code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Microbiome data normalization tutorial
# This is a jupyter notebook example of the different ways to normalize the reads (i.e. TSS, rarefaction, compositionality correction, etc).
# ## Setup
import calour as ca
ca.set_log_level(11)
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib notebook
# ## Load the data
# we use the chronic fatigue syndrome data from:
#
# <NAME>., <NAME>., <NAME>., <NAME>., <NAME>. and <NAME>., 2016.
#
# Reduced diversity and altered composition of the gut microbiome in individuals with myalgic encephalomyelitis/chronic fatigue syndrome.
#
# Microbiome, 4(1), p.30.
# ### Standard load with TSS normalization
# for each sample we normalize to 10000 reads/sample.
#
# This is done by dividing the number of reads of each feature in a sample by the total sum of reads (of all features) in the sample, and then multiplying by the desired number of reads (i.e. 10000).
#
# After this normalization, the sum of (normalized) reads in each sample will be 10000.
#
# This is different from rarefaction, since each feature can have a non-integer number of reads, and less information is thrown away. However, you need to be careful not to have a bias by the original number of reads (mostly in binary methods). dsFDR works fine with this normalization.
#
# Note that we also throw away samples with less than min_reads=1000 reads total (before normalization). This is in order to reduce the discretization effect in samples with low number of reads.
cfs_normalized=ca.read_amplicon('data/chronic-fatigue-syndrome.biom',
'data/chronic-fatigue-syndrome.sample.txt',
normalize=10000,min_reads=1000)
print(cfs_normalized)
# The sum of reads per sample should be 10000
cfs_normalized.get_data(sparse=False).sum(axis=1)
# The original number of reads per sample (before normalization) is stored in the sample_metadata table in the field "_calour_original_abundance"
res=plt.hist(cfs_normalized.sample_metadata['_calour_original_abundance'],50)
plt.xlabel('original number of reads')
plt.ylabel('number of samples')
# ### load with no normalization
# we can load the data without normalizing the reads per sample by setting the parameter `normalize=None`
#
# This is not recommended for typical microbiome experiments since the number of reads per sample is arbitrary and does not reflect the number of bacteria in the sample.
#
# We still chose to remove all samples with less than 1000 reads total.
cfs_not_normalized=ca.read_amplicon('data/chronic-fatigue-syndrome.biom',
'data/chronic-fatigue-syndrome.sample.txt',
normalize=None,min_reads=1000)
cfs_not_normalized.get_data(sparse=False).sum(axis=1)
# ## TSS normalization (`normalize`)
# We can always normalize to constant sum per sample (similar to the `read_amplicon` normaliztion)
tt = cfs_not_normalized.normalize(5000)
tt.get_data(sparse=False).sum(axis=1)
# ## Compositional normalization (`normalize_compositional`)
# In some cases, a plausible biological scenario is that a few bacteria have a very large number of reads. Increase in the frequency of such a bacteria will cause a decrease in the frequencies of all other bacteria (even if in reality their total number remains constant in the sample) since data is normalized to constant sum per sample.
#
# Under the assumption that most bacteria do not change in total number between the samples, we can normalize to constant sum when ignoring the set of high frequency bacteria.
# We will demonstrate using a synthetic example:
#
# In the original dataset we have a few tens of bacteria separating between healthy and sick
dd=cfs_normalized.diff_abundance('Subject','Control','Patient', random_seed=2018)
# ### Effect of a high frequency artificial bacteria
# let's make the first bactertia high frequency only in the Healthy (Control) and not in the Sick (Patient).
#
# And renormalize to 10000 reads/sample
# +
tt=cfs_normalized.copy()
tt.sparse=False
tt.data[tt.sample_metadata['Subject']=='Control',0] = 50000
tt=tt.normalize(10000)
# -
dd=tt.diff_abundance('Subject','Control','Patient', random_seed=2018)
# We get more bacteria which are higher in 'Patient' since bacteria 0 is now very high in controls, and data is TSS normalized
# ### Let's fix by doing the compositional normalization
yy=tt.normalize_compositional()
dd=yy.diff_abundance('Subject','Control','Patient', random_seed=2018)
# so we reduced the inflation of false differentially abundant bacteria due to data compositionallity
# ## Normalization on part of the features (`normalize_by_subset_features`)
#
# Sometimes we want to normalize while ignoring some features (say ignoring all mitochondrial sequences), but we still want to keep these features - just not use them in the normalization.
#
# Note the sum of reads per sample will not be constant (since samples also contain the ignored features).
# Lets ignore the bacteria that don't have a good taxonomy assignment
bad_seqs=[cseq for cseq,ctax in cfs_not_normalized.feature_metadata['taxonomy'].iteritems() if len(ctax)<13]
tt = cfs_not_normalized.normalize_by_subset_features(bad_seqs, total=10000)
tt.get_data(sparse=False).sum(axis=1)
| doc/source/notebooks/microbiome_normalization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# __Chapter 13 - Parallelizing Neural Network Training with TensorFlow__
#
# 1. [Import](#Import)
# 1. [First steps with TensoFlow](#First-steps-with-TensoFlow)
# 1. [Working with array structures](#Working-with-array-structures)
# 1. [Developing a simple model with the low-level TensorFlow API](#Developing-a-simple-model-with-the-low-level-TensorFlow-API)
# 1. [Training neural networks efficiently with high-level TensorFlow APIs](#Training-neural-networks-efficiently-with-high-level-TensorFlow-APIs)
# 1. [Building multilayer neural networks using TensorFlow's Layers API](#Building-multilayer-neural-networks-using-TensorFlows-Layers-API)
# 1. [Developing a multilayer neural network with Keras](#Developing-a-multilayer-neural-network-with-Keras)
# 1. [Choosing activation functions for multilayer networks](#Choosing-activation-functions-for-multilayer-networks)
# 1. [Logistic function recap](#Logistic-function-recap)
# 1. [Estimating class probabilities in multiclass classification via the softmax function](#Estimating-class-probabilities-in-multiclass-classification-via-the-softmax-function)
# 1. [Broadening the output spectrum using a hyperbolic tangent](#Broadening-the-output-spectrum-using-a-hyperbolic-tangent)
# 1. [Rectified linear unit activation](#Rectified-linear-unit-activation)
#
#
#
# # Import
# <a id = 'Import'></a>
# +
# standard libary and settings
import os
import sys
import importlib
import itertools
from io import StringIO
import warnings
warnings.simplefilter("ignore")
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:95% !important; }</style>"))
# data extensions and settings
import numpy as np
np.set_printoptions(threshold=np.inf, suppress=True)
import pandas as pd
pd.set_option("display.max_rows", 500)
pd.options.display.float_format = "{:,.6f}".format
# modeling extensions
from sklearn.base import TransformerMixin, BaseEstimator
from sklearn.cluster import KMeans, DBSCAN, AgglomerativeClustering
from sklearn.datasets import load_boston, load_wine, load_iris, load_breast_cancer, make_blobs, make_moons
from sklearn.decomposition import PCA, LatentDirichletAllocation
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier, AdaBoostClassifier, ExtraTreesClassifier, IsolationForest
from sklearn.feature_extraction.text import CounterVectorizer, TfidfTransformer, TfidfVectorizer, HashingVectorizer
from sklearn.feature_selection import f_classif, f_regression, VarianceThreshold, SelectFromModel, SelectKBest
from sklearn.linear_model import Lasso, Ridge, ElasticNet, LinearRegression, LogisticRegression, SGDRegressor
from sklearn.metrics import precision_score, recall_score, f1_score, explained_variance_score, mean_squared_log_error, mean_absolute_error, median_absolute_error, mean_squared_error, r2_score, confusion_matrix, roc_curve, accuracy_score, roc_auc_score, homogeneity_score, completeness_score, classification_report, silhouette_samples
from sklearn.model_selection import KFold, train_test_split, GridSearchCV, StratifiedKFold, cross_val_score, RandomizedSearchCV
from sklearn.neighbors import KNeighborsClassifier, KNeighborsRegressor
from sklearn.pipeline import make_pipeline, Pipeline, FeatureUnion
from sklearn.preprocessing import StandardScaler, RobustScaler, PolynomialFeatures, OrdinalEncoder, LabelEncoder, OneHotEncoder, KBinsDiscretizer, QuantileTransformer, PowerTransformer, MinMaxScaler
from sklearn.svm import SVC, SVR
from sklearn.tree import DecisionTreeRegressor, DecisionTreeClassifier
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
import sklearn.utils as utils
# visualization extensions and settings
import seaborn as sns
import matplotlib.pyplot as plt
# custom extensions and settings
sys.path.append("/home/mlmachine") if "/home/mlmachine" not in sys.path else None
sys.path.append("/home/prettierplot") if "/home/prettierplot" not in sys.path else None
import mlmachine as mlm
from prettierplot.plotter import PrettierPlot
import prettierplot.style as style
# magic functions
# %matplotlib inline
# -
# # First steps with TensoFlow
#
# TensorFlow is built around a computation graphs composed of a node set. Each node represents and operation that may have zero or more input or output. The values that flow through the edges of the computation graph are referred to as tensors.
#
# Tensors can be scalars, vectors, matrices, and so on. A scalar is a rank-0 tensor, a vector is a rank-1 tensor, a matrix is a rank-2 tensor, and matrices stacked to form a three dimensional array are rank-3 tensors.
#
# We need to build the computation graph and launch it in a TensorFlow session. As a simple warm-up, we start with the use of scalars from TensorFlow to compute a net input $z$ of a sample point $x$ in a 1-dimensional dataset with weight $w$ and bias $b$.
#
# $$
# z = wx + b
# $$
#
# The following code shows how to implement this equation in the lower-level TensorFlow API. In general, the low-level API requires that we define placeholders for input data (x, y and sometimes other tunable parameters). Then we define the weight matrices, and if this is an optimization algorithm we need to define the loss or cost function to use. TensoFlow creates a graph that contains all the sybmols we define as nodes in this graph
# <a id = 'First-steps-with-TensoFlow'></a>
# +
# load TensorFlow and check version
import tensorflow as tf
print(tf.__version__)
# +
# create a graph
g = tf.Graph()
with g.as_default():
x = tf.placeholder(dtype=tf.float32, shape=(None), name="x")
w = tf.Variable(2.0, name="weight")
b = tf.Variable(0.7, name="bias")
z = w * x + b
init = tf.global_variables_initializer()
# create a session and pass in graph g
with tf.Session(graph=g) as sess:
# initialize w and b
sess.run(init)
# evaluate z:
for t in [1.0, 0.6, -1.8]:
print("x = {:.2f} --> {:.2f}".format(t, sess.run(z, feed_dict={x: t})))
# -
# the placeholder for the variable x with shape = (None) allows us to feed the values in an element-by-element form, as well as in batch form where we feed in all the input data at once. The latter implementation would be as follows:
# batch form
with tf.Session(graph=g) as sess:
sess.run(init)
print(sess.run(z, feed_dict={x: [1.0, 2.0, 3.0]}))
# ## Working with array structures
#
# As an example of how to use array structures, we will create a simple rank-3 tensor of size _batchsize_ x 2 x 3, reshape it, and calculate the column sums using TensorFlow's optimized expression. We generally won't know the batch size beforehand, so we specify None for the batch size in the argument for the shape parameter of the placeholder x.
#
#
# <a id = 'Working-with-array-structures'></a>
# +
# create graph and session
g = tf.Graph()
with g.as_default():
x = tf.placeholder(dtype=tf.float32, shape=(None, 2, 3), name="input_x")
x2 = tf.reshape(x, shape=(-1, 6), name="x2")
# calculate the sum of each column
xsum = tf.reduce_sum(x2, axis=0, name="col_sum")
# calculate the mean of each column
xmean = tf.reduce_mean(x2, axis=0, name="col_sum")
with tf.Session(graph=g) as sess:
x_array = np.arange(18).reshape(3, 2, 3)
print("Original array:\n {}".format(x_array))
print("input shape: {}".format(x_array.shape))
print("Reshaped: \n {0}".format(sess.run(x2, feed_dict={x: x_array})))
print("Column sums: \n {0}".format(sess.run(xsum, feed_dict={x: x_array})))
print("Column means: \n {0}".format(sess.run(xmean, feed_dict={x: x_array})))
# -
# We used three functions - tf.reshape, tf.reduce_mean, tf.reduce_sum. The value -1 is used when reshaping because we don't know the batch size. Using -1 in TensorFlow when reshaping means the size of that dimensions will be computed according to the total size of the tensor and the other dimensions. Something like tf.reshape(tensor, shape = (-1,) can be used to flatten a tensor.
# # Developing a simple model with the low-level TensorFlow API
#
# Implement an ordinary least squares regressioin model. Let's create a small 1-dimensional dataset with 10 training samples. We want to train a linear regression model to predict the output $y$ from the input $x$, per usual. We will implement this model in a class that we will call TfLinreg. We need two placeholders, one for the input $x$ and one for the output $y$ in order to feed the data into ourmodel. We also need the trainable variables $w$ and $b$ for the weight and bias.
#
# With these, we can then define the linear regression model $z= w \times x + b$, as well as the cost function. For this function we will use Mean Squared Error. To learn the weights, we will use gradient descent.
# <a id = 'Developing-a-simple-model-with-the-low-level-TensorFlow-API'></a>
# create train and test sets
X_train = np.arange(10).reshape((10, 1))
y_train = np.array([1.0, 1.3, 3.1, 2.0, 5.0, 6.3, 6.6, 7.4, 8.0, 9.0])
# custom class implementing linear regression with TensorFlow
class TfLinReg:
def __init__(self, x_dim, learning_rate=0.01, random_seed=None):
self.x_dim = x_dim
self.learning_rate = learning_rate
self.g = tf.Graph()
# build model
with self.g.as_default():
# set graph-level random-seed
tf.set_random_seed(random_seed)
self.build()
# create initializer
self.init_op = tf.global_variables_initializer()
def build(self):
# define placeholders for inputs
self.X = tf.placeholder(
dtype=tf.float32, shape=(None, self.x_dim), name="x_input"
)
self.y = tf.placeholder(dtype=tf.float32, shape=(None), name="y_input")
print(self.X)
print(self.y)
# define weight matrix and bias vector
w = tf.Variable(tf.zeros(shape=(1)), name="weight")
b = tf.Variable(tf.zeros(shape=(1)), name="bias")
print(w)
print(b)
self.z_net = tf.squeeze(w * self.X + b, name="z_net")
print(self.z_net)
sqr_errors = tf.square(self.y - self.z_net, name="sqr_errors")
print(sqr_errors)
self.mean_cost = tf.reduce_mean(sqr_errors, name="mean_cost")
optimizer = tf.train.GradientDescentOptimizer(
learning_rate=self.learning_rate, name="GradientDescent"
)
self.optimizer = optimizer.minimize(self.mean_cost)
# fit model
lrmodel = TfLinReg(x_dim=X_train.shape[1], learning_rate=0.01)
# The print statements display information about the six nodes in the graph $X$, $y$, $w$, $b$, $z\_net$, $sqr\_errors$. Now we need to implement a training function to learn the weights of the linear regression model. To train the model, we implement a separate function that requires a TensorFlow session, model instance, training data and the number of epochs. In the following function, we initialize the variables in the TensorFlow session using the init_op operation defined in the mode. Then we iterate and call the optimizer operation of the model while feeding the training data. This functions returns a list of training costs.
# custom function for capturing training costs
def train_linreg(sess, model, X_train, y_train, num_epochs=10):
# initialize all variables: W and b
sess.run(model.init_op)
training_costs = []
for i in range(num_epochs):
_, cost = sess.run(
[model.optimizer, model.mean_cost],
feed_dict={model.X: X_train, model.y: y_train},
)
training_costs.append(cost)
return training_costs
# +
# create a new TensorFlwo session to launch the lrmodel.g graph
sess = tf.Session(graph=lrmodel.g)
# pass all required arguments to the train_linreg function for training
training_costs = train_linreg(sess, lrmodel, X_train, y_train)
# -
np.arange(1, len(training_costs) + 1)
# visualize training cost by epoch
p = PrettierPlot()
ax = p.make_canvas(
title="Cost reduction over time",
x_label="Epochs",
y_label="Training cost",
y_shift=0.6,
)
p.line(
x=np.arange(1, len(training_costs) + 1),
y=np.array(training_costs),
label="training cost",
linecolor=style.style_hex_mid[0],
x_ticks=np.arange(0, 11, 1),
bbox=(1.2, 0.9),
marker_on=True,
ax=ax,
)
# We need to add a function to make predictions based on input features. This function will need the TensorFlow session, model and the test data.
# function to make predictions. Simply run z_net in the previously defined graph
def predict_linreg(sess, model, X_test):
y_pred = sess.run(model.z_net, feed_dict={model.X: X_test})
return y_pred
# +
# visualize line of best fit
p = PrettierPlot()
ax = p.make_canvas(title="Basic linear regression", x_label="X", y_label="y", y_shift=0.95)
p.scatter_2d(x=X_train, y=y_train, x_units="f", y_units="f", ax=ax)
p.line(
x=np.arange(X_train.shape[0]),
y=predict_linreg(sess, lrmodel, X_train),
label="Line of best fit",
linecolor=style.style_hex_mid[0],
bbox=(1.2, 0.9),
marker_on=False,
ax=ax,
)
# -
# # Training neural networks efficiently with high-level TensorFlow APIs
#
# We will review two high-level TensorFlow APIs, the Layers API and Keras API
# <a id = 'Training-neural-networks-efficiently-with-high-level-TensorFlow-APIs'></a>
# ## Building multilayer neural networks using TensorFlow's Layers API
# <a id = 'Building-multilayer-neural-networks-using-TensorFlows-Layers-API'></a>
# +
# Load data and print dimensions
df_train = pd.read_csv("s3://tdp-ml-datasets/kaggle-mnist//train.csv", sep=",")
df_test = pd.read_csv("s3://tdp-ml-datasets/kaggle-mnist//test.csv", sep=",")
print("Training data dimensions: {}".format(df_train.shape))
print("Test data dimensions: {}".format(df_test.shape))
# separate
df_train_label = df_train["label"]
df_train = df_train.drop(labels="label", axis=1)
# train/test split
X_train, X_test, y_train, y_test = train_test_split(
df_train, df_train_label, test_size=0.2
)
X_train = X_train.values
X_test = X_test.values
y_train = y_train.values
y_test = y_test.values
# -
# split into train/test data
print("Training data dimensions: {}".format(X_train.shape))
print("Test data dimensions: {}".format(X_test.shape))
# standardize data
mean_vals = np.mean(X_train, axis=0)
std_val = np.std(X_train)
X_train_centered = (X_train - mean_vals) / std_val
X_test_centered = (X_test - mean_vals) / std_val
print(
"X_train_centered dimensions: {}, training label shape: {}".format(
X_train_centered.shape, y_train.shape
)
)
print(
"X_test_centered dimensions: {}, training label shape: {}".format(
X_test_centered.shape, y_test.shape
)
)
# With this data in hand, we will start building the model by creating two placeholders, named tf_x and tf_y, and then build an MLP with three fully connected layers. For the activation functions, we will use hyperbolic tangent activation functions (tanh) in the hidden layer, and in the output layer we will use softmax. We will also add an additional hidden layer to get us to three fully connected layers.
# +
# create TensorFlow graph
n_features = X_train_centered.shape[1]
n_classes = 10
random_seed = 123
np.random.seed(random_seed)
g = tf.Graph()
with g.as_default():
tf.set_random_seed(random_seed)
tf_x = tf.placeholder(dtype=tf.float32, shape=(None, n_features), name="tf_x")
tf_y = tf.placeholder(dtype=tf.int32, shape=None, name="tf_y")
y_onehot = tf.one_hot(indices=tf_y, depth=n_classes)
h1 = tf.layers.dense(inputs=tf_x, units=50, activation=tf.tanh, name="layer1")
h2 = tf.layers.dense(inputs=h1, units=50, activation=tf.tanh, name="layer2")
logits = tf.layers.dense(inputs=h2, units=10, activation=None, name="layer3")
predictions = {
"classes": tf.argmax(logits, axis=1, name="predicted_classes"),
"probabilities": tf.nn.softmax(logits, name="softmax_tensor"),
}
with g.as_default():
cost = tf.losses.softmax_cross_entropy(onehot_labels=y_onehot, logits=logits)
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.001)
train_op = optimizer.minimize(loss=cost)
init_op = tf.global_variables_initializer()
# -
# In order to train the network, we need a way to generate batches of data. To do this, we implement a function that returns a generator.
# custom function to generate samples for model
def create_batch_generator(X, y, batch_size=128, shuffle=False):
X_copy = np.array(X)
y_copy = np.array(y)
if shuffle:
data = np.column_stack((X_copy, y_copy))
np.random.shuffle(data)
X_copy = data[:, :-1]
y_copy = data[:, -1].astype(int)
for i in range(0, X.shape[0], batch_size):
yield (X_copy[i : i + batch_size, :], y_copy[i : i + batch_size])
# +
# create a new TensorFlow session, initialize variables in our network and train
# create a new session to launch the graph
sess = tf.Session(graph=g)
# run the variable initialization operator
sess.run(init_op)
# 50 epochs of training
for epoch in range(50):
training_costs = []
batch_generator = create_batch_generator(X_train_centered, y_train, batch_size=64)
for batch_X, batch_y in batch_generator:
# prepare a dict to feed data to the network
feed = {tf_x: batch_X, tf_y: batch_y}
_, batch_cost = sess.run([train_op, cost], feed_dict=feed)
training_costs.append(batch_cost)
print("Epoch {0} training loss: {1}".format(epoch + 1, training_costs[-1]))
# -
# lastly, make predictions with the trained model
feed = {tf_x: X_test_centered}
y_pred = sess.run(predictions["classes"], feed_dict=feed)
print("Test accuracy: {:.4f}%".format(100 * np.sum(y_pred == y_test) / y_test.shape[0]))
# # Developing a multilayer neural network with Keras
# <a id = 'Developing-a-multilayer-neural-network-with-Keras'></a>
# +
# load keras
import tensorflow.contrib.keras as keras
np.random.seed(123)
tf.set_random_seed(123)
# -
# one-hot encode traiing data
y_train_onehot = keras.utils.to_object(y_train)
print("First 5 labels: {}".format(y_train[:5]))
print("\nFirst 5 labels one-hot encoded\n".format(y_train_onehot[:5]))
# Build a neural network with three layers, where the first two layers each ahve 50 hidden units with a tanh activation function, and the last layer has 10 layers for each of the 10 class labels and uses softmax to give the probability of each class.
#
# First we initialize a new model using the Sequential class to implement a feedforward neural network. Then we can add as many layers as we want to. The kernel_initializer 'glorot uniform' is a more robust way of initializing weights for deep neural networks.
#
# We also define an optimizer, which is gradient descent. Binary cross-entropy is merely a technical term for the cost function in logistc regression, and the object cross-entropy is its generalization for multiclass predictions via softmax.
#
# Once the model is compiled, we can train it by calling the fit method. In the implementation below we use a batch size of 64 and 50 epochs. The validation_split parameter automatically reserves 10 percent of the training data for validation in each epoch so that can monitor if the model is overfitting during training.
# +
# create keras model
model = keras.models.Sequential()
model.add(
keras.layers.Dense(
units=50,
input_dim=X_train_centered.shape[1],
kernel_initializer="glorot_uniform",
bias_initializer="zeros",
activation="tanh",
)
)
model.add(
keras.layers.Dense(
units=50,
input_dim=50,
kernel_initializer="glorot_uniform",
bias_initializer="zeros",
activation="tanh",
)
)
model.add(
keras.layers.Dense(
units=y_train_onehot.shape[1],
input_dim=50,
kernel_initializer="glorot_uniform",
bias_initializer="zeros",
activation="softmax",
)
)
sgd_optimizer = keras.optimizers.SGD(lr=0.001, decay=1e-7, momentum=0.9)
model.compile(optimizer=sgd_optimizer, loss="object_crossentropy")
# -
# fit model
history = model.fit(
X_train_centered,
y_train_onehot,
batch_size=64,
epochs=50,
verbose=1,
validation_split=0.1,
)
# predict class labels
y_train_pred = model.predict_classes(X_train_centered, verbose=0)
print("First 5 predictions: {0}".format(y_train_pred[:5]))
# +
# generate predictions and evaluate accuracy
y_train_pred = model.predict_classes(X_train_centered, verbose=0)
correct_preds = np.sum(y_train == y_train_pred, axis=0)
train_acc = correct_preds / y_train.shape[0]
print("First 5 predictions: {0}".format(y_train_pred[:5]))
print("Training accuracy: {0}".format(train_acc))
y_test_pred = model.predict_classes(X_test_centered, verbose=0)
correct_preds = np.sum(y_test == y_test_pred, axis=0)
test_acc = correct_preds / y_test.shape[0]
print("First 5 predictions: {0}".format(y_test_pred[:5]))
print("Test accuracy: {0}".format(test_acc))
# -
# # Choosing activation functions for multilayer networks
#
# To this point in the book, we have only discussed the sigmoid activation function in the context of MLPs. there are many alternative sigmoidal functions that are useful for implementin gin MLPs.
#
# We can technically use an function as an activation function as long as it is differentiable. We could even use a linear activation function such as Adaline. Linear function are not useful in practice within hidden and output layers however since we want to introduce nonlinearity in a typical neural network to be able to tackle complex problems.
#
# The logistic activation funciton covered previously can be problematic if we have highly negative input since the output of the sigmoid function would be close to zero in this case. If the sigmoid function returns outputs that are close to zero, then the neural network will learn very slowly and it becomes more likely that it will get trapped in the local minima during training.
# <a id = 'Choosing-activation-functions-for-multilayer-networks'></a>
# ## Logistic function recap
#
# In a binary classification problem, the logistic function is used to model the probability that sample $x$ belongs to the positive class (class 1). the net input $z$ is shown in the following equation:
#
# $$
# z = w_0x_0 + w_1x_1 + ... + w_mx_m = \sum^m_{i=0}w_ix_i = w^Tx
# $$
#
# In this equation $w_0$ is the bias unit (the y-intercept, which means $x_0$ = 1. And this net input is put into the logistic function:
#
# $$
# \phi{logistic}(z) = \frac{1}{1+e^{-z}}
# $$
# <a id = 'Logistic-function-recap'></a>
# +
# basic logistic regression example
X = np.array([1.0, 1.4, 2.5]) # first value must be 1
w = np.array([0.4, 0.3, 0.5])
def net_input(X, w):
return np.dot(X, w)
def logistic(z):
return 1.0 / (1.0 + np.exp(-z))
def logistic_activation(X, w):
z = net_input(X, w)
return logistic(z)
print("P(y = w| x) = {:.3f}".format(logistic_activation(X, w)))
# -
# > Remarks - In the context of a neural network, we would calculate the net input and use it to activate a logistic neuron, we get a value of 0.888, meaning there is an 88.8 percent probability that this particular sample $x$ belongs to the positive class.
# In chapter 12, one-hot encoding to compute the values in the output layer consisting of multiple logistic activation units. However, an output layer consisting of multiple logistic activation units does not produce meaningful, interpretable probability values.
# +
# W : array with shape = (n_output_units, n_hidden_units + 1)
# the first column values are the bias units
W = np.array([[1.1, 1.2, 0.8, 0.4], [0.2, 0.4, 1.0, 0.2], [0.6, 1.5, 1.2, 0.7]])
# A : data array with shape = (n_hidden_units + 1, n_samples)
# the first column of this array must be 1
A = np.array([[1, 0.1, 0.4, 0.6]])
Z = np.dot(W, A[0])
y_probas = logistic(Z)
print("Net input: {}".format(Z))
print("Output units: {}".format(y_probas))
# -
# > Remarks - The resulting values cannot be interpreted as probabilities of a three class problem because the values do not sum to 1. This is not a problem if our only goal is to predict the class labels and we do not care about the class membership probabilities. In this scenario, we just need to observe the index of the highest value.
# generate prediction
y_class = np.argmax(Z, axis=0)
print("Predicted class label: {:d}".format(y_class))
# ## Estimating class probabilities in multiclass classification via the softmax function
#
# If simplys obtaining the class label through the argmax function isn't enough, we can use the softmax function. It is a soft form of the argmax function that provides the probability of each class rather than a single class index. With this tool, we can compute meaningful class probabilities in a multiclass setting.
#
# The probability of a particular sample with net input $z$ belonging the the $i$th class can be computed with a normalization term in the denominator. This is the sum of all $M$ linear functions:
#
# $$
# p(y = i|z) = \phi(z) = \frac{e^{z_i}}{\sum^M_{i=1}e^{z_j}}
# $$
# <a id = 'Estimating-class-probabilities-in-multiclass-classification-via-the-softmax-function'></a>
# +
# example probabilities using softmax
def softmax(z):
return np.exp(z) / np.sum(np.exp(z))
y_probas = softmax(Z)
print("Probabilities: {}".format(y_probas))
print(np.sum(y_probas))
# -
# ## Broadening the output spectrum using a hyperbolic tangent
#
# The hyperbolic tangent, commonly referred to as tanh, is another sigmoid function that is often used in the hidden layers of neural networks. It is a rescaled version of the logistic functions:
#
# $$
# \phi_{tanh}(z) = 2 \times \phi_{logistic}(2z) - 1 = \frac{e^z - e^{-z}}{e^z + e^{-z}}
# $$
#
# The advantage of the hyuperbolic tangent over the logistic function is that is has a broder output spectrum that ranges over the open interval (-1,1), which can improve the convergence of the back propagation algorithms. This contrasts with the logistic function which returns an output signal that ranges over the interval (0,1)
# <a id = 'Broadening-the-output-spectrum-using-a-hyperbolic-tangent'></a>
# ## Rectified linear unit activation
#
# The rectified linear unit, or ReLU, is anohter activation function that is often used in deep neural networks. This activation may help to address the vanishing gradient problem of tanh and logistic activations. This problem occurs for very large values of $z$. As an example, the activaiton of $z_1$ = 20 and $z_2$ = 25 are both about equal to 1.0 on the logistic and tanh curves. This effectively shows no change in output in the eyes of these activation functions. This means that the derivative of the activations with respect to the net input diminishes as $z$ becomes large. As a result, learing weights during the training phase becomes very slow because the gradient terms may be very close to zero. ReLU addresses this issues. It is mathematically defined as:
#
# $$
# \phi(z) = max(0,z)
# $$
#
# It is still a nonlinear function that is effective at learning complex functions within neural networks. Furhter, the derivative of ReLU, with respect to its input, is always 1 for positive input values. Therefore, it solves the problem of vanishing gradients, making it suitable for deep neural networks.
# <a id = 'Rectified-linear-unit-activation'></a>
| textbooks/PythonMachineLearning/ch13_Parallelizing_Neural_Network_Training_with_TensorFlow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [<img src="https://mybinder.org/badge_logo.svg">](https://mybinder.org/v2/gh/NumEconCopenhagen/NumEconNotebooks/master?urlpath=lab/tree/micro/04_Substitution_income_and_wealth_effects.ipynb)
# # Setup
# + jupyter={"source_hidden": true}
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
prop_cycle = plt.rcParams['axes.prop_cycle']
colors = prop_cycle.by_key()['color']
from mpl_toolkits.mplot3d import Axes3D
from mpl_toolkits.mplot3d.art3d import Poly3DCollection, Line3DCollection
import ipywidgets as widgets
import numecon.micro.consumption as consumption
# -
# # Static: Decomposition with exogenous income
# The budget set is
#
# $$
# C(p_1,p_2,I) = \{(x_1,x_2) \in \mathbb{R}_{+}^2 \,\, | \,\, p_1 x_1 + p_2 x_2 \leq I\}
# $$
#
# We normalize with $p_2 = 1$ and consider a change in $p_1$ to $p_1^{\prime}$.
# +
consumer = consumption.ConsumerClass(preferences='cobb_douglas')
fig,ax = consumer.figure()
A,B,C = consumer.plot_decomposition_exogenous(ax,p1_old=1,p1_new=2,p2=1)
print(f'A = ({A[0]:5.2f},{A[1]:5.2f})')
print(f'B = ({B[0]:5.2f},{B[1]:5.2f})')
print(f'C = ({C[0]:5.2f},{C[1]:5.2f})')
# -
# # Static: Decomposition with endogenous income
# The budget set is
#
# $$
# C(p_1,p_2,e_1,e_2) = \{(x_1,x_2) \in \mathbb{R}_{+}^2 \,\, | \,\, p_1 x_1 + p_2 x_2 \leq p_1 e_1 + p_2 e_2\}
# $$
#
# We normalize with $p_2 = 1$ and consider a change in $p_1$ to $p_1^{\prime}$.
# +
consumer = consumption.ConsumerClass(preferences='cobb_douglas')
fig,ax = consumer.figure()
A,B,C1,C2 = consumer.plot_decomposition_endogenous(ax,p1_old=1,p1_new=2,p2=1,e1=6,e2=4)
print(f'A = ({A[0]:5.2f},{A[1]:5.2f})')
print(f'B = ({B[0]:5.2f},{B[1]:5.2f})')
print(f'C1 = ({C1[0]:5.2f},{C1[1]:5.2f})')
print(f'C2 = ({C2[0]:5.2f},{C2[1]:5.2f})')
# -
# # Interactive: Cobb-Douglas
consumption.interactive_slutsky_exogenous(preferences='cobb_douglas')
consumption.interactive_slutsky_endogenous(preferences='cobb_douglas')
# # Interactive: CES
consumption.interactive_slutsky_exogenous(preferences='ces')
consumption.interactive_slutsky_endogenous(preferences='ces')
# # Deadweight loss
# ## Integration
# + jupyter={"source_hidden": true}
# a. calculations
x = np.linspace(1e-8,10,1000)
y = 1/x
I = x <= 1
# a. figure
fig = plt.figure(figsize=(6,4),dpi=100)
ax = fig.add_subplot(1,1,1)
ax.plot(x,y,color='black',lw=2)
ax.fill_between(x[I],y[I],np.ones(np.sum(I)),color='black',alpha=0.3)
ax.set_xlim([0,3])
ax.set_ylim([0,5])
ax.set_xlabel('x')
ax.set_ylabel('y');
# -
# ## Midpoint approximation of integral
# + jupyter={"source_hidden": true}
import numpy as np
import matplotlib.pyplot as plt
# a. interval
x0 = 1
x1 = 2
z0 = 1.2
z1 = 1.6
# b. functions
f = lambda x: np.log(x)
f_prime = lambda x: 1/x
# c. points
x = np.linspace(1,2,1000)
y = f(x)
# d. box
width = z1-z0
midpoint = (z0+z1)/2
height = f(z0) + f_prime(z0)*0.5*width
# e. figure
fig = plt.figure(figsize=(6,4),dpi=100)
ax = fig.add_subplot(1,1,1)
ax.plot(x,y,lw=2,color='navy')
ax.plot([z0,z0,z1,z1],[0,height,height,0],color='firebrick')
ax.plot(x,f(z0)+f_prime(z0)*(x-z0),color='black')
ax.plot([midpoint,midpoint],[0,height],color='black',ls='--')
ax.scatter(z0,0,s=5,color='black')
ax.text(z0*0.97,0.01,'$z_0$',fontsize=10)
ax.scatter(z1,0,s=5,color='black')
ax.text(z1*1.01,0.01,'$z_1$',fontsize=10)
ax.scatter(z0,f(z0),s=5,color='black')
ax.text(z0*0.94,f(z0),'$f(z_0)$',fontsize=10)
ax.text(1.5,0.7,'$f(z_0)+f^{\\prime}(z_0)(x-z_0)$',fontsize=10)
I = (x >= z0) & (x <= z1)
ax.fill_between(x[I],y[I],color='black',alpha=0.25)
ax.set_xlim([1,2])
ax.set_ylim([0.0,0.8])
# f. box
width = z1-z0
midpoint = (z0+z1)/2
height = f(z1) - f_prime(z1)*0.5*width
# g. figure
fig = plt.figure(figsize=(6,4),dpi=100)
ax = fig.add_subplot(1,1,1)
ax.plot(x,y,lw=2,color='navy')
ax.plot([z0,z0,z1,z1],[0,height,height,0],color='firebrick')
ax.plot(x,f(z1)+f_prime(z1)*(x-z1),color='black')
ax.plot([midpoint,midpoint],[0,height],color='black',ls='--')
ax.scatter(z0,0,s=5,color='black')
ax.text(z0*0.97,0.01,'$z_0$',fontsize=10)
ax.scatter(z1,0,s=5,color='black')
ax.text(z1*1.01,0.01,'$z_1$',fontsize=10)
ax.scatter(z1,f(z1),s=5,color='black')
ax.text(z1*0.95,f(z1),'$f(z_1)$',fontsize=10)
ax.text(1.55,0.65,'$f(z_1)+f^{\\prime}(z_1)(x-z_1)$',fontsize=10)
I = (x >= z0) & (x <= z1)
ax.fill_between(x[I],y[I],color='black',alpha=0.25)
ax.set_xlim([1,2])
ax.set_ylim([0.0,0.8]);
| micro/04_Substitution_income_and_wealth_effects.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
with open('day_05_input.txt', 'r') as inputfile:
fabric = inputfile.readlines()
code = fabric[0][:-1]
print(len(code))
# +
# ----- Part 1 -----
finished = False
while not finished:
for idx in range(len(code)):
try:
if (ord(code[idx]) == ord(code[idx+1])+32) or (ord(code[idx])+32 == ord(code[idx+1])):
temp = code[:idx] + code[idx+2:]
code = temp
break
except:
pass
finished = idx == len(code) - 1
print(len(code))
# +
----- Part 2 -----
import time
with open('day_05_input.txt', 'r') as inputfile:
fabric = inputfile.readlines()
for letter in string.ascii_uppercase:
starttime = time.time()
code = fabric[0][:-1]
newcode = code.replace(letter, "")
code = newcode.replace(chr(ord(letter)+32), "")
assert len(code) == (50000 - letters[letter] - letters[letter.upper()])
finished = False
while not finished:
for idx in range(len(code)):
try:
if (ord(code[idx]) == ord(code[idx+1])+32) or (ord(code[idx])+32 == ord(code[idx+1])):
temp = code[:idx] + code[idx+2:]
code = temp
break
except:
pass
finished = idx == len(code) - 1
print(f'Letter {letter} Length:{len(code)} {time.time()-starttime}')
# 5524
# -
| day_05_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "-"}
# # Exploring BFS
#
# First, include some libraries
# +
# Run boilerplate code to set up environment
# %run ../prelude.py
# -
# ## Graph Inputs
# +
#
# Function to create graph inputs
#
def create_inputs(display=True):
# Adjacency matrix - Ranks "S" (source) and "D" (destination)
a = Tensor.fromUncompressed([ "S", "D"],
[ [ 0, 1, 1, 0, 0, 0 ],
[ 0, 0, 1, 1, 0, 0 ],
[ 0, 0, 0, 1, 1, 0 ],
[ 0, 0, 0, 0, 1, 1 ],
[ 1, 0, 0, 0, 0, 1 ],
[ 1, 1, 0, 0, 0, 0 ] ])
# Fringe (current) - Rank "V" (either source or destination)
f0 = Tensor.fromUncompressed([ "V" ], [ 1, 0, 0, 0, 0, 0 ])
# Distance - Rank "V" (either source or destination)
d = Tensor.fromUncompressed([ "V" ], [1, 0, 0, 0, 0, 0])
print("Adjacency Matrix")
displayTensor(a)
print("Distance Vector")
displayTensor(d)
print("Current Fringe")
displayTensor(f0)
return (a, f0, d)
# -
# # Naive BFS - source stationary (push)
#
# This version traverses all neighbors of each source node, even if there already is a distance. So there is a check to not create a new distance.
# +
# Create inputs
(a, f0, d) = create_inputs()
# Setup for traversing roots
# Get root fibers
a_s = a.getRoot()
f0_s = f0.getRoot()
d_d = d.getRoot()
# +
level = 2
while (f0_s.countValues() > 0):
print("\n\n")
print(f"Level {level} fringe")
displayTensor(f0_s.nonEmpty())
print(f"Level {level} distances")
displayTensor(d_d)
# Create a new next fringe (f1)
f1 = Tensor(rank_ids=[ "V" ])
f1_d = f1.getRoot()
# For each source in fringe get destinations (neighbors)
for s, (_, a_d) in f0_s & a_s:
print(f"Processing source {s}")
print(f"Neighbors:\n {a_d}")
# For each neighboring destination
# prepare to update distance and next fringe
for d, (f1_d_ref, (d_d_ref, _)) in f1_d << (d_d << a_d):
print(f" Processing destination {d} = {d_d_ref}")
# Only update distance and fringe for "empty" destinations,
# i.e., without a distance (unvisited)
if Payload.isEmpty(d_d_ref):
print(f" Adding destination {d}")
# Update next fringe
f1_d_ref += 1
# Update destination's distance
d_d_ref += level
# Move to next level
level += 1
# Copy next fringe to current fringe
f0 = f1
f0_s = f0.getRoot()
print("\n\n")
print("Final Distances")
displayTensor(d_d)
# -
# ## Optimized BFS - source stationary (push)
#
# Avoid processing of any destination node that already has a distance by subtracting the distance array from the neighbors
# +
# Create inputs
(a, f0, d) = create_inputs()
# Setup for traversing roots
# Get root fibers
a_s = a.getRoot()
f0_s = f0.getRoot()
d_d = d.getRoot()
# +
level = 2
while (f0_s.countValues() > 0):
print("\n\n")
print(f"Level {level} fringe")
displayTensor(f0_s.nonEmpty())
print(f"Level {level} distances")
displayTensor(d_d)
# Create a new next fringe (f1)
f1 = Tensor(rank_ids=[ "D" ])
f1_d = f1.getRoot()
# For each source in fringe get destinations (neighbors)
for s, (_, a_d) in f0_s & a_s:
print(f"Processing source {s}")
print(f"Neighbors:\n {a_d}")
# For each neighboring destination without a distance
# prepare to update distance and next fringe
for d, (f1_d_ref, (d_d_ref, _)) in f1_d << (d_d << (a_d - d_d)):
print(f" Processing destination {d} = {d_d_ref}")
print(f" Adding destination {d}")
# Update next frige (note no "if" statement)
f1_d_ref += 1
# Update destination's distance
d_d_ref += level
# Move to next level
level += 1
# Copy next fringe to current fringe
f0 = f1
f0_s = f0.getRoot()
print("\n\n")
print("Final Distances")
displayTensor(d_d)
# -
# ### Destination stationary BFS (pull)
# +
# Create inputs
(a, f0, d) = create_inputs()
# Setup for traversing roots
# Get root fibers
a_s = a.getRoot()
f0_s = f0.getRoot()
d_d = d.getRoot()
# Transpose the adjacency matrix
at_d = a_s.swapRanks()
print("Transposed adjaceny matrix")
at = Tensor.fromFiber(["D", "S"], at_d)
displayTensor(at)
# +
# Destination Stationary
iteration = 1
while (f0_s.countValues() > 0):
print("\n\n")
print(f"Iteration {iteration} fringe")
displayTensor(f0_s.nonEmpty())
print(f"Iteration {iteration} distances")
displayTensor(d_d)
# Create a new next fringe (f1)
f1 = Tensor(rank_ids=[ "V" ])
f1_d = f1.getRoot()
# For destinations without a distance get incoming neighbors
# and prepare for updates to distances and next fringe
for d, (f1_d_ref, (d_d_ref, at_s)) in f1_d << (d_d << (at_d - d_d)):
print(f"Processing destination {d}")
print(f"Incoming neighbors:\n {at_s}")
# For incoming sources in fringe with a distance
# pick any source to assign a distance (we use the first)
# Note, because all the sources are in the fringe they have the same distance!
for s, ((_, _), d_s_val) in (at_s & f0_s) & d_d:
print(f" Processing source {s} = {d_s_val}")
print(f" Adding destination {d}")
assert d_s_val != 0
print(f"Debug fringe {f0_s!r}")
# Update next fringe
f1_d_ref += 1
# Update destination's distance
d_d_ref += d_s_val + 1
break
# Move to next iteration
iteration += 1
# Copy next fringe to current fringe
f0 = f1
f0_s = f0.getRoot()
print("\n\n")
print("Final Distances")
displayTensor(d_d)
# -
# ## First source then destination stationary BFS - Push Pull
# +
# Create inputs
(a, f0, d) = create_inputs()
# Setup for traversing roots
# Get root fibers
a_s = a.getRoot()
f0_s = f0.getRoot()
d_d = d.getRoot()
# +
# Destination stationary (push) stage
level = 2
while (f0_s.countValues() > 0 and d_d.countValues() < 3):
print("\n\n")
print(f"Level {level} fringe")
displayTensor(f0_s.nonEmpty())
print(f"Level {level} distances")
displayTensor(d_d)
# Create a new next fringe (f1)
f1 = Tensor(rank_ids=[ "D" ])
f1_d = f1.getRoot()
# For each source in fringe get destinations (neighbors)
for s, (_, a_d) in f0_s & a_s:
print(f"Processing source {s}")
print(f"Neighbors:\n {a_d}")
# For each neighboring destination without a distance
# prepare to update distance and next fringe
for d, (f1_d_ref, (d_d_ref, _)) in f1_d << (d_d << (a_d - d_d)):
print(f" Processing destination {d} = {d_d_ref}")
print(f" Adding destination {d}")
# Update next frige (note no "if" statement)
f1_d_ref += 1
# Update destination's distance
d_d_ref += level
# Move to next level
level += 1
# Copy next fringe to current fringe
f0 = f1
f0_s = f0.getRoot()
print("\n\n")
print("Final destination stationary fringe")
displayTensor(f0)
print("Final destination stationary distances")
displayTensor(d_d)
# +
# Prepare for destination stations part
# Transpose the adjacency matrix
at_d = a_s.swapRanks()
print("Transposed adjaceny matrix")
at = Tensor.fromFiber(["D", "S"], at_d)
displayTensor(at)
# +
# Destination Stationary
iteration = 1
while (f0_s.countValues() > 0):
print("\n\n")
print(f"Iteration {iteration} fringe")
displayTensor(f0_s.nonEmpty())
print(f"Iteration {iteration} distances")
displayTensor(d_d)
# Create a new next fringe (f1)
f1 = Tensor(rank_ids=[ "V" ])
f1_d = f1.getRoot()
# For destinations without a distance get incoming neighbors
# and prepare for updates to distances and next fringe
for d, (f1_d_ref, (d_d_ref, at_s)) in f1_d << (d_d << (at_d - d_d)):
print(f"Processing destination {d}")
print(f"Incoming neighbors:\n {at_s}")
# For incoming sources in fringe with a distance
# pick any source to assign a distance (we use the first)
# Note, because all the sources are in the fringe they have the same distance!
xxx = at_s & f0_s
print(f"Debug intersection: {xxx!r}")
for s, ((_, _), d_s_val) in (at_s & f0_s) & d_d:
print(f" Processing source {s} = {d_s_val}")
print(f" Adding destination {d}")
assert d_s_val != 0
print(f"Debug fringe {f0_s!r}")
# Update next fringe
f1_d_ref += 1
# Update destination's distance
d_d_ref += d_s_val + 1
break
# Move to next iteration
iteration += 1
# Copy next fringe to current fringe
f0 = f1
f0_s = f0.getRoot()
print("\n\n")
print("Final Distances")
displayTensor(d_d)
# -
# ## Testing area
#
# For running alternative algorithms
| notebooks/graphs/bfs.ipynb |
# ##### Copyright 2021 Google LLC.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# # strimko2
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/google/or-tools/blob/master/examples/notebook/contrib/strimko2.ipynb"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/colab_32px.png"/>Run in Google Colab</a>
# </td>
# <td>
# <a href="https://github.com/google/or-tools/blob/master/examples/contrib/strimko2.py"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/github_32px.png"/>View source on GitHub</a>
# </td>
# </table>
# First, you must install [ortools](https://pypi.org/project/ortools/) package in this colab.
# !pip install ortools
# +
# Copyright 2010 <NAME> <EMAIL>
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""Strimko problem in Google CP Solver.
From
360: A New Twist on Latin Squares
http://threesixty360.wordpress.com/2009/08/04/a-new-twist-on-latin-squares/
'''
The idea is simple: each row and column of an nxn grid must contain
the number 1, 2, ... n exactly once (that is, the grid must form a
Latin square), and each "stream" (connected path in the grid) must
also contain the numbers 1, 2, ..., n exactly once.
'''
For more information, see:
* http://www.strimko.com/
* http://www.strimko.com/rules.htm
* http://www.strimko.com/about.htm
* http://www.puzzlersparadise.com/Strimko.htm
I have blogged about this (using MiniZinc model) in
'Strimko - Latin squares puzzle with "streams"'
http://www.hakank.org/constraint_programming_blog/2009/08/strimko_latin_squares_puzzle_w_1.html
Compare with the following models:
* MiniZinc: http://hakank.org/minizinc/strimko2.mzn
* ECLiPSe: http://hakank.org/eclipse/strimko2.ecl
* SICStus: http://hakank.org/sicstus/strimko2.pl
* Gecode: http://hakank.org/gecode/strimko2.cpp
This model was created by <NAME> (<EMAIL>)
See my other Google CP Solver models: http://www.hakank.org/google_or_tools/
"""
import sys
from ortools.constraint_solver import pywrapcp
# Create the solver.
solver = pywrapcp.Solver('Strimko')
#
# default problem
#
if streams == '':
streams = [[1, 1, 2, 2, 2, 2, 2], [1, 1, 2, 3, 3, 3, 2],
[1, 4, 1, 3, 3, 5, 5], [4, 4, 3, 1, 3, 5, 5],
[4, 6, 6, 6, 7, 7, 5], [6, 4, 6, 4, 5, 5, 7],
[6, 6, 4, 7, 7, 7, 7]]
# Note: This is 1-based
placed = [[2, 1, 1], [2, 3, 7], [2, 5, 6], [2, 7, 4], [3, 2, 7], [3, 6, 1],
[4, 1, 4], [4, 7, 5], [5, 2, 2], [5, 6, 6]]
n = len(streams)
num_placed = len(placed)
print('n:', n)
#
# variables
#
x = {}
for i in range(n):
for j in range(n):
x[i, j] = solver.IntVar(1, n, 'x[%i,%i]' % (i, j))
x_flat = [x[i, j] for i in range(n) for j in range(n)]
#
# constraints
#
# all rows and columns must be unique, i.e. a Latin Square
for i in range(n):
row = [x[i, j] for j in range(n)]
solver.Add(solver.AllDifferent(row))
col = [x[j, i] for j in range(n)]
solver.Add(solver.AllDifferent(col))
#
# streams
#
for s in range(1, n + 1):
tmp = [x[i, j] for i in range(n) for j in range(n) if streams[i][j] == s]
solver.Add(solver.AllDifferent(tmp))
#
# placed
#
for i in range(num_placed):
# note: also adjust to 0-based
solver.Add(x[placed[i][0] - 1, placed[i][1] - 1] == placed[i][2])
#
# search and solution
#
db = solver.Phase(x_flat, solver.INT_VAR_DEFAULT, solver.INT_VALUE_DEFAULT)
solver.NewSearch(db)
num_solutions = 0
while solver.NextSolution():
for i in range(n):
for j in range(n):
print(x[i, j].Value(), end=' ')
print()
print()
num_solutions += 1
solver.EndSearch()
print()
print('num_solutions:', num_solutions)
print('failures:', solver.Failures())
print('branches:', solver.Branches())
print('WallTime:', solver.WallTime(), 'ms')
| examples/notebook/contrib/strimko2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Pipeline Preparation
#
# This Jupyter Notebook demonstrates how data pipeline is prepared.
# * text will be extracted from PDF files and saved into SQLite database
# ## Import Libraries
# +
import numpy as np
import pandas as pd
from glob import glob
# Python package to extract text from pdf
from pdfminer.high_level import extract_text
# -
# Save file path in numpy array
# load filenames for pdf files
pdf_files = np.array(glob("pdf/*"))
# number of files in the 'PDF' folder
len(pdf_files)
# ## Extract text from PDF files
# +
# create empty dictionary called text_dict
text_dict = {}
# loop through the filenames, extract text from PDF files and save to the dictionary
for file in pdf_files:
text = extract_text(file)
text_dict[file] = text
# -
# convert dictionary to pandas DataFrame
df = pd.DataFrame(list(text_dict.items()), columns = ['file_path', "raw_text"])
# File paths and raw text from PDF files in PDF folder have been saved to a dataframe
df
# ## Save Data
#
# Save dataset into a SQLite database
# +
# import SQLAlchemy library
from sqlalchemy import create_engine
engine = create_engine('sqlite:///Text.db')
df.to_sql('Text_table', engine, if_exists = 'replace', index=False)
# -
| data/Data Pipeline Preparation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
from sklearn.metrics import log_loss
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.datasets import make_hastie_10_2
from sklearn.model_selection import train_test_split
# -
# Use this sigmoid function to turn probabilities into classifications
def sigmoid(x):
return 1 / (1 + np.exp(-x))
# A simple synthetic dataset is created below. It has a 1D normal distribution, the goal is to predict Class '1' if the item is positive and '0' otherwise.
# Generate an array of 5000 random numbers that are normally distributed
X_all = np.random.randn(5000, 1)
# Based on whether the item is greater or less than zero, the mask will return true or false. The result is multiplied by 2 and 1 is subtracted to convert booleans into 1 or 0
y_all = (X_all[:, 0] > 0)*2 - 1
# Use sci-kit learn's famous [train-test-split](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) function. Half the data is used for training data and the remaining hald if used for testing purposes.
X_train, X_test, y_train, y_test = train_test_split(X_all, y_all, test_size=0.5, random_state=42)
# You will notice this dataset can be solved with one tree stump! Thus, we will choose max depth as 1
# +
clf = DecisionTreeClassifier(max_depth=1)
clf.fit(X_train, y_train)
print ('Accuracy for a single decision stump: {}'.format(clf.score(X_test, y_test)))
# -
# The Decision Tree Classifier only needed 1 stump but for the Gradient Boosting Classifier, we will need 800 trees to classify the data correctly. We shall use sklearn's GradientBoostingClassifier
clf = GradientBoostingClassifier(n_estimators=5000, learning_rate=0.01, max_depth=3, random_state=0)
clf.fit(X_train, y_train)
print('Accuracy for Gradient Booing: {}'.format(clf.score(X_test, y_test)))
# The predict_proba method gives the probabilities on a data-point belonging to a class. As our metric, we will use the logloss method from sklearn. Find more about it [here](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.log_loss.html)
y_pred = clf.predict_proba(X_test)[:, 1]
print("Test logloss: {}".format(log_loss(y_test, y_pred)))
# Some helper functions below
# +
def compute_loss(y_true, scores_pred):
'''
Since we use raw scores we will wrap log_loss
and apply sigmoid to our predictions before computing log_loss itself
'''
return log_loss(y_true, sigmoid(scores_pred))
'''
Get cummulative sum of *decision function* for trees. i-th element is a sum of trees 0...i-1.
We cannot use staged_predict_proba, since we want to manipulate raw scores
(not probabilities). And only in the end convert the scores to probabilities using sigmoid
'''
cum_preds = np.array([x for x in clf.staged_decision_function(X_test)])[:, :, 0]
print ("Logloss using all trees: {}".format(compute_loss(y_test, cum_preds[-1, :])))
print ("Logloss using all trees but last: {}".format(compute_loss(y_test, cum_preds[-2, :])))
print ("Logloss using all trees but first: {}".format(compute_loss(y_test, cum_preds[-1, :] - cum_preds[0, :])))
# -
# While there is a LogLoss difference, it is not as significant as you would expect. If you get rid of the first tree, the model still works! Let's take a plot of the cummulative decision function depending on the number of trees to understand this further
# +
# Pick an object of class 1 for visualisation
plt.figure(figsize=(8, 6))
plt.plot(cum_preds[:, y_test == 1][:, 0], color='r')
plt.xlabel('n_trees')
plt.ylabel('Cumulative decision score')
plt.title('Score vs. Trees')
plt.savefig('plots/score_vs_trees.png', transparent=False)
plt.show()
# -
# You can notice that the decision function improves ~ linearly till about the 800 iteration and then stops. This is actually connected to the learning rate of 0.01 which we set!
#
# The main formula of boosting is:
# $$ F(x) = const + \sum\limits_{i=1}^{n}\gamma_i h_i(x) $$
#
# In our case, $\gamma_i$ are constant and equal to learning rate $\eta = 0.01$. It takes about $800$ iterations to get the score $8$, which means at every iteration score goes up for about $0.01$. It means that first 800 terms are approximately equal to $0.01$, and the following are almost $0$.
#
# So if we drop the last tree, we lower $F(x)$ by $0$ and if we drop the first tree we lower $F(x)$ by $0.01$, which results in a very very little performance drop!
#
# So, even in the case of simple dataset which can be solved with single decision stump, with a Gradient Boosting Machine we need to sum a lot of trees (roughly $\frac{1}{\eta}$) to approximate this golden single decision stump.
# **What if we use a larger learning rate?** Let's use the learning rate of 10
clf = GradientBoostingClassifier(n_estimators=5000, learning_rate=10, max_depth=3, random_state=0)
clf.fit(X_train, y_train)
y_pred = clf.predict_proba(X_test)[:, 1]
print("Test logloss: {}".format(log_loss(y_test, y_pred)))
print('Accuracy for High Learning Rate GBM: {}'.format(clf.score(X_test, y_test)))
# +
cum_preds = np.array([x for x in clf.staged_decision_function(X_test)])[:, :, 0]
print ("Logloss using all trees: {}".format(compute_loss(y_test, cum_preds[-1, :])))
print ("Logloss using all trees but last: {}".format(compute_loss(y_test, cum_preds[-2, :])))
print ("Logloss using all trees but first: {}".format(compute_loss(y_test, cum_preds[-1, :] - cum_preds[0, :])))
# -
# As you can see, the significance of the first tree in a GBM is clear. The logloss without the first tree is much higher! Let's plot the score vs. trees like we did earlier
# +
# Pick an object of class 1 for visualisation
plt.figure(figsize=(8, 6))
plt.plot(cum_preds[:, y_test == 1][:, 0])
plt.xlabel('n_trees')
plt.ylabel('Cumulative decision score')
plt.title('Score vs. Trees')
plt.savefig('plots/score_vs_trees_high_learn.png', transparent=False)
plt.show()
# -
from xgboost import XGBClassifier
xg_model = XGBClassifier(n_estimators=5000, learning_rate=0.01)
xg_model.fit(X_train, y_train)
print('Accuracy for XGBoost: {}'.format(xg_model.score(X_train, y_train)))
| Gradient-Boosting/Gradient-Boosting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/lauramenicacci/MLProject_KenyaFinancial/blob/main/RandomForest_Final.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="LWQ7N7E6pvSk"
#Import packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import pickle
from sklearn import tree
from sklearn import preprocessing
from sklearn import metrics
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
from sklearn.metrics import roc_auc_score
# + colab={"base_uri": "https://localhost:8080/"} id="yRVwHCIIBHHW" outputId="6cb32162-ae85-4ada-8fae-5e1933a83bb4"
#Mount Drive
from google.colab import drive
drive.mount('/content/drive')
# %cd /content/drive/MyDrive/MLProject_KenyaFinancial
# + id="R3GDnkJeBHFF"
#X and Y paths based on data
x_path = "/content/drive/MyDrive/MLProject_KenyaFinancial/clean_data/XoheImp.csv"
y_path = "/content/drive/MyDrive/MLProject_KenyaFinancial/clean_data/Y.csv"
# + id="Zbll08QgBHCK"
X = pd.read_csv(x_path)
y = pd.read_csv(y_path)
# + id="N325WzsZBG_p" colab={"base_uri": "https://localhost:8080/"} outputId="2a00eead-3c2e-4c3e-eaf5-191143a6186d"
#Split training and validation set
# %run Splitting_data.ipynb
X_train, X_val, y_train, y_val = training_set(x_path, y_path)
# + id="Uh2QvHXVBG9J"
#Random Forest (50 trees)
# + colab={"base_uri": "https://localhost:8080/"} id="REGKOZwKBG2m" outputId="a7d2d4e4-a950-40dd-c44a-70937e1922a5"
clf = RandomForestClassifier(n_estimators = 50)
clf.fit(X_train, y_train)
# + id="qenpcrQgBGoE"
y_pred_val50 = clf.predict(X_val)
# + colab={"base_uri": "https://localhost:8080/"} id="ptVGRE42BUBu" outputId="21bbba49-fb57-4440-b91b-eea4ace3f899"
#Print Random Forest (50) results
print("Accuracy:",metrics.accuracy_score(y_val, y_pred_val50))
print("F1:",metrics.f1_score(y_val, y_pred_val50, average="weighted"))
print("Matthew's Correlation Coefficient:",metrics.matthews_corrcoef(y_val, y_pred_val50))
# + id="mwpC5n3RJhaJ"
with open('RandomForest_Final', 'wb') as files:
pickle.dump(clf, files)
# + id="G5c6S6tLB_A8"
#Label encoder for ROC AUC
from sklearn import preprocessing
le = preprocessing.LabelEncoder()
# + id="nVTq2ESoyxWA"
trans_y_pred_val50 = le.fit(y_pred_val50)
# + id="sa9HuzYTCPar"
roc_auc_score(y_val, trans_y_pred_val50, average = "macro", multi_class = "ovr")
# + id="H5BYVLIZIaNX"
#Random Forest (100 trees)
# + id="c2QCwuimItJx"
forest100 = RandomForestClassifier(n_estimators = 100)
forest100.fit(X_train, y_train)
# + id="3AAft4MlIwVC"
y_pred_val100 = forest100.predict(X_val)
# + id="cMN0ptOvIyFH"
#Print Random Forest (100) Results
print("Accuracy:",metrics.accuracy_score(y_val, y_pred_val100))
print("F1:",metrics.f1_score(y_val, y_pred_val100, average="weighted"))
print("Matthew's Correlation Coefficient:",metrics.matthews_corrcoef(y_val, y_pred_val100))
# + colab={"base_uri": "https://localhost:8080/", "height": 599} id="ITCDgGkrlDvR" outputId="d66076d0-c30f-4291-c36e-02c4994dfd9b"
#Variable importance graph
importances = forest50.feature_importances_
std = np.std([tree.feature_importances_ for tree in forest50.estimators_], axis=0)
forest_importances = pd.Series(importances, index=X.columns)
fig, ax = plt.subplots()
forest_importances.plot.bar(yerr=std, ax=ax)
ax.set_title("Feature importances using Random Forest")
ax.set_ylabel("Mean decrease in impurity")
fig.tight_layout()
# + id="2EJGjv5ypyGU"
from sklearn.metrics import confusion_matrix
import seaborn as sns
# + colab={"base_uri": "https://localhost:8080/", "height": 469} id="gJH7M_nHpfjI" outputId="2aa6dbc4-65ed-492d-f6ad-9a002a139bf8"
# Create a confusion matrix
cnf_matrix = confusion_matrix(y_val, y_pred_val50)
# Create heatmap from the confusion matrix
# %matplotlib inline
fig, ax = plt.subplots(figsize=(7, 6))
sns.heatmap(pd.DataFrame(cnf_matrix), annot=True, cmap="YlGnBu", fmt='g')
ax.xaxis.set_label_position("top")
plt.tight_layout()
plt.title('Confusion matrix')
plt.ylabel('Actual label')
plt.xlabel('Predicted label')
tick_marks = [0.5, 1.5]
# + id="stFZ_lktIyum"
#Score tabulation
# + id="nAkmDCMKI1Bv"
#Must fill in values
Scores = {}
Scores[50] = {"Accuracy" : 0.985181492099104, "F1" : 0.9848746348842544, "MCC" : 0.9352177479090404}
# + id="F0rzjV6NI3NF"
Scores[100] = {"Accuracy": 0.9852438129267246, "F1": 0.9849617541345709, "MCC": 0.9355360606936755}
# + id="osL9uEUsI5bo"
ScoresDF = pd.DataFrame([[50, 0.985181492099104, 0.9848746348842544, 0.9352177479090404],[100, 0.9852438129267246, 0.9849617541345709, 0.9355360606936755]], columns = ["Trees", "Accuracy", "F1", "MCC"])
# + id="zjSnZCp3I-cd"
ScoresDF.to_csv("treeScores.csv", sep=";")
# + id="sQ8ZxJo5JA8S"
plt.plot(ScoresDF.Trees, ScoresDF.Accuracy, label = "Accuracy")
plt.plot(ScoresDF.Trees, ScoresDF.F1, label = "F1 score")
plt.plot(ScoresDF.Trees, ScoresDF.MCC, label = "Matthew's Correlation Coefficient")
plt.legend()
plt.show()
# + id="v-Ebe8lmJwIB"
# Define Parameters
#max_depth=[2, 8, 16]
n_estimators = [25, 50, 100]
param_grid = dict(n_estimators=n_estimators)
# + id="dnkY4q0tJ_ef"
from sklearn.model_selection import GridSearchCV
# + colab={"base_uri": "https://localhost:8080/"} id="4gnMqGzhJwBH" outputId="de7b9d17-de65-4f9a-d221-e6c2889ffc85"
# Build the grid search
dfrst = RandomForestClassifier(n_estimators=n_estimators)
grid = GridSearchCV(estimator=dfrst, param_grid=param_grid, cv = 5)
grid_results = grid.fit(X_train, y_train)
# + colab={"base_uri": "https://localhost:8080/", "height": 278} id="oEWepKLzJzbS" outputId="e694cc0c-08ee-4557-a142-46e336e382dd"
# Summarize the results in a readable format
print("Best: {0}, using {1}".format(grid_results.cv_results_['mean_test_score'], grid_results.best_params_))
results_df = pd.DataFrame(grid_results.cv_results_)
results_df
| code/RandomForest_Final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn.datasets import fetch_openml
mnist = fetch_openml('mnist_784',version=1)
mnist.keys()
X, y = mnist['data'], mnist['target']
# +
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
some_digit = X[0]
some_digit_image = some_digit.reshape(28,28)
plt.imshow(some_digit_image, cmap=mpl.cm.binary)
plt.axis('off')
plt.show()
# -
y[0]
# tenemos que cambiar los objetivos de string a numero
# +
import numpy as np
import pandas as pd
y = y.astype(np.int8)
y[0]
# -
X.shape
# Sabemos que este data set ya no trae dividido el train del test, los primeros 60k de instancias son para el train, las ultimas 10k son para el train
X_train, X_test, y_train, y_test = X[:60000], X[:-10000], y[:60000], y[:-10000]
# +
from sklearn.metrics import confusion_matrix, precision_score, recall_score, precision_recall_curve
from sklearn.metrics import f1_score, roc_curve, roc_auc_score
from sklearn.neighbors import KNeighborsClassifier
kng_cla = KNeighborsClassifier()
kng_cla.fit(X_train,y_train)
# -
y_knn_pred = kng_cla.predict(X_train)
TPR = precision_score(y_train, y_knn_pred)
TPR
# +
from sklearn.metrics import accuracy_score
accuracy = accuracy_score(y_train, y_knn_pred)
accuracy
# -
from scipy.ndimage.interpolation import shift
def shift_image(image, dx, dy):
image = image.reshape((28, 28))
shifted_image = shift(image, [dy, dx], cval=0, mode="constant")
return shifted_image.reshape([-1])
# +
X_train_augmented = [image for image in X_train]
y_train_augmented = [label for label in y_train]
for dx, dy in ((1, 0), (-1, 0), (0, 1), (0, -1)):
for image, label in zip(X_train, y_train):
X_train_augmented.append(shift_image(image, dx, dy))
y_train_augmented.append(label)
X_train_augmented = np.array(X_train_augmented)
y_train_augmented = np.array(y_train_augmented)
# -
X_train_augmented.shape()
| Capitulo 3 - Exercises .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualization with Bitbrains Data
#
# # Data Science Consulting Project
#
# ### Modeling System Resource Usage for Predictive Scheduling
# +
# Import packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import os
import glob
from pandas import read_csv, datetime
from pandas.tools.plotting import autocorrelation_plot
from dateutil.relativedelta import relativedelta
from scipy.optimize import minimize
import statsmodels.formula.api as smf
import statsmodels.tsa.api as smt
import statsmodels.api as sm
import scipy.stats as scs
from sklearn.linear_model import LassoCV, RidgeCV
from itertools import product
from tqdm import tqdm_notebook
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
# -
# ## Read in data
# +
path = r'rnd/2013-7/' # use your path
all_files = glob.glob(os.path.join(path, "*.csv")) # advisable to use os.path.join as this makes concatenation OS independent
df_from_each_file = (pd.read_csv(f, sep = ';\t').assign(VM=os.path.basename(f).split('.')[0]) for f in all_files)
concatenated_df = pd.concat(df_from_each_file)
# +
path = r'rnd/2013-8/' # use your path
all_files = glob.glob(os.path.join(path, "*.csv")) # advisable to use os.path.join as this makes concatenation OS independent
df_from_each_file = (pd.read_csv(f, sep = ';\t').assign(VM=os.path.basename(f).split('.')[0]) for f in all_files)
concatenated_df8 = pd.concat(df_from_each_file)
# +
path = r'rnd/2013-9/' # use your path
all_files = glob.glob(os.path.join(path, "*.csv")) # advisable to use os.path.join as this makes concatenation OS independent
df_from_each_file = (pd.read_csv(f, sep = ';\t').assign(VM=os.path.basename(f).split('.')[0]) for f in all_files)
concatenated_df9 = pd.concat(df_from_each_file)
# -
# ## Create Data Frame
newdat = concatenated_df.append(concatenated_df8)
newerdat = newdat.append(concatenated_df9)
concatenated_df = newerdat
# Check it out
concatenated_df.head()
# ## Feature engineering and converting pandas into a timeseries
# ### Timestamp is in UNIX epochs
# +
concatenated_df['Timestamp'] = pd.to_datetime(concatenated_df['Timestamp [ms]'], unit = 's')
concatenated_df.apply(pd.to_numeric, errors='ignore')
# Date Feature Engineering
concatenated_df['weekday'] = concatenated_df['Timestamp'].dt.dayofweek
concatenated_df['weekend'] = ((concatenated_df.weekday) // 5 == 1).astype(float)
concatenated_df['month']=concatenated_df.Timestamp.dt.month
concatenated_df['day']=concatenated_df.Timestamp.dt.day
concatenated_df.set_index('Timestamp',inplace=True)
# Other Feature Engineering
concatenated_df["CPU usage prev"] = concatenated_df['CPU usage [%]'].shift(1)
concatenated_df["CPU_diff"] = concatenated_df['CPU usage [%]'] - concatenated_df["CPU usage prev"]
concatenated_df["received_prev"] = concatenated_df['Network received throughput [KB/s]'].shift(1)
concatenated_df["received_diff"] = concatenated_df['Network received throughput [KB/s]']- concatenated_df["received_prev"]
concatenated_df["transmitted_prev"] = concatenated_df['Network transmitted throughput [KB/s]'].shift(1)
concatenated_df["transmitted_diff"] = concatenated_df['Network transmitted throughput [KB/s]']- concatenated_df["transmitted_prev"]
# -
# ## Fill in missing values using forward propagating function from pandas
concatenated_df = concatenated_df.fillna(method='ffill')
# ## Create new data frame: resampled & aggregated over each hour for all VMs
hourlydat = concatenated_df.resample('H').sum()
# ## Examine autocorrelations of hourly CPU usage
## Hourly resampled means
plt.figure(figsize=(15,5))
pd.plotting.autocorrelation_plot(hourlydat['CPU usage [MHZ]']);
# ## Is CPU Capacity Ever Met? If so, how often?
# +
overprovision = pd.DataFrame(hourlydat['CPU usage [MHZ]'])
overprovision['CPU capacity provisioned'] = pd.DataFrame(hourlydat['CPU capacity provisioned [MHZ]'])
plt.style.use('seaborn-white')
overprovision.plot(figsize = (12,10),linewidth=2.5, fontsize=20)
plt.title('Is CPU Capacity Ever Met?',fontsize=22)
plt.ylabel((r'CPU [MHz] $e^{7}$'), fontsize=20);
plt.xlabel('Date', fontsize=20);
plt.tick_params(labelsize=15)
plt.xticks( fontsize = 15)
plt.legend(loc="best", fontsize =14)
plt.ticklabel_format(axis = 'y', style = 'sci', scilimits = (1,6))
plt.savefig('CPU_cap_under.png')
plt.show()
# +
## percent CPU used across network
# -
print("The Average CPU Percent Usage is only: " + str(round(concatenated_df['CPU usage [%]'].mean(),2)) + "%!!")
print("The Minimum CPU Percent Usage is: " + str(round(concatenated_df['CPU usage [%]'].min(),2)) + "%!!")
print("The Maximum CPU Percent Usage is: " + str(round(concatenated_df['CPU usage [%]'].max(),2)) + "%, possibly inidcating the system crashed?")
# ## What might cause over provision? Spikes in Network throughput?
#
# ### Graphs below are aggregated (summed)
# +
cpu = concatenated_df[['CPU usage [MHZ]']]
receive = concatenated_df[['Network received throughput [KB/s]']]
transmit = concatenated_df[['Network transmitted throughput [KB/s]']]
provisioned = concatenated_df[['CPU capacity provisioned [MHZ]']]
hourlycpu = cpu.resample('H').sum()
hourlytransmit = transmit.resample('H').sum()
hourlyreceive = receive.resample('H').sum()
hourlyprovisioned = provisioned.resample('H').sum()
# +
hourlytransmit.plot(color = "purple",linewidth = 4, figsize=(10, 5))
plt.title('Transmitted Throughput [KB/s] Totals \n Resampled & Aggregated Hourly',fontsize=15);
plt.ylabel('Transmitted Throughput [KB/s]', fontsize=15);
plt.xlabel('', fontsize=15);
hourlyreceive.plot( linewidth = 4, figsize=(10, 5))
plt.title('Received Throughput [KB/s] Totals \n Resampled & Aggregated Hourly',fontsize=15);
plt.ylabel('Received Throughput [KB/s]', fontsize=15);
plt.xlabel('', fontsize=15);
plt.yticks(fontsize=15);
plt.xticks(fontsize=15);
hourlyprov.plot(color = "g", linewidth = 4, figsize=(10, 5))
plt.title('CPU Provisioned Totals \n Resampled & Aggregated Hourly',fontsize=15);
plt.ylabel('CPU Capacity Provisioned [MHz] $e^{7}$', fontsize=15);
plt.xlabel('', fontsize=15);
plt.yticks(fontsize=15);
plt.xticks(fontsize=15);
hourlycpu.plot(linewidth = 4, figsize=(10, 5))
plt.title('CPU Usage Totals \n Resampled & Aggregated Hourly',fontsize=15);
plt.ylabel('CPU usage [MHz] $e^{7}$', fontsize=15);
plt.xlabel('', fontsize=15);
plt.yticks(fontsize=15);
plt.xticks(fontsize=15);
# -
# ### Graphs below are max values across network
hourlycpu = cpu.resample('H').max()
hourlytransmit = transmit.resample('H').max()
hourlyreceive = receive.resample('H').max()
hourlyprovisioned = provisioned.resample('H').max()
# +
hourlytransmit.plot(color = "purple",linewidth = 4, figsize=(10, 5))
plt.title('Transmitted Throughput [KB/s] Max',fontsize=15);
plt.ylabel('Transmitted Throughput [KB/s]', fontsize=15);
plt.xlabel('', fontsize=15);
hourlyreceive.plot( linewidth = 4, figsize=(10, 5))
plt.title('Received Throughput [KB/s] Max',fontsize=15);
plt.ylabel('Received Throughput [KB/s]', fontsize=15);
plt.xlabel('', fontsize=15);
plt.yticks(fontsize=15);
plt.xticks(fontsize=15);
hourlyprovisioned.plot(color = "g", linewidth = 4, figsize=(10, 5))
plt.title('CPU Provisioned Max',fontsize=15);
plt.ylabel('CPU Capacity Provisioned [MHz] $e^{7}$', fontsize=15);
plt.xlabel('', fontsize=15);
plt.yticks(fontsize=15);
plt.xticks(fontsize=15);
hourlycpu.plot(linewidth = 4, figsize=(10, 5))
plt.title('CPU Usage Max',fontsize=15);
plt.ylabel('CPU usage [MHz] $e^{7}$', fontsize=15);
plt.xlabel('', fontsize=15);
plt.yticks(fontsize=15);
plt.xticks(fontsize=15);
# -
# ## Visualize rolling mean trends over time, large spike, keep in model
df_rm = pd.concat([receive.rolling(12).mean(), transmit.rolling(12).mean()], axis=1)
df_rm.plot(figsize=(15,5), linewidth=2, fontsize=20)
plt.xlabel('Date', fontsize=20);
df_rm = pd.concat([cpu.rolling(24).sum()], axis=1)
df_rm.plot(figsize=(15,5), linewidth=2, fontsize=20)
plt.xlabel('Date', fontsize=20);
# ## Zoom in to look at hourly trends in cpu usage
hourlycpu[60:120].plot(style=[':', '--', '-'])
plt.ylabel('CPU Usage Avg [MHZ]');
# ## Plots of CPU Usage Across the Week- Highly Variable!
hourlydat = concatenated_df.resample('H').sum()
hourlydat['Date_Time'] = hourlydat.index
hourlydat['weekday'] = hourlydat['Date_Time'].dt.dayofweek
hourlydat['weekend'] = ((hourlydat.weekday) // 5 == 1).astype(float)
# ### Feature engineering with the date
hourlydat['month']=hourlydat['Date_Time'].dt.month
hourlydat['day']=hourlydat['Date_Time'].dt.day
hourlydat.drop('Date_Time', axis=1, inplace=True)
hourlydat.drop('Timestamp [ms]', axis=1, inplace=True)
plotdays = hourlydat.groupby('weekday').agg({'CPU usage [MHZ]': ['mean']})
plotdays = pd.DataFrame(plotdays)
plotdays.plot(linewidth = 4, figsize=(7, 7),legend=None)
plt.title('CPU Usage Totals \n Across Days',fontsize=20);
plt.ylabel('CPU usage [MHZ]', fontsize=15);
plt.xlabel('', fontsize=15);
plt.xticks(np.arange(7), ('Mon', 'Tues', 'Wed', 'Thurs', 'Fri', 'Sat', 'Sun'), fontsize=15);
plt.yticks(fontsize=15);
plt.figure(figsize=(7,7))
plt.title('CPU Usage Totals \n Across Days',fontsize=20);
plt.ylabel('CPU usage [MHZ]', fontsize=15);
plt.yticks(fontsize=15);
plt.xlabel('', fontsize=15);
sns.boxplot(y=hourlydat['CPU usage [MHZ]'],x = hourlydat.weekday,
whis=np.inf, palette="vlag",linewidth=3)
plt.xticks(np.arange(7), ('Mon', 'Tues', 'Wed', 'Thurs', 'Fri', 'Sat', 'Sun'), fontsize=15);
plt.figure(figsize=(7,7))
plt.title('CPU Usage Lower on Weekends',fontsize=20);
plt.ylabel('CPU usage [MHZ]', fontsize=15);
plt.yticks(fontsize=15);
sns.boxplot(y=hourlydat['CPU usage [MHZ]'],x = hourlydat.weekend,
whis=np.inf, palette="vlag",linewidth=3)
plt.xticks(np.arange(2), ('Weekday', 'Weekend'), fontsize=15);
plt.xlabel('', fontsize=15);
# ## Visualize Correlations in Data (hourlydat)
plt.figure(figsize=(10, 8))
sns.heatmap(hourlydat.corr())
| Visualize_Initial_Explore.py.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Indentation, indentation, indentation
import numpy as np
# Python cares a lot about indentation.
#
# You are going see this often in Python code.
# `for` loops are one of many places that Python depends on indentation. The indentation tells Python which statements are in the loop, and which are outside the loop.
#
# Remember that the `for` statement:
#
# * starts with the word `for`, followed by
# * a variable name (the *loop variable*) followed by
# * the word `in` followed by
# * an expression that gives sequence of values followed by
# * the character `:` followed by
# * an indented block of one or more statements. This is the *body* of
# the `for` loop.
#
# Here was our first `for` loop:
for i in np.arange(3):
print(i)
# Following the sequence above, we have:
#
# * `for`
# * `i` (the loop variable name)
# * `in`
# * `np.arange(3)` (a sequence with three values \- 0, 1, 2)
# * `:`
# * ` print(i)` (the indented block, consisting of one statement.
# If we want to execute more than one statement in the loop, we need to indent each statement:
for i in np.arange(3):
print(i)
print('Finished this iteration of the loop')
# In the above, both statements are indented, so Python runs both statements for each run through the loop.
#
# The first not-indented statement signals that the `for` loop body is over:
for i in np.arange(3):
print(i)
print('Finished this iteration of the loop')
print('Now the loop has finished')
# The lines in the `for` block must have the same indentation. Try
# knocking one space off the indentation in one of the lines in the loop above, and see what happens.
#
# The first line must end with a colon character. Try knocking the
# colon off the line beginning `for` above, and see what happens.
#
# There must be a `for` block. Try removing all the indentation from the line `print(i)` above, and see what happens.
#
# Now try the [fix my fors exercise](../../exercises/fix_my_fors.zip).
| notebooks/03/indentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="RJhCQX-j29vY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="bc5ed941-b7b3-44a3-ff79-95c0437bc277" executionInfo={"status": "ok", "timestamp": 1534764320646, "user_tz": -60, "elapsed": 8882, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-udmeYOwOAM8/AAAAAAAAAAI/AAAAAAAACcE/FeHoxk7zwGo/s50-c-k-no/photo.jpg", "userId": "110898134925759090027"}}
# TensorFlow and tf.keras
import tensorflow as tf
from tensorflow import keras
# Helper libraries
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
# + id="jqOKprIB3FE4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="2602c970-19b8-4f44-90cb-d641fe71c7fd" executionInfo={"status": "ok", "timestamp": 1534764322478, "user_tz": -60, "elapsed": 1364, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-udmeYOwOAM8/AAAAAAAAAAI/AAAAAAAACcE/FeHoxk7zwGo/s50-c-k-no/photo.jpg", "userId": "110898134925759090027"}}
mnist = keras.datasets.mnist
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
# + id="90C0TJVi4dGC" colab_type="code" colab={}
class_names = ['0', '1', '2', '3', '4',
'5', '6', '7', '8', '9']
# + id="I94A87iO5qhu" colab_type="code" colab={}
train_images = train_images / 255
test_images = test_images / 255
# + id="cxarLtRR9VvP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 589} outputId="9e425488-c18d-4e3d-b86a-eab2290a0eb7" executionInfo={"status": "ok", "timestamp": 1534764329753, "user_tz": -60, "elapsed": 2200, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-udmeYOwOAM8/AAAAAAAAAAI/AAAAAAAACcE/FeHoxk7zwGo/s50-c-k-no/photo.jpg", "userId": "110898134925759090027"}}
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
plt.xlabel(train_labels[i])
# + id="xiaOVh7pCk6Y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 351} outputId="d056eb54-9001-4629-a249-75f37442429b" executionInfo={"status": "ok", "timestamp": 1534764767501, "user_tz": -60, "elapsed": 755, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-udmeYOwOAM8/AAAAAAAAAAI/AAAAAAAACcE/FeHoxk7zwGo/s50-c-k-no/photo.jpg", "userId": "110898134925759090027"}}
plt.figure()
plt.imshow(train_images[0])
plt.colorbar()
plt.grid(False)
# + id="iyaLOQ9J-O4_" colab_type="code" colab={}
#training loop to test learning rates - this is overfitting to test set though
learning_rates = [0.001, 0.005, 0.0001, 0.0005]
test_accuracies = []
for i in range(len(learning_rates)):
model = keras.Sequential([
keras.layers.Flatten(input_shape=(28, 28)),
keras.layers.Dense(128, activation=tf.nn.relu),
keras.layers.Dense(10, activation=tf.nn.softmax)
])
model.compile(optimizer=tf.train.AdamOptimizer(learning_rate=learning_rates[i]),
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
model.fit(train_images, train_labels, epochs=7, verbose=0)
test_loss, test_acc = model.evaluate(test_images, test_labels)
test_accuracies.append(test_acc)
keras.backend.clear_session()
# + id="RZH9xDkkJzGS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 374} outputId="46288da4-1c11-4a92-a937-0184ce4db5df" executionInfo={"status": "ok", "timestamp": 1534764849551, "user_tz": -60, "elapsed": 77755, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-udmeYOwOAM8/AAAAAAAAAAI/AAAAAAAACcE/FeHoxk7zwGo/s50-c-k-no/photo.jpg", "userId": "110898134925759090027"}}
# best learning rate
model = keras.Sequential([
keras.layers.Flatten(input_shape=(28, 28)),
keras.layers.Dense(128, activation=tf.nn.relu),
keras.layers.Dense(10, activation=tf.nn.softmax)
])
model.compile(optimizer=tf.train.AdamOptimizer(learning_rate=0.0007),
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
model.fit(train_images, train_labels, epochs=10)
# + id="fSe6uJk4Lfaq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="03d780f2-0f6b-4a1d-dc34-2bc7fcc4fe21" executionInfo={"status": "ok", "timestamp": 1534765532578, "user_tz": -60, "elapsed": 938, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-udmeYOwOAM8/AAAAAAAAAAI/AAAAAAAACcE/FeHoxk7zwGo/s50-c-k-no/photo.jpg", "userId": "110898134925759090027"}}
test_loss, test_acc = model.evaluate(test_images, test_labels)
print('Test accuracy:', test_acc)
# + id="sPKOe3wXNuyv" colab_type="code" colab={}
predictions = model.predict(test_images)
# + id="6gW6J8-lNze-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="01e42fbc-5fcc-49a3-a854-f179df337c9c" executionInfo={"status": "ok", "timestamp": 1534766581775, "user_tz": -60, "elapsed": 425, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-udmeYOwOAM8/AAAAAAAAAAI/AAAAAAAACcE/FeHoxk7zwGo/s50-c-k-no/photo.jpg", "userId": "110898134925759090027"}}
print(np.argmax(predictions[0]))
print(test_labels[0])
# + id="Tse2s3u5NMdK" colab_type="code" colab={}
def plot_image(i, predictions_array, true_label, img):
predictions_array, true_label, img = predictions_array[i], true_label[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.imshow(img, cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label:
color = 'blue'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(predicted_label,
100*np.max(predictions_array),
true_label),
color=color)
def plot_value_array(i, predictions_array, true_label):
predictions_array, true_label = predictions_array[i], true_label[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
thisplot = plt.bar(range(10), predictions_array, color="#777777")
plt.ylim([0, 1])
predicted_label = np.argmax(predictions_array)
thisplot[predicted_label].set_color('red')
thisplot[true_label].set_color('blue')
# + id="wI-Vf0XgWFKN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 203} outputId="011bad6f-e948-4452-c141-9d9886766992" executionInfo={"status": "ok", "timestamp": 1534769005824, "user_tz": -60, "elapsed": 811, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-udmeYOwOAM8/AAAAAAAAAAI/AAAAAAAACcE/FeHoxk7zwGo/s50-c-k-no/photo.jpg", "userId": "110898134925759090027"}}
i = 764
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(1,2,2)
plot_value_array(i, predictions, test_labels)
# + id="1euwITK2NUOz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 589} outputId="9fcf7c58-552a-4443-f701-75ff4d0c882d" executionInfo={"status": "ok", "timestamp": 1534769150968, "user_tz": -60, "elapsed": 2672, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-udmeYOwOAM8/AAAAAAAAAAI/AAAAAAAACcE/FeHoxk7zwGo/s50-c-k-no/photo.jpg", "userId": "110898134925759090027"}}
# Plot the first X test images, their predicted label, and the true label
# Color correct predictions in blue, incorrect predictions in red
num_rows = 5
num_cols = 3
num_images = num_rows*num_cols
plt.figure(figsize=(2*2*num_cols, 2*num_rows))
for i in range(num_images):
plt.subplot(num_rows, 2*num_cols, 2*i+1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(num_rows, 2*num_cols, 2*i+2)
plot_value_array(i, predictions, test_labels)
# + id="ZJM22ceMUyS0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d309c6c3-0dae-4fe3-b48e-f0965e67c52b" executionInfo={"status": "ok", "timestamp": 1534769041572, "user_tz": -60, "elapsed": 562, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-udmeYOwOAM8/AAAAAAAAAAI/AAAAAAAACcE/FeHoxk7zwGo/s50-c-k-no/photo.jpg", "userId": "110898134925759090027"}}
i=764
img = test_images[i]
# Add the image to a batch where it's the only member.
img = (np.expand_dims(img,0))
print(img.shape)
# + id="HH2jaoEFVENL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 347} outputId="956944d1-1cf9-487d-8fe3-a71f6d7ddc30" executionInfo={"status": "ok", "timestamp": 1534769142622, "user_tz": -60, "elapsed": 634, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-udmeYOwOAM8/AAAAAAAAAAI/AAAAAAAACcE/FeHoxk7zwGo/s50-c-k-no/photo.jpg", "userId": "110898134925759090027"}}
predictions_single = model.predict(img)
plot_value_array(i, predictions, test_labels)
_ = plt.xticks(range(10))
| MNIST_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# What if we need to scale the intensity of the galaxy to the intensity of the star in each image? That could explain why our fluxes, even post-galaxy correction, are correlated!
#
# I should:
#
# * think about a more carefully background subtracted galaxy series
#
# * then measure the flux of the comparison star in each frame, normalize that light curve by its mean
#
# * then take that normalized comp star time series and apply it to the galaxy, making its flux modulate by the same telluric effects, with the same fractional amplitude
#
# * then add the galaxy back into each image with that **galaxy time series**
#
# Support for this idea:
#
# * the correlated noise in the light curve is strongest in the channels where there is most flux from the galaxy. This makes sense I guess for any explanation.
# +
# %pylab inline
import pyfits
import numpy as np
import matplotlib
matplotlib.rcParams['font.size'] = 15
from matplotlib import pyplot as plt
import sys
sys.path.append('../')
from astropy.time import Time
Nbins = 8
paddingbounds = [210, 2130]
spectralbinbounds = np.linspace(paddingbounds[0], paddingbounds[1], Nbins+1, dtype=int)
galaxyimagepath = '/astro/users/bmmorris/git/research/keck/2014september/analysis/rightnod/galaxy/wholeframegalaxyimg.fits'
wavelengthbincenters = np.load('photoutputs/wavelengthbincenters.npy')
#spectralbinbounds = np.linspace(paddingbounds[0], paddingbounds[1], Nbins+1, dtype=int)
#print spectralbinbounds
firstlines = \
'''
# Region file format: DS9 version 4.1
global color=green dashlist=8 3 width=1 font="helvetica 10 normal roman" select=1 highlite=1 dash=0 fixed=0 edit=1 move=1 delete=1 include=1 source=1
physical
'''
# Box format:
# centerx centery widthx widthy rot
with open('binregions.reg','w') as reg:
for i in range(len(spectralbinbounds)-1):
centerx = 0.5*(spectralbinbounds[i] + spectralbinbounds[i+1])
centery = 2024/2
widthx = spectralbinbounds[i+1] - spectralbinbounds[i]
widthy = 2024
angle = 0
linewidth = 3
wavelength = wavelengthbincenters[i]
reg.write("box({0:f},{1:f},{2:f},{3:f},{4:f}) # width={5} text={{{6:.3f}}} \n".format(
centerx, centery, widthx, widthy, angle, linewidth, wavelength))
# opends9 = True
# if opends9:
# import os
# os.system('ds9 '+galaxyimagepath+' -regions binregions.reg &')
print 'Run:\n'+'ds9 '+galaxyimagepath+' -regions binregions.reg &'
# + language="bash"
# ds9 /astro/users/bmmorris/git/research/keck/2014september/analysis/rightnod/galaxy/wholeframegalaxyimg.fits -regions binregions.reg &
# -
| analysis/galaxyproblem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploring Ebay Car Sales Data
#
# ---
#
# ## 1. Introduction
#
#
# This project is created to demonstrate the basics of data cleaning and data exploration using *pandas*.
#
# The dataset used in this project contain records of used cars from eBay-Kleinanzeigen, a classifieds section of the German eBay website.
#
# The dataset was originally scraped and uploaded to [Kaggle](https://www.kaggle.com/orgesleka/used-cars-database/data), but a few modifications have been made by DataQuest to the original dataset. About 50,000 data points were sampled from the full dataset, and the data was dirtied to make it more closely resemble what would be expected from a scraped dataset (the version uploaded to Kaggle was cleaned to be easier to work with).
#
# The data dictionary provided with data is as follows:
# - **dateCrawled** - When this ad was first crawled. All field-values are taken from this date.
# - **name** - Name of the car.
# - **seller** - Whether the seller is private or a dealer.
# - **offerType** - The type of listing
# - **price** - The price on the ad to sell the car.
# - **abtest** - Whether the listing is included in an A/B test.
# - **vehicleType** - The vehicle Type.
# - **yearOfRegistration** - The year in which the car was first registered.
# - **gearbox** - The transmission type.
# - **powerPS** - The power of the car in PS.
# - **model** - The car model name.
# - **kilometer** - How many kilometers the car has driven.
# - **monthOfRegistration** - The month in which the car was first registered.
# - **fuelType** - What type of fuel the car uses.
# - **brand** - The brand of the car.
# - **notRepairedDamage** - If the car has a damage which is not yet repaired.
# - **dateCreated** - The date on which the eBay listing was created.
# - **nrOfPictures** - The number of pictures in the ad.
# - **postalCode** - The postal code for the location of the vehicle.
# - **lastSeenOnline** - When the crawler saw this ad last online.
#
# Note: The fields **lastSeenOnline** and **dateCreated** could be used to estimate how long a car will be at least online before it is sold.
#
# ## 2. Goal & Objective of This Project
#
# This project aims to clean the data and analyze the included used car listings.
#
#
#
#
#
#
#
# ## 3. General Observation of the Dataset
# +
### Import the pandas and NumPy libraries.
### Reading the dataset into pandas.
import numpy as np
import pandas as pd
autos = pd.read_csv("autos.csv", encoding = "Latin-1")
### Note: Common encoding is "UTF-8" / "Latin-1" / "Windows-1252".
# +
### Showing a preview of the dataset.
autos
# +
### Showing the first 5 rows of data.
autos.head()
# +
### Getting the overview of all the dtypes used in the dataset, along with its shape, columns, and other information.
autos.info()
# +
### Check columns that have null values.
check_null = autos.isnull().sum()
check_null
# -
# **Observations from the above:**
#
# - The dataset contains 20 columns, most of which are strings (objects).
#
# - Some columns have null values, but none have more than ~20% null values.
#
# - Most of the dates columns are stored as strings (objects).
#
# - The column names use [camelcase](https://en.wikipedia.org/wiki/Camel_case) instead of Python's preferred [snakecase](https://en.wikipedia.org/wiki/Snake_case), which means we can't just replace spaces with underscores.
#
# Let's start by cleaning the column names so that the dataset is easier to work with.
#
# We will use snakecase for the column names.
# ## 4. Cleaning Column Names
print("Original Column Names:")
print("----------------------")
print(autos.columns)
# +
### Define a customized function to rename the column of this dataset.
### There are other more advanced technic to do this, but we will just use a simple basic function.
def rename_column(column_name):
if column_name == "yearOfRegistration":
new_name = "registration_year"
elif column_name == "monthOfRegistration":
new_name = "registration_month"
elif column_name == "notRepairedDamage":
new_name = "unrepaired_damage"
elif column_name == "dateCreated":
new_name = "ad_created"
elif column_name == "dateCrawled":
new_name = "date_crawled"
elif column_name == "offerType":
new_name = "offer_type"
elif column_name == "abtest":
new_name = "ab_test"
elif column_name == "vehicleType":
new_name = "vehicle_type"
elif column_name == "gearbox":
new_name = "gear_box"
elif column_name == "powerPS":
new_name = "power_PS"
elif column_name == "fuelType":
new_name = "fuel_type"
elif column_name == "nrOfPictures":
new_name = "no_of_pictures"
elif column_name == "postalCode":
new_name = "postal_code"
elif column_name == "lastSeen":
new_name = "last_seen"
else:
new_name = column_name
return new_name
### Looping through each column in the dataset and call the function to rename the column.
new_columns = []
for c in autos.columns:
new_name = rename_column(c)
new_columns.append(new_name)
autos.columns = new_columns
# +
### Alternatively, can also directly assign the column names as shown below.
autos.columns = ['date_crawled', 'name', 'seller', 'offer_type', 'price', 'ab_test',
'vehicle_type', 'registration_year', 'gear_box', 'power_PS', 'model',
'odometer', 'registration_month', 'fuel_type', 'brand',
'unrepaired_damage', 'ad_created', 'no_of_pictures', 'postal_code',
'last_seen']
# -
print("Renamed Column Names:")
print("---------------------")
print(autos.columns)
# +
### Showing the first 5 rows of data with new column names.
autos.head()
# -
# ## 5. Initial Exploration - Checking General Statistics of the Dataset
# +
### Check statistics for only numeric columns.
autos.describe()
# +
### Check statistics for object columns (non-numeric columns).
autos.describe(include=['O'])
# +
### Check statistics for all columns.
autos.describe(include='all')
# -
# **Observation:**
#
# Columns that have mostly one value (or where almost all of the values are the same) are candidates to be dropped:
#
# - `seller` (almost all records are "privat")
# - `offer_type` (almost all records are "Angebot")
#
# The `no_of_pictures` column looks odd, we'll need to investigate this further.
# +
### Further investigation on the 'no_of_pictures' column.
### All the values in this column is 0.
autos["no_of_pictures"].value_counts()
# -
# ## 6. Dropping Columns
#
# Drop the following columns that have mostly one value (or where almost all of the values are the same):
# - `seller` (almost all records are "privat")
# - `offer_type` (almost all records are "Angebot")
# - `no_of_pictures` (all records are "0")
autos = autos.drop(["seller", "offer_type", "no_of_pictures"], axis=1)
# ## 7. Cleaning Numeric Data Stored As Text
#
# ### 7.1 Converting the 'price' and 'odometer' column to numeric type
# +
### Replace dollar sign and comma with empty string before converting the 'price' column to int type.
autos["price"] = autos["price"].str.replace("$","").str.replace(",","").astype(int)
### Replace 'km' with empty string before converting the 'odometer' column to int type.
autos["odometer"] = autos["odometer"].str.replace("km","").str.replace(",","").astype(int)
# -
# ### 7.2 Renaming the 'odometer' column to 'odometer_km'
# +
### Renaming the 'odometer' column to 'odometer_km' so that we know the numeric values are in km.
autos.rename({"odometer":"odometer_km"}, axis=1, inplace=True)
### Note: Either use inplace=True or assign the result back to the dataframe; otherwise, the modifications will be lost.
# -
# ## 8. Identify Outliers (values that look unrealistically high or low)
#
# ### 8.1 Inspecting the 'price' Column
# +
### Checking the number of unique price values.
autos["price"].unique().shape
# +
### Checking statistics of the price column.
autos["price"].describe()
# +
### Checking the count for each unique price value.
autos["price"].value_counts().sort_index(ascending=True)
# -
autos["price"].value_counts(normalize=True).sort_index().head(10)
# +
### Take a closer look on prices that are on the high end (sort by descending order).
autos["price"].value_counts().sort_index(ascending=False).head(20)
# -
# **Observation:**
#
# - There are 2357 unique price values.
#
# - 1421 records are with 0 price (which is about 2.8% of the listing), and the maximum price is one hundred million (99,999,999).
#
# - For prices that are on the high end, it seems like prices increased steadily until 350,000 and from there jump to unrealistically high values. There are 14 records with prices greater than 350,000.
#
# - These unrealistic records, which are less than 3% of the listing, shall be removed later.
# ### 8.2 Inspecting the 'odometer_km' Column
# +
### Checking the number of unique mileage values.
autos["odometer_km"].unique().shape
# +
### Checking statistics of the mileage.
autos["odometer_km"].describe()
# +
### Checking the count for each unique mileage value.
autos["odometer_km"].value_counts().sort_index(ascending=True)
# -
# **Observation:**
#
# There seems to be more records with high mileage than low mileage in the listing.
# ### 8.3 Inspecting Dates
#
# As seen below, dates for `date_crawled`, `ad_created`, and `last_seen` are all identified as string values by pandas.
# We need to convert the data into a numerical representation so we can understand it quantitatively.
#
# +
### Showing the first 5 records of 'date_crawled', 'ad_created', and 'last_seen'.
autos[['date_crawled','ad_created','last_seen']][0:5]
# -
# ### 8.3.1 Inspecting 'date_crawled'
# +
### Select only the first 10 characters of the date to generate a distribution, and then sort by the index.
### To include missing values in the distribution with 'dropna=False'
### Use sort_index() to rank by date in ascending order.
autos["date_crawled"].str[:10].value_counts(dropna=False).sort_index()
# Following code is to use percentages instead of counts with 'normalize=True'.
# autos["date_crawled"].str[:10].value_counts(normalize=True, dropna=False).sort_index()
# -
autos["date_crawled"].str[:10].describe()
# **Observation:**
#
# - There are 34 unique date value for `date_crawled`.
# - Looks like the website is crawled daily for about a month in March and April 2016 (from 2016-03-05 to 2016-04-07).
# - The distribution of listings crawled on each day is roughly uniform.
# - The date with the highest number of ads crawled, 1934, is on 2016-04-03.
# ### 8.3.2 Inspecting 'ad_created' Date
autos["ad_created"].str[:10].describe()
# +
### Very few ads are created on 2015.
### Let's check further below.
autos["ad_created"].str[:10].value_counts(dropna=False).sort_index()
# +
### Noticed that there are significant number of ads created on and after March 2016.
### Very few ads are created before March 2016.
autos["ad_created"].str[:10].value_counts(dropna=False).sort_index().head(50)
# -
autos["ad_created"].str[:10].value_counts(dropna=False).sort_index().tail(50)
# **Observation:**
#
# - There are 76 unique date value for `ad_created`.
#
# - Noticed that there are significant number of ads created on and after March 2016, particularly during the crawling period which is from 2016-03-05 to 2016-04-07.
#
# - Very few ads are created before March 2016.
#
# - The date with the highest number of ads created, 1946, is on 2016-04-03.
#
# - This is also the same date where the highest number of ads are crawled.
# ### 8.3.3 Inspecting 'last_seen' Date
autos["last_seen"].str[:10].describe()
# +
### There is significant number of ads that are last seen on 2016-04-06.
autos["last_seen"].str[:10].value_counts(dropna=False).sort_index()
# -
# **Observation:**
#
# - There are 34 unique date value for `last_seen`.
#
# - There is a spike of records (6214, 11050, 6546) with `last seen` date on the last 3 days of the crawling period.
#
# - The highest number of records (11050) are last seen on 2016-04-06, a day before the crawling period ends.
#
# - This is unlikely due to a spike in sales.
# ### 8.4 Inspecting 'registration_year'
autos["registration_year"].describe()
autos["registration_year"].value_counts()
autos["registration_year"].value_counts().sort_index()
# **Observation:**
#
# The year that the car was first registered will likely indicate the age of the car. Looking at this column, we note some odd values. The minimum value is 1000, long before cars were invented and the maximum is 9999, many years into the future.
#
# Because a car can't be first registered after the listing was seen, any vehicle with a registration year above 2016 is definitely inaccurate. Determining the earliest valid year is more difficult. Realistically, it could be somewhere in the first few decades of the 1900s.
#
# Let's count the number of listings with cars that fall outside the 1900 - 2016 interval and see if it's safe to remove those rows entirely, or if we need more custom logic.
# +
### Checking the number of records where 'registration_year' is before 1900 OR after 2016.
autos[(autos["registration_year"] < 1900) | (autos["registration_year"] > 2016)]
# -
# There are 1972 records where 'registration_year' is before 1900 OR after 2016, which is about 4%.
#
# We will remove these outliers in the following section.
# ## 9. Remove Outliers
#
# ### 9.1 Remove records with unrealistic 'registration_year'
# +
### Filter records where 'registration_year' is between 1900 and 2016.
boo = autos["registration_year"].between(1900, 2016)
autos2 = autos.loc[boo]
autos2.describe()
# -
autos2["registration_year"].value_counts(normalize=True)
# ### 9.2 Remove records with unrealistic 'price'
# +
### Filter records where 'price' is not 0, and not greater than 350,000.
boo = autos2["price"].between(1, 350000)
autos2 = autos2.loc[boo]
autos2.describe()
# -
# ## 10. Find Popular Brands
# +
### Create an array of unique brands
brands = autos2["brand"].unique()
print(brands)
# +
### Getting the count for each unique brand.
autos2["brand"].value_counts()
# +
### Selecting the top 10 brands to be used for further analysis.
top10_brands = autos2["brand"].value_counts().head(10)
print(top10_brands)
# +
### Getting the index label of the top 10 brands.
top10_brands = top10_brands.index
print(top10_brands)
# -
# **Observation:**
#
# Volkswagen is the most popular brand, having 9862 records, which is almost double of it's first rival, BMW (5137 records).
# ## 11. Analyze the Top 10 Brands
# +
### Create a general function to sort a given dictionary in descending order.
### This function will return a sorted list of tuple.
### Note that sorted() built-in function doesn't work too well with dictionaries because
### it only considers and returns the dictionary keys (instead of the key+value pair).
### We need to transform the dictionary into a list of tuples in order to do sorting.
def sort_dict(dictionary):
### Initialise a blank list to store tuples.
list_of_tuple = []
for key in dictionary:
val_key_tuple = (dictionary[key], key)
list_of_tuple.append(val_key_tuple)
list_of_tuple_sorted = sorted(list_of_tuple, reverse = True)
# Either print the sorted records, or return the sorted list of tuple.
#for item in list_of_tuple_sorted:
# print(item[1], ':', item[0])
return list_of_tuple_sorted
# -
# ### 11.1 Exploring Price by Brand
# +
### Getting the average price for each of the top 10 brands.
avg_price_by_brand = {}
for b in top10_brands:
selected_rows = autos2[autos2["brand"] == b]
mean = selected_rows["price"].mean()
avg_price_by_brand[b] = int(mean)
print("### Average Price by Brand ###")
avg_price_by_brand
# +
### Call the sort_dict() function to sort the records.
print("### Average Price by Brand (Sorted) ###")
sort_dict(avg_price_by_brand)
# -
# **Observation:**
#
# In the top 10 brands, there's a distinct price gap.
#
# - Audi, <NAME>, and BMW are more expensive.
# - Opel, Fiat, and Renault are less expensive.
# - Volkswagen, Seat, Ford and Peugeot are in between.
#
# This may explain why Volkswagen is the most popular brand, as it's price is in between.
# ### 11.2 Exploring Mileage by Brand
# +
### Getting the average mileage for each of the top 10 brands.
avg_mileage_by_brand = {}
for b in top10_brands:
selected_rows = autos2[autos2["brand"] == b]
mean = selected_rows["odometer_km"].mean()
avg_mileage_by_brand[b] = int(mean)
#print("### Average Mileage by Brand ###")
#avg_mileage_by_brand
### Call the sort_dict() function to sort the records.
print("### Average Mileage by Brand (Sorted) ###")
sort_dict(avg_mileage_by_brand)
# -
# **Observation:**
#
# All the top 10 brands have high mileage.
# ### 11.3 Analyzing Price & Mileage by Brand (Part 1)
#
# Let's use aggregation to understand the average mileage for those cars and if there's any visible link with mean price.
#
# The following will combine the data from both series objects into a single dataframe (with a shared index) and display the dataframe directly.
# +
### Convert the 'avg_price_by_brand' dictionary to series objects, using the series constructor.
### The keys in the dictionary became the index in the series object.
bmp_series = pd.Series(avg_price_by_brand)
print(bmp_series)
### We can then create a single-column dataframe from this series object.
### We need to use the columns parameter when calling the dataframe constructor (which accepts a array-like object)
### to specify the column name (or the column name will be set to 0 by default).
df = pd.DataFrame(bmp_series, columns=['mean_price'])
df
# +
### Adding 'mean_mileage' new column to the dataframe.
df["mean_mileage"] = pd.Series(avg_mileage_by_brand)
df
# -
# **Observation:**
#
# The average mileage seems to have no visible link with mean price.
#
# There is no significant difference on the average mileage for cars with expensive or cheaper price.
#
# ### 11.4 Analyzing Price & Mileage by Brand (Part 2)
#
# Split the mileage (odometer_km) into groups, and use aggregation to see if average prices follows any patterns based on the mileage.
# +
### Checking the count and unique values of 'odometer_km'.
autos2["odometer_km"].value_counts().sort_index()
# +
### Split the odometer_km into 3 groups.
### Getting the average price for each of the top 10 brands, for each of the 'odometer_km' groups.
avg_price_group1 = {}
avg_price_group2 = {}
avg_price_group3 = {}
for b in top10_brands:
selected_rows_group1 = autos2[(autos2["brand"] == b) & (autos2["odometer_km"] <= 50000)]
selected_rows_group2 = autos2[(autos2["brand"] == b) & (autos2["odometer_km"].between(50001, 100000))]
selected_rows_group3 = autos2[(autos2["brand"] == b) & (autos2["odometer_km"] > 100000)]
mean_group1 = selected_rows_group1["price"].mean()
mean_group2 = selected_rows_group2["price"].mean()
mean_group3 = selected_rows_group3["price"].mean()
avg_price_group1[b] = int(mean_group1)
avg_price_group2[b] = int(mean_group2)
avg_price_group3[b] = int(mean_group3)
### Convert the 'avg_price_with_damage' dictionary to series objects, using the series constructor.
### The keys in the dictionary became the index in the series object.
mp_series = pd.Series(avg_price_group1)
### We can then create a single-column dataframe from this series object.
### We need to use the columns parameter when calling the dataframe constructor (which accepts a array-like object)
### to specify the column name (or the column name will be set to 0 by default).
df = pd.DataFrame(mp_series, columns=['mean_price_odo_up_to_50km'])
### Adding 'mean_price_non_damage' new column to the dataframe.
df["mean_price_odo_50_to_100km"] = pd.Series(avg_price_group2)
df["mean_price_odo_above_100km"] = pd.Series(avg_price_group3)
df
# -
# **Observation:**
#
# For the same brand, average price is reduced when the mileage is increased.
# ### 11.5 Analyzing Price For Cars With Damaged and Non-Damage
#
# Here we would want to find out how much cheaper are cars with damage than their non-damaged counterparts.
# +
### Checking the unique values for the 'unrepaired_damage' column.
autos2["unrepaired_damage"].value_counts()
# +
### Getting the average price for each of the top 10 brands, for cars with damage and without damage.
avg_price_with_damage = {}
avg_price_non_damaged = {}
for b in top10_brands:
selected_rows_with_damage = autos2[(autos2["brand"] == b) & (autos2["unrepaired_damage"] == "ja")]
selected_rows_non_damage = autos2[(autos2["brand"] == b) & (autos2["unrepaired_damage"] == "nein")]
mean_with_damage = selected_rows_with_damage["price"].mean()
mean_non_damage = selected_rows_non_damage["price"].mean()
avg_price_with_damage[b] = int(mean_with_damage)
avg_price_non_damaged[b] = int(mean_non_damage)
### Convert the 'avg_price_with_damage' dictionary to series objects, using the series constructor.
### The keys in the dictionary became the index in the series object.
mp_series = pd.Series(avg_price_with_damage)
### We can then create a single-column dataframe from this series object.
### We need to use the columns parameter when calling the dataframe constructor (which accepts a array-like object)
### to specify the column name (or the column name will be set to 0 by default).
df = pd.DataFrame(mp_series, columns=['mean_price_with_damage'])
### Adding 'mean_price_non_damage' new column to the dataframe.
df["mean_price_non_damage"] = pd.Series(avg_price_non_damaged)
df
# +
### Calculate the percentage and adding it as column to the dataframe.
df["percent_cheaper"] = (df["mean_price_with_damage"] / df["mean_price_non_damage"]) * 100
df
# -
# **Observation:**
#
# For the top 10 brands, cars with damage are around 30-40% cheaper than their non-damaged counterparts.
# ## 12. Find the most common brand/model combinations
# +
### Create a new column 'brand_and_model', which combines the brand and the model columns.
autos2["brand_and_model"] = autos2["brand"] + "/" + autos2["model"]
autos2
# +
### Getting the count for each unique brand/model combination.
autos2["brand_and_model"].value_counts()
# -
# **Observation:**
#
# The most common brand/model combination is Volkswagen/Golf (3707 records), followed by BMW/3er (2615 records).
| Project_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Fast PDE/IE course, Skoltech, Spring 2015
# + [markdown] slideshow={"slide_type": "slide"}
# ## Team
# <NAME>, Associate Professor, Skoltech
# <NAME>, Assistant Professor, Skoltech
# <NAME>, TA
# <NAME>, TA
# <NAME>, TA
# <NAME>, TA
# + [markdown] slideshow={"slide_type": "slide"}
# ## How do we grade
#
# - **50%** homework
# - **20%** 2 written tests
# - **30%** Application Period
# - **pass-fail** ping-pong test
#
# **Strict** deadlines with **5 days** of additional budget. 50% max for late submissions
#
# Attendance control **not strict**
#
# (as usual do not disappoint us: the negative bonus can be large)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Technical details
#
# 1. We will try to use [Python](www.python.org) and [IPython/Jupyter notebooks](http://nbviewer.ipython.org/github/ellisonbg/talk-2014-summer/blob/master/Jupyter%20and%20IPython.ipynb)
# for everything (lecture notes & homework assignments)
# 2. I highly recommend you to install [Anaconda Python Distribution](http://continuum.io/downloads)
# (Skoltech email is enough for the free Academic license for the full version)
# 3. Python 2.7 is used
# 4. Feel free to ask TAs for help
# + [markdown] slideshow={"slide_type": "slide"}
# ## Course description
#
# **PDE** (Partial differential equations) and **IE** (integral equations) are the barebones for physical simulations.
#
# We will discuss how to solve them, and how to solve them fast.
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Learning outcomes
#
# - Finite elements & meshes for elliptic & parabolic problems
# - Sparse linear systems (direct solvers, preconditioners, multigrid)
# - Software packages for solving PDEs
# - Basic integral equations for exterior problems
# - High-frequency problems & convolution & FFT
# + [markdown] slideshow={"slide_type": "slide"}
# ## (Approximate) Syllabus
# - **Week 1:** Physical models & PDE & IE, finite differences 1D/2D
# - **Week 2:** FEM (basic elements, meshes, tools for meshing)
# - **Week 3:** Sparse matrices/iterative methods/multigrid (geom & AMG)
# - **Week 4:** Exterior problems & IE: discretization, quadrature, multipole, hierarchical matrices + test)
# - **Week 5:** High-frequency & Fourier
# - **Week 6:** Software packages for PDEs
# - **Week 7:** Exam & test
# - **Week 8:** App Period
# + [markdown] slideshow={"slide_type": "slide"}
# ## Homework
# - 4 problem sets
# - Each Friday (starting from the **week 2**) will be the deadline
# - Everybody has a resource of **5 days** to postpone problem sets deadlines
#
# ## Tests
# - 2 written tests (**week 4** and **week 7**)
# - Ping-pong test (**week 7**)
# + [markdown] slideshow={"slide_type": "slide"}
# ##Materials
#
# - Lecture notes
# - Bother TAs with questions
# - We will point out important books/papers when appropriate
# + [markdown] slideshow={"slide_type": "slide"}
# ## Starting
# Now let us start the [First lecture](Lecture-1.ipynb)
#
# + slideshow={"slide_type": "skip"}
from IPython.core.display import HTML
def css_styling():
styles = open("./styles/custom.css", "r").read()
return HTML(styles)
css_styling()
| PDE_start.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Copyright 2020 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# -
# <a target="_blank" href="https://colab.research.google.com/github/GoogleCloudPlatform/keras-idiomatic-programmer/blob/master/books/deep-learning-design-patterns/Workshops/Junior/Deep%20Learning%20Design%20Patterns%20-%20Workshop%20-%20Chapter%202.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# ## Deep Learning Design Patterns - Code Labs
#
# ## Lab Exercise #6 - Get Familiar with Wide Convolutional Models
#
# ## Prerequistes:
#
# 1. Familiar with Python
# 2. Completed Chapter 2: Wide Convolutional Models
#
# ## Objectives:
#
# 1. Code a Naive Inception module
# 2. Code a Inception V1 block
# 3. Refactor an Inception V1 block.
# 4. Code a mini Wide Residual Network (WRN)
# ## Code a Naive Inception Module
#
# Let's code an naive inception module:
#
# <img src='https://github.com/GoogleCloudPlatform/keras-idiomatic-programmer/blob/master/books/deep-learning-design-patterns/Workshops/Junior/naive-inception.jpg?raw=true'>
#
#
# You fill in the blanks (replace the ??), make sure it passes the Python interpreter.
#
# You will need to:
#
# 1. Create 4 branches.
# 2. Implement each parallel branch
# 3. Concatenate the output from each branch into a single output for the module.
# +
from tensorflow.keras import Input, Model
from tensorflow.keras.layers import Conv2D, ReLU, BatchNormalization, MaxPooling2D, Concatenate, SeparableConv2D
def naive_inception(inputs):
# pooling branch
# HINT: The feature map output must stay the same, so don't downsample it, and remember the padding
x1 = MaxPooling2D((2, 2), ??)(inputs)
# 1x1 branch
x2 = Conv2D(64, (1, 1), strides=1, padding='same', activation='relu')(inputs)
# 3x3 branch
# HINT: should look like the 1x1 convolution, except it uses a 3x3
x3 = ??
# 5x5 branch
x4 = Conv2D(64, (5, 5), strides=1, padding='same', activation='relu')(inputs)
# Concatenate the output from the four branches together
# HINT: Should be a list of the four branches outputs (x...)
outputs = Concatenate()([??])
return outputs
inputs = Input((32, 32, 3))
outputs = naive_inception(inputs)
model = Model(inputs, outputs)
# -
# ### Verify the module using summary method
#
# It should look like below:
#
# ```
# Layer (type) Output Shape Param # Connected to
# ==================================================================================================
# input_3 (InputLayer) [(None, 32, 32, 3)] 0
# __________________________________________________________________________________________________
# max_pooling2d_2 (MaxPooling2D) (None, 32, 32, 3) 0 input_3[0][0]
# __________________________________________________________________________________________________
# conv2d_6 (Conv2D) (None, 32, 32, 64) 256 input_3[0][0]
# __________________________________________________________________________________________________
# conv2d_7 (Conv2D) (None, 32, 32, 64) 1792 input_3[0][0]
# __________________________________________________________________________________________________
# conv2d_8 (Conv2D) (None, 32, 32, 64) 4864 input_3[0][0]
# __________________________________________________________________________________________________
# concatenate_2 (Concatenate) (None, 32, 32, 195) 0 max_pooling2d_2[0][0]
# conv2d_6[0][0]
# conv2d_7[0][0]
# conv2d_8[0][0]
# ==================================================================================================
# Total params: 6,912
# Trainable params: 6,912
# Non-trainable params: 0
# ```
model.summary()
# ## Code and Inception V1 Block
#
# Let's now code an inception V1 block (referred to as module in paper). Remember, this V1 module used factorization to reduce complexity (parameters) while maintaining representational equivalence.
#
# <img src='https://github.com/GoogleCloudPlatform/keras-idiomatic-programmer/blob/master/books/deep-learning-design-patterns/Workshops/Junior/block-v1.jpg?raw=true'>
#
# You will need to:
#
# 1. Add 1x1 bottleneck convolutions
# +
def inception_block(inputs):
# pooling branch
x1 = MaxPooling2D((3, 3), strides=(1, 1), padding='same')(inputs)
# Add a 1x1 bottleneck convolution with 64 filters
# HINT: the output shape should not change (think of strides and padding)
x1 = Conv2D(64, (1, 1), ??)
# 1x1 branch
x2 = Conv2D(64, (1, 1), strides=(1, 1), padding='same', activation='relu')(inputs)
# 3x3 branch
# Add 1x1 bottleneck convolution of 64 filters
# HINT: the input should be the input to the block
x3 = ??
x3 = Conv2D(96, (3, 3), strides=(1, 1), padding='same', activation='relu')(x3)
# 5x5 branch
# Add 1x1 bottleneck convolution of 64 filters
# HINT: the input should be the input to the block
x4 = ??
x4 = Conv2D(48, (5, 5), strides=(1, 1), padding='same', activation='relu')(x4)
outputs = Concatenate()([x1, x2, x3, x4])
return outputs
inputs = Input((32, 32, 3))
outputs = inception_block(inputs)
model = Model(inputs, outputs)
# -
# ### Verify the module using summary method
#
# It should look like below:
#
# ```
# Model: "model_3"
# __________________________________________________________________________________________________
# Layer (type) Output Shape Param # Connected to
# ==================================================================================================
# input_8 (InputLayer) [(None, 32, 32, 3)] 0
# __________________________________________________________________________________________________
# max_pooling2d_7 (MaxPooling2D) (None, 32, 32, 3) 0 input_8[0][0]
# __________________________________________________________________________________________________
# conv2d_29 (Conv2D) (None, 32, 32, 64) 256 input_8[0][0]
# __________________________________________________________________________________________________
# conv2d_31 (Conv2D) (None, 32, 32, 64) 256 input_8[0][0]
# __________________________________________________________________________________________________
# conv2d_27 (Conv2D) (None, 32, 32, 64) 256 max_pooling2d_7[0][0]
# __________________________________________________________________________________________________
# conv2d_28 (Conv2D) (None, 32, 32, 64) 256 input_8[0][0]
# __________________________________________________________________________________________________
# conv2d_30 (Conv2D) (None, 32, 32, 96) 55392 conv2d_29[0][0]
# __________________________________________________________________________________________________
# conv2d_32 (Conv2D) (None, 32, 32, 48) 76848 conv2d_31[0][0]
# __________________________________________________________________________________________________
# concatenate_7 (Concatenate) (None, 32, 32, 272) 0 conv2d_27[0][0]
# conv2d_28[0][0]
# conv2d_30[0][0]
# conv2d_32[0][0]
# ==================================================================================================
# Total params: 133,264
# Trainable params: 133,264
# Non-trainable params: 0
# ```
model.summary()
# ## Refactor an Inception V1 Block
#
# Let's refactor the Inception V1 block, where:
#
# 1. The 5x5 parallel convolution is replaced by two 3x3 (B(3,3))
# 2. Replace the 3x3 convolution with a spatially separable convolution (3x1, 1x3)
#
#
# You will need to:
#
# 1. Add the parallel spatially separable 3x1 and 1x3 convolutions.
# 2. Concatenate the outputs together from the separable convolutions.
# 3. Add the sequential two 3x3 convolutions.
# +
def inception_block(inputs):
# pooling branch
x1 = MaxPooling2D((3, 3), strides=(1, 1), padding='same')(inputs)
x1 = Conv2D(64, (1, 1), strides=(1, 1), padding='same', activation='relu')(inputs)
# 1x1 branch
x2 = Conv2D(64, (1, 1), strides=(1, 1), padding='same', activation='relu')(inputs)
# 3x3 branch
x3 = Conv2D(64, (1, 1), strides=(1, 1), padding='same', activation='relu')(inputs)
# Add two parallel spatially separable convolutions for 3x1 and 1x3 with 96 filters
# HINT: Use SeparableConv2D. The input to both convolutions is the same, i.e., the output from
# the prior 1x1 bottleneck.
x3_a = ??
x3_b = ??
# Concatenate the outputs together from the spatially separable convolutions
# HINT: x3 was split into a and b, let's put them back together.
x3 = Concatenate()([??])
# 5x5 branch replaced by two 3x3
x4 = Conv2D(64, (1, 1), strides=(1, 1), padding='same', activation='relu')(inputs)
# Add two sequential 3x3 normal convolutions with 48 filters
# HINT: both should have x4 as input.
x4 = ??
x4 = ??
outputs = Concatenate()([x1, x2, x3, x4])
return outputs
inputs = Input((32, 32, 3))
outputs = inception_block(inputs)
model = Model(inputs, outputs)
# -
# ### Verify the module using summary method
#
# It should look like below. Note how the number of parameters after refactoring is about 1/2.
#
# ```
# Layer (type) Output Shape Param # Connected to
# ==================================================================================================
# input_3 (InputLayer) [(None, 32, 32, 3)] 0
# __________________________________________________________________________________________________
# conv2d_11 (Conv2D) (None, 32, 32, 64) 256 input_3[0][0]
# __________________________________________________________________________________________________
# conv2d_12 (Conv2D) (None, 32, 32, 64) 256 input_3[0][0]
# __________________________________________________________________________________________________
# separable_conv2d (SeparableConv (None, 32, 32, 96) 6432 conv2d_11[0][0]
# __________________________________________________________________________________________________
# separable_conv2d_1 (SeparableCo (None, 32, 32, 96) 6432 conv2d_11[0][0]
# __________________________________________________________________________________________________
# conv2d_13 (Conv2D) (None, 32, 32, 48) 27696 conv2d_12[0][0]
# __________________________________________________________________________________________________
# conv2d_9 (Conv2D) (None, 32, 32, 64) 256 input_3[0][0]
# __________________________________________________________________________________________________
# conv2d_10 (Conv2D) (None, 32, 32, 64) 256 input_3[0][0]
# __________________________________________________________________________________________________
# concatenate_1 (Concatenate) (None, 32, 32, 192) 0 separable_conv2d[0][0]
# separable_conv2d_1[0][0]
# __________________________________________________________________________________________________
# conv2d_14 (Conv2D) (None, 32, 32, 48) 20784 conv2d_13[0][0]
# __________________________________________________________________________________________________
# concatenate_2 (Concatenate) (None, 32, 32, 368) 0 conv2d_9[0][0]
# conv2d_10[0][0]
# concatenate_1[0][0]
# conv2d_14[0][0]
# ==================================================================================================
# Total params: 62,368
# Trainable params: 62,368
# Non-trainable params: 0
# __________________________________________________________________________________________________
# ```
model.summary()
# ## Code a Wide Residual Network
#
# Let's now code a mini version of a WRN:
#
# 1. Stem
# 2. Single Group of two residual blocks
# 3. Classifier
#
# You will need to:
#
# 1. Get the value for k (width factor) from kwargs
# 2. Pass the width factor along with block params to the block method.
# 3. Determine the number of input channels (feature maps) for the block.
# 4. Complete the residual link.
# 5. Add the activation function for the classifier.
# +
from tensorflow.keras import Input, Model
from tensorflow.keras.layers import Conv2D, BatchNormalization, ReLU, GlobalAveragePooling2D, Dense, Add
def stem(inputs):
# 3x3 16 filter stem convolution with post-activation batch norm (CONV-BN-RE)
outputs = Conv2D(16, (3, 3), strides=(1, 1), padding='same')(inputs)
outputs = BatchNormalization()(outputs)
outputs = ReLU()(outputs)
return outputs
def group(inputs, **params):
# Get the kwarg blocks info.
blocks = params['blocks']
# Get the kwarg k (width factor)
# HINT: its the value of the key 'k'
k = params[??]
# Construct each block for this group
outputs = inputs
for block_params in blocks:
# Pass the global width parameter along with the block paramters
# HINT: You extracted the key-value above
outputs = block(outputs, **block_params, k=??)
return outputs
def block(inputs, **params):
n_filters = params['n_filters']
k = params['k']
# input will not match output shape.
# do 1x1 linear projection to match the shapes
# HINT: the channels is the last dimension. Input is a 4D tensor: (batch, height, width, channels)
in_channels = inputs.shape[??]
if in_channels != n_filters:
inputs = BatchNormalization()(inputs)
inputs = Conv2D(n_filters, (1, 1), strides=(1, 1), padding='same')(inputs)
# Dimensionality expansion
outputs = BatchNormalization()(inputs)
outputs = ReLU()(outputs)
# Set the number of expanded filters
# HINT: multiply the number of filters for the block by the width factor
outputs = Conv2D(??, (3, 3), strides=(1, 1), padding='same')(outputs)
# Dimensionality reduction
outputs = BatchNormalization()(outputs)
outputs = ReLU()(outputs)
outputs = Conv2D(n_filters, (3, 3), strides=(1, 1), padding='same')(outputs)
# Add the residual link to the outputs
# HINT: the residual link is the inputs to the block
outputs = Add()([??])
return outputs
def classifier(inputs, n_classes):
# Pool and Flatten into 1x1xC
outputs = GlobalAveragePooling2D()(inputs)
# Add the activation method to the classifier
# HINT: what activation is used for a multi-class classifier?
outputs = Dense(n_classes, activation=??)(outputs)
return outputs
inputs = Input((32, 32, 3))
outputs = stem(inputs)
outputs = group(outputs, **{ 'blocks': [ { 'n_filters': 32 }, { 'n_filters': 64 }], 'k': 4 })
outputs = classifier(outputs, 10)
model = Model(inputs, outputs)
# -
# ### Verify the module using summary method
#
# It should look like below:
#
# ```
# Layer (type) Output Shape Param # Connected to
# ==================================================================================================
# input_6 (InputLayer) [(None, 32, 32, 3)] 0
# __________________________________________________________________________________________________
# conv2d_18 (Conv2D) (None, 32, 32, 16) 448 input_6[0][0]
# __________________________________________________________________________________________________
# batch_normalization_18 (BatchNo (None, 32, 32, 16) 64 conv2d_18[0][0]
# __________________________________________________________________________________________________
# re_lu_17 (ReLU) (None, 32, 32, 16) 0 batch_normalization_18[0][0]
# __________________________________________________________________________________________________
# batch_normalization_19 (BatchNo (None, 32, 32, 16) 64 re_lu_17[0][0]
# __________________________________________________________________________________________________
# conv2d_19 (Conv2D) (None, 32, 32, 32) 544 batch_normalization_19[0][0]
# __________________________________________________________________________________________________
# batch_normalization_20 (BatchNo (None, 32, 32, 32) 128 conv2d_19[0][0]
# __________________________________________________________________________________________________
# re_lu_18 (ReLU) (None, 32, 32, 32) 0 batch_normalization_20[0][0]
# __________________________________________________________________________________________________
# conv2d_20 (Conv2D) (None, 32, 32, 128) 36992 re_lu_18[0][0]
# __________________________________________________________________________________________________
# batch_normalization_21 (BatchNo (None, 32, 32, 128) 512 conv2d_20[0][0]
# __________________________________________________________________________________________________
# re_lu_19 (ReLU) (None, 32, 32, 128) 0 batch_normalization_21[0][0]
# __________________________________________________________________________________________________
# conv2d_21 (Conv2D) (None, 32, 32, 32) 36896 re_lu_19[0][0]
# __________________________________________________________________________________________________
# add_6 (Add) (None, 32, 32, 32) 0 conv2d_19[0][0]
# conv2d_21[0][0]
# __________________________________________________________________________________________________
# batch_normalization_22 (BatchNo (None, 32, 32, 32) 128 add_6[0][0]
# __________________________________________________________________________________________________
# conv2d_22 (Conv2D) (None, 32, 32, 64) 2112 batch_normalization_22[0][0]
# __________________________________________________________________________________________________
# batch_normalization_23 (BatchNo (None, 32, 32, 64) 256 conv2d_22[0][0]
# __________________________________________________________________________________________________
# re_lu_20 (ReLU) (None, 32, 32, 64) 0 batch_normalization_23[0][0]
# __________________________________________________________________________________________________
# conv2d_23 (Conv2D) (None, 32, 32, 256) 147712 re_lu_20[0][0]
# __________________________________________________________________________________________________
# batch_normalization_24 (BatchNo (None, 32, 32, 256) 1024 conv2d_23[0][0]
# __________________________________________________________________________________________________
# re_lu_21 (ReLU) (None, 32, 32, 256) 0 batch_normalization_24[0][0]
# __________________________________________________________________________________________________
# conv2d_24 (Conv2D) (None, 32, 32, 64) 147520 re_lu_21[0][0]
# __________________________________________________________________________________________________
# add_7 (Add) (None, 32, 32, 64) 0 conv2d_22[0][0]
# conv2d_24[0][0]
# __________________________________________________________________________________________________
# global_average_pooling2d (Globa (None, 64) 0 add_7[0][0]
# __________________________________________________________________________________________________
# dense (Dense) (None, 10) 650 global_average_pooling2d[0][0]
# ==================================================================================================
# Total params: 375,050
# Trainable params: 373,962
# Non-trainable params: 1,088
# __________________________________________________________________________________________________
# ```
model.summary()
# ## Training
#
# Finally, let's do a bit of training with your WRN model.
#
# ### Dataset
#
# Let's get the tf.Keras builtin dataset for CIFAR-10. These are 32x32 color images (3 channels) of 10 classes (airplanes, cars, birds, cats, deer, dogs, frogs, horses, ships, and trucks). We will preprocess the image data (not covered yet).
# +
from tensorflow.keras.datasets import cifar10
import numpy as np
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
x_train = (x_train / 255.0).astype(np.float32)
x_test = (x_test / 255.0).astype(np.float32)
# -
# ### Results
# Let's train the model for 3 epochs.
#
# Because it just a few epochs, you test accuracy may vary from run to run. For me, it was 52.8%
model.compile(loss='sparse_categorical_crossentropy', optimizer='adam', metrics=['acc'])
model.fit(x_train, y_train, epochs=3, batch_size=32, validation_split=0.1, verbose=1)
model.evaluate(x_test, y_test)
# ## End of Lab Exercise
| books/deep-learning-design-patterns/Workshops/Junior/Deep Learning Design Patterns - Workshop - Chapter 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Doubly Linked List Challenge
#
# Since it is a doubly linked list, we are going to change the pointers for both next and prev.
#
#
# +
# 1<-->2 <-->3 <-->4
# we need to make changes to both the next pointer and previous pointer
# when we start at the first node of 1:
# 1.prev would be None and 1.next would be 2.
# We swap it so that 1.next would be none and 1.prev would be 2.
#
def reverse(head):
if head.next != None:
head.next, head.prev, head = head.prev, head.next, head.next
# for the tail node i.e. NULL, the head.next would be head.prev i.e.3
# so we will take 3 and make it the next Node but head.prev would point to None
head.next, head.prev = head.prev, None
# -
| .ipynb_checkpoints/LinkedListChall-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.3 64-bit (''base'': conda)'
# name: python373jvsc74a57bd0f0396a0f98e081442f6005f4438dae70905c4dba32e635697d7a979ca5a56ea2
# ---
# # Paper: Backward bifurcations in dengue transmission dynamics
import numpy as np
from scipy.integrate import odeint
import matplotlib.pyplot as plt
# +
# Initial conditions
Sh0 = 100
Eh0 = 0
Ih = 0
Rh0 = 0
Sv0 = 1
Ev0 = 0
Iv0 = 0
# A grid of time points (in days)
t = np.linspace(0, 200, 50)
#parameters
pi_h = 10
pi_v = 30
lambdda_h = 0.05530
lambdda_v = 0.05
delta_h = 0.99
delta_v = 0.057
mu_h = 0.0195
mu_v = 0.016
sigma_h = 0.53
sigma_v = 0.2
tao_h = 0.1
# The SIR model differential equations.
def deriv(y, t, pi_h ,pi_v ,lambdda_h ,lambdda_v ,delta_h ,delta_v ,mu_h ,mu_v ,sigma_h ,sigma_v ,tao_h):
Sh ,Eh ,Ih ,Rh ,Sv ,Ev ,Iv = y
dShdt = pi_h - lambdda_h * Sh - mu_h * Sh
dEhdt = lambdda_h * Sh - (sigma_h * mu_h) * Eh
dIhdt = sigma_h * Eh - (tao_h + mu_h + delta_h) * Ih
dRhdt = tao_h * Ih - mu_h * Rh
dSvdt = pi_v - delta_v * Sv - mu_v * Sv
dEvdt = delta_v * Sv - (sigma_v + mu_v) * Ev
dIvdt = sigma_v * Ev - (mu_v + delta_v) * Iv
return dShdt,dEhdt,dIhdt,dRhdt,dSvdt,dEvdt, dIvdt
# Initial conditions vector
y0 = Sh0 ,Eh0 ,Ih ,Rh0 ,Sv0 ,Ev0 ,Iv0
# Integrate the SIR equations over the time grid, t.
ret = odeint(deriv, y0, t, args=(pi_h ,pi_v ,lambdda_h ,lambdda_v ,delta_h ,delta_v ,mu_h ,mu_v ,sigma_h ,sigma_v ,tao_h))
Sh ,Eh ,Ih ,Rh ,Sv ,Ev ,Iv = ret.T
# Plot the data on two separate curves for S(t), I(t)
fig = plt.figure(facecolor='w')
ax = fig.add_subplot(111, facecolor='#dddddd', axisbelow=True)
ax.plot(t, Sh, 'violet', alpha=0.5, lw=2, label='Sh', linestyle='dashed')
ax.plot(t, Eh, 'darkgreen', alpha=0.5, lw=2, label='Eh', linestyle='dashed')
ax.plot(t, Ih, 'blue', alpha=0.5, lw=2, label='Ih1', linestyle='dashed')
ax.plot(t, Rh, 'teal', alpha=0.5, lw=2, label='Rh', linestyle='dashed')
ax.plot(t, Sv, 'black', alpha=0.5, lw=2, label='Sv', linestyle='dashed')
ax.plot(t, Ev, 'green', alpha=0.5, lw=2, label='Ev', linestyle='dashed')
ax.plot(t, Iv, 'purple', alpha=0.5, lw=2, label='Iv', linestyle='dashed')
ax.set_xlabel('Time /days')
ax.yaxis.set_tick_params(length=0)
ax.xaxis.set_tick_params(length=0)
ax.grid(b=True, which='major', c='w', lw=2, ls='-')
legend = ax.legend()
legend.get_frame().set_alpha(0.5)
for spine in ('top', 'right', 'bottom', 'left'):
ax.spines[spine].set_visible(False)
plt.show()
# +
#save to csv file
a = np.asarray([t, Sh ,Eh ,Ih ,Rh ,Sv ,Ev ,Iv])
# np.savetxt("dengue.csv", a, delimiter=",")
# -
| Diseases/dengue/generate_dengue_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### <b> Course: Machine Learning for Absolute Beginners - Level 2
# #### By: <NAME>
# #### Exercise: #1 - Python Fundamentals
# *****************************
# ### Step 1 - Variables and Data Types
# Create the following variables:<br>
# num1=100<br>
# num2=300.0<br>
# str1="I would to learn data science"<br>
# list1=['A', 'B', 'C']<br>
# bol1=True
# Check the type of each variable that you created using the type() function. <br>
# Print the value of each variable using the print() function.
# Using one line assign the same value "100" to the following variables: num1, num2, num3.
# Using one line assign the following values to the following variables: num1=10, num2=20, num3=30.
# Check the size of the str1 string variable.
# Print in upper case and also in lower case the str1 variable (tip - use the tab completion to see the list of methods).
# Print the result (True or False) of checking if the text "data" exist in str1 using the in operator.
# Split str1 into substrings using the split method and store the result in a list variable called list1. Print list1.
# Define the following variables: Name='David', Phone='12345678'
# and then print the following message using the the f-string formatting: "My name is David and my phone number is 12345678"
# ### Step 2 - Lists
# Create the following list: colors = ['Red', 'Yellow', 'Green', 'Black', 'White'] and print the new list.
# Print the second color in the colors list.
# Sort and print the colors list in alphabetical order using the sort method.
# Add the the "Blue" color to the colors list and print the updated list.
# Remove the color 'Green' from the list by using the value and print the updated list.
# Check the size of a list.
# Create the following two lists: list1 = ['A','B','C'], list2 = ['D','E','F','G'] and then concatenate the two lists into list3 by using the + operator. Print list3.
# Print items 'B', 'C' and 'D' from list3 using the slicing notation.
# Print items 'C' until the end of list3 using the slicing notation.
# Copy list3 into a list4 using the slicing notation.
# Create the following tuple variable: tuple1=(1,2,3,4,5) and print it.
# Try to change the first item in the tuple list to the number 7.
# ### Step 3 - IF and For-Loop Statements
# Create the following list: colors = ['Red', 'Yellow', 'Green', 'Black', 'White'] and print the new list.
# Print "Yes" if the "Black" value inside the colors list.
# Print "Yes" if the "Black" value inside the colors list and the "Pink" value is not inside the colors list, otherwise print "No".
# Create the following variable: num1=50. Print 'More than 100' if the num1>=100. Print 'More than 40' if num1>=40.
# Scan the colors list and per each item in the list print the value and the index location.
# Print the numbers 1 to 30 using the range() function.
# Print the calculation of multiplying the numbers 10,15,20 with the numbers 1,2,3,4,5,6,7,8,9,10 (tip - use nested loops and the range function).
# ### Step 4 - Functions
# Define a function called check_color that is getting as input a list of colors and also a color value. If the color value exist in the list it will return 'Yes', otherwise it will return 'No'.
# Create the following list: colors = ['Red', 'Yellow', 'Green', 'Black', 'White'] and then call the function check_color to check if the 'Green' color in that list.
# Call the function check_color to check if the 'Pink' color in that list but this time while passing a copy of that list (tip - use the slice notation).
# Add a docstring comment to the check_color function definion and then ask Python to display information about the function (tip - use the ? mark).
# ### Step 5 - Dictionaries
# Create the following dictionaries friend1, friend2, friend3 with the following info:<br>friend1 with 'Name':'David', 'Age':34, 'Phone':556677.<br>friend2 with 'Name':'Marta', 'Age':29, 'Phone':223344.<br>friend3 with 'Name':'Mark', 'Age':45, 'Phone':333444.
# Print the 'Name' and 'Age' value for friend1.
# Print the complete friend2 dictionary.
# Add the following information to friend1, friend2, friend3: David 'Birthday' is '3.6', Marth 'Birthday' is '22.4', Mark 'Birthday' is '20.1'.
# Print friend1, friend2, friend3.
# Update friend2 phone number to: '112233'.
# Remove 'Phone' from friend1 and print the updated dictionaries.
# Create an empty list variable called myFriends.
# Add the three dictionaries: friend1, friend2, friend3 to the myFriends list.
# Print the first item in the myFriends list and then print just the 'Name' of the second item in the list."
# Loop through the myFriends list and check which friend is the oldest and print his/her name.
# ### Step 6 - Classes, Objects, Attributes, and Methods
# Define a new class called: Book with the following definition:
# 1. Attributes to be initalized during a new object creation: name, author, pages, price.
# 2. One method called update_price that will get as a parameter an updated price and adjust the object price attribute.
# Create two objects from the new class: book1, book2 with the following info:
# 1. book1: 'Python for Begineers', 'David', 240, 99
# 2. book2: 'Machine Learning for Begineers', 'Marta',320, 199
# Print the name and price of book1.
# Update the price of book1 to 79 using the object method and print the name and price again.
# ### Step 7 - Importing Modules
# Import the following modules:
# 1. random module with alias rd
# 2. math module
# 3. statistics module
# Generate 10 random integer numbers between 0 to 20 and store them in a list variable called myrandom. Print the new list.
# Calculate and print the following calculation : sin(0.5*pi) * (2^8).
# Print the arithmetic mean and the standard deviation of the numbers in the myrandom list.
# ### End of document
# ************************
| Exercise 1/Exercise #1 - Python Fundamentals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import urllib.request as urlreq
import urllib.error as urlerr
import urllib.parse as urlparse
import urllib.robotparser as urlrp
import datetime
import time
import sys
sys.path.append('../')
from common.utils import *
download("http://httpstat.us/500")
download("http://www.meetup.com")
# +
import re
def crawl_sitemap(url):
sitemap = download(url)
# decode from byte object to string
links = re.findall('<loc>(.*?)</loc>', sitemap.decode())
for link in links:
html = download(link)
# -
crawl_sitemap("http://www.mapscape.eu/sitemap.xml")
def link_crawler(seed_url, link_regex, max_depth=2):
crawl_queue = [seed_url]
seen = {seed_url:0}
throttle = Throttle(3)
user_agent = 'victor'
rp = urlrp.RobotFileParser()
rp.set_url("http://example.webscraping.com/robots.txt")
rp.read()
while crawl_queue:
url = crawl_queue.pop()
depth = seen[url]
if depth != max_depth:
if rp.can_fetch(user_agent, url):
throttle.wait(url)
html = download(url, user_agent)
for link in get_links(html.decode()):
if re.search(link_regex, link):
# form absolute link
link = urlparse.urljoin(seed_url, link)
# check if this link is already seen
if link not in seen:
seen[link] = depth + 1
crawl_queue.append(link)
else:
print('blocked by robots.txt, ', url)
return seen
all_links = link_crawler('http://example.webscraping.com', '/(index|view)/')
| chapter 1/download website.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/esohman/Waseda-DH/blob/main/JDMDH_RedditScrape.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Fpmpa8SEvog_"
# # Scraping Reddit Data
# + id="-5za1HPNvohF" colab={"base_uri": "https://localhost:8080/"} outputId="b9a00d34-05aa-4645-a2a6-fa6ac55eda54"
# !pip install praw
import praw
# + id="Jy_qM67VvohH"
reddit = praw.Reddit(client_id='anon',
client_secret='anon',
user_agent='anon')
# + id="29i-ZD47vohO"
import pandas as pd
submission = reddit.submission(url="https://www.reddit.com/r/announcements/comments/9jf8nh/revamping_the_quarantine_function/") #replace with target post
# or
#submission = reddit.submission(id="3djjxw") #id comes after comments/
# + id="2pvCzUc7vohQ" outputId="babe7f63-30fe-436a-a65b-b60a62eb18f8" colab={"base_uri": "https://localhost:8080/", "height": 1000}
submission.comments.replace_more(limit=None)
comments = []
for comment in submission.comments.list():
comments.append([comment.body, comment.score, comment.id, comment.author, comment.replies, comment.parent_id, comment.created_utc])
comments = pd.DataFrame(comments,columns=['comment', 'score', 'id', 'author', 'replies', 'parent', 'created'])
comments
# + id="XJhSsg3P1myz"
comments.to_csv("content18.csv") #change name as appropriate
| Reddit Content Moderation/JDMDH_RedditScrape.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Convolutional Variational Autoencoder - MNIST
#
# Slightly modified from variational_autoencoder_deconv.py in the Keras examples folder:
#
# **https://github.com/fchollet/keras/blob/master/examples/variational_autoencoder_deconv.py**
WEIGHTS_FILEPATH = 'mnist_vae.hdf5'
MODEL_ARCH_FILEPATH = 'mnist_vae.json'
# +
import numpy as np
np.random.seed(1337) # for reproducibility
import matplotlib.pyplot as plt
from keras.layers import Input, Dense, Lambda, Flatten, Reshape
from keras.layers import Convolution2D, Deconvolution2D
from keras.models import Model
from keras import backend as K
from keras import objectives
from keras.datasets import mnist
from keras.callbacks import EarlyStopping, ModelCheckpoint
# +
# input image dimensions
img_rows, img_cols, img_chns = 28, 28, 1
# number of convolutional filters to use
nb_filters = 64
# convolution kernel size
nb_conv = 3
batch_size = 200
original_img_size = (img_rows, img_cols, img_chns)
latent_dim = 2
intermediate_dim = 128
epsilon_std = 0.01
x = Input(batch_shape=(batch_size,) + original_img_size)
conv_1 = Convolution2D(img_chns, 2, 2, border_mode='same', activation='relu', dim_ordering='tf')(x)
conv_2 = Convolution2D(nb_filters, 2, 2,
border_mode='same', activation='relu', dim_ordering='tf',
subsample=(2, 2))(conv_1)
conv_3 = Convolution2D(nb_filters, nb_conv, nb_conv,
border_mode='same', activation='relu', dim_ordering='tf',
subsample=(1, 1))(conv_2)
conv_4 = Convolution2D(nb_filters, nb_conv, nb_conv,
border_mode='same', activation='relu', dim_ordering='tf',
subsample=(1, 1))(conv_3)
flat = Flatten()(conv_4)
hidden = Dense(intermediate_dim, activation='relu')(flat)
z_mean = Dense(latent_dim)(hidden)
z_log_var = Dense(latent_dim)(hidden)
def sampling(args):
z_mean, z_log_var = args
epsilon = K.random_normal(shape=(batch_size, latent_dim),
mean=0., std=epsilon_std)
return z_mean + K.exp(z_log_var) * epsilon
# note that "output_shape" isn't necessary with the TensorFlow backend
# so you could write `Lambda(sampling)([z_mean, z_log_var])`
z = Lambda(sampling, output_shape=(latent_dim,))([z_mean, z_log_var])
# we instantiate these layers separately so as to reuse them later
decoder_hid = Dense(intermediate_dim, activation='relu')
decoder_upsample = Dense(nb_filters * 14 * 14, activation='relu')
output_shape = (batch_size, 14, 14, nb_filters)
decoder_reshape = Reshape(output_shape[1:])
decoder_deconv_1 = Deconvolution2D(nb_filters, nb_conv, nb_conv,
output_shape,
border_mode='same',
subsample=(1, 1),
activation='relu', dim_ordering='tf')
decoder_deconv_2 = Deconvolution2D(nb_filters, nb_conv, nb_conv,
output_shape,
border_mode='same',
subsample=(1, 1),
activation='relu', dim_ordering='tf')
output_shape = (batch_size, 29, 29, nb_filters)
decoder_deconv_3_upsamp = Deconvolution2D(nb_filters, 2, 2,
output_shape,
border_mode='valid',
subsample=(2, 2),
activation='relu', dim_ordering='tf')
decoder_mean_squash = Convolution2D(img_chns, 2, 2,
border_mode='valid',
activation='sigmoid', dim_ordering='tf')
hid_decoded = decoder_hid(z)
up_decoded = decoder_upsample(hid_decoded)
reshape_decoded = decoder_reshape(up_decoded)
deconv_1_decoded = decoder_deconv_1(reshape_decoded)
deconv_2_decoded = decoder_deconv_2(deconv_1_decoded)
x_decoded_relu = decoder_deconv_3_upsamp(deconv_2_decoded)
x_decoded_mean_squash = decoder_mean_squash(x_decoded_relu)
def vae_loss(x, x_decoded_mean):
# NOTE: binary_crossentropy expects a batch_size by dim
# for x and x_decoded_mean, so we MUST flatten these!
x = K.flatten(x)
x_decoded_mean = K.flatten(x_decoded_mean)
xent_loss = img_rows * img_cols * objectives.binary_crossentropy(x, x_decoded_mean)
kl_loss = - 0.5 * K.mean(1 + z_log_var - K.square(z_mean) - K.exp(z_log_var), axis=-1)
return xent_loss + kl_loss
vae = Model(x, x_decoded_mean_squash)
vae.compile(optimizer='adam', loss=vae_loss)
# +
nb_epoch = 100
# train the VAE on MNIST digits
(x_train, _), (x_test, y_test) = mnist.load_data()
x_train = x_train.astype('float32') / 255.
x_train = x_train.reshape((x_train.shape[0],) + original_img_size)
x_test = x_test.astype('float32') / 255.
x_test = x_test.reshape((x_test.shape[0],) + original_img_size)
print('x_train.shape:', x_train.shape)
# Early stopping
early_stopping = EarlyStopping(monitor='val_loss', verbose=1, patience=5)
vae.fit(x_train, x_train,
validation_data=(x_test, x_test),
shuffle=True, nb_epoch=nb_epoch, batch_size=batch_size, verbose=2,
callbacks=[early_stopping])
# -
# %matplotlib inline
# +
# build a model to project inputs on the latent space
encoder = Model(x, z_mean)
# display a 2D plot of the digit classes in the latent space
x_test_encoded = encoder.predict(x_test, batch_size=batch_size)
plt.figure(figsize=(6, 6))
plt.scatter(x_test_encoded[:, 0], x_test_encoded[:, 1], c=y_test)
plt.colorbar()
plt.show()
# -
# **Decoder generator**
#
# To make the decoder generator serializable, we will redefine new layers and transfer weights over, rather than sharing the layers. Sharing layers will create new nodes, some with different output shapes, which causes problems for serialization.
#
# Here we also set batch_size to 1
# +
batch_size = 1
_hid_decoded = Dense(intermediate_dim, activation='relu')
_up_decoded = Dense(nb_filters * 14 * 14, activation='relu')
_reshape_decoded = Reshape((14, 14, nb_filters))
_deconv_1_decoded = Deconvolution2D(nb_filters, nb_conv, nb_conv, (batch_size, 14, 14, nb_filters),
border_mode='same', subsample=(1, 1), activation='relu', dim_ordering='tf')
_deconv_2_decoded = Deconvolution2D(nb_filters, nb_conv, nb_conv, (batch_size, 14, 14, nb_filters),
border_mode='same', subsample=(1, 1), activation='relu', dim_ordering='tf')
_x_decoded_relu = Deconvolution2D(nb_filters, 2, 2, (batch_size, 29, 29, nb_filters),
border_mode='valid', subsample=(2, 2), activation='relu', dim_ordering='tf')
_x_decoded_mean_squash = Convolution2D(img_chns, 2, 2, border_mode='valid', activation='sigmoid', dim_ordering='tf')
decoder_input = Input(shape=(latent_dim,))
layer1 = _hid_decoded(decoder_input)
layer2 = _up_decoded(layer1)
layer3 = _reshape_decoded(layer2)
layer4 = _deconv_1_decoded(layer3)
layer5 = _deconv_2_decoded(layer4)
layer6 = _x_decoded_relu(layer5)
layer7 = _x_decoded_mean_squash(layer6)
generator = Model(decoder_input, layer7)
_hid_decoded.set_weights(decoder_hid.get_weights())
_up_decoded.set_weights(decoder_upsample.get_weights())
_deconv_1_decoded.set_weights(decoder_deconv_1.get_weights())
_deconv_2_decoded.set_weights(decoder_deconv_2.get_weights())
_x_decoded_relu.set_weights(decoder_deconv_3_upsamp.get_weights())
_x_decoded_mean_squash.set_weights(decoder_mean_squash.get_weights())
# display a 2D manifold of the digits
n = 15 # figure with 15x15 digits
digit_size = 28
figure = np.zeros((digit_size * n, digit_size * n))
# we will sample n points within [-1.5, 1.5] standard deviations
grid_x = np.linspace(-1.5, 1.5, n)
grid_y = np.linspace(-1.5, 1.5, n)
for i, yi in enumerate(grid_x):
for j, xi in enumerate(grid_y):
z_sample = np.array([[xi, yi]])
z_sample = np.tile(z_sample, batch_size).reshape(batch_size, 2)
x_decoded = generator.predict(z_sample, batch_size=batch_size)
digit = x_decoded[0].reshape(digit_size, digit_size)
figure[i * digit_size: (i + 1) * digit_size,
j * digit_size: (j + 1) * digit_size] = digit
plt.figure(figsize=(10, 10))
plt.imshow(figure)
plt.show()
# +
generator.save_weights(WEIGHTS_FILEPATH)
with open(MODEL_ARCH_FILEPATH, 'w') as f:
f.write(generator.to_json())
# -
| demos/notebooks/mnist_vae.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# https://www.kaggle.com/ibtesama/getting-started-with-a-movie-recommendation-system
# + [markdown] _uuid="d7aa1b0ad6979877450f9cd89e1e37289b51cf6e"
# # **The Age of Recommender Systems**
# + [markdown] _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# The rapid growth of data collection has led to a new era of information. Data is being used to create more efficient systems and this is where Recommendation Systems come into play. Recommendation Systems are a type of **information filtering systems** as they improve the quality of search results and provides items that are more relevant to the search item or are realted to the search history of the user.
#
# + [markdown] _uuid="65dbae55f1e6e06c5fa7251f8ddae887d3fbf480"
# They are used to predict the **rating** or **preference** that a user would give to an item. Almost every major tech company has applied them in some form or the other: Amazon uses it to suggest products to customers, YouTube uses it to decide which video to play next on autoplay, and Facebook uses it to recommend pages to like and people to follow.
# Moreover, companies like Netflix and Spotify depend highly on the effectiveness of their recommendation engines for their business and sucees.
# + [markdown] _uuid="af1e8d514096cde4abeae2d65a44cfdb01228d77"
# 
# + [markdown] _uuid="f34fc4dcfba717692c620e1fdfa502ee910c6365"
# In this kernel we'll be building a baseline Movie Recommendation System using [TMDB 5000 Movie Dataset](https://www.kaggle.com/tmdb/tmdb-movie-metadata). For novices like me this kernel will pretty much serve as a foundation in recommendation systems and will provide you with something to start with.
# + [markdown] _uuid="1fa8fc3cb8348853bb51472b248134a59c24bf60"
# **So let's go!**
# + [markdown] _uuid="f9a9405b7e81c1da449bd2e96c2849fb86caa614"
# There are basically three types of recommender systems:-
#
# > * **Demographic Filtering**- They offer generalized recommendations to every user, based on movie popularity and/or genre. The System recommends the same movies to users with similar demographic features. Since each user is different , this approach is considered to be too simple. The basic idea behind this system is that movies that are more popular and critically acclaimed will have a higher probability of being liked by the average audience.
#
#
# + [markdown] _uuid="60a2df15abf82ba21918e3a42cb0ee46d22fa764"
# > * **Content Based Filtering**- They suggest similar items based on a particular item. This system uses item metadata, such as genre, director, description, actors, etc. for movies, to make these recommendations. The general idea behind these recommender systems is that if a person liked a particular item, he or she will also like an item that is similar to it.
# + [markdown] _uuid="b7e719fcc502c59f098a51ae35e2ceb6d7cdfe4e"
# > * **Collaborative Filtering**- This system matches persons with similar interests and provides recommendations based on this matching. Collaborative filters do not require item metadata like its content-based counterparts.
# + [markdown] _uuid="6b418588e3f9139f74cb3a9546f5dca49729579b"
# Let's load the data now.
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
pd.set_option('display.max_columns', 500)
pd.set_option('display.max_colwidth', 500)
pd.set_option('display.expand_frame_repr', True)
# + _uuid="c1fdd129c1cbab68ae3e6bf2062575f01f80b87c"
df1 = pd.read_csv('tmdb-5000-movie-dataset/tmdb_5000_credits.csv')
df2 = pd.read_csv('tmdb-5000-movie-dataset/tmdb_5000_movies.csv')
df1.shape, df2.shape
# -
df1.head(2)
df2.head(2)
# + [markdown] _uuid="402a28d17c13bba3f2060d72c2ff75f5377a9f01"
# The first dataset contains the following features:-
#
# * movie_id - A unique identifier for each movie.
# * cast - The name of lead and supporting actors.
# * crew - The name of Director, Editor, Composer, Writer etc.
#
# The second dataset has the following features:-
#
# * budget - The budget in which the movie was made.
# * genre - The genre of the movie, Action, Comedy ,Thriller etc.
# * homepage - A link to the homepage of the movie.
# * id - This is infact the movie_id as in the first dataset.
# * keywords - The keywords or tags related to the movie.
# * original_language - The language in which the movie was made.
# * original_title - The title of the movie before translation or adaptation.
# * overview - A brief description of the movie.
# * popularity - A numeric quantity specifying the movie popularity.
# * production_companies - The production house of the movie.
# * production_countries - The country in which it was produced.
# * release_date - The date on which it was released.
# * revenue - The worldwide revenue generated by the movie.
# * runtime - The running time of the movie in minutes.
# * status - "Released" or "Rumored".
# * tagline - Movie's tagline.
# * title - Title of the movie.
# * vote_average - average ratings the movie recieved.
# * vote_count - the count of votes recieved.
#
# Let's join the two dataset on the 'id' column
#
# + _uuid="c87bda9d56a936be126d03eda0bc743ee35be461"
df1.columns = ['id','tittle','cast','crew']
df2 = df2.merge(df1,on='id')
df2.shape
# + [markdown] _uuid="e88ed16f798ec3a094fce8aaf8a971f7b2aae83e"
# Just a peak at our data.
# + _uuid="71d266ed92947c51acf07189d3b42379134ef6e7"
df2.head(5)
# + [markdown] _uuid="ee603279675033fc397f0c94738e20b34f35312b"
# # **Demographic Filtering** -
# Before getting started with this -
# * we need a metric to score or rate movie
# * Calculate the score for every movie
# * Sort the scores and recommend the best rated movie to the users.
#
# We can use the average ratings of the movie as the score but using this won't be fair enough since a movie with 8.9 average rating and only 3 votes cannot be considered better than the movie with 7.8 as as average rating but 40 votes.
# So, I'll be using IMDB's weighted rating (wr) which is given as :-
#
# 
# where,
# * v is the number of votes for the movie;
# * m is the minimum votes required to be listed in the chart;
# * R is the average rating of the movie; And
# * C is the mean vote across the whole report
#
# We already have v(**vote_count**) and R (**vote_average**) and C can be calculated as
# + _uuid="5799b99c5e5ed5b7723ae8b31e1fc9fb1e7b89ec"
C = df2['vote_average'].mean()
C
# + [markdown] _uuid="02fa8642f75bcef6d4a79e029af6dfeebf19e3a5"
# So, the mean rating for all the movies is approx 6 on a scale of 10.The next step is to determine an appropriate value for m, the minimum votes required to be listed in the chart. We will use 90th percentile as our cutoff. In other words, for a movie to feature in the charts, it must have more votes than at least 90% of the movies in the list.
# + _uuid="f2f1eaff1e4349b5d2d11dd5ce79c19a85561148"
m = df2['vote_count'].quantile(0.9)
m
# + [markdown] _uuid="b77dea5a38ca2c399e3abeac1487e784fe146078"
# Now, we can filter out the movies that qualify for the chart
# + _uuid="a22008df6d81d3b716d39a56efd3d547345bfbce"
q_movies = df2.copy().loc[df2['vote_count'] >= m]
q_movies.shape
# + [markdown] _uuid="cf1fe5a3692caee41a6e7a74c3fde6aeb8a62947"
# We see that there are 481 movies which qualify to be in this list. Now, we need to calculate our metric for each qualified movie. To do this, we will define a function, **weighted_rating()** and define a new feature **score**, of which we'll calculate the value by applying this function to our DataFrame of qualified movies:
# + _uuid="bb680ed0fb1c3020785d34152c57c6e2279d4424"
def weighted_rating(x, m=m, C=C):
v = x['vote_count']
R = x['vote_average']
# Calculation based on the IMDB formula
return (v/(v+m) * R) + (m/(m+v) * C)
# + _uuid="d2d189929715237ab19a18fb8747239b86092968"
# Define a new feature 'score' and calculate its value with `weighted_rating()`
q_movies['score'] = q_movies.apply(weighted_rating, axis=1)
# + [markdown] _uuid="4cdd60c146173606146ec4fc3a1c9d8c184cb81c"
# Finally, let's sort the DataFrame based on the score feature and output the title, vote count, vote average and weighted rating or score of the top 10 movies.
# + _uuid="a9a9fc3810ea67c31908bbdf8bb930daa918102b"
#Sort movies based on score calculated above
q_movies = q_movies.sort_values('score', ascending=False)
#Print the top 15 movies
q_movies[['title', 'vote_count', 'vote_average', 'score']].head(10)
# + [markdown] _uuid="7625c6d5bd647f4f75eafd7880b7ca2d07938cf7"
# Hurray! We have made our first(though very basic) recommender.
# Under the **Trending Now** tab of these systems we find movies that are very popular and they can just be obtained by sorting the dataset by the popularity column.
# + _uuid="207f7058f92698b5fd776f7771a3ac0cc2928bf1"
pop = df2.sort_values('popularity', ascending=False)
plt.figure(figsize=(12,4))
plt.barh(pop['title'].head(6), pop['popularity'].head(6), align='center', color='skyblue')
plt.gca().invert_yaxis()
plt.xlabel('Popularity')
plt.title('Popular Movies')
plt.show()
# + [markdown] _uuid="d1cf28fc0c22afa80edc6d464177035357721d35"
# Now something to keep in mind is that these demographic recommender provide a general chart of recommended movies to all the users. They are not sensitive to the interests and tastes of a particular user.
# This is when we move on to a more refined system- Content Basesd Filtering.
# + [markdown] _uuid="fe716df6e5e5a354ac53d556087147c0a64df2cc"
# # **Content Based Filtering**
# In this recommender system the content of the movie (overview, cast, crew, keyword, tagline etc) is used to find its similarity with other movies. Then the movies that are most likely to be similar are recommended.
#
# 
# + [markdown] _uuid="b0a813c803b0ba1f0204188ab2a63dc7f59ce2eb"
# ## **Plot description based Recommender**
#
# We will compute pairwise similarity scores for all movies based on their plot descriptions and recommend movies based on that similarity score. The plot description is given in the **overview** feature of our dataset.
# Let's take a look at the data. ..
# + _uuid="5e676c38ace04a24205b76b16dac0fa3e058027f"
df2['overview'].head(5)
# + [markdown] _uuid="277a9bb5b00a6bd2469c45777f9c659066f402b3"
# For any of you who has done even a bit of text processing before knows we need to convert the word vector of each overview.
# Now we'll compute Term Frequency-Inverse Document Frequency (TF-IDF) vectors for each overview.
#
# Now if you are wondering what is term frequency , it is the relative frequency of a word in a document and is given as
# **(term instances/total instances)**.
# Inverse Document Frequency is the relative count of documents containing the term is given as
# **log(number of documents/documents with term)**
# The overall importance of each word to the documents in which they appear is equal to **TF * IDF**
#
# This will give you a matrix where each column represents a word in the overview vocabulary (all the words that appear in at least one document) and each column represents a movie, as before.This is done to reduce the importance of words that occur frequently in plot overviews and therefore, their significance in computing the final similarity score.
#
# Fortunately, scikit-learn gives you a built-in TfIdfVectorizer class that produces the TF-IDF matrix in a couple of lines. That's great, isn't it?
# + _uuid="a92da8cde39c61deef5a1b8efa31ed84cda7f5fe"
#Import TfIdfVectorizer from scikit-learn
from sklearn.feature_extraction.text import TfidfVectorizer
#Define a TF-IDF Vectorizer Object. Remove all english stop words such as 'the', 'a'
tfidf = TfidfVectorizer(stop_words='english')
#Replace NaN with an empty string
df2['overview'] = df2['overview'].fillna('')
#Construct the required TF-IDF matrix by fitting and transforming the data
tfidf_matrix = tfidf.fit_transform(df2['overview'])
#Output the shape of tfidf_matrix
tfidf_matrix.shape
# -
tfidf_matrix[0]
# + [markdown] _uuid="6bde57434bf9a0e8f8b229d36901d75b77ff962f"
# We see that over 20,000 different words were used to describe the 4800 movies in our dataset.
#
# With this matrix in hand, we can now compute a similarity score. There are several candidates for this; such as the euclidean, the Pearson and the [cosine similarity scores](https://en.wikipedia.org/wiki/Cosine_similarity). There is no right answer to which score is the best. Different scores work well in different scenarios and it is often a good idea to experiment with different metrics.
#
# We will be using the cosine similarity to calculate a numeric quantity that denotes the similarity between two movies. We use the cosine similarity score since it is independent of magnitude and is relatively easy and fast to calculate. Mathematically, it is defined as follows:
# 
# + [markdown] _uuid="c1c138ae64648cb5a94127a06441d88dddc2bd9a"
# Since we have used the TF-IDF vectorizer, calculating the dot product will directly give us the cosine similarity score. Therefore, we will use sklearn's **linear_kernel()** instead of cosine_similarities() since it is faster.
# + _uuid="5eb17d12220eecab4faf01bbfd13e79d8e446537"
# Import linear_kernel
from sklearn.metrics.pairwise import linear_kernel
# Compute the cosine similarity matrix
cosine_sim = linear_kernel(tfidf_matrix, tfidf_matrix)
type(cosine_sim)
# -
cosine_sim.shape
cosine_sim[:5]
# + [markdown] _uuid="f5ca112fbbe25b11f0f3356a31d1604727242700"
# We are going to define a function that takes in a movie title as an input and outputs a list of the 10 most similar movies. Firstly, for this, we need a reverse mapping of movie titles and DataFrame indices. In other words, we need a mechanism to identify the index of a movie in our metadata DataFrame, given its title.
# + _uuid="55df2df36be98e6dec5f617a5aa51b77c500faa4"
#Construct a reverse map of indices and movie titles
indices = pd.Series(df2.index, index=df2['title']).drop_duplicates()
indices.shape
# -
indices[:5]
"""The 1st tuple (t1) has t1[1]=1.0 similarity score because it is comparing with itself"""
list(enumerate(cosine_sim[0]))[:5]
# + [markdown] _uuid="da5896c6ccfd44c3347af3097275d0aa707c1001"
# We are now in a good position to define our recommendation function. These are the following steps we'll follow :-
# * Get the index of the movie given its title.
# * Get the list of cosine similarity scores for that particular movie with all movies. Convert it into a list of tuples where the first element is its position and the second is the similarity score.
# * Sort the aforementioned list of tuples based on the similarity scores; that is, the second element.
# * Get the top 10 elements of this list. Ignore the first element as it refers to self (the movie most similar to a particular movie is the movie itself).
# * Return the titles corresponding to the indices of the top elements.
# + _uuid="9c383fcbb916dce464b01adf980d26ad96aebe0e"
# Function that takes in movie title as input and outputs most similar movies
def get_recommendations(title, cosine_sim=cosine_sim):
# Get the index of the movie that matches the title
idx = indices[title]
# Get the pairwsie similarity scores of all movies with that movie
sim_scores = list(enumerate(cosine_sim[idx]))
# Sort the movies based on the similarity scores
sim_scores = sorted(sim_scores, key=lambda x: x[1], reverse=True)
# Get the scores of the 10 most similar movies
sim_scores = sim_scores[1:11]
# Get the movie indices
movie_indices = [i[0] for i in sim_scores]
# Return the top 10 most similar movies
return df2['title'].iloc[movie_indices]
# + _uuid="14d722124f82e69cb444adcc589e396c75cbb4ff"
get_recommendations('The Dark Knight Rises')
# + _uuid="902b9f1ab91921889c85e9008818dcc0b4710ccd"
get_recommendations('The Avengers')
# + [markdown] _uuid="146302c25776b2c0076e64663a5e2e41e977fd2c"
# While our system has done a decent job of finding movies with similar plot descriptions, the quality of recommendations is not that great. "The Dark Knight Rises" returns all Batman movies while it is more likely that the people who liked that movie are more inclined to enjoy other Christopher Nolan movies. This is something that cannot be captured by the present system.
# + [markdown] _uuid="fcfe9db9c2fdd9334538256d233c6acf33c1c049"
# ## **Credits, Genres and Keywords Based Recommender**
# It goes without saying that the quality of our recommender would be increased with the usage of better metadata. That is exactly what we are going to do in this section. We are going to build a recommender based on the following metadata: the 3 top actors, the director, related genres and the movie plot keywords.
#
# From the cast, crew and keywords features, we need to extract the three most important actors, the director and the keywords associated with that movie. Right now, our data is present in the form of "stringified" lists , we need to convert it into a safe and usable structure
# + _uuid="59a8d0991e3cae9a44a4b351e154fd1000724448"
# Parse the stringified features into their corresponding python objects
from ast import literal_eval
features = ['cast', 'crew', 'keywords', 'genres']
for feature in features:
df2[feature] = df2[feature].apply(literal_eval)
# -
df2.head(2)
# + [markdown] _uuid="47d6062d1622a163f2bcf80b79eb7b1454003739"
# Next, we'll write functions that will help us to extract the required information from each feature.
# + _uuid="783b0e89f1c04a12ff51eb29cc68e93c818896cd"
# Get the director's name from the crew feature. If director is not listed, return NaN
def get_director(x):
for i in x:
if i['job'] == 'Director':
return i['name']
return np.nan
# + _uuid="86c4e9f4e6ef1e5ff287f58f3a1119fbddbdae09"
# Returns the list top 3 elements or entire list; whichever is more.
def get_list(x):
if isinstance(x, list):
names = [i['name'] for i in x]
#Check if more than 3 elements exist. If yes, return only first three. If no, return entire list.
if len(names) > 3:
names = names[:3]
return names
#Return empty list in case of missing/malformed data
return []
# + _uuid="dd060c3c1d724de71555218f30cccafd4a8ad6af"
# Define new director, cast, genres and keywords features that are in a suitable form.
df2['director'] = df2['crew'].apply(get_director)
features = ['cast', 'keywords', 'genres']
for feature in features:
df2[feature] = df2[feature].apply(get_list)
# + _uuid="87a96f835470aa3df590b74322c2717ff529d6ae"
# Print the new features of the first 3 films
df2[['title', 'cast', 'director', 'keywords', 'genres']].head(3)
# + [markdown] _uuid="0bcb2c2e99ffd9ce73205c9c6ef6687d16caa31f"
# The next step would be to convert the names and keyword instances into lowercase and strip all the spaces between them. This is done so that our vectorizer doesn't count the Johnny of "<NAME>" and "<NAME>" as the same.
# + _uuid="86af764c406a8b6184b37b57cfe499d20ce45f9c"
# Function to convert all strings to lower case and strip names of spaces
def clean_data(x):
if isinstance(x, list):
return [str.lower(i.replace(" ", "")) for i in x]
else:
#Check if director exists. If not, return empty string
if isinstance(x, str):
return str.lower(x.replace(" ", ""))
else:
return ''
# + _uuid="5728cc017ff6ed1dcd79da05b1dd57a60557e853"
# Apply clean_data function to your features.
features = ['cast', 'keywords', 'director', 'genres']
for feature in features:
df2[feature] = df2[feature].apply(clean_data)
# + _uuid="87a96f835470aa3df590b74322c2717ff529d6ae"
# Print the new features of the first 3 films
df2[['title', 'cast', 'director', 'keywords', 'genres']].head(3)
# + [markdown] _uuid="b6b3e1c480a7c280fbe81e63c5c4cf3ce308dc28"
# We are now in a position to create our "metadata soup", which is a string that contains all the metadata that we want to feed to our vectorizer (namely actors, director and keywords).
# + _uuid="20aef87703c408926f7617573ed043605207767f"
def create_soup(x):
return ' '.join(x['keywords']) + ' ' + ' '.join(x['cast']) + ' ' + x['director'] + ' ' + ' '.join(x['genres'])
df2['soup'] = df2.apply(create_soup, axis=1)
# -
df2[['title', 'cast', 'director', 'keywords', 'genres', 'soup']].head(3)
# + [markdown] _uuid="7b79886883806b8fb58098f9f803dabeaa0cadf6"
# The next steps are the same as what we did with our plot description based recommender. One important difference is that we use the **CountVectorizer()** instead of TF-IDF. This is because we do not want to down-weight the presence of an actor/director if he or she has acted or directed in relatively more movies. It doesn't make much intuitive sense.
# + _uuid="b66a1afc1083917d5ef136ccdcd9b50cca087e2b"
# Import CountVectorizer and create the count matrix
from sklearn.feature_extraction.text import CountVectorizer
count = CountVectorizer(stop_words='english')
count_matrix = count.fit_transform(df2['soup'])
# -
type(count_matrix)
count_matrix.shape
# + _uuid="3fa5539ed1680ed5323f8351ac7e4840f629e958"
# Compute the Cosine Similarity matrix based on the count_matrix
from sklearn.metrics.pairwise import cosine_similarity
cosine_sim2 = cosine_similarity(count_matrix, count_matrix)
type(cosine_sim2)
# -
cosine_sim2.shape
# + _uuid="b2b8565a04f4bda92d3ba9d15c348af1cd8f8b4d"
# Reset index of our main DataFrame and construct reverse mapping as before
df2 = df2.reset_index()
df2.head(1)
# -
indices = pd.Series(df2.index, index=df2['title'])
indices.head(1)
# + [markdown] _uuid="3c5a3a44893f63aa558030d03e228a365d10d91f"
# We can now reuse our **get_recommendations()** function by passing in the new **cosine_sim2** matrix as your second argument.
# + _uuid="d1e0e02be7a9e71422d3a492834cb4f8434d1464"
get_recommendations('The Dark Knight Rises', cosine_sim2)
# + _uuid="d6c4df85a80d830b2905f69e0e59ebb3461db3b7"
get_recommendations('The Godfather', cosine_sim2)
# + [markdown] _uuid="4d963ff547ee4980c0f23840394046d805fda574"
# We see that our recommender has been successful in capturing more information due to more metadata and has given us (arguably) better recommendations. It is more likely that Marvels or DC comics fans will like the movies of the same production house. Therefore, to our features above we can add *production_company* .
# We can also increase the weight of the director , by adding the feature multiple times in the soup.
# + [markdown] _uuid="71b15b5c090694303fa5e8d67b8bf394e07f45d6"
# # **Collaborative Filtering**
#
# Our content based engine suffers from some severe limitations. It is only capable of suggesting movies which are close to a certain movie. That is, it is not capable of capturing tastes and providing recommendations across genres.
#
# Also, the engine that we built is not really personal in that it doesn't capture the personal tastes and biases of a user. Anyone querying our engine for recommendations based on a movie will receive the same recommendations for that movie, regardless of who she/he is.
#
# Therefore, in this section, we will use a technique called Collaborative Filtering to make recommendations to Movie Watchers.
# It is basically of two types:-
#
# * **User based filtering**- These systems recommend products to a user that similar users have liked. For measuring the similarity between two users we can either use pearson correlation or cosine similarity.
# This filtering technique can be illustrated with an example. In the following matrixes, each row represents a user, while the columns correspond to different movies except the last one which records the similarity between that user and the target user. Each cell represents the rating that the user gives to that movie. Assume user E is the target.
# 
#
# Since user A and F do not share any movie ratings in common with user E, their similarities with user E are not defined in Pearson Correlation. Therefore, we only need to consider user B, C, and D. Based on Pearson Correlation, we can compute the following similarity.
# 
#
# From the above table we can see that user D is very different from user E as the Pearson Correlation between them is negative. He rated Me Before You higher than his rating average, while user E did the opposite. Now, we can start to fill in the blank for the movies that user E has not rated based on other users.
# 
#
# Although computing user-based CF is very simple, it suffers from several problems. One main issue is that users’ preference can change over time. It indicates that precomputing the matrix based on their neighboring users may lead to bad performance. To tackle this problem, we can apply item-based CF.
#
# * **Item Based Collaborative Filtering** - Instead of measuring the similarity between users, the item-based CF recommends items based on their similarity with the items that the target user rated. Likewise, the similarity can be computed with Pearson Correlation or Cosine Similarity. The major difference is that, with item-based collaborative filtering, we fill in the blank vertically, as oppose to the horizontal manner that user-based CF does. The following table shows how to do so for the movie Me Before You.
# 
#
# It successfully avoids the problem posed by dynamic user preference as item-based CF is more static. However, several problems remain for this method. First, the main issue is ***scalability***. The computation grows with both the customer and the product. The worst case complexity is O(mn) with m users and n items. In addition, ***sparsity*** is another concern. Take a look at the above table again. Although there is only one user that rated both Matrix and Titanic rated, the similarity between them is 1. In extreme cases, we can have millions of users and the similarity between two fairly different movies could be very high simply because they have similar rank for the only user who ranked them both.
#
#
# + [markdown] _uuid="4307f75107f9c5e5f911d52a6f1dc5530990c75e"
# ### **Single Value Decomposition**
# One way to handle the scalability and sparsity issue created by CF is to leverage a **latent factor model** to capture the similarity between users and items. Essentially, we want to turn the recommendation problem into an optimization problem. We can view it as how good we are in predicting the rating for items given a user. One common metric is Root Mean Square Error (RMSE). **The lower the RMSE, the better the performance**.
#
# Now talking about latent factor you might be wondering what is it ?It is a broad idea which describes a property or concept that a user or an item have. For instance, for music, latent factor can refer to the genre that the music belongs to. SVD decreases the dimension of the utility matrix by extracting its latent factors. Essentially, we map each user and each item into a latent space with dimension r. Therefore, it helps us better understand the relationship between users and items as they become directly comparable. The below figure illustrates this idea.
#
#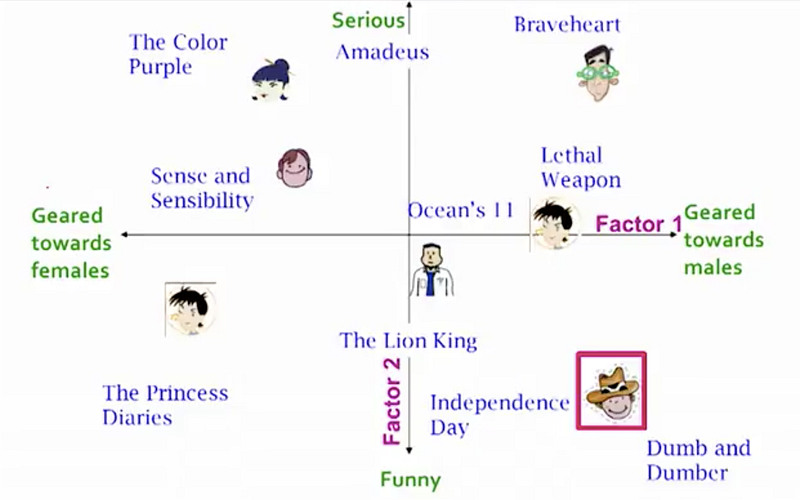 
# + [markdown] _uuid="defca8163cfc24a97bee620d6a3d501aa2ec95ae"
# Now enough said , let's see how to implement this.
# Since the dataset we used before did not have userId(which is necessary for collaborative filtering) let's load another dataset. We'll be using the [**Surprise**](https://surprise.readthedocs.io/en/stable/index.html) library to implement SVD.
# -
from surprise import Reader, Dataset, SVD, evaluate
from surprise.model_selection.search import GridSearchCV
# + _uuid="9a7faf48bf42293d18b29efac95e15010f6c900e"
reader = Reader()
ratings = pd.read_csv('the-movies-dataset/ratings_small.csv')
ratings.head()
# + [markdown] _uuid="f8a5cd5580510c27846a564bfa6d13f1a6dfa6de"
# Note that in this dataset movies are rated on a scale of 5 unlike the earlier one.
# + _uuid="75166cecd9821bab4299605b66ea2a7787a4c3b7"
data = Dataset.load_from_df(ratings[['userId', 'movieId', 'rating']], reader)
data.split(n_folds=5)
type(data)
# + _uuid="17880aee6e7750afed98002593251010dcf5fb20"
svd = SVD()
evaluate(svd, data, measures=['RMSE', 'MAE'])
# GridSearchCV(svd, data, measures=['RMSE', 'MAE'])
# + [markdown] _uuid="81f6d5f460d4cbaaa43b6bb86a0abd9bce1a3134"
# We get a mean Root Mean Sqaure Error of 0.89 approx which is more than good enough for our case. Let us now train on our dataset and arrive at predictions.
# + _uuid="58007ee500ce1735d173c247d188a5cd603b803c"
trainset = data.build_full_trainset()
svd.fit(trainset)
# + [markdown] _uuid="e130ef65d2c6a59823869d5b08c44e96e06f3b94"
# Let us pick user with user Id 1 and check the ratings she/he has given.
# + _uuid="5a526fddacb2f7234e524e71224fdd1aecdd6ec0"
ratings[ratings['userId'] == 1]
# + _uuid="0cb981abe36b30f8f27b4c5dc1b1d0e090431651"
svd.predict(1, 302, 3)
# + [markdown] _uuid="50cf59d88c55c17de150e6a84a190e179bec2d33"
# For movie with ID 302, we get an estimated prediction of **2.618**. One startling feature of this recommender system is that it doesn't care what the movie is (or what it contains). It works purely on the basis of an assigned movie ID and tries to predict ratings based on how the other users have predicted the movie.
# + [markdown] _uuid="aece2ca6c5dcfcc8287562733f74a31fa115605c"
# ## **Conclusion**
# We create recommenders using demographic , content- based and collaborative filtering. While demographic filtering is very elemantary and cannot be used practically, **Hybrid Systems** can take advantage of content-based and collaborative filtering as the two approaches are proved to be almost complimentary.
# This model was very baseline and only provides a fundamental framework to start with.
#
# I would like to mention some excellent refereces that I learned from
# 1. [https://hackernoon.com/introduction-to-recommender-system-part-1-collaborative-filtering-singular-value-decomposition-44c9659c5e75](https://hackernoon.com/introduction-to-recommender-system-part-1-collaborative-filtering-singular-value-decomposition-44c9659c5e75)
# 2. [https://www.kaggle.com/rounakbanik/movie-recommender-systems](https://www.kaggle.com/rounakbanik/movie-recommender-systems)
# 3. [http://trouvus.com/wp-content/uploads/2016/03/A-hybrid-movie-recommender-system-based-on-neural-networks.pdf](http://trouvus.com/wp-content/uploads/2016/03/A-hybrid-movie-recommender-system-based-on-neural-networks.pdf)
#
# If you enjoyed reading the kernel , hit the upvote button !
# Please leave the feedback or suggestions below.
| film-recommendation-engine/getting-started-with-a-movie-recommendation-system.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 5. Estimation – Monte Carlo Augmented Data
#
# <div class="alert alert-block alert-info">
# <b>About:</b>
# This notebook refers to the studies presented in <b>Chapter 5.5</b> of the Ph.D. thesis [3].
# We can not guarantee completeness or correctness of the code.
# If you find bugs or if you have suggestions on how to improve the code, we encourage you to post your ideas as <a href="https://github.com/felixriese/alpaca-processing/issues">GitHub issue</a>.
# </div>
#
# ## Imports
# %load_ext autoreload
# %autoreload 2
# +
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import train_test_split
import sklearn.metrics as me
import numpy as np
import matplotlib.pyplot as plt
from tqdm.notebook import tqdm
import susi
import pandas as pd
import pickle
import utils
# -
def fit_model(model, data):
X_train, X_test, y_train, y_test = data
# fit and predict
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
# evaluate
r2 = me.r2_score(y_test, y_pred)
rmse = np.sqrt(me.mean_squared_error(y_test, y_pred))
mae = me.mean_absolute_error(y_test, y_pred)
return r2, rmse, mae
# ## Regression over number of datapoints
# ### Generate estimations
# +
# CHANGE
area = ["1", "3"]
# postfix = area[0][0]
postfix = "1+3"
test_sizes = np.round(np.arange(0.15, 0.35, 0.05), 2)
mc_args = {"std": 2, "n_new": 10, "area": area, "max_sm": 40}
# load data for checks only
_, y = utils.loadCSVData(area, max_sm=mc_args["max_sm"])
print("Area {0} with {1} datapoints and soil moisture of {2:.2f} ± {3:.2f} %.".format(
area, len(y), np.mean(y), np.std(y)))
# for plots only
meas_error = 4.
meas_error_std = 2.2/2
# plot histo
fig, ax = plt.subplots(1, 1)
n, _, _ = plt.hist(y)
ax.set_ylim(0, max(n)*1.5)
ax.set_xlabel("Soil moisture in %")
ax.set_ylabel("Number of datapoints")
# +
results = {
"model": [],
"r2": [],
"rmse": [],
"mae": [],
"test_size": [],
"random_state": []}
for i, test_size in enumerate(tqdm(test_sizes)):
for random_state in range(40):
data = utils.getMCSoilMoistureData(
test_size=test_size, random_state=random_state, **mc_args)
# --- SOM
model = susi.SOMRegressor(n_rows=10, n_columns=10,
n_iter_unsupervised=2000, n_iter_supervised=1000)
r2, rmse, mae = fit_model(model, data)
results["r2"].append(r2)
results["rmse"].append(rmse)
results["mae"].append(mae)
results["test_size"].append(test_size)
results["random_state"].append(random_state)
results["model"].append("SOM")
# --- RF
model = RandomForestRegressor(n_estimators=100, n_jobs=-1)
r2, rmse, mae = fit_model(model, data)
results["r2"].append(r2)
results["rmse"].append(rmse)
results["mae"].append(mae)
results["test_size"].append(test_size)
results["random_state"].append(random_state)
results["model"].append("RF")
results_df = pd.DataFrame(results)
# + active=""
# results_df.to_csv("results/mc_som_rf.csv")
# print(results_df.shape)
# results_df.head()
# -
# ## Plots
color_list = ["tab:blue", "tab:orange"]
# +
fontsize = 18
fig, ax = plt.subplots(1, 1, figsize=(6, 5))
a_min = -0.5 * 100
a_max = 1. * 100
y_max = 60
bins = np.arange(a_min, a_max, 0.1*100)
for i, m in enumerate(np.unique(results_df.model)):
r2_list = results_df[results_df["model"]==m]["r2"].values
ax.hist(np.clip(r2_list*100, a_min=a_min, a_max=a_max), bins=bins, label=m, alpha=0.5,
color=color_list[i])
print("{0:10} |\tOf {1} are {2} ({3:.1f} %) above 0. Median = {4:.1f} %. Mean = {5:.1f} %.".format(
m, len(np.ravel(r2_list)), sum(np.ravel(r2_list)>0.),
sum(np.ravel(r2_list)>0.)/len(np.ravel(r2_list))*100,
np.median(r2_list)*100, np.mean(r2_list)*100))
ax.axvline(np.median(r2_list)*100, label=m+" median",
color=color_list[i], linestyle="dashed")
ax.set_xlim(a_min, a_max)
ax.set_ylim(0, y_max)
ax.set_xlabel("$R^2$ in %", fontsize=fontsize)
ax.set_ylabel("Number of experiments", fontsize=fontsize, labelpad=10)
for tick in ax.xaxis.get_major_ticks():
tick.label.set_fontsize(fontsize)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(fontsize)
plt.legend(fontsize=fontsize*0.8)
plt.tight_layout()
plt.savefig("plots/mc_area"+postfix+"_hist_r2.pdf", bbox_inches="tight")
# +
fontsize = 18
fig, ax = plt.subplots(1, 1, figsize=(6, 5))
a_min = -0.5
y_max = 120
bins = np.arange(1, 12, 1)
for i, m in enumerate(np.unique(results_df.model)):
mae_list = results_df[results_df["model"]==m]["mae"].values
ax.hist(mae_list, label=m, alpha=0.5, bins=bins, color=color_list[i])
print("{0:10} |\t Median = {1:.1f} %. Mean = {2:.1f} %.".format(
m, np.median(mae_list), np.mean(mae_list)))
ax.axvline(np.median(mae_list), label=m+" median",
color=color_list[i], linestyle="dashed")
ax.set_xlim(bins[0], bins[-1])
ax.set_ylim(0, y_max)
ax.set_xlabel("MAE in % soil moisture", fontsize=fontsize)
ax.set_ylabel("Number of experiments", fontsize=fontsize, labelpad=10)
for tick in ax.xaxis.get_major_ticks():
tick.label.set_fontsize(fontsize)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(fontsize)
plt.legend(fontsize=fontsize*0.8)
plt.tight_layout()
plt.savefig("plots/mc_area"+postfix+"_hist_mae.pdf", bbox_inches="tight")
# +
fontsize = 18
fig, ax = plt.subplots(1, 1, figsize=(6, 5))
y_max = 120
bins = np.arange(1, 13, 1)
for i, m in enumerate(np.unique(results_df.model)):
rmse_list = results_df[results_df["model"]==m]["rmse"].values
ax.hist(rmse_list, label=m, alpha=0.5, bins=bins, color=color_list[i])
print("{0:10} |\t Median = {1:.1f} %. Mean = {2:.1f} %.".format(
m, np.median(rmse_list), np.mean(rmse_list)))
ax.axvline(np.median(rmse_list), label=m+" median",
color=color_list[i], linestyle="dashed")
ax.set_xlim(bins[0], bins[-1])
ax.set_ylim(0, y_max)
ax.set_xlabel("RMSE in % soil moisture", fontsize=fontsize)
ax.set_ylabel("Number of experiments", fontsize=fontsize, labelpad=10)
for tick in ax.xaxis.get_major_ticks():
tick.label.set_fontsize(fontsize)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(fontsize)
plt.legend(fontsize=fontsize*0.8)
plt.tight_layout()
plt.savefig("plots/mc_area"+postfix+"_hist_rmse.pdf", bbox_inches="tight")
# -
# ## MC data histogram plot
# +
X, y = utils.loadCSVData(areas=area, max_sm=mc_args["max_sm"])
X_new, y_new = utils.generateMCData(X=X, y=y, std=mc_args["std"], n_new=mc_args["n_new"], verbose=0)
normed = False
bins = np.arange(int(min(y_new)), int(max(y_new))+2, 2)
fontsize = 15
fig, ax = plt.subplots(1, 1, figsize=(7, 4))
n, _, _ = ax.hist(y_new, density=normed, alpha=1.0, label="Monte Carlo", bins=bins)
ax.hist(y, density=normed, alpha=1.0, label="Original", bins=bins, histtype="step", linewidth=3)
# ax.set_title("Area "+area[0][0], fontsize=fontsize)
ax.legend(fontsize=fontsize*1.0, frameon=False)
ax.set_ylim(0, max(n)*1.5)
ax.set_xlabel("Soil moisture in %", fontsize=fontsize)
ax.set_ylabel("Number of datapoints", fontsize=fontsize)
for tick in ax.xaxis.get_major_ticks():
tick.label.set_fontsize(fontsize)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(fontsize)
plt.tight_layout()
plt.savefig("plots/mc_area"+postfix+"_hist.pdf", bbox_inches="tight")
# -
# ## Scatter plot
# + active=""
# # results_df.iloc[309] # [(results_df["test_size"]==0.3) & (results_df["model"]=="SOM")]
# results_df["r2"].tail(40)
# +
# CHANGE index for plot
i = 312
model_single = susi.SOMRegressor(n_rows=10, n_columns=10,
n_iter_unsupervised=2000, n_iter_supervised=1000)
X_train, X_test, y_train, y_test = utils.getMCSoilMoistureData(
test_size=results_df.iloc[i]["test_size"],
random_state=results_df.iloc[i]["random_state"], **mc_args)
# fit and predict
model_single.fit(X_train, y_train)
y_pred_single = model_single.predict(X_test)
print("R2 = {0:.1f} %".format(100*me.r2_score(y_test, y_pred_single)))
# plot
fontsize = 15
fig, ax = plt.subplots(1, 1, figsize=(6, 6))
factor = mc_args["n_new"]+1
ax.scatter(y_test, y_pred_single, alpha=0.4, label="Monte Carlo")
ax.scatter(y_test[::factor], y_pred_single[::factor], marker="x", s=100, label="Measurements")
ax.set_xlabel("Soil moisture (ground truth) in %", fontsize=fontsize, labelpad=10)
ax.set_ylabel("Soil moisture (estimated) in %", fontsize=fontsize, labelpad=10)
for tick in ax.xaxis.get_major_ticks():
tick.label.set_fontsize(fontsize)
for tick in ax.yaxis.get_major_ticks():
tick.label.set_fontsize(fontsize)
# min_plot = min(min(y_test), min(y_pred_single))
# max_plot = max(max(y_test), max(y_pred_single))
min_plot = 1
max_plot = 34
ax.set_xlim(min_plot-1, max_plot+1)
ax.set_ylim(min_plot-1, max_plot+1)
ax.plot([min_plot-2, max_plot+2], [min_plot-2, max_plot+2], color="grey", linewidth=1)
ax.legend(fontsize=fontsize)
plt.savefig("plots/mc_area"+postfix+"_scatter.pdf", bbox_inches="tight")
# -
# ## SOM estimation map
def plot_estimation_map_regression(estimation_map, sm_range=None,
title="", fontsize=20):
plt.figure(figsize=(7,5))
if sm_range:
plt.imshow(estimation_map, cmap="viridis_r",
vmin=sm_range[0], vmax=sm_range[1])
else:
plt.imshow(estimation_map, cmap="viridis_r")
plt.xlabel("SOM columns", fontsize=fontsize)
plt.ylabel("SOM rows", fontsize=fontsize)
plt.xticks(fontsize=fontsize)
plt.yticks(fontsize=fontsize)
cbar = plt.colorbar()
cbar.ax.tick_params(labelsize=fontsize)
cbar.ax.set_ylabel('Soil moisture in %', fontsize=fontsize, labelpad=10)
for label in cbar.ax.xaxis.get_ticklabels()[::2]:
label.set_visible(False)
plt.grid(b=False)
plt.tight_layout()
plt.savefig("plots/mc_area"+postfix+"_estimationmap"+title+".pdf",
bbox_inches="tight")
esti_map = model_single.get_estimation_map()
plot_estimation_map_regression(np.squeeze(esti_map), sm_range=(9., 26.))
# ## Plot Estimation Map Area 1
hyp_data_map = utils.predictSoilmoistureMap(
area=area[0], model=model_single, dim_red_mode=None,
# sm_range=(12., 29.),
postfix="mc", verbose=1)
pickle.dump(hyp_data_map, open("estimations/hyp_data_map_mc_area1.p", "wb"))
hyp_data_map = pickle.load(open("estimations/hyp_data_map_mc_area1.p", "rb"))
print("Soil moisture between {0:.1f} and {1:.1f} %".format(
np.min(hyp_data_map[hyp_data_map !=0]), np.max(hyp_data_map)))
utils.plotSoilMoistureMap(hyp_data_map, area=area[0][0], sm_range=(9., 26.), postfix="mc")
| py/5_Estimation_MonteCarloAugmentedData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import ete2
import skbio
import numpy as np
import matplotlib.pyplot as plt
# %pylab inline
# -
# ## Load a tree
# it'd be nice if this expandvars call happened interally
from os.path import expandvars
et = ete2.Tree(expandvars('$HOME/data/gg_13_8_otus/trees/61_otus.tree'))
st = skbio.TreeNode.read(expandvars('$HOME/data/gg_13_8_otus/trees/61_otus.tree'))
# ## repr the tree
et
st
print et
et.describe()
print st
# ## Traversing the tree and time it
# %%timeit
# load the tree on each loop to avoid caching of results
ete2.Tree(expandvars('$HOME/data/gg_13_8_otus/trees/85_otus.tree')).traverse("postorder")
# %%timeit
# load the tree on each loop to avoid caching of results
skbio.TreeNode.read(expandvars('$HOME/data/gg_13_8_otus/trees/85_otus.tree')).postorder()
# ## Implement and compute PD
# Phylogenetic Diversity (PD) is the amount of branch length in a phylogenetic tree that is observed in a given biological community. It is a phylogenetic estimator of community richness. I describe it in some more detail in [IAB](http://nbviewer.ipython.org/github/gregcaporaso/An-Introduction-To-Applied-Bioinformatics/blob/0.1.0/applications/biological-diversity.ipynb#Phylogenetic-Diversity-(PD)).
from random import shuffle
tip_names = [t.name for t in st.tips()]
shuffle(tip_names)
observed_tips = tip_names[:10]
# %timeit et.get_leaves_by_name('801940')[0]
# %timeit st.find('801940')
# %timeit et.get_leaves_by_name('801940')[0].get_ancestors()
# %timeit st.find('801940').ancestors()
def et_pd(tree, observed_tips):
observed_nodes = set()
for tip_name in observed_tips:
observed_nodes |= set(tree.get_leaves_by_name(tip_name)[0].get_ancestors())
pd = sum([o.dist for o in observed_nodes])
ds = [(o.dist, o.name) for o in observed_nodes]
ds.sort()
print len(ds)
return ds, pd
ds, pd = et_pd(et, observed_tips)
print pd
_ = plt.hist([e[0] for e in ds])
def sk_pd(tree, observed_tips):
observed_nodes = set()
for tip_name in observed_tips:
observed_nodes |= set(tree.find(tip_name).ancestors())
pd = sum([o.length for o in observed_nodes if o.length != None])
ds = [(o.length, o.name) for o in observed_nodes if o.length != None]
print len(ds)
return ds, pd
ds, pd = sk_pd(st, observed_tips)
print pd
_ = plt.hist([e[0] for e in ds])
# ## **Question**: where are all of the branch lengths of 1.0 coming from in the ete2 tree??
# let's just parse the floats from the newick tree now and see what we get...
import re
s = open(expandvars("$HOME/data/gg_13_8_otus/trees/61_otus.tree")).read()
print s
lengths = map(float,[e[1:] for e in re.findall(':\d\.\d+', s)])
lengths.sort()
print lengths
print len(lengths), np.min(lengths), np.median(lengths), np.max(lengths)
_ = plt.hist(lengths)
# and compare that with the skbio lengths
lengths = [n.length for n in st.postorder() if n.length is not None]
lengths.sort()
print lengths
print len(lengths), np.min(lengths), np.median(lengths), np.max(lengths)
_ = plt.hist(lengths)
# and the ete2 lengths
lengths = [n.dist for n in et.traverse("postorder")]
lengths.sort()
print lengths
print len(lengths), np.min(lengths), np.median(lengths), np.max(lengths)
_ = plt.hist(lengths)
# This is bad... it looks like ete2 is not recognizing the node names, but silently converting them to ``1.0``. This should raise an exception as it can give very misleading results (as is the case here).
#
# Loading with ``format=1`` gets us closer, but there is one label that is still being converted to ``1.0``. This is still very bad, but there also may be an issue with that node in the tree... Is the name of the node really supposed to be ``0.081``?
et1 = ete2.Tree(expandvars('$HOME/data/gg_13_8_otus/trees/61_otus.tree'), format=1)
ds, pd = et_pd(et1, observed_tips)
print pd
_ = plt.hist([e[0] for e in ds])
| ete2-experiments/experiments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.3 64-bit (''base'': conda)'
# metadata:
# interpreter:
# hash: dca0ade3e726a953b501b15e8e990130d2b7799f14cfd9f4271676035ebe5511
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from model import *
from model import Model_Rescorla_Wagner
# +
# -------------------
# Blocking Design
# -------------------
# Define the model
experiment = Model_Rescorla_Wager(experiment_name="Blocking", lambda_US=1, beta_US=0.5)
# Define the predictors
A = Predictor(name='A', alpha = 0.2)
B = Predictor(name='B',alpha = 0.2)
C = Predictor(name='C',alpha = 0.2)
# Define the experiment groups
experiment_group = Group(name="Experiment Group")
experiment_group.add_phase_for_group(phase_name='Conditioning', predictors=[A], outcome=True, number_of_trial=10)
experiment_group.add_phase_for_group(phase_name='Blocking', predictors=[A], outcome=True, number_of_trial=10)
experiment.add_group(experiment_group)
control_group = Group(name="Control Group")
control_group.add_phase_for_group(phase_name='Conditioning', predictors=[C], outcome=True, number_of_trial=10)
control_group.add_phase_for_group(phase_name='Blocking', predictors=[A,B], outcome=True, number_of_trial=10)
experiment.add_group(control_group)
# Run the model
experiment.model_run()
experiment.display_results(save_to_file=True)
# -
experiment.result.head()
# +
# -------------------
# Extinction Design
# -------------------
# Define the model
exitinction = Model_Rescorla_Wager(experiment_name="Extinction", lambda_US=1, beta_US=0.5)
# Define the predictors
A = Predictor(name='A', alpha = 0.2)
# Define the experiment groups
exitinction_group = Group(name="Experiment Group")
exitinction_group.add_phase_for_group(phase_name='Conditioning', predictors=[A], outcome=True, number_of_trial=10)
exitinction_group.add_phase_for_group(phase_name='Extinction', predictors=[A], outcome=False, number_of_trial=10)
exitinction.add_group(exitinction_group)
# Run the model
exitinction.model_run()
exitinction.display_results(save_to_file=True)
| model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Activity 3: Dealing with online JSON files
# Extract comments from https://jsonplaceholder.typicode.com/comments and evaluate sentiment scores of each of them using TextBlob <br>
# Collect 15 author names and titles by parsing JSON files available from http://libgen.io/json.php
import json
import urllib3
from textblob import TextBlob
from pprint import pprint
import pandas as pd
http = urllib3.PoolManager()
rr = http.request('GET', 'https://jsonplaceholder.typicode.com/comments')
rr.status
data = json.loads(rr.data.decode('utf-8'))
import pandas as pd
df = pd.DataFrame(data).head(15)
df.head()
df['body_english'] = df['body'].apply(lambda x: str(TextBlob('u'+str(x)).translate(to='en')))
df[['body', 'body_english']].head()
df['sentiment_score'] = df['body_english'].apply(lambda x: str(TextBlob('u'+str(x)).sentiment.polarity))
df[['body_english', 'sentiment_score']]
# # Collecting JSON data online by multiple requests
# Collect 15 author names and titles by parsing JSON files available from http://libgen.io/json.php
http = urllib3.PoolManager()
df = pd.DataFrame({'author':[], 'title':[]})
for i in range(1,16):
rrd = http.request('GET', 'http://libgen.io/json.php?ids='+str(i)+'&fields=Title,Author')
if rrd.status == 200:
df = df.append(pd.DataFrame(json.loads(rrd.data.decode('utf-8'))))
rrd.data
df
| Lesson04/Activity8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import malaya
# ## List available deep learning POS models
malaya.pos.available_deep_model()
# ## Describe supported POS
malaya.describe_pos()
# ## Load CRF Model
crf = malaya.pos.crf()
string = 'KUALA LUMPUR: Sempena sambutan Aidilfitri minggu depan, Perdana Menteri Tun Dr <NAME> dan <NAME> <NAME> <NAME> menitipkan pesanan khas kepada orang ramai yang mahu pulang ke kampung halaman masing-masing. Dalam video pendek terbitan Jabatan Keselamatan Jalan Raya (JKJR) itu, Dr Mahathir menasihati mereka supaya berhenti berehat dan tidur sebentar sekiranya mengantuk ketika memandu.'
crf.predict(string)
# ## Print important features CRF model
crf.print_features(10)
# ## Print important transitions CRF model
crf.print_transitions(10)
# ## Load deep learning models
for i in malaya.pos.available_deep_model():
print('Testing %s model'%(i))
model = malaya.pos.deep_model(i)
print(model.predict(string))
print()
# ## Voting stack model
entity_network = malaya.pos.deep_model('entity-network')
bahdanau = malaya.pos.deep_model('bahdanau')
luong = malaya.pos.deep_model('luong')
malaya.stack.voting_stack([entity_network, bahdanau, crf], string)
| example/part-of-speech/load-pos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Vanilla Python
# language: vanilla_python
# name: vanilla_python
# ---
# ***
# ## Topic 4.2.2 Explain the role of sub-procedures in solving a problem
#
# Sub-procedures, also called functions, or sub-routines, is an important aspect to programming. Whenever you find yourself writing code over and over again, you can package it up into a function. It helps us to organize our code, and can be used again later, so you're not constantly re-writing when you've found a solution.
#
# Let's learn the syntax of sub-procedure by viewing this quirky program:
# Things to notice:
#
# - You start a sub-procedure definition with the `sub` keyword and end it with `end sub`
# - You call a sub-procedure by typing its name with parentheses
# - You can send it parameters, in order
# - You can define sub-procedures in a different order than they are called
#
# There is an additional facet of the syntax that must be learned as well, and that is its **return value**. This is actually what they are used for most often, as it can save a lot of time. In fact, large, sophisticated programs are impossible without functions. What does `RESULT` equal to?
# +
sub Increment(PARAM)
return PARAM + 1
end sub
RESULT = Increment(41)
# -
# ***
# ## Thinking Like a Computer Scientist: Topics 4.1.4 - 4.1.8
#
# - 4.1.4 Identify when decision-making is required in a specified situation
# - 4.1.5 Identify the decisions required for the solution to a specified problem
# - 4.1.6 Identify the condition associated with a given decision in a specified problem
# - 4.1.7 Explain the relationship between the decisions and conditions of a system
# - 4.1.8 Deduce logical rules for real-world situations
#
# This section of the curriculum could aply be called "Thinking like a Computer Scientist."
# + language="html"
# <img src="https://docs.google.com/drawings/d/e/2PACX-1vSAFs6OGJtRgXSflnIJiWIf7ioH0h_yc5m5he-MqOFGFJciV87W1REJo-_qfMCWIItXwhndWEJDjViC/pub?w=960&h=720">
# -
# ***
# ## Topic 4.1.9 Identify the inputs and outputs required in a solution
#
# Solutions to problems can eventually be wrapped into a function, so that by providing any input, the correct output is provided. Let's build a way for us to calculate the quadratic equation.
#
# $ ax^2 + bx + c = 0 $
#
# Let's remember that in order to solve for $x$, we'll need the square root function. So how do we do that in code? What do we have to replace the `"?"` with in order to make a function that always returns the square root of a number `V`?
# +
sub Sqrt(V)
return "?"
end sub
output Sqrt(16)
# -
# How do we take the power of something? And can we take the `1/2` power?
# +
sub Power(X, Y)
return X ** Y // x to the power of y
end sub
output Power(2, 2)
# -
# So if we understand this so far, making a square root function, we can use the `Power` function and send it `0.5` for the `Y` value.
# +
sub Sqrt(Z)
return Power(Z, 0.5)
end sub
sub Power(X, Y)
return X ** Y // x to the power of y
end sub
output "Power:"
output Power(2, 2)
output ""
output "Sqrt:"
output Sqrt(16)
# -
# We also need to understand why the first function returns `4` bu tthe second returns `4.0`, and it has to do with how numbers are represented, which is an interesting rabbit hole. But for our purposes, the main thing is to get used to thinking of writing computer programs as a series of little functions, each of which has different inputs and produces an expected output.
# ***
# ## Workflow & Design: 4.1.10 - 4.1.13
#
# ## Topic 4.1.10 Identify pre-planning in a suggested problem and solution
#
# Once you've got a solution to a problem, you need to zoom out of the code and see what is needed in order to make sure that a suggestion solution will be able to work for you, or for the company. You'll be able to maintain the suggested software that you are going to build. Let's say for exmaple our school wanted t
# ***
# ## 4.1.11 Explain the need for pre-conditions when executing an algorithm
#
# Assumptions are a basic part of computers, as we saw from our calculation of fibonocci numbers above. In that case, we were given some assumptions. There is also the case where
# ***
# ## 4.1.12 Outline the pre- and post-conditions to a specified problem
#
# Preconditions and input, post-conditions and output are not identical. The input are the given data points and pre-condition are expectations on the data going in. The output is the effect on the data that the program produces, and the post-conditions refer to the expections on the output, for example "the output is consistent with xyz."
#
# For this problem, we're going to write a module (a series of functions) that operates on the data, and passes all tests.
#
# **Input:** A csv file with student information: Name, Grade, Gender, Nationality
#
# **Precondition:** The module must be able to parse a csv and run analytical operations without any errors and complete within seconds
#
# **Ouptut:** A breakdown of most common last name, male to female ratios, and nationality by grades
#
# **Postconditions:** The output is free of numerical errors
# ***
# ## 4.1.13 Identify exceptions that need to be considered in a specified problem solution
#
# Exceptions in code happen all the time. Some of them are accidental due to programmer error, but sometimes they are a part of the process. The same is true for a workflow.
#
# Take for example a factory. Most of the time, fantastic products are produced on an assembly line that can be put into stores. However, due to limitations, perhaps 1 out of 10 items cannot go to stores without another more careful inspection. Or perahps you need to make an exception for every 100 items in order to do safety inspections.
#
# Programs are the same way. You can have programs that expect there to be exceptions, and handle it in a different way. You can also have programs that have bugs, which are exceptions you hadn't thought ahead about.
#
# Pseudocode doesn't really have exceptions specified and you won't need to accommodate them , so we'll use straight Python on line 3, just to illustrate:
#
# +
sub Divide(DIVIDEND, DIVISOR)
if DIVISOR = 0 then
raise Exception("Cannot divide by zero, dummy!") // py
end if
return DIVIDEND div DIVISOR
end sub
output Divide(10, 5) // 2.0
output Divide(10, 0) // "Exception: Cannot divide by zero, dummy!"
# -
# Just like in a workflow or design specificiation, exceptions in programming have the same effect. They don't happen very often, but when they do they potentially break things.
#
# The number `0` is just one number and it is not often that you would run into a situation where you could divide by zero, right? Actually, what about calculating averages?
# +
sub Average(COLLECTION)
SUM = 0
COUNT = 0
loop while COLLECTION.hasNext()
VALUE = COLLECTION.getNext()
SUM = SUM + VALUE
COUNT = COUNT + 1
end loop
COLLECTION.resetNext()
return SUM div COUNT
end sub
GRADES = Collection.from_list([5, 6, 4, 6, 6, 5])
output Average(GRADES)
# -
# But what if we our `COLLECTION` is empty? The program throws an error for us.
# +
sub Average(COLLECTION)
SUM = 0
COUNT = 0
loop while COLLECTION.hasNext()
VALUE = COLLECTION.getNext()
SUM = SUM + VALUE
COUNT = COUNT + 1
end loop
COLLECTION.resetNext()
return SUM div COUNT
end sub
GRADES = Collection() // empty
output Average(GRADES)
# -
# It is common and normal practice that functions check that pre-conditions are met, and returns values depending on those exceptions. So, in this case, we need to make sure that we won't divide by zero. There are two ways to do this, which one is better, and why?
# +
// method #1: count the members, check for zero after counting
sub Average(COLLECTION)
SUM = 0
COUNT = 0
loop while COLLECTION.hasNext()
VALUE = COLLECTION.getNext()
SUM = SUM + VALUE
COUNT = COUNT + 1
end loop
COLLECTION.resetNext()
if COUNT = 0 then
return None // py
end if
return SUM div COUNT
end sub
GRADES = Collection()
output Average(GRADES)
# +
// method #2: use the "isEmpty" method
sub Average(COLLECTION)
if COLLECTION.isEmpty() then
return None // py
end if
SUM = 0
COUNT = 0
loop while COLLECTION.hasNext()
VALUE = COLLECTION.getNext()
SUM = SUM + VALUE
COUNT = COUNT + 1
end loop
COLLECTION.resetNext()
return SUM div COUNT
end sub
GRADES = Collection()
output Average(GRADES)
# -
# Both functions will return `None` if the `COLLECTION` is empty, but it does it in two different ways. The first way is the more manual way, where it sets the variable to `0`, and since `.hasNext` returns `False` on the first attempt, it'll still be `0` on line 12 when we check for it.
#
# The second way is better, because it immediately uses a built-in method that return `True` if there are zero elements, and we can exit early. Not only is it better because there is less code involved, and therefore faster, but it is also better organized. The idea that the first thing a function should do is check the parameters for validity is something very commonly done in programming.
#
# So, both function return `None` but one does it better than the other, and hopefully it is clear why.
#
# > NOTE: `None` in Python is a special value that means roughly "nothing" or "null" or "nadda". When a value is nonsensical, or does not apply, we often use `None` to mean exactly this. Pseudocode doesn't really have this concept, as you are not tested in code on this. But the code helps us understand the concept.
#
# > CHALLENGE: Why do we do `.resetNext` on line 10 (first example) and line 14 (second example)?
# +
sub Calculate(START, BY, UNTIL)
NUM = START
loop NUM from START to UNTIL
NUM = NUM + BY
end loop
output NUM + 2
end sub
sub Enter(QUESTION)
output QUESTION
end sub
sub Exit()
output "*rimshot*"
end sub
Enter("What is the answer to all things?")
Calculate(1, 10, 30)
Exit()
| Topic 4 Computational Thinking, Problem-solving and Programming/5) Connecting Computational Thinking and Program Design (4.2.1 - 4.2.9).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # YOLOFace
#
# Uses code from:
# * https://github.com/sthanhng/yoloface
# * https://github.com/fyr91/face_detection/blob/master/detect_yolov3.py
#
# Useful:
# * https://towardsdatascience.com/real-time-face-recognition-with-cpu-983d35cc3ec5
# * https://github.com/fyr91/face_detection
# * https://github.com/sthanhng/yoloface
# * https://github.com/dannyblueliu/YOLO-Face-detection
# * https://www.baseapp.com/deepsight-image-recognition-sdk/deepsight-face-sdk-download/
import cv2
import os
import numpy as np
import glob
import datetime
# ### Parameters
CONF_THRESHOLD = 0.5
NMS_THRESHOLD = 0.4
IMG_WIDTH = 416
IMG_HEIGHT = 416
# ### Helper Functions
# +
# Get the names of the output layers
def get_outputs_names(net):
# Get the names of all the layers in the network
layers_names = net.getLayerNames()
# Get the names of the output layers, i.e. the layers with unconnected
# outputs
return [layers_names[i[0] - 1] for i in net.getUnconnectedOutLayers()]
# Draw the predicted bounding box
def draw_predict(frame, conf, left, top, right, bottom):
# Draw a bounding box.
cv2.rectangle(frame, (left, top), (right, bottom), (0, 0, 255), 2)
text = '{:.2f}'.format(conf)
# Display the label at the top of the bounding box
label_size, base_line = cv2.getTextSize(text, cv2.FONT_HERSHEY_SIMPLEX, 0.5, 1)
top = max(top, label_size[1])
cv2.putText(frame, text, (left, top - 4), cv2.FONT_HERSHEY_SIMPLEX, 0.4,
(255, 255, 255), 1)
# Add the blurring in
roi = frame[top : bottom, left : right]
# Blur the coloured image
blur = cv2.GaussianBlur(roi, (101, 101), 0)
# Insert the blurred section back into image
frame[top : bottom, left : right] = blur
def post_process(frame, outs, conf_threshold, nms_threshold):
frame_height = frame.shape[0]
frame_width = frame.shape[1]
# Scan through all the bounding boxes output from the network and keep only
# the ones with high confidence scores. Assign the box's class label as the
# class with the highest score.
confidences = []
boxes = []
final_boxes = []
for out in outs:
for detection in out:
scores = detection[5:]
class_id = np.argmax(scores)
confidence = scores[class_id]
if confidence > conf_threshold:
center_x = int(detection[0] * frame_width)
center_y = int(detection[1] * frame_height)
width = int(detection[2] * frame_width)
height = int(detection[3] * frame_height)
left = int(center_x - width / 2)
top = int(center_y - height / 2)
confidences.append(float(confidence))
boxes.append([left, top, width, height])
# Perform non maximum suppression to eliminate redundant
# overlapping boxes with lower confidences.
indices = cv2.dnn.NMSBoxes(boxes, confidences, conf_threshold,
nms_threshold)
for i in indices:
i = i[0]
box = boxes[i]
left = box[0]
top = box[1]
width = box[2]
height = box[3]
final_boxes.append(box)
left, top, right, bottom = refined_box(left, top, width, height)
# draw_predict(frame, confidences[i], left, top, left + width,
# top + height)
draw_predict(frame, confidences[i], left, top, right, bottom)
return final_boxes
def refined_box(left, top, width, height):
right = left + width
bottom = top + height
original_vert_height = bottom - top
top = int(top + original_vert_height * 0.15)
bottom = int(bottom - original_vert_height * 0.05)
margin = ((bottom - top) - (right - left)) // 2
left = left - margin if (bottom - top - right + left) % 2 == 0 else left - margin - 1
right = right + margin
return left, top, right, bottom
def blurFacesYOLO(frame, CONF_THRESHOLD):
# Create a 4D blob from a frame.
blob = cv2.dnn.blobFromImage(frame, 1 / 255, (IMG_WIDTH, IMG_HEIGHT),[0, 0, 0], 1, crop=False)
# Sets the input to the network
net.setInput(blob)
# Runs the forward pass to get output of the output layers
outs = net.forward(get_outputs_names(net))
# Remove the bounding boxes with low confidence
faces = post_process(frame, outs, CONF_THRESHOLD, NMS_THRESHOLD)
return frame.astype(np.uint8)
def blursingle(image, CONF_THRESHOLD):
cv2.imwrite(os.path.splitext(image)[0] + "_blurred.jpg",
blurFacesYOLO(cv2.imread(image), CONF_THRESHOLD))
def blurfolder(folder, CONF_THRESHOLD):
files = glob.glob(os.path.join(folder,"*.jpg"))
for file in files:
blursingle(file, CONF_THRESHOLD)
def blurvideo(video, CONF_THRESHOLD):
# Get first frame
cap = cv2.VideoCapture(video)
video_writer = cv2.VideoWriter(os.path.join(os.path.splitext(video)[0] + "_blurred.avi"),
cv2.VideoWriter_fourcc('M', 'J', 'P', 'G'),
cap.get(cv2.CAP_PROP_FPS), (
round(cap.get(cv2.CAP_PROP_FRAME_WIDTH)),
round(cap.get(cv2.CAP_PROP_FRAME_HEIGHT))))
# While there are still frames to process...
while True:
has_frame, frame = cap.read()
# Stop the program if reached end of video
if not has_frame:
break
video_writer.write(blurFacesYOLO(frame,CONF_THRESHOLD))
# -
# ### Load Model
# +
# Load the model config
net = cv2.dnn.readNetFromDarknet(os.path.abspath("../weights/yolov3-face.cfg"),
os.path.abspath("../weights/yolov3-wider_16000.weights")) # Weights from yoloface github page
net.setPreferableBackend(cv2.dnn.DNN_BACKEND_OPENCV)
net.setPreferableTarget(cv2.dnn.DNN_TARGET_CPU)
# -
# ### Run on Single File
path_to_image = "/path_to_image/image.jpg"
blursingle(path_to_image, 0.25)
# ### Run on folder
path_to_folder = "/path_to_folder"
blurfolder(path_to_folder,0.25)
# ### Run on video
path_to_video = "/path_to_video/video.mp4"
blurvideo(path_to_video, 0.25)
| notebooks/YOLOFace Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (cvxpy)
# language: python
# name: cvxpy
# ---
# + [markdown] nbpresent={"id": "196b8a50-3d29-45c3-82b9-f4a09b49491d"} slideshow={"slide_type": "slide"}
# # Введение в численные методы оптимизации (Ю. Е. Нестеров Введение в выпуклую оптимизацию, гл. 1 $\S$ 1.1)
#
# 1. Обзор материала весеннего семестра
# 2. Постановка задачи
# 3. Общая схема решения
# 4. Сравнение методов оптимизации
# 5. Методы одномерной минимизации
#
# + [markdown] nbpresent={"id": "2a573842-172b-4931-b0dd-9c9d3c47a450"} slideshow={"slide_type": "slide"}
# ## Обзор материала весеннего семестра
#
# Также на [странице курса](https://github.com/amkatrutsa/MIPT-Opt/blob/master/Spring2021/README.md).
#
# 1. Методы решения задач **безусловной** оптимизации
# - Одномерная минимизация (**уже сегодня!**)
# - Градиентный спуск и способы его ускорения
# - Метод Ньютона
# - Квазиньютоновские методы
# - Метод сопряжённых градиентов
# - Решение задачи наименьших квадратов
# 2. Методы решения задач **условной** оптимизации
# - Методы проекции градиента и условного градиента
# - Проксимальные методы
# - Методы штрафных и барьерных функций
# - Метод модифицированой функции Лагранжа
# + [markdown] slideshow={"slide_type": "slide"}
# ## Постановка задачи
#
# \begin{equation}
# \begin{split}
# & \min_{x \in S} f_0(x)\\
# \text{s.t. } & f_j(x) = 0, \; j = 1,\ldots,m\\
# & g_k(x) \leq 0, \; k = 1,\ldots,p
# \end{split}
# \end{equation}
# где $S \subseteq \mathbb{R}^n$, $f_j: S \rightarrow \mathbb{R}, \; j = 0,\ldots,m$, $g_k: S \rightarrow \mathbb{R}, \; k=1,\ldots,p$
#
# Все функции как минимум непрерывны.
#
# Важный факт</span>: задачи **нелинейной** оптимизации
#
# в их самой общей форме являются **численно неразрешимыми**!
# + [markdown] slideshow={"slide_type": "slide"}
# ## Аналитические результаты
# - Необходимое условие первого порядка:
#
# если $x^*$ точка локального минимума дифференцируемой функции $f(x)$, тогда
#
# $$
# f'(x^*) = 0
# $$
#
# - Необходимое условие второго порядка
#
# если $x^*$ точка локального минимума дважды дифференцируемой функции $f(x)$, тогда
#
# $$
# f'(x^*) = 0 \quad \text{и} \quad f''(x^*) \succeq 0
# $$
#
# - Достаточное условие:
#
# пусть $f(x)$ дважды дифференцируемая функция, и пусть точка $x^*$ удовлетворяет условиям
#
# $$
# f'(x^*) = 0 \quad f''(x^*) \succ 0,
# $$
#
# тогда $x^*$ является точкой строго локального минимума функции $f(x)$.
#
# **Замечание**: убедитесь, что Вы понимаете, как доказывать эти
#
# результаты!
# + [markdown] slideshow={"slide_type": "slide"}
# ## Особенности численного решения
#
# 1. Точно решить задачу принципиально невозможно из-за погрешности машинной арифметики
# 2. Необходимо задать критерий обнаружения решения
# 3. Необходимо определить, какую информацию о задаче использовать
# + [markdown] slideshow={"slide_type": "slide"}
# ## Общая итеративная схема
#
# Дано: начальное приближение $x$, требуемая точность $\varepsilon$.
#
# ```python
# def GeneralScheme(x, epsilon):
#
# while StopCriterion(x) > epsilon:
#
# OracleResponse = RequestOracle(x)
#
# UpdateInformation(I, x, OracleResponse)
#
# x = NextPoint(I, x)
#
# return x
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# ### Вопросы
# 1. Какие критерии остановки могут быть?
# 2. Что такое оракул и зачем он нужен?
# 3. Что такое информационная модель?
# 4. Как вычисляется новая точка?
# + [markdown] slideshow={"slide_type": "slide"}
# #### Критерии остановки
# 1. Сходимость по аргументу:
# $$
# \| x_k - x^* \|_2 < \varepsilon
# $$
# 2. Сходимость по функции:
# $$
# \| f_k - f^* \|_2 < \varepsilon
# $$
# 3. Выполнение необходимого условия
# $$
# \| f'(x_k) \|_2 < \varepsilon
# $$
#
# Но ведь $x^*$ неизвестна!
#
# Тогда
#
# \begin{align*}
# & \|x_{k+1} - x_k \| = \|x_{k+1} - x_k + x^* - x^* \| \leq \\
# & \|x_{k+1} - x^* \| + \| x_k - x^* \| \leq 2\varepsilon
# \end{align*}
#
# Аналогично для сходимости по функции,
#
# однако иногда можно оценить $f^*$!
#
# **Замечание**: лучше использовать относительные изменения
#
# этих величин!
#
# Например $\dfrac{\|x_{k+1} - x_k \|_2}{\| x_k \|_2}$
#
# + [markdown] slideshow={"slide_type": "slide"}
# #### Что такое оракул?
# **Определение**: оракулом называют некоторое абстрактное
#
# устройство, которое отвечает на последовательные вопросы
#
# метода
#
# Аналогия из ООП:
#
# - оракул - это виртуальный метод базового класса
# - каждая задача - производный класс
# - оракул определяется для каждой задачи отдельно согласно общему определению в базовом классе
#
# **Концепция чёрного ящика**
# 1. Единственной информацией, получаемой в ходе работы итеративного метода, являются ответы оракула
# 2. Ответы оракула являются *локальными*
# + [markdown] slideshow={"slide_type": "slide"}
# #### Информация о задаче
# 1. Каждый ответ оракула даёт **локальную** информацию о поведении функции в точке
# 2. Агрегируя все полученные ответы оракула, обновляем информацию о **глобальном** виде целевой функции:
# - кривизна
# - направление убывания
# - etc
# + [markdown] slideshow={"slide_type": "slide"}
# #### Вычисление следующей точки
#
# $$
# x_{k+1} = x_{k} + \alpha_k h_k
# $$
#
# - **Линейный поиск**: фиксируется направление $h_k$ и производится поиск по этому направлению "оптимального" значения $\alpha_k$
# - **Метод доверительных областей**: фиксируется допустимый размер *области* по некоторой норме $\| \cdot \| \leq \alpha$ и *модель* целевой функции, которая хорошо её аппроксимирует в выбранной области.
#
# Далее производится поиск направления $h_k$, минимизирующего модель целевой функции и не выводящего точку $x_k + h_k$ за пределы доверительной области
# + [markdown] slideshow={"slide_type": "slide"}
# #### Вопросы
# 1. Как выбрать $\alpha_k$?
# 2. Как выбрать $h_k$?
# 3. Как выбрать модель?
# 4. Как выбрать область?
# 5. Как выбрать размер области?
#
# <span style="color:red">
# В курсе рассматривается только линейный поиск!</span>
#
# Однако несколько раз копцепция метода доверительных областей
#
# будет использована.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Как сравнивать методы оптимизации?
# Для заданного класса задач сравнивают следующие величины:
# 1. Сложность
# - аналитическая: число обращений к оракулу для решения задачи с точностью $\varepsilon$
# - арифметическая: общее число всех вычислений, необходимых для решения задачи с точностью $\varepsilon$
# 2. Скорость сходимости
# 3. Эксперименты
# + [markdown] slideshow={"slide_type": "slide"}
# ### Скорости сходимости
#
# _1._ Сублинейная
#
# $$
# \| x_{k+1} - x^* \|_2 \leq C k^{\alpha},
# $$
#
# где $\alpha < 0$ и $ 0 < C < \infty$
#
# _2._ Линейная (геометрическая прогрессия)
#
# $$
# \| x_{k+1} - x^* \|_2 \leq Cq^k,
# $$
#
# где $q \in (0, 1)$ и $ 0 < C < \infty$
#
# _3._ Сверхлинейная
#
# $$
# \| x_{k+1} - x^* \|_2 \leq Cq^{k^p},
# $$
#
# где $q \in (0, 1)$, $ 0 < C < \infty$ и $p > 1$
#
# _4._ Квадратичная
#
# $$
# \| x_{k+1} - x^* \|_2 \leq C\| x_k - x^* \|^2_2, \qquad \text{или} \qquad \| x_{k+1} - x^* \|_2 \leq C q^{2^k}
# $$
#
# где $q \in (0, 1)$ и $ 0 < C < \infty$
# + slideshow={"slide_type": "slide"}
# %matplotlib inline
import matplotlib.pyplot as plt
USE_COLAB = False
if not USE_COLAB:
plt.rc("text", usetex=True)
import numpy as np
C = 10
alpha = -0.5
q = 0.9
num_iter = 50
sublinear = np.array([C * k**alpha for k in range(1, num_iter + 1)])
linear = np.array([C * q**k for k in range(1, num_iter + 1)])
superlinear = np.array([C * q**(k**2) for k in range(1, num_iter + 1)])
quadratic = np.array([C * q**(2**k) for k in range(1, num_iter + 1)])
plt.figure(figsize=(12,8))
plt.semilogy(np.arange(1, num_iter+1), sublinear,
label=r"Sublinear, $\alpha = -0.5$", linewidth=5)
# plt.semilogy(np.arange(1, num_iter+1), superlinear, linewidth=5,
# label=r"Superlinear, $q = 0.5, p=2$")
plt.semilogy(np.arange(1, num_iter+1), linear,
label=r"Linear, $q = 0.5$", linewidth=5)
# plt.semilogy(np.arange(1, num_iter+1), quadratic,
# label=r"Quadratic, $q = 0.5$", linewidth=5)
plt.xlabel("Number of iterations, $k$", fontsize=28)
plt.ylabel("Error rate upper bound", fontsize=28)
plt.legend(loc="best", fontsize=26)
plt.xticks(fontsize = 28)
_ = plt.yticks(fontsize = 28)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Значение теорем сходимости (Б.Т. Поляк Введение в оптимизацию, гл. 1, $\S$ 6)
# 1. Что дают теоремы сходимости
# - класс задач, для которых можно рассчитывать на применимость метода (важно не завышать условия!)
# - выпуклость
# - гладкость
# - качественное поведение метода
# - существенно ли начальное приближение
# - по какому функционалу есть сходимость
# - оценку скорости сходимости
# - теоретическая оценка поведения метода без проведения экспериментов
# - определение факторов, которые влияют на сходимость (обусловленность, размерность, etc)
# - иногда заранее можно выбрать число итераций для достижения заданной точности
#
# 2. Что **НЕ** дают теоремы сходимости
# - сходимость метода **ничего не говорит** о целесообразности его применения
# - оценки сходимости зависят от неизвестных констант - неконструктивный характер
# - учёт ошибок округления и точности решения вспомогательных задач
#
# **Мораль**: нужно проявлять разумную осторожность
#
# и здравый смысл!
# + [markdown] slideshow={"slide_type": "slide"}
# ## Классификация задач
# 1. Безусловная оптимизация
# - целевая функция липшицева
# - градиент целевой функции липшицев
# 2. Условная оптимизация
# - многогранник
# - множество простой структуры
# - общего вида
# + [markdown] slideshow={"slide_type": "slide"}
# ## Классификация методов
#
# ### Какой размер истории нужно хранить для обновления?
#
# 1. Одношаговые методы
#
# $$
# x_{k+1} = \Phi(x_k)
# $$
#
# 2. Многошаговые методы
#
# $$
# x_{k+1} = \Phi(x_k, x_{k-1}, ...)
# $$
#
# ### Какой порядок поизводных нужно вычислить?
# 1. Методы нулевого порядка: оракул возвращает только значение функции $f(x)$
#
# 2. Методы первого порядка: оракул возвращает значение функции $f(x)$ и её градиент $f'(x)$
#
# 3. Методы второго порядка: оракул возвращает значение функции $f(x)$, её градиент $f'(x)$ и гессиан $f''(x)$.
#
# **Q**: существуют ли методы более высокого порядка?
#
# + [markdown] slideshow={"slide_type": "slide"}
# **А**: [Implementable tensor methods in unconstrained convex optimization](https://link.springer.com/content/pdf/10.1007/s10107-019-01449-1.pdf) by <NAME>, 2019
# + [markdown] slideshow={"slide_type": "slide"}
# ## Одномерная минимизация
# **Определение**. Функция $f(x)$ называется унимодальной на $[a, b]$, если существует такая точка $x^* \in [a, b]$, что
# - $f(x_1) > f(x_2)$ для любых $a \leq x_1 < x_2 < x^*$,
#
# и
# - $f(x_1) < f(x_2)$ для любых $x^* < x_1 < x_2 \leq b$.
#
# **Вопрос**: какая геометрия унимодальных функций?
# + [markdown] slideshow={"slide_type": "slide"}
# ### Метод дихотомии
#
# Идея из информатики первого семестра:
#
# делим отрезок $[a,b]$ на две равные части
#
# пока не найдём минимум унимодальной функции.
#
# - $N$ - число вычислений функции $f$
# - $K = \frac{N - 1}{2}$ - число итераций
#
# Тогда
#
# $$
# |x_{K+1} - x^*| \leq \frac{b_{K+1} - a_{K+1}}{2} = \left( \frac{1}{2} \right)^{\frac{N-1}{2}} (b - a) \approx 0.5^{K} (b - a)
# $$
# + slideshow={"slide_type": "fragment"}
def binary_search(f, a, b, epsilon, callback=None):
c = (a + b) / 2.0
while abs(b - a) > epsilon:
# Check left subsegment
y = (a + c) / 2.0
if f(y) <= f(c):
b = c
c = y
else:
# Check right subsegment
z = (b + c) / 2.0
if f(c) <= f(z):
a = y
b = z
else:
a = c
c = z
if callback is not None:
callback(a, b)
return c
# -
def my_callback(a, b, left_bound, right_bound, approximation):
left_bound.append(a)
right_bound.append(b)
approximation.append((a + b) / 2.0)
# + slideshow={"slide_type": "slide"}
import numpy as np
left_boud_bs = []
right_bound_bs = []
approximation_bs = []
callback_bs = lambda a, b: my_callback(a, b,
left_boud_bs, right_bound_bs, approximation_bs)
# Target unimodal function on given segment
f = lambda x: (x - 2) * x * (x + 2)**2 # np.power(x+2, 2)
# f = lambda x: -np.sin(x)
x_true = -2
# x_true = np.pi / 2.0
a = -3
b = -1.5
epsilon = 1e-8
x_opt = binary_search(f, a, b, epsilon, callback_bs)
print(np.abs(x_opt - x_true))
plt.figure(figsize=(10,6))
plt.plot(np.linspace(a,b), f(np.linspace(a,b)))
plt.title("Objective function", fontsize=28)
plt.xticks(fontsize = 28)
_ = plt.yticks(fontsize = 28)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Метод золотого сечения
# Идея:
#
# делить отрезок $[a,b]$ не на две равные насти,
#
# а в пропорции "золотого сечения".
#
# Оценим скорость сходимости аналогично методу дихотомии:
#
# $$
# |x_{K+1} - x^*| \leq b_{K+1} - a_{K+1} = \left( \frac{1}{\tau} \right)^{N-1} (b - a) \approx 0.618^K(b-a),
# $$
# где $\tau = \frac{\sqrt{5} + 1}{2}$.
#
# - Константа геометрической прогрессии **больше**, чем у метода дихотомии
# - Количество вызовов функции **меньше**, чем у метода дихотомии
# + slideshow={"slide_type": "fragment"}
def golden_search(f, a, b, tol=1e-5, callback=None):
tau = (np.sqrt(5) + 1) / 2.0
y = a + (b - a) / tau**2
z = a + (b - a) / tau
while b - a > tol:
if f(y) <= f(z):
b = z
z = y
y = a + (b - a) / tau**2
else:
a = y
y = z
z = a + (b - a) / tau
if callback is not None:
callback(a, b)
return (a + b) / 2.0
# + slideshow={"slide_type": "slide"}
left_boud_gs = []
right_bound_gs = []
approximation_gs = []
cb_gs = lambda a, b: my_callback(a, b, left_boud_gs, right_bound_gs, approximation_gs)
x_gs = golden_search(f, a, b, epsilon, cb_gs)
print(f(x_opt))
print(f(x_gs))
print(np.abs(x_opt - x_true))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Сравнение методов одномерной минимизации
# + slideshow={"slide_type": "fragment"}
plt.figure(figsize=(10,6))
plt.semilogy(np.arange(1, len(approximation_bs) + 1), np.abs(x_true - np.array(approximation_bs, dtype=np.float64)), label="Binary search")
plt.semilogy(np.arange(1, len(approximation_gs) + 1), np.abs(x_true - np.array(approximation_gs, dtype=np.float64)), label="Golden search")
plt.xlabel(r"Number of iterations, $k$", fontsize=26)
plt.ylabel("Error rate upper bound", fontsize=26)
plt.legend(loc="best", fontsize=26)
plt.xticks(fontsize = 26)
_ = plt.yticks(fontsize = 26)
# + slideshow={"slide_type": "fragment"}
# %timeit binary_search(f, a, b, epsilon)
# %timeit golden_search(f, a, b, epsilon)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Пример иного поведения методов
#
# $$
# f(x) = \sin(\sin(\sin(\sqrt{x}))), \; x \in [2, 60]
# $$
# + slideshow={"slide_type": "fragment"}
f = lambda x: np.sin(np.sin(np.sin(np.sqrt(x))))
x_true = (3 * np.pi / 2)**2
a = 2
b = 60
epsilon = 1e-8
plt.plot(np.linspace(a,b), f(np.linspace(a,b)))
plt.xticks(fontsize = 28)
_ = plt.yticks(fontsize = 28)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Сравнение скорости сходимости и времени работы методов
# + [markdown] slideshow={"slide_type": "slide"}
# ### Метод дихотомии
# + slideshow={"slide_type": "fragment"}
left_boud_bs = []
right_bound_bs = []
approximation_bs = []
callback_bs = lambda a, b: my_callback(a, b,
left_boud_bs, right_bound_bs, approximation_bs)
x_opt = binary_search(f, a, b, epsilon, callback_bs)
print(np.abs(x_opt - x_true))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Метод золотого сечения
# + slideshow={"slide_type": "fragment"}
left_boud_gs = []
right_bound_gs = []
approximation_gs = []
cb_gs = lambda a, b: my_callback(a, b, left_boud_gs, right_bound_gs, approximation_gs)
x_gs = golden_search(f, a, b, epsilon, cb_gs)
print(np.abs(x_opt - x_true))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Сходимость
# + slideshow={"slide_type": "fragment"}
plt.figure(figsize=(8,6))
plt.semilogy(np.abs(x_true - np.array(approximation_bs, dtype=np.float64)), label="Binary")
plt.semilogy(np.abs(x_true - np.array(approximation_gs, dtype=np.float64)), label="Golden")
plt.legend(fontsize=28)
plt.xticks(fontsize=28)
_ = plt.yticks(fontsize=28)
plt.xlabel(r"Number of iterations, $k$", fontsize=26)
plt.ylabel("Error rate upper bound", fontsize=26)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Время работы
# + slideshow={"slide_type": "fragment"}
# %timeit binary_search(f, a, b, epsilon)
# %timeit golden_search(f, a, b, epsilon)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Резюме
# 1. Введение в численные методы оптимизации
# 2. Общая схема работы метода
# 3. Способы сравнения методов оптимизации
# 4. Зоопарк задач и методов
# 5. Одномерная минимизация
# + [markdown] slideshow={"slide_type": "slide"}
# ## Методы спуска. Градиентный спуск и его ускоренные модификации
# + [markdown] slideshow={"slide_type": "fragment"}
# ## Что такое методы спуска?
#
# Последовательность $x_k$ генерируется по правилу
#
# $$
# x_{k+1} = x_k + \alpha_k h_k
# $$
#
# так что
#
# $$
# f(x_{k+1}) < f(x_k)
# $$
#
# Направление $h_k$ называется *направлением убывания*.
#
# **Замечание**: существуют методы, которые не требуют монотонного убывания функции от итерации к итерации.
# + [markdown] slideshow={"slide_type": "fragment"}
# ```python
# def DescentMethod(f, x0, epsilon, **kwargs):
#
# x = x0
#
# while StopCriterion(x, f, **kwargs) > epsilon:
#
# h = ComputeDescentDirection(x, f, **kwargs)
#
# alpha = SelectStepSize(x, h, f, **kwargs)
#
# x = x + alpha * h
#
# return x
#
#
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# ## Способ 1: направление убывания
# Рассмотрим линейную аппроксимацию дифференцируемой функции $f$ вдоль некоторого направления убывания $h, \|h\|_2 = 1$:
#
# $$
# f(x + \alpha h) = f(x) + \alpha \langle f'(x), h \rangle + o(\alpha)
# $$
#
# Из условия убывания
#
# $$
# f(x) + \alpha \langle f'(x), h \rangle + o(\alpha) < f(x)
# $$
#
# и переходя к пределу при $\alpha \rightarrow 0$:
#
# $$
# \langle f'(x), h \rangle \leq 0
# $$
#
# Также из неравенства Коши-Буняковского-Шварца
#
# $$
# \langle f'(x), h \rangle \geq -\| f'(x) \|_2 \| h \|_2 = -\| f'(x) \|_2
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# Таким образом, направление антиградиента
#
# $$
# h = -\dfrac{f'(x)}{\|f'(x)\|_2}
# $$
#
# даёт направление **наискорейшего локального** убывания функции$~f$.
#
# В итоге метод имеет вид
#
# $$
# x_{k+1} = x_k - \alpha f'(x_k)
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# ## Способ 2: схема Эйлера решения ОДУ
#
# Рассмотрим обыкновенное диференциальное уравнение вида:
#
# $$
# \frac{dx}{dt} = -f'(x(t))
# $$
#
# и дискретизуем его на равномерной сетке с шагом $\alpha$:
#
# $$
# \frac{x_{k+1} - x_k}{\alpha} = -f'(x_k),
# $$
#
# где $x_k \equiv x(t_k)$ и $\alpha = t_{k+1} - t_k$ - шаг сетки.
#
# Отсюда получаем выражение для $x_{k+1}$
#
# $$
# x_{k+1} = x_k - \alpha f'(x_k),
# $$
#
# которое в точности совпадает с выражением для градиентного спуска.
#
# Такая схема называется явной или прямой схемой Эйлера.
#
# **Q:** какая схема называется неявной или обратной?
# + [markdown] slideshow={"slide_type": "slide"}
# ## Способ 3: минимизация квадратичной оценки сверху
# #### (<NAME> "Метод универсального градиентного спуска" https://arxiv.org/abs/1711.00394)
#
# Глобальная оценка сверху на функцию $f$ в точке $x_k$:
#
# $$
# f(y) \leq f(x_k) + \langle f'(x_k), y - x_k \rangle + \frac{L}{2} \|y - x_k \|_2^2 = g(y),
# $$
#
# где $\lambda_{\max}(f''(x)) \leq L$ для всех допустимых $x$.
#
# Справа — квадратичная форма, точка минимума которой имеет аналитическое выражение:
#
# \begin{align*}
# & g'(y^*) = 0 \\
# & f'(x_k) + L (y^* - x_k) = 0 \\
# & y^* = x_k - \frac{1}{L}f'(x_k) = x_{k+1}
# \end{align*}
#
# Этот способ позволяет оценить значение шага как $\frac{1}{L}$. Однако часто константа $L$ неизвестна.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Итого: метод градиентного спуска — дёшево и сердито
#
# ```python
# def GradientDescentMethod(f, x0, epsilon, **kwargs):
#
# x = x0
#
# while StopCriterion(x, f, **kwargs) > epsilon:
#
# h = ComputeGradient(x, f, **kwargs)
#
# alpha = SelectStepSize(x, h, f, **kwargs)
#
# x = x - alpha * h
#
# return x
#
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# ## Как выбрать шаг $\alpha_k$? (<NAME>, <NAME> Numerical Optimization, $\S$ 3.1.)
#
# Список подходов:
# - Постоянный шаг
#
# $$
# \alpha_k = \overline{\alpha}
# $$
#
# - Априорно заданная последовательность, например
#
# $$
# \alpha_k = \dfrac{\overline{\alpha}}{\sqrt{k+1}}
# $$
#
# - Наискорейший спуск
#
# $$
# \alpha_k = \arg\min_{\alpha \geq 0} f(x_k - \alpha f'(x_k))
# $$
#
# - Требование **достаточного** убывания, требование **существенного** убывания и условие кривизны: для некоторых $\beta_1, \beta_2$, таких что $0 < \beta_1 < \beta_2 < 1$ найти $x_{k+1}$ такую что
#
# - Достаточное убывание: $f(x_{k+1}) \leq f(x_k) + \beta_1 \alpha_k \langle f'(x_k), h_k \rangle$ или
# $ f(x_k) - f(x_{k+1}) \geq \beta_1 \alpha_k \langle f'(x_k), h_k \rangle
# $
# - Существенное убывание: $f(x_{k+1}) \geq f(x_k) + \beta_2 \alpha_k \langle f'(x_k), h_k \rangle$ или
# $
# f(x_k) - f(x_{k+1}) \leq \beta_2 \alpha_k \langle f'(x_k), h_k \rangle
# $
# - Условие кривизны: $\langle f'(x_{k+1}), h_k \rangle \geq \beta_2 \langle f'(x_k), h_k \rangle$
#
# Обычно коэффициенты выбирают так: $\beta_1 \in (0, 0.3)$, а $\beta_2 \in (0.9, 1)$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Анализ и мотивация подходов к выбору шага $\alpha_k$
# - Постоянный шаг: самое простое и неэффективное решение
# - Априорно заданная последовательность: немногим лучше постоянного шага
# - Наискорейший спуск: самое лучшее решение, но применимо только если вспомогательная задача решается аналитически или ооооооочень быстро. <br></br>
# То есть почти всегда неприменимо :)
# - Требование достаточного убывания, требование существенного убывания и условие кривизны:
# - требование достаточного убывания гарантирует, что функция в точке $x_{k+1}$ не превосходит линейной аппроксимации с коэффициентом наклона $\beta_1$
# - требование существенного убывания гарантирует, что функция в точке $x_{k+1}$ убывает не меньше, чем линейная аппроксимация c коэффициентом наклона $\beta_2$
# - условие кривизны гарантирует, что угол наклона касательной в точке $x_{k+1}$ не меньше, чем угол наклона касательной в точке $x_k$, <br></br>
# умноженный на $\beta_2$
#
# Требование существенного убывания и условие кривизны обеспечивают убывание функции по выбранному направлению $h_k$. Обычно выбирают одно из них.
# + [markdown] slideshow={"slide_type": "fragment"}
# #### Альтернативные названия
# - Требование достаточного убывания $\equiv$ правило Армихо
# - Требование достаточного убывания + условие кривизны $\equiv$ правило Вольфа
# - Требование достаточного убывания + требование существенного убывания $\equiv$ правило Гольдштейна
# + [markdown] slideshow={"slide_type": "slide"}
# ## Зачем нужно условие существенного убывания?
# + slideshow={"slide_type": "slide"}
# %matplotlib notebook
import matplotlib.pyplot as plt
plt.rc("text", usetex=True)
import ipywidgets as ipywidg
import numpy as np
import liboptpy.unconstr_solvers as methods
import liboptpy.step_size as ss
from tqdm import tqdm
# + slideshow={"slide_type": "slide"}
f = lambda x: np.power(x, 2)
gradf = lambda x: 2 * x
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1)
def update(x0, step):
gd = methods.fo.GradientDescent(f, gradf, ss.ConstantStepSize(step))
_ = gd.solve(np.array([x0]), max_iter=10)
x_hist = gd.get_convergence()
x = np.linspace(-5, 5)
ax.clear()
ax.plot(x, f(x), color="r", label="$f(x) = x^2$")
y_hist = np.array([f(x) for x in x_hist])
x_hist = np.array(x_hist)
plt.quiver(x_hist[:-1], y_hist[:-1], x_hist[1:]-x_hist[:-1], y_hist[1:]-y_hist[:-1],
scale_units='xy', angles='xy', scale=1, width=0.005, color="green", label="Descent path")
ax.legend()
fig.canvas.draw()
step_slider = ipywidg.FloatSlider(value=0.8, min=0, max=1.2, step=0.1, description="Step")
x0_slider = ipywidg.FloatSlider(value=1.5, min=-4, max=4, step=0.1, description="Initial point")
_ = ipywidg.interact(update, x0=x0_slider, step=step_slider)
# + slideshow={"slide_type": "slide"}
def plot_alpha(f, grad, x, h, alphas, beta1, beta2):
df = np.zeros_like(alphas)
for i, alpha in enumerate(alphas):
df[i] = f(x + alpha * h)
upper_bound = f(x) + beta1 * alphas * grad(x) * h
lower_bound = f(x) + beta2 * alphas * grad(x) * h
plt.plot(alphas, df, label=r"$f(x + \alpha h)$")
plt.plot(alphas, upper_bound, label="Upper bound")
plt.plot(alphas, lower_bound, label="Lower bound")
plt.xlabel(r"$\alpha$", fontsize=18)
plt.legend(loc="best", fontsize=18)
# + slideshow={"slide_type": "slide"}
f = lambda x: x**2
grad = lambda x: 2 * x
beta1 = 0.1
beta2 = 0.9
x0 = 0.5
plot_alpha(f, grad, x0, -grad(x0), np.linspace(1e-3, 1.01, 10), beta1, beta2)
# + [markdown] slideshow={"slide_type": "slide"}
# ## $f(x) = x\log x$
# + slideshow={"slide_type": "slide"}
x_range = np.linspace(1e-10, 4)
plt.plot(x_range, x_range * np.log(x_range))
# + slideshow={"slide_type": "slide"}
x0 = 1
f = lambda x: x * np.log(x)
grad = lambda x: np.log(x) + 1
beta1 = 0.3
beta2 = 0.7
plot_alpha(f, grad, x0, -grad(x0), np.linspace(1e-3, 0.9, 10), beta1, beta2)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Backtracking
#
# ```python
# def SelectStepSize(x, f, h, rho, alpha0, beta1, beta2):
#
# # 0 < rho < 1
#
# # alpha0 - initial guess of step size
#
# # beta1 and beta2 - constants from conditions
#
# alpha = alpha0
#
# # Check violating sufficient decrease and curvature conditions
#
# while (f(x - alpha * h) >= f(x) + beta1 * alpha grad_f(x_k).dot(h)) and
#
# (grad_f(x - alpha * h).dot(h) <= beta2 * grad_f(x_k).dot(h)):
#
# alpha *= rho
#
# return alpha
#
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# ## Теоремы сходимости (<NAME>ведение в оптимизацию, гл. 1, $\S$ 4; гл. 3, $\S$ 1; Ю.Е. Нестеров Введение в выпуклую оптимизацию, $\S$ 2.2)
# От общего к частному:
# + [markdown] slideshow={"slide_type": "fragment"}
# **Теорема 1.**
# Пусть
#
# - $f(x)$ дифференцируема на $\mathbb{R}^n$,
# - градиент $f(x)$ удовлетворяет условию Липшица с константой $L$
# - $f(x)$ ограничена снизу
# - $\alpha = const$ и $0 < \alpha < \frac{2}{L}$
#
# Тогда для градиентного метода выполнено:
#
# $$
# \lim\limits_{k \to \infty} f'(x_k) = 0,
# $$
#
# а функция монотонно убывает $f(x_{k+1}) < f(x_k)$.
# + [markdown] slideshow={"slide_type": "fragment"}
# **Теорема 2.** Пусть
# - $f(x)$ дифференцируема на $\mathbb{R}^n$
# - $f(x)$ выпукла
# - $f'(x)$ удовлетворяет условию Липшица с константой $L$
# - $\alpha = \dfrac{1}{L}$
#
# Тогда
#
# $$
# f(x_k) - f^* \leq \dfrac{2L \| x_0 - x^*\|^2_2}{k+4}
# $$
#
# + [markdown] slideshow={"slide_type": "fragment"}
# **Теорема 3.**
# Пусть
# - $f(x)$ дважды дифференцируема и $\mu\mathbf{I} \preceq f''(x) \preceq L\mathbf{I}$ для всех $x$
# - $\alpha = const$ и $0 < \alpha < \frac{2}{L}$
#
# Тогда
#
# $$
# \| x_k - x^*\|_2 \leq \|x_0 - x^*\|_2 q^k, \qquad q = \max(|1 - \alpha l|, |1 - \alpha L|) < 1
# $$
#
# и минимальное $q^* = \dfrac{L - \mu}{L + \mu}$ при $\alpha^* = \dfrac{2}{L + \mu}$
# + [markdown] slideshow={"slide_type": "slide"}
# ### От чего зависит $q^*$ и как это использовать?
# Из Теоремы 3 имеем
#
# $$
# q^* = \dfrac{L - \mu}{L + \mu} = \dfrac{L/\mu - 1}{L/\mu + 1} = \dfrac{M - 1}{M + 1},
# $$
#
# где $M$ - оценка числа обусловленности $f''(x)$.
#
# **Вопрос**: что такое число обусловленности матрицы?
#
# - При $M \gg 1$, $q^* \to 1 \Rightarrow$ оооочень **медленная** сходимости градиентного метода. Например при $M = 100$: $q^* \approx 0.98 $
# - При $M \simeq 1$, $q^* \to 0 \Rightarrow$ **ускорение** сходимости градиентного метода. Например при $M = 4$: $q^* = 0.6 $
#
# **Вопрос**: какая геометрия у этого требования?
#
# **Мораль**: необходимо сделать оценку $M$ как можно ближе к 1!
#
# О том, как это сделать, Вам будет предложено подумать в домашнем задании :)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Вычислительный аспект и эксперименты
# 1. Для каждого шага метода нужно хранить только текущую точку и вектор градиента: $O(n)$ памяти
# 2. Поиск $\alpha_k$:
#
# - дан априори
# - ищется из аналитического решения задачи наискорейшего спуска
# - заканчивается за конечное число шагов
# 3. Для каждого шага метода нужно вычислять линейную комбинацию векторов: $O(n)$ вычислений + высокопроизводительные реализации
# + [markdown] slideshow={"slide_type": "slide"}
# ### Pеализация градиентного спуска
# + slideshow={"slide_type": "fragment"}
def GradientDescent(f, gradf, x0, epsilon, num_iter, line_search,
disp=False, callback=None, **kwargs):
x = x0.copy()
iteration = 0
opt_arg = {"f": f, "grad_f": gradf}
for key in kwargs:
opt_arg[key] = kwargs[key]
while True:
gradient = gradf(x)
alpha = line_search(x, -gradient, **opt_arg)
x = x - alpha * gradient
if callback is not None:
callback(x)
iteration += 1
if disp:
print("Current function val =", f(x))
print("Current gradient norm = ", np.linalg.norm(gradf(x)))
if np.linalg.norm(gradf(x)) < epsilon:
break
if iteration >= num_iter:
break
res = {"x": x, "num_iter": iteration, "tol": np.linalg.norm(gradf(x))}
return res
# + [markdown] slideshow={"slide_type": "slide"}
# ### Выбор шага
#
# Реализации различных способов выбора шага приведены [тут](https://github.com/amkatrutsa/liboptpy/blob/master/step_size.py)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Зависимость от обусловленности матрицы $f''(x)$
# Рассмотрим задачу
# $$
# \min f(x),
# $$
# где
# $$ f(x) = x^{\top}Ax, \; A = \begin{bmatrix} 1 & 0\\ 0 & \gamma \end{bmatrix} $$
#
# $$
# f'(x) = 2Ax
# $$
# + slideshow={"slide_type": "fragment"}
def my_f(x, A):
return 0.5 * x.dot(A.dot(x))
def my_gradf(x, A):
return A.dot(x)
# + slideshow={"slide_type": "slide"}
plt.rc("text", usetex=True)
gammas = [0.1, 0.5, 1, 2, 3, 4, 5, 10, 20, 50, 100, 1000, 5000, 10000]
# gammas = [1]
num_iter_converg = []
for g in gammas:
A = np.array([[1, 0],
[0, g]], dtype=np.float64)
f = lambda x: my_f(x, A)
gradf = lambda x: my_gradf(x, A)
# x0 = np.random.rand(A.shape[0])
# x0 = np.sort(x0)
# x0 = x0[::-1]
x0 = np.array([g, 1], dtype=np.float64)
# print x0[1] / x0[0]
gd = methods.fo.GradientDescent(f, gradf, ss.ExactLineSearch4Quad(A))
x = gd.solve(x0, tol=1e-7, max_iter=100)
num_iter_converg.append(len(gd.get_convergence()))
plt.figure(figsize=(8, 6))
plt.loglog(gammas, num_iter_converg)
plt.xticks(fontsize = 20)
plt.yticks(fontsize = 20)
plt.xlabel(r"$\gamma$", fontsize=20)
plt.ylabel(r"Number of iterations with $\varepsilon = 10^{-7}$", fontsize=20)
# + [markdown] slideshow={"slide_type": "slide"}
# - При неудачном начальном приближении сходимость для плохо обусловенной задачи очень медленная
# - При случайном начальном приближении сходимость может быть гораздо быстрее теоретических оценок
# + [markdown] slideshow={"slide_type": "slide"}
# ## Эксперимент на многомерной задаче
# Пусть $A \in \mathbb{R}^{m \times n}$. Рассмотрим систему линейных неравенств: $Ax \leq 1$ при условии $|x_i| \leq 1$ для всех $i$.
#
# **Определение.** Аналитическим центром системы неравенств $Ax \leq 1$ при условии $|x_i| \leq 1$ является решение задачи
# $$
# f(x) = - \sum_{i=1}^m \log(1 - a_i^{\top}x) - \sum_{i = 1}^n \log (1 - x^2_i) \to \min_x
# $$
# $$
# f'(x) - ?
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Точное решение с помощью CVXPy
# + slideshow={"slide_type": "fragment"}
import numpy as np
n = 1000
m = 2000
A = np.random.rand(n, m)
x = cvx.Variable(n)
obj = cvx.Minimize(cvx.sum(-cvx.log(1 - A.T * x)) -
cvx.sum(cvx.log(1 - cvx.square(x))))
prob = cvx.Problem(obj)
prob.solve(solver="SCS", verbose=True)
x = x.value
print("Optimal value =", prob.value)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Решение с помощью градиентного спуска
# + slideshow={"slide_type": "fragment"}
import cvxpy as cvx
print(cvx.installed_solvers())
# # !pip install jax
# # !pip install jaxlib
import jax.numpy as jnp
import jax
# from jax.config import config
# config.update("jax_enable_x64", True)
A = jnp.array(A)
print(A.dtype)
x0 = jnp.zeros(n)
f = lambda x: -jnp.sum(jnp.log(1 - A.T@x)) - jnp.sum(jnp.log(1 - x*x))
grad_f = lambda x: jnp.sum(A @ (jnp.diagflat(1 / (1 - A.T @ x))), \
axis=1) + 2 * x / (1 - jnp.power(x, 2))
grad_f_jax = jax.grad(f)
print(jnp.linalg.norm(grad_f(x0) - grad_f_jax(x0)))
# + [markdown] slideshow={"slide_type": "fragment"}
# Подробнее про jax, его возможности и особенности можно посмотреть например [тут](https://github.com/amkatrutsa/MIPT-Opt/blob/master/Fall2020/03-MatrixCalculus/jax_autodiff_tutorial.ipynb)
# + slideshow={"slide_type": "slide"}
gd = methods.fo.GradientDescent(f, grad_f_jax, ss.Backtracking("Armijo", rho=0.5, beta=0.1, init_alpha=1.))
x = gd.solve(x0, tol=1e-5, max_iter=100, disp=True)
x_conv = gd.get_convergence()
grad_conv = [jnp.linalg.norm(grad_f_jax(x)) for x in x_conv]
plt.figure(figsize=(8,6))
plt.semilogy(grad_conv, label=r"$\| f'(x_k) \|_2$")
plt.semilogy([np.linalg.norm(x - np.array(x_k)) for x_k in x_conv], label=r"$\|x_k - x^*\|_2$")
plt.semilogy([np.linalg.norm(prob.value - f(np.array(x_k))) for x_k in x_conv], label=r"$\|f(x_k) - f^*\|_2$")
plt.semilogy([np.linalg.norm(np.array(x_conv[i]) - np.array(x_conv[i+1])) for i in range(len(x_conv) - 1)], label=r"$\|x_k - x_{k+1}\|_2$")
plt.semilogy([np.linalg.norm(f(np.array(x_conv[i])) - f(np.array(x_conv[i+1]))) for i in range(len(x_conv) - 1)], label=r"$\|f(x_k) - f(x_{k+1})\|_2$")
plt.xlabel(r"Number of iteration, $k$", fontsize=20)
plt.ylabel(r"Convergence rate", fontsize=20)
plt.xticks(fontsize = 20)
plt.yticks(fontsize = 20)
plt.legend(loc="best", fontsize=20)
plt.tight_layout()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Pro & Contra
#
# Pro
# - легко реализовать
# - сходимость как минимум к стационарной точке
# - параметры при выборе шага влияют на сходимость не столь сильно
# - имеет многочисленные вариации
#
# Contra
# - линейная сходимость для сильно выпуклых функций
# - очень сильно зависит от числа обусловленности $f''(x)$, выбор начального приближения может помочь
# - не является оптимальным для выпуклых функций с липшицевым градиентом и сильновыпуклых функций (см. [ускорение Нестерова](https://blogs.princeton.edu/imabandit/2013/04/01/acceleratedgradientdescent/))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Резюме
# 1. Методы спуска
# 2. Направление убывания
# 3. Метод градиентного спуска
# 4. Правила выбора шага
# 5. Теоремы сходимости
# 6. Эксперименты
| Spring2021/intro_gd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="QHMl5iOcxiQm"
# [](https://colab.research.google.com/github/MobleyLab/drug-computing/blob/master/uci-pharmsci/Getting_Started_condacolab.ipynb)
# + [markdown] id="7uWUhEZk_BKQ"
# # Getting Started with Condacolab
# This notebook describes the steps to set up condacolab, which is another method of using the conda package manager on google colab notebooks.
#
# Here is a link to the package [repository](https://github.com/conda-incubator/condacolab/)
# + colab={"base_uri": "https://localhost:8080/"} id="R4srDnu0_Utk" outputId="c676025b-b834-46e2-bbce-50f3b082f42f"
# !pip install -q condacolab
import condacolab
condacolab.install()
# + [markdown] id="W69bjdHkAe2n"
# Make sure that the kernel restarts so that you can run the cell below. It should be ensured that the above code block should be run first before running any other cells since it restarts the kernel and resets the variables.
# + colab={"base_uri": "https://localhost:8080/"} id="A30gwGyfAjhd" outputId="89a57efc-f21d-4c64-dae7-3f914b995995"
import condacolab
condacolab.check()
# + [markdown] id="1oyPEmeYA7YJ"
# # Package Installation
# The section below will describe the necessary steps to install packages required for the course. Please make sure to run them in the appropriate order.
#
# Note: **We will be using mamba here instead of conda since it is faster. Mamba is more or less a drop-in replacement for conda, so if you are using mamba in lectures/assignments you should switch the `conda` command to the `mamba` command everywhere it occurs.**
#
# In this section we will install libgfortran and openeye toolkits. **Depending on the lecture and/or assignment, you may or may not need either or both of these**.
#
# `libgfortran` is needed mainly for the energy minimization, MD and MC assignments and the MD and MC lectures, where we compile fortran code. If you are not working with `f2py3`, you will not need libgfortran.
#
# The OpenEye toolkits are used more frequently, for general cheminformatics work. Still, they are only used in some notebooks, so it's only necessary to install them if the assignment or lecture contains `import openeye` or similar.
# + [markdown] id="wj4m2U8uBXUm"
# ## Installation of Fortran
# This involves the installation of 3 packages, gcc (fortran compiler for conda), libgfortran (fortran library for conda) and f2py (which allows us to convert fortran code to python outputs).
# + colab={"base_uri": "https://localhost:8080/"} id="PoMiN1kMBWEn" outputId="89999ec6-9c04-4a4f-b6ec-97cf23876c37"
# !mamba install gcc -c conda-forge
# + colab={"base_uri": "https://localhost:8080/"} id="iJMQadvlCre5" outputId="38f89a37-8847-4b5b-ba78-12f58b29d881"
# !mamba install gcc libgfortran
# + colab={"base_uri": "https://localhost:8080/"} id="ZQ1PWl65DDWv" outputId="461b6bef-3f50-44fe-f491-45a9c39fb3db"
# !mamba install numpy
# + [markdown] id="VfhHqvnzD5Ae"
# ## Installation of Plotting libraries and MD related packages.
# There are a number of additional packages that need to be installed for running MD. In this section we will install them using mamaba and the appropriate channels.
# + colab={"base_uri": "https://localhost:8080/"} id="Ow7Z08eUEVV1" outputId="7acd65b9-53cc-4ddc-c0e6-1884e31dd38b"
# !mamba install openmm openforcefield parmed yank openmoltools pdbfixer solvationtoolkit -c omnia
# + colab={"base_uri": "https://localhost:8080/"} id="8mWaYJh-GIYQ" outputId="f9ff44af-a10e-4a21-da3f-a08e980ff57d"
# !mamba install nb_conda mpld3 scikit-learn seaborn numpy matplotlib bokeh -c conda-forge
# + colab={"base_uri": "https://localhost:8080/"} id="G8ZOgWQYGXN4" outputId="f78ccca6-92c3-4281-d653-e7835b67a93d"
# !mamba install nglview -c bioconda
# + [markdown] id="9s9a84nIGhBy"
# ## Installation of OpenEye-Toolkits
# These toolkits will be utilized quite often in the course and rely on a license file to be used. Please make sure to have the license file uploaded onto google drive.
# + colab={"base_uri": "https://localhost:8080/"} id="wqu009PiGoLS" outputId="807801f7-6487-42dc-be6c-486954d6b6e1"
# !mamba install openeye-toolkits -c openeye
# + [markdown] id="DBtyRrrWHBq7"
# The code block below checks if your openeye-toolkits are activated. Make sure to place the license file on your google drive.
# + colab={"base_uri": "https://localhost:8080/"} id="3jUwtIRYG58b" outputId="bad3e5df-33fd-4bfd-8cd7-e68bede872cd"
# Mount Google Drive. NOTE: Do NOT move this before install of miniconda and other packages above or you may encounter path issues.
from google.colab import drive
drive.mount('/content/drive',force_remount = True)
license_filename = '/content/drive/MyDrive/oe_license.txt'
import os
import openeye
if os.path.isfile(license_filename):
license_file = open(license_filename, 'r')
openeye.OEAddLicenseData(license_file.read())
license_file.close()
else:
print("Error: Your OpenEye license is not readable; please check your filename and that you have mounted your Google Drive")
licensed = openeye.oechem.OEChemIsLicensed()
print("Was your OpenEye license correctly installed (True/False)? " + str(licensed))
if not licensed:
print("Error: Your OpenEye license is not correctly installed.")
raise Exception("Error: Your OpenEye license is not correctly installed.")
# -
| uci-pharmsci/Getting_Started_condacolab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Predefined colors in Python
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.colors as mpl_colors
# %matplotlib notebook
# ### What methods/attributes can be used?
dir(mpl_colors)
# ## Plotting the available base colors
bcolors = mpl_colors.BASE_COLORS
# +
fig, ax = plt.subplots()
# x, y = np.random.rand(len(bcolors)), np.random.rand(len(bcolors))
mu, sigma = 0, 0.5 # mean and standard deviation
x = np.random.normal(mu, sigma, len(bcolors))
y = np.random.normal(mu, sigma, len(bcolors))
s = np.random.randint(1000, high=2000, size=len(bcolors))
ax.scatter(x, y, s=s, c=list(bcolors.keys()), edgecolors='k')
ax.set_title('Base colors in matplotlib', fontsize=12, color='dodgerblue', fontweight='bold')
# ax.axes.get_xaxis().set_visible(False)
# ax.axes.get_yaxis().set_visible(False)
ax.axis('off')
if False:
plt.savefig('base_colors.png', dpi=300)
# -
# ### Create a dictionary to harbor all the available colors, bothe the base colors and the larger list of named colors
ncolors = mpl_colors.CSS4_COLORS
bn_colors = dict(**bcolors, **ncolors)
# ### Sort colors using: hue, saturation, value, name
func = lambda c: mpl_colors.rgb_to_hsv(mpl_colors.to_rgba(c)[:3])
aux_hsv = [[list(func(color_x)), name_x] for name_x, color_x in bn_colors.items()]
aux_hsv = sorted(aux_hsv)
cnames_sorted = [name for hsv, name in aux_hsv]
# ### Let's do a quick inpection
fig, ax = plt.subplots(figsize=(8, 8))
x = np.linspace(-10, 10, len(cnames_sorted))
y = np.power(x, 2)
ax.scatter(x, y, c=cnames_sorted, s=40)
# ### Now, plot of the available color, getting some ideas from the matplotlib example
# +
n = len(cnames_sorted)
ncols = 4
nrows = n // ncols + 1
fig, ax = plt.subplots(figsize=(10, 10))
X, Y = fig.get_dpi() * fig.get_size_inches()
h = Y / (nrows + 1)
w = X / ncols
for i, name in enumerate(cnames_sorted):
col = i % ncols
row = i // ncols
y = Y - (row * h) - h
xi = w * (col + 0.05)
# xf_line = w * (col + 0.25)
xi_text = w * (col + 0.13)
ax.text(xi_text, y, name, fontsize=(h * 0.8),
horizontalalignment='left',
verticalalignment='center')
ax.scatter(xi, y, c=name, s=np.random.randint(150, high=250),
edgecolor='k', lw=0.2)
ax.set_xlim(0, X)
ax.set_ylim(0, Y)
ax.set_axis_off()
fig.subplots_adjust(left=0, right=1,
top=1, bottom=0,
hspace=0.2, wspace=0.2)
if False:
fig.savefig('named_colors.png', dpi=300)
# -
| code/p01_python_colors/python_colors.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Q#
# language: qsharp
# name: iqsharp
# ---
# # Single-Qubit Gates
#
# This tutorial introduces you to single-qubit gates. Quantum gates are the quantum counterpart to classical logic gates, acting as the building blocks of quantum algorithms. Quantum gates transform qubit states in various ways, and can be applied sequentially to perform complex quantum calculations. Single-qubit gates, as their name implies, act on individual qubits. You can learn more at [Wikipedia](https://en.wikipedia.org/wiki/Quantum_logic_gate).
#
# We recommend to go through the [tutorial that introduces the concept of qubit](../Qubit/Qubit.ipynb) before starting this one.
#
# This tutorial covers the following topics:
#
# * Matrix representation
# * Ket-bra representation
# * The most important single-qubit gates
#
# A quick summary of the gates can also be found in [this cheatsheet](https://github.com/microsoft/QuantumKatas/blob/main/quickref/qsharp-quick-reference.pdf).
# # The Basics
#
# There are certain properties common to all quantum gates. This section will introduce those properties, using the $X$ gate as an example.
# ## Matrix Representation
#
# Quantum gates are represented as $2^N \times 2^N$ [unitary matrices](../LinearAlgebra/LinearAlgebra.ipynb#Unitary-Matrices), where $N$ is the number of qubits the gate operates on.
# As a quick reminder, a unitary matrix is a square matrix whose inverse is its adjoint.
# Single-qubit gates are represented by $2 \times 2$ matrices.
# Our example for this section, the $X$ gate, is represented by the following matrix:
#
# $$\begin{bmatrix} 0 & 1 \\ 1 & 0 \end{bmatrix}$$
#
# You may recall that the state of a qubit is represented by a vector of size $2$. You can apply a gate to a qubit by [multiplying](../LinearAlgebra/LinearAlgebra.ipynb#Matrix-Multiplication) the gate's matrix by the qubit's state vector. The result will be another vector, representing the new state of the qubit. For example, applying the $X$ gate to the computational basis states looks like this:
#
# $$X|0\rangle =
# \begin{bmatrix} 0 & 1 \\ 1 & 0 \end{bmatrix}
# \begin{bmatrix} 1 \\ 0 \end{bmatrix} =
# \begin{bmatrix} 0 \cdot 1 + 1 \cdot 0 \\ 1 \cdot 1 + 0 \cdot 0 \end{bmatrix} =
# \begin{bmatrix} 0 \\ 1 \end{bmatrix} \\
# X|1\rangle =
# \begin{bmatrix} 0 & 1 \\ 1 & 0 \end{bmatrix}
# \begin{bmatrix} 0 \\ 1 \end{bmatrix} =
# \begin{bmatrix} 0 \cdot 0 + 1 \cdot 1 \\ 1 \cdot 0 + 0 \cdot 1 \end{bmatrix} =
# \begin{bmatrix} 1 \\ 0 \end{bmatrix}$$
#
# The general case:
#
# $$|\psi\rangle = \alpha|0\rangle + \beta|1\rangle \\
# X|\psi\rangle =
# \begin{bmatrix} 0 & 1 \\ 1 & 0 \end{bmatrix}
# \begin{bmatrix} \alpha \\ \beta \end{bmatrix} =
# \begin{bmatrix} 0 \cdot \alpha + 1 \cdot \beta \\ 1 \cdot \alpha + 0 \cdot \beta \end{bmatrix} =
# \begin{bmatrix} \beta \\ \alpha \end{bmatrix}$$
#
# > If you need a reminder of what $|0\rangle$, $|1\rangle$, and $|\psi\rangle$ mean, you can review the section on [Dirac notation](../Qubit/Qubit.ipynb#Dirac-Notation) in the previous tutorial.
#
# Because this is the most common way to represent quantum gates, the terms "gate" and "gate matrix" will be used interchangeably in this tutorial.
#
# Applying several quantum gates in sequence is equivalent to performing several of these multiplications.
# For example, if you have gates $A$ and $B$ and a qubit in state $|\psi\rangle$, the result of applying $A$ followed by $B$ to that qubit would be $B\big(A|\psi\rangle\big)$ (the gates closest to the qubit state get applied first).
# Matrix multiplication is associative, so this is equivalent to multiplying the $B$ matrix by the $A$ matrix, producing a compound gate of the two, and then applying that to the qubit: $\big(BA\big)|\psi\rangle$.
#
# All quantum gates are reversible - there is another gate which will undo any given gate's transformation, returning the qubit to its original state.
# This means that when dealing with quantum gates, information about qubit states is never lost, as opposed to classical logic gates, some of which destroy information.
# Quantum gates are represented by unitary matrices, so the inverse of a gate is its adjoint; these terms are also used interchangeably in quantum computing.
# ## Effects on Basis States (Dirac Notation, Continued)
#
# There is a simple way to find out what a gate does to the two computational basis states ($|0\rangle$ and $|1\rangle$) from looking at its matrix that comes in handy when you want to work with states in Dirac notation. Consider an arbitrary gate:
#
# $$A = \begin{bmatrix} \epsilon & \zeta \\ \eta & \mu \end{bmatrix}$$
#
# Watch what happens when we apply it to these states:
#
# $$A|0\rangle =
# \begin{bmatrix} \epsilon & \zeta \\ \eta & \mu \end{bmatrix}
# \begin{bmatrix} 1 \\ 0 \end{bmatrix} =
# \begin{bmatrix} \epsilon \cdot 1 + \zeta \cdot 0 \\ \eta \cdot 1 + \mu \cdot 0 \end{bmatrix} =
# \begin{bmatrix} \epsilon \\ \eta \end{bmatrix} = \epsilon|0\rangle + \eta|1\rangle \\
# A|1\rangle =
# \begin{bmatrix} \epsilon & \zeta \\ \eta & \mu \end{bmatrix}
# \begin{bmatrix} 0 \\ 1 \end{bmatrix} =
# \begin{bmatrix} \epsilon \cdot 0 + \zeta \cdot 1 \\ \eta \cdot 0 + \mu \cdot 1 \end{bmatrix} =
# \begin{bmatrix} \zeta \\ \mu \end{bmatrix} = \zeta|0\rangle + \mu|1\rangle$$
#
# Notice that applying the gate to the $|0\rangle$ state transforms it into the state written as the first column of the gate's matrix. Likewise, applying the gate to the $|1\rangle$ state transforms it into the state written as the second column. This holds true for any quantum gate, including, of course, the $X$ gate:
#
# $$X = \begin{bmatrix} 0 & 1 \\ 1 & 0 \end{bmatrix} \\
# X|0\rangle = \begin{bmatrix} 0 \\ 1 \end{bmatrix} = |1\rangle \\
# X|1\rangle = \begin{bmatrix} 1 \\ 0 \end{bmatrix} = |0\rangle$$
#
# Once you understand how a gate affects the computational basis states, you can easily find how it affects any state.
# Recall that any qubit state vector can be written as a linear combination of the basis states:
#
# $$|\psi\rangle = \begin{bmatrix} \alpha \\ \beta \end{bmatrix} = \alpha|0\rangle + \beta|1\rangle$$
#
# Because matrix multiplication distributes over addition, once you know how a gate affects those two basis states, you can calculate how it affects any state:
#
# $$X|\psi\rangle = X\big(\alpha|0\rangle + \beta|1\rangle\big) = X\big(\alpha|0\rangle\big) + X\big(\beta|1\rangle\big) = \alpha X|0\rangle + \beta X|1\rangle = \alpha|1\rangle + \beta|0\rangle$$
#
# That is, applying a gate to a qubit in superposition is equivalent to applying that gate to the basis states that make up that superposition and adding the results with appropriate weights.
# ## Ket-bra Representation
#
# There is another way to represent quantum gates, this time using Dirac notation. However, the kets we've been using aren't enough to represent arbitrary matrices. We need to introduce another piece of notation: the **bra** (this is why Dirac notation is sometimes called **bra-ket notation**).
#
# Recall that kets represent column vectors; a bra is a ket's row vector counterpart. For any ket $|\psi\rangle$, the corresponding bra is its adjoint (conjugate transpose): $\langle\psi| = |\psi\rangle^\dagger$.
#
# > As a quick reminder, the [adjoint](../LinearAlgebra/LinearAlgebra.ipynb#Unary-Operations), also known as the conjugate transpose of a matrix, well, the conjugate of that matrix's transpose.
#
# Some examples:
#
# <table style="border:1px solid">
# <col width=150>
# <col width=150>
# <tr>
# <th style="text-align:center; border:1px solid">Ket</th>
# <th style="text-align:center; border:1px solid">Bra</th>
# </tr>
# <tr>
# <td style="text-align:center; border:1px solid">$|0\rangle = \begin{bmatrix} 1 \\ 0 \end{bmatrix}$</td>
# <td style="text-align:center; border:1px solid">$\langle0| = \begin{bmatrix} 1 & 0 \end{bmatrix}$</td>
# </tr>
# <tr>
# <td style="text-align:center; border:1px solid">$|1\rangle = \begin{bmatrix} 0 \\ 1 \end{bmatrix}$</td>
# <td style="text-align:center; border:1px solid">$\langle1| = \begin{bmatrix} 0 & 1 \end{bmatrix}$</td>
# </tr>
# <tr>
# <td style="text-align:center; border:1px solid">$|i\rangle = \begin{bmatrix} \frac{1}{\sqrt{2}} \\ \frac{i}{\sqrt{2}} \end{bmatrix}$</td>
# <td style="text-align:center; border:1px solid">$\langle i| = \begin{bmatrix} \frac{1}{\sqrt{2}} & -\frac{i}{\sqrt{2}} \end{bmatrix}$</td>
# </tr>
# <tr>
# <td style="text-align:center; border:1px solid">$|\psi\rangle = \begin{bmatrix} \alpha \\ \beta \end{bmatrix}$</td>
# <td style="text-align:center; border:1px solid">$\langle\psi| = \begin{bmatrix} \overline{\alpha} & \overline{\beta} \end{bmatrix}$</td>
# </tr>
# <tr>
# <td style="text-align:center; border:1px solid">$|\psi\rangle = \alpha|0\rangle + \beta|1\rangle$</td>
# <td style="text-align:center; border:1px solid">$\langle\psi| = \overline{\alpha}\langle0| + \overline{\beta}\langle1|$</td>
# </tr>
# </table>
# Kets and bras give us a neat way to express [inner](../LinearAlgebra/LinearAlgebra.ipynb#Inner-Product) and [outer](../LinearAlgebra/LinearAlgebra.ipynb#Outer-Product) products. The inner product of $|\phi\rangle$ and $|\psi\rangle$ is the matrix product of $\langle\phi|$ and $|\psi\rangle$, denoted as $\langle\phi|\psi\rangle$, and their outer product is the matrix product of $|\phi\rangle$ and $\langle\psi|$, denoted as $|\phi\rangle\langle\psi|$. Notice that the norm of $|\psi\rangle$ is $\sqrt{\langle\psi|\psi\rangle}$.
#
# This brings us to representing matrices. Recall that the outer product of two vectors of the same size produces a square matrix. We can use a linear combination of several outer products of simple vectors (such as basis vectors) to express any square matrix. For example, the $X$ gate can be expressed as follows:
#
# $$X = |0\rangle\langle1| + |1\rangle\langle0| \\
# |0\rangle\langle1| + |1\rangle\langle0| =
# \begin{bmatrix} 1 \\ 0 \end{bmatrix}\begin{bmatrix} 0 & 1 \end{bmatrix} +
# \begin{bmatrix} 0 \\ 1 \end{bmatrix}\begin{bmatrix} 1 & 0 \end{bmatrix} =
# \begin{bmatrix} 0 & 1 \\ 0 & 0 \end{bmatrix} + \begin{bmatrix} 0 & 0 \\ 1 & 0 \end{bmatrix} =
# \begin{bmatrix} 0 & 1 \\ 1 & 0 \end{bmatrix}$$
#
# This representation can be used to carry out calculations in Dirac notation without ever switching back to matrix representation:
#
# $$X|0\rangle = \big(|0\rangle\langle1| + |1\rangle\langle0|\big)|0\rangle = |0\rangle\langle1|0\rangle + |1\rangle\langle0|0\rangle = |0\rangle\big(\langle1|0\rangle\big) + |1\rangle\big(\langle0|0\rangle\big) = |0\rangle(0) + |1\rangle(1) = |1\rangle$$
#
# > That last step may seem a bit confusing. Recall that $|0\rangle$ and $|1\rangle$ form an **orthonormal basis**. That is, they are both normalized, and they are orthogonal to each other.
# >
# > A vector is normalized if its norm is equal to $1$, which only happens if its inner product with itself is equal to $1$. This means that $\langle0|0\rangle = \langle1|1\rangle = 1$
# >
# > Two vectors are orthogonal to each other if their inner product equals $0$. This means that $\langle0|1\rangle = \langle 1|0\rangle = 0$.
# In general case, a matrix
# $$A = \begin{bmatrix} a_{00} & a_{01} \\ a_{10} & a_{11} \end{bmatrix}$$
# will have the following ket-bra representation:
# $$A = a_{00} |0\rangle\langle0| + a_{01} |0\rangle\langle1| + a_{10} |1\rangle\langle0| + a_{11} |1\rangle\langle1|$$
# > ## Ket-bra decomposition
# >
# > This section describes a more formal process of finding the ket-bra decompositions of quantum gates. This section is not necessary to start working with quantum gates, so feel free to skip it for now, and come back to it later.
# >
# > You can use the properties of [eigenvalues and eigenvectors](../LinearAlgebra/LinearAlgebra.ipynb#Part-III:-Eigenvalues-and-Eigenvectors) to find the ket-bra decomposition of any gate. Given a gate $A$, and its orthogonal eigenvectors $|\phi\rangle$ and $|\psi\rangle$, if:
# >
# > $$A|\phi\rangle = x_\phi|\phi\rangle \\
# A|\psi\rangle = x_\psi|\psi\rangle$$
# >
# > Then:
# >
# > $$A = x_\phi|\phi\rangle\langle\phi| + x_\psi|\psi\rangle\langle\psi|$$
# >
# > Let's use our $X$ gate as a simple example. The $X$ gate has two eigenvectors: $|+\rangle = \frac{1}{\sqrt{2}}\big(|0\rangle + |1\rangle\big)$ and $|-\rangle = \frac{1}{\sqrt{2}}\big(|0\rangle - |1\rangle\big)$. Their eigenvalues are $1$ and $-1$ respectively:
# >
# > $$X|+\rangle = |+\rangle \\
# X|-\rangle = -|-\rangle$$
# >
# > Here's what the decomposition looks like:
# >
# > $$X = |+\rangle\langle+| - |-\rangle\langle-| = \\
# = \frac{1}{2}\big[\big(|0\rangle + |1\rangle\big)\big(\langle0| + \langle1|\big) - \big(|0\rangle - |1\rangle\big)\big(\langle0| - \langle1|\big)\big] = \\
# = \frac{1}{2}\big(\color{red}{|0\rangle\langle0|} + |0\rangle\langle1| + |1\rangle\langle0| + \color{red}{|1\rangle\langle1|} - \color{red}{|0\rangle\langle0|} + |0\rangle\langle1| + |1\rangle\langle0| - \color{red}{|1\rangle\langle1|}\big) = \\
# = \frac{1}{2}\big(2|0\rangle\langle1| + 2|1\rangle\langle0|\big) = |0\rangle\langle1| + |1\rangle\langle0|$$
# # Important Gates
#
# This section introduces some of the common single-qubit gates, including their matrix form, their ket-bra decomposition, and a brief "cheatsheet" listing their effect on some common qubit states.
#
# You can use a tool called [Quirk](https://algassert.com/quirk) to visualize how these gates interact with various qubit states.
#
# This section relies on the following notation:
#
# <table>
# <col width=180>
# <col width=180>
# <tr>
# <td style="text-align:center; border:1px solid">$|+\rangle = \frac{1}{\sqrt{2}}\big(|0\rangle + |1\rangle\big)$</td>
# <td style="text-align:center; border:1px solid">$|-\rangle = \frac{1}{\sqrt{2}}\big(|0\rangle - |1\rangle\big)$</td>
# </tr>
# <tr>
# <td style="text-align:center; border:1px solid">$|i\rangle = \frac{1}{\sqrt{2}}\big(|0\rangle + i|1\rangle\big)$</td>
# <td style="text-align:center; border:1px solid">$|-i\rangle = \frac{1}{\sqrt{2}}\big(|0\rangle - i|1\rangle\big)$</td>
# </tr>
# </table>
# ## Pauli Gates
#
# The Pauli gates, named after [Wolfgang Pauli](https://en.wikipedia.org/wiki/Wolfgang_Pauli), are based on the so-called **Pauli matrices**. All three Pauli gates are **self-adjoint**, meaning that each one is its own inverse.
#
# <table style="border:1px solid">
# <col width=50>
# <col width=50>
# <col width=150>
# <col width=200>
# <col width=150>
# <col width=50>
# <tr>
# <th style="text-align:center; border:1px solid">Gate</th>
# <th style="text-align:center; border:1px solid">Matrix</th>
# <th style="text-align:center; border:1px solid">Ket-Bra</th>
# <th style="text-align:center; border:1px solid">Applying to $|\psi\rangle = \alpha|0\rangle + \beta|1\rangle$</th>
# <th style="text-align:center; border:1px solid">Applying to basis states</th>
# <th style="text-align:center; border:1px solid">Q# Documentation</th>
# </tr>
# <tr>
# <td style="text-align:center; border:1px solid">$X$</td>
# <td style="text-align:center; border:1px solid">$\begin{bmatrix} 0 & 1 \\ 1 & 0 \end{bmatrix}$</td>
# <td style="text-align:center; border:1px solid">$|0\rangle\langle1| + |1\rangle\langle0|$</td>
# <td style="text-align:center; border:1px solid">$X|\psi\rangle = \alpha|1\rangle + \beta|0\rangle$</td>
# <td style="text-align:center; border:1px solid">$X|0\rangle = |1\rangle \\
# X|1\rangle = |0\rangle \\
# X|+\rangle = |+\rangle \\
# X|-\rangle = -|-\rangle \\
# X|i\rangle = i|-i\rangle \\
# X|-i\rangle = -i|i\rangle$ </td>
# <td style="text-align:center; border:1px solid"><a href="https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic.x">X</a></td>
# </tr>
# <tr>
# <td style="text-align:center; border:1px solid">$Y$</td>
# <td style="text-align:center; border:1px solid">$\begin{bmatrix} 0 & -i \\ i & 0 \end{bmatrix}$</td>
# <td style="text-align:center; border:1px solid">$i(|1\rangle\langle0| - |0\rangle\langle1|)$</td>
# <td style="text-align:center; border:1px solid">$Y|\psi\rangle = i\big(\alpha|1\rangle - \beta|0\rangle\big)$</td>
# <td style="text-align:center; border:1px solid">$Y|0\rangle = i|1\rangle \\
# Y|1\rangle = -i|0\rangle \\
# Y|+\rangle = -i|-\rangle \\
# Y|-\rangle = i|+\rangle \\
# Y|i\rangle = |i\rangle \\
# Y|-i\rangle = -|-i\rangle$</td>
# <td style="text-align:center; border:1px solid"><a href="https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic.y">Y</a></td>
# </tr>
# <tr>
# <td style="text-align:center; border:1px solid">$Z$</td>
# <td style="text-align:center; border:1px solid">$\begin{bmatrix} 1 & 0 \\ 0 & -1 \end{bmatrix}$</td>
# <td style="text-align:center; border:1px solid">$|0\rangle\langle0| - |1\rangle\langle1|$</td>
# <td style="text-align:center; border:1px solid">$Z|\psi\rangle = \alpha|0\rangle - \beta|1\rangle$</td>
# <td style="text-align:center; border:1px solid">$Z|0\rangle = |0\rangle \\
# Z|1\rangle = -|1\rangle \\
# Z|+\rangle = |-\rangle \\
# Z|-\rangle = |+\rangle \\
# Z|i\rangle = |-i\rangle \\
# Z|-i\rangle = |i\rangle$ </td>
# <td style="text-align:center; border:1px solid"><a href="https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic.z">Z</a></td>
# </tr>
# </table>
# > The $X$ gate is sometimes referred to as the **bit flip** gate, or the **NOT** gate, because it acts like the classical NOT gate on the computational basis.
# >
# > The $Z$ gate is sometimes referred to as the **phase flip** gate.
#
# Here are several properties of the Pauli gates that are easy to verify and convenient to remember:
#
# * Different Pauli gates *anti-commute*:
# $$XZ = -ZX, XY = -YX, YZ = -ZY$$
# * A product of any two Pauli gates equals the third gate, with an extra $i$ (or $-i$) phase:
# $$XY = iZ, YZ = iX, ZX = iY$$
# * A product of all three Pauli gates equals identity (with an extra $i$ phase):
# $$XYZ = iI$$
# ### <span style="color:blue">Demo: Pauli Gates</span>
#
# The following cell contains code demonstrating how to apply gates in Q#, using the Pauli $X$ gate as an example. It sets up a series of quantum states, and then shows the result of applying the $X$ gate to each one. To run the demo, run the next cell using `Ctrl+Enter` (`⌘+Enter` on a Mac).
#
# In the previous tutorial we discussed that the qubit state in Q# cannot be directly assigned or accessed. The same logic is extended to the quantum gates: applying a gate to a qubit modifies the internal state of that qubit but doesn't return the resulting state of the qubit. This is why we never assign the output of these gates to any variables in this demo - they don't produce any output.
#
# Applying several gates in a row follows the same principle. In the mathematical notation applying an $X$ gate followed by a $Z$ gate to a state $|\psi\rangle$ is denoted as $Z(X(|\psi\rangle))$, because the result of applying a gate to a state is another state. In Q#, applying a gate doesn't return anything, so you can't use its output as an input to another gate - something like `Z(X(q))` will not produce expected result. Instead, to apply several gates to the same qubit, you need to call them separately in the order in which they are applied:
#
# ```
# X(q);
# Z(q);
# ```
#
# All the basic gates we will be covering in this tutorial are part of the [Intrinsic](https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic) namespace. We're also using the function [DumpMachine](https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.diagnostics.dumpmachine) to print the state of the quantum simulator.
# +
// Run this cell using Ctrl+Enter (⌘+Enter on Mac)
// Run the next cell to see the output
// To use a namespace, you need to use the `open` keyword to access it
open Microsoft.Quantum.Diagnostics;
operation PauliGatesDemo () : Unit {
// This allocates a qubit for us to work with
use q = Qubit();
// This will put the qubit into an uneven superposition |𝜓❭,
// where the amplitudes of |0⟩ and |1⟩ have different moduli
Ry(1.0, q);
Message("Qubit in state |𝜓❭:");
DumpMachine();
// Let's apply the X gate; notice how it swaps the amplitudes of the |0❭ and |1❭ basis states
X(q);
Message("Qubit in state X|𝜓❭:");
DumpMachine();
// Applying the Z gate adds -1 relative phase to the |1❭ basis states
Z(q);
Message("Qubit in state ZX|𝜓❭:");
DumpMachine();
// Finally, applying the Y gate returns the qubit to its original state |𝜓❭, with an extra global phase of i
Y(q);
Message("Qubit in state YZX|𝜓❭:");
DumpMachine();
// This returns the qubit into state |0❭
Reset(q);
}
# -
# In the previous tutorials we used `%simulate` command to run the Q# code on the full-state simulator. Here we will use an additional `%trace` command: it will print the circuit diagram of the run after the output.
%simulate PauliGatesDemo
%trace PauliGatesDemo
# ## Exercises
#
# The following exercises are designed to test your understanding of the concepts you've learned so far.
# In each exercise your task is to implement an operation that applies a particular transformation to a qubit.
# Unlike the demos you have seen so far, you don't have to allocate the qubit or to put it into a certain initial state - the qubit is already allocated, prepared in some state and provided to you as an input to the operation.
#
# ### <span style="color:blue">Exercise 1</span>: The $Y$ gate
#
# **Input:** A qubit in an arbitrary state $|\psi\rangle = \alpha|0\rangle + \beta|1\rangle$.
#
# **Goal:** Apply the $Y$ gate to the qubit, i.e., transform the given state into $i\alpha|1\rangle - i\beta|0\rangle$.
# +
%kata T1_ApplyY
operation ApplyY (q : Qubit) : Unit is Adj+Ctl {
// Fill in your code here, then run the cell to test your work.
// For this exercise, just apply the Y gate.
Y(q);
}
# -
# *Can't come up with a solution? See the explained solution in the [Single-Qubit Gates Workbook](./Workbook_SingleQubitGates.ipynb#Exercise-1:-The-$Y$-gate).*
# ### <span style="color:blue">Exercise 2</span>: Applying a global phase $i$
#
# **Input:** A qubit in an arbitrary state $|\psi\rangle = \alpha|0\rangle + \beta|1\rangle$.
#
# **Goal:** Use several Pauli gates to change the qubit state to $i|\psi\rangle = i\alpha|0\rangle + i\beta|1\rangle$.
# +
%kata T2_GlobalPhaseI
operation GlobalPhaseI (q : Qubit) : Unit is Adj+Ctl {
Z(q);
Y(q);
X(q);
}
# -
# *Can't come up with a solution? See the explained solution in the [Single-Qubit Gates Workbook](./Workbook_SingleQubitGates.ipynb#Exercise-2:-Applying-a-global-phase-$i$).*
# ### <span style="color:blue">Exercise 3</span>*: Applying a $-1$ phase to $|0\rangle$ state
#
# **Input:** A qubit in an arbitrary state $|\psi\rangle = \alpha|0\rangle + \beta|1\rangle$.
#
# **Goal:** Use several Pauli gates to change the qubit state to $- \alpha|0\rangle + \beta|1\rangle$, i.e., apply the transformation represented by the following matrix::
#
# $$\begin{bmatrix} -1 & 0 \\ 0 & 1 \end{bmatrix}$$
#
# <br/>
# <details>
# <summary><strong>Need a hint? Click here</strong></summary>
# Experiment with different sequences of Pauli gates and observe their effect on the state.
# </details>
# +
%kata T3_SignFlipOnZero
operation SignFlipOnZero (q : Qubit) : Unit is Adj+Ctl {
X(q);
Z(q);
X(q);
}
# -
# *Can't come up with a solution? See the explained solution in the [Single-Qubit Gates Workbook](./Workbook_SingleQubitGates.ipynb#exercise-3).*
# ## Identity
#
# The identity gate is mostly here for completeness, at least for now. It will come in handy when dealing with multi-qubit systems and multi-qubit gates. It is represented by the identity matrix, and does not affect the state of the qubit.
#
# <table style="border:1px solid">
# <col width=50>
# <col width=50>
# <col width=150>
# <col width=200>
# <col width=50>
# <tr>
# <th style="text-align:center; border:1px solid">Gate</th>
# <th style="text-align:center; border:1px solid">Matrix</th>
# <th style="text-align:center; border:1px solid">Ket-Bra</th>
# <th style="text-align:center; border:1px solid">Applying to $|\psi\rangle = \alpha|0\rangle + \beta|1\rangle$</th>
# <th style="text-align:center; border:1px solid">Q# Documentation</th>
# </tr>
# <tr>
# <td style="text-align:center; border:1px solid">$I$</td>
# <td style="text-align:center; border:1px solid">$\begin{bmatrix} 1 & 0 \\ 0 & 1 \end{bmatrix}$</td>
# <td style="text-align:center; border:1px solid">$|0\rangle\langle0| + |1\rangle\langle1|$</td>
# <td style="text-align:center; border:1px solid">$I|\psi\rangle = |\psi\rangle$</td>
# <td style="text-align:center; border:1px solid"><a href="https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic.i">I</a></td>
# </tr>
# </table>
# ## Hadamard
#
# The **Hadamard** gate is an extremely important quantum gate. Unlike the previous gates, applying the Hadamard gate to a qubit in a computational basis state puts that qubit into a superposition.
# Like the Pauli gates, the Hadamard gate is self-adjoint.
#
# <table style="border:1px solid">
# <col width=50>
# <col width=200>
# <col width=150>
# <col width=260>
# <col width=150>
# <col width=50>
# <tr>
# <th style="text-align:center; border:1px solid">Gate</th>
# <th style="text-align:center; border:1px solid">Matrix</th>
# <th style="text-align:center; border:1px solid">Ket-Bra</th>
# <th style="text-align:center; border:1px solid">Applying to $|\psi\rangle = \alpha|0\rangle + \beta|1\rangle$</th>
# <th style="text-align:center; border:1px solid">Applying to basis states</th>
# <th style="text-align:center; border:1px solid">Q# Documentation</th>
# </tr>
# <tr>
# <td style="text-align:center; border:1px solid">$H$</td>
# <td style="text-align:center; border:1px solid">$\begin{bmatrix} \frac{1}{\sqrt{2}} & \frac{1}{\sqrt{2}} \\ \frac{1}{\sqrt{2}} & -\frac{1}{\sqrt{2}} \end{bmatrix} = \frac{1}{\sqrt{2}}\begin{bmatrix} 1 & 1 \\ 1 & -1 \end{bmatrix}$</td>
# <td style="text-align:center; border:1px solid">$|0\rangle\langle+| + |1\rangle\langle-|$</td>
# <td style="text-align:center; border:1px solid">$H|\psi\rangle = \alpha|+\rangle + \beta|-\rangle = \frac{\alpha + \beta}{\sqrt{2}}|0\rangle + \frac{\alpha - \beta}{\sqrt{2}}|1\rangle$</td>
# <td style="text-align:center; border:1px solid">$H|0\rangle = |+\rangle \\
# H|1\rangle = |-\rangle \\
# H|+\rangle = |0\rangle \\
# H|-\rangle = |1\rangle \\
# H|i\rangle = e^{i\pi/4}|-i\rangle \\
# H|-i\rangle = e^{-i\pi/4}|i\rangle$</td>
# <td style="text-align:center; border:1px solid"><a href="https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic.h">H</a></td>
# </tr>
# </table>
#
# > As a reminder, $e^{i\pi/4} = \frac{1}{\sqrt2} (1 + i)$ and $e^{-i\pi/4} = \frac{1}{\sqrt2} (1 - i)$.
# > If you need a refresher on calculating expressions like $e^{i\theta}$, you should review the section on [complex exponentiation](../ComplexArithmetic/ComplexArithmetic.ipynb#Imaginary-Exponents).
# ### <span style="color:blue">Exercise 4</span>: Preparing a $|-\rangle$ state
#
# **Input:** A qubit in state $|0\rangle$.
#
# **Goal:** Transform the qubit into state $|-\rangle$.
# +
%kata T4_PrepareMinus
operation PrepareMinus (q : Qubit) : Unit is Adj+Ctl {
X(q);
H(q);
}
# -
# *Can't come up with a solution? See the explained solution in the [Single-Qubit Gates Workbook](./Workbook_SingleQubitGates.ipynb#Exercise-4:-Preparing-a-$|-\rangle$-state).*
# ## Phase Shift Gates
#
# The next two gates are known as phase shift gates. They apply a phase to the $|1\rangle$ state, and leave the $|0\rangle$ state unchanged.
#
# <table style="border:1px solid">
# <col width=50>
# <col width=50>
# <col width=150>
# <col width=200>
# <col width=150>
# <col width=50>
# <tr>
# <th style="text-align:center; border:1px solid">Gate</th>
# <th style="text-align:center; border:1px solid">Matrix</th>
# <th style="text-align:center; border:1px solid">Ket-Bra</th>
# <th style="text-align:center; border:1px solid">Applying to $|\psi\rangle = \alpha|0\rangle + \beta|1\rangle$</th>
# <th style="text-align:center; border:1px solid">Applying to basis states</th>
# <th style="text-align:center; border:1px solid">Q# Documentation</th>
# </tr>
# <tr>
# <td style="text-align:center; border:1px solid">$S$</td>
# <td style="text-align:center; border:1px solid">$\begin{bmatrix} 1 & 0 \\ 0 & i \end{bmatrix}$</td>
# <td style="text-align:center; border:1px solid">$|0\rangle\langle0| + i|1\rangle\langle1|$</td>
# <td style="text-align:center; border:1px solid">$S|\psi\rangle = \alpha|0\rangle + i\beta|1\rangle$</td>
# <td style="text-align:center; border:1px solid">$S|0\rangle = |0\rangle \\
# S|1\rangle = i|1\rangle \\
# S|+\rangle = |i\rangle \\
# S|-\rangle = |-i\rangle \\
# S|i\rangle = |-\rangle \\
# S|-i\rangle = |+\rangle$</td>
# <td style="text-align:center; border:1px solid"><a href="https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic.s">S</a></td>
# </tr>
# <tr>
# <td style="text-align:center; border:1px solid">$T$</td>
# <td style="text-align:center; border:1px solid">$\begin{bmatrix} 1 & 0 \\ 0 & e^{i\pi/4} \end{bmatrix}$</td>
# <td style="text-align:center; border:1px solid">$|0\rangle\langle0| + e^{i\pi/4}|1\rangle$$\langle1|$</td>
# <td style="text-align:center; border:1px solid">$T|\psi\rangle = \alpha|0\rangle + e^{i\pi/4} \beta |1\rangle$</td>
# <td style="text-align:center; border:1px solid">$T|0\rangle = |0\rangle \\
# T|1\rangle = e^{i\pi/4}|1\rangle$</td>
# <td style="text-align:center; border:1px solid"><a href="https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic.t">T</a></td>
# </tr>
# </table>
#
# > Notice that applying the $T$ gate twice is equivalent to applying the $S$ gate, and applying the $S$ gate twice is equivalent to applying the $Z$ gate:
# $$T^2 = S, S^2 = Z$$
# ### <span style="color:blue">Exercise 5</span>: Three-fourths phase
#
# **Input:** A qubit in an arbitrary state $|\psi\rangle = \alpha|0\rangle + \beta|1\rangle$.
#
# **Goal:** Use several phase shift gates to apply the transformation represented by the following matrix to the given qubit:
#
# $$\begin{bmatrix} 1 & 0 \\ 0 & e^{3i\pi/4} \end{bmatrix}$$
#
# <br/>
# <details>
# <summary><strong>Need a hint? Click here</strong></summary>
# As a reminder, $i = e^{i\pi/2}$.
# </details>
# +
%kata T5_ThreeQuatersPiPhase
operation ThreeQuatersPiPhase (q : Qubit) : Unit is Adj+Ctl {
T(q);
T(q);
T(q);
}
# -
# *Can't come up with a solution? See the explained solution in the [Single-Qubit Gates Workbook](./Workbook_SingleQubitGates.ipynb#Exercise-5:-Three-fourths-phase).*
# ## Rotation Gates
#
# The next few gates are parametrized: their exact behavior depends on a numeric parameter - an angle $\theta$, given in radians.
# These gates are the $X$ rotation gate $R_x(\theta)$, $Y$ rotation gate $R_y(\theta)$, $Z$ rotation gate $R_z(\theta)$, and the arbitrary phase gate $R_1(\theta)$.
# Note that for the first three gates the parameter $\theta$ is multiplied by $\frac{1}{2}$ within the gate's matrix.
#
# > These gates are known as rotation gates, because they represent rotations around various axes on the Bloch sphere. The Bloch sphere is a way of representing the qubit states visually, mapping them onto the surface of a sphere.
# > Unfortunately, this visualization isn't very useful beyond single-qubit states, which is why we have opted not to go into details in this tutorial series.
# > If you are curious about it, you can learn more in [this slide deck](http://www.vcpc.univie.ac.at/~ian/hotlist/qc/talks/bloch-sphere.pdf) or in [Wikipedia article](https://en.wikipedia.org/wiki/Bloch_sphere).
#
# <table style="border:1px solid">
# <col width=50>
# <col width=100>
# <col width=400>
# <col width=250>
# <col width=50>
# <tr>
# <th style="text-align:center; border:1px solid">Gate</th>
# <th style="text-align:center; border:1px solid">Matrix</th>
# <th style="text-align:center; border:1px solid">Applying to $|\psi\rangle = \alpha|0\rangle + \beta|1\rangle$</th>
# <th style="text-align:center; border:1px solid">Applying to basis states</th>
# <th style="text-align:center; border:1px solid">Q# Documentation</th>
# </tr>
# <tr>
# <td style="text-align:center; border:1px solid">$R_x(\theta)$</td>
# <td style="text-align:center; border:1px solid">$\begin{bmatrix} \cos\frac{\theta}{2} & -i\sin\frac{\theta}{2} \\ -i\sin\frac{\theta}{2} & \cos\frac{\theta}{2} \end{bmatrix}$</td>
# <td style="text-align:center; border:1px solid">$R_x(\theta)|\psi\rangle = (\alpha\cos\frac{\theta}{2} - i\beta\sin\frac{\theta}{2})|0\rangle + (\beta\cos\frac{\theta}{2} - i\alpha\sin\frac{\theta}{2})|1\rangle$</td>
# <td style="text-align:center; border:1px solid">$R_x(\theta)|0\rangle = \cos\frac{\theta}{2}|0\rangle - i\sin\frac{\theta}{2}|1\rangle \\
# R_x(\theta)|1\rangle = \cos\frac{\theta}{2}|1\rangle - i\sin\frac{\theta}{2}|0\rangle$</td>
# <td style="text-align:center; border:1px solid"><a href="https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic.rx">Rx</a></td>
# </tr>
# <tr>
# <td style="text-align:center; border:1px solid">$R_y(\theta)$</td>
# <td style="text-align:center; border:1px solid">$\begin{bmatrix} \cos\frac{\theta}{2} & -\sin\frac{\theta}{2} \\ \sin\frac{\theta}{2} & \cos\frac{\theta}{2} \end{bmatrix}$</td>
# <td style="text-align:center; border:1px solid">$R_y(\theta)|\psi\rangle = (\alpha\cos\frac{\theta}{2} - \beta\sin\frac{\theta}{2})|0\rangle + (\beta\cos\frac{\theta}{2} + \alpha\sin\frac{\theta}{2})|1\rangle$</td>
# <td style="text-align:center; border:1px solid">$R_y(\theta)|0\rangle = \cos\frac{\theta}{2}|0\rangle + \sin\frac{\theta}{2}|1\rangle \\
# R_y(\theta)|1\rangle = \cos\frac{\theta}{2}|1\rangle - \sin\frac{\theta}{2}|0\rangle$</td>
# <td style="text-align:center; border:1px solid"><a href="https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic.ry">Ry</a></td>
# </tr>
# <tr>
# <td style="text-align:center; border:1px solid">$R_z(\theta)$</td>
# <td style="text-align:center; border:1px solid">$\begin{bmatrix} e^{-i\theta/2} & 0 \\ 0 & e^{i\theta/2} \end{bmatrix}$</td>
# <td style="text-align:center; border:1px solid">$R_z(\theta)|\psi\rangle = \alpha e^{-i\theta/2}|0\rangle + \beta e^{i\theta/2}|1\rangle$</td>
# <td style="text-align:center; border:1px solid">$R_z(\theta)|0\rangle = e^{-i\theta/2}|0\rangle \\
# R_z(\theta)|1\rangle = e^{i\theta/2}|1\rangle$</td>
# <td style="text-align:center; border:1px solid"><a href="https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic.rz">Rz</a></td>
# </tr>
# <tr>
# <td style="text-align:center; border:1px solid">$R_1(\theta)$</td>
# <td style="text-align:center; border:1px solid">$\begin{bmatrix} 1 & 0 \\ 0 & e^{i\theta} \end{bmatrix}$</td>
# <td style="text-align:center; border:1px solid">$R_1(\theta)|\psi\rangle = \alpha|0\rangle + \beta e^{i\theta}|1\rangle$</td>
# <td style="text-align:center; border:1px solid">$R_1(\theta)|0\rangle = |0\rangle \\
# R_1(\theta)|1\rangle = e^{i\theta}|1\rangle$</td>
# <td style="text-align:center; border:1px solid"><a href="https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic.r1">R1</a></td>
# </tr>
# </table>
#
# You have already encountered some special cases of the $R_1$ gate:
#
# $$T = R_1(\frac{\pi}{4}), S = R_1(\frac{\pi}{2}), Z = R_1(\pi)$$
#
# In addition, this gate is closely related to the $R_z$ gate: applying $R_1$ gate is equivalent to applying the $R_z$ gate, and then applying a global phase:
#
# $$R_1(\theta) = e^{i\theta/2}R_z(\theta)$$
#
# In addition, the rotation gates are very closely related to their respective Pauli gates:
#
# $$X = iR_x(\pi), Y = iR_y(\pi), Z = iR_z(\pi)$$
# ### <span style="color:blue">Exercise 6</span>: Preparing a rotated state
#
# **Inputs:**
#
# 1. Real numbers $\alpha$ and $\beta$ such that $\alpha^2 + \beta^2 = 1$.
# 3. A qubit in state $|0\rangle$.
#
# **Goal:** Use a rotation gate to transform the qubit into state $\alpha|0\rangle -i\beta|1\rangle$.
#
# > You will probably need functions from the [Math](https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.math) namespace, specifically [ArcTan2](https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.math.arctan2).
# >
# > You can assign variables in Q# by using the `let` keyword: `let num = 3;` or `let result = Function(input);`
#
# <details>
# <summary><strong>Need a hint? Click here</strong></summary>
# Don't forget, you can tell what a matrix does to the basis states by looking at its matrix: the first column of the matrix is the state into which it will transform the $|0\rangle$ state.
# </details>
# +
%kata T6_PrepareRotatedState
open Microsoft.Quantum.Math;
operation PrepareRotatedState (alpha : Double, beta : Double, q : Qubit) : Unit is Adj+Ctl {
// need the angle where cos(theta) = alpha, sin(theta) = beta
// or tan(theta) = sin(theta)/cos(theta)
// Then apply Rx gate.
let theta = ArcTan2(beta, alpha);
Rx(2.0 * theta, q);
}
# -
# *Can't come up with a solution? See the explained solution in the [Single-Qubit Gates Workbook](./Workbook_SingleQubitGates.ipynb#Exercise-6:-Preparing-a-rotated-state).*
# ### <span style="color:blue">Exercise 7</span>**: Preparing an arbitrary state
#
# **Inputs:**
#
# 1. A non-negative real number $\alpha$.
# 2. A non-negative real number $\beta = \sqrt{1 - \alpha^2}$.
# 3. A real number $\theta$.
# 4. A qubit in state $|0\rangle$.
#
# **Goal:** Transform the qubit into state $\alpha|0\rangle + e^{i\theta}\beta|1\rangle$.
#
# > Since only the relative amplitudes and relative phase have any physical meaning, this allows us to prepare any single-qubit quantum state we want to.
# +
%kata T7_PrepareArbitraryState
open Microsoft.Quantum.Math;
operation PrepareArbitraryState (alpha : Double, beta : Double, theta : Double, q : Qubit) : Unit is Adj+Ctl {
// PrepareRotatedState would get us alpha|0> - i * beta|1>
// So can I leave alpha untouched and turn i * beta into e^(i * theta) * beta?
// Applying S would give me alpha|0> + beta|1>
// Then apply R1(theta)
PrepareRotatedState(alpha, beta, q);
S(q);
R1(theta, q);
}
# -
# *Can't come up with a solution? See the explained solution in the [Single-Qubit Gates Workbook](./Workbook_SingleQubitGates.ipynb#Exercise-7**:-Preparing-an-arbitrary-state).*
# ## Conclusion
#
# Congratulations! You have learned enough to try solving the first part of the [Basic Gates kata](../../BasicGates/BasicGates.ipynb).
# When you are done with that, you can continue to the next tutorials in the series to learn about the [multi-qubit systems](../MultiQubitSystems/MultiQubitSystems.ipynb) and the [multi-qubit gates](../MultiQubitGates/MultiQubitGates.ipynb).
| tutorials/SingleQubitGates/SingleQubitGates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Demo target data generator
#
# This notebook shows how to use the target data generator to supply training data to use when fitting the PDF model. The `target_data_generator` implemented in `fit_target_data_model.py` yields an infinite stream of target data, by randomly generating new arrays of $p_{50\%}$ and $\log_{10}M_{\rm p},$ and computing the mean and variance of the input concentrations across time.
# %matplotlib inline
import matplotlib.cm as cm
from time import time
from matplotlib import lines as mlines
# ### Load the simulation data that includes best-fitting diffprof parameters
import os
from astropy.table import Table
diffprof_drn = "/Users/aphearin/work/DATA/diffprof_data"
mdpl2 = Table.read(os.path.join(diffprof_drn, "MDPL2_halo_table.hdf5"))
bpl = Table.read(os.path.join(diffprof_drn, "BPL_halo_table.hdf5"))
print(bpl.keys())
# ### Calculate ${\rm conc}(t_{\rm fit})$ based on the diffprof parameters
# +
from diffprof.nfw_evolution import lgc_vs_lgt
lgc_vs_lgt_vmap = jjit(jvmap(lgc_vs_lgt, in_axes=(None, 0, 0, 0, 0)))
N_T = 25
TARR_FIT = np.linspace(2, 13.8, N_T)
lgconc_history_bpl = np.array(lgc_vs_lgt_vmap(
np.log10(TARR_FIT), bpl["conc_lgtc"], bpl["conc_k"], bpl["conc_beta_early"], bpl["conc_beta_late"]))
lgconc_history_mdpl2 = np.array(lgc_vs_lgt_vmap(
np.log10(TARR_FIT), mdpl2["conc_lgtc"], mdpl2["conc_k"], mdpl2["conc_beta_early"], mdpl2["conc_beta_late"]))
# -
# ### Calculate downsampling masks
# +
from diffprof.latin_hypercube_sampler import get_scipy_kdtree, retrieve_lh_sample_indices
tree_bpl = get_scipy_kdtree(bpl['logmp'])
tree_mdpl2 = get_scipy_kdtree(mdpl2['logmp'])
indx_bpl = retrieve_lh_sample_indices(tree_bpl, 11.35, 13.65, 1, 100_000)
indx_mdpl2 = retrieve_lh_sample_indices(tree_mdpl2, 13.4, 14.6, 1, 100_000)
fig, ax = plt.subplots(1, 1)
__=ax.hist(bpl['logmp'][indx_bpl], bins=30, density=True, alpha=0.7)
__=ax.hist(mdpl2['logmp'][indx_mdpl2], bins=30, density=True, alpha=0.7)
# -
# ### Calculate the target data generator for this downsample of halos
#
#
# +
from diffprof.get_target_simdata import target_data_generator
N_MH_TARGETS, N_P_TARGETS = 20, 15
args = (bpl['logmp'][indx_bpl],
mdpl2['logmp'][indx_mdpl2],
lgconc_history_bpl[indx_bpl],
lgconc_history_mdpl2[indx_mdpl2],
bpl['p_tform_50'][indx_bpl],
mdpl2['p_tform_50'][indx_mdpl2],
N_MH_TARGETS,
N_P_TARGETS
)
gen = target_data_generator(*args)
# -
# ### Plot average concentration vs. mass
# +
target_data = next(gen)
lgmhalo_targets, p50_targets = target_data[0:2]
lgc_mean_targets_lgm0, lgc_std_targets_lgm0 = target_data[2:4]
lgc_mean_targets_lgm0_p50, lgc_std_targets_lgm0_p50 = target_data[4:]
lgm_colors=cm.coolwarm(np.linspace(1,0,N_MH_TARGETS))
fig, ax = plt.subplots(1, 1)
xlim = ax.set_xlim(0.5, 13.9)
xlabel = ax.set_xlabel(r'${\rm cosmic\ time\ [Gyr]}$')
ylabel = ax.set_ylabel(r'$\langle{\rm conc}(t)\ \vert\ M_{\odot},\ p_{50\%}\rangle$')
for im in range(N_MH_TARGETS):
__=ax.plot(TARR_FIT, lgc_mean_targets_lgm0[im, :], color=lgm_colors[im])
red_line=mlines.Line2D([],[],ls='-',c=mred,label=r'$M_{\rm halo}=10^{14.5}M_{\odot}$')
blue_line=mlines.Line2D([],[],ls='-',c=mblue,label=r'$M_{\rm halo}=10^{11.5}M_{\odot}$')
leg=ax.legend(handles=[red_line, blue_line])
fig.savefig('mean_lgm0_targets.png', bbox_extra_artists=[xlabel, ylabel], bbox_inches='tight', dpi=200)
# +
lgm_colors=cm.coolwarm(np.linspace(1,0,N_MH_TARGETS))
fig, ax = plt.subplots(1, 1)
xlim = ax.set_xlim(0.5, 13.9)
xlabel = ax.set_xlabel(r'${\rm cosmic\ time\ [Gyr]}$')
ylabel = ax.set_ylabel(r'$\sigma({\rm conc}(t)\ \vert\ M_{\odot},\ p_{50\%})$')
for im in range(N_MH_TARGETS):
__=ax.plot(TARR_FIT, lgc_std_targets_lgm0[im, :], color=lgm_colors[im])
red_line=mlines.Line2D([],[],ls='-',c=mred,label=r'$M_{\rm halo}=10^{14.5}M_{\odot}$')
blue_line=mlines.Line2D([],[],ls='-',c=mblue,label=r'$M_{\rm halo}=10^{11.5}M_{\odot}$')
leg=ax.legend(handles=[red_line, blue_line])
fig.savefig('std_lgm0_targets.png', bbox_extra_artists=[xlabel, ylabel], bbox_inches='tight', dpi=200)
# -
# +
p50_colors=cm.coolwarm(np.linspace(1, 0, N_P_TARGETS))
fig, ((ax0, ax1), (ax2, ax3)) = plt.subplots(2, 2, sharex=True, sharey=True, figsize=(10, 8))
# fig.subplots_adjust(wspace=0, hspace=0)
xlim = ax0.set_xlim(0.5, 13.9)
axes = ax0, ax1, ax2, ax3
for ax in ax2, ax3:
xlabel = ax.set_xlabel(r'${\rm cosmic\ time\ [Gyr]}$')
for ax in ax0, ax2:
ylabel = ax.set_ylabel(r'$\langle{\rm conc}(t)\ \vert\ M_{\odot},\ p_{50\%}\rangle$')
red_line=mlines.Line2D([],[],ls='-',c=mred,label=r'$p_{50\%}=0$')
blue_line=mlines.Line2D([],[],ls='-',c=mblue,label=r'$p_{50\%}=1$')
for ax, lgm in zip(axes, (11.5, 12.5, 13.5, 14.5)):
im = np.argmin(np.abs(lgmhalo_targets - lgm))
for ip, p50 in enumerate(p50_targets):
__=ax.plot(TARR_FIT, lgc_mean_targets_lgm0_p50[im, ip, :], color=p50_colors[ip])
leg=ax.legend(handles=[red_line, blue_line])
titles = (r'$M_{\rm halo}=10^{11.5}M_{\odot}$',r'$M_{\rm halo}=10^{12.5}M_{\odot}$',
r'$M_{\rm halo}=10^{13.5}M_{\odot}$', r'$M_{\rm halo}=10^{14.5}M_{\odot}$')
for ax, title in zip(axes, titles):
__=ax.set_title(title)
fig.savefig('mean_p50_targets.png', bbox_extra_artists=[xlabel, ylabel], bbox_inches='tight', dpi=200)
# -
# +
start = time()
target_data = next(gen)
lgmhalo_targets, p50_targets = target_data[0:2]
lgc_mean_targets_lgm0, lgc_std_targets_lgm0 = target_data[2:4]
lgc_mean_targets_lgm0_p50, lgc_std_targets_lgm0_p50 = target_data[4:]
end = time()
print("Runtime to generate target data = {0:.1f} seconds".format(end-start))
p50_colors=cm.coolwarm(np.linspace(1, 0, N_P_TARGETS))
fig, ((ax0, ax1), (ax2, ax3)) = plt.subplots(2, 2, sharex=True, sharey=True, figsize=(10, 8))
xlim = ax0.set_xlim(0.5, 13.9)
ylim = ax0.set_ylim(0, 0.2)
axes = ax0, ax1, ax2, ax3
red_line=mlines.Line2D([],[],ls='-',c=mred,label=r'$p_{50\%}=0$')
blue_line=mlines.Line2D([],[],ls='-',c=mblue,label=r'$p_{50\%}=1$')
for ax, lgm in zip(axes, (11.5, 12.5, 13.5, 14.5)):
im = np.argmin(np.abs(lgmhalo_targets - lgm))
for ip, p50 in enumerate(p50_targets):
__=ax.plot(TARR_FIT, lgc_std_targets_lgm0_p50[im, ip, :], color=p50_colors[ip])
leg=ax.legend(handles=[red_line, blue_line])
for ax in ax2, ax3:
xlabel = ax.set_xlabel(r'${\rm cosmic\ time\ [Gyr]}$')
for ax in ax0, ax2:
ylabel = ax.set_ylabel(r'$\sigma({\rm conc}(t)\ \vert\ M_{\odot},\ p_{50\%})$')
titles = (r'$M_{\rm halo}=10^{11.5}M_{\odot}$',r'$M_{\rm halo}=10^{12.5}M_{\odot}$',
r'$M_{\rm halo}=10^{13.5}M_{\odot}$', r'$M_{\rm halo}=10^{14.5}M_{\odot}$')
for ax, title in zip(axes, titles):
__=ax.set_title(title)
fig.savefig('std_p50_targets.png', bbox_extra_artists=[xlabel, ylabel], bbox_inches='tight', dpi=200)
# -
# ## Now build a simple toy model for $\sigma({\rm conc}(t)\ \vert\ M_{\odot}, p_{50\%})$
#
# The goal is simply to roughly capture the curves in the previous plot using a simple, smooth parametric model.
#
# When I inspect those curves, what I see are three basic things:
# 1. There's not a lot of time-dependence, especially for $t\gtrsim2.5\ {\rm Gyr},$ so we can probably assume time-independent scatter.
# 2. For halos with $p_{50\%}$ of all mass, the scatter looks about constant of 0.05 dex
# 3. For lower-mass halos, there is a $p_{50\%}$-dependent _width_ in the scatter that is roughly 0.04 dex, and this width practically vanishes in cluster-mass halos.
#
# So let's first focus on (3) and build a simple model for the width function, $W(m),$ where I'm writing $m\equiv\log_{10}M_{\rm h}$ for notational convenience. To get us started, let's try out a sigmoid function for the shape of $W(m).$ To make the plot below, I just fiddled for a minute with the 4 sigmoid parameters until I got a curve that looks about right.
# +
@jjit
def _sigmoid(x, x0, k, ylo, yhi):
height_diff = yhi - ylo
return ylo + height_diff / (1 + jnp.exp(-k * (x - x0)))
fig, ax = plt.subplots(1, 1)
ylim = ax.set_ylim(0, 0.1)
lgmarr = np.linspace(10, 15.5, 500)
delta_scatter_p50 = _sigmoid(lgmarr, 13, 1, 0.05, 0)
__=ax.plot(lgmarr, delta_scatter_p50)
xlabel = ax.set_xlabel(r'$\log_{10}M_{\rm halo}$')
xlabel = ax.set_ylabel(r'${\rm scatter\ width}$')
# -
# Ok, now that we have our rough sigmoid-type behavior to the width of the scatter, let's port that into a callable function.
@jjit
def _scatter_p50_width_vs_lgmhalo(
lgmh, p50_sig_width_x0=13, p50_sig_width_k=1, p50_sig_width_ylo=0.05, p50_sig_width_yhi=0):
return _sigmoid(lgmh, p50_sig_width_x0, p50_sig_width_k, p50_sig_width_ylo, p50_sig_width_yhi)
# Now that we have our model for the width, then we can use this model together with observation (1) to build a simple model for the value of the scatter, $\sigma(p, m).$ So we want to write down some function that returns $\sigma(p\rightarrow0, m)\approx0.05,$ and also $\sigma(p\rightarrow1, m)\approx0.05+W(m),$ where the $W(m)$ is the width function we calibrated above. To implement such a function for $\sigma(p, m),$ what functional form should we use? Let's try...you guessed it, a sigmoid function!
#
# The last ingredient we need is just to know how to set our value of $k$ for how rapidly the sigmoid function will transition the scatter from its lower-bound value of $\sigma\approx0.05$ to its upper value of $\sigma\approx0.05+W(m).$ In the figure below, I show that $k\approx5$ is about right.
# +
fig, ax = plt.subplots(1, 1)
ylim = ax.set_ylim(0, 0.2)
pp = np.linspace(0, 1, 500)
sig_lo, sig_hi = 0.05, 0.1
k_p = 5
__=ax.plot(pp, _sigmoid(pp, 0.5, k_p, sig_lo, sig_hi))
# -
@jjit
def _scatter_vs_p50_and_lgmhalo(
lgmh, p, p_lo=0.05, p50_sig_width_x0=13, p50_sig_width_k=1,
p50_sig_width_ylo=0.05, p50_sig_width_yhi=0):
width = _scatter_p50_width_vs_lgmhalo(
lgmh, p50_sig_width_x0, p50_sig_width_k,
p50_sig_width_ylo, p50_sig_width_yhi)
return _sigmoid(p, 0.5, 5, p_lo, p_lo+width)
# Now let's make our 4-panel plot of $\sigma(p, m)$ and overplot our new simple model.
# +
start = time()
target_data = next(gen)
lgmhalo_targets, p50_targets = target_data[0:2]
lgc_mean_targets_lgm0, lgc_std_targets_lgm0 = target_data[2:4]
lgc_mean_targets_lgm0_p50, lgc_std_targets_lgm0_p50 = target_data[4:]
end = time()
print("Runtime to generate target data = {0:.1f} seconds".format(end-start))
p50_colors=cm.coolwarm(np.linspace(1, 0, N_P_TARGETS))
fig, ((ax0, ax1), (ax2, ax3)) = plt.subplots(2, 2, sharex=True, sharey=True, figsize=(10, 8))
xlim = ax0.set_xlim(0.5, 13.9)
ylim = ax0.set_ylim(0, 0.15)
axes = ax0, ax1, ax2, ax3
red_line=mlines.Line2D([],[],ls='-',c=mred,label=r'$p_{50\%}=0$')
blue_line=mlines.Line2D([],[],ls='-',c=mblue,label=r'$p_{50\%}=1$')
for ax, lgm in zip(axes, (11.5, 12.5, 13.5, 14.5)):
im = np.argmin(np.abs(lgmhalo_targets - lgm))
for ip, p50 in enumerate(p50_targets):
__=ax.plot(TARR_FIT, lgc_std_targets_lgm0_p50[im, ip, :], color=p50_colors[ip])
for ax in ax2, ax3:
xlabel = ax.set_xlabel(r'${\rm cosmic\ time\ [Gyr]}$')
for ax in ax0, ax2:
ylabel = ax.set_ylabel(r'$\sigma({\rm conc}(t)\ \vert\ M_{\odot},\ p_{50\%})$')
titles = (r'$M_{\rm halo}=10^{11.5}M_{\odot}$',r'$M_{\rm halo}=10^{12.5}M_{\odot}$',
r'$M_{\rm halo}=10^{13.5}M_{\odot}$', r'$M_{\rm halo}=10^{14.5}M_{\odot}$')
for ax, title in zip(axes, titles):
__=ax.set_title(title)
tarr_plot = np.linspace(-100, 100, 5000)
for ax, lgm in zip(axes, (11.5, 12.5, 13.5, 14.5)):
for ip, p50 in enumerate(p50_targets):
scatter = _scatter_vs_p50_and_lgmhalo(lgm, p50)
__=ax.plot(tarr_plot, np.zeros_like(tarr_plot)+scatter, '--', color=p50_colors[ip])
from matplotlib import lines as mlines
red_line=mlines.Line2D([],[],ls='-',c=mred,label=r'$p_{50\%}=0$')
blue_line=mlines.Line2D([],[],ls='-',c=mblue,label=r'$p_{50\%}=1$')
solid_line=mlines.Line2D([],[],ls='-',c='gray',label=r'${\rm simulation}$')
dashed_line=mlines.Line2D([],[],ls='--',c='gray',label=r'${\rm target\ data\ model}$')
leg=ax0.legend(handles=[blue_line, red_line])
leg=ax1.legend(handles=[solid_line, dashed_line])
fig.savefig('std_p50_target_data_model_demo.png', bbox_extra_artists=[xlabel, ylabel], bbox_inches='tight', dpi=200)
# -
# ### Not bad!
#
# All the basic trends are captured, and our function $\sigma(m, p, t)$ is relatively simple. Looking at this last plot, we could probably improve on it slightly if we had set our lower bound to something more like $0.04,$ and maybe the width should be slightly larger in lower-mass halos as well. So the final step towards building this target data model is just to optimize the parameters we used to define $\sigma(m, p, t).$ To do that, these functions should be ported into a module, you'll need to implement them such that the relevant parameters are not just hard-coded default values, and instead are allowed to vary freely. Then you'll need to define an MSE loss function that calculates the difference between some input targets and the model predictions, and this is the MSE loss that you'll use with the usual `jax_adam_wrapper` function. For the targets, you can get them using the same `target_data_generator` used to make this plot. And just like you have been doing with the actual `diffprof` model, you should write a simple for loop where you generate some target data, to gradient descent on the parameters for ~100 steps, then generate a new variation on the target data, burn for another ~100 steps, and proceed until your model parameters settle into something that gives a good description for what we see in this plot. Probably the best-fit parameters will not be too far off from the default values in this notebook.
#
# Once that is done, then you can return to optimizing the actual `diffprof` model, and instead of using the `target_data_generator`, you can use the newly-developed target data model.
| notebooks/demo_target_data_generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/LuisFelipeUrena/DS-Unit-3-Sprint-1-Software-Engineering/blob/master/Assignments/DS14_TL_LS_LUIS_URENA_UNIT3_1-1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="B9MPm1vMjVrb" colab_type="code" colab={}
import pandas as pd
cali = '/content/sample_data/california_housing_train.csv'
df = pd.read_csv(cali)
# + id="Ywr8OI2AjjeS" colab_type="code" outputId="157a8420-fca1-4c78-b465-b2f12fc67113" colab={"base_uri": "https://localhost:8080/", "height": 136}
df.head(3
)
# + id="3Nq7aO5uj0MT" colab_type="code" colab={}
values = pd.Series(df.isnull().sum())
# + id="jxOHwucyj5TS" colab_type="code" outputId="631ee155-bc1c-4d04-84bf-adb1fa08891f" colab={"base_uri": "https://localhost:8080/", "height": 185}
values
# + id="wN0eTA5EkGO4" colab_type="code" outputId="bdb6c19b-770a-42a7-8eea-90cb2590baf1" colab={"base_uri": "https://localhost:8080/", "height": 168}
for i in values:
print(df.columns[0])
# + id="cNwESjRWkafR" colab_type="code" outputId="cb1189d3-9cf6-4cb9-ac15-76a40a3f9984" colab={"base_uri": "https://localhost:8080/", "height": 235}
print('Null Values in each Column')
print('---------------------------')
print(values)
print('---------------------------')
# + id="Oj2VUZysmjyM" colab_type="code" colab={}
# + id="pSOLxr9snCPx" colab_type="code" colab={}
# + id="-pYMm14msTPb" colab_type="code" colab={}
result = df.astype('object').describe()
# + id="Rn2oSVp0vEBF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="874cc2a6-2a2a-41ed-d6c4-5e0a2f080384"
print(result)
# + id="IjQmm_wevgqR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="8670ce73-b50f-410c-9dde-206f10b43e03"
# !pip install -i https://test.pypi.org/simple/ lambdata-luife
# + id="VTqLTUXcNZpX" colab_type="code" colab={}
from my_lamdata.assign import null_val
# + id="i8nOCeXdNilA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="b8261b8c-31b4-44c5-8155-d536d5de98a0"
null_val(df)
# + id="MALy547VOXpG" colab_type="code" colab={}
from my_lamdata.assign import cat_variables
# + id="qQlxxkBrOj9q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 166} outputId="b102767b-eabc-42e4-c42b-046dca83208c"
cat_variables(df)
# + id="g3H_EtvXPWRi" colab_type="code" colab={}
| Assignments/DS14_TL_LS_LUIS_URENA_UNIT3_1-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Imports
# +
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import json
import copy
from collections import defaultdict
import os
import sys
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
module_path = os.path.abspath(os.path.join('../..'))
if module_path not in sys.path:
sys.path.append(module_path)
from src import runner
from src import gen_spectra
# -
# # Correct sequence fall off detection
# We have a property in the program that lets us pass in a "truth" set. This truth set we will pull from Delong lab and will primarily be used for debugging our hybrids. The steps we need to take:
# 1. Load in SpectrumMill results and create a "truth" set where each entry in this json file has the form:
# ```json
# {
# spectrum_id: {
# "sequence": str,
# "hybrid": bool,
# "parent": str
# }
# }
# ```
# 2. Run hyped search with the "truth_set" param set to the file generated in step 1
# 3. Load in the file created (output_dir + 'fall_off.json')
# 4. Run all analysis
# ## Constants
# +
spec_file = '/Users/zacharymcgrath/Desktop/nod2 data/filteredSpec/'
db_file = '/Users/zacharymcgrath/Desktop/nod2 data/filteredNOD2.fasta'
output_dir = '/Users/zacharymcgrath/Desktop/Experiment output/fall_off/'
specmil_truth_set = '/Users/zacharymcgrath/Downloads/NOD2_E3_results.ssv'
minPep = 3
maxPep = 30
tolerance = 20
relative_abundance_filter = 0.0
precursor_tolerance = 10
peak_filter = 25
verbose = True
# -
# ## 1. Load in SpectrumMill and create truth file
# first load in the results
specmil_results = pd.read_csv(specmil_truth_set, sep=';')
specmil_results.head(5)
def get_hybrid_seq(peptide: str, protein: str) -> str:
'''
From a hybrid protein that looks like
ABCDE-FGHI
and a peptide that looks like
DEFGH
extract
DE-FGH
Inputs:
peptide: (str) the desired subseq
protein: (str) the full string with the hybrid character
Outputs:
new peptide string
'''
# copy the protein
prot_cp = copy.deepcopy(protein)
# find the subseq
peptide_idx = protein.replace('-', '').replace('(', '').replace(')', '').index(peptide)
# get that stretch of prot
add_on = len(peptide) + (1 if '-' in protein else 2)
return protein[peptide_idx:peptide_idx+add_on]
# +
# json to store results in
json_truth_set = {}
# go through every entry in the database, find out if its a hybrid and get the sequence, ided by the filename
for idx, row in specmil_results.iterrows():
# all the ids seem to have .pkl at the end of them so add that
key = row['filename'] + '.pkl'
hybrid = 'HYBRID' in row['entry_name']
seq = row['sequence'] if not hybrid else get_hybrid_seq(row['sequence'], row['entry_name'])
json_truth_set[key] = {
'hybrid': hybrid,
'sequence': seq,
'parent': row['entry_name']
}
full_truth_path = output_dir + 'specmil_truth_set.json'
json.dump(json_truth_set, open(full_truth_path, 'w'))
# -
# ## 2. Run hypedsearch with the truth set
# +
truth_run_params = {
'spectra_folder': spec_file,
'database_file': db_file,
'output_dir': output_dir,
'min_peptide_len': minPep,
'max_peptide_len': maxPep,
'tolerance': tolerance,
'precursor_tolerance': precursor_tolerance,
'peak_filter': peak_filter,
'relative_abundance_filter': relative_abundance_filter,
'digest': 'trypsin',
'missed_cleavages': 2,
'verbose': verbose,
'DEBUG': False,
'cores': 16,
'truth_set': full_truth_path
}
runner.run(truth_run_params)
# -
# ## 3. Load in the fall off results
fall_off_results = json.load(open(output_dir + 'fall_off.json'))
len(fall_off_results)
# ## 4. Run all analysis
# ### Plot the raw results
# +
# first load them into dictionaries by {type: [seq]}
typed_fall_off = defaultdict(list)
for _id, entry in fall_off_results.items():
typed_fall_off[entry['fall_off_operation']].append((_id, entry))
# +
# bar graph it
plt.figure(figsize=(15, 10))
# get parallel x and y lists
x = []
y = []
xlabels = []
for i, (op, entries) in enumerate(typed_fall_off.items()):
x.append(i)
y.append(len(entries))
xlabels.append(op)
plt.bar(x, y, tick_label=xlabels)
# -
# ### Just hybrid results
# +
hybrid_typed_fall_off = defaultdict(list)
nonhybrid_typed_fall_off = defaultdict(list)
for _id, entry in fall_off_results.items():
if entry['hybrid']:
hybrid_typed_fall_off[entry['fall_off_operation']].append((_id, entry))
else:
nonhybrid_typed_fall_off[entry['fall_off_operation']].append((_id, entry))
# +
# bar graph it
plt.figure(figsize=(15, 10))
# get parallel x and y lists
x = []
y = []
xlabels = []
for i, (op, entries) in enumerate(hybrid_typed_fall_off.items()):
x.append(i)
y.append(len(entries))
xlabels.append(op)
plt.bar(x, y, tick_label=xlabels)
plt.title('Hybrid fall off positions')
# +
# bar graph it
plt.figure(figsize=(15, 10))
# get parallel x and y lists
x = []
y = []
xlabels = []
for i, (op, entries) in enumerate(nonhybrid_typed_fall_off.items()):
x.append(i)
y.append(len(entries))
xlabels.append(op)
plt.bar(x, y, tick_label=xlabels)
plt.title('Non hybrid fall off positions')
# -
# ### Hybrid analysis
precursor_fall_offs = [x for x in hybrid_typed_fall_off['precursor_filling']]
for pfo in precursor_fall_offs:
t_seq = pfo[1]['truth_sequence']
print(f'Truth sequence: {t_seq}')
print('Before\n===========================================')
for x in pfo[1]['meta_data']['sequences_before_precursor_filling']:
if x[0][:2] == t_seq[:2] and x[0][-2:] == t_seq[-2:]:
print(x)
print('After\n===========================================')
for x in pfo[1]['meta_data']['sequences_after_precursor_filling']:
if x[0][:1] == t_seq[:1] and x[0][-1:] == t_seq[-1:]:
print(x)
print()
top_x_filtering = [x for x in hybrid_typed_fall_off['top_x_filtering']]
# +
def is_close(truth, tried, ion, close_dist=2):
if len(tried) > len(truth):
return False
if ion == 'b':
return truth[:close_dist].replace('I', 'B').replace('L', 'B') == tried[:close_dist].replace('I', 'B').replace('L', 'B')
else:
return truth[-close_dist:].replace('I', 'B').replace('L', 'B') == tried[-close_dist:].replace('I', 'B').replace('L', 'B')
for _id, txf in top_x_filtering:
t_seq = txf['truth_sequence']
print(f'Truth sequence: {t_seq}')
print('Kept b hits close to correct half\n============================')
for x in txf['meta_data']['top_x_b_hits']:
if is_close(t_seq, x, 'b', 3):
print(x)
print('Kept y hits close to correct half\n============================')
for x in txf['meta_data']['top_x_y_hits']:
if is_close(t_seq, x, 'y', 3):
print(x)
print('Lost b hits close to correct half\n============================')
for x in txf['meta_data']['excluded_b_hits']:
if is_close(t_seq, x, 'b', 3):
print(x)
print('Lost y hits close to correct half\n============================')
for x in txf['meta_data']['excluded_y_hits']:
if is_close(t_seq, x, 'y', 3):
print(x)
print()
# -
# ### Non hybrid analysis
# #### Precursor filling
# get the precursor filling subjects
nh_precursor_fall_off = [x for x in nonhybrid_typed_fall_off['precursor_filling']]
# +
# go through every entry and find the sequences that were closest to the real sequence as possible
# closeness is:
# 1. Having a lot of the right amino acids
# 2. Not have too many amino acids or too few
# Prioritize 1, rank by 2
def closeness(t_seq, trying, observed_precursor, observed_precursor_charge) -> (int, int):
from_right = 0
from_left = 0
prec_distance = abs(gen_spectra.get_precursor(trying, observed_precursor_charge) - observed_precursor)
# first check if they ARE the same
if t_seq == trying:
return (len(t_seq), 0, prec_distance)
# go from left to right
i = 0
while i < len(t_seq) and i < len(trying) and t_seq[i] == trying[i]:
i += 1
from_left += 1
# now right to left
i = -1
while abs(i) < len(t_seq) + 1 and abs(i) < len(trying) + 1 and t_seq[i] == trying[i]:
i -= 1
from_right += 1
return (from_left + from_right, abs(len(t_seq) - len(trying)), prec_distance)
for _id, nhpfo in nh_precursor_fall_off:
# get the real sequence
t_seq = nhpfo['truth_sequence']
# keep track of the best of the close ones
overlapped_idxed_close_hits = defaultdict(list)
observed_prec = nhpfo['meta_data']['observed_precursor_mass']
observed_prec_charge = nhpfo['meta_data']['observed_percursor_charge']
for non_hyb, hyb in nhpfo['meta_data']['sequences_before_precursor_filling']:
overlap, dist, prec_dist = closeness(t_seq, non_hyb, observed_prec, observed_prec_charge)
overlapped_idxed_close_hits[overlap].append((non_hyb, hyb, dist, prec_dist))
# now get the hightest key
best_key = max(list(overlapped_idxed_close_hits.keys()))
# show the best results sorted by their distance
print(f'({_id}) Hits with the most overlap ({best_key} AAs) for sequence {t_seq} with allowed gap {nhpfo["meta_data"]["allowed_gap"]}')
print('=====================================================================================')
for seq, hyb_seq, dist, prec_distance in sorted(overlapped_idxed_close_hits[best_key], key=lambda x: x[2]):
print(f'{seq} \t {dist} \t {prec_distance} \t {hyb_seq}')
print()
# -
# #### Taking the top n alignments
top_n = nh_precursor_fall_off = [x for x in nonhybrid_typed_fall_off['taking_top_n_alignments']]
for _id, top_n_spec in top_n:
t_seq = top_n_spec['truth_sequence']
print(f'Top n alignments for sequence {t_seq}')
print('============================================================')
print('sequence \t b score \t y score \t total score \t precursor distance \t hybrid')
print('------------------------------------------------------------')
[print(f'{x["sequence"]} \t {x["b_score"]} \t {x["y_score"]} \t {x["total_score"]} \t {x["precursor_distance"]} \t {"hybrid_sequence" in x}') for x in top_n_spec['meta_data']['top_n']]
print('------------------------------------------------------------')
print('Missed alignments for this sequence')
print('------------------------------------------------------------')
[print(f'{x["sequence"]} \t {x["b_score"]} \t {x["y_score"]} \t {x["total_score"]} \t {x["precursor_distance"]} \t {"hybrid_sequence" in x}') for x in top_n_spec['meta_data']['not_top_n'][:10]]
print('------------------------------------------------------------')
all_alignments = [x for x in top_n_spec['meta_data']['top_n']] + [x for x in top_n_spec['meta_data']['not_top_n']]
num_hyb = len([0 for x in all_alignments if 'hybrid_sequence' in x])
p_hyb = int(100 * (float(num_hyb) / float(len(all_alignments))))
scores = [x['total_score'] for x in all_alignments]
avg_score = np.mean(scores)
min_score = min(scores)
print(f'Stats: \t Number alignments: {len(all_alignments)} \t % hybrid: {p_hyb} \t Average score: {avg_score} \t min score: {min_score}')
print()
# #### first alignment round
first_as = [x for x in nonhybrid_typed_fall_off['first_alignment_round']]
# +
def most_aa_overlap(truth, trying):
overlap = 0
if truth == trying:
return len(truth)
i = 0
while i < len(truth) and i < len(trying) and trying[i] == truth[i]:
i += 1
overlap += 1
i = -1
while i > -1 * min(len(trying), len(truth)) and truth[i] == trying[i]:
i -= 1
overlap += 1
return overlap - max(0, len(trying) - len(truth))
for _id, fa in first_as:
t_seq = fa['truth_sequence']
most_overlap = defaultdict(list)
for non_hyb, hyb in fa['meta_data']['alignments']:
most_overlap[most_aa_overlap(t_seq, non_hyb)].append((non_hyb, hyb))
most_aas = max(list(most_overlap.keys()))
closest_matches = most_overlap[most_aas]
print(f'({_id}) Closest alignments to sequence {t_seq} at overlap {most_aas}')
print('=================================================================')
[print(f'{non_hyb} \t {hyb}') for non_hyb, hyb in closest_matches]
print()
# -
ss = ['SSEPTQGSYKVVIRT-PEGATE', 'SSEELLV-EAGGAFGKREKAEE', 'SSEELLVAE-GGAFGKREKAEE', 'SSEEGKLF-PEGCVVAAVAARSE', 'SSLDSGVPKRFSGS-TTAANPKE', 'STED-YQKYKFMELNLAQK', 'SVRSGTPHVAEAA-EELDPENK', 'SVRSGTPHVAEAA-EEDLPENK', 'SSLDSGVPKRFSG-LKDNATQE', 'SYKALLDSQSIPT-LECNEPK', 'SEES-YQKYKFMELNLAQK', 'SEESY-QKYKFMELNLAQK', 'SEESYQKYKFMELNLAQK', 'SEES-YQKYKFMELNLAQK', 'SEES-YQKYKFMELNLAQK', 'SEES-YQKYKFMELNLAQK', 'SEES-YQKYKFMELNLAQK', 'SEES-YQKYKFMELNLAQK', 'SEESY-QKYKFMELNLAQK', 'SEESYQKYKFMELNLAQK', 'SEES-YQKYKFMELNLAQK', 'SEES-YQKYKFMELNLAQK', 'SEES-YQKYKFMELNLAQK', 'SEES-YQKYKFMELNLAQK', 'SEESY-QKYKFMELNLAQK', 'SEESYQKYKFMELNLAQK', 'AGQVRLTYSTGE-REGILQEE', 'SSLDSGVPKRFSGSRS-PNQTE', 'QQYGVRGYPTIKF-ASYQTE', 'SSEPTQGSYKVVIRTE-PQTE', 'SSEPTQGSYKVVIRTE-PQTE', 'SESE-YQKYKFMELNLAQK', 'QQSLFQRLDF-EELDPENK', 'QQSLFQRLDF-EEDLPENK', 'STRIIYGGSVTGATC-GAPGNKPE', 'QQELPSLSVGPSLH-VATDQTE', 'SSLEKSYE-LPEATGLSPLSVE', 'SSLEKSYEL-PEATGLSPLSVE', 'SSLEKSYELPEATGLSPLSVE', 'SSLEKSYELP-EATGLSPLSVE', 'SSLEKSYELPEATGLSPLSVE', 'SSLEKSYELPEATGLSPLSVE', 'SSLEKSYE-LPEATGLSPLSVE', 'SSLEKSYEL-PEATGLSPLSVE', 'SSLEKSYELPEATGLSPLSVE', 'SSLEKSYELP-EATGLSPLSVE', 'SSLEKSYELPEATGLSPLSVE', 'SSLEKSYELPEATGLSPLSVE', 'SYDIVLVKEESLE-SGIPQTE', 'SYDIVLVKEESLEV-ASPQTE', 'SYDIVLVKEESLEV-PASQTE', 'SLVSNYLQTQE-GEAKSPGEAK', 'SSEE-YQKYKFMELNLAQK', 'SSEE-YQKYKFMELNLAQK', 'SSEE-YQKYKFMELNLAQK', 'SSEE-YQKYKFMELNLAQK', 'SSEE-YQKYKFMELNLAQK', 'SSEE-YQKYKFMELNLAQK', 'SSEE-YQKYKFMELNLAQK', 'SSEE-YQKYKFMELNLAQK', 'SSEE-YQKYKFMELNLAQK', 'SSEE-YQKYKFMELNLAQK', 'SSEE-YQKYKFMELNLAQK', 'SSEELQE-LPALAIYESVDDK', 'SSLDSGVPKRFSG-SLIGGNASAE', 'SSLDSGVPKRFSGS-LIGGNASAE', 'SSLDSGVPKRFSGSLIGGNASAE', 'QQYLRQEV-EKKKGDEDDK', 'SSEERAAPF-EALSRTGRSRE', 'SSEPTQGSYKVVIRTE-PDDK', 'SSEPTQGSYKVVIRTE-PDDK', 'SYGDLGGPIITT-DAATAVEQEK', 'SSQVVLPAPTGIIHQ-SEDDDK', 'SEDT-YQKYKFMELNLAQK', 'SEDT-YQKYKFMELNLAQK', 'SSEPTQGSYKVV-REGILQEE', 'STIAIVSTFTCH-TVIVEGQTE', 'AGQVRLTYSTGESN-PKVAAGTE', 'QQSLFQRLDF-AEVPEAASAE', 'STCFRPACVKLGAGAG-RPSTQE', 'QQYGVRGYPTI-EELDPENK', 'QQYGVRGYPTI-EEDLPENK', 'STCFRPACVKLG-REGILQEE', 'SSEPTQGSYKVVIRTE-PAGTE', 'SSEELQE-PLYRLNTKAASAE', 'SETD-YQKYKFMELNLAQK', 'SSEEGKLF-PRATDLTARQTE', 'SSEERAAPF-RATDLTARQTE', 'STDE-YQKYKFMELNLAQK', 'SSIPSHPSQSVR-EELDPENK', 'SSIPSHPSQSVR-EEDLPENK', 'SYDIVLVKEES-AEVPEAASAE', 'SSARFRKVDVDE-PSRETAGE', 'SEVKTDVNKIEE-VYDPKNE', 'SSIPSHPSQSVRSVN-DANPEK', 'SSLEKSYELP-EREGILQEE', 'SSLEKSYELP-EREGILQEE', 'SYDIVLVKEES-EELDPENK', 'SYDIVLVKEES-EEDLPENK', 'SLVSNYLQTQE-REGILQEE', 'QQYGVRGYPTIKF-SAYGATE', 'QQYGVRGYPTI-AEVPEAASAE', 'QTDAKKGTITIQDTGIGMTQE', 'SSEPTQGSYKVVIRTE-PTGAE', 'SSIPSHPSQSVR-AEVPEAASAE', 'SYKALLDSQSIPTD-RPSTQE', 'STRIIYGGSVTGA-EELDPENK', 'STRIIYGGSVTGA-EEDLPENK', 'SSEERAAPF-VESKHKSDFGK', 'SSEPTQGSYKVV-GEAKSPGEAK', 'SEAAKVNTD-PALAIYESVDDK', 'SSQVVLPAPTGIIH-NESESAAE']
[x for x in ss if 'AGSIREAGGAFGKREKAEE' in x]
| testing framework/jupyter notebooks/.ipynb_checkpoints/Correct sequence fall off detection-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="aYjtnkWINc2Y"
# In this notebook we demonstrate a basic text classifier using [Textblob.](https://textblob.readthedocs.io/en/dev/) .
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="spAkPPa0LJme" outputId="2ad2c9cd-e5f4-4510-e1dd-621161c02032"
# Load the dataset.
import pandas as pd
df = pd.read_csv("Data/sts_gold_tweet.csv",error_bad_lines=False,delimiter=";")
print(df.columns)
# Make a list of all the tweets.
tweets_text_collection = list(df['tweet'])
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="mBPraZ2FLnX7" outputId="c719024a-b44a-48f7-dcc6-59923e0f7740"
# !pip install textblob
from textblob import TextBlob
for tweet_text in tweets_text_collection:
print(tweet_text)
analysis = TextBlob(tweet_text)
# Analyse the sentitment.
print(analysis.sentiment)
# Polarity is a value between [-1.0, 1.0] and tells how positive or negative the text is.
# Subjectivity is within the range [0.0, 1.0] where 0.0 is very objective and 1.0 is very subjective.
print("-"*20)
# + [markdown] colab_type="text" id="Hmq64mfYO0NQ"
# There are many more features in textblob you can refer the official documentation for them.
| Ch8/04_Sentiment_Analysis_Textblob.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# ## STATISTICS WORKSHOP
#
# __Version: March 2022__
# -
# __USING THE NOTEBOOK__
# The present notebook is composed of text and code cells. The former include the instructions for the activity and look just like regular text in a webpage. Cells that have "Answer:" at the beginning of them are also text cells. To write your answer just double click on them so the cursor appears and you can type your answer. When you are done click "shift" + "enter".
# The code cells look like gray squares with empty square brackets to their left ([ ]). To run the code inside a code cell you'll need to hover on the top left corner of the box, and when the empty square brackets change to a "play" sign just click on it (alternatively: click on the code cell and then click "shift" + "enter"), this will make the outcome of the code to appear underneath the cell.
#
# The following code cell will upload all the libraries and functions we'll need for the workshop. Please run it.
# +
# Loading Python libraries
import numpy as np
import pandas as pd
import scipy.stats as stats
import statsmodels.api as sm
import statsmodels.stats.multicomp as multi
from statsmodels.formula.api import ols
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
pd.options.display.float_format = '{:.3f}'.format
np.set_printoptions(precision=3, suppress=True)
# Statistics functions
def parammct(data=None, independent=None, dependent=None):
independent = str(independent)
dependent = str(dependent)
if input_check_numerical_categorical(data, independent, dependent):
return
parammct_df = pd.DataFrame()
for value in pd.unique(data[independent]):
mean = data[dependent][data[independent]==value].mean()
stdev = data[dependent][data[independent]==value].std()
n = data[dependent][data[independent]==value].count()
sdemean = stdev/np.sqrt(n)
ci = 1.96*sdemean
lowerboundci = mean-ci
upperboundci = mean+ci
parammct_df[value] = pd.Series([mean, stdev, n, sdemean, lowerboundci, upperboundci],
index = ['Mean','SD','n','SEM','Lower bound CI', 'Upper bound CI'])
return parammct_df
def non_parammct(data=None, independent=None, dependent=None):
independent = str(independent)
dependent = str(dependent)
if input_check_numerical_categorical(data, independent, dependent):
return
non_parammct_df = pd.DataFrame()
for value in pd.unique(data[independent]):
median = data[dependent][data[independent]==value].median()
minimum = data[dependent][data[independent]==value].quantile(0)
q25 = data[dependent][data[independent]==value].quantile(0.25)
q75 = data[dependent][data[independent]==value].quantile(0.75)
maximum = data[dependent][data[independent]==value].quantile(1)
non_parammct_df[value] = pd.Series([median, minimum, q25,q75, maximum],
index = ['Median', 'Minimum', 'Lower bound IQR', 'Upper bound IQR',
'Maximum'])
return non_parammct_df
def histograms(data=None, independent=None, dependent=None):
independent = str(independent)
dependent = str(dependent)
if input_check_numerical_categorical(data, independent, dependent):
return
for value in pd.unique(data[independent]):
sns.distplot(data[dependent][data[independent]==value], fit=stats.norm, kde=False)
plt.title(dependent + ' by ' + independent + '(' + str(value).lower() + ')',
fontweight='bold', fontsize=16)
plt.ylabel('Frequency', fontsize=14)
plt.xlabel(dependent, fontsize=14)
plt.show()
return
def t_test(data=None, independent=None, dependent=None):
pd.set_eng_float_format(accuracy=3, use_eng_prefix=False)
independent_groups = pd.unique(data[independent])
if len(independent_groups)>2:
print('There are more than 2 groups in the independent variable')
print('t-test is not the correct statistical test to run in that circumstance,')
print('consider running an ANOVA')
return
mct = parammct(data=data, independent=independent, dependent=dependent)
t_test_value, p_value = stats.ttest_ind(data[dependent][data[independent] == independent_groups[0]],
data[dependent][data[independent] == independent_groups[1]])
difference_mean = np.abs(mct.loc['Mean'][0] - mct.loc['Mean'][1])
pooled_sd = np.sqrt( ( ((mct.loc['n'][0]-1)*mct.loc['SD'][0]**2) + ((mct.loc['n'][1]-1)*mct.loc['SD'][1]**2) ) /
(mct.loc['n'][0] + mct.loc['n'][1] - 2) )
sedifference = pooled_sd * np.sqrt( (1/mct.loc['n'][0]) + (1/mct.loc['n'][1]) )
difference_mean_ci1 = difference_mean + (t_test_value * sedifference)
difference_mean_ci2 = difference_mean - (t_test_value * sedifference)
if difference_mean_ci1>difference_mean_ci2:
difference_mean_cilower = difference_mean_ci2
difference_mean_ciupper = difference_mean_ci1
else:
difference_mean_cilower = difference_mean_ci1
difference_mean_ciupper = difference_mean_ci2
cohend = difference_mean / pooled_sd
t_test_result= pd.Series ([difference_mean, sedifference, t_test_value, p_value,
difference_mean_cilower, difference_mean_ciupper, cohend],
index = ['Difference between means', 'SE difference', 't-test', 'p-value',
'Lower bound difference CI', 'Upper bound difference CI', 'Cohen\'s d'])
return t_test_result
def anova(data=None, independent=None, dependent=None):
pd.set_eng_float_format(accuracy=3, use_eng_prefix=False)
independent = str(independent)
dependent = str(dependent)
if input_check_numerical_categorical(data, independent, dependent):
return
formula = dependent + ' ~ ' + independent
model = ols(formula, data=data).fit()
aov_table = sm.stats.anova_lm(model, typ=2)
aov_table.rename(columns={'PR(>F)':'p'}, inplace=True)
aov_table['F'] = pd.Series([aov_table['F'][0], ''], index = [independent, 'Residual'])
aov_table['p'] = pd.Series([aov_table['p'][0], ''], index = [independent, 'Residual'])
eta_sq = aov_table['sum_sq'][0]/(aov_table['sum_sq'][0]+aov_table['sum_sq'][1])
aov_table['Eta squared'] = pd.Series([eta_sq, ''], index = [independent, 'Residual'])
return aov_table
def tukey(data=None, independent=None, dependent=None):
pd.set_eng_float_format(accuracy=3, use_eng_prefix=False)
independent = str(independent)
dependent = str(dependent)
if input_check_numerical_categorical(data, independent, dependent):
return
test = multi.MultiComparison(data[dependent], data[independent])
res = test.tukeyhsd()
print(res.summary())
return
def chi_square(data=None, variable1=None, variable2=None):
pd.set_eng_float_format(accuracy=3, use_eng_prefix=False)
variable1 = str(variable1)
variable2 = str(variable2)
if input_check_categorical_categorical(data, variable1, variable2):
return
values_var1=pd.unique(data[variable1])
values_var2=pd.unique(data[variable2])
problem_found=False
for variable in [values_var1, values_var2]:
if len(variable)<2:
print(variable, 'has less than two categories. It has:', len(variable))
problem_found=True
if problem_found:
return
contingency_table = pd.crosstab(data[variable1], data[variable2])
print('\033[1m' + 'Contingency Table' + '\033[0m')
print(contingency_table, '\n\n')
print('\033[1m' + 'Chi-square results' + '\033[0m')
chi2_test=stats.chi2_contingency(contingency_table, correction=False)
chi2_result= pd.Series ([chi2_test[0], chi2_test[1], chi2_test[2], chi2_test[3]],
index = ['Chi-square value', 'p-value', 'Degrees of freedom', 'Expected frequencies'])
return chi2_result
def logistic_reg(data=None, independent=None, dependent=None):
pd.set_eng_float_format(accuracy=3, use_eng_prefix=False)
independent = str(independent)
dependent = str(dependent)
if input_check_categorical(data, independent, dependent):
return
if not len(pd.unique(data[dependent]))==2:
print('Dependent variable must have two categories')
print(dependent, 'variable has', len(pd.unique(data[dependent])), 'categories')
return
data['interceptant']=1
independent=[independent, 'interceptant']
logReg = sm.Logit(data[dependent], data[independent])
regression = logReg.fit()
print(regression.summary(), '\n')
print('\033[1m' + 'Coefficients confidence intervals' + '\033[0m')
print(regression.conf_int())
predicted_values =regression.predict()
plt.plot(data['age'], data['osas'], 'o', label='Actual values')
plt.plot(data['age'], predicted_values, 'ok', label='Predicted probabilities')
plt.xlabel('Age', fontsize=14)
plt.ylabel('OSAS', fontsize=14)
plt.ylim(-0.05, 1.05)
plt.legend()
plt.show()
return
# Functions to validate statistical functions inputs
def input_check_numerical_categorical(data, independent, dependent):
problem_found=check_input_dataframe(data)
if check_variable_specified(independent):
print ('An independent variable was not specified')
problem_found=True
if check_variable_specified(dependent):
print ('A dependent variable was not specified')
problem_found=True
if problem_found:
return problem_found
if check_variables_are_columns(data, independent, dependent):
return True
if check_variable_types(data, dependent, ['int', 'float']):
problem_found=True
if check_variable_types(data, independent, ['bool', 'category']):
problem_found=True
return problem_found
def input_check_numerical_numerical(data, variable1, variable2):
problem_found=check_input_dataframe(data)
if check_variable_specified(variable1) or check_variable_specified(variable2):
print ('Two variables must be specified')
problem_found=True
if problem_found:
return problem_found
if check_variables_are_columns(data, variable1, variable2):
return True
for variable in [variable1, variable2]:
if check_variable_types(data, variable, ['int', 'float']):
problem_found=True
return problem_found
def input_check_categorical_categorical(data, variable1, variable2):
problem_found=check_input_dataframe(data)
if check_variable_specified(variable1) or check_variable_specified(variable2):
print ('Two variables must be specified')
problem_found=True
if problem_found:
return problem_found
if check_variables_are_columns(data, variable1, variable2):
return True
for variable in [variable1, variable2]:
if check_variable_types(data, variable, ['bool', 'category']):
problem_found=True
return problem_found
def input_check_categorical(data, independent, dependent):
problem_found=check_input_dataframe(data)
if check_variable_specified(independent):
print ('An independent variable was not specified')
problem_found=True
if check_variable_specified(dependent):
print ('A dependent variable was not specified')
problem_found=True
if problem_found:
return problem_found
if check_variables_are_columns(data, independent, dependent):
return True
if check_variable_types(data, dependent, ['bool', 'category']):
problem_found=True
return problem_found
# Functions to validate individual inputs
def check_input_dataframe(data):
if not str(type(data))=='<class \'pandas.core.frame.DataFrame\'>':
print (data, 'is not a DataFrame')
return True
else:
return False
def check_variable_specified(variable):
if variable==None:
return True
else:
return False
def check_variable_is_column(data, variable):
if variable not in data.columns:
print (variable, 'is not a column of', data, 'dataset')
return True
else:
return False
def check_variables_are_columns(data, variable1, variable2):
problem_found=False
for variable in [variable1, variable2]:
if check_variable_is_column(data, variable):
problem_found=True
return problem_found
def check_variable_types(data, variable, data_types):
if data[variable].dtypes not in data_types:
print (variable, 'is not of', data_types, 'type')
return True
else:
return False
# -
# __LOADING THE DATABASE__
# In this exercise we will use a database of patients evaluated for obstructive sleep apnea syndrome (OSAS). Each patient filled out a survey where epidemiological characteristics and symptoms were recorded. The database will contain some of those characteristics along with whether they had OSAS or not, and its severity, based on a measure of how frequently the patient stops breathing through the nigh called the Apnea-Hypopnea Index (ahi).
#
# We will upload the data we'll work into memory from a CSV file in the website GitHub and put it in a variable called "data". Please execute the following code cells.
data = pd.read_csv("https://raw.githubusercontent.com/gapatino/stats-notebooks/master/stats_workshop_database.csv")
# Then define some of the columns in the database as categorical variables
data['gender']=data['gender'].astype('category')
data['osas_severity']=data['osas_severity'].astype('category')
# Let's look at the data by displaying the first 10 rows of it
data.head(10)
# __APPLICATION EXERCISE__
# Below you will find questions about analyzing this data. After each question you will find a code cell and a text cell. Please enter the code for the appropriate statistical test in the code cell below it and run it, based on the output of the test answer the question in the text cell.
# If you need additional code cells you can add them by clicking on the button with the plus sign at the top of the page.
# __Question 1__
# What is the type of each variable (column) in the dataset table?
# Hint: You don't need to run any functions to answer this
# + active=""
# ANSWER:
# -
#
# __Question 2__
# What is the mean and standard deviation of the age of male subjects?
# + active=""
# ANSWER:
# -
#
# __Question 3__
# Does the BMI values have a normal distribution across OSAS patients and controls?
# + active=""
# ANSWER:
# -
#
# __Question 4__
# What is the median and interquartile range of BMI among smokers?
# + active=""
# ANSWER:
# -
#
# __Question 5__
# What is the range of AHI among subjects that snore?
# + active=""
# ANSWER:
# -
#
# __Question 6__
# How many levels of OSAS severity are there and how many subjects are in each of them?
# + active=""
# ANSWER:
# -
#
# __Question 7__
# Is there a difference in the mean age of subjects with and without OSAS?
# + active=""
# ANSWER:
# -
#
# __Question 8__
# Is there a difference in the mean BMI of subjects across the severity levels of OSAS?
# + active=""
# ANSWER:
# -
#
# __Question 9__
# Is there a difference in the number of subjects with apnea between those with and without OSAS?
# + active=""
# ANSWER:
# -
#
# __Question 10__
# Can the age predict if a subject will have OSAS?
# + active=""
# ANSWER:
# -
#
# __Question 11__
# Did you find this session useful?
import ipywidgets as widgets
widgets.RadioButtons(
options=['Yes', 'No'],
description=' ',
disabled=False
)
#
# __Question 12__
# Would you prefer to have future statistics sessions delivered as regular lectures or hands-on exercises like this one?
widgets.RadioButtons(
options=['Yes', 'No'],
description=' ',
disabled=False
)
| Stats_notebook_Python_full.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from absl import logging
import tensorflow.compat.v1 as tf
from open_spiel.python import policy
from open_spiel.python import rl_environment
from open_spiel.python.algorithms import exploitability
from open_spiel.python.algorithms import nfsp
from open_spiel.python.pytorch import nfsp as nfsp_pt
class NFSPPolicies(policy.Policy):
"""Joint policy to be evaluated."""
def __init__(self, env, nfsp_policies, mode):
game = env.game
player_ids = [0, 1]
super(NFSPPolicies, self).__init__(game, player_ids)
self._policies = nfsp_policies
self._mode = mode
self._obs = {"info_state": [None, None], "legal_actions": [None, None]}
def action_probabilities(self, state, player_id=None):
cur_player = state.current_player()
legal_actions = state.legal_actions(cur_player)
self._obs["current_player"] = cur_player
self._obs["info_state"][cur_player] = (
state.information_state_tensor(cur_player))
self._obs["legal_actions"][cur_player] = legal_actions
info_state = rl_environment.TimeStep(
observations=self._obs, rewards=None, discounts=None, step_type=None)
with self._policies[cur_player].temp_mode_as(self._mode):
p = self._policies[cur_player].step(info_state, is_evaluation=True).probs
prob_dict = {action: p[action] for action in legal_actions}
return prob_dict
def tf_main(game,
env_config,
num_train_episodes,
eval_every,
hidden_layers_sizes,
replay_buffer_capacity,
reservoir_buffer_capacity,
anticipatory_param):
env = rl_environment.Environment(game, **env_configs)
info_state_size = env.observation_spec()["info_state"][0]
num_actions = env.action_spec()["num_actions"]
hidden_layers_sizes = [int(l) for l in hidden_layers_sizes]
kwargs = {
"replay_buffer_capacity": replay_buffer_capacity,
"epsilon_decay_duration": num_train_episodes,
"epsilon_start": 0.06,
"epsilon_end": 0.001,
}
expl_list = []
with tf.Session() as sess:
# pylint: disable=g-complex-comprehension
agents = [
nfsp.NFSP(sess, idx, info_state_size, num_actions, hidden_layers_sizes,
reservoir_buffer_capacity, anticipatory_param,
**kwargs) for idx in range(num_players)
]
expl_policies_avg = NFSPPolicies(env, agents, nfsp.MODE.average_policy)
sess.run(tf.global_variables_initializer())
for ep in range(num_train_episodes):
if (ep + 1) % eval_every == 0:
losses = [agent.loss for agent in agents]
print("Losses: %s" %losses)
expl = exploitability.exploitability(env.game, expl_policies_avg)
expl_list.append(expl)
print("[%s] Exploitability AVG %s" %(ep + 1, expl))
print("_____________________________________________")
time_step = env.reset()
while not time_step.last():
player_id = time_step.observations["current_player"]
agent_output = agents[player_id].step(time_step)
action_list = [agent_output.action]
time_step = env.step(action_list)
# Episode is over, step all agents with final info state.
for agent in agents:
agent.step(time_step)
return expl_list
def pt_main(game,
env_config,
num_train_episodes,
eval_every,
hidden_layers_sizes,
replay_buffer_capacity,
reservoir_buffer_capacity,
anticipatory_param):
env = rl_environment.Environment(game, **env_configs)
info_state_size = env.observation_spec()["info_state"][0]
num_actions = env.action_spec()["num_actions"]
hidden_layers_sizes = [int(l) for l in hidden_layers_sizes]
kwargs = {
"replay_buffer_capacity": replay_buffer_capacity,
"epsilon_decay_duration": num_train_episodes,
"epsilon_start": 0.06,
"epsilon_end": 0.001,
}
expl_list = []
agents = [
nfsp_pt.NFSP(idx, info_state_size, num_actions, hidden_layers_sizes,
reservoir_buffer_capacity, anticipatory_param,
**kwargs) for idx in range(num_players)
]
expl_policies_avg = NFSPPolicies(env, agents, nfsp_pt.MODE.average_policy)
for ep in range(num_train_episodes):
if (ep + 1) % eval_every == 0:
losses = [agent.loss.item() for agent in agents]
print("Losses: %s" %losses)
expl = exploitability.exploitability(env.game, expl_policies_avg)
expl_list.append(expl)
print("[%s] Exploitability AVG %s" %(ep + 1, expl))
print("_____________________________________________")
time_step = env.reset()
while not time_step.last():
player_id = time_step.observations["current_player"]
agent_output = agents[player_id].step(time_step)
action_list = [agent_output.action]
time_step = env.step(action_list)
# Episode is over, step all agents with final info state.
for agent in agents:
agent.step(time_step)
return expl_list
# -
game = "kuhn_poker"
num_players = 2
env_configs = {"players": num_players}
num_train_episodes = int(3e6)
eval_every = 10000
hidden_layers_sizes = [128]
replay_buffer_capacity = int(2e5)
reservoir_buffer_capacity = int(2e6)
anticipatory_param = 0.1
tf_kuhn_result = tf_main(game,
env_configs,
num_train_episodes,
eval_every,
hidden_layers_sizes,
replay_buffer_capacity,
reservoir_buffer_capacity,
anticipatory_param)
pt_kuhn_result = pt_main(game,
env_configs,
num_train_episodes,
eval_every,
hidden_layers_sizes,
replay_buffer_capacity,
reservoir_buffer_capacity,
anticipatory_param)
# +
import matplotlib.pyplot as plt
x = [i*1000 for i in range(len(tf_kuhn_result))]
plt.plot(x, tf_kuhn_result, label='tensorflow')
plt.plot(x, pt_kuhn_result, label='pytorch')
plt.title('Kuhn Poker')
plt.xlabel('Episodes')
plt.ylabel('Exploitability')
plt.legend()
plt.show()
# -
game = "leduc_poker"
num_players = 2
env_configs = {"players": num_players}
num_train_episodes = int(3e6)
eval_every = 100000
hidden_layers_sizes = [128]
replay_buffer_capacity = int(2e5)
reservoir_buffer_capacity = int(2e6)
anticipatory_param = 0.1
tf_leduc_result = tf_main(game,
env_configs,
num_train_episodes,
eval_every,
hidden_layers_sizes,
replay_buffer_capacity,
reservoir_buffer_capacity,
anticipatory_param)
pt_leduc_result = pt_main(game,
env_configs,
num_train_episodes,
eval_every,
hidden_layers_sizes,
replay_buffer_capacity,
reservoir_buffer_capacity,
anticipatory_param)
# +
x = [i * 10000 for i in range(len(tf_leduc_result))]
plt.plot(x, tf_leduc_result, label='tensorflow')
plt.plot(x, pt_leduc_result, label='pytorch')
plt.title('Leduc Poker')
plt.xlabel('Episodes')
plt.ylabel('Exploitability')
plt.legend()
plt.show()
# -
| open_spiel/colabs/research_nfsp_tf_pt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Credit card fraud detector using Amazon Fraud Detector
# ## Investigate and process the data
# +
from IPython.display import clear_output
from datetime import datetime
from io import StringIO
from sklearn.model_selection import train_test_split
from imblearn.under_sampling import RandomUnderSampler
from imblearn.over_sampling import SMOTENC
from collections import Counter
from sklearn.metrics import roc_curve, roc_auc_score, auc, roc_auc_score
# %matplotlib inline
import os
import sys
import json
import uuid
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import boto3
import time
# -
# Let's start by downloading and reading in the credit card fraud data set.
# + language="bash"
# wget https://fraud-detector-blog-assets.s3.amazonaws.com/creditcard.csv
# +
# Resources and env variables setup
s3_resource = boto3.resource('s3')
afd_resource = boto3.client('frauddetector')
# suffix is appended to detector and model name for uniqueness
sufx = datetime.now().strftime("%Y%m%d")
# replace with the bucket created in the CloudFormation
S3_BUCKET = "afd-poc-trainingbucket-1i37svk9elcoe"
# Replace the ARN Role with the resources created in CloudFormation stack
ARN_ROLE = "arn:aws:iam::387461613214:role/afd-poc-NotebookInstanceExecutionRole-1FNQ41S8H2G68"
# +
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
data = pd.read_csv('creditcard.csv', delimiter=',')
# -
# Let's take a peek at our data (we only show a subset of the columns in the table):
print(data.columns)
data.describe()
nonfrauds, frauds = data.groupby('Class').size()
print('Number of frauds: ', frauds)
print('Number of non-frauds: ', nonfrauds)
print('Percentage of fradulent data:', 100.*frauds/(frauds + nonfrauds))
# The class column corresponds to whether or not a transaction is fradulent. We see that the majority of data is non-fraudulent with only $492$ ($.172\%$), check the Class column mean, of the data corresponding to fraudulent examples.
# A PCA have been made lets check the mean and standard deviation of the features.
data.hist(bins=50,figsize=(20,15))
plt.show()
# Looks good, columns have been normalized to have 0 mean and unit standard deviation as the result of a PCA. Now, lets change the data to be Amazon Fraud Detector compatible.
# to lowercase
data.columns = map(str.lower, data.columns)
print(data.columns)
# +
# mapping column names numbers to letters
def standardize_headers(x):
if any(char.isdigit() for char in x):
if int(x[1:]) > 26:
return 'va'+chr(int(x[1:])+70)
return 'v'+chr(int(x[1:])+96)
return x
data.rename(columns=standardize_headers, inplace=True)
print(data.columns)
# -
# Then change the timestamp and label column names
# rename to the Amazon Fraud Detector name conventions
data.rename(columns={'time':'EVENT_TIMESTAMP','class':'EVENT_LABEL'}, inplace=True)
data.head()
# The date column is represented as incremental seconds, lets translate that to real dates.
# +
# Get epoch time for the initial dataset date
epoch = datetime.utcfromtimestamp(0)
def unix_time_seconds(dt):
return (dt - epoch).total_seconds()
# Lets pretend that the data is from yesterday and could can test at the end with todays date.
start_dt = datetime.strptime('Sep 22 2020 12:00AM', '%b %d %Y %I:%M%p')
start_dt = datetime.now()
start_ep = unix_time_seconds(start_dt)
print(start_ep)
# -
# Datetime parse test
date_str = '9/22/2020 12:00:00 AM'
date = datetime.strptime(date_str, "%m/%d/%Y %I:%M:%S %p")
print(date)
# Translate the current timestamp format (increasing seconds) to ISO 8601 standard
# +
# translate seconds delta to actual datetimes in ISO 8601
def to_datetime(x):
current_ep = start_ep + x
current_dt = time.strftime('%Y-%m-%dT%H:%M:%SZ', time.localtime(current_ep))
return current_dt
data['EVENT_TIMESTAMP'] = data['EVENT_TIMESTAMP'].apply(to_datetime)
data.head()
# -
# Lets check for null values
data.isnull().sum()
# We will split our dataset into a train and test to evaluate the performance of our models. It's important to do so before any techniques meant to alleviate the class imbalance are used. This ensures that we don't leak information from the test set into the train set.
features = data.drop('EVENT_LABEL', axis=1).values
labels = (data['EVENT_LABEL'].values).astype('float32')
X, X_test, y, y_test = train_test_split(
features, labels, test_size=0.1, random_state=42)
counter = Counter(y)
print(counter)
# Getting the train DataFrame back together
saved_cols = data.drop('EVENT_LABEL', axis=1).columns
print(saved_cols)
data = pd.DataFrame(X, columns = saved_cols)
data['EVENT_LABEL']=y.astype(int)
data.head(10)
test = pd.DataFrame(X_test, columns = saved_cols)
test.head(10)
# The testing dataset with the labels to perform evaluations latter on
test_label = pd.DataFrame(X_test, columns = saved_cols)
test_label['EVENT_LABEL']=y_test.astype(int)
test_label.head()
#validating the test dataset with labels
nonfrauds, frauds = test_label.groupby('EVENT_LABEL').size()
print('Number of frauds in test data: ', frauds)
print('Number of non-frauds in test data: ', nonfrauds)
print('Percentage of fradulent data:', 100.*frauds/(frauds + nonfrauds))
# +
#validating the training dataset
nonfrauds, frauds = data.groupby('EVENT_LABEL').size()
print('Number of frauds: ', frauds)
print('Number of non-frauds: ', nonfrauds)
print('Percentage of fradulent data:', 100.*frauds/(frauds + nonfrauds))
count_class_0, count_class_1 = data.EVENT_LABEL.value_counts()
data.EVENT_LABEL.value_counts().plot(kind='bar', title='Count (EVENT_LABEL)');
# -
# Uploading the data for training
csv_buffer = StringIO()
data.to_csv(csv_buffer, index=False)
s3_resource.Object(S3_BUCKET, 'dataset-training.csv').put(Body=csv_buffer.getvalue())
# Uploading the data for testing
# +
csv_buffer = StringIO()
test.to_csv(csv_buffer, index=False)
s3_resource.Object(S3_BUCKET, 'dataset-test.csv').put(Body=csv_buffer.getvalue())
# -
# Once we have the datasets ready we need create the necesary entities for build and deploy the fraud detection model. This can be done within the Amazon Fraud Detector console visually or through the API as shown in the following seccion.
# -- This is all you need to fill out. Once complete simply interactively run each code cell. --
# your_entity_name
ENTITY_TYPE = "creditcardtrans{0}".format(sufx)
ENTITY_DESC = "creditcard transactions: {0}".format(sufx)
# your_event_type
EVENT_TYPE = "creditcard{0}".format(sufx)
EVENT_DESC = "creditcard card payment events: {0}".format(sufx)
# your_model_name
MODEL_NAME = "fraud_detector_model{0}".format(sufx)
MODEL_DESC = "model trained on: {0}".format(sufx)
# your_detector_name
DETECTOR_NAME = "fraud_detector_endpoint{0}".format(sufx)
DETECTOR_DESC = "detects synthetic fraud events created: {0}".format(sufx)
# ### 1. Profile Your Dataset
# -----
#
#
# <div class="alert alert-info"> 💡 <strong> Profiling </strong>
#
# The function below will: 1. profile your data, creating descriptive statistics, 2. perform basic data quality checks (nulls, unique variables, etc.), and 3. return summary statistics and the EVENT and MODEL schemas used to define your EVENT_TYPE and TRAIN your MODEL.
#
#
# </div>
# +
# --- no changes; just run this code block ---
def summary_stats(df):
""" Generate summary statistics for a panda's data frame
Args:
df (DataFrame): panda's dataframe to create summary statistics for.
Returns:
DataFrame of summary statistics, training data schema, event variables and event lables
"""
df = df.copy()
rowcnt = len(df)
df_s1 = df.agg(['count', 'nunique']).transpose().reset_index().rename(columns={"index":"feature_name"})
df_s1["null"] = (rowcnt - df_s1["count"]).astype('int64')
df_s1["not_null"] = rowcnt - df_s1["null"]
df_s1["null_pct"] = df_s1["null"] / rowcnt
df_s1["nunique_pct"] = df_s1['nunique']/ rowcnt
dt = pd.DataFrame(df.dtypes).reset_index().rename(columns={"index":"feature_name", 0:"dtype"})
df_stats = pd.merge(dt, df_s1, on='feature_name', how='inner').round(4)
df_stats['nunique'] = df_stats['nunique'].astype('int64')
df_stats['count'] = df_stats['count'].astype('int64')
# -- variable type mapper --
df_stats['feature_type'] = "UNKOWN"
df_stats.loc[df_stats["dtype"] == object, 'feature_type'] = "CATEGORY"
df_stats.loc[(df_stats["dtype"] == "int64") | (df_stats["dtype"] == "float64"), 'feature_type'] = "NUMERIC"
df_stats.loc[df_stats["feature_name"].str.contains("ipaddress|ip_address|ipcli"), 'feature_type'] = "IP_ADDRESS"
df_stats.loc[df_stats["feature_name"].str.contains("email|email_address|emailaddr"), 'feature_type'] = "EMAIL_ADDRESS"
df_stats.loc[df_stats["feature_name"].str.contains("canal|channel"), 'feature_type'] = "USERAGENT"
df_stats.loc[df_stats["feature_name"].str.contains("monto|amount"), 'feature_type'] = "PRICE"
df_stats.loc[df_stats["feature_name"].str.contains("nomdes|name"), 'feature_type'] = "BILLING_NAME"
df_stats.loc[df_stats["feature_name"] == "EVENT_LABEL", 'feature_type'] = "TARGET"
df_stats.loc[df_stats["feature_name"] == "EVENT_TIMESTAMP", 'feature_type'] = "EVENT_TIMESTAMP"
# -- variable warnings --
df_stats['feature_warning'] = "NO WARNING"
df_stats.loc[(df_stats["nunique"] != 2) & (df_stats["feature_name"] == "EVENT_LABEL"),'feature_warning' ] = "LABEL WARNING, NON-BINARY EVENT LABEL"
df_stats.loc[(df_stats["nunique_pct"] > 0.97) & (df_stats['feature_type'] == "CATEGORY") ,'feature_warning' ] = "EXCLUDE, GT 97% UNIQUE"
df_stats.loc[(df_stats["null_pct"] > 0.2) & (df_stats["null_pct"] <= 0.5), 'feature_warning' ] = "NULL WARNING, GT 20% MISSING"
df_stats.loc[df_stats["null_pct"] > 0.5,'feature_warning' ] = "EXCLUDE, GT 50% MISSING"
df_stats.loc[((df_stats['dtype'] == "int64" ) | (df_stats['dtype'] == "float64" ) ) & (df_stats['nunique'] < 0.2), 'feature_warning' ] = "LIKELY CATEGORICAL, NUMERIC w. LOW CARDINALITY"
# -- target check --
exclude_fields = df_stats.loc[(df_stats['feature_warning'] != 'NO WARNING')]['feature_name'].to_list()
event_variables = df_stats.loc[(~df_stats['feature_name'].isin(['EVENT_LABEL', 'EVENT_TIMESTAMP']))]['feature_name'].to_list()
event_labels = df["EVENT_LABEL"].unique().tolist()
trainingDataSchema = {
'modelVariables' : df_stats.loc[(df_stats['feature_type'].isin(['IP_ADDRESS', 'EMAIL_ADDRESS', 'CATEGORY', 'NUMERIC' ]))]['feature_name'].to_list(),
'labelSchema' : {
'labelMapper' : {
'FRAUD' : [str(df["EVENT_LABEL"].value_counts().idxmin())],
'LEGIT' : [str(df["EVENT_LABEL"].value_counts().idxmax())]
}
}
}
model_variables = df_stats.loc[(df_stats['feature_type'].isin(['IP_ADDRESS', 'EMAIL_ADDRESS', 'CATEGORY', 'NUMERIC' ]))]['feature_name'].to_list()
# -- label schema --
label_map = {
'FRAUD' : [df["EVENT_LABEL"].value_counts().idxmin()],
'LEGIT' : [df["EVENT_LABEL"].value_counts().idxmax()]
}
print("--- summary stats ---")
print(df_stats)
print("\n")
print("--- event variables ---")
print(event_variables)
print("\n")
print("--- event labels ---")
print(event_labels)
print("\n")
print("--- training data schema ---")
print(trainingDataSchema)
print("\n")
return df_stats, trainingDataSchema, event_variables, event_labels
# -- connect to S3, snag file, and convert to a panda's dataframe --
#s3 = boto3.resource('s3')
#obj = s3.Object(S3_BUCKET, S3_FILE)
#body = obj.get()['Body']
#df = pd.read_csv(body)
# -- call profiling function --
df_stats, trainingDataSchema, eventVariables, eventLabels = summary_stats(data)
# -
# ### 2. Create Variables
# -----
#
# <div class="alert alert-info"> 💡 <strong> Create Variables. </strong>
#
# The following section will automatically create your modeling input variables and your model scoring variable for you.
#
# </div>
# +
# --- no changes just run this code block ---
def create_label(df, FRAUD_LABEL):
"""
Returns a dictionary for the model labelSchema, by identifying the rare event as fraud / and common as not-fraud
Arguments:
df -- input dataframe
FRAUD_LABEL -- the name of the field that contains fraud label
Returns:
labelSchema -- a dictionary containing labelKey & labelMapper
"""
label_summary = df[FRAUD_LABEL].value_counts()
labelSchema = {'labelKey': FRAUD_LABEL,
"labelMapper" : { "FRAUD": [str(label_summary.idxmin())],
"LEGIT": [str(label_summary.idxmax())]}
}
afd_resource.put_label(
name = str(label_summary.idxmin()),
description = 'FRAUD')
afd_resource.put_label(
name = str(label_summary.idxmax()),
description = 'LEGIT')
return labelSchema
# -- function to create all your variables ---
def create_variables(df_stats, MODEL_NAME):
"""
Returns a variable list of model input variables, checks to see if variable exists,
and, if not, then it adds the variable to Fraud Detector
Arguments:
enrichment_features -- dictionary of optional features, mapped to specific variable types enriched (CARD_BIN, USERAGENT)
numeric_features -- optional list of numeric field names
categorical_features -- optional list of categorical features
Returns:
variable_list -- a list of variable dictionaries
"""
enrichment_features = df_stats.loc[(df_stats['feature_type'].isin(['IP_ADDRESS', 'EMAIL_ADDRESS', 'USERAGENT', 'BILLING_NAME', 'PRICE']))]['feature_name'].to_dict()
enrichment_type = df_stats.loc[(df_stats['feature_type'].isin(['IP_ADDRESS', 'EMAIL_ADDRESS', 'USERAGENT', 'BILLING_NAME', 'PRICE']))]['feature_type'].to_dict()
numeric_features = df_stats.loc[(df_stats['feature_type'].isin(['NUMERIC']))]['feature_name'].to_dict()
categorical_features = df_stats.loc[(df_stats['feature_type'].isin(['CATEGORY']))]['feature_name'].to_dict()
variable_list = []
# -- first do the enrichment features
for feature in enrichment_features.keys():
variable_list.append( {'name' : enrichment_features[feature]+""})
try:
varname = enrichment_features[feature]+""
afd_resource.get_variables(name=varname)
except:
print("Creating variable: {0}".format(enrichment_features[feature]))
if enrichment_type[feature] == "PRICE":
resp = afd_resource.create_variable(
name = varname,
dataType = 'FLOAT',
dataSource ='EVENT',
defaultValue = '0',
description = enrichment_features[feature],
variableType = enrichment_type[feature] )
else:
resp = afd_resource.create_variable(
name = varname,
dataType = 'STRING',
dataSource ='EVENT',
defaultValue = '<unknown>',
description = enrichment_features[feature],
variableType = enrichment_type[feature] )
# -- check and update the numeric features
for feature in numeric_features:
variable_list.append( {'name' : numeric_features[feature]+""})
try:
varname = numeric_features[feature]+""
afd_resource.get_variables(name=varname)
except:
print("Creating variable: {0}".format(numeric_features[feature]))
resp = afd_resource.create_variable(
name = varname,
dataType = 'FLOAT',
dataSource ='EVENT',
defaultValue = '0.0',
description = numeric_features[feature],
variableType = 'NUMERIC' )
# -- check and update the categorical features
for feature in categorical_features:
variable_list.append( {'name' : categorical_features[feature]+""})
try:
varname = categorical_features[feature]+""
afd_resource.get_variables(name=varname)
except:
print("Creating variable: {0}".format(categorical_features[feature]))
resp = afd_resource.create_variable(
name = varname,
dataType = 'STRING',
dataSource ='EVENT',
defaultValue = '<unknown>',
description = categorical_features[feature],
variableType = 'CATEGORICAL' )
# -- create a model score feature
model_feature = "{0}_insightscore".format(MODEL_NAME)
# variable_list.append( {'name' : model_feature})
try:
afd_resource.get_variables(name=model_feature)
except:
print("Creating variable: {0}".format(model_feature))
resp = afd_resource.create_variable(
name = model_feature,
dataType = 'FLOAT',
dataSource ='MODEL_SCORE',
defaultValue = '0.0',
description = model_feature,
variableType = 'NUMERIC' )
return variable_list
model_variables = create_variables(df_stats, MODEL_NAME)
print("\n --- model variable dict --")
print(model_variables)
model_label = create_label(data, "EVENT_LABEL")
print("\n --- model label schema dict --")
print(model_label)
# -
# ### 3. Create Entity and Event Types
# -----
#
# <div class="alert alert-info"> 💡 <strong> Entity and Event. </strong>
#
# The following code block will automatically create your entity and event types for you.
#
# </div>
#Amazon Fraud Detector expect the labels to be strings.
eventLabels = list(map(str, eventLabels))
print(eventLabels)
# +
# --- no changes just run this code block ---
response = afd_resource.put_entity_type(
name = ENTITY_TYPE,
description = ENTITY_DESC
)
print("-- create entity --")
print(response)
response = afd_resource.put_event_type (
name = EVENT_TYPE,
eventVariables = eventVariables,
labels = eventLabels,
entityTypes = [ENTITY_TYPE])
print("-- create event type --")
print(response)
# -
# ### 4. Create & Train your Model
# -----
#
# <div class="alert alert-info"> 💡 <strong> Train Model. </strong>
#
# The following section will automatically train and activate your model for you.
#
# </div>
# -- create our model --
response = afd_resource.create_model(
description = MODEL_DESC,
eventTypeName = EVENT_TYPE,
modelId = MODEL_NAME,
modelType = 'ONLINE_FRAUD_INSIGHTS')
print("-- initalize model --")
print(response)
# +
# -- initializes the model, it's now ready to train --
S3_FILE = "dataset-training.csv"
S3_FILE_LOC = "s3://{0}/{1}".format(S3_BUCKET,S3_FILE)
response = afd_resource.create_model_version(
modelId = MODEL_NAME,
modelType = 'ONLINE_FRAUD_INSIGHTS',
trainingDataSource = 'EXTERNAL_EVENTS',
trainingDataSchema = trainingDataSchema,
externalEventsDetail = {
'dataLocation' : S3_FILE_LOC,
'dataAccessRoleArn': ARN_ROLE
}
)
print("-- model training --")
print(response)
# +
# -- model training takes time, we'll loop until it's complete --
print("-- wait for model training to complete --")
stime = time.time()
while True:
clear_output(wait=True)
response = afd_resource.get_model_version(modelId=MODEL_NAME, modelType = "ONLINE_FRAUD_INSIGHTS", modelVersionNumber = '1.0')
if response['status'] == 'TRAINING_IN_PROGRESS':
print(f"current progress: {(time.time() - stime)/60:{3}.{3}} minutes")
time.sleep(60) # -- sleep for 60 seconds
if response['status'] != 'TRAINING_IN_PROGRESS':
print("Model status : " + response['status'])
break
etime = time.time()
# -- summarize --
print("\n --- model training complete --")
print("Elapsed time : %s" % (etime - stime) + " seconds \n" )
print(response)
# -
# ### 5. Activate the Model and evaluate the performance
# -----
#
# <div class="alert alert-info"> 💡 <strong> Train Model. </strong>
#
# The following section will automatically train and activate your model for you.
#
# </div>
# +
response = afd_resource.update_model_version_status (
modelId = MODEL_NAME,
modelType = 'ONLINE_FRAUD_INSIGHTS',
modelVersionNumber = '1.0',
status = 'ACTIVE'
)
print("-- activating model --")
print(response)
#-- wait until model is active
print("--- waiting until model status is active ")
stime = time.time()
while True:
clear_output(wait=True)
response = afd_resource.get_model_version(modelId=MODEL_NAME, modelType = "ONLINE_FRAUD_INSIGHTS", modelVersionNumber = '1.0')
if response['status'] != 'ACTIVE':
print(f"current progress: {(time.time() - stime)/60:{3}.{3}} minutes")
time.sleep(60) # sleep for 1 minute
if response['status'] == 'ACTIVE':
print("Model status : " + response['status'])
break
etime = time.time()
print("Elapsed time : %s" % (etime - stime) + " seconds \n" )
print(response)
# +
# -- model performance summary --
auc = afd_resource.describe_model_versions(
modelId= MODEL_NAME,
modelVersionNumber='1.0',
modelType='ONLINE_FRAUD_INSIGHTS',
maxResults=10
)['modelVersionDetails'][0]['trainingResult']['trainingMetrics']['auc']
df_model = pd.DataFrame(afd_resource.describe_model_versions(
modelId= MODEL_NAME,
modelVersionNumber='1.0',
modelType='ONLINE_FRAUD_INSIGHTS',
maxResults=10
)['modelVersionDetails'][0]['trainingResult']['trainingMetrics']['metricDataPoints'])
plt.figure(figsize=(10,10))
plt.plot(df_model["fpr"], df_model["tpr"], color='darkorange',
lw=2, label='ROC curve (area = %0.3f)' % auc)
plt.plot([0, 1], [0, 1], color='navy', lw=2, linestyle='--')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title( MODEL_NAME + ' ROC Chart')
plt.legend(loc="lower right",fontsize=12)
plt.axvline(x = 0.02 ,linewidth=2, color='r')
plt.axhline(y = 0.73 ,linewidth=2, color='r')
plt.show()
# -
# ### 6. Create Detector, generate Rules and assemble your Detector
#
# -----
#
# <div class="alert alert-info"> 💡 <strong> Generate Rules, Create and Publish a Detector. </strong>
#
# The following section will automatically generate a number of fraud, investigate and approve rules based on the false positive rate and score thresholds of your model. These are just example rules that you could create, it is recommended that you fine tune your rules specifically to your business use case.
# </div>
# +
# -- initialize your detector --
response = afd_resource.put_detector(detectorId = DETECTOR_NAME,
description = DETECTOR_DESC,
eventTypeName = EVENT_TYPE )
print(response)
# +
# -- make rules --
model_stat = df_model.round(decimals=2)
m = model_stat.loc[model_stat.groupby(["fpr"])["threshold"].idxmax()]
def make_rule(x):
rule = ""
if x['fpr'] <= 0.05:
rule = "${0}_insightscore > {1}".format(MODEL_NAME,x['threshold'])
if x['fpr'] == 0.06:
rule = "${0}_insightscore <= {1}".format(MODEL_NAME,x['threshold_prev'])
return rule
m["threshold_prev"] = m['threshold'].shift(1)
m['rule'] = m.apply(lambda x: make_rule(x), axis=1)
m['outcome'] = "approve"
m.loc[m['fpr'] <= 0.03, "outcome"] = "fraud"
m.loc[(m['fpr'] > 0.03) & (m['fpr'] <= 0.05), "outcome"] = "investigate"
print (" --- score thresholds 1% to 6% --- ")
print(m[["fpr", "tpr", "threshold", "rule", "outcome"]].loc[(m['fpr'] > 0.0 ) & (m['fpr'] <= 0.06)].reset_index(drop=True))
# +
# -- create outcomes --
def create_outcomes(outcomes):
""" create Fraud Detector Outcomes
"""
for outcome in outcomes:
print("creating outcome variable: {0} ".format(outcome))
response = afd_resource.put_outcome(
name=outcome,
description=outcome)
# -- get distinct outcomes
outcomes = m["outcome"].unique().tolist()
create_outcomes(outcomes)
# -
rule_set = m[(m["fpr"] > 0.0) & (m["fpr"] <= 0.06)][["outcome", "rule"]].to_dict('records')
rule_list = []
for i, rule in enumerate(rule_set):
ruleId = "rule{0}_{1}".format(i, MODEL_NAME)
rule_list.append({"ruleId": ruleId,
"ruleVersion" : '1',
"detectorId" : DETECTOR_NAME
})
print("creating rule: {0}: IF {1} THEN {2}".format(ruleId, rule["rule"], rule['outcome']))
try:
response = afd_resource.create_rule(
ruleId = ruleId,
detectorId = DETECTOR_NAME,
expression = rule['rule'],
language = 'DETECTORPL',
outcomes = [rule['outcome']]
)
except:
print("this rule already exists in this detector")
rule_list
# +
afd_resource.create_detector_version(
detectorId = DETECTOR_NAME,
rules = rule_list,
modelVersions = [{"modelId":MODEL_NAME,
"modelType" : "ONLINE_FRAUD_INSIGHTS",
"modelVersionNumber" : "1.0"}],
ruleExecutionMode = 'FIRST_MATCHED'
)
print("\n -- detector created -- ")
print(response)
# -
response = afd_resource.update_detector_version_status(
detectorId= DETECTOR_NAME,
detectorVersionId='1',
status='ACTIVE'
)
print("\n -- detector activated -- ")
print(response)
# Testing our model endpoint
# +
S3_FILE = "dataset-test.csv"
S3_FILE_LOC = "s3://{0}/{1}".format(S3_BUCKET,S3_FILE)
s3_resource.Bucket(S3_BUCKET).download_file(S3_FILE, 'dataset-test.csv')
# +
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
test = pd.read_csv('dataset-test.csv', delimiter=',')
test.head(10)
# -
# Cleaning the test dataset from training columns and defining the start datetime.
record_count = 400
model_variables = [column for column in test.columns if column not in ['EVENT_LABEL', 'EVENT_TIMESTAMP']]
#dateTimeObj = datetime.strptime('Sep 3 2013 12:00AM', '%b %d %Y %I:%M%p')
dateTimeObj = datetime.now()
timestampStr = dateTimeObj.strftime("%Y-%m-%dT%H:%M:%SZ")
print(' '.join(model_variables))
# +
import uuid
# test the endpoint with a single prediction.
eventId = uuid.uuid1()
testrecord = test[model_variables].head(15).astype(str).to_dict(orient='records')[6]
pred = afd_resource.get_event_prediction(detectorId=DETECTOR_NAME,
detectorVersionId='1',
eventId = str(eventId),
eventTypeName = EVENT_TYPE,
eventTimestamp = timestampStr,
entities = [{'entityType': ENTITY_TYPE, 'entityId':str(eventId.int)}],
eventVariables= testrecord)
print(pred)
# -
# The next block will use some parallelization to run several test against the fraud detector endpoint.
# +
import dask
import time
from IPython.core.display import display, HTML
#display(HTML("<style>.container { width:90% }</style>"))
start = time.time()
@dask.delayed
def _predict(record):
eventId = uuid.uuid1()
try:
pred = afd_resource.get_event_prediction(detectorId=DETECTOR_NAME,
detectorVersionId='1',
eventId = str(eventId),
eventTypeName = EVENT_TYPE,
eventTimestamp = timestampStr,
entities = [{'entityType': ENTITY_TYPE, 'entityId':str(eventId.int)}],
eventVariables= record)
record["score"] = pred['modelScores'][0]['scores']["{0}_insightscore".format(MODEL_NAME)]
if len(pred['ruleResults']) > 0:
record["outcomes"]= pred['ruleResults'][0]['outcomes']
else:
record["outcomes"]= 'approve'
return record
except:
pred = afd_resource.get_event_prediction(detectorId=DETECTOR_NAME,
detectorVersionId='1',
eventId = str(eventId),
eventTypeName = EVENT_TYPE,
eventTimestamp = timestampStr,
entities = [{'entityType': ENTITY_TYPE, 'entityId':str(eventId.int)}],
eventVariables= record)
record["score"] = "-999"
record["outcomes"]= "error"
return record
predict_data = test[model_variables].head(record_count).astype(str).to_dict(orient='records')
predict_score = []
i=0
for record in predict_data:
clear_output(wait=True)
rec = dask.delayed(_predict)(record)
predict_score.append(rec)
i += 1
print("current progress: ", round((i/record_count)*100,2), "%" )
predict_recs = dask.compute(*predict_score)
# Calculate time taken and print results
time_taken = time.time() - start
tps = len(predict_recs) / time_taken
print ('Process took %0.2f seconds' %time_taken)
print ('Scored %d records' %len(predict_recs))
# -
# lets take a look to the predicted frauds
predictions = pd.DataFrame.from_dict(predict_recs, orient='columns')
predictions.loc[predictions['score'].astype('float32') > 900]
data[(data["NomDes"]=="victoria") & (data["RUTDES"]== 'b2d3f8b557944f6780d6d60f7b923eaf')]
data[(data["NomDes"]=="alan cerda") & (data["RUTDES"]== '781c1bcc58864eab9c42b5e2a8b40660')]
# See the model metrics on CloudWatch and the prediction history in Fraud Detector.
# save the results to a csv file and upload it to the output s3 bucket
csv_buffer = StringIO()
predictions.to_csv(csv_buffer, index=False)
s3_resource.Object(S3_BUCKET, MODEL_NAME + "precictions{}.csv".format(sufx)).put(Body=csv_buffer.getvalue())
# +
#data.loc[data['vaa'] == 0.14205158164005, 'vaa':'EVENT_LABEL']
# -
# Finish
| notebooks/frauddetector_fraud_detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Colorbot
#
# **Special thanks @MarkDaoustthat helped us with this material**
#
# In order to have a better experience follow these steps:
#
# 1. Read all the notebook, try to understand what each part of the code is doing and get familiar with the implementation;
# 2. For each exercise in this notebook make a copy of this notebook and try to implement what is expected. We suggest the following order for the exercises: *EXERCISE: HYPERPARAMETERS*, *EXERCISE: EXPERIMENT*, *EXERCISE: DATASET*;
# 3. Troubles or doubts about the code/exercises? Ask the instructor about it or go to the end of this notebook for a possible implementation/instruction for the exercises.
#
# ## Content of this notebook
#
# In this notebook you'll find a full implementation of a RNN model using the TensorFlow Estimators including comments and details about how to do it.
#
# Once you finish this notebook, you'll have a better understanding of:
# * [TensorFlow Estimators](https://www.tensorflow.org/extend/estimators)
# * [TensorFlow DataSets](https://github.com/tensorflow/tensorflow/tree/r1.2/tensorflow/contrib/data)
# * [RNNs](http://colah.github.io/posts/2015-08-Understanding-LSTMs/)
#
#
# ## What is colorbot?
#
# Colorbot is a RNN model that receives a word (sequence of characters) as input and learns to predict a rgb value that better represents this word. As a result we have a color generator!
#
# 
# ## Dependencies
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="mDT8S9C9CYtr"
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
# Tensorflow
import tensorflow as tf
print('Expected version: 1.2.0 or higher')
print('Your TensorFlow version:', tf.__version__)
# Feeding function for enqueue data
from tensorflow.python.estimator.inputs.queues import feeding_functions as ff
# Rnn common functions
from tensorflow.contrib.learn.python.learn.estimators import rnn_common
# Run an experiment
from tensorflow.contrib.learn.python.learn import learn_runner
# Model builder
from tensorflow.python.estimator import model_fn as model_fn_lib
# Plot images with pyplot
# %matplotlib inline
from matplotlib import pyplot as plt
# Helpers for data processing
import pandas as pd
import numpy as np
import argparse
# -
# ## Parameters
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="UrAyWt23AtCM"
# Data files
TRAIN_INPUT = 'data/train.csv'
TEST_INPUT = 'data/test.csv'
MY_TEST_INPUT = 'data/mytest.csv'
# Parameters for training
BATCH_SIZE = 64
# Parameters for data processing
VOCAB_SIZE = 256
CHARACTERS = [chr(i) for i in range(VOCAB_SIZE)]
SEQUENCE_LENGTH_KEY = 'sequence_length'
COLOR_NAME_KEY = 'color_name'
# -
# ## Helper functions
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="0dlZ9C27M-bS"
# Returns the column values from a CSV file as a list
def _get_csv_column(csv_file, column_name):
with open(csv_file, 'r') as f:
df = pd.read_csv(f)
return df[column_name].tolist()
# Plots a color image
def _plot_rgb(rgb):
data = [[rgb]]
plt.figure(figsize=(2,2))
plt.imshow(data, interpolation='nearest')
plt.show()
# -
# ## Input function
#
# Here we are defining the input pipeline using the [Dataset API](https://github.com/tensorflow/tensorflow/tree/master/tensorflow/contrib/data).
#
# One more complex transformation that we're using is called **group_by_window**, what this function does is to map each consecutive element in this dataset to a key using `key_func` and then groups the elements by key. It then applies `reduce_func` to at most `window_size` elements matching the same key. All except the final window for each key will contain `window_size` elements; the final window may be smaller.
#
# In the code below what we're doing is using the **group_by_window** transformation to batch color names that have similar length together, this makes the code more efficient since the RNN will be unrroled (approximately) the same number of steps in each batch. This avoids that we waist space and computing time :)!
#
# 
# *Image from [Sequence Models and the RNN API (TensorFlow Dev Summit 2017)](https://www.youtube.com/watch?v=RIR_-Xlbp7s)*
#
# ** *EXERCISE DATASET (first complete the EXERCISE EXPERIMENT: change the input function bellow so it will just use normal padded_batch instead sorting the batches. Then run each model using experiments and compare the efficiency (time, global_step/sec) using TensorBoard.
# hint: to compare the implementations using tensorboard just copy the model_dir folder of both executions to the same directory (the model dir should be different at each time you run the model) and point tensorboard to it with: tensorboard --logdir=path_to_model_dirs_par)* **
def get_input_fn(csv_file, batch_size, num_epochs=1, shuffle=True):
def _parse(line):
# each line: name, red, green, blue
# split line
items = tf.string_split([line],',').values
# get color (r, g, b)
color = tf.string_to_number(items[1:], out_type=tf.float32) / 255.0
# split color_name into a sequence of characters
color_name = tf.string_split([items[0]], '')
length = color_name.indices[-1, 1] + 1 # length = index of last char + 1
color_name = color_name.values
return color, color_name, length
def _length_bin(length, cast_value=5, max_bin_id=10):
'''
Chooses a bin for a word given it's length.
The goal is to use group_by_window to group words
with the ~ same ~ length in the same bin.
Each bin will have the size of a batch, so it can train faster.
'''
bin_id = tf.cast(length / cast_value, dtype=tf.int64)
return tf.minimum(bin_id, max_bin_id)
def _pad_batch(ds, batch_size):
return ds.padded_batch(batch_size,
padded_shapes=([None], [None], []),
padding_values=(0.0, chr(0), tf.cast(0, tf.int64)))
def input_fn():
# https://github.com/tensorflow/tensorflow/tree/master/tensorflow/contrib/data
dataset = (
tf.contrib.data.TextLineDataset(csv_file) # reading from the HD
.skip(1) # skip header
.repeat(num_epochs) # repeat dataset the number of epochs
.map(_parse) # parse text to variables
.group_by_window(key_func=lambda color, color_name, length: _length_bin(length), # choose a bin
reduce_func=lambda key, ds: _pad_batch(ds, batch_size), # apply reduce funtion
window_size=batch_size)
)
# for our "manual" test we don't want to shuffle the data
if shuffle:
dataset = dataset.shuffle(buffer_size=100000)
# create iterator
color, color_name, length = dataset.make_one_shot_iterator().get_next()
features = {
COLOR_NAME_KEY: color_name,
SEQUENCE_LENGTH_KEY: length,
}
return features, color
return input_fn
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="m5UJyvW5P0Sy"
train_input_fn = get_input_fn(TRAIN_INPUT, BATCH_SIZE)
test_input_fn = get_input_fn(TEST_INPUT, BATCH_SIZE)
# -
# Testing the input function
# +
x, y = get_input_fn(TRAIN_INPUT, 1)()
with tf.Session() as s:
print(s.run(x))
print(s.run(y))
# -
# ## Creating the Estimator model
#
# 
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="VxXAUrYN7TvR"
def get_model_fn(rnn_cell_sizes,
label_dimension,
dnn_layer_sizes=[],
optimizer='SGD',
learning_rate=0.01):
def model_fn(features, labels, mode):
color_name = features[COLOR_NAME_KEY]
sequence_length = tf.cast(features[SEQUENCE_LENGTH_KEY], dtype=tf.int32) # int64 -> int32
# ----------- Preparing input --------------------
# Creating a tf constant to hold the map char -> index
mapping = tf.constant(CHARACTERS, name="mapping")
table = tf.contrib.lookup.index_table_from_tensor(mapping, dtype=tf.string)
int_color_name = table.lookup(color_name)
# Converting color names to one hot representation
color_name_onehot = tf.one_hot(int_color_name, depth=len(CHARACTERS) + 1)
# ---------- RNN -------------------
# Each RNN layer will consist of a LSTM cell
rnn_layers = [tf.nn.rnn_cell.LSTMCell(size) for size in rnn_cell_sizes]
# Construct the layers
multi_rnn_cell = tf.nn.rnn_cell.MultiRNNCell(rnn_layers)
# Runs the RNN model dynamically
# more about it at:
# https://www.tensorflow.org/api_docs/python/tf/nn/dynamic_rnn
outputs, final_state = tf.nn.dynamic_rnn(cell=multi_rnn_cell,
inputs=color_name_onehot,
sequence_length=sequence_length,
dtype=tf.float32)
# Slice to keep only the last cell of the RNN
last_activations = rnn_common.select_last_activations(outputs,
sequence_length)
# ------------ Dense layers -------------------
# Construct dense layers on top of the last cell of the RNN
for units in dnn_layer_sizes:
last_activations = tf.layers.dense(
last_activations, units, activation=tf.nn.relu)
# Final dense layer for prediction
predictions = tf.layers.dense(last_activations, label_dimension)
# ----------- Loss and Optimizer ----------------
loss = None
train_op = None
if mode != tf.estimator.ModeKeys.PREDICT:
loss = tf.losses.mean_squared_error(labels, predictions)
if mode == tf.estimator.ModeKeys.TRAIN:
train_op = tf.contrib.layers.optimize_loss(
loss,
tf.contrib.framework.get_global_step(),
optimizer=optimizer,
learning_rate=learning_rate)
return model_fn_lib.EstimatorSpec(mode,
predictions=predictions,
loss=loss,
train_op=train_op)
return model_fn
# -
# ** *EXERCISE HYPERPARAMETERS: try making changes to the model and see if you can improve the results.
# Run the original model, run yours and compare them using Tensorboard. What improvements do you see?
# hint 0: change the type of RNNCell, maybe a GRUCell? Change the number of hidden layers, or add dnn layers.
# hint 1: to compare the implementations using tensorboard just copy the model_dir folder of both executions to the same directory (the model dir should be different at each time you run the model) and point tensorboard to it with: tensorboard --logdir=path_to_model_dirs_par)* **
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="gUHR3Mzc7Tvb"
model_fn = get_model_fn(rnn_cell_sizes=[256, 128], # size of the hidden layers
label_dimension=3, # since is RGB
dnn_layer_sizes=[128], # size of units in the dense layers on top of the RNN
optimizer='Adam', # changing optimizer to Adam
learning_rate=0.01)
estimator = tf.estimator.Estimator(model_fn=model_fn, model_dir='colorbot')
# -
# ## Trainning and Evaluating
#
# ** *EXERCISE EXPERIMENT: The code below works, but we can use an experiment instead. Add a cell that runs an experiment instead of interacting directly with the estimator.
# hint 0: you'll need to change the train_input_fn definition, think about it...
# hint 1: the change is related with the for loop* **
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="DUZEKQrdGgZE"
NUM_EPOCHS = 40
for i in range(NUM_EPOCHS):
print('Training epoch %d' % i)
print('-' * 20)
estimator.train(input_fn=train_input_fn)
print('Evaluating epoch %d' % i)
print('-' * 20)
estimator.evaluate(input_fn = test_input_fn)
# -
# ## Making Predictions
def predict(estimator, input_file):
preds = estimator.predict(input_fn=get_input_fn(input_file, 1, shuffle=False))
color_names = _get_csv_column(input_file, 'name')
print()
for p, name in zip(preds, color_names):
color = tuple(map(int, p * 255))
print(name + ',', 'rgb:', color)
_plot_rgb(p)
predict(estimator, MY_TEST_INPUT)
# ## Pre-trained model predictions
#
# In order to load the pre-trained model we can just create an estimator using the model_fn and use the model_dir that contains the pre-trained model files in this case it's 'pretrained'
pre_estimator = tf.estimator.Estimator(model_dir='pretrained', model_fn=model_fn)
predict(pre_estimator, MY_TEST_INPUT)
# # Colorbot Solutions
#
# Here are the solutions to the exercises available at the colorbot notebook.
#
# In order to compare the models we encourage you to use Tensorboard and also use play_colorbot.py --model_dir=path_to_your_model to play with the models and check how it does with general words other than color words.
# ## *EXERCISE EXPERIMENT*
#
# When using experiments you should make sure you repeat the datasets the number of epochs desired since the experiment will "run the for loop for you". Also, you can add a parameter to run a number of steps instead, it will run until the dataset ends or the number of steps.
#
# You can add this cell to your colorbot notebook and run it.
# +
# small important detail, to train properly with the experiment you need to
# repeat the dataset the number of epochs desired
train_input_fn = get_input_fn(TRAIN_INPUT, BATCH_SIZE, num_epochs=40)
# create experiment
def generate_experiment_fn(run_config, hparams):
estimator = tf.estimator.Estimator(model_fn=model_fn, config=run_config)
return tf.contrib.learn.Experiment(
estimator,
train_input_fn=train_input_fn,
eval_input_fn=test_input_fn
)
learn_runner.run(generate_experiment_fn, run_config=tf.contrib.learn.RunConfig(model_dir='model_dir'))
# -
# ## *EXERCISE DATASET*
#
# 0. Run the colorbot experiment and notice the choosen model_dir
# 1. Below is the input function definition,we don't need some of the auxiliar functions anymore
# 2. Add this cell and then add the solution to the EXERCISE EXPERIMENT
# 3. choose a different model_dir and run the cells
# 4. Copy the model_dir of the two models to the same path
# 5. tensorboard --logdir=path
def get_input_fn(csv_file, batch_size, num_epochs=1, shuffle=True):
def _parse(line):
# each line: name, red, green, blue
# split line
items = tf.string_split([line],',').values
# get color (r, g, b)
color = tf.string_to_number(items[1:], out_type=tf.float32) / 255.0
# split color_name into a sequence of characters
color_name = tf.string_split([items[0]], '')
length = color_name.indices[-1, 1] + 1 # length = index of last char + 1
color_name = color_name.values
return color, color_name, length
def input_fn():
# https://github.com/tensorflow/tensorflow/tree/master/tensorflow/contrib/data
dataset = (
tf.contrib.data.TextLineDataset(csv_file) # reading from the HD
.skip(1) # skip header
.map(_parse) # parse text to variables
.padded_batch(batch_size, padded_shapes=([None], [None], []),
padding_values=(0.0, chr(0), tf.cast(0, tf.int64)))
.repeat(num_epochs) # repeat dataset the number of epochs
)
# for our "manual" test we don't want to shuffle the data
if shuffle:
dataset = dataset.shuffle(buffer_size=100000)
# create iterator
color, color_name, length = dataset.make_one_shot_iterator().get_next()
features = {
COLOR_NAME_KEY: color_name,
SEQUENCE_LENGTH_KEY: length,
}
return features, color
return input_fn
# As a result you will see something like:
#
# 
#
# We called the original model "sorted_batch" and the model using the simplified input function as "simple_batch"
#
# Notice that both models have basically the same loss in the last step, but the "sorted_batch" model runs way faster , notice the `global_step/sec` metric, it measures how many steps the model executes per second. Since the "sorted_batch" has a larger `global_step/sec` it means it trains faster.
#
# If you don't belive me you can change Tensorboard to compare the models in a "relative" way, this will compare the models over time. See result below.
#
# 
# ## *EXERCISE HYPERPARAMETERS*
#
# This one is more personal, what you see will depends on what you change in the model.
# Below is a very simple example we just changed the model to use a GRUCell, just in case...
def get_model_fn(rnn_cell_sizes,
label_dimension,
dnn_layer_sizes=[],
optimizer='SGD',
learning_rate=0.01):
def model_fn(features, labels, mode):
color_name = features[COLOR_NAME_KEY]
sequence_length = tf.cast(features[SEQUENCE_LENGTH_KEY], dtype=tf.int32) # int64 -> int32
# ----------- Preparing input --------------------
# Creating a tf constant to hold the map char -> index
# this is need to create the sparse tensor and after the one hot encode
mapping = tf.constant(CHARACTERS, name="mapping")
table = tf.contrib.lookup.index_table_from_tensor(mapping, dtype=tf.string)
int_color_name = table.lookup(color_name)
# representing colornames with one hot representation
color_name_onehot = tf.one_hot(int_color_name, depth=len(CHARACTERS) + 1)
# ---------- RNN -------------------
# Each RNN layer will consist of a GRU cell
rnn_layers = [tf.nn.rnn_cell.GRUCell(size) for size in rnn_cell_sizes]
# Construct the layers
multi_rnn_cell = tf.nn.rnn_cell.MultiRNNCell(rnn_layers)
# Runs the RNN model dynamically
# more about it at:
# https://www.tensorflow.org/api_docs/python/tf/nn/dynamic_rnn
outputs, final_state = tf.nn.dynamic_rnn(cell=multi_rnn_cell,
inputs=color_name_onehot,
sequence_length=sequence_length,
dtype=tf.float32)
# Slice to keep only the last cell of the RNN
last_activations = rnn_common.select_last_activations(outputs,
sequence_length)
# ------------ Dense layers -------------------
# Construct dense layers on top of the last cell of the RNN
for units in dnn_layer_sizes:
last_activations = tf.layers.dense(
last_activations, units, activation=tf.nn.relu)
# Final dense layer for prediction
predictions = tf.layers.dense(last_activations, label_dimension)
# ----------- Loss and Optimizer ----------------
loss = None
train_op = None
if mode != tf.estimator.ModeKeys.PREDICT:
loss = tf.losses.mean_squared_error(labels, predictions)
if mode == tf.estimator.ModeKeys.TRAIN:
train_op = tf.contrib.layers.optimize_loss(
loss,
tf.contrib.framework.get_global_step(),
optimizer=optimizer,
learning_rate=learning_rate)
return model_fn_lib.EstimatorSpec(mode,
predictions=predictions,
loss=loss,
train_op=train_op)
return model_fn
| archive/extras/colorbot/colorbot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pylab inline
# %config InlineBackend.figure_format = 'retina'
from ipywidgets import interact
# # Brownian motion: First passage time to exit the interval (0, L)
# Assume 1D Brownian motion with diffusivity $D>0$.
# Consider the mean first passage time $T(x_0)$, which satisfies
# $$ DT'' = -1,\qquad x_0\in (0, L)$$
# with boundary conditions
# $$T(0) = T(L) = 0.$$
# The solution is given by
# $$T(x_0) = \frac{x_0(L-x_0)}{2D}$$
# +
D = 2
sigma = sqrt(2.*D)
L = 50.
x0 = L/3. # must be between zero and L
def first_passage_time_FAST(): ## this is a faster version
dt = 0.1
dW_stdev = sigma*sqrt(dt)
Nbuffer = 10000
# Z = zeros(Nsteps) # we do not need to store the entire path
# Z[0] = z0
x = x0
t = 0
j = 0
while True: # this will simply loop forever unless otherwise stopped in the loop
## if a bug ever causes a loop to continue forever, you can halt the process by
## pushing the 'interupt' button (looks like a stop button in the toolbar)
if j%Nbuffer == 0:
dW = normal(0, dW_stdev, Nbuffer)
j = 0
x += dW[j]
t += dt
j += 1
## check to see if x=0 or x=L has been reached
if x <= 0 or x >= L:
return t ## this stops the loop and returns the time
## we want to collect many samples of the first passage time in an array
Nsamples = 1000
## Way 1: get samples using a for loop
# T = zeros(Nsamples) ## to store samples of first passage time
# for n in arange(Nsamples):
# T[n] = first_passage_time_FAST()
## Way 2: this is a short hand way of computing T, same as the three lines in Way 1
T = array([first_passage_time_FAST() for n in arange(Nsamples)])
## Figure
figure(1, [6, 4])
hist(T, bins=20)
xlabel(r'$\tau$', fontsize=24)
ylabel('count', fontsize=24);
print('Mean first passage time (simulations):', T.mean())
print('Mean first passage time (exact):', x0*(L-x0)/(2*D))
# -
# # Brownian motion: First passage time to exit the interval $(0, L)$ through $x=L$ (with $x=0$ reflecting)
# Assume 1D Brownian motion with diffusivity $D>0$.
# Consider the mean first passage time $T(x_0)$, which satisfies
# $$ DT'' = -1,\qquad x_0\in (0, L)$$
# with boundary conditions
# $$T'(0) = T(L) = 0.$$
# The solution is given by
# $$T(x_0) = \frac{L^2 - x_0^2}{2D}.$$
# +
D = 2
sigma = sqrt(2.*D)
L = 50.
x0 = 0.7*L # must be between zero and L
def first_passage_time_FAST(): ## this is a faster version
dt = 0.1
dW_stdev = sigma*sqrt(dt)
Nbuffer = 10000
# Z = zeros(Nsteps) # we do not need to store the entire path
# Z[0] = z0
x = x0
t = 0
j = 0
while True: # this will simply loop forever unless otherwise stopped in the loop
## if a bug ever causes a loop to continue forever, you can halt the process by
## pushing the 'interupt' button (looks like a stop button in the toolbar)
if j%Nbuffer == 0:
dW = normal(0, dW_stdev, Nbuffer)
j = 0
x += dW[j]
t += dt
j += 1
## reflecting boundary at x=0
if x < 0:
x = -x
## check to see if x=L has been reached
if x <= 0 or x >= L:
return t ## this stops the loop and returns the time
## we want to collect many samples of the first passage time in an array
Nsamples = 1000
## Way 1: get samples using a for loop
# T = zeros(Nsamples) ## to store samples of first passage time
# for n in arange(Nsamples):
# T[n] = first_passage_time_FAST()
## Way 2: this is a short hand way of computing T, same as the three lines in Way 1
T = array([first_passage_time_FAST() for n in arange(Nsamples)])
## Figure
figure(1, [6, 4])
hist(T, bins=20)
xlabel(r'$\tau$', fontsize=24)
ylabel('count', fontsize=24);
print('Mean first passage time (simulations):', T.mean())
print('Mean first passage time (exact):', (L**2 - x0**2)/(2*D))
| Week 10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Data loading and churn label creation
# The raw data contains all the users' activities from March 30 to May 12, 2017. (Fresh data!!) Since the data amount is so huge for a PC, reading all data into memory is a mission impossible. On the other hand, it's not necessary to get all the activity information to determine if a user is a churn. To be more specific, whether one has activity is important, while the detail of the activities shall not be considered for the moment.
# ### Procedure:
#
# 1. Unzip all the "play" activity data in a batch.
#
# 2. Choose a cut-off date for the churn labeling: Any user who is active before this cut-off date but has no activity after the date shall be labeled as a churn.
#
# 3. Read only "user_id","date/time" of each activity log before and after the cut-off date, and save them in two new file respectively.
#
# 4. Get two sets of active users, before and after the cut-off date: {active_before}, {active_after}
#
# 5. Output the churns to a .txt file. {Churn} = {active_before} - {active_before} & {active_after}
#
# ##### 1. Unzip the 400+ .tar.gz compressed files of raw data
#
# 7z can only unzip the .tar.gz files to .tar, so the complete unzip takes two steps:
#
# 1. Batch unzip from .tar.gz --> .tar
# 2. Batch unzip from .tar --> log files
## In windows powershell, run the following iteration commands in the raw data directory:
'''
$files = Get-ChildItem "../data/raw/" -Filter *_play.log.tar.gz
foreach ($f in $files) {7z e $f -oC../data/raw/unzip}
cd unzip
$files = Get-ChildItem "../raw/unzip" -Filter *_play.log.tar
foreach ($f in $files) {7z e $f}
'''
# References:
#
# __[for loop in windows powershell](https://stackoverflow.com/questions/18847145/loop-through-files-in-a-directory-using-powershell)__
#
#
# __[Unzip .tar.gz files using 7z commands](https://stackoverflow.com/questions/1359793/programmatically-extract-tar-gz-in-a-single-step-on-windows-with-7zip)__
# ##### 2. Cut-off date = April 21st.
#
# So that there are three weeks before cut-off, and three weeks after. The cut off might need changing later, depending on the model's performance.
# ##### 3. File operations on the play logs
#
# 1. Open *play.log files one by one
#
# 2. "Read - Append" for each line
#
# 3. Write the new line to a new file for all lines of all files.
#
# 4. Meawhile, extract the user_id (first item) in each line, and save them into two python sets for logs before and after the "snapshot date"
#
# 5. Save the sets into .log files.
# 1. Open all play logs, using * wildcard
# +
import glob
filepath = 'C:\\Users\\Sean\\Documents\\BitTiger\\Capston_music_player_python\\*play.log'
files = glob.glob(filepath)
len(files)
# -
files
i, log_amounts = 0, []
for the_file in files:
f = open(the_file, 'r')
lines = f.readlines()
log_amounts.append(len(lines))
f.close()
log_amounts
# 2. Read the files, append the date to each line
with open('C:\\Users\\Sean\\Documents\\BitTiger\\Capston_music_player_python\\20170512_3_play.log','r') as f:
content = f.readlines()
len(content)
first_line = content[0]
first_line
first_line_fields = content[0].strip('\n').split('\t')
#first_line_fields.append(f.name.split('\\')[-1][8])
first_line_fields
first_line_fields.append(f.name.split('\\')[-1][:8])
'\t'.join(first_line_fields)
new_line = '\t'.join(first_line_fields) + '\n'
new_line
# 3. write into a new file
#
# Using the f.write() method
output = open('C:\\Users\\Sean\\Documents\\BitTiger\\Capston_music_player_python\\output\\all.log','a')
# +
for the_file in files:
with open(the_file, 'r') as f:
lines = f.readlines()
for line in lines:
contents = line.strip('\n').split('\t')
contents.append(f.name.split('\\')[-1][:8])
output.write('\t'.join(contents)+'\n')
output.close()
# -
with open('C:\\Users\\Sean\\Documents\\BitTiger\\Capston_music_player_python\\output\\all.log','r') as output:
lines = output.readlines()
len(lines)
sum(log_amounts)
# 6.
# +
from sets import Set
list_of_sets = []
# -
with open('C:\\Users\\Sean\\Documents\\BitTiger\\Capston_music_player_python\\output\\all.log','a') as output:
for the_file in files:
with open(the_file, 'r') as f:
lines = f.readlines()
list_of_sets.append(Set([line.split('\t')[0] for line in lines]))
for line in lines:
contents = line.strip('\n').split('\t')
contents.append(f.name.split('\\')[-1][:8])
output.write('\t'.join(contents)+'\n')
[len(each_set) for each_set in list_of_sets]
# 5. Churn labeling and file saving
#
# Save the user_id of churns into a new file.
active_before, active_after = list_of_sets[0],list_of_sets[1]
loyal_users = active_before.intersection(active_after)
len(loyal_users)
churn = active_before.difference(active_after)
len(churn)
new_users = active_after.difference(active_before)
len(new_users)
# +
# Use loyal_user as an example
with open('C:\\Users\\Sean\\Documents\\BitTiger\\Capston_music_player_python\\loyal.log','a') as loyal_file:
loyal_file.write('\n'.join(list(loyal_users))+'\n')
# +
# check loyal_file
with open('C:\\Users\\Sean\\Documents\\BitTiger\\Capston_music_player_python\\loyal.log','r') as loyal_file:
lines = loyal_file.readlines()
lines
# -
| .ipynb_checkpoints/Data_loading_and_churn_label_creation-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="CRNVzhoo1clG" colab_type="text"
# ## **1. Mount google drive**
# ---
# + id="5W39GXyk1hME" colab_type="code" colab={}
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="dOQ-2xS_MYHi" colab_type="text"
# ## **2. Import the necessary libraries**
# ---
# + id="yoi4gWDELtek" colab_type="code" colab={}
import matplotlib
import sklearn
import numpy as np
import pandas as pd
import sklearn.metrics as metrics
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow.keras.callbacks import ModelCheckpoint,CSVLogger,LearningRateScheduler
from tensorflow.keras.models import Model
from tensorflow.keras.layers import Input
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import Flatten
from tensorflow.keras.layers import Conv2D
from tensorflow.keras.layers import BatchNormalization
from tensorflow.keras.layers import Activation
from tensorflow.keras.layers import AveragePooling2D
from tensorflow.keras.layers import add
from tensorflow.keras.regularizers import l2
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.datasets import cifar10
from tensorflow.keras import optimizers
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.utils import plot_model
print("Versions of key libraries")
print("---")
print("tensorflow: ", tf.__version__)
print("numpy: ", np.__version__)
print("matplotlib: ", matplotlib.__version__)
print("sklearn: ", sklearn.__version__)
# + [markdown] id="0amhwjZ4M2m-" colab_type="text"
# ## **3.Create a function to plot image without axis**
# ---
# + id="OFxsBB1mNXl2" colab_type="code" colab={}
def implt(img):
plt.figure()
plt.imshow(img)
plt.axis('off')
print(implt)
# + [markdown] id="X3N41iixORPz" colab_type="text"
# ## **4. Set matplotlib to have seaborn plot style**
# ---
# + id="OyO5OsUrOYNQ" colab_type="code" colab={}
plt.style.use('seaborn') # if want to use the default style, set 'classic'
plt.rcParams['ytick.right'] = True
plt.rcParams['ytick.labelright']= True
plt.rcParams['ytick.left'] = False
plt.rcParams['ytick.labelleft'] = False
plt.rcParams['figure.figsize'] = [7,7] # Set the figure size to be 7 inch for (width,height)
print("Matplotlib setup completes.")
# + [markdown] id="5w2jAiKZOmgP" colab_type="text"
# ## **5. Prepare Cifar10 data for training and testing**
# ---
# * Step 1: Load the cifar10
# * Step 2: Check the shape and type of the data
# * Step 3: Convert the data into float32 and rescale the values from the range of 0\~255 into 0\~1
# * Step 4: Retrieve the row size and the column size of each image
# * Step 5: Perform one-hot enconding on the labels
# * Step 6: Retrieve the number of classes in this problem
# + id="v3Ad2V0pO1TX" colab_type="code" colab={}
# Step 1
data = cifar10.load_data()
(trDat, trLbl) = data[0]
(tsDat, tsLbl) = data[1]
# Step 2
print("The shape of trDat is", trDat.shape, "and the type of trDat is", trDat.dtype)
print("The shape of tsDat is", tsDat.shape, "and the type of tsDat is", tsDat.dtype)
print("")
print("The shape of trLbl is", trLbl.shape, "and the type of trLbl is", trLbl.dtype)
print("The shape of tsLbl is", tsLbl.shape, "and the type of tsLbl is", tsLbl.dtype)
# Step 3
trDat = trDat.astype('float32')/255
tsDat = tsDat.astype('float32')/255
# Step 4
imgrows = trDat.shape[1]
imgclms = trDat.shape[2]
channel = trDat.shape[3]
# Step 5
trLbl = to_categorical(trLbl)
tsLbl = to_categorical(tsLbl)
num_classes = tsLbl.shape[1] # Step 6
# + [markdown] id="DoP3WcoJW-jZ" colab_type="text"
# ## **6. Define the resnet model (to be completed)**
# ___
# * Step 1: Setup the optimizer to be used for training
# * Step 2: Set a name for the coming model (required for saving)
# * Step 3: Function to create layers for the resnet
# * Step 4: Function to create residual blocks
# * Step 5: Define the resnet model (to be completed)
# * Step 6: Create models for training and testing
# * Step 7: Display the summary of the model of interest
# + id="0HMOes0kXCPd" colab_type="code" colab={}
optmz = optimizers.Adam(lr=0.001) # Step 1
modelname = 'cifar10ResV1Cfg5' # Step 2
# Step 3
def resLyr(inputs,
numFilters=16,
kernelSz=3,
strides=1,
activation='relu',
batchNorm=True,
convFirst=True,
lyrName=None):
convLyr = Conv2D(numFilters,
kernel_size=kernelSz,
strides=strides,
padding='same',
kernel_initializer='he_normal',
kernel_regularizer=l2(1e-4),
name=lyrName+'_conv' if lyrName else None)
x = inputs
if convFirst:
x = convLyr(x)
if batchNorm:
x = BatchNormalization(name=lyrName+'_bn' if lyrName else None)(x)
if activation is not None:
x = Activation(activation,
name=lyrName+'_'+activation if lyrName else None)(x)
else:
if batchNorm:
x = BatchNormalization(name=lyrName+'_bn' if lyrName else None)(x)
if activation is not None:
x = Activation(activation,
name=lyrName+'_'+activation if lyrName else None)(x)
x = convLyr(x)
return x
# Step 4
def resBlkV1(inputs,
numFilters=16,
numBlocks=3,
downsampleOnFirst=True,
names=None):
x = inputs
for run in range(0,numBlocks):
strides = 1
blkStr = str(run+1)
if downsampleOnFirst and run == 0:
strides = 2
y = resLyr(inputs=x,
numFilters=numFilters,
strides=strides,
lyrName=names+'_Blk'+blkStr+'_Res1' if names else None)
y = resLyr(inputs=y,
numFilters=numFilters,
activation=None,
lyrName=names+'_Blk'+blkStr+'_Res2' if names else None)
if downsampleOnFirst and run == 0:
x = resLyr(inputs=x,
numFilters=numFilters,
kernelSz=1,
strides=strides,
activation=None,
batchNorm=False,
lyrName=names+'_Blk'+blkStr+'_lin' if names else None)
x = add([x,y],
name=names+'_Blk'+blkStr+'_add' if names else None)
x = Activation('relu',
name=names+'_Blk'+blkStr+'_relu' if names else None)(x)
return x
# Step 5
def createResNetV1(inputShape=(32,32,3),
numClasses=10):
return model
# Step 6
model = createResNetV1() # This is meant for training
modelGo = createResNetV1() # This is used for final testing
model.summary() # Step 7
# + [markdown] id="DlquJEaFZxV9" colab_type="text"
# ## **7. Create the callbacks to be applied during training**
# ---
# * Step 1: Create a callback to save the model from an epoch when validation accuracy is the highest
# * Step 2: Create a callback to save the training loss, training accuracy, validation loss and validation accuracy of each epoch into a csv file
# * Step 3: Put the two callback objects into a list
# + id="9-a1LSCbahKy" colab_type="code" colab={}
# Step 1
def lrSchedule(epoch):
lr = 1e-3
if epoch > 160:
lr *= 0.5e-3
elif epoch > 140:
lr *= 1e-3
elif epoch > 120:
lr *= 1e-2
elif epoch > 80:
lr *= 1e-1
print('Learning rate: ', lr)
return lr
LRScheduler = LearningRateScheduler(lrSchedule)
# Step 2
folderpath = '/content/gdrive/My Drive/iss/prumls/colab/'
filepath = folderpath + modelname + ".hdf5"
checkpoint = ModelCheckpoint(filepath,
monitor='val_accuracy',
verbose=0,
save_best_only=True,
mode='max')
csv_logger = CSVLogger(folderpath+modelname +'.csv') # Step 2
callbacks_list = [checkpoint,csv_logger,LRScheduler] # Step 3
print("Callbacks created:")
print(callbacks_list[0])
print(callbacks_list[1])
print(callbacks_list[2])
print('')
print("Path to model:", filepath)
print("Path to log: ", folderpath+modelname+'.csv')
# + [markdown] id="1mKgjQsmfOBz" colab_type="text"
# ## **8. Train the deep learning model with image augmentation (to be completed)**
# ___
# * Step 1: Create the image data generator (for image augmentation)
# * Step 2: Train the model with generator
# + id="23lUNwpGfV0A" colab_type="code" colab={}
# Step 1
# Step 2
# + [markdown] id="TevfJTd-s0nk" colab_type="text"
# ## **9. Validate the deep learning model**
# ---
# * Step 1: Load the trained weights and compile the model
# * Step 2: Make prediction
#
# + id="2sVtWmcVtiV5" colab_type="code" colab={}
# Step 1
modelGo.load_weights(filepath)
modelGo.compile(loss='categorical_crossentropy',
optimizer=optmz,
metrics=['accuracy'])
predicts = modelGo.predict(tsDat) # Step 2
print("Prediction completes.")
# + [markdown] id="0aOCUljp5qq4" colab_type="text"
# ## **10. Report classification metrics**
# ---
# * Step 1: Setup the label
# * Step 2: Convert label from one-hot to integer
# * Step 3: Calculate the accuracy score
# * Step 4: Generate classification report
# + id="2tI4hBmk5uRh" colab_type="code" colab={}
# Step 1
labelname = ['airplane', # The label for reporting metrics
'automobile',
'bird',
'cat',
'deer',
'dog',
'frog',
'horse',
'ship',
'truck']
# Step 2
predout = np.argmax(predicts,axis=1)
testout = np.argmax(tsLbl,axis=1)
testScores = metrics.accuracy_score(testout,predout) # Step 3
# Step 4
print("Best accuracy (on testing dataset): %.2f%%" % (testScores*100))
print(metrics.classification_report(testout,
predout,
target_names=labelname,
digits=4))
# + [markdown] id="gEK_4UXN6IVa" colab_type="text"
# ## **11. Print confusion matrix**
# ---
# + id="UCBJCYp26L1t" colab_type="code" colab={}
confusion = metrics.confusion_matrix(testout,predout)
print(confusion)
# + [markdown] id="2QMIDPD46UGT" colab_type="text"
# ## **12. Plot curves on validation loss and accuracy**
# ---
# + id="qr2ZbvUi6YHf" colab_type="code" colab={}
records = pd.read_csv(folderpath+modelname +'.csv')
plt.figure()
plt.subplot(211)
plt.plot(records['val_loss'], label="validation")
plt.plot(records['loss'],label="training")
plt.yticks([0.00,0.50,1.00,1.50])
plt.title('Loss value',fontsize=12)
ax = plt.gca()
ax.set_xticklabels([])
plt.subplot(212)
plt.plot(records['val_accuracy'],label="validation")
plt.plot(records['accuracy'],label="training")
plt.yticks([0.5,0.6,0.7,0.8])
plt.title('Accuracy',fontsize=12)
ax.legend()
plt.show()
# + [markdown] id="PWoTz-bLug3X" colab_type="text"
# ## **13. Save the model plot**
# ---
# + id="Tz1YfuV1ujcE" colab_type="code" colab={}
plotpath = folderpath+modelname+'_plot.png'
plot_model(model,
to_file=plotpath,
show_shapes=True,
show_layer_names=False,
rankdir='TB')
print("Path to plot:", plotpath)
| Sessions/SESSION_5/WORKSHOPS/workshops/prumls-wks4-2-_yourname_.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# <a id='back'></a>
# # Statistical Tests
#
# ### Intro
#
# ### Table of Contents
# * <a href='#samplez'>Large Sample z-test for a population proportion</a>
# * <a href='#samplez_hype'>Hypotheis Test</a>
# * <a href='#samplez_prop'>prop.test function in R</a>
#
#
# * <a href='#samplez_diff'>Large Sample z-test for Difference in Proportion</a>
# * <a href='#hype_samplez_diff'>Hypotheis Test</a>
# * <a href='#prop_samplez_diff'>prop.test function</a>
#
#
# * <a href='#samp_mean'>One sample t-test for population mean</a>
# * <a href='#samp_mean_hype'>Hypothesis test</a>
#
#
# * <a href='#two_samp'>Two-sample tests</a>
# * <a href='#two_samp_ue'>Two sample independent t-test for unequal variance</a>
# * <a href='#two_samp_ue_hype'>Hypothesis Test</a>
#
# * <a href='#paired'>Paired t-test</a>
#
#
# * <a href='#boot'>Bootstrap confidence intervals</a>
#
#
# * <a href='#rand'>Randomization tests</a>
#
#
# * <a href='#anova'>One-Way ANOVA</a>
# * <a href='#anova'>ANOVA Hyptothesis Testing</a>
# * <a href='#tukey'>Tukey HSD</a>
# * <a href='#anova_val'>Checking ANOVA assumptions are met</a>
# * <a href='#anova_res'>Results of ANOVA</a>
#
#
# * <a href='#two_anova'>Two-Way ANOVA</a>
#
#
# * <a href='#ancova'>ANCOVA</a>
#
#
# * <a href='#lm'>Linear Regression</a>
#
#
# * <a href='#logistic'>Logistic Regression</a>
# Packages to load
library(tidyverse)
library(Lock5Data)
library(car)
library(ISLR)
# <a id='samplez'></a>
# ## Large Sample z-test for a population proportion
#
# The data set we are looking at is the ChickWeight data set from the datasets package. The columns are:
#
# weight - a numeric vector giving the body weight of the chick (gm)
#
# Time - a numeric vector giving the number of days since birth when the measurement was made.
#
# Chick - an ordered factor with levels 18 < ... < 48 giving a unique identifier for the chick. The ordering of the levels groups chicks on the same diet together and orders them according to their final weight (lightest to heaviest) within diet.
#
# Diet - a factor with levels 1, ..., 4 indicating which experimental diet the chick received.
#
# The problem that we are going to be tackling is that in a sample of 220 chicks, after going through diet 1 it was found that a certain number of chicks had a weight greater than 130. The people that provided the diet for the farmer claims that more than 30% of the chicks will weight greater than 130 gm. Find the amount of chicks with a weight greater than 130 and using a significance level equal to .05, test whether this claim is accurate or not.
#Load in the data and look over a little bit of the data
data(ChickWeight)
head(ChickWeight)
str(ChickWeight)
#Filter the data so we only are using chicks that went through diet 1 and chicks greater than 130
chick <- filter(ChickWeight, Diet == 1)
chick130 <- filter(ChickWeight, Diet == 1 & weight > 130)
sample_size <- nrow(chick)
greater130_size <- nrow(chick130)
alpha <- .05
prop <- .30
test_prop <- greater130_size / sample_size
test_stat <- (test_prop - prop) / sqrt((prop * (1 - prop)) / sample_size)
p_value <- 1 - pnorm(test_stat)
conf_int <- c((test_prop - qnorm(1 - (alpha / 2)) * sqrt((test_prop * (1 - test_prop)) / sample_size)), (test_prop + qnorm(1 - (alpha / 2)) * sqrt((test_prop * (1 - test_prop)) / sample_size)))
# Results
test_prop
test_stat
p_value
conf_int
# <a id='samplez_hype'></a>
# ### Hypotheis Test for Large Sample z-test for a population proportion
#
# #### Assumptions:
# For this test, observations $x_{1}$, . . . , $x_{n}$ (a sequence of 0’s and 1’s) are a
# random sample from Bern(p) with p unknown and n is equal to the sample size.
#
# The sample size n is large enough to ensure that n$p_{0}$ ≥ 15 and
# n(1 − $p_{0}$) ≥ 15
#
# #### Hypothesis:
# $$H_{0}: p = .30$$
#
# $$H_{a}: p > .30$$
#
#
# #### Test Statistic:
#
# $$z=\frac{\hat{p}-p_{0}}{\sqrt{\frac{p_{0}(1-p_{0})}{n}}}$$
#
# $$z=-1.0298$$
#
# #### P_value:
# $$p-value = 0.848461501351687$$
#
# #### Conclusion:
# We fail to reject the null hypothesis at significance level of .05. This however does not mean we accept the alternative hypothesis and more testing need to be done
#
# #### Confidence Interval:
#
# $$\hat{p}\pm z_{(1-\alpha/2)}\sqrt{\frac{\hat{p}(1-\hat{p})}{n}}$$
#
# For a 95% confidence interval, we are 95% confident that p is in the interval (0.209641778346864, 0.326721858016773), however, the probability that p is in this interval is either 0 or 1. This means that if we were to do this experiment over and over again we are confident that 95% of the time the true population proportion is in this range.
# <a id='samplez_prop'></a>
# ## Using R's prop.test
#
# We can also use R's built in prop.test function if we know the probability ahead of time. The example we will use this time is tossing a coin. Lets say you toss the coin 500 times and it only landed heads 200 times. Test at a .05 significance level if the coin is fair or in other words if the probability of landing heads is not 50%.
## Prop.test
prop.test(x = 200, n = 500, p = 0.5, alternative = "two.sided", conf.level = .95)
# #### Explanation of the results
#
# From the prop.test we get a nice layout of the results for the test. We see alternative hypothesis, p-value and the confidence interval as well. From this we can see that the p-value is much less than the .05 significance level and thus we reject the null hypothesis, and we conclude that there is enough statistical evidence to infer that the alternative hypothesis is true.
# <a id='samplez_diff'></a>
# ## Large Sample z-test for Difference in Proportion
# The dataset we are looking at is the StatGrades dataset found in the Lock5Data package and it contains Stats test scores. The columns are:
#
# Exam1: Score (out of 100 points) on the first exam
#
# Exam2: Score (out of 100 points) on the second exam
#
# Final: Score (out of 100 points) on the final exam
#
# The problem that we are going to answer is if there is a greater chance that you will pass the first exam then the second exam where a passing grade is greater than or equal to 75.
#Look over the data
data(StatGrades)
head(StatGrades)
str(StatGrades)
summary(StatGrades)
#Filter data
alpha <- .05
exam1_pass <- StatGrades %>% filter(Exam1 >= 75) %>% select(Exam1)
exam2_pass <- StatGrades %>% filter(Exam2 >= 75) %>% select(Exam1)
size_pass1 <- nrow(exam1_pass)
size_pass2 <- nrow(exam2_pass)
samp_size <- nrow(StatGrades)
samp_prop1 <- size_pass1 / samp_size
samp_prop2 <- size_pass2 / samp_size
p_hat <- (size_pass1*samp_prop1 + size_pass2*samp_prop2)/(size_pass1 +size_pass2)
test_stat <- (samp_prop1 - samp_prop2) / sqrt(p_hat*(1-p_hat)*((1/size_pass1)+(1/size_pass2)))
p_value <- 1 - pnorm(test_stat)
conf_int2 <- c((samp_prop1 - samp_prop2) - qnorm(1 - (alpha/2))*sqrt(((samp_prop1*(1-samp_prop1))/size_pass1) + ((samp_prop2*(1-samp_prop2))/size_pass2)), (samp_prop1 - samp_prop2) + qnorm(1 - (alpha/2))*sqrt(((samp_prop1*(1-samp_prop1))/size_pass1) + ((samp_prop2*(1-samp_prop2))/size_pass2)))
conf_int <- c((samp_prop1 - samp_prop2) - qnorm(1 - (alpha/2))*sqrt(p_hat*(1-p_hat)*((1/size_pass1)+(1/size_pass2))), (samp_prop1 - samp_prop2) + qnorm(1 - (alpha/2))*sqrt(p_hat*(1-p_hat)*((1/size_pass1)+(1/size_pass2))))
#Results
test_stat
p_value
conf_int
conf_int2
# <a id='hype_samplez_diff'></a>
# ### Hypotheis Test for Large Sample z-test for difference in proportions
#
# #### Assumptions:
# $x_{11}$, . . . , $x_{1n_{1}}$ and $x_{21}$, . . . , $x_{2n_{2}}$
# are random samples from two independent Bernoulli populations Bern($p_{1}$) and Bern($p_{2}$)
# respectively with at least 10 successes and 10 failures in both groups.
#
# $$ n_{1}\hat{p_{1}}\geq 10,\quad n_{1}(1−\hat{p_{1}})\geq 10 \quad and \quad n_{2}\hat{p_{2}}\geq 10,\quad n_{2}(1−\hat{p_{2}})\geq 10$$
#
# #### Hypothesis
# $$H_{0}: p_{1} - p_{2} = 0$$
#
# $$H_{a}: p_{1} - p_{2} > 0$$
#
# #### Test Statistic
#
# $$z=\frac{\hat{p_{1}}-\hat{p_{2}}-0}{\sqrt{\hat{p}(1 - \hat{p})(1/n_{1} + 1/n_{2})}}$$
#
# where $\hat{p} = (n_{1}\hat{p_{1}} + n_{2}\hat{p_{2}})/(n_{1} + n_{2})$
#
# $$ z= -0.471763853242795$$
#
# #### P-value
#
# $$p-value = 0.68145232283435$$
#
# #### Conclusion
#
# Because the p-value is found to be greater than the .05 significance level, we fail to reject the null hypothesis and we can not conclude whether or not the their is a higher chance to pass the first exam compared to the second one.
#
# #### Confidence Interval
#
# $$(\hat{p_{1}}-\hat{p_{2}})\pm z_{(1-\alpha/2)}SE$$
#
# Where SE is the denominator of the test statistics (didn't want to write the Latex of it :p)
# <a id='prop_samplez_diff'></a>
# ## Using prop.test
#
# Just like in the large sample z test for population proportion we can do most of these calculation using prop.test except this time the parameters will be taking a list of the proportions and also each of the proportions sample size. We will have a nicely laid out example with all the values we need. Lets say we want to see if mean and women are equally likely to go to a a 4 year college after high school. 12000 people are sampled where 5622 are men and 6378 are women and out of those men 3004 went to college while 4234 went to college on the womens side. Test whether this claim is true.
men_coll <- 3004
women_coll <- 4234
samp_men <- 5622
samp_women <- 6378
prop.test(x = c(men_coll, women_coll), n = c(samp_men, samp_women), alternative = 'two.sided')
# #### Results
# From this we can now make a conclusion that with p-value less than the .05 significance value, we can reject the null hypothesis and conclude that there is statistical evidence towards the two population proportions not being equal. In other words males and females seem to not have the same proportion of going to college.
# <a id='samp_mean'></a>
# #### Disclaimer
# In the following sections we will be focusing more on the functions and will not be writing out the formulas anymore (I got tired writing so much Latex haha). If you want to know the formulas pleas google them.
# ## One sample t-test for a population mean
#
# The data set we will be using is the FloridaLakes dataset in the Lock5Data package and we will test to see if this sample provides evidence that the average alkalinity of all Florida lakes is greater than 40 mg/L. We will be using the t.test function in R
data(FloridaLakes)
head(FloridaLakes)
t.test(FloridaLakes$Alkalinity, alternative = 'greater', mu = 25)
# <a id='samp_mean_hype'></a>
# ### Hypothesis test
#
# #### Assumptions:
#
# Observations $x_{1}$, . . . , $x_{n}$ are a random sample from the normal distribution N(µ, σ)
#
# #### Hypothesis:
#
# $$H_{0}: µ > 30$$
# $$H_{a}: µ > 30$$
#
# #### T-test:
#
# $$ t = 2.3878$$
#
# *We use this test when σ is not known
#
# #### P-value:
#
# $$ p-value = 0.01031$$
#
# #### Conclusion:
#
# Because the p-value is less than the .05 significance level, we can reject the null hypothesis and conclude that we have enough statistical evidence that the claim is true.
#
# <a id='two_samp'></a>
# ## Two Sample t-test for difference in means
#
# We will belooking mainly at two different ways to do this kind of test. The first will focus on when the two samples are independent and they both have unequal variance(Two-sample independent t-test). One example is the exam score of students trained with method 1 to those trained with method 2.
#
# The second test we will be looking at is when the two samples are not independent and are connected with some sort of treatment or effect. For example, the before and after of students that got a certain learning program and see how well this program worked. This test is called the paired t-test.
# <a id='two_samp_ue'></a>
# ### Two sample independent t-test for unequal variance
#
# #### Rationale
# Usually we would choose the test for unequal variance when there were unequal sample size between the independent sample and choose equal when there were equal sample sizes. However much research has been done on this topic and it has shown that the t-test for unequal variance for two independent samples(Welch Test) performs better than Student's t-test whenever sample sizes and variances are unequal between groups, and gives the same result when sample sizes and variances are equal. except when the sample sizes are very small(5 subjects or less).
#
# Great resource that looks more into this: http://daniellakens.blogspot.com.es/2015/01/always-use-welchs-t-test-instead-of.html
#
# #### Example
#
# We will be looking at the ColaCalcium dataset in the Lock5Data and the questions we will try to answer is if the calcium level in diet cola is greater than the calcium level in water.
##Review data and conduct test
data(ColaCalcium)
head(ColaCalcium)
str(ColaCalcium)
t.test(Calcium~Drink, ColaCalcium, alternative = 'greater')
# <a id='two_samp_ue_hype'></a>
# ### Hypothesis test
#
# #### Assumptions:
# Observations $x_{11}$, . . . , $x_{1n_{1}}$ and $x_{21}$, . . . , $x_{2n_{2}}$
# are random samples from two distinct normal populations N($µ_{1}$, $σ_{1}$) and N($µ_{2}$, $σ_{2}$).
#
# #### Hypothesis:
#
# $$H_{0}: \mu_{1} - \mu_{2} = 0$$
# $$H_{a}: \mu_{1} - \mu_{2} > 0$$
#
# #### T-statistic:
#
# $$t = 3.1732$$
#
# #### P-value
#
# $$p-value = 0.003703$$
#
# #### Conclusion:
#
#
#
#
#
#
# <a id="paired"></a>
# ### Paired t-test for difference in means
#
# We will be using the Wetsuits dataset from the Lock5Data package. From this we will see whether their is a difference between the max velocity of swimmers swimming with a wetsuit and the smae swimmers swimming without one.
data(Wetsuits)
head(Wetsuits)
str(Wetsuits)
t.test(Wetsuits$Wetsuit, Wetsuits$NoWetsuit, paired = TRUE)
# ### Conclusion
#
# The p-value found was less than the .05 significane level, thus we can reject the null hypotheis that the difference in means is equal to 0 and conclude that the true differnece between the two methods is not equal to 0. We see that their is a difference when swimming withthe wetsuit and without the wetsuit.
# <a id='boot'></a>
# ## Randomization Test(Monte Carlo Permutation test)
#
# Randomization tests rely on fewer assumptions than do common parametric tests (such as the t-test) and so can be used when
# requirements for parametric tests are not satisfied, and they can sometimes be more powerful than common
# rank-based nonparametric tests and so also can be used when typical nonparametric tests are not desired.
#
# We will be seing how powerful the test is in comparison to a one sample t-test and also look at how to implment a randomization test and the bootstrap confidence interval in R.
# ### Bootstrap confidence interval
#
# We use the bootstrap confidence interval on some values of samples such as st. dev., correlation, and other metrics in order to get the sampling distribution from one samaple without the need to have assumptions met that usually are very loosely met. We will be looking at how to code this confidence interval in R.
#
# (1-α)% confidence interval using t distribtution(we can use other distributions too:
#
# $$Stat_{PE}\pm t_{1-\alpha/2,n-1}*SE_{boot}$$
#By Hand
#### First get Freedman dataset from cars library
data(Freedman)
summary(Freedman)
Freedman <- na.omit(Freedman)
#We have omitted all the NA values now lets find the correlation between density and the crime for a city
summary(Freedman)
cor(Freedman$density, Freedman$crime)
nrow(Freedman)
# +
#Not that much of a correlation now lets make a bootstrap confidence interval for the correlation
cor_boot <- function(num_boot, samp_size, x, y){
boot_dist <- rep(NA, num_boot)
#Dont usually want to use for loops in R, will later fix to apply function
for (i in 1:num_boot){
#Sample with replacement the sample size
boot_sample<-sample(samp_size, replace = TRUE)
#For each of the samples we find the correlations and put it into the list
boot_dist[i]<-cor(x[boot_sample], y[boot_sample])
}
hist(boot_dist)
return(boot_dist)
}
boot_dist <- cor_boot(5000, nrow(Freedman), Freedman$density, Freedman$crime)
# -
#Sample correlation +- t * se of the bootstrap dist
cor(Freedman$density,Freedman$crime)+c(-1,1)*qt(0.975,nrow(Freedman))*sd(boot_dist)
# #### Interpretation of results
# We are 95% confident that the true correlation between density and crime is between the values -0.1514 and 0.3758.
# <a id='rand'></a>
# ### Randomization test
#
# Why use it - Dont have to assume a distribution and does not need to be randomly sampled however it nevertheless remains important for randomization tests that the data come from an experiment in which experimental units have been randomly assigned to treatments. We will first go over some topics such as Type 1 and Type 2 error and then do a power analysis to see how it does compared to its conunterpart test.
#
# #### Formal Definitions
#
# Type 1 error - the error of rejecting a null
# hypothesis when it is actually true.
#
# Type 2 error - the error of not rejecting a null
# hypothesis when the alternative hypothesis is true.
#
# Statistical Power - The power of a study is the likelihood that it will distinguish an effect of a certain size from pure luck. Also 1 - Type 2 error.
# #### How to do a randomization test for one sample
#
# The problem we are looking at is levels of toxins in different food products. We only get a couple of samples (6) and we need to find out if the products have a mean level greater than 80. Because the one sample t-test for population mean has its assumption of sample size violated so we use the randomization test
# +
# Find the sample mean of the samples
toxin <- c(68, 75, 81, 93, 95, 134)
set.seed(123)
doRandTestOne <- function(samples, times, null){
rand_dist <- rep(NA,times)
#Adjust value in the original sample by 91-80 = 11 to center
adjusted <- samples - (mean(samples) - null)
for(i in 1:times){
rand_sample <- sample(length(samples),replace = TRUE)
rand_dist[i] <- mean(adjusted[rand_sample])
}
#Determine the proportion of samples greater than the original mean, 91
p_value <- mean(rand_dist >= mean(samples))
return(p_value)
}
doRandTestOne(toxin, 10000, 80)
# -
####Using the function
# <a id='anova'></a>
# ## ANOVA
#
# ANOVA is a statistical test where we want to figure out if there is a difference in means between three or more populations. We do this instead of doing multiple t-tests because of the type 1 error that increases when you do multiple testing.
#
# Some assumptions that need to be met are:
# • Each group is a random sample from their population
# • Groups are independent and observations are independent
# • Observations within each group are normally distributed
# • Variance of each group is equal (homoscedasticity)
#
# Assumptions however can be
#
# For hypothesis testing we will be using the F-statistic which is the ratio of the variability between groups and variability within groups. This follows a F distribution that is right skewed, non-negative, and has 2 df's.
#
# We will be lookin at a example where we compare differnent kinds of fillings to how many ants come to the sandwich with that filling and see if there is a noticeable difference between filling types
data(SandwichAnts)
head(SandwichAnts)
one.way.anova <- aov(Ants ~ Filling, data = SandwichAnts)
summary(one.way.anova)
summary.lm(one.way.anova)
# <a id='tukey'></a>
# ### Tukey HSD Test for Multiple pairwise-comparison between the means of groups
## Rationale is to determine if the mean difference between specific pairs of group are statistically significant
TukeyHSD(one.way.anova)
# <a id='anova_val'></a>
# ### Checking Assumptions
#Equal variance for all filling groups
plot(one.way.anova, 1)
##From graph it looks like it might not be supported so we do the Levene test from the car library to make sure
leveneTest(Ants~Filling, data = SandwichAnts)
# Now to check if the residuals follow a normal distribtution
# Extract the residuals
res <- residuals(one.way.anova)
qqnorm(res, main = "QQ Plot of Residuals")
qqline(res)
# Does not look like it does so we do the Shapiro-Wilks test to make sure
# Run Shapiro-Wilk test
shapiro.test(res)
# <a id='anova_res'></a>
# ### Results
# From looking at the table after conducting the ANOVA test on the different groups we get a p-value that is less than the .05 significance level thus we can reject the null hypothesis an conclude their is statistical evidence that theres a difference in means in the different filling types.
#
# In terms of the specific kinds of pairing between fillings we see from the Tukey HSD test that Peanut Butter-Ham vs Pickles and
# Vegemite-Ham vs Pickles had p-values that were less than the .05 significance level meaning there was seen to be a significant difference between those pairs means while the Vegemite vs Peanut Butter did not have enough statistical evidence to find a difference in the sample means
#
# When validating the assumptions of ANOVA we need to see if the variances are homoschedastic or equal between the different filling types. Using the Levene test we find a p-value greater than the significance level thus we cannot reject the null hypothesis that the variancees are equal. We also look at the residuals and we need to find that the distribution is normal. We do the Shapiro-Wilks normality test and we find that the p-value is greater than the .05 significance level thus we cannot reject the null hypothesis that the residual are normal.
# <a id='two_anova'></a>
# ## Two Way ANOVA
# ## (factorial treatment structure or a factorial design)
# What it is: compares the levels of two categorical variables
# on a single continuous response.
#
# Why do we use it: Because we want to see not just the effects of each of the categorical variables but also want to see the itneraction between them
#
# Assumptions are the same as the One-Way ANOVA where the groups have equal variance and the residuals follow a normal distribtuion.
#
# We will be looking at two different kinds of factorial designs, balanced and unbalanced. We use balanced when groups have equal samples and we can use the standard Two-Way ANOVA. For unbalanced this is not true and we will use Type 3 Two-Way ANOVA
# ### Balanced Design
# Load in the data
data(ToothGrowth)
head(ToothGrowth)
#Change into factors and see that the groups have equal sample size
ToothGrowth$supp <- as.factor(ToothGrowth$supp)
ToothGrowth$dose <- as.factor(ToothGrowth$dose)
str(ToothGrowth)
table(ToothGrowth$supp, ToothGrowth$dose)
#Visualize the data
ggplot(ToothGrowth, aes(x = dose, y = len, color = supp))+
geom_boxplot()
#Conduct the Two Way ANOVA test
aov2 <- aov(len ~ supp * dose, ToothGrowth)
summary(aov2)
#Conduct Tukey HSD to see if any specific inferences can be made
TukeyHSD(aov2)
# ### Check Assumptions
# +
#Check homogenity og variances by checking residual vs fitted values
plot(aov2, 1)
# Outliers might invalidate our assumption so we do the levene test to make sure
leveneTest(len ~ supp * dose, ToothGrowth )
# +
# Check if residual follow normal distribution
plot(aov2, 2)
#Use shapiro wilks to make sure
shapiro.test(residuals(aov2))
# -
# ### Results
#
# We have three hypotheses that we tested. First is if there is an effect with supplement, dosage has an effect and if the interation between dosage and supplement has an effect.
#
# From looking at the table after we do the two way anova test, we see that the p-values for all of the things we wantwed to test for are seen to be significant thus we can eject the null hypotheses and conclude there is statistical evidence that suggest that there are significant difference between levels of the supplement, dosage and the interaction in terms of amount of ants.
#
# We also see that all the assumptions are found to be met as well thus validating our test.
#
# ## Unbalanced
# Before we do the analysis we will look at all the different types of the Two Way ANOVA test.
#
# Type 1: Used mainly when we have balanced factorial design. This uses type 1 sum of squares that is the incremental reduction of residual sum of squares for each factor added into the model. This approach test for main effects of factor a followed by factor b and then followed by the interaction ab.
#
# Type 2: Order does not matter like in type 1 and the incremental reduction of sum of squares is due to adding all other terms containing the effect being tested.
#
# Type 3: It is the incremental reduction of residual sum of
# squares due to adding that term to the model consisting of all
# other terms including the interaction and is computed after all the other effect are in the model.
#
# We usually just want to use type 3 because it works in a wide variety of cases with unbalanced data while type 2 works better when the unbalanced data does not have the interaction term in the model
# Load Data
data(Cars2015)
head(Cars2015)
# Choose Type, Drive and High price and put them in their own df
df <- Cars2015[c('Type', 'Drive', 'HighPrice')]
table(df$Type, df$Drive)
#we see that the groups are not balanced from the table we will now do two way anova with type 3 sum of squares
aov3 <- aov(HighPrice ~ Type * Drive, df)
Anova(aov3, type = 'III')
# <a id='ancova'></a>
# ## ANCOVA
#
# We have been mainly looking at ANOVA models with without any covariates or a quantitative variable that we are not very interested in but still might have an effect on our response.
#
# The model is very similar to a linear model and here is a link to what the model looks like: https://image.slidesharecdn.com/t15-ancova-120411124946-phpapp01/95/t15-ancova-7-728.jpg?cb=1334148735
#
# We will be looking at the Orange data set to figure out how to do this in R.
#Read in the data and take a look at it
data(Orange)
head(Orange)
Orange$Tree = factor(as.numeric(Orange$Tree))
str(Orange)
#We will be using the circumference of the orange as the response and see if the type of tree has any effect on it
#Covariate will be the age of the tree and we will see if this has any effect on the circumference.
#We will first see the a model with just the covariate in it
lm1 <- lm(Orange$circumference ~ Orange$age)
summary(lm1)
# We see that the covariate is significant now lets check the type of trees category
lm2 <- lm(Orange$circumference ~ Orange$Tree)
anova(lm2)
summary(lm2)
#We see that the type of tree does not have much difference in circumference from one another now lets check the ANCOVA model
lm3 <- lm(Orange$circumference ~ Orange$age + Orange$Tree)
summary(lm3)
#We see that all of the variables are significant except for tree 2
#Because we have nested models we can now use anova to determine which model is more significant
anova(lm1, lm3)
# <a id='dist'></a>
# ## Distributions
#
# Definition - The distribution of a variable is a description of the relative numbers of times each possible outcome will occur in a number of trials.
#
# Uniform Distribution
#
# Normal Distribution
#
# t - Distribution:
#
# F-Distribution
#
# Chi- Square Distribution
#
# Bernoulli Distribution
#
# Binomial Distribution
#
# Poisson Distribtuion
#
# And there are many more but we will stick to these only.
# <a id='lm'></a>
# ## Linear Regression
#
# ### Why do we use it?
#
# We use it because we want to be able to see if there is a strong linear relationship between the quantitative variables and also to be able to predict a quantitative response
#
#
# ### What are the assumptions required for linear regression?
#
# - There is a linear relationship between the dependent variables and the regressors, meaning the model you are creating actually fits the data,
# - The errors or residuals of the data are normally distributed and independent from each other,
# - There is minimal multicollinearity between explanatory variables, and
# - Homoscedasticity. This means the variance around the regression line is the same for all values of the predictor variable.
#
# ### What method do we use to assess a linear relationship?
#
# We use the least squares method which pretty much finds the equation of the line that best fits the date. How it does that is it find values of the intercept and the slope that best minimizes the residual sum of squares value or the summation of error values squared. The intercept is the expected value of y when X = 0 an the slope is the average increase in Y associated with a one unit increase in X.
#
# ### Why do we do multiple linear regression instead of multiple linear regression?
#
# - Prediction is not possible because three regression equations are made
# - Each of the three regression equations ignores the other two media in forming estimates for the regression coefficients
# Linear model in r
advert <- read.csv('advert.csv')
str(advert)
head(advert)
summary(advert)
fit1 <- lm(Sales~., data = advert)
summary(fit1)
fit2 <- lm(Sales ~ TV*Radio*Newspaper , data = advert)
summary(fit2)
fit3 <- lm(Sales ~.+ TV:Newspaper , data = advert)
summary(fit3)
# #### Why is newspaper now significant with the interaction term in the model?
#
anova(fit1, fit3)
#Accuracy of the fit
# +
#We will be looking at the UN data set in the cars library to see if infant mortality has linear relationship \
#with the countries gdp
data(UN)
str(UN)
summary(UN)
## Not that many NA's in each column so just omit them
UN <- na.omit(UN)
# +
#Check out the relationship
ggplot(UN, aes(UN$infant.mortality, UN$gdp)) +
geom_point()
#We need to do a transformation to get that linear transformation
ggplot(UN, aes(log(UN$infant.mortality), log(UN$gdp))) +
geom_point()
# -
fit1 <- lm(log(UN$gdp) ~ log(UN$infant.mortality))
summary(fit1)
plot(fit1)
# <a id='logistic'></a>
# ## Logistic Regression
#
# ### Why do we use it?
#
# Because we have a indepnedent variable dependent variable that has two levels and we want to predcit the probability that the response will happen. Ex. Male vs Female, Yes vs No.
#
# ### What is the difference in models?
#
# Predicted values are probabilities and not numbers. Probabilities are whether or not 0 or 1, yes, or no, or any other binary outcome. Linear regression uses least squares to arrive at a best possible fit while logistic regression uses maximum likelihood method to arrive at the solution of best estimating coefficients
#
# The equation for just one variable logistic regression(Transforms linear equation using the logistic function):
#
# $$p(X)=\frac{e^{β_{0}+β_{1}X}}{1+e^{β_{0}+β_{1}X}}$$
#
# Also after manipulation you can get the odds and log odds model.
# After getting the log odds model we can get the linear equation:
#
# $$log(\frac{p(X)}{1-p(X)})=β_{0}+β_{1}X$$
#
# Now we can make some sense of what it actually means in context. Regardless of the value of X, if β1 is positive then increasing X will be associated with increasing p(X), and if β1 is negative then increasing X will be associated with decreasing p(X). We cannot assume a straight line relationship between X and p(X).
#
# ### Using Qualitative Variable
#
# We have the same model but just code the variables 1 for yes or 0 for no and then using maximum likelihoof estimator we get a coefficient and computation is doe normally when predicting probability of the observation being a certain class.
#We look at the data and check out the distributions
#of each of the variables
data(Default)
str(Default)
barplot(table(Default$student))
barplot(table(Default$default))
boxplot(Default$balance)
boxplot(Default$income)
# We get the fit and we find that income is the only variable
# that doesn't
fit1 <- glm(default ~ ., Default, family = binomial)
summary(fit1)
# <a href='#back'>Back to the top</a>
data()
| Stats-Review.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="1fkNjVSEyCbC"
# # Algorithm
# 1. Take 2 integers
# 2. See the higher integer.
# 3. Put the higher one before the '='.
# 4. then put the smaller after the '='.
# 5. If they are equal, say that the no. are it's H.C.F.
# 6. print(larger int = smaller int * some no. + the reaminder)
# 7. repeat step 6 until r=0
# 8. when r=0, the devisor is the H.C.F.
# + id="UtZsDwvSx_ZX"
devident=input("Enter the devident: ")
devisor=input("Enter the devisor: ")
r = int(devident) % int(devisor)
q = int(devident) // int(devisor)
print(str(devident),"=", str(devisor),"×",str(q),"+",str(r))
# + [markdown] id="G00KmrONyIGi"
# # Lessons Learnt
# * Foucs on making the most basic version of the project first. Don't go for advanced stuff in the beginning.
# * Errors are a significant part of programming.
| Euclid's Division Lemma/Euclid's Division Lemma.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/vikaskumar04121998/practical-for-deep-learning/blob/main/AlexNet_with_transfer_learning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="9vlUConvm4tx"
https://github.com/vikaskumar04121998/practical-for-deep-learning/blob/main/AlexNet%20with%20transfer%20learning.ipynb
# + id="VT03zMO6lTLy" outputId="36f7fa2c-a67e-4765-9916-aba4d512f6a3"
# %matplotlib inline
from keras.preprocessing.image import ImageDataGenerator
from keras.optimizers import SGD
from alexnet_base import *
from utils import *
# + id="LN4a_D7nlTL2"
batch_size = 16
input_size = (3,227,227)
nb_classes = 2
mean_flag = True # if False, then the mean subtraction layer is not prepended
# + id="w5AE3enllTL4" outputId="c3ad27f6-2a64-42d3-b98d-e95eca77b917"
#code ported from https://blog.keras.io/building-powerful-image-classification-models-using-very-little-data.html
# this is the augmentation configuration we will use for training
train_datagen = ImageDataGenerator(
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True)
test_datagen = ImageDataGenerator()
train_generator = train_datagen.flow_from_directory(
'../Data/Train',
batch_size=batch_size,
shuffle=True,
target_size=input_size[1:],
class_mode='categorical')
validation_generator = test_datagen.flow_from_directory(
'../Data/Test',
batch_size=batch_size,
target_size=input_size[1:],
shuffle=True,
class_mode='categorical')
# + [markdown] id="oVLtwVAtlTL6"
# ## Task 1 : Training AlexNet from scratch
# + id="TMU0WBfOlTL8" outputId="805e8383-3646-4d13-fbb9-7473fd2317a7"
alexnet = get_alexnet(input_size,nb_classes,mean_flag)
print alexnet.summary()
# + id="j2ZFVHgBlTL-" outputId="8efc53e3-ef4c-4de5-8675-ee3ad7a86c62"
sgd = SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
alexnet.compile(loss='mse',
optimizer=sgd,
metrics=['accuracy'])
history = alexnet.fit_generator(train_generator,
samples_per_epoch=2000,
validation_data=validation_generator,
nb_val_samples=800,
nb_epoch=80,
verbose=1)
# + id="pfOvHifplTMA" outputId="688faa20-59f6-4b58-e045-5fb853cff94d"
plot_performance(history)
# + [markdown] id="uZ4Z0Fz8lTMC"
# ### Task 2 : Fine-Tuning Alexnet
# + id="e9FkEYA9lTMD" outputId="4894b06a-f3a2-48a3-c87e-e7d33bc52ccf"
alexnet = get_alexnet(input_size,nb_classes,mean_flag)
alexnet.load_weights('../convnets-keras/weights/alexnet_weights.h5', by_name=True)
print alexnet.summary()
# + id="nVBqlFBIlTMF" outputId="b29666d1-31e9-4a72-e79c-42773e0cf125"
layers = ['dense_3_new','dense_2','dense_1','conv_5_1','conv_4_1','conv_3','conv_2_1','conv_1']
epochs = [10,10,10,10,10,10,10,10]
lr = [1e-2,1e-3,1e-3,1e-3,1e-3,1e-3,1e-3,1e-3]
history_finetune = []
for i,layer in enumerate(layers):
alexnet = unfreeze_layer_onwards(alexnet,layer)
sgd = SGD(lr=lr[i], decay=1e-6, momentum=0.9, nesterov=True)
alexnet.compile(loss='mse',
optimizer=sgd,
metrics=['accuracy'])
for epoch in range(epochs[i]):
h = alexnet.fit_generator(train_generator,
samples_per_epoch=2000,
validation_data=validation_generator,
nb_val_samples=800,
nb_epoch=1,
verbose=1)
history_finetune = append_history(history_finetune,h)
# + id="Awd92L4UlTMG" outputId="30141712-816b-4ed7-d76f-6f8cad544391"
plot_performance(history_finetune)
# + [markdown] id="njpu19V3lTMJ"
# ### Task 3 : Using AlexNet as a feature extractor
#
# + id="Svpr3ty4lTMK"
alexnet = get_alexnet(input_size,nb_classes,mean_flag)
alexnet.load_weights('../convnets-keras/weights/alexnet_weights.h5', by_name=True)
alexnet_convolutional_only = Model(input=alexnet.input, output=alexnet.get_layer('convpool_5').output)
# + id="lJSiezkdlTMM" outputId="1a915aff-e276-420a-d4a3-7b0d8a124ace"
#code ported from https://blog.keras.io/building-powerful-image-classification-models-using-very-little-data.html
import numpy as np
generator = train_datagen.flow_from_directory(
'../Data/Train',
target_size=(227, 227),
batch_size=batch_size,
class_mode=None, # this means our generator will only yield batches of data, no labels
shuffle=False) # our data will be in order, so all first 1000 images will be cats, then 1000 dogs
# the predict_generator method returns the output of a model, given
# a generator that yields batches of numpy data
train_data = alexnet_convolutional_only.predict_generator(generator, 2000)
train_labels = np.array([[1, 0]] * 1000 + [[0, 1]] * 1000)
generator = test_datagen.flow_from_directory(
'../Data/Test',
target_size=(227, 227),
batch_size=batch_size,
class_mode=None,
shuffle=False)
validation_data = alexnet_convolutional_only.predict_generator(generator, 800)
validation_labels = np.array([[1,0]] * 400 + [[0,1]] * 400)
# + id="YECLSpB_lTMO" outputId="12eb854c-6f23-46c4-90e0-c01aa5858136"
from keras.models import Sequential
model = Sequential()
model.add(Flatten(input_shape=train_data.shape[1:]))
model.add(Dense(256, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(2, activation='softmax'))
sgd = SGD(lr=0.001, decay=1e-6, momentum=0.9, nesterov=True)
model.compile(optimizer=sgd,
loss='mse',
metrics=['accuracy'])
history_convpool_5 = model.fit(train_data, train_labels,
nb_epoch=80,
batch_size=batch_size,
validation_data=(validation_data, validation_labels),
verbose=2)
# + id="NqdoAnTclTMQ" outputId="56b68b2e-f28b-45d0-e5b5-497cb38f00c0"
plot_performance(history_convpool_5)
| AlexNet_with_transfer_learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# Which model is most accurate? Support Vector Machine (SVM), Decision Tree (C4.5), Naive Bayes (NB) and k Nearest Neighbors (k-NN), random forest, neural network
# # Logistic Regression
# +
from sklearn.datasets import load_breast_cancer
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
cancer = pd.read_csv('breast_cancer.csv')
X_train, X_test, y_train, y_test = train_test_split(cancer.loc[:,"mean radius":"worst fractal dimension"],cancer['target'], stratify=cancer['target'], random_state=42)
log_reg_model = LogisticRegression()
log_reg_model.fit(X_train, y_train)
# +
print('Accuracy of training set: {:.3f}'.format(log_reg_model.score(X_train, y_train)))
print('Accuracy of testing set: {:.3f}'.format(log_reg_model.score(X_test, y_test)))
# -
# ### there are some underfitting !!
#
# ### bying change the paramter to boost the accuracy and correct the overfitting
log_reg_model
# +
logReg100_model = LogisticRegression(C=100)
logReg100_model.fit(X_train, y_train)
print('Accuracy of training set: {:.3f}'.format(logReg100_model.score(X_train, y_train)))
print('Accuracy of Testing set: {:.3f}'.format(logReg100_model.score(X_test, y_test)))
# -
# ## prediction
prediction = logReg100_model.predict(X_test)
prediction
# ## uncertainty Estimation
print('The decision function is(the confidence of each result predicted(pos - Benign, neg - Malignant): \n\n{}'.format(logReg100_model.decision_function(X_test)))
print("------------------------------------------------")
print('Thresholded decision function (whether result is predicted as Benign) :\n\n{}'.format(logReg100_model.decision_function(X_test)>0))
# ### Logistic Regression Visualization
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
benign_count = len(prediction[prediction == 1])
malignant_count = len(prediction[prediction == 0])
objects = ('Benign','Malignent')
y_pos = np.arange(len(objects))
performance = [benign_count, malignant_count ]
plt.bar(y_pos, performance, align='center', alpha=0.5)
plt.xticks(y_pos, objects)
plt.ylabel('Count')
plt.title('Predicted Results')
plt.show()
# -
predict = logReg100_model.decision_function(X_test)
predict
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
benign = list(predict[predict > 0])
malignant = list(predict[predict < 0])
malignantIndex = []
for i in malignant:
malignantIndex.append(malignant.index(i))
benignIndex = []
for i in benign:
benignIndex.append(benign.index(i))
plt.scatter(benignIndex, benign, color = "green", alpha=0.5)
plt.ylabel('Confident Value')
plt.title(f'Predicted Results (benign): Count of value: {len(benignIndex)}')
plt.show()
plt.scatter(malignantIndex,malignant , color = "Red", alpha=0.5)
plt.ylabel('Confident Value')
plt.title(f'Predicted Results (malignant): Count of value: {len(malignantIndex)}')
plt.show()
# -
from sklearn.metrics import classification_report
print(classification_report(y_test, predictions,
target_names=["blue", "red"]))
# # Support Vector Machine
# +
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
cancer = pd.read_csv('breast_cancer.csv')
X_train, X_test, y_train, y_test = train_test_split(cancer.loc[:,"mean radius":"worst fractal dimension"],cancer['target'], stratify=cancer['target'], random_state=0)
svm_model = SVC()
svm_model.fit(X_train, y_train)
print('The accuracy on the training subset: {:.3f}'.format(svm_model.score(X_train, y_train)))
print('The accuracy on the testing subset: {:.3f}'.format(svm_model.score(X_test, y_test)))
# +
import matplotlib.pyplot as plt
# %matplotlib inline
plt.plot(X_train.min(axis=0), 'o', label = "Min")
plt.plot(X_train.max(axis=0), 'v', label = "Max")
plt.xlabel("Feature Index")
plt.ylabel("feature mag in log scale")
plt.yscale("log")
plt.legend()
# -
# #### we have a overfitting Siuation due to scaling issue
#
# Scale the data feature and modify the paramaters
# +
min_train = X_train.min(axis=0)
range_train = (X_train - min_train).max(axis=0)
X_train_scaled = (X_train - min_train)/range_train
print('Minimum per feature\n{}'.format(X_train_scaled.min(axis=0)))
print('Maximum per feature\n{}'.format(X_train_scaled.max(axis=0)))
# +
X_test_scaled = (X_test - min_train)/range_train
svm = SVC()
svm.fit(X_train_scaled, y_train)
print('The accuracy on the training subset: {:.3f}'.format(svm.score(X_train_scaled, y_train)))
print('The accuracy on the test subset: {:.3f}'.format(svm.score(X_test_scaled, y_test)))
# -
# now the underfitting is fixed now I want to change the parameter to see if I could boost up the accuracy
# +
svm = SVC(C=800)
svm.fit(X_train_scaled, y_train)
print('The accuracy on the training subset: {:.3f}'.format(svm.score(X_train_scaled, y_train)))
print('The accuracy on the test subset: {:.3f}'.format(svm.score(X_test_scaled, y_test)))
# +
svm = SVC(C=1000)
svm.fit(X_train_scaled, y_train)
print('The accuracy on the training subset: {:.3f}'.format(svm.score(X_train_scaled, y_train)))
print('The accuracy on the test subset: {:.3f}'.format(svm.score(X_test_scaled, y_test)))
# -
# ### Prediction
from sklearn.metrics import classification_report
print(classification_report(y_test, predictions,
target_names=["blue", "red"]))
prediction = svm.predict(X_test_scaled)
prediction
# # Decison Tree
# # Random Forest
| Final_Project/ML-models/Breast Cancer ML Modeling Logistic_Regression & SVM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from sklearn.linear_model import LinearRegression
import matplotlib.pyplot as plt
#read data
bmi_life_data = pd.read_csv('bmi_and_life_expectancy.csv')
x_values = bmi_life_data[['BMI']]
y_values = bmi_life_data[['Life expectancy']]
#train model on data
bmi_life_model = LinearRegression()
bmi_life_model.fit(x_values, y_values)
laos_life_exp = bmi_life_model.predict(21.07931)
print(laos_life_exp)
#visualize results
plt.scatter(x_values, y_values)
plt.plot(x_values, bmi_life_model.predict(x_values))
plt.show()
| DeepLearning - Udacity/code/LinearRegression/LinearRegression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
import torch
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
import seaborn as sns
from selective_gp.models import DeepGPModel
from selective_gp.layers import SVGP
from selective_gp.utils import (
load_data, get_model, remove_points, fit_layerwise)
from selective_gp.utils.visualization import (
plot_density, plot_samples, plot_deep_latent)
sns.set(
font_scale=1.5,
style="whitegrid",
)
fig_width = 16
# -
def plot_all(model):
fig, axes = plt.subplots(1, len(model.gps), figsize=(16, 5))
for i, ax in enumerate(axes, 1):
ax.set_title(f"Layer {i}")
ax.margins(x=0)
plot_deep_latent(model, axes=axes)
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(16, 5))
for ax in (ax1, ax2):
ax.set(xlim=(-5, 25), ylim=(-1, 2))
ax1.set_title("Posterior samples")
ax2.set_title("Posterior density")
plot_samples(model, ax=ax1)
plot_density(model, ax=ax2)
# # Probabilistic selection of inducing points in deep Gaussian processes
#
# In this notebook we demonstrate how the selection of inducing points through variational inference is extended to the more general setting of deep Gaussian processes.
#
# Consider the case of wanting to model a square wave with a sparse Gaussian process with a radial basis function (RBF) kernel (which is generally a bad idea).
dataset = load_data("square_wave", n_observations=200, test_size=0.5)
X, Y = dataset.X_train, dataset.Y_train
fig, ax = plt.subplots(figsize=(fig_width, 5))
ax.set(title="Observed data", xlim=(-5, 25), ylim=(-1, 2))
ax.plot(X.flatten(), Y.flatten(), "x", color=plt.cm.Greys(0.9, 0.5));
M = 80
model = DeepGPModel(add_input=False)
model.eval()
gp = SVGP(1, 1, n_inducing=M)
model.add_gp(gp)
gp.inducing_inputs = torch.linspace(-5, 25, M)
model.fit(X=X, Y=Y, max_epochs=200)
# +
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(16, 5), sharey=True)
ax1.set_title("Function posterior (noise excluded)")
ax2.set_title("Model posterior (noise included)")
for ax in fig.axes:
ax.margins(x=0)
ax.plot(X.flatten(), Y.flatten(), "kx")
cm = plt.cm.Set1
x_in = torch.linspace(-5, 25)
with torch.no_grad():
f_dist = gp(x_in[:, None])
y_dist = model.likelihood(f_dist)
f_m, f_s = f_dist.mean, f_dist.stddev
y_m, y_s = y_dist.mean, y_dist.stddev
ax1.plot(x_in, f_m, color=cm(1))
ax1.fill_between(x_in, f_m - f_s, f_m + f_s, color=cm(1, 0.3))
ax2.plot(x_in, y_m, color=cm(0))
ax2.fill_between(x_in, y_m - y_s, y_m + y_s, color=cm(0, 0.3)),
# -
# Although not entirely unreasonable, the Gaussian process finds it difficult to handle the discontinuities. This is largely due to a critical assumption, that we have baked into our model through our choice of kernel: that the covariance between any pair of observations is solely determined by their distance (stationarity). In particular, two points in the same, vertical segment (e.g. $x = 0$ and $x = 2$) must have the same covariance as two points with same distance but in different segments (e.g. $x = 2$ and $x = 4$), even though the former pair should obviously be considered more informative about one another. The model is now forced to use a lengthscale that enables "jumping" between segments, but with the result of too much flexibility within segments and high kernel variance and likelihood noise.
#
# One way of circumventing the stationarity assumption is by making multiple compositions of Gaussian processes, thus creating a deep, hierarchical model. That is, the (uncertain) outputs of the first function becomes the (uncertain) inputs to the next function, and so on. Now, two points, that have the same distance in one layer, may not have so in the next, meaning that the stationarity assumption is effectively lifted.
# To see how this increases expressiveness, we will add a single layer and re-fit. We initialise the inducing points for the new layer by taking 10 samples from the current model and use the minimum and maximum values as the range limits.
outputs = torch.stack([model.sample(X) for _ in range(10)])
zmin, zmax = outputs.min(), outputs.max()
gp = SVGP(1, 1, n_inducing=M)
model.add_gp(gp)
gp.inducing_inputs = torch.linspace(zmin, zmax, M)
model.fit(X=X, Y=Y, max_epochs=500)
plot_all(model)
# The resulting model has learned an approximate step function in the second layer that "pulls apart" 0- and 1-evaluations. Note that the uncertainty outside of the input range $x \in (0, 20)$ now resembles a bi-modal distribution.
#
# We can further refine the estimation by adding another layer.
outputs = torch.stack([model.sample(X) for _ in range(10)])
zmin, zmax = outputs.min(), outputs.max()
gp = SVGP(1, 1, n_inducing=M)
model.add_gp(gp)
gp.inducing_inputs = torch.linspace(zmin, zmax, M)
model.fit(X=X, Y=Y, max_epochs=500)
plot_all(model)
# At this point, it is informative to consider the complexity of each function in our deep model. While the first function still approximates a square wave and thus needs to cover the entire input range $x = (0, 20)$, the second and third functions are considerably simpler. It may then seem reasonable that quite a few inducing points can be removed from the last two layers without it having severe impact on the predictive capabilities of the model, although it may affect computation time.
#
# To test this hypothesis, we will measure sample time and RSME for the current model, use the variational point processes to prune the layers, and compare.
# pre_prune_time = %timeit -o model.sample(dataset.X_test, True)
samples = torch.stack([model.sample(dataset.X_test) for _ in range(50)])
pre_prune_rmse = (((samples.T - dataset.Y_test)**2).mean()).sqrt()
# +
for gp in model.gps:
gp.prior_point_process.rate.fill_(0.5)
gp.variational_point_process.probabilities = 0.2
model.fit_score_function_estimator(
X=X, Y=Y, learning_rate=0.3, max_epochs=1000,
n_mcmc_samples=8)
# -
fig, axes = plt.subplots(1, 3, figsize=(fig_width, 5))
for ax in axes:
ax.margins(x=0)
plot_deep_latent(model, axes=axes)
# The filling of each pseudo-point in the above figure is the latent probability of inclusion as inferred by the variational points processes. Note that more points are deemed redundant in the 2nd and 3rd layer.
#
# We remove those points by sampling from the point processes, and then re-fit the pruned model.
# +
for gp in model.gps:
remove_points(gp)
for i, gp in enumerate(model.gps, 1):
print(f"Number of pseudo-points in layer {i}: {gp.n_inducing}")
model.fit(X=X, Y=Y, max_epochs=500, hp_learning_rate=0.005,
var_learning_rate=0.01)
# -
plot_all(model)
# Notice that there does not seem to be much, if any, detoriation in the predictive capabilities. Comparing to the pre-pruned model:
# +
# post_prune_time = %timeit -o model.sample(dataset.X_test, True)
samples = torch.stack([model.sample(dataset.X_test) for _ in range(50)])
post_prune_rmse = (((samples.T - dataset.Y_test)**2).mean()).sqrt()
# -
print(f"Pre-prune RMSE: {pre_prune_rmse:.2e}\n"
f"Pre-prune time: {pre_prune_time}\n\n"
f"Post-prune RMSE: {post_prune_rmse:.2e}\n"
f"Post-prune time: {post_prune_time}")
| notebooks/02_DeepGaussianProcess.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Deep learning for identifying the orientation Scanned images
#
# First we will load the train and test data and create a CTF file
# +
import os
from PIL import Image
import numpy as np
import itertools
import random
import time
import matplotlib.pyplot as plt
import cntk as C
def split_line(line):
splits = line.strip().split(',')
return (splits[0], int(splits[1]))
def load_labels_dict(labels_file):
with open(labels_file) as f:
return dict([split_line(line) for line in f.readlines()[1:]])
def load_data(data_dir, labels_dict):
for f in os.listdir(data_dir):
key = f[:-4]
label = labels_dict[key]
image = np.array(Image.open(os.path.join(data_dir, f)), dtype = np.int16).flatten()
yield np.hstack([image, int(label)])
def write_to_ctf_file(generator, test_file_name, train_file_name, pct_train = 0.9, rng_seed = 0):
random.seed(rng_seed)
labels = [l for l in map(' '.join, np.eye(4, dtype = np.int16).astype(str))]
with open(test_file_name, 'w') as testf:
with open(train_file_name, 'w') as trainf:
lines = 0
for entry in generator:
rand_num = random.random()
formatted_line = '|labels {} |features {}\n'.format(labels[int(entry[-1])], ' '.join(entry[:-1].astype(str)))
if rand_num <= pct_train:
trainf.write(formatted_line)
else:
testf.write(formatted_line)
lines += 1
if lines % 1000 == 0:
print('Processed {} entries'.format(str(lines)))
# +
train_data_dir = os.path.join('data', 'train')
labels_file = os.path.join('data', 'train_labels.csv')
train_file = 'train_data.ctf'
test_file = 'test_data.ctf'
all_data_file = 'all_data.ctf'
labels_dict = load_labels_dict(labels_file)
if os.path.exists(train_file) and os.path.exists(test_file):
print("Test and training CTF Files exists, not recreating them again")
else:
generator = load_data(train_data_dir, labels_dict)
write_to_ctf_file(generator, test_file, train_file)
#Created only to enable testing on entire test data to hoping to improve the submission score
if os.path.exists(all_data_file):
print("All data CTF Files exists, not recreating it again")
else:
generator = load_data(train_data_dir, labels_dict)
labels = [l for l in map(' '.join, np.eye(4, dtype = np.int16).astype(str))]
with open(all_data_file, 'w') as f:
lines = 0
for entry in generator:
formatted_line = '|labels {} |features {}\n'.format(labels[int(entry[-1])], ' '.join(entry[:-1].astype(str)))
f.write(formatted_line)
lines += 1
if lines % 1000 == 0:
print('Processed {} entries'.format(str(lines)))
# +
np.random.seed(0)
C.cntk_py.set_fixed_random_seed(1)
C.cntk_py.force_deterministic_algorithms()
num_output_classes = 4
input_dim_model = (1, 64, 64)
def create_reader(file_path, is_training):
print('Creating reader from file ' + file_path)
ctf = C.io.CTFDeserializer(file_path, C.io.StreamDefs(
labels = C.io.StreamDef(field='labels', shape = 4, is_sparse=False),
features = C.io.StreamDef(field='features', shape = 64 * 64, is_sparse=False),
))
return C.io.MinibatchSource(ctf, randomize = is_training, max_sweeps = C.io.INFINITELY_REPEAT if is_training else 1)
# +
x = C.input_variable(input_dim_model)
y = C.input_variable(num_output_classes)
def create_model(features):
with C.layers.default_options(init = C.glorot_uniform(), activation = C.relu):
h = features
h = C.layers.Convolution2D(filter_shape=(5, 5),
num_filters = 32,
strides=(2, 2),
pad=True, name='first_conv')(h)
h = C.layers.MaxPooling(filter_shape = (5, 5), strides = (2, 2), name = 'pool1')(h)
h = C.layers.Convolution2D(filter_shape=(5, 5),
num_filters = 64,
strides=(2, 2),
pad=True, name='second_conv')(h)
h = C.layers.MaxPooling(filter_shape = (3, 3), strides = (2, 2), name = 'pool2')(h)
r = C.layers.Dense(num_output_classes, activation = None, name='classify')(h)
return r
# -
def print_training_progress(trainer, mb, frequency, verbose=1):
training_loss = "NA"
eval_error = "NA"
if mb % frequency == 0:
training_loss = trainer.previous_minibatch_loss_average
eval_error = trainer.previous_minibatch_evaluation_average
if verbose:
print ("Minibatch: {0}, Loss: {1:.4f}, Error: {2:.2f}%".format(mb, training_loss, eval_error*100))
# +
def train_test(train_reader, test_reader, model_func, num_sweeps_to_train_with=10):
model = model_func(x/255)
# Instantiate the loss and error function
loss = C.cross_entropy_with_softmax(model, y)
label_error = C.classification_error(model, y)
# Initialize the parameters for the trainer
minibatch_size = 64
num_samples_per_sweep = 60000
num_minibatches_to_train = (num_samples_per_sweep * num_sweeps_to_train_with) / minibatch_size
learning_rate = 0.1
lr_schedule = C.learning_rate_schedule(learning_rate, C.UnitType.minibatch)
learner = C.sgd(model.parameters, lr_schedule)
trainer = C.Trainer(model, (loss, label_error), [learner])
input_map={
y : train_reader.streams.labels,
x : train_reader.streams.features
}
training_progress_output_freq = 500
start = time.time()
for i in range(0, int(num_minibatches_to_train)):
data=train_reader.next_minibatch(minibatch_size, input_map = input_map)
trainer.train_minibatch(data)
print_training_progress(trainer, i, training_progress_output_freq, verbose=1)
print("Training took {:.1f} sec".format(time.time() - start))
test_input_map = {
y : test_reader.streams.labels,
x : test_reader.streams.features
}
test_minibatch_size = 64
num_samples = 2000
num_minibatches_to_test = num_samples // test_minibatch_size
test_result = 0.0
for i in range(num_minibatches_to_test):
data = test_reader.next_minibatch(test_minibatch_size, input_map=test_input_map)
eval_error = trainer.test_minibatch(data)
test_result = test_result + eval_error
# Average of evaluation errors of all test minibatches
print("Average test error: {0:.2f}%".format(test_result*100 / num_minibatches_to_test))
# -
def do_train_test(model, train_on_all_data = False):
if train_on_all_data:
reader_train = create_reader(all_data_file, True)
else:
reader_train = create_reader(train_file, True)
reader_test = create_reader(test_file, False)
train_test(reader_train, reader_test, model)
# +
C.cntk_py.set_fixed_random_seed(1)
C.cntk_py.force_deterministic_algorithms()
model = create_model(x)
print('pool2 shape is ' + str(model.pool2.shape))
C.logging.log_number_of_parameters(model)
do_train_test(model, train_on_all_data = False)
#Test data not relevant here in case we use all data, the tests won't be out of sample
#Just done as an attempt improve the submission score using all possible test data after we find the best model
#that gave minimum error on validation set
#Surprisingly, it didn't improve the score but reduced the score by a fraction.
#do_train_test(model, train_on_all_data = True)
# +
#Accumulate and display the misclassified
#TODO: FIX this
test_reader = create_reader(test_file, False)
labels = []
predictions = []
all_images = []
for i in range(0, 2000, 500):
validation_data = test_reader.next_minibatch(500)
features = validation_data[test_reader.streams.features].as_sequences()
all_images += features
l = validation_data[test_reader.streams.labels].as_sequences()
labels += [np.argmax(i.flatten()) for i in l]
images = [i.reshape(1, 64, 64) for i in features]
preds = model(images)
predictions += [np.argmax(i.flatten()) for i in preds]
predictions = np.array(predictions)
labels = np.array(labels)
mask = predictions != labels
mismatch = np.array(all_images)[mask]
expected_label = labels[mask]
mismatch_pred = predictions[mask]
mismatch_images = np.array(all_images)[mask]
# +
# %matplotlib inline
for i in range(len(expected_label)):
fig = plt.figure(figsize = (8, 6))
ax = fig.gca()
ax.set_title('Expected label ' + str(expected_label[i]) + ', got label ' + str(mismatch_pred[i]))
image = mismatch_images[i]
plt.imshow(image.reshape(64, 64), cmap = 'gray')
plt.axis('off')
# +
submission_data_dir = os.path.join('data', 'test')
submission_file = 'submission_data.ctf'
def file_to_ndarray(file_root, imfile):
return (imfile[:-4], np.array(Image.open(os.path.join(file_root, imfile))).reshape((-1, 64, 64)))
submission_images = [file_to_ndarray(submission_data_dir, f) for f in os.listdir(submission_data_dir)]
submission_images = sorted(submission_images, key = lambda x: x[0])
input_images = [x[1].astype(np.float32) / 255 for x in submission_images]
all_predictions = []
submission_mini_batch_size = 50
for i in range(0, 20000, submission_mini_batch_size):
predictions = model(input_images[i:(i + submission_mini_batch_size)])
all_predictions.append(np.argmax(predictions, axis = 1))
all_predictions = [item for sl in all_predictions for item in sl]
with open('submission.csv', 'w') as f:
f.write('id,orientation\n')
for i in range(20000):
f.write(submission_images[i][0] + "," + str(all_predictions[i]) + "\n")
# -
| 8 - Microsoft Professional Capstone/Medical+Image+Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Moran Processes
#
# The evolutionary models discussed in the previous chapters assume an infinite population that can be divided in to infinitessimal parts. Finite populations can also be studied using a model called a Moran Process (first described in 1958).
#
# ---
#
# ## Moran process with neutral drift
#
# [Video](https://youtu.be/OeMku85hwEc?list=PLnC5h3PY-znxMsG0TRYGOyrnEO-QhVwLb)
#
# Consider a population of two types of fixed size $N$. This can be represented as a vector of the form: $(i, N-i)$ where $i\geq 0$ represents the number of individuals of the first type.
#
# The term **neutral** drift refers to the fact that the two types reproduce at the same rate.
#
# The Moran process is as follows:
#
# - At a given time step: select a random individual for reproduction and a random individual for elimination
# - The eliminated individual is replaced by a new individual of the same type as the individual chosen for reproduction.
# - Proceed to the next time step.
# - The process terminates when there is only one type of individual in the population.
#
# ---
#
# Here is some simple Python code that simulates such a Process assuming an initial population of $(3, 3)$:
# + tags=["nbval-ignore-output"]
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
def neutral_moran(N, i=1, seed=0):
"""
Return the population counts for the Moran process with neutral drift.
"""
population = [0 for _ in range(i)] + [1 for _ in range(N - i)]
counts = [(population.count(0), population.count(1))]
np.random.seed(seed)
while len(set(population)) == 2:
reproduce_index = np.random.randint(N)
eliminate_index = np.random.randint(N)
population[eliminate_index] = population[reproduce_index]
counts.append((population.count(0), population.count(1)))
return counts
N = 6
plt.plot(neutral_moran(N=N, i=3, seed=6));
# -
# For different seeds we see we obtain different results. What becomes of interest is not the path but the end result: which strategy overcomes the presence of the other?
def neutral_fixation(N, i=None, repetitions=10):
"""
Repeat the neutral Moran process and calculate the fixation probability
"""
fixation_count = 0
for seed in range(repetitions):
final_counts = neutral_moran(N=N, i=i, seed=seed)
if final_counts[-1][0] > 0:
fixation_count += 1
return fixation_count / repetitions
# Let us take a look at probability of the first strategy taking over for different starting populations:
# + tags=["nbval-ignore-output"]
probabilities = [neutral_fixation(N, i=i, repetitions=500) for i in range(1, N)]
plt.scatter(range(1, N), probabilities)
plt.xlabel("$i$")
plt.ylabel("$x_i$");
# -
# We see that as the initial population starts with more of a given type, the chance that that type "takes over" (becomes fixed) grows.
#
# This Moran Process is a specific case of a Markov Process:
#
# - A given state of the system can be described by a single integer $0\leq i\leq N$;
# - The state to state transition probabilities are given by:
#
# $$
# \begin{aligned}
# p_{i, i-1}&=\frac{i(N - i)}{N^2}\\
# p_{i, i+1}&=\frac{i(N - i)}{N^2}\\
# p_{i, i}&=1 - p_{i, i-1} - p_{i, i+1}
# \end{aligned}
# $$
#
# We also have two absorbing states (when the Moran process ends):
#
# $$p_{00}=1\qquad p_{0i}=0\text{ for all }i>0$$
#
# $$
# p_{NN}=1\qquad p_{Ni}=0\text{ for all } N>i
# $$
#
# these transitions can be represented as a matrix. Here for example is the matrix for $N=6$:
N = 6
p = np.zeros((N + 1, N + 1))
p[0, 0] = 1
p[N, N] = 1
for i in range(1, N):
for j in [i - 1, i + 1]:
p[i, j] = i * (N - i) / (N ** 2)
p[i, i] = 1 - sum(p[i, :])
p.round(2)
# The above corresponds to a particular type of Markov process called a Birth-Death process
#
# ---
#
# ## Birth death process
#
# [Video](https://youtu.be/zJQQF2tq9AA?list=PLnC5h3PY-znxMsG0TRYGOyrnEO-QhVwLb)
#
# A birth death process is a Markov process with the following properties:
#
# - $p_{i,i+1}+p_{i,i-1}\leq 1$
# - $p_{ii}=1-p_{i,i+1}-p_{i,i-1}$
# - $p_{00}=1$ and $p_{NN}=1$
#
# ---
#
# Thus we have two absorbing states: $\{0, N\}$. Let us denote by $x_i$ the probability of being in $state$ $i$ and eventually reaching state $N$.
#
# We have the following linear system:
#
# \begin{align}
# x_0&=0\\
# x_i&=p_{i,i-1}x_{i-1}+p_{ii}x_i+p_{i,i+1}x_{i+1}\text{ for all }0< i< N-1\\
# x_N&=1\\
# \end{align}
#
# ---
#
# ## Theorem: Fixation probabilities for the birth death process
#
# Given a birth death process as defined above, the fixation probability $x_i$ is given by:
#
# $$x_i=\frac{1+\sum_{j=1}^{i-1}\prod_{k=1}^j\gamma_k}{1+\sum_{j=1}^{N-1}\prod_{k=1}^j\gamma_k}$$
#
# where:
#
# $$
# \gamma_k = \frac{p_{k,k-1}}{p_{k,k+1}}
# $$
#
# ### Proof
#
# We have:
#
# $$
# \begin{aligned}
# p_{i,i+1}x_{i+1} & = -p_{i,i-1}x_{i-1} + x_i(1 - p_{ii}) \\
# p_{i,i+1}x_{i+1} & = p_{i,i-1}(x_{i} - x_{i-1}) + x_ip_{i,i+1} \\
# x_{i+1} - x_i & = \frac{p_{i, i-1}}{p_{i, i+1}}(x_i-x_{i-1})=\gamma_i(x_i-x_{i-1})
# \end{aligned}
# $$
#
# We observe that:
#
# $$
# \begin{aligned}
# x_2 - x_1 &= \gamma_1(x_1-x_{0})=\gamma_1x_1\\
# x_3 - x_2 &= \gamma_2(x_2-x_1)=\gamma_2\gamma_1x_1\\
# x_4 - x_3 &= \gamma_3(x_3-x_2)=\gamma_3\gamma_2\gamma_1x_1\\
# &\; \vdots & \\
# x_{i+1} - x_i &= \gamma_i(x_i-x_{i-1})=\prod_{k=1}^i\gamma_kx_1\\
# &\; \vdots & \\
# x_{N} - x_{N-1} &= \gamma_{N-1}(x_{N-1}-x_{N-2})=\prod_{k=1}^{N-1}\gamma_kx_1\\
# \end{aligned}
# $$
#
# thus we have:
#
# $$x_i=\sum_{j=0}^{i-1}x_{j+1}-x_j=\left(1+\sum_{j=1}^{i-1}\prod_{k=1}^j\gamma_k\right)x_1$$
#
# we complete the proof by solving the following equation to obtain $x_1$:
#
# $$x_N=1=\left(1+\sum_{j=1}^{N-1}\prod_{k=1}^j\gamma_k\right)x_1$$
#
# ---
#
# In the case of neutral drift (considered above) we have:
#
# $$p_{i,i-1}=p_{i,i+1}$$
#
# thus:
#
# $$
# \gamma_i=1
# $$
#
# so:
#
# $$
# x_i=\frac{1+\sum_{j=1}^{i-1}\prod_{k=1}^j\gamma_k}{1+\sum_{j=1}^{N-1}\prod_{k=1}^j\gamma_k}=\frac{1+i-1}{1+N-1}=\frac{i}{N}
# $$
# + tags=["nbval-ignore-output"]
probabilities = [neutral_fixation(N, i=i, repetitions=500) for i in range(1, N)]
plt.scatter(range(1, N), probabilities, label="Simulated")
plt.plot(range(1, N), [i / N for i in range(1, N)], label="Theoretic: $i/N$", linestyle="dashed")
plt.xlabel("$i$")
plt.ylabel("$x_i$")
plt.legend();
# -
# ---
#
# ## Fixation probability
#
# The fixation probability in a Moran process is the probability that a give type starting with $i=1$ individuals takes over an entire population. We denote the fixation probabilities of the first/second type as $\rho_1$ and $\rho_2$ respectively and we have:
#
# $$
# \rho_1=x_1
# $$
#
# $$
# \rho_2=1-x_{N-1}
# $$
#
# ---
#
# We will now consider a Moran process on a game:
#
# ---
#
# ## Moran process on a game
#
# [Video](https://www.youtube.com/watch?v=TpqVoF1fBF8&index=43&list=PLnC5h3PY-znxMsG0TRYGOyrnEO-QhVwLb)
#
#
# Consider a matrix $A\in\mathbb{R}^{m\times n}$ representing a game with two strategies.
#
# $$
# A=
# \begin{pmatrix}
# a & b\\
# c & d
# \end{pmatrix}
# $$
#
# The Moran process is as follows:
#
# - At a given time step: all individuals play all other individuals.
# - Obtain their fitness as given by the game.
# - Randomly select an individual proportional to their fitness as an individual to be reproduced
# - Uniformly select an individual to be replaced
# - Proceed to the next time step.
# - The process terminates when there is only one type of individual in the population.
#
#
# Assuming $i$ individuals of the first type, the fitness of both types is given respectively by:
#
# $$f_{1i}=\frac{a(i-1)+b(N-i)}{N-1}$$
# $$f_{2i}=\frac{c(i)+d(N-i-1)}{N-1}$$
#
# The transition probabilities are then given by:
#
# $$p_{i,i+1}=\frac{if_{1i}}{if_{1i} + (N-i)f_{2i}}\frac{N-i}{N}$$
# $$p_{i,i-1}=\frac{(N-i)f_{2i}}{if_{1i} + (N-i)f_{2i}}\frac{i}{N}$$
#
# which gives:
#
# $$\gamma_i=\frac{f_{2i}}{f_{1i}}$$
#
# thus:
#
# $$
# x_i=\frac{1+\sum_{j=1}^{i-1}\prod_{k=1}^j\gamma_k}{1+\sum_{j=1}^{N-1}\prod_{k=1}^j\gamma_k}
# $$
#
# ---
# Here is some code to carry out this calculation:
def theoretic_fixation(N, game, i=1):
"""
Calculate x_i as given by the above formula
"""
f_ones = np.array([(game[0, 0] * (i - 1) + game[0, 1] * (N - i)) / (N - 1) for i in range(1, N)])
f_twos = np.array([(game[1, 0] * i + game[1, 1] * (N - i - 1)) / (N - 1) for i in range(1, N)])
gammas = f_twos / f_ones
return (1 + np.sum(np.cumprod(gammas[:i-1]))) / (1 + np.sum(np.cumprod(gammas)))
# Here is an example of calculating $x_1$ for the following game for $N=4$:
#
# [Video](https://www.youtube.com/watch?v=3sBVrnQhemE&index=44&list=PLnC5h3PY-znxMsG0TRYGOyrnEO-QhVwLb)
#
# $$
# A =
# \begin{pmatrix}
# 4 & 1\\
# 1 & 4
# \end{pmatrix}
# $$
A = np.array([[4, 1],
[1, 4]])
theoretic_fixation(N=4, i=1, game=A)
# Applying the theorem gives:
#
# $$
# \begin{aligned}
# f_{1i}&=\frac{4(i - 1) + 4 - i}{3} = \frac{4i-4+4-i}{3}=i\\
# f_{2i}&=\frac{i + 4(3 - i)}{3} = \frac{12-3i}{3}=4-i
# \end{aligned}
# $$
#
# $$
# \gamma_i = \frac{f_{2i}}{f_{1i}}=\frac{4-i}{i}=\frac{4}{i}-1
# $$
#
# Thus:
#
# $$
# \begin{aligned}
# x_1 & =\frac{1 + \sum_{j=1}^{0}\prod_{k=1}^{j}\gamma_k}{1 + \sum_{j=1}^{4 - 1}\prod_{k=1}^{j}\gamma_k}\\
# & =\frac{1}{1 + \sum_{j=1}^{3}\prod_{k=1}^{j}\gamma_k}\\
# & =\frac{1}{1 + \gamma_1 + \gamma_1\times \gamma_2 + \gamma_1 \times \gamma_2 \times \gamma_3}\\
# & =\frac{1}{1+3+3\times 1 + 3 \times 1\times \frac{1}{3}} = \frac{1}{1 + 3 + 3 + 1}=\frac{1}{8}\\
# \end{aligned}
# $$
# Nashpy has the ability to run a single Moran process:
# +
import nashpy as nash
game = nash.Game(A)
initial_population = np.array((0, 1, 1, 1))
np.random.seed(0)
generations = game.moran_process(
initial_population=initial_population
)
for population in generations:
print(population)
# -
# We see there that in a population of 4 individuals, a single individual of the first type (`0`) does not become fixed. That is just for a single run, to be able to approximate the fixation probability the process needs to be repeated, this can be done in Nashpy:
# +
import collections
def approximate_fixation(N, A, i=None, repetitions=10):
"""
Repeat the Moran process and calculate the fixation probability
This is done by carrying out the following steps:
1. Creating a game
2. Building an initial population with i individuals
of the first type
3. Getting the fixation probabilities of both types
4. Returning the probability of the first type
"""
game = nash.Game(A)
initial_population = i * [0] + (N - i) * [1]
probabilities = game.fixation_probabilities(
initial_population=initial_population,
repetitions=repetitions
)
return probabilities[0]
# -
# Here is how the fixation probabilities vary for different initial populations:
# + tags=["nbval-ignore-output"]
probabilities = [approximate_fixation(N, i=i, A=A, repetitions=500) for i in range(1, N)]
plt.scatter(range(1, N), probabilities, label="Simulated")
plt.plot(range(1, N), [i / N for i in range(1, N)], label="Neutral: $i/N$", linestyle="dashed")
plt.plot(range(1, N), [theoretic_fixation(N=N, i=i, game=A) for i in range(1, N)], label="Theoretic")
plt.xlabel("$i$")
plt.ylabel("$x_i$")
plt.legend();
| nbs/chapters/09-Moran-processes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Diarie Linköpings kommun
# * Denna [Jupyter Notebook](https://github.com/salgo60/open-data-examples/blob/master/Diarie%20Link%C3%B6pings%20kommun.ipynb)
#
# * om [ärenden Linköping](https://www.linkoping.se/open/data/arende/)
#
# OBS unik nyckel måste begäras se [open](https://www.linkoping.se/open/)
#
# GITHUB [open-data-examples](https://github.com/salgo60/open-data-examples)
#
from IPython.display import JSON
import requests
import json
key = "<KEY>"
baseUrl = "http://arendedata.linkoping.se/api/v1/"
endpoint = "Cases?maxItemsInResult=1000&api_key="
headers = {'content-type': 'application/json'}
url = baseUrl + endpoint + key
url
req = requests.get(url,headers=headers)
data = json.loads(req.text)
import pandas as pd
df = pd.DataFrame(data)
df.head(n=40)
req
import matplotlib.pyplot as plt
# %matplotlib inline
df.info()
df.head(20)
# ### Ärenden
endpointEvent = "Events?maxItemsInResult=1000&api_key=f3d212d2700f4c80aba7d2a88bc94529"
urlEvent = baseUrl + endpointEvent
dataEvent = json.loads(requests.get(urlEvent).text)
import pandas as pd
dfEvent = pd.DataFrame(dataEvent)
dfEvent.head(n=10)
pd.set_option('display.max_rows', None)
dfEvent["diary"].value_counts()
| Diarie Linköpings kommun.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Gradient Checking
#
# Welcome to the final assignment for this week! In this assignment you'll be implementing gradient checking.
#
# By the end of this notebook, you'll be able to:
#
# Implement gradient checking to verify the accuracy of your backprop implementation
# ## Table of Contents
# - [1 - Packages](#1)
# - [2 - Problem Statement](#2)
# - [3 - How does Gradient Checking work?](#3)
# - [4 - 1-Dimensional Gradient Checking](#4)
# - [Exercise 1 - forward_propagation](#ex-1)
# - [Exercise 2 - backward_propagation](#ex-2)
# - [Exercise 3 - gradient_check](#ex-3)
# - [5 - N-Dimensional Gradient Checking](#5)
# - [Exercise 4 - gradient_check_n](#ex-4)
# <a name='1'></a>
# ## 1 - Packages
# +
import numpy as np
from testCases import *
from public_tests import *
from gc_utils import sigmoid, relu, dictionary_to_vector, vector_to_dictionary, gradients_to_vector
# %load_ext autoreload
# %autoreload 2
# -
# <a name='2'></a>
# ## 2 - Problem Statement
#
# You are part of a team working to make mobile payments available globally, and are asked to build a deep learning model to detect fraud--whenever someone makes a payment, you want to see if the payment might be fraudulent, such as if the user's account has been taken over by a hacker.
#
# You already know that backpropagation is quite challenging to implement, and sometimes has bugs. Because this is a mission-critical application, your company's CEO wants to be really certain that your implementation of backpropagation is correct. Your CEO says, "Give me proof that your backpropagation is actually working!" To give this reassurance, you are going to use "gradient checking."
#
# Let's do it!
# <a name='3'></a>
# ## 3 - How does Gradient Checking work?
# Backpropagation computes the gradients $\frac{\partial J}{\partial \theta}$, where $\theta$ denotes the parameters of the model. $J$ is computed using forward propagation and your loss function.
#
# Because forward propagation is relatively easy to implement, you're confident you got that right, and so you're almost 100% sure that you're computing the cost $J$ correctly. Thus, you can use your code for computing $J$ to verify the code for computing $\frac{\partial J}{\partial \theta}$.
#
# Let's look back at the definition of a derivative (or gradient):$$ \frac{\partial J}{\partial \theta} = \lim_{\varepsilon \to 0} \frac{J(\theta + \varepsilon) - J(\theta - \varepsilon)}{2 \varepsilon} $$
#
# If you're not familiar with the "$\displaystyle \lim_{\varepsilon \to 0}$" notation, it's just a way of saying "when $\varepsilon$ is really, really small."
#
# You know the following:
#
# $\frac{\partial J}{\partial \theta}$ is what you want to make sure you're computing correctly.
# You can compute $J(\theta + \varepsilon)$ and $J(\theta - \varepsilon)$ (in the case that $\theta$ is a real number), since you're confident your implementation for $J$ is correct.
# Let's use equation (1) and a small value for $\varepsilon$ to convince your CEO that your code for computing $\frac{\partial J}{\partial \theta}$ is correct!
# <a name='4'></a>
# ## 4 - 1-Dimensional Gradient Checking
#
# Consider a 1D linear function $J(\theta) = \theta x$. The model contains only a single real-valued parameter $\theta$, and takes $x$ as input.
#
# You will implement code to compute $J(.)$ and its derivative $\frac{\partial J}{\partial \theta}$. You will then use gradient checking to make sure your derivative computation for $J$ is correct.
#
# <img src="images/1Dgrad_kiank.png" style="width:600px;height:250px;">
# <caption><center><font color='purple'><b>Figure 1</b>:1D linear model </font></center></caption>
#
# The diagram above shows the key computation steps: First start with $x$, then evaluate the function $J(x)$ ("forward propagation"). Then compute the derivative $\frac{\partial J}{\partial \theta}$ ("backward propagation").
#
# <a name='ex-1'></a>
# ### Exercise 1 - forward_propagation
#
# Implement `forward propagation`. For this simple function compute $J(.)$
# + deletable=false nbgrader={"cell_type": "code", "checksum": "0f934d7a5ec9e6a41fc9ece5ec6a07fa", "grade": false, "grade_id": "cell-a4be88c5c0419ab7", "locked": false, "schema_version": 3, "solution": true, "task": false}
# GRADED FUNCTION: forward_propagation
def forward_propagation(x, theta):
"""
Implement the linear forward propagation (compute J) presented in Figure 1 (J(theta) = theta * x)
Arguments:
x -- a real-valued input
theta -- our parameter, a real number as well
Returns:
J -- the value of function J, computed using the formula J(theta) = theta * x
"""
# (approx. 1 line)
# J =
# YOUR CODE STARTS HERE
J = theta * x
# YOUR CODE ENDS HERE
return J
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "c775107f8d8491592913f1991d0fc3da", "grade": true, "grade_id": "cell-805a7fd19d554221", "locked": true, "points": 10, "schema_version": 3, "solution": false, "task": false}
x, theta = 2, 4
J = forward_propagation(x, theta)
print ("J = " + str(J))
forward_propagation_test(forward_propagation)
# -
# <a name='ex-2'></a>
# ### Exercise 2 - backward_propagation
#
# Now, implement the `backward propagation` step (derivative computation) of Figure 1. That is, compute the derivative of $J(\theta) = \theta x$ with respect to $\theta$. To save you from doing the calculus, you should get $dtheta = \frac { \partial J }{ \partial \theta} = x$.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "7315e45824efc41770654b46c64c1c14", "grade": false, "grade_id": "cell-c06a1275399b210f", "locked": false, "schema_version": 3, "solution": true, "task": false}
# GRADED FUNCTION: backward_propagation
def backward_propagation(x, theta):
"""
Computes the derivative of J with respect to theta (see Figure 1).
Arguments:
x -- a real-valued input
theta -- our parameter, a real number as well
Returns:
dtheta -- the gradient of the cost with respect to theta
"""
# (approx. 1 line)
# dtheta =
# YOUR CODE STARTS HERE
dtheta = x
# YOUR CODE ENDS HERE
return dtheta
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "79ac4ec84141d381d3f9fffccc19b723", "grade": true, "grade_id": "cell-7b67ed84ac8bfd91", "locked": true, "points": 10, "schema_version": 3, "solution": false, "task": false}
x, theta = 2, 4
dtheta = backward_propagation(x, theta)
print ("dtheta = " + str(dtheta))
backward_propagation_test(backward_propagation)
# -
# <a name='ex-3'></a>
# ### Exercise 3 - gradient_check
#
# To show that the `backward_propagation()` function is correctly computing the gradient $\frac{\partial J}{\partial \theta}$, let's implement gradient checking.
#
# **Instructions**:
# - First compute "gradapprox" using the formula above (1) and a small value of $\varepsilon$. Here are the Steps to follow:
# 1. $\theta^{+} = \theta + \varepsilon$
# 2. $\theta^{-} = \theta - \varepsilon$
# 3. $J^{+} = J(\theta^{+})$
# 4. $J^{-} = J(\theta^{-})$
# 5. $gradapprox = \frac{J^{+} - J^{-}}{2 \varepsilon}$
# - Then compute the gradient using backward propagation, and store the result in a variable "grad"
# - Finally, compute the relative difference between "gradapprox" and the "grad" using the following formula:
# $$ difference = \frac {\mid\mid grad - gradapprox \mid\mid_2}{\mid\mid grad \mid\mid_2 + \mid\mid gradapprox \mid\mid_2} \tag{2}$$
# You will need 3 Steps to compute this formula:
# - 1'. compute the numerator using np.linalg.norm(...)
# - 2'. compute the denominator. You will need to call np.linalg.norm(...) twice.
# - 3'. divide them.
# - If this difference is small (say less than $10^{-7}$), you can be quite confident that you have computed your gradient correctly. Otherwise, there may be a mistake in the gradient computation.
#
# + deletable=false nbgrader={"cell_type": "code", "checksum": "5545d5ca718b8580e72da217b49516ae", "grade": false, "grade_id": "cell-ed57ede577f9d607", "locked": false, "schema_version": 3, "solution": true, "task": false}
# GRADED FUNCTION: gradient_check
def gradient_check(x, theta, epsilon=1e-7, print_msg=False):
"""
Implement the backward propagation presented in Figure 1.
Arguments:
x -- a float input
theta -- our parameter, a float as well
epsilon -- tiny shift to the input to compute approximated gradient with formula(1)
Returns:
difference -- difference (2) between the approximated gradient and the backward propagation gradient. Float output
"""
# Compute gradapprox using left side of formula (1). epsilon is small enough, you don't need to worry about the limit.
# (approx. 5 lines)
# theta_plus = # Step 1
# theta_minus = # Step 2
# J_plus = # Step 3
# J_minus = # Step 4
# gradapprox = # Step 5
# YOUR CODE STARTS HERE
theta_plus = theta + epsilon # Step 1
theta_minus = theta - epsilon # Step 2
J_plus = forward_propagation(x, theta_plus) # Step 3
J_minus = forward_propagation(x, theta_minus) # Step 4
gradapprox = (J_plus - J_minus)/(2*epsilon) # Step 5
# YOUR CODE ENDS HERE
# Check if gradapprox is close enough to the output of backward_propagation()
#(approx. 1 line) DO NOT USE "grad = gradapprox"
# grad =
# YOUR CODE STARTS HERE
grad = backward_propagation(x, theta)
# YOUR CODE ENDS HERE
#(approx. 1 line)
# numerator = # Step 1'
# denominator = # Step 2'
# difference = # Step 3'
# YOUR CODE STARTS HERE
numerator = np.linalg.norm(grad - gradapprox) # Step 1'
denominator = np.linalg.norm(grad) + np.linalg.norm(gradapprox) # Step 2'
difference = numerator/denominator # Step 3'
# YOUR CODE ENDS HERE
if print_msg:
if difference > 2e-7:
print ("\033[93m" + "There is a mistake in the backward propagation! difference = " + str(difference) + "\033[0m")
else:
print ("\033[92m" + "Your backward propagation works perfectly fine! difference = " + str(difference) + "\033[0m")
return difference
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "d337f8314fabea08c817cdf32fdbcfb3", "grade": true, "grade_id": "cell-be0338be7d50dd11", "locked": true, "points": 10, "schema_version": 3, "solution": false, "task": false}
x, theta = 2, 4
difference = gradient_check(2,4, print_msg=True)
#gradient_check_test(gradient_check)
# -
# Congrats, the difference is smaller than the $10^{-7}$ threshold. So you can have high confidence that you've correctly computed the gradient in `backward_propagation()`.
#
# Now, in the more general case, your cost function $J$ has more than a single 1D input. When you are training a neural network, $\theta$ actually consists of multiple matrices $W^{[l]}$ and biases $b^{[l]}$! It is important to know how to do a gradient check with higher-dimensional inputs. Let's do it!
# <a name='5'></a>
# ## 5 - N-Dimensional Gradient Checking
# The following figure describes the forward and backward propagation of your fraud detection model.
#
# <img src="images/NDgrad_kiank.png" style="width:600px;height:400px;">
# <caption><center><font color='purple'><b>Figure 2</b>: Deep neural network. LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SIGMOID</font></center></caption>
#
# Let's look at your implementations for forward propagation and backward propagation.
def forward_propagation_n(X, Y, parameters):
"""
Implements the forward propagation (and computes the cost) presented in Figure 3.
Arguments:
X -- training set for m examples
Y -- labels for m examples
parameters -- python dictionary containing your parameters "W1", "b1", "W2", "b2", "W3", "b3":
W1 -- weight matrix of shape (5, 4)
b1 -- bias vector of shape (5, 1)
W2 -- weight matrix of shape (3, 5)
b2 -- bias vector of shape (3, 1)
W3 -- weight matrix of shape (1, 3)
b3 -- bias vector of shape (1, 1)
Returns:
cost -- the cost function (logistic cost for one example)
cache -- a tuple with the intermediate values (Z1, A1, W1, b1, Z2, A2, W2, b2, Z3, A3, W3, b3)
"""
# retrieve parameters
m = X.shape[1]
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
W3 = parameters["W3"]
b3 = parameters["b3"]
# LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SIGMOID
Z1 = np.dot(W1, X) + b1
A1 = relu(Z1)
Z2 = np.dot(W2, A1) + b2
A2 = relu(Z2)
Z3 = np.dot(W3, A2) + b3
A3 = sigmoid(Z3)
# Cost
log_probs = np.multiply(-np.log(A3),Y) + np.multiply(-np.log(1 - A3), 1 - Y)
cost = 1. / m * np.sum(log_probs)
cache = (Z1, A1, W1, b1, Z2, A2, W2, b2, Z3, A3, W3, b3)
return cost, cache
# Now, run backward propagation.
def backward_propagation_n(X, Y, cache):
"""
Implement the backward propagation presented in figure 2.
Arguments:
X -- input datapoint, of shape (input size, 1)
Y -- true "label"
cache -- cache output from forward_propagation_n()
Returns:
gradients -- A dictionary with the gradients of the cost with respect to each parameter, activation and pre-activation variables.
"""
m = X.shape[1]
(Z1, A1, W1, b1, Z2, A2, W2, b2, Z3, A3, W3, b3) = cache
dZ3 = A3 - Y
dW3 = 1. / m * np.dot(dZ3, A2.T)
db3 = 1. / m * np.sum(dZ3, axis=1, keepdims=True)
dA2 = np.dot(W3.T, dZ3)
dZ2 = np.multiply(dA2, np.int64(A2 > 0))
dW2 = 1. / m * np.dot(dZ2, A1.T)
db2 = 1. / m * np.sum(dZ2, axis=1, keepdims=True)
dA1 = np.dot(W2.T, dZ2)
dZ1 = np.multiply(dA1, np.int64(A1 > 0))
dW1 = 1. / m * np.dot(dZ1, X.T)
db1 = 1. / m * np.sum(dZ1, axis=1, keepdims=True)
gradients = {"dZ3": dZ3, "dW3": dW3, "db3": db3,
"dA2": dA2, "dZ2": dZ2, "dW2": dW2, "db2": db2,
"dA1": dA1, "dZ1": dZ1, "dW1": dW1, "db1": db1}
return gradients
# You obtained some results on the fraud detection test set but you are not 100% sure of your model. Nobody's perfect! Let's implement gradient checking to verify if your gradients are correct.
# **How does gradient checking work?**.
#
# As in Section 3 and 4, you want to compare "gradapprox" to the gradient computed by backpropagation. The formula is still:
#
# $$ \frac{\partial J}{\partial \theta} = \lim_{\varepsilon \to 0} \frac{J(\theta + \varepsilon) - J(\theta - \varepsilon)}{2 \varepsilon} \tag{1}$$
#
# However, $\theta$ is not a scalar anymore. It is a dictionary called "parameters". The function "`dictionary_to_vector()`" has been implemented for you. It converts the "parameters" dictionary into a vector called "values", obtained by reshaping all parameters (W1, b1, W2, b2, W3, b3) into vectors and concatenating them.
#
# The inverse function is "`vector_to_dictionary`" which outputs back the "parameters" dictionary.
#
# <img src="images/dictionary_to_vector.png" style="width:600px;height:400px;">
# <caption><center><font color='purple'><b>Figure 2</b>: dictionary_to_vector() and vector_to_dictionary(). You will need these functions in gradient_check_n()</font></center></caption>
#
# The "gradients" dictionary has also been converted into a vector "grad" using gradients_to_vector(), so you don't need to worry about that.
#
# Now, for every single parameter in your vector, you will apply the same procedure as for the gradient_check exercise. You will store each gradient approximation in a vector `gradapprox`. If the check goes as expected, each value in this approximation must match the real gradient values stored in the `grad` vector.
#
# Note that `grad` is calculated using the function `gradients_to_vector`, which uses the gradients outputs of the `backward_propagation_n` function.
#
# <a name='ex-4'></a>
# ### Exercise 4 - gradient_check_n
#
# Implement the function below.
#
# **Instructions**: Here is pseudo-code that will help you implement the gradient check.
#
# For each i in num_parameters:
# - To compute `J_plus[i]`:
# 1. Set $\theta^{+}$ to `np.copy(parameters_values)`
# 2. Set $\theta^{+}_i$ to $\theta^{+}_i + \varepsilon$
# 3. Calculate $J^{+}_i$ using to `forward_propagation_n(x, y, vector_to_dictionary(`$\theta^{+}$ `))`.
# - To compute `J_minus[i]`: do the same thing with $\theta^{-}$
# - Compute $gradapprox[i] = \frac{J^{+}_i - J^{-}_i}{2 \varepsilon}$
#
# Thus, you get a vector gradapprox, where gradapprox[i] is an approximation of the gradient with respect to `parameter_values[i]`. You can now compare this gradapprox vector to the gradients vector from backpropagation. Just like for the 1D case (Steps 1', 2', 3'), compute:
# $$ difference = \frac {\| grad - gradapprox \|_2}{\| grad \|_2 + \| gradapprox \|_2 } \tag{3}$$
#
# **Note**: Use `np.linalg.norm` to get the norms
# + deletable=false nbgrader={"cell_type": "code", "checksum": "e202e41c23c49198f7bf3abb69ed7e1a", "grade": false, "grade_id": "cell-1e5a768bc4e28e66", "locked": false, "schema_version": 3, "solution": true, "task": false}
# GRADED FUNCTION: gradient_check_n
def gradient_check_n(parameters, gradients, X, Y, epsilon=1e-7, print_msg=False):
"""
Checks if backward_propagation_n computes correctly the gradient of the cost output by forward_propagation_n
Arguments:
parameters -- python dictionary containing your parameters "W1", "b1", "W2", "b2", "W3", "b3":
grad -- output of backward_propagation_n, contains gradients of the cost with respect to the parameters.
x -- input datapoint, of shape (input size, 1)
y -- true "label"
epsilon -- tiny shift to the input to compute approximated gradient with formula(1)
Returns:
difference -- difference (2) between the approximated gradient and the backward propagation gradient
"""
# Set-up variables
parameters_values, _ = dictionary_to_vector(parameters)
grad = gradients_to_vector(gradients)
num_parameters = parameters_values.shape[0]
J_plus = np.zeros((num_parameters, 1))
J_minus = np.zeros((num_parameters, 1))
gradapprox = np.zeros((num_parameters, 1))
# Compute gradapprox
for i in range(num_parameters):
# Compute J_plus[i]. Inputs: "parameters_values, epsilon". Output = "J_plus[i]".
# "_" is used because the function you have to outputs two parameters but we only care about the first one
#(approx. 3 lines)
# theta_plus = # Step 1
# theta_plus[i] = # Step 2
# J_plus[i], _ = # Step 3
# YOUR CODE STARTS HERE
theta_plus = np.copy(parameters_values) # Step 1
theta_plus[i][0] = theta_plus[i][0] + epsilon # Step 2
J_plus[i], _ = forward_propagation_n(X, Y, vector_to_dictionary(theta_plus)) # Step 3
# YOUR CODE ENDS HERE
# Compute J_minus[i]. Inputs: "parameters_values, epsilon". Output = "J_minus[i]".
#(approx. 3 lines)
# theta_minus = # Step 1
# theta_minus[i] = # Step 2
# J_minus[i], _ = # Step 3
# YOUR CODE STARTS HERE
theta_minus = np.copy(parameters_values) # Step 1
theta_minus[i][0] = theta_minus[i][0] - epsilon # Step 2
J_minus[i], _ = forward_propagation_n(X, Y, vector_to_dictionary(theta_minus)) # Step 3
# YOUR CODE ENDS HERE
# Compute gradapprox[i]
# (approx. 1 line)
# gradapprox[i] =
# YOUR CODE STARTS HERE
gradapprox[i] = (J_plus[i] - J_minus[i])/(2*epsilon)
# YOUR CODE ENDS HERE
# Compare gradapprox to backward propagation gradients by computing difference.
# (approx. 1 line)
# numerator = # Step 1'
# denominator = # Step 2'
# difference = # Step 3'
# YOUR CODE STARTS HERE
numerator = np.linalg.norm(grad - gradapprox) # Step 1'
denominator = np.linalg.norm(grad) + np.linalg.norm(gradapprox) # Step 2'
difference = numerator/denominator # Step 3'
# YOUR CODE ENDS HERE
if print_msg:
if difference > 2e-7:
print ("\033[93m" + "There is a mistake in the backward propagation! difference = " + str(difference) + "\033[0m")
else:
print ("\033[92m" + "Your backward propagation works perfectly fine! difference = " + str(difference) + "\033[0m")
return difference
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "e119ddabcb075e6e3391464b48e11234", "grade": true, "grade_id": "cell-0d7896ce7c954fc9", "locked": true, "points": 20, "schema_version": 3, "solution": false, "task": false}
X, Y, parameters = gradient_check_n_test_case()
cost, cache = forward_propagation_n(X, Y, parameters)
gradients = backward_propagation_n(X, Y, cache)
difference = gradient_check_n(parameters, gradients, X, Y, 1e-7, True)
expected_values = [0.2850931567761623, 1.1890913024229996e-07]
assert not(type(difference) == np.ndarray), "You are not using np.linalg.norm for numerator or denominator"
assert np.any(np.isclose(difference, expected_values)), "Wrong value. It is not one of the expected values"
# -
# **Expected output**:
#
# <table>
# <tr>
# <td> <b> There is a mistake in the backward propagation!</b> </td>
# <td> difference = 0.2850931567761623 </td>
# </tr>
# </table>
# It seems that there were errors in the `backward_propagation_n` code! Good thing you've implemented the gradient check. Go back to `backward_propagation` and try to find/correct the errors *(Hint: check dW2 and db1)*. Rerun the gradient check when you think you've fixed it. Remember, you'll need to re-execute the cell defining `backward_propagation_n()` if you modify the code.
#
# Can you get gradient check to declare your derivative computation correct? Even though this part of the assignment isn't graded, you should try to find the bug and re-run gradient check until you're convinced backprop is now correctly implemented.
#
# **Notes**
# - Gradient Checking is slow! Approximating the gradient with $\frac{\partial J}{\partial \theta} \approx \frac{J(\theta + \varepsilon) - J(\theta - \varepsilon)}{2 \varepsilon}$ is computationally costly. For this reason, we don't run gradient checking at every iteration during training. Just a few times to check if the gradient is correct.
# - Gradient Checking, at least as we've presented it, doesn't work with dropout. You would usually run the gradient check algorithm without dropout to make sure your backprop is correct, then add dropout.
#
# Congrats! Now you can be confident that your deep learning model for fraud detection is working correctly! You can even use this to convince your CEO. :)
# <br>
# <font color='blue'>
#
# **What you should remember from this notebook**:
# - Gradient checking verifies closeness between the gradients from backpropagation and the numerical approximation of the gradient (computed using forward propagation).
# - Gradient checking is slow, so you don't want to run it in every iteration of training. You would usually run it only to make sure your code is correct, then turn it off and use backprop for the actual learning process.
| C02W01/1.3 Gradient_Checking/Gradient_Checking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import sys
from itertools import combinations, groupby
from collections import Counter
from IPython.display import display
# Function that returns the size of an object in MB
def size(obj):
return "{0:.2f} MB".format(sys.getsizeof(obj) / (1000 * 1000))
orders = pd.read_csv('order_products__prior.csv')
print('orders -- dimensions: {0}; size: {1}'.format(orders.shape, size(orders)))
display(orders.head())
# Convert from DataFrame to a Series, with order_id as index and item_id as value
orders = orders.set_index('order_id')['product_id'].rename('item_id')
display(orders.head(10))
type(orders)
print('dimensions: {0}; size: {1}; unique_orders: {2}; unique_items: {3}'
.format(orders.shape, size(orders), len(orders.index.unique()), len(orders.value_counts())))
#orders.index.unique gives unique number of orders
#orders.value_counts gives unique values of item_id
# Returns frequency counts for items and item pairs
def freq(iterable):
if type(iterable) == pd.core.series.Series:
return iterable.value_counts().rename("freq")
else:
return pd.Series(Counter(iterable)).rename("freq")
# +
#freq(orders)
# -
# Returns number of unique orders
def order_count(order_item):
return len(set(order_item.index))
# Returns generator that yields item pairs, one at a time
def get_item_pairs(order_item):
order_item = order_item.reset_index().as_matrix()
for order_id, order_object in groupby(order_item, lambda x: x[0]):
item_list = [item[1] for item in order_object]
for item_pair in combinations(item_list, 2):
yield item_pair
# Returns frequency and support associated with item
def merge_item_stats(item_pairs, item_stats):
return (item_pairs
.merge(item_stats.rename(columns={'freq': 'freqA', 'support': 'supportA'}), left_on='item_A', right_index=True)
.merge(item_stats.rename(columns={'freq': 'freqB', 'support': 'supportB'}), left_on='item_B', right_index=True))
# Returns name associated with item
def merge_item_name(rules, item_name):
columns = ['itemA','itemB','freqAB','supportAB','freqA','supportA','freqB','supportB',
'confidenceAtoB','confidenceBtoA','lift']
rules = (rules
.merge(item_name.rename(columns={'item_name': 'itemA'}), left_on='item_A', right_on='item_id')
.merge(item_name.rename(columns={'item_name': 'itemB'}), left_on='item_B', right_on='item_id'))
return rules[columns]
def association_rules(order_item, min_support):
print("Starting order_item: {:22d}".format(len(order_item)))
# Calculate item frequency and support
item_stats = freq(order_item).to_frame("freq")
item_stats['support'] = item_stats['freq'] / order_count(order_item) * 100
# Filter from order_item items below min support
qualifying_items = item_stats[item_stats['support'] >= min_support].index
order_item = order_item[order_item.isin(qualifying_items)]
print("Items with support >= {}: {:15d}".format(min_support, len(qualifying_items)))
print("Remaining order_item: {:21d}".format(len(order_item)))
# Filter from order_item orders with less than 2 items
order_size = freq(order_item.index)
qualifying_orders = order_size[order_size >= 2].index
order_item = order_item[order_item.index.isin(qualifying_orders)]
print("Remaining orders with 2+ items: {:11d}".format(len(qualifying_orders)))
print("Remaining order_item: {:21d}".format(len(order_item)))
# Recalculate item frequency and support
item_stats = freq(order_item).to_frame("freq")
item_stats['support'] = item_stats['freq'] / order_count(order_item) * 100
# Get item pairs generator
item_pair_gen = get_item_pairs(order_item)
# Calculate item pair frequency and support
item_pairs = freq(item_pair_gen).to_frame("freqAB")
item_pairs['supportAB'] = item_pairs['freqAB'] / len(qualifying_orders) * 100
print("Item pairs: {:31d}".format(len(item_pairs)))
# Filter from item_pairs those below min support
item_pairs = item_pairs[item_pairs['supportAB'] >= min_support]
print("Item pairs with support >= {}: {:10d}\n".format(min_support, len(item_pairs)))
# Create table of association rules and compute relevant metrics
item_pairs = item_pairs.reset_index().rename(columns={'level_0': 'item_A', 'level_1': 'item_B'})
item_pairs = merge_item_stats(item_pairs, item_stats)
item_pairs['confidenceAtoB'] = item_pairs['supportAB'] / item_pairs['supportA']
item_pairs['confidenceBtoA'] = item_pairs['supportAB'] / item_pairs['supportB']
item_pairs['lift'] = item_pairs['supportAB'] / (item_pairs['supportA'] * item_pairs['supportB'])
# Return association rules sorted by lift in descending order
return item_pairs.sort_values('lift', ascending=False)
# %%time
rules = association_rules(orders, 0.01)
# Replace item ID with item name and display association rules
item_name = pd.read_csv('products.csv')
item_name = item_name.rename(columns={'product_id':'item_id', 'product_name':'item_name'})
rules_final = merge_item_name(rules, item_name).sort_values('lift', ascending=False)
display(rules_final)
# The text in the document by 'Instacart Kaggle Competition' is licensed under CC BY 3.0 https://creativecommons.org/licenses/by/3.0/us/
#
# The code in the document by 'datatheque' Kaggle is licensed under the MIT License https://opensource.org/licenses/MIT
| Apriori Instacart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
import statsapi
# + pycharm={"name": "#%%\n"}
astros_data = statsapi.get('team_roster', {'teamId':117})
player_list = []
print('firstName,lastName,fullName,birthDate,twitter,instagram,youtube')
for player in astros_data['roster']:
player_data = statsapi.get('person', {'personId': player['person']['id'],'hydrate':'social'})
# player_data = statsapi.get('person', {'personId': player['person']['id'], 'fields':'people,id,fullName,firstName,lastName,primaryNumber,currentTeam,id,primaryPosition,code,abbreviation,useName,boxscoreName,nickName,mlbDebutDate,nameFirstLast,firstLastName,lastFirstName,lastInitName,initLastName,fullFMLName,fullLFMName,social'})
info = player_data["people"][0]
social = info.get('social', {})
twitter = social.get('twitter', [''])
instagram = social.get('instagram', [''])
youtube = social.get('youtube', [''])
print(f'{info["firstName"]},{info["lastName"]},{info["fullName"]},{info["birthDate"]},{twitter[0]},{instagram[0]},{youtube[0]}')
# print(player_data)
# player = statsapi.player_stats(621043, type="season")
# print(player)
| astros-roster.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Datajoint Quick Introduction
# The goal of this notebook is to quickly go over some the most commonly used datajoint features in relationship to typical project.
# ## Why use database?
# For small projects, just writing stuff to a file or something along that manner would be sufficient, however when it comes to collobration between large amount of people or when there is a need to store everything in a single location where it can be access by other people as well as being backed up constantly, a database is the way to go.
#
# A database, without going too technical, is just a system that store tables of information and their relationship between each other. There is a whole process that goes into how to design these tables and relationship to be express in the database, but we will go over that later, or at least the basic of it.
# ## Why use datajoint?
# Datajoint is a useful python libaray that allows anyone who knows basic python to interact with a traditional relational database, which in this case is MYSQL.
#
# Without datajoint, to interact with a database, the user must use SQL statements like below which isn't always pretty nor easy to learn without some background knowledge on how database works.
#
# Datajoint is meant to bridge this gap and make the knowledge barrier more tolerable for people don't have time to spend half a year studying about database design and etc.
SELECT column_list
FROM table_1
INNER JOIN table_2 ON join_condition;
# ## Part 1: Installing Datajoint
# Run:
# pip3 install datajoint
# ## Part 2: Setting up datajoint login
# +
import datajoint as dj
# To get your datajoint login, please contact <NAME> on slack, he will be able to create your database credential.
# Datajoint will then use those credentials to login into the database and allow you to use it.
# This is the basic setup of the login information for datajoint. This only needs to be run once per computer.
# In Docker this is done by setting the enviorment variables which datajoint will use (more details on K8 stuff)
dj.config['database.host'] = 'at-database1.ad.bcm.edu' # This the database server, yes you must vpn into Baylor
dj.config['database.username'] = 'kijung' # Provided when you setup an account with Daniel
dj.config['database.password'] = '<PASSWORD>' # Provided when you setup an account with Daniel
dj.config.save_global() # Save the information above on your machine so you don't have to do this again.
# -
# ## Part 3: dj.Manual Tables
# +
import datajoint as dj
# There 3 types of tables that are mainly used in datajoint, dj.Manual, dj.Compute, and dj.Lookup.
# We will focus on dj.Manual for this part.
# Every table needs to be created in a schema, which is where all the tables we will create will go under
# By default, you can only create schema with your username as a prefix, i.e. yourusername_exampleschema
# This is how you set the schema
# schema = dj.schema('kijung_datajoint_tutorial')
# schema = dj.schema('kijung_test_II')
schema = dj.schema('kijung-gnn')
# schema = dj.schema('kijung-gnn-02')
# schema = dj.schema('kijung-gnn-03')
# This is requried during table creation, as you will decreate the table class with this schema so datajoint will know
# where to create the table
# -
@schema
class Sample(dj.Manual):
definition = """
sample_id : int unsigned
"""
schema.jobs.delete_quick()
# temp = dj.create_virtual_module('kijung_datajoint_tutorial', 'kijung_datajoint_tutorial')
# temp = dj.create_virtual_module('kijung_test_II', 'kijung_test_II')
temp = dj.create_virtual_module('kijung-gnn', 'kijung-gnn')
# temp = dj.create_virtual_module('kijung-gnn-02', 'kijung-gnn-02')
# temp = dj.create_virtual_module('kijung-gnn-03', 'kijung-gnn-03')
temp.Sample().delete_quick()
temp.SampleComputed.delete_quick()
# +
import numpy as np
# Sid = np.array([1201, 1202, 1204, 1205, 1207, 1208, 1210, 1213, 1215, 1217, 1220,
# 1221, 1222, 1223, 1226, 1227, 1229, 1230, 1231, 1235, 1238, 1239,
# 1240, 1241, 1242, 1244, 1247, 1257, 1259, 1261, 1267, 1269, 1270,
# 1272, 1273, 1276, 1282, 1283, 1284, 1285, 1291, 1292, 1294, 1295,
# 1296, 1297, 1299, 1300])
# for i in Sid:
# Sample.insert1(dict(sample_id=i))
for i in range(1,37):
Sample.insert1(dict(sample_id=i))
# -
Sample()
schema.jobs
@schema
class SampleComputed(dj.Computed):
definition = """
-> Sample
"""
def make(self, key):
sample_id = key['sample_id']
# Call main function to start computation
main(sample_id)
self.insert1(key) # Just to mark it as completed
SampleComputed()
StudentScore().populate(reserve_jobs=True, suppress_errors=True)
(schema.jobs & dict(status = 'error')).delete_quick()
# +
# Datajoint tables are created via classes with a definition field like this
# dj.Manual creation
@schema
class Student(dj.Manual):
definition = """
student_id : int unsigned
---
first_name : varchar(64)
last_name : varchar(64)
"""
# So a quick run down what all of this syntax means
# Looking at the definiton field, anything values --- is considered to the primary key of the table, or in simpler terms
# the column of values that uniquely identify each row of a table, in this case it is student_id
# For every attribute (column, but in Database desgin we call it attribute) of the table the follwing syntax is a lowercase
# seperated by underscore attirubte name, i.e student_id follow by a colon sepeartor : with a space before and after
# and then the variable type. Datajoint support a couple of variable types, you can look them up here:
# https://github.com/datajoint/wiki/wiki/Data-types
# +
# After creation the table my defining it, you can then access like this:
Student()
# Which of course will be empty since there is nothing in it
# +
# To insert values into the table, you can do them via dicts like this
# Note that the key values of the dict must match the attribute names in the table
# -
# Querying the stduent table again, we should see one value now
Student()
# To restrict by a condition
Student() & 'student_id < 5'
# To restrict by a specific match
Student() & 'student_id = 1'
# or
Student() & dict(student_id = 1)
# fetching a specific value as dict
(Student() & dict(student_id = 1)).fetch1()
# fetching only the primary key of the tuple
(Student() & dict(student_id = 1)).fetch1('KEY')
# +
# If there are buip
# +
# to delete a specific value
(Student() & dict(student_id = 1)).delete_quick()
Student()
# +
# To wipe the table from the DB we run drop_quick()
# Note that this is not recoverabe, same thing for Stduent().delete_quick(), the differnce is that one drop dumps
# the entire table out of the database allowing you to redefine the defintion, and delete just delete the rows only
Student().drop_quick()
# This basically covers the basic of datajoint tables
# -
# ## Part 4 dj.Computed Tables
# +
'''
dj.Computed tables are very similar to dj.Manual execpt that instead of you inserting each entry manually, the entry
will be instead computed from a pre-define function from the user, hence the computed parts
The other advantage of dj.Computed tables is that it can serve as a job queue for multiple instances of your application via
the populate function which is called to start the computation of each row. This is particularly useful in cases with
kubernetes cluster deployment where you can deploy like a 1000 instances of the application and datajoint will handle
in an atomic fashion where each instance is populating 1 unique tuple.
Let begin with creating with recreating the student table
'''
@schema
class Student(dj.Manual):
definition = """
student_id : int unsigned
---
first_name : varchar(64)
last_name : varchar(64)
"""
# -
for i in range(10):
dict_to_insert = dict(
student_id = i,
first_name = 'Daniel' + str(i),
last_name = 'Sitonic' + str(i))
Student.insert1(dict_to_insert)
# +
# Creating the compute table that refernces the student table
@schema
class StudentScore(dj.Computed):
definition = """
-> Student
---
test_score : float
"""
def make(self, key):
'''
This function is what is called during populate, and is where the user put his or her computation code and details
on what to store once the computation is done.
'''
# Key here is a dictionary containing the primary key of Student which is student_id
# If you want to get the attribute then you will need to use fetch1 like the exmaple I put above
key['test_score'] = key['student_id'] * 100
self.insert1(key)
'''
The -> Student is what we call a forgien key refernce as in it reference the primary key of another table which in this
case is Student. This means for each key in Student, there will be a computation tuple in StudentScore that corresponds
with that tuple. In general the dj.Computed table is like your computation code / result storage table.
'''
# -
# Querying the StudentScore table, which should be empty
StudentScore()
# +
'''
Populating the StudentScore table with reserved_jobs=True
reserve_jobs, basically tells the database that whatever tuple it is working on to reserve it and not let other instances
of populate work on that tuple. This is what allow coredenation among mutiple instances.
In general the work flow of the populate function works like this:
Get a forigen key that has not been reserved or computed already -> send to make function -> make function insert into
the computed table -> repeat
suppress_errors=True means not to throw an exception if there is a problem and just continue the loop above
'''
StudentScore().populate(reserve_jobs=True, suppress_errors=True)
# -
# Querying the table again, we shall see that there are computed entires now
StudentScore()
# +
# To make get useful data there is a join command that can be done between the Student and StudentScore table
Student * StudentScore
# This will combine both tables where the student_id matches, after this you can use fetch() to fetch all the tuples all
# at once which will be a list of dicts, then dump it in pandas or something to make format it and plot it
# +
'''
In order to see that status of your currently running jobs when let say you deploy to the cluster. Datajoint records all
reserved jobs and failed jobs along with there error message in schema.jobs
If we query it is now, it will be empty since there is nothing in there, however if there is an error job or
one of the reserved_job instances was stop arrupptly without SIGTERM signal, then it stay in that table until deleted.
The problem is that until it is deleted no other instances of populate can fetch that key to populate thus you must
delete it so the database knows it is clear to allow computation of that key again
'''
schema.jobs
# -
# ## Part 5: Basic Table Design
| DatajointQuickIntroduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: env-tf2
# language: python
# name: env-tf2
# ---
import logging; logging.basicConfig(level=logging.INFO)
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import logictensornetworks as ltn
import pandas as pd
# # Data
#
# Crabs dataset from: http://www.stats.ox.ac.uk/pub/PRNN/
#
# The crabs data frame has 200 rows and 8 columns, describing 5 morphological measurements on 50 crabs each of two colour forms and both sexes, of the species Leptograpsus variegatus collected at Fremantle, W. Australia.
#
# - Multi-class: Male, Female, Blue, Orange.
# - Multi-label: Only Male-Female and Blue-Orange are mutually exclusive.
#
df = pd.read_csv("crabs.dat",sep=" ", skipinitialspace=True)
df = df.sample(frac=1) #shuffle
print(df.head(5))
# We use 160 samples for training and 40 samples for testing.
# +
features = df[['FL','RW','CL','CW','BD']]
labels_sex = df['sex']
labels_color = df['sp']
batch_size=64
ds_train = tf.data.Dataset.from_tensor_slices((features[:160],labels_sex[:160],labels_color[:160])).batch(batch_size)
ds_test = tf.data.Dataset.from_tensor_slices((features[160:],labels_sex[160:],labels_color[160:])).batch(batch_size)
# -
# # LTN
#
# ### Predicate
#
# | index | class |
# | --- | --- |
# | 0 | Male |
# | 1 | Female |
# | 2 | Blue |
# | 3 | Orange |
#
# Let's note that, since the classes are not mutually exclusive, the last layer of the model will be a `sigmoid` and not a `softmax`.
# +
class MLP(tf.keras.Model):
"""Model that returns logits."""
def __init__(self, n_classes, hidden_layer_sizes=(16,16,8)):
super(MLP, self).__init__()
self.denses = [tf.keras.layers.Dense(s, activation="elu") for s in hidden_layer_sizes]
self.dense_class = tf.keras.layers.Dense(n_classes)
def call(self, inputs):
x = inputs
for dense in self.denses:
x = dense(x)
return self.dense_class(x)
logits_model = MLP(4)
p = ltn.Predicate(ltn.utils.LogitsToPredicateModel(logits_model,single_label=False))
# -
# Constants to index the classes
class_male = ltn.constant(0)
class_female = ltn.constant(1)
class_blue = ltn.constant(2)
class_orange = ltn.constant(3)
# ### Axioms
#
# ```
# forall x_blue: C(x_blue,blue)
# forall x_orange: C(x_orange,orange)
# forall x_male: C(x_male,male)
# forall x_female: C(x_female,female)
# forall x: ~(C(x,male) & C(x,female))
# forall x: ~(C(x,blue) & C(x,orange))
# ```
Not = ltn.Wrapper_Connective(ltn.fuzzy_ops.Not_Std())
And = ltn.Wrapper_Connective(ltn.fuzzy_ops.And_Prod())
Or = ltn.Wrapper_Connective(ltn.fuzzy_ops.Or_ProbSum())
Implies = ltn.Wrapper_Connective(ltn.fuzzy_ops.Implies_Reichenbach())
Forall = ltn.Wrapper_Quantifier(ltn.fuzzy_ops.Aggreg_pMeanError(p=2),semantics="forall")
# +
formula_aggregator = ltn.fuzzy_ops.Aggreg_pMeanError(p=2)
@tf.function
def axioms(features,labels_sex,labels_color):
x = ltn.variable("x",features)
x_blue = ltn.variable("x_blue",features[labels_color=="B"])
x_orange = ltn.variable("x_orange",features[labels_color=="O"])
x_male = ltn.variable("x_blue",features[labels_sex=="M"])
x_female = ltn.variable("x_blue",features[labels_sex=="F"])
axioms = [
Forall(x_blue, p([x_blue,class_blue])),
Forall(x_orange, p([x_orange,class_orange])),
Forall(x_male, p([x_male,class_male])),
Forall(x_female, p([x_female,class_female])),
Forall(x,Not(And(p([x,class_blue]),p([x,class_orange])))),
Forall(x,Not(And(p([x,class_male]),p([x,class_female]))))
]
axioms = tf.stack(axioms)
sat_level = formula_aggregator(axioms)
return sat_level, axioms
# -
# Initialize all layers and the static graph.
for features, labels_sex, labels_color in ds_train:
print("Initial sat level %.5f"%axioms(features,labels_sex,labels_color)[0])
break
# # Training
#
# Define the metrics. While training, we measure:
# 1. The level of satisfiability of the Knowledge Base of the training data.
# 1. The level of satisfiability of the Knowledge Base of the test data.
# 3. The training accuracy.
# 4. The test accuracy.
# 5. The level of satisfiability of a formula phi_1 we expect to have a high truth value.
# forall x (p(x,blue)->~p(x,orange))
# 6. The level of satisfiability of a formula phi_1 we expect to have a low truth value.
# forall x (p(x,blue)->p(x,orange))
# 7. The level of satisfiability of a formula phi_1 we expect to have a neither high neither low truth value.
# forall x (p(x,blue)->p(x,male))
#
# +
metrics_dict = {
'train_sat_kb': tf.keras.metrics.Mean(name='train_sat_kb'),
'test_sat_kb': tf.keras.metrics.Mean(name='test_sat_kb'),
'train_accuracy': tf.keras.metrics.Mean(name="train_accuracy"),
'test_accuracy': tf.keras.metrics.Mean(name="test_accuracy"),
'test_sat_phi1': tf.keras.metrics.Mean(name='test_sat_phi1'),
'test_sat_phi2': tf.keras.metrics.Mean(name='test_sat_phi2'),
'test_sat_phi3': tf.keras.metrics.Mean(name='test_sat_phi3')
}
@tf.function()
def phi1(features):
x = ltn.variable("x",features)
return Forall(x, Implies(p([x,class_blue]),Not(p([x,class_orange]))),p=5)
@tf.function()
def phi2(features):
x = ltn.variable("x",features)
return Forall(x, Implies(p([x,class_blue]),p([x,class_orange])),p=5)
@tf.function()
def phi3(features):
x = ltn.variable("x",features)
return Forall(x, Implies(p([x,class_blue]),p([x,class_male])),p=5)
def multilabel_hamming_loss(y_true, y_pred, threshold=0.5,from_logits=False):
if from_logits:
y_pred = tf.math.sigmoid(y_pred)
y_pred = y_pred > threshold
y_true = tf.cast(y_true, tf.int32)
y_pred = tf.cast(y_pred, tf.int32)
nonzero = tf.cast(tf.math.count_nonzero(y_true-y_pred,axis=-1),tf.float32)
return nonzero/y_true.get_shape()[-1]
# +
optimizer = tf.keras.optimizers.Adam(learning_rate=0.001)
@tf.function
def train_step(features, labels_sex, labels_color):
# sat and update
with tf.GradientTape() as tape:
sat = axioms(features, labels_sex, labels_color)[0]
loss = 1.-sat
gradients = tape.gradient(loss, p.trainable_variables)
optimizer.apply_gradients(zip(gradients, p.trainable_variables))
metrics_dict['train_sat_kb'](sat)
# accuracy
predictions = logits_model(features)
labels_male = (labels_sex == "M")
labels_female = (labels_sex == "F")
labels_blue = (labels_color == "B")
labels_orange = (labels_color == "O")
onehot = tf.stack([labels_male,labels_female,labels_blue,labels_orange],axis=-1)
metrics_dict['train_accuracy'](1-multilabel_hamming_loss(onehot,predictions,from_logits=True))
@tf.function
def test_step(features, labels_sex, labels_color):
# sat
sat_kb = axioms(features, labels_sex, labels_color)[0]
metrics_dict['test_sat_kb'](sat_kb)
sat_phi1 = phi1(features)
metrics_dict['test_sat_phi1'](sat_phi1)
sat_phi2 = phi2(features)
metrics_dict['test_sat_phi2'](sat_phi2)
sat_phi3 = phi3(features)
metrics_dict['test_sat_phi3'](sat_phi3)
# accuracy
predictions = logits_model(features)
labels_male = (labels_sex == "M")
labels_female = (labels_sex == "F")
labels_blue = (labels_color == "B")
labels_orange = (labels_color == "O")
onehot = tf.stack([labels_male,labels_female,labels_blue,labels_orange],axis=-1)
metrics_dict['test_accuracy'](1-multilabel_hamming_loss(onehot,predictions,from_logits=True))
# +
import commons
EPOCHS = 200
commons.train(
EPOCHS,
metrics_dict,
ds_train,
ds_test,
train_step,
test_step,
csv_path="crabs_results.csv",
track_metrics=20
)
# -
df_results = pd.read_csv("crabs_results.csv")
df_results.plot(x="Epoch")
| examples/multiclass_classification/multiclass-multilabel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Classification: Instant Recognition with Caffe
#
# In this example we'll classify an image with the bundled CaffeNet model (which is based on the network architecture of Krizhevsky et al. for ImageNet).
#
# We'll compare CPU and GPU modes and then dig into the model to inspect features and the output.
# ### 1. Setup
#
# * First, set up Python, `numpy`, and `matplotlib`.
# +
# set up Python environment: numpy for numerical routines, and matplotlib for plotting
import numpy as np
import matplotlib.pyplot as plt
# display plots in this notebook
# %matplotlib inline
# set display defaults
plt.rcParams['figure.figsize'] = (10, 10) # large images
plt.rcParams['image.interpolation'] = 'nearest' # don't interpolate: show square pixels
plt.rcParams['image.cmap'] = 'gray' # use grayscale output rather than a (potentially misleading) color heatmap
# -
# * Load `caffe`.
# +
# The caffe module needs to be on the Python path;
# we'll add it here explicitly.
import sys
caffe_root = '../' # this file should be run from {caffe_root}/examples (otherwise change this line)
sys.path.insert(0, caffe_root + 'python')
import caffe
# If you get "No module named _caffe", either you have not built pycaffe or you have the wrong path.
# -
# * If needed, download the reference model ("CaffeNet", a variant of AlexNet).
import os
if os.path.isfile(caffe_root + 'models/bvlc_reference_caffenet/bvlc_reference_caffenet.caffemodel'):
print('CaffeNet found.')
else:
print('Downloading pre-trained CaffeNet model...')
# !../scripts/download_model_binary.py ../models/bvlc_reference_caffenet
# ### 2. Load net and set up input preprocessing
#
# * Set Caffe to CPU mode and load the net from disk.
# +
caffe.set_mode_gpu()
caffe.set_device(5) # if we have multiple GPUs, pick the first one
model_def = caffe_root + 'models/bvlc_reference_caffenet/deploy.prototxt'
model_weights = caffe_root + 'models/bvlc_reference_caffenet/bvlc_reference_caffenet.caffemodel'
net = caffe.Net(model_def, # defines the structure of the model
caffe.TEST, # use test mode (e.g., don't perform dropout)
weights=model_weights) # contains the trained weights
# -
# * Set up input preprocessing. (We'll use Caffe's `caffe.io.Transformer` to do this, but this step is independent of other parts of Caffe, so any custom preprocessing code may be used).
#
# Our default CaffeNet is configured to take images in BGR format. Values are expected to start in the range [0, 255] and then have the mean ImageNet pixel value subtracted from them. In addition, the channel dimension is expected as the first (_outermost_) dimension.
#
# As matplotlib will load images with values in the range [0, 1] in RGB format with the channel as the _innermost_ dimension, we are arranging for the needed transformations here.
# +
# load the mean ImageNet image (as distributed with Caffe) for subtraction
mu = np.load(caffe_root + 'python/caffe/imagenet/ilsvrc_2012_mean.npy')
mu = mu.mean(1).mean(1) # average over pixels to obtain the mean (BGR) pixel values
print('mean-subtracted values:', zip('BGR', mu))
# create transformer for the input called 'data'
transformer = caffe.io.Transformer({'data': net.blobs['data'].data.shape})
transformer.set_transpose('data', (2,0,1)) # move image channels to outermost dimension
transformer.set_mean('data', mu) # subtract the dataset-mean value in each channel
transformer.set_raw_scale('data', 255) # rescale from [0, 1] to [0, 255]
transformer.set_channel_swap('data', (2,1,0)) # swap channels from RGB to BGR
# -
# ### 3. CPU classification
#
# * Now we're ready to perform classification. Even though we'll only classify one image, we'll set a batch size of 50 to demonstrate batching.
# set the size of the input (we can skip this if we're happy
# with the default; we can also change it later, e.g., for different batch sizes)
net.blobs['data'].reshape(50, # batch size
3, # 3-channel (BGR) images
227, 227) # image size is 227x227
# * Load an image (that comes with Caffe) and perform the preprocessing we've set up.
image = caffe.io.load_image(caffe_root + 'examples/images/cat.jpg')
transformed_image = transformer.preprocess('data', image)
plt.imshow(image)
# * Adorable! Let's classify it!
# +
# # copy the image data into the memory allocated for the net
net.blobs['data'].data[...] = transformed_image
### perform classification
output = net.forward()
output_prob = output['prob'][0] # the output probability vector for the first image in the batch
print('predicted class is:', output_prob.argmax())
# -
# * The net gives us a vector of probabilities; the most probable class was the 281st one. But is that correct? Let's check the ImageNet labels...
# +
# load ImageNet labels
labels_file = caffe_root + 'data/ilsvrc12/synset_words.txt'
if not os.path.exists(labels_file):
# !../data/ilsvrc12/get_ilsvrc_aux.sh
labels = np.loadtxt(labels_file, str, delimiter='\t')
print('output label:', labels[output_prob.argmax()])
# -
# * "Tabby cat" is correct! But let's also look at other top (but less confident predictions).
# +
# sort top five predictions from softmax output
top_inds = output_prob.argsort()[::-1][:5] # reverse sort and take five largest items
print('probabilities and labels:')
zip(output_prob[top_inds], labels[top_inds])
# -
# * We see that less confident predictions are sensible.
# ### 4. Switching to GPU mode
#
# * Let's see how long classification took, and compare it to GPU mode.
# %timeit net.forward()
# * That's a while, even for a batch of 50 images. Let's switch to GPU mode.
net.forward() # run once before timing to set up memory
# %timeit net.forward()
# * That should be much faster!
# ### 5. Examining intermediate output
#
# * A net is not just a black box; let's take a look at some of the parameters and intermediate activations.
#
# First we'll see how to read out the structure of the net in terms of activation and parameter shapes.
#
# * For each layer, let's look at the activation shapes, which typically have the form `(batch_size, channel_dim, height, width)`.
#
# The activations are exposed as an `OrderedDict`, `net.blobs`.
# for each layer, show the output shape
for layer_name, blob in net.blobs.iteritems():
print(layer_name + '\t' + str(blob.data.shape))
# * Now look at the parameter shapes. The parameters are exposed as another `OrderedDict`, `net.params`. We need to index the resulting values with either `[0]` for weights or `[1]` for biases.
#
# The param shapes typically have the form `(output_channels, input_channels, filter_height, filter_width)` (for the weights) and the 1-dimensional shape `(output_channels,)` (for the biases).
for layer_name, param in net.params.iteritems():
print(layer_name + '\t' + str(param[0].data.shape), str(param[1].data.shape))
# * Since we're dealing with four-dimensional data here, we'll define a helper function for visualizing sets of rectangular heatmaps.
def vis_square(data):
"""Take an array of shape (n, height, width) or (n, height, width, 3)
and visualize each (height, width) thing in a grid of size approx. sqrt(n) by sqrt(n)"""
# normalize data for display
data = (data - data.min()) / (data.max() - data.min())
# force the number of filters to be square
n = int(np.ceil(np.sqrt(data.shape[0])))
padding = (((0, n ** 2 - data.shape[0]),
(0, 1), (0, 1)) # add some space between filters
+ ((0, 0),) * (data.ndim - 3)) # don't pad the last dimension (if there is one)
data = np.pad(data, padding, mode='constant', constant_values=1) # pad with ones (white)
# tile the filters into an image
data = data.reshape((n, n) + data.shape[1:]).transpose((0, 2, 1, 3) + tuple(range(4, data.ndim + 1)))
data = data.reshape((n * data.shape[1], n * data.shape[3]) + data.shape[4:])
plt.imshow(data); plt.axis('off')
# * First we'll look at the first layer filters, `conv1`
# the parameters are a list of [weights, biases]
filters = net.params['conv1'][0].data
vis_square(filters.transpose(0, 2, 3, 1))
# * The first layer output, `conv1` (rectified responses of the filters above, first 36 only)
feat = net.blobs['conv1'].data[0, :36]
vis_square(feat)
# * The fifth layer after pooling, `pool5`
feat = net.blobs['pool5'].data[0]
vis_square(feat)
# * The first fully connected layer, `fc6` (rectified)
#
# We show the output values and the histogram of the positive values
feat = net.blobs['fc6'].data[0]
plt.subplot(2, 1, 1)
plt.plot(feat.flat)
plt.subplot(2, 1, 2)
_ = plt.hist(feat.flat[feat.flat > 0], bins=100)
# * The final probability output, `prob`
feat = net.blobs['prob'].data[0]
plt.figure(figsize=(15, 3))
plt.plot(feat.flat)
# Note the cluster of strong predictions; the labels are sorted semantically. The top peaks correspond to the top predicted labels, as shown above.
# ### 6. Try your own image
#
# Now we'll grab an image from the web and classify it using the steps above.
#
# * Try setting `my_image_url` to any JPEG image URL.
# +
# download an image
my_image_url = "..." # paste your URL here
# for example:
# my_image_url = "https://upload.wikimedia.org/wikipedia/commons/b/be/Orang_Utan%2C_Semenggok_Forest_Reserve%2C_Sarawak%2C_Borneo%2C_Malaysia.JPG"
# !wget -O image.jpg $my_image_url
# transform it and copy it into the net
image = caffe.io.load_image('image.jpg')
net.blobs['data'].data[...] = transformer.preprocess('data', image)
# perform classification
net.forward()
# obtain the output probabilities
output_prob = net.blobs['prob'].data[0]
# sort top five predictions from softmax output
top_inds = output_prob.argsort()[::-1][:5]
plt.imshow(image)
print('probabilities and labels:')
zip(output_prob[top_inds], labels[top_inds])
| examples/00-classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# fundamentals
import os, glob, sys
import numpy as np
import pandas as pd
from calendar import monthrange, month_name
import datetime
import imp
# plotting libraries and setup
from matplotlib.colors import BoundaryNorm
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rc('font', family='serif')
plt.rc('font', size=14)
plt.rc('facecolor', )
# met mast functions and utilities
sys.path.append('../')
import met_funcs as MET
import vis
import utils
# -
# paths (must mount volume smb://nrel.gov/shared/wind/WindWeb/MetData/135mData/)
towerID = 'M5'
datapath = '../example_data/'
figPath = '../../figs/{}'.format(towerID)
try:
os.makedirs(figPath)
except:
pass
# # Data loading
#
# Data located in '../example_data/' has already been filtered orgainzed and qc'd.
#
# If loading raw data (still needing to be qc'd and filtered), refer to 'M5_figs_process.ipyng' for proceedure.
# +
fPaths = os.listdir(datapath)
for ifile, file in enumerate(fPaths):
loadfile = os.path.join(datapath,file)
temp = pd.read_csv(loadfile)
if ifile==0:
metdat = temp
else:
metdat = metdat.append(temp)
# Reindex with date
metdat.set_index('Date', inplace=True)
metdat.index = pd.to_datetime(metdat.index)
metdat.sort_index(inplace=True)
catinfo = MET.get_catinfo(metdat)
# -
nrelcolors = utils.get_nrelcolors()
nrelblue = nrelcolors['blue'][0]
nrelred = nrelcolors['red'][1]
vloc = [40, 87, 130]
for loc in vloc:
fig,ax = vis.hist(metdat, catinfo, 'speed', vertloc=loc, fit='Weibull')
# fig.savefig(os.path.join(figPath,'speed_hist_{}m_fit.png'.format(loc)), bbox_inches='tight', dpi=200)
# +
spdcol, vertloc, _= utils.get_vertical_locations(catinfo['columns']['speed'], location=87)
spdbymonth = metdat[spdcol].groupby(metdat.index.month)
for month in range(1,2):
temp = spdbymonth.get_group(month).dropna()
fig, ax = plt.subplots()
temp.hist(ax=ax, bins=35, color=nrelblue, edgecolor='k')
# fig,ax = vis.hist(temp, catinfo, 'speed',87)
# -
fig,ax = vis.monthly_hist(metdat, catinfo, 'speed', vertloc=87)
imp.reload(vis)
fig,ax = vis.monthly_hist(metdat, catinfo, 'speed', vertloc=87, fit='Weibull')
fig.savefig(os.path.join(figPath,'monthly_speed_hist_fits.png'), bbox_inches='tight', dpi=200)
# +
# fig,ax = vis.hist(metdat, catinfo, 'air pressure', vertloc=38, fit='skewedgaussian')
# -
fig,ax = vis.hist(metdat, catinfo, 'speed', vertloc=87, fit='Weibull')
fig.savefig(os.path.join(figPath,'{}_hist_fits.png'.format(catinfo['save']['speed'])), bbox_inches='tight', dpi=200)
cats = ['air pressure','air temperature','air density']
for cat in cats:
fig,ax = vis.hist(metdat, catinfo, cat, vertloc=87, fit='skewedgaussian')
fig.savefig(os.path.join(figPath,'{}_hist_fits.png'.format(catinfo['save'][cat])), bbox_inches='tight', dpi=200)
# +
months = utils.monthnames()
winter = metdat[(metdat.index.month >= 1) & (metdat.index.month <= 3)]
spring = metdat[(metdat.index.month >= 4) & (metdat.index.month <= 6)]
summer = metdat[(metdat.index.month >= 7) & (metdat.index.month <= 9)]
autumn = metdat[(metdat.index.month >= 10) & (metdat.index.month <= 12)]
# +
fig, ax = plt.subplots(2,2, sharex=True, sharey=True, figsize=(8,6))
dcol,_,_ = utils.get_vertical_locations(catinfo['columns']['speed'], location=87)
for ii in range(4):
startmonth = ii*3+1
stopmonth = startmonth+2
subdat = metdat[(metdat.index.month >= startmonth) & (metdat.index.month <= stopmonth)]
dat1 = subdat[dcol].dropna()
dat1.hist(ax=ax.flatten()[ii], bins=35, facecolor=nrelblue, edgecolor='k', weights=np.ones(len(dat1))/len(dat1))
vis.fit_weibull(dat1, ax=ax.flatten()[ii])
ax.flatten()[ii].grid(False)
ax.flatten()[ii].set_title(r'Q{}'.format(ii+1))
fig.text(0.5,-0.025, catinfo['labels']['speed'], ha='center')
fig.text(-0.025,0.5, 'Frequency [%]', rotation='vertical', va='center')
fig.tight_layout()
# fig.savefig(os.path.join(figPath,'speed_hist_seasonal_fits.png'), bbox_inches='tight', dpi=200)
# +
fig, ax = plt.subplots(2,2, sharex=True, sharey=True, figsize=(8,6))
dcol,_,_ = utils.get_vertical_locations(catinfo['columns']['speed'], location=87)
dat1 = winter[dcol].dropna()
dat1.hist(ax=ax.flatten()[0], bins=35, facecolor=nrelblue, edgecolor='k', weights=np.ones(len(dat1))/len(dat1))
vis.fit_weibull(dat1, ax=ax.flatten()[0])
ax.flatten()[0].grid(False)
ax.flatten()[0].set_title('Jan-Mar')
dat1 = spring[dcol].dropna()
dat1.hist(ax=ax.flatten()[1], bins=35, facecolor=nrelblue, edgecolor='k', weights=np.ones(len(dat1))/len(dat1))
vis.fit_weibull(dat1, ax=ax.flatten()[1])
ax.flatten()[1].grid(False)
ax.flatten()[1].set_title('Apr-Jun')
dat1 = summer[dcol].dropna()
dat1.hist(ax=ax.flatten()[2], bins=35, facecolor=nrelblue, edgecolor='k', weights=np.ones(len(dat1))/len(dat1))
vis.fit_weibull(dat1, ax=ax.flatten()[2])
ax.flatten()[2].grid(False)
ax.flatten()[2].set_title('Jul-Sep')
dat1 = autumn[dcol].dropna()
dat1.hist(ax=ax.flatten()[3], bins=35, facecolor=nrelblue, edgecolor='k', weights=np.ones(len(dat1))/len(dat1))
vis.fit_weibull(dat1, ax=ax.flatten()[3])
ax.flatten()[3].grid(False)
ax.flatten()[3].set_title('Oct-Dec')
fig.text(0.5,-0.025, catinfo['labels']['speed'], ha='center')
fig.text(-0.025,0.5, 'Frequency [%]', rotation='vertical', va='center')
fig.tight_layout()
fig.savefig(os.path.join(figPath,'speed_hist_seasonal_fits.png'), bbox_inches='tight', dpi=200)
# -
imp.reload(vis)
fig,ax = vis.hist(metdat, catinfo, 'direction', vertloc=87, bins=72)
fig.savefig(os.path.join(figPath,'dir_hist.png'), bbox_inches='tight', dpi=200)
# +
fig, ax = plt.subplots(2,2, sharex=True, sharey=True, figsize=(8,6))
dcol,_,_ = utils.get_vertical_locations(catinfo['columns']['direction'], location=87)
dat1 = winter[dcol].dropna()
dat1.hist(ax=ax.flatten()[0], bins=35, facecolor=nrelblue, edgecolor='k', weights=np.ones(len(dat1))/len(dat1))
# vis.fit_weibull(dat1, ax=ax.flatten()[0])
ax.flatten()[0].grid(False)
ax.flatten()[0].set_title('Jan-Mar')
dat1 = spring[dcol].dropna()
dat1.hist(ax=ax.flatten()[1], bins=35, facecolor=nrelblue, edgecolor='k', weights=np.ones(len(dat1))/len(dat1))
# vis.fit_weibull(dat1, ax=ax.flatten()[1])
ax.flatten()[1].grid(False)
ax.flatten()[1].set_title('Apr-Jun')
dat1 = summer[dcol].dropna()
dat1.hist(ax=ax.flatten()[2], bins=35, facecolor=nrelblue, edgecolor='k', weights=np.ones(len(dat1))/len(dat1))
# vis.fit_weibull(dat1, ax=ax.flatten()[2])
ax.flatten()[2].grid(False)
ax.flatten()[2].set_title('Jul-Sep')
dat1 = autumn[dcol].dropna()
dat1.hist(ax=ax.flatten()[3], bins=35, facecolor=nrelblue, edgecolor='k', weights=np.ones(len(dat1))/len(dat1))
# vis.fit_weibull(dat1, ax=ax.flatten()[3])
ax.flatten()[3].grid(False)
ax.flatten()[3].set_title('Oct-Dec')
fig.text(0.5,-0.025, catinfo['labels']['direction'], ha='center')
fig.text(-0.025,0.5, 'Frequency [%]', rotation='vertical', va='center')
fig.tight_layout()
fig.savefig(os.path.join(figPath,'direction_hist_seasonal.png'), bbox_inches='tight', dpi=200)
# -
fig,ax = vis.monthly_hist(metdat, catinfo, 'direction', vertloc=87)
fig.savefig(os.path.join(figPath,'direction_hist_monthly.png'), bbox_inches='tight', dpi=200)
# +
years = metdat.index.year.value_counts().index.values
years.sort()
dcol,_,_ = utils.get_vertical_locations(catinfo['columns']['direction'], location=87)
fig, ax = plt.subplots(3,2, sharex=True, sharey=True, figsize=(8,9))
yeardat = metdat[dcol].groupby(metdat.index.year)
for iy, year in enumerate(years):
dat1 = yeardat.get_group(year).dropna()
dat1.hist(ax=ax.flatten()[iy], bins=35, facecolor=nrelblue, edgecolor='k', weights=np.ones(len(dat1))/len(dat1))
ax.flatten()[iy].set_title(year)
ax.flatten()[iy].grid(False)
ax.flatten()[iy].annotate('N={}'.format(len(dat1)), xy=(0.05,0.8), xycoords='axes fraction')
fig.text(0.5,-0.025, catinfo['labels']['direction'], ha='center')
fig.text(-0.025,0.5, 'Frequency [%]', rotation='vertical', va='center')
fig.tight_layout()
fig.savefig(os.path.join(figPath,'direction_hist_annual.png'), bbox_inches='tight', dpi=200)
# -
imp.reload(vis)
# +
years = metdat.index.year.value_counts().index.values
years.sort()
dcol,_,_ = utils.get_vertical_locations(catinfo['columns']['speed'], location=87)
fig, ax = plt.subplots(3,2, sharex=True, sharey=True, figsize=(8,9))
yeardat = metdat[dcol].groupby(metdat.index.year)
for iy, year in enumerate(years):
dat1 = yeardat.get_group(year).dropna()
dat1.hist(ax=ax.flatten()[iy], bins=35, facecolor=nrelblue, edgecolor='k', weights=np.ones(len(dat1))/len(dat1))
vis.fit_weibull(dat1, ax=ax.flatten()[iy])
ax.flatten()[iy].set_title(year)
ax.flatten()[iy].grid(False)
# ax.flatten()[iy].annotate('N={}'.format(len(dat1)), xy=(0.05,0.8), xycoords='axes fraction')
fig.text(0.5,-0.025, catinfo['labels']['speed'], ha='center')
fig.text(-0.025,0.5, 'Frequency [%]', rotation='vertical', va='center')
fig.tight_layout()
fig.savefig(os.path.join(figPath,'speed_hist_annual_fits.png'), bbox_inches='tight', dpi=200)
# -
# # Cut by TI
#
# +
turbclasses = np.linspace(0,50,6)
turbcol, probe_heights, _ = utils.get_vertical_locations(catinfo['columns']['ti'], 87)
metdat['turbclass'] = pd.cut(metdat[turbcol], turbclasses, include_lowest=False).astype(str)
metdat['turbclass'].value_counts()
temp = metdat.groupby('turbclass')
turbclasses = list(temp.groups.keys())[:-1]
lowTI = temp.get_group(turbclasses[0])
# -
plotcats = ['air density',
'air pressure',
'air temperature',
'direction',
'relative humidity',
'speed',
'wind shear',
'wind veer']
lowtifigpath = '../../figs/lowTI'
try:
os.makedirs(lowtifigpath)
except:
pass
# +
nrelcolors = utils.get_nrelcolors()
imp.reload(vis)
plotcats = ['air density',
'air pressure',
'air temperature',
'direction',
'relative humidity',
'speed',
'wind shear',
'wind veer']
for cat in ['speed']:#plotcats:
height = 87
if 'shear' in cat.lower():
height = 110
plotvar, _, _ = utils.get_vertical_locations(catinfo['columns'][cat], height)
fulldat = metdat[plotvar].dropna()#.sort_values()
# fulldat = MET.reject_outliers(fulldat,m=4)
lowtidat = lowTI[plotvar].dropna()#.sort_values()
# lowtidat = MET.reject_outliers(lowtidat,m=4)
result = pd.concat([fulldat, lowtidat], axis=1)
fig, ax = plt.subplots(figsize=(5,3))
binwidth = np.round((fulldat.max()-fulldat.min())/35.0,decimals=3)
bins = np.arange(fulldat.min(), fulldat.max(), binwidth)
h1 = fulldat.plot.hist(bins = bins,
color=nrelcolors['blue'][0],
edgecolor='k',
alpha=0.5,
ax=ax,
weights=np.ones(len(fulldat))/len(fulldat),
legend=False, label='fulldat')
vis.fit_weibull(fulldat, ax=ax, basecolor='blue', xy = (1.35,0.90))
h2 = lowtidat.plot.hist(bins = bins,
color=nrelcolors['red'][0],
edgecolor='k',
alpha=0.5,
ax=ax,
weights=np.ones(len(lowtidat))/len(lowtidat),
legend=False, label='lowti')
vis.fit_weibull(lowtidat, ax=ax, basecolor='red', xy = (1.35,0.35))
# proxy artists
from matplotlib.patches import Rectangle
handles = [Rectangle((0,0),1,1,color=c,ec="k",alpha=0.5) for c in [nrelblue,nrelred]]
labels= ['Full Data', 'Low TI']
plt.legend(handles, labels,frameon=False, loc=6, bbox_to_anchor=(1,0.5))
ax.set_xlabel(catinfo['labels'][cat])
ax.set_ylabel('Frequency [%]')
# leg = ax.legend((h1, h2),('Full Data', 'Low TI'), frameon=False, loc=6, bbox_to_anchor=(1,0.5))
fig.savefig(os.path.join(lowtifigpath,'LOWTI_{}_hist_comp.png'.format(catinfo['save'][cat])),dpi=200,bbox_inches='tight')
# plt.clf()
# +
nrelcolors = utils.get_nrelcolors()
imp.reload(vis)
plotcats = ['air density',
'air pressure',
'air temperature',
'relative humidity']
for cat in plotcats[2:3]:
height = 87
if 'shear' in cat.lower():
height = 110
plotvar, _, _ = utils.get_vertical_locations(catinfo['columns'][cat], height)
fulldat = metdat[plotvar].dropna().sort_values()
lowtidat = lowTI[plotvar].dropna().sort_values()
result = pd.concat([fulldat, lowtidat], axis=1)
fig, ax = plt.subplots(figsize=(5,3))
binwidth = np.round((fulldat.max()-fulldat.min())/35.0,decimals=3)
bins = np.arange(fulldat.min(), fulldat.max(), binwidth)
h1 = fulldat.plot.hist(bins = bins,
color=nrelcolors['blue'][0],
edgecolor='k',
alpha=0.5,
ax=ax,
weights=np.ones(len(fulldat))/len(fulldat),
legend=False, label='fulldat')
vis.fit_skewedgaussian(fulldat, bins=bins, ax=ax, basecolor='blue', xy = (1.35,1.025), gamma=10)
h2 = lowtidat.plot.hist(bins = bins,
color=nrelcolors['red'][0],
edgecolor='k',
alpha=0.5,
ax=ax,
weights=np.ones(len(lowtidat))/len(lowtidat),
legend=False, label='lowti')
vis.fit_skewedgaussian(lowtidat, bins=bins, ax=ax, basecolor='red', xy = (1.35,0.35))
# proxy artists
from matplotlib.patches import Rectangle
handles = [Rectangle((0,0),1,1,color=c,ec="k",alpha=0.5) for c in [nrelblue,nrelred]]
labels= ['Full Data', 'Low TI']
plt.legend(handles, labels,frameon=False, loc=6, bbox_to_anchor=(1,0.5))
ax.set_xlabel(catinfo['labels'][cat])
ax.set_ylabel('Frequency [%]')
fig.savefig(os.path.join(lowtifigpath,'LOWTI_{}_hist_comp.png'.format(catinfo['save'][cat])),dpi=200,bbox_inches='tight')
# plt.clf()
# +
nrelcolors = utils.get_nrelcolors()
imp.reload(vis)
plotcats = ['air density',
'air pressure',
'air temperature',
'relative humidity']
for cat in plotcats[1:2]:
height = 87
if 'shear' in cat.lower():
height = 110
plotvar, _, _ = utils.get_vertical_locations(catinfo['columns'][cat], height)
fulldat = metdat[plotvar].dropna().sort_values()
lowtidat = lowTI[plotvar].dropna().sort_values()
result = pd.concat([fulldat, lowtidat], axis=1)
fig, ax = plt.subplots(figsize=(5,3))
binwidth = np.round((fulldat.max()-fulldat.min())/35.0,decimals=3)
bins = np.arange(fulldat.min(), fulldat.max(), binwidth)
h1 = fulldat.plot.hist(bins = bins,
color=nrelcolors['blue'][0],
edgecolor='k',
alpha=0.5,
ax=ax,
weights=np.ones(len(fulldat))/len(fulldat),
legend=False, label='fulldat')
vis.fit_skewedgaussian(fulldat, bins=bins, ax=ax, basecolor='blue', xy = (1.35,1.025), gamma=-0.5)
h2 = lowtidat.plot.hist(bins = bins,
color=nrelcolors['red'][0],
edgecolor='k',
alpha=0.5,
ax=ax,
weights=np.ones(len(lowtidat))/len(lowtidat),
legend=False, label='lowti')
vis.fit_skewedgaussian(lowtidat, bins=bins, ax=ax, basecolor='red', xy = (1.35,0.35), gamma=-2)
# proxy artists
from matplotlib.patches import Rectangle
handles = [Rectangle((0,0),1,1,color=c,ec="k",alpha=0.5) for c in [nrelblue,nrelred]]
labels= ['Full Data', 'Low TI']
plt.legend(handles, labels,frameon=False, loc=6, bbox_to_anchor=(1,0.5))
ax.set_xlabel(catinfo['labels'][cat])
ax.set_ylabel('Frequency [%]')
fig.savefig(os.path.join(lowtifigpath,'LOWTI_{}_hist_comp.png'.format(catinfo['save'][cat])),dpi=200,bbox_inches='tight')
# plt.clf()
# +
nrelcolors = utils.get_nrelcolors()
imp.reload(vis)
plotcats = ['air density',
'air pressure',
'air temperature',
'relative humidity']
for cat in plotcats[0:1]:
height = 87
if 'shear' in cat.lower():
height = 110
plotvar, _, _ = utils.get_vertical_locations(catinfo['columns'][cat], height)
fulldat = metdat[plotvar].dropna().sort_values()
lowtidat = lowTI[plotvar].dropna().sort_values()
result = pd.concat([fulldat, lowtidat], axis=1)
fig, ax = plt.subplots(figsize=(5,3))
binwidth = np.round((fulldat.max()-fulldat.min())/35.0,decimals=3)
bins = np.arange(fulldat.min(), fulldat.max(), binwidth)
h1 = fulldat.plot.hist(bins = bins,
color=nrelcolors['blue'][0],
edgecolor='k',
alpha=0.5,
ax=ax,
weights=np.ones(len(fulldat))/len(fulldat),
legend=False, label='fulldat')
vis.fit_skewedgaussian(fulldat, bins=bins, ax=ax, basecolor='blue', xy = (1.35,1.025), gamma=0.5)
h2 = lowtidat.plot.hist(bins = bins,
color=nrelcolors['red'][0],
edgecolor='k',
alpha=0.5,
ax=ax,
weights=np.ones(len(lowtidat))/len(lowtidat),
legend=False, label='lowti')
vis.fit_skewedgaussian(lowtidat, bins=bins, ax=ax, basecolor='red', xy = (1.35,0.35), gamma=0)
# proxy artists
from matplotlib.patches import Rectangle
handles = [Rectangle((0,0),1,1,color=c,ec="k",alpha=0.5) for c in [nrelblue,nrelred]]
labels= ['Full Data', 'Low TI']
plt.legend(handles, labels,frameon=False, loc=6, bbox_to_anchor=(1,0.5))
ax.set_xlabel(catinfo['labels'][cat])
ax.set_ylabel('Frequency [%]')
fig.savefig(os.path.join(lowtifigpath,'LOWTI_{}_hist_comp.png'.format(catinfo['save'][cat])),dpi=200,bbox_inches='tight')
# plt.clf()
# +
nrelcolors = utils.get_nrelcolors()
imp.reload(vis)
plotcats = ['air density',
'air pressure',
'air temperature',
'relative humidity']
for cat in ['relative humidity']:
height = 87
if 'shear' in cat.lower():
height = 110
plotvar, _, _ = utils.get_vertical_locations(catinfo['columns'][cat], height)
fulldat = metdat[plotvar].dropna().sort_values()
lowtidat = lowTI[plotvar].dropna().sort_values()
result = pd.concat([fulldat, lowtidat], axis=1)
fig, ax = plt.subplots(figsize=(5,3))
binwidth = np.round((fulldat.max()-fulldat.min())/35.0,decimals=3)
bins = np.arange(fulldat.min(), fulldat.max(), binwidth)
h1 = fulldat.plot.hist(bins = bins,
color=nrelcolors['blue'][0],
edgecolor='k',
alpha=0.5,
ax=ax,
weights=np.ones(len(fulldat))/len(fulldat),
legend=False, label='fulldat')
vis.fit_skewedgaussian(fulldat, bins=bins, ax=ax, basecolor='blue', xy = (1.35,1.025), gamma=0)
h2 = lowtidat.plot.hist(bins = bins,
color=nrelcolors['red'][0],
edgecolor='k',
alpha=0.5,
ax=ax,
weights=np.ones(len(lowtidat))/len(lowtidat),
legend=False, label='lowti')
vis.fit_skewedgaussian(lowtidat, bins=bins, ax=ax, basecolor='red', xy = (1.35,0.35), gamma=5)
# proxy artists
from matplotlib.patches import Rectangle
handles = [Rectangle((0,0),1,1,color=c,ec="k",alpha=0.5) for c in [nrelblue,nrelred]]
labels= ['Full Data', 'Low TI']
plt.legend(handles, labels,frameon=False, loc=6, bbox_to_anchor=(1,0.5))
ax.set_xlabel(catinfo['labels'][cat])
ax.set_ylabel('Frequency [%]')
fig.savefig(os.path.join(lowtifigpath,'LOWTI_{}_hist_comp.png'.format(catinfo['save'][cat])),dpi=200,bbox_inches='tight')
# plt.clf()
# +
nrelcolors = utils.get_nrelcolors()
height = 87
if 'shear' in cat.lower():
height = 110
cat = 'wind shear'
plotvar = 'Wind Shear (3_ 122 m)'
fulldat = metdat[plotvar].dropna().sort_values()
lowtidat = lowTI[plotvar].dropna().sort_values()
result = pd.concat([fulldat, lowtidat], axis=1)
fig, ax = plt.subplots(figsize=(5,3))
binwidth = np.round((fulldat.max()-fulldat.min())/35.0,decimals=3)
bins = np.arange(fulldat.min(), fulldat.max(), binwidth)
h1 = fulldat.plot.hist(bins = bins,
color=nrelcolors['blue'][0],
edgecolor='k',
alpha=0.5,
ax=ax,
weights=np.ones(len(fulldat))/len(fulldat),
legend=False, label='fulldat')
# vis.fit_skewedgaussian(fulldat, bins=bins, ax=ax, basecolor='blue', xy = (1.35,1.025), gamma=0)
h2 = lowtidat.plot.hist(bins = bins,
color=nrelcolors['red'][0],
edgecolor='k',
alpha=0.5,
ax=ax,
weights=np.ones(len(lowtidat))/len(lowtidat),
legend=False, label='lowti')
# vis.fit_skewedgaussian(lowtidat, bins=bins, ax=ax, basecolor='red', xy = (1.35,0.35), gamma=5)
# proxy artists
from matplotlib.patches import Rectangle
handles = [Rectangle((0,0),1,1,color=c,ec="k",alpha=0.5) for c in [nrelblue,nrelred]]
labels= ['Full Data', 'Low TI']
plt.legend(handles, labels,frameon=False, loc=6, bbox_to_anchor=(1,0.5))
ax.set_xlabel(catinfo['labels'][cat])
ax.set_ylabel('Frequency [%]')
fig.savefig(os.path.join(lowtifigpath,'LOWTI_{}_hist_comp.png'.format(catinfo['save'][cat])),dpi=200,bbox_inches='tight')
# plt.clf()
# +
nrelcolors = utils.get_nrelcolors()
height = 87
if 'shear' in cat.lower():
height = 110
cat = 'wind veer'
plotvar = 'Wind Veer (3_ 122 m)'
fulldat = metdat[plotvar].dropna().sort_values()
lowtidat = lowTI[plotvar].dropna().sort_values()
result = pd.concat([fulldat, lowtidat], axis=1)
fig, ax = plt.subplots(figsize=(5,3))
binwidth = np.round((fulldat.max()-fulldat.min())/35.0,decimals=3)
bins = np.arange(fulldat.min(), fulldat.max(), binwidth)
h1 = fulldat.plot.hist(bins = bins,
color=nrelcolors['blue'][0],
edgecolor='k',
alpha=0.5,
ax=ax,
weights=np.ones(len(fulldat))/len(fulldat),
legend=False, label='fulldat')
# vis.fit_skewedgaussian(fulldat, bins=bins, ax=ax, basecolor='blue', xy = (1.35,1.025), gamma=0)
h2 = lowtidat.plot.hist(bins = bins,
color=nrelcolors['red'][0],
edgecolor='k',
alpha=0.5,
ax=ax,
weights=np.ones(len(lowtidat))/len(lowtidat),
legend=False, label='lowti')
# vis.fit_skewedgaussian(lowtidat, bins=bins, ax=ax, basecolor='red', xy = (1.35,0.35), gamma=5)
# proxy artists
from matplotlib.patches import Rectangle
handles = [Rectangle((0,0),1,1,color=c,ec="k",alpha=0.5) for c in [nrelblue,nrelred]]
labels= ['Full Data', 'Low TI']
plt.legend(handles, labels,frameon=False, loc=6, bbox_to_anchor=(1,0.5))
ax.set_xlabel(catinfo['labels'][cat])
ax.set_ylabel('Frequency [%]')
fig.savefig(os.path.join(lowtifigpath,'LOWTI_{}_hist_comp.png'.format(catinfo['save'][cat])),dpi=200,bbox_inches='tight')
# plt.clf()
# -
params.keys()
# +
nbins = 100
data = metdat['Wind Veer (3_ 87 m)'].dropna()
binwidth = np.round((data.max()-data.min())/nbins,decimals=3)
bins = np.arange(data.min(), data.max(), binwidth)
yvals, xvals = np.histogram(data, bins=bins)
xvals = np.array([(xvals[i]+xvals[i+1])/2 for i in range(len(xvals)-1)])
params = model.make_params(sigma=np.std(data), center=np.mean(data), amplitude=1000000)
result = model.fit(yvals, params, x=xvals)
fig, ax = plt.subplots()
data.hist(ax=ax, color = nrelblue, edgecolor='k', weights=np.ones(len(data))/len(data), bins=nbins)
ax.plot(xvals, result.best_fit* 1.0/float(len(data)), color=nrelred, linewidth=2.5)
# +
cats = catinfo['columns']['wind shear']
data = metdat[cats[1]].dropna()
fig, ax = plt.subplots(figsize=(5,3))
data.hist(ax=ax, color = nrelblue, edgecolor='k', weights=np.ones(len(data))/len(data), bins=35)
# ax.set_title(cat)
ax.grid(False)
ax.set_xlabel(catinfo['labels']['wind shear'])
ax.set_ylabel('Frequency [%]')
fig.savefig(os.path.join(figPath,'{}_3_122.png'.format(catinfo['save']['wind shear'])), dpi=200, bbox_inches='tight')
# fig, ax = vis.hist(metdat, catinfo, 'wind shear', vertloc=38)
# yvals, xvals = np.histogram(data, bins=100)
# x = np.array([np.sum(xvals[ii:ii+1]) for ii in range(len(xvals[:-1]))])
# lower = np.argwhere(yvals < data.mean()-data.std())
# test = np.trapz(data)
# -
lower = np.argmin(np.abs(yvals - (data.mean()-data.std())))
data.mean()
print(np.round(data.mean(),3),np.round(data.std(),3))
# +
cats = catinfo['columns']['wind veer']
data = metdat[cats[2]].dropna()
fig, ax = plt.subplots(figsize=(5,3))
data.hist(ax=ax, color = nrelblue, edgecolor='k', weights=np.ones(len(data))/len(data), bins=35)
# ax.set_title(cat)
ax.grid(False)
ax.set_xlabel(catinfo['labels']['wind veer'])
ax.set_ylabel('Frequency [%]')
fig.savefig(os.path.join(figPath,'{}_3_122.png'.format(catinfo['save']['wind veer'])), dpi=200, bbox_inches='tight')
# -
catinfo['save']['wind shear']
imp.reload(vis)
fig, ax = vis.normalized_monthly_hist_by_stability(metdat,catinfo)
| notebooks/.ipynb_checkpoints/M5_hist_w_fits-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import pyNN.nest as p
from pyNN.random import NumpyRNG, RandomDistribution
from pyNN.utility import Timer
import pylab
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
timer = Timer()
ts = 0.1
simulation_time = 100
# +
def setup(simulation_time, ts):
#timer = Timer()
#p.setup(timestep=ts, min_delay=ts, max_delay=2.0*ts) # 0.1ms
p.setup(timestep=ts) # 0.1ms
# -
def compute_network_state(n_res, w_exc_b, w_inh_b, rout_w_exc, rout_w_inh, inp_rate):
n_input_neurons = 10
n_readout_neurons = 2
n_reservoir_neurons = n_res
exc_rate = 0.8 # % of excitatory neurons in reservoir
n_exc = int(round(n_reservoir_neurons*exc_rate))
n_inh = n_reservoir_neurons-n_exc
izh_celltype = p.native_cell_type('izhikevich')
if_celltype = p.IF_curr_exp
celltype = if_celltype
spike_source = p.native_cell_type('poisson_generator')
inp_pop=p.Population(n_input_neurons,spike_source,{'rate':inp_rate, 'start':0.0,'stop':40.0})
#exc_cells = p.Population(n_exc, celltype,cellparams={'tau_refrac':1.}, label="Excitatory_Cells")
exc_cells = p.Population(n_exc, celltype, label="Excitatory_Cells")
inh_cells = p.Population(n_inh, celltype, label="Inhibitory_Cells")
# initialize with a uniform random distributin
# use seeding for reproducability
rngseed = 98766987
parallel_safe = True
rng = NumpyRNG(seed=rngseed, parallel_safe=parallel_safe)
unifDistr = RandomDistribution('uniform', (-70,-65), rng=rng)
inh_cells.initialize('V_m',unifDistr)
exc_cells.initialize('V_m',unifDistr)
readout_neurons = p.Population(2, celltype, label="readout_neuron")
inp_weight=3.
inp_delay =1
inp_weight_distr = RandomDistribution('normal', [inp_weight, 1e-3], rng=rng)
# connect each input neuron to 30% of the reservoir neurons
inp_conn = p.FixedProbabilityConnector(p_connect=0.3/np.sqrt(n_res)*np.sqrt(200),weights =inp_weight_distr, delays=inp_delay)
connections = {}
connections['inp2e'] = p.Projection(inp_pop, exc_cells, inp_conn)
connections['inp2i'] = p.Projection(inp_pop, inh_cells, inp_conn)
pconn = 0.05 # sparse connection probability
# scale the weights w.r.t. the network to keep it stable
w_exc = w_exc_b/np.sqrt(n_res) # nA
w_inh = w_inh_b/np.sqrt(n_res) # nA
# without weight scaling
#w_exc = w_exc_b/np.sqrt(200) # nA
#w_inh = w_inh_b/np.sqrt(200) # nA
delay_exc = 1 # defines how long (ms) the synapse takes for transmission
delay_inh = 1
weight_distr_exc = RandomDistribution('normal', [w_exc, 1/n_res], rng=rng)
weight_distr_inh = RandomDistribution('normal', [w_inh, 1/n_res], rng=rng)
exc_conn = p.FixedProbabilityConnector(pconn, weights=weight_distr_exc, delays=delay_exc)
inh_conn = p.FixedProbabilityConnector(pconn, weights=weight_distr_inh, delays=delay_inh)
connections['e2e'] = p.Projection(exc_cells, exc_cells, exc_conn, target='excitatory')
connections['e2i'] = p.Projection(exc_cells, inh_cells, exc_conn, target='excitatory')
connections['i2e'] = p.Projection(inh_cells, exc_cells, inh_conn, target='inhibitory')
connections['i2i'] = p.Projection(inh_cells, inh_cells, inh_conn, target='inhibitory')
# Eigenvalue analysis for stability
weights_e2e = connections['e2e'].getWeights(format='array')
weights_e2i = connections['e2i'].getWeights(format='array')
weights_i2e = connections['i2e'].getWeights(format='array')
weights_i2i = connections['i2i'].getWeights(format='array')
a=np.concatenate((weights_e2e,weights_i2e), axis=0)
b=np.concatenate((weights_e2i,weights_i2i), axis=0)
weight_matrix=np.concatenate((a,b), axis=1)
np.nan_to_num(weight_matrix, copy=False)
eigenvalues, eigenvectors=np.linalg.eig(weight_matrix)
# uncomment the next line if you want to remove the oulier on the real-axis
#eigenvalues = eigenvalues[np.where(eigenvalues.imag != 0)]
plt.scatter(eigenvalues.real, eigenvalues.imag)
'''
weights_e2e = connections['e2e'].getWeights(format='array')
np.nan_to_num(weights_e2e, copy=False)
eigenvalues, eigenvectors=np.linalg.eig(weights_e2e)
plt.scatter(eigenvalues.real, eigenvalues.imag)
'''
rout_conn_exc = p.AllToAllConnector(weights=rout_w_exc, delays=delay_exc)
rout_conn_inh = p.AllToAllConnector(weights=rout_w_inh, delays=delay_exc)
connections['e2rout'] = p.Projection(exc_cells, readout_neurons, rout_conn_exc, target='excitatory')
connections['i2rout'] = p.Projection(inh_cells, readout_neurons, rout_conn_inh, target='inhibitory')
readout_neurons.record()
exc_cells.record()
inh_cells.record()
inp_pop.record()
p.run(simulation_time)
r_spikes = readout_neurons.getSpikes()
exc_spikes = exc_cells.getSpikes()
inh_spikes = inh_cells.getSpikes()
inp_spikes = inp_pop.getSpikes()
#p.end()
return r_spikes, exc_spikes, inh_spikes, inp_spikes
def plot_spikes(spiketrain):
st = [spike[1] for spike in spiketrain] # spike times
sid = [spike[0] for spike in spiketrain]
plt.plot(st, sid, "|")
plt.axis([np.min(st), 100, np.min(sid)-1, np.max(sid)+1])
plt.xlabel('Time (ms)')
plt.ylabel('Neuron ID')
plt.title('Spike Plot')
plt.xlim(xmin=0)
plt.show()
# # Define and Run Network
# +
n_res=2000
w_exc_b=2.828
w_inh_b=-11.312
rout_w_exc=20
rout_w_inh=-80
inp_rate=100.0
ts = 0.1
simulation_time = 100
setup(simulation_time,ts)
timer = Timer()
r_spikes, exc_spikes, inh_spikes, inp_spikes = compute_network_state(n_res, w_exc_b, w_inh_b, rout_w_exc, rout_w_inh, inp_rate)
timer.elapsedTime()
# -
plot_spikes(exc_spikes)
plot_spikes(r_spikes)
plot_spikes(inp_spikes)
st = [spike[1] for spike in r_spikes]
st[:10]
r_spikes[:10]
np.shape(r_spikes)
np.shape(exc_spikes)
def get_spike_rates(spikes):
"""returns the rates of all neurons in the recorded population"""
shape=np.shape(spikes)
spike_rates=np.zeros((int(np.max(spikes[:,0]))+1,))
for i in range(shape[0]): # for each spikes
for j in range(int(np.max(spikes[:,0]))+1): #for each neuron
if spikes[i,0]==j:
spike_rates[j]+=1
return spike_rates
spike_rates=get_spike_rates(r_spikes)
spike_rates
res_rates=get_spike_rates(exc_spikes)
res_rates
# +
# separability: different input signals (1. rates, 2. time series of rates)
# must lead to different reservoir state vectors
# needs a distance metric
# +
n_res=200
w_exc_b=2
w_inh_b=-8
rout_w_exc=20
rout_w_inh=-80
inp_rate=20.0
r_spikes2, exc_spikes2 = compute_network_state(n_res, w_exc_b, w_inh_b, rout_w_exc, rout_w_inh, inp_rate)
# -
| src/experimental_code/stability_lsm/.ipynb_checkpoints/compute_reservoir_state-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import gym
env = gym.make("FrozenLake8x8-v0").env
"""
Documentation from https://gym.openai.com/envs/FrozenLake-v0/
SFFF (S: starting point, safe)
FHFH (F: frozen surface, safe)
FFFH (H: hole, fall to your doom)
HFFG (G: goal, where the frisbee is located)
"""
# reset and display environment
env.reset()
env.render()
# actions: 4 possible actions ["Left", "Down", "Right", "Up"]
# https://github.com/openai/gym/blob/master/gym/envs/toy_text/frozen_lake.py
print("Action Space {}".format(env.action_space))
# states: 8x8=64 possible positions for the agent
print("State Space {}".format(env.observation_space))
action = 1
state, reward, done, info = env.step(action)
env.render()
print(state, reward, done, info)
# Reward table called P: states x actions
# {action: [(probability, nextstate, reward, done)]}
# probability of taking the action can be lower than one:
# As the ice is slippery, the move won't always be the intended one
env.P[19]
# ## Run random moves and print animated result
# +
env.reset()
imgs = []
done = False
while not done:
action = env.action_space.sample()
state, reward, done, info = env.step(action)
imgs.append({
'frame': env.render(mode='ansi'),
'state': state,
'action': action,
'reward': reward,
'done': done
}
)
# +
from IPython.display import clear_output
from time import sleep
def print_frames(frames, sleep_time=0.1):
for i, img in enumerate(imgs):
clear_output(wait=True)
print(img['frame'])
print("Step: {}".format(i))
print("State: {}".format(img['state']))
print("Action: {}".format(img['action']))
print("Reward: {}".format(img['reward']))
print("Done: {}".format(img['done']))
sleep(sleep_time)
# -
print_frames(imgs)
# ## Create q-table
import numpy as np
q_table = np.zeros([env.observation_space.n, env.action_space.n])
# ## Fill q-table
# +
import random
# parameter for q-table
alpha = 0.1
gamma = 0.9
epsilon = 0.4
for i in range(200000):
state = env.reset()
done = False
while not done:
# probability of taking a random action
if random.uniform(0, 1) < epsilon:
action = env.action_space.sample()
# take the action that lead to better reward
else:
action = np.argmax(q_table[state])
# do a move
next_state, reward, done, info = env.step(action)
## set specific reward to add penalties
# if in a hole
if done and reward != 1:
reward = -10
# if on a frozen position
elif reward == 0:
reward = -0.5
# if goal reached
else:
reward = 20
# get current reward for the given action in the current state
current_expected_reward = q_table[state, action]
# get max possible calculated reward from next state
next_state_max_reward = np.max(q_table[next_state])
# update q-table with formula
updated_reward = (1 - alpha) * current_expected_reward + alpha * (reward + gamma * next_state_max_reward)
q_table[state, action] = updated_reward
state = next_state
if i % 100 == 0:
clear_output(wait=True)
print(f"Try n°{i}")
# +
env.reset()
imgs = []
done = False
while not done:
action = np.argmax(q_table[state])
state, reward, done, info = env.step(action)
imgs.append({
'frame': env.render(mode='ansi'),
'state': state,
'action': action,
'reward': reward,
'done': done
}
)
# -
print_frames(imgs)
| q_learning/qlearning_FrozenLake.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Quantum Variational Simulator
# ## Summary
# The usual method to simulate quantum evolution in quantum systems is using the Trotter expansion. However, this usually requires circuits with many quantum operations and a large depth, which is not desirable in the current noisy era of quantum computation. An alternative is to use Variational Quantum Simulation, since the depth of the circuit is fixed during all the computation and we can choose variational forms that are efficient for different hardwares.
#
# In this project, we implement a Python package that, given a Hamiltonian, a variational form and a state at $t=0$, gives us the state of the system for a future state at time $t = T$ using a variational method. In the tutorial, we present the theory of variational quantum simulation and continue with the details of our implementation using Qiskit. Then, we show simulations for the evolution of quantum Isings models of 2 and 3 qubits and compare with experimental realizations using IBM's quantum processors and Qiskit Runtime. The results show an excellent agreement between theory and experiment.
# ## 1. Introduction
# The problem of simulating quantum many-body systems is well-known to be extremely challenging for classical computers. This is due to the exponential dimension of the associated Hilbert space. For these systems, it is impossible to diagonalize the Hamiltonian to find the ground state energy, or to solve the Schrödinger equation to obtain its time evolution. Variational Classical Simulation (VCE) $[$[1](https://en.wikipedia.org/wiki/Variational_method_(quantum_mechanics)), [5](https://www.sciencedirect.com/science/article/abs/pii/0003491688902801?via%3Dihub), [6](https://doi.org/10.1103/PhysRevLett.107.070601)$]$ is an alternative to these direct methods because effectively reduces the dimension of systems using efficient classical representations. It can be used to find the ground state energy and to simulate time evolution of quantum systems. However, for highly-entangled systems VQE is usually infeasible, because there are not good classical representations of these states.
#
# The efficient alternative is using quantum systems to simulate quantum systems. The Trotterization technique is the usual method to perform quantum evolution, and approximates the time evolution under a Hamiltonian $H = \sum_j H_j$ from $0$ to $T$ by
#
# $$
# e^{-i H T} \approx \left( \prod_j e^{-i H_j \Delta t}\right)^N,
# $$
#
# where $e^{-i H_j \Delta t}$ is the unitary generated by $H_j$ for a short time $\Delta t = T/N$. Variational Quantum Algorithms (VQA) are the quantum version of VCE and they can be used to find the ground state energy of relevant Hamiltonians using efficient variational forms and classical optimization. These algorithms can also be used as an alternative for Trotterization, that usually requires circuits with many quantum operations and a large depth. This is not desirable in the current era of quantum computation, because quantum processors are still noisy and the depth of the circuits is restricted by the decoherence time of the system. For VQA, the depth of the circuit is fixed during all the computation and we can choose variational forms that are efficient for different hardwares.
#
# In this project, we present a Python package that, given a Hamiltonian, a variational form and a state at $t=0$, gives us the state of the system for a future state at time $t = T$ using the variational method presented in the paper [Efficient Variational Quantum Simulator Incorporating Active Error Minimization](https://doi.org/10.1103/PhysRevX.7.021050). We start this tutorial with the theory of variational quantum simulation and continue with the details of our implementation using Qiskit. Then, we show simulations for the evolution of quantum Isings models of 2 and 3 qubits and compare with experimental realizations using IBM's quantum processors and Qiskit Runtime. Finally, we discuss our results and propose future improvements of the package.
# ## 2. The time-dependent variational principle
# This a variational principle that, when applied to a system, yields the equations of motion for that system $[$[2](https://en.wikipedia.org/wiki/Stationary-action_principle)$]$. It is given by
#
# $$\delta \int_{t_{i}}^{t_{f}} dt L=0,$$
#
# where in the case of the Schrödinger equation the Lagrangian is $L=\langle\Psi(t)|\left(i \frac{\partial}{\partial t}-H\right)| \Psi(t)\rangle$. Now, we assume that the state $| \Psi(t)\rangle$ is determined by a set of real parameters $\left\{\lambda_{k}(t)\right\}$, that is $|\Psi(t)\rangle \equiv\left|\Psi\left(\lambda_{1}, \lambda_{2}, ..., \lambda_{N}\right)\right\rangle$. We want to obtain differential equations for the evolution of the system as a function of these parameters. The Lagrangian can be rewritten as
#
# $$
# L=i \sum_{k}\langle\Psi| \frac{\partial|\Psi\rangle}{\partial \lambda_{k}} \dot{\lambda}_{k}-\langle\Psi|H| \Psi\rangle
# $$
#
# which is a function of parameters $\{\lambda_{k}\}$ and their time derivatives $\left\{\dot{\lambda}_{k}=\left[d \lambda_{k} / d t\right]\right\}$. After applying the [Euler-Lagrange equations](https://en.wikipedia.org/wiki/Euler–Lagrange_equation#Statement), the evolution of the parameters is given by
#
# $$
# \sum_{q} M_{k, q} \dot{\lambda}_{q}=V_{k}
# $$
#
# where
#
# $$
# M_{k, q}=i \frac{\partial\langle\Psi|}{\partial \lambda_{k}} \frac{\partial|\Psi\rangle}{\partial \lambda_{q}}+\text { H.c. } \quad \text{and} \quad V_{k}= \frac{\partial\langle\Psi|}{\partial \lambda_{k}} H|\Psi\rangle+\text { H.c. }
# $$
#
# We now suppose that we know all the coefficients $M_{k, q}$ and $V_{k}$ for all times. Then, given an initial state $|\Psi(0)\rangle \equiv\left|\Psi\left(\lambda_{1} (0), \lambda_{2} (0), ..., \lambda_{N} (0)\right)\right\rangle$, we can solve the equations for $\{ \dot{\lambda}_k \}$ to find the parameters $\{ \lambda_{k} \}$ at a given time $T$, and then, the evolved state $|\Psi(T)\rangle \equiv\left|\Psi\left(\lambda_{1} (T), \lambda_{2} (T), ..., \lambda_{N} (T)\right)\right\rangle$.
# ## 3. Variational algorithm on a hybrid computer
#
# The variational method avoids the exponential space problem that classical methods have by considering trial states from a physically motivated small subset of the exponentially large Hilbert space. Here we consider trial states that can be directly prepared in the quantum computer; i.e., states can be expressed as $|\Psi\rangle=R|\overline{0}\rangle$, where $|\overline{0}\rangle$ is an initial state of the quantum computer and $R$ is a sequence of $N$ quantum gates determined by parameters $\left\{\lambda_{k}\right\}$, that is
#
# $$
# R=R_{N}\left(\lambda_{N}\right) \ldots R_{k}\left(\lambda_{k}\right) \ldots R_{1}\left(\lambda_{1}\right).
# $$
#
# <!-- If N is smaller than the dimension of the Hilbert space, trial states $|\Psi\rangle$ only span a submanifold of the Hilbert space. Therefore, Eq.\eqref{main} approximates -->
# In conventional quantum computing, each unitary gate involves a small subset of qubits, typically one or two. Thefore, the derivative of the unitary $R_k = \exp \left(\lambda_k {\sum_{i} f_{k, i} \sigma_{k, i}} \right)$ can be efficiently decomposed via
#
# $$
# \frac{d R_{k}}{d \lambda_{k}}=\sum_{i} f_{k, i} R_{k} \sigma_{k, i},
# $$
#
# where $\sigma_{k, i}$ are also unitary operators and $f_{k, i}$ are complex coefficients. For most single- and two-qubit gates $R_{k}$, we find that there are only one or two terms in this expression, and $\sigma_{k, i}$ is also a one-qubit or two-qubit gate.
# <details>
# <summary>Click to Expand: Example</summary>
# For the case when
# $$
# R_{k}(\lambda_{k})=e^{-i \lambda_{k} \sigma / 2},$$
# with a one- or two-qubit Hermitian operator $\sigma$, we have
# $$
# \frac{\partial R_{k}(\lambda_{k})}{\partial \lambda_{k}}=-\frac{i}{2} \cdot \sigma \cdot e^{-i \lambda_{k} \sigma / 2},
# $$
# where $f_{k}=-i / 2$ and $\sigma_{k}=\sigma$.
# </details>
#
# With the definition of the trial state we can rewrite the derivative of the state as
#
# $$
# \frac{\partial|\Psi\rangle}{\partial \lambda_{k}}=\sum_{i} f_{k, i} R_{k, i}|\overline{0}\rangle,
# $$
#
# where
# $$
# R_{k, i}=R_{N} R_{N-1} \cdots R_{k+1} R_{k} \sigma_{k, i} \cdots R_{2} R_{1}.
# $$
#
# Then, based on the definition of the Hamiltonian, the elements of $M$ and $V$ can be expressed as
#
# $$
# M_{k, q}=\sum_{i, j}i\left(f_{k, i}^{*} f_{q, j}\langle\overline{0}|R_{k, i}^{\dagger} R_{q, j}| \overline{0}\rangle\right) \quad \text{and} \quad V_{k}=\sum_{i, j}\left(f_{k, i}^{*} h_{j}\langle\overline{0}|R_{k, i}^{\dagger} \sigma_{j} R| \overline{0}\rangle\right).
# $$
#
# Note that each term is in the form $a \operatorname{Re}(e^{i \theta}\langle\overline{0}|U| \overline{0}\rangle)$ where the amplitude $a$ and phase $\theta$ are determined by either $i f_{k, i}^{*} f_{q, j}$ or $f_{k, i}^{*} h_{j}$, and $U$ is a unitary operator equal to either $R_{k, i}^{\dagger} R_{q, j}$ or $R_{k, i}^{\dagger} \sigma_{j} R$. Such a term can be evaluated using the following quantum circuit
# <img src="img/circuit.png" alt="Drawing" style="width: 600px;"/>
# <font size="2"><center> Figure extracted from the paper "Theory of variational quantum simulation" [8] </center> <font size="5"></font>
# This circuit needs an ancillary qubit initialized in the state $\left(|0\rangle+e^{i \theta}|1\rangle\right) / \sqrt{2}$ and a register initialized in the state $|\overline{0}\rangle$. The ancillary qubit is measured in the $\{|+\rangle,|-\rangle\}$ basis after a sequence of quantum gates on the register and two controlled gates, in which the ancillary qubit is the control qubit. The value of each term is given by $\operatorname{Re}(e^{i \theta}\langle\overline{0}|U| \overline{0}\rangle)= 2P_{+} - 1$, where $P_{+}$ is the probability that the qubit is in the state $| + \rangle$.
# ## 4. Implementation using Qiskit
# ### 4.1 Introduction
#
# The Variational Quantum Simulation theory allows to describe the time evolution of a parameterized state $ |\Psi(t)\rangle = |\Psi (\{\lambda_i(t)\}_{i=1}^N) \rangle$ through the ordinary differential equation (ODE),
# $$ \frac{d\lambda_i}{dt} = \left[M^{-1}\right]_{ik} V_k $$
# where the matrix $M$ and the vector $V$ can be computed by means of the Hamiltonian of the system, and a quantum circuit, as presented on Sec. 2.
#
# In this spirit, we implemented a python library which can solve this ODE by computing the relevant quantities on a real quantum device provided by IBM through Qiskit, and perform a fully working simulation for a quantum system of interest.
# To accomplish this goal, the library which we named `VarQuS`, is organized as follows
# ```
# varqus
# └─── ode.py
# └─── variational_simulation.py
# └─── analytic.py
# └─── integrators.py
# └─── states.py
# └─── utils.py
# ```
# ### 4.2 Define a problem
# In the following, we will briefly describe each part of the library, but in order to give exemples as we go through it, we will begin by defining a system to solve
# ```python
# import numpy as np
#
# # Circuit information
# circuit_coefs = [[-0.5j], [-0.5j, -0.5j]]
# circuit_operators = [["ZZ"], [ "XI", "IX"]]
#
# # System information
# hamiltonian_coefs = [-1j, -0.5j, -0.5j]
# hamiltonian_operators = ["ZZ", "XI", "IX"]
#
# # Initial state and parameters
# initial_state = np.ones(4, dtype=complex)/2
# initial_params = np.array([1.0, 1.0])
# ```
# Note that a time-dependent Hamiltonian could also be provided, for example, with coefficients:
# ```python
# hamiltonian_coefs = lambda t: [0.5*np.sin(t), 2.0*np.cos(t)]
# ```
# ### 4.3 ODE
# The first file, named `ode.py` provides an interface for the final user, a function called `define_vqs_ode`, which takes the information of the circuit and the Hamiltonian of the system, and defines another function, representing the ODE to solve, which can be evaluated for a group of parameters, $\lambda_i(t_n)$ and its corresponding time, $t_n$, to obtain $d\lambda_i/dt$.
# ```python
# from varqus.ode import define_vqs_ode
#
# vqs_analytic = define_vqs_ode(circuit_operators, hamiltonian_operators, circuit_coefs, hamiltonian_coefs, initial_state, backend='analytic')
# vqs_simulated = define_vqs_ode(circuit_operators, hamiltonian_operators, circuit_coefs, hamiltonian_coefs, initial_state, shots=2**13)
# ```
# Note that this function takes a keyword `backend`, which accepts a qiskit backend and defaults to the Qiskit Aer simulator, but can be changed to run on a real quantum device. If `backend='analytic'` is passed, then the system is solved exactly by means of linear algebra operations instead of running in a backend. For any backend different to `'analytic'`, the number of shots must be provided with the `shots` keyword.
# Similarly, `ode.py` also provides a function called `define_schrodinger_ode` which takes the information of the system and defines an ODE, corresponding to the time-dependent Schrodinger equation.
# ```python
# from varqus.ode import define_schrodinger_ode
#
# schrodinger_ode = define_schrodinger_ode(hamiltonian_operators, hamiltonian_coefs)
# ```
# ### 4.4 Variational simulation
#
# The file `variational_simulation.py` provides all the methods required to calculate the matrix `M` and the vector `V` in a quantum circuit. This tools are managed internally by the ODE defined, and therefore, they are not of interest for the final user.
# ### 4.5 Analytic
#
# The file `analytic.py` provides the methods to calculate `M` and `V` analytically. That is, the Variational Quantum Simulation theory is used and the equations are directly solved, instead of running them in a backend. Just like `variational_simulation.py`, the methods defined on this file are not of interest for the final user.
# ### 4.6 Integrators
#
# The file `integrators.py` provide integrators which can take the ODEs previously defined and solve it for some time span. As of now, two integrators are implemented: the first order `euler` and the forth order Runge-Kutta `rk4`. Both integrators accept the same arguments.
# ```python
# # Set time discretization
# dt = 0.01 # time step
# Nt = 10 # number of time steps
#
# from varqus.integrators import euler, rk4
#
# # Here we solve the previously posed ODE's
# vqs_analytic_evolved = euler(vqs_analytic, initial_params, dt, Nt)
# vqs_simulated_evolved = euler(vqs_simulated, initial_params, dt, Nt)
#
# # We can solve the Schrodinger ODE too (Remember the Schrodinger equation acts over the state, not the parameters)
# schrodinger_evolved = euler(schrodinger_ode, initial_state, dt, Nt)
# ```
# ### 4.7 States
# The file `states.py` provides some basic tools to work on states. They are useful to the end user.
# ```python
# from varqus.states import state_infidelity, state_from_parameters
#
# # With 'state_from_parameters' we can recover the parameters
# simulated_states = [state_from_parameters(params, circuit_operators, circuit_coefs, initial_state) for params in vqs_simulated_evolved]
# analytic_states = [state_from_parameters(params, circuit_operators, circuit_coefs, initial_state) for params in vqs_analytic_evolved]
#
# # State infidelity is a distance between states. If both states are the same, their infidelity will be zero.
# vqs_infidelities = [state_infidelity(simulated_states[t], schrodinger_evolved[t]) for t in range(Nt)]
# ```
# ### 4.8 Utils
#
# The file `utils.py` provides utilities needed by the rest of the library, for example, to parse the matrix form of the operator from their name.
# ## 5. Variational Quantum Simulations for the 2 qubits quantum Ising model
# +
import numpy as np
import matplotlib.pyplot as plt
import h5py
from varqus.ode import define_vqs_ode, define_schrodinger_ode
from varqus.integrators import euler
from varqus.states import state_from_parameters, state_infidelity
# -
# Following $[$[8](https://doi.org/10.1103/PhysRevLett.107.070601)$]$, we test our variational algorithm for the quantum Ising model $[$[3](https://en.wikipedia.org/wiki/Ising_model)$]$. The Hamiltonian of the model is given by:
#
# $$
# H = H_{z} + H_{x} = -J \sum_{j=1}^{n_{\mathrm{s}}} \sigma_{j}^{\mathrm{z}} \sigma_{j+1}^{\mathrm{z}} - B\sum_{j=1}^{n_{\mathrm{s}}} \sigma_{j}^{\mathrm{x}}
# $$
#
# where $n_s$ is the number of spins, $J$ is the interaction between nearest neighbours spins and $B$ is an external magnetic field. The operators $\sigma^{x}_{j}$ and $\sigma^{z}_{j}$ are the $X$ and $Z$ Pauli operators acting on the $j$ site, and every pair is counted once. For the 2 spins case $n_{s}=2$, and taking the constants $J=B=\frac{1}{2}$, the Hamiltonian becomes:
#
# $$
# H = H_{z} + H_{x} = -\frac{1}{2} \left( ZZ\right) -\frac{1}{2} \left(XI + IX \right)
# $$
# The trial state $|\Psi\rangle$ for the hybrid algorithm is
#
# $$
# |\Psi (\lambda_1, \lambda_2) \rangle = R(\lambda_1, \lambda_2) |\Phi(0)\rangle = e^{i \lambda_{1} H_{x}} e^{i \lambda_{2} H_{z}} |\Phi(0)\rangle
# $$
#
# where $R(\lambda_1, \lambda_2)$ is a sequence of quantum gates determined by parameters $(\lambda_1, \lambda_2)$ and the initial state $|\Phi(0)\rangle$ is a one-dimensional cluster state given by
#
# $$
# |\Phi(0)\rangle = H_{1} H_{2} |00\rangle = \frac{1}{2} \left( |00\rangle + |01\rangle + |10\rangle + |11\rangle \right)
# $$
#
# where $H_i$ is the Hadamard gates acting on the $i$-th qubit.
# First, we define the proper parameters and operators to construct the Hamiltonian $H$, the trial state $|\Psi\rangle$ and the initial state $|\Phi(0)\rangle$.
# +
J = 1/2
B = 1/2
# Hamiltonian parameters and operators
hs = [-J, -B, -B]
opsH = ["ZZ", "XI", "IX"]
# Rotations parameters and operators
fs = [[-1j*J], [-1j*B, -1j*B]]
ops = [["ZZ"], ["XI", "IX"]]
# Initial state
ini_state = np.ones(4, dtype=complex)/2
# -
# Then, we fix the time step $\delta t$, the number of steps $N_t$ and the initial parameters $\lambda_{1}(0), \lambda_{2}(0)$ for the trial state
# +
# Time discretization and initial parameters for the trial state
params_init = np.array([1.0, 1.0])
dt = 0.01
Nt = 100
# -
# Now, we execute our algorithm, first defining the ordinary diferential equation using the variational quantum simulator, and then solving it using a numerical Euler's method, to obtain the parameters in a posterior time $\left\{\lambda_{k}\left(t_{n}\right)\right\}$.
# +
# Define the ode as a function of the parameters
# ode = define_vqs_ode(ops, opsH, fs, hs, ini_state, shots=2**13)
# # Integrate in time using Euler's method
# lambda_k_t = euler(ode, params_init, dt, Nt)
# +
# If you don't want to run the full algorithm, an example is attached on the file "params_2qubits.hdf5".
# Comment the two previous line and uncomment the next two lines
# Load simulated parameters
params_sim = h5py.File('data/params_2qubits.hdf5', 'r')
lambda_k_t = params_sim["params"][:]
# +
# Plot parameter evolution
plt.plot(dt*np.arange(Nt), lambda_k_t, label = [f"param {i}" for i in range(lambda_k_t.shape[1])])
plt.xlabel("Time")
plt.ylabel("Parameters")
plt.legend(loc="best")
plt.title("Parameter Evolution using the Variational Quantum Simulator")
# -
# We want to compare our results to the actual time evolution of the state obtained solving the Schrödinger equation. First, we need to make sure that for the initial parameters, the trial state $|\Psi (\lambda_1(0), \lambda_2(0)) \rangle$ is equivalent to the initial state for $t=0$. We will fix then as initial state for the evolution $|\Phi_s (0) \rangle$ such that
#
# $$
# |\Phi_s (0) \rangle = e^{i \lambda_1(0) H_{x}} e^{i \lambda_2(0) H_{z}} |\Phi (0) \rangle = |\Psi (\lambda_1(0), \lambda_2(0)) \rangle
# $$
#
# and the state evolution is then given by
#
# $$
# |\Phi_s (t) \rangle = e^{-i H t} |\Phi_s (0) \rangle
# $$
initial_state_s = state_from_parameters(params_init, ops, fs, ini_state)
# The Schrödinger equation is then solved numerically using the Euler's method
# +
# Solve the schrodinger equation numerically
schrodinger = define_schrodinger_ode(opsH, hs)
numeric_evolution = euler(schrodinger, initial_state_s, dt, Nt)
# -
# Lastly, we compare how close are the states obtained from solving the Schrödinger equation to the states obtained through the variational algorithm using the quantum state fidelity [[4]](https://en.wikipedia.org/wiki/Fidelity_of_quantum_states), which for pure states is given by
#
# $$
# F(|\psi\rangle, |\phi\rangle) = \left|\left\langle\psi | \phi\right\rangle\right|^{2}
# $$
#
# Then, we define the infidelity between two quantum states simply as
#
# $$
# I(|\psi\rangle, |\phi\rangle) = 1 - F(|\psi\rangle, |\phi\rangle) = 1 - \left|\left\langle\psi | \phi\right\rangle\right|^{2}
# $$
#
# As two states are equivalent up to a global phase if the infidelity between them is 0, we can say that two states are closer as the infidelity gets lower.
# +
# From params to the state psi(lambda)
psi_lambda_t = [state_from_parameters(lambda_k_t[t], ops, fs, ini_state) for t in range(Nt)]
# Calculate infidelity from the circuit vqs with the schrodinger equation solution using matrix exponentiation
inf_array = [state_infidelity(numeric_evolution[t], psi_lambda_t[t]) for t in range(Nt)]
plt.plot(dt*np.arange(Nt), inf_array)
plt.xlabel("Time")
plt.ylabel("Infidelity")
plt.yscale("log")
plt.title("Infidelity between the obtained state from the VQS and the Schrödinger evolution")
# -
# From the above figure, it's possible to see that our algorithm successfully simulates the state evolution, achieving at worst an infidelity of the order of $10^{-5}$. For the 3 qubits simulation, refer to the `3qubits_simulation_tutorial.ipynb` notebook.
# ## 6. Experiments for quantum Ising model
# Finally, we performed an experiment on the ibmq_lima device with the qiskit-runtime architecture, using the same parameters as in the simulation.
# +
# First, we load the experimental data
exp_lambda_k_t = np.load('data/params_2qubits_experiment.npy')
# -
# Next, we plot the parameters evolution
plt.plot(dt*np.arange(Nt), exp_lambda_k_t, label = [f"param {i}" for i in range(exp_lambda_k_t.shape[1])])
plt.xlabel("Time")
plt.ylabel("Parameters")
plt.legend(loc="best")
plt.title("Parameter Evolution using the Variational Quantum Simulator")
# Then we calculate the infidelity between the state from the VQS experiment and the Schrödinger evolution, and compare it to the simulation results.
# +
psi_lambda_exp_t = [state_from_parameters(exp_lambda_k_t[t], ops, fs, ini_state) for t in range(Nt)]
inf_array_exp = [state_infidelity(numeric_evolution[t], psi_lambda_exp_t[t]) for t in range(Nt)]
plt.plot(dt*np.arange(Nt), inf_array, label ="Simulation")
plt.plot(dt*np.arange(Nt), inf_array_exp, label = "Experiment")
plt.xlabel("Time")
plt.ylabel("Infidelity")
plt.yscale("log")
plt.legend()
plt.title("Infidelity between the obtained state from the VQS and the Schrödinger evolution")
# -
# Even considering the errors asociated with the quantum hardware, our algorithm is capable of correctly simulating the state evolution, achieving at worst an infidelity of the order of $10^{-2}$.
# ## 7. Discussion and outlook
# We implemented a package using Qiskit to simulate the time evolution of quantum systems. The algorithm is of the variational type, so it can be implemented with quantum circuits of much less depth compared to the Trotter algorithm. The evolution of the state using both the simulator and the experimental results show a close agreement with the real state of the system. This is shown using the fidelity between states.
# There are some improvements that could be made on the package.
# - We are currently implementing the variational forms using the Qiskit function <i>HamiltonianGate </i> applied to a string of Paulis $\sigma_k$ and a parameter $\lambda_k$. This function constructs $e^{-i \lambda_k \sigma_k}$ and decomposes it into basic gates. This is not necessarly efficient when the number of qubits is high, so we could improve this constructing variational forms that are efficient for specific each problem.
# - The same algorithm can be used to perform imaginary time evolution. This is useful to find ground states of Hamiltonians.
# - We could test our package with different Hamiltonians and with a higher number of qubits.
# - We could modify the parameters in the Hamiltonians to find phase transitions in the systems.
# ## References
#
#
# [1] [Variational Method](https://en.wikipedia.org/wiki/Variational_method_(quantum_mechanics))
#
# [2] [Stationary-action principle](https://en.wikipedia.org/wiki/Stationary-action_principle)
#
# [3] [Ising Model](https://en.wikipedia.org/wiki/Ising_model)
#
# [4] [Fidelity of quantum states](https://en.wikipedia.org/wiki/Fidelity_of_quantum_states)
#
# [5] <NAME> and <NAME>. Static and dynamic variational principles for expectation values of observables.$\textit{ Annals of Physics}$, 187 (1988). DOI: https://doi.org/10.1016/0003-4916(88)90280-1
#
# [6] <NAME> and <NAME> and <NAME>. and <NAME> <NAME>. Time-Dependent Variational Principle for Quantum Lattices. $\textit{Phys. Rev. Lett.}$, 107 (2011). DOI: https://doi.org/10.1103/PhysRevLett.107.070601
#
# [7] <NAME>., <NAME>., <NAME>., <NAME>. & <NAME>. Theory of variational quantum simulation. $\textit{Quantum 3}$, 191 (2019). DOI: https://doi.org/10.22331/q-2019-10-07-191. URL:https://quantum-journal.org/papers/q-2019-10-07-191/
#
# [8] <NAME> and <NAME>. Efficient variational quantum simulator incorporating active error minimization. $\textit{Phys. Rev. X}$, 7:021050, Jun 2017. DOI: https://doi.org/10.1103/PhysRevX.7.021050. URL: https://link.aps.org/doi/10.1103/PhysRevX.7.021050.
#
#
#
#
#
#
| Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
# +
with open("../data/trivia10k13train.bio", "r") as f:
text_trivia = f.read()
with open("../data/engtrain.bio", "r") as f:
text_eng = f.read()
with open("../data/trivia10k13test.bio", "r") as f:
text_trivia = f.read()
with open("../data/engtest.bio", "r") as f:
text_eng = f.read()
# -
def parse_text(text):
lines = text.split("\n")
new_lines = []
ents = []
for line in lines:
if line:
e, t = line.split("\t")
ents.append(e)
new_lines.append("\t".join([t, e]))
else:
new_lines.append("")
text = "\n".join(new_lines)
return text, list(set(ents))
# +
new_eng, labels_e = parse_text(text_eng)
new_trivia, labels_t = parse_text(text_trivia)
labels_e.sort()
labels_t.sort()
labels = list(set(labels_e+labels_t))
# -
print("Trivia:", labels_t[:12], "\n", "Eng:", labels_e[:11])
# +
# Replace similar labels
replace_dict = {
'Actor': "ACTOR",
'Character_Name': "CHARACTER",
'Director': "DIRECTOR",
'Genre': "GENRE",
'Plot': "PLOT",
'Opinion': "REVIEW",
'Soundtrack': "SONG",
'Year': "YEAR",
'Award': "AWARD",
'Relationship': "RELATIONSHIP",
'Origin': "ORIGIN",
'Quote': "QUOTE"
}
new_labels_t = []
for label in labels_t:
label_ = label.split("-")
new_label = ""
if len(label_) > 1:
if label_[1] in replace_dict:
new_label = f"{label_[0]}-{replace_dict[label_[1]]}"
else:
new_label = label
else:
new_label = label
new_labels_t.append(new_label)
# -
labels = list(set(labels_e+new_labels_t))
print(f"Found {int((len(labels)-1)/2)} labels")
print(f"Example labels: {labels[:5]}")
def replace_text(text):
lines = text.split("\n")
new_lines = []
for line in lines:
if line:
t, label = line.split("\t")
label_ = label.split("-")
e = ""
if len(label_) > 1:
if label_[1] in replace_dict:
e = f"{label_[0]}-{replace_dict[label_[1]]}"
else:
e = label
else:
e = label
new_lines.append("\t".join([t, e]))
else:
new_lines.append("")
text = "\n".join(new_lines)
return text
new_trivia = replace_text(new_trivia)
# +
with open("../data/engtest.txt", "w")as f:
f.write(new_eng)
with open("../data/trivia10k13test.txt", "w")as f:
f.write(new_trivia)
# -
def merge(text1, text2):
text = "-DOCSTART-\tO\n"
text += text1
text += text2
return text
# +
# text = "-DOCSTART-\tO\n\n" + new_eng
# with open("../data/train.txt", "w") as f:
# f.write(text)
# +
# text = merge(new_eng, new_trivia)
# with open("../data/train.txt", "w") as f:
# f.write(text)
# with open("../data/labels.txt", "w") as f:
# for label in labels:
# f.write(label+"\n")
# -
| notebooks/data_preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# ==============================================
# Feature agglomeration vs. univariate selection
# ==============================================
#
# This example compares 2 dimensionality reduction strategies:
#
# - univariate feature selection with Anova
#
# - feature agglomeration with Ward hierarchical clustering
#
# Both methods are compared in a regression problem using
# a BayesianRidge as supervised estimator.
#
#
# +
# Author: <NAME> <<EMAIL>>
# License: BSD 3 clause
print(__doc__)
import shutil
import tempfile
import numpy as np
import matplotlib.pyplot as plt
from scipy import linalg, ndimage
from joblib import Memory
from sklearn.feature_extraction.image import grid_to_graph
from sklearn import feature_selection
from sklearn.cluster import FeatureAgglomeration
from sklearn.linear_model import BayesianRidge
from sklearn.pipeline import Pipeline
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import KFold
# #############################################################################
# Generate data
n_samples = 200
size = 40 # image size
roi_size = 15
snr = 5.
np.random.seed(0)
mask = np.ones([size, size], dtype=np.bool)
coef = np.zeros((size, size))
coef[0:roi_size, 0:roi_size] = -1.
coef[-roi_size:, -roi_size:] = 1.
X = np.random.randn(n_samples, size ** 2)
for x in X: # smooth data
x[:] = ndimage.gaussian_filter(x.reshape(size, size), sigma=1.0).ravel()
X -= X.mean(axis=0)
X /= X.std(axis=0)
y = np.dot(X, coef.ravel())
noise = np.random.randn(y.shape[0])
noise_coef = (linalg.norm(y, 2) / np.exp(snr / 20.)) / linalg.norm(noise, 2)
y += noise_coef * noise # add noise
# #############################################################################
# Compute the coefs of a Bayesian Ridge with GridSearch
cv = KFold(2) # cross-validation generator for model selection
ridge = BayesianRidge()
cachedir = tempfile.mkdtemp()
mem = Memory(location=cachedir, verbose=1)
# Ward agglomeration followed by BayesianRidge
connectivity = grid_to_graph(n_x=size, n_y=size)
ward = FeatureAgglomeration(n_clusters=10, connectivity=connectivity,
memory=mem)
clf = Pipeline([('ward', ward), ('ridge', ridge)])
# Select the optimal number of parcels with grid search
clf = GridSearchCV(clf, {'ward__n_clusters': [10, 20, 30]}, n_jobs=1, cv=cv)
clf.fit(X, y) # set the best parameters
coef_ = clf.best_estimator_.steps[-1][1].coef_
coef_ = clf.best_estimator_.steps[0][1].inverse_transform(coef_)
coef_agglomeration_ = coef_.reshape(size, size)
# Anova univariate feature selection followed by BayesianRidge
f_regression = mem.cache(feature_selection.f_regression) # caching function
anova = feature_selection.SelectPercentile(f_regression)
clf = Pipeline([('anova', anova), ('ridge', ridge)])
# Select the optimal percentage of features with grid search
clf = GridSearchCV(clf, {'anova__percentile': [5, 10, 20]}, cv=cv)
clf.fit(X, y) # set the best parameters
coef_ = clf.best_estimator_.steps[-1][1].coef_
coef_ = clf.best_estimator_.steps[0][1].inverse_transform(coef_.reshape(1, -1))
coef_selection_ = coef_.reshape(size, size)
# #############################################################################
# Inverse the transformation to plot the results on an image
plt.close('all')
plt.figure(figsize=(7.3, 2.7))
plt.subplot(1, 3, 1)
plt.imshow(coef, interpolation="nearest", cmap=plt.cm.RdBu_r)
plt.title("True weights")
plt.subplot(1, 3, 2)
plt.imshow(coef_selection_, interpolation="nearest", cmap=plt.cm.RdBu_r)
plt.title("Feature Selection")
plt.subplot(1, 3, 3)
plt.imshow(coef_agglomeration_, interpolation="nearest", cmap=plt.cm.RdBu_r)
plt.title("Feature Agglomeration")
plt.subplots_adjust(0.04, 0.0, 0.98, 0.94, 0.16, 0.26)
plt.show()
# Attempt to remove the temporary cachedir, but don't worry if it fails
shutil.rmtree(cachedir, ignore_errors=True)
| 01 Machine Learning/scikit_examples_jupyter/cluster/plot_feature_agglomeration_vs_univariate_selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="8yHl0Z8FPPXm" colab_type="text"
# ##First we make our imports.
# + id="z8i_nl7PK85c" colab_type="code" outputId="c60bd03b-daff-4e39-bd66-9be99aae4181" executionInfo={"status": "ok", "timestamp": 1586451087593, "user_tz": 300, "elapsed": 2451, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0LpL-hu169RD5uOvSoMS5Ja4MNM6gnH89MbKU=s64", "userId": "03990109818721859613"}} colab={"base_uri": "https://localhost:8080/", "height": 199}
import pandas as pd
import numpy as np
import nltk
import random
from nltk.corpus import wordnet
from nltk.stem.wordnet import WordNetLemmatizer
from nltk import word_tokenize
from nltk.corpus import stopwords
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics.pairwise import cosine_similarity
nltk.download('punkt')
nltk.download('averaged_perceptron_tagger')
nltk.download('stopwords')
nltk.download('wordnet')
# + [markdown] id="12w55jzCPy-O" colab_type="text"
# ##Read in our data.
# + id="GW0dpWheLShw" colab_type="code" colab={}
df = pd.read_csv('https://raw.githubusercontent.com/danielmoore19/COVID-Chat-Bot/master/covid_data/covid_df.csv')
# + [markdown] id="LWs_7mZVP2_Y" colab_type="text"
# ##Make sure our data looks accurate.
# + id="aY5Y_Q3bLkX3" colab_type="code" outputId="b1d47dc3-eec4-4e04-fdfb-81e84c386fef" executionInfo={"status": "ok", "timestamp": 1586451088533, "user_tz": 300, "elapsed": 3374, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0LpL-hu169RD5uOvSoMS5Ja4MNM6gnH89MbKU=s64", "userId": "03990109818721859613"}} colab={"base_uri": "https://localhost:8080/", "height": 198}
df.head()
# + [markdown] id="8OEmNoMlQF2d" colab_type="text"
# ##Everything looks right as rain.
# + [markdown] id="HRJ_NvSzQKWx" colab_type="text"
# ##Define our tagging function for the lemmatizer.
# + id="t_VWBenqL7w3" colab_type="code" colab={}
def get_wordnet_pos(word):
#Map POS tag to first character lemmatize() accepts
tag = nltk.pos_tag([word])[0][1][0].upper()
tag_dict = {"J": wordnet.ADJ,
"N": wordnet.NOUN,
"V": wordnet.VERB,
"R": wordnet.ADV}
return tag_dict.get(tag, wordnet.NOUN)
# + [markdown] id="G9nWFepVQSkr" colab_type="text"
# ##Define our preprocessing function to tokenize the words in our question bank. After serveral rounds of testing, I decided to use a ranked answering system which functions better if we do not remove stop words. Thus that line has been commented out.
# + id="5JYpI2BKMMyw" colab_type="code" colab={}
def preprocess(docs):
lemmer = WordNetLemmatizer()
prepd = []
for doc in docs:
tokenized = word_tokenize(doc)
cleaned = [lemmer.lemmatize(token.lower(), get_wordnet_pos(token))
for token in tokenized
#if token.lower() not in stopwords.words('english')
if token.isalpha()]
untokenized = ' '.join(cleaned)
prepd.append(untokenized)
return prepd
# + [markdown] id="ocBzKNZHQsEF" colab_type="text"
# ##Process our questions.
# + id="ebLcraVuMWFy" colab_type="code" colab={}
lemm = preprocess(df['questions'])
# + [markdown] id="afEKpW7mQxaL" colab_type="text"
# ##Now we instantiate our model and fit/transform our data and make it a numpy array for our cosine similarity feature.
# + id="eRldrFJ2MlQQ" colab_type="code" colab={}
model = TfidfVectorizer()
tfidf = model.fit_transform(lemm).toarray()
# + [markdown] id="PueAYrJhRBdb" colab_type="text"
# ##The heavy lifting, here we design the engine of the bot. We will tranform the user question to tfidf and set it to an array, then create a new df column called similarity that is populated with the similarity scores of the user questions to our question bank. Next, we rank by descending values and--unless the questions are nearly identical--offer the top three matches.
# + id="yL31SCRjM7x9" colab_type="code" colab={}
def COVID2bot(user_response):
text = model.transform([user_response]).toarray()
df['similarity'] = cosine_similarity(tfidf, text)
scores = df.sort_values(['similarity'], ascending=False)
if scores.iloc[0]['similarity'] >= 0.8:
return (scores.iloc[0]['answers'])
else:
first = scores.iloc[0]['questions']
second = scores.iloc[1]['questions']
third = scores.iloc[2]['questions']
user_input = input(
f'These are the top 3 matches to your question:\n1. "{first}"\n2. "{second}"\n3. "{third}"\nPlease type the number that matches your question, or hit return to ask a different question.\n')
if user_input == '1':
return (scores.iloc[0]['answers'])
elif user_input == '2':
return (scores.iloc[1]['answers'])
elif user_input == '3':
return (scores.iloc[2]['answers'])
else:
return ('Please ask another question.')
# + [markdown] id="D3dajf_qR7RY" colab_type="text"
# ##Create our greeting.
# + id="sBtLw110NXKS" colab_type="code" colab={}
welcome_input = ("hello", "hi", "greetings", "sup", "what's up","hey",)
welcome_response = ["hi", "hey", "*nods*", "hi there", "hello", "I am glad you are talking to me!"]
def welcome(user_response):
for word in user_response.split():
if word.lower() in welcome_input:
return random.choice(welcome_response)
# + [markdown] id="jDaHp5VXR-lz" colab_type="text"
# ##Lastly, we initiate the chatbot. We greet the user and let them know they are speaking to a bot. We let them know what type of bot we are, and how do they end the chat session. Viola!
# + id="I-AO8EXgNajv" colab_type="code" outputId="3c2ac948-e83a-4f09-f86f-8e80ca4c55b0" executionInfo={"status": "ok", "timestamp": 1586451352810, "user_tz": 300, "elapsed": 267602, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj0LpL-hu169RD5uOvSoMS5Ja4MNM6gnH89MbKU=s64", "userId": "03990109818721859613"}} colab={"base_uri": "https://localhost:8080/", "height": 583}
flag=True
print("Greetings! I am a chatbot, and I will try to answer your questions about COVID-19. If you want to exit, type Bye!")
while(flag==True):
user_response = input()
user_response = user_response.lower()
if(user_response not in ['bye','shutdown','exit', 'quit']):
if(user_response=='thanks' or user_response=='thank you'):
flag=False
print("Chatbot : You are welcome..")
else:
if(welcome(user_response)!=None):
print("Chatbot : "+welcome(user_response))
else:
print("Chatbot : ",end="")
print(COVID2bot(user_response))
else:
flag=False
print("Chatbot: Stay safe, and wash your hands!!! ")
| Final COVID Chatbot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="bfa5334d75ac4497808d5ff0a442b91a3fb101e1"
# **Βήμα10:**
# Παρακάτω έχουμε υλοποιήσει τα μοντέλα που ζητήθηκαν για το βημα 10 (3 για κάθε κατηγορία - valence,energy,danceability)
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
import os
print(os.listdir("../input"))
# Any results you write to the current directory are saved as output.
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
import numpy as np
import gzip
import copy
from sklearn.preprocessing import LabelEncoder
from torch.utils.data import Dataset
from torch.utils.data import SubsetRandomSampler, DataLoader
import os
class_mapping = {
'Rock': 'Rock',
'Psych-Rock': 'Rock',
'Indie-Rock': None,
'Post-Rock': 'Rock',
'Psych-Folk': 'Folk',
'Folk': 'Folk',
'Metal': 'Metal',
'Punk': 'Metal',
'Post-Punk': None,
'Trip-Hop': 'Trip-Hop',
'Pop': 'Pop',
'Electronic': 'Electronic',
'Hip-Hop': 'Hip-Hop',
'Classical': 'Classical',
'Blues': 'Blues',
'Chiptune': 'Electronic',
'Jazz': 'Jazz',
'Soundtrack': None,
'International': None,
'Old-Time': None
}
def torch_train_val_split(
dataset, batch_train, batch_eval,
val_size=.2, shuffle=True, seed=42):
# Creating data indices for training and validation splits:
dataset_size = len(dataset)
indices = list(range(dataset_size))
val_split = int(np.floor(val_size * dataset_size))
if shuffle:
np.random.seed(seed)
np.random.shuffle(indices)
train_indices = indices[val_split:]
val_indices = indices[:val_split]
# Creating PT data samplers and loaders:
train_sampler = SubsetRandomSampler(train_indices)
val_sampler = SubsetRandomSampler(val_indices)
train_loader = DataLoader(dataset,
batch_size=batch_train,
sampler=train_sampler)
val_loader = DataLoader(dataset,
batch_size=batch_eval,
sampler=val_sampler)
return train_loader, val_loader
def read_spectrogram(spectrogram_file, chroma=True):
with gzip.GzipFile(spectrogram_file, 'r') as f:
spectrograms = np.load(f)
# spectrograms contains a fused mel spectrogram and chromagram
# Decompose as follows
return spectrograms.T
class LabelTransformer(LabelEncoder):
def inverse(self, y):
try:
return super(LabelTransformer, self).inverse_transform(y)
except:
return super(LabelTransformer, self).inverse_transform([y])
def transform(self, y):
try:
return super(LabelTransformer, self).transform(y)
except:
return super(LabelTransformer, self).transform([y])
class PaddingTransform(object):
def __init__(self, max_length, padding_value=0):
self.max_length = max_length
self.padding_value = padding_value
def __call__(self, s):
if len(s) == self.max_length:
return s
if len(s) > self.max_length:
return s[:self.max_length]
if len(s) < self.max_length:
s1 = copy.deepcopy(s)
pad = np.zeros((self.max_length - s.shape[0], s.shape[1]), dtype=np.float32)
s1 = np.vstack((s1, pad))
return s1
class SpectrogramDataset(Dataset):
def __init__(self, path, class_mapping=None, train=True, max_length=-1):
t = 'train' if train else 'test'
p = os.path.join(path, t)
self.index = os.path.join(path, "{}_labels.txt".format(t))
self.files, labels = self.get_files_labels(self.index, class_mapping)
#print(self.files)
self.feats = [read_spectrogram(os.path.join(p, f+".fused.full.npy.gz")) for f in self.files]
self.feat_dim = self.feats[0].shape[1]
self.lengths = [len(i) for i in self.feats]
self.max_length = max(self.lengths) if max_length <= 0 else max_length
self.zero_pad_and_stack = PaddingTransform(self.max_length)
#self.label_transformer = LabelTransformer()
#if isinstance(labels, (list, tuple)):
#self.labels = np.array(self.label_transformer.fit_transform(labels)).astype('int64')
self.labels=labels
def get_files_labels(self, txt, class_mapping):
with open(txt, 'r') as fd:
lines = [l.rstrip().split('\t') for l in fd.readlines()[1:]]
files, labels = [], []
for l in lines:
l=l[0].split(",")
b=l[1:]
b = list(map(float,b))
files.append(l[0])
labels.append(b)
return files, labels
def __getitem__(self, item):
l = min(self.lengths[item], self.max_length)
return self.zero_pad_and_stack(self.feats[item]), self.labels[item], l
def __len__(self):
return len(self.labels)
# + _uuid="d41fa2bca26c8c21ee72a4cb0705a369903f4784"
BATCH_SZ=32
specs = SpectrogramDataset('../input/data/data/multitask_dataset/', train=True, class_mapping=class_mapping, max_length=-1)
train_loader, val_loader = torch_train_val_split(specs, BATCH_SZ ,BATCH_SZ, val_size=.33)
#test_loader = DataLoader(SpectrogramDataset('../input/data/data/multitask_dataset/', train=False, class_mapping=class_mapping, max_length=-1))
# + [markdown] _uuid="3fee0ca9a98e7102182c6e9d6d1ed0ae6ac2ceb7"
# **REGRESSION FOR VALENCE**
# + [markdown] _uuid="769ed9e2db6e0e7ec78f572dca088647d6265504"
# 1. LSTM for regression
# + _uuid="756b59bcd0266f03ceeb5362c11ee2848ad10bf6"
import numpy as np
import torch
from torch.utils.data import Dataset
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from scipy import stats
class BasicLSTM(nn.Module):
def __init__(self, input_dim,rnn_size, output_dim, num_layers,dropout_type=None,dropout=0, bidirectional=False):
super(BasicLSTM, self).__init__()
self.bidirectional = bidirectional
self.feature_size = rnn_size * 2 if self.bidirectional else rnn_size
self.dropout=dropout
'''
if dropout_type is not None:
if dropout_type = 'Variational_Locked_Dropout'
self.dropout = Variational_LockedDropout(dropout=dropout_prob)
'''
# --------------- Insert your code here ---------------- #
# Initialize the LSTM, Dropout, Output layers
#for non-bidirectional:
#we assume that rnn-size is the number of lstm-units...
#input_dim is the vector that each unit will receive as input..
#hidden_dim at basic lstm is the same as hidden_dim...
#so we have...
self.num_layers=num_layers
self.hidden_dim = rnn_size # OR self.hidden_dim = self.feature_size
if self.bidirectional:
self.lstm = nn.LSTM(input_dim,self.hidden_dim//2,num_layers=num_layers,bidirectional=bidirectional,
dropout=self.dropout,batch_first=True)
else:
self.lstm = nn.LSTM(input_dim,self.hidden_dim,num_layers=num_layers,bidirectional=bidirectional,
dropout=self.dropout,batch_first=True)
#using batch_first=True affects only the input!
# if the input is at form seq_len,batch,features batch_first=True is not needed
#self.hidden = self.init_hidden() #initialize hidden state(and cell state)
self.output_set_size = output_dim
self.hidden2output = nn.Linear(self.hidden_dim,self.output_set_size)
def forward(self, x, lengths):
"""
x : 3D numpy array of dimension N x L x D
N: batch index
L: sequence index
D: feature index
lengths: N x 1
"""
# --------------- Insert your code here ---------------- #
# You must have all of the outputs of the LSTM, but you need only the last one (that does not exceed the sequence length)
# To get it use the last_timestep method
# Then pass it through the remaining network
lstm_out,_ = self.lstm(x)
last_lstm_out = self.last_timestep(lstm_out,lengths)
out_space = self.hidden2output(last_lstm_out)
return out_space
def last_timestep(self, outputs, lengths, bidirectional=False):
"""
Returns the last output of the LSTM taking into account the zero padding
"""
if self.bidirectional:
forward, backward = self.split_directions(outputs)
last_forward = self.last_by_index(forward, lengths)
last_backward = backward[:, 0, :]
# Concatenate and return - maybe add more functionalities like average
return torch.cat((last_forward, last_backward), dim=-1)
else:
return self.last_by_index(outputs, lengths)
@staticmethod
def split_directions(outputs):
direction_size = int(outputs.size(-1) / 2)
forward = outputs[:, :, :direction_size]
backward = outputs[:, :, direction_size:]
return forward, backward
@staticmethod
def last_by_index(outputs, lengths):
# Index of the last output for each sequence.
idx = (lengths - 1).view(-1, 1).expand(outputs.size(0),
outputs.size(2)).unsqueeze(1)
return outputs.gather(1, idx).squeeze()
# + _uuid="51e394790c571013e6738c129e32fc27b491b071"
INPUT_SZ=140
NUM_CLASSES=1
NUM_LAYERS=1
HIDDEN_SIZE=20
DROPOUT=0
num_epochs=30
device=torch.device("cuda")
model_lstm_regr_val = BasicLSTM(INPUT_SZ,HIDDEN_SIZE,NUM_CLASSES,NUM_LAYERS,dropout=DROPOUT,bidirectional=False)
model_lstm_regr_val.to(device)
# + _uuid="a821f1344a70c5c56e73a399973a193c6ef3cce1"
# Loss and optimizer
criterion = nn.MSELoss()
optimizer = torch.optim.SGD(model_lstm_regr_val.parameters(),lr=0.01)
for epoch in range(num_epochs):
#no need to set requires_grad=True for parameters(weights) as it done by default. Also for input requires_grad is not
#always necessary. So we comment the following line.
#with torch.autograd():
model_lstm_regr_val.train()
running_average_loss = 0
#train model in each epoch
for index,instance in enumerate(train_loader):
# Step 1. Remember that Pytorch accumulates gradients.
# We need to clear them out before each instance
#features,labels,lengths=instance
#print(instance)
features = instance[:][0].to(device)
labels = instance[:][1]
valence_labels = labels[0].type(torch.FloatTensor).to(device)
energy_labels = labels[1].type(torch.FloatTensor).to(device)
dance_labels = labels[2].type(torch.FloatTensor).to(device)
lengths = instance[:][2].to(device)
features = features.type(torch.FloatTensor).to(device)
optimizer.zero_grad()
# Step 3. Run our forward pass.
prediction_vec = model_lstm_regr_val(features,lengths)
prediction_vec.to(device)
#print(prediction_vec.shape)
# Step 4. Compute the loss, gradients, and update the parameters by
# calling optimizer.step()
valence_labels=valence_labels.unsqueeze(1)
loss = criterion(prediction_vec,valence_labels)
loss.backward(retain_graph=True)
optimizer.step()
running_average_loss += loss.detach().item()
print("Epoch: {} \t \t Training Loss {}".format(epoch, float(running_average_loss) / (index + 1)))
# + _uuid="31d68b67268115d56fa7b46329b65151cb7e6e38"
model_lstm_regr_val.eval()
n_samples = 0
SE = 0
spearman=[]
running_average_loss=0
with torch.no_grad():
for index, instance in enumerate(val_loader):
features = instance[:][0].to(device)
labels = instance[:][1]
valence_labels = labels[0].type(torch.FloatTensor).to(device)
energy_labels = labels[1].type(torch.FloatTensor).to(device)
dance_labels = labels[2].type(torch.FloatTensor).to(device)
lengths = instance[:][2].to(device)
features = features.type(torch.FloatTensor).to(device)
out = model_lstm_regr_val(features,lengths)
out = out.to(device)
#print(out)
#print(valence_labels)
valence_labels = valence_labels.unsqueeze(1)
E = valence_labels-out
SE = pow(E,2).sum().item() + SE
#print(SE)
n_samples+=features.shape[0]
spearman.append(stats.spearmanr(valence_labels.cpu().squeeze(),out.cpu().squeeze(),axis=0)[0])
print("Spearnman's correlation for LSTM in validation set (predicting valence): " , np.mean(spearman) )
# + [markdown] _uuid="6585b2cb3f767c97ac5a4a4c4b5ea3ece0df99ab"
# 2. CNN Regression
# + _uuid="e56b1a650a4a7260bbe9a022d9f07e8cb567c07a"
import numpy as np
import torch
from torch.utils.data import Dataset
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
class ConvNet(nn.Module):
def __init__(self,input_channels,out_channels,kernel_sz,stride,padding, num_classes):
super(ConvNet, self).__init__()
self.layer1 = nn.Sequential(
nn.Conv2d(input_channels, 4, kernel_size=(3,3), stride=1, padding=1),
nn.ReLU(),
nn.BatchNorm2d(4),
nn.MaxPool2d(kernel_size=2, stride=2)
)
self.layer2 = nn.Sequential(
nn.Conv2d(4, 16, kernel_size=(3,3), stride=1, padding=1),
nn.ReLU(),
nn.BatchNorm2d(16),
nn.MaxPool2d(kernel_size=2, stride=2)
)
self.layer3 = nn.Sequential(
nn.Conv2d(16 , 32 , kernel_size=(3,3), stride=1, padding=1),
nn.ReLU(),
nn.BatchNorm2d(32),
nn.MaxPool2d(kernel_size=3, stride=3)
)
self.layer4 = nn.Sequential(
nn.Conv2d(32, 64, kernel_size=(3,3), stride=1, padding=1),
nn.ReLU(),
nn.BatchNorm2d(64),
nn.MaxPool2d(kernel_size=3, stride=3)
)
self.dense1= nn.Linear(6720,500)
self.dense2 = nn.Linear(500,1)
def forward(self, x,lengths):
#print(x.shape)
x = x.transpose(1, 2)
#print(x.shape)
x.unsqueeze_(1)
#print(x.shape)
out1 = self.layer1(x)
#print(out1.shape)
out2= self.layer2(out1)
#print(out2.shape)
out3= self.layer3(out2)
#print(out3.shape)
out4= self.layer4(out3)
#print(out4.shape)
out_flat=out4.reshape(-1,out4.size(1)*out4.size(2)*out4.size(3))
#print(out_flat.shape)
#implementing fully connected layers
hidden_out = self.dense1(out_flat)
final_out = self.dense2(hidden_out)
return final_out
# + _uuid="d1abfefddd24ca503636e41ff804fb29395ec37c"
num_epochs=35
kernel_sz=3
input_channels=1
out_channels=1
stride=2
padding=2
num_classes=1
device=torch.device("cuda")
model_cnn_regr_val = ConvNet(input_channels,out_channels,kernel_sz,stride,padding ,num_classes)
model_cnn_regr_val.to(device)
# + _uuid="2f03e156bb7f53056db6dd6c71bb7e93298427a5"
# Loss and optimizer
criterion = nn.MSELoss()
optimizer = torch.optim.Adam(model_cnn_regr_val.parameters())
for epoch in range(num_epochs):
#no need to set requires_grad=True for parameters(weights) as it done by default. Also for input requires_grad is not
#always necessary. So we comment the following line.
#with torch.autograd():
model_cnn_regr_val.train()
#scheduler.step()
running_average_loss = 0
#train model in each epoch
for index,instance in enumerate(train_loader):
# Step 1. Remember that Pytorch accumulates gradients.
# We need to clear them out before each instance
#features,labels,lengths=instance
features = instance[:][0].to(device)
labels = instance[:][1]
valence_labels = labels[0].type(torch.FloatTensor).to(device)
energy_labels = labels[1].type(torch.FloatTensor).to(device)
dance_labels = labels[2].type(torch.FloatTensor).to(device)
lengths = instance[:][2].to(device)
features = features.type(torch.FloatTensor).to(device)
optimizer.zero_grad()
# Step 3. Run our forward pass.
prediction_vec = model_cnn_regr_val(features,lengths)
prediction_vec.to(device)
# Step 4. Compute the loss, gradients, and update the parameters by
# calling optimizer.step()
valence_labels = valence_labels.unsqueeze(1)
loss = criterion(prediction_vec,valence_labels)
loss.backward(retain_graph=True)
optimizer.step()
running_average_loss += loss.detach().item()
print("Epoch: {} \t \t Training Loss {}".format(epoch, float(running_average_loss) / (index + 1)))
# + _uuid="32339d286fbcedc7edd6dba96a18443ddaf8980f"
model_cnn_regr_val.eval()
n_samples = 0
SE = 0
spearman=[]
with torch.no_grad():
for index, instance in enumerate(val_loader):
features = instance[:][0].to(device)
labels = instance[:][1]
valence_labels = labels[0].type(torch.FloatTensor).to(device)
energy_labels = labels[1].type(torch.FloatTensor).to(device)
dance_labels = labels[2].type(torch.FloatTensor).to(device)
lengths = instance[:][2].to(device)
features = features.type(torch.FloatTensor).to(device)
out = model_cnn_regr_val(features,lengths)
out = out.to(device)
valence_labels = valence_labels.unsqueeze(1)
E = valence_labels-out
SE = pow(E,2).sum().item() + SE
#print(SE)
n_samples+=features.shape[0]
spearman.append(stats.spearmanr(valence_labels.cpu().squeeze(),out.cpu().squeeze(),axis=0)[0])
print("Spearnman's correlation for CNN-2D in validation set (predicting valence): " , np.mean(spearman) )
# + [markdown] _uuid="1ff8fcc069b15a99b4aa410be93ac30c494df4d3"
# 3. CNN-LSTM regression
# + _uuid="5b5d632cf33cb8b7085e4b006b11f923b658f317"
import numpy as np
import torch
from torch.utils.data import Dataset
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
class ConvLSTM(nn.Module):
def __init__(self,input_channels,out_channels,kernel_sz,stride,padding, num_classes,input_dim,rnn_size, output_dim, num_layers,dropout_type=None,dropout=0, bidirectional=False):
super(ConvLSTM, self).__init__()
self.layer1 = nn.Sequential(
nn.Conv1d(input_channels, 256, kernel_size=kernel_sz, stride=stride, padding=padding),
nn.ReLU(),
nn.BatchNorm1d(256),
nn.MaxPool1d(kernel_size=2, stride=2)
)
self.layer2 = nn.Sequential(
nn.Conv1d(256, 512, kernel_size=kernel_sz, stride=stride, padding=padding),
nn.ReLU(),
nn.BatchNorm1d(512),
nn.MaxPool1d(kernel_size=2, stride=2)
)
self.bidirectional = bidirectional
self.feature_size = rnn_size * 2 if self.bidirectional else rnn_size
self.dropout=dropout
'''
if dropout_type is not None:
if dropout_type = 'Variational_Locked_Dropout'
self.dropout = Variational_LockedDropout(dropout=dropout_prob)
'''
# --------------- Insert your code here ---------------- #
# Initialize the LSTM, Dropout, Output layers
#for non-bidirectional:
#we assume that rnn-size is the number of lstm-units...
#input_dim is the vector that each unit will receive as input..
#hidden_dim at basic lstm is the same as hidden_dim...
#so we have...
self.num_layers=num_layers
self.hidden_dim = rnn_size # OR self.hidden_dim = self.feature_size
if self.bidirectional:
self.lstm = nn.LSTM(input_dim,self.hidden_dim//2,num_layers=num_layers,bidirectional=bidirectional,
dropout=self.dropout,batch_first=True)
else:
self.lstm = nn.LSTM(input_dim,self.hidden_dim,num_layers=num_layers,bidirectional=bidirectional,
dropout=self.dropout,batch_first=True)
#using batch_first=True affects only the input!
# if the input is at form seq_len,batch,features batch_first=True is not needed
#self.hidden = self.init_hidden() #initialize hidden state(and cell state)
#use a linear transformation from lstm hidden_state space to ouput space..
#for digit classification we want to classify a sequence to 0-9 digits. So we will use as output dim the number 10.
self.output_set_size = output_dim
self.hidden2output = nn.Linear(self.hidden_dim,self.output_set_size)
def forward(self, x,lengths):
#print(x.shape)
x = x.transpose(1, 2)
#print(x.shape)
out = self.layer1(x)
out = self.layer2(out)
out = out.transpose(1, 2)
#print(out.shape)
#newlengths = torch.ones(out.shape[0])
#newlengths=newlengths * (out.shape[1])
newlengths = lengths//4-1
#print(newlengths)
newlengths = newlengths.type(torch.LongTensor).to(device)
#print(out.shape)
lstm_out,_ = self.lstm(out)
#print(lstm_out.shape)
last_lstm_out = self.last_timestep(lstm_out,newlengths)
out_space = self.hidden2output(last_lstm_out)
#print(out_space.shape)
#out_space = self.hidden2output(lstm_out)
return out_space
def last_timestep(self, outputs, lengths):
"""
Returns the last output of the LSTM taking into account the zero padding
"""
if self.bidirectional:
forward, backward = self.split_directions(outputs)
last_forward = self.last_by_index(forward, lengths)
last_backward = backward[:, 0, :]
# Concatenate and return - maybe add more functionalities like average
return torch.cat((last_forward, last_backward), dim=-1)
else:
return self.last_by_index(outputs, lengths)
@staticmethod
def split_directions(outputs):
direction_size = int(outputs.size(-1) / 2)
forward = outputs[:, :, :direction_size]
backward = outputs[:, :, direction_size:]
return forward, backward
@staticmethod
def last_by_index(outputs, lengths):
# Index of the last output for each sequence.
idx = (lengths - 1).view(-1, 1).expand(outputs.size(0),
outputs.size(2)).unsqueeze(1)
return outputs.gather(1, idx).squeeze()
# + _uuid="9dd0175971fc4fcbf67c400907c7d7a41666c152"
num_epochs=35
kernel_sz=3
input_channels=140
out_channels=140
stride=1
padding=1
num_classes=1
INPUT_SZ=512
NUM_CLASSES=1
NUM_LAYERS=1
HIDDEN_SIZE=20
DROPOUT=0
device=torch.device("cuda")
model_cnn_lstm_regr_val = ConvLSTM(input_channels,out_channels,kernel_sz,stride,padding ,num_classes,INPUT_SZ,HIDDEN_SIZE,NUM_CLASSES,NUM_LAYERS,dropout=DROPOUT,bidirectional=False)
model_cnn_lstm_regr_val.to(device)
# + _uuid="de76ccbc3e9220f673e09795a4876c3d0bdc3485"
# Loss and optimizer
criterion = nn.MSELoss()
optimizer = torch.optim.Adam(model_cnn_lstm_regr_val.parameters())
for epoch in range(num_epochs):
#no need to set requires_grad=True for parameters(weights) as it done by default. Also for input requires_grad is not
#always necessary. So we comment the following line.
#with torch.autograd():
model_cnn_lstm_regr_val.train()
#scheduler.step()
running_average_loss = 0
#train model in each epoch
for index,instance in enumerate(train_loader):
# Step 1. Remember that Pytorch accumulates gradients.
# We need to clear them out before each instance
#features,labels,lengths=instance
features = instance[:][0]
labels = instance[:][1]
valence_labels = labels[0].type(torch.FloatTensor).to(device)
energy_labels = labels[1].type(torch.FloatTensor).to(device)
dance_labels = labels[2].type(torch.FloatTensor).to(device)
lengths = instance[:][2].to(device)
features = features.type(torch.FloatTensor).to(device)
optimizer.zero_grad()
# Step 3. Run our forward pass.
prediction_vec = model_cnn_lstm_regr_val(features,lengths)
prediction_vec.to(device)
# Step 4. Compute the loss, gradients, and update the parameters by
# calling optimizer.step()
valence_labels = valence_labels.unsqueeze(1)
loss = criterion(prediction_vec,valence_labels)
loss.backward(retain_graph=True)
optimizer.step()
running_average_loss += loss.detach().item()
print("Epoch: {} \t \t Training Loss {}".format(epoch, float(running_average_loss) / (index + 1)))
# + _uuid="775a582575c8f3945d17d9a93c320b3a230aaa5e"
model_cnn_lstm_regr_val.eval()
n_samples = 0
SE = 0
spearman=[]
with torch.no_grad():
for index, instance in enumerate(val_loader):
features = instance[:][0].to(device)
labels = instance[:][1]
valence_labels = labels[0].type(torch.FloatTensor).to(device)
energy_labels = labels[1].type(torch.FloatTensor).to(device)
dance_labels = labels[2].type(torch.FloatTensor).to(device)
lengths = instance[:][2].to(device)
features = features.type(torch.FloatTensor).to(device)
out = model_cnn_lstm_regr_val(features,lengths)
out = out.to(device)
valence_labels = valence_labels.unsqueeze(1)
E = valence_labels-out
SE = pow(E,2).sum().item() + SE
#print(SE)
n_samples+=features.shape[0]
spearman.append(stats.spearmanr(valence_labels.cpu().squeeze(),out.cpu().squeeze(),axis=0)[0])
print("Spearnman's correlation for CNN-LSTM in validation set (predicting valence): " , np.mean(spearman) )
# + [markdown] _uuid="b195f65a93b0a42733bc6dce6d73dea93266cbcb"
# **REGRESSION FOR ENERGY**
# + [markdown] _uuid="cd184993a3c150c84936594a846513fd2906ec99"
# 1. LSTM Regression
# + _uuid="69b37dd279e41641e4b0eb0facee0f7fa65386f8"
INPUT_SZ=140
NUM_CLASSES=1
NUM_LAYERS=1
HIDDEN_SIZE=40
DROPOUT=0
num_epochs=30
device=torch.device("cuda")
model_lstm_regr_energy = BasicLSTM(INPUT_SZ,HIDDEN_SIZE,NUM_CLASSES,NUM_LAYERS,dropout=DROPOUT,bidirectional=False)
model_lstm_regr_energy.to(device)
# + _uuid="eb7c1dfd9cef8501831a78d439c6fb3d8def2ecb"
# Loss and optimizer
criterion = nn.MSELoss()
optimizer = torch.optim.SGD(model_lstm_regr_energy.parameters(),lr=0.001)
for epoch in range(num_epochs):
#no need to set requires_grad=True for parameters(weights) as it done by default. Also for input requires_grad is not
#always necessary. So we comment the following line.
#with torch.autograd():
model_lstm_regr_energy.train()
running_average_loss = 0
#train model in each epoch
for index,instance in enumerate(train_loader):
# Step 1. Remember that Pytorch accumulates gradients.
# We need to clear them out before each instance
#features,labels,lengths=instance
#print(instance)
features = instance[:][0].to(device)
labels = instance[:][1]
valence_labels = labels[0].type(torch.FloatTensor).to(device)
energy_labels = labels[1].type(torch.FloatTensor).to(device)
dance_labels = labels[2].type(torch.FloatTensor).to(device)
lengths = instance[:][2].to(device)
features = features.type(torch.FloatTensor).to(device)
optimizer.zero_grad()
# Step 3. Run our forward pass.
prediction_vec = model_lstm_regr_energy(features,lengths)
prediction_vec.to(device)
#print(prediction_vec.shape)
# Step 4. Compute the loss, gradients, and update the parameters by
# calling optimizer.step()
energy_labels=energy_labels.unsqueeze(1)
loss = criterion(prediction_vec,energy_labels)
loss.backward(retain_graph=True)
optimizer.step()
running_average_loss += loss.detach().item()
print("Epoch: {} \t \t Training Loss {}".format(epoch, float(running_average_loss) / (index + 1)))
# + _uuid="bae487fef24f4c4c7687190ee890b9937ae49d78"
model_lstm_regr_energy.eval()
n_samples = 0
SE = 0
spearman=[]
running_average_loss=0
with torch.no_grad():
for index, instance in enumerate(val_loader):
features = instance[:][0].to(device)
labels = instance[:][1]
valence_labels = labels[0].type(torch.FloatTensor).to(device)
energy_labels = labels[1].type(torch.FloatTensor).to(device)
dance_labels = labels[2].type(torch.FloatTensor).to(device)
lengths = instance[:][2].to(device)
features = features.type(torch.FloatTensor).to(device)
out = model_lstm_regr_energy(features,lengths)
out = out.to(device)
#print(out)
#print(valence_labels)
energy_labels = energy_labels.unsqueeze(1)
E = energy_labels-out
SE = pow(E,2).sum().item() + SE
n_samples+=features.shape[0]
spearman.append(stats.spearmanr(energy_labels.cpu().squeeze(),out.cpu().squeeze(),axis=0)[0])
print("Spearnman's correlation for CNN-LSTM in validation set (predicting energy): " , np.mean(spearman) )
# + [markdown] _uuid="b4591b2eb1707e0d3c70e3664c41e11a82f0d285"
# 2.CNN regression
# + _uuid="89a1dfb8ce79cb0cbd373e10c0ad9966b148328d"
num_epochs=35
kernel_sz=3
input_channels=1
out_channels=1
stride=2
padding=2
num_classes=10
device=torch.device("cuda")
model_cnn_regr_energy = ConvNet(input_channels,out_channels,kernel_sz,stride,padding ,num_classes)
model_cnn_regr_energy.to(device)
# + _uuid="c88757826fd269cc3c56f5ffd7c98277e667dad7"
# Loss and optimizer
criterion = nn.MSELoss()
optimizer = torch.optim.Adam(model_cnn_regr_energy.parameters())
for epoch in range(num_epochs):
#no need to set requires_grad=True for parameters(weights) as it done by default. Also for input requires_grad is not
#always necessary. So we comment the following line.
#with torch.autograd():
model_cnn_regr_energy.train()
#scheduler.step()
running_average_loss = 0
#train model in each epoch
for index,instance in enumerate(train_loader):
# Step 1. Remember that Pytorch accumulates gradients.
# We need to clear them out before each instance
#features,labels,lengths=instance
features = instance[:][0].to(device)
labels = instance[:][1]
valence_labels = labels[0].type(torch.FloatTensor).to(device)
energy_labels = labels[1].type(torch.FloatTensor).to(device)
dance_labels = labels[2].type(torch.FloatTensor).to(device)
lengths = instance[:][2].to(device)
features = features.type(torch.FloatTensor).to(device)
optimizer.zero_grad()
# Step 3. Run our forward pass.
prediction_vec = model_cnn_regr_energy(features,lengths)
prediction_vec.to(device)
# Step 4. Compute the loss, gradients, and update the parameters by
# calling optimizer.step()
energy_labels = energy_labels.unsqueeze(1)
loss = criterion(prediction_vec,energy_labels)
loss.backward(retain_graph=True)
optimizer.step()
running_average_loss += loss.detach().item()
print("Epoch: {} \t \t Training Loss {}".format(epoch, float(running_average_loss) / (index + 1)))
# + _uuid="4a1b9683e4929d5f96ddfb362b88a8ef827ecb8c"
model_cnn_regr_energy.eval()
n_samples = 0
SE = 0
spearman=[]
running_average_loss=0
with torch.no_grad():
for index, instance in enumerate(val_loader):
features = instance[:][0].to(device)
labels = instance[:][1]
valence_labels = labels[0].type(torch.FloatTensor).to(device)
energy_labels = labels[1].type(torch.FloatTensor).to(device)
dance_labels = labels[2].type(torch.FloatTensor).to(device)
lengths = instance[:][2].to(device)
features = features.type(torch.FloatTensor).to(device)
out = model_cnn_regr_energy(features,lengths)
out = out.to(device)
#print(out)
#print(valence_labels)
energy_labels = energy_labels.unsqueeze(1)
E = energy_labels-out
SE = pow(E,2).sum().item() + SE
n_samples+=features.shape[0]
spearman.append(stats.spearmanr(energy_labels.cpu().squeeze(),out.cpu().squeeze(),axis=0)[0])
print("Spearnman's correlation for CNN-2d in validation set (predicting energy): " , np.mean(spearman) )
# + [markdown] _uuid="036d270b657ba5adcce175097e046a9949c39479"
# 3.CNN-LSTM Regression
# + _uuid="925d30454814a2cc0ef46a2a36f68d63fcf44243"
num_epochs=35
kernel_sz=3
input_channels=140
out_channels=140
stride=1
padding=1
num_classes=1
INPUT_SZ=512
NUM_CLASSES=1
NUM_LAYERS=1
HIDDEN_SIZE=20
DROPOUT=0
device=torch.device("cuda")
model_cnn_lstm_regr_energy = ConvLSTM(input_channels,out_channels,kernel_sz,stride,padding ,num_classes,INPUT_SZ,HIDDEN_SIZE,NUM_CLASSES,NUM_LAYERS,dropout=DROPOUT,bidirectional=False)
model_cnn_lstm_regr_energy.to(device)
# + _uuid="e7565e64f10bd2e3e014f4ff135bf5fc8ac36ef7"
# Loss and optimizer
criterion = nn.MSELoss()
optimizer = torch.optim.Adam(model_cnn_regr_energy.parameters())
for epoch in range(num_epochs):
#no need to set requires_grad=True for parameters(weights) as it done by default. Also for input requires_grad is not
#always necessary. So we comment the following line.
#with torch.autograd():
model_cnn_lstm_regr_energy.train()
#scheduler.step()
running_average_loss = 0
#train model in each epoch
for index,instance in enumerate(train_loader):
# Step 1. Remember that Pytorch accumulates gradients.
# We need to clear them out before each instance
#features,labels,lengths=instance
features = instance[:][0].to(device)
labels = instance[:][1]
valence_labels = labels[0].type(torch.FloatTensor).to(device)
energy_labels = labels[1].type(torch.FloatTensor).to(device)
dance_labels = labels[2].type(torch.FloatTensor).to(device)
lengths = instance[:][2].to(device)
features = features.type(torch.FloatTensor).to(device)
optimizer.zero_grad()
# Step 3. Run our forward pass.
prediction_vec = model_cnn_lstm_regr_energy(features,lengths)
prediction_vec.to(device)
# Step 4. Compute the loss, gradients, and update the parameters by
# calling optimizer.step()
energy_labels = energy_labels.unsqueeze(1)
loss = criterion(prediction_vec,energy_labels)
loss.backward(retain_graph=True)
optimizer.step()
running_average_loss += loss.detach().item()
print("Epoch: {} \t \t Training Loss {}".format(epoch, float(running_average_loss) / (index + 1)))
# + _uuid="24cba6c3dc2673c95ead3b3f3dcaf07e538df2f4"
model_cnn_lstm_regr_energy.eval()
n_samples = 0
SE = 0
spearman=[]
running_average_loss=0
with torch.no_grad():
for index, instance in enumerate(val_loader):
features = instance[:][0].to(device)
labels = instance[:][1]
valence_labels = labels[0].type(torch.FloatTensor).to(device)
energy_labels = labels[1].type(torch.FloatTensor).to(device)
dance_labels = labels[2].type(torch.FloatTensor).to(device)
lengths = instance[:][2].to(device)
features = features.type(torch.FloatTensor).to(device)
out = model_cnn_lstm_regr_energy(features,lengths)
out = out.to(device)
#print(out)
#print(valence_labels)
energy_labels = energy_labels.unsqueeze(1)
E = energy_labels-out
SE = pow(E,2).sum().item() + SE
n_samples+=features.shape[0]
spearman.append(stats.spearmanr(energy_labels.cpu().squeeze(),out.cpu().squeeze(),axis=0)[0])
print("Spearnman's correlation for CNN-LSTM in validation set (predicting energy): " , np.mean(spearman) )
# + [markdown] _uuid="d9c2b156a582f44fe8dcb261bc2ef8c2ba012a63"
# **REGRESSION FOR DANCEABILITY**
# + [markdown] _uuid="fd9a5018aca7e4eed2d11f8c4561a316e0592ad4"
# 1.LSTM Regression
# + _uuid="c3f1e26af853104c477e98a55e813389ee151370"
INPUT_SZ=140
NUM_CLASSES=1
NUM_LAYERS=1
HIDDEN_SIZE=40
DROPOUT=0
num_epochs=30
device=torch.device("cuda")
model_lstm_regr_dance = BasicLSTM(INPUT_SZ,HIDDEN_SIZE,NUM_CLASSES,NUM_LAYERS,dropout=DROPOUT,bidirectional=False)
model_lstm_regr_dance.to(device)
# + _uuid="a4bf4f5e7d37c2d962cb01b2f84e0f231631bb1e"
# Loss and optimizer
criterion = nn.MSELoss()
optimizer = torch.optim.SGD(model_lstm_regr_dance.parameters(),lr=0.001)
for epoch in range(num_epochs):
#no need to set requires_grad=True for parameters(weights) as it done by default. Also for input requires_grad is not
#always necessary. So we comment the following line.
#with torch.autograd():
model_lstm_regr_dance.train()
running_average_loss = 0
#train model in each epoch
for index,instance in enumerate(train_loader):
# Step 1. Remember that Pytorch accumulates gradients.
# We need to clear them out before each instance
#features,labels,lengths=instance
#print(instance)
features = instance[:][0].to(device)
labels = instance[:][1]
valence_labels = labels[0].type(torch.FloatTensor).to(device)
energy_labels = labels[1].type(torch.FloatTensor).to(device)
dance_labels = labels[2].type(torch.FloatTensor).to(device)
lengths = instance[:][2].to(device)
features = features.type(torch.FloatTensor).to(device)
optimizer.zero_grad()
# Step 3. Run our forward pass.
prediction_vec = model_lstm_regr_dance(features,lengths)
prediction_vec.to(device)
#print(prediction_vec.shape)
# Step 4. Compute the loss, gradients, and update the parameters by
# calling optimizer.step()
dance_labels=dance_labels.unsqueeze(1)
loss = criterion(prediction_vec,dance_labels)
loss.backward(retain_graph=True)
optimizer.step()
running_average_loss += loss.detach().item()
print("Epoch: {} \t \t Training Loss {}".format(epoch, float(running_average_loss) / (index + 1)))
# + _uuid="beb98bc04079736d7a778e827e1a6bb72c076263"
model_lstm_regr_dance.eval()
n_samples = 0
SE = 0
spearman=[]
running_average_loss=0
with torch.no_grad():
for index, instance in enumerate(val_loader):
features = instance[:][0].to(device)
labels = instance[:][1]
valence_labels = labels[0].type(torch.FloatTensor).to(device)
energy_labels = labels[1].type(torch.FloatTensor).to(device)
dance_labels = labels[2].type(torch.FloatTensor).to(device)
lengths = instance[:][2].to(device)
features = features.type(torch.FloatTensor).to(device)
out = model_lstm_regr_dance(features,lengths)
out = out.to(device)
#print(out)
#print(valence_labels)
dance_labels = dance_labels.unsqueeze(1)
E = dance_labels-out
SE = pow(E,2).sum().item() + SE
#print(SE)
n_samples+=features.shape[0]
spearman.append(stats.spearmanr(dance_labels.cpu().squeeze(),out.cpu().squeeze(),axis=0)[0])
print("Spearnman's correlation for LSTM in validation set (predicting dance): " , np.mean(spearman) )
# + [markdown] _uuid="20a8070ec5529dbfdadb5a64f8269676cb5f3fc6"
# CNN regression
# + _uuid="84e1bd0d57b6dc2c0f938d84956dd1bf04889f1a"
num_epochs=35
kernel_sz=3
input_channels=1
out_channels=1
stride=2
padding=2
num_classes=10
device=torch.device("cuda")
model_cnn_regr_dance = ConvNet(input_channels,out_channels,kernel_sz,stride,padding ,num_classes)
model_cnn_regr_dance.to(device)
# + _uuid="bb134477e4ef2ba7e4ccfb6a011cacf2a2436dd5"
# Loss and optimizer
criterion = nn.MSELoss()
optimizer = torch.optim.Adam(model_cnn_regr_dance.parameters())
for epoch in range(num_epochs):
#no need to set requires_grad=True for parameters(weights) as it done by default. Also for input requires_grad is not
#always necessary. So we comment the following line.
#with torch.autograd():
model_cnn_regr_dance.train()
#scheduler.step()
running_average_loss = 0
#train model in each epoch
for index,instance in enumerate(train_loader):
# Step 1. Remember that Pytorch accumulates gradients.
# We need to clear them out before each instance
#features,labels,lengths=instance
features = instance[:][0].to(device)
labels = instance[:][1]
valence_labels = labels[0].type(torch.FloatTensor).to(device)
energy_labels = labels[1].type(torch.FloatTensor).to(device)
dance_labels = labels[2].type(torch.FloatTensor).to(device)
lengths = instance[:][2].to(device)
features = features.type(torch.FloatTensor).to(device)
optimizer.zero_grad()
# Step 3. Run our forward pass.
prediction_vec = model_cnn_regr_dance(features,lengths)
prediction_vec.to(device)
# Step 4. Compute the loss, gradients, and update the parameters by
# calling optimizer.step()
dance_labels = dance_labels.unsqueeze(1)
loss = criterion(prediction_vec,dance_labels)
loss.backward(retain_graph=True)
optimizer.step()
running_average_loss += loss.detach().item()
print("Epoch: {} \t \t Training Loss {}".format(epoch, float(running_average_loss) / (index + 1)))
# + _uuid="d8c6508bfa5bbef94945f88f63dfb07334b85a53"
model_cnn_regr_dance.eval()
n_samples = 0
SE = 0
spearman=[]
running_average_loss=0
with torch.no_grad():
for index, instance in enumerate(val_loader):
features = instance[:][0].to(device)
labels = instance[:][1]
valence_labels = labels[0].type(torch.FloatTensor).to(device)
energy_labels = labels[1].type(torch.FloatTensor).to(device)
dance_labels = labels[2].type(torch.FloatTensor).to(device)
lengths = instance[:][2].to(device)
features = features.type(torch.FloatTensor).to(device)
out = model_cnn_regr_dance(features,lengths)
out = out.to(device)
#print(out)
#print(valence_labels)
dance_labels = dance_labels.unsqueeze(1)
E = dance_labels-out
SE = pow(E,2).sum().item() + SE
n_samples+=features.shape[0]
spearman.append(stats.spearmanr(dance_labels.cpu().squeeze(),out.cpu().squeeze(),axis=0)[0])
print("Spearnman's correlation for CNN in validation set (predicting dance): " , np.mean(spearman) )
# + [markdown] _uuid="2af8f66ca3a46a522e8476152e3f164590e9dd45"
# 3. CNN-LSTM regression
#
# + _uuid="2cb02f8463d3fcd2adc4f655934d42d042468078"
num_epochs=35
kernel_sz=3
input_channels=140
out_channels=140
stride=1
padding=1
num_classes=1
INPUT_SZ=512
NUM_CLASSES=1
NUM_LAYERS=1
HIDDEN_SIZE=20
DROPOUT=0
device=torch.device("cuda")
model_cnn_lstm_regr_dance = ConvLSTM(input_channels,out_channels,kernel_sz,stride,padding ,num_classes,INPUT_SZ,HIDDEN_SIZE,NUM_CLASSES,NUM_LAYERS,dropout=DROPOUT,bidirectional=False)
model_cnn_lstm_regr_dance.to(device)
# + _uuid="6e4b8f454ef2b14ea03ff02a8cb4ce23bee177cb"
# Loss and optimizer
criterion = nn.MSELoss()
optimizer = torch.optim.Adam(model_cnn_regr_dance.parameters())
for epoch in range(num_epochs):
#no need to set requires_grad=True for parameters(weights) as it done by default. Also for input requires_grad is not
#always necessary. So we comment the following line.
#with torch.autograd():
model_cnn_lstm_regr_dance.train()
#scheduler.step()
running_average_loss = 0
#train model in each epoch
for index,instance in enumerate(train_loader):
# Step 1. Remember that Pytorch accumulates gradients.
# We need to clear them out before each instance
#features,labels,lengths=instance
features = instance[:][0].to(device)
labels = instance[:][1]
valence_labels = labels[0].type(torch.FloatTensor).to(device)
energy_labels = labels[1].type(torch.FloatTensor).to(device)
dance_labels = labels[2].type(torch.FloatTensor).to(device)
lengths = instance[:][2].to(device)
features = features.type(torch.FloatTensor).to(device)
optimizer.zero_grad()
# Step 3. Run our forward pass.
prediction_vec = model_cnn_lstm_regr_dance(features,lengths)
prediction_vec.to(device)
# Step 4. Compute the loss, gradients, and update the parameters by
# calling optimizer.step()
dance_labels = dance_labels.unsqueeze(1)
loss = criterion(prediction_vec,dance_labels)
loss.backward(retain_graph=True)
optimizer.step()
running_average_loss += loss.detach().item()
print("Epoch: {} \t \t Training Loss {}".format(epoch, float(running_average_loss) / (index + 1)))
# + _uuid="3728e7daf85aadc751bf8aa96d0ddc958769d864"
model_cnn_lstm_regr_dance.eval()
n_samples = 0
SE = 0
spearman = []
running_average_loss=0
with torch.no_grad():
for index, instance in enumerate(val_loader):
features = instance[:][0].to(device)
labels = instance[:][1]
valence_labels = labels[0].type(torch.FloatTensor).to(device)
energy_labels = labels[1].type(torch.FloatTensor).to(device)
dance_labels = labels[2].type(torch.FloatTensor).to(device)
lengths = instance[:][2].to(device)
features = features.type(torch.FloatTensor).to(device)
out = model_cnn_lstm_regr_dance(features,lengths)
out = out.to(device)
#print(out)
#print(valence_labels)
dance_labels = dance_labels.unsqueeze(1)
#E = dance_labels-out
#SE = pow(E,2).sum().item() + SE
#n_samples+=features.shape[0]
spearman.append(stats.spearmanr(dance_labels.cpu().squeeze(),out.cpu().squeeze(),axis=0)[0])
print("Spearnman's correlation for CNN-LSTM in validation set (predicting danceability): " , np.mean(spearman) )
# + _uuid="1ce90be411ad80f2efa8f883de5edecb1828515c"
# + _uuid="ce31f0a2510dc56536a9b954924a1270c4aa9b02"
# + _uuid="d122d9d6d915fd5126dab2f346af7f6b62309b95"
| lab3/step_10_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
def strStr(haystack, needle):
if needle in haystack:
return haystack.index(needle)
else:
return -1
print(strStr("hello", "llso"))
# -
| Anjani/Leetcode/Implement strStr().ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="OLN8UWNM4gO5"
# # Conditional Logic
#
# In the last lesson, we learned about logic with booleans. Booleans can be used to determine if a certain line of code should be run. How would we do this? This is where we can use something called an if statement.
#
# If statements follow the following formula:
#
# ```
# if [condition]:
# [code]
# ```
#
# The __condition__ is a boolean, and the code under the condition only runs if the condition is true. These are called **if statements**
#
# For example, say we had a list that we wanted to print out, but we don't want to print the list if it's too long. We could run the following:
# + colab={} colab_type="code" id="w0vYiQVk4gO-" outputId="6da5cb4d-5732-4b83-93c3-312e55faf3c7"
# create a list called my_list containing the numbers 0 to 7
# write an if statement that only prints my_list if it has less than 10 items
# -
# Let's test the conditional statement on its own to see what the result is:
# + colab={} colab_type="code" id="q3hSCO_i4gPI" outputId="adbf2d2c-9330-4783-87aa-ff9e9f64e9d3"
# print the boolean value of the condition in the if statement above
# + [markdown] colab_type="text" id="eLWm7_7f4gPP"
# We see that the condition is `True`, the code below runs.
#
# What if we changed the condition above? How would this change our code?
# + colab={} colab_type="code" id="Yx_5-0Fq4gPR"
# write an if statement that only prints my_list if it has 10 or more items
# + [markdown] colab_type="text" id="4zGhearv4gPV"
# Nothing happened! Let's see why this is.
# + colab={} colab_type="code" id="aa6JU-xC4gPW" outputId="7375ac9c-6d78-481e-e926-697d86ddef9a"
# print the boolean value of the above if statement
# + [markdown] colab_type="text" id="KHYd0tUm4gPc"
# In this case, the condition is `False`, so the code within the conditional did not run.
# + [markdown] colab_type="text" id="LuaRXfzy4gPd"
# What if we wanted to run one block of code if the condition is `True`, and a different line of code if the condition is `False`? In other words, what if we wanted to run one block of code if our list has less than 10 items, and another block of code if the list has more than 10? This is where we use `if/else`.
#
# An `if/else` looks like this:
#
# ```
# if [condition]:
# [command 1]
# else:
# [command 2]
# ```
#
# The first two lines should look familiar; it's an if statement, which we just used! But after the if statement, we write `else:` and then some more python code. The code that comes after `else:` will run if the value of `condition` is false.
#
# Let's see how this works! Let's write code that prints my_list if it has less than 10 items, and otherwise prints a message letting us know the list is too long to print.
# + colab={} colab_type="code" id="JTbTyDzR4gPe" outputId="2f0ee061-cd24-46b0-fb39-2ef577086d57"
# write an if/else statement that prints my_list if my_list has less than 10 items
# and if my_list has 10 or more items, prints a message
# + [markdown] colab_type="text" id="Nf15XEXN4gPj"
# Because the list has less than 10 items, the list printed. But let's say we added more items to the list, making it longer than 10 items.
# + colab={} colab_type="code" id="V6zs4yeO4gPl"
# append the numbers 8, 9, 10, and 11 to my_list
# + colab={} colab_type="code" id="f2kJl8cf4gPp" outputId="b1b2a8e5-4907-44a8-c98e-7b55590d9a86"
# print the length of my_list
# + [markdown] colab_type="text" id="MVTxM5e54gPs"
# Now, the length of my_list is above 10. How would the above line of code run now?
# + colab={} colab_type="code" id="4PJ8LJSG4gPu" outputId="ecd36b06-1017-4a67-b7c6-659f08c3ed30"
# # copy and paste the if/else statement we just wrote
# + [markdown] colab_type="text" id="TpJm2bZ44gPy"
# What if there were more than two different conditions? Using `if/elif/else` statements, we can command Python to run different code for any number of conditions. An `if/elif/else` statement looks like this:
#
# ```
# if [condition 1]:
# [command 1]
# elif [condition 2]:
# [command 2]
# else:
# [command 3]
#
# [more python code]
# ```
#
# How does Python run this code? First it checks to see if `condition 1` is true. If it is, then it runs `command 1`, skips `command 2` and `command 3`, then continues running whatever code that comes after the `if/elif/else`.
#
# If `condition 1` is false, but `condition 2` is true, then Python runs `command 2`, skips `command 3`, and continues with the code that comes after the `if/elif/else`.
#
# Finally, if `condition 1` AND `condition 2` are both false, then `command 3` runs, and continues with the code that comes after the `if/elif/else`.
#
# Python is doing a lot to make these statements work! Here is a diagram of how these statements work:
#
# 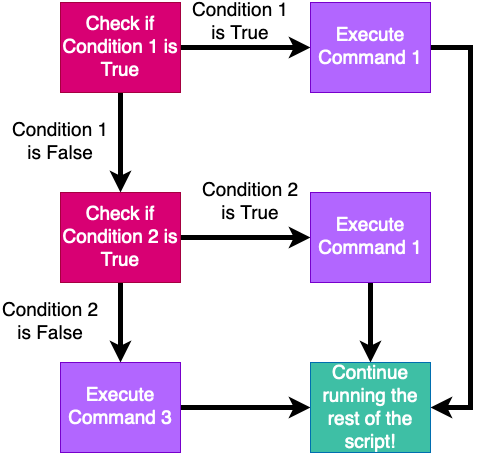
# -
# We can do a lot of really powerful things with `if/elif/else` statements! Let's start by writing code that does the following:
# - If my_list has less than 10 items, print a message saying the list is "short"
# - If my_list has at least 10 items, but fewer than 15 items (so 10, 11, 12, 13, or 14 items), print a message saying the list is "medium"
# - Otherwise, print a message saying the list is "long"
# + colab={} colab_type="code" id="JK80pu224gPz" outputId="9f0670d5-4abd-48e6-aed9-5d69d36c3c20"
# use if/elif/else to print the length of my_list
# + [markdown] colab_type="text" id="y-fBqabh4gP2"
# So what happened here? `my_list` is 12 items long. We got to the first `if` statement, and because the length was not less than 10, we did not execute the line that says `This list is short`. The next line tests if the length is between 10 and 15. This conditional is true, so it executes the line saying `This list is medium length`. Because we got to this line, we skip the `else` statement.
#
# But what if our list was longer? Then what would happen?
# + colab={} colab_type="code" id="AuTSXB1w4gP4"
# append numbers 12, 13, and 14 to my_list
# + colab={} colab_type="code" id="E0NFUr0P4gP7" outputId="4a7d7209-412b-4baf-c062-3d934714762a"
# print the length of my_list
# -
# Let's copy and paste our `if/elif/else` statement from before and see how our code works now that my_list is longer!
# + colab={} colab_type="code" id="v_k76Tfr4gP_" outputId="e70d2b77-c396-42f5-846a-3deb0c4ea3db"
# # copy and paste if/elif/else from before
# + [markdown] colab={} colab_type="code" id="yaPkTJ3h4gQC"
# Great job! You just learned how to use conditionals in Python. You learned:
# - How to use a boolean to only execute code under certain conditions using `if` statements
# - How to further control how code runs using `if/else` and `if/elif/else` statements.
| Lessons/Lesson09_Conditionals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
class RandomForest():
def __init__(self, x, y, n_trees, n_features, sample_sz, depth=10, min_leaf=5):
np.random.seed(12)
if n_features == 'sqrt':
self.n_features = int(np.sqrt(x.shape[1]))
elif n_features == 'log2':
self.n_features = int(np.log2(x.shape[1]))
else:
self.n_features = n_features
# print(self.n_features, "sha: ",x.shape[1])
self.x, self.y, self.sample_sz, self.depth, self.min_leaf = x, y, sample_sz, depth, min_leaf
self.trees = [self.create_tree() for i in range(n_trees)]
def create_tree(self):
idxs = np.random.permutation(len(self.y))[:self.sample_sz]
f_idxs = np.random.permutation(self.x.shape[1])[:self.n_features]
return DecisionTree(self.x.iloc[idxs], self.y[idxs], self.n_features, f_idxs,
idxs=np.array(range(self.sample_sz)),depth = self.depth, min_leaf=self.min_leaf)
def predict(self, x):
return np.mean([t.predict(x) for t in self.trees], axis=0)
def std_agg(cnt, s1, s2): return math.sqrt((s2/cnt) - (s1/cnt)**2)
class DecisionTree():
def __init__(self, x, y, n_features, f_idxs,idxs,depth=10, min_leaf=5):
self.x, self.y, self.idxs, self.min_leaf, self.f_idxs = x, y, idxs, min_leaf, f_idxs
self.depth = depth
# print(f_idxs)
# print(self.depth)
self.n_features = n_features
self.n, self.c = len(idxs), x.shape[1]
self.val = np.mean(y[idxs])
self.score = float('inf')
self.find_varsplit()
def find_varsplit(self):
for i in self.f_idxs: self.find_better_split(i)
if self.is_leaf: return
x = self.split_col
lhs = np.nonzero(x<=self.split)[0]
rhs = np.nonzero(x>self.split)[0]
lf_idxs = np.random.permutation(self.x.shape[1])[:self.n_features]
rf_idxs = np.random.permutation(self.x.shape[1])[:self.n_features]
self.lhs = DecisionTree(self.x, self.y, self.n_features, lf_idxs, self.idxs[lhs], depth=self.depth-1, min_leaf=self.min_leaf)
self.rhs = DecisionTree(self.x, self.y, self.n_features, rf_idxs, self.idxs[rhs], depth=self.depth-1, min_leaf=self.min_leaf)
def find_better_split(self, var_idx):
x, y = self.x.values[self.idxs,var_idx], self.y[self.idxs]
sort_idx = np.argsort(x)
sort_y,sort_x = y[sort_idx], x[sort_idx]
rhs_cnt,rhs_sum,rhs_sum2 = self.n, sort_y.sum(), (sort_y**2).sum()
lhs_cnt,lhs_sum,lhs_sum2 = 0,0.,0.
for i in range(0,self.n-self.min_leaf-1):
xi,yi = sort_x[i],sort_y[i]
lhs_cnt += 1; rhs_cnt -= 1
lhs_sum += yi; rhs_sum -= yi
lhs_sum2 += yi**2; rhs_sum2 -= yi**2
if i<self.min_leaf or xi==sort_x[i+1]:
continue
lhs_std = std_agg(lhs_cnt, lhs_sum, lhs_sum2)
rhs_std = std_agg(rhs_cnt, rhs_sum, rhs_sum2)
curr_score = lhs_std*lhs_cnt + rhs_std*rhs_cnt
if curr_score<self.score:
self.var_idx,self.score,self.split = var_idx,curr_score,xi
@property
def split_name(self): return self.x.columns[self.var_idx]
@property
def split_col(self): return self.x.values[self.idxs,self.var_idx]
@property
def is_leaf(self): return self.score == float('inf') or self.depth <= 0
def predict(self, x):
return np.array([self.predict_row(xi) for xi in x])
def predict_row(self, xi):
if self.is_leaf: return self.val
t = self.lhs if xi[self.var_idx]<=self.split else self.rhs
return t.predict_row(xi)
# -
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import matplotlib.dates as mdates
from pandas import DatetimeIndex
import math
import seaborn as sns
sns.set(style='darkgrid', context='talk', palette='Dark2')
my_year_month_fmt = mdates.DateFormatter('%m/%y')
plt.rcParams['figure.figsize'] = (15, 9)
from datetime import datetime
import datetime
import pandas_datareader as web
start = datetime.datetime(2014, 1, 1)
end = datetime.datetime(2020, 1, 1)
df_amzn = web.DataReader('AMZN', 'yahoo', start, end)
amzn_df = df_amzn
amzn_df.head()
amzn_df=amzn_df.reset_index()
amzn_df['Date'] = pd.to_datetime(amzn_df['Date'])
amzn_df
amzn_df.info()
start = datetime.datetime(2014, 1, 1)
end = datetime.datetime(2020, 1, 1)
gspc = web.DataReader('^GSPC', 'yahoo', start, end)
snp = gspc
snp.head()
snp=snp.reset_index()
snp['Date'] = pd.to_datetime(snp['Date'])
snp.info()
snp.head()
amzn_df.index=DatetimeIndex(amzn_df['Date'])
snp.index=DatetimeIndex(snp['Date'])
X = amzn_df
print(X)
X['Stock_Price'] = (2*amzn_df.High + amzn_df.Low + amzn_df.Close)/4
X['Stock_Price']
snp['High']
X['SP500'] = (2*snp.High + snp.Low + snp.Close)/4
X
from sklearn.preprocessing import StandardScaler
col_names = ['SP500']
features = X[col_names]
scaler = StandardScaler().fit(features.values)
features = scaler.transform(features.values)
X[col_names] = features
X.drop(columns=['Date', 'Open', 'High', 'Low', 'Close','Adj Close'],inplace=True)
X.head()
fig, (ax1) = plt.subplots(1, 1, figsize=(16,5))
ax1.plot(X.index,X['Stock_Price'])
ax1.set_ylabel('Stock_Price')
y = X.Stock_Price
X.drop(['Stock_Price'],inplace = True,axis=1)
rf = RandomForest(x=X,y=y,n_trees=30,n_features=3,sample_sz=100)
tree = rf.create_tree()
y_pred = rf.predict(X.tail(90).values)
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(25,9))
ax1.plot(X.tail(90).index,y_pred,label="Predicted stock price(USD)")
ax1.set_ylabel("Predictions")
ax2.set_ylabel("Actual")
ax1.legend(loc='best')
ax2.plot(X.tail(90).index,y.tail(90),label="Actual stock prices(USD)")
ax2.legend(loc='best')
# exponential weighted functions
ema_short = X.ewm(span=7, adjust=False).mean()
y_short = y.ewm(span=7, adjust=False).mean()
rf2 = RandomForest(x=ema_short,y=y_short,n_trees=30,n_features=3,sample_sz=100)
y_pred_short = rf.predict(ema_short.tail(90).values)
rf2.create_tree()
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(25,9))
ax1.plot(ema_short.tail(90).index,y_pred_short,label="Predicted stock price(USD)")
ax2.plot(X.tail(90).index,y_short.tail(90),label="Actual stock prices(USD)")
ax1.set_ylabel("Predictions")
ax2.set_ylabel("Actual")
ax1.legend(loc='best')
ax2.legend(loc='best')
trading_signal_week = pd.DataFrame(np.sign(np.diff(y_pred_short)),index=X.tail(89).index,columns=["signal"])
trading_signal_week.head()
X.head()
trading_signal_week = trading_signal_week.shift(1)
# +
fig, (ax2) = plt.subplots(1, 1, figsize=(16,7))
duration = 89
y_temp = pd.DataFrame(y_short.tail(duration),index =X.tail(duration).index)
ax2.plot(X.tail(duration).index,y.tail(duration),label="Stock Price")
ax2.set_ylabel("Actual Price for last 90 days")
ax2.legend(loc='best')
# # Plot the buy signals
ax2.plot(trading_signal_week.tail(duration).loc[trading_signal_week.signal == 1.0].index,
y.tail(duration)[trading_signal_week.signal == 1.0],'^', markersize=10, color='y')
# Plot the sell signals
ax2.plot(trading_signal_week.tail(duration).loc[trading_signal_week.signal == -1.0].tail(duration).index,
y.tail(duration)[trading_signal_week.signal == -1.0],'v', markersize=10, color='r')
# +
# Set the initial capital
initial_capital= float(10000.0)
# Create a DataFrame 'positions'
positions = pd.DataFrame(index=trading_signal_week.index).fillna(0.0)
# Buy
positions['stock_price'] = trading_signal_week['signal']
# Initialize the portfolio with value owned
portfolio = positions.multiply(y_short.tail(90), axis=0)
# Store the difference in shares owned
pos_diff = positions.diff()
# Add 'holdings' to portfolio
portfolio['holdings'] = (positions.multiply(y_short.tail(90), axis=0)).sum(axis=1)
# Add 'cash' to portfolio
portfolio['cash'] = initial_capital - (pos_diff.multiply(y_short.tail(90), axis=0)).sum(axis=1).cumsum()
# Add 'total' & 'returns' to portfolio
portfolio['total'] = portfolio['cash'] + portfolio['holdings']
portfolio['returns'] = portfolio['total'].pct_change()
# -
portfolio = portfolio.loc["2019-08-01":]
portfolio
fig, ax1 = plt.subplots(1, 1, figsize=(30,7))
duration = 89
ax1.plot(portfolio['total'])
# +
# Isolate the returns of the strategy
returns = portfolio['returns']
print(returns)
# 90 day Sharpe ratio
#sharpe_ratio = np.sqrt(90) * (returns.mean() / returns.std())
# Print the Sharpe ratio
#print(sharpe_ratio)
# +
from sklearn.metrics import mean_squared_error
from math import sqrt
rms = sqrt(mean_squared_error(y_short.tail(90), y_pred_short))
# -
rms
| AmazonStockAnalysis-main/random_forest_amzn.ipynb |
# +
try:
from tensorflow import keras
except ModuleNotFoundError:
# %pip install -qq tensorflow
from tensorflow import keras
import tensorflow as tf
import numpy as np
import scipy
import matplotlib.pyplot as plt
try:
import pandas as pd
except ModuleNotFoundError:
# %pip install -qq pandas
import pandas as pd
try:
import sklearn
except ModuleNotFoundError:
# %pip install -qq scikit-learn
import sklearn
from time import time
import os
figdir = "figures"
def savefig(fname): plt.savefig(os.path.join(figdir, fname))
# print(tf.__version__)
np.random.seed(0)
mnist = keras.datasets.mnist
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
train_images = train_images / 255.0
test_images = test_images / 255.0
# print(np.shape(train_images))
# print(np.shape(test_images))
#(60000, 28, 28)
#(10000, 28, 28)
plt.figure(figsize=(10, 10))
for i in range(25):
plt.subplot(5, 5, i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
savefig("mnist-data.pdf")
plt.show()
| notebooks/book1/01/mnist_viz_tf.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .js
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Spark 1.6.0 (EclairJS)
// language: javascript
// name: eclair
// ---
// # Movie recommder example
// this example is a EclairJS (JavaScript) implementation of [movie recommending](https://github.com/jadianes/spark-movie-lens/blob/master/notebooks/building-recommender.ipynb).
// #### This notebook requires the following additional setup
// * Download the [movieLens rating dataset](http://grouplens.org/datasets/movielens/) and unzip is a location that is
// * [ml-latest-small.zip](http://files.grouplens.org/datasets/movielens/ml-latest-small.zip)
// * [ml-latest.zip](http://files.grouplens.org/datasets/movielens/ml-latest.zip) accessible by Spark.
//
// ## Create the spark context and globals
// +
var SparkContext = require('eclairjs/SparkContext');
var SparkConf = require('eclairjs/SparkConf');
var sparkConf = new SparkConf()
.set("spark.executor.memory", "10g")
.set("spark.driver.memory", "6g")
.setMaster("local[*]")
.setAppName("movie_recommender");
var sc = new SparkContext(sparkConf);
var Tuple2 = require('eclairjs/Tuple2');
var Tuple3 = require('eclairjs/Tuple3');
var ALS = require('eclairjs/mllib/recommendation/ALS');
var Rating = require('eclairjs/mllib/recommendation/Rating');
// -
// ## Set the path to the movieLens Datasets
//
var pathToSmallDataset = '../data/mllib/ml-latest-small';
var pathToCompleteDataset = '../data/mllib/ml-latest-small';
// ## load the small rating dataset
// +
var small_ratings_raw_data = sc.textFile(pathToSmallDataset + '/ratings.csv');
var small_ratings_raw_data_header = small_ratings_raw_data.take(1)[0];
var small_ratings_data = small_ratings_raw_data
.filter(function(line, small_ratings_raw_data_header) {
// filters out the header
return line != small_ratings_raw_data_header;
}, [small_ratings_raw_data_header])
.map(function(line, Rating) {
var tokens = line.split(",");
return new Rating(tokens[0],tokens[1],tokens[2]);
},[Rating])
.cache();
JSON.stringify(small_ratings_data.take(3));
// -
// ## load the small moving dataset
// +
var small_movies_raw_data = sc.textFile(pathToSmallDataset + '/movies.csv');
var small_movies_raw_data_header = small_movies_raw_data.take(1)[0];
var small_movies_data = small_movies_raw_data
.filter(function(line, small_movies_raw_data_header) {
// filters out the header
return line != small_movies_raw_data_header;
}, [small_movies_raw_data_header])
.map(function(line, Tuple2) {
var fields = line.split(",");
return new Tuple2(parseInt(fields[0]), fields[1]);
}, [Tuple2])
.cache();
var small_movies_titles = small_movies_data
.mapToPair(function( tuple2, Tuple2) { // Tuple2
return new Tuple2(tuple2[0], tuple2[1]);
}, [Tuple2]);
JSON.stringify(small_movies_data.take(3));
// -
// ## Selecting ALS parameters using the small dataset
// In order to determine the best ALS parameters, we will use the small dataset. We need first to split it into train, validation, and test datasets.
// +
var seed = 0;
var split = small_ratings_data.randomSplit([0.6, 0.2, 0.2], seed)
var training_RDD = split[0];
var validation_RDD = split[1];
var test_RDD = split[2];
var validation_for_predict_RDD = validation_RDD
.map(function(rating, Tuple2) {
return new Tuple2(rating.user(), rating.product());
}, [Tuple2]);
JSON.stringify(validation_for_predict_RDD.take(3));
// -
// ## Proceed with the training phase.
// +
seed = 5
var iterations = 10
var regularization_parameter = 0.1
var ranks = [4, 8, 12];
var errors = [0, 0, 0];
var err = 0
var tolerance = 0.02
var min_error = Number.POSITIVE_INFINITY
var best_rank = -1
var best_iteration = -1
var blocks = -1;
var lambda = regularization_parameter;
ranks.forEach(function(rank) {
var model = ALS.train(training_RDD, rank, iterations, regularization_parameter, blocks, seed);
var predictions = model.predict(validation_for_predict_RDD)
.mapToPair(function(rating, Tuple2) {
return new Tuple2(new Tuple2(rating.user(), rating.product()), rating.rating());
}, [Tuple2]);
var rates_and_preds = validation_RDD
.mapToPair(function(rating, Tuple2) {
return new Tuple2(new Tuple2(rating.user(), rating.product()), rating.rating());
}, [Tuple2])
.join(predictions);
var t = rates_and_preds
.mapToFloat(function(tuple) {
var y =tuple._2()._1() - tuple._2()._2();
return Math.pow(y, 2);
});
var error = Math.sqrt(t.mean());
errors[err] = error;
err += 1;
if (error < min_error) {
min_error = error;
best_rank = rank;
}
});
"The best model was trained with rank " +best_rank;
// -
// ## To build our recommender model, we will use the complete dataset.
// +
var complete_ratings_raw_data =
sc.textFile(pathToCompleteDataset + '/ratings.csv');
var complete_ratings_raw_data_header = complete_ratings_raw_data.take(1)[0];
var complete_ratings_data = complete_ratings_raw_data
.filter(function (line, complete_ratings_raw_data_header) {
return line != complete_ratings_raw_data_header;
}, [complete_ratings_raw_data_header])
.map(function( line, Rating) {
var fields = line.split(",");
var userId = parseInt(fields[0]);
var movieId = parseInt(fields[1]);
var rating = parseFloat(fields[2]);
return new Rating(userId, movieId, rating);
}, [Rating])
.cache();
JSON.stringify("There are recommendations in the complete dataset: " + complete_ratings_data.count());
// -
// ## We test on our testing set.
// +
var splits2 = complete_ratings_data.randomSplit([0.7, 0.3], 0);
training_RDD = splits2[0];
test_RDD = splits2[1];
var complete_model = ALS.train(training_RDD, best_rank, iterations, regularization_parameter, blocks, seed);
var test_for_predict_RDD = test_RDD
.map(function (rating, Tuple2) {
return new Tuple2(rating.user(), rating.product());
}, [Tuple2]);
var predictions = complete_model.predict(test_for_predict_RDD)
.mapToPair(function( rating, Tuple2) {
return new Tuple2(new Tuple2(rating.user(), rating.product()), rating.rating());
}, [Tuple2]);
var rates_and_preds = test_RDD
.mapToPair(function( rating, Tuple2) {
return new Tuple2(new Tuple2(rating.user(), rating.product()), rating.rating());
}, [Tuple2])
.join(predictions);
var t = rates_and_preds
.mapToFloat( function( x) {
return Math.pow(x._2()._1() - x._2()._2(), 2);
});
var error = Math.sqrt(t.mean());
JSON.stringify("For testing data the RMSE is " + error);
// -
// ## How to make recommendations
// So let's first load the movies complete file for later use.
// +
var complete_movies_raw_data =
sc.textFile(pathToCompleteDataset + '/movies.csv');
var complete_movies_raw_data_header = complete_movies_raw_data.take(1)[0];
var complete_movies_data = complete_movies_raw_data
.filter(function(line, complete_movies_raw_data_header) {
// filters out the header
return line != complete_movies_raw_data_header;
}, [complete_movies_raw_data_header])
.map(function(line, Tuple2) {
var fields = line.split(",");
return new Tuple2(parseInt(fields[0]), fields[1]);
}, [Tuple2]).cache();
var complete_movies_titles = complete_movies_data
.mapToPair(function( tuple2, Tuple2) { // Tuple2
return new Tuple2(tuple2._1(), tuple2._2());
}, [Tuple2]);
JSON.stringify("There are movies in the complete dataset " + complete_movies_titles.count());
// -
// ## Give recommendations of movies
// Another thing we want to do, is give recommendations
// of movies with a certain minimum number of ratings. For that, we need to count the number of ratings per movie.
//
// +
var movie_ID_with_ratings_RDD = complete_ratings_data
.mapToPair(function( rating, Tuple2) {
return new Tuple2(rating.product(), rating.rating());
}, [Tuple2])
.groupByKey();
var movie_ID_with_avg_ratings_RDD = movie_ID_with_ratings_RDD
.mapToPair(function( ID_and_ratings_tuple, Tuple2) {
var w = ID_and_ratings_tuple._2();
var count = 0;
var sum = 0;
for (var i = 0; i < w.length; i++) {
var r = w[i];
sum += r;
count++;
}
var avgRating = sum / count;
return new Tuple2(ID_and_ratings_tuple._1(), new Tuple2(count, avgRating));
}, [Tuple2]);
var movie_rating_counts_RDD = movie_ID_with_avg_ratings_RDD
.mapToPair(function( ID_with_avg_ratings, Tuple2) {
return new Tuple2(ID_with_avg_ratings._1(), ID_with_avg_ratings._2()._1()); // movieID, rating count
}, [Tuple2]);
JSON.stringify("movie_ID_with_avg_ratings_RDD " + movie_ID_with_avg_ratings_RDD.take(10));
// -
// ## Rate some movies for the new user.
// +
var new_user_ID = 0;
// The format of each line is (userID, movieID, rating)
var new_user_ratings = [
new Rating(0, 260, 9), // Star Wars (1977)
new Rating(0, 1, 8), // Toy Story (1995)
new Rating(0, 16, 7), // Casino (1995)
new Rating(0, 25, 8), // Leaving Las Vegas (1995)
new Rating(0, 32, 9), // Twelve Monkeys (a.k.a. 12 Monkeys) (1995)
new Rating(0, 335, 4), // Flintstones, The (1994)
new Rating(0, 379, 3), // Timecop (1994)
new Rating(0, 296, 7), // Pulp Fiction (1994)
new Rating(0, 858, 10), // Godfather, The (1972)
new Rating(0, 50, 8) // Usual Suspects, The (1995)
];
var new_user_ratings_RDD = sc.parallelize(new_user_ratings);
JSON.stringify("New user ratings: " + new_user_ratings_RDD.take(10));
// -
// ## Add them to the data we will use to train our recommender model.
// +
var complete_data_with_new_ratings_RDD = complete_ratings_data.union(new_user_ratings_RDD);
var new_ratings_model =
ALS.train(complete_data_with_new_ratings_RDD, best_rank,
iterations, regularization_parameter, blocks, seed);
/*
Let's now get some recommendations
*/
// get just movie IDs
var new_user_ratings_ids = [];
for (var i = 0; i < new_user_ratings.length; i++) {
new_user_ratings_ids.push(new_user_ratings[i].product());
}
// keep just those not on the ID list
var new_user_unrated_movies_RDD = complete_movies_data.filter(function( tuple, new_user_ratings_ids) {
if (new_user_ratings_ids.indexOf(tuple._1()) < 0) {
return true;
} else {
return false;
}
}, [new_user_ratings_ids])
.map(function( tuple, new_user_ID, Tuple2) {
return new Tuple2(new_user_ID, tuple._1());
}, [new_user_ID, Tuple2]);
// Use the input RDD, new_user_unrated_movies_RDD,
//with new_ratings_model.predictAll() to predict new ratings for the movies
var new_user_recommendations_RDD = new_ratings_model.predict(new_user_unrated_movies_RDD);
// Transform new_user_recommendations_RDD into pairs of the form (Movie ID, Predicted Rating)
var new_user_recommendations_rating_RDD = new_user_recommendations_RDD.mapToPair( function( rating, Tuple2) {
return new Tuple2(rating.product(), rating.rating());
}, [Tuple2]);
var new_user_recommendations_rating_title_and_count_RDD = new_user_recommendations_rating_RDD
.join(complete_movies_titles)
.join(movie_rating_counts_RDD);
"new_user_recommendations_rating_title_and_count_RDD " + new_user_recommendations_rating_title_and_count_RDD.count();
// -
// ## Flatten the RDD
// we need to flat this down a bit in order to have (Title, Rating, Ratings Count).
// +
var new_user_recommendations_rating_title_and_count_RDD2 =
new_user_recommendations_rating_title_and_count_RDD.map(function( t, Tuple3) {
var x = new Tuple3(t._2()._1()._2(), t._2()._1()._1(), t._2()._2());
return x;
}, [Tuple3]);
"count" + new_user_recommendations_rating_title_and_count_RDD2.count();
"new_user_recommendations_rating_title_and_count_RDD2" +
JSON.stringify(new_user_recommendations_rating_title_and_count_RDD2.take(3));
// -
// ## Get highest rated recommendations
// Finally, get the highest rated recommendations for the new user, filtering out movies with less than 25 ratings.
// +
var new_user_recommendations_rating_title_and_count_RDD2_filtered =
new_user_recommendations_rating_title_and_count_RDD2.filter(function( tuple3) {
if (tuple3._3() < 25) {
return false;
} else {
return true;
}
});
/*
list top 25
*/
var top_movies = new_user_recommendations_rating_title_and_count_RDD2_filtered.takeOrdered(25,
function(tuple3_a, tuple3_b){
var aRate = tuple3_a._2();
var bRate = tuple3_b._2();
return aRate > bRate ? -1 : aRate == bRate? 0 : 1;
});
var str = "TOP recommended movies (with more than 25 reviews):\n\n";
for (var i = 0; i < top_movies.length; i++) {
str += top_movies[i]._1() + " average rating " +
top_movies[i]._2() + " number of ratings " + top_movies[i]._3() + "\n";
}
// -
// ## Look up movie Title from ID
//
// +
var DataTypes = require('eclairjs/sql/types/DataTypes');
var RowFactory = require('eclairjs/sql/RowFactory');
var SQLContext = require('eclairjs/sql/SQLContext');
var movieID = "1";
//Generate the schema
var sqlContext = new SQLContext(sc);
var fields = [];
fields.push(DataTypes.createStructField("id", DataTypes.IntegerType, true));
fields.push(DataTypes.createStructField("title", DataTypes.StringType, true));
var schema = DataTypes.createStructType(fields);
var rowRDD = complete_movies_data.map(function (tuple2, RowFactory) {
return RowFactory.create([tuple2._1(), tuple2._2()]);
}, [RowFactory]);
//Apply the schema to the RDD.
var complete_movies_titlesDF = sqlContext.createDataFrame(rowRDD, schema);
var col = complete_movies_titlesDF.col("id");
//var col2 = complete_movies_titlesDF.col("title");
var testCol = col.equalTo(movieID);
var result = complete_movies_titlesDF.filter(testCol).collect();
JSON.stringify(result);
// -
// ## Lookup movie id from title
// +
var movieTitle = "Toy Story";
//var col = complete_movies_titlesDF.col("id");
var col2 = complete_movies_titlesDF.col("title");
var testCol = col2.contains(movieTitle);
var result = complete_movies_titlesDF.filter(testCol).collect();
JSON.stringify(result);
// -
// ## Predicted rating for a particular movie for a given user.
// +
var my_movie = sc.parallelizePairs([new Tuple2(0, 500)]); // Quiz Show (1994)
var individual_movie_rating_RDD = new_ratings_model.predict(my_movie);
"Predicted rating for movie " + individual_movie_rating_RDD.take(1);
// -
| server/examples/notebooks/movie_recomennder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from IPython.core.display import display, HTML
display(HTML("<style>.container{ width:95%}</style>"))
import sys
print(sys.executable)
#import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
from IPython.display import set_matplotlib_formats
set_matplotlib_formats('retina')
import seaborn as sns
from sklearn import preprocessing
from IPython.display import HTML
HTML('''<script>
code_show=false;
function code_toggle() {
if (code_show){
$('div.input').hide();
} else {
$('div.input').show();
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
El codigo Python de este cuaderno está oculto para facilitar la lectura.
Para mostrar/ocultar el código, haz click <a href="javascript:code_toggle()">aquí.</a>.''')
# -
# ## Load metrics
# +
import pandas as pd
df_alg=pd.read_csv('output/metrics/merged_output_genetic.txt')
convert_dict = {'Dataset': "string",
'Algorithm': "string",
'Population Length': "int64",
'Generations': "int64",
'Selection Scheme': "string",
'Selection Candidates': "int64",
'Crossover Scheme': "string",
'Crossover Probability': "float64",
'Mutation Scheme': "string",
'Mutation Probability': "float64",
'Replacement Scheme': "string",
'Time(s)': "float64",
'AvgValue': "float64",
'BestAvgValue': "float64",
'BestGeneration': "int64",
'HV': "float64",
'Spread': "float64",
'NumSolutions': "float64",
'Spacing': "float64",
'NumGenerations': "int64"
}
df_alg = df_alg.astype(convert_dict)
df_alg.loc[(df_alg.Dataset == 'dataset1'),'Dataset']='1'
df_alg.loc[(df_alg.Dataset == 'dataset2'),'Dataset']='2'
#display(df_alg.head(2))
df_alg2=pd.read_csv('output/metrics/merged_output_grasp.txt',header=0)
convert_dict = {'Dataset': "string",
'Algorithm': "string",
'Iterations': "float64",
'Solutions per Iteration': "int64",
'Initialization Type': "string",
'Local Search Type': "string",
'Path Relinking': "string",
'Time(s)': "float64",
'AvgValue': "float64",
'BestAvgValue': "float64",
'HV': "float64",
'Spread': "float64",
'NumSolutions': "int64",
'Spacing': "float64",
'NumGenerations': "int64"
}
df_alg2 = df_alg2.astype(convert_dict)
#display(df_alg2.head(200))
df_alg = df_alg.append(df_alg2)
#display(df_alg.head(200))
# -
# ## Pareto analysis
# Change ```dataset``` value to load different dataset Paretos
# +
import matplotlib.pyplot as plt
import numpy as np
from algorithms.GRASP.GRASP import GRASP
from algorithms.genetic.nsgaii.nsgaii_algorithm import NSGAIIAlgorithm
from algorithms.genetic.geneticnds.geneticnds_algorithm import GeneticNDSAlgorithm
sizes=[30,25,20,15,10,7,5]
markers=["+","x","s","v","h","o"]
labels=["Random","Single-Objective GA","NSGA-II","GPPR"]
#labels=["Random","Mono-Objective GA","NSGA-II","GPPR-noLocal-PR","GPPR-SO-PR","GPPR-MO-PR"]
datasets=["1","2","s1","s2","s3"]
dataset="2"
seed=10
generations=100
solutions_per_iteration=100
population_length=100
gens_genetic=100
algorithms = [
GRASP(dataset=dataset,iterations=generations,solutions_per_iteration=solutions_per_iteration,seed=seed,
init_type="uniform",local_search_type="None",
path_relinking_mode="None"),
GeneticNDSAlgorithm(dataset_name=dataset,random_seed=seed,population_length=100,max_generations=100,crossover_prob=0.8,
crossover="onepoint",mutation_prob=1.0,mutation="flip1bit",replacement="elitism"),
NSGAIIAlgorithm(dataset_name=dataset,random_seed=seed,population_length=population_length,max_generations=gens_genetic
,crossover_prob=0.6,crossover="onepoint",mutation_prob=1.0,mutation="flip1bit",replacement="elitism"),
#GRASP(dataset=dataset,iterations=generations,solutions_per_iteration=solutions_per_iteration,seed=seed,
# init_type="stochastically",local_search_type="None",
# path_relinking_mode="after_local"),
#GRASP(dataset=dataset,iterations=500,solutions_per_iteration=200,seed=seed,
# init_type="stochastically",local_search_type="best_first_neighbor_sorted_score",
# path_relinking_mode="after_local"),
#GRASP(dataset=dataset,iterations=generations,solutions_per_iteration=solutions_per_iteration,seed=seed,
# init_type="stochastically",local_search_type="best_first_neighbor_sorted_domination",
# path_relinking_mode="after_local"),
GRASP(dataset=dataset,iterations=generations,solutions_per_iteration=solutions_per_iteration,seed=seed,
init_type="stochastically",local_search_type="best_first_neighbor_random_domination",
path_relinking_mode="after_local"),
]
for i in range(len(algorithms)):
if "GRASP" in algorithms[i].file:
file = "output/paretos/pareto-grasp-"+algorithms[i].file
else:
file = "output/paretos/pareto-genetic-"+algorithms[i].file
data = np.loadtxt(file,delimiter=',', dtype=float)
x,y=data.T
plt.scatter(x,y,label=labels[i],s=50,marker=markers[i])
#file = "output/backtracking.txt"
#data = np.loadtxt(file,delimiter=',', dtype=float)
#x,y=data.T
#plt.scatter(x,y,label="optimo",s=10)
plt.title(dataset)
plt.xlabel('Effort', fontsize=12)
plt.ylabel('Satisfaction', fontsize=12)
plt.legend(loc="lower right")
plt.title("Dataset "+dataset)
plt.grid(True)
plt.rcParams['figure.figsize'] = [16, 10]
plt.rcParams['figure.dpi'] = 200 # 200 e.g. is really fine, but slower
plt.show()
# -
# ## Metrics analysis
# +
from sklearn import preprocessing
from scipy.stats import ranksums
import numpy as np
import plotly.graph_objects as go
import plotly.offline as pyo
import math
class AlgorithmDataGenetic():
def __init__(self,a,rs,d,p,g,ss,sc,cs,cp,ms,mp):
self.a=a
self.rs=rs
self.d=d
self.p=p
self.g=g
self.ss=ss
self.sc=sc
self.cs=cs
self.cp=cp
self.ms=ms
self.mp=mp
def findConfigurationData(self,df):
return df[(df["Population Length"]==self.p)&(df["Generations"]==self.g)
&(df["Selection Scheme"]==self.ss)&(df["Selection Candidates"]==self.sc)
&(df["Crossover Scheme"]==self.cs)&(df["Crossover Probability"]==self.cp)
&(df["Mutation Scheme"]==self.ms)&(df["Mutation Probability"]==self.mp)
&(df["Algorithm"]==self.a)&(df["Replacement Scheme"]==self.rs)
&(df["Dataset"]==self.d)
]
class AlgorithmDataGrasp():
def __init__(self,a,d,it,so,ini,ls,pr):
self.a=a
self.it=it
self.so=so
self.ls=ls
self.d=d
self.ini=ini
self.pr=pr
def findConfigurationData(self,df):
return df[(df["Iterations"]==self.it)&(df["Solutions per Iteration"]==self.so)
&(df["Local Search Type"]==self.ls)&(df["Initialization Type"]==self.ini)
&(df["Algorithm"]==self.a)&(df["Dataset"]==self.d)&(df["Path Relinking"]==self.pr)
]
dat="1"
datasets=["1","2","s1","s2","s3"]
cols=["HV","Spread","Spacing","NumSolutions","Time(s)"]
maxmin=[1,-1,1,1,-1]
for dat in datasets:
print("------Dataset "+dat+"-----")
algs = [
AlgorithmDataGenetic("GeneticNDSAlgorithm",'elitism',dat,100,100,"tournament",2,"onepoint",0.8,"flip1bit",1.0),
AlgorithmDataGenetic("NSGAIIAlgorithm",'elitism',dat,100,100,"tournament",2,"onepoint",0.6,"flip1bit",1.0),
AlgorithmDataGrasp("GRASP",dat,100,100,"stochastically","best_first_neighbor_random_domination","after_local"),
]
for j in range(len(cols)):
print(cols[j])
results=list()
best_avg=0
best_avgn=10**9
best_alg_index=None
for i in range(len(algs)):
avg=np.mean((algs[i].findConfigurationData(df_alg)[cols])[cols[j]].values)
results.append("{:.3f}".format(avg))
if maxmin[j]>0 and avg>best_avg:
best_avg=avg
best_alg_index=i
elif maxmin[j]<0 and avg<best_avgn:
best_avgn=avg
best_alg_index=i
p_best=True
p_list=[]
for i in range(len(algs)):
if i!=best_alg_index:
dataA=(algs[best_alg_index].findConfigurationData(df_alg)[cols])[cols[j]].values
dataB=(algs[i].findConfigurationData(df_alg)[cols])[cols[j]].values
_, p = ranksums(dataA, dataB)
print("p:",p)
if p>=0.05:
#print(dataA)
#print(dataB)
p_best=False
else:
p_list.append(i)
if p_best:
mark = '*'
else:
mark = ''
for index in p_list:
results[index]=results[index]+'-'
#results[best_alg_index]=results[best_alg_index]+mark
results.insert(0,cols[j])
print(results)
| experimentation_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## String concatenation
part1 = "I want"
amount = 5
what = "eggs"
# We can also use `+` for concatenation!
part1 + amount + what
part1 + str(amount) + what
part1 + " " + str(amount) + " " + what
f"{part1} {amount} {what}"
| Learn Python/09. String Concatenation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# C:\Users\ksg\py_tutorial
from selenium import webdriver
from bs4 import BeautifulSoup
from pprint import pprint
def keyword_setting(word):
change_word = list(word)
for i in range(len(change_word)):
if(change_word[i]==' '):
change_word[i]='+'
change_word = ''.join(change_word)
return change_word
# +
keyword = 'linux 동영상 녹화'
URL = 'https://www.google.com/search?q=' +keyword_setting(keyword)
driver = webdriver.Chrome("C:/Users/ksg/py_tutorial/chromedriver.exe")
driver.implicitly_wait(1)
driver.get(URL)
driver.implicitly_wait(2)
html = driver.page_source
soup = BeautifulSoup(html, 'html.parser')
title_list = soup.find_all(name='div',attrs={'class':'r'})
# -
title_list
for i in title_list:
print(i.text,end=" : ")
print(str(i).split("\"")[3])
print("")
print("http://blog.naver.com/PostView.nhn?blogId=doksg&logNo=221454150493&parentCategoryNo=&categoryNo=19&viewDate=&isShowPopularPosts=true&from=search")
def keyword_setup(keyword):
URL = 'https://www.google.com/search?q=' +keyword_setting(keyword)
driver = webdriver.Chrome("C:/Users/ksg/py_tutorial/chromedriver.exe")
driver.implicitly_wait(1)
driver.get(URL)
driver.implicitly_wait(2)
html = driver.page_source
soup = BeautifulSoup(html, 'html.parser')
title_list = soup.find_all(name='div',attrs={'class':'r'})
return title_list
keyword = "<NAME>"
title_list = keyword_setup(keyword)
for i,val in enumerate(title_list):
title=str(val).split("\"")[3]
if(title[0:36]=="https://m.blog.naver.com/fastcampus/"):
print(i)
print("")
channel_list=["https://www.fastcampus.co.kr/",
"https://m.blog.naver.com/fastcampus/",
"https://m.post.naver.com/fastcampus/",
"http://media.fastcampus.co.kr/"]
for i in channel_list:
print(len(i))
keyword_set=['업무자동화',
'컴퓨터공학 올인원',
'올인원 패키지',
'패스트캠퍼스',
'UX/UI',
'자바스크립트 인강',
'모션그래픽 디자인',
'재무회계 실무',
'부동산 대체투자',
'신사업 발굴',
'디자인 툴',
'데이터 분석',
'딥러닝 강의',
'인공지능 강의',
'HTML/CSS 인강',
'iOS 강의',
'OpenCV 강의',
'디지털 마케팅']
for keyword in keyword_set:
title_list = keyword_setup(keyword)
for i,val in enumerate(title_list):
for channel in channel_list:
title=str(val).split("\"")[3]
if(title[0:len(channel)]==channel):
print(keyword,end=" : ")
print(i,end=" => ")
print(val.text[0:40])
print("")
| .ipynb_checkpoints/contents_ranking-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import numpy as np
import pandas as pd
import sys
import os
import pickle
from sklearn.model_selection import train_test_split
from sklearn.metrics import roc_auc_score
from sklearn.metrics import log_loss
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import label_binarize
from sklearn.ensemble import RandomForestClassifier
import scipy.stats as ss
# -
sys.path.append('../utils')
from simple_impute import simple_imputer
# # Task Specifics
INTERVENTION = 'vent'
RANDOM = 0
MAX_LEN = 240
SLICE_SIZE = 6
GAP_TIME = 6
PREDICTION_WINDOW = 4
OUTCOME_TYPE = 'all'
NUM_CLASSES = 4
CHUNK_KEY = {'ONSET': 0, 'CONTROL': 1, 'ON_INTERVENTION': 2, 'WEAN': 3}
# # Load Data
DATAFILE = '../data/all_hourly_data.h5'
X = pd.read_hdf(DATAFILE,'vitals_labs')
Y = pd.read_hdf(DATAFILE,'interventions')
static = pd.read_hdf(DATAFILE,'patients')
Y = Y[[INTERVENTION]]
print 'Shape of X : ', X.shape
print 'Shape of Y : ', Y.shape
print 'Shape of static : ', static.shape
# # Preprocessing Data
# ## Train-Test Split, Stratified
train_ids, test_ids = train_test_split(static.reset_index(), test_size=0.2,
random_state=RANDOM, stratify=static['mort_hosp'])
split_train_ids, val_ids = train_test_split(train_ids, test_size=0.125,
random_state=RANDOM, stratify=train_ids['mort_hosp'])
# ## Imputation and Standardization of Time Series Features
X_clean = simple_imputer(X,train_ids['subject_id'])
def minmax(x):# normalize
mins = x.min()
maxes = x.max()
x_std = (x - mins) / (maxes - mins)
return x_std
def std_time_since_measurement(x):
idx = pd.IndexSlice
x = np.where(x==100, 0, x)
means = x.mean()
stds = x.std()
x_std = (x - means)/stds
return x_std
idx = pd.IndexSlice
X_std = X_clean.copy()
X_std.loc[:,idx[:,'mean']] = X_std.loc[:,idx[:,'mean']].apply(lambda x: minmax(x))
X_std.loc[:,idx[:,'time_since_measured']] = X_std.loc[:,idx[:,'time_since_measured']].apply(lambda x: std_time_since_measurement(x))
X_std.columns = X_std.columns.droplevel(-1)
del X
# ## Categorization of Static Features
# +
def categorize_age(age):
if age > 10 and age <= 30:
cat = 1
elif age > 30 and age <= 50:
cat = 2
elif age > 50 and age <= 70:
cat = 3
else:
cat = 4
return cat
def categorize_ethnicity(ethnicity):
if 'AMERICAN INDIAN' in ethnicity:
ethnicity = 'AMERICAN INDIAN'
elif 'ASIAN' in ethnicity:
ethnicity = 'ASIAN'
elif 'WHITE' in ethnicity:
ethnicity = 'WHITE'
elif 'HISPANIC' in ethnicity:
ethnicity = 'HISPANIC/LATINO'
elif 'BLACK' in ethnicity:
ethnicity = 'BLACK'
else:
ethnicity = 'OTHER'
return ethnicity
# -
# use gender, first_careunit, age and ethnicity for prediction
static_to_keep = static[['gender', 'age', 'ethnicity', 'first_careunit', 'intime']]
static_to_keep.loc[:, 'intime'] = static_to_keep['intime'].astype('datetime64').apply(lambda x : x.hour)
static_to_keep.loc[:, 'age'] = static_to_keep['age'].apply(categorize_age)
static_to_keep.loc[:, 'ethnicity'] = static_to_keep['ethnicity'].apply(categorize_ethnicity)
static_to_keep = pd.get_dummies(static_to_keep, columns = ['gender', 'age', 'ethnicity', 'first_careunit'])
# ## Create Feature Matrix
# merge time series and static data
X_merge = pd.merge(X_std.reset_index(), static_to_keep.reset_index(), on=['subject_id','icustay_id','hadm_id'])
# add absolute time feature
abs_time = (X_merge['intime'] + X_merge['hours_in'])%24
X_merge.insert(4, 'absolute_time', abs_time)
X_merge.drop('intime', axis=1, inplace=True)
X_merge = X_merge.set_index(['subject_id','icustay_id','hadm_id','hours_in'])
del X_std, X_clean
# ## Make Tensors
# +
def create_x_matrix(x):
zeros = np.zeros((MAX_LEN, x.shape[1]-4))
x = x.values
x = x[:(MAX_LEN), 4:]
zeros[0:x.shape[0], :] = x
return zeros
def create_y_matrix(y):
zeros = np.zeros((MAX_LEN, y.shape[1]-4))
y = y.values
y = y[:,4:]
y = y[:MAX_LEN, :]
zeros[:y.shape[0], :] = y
return zeros
# -
x = np.array(list(X_merge.reset_index().groupby('subject_id').apply(create_x_matrix)))
y = np.array(list(Y.reset_index().groupby('subject_id').apply(create_y_matrix)))[:,:,0]
lengths = np.array(list(X_merge.reset_index().groupby('subject_id').apply(lambda x: x.shape[0])))
keys = pd.Series(X_merge.reset_index()['subject_id'].unique())
print("X tensor shape: ", x.shape)
print("Y tensor shape: ", y.shape)
print("lengths shape: ", lengths.shape)
# ## Stratified Sampling
train_indices = np.where(keys.isin(train_ids['subject_id']))[0]
test_indices = np.where(keys.isin(test_ids['subject_id']))[0]
train_static = train_ids
split_train_indices = np.where(keys.isin(split_train_ids['subject_id']))[0]
val_indices = np.where(keys.isin(val_ids['subject_id']))[0]
X_train = x[split_train_indices]
Y_train = y[split_train_indices]
X_test = x[test_indices]
Y_test = y[test_indices]
X_val = x[val_indices]
Y_val = y[val_indices]
lengths_train = lengths[split_train_indices]
lengths_val = lengths[val_indices]
lengths_test = lengths[test_indices]
print("Training size: ", X_train.shape[0])
print("Validation size: ", X_val.shape[0])
print("Test size: ", X_test.shape[0])
# ## Make Windows
def make_3d_tensor_slices(X_tensor, Y_tensor, lengths):
num_patients = X_tensor.shape[0]
timesteps = X_tensor.shape[1]
num_features = X_tensor.shape[2]
X_tensor_new = np.zeros((lengths.sum(), SLICE_SIZE, num_features + 1))
Y_tensor_new = np.zeros((lengths.sum()))
current_row = 0
for patient_index in range(num_patients):
x_patient = X_tensor[patient_index]
y_patient = Y_tensor[patient_index]
length = lengths[patient_index]
for timestep in range(length - PREDICTION_WINDOW - GAP_TIME - SLICE_SIZE):
x_window = x_patient[timestep:timestep+SLICE_SIZE]
y_window = y_patient[timestep:timestep+SLICE_SIZE]
x_window = np.concatenate((x_window, np.expand_dims(y_window,1)), axis=1)
result_window = y_patient[timestep+SLICE_SIZE+GAP_TIME:timestep+SLICE_SIZE+GAP_TIME+PREDICTION_WINDOW]
result_window_diff = set(np.diff(result_window))
#if 1 in result_window_diff: pdb.set_trace()
gap_window = y_patient[timestep+SLICE_SIZE:timestep+SLICE_SIZE+GAP_TIME]
gap_window_diff = set(np.diff(gap_window))
#print result_window, result_window_diff
if OUTCOME_TYPE == 'binary':
if max(gap_window) == 1:
result = None
elif max(result_window) == 1:
result = 1
elif max(result_window) == 0:
result = 0
if result != None:
X_tensor_new[current_row] = x_window
Y_tensor_new[current_row] = result
current_row += 1
else:
if 1 in gap_window_diff or -1 in gap_window_diff:
result = None
elif (len(result_window_diff) == 1) and (0 in result_window_diff) and (max(result_window) == 0):
result = CHUNK_KEY['CONTROL']
elif (len(result_window_diff) == 1) and (0 in result_window_diff) and (max(result_window) == 1):
result = CHUNK_KEY['ON_INTERVENTION']
elif 1 in result_window_diff:
result = CHUNK_KEY['ONSET']
elif -1 in result_window_diff:
result = CHUNK_KEY['WEAN']
else:
result = None
if result != None:
X_tensor_new[current_row] = x_window
Y_tensor_new[current_row] = result
current_row += 1
X_tensor_new = X_tensor_new[:current_row,:,:]
Y_tensor_new = Y_tensor_new[:current_row]
return X_tensor_new, Y_tensor_new
x_train, y_train = make_3d_tensor_slices(X_train, Y_train, lengths_train)
x_val, y_val = make_3d_tensor_slices(X_val, Y_val, lengths_val)
x_test, y_test = make_3d_tensor_slices(X_test, Y_test, lengths_test)
y_train_classes = label_binarize(y_train, classes=range(NUM_CLASSES))
y_val_classes = label_binarize(y_val, classes=range(NUM_CLASSES))
y_test_classes = label_binarize(y_test, classes=range(NUM_CLASSES))
del X_train, Y_train, X_test, Y_test, X_val, Y_val
print('shape of x_train: ', x_train.shape)
print('shape of x_val: ', x_val.shape)
print('shape of x_test: ', x_test.shape)
# # Random Forest and Logistic Regression
# ## Prepare data
static_col = 17 #static_to_keep.shape[1] - 1
time_series_col = 124 #X_merge.shape[1] - static_col
def remove_duplicate_static(x):
x_static = x[:,0,time_series_col:x.shape[2]-1]
x_timeseries = np.reshape(x[:,:,:time_series_col],(x.shape[0], -1))
x_int = x[:,:,-1]
x_concat = np.concatenate((x_static, x_timeseries, x_int), axis=1)
return x_concat
# concatenate hourly features
x_train_concat = remove_duplicate_static(x_train)
x_val_concat = remove_duplicate_static(x_val)
x_test_concat = remove_duplicate_static(x_test)
print(x_train_concat.shape)
print(x_val_concat.shape)
print(x_test_concat.shape)
# ## Hyperparameter Generation
# +
class DictDist():
def __init__(self, dict_of_rvs): self.dict_of_rvs = dict_of_rvs
def rvs(self, n):
a = {k: v.rvs(n) for k, v in self.dict_of_rvs.items()}
out = []
for i in range(n): out.append({k: vs[i] for k, vs in a.items()})
return out
class Choice():
def __init__(self, options): self.options = options
def rvs(self, n): return [self.options[i] for i in ss.randint(0, len(self.options)).rvs(n)]
# +
N = 10
np.random.seed(RANDOM)
LR_dist = DictDist({
'C': Choice(np.geomspace(1e-3, 1e3, 10000)),
'penalty': Choice(['l2']),
'solver': Choice(['sag']),
'max_iter': Choice([100, 200]),
'class_weight': Choice(['balanced']),
'multi_class': Choice(['multinomial']),
'random_state': Choice([RANDOM])
})
LR_hyperparams_list = LR_dist.rvs(N)
RF_dist = DictDist({
'n_estimators': ss.randint(50, 200),
'max_depth': ss.randint(2, 10),
'min_samples_split': ss.randint(2, 75),
'min_samples_leaf': ss.randint(1, 50),
'class_weight': Choice(['balanced']),
'random_state': Choice([RANDOM])
})
RF_hyperparams_list = RF_dist.rvs(N)
# -
# ## Fit model
# +
def run_basic(model, hyperparams_list, X_train, X_val, X_test):
best_s, best_hyperparams = -np.Inf, None
for i, hyperparams in enumerate(hyperparams_list):
print("On sample %d / %d (hyperparams = %s)" % (i+1, len(hyperparams_list), repr((hyperparams))))
M = model(**hyperparams)
M.fit(X_train, y_train)
s = roc_auc_score(y_val_classes, M.predict_proba(X_val),average='macro')
if s > best_s:
best_s, best_hyperparams = s, hyperparams
print("New Best Score: %.2f @ hyperparams = %s" % (100*best_s, repr((best_hyperparams))))
return run_only_final(model, best_hyperparams, X_train, X_val, X_test)
def run_only_final(model, best_hyperparams, X_train, X_val, X_test):
best_M = model(**best_hyperparams)
best_M.fit(np.concatenate((X_train, X_val)), np.concatenate((y_train, y_val)))
y_pred = best_M.predict_proba(X_test)
auc = roc_auc_score(y_test_classes, y_pred, average=None)
aucmacro = roc_auc_score(y_test_classes, y_pred, average='macro')
return best_M, best_hyperparams, auc, aucmacro
# -
results = {}
for model_name, model, hyperparams_list in [('RF', RandomForestClassifier, RF_hyperparams_list),
('LR', LogisticRegression, LR_hyperparams_list)]:
if model_name not in results: results[model_name] = {}
print("Running model %s " % (model_name))
results[model_name] = run_basic(
model, hyperparams_list, x_train_concat, x_val_concat, x_test_concat)
print("Final results for model %s " % (model_name))
print(results[model_name])
# # CNN
import tensorflow as tf
import keras
from keras.models import Sequential, load_model
from keras.layers import Dense, Dropout, Flatten, Reshape, RepeatVector, Lambda
from keras.layers import Input, Conv2D, Conv1D, Conv3D, MaxPooling2D, MaxPooling1D
from keras.layers import Concatenate
from keras import backend as K
from keras.callbacks import EarlyStopping
from tensorflow import set_random_seed
set_random_seed(RANDOM)
BATCH_SIZE = 128
EPOCHS = 12
DROPOUT = 0.5
from sklearn.utils import class_weight
class_weight = class_weight.compute_class_weight('balanced', np.unique(y_train), y_train)
class_weight = dict(zip(range(len(class_weight)), class_weight))
# +
input_shape = (x_train.shape[1], x_train.shape[2])
inputs = Input(shape=input_shape)
model = Conv1D(64, kernel_size=3,
strides=1,
activation='relu',
input_shape=input_shape,
padding='same',
name='conv2')(inputs)
model = (MaxPooling1D(pool_size=3, strides=1))(model)
model2 = Conv1D(64, kernel_size=4,
strides=1,
activation='relu',
input_shape=input_shape,
padding='same',
name='conv3')(inputs)
model2 = MaxPooling1D(pool_size=3, strides=1)(model2)
model3 = Conv1D(64, kernel_size=5,
strides=1,
activation='relu',
input_shape=input_shape,
padding='same',
name='conv4')(inputs)
model3 = MaxPooling1D(pool_size=3, strides=1)(model3)
models = [model, model2, model3]
full_model = keras.layers.concatenate(models)
full_model = Flatten()(full_model)
full_model = Dense(128, activation='relu')(full_model)
full_model = Dropout(DROPOUT)(full_model)
full_model = Dense(NUM_CLASSES, activation='softmax')(full_model)
full_model = keras.models.Model(input=inputs, outputs=full_model)
full_model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adam(lr=.0005),
metrics=['accuracy'])
early_stopping = EarlyStopping(monitor='val_loss', patience=2)
full_model.fit(x_train, y_train_classes,
batch_size=BATCH_SIZE,
epochs=EPOCHS,
verbose=1,
class_weight=class_weight,
callbacks=[early_stopping],
validation_data=(x_val, y_val_classes))
# -
test_preds_cnn = full_model.predict(x_test, batch_size=BATCH_SIZE)
print(roc_auc_score(y_test_classes, test_preds_cnn, average=None))
print(roc_auc_score(y_test_classes, test_preds_cnn, average='macro'))
print(roc_auc_score(y_test_classes, test_preds_cnn, average='micro'))
# # LSTM
import tensorflow as tf
import functools
BATCH_SIZE = 128
EPOCHS = 12
KEEP_PROB = 0.8
REGULARIZATION = 0.001
NUM_HIDDEN = [512, 512]
# +
def lazy_property(function):
attribute = '_' + function.__name__
@property
@functools.wraps(function)
def wrapper(self):
if not hasattr(self, attribute):
setattr(self, attribute, function(self))
return getattr(self, attribute)
return wrapper
class VariableSequenceLabelling:
def __init__(self, data, target, dropout_prob, reg, num_hidden=[256], class_weights=[1,1,1,1]):
self.data = data
self.target = target
self.dropout_prob = dropout_prob
self.reg = reg
self._num_hidden = num_hidden
self._num_layers = len(num_hidden)
self.num_classes = len(class_weights)
self.attn_length = 0
self.class_weights = class_weights
self.prediction
self.error
self.optimize
@lazy_property
def make_rnn_cell(self,
attn_length=0,
base_cell=tf.nn.rnn_cell.BasicLSTMCell,
state_is_tuple=True):
attn_length = self.attn_length
input_dropout = self.dropout_prob
output_dropout = self.dropout_prob
cells = []
for num_units in self._num_hidden:
cell = base_cell(num_units, state_is_tuple=state_is_tuple)
cell = tf.nn.rnn_cell.DropoutWrapper(cell, input_keep_prob=input_dropout, output_keep_prob=output_dropout)
cells.append(cell)
cell = tf.nn.rnn_cell.MultiRNNCell(cells, state_is_tuple=state_is_tuple)
if attn_length > 0:
sys.path.insert(0, 'attention')
import attention_cell_wrapper_single
cell = attention_cell_wrapper_single.AttentionCellWrapper(
cell, attn_length, input_size=int(self.data.get_shape().as_list()[2]), state_is_tuple=state_is_tuple)
print cell
return cell
# predictor for slices
@lazy_property
def prediction(self):
cell = self.make_rnn_cell
# Recurrent network.
output, final_state = tf.nn.dynamic_rnn(cell,
self.data,
dtype=tf.float32
)
with tf.variable_scope("model") as scope:
tf.get_variable_scope().reuse_variables()
# final weights
num_classes = self.num_classes
weight, bias = self._weight_and_bias(self._num_hidden[-1], num_classes)
# flatten + sigmoid
if self.attn_length > 0:
logits = tf.matmul(final_state[0][-1][-1], weight) + bias
else:
logits = tf.matmul(final_state[-1][-1], weight) + bias
prediction = tf.nn.softmax(logits)
return logits, prediction
@lazy_property
def cross_ent(self):
predictions = self.prediction[0]
real = tf.cast(tf.squeeze(self.target), tf.int32)
class_weight = tf.expand_dims(tf.cast(self.class_weights, tf.int32), axis=0)
print("class_weights", class_weight)
one_hot_labels = tf.cast(tf.one_hot(real, depth=self.num_classes), tf.int32)
weight_per_label = tf.cast(tf.transpose(tf.matmul(one_hot_labels, tf.transpose(class_weight))), tf.float32) #shape [1, batch_size]
xent = tf.multiply(weight_per_label, tf.nn.sparse_softmax_cross_entropy_with_logits(labels=real, logits=predictions, name="xent_raw")) #shape [1, batch_size]
loss = tf.reduce_mean(xent) #shape 1
ce = loss
l2 = self.reg * sum(tf.nn.l2_loss(tf_var) for tf_var in tf.trainable_variables())
ce += l2
return ce
@lazy_property
def optimize(self):
learning_rate = 0.0003
optimizer = tf.train.AdamOptimizer(learning_rate)
return optimizer.minimize(self.cross_ent)
@lazy_property
def error(self):
prediction = tf.argmax(self.prediction[1], 1)
real = tf.cast(self.target, tf.int32)
prediction = tf.cast(prediction, tf.int32)
mistakes = tf.not_equal(real, prediction)
mistakes = tf.cast(mistakes, tf.float32)
mistakes = tf.reduce_sum(mistakes, reduction_indices=0)
total = 128
mistakes = tf.divide(mistakes, tf.to_float(total))
return mistakes
@staticmethod
def _weight_and_bias(in_size, out_size):
weight = tf.truncated_normal([in_size, out_size], stddev=0.01)
bias = tf.constant(0.1, shape=[out_size])
return tf.Variable(weight), tf.Variable(bias)
@lazy_property
def summaries(self):
tf.summary.scalar('loss', tf.reduce_mean(self.cross_ent))
tf.summary.scalar('error', self.error)
merged = tf.summary.merge_all()
return merged
# +
tf.reset_default_graph()
config = tf.ConfigProto(allow_soft_placement = True)
# if attn_length > 0:
# # weights file initialized
# weight_file = 'weights.txt'
# with open(weight_file, 'a') as the_file:
# pass
with tf.Session(config = config) as sess, tf.device('/cpu:0'):
_, length, num_features = x_train.shape
num_data_cols = num_features
print "num features", num_features
print "num_data cols", num_data_cols
# placeholders
data = tf.placeholder(tf.float32, [None, length, num_data_cols])
target = tf.placeholder(tf.float32, [None])
dropout_prob = tf.placeholder(tf.float32)
reg = tf.placeholder(tf.float32)
# initialization
model = VariableSequenceLabelling(data, target, dropout_prob, reg, num_hidden=NUM_HIDDEN, class_weights=class_weight)
sess.run(tf.global_variables_initializer())
print('Initialized Variables...')
batch_size = BATCH_SIZE
dp = KEEP_PROB
rp = REGULARIZATION
train_samples = x_train.shape[0]
indices = range(train_samples)
num_classes = NUM_CLASSES
# for storing results
test_data = x_test
val_data = x_val
val_aucs = []
test_aucs = []
val_aucs_macro = []
test_aucs_macro = []
epoch = -1
print('Beginning Training...')
while (epoch < 3 or max(np.diff(early_stop[-3:])) > 0):
epoch += 1
np.random.shuffle(indices)
num_batches = train_samples/batch_size
for batch_index in range(num_batches):
sample_indices = indices[batch_index*batch_size:batch_index*batch_size+batch_size]
batch_data = x_train[sample_indices, :, :num_data_cols]
batch_target = y_train[sample_indices]
_, loss = sess.run([model.optimize, model.cross_ent], {data: batch_data, target: batch_target, dropout_prob: dp, reg: rp})
# write train accuracy to log files every 10 batches
#if batch_index % 2000 == 0:
# loss, prediction, error = sess.run([model.cross_ent, model.prediction, model.error], {data: batch_data, target: batch_target, dropout_prob: dp, reg: rp})
# #train_writer.add_summary(summaries, global_step=epoch*batch_index)
# print('Epoch {:2d} Batch {:2d}'.format(epoch+1, batch_index))
# print('Loss = ', np.mean(loss))
# print('Error = ', error)
cur_val_preds = sess.run(model.prediction, {data: x_val, target: y_val, dropout_prob: 1, reg: rp})
val_preds = cur_val_preds[1]
cur_test_preds = sess.run(model.prediction, {data: x_test, target: y_test, dropout_prob: 1, reg: rp})
test_preds = cur_test_preds[1]
val_auc_macro = roc_auc_score(y_val_classes, val_preds, average='macro')
test_auc_macro = roc_auc_score(y_test_classes, test_preds, average='macro')
val_aucs_macro.append(val_auc_macro)
test_aucs_macro.append(test_auc_macro)
val_auc = roc_auc_score(y_val_classes, val_preds, average=None)
test_auc = roc_auc_score(y_test_classes, test_preds, average=None)
val_aucs.append(val_auc)
test_aucs.append(test_auc)
if isinstance(val_aucs_macro[-1], dict):
early_stop = [val_auc_macro for val_auc_macro in val_aucs_macro]
else:
early_stop = val_aucs_macro
print "Val AUC = ", val_auc
print "Test AUC = ", test_auc
if isinstance(val_aucs_macro[-1], dict):
best_epoch = np.argmax(np.array([val_auc_macro for val_auc_macro in val_aucs_macro]))
else:
best_epoch = np.argmax(val_aucs_macro)
best_val_auc = val_aucs[best_epoch]
best_test_auc = test_aucs[best_epoch]
best_test_auc_macro = test_aucs_macro[best_epoch]
print 'Best Test AUC: ', best_test_auc, 'at epoch ', best_epoch
print 'Best Test AUC Macro: ', best_test_auc_macro, 'at epoch ', best_epoch
| notebooks/Baselines for Intervention Prediction - Mechanical Ventilation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Weekend Movie Trip
#
# <NAME> (2762306)
# ## MovieLens Datasets
#
# MovieLens Latest-Small Dataset
# http://files.grouplens.org/datasets/movielens/ml-latest-small.zip
import pandas as pd
import numpy as np
import datetime as dt
import seaborn as sns
import matplotlib.pyplot as plt
import math
from statistics import mean, stdev
from sklearn.cluster import KMeans
from sklearn.decomposition import PCA
# ## Read in the Processed Data
df = pd.read_csv("../data/processed/movies_processed.csv")
df.head()
# ### Need to clean up years column some, seem to be getting some errros
df = df.drop(['Unnamed: 0'], axis=1)
df.head()
unique_years = df.Year.unique()
print(unique_years)
for index, row in df.iterrows():
if row['Year'] < 1800:
df.drop(index, inplace=True)
# ## Encountered an error with the way I was previously extracting years from the title row, have fixed in processing notebook
# Dropping NaNs from Years
df = df.dropna()
# ### Struggling to apply k-means with genres in current form, will one-hot encode them
# +
#one_hot = df.GenreCodes.str.get_dummies()
#df = df.join(one_hot)
#df.head()
# INSTEAD OF DOING ONE-HOT HERE, WENT BACK AND ADDED IT TO PROCESSING NOTEBOOK
# -
# ## Applying K-Means
# +
# Using https://medium.com/hanman/data-clustering-what-type-of-movies-are-in-the-imdb-top-250-7ef59372a93b
#for inspiration
kmeans_model = KMeans(20)
# Need to drop all non-numeric data to perform fitting
kmeans_model.fit(df.drop(['movieId', 'title', 'Genres', 'GenreCodes'], axis=1))
clust_labels = kmeans_model.predict(df.drop(['movieId', 'title', 'Genres', 'GenreCodes'], axis=1))
cent = kmeans_model.cluster_centers_
# -
kmeans = pd.DataFrame(clust_labels)
# +
fig = plt.figure(figsize=[25,10])
ax = fig.add_subplot(111)
scatter = ax.scatter(df['AvRate'],df['Year'], c=kmeans[0], s=50)
plt.xticks(rotation='vertical')
plt.colorbar(scatter)
# -
# ## CONCLUSIONS
# There appears to be a fairly even distribution of rating scores across movies released over many years. While there are certainly fewer movies in this dataset from the earlier years, it doesn't seem that there is any significance over whether users felt that a given movie was deserving of a high score or not. Using this clustering methodology, we can recommend to users that liked a certain movie other movies near that same rating. However, we can also extend this methodology to inclusion of genre data as well. Say a user rated a Romantic-Comedy movie very highly, we could recommend a similarly rated Romantic-Comedy to that user in the hopes that it would be enjoyable for them as well.
| notebooks/02-weekend_movie_trip.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
#
# # Predicting Loan Repayment with Automated Feature Engineering in Featuretools
#
# Feature engineering is the process of creating new features (also called predictors or explanatory variables) out of an existing dataset. Traditionally, this process is done by hand using domain knowledge to build new features one at a time. Feature engineering is crucial for a data science problem and a manual approach is time-consuming, tedious, error-prone, and must be re-done for each problem. Automated feature engineering aims to aid the data scientist in this critical process by automatically creating hundreds or thousands of new features from a set of related tables in a fraction of the time as the manual approach. In this notebook, we will apply automated feature engineering to the Home Credit Default Risk loan dataset using [Featuretools, an open-source Python library](https://www.featuretools.com/) for automated feature engineering.
#
# This problem is a machine learning competition on Kaggle where the objective is to predict if an applicant will default on a loan given comprehensive data on past loans and applicants. The data is spread across seven different tables making this an ideal problem for automated feature engineering: all of the data must be gathered into a single dataframe for training (and one for testing) with the aim of capturing as much usable information for the prediction problem as possible. As we will see, featuretools can efficiently carry out the tedious process of using all of these tables to make new features with only a few lines of code. Moreover, this code is generally applicable to any data science problem!
#
# The general idea of automated feature engineering is picture below:
#
# 
#
#
# ## Approach
#
# In this notebook, we will implement an automated feature engineering approach to the loan repayment problem. While Featuretools allows plenty of options for customization of the library to improve accuracy, we'll focus on a fairly high-level implementation.
#
# Our approach will be as follows with the background covered as we go:
#
# 1. Read in the set of related data tables
# 2. Create a featuretools `EntitySet` and add `entities` to it
# * Identify correct variable types as required
# * Identify indices in data
# 3. Add relationships between `entities`
# 4. Select feature primitives to use to create new features
# * Use basic set of primitives
# * Examine features that will be created
# 5. Run Deep Feature Synthesis to generate thousands of new features
#
#
# ## Problem and Dataset
#
# The [Home Credit Default Risk competition](https://www.kaggle.com/c/home-credit-default-risk) currently running on Kaggle is a supervised classification task where the objective is to predict whether or not an applicant for a loan (known as a client) will default on the loan. The data comprises socio-economic indicators for the clients, loan specific financial information, and comprehensive data on previous loans at Home Credit (the institution sponsoring the competition) and other credit agencies. The metric for this competition is Receiver Operating Characteristic Area Under the Curve (ROC AUC) with predictions made in terms of the probability of default. We can evaluate our submissions both through cross-validation on the training data (for which we have the labels) or by submitting our test predictions to Kaggle to see where we place on the public leaderboard (which is calculated with only 10% of the testing data).
#
# The Home Credit Default Risk dataset ([available for download here](https://www.kaggle.com/c/home-credit-default-risk/data)) consists of seven related tables of data:
#
# * application_train/application_test: the main training/testing data for each client at Home Credit. The information includes both socioeconomic indicators for the client and loan-specific characteristics. Each loan has its own row and is uniquely identified by the feature `SK_ID_CURR`. The training application data comes with the `TARGET` indicating 0: the loan was repaid or 1: the loan was not repaid.
# * bureau: data concerning client's previous credits from other financial institutions (not Home Credit). Each previous credit has its own row in bureau, but one client in the application data can have multiple previous credits. The previous credits are uniquely identified by the feature `SK_ID_BUREAU`.
# * bureau_balance: monthly balance data about the credits in bureau. Each row has information for one month about a previous credit and a single previous credit can have multiple rows. This is linked backed to the bureau loan data by `SK_ID_BUREAU` (not unique in this dataframe).
# * previous_application: previous applications for loans at Home Credit of clients who have loans in the application data. Each client in the application data can have multiple previous loans. Each previous application has one row in this dataframe and is uniquely identified by the feature `SK_ID_PREV`.
# * POS_CASH_BALANCE: monthly data about previous point of sale or cash loans from the previous loan data. Each row is one month of a previous point of sale or cash loan, and a single previous loan can have many rows. This is linked backed to the previous loan data by `SK_ID_PREV` (not unique in this dataframe).
# * credit_card_balance: monthly data about previous credit cards loans from the previous loan data. Each row is one month of a credit card balance, and a single credit card can have many rows. This is linked backed to the previous loan data by `SK_ID_PREV` (not unique in this dataframe).
# * installments_payment: payment history for previous loans at Home Credit. There is one row for every made payment and one row for every missed payment. This is linked backed to the previous loan data by `SK_ID_PREV` (not unique in this dataframe).
#
# The image below shows the seven tables and the variables linking them:
#
# 
#
# The variables that tie the tables together will be important to understand when it comes to adding `relationships` between entities. __The only domain knowledge we need for a full Featuretools approach to the problem is the indexes of the tables and the relationships between the tables.__
# +
# pandas and numpy for data manipulation
import pandas as pd
import numpy as np
# featuretools for automated feature engineering
import featuretools as ft
# -
# ### Read in Data
#
# First we can read in the seven data tables. We also replace the anomalous values previously identified (we did the same process with manual feature engineering).
# Read in the datasets and replace the anomalous values
app_train = pd.read_csv('input/application_train.csv').replace({365243: np.nan})
app_test = pd.read_csv('input/application_test.csv').replace({365243: np.nan})
bureau = pd.read_csv('input/bureau.csv').replace({365243: np.nan})
bureau_balance = pd.read_csv('input/bureau_balance.csv').replace({365243: np.nan})
cash = pd.read_csv('input/POS_CASH_balance.csv').replace({365243: np.nan})
credit = pd.read_csv('input/credit_card_balance.csv').replace({365243: np.nan})
previous = pd.read_csv('input/previous_application.csv').replace({365243: np.nan})
installments = pd.read_csv('input/installments_payments.csv').replace({365243: np.nan})
# We will join together the training and testing datasets to make sure we build the same features for each set. Later, after the feature matrix is built, we can separate out the two sets.
# +
app_test['TARGET'] = np.nan
# Join together training and testing
app = app_train.append(app_test, ignore_index = True, sort = True)
# -
# Several of the indexes are an incorrect data type (floats) so we need to make these all the same (integers) for adding relationships.
for index in ['SK_ID_CURR', 'SK_ID_PREV', 'SK_ID_BUREAU']:
for dataset in [app, bureau, bureau_balance, cash, credit, previous, installments]:
if index in list(dataset.columns):
dataset[index] = dataset[index].fillna(0).astype(np.int64)
# # Featuretools Basics
#
# [Featuretools](https://docs.featuretools.com/#minute-quick-start) is an open-source Python library for automatically creating features out of a set of related tables using a technique called [Deep Feature Synthesis](http://www.jmaxkanter.com/static/papers/DSAA_DSM_2015.pdf). Automated feature engineering, like many topics in machine learning, is a complex subject built upon a foundation of simpler ideas. By going through these ideas one at a time, we can build up our understanding of Featuretools which will later allow for us to get the most out of it.
#
# There are a few concepts that we will cover along the way:
#
# * [Entities and EntitySets](https://docs.featuretools.com/en/stable/loading_data/using_entitysets.html): our tables and a data structure for keeping track of them all
# * [Relationships between tables](https://docs.featuretools.com/en/stable/loading_data/using_entitysets.html#adding-a-relationship): how the tables can be related to one another
# * [Feature primitives](https://docs.featuretools.com/en/stable/automated_feature_engineering/primitives.html): aggregations and transformations that are stacked to build features
# * [Deep feature synthesis](https://docs.featuretools.com/en/stable/automated_feature_engineering/afe.html): the method that uses feature primitives to generate thousands of new features
#
# # Entities and Entitysets
#
# An entity is simply a table or in Pandas, a `dataframe`. The observations must be in the rows and the features in the columns. An entity in featuretools must have a unique index where none of the elements are duplicated. Currently, only `app`, `bureau`, and `previous` have unique indices (`SK_ID_CURR`, `SK_ID_BUREAU`, and `SK_ID_PREV` respectively). For the other dataframes, when we create entities from them, we must pass in `make_index = True` and then specify the name of the index.
#
# Entities can also have time indices that represent when the information in the row became known. (There are not datetimes in any of the data, but there are relative times, given in months or days, that could be treated as time variables, although we will not use them as time in this notebook).
#
# An [EntitySet](https://docs.featuretools.com/en/stable/loading_data/using_entitysets.html) is a collection of tables and the relationships between them. This can be thought of a data structure with its own methods and attributes. Using an EntitySet allows us to group together multiple tables and will make creating the features much simpler than keeping track of individual tables and relationships. __EntitySets and entities are abstractions that can be applied to any dataset because they do not depend on the underlying data.__
#
# First we'll make an empty entityset named clients to keep track of all the data.
# Entity set with id applications
es = ft.EntitySet(id = 'clients')
# ### Variable Types
#
# Featuretools will automatically infer the variable types. However, there may be some cases where we need to explicitly tell featuretools the variable type such as when a boolean variable is represented as an integer. Variable types in featuretools can be specified as a dictionary.
#
# We will first work with the `app` data to specify the proper variable types. To identify the `Boolean` variables that are recorded as numbers (1.0 or 0.0), we can iterate through the data and find any columns where there are only 2 unique values and the data type is numeric. We can also use the column definitions to find any other data types that should be identified, such as `Ordinal` variables. Identifying the correct variable types is important because Featuretools applies different operations to different data types (just as we do when manual feature engineering).
import featuretools.variable_types as vtypes
# +
app_types = {}
# Handle the Boolean variables:
for col in app:
if (app[col].nunique() == 2) and (app[col].dtype == float):
app_types[col] = vtypes.Boolean
# Remove the `TARGET`
del app_types['TARGET']
print('There are {} Boolean variables in the application data.'.format(len(app_types)))
# -
# Ordinal variables
app_types['REGION_RATING_CLIENT'] = vtypes.Ordinal
app_types['REGION_RATING_CLIENT_W_CITY'] = vtypes.Ordinal
app_types['HOUR_APPR_PROCESS_START'] = vtypes.Ordinal
# The `previous` table is the only other `entity` that has features which should be recorded as Boolean. Correctly identifying the type of column will prevent featuretools from making irrelevant features such as the mean or max of a `Boolean`.
# +
previous_types = {}
# Handle the Boolean variables:
for col in previous:
if (previous[col].nunique() == 2) and (previous[col].dtype == float):
previous_types[col] = vtypes.Boolean
print('There are {} Boolean variables in the previous data.'.format(len(previous_types)))
# -
# In addition to identifying Boolean variables, we want to make sure featuretools does not create nonsense features such as statistical aggregations (mean, max, etc.) of ids. The `credit`, `cash`, and `installments` data all have the `SK_ID_CURR` variable. However, we do not actually need this variable in these dataframes because we link them to `app` through the `previous` dataframe with the `SK_ID_PREV` variable.
#
# We don't want to make features from `SK_ID_CURR` since it is an arbitrary id and should have no predictive power.
# Our options to handle these variables is either to tell featuretools to ignore them, or to drop the features before including them in the entityset. We will take the latter approach.
installments = installments.drop(columns = ['SK_ID_CURR'])
credit = credit.drop(columns = ['SK_ID_CURR'])
cash = cash.drop(columns = ['SK_ID_CURR'])
# ## Adding Entities
#
# Now we define each entity, or table of data, and add it to the `EntitySet`. We need to pass in an index if the table has one or `make_index = True` if not. In the cases where we need to make an index, we must supply a name for the index. We also need to pass in the dictionary of variable types if there are any specific variables we should identify. The following code adds all seven tables to the `EntitySet`.
# +
# Entities with a unique index
es = es.entity_from_dataframe(entity_id = 'app', dataframe = app, index = 'SK_ID_CURR',
variable_types = app_types)
es = es.entity_from_dataframe(entity_id = 'bureau', dataframe = bureau, index = 'SK_ID_BUREAU')
es = es.entity_from_dataframe(entity_id = 'previous', dataframe = previous, index = 'SK_ID_PREV',
variable_types = previous_types)
# Entities that do not have a unique index
es = es.entity_from_dataframe(entity_id = 'bureau_balance', dataframe = bureau_balance,
make_index = True, index = 'bureaubalance_index')
es = es.entity_from_dataframe(entity_id = 'cash', dataframe = cash,
make_index = True, index = 'cash_index')
es = es.entity_from_dataframe(entity_id = 'installments', dataframe = installments,
make_index = True, index = 'installments_index')
es = es.entity_from_dataframe(entity_id = 'credit', dataframe = credit,
make_index = True, index = 'credit_index')
# -
# Display entityset so far
es
# The `EntitySet` allows us to group together all of our tables as one data structure. This is much easier than manipulating the tables one at a time (as we have to do in manual feature engineering).
# # Relationships
#
# Relationships are a fundamental concept not only in featuretools, but in any relational database. The most common type of relationship is one-to-many. The best way to think of a one-to-many relationship is with the analogy of parent-to-child. A parent is a single individual, but can have mutliple children. In the context of tables, a parent table will have one row (observation) for every individual while a child table can have many observations for each parent. In a _parent table_, each individual has a single row and is uniquely identified by an index (also called a key). Each individual in the parent table can have multiple rows in the _child table_. Things get a little more complicated because children tables can have children of their own, making these grandchildren of the original parent.
#
# As an example of a parent-to-child relationship, the `app` dataframe has one row for each client (identified by `SK_ID_CURR`) while the `bureau` dataframe has multiple previous loans for each client. Therefore, the `bureau` dataframe is the child of the `app` dataframe. The `bureau` dataframe in turn is the parent of `bureau_balance` because each loan has one row in `bureau` (identified by `SK_ID_BUREAU`) but multiple monthly records in `bureau_balance`. When we do manual feature engineering, keeping track of all these relationships is a massive time investment (and a potential source of error), but we can add these relationships to our `EntitySet` and let featuretools worry about keeping the tables straight!
print('Parent: app, Parent Variable of bureau: SK_ID_CURR\n\n', app.iloc[:, 111:115].head())
print('\nChild: bureau, Child Variable of app: SK_ID_CURR\n\n', bureau.iloc[:, :5].head())
# The `SK_ID_CURR` 215354 has one row in the parent table and multiple rows in the child.
#
# Two tables are linked via a shared variable. The `app` and `bureau` dataframe are linked by the `SK_ID_CURR` variable while the `bureau` and `bureau_balance` dataframes are linked with the `SK_ID_BUREAU`. The linking variable is called the `parent` variable in the parent table and the `child` variable in the child table.
print('Parent: bureau, Parent Variable of bureau_balance: SK_ID_BUREAU\n\n', bureau.iloc[:, :5].head())
print('\nChild: bureau_balance, Child Variable of bureau: SK_ID_BUREAU\n\n', bureau_balance.head())
# Traditionally, we use the relationships between parents and children to aggregate data by grouping together all the children for a single parent and calculating statistics. For example, we might group together all the loans for a single client and calculate the average loan amount. This is straightforward, but can grow extremely tedious when we want to make hundreds of these features. Doing so one at a time is extremely inefficient especially because we end up re-writing much of the code over and over again and this code cannot be used for any different problem!
#
# Things get even worse when we have to aggregate the grandchildren because we have to use two steps: first aggregate at the parent level, and then at the grandparent level. Soon we will see that Featuretools can do this work automatically for us, generating thousands of features from __all__ of the data tables. When we did this manually it took about 15 minutes per feature so Featuretools potentially saves us hundreds of hours.
#
# ### Adding Relationships
#
# Defining the relationships is straightforward using the diagram for the data tables. For each relationship, we need to first specify the parent variable and then the child variable. Altogether, there are a total of 6 relationships between the tables (counting the training and testing relationships as one). Below we specify these relationships and then add them to the EntitySet.
# +
# Relationship between app_train and bureau
r_app_bureau = ft.Relationship(es['app']['SK_ID_CURR'], es['bureau']['SK_ID_CURR'])
# Relationship between bureau and bureau balance
r_bureau_balance = ft.Relationship(es['bureau']['SK_ID_BUREAU'], es['bureau_balance']['SK_ID_BUREAU'])
# Relationship between current app and previous apps
r_app_previous = ft.Relationship(es['app']['SK_ID_CURR'], es['previous']['SK_ID_CURR'])
# Relationships between previous apps and cash, installments, and credit
r_previous_cash = ft.Relationship(es['previous']['SK_ID_PREV'], es['cash']['SK_ID_PREV'])
r_previous_installments = ft.Relationship(es['previous']['SK_ID_PREV'], es['installments']['SK_ID_PREV'])
r_previous_credit = ft.Relationship(es['previous']['SK_ID_PREV'], es['credit']['SK_ID_PREV'])
# -
# Add in the defined relationships
es = es.add_relationships([r_app_bureau, r_bureau_balance, r_app_previous,
r_previous_cash, r_previous_installments, r_previous_credit])
# Print out the EntitySet
es
# Again, we can see the benefits of using an `EntitySet` that is able to track all of the relationships for us. This allows us to work at a higher level of abstraction, thinking about the entire dataset rather than each individual table, greatly increasing our efficiency.
# __Slightly advanced note__: we need to be careful to not create a [diamond graph](https://en.wikipedia.org/wiki/Diamond_graph) where there are multiple paths from a parent to a child. If we directly link `app` and `cash` via `SK_ID_CURR`; `previous` and `cash` via `SK_ID_PREV`; and `app` and `previous` via `SK_ID_CURR`, then we have created two paths from `app` to `cash`. This results in ambiguity, so the approach we have to take instead is to link `app` to `cash` through `previous`. We establish a relationship between `previous` (the parent) and `cash` (the child) using `SK_ID_PREV`. Then we establish a relationship between `app` (the parent) and `previous` (now the child) using `SK_ID_CURR`. Then featuretools will be able to create features on `app` derived from both `previous` and `cash` by stacking multiple primitives.
#
# If this doesn't make too much sense, then just remember to only include one path from a parent to any descendents. For example, link a grandparent to a grandchild through the parent instead of directly through a shared variable.
# All entities in the entity can be linked through these relationships. In theory this allows us to calculate features for any of the entities, but in practice, we will only calculate features for the `app` dataframe since that will be used for training/testing. The end outcome will be a dataframe that has one row for each client in `app` with thousands of features for each individual.
#
# We are almost to the point where we can start creating thousands of features but we still have a few foundational topics to understand. The next building block to cover is feature primitives.
# ## Visualize EntitySet
es.plot()
# # Feature Primitives
#
# A [feature primitive](https://docs.featuretools.com/en/stable/automated_feature_engineering/primitives.html) is an operation applied to a table or a set of tables to create a feature. These represent simple calculations, many of which we already use in manual feature engineering, that can be stacked on top of each other to create complex deep features. Feature primitives fall into two categories:
#
# * __Aggregation__: function that groups together children for each parent and calculates a statistic such as mean, min, max, or standard deviation across the children. An example is the maximum previous loan amount for each client. An aggregation covers multiple tables using relationships between tables.
# * __Transformation__: an operation applied to one or more columns in a single table. An example would be taking the absolute value of a column, or finding the difference between two columns in one table.
#
# A list of the available features primitives in featuretools can be viewed below.
# +
# List the primitives in a dataframe
primitives = ft.list_primitives()
pd.options.display.max_colwidth = 100
primitives[primitives['type'] == 'aggregation'].head(10)
# -
primitives[primitives['type'] == 'transform'].head(10)
# # Deep Feature Synthesis
#
# [Deep Feature Synthesis (DFS)](https://docs.featuretools.com/en/stable/automated_feature_engineering/afe.html) is the method Featuretools uses to make new features. DFS stacks feature primitives to form features with a "depth" equal to the number of primitives. For example, if we take the maximum value of a client's previous loans (say `MAX(previous.loan_amount)`), that is a "deep feature" with a depth of 1. To create a feature with a depth of two, we could stack primitives by taking the maximum value of a client's average monthly payments per previous loan (such as `MAX(previous(MEAN(installments.payment)))`). In manual feature engineering, this would require two separate groupings and aggregations and took more than 15 minutes to write the code per feature.
#
# Deep Feature Synthesis is an extremely powerful method that allows us to overcome our human limitations on time and creativity by building features that we would never be able to think of on our own (or would not have the patience to implement). Furthermore, DFS is applicable to any dataset with only very minor changes in syntax. In feature engineering, we generally apply the same functions to multiple datasets, but when we do it by hand, we have to re-write the code because it is problem-specific. Featuretools code can be applied to any dataset because it is written at a higher level of abstraction.
#
# The [original paper on automated feature engineering using Deep Feature Synthesis](https://dai.lids.mit.edu/wp-content/uploads/2017/10/DSAA_DSM_2015.pdf) is worth a read if you want to understand the concepts at a deeper level.
#
# To perform DFS in featuretools, we use the `dfs` function passing it an `entityset`, the `target_entity` (where we want to make the features), the `agg_primitives` to use, the `trans_primitives` to use, the `max_depth` of the features, and a number of other arguments depending on our use case. There are also options for multi-processing with `njobs` and the information that is printed out with `verbose`.
#
# One other important argument is __`features_only`__. If we set this to `True`, `dfs` will only make the feature names and not calculate the actual values of the features (called the feature matrix). This is useful when we want to inspect the feature that will be created and we can also save the features to use with a different dataset (for example when we have training and testing data).
# ## Deep Feature Synthesis with Default Primitives
#
# Without using any domain knowledge we can make thousands of features by using the default primitives in featuretools. This first call will use the default aggregation and transformation primitives, a max depth of 2, and calculate primitives for the `app` entity. We will only generate the features themselves (the names and not the values) which we can save and inspect.
# +
# Default primitives from featuretools
default_agg_primitives = ["sum", "std", "max", "skew", "min", "mean", "count", "percent_true", "num_unique", "mode"]
default_trans_primitives = ["day", "year", "month", "weekday", "haversine", "num_words", "num_characters"]
# DFS with specified primitives
feature_names = ft.dfs(entityset = es, target_entity = 'app',
trans_primitives = default_trans_primitives,
agg_primitives=default_agg_primitives,
where_primitives = [], seed_features = [],
max_depth = 2, n_jobs = -1, verbose = 1,
features_only=True)
# -
# Even a basic call to deep feature synthesis gives us over 2000 features to work with. Granted, not all of these will be important, but this still represents hundreds of hours that we saved. Moreover, `dfs` might be able to find important features that we would never have thought of in the first place.
#
# We can look at the some of the feature names:
feature_names[1000:1020]
# Notice how featuretools stacks multiple primitives on top of each other. This one of the ideas behind Deep Feature Synthesis and automated feature engineering. Rather than having to do these groupings and aggregations by ourselves, Featuretools is able to handle it all using the framework (`entities`, `relationships`, and `primitives`) that we provide. We can also use Featuretools to expand on our domain knowledge.
# # Building on Top of Domain Features
#
# Featuretools will automatically build thousands of features for us, but that does not mean we can't use our own knowledge to improve the predictive performance. Featuretools is able to augment our domain knowledge by stacking additional features on top of our domain knowledge based features. We identified and created numerous useful features in the manual feature engineering notebook, based on our own knowledge and that of thousands of data scientists working on this problem on Kaggle. Rather than getting only one domain knowledge feature, we can effectively get dozens or even hundreds. __Here we'll explain the options for using domain knowledge, but we'll stick with the simple implementation of Featuretools for comparison purposes.__
#
# For more information on any of these topics, see the [documentation](https://docs.featuretools.com/en/stable/guides/tuning_dfs.html) or the other notebooks in this repository.
#
# ### Seed Features
#
# Seed features are domain features that we make in the data that Featuretools is then able to build on top of. For example, we saw that the rate of a loan is an important feature because a higher rate loan is likely more risky. In Featuretools, we can encode the loan rate (both for the current loan and for previous loans) as a seed feature and Featuretools will build additional explanatory variables on this domain knowledge wherever possible.
#
# ### Interesting Values
#
# Interesting values have a similar idea to seed features except they allow us to make conditional features. For example, we might want to find for each client the mean amount of previous loans that have been closed and the mean amount of previous loans that are still active. By specifying interesting values in `bureau` on the `CREDIT_ACTIVE` variable we can have Featuretools do exactly that! Carrying this out by hand would be extremely tedious and present numerous opportunities for errors.
#
# ### Custom Primitives
#
# If we aren't satisfied with the primitives available to use in Featuretools, we can write our own functions to transform or aggregate the data. This is one of the most powerful capabilities in featuretools because it allows us to make very specific operations that can then be applied to multiple datasets.
#
# __In this notebook we concentrate on a basic implementation of Featuretools, but keep in mind these capabilities are available for optimizing the library and using domain knowledge!__
# # Selecting Primitives
#
# For our actual set of features, we will use a select group of primitives rather than just the defaults. This will generate over 2100 features to use for modeling.
# Specify primitives
agg_primitives = ["sum", "max", "min", "mean", "count", "percent_true", "num_unique", "mode"]
trans_primitives = ['percentile', 'and']
# Deep feature synthesis
feature_names = ft.dfs(entityset=es, target_entity='app',
agg_primitives = agg_primitives,
trans_primitives = trans_primitives,
n_jobs = -1, verbose = 1,
features_only = True,
max_depth = 2)
ft.save_features(feature_names, 'input/features.txt')
# If we save the features, we can then use them with `calculate_feature_matrix`. This is useful when we want to apply the same features across datasets (such as if we have separate trainig/testing).
# ## Run Full Deep Feature Synthesis
#
# If we are content with the features that will be built, we can run deep feature synthesis and create the feature matrix. The following call runs the full deep feature synthesis. This might take a long time depending on your machine. Featuretools does allow for parallel processing, but each core must be able to handle the entire entityset.
#
# __An actual run of this code was completed using Dask which can be seen in the [Featuretools on Dask notebook](https://github.com/Featuretools/Automated-Manual-Comparison/blob/master/Loan%20Repayment/notebooks/Featuretools%20on%20Dask.ipynb).__ The Dask code takes under 2 hours to run and is a great example of how we can use parallel processing to use our resouces in the most efficient manner.
import sys
print('Total size of entityset: {:.5f} gb.'.format(sys.getsizeof(es) / 1e9))
# +
import psutil
print('Total number of cpus detected: {}.'.format(psutil.cpu_count()))
print('Total size of system memory: {:.5f} gb.'.format(psutil.virtual_memory().total / 1e9))
# -
feature_matrix, feature_names = ft.dfs(entityset=es, target_entity='app',
agg_primitives = agg_primitives,
trans_primitives = trans_primitives,
n_jobs = 1, verbose = 1, features_only = False,
max_depth = 2, chunk_size = 100)
# +
# feature_matrix.reset_index(inplace = True)
# feature_matrix.to_csv('../input/feature_matrix.csv', index = False)
# -
# To download the feature matrix, head to https://www.kaggle.com/willkoehrsen/home-credit-default-risk-feature-tools and select the `feature_matrix_article.csv`. There are several other versions of automatically engineered feature matrices available there as well.
# # Conclusions
#
# In this notebook, we saw how to implement automated feature engineering for a data science problem. __Automated feature engineering allows us to create thousands of new features from a set of related data tables, significantly increasing our efficiency as data scientists.__ Moreover, we can still use domain knowledge in our features and even augment our domain knowledge by building on top of our own hand-built features. The main takeaways are:
#
# * Automated feature engineering took 1 hour to implement compared to 10 hours for manual feature engineering
# * Automated feature engineering built thousands of features in a few lines of code compared to dozens of lines of code per feature for manual engineering.
# * Overall, performance of the automated features are comparable or better than those of the manual features (see the Results notebook)
#
# The benefits of automated feature engineering are significant and will considerably help us in our role as data scientists. It won't alleviate the need for data scientists, but rather will make us more efficient and build better predictive pipelines in less time.
# ## Next Steps
#
# After creating a full set of features, we can apply feature selection and then proceed with modeling. To optimize the model for the features, we use random search for 100 iterations over a grid of hyperparamters. To see how to use Dask to run Featuretools in parallel, refer to the Featuretools Implementation with Dask notebook. For feature selection refer to the Feature Selection notebook. Final results are presented in the Results notebook.
# <p>
# <img src="https://www.featurelabs.com/wp-content/uploads/2017/12/logo.png" alt="Featuretools" />
# </p>
#
# Featuretools was created by the developers at [Feature Labs](https://www.featurelabs.com/). If building impactful data science pipelines is important to you or your business, please [get in touch](https://www.featurelabs.com/contact).
| archived-notebooks/predict-loan-repayment/Automated Loan Repayment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (with sys packages)
# language: python
# name: py3syspck
# ---
import numpy as np
from keras.models import Model
from keras.layers import Input
from keras.layers.convolutional import Cropping2D
from keras import backend as K
def format_decimal(arr, places=6):
return [round(x * 10**places) / 10**places for x in arr]
# ### Cropping2D
# **[convolutional.Cropping2D.0] cropping (1,1),(1, 1) on 3x5x4 input, dim_ordering=tf**
# +
data_in_shape = (3, 5, 4)
L = Cropping2D(cropping=((1,1),(1, 1)), dim_ordering='tf')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(250)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
# **[convolutional.Cropping2D.1] cropping (1,1),(1, 1) on 3x5x4 input, dim_ordering=th**
# +
data_in_shape = (3, 5, 4)
L = Cropping2D(cropping=((1,1),(1, 1)), dim_ordering='th')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(250)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
# **[convolutional.Cropping2D.2] cropping (4,2),(3,1) on 8x7x6 input, dim_ordering=tf**
# +
data_in_shape = (8, 7, 6)
L = Cropping2D(cropping=((4,2),(3,1)), dim_ordering='tf')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(252)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
# **[convolutional.Cropping2D.3] cropping (4,2),(3,1) on 8x7x6 input, dim_ordering=th**
# +
data_in_shape = (8, 7, 6)
L = Cropping2D(cropping=((4,2),(3,1)), dim_ordering='th')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(252)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
| notebooks/layers/convolutional/Cropping2D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <div align="center">Grid Search for model tuning</div>
# ---------------------------------------------------------------------
#
# you can Find me on Github:
# > ###### [ GitHub](https://github.com/lev1khachatryan)
#
# <img src="pics/main.gif" />
# A model hyperparameter is a characteristic of a model that is external to the model and whose value cannot be estimated from data. The value of the hyperparameter has to be set before the learning process begins. For example, c in Support Vector Machines, k in k-Nearest Neighbors, the number of hidden layers in Neural Networks.
#
# In contrast, a parameter is an internal characteristic of the model and its value can be estimated from data. Example, beta coefficients of linear/logistic regression or support vectors in Support Vector Machines.
#
# Grid-search is used to find the optimal hyperparameters of a model which results in the most ‘accurate’ predictions.
#
# Let’s look at Grid-Search by building a classification model on the Breast Cancer dataset.
#
# +
'''
Load packages for working with data
'''
import numpy as np
import pandas as pd
'''
Suppress warnings
'''
import warnings
warnings.filterwarnings('ignore')
# +
#import data
data = pd.read_csv('data/breast-cancer-wisconsin.data',header=None)
#set column names
data.columns = ['Sample Code Number','Clump Thickness','Uniformity of Cell Size',
'Uniformity of Cell Shape','Marginal Adhesion','Single Epithelial Cell Size',
'Bare Nuclei','Bland Chromatin','Normal Nucleoli','Mitoses','Class']
#view top 10 rows
data.head(10)
# -
# Each row in the dataset have one of two possible classes: benign (represented by 2) and malignant (represented by 4). Also, there are 10 attributes in this dataset (shown above) which will be used for prediction, except Sample Code Number which is the id number.
data = data.drop(['Sample Code Number'],axis=1) #Drop 1st column
data = data[data['Bare Nuclei'] != '?'] #Remove rows with missing data
data['Class'] = np.where(data['Class'] ==2,0,1) #Change the Class representation
data['Class'].value_counts() #Class distribution
# Before building a classification model, let’s build a Dummy Classifier to determine the ‘baseline’ performance. This answers the question — ‘What would be the success rate of the model, if one were simply guessing?’ The dummy classifier we are using will simply predict the majority class.
# +
#Split data into attributes and class
X = data.drop(['Class'],axis=1)
y = data['Class']
#perform training and test split
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
#Dummy Classifier
from sklearn.dummy import DummyClassifier
clf = DummyClassifier(strategy= 'most_frequent').fit(X_train,y_train)
y_pred = clf.predict(X_test)
#Distribution of y test
print('y actual : \n' + str(y_test.value_counts()))
#Distribution of y predicted
print('y predicted : \n' + str(pd.Series(y_pred).value_counts()))
# -
# From the output, we can observe that there are 68 malignant and 103 benign cases in the test dataset. However, our classifier predicts all cases as benign (as it is the majority class).
# +
# Model Evaluation metrics
from sklearn.metrics import accuracy_score,recall_score,precision_score,f1_score
print('Accuracy Score : ' + str(accuracy_score(y_test,y_pred)))
print('Precision Score : ' + str(precision_score(y_test,y_pred)))
print('Recall Score : ' + str(recall_score(y_test,y_pred)))
print('F1 Score : ' + str(f1_score(y_test,y_pred)))
#Dummy Classifier Confusion matrix
from sklearn.metrics import confusion_matrix
print('Confusion Matrix : \n' + str(confusion_matrix(y_test,y_pred)))
# -
# The accuracy of the model is 60.2%, but this is a case where accuracy may not be the best metric to evaluate the model. So, let’s take a look at the other evaluation metrics.
#
#
# To summarize the confusion matrix : TRUE POSITIVES (TP)= 0,TRUE NEGATIVES (TN)= 103,FALSE POSITIVES (FP)= 0, FALSE NEGATIVES (FN)= 68. The formulae for the evaluation metrics are as follows :
# <img src="pics/pics.png" />
# Since the model does not classify any malignant case correctly, the recall and precision metrics are 0.
# Now that we have the baseline accuracy, let’s build a Logistic regression model with default parameters and evaluate the model.
# +
#Logistic regression
from sklearn.linear_model import LogisticRegression
clf = LogisticRegression().fit(X_train,y_train)
y_pred = clf.predict(X_test)
# Model Evaluation metrics
from sklearn.metrics import accuracy_score,recall_score,precision_score,f1_score
print('Accuracy Score : ' + str(accuracy_score(y_test,y_pred)))
print('Precision Score : ' + str(precision_score(y_test,y_pred)))
print('Recall Score : ' + str(recall_score(y_test,y_pred)))
print('F1 Score : ' + str(f1_score(y_test,y_pred)))
#Logistic Regression Classifier Confusion matrix
from sklearn.metrics import confusion_matrix
print('Confusion Matrix : \n' + str(confusion_matrix(y_test,y_pred)))
# -
# By fitting the Logistic Regression model with the default parameters, we have a much ‘better’ model. The accuracy is 94.7% and at the same time, the Precision is a staggering 98.3%. Now, let’s take a look at the confusion matrix again for this model results again :
#
#
# Looking at the misclassified instances, we can observe that 8 malignant cases have been classified incorrectly as benign (False negatives). Also, just one benign case has been classified as malignant (False positive).
#
#
# A false negative is more serious as a disease has been ignored, which can lead to the death of the patient. At the same time, a false positive would lead to an unnecessary treatment — incurring additional cost.
# Let’s try to minimize the false negatives by using Grid Search to find the optimal parameters. Grid search can be used to improve any specific evaluation metric.
#
# The metric we need to focus on to reduce false negatives is Recall.
# +
#Grid Search
from sklearn.model_selection import GridSearchCV
clf = LogisticRegression()
grid_values = {'penalty': ['l1', 'l2'],'C':[0.001,.009,0.01,.09,1,5,10,25]}
grid_clf_acc = GridSearchCV(clf, param_grid = grid_values,scoring = 'recall')
grid_clf_acc.fit(X_train, y_train)
#Predict values based on new parameters
y_pred_acc = grid_clf_acc.predict(X_test)
# New Model Evaluation metrics
print('Accuracy Score : ' + str(accuracy_score(y_test,y_pred_acc)))
print('Precision Score : ' + str(precision_score(y_test,y_pred_acc)))
print('Recall Score : ' + str(recall_score(y_test,y_pred_acc)))
print('F1 Score : ' + str(f1_score(y_test,y_pred_acc)))
#Logistic Regression (Grid Search) Confusion matrix
confusion_matrix(y_test,y_pred_acc)
# -
# You can further tune the model to strike a balance between precision and recall by using ‘f1’ score as the evaluation metric
# Grid search builds a model for every combination of hyperparameters specified and evaluates each model. A more efficient technique for hyperparameter tuning is the Randomized search — where random combinations of the hyperparameters are used to find the best solution.
| Lectures/Levon/GridSearch_for_model_tuning/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="l1rVFB-tpvjo"
import numpy as np
import pandas as pd
import nltk
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics import accuracy_score, confusion_matrix, classification_report, plot_roc_curve
import matplotlib.pyplot as plt
#import string
#punc=string.punctuation
# + id="UtThXGaerQgM" colab={"base_uri": "https://localhost:8080/"} outputId="af9d758d-a6b1-4113-e718-40242d410ca4"
from google.colab import drive
drive.mount('/content/drive')
# + id="V39_8W4Ppvj2"
df=pd.read_csv("/content/drive/MyDrive/NLP/train.csv")
df.dropna(inplace=True)
# + id="5I3mcJsctFoX"
df=df.drop(100)
# + id="0Fv3Fjxfpvj5"
df["sex"]=df["sex"].replace("M",1)
df["sex"]=df["sex"].replace("F",0)
# + id="9voiiA38rQc6"
df = df[0:100]
# + colab={"base_uri": "https://localhost:8080/", "height": 418} id="GzWuqzq1rktf" outputId="46b469aa-eb20-43eb-ffd1-9b595aead439"
df
# + [markdown] id="gKc9Atzrpvj6"
# # CLEAN TRAIN
# Segmentation(count format to words format), Cleaning(clean stop_words) & Normalization (apply porter stem etc.)
# (We've tried clean the punctuations and numerical words in keywords, but it shows less accuracy in result so we don't use it.)
# (There are repeated ID in the dataset, but as all the data is daily based include the test file, some ID has the data of over 2200 days and most of them only has 1-2 days, we don't want influence the model through visiting frenquency, so we decide not aggregate the data by ID)
# + id="zj4wiWGYrviJ" colab={"base_uri": "https://localhost:8080/"} outputId="2453c843-e232-46ec-a3b5-0d998e0082f4"
nltk.download('stopwords')
# + id="qofH9fLwpvj7"
stop_words = stopwords.words()
porter = PorterStemmer()
def cleantext(counttext):
wordcounts=counttext.split(";")
text=""
i = 0
for words in wordcounts:
wordocr=words.split(":")
print(i)
i = i+1
if len(wordocr)>1 and not wordocr[0] in stop_words :
wordocr[0]=porter.stem(wordocr[0])
text+=(wordocr[0]+" ")*int(wordocr[1])
return text
# + id="ajpp6IU1pvj8"
df=df[df["keywords"].str.contains(":")]
# + id="HsEDsPoNpvj9" colab={"base_uri": "https://localhost:8080/"} outputId="78ee6c85-a41b-42c5-f40d-9462fd66a851"
df["keywords"]=df["keywords"].map(lambda x: cleantext(x))
# + id="fBTTdf-kpvj-"
df.dropna(inplace=True)
df.to_csv("train_clean.csv",index=False,sep=',')
# + id="7qUxvwq_pvj_" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="d44942d7-fdbb-43e1-c01e-350b3585c40a"
df.head()
# + [markdown] id="_Our1RHQpvkA"
# # CLEAN TEST
# Segmentation, Cleaning & Normalization (there is no repeated ID in test data)
# + id="apgQHRaRpvkB"
test=pd.read_csv("/content/drive/MyDrive/NLP/test.csv") # 311M original
test.dropna(subset=["keywords"], inplace=True) # 285M rows after dropna in keywords
test=test[test["keywords"].str.contains(":")] #only 7 rows lost in this filter
# + id="bfeeOTyWr0Gj"
test = test[0:100]
# + id="Lds6xQl1pvkC" colab={"base_uri": "https://localhost:8080/"} outputId="5a8a2780-9e70-4a2d-84ce-4a887191133e"
test["keywords"]=test["keywords"].map(lambda x: cleantext(x))
# + id="wOAflHyRpvkC" colab={"base_uri": "https://localhost:8080/"} outputId="b1bbe5e3-7104-46af-d9bc-81ae808dd82e"
test.dropna(subset=["keywords"], inplace=True)
test.to_csv("test_clean.csv",index=False,sep=',')
# + [markdown] id="ppnN0sCBpvkD"
# # Apply TfidfVectorizer
# + id="2woUVAJFpvkD"
df=pd.read_csv("train_clean.csv")
test=pd.read_csv("test_clean.csv")
# + id="v5RkX3rbpvkE" colab={"base_uri": "https://localhost:8080/"} outputId="b58a8d32-e08a-408c-dc37-b0e05824e8fd"
X_train,X_test,y_train,y_test,z_train,z_test = train_test_split(df["keywords"],df["sex"],df["age"], test_size = 0.2, random_state = 42)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
print(z_train.shape)
print(z_test.shape)
# + id="smCWST_rpvkF" colab={"base_uri": "https://localhost:8080/"} outputId="b8431dd4-66c2-45af-a785-b7e573abd3e5"
vc_tf_idf = TfidfVectorizer()
vc_tf_idf.fit(X_train.apply(lambda x: np.str_(x)))
# + id="HNxSxeDipvkG" colab={"base_uri": "https://localhost:8080/"} outputId="5a88d5e3-cb35-4f2a-9d4d-6cea36c3a237"
print("Vocabulary size: {}".format(len(vc_tf_idf.vocabulary_)))
# + id="10JCbUFUpvkG" colab={"base_uri": "https://localhost:8080/"} outputId="8f7f9703-0cd9-4151-c21f-7c498eafa5c5"
X_train_tf = vc_tf_idf.transform(X_train.apply(lambda x: np.str_(x)))
X_train_tf[:3].nonzero()
# + [markdown] id="QoUkfeh9pvkH"
# # Build model for "Sex" and store result
# 3 models are tried: MultinomialNB, Logistic regression and Random forest classifier, and we finally chose Logistic Regression model as it has the best AUC and accuracy.
# + id="sJ3zFLxYpvkI" colab={"base_uri": "https://localhost:8080/"} outputId="b63b5326-8f2f-4c27-b1a4-616a929d09d9"
from sklearn.naive_bayes import MultinomialNB
mnb = MultinomialNB()
mnb.fit(X_train_tf,y_train)
# + id="CDEtstZVpvkJ" colab={"base_uri": "https://localhost:8080/"} outputId="bef93a5b-4e57-4c21-8d44-061103ea68bd"
from sklearn.linear_model import LogisticRegression
logreg = LogisticRegression(max_iter=10000)
logreg.fit(X_train_tf,y_train)
# + id="T5oMvifspvkK" colab={"base_uri": "https://localhost:8080/"} outputId="1f439642-6343-4e56-9e88-070bd831572f"
from sklearn.ensemble import RandomForestClassifier
rfc= RandomForestClassifier(n_estimators=2000,max_depth=8)
rfc.fit(X_train_tf,y_train)
# + id="lFXi-bYmpvkK"
X_test_tf = vc_tf_idf.transform(X_test.apply(lambda x: np.str_(x)))
# + id="jZrZeqYipvkL"
predictionsex = dict()
predictionsex["mnb"] = mnb.predict(X_test_tf)
predictionsex["logreg"] = logreg.predict(X_test_tf)
predictionsex["rfc"] = rfc.predict(X_test_tf)
# + id="DNXA93QzpvkL" colab={"base_uri": "https://localhost:8080/"} outputId="31a80fe2-dfb9-41e9-8eda-20174aef4041"
print(classification_report(y_test,predictionsex["mnb"]))
#Test result on MultinomialNB
# + id="FsfHUhSVpvkM" colab={"base_uri": "https://localhost:8080/"} outputId="3489641a-c2d7-46d5-da18-18e5216fd51d"
print(classification_report(y_test,predictionsex["logreg"]))
#Test result on LogisticRegression
# + id="ppxssd2wpvkN" colab={"base_uri": "https://localhost:8080/"} outputId="4c46d9fb-f440-428f-913a-0a651c544e8e"
print(classification_report(y_test,predictionsex["rfc"]))
#Test result on Randomforest Classifier
# + id="_W9XYmN1pvkO" colab={"base_uri": "https://localhost:8080/"} outputId="5b4b7752-5d5e-4cb2-ecec-72d675f528f6"
accuracy_score(y_test,predictionsex["logreg"])
#Logistic regression has the best accuracy
# + id="_VibdESopvkO" colab={"base_uri": "https://localhost:8080/", "height": 278} outputId="a6c95b73-e411-4893-ca0f-5c480d6f0c80"
ax = plt.gca()
test_rf = plot_roc_curve(logreg, X_test_tf, y_test,ax=ax, alpha=0.8, name="test")
train_rf = plot_roc_curve(logreg,X_train_tf, y_train,ax=ax, alpha=0.8, name="train")
plt.show()
#AUC curve for Logistic Regression
# + id="z49PhPNGpvkP" colab={"base_uri": "https://localhost:8080/", "height": 278} outputId="0cae7732-fa32-4527-ba3a-58664d9e4f0c"
ax = plt.gca()
test_rf = plot_roc_curve(mnb, X_test_tf, y_test,ax=ax, alpha=0.8, name="test")
train_rf = plot_roc_curve(mnb,X_train_tf, y_train,ax=ax, alpha=0.8, name="train")
plt.show()
#AUC curve for MultinomialNB
# + id="G_7FkAOlpvkQ" colab={"base_uri": "https://localhost:8080/", "height": 278} outputId="40be5d28-f9f7-4f2a-b884-7b92f3b23b54"
ax = plt.gca()
test_rf = plot_roc_curve(rfc, X_test_tf, y_test,ax=ax, alpha=0.8, name="test")
train_rf = plot_roc_curve(rfc,X_train_tf, y_train,ax=ax, alpha=0.8, name="train")
plt.show()
#AUC curve for RandomForest Classifier
# + id="jJ6syK5ZpvkR"
X_test_final = vc_tf_idf.transform(test["keywords"].apply(lambda x: np.str_(x)))
# + id="VlMByIP-pvkS"
result=pd.DataFrame()
result["ID"]=test["ID"]
result["sex_pred"]=logreg.predict(X_test_final)
# + id="KWzinKycpvkS"
result["sex_pred"]=result["sex_pred"].replace(1, "M")
result["sex_pred"]=result["sex_pred"].replace(0, "F")
# + id="drvdbr1GpvkT"
result.to_csv("test_result.csv",index=False,sep=',')
# + [markdown] id="2UFtuUfRpvkT"
# # Build model for "Age" and store results
# Compare to 6 other regressors that we have tried with a small dataset with 50K of rows.
# SGD regressor has the best performance in test.
# 6 other regressors are: Logistic Regression, Linear Regression, multinomialNB, Random Forest Regressor, Adabooster Regressor, and Gradientboosting Regressor.
# So for the whole dataset, we only apply SGD regressor to save time.
# + id="UzlRojiHpvkU" colab={"base_uri": "https://localhost:8080/"} outputId="480512ed-ba3d-427e-fd40-02994c876a4a"
from sklearn.linear_model import SGDRegressor
sgd = SGDRegressor(alpha=0.00001,max_iter=10000,random_state=42)
sgd.fit(X_train_tf,z_train)
# + id="8MLq2nMBpvkV" colab={"base_uri": "https://localhost:8080/"} outputId="2ad4c374-0925-474a-f99a-d00e3a311b41"
from sklearn.metrics import mean_squared_error
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import r2_score
predictionage = dict()
predictionage["sgd"] = sgd.predict(X_test_tf)
print("Mean square error:%.4f"%(mean_squared_error(z_test,predictionage["sgd"])))
print("Mean absolute error:%.4f"%(mean_absolute_error(z_test,predictionage["sgd"])))
print("R square: ", '{:.2%}'.format(r2_score(z_test,predictionage["sgd"])))
# + id="GT0BFIYrpvkW" colab={"base_uri": "https://localhost:8080/"} outputId="c8f0a0a5-b1e8-4638-dd28-d505c4eee8d1"
print(classification_report(np.floor((z_test+5)/10)*10,np.floor((predictionage["sgd"]+5)/10)*10))
# 10 stands for age group between 5-15, 20 stands for age group between 15-25 and so on
# + id="M6sw91jYpvkX" colab={"base_uri": "https://localhost:8080/"} outputId="f5498636-3cca-4e79-d6de-1f4fe4c8b635"
predictionage["dif"]=predictionage["sgd"]-z_test
predictionage["dif"]=predictionage["dif"].map(lambda x: 0 if abs(x)>5 else 1)
print("1 stands for the precision rate of the whole test data")
print(predictionage["dif"].value_counts(1))
# + id="rKAkJg4RpvkX"
result=pd.read_csv("test_result.csv")
result["age_pred"]=sgd.predict(X_test_final)
result.to_csv("test_result.csv",index=False,sep=',')
# + id="X0QrcFi0pvkY"
| 19_April(Co.Jo frnd)/NLP(Demographic Prediction)/Demographic_prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import linac3
import matplotlib.pyplot as plt
import math
# +
df = linac3.read_csv("../data/raw/Sep2018.csv").ffill()
sampling = 10
current = df[linac3.SourceFeatures.BCT05_CURRENT].resample(f"{sampling}s").mean().ffill().values
current = current[~np.isnan(current)]
htv = df[linac3.SourceFeatures.SOURCEHTAQNV].resample(f"{sampling}s").mean().values
htv = htv[~np.isnan(htv)]
current.size
# -
num_hours = 24 * 15
T = current[-num_hours * 60 * 60 // sampling:] * 1000
V = htv[-num_hours * 60 * 60 // sampling:]
T.size
# +
mask_ranges = [
(3312, 3355),
(7217, 7240),
(11201, 11222),
(14550, 14563),
(16152, 16173),
(17281, 17293),
(18343, 18361),
(18847, 18869),
(20306, 20336)
]
mask_ranges = [
(2038, 2073),
(9770, 9840),
(13710, 13790),
(16670, 16730),
(19330, 19390),
(20350, 20420),
(23280, 23330),
(28890, 28990),
(29490, 29560)
]
mask = np.concatenate([np.arange(start, stop) for start, stop in mask_ranges])
T=T[~np.isin(np.arange(T.size), mask)].copy()
T.size
# +
# %matplotlib widget
fig, ax = plt.subplots(1, 1, figsize=(20,5))
ax.plot(T[13000:16000], marker='.', ls='', ms=1, c='black')
ax.set_ylim(50, 200)
#ax.plot(np.arange(T.size)[V < 18500], T[V < 18500], color="red", marker=".", ls="", markersize="4")
ax2 = ax.twinx()
#ax2.plot(V, color="red")
plt.show()
# -
init_size = 60 * 60 // sampling # Take the first hour to estimate mean and variance of likelihood
mean_changepoint_distance = 6 * 60 * 60 // sampling # On average, one change point every six hours
# +
from bocp_detector import BayesianDetector, ConstantHazard, StudentT
def create_detector(init_size, mean_changepoint_distance):
hazard = ConstantHazard(mean_changepoint_distance)
init_mean, init_var = T[:init_size].mean(), T[:init_size].var()
if init_var < 1e-7:
init_var = 0.01
kappa = 1#init_size
alpha = 0.5#init_size * 0.5
mu = init_mean
beta = 0.5 * init_size * init_var
observation_likelihood = StudentT(alpha, beta, kappa, mu)
tail_threshold = 1e-4
detector = BayesianDetector(hazard, observation_likelihood, tail_threshold)
print(init_mean, np.sqrt(init_var))
return detector
# +
# %matplotlib inline
import matplotlib.pyplot as plt
fig, ax = plt.subplots(1, 1)
ax.plot(T[:init_size])
ax.set_ylim(0, 250)
# +
np.seterr(all="raise")
stepX, stepY = 6, 20
detector = create_detector(init_size, mean_changepoint_distance)
end = 20000#T.size
rec_probs = 3*mean_changepoint_distance
growth_matrix = np.zeros(((end - init_size) // stepX, rec_probs), dtype=float)
predictive_mean = np.zeros((end - init_size) // stepX, dtype=float)
predictive_std = np.zeros((end - init_size) // stepX, dtype=float)
win_size = 250
win_mean = np.zeros((end - init_size) // stepX, dtype=float)
win_std = np.zeros((end - init_size) // stepX, dtype=float)
cp = set([0])
for i, (x, v) in enumerate(zip(T[init_size:end], V[init_size:end])):
if i % 1000 == 0:
print(f"{i}/{end - init_size}")
#if i == 1000 - init_size:
# import pdb; pdb.set_trace()
detector.update(x)
new_cp = detector.detect(0.8, 0.1)
if new_cp >= 0:
#import pdb; pdb.set_trace()
cp.add(i - new_cp)
growth_probs = detector.get_growth_probabilities()
l = min(growth_probs.size, rec_probs)
if i % stepX == 0:
growth_matrix[i // stepX, :l] = growth_probs[:l]
predictive_mean[i // stepX] = detector.predictive_mean()
predictive_std[i // stepX] = detector.predictive_std()
if i > 1:
win_mean[i // stepX] = T[max(0, i-win_size):i].mean()
win_std[i // stepX] = T[max(0, i-win_size):i].std()
print(f"{end - init_size}/{end - init_size}")
# +
#cp = [0, 886, 1270, 1718, 1988, 2368, 2810, 3149, 3280, 3565, 3663, 3963, 4495, 4930, 5050, 5463]
# +
# %matplotlib widget
from mpl_toolkits.axes_grid1 import make_axes_locatable
import matplotlib.ticker as ticker
fig, ax = plt.subplots(2, 1, sharex=True, figsize=(15, 5), constrained_layout=True)
ax[0].plot(T[init_size:end:stepX], color="black")#, ls="", marker=".", ms=2)
#ax[0].plot(predictive_mean, color="firebrick")
#ax[0].plot(predictive_mean + predictive_std, color="firebrick", ls="--")
#ax[0].plot(predictive_mean - predictive_std, color="firebrick", ls="--")
ax[0].set_ylim(50, 200)
ax[0].vlines([c / stepX for c in cp], ymin=0, ymax=1, transform=ax[0].get_xaxis_transform(), color="blue")
ax0_twin = ax[0].twinx()
#ax0_twin.plot(growth_matrix.argmax(axis=1), color="red")
ax[0].set_ylabel("BCT05 current [uA]")
cmap = plt.get_cmap("Greys")
gmplot = growth_matrix.T.copy()
gmplot = np.log(gmplot, where=gmplot > 0, out=np.full(gmplot.shape, -np.inf))
im = ax[1].pcolormesh(gmplot[:10000:stepY, ::], cmap=cmap, vmin=-20, vmax=0)
#ax[1].plot(np.argmax(growth_matrix, axis=1) / stepY, ls="--", color="red")
cb = fig.colorbar(im, ax=ax[1])
def label_exp(x,pos):
return "{:.0E}".format(np.exp(x))
cb.set_ticks([np.log(t) for t in [1, 0.1, 0.01, 0.001, 0.0001, 1e-5, 1e-6, 1e-7]])
cb.formatter = ticker.FuncFormatter(label_exp)
#cb.set_ticks([0], update_ticks=True)
cb.update_ticks()
ax[1].set_ylabel("Posterior run length distribution")
#ax[2].plot(win_mean)
#ax[2].set_ylim(150, 180)
#ax[2].set_ylim(0, 20)
#ax2_twin = ax[2].twinx()
#ax[2].vlines([c // stepX for c in cp], ymin=0, ymax=1, transform=ax[2].get_xaxis_transform())
plt.show()
# -
cpl = sorted(cp)
cpl.append(end - init_size)
segments = [(cpl[i-1], cpl[i], T[init_size+cpl[i-1]:init_size+cpl[i]]) for i in range(1, len(cpl))]
segments[3]
# +
# %matplotlib widget
from mpl_toolkits.axes_grid1 import make_axes_locatable
import matplotlib.ticker as ticker
fig, ax = plt.subplots(2, 1, sharex=True, figsize=(25, 10), constrained_layout=True)
ax[0].plot(T[init_size:end:stepX], color="black", ls="", marker=".", ms=3)
#ax[0].plot(predictive_mean, color="firebrick")
#ax[0].plot(predictive_mean + predictive_std, color="firebrick", ls="--")
#ax[0].plot(predictive_mean - predictive_std, color="firebrick", ls="--")
ax[0].set_ylim(50, 200)
ax[0].vlines([c // stepX for c in cp], ymin=0, ymax=1, transform=ax[0].get_xaxis_transform())
ax0_twin = ax[0].twinx()
#ax0_twin.plot(growth_matrix.argmax(axis=1), color="red")
for s, e, seg in segments:
ax[0].hlines(seg.mean(), xmin=s, xmax=e, color="firebrick")
ax[0].hlines(seg.mean()+seg.std(), xmin=s, xmax=e, color="firebrick", ls="--")
ax[0].hlines(seg.mean()-seg.std(), xmin=s, xmax=e, color="firebrick", ls="--")
cmap = plt.get_cmap("Greys")
gmplot = growth_matrix.T.copy()
gmplot = np.log(gmplot, where=gmplot > 0, out=np.full(gmplot.shape, -np.inf))
im = ax[1].pcolormesh(gmplot[:1000:stepY, ::], cmap=cmap, vmin=-20, vmax=0)
cb = fig.colorbar(im, ax=ax[1])
def label_exp(x,pos):
return "{:.0E}".format(np.exp(x))
cb.set_ticks([np.log(t) for t in [1, 0.1, 0.01, 0.001, 0.0001, 1e-5, 1e-6, 1e-7]])
cb.formatter = ticker.FuncFormatter(label_exp)
#cb.set_ticks([0], update_ticks=True)
cb.update_ticks()
#ax[2].plot(win_mean)
#ax[2].set_ylim(150, 180)
#ax[2].set_ylim(0, 20)
#ax2_twin = ax[2].twinx()
#ax[2].vlines([c // stepX for c in cp], ymin=0, ymax=1, transform=ax[2].get_xaxis_transform())
plt.show()
# +
# %matplotlib inline
import matplotlib.animation as animation
plt.rcParams["animation.html"] = "jshtml"
from IPython.display import HTML
fig, ax = plt.subplots(1, 1, figsize=(25,6))
ax.set_xlim((0, growth_matrix.shape[1]))
line, = ax.plot([], [], lw=2)
def init():
line.set_data([], [])
return line,
def animate(i):
y = growth_matrix[i+850:]
x = np.arange(y.size)
line.set_data(x, y)
print(i)
return line,
anim = animation.FuncAnimation(fig, animate, init_func=init, frames=200, interval=1, blit=True)
anim
# +
import scipy.stats as stats
np.seterr(all='raise')
stats.t.pdf(x=100, df=1000, loc=100, scale=1)
# -
| changepoints/bocd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.2.0
# language: julia
# name: julia-1.2
# ---
include("leap_frog_definitions.jl")
using PyCall
# pygui(:qt)
using PyPlot
# pygui(true)
using PolynomialRoots
@time P1_find_dimer(1,1,1/4)
poly_dimer(X) = (-16)*H^2 + 32*H^2*P^2 + (-24)*H^2*P^4 +8*H^2*P^6 + (-1)*H^2*P^8 + (-32)*H^2*Q^2 + 16*H^2*P^2*Q^2 + 8*H^2*P^4*Q^2 + (-4)*H^2*P^6*Q^2 + (-24)*H^2*Q^4 + (-8)*H^2*P^2*Q^4 + (-6)*H^2*P^4*Q^4 + (-8)*H^2*Q^6 + (-4)*H^2*P^2*Q^6 + (-1)*H^2*Q^8 +
(32*H^2 + 16*P^2 + (-16)*H^2*P^2 + (-8)*H^2*P^4 + 4*H^2*P^6 + 16*Q^2 + 16*H^2*Q^2 + (-80)*H^2*P^2*Q^2 + 4*H^2*P^4*Q^2 + (-8)*H^2*Q^4 + (-4)*H^2*P^2*Q^4 + (-4)*H^2*Q^6)* X^2 +
((-24)*H^2 + (-8)*H^2*P^2 + (-6)*H^2*P^4 + 8*H^2*Q^2 + 4*H^2*P^2*Q^2 + (-6)*H^2*Q^4)* X^4 +
(8*H^2 + 4*H^2*P^2 + (-4)*H^2*Q^2)*X^6+
(-1)*H^2*X^8
# +
Q=.5;
P=0;
H=1/4;
coeff=(-16)*H^2 + 32*H^2*P^2 + (-24)*H^2*P^4 +8*H^2*P^6 + (-1)*H^2*P^8 + (-32)*H^2*Q^2 + 16*H^2*P^2*Q^2 + 8*H^2*P^4*Q^2 + (-4)*H^2*P^6*Q^2 + (-24)*H^2*Q^4 + (-8)*H^2*P^2*Q^4 + (-6)*H^2*P^4*Q^4 + (-8)*H^2*Q^6 + (-4)*H^2*P^2*Q^6 + (-1)*H^2*Q^8
coeff2=(32*H^2 + 16*P^2 + (-16)*H^2*P^2 + (-8)*H^2*P^4 + 4*H^2*P^6 + 16*Q^2 + 16*H^2*Q^2 + (-80)*H^2*P^2*Q^2 + 4*H^2*P^4*Q^2 + (-8)*H^2*Q^4 + (-4)*H^2*P^2*Q^4 + (-4)*H^2*Q^6)
coeff4=((-24)*H^2 + (-8)*H^2*P^2 + (-6)*H^2*P^4 + 8*H^2*Q^2 + 4*H^2*P^2*Q^2 + (-6)*H^2*Q^4)
coeff6=(8*H^2 + 4*H^2*P^2 + (-4)*H^2*Q^2)
coeff8=(-1)*H^2;
poynomial8=[coeff,0,coeff2,0,coeff4,0,coeff6,0,coeff8];
poynomial4=[coeff,coeff2,coeff4,coeff6,coeff8];
# -
@time sol=roots(poynomial4,polish=true)
for k in 1:length(sol)
if abs(imag(sol[k]))<1e-15
sol[k]=real(sol[k])
end
end
sol
sol=roots(poynomial4)
for k in 1:length(sol)
if abs(imag(sol[k]))<1e-15
sol[k]=real(sol[k])
end
end
sol
function P1_poly(Q,P,H)
coeff=(-16)*H^2 + 32*H^2*P^2 + (-24)*H^2*P^4 +8*H^2*P^6 + (-1)*H^2*P^8 + (-32)*H^2*Q^2 + 16*H^2*P^2*Q^2 + 8*H^2*P^4*Q^2 + (-4)*H^2*P^6*Q^2 + (-24)*H^2*Q^4 + (-8)*H^2*P^2*Q^4 + (-6)*H^2*P^4*Q^4 + (-8)*H^2*Q^6 + (-4)*H^2*P^2*Q^6 + (-1)*H^2*Q^8
coeff2=(32*H^2 + 16*P^2 + (-16)*H^2*P^2 + (-8)*H^2*P^4 + 4*H^2*P^6 + 16*Q^2 + 16*H^2*Q^2 + (-80)*H^2*P^2*Q^2 + 4*H^2*P^4*Q^2 + (-8)*H^2*Q^4 + (-4)*H^2*P^2*Q^4 + (-4)*H^2*Q^6)
coeff4=((-24)*H^2 + (-8)*H^2*P^2 + (-6)*H^2*P^4 + 8*H^2*Q^2 + 4*H^2*P^2*Q^2 + (-6)*H^2*Q^4)
coeff6=(8*H^2 + 4*H^2*P^2 + (-4)*H^2*Q^2)
coeff8=(-1)*H^2;
poynomial4=[coeff,coeff2,coeff4,coeff6,coeff8];
p1=[]
sol=roots(poynomial4,polish=true)
for k in 1:length(sol)
if abs(imag(sol[k]))<1e-15
push!(p1,sqrt(abs(sol[k])))
end
end
return p1
end
@time P1_poly(Q,P,H)[2]
@time P1_find_dimer_second(Q,P,H)
# +
function f(Q)
sol=P1_poly(Q,0,H)
if isempty(sol)
return 0
else
return minimum(sol)
end
end
# function f(Q)
# sol=P1_poly(Q,0,H)
# if length(sol)==2
# return sol[2]
# else
# return 0
# end
# end
# -
X=range(0, stop = 3, length = 100)
N=length(X)
Y=zeros(N)
for k in 1:N
Y[k]=f(X[k])
end
plot(X,Y)
X=range(0, stop = .1, length = 10000)
N=length(X)
Y=zeros(N)
for k in 1:N
Y[k]=f(X[k])
end
plot(X,Y)
f(X)
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:iblenv] *
# language: python
# name: conda-env-iblenv-py
# ---
# +
from ibl_pipeline import acquisition, behavior, ephys, subject
from ibl_pipeline.analyses import behavior as behavior_analyses
import datajoint as dj
import pandas as pd
from uuid import UUID
from oneibl.one import ONE
one = ONE()
# -
# grab insertions and behavioral QC from datajoint
sessions = acquisition.Session & (acquisition.SessionProject & 'session_project like "%brainwide%"') & \
ephys.DefaultCluster()
insertions_qc = subject.Subject * subject.SubjectLab * ephys.ProbeInsertion * sessions * (behavior_analyses.SessionTrainingStatus)
insertions_qc = insertions_qc.proj('session_uuid', 'session_lab',
'subject_nickname', behavior_qc_passed='good_enough_for_brainwide_map')
insertions = insertions_qc.fetch(format='frame').reset_index()
insertions['eid'] = [eid.urn[9:] for eid in insertions.session_uuid] # convert to string
insertions.shape
# +
# # THIS IS SUPER SLOW, SEE https://github.com/int-brain-lab/iblenv/issues/15#issuecomment-682150015
# for eid, dat in insertions.groupby(['session_uuid']):
# # bit awkward, have to extract string from UUID object
# sess_info = one.alyx.rest('sessions', 'read', id=eid.urn[9:])
# insertions.loc[insertions.session_uuid == eid, 'alyx_qc'] = sess_info['qc']
# print(eid, ': ', sess_info['qc'])
# -
insertions['alyx_qc'] = 'empty'
qcs = ['critical', 'error', 'warning','not_set']
for qc in qcs:
# grab those eids
sess = list(one.alyx.rest('sessions', 'list', qc=qc))
eids = [s['url'] for s in sess]
eids = [e.split('/')[-1] for e in eids]
insertions.loc[insertions['eid'].isin(eids), 'alyx_qc'] = qc
# SUMMARIZE PER LAB/USER
insertions.groupby(['alyx_qc', 'behavior_qc_passed', 'session_lab'])['session_start_time'].count()
# SHOW EACH INSERTION WITH ITS QC VALUES
pd.set_option('display.max_rows', 1000)
insertions
# +
from oneibl.one import ONE
one = ONE()
# find projects: proj = one.alyx.rest('projects', 'list')
# proj_list = [p['name'] for p in proj]
traj = one.alyx.rest('trajectories', 'list', provenance='Planned',
x=-2243, y=-2000, # repeated site coordinate
project='ibl_neuropixel_brainwide_01')
# Print which sessions have not been hist-ephys aligned
for i_tr in range(0, len(traj)):
eid = traj[i_tr]['session']['id']
probe_label = traj[i_tr]['probe_name']
traj_align = one.alyx.rest('trajectories', 'list',
provenance='Ephys aligned histology track',
session=eid, probe=probe_label)
if len(traj_align) == 0:
subject = traj[i_tr]['session']['subject']
date = traj[i_tr]['session']['start_time'][0:10]
sess_info = one.alyx.rest('sessions', 'read', id=eid)
qc = sess_info['qc']
print(f'Not aligned: {eid} - {subject} - {date} - Session QC: {qc}')
# Print which sessions have not been hist-ephys aligned
for i_tr in range(0, len(traj)):
eid = traj[i_tr]['session']['id']
probe_label = traj[i_tr]['probe_name']
traj_align = one.alyx.rest('trajectories', 'list',
provenance='Ephys aligned histology track',
session=eid, probe=probe_label)
if len(traj_align) == 1:
subject = traj[i_tr]['session']['subject']
date = traj[i_tr]['session']['start_time'][0:10]
sess_info = one.alyx.rest('sessions', 'read', id=eid)
qc = sess_info['qc']
print(f'Aligned: {eid} - {subject} - {date} - Session QC: {qc}')
# -
| python/.ipynb_checkpoints/session_qc_overview-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# ## Introduction to Geospatial Data
# ### Part 4 of 5
# # Indirect georeferencing
# + [markdown] slideshow={"slide_type": "slide"}
#
# ## Reminder
# <a href="#/slide-2-0" class="navigate-right" style="background-color:blue;color:white;padding:8px;margin:2px;font-weight:bold;">Continue with the lesson</a>
#
# <font size="+1">
#
# By continuing with this lesson you are granting your permission to take part in this research study for the Hour of Cyberinfrastructure: Developing Cyber Literacy for GIScience project. In this study, you will be learning about cyberinfrastructure and related concepts using a web-based platform that will take approximately one hour per lesson. Participation in this study is voluntary.
#
# Participants in this research must be 18 years or older. If you are under the age of 18 then please exit this webpage or navigate to another website such as the Hour of Code at https://hourofcode.com, which is designed for K-12 students.
#
# If you are not interested in participating please exit the browser or navigate to this website: http://www.umn.edu. Your participation is voluntary and you are free to stop the lesson at any time.
#
# For the full description please navigate to this website: <a href="../../gateway-lesson/gateway/gateway-1.ipynb">Gateway Lesson Research Study Permission</a>.
#
# </font>
# + hide_input=true init_cell=true slideshow={"slide_type": "skip"} tags=["Hide"]
# This code cell starts the necessary setup for Hour of CI lesson notebooks.
# First, it enables users to hide and unhide code by producing a 'Toggle raw code' button below.
# Second, it imports the hourofci package, which is necessary for lessons and interactive Jupyter Widgets.
# Third, it helps hide/control other aspects of Jupyter Notebooks to improve the user experience
# This is an initialization cell
# It is not displayed because the Slide Type is 'Skip'
from IPython.display import HTML, IFrame, Javascript, display
from ipywidgets import interactive
import ipywidgets as widgets
from ipywidgets import Layout
import getpass # This library allows us to get the username (User agent string)
# import package for hourofci project
import sys
sys.path.append('../../supplementary') # relative path (may change depending on the location of the lesson notebook)
import hourofci
# load javascript to initialize/hide cells, get user agent string, and hide output indicator
# hide code by introducing a toggle button "Toggle raw code"
HTML('''
<script type="text/javascript" src=\"../../supplementary/js/custom.js\"></script>
<style>
.output_prompt{opacity:0;}
</style>
<input id="toggle_code" type="button" value="Toggle raw code">
''')
# + [markdown] slideshow={"slide_type": "slide"}
# # Indirect georeferencing
#
# Relates the data to geography via an identifier.
#
# Some examples of referenced geography are:
# - Place names
# - Census zones
# - Zip codes, Postal codes
# - Administrative districts
# - Telephone area codes
# - Addresses
# - Linear Referencing (e.g. mileposts)
# - Geographic references in tweets
# + [markdown] slideshow={"slide_type": "slide"}
# Indirect georeferencing is really important! It allows you to connect tables of data to geography.
# <table>
# <tr style="background: #fff">
# <td width=75%> <img src='supplementary/indirect.png' alt='Indirect Georeferencing'></td>
# </tr>
# </table>
# + [markdown] slideshow={"slide_type": "slide"}
# # Let's try that with code.
#
# Open this notebook and run each of the code chunks sequentially by clicking the arrow beside the code chunk.
#
# <a href="gd-example_2.ipynb">Indirect georeferencing</a>
| beginner-lessons/geospatial-data/gd-5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + cell_id="00003-6c6aae3c-9544-40bd-863d-b30d731618ff" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=3711 execution_start=1628865583263 source_hash="64cb0142" tags=[]
# Packages that need installing on startup
# #!pip install ipywidgets
#if there are futher issues with librosa complaining about sndfile on import, run the following
# #!apt-get -y update && apt-get -y install libsndfile1
# + cell_id="00004-fa575087-dedb-4dec-86e6-df7d22ca4bef" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=5 execution_start=1628865629184 source_hash="10b5cae7" tags=[]
import os
import seaborn as sns
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import IPython.display as ipd
import librosa as lb
# Issue with installing librosa, using wavfile for visualization instead
#import ipywidgets as widgets
from scipy.io import wavfile
from sklearn.model_selection import StratifiedShuffleSplit
from pathlib import Path
plt.rcParams['figure.figsize'] = [12, 8]
# +
# Setup relative data paths, will need to be changed if running in deepnote
RAW_FOLDER = "../data/raw"
METADATA_FOLDER = "../data/metadata"
PROCESSED_FOLDER = "../data/processed"
# + cell_id="00004-15e702f7-554c-494d-a86c-fd8069ad2ccd" deepnote_cell_type="code" deepnote_to_be_reexecuted=true execution_millis=1093 execution_start=1628545902008 source_hash="31e51c06" tags=[]
df = pd.read_excel(f"{METADATA_FOLDER}/Age-sex calls- Dzanga Bai.xlsx",sheet_name="context")
df.head()
# + [markdown] cell_id="00004-7286800e-24a5-4edb-bbca-80092a8d1bc3" deepnote_cell_type="markdown" tags=[]
# ## Missingness by strata
# + cell_id="00005-8237d25d-29a2-48b1-ae9e-aecbe4be7a5e" deepnote_cell_type="code" deepnote_to_be_reexecuted=true execution_millis=33 execution_start=1628545903593 source_hash="2ca89ed0" tags=[]
n_records = len(df)
print(n_records)
valid_age = len(df[df.age != "un"]) / n_records
valid_sex = len(df[df.sex != "un"]) / n_records
valid_agsex = len(df[df.agsex != "un"]) / n_records
print(f"Proportion of data available: \nAge: {valid_age}\nSex: {valid_sex}\nAgeSex: {valid_agsex}")
# + cell_id="00003-f924168d-5468-4f40-a454-065e0ff8b695" deepnote_cell_type="code" deepnote_to_be_reexecuted=true execution_millis=215 execution_start=1628545903595 source_hash="7f2142ea" tags=[]
plt.hist(df.age)
plt.title("Distribution of labels Age")
plt.show()
# + cell_id="00004-0d402caf-6026-44e9-ba5b-e215bc487570" deepnote_cell_type="code" deepnote_to_be_reexecuted=true execution_millis=162 execution_start=1628545903798 source_hash="656ee246" tags=[]
plt.hist(df.sex)
plt.title("Distribution of labels Sex")
plt.show()
# + cell_id="00005-ed8e22b5-4bc4-4a33-9b9a-08c39ea535be" deepnote_cell_type="code" deepnote_to_be_reexecuted=true execution_millis=206 execution_start=1628545903993 source_hash="1b86dc50" tags=[]
plt.hist(df.agsex)
plt.title("Distribution of labels Age Sex")
plt.show()
# + [markdown] cell_id="00007-53005912-4566-4707-8a03-1950567aa81e" deepnote_cell_type="markdown" tags=[]
#
# + cell_id="00006-5795e386-7518-4d55-b989-295da6bc703e" deepnote_cell_type="code" deepnote_to_be_reexecuted=true execution_millis=8 execution_start=1628545904173 source_hash="b8e61c93" tags=[]
y = df.age.values
sss = StratifiedShuffleSplit(n_splits=1,test_size=0.3, random_state=42)
for train_idx, test_idx in sss.split(np.zeros(len(y)), y):
y_train=y[train_idx]
y_test=y[test_idx]
# + cell_id="00011-c175ad37-f815-43be-b7ca-bebe83d35b79" deepnote_cell_type="code" deepnote_to_be_reexecuted=true execution_millis=388 execution_start=1628545904181 source_hash="bf20ad6a" tags=[]
plt.subplot(1, 3, 1)
plt.title("Age test set")
plt.hist(sorted(y_test))
plt.subplot(1, 3, 2)
plt.title("Age train set")
plt.hist(sorted(y_train))
plt.subplot(1, 3, 3)
plt.title("Age population")
plt.hist(sorted(y))
plt.show()
# + [markdown] cell_id="00013-0c9eaaf8-817a-4cec-a308-6fcac34afe83" deepnote_cell_type="markdown" tags=[]
# ## Creating spectograms
# + cell_id="00014-836a5ad4-a882-4460-815f-f00371d36579" deepnote_cell_type="code" deepnote_to_be_reexecuted=true execution_millis=654 execution_start=1628545904559 source_hash="437ac02b" tags=[]
# Testing with a single wav file
audio_data = f'{RAW_FOLDER}/opp_329_ro.wav'
#
#ipd.Audio(audio_data,rate=60)
samplingFrequency, signalData = wavfile.read(audio_data)
# Plot the signal read from wav file
plt.subplot(121)
plt.title('Spectrogram of a wav file')
plt.plot(signalData)
plt.xlabel('Sample')
plt.ylabel('Amplitude')
plt.subplot(122)
plt.specgram(signalData,Fs=samplingFrequency)
plt.xlabel('Time')
plt.ylabel('Frequency')
plt.show()
# + [markdown] cell_id="00014-c750f5b0-d232-4466-84b5-421abf080fdc" deepnote_cell_type="markdown" tags=[] pycharm={"name": "#%% md\n"}
# ##### This is an attempt to view individual spectrograms, but doesnt seem to work in this environment
#
# Looks like this feature is not supported in deepnote https://community.deepnote.com/c/ask-anything/when-will-ipywidgets-support-be-added
# + [markdown] cell_id="00018-cf06cbda-29a0-4884-801b-ac8e990ce1a7" deepnote_cell_type="markdown" tags=[] pycharm={"name": "#%% md\n"}
# ## Create spectrograms - first pass
#
# We will create spectrograms and save to spectorgrams folder for use in CNN models
#
# + cell_id="00019-213fcc7d-9339-4a2d-89ed-4ac66fa39d9f" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=11 execution_start=1628865643996 source_hash="e9b535cc" tags=[] pycharm={"name": "#%%\n"}
def wav_to_spectrogram(path):
samplingFrequency, signalData = wavfile.read(path)
plt.specgram(signalData,Fs=samplingFrequency)
#plt.show()
plt.savefig(f"{PROCESSED_FOLDER}/{path.stem}.png")
# Store all wav files as Path objects
wav_paths = sorted(Path(RAW_FOLDER).glob('*.wav'))
# Superceded by mel spectograms, dont run
#list(map(wav_to_spectrogram,wav_paths))
# + [markdown] cell_id="00022-ec11fef9-7dcb-4303-ba96-cfb259cac3f9" deepnote_cell_type="markdown" tags=[]
# ### TODO: Create Mel spectrograms to replace wav spectrograms using librosa
#
# Only a fraction of the spectrograms are created as the machine timed out. Will have to create in batch
# + cell_id="00023-fc181053-9be8-4f46-a3b5-78b81b877b1d" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=894 execution_start=1628866189116 source_hash="68ae0a4b" tags=[]
import librosa
import librosa.display
# Testing with a single wav file
audio_data = f'{RAW_FOLDER}/opp_329_ro.wav'
#
#ipd.Audio(audio_data,rate=60)
y, sr = librosa.load(audio_data)
# Passing through arguments to the Mel filters
S = librosa.feature.melspectrogram(y=y, sr=sr)#, n_mels=512,
# fmax=8000)
fig, ax = plt.subplots()
S_dB = librosa.power_to_db(S, ref=np.max)
img = librosa.display.specshow(S_dB, x_axis='time',
y_axis='mel', sr=sr, ax=ax)
fig.colorbar(img, ax=ax, format='%+0.0f dB')
ax.set(title='Mel-frequency spectrogram')
# + [markdown] cell_id="00025-77433c3f-38ec-497b-9363-eb2da23c4394" deepnote_cell_type="markdown" tags=[]
# S = librosa.feature.melspectrogram(y=y, sr=sr, n_mels=512)
#
# fig, ax = plt.subplots()
# S_dB = librosa.power_to_db(S, ref=np.max)
# img = librosa.display.specshow(S_dB, x_axis='time',
# y_axis='mel', sr=sr, ax=ax)
# fig.colorbar(img, ax=ax, format='%+0.0f dB')
# ax.set(title='Mel-frequency spectrogram')
# -
# Here we are exploring the features provided in the measurement tab of the metadata. We will look to see if there is any apparent structure to the data by performing a PCA.
# + cell_id="00025-92bf6550-bd51-40ca-8572-953528cfe69e" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=583 execution_start=1628589500031 source_hash="ead60dbc" tags=[]
features_df = pd.read_excel(f"{METADATA_FOLDER}/Age-sex calls- Dzanga Bai.xlsx", sheet_name="measurements")
features_df.head()
# + cell_id="00027-4c485750-80b5-42d9-8c87-14d9c5686df4" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=14 execution_start=1628589502213 source_hash="fc82a4cb" tags=[]
from sklearn.preprocessing import StandardScaler
# Separating out the features
# Removing text features, not clear what these are for
# call_type and call have 0 variance, analysis looks like truncated text
features_df.drop(["call_type","call","analysis"], axis=1, inplace=True)
features_df.reset_index(inplace=True)
features_df.dropna(inplace=True)
id_col = 'unique_ID'
x = features_df.loc[:, features_df.columns != id_col].values
# Separating out the target
y = features_df.loc[:,[id_col]].values
# Standardizing the features for use in PCA
x = StandardScaler().fit_transform(x)
# + cell_id="00028-663eeafc-dc70-49f8-81e4-ced19a868edd" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=6 execution_start=1628589505048 source_hash="577d5527" tags=[]
x
# + cell_id="00026-7b88f33e-8865-4767-b3ad-e74264f1997a" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=346 execution_start=1628589507697 source_hash="b28ee246" tags=[]
from sklearn.decomposition import PCA
pca = PCA().fit(x)
plt.plot(np.cumsum(pca.explained_variance_ratio_))
plt.xlabel('number of components')
plt.ylabel('cumulative explained variance')
# + [markdown] cell_id="00030-4688cc08-9d4b-473a-acff-e7d4737e0cd7" deepnote_cell_type="markdown" tags=[]
# Looks like we need about 9 principal components to account for around 90% of the variance using the provided features alone. Features may not be a good way to predict labels.
# + [markdown] created_in_deepnote_cell=true deepnote_cell_type="markdown" tags=[]
# <a style='text-decoration:none;line-height:16px;display:flex;color:#5B5B62;padding:10px;justify-content:end;' href='https://deepnote.com?utm_source=created-in-deepnote-cell&projectId=7aa10abb-09ba-4e5c-b109-d1b9cd39afe4' target="_blank">
# <img alt='Created in deepnote.com' style='display:inline;max-height:16px;margin:0px;margin-right:7.5px;' src='data:image/svg+xml;base64,<KEY> > </img>
# Created in <span style='font-weight:600;margin-left:4px;'>Deepnote</span></a>
| notebooks/1.0-kdt-initial-data-exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/96jonesa/CSE-517-Project/blob/main/testing.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Qp0_axn_A0RP"
# #Imports
# + id="IouH5zC4C6tM"
# !pip3 install --quiet "tensorflow-hub>=0.7.0"
# !pip3 install --quiet seaborn
# !pip3 install --quiet pandas-market-calendars
# + id="hkbW7y5PAL4E"
import torch
import torch.nn as nn
from torch import optim
import torch.nn.functional as F
from absl import logging
import tensorflow.compat.v1 as tf
import tensorflow_hub as hub
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import os
import re
import seaborn as sns
import json
import itertools
import pandas as pd
import torch
import pandas_market_calendars as mcal
import datetime
from torch.utils.data import Dataset, DataLoader
from tqdm.notebook import tqdm
# + [markdown] id="B3_bjmyzAy67"
# #Layers
# + id="V_I8XFDzAQaW"
class GRU(nn.Module):
def __init__(self, input_size, hidden_size, batch_first=False):
super(GRU, self).__init__()
self.input_size = input_size
self.hidden_size = hidden_size
self.batch_first = batch_first
self.gru = nn.GRU(input_size, hidden_size, batch_first=self.batch_first)
def forward(self, input):
output, hn = self.gru(input)
return output, hn
# + id="8pc_MLjpAQsN"
# attention weights are softmax(u^T tanh(W input + b)) where W is learned parameter matrix, u is a learned parameter vector, and b is a learned offset
class LinearAttention(nn.Module):
def __init__(self, input_size, intermediate_size, weights_size):
super(LinearAttention, self).__init__()
self.input_size = input_size
self.intermediate_size = intermediate_size
self.weights_size = weights_size
self.linear_1 = nn.Linear(self.input_size, self.intermediate_size, bias=True)
self.linear_2 = nn.Linear(self.intermediate_size, self.weights_size, bias=False)
self.tanh = nn.Tanh()
self.softmax = nn.Softmax(dim=2)
def forward(self, input, mask=None):
intermediate = self.tanh(self.linear_1(input))
pre_attention = self.linear_2(intermediate)
if mask is not None:
zero_vec = -9e15*torch.ones_like(pre_attention)
pre_attention = torch.where(mask > 0, pre_attention, zero_vec)
attention_weights = self.softmax(pre_attention)
attention_weights = attention_weights.permute(0, 2, 1)
output_features = torch.bmm(attention_weights, input)
return output_features
# + id="eZ4wJCUJATJ5"
# output is ReLU(left^T W right + b) where W is a learned paramater matrix
# and b is a learned bias
class Blend(nn.Module):
def __init__(self, left_size, right_size, output_size):
super(Blend, self).__init__()
self.left_size = left_size
self.right_size = right_size
self.output_size = output_size
self.bilinear = nn.Bilinear(self.left_size, self.right_size, output_size, bias=True)
self.relu = nn.ReLU()
def forward(self, left, right):
output = self.relu(self.bilinear(left, right))
return output
# + id="UOSXvN33Ae-Q"
# https://github.com/Diego999/pyGAT/blob/master/layers.py
class SGAT(nn.Module):
def __init__(self, input_size, output_size, leakyrelu_slope=0.01):
super(SGAT, self).__init__()
self.input_size = input_size
self.output_size = output_size
self.leakyrelu_slope = leakyrelu_slope
self.W = nn.Parameter(torch.empty(size=(input_size, output_size)))
nn.init.xavier_uniform_(self.W.data, gain=1.414)
self.a = nn.Parameter(torch.empty(size=(2*output_size, 1)))
nn.init.xavier_uniform_(self.a.data, gain=1.414)
self.leakyrelu = nn.LeakyReLU(self.leakyrelu_slope)
def forward(self, h, adj):
Wh = torch.mm(h, self.W)
a_input = self._prepare_attentional_mechanism_input(Wh)
e = self.leakyrelu(torch.matmul(a_input, self.a).squeeze(2))
zero_vec = -9e15*torch.ones_like(e)
attention = torch.where(adj > 0, e, zero_vec)
attention = F.softmax(attention, dim=1)
h_prime = torch.matmul(attention, Wh)
return h_prime
def _prepare_attentional_mechanism_input(self, Wh):
N = Wh.size()[0] # number of nodes
Wh_repeated_in_chunks = Wh.repeat_interleave(N, dim=0)
Wh_repeated_alternating = Wh.repeat(N, 1)
all_combinations_matrix = torch.cat([Wh_repeated_in_chunks, Wh_repeated_alternating], dim=1)
return all_combinations_matrix.view(N, N, 2 * self.output_size)
# + id="SgbnNp3nAh47"
class MANSF(nn.Module):
def __init__(self, T, gru_hidden_size, attn_inter_size, use_embed_size,
blend_size, gat_1_inter_size, gat_2_inter_size, leakyrelu_slope, elu_alpha, U):
super(MANSF, self).__init__()
self.T = T
self.gru_hidden_size = gru_hidden_size
self.attn_inter_size = attn_inter_size
self.use_embed_size = use_embed_size
self.blend_size = blend_size
self.gat_1_inter_size = gat_1_inter_size
self.gat_2_inter_size = gat_2_inter_size
self.leakyrelu_slope = leakyrelu_slope
self.elu_alpha = elu_alpha
self.U = U
self.gru_p = GRU(3, gru_hidden_size, batch_first=True)
self.gru_m = GRU(use_embed_size, gru_hidden_size, batch_first=True)
self.gru_s = GRU(gru_hidden_size, gru_hidden_size, batch_first=True)
self.attn_p = LinearAttention(gru_hidden_size, attn_inter_size, 1)
self.attn_m = LinearAttention(gru_hidden_size, attn_inter_size, 1)
self.attn_s = LinearAttention(gru_hidden_size, attn_inter_size, 1)
self.blend = Blend(gru_hidden_size, gru_hidden_size, blend_size)
self.mgat_1 = nn.ModuleList([SGAT(blend_size, gat_1_inter_size, leakyrelu_slope=leakyrelu_slope) for u in range(U)])
self.mgat_2 = nn.ModuleList([SGAT(U * gat_1_inter_size, gat_2_inter_size, leakyrelu_slope=leakyrelu_slope) for u in range(U)])
self.sigmoid = nn.Sigmoid()
self.elu = nn.ELU(elu_alpha)
self.final_linear = nn.Linear(U * gat_2_inter_size, 1, bias=True)
# p is price data tensor of shape (num_stocks, T, 3), for the day under consideration
# m is smi data list of tensors of shape (num_stocks, K, use_embed_size) of length T,
# where K is the number of tweets for the given stock on the day under consideration
# neighorhoods is a list of adjacency lists, where each stock is indexed with the same
# indices they have in p and m
def forward(self, p, m, m_mask, neighborhoods):
## price encoding
h_p, _ = self.gru_p(p)
q = self.attn_p(h_p)
## smi encoding (day level)
r = torch.zeros(p.shape[0], 0, self.gru_hidden_size)
r = r.to(device)
for t in range(self.T):
h_m, _ = self.gru_m(m[t])
r_t = self.attn_m(h_m, m_mask[t])
r = torch.cat((r, r_t), 1)
## smi encoding (aggregate)
h_s, _ = self.gru_s(r)
c = self.attn_s(h_s)
## blending
x = self.blend(q, c)
## reshaping (eliminating superfluous dimension)
x = x.view(x.shape[0], x.shape[2])
## first gat layer
# first head
sgat = self.mgat_1[0]
z = sgat(x, neighborhoods)
z = self.elu(z)
# remaining heads
for u in range(1, self.U):
sgat = self.mgat_1[u]
z_u = sgat(x, neighborhoods)
z_u = self.elu(z_u)
z = torch.cat((z, z_u), 1)
## second gat layer
# first head
sgat = self.mgat_2[0]
new_z = sgat(z, neighborhoods)
new_z = self.sigmoid(new_z)
# remaining heads
for u in range(1, self.U):
sgat = self.mgat_2[u]
new_z_u = sgat(z, neighborhoods)
new_z_u = self.sigmoid(new_z_u)
new_z = torch.cat((new_z, new_z_u), 1)
## final layer
y = self.sigmoid(self.final_linear(new_z))
## return result
return y
# + [markdown] id="TyyPL-psA4cU"
# #Data Processing
# + id="JZhP9VnQCyZm"
# #!wget https://github.com/yumoxu/stocknet-dataset/archive/master.zip
# + id="aW75dPMQCzQ_"
# #!unzip master.zip
# + id="xqxlViAXArti"
module_url = "https://tfhub.dev/google/universal-sentence-encoder/2" #@param ["https://tfhub.dev/google/universal-sentence-encoder/2", "https://tfhub.dev/google/universal-sentence-encoder-large/3"]
# + id="DRdIAHpeBJvw"
tf.disable_v2_behavior()
tf.compat.v1.disable_eager_execution()
# + id="uK1ab2p7BLDm"
stocknet_dataset_filepath = './stocknet-dataset-master'
train_start_date = '2014-01-01'
train_end_date = '2015-07-31'
val_start_date = '2015-08-01'
val_end_date = '2015-09-30'
test_start_date = '2015-10-01'
test_end_date = '2016-01-01'
# + id="oIRBbhAZy9ls"
def prep_dataset(dataset_filepath, start_date, end_date):
cache = {}
calendar = mcal.get_calendar('NYSE')
def next_trading_day(start_day=None, SAFE_DELTA = 4):
"""Returns the next/previous trading date separated by a certain number of
trading days.
"""
if start_day is None:
start_day = datetime.datetime.utcnow().date()
if start_day in cache:
return cache[start_day]
start = pd.to_datetime(start_day)
end = start + np.timedelta64(SAFE_DELTA, 'D')
business_days = calendar.valid_days(start_date=start, end_date=end)
next_day = business_days[1].date()
next_day = next_day.strftime("%Y-%m-%d")
cache[start_day] = next_day
return next_day
raw_prices_filepath = stocknet_dataset_filepath + '/price/raw'
preprocessed_tweets_filepath = stocknet_dataset_filepath + '/tweet/preprocessed'
company_to_price_df = {}
company_to_tweets = {}
for filename in os.listdir(raw_prices_filepath):
with open(raw_prices_filepath + '/' + filename) as file:
company_name = filename.split('.')[0]
# Not enough data for GMRE
if company_name == 'GMRE':
continue
df = pd.read_csv(file)
df.columns = ['date', 'open', 'high', 'low', 'close', 'adjust_close', 'volume']
mask = (df['date'] >= start_date) & (df['date'] <= end_date)
df = df.loc[mask]
company_to_price_df[company_name] = df.dropna()
for filename in tqdm(os.listdir(preprocessed_tweets_filepath)):
company_name = filename.split('.')[0]
dates_to_tweets = {}
for tweet_filename in os.listdir(preprocessed_tweets_filepath + '/' + filename):
if tweet_filename < start_date or tweet_filename > end_date:
continue
with open(preprocessed_tweets_filepath + '/' + filename + '/' + tweet_filename) as file:
list_of_tweets = []
for line in file:
tweet_json = json.loads(line)
list_of_tweets.append(tweet_json)
date_idx = next_trading_day(tweet_filename)
if date_idx not in dates_to_tweets:
dates_to_tweets[date_idx] = list_of_tweets
else:
dates_to_tweets[date_idx] += list_of_tweets
company_to_tweets[company_name] = dates_to_tweets
# Reduce logging output.
logging.set_verbosity(logging.ERROR)
tf.get_logger().setLevel(logging.ERROR)
tf.compat.v1.logging.set_verbosity(tf.compat.v1.logging.ERROR)
# Import the Universal Sentence Encoder's TF Hub module
def embed_useT(module):
with tf.Graph().as_default():
sentences = tf.placeholder(tf.string)
embed = hub.Module(module)
embeddings = embed(sentences)
session = tf.train.MonitoredSession()
return lambda x: session.run(embeddings, {sentences: x})
embed_fn = embed_useT(module_url)
# Generate embeddings
for company in tqdm(company_to_tweets.keys()):
for date in company_to_tweets[company].keys():
messages = []
for j in range(len(company_to_tweets[company][date])):
messages.append(' '.join(company_to_tweets[company][date][j]['text']))
message_embeddings = embed_fn(messages)
for k in range(len(company_to_tweets[company][date])):
company_to_tweets[company][date][k]['embedding'] = list(message_embeddings[k])
# Create date mapping
date_universe = set()
for company in company_to_price_df.keys():
date_universe = date_universe.union(set(company_to_price_df[company].date))
for company in company_to_tweets.keys():
date_universe = date_universe.union(set(company_to_tweets[company].keys()))
date_universe = sorted(list(date_universe))
index_to_date = {i-5:d for i,d in enumerate(date_universe)}
date_to_index = {d:i-5 for i,d in enumerate(date_universe)}
# Calculate dimensions for tensor
n_stocks = len(company_to_tweets.keys())
n_days = len(date_universe)
max_tweets = 0
for c,d in itertools.product(company_to_tweets.keys(), date_universe):
if d in company_to_tweets[c]:
max_tweets = max(max_tweets, len(company_to_tweets[c][d]))
# Create index mapping for stocks alphabetically
company_to_index = {c:i for i,c in enumerate(sorted(list(company_to_tweets.keys())))}
return company_to_price_df, company_to_tweets, date_universe, n_days, n_stocks, max_tweets
# + id="7VwHfOnq19zW"
train_company_to_price_df, train_company_to_tweets, train_date_universe, train_n_days, train_n_stocks, train_max_tweets = prep_dataset(stocknet_dataset_filepath, train_start_date, train_end_date)
val_company_to_price_df, val_company_to_tweets, val_date_universe, val_n_days, val_n_stocks, val_max_tweets = prep_dataset(stocknet_dataset_filepath, val_start_date, val_end_date)
test_company_to_price_df, test_company_to_tweets, test_date_universe, test_n_days, test_n_stocks, test_max_tweets = prep_dataset(stocknet_dataset_filepath, test_start_date, test_end_date)
# + [markdown] id="Td7nHz4py1wv"
# #Dataset and DataLoader
# + id="8XrZt8s1Kys5"
class StockDataset(Dataset):
"""Price dataset"""
def __init__(self, company_to_price_df, company_to_tweets, date_universe, n_days, n_stocks, max_tweets):
# Initialize class members
self.n_stocks = n_stocks
self.n_days = n_days
self.max_tweets = max_tweets
self.window = 6
window = self.window
# Build maps
self.company_to_index = {c:i for i,c in enumerate(sorted(list(company_to_tweets.keys())))}
self.date_to_index = {d:i for i,d in enumerate(date_universe)}
self.index_to_date = {i:d for i,d in enumerate(date_universe)}
# Store data
self.company_to_price_df = company_to_price_df
self.company_to_tweets = company_to_tweets
# Get price data tensor: n_stocks, n_days, 3
self.price_data = np.zeros((n_stocks, n_days, 3))
for company in company_to_price_df.keys():
df = company_to_price_df[company]
df.reset_index(inplace=True, drop=True)
# Look up specific rows in DF
for index, row in df.iterrows():
# Grab row with particular date
if index != 0:
d_index = self.date_to_index[row['date']]
c_index = self.company_to_index[company]
self.price_data[c_index, d_index, 0] = row['high'] / prev_close
self.price_data[c_index, d_index, 1] = row['low'] / prev_close
self.price_data[c_index, d_index, 2] = row['close'] / prev_close
prev_close = row['close']
# Which stocks are usable for these dates, shape n_days n_stocks
self.usable_stocks = torch.ones((self.n_days-7, self.n_stocks))
# Labels of shape n_days, n_stocks
self.labels = torch.zeros((self.n_days-7, self.n_stocks))
# Get labels
for i in range(self.n_days-7):
# Day after (for label)
day_after = self.index_to_date[i + window + 1]
# Current day
current_day = self.index_to_date[i + window]
for company in self.company_to_price_df.keys():
df = self.company_to_price_df[company]
# Grab row with particular date
post_row = df.loc[df['date'] == day_after]
row = df.loc[df['date'] == current_day]
c_index = self.company_to_index[company]
if (len(post_row['close']) > 0) and (len(row['close']) > 0):
close = np.zeros((1))
close[0] = post_row['close']
close[0] /= row['close']
if close >= 1.0055:
self.labels[i, c_index] = 1
elif close <= 0.995:
self.labels[i, c_index] = 0
else:
self.usable_stocks[i, c_index] = 0
else:
self.usable_stocks[i, c_index] = 0
def __len__(self):
return self.n_days-7
def __getitem__(self, idx):
"""
gets a price tensor of shape (n_stocks, 6, 3)
gets a smi tensor of shape (n_stocks, 6, K, 512)
"""
if torch.is_tensor(idx):
idx = idx.tolist()
# Size of sliding window
window = self.window
# Current day's usable stocks from price filter
usable_stocks = self.usable_stocks[idx]
# Labels from price day
labels = self.labels[idx]
# Dates that we need to look up
dates_range = [self.index_to_date[i] for i in range(idx + 1, idx + window + 1)]
# Day after (for label)
day_after = self.index_to_date[idx + window + 1]
# Current day
current_day = self.index_to_date[idx + window]
# Get price data tensor: n_stocks, window, 3
price_data = self.price_data[:, idx+1:idx+window+1, :]
# Extract tweets for specific window
smi_data = np.zeros((self.n_stocks, window, self.max_tweets, 512))
tweet_counts = np.zeros((self.n_stocks, window))
for company in self.company_to_tweets.keys():
# Look up tweets from specific days
for date_idx, date in enumerate(dates_range):
n_tweets = 0
tweets = []
c_index = self.company_to_index[company]
if date in self.company_to_tweets[company]:
n_tweets = len(self.company_to_tweets[company][date])
tweets = [self.company_to_tweets[company][date][k]['embedding'] for k in range(n_tweets)]
else:
usable_stocks[c_index] = 0
tweet_counts[c_index, date_idx] = n_tweets
if n_tweets == 0:
usable_stocks[c_index] = 0
for i,embedding in enumerate(tweets):
#stocks, day, lags, tweet, embedding
smi_data[c_index, date_idx, i, :] = embedding[:]
usable_stocks = (usable_stocks == 1)
m_mask = torch.zeros(6, self.n_stocks, self.max_tweets, 1)
for t in range(6):
for i in range(self.n_stocks):
m_mask[t, i, 0:int(round(tweet_counts[i][t])), 0] = 1
price_output = price_data[usable_stocks,:,:]
smi_output = smi_data[usable_stocks,:,:,:]
tweet_count = tweet_counts[usable_stocks,:]
m_mask = m_mask[:,usable_stocks,:,:]
labels = labels[usable_stocks]
# construct output
return price_output, smi_output, tweet_count, usable_stocks, labels, m_mask
# + id="8W_Y3YxELDp6"
train_dataset = StockDataset(train_company_to_price_df, train_company_to_tweets, train_date_universe, train_n_days, train_n_stocks, train_max_tweets)
val_dataset = StockDataset(val_company_to_price_df, val_company_to_tweets, val_date_universe, val_n_days, val_n_stocks, val_max_tweets)
test_dataset = StockDataset(test_company_to_price_df, test_company_to_tweets, test_date_universe, test_n_days, test_n_stocks, test_max_tweets)
# + id="qV_GvReJLFM5"
train_dataloader = DataLoader(train_dataset, batch_size=1,
shuffle=True, num_workers=0)
val_dataloader = DataLoader(val_dataset, batch_size=1,
shuffle=False, num_workers=0)
test_dataloader = DataLoader(test_dataset, batch_size=1,
shuffle=False, num_workers=0)
# + [markdown] id="tlOeo8caLWKK"
# #Separator
# + colab={"base_uri": "https://localhost:8080/"} id="ANYU8fO-BdS6" outputId="8a8108c0-2e77-4b7e-ffee-2b12955a7925"
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(device)
# + [markdown] id="tzGtCej6Bu_n"
# #Training
# + id="etylmeyDBtQ4"
mansf = MANSF(T=6,
gru_hidden_size=64,
attn_inter_size=32,
use_embed_size=512,
blend_size=32,
gat_1_inter_size=32,
gat_2_inter_size=32,
leakyrelu_slope=0.01,
elu_alpha=1.0,
U=8)
# + id="eNy0SuSXB0H2"
mansf = mansf.to(device)
# + id="ELS9f6JKCDCU"
optimizer = optim.Adam(mansf.parameters(), lr=5e-4)
loss_fn = nn.BCELoss(reduction='mean')
# + id="TeuL7Yt_shSW"
train_acc_list = []
val_acc_list = []
# + id="GhxL1HqYCEPD"
for epoch in range(18):
mansf.train()
correct = 0.0
total = 0.0
running_loss = 0.0
for price, smi, n_tweets, usable_stocks, labels, m_mask in tqdm(train_dataloader):
price = price.type(torch.FloatTensor)
smi = smi.type(torch.FloatTensor)
price = price.to(device).squeeze(axis=0)
smi = smi.to(device).squeeze(axis=0).permute(1, 0, 2, 3)
n_tweets = n_tweets.to(device).squeeze(axis=0)
usable_stocks = usable_stocks.to(device).squeeze(axis=0)
labels = labels.to(device)
m_mask = m_mask.to(device).squeeze(axis=0)
#print(smi.shape, m_mask.shape)
#print(smi[:, :, :, :1])
print(m_mask)
print(np.ma.masked_where(m_mask.cpu() == False, smi[:, :, :, :1].cpu()))
m = []
for t in range(6):
m.append(smi[t])
neighborhoods = torch.eye(87, 87)
neighborhoods = neighborhoods.to(device)
neighborhoods = neighborhoods[usable_stocks, :]
neighborhoods = neighborhoods[:, usable_stocks]
if price.shape[0] != 0:
y = mansf(price, smi, m_mask, neighborhoods)
loss = loss_fn(y.view(-1), labels.view(-1))
loss.backward()
optimizer.step()
optimizer.zero_grad()
correct += torch.sum(((y > 0.5).view(-1) == labels.view(-1))).item()
total += len(y)
running_loss = loss.item() * len(y)
train_acc = correct / total
train_acc_list.append(train_acc)
mansf.eval()
correct = 0.0
total = 0.0
for price, smi, n_tweets, usable_stocks, labels, m_mask in tqdm(val_dataloader):
price = price.type(torch.FloatTensor)
smi = smi.type(torch.FloatTensor)
price = price.to(device).squeeze(axis=0)
smi = smi.to(device).squeeze(axis=0).permute(1, 0, 2, 3)
n_tweets = n_tweets.to(device).squeeze(axis=0)
usable_stocks = usable_stocks.to(device).squeeze(axis=0)
labels = labels.to(device)
m_mask = m_mask.to(device).squeeze(axis=0)
m = []
for t in range(6):
m.append(smi[t])
neighborhoods = torch.eye(87, 87)
neighborhoods = neighborhoods.to(device)
neighborhoods = neighborhoods[usable_stocks, :]
neighborhoods = neighborhoods[:, usable_stocks]
if price.shape[0] != 0:
y = mansf(price, smi, m_mask, neighborhoods)
correct += torch.sum((y > 0.5).view(-1) == labels.view(-1)).item()
total += len(y)
val_acc = correct / total
val_acc_list.append(val_acc)
print('epoch:', epoch, 'loss:', running_loss, 'train_acc:', train_acc, 'val_acc:', val_acc)
# + id="cxKSr2BV4k_z"
mansf.eval()
price, smi, n_tweets, usable_stocks, labels, m_mask = next(iter(val_dataloader))
price = price.type(torch.FloatTensor)
smi = smi.type(torch.FloatTensor)
price = price.to(device)
smi = smi.to(device)
n_tweets = n_tweets.to(device)
usable_stocks = usable_stocks.to(device)
labels = labels.to(device)
m_mask = m_mask.to(device)
price = price.view(price.shape[1], price.shape[2], price.shape[3])
smi = smi.view(smi.shape[1], smi.shape[2], smi.shape[3], smi.shape[4])
n_tweets = n_tweets.view(n_tweets.shape[1], n_tweets.shape[2])
usable_stocks = usable_stocks.view(usable_stocks.shape[1])
m_mask = m_mask.view(m_mask.shape[1], m_mask.shape[2], m_mask.shape[3], m_mask.shape[4])
smi = smi.permute(1, 0, 2, 3)
m = []
for t in range(6):
m.append(smi[t])
neighborhoods = torch.eye(87, 87)
neighborhoods = neighborhoods.to(device)
neighborhoods = neighborhoods[usable_stocks, :]
neighborhoods = neighborhoods[:, usable_stocks]
y = mansf(price, smi, m_mask, neighborhoods)
# + colab={"base_uri": "https://localhost:8080/"} id="XqAGN_ND12G4" outputId="8a415adf-a839-4632-d3c4-32a8bd296471"
print(y)
# + colab={"base_uri": "https://localhost:8080/"} id="j2S1py5m47Ts" outputId="eb6cc1ee-369c-41b6-e019-c8b57d1e9148"
print(labels)
# + [markdown] id="64uHrKiQrq4X"
# #Figures
# + id="yGV8hlffrr9k"
def plot(X, Y, xlabel, ylabel, legend, title):
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1)
for i in range(len(Y)):
ax.plot(X, Y[i], label=legend[i])
plt.grid(color='0.95')
plt.legend()
ax.set(xlabel=xlabel, ylabel=ylabel, title=title)
# + id="M6pHZ4GIr1BI"
"""plot(range(18),
[train_acc_list, val_acc_list],
'epoch',
'accuracy',
['training accuracy', 'validation accuracy'],
'accuracy vs. epoch')"""
| model_notebooks/testing_speedup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import dgl
import torch
import torch.nn as nn
import torch.nn.functional as F
# +
# Generate a synthetic dataset with 10000 graphs, ranging from 10 to 500 nodes
import dgl.data
dataset = dgl.data.GINDataset('PROTEINS', self_loop=True)
print('Node feature dimensionality:', dataset.dim_nfeats)
print('Number of graph categories:', dataset.gclasses)
# +
#Load data with mini-batches
from dgl.dataloading import GraphDataLoader
from torch.utils.data.sampler import SubsetRandomSampler
num_samples = len(dataset)
num_train_samples = int(0.8 * num_samples)
num_valid_samples = int(0.1 * num_samples)
num_test_samples = int(0.1 * num_samples)
# Choose a data sampler
# There are many other options, see details below:
# https://pytorch.org/docs/stable/data.html#data-loading-order-and-sampler
train_sampler = SubsetRandomSampler(torch.arange(num_train_samples))
valid_sampler = SubsetRandomSampler(torch.arange(num_train_samples, num_train_samples+num_valid_samples))
test_sampler = SubsetRandomSampler(torch.arange(num_train_samples+num_valid_samples, num_train_samples+num_valid_samples+num_test_samples))
train_loader = GraphDataLoader(
dataset, sampler=train_sampler, batch_size=16, drop_last=False)
valid_loader = GraphDataLoader(
dataset, sampler=valid_sampler, batch_size=16, drop_last=False)
test_loader = GraphDataLoader(
dataset, sampler=test_sampler, batch_size=16, drop_last=False)
# Check datapoint
item = iter(train_loader)
batch = next(item)
print(batch)
# +
graph_batch, labels = batch
print('Number of nodes for each graph:', graph_batch.batch_num_nodes())
print('Number of edges for each graph:', graph_batch.batch_num_edges())
# Reconstruct original graphs in a mini-batch
ori_graphs = dgl.unbatch(graph_batch)
print('The original graphs in the minibatch:', ori_graphs)
# +
# Build models
from dgl.nn import GraphConv, GATConv, GINConv
# Graph Convolutional Networks
# TODO: more options for graph pooling
class GCN(nn.Module):
def __init__(self, in_feats, h_feats, num_classes):
super(GCN, self).__init__()
self.conv1 = GraphConv(in_feats, h_feats)
self.conv2 = GraphConv(h_feats, num_classes)
def forward(self, g, in_feat):
h = self.conv1(g, in_feat)
h = F.relu(h)
h = self.conv2(g, h)
g.ndata['h'] = h
return dgl.mean_nodes(g, 'h')
# Graph Attention Networks
class GAT(nn.Module):
def __init__(self, in_feats, h_feats, num_classes):
super(GAT, self).__init__()
self.conv1 = GATConv(in_feats, h_feats, num_heads=3)
self.conv2 = GATConv(h_feats, num_classes, num_heads=3)
def forward(self, g, in_feat):
h = self.conv1(g, in_feat)
h = F.relu(h)
h = torch.mean(h, dim=1) # Average value over three attention heads
h = self.conv2(g, h)
h = torch.mean(h, dim=1) # Average value over three attention heads
g.ndata['h'] = h
return dgl.mean_nodes(g, 'h')
# Graph Isomorphism Networks
class GIN(nn.Module):
def __init__(self, in_feats, h_feats, num_classes):
super(GIN, self).__init__()
lin1 = torch.nn.Linear(in_feats, h_feats)
lin2 = torch.nn.Linear(h_feats, num_classes)
self.conv1 = GINConv(lin1, 'sum')
self.conv2 = GINConv(lin2, 'sum')
def forward(self, g, in_feat):
h = self.conv1(g, in_feat)
h = F.relu(h)
h = self.conv2(g, h)
g.ndata['h'] = h
return dgl.mean_nodes(g, 'h')
# +
# Train models
model_gcn = GCN(dataset.dim_nfeats, 16, dataset.gclasses)
optimizer = torch.optim.Adam(model_gcn.parameters(), lr=0.01)
for epoch in range(20):
for graph_batch, labels in train_loader:
pred = model_gcn(graph_batch, graph_batch.ndata['attr'].float())
loss = F.cross_entropy(pred, labels)
optimizer.zero_grad()
loss.backward()
optimizer.step()
num_correct = 0
num_valids = 0
for graph_batch, labels in valid_loader:
pred = model_gcn(graph_batch, graph_batch.ndata['attr'].float())
num_correct += (pred.argmax(1) == labels).sum().item()
num_valids += len(labels)
print('Valid accuracy:', num_correct / num_valids)
num_correct = 0
num_tests = 0
for graph_batch, labels in test_loader:
pred = model_gcn(graph_batch, graph_batch.ndata['attr'].float())
num_correct += (pred.argmax(1) == labels).sum().item()
num_tests += len(labels)
print('Test accuracy:', num_correct / num_tests)
# +
# Train GAT model
model_gat = GAT(dataset.dim_nfeats, 16, dataset.gclasses)
optimizer = torch.optim.Adam(model_gat.parameters(), lr=0.01)
for epoch in range(20):
for graph_batch, labels in train_loader:
pred = model_gat(graph_batch, graph_batch.ndata['attr'].float())
loss = F.cross_entropy(pred, labels)
optimizer.zero_grad()
loss.backward()
optimizer.step()
num_correct = 0
num_valids = 0
for graph_batch, labels in valid_loader:
pred = model_gat(graph_batch, graph_batch.ndata['attr'].float())
num_correct += (pred.argmax(1) == labels).sum().item()
num_valids += len(labels)
print('Valid accuracy:', num_correct / num_valids)
num_correct = 0
num_tests = 0
for graph_batch, labels in test_loader:
pred = model_gat(graph_batch, graph_batch.ndata['attr'].float())
num_correct += (pred.argmax(1) == labels).sum().item()
num_tests += len(labels)
print('Test accuracy:', num_correct / num_tests)
# +
# Train GIN model
model_gin = GIN(dataset.dim_nfeats, 16, dataset.gclasses)
optimizer = torch.optim.Adam(model_gin.parameters(), lr=0.01)
for epoch in range(20):
for graph_batch, labels in train_loader:
pred = model_gin(graph_batch, graph_batch.ndata['attr'].float())
loss = F.cross_entropy(pred, labels)
optimizer.zero_grad()
loss.backward()
optimizer.step()
num_correct = 0
num_valids = 0
for graph_batch, labels in valid_loader:
pred = model_gin(graph_batch, graph_batch.ndata['attr'].float())
num_correct += (pred.argmax(1) == labels).sum().item()
num_valids += len(labels)
print('Valid accuracy:', num_correct / num_valids)
num_correct = 0
num_tests = 0
for graph_batch, labels in test_loader:
pred = model_gin(graph_batch, graph_batch.ndata['attr'].float())
num_correct += (pred.argmax(1) == labels).sum().item()
num_tests += len(labels)
print('Test accuracy:', num_correct / num_tests)
| codes/models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Worksheet 0.0.2: Getting comfortable with the terminal
#
# <div class="alert alert-block alert-warning">
# <p>This activity only works if we're in our JupyterLab environment. If you haven't finished the <a href = '1_week-0-worksheet-github-clone.md'>"Cloning" a repository</a> worksheet, please do so now.
# </div>
#
# We'll gain some experience with our terminal and Jupyter notebooks in this activity.
#
# ## Table of contents
# ---
#
# * [Understanding the terminal](#Understanding-the-terminal)
# * [Basic commands](#Basic-commands)
# * [Finishing this activity](#Finishing-this-activity)
#
# ## Understanding the terminal
#
# ---
#
# <div class="alert alert-block alert-warning">
# <p>If you do not have a terminal tab open in your JupyterLab workspace, open one now.</p>
# <p>Need a refresher on how to do that? <a href = "../README.md#Using-our-terminal">Check out this part of the README</a></p>
# </div>
#
# In our [README](../README.md), we encountered a bit of history about the terminal. Here, we're more concerned with understanding _what the terminal is telling us_.
#
# We can tell a bit from the following example:
#
# ```
# dluman@jupyter-cs-allegheny-edu:~$
# ```
#
# Here, we see the anatomy of a basic **command prompt** (the line up to, and including the `$` sign). Here's a breakdown of the parts:
#
# |Text |Meaning |
# |--------------|----------------------|
# |dluman | User name |
# |jupyter-cs-allegheny-edu| Server name|
# |~ | User `dluman` is in their "home" directory|
#
# We can tell a bit from this prompt. Keep track of how it will change as we move through this exercise.
#
# ## Basic commands
#
# ---
#
# In this worksheet, we'll run through:
#
# * `cd`
# * `pwd`
# * `ls`
# <div class="alert alert-block alert-info">
# <p>
# Run each of the commands in the empty space provided below the question/prompt. These are referred to as "cells" in JupyterLab vocabulary. When you're finished typing a command, press the <b>SHIFT</b> + <b>ENTER</b> keys to run it.
# </p>
# <p>
# Also -- important note -- in this notebook, stick to <em>one command per cell</em> -- don't get fancy on me. (It won't work.) This is not like code -- when we use code in JupyterLab, we can combine it all together in a big ol' block. But, here, Jupyter gets confused if we try too much.
# </p>
# </div>
#
# #### 1. Home is getting kind-of boring: let's go somewhere else -- type `cd /mansion`
# +
# TODO
# -
# `mansion` is just a directory that we have on our system -- one that lives at the _lowest level_ of our server's hard drive -- that's why there's only a `/` in front of it.
#
# #### 2. What is this run-down mansion? Let's have a look around. Type `ls`.
# +
# TODO
# -
# #### 3. 👻 It's a ghost! Let's run for it: go back `home` by typing `cd ~`
# +
# TODO
# -
# #### 4. Ok. We're back `~`. If we use the `pwd` command, we can further prove it to ourselves. Type it below.
# +
# TODO
# -
# #### 5. Do the outputs from the two commands match? That means you are, in fact `~`. Let's look around again and see what we can see: type `ls`
# +
# TODO
# -
# #### 6. We see that we have our `cmpsc-100-jan-2021-week-0-YOUR USER NAME` directory. Let's `cd` to it.
#
# <div class="alert alert-block alert-info">
# <p><b>Tip:</b> A lot of the directories and file names we use in our class are pretty long. Blame your instructor for that. But there's a trick we can use.
# </p>
# <p>
# If you know (or want to complete) only _part_ of the name of the destination or file, you can "completion match" on the rest of the name by pressing the <b>TAB</b> key.
# </p>
# </div>
#
# Type `cd cmpsc` and then the `TAB` key and run the command.
# +
# TODO
# -
# Nifty, right? It'll save you a lot of time and typos. Or, you can just save your typos for other things. Like code.
#
# #### 7. Let's look around again and see what we see. What's the command for that? Type and run it in the space below.
# +
# TODO
# -
# #### 8. Hm. Bunch of stuff here.
#
# * The items appended with a `/` are directories (just like `/mansion` and just like `cmpsc-100-jan-2021...`)
# * The other stuff: files of no particular importance to you. Nope. Don't mean anything...yet.
#
# Let's snoop on ourselves. Move into the `day-1` directory and look around. (I've provided you with two cells to run the commands separately.)
# +
# TODO
# +
# TODO
# -
# #### 9. This is all well and good, but there are other "rooms" in this repository -- like other days, including our lab day, `day-3`. How can we get there from here?
#
# <div class="alert alert-block alert-info">
# <p>
# <b>Tip:</b> what goes up can also come down.
# </p>
# <p>
# Using <b>..</b> after our <b>cd</b> commands allows us to return down a single level. Later, we'll see that they can be <em>chained</em>.
# </p>
# </div>
#
# Let's go down a level (back a "step"). Type and run `cd ..` in the space below.
# +
# TODO
# -
# #### 10. We're back where we were a minute ago. Let's go into the `day-3` and back home using some of our new tricks.
#
# This is a multi-step process:
#
# 1. In the first cell, enter the `day-3` folder
# 2. In the second cell, go back home by using `cd ../../`
# * `Tidbit`: "down" or "back" commands can be _chained together_ to perform mutliple steps at once
# 3. In the third cell, verify you're home by using `pwd`
#
# If the output to `2` and `3` are the same -- you're done!
# +
# TODO
# +
# TODO
# +
# TODO
# -
# ## Finishing this activity
#
# If you'd like some _direct_ practice with these commands, you can run through them in your terminal tab. Don't worry: with `cd`, `pwd`, and `ls`, you won't hurt anything. Take some time to do this now, and you'll thank yourself later.
| day-0/platforms/2_week-0-worksheet-basic-terminal.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CS 109A/STAT 121A/AC 209A/CSCI E-109A:
# # Midterm - 2017
#
# **Harvard University**<br/>
# **Fall 2017**<br/>
# **Instructors**: <NAME>, <NAME>, <NAME>, <NAME>
#
# ---
#
# ### INSTRUCTIONS
#
#
# - You must submit the Midterm on your own. ** No group submissions are allowed**. You may use any print or online resources but ** you may not work or consult with others**.
# - Restart the kernel and run the whole notebook again before you submit.
# - Please submit both a notebook and a pdf.
#
#
# ---
#
# ## Flight Delays
#
# The U.S. Department of Transportation's (DOT) Bureau of Transportation Statistics tracks the on-time performance of domestic flights operated by large air carriers. Summary information on the number of on-time, delayed, canceled, and diverted flights are published in DOT's monthly Air Travel Consumer Report and in this dataset of 2015 flight delays and cancellations.
#
#
# ## Data
# Each entry of the flights.csv file corresponds to a flight. More than 5,800,000 flights were recorded in 2015. These flights are described according to 31 variables. Further details of these variables can be found <a href='https://www.transtats.bts.gov/DL_SelectFields.asp?Table_ID=236&DB_Short_Name=On-Time'>here</a>, if you are interested (not needed to answer these questions).
#
#
#
#
# | Name | Type | DESCRIPTION |
# |--------------------------------|---|----------------------------------------------------------------------|
# | DATE | object | The date in python datetime format |
# | MONTH | int64 | The month of the year(1-12) |
# | DAY | int64 | The day of the month |
# | DAY_OF_WEEK | int64 | The day of the week(1-7, MON-SUN) |
# | AIRLINE | object | An identifier for the airline |
# | FLIGHT_NUMBER | int64 | The flight number |
# | TAIL_NUMBER | object | The tail number (aircraft) corresponding to this flight |
# | ORIGIN_AIRPORT | object | The code for origin airport |
# | DESTINATION_AIRPORT | object | The code for destination airport |
# | SCHED_DEP | object | The departure time in python datetime.time format |
# | SCHED_ARR | object | The arrival time in python datetime.time format |
# | DEPARTURE_DELAY | float64| The delay incurred at the origin (mins) |
# | ARRIVAL_DELAY | float64 | The delay when the flight reached the (mins) destination |
# | DISTANCE | int64 | Distance in miles between origin and destination |
# | SCHEDULED_TIME | float64 | Scheduled time of flight (minutes) |
# | ELAPSED_TIME | float64 | Actual time of flight (minutes) |
# | AIR_SYSTEM_DELAY | float64 | What part of the delay was NASD?(mins) |
# | SECURITY_DELAY | float64 | What part of the delay was due to security problems? (mins) |
# | AIRLINE_DELAY | float64 | What part of the delay is due to the airline? (mins) |
# | LATE_AIRCRAFT_DELAY | float64 | What part of the delay is due to previous flight(s) being late(mins) |
# | WEATHER_DELAY | float64 | Delay due to extreme weather events(min) |
#
# You can read more about the various weather delays [here](https://www.rita.dot.gov/bts/help/aviation/html/understanding.html) if you are so inclined.
#
#
#
# ## Data/Caveats
#
# The data file, flights.csv, is found <a href="https://drive.google.com/file/d/0B9dVesTppCgHY0IwZHk3SGhjd00/view?usp=sharing">here</a> (note, it is about 70MB).
#
# This data is already preprocessed, reduced, partially cleaned and therefore not identical to the original dataset.
import numpy as np
import pandas as pd
from datetime import datetime
import time
import matplotlib
import matplotlib.pyplot as plt
from sklearn.pipeline import Pipeline
from sklearn.linear_model import LogisticRegression
from sklearn.linear_model import LogisticRegressionCV
import sklearn.metrics as metrics
from sklearn.preprocessing import PolynomialFeatures
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis as QDA
from sklearn.neighbors import KNeighborsClassifier as KNN
from sklearn.tree import DecisionTreeClassifier as DecisionTree
from sklearn.model_selection import cross_val_score
from sklearn.metrics import accuracy_score
from sklearn import preprocessing
from sklearn.tree import export_graphviz
from IPython.display import Image
from IPython.display import display
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import LassoCV
from sklearn.linear_model import RidgeCV
from sklearn.decomposition import PCA
from sklearn.model_selection import train_test_split
import warnings
warnings.filterwarnings("ignore")
# %matplotlib inline
# ## Problem Description
# We will build two separate models: one model that classifies whether a flight will be delayed and a second model that predicts the length of delay given that a flight is truly delayed. Only consider models taught in class so far.
#
# ** Consider the following: **
# This is a large dataset; think of strategies on how to solve this problem. Create a manageable subsample of the data that you can use to train and test/validate, but eventually you should predict on all the data (excluding the training set).
#
# ### Questions
# 1. (5pts) Create a new variable, `DELAY_OR_NOT`: a boolean/indicator variable which indicates any arrival delay under 15 mins as a 0, and any delay at or above 15 mins as a 1 (`ARRIVAL_DELAY >= 15`).
# 2. (5pts) Make sure you understand the data variable descriptions before you start the analysis. Consider all the columns and determine and list which of these predictors should not be used.
# 3. (15pts) Perform EDA to gain intuition of the factors that affect delay and provide visuals: do delays vary across airlines, or time of departure, or airport (do, at the very least, Chicago (ORD), Boston (BOS), and your favorite another airport), or airport traffic?
# 4. (20pts) Build a classification model that classifies delays according to `DELAY_OR_NOT`. This is an unbalanced dataset, thus consider the appropriate performance metric when reporting your results.
# 5. (5pts) Given your model, comment on the importance of factors as related to whether a flight is delayed.
# 6. (5pts) Evaluate your model(s) on your test set, and finally provide a visual to show which airlines are predicted to have the most delays using all the data excluding the training and test set.
# 7. (15pts) Build a regression model that predicts the length of delay (on the log scale) given that a flight is truly delayed.
# 8. (20pts) Write a report (in the last markdown cell in your notebook with your findings (without code)). Describe the main design decisions you have made with justifications. Clearly explain your methodology and results. This should not be more than 300 words. You may use up to 5 diagrams.
#
#
# **Question 1**<br>
# (5pts) Create a new variable, DELAY_OR_NOT: a boolean/indicator variable which indicates any arrival delay under 15 mins as a 0, and any delay at or above 15 mins as a 1 (ARRIVAL_DELAY >= 15).
#Load the data
df = pd.read_csv("cs109a_midterm.csv")
#Display the first couple columns
df.head()
#Let's first clean our dataframe
#See where the missing values are
cols_missing_values = df.columns[df.isnull().any()]
cols_missing_values
#Let's see if we have any missing values in the 'ARRIVAL_DELAY' column
np.sum([df.ARRIVAL_DELAY.isnull()])
#Create the variable 'DELAY_OR_NOT' in our DataFrame based on if 'ARRIVAL_DELAY_>=15 (min)
#Since we saw we do not have any missing values in ARRIVAL_DELAY column, we can proceed.
DELAY_OR_NOT = (df.ARRIVAL_DELAY >= 15).astype(int)
df['DELAY_OR_NOT'] = DELAY_OR_NOT
# **Question 2**<br>(5pts) Make sure you understand the data variable descriptions before you start the analysis. Consider all the columns and determine and list which of these predictors should not be used.
# **I believe there is several columns which do not help in our analysis**
#
# # DATE
# since we already have variables MONTH and DAY and we know all the data points are from 2015
# # SCHEDULED_TIME
# is duplicate information since SCHEDULED_TIME = SCHED_ARRIVAL - SCHED_DEPARTURE
# # ELAPSED_TIME
# is duplicate information since ELAPSED_TIME = SCHED_ARRIVAL + ARRIVAL_DELAY - (SCHED_DEPARTURE + DEPARTURE DELAY)
# # FLIGHT NUMBER
# It is redundant since we have info about origin/destination and scheduled dep/arr time
# # TAIL NUMBER
# This is indicative of each unique plane. When we have a dataset with 800,000 observations and 4819 planes, it is very unlikely our regression model would be able to recognize if there is a particular plane which causes delay.
#
# # All the delay features
# Given the fact we already have a "delay_or_not" response we are seeking, our "types" of delays are not predictors of whether there will be a delay or not. In other words, a type of "delay" will predict the length of the delay but not whether there will be an event of a delay.
# **So, we will get rid of those in our analysis in this subproblem. I already looked for missing values in the previous subquestion and the missing values happen to be only in the columns I am dropping.**
# +
#We copy the dataframe, since it's 2017 and we all have enough memory,right?
new_df = df.copy(deep=True)
#drop all the columns based on the analysis in the cell above
del new_df['DATE']
del new_df['SCHEDULED_TIME']
del new_df['ELAPSED_TIME']
del new_df['FLIGHT_NUMBER']
del new_df['TAIL_NUMBER']
del new_df['ARRIVAL_DELAY']
del new_df['DEPARTURE_DELAY']
del new_df['AIR_SYSTEM_DELAY']
del new_df['AIRLINE_DELAY']
del new_df['SECURITY_DELAY']
del new_df['LATE_AIRCRAFT_DELAY']
del new_df['WEATHER_DELAY']
# -
# **Question 3** <br>
# (15pts) Perform EDA to gain intuition of the factors that affect delay and provide visuals: do delays vary across airlines, or time of departure, or airport (do, at the very least, Chicago (ORD), Boston (BOS), and your favorite another airport), or airport traffic?
# +
#We will ONLY look at the visualisation in the training data we will later use to build our model
#(I am splitting train/test here just for purposes for visualisation, I am later using the same random seed
#to do it again after preproccessing)
#Subsample the data to reduce the # of observations.
np.random.seed(9001)
new_df_for_viz = new_df.sample(n=100000,random_state=6)
#Split the data in test and training (see Lecture 5)
train_df_viz, data_test_NOT_USE = train_test_split(new_df_for_viz,test_size=0.33, random_state=0)
# +
#Let's see what airlines had the highest percentage of delays in 2015
list_of_airlines = [i for i in train_df_viz.AIRLINE.value_counts().index]
airlines_delays = []
for i in list_of_airlines:
delayed = train_df_viz[train_df_viz.AIRLINE==i][train_df_viz.DELAY_OR_NOT==1].shape[0]
not_delayed = train_df_viz[train_df_viz.AIRLINE==i][train_df_viz.DELAY_OR_NOT==0].shape[0]
airlines_delays.append(delayed/(delayed+not_delayed))
plt.figure(figsize=(20,10))
plt.bar([i for i in range(len(airlines_delays))],airlines_delays,color='red',label = "Percentage of delayed flights by Airline")
plt.xticks([i for i in range(len(airlines_delays))],list_of_airlines);
mean_delays=np.mean(airlines_delays)
plt.axhline(mean_delays,label='Mean percentage of delay accross airlines')
plt.legend();
plt.xlabel("Airline")
plt.ylabel("Ratio of delayed flights");
# -
# **We see significant differences in the ratio of delayed flights among airlines. For example, "HA" seems to have relatively less delayed flights than "NK"**
#Let's see which airline had the most delays in 2015 as well as how many flights were not delayed for that airlines (total number)
import seaborn as sns
plt.figure(figsize=(20,10))
sns.set(style="darkgrid")
sns.countplot(x='AIRLINE',hue='DELAY_OR_NOT',data=train_df_viz,palette="Set2");
# **This graph is related to the one above. However, we are just plotting the absolute number of flights here.**
# +
#Let's see what days of the week had the most delays in 2015
weekday_delays = []
for i in range(1,8):
delays = train_df_viz[train_df_viz.DAY_OF_WEEK==i][train_df_viz.DELAY_OR_NOT==1].shape[0]
total = train_df_viz[train_df_viz.DAY_OF_WEEK==i].shape[0]
weekday_delays.append(delays/total)
plt.figure(figsize=(20,10))
plt.bar([i for i in range(7)],weekday_delays,color='green',label = "Delay in a weekday")
plt.xticks([i for i in range(7)],['Monday','Tuesday','Wednesday','Thursday','Friday','Saturday','Sunday']);
mean_delays=np.mean(weekday_delays)
plt.axhline(mean_delays,label='Mean delay across weekdays',color='gold')
plt.legend();
plt.xlabel("Day of the week")
plt.ylabel("Percentage of delays");
# -
# **This visualisation indicates that certain days have a higher ratio of delayed flights than others. For example, Thursdays have the highest percentage of delayed flights while Saturdays the smallest.**
#Let's visualize the total delays per day of the week
sns.set(style="darkgrid")
plt.figure(figsize=(20,10))
sns.countplot(x='DAY_OF_WEEK',hue='DELAY_OR_NOT',data=train_df_viz,palette="Set1")
plt.xticks([i for i in range(7)],['Monday','Tuesday','Wednesday','Thursday','Friday','Saturday','Sunday']);
# **This graph is related to the one above. However, we are just plotting the absolute number of flights per day of the week here.**
#
#Let's take a look at how many flights in total were delayed in each month
sns.set(style="darkgrid")
plt.figure(figsize=(20,10))
sns.countplot(x='MONTH',data=train_df_viz[train_df_viz.DELAY_OR_NOT==1],palette="Set1");
# **In this graph, we plot the absolute number of delayed flights in any given month. We see that the lowest number of delayed flights is in September/October, while the most delayed flights are in June/July timeframe.**
#Let's take a look at the percentage of flights delayed in each month
months = train_df_viz.groupby('MONTH').mean()
plt.figure(figsize=(20,10))
plt.bar(months.index,months.DELAY_OR_NOT,color='gray',label = "Percentage delay in a month")
plt.axhline(mean_delays,label='Mean percentage delay across months',color='gold')
plt.legend(bbox_to_anchor=(1, 1));
plt.xlabel("Month")
plt.ylabel("Percentage of delays");
# **Here, we plot the ratio of flights delayed each month. The trends reflect what the previous graph with absolute numbers displayed.**
# +
#Number of delayed/not delayed flights departing every hour, also showing the overall traffic at the airports at a given hour
from datetime import datetime
import time
# string to time tuple
date_str = list(train_df_viz.SCHED_DEP)
time_tuples = []
for i in date_str:
time_tuple = time.strptime(i, "%H:%M:%S")
time_tuples.append(time_tuple.tm_hour)
train_df_viz['hour_of_dep'] = np.array(time_tuples)
plt.figure(figsize=(20,10))
sns.countplot(x='hour_of_dep',hue='DELAY_OR_NOT',data=train_df_viz,palette="Set1");
plt.xlabel("hour of the day");
# -
# **Here we can see the highest number of flights departs in the morning while being the lest delayed, while evening flights are the most delayed and there is less of them than in the morning hours**
# +
#Now, let's plot the percentage of delayed flights dependent on the time departure
hour_delays=[]
for i in range(24):
delays = train_df_viz[train_df_viz.hour_of_dep==i][train_df_viz.DELAY_OR_NOT==1].shape[0]
total = train_df_viz[train_df_viz.hour_of_dep==i].shape[0]
hour_delays.append(delays/total)
plt.figure(figsize=(20,10))
plt.bar([i for i in range(24)],hour_delays,color='gray',label = "Percentage delay in a month")
plt.xticks([i for i in range(24)],[i for i in range(24)]);
mean_delays=np.mean(weekday_delays)
plt.axhline(mean_delays,label='Mean percentage delay across day hours',color='gold')
plt.legend();
plt.xlabel("Hour of the day")
plt.ylabel("Percentage of delays")
del train_df_viz['hour_of_dep'];
# -
# **Seems like most flights are delayed in the evening hours with the peak around 8pm, while morning flights have the least delays.**
# +
#Let's look at delays at 15 most busy airports (from left as the busiest)
frequent_airports = train_df_viz.ORIGIN_AIRPORT.value_counts().index[0:15]
airport_delays = []
for i in frequent_airports:
delays = train_df_viz[train_df_viz.ORIGIN_AIRPORT==i][train_df_viz.DELAY_OR_NOT==1].shape[0]
total = train_df_viz[train_df_viz.ORIGIN_AIRPORT==i].shape[0]
airport_delays.append(delays/total)
plt.figure(figsize=(20,10))
plt.bar([i for i in range(len(frequent_airports))],airport_delays,color='orange',label = "Percentage delay origin aiport")
plt.xticks([i for i in range(len(frequent_airports))],[i for i in frequent_airports]);
mean_delays=np.mean(weekday_delays)
plt.axhline(mean_delays,label='Mean percentage delay across {} most frequent airports'.format(len(frequent_airports)),color='black')
plt.legend();
plt.xlabel("Origin airport")
plt.ylabel("Percentage of delays");
# -
# **Here, we look at the 15 busiest airports (defined as how many flights depart from it in total in 2015). We can see that ORD or MCO have a high percentage of delayed flights while ATL or SEA have a low percentage of delayed flights.**
# **Question 4**<br>
# (20pts) Build a classification model that classifies delays according to DELAY_OR_NOT. This is an unbalanced dataset, thus consider the appropriate performance metric when reporting your results.
# +
#Now, we need to convert our categorical variables
cols_to_change =['MONTH','DAY','DAY_OF_WEEK','AIRLINE','ORIGIN_AIRPORT','DESTINATION_AIRPORT']
#hot-encode
new_df_dummy = pd.get_dummies(new_df,columns=cols_to_change)
#Subsample the data to reduce the # of observations.
new_df_sample = new_df_dummy.sample(n=100000,random_state=6)
#Split the data in test and training (see Lecture 5)
np.random.seed(9001)
data_train, data_test = train_test_split(new_df_sample,test_size=0.33, random_state=0)
#Check we actually got the right shapes
print(new_df_sample.shape,df.shape,data_train.shape,data_test.shape)
# -
#now create a remaining df to test in Question 6
remaining_df = new_df_dummy.drop(new_df_sample.index)
# **Change the scheduled time of dep and arr to decimals**
# string to time tuple TRAIN
date_str_dep = list(data_train.SCHED_DEP)
date_str_arr = list(data_train.SCHED_ARR)
time_tuples_dep = []
time_tuples_arr = []
for i,j in zip(date_str_dep,date_str_arr):
time_tuple_dep = time.strptime(i, "%H:%M:%S")
time_tuple_arr = time.strptime(j, "%H:%M:%S")
time_tuples_dep.append(time_tuple_dep.tm_hour+time_tuple_dep.tm_min/60)
time_tuples_arr.append(time_tuple_arr.tm_hour+time_tuple_arr.tm_min/60)
data_train['SCHED_DEP'] = np.array(time_tuples_dep)
data_train['SCHED_ARR'] = np.array(time_tuples_arr)
# string to time tuple TEST
date_str_dep = list(data_test.SCHED_DEP)
date_str_arr = list(data_test.SCHED_ARR)
time_tuples_dep = []
time_tuples_arr = []
for i,j in zip(date_str_dep,date_str_arr):
time_tuple_dep = time.strptime(i, "%H:%M:%S")
time_tuple_arr = time.strptime(j, "%H:%M:%S")
time_tuples_dep.append(time_tuple_dep.tm_hour+time_tuple_dep.tm_min/60)
time_tuples_arr.append(time_tuple_arr.tm_hour+time_tuple_arr.tm_min/60)
data_test['SCHED_DEP'] = np.array(time_tuples_dep)
data_test['SCHED_ARR'] = np.array(time_tuples_arr)
# string to time REMAINING df
date_str_dep = list(remaining_df.SCHED_DEP)
date_str_arr = list(remaining_df.SCHED_ARR)
time_tuples_dep = []
time_tuples_arr = []
for i,j in zip(date_str_dep,date_str_arr):
time_tuple_dep = time.strptime(i, "%H:%M:%S")
time_tuple_arr = time.strptime(j, "%H:%M:%S")
time_tuples_dep.append(time_tuple_dep.tm_hour+time_tuple_dep.tm_min/60)
time_tuples_arr.append(time_tuple_arr.tm_hour+time_tuple_arr.tm_min/60)
remaining_df['SCHED_DEP'] = np.array(time_tuples_dep)
remaining_df['SCHED_ARR'] = np.array(time_tuples_arr)
#check we converted time to decimals correctly in SCHED_DEP and SCHED_ARR features
data_train.head()
# **Let's take a look at how unbalanced our data (delayed/not delayed) is - I see that only ~10% of observations are delayed flights.**
print(data_train[data_train.DELAY_OR_NOT==1].shape)
print(data_train[data_train.DELAY_OR_NOT==0].shape)
# **Finally, before I start building out models, I need to standardize all of our continuous variables**
# +
p_data_train=data_train.copy(deep=True)
p_data_train.loc[:,"SCHED_DEP":"DISTANCE"]=(data_train.loc[:,"SCHED_DEP":"DISTANCE"]-data_train.loc[:,"SCHED_DEP":"DISTANCE"].mean())/data_train.loc[:,"SCHED_DEP":"DISTANCE"].std()
#test set - use mean and std of the training set
p_data_test=data_test.copy(deep=True)
p_data_test.loc[:,"SCHED_DEP":"DISTANCE"]=(data_test.loc[:,"SCHED_DEP":"DISTANCE"]-data_train.loc[:,"SCHED_DEP":"DISTANCE"].mean())/data_train.loc[:,"SCHED_DEP":"DISTANCE"].std()
p_remaining_df = remaining_df.copy(deep=True)
p_remaining_df.loc[:,"SCHED_DEP":"DISTANCE"]=(remaining_df.loc[:,"SCHED_DEP":"DISTANCE"]-data_train.loc[:,"SCHED_DEP":"DISTANCE"].mean())/data_train.loc[:,"SCHED_DEP":"DISTANCE"].std()
# -
p_data_train.shape
# **Now, I am ready to try different classification models** <br>
# +
x_train = p_data_train.copy(deep=True).drop('DELAY_OR_NOT',axis=1)
y_train = p_data_train.DELAY_OR_NOT
x_test = p_data_test.copy(deep=True).drop('DELAY_OR_NOT',axis=1)
y_test = p_data_test.DELAY_OR_NOT
# -
# **I will use the classification models comparison code from Tuesday section 8 which reviewed last years midterm. However, I will throw out the kNN classification since calculating Euclidean distances between over 1000 predictors would be too computationally expensive**
#following lamda function scores predictions for two-way classifier
score = lambda model, x_train, y_train: pd.Series([model.score(x_train, y_train),
model.score(x_train[y_train==0], y_train[y_train==0]),
model.score(x_train[y_train==1], y_train[y_train==1])],
index=['overall accuracy', 'accuracy on class 0', 'accuracy on class 1'])
def out_of_box_class(x,y):
#Unweighted logistic regression
unweighted_logistic = LogisticRegression(C=1000)
unweighted_logistic.fit(x, y)
unweighted_log_scores = score(unweighted_logistic, x, y)
print('unweighted log')
#Weighted logistic regression
weighted_logistic = LogisticRegression(C=1000, class_weight='balanced')
weighted_logistic.fit(x, y)
weighted_log_scores = score(weighted_logistic, x, y)
print('weighted log')
#LDA
lda = LDA()
lda.fit(x, y)
lda_scores = score(lda, x, y)
print('lda')
#QDA
qda = QDA()
qda.fit(x, y)
qda_scores = score(qda, x, y)
print('qda')
#Decision Tree
tree = DecisionTree(max_depth=50, class_weight='balanced', criterion='entropy')
tree.fit(x, y)
tree_scores = score(tree, x, y)
print('tree')
score_df = pd.DataFrame({'unweighted logistic': unweighted_log_scores,
'weighted logistic': weighted_log_scores,
'lda': lda_scores,
'qda': qda_scores,
'tree': tree_scores})
return score_df
out_of_box_class(x_train,y_train)
# **From running these accuracy tests, I can see that the data is likely not multivariate normally (MVN) distributed (see LDA/QDA accuracy score on the class 0/1). Thus, LDA/QDA is not a good fit given the fact they require MVN property. Additionally, the decision tree is clearly overfitting. Thus we will choose weighted logistic regression as our classification model. Further justification is that only ~10% of my training data has DELAY==1 response, thus there needs to be weight assigned to the "positive DELAY" samples to ensure better accuracy of our model. Finally, the overall score of the weighted logit regression (still with non-tuned C parameter) is fairly high.**
# +
# Fit cross-validated weighted L2 logistic regression
clf = LogisticRegressionCV(fit_intercept=True, penalty='l2',class_weight='balanced')
clf.fit(x_train, y_train)
y_train_hat = clf.predict(x_train)
print("Training set prediction accuracy of logistic regression =", accuracy_score(y_train, y_train_hat))
# -
clf.Cs
# **After tuning the hyperparameter C on my weighted logistic regression, we got C = 10. In logistic regression, as seen in the class, C is the inverse of the regularization parameter $\lambda$.**
print(x_test.shape,x_train.shape)
clf_test_score = score(clf,x_test,y_test)
print("Accuracy on the test set achieved:")
print(clf_test_score)
auc_logreg_test = metrics.roc_auc_score(y_test, clf.predict_proba(x_test)[:,1])
print("The AUC score on the test set auc_logreg",auc_logreg_test)
# **I was able to beat the 60% benchmark AUC accuracy on the test set with the chosen weighted logistic model (C=10).**
# **Question 5**<br>
# (5pts) Given your model, comment on the importance of factors as related to whether a flight is delayed.
#Let's display some coefficients of our weighted logistic regression
clf.coef_[0][0:7]
#the order of coefficients is the same as the order of columns in my x_train data set
x_train.head()
# **The importance of each feature on flight delays can be shown from the coefficient values. We have 1322 coefficients for each of 1322 features. Each coefficient should represent how important each feature is in predicting whether the flight will be delayed or not. For example, looking at the first 6 coefficients, we have **<br>
#
#
# | SCHED_DEP |SCHED_ARR|DISTANCE|MONTH_1|MONTH_2|MONTH_3|MONTH_4
# |---|---|---|
# | 0.94856692|0.03805954| 0.02356447|0.42227514|0.60139052|0.11210393|-0.21800069
# <br>
#
# **And we can clearly see that the SCHED_DEP time is the most important predictor for the events of delayed flights among the displayed features while SCHED_ARR and DISTANCE are not that important and MONTH_4 is the least important. This makes sense with my visualisation that the percentage of delayed flights is much higher in the evening than in the morning. If the coefficient is negative, that means that the particular predictor is the least important (the least contribution of the beta coefficient in the exponential form) **
#
# **We could do a bootstrapping of our train data to get e.g. 100 values for each coefficient to get confidence intervals for our coefficients, but this would be too computationally expensive (we have >1,300 features and doing a LogisticRegressionCV one time takes ~10 minutes and I have only 36 hours (960 minutes) to complete the midterm!!)**
# +
#Now let's take a look at which predictors are most important in predicting a delay
#and which ones are the least importnat
X = x_train.columns
Y= clf.coef_[0]
Z = [x for _,x in sorted(zip(Y,X))]
print("Several features the least important for prediction of delay \n(chose most negative coefficients in the logit regression)")
print(Z[0:10])
print("\n")
print("Most important features predicting delay events (chose most positive coefficients in the logit regression)")
print(Z[-10:])
# -
# **As we can see from above, we again confirm our visualisation that month 9 has little importance in predicting delay events (on par with month 10) in accordance with the outcome of the weighted logistic model.**
# **QUESTION 6**<br>
# (5pts) Evaluate your model(s) on your test set, and finally provide a visual to show which airlines are predicted to have the most delays using all the data excluding the training and test set.
# **The model was already evaluated on the test set in the end of questions 4 but I will list it here again.**
clf_test_score = score(clf,x_test,y_test)
print("Accuracy on the test set achieved:")
print("Accuracy", clf_test_score)
print("The AUC score on the test set auc_logreg",auc_logreg_test)
# +
#Visual showing which airlines are predicted to have the most delays using all data except training and test set
p_remaining_df.head()
y_test_remain = p_remaining_df.DELAY_OR_NOT
x_test_remain = p_remaining_df.drop('DELAY_OR_NOT',axis=1)
#now predict on the full remaining data_set
y_pred_remain = clf.predict(x_test_remain)
# -
score(clf,x_test_remain,y_test_remain)
auc_logreg_rem = metrics.roc_auc_score(y_test_remain, clf.predict_proba(x_test_remain)[:,1])
print("The AUC score for the remaining whole data set is",auc_logreg_rem)
# **The accuracy of prediction achieved on the remaining full dataset is 0.754205 and AUC is 0.823**
print("Predicted delayed",sum(y_pred_remain==1),"; true delayed",sum(y_test_remain))
original_dataframe = new_df.drop(new_df_sample.index) #getting the rest of indices not used in train/test
predicted_dataframe = new_df.drop(new_df_sample.index) #getting the rest of indices not used in train/test
sns.countplot(x='AIRLINE',hue='DELAY_OR_NOT',data=original_dataframe, palette="Set1");
plt.xlabel("Airline")
plt.ylabel("Truth value count of delayed flights")
plt.title("Truth values");
#Let's look at the mean delay % for each airline (truth)
original_dataframe.groupby("AIRLINE").mean()
predicted_dataframe['DELAY_OR_NOT'] = y_pred_remain
sns.countplot(x='AIRLINE',hue='DELAY_OR_NOT',data=predicted_dataframe,palette="Set1");
plt.xlabel("Airline")
plt.ylabel("Predicted count of delayed flights")
plt.title("Prediction");
#Let's look at the mean delay % for each airline (PREDICTION)
predicted_dataframe.groupby("AIRLINE").mean()
# +
#Now take a look at how many flights we classify FALSE POSITIVE and FALSE NEGATIVE
from sklearn.metrics import confusion_matrix
from sklearn.metrics import roc_curve
print(confusion_matrix(y_test_remain,y_pred_remain))
tn_logreg1, fp_logreg1, fn_logreg1, tp_logreg1 = metrics.confusion_matrix(y_test_remain,y_pred_remain).astype('float').ravel()
print("\nThe true positive rate for the weighted logistic regression model:",tp_logreg1/(tp_logreg1 + fn_logreg1))
# -
# **Comparing the two plots above (truth and prediction, respectively) we see that our model predicts more delayed flights than there actually are: Predicted delayed 207829 (TP:53203,FP:154626), True delayed total 71,848 out of 700,000 flights in the remaining dataset (see the confusion matrix above). In other words, we have more FPRs than FNRs which makes sense as our data is highly unbalanced towards not delayed flights and our model is balanced in a way to put extra weight on the positive classification.** <br>
#
# **E.g. for airlines "NK" and "F9", more flights are predicted to be delayed than on time, than in the truth baseline. However, this is to be expected given our AUC of the classification model is ~80%.**
#
# **Airlines with most delays: 'NK' has 76%/37% delayed flights (predicted/ground truth) followed by 'F9': 60%/22% and 'B6': 47%/16% . We can see that the percentage of delayed flights predicted is higher than ground truth in all cases, which is consistent with the high false positive ratio I have with my model.**
#
# **Speculation: if the user of my model preferred to get more false positives/false negatives (depending whether the user is more concerned about learning that a flight is predicted to be delayed when it's on time (FPR) or a flight is predicted to be on time when it is delayed - FNR. I would imagine for e.g., a businessman, being on the side of caution and having more FPRs than FNRs and taking precations to not e.g., miss a business meeting, might be a more useful use-case.**
# **Question 7**<br>
# (15pts) Build a regression model that predicts the length of delay (on the log scale) given that a flight is truly delayed.
# +
#First, we want to look at lenght of flight GIVEN the flight is delayed - only select observations when a flight is truly delayed
new_df_regre = df.copy(deep=True)[df.DELAY_OR_NOT==1]
#drop everything as it was, but keep ARRIVAL_TIME as that will be our response variable
del new_df_regre['DATE']
del new_df_regre['SCHEDULED_TIME']
del new_df_regre['ELAPSED_TIME']
del new_df_regre['FLIGHT_NUMBER']
del new_df_regre['TAIL_NUMBER']
del new_df_regre['DEPARTURE_DELAY']
del new_df_regre['AIR_SYSTEM_DELAY']
del new_df_regre['AIRLINE_DELAY']
del new_df_regre['SECURITY_DELAY']
del new_df_regre['LATE_AIRCRAFT_DELAY']
del new_df_regre['WEATHER_DELAY']
# -
new_df_regre.head()
new_df_regre.shape
#Change time to decimal in scheduled dep and arr
date_str_dep = list(new_df_regre.SCHED_DEP)
date_str_arr = list(new_df_regre.SCHED_ARR)
time_tuples_dep = []
time_tuples_arr = []
for i,j in zip(date_str_dep,date_str_arr):
time_tuple_dep = time.strptime(i, "%H:%M:%S")
time_tuple_arr = time.strptime(j, "%H:%M:%S")
time_tuples_dep.append(time_tuple_dep.tm_hour+time_tuple_dep.tm_min/60)
time_tuples_arr.append(time_tuple_arr.tm_hour+time_tuple_arr.tm_min/60)
new_df_regre['SCHED_DEP'] = np.array(time_tuples_dep)
new_df_regre['SCHED_ARR'] = np.array(time_tuples_arr)
#Check we have continuos time in decimals (hours) and that we only have DELAYED == True observations
new_df_regre.head()
#create a log response variable ARRIVAL TIME
new_df_regre['ARRIVAL_DELAY']=np.log(new_df_regre['ARRIVAL_DELAY'])
# +
#Now, we need to convert our categorical variables
cols_to_change =['MONTH','DAY','DAY_OF_WEEK','AIRLINE','ORIGIN_AIRPORT','DESTINATION_AIRPORT']
new_df_dummy_regre = pd.get_dummies(new_df_regre,columns=cols_to_change)
#Subsample the data to reduce the # of observations.
new_df_sample_regre = new_df_dummy_regre.sample(n=16000,random_state=6)
#Split the data in test and training (see Lecture 5)
np.random.seed(9001)
data_train_regre, data_test_regre = train_test_split(new_df_sample_regre,test_size=0.33, random_state=0)
#Check we actually got the right shapes
print(new_df_sample_regre.shape,df.shape,data_train_regre.shape,data_test_regre.shape)
data_train_regre.head()
# +
#now we standardize our continuous variables
p_data_train_regre = data_train_regre.copy(deep=True)
p_data_train_regre.loc[:,"SCHED_DEP":"DISTANCE"] = (data_train_regre.loc[:,"SCHED_DEP":"DISTANCE"] - data_train_regre.loc[:,"SCHED_DEP":"DISTANCE"].mean())/data_train_regre.loc[:,"SCHED_DEP":"DISTANCE"].std()
#test set - use mean and std of the training set
p_data_test_regre=data_test_regre.copy(deep=True)
p_data_test_regre.loc[:,"SCHED_DEP":"DISTANCE"]=(data_test_regre.loc[:,"SCHED_DEP":"DISTANCE"]-data_train_regre.loc[:,"SCHED_DEP":"DISTANCE"].mean())/data_train_regre.loc[:,"SCHED_DEP":"DISTANCE"].std()
# +
#create x-train, y-train
x_train_regre = p_data_train_regre.copy(deep=True).drop('ARRIVAL_DELAY',axis=1)
y_train_regre = p_data_train_regre.ARRIVAL_DELAY
x_test_regre = p_data_test_regre.copy(deep=True).drop('ARRIVAL_DELAY',axis=1)
y_test_regre = p_data_test_regre.ARRIVAL_DELAY
# +
#We create our regression model by performing regularized linear regression - Lasso.
Lasso_pipe = Pipeline([ ('mlr', LassoCV(cv=3)
)])
Lasso_pipe.fit(x_train_regre, y_train_regre)
# +
from sklearn.metrics import r2_score
y_pred_train = Lasso_pipe.predict(x_train_regre)
y_pred = Lasso_pipe.predict(x_test_regre)
print("R^2 accuracy prediction on the training set")
print(r2_score(y_train_regre,y_pred_train))
print("R^2 accuracy prediction on the test set")
print(r2_score(y_test_regre,y_pred))
# -
# **I achieved R^2 over 3% on the test set so I am in the clear! I used Lasso regression with cross validation as it allows to regularize features coefficients and also removes not-significant coefficients.**
# # 209 Additional questions
# (10pts) Engineer two additional features that will help improve the classification model's performance.<br>
#
#
#
# # First engineered feature
# +
#Let's take a look to find how some of my features are correlated.
x_train.columns[53:67]
add = [i for i in range(53,100)]
features_to_look_at = [i for i in range(15)]
for i in add:
features_to_look_at.append(i)
a=np.corrcoef(x_train.iloc[:,features_to_look_at].T)
plt.figure(figsize=(20,15))
sns.heatmap(a,xticklabels=list(x_train.iloc[:,features_to_look_at].columns), yticklabels=list(x_train.iloc[:,features_to_look_at].columns));
# -
# **It is hard to tell which features are correlated from the correlation matrix above (we can only display a small subset of the predictors given our predictor space is too large. We get some blank values since certain airports do not have enough flights to calculate the correlation matrix**
# +
#We will add the interaction feature between day of the week - Thursday and scheduled departure time.
#We saw in the visualisation that Thursday has the largest % of delayed flights
#and also that evening-hours flights have a higher percentage of delayed than morning ones
#It is computationally not viable to go through all the different interaction terms (even of degree 2), given our large feature space
#So we have to think about which combinations might make logical sense.
x_train_eng1 = p_data_train.copy(deep=True).drop('DELAY_OR_NOT',axis=1)
y_train_eng1 = p_data_train.DELAY_OR_NOT
x_test_eng1 = p_data_test.copy(deep=True).drop('DELAY_OR_NOT',axis=1)
y_test_eng1 = p_data_test.DELAY_OR_NOT
# -
#Creating the interaction featured between Thursday and time of departure
x_train_eng1['inter_time_day']= x_train_eng1.DAY_OF_WEEK_4*x_train_eng1.SCHED_DEP
x_test_eng1['inter_time_day']= x_test_eng1.DAY_OF_WEEK_4*x_test_eng1.SCHED_DEP
# +
# Fit cross-validated weighted 'L2'-penalized logistic regression
clf_eng = LogisticRegressionCV(fit_intercept=True, penalty='l2',class_weight='balanced')
clf_eng.fit(x_train_eng1, y_train_eng1)
y_train_hat = clf_eng.predict(x_train_eng1)
print("Training set prediction accuracy of logistic regression =", accuracy_score(y_train_eng1, y_train_hat))
# -
auc_logreg_eng = metrics.roc_auc_score(y_test_eng1, clf_eng.predict_proba(x_test_eng1)[:,1])
print("The AUC score without engineered features on the test set ",auc_logreg_test)
print("The AUC score with a engineered feature on the test set",auc_logreg_eng)
# **We achieved slight improvemenent from AUC of 0.821 to 0.822 on the test set, which may be small, but given the fact we have over 1300 features, this interaction feature still adds a bit of accuracy. I observed most delayed flights happen on Thursdays and also in the evening from the visualisation, thus combining these two features added a bit of classification accuracy makes a logical sense.**
#Now test the model with the engineered feature on the entire dataset
x_test_remain['inter_time_day'] = x_test_remain.DAY_OF_WEEK_4*x_test_remain.SCHED_DEP
#Calculate the AUC score
auc_logreg_eng_rem = metrics.roc_auc_score(y_test_remain, clf_eng.predict_proba(x_test_remain)[:,1])
print("AUC on the remaining data without engineered features",auc_logreg_rem)
print("AUC on the remaining data WITH a engineered features",auc_logreg_eng_rem)
# **Despite achieving a minimal improvement with the first engineered feature on the test set, we actually did slightly worse on the remaining dataset.**
# # Second engineered feature
# +
#Let's get rid of the airports which have less than 100 flights in a year
# - to reduce the complexity of our model by engineering the ORIGIN_AIRPORT feature
new_df_eng = df.copy(deep=True)
del new_df_eng['DATE']
del new_df_eng['SCHEDULED_TIME']
del new_df_eng['ELAPSED_TIME']
del new_df_eng['FLIGHT_NUMBER']
del new_df_eng['TAIL_NUMBER']
del new_df_eng['ARRIVAL_DELAY']
del new_df_eng['DEPARTURE_DELAY']
del new_df_eng['AIR_SYSTEM_DELAY']
del new_df_eng['AIRLINE_DELAY']
del new_df_eng['SECURITY_DELAY']
del new_df_eng['LATE_AIRCRAFT_DELAY']
del new_df_eng['WEATHER_DELAY']
# -
#Change time to decimal in scheduled dep and arr
date_str_dep = list(new_df_eng.SCHED_DEP)
date_str_arr = list(new_df_eng.SCHED_ARR)
time_tuples_dep = []
time_tuples_arr = []
for i,j in zip(date_str_dep,date_str_arr):
time_tuple_dep = time.strptime(i, "%H:%M:%S")
time_tuple_arr = time.strptime(j, "%H:%M:%S")
time_tuples_dep.append(time_tuple_dep.tm_hour+time_tuple_dep.tm_min/60)
time_tuples_arr.append(time_tuple_arr.tm_hour+time_tuple_arr.tm_min/60)
new_df_eng['SCHED_DEP'] = np.array(time_tuples_dep)
new_df_eng['SCHED_ARR'] = np.array(time_tuples_arr)
# +
#Now, we need to convert our categorical variables
cols_to_change =['MONTH','DAY','DAY_OF_WEEK','AIRLINE','ORIGIN_AIRPORT','DESTINATION_AIRPORT']
new_df_dummy = pd.get_dummies(new_df_eng,columns=cols_to_change)
#get the airports with only more than 100 flights
indices_to_keep = new_df.ORIGIN_AIRPORT.value_counts()[list(new_df.ORIGIN_AIRPORT.value_counts()>100)].index
collection = []
for c in new_df_dummy.columns:
if "ORIGIN_AIRPORT" in c or "DESTINATION_AIRPORT" in c:
tag = c.replace("ORIGIN_AIRPORT_", "").replace("DESTINATION_AIRPORT_","")
if not tag in indices_to_keep:
collection.append(c)
new_df_dummy=new_df_dummy.drop(collection, axis=1)
#Subsample the data to reduce the # of observations.
new_df_sample = new_df_dummy.sample(n=100000,random_state=6)
#Split the data in test and training (see Lecture 5)
np.random.seed(9001)
data_train_eng, data_test_eng = train_test_split(new_df_sample,test_size=0.33, random_state=0)
#Check we actually got the right shapes
print(new_df_sample.shape,df.shape,data_train_eng.shape,data_test_eng.shape)
data_train_eng.head()
# +
#now we standardize our continuous variables
p_data_train_eng = data_train_eng.copy(deep=True)
p_data_train_eng.loc[:,"SCHED_DEP":"DISTANCE"] = (data_train_eng.loc[:,"SCHED_DEP":"DISTANCE"] - data_train_eng.loc[:,"SCHED_DEP":"DISTANCE"].mean())/data_train_eng.loc[:,"SCHED_DEP":"DISTANCE"].std()
#test set - use mean and std of the training set
p_data_test_eng=data_test_eng.copy(deep=True)
p_data_test_eng.loc[:,"SCHED_DEP":"DISTANCE"]=(data_test_eng.loc[:,"SCHED_DEP":"DISTANCE"]-data_train_eng.loc[:,"SCHED_DEP":"DISTANCE"].mean())/data_train_eng.loc[:,"SCHED_DEP":"DISTANCE"].std()
# -
p_data_train_eng.head()
# +
x_train_eng2 = p_data_train_eng.copy(deep=True).drop('DELAY_OR_NOT',axis=1)
y_train_eng2 = p_data_train_eng.DELAY_OR_NOT
x_test_eng2 = p_data_test_eng.copy(deep=True).drop('DELAY_OR_NOT',axis=1)
y_test_eng2 = p_data_test_eng.DELAY_OR_NOT
# +
# Fit cross-validated weighted L2 logistic regression
clf_eng2 = LogisticRegressionCV(fit_intercept=True, penalty='l2',class_weight='balanced')
clf_eng2.fit(x_train_eng2, y_train_eng2)
y_train_hat = clf_eng2.predict(x_train_eng2)
print("Training set prediction accuracy of logistic regression =", accuracy_score(y_train_eng2, y_train_hat))
# -
auc_logreg_eng2 = metrics.roc_auc_score(y_test_eng2, clf_eng2.predict_proba(x_test_eng2)[:,1])
print("The AUC score without engineered features on the test set ",auc_logreg_test)
print("The AUC score with a engineered feature (number of airports) on the test set",auc_logreg_eng2)
# **We can see we improved our accuracy by engineering the "ORIGIN_AIRPORTS" feature by a bit, but most importantly we reduced the feature space from 1,300 to about 700, which improves the computational speed significantly.**
# (5pts) Add one additional feature from a data source not given to you. Do this only after you complete the rest of the exam.
# **Given lack of time, I will just add the "diverted" occurences feature and only look at January as I won't have enough time to clean and concatenate all of the months. I will add the feature called "DIVERTED" to see whether DIVERTED affected delay flights**
#read the dataset I downloaded from https://www.transtats.bts.gov/DL_SelectFields.asp?Table_ID=236&DB_Short_Name=On-Time
jan_df = pd.read_csv("Jan_15_flights.csv")
jan_df.head()
#fill in for 0s, we will only be using ARR_DEL15 as our response which has NaNs, other variables with NaN;s will be dropped.
my_df = jan_df.fillna(0)
my_df.columns
del my_df['ACTUAL_ELAPSED_TIME']
del my_df['DIV_AIRPORT_LANDINGS'] #duplicate to DIVERTED
del my_df['Unnamed: 13']
my_df.head()
# +
cols_to_change =['MONTH','DAY_OF_MONTH','DAY_OF_WEEK','AIRLINE_ID','ORIGIN','DEST']
#hot-encode
new_df_dummy = pd.get_dummies(my_df,columns=cols_to_change)
#Subsample the data to reduce the # of observations.
new_df_sample = new_df_dummy.sample(n=50000,random_state=6)
#Split the data in test and training (see Lecture 5)
np.random.seed(9001)
data_train, data_test = train_test_split(new_df_sample,test_size=0.33, random_state=0)
#Check we actually got the right shapes
print(new_df_sample.shape,df.shape,data_train.shape,data_test.shape)
data_train.head()
# +
#standardize the data
p_data_train=data_train.copy(deep=True)
p_data_train.iloc[:,[0,1,4]]=(data_train.iloc[:,[0,1,4]]-data_train.iloc[:,[0,1,4]].mean())/data_train.iloc[:,[0,1,4]].std()
#test set - use mean and std of the training set
p_data_test=data_test.copy(deep=True)
p_data_test.iloc[:,[0,1,4]]=(data_test.iloc[:,[0,1,4]]-data_train.iloc[:,[0,1,4]].mean())/data_train.iloc[:,[0,1,4]].std()
# -
p_data_train.head()
# +
x_train = p_data_train.copy(deep=True).drop('ARR_DEL15',axis=1)
y_train = p_data_train.ARR_DEL15
x_test = p_data_test.copy(deep=True).drop('ARR_DEL15',axis=1)
y_test = p_data_test.ARR_DEL15
# +
# Fit cross-validated weighted L2 logistic regression
clf = LogisticRegressionCV(fit_intercept=True, penalty='l2',class_weight='balanced')
clf.fit(x_train, y_train)
y_train_hat = clf.predict(x_train)
print("Training set prediction accuracy of logistic regression =", accuracy_score(y_train, y_train_hat))
# -
clf_test_score = score(clf,x_test,y_test)
print("Accuracy on the test set achieved:")
print(clf_test_score)
auc_logreg_test = metrics.roc_auc_score(y_test, clf.predict_proba(x_test)[:,1])
print("The AUC score on the test set auc_logreg",auc_logreg_test)
# **In this last 209a subproblem, I added DIVERT feature. It is inconclusive to say whether it improved since I was not able to pull in the whole 2015 year in time from the website. I still beat the benchmark AUC provided by the Hint on this Midterm!.**
# **Question 8**<br>
# (20pts) Write a report (in the last markdown cell in your notebook with your findings (without code)). Describe the main design decisions you have made with justifications. Clearly explain your methodology and results. This should not be more than 300 words. You may use up to 5 diagrams.
# Summary: After pre-processing and exploration of the data I have decided to eliminate certain features as described above. I believe the various "delay" features are not useful in predicting the delay events since they are sort of after-the-fact (e.g., delayed departure is already after-the-fact feature). In visualisation, I found that select features (Airline, Hour of the day, month, day of the week) likely heavily determine the probability of a delayed flight - see diagrams. Then, I decided to use the weighted logistic regression for classification. I based this decision mainly on the fact we have an unbalanced training set (only 10% of observations are delayed flight events), lack of assumption that we have multivariate normally distributed data (exclusion of LDA/QDA), and also finding the best score with weight logistic regression among classification models (kNN too computationally expensive, decision tree clearly overfitting on the training set). My model performed fairly well, with AUC of 82% on the whole dataset (exc. sample). Of note, the coefficients of features in weighted logistic regression dictate their importance to a classification decision (the lower the coefficient, including negative, the less contributing it is). Next up, with my model, I had a high False Positive classification decisions rate (~75%) but I was limited by an unbalanced dataset. Additionally, the False Negatives rate was better (~4%). Also, a high FPR might be better than high FNR if our model were to be used in practice - people err on the side of caution whether a flight will be delayed or not to make arrangements given they will likely be late.
#
# In the second part, I chose the logarithm of "ARRIVAL_TIME" as a response variable to build a predictive model for the length of the delay. I chose the cross-validated L2-penalized Lasso regression to build my model. I chose Lasso because it eliminates features non-important for predicting the response variable and I also wanted to regularize to avoid overfitting on the training set (in contrast, Ridge does not eliminate features completely, only minimizes their weight). I achieved R^2 of ~3.4% which is above the given baseline. Further improvements could be made by e.g.,expanding the training set with Polynomial interaction terms before Lasso regression, however, in case of >1000 predictors, this is too computationally expensive. I also found PCA was not useful in case of reducing dimensionality when dealing with categorical variables (which we have many in this hot-encoded dataset).
#
# Finally, for 209a, after looking at a subset of predictors in a correlation matrix, I was not able to clearly see which ones are correlated so I used "common sense" and tried a combination of a day with worst percentage of delayed flights (Thursday) with scheduled departure time - which had fairly high coefficient in logistic regression. I was able to achieve only very slight AUC improvement.
#
# The second feature engineering was adjusting one existing feature - dropping airports with less than 100 flights from the dataset. This also did not significantly improved AUC, but it majorly reduced the complexity of the model (from 1300 to 700 features).
#
# Finally, for the additional added feature, I used DIVERTED (bool: 0/1) from the Transportation Bureau website to see whether DIVERTED flights significantly affect prediction of delays. I followed the same steps as in previous analyses and got an AUC of 71%. This is caused by the fact I only had time to clean January 2015 and not concatenate months.
| Midterm - Predicting Airline Delays/Midterm_Predicting_Airline_Delays.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import keras
import numpy
x = numpy.array([0, 1, 2, 3, 4])
y = [1, 3, 5, 7, 9]
model = keras.models.Sequential()
model.add(keras.layers.Dense(1, input_shape=(1, )))
model.compile('SGD', 'mse')
model.fit(x[:2], y[:2], epochs=1000, verbose=1)
model.save('model_1.h5')
| jupyter/eMachinLearning/k_keras_save_restore/Ex03_k_model_saver.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Image of a Black Hole
# <NAME> /
# January 2021
# ## Photons Geodesics
# First of all, the trajectory of photons is studied. The Black hole has a mass M, it is a non rotative one and not charged one, called : Schwarzchild Black hole. In polar coordinate one can express geodesic equation as :
# \begin{align} \left(\frac{1}{r^2}\frac{\partial r}{\partial \theta} \right)^2 +\frac{1}{r^2}\left(1-\frac{r_s}{r} \right) = \frac{1}{b^2}\end{align}
# Where b is the infinite impact parameter and $r_s = 2GM/c^2$ the Schwarzschild radius.
#
# This equation can be re written using $u=r_s/r$ and $u_c = b_c/b$ with $b_c = 3\sqrt(3)/2$ :
#
# \begin{align} \left(\frac{\partial u}{\partial \theta}\right)^2 + V(u) = 0 \end{align}
# With \begin{align} V(u) = u^2(1-u)-\frac{4}{27}u_c^2 \end{align}
# +
# Packages
import numpy as np
import matplotlib.pyplot as plt
from scipy import *
from pylab import *
from scipy.integrate import odeint
from scipy.interpolate import interp1d
from matplotlib import ticker,cm
###########################
# Constants
###########################
G = 6.67*10**(-11) # Gravitation constant
Msoleil = 1.989*10**(30) # Solar mass
M = 30*Msoleil # We consider a 30 solar mass black hole
c = 3.00*10**8 # Speed light
rs = 2*G*M/c**2 # Schwarzschild radius
bc = 3*np.sqrt(3)*rs/2
# -
# It is then possible to plot V(u) for different values of b the impact parameter.
# +
############################
# V(u) Definition
############################
def V(u,b):
uc = bc/b
return u**2*(1-u)-4/27*uc**2
def const(u):
return 0*u
# Plotting V(u)
b=[0.8*bc,0.6*bc,0.9*bc,bc,1.3*bc,1.6*bc,2*bc,3*bc]
u = np.linspace(-1,2,2000)
for elem in b:
plt.plot(u,V(u,elem),label='b='+str(elem/bc)+'bc')
plt.plot(u,const(u),'k')
plt.xlabel('u = rs/r')
plt.ylabel('V(u)')
plt.ylim([-0.5,0.2])
plt.xlim([0,1.5])
plt.legend()
plt.show()
# -
# It is possible to distinguish 2 different regimes :
# - One for $b < b_c$, the arriving photon goes nearer than the limit trapping radius of the black hole. In this case, the arriving photons are trapped by the black hole.
# - On the other hand, for $ b > b_c $, the arriving photons are facing a potential barrier and are defleted by the black hole and go back to infinity.
# In order to find the photon geodesics, the differential equation needs to be solved. It is easier to work with the derived equation
# \begin{align} u'' = 3/2u^2-u \end{align}
# In order to solve this equation, the 4th order Runge Kutta method for second derivative differential equation is computed. The method is the following, if the equation to solve is $ u''=f(\theta,u,u') $ with $ f(\theta,u,u') = \frac{3}{2}u^2-u $. We then define :
# $$ k_1 = f(\theta_n,u_n,u'_n) $$
# $$ k_2 = f(\theta_n +\frac{h}{2},u_n+\frac{h}{2}u'_n,u'_n+\frac{h}{2}k_1) $$
# $$ k_3 = f(\theta_n +\frac{h}{2},u_n+\frac{h}{2}u'_n+\frac{h^2}{4}k_1,u'_n+\frac{h}{2}k_2)$$
# $$ k_4 = f(\theta_n +h,u_n+h*u'_n+\frac{h^2}{2}k_2,u'_n+h*k_3) $$
# And then the solution will be deduced by :
# $$ u_{n+1} = u_n + h*u'_n + \frac{h^2}{6}(k_1+k_2+k_3) $$
# and
# $$ u'_{n+1} = u'_n + \frac{h}{6}(k_1+2k_2+2k_3+k_4) $$
# For the resolution we consider that if a photon is deviated by the black hole at the point that it goes back to infinity we consider that it stayes at infinity and don't go back to the black hole (*).
# Function definition
def fonct(U):
dudtheta = 3/2*U**2-U
return dudtheta
# Parameter to define theta start and theta end in order to solve the equation between theta start and stop
start = 0
stop = 20
N=1000
def solve_equadiff(b):
'''
Solve differential equation u''=3/2u**2-u using 4th order Runge Kutta method for 2nd order differential equations
Input : - b = impact parameter
Output : ulist, uprimelist, theta
- ulist = list of values of function u
- uprimelist = list of values of derivative of u function
- theta = list of theta on which the differential equation is solved
'''
uc = bc/b
# Initialisation
u0 = 0
uprime0 = 2*uc/(3*np.sqrt(3))
# Step
h = (-start+stop)/(N-1)
u=u0
uprime=uprime0
theta=np.linspace(start,stop,N)
# Creation of ulist and uprimelist
u_list=[]
uprime_list=[]
u_list.append(u)
uprime_list.append(uprime)
for i in range(len(theta)-1):
k1 = fonct(u)
k2 = fonct(u+h/2*uprime)
k3 = fonct(u+h/2*uprime+h**2/4*k1)
k4 = fonct(u+h*uprime+h**2/2*k2)
u = u + h*uprime + h**2/6*(k1+k2+k3)
uprime = uprime + h/6*(k1+2*k2+2*k3+k4)
u_list.append(u)
uprime_list.append(uprime)
# when a photo goes back to infinity it stayes at infinity (*)
t=0
while ( u_list[t]>=0 and t<len(u_list)-1):
t=t+1
for k in range(t,len(u_list)):
u_list[k] = 0
# we set the maximum value for u to u = rs / r = 10
t=0
while ( u_list[t]<=10 and t<len(u_list)-1):
t=t+1
for k in range(t,len(u_list)):
u_list[k] = 10
return u_list, uprime_list, theta
# It is then possible to plot u the solution of the differential equation for different values of b and to determine how many times the photon turns around the Black hole by looking at the support of the u function that is to say the length of the segment such as { $ \theta | u>0 $ }.
#
#
# One can highlight the fact that the more b is close to bc te more the photon will turn around the Black hole.
# +
ul1 , upl, th = solve_equadiff(bc)
ul2 , upl, th = solve_equadiff(1.2*bc)
ul3 , upl, th = solve_equadiff(1.5*bc)
ul4 , upl, th = solve_equadiff(2*bc)
ul5 , upl, th = solve_equadiff(10**(10)*bc)
plt.figure()
#plt.plot(th[1],ul1,label='b/bc = {}'.format(str(1)))
plt.plot(th,ul2,label='b/bc = {}'.format(str(1.2)))
plt.plot(th,ul3,label='b/bc = {}'.format(str(1.5)))
plt.plot(th,ul4,label='b/bc = {}'.format(str(2)))
plt.plot(th,ul5,label='b/bc = Inf')
plt.ylim([0,0.5])
plt.xlabel('Theta')
plt.ylabel('u')
plt.title('u(theta)')
plt.legend()
plt.show()
# -
# One can compute now the phase portrait of the u fonction
# +
plt.figure()
b=[0.2*bc,0.6*bc,1*bc,3*bc]
for i in range(len(b)):
ulist, uprimelist,theta = solve_equadiff(b[i])
plt.plot(ulist,uprimelist,label='b='+str(b[i]/bc)+'bc')
plt.xlim([0,2])
plt.ylim([0,3])
plt.title('Phase portrait',loc='center')
plt.ylabel('u')
plt.ylabel('du/dtheta')
plt.legend()
plt.show()
# -
# Looking at the previous graph one can validate the fact that for $b<b_c$ the photon will fall on the Black hole, on the other hand for $b>b_c$ the photon are attracted and the goes bah to infinty with a null speed.
# ## Accretion Disk
# Now that we know how photons propagate we will try to display images of a black hole and its properties.
from IPython.display import Image
Image("disque_enonce.png")
# We consider a black hole inclined with an angle i compared to the observer. The reverse return of light is valid for Schwaerzchild black hole. Therefore it is possible to goes back from the observer plan, follow back geodesics until finding the black hole disk at radius $r_d$.
#
# In order to simplify we will assume that the unit vector nomal to the accretion disk plan is included in the plan $\alpha=\frac{\pi}{2}$.
#
# A photon leaves the disk at a radius $r_d$ and angle $\theta_d$.
# Three dimensional trigonometry gives us :
# $$ cos(\theta_d) = \frac{-sin(\alpha)cos(i)}{\sqrt{1-cos^2(\alpha)cos^2(i))}} $$
# and as we know that the photons are following the geodesics from the moment they leave the disk to the moment they arrive on observer plan one can write :
# $$ r_d=u(\theta_d) $$
# with u defined earlier.
# In order to consider the computed quantities ( iso-radius, redshift, flux ) we introduces 2 variables x and y defined as following :
# $$ x=b*cos(\alpha) $$ and $$ y=b*sin(\alpha) $$
# Parameter to compute the accretion disk
pas = 100
i_list = [ 10*np.pi/180, 40*np.pi/180, 80*np.pi/180]
# We first focus on the primary images, that is to say the images made by photons that didn't made a round around the black hole.
# +
def compute_primary(x, y):
'''
Fonction in order to compute the photons from primary image, direct photons coming
'''
x=x*bc
y=y*bc
b = np.sqrt(x**2+y**2)
alpha = (y/b)/(np.abs(y/b))*np.arccos(x/b)
thetad = np.arccos( -np.sin(alpha)*np.cos(i)/np.sqrt(1-np.cos(alpha)**2*np.cos(i)**2) )
if len(thetad)>1:
fct_u = np.ones((pas,pas))
for k in range(pas):
for l in range(pas):
ul , upl, th = solve_equadiff(b[k][l])
fct_u[k][l] = abs(1/(interp1d(th,ul,kind='cubic')(thetad[k][l])))
else :
fct_u = interp1d(th,ul,kind='cubic')(theta)
return fct_u
# -
# Plot the map for different inclinaision ( 10, 40, 80 )
for a in range(len(i_list)) :
i = i_list[a]
x = np.linspace(-5,5,pas)
y = np.linspace(-5,5,pas)
X,Y = np.meshgrid(x, y)
rd = compute_primary(X,Y)
fig, ax = plt.subplots()
plt.pcolor(X, Y, rd, cmap = 'inferno')
#cset2 = ax.contour(X, Y, rd, colors='k')
#cs = ax.contourf(X, Y, rd, locator=ticker.LogLocator(), cmap=cm.PuBu_r)
level = [2,6,10,15,20,30,50]
cs = ax.contourf(X, Y, rd,level)
cbar = fig.colorbar(cs)
cbar.ax.set_ylabel('rd/rs')
plt.ylabel(' y ')
plt.xlabel(' x ')
plt.title('rd computation, inclinaison : {}'.format(str(i*180/np.pi)))
plt.show()
# - For low disk inclinaision ( 10 degree ) the rays are distorded above the black hole. We can make the hypothesis that these photon come from behind the black hole because of distorded images.
# - For higher inclianision ( 80 degree ) iso-radius look like circular and non distorded. This confirm what is said in the previous point.
# It is also to compute the secondary images of the black hole, obtained by replacing $\alpha$ by $\alpha + \pi$, that is to says $\theta_d = \pi + arccos \left (\frac{sin(\alpha)cos(i)}{\sqrt{1-cos^2(\alpha)cos^2(i))}} \right )$ but it won't be done here.
# The emitted flux by disk surface unity at a distance r from the black hole is :
#
# \begin{align}
# \dfrac{F_{emis}(x)}{F_0} = \dfrac{x^{-5/2}}{x - 3/2} \left[ \sqrt{x} - \sqrt{3} + \sqrt{\dfrac{3}{8}} \ ln \left ( \dfrac{\sqrt{x} + \sqrt{3/2}}{\sqrt{x} - \sqrt{3/2}} \dfrac{\sqrt{2} - 1}{\sqrt{2} + 1} \right )\right ]
# \end{align}
# with $x=r/r_s$
# ## Accretion disk luminosity
# +
def Femis_F0(x):
facteur = x**(-5/2)/(x-3/2)
quotient1 =(np.sqrt(x)+np.sqrt(3/2))/(np.sqrt(x)-np.sqrt(3/2))
quotient2 = (np.sqrt(2)-1)/(np.sqrt(2)+1)
return facteur*(np.sqrt(x)-np.sqrt(3)+np.sqrt(3/8)*np.log(quotient1*quotient2))
x_1 = np.linspace(0,5,1000)
x_l = np.linspace(0.1,25,1000)
plt.semilogy(x_l,Femis_F0(x_l))
plt.xlabel('r/rs')
plt.ylabel('log (Femis/F0)')
plt.show()
plt.plot(x_1,Femis_F0(x_1))
plt.xlabel('r/rs')
plt.ylabel('Femis/F0')
plt.ylim([-1,60])
plt.show()
# -
# #### Redshift
# We then compute the redshift map around the black hole. As its density is very high the redshift around the black hole is modified. We have :
# \begin{align}
# 1+z = \frac{1}{\sqrt{1-\frac{3}{2x}}}* \left [ 1 + \left ( \dfrac{3}{2x} \right )^{3/2} \dfrac{b}{b_c}\right ]cos(i)cos(\alpha)
# \end{align}
# The considered redshift map will be computed for a disk between $r_{min}=3r_s$ and $r_{max} >> r_{min}$.
# +
def comparex_3(x,min_x,max_x):
'''
Fonction in order to consider only a disk starting at 3rs
'''
bool_l = np.ones((len(x[0]), len(x[0])))
for k in range(len(x[0])):
for l in range(len(x[0])):
if x[k][l] < min_x :
bool_l[k][l] = 10**(20)
if x[k][l] > max_x :
bool_l[k][l] = 10**(20)
return bool_l
# -
def compute_zbis(x1,y,rad):
'''
Fonction in order to compute redshift from formula
'''
x1=x1*bc
y=y*bc
b = np.sqrt(x1**2+y**2)
alpha = (y/b)/(np.abs(y/b))*np.arccos(x1/b)
x = rad
facteur1 = 1/np.sqrt(1-3/(2*x))
facteur2 = ( 1 + (3/(2*x))**(3/2)*b/bc*np.cos(i)*np.cos(alpha))
return comparex_3(x,rmin,rmax)*(facteur1*facteur2-1)
# +
rmin = 3
rmax = 25
# Plot the map for different inclinaision ( 10, 40, 80 )
for a in range(len(i_list)) :
i = i_list[a]
x = np.linspace(-3,3,pas)
y = np.linspace(-3,3,pas)
X,Y = np.meshgrid(x, y)
rd = compute_primary(X,Y)
zbis = compute_zbis(X,Y,rd)
fig, ax = plt.subplots()
plt.pcolor(X, Y, zbis, cmap = 'inferno')
#cset2 = ax.contour(X, Y, zbis, colors='k')
#cs = ax.contourf(X, Y, rd, locator=ticker.LogLocator(), cmap=cm.PuBu_r)
level = [-0.25,-0.1,0,0.05,0.10,0.15,0.25,0.5,0.75]
cs = ax.contourf(X, Y, zbis,level)
cbar = fig.colorbar(cs)
cbar.ax.set_ylabel('z')
plt.ylabel(' y ')
plt.xlabel('x')
plt.title('z computation, inclinaison : {}'.format(str(i*180/np.pi)))
plt.show()
# -
# - There is a huge difference between low inclinaision ( 10 degree ) where a redshift can be spotted on the right and a blueshift ($z<0$) on the left due to rotation of the accretion disk ( Doppler shift due to rotation ). On the other hand for high inclinasion ( 80 degree ) only redshift can be spotted.
# - Redshift values are high above blueshift values. We will then assume that the distorsion induced by the disk rotation can be considered as a little pertubation compared to the distorsion induced by the gravitationnal field.
# When the redshift is positive on the redshift map, the flux will decrease meanwhile when the redshift is negative the flux will increase which is coherent with the formula $F_{obs} = \frac{F_{emis}}{(1+z)^4} $. Thanks to this formula it is then possible to compute the flux.
# +
pas = 200
i_list=[10*np.pi/180,40*np.pi/180,80*np.pi/180]
# Comme image visible fonction de x et y
def Fobsbis(rd,z):
Femis = np.ones((pas,pas))*0
#F0 = 3/4*debm*c**2/(4*np.pi*rs**2)
#thetad = np.arccos( -np.sin(alpha)*np.cos(i)/np.sqrt(1-np.cos(alpha)**2*np.cos(i)**2) )
for k in range(pas):
for l in range(pas):
if not math.isnan(rd[k][l]):
if rd[k][l] >= rmin-3 and rd[k][l] <= rmax +150 :
facteur = rd[k][l]**(-5/2)/(rd[k][l]-3/2)
quotient1 =(np.sqrt(rd[k][l])+np.sqrt(3/2))/(np.sqrt(rd[k][l])-np.sqrt(3/2))
quotient2 = (np.sqrt(2)-1)/(np.sqrt(2)+1)
Femis[k][l] = (facteur*(np.sqrt(rd[k][l])-np.sqrt(3)+np.sqrt(3/8)*np.log(quotient1*quotient2)))/(z[k][l]+1)**4
Femis = Femis.astype('float64')
NAN = np.argwhere(np.isnan(Femis))
for elem in NAN :
Femis[elem[0]][elem[1]]=0
return Femis
# -
# We now obtain an numerical image of a black hole.
for a in range(len(i_list)) :
i = i_list[a]
x = np.linspace(-7,7,pas)
y = np.linspace(-7,7,pas)
X,Y = np.meshgrid(x, y)
rd = compute_primary(X,Y)
zbis = compute_zbis(X,Y,rd)
F = Fobsbis(rd,zbis)
fig, ax = plt.subplots()
plt.pcolor(X, Y, F,cmap = 'gist_heat')
#level = [0.05,0.10,0.15,0.2,0.3,0.5,0.6,0.9,1.2,1.5,1.8,2.1]*10**(-3)
#cs = ax.contourf(X, Y, F,level)
#cset2 = ax.contour(X, Y, F, colors='k')
#cs = ax.contourf(X, Y, F, locator=ticker.LogLocator(), cmap=cm.Wistia)
#cbar = fig.colorbar(cs)
cbar.ax.set_ylabel('Fobs/F0')
plt.ylabel(' y ')
plt.xlabel('x')
plt.title('F/F0 computation, inclinaison : {}'.format(str(i*180/np.pi)))
plt.show()
| Black_hole_image.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # <center> COMP 562 : Inroduction to Machine Learning </center>
# ## <center> <font color='red'>Lecture 3 : Maximum Likeihood Estimation, Linear Regression </font> </center>
# ### <center> <NAME> </center>
# ### <center> Department of Computer Science -- UNC Chapel Hill</center>
# ### <center> August 29, 2018 </center>
# + [markdown] slideshow={"slide_type": "slide"}
# # COMP 562 – Lecture 3
#
# Plan for today
#
# 1. Review likelihood, log-likelihood, maximum likelihood estimates
# 2. Compute maximum likelihood estimate for a Gaussian model
# 3. Introduce linear regression
# 4. Introduce gradient ascent/descent methods
#
# $$
# \renewcommand{\xx}{\mathbf{x}}
# \renewcommand{\yy}{\mathbf{y}}
# \renewcommand{\loglik}{\log\mathcal{L}}
# \renewcommand{\likelihood}{\mathcal{L}}
# \renewcommand{\Data}{\textrm{Data}}
# \renewcommand{\given}{ | }
# \renewcommand{\MLE}{\textrm{MLE}}
# \renewcommand{\Gaussian}[2]{\mathcal{N}\left(#1,#2\right)}
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# # Last Time -- Distributions
#
# Last time we reviewed probability distributions of one dimensional random variables:
# * Bernoulli (coin toss), Binomial (count of heads in multiple coin tosses)
# * Categorical (die roll), Multinomial (count of different die sides in multiple rolls)
#
# Continuous distributions:
# * Gaussian
# * Laplace
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Last Time -- Likelihood
#
# We introduced **likelihood** function, which tells us how well our model fits the data for particular parameters
#
# $$
# \likelihood(\theta|\xx) = p(\underbrace{\xx}_{\Data} \given \underbrace{\theta}_{\textrm{parameters}})
# $$
#
# It is common to assume the training examples are independent and identically distributed,commonly abbreviated to **i.i.d.**, hence:
#
# $$
# \likelihood(\theta|\xx) = \prod_i p(x_i | \theta)
# $$
#
# An example of model
#
# $$
# p(x|\theta) = \theta^x(1-\theta)^{1-x}
# $$
#
# Given data $\xx = \{0,1,1,1\}$, the likelihood function is defined as
#
# $$
# \likelihood(\theta\given\xx) = \prod_i p(x_i\given\theta) = p(x_1\given\theta)p(x_2\given\theta)p(x_3\given\theta)p(x_4\given\theta) = (1-\theta)\theta^3
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# # Last Time -- Log-Likelihood
#
# We introduced log-likelihood function
#
# $$
# \loglik(\theta\given\Data).
# $$
#
# Given data $\Data=\{0,0,0,1\}$ and model $p(x\given\theta) = \theta^x(1-\theta)^{1-x}$
#
# **<font color='red'> Q: Which is more likely (better fit the data) $\theta = 0.25$ or $\theta = 0.99$ </font>**
# + slideshow={"slide_type": "slide"}
import numpy
from __future__ import print_function
def likelihood(theta,xs):
p = 1.0
for x in xs:
p = p*theta**x * (1-theta)**(1-x)
return p
xs = [0,0,0,1]
theta1 = 0.99
theta2 = 0.25
loglik1 = numpy.log(likelihood(theta1,xs))
loglik2 = numpy.log(likelihood(theta2,xs))
print("Log-Likelihood(",theta1,"|",xs, ")=",loglik1)
print("Log-Likelihood(",theta2,"|",xs, ")=",loglik2)
# + [markdown] slideshow={"slide_type": "fragment"}
# $$
# -2.25= \loglik(0.25\given\xx) > \loglik(0.99\given\Data) = -13.83
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# # Last Time -- Maximum Likelihood Estimate (MLE)
#
# We talked about finding a mazimizer of the log-likelihood called **Maximum Likelihood Estimate** (MLE)
#
# $$
# \theta^{\MLE} = \arg\max_{\theta} \loglik(\theta\given\Data)
# $$
#
# which can be interepreted as the parameter for which the data is most probable under the model
# + [markdown] slideshow={"slide_type": "slide"}
# We recalled that in order to maximize a functions, $f(x)$, we finding $x$ for which
#
# $$
# \frac{\partial}{\partial x} f(x) = 0.
# $$
#
# We used this approach to find MLE for a Bernoulli model
#
# $$
# p(x\given\theta) = \theta^x(1-\theta)^{1-x}
# $$
#
# and came up with
#
# $$
# \theta^{\MLE} = \frac{
# \overbrace{ \sum_i [x_i = 1] }^{ \textrm{count of 1s in data} } }{ \underbrace{
# \sum_i [x_i = 1]
# }_{\textrm{count of 1s in data} } +
# \underbrace{
# \sum_i [x_i = 0]
# }_{\textrm{count of 0s in data}}}
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# # Finding $\mu^{MLE}$ of Gaussian Distribution
#
# We left as an exercise a problem to come up with a maximum likelihood estimate for parameter $\mu$ of a Gaussian distribution
#
# $$
# p(x\given\mu,\sigma^2)= \frac{1}{\sqrt{2\pi\sigma^2}} e^{-\frac{1}{2\sigma^2}(x-\mu)^2}
# $$
#
# So we will do that now
# + [markdown] slideshow={"slide_type": "slide"}
# Likelihood function is
#
# $$
# \likelihood(\mu,\sigma^2\given\xx) = \prod_{i=1}^N p(x_i\given\mu,\sigma^2) = \prod_{i=1}^N \frac{1}{\sqrt{2\pi\sigma^2}} e^{-\frac{1}{2\sigma^2}(x_i-\mu)^2}
# $$
#
# Log-likelihood function is
#
# $$
# \log\likelihood(\mu,\sigma^2\given\xx) = \log \prod_{i=1}^N \frac{1}{\sqrt{2\pi\sigma^2}} e^{-\frac{1}{2\sigma^2}(x_i-\mu)^2} = \sum_{i=1}^N \left[-\frac{1}{2}\log{2\pi\sigma^2} -\frac{1}{2\sigma^2}(x_i-\mu)^2\right]
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# # Finding $\mu^{MLE}$ of Gaussian Distribution
#
# Our recipe is:
#
# 1. Take the function you want to maximize:
#
# $$
# f(\mu) = \sum_{i=1}^N \left[-\frac{1}{2}\log{2\pi\sigma^2}-\frac{1}{2\sigma^2}(x_i-\mu)^2\right]
# $$
#
# 2. Compute its first derivative: $\frac{\partial}{\partial \mu} f(\mu)$
# 3. Equate that derivative to zero and solve: $\frac{\partial}{\partial \mu} f(\mu) = 0$
#
# + [markdown] slideshow={"slide_type": "slide"}
#
# The first derivative is
#
# $$
# \frac{\partial}{\partial \mu} f(\mu) = \sum_{i=1}^N \left[ \frac{1}{\sigma^2}(x_i - \mu)\right]
# $$
#
# We equate it to zero and solve
#
# $$
# \sum_{i=1}^N \left[ \frac{1}{\sigma^2}(x_i - \mu)\right] = 0
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# # Finding $\mu^{MLE}$ of Gaussian Distribution
#
# $$
# \begin{aligned}
# \sum_{i=1}^N \left[ - \frac{1}{\sigma^2}(x_i - \mu)\right] &= 0\\
# \sum_{i=1}^N \frac{1}{\sigma^2}(x_i - \mu) &= 0\\
# \sum_{i=1}^N (x_i - \mu) &= 0\\
# \end{aligned}
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# $$
# \begin{aligned}
# \sum_{i=1}^N x_i &= \sum_i \mu \\
# \sum_{i=1}^N x_i &= \mu \sum_i 1 \\
# \frac{\sum_{i=1}^N x_i}{\sum_{i=1}^N 1} &= \mu \\
# \frac{\sum_{i=1}^N x_i}{N} &= \mu
# \end{aligned}
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# # Finding ${\sigma^{2}}^{MLE}$ of Gaussian Distribution
#
# Our recipe is:
#
# 1. Take the function you want to maximize:
#
# $$
# f(\sigma^{2}) = \sum_{i=1}^N \left[-\frac{1}{2}\log{2\pi\sigma^2}-\frac{1}{2\sigma^2}(x_i-\mu)^2\right]
# $$
#
# 2. Compute its first derivative: $\frac{\partial}{\partial \sigma^{2}} f(\sigma^{2})$
# 3. Equate that derivative to zero and solve: $\frac{\partial}{\partial \sigma^{2}} f(\sigma^{2}) = 0$
#
# + [markdown] slideshow={"slide_type": "slide"}
# $$
# f(\sigma^{2}) = \sum_{i=1}^N \left[-\frac{1}{2}\log{2\pi\sigma^2}-\frac{1}{2\sigma^2}(x_i-\mu)^2\right] = - \frac{N}{2}\log{2\pi} - \frac{N}{2}\log{\sigma^2} - \frac{1}{2\sigma^2} \sum_{i=1}^N \left[(x_i-\mu)^2\right]
# $$
#
# The first derivative is
#
# $$
# \frac{\partial}{\partial \sigma^{2}} f(\sigma^{2}) = - \frac{N}{2\sigma^{2}} - \left(\frac{1}{2} \sum_{i=1}^N \left[{(x_i - \mu)}^{2}\right]\right)\frac{\partial}{\partial \sigma^{2}}\left(\frac{1}{\sigma^{2}}\right) \\
# = - \frac{N}{2\sigma^{2}} - \left(\frac{1}{2} \sum_{i=1}^N \left[{(x_i - \mu)}^{2}\right]\right)\left(-\frac{1}{{(\sigma^{2})}^{2}}\right) = \frac{1}{2\sigma^{2}} \left(\frac{1}{\sigma^{2}} \sum_{i=1}^N \left[{(x_i - \mu)}^{2} \right] - N \right)
# $$
#
# Which, if we rule out $\sigma^{2} = 0$, is equal to zero only if
#
# $$
# \sigma^{2} = \frac{1}{N} \sum_{i=1}^N \left[{(x_i - \mu)}^{2} \right]
# $$
#
# **<font color='red'> Please Verify both ${\mu}^{MLE}$ and ${\sigma^{2}}^{MLE}$ using seconed derivative test </font>**
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Using MLE Estimates in Prediction
#
# What can we do with maximum likelihood estimates?
#
# We can predict outcome of the next experiment:
#
# * Given data $\{1,0,1,1,0,0,0,0,0\}$ and assuming Bernoulli model
#
# * $\theta^{\MLE}$ is $\frac{3}{3 + 5} = \frac{3}{8}$
#
# * Then probability that the next experiment will yield $x=1$ is $\frac{3}{8}$
# + [markdown] slideshow={"slide_type": "slide"}
# * Given heights of 6th graders in inches $\{56,57,59,63,55,61\}$ and assuming Gaussian model
#
# * $\mu^{\MLE} = 58.5$ and ${\sigma}^{MLE} = 3.082$
#
# * Hence, probability that the next 6th grader will be of height 58.5 $\pm$ 2*3.082 is 0.95
#
# <img src="./Images/Standard_deviation_diagram.png" width="600" align="center"/>
#
# Given realizations of the same random variable we can estimate what the next draw might look like
#
# **<font color='red'> Q: Suppose you wanted to predict a student's grade in COMP 562? What would you do? </font>**
# + [markdown] slideshow={"slide_type": "slide"}
# # Linear Regression
#
# One of the simplest examples of supervised learning is **linear regression**
#
# Aim to model the relationship between an outcome variable $y$ (a.k.a dependent variable or response variable), and a vector of explanatory variables $\mathbf{x}=(x_1,x_2,…,x_N)$ (independent variables, covariates, or features)
#
# For example, we can try to predict student's grade in COMP562 class using grades from prereq's and number of beers a student has each week
#
# $$
# \begin{aligned}
# \textrm{COMP562} &= 0.25 + 0.2*\textrm{COMP410} + 0.3*\textrm{MATH233} + 0.5*\textrm{STOR435} - 0.1* \textrm{beers} + \epsilon
# \end{aligned}
# $$
#
# Where,
#
# $$
# \begin{aligned}
# \epsilon &\sim \mathcal{N}(0, 0.1)
# \end{aligned}
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# Formally, we would write
#
# $$
# \begin{aligned}
# y &= \beta_0 + \sum_j x_j \beta_j + \epsilon \\
# \epsilon &\sim \Gaussian{0}{\sigma^2}
# \end{aligned}
# $$
#
# or more compactly
#
# $$
# y \given \xx \sim \Gaussian{\beta_0 + \sum_j x_j \beta_j}{ \sigma^2}
# $$
#
# Notice that the function is linear in the parameters $\beta=(\beta_0,\beta_1,…,\beta_p)$, not necessarily in terms of the covariates
# + [markdown] slideshow={"slide_type": "slide"}
# # Linear Regression
#
# Taking a closer look at the model:
#
# $$
# y \given \xx \sim \Gaussian{\beta_0 + \sum_{j=1}^p x_j \beta_j}{ \sigma^2}
# $$
#
# * $y$ is a target variable we are modeling
# * $\xx$ is a vector of $p$ features (aka predictors and covariates)
# * $\beta_0$ is a **bias** which does not depent on the features
# * $\beta_1,\dots,\beta_p$ is a vector of weights (one per feature)
# * $\sigma^2$ is variance -- how far can we expect $y$ to be away from $\beta_0 + \sum_{j=1}^p x_j \beta_j$
#
# **<font color='red'> Q: Which of these items above are parameters? Hint: Which values can we adjust to get a better prediction?</font>**
#
# $$
# \begin{aligned}
# \textrm{COMP562} &= 0.25 + 0.2*\textrm{COMP410} + 0.3*\textrm{MATH233} + 0.5*\textrm{STOR435} - 0.1* \textrm{beers} + \epsilon \\
# \end{aligned}
# $$
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Linear Regression
#
# Probability of target variable $y$
#
# $$
# p(y\given\xx,\beta_0,\beta,\sigma^2) = \frac{1}{\sqrt{2\pi\sigma^2}} \exp\left\{-\frac{1}{2\sigma^2}\left(y_i-\underbrace{(\beta_0 + \sum_j x_j \beta_j)}_{\textrm{mean of the Gaussian}}\right)^2\right\}
# $$
#
# In the case of the 6th grader's height, we made **the same** prediction for any other 6th grader (58.5 inches)
#
# In our COMP 562 grade example, we compute a potentially different mean for every student
#
# $$
# \beta_0 + \beta_{\textrm{COMP410}}*\textrm{COMP410} + \beta_{\textrm{MATH233}}*\textrm{MATH233} + \beta_{\textrm{STOR435}}*\textrm{STOR435} + \beta_{\textrm{beers}}* \textrm{beers}
# $$
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Linear Regression -- Toy Example
#
# We can try to fit this model to some data (**not real UNC data**)
# + slideshow={"slide_type": "fragment"}
import pandas
X = numpy.asarray([[3.0,3.0,4.0,0.0],[3.0,3.0,3.0,0.0],[3.0,2.0,2.0,5.0],
[2.0,2.0,4.0,0.0],[3.0,3.0,4.0,4.0],[4.0,4.0,3.0,0.0],
[2.0,2.0,4.0,0.0],[2.0,2.0,2.0,0.0],[3.0,4.0,2.0,5.0],
[2.0,2.0,3.0,0.0],[2.0,4.0,2.0,0.0],[3.0,3.0,3.0,0.0],
[2.0,2.0,4.0,5.0],[4.0,3.0,2.0,6.0],[3.0,4.0,4.0,4.0],
[3.0,2.0,2.0,6.0],[3.0,3.0,3.0,0.0],[3.0,2.0,4.0,0.0],
[3.0,3.0,4.0,4.0],[3.0,4.0,2.0,0.0]])
y = numpy.asarray([4.0,3.0,2.0,3.0,3.0,4.0,3.0,2.0,3.0,
3.0,3.0,3.0,2.0,2.0,3.0,2.0,3.0,3.0,3.0,3.0])
#Creating pandas dataframe from numpy array
dataset = pandas.DataFrame({'x_1:COMP410':X[:,0],'x_2:MATH233':X[:,1],'x_3:STOR345':X[:,2],'x_4:#beers':X[:,3],'y:COMP562':y})
dataset.head()
# + slideshow={"slide_type": "slide"}
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy
features = ['COMP410','MATH233','STOR435','Beers']
for j in range(X.shape[1]):
plt.subplot(2,3,j+1)
plt.hist(X[:,j])
plt.title(features[j])
plt.subplot(2,3,5)
print(y)
plt.hist(y)
plt.title('COMP562 -- target')
plt.tight_layout() # ensure sensible layout of subplots
# + [markdown] slideshow={"slide_type": "slide"}
# # Linear Regression -- Toy Example
#
# Our goal is to fit the model that predicts COMP 562 grade
#
# Our prediction is based on
#
# $$
# \begin{aligned}
# \textrm{COMP562} &= \beta_0 + \beta_{\textrm{COMP410}}*\textrm{COMP410} +
# \beta_{\textrm{MATH233}}*\textrm{MATH233} \\
# &+ \beta_{\textrm{STOR435}}*\textrm{STOR435} +
# \beta_{\textrm{beers}}* \textrm{beers} + \epsilon \\
# \end{aligned}
# $$
#
# In order to make a prediction we need
#
# * $\beta_0$ -- bias or grade you get regardless of your other grades
# * $\beta_{\textrm{COMP410}}$
# * $\beta_{\textrm{MATH233}}$
# * $\beta_{\textrm{STOR435}}$
# * $\beta_{\textrm{beers}}$
#
# Note that having $\sigma^2$ would enable us to give a range of grades that cover 95% of the probability
# + [markdown] slideshow={"slide_type": "slide"}
# # Linear Regression -- Likelihood
#
# We start by writing out a likelihood for linear regression is
#
# $$
# \likelihood(\beta_0,\beta,\sigma^2\given\xx,\yy) =
# \prod_{i=1}^N p(y\given\xx,\beta_0,\beta,\sigma^2) =
# \prod_{i=1}^N \frac{1}{\sqrt{2\pi\sigma^2}} \exp\left\{-\frac{1}{2\sigma^2}\left(y_i-(\beta_0 + \sum_j x_j \beta_j)\right)^2\right\}
# $$
#
# Log-likelihood for linear regression is
#
# $$
# \log\likelihood(\beta_0,\beta,\sigma^2\given\xx,\yy) = \sum_{i=1}^N \left[ -\frac{1}{2}\log 2\pi\sigma^2 -\frac{1}{2\sigma^2}\left(y_i-(\beta_0 + \sum_j x_j \beta_j)\right)^2\right] \\ = - \frac{N}{2}\log(2\pi\sigma^2) -\frac{1}{2\sigma^2} \sum_{i=1}^N \left(y_i-(\beta_0 + \sum_j x_{i,j} \beta_j)\right)^2 = - \frac{N}{2}\log(2\pi\sigma^2) -\frac{RSS}{2\sigma^2}
# $$
#
# + [markdown] slideshow={"slide_type": "slide"}
# We will refer to expression $y_i-(\beta_0 + \sum_j x_j \beta_j)$ as **residual**, and hence **RSS** stands for **residual sum of squares** or **sum of squared errors** and is defined by
#
# $$
# RSS = \sum_{i=1}^N \left(y_i-(\beta_0 + \sum_j x_{i,j} \beta_j)\right)^2
# $$
#
# And RSS/N is called the **mean squared error** or **MSE**
#
# $$
# MSE = \frac{1}{N}\sum_{i=1}^N \left(y_i-(\beta_0 + \sum_j x_{i,j} \beta_j)\right)^2
# $$
#
# Hence, maximizing log-likelihood is equivalent to minimizing RSS or MSE
# + [markdown] slideshow={"slide_type": "slide"}
# Our goal is still to find $\beta_0,\beta$ such that
#
# $$
# \begin{aligned}
# \frac{\partial}{\partial \beta_0} \log\likelihood(\beta_0,\beta,\sigma^2\given\xx,\yy) &= 0 \\
# \frac{\partial}{\partial \beta_1} \log\likelihood(\beta_0,\beta,\sigma^2\given\xx,\yy) &= 0 \\
# \cdots&\\
# \frac{\partial}{\partial \beta_p} \log\likelihood(\beta_0,\beta,\sigma^2\given\xx,\yy) &= 0
# \end{aligned}
# $$
#
# Because that will guarantee the parameters can not be further changed to improve the likelihood
# + [markdown] slideshow={"slide_type": "slide"}
# # Introducing Gradient Ascent
#
# Previously, we solved equations of type $\frac{\partial}{\partial \theta} \loglik(\theta\given\Data) = 0$ in a closed-form
#
# Here, we will develop a different approach using numerical optimization
#
# Let's first consider how we can maximize a univariate differentiable function $f(x)$ iteratively
# + slideshow={"slide_type": "slide"}
x = numpy.arange(0.0,10.0,0.1)
f = -(x-4.0)**2.0 - 0.5*(x-6.0)**2.0
dfdx = -(x-4.0)*2 - (x-6.0)
plt.plot(x,f)
plt.xlabel('x')
plt.ylabel('f(x)')
plt.plot(x[10],f[10],'ro')
plt.annotate('Starting point',(x[10]+0.25,f[10]))
x1 = x[10]-1
x2 = x[10]+1.5
f1 = f[10]-dfdx[10]*1
f2 = f[10]+dfdx[10]*1.5
plt.plot([x1,x2],[f1,f2])
plt.annotate('Linear\napproximation',(x2,f2-5))
# + [markdown] slideshow={"slide_type": "fragment"}
# **<font color='red'> Q: Starting from the labeled point, should we increase or decrease $x$ to get to the maximum of function $f(x)$? Why? </font>**
# + [markdown] slideshow={"slide_type": "slide"}
# Another way to see this is to consider a very simplified version of Taylor's theorem
#
# **Theorem.** Given a function $f(\cdot)$ which is smooth at $x$
#
# $$
# f(x + d) = f(x) + f'(x)d + O(d^2)
# $$
#
# In words, close to $x$ function $f(\cdot)$ is very close to being a linear function of $d$
#
# $$
# f(x + \color{blue}{d}) = f(x) + f'(x)\color{blue}{d}
# $$
#
# Slope of the best linear approximation is $f'(x)$, i.e.,$\hspace{0.5em}$$f'(x)$ tells us in which direction function grows
#
# <img src="./Images/Derivative.gif" width="700" align="center"/>
#
# + [markdown] slideshow={"slide_type": "slide"}
# Multivariate functions are a bit harder to visualize, so we view them like topographical maps
#
# Contours connect nearby points with the same altitutde (function value)
#
# <img src="./Images/Topographic_Image.png" width="1400" align="center"/>
# + slideshow={"slide_type": "slide"}
import matplotlib.cm as cm
x = numpy.arange(-5.0,10.0,0.1)
y = numpy.arange(-5.0,10.0,0.1)
X,Y = numpy.meshgrid(x,y)
F = -(X-1.0)**2.0 - (Y-2.0)**2.0 + 0.5*X*Y
fig = plt.figure(figsize=plt.figaspect(0.35))
from mpl_toolkits.mplot3d import Axes3D
ax = fig.add_subplot(1, 2, 1, projection='3d')
ax.plot_surface(X,Y,F)
plt.tight_layout(6.0)
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_zlabel('f(x,y)')
ax.plot([x[130]],[y[10]],[F[130,10]],'ro')
ax.plot([x[70]],[y[70]],[F[70,70]],'go')
ax.plot([x[100]],[y[30]],[F[100,30]],'yo')
ax = fig.add_subplot(1,2,2)
C = ax.contour(X,Y,F,levels=numpy.arange(numpy.min(F),numpy.max(F),3),cmap=plt.cm.rainbow)
ax.clabel(C, inline=4, fontsize=10)
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.plot(x[130],y[10],'ro')
ax.plot(x[70],y[70],'go')
ax.plot(x[100],y[30],'yo')
# + slideshow={"slide_type": "slide"}
import matplotlib.cm as cm
x = numpy.arange(-5.0,10.0,0.1)
y = numpy.arange(-5.0,10.0,0.1)
X,Y = numpy.meshgrid(x,y)
F = -(X-1.0)**2.0 - (Y-2.0)**2.0 + 0.5*X*Y
dfdx = -2.0*(X-1.0) + 0.5*Y
dfdy = -2.0*(Y-2.0) + 0.5*X
C = plt.contour(X,Y,F,levels=numpy.arange(numpy.min(F),numpy.max(F),2),cmap=plt.cm.rainbow)
#ax.clabel(C, inline=4, fontsize=10)
plt.annotate('Starting point',(x[20]+0.25,y[20]),fontsize=12,fontweight='bold')
plt.plot(x[20],y[20],'ro')
x1 = x[20] -0.2*dfdx[20,20]
y1 = y[20] -0.2*dfdy[20,10]
x2 = x[20] +0.2*dfdx[20,20]
y2 = y[20] +0.2*dfdy[20,20]
plt.annotate('Graident is the direction of the \nat largest change the starting point',(x2+0.25,y2-0.5),fontsize=12,fontweight='bold')
plt.xlabel('x')
plt.ylabel('y')
plt.plot([x1,x2],[y1,y2])
# + [markdown] slideshow={"slide_type": "slide"}
# Much as in the case of univariate function, the direction in which the function changes is described by derivatives
#
# A vector composed of partial derivatives of a function is called gradient
#
# $$
# \nabla f(\xx) = \left[\begin{array}{c} \frac{\partial}{\partial x_1} f(\xx) \\ \vdots \\ \frac{\partial}{\partial x_p} f(\xx) \end{array}\right]
# $$
#
# For example,
#
# $$
# f(x,y) = -(x-1)^2 - (y- 2)^2 + \frac{1}{2}xy
# $$
#
# has gradient
#
# $$
# \nabla f(x,y) = \left[\begin{array}{c} -2(x-1) + \frac{1}{2}y \\ -2(y-2) + \frac{1}{2}x \end{array}\right]
# $$
#
# Gradient points in the direction of the largest change in the function
#
# **<font color='red'> Q: Can you come up with an algorithm that would use the gradients to maximize a function? </font>**
# + [markdown] slideshow={"slide_type": "slide"}
# Gradients are important because:
#
# 1. They can tell us when we have reached an optimum ($\nabla f(\xx) = 0$)
# 2. Point in the direction in which function is changing the most
#
# So, we shouldn't mess them up
#
# To check your gradient go back to the definition:
#
# $$
# \frac{\partial}{\partial x_1} f(x_1,\dots,x_p) = \lim_{\delta \rightarrow 0} \frac{f(x_1 + \delta/2,\dots,x_p) - f(x_1 - \delta/2,\dots,x_p)}{\delta}
# $$
#
# Left side can be numerically approximated using a small $\delta=10^{-6}$:
#
# $$
# \frac{f(x_1 + 0.5\delta,\dots,x_p) - f(x_1 - 0.5\delta,\dots,x_p)}{\delta}
# $$
#
# This is called **finite difference approximation**, let's see this in practice
# + slideshow={"slide_type": "slide"}
def grad_check(f,xy0,delta=1e-6,tolerance=1e-7):
f0,g0 = f(xy0)
p = len(xy0)
finite_diff = numpy.zeros(p)
gradient_correct = True
for i in range(p):
xy1 = numpy.copy(xy0)
xy2 = numpy.copy(xy0)
xy1[i] = xy1[i] - 0.5*delta
xy2[i] = xy2[i] + 0.5*delta
f1,_ = f(xy1)
f2,_ = f(xy2)
finite_diff = (f2 - f1)/(delta)
if (abs(finite_diff - g0[i])>tolerance):
print("Broken partial",i," Finite Diff: ",finite_diff," Partial: ",g0[i])
gradient_correct = False
return gradient_correct
def f_broken(xy):
x = xy[0]
y = xy[1]
f = -(x-1.0)**2.0 - (y-2.0)**2.0 + 0.5*x*y
dfdx = -2.0*(x-1.0) + 0.5*y
dfdy = -2.0*(y-2.0) - 0.5*x
grad = [dfdx,dfdy]
return f,grad
def f_correct(xy):
x = xy[0]
y = xy[1]
f = -(x-1.0)**2.0 - (y-2.0)**2.0 + 0.5*x*y
dfdx = -2.0*(x-1.0) + 0.5*y
dfdy = -2.0*(y-2.0) + 0.5*x
grad = [dfdx,dfdy]
return f,grad
xy = numpy.asarray([1.0,1.0])
print("f_broken has correct gradient:", grad_check(f_broken,[1.0,1.0]))
print("f_correct has correct gradient:", grad_check(f_correct,[1.0,1.0]))
# + [markdown] slideshow={"slide_type": "slide"}
# Once we are sure that the gradients are correct, we can proceed to optimize the function
#
# To do that, we use **gradient ascent (GA)** or **gradient descent (GD)** depending on whether we are maximizing or minimizing a function
#
# Recall that we have $f : \mathbb{R}^{n} \rightarrow \mathbb{R}$, convex and differentiable
#
# We want to solve
# $$
# \begin{aligned}
# GA: \, \mathop{\textrm{argmax}}_{\mathbf{\theta} \in \mathbb{R}^{n}} f(\mathbf{\theta}) \; \; \; \; \; \; GD: \, \mathop{\textrm{argmin}}_{\mathbf{\theta} \in \mathbb{R}^{n}} f(\mathbf{\theta})
# \end{aligned}
# $$
#
# <img src="./Images/OptimizationProblems.png" align="center"/>
#
# + [markdown] slideshow={"slide_type": "slide"}
# * Gradient Ascent\Descent: Choose initial ${\mathbf{\theta}^{(0)}} \in \mathbb{R}^{n}$, repeat:
# $$ \;
# \begin{aligned}
# {\mathbf{\theta}^{(k)}} = {\mathbf{\theta}^{(k-1)}} \pm t_{k}.\nabla f({\mathbf{\theta}^{(k-1)}}), k =1,2,3,\ldots
# \end{aligned}
# $$
# Where $t_{k}$ is the step size (learning rate) at step $k$
#
# * Stop at some point using a stopping criteria (depend on the problem we are solving), for example:
# * Maximum number of iterations reached
# * $| f({\mathbf{\theta}^{(k)}}) − f({\mathbf{\theta}^{(k-1)}}) | < \epsilon$
#
# <img src="./Images/GD.gif" align="center"/>
# + [markdown] slideshow={"slide_type": "slide"}
# **<font color='red'> Q: What would you change for gradient descent? How would you specify the learning rate $(t_{k})$?</font>**
# + [markdown] slideshow={"slide_type": "fragment"}
# Choosing a step size or learning rate ($t_{k}$)
#
# 1. Fixed step size
# * Simply take $t_{k} = t$ $\forall$ $k$
# * However, can diverge if $t$ is too large, and convergence can be slow if $t$ is too small
#
# <img src="./Images/LearningRate.gif" align="center"/>
# + [markdown] slideshow={"slide_type": "slide"}
# 2. Use Line search Strategy
# * At each iteration, do the best you can along the direction of the gradient,
#
# $$
# \begin{aligned}
# t = \mathop{\textrm{argmax}}_{s \geq 0} f(\mathbf{\theta} + s.\nabla f({\mathbf{\theta}}))
# \end{aligned}
# $$
#
# <img src="./Images/Line_Search1.gif" align="center"/>
#
# * Usually, it is not possible to do this minimization exactly, and approximation methods are used
# * Backtracking Line Search:
# * Choose an initial learning rate ($t_{k} = t_{init})$, and update your parameters ${\mathbf{\theta}^{(k)}} = {\mathbf{\theta}^{(k-1)}} \pm t_{k}.\nabla f({\mathbf{\theta}^{(k-1)}})$
# * Reduce learning rate $t_{k} = \alpha . t_{init}$, where $0< \alpha <1 $
# * Repeat by reducing $\alpha$ till you see an improvmnet in $f({\mathbf{\theta}^{(k)}})$
# + slideshow={"slide_type": "slide"}
def gradient_ascent(f,theta,init_step,iterations):
f_val,grad = f(theta) # compute function value and gradient
f_vals = [f_val]
for it in range(iterations): # iterate for a fixed number of iterations
done = False # initial condition for done
line_search_it = 0 # how many times we tried to shrink the step
step = init_step # reset step size to the initial size
while not done and line_search_it<100: # are we done yet?
new_theta = theta + step*grad # take a step along the gradient
new_f_val,new_grad = f(new_theta) # evaluate function value and gradient
if new_f_val<f_val: # did we go too far?
step = step*0.95 # if so, shrink the step-size
line_search_it += 1 # how many times did we shrank the step
else:
done = True # better than the last theta, so we move on
if not done: # did not find right step size
print("Line Search failed.")
else:
f_val = new_f_val # ah, we are ok, accept the new theta
theta = new_theta
grad = new_grad
f_vals.append(f_val)
plt.plot(f_vals)
plt.xlabel('Iterations')
plt.ylabel('Function value')
return f_val, theta
# + [markdown] slideshow={"slide_type": "slide"}
# # Linear Regression -- Likelihood
#
# We start by writing out a likelihood for linear regression is
#
# $$
# \likelihood(\beta_0,\beta,\sigma^2\given\xx,\yy) =
# \prod_{i=1}^N p(y\given\xx,\beta_0,\beta,\sigma^2) =
# \prod_{i=1}^N \frac{1}{\sqrt{2\pi\sigma^2}} \exp\left\{-\frac{1}{2\sigma^2}\left(y_i-(\beta_0 + \sum_j x_j \beta_j)\right)^2\right\}
# $$
#
# Log-likelihood for linear regression is
#
# $$
# \log\likelihood(\beta_0,\beta,\sigma^2\given\xx,\yy) = \sum_{i=1}^N \left[ -\frac{1}{2}\log 2\pi\sigma^2 -\frac{1}{2\sigma^2}\left(y_i-(\beta_0 + \sum_j x_j \beta_j)\right)^2\right].
# $$
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Linear Regression -- Gradient of Log-Likelihood
#
# Partial derivatives
#
# $$
# \begin{aligned}
# \frac{\partial}{\partial \beta_0} \log\likelihood(\beta_0,\beta,\sigma^2\given\xx,\yy) &= \sum_{i=1}^N -\frac{1}{\sigma^2}\left(y_i-(\beta_0 + \sum_j x_j \beta_j)\right)(-1)\\
# \frac{\partial}{\partial \beta_k} \log\likelihood(\beta_0,\beta,\sigma^2\given\xx,\yy) &= \sum_{i=1}^N -\frac{1}{\sigma^2}\left(y_i-(\beta_0 + \sum_j x_j \beta_j)\right)(-x_k)&,k\in\{1,\dots,p\}
# \end{aligned}
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# Hence gradient (with respect to $\beta$s)
# $$
# \nabla \loglik(\beta_0,\beta,\sigma^2\given\xx,\yy) = \left[\begin{array}{c}
# \sum_{i=1}^N -\frac{1}{\sigma^2}\left(y_i-(\beta_0 + \sum_j x_j \beta_j)\right)(-1) \\
# \sum_{i=1}^N -\frac{1}{\sigma^2}\left(y_i-(\beta_1 + \sum_j x_j \beta_j)\right)(-x_1) \\
# \vdots\\
# \sum_{i=1}^N -\frac{1}{\sigma^2}\left(y_i-(\beta_0 + \sum_j x_j \beta_j)\right)(-x_p)
# \end{array}
# \right]
# $$
# + slideshow={"slide_type": "slide"}
X = numpy.asarray([[3.0,3.0,4.0,0.0],[3.0,3.0,3.0,0.0],[3.0,2.0,2.0,5.0],
[2.0,2.0,4.0,0.0],[3.0,3.0,4.0,4.0],[4.0,4.0,3.0,0.0],
[2.0,2.0,4.0,0.0],[2.0,2.0,2.0,0.0],[3.0,4.0,2.0,5.0],
[2.0,2.0,3.0,0.0],[2.0,4.0,2.0,0.0],[3.0,3.0,3.0,0.0],
[2.0,2.0,4.0,5.0],[4.0,3.0,2.0,6.0],[3.0,4.0,4.0,4.0],
[3.0,2.0,2.0,6.0],[3.0,3.0,3.0,0.0],[3.0,2.0,4.0,0.0],
[3.0,3.0,4.0,4.0],[3.0,4.0,2.0,0.0]])
Y = numpy.asarray([4.0,3.0,2.0,3.0,3.0,4.0,3.0,2.0,3.0,
3.0,3.0,3.0,2.0,2.0,3.0,2.0,3.0,3.0,3.0,3.0])
def linear_regression_log_likelihood(Y,X,betas,sigma2=1.0):
ll = 0
beta0 = betas[0]
beta = betas[1:]
dlldbeta0 = 0
dlldbeta = numpy.zeros(len(beta))
for (x,y) in zip(X,Y):
ll = ll -0.5*numpy.log(2*numpy.pi*sigma2)
res = y - beta0 - numpy.sum(x*beta)
ll = ll - 1.0/(2.0*sigma2)*(res**2.0)
dlldbeta0 = dlldbeta0 - 1.0/sigma2*res*(-1)
dlldbeta = dlldbeta - 1.0/sigma2*(res*(-x))
grad = numpy.zeros(len(beta)+1)
grad[0] = dlldbeta0
grad[1:] = dlldbeta
return ll, grad
init_beta = [0.1]*5
f = lambda betas: linear_regression_log_likelihood(Y,X,betas)
grad_check(f,init_beta)
# + [markdown] slideshow={"slide_type": "slide"}
# # Fitting Linear Regression Using Gradient Ascent
#
# We are now ready to optimize our model
#
# We will aim to find maximum of the log-likelihood function using the gradient ascent algorithm
# we implemented
# + slideshow={"slide_type": "fragment"}
init_beta = [0.1]*5
init_step = 0.01
iterations = 1000
f = lambda betas: linear_regression_log_likelihood(Y,X,betas)
[f_best,betas_mle] = gradient_ascent(f,init_beta,0.01,100)
names = ['0','COMP410','MATH233','STOR435','Beers']
for (name,beta) in zip(names,betas_mle):
print('Beta',name,'=',beta)
# + [markdown] slideshow={"slide_type": "slide"}
# # Today
#
# * Introduced linear regression
# * Introduced gradient ascent/descent
# * Implemented linear regression log-likelihood and gradient ascent
# * Fit a simple model for COMP 562 grades
| CourseMaterial/COMP562_Lect3/COMP562_Lect3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/FUDGEMASTER/Breast_cancer_case-study/blob/master/Hackathon.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="n9GCHQJlWW45"
# **Importing Libraries**
#
#
# + id="n2tHFaBqWc_M" outputId="ef77aa98-88de-4bb8-9f25-b523d6af13f8" colab={"base_uri": "https://localhost:8080/"}
import pandas as pd
import numpy as np
from sklearn.impute import SimpleImputer
from scipy import stats
from mlxtend.preprocessing import minmax_scaling
import seaborn as sns
import matplotlib.pyplot as plt
from scipy.stats import kurtosis
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
from sklearn.dummy import DummyClassifier
from imblearn.over_sampling import SMOTE
from sklearn.model_selection import RandomizedSearchCV
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import make_scorer
from sklearn.metrics import accuracy_score
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
from sklearn.metrics import f1_score
from sklearn.model_selection import cross_validate
from sklearn.linear_model import LogisticRegression
from sklearn.svm import LinearSVC
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.feature_selection import SelectKBest, f_classif
from sklearn.model_selection import cross_val_score
from sklearn.neural_network import MLPClassifier
from sklearn.metrics import confusion_matrix
from xgboost import XGBClassifier
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import MinMaxScaler
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.decomposition import PCA
from sklearn.svm import SVC
# + id="_gVbJNrNSNpK"
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
# + id="F3xS0g4qSVxa"
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
# + id="bK_VzTQoSa2k"
downloaded = drive.CreateFile({'id':"1FttBvXlR5lC45E2Y3INmpJeuiAXhytfj"})
downloaded.GetContentFile('hackathon dataset.csv')
# + id="QBowvJlIWhsH" outputId="44eeb16d-7cd6-4144-834e-be07e562d04f" colab={"base_uri": "https://localhost:8080/", "height": 195}
hackathon = pd.read_csv('hackathon dataset.csv') #Load the dataset
hackathon.head()
# + [markdown] id="NGGWJs2gcrU0"
# ## **Renaming Columns**
# + id="bgltlcSiXWQk" outputId="cfcf7fdd-e265-41c7-e8e1-aacc0d2016fb" colab={"base_uri": "https://localhost:8080/", "height": 195}
new_row = pd.DataFrame({'1':1, '39':39, '4':4, '0':0, '0.1':0.1, '0.2':0.2, '0.3':0.3, '0.4':0.4, '0.5':0.5, '195':195, '106':106, '70':70, '26.97':26.97, '80':80,'77':77, '0.6':0.6 }, index =[0])
hackathon = pd.concat([new_row, hackathon]).reset_index(drop = True)
#Appending a row at the beginning of the dataset with the same value as columnname
hackathon.head(5)
# + id="KosyfXDyZlIm" outputId="cfcf31dd-b926-49e8-ffcb-a48fecba0cb5" colab={"base_uri": "https://localhost:8080/", "height": 195}
hackathon = hackathon.rename(columns = {'1':1, '39':'2', '4':'3',
'0':'4', '0.1':'5', '0.2':'6',
'0.3':'7', '0.4':'8', '0.5':'9', '195':'10', '106':'11', '70':'12', '26.97':'13', '80':'14','77':'15', '0.6':'16' })
hackathon.head()
# + id="zJx0IINTbTpm" outputId="2a863bae-aa2d-43db-fa75-8cd674aec7ed" colab={"base_uri": "https://localhost:8080/", "height": 402}
#hackathon = hackathon.drop(1) #dropping duplicated rows
hackathon
# + id="EYYdEqjicJqq" outputId="dbf15038-53af-406c-8358-957f658f9896" colab={"base_uri": "https://localhost:8080/", "height": 419}
hackathon = hackathon.reset_index(drop = True) #resetting index due to dropping duplicate rows
hackathon
# + id="d11Z65XMolvi" outputId="467f2ce6-0d97-4f15-c3cd-e1ff9fd810a4" colab={"base_uri": "https://localhost:8080/", "height": 419}
hackathon.iloc[0,4:9] = 0
hackathon.iloc[0,15] = 0
hackathon
# + [markdown] id="tJcCvttQdBKn"
# # **Step 1: Check for Missing Values and Edit**
#
#
# + id="Xb1HzUb6cX7d" outputId="862d68e9-69ca-4ec9-f8a2-a53acbc33e33" colab={"base_uri": "https://localhost:8080/"}
missing_values_count = hackathon.isnull().sum() #count the no. of missing values per column
missing_values_count
# + id="dIPeFK1ifyMc"
total_cells = np.product(hackathon.shape) #total cells in the dataset
total_missing = missing_values_count.sum() #cells with missing values(NaN)
# + id="7Ha7okXIg1HH" outputId="110b9e03-0ddc-4334-8a04-c56b6c7d72b4" colab={"base_uri": "https://localhost:8080/"}
percent_missing = (total_missing/total_cells)*100
percent_missing #percentage of values missing
# + id="PJTqh4AqhHcn"
#as data missing is the dataset is a low percentage, we use imputation as dropping can lead to loss of data
my_imputer = SimpleImputer()
# + id="VcaUGSE8hh4Y"
imputed_hackathon = pd.DataFrame(my_imputer.fit_transform(hackathon)) #Checkpoint 1
# + id="xxAOCsVlhlAc" outputId="ee77facf-62bf-43dc-847e-0e3efdde4d63" colab={"base_uri": "https://localhost:8080/", "height": 419}
imputed_hackathon.columns = hackathon.columns #we put back column names removed by imputation
imputed_hackathon
# + id="rQAlhTy-iu5r" outputId="d8056afa-59a9-4021-a522-3e614ac899de" colab={"base_uri": "https://localhost:8080/"}
#Checking Imputation Effect
missing_values_count = imputed_hackathon.isnull().sum() #count the no. of missing values per column
missing_values_count
# + [markdown] id="-Nr_ZsVy88WX"
# ## **Detecting and Removing Outliers**
# + id="YEWOxYPF9C3Q" outputId="dfa3a165-a98e-41f5-d469-7cfe0cbc7d8c" colab={"base_uri": "https://localhost:8080/", "height": 279}
#Column 3
#Comparing scatterplot of every numerical data column with the output column
fig,ax = plt.subplots()
ax.scatter(x = imputed_hackathon['2'], y = imputed_hackathon['16'])
plt.ylabel('Prediction')
plt.xlabel('3')
plt.show()
#no outliers
# + id="qmt17zPu9M_g" outputId="67a75f4d-a59a-498e-fbd8-c2afc238dd87" colab={"base_uri": "https://localhost:8080/", "height": 279}
#Column 10
fig,ax = plt.subplots()
ax.scatter(x = imputed_hackathon['10'], y = imputed_hackathon['16'])
plt.ylabel('Prediction')
plt.xlabel('10')
plt.show()
# + id="-m8mu0bz9e37" outputId="1dde90fb-8e27-472b-ec4d-d7645f000030" colab={"base_uri": "https://localhost:8080/", "height": 111}
#from the scatterplot, we see that values greater then 500 are anomalies
imputed_hackathon[imputed_hackathon['10'] > 500]
# + id="G3eK-3TD9h94"
#taking mean value of the column to replace the outlier values
mean10 = np.mean(imputed_hackathon['10'])
# + id="mWjBechF9r3L"
#replacing outliers
imputed_hackathon.iloc[1111,9] = mean10
imputed_hackathon.iloc[3160,9] = mean10
# + id="0jDcuVUS9zNx" outputId="705a80ea-141e-4eb4-9257-20227f3c04a2" colab={"base_uri": "https://localhost:8080/", "height": 279}
fig,ax = plt.subplots()
ax.scatter(x = imputed_hackathon['10'], y = imputed_hackathon['16'])
plt.ylabel('Prediction')
plt.xlabel('10')
plt.show()
#the plot after outliers have been eliminated
# + id="qASIp0qw934l" outputId="d4a2869f-da1e-4b9f-d134-782b853ac10d" colab={"base_uri": "https://localhost:8080/", "height": 279}
#Column 11
fig,ax = plt.subplots()
ax.scatter(x = imputed_hackathon['11'], y = imputed_hackathon['16'])
plt.ylabel('Prediction')
plt.xlabel('11')
plt.show()
# + id="2vi0Xl71-HW3" outputId="d494634c-35d9-47f6-951c-29ecd0e1de79" colab={"base_uri": "https://localhost:8080/", "height": 80}
imputed_hackathon[imputed_hackathon['11']>250]
# + id="f4OU2utA-Kx3" outputId="7e75369a-e958-4b15-b12b-b41cc5af6fd6" colab={"base_uri": "https://localhost:8080/"}
mean11 = np.mean(imputed_hackathon['11'])
mean11
# + id="8dF_zSnT-YVv"
imputed_hackathon.iloc[481,10] = mean11
# + id="wBQXH7Sf-QHD" outputId="caadd5f0-dcca-4197-cbd5-5404ba63572b" colab={"base_uri": "https://localhost:8080/", "height": 279}
fig,ax = plt.subplots()
ax.scatter(x = imputed_hackathon['11'], y = imputed_hackathon['16'])
plt.ylabel('Prediction')
plt.xlabel('11')
plt.show()
# + id="4CmP1ByR-cye" outputId="86c0d49c-8eb8-4391-b0b6-0e10613491ea" colab={"base_uri": "https://localhost:8080/", "height": 279}
#Column 12
fig,ax = plt.subplots()
ax.scatter(x = imputed_hackathon['12'], y = imputed_hackathon['16'])
plt.ylabel('Prediction')
plt.xlabel('12')
plt.show()
#no outliers
# + id="tOKPTjuQ-i7n" outputId="ffbc2d33-10e3-4ff5-c362-ada45734116f" colab={"base_uri": "https://localhost:8080/", "height": 279}
#Column 13
fig,ax = plt.subplots()
ax.scatter(x = imputed_hackathon['13'], y = imputed_hackathon['16'])
plt.ylabel('Prediction')
plt.xlabel('13')
plt.show()
# + id="xbvgeBrr-m6r" outputId="f3da3c95-f8bb-46f5-e5ea-e63f38d43442" colab={"base_uri": "https://localhost:8080/", "height": 111}
imputed_hackathon[imputed_hackathon['13']>50]
# + id="1WX_GeUX-pWQ" outputId="52bb93f1-4e2e-4c8a-9e42-c15c6eaca8b4" colab={"base_uri": "https://localhost:8080/"}
mean13 = np.mean(imputed_hackathon['13'])
mean13
# + id="cnq-QP4V-usT"
imputed_hackathon.iloc[2657, 12] = mean13
imputed_hackathon.iloc[3927, 12] = mean13
# + id="vL1h6sK9-w-6" outputId="569d4774-b878-4280-9fb5-50cab0f72ff1" colab={"base_uri": "https://localhost:8080/", "height": 279}
fig,ax = plt.subplots()
ax.scatter(x = imputed_hackathon['13'], y = imputed_hackathon['16'])
plt.ylabel('Prediction')
plt.xlabel('13')
plt.show()
# + id="qyr3Xq9m-zFl" outputId="17200e30-afa6-45d2-a190-e05783f429ea" colab={"base_uri": "https://localhost:8080/", "height": 279}
#Column 14
fig,ax = plt.subplots()
ax.scatter(x = imputed_hackathon['14'], y = imputed_hackathon['16'])
plt.ylabel('Prediction')
plt.xlabel('14')
plt.show()
# + id="h2CmLPhK-899" outputId="5b8cd4fa-5a67-4575-8b0d-54769b0b541a" colab={"base_uri": "https://localhost:8080/", "height": 111}
imputed_hackathon[imputed_hackathon['14']>130]
# + id="HUhWbwjs_EWZ"
mean14 = np.mean(imputed_hackathon['14'])
# + id="avfR4iPd-_k0"
imputed_hackathon.iloc[339,13] = mean14
imputed_hackathon.iloc[3142,13] = mean14
# + id="CdWtGypE_CBP" outputId="e6b8ae68-1a91-4916-b9ff-3dd3ac9ca9d9" colab={"base_uri": "https://localhost:8080/", "height": 279}
fig,ax = plt.subplots()
ax.scatter(x = imputed_hackathon['14'], y = imputed_hackathon['16'])
plt.ylabel('Prediction')
plt.xlabel('14')
plt.show()
# + id="P9LSfd3Q_KU2" outputId="a63bc14b-6ae4-448e-f445-d4c693ac76a8" colab={"base_uri": "https://localhost:8080/", "height": 279}
fig,ax = plt.subplots()
ax.scatter(x = imputed_hackathon['15'], y = imputed_hackathon['16'])
plt.ylabel('Prediction')
plt.xlabel('15')
plt.show()
# + id="kc6I7ftM_NN_" outputId="11e353df-0fbd-4992-fee2-810b84acea68" colab={"base_uri": "https://localhost:8080/", "height": 204}
imputed_hackathon[imputed_hackathon['15']>350]
# + id="egtc_saE_Pg5" outputId="431716ad-6e55-4baa-e1cf-f9abc61c2a8b" colab={"base_uri": "https://localhost:8080/"}
mean15 = np.mean(imputed_hackathon['15'])
mean15
# + id="HGgFBhMl_SLR"
imputed_hackathon.iloc[2406, 14] = mean15
imputed_hackathon.iloc[2893, 14] = mean15
imputed_hackathon.iloc[2909, 14] = mean15
imputed_hackathon.iloc[3844, 14] = mean15
imputed_hackathon.iloc[3971, 14] = mean15
# + id="PMc5S9bh_Uf1" outputId="5f997eb5-1c13-4980-d53f-f99ad3ed5a4d" colab={"base_uri": "https://localhost:8080/", "height": 279}
fig,ax = plt.subplots()
ax.scatter(x = imputed_hackathon['15'], y = imputed_hackathon['16'])
plt.ylabel('Prediction')
plt.xlabel('15')
plt.show()
# + [markdown] id="kKzUB3iQzKn4"
# ## **Skewness and Transformation(Normalization)**
# + id="n0qKwkx0_XNx" outputId="3db0f815-ecf5-4854-901d-a73cf2cd3be4" colab={"base_uri": "https://localhost:8080/"}
skewValue = imputed_hackathon.skew()
skewValue
# + id="MMbnY0qtzgyU"
#Skewness value is large for 6,7,9,11,15,16
#Only Columns with numerical data are considered for normalization
# + id="usefRDOM8ozS" outputId="1798a1f6-c9c0-4dd8-fbd9-d78dd1f782eb" colab={"base_uri": "https://localhost:8080/", "height": 400}
#Column 11
normalized_data11 = stats.boxcox(imputed_hackathon['11']) #applying boxcox transformation
fig, ax=plt.subplots(1,2)
sns.distplot(imputed_hackathon['11'], ax=ax[0]) #Original data visualization
ax[0].set_title("Original Data")
sns.distplot(normalized_data11[0] , ax=ax[1]) #After normalization visualization
ax[1].set_title("Normalized Data")
# + id="E3Etr_z58sFE" outputId="4e4b44b4-cec4-429c-b069-2e1ff25a64b2" colab={"base_uri": "https://localhost:8080/"}
new_11 = pd.Series(normalized_data11[0])
new_11
# + id="1H96AVtl83rt" outputId="9c7512c3-ee98-4b03-8e53-b3a5d1e6ed6f" colab={"base_uri": "https://localhost:8080/", "height": 419}
imputed_hackathon['11'] = new_11 #Replacing with transformed values
imputed_hackathon
# + id="lpMqEDOJ86H8" outputId="43ef1a87-2064-4440-ab45-79d4176a28a6" colab={"base_uri": "https://localhost:8080/", "height": 400}
#Column 15
normalized_data15 = stats.boxcox(imputed_hackathon['15']) #applying boxcox transformation
fig, ax=plt.subplots(1,2)
sns.distplot(imputed_hackathon['15'], ax=ax[0]) #Original data visualization
ax[0].set_title("Original Data")
sns.distplot(normalized_data15[0] , ax=ax[1])
ax[1].set_title("Normalized Data")
# + id="3apmK3av89jM" outputId="a5c37e16-b379-4d45-f797-dd1d3466cab0" colab={"base_uri": "https://localhost:8080/"}
new_15 = normalized_data15[0]
new_15
# + id="yfDn4IKu9Caj" outputId="83ec2099-70bc-45d7-cfe9-8ce25e9ac2a8" colab={"base_uri": "https://localhost:8080/", "height": 419}
imputed_hackathon['15'] = new_15
imputed_hackathon
# + id="jQhc2Yvm9EdW" outputId="ff40e633-ad2e-414c-dabc-61ae3b7e9971" colab={"base_uri": "https://localhost:8080/", "height": 400}
#Column 12
normalized_data12 = stats.boxcox(imputed_hackathon['12']) #applying boxcox transformation
fig, ax=plt.subplots(1,2)
sns.distplot(imputed_hackathon['12'], ax=ax[0]) #Original data visualization
ax[0].set_title("Original Data")
sns.distplot(normalized_data12[0] , ax=ax[1])
ax[1].set_title("Normalized Data")
# + id="nZsQGGsj9G2f" outputId="a2e49e94-f8df-4ff1-e619-497f3ac9e72b" colab={"base_uri": "https://localhost:8080/"}
new_12 = normalized_data12[0]
new_12
# + id="RIR9tg0D9JRK" outputId="ac3a5c39-019c-4cdc-8a5d-2b3b90eb0c56" colab={"base_uri": "https://localhost:8080/", "height": 419}
imputed_hackathon['12'] = new_12
imputed_hackathon
# + id="EPT6zvSq9Lns" outputId="1671a822-b7fe-4145-a9c6-502fbe4244b7" colab={"base_uri": "https://localhost:8080/", "height": 400}
#Column 13
normalized_data13 = stats.boxcox(imputed_hackathon['13']) #applying boxcox transformation
fig, ax=plt.subplots(1,2)
sns.distplot(imputed_hackathon['13'], ax=ax[0]) #Original data visualization
ax[0].set_title("Original Data")
sns.distplot(normalized_data13[0] , ax=ax[1])
ax[1].set_title("Normalized Data")
# + id="COMjejVr9OaO" outputId="9c455731-2b19-48d5-afa8-9ec7eddc5414" colab={"base_uri": "https://localhost:8080/"}
new_13 = normalized_data13[0]
new_13
# + id="uDYt596D9RpF" outputId="e22f5ade-c669-42c5-e470-ff6612be255e" colab={"base_uri": "https://localhost:8080/", "height": 419}
imputed_hackathon['13'] = new_13
imputed_hackathon
# + id="9RbHFm0c9UQh" outputId="0da439d8-7eaa-4d45-b1a5-c69f9ac4d981" colab={"base_uri": "https://localhost:8080/", "height": 400}
#Column 14
normalized_data14 = stats.boxcox(imputed_hackathon['14']) #applying boxcox transformation
fig, ax=plt.subplots(1,2)
sns.distplot(imputed_hackathon['14'], ax=ax[0]) #Original data visualization
ax[0].set_title("Original Data")
sns.distplot(normalized_data14[0] , ax=ax[1])
ax[1].set_title("Normalized Data")
# + id="3sOHweid9WlM"
new_14 = normalized_data14[0]
# + id="fnTzi_zu9Y3V" outputId="c796f255-4ce8-49a9-9a5a-3cd411614a21" colab={"base_uri": "https://localhost:8080/", "height": 419}
imputed_hackathon['14'] = new_14
imputed_hackathon
# + id="jPlm-3Q89bEN" outputId="1a97cfbb-702c-41c2-e9ff-2a1bf2055a0d" colab={"base_uri": "https://localhost:8080/", "height": 400}
#Column 15
normalized_data10 = stats.boxcox(imputed_hackathon['10']) #applying boxcox transformation
fig, ax=plt.subplots(1,2)
sns.distplot(imputed_hackathon['10'], ax=ax[0]) #Original data visualization
ax[0].set_title("Original Data")
sns.distplot(normalized_data10[0] , ax=ax[1])
ax[1].set_title("Normalized Data")
# + id="YqTN7eyF9dHa"
new_10 = normalized_data10[0]
# + id="LqqpceiQ9f1x" outputId="9b7a34c4-dc82-4dba-fe01-fbb168c0dc46" colab={"base_uri": "https://localhost:8080/", "height": 419}
imputed_hackathon['10'] = new_10
imputed_hackathon
# + id="Zv3B5Yfk9iAQ" outputId="9cbb4e88-04f7-449e-fb9d-36a5d937ef8d" colab={"base_uri": "https://localhost:8080/", "height": 400}
#Column 2
normalized_data2 = stats.boxcox(imputed_hackathon['2']) #applying boxcox transformation
fig, ax=plt.subplots(1,2)
sns.distplot(imputed_hackathon['2'], ax=ax[0]) #Original data visualization
ax[0].set_title("Original Data")
sns.distplot(normalized_data2[0] , ax=ax[1])
ax[1].set_title("Normalized Data")
# + id="PLLIld3T9kbw" outputId="e9957b3e-4c8f-4e1b-95b7-6bc7d177f7a7" colab={"base_uri": "https://localhost:8080/"}
new_2 = normalized_data2[0]
new_2
# + id="EBlbCBOx9nCk" outputId="7b7e3626-a17f-43a8-a519-9d5b080305fe" colab={"base_uri": "https://localhost:8080/", "height": 419}
imputed_hackathon['2'] = new_2
imputed_hackathon
# + id="rW1UmVQI9pEO" outputId="d4e45ab6-44f1-48de-c06f-b26c63ffb0fa" colab={"base_uri": "https://localhost:8080/"}
skewValue = imputed_hackathon.skew() #Checking for Skewness
skewValue
# + id="cizESAjj9tFQ"
normal_hackathon = imputed_hackathon #Checkpoint 2
# + [markdown] id="IT2tNPIZm9l8"
# ## **Checking Kurtosis**
# + id="oQ0zuvZkRlpA" outputId="b09de941-08f5-4e04-cfcb-c458af9ad34e" colab={"base_uri": "https://localhost:8080/"}
kur = kurtosis(normal_hackathon, fisher = True)
print(kur)
# + id="eFogU9l6SJgh"
#As Kurtosis values are in proper range, we do not do any more normalization
# + id="nW12DLWgVhgt"
# + [markdown] id="nHcuRDHUrUea"
# ## **Checking Correlation Matrix for Feature Selection**
# + id="GlxkyZzSq9TE" outputId="ff916dc9-f9a2-4e87-9da7-9727955d7435" colab={"base_uri": "https://localhost:8080/", "height": 545}
# Create correlation matrix
corr_matrix = normal_hackathon.corr().abs()
#Selecting upper traingular region
upper = corr_matrix.where(np.triu(np.ones(corr_matrix.shape), k=1).astype(np.bool))
upper
# + id="Dgg4MOc8sAQr"
#First we check for inter correlated columns. Those with high correlation were then compared with the correlation values of the final column.
#Those which had the minimum impact on final output were then dropped and evaluation was checked for different permutations and combinations.
# + [markdown] id="MrXrxLr0V-_i"
# ## **Scaling (StandardScaler)**
# + id="4lqXLYyhWEpW"
scaler = StandardScaler()
# + id="xyjiJNVbWXBc" outputId="79229980-8e9e-4082-fef8-6606055804da" colab={"base_uri": "https://localhost:8080/", "height": 419}
numerical = normal_hackathon[['2','10','11','12','13','14','15']]
categorical = normal_hackathon[[1,'3','4','5','6','7','8','9','16']]
# + id="ZqzGwZNeW7dY" outputId="48d3bd44-5b1c-49b9-d5f8-989cbff95748" colab={"base_uri": "https://localhost:8080/", "height": 419}
numerical_scaled = scaler.fit_transform(numerical)
numerical_scaled = pd.DataFrame(numerical_scaled)
numerical_scaled
# + id="UhD5F8hlXaoL" outputId="4c0eb3f4-771f-403e-f9e6-df1cfcadadcc" colab={"base_uri": "https://localhost:8080/", "height": 419}
final = pd.concat([numerical_scaled, categorical], ignore_index = True, axis = 1)
final
# + id="l7yqNAgpXk4d"
# + [markdown] id="c0Dwk0KEsLlY"
# ## **Encoding**
# + id="gE5DWbwlsAyn"
le = LabelEncoder()
#Column 3 has categorical data, 0,1,1.974,2,3
#We Encode this to 0,1,2,3,4
# + id="LoSzakAjsVxM" outputId="eee435d4-b001-40b1-a38c-da0e6aeceb28" colab={"base_uri": "https://localhost:8080/"}
normal_hackathon['3'] = le.fit_transform(normal_hackathon['3'])
normal_hackathon['3'].unique()
# + [markdown] id="I9Z6-23hsuCT"
# ## **Train Test Split**
# + id="RUktnemztErh"
X = final.drop(15, axis = 1)
y = final[15]
# + id="-Rr9IYMusYYR"
X_train, X_test, y_train, y_test = train_test_split(X,y,random_state = 42)
# + [markdown] id="ESKhkF3Utdyu"
# ## **Baseline Model - Dummy Classifier**
# + id="HuIZPyeytLs6"
dummy = DummyClassifier(strategy = 'most_frequent').fit(X_train, y_train)
# + id="ZExYg2xvuFuR" outputId="59838743-8d71-436e-adf1-2f117f012ff0" colab={"base_uri": "https://localhost:8080/"}
dummy_prediction = dummy.predict(X_test)
dummy_prediction
# + id="A0DPHP3OuIX6" outputId="6dd0d07a-1746-49e3-cfd8-7ad4630d0973" colab={"base_uri": "https://localhost:8080/"}
accuracy_score(y_test, dummy_prediction)
# + [markdown] id="vI_TRNezu_nW"
# # **Creating a Model Comparison Table**
# + id="M-5gfij1uLAM"
scoring = {'accuracy':make_scorer(accuracy_score),
'precision':make_scorer(precision_score),
'recall':make_scorer(recall_score),
'f1_score':make_scorer(f1_score)}
# + id="vbQInn1SvIcP"
log_model = LogisticRegression(max_iter=10000)
svc_model = LinearSVC(dual=False)
dtr_model = DecisionTreeClassifier()
rfc_model = RandomForestClassifier()
gnb_model = GaussianNB()
mlp_model = MLPClassifier()
# + id="GLUuKzkKvLPX"
def models_evaluation(X, y, folds):
'''
X : data set features
y : data set target
folds : number of cross-validation folds
'''
# Perform cross-validation to each machine learning classifier
log = cross_validate(log_model, X, y, cv=folds, scoring=scoring)
svc = cross_validate(svc_model, X, y, cv=folds, scoring=scoring)
dtr = cross_validate(dtr_model, X, y, cv=folds, scoring=scoring)
rfc = cross_validate(rfc_model, X, y, cv=folds, scoring=scoring)
gnb = cross_validate(gnb_model, X, y, cv=folds, scoring=scoring)
mlp = cross_validate(gnb_model, X, y, cv=folds, scoring=scoring)
# Create a data frame with the models perfoamnce metrics scores
models_scores_table = pd.DataFrame({'Logistic Regression':[log['test_accuracy'].mean(),
log['test_precision'].mean(),
log['test_recall'].mean(),
log['test_f1_score'].mean()],
'Support Vector Classifier':[svc['test_accuracy'].mean(),
svc['test_precision'].mean(),
svc['test_recall'].mean(),
svc['test_f1_score'].mean()],
'Decision Tree':[dtr['test_accuracy'].mean(),
dtr['test_precision'].mean(),
dtr['test_recall'].mean(),
dtr['test_f1_score'].mean()],
'Random Forest':[rfc['test_accuracy'].mean(),
rfc['test_precision'].mean(),
rfc['test_recall'].mean(),
rfc['test_f1_score'].mean()],
'Gaussian Naive Bayes':[gnb['test_accuracy'].mean(),
gnb['test_precision'].mean(),
gnb['test_recall'].mean(),
gnb['test_f1_score'].mean()],
'Multi Layer Perceptron':[mlp['test_accuracy'].mean(),
mlp['test_precision'].mean(),
mlp['test_recall'].mean(),
mlp['test_f1_score'].mean()]},
index=['Accuracy', 'Precision', 'Recall', 'F1 Score'])
# Add 'Best Score' column
models_scores_table['Best Score'] = models_scores_table.idxmax(axis=1)
# Return models performance metrics scores data frame
return(models_scores_table)
# + id="xynOArwrvgGw" outputId="5ce3706d-5606-4574-b2fe-63f3245eacb9" colab={"base_uri": "https://localhost:8080/", "height": 173}
models_evaluation(X_test, y_test , 5)
# + id="aWlxUhtbvk9M"
#F1 scores are very low
#Here, we see that accuracy is almost same as the Dummy Classfier Accuracy.
#This means that Data Imbalance exists
# + id="VLHJtm_uwJvu"
rfc_model.fit(X_train, y_train) #Creating a sample RF model for Imbalance Check
pred = rfc_model.predict(X_test)
# + id="h7HBLVzbvxP7" outputId="4cbdd2b7-c8a8-461a-e8c0-bf33326b54ac" colab={"base_uri": "https://localhost:8080/"}
confusion = confusion_matrix(y_test, pred )
confusion #Here, we see that imbalance exists
#Thus, Data Resampling is required
# + [markdown] id="_mrBkMmoxGaR"
# ## **Handling Imbalance**
# + id="8xqSmAOqwlIc"
sm = SMOTE(sampling_strategy = 'auto')
# + id="4ORtMqMSxOgT" outputId="a41eb0e8-7386-48f8-f7e2-35369d84f763" colab={"base_uri": "https://localhost:8080/"}
#Using Oversampling strategy
X_res, y_res = sm.fit_resample(X,y)
X_res = pd.DataFrame(X_res)
y_res = pd.DataFrame(y_res)
y_res.columns = ['15']
# + id="00F6t9VcxR7U" outputId="ebabf2b5-dbe6-4753-8ee1-f64623379a6e" colab={"base_uri": "https://localhost:8080/", "height": 419}
res_hackathon = pd.concat([X_res, y_res], axis = 1)
res_hackathon
# + id="tOWhAUjwxblj"
X_train, X_test, y_train, y_test = train_test_split(X_res,y_res,random_state = 42) #Train test split for resampled
# + id="XNhJetNCzyd1" outputId="2df23506-4d32-4e25-ec77-eb6023a2bafa" colab={"base_uri": "https://localhost:8080/", "height": 1000}
#Checking accuracies in model comparison table
models_evaluation(X_test, y_test , 5)
# + id="kEzN_19tCgyM"
dummy = DummyClassifier(strategy = 'most_frequent').fit(X_train, y_train)
# + id="Twc-8VFhHDxP" outputId="3e435cb6-ad3b-4cda-9d79-80c03359471f" colab={"base_uri": "https://localhost:8080/"}
dummy_prediction = dummy.predict(X_test)
dummy.score(X_test, y_test) #Reduced Dummy Classifier accuracy indicates that less imbalance Exists
# + [markdown] id="0uHGJbqW0Bch"
# **We see that accuracy in the model comparison table has significantly dropped after resampling. This means that, accuracy is not the correct evalutation metric.
# On the other hand, f1 score has shown an improvement.
# So, we choose f1_score as our evaluation metric.**
# + id="kiBZMyeRAYdO" outputId="c9894065-1c2e-4af5-b6b6-293dbf1bc59e" colab={"base_uri": "https://localhost:8080/"}
#Random Forests Classifier gives the best F1 Score (0.7652)
#Trying out more Algorithms for better F1_score
#XGBoost
xgb = XGBClassifier(objective= 'binary:logistic', n_estimators=70, seed=101).fit(X_train, y_train)
# + id="VAVd47fE0oCt"
xgpred = xgb.predict(X_test)
# + id="vy532lXBJTkn" outputId="1467edef-4b01-4ff2-9e88-ae659e84cf8d" colab={"base_uri": "https://localhost:8080/"}
f1_score(xgpred, y_test) #Better accuracy than RandomForestClassifier
# + [markdown] id="1Moy2H7hKY_x"
# ## **Hyperparameter Tuning to improve the model**
# + id="zPqOzuh-JdI2"
params = {
'learning_rate' :[0.05, 0.10, 0.15, 0.20, 0.25, 0.30],
'max_depth' :[3,4,5,6,8,10,12,15],
'min_child_weight' :[1,3,5,7],
'gamma' :[0.0, 0.1, 0.2, 0.3, 0.4],
'colsample_bytree' :[0.3,0.4,0.5,0.7]
}
# + id="Fsc-Mf0OKBGt"
random_search = RandomizedSearchCV(xgb, param_distributions=params, n_iter = 5, scoring = 'roc_auc', n_jobs = -1, cv=5,verbose = 3)
# + id="ay11qjzvKn_-" outputId="e90ff940-d78b-434a-bb97-77f85f098a67" colab={"base_uri": "https://localhost:8080/"}
random_search.fit(X_res, y_res)
# + id="aokq_0vpKq_R" outputId="d1daaa31-d8a6-4964-8746-caa10c163d48" colab={"base_uri": "https://localhost:8080/"}
random_search.best_estimator_ #gives the best estimator values for xgboost
# + id="ySjsCk__KwQm"
xgb = XGBClassifier(base_score=0.5, booster='gbtree', colsample_bylevel=1,
colsample_bynode=1, colsample_bytree=0.7, gamma=0.1,
learning_rate=0.2, max_delta_step=0, max_depth=15,
min_child_weight=1, missing=None, n_estimators=70, n_jobs=1,
nthread=None, objective='binary:logistic', random_state=0,
reg_alpha=0, reg_lambda=1, scale_pos_weight=1, seed=101,
silent=None, subsample=1, verbosity=1)
# + id="o31lg0bBLDI1" outputId="2549ce52-ce04-4b03-d277-b3bcc33ed7b4" colab={"base_uri": "https://localhost:8080/"}
xgb.fit(X_train, y_train)
# + id="5GfoarT8LGKC" outputId="2a69cb68-0e54-49fc-eb67-28f81407547a" colab={"base_uri": "https://localhost:8080/"}
predicted = xgb.predict(X_test)
predicted
# + id="p6KKk3mnLTUJ" outputId="06a3df24-c47e-406a-cc73-f65617fa611a" colab={"base_uri": "https://localhost:8080/"}
f1_score(predicted, y_test) #F1 score increased by 10% !!!
#This is probably our FINAL SCORE !!!
# + [markdown] id="9ieHBxMad5EI"
# ## **Creating Pipelines for different methods to compare with different normalization and Scaling Techniques**
# + id="W6_6lV7-Lbk4"
pipeline_lr = Pipeline([('scaler1', StandardScaler()),
('pca1',PCA(n_components = 2)),
('lr_classifier',LogisticRegression(random_state = 0))])
# + id="IUof8AoTednc"
pipeline_dt = Pipeline([('scaler2', MinMaxScaler()),
('lda2', LinearDiscriminantAnalysis(n_components = 1)),
('dt_classifier', DecisionTreeClassifier())])
# + id="un6sKnojeoyd"
pipeline_rf = Pipeline([('scaler3', StandardScaler()),
('pca2', PCA(n_components = 2)),
('rf_classifier', RandomForestClassifier())])
# + id="g6FgAlM0eq_d"
pipeline_svm = Pipeline([('scaler4', StandardScaler()),
('lda2', LinearDiscriminantAnalysis(n_components = 1)),
('svm_classifier', SVC())])
# + id="YKAAZS1BetG4"
pipelines = [pipeline_lr, pipeline_dt, pipeline_rf, pipeline_svm]
# + id="LEl8ihh8fDIx" outputId="f424225b-a1ed-4b08-f87e-ddfbf0fd7aaa" colab={"base_uri": "https://localhost:8080/"}
pipe_dict = {0: 'Logistic Regression', 1: 'Decision Tree', 2:'Random Classifier', 3: 'Support Vector Classifier', 4: 'XGB Classifier'}
for pipe in pipelines:
pipe.fit(X_train, y_train)
# + id="7ass_pBefEDN" outputId="a9dc684d-4fe4-4745-fef7-9dd37bcc7b14" colab={"base_uri": "https://localhost:8080/"}
for i, model in enumerate(pipelines):
pred = model.predict(X_test)
print('{} F1 scores: {}'.format(pipe_dict[i], f1_score(pred, y_test)))
# + id="bd2rnmgkfRp4"
#Thus, we see that all these F1 scores are less than XGBoost Classifier used above.
#Thus XGBoost remains our final model.
# + id="KmvBm6yhfn-j"
#Now, we drop rows, which have been determined by using various permutations upon abserving the correlation matrix, to improve the score.
# + id="2JW31T1vf1GF" outputId="f155ba11-7235-4836-8858-2562b4614c99" colab={"base_uri": "https://localhost:8080/", "height": 419}
X_res = X_res.drop([2,13],axis = 1)
X_res
# + id="4lZNoguMf62y"
Xd_train, Xd_test, yd_train, yd_test = train_test_split(X_res, y_res, random_state =42)
# + id="3GkCXuI2gBlZ" outputId="6136245c-34fd-41ab-ac90-527e593d5859" colab={"base_uri": "https://localhost:8080/"}
xgb.fit(Xd_train, yd_train)
# + id="MgvdBKiggH8x"
predict = xgb.predict(Xd_test)
# + id="euzjdfUkgL59" outputId="f500e939-b460-43d2-e3d9-da34ae861865" colab={"base_uri": "https://localhost:8080/"}
f1_score(predict, yd_test)
# + id="MmjeSPTGgOyL"
#After trying out feature selection using correlation matrix, we find out that the score does not increase.
#So, our final score using XGBClassifier is 90.07 %.
# + [markdown] id="DJp-ZMxmalhr"
# ## **Final Score = 90.07%**
# + [markdown] id="FKnlcjQepvE8"
# ### **Neural Network**
# + id="ePWmszCIoPHR"
import pandas as pd
import numpy as np
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
# + id="RxkDEJzooP9E" outputId="ac96c2bc-c64e-45ac-d417-3530451f91e6" colab={"base_uri": "https://localhost:8080/"}
from google.colab import drive
drive.mount('/content/drive')
# + id="najlsXRVoQAM" outputId="4f0b4d64-5046-40d7-afd0-6d0cb5f0f61d" colab={"base_uri": "https://localhost:8080/", "height": 304}
hack = pd.read_csv("/content/drive/My Drive/ML projects/reshackathon.csv")
hack.describe()
# + id="IizRxc3loQDZ" outputId="338228c6-b9e2-48d1-a035-a27fca12dc30" colab={"base_uri": "https://localhost:8080/"}
dataset = hack.values
x = dataset[:,1:16]
x
# + id="E33vOSK4oQGO" outputId="91593119-0a46-444f-f4d6-ded72ace03ed" colab={"base_uri": "https://localhost:8080/"}
y = dataset[:,16]
y
# + id="qROlxThloQI_" outputId="206bb7d0-1877-4bc4-c7d7-522c821d190b" colab={"base_uri": "https://localhost:8080/"}
min_max_scaler = preprocessing.MinMaxScaler()
X_scale = min_max_scaler.fit_transform(x)
X_scale
# + id="id4y_4BToQLw" outputId="beedcc09-2142-4a6e-c386-e49f418e6e1f" colab={"base_uri": "https://localhost:8080/"}
X_train, X_val_and_test, Y_train, Y_val_and_test = train_test_split(X_scale, y, test_size=0.3,random_state = 42)
X_val, X_test, Y_val, Y_test = train_test_split(X_val_and_test, Y_val_and_test, test_size=0.5,random_state=42)
print(X_train.shape, X_val.shape, X_test.shape, Y_train.shape, Y_val.shape, Y_test.shape)
# + id="1T3gRXgcoQOT"
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Dropout
from keras import regularizers
from keras.layers import BatchNormalization
# + id="12xS8UncsQ9z"
from keras.callbacks import EarlyStopping
early_stopping = EarlyStopping(monitor='val_loss',
patience=8,
min_delta=0.001,
mode='min')
# + id="7ydgz3TnoQRQ"
model_3 = Sequential()
model_3.add(Dense(64, activation='relu',kernel_regularizer=regularizers.l2(0.01), input_shape=(15,)))
model_3.add(BatchNormalization())
model_3.add(Dropout(0.3))
model_3.add(Dense(64, activation='relu',kernel_regularizer=regularizers.l2(0.01)))
model_3.add(BatchNormalization())
#model_3.add(Dropout(0.3))
model_3.add(Dense(64, activation='relu',kernel_regularizer=regularizers.l2(0.01)))
model_3.add(BatchNormalization())
model_3.add(Dense(1, activation='sigmoid'))
# + id="kFZUlDnUoQUO"
model_3.compile(optimizer='adam',loss='binary_crossentropy',metrics=['BinaryAccuracy'])
# + id="6BVrGtJboQcI" outputId="1223e767-c072-4f17-b771-0cf7fb52ab18" colab={"base_uri": "https://localhost:8080/"}
model_3.summary()
# + id="llQ3IERbuDGu" outputId="446b7821-7893-45e2-8d00-e8b85c339dc9" colab={"base_uri": "https://localhost:8080/"}
hist = model_3.fit(X_train, Y_train,batch_size=64, epochs=150,validation_data=(X_val, Y_val))
# + id="laNWAAz0uDY-" outputId="78da6908-de16-4752-e12d-4798150a2a23" colab={"base_uri": "https://localhost:8080/", "height": 295}
plt.plot(hist.history['loss'])
plt.plot(hist.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Val'], loc='upper right')
plt.show()
# + id="k9WRqFFNuDyh" outputId="e2e3994f-717b-4940-8c6b-5bd932c6f0b5" colab={"base_uri": "https://localhost:8080/", "height": 295}
plt.plot(hist.history['binary_accuracy'])
plt.plot(hist.history['val_binary_accuracy'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Val'], loc='lower right')
plt.show()
# + id="P2wsWtNWuEFy" outputId="140b0eeb-d20e-4ae4-b9e9-5226cd663b35" colab={"base_uri": "https://localhost:8080/"}
model_3.evaluate(X_test, Y_test)[1]
#Deep learning gives a lower accuracy
# + [markdown] id="44PNXE--zxEx"
# ## **Final Score = 90.07**
# + id="68U4Bmv2uEL7"
# + id="gwbFu-z1uEO-"
| Hackathon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import os
import numpy as np
import pandas as pd
import seaborn as sns
df = pd.read_csv('heart.csv')
df.head()
# -
# ## Distrubution Plot for Numeric Columns
sns.displot(df['Age'], kde = True, color = 'blue')
sns.displot(df['RestingBP'], kde = True, color = 'blue')
sns.displot(df['Cholesterol'], kde = True, color = 'blue')
sns.displot(df['MaxHR'], kde = True, color = 'blue')
# ## Pie Charts for Categorical
df.groupby('Sex').size().plot(kind = 'pie', autopct = '%.1f')
df.groupby('ChestPainType').size().plot(kind = 'pie', autopct = '%.1f')
df.groupby('RestingECG').size().plot(kind = 'pie', autopct = '%.1f')
df.groupby('ST_Slope').size().plot(kind = 'pie', autopct = '%.1f')
df.groupby('HeartDisease').size().plot(kind = 'pie', autopct = '%.1f')
# ## ViolinPlot
sns.violinplot(df['Age'])
sns.violinplot(y = df['Sex'], x = df['HeartDisease'])
sns.violinplot(y = df['Age'], x = df['HeartDisease'])
sns.violinplot(y = df['Cholesterol'], x = df['HeartDisease'])
# # Correlation - Heatmap
df.corr()
sns.heatmap(df.corr())
# # JointPlot
sns.jointplot(x = 'Age' , y = 'MaxHR', data = df, kind = 'hex')
sns.jointplot(x = 'Age' , y = 'MaxHR', data = df, kind = 'reg')
sns.jointplot(x = 'Cholesterol' , y = 'MaxHR', data = df, kind = 'reg')
df.head()
sns.jointplot(x = 'HeartDisease' , y = 'MaxHR', data = df, kind = 'reg')
# # Pairplot
sns.pairplot(df)
| Heart Disease EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
from tensorflow.python.framework import ops
ops.reset_default_graph()
sess = tf.Session()
x_vals = np.random.normal(1, 0.1, 100)
y_vals = np.repeat(10., 100)
x_data = tf.placeholder(shape=[1], dtype=tf.float32)
y_target = tf.placeholder(shape=[1], dtype=tf.float32)
A = tf.Variable(tf.random_normal(shape=[1]))
my_output = tf.multiply(x_data, A)
loss = tf.square(my_output - y_target)
init = tf.global_variables_initializer()
sess.run(init)
my_opt = tf.train.GradientDescentOptimizer(0.02)
train_step = my_opt.minimize(loss)
for i in range(100):
rand_index = np.random.choice(100)
rand_x = [x_vals[rand_index]]
rand_y = [y_vals[rand_index]]
sess.run(train_step, feed_dict={x_data: rand_x, y_target: rand_y})
if (i+1)%25==0:
print('Step #' + str(i+1) + ' A = ' + str(sess.run(A)))
print('Loss = ' + str(sess.run(loss, feed_dict={x_data: rand_x, y_target: rand_y})))
ops.reset_default_graph()
sess = tf.Session()
x_vals = np.concatenate((np.random.normal(-1, 1, 50), np.random.normal(3, 1, 50)))
y_vals = np.concatenate((np.repeat(0., 50), np.repeat(1., 50)))
x_data = tf.placeholder(shape=[1], dtype=tf.float32)
y_target = tf.placeholder(shape=[1], dtype=tf.float32)
A = tf.Variable(tf.random_normal(mean=10, shape=[1]))
my_output = tf.add(x_data, A)
my_output_expanded = tf.expand_dims(my_output, 0)
y_target_expanded = tf.expand_dims(y_target, 0)
xentropy = tf.nn.sigmoid_cross_entropy_with_logits(logits=my_output_expanded,labels=y_target_expanded)
my_opt = tf.train.GradientDescentOptimizer(0.05)
train_step = my_opt.minimize(xentropy)
init = tf.global_variables_initializer()
sess.run(init)
for i in range(1400):
rand_index = np.random.choice(100)
rand_x = [x_vals[rand_index]]
rand_y = [y_vals[rand_index]]
sess.run(train_step, feed_dict={x_data: rand_x, y_target: rand_y})
if (i+1)%200==0:
print('Step #' + str(i+1) + ' A = ' + str(sess.run(A)))
print('Loss = ' + str(sess.run(xentropy, feed_dict={x_data: rand_x, y_target: rand_y})))
# +
predictions = []
for i in range(len(x_vals)):
x_val = [x_vals[i]]
prediction = sess.run(tf.round(tf.sigmoid(my_output)), feed_dict={x_data: x_val})
predictions.append(prediction[0])
accuracy = sum(x==y for x,y in zip(predictions, y_vals))/100.
print('Ending Accuracy = ' + str(np.round(accuracy, 2)))
# -
| Section 2/Implementing Back Propagation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python388jvsc74a57bd0e4a0114c34cb418e7d5a3697731935c08a51df47520424022bf18e448f4ec53a
# ---
# + [markdown] id="vN99YjPTDena" colab_type="text"
# # Polynomial Regression
# + [markdown] id="ZIx_naXnDyHd" colab_type="text"
# ## Importing the libraries
# + id="FjnmdyPLD2tS" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# + [markdown] id="6c8YExmOD5x5" colab_type="text"
# ## Importing the dataset
# + id="nQOdXhjXD_AE" colab_type="code" colab={}
dataset = pd.read_csv('Position_Salaries.csv')
X = dataset.iloc[:, 1:-1].values
y = dataset.iloc[:, -1].values
# + [markdown] id="Le8SEL-YEOLb" colab_type="text"
# ## Training the Linear Regression model on the whole dataset
# + id="2eZ4xxbKEcBk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="41074f6d-44c7-4a04-fd49-14bda9fb2885" executionInfo={"status": "ok", "timestamp": 1587538036471, "user_tz": -240, "elapsed": 1084, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhEuXdT7eQweUmRPW8_laJuPggSK6hfvpl5a6WBaA=s64", "userId": "15047218817161520419"}}
from sklearn.linear_model import LinearRegression
lin_reg = LinearRegression()
lin_reg.fit(X, y)
# + [markdown] id="Rb5nWuSHEfBV" colab_type="text"
# ## Training the Polynomial Regression model on the whole dataset
# + id="HYplp4pTEm0O" colab_type="code" outputId="4c3c03dd-0def-4584-a893-aa2e72629e8f" executionInfo={"status": "ok", "timestamp": 1587538038634, "user_tz": -240, "elapsed": 693, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhEuXdT7eQweUmRPW8_laJuPggSK6hfvpl5a6WBaA=s64", "userId": "15047218817161520419"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
from sklearn.preprocessing import PolynomialFeatures
poly_reg = PolynomialFeatures(degree = 6) # If coefficients are constant, the regression is linear
# degree is the highest degree(needs to be predicted)
X_poly = poly_reg.fit_transform(X)
lin_reg_2 = LinearRegression()
lin_reg_2.fit(X_poly, y)
# + [markdown] id="0O8R0tzbEpvy" colab_type="text"
# ## Visualising the Linear Regression results
# + id="dcTIBAEdEyve" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="c242f259-d9e6-442a-f026-79dffab85972" executionInfo={"status": "ok", "timestamp": 1587538041090, "user_tz": -240, "elapsed": 753, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhEuXdT7eQweUmRPW8_laJuPggSK6hfvpl5a6WBaA=s64", "userId": "15047218817161520419"}}
plt.scatter(X, y, color = 'red')
plt.plot(X, lin_reg.predict(X), color = 'blue')
plt.title('Truth or Bluff (Linear Regression)')
plt.xlabel('Position Level')
plt.ylabel('Salary')
plt.show()
# + [markdown] id="stOnSo74E52m" colab_type="text"
# ## Visualising the Polynomial Regression results
# + id="UCOcurIQE7Zv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="93927499-de98-4a31-a619-c373926cbe56" executionInfo={"status": "ok", "timestamp": 1587538044032, "user_tz": -240, "elapsed": 827, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhEuXdT7eQweUmRPW8_laJuPggSK6hfvpl5a6WBaA=s64", "userId": "15047218817161520419"}}
plt.scatter(X, y, color = 'red')
plt.plot(X, lin_reg_2.predict(poly_reg.fit_transform(X)), color = 'blue')
plt.title('Truth or Bluff (Polynomial Regression)')
plt.xlabel('Position level')
plt.ylabel('Salary')
plt.show()
# + [markdown] id="U_qsAMKnE-PJ" colab_type="text"
# ## Visualising the Polynomial Regression results (for higher resolution and smoother curve)
# + id="iE6EnC3fFClE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="6ecb5687-3c8a-4b46-db4a-c4955c24b9de" executionInfo={"status": "ok", "timestamp": 1587538047267, "user_tz": -240, "elapsed": 1035, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhEuXdT7eQweUmRPW8_laJuPggSK6hfvpl5a6WBaA=s64", "userId": "15047218817161520419"}}
X_grid = np.arange(min(X), max(X), 0.01)
X_grid = X_grid.reshape((len(X_grid), 1))
plt.scatter(X, y, color = 'red')
plt.plot(X_grid, lin_reg_2.predict(poly_reg.fit_transform(X_grid)), color = 'blue')
plt.title('Truth or Bluff (Polynomial Regression)')
plt.xlabel('Position level')
plt.ylabel('Salary')
plt.show()
# + [markdown] id="diyJFZHhFFeK" colab_type="text"
# ## Predicting a new result with Linear Regression
# + id="Blmp6Hn7FJW6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f01610bc-b077-4df0-cae4-ea37c8b0037f" executionInfo={"status": "ok", "timestamp": 1587538054192, "user_tz": -240, "elapsed": 872, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhEuXdT7eQweUmRPW8_laJuPggSK6hfvpl5a6WBaA=s64", "userId": "15047218817161520419"}}
lin_reg.predict([[6.5]]) # 2D array
# + [markdown] id="DW7I7ZVDFNkk" colab_type="text"
# ## Predicting a new result with Polynomial Regression
# + id="uQmtnyTHFRGG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="2739bf8a-6dfb-4226-b200-252ee8857097" executionInfo={"status": "ok", "timestamp": 1587538056906, "user_tz": -240, "elapsed": 774, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhEuXdT7eQweUmRPW8_laJuPggSK6hfvpl5a6WBaA=s64", "userId": "15047218817161520419"}}
lin_reg_2.predict(poly_reg.fit_transform([[6.5]])) # needs to fit transform for poly reg model
| Python Tutorial Machine Learning/Part 2 - Regression/polynomial_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Kaggle Titanic survival - logistic regression model
#
# In this notebook we repeat our basic logistic regression model as previously described:
#
# https://github.com/MichaelAllen1966/1804_python_healthcare/blob/master/titanic/02_logistic_regression.ipynb
#
# We will extend the model to report a range of accuracy measures, as described:
#
# https://github.com/MichaelAllen1966/1804_python_healthcare/blob/master/titanic/05_accuracy_standalone.ipynb
# We will go through the following steps:
#
# * Download and save pre-processed data
# * Split data into features (X) and label (y)
# * Split data into training and test sets (we will test on data that has not been used to fit the model)
# * Standardise data
# * Fit a logistic regression model (from sklearn learn)
# * Predict survival of the test set
# * Define a function to calculate a range of accuracy measure (and return as a dictionary)
# * Report multipmultipleel accuracy scores for model
# ## Load modules
#
# A standard Anaconda install of Python (https://www.anaconda.com/distribution/) contains all the necessary modules.
import numpy as np
import pandas as pd
# Import machine learning methods
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
# ## Load data
#
# The section below downloads pre-processed data, and saves it to a subfolder (from where this code is run).
# If data has already been downloaded that cell may be skipped.
#
# Code that was used to pre-process the data ready for machine learning may be found at:
# https://github.com/MichaelAllen1966/1804_python_healthcare/blob/master/titanic/01_preprocessing.ipynb
# +
download_required = True
if download_required:
# Download processed data:
address = 'https://raw.githubusercontent.com/MichaelAllen1966/' + \
'1804_python_healthcare/master/titanic/data/processed_data.csv'
data = pd.read_csv(address)
# Create a data subfolder if one does not already exist
import os
data_directory ='./data/'
if not os.path.exists(data_directory):
os.makedirs(data_directory)
# Save data
data.to_csv(data_directory + 'processed_data.csv', index=False)
# -
data = pd.read_csv('data/processed_data.csv')
# Make all data 'float' type
data = data.astype(float)
# The first column is a passenger index number. We will remove this, as this is not part of the original Titanic passenger data.
# +
# Drop Passengerid (axis=1 indicates we are removing a column rather than a row)
# We drop passenger ID as it is not original data
data.drop('PassengerId', inplace=True, axis=1)
# -
# ## Divide into X (features) and y (labels)
#
# We will separate out our features (the data we use to make a prediction) from our label (what we are truing to predict).
# By convention our features are called `X` (usually upper case to denote multiple features), and the label (survive or not) `y`.
X = data.drop('Survived',axis=1) # X = all 'data' except the 'survived' column
y = data['Survived'] # y = 'survived' column from 'data'
# ## Divide into training and tets sets
#
# When we test a machine learning model we should always test it on data that has not been used to train the model.
# We will use sklearn's `train_test_split` method to randomly split the data: 75% for training, and 25% for testing.
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.25)
# ## Standardise data
#
# We want all of out features to be on roughly the same scale. This generally leads to a better model, and also allows us to more easily compare the importance of different features.
#
# One simple method is to scale all features 0-1 (by subtracting the minimum value for each value, and dividing by the new remaining maximum value).
#
# But a more common method used in many machine learning methods is standardisation, where we use the mean and standard deviation of the training set of data to normalise the data. We subtract the mean of the test set values, and divide by the standard deviation of the training data. Note that the mean and standard deviation of the training data are used to standardise the test set data as well.
#
# Here we will use sklearn's `StandardScaler method`. This method also copes with problems we might otherwise have (such as if one feature has zero standard deviation in the training set).
def standardise_data(X_train, X_test):
# Initialise a new scaling object for normalising input data
sc = StandardScaler()
# Set up the scaler just on the training set
sc.fit(X_train)
# Apply the scaler to the training and test sets
train_std=sc.transform(X_train)
test_std=sc.transform(X_test)
return train_std, test_std
X_train_std, X_test_std = standardise_data(X_train, X_test)
# ## Fit logistic regression model
#
# Now we will fir a logistic regression model, using sklearn's `LogisticRegression` method. Our machine learning model fitting is only two lines of code! By using the name `model` for our logistic regression model we will make our model more interchangeable later on.
model = LogisticRegression(solver='lbfgs')
model.fit(X_train_std,y_train)
# ## Predict values
#
# Now we can use the trained model to predict survival. We will test the accuracy of both the training and test data sets.
# Predict training and test set labels
y_pred_train = model.predict(X_train_std)
y_pred_test = model.predict(X_test_std)
# ## Calculate accuracy
#
# Here we define a function that will calculate a range of accuracy scores.
def calculate_accuracy(observed, predicted):
"""
Calculates a range of accuracy scores from observed and predicted classes.
Takes two list or NumPy arrays (observed class values, and predicted class
values), and returns a dictionary of results.
1) observed positive rate: proportion of observed cases that are +ve
2) Predicted positive rate: proportion of predicted cases that are +ve
3) observed negative rate: proportion of observed cases that are -ve
4) Predicted negative rate: proportion of predicted cases that are -ve
5) accuracy: proportion of predicted results that are correct
6) precision: proportion of predicted +ve that are correct
7) recall: proportion of true +ve correctly identified
8) f1: harmonic mean of precision and recall
9) sensitivity: Same as recall
10) specificity: Proportion of true -ve identified:
11) positive likelihood: increased probability of true +ve if test +ve
12) negative likelihood: reduced probability of true +ve if test -ve
13) false positive rate: proportion of false +ves in true -ve patients
14) false negative rate: proportion of false -ves in true +ve patients
15) true positive rate: Same as recall
16) true negative rate
17) positive predictive value: chance of true +ve if test +ve
18) negative predictive value: chance of true -ve if test -ve
"""
# Converts list to NumPy arrays
if type(observed) == list:
observed = np.array(observed)
if type(predicted) == list:
predicted = np.array(predicted)
# Calculate accuracy scores
observed_positives = observed == 1
observed_negatives = observed == 0
predicted_positives = predicted == 1
predicted_negatives = predicted == 0
true_positives = (predicted_positives == 1) & (observed_positives == 1)
false_positives = (predicted_positives == 1) & (observed_positives == 0)
true_negatives = (predicted_negatives == 1) & (observed_negatives == 1)
accuracy = np.mean(predicted == observed)
precision = (np.sum(true_positives) /
(np.sum(true_positives) + np.sum(false_positives)))
recall = np.sum(true_positives) / np.sum(observed_positives)
sensitivity = recall
f1 = 2 * ((precision * recall) / (precision + recall))
specificity = np.sum(true_negatives) / np.sum(observed_negatives)
positive_likelihood = sensitivity / (1 - specificity)
negative_likelihood = (1 - sensitivity) / specificity
false_positive_rate = 1 - specificity
false_negative_rate = 1 - sensitivity
true_positive_rate = sensitivity
true_negative_rate = specificity
positive_predictive_value = (np.sum(true_positives) /
np.sum(observed_positives))
negative_predictive_value = (np.sum(true_negatives) /
np.sum(observed_negatives))
# Create dictionary for results, and add results
results = dict()
results['observed_positive_rate'] = np.mean(observed_positives)
results['observed_negative_rate'] = np.mean(observed_negatives)
results['predicted_positive_rate'] = np.mean(predicted_positives)
results['predicted_negative_rate'] = np.mean(predicted_negatives)
results['accuracy'] = accuracy
results['precision'] = precision
results['recall'] = recall
results['f1'] = f1
results['sensitivity'] = sensitivity
results['specificity'] = specificity
results['positive_likelihood'] = positive_likelihood
results['negative_likelihood'] = negative_likelihood
results['false_positive_rate'] = false_positive_rate
results['false_negative_rate'] = false_negative_rate
results['true_positive_rate'] = true_positive_rate
results['true_negative_rate'] = true_negative_rate
results['positive_predictive_value'] = positive_predictive_value
results['negative_predictive_value'] = negative_predictive_value
return results
# +
# Call calculate_accuracy function
accuracy = calculate_accuracy(y_test, y_pred_test)
# Print results up to three decimal places
for key, value in accuracy.items():
print (key, "{0:0.3}".format(value))
# -
# We can see from the accuracy scores that overall accuracy is about 80%, but that accuracy is imbalanced between survivors and non-survivors. We can see the model is biased towards predicting fewer survivors than actually occurred, and this gives higher specificity (the proportion of non-survivors correctly identified) than sensitivity (the proportion of survivors correctly identified). In the next notebook we will look at adjusting the balance between sensitivity and specificity.
#
# Note: To keep this example simple we have used a single random split between training and test data. A more thorough analysis would use repeated measurement using stratified k-fold validation (see https://github.com/MichaelAllen1966/1804_python_healthcare/blob/master/titanic/03_k_fold.ipynb).
| jupyter_notebooks/05a_accuracy_logistic_regression.ipynb |