'mean'),
('BASEMENTAREA_AVG', 'mean'),
('EXT_SOURCE_1', 'mean'),
('EXT_SOURCE_2', 'mean'),
('EXT_SOURCE_3', 'mean'),
('NONLIVINGAREA_AVG', 'mean'),
('OWN_CAR_AGE', 'mean'),
('YEARS_BUILD_AVG', 'mean')]),
(['NAME_EDUCATION_TYPE', 'OCCUPATION_TYPE', 'REG_CITY_NOT_WORK_CITY'], [('ELEVATORS_AVG', 'mean'),
('EXT_SOURCE_1', 'mean')]),
(['OCCUPATION_TYPE'], [('AMT_ANNUITY', 'mean'),
('CNT_CHILDREN', 'mean'),
('CNT_FAM_MEMBERS', 'mean'),
('DAYS_BIRTH', 'mean'),
('DAYS_EMPLOYED', 'mean'),
('DAYS_ID_PUBLISH', 'mean'),
('DAYS_REGISTRATION', 'mean'),
('EXT_SOURCE_1', 'mean'),
('EXT_SOURCE_2', 'mean'),
('EXT_SOURCE_3', 'mean')]),
]
# + _uuid="66dbb34a6215465bd086d1ce5eb987100d747ff0"
groupby_aggregate_names = []
for groupby_cols, specs in tqdm(AGGREGATION_RECIPIES):
group_object = X.groupby(groupby_cols)
for select, agg in tqdm(specs):
groupby_aggregate_name = '{}_{}_{}'.format('_'.join(groupby_cols), agg, select)
X = X.merge(group_object[select]
.agg(agg)
.reset_index()
.rename(index=str,
columns={select: groupby_aggregate_name})
[groupby_cols + [groupby_aggregate_name]],
on=groupby_cols,
how='left')
groupby_aggregate_names.append(groupby_aggregate_name)
# + _uuid="c31f24c16238e003b02168e935ec80d4cc9e7853"
X_agg = X[groupby_aggregate_names + ['TARGET']]
X_agg_corr = abs(X_agg.corr())
# + _uuid="5c8f10e04cb4131e4b65eece35b28b9f5d167c76"
X_agg_corr.sort_values('TARGET', ascending=False)['TARGET']
# + _uuid="64f7e1b02336b34a6ad004864b5888af10f8267f"
sns.heatmap(X_agg_corr,
xticklabels=X_agg_corr.columns,
yticklabels=X_agg_corr.columns)
# -
dsccb = pd.read_csv('./data/credit_card_balance.csv')
dsccb_g = dsccb.groupby('SK_ID_CURR').mean()
dsccb_g['AMT_BALANCE_LIMIT_RATIO'] = dsccb_g['AMT_BALANCE'] / dsccb_g['AMT_CREDIT_LIMIT_ACTUAL']
dsccb_g = dsccb_g.reset_index('SK_ID_CURR')
ccb_columns = ['AMT_BALANCE_LIMIT_RATIO','CNT_DRAWINGS_ATM_CURRENT','AMT_BALANCE','AMT_TOTAL_RECEIVABLE','AMT_RECIVABLE','AMT_RECEIVABLE_PRINCIPAL','MONTHS_BALANCE','AMT_DRAWINGS_ATM_CURRENT','AMT_DRAWINGS_CURRENT','CNT_DRAWINGS_POS_CURRENT']
# dsccb_g = dsccb_g[ccb_columns]
dsbb = pd.read_csv('./data/bureau.csv')
dsbb_g = dsbb.groupby('SK_ID_CURR').mean()
dsbb_g = dsbb_g.reset_index()
bb_columns = ['DAYS_CREDIT','CREDIT_DAY_OVERDUE','DAYS_CREDIT_ENDDATE','DAYS_ENDDATE_FACT','AMT_CREDIT_MAX_OVERDUE','CNT_CREDIT_PROLONG','AMT_CREDIT_SUM','AMT_CREDIT_SUM_DEBT','AMT_CREDIT_SUM_LIMIT','AMT_CREDIT_SUM_OVERDUE','DAYS_CREDIT_UPDATE']
# +
# X_final2 = X[engineered_numerical_columns + groupby_aggregate_names + ['TARGET']]
X_final2 = X[['SK_ID_CURR'] + engineered_numerical_columns + groupby_aggregate_names]
X_final2 = pd.merge(X_final2, dsbb_g, how='left', on=['SK_ID_CURR'])
X_final2 = X_final2[bb_columns + engineered_numerical_columns + groupby_aggregate_names]
X_final2 = X_final2.replace(-np.inf, np.nan)
# X_final2 = X_final2.fillna(X_final2.median())
X_final2 = pd.get_dummies(X_final2, columns=['NAME_EDUCATION_TYPE', 'OCCUPATION_TYPE'])
arr = X_final2.as_matrix()
Y2_ = X['TARGET']
# Y2_ = arr[:,-1:]
X2_ = X_final2
features = X_final2.columns
import xgboost as xgb
param_dist = {'objective':'binary:logistic',
'n_estimators': 600,
'scale_pos_weight': 10,
'eval_metric': 'auc',
'max_depth': 5,
'gamma': 0.1,
'nthread': 4,
'subsample': 0.9
}
clf = xgb.XGBModel(**param_dist)
# dm = xgb.DMatrix(X2_, label=Y2_, feature_names=features)
# bst = xgb.train(param_dist, dm, num_boost_round=50)
# scores = bst.get_fscore()
X_train, X_val, y_train, y_val = train_test_split(X2_, Y2_, test_size=0.3, random_state=1)
clf.fit(X_train, y_train,
eval_set=[(X_train, y_train), (X_val, y_val)],
eval_metric='auc',
verbose=True)
# clf.fit(X2_, Y2_,
# verbose=True)
# Load evals result by calling the evals_result() function
# evals_result = clf.evals_result()
# evals_result
# -
Y2_[Y2_==1].count()
# X_final2.columns
X_train_test = X[['SK_ID_CURR'] + engineered_numerical_columns + groupby_aggregate_names + ['TARGET']]
X_train_test2 = pd.merge(X_train_test, dsccb_g, how='left', on=['SK_ID_CURR'])
X_train_test2.count()
# scores = sorted(scores.items(), key=lambda x:x[1],reverse = True)
scores
# +
X_test = pd.read_csv('./data/application_test.csv', nrows=None)
X_test['CODE_GENDER'].replace('XNA', np.nan, inplace = True)
X_test['annuity_income_percentage'] = X_test['AMT_ANNUITY'] / X_test['AMT_INCOME_TOTAL']
X_test['car_to_birth_ratio'] = X_test['OWN_CAR_AGE'] / X_test['DAYS_BIRTH']
X_test['car_to_employ_ratio'] = X_test['OWN_CAR_AGE'] / X_test['DAYS_EMPLOYED']
X_test['children_ratio'] = X_test['CNT_CHILDREN'] / X_test['CNT_FAM_MEMBERS']
X_test['credit_to_annuity_ratio'] = X_test['AMT_CREDIT'] / X_test['AMT_ANNUITY']
X_test['credit_to_goods_ratio'] = X_test['AMT_CREDIT'] / X_test['AMT_GOODS_PRICE']
X_test['credit_to_income_ratio'] = X_test['AMT_CREDIT'] / X_test['AMT_INCOME_TOTAL']
X_test['days_employed_percentage'] = X_test['DAYS_EMPLOYED'] / X_test['DAYS_BIRTH']
X_test['income_credit_percentage'] = X_test['AMT_INCOME_TOTAL'] / X_test['AMT_CREDIT']
X_test['income_per_child'] = X_test['AMT_INCOME_TOTAL'] / (1 + X_test['CNT_CHILDREN'])
X_test['income_per_person'] = X_test['AMT_INCOME_TOTAL'] / X_test['CNT_FAM_MEMBERS']
X_test['payment_rate'] = X_test['AMT_ANNUITY'] / X_test['AMT_CREDIT']
X_test['phone_to_birth_ratio'] = X_test['DAYS_LAST_PHONE_CHANGE'] / X_test['DAYS_BIRTH']
X_test['phone_to_employ_ratio'] = X_test['DAYS_LAST_PHONE_CHANGE'] / X_test['DAYS_EMPLOYED']
X_test['occupation_type'] = X_test['OCCUPATION_TYPE'].replace({'Laborers':0, 'Core staff':1, 'Accountants':2, 'Managers':3,
'Drivers':4, 'Sales staff':5, 'Cleaning staff':6, 'Cooking staff':7,
'Private service staff':8, 'Medicine staff':9, 'Security staff':10,
'High skill tech staff':11, 'Waiters/barmen staff':12,
'Low-skill Laborers':13, 'Realty agents':14, 'Secretaries':15, 'IT staff':16,
'HR staff':17})
X_test['name_education_type'] = X_test['NAME_EDUCATION_TYPE'].replace({'Secondary / secondary special':0, 'Higher education':1,
'Incomplete higher':2, 'Lower secondary':3, 'Academic degree':4})
# External sources
X_test['external_sources_weighted'] = X_test.EXT_SOURCE_1 * 2 + X_test.EXT_SOURCE_2 * 3 + X_test.EXT_SOURCE_3 * 4
for function_name in ['min', 'max', 'sum', 'mean', 'nanmedian']:
X_test['external_sources_{}'.format(function_name)] = eval('np.{}'.format(function_name))(
X_test[['EXT_SOURCE_1', 'EXT_SOURCE_2', 'EXT_SOURCE_3']], axis=1)
groupby_aggregate_names = []
for groupby_cols, specs in tqdm(AGGREGATION_RECIPIES):
group_object = X_test.groupby(groupby_cols)
for select, agg in tqdm(specs):
groupby_aggregate_name = '{}_{}_{}'.format('_'.join(groupby_cols), agg, select)
X_test = X_test.merge(group_object[select]
.agg(agg)
.reset_index()
.rename(index=str,
columns={select: groupby_aggregate_name})
[groupby_cols + [groupby_aggregate_name]],
on=groupby_cols,
how='left')
groupby_aggregate_names.append(groupby_aggregate_name)
# X_test_final = X_test[engineered_numerical_columns + groupby_aggregate_names]
X_eng_test = X_test[['SK_ID_CURR'] + engineered_numerical_columns + groupby_aggregate_names]
X_test_final = X_eng_test
X_test_final = pd.merge(X_test_final, dsbb_g, how='left', on=['SK_ID_CURR'])
X_test_final = X_test_final[bb_columns + engineered_numerical_columns + groupby_aggregate_names]
X_test_final = X_test_final.replace(-np.inf, np.nan)
# X_test_final = X_test_final.fillna(X_test_final.median())
y_test = clf.predict(X_test_final.values)
X_raw = X_test.values
f = open("./result2.csv", "w")
result = 'SK_ID_CURR,TARGET\n'
for i in range(len(y_test)):
result += (str(X_raw[i][0]) + ',' + str(round(y_test[i])) + '\n')
f.write(result)
f.close()
print('finished')
# -
X_test_final.count()
# + [markdown] _uuid="1b95ed77157b03b1a901bd828e0221a3f0669098"
# ## Solution 4
# ### Hand crafted features
# * diff features
# + _uuid="f26b73aba8f8247feb059bd6036e9c880d2c9362"
diff_feature_names = []
for groupby_cols, specs in tqdm(AGGREGATION_RECIPIES):
for select, agg in tqdm(specs):
if agg in ['mean','median','max','min']:
groupby_aggregate_name = '{}_{}_{}'.format('_'.join(groupby_cols), agg, select)
diff_name = '{}_diff'.format(groupby_aggregate_name)
abs_diff_name = '{}_abs_diff'.format(groupby_aggregate_name)
X[diff_name] = X[select] - X[groupby_aggregate_name]
X[abs_diff_name] = np.abs(X[select] - X[groupby_aggregate_name])
diff_feature_names.append(diff_name)
diff_feature_names.append(abs_diff_name)
# + _uuid="c679a8f013da2fd5702f9d78afa8f678b137f179"
X_diff = X[diff_feature_names + ['TARGET']]
X_diff_corr = abs(X_diff.corr())
X_diff_corr.sort_values('TARGET', ascending=False)['TARGET']
# + [markdown] _uuid="77e0d4b4d2aeb397c681b55fed60e5009aea35aa"
# * unemployed
# + _uuid="ac608922e66f2412d32e23918ac207c1fd62e56d"
X['DAYS_EMPLOYED'].replace(365243,np.nan, inplace=True)
# + _uuid="78cf2d1aac56643cd22f6ffb0c15e722b9753f1e"
X_clean = X[~pd.isnull(X['DAYS_EMPLOYED'])]
# + _uuid="5391b001fce17a31018bf4eac19a4b6de78c3d97"
sns.distplot(X_clean['DAYS_EMPLOYED'])
# + _uuid="f211cc0c6758f2d52476a28c471e4dfc5bf92ae9"
X['DAYS_EMPLOYED'].describe()
# + _uuid="1613168567862ccf2424520620943d32f970debc"
X['long_employment'] = (X['DAYS_EMPLOYED'] > -2000).astype(int)
feature_names = ['long_employment']
# + _uuid="3de182485ea6ca83cec70165ddcad9c68c234a3c"
X_employment = X[feature_names +['DAYS_EMPLOYED'] + ['TARGET']]
X_employment_corr = abs(X_employment.corr())
X_employment_corr.sort_values('TARGET', ascending=False)['TARGET']
# + _uuid="0df5576e1130087059ebb884fea64e2cf1bfc13d"
sns.heatmap(X_employment_corr,
xticklabels=X_employment_corr.columns,
yticklabels=X_employment_corr.columns)
# + [markdown] _uuid="5de454dc737f188ca0745b685c51f3496edcaf11"
# * age binns
# + _uuid="3a34ccfb0b51bf1246729ec92139a0ed25c445d6"
sns.distplot(X['DAYS_BIRTH'])
# + _uuid="5f2b918fcd7b021bef4aaf93b63ee8df7f864c53"
X['retirement_age'] = (X['DAYS_BIRTH'] > -14000).astype(int)
feature_names = ['DAYS_BIRTH','retirement_age']
# + _uuid="5d77c7d0bb260ced7eeea08e7045b7255ae1b3a4"
X_age = X[feature_names + ['TARGET']]
X_age_corr = abs(X_age.corr())
X_age_corr.sort_values('TARGET', ascending=False)['TARGET']
# + _uuid="92ddd1b98e5204235cc6e398fb9bcf09cf7739b4"
sns.heatmap(X_age_corr,
xticklabels=X_age_corr.columns,
yticklabels=X_age_corr.columns)
# + _uuid="790ef1bd4cec40cb02e19015036711dba3ed6d49"
| home_credit_default_risk/public.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from numpy import array, exp, sqrt
import pandas as pd
from copy import deepcopy
from mkt_data import mkt_data as md
from scipy.stats import norm, normaltest, probplot
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import seaborn as sns
sns.set_context("poster")
sns.set(rc={'figure.figsize': (16, 9.)})
sns.set_style("whitegrid")
np.set_printoptions(precision=5)
% load_ext autoreload
% autoreload 2
# -
csv = 'csv/eurusd.csv'
index = pd.read_csv(csv, parse_dates=['Date'])
index['dlogS'] = index.Close.diff()/index.Close
index.dropna(inplace=True)
index.head(2)
# +
fig, axs = plt.subplots(4, figsize=(20,30))
axs[0].plot(index.Date, index.Close);
axs[0].set_title('Spot')
axs[1].plot(index.Date, index.dlogS);
axs[1].set_title('dLogS')
axs[2].hist(index.dlogS, bins=20, density=True, alpha=0.6, color='g');
mu, std = norm.fit(index.dlogS)
xmin, xmax = axs[2].get_xlim()
x = np.linspace(xmin, xmax, 100)
p = norm.pdf(x, mu, std)
axs[2].plot(x, p, 'k', linewidth=2);
axs[2].set_title('dlogS hist')
probplot(index.dlogS, dist="norm", plot=axs[3]);
axs[3].set_title('Q-Q plot')
k2, p = normaltest(index.dlogS)
alpha = 1e-3
if p < alpha: # null hypothesis: x comes from a normal distribution
print(f"The null hypothesis can be rejected p = {p}")
else:
print(f"The null hypothesis cannot be rejected p = {p}")
# +
indexold = index[index.Date < np.datetime64('2020-02-14')]
fig, axs = plt.subplots(4, figsize=(20, 30))
axs[0].plot(indexold.Date, indexold.Close);
axs[1].plot(indexold.Date, indexold.dlogS);
axs[2].hist(indexold.dlogS, bins=25, density=True, alpha=0.6, color='g');
mu, std = norm.fit(indexold.dlogS)
xmin, xmax = axs[2].get_xlim()
x = np.linspace(xmin, xmax, 100)
p = norm.pdf(x, mu, std)
axs[2].plot(x, p, 'k', linewidth=2);
probplot(indexold.dlogS, dist="norm", plot=axs[3]);
k2, p = normaltest(indexold.dlogS)
alpha = 1e-3
if p < alpha: # null hypothesis: x comes from a normal distribution
print(f"The null hypothesis can be rejected p = {p}")
else:
print(f"The null hypothesis cannot be rejected p = {p}")
# -
| notebooks/[underDev]Normal hyptohesis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Solving an electric circuit using Particle Swarm Optimization
# ## Introduction
#
# PSO can be utilized in a wide variety of fields. In this example, the problem consists of analysing a given electric circuit and finding the electric current that flows through it. To accomplish this, the ```pyswarms``` library will be used to solve a non-linear equation by restructuring it as an optimization problem. The circuit is composed by a source, a resistor and a diode, as shown below.
#
# 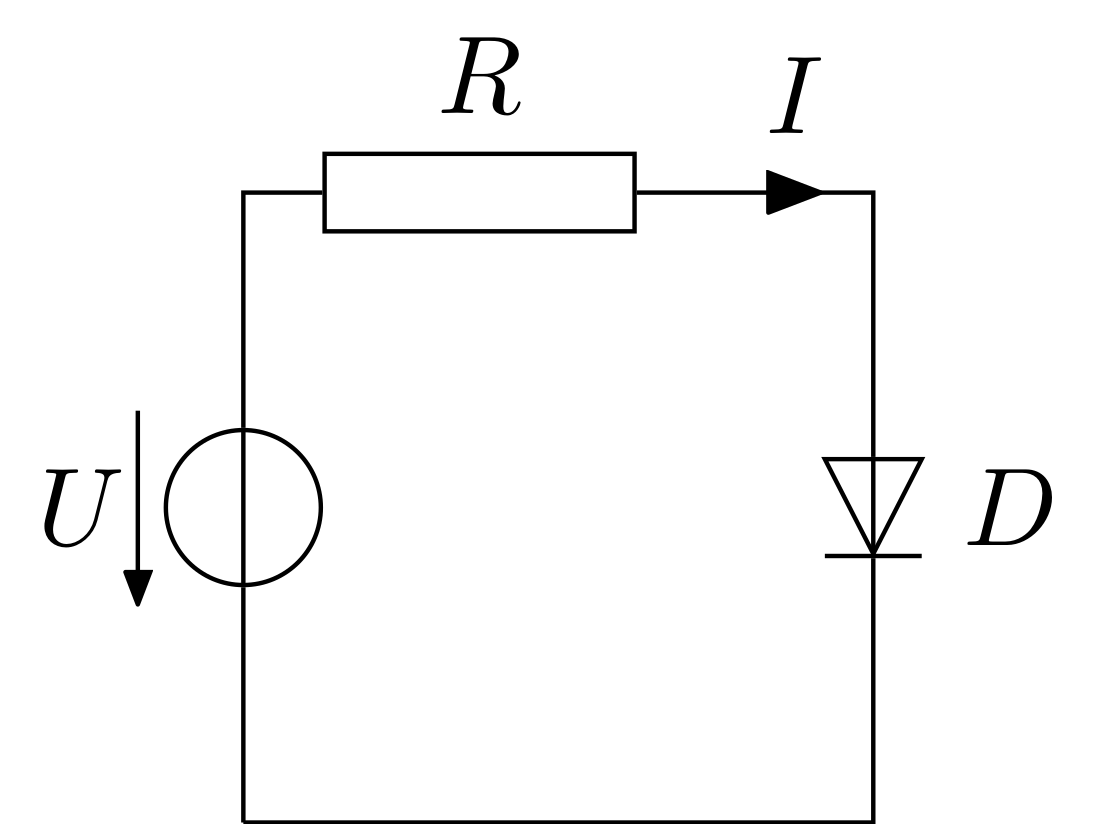
#
# ### Mathematical Formulation
#
# Kirchhoff's voltage law states that the directed sum of the voltages around any closed loop is zero. In other words, the sum of the voltages of the passive elements must be equal to the sum of the voltages of the active elements, as expressed by the following equation:
#
# $ U = v_D + v_R $, where $U$ represents the voltage of the source and, $v_D$ and $v_R$ represent the voltage of the diode and the resistor, respectively.
#
# To determine the current flowing through the circuit, $v_D$ and $v_R$ need to be defined as functions of $I$. A simplified Shockley equation will be used to formulate the current-voltage characteristic function of the diode. This function relates the current that flows through the diode with the voltage across it. Both $I_s$ and $v_T$ are known properties.
#
# $I = I_s e^{\frac{v_D}{v_T}}$
#
# Where:
#
# - $I$ : diode current
# - $I_s$ : reverse bias saturation current
# - $v_D$ : diode voltage
# - $v_T$ : thermal voltage
#
# Which can be formulated over $v_D$:
#
# $v_D = v_T \log{\left |\frac{I}{I_s}\right |}$
#
# The voltage over the resistor can be written as a function of the resistor's resistance $R$ and the current $I$:
#
# $v_R = R I$
#
# And by replacing these expressions on the Kirschhoff's voltage law equation, the following equation is obtained:
#
# $ U = v_T \log{\left |\frac{I}{I_s}\right |} + R I $
#
# To find the solution of the problem, the previous equation needs to be solved for $I$, which is the same as finding $I$ such that the cost function $c$ equals zero, as shown below. By doing this, solving for $I$ is restructured as a minimization problem. The absolute value is necessary because we don't want to obtain negative currents.
#
# $c = \left | U - v_T \log{\left | \frac{I}{I_s} \right |} - RI \right |$
#
#
# ### Known parameter values
#
# The voltage of the source is $ 10 \space V $ and the resistance of the resistor is $ 100 \space \Omega $. The diode is a silicon diode and it is assumed to be at room temperature.
#
# $U = 10 \space V $
#
# $R = 100 \space \Omega $
#
# $I_s = 9.4 \space pA = 9.4 \times 10^{-12} \space A$ (reverse bias saturation current of silicon diodes at room temperature, $T=300 \space K$)
#
# $v_T = 25.85 \space mV = 25.85 \times 10^{-3} \space V $ (thermal voltage at room temperature, $T=300 \space K$)
# ## Optimization
# +
# Import modules
import sys
import numpy as np
import matplotlib.pyplot as plt
# Import PySwarms
import pyswarms as ps
# -
print('Running on Python version: {}'.format(sys.version))
# ### Defining the cost fuction
#
# The first argument of the cost function is a ```numpy.ndarray```. Each dimension of this array represents an unknown variable. In this problem, the unknown variable is just $I$, thus the first argument is a unidimensional array. As default, the thermal voltage is assumed to be $25.85 \space mV$.
def cost_function(I):
#Fixed parameters
U = 10
R = 100
I_s = 9.4e-12
v_t = 25.85e-3
c = abs(U - v_t * np.log(abs(I[:, 0] / I_s)) - R * I[:, 0])
return c
# ### Setting the optimizer
#
# To solve this problem, the global-best optimizer is going to be used.
# +
# %%time
# Set-up hyperparameters
options = {'c1': 0.5, 'c2': 0.3, 'w':0.3}
# Call instance of PSO
optimizer = ps.single.GlobalBestPSO(n_particles=10, dimensions=1, options=options)
# Perform optimization
cost, pos = optimizer.optimize(cost_function, iters=30)
# -
print(pos[0])
print(cost)
# ### Checking the solution
#
# The current flowing through the circuit is approximately $ 0.094 \space A$ which yields a cost of almost zero. The graph below illustrates the relationship between the cost $c$ and the current $I$. As shown, the cost reaches its minimum value of zero when $I$ is somewhere close to $0.09$.
#
# The use of ```reshape(100, 1)``` is required since ```np.linspace(0.001, 0.1, 100)``` returns an array with shape ```(100,)``` and first argument of the cost function must be a unidimensional array, that is, an array with shape ```(100, 1)```.
# +
x = np.linspace(0.001, 0.1, 100).reshape(100, 1)
y = cost_function(x)
plt.plot(x, y)
plt.xlabel('Current I [A]')
plt.ylabel('Cost');
# -
# Another way of solving non-linear equations is by using non-linear solvers implemented in libraries such as ```scipy```. There are different solvers that one can choose which correspond to different numerical methods. We are going to use ```fsolve```, which is a general non-linear solver that finds the root of a given function.
#
# Unlike ```pyswarms```, the function (in this case, the cost function) to be used in ```fsolve``` must have as first argument a single value. Moreover, numerical methods need an initial guess for the solution, which can be made from the graph above.
# Import non-linear solver
from scipy.optimize import fsolve
# +
c = lambda I: abs(10 - 25.85e-3 * np.log(abs(I / 9.4e-12)) - 100 * I)
initial_guess = 0.09
current_I = fsolve(func=c, x0=initial_guess)
print(current_I[0])
# -
# The best solution value found using the PSO method was approximately the same as the one found using a non-linear solver, about $0.094 \space A$. In fact, the relative error was less than $1 \times 10^{-5}$.
| docs/examples/usecases/electric_circuit_problem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basic Usage of py_SBeLT
# ## Installation
# !pip install sbelt
# Once `sbelt` has been installed, we can import the `sbelt_runner` module. The sbelt_runner module is responsible for executing an instance of an py_SBeLT as it is described in py_SBeLT's [paper.md](https://github.com/szwiep/py_SBeLT/blob/master/paper/paper.md).
from sbelt import sbelt_runner
# ## Parameters and Running
# + [markdown] tags=[]
# The `sbelt_runner` module contains a function `run` which we will use to execute a run of py_SBeLT. `run` takes 13 parameters/arguments. The project's [API documentation](https://github.com/szwiep/py_SBeLT/tree/master/docs/API) describes these parameters but we also provide the table below:
#
#
# | Parameter | Type | Tested Range | Default | Description |
# | ----------- | ------- | ------ | ----------- | ----------- |
# | particle_pack_dens | float | 0.50, 0.80 | 0.78 | The packing fraction of the model particles (-) |
# | bed_length | int | 100, 1000 | 100 | Length of the domain in the streamwise direction (mm) |
# | particle_diam | int or float =+ 0.5 | 0.5, 10 | 0.5 | Grain diameter (mm) |
# | num_subregions | int | 1, 10 | 4 | The number of bed subregions |
# | level_limit | int | 1, 3 | 3 | The maximum number of levels permitted (i.e how many particles high to stack) |
# | iterations | int | 1, 1000000 | 1000 | The number of iterations to run |
# | poiss_lambda | int | 1, 5 | 5 | Lamba for poisson dist., used to determine the number of entrainment events |
# | gauss | boolean | - | False | Flag for which distribution to sample from for hop calculations. True=Normal, False=logNormal |
# | gauss_mu | float | 0.25, 2 | 1.0 | Mean/expectation of the logNormal/Normal distribution for hop calculations |
# | gauss_sigma | float | 0.25, 0.50 | 0.25 | Standard deviation of logNormal/Normal distribution for hop calculations|
# | data_save_interval | int | 1, 2 | 1 | How often to record model particle arrays (e.g 1=save every iteration, 2=save every other) |
# | height_dependant_entr | boolean | - | False | Flag indicating whether model automatically entrains particles that are at the height/level limit |
# | out_path | string | - | '.' | The location/directory to save model run data |
# | out_name | string | - | 'sbelt-out' | Filename for model run data |
#
#
# If we do not pass any arguments to `run` then the default parameters (described in the [project's DEFAULT_PARAMS.md](https://github.com/szwiep/py_SBeLT/blob/master/docs/DEFAULT_PARAMS.md)) will be used. Let's start by using the default parameters!
# -
# ### Running with Default Parameters
sbelt_runner.run()
# And that's all it takes! We can see that there is now a file `./sbelt-out.hdf5` located in our directory. This file contains information from the sbelt run including the placement of all particles in the stream for each iteration and metrics such as average age over all particles each iteration. For more detail regarding the information stored and how to interact with it, see the project documentation and additional notebooks.
#
#
#
# ### Running with User-defined Parameters
# But what if we don't want to use the default parameters but instead what to try our own? Let's try!
#
# Instead of using the default parameters, let's execute a run of sbelt over _3000 iterations_, with a _bed length of 150 mm_, _3 subregions_, and the number of entrainment events per-iteration being sample from a poissoin distribution parameterized by _$\lambda$ = 2_. Note that we will also need to create a new filename since we have already written to the default filename (`./sbelt-out.hdf5`) and run will not overwrite model files. Let's use `user-defined-sbelt`.
sbelt_runner.run(iterations=3000, bed_length=150, num_subregions=3, poiss_lambda=2, out_name='user-defined-sbelt')
# Now that we've got two files full of particle-related information, let's plot some of it!
# ## Plotting
# The `sbelt` package comes with some basic plotting logic. We can access these functions with the following import:
from sbelt.plots import plotting
# Each function in `plotting` will require information derived from the `sbelt.hdf5` files (as well a file names and save locations, if desired) which we created in the **Parameters and Running** section. We will need to import both `numpy` and `h5py` into our envrionment to allow us to open/handle the `sbelt.hdf5` files and their stored data structures.
import numpy as np
import h5py
# ### Plotting the Stream
# Let's start by plotting the stream at iteration 300 for both of our sbelt runs from **Parameters and Running** (default and user-defined parameters). Looking at the [API documentation](https://github.com/szwiep/py_SBeLT/blob/update_docs/docs/API/plotting.html), we can see that `plotting.stream` requires 5 arguments: `iteration`, `bed_particles`, `model_particles`, `x_lim`, and `y_lim`.
#
# For more information regarding the data in the HDF5 file, see the data_storage_sbelt notebook.
# +
iteration = 300
# y_lim is up to us and how we want the plot to look - not derived from hdf5 file!
y_lim = 10
with h5py.File('sbelt-out.hdf5', 'r') as f: # open the hdf5 file to read
# bed particles are stored in the initial_values group
default_bed_particles = np.array(f['initial_values']['bed'])
# model particles at the end of iteration i are stored with the key `iteration_i-1`
default_model_particles_300 = np.array(f['iteration_299']['model'])
# We want to plot the whole stream so let x_lim = length of the bed
default_x_lim = f['params']['bed_length'][()]
# -
plotting.stream(iteration, default_bed_particles, default_model_particles_300, default_x_lim, y_lim)
# +
# For user-defined (ud) run
with h5py.File('user-defined-sbelt.hdf5', 'r') as f: # open the hdf5 file to read
# bed particles are stored in the initial_values group
ud_bed_particles = np.array(f['initial_values']['bed'])
# model particles at the end of iteration i are stored with the key `iteration_i-1`
ud_model_particles_300 = np.array(f['iteration_299']['model'])
# We want to plot the whole stream so let x_lim = length of the bed
ud_x_lim = f['params']['bed_length'][()]
# -
plotting.stream(iteration, ud_bed_particles, ud_model_particles_300, ud_x_lim, y_lim)
# ### Plotting the Downstream Particle Crossings
# Next, let's plot a histogram and time-series of the particles crossings at the downstream boundary using the `downstream_boundary_hist` and `downstream_boundary_ts` functions, respectively. We will grab the required information (see [API documentation]()) from the `.hdf5` files similarly to how we did in **Plotting the Stream**.
# +
# For the default run (change filename to try another run)
with h5py.File('sbelt-out.hdf5', 'r') as f:
# Find how many subregions there are:
default_num_subregions = f['params']['num_subregions'][()]
# id of the downstream boundary (final subregion's right boundary) is `subregion_num_subregion-1`
downstream_key = default_num_subregions - 1 # (because subregions are named 0-(N-1))
# Get the crossings at the final subregion:
default_particle_crossing_list = np.array(f['final_metrics']['subregions'][f'subregion-{downstream_key}-flux'])
# Total number of iterations is stored in params group
default_iterations = f['params']['iterations'][()]
# -
plotting.downstream_boundary_hist(default_particle_crossing_list, default_iterations)
plotting.downstream_boundary_ts(default_particle_crossing_list, default_iterations, 1)
# ### Plotting the Downstream Particle Crossings with Age
# Finally, let's use the final plotting function provided by `sbelt`, `crossing_info_age`.
# +
# For the default run (change filename to try another run)
with h5py.File('sbelt-out.hdf5', 'r') as f:
# Find how many subregions there are:
default_num_subregions = f['params']['num_subregions'][()]
# id of the downstream boundary (final subregion's right boundary) is `subregion_num_subregion-1`
downstream_key = default_num_subregions - 1
# Get the crossings at the final subregion:
default_particle_crossing_list = np.array(f['final_metrics']['subregions'][f'subregion-{downstream_key}-flux'])
# Get average age and # of iterations
default_avg_age = np.array(f['final_metrics']['avg_age'])
default_iterations = f['params']['iterations'][()]
# -
plotting.crossing_info_age(default_particle_crossing_list, default_avg_age, default_iterations, 1)
| docs/notebooks/basic_usage_sbelt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import sys
sys.path.append('./')
from collections import Counter
from linear_algebra import distance
from stats import mean
import random
import matplotlib.pyplot as plt
from plot_state_borders import plot_state_borders
def raw_majority_vote(labels):
votes = Counter(labels)
winner, _ = votes.most_common(1)[0]
return winner
def majority_vote(labels):
"""assumes that labels are ordered from nearest to farthest"""
vote_counts = Counter(labels)
winner, winner_count = vote_counts.most_common(1)[0]
num_winners = len([count
for count in vote_counts.values()
if count == winner_count])
if num_winners == 1:
return winner # unique winner, so return it
else:
return majority_vote(labels[:-1]) # try again without the farthest
def knn_classify(k, labeled_points, new_point):
"""each labeled point should be a pair (point, label)"""
# order the labeled points from nearest to farthest
by_distance = sorted(labeled_points,
key=lambda point_label: distance(point_label[0], new_point))
# find the labels for the k closest
k_nearest_labels = [label for _, label in by_distance[:k]]
# and let them vote
return majority_vote(k_nearest_labels)
cities = [(-86.75, 33.5666666666667, 'Python'), (-88.25, 30.6833333333333, 'Python'), (-112.016666666667, 33.4333333333333, 'Java'), (-110.933333333333, 32.1166666666667, 'Java'), (-92.2333333333333, 34.7333333333333, 'R'), (-121.95, 37.7, 'R'), (-118.15, 33.8166666666667, 'Python'), (-118.233333333333, 34.05, 'Java'), (-122.316666666667, 37.8166666666667, 'R'), (-117.6, 34.05, 'Python'), (-116.533333333333, 33.8166666666667, 'Python'), (-121.5, 38.5166666666667, 'R'), (-117.166666666667, 32.7333333333333, 'R'), (-122.383333333333, 37.6166666666667, 'R'), (-121.933333333333, 37.3666666666667, 'R'), (-122.016666666667, 36.9833333333333, 'Python'), (-104.716666666667, 38.8166666666667, 'Python'), (-104.866666666667, 39.75, 'Python'), (-72.65, 41.7333333333333, 'R'), (-75.6, 39.6666666666667, 'Python'), (-77.0333333333333, 38.85, 'Python'), (-80.2666666666667, 25.8, 'Java'), (-81.3833333333333, 28.55, 'Java'), (-82.5333333333333, 27.9666666666667, 'Java'), (-84.4333333333333, 33.65, 'Python'), (-116.216666666667, 43.5666666666667, 'Python'), (-87.75, 41.7833333333333, 'Java'), (-86.2833333333333, 39.7333333333333, 'Java'), (-93.65, 41.5333333333333, 'Java'), (-97.4166666666667, 37.65, 'Java'), (-85.7333333333333, 38.1833333333333, 'Python'), (-90.25, 29.9833333333333, 'Java'), (-70.3166666666667, 43.65, 'R'), (-76.6666666666667, 39.1833333333333, 'R'), (-71.0333333333333, 42.3666666666667, 'R'), (-72.5333333333333, 42.2, 'R'), (-83.0166666666667, 42.4166666666667, 'Python'), (-84.6, 42.7833333333333, 'Python'), (-93.2166666666667, 44.8833333333333, 'Python'), (-90.0833333333333, 32.3166666666667, 'Java'), (-94.5833333333333, 39.1166666666667, 'Java'), (-90.3833333333333, 38.75, 'Python'), (-108.533333333333, 45.8, 'Python'), (-95.9, 41.3, 'Python'), (-115.166666666667, 36.0833333333333, 'Java'), (-71.4333333333333, 42.9333333333333, 'R'), (-74.1666666666667, 40.7, 'R'), (-106.616666666667, 35.05, 'Python'), (-78.7333333333333, 42.9333333333333, 'R'), (-73.9666666666667, 40.7833333333333, 'R'), (-80.9333333333333, 35.2166666666667, 'Python'), (-78.7833333333333, 35.8666666666667, 'Python'), (-100.75, 46.7666666666667, 'Java'), (-84.5166666666667, 39.15, 'Java'), (-81.85, 41.4, 'Java'), (-82.8833333333333, 40, 'Java'), (-97.6, 35.4, 'Python'), (-122.666666666667, 45.5333333333333, 'Python'), (-75.25, 39.8833333333333, 'Python'), (-80.2166666666667, 40.5, 'Python'), (-71.4333333333333, 41.7333333333333, 'R'), (-81.1166666666667, 33.95, 'R'), (-96.7333333333333, 43.5666666666667, 'Python'), (-90, 35.05, 'R'), (-86.6833333333333, 36.1166666666667, 'R'), (-97.7, 30.3, 'Python'), (-96.85, 32.85, 'Java'), (-95.35, 29.9666666666667, 'Java'), (-98.4666666666667, 29.5333333333333, 'Java'), (-111.966666666667, 40.7666666666667, 'Python'), (-73.15, 44.4666666666667, 'R'), (-77.3333333333333, 37.5, 'Python'), (-122.3, 47.5333333333333, 'Python'), (-89.3333333333333, 43.1333333333333, 'R'), (-104.816666666667, 41.15, 'Java')]
cities = [([longitude, latitude], language) for longitude, latitude, language in cities]
def plot_cities():
# key is language, value is pair (longitudes, latitudes)
plots = { "Java" : ([], []), "Python" : ([], []), "R" : ([], []) }
# we want each language to have a different marker and color
markers = { "Java" : "o", "Python" : "s", "R" : "^" }
colors = { "Java" : "r", "Python" : "b", "R" : "g" }
for (longitude, latitude), language in cities:
plots[language][0].append(longitude)
plots[language][1].append(latitude)
# create a scatter series for each language
for language, (x, y) in plots.items():
plt.scatter(x, y, color=colors[language], marker=markers[language],
label=language, zorder=10)
plot_state_borders() # assume we have a function that does this
plt.legend(loc=0) # let matplotlib choose the location
plt.axis([-130,-60,20,55]) # set the axes
plt.title("Favorite Programming Languages")
plt.show()
def classify_and_plot_grid(k=1):
plots = { "Java" : ([], []), "Python" : ([], []), "R" : ([], []) }
markers = { "Java" : "o", "Python" : "s", "R" : "^" }
colors = { "Java" : "r", "Python" : "b", "R" : "g" }
for longitude in range(-130, -60):
for latitude in range(20, 55):
predicted_language = knn_classify(k, cities, [longitude, latitude])
plots[predicted_language][0].append(longitude)
plots[predicted_language][1].append(latitude)
# create a scatter series for each language
for language, (x, y) in plots.items():
plt.scatter(x, y, color=colors[language], marker=markers[language],
label=language, zorder=0)
plot_state_borders(color='0') # assume we have a function that does this
plt.legend(loc=0) # let matplotlib choose the location
plt.axis([-130,-60,20,55]) # set the axes
plt.title(str(k) + "-Nearest Neighbor Programming Languages")
plt.show()
#
# the curse of dimensionality
#
def random_point(dim):
return [random.random() for _ in range(dim)]
def random_distances(dim, num_pairs):
return [distance(random_point(dim), random_point(dim))
for _ in range(num_pairs)]
if __name__ == "__main__":
# try several different values for k
for k in [1, 3, 5, 7]:
num_correct = 0
for location, actual_language in cities:
other_cities = [other_city
for other_city in cities
if other_city != (location, actual_language)]
predicted_language = knn_classify(k, other_cities, location)
if predicted_language == actual_language:
num_correct += 1
print(k, "neighbor[s]:", num_correct, "correct out of", len(cities))
dimensions = range(1, 101, 5)
avg_distances = []
min_distances = []
random.seed(0)
for dim in dimensions:
distances = random_distances(dim, 10000) # 10,000 random pairs
avg_distances.append(mean(distances)) # track the average
min_distances.append(min(distances)) # track the minimum
print(dim, min(distances), mean(distances), min(distances) / mean(distances))
classify_and_plot_grid()
plot_cities()
| first-edition/code-python3/NearestNeighbors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: parcels-container_2021.03.17-6c459b7
# language: python
# name: parcels-container_2021.03.17-6c459b7
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # OSNAP line Lagrangian particle tracking investigation of the cold/fresh blob
#
# -
# redo other sourcetime calculation because it was wrong first time round
# + [markdown] slideshow={"slide_type": "skip"}
# ## Technical preamble
# + slideshow={"slide_type": "skip"}
# import matplotlib.colors as colors
import numpy as np
from pathlib import Path
import matplotlib.pyplot as plt
import xarray as xr
from datetime import datetime, timedelta
import seaborn as sns
# from matplotlib.colors import ListedColormap
import cmocean as co
import pandas as pd
import matplotlib.dates as mdates
import cartopy.crs as ccrs
import cartopy
import seawater as sw
from matplotlib import colors as c
from matplotlib import ticker
# from xhistogram.xarray import histogram
# + slideshow={"slide_type": "skip"}
sns.set(style="darkgrid")
xr.set_options(keep_attrs=True)
np.warnings.filterwarnings('ignore')
sns.set_palette("colorblind")
xr.set_options(keep_attrs=True);
plt.rc('font', size=14) #controls default text size
plt.rc('axes', titlesize=14) #fontsize of the title
plt.rc('axes', labelsize=14) #fontsize of the x and y labels
plt.rc('xtick', labelsize=14) #fontsize of the x tick labels
plt.rc('ytick', labelsize=14) #fontsize of the y tick labels
plt.rc('legend', fontsize=14) #fontsize of the legend
plt.rc('savefig', dpi=300) # higher res outputs
# +
# from dask.distributed import Client
# +
# client = Client(n_workers=1, threads_per_worker=8, memory_limit=48e9)
# client
# + [markdown] slideshow={"slide_type": "skip"}
# _(Click on the link above if you want to see the Dask cluster in action.)_
# + [markdown] slideshow={"slide_type": "skip"}
# ## Set up paths and read in trajectory data
# + tags=["parameters"]
# parameters
project_path = Path.cwd() / '..' / '..'
project_path = project_path.resolve()
interim_data_path = Path('/data/spg_fresh_blob_202104_data/interim/')
outputPath = Path('data/interim/sumsAndMeans/')
output_data_path = project_path / outputPath
sectionPath = Path('data/external/')
sectionFilename = 'osnap_pos_wp.txt'
sectionname = 'osnap'
figure_path = Path('reports/figures/subsets/')
# output figures path
figures_path = Path("reports/figures/ColdBlob/")
# do this year-by-year because of filesizes
year = 1990
nsubsets = 32
# proportion of data in subset
subset = 1.0
# -
yearstr = str(year)
# + slideshow={"slide_type": "skip"}
# model mask file
data_path = Path("data/external/iAtlantic/")
experiment_name = "VIKING20X.L46-KKG36107B"
mesh_mask_file = project_path / data_path / "mask" / experiment_name / "1_mesh_mask.nc"
#section lonlat file
sectionPath = Path('data/external/')
sectionFilename = 'osnap_pos_wp.txt'
sectionname = 'osnap'
gsrsectionFilename = 'gsr_pos_wp.txt'
degree2km = 1.852*60.0
# some transport values specific to osnap runs
# randomly seeded 39995 particles, 19886 were in ocean points (the rest were land)
osnap_section_length = 3594572.87839 # m
osnap_subsection_length = 2375914.29783 # m
osnap_section_depth = 4000 # m over which particles launched
osnap_subsection_depth = 1000 # m over which particles launched
osnap_subsection_ocean_area = osnap_subsection_length * osnap_subsection_depth * 2100000 / 2643886
# this is to compensate for not using all the particles. 1 in 10 particles selected.
max_current = 2.0
particle_section_area = max_current * osnap_subsection_length * osnap_subsection_depth / (2643886 * subset)
# + [markdown] slideshow={"slide_type": "skip"}
# ## Load data
# + [markdown] slideshow={"slide_type": "skip"}
# ### mesh and masks
# + slideshow={"slide_type": "skip"}
mesh_mask = xr.open_dataset(mesh_mask_file)
mesh_mask = mesh_mask.squeeze()
mesh_mask = mesh_mask.set_coords(["nav_lon", "nav_lat", "nav_lev"])
bathy = mesh_mask.mbathy.rename("number of water filled points")
depth = (mesh_mask.e3t_0 * mesh_mask.tmask).sum("z")
# display(mesh_mask)
# + [markdown] slideshow={"slide_type": "skip"}
# ### section position data
# + slideshow={"slide_type": "skip"}
lonlat = xr.Dataset(pd.read_csv(project_path / sectionPath / sectionFilename,delim_whitespace=True))
# + slideshow={"slide_type": "skip"}
lonlat.lon.attrs['long_name']='Longitude'
lonlat.lat.attrs['long_name']='Latitude'
lonlat.lon.attrs['standard_name']='longitude'
lonlat.lat.attrs['standard_name']='latitude'
lonlat.lon.attrs['units']='degrees_east'
lonlat.lat.attrs['units']='degrees_north'
lonlat2mean= lonlat.rolling({'dim_0':2}).mean()
lonlatdiff = (lonlat.diff('dim_0'))
lonlatdiff = lonlatdiff.assign({'y':lonlatdiff['lat']*degree2km})
lonlatdiff = lonlatdiff.assign({'x':lonlatdiff['lon']*degree2km*np.cos(np.radians(lonlat2mean.lat.data[1:]))})
lonlatdiff=lonlatdiff.assign({'length':np.sqrt(lonlatdiff['x']**2+lonlatdiff['y']**2)})
lonlatdiff=lonlatdiff.assign({'costheta':lonlatdiff['x']/lonlatdiff['length']})
lonlatdiff=lonlatdiff.assign({'sintheta':lonlatdiff['y']/lonlatdiff['length']})
total_length = lonlatdiff.length.sum().data
total_osnap_length = lonlatdiff.length[0:12].sum().data; # exclude section across UK - just there for testing north/south
length_west = xr.concat((xr.DataArray([0],dims=("dim_0"),coords={"dim_0": [0]}),lonlatdiff.length.cumsum()),dim='dim_0')
# -
lonlat
# + [markdown] slideshow={"slide_type": "skip"}
# ### tracks
# + [markdown] slideshow={"slide_type": "skip"}
# ## Load VIKING20X data
#
# We'll first find all the relevant files and then open them as a virtual contiguous dataset.
# + slideshow={"slide_type": "skip"}
# data_stores_subsets = list(sorted(Path(data_path).glob("*_????_subset.zarr/")))[:use_number_subset_years]
data_trackends_subsets = list(sorted(Path(interim_data_path).glob(f"*{yearstr}*.nc/")))
# -
print(data_trackends_subsets)
# +
# ds = xr.concat(
# [xr.open_dataset(store,chunks={
# "ends": 1, "traj": 1024
# }) for store in data_trackends_subsets],
# dim="traj",
# )
ds = xr.concat(
[xr.open_dataset(store) for store in data_trackends_subsets],
dim="traj",
)
display(ds)
print(ds.nbytes / 1e9, "GiB")
# -
# ### 32 subsets, run separately
display(ds.time.isel(ends=1))
# #### Subset tracks by OSNAP line cross longitude and depth range
lonRange=[-37,0]
depthRange=[0,500]
range_str = 'OsnapE_test'
ds = ds.where((ds.isel(ends=0).lon > lonRange[0]) & (ds.isel(ends=0).lon < lonRange[1]))
ds = ds.where((ds.isel(ends=0).z > depthRange[0]) & (ds.isel(ends=0).z < depthRange[1]))
ds = ds.dropna('traj', how='all')
# #### Add density (sigma0) to variables
ds = ds.assign({'rho0':xr.apply_ufunc(
sw.dens,
ds.salt,ds.temp,0,
dask="parallelized",
output_dtypes=[float, ])})
ds.rho0.attrs = {'units':'kg/m3','long_name':'potential density $\rho_0$'}
# + [markdown] slideshow={"slide_type": "skip"}
# #### Velocity conversions from degrees lat/lon per second to m/s
# + slideshow={"slide_type": "skip"}
ds=ds.assign({'uvel_ms':ds.uvel * degree2km * 1000.0 * np.cos(np.radians(ds.lat))})
ds=ds.assign({'vvel_ms':ds.vvel * degree2km * 1000.0})
# -
ds = ds.assign({'section_index':(ds.isel(ends=0).lon > lonlat.lon).sum(dim='dim_0')-1})
costheta = lonlatdiff.costheta[ds.section_index]
sintheta = lonlatdiff.sintheta[ds.section_index]
ds = ds.assign({'u_normal':ds.isel(ends=0).vvel_ms * costheta -
ds.isel(ends=0).uvel_ms * sintheta})
ds = ds.assign({'u_along':ds.isel(ends=0).vvel_ms * sintheta +
ds.isel(ends=0).uvel_ms * costheta})
# #### Find along-section distances of initial points
ds = ds.assign({'x':xr.DataArray(length_west[ds.section_index] + lonlatdiff.length[ds.section_index]*
(ds.isel(ends=0).lon - lonlat.lon[ds.section_index])/lonlatdiff.lon[ds.section_index],dims='traj')})
# ### volume, temperature and salt transports along track
# at osnap line
ds = ds.assign({'vol_trans_normal':np.sign(ds.u_normal) * particle_section_area/1.0e06})
ds = ds.assign({'particle_vol':ds.vol_trans_normal/ds.u_normal})
# #### Temporary fix for particle numbers in subset, based on constant total particle volume (basically area of section, use annual total)
# at osnap line
ds = ds.assign({'temp_transport':ds.temp * ds.vol_trans_normal})
ds = ds.assign({'salt_transport':ds.salt * ds.vol_trans_normal})
ds = ds.assign({'depth_transport':ds.z * ds.vol_trans_normal})
ds = ds.assign({'lon_transport':ds.lon * ds.vol_trans_normal})
ds = ds.assign({'tempxvol':ds.temp * ds.particle_vol})
ds = ds.assign({'saltxvol':ds.salt * ds.particle_vol})
ds = ds.assign({'depthxvol':ds.z * ds.particle_vol})
ds = ds.assign({'lonxvol':ds.lon * ds.particle_vol})
ds
# + [markdown] slideshow={"slide_type": "skip"}
# ## Plot section
# + slideshow={"slide_type": "skip"}
sns.set(style="whitegrid")
central_lon, central_lat = -30, 55
fig, ax = plt.subplots(subplot_kw={'projection': ccrs.Orthographic(central_lon, central_lat)})
extent = [-60, 0, 40, 70]
ax.set_extent(extent)
ax.gridlines()
ax.coastlines(resolution='50m')
lonlat.plot.scatter(ax=ax,transform=ccrs.PlateCarree(),x='lon',y='lat')
lonlat2mean.plot.scatter(ax=ax,transform=ccrs.PlateCarree(),x='lon',y='lat');
# -
# ### group by times at source
starttime = np.datetime64('1980-01-01T00:00')
deltat = np.timedelta64(5,'D')
times = np.array([starttime + i * deltat for i in range(2923)])
other_sum_sourcetime = xr.concat(
[ds.isel(ends=1,traj=slice(subsetno,None,nsubsets))
.where(ds.isel(ends=0,traj=slice(subsetno,None,nsubsets)).other_is_source)
.groupby_bins("time",times).sum()
for subsetno in range(0,nsubsets)],
dim="subsetno",
)
other_mean_sourcetime = xr.concat(
[ds.isel(ends=1,traj=slice(subsetno,None,nsubsets))
.where(ds.isel(ends=0,traj=slice(subsetno,None,nsubsets)).other_is_source)
.groupby_bins("time",times).mean()
for subsetno in range(0,nsubsets)],
dim="subsetno",
)
time_mid = [v.mid for v in other_sum_sourcetime.time_bins.values]
other_sum_sourcetime["time_bins"]=time_mid
time_mid = [v.mid for v in other_mean_sourcetime.time_bins.values]
other_mean_sourcetime["time_bins"]=time_mid
other_sum_sourcetime.to_netcdf(output_data_path / str('other_sum_sourcetime_'+yearstr+'.nc'))
other_mean_sourcetime.to_netcdf(output_data_path / str('other_mean_sourcetime_'+yearstr+'.nc'))
# + slideshow={"slide_type": "skip"}
conda list
# -
| notebooks/exploratory/107_afox_trackendsandpaths_fullstats_forpaper_fix2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generate the AudioSet ontology
#
# Using [OwlReady2](https://owlready2.readthedocs.io/en/latest/index.html) package. Ontology documentation published at https://maastrichtu-ids.github.io/audioset-owl
#
# First define the Notebook parameters for [papermill](https://papermill.readthedocs.io/en/latest/usage-parameterize.html)
# + tags=["parameters"]
# Papermill parameters. Do not delete this cell.
output_format = 'rdfxml'
audioset_ontology_uri = 'https://w3id.org/audioset'
# -
# Import the library and define the local `ontologies` folder. If an URL is given, first searches for a local copy of the OWL file and, if not found, tries to download it from the Internet.
# +
from owlready2 import *
import types
if output_format == 'ntriples':
output_extension = 'nt'
else:
output_extension = 'rdf'
global audioset_onto
global audioset_curated_hash
onto_path.append("/notebooks/ontologies")
# -
# ### Create and load ontologies
#
# Create the AudioSet ontology and load the Pizza ontology from the Internet (for example purpose)
# +
audioset_onto = get_ontology(audioset_ontology_uri)
pizza_onto = get_ontology("http://www.lesfleursdunormal.fr/static/_downloads/pizza_onto.owl").load()
# -
# ### Create AudioSet OWL ontology from the JSON
#
# * Get [AudioSet ontology JSON from GitHub](https://github.com/audioset/ontology)
# * [AudioSet Top classes](https://research.google.com/audioset/ontology/index.html): Human sounds, Animal, Music, Sounds of things, Natural sounds, source-ambiguous things, "Channel, environment and background"
# * Add classes respecting hierarchy provided in the JSON through the `child_ids` field
#
# See [OwlReady2 documentation](https://owlready2.readthedocs.io/en/latest/index.html) for:
# * [Dynamic Classes](https://owlready2.readthedocs.io/en/latest/class.html#creating-classes-dynamically)
# * [Add annotations to a Class](https://owlready2.readthedocs.io/en/latest/annotations.html?highlight=comment#adding-an-annotation): `comment`, `isDefinedBy`, `label`, `seeAlso`, `backwardCompatibleWith`, `deprecated`, `incompatibleWith`, `priorVersion`, `versionInfo`
# * [Properties](https://owlready2.readthedocs.io/en/latest/properties.html)
#
# Note: classes with multiple parents are properly defined, see `ChirpTweet` or the graph visualization as example
import requests, json
audioset_json = json.loads(requests.get("https://raw.githubusercontent.com/audioset/ontology/master/ontology.json").text)
def generate_owl_class(class_json, parent_class):
"""Recursively generates OWL classes and instances, original hierarchy respected using child_ids."""
with audioset_onto:
NewClass = types.new_class(class_json['uri_id'], (parent_class,))
NewClass.label = locstr(class_json['name'], lang = "en")
NewClass.comment = locstr(class_json['description'], lang = "en")
NewClass.comment = class_json['id']
if class_json['citation_uri']:
NewClass.comment = class_json['citation_uri']
if class_json['positive_examples']:
# Generate instances
for youtube_example in class_json['positive_examples']:
NewClass(comment = 'https://' + youtube_example)
for child in class_json['child_ids']:
generate_owl_class(audioset_curated_hash[child], NewClass)
# +
c = 0
# Create a hash using google audioset ID as key
audioset_curated_hash = {}
for row in audioset_json:
# Generate the ID that will be used for the ontology URI
uri_id = row['name'].replace(',', '').replace(')', '').replace('(', '').replace('.', '').replace("'", '').replace(";", '')
uri_id = uri_id.title().replace(' ', '').replace('-', '')
audioset_curated_hash[row['id']] = row
audioset_curated_hash[row['id']]['uri_id'] = uri_id
c += 1
print('Number of classes in the original AudioSet JSON: ' + str(c))
# Recursively generates classes starting from AudioSet top classes
audioset_top_classes = ['/m/0dgw9r', '/m/0jbk', '/m/04rlf', '/t/dd00041', '/m/059j3w', '/t/dd00098', '/t/dd00123']
for top_class in audioset_top_classes:
generate_owl_class(audioset_curated_hash[top_class], Thing)
# -
# ### Example to generate properties with domain and ranges
# +
# with audioset_onto:
# class Accent(Thing):
# pass
# class has_accent(ObjectProperty):
# domain = [HumanVoice]
# range = [Accent]
# class description(ObjectProperty):
# range = [str]
# -
# ### Add metadata to the ontology
audioset_onto.metadata.comment.append("OWL Ontology for the AudioSet ontology from Google defined in JSON.")
# ### Save the ontology file
#
# Ontology files saved in the `ontologies` folder.
#
# 2 formats available, defined in the papermill parameters (at the start of the notebook or in the `papermill-config.json` file):
# * `rdfxml`
# * `ntriples`
audioset_onto.save(file = "ontologies/audioset." + output_extension, format = output_format)
# # Explore the ontology
#
# **With OwlReady2**, e.g. list an ontology classes and properties.
# Get a class IRI:
print(audioset_onto.HumanVoice.iri)
# List all 682 classes:
#print(list(audioset_onto.classes()))
# List object properties:
print(list(audioset_onto.object_properties()))
# List a class instances:
for i in audioset_onto.InsideSmallRoom.instances(): print(i)
# ### Use Ontospy to analyze the ontology
#
# Load the ontology file with `ontospy`, then:
# * print top classes and the class tree
# * print instances of a class
import ontospy
audioset_spy = ontospy.Ontospy("ontologies/audioset.rdf", verbose=True)
# audioset_spy.printClassTree()
audioset_spy.toplayer_classes
# Print instances of Sigh class
audioset_spy.get_class('Sigh')[0]
for instance in audioset_spy.get_class('Sigh')[0].instances:
print(instance.uri, instance.qname)
instance.printTriples()
# ### Visualize with Ontospy docs
#
# Experimental, it is recommended to generate the documentation from the commandline (cf. `README.md` file)
# +
# from ontospy.ontodocs.viz.viz_html_single import *
# v = HTMLVisualizer(audioset_spy) # => instantiate the visualization object
# v.build("/notebooks/docs") # => render visualization. You can pass an 'output_path' parameter too
# v.preview() # => open in browser
# -
# ### Visualize with WebVOWL
#
# Use the URL to the ontology file:
#
# [http://www.visualdataweb.de/webvowl/#iri=https://raw.githubusercontent.com/MaastrichtU-IDS/audioset-owl/master/ontologies/audioset.rdf](http://www.visualdataweb.de/webvowl/#iri=https://raw.githubusercontent.com/MaastrichtU-IDS/audioset-owl/master/ontologies/audioset.rdf)
# ### Load the ontology RDF with `rdflib`
#
# Use `rdflib` and `networkx` to load the data in the graph and display it (not working with the ontology size, to be improved.
#
# Visualize as graph using networkx
# + tags=["outputPrepend"]
import rdflib
from rdflib import Graph, ConjunctiveGraph, plugin, Literal, RDF, URIRef, Namespace
from rdflib.serializer import Serializer
from rdflib.namespace import RDFS, XSD, DC, DCTERMS, VOID, OWL, SKOS
# from rdflib.plugins.sparql.parser import Query, UpdateUnit
# from rdflib.plugins.sparql.processor import translateQuery
# from rdflib.extras.external_graph_libs import rdflib_to_networkx_multidigraph
# import networkx as nx
# import matplotlib.pyplot as plt
g = rdflib.Graph()
result = g.parse('ontologies/audioset.rdf', format='xml')
for owl_class in g.subjects(RDF.type, OWL.Class):
print(owl_class)
# G = rdflib_to_networkx_multidigraph(result)
# # # Plot Networkx instance of RDF Graph
# pos = nx.spring_layout(G, scale=3)
# edge_labels = nx.get_edge_attributes(G, 'r')
# nx.draw_networkx_edge_labels(G, pos, labels=edge_labels)
# nx.draw(G, with_labels=True)
# -
| EditAudioSet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from scipy.stats import norm
# ## Brownian Limit
#
# In the Brownian limit, the ratio of the mass $m$ of the background particles to that of the selected heavy B particle $M_B$, $\lambda = \frac{m}{M_B}$, becomes small, it is then convenient to divide the particles up into two subgroups because of hte enormous difference in time scales of motion of the B and bath particles.
#
# In the Brownian limit $\lambda = \sqrt{\frac{m}{M_B}} \rightarrow 0$, memory function for heavy particles given by delta function in time,
#
# $$
# K_v(t) = \lambda_1 \delta(t)
# $$
#
# or
#
# $$
# \tilde{K_v}(s) = \lambda_1 = \dfrac{\zeta}{M_B} = \gamma
# $$
#
# where $\gamma$ is friction coeff and $\zeta$ the friction factor $\zeta = M_B \gamma$.
# ## Stokes Einstein
#
# If Stokes-Einstein holds, then friction factor $\gamma$ is
#
# $$
# \gamma = 6 \pi m_i \eta a_i
# $$
#
# $$
# \gamma = \dfrac{k_B T}{m_i D_s}
# $$
#
# Now writing chosen particle's velocity $v_i$ as $V_B$ and mass as $M_B$ gives
#
# $$
# M_B \dfrac{d}{dt} V_B(t) = - \zeta V_B(t) + F_{B}^{R}(t)
# $$
#
# and
#
# $$
# \langle F_B^R(0) \rangle = 0 \\
# \langle F_B^R(0) \cdot F_B^R(t) \rangle = 3 \gamma M_B k_B T \delta(t)
# $$
#
# or
#
# $$
# \langle v_i \cdot v_i \rangle = \dfrac{3k_B T}{m_i}
# $$
# +
Ndim = 2
N = 10000
dp = 1e-6
nu = 8.9e-4
T = 293
kB = 1.38e-23
pi = np.pi
T = 10000.0
dt = T/N
# +
def get_Dtheor(T, Ndim, dp, nu):
Dtheor = (kB*T)/(3*Ndim*pi*dp*nu)
return Dtheor
Dtheor = get_Dtheor(T,Ndim,dp,nu)
print(Dtheor)
# -
# Variance of step size distribution
# (units of m)
var = 2*Dtheor*dt
stdev = np.sqrt(var)
print(stdev)
# ## Verification of the Diffusion Coefficient
#
# We are simulating random walks (integrating a single random realization of a random diffusion walk) using some parameter to control the distribution of step size. This distribution results in a diffusion coefficient.
#
# We can verify that the diffusion coefficient we back out from the realizations of random walks matches the theoretical diffusion coefficient.
#
# To back out the diffusion coefficient from MSD:
#
# * Compute MSD versus lag time
# * Plot MSD versus lag time
# * Fit data to line - displacement vs. time
# * This velocity is proportional to $v \sim \dfrac{2D}{\delta t}$
#
# [This page](https://tinevez.github.io/msdanalyzer/tutorial/MSDTuto_brownian.html) mentions a reference for the 2D/t relation, which is also derived in the stat mech textbook mentioned in notebook 4, and is also derived (third method) in the brownian motion notes Z sent me.
# +
# Single random diffusion walk
# mean 0, std dev computed above
dx = stdev*np.random.randn(N,)
dy = stdev*np.random.randn(N,)
x = np.cumsum(dx)
y = np.cumsum(dy)
# -
plt.plot(x, y, '-')
plt.xlabel('x'); plt.ylabel('y');
plt.title("Brownian Motion 2D Walk")
plt.show()
# +
# Compute MSD versus lag time
# 0 to sqrt(N) avoids bias of longer lag times
upper = int(round(np.sqrt(N)))
msd = np.zeros(upper,)
lag = np.zeros(upper,)
for i, p in enumerate(range(1,upper+1)):
lagtime = dt*p
delx = ( x[p:] - x[:-p] )
dely = ( y[p:] - y[:-p] )
msd[i] = np.mean(delx*delx + dely*dely)
lag[i] = lagtime
# -
m, b = np.polyfit(lag, msd, 1)
# +
plt.loglog(lag, msd, 'o')
plt.loglog(lag, m*lag+b, '--k')
plt.xlabel('Lag time (s)')
plt.ylabel('MSD (m)')
plt.title('Linear Fit: MSD vs. Lag Time')
plt.show()
print("linear fit:")
print("Slope = %0.2g"%(m))
print("Intercept = %0.2g"%(b))
# -
# **NOTE:** If the total time being simulated *decreases* such that timesteps are on the order of $10^{-1}$ or $10^{-2}$, the scale of the MSD becomes $10^{-14}$ and numerical error becomes significant.
# +
# Slope is:
# v = dx / dt
# v = 2 D / dt
# Rearrange:
# D = v * dt / 2
v = m
Dempir = (v*dt)/2
err = (np.abs(Dtheor-Dempir)/Dtheor)*100
print("Theoretical D:\t%0.4g"%(Dtheor))
print("Empirical D:\t%0.4g"%(Dempir))
print("Percent Error:\t%0.4g"%(err))
print("\nNote: this result is from a single realization. Taking an ensemble yields a more accurate predicted D.")
# -
def msd_ensemble(T, Ndim, dp, nu, N, Nwalks):
Dtheor = get_Dtheor(T, Ndim, dp, nu)
ms = []
msds = []
msdxs = []
msdys = []
lags = []
for w in range(Nwalks):
# Single random diffusion walk
# mean 0, std dev computed above
dx = stdev*np.random.randn(N,)
dy = stdev*np.random.randn(N,)
# accumulate
x = np.cumsum(dx)
y = np.cumsum(dy)
# Compute MSD versus lag time
# 0 to sqrt(N) avoids bias of longer lag times
upper = int(round(np.sqrt(N)))
msd = np.zeros(upper,)
msdx = np.zeros(upper,)
msdy = np.zeros(upper,)
lag = np.zeros(upper,)
for i, p in enumerate(range(1,upper+1)):
lagtime = dt*p
delx = ( x[p:] - x[:-p] )
dely = ( y[p:] - y[:-p] )
msd[i] = np.mean((delx*delx + dely*dely)/2)
msdx[i] = np.mean(delx*delx)
msdy[i] = np.mean(dely*dely)
lag[i] = lagtime
slope, _ = np.polyfit(lag, msd, 1)
ms.append( slope )
msds.append( msd )
msdxs.append(msdx)
msdys.append(msdy)
lags.append( lag )
return (ms, msds, msdxs, msdys, lags)
# +
Ndim = 2
N = 10000
dp = 1e-6
nu = 8.9e-4
T = 293
kB = 1.38e-23
pi = np.pi
T = 10000.0
dt = T/N
Nwalks = 1000
# -
slopes, msds, msdxs, msdys, lags = msd_ensemble(T, Ndim, dp, nu, N, Nwalks)
# +
Dempir = np.mean((np.array(slopes)*dt)/2)
err = (np.abs(Dtheor-Dempir)/Dtheor)*100
print("Theoretical D:\t%0.4g"%(Dtheor))
print("Empirical D:\t%0.4g"%(Dempir))
print("Percent Error:\t%0.4g%%"%(err))
print("\nUsing an ensemble of %d particles greatly improves accuracy of predicted D."%(N))
# +
for i, (msd, lag) in enumerate(zip(msdxs,lags)):
if(i>200):
break
plt.loglog(lag,msd,'b',alpha=0.1)
for i, (msd, lag) in enumerate(zip(msdys,lags)):
if(i>200):
break
plt.loglog(lag,msd,'r',alpha=0.1)
for i, (msd, lag) in enumerate(zip(msds,lags)):
if(i>200):
break
plt.loglog(lag,msd,'k',alpha=0.1)
plt.xlabel('Lag Time (m)')
plt.ylabel('MSD (s)')
plt.title('MSD vs Lag Time: \nMSD X (blue), MSD Y (red), MSD MAG (black)')
plt.show()
# -
# **NOTE:** As mentioned above, the total time being simulated needs to be large enough that the lag time is larger than $10^{-2}$. At lag times of $10^{-2}$ the MSD values are near machine truncation values, so the solution is succeptible to roundoff error.
| Statistical Mechanics 6 MSD to Diffusion Coefficient.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.8 ('base')
# language: python
# name: python3
# ---
# #### Análise Exploratória Incial
# Importar bibliotecas
import numpy as np
import pandas as pd
# Carregar datasets
olist_orders = pd.read_csv("../data/raw/olist_orders_dataset.csv")
olist_order_items = pd.read_csv("../data/raw/olist_order_items_dataset.csv")
olist_customers = pd.read_csv("../data/raw/olist_customers_dataset.csv")
# ##### Análise Exploratória de Dados
# - Verificação de dados nulos
# - Verificação da tipagem de dados
# - Verificação de dados duplicados
# - Análise da distribuição de cada variável
# Imprime base olist_orders
olist_orders
# +
# Mostra o tipo dos dados e quantidade de non-null
olist_orders.info()
# Possuem valores nulos:
# order_approved_at,
# order_delivered_carrier_date e
# order_delivered_customer_date
# -
# Calcula quantidade de valores nulos
olist_orders.isnull().sum()
# Investiga valores nulos order_approved_at
olist_orders[olist_orders.order_approved_at.isnull()].order_status.value_counts()
# Investiga valores nulos order_delivered_carrier_date
olist_orders[olist_orders.order_delivered_carrier_date.isnull()].order_status.value_counts()
# Investiga valores nulos order_delivered_customer_date
olist_orders[olist_orders.order_delivered_customer_date.isnull()].order_status.value_counts()
# Verifica registros duplicados
olist_orders.duplicated().value_counts()
# Verifica frequência de cada order_status
olist_orders.order_status.value_counts()
# Imprime base olist_customers
olist_customers
# Mostra o tipo dos dados e quantidade de non-null
olist_customers.info()
# Verifica registros duplicados
olist_customers.duplicated().value_counts()
# Verifica quantidade de clientes
olist_customers.customer_unique_id.unique().shape
# Calcula quantidade de pedidos feitos por clientes repetidos
99441-olist_customers.customer_unique_id.unique().shape[0]
# Verifica frequência de Estados dos clientes
olist_customers.customer_state.value_counts()
# Imprime base olist_order_items
olist_order_items
# Mostra o tipo dos dados e quantidade de non-null
olist_order_items.info()
# Verifica registros duplicados
olist_order_items.duplicated().value_counts()
# Gera resumo estatístico descritivo
olist_order_items.order_item_id.describe()
# Gera resumo estatístico descritivo
olist_order_items.price.describe()
# Gera resumo estatístico descritivo
olist_order_items.freight_value.describe()
# Verifica item com 409.68 de frete
olist_order_items[olist_order_items.freight_value == 409.68]
| notebooks/analise-exploratoria-inicial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Hypothesis Testing
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(font_scale=2, palette= "viridis")
import researchpy as rp
# 
data = pd.read_csv('../data/pulse_data.csv')
data.head()
# ## Inspecting Variables
data.info()
# ## Summarize and Visualize Data
# summary stats of numerical variables
data.describe()
# summary stats of numerical variables
rp.summary_cat(data[['Gender', 'Smokes', 'Alcohol', 'Exercise','Ran','BMICat']])
# ## One Categorical Variable
# __Questions: Is there any difference in the number of men and women?__
data.Gender.value_counts()
data.Gender.value_counts(normalize=True)
plt.figure(figsize=(10,8))
sns.countplot(data=data, x='Gender')
plt.show()
# ## One Numeric Variable
# __Question: Is the average height different from a established height?__
data.Height.head()
data.Height.describe()
plt.figure(figsize=(10,8))
sns.boxplot(data=data, x='Height')
plt.show()
# ## Two Categorical Variables(C-C)
# __Question: Does the porportion of men and women differ across the BMI Category?__
data.Gender.unique()
data.BMICat.unique()
pd.crosstab(data['Gender'], data['BMICat'])
pd.crosstab(data['Gender'], data['BMICat'], normalize=True)
plt.figure(figsize=(10,8))
sns.countplot(data=data, x='BMICat', hue='Gender')
plt.show()
# ## One Quantitative and One Categorical Variables(Q-C)
# __Question: Is there a difference in the height between men and women?__
#
data.groupby('Gender')['Height'].mean()
plt.figure(figsize=(10,8))
sns.boxplot(data=data, x='Height', y="Gender")
plt.show()
# ## Two Quantitative Variables(Q-Q)
# __Question: Is there any relationship between height and weight?__
data.Height.corr(data.Weight)
plt.figure(figsize=(10,8))
sns.scatterplot(data=data, x='Height', y="Weight")
plt.show()
plt.figure(figsize=(10,8))
sns.regplot(data=data, x='Height', y="Weight")
plt.show()
# ## What is Inferential Statistics
# Assess the strength of evidence for/against a hypothesis; evaluate the data
#
# - Inferential statistical methods provide a confirmatory data analysis.
# - Generalize conclusions from data from part of a group (sample) to the whole group (population)
# - Assess the strength of the evidence
# - Make comparisons
# - Make predictions
# Inferential statistical methods divide into 2 categories.
#
# - **Hypothesis Testing:** Hypothesis testing is a formal procedure for investigating our ideas about the world using statistics. It is most often used by scientists to test specific predictions, called hypotheses, that arise from theories.
#
# - **Model Fitting:** Model fitting is a measure of how well a statistical learning model generalizes to similar data to that on which it was trained. A model that is well-fitted produces more accurate outcomes.
# ## What is Inference?
# The process of drawing conclusions about population parameters based on a sample taken from the population.
#
# - A sample is likely to be a good representation of the population.
# - There is an element of uncertainty as to how well the sample represents the population.
# - The way the sample is taken matters.
# ## What is Hypothesis?
# - Proposed explanation for a phenomenon.
# - A hypothesis is an educated guess about something in the world around you. It should be testable, either by experiment or observation.
#
# - Proposed explanation
# - Objectively testable
# - Singular - hypothesis
# - Plural - hypotheses
#
# __Examples__
#
# - A new medicine you think might work.
# - A way of teaching you think might be better.
# - A possible location of new species.
#
#
#
# ## Hypothesis and Study Design
# - **Hypothesis:** seat belts decreases the fatality rate
# - **Study design:** cross-sectional study of fatality outcome and seat-belt use of victims of motor vehicle accidents during a one-month time period in a large city
# 
# ## Effect of Seat Belt Use on Accident Fatality
# What is your conclusion?
# - The fatality rate is:
# - 40% in the group of drivers who did not wear seat belts
# - 20% in drivers who did wear seat belts
#
# - Seat belts appear to save lives
# ## The Inferential Questions of Interest
# - The inferential questions of interest are:
# - Are results applicable to the population of all drivers?
# (generalization)
# - Does wearing seat belts decreases fatality rate? (assess strength of evidence)
# - Is the fatality rate of those not wearing seat belts higher than the fatality rate of those wearing seat belts? (comparison)
# - How many lives can be saved by wearing seat belts? (prediction)
# - Do other variables influence the conclusion?
# - For example: the age of driver, alcohol use, type of car, speed at impact (ask more questions)
# ## Speed at Impact
# 
# ## How Does This Influence Your Conclusion?
# - How does this influence your conclusion?
# - The fatality rate is 10% at low-impact speeds regardless of seat-belt use
#
# - The fatality rate at high impact speeds is:
# - 60% in drivers not wearing seat belts
# - 35% in drivers wearing seat belts
# ## Null and Alternative Hypothesis
# - **Hypothesis 0 (H0):** Assumption of the test holds and is failed to be rejected at some level of significance.
# - **Hypothesis 1 (Ha):** Assumption of the test does not hold and is rejected at some level of significance.
# ## Errors in Statistical Tests
# - **Type I Error:** The incorrect rejection of a true null hypothesis or a false positive.
# - **Type II Error:** The incorrect failure of rejection of a false null hypothesis or a false negative.
#
# 
# ## Alpha($\alpha$)
# - $\alpha$ is probability of rejecting H0 when H0 is true.
# - $\alpha$ = Probability of Type-I error.
# - Ranges from 0 to 1
# - **High α is not good**
# ## p-value
# In statistics, the p-value is the probability of obtaining results at least as extreme as the observed results of a statistical hypothesis test, assuming that the null hypothesis is correct.
#
# generally cut off value of alpha 0.05
# - If p-value > alpha: Fail to reject the null hypothesis (i.e. not significant result).
# - If p-value <= alpha: Reject the null hypothesis (i.e. significant result).
# ## Hypothesis Testing Process
# - **Step-1:** Null Hypothesis H0
# - True until proven false
# - Usually posits no relationship
#
# - **Step-2:** Select Test
# - Pick from vast library
# - Know which one to choose
#
# - **Step-3:** Significance Level
# - Usually 1% or 5%
# - What threshold for luck?
#
# - **Step-4:** Alternative Hypothesis
# - Negation of null hypothesis
# - Usually asserts specific relationship
#
# - **Step-5:** Test Statistic
# - Convert to p-value
# - How likely it was just luck?
#
#
# - **Step-6:** Accept or Reject
# - Small p-value? Reject H0
# - Small: Below significance level
#
# 
# # Common Statistical Tests
# ### Variable Distribution Type Tests (Gaussian)
# - Shapiro-Wilk Test
# - D’Agostino’s K^2 Test
# - Anderson-Darling Test
#
# ### Variable Relationship Tests (correlation)
# - Pearson’s Correlation Coefficient
# - Spearman’s Rank Correlation
# - Kendall’s Rank Correlation
# - Chi-Squared Test
#
# ### Compare Sample Means (parametric)
# - Student’s t-test
# - Paired Student’s t-test
# - Analysis of Variance Test (ANOVA)
# - Repeated Measures ANOVA Test
#
# ### Compare Sample Means (nonparametric)
# - Mann-Whitney U Test
# - Wilcoxon Signed-Rank Test
# - Kruskal-Wallis H Test
# - Friedman Test
# ## Statistical Test Selection
# What we observe in our sample data | Is it real?(statistical test)|
# ----------|------|
# 1 categorical variable| 1 sample proportion test
# 2 categorical variables | chi squared test
# 1 numeric variable | t-test
# 1 numeric and 1 categorical variable | t-test or ANOVA
# more than 2 categorical variables | ANOVA
# 2 numeric variables | correlation test
# ## References
# - https://machinelearningmastery.com/statistical-hypothesis-tests/
# - https://www.statisticshowto.com/probability-and-statistics/hypothesis-testing/
| 07 - Inferential Stats with Python/notebooks/01_IntroductionToHypothesisTesting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="LjL6WmHnFsDm"
# # Rethinking Statistics course in NumPyro - Week 1
# + [markdown] id="wIiAZbVJFsDo"
# Lecture 1: The Golem of Prague
#
# - [Video](https://youtu.be/cclUd_HoRlo)
# - [Slides](https://speakerdeck.com/rmcelreath/statistical-rethinking-2022-lecture-01)
#
# Lecture 2: Bayesian Inference
#
# - [Video](https://www.youtube.com/watch?v=guTdrfycW2Q&list=PLDcUM9US4XdMROZ57-OIRtIK0aOynbgZN&index=2)
# - [Slides](https://speakerdeck.com/rmcelreath/statistical-rethinking-2022-lecture-02)
#
# [Proposed problems](https://github.com/rmcelreath/stat_rethinking_2022/blob/main/homework/week01.pdf) and [solutions in R](https://github.com/rmcelreath/stat_rethinking_2022/blob/main/homework/week01_solutions.pdf) for the exercises of the week.
# + executionInfo={"elapsed": 1511, "status": "ok", "timestamp": 1613851213838, "user": {"displayName": "<NAME>\u00e1rez", "photoUrl": "https://lh5.googleusercontent.com/-s0kzcIwylzA/AAAAAAAAAAI/AAAAAAAAQXA/v8Sc6WgQy7c/s64/photo.jpg", "userId": "06409440331868776168"}, "user_tz": -60} id="X3Nnt0q_0qUK"
import os
import arviz as az
import numpy as np
import jax.numpy as jnp
from jax import random
import numpyro
import numpyro.distributions as dist
import matplotlib.pyplot as plt
if "SVG" in os.environ:
# %config InlineBackend.figure_formats = ["svg"]
az.style.use("arviz-darkgrid")
numpyro.set_platform("cpu")
# -
# %load_ext watermark
# %watermark -n -u -v -iv -w
# The solutions are largely based on code examples provided by [Du Phan](https://fehiepsi.github.io/rethinking-numpyro/) that accompany the Statistical Rethinking [book](https://xcelab.net/rm/statistical-rethinking/) book.
# + [markdown] id="c9QACe2VFsEk"
# ## Exercise 1
# + [markdown] id="x9U5N9Y1FsEk"
# >Suppose the globe tossing data (Chapter 2) had turned out to be 4 water and 11 land. Construct the posterior distribution, using grid approximation. Use the same flat prior as in the book.
# + executionInfo={"elapsed": 521, "status": "ok", "timestamp": 1613851615338, "user": {"displayName": "Andr\u00e9<NAME>\u00e1rez", "photoUrl": "https://lh5.googleusercontent.com/-s0kzcIwylzA/AAAAAAAAAAI/AAAAAAAAQXA/v8Sc6WgQy7c/s64/photo.jpg", "userId": "06409440331868776168"}, "user_tz": -60} id="MeRFU-XmFsEl"
water = 4
land = 11
grid_size = 100
# define grid
p_grid = jnp.linspace(start=0, stop=1, num=grid_size)
# define prior
prior = jnp.repeat(1, grid_size)
# compute likelihood at each value in grid
likelihood = jnp.exp(dist.Binomial(total_count=water+land, probs=p_grid).log_prob(water))
# compute product of likelihood and prior
unstd_posterior = likelihood * prior
# standardize the posterior, so it sums to 1
posterior = unstd_posterior / jnp.sum(unstd_posterior)
# -
# Let's plot the posterior distribution
plt.plot(p_grid, posterior, "-o")
plt.xlabel("probability of water")
plt.ylabel("posterior probability")
plt.show()
samples = p_grid[dist.Categorical(posterior).sample(random.PRNGKey(0), (10000,))]
jnp.mean(samples)
# + [markdown] id="GQxAeX3FFsEz"
# ## Exercise 2
# + [markdown] id="PONtBDo-FsE0"
# >Now suppose the data are 4 water and 2 land. Compute the posterior again, but this time use a prior that is zero below $p = 0.5$ and a constant above $p = 0.5$. This corresponds to prior information that a majority of the Earth's surface is water.
# + executionInfo={"elapsed": 529, "status": "ok", "timestamp": 1613851626167, "user": {"displayName": "Andr\u0<NAME>\u00e1rez", "photoUrl": "https://lh5.googleusercontent.com/-s0kzcIwylzA/AAAAAAAAAAI/AAAAAAAAQXA/v8Sc6WgQy7c/s64/photo.jpg", "userId": "06409440331868776168"}, "user_tz": -60} id="BM5bu4-7FsE1"
water = 4
land = 2
grid_size = 100
# define grid
p_grid = jnp.linspace(start=0, stop=1, num=grid_size)
# define prior
new_prior = p_grid > 0.5
# compute likelihood at each value in grid
likelihood = jnp.exp(dist.Binomial(total_count=water+land, probs=p_grid).log_prob(water))
# compute product of likelihood and prior
unstd_posterior_new = likelihood * new_prior
# standardize the posterior, so it sums to 1
posterior_new = unstd_posterior_new / jnp.sum(unstd_posterior_new)
# + colab={"base_uri": "https://localhost:8080/", "height": 142} executionInfo={"elapsed": 553, "status": "ok", "timestamp": 1613851631803, "user": {"displayName": "Andr\u00e9<NAME>\u00e1rez", "photoUrl": "https://lh5.googleusercontent.com/-s0kzcIwylzA/AAAAAAAAAAI/AAAAAAAAQXA/v8Sc6WgQy7c/s64/photo.jpg", "userId": "06409440331868776168"}, "user_tz": -60} id="qtdbt7_cFsFC" outputId="225473dc-a9e3-4209-8043-27a847cbf160"
plt.plot(p_grid, posterior_new, "-o")
plt.xlabel("probability of water")
plt.ylabel("posterior probability")
plt.show()
# -
samples_new = p_grid[dist.Categorical(posterior_new).sample(random.PRNGKey(1), (10000,))]
jnp.mean(samples_new)
# + [markdown] id="-X7n4j-FFsFI"
# ## Exercise 3
# + [markdown] id="8bN142K_FsFJ"
# >For the posterior distribution from 2, compute 89% percentile and HPDI intervals. Compare the widths of these intervals. Which is wider? Why? If you had only the information in the interval, what might you misunderstand about the shape of the posterior distribution?
# -
a = (100 - 89) / 2
b = 100 - a
jnp.percentile(samples_new, jnp.array([a, b]))
numpyro.diagnostics.hpdi(samples_new, prob=0.89)
# We can see that the percentile interval (the top one) is wider, while the HDPI interval is narrower. HDPI should be narrower and include the point with highest posterior probability. When these intervals are very different, it is better to plot and report the whole posterior distribution to see what is going on.
# ## Exercise 4
# >Suppose there is bias in sampling so that Land is more likely than Water to be recorded. Specifically, assume that 1-in-5 (20%) of Water samples are accidentally recorded instead as ”Land”. First, write a generative simulation of this sampling process. Assuming the true proportion of Water is 0.70, what proportion does your simulation tend to produce instead? Second, using a simulated sample of 20 tosses, compute the unbiased posterior distribution of the true proportion of water.
# First, let's see how an unbiased sample would look like
# +
# true proportion of water
p = 0.7
num_tosses = 20
seed = 100
num_samples = 1000
key = random.PRNGKey(seed)
num_waters = dist.Binomial(total_count=num_tosses, probs=p).sample(key, (num_samples,))
# -
jnp.mean(num_waters/num_tosses)
az.plot_dist(num_waters, kind="hist", hist_kwargs={"rwidth": 0.1})
# Now let's generate a biased sample
def generate_samples(num_tosses, bias=0.2, seed=100, key=None):
# true proportion of water
p = 0.7
sample = []
if key is None:
key = random.PRNGKey(seed)
for i in range(num_tosses):
key, subkey1, subkey2 = random.split(key, 3)
sample_i = "L" if random.uniform(subkey1) > p else "W"
if sample_i == "W":
sample_i = sample_i if random.uniform(subkey2) > bias else "L"
sample.append(sample_i)
return np.array(sample)
def count_water(sample):
return sum(sample == "W")
# +
num_tosses = 20
num_samples = 1000
key = random.PRNGKey(101)
keys = random.split(key, num_samples)
num_waters = []
for i in range(num_samples):
sample = generate_samples(num_tosses, bias=0.2, key=keys[i])
water = count_water(sample)
num_waters.append(water)
# -
np.mean(np.array(num_waters)/num_tosses)
az.plot_dist(num_waters, kind="hist", hist_kwargs={"rwidth": 0.1})
# The same can be more succintly and efficiently computed as follows
# +
# true proportion of water
p = 0.7
# sampling bias
bias = 0.2
num_tosses = 20
seed = 100
num_samples = 1000
key = random.PRNGKey(seed)
key, subkey = random.split(key)
num_waters_unbiased = dist.Binomial(total_count=num_tosses, probs=p).sample(key, (num_samples,))
num_waters = dist.Binomial(total_count=num_waters_unbiased, probs=(1-bias)).sample(subkey, (num_samples,))
# -
jnp.mean(num_waters/num_tosses)
# In one step, the biased proportion of water can be computed as $p\times(1-bias)$. Let's check
# +
# true proportion of water
p = 0.7
# sampling bias
bias = 0.2
num_tosses = 20
seed = 100
num_samples = 1000
key = random.PRNGKey(seed)
num_waters = dist.Binomial(total_count=num_tosses, probs=p*(1-bias)).sample(key, (num_samples,))
jnp.mean(num_waters/num_tosses)
# -
# Finally, let's compute an unbiased posterior distribution
num_tosses = 20
sample = generate_samples(num_tosses, bias=0.2, seed=102)
print(sample)
# +
water = count_water(sample)
# Alternatively:
# water = dist.Binomial(total_count=num_tosses, probs=p*(1-bias)).sample(random.PRNGKey(105), (1,))
grid_size = 100
# define grid
p_grid = jnp.linspace(start=0, stop=1, num=grid_size)
# define prior
prior = jnp.repeat(1, grid_size)
# compute likelihood at each value in grid
likelihood_biased = jnp.exp(dist.Binomial(total_count=num_tosses, probs=p_grid).log_prob(water))
# account for the sampling bias
likelihood_unbiased = jnp.exp(dist.Binomial(total_count=num_tosses, probs=p_grid*(1-bias)).log_prob(water))
# compute product of likelihood and prior
unstd_posterior_biased = likelihood_biased * prior
unstd_posterior_unbiased = likelihood_unbiased * prior
# standardize the posterior, so it sums to 1
posterior_biased = unstd_posterior_biased / jnp.sum(unstd_posterior_biased)
posterior_unbiased = unstd_posterior_unbiased / jnp.sum(unstd_posterior_unbiased)
# -
plt.plot(p_grid, posterior_biased, color="red")
plt.plot(p_grid, posterior_unbiased, color="blue")
plt.xlabel("probability of water")
plt.ylabel("posterior probability")
plt.show()
| statrethink_numpyro_w01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# Word Representations of Words
embeddings = tf.Variable(
tf.random_uniform([vocabulary_size, embedding_size], -1.0, 1.0))
nce_weights = tf.Variable(
tf.truncated_normal([vocabulary_size, embedding_size],
stddev=1.0 / math.sqrt(embedding_size)))
nce_biases = tf.Variable(tf.zeros([vocabulary_size]))
# -
# Placeholders for inputs
train_inputs = tf.placeholder(tf.int32, shape=[batch_size])
train_labels = tf.placeholder(tf.int32, shape=[batch_size, 1])
embed = tf.nn.embedding_lookup(embeddings, train_inputs)
# Compute the NCE loss, using a sample of the negative labels each time.
loss = tf.reduce_mean(
tf.nn.nce_loss(weights=nce_weights,
biases=nce_biases,
labels=train_labels,
inputs=embed,
num_sampled=num_sampled,
num_classes=vocabulary_size))
# We use the SGD optimizer.
optimizer = tf.train.GradientDescentOptimizer(learning_rate=1.0).minimize(loss)
for inputs, labels in generate_batch(...):
feed_dict = {train_inputs: inputs, train_labels: labels}
_, cur_loss = session.run([optimizer, loss], feed_dict=feed_dict)
| crackingcode/day10/cc_tf_day10_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import glob, serial, time, math
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
from skimage.transform import iradon
from IPython.display import display
import random
ports = glob.glob('/dev/tty[A-Za-z]*') # Modify if on a platform other than Linux
ports[0]
ser = serial.Serial(ports[0], 115200)
# +
ds = {0:b'i',
7:b'j',
3:b'k',
4:b'l',
2:b'm',
5:b'n',
6:b'o',
1:b'p'}
es = {0:b'a',
7:b'b',
1:b'c',
6:b'd',
2:b'e',
4:b'f',
3:b'g',
5:b'h'}
def read_d(d):
ser.write(ds[d])
while ser.in_waiting < 1:
pass # wait for a response
return ser.read_all()
def set_l(l):
ser.write(es[l])
while ser.in_waiting < 1:
pass # wait for a response
return ser.read_all()
def loff():
ser.write(b'A')
while ser.in_waiting < 1:
pass # wait for a response
return ser.read_all()
def read_all():
a = []
# Take a base set of readings
loff()
for d in range(8):
a.append(int(read_d(d).strip()))
for l in range(8):
loff()
set_l(l)
for d in range(8):
a.append(int(read_d(d).strip()))
loff()
return a
# -
# %matplotlib inline
# +
from IPython.display import clear_output
X = []
y = []
# -
for i in range(300):
# fig, ax = plt.subplots()
readings = read_all()
X.append(readings)
y.append([0,0]) # 0 = empty, 1-4 = quadrants. # 0 = nothing, 1=pen back, 2=cutter back, 3=shiny pen, 4=finger
# +
# df1 = pd.DataFrame(X)
# df1['quadrant'] = [m[0] for m in y]
# df1['object'] = [m[1] for m in y]
# objects = {0:'empty', 1:'black_pen_back', 2:'cutter_back', 3:'shiny_pen', 4:'finger'}
# df1['object_name'] = [objects[m[1]] for m in y]
# df1.to_csv('dataset_2qr.csv')
df1 = pd.read_csv('dataset_2q.csv')
df1 = df1.drop('Unnamed: 0', axis=1)
# -
df1.head()
# +
X_train, X_test, y_train, y_test = train_test_split(df1[[str(i) for i in range(72)]], df1['object'])
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
mlpc = MLPClassifier(hidden_layer_sizes=(20, 20, 20), max_iter=1000)
mlpc.fit(X_train, y_train)
mlpc.score(X_test, y_test)
# +
from sklearn.neural_network import MLPClassifier
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
df3 = df1.loc[df1['object']%4 < 2] # What a hack. This excludes classes 2 and 3, but not 0, 1 and 4!
X_train, X_test, y_train, y_test = train_test_split(df3[[str(i) for i in range(72)]], df3['object'])
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
mlpc = MLPClassifier(hidden_layer_sizes=(20, 20, 20), max_iter=1000)
mlpc.fit(X_train, y_train)
mlpc.score(X_test, y_test)
# -
# Do confusion matrix (can do for prev as well)
from sklearn.metrics import confusion_matrix
confusion_matrix(y_test, mlpc.predict(X_test))
# +
from sklearn.neural_network import MLPClassifier
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(df1[[str(i) for i in range(72)]], df1['quadrant'])
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
scores = []
ns = []
for i in range(10, 1000, 10):
mlpc = MLPClassifier(hidden_layer_sizes=(20, 20, 20), max_iter=1000)
mlpc.fit(X_train[:i], y_train[:i])
scores.append(mlpc.score(X_test, y_test))
ns.append(i)
# mlpc = MLPClassifier(hidden_layer_sizes=(20, 20, 20), max_iter=1000)
# mlpc.fit(X_train, y_train)
# mlpc.score(X_test, y_test)
# +
df3 = df1.loc[df1['object']%4 < 2] # What a hack. This excludes classes 2 and 3, but not 0, 1 and 4!
X_train, X_test, y_train, y_test = train_test_split(df3[[str(i) for i in range(72)]], df3['object'])
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
scores = []
ns = []
for i in range(10, 1000, 10):
mlpc = MLPClassifier(hidden_layer_sizes=(20, 20, 20), max_iter=1000)
mlpc.fit(X_train[:i], y_train[:i])
scores.append(mlpc.score(X_test, y_test))
ns.append(i)
# +
X_train, X_test, y_train, y_test = train_test_split(df1[[str(i) for i in range(72)]], df1['object'])
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
scores2 = []
ns = []
for i in range(10, 1000, 10):
mlpc = MLPClassifier(hidden_layer_sizes=(20, 20, 20), max_iter=1000)
mlpc.fit(X_train[:i], y_train[:i])
scores2.append(mlpc.score(X_test, y_test))
ns.append(i)
# -
plt.plot(ns, scores, label='All classes')
plt.plot(ns, scores2, label='Finger vs Pen')
plt.xlabel('Number of training samples')
plt.ylabel('Score')
plt.savefig('ring_of_8_classification.png')
sdf = pd.read_csv('scores_simulation_ring_of_8_overlap_varied.csv')
sdf['Observed'] = scores
ax = sdf.plot(x='Number of Samples')
ax.set_ylabel('Score')
# +
plt.plot(ns, sdf['Low Overlap'], alpha=0.4, label = 'Low Overlap (simulation)')
plt.plot(ns, sdf['Partial Overlap'], alpha=0.4, label = 'Partial Overlap (simulation)')
plt.plot(ns, sdf['Full Overlap'], alpha=0.4, label = 'Full Overlap (simulation)')
plt.plot(ns, sdf['Observed'])
plt.xlabel('Number of Training Samples')
plt.ylabel('Prediction Accuracy')
plt.legend()
plt.savefig('ring_of_8_observed_accuracy_vs_prediction_alpha.png')
# -
import pandas as pd
Xdf = pd.DataFrame(X)
Xdf['class'] = y
Xdf.to_csv('quadrants.csv')
while True:
readings = read_all()
Xt2 = scaler.transform([readings])
print(mlpc.predict(Xt2))
clear_output(wait=True)
X = []
y = []
for i in range(40):
readings = read_all()
X.append(readings)
y.append(2) # 0 empty, 1 finger, 2 pen
# +
X_train2, X_test2, y_train2, y_test2 = train_test_split(X[-120:], y[-120:])
scaler = StandardScaler()
scaler.fit(X_train2)
X_train2 = scaler.transform(X_train2)
X_test2 = scaler.transform(X_test2)
mlpc = MLPClassifier(hidden_layer_sizes=(20, 20, 20), max_iter=1000)
mlpc.fit(X_train2, y_train2)
mlpc.score(X_test2, y_test2)
# -
readings = read_all()
Xt2 = scaler.transform([readings])
p = mlpc.predict(Xt2)[0]
if p == 0:
print('Empty')
elif p == 1:
print('Finger')
else:
print('Pen')
y_test2
while True:
fig, (ax1, ax2) = plt.subplots(1, 2, sharey=True)
a = read_all()
for e in range(8):
for d in range(8):
a[(e+1)*8 + d] -= a[d]
ax2.plot(a[(e+1)*8:][:8])
ax1.plot(a[8:])
ax1.set_ylim(-500, 200)
plt.show()
# time.sleep(0.3)
clear_output(wait=True)
a = []
for i in range(100):
time.sleep(0.2)
s = ''
vals = []
v2 = []
for i in range(8):
v = int(read_d(0).strip())
s += ", "
s += str(v)
vals.append(v)
set_l(0) # Turn on LED
time.sleep(0.2)
for i in range(8):
v = int(read_d(0).strip())
v2.append(v)
s += ", "
s += str(v)
loff(0) # LEDs off
arr.append(vals)
arr2.append(v2)
a1 = np.asarray(vals)
a2 = np.asarray(v2)
print(a1-a2)
plt.plot(a1-a2)
clear_output(wait=True)
ser.read_all()
a1 = np.asarray(arr)
a2 = np.asarray(arr2)
print(a1-a2)
a1 = np.asarray(arr)
a2 = np.asarray(arr2)
print(a1-a2)
| Misc Notebooks/Ring of 8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Random Forest
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# ## Dataset
dataset = pd.read_csv('Social_Network_Ads.csv')
dataset.head()
X = dataset.iloc[:,2:4].values
y = dataset.iloc[:,-1].values
# ## Split data
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.25,random_state=0)
# ## Feature Scaling
from sklearn.preprocessing import StandardScaler
sc_X = StandardScaler()
X_train = sc_X.fit_transform(X_train)
X_test = sc_X.transform(X_test)
# ## Fit model
from sklearn.ensemble import RandomForestClassifier
classifier = RandomForestClassifier(n_estimators=10, criterion='entropy', random_state=0)
classifier.fit(X_train, y_train)
# ## Predict
y_pred = classifier.predict(X_test)
y_pred
# ## Confusion matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
cm
# ## Visualization
# +
from matplotlib.colors import ListedColormap
X_set, y_set = X_train, y_train
X1, X2 = np.meshgrid(
np.arange(start=X_set[:,0].min() - 1, stop=X_set[:,0].max() + 1, step=0.01),
np.arange(start=X_set[:,1].min() - 1, stop=X_set[:,1].max() + 1, step=0.01)
)
plt.contourf(
X1,
X2,
classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha=0.75,
cmap=ListedColormap(('red', 'green'))
)
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i,j in enumerate(np.unique(y_set)):
plt.scatter(
X_set[y_set==j, 0],
X_set[y_set==j, 1],
c=ListedColormap(('red', 'green'))(i),
label=j
)
plt.title('Random Forest (Training set)')
plt.xlabel('Age')
plt.ylabel('Estimated Salary')
plt.legend()
plt.show()
# +
from matplotlib.colors import ListedColormap
X_set, y_set = X_test, y_test
X1, X2 = np.meshgrid(
np.arange(start=X_set[:,0].min() - 1, stop=X_set[:,0].max() + 1, step=0.01),
np.arange(start=X_set[:,1].min() - 1, stop=X_set[:,1].max() + 1, step=0.01)
)
plt.contourf(
X1,
X2,
classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha=0.75,
cmap=ListedColormap(('red', 'green'))
)
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i,j in enumerate(np.unique(y_set)):
plt.scatter(
X_set[y_set==j, 0],
X_set[y_set==j, 1],
c=ListedColormap(('red', 'green'))(i),
label=j
)
plt.title('Random Forest (Training set)')
plt.xlabel('Age')
plt.ylabel('Estimated Salary')
plt.legend()
plt.show()
| z_Miscellaneous/ML2/Classification/13_RandomForestClassification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# This reads in Balmer line widths and plots the scaling with respect to H-delta
# -
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
file_name = "/Users/bandari/Documents/git.repos/rrlfe/ew_products/all_data_input_mcmc_20220130_run_1.csv"
df = pd.read_csv(file_name)
df.keys()
plt.figure(figsize=(10,7))
plt.scatter(df["EW_Hdelta"], df["EW_Hgamma"], s=12, label="W"+r"$\gamma$")
plt.scatter(df["EW_Hdelta"], np.add(df["EW_Hbeta"],5), s=12, label="W"+r"$\beta$ + 5")
plt.scatter(df["EW_Hdelta"], np.add(df["EW_Heps"],10), s=12, label="W"+r"$\epsilon$ + 10")
plt.xlabel(r"$W_{\delta}$"+" "+"($\AA$)", fontsize=25)
plt.ylabel("$W$"+" "+"($\AA$)", fontsize=25)
plt.legend(fontsize=20)
plt.xticks(fontsize=20)
plt.yticks(fontsize=20)
plt.show()
# +
# sanity check of the errors
# -
plt.clf()
plt.hist(np.divide(df["err_EW_Hdelta_from_robo"],df["EW_Hdelta"]))
plt.show()
plt.clf()
plt.hist(np.divide(df["err_EW_Hgamma_from_robo"],df["EW_Hgamma"]))
plt.show()
plt.clf()
plt.hist(np.divide(df["err_EW_Hbeta_from_robo"],df["EW_Hbeta"]))
plt.show()
plt.clf()
plt.hist(np.divide(df["err_EW_Heps_from_robo"],df["EW_Heps"]))
plt.show()
| notebooks_for_development/plot_scaling_balmer_lines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# +
import os
import matplotlib.pyplot as plt
import pandas as pd
# -
DATA_ROOT = "./data/csv"
os.listdir(DATA_ROOT)
# +
resnet18_train_loss_path = os.path.join(DATA_ROOT, "resnet18", "train-loss.csv")
resnet18_val_loss_path = os.path.join(DATA_ROOT, "resnet18", "val-loss.csv")
resnet50_train_loss_path = os.path.join(DATA_ROOT, "resnet50", "train-loss.csv")
resnet50_val_loss_path = os.path.join(DATA_ROOT, "resnet50", "val-loss.csv")
resnet18_train_acc_path = os.path.join(DATA_ROOT, "resnet18", "train-acc.csv")
resnet18_val_acc_path = os.path.join(DATA_ROOT, "resnet18", "val-acc.csv")
resnet50_train_acc_path = os.path.join(DATA_ROOT, "resnet50", "train-acc.csv")
resnet50_val_acc_path = os.path.join(DATA_ROOT, "resnet50", "val-acc.csv")
resnet18_augmentation_train_loss_path = os.path.join(DATA_ROOT, "resnet18-augmentation", "train-loss.csv")
resnet18_augmentation_val_loss_path = os.path.join(DATA_ROOT, "resnet18-augmentation", "val-loss.csv")
resnet50_augmentation_train_loss_path = os.path.join(DATA_ROOT, "resnet50-augmentation", "train-loss.csv")
resnet50_augmentation_val_loss_path = os.path.join(DATA_ROOT, "resnet50-augmentation", "val-loss.csv")
resnet18_augmentation_train_acc_path = os.path.join(DATA_ROOT, "resnet18-augmentation", "train-acc.csv")
resnet18_augmentation_val_acc_path = os.path.join(DATA_ROOT, "resnet18-augmentation", "val-acc.csv")
resnet50_augmentation_train_acc_path = os.path.join(DATA_ROOT, "resnet50-augmentation", "train-acc.csv")
resnet50_augmentation_val_acc_path = os.path.join(DATA_ROOT, "resnet50-augmentation", "val-acc.csv")
resnet18_augmentation2_train_loss_path = os.path.join(DATA_ROOT, "resnet18-augmentation2", "train-loss.csv")
resnet18_augmentation2_val_loss_path = os.path.join(DATA_ROOT, "resnet18-augmentation2", "val-loss.csv")
resnet50_augmentation2_train_loss_path = os.path.join(DATA_ROOT, "resnet50-augmentation2", "train-loss.csv")
resnet50_augmentation2_val_loss_path = os.path.join(DATA_ROOT, "resnet50-augmentation2", "val-loss.csv")
resnet18_augmentation2_train_acc_path = os.path.join(DATA_ROOT, "resnet18-augmentation2", "train-acc.csv")
resnet18_augmentation2_val_acc_path = os.path.join(DATA_ROOT, "resnet18-augmentation2", "val-acc.csv")
resnet50_augmentation2_train_acc_path = os.path.join(DATA_ROOT, "resnet50-augmentation2", "train-acc.csv")
resnet50_augmentation2_val_acc_path = os.path.join(DATA_ROOT, "resnet50-augmentation2", "val-acc.csv")
# +
resnet18_train_acc_df = pd.read_csv(resnet18_train_acc_path)
resnet18_val_acc_df = pd.read_csv(resnet18_val_acc_path)
resnet50_train_acc_df = pd.read_csv(resnet50_train_acc_path)
resnet50_val_acc_df = pd.read_csv(resnet50_val_acc_path)
resnet18_train_loss_df = pd.read_csv(resnet18_train_loss_path)
resnet18_val_loss_df = pd.read_csv(resnet18_val_loss_path)
resnet50_train_loss_df = pd.read_csv(resnet50_train_loss_path)
resnet50_val_loss_df = pd.read_csv(resnet50_val_loss_path)
resnet18_augmentation_train_acc_df = pd.read_csv(resnet18_augmentation_train_acc_path)
resnet18_augmentation_val_acc_df = pd.read_csv(resnet18_augmentation_val_acc_path)
resnet50_augmentation_train_acc_df = pd.read_csv(resnet50_augmentation_train_acc_path)
resnet50_augmentation_val_acc_df = pd.read_csv(resnet50_augmentation_val_acc_path)
resnet18_augmentation_train_loss_df = pd.read_csv(resnet18_augmentation_train_loss_path)
resnet18_augmentation_val_loss_df = pd.read_csv(resnet18_augmentation_val_loss_path)
resnet50_augmentation_train_loss_df = pd.read_csv(resnet50_augmentation_train_loss_path)
resnet50_augmentation_val_loss_df = pd.read_csv(resnet50_augmentation_val_loss_path)
resnet18_augmentation2_train_acc_df = pd.read_csv(resnet18_augmentation2_train_acc_path)
resnet18_augmentation2_val_acc_df = pd.read_csv(resnet18_augmentation2_val_acc_path)
resnet50_augmentation2_train_acc_df = pd.read_csv(resnet50_augmentation2_train_acc_path)
resnet50_augmentation2_val_acc_df = pd.read_csv(resnet50_augmentation2_val_acc_path)
resnet18_augmentation2_train_loss_df = pd.read_csv(resnet18_augmentation2_train_loss_path)
resnet18_augmentation2_val_loss_df = pd.read_csv(resnet18_augmentation2_val_loss_path)
resnet50_augmentation2_train_loss_df = pd.read_csv(resnet50_augmentation2_train_loss_path)
resnet50_augmentation2_val_loss_df = pd.read_csv(resnet50_augmentation2_val_loss_path)
# -
# ## Exp 1
#
# - ResNet18 & ResNet50
# - No augmentation
# +
plt.figure(figsize=(16, 6))
plt.subplot(1, 2, 1)
plt.title("Loss")
plt.xlabel("global step")
plt.ylabel("loss")
plt.ylim(0.5, 2)
plt.plot(
resnet18_train_loss_df["Step"],
resnet18_train_loss_df["Value"],
label="ResNet18 train loss",
marker=".",
)
plt.plot(
resnet18_val_loss_df["Step"],
resnet18_val_loss_df["Value"],
label="ResNet18 val loss",
marker=".",
)
plt.plot(
resnet50_train_loss_df["Step"],
resnet50_train_loss_df["Value"],
label="ResNet50 train loss",
marker=".",
)
plt.plot(
resnet50_val_loss_df["Step"],
resnet50_val_loss_df["Value"],
label="ResNet50 val loss",
marker=".",
)
plt.legend(loc="upper right")
plt.plot()
plt.subplot(1, 2, 2)
plt.title("Accuracy")
plt.xlabel("global step")
plt.ylabel("acc")
plt.ylim(0, 1)
plt.plot(
resnet18_train_acc_df["Step"],
resnet18_train_acc_df["Value"],
label="ResNet18 train acc",
marker=".",
)
plt.plot(
resnet18_val_acc_df["Step"],
resnet18_val_acc_df["Value"],
label="ResNet18 val acc",
marker=".",
)
plt.plot(
resnet50_train_acc_df["Step"],
resnet50_train_acc_df["Value"],
label="ResNet50 train acc",
marker=".",
)
plt.plot(
resnet50_val_acc_df["Step"],
resnet50_val_acc_df["Value"],
label="ResNet50 val acc",
marker=".",
)
plt.legend(loc="lower right")
plt.savefig("01.png")
plt.plot()
# -
# ## Exp 2
#
# - ResNet18 & ResNet50
# - horizontal_flip: true
# - horizontal_flip_rate: 0.5
# - random_rotation: true
# - random_rotation_degrees: 30
# +
plt.figure(figsize=(16, 6))
plt.subplot(1, 2, 1)
plt.title("Loss")
plt.xlabel("global step")
plt.ylabel("loss")
plt.ylim(0.5, 2)
plt.plot(
resnet18_augmentation_train_loss_df["Step"],
resnet18_augmentation_train_loss_df["Value"],
label="ResNet18 train loss",
marker=".",
)
plt.plot(
resnet18_augmentation_val_loss_df["Step"],
resnet18_augmentation_val_loss_df["Value"],
label="ResNet18 val loss",
marker=".",
)
plt.plot(
resnet50_augmentation_train_loss_df["Step"],
resnet50_augmentation_train_loss_df["Value"],
label="ResNet50 train loss",
marker=".",
)
plt.plot(
resnet50_augmentation_val_loss_df["Step"],
resnet50_augmentation_val_loss_df["Value"],
label="ResNet50 val loss",
marker=".",
)
plt.legend(loc="upper right")
plt.plot()
plt.subplot(1, 2, 2)
plt.title("Accuracy")
plt.xlabel("global step")
plt.ylabel("acc")
plt.ylim(0, 1)
plt.plot(
resnet18_augmentation_train_acc_df["Step"],
resnet18_augmentation_train_acc_df["Value"],
label="ResNet18 train acc",
marker=".",
)
plt.plot(
resnet18_augmentation_val_acc_df["Step"],
resnet18_augmentation_val_acc_df["Value"],
label="ResNet18 val acc",
marker=".",
)
plt.plot(
resnet50_augmentation_train_acc_df["Step"],
resnet50_augmentation_train_acc_df["Value"],
label="ResNet50 train acc",
marker=".",
)
plt.plot(
resnet50_augmentation_val_acc_df["Step"],
resnet50_augmentation_val_acc_df["Value"],
label="ResNet50 val acc",
marker=".",
)
plt.legend(loc="lower right")
plt.savefig("02.png")
plt.plot()
# -
# ## Exp 3
#
# - ResNet18 & ResNet50
# - horizontal_flip: true
# - horizontal_flip_rate: 0.5
# - random_rotation: true
# - random_rotation_degrees: 10
# +
plt.figure(figsize=(16, 6))
plt.subplot(1, 2, 1)
plt.title("Loss")
plt.xlabel("global step")
plt.ylabel("loss")
plt.ylim(0.5, 2)
plt.plot(
resnet18_augmentation2_train_loss_df["Step"],
resnet18_augmentation2_train_loss_df["Value"],
label="ResNet18 train loss",
marker=".",
)
plt.plot(
resnet18_augmentation2_val_loss_df["Step"],
resnet18_augmentation2_val_loss_df["Value"],
label="ResNet18 val loss",
marker=".",
)
plt.plot(
resnet50_augmentation2_train_loss_df["Step"],
resnet50_augmentation2_train_loss_df["Value"],
label="ResNet50 train loss",
marker=".",
)
plt.plot(
resnet50_augmentation2_val_loss_df["Step"],
resnet50_augmentation2_val_loss_df["Value"],
label="ResNet50 val loss",
marker=".",
)
plt.legend(loc="upper right")
plt.plot()
plt.subplot(1, 2, 2)
plt.title("Accuracy")
plt.xlabel("global step")
plt.ylabel("acc")
plt.ylim(0, 1)
plt.plot(
resnet18_augmentation2_train_acc_df["Step"],
resnet18_augmentation2_train_acc_df["Value"],
label="ResNet18 train acc",
marker=".",
)
plt.plot(
resnet18_augmentation2_val_acc_df["Step"],
resnet18_augmentation2_val_acc_df["Value"],
label="ResNet18 val acc",
marker=".",
)
plt.plot(
resnet50_augmentation2_train_acc_df["Step"],
resnet50_augmentation2_train_acc_df["Value"],
label="ResNet50 train acc",
marker=".",
)
plt.plot(
resnet50_augmentation2_val_acc_df["Step"],
resnet50_augmentation2_val_acc_df["Value"],
label="ResNet50 val acc",
marker=".",
)
plt.legend(loc="lower right")
plt.savefig("03.png")
plt.plot()
| visualize/visualize.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import pandas as pd
import os
import numpy as np
import datetime
import glob
from pathlib import Path
from sklearn import datasets, linear_model
from sklearn.metrics import mean_squared_error, r2_score
DOWNLOAD_DIR = 'c:/users/MertKeles/scientific computing/homework-scientific-computing/entsoe-data'
# -
# This will import the data, you have to run it to be able to solve the homework.
# +
def read_single_csv_entso_e(file):
return pd.read_csv(file, sep='\t', encoding='utf-16', parse_dates=["DateTime"])
def load_complete_entso_e_data(directory):
pattern = Path(directory) / '*.csv'
files = glob.glob(str(pattern))
if not files:
raise ValueError(f"No files found when searching in {pattern}, wrong directory?")
print(f'Concatenating {len(files)} csv files...')
each_csv_file = [read_single_csv_entso_e(file) for file in files]
data = pd.concat(each_csv_file, ignore_index=True)
data = data.sort_values(by=["AreaName", "DateTime"])
data = data.set_index("DateTime")
print("Loading done.")
return data
power_demand = load_complete_entso_e_data(DOWNLOAD_DIR)
# -
# # Exercise 1 - Calculate the relation of Wednesday average consumption to Sunday average consumption for selected countries
#
# In this exercise, calculate the relation of Wednesday average consumption to Sunday average consumption for the following countries: Austria, Germany, United Kingdom, Spain, Sweden, Italy, Croatia.
#
# (1) First create a variable that contains only power consumption data for these countries. The pandas command ```isin()``` may be very helpful here. Reduce the data to only consider the period 2015-01-01 until 2019-12-31. The lecture slides may contain relevant code here.
#
# (2) Then, group the data by weekday and country (i.e. AreaName). Use ```groupby``` and ```mean```for that purpose.
#
# (3) Calculate for all countries the proportion of Wednesday (day 2) and Sunday (day 6) by dividing the two values.
#
# (4) For which country, this relative value is highest? What could this indicate?
power_demand
power_demand["AreaName"].unique()
filtercountry = power_demand['AreaName'].isin(['Austria', 'Germany', 'United Kingdom', 'Spain', 'Sweden', 'Italy', 'Croatia'])
power_demand_country = power_demand[filtercountry]
power_demand_country_time = power_demand_country['2015-01-01':'2019-12-31']
power_demand_country_time
power_demand_weekday = power_demand_country_time.groupby([power_demand_country_time.index.weekday, 'AreaName']).mean()
power_demand_weekday
# +
power_demand_wednesday = power_demand_weekday.loc[2, 'TotalLoadValue']
power_demand_sunday = power_demand_weekday.loc[6, 'TotalLoadValue']
relation = power_demand_wednesday/power_demand_sunday
relation
# -
More conumption on Wednesdays --> no shops, industry, everything is closed -> religious country
# # Exercise 2 - Calculate the monthly average consumption as deviation from mean consumption
#
# For the same countries as in the above dataset, calculate the monthly mean consumption as deviation from the mean of consumption over the whole time. Plot the curves for all countries.
#
# (1) First create a variable that contains only power consumption data for the selected countries. The pandas command ```isin()``` may be very helpful here. If you did Exercise 1, you can use the same dataset.
#
# (2) Then, aggregate the data by country (i.e. AreaName) and month. Use ```groupby``` and ```mean``` for that purpose. Select the column ```TotalLoadValue``` from the result.
#
# (3) Aggregate the data by country (i..e AreaName) only, i.e. calculate the average consumption by country using ```groupby``` and ```mean```. Select the column ```TotalLoadValue``` from the result.
#
# (4) Divide the result of (2) by (3) and observe how well broadcasting works here.
#
# (5) Use the command ```unstack``` on the result. How does the table look now? Plot the result. If your resulting, unstacked dataframe is called ```result```, you may use ```result.plot()``` to get a nice plot.
#
# (6) How would you explain the difference in the curve between Croatia and Sweden?
#
# +
# filtercountry = power_demand['AreaName'].isin(['Austria', 'Germany', 'United Kingdom', 'Spain', 'Sweden', 'Italy', 'Croatia'])
# power_demand_country = power_demand[filtercountry]
# -
power_demand_month = power_demand_country.groupby([power_demand_country.index.month, 'AreaName']).mean()
power_demand_month
TotalLoadValue1 = power_demand_month['TotalLoadValue']
TotalLoadValue1
# +
average_consumption = power_demand_country.groupby('AreaName').mean()
TotalLoadValue2 = average_consumption['TotalLoadValue']
TotalLoadValue2
# -
monthly_average_consumption = TotalLoadValue1 / TotalLoadValue2
# +
result = monthly_average_consumption.unstack(level=-1)
result.plot()
# -
plt.plot(result)
plt.xlabel('months')
plt.ylabel('monthly average consumtion')
# # Exercise 3 - calculate the hourly average consumption as deviation from mean consumption
#
# Do the same as in exercise 2, but now for the hourly average consumption. I.e. how much is consumed on each of the 24 hours of a day?
#
# Which country has the lowest, which the highest variability? What may be the reason for it?
# +
# filtercountry = power_demand['AreaName'].isin(['Austria', 'Germany', 'United Kingdom', 'Spain', 'Sweden', 'Italy', 'Croatia'])
# power_demand_country = power_demand[filtercountry]
# -
power_demand_hour = power_demand_country.groupby([power_demand_country.index.hour, 'AreaName']).mean()
power_demand_hour
TotalLoadValue3 = power_demand_hour['TotalLoadValue']
# +
# average_consumption = power_demand_country.groupby('AreaName').mean()
# TotalLoadValue2 = average_consumption['TotalLoadValue']
# TotalLoadValue2
# -
hourly_average_consumption = TotalLoadValue3 / TotalLoadValue2
result = hourly_average_consumption.unstack(level=-1)
result.plot()
# # Exercise 4 - Calculate the average load per capita
#
# Below you find a table with population data for our selected countries. You should use it to calculate per capita consumption.
#
# (1) Calculate the average load in all countries using ```groupby``` and ```mean``` and select the column ```TotalLoadValue``` from the result.
#
# (2) Divide the result by the ```Population``` column of the dataframe ```population```. Observe, how broadcasting helps here nicely.
#
# (3) Plot the result. Which country has the highest load, which the lowest? What may be the reason? In which unit is this value? How could we convert it to MWh per year?
# +
population = pd.DataFrame({'Country': ["Austria", "Croatia", "Germany", "Italy", "Spain", "Sweden", "United Kingdom"],
'Population': [8840521, 4087843, 82905782, 60421760, 46796540, 10175214, 66460344]})
population.index = population["Country"]
population
# -
average_load = power_demand_country.groupby('AreaName').mean()
average_load
Total_Load_Value = average_load['TotalLoadValue']
average_load_per_capita = Total_Load_Value / population['Population']
average_load_per_capita
plt.plot(average_load_per_capita)
| homework06-pandas/homework06_haemmerlemert.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# +
import mcnultymod
import pandas as pd
import numpy as np
# import matplotlib.pyplot as plt
# from mpl_toolkits.basemap import Basemap
# import seaborn as sns
# plt.style.use('seaborn')
# # %matplotlib inline
# from geopy.distance import vincenty
import pickle
# -
auctions = pd.read_pickle('../data/auctionsclosed.pkl')
auctions_test = pd.read_pickle('../data/auctionsclosed_test.pkl')
# Performs feature engineering on both the train and test set. **All transforms are fit on the training set, exactly like with Standard Scaler — the test features never "see" their corresponding targets.**
auctions_v2, auctions_test_v2 = mcnultymod.feature_eng([auctions, auctions_test])
auctions_v2.to_pickle("../data/auctionsclosed_v2.pkl")
auctions_test_v2.to_pickle("../data/auctionsclosed_test_v2.pkl")
| 3-detroit-auctions/code/10-mcnulty-feature-engineering-final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sympy import *
from sympy.abc import m,M,l,b,c,g,t
from sympy.physics.mechanics import dynamicsymbols, init_vprinting
th = dynamicsymbols('theta')
x = dynamicsymbols('x')
dth = diff(th)
dx = diff(x)
ddth = diff(dth)
ddx = diff(dx)
init_vprinting()
ddth = (-(1/2)*m*l cos(th)ddth - b*dx +(1/2)*m*l*sin(th)*dth*dx)/((m/12)(3l+l^2))
ddx = (-(1/2)*m*l cos(th)ddth - b*dx +(1/2)*m*l*sin(th)*dth^2)/(M+m)
| notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from sklearn.cross_validation import cross_val_score, StratifiedShuffleSplit
from sklearn.metrics import log_loss
folder_id = 1
fff = 'tmpsubs/' + str(folder_id) + '/'
def convert_class_label_to_int(class_label):
return int(class_label[6:]) - 1
# +
base_names = ['nn_garf1_', 'nn_katz1_', 'nnlasagna3_', 'cb_rf5_',
'nn_dbn2_', 'gb_lr05_4_', 'xgb1_', 'xgb3_']
base_names = ['garfs1_', 'katzs1_', 'lass1_', 'cb_rf5_',
'nn_dbn2_', 'gb_lr05_4_', 'xgb1_', 'kittys1_', 'xgb3_']
# these are the prefixes of the files to be loaded, there are
# 3 for each classifier
tot_subs = len(base_names)
data_names = [[], [], []]
data_names[0] = [fff + name + '1.csv' for name in base_names]
data_names[1] = [fff + name + '2.csv' for name in base_names]
data_names[2] = [fff + name + '3.csv' for name in base_names]
w_shape = [7] * (tot_subs - 1) # the weights of each submission (except the last one) go from 0.0 to 0.35
w_shape = [5, 6, 5, 5, 5, 5, 7, 7] # or we can manually specify the max weight of each one
w_shape.append(3) # we're using 3-fold CV
start_vals = [0.] * (tot_subs - 1)
step_size = 0.05
weight_results_array = np.ones(w_shape) # we store the results in this array
# -
data_dfs = []
for data_fold in data_names:
tmp = []
for i in data_fold:
tmp.append(pd.read_csv(i))
data_dfs.append(tuple(tmp))
df_train = pd.read_csv('raw_data/train.csv')
renamed_labels = [convert_class_label_to_int(i) for i in df_train['target'].values]
df_train['renamed_labels'] = renamed_labels
curr_id = 0
sss = StratifiedShuffleSplit(df_train['renamed_labels'], n_iter=3, test_size=0.2, random_state=42)
for train_index, test_index in sss:
print(curr_id)
it = np.nditer(weight_results_array[..., curr_id], flags=['multi_index'], op_flags=['writeonly'])
labels_test = df_train['renamed_labels'].values[test_index]
while not it.finished:
weights = [start_vals[mi[0]] + mi[1] * step_size for mi in enumerate(it.multi_index)]
if sum(weights) <= 1.:
# weights.append(1. - sum(weights))
pred = data_dfs[curr_id][0].values * weights[0]
for di, w in enumerate(weights[1:]):
pred += w * data_dfs[curr_id][di+1].values
pred += (1. - sum(weights)) * data_dfs[curr_id][-1].values
it[0] = log_loss(labels_test, pred)
it.iternext()
curr_id += 1
summed_w = np.sum(weight_results_array, axis=tot_subs-1) / 3
print(summed_w.shape)
print(np.min(summed_w))
min_arg = np.argmin(summed_w)
min_point = np.unravel_index(min_arg, summed_w.shape)
print(np.min(weight_results_array[...,0]), np.min(weight_results_array[...,1]), np.min(weight_results_array[...,2]))
print(min_point)
min_point = (0, 2, 3, 2, 0, 2, 1, 2)
step_size = 0.05
for j in range(tot_subs-1):
start_vals[j] = max(0., min_point[j] * step_size - 0.03)
weights = [0.] * (tot_subs - 1)
step_size = 0.01
print(start_vals)
w_shape = [7] * (tot_subs - 1)
w_shape.append(3)
weight_results_array = np.ones(w_shape)
curr_id = 0
sss = StratifiedShuffleSplit(df_train['renamed_labels'], n_iter=3, test_size=0.2, random_state=42)
for train_index, test_index in sss:
print(curr_id)
it = np.nditer(weight_results_array[..., curr_id], flags=['multi_index'], op_flags=['writeonly'])
labels_test = df_train['renamed_labels'].values[test_index]
while not it.finished:
weights = [start_vals[mi[0]] + mi[1] * step_size for mi in enumerate(it.multi_index)]
if sum(weights) <= 1.:
# weights.append(1. - sum(weights))
pred = data_dfs[curr_id][0].values * weights[0]
for di, w in enumerate(weights[1:]):
pred += w * data_dfs[curr_id][di+1].values
pred += (1. - sum(weights)) * data_dfs[curr_id][-1].values
it[0] = log_loss(labels_test, pred)
it.iternext()
curr_id += 1
summed_w = np.sum(weight_results_array, axis=tot_subs-1) / 3
print(summed_w.shape)
print(np.min(summed_w))
min_arg = np.argmin(summed_w)
min_point = np.unravel_index(min_arg, summed_w.shape)
last_w = 0.
for j in range(tot_subs-1):
last_w += start_vals[j] + min_point[j] * step_size
print(start_vals[j] + min_point[j] * step_size)
print(1. - last_w)
| best-avgs-new.ipynb |
# # Fisher's method vs. min (after multiple comparison's correction)
# +
from pkg.utils import set_warnings
set_warnings()
import datetime
import time
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from myst_nb import glue as default_glue
from pkg.data import load_network_palette, load_node_palette, load_unmatched
from pkg.io import savefig
from pkg.plot import set_theme
from pkg.stats import stochastic_block_test
from graspologic.simulations import sbm
from tqdm import tqdm
import matplotlib.colors as colors
from scipy.stats import binom, combine_pvalues
from pkg.stats import binom_2samp
import matplotlib.colors as colors
from pathlib import Path
DISPLAY_FIGS = False
FILENAME = "compare_sbm_methods_sim"
def gluefig(name, fig, **kwargs):
savefig(name, foldername=FILENAME, **kwargs)
glue(name, fig, prefix="fig")
if not DISPLAY_FIGS:
plt.close()
def glue(name, var, prefix=None):
savename = f"{FILENAME}-{name}"
if prefix is not None:
savename = prefix + ":" + savename
default_glue(savename, var, display=False)
t0 = time.time()
set_theme()
rng = np.random.default_rng(8888)
network_palette, NETWORK_KEY = load_network_palette()
node_palette, NODE_KEY = load_node_palette()
fisher_color = sns.color_palette("Set2")[2]
min_color = sns.color_palette("Set2")[3]
method_palette = {"fisher": fisher_color, "min": min_color}
GROUP_KEY = "simple_group"
left_adj, left_nodes = load_unmatched(side="left")
right_adj, right_nodes = load_unmatched(side="right")
left_labels = left_nodes[GROUP_KEY].values
right_labels = right_nodes[GROUP_KEY].values
# -
stat, pvalue, misc = stochastic_block_test(
left_adj,
right_adj,
labels1=left_labels,
labels2=right_labels,
method="fisher",
combine_method="fisher",
)
# ## Model for simulations (alternative)
# We have fit a stochastic block model to the left and right hemispheres. Say the
# probabilities of group-to-group connections *on the left* are stored in the matrix
# $B$, so that $B_{kl}$ is the probability of an edge from group $k$ to $l$.
#
# Let $\tilde{B}$ be a *perturbed* matrix of probabilities. We are interested in testing
# $H_0: B = \tilde{B}$ vs. $H_a: ... \neq ...$. To do so, we compare each
# $H_0: B_{kl} = \tilde{B}_{kl}$ using Fisher's exact test. This results in p-values for
# each $(k,l)$ comparison, $\{p_{1,1}, p_{1,2}...p_{K,K}\}$.
#
# Now, we still are after an overall test for the equality $B = \tilde{B}$. Thus, we
# need a way to combine p-values $\{p_{1,1}, p_{1,2}...p_{K,K}\}$ to get an *overall*
# p-value for our test comparing the stochastic block model probabilities. One way is
# Fisher's method; another is to take the
# minimum p-value out of a collection of p-values which have been corrected for multiple
# comparisons (say, via Bonferroni or Holm-Bonferroni).
#
# To compare how these two alternative methods of combining p-values work, we did the
# following simulation:
#
# - Let $t$ be the number of probabilities to perturb.
# - Let $\delta$ represent the strength of the perturbation (see model below).
# - For each trial:
# - Randomly select $t$ probabilities without replacement from the elements of $B$
# - For each of these elements, $\tilde{B}_{kl} = TN(B_{kl}, \delta B_{kl})$ where
# $TN$ is a truncated normal distribution, such that probabilities don't end up
# outside of [0, 1].
# - For each element *not* perturbed, $\tilde{B}_{kl} = B_{kl}$
# - Sample the number of edges from each block under each model. In other words, let
# $m_{kl}$ be the number of edges in the $(k,l)$-th block, and let $n_k, n_l$ be
# the number of edges in the $k$-th and $l$-th blocks, respectively. Then, we have
#
# $$m_{kl} \sim Binomial(n_k n_l, B_{kl})$$
#
# and likewise but with $\tilde{B}_{kl}$ for $\tilde{m}_{kl}$.
# - Run Fisher's exact test to generate a $p_{kl}$ for each $(k,l)$.
# - Run Fisher's method for combining p-values, or take the minimum p-value after
# Bonferroni correction.
# - These trials were repeated for $\delta \in \{0.1, 0.2, 0.3, 0.4, 0.5\}$ and
# $t \in \{25, 50, 75, 100, 125\}$. For each $(\delta, t)$ we ran 100 replicates of the
# model/test above.
# ## P-values under the null
# +
B_base = misc["probabilities1"].values
inds = np.nonzero(B_base)
base_probs = B_base[inds]
n_possible_matrix = misc["possible1"].values
ns = n_possible_matrix[inds]
n_null_sims = 100
RERUN_NULL = False
save_path = Path(
"/Users/bpedigo/JHU_code/bilateral/bilateral-connectome/results/"
"outputs/compare_sbm_methods_sim/null_results.csv"
)
if RERUN_NULL:
null_rows = []
for sim in tqdm(range(n_null_sims)):
base_samples = binom.rvs(ns, base_probs)
perturb_samples = binom.rvs(ns, base_probs)
# test on the new data
def tester(cell):
stat, pvalue = binom_2samp(
base_samples[cell],
ns[cell],
perturb_samples[cell],
ns[cell],
null_odds=1,
method="fisher",
)
return pvalue
pvalue_collection = np.vectorize(tester)(np.arange(len(base_samples)))
n_overall = len(pvalue_collection)
pvalue_collection = pvalue_collection[~np.isnan(pvalue_collection)]
n_tests = len(pvalue_collection)
n_skipped = n_overall - n_tests
row = {
"sim": sim,
"n_tests": n_tests,
"n_skipped": n_skipped,
}
for method in ["fisher", "min"]:
row = row.copy()
if method == "min":
overall_pvalue = min(pvalue_collection.min() * n_tests, 1)
row["pvalue"] = overall_pvalue
elif method == "fisher":
stat, overall_pvalue = combine_pvalues(
pvalue_collection, method="fisher"
)
row["pvalue"] = overall_pvalue
row["method"] = method
null_rows.append(row)
null_results = pd.DataFrame(null_rows)
null_results.to_csv(save_path)
else:
null_results = pd.read_csv(save_path, index_col=0)
# +
from giskard.plot import subuniformity_plot
fig, axs = plt.subplots(1, 2, figsize=(10, 5))
for i, method in enumerate(["fisher", "min"]):
ax = axs[i]
method_null_results = null_results[null_results["method"] == method]
subuniformity_plot(
method_null_results["pvalue"],
ax=ax,
color=method_palette[method],
element="step",
)
ax.set_title(method.capitalize())
gluefig("null_distributions", fig)
# -
# ```{glue:figure} fig:compare_sbm_methods_sim-null_distributions
#
# Distributions of p-values under the null for Fisher's method (left) and the Min method
# (right) from a simulation with 100 resamples under the null. Dotted line indicates
# the CDF of a $Uniform(0,1)$ random variable. The
# p-values in the upper left of each panel is for a 1-sample KS test, where the null is
# that the variable is distributed $Uniform(0,1)$ against the alternative that its CDF
# is larger than that of a $Uniform(0,1)$ random variable (i.e. that it is superuniform).
# Note that both methods appear empirically valid, but Fisher's appears highly conservative.
# ```
# ## P-values under the alternative
# +
n_sims = 100
n_perturb_range = np.linspace(0, 125, 6, dtype=int)[1:]
perturb_size_range = np.round(np.linspace(0, 0.5, 6), decimals=3)[1:]
print(f"Perturb sizes: {perturb_size_range}")
print(f"Perturb number range: {n_perturb_range}")
n_runs = n_sims * len(n_perturb_range) * len(perturb_size_range)
print(f"Number of runs: {n_runs}")
# +
RERUN_SIM = False
save_path = Path(
"/Users/bpedigo/JHU_code/bilateral/bilateral-connectome/results/"
"outputs/compare_sbm_methods_sim/results.csv"
)
if RERUN_SIM:
t0 = time.time()
mean_itertimes = 0
n_time_first = 5
progress_steps = 0.05
progress_counter = 0
last_progress = -0.05
simple_rows = []
example_perturb_probs = {}
for perturb_size in perturb_size_range:
for n_perturb in n_perturb_range:
for sim in range(n_sims):
itertime = time.time()
# just a way to track progress
progress_counter += 1
progress_prop = progress_counter / n_runs
if progress_prop - progress_steps > last_progress:
print(f"{progress_prop:.2f}")
last_progress = progress_prop
# choose some elements to perturb
currtime = time.time()
perturb_probs = base_probs.copy()
choice_indices = rng.choice(
len(perturb_probs), size=n_perturb, replace=False
)
# pertub em
for index in choice_indices:
prob = base_probs[index]
new_prob = -1
while new_prob <= 0 or new_prob >= 1:
new_prob = rng.normal(prob, scale=prob * perturb_size)
perturb_probs[index] = new_prob
if sim == 0:
example_perturb_probs[(perturb_size, n_perturb)] = perturb_probs
perturb_elapsed = time.time() - currtime
# sample some new binomial data
currtime = time.time()
base_samples = binom.rvs(ns, base_probs)
perturb_samples = binom.rvs(ns, perturb_probs)
sample_elapsed = time.time() - currtime
currtime = time.time()
# test on the new data
def tester(cell):
stat, pvalue = binom_2samp(
base_samples[cell],
ns[cell],
perturb_samples[cell],
ns[cell],
null_odds=1,
method="fisher",
)
return pvalue
pvalue_collection = np.vectorize(tester)(np.arange(len(base_samples)))
pvalue_collection = np.array(pvalue_collection)
n_overall = len(pvalue_collection)
pvalue_collection = pvalue_collection[~np.isnan(pvalue_collection)]
n_tests = len(pvalue_collection)
n_skipped = n_overall - n_tests
test_elapsed = time.time() - currtime
# combine pvalues
currtime = time.time()
row = {
"perturb_size": perturb_size,
"n_perturb": n_perturb,
"sim": sim,
"n_tests": n_tests,
"n_skipped": n_skipped,
}
for method in ["fisher", "min"]:
row = row.copy()
if method == "min":
overall_pvalue = min(pvalue_collection.min() * n_tests, 1)
row["pvalue"] = overall_pvalue
elif method == "fisher":
stat, overall_pvalue = combine_pvalues(
pvalue_collection, method="fisher"
)
row["pvalue"] = overall_pvalue
row["method"] = method
simple_rows.append(row)
combine_elapsed = time.time() - currtime
if progress_counter < n_time_first:
print("-----")
print(f"Perturb took {perturb_elapsed:0.3f}s")
print(f"Sample took {sample_elapsed:0.3f}s")
print(f"Test took {test_elapsed:0.3f}s")
print(f"Combine took {combine_elapsed:0.3f}s")
print("-----")
iter_elapsed = time.time() - itertime
mean_itertimes += iter_elapsed / n_time_first
elif progress_counter == n_time_first:
projected_time = mean_itertimes * n_runs
projected_time = datetime.timedelta(seconds=projected_time)
print("---")
print(f"Projected time: {projected_time}")
print("---")
total_elapsed = time.time() - t0
print("Done!")
print(f"Total experiment took: {datetime.timedelta(seconds=total_elapsed)}")
results = pd.DataFrame(simple_rows)
results.to_csv(save_path)
else:
results = pd.read_csv(save_path, index_col=0)
# -
if RERUN_SIM:
fig, axs = plt.subplots(
len(perturb_size_range), len(n_perturb_range), figsize=(20, 20), sharey=True
)
for i, perturb_size in enumerate(perturb_size_range):
for j, n_perturb in enumerate(n_perturb_range):
ax = axs[i, j]
perturb_probs = example_perturb_probs[(perturb_size, n_perturb)]
mask = base_probs != perturb_probs
show_base_probs = base_probs[mask]
show_perturb_probs = perturb_probs[mask]
sort_inds = np.argsort(-show_base_probs)
show_base_probs = show_base_probs[sort_inds]
show_perturb_probs = show_perturb_probs[sort_inds]
sns.scatterplot(
x=np.arange(len(show_base_probs)), y=show_perturb_probs, ax=ax, s=10
)
sns.lineplot(
x=np.arange(len(show_base_probs)),
y=show_base_probs,
ax=ax,
linewidth=1,
zorder=-1,
color="orange",
)
ax.set(xticks=[])
ax.set(yscale="log")
gluefig("example-perturbations", fig)
# +
fisher_results = results[results["method"] == "fisher"]
min_results = results[results["method"] == "min"]
fisher_means = fisher_results.groupby(["perturb_size", "n_perturb"]).mean()
min_means = min_results.groupby(["perturb_size", "n_perturb"]).mean()
mean_diffs = fisher_means["pvalue"] - min_means["pvalue"]
mean_diffs = mean_diffs.to_frame().reset_index()
mean_diffs_square = mean_diffs.pivot(
index="perturb_size", columns="n_perturb", values="pvalue"
)
# v = np.max(np.abs(mean_diffs_square.values))
# fig, ax = plt.subplots(1, 1, figsize=(8, 8))
# sns.heatmap(
# mean_diffs_square,
# cmap="RdBu",
# ax=ax,
# yticklabels=perturb_size_range,
# xticklabels=n_perturb_range,
# square=True,
# center=0,
# vmin=-v,
# vmax=v,
# cbar_kws=dict(shrink=0.7),
# )
# ax.set(xlabel="Number of perturbed blocks", ylabel="Size of perturbation")
# cax = fig.axes[1]
# cax.text(4, 1, "Min more\nsensitive", transform=cax.transAxes, va="top")
# cax.text(4, 0, "Fisher more\nsensitive", transform=cax.transAxes, va="bottom")
# ax.set_title("(Fisher - Min) pvalues", fontsize="x-large")
# DISPLAY_FIGS = True
# gluefig("pvalue_diff_matrix", fig)
# +
fig, axs = plt.subplots(2, 3, figsize=(15, 10))
for i, perturb_size in enumerate(perturb_size_range):
ax = axs.flat[i]
plot_results = results[results["perturb_size"] == perturb_size]
sns.lineplot(
data=plot_results,
x="n_perturb",
y="pvalue",
hue="method",
style="method",
palette=method_palette,
ax=ax,
)
ax.set(yscale="log")
ax.get_legend().remove()
ax.axhline(0.05, color="dimgrey", linestyle=":")
ax.axhline(0.005, color="dimgrey", linestyle="--")
ax.set(ylabel="", xlabel="", title=f"{perturb_size}")
ylim = ax.get_ylim()
if ylim[0] < 1e-25:
ax.set_ylim((1e-25, ylim[1]))
handles, labels = ax.get_legend_handles_labels()
ax.annotate(
0.05,
xy=(ax.get_xlim()[1], 0.05),
xytext=(30, 10),
textcoords="offset points",
arrowprops=dict(arrowstyle="-"),
)
ax.annotate(
0.005,
xy=(ax.get_xlim()[1], 0.005),
xytext=(30, -40),
textcoords="offset points",
arrowprops=dict(arrowstyle="-"),
)
axs.flat[-1].axis("off")
[ax.set(ylabel="p-value") for ax in axs[:, 0]]
[ax.set(xlabel="Number perturbed") for ax in axs[1, :]]
axs[0, -1].set(xlabel="Number perturbed")
axs[0, 0].set_title(f"Perturbation size = {perturb_size_range[0]}")
for i, label in enumerate(labels):
labels[i] = label.capitalize()
axs.flat[-1].legend(handles=handles, labels=labels, title="Method")
gluefig("perturbation_pvalues_lineplots", fig)
# -
# ```{glue:figure} fig:compare_sbm_methods_sim-perturbation_pvalues_lineplots
#
# p-values under the alternative for two different methods for combining p-values:
# [**Fisher's method**](https://en.wikipedia.org/wiki/Fisher%27s_method) (performed on the
# *uncorrected* p-values) and simply taking
# the minimum p-value after [Bonferroni correction](https://en.wikipedia.org/wiki/Bonferroni_correction) (here, called **Min**).
# The alternative is specified by changing the number of probabilities which are perturbed
# (x-axis in each panel) as well as the size of the perturbations which are done
# to each probability (panels show increasing perturbation size). Dotted and dashed
# lines indicate significance thresholds for $\alpha = \{0.05, 0.005\}$, respectively.
# Note that in this simulation, even for large numbers of small perturbations (i.e. upper
# left panel), the Min method has smaller p-values. Fisher's method displays smaller p-values
# than Min only when there are many (>50) large perturbations, but by this point both
# methods yield extremely small p-values.
# ```
# ## Power under the alternative
alpha = 0.05
results["detected"] = 0
results.loc[results[(results["pvalue"] < alpha)].index, "detected"] = 1
# +
fisher_results = results[results["method"] == "fisher"]
min_results = results[results["method"] == "min"]
fisher_means = fisher_results.groupby(["perturb_size", "n_perturb"]).mean()
min_means = min_results.groupby(["perturb_size", "n_perturb"]).mean()
fisher_power_square = fisher_means.reset_index().pivot(
index="perturb_size", columns="n_perturb", values="detected"
)
min_power_square = min_means.reset_index().pivot(
index="perturb_size", columns="n_perturb", values="detected"
)
mean_diffs = fisher_means["detected"] / min_means["detected"]
mean_diffs = mean_diffs.to_frame().reset_index()
ratios_square = mean_diffs.pivot(
index="perturb_size", columns="n_perturb", values="detected"
)
v = np.max(np.abs(mean_diffs_square.values))
# fig, axs = plt.subplots(1, 3, figsize=(12, 4), sharex=True, sharey=True)
from matplotlib.transforms import Bbox
set_theme(font_scale=1.5)
# set up plot
pad = 0.5
width_ratios = [1, pad * 1.2, 10, pad, 10, 1.3 * pad, 10, 1]
fig, axs = plt.subplots(
1,
len(width_ratios),
figsize=(30, 10),
gridspec_kw=dict(
width_ratios=width_ratios,
),
)
fisher_col = 2
min_col = 4
ratio_col = 6
def shrink_axis(ax, scale=0.7):
pos = ax.get_position()
mid = (pos.ymax + pos.ymin) / 2
height = pos.ymax - pos.ymin
new_pos = Bbox(
[
[pos.xmin, mid - scale * 0.5 * height],
[pos.xmax, mid + scale * 0.5 * height],
]
)
ax.set_position(new_pos)
def power_heatmap(
data, ax=None, center=0, vmin=0, vmax=1, cmap="RdBu_r", cbar=False, **kwargs
):
out = sns.heatmap(
data,
ax=ax,
yticklabels=perturb_size_range,
xticklabels=n_perturb_range,
square=True,
center=center,
vmin=vmin,
vmax=vmax,
cbar_kws=dict(shrink=0.7),
cbar=cbar,
cmap=cmap,
**kwargs,
)
ax.invert_yaxis()
return out
ax = axs[fisher_col]
im = power_heatmap(fisher_power_square, ax=ax)
ax.set_title("Fisher's method", fontsize="large")
ax = axs[0]
shrink_axis(ax, scale=0.5)
_ = fig.colorbar(
im.get_children()[0],
cax=ax,
fraction=1,
shrink=1,
ticklocation="left",
)
ax.set_title("Power\n" + r"($\alpha=0.05$)", pad=25)
ax = axs[min_col]
power_heatmap(min_power_square, ax=ax)
ax.set_title("Min method", fontsize="large")
ax.set(yticks=[])
pal = sns.diverging_palette(145, 300, s=60, as_cmap=True)
ax = axs[ratio_col]
im = power_heatmap(np.log10(ratios_square), ax=ax, vmin=-2, vmax=2, center=0, cmap=pal)
# ax.set_title(r'$log_10(\frac{\text{Power}_{Fisher}}{\text{Power}_{Min}})$')
# ax.set_title(
# r"$log_{10}($Fisher power$)$" + "\n" + r" - $log_{10}($Min power$)$",
# fontsize="large",
# )
ax.set(yticks=[])
ax = axs[-1]
shrink_axis(ax, scale=0.5)
_ = fig.colorbar(
im.get_children()[0],
cax=ax,
fraction=1,
shrink=1,
ticklocation="right",
)
ax.text(2, 1, "Fisher more\nsensitive", transform=ax.transAxes, va="top")
ax.text(2, 0.5, "Equal power", transform=ax.transAxes, va="center")
ax.text(2, 0, "Min more\nsensitive", transform=ax.transAxes, va="bottom")
ax.set_title("Log10\npower\nratio", pad=20)
# remove dummy axes
for i in range(len(width_ratios)):
if not axs[i].has_data():
axs[i].set_visible(False)
xlabel = r"# perturbed blocks $\rightarrow$"
ylabel = r"Perturbation size $\rightarrow$"
axs[fisher_col].set(
xlabel=xlabel,
ylabel=ylabel,
)
axs[min_col].set(xlabel=xlabel, ylabel="")
axs[ratio_col].set(xlabel=xlabel, ylabel="")
fig.text(0.09, 0.86, "A)", fontweight="bold", fontsize=50)
fig.text(0.64, 0.86, "B)", fontweight="bold", fontsize=50)
gluefig("relative_power", fig)
# -
# ```{glue:figure} fig:compare_sbm_methods_sim-relative_power
#
# Comparison of power for Fisher's and the Min method. **A)** The power under the
# alternative described in the text for both Fisher's method and the Min method. In both
# heatmaps, the x-axis represents an increasing number of blocks which are perturbed,
# and the y-axis represents an increasing magnitude for each perturbation. **B)** The
# log of the ratio of powers (Fisher's / Min) for each alternative. Note that positive
# (purple) values would represent that Fisher's is more powerful, and negative (green)
# represent that the Min method is more powerful. Notice that the Min method appears
# to have more power for subtler (fewer or smaller perturbations) alternatives, and
# nearly equal power for more obvious alternatives.
# ```
| docs/compare_sbm_methods_sim.ipynb |