code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.0
# language: julia
# name: julia-1.1
# ---
# # MeshCat Animations
#
# MeshCat.jl also provides an animation interface, built on top of the [three.js animation system](https://threejs.org/docs/#manual/introduction/Animation-system). While it is possible to construct animation clips and tracks manually, just as you would in Three.js, it's generally easier to use the MeshCat `Animation` type.
#
# Let's show off building a simple animation. We first have to create our scene:
# +
# Optional:
# These commands tell the Julia package manager to use the exact
# set of dependencies specified in the Project.toml and
# Manifest.toml files in this folder. That should
# give you a nice, reproducible environment for testing.
using Pkg
Pkg.activate(@__DIR__)
Pkg.instantiate()
# -
using MeshCat, GeometryTypes, CoordinateTransformations
vis = Visualizer()
# +
## To open the visualizer in a new browser tab, do:
# open(vis)
## To open the visualizer inside this jupyter notebook, do:
# IJuliaCell(vis)
## To open this visualizer in a standalone window, do:
# using Blink
# AtomShell.isinstalled() || AtomShell.install()
# open(vis, Window())
# -
setobject!(vis[:box1],
HyperRectangle(Vec(0., 0, 0), Vec(0.1, 0.2, 0.3)))
# ### Building an Animation
#
# We construct an animation by first creating a blank `Animation()` object. We can then use the `atframe` function to set properties or transforms of the animation at specific frames of the animation. Three.js will automatically interpolate between whatever values we provide.
#
# For example, let's animate moving the box from [0, 0, 0] to [0, 1, 0]:
# +
anim = Animation()
atframe(anim, 0) do
# within the context of atframe, calls to
# `settransform!` and `setprop!` are intercepted
# and recorded in `anim` instead of having any
# effect on `vis`.
settransform!(vis[:box1], Translation(0., 0, 0))
end
atframe(anim, 30) do
settransform!(vis[:box1], Translation(0., 1, 0))
end
# `setanimation!()` actually sends the animation to the
# viewer. By default, the viewer will play the animation
# right away. To avoid that, you can also pass `play=false`.
setanimation!(vis, anim)
# -
# You should see the box slide 1 meter to the right in the viewer. If you missed the animation, you can run it again from the viewer. Click "Open Controls", find the "Animations" section, and click "play".
# ### Animating the Camera
#
# The camera is just another object in the MeshCat scene. To set its transform, we just need to index into the visualizer with the right path (note the leading `/`):
settransform!(vis["/Cameras/default"], Translation(0, 0, 1))
# To animate the camera, we just have to do that same kind of `settransform!` to individual frames in an animation:
# +
anim = Animation()
atframe(anim, 0) do
settransform!(vis["/Cameras/default"], Translation(0., 0, 0))
end
atframe(anim, 30) do
settransform!(vis["/Cameras/default"], Translation(0., 0, 1))
end
setanimation!(vis, anim)
# -
# We can also animate object properties. For example, let's animate the camera's `zoom` property to smoothly zoom out and then back in. Note that to do this, we have to access a deeper path in the visualizer to get to the actual camera object. For more information, see: https://github.com/rdeits/meshcat#camera-control
# +
anim = Animation()
atframe(anim, 0) do
setprop!(vis["/Cameras/default/rotated/<object>"], "zoom", 1)
end
atframe(anim, 30) do
setprop!(vis["/Cameras/default/rotated/<object>"], "zoom", 0.5)
end
atframe(anim, 60) do
setprop!(vis["/Cameras/default/rotated/<object>"], "zoom", 1)
end
setanimation!(vis, anim)
# -
# ### Recording an Animation
#
# To record an animation at a smooth, fixed frame rate, click on "Open Controls" in the viewer, and then go to "Animations" -> "default" -> "Recording" -> "record". This will play the entire animation, recording every frame and then let you download the resulting frames to your computer.
#
# To record activity in the MeshCat window that isn't a MeshCat animation, we suggest using a screen-capture tool like Quicktime for macOS or RecordMyDesktop for Linux.
# ### Converting the Animation into a Video
#
# Currently, meshcat can only save an animation as a `.tar` file consisting of a list of `.png` images, one for each frame. To convert that into a video, you will need to install the `ffmpeg` program, and then you can run:
# +
# MeshCat.convert_frames_to_video("/home/rdeits/Downloads/meshcat_1528401494656.tar", overwrite=true)
# -
| notebooks/animation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Normalize data
p = [2, 4, 10, 6, 8, 4]
pmin, pmax = min(p), max(p)
delta = pmax - pmin
for i, _ in enumerate(p):
p[i] = (p[i] - pmin) / delta
print(p)
# ## FizzBuzz
nmax = 100
for n in range(1, nmax + 1):
message = ''
if not n % 3:
message = 'Fizz'
if not n % 5:
message += 'Buzz'
print(message or n)
# ## Hamming distance
# +
s = 'hello'
t = 'hallu'
d = 0
for a, b in zip(s, t):
if a != b:
d += 1
print("hamming(", s, ", ", t, ") = ", d, sep='')
# -
# ## 2D Arrays
n = 3
m = 3
a = []
val = 0
for i in range(n):
row = []
for j in range(m):
row.append(val)
val += 1
a.append(row)
print(*a, sep='\n')
# ## The Luhn formula
# +
ccn = '4799 2739 8713 6272'
ccn = ''.join(ccn.split())[::-1]
checksum = 0
for index, digit in enumerate(ccn):
d = int(digit)
if index % 2:
d *= 2
a, b = d//10, d%10
checksum += a + b
print(not checksum%10)
# -
# ## Redwood data
# +
f = open('data/redwood.txt')
# read header information
f.readline()
f.readline()
max_h = 0
max_d = 0
info_h = ''
info_d = ''
for line in f:
data = line.split()
val = float(data[-1])
if max_h < val:
max_h = val
info_h = " ".join(data)
val = float(data[-2])
if max_d < val:
max_d = val
info_d = " ".join(data)
print("maximum height : ", max_h)
print("info about tallest tree :", info_h)
print("maximum diameter : ", max_d)
print("info about widest tree :", info_d)
# -
| ue7/ue7_project/notebooks/ue07.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import yahoo_fin.stock_info as si
import pandas as pd
import numpy as np
import statsmodels.tsa.stattools as ts
import statsmodels.api as sm
list_banks = ['ZION','SBNY','NYCB','WAL','BKU','CMA','EWBC','BAC','C','AXP','COF']
def get_price_data(ticker):
price_data = si.get_data(ticker,start_date = '01/01/2009')
df = pd.DataFrame(price_data)
df = df[['adjclose']]
df['pct_change'] = df.adjclose.pct_change()
df['log_return'] = np.log(1 + df['pct_change'].astype(float))
df['ticker'] = ticker
return df
def get_portfolio_return(df,list_stocks,n):
df = df.iloc[-n:,].copy()
for col in df.columns:
if col in list_stocks:
df[col+'_cmltv_ret'] = np.exp(np.log1p(df[col]).cumsum())-1
list_cols = []
for col in df.columns:
if 'cmltv' in col and 'SPY' not in col:
list_cols.append(col)
df['portfolio_return'] = df[list_cols].mean(axis=1)
return df
def get_betas(x,y,n=0):
if n > 0:
x = x.iloc[-n:,]
y = y.iloc[-n:,]
res = sm.OLS(y, x).fit()
ticker = col.split('_')[0]
beta = res.params[0]
r2 = res.rsquared
n = len(x)
return [beta,r2,n]
list_df = []
for item in list_banks:
df = get_price_data(item)
list_df.append(df)
# +
df_long = pd.concat(list_df,axis = 0).reset_index()
df_wide = df_long.pivot_table(index=["index"],
columns='ticker',
values='pct_change')
spy = get_price_data('TLT').reset_index()
dataset = pd.merge(spy,df_wide,how = 'inner',left_on='index',right_on='index')
betas = {}
ns = {}
r2s = {}
for col in dataset.columns:
if col in list_banks:
df = dataset[['pct_change',col]].dropna()
beta,r2,n = get_betas(df['pct_change'],df[col])
betas[col]=beta
ns[col]=n
r2s[col]=r2
# -
# PRINT LIST OF BETAS VS the TLT... INVERSE BETAS = GREATER ASSET SENSITIVITY
# LIST SORTED FROM MOST ASSET SENSITIVE TO LEAST
{k: v for k, v in sorted(betas.items(), key=lambda item: item[1])}
ns
dataset_stocks = dataset.drop(columns = ['adjclose','pct_change','log_return','ticker'])
portfolio_return = get_portfolio_return(dataset_stocks,list_banks,n=252)
spy_return = get_portfolio_return(dataset[['pct_change']],'pct_change',n=252)
spy_return = spy_return.drop(columns = ['pct_change','pct_change_cmltv_ret'])
spy_return.columns = ['SPY_return']
df_plot = pd.concat([portfolio_return,spy_return],axis = 1)
df_plot.set_index('index')[['portfolio_return','SPY_return']].plot(title = '252 day cumulative return')
| banks_book_eda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Character-level Language Modeling with LSTMs
#
# This notebook is adapted from [Keras' lstm_text_generation.py](https://github.com/fchollet/keras/blob/master/examples/lstm_text_generation.py).
#
# Steps:
#
# - Download a small text corpus and preprocess it.
# - Extract a character vocabulary and use it to vectorize the text.
# - Train an LSTM-based character level langague model.
# - Use the trained model to sample random text with varying entropy levels.
# - Implement a beam-search deterministic decoder.
#
#
# **Note**: fitting language models is compute intensive. It is recommended to do this notebook on a server with a GPU or powerful CPUs that you can leave running for several hours at once.
# + deletable=true editable=true
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# + [markdown] deletable=true editable=true
# ## Loading some text data
#
# Let's use some publicly available philosopy:
# + deletable=true editable=true
from keras.utils.data_utils import get_file
URL = "https://s3.amazonaws.com/text-datasets/nietzsche.txt"
corpus_path = get_file('nietzsche.txt', origin=URL)
text = open(corpus_path).read().lower()
print('Corpus length: %d characters' % len(text))
# + deletable=true editable=true
print(text[:600], "...")
# + deletable=true editable=true
text = text.replace("\n", " ")
split = int(0.9 * len(text))
train_text = text[:split]
test_text = text[split:]
# + [markdown] deletable=true editable=true
# ## Building a vocabulary of all possible symbols
#
# To simplifly things, we build a vocabulary by extracting the list all possible characters from the full datasets (train and validation).
#
# In a more realistic setting we would need to take into account that the test data can hold symbols never seen in the training set. This issue is limited when we work at the character level though.
#
# Let's build the list of all possible characters and sort it to assign a unique integer to each possible symbol in the corpus:
# + deletable=true editable=true
chars = sorted(list(set(text)))
print('total chars:', len(chars))
char_indices = dict((c, i) for i, c in enumerate(chars))
indices_char = dict((i, c) for i, c in enumerate(chars))
# + [markdown] deletable=true editable=true
# `char_indices` is a mapping to from characters to integer identifiers:
# + deletable=true editable=true
len(char_indices)
# + deletable=true editable=true
sorted(char_indices.items())[:15]
# + [markdown] deletable=true editable=true
# `indices_char` holds the reverse mapping:
# + deletable=true editable=true
len(indices_char)
# + deletable=true editable=true
indices_char[52]
# + [markdown] deletable=true editable=true
# While not strictly required to build a language model, it's a good idea to have a look a the distribution of relative frequencies of each symbol in the corpus:
# + deletable=true editable=true
from collections import Counter
counter = Counter(text)
chars, counts = zip(*counter.most_common())
indices = np.arange(len(counts))
plt.figure(figsize=(14, 3))
plt.bar(indices, counts, 0.8)
plt.xticks(indices, chars);
# + [markdown] deletable=true editable=true
# Let's cut the dataset into fake sentences at random with some overlap. Instead of cutting at random we could use a English specific sentence tokenizer. This is explained at the end of this notebook. In the mean time random substring will be good enough to train a first language model.
# + deletable=true editable=true
max_length = 40
step = 3
def make_sequences(text, max_length=max_length, step=step):
sequences = []
next_chars = []
for i in range(0, len(text) - max_length, step):
sequences.append(text[i: i + max_length])
next_chars.append(text[i + max_length])
return sequences, next_chars
sequences, next_chars = make_sequences(train_text)
sequences_test, next_chars_test = make_sequences(test_text, step=10)
print('nb train sequences:', len(sequences))
print('nb test sequences:', len(sequences_test))
# + [markdown] deletable=true editable=true
# Let's shuffle the sequences to break some of the dependencies:
# + deletable=true editable=true
from sklearn.utils import shuffle
sequences, next_chars = shuffle(sequences, next_chars,
random_state=42)
# + deletable=true editable=true
sequences[0]
# + deletable=true editable=true
next_chars[0]
# + [markdown] deletable=true editable=true
# ## Converting the training data to one-hot vectors
#
# Unfortunately the LSTM implementation in Keras does not (yet?) accept integer indices to slice columns from an input embedding by it-self. Let's use one-hot encoding. This is slightly less space and time efficient than integer coding but should be good enough when using a small character level vocabulary.
#
# **Exercise:**
#
# One hot encoded the training `data sequences` as `X` and `next_chars` as `y`:
# + deletable=true editable=true
n_sequences = len(sequences)
n_sequences_test = len(sequences_test)
voc_size = len(chars)
X = np.zeros((n_sequences, max_length, voc_size),
dtype=np.float32)
y = np.zeros((n_sequences, voc_size), dtype=np.float32)
X_test = np.zeros((n_sequences_test, max_length, voc_size),
dtype=np.float32)
y_test = np.zeros((n_sequences_test, voc_size), dtype=np.float32)
# TODO
# + deletable=true editable=true
# # %load solutions/language_model_one_hot_data.py
n_sequences = len(sequences)
n_sequences_test = len(sequences_test)
voc_size = len(chars)
X = np.zeros((n_sequences, max_length, voc_size),
dtype=np.float32)
y = np.zeros((n_sequences, voc_size), dtype=np.float32)
X_test = np.zeros((n_sequences_test, max_length, voc_size),
dtype=np.float32)
y_test = np.zeros((n_sequences_test, voc_size), dtype=np.float32)
for i, sequence in enumerate(sequences):
for t, char in enumerate(sequence):
X[i, t, char_indices[char]] = 1
y[i, char_indices[next_chars[i]]] = 1
for i, sequence in enumerate(sequences_test):
for t, char in enumerate(sequence):
X_test[i, t, char_indices[char]] = 1
y_test[i, char_indices[next_chars_test[i]]] = 1
# + deletable=true editable=true
X.shape
# + deletable=true editable=true
y.shape
# + deletable=true editable=true
X[0]
# + deletable=true editable=true
y[0]
# + [markdown] deletable=true editable=true
# ## Measuring per-character perplexity
#
# The NLP community measures the quality of probabilistic model using [perplexity](https://en.wikipedia.org/wiki/Perplexity).
#
# In practice perplexity is just a base 2 exponentiation of the average negative log2 likelihoods:
#
# $$perplexity_\theta = 2^{-\frac{1}{n} \sum_{i=1}^{n} log_2 (p_\theta(x_i))}$$
#
# **Note**: here we define the **per-character perplexity** (because our model naturally makes per-character predictions). **It is more common to report per-word perplexity**. Note that this is not as easy to compute the per-world perplexity as we would need to tokenize the strings into a sequence of words and discard whitespace and punctuation character predictions. In practice the whitespace character is the most frequent character by far making our naive per-character perplexity lower than it sould be if we ignored those.
#
# **Exercise**: implement a Python function that computes the per-character perplexity with model predicted probabilities `y_pred` and `y_true` for the encoded ground truth:
# + deletable=true editable=true
def perplexity(y_true, y_pred):
"""Compute the per-character perplexity of model predictions.
y_true is one-hot encoded ground truth.
y_pred is predicted likelihoods for each class.
2 ** -mean(log2(p))
"""
# TODO
return 1.
# + deletable=true editable=true
# # %load solutions/language_model_perplexity.py
def perplexity(y_true, y_pred):
"""Compute the perplexity of model predictions.
y_true is one-hot encoded ground truth.
y_pred is predicted likelihoods for each class.
2 ** -mean(log2(p))
"""
likelihoods = np.sum(y_pred * y_true, axis=1)
return 2 ** -np.mean(np.log2(likelihoods))
# + deletable=true editable=true
y_true = np.array([
[0, 1, 0],
[0, 0, 1],
[0, 0, 1],
])
y_pred = np.array([
[0.1, 0.9, 0.0],
[0.1, 0.1, 0.8],
[0.1, 0.2, 0.7],
])
perplexity(y_true, y_pred)
# + [markdown] deletable=true editable=true
# A perfect model has a minimal perplixity of 1.0 (negative log likelihood of 0.0):
# + deletable=true editable=true
perplexity(y_true, y_true)
# + [markdown] deletable=true editable=true
# ## Building recurrent model
#
# Let's build a first model and train it on a very small subset of the data to check that it works as expected:
# + deletable=true editable=true
from keras.models import Sequential
from keras.layers import LSTM, Dense
from keras.optimizers import RMSprop
model = Sequential()
model.add(LSTM(128, input_shape=(max_length, voc_size)))
model.add(Dense(voc_size, activation='softmax'))
optimizer = RMSprop(lr=0.01)
model.compile(optimizer=optimizer, loss='categorical_crossentropy')
# + [markdown] deletable=true editable=true
# Let's measure the perplexity of the randomly initialized model:
# + deletable=true editable=true
def model_perplexity(model, X, y, verbose=0):
predictions = model.predict(X, verbose=verbose)
return perplexity(y, predictions)
# + deletable=true editable=true
model_perplexity(model, X_test, y_test)
# + [markdown] deletable=true editable=true
# Let's train the model for one epoch on a very small subset of the training set to check that it's well defined:
# + deletable=true editable=true
small_train = slice(0, None, 40)
model.fit(X[small_train], y[small_train], validation_split=0.1,
batch_size=128, nb_epoch=1)
# + deletable=true editable=true
model_perplexity(model, X[small_train], y[small_train])
# + deletable=true editable=true
model_perplexity(model, X_test, y_test)
# + [markdown] deletable=true editable=true
# ## Sampling random text from the model
#
# Recursively generate one character at a time by sampling from the distribution parameterized by the model:
#
# $$
# p_{\theta}(c_n | c_{n-1}, c_{n-2}, \ldots, c_0) \cdot p_{\theta}(c_{n-1} | c_{n-2}, \ldots, c_0) \cdot \ldots \cdot p_{\theta}(c_{0})
# $$
#
# The temperature parameter makes it possible to remove additional entropy (bias) into the parmeterized multinoulli distribution of the output of the model:
# + deletable=true editable=true
def sample_one(preds, temperature=1.0):
"""Sample the next character according to the network output.
Use a lower temperature to force the model to output more
confident predictions: more peaky distribution.
"""
preds = np.asarray(preds).astype('float64')
preds = np.log(preds) / temperature
exp_preds = np.exp(preds)
preds = exp_preds / np.sum(exp_preds)
# Draw a single sample (size=1) from a multinoulli distribution
# parameterized by the output of the softmax layer of our
# network. A multinoulli distribution is a multinomial
# distribution with a single trial with n_classes outcomes.
probs = np.random.multinomial(1, preds, size=1)
return np.argmax(probs)
def generate_text(model, seed_string, length=300, temperature=1.0):
"""Recursively sample a sequence of chars, one char at a time.
Each prediction is concatenated to the past string of predicted
chars so as to condition the next prediction.
Feed seed string as a sequence of characters to condition the
first predictions recursively. If seed_string is lower than
max_length, pad the input with zeros at the beginning of the
conditioning string.
"""
generated = seed_string
prefix = seed_string
for i in range(length):
# Vectorize prefix string to feed as input to the model:
x = np.zeros((1, max_length, voc_size))
shift = max_length - len(prefix)
for t, char in enumerate(prefix):
x[0, t + shift, char_indices[char]] = 1.
preds = model.predict(x, verbose=0)[0]
next_index = sample_one(preds, temperature)
next_char = indices_char[next_index]
generated += next_char
prefix = prefix[1:] + next_char
return generated
# + deletable=true editable=true
generate_text(model, 'philosophers are ', temperature=0.1)
# + deletable=true editable=true
generate_text(model, 'atheism is the root of ', temperature=0.8)
# + [markdown] deletable=true editable=true
# ## Training the model
#
# Let's train the model and monitor the perplexity after each epoch and sample some text to qualitatively evaluate the model:
# + deletable=true editable=true
nb_epoch = 30
seed_strings = [
'philosophers are ',
'atheism is the root of ',
]
for epoch in range(nb_epoch):
print("# Epoch %d/%d" % (epoch + 1, nb_epoch))
print("Training on one epoch takes ~90s on a K80 GPU")
model.fit(X, y, validation_split=0.1, batch_size=128, nb_epoch=1,
verbose=2)
print("Computing perplexity on the test set:")
test_perplexity = model_perplexity(model, X_test, y_test)
print("Perplexity: %0.3f\n" % test_perplexity)
for temperature in [0.1, 0.5, 1]:
print("Sampling text from model at %0.2f:\n" % temperature)
for seed_string in seed_strings:
print(generate_text(model, seed_string, temperature=temperature))
print()
# + [markdown] deletable=true editable=true
# ## Beam search for deterministic decoding
#
# **Exercise**: adapt the sampling decoder to implement a deterministic decoder with a beam of k=30 sequences that are the most likely sequences based on the model predictions.
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# ## Better handling of sentence boundaries
#
# To simplify things we used the lower case version of the text and we ignored any sentence boundaries. This prevents our model to learn when to stop generating characters. If we want to train a model that can start generating text at the beginning of a sentence and stop at the end of a sentence, we need to provide it with sentency boundary markers in the training set and use those special markers when sampling.
#
# The following give an example of how to use NLTK to detect sentence boundaries in English text.
#
# This could be used to insert an explicit "end_of_sentence" (EOS) symbol to mark separation between two consecutive sentences. This should make it possible to train a language model that explicitly generates complete sentences from start to end.
#
# Use the following command (in a terminal) to install nltk before importing it in the notebook:
#
# ```
# $ pip install nltk
# ```
# + deletable=true editable=true
text_with_case = open(corpus_path).read().replace("\n", " ")
# + deletable=true editable=true
import nltk
nltk.download('punkt')
from nltk.tokenize import sent_tokenize
sentences = sent_tokenize(text_with_case)
# + deletable=true editable=true
plt.hist([len(s.split()) for s in sentences], bins=30);
plt.title('Distribution of sentence lengths')
plt.xlabel('Approximate number of words');
# + [markdown] deletable=true editable=true
# The first few sentences detected by NLTK are too short to be considered real sentences. Let's have a look at short sentences with at least 20 characters:
# + deletable=true editable=true
sorted_sentences = sorted([s for s in sentences if len(s) > 20], key=len)
for s in sorted_sentences[:5]:
print(s)
# + [markdown] deletable=true editable=true
# Some long sentences:
# + deletable=true editable=true
for s in sorted_sentences[-3:]:
print(s)
# + [markdown] deletable=true editable=true
# The NLTK sentence tokenizer seems to do a reasonable job despite the weird casing and '--' signs scattered around the text.
#
# Note that here we use the original case information because it can help the NLTK sentence boundary detection model make better split decisions. Our text corpus is probably too small to train a good sentence aware language model though, especially with full case information. Using larger corpora such as a large collection of [public domain books](http://www.gutenberg.org/) or Wikipedia dumps. The NLTK toolkit also comes from [corpus loading utilities](http://www.nltk.org/book/ch02.html).
#
# The following loads a selection of famous books from the Gutenberg project archive:
# + deletable=true editable=true
import nltk
nltk.download('gutenberg')
book_selection_text = nltk.corpus.gutenberg.raw().replace("\n", " ")
# + deletable=true editable=true
print(book_selection_text[:300])
# + deletable=true editable=true
print("Book corpus length: %d characters" % len(book_selection_text))
# + [markdown] deletable=true editable=true
# Let's do an arbitrary split. Note the training set will have a majority of text that is not authored by the author(s) of the validation set:
# + deletable=true editable=true
split = int(0.9 * len(book_selection_text))
book_selection_train = book_selection_text[:split]
book_selection_validation = book_selection_text[split:]
# + [markdown] deletable=true editable=true
# ## Bonus exercises
#
# - Adapt the previous language model to handle explicitly sentence boundaries with a special EOS character.
# - Train a new model on the random sentences sampled from the the book selection corpus with full case information.
# - Adapt the random sampling code to start sampling at the beginning of sentence and stop when the sentence ends.
# - Train a deep GRU (e.g. two GRU layers instead of one LSTM) to see if you can improve the validation perplexity.
# - Git clone the source code of the [Linux kernel](https://github.com/torvalds/linux) and train a C programming language model on it. Instead of sentence boundary markers, we could use source file boundary markers for this exercise.
# - Try to increase the vocabulary size to 256 using a [Byte Pair Encoding](https://arxiv.org/abs/1508.07909) strategy.
# + deletable=true editable=true
| lectures-labs/labs/06_deep_nlp/Character_Level_Language_Model_rendered.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.2 64-bit (''in1910-book'': conda)'
# name: python3
# ---
# # Week 3
# ## Writing reliable and understandable code
#
# The goal of this weeks exercises is to give you some exercise with testing and documentation.
# ### Exercise 1) Checking primes
#
# Write a function `is_prime(n)` that takes in a positive integer and returns a boolean saying if the input is a prime or not. The function should raise an error if the input is invalid.
#
# Add a descriptive docstring to your function. Now write a few unit tests for your function using `pytest`. Your tests should test a few different numbers, especially 1, 2, and 3. Also write a test to verify that your function raises exceptions as expected (use `pytest.raises`, read more [here](https://docs.pytest.org/en/latest/assert.html) under 'Assertions about expected exceptions').
#
#
#
#
# ### Exercise 2) Writing Bubble Sort
#
# You will now write a function implementing the "bubble sort" algorithm, which is one of the simplest sorting algorithms. Assuming we have a list with elements we can compare using a "less than" operator. We want to sort the list in increasing order, so for example:
# * `[4, 2, 3, 7, 1, 5] -> [1, 2, 3, 4, 5, 7]`
#
# The steps of bubble sort are as follows:
#
# 1. Start with the first element in the list, compare it with the second.
# * If they are in the wrong order, exchange them.
# 2. Now compare the second and third element
# * If they are in the wrong order, exchange them.
# 3. Proceed in the same manner, comparing the third and fourth, then the fourth and fifth, etc
#
# At this point, after comparing the second to last and the last elements and flipping if necessary, the largest element of the list will be at the end of the list (Why is this?).
#
# Now, start again from the beginning, but stop once you reach the second-last element in the list. Then start again from the beginning, but stop once you reach the third-last element and so on, untill the whole list is sorted.
#
# If you need a better description, you can read more about bubble sort on [wikipedia](https://en.wikipedia.org/wiki/Bubble_sort) where they also have pseudocode examples.
#
# **Write a `bubble_sort` function that takes in a tuple or list and returns a new list with the same data in sorted order. The function should *not* change the original list. Include a docstring with your function**
# ### Exercise 3) Testing Bubble Sort
#
# You should now write a few unit tests of your bubble sort function. Include at least these tests:
# - An empty list should return an empty list.
# - A list with just one element should return a list with just that element.
# - Try sorting a few example inputs and check the output is sorted.
# - Verify that the input list is unchanged.
#
# If any tests fail, go back and improve your bubble sort.
# ### Exercise 4) Testing and correcting a median function
#
# The following function finds the median of a given dataset:
# +
def median(data):
"""Returns the median of a dataset."""
data.sort()
return data[len(data)//2]
print(median([11, 3, 1, 5, 3]))
# -
# The function returns the correct answer for the example given, but is the function any good? To find out, implement the following tests and run them. If any of the tests fails, improve the function so that it passes:
# - A test checking the output for a one-element list
# - A test checking the output for a two-element list
# - A test ensuring the original data is unchanged
# - A test checking that an error is raised if the input is an empty list/tuple.
| book/docs/exercises/week3/E3_testing_and_docstrings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Programmieren mit IfcOpenShell: Einführung
#
# Wie in jedes andere Pythonmodul, muss IfcOpenshell zunächst import werden um die Funktionalitäten zur Verfügung zu haben
import ifcopenshell
# Dann kann eines der verfügbaren Modelle geladen werden. Im `data` stehen eine paar Testmodelle bereit.
m = ifcopenshell.open("./data/hello_reiff_2021.ifc")
# Mit der in diesem Notebook verfügbaren Helfer-Klasssen `JupyterIFCRenderer` kann das geladene Modell eingebettet im Notebook angezeigt werden.
from utils.JupyterIFCRenderer import JupyterIFCRenderer
viewer = JupyterIFCRenderer(m, size=(400,300))
viewer
# ## Auswahl von Objekten
# Durch klicken mit der Maus auf ein Objekt mit dem im Viewer-Fenster können wir ein Objekt auswählen. Rechts vom 3D-Fenster erscheinen entsprechende Informationen zur Art des Objecktes und seinen Attributen. Das momentan selektierte Objekt können wir einer Variablen zuweisen.
#
auswahl = viewer.getSelectedProduct()
auswahl
# ## Attribute
#
# Die ausgegebene Information ist genau die Zeile, mit der das entsprechende Objekt in der `.ifc` Datei (im sog. "STEP Physical File Format" nach der ISO 10303 part 21) definiert wurde.
#
# IfcOpenshell bietet verschiedene Möglichkeiten, um Informationen des Objektes anzuzeigen. `get_info()` liefert ein dictionary aller attribute, und ihrer Werte für das entsprechenden Objekt.
auswahl.get_info()
# Auf jedes Attribut kann auf verschieden Arten im Skript zugegriffen werden:
# Alle Attribute können in der Reihenfolge ihrer Schema-Definition geladen werden: Das 0te Element ist bei allen von `IfcRoot` Elementen immer die GlobalId, also eine üseudo-zufällig erzeugte Zeichenkette mit einem einzigartigen Identifikator. Das dritte Element (indexzummer 2) ist der `Name` (wenn der vergeben wurde).
auswahl[0]
auswahl[2]
# Wenn wir sicher sind, um welche Instanz es sich handelt, können wir auch per .Attrubutnamen - Notation direkt auf das den Wert des entsprechenen Parameters zugreifen.
auswahl.GlobalId
# ## Verbindungen und Beziehungen
#
# 
#
# 
## Schema und Intropsektion
import ifcopenshell.util
import ifcopenshell.util.element
print(ifcopenshell.util.element.get_psets(auswahl))
help(m.wrapped_data.entity_names())
import ifcopenshell
schema = ifcopenshell.ifcopenshell_wrapper.schema_by_name("IFC4")
wall_decl = schema.declaration_by_name("IfcWall")
dir(ifcopenshell.ifcopenshell_wrapper)
# Mit der Funktion `m.wrapped_data.types()` können wir uns alle im Modell verwendeten Datenypen anzeigen lassen:
m.wrapped_data.types()
viewer.setDefaultColors()
#
viewer.getSelectedProduct()
viewer._bb.xmax
dir(viewer._renderer)
settings = ifcopenshell.geom.settings()
settings.set(settings.USE_PYTHON_OPENCASCADE, True)
for wall in m.by_type("IfcDoor"):
pdtc_shape = ifcopenshell.geom.create_shape(settings, wall)
print(pdtc_shape.styles[0])
for p in m.by_type("IfcProduct"):
viewer.setColorProduct(p, "#BBBBBB")
from IPython.display import display, HTML
js = "<script>alert('Hello World!');</script>"
viewer.html.value+=js
viewer.setDefaultColors()
tree_settings = ifcopenshell.geom.settings()
tree_settings.set(tree_settings.DISABLE_OPENING_SUBTRACTIONS, True)
t = ifcopenshell.geom.tree(m, tree_settings)
# +
wall = m.by_type("IfcWall")[0]
print("Intersecting with wall 2O2Fr$t4X7Zf8NOew3FLPP")
print(t.select(wall))
# -
sel = t.select(wall)
type(sel[0])
list(viewer._meshdict.values())[0].material.metallness=0.0
list(viewer._meshdict.values())[0].material.
# + active=""
# viewer.DisplayShapeAsSVG(viewer.elementdict.get(m.by_type("IfcWall")[5]))
# +
list(viewer._meshdict.values())[0].material.__dict__
# -
viewer.layout
help(viewer.layout)
| 00_hello_wall.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Import the codes
import sys
sys.path.insert(0, '/Users/ageller/WORK/LSST/onGitHub/EBLSST/code')
from EBLSST import BreivikGalaxy, EclipsingBinary
# ### Set up Katie's model
# +
g = BreivikGalaxy()
#define the correct paths to the input files and db
g.GalaxyFile ='/Users/ageller/WORK/LSST/onGitHub/EBLSST/input/Breivik/dat_ThinDisk_12_0_12_0.h5' #for Katie's model
g.GalaxyFileLogPrefix ='/Users/ageller/WORK/LSST/onGitHub/EBLSST/input/Breivik/fixedPopLogCm_'
g.setKernel()
# -
# ### Draw binaries from the model\
#
# *And add on the Teff values*
# +
nSample = 1 #number of binaries to draw
BreivikBin = g.GxSample(nSample)
EB = EclipsingBinary()
BreivikBin['Teff1'] = EB.getTeff(BreivikBin['L1'].values[0], BreivikBin['r1'].values[0])
BreivikBin['Teff2'] = EB.getTeff(BreivikBin['L2'].values[0], BreivikBin['r2'].values[0])
print(BreivikBin)
# -
# *Below you can use this to define the binaries for the light curves*
#
# For the light curve:
# * r_1 = the stellar radius_1 / semi-major axis (careful about units)
# * surface brightness is proportional to luminosity / radius$^2$
# * the mass ratio, q = m2/m1
| testing/limbDarkening/testLDC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Blockchain data
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import requests
import json
# ## Bitcoin blockchain, what's in a block?
#
# APIs are available to retrieve the blockchain data like [blockchain.com](https://www.blockchain.com/explorer?utm_campaign=expnav_explorer)
r = requests.get('https://blockchain.info/rawblock/1')
block = dict(r.json())
block
# ["tx"]
# ["hash"]
# ## Bitcoin price index
#
# We are going to download the bitcoin exchange rate information to USD and make some plots. Let us start with the bitcoin prices. The data is retrieved from the website [blockchain.com](https://www.blockchain.com/charts)
# +
import datetime
with open('../Data/market-price.json') as f:
exchange_rate = json.load(f)
exchange_rate_data = dict(exchange_rate)
exchange_rate_df = pd.DataFrame.from_records(exchange_rate_data["values"])
exchange_rate_df['Date'] = np.array([datetime.datetime(1970, 1, 1) + datetime.timedelta(seconds=int(exchange_rate_df["x"].iloc[k])) for k in range(len(exchange_rate_df["x"]))])
exchange_rate_df
fig, ax = plt.subplots(figsize=(6, 4))
ax.plot(exchange_rate_df['Date'], exchange_rate_df['y'], color=(0,0.4,0.5))
ax.set_xlabel('Date')
ax.set_title('Bitcoin price (in USD)')
sns.despine()
plt.savefig("../Figures/btc_price.pdf")
# -
# ## Bitcoin Network Hashrate
# +
with open('../Data/hash-rate.json') as f:
hash_rate = json.load(f)
hash_rate_data = dict(hash_rate)
hash_rate_df = pd.DataFrame.from_records(hash_rate_data["values"])
hash_rate_df['Date'] = np.array([datetime.datetime(1970, 1, 1) + datetime.timedelta(seconds=int(hash_rate_df["x"].iloc[k])) for k in range(len(hash_rate_df["x"]))])
fig, ax = plt.subplots(figsize=(6, 4))
ax.plot(hash_rate_df['Date'], hash_rate_df['y'] / 1e6, color=(0,0.4,0.5))
ax.set_xlabel('Date')
ax.set_title('Daily Network Hashrate (in millions of TH per second)')
sns.despine()
plt.savefig("../Figures/btc_hahsrate.pdf")
# -
# ## Number of transactions
# +
with open('../Data/n-transactions.json') as f:
n_transaction = json.load(f)
n_transaction_data = dict(n_transaction)
n_transaction_df = pd.DataFrame.from_records(n_transaction_data["values"])
n_transaction_df['Date'] = np.array([datetime.datetime(1970, 1, 1) + datetime.timedelta(seconds=int(n_transaction_df["x"].iloc[k])) for k in range(len(n_transaction_df["x"]))])
fig, ax = plt.subplots(figsize=(6, 4))
ax.plot(n_transaction_df['Date'], n_transaction_df['y'], color=(0,0.4,0.5))
ax.set_xlabel('Date')
ax.set_title('Daily number of transactions')
sns.despine()
plt.savefig("../Figures/btc_n_transaction.pdf")
# -
# ## Bitcoin Network Electricity consumption
#
# The electricity consumption of the bitcoin network is estimated by the Cambridge Bitcoin Electricity Consumption Index [CBECI](https://cbeci.org/)
# +
elec_conso_df = pd.read_csv("../Data/BTC_electricity_consumption.csv")
fig, ax = plt.subplots(figsize=(6, 4))
ax.plot(pd.to_datetime(elec_conso_df['Date and Time']), elec_conso_df['GUESS'], color=(0,0.4,0.5))
ax.axhline(y=48, color = "black", linestyle = "--")
ax.text(16200, 52, "Portugal", fontsize=12)
ax.axhline(y=66.85, color = "black", linestyle = "--")
ax.text(16200, 70.85, "Austria", fontsize=12)
ax.axhline(y=131.8, color = "black", linestyle = "--")
ax.text(16200, 135.8, "Sweden", fontsize=12)
ax.set_xlabel('Date')
ax.set_title('Estimated yearly electricity consumption of the network in TWh')
sns.despine()
plt.savefig("../Figures/btc_elec_conso.pdf")
# -
| Python/Lecture1_blockchain_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (ptenv2)
# language: python
# name: ptenv
# ---
# +
from tqdm import tqdm
import numpy as np
import os, glob
import shutil
import matplotlib.pyplot as plt
import cv2
from copy import deepcopy
import sys
sys.path.append('/raid-dgx1/Hasnat/Covoiturage/')
from my_utilities_display_related import show_image, show_image_from_file, show_image_subplots
# -
#reads the yolo formatted bboxes and makes them into the required format
def get_formatted_info_from_file(img_size, labels_path, padding=0):
(height, width, _) = img_size
info = []
with open(labels_path) as lp:
for line in lp:
parsed = [float(x) for x in line.strip().split(' ')]
bb_center_x = round(parsed[1] * width)
bb_center_y = round(parsed[2] * height)
bb_width = round(parsed[3] * width)
bb_height = round(parsed[4] * height)
x1=round(bb_center_x - bb_width/2 + padding)
y1=round(bb_center_y - bb_height/2 + padding)
x2=round(bb_center_x + bb_width/2 + padding)
y2=round(bb_center_y + bb_height/2 + padding)
info.append([x1, y1, x2, y2, int(parsed[0])])
return info
# +
image_set = 'test' # 'trainval' #
#"__background__",
classes = ["aeroplane", "bicycle",
"bird", "boat", "bottle", "bus", "car",
"cat", "chair", "cow", "diningtable", "dog",
"horse", "motorbike", "person", "pottedplant",
"sheep", "sofa", "train", "tvmonitor"
]
save_path_images = '/raid-dgx1/Hasnat/ptlrn/efrp/data/VOC_yolo/'+image_set+'/images/'
save_path_anno = '/raid-dgx1/Hasnat/ptlrn/efrp/data/VOC_yolo/'+image_set+'/annotations/'
list_images = os.listdir(save_path_images)
#for jj in tqdm(list_images[:2]):
#in_file_name = _annotations[jj]
#out_file_name = save_path_anno + in_file_name.split('/')[-1].split('.')[0]+'.txt'
#convert_annotation_and_save(in_file_name, out_file_name)
#shutil.copyfile(_images[jj], save_path_images+_images[jj].split('/')[-1])
# +
fid = open(image_set+'.txt', 'w')
#jj = 0
for jj in tqdm(range(len(list_images))):
t_img_file = save_path_images + list_images[jj]
t_img = cv2.imread(t_img_file)
t_ext = '.' + t_img_file.split('.')[-1]
t_anno = save_path_anno + list_images[jj].replace(t_ext, '.txt')
t_info = get_formatted_info_from_file(t_img.shape, t_anno)
t_anno_str = list_images[jj] + ' '
for t_ob in t_info:
t_anno_str += ','.join([str(t_) for t_ in t_ob]) + ' '
t_anno_str += '\n'
fid.writelines(t_anno_str)
fid.close()
# -
#
target_dir = '/raid-dgx1/Hasnat/pytorch-YOLOv4/voc_images/'
for t_ in tqdm(list_images):
shutil.copyfile(save_path_images+t_, target_dir+t_)
| Write_YPT_style_annotation_file_VOC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Ic4_occAAiAT" colab_type="text"
# ##### Copyright 2018 The TensorFlow Authors.
# + id="ioaprt5q5US7" colab_type="code" colab={} cellView="form"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + id="yCl0eTNH5RS3" colab_type="code" colab={} cellView="form"
#@title MIT License
#
# Copyright (c) 2017 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a
# # copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
# + [markdown] id="ItXfxkxvosLH" colab_type="text"
# # Text classification with movie reviews
# + [markdown] id="NBM1gMpEGN_d" colab_type="text"
# This file has moved.
# + [markdown] id="hKY4XMc9o8iB" colab_type="text"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/keras/basic_text_classification"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/keras/basic_text_classification.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/keras/basic_text_classification.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
| samples/core/tutorials/keras/basic_text_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Prepare Data for Training & Validating & Testing
# ## Dataset
# - [**IMDB Large Movie Review Dataset**](http://ai.stanford.edu/~amaas/data/sentiment/)
# - Binary sentiment classification
# - Citation: [<NAME> et al., 2011](http://ai.stanford.edu/~amaas/papers/wvSent_acl2011.pdf)
# - 50,000 movie reviews for training and testing
# - Average review length: 231 vocab
# ---
# - [**Yelp reviews-full**](http://xzh.me/docs/charconvnet.pdf)
# - Multiclass sentiment classification (5 stars)
# - Citation: [Xiang Zhang et al., 2015](https://arxiv.org/abs/1509.01626)
# - 650,000 training samples and 50,000 testing samples (Nums of each star are equal)
# - Average review length: 140 vocab
# ---
# - [**Yelp reviews-polarity**](http://xzh.me/docs/charconvnet.pdf)
# - Binary sentiment classification
# - Citation: [Xiang Zhang et al., 2015](https://arxiv.org/abs/1509.01626)
# - 560,000 training samples and 38,000 testing samples (Nums of positive or negative samples are equal)
# - Average review length: 140 vocab
# ---
# - [**Douban Movie Reviews**](https://drive.google.com/open?id=1DsmQfB1Ff_BUoxOv4kfUMg7Y8M7tHB9F)
# - My Custom Chinese movie reviews dataset scraped by python extension package requests
# - Binary sentiment classification
# - 750,000 movie reviews from [Douban](https://movie.douban.com/), 650,000 samples for training and 100,000 samples for testing (Num of positive or negative samples is equal)
# - Average review length: 52 character, 28 vocab
# +
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings('ignore')
from utils import clean_text, clean_text_zh
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# -
# ## IMDB
df = pd.read_csv('dataset/IMDB/imdb_master.csv', encoding='latin-1')
df.head()
df = df[df['label'] != 'unsup']
df['label'] = df['label'].map({'neg':0, 'pos':1})
df.head()
df['Processed_Reviews'] = df['review'].apply(clean_text)
df_train, df_test = train_test_split(df, test_size=0.1, random_state=0, stratify=df.label)
df_train, df_validate = train_test_split(df_train, test_size=5000, random_state=0, stratify=df_train.label)
print("Train set size: ", len(df_train))
print("Validation set size: ", len(df_validate))
print("Test set size: ", len(df_test))
plt.figure()
_ = plt.hist(df_train.Processed_Reviews.apply(lambda x: len(x.split())), bins = 100)
plt.title('Distribution of sentence length')
plt.xlabel('Length')
plt.ylabel('Counts')
# ### Word sequence to num sequence
word_count = {}
def count(x):
for word in x.split():
word_count[word] = word_count.get(word, 0) + 1
_ = df_train.Processed_Reviews.apply(count)
df_word_count = pd.DataFrame(list(word_count.items()), columns=['word', 'count'])
df_word_count.sort_values(by='count', ascending=False, inplace=True)
df_word_count.set_index('word', inplace=True)
df_word_count.drop(['the', 'a', 'and', 'of', 'to', 'br', 'in', 'this', 'that', 's'], inplace=True)
vocab_size = 20000
df_word_count = df_word_count[:vocab_size-3]
word2num_series = df_word_count.reset_index().reset_index().set_index('word')['index'] + 3 # reverse 0, 1, 2 for padding, BOS, EOS
X_train = df_train.Processed_Reviews.apply(lambda x: np.array([word2num_series[word] for word in x.split() if word in word2num_series], dtype=np.int32))
y_train = df_train.label
# ### Process validation set and test set
X_val = df_validate.Processed_Reviews.apply(lambda x: np.array([word2num_series[word] for word in x.split() if word in word2num_series], dtype=np.int32))
X_test = df_test.Processed_Reviews.apply(lambda x: np.array([word2num_series[word] for word in x.split() if word in word2num_series], dtype=np.int32))
y_val = df_validate.label
y_test = df_test.label
# ### Save the data into h5 file
# +
X_train.to_hdf('dataset/IMDB/X_train.h5', key='s', model='w')
y_train.to_hdf('dataset/IMDB/y_train.h5', key='s', model='w')
X_val.to_hdf('dataset/IMDB/X_val.h5', key='s', model='w')
y_val.to_hdf('dataset/IMDB/y_val.h5', key='s', model='w')
X_test.to_hdf('dataset/IMDB/X_test.h5', key='s', model='w')
y_test.to_hdf('dataset/IMDB/y_test.h5', key='s', model='w')
word2num_series.to_hdf('dataset/IMDB/word2num_series.h5', key='s', model='w')
# -
# # Yelp Reviews Polarity
df = pd.read_csv('dataset/yelp_review_polarity_csv/train.csv', header=None)
df_test = pd.read_csv('dataset/yelp_review_polarity_csv/test.csv', header=None)
df.columns = ['label', 'review']
df['label'] = df['label'].map({1:0, 2:1})
df_test.columns = ['label', 'review']
df_test['label'] = df_test['label'].map({1:0, 2:1})
df.count()
df_test.count()
df['Processed_Reviews'] = df['review'].apply(lambda x: clean_text(x))
df_test['Processed_Reviews'] = df_test['review'].apply(lambda x: clean_text(x))
df['Processed_Reviews'].apply(lambda x: len(x.split())).mean()
df_train, df_validate = train_test_split(df, test_size=40000, random_state=0, stratify=df.label)
print("Train set size: ", len(df_train))
print("Validation set size: ", len(df_validate))
print("Test set size: ", len(df_test))
plt.figure()
_ = plt.hist(df_train.Processed_Reviews.apply(lambda x: len(x.split())), bins = 100)
plt.title('Distribution of sentence length')
plt.xlabel('Length')
plt.ylabel('Counts')
word_count = {}
def count(x):
for word in x.split():
word_count[word] = word_count.get(word, 0) + 1
_ = df_train.Processed_Reviews.apply(count)
df_word_count = pd.DataFrame(list(word_count.items()), columns=['word', 'count'])
df_word_count.sort_values(by='count', ascending=False, inplace=True)
df_word_count.set_index('word', inplace=True)
df_word_count.drop(['the', 'a', 'and', 'of', 'to', 'br', 'in', 'this', 'that', 's', 'n'], inplace=True)
vocab_size = 40000
df_word_count = df_word_count[:vocab_size-3]
word2num_series = df_word_count.reset_index().reset_index().set_index('word')['index'] + 3 # reverse 0, 1, 2 for padding, BOS, EOS
X_train = df_train.Processed_Reviews.apply(lambda x: np.array([word2num_series[word] for word in x.split() if word in word2num_series], dtype=np.int32))
y_train = df_train.label
X_val = df_validate.Processed_Reviews.apply(lambda x: np.array([word2num_series[word] for word in x.split() if word in word2num_series], dtype=np.int32))
X_test = df_test.Processed_Reviews.apply(lambda x: np.array([word2num_series[word] for word in x.split() if word in word2num_series], dtype=np.int32))
y_val = df_validate.label
y_test = df_test.label
# +
X_train.to_hdf('dataset/yelp_review_polarity_csv/X_train.h5', key='s', model='w')
y_train.to_hdf('dataset/yelp_review_polarity_csv/y_train.h5', key='s', model='w')
X_val.to_hdf('dataset/yelp_review_polarity_csv/X_val.h5', key='s', model='w')
y_val.to_hdf('dataset/yelp_review_polarity_csv/y_val.h5', key='s', model='w')
X_test.to_hdf('dataset/yelp_review_polarity_csv/X_test.h5', key='s', model='w')
y_test.to_hdf('dataset/yelp_review_polarity_csv/y_test.h5', key='s', model='w')
word2num_series.to_hdf('dataset/yelp_review_polarity_csv/word2num_series.h5', key='s', model='w')
# -
# ## Yelp Reviews Full
df = pd.read_csv('dataset/yelp_review_full_csv/train.csv', header=None)
df_test = pd.read_csv('dataset/yelp_review_full_csv/test.csv', header=None)
df.columns = ['label', 'review']
df_test.columns = ['label', 'review']
df['Processed_Reviews'] = df['review'].apply(lambda x: clean_text(x))
df_test['Processed_Reviews'] = df_test['review'].apply(lambda x: clean_text(x))
df['Processed_Reviews'].apply(lambda x: len(x.split())).mean()
df_train, df_validate = train_test_split(df, test_size=50000, random_state=0, stratify=df.label)
print("Train set size: ", len(df_train))
print("Validation set size: ", len(df_validate))
print("Test set size: ", len(df_test))
plt.figure()
_ = plt.hist(df_train.Processed_Reviews.apply(lambda x: len(x.split())), bins = 100)
plt.title('Distribution of sentence length')
plt.xlabel('Length')
plt.ylabel('Counts')
word_count = {}
def count(x):
for word in x.split():
word_count[word] = word_count.get(word, 0) + 1
_ = df_train.Processed_Reviews.apply(count)
df_word_count = pd.DataFrame(list(word_count.items()), columns=['word', 'count'])
df_word_count.sort_values(by='count', ascending=False, inplace=True)
df_word_count.set_index('word', inplace=True)
df_word_count.drop(['the', 'a', 'and', 'of', 'to', 'br', 'in', 'this', 'that', 's', 'n'], inplace=True)
vocab_size = 40000
df_word_count = df_word_count[:vocab_size-3]
word2num_series = df_word_count.reset_index().reset_index().set_index('word')['index'] + 3 # reverse 0, 1, 2 for padding, BOS, EOS
X_train = df_train.Processed_Reviews.apply(lambda x: np.array([word2num_series[word] for word in x.split() if word in word2num_series], dtype=np.int32))
y_train = df_train.label
X_val = df_validate.Processed_Reviews.apply(lambda x: np.array([word2num_series[word] for word in x.split() if word in word2num_series], dtype=np.int32))
X_test = df_test.Processed_Reviews.apply(lambda x: np.array([word2num_series[word] for word in x.split() if word in word2num_series], dtype=np.int32))
y_val = df_validate.label
y_test = df_test.label
# +
X_train.to_hdf('dataset/yelp_review_full_csv/X_train.h5', key='s', model='w')
y_train.to_hdf('dataset/yelp_review_full_csv/y_train.h5', key='s', model='w')
X_val.to_hdf('dataset/yelp_review_full_csv/X_val.h5', key='s', model='w')
y_val.to_hdf('dataset/yelp_review_full_csv/y_val.h5', key='s', model='w')
X_test.to_hdf('dataset/yelp_review_full_csv/X_test.h5', key='s', model='w')
y_test.to_hdf('dataset/yelp_review_full_csv/y_test.h5', key='s', model='w')
word2num_series.to_hdf('dataset/yelp_review_full_csv/word2num_series.h5', key='s', model='w')
# -
# ## Douban Dataset
df = pd.read_csv('dataset/Douban/train.csv')
df_test = pd.read_csv('dataset/Douban/test.csv')
len(df)
len(df_test)
df['Processed_Reviews'] = df['content'].apply(clean_text_zh)
df_test['Processed_Reviews'] = df_test['content'].apply(clean_text_zh)
import jieba
df['Processed_Reviews'].apply(lambda x: len([word for word in jieba.cut(x) if word.strip()])).mean()
df['content'].apply(lambda x: len(x)).mean()
len(df)
df_train, df_validate = train_test_split(df, test_size=50000, random_state=0, stratify=df.label)
print("Train set size: ", len(df_train))
print("Validation set size: ", len(df_validate))
print("Test set size: ", len(df_test))
plt.figure()
_ = plt.hist(df_train.Processed_Reviews.apply(lambda x: len([word for word in jieba.cut(x) if word.strip()])), bins = 100)
plt.title('Distribution of sentence length')
plt.xlabel('Length')
plt.ylabel('Counts')
word_count = {}
def count(x):
for word in jieba.cut(x):
word_count[word] = word_count.get(word, 0) + 1
_ = df_train.Processed_Reviews.apply(count)
df_word_count = pd.DataFrame(list(word_count.items()), columns=['word', 'count'])
df_word_count.sort_values(by='count', ascending=False, inplace=True)
df_word_count.set_index('word', inplace=True)
df_word_count.drop([' ', '的', '了', '是', '都', '在', '就'], inplace=True)
vocab_size = 40000
df_word_count = df_word_count[:vocab_size-3]
word2num_series = df_word_count.reset_index().reset_index().set_index('word')['index'] + 3 # reverse 0, 1, 2 for padding, BOS, EOS
X_train = df_train.Processed_Reviews.apply(lambda x: np.array([word2num_series[word] for word in jieba.cut(x) if word in word2num_series], dtype=np.int32))
y_train = df_train.label
X_val = df_validate.Processed_Reviews.apply(lambda x: np.array([word2num_series[word] for word in jieba.cut(x) if word in word2num_series], dtype=np.int32))
X_test = df_test.Processed_Reviews.apply(lambda x: np.array([word2num_series[word] for word in jieba.cut(x) if word in word2num_series], dtype=np.int32))
y_val = df_validate.label
y_test = df_test.label
# +
X_train.to_hdf('dataset/Douban/X_train.h5', key='s', model='w')
y_train.to_hdf('dataset/Douban/y_train.h5', key='s', model='w')
X_val.to_hdf('dataset/Douban/X_val.h5', key='s', model='w')
y_val.to_hdf('dataset/Douban/y_val.h5', key='s', model='w')
X_test.to_hdf('dataset/Douban/X_test.h5', key='s', model='w')
y_test.to_hdf('dataset/Douban/y_test.h5', key='s', model='w')
word2num_series.to_hdf('dataset/Douban/word2num_series.h5', key='s', model='w')
# -
# ## Amazon Review Polarity
#
df = pd.read_csv('dataset/amazon_review_polarity_csv/train.csv', header=None)
df_test = pd.read_csv('dataset/amazon_review_polarity_csv/test.csv', header=None)
len(df)
df.columns = ['label', 'title', 'review']
df_test.columns = ['label', 'title', 'review']
df['label'] = df['label'].map({1:0, 2:1})
df_test['label'] = df_test['label'].map({1:0, 2:1})
df.head()
df['review'].apply(lambda x: len(x.split())).mean()
df['Processed_Reviews'] = df['review'].apply(lambda x: clean_text(x))
df_test['Processed_Reviews'] = df_test['review'].apply(lambda x: clean_text(x))
df_train, df_validate = train_test_split(df, test_size=400000, random_state=0, stratify=df.label)
print("Train set size: ", len(df_train))
print("Validation set size: ", len(df_validate))
print("Test set size: ", len(df_test))
word_count = {}
def count(x):
for word in x.split():
word_count[word] = word_count.get(word, 0) + 1
_ = df_train.Processed_Reviews.apply(count)
df_word_count = pd.DataFrame(list(word_count.items()), columns=['word', 'count'])
df_word_count.sort_values(by='count', ascending=False, inplace=True)
df_word_count.set_index('word', inplace=True)
df_word_count.drop(['the', 'a', 'and', 'of', 'to', 'in', 'this', 'that', 's', 'n', 'for'], inplace=True)
vocab_size = 40000
df_word_count = df_word_count[:vocab_size-3]
word2num_series = df_word_count.reset_index().reset_index().set_index('word')['index'] + 3 # reverse 0, 1, 2 for padding, BOS, EOS
X_train = df_train.Processed_Reviews.apply(lambda x: np.array([word2num_series[word] for word in x.split() if word in word2num_series], dtype=np.int32))
y_train = df_train.label
X_val = df_validate.Processed_Reviews.apply(lambda x: np.array([word2num_series[word] for word in x.split() if word in word2num_series], dtype=np.int32))
X_test = df_test.Processed_Reviews.apply(lambda x: np.array([word2num_series[word] for word in x.split() if word in word2num_series], dtype=np.int32))
y_val = df_validate.label
y_test = df_test.label
# +
X_train.to_hdf('dataset/amazon_review_polarity_csv/X_train.h5', key='s', model='w')
y_train.to_hdf('dataset/amazon_review_polarity_csv/y_train.h5', key='s', model='w')
X_val.to_hdf('dataset/amazon_review_polarity_csv/X_val.h5', key='s', model='w')
y_val.to_hdf('dataset/amazon_review_polarity_csv/y_val.h5', key='s', model='w')
X_test.to_hdf('dataset/amazon_review_polarity_csv/X_test.h5', key='s', model='w')
y_test.to_hdf('dataset/amazon_review_polarity_csv/y_test.h5', key='s', model='w')
word2num_series.to_hdf('dataset/amazon_review_polarity_csv/word2num_series.h5', key='s', model='w')
# -
| Sentiment-Analylsis-based-on-Attention-Mechanism/Data Preprocess.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Highcharts Demos
# ================
# Bubble chart: http://www.highcharts.com/demo/bubble
# ----------------------------------------------------
# +
from highcharts import Highchart
H = Highchart(width=850, height=400)
options = {
'chart': {
'type': 'bubble',
'zoomType': 'xy'
},
'title': {
'text': 'Highcharts Bubbles'
},
}
data1 = [[97, 36, 79], [94, 74, 60], [68, 76, 58], [64, 87, 56], [68, 27, 73], [74, 99, 42], [7, 93, 87], [51, 69, 40], [38, 23, 33], [57, 86, 31]]
data2 = [[25, 10, 87], [2, 75, 59], [11, 54, 8], [86, 55, 93], [5, 3, 58], [90, 63, 44], [91, 33, 17], [97, 3, 56], [15, 67, 48], [54, 25, 81]]
data3 = [[47, 47, 21], [20, 12, 4], [6, 76, 91], [38, 30, 60], [57, 98, 64], [61, 17, 80], [83, 60, 13], [67, 78, 75], [64, 12, 10], [30, 77, 82]]
H.set_dict_options(options)
H.add_data_set(data1, 'bubble')
H.add_data_set(data2, 'bubble')
H.add_data_set(data3, 'bubble')
H
| examples/ipynb/highcharts/bubble.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/computacao-io/estrutura-de-dados/blob/master/estrutura_de_dados.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="tmbSu-DX5rOF" colab_type="text"
# # Estrutura de Dados com python
# + [markdown] id="LkOxfOYz56SW" colab_type="text"
# ## Estruturas Lineares
# + [markdown] id="MFGMLf3mEbXJ" colab_type="text"
# ### Tuple - São imutáveis
# + id="BqjsNxTtJuUw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="4fcb3b0c-162a-4aca-aeff-066f07266c23"
tuple1 = (1, 2, 3, 4)
tuple2 = tuple([1, 2, 3, 4])
print(tuple1, tuple2)
# + [markdown] id="TB4-iVMb6qod" colab_type="text"
# ### List - São mutáveis
# + id="FYOrkktXKbrL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="37631286-e5a8-42b5-81c7-75fd02c612ba"
list1 = [1, 2, 3, 4]
list2 = list([1, 2, 3, 4])
print(list1, list2)
# + id="zfKuzd8YLCfS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="90f40c42-1c78-4b79-d1a1-d3f1c540cdae"
list1[1] = 3
print(list1)
# + id="TF1X5p5nLOWx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="8ebf7023-814f-4d64-db57-1cc1c30b95ef"
list1.append(5)
print(list1)
# + id="Z5RBH4npLg0o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a6195118-8feb-4cd1-e19b-d23febdf05dd"
list1.pop()
# + id="FwkHzN-bLnxn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="b1387a91-2232-4a52-8660-3a2ddcff0577"
print(list1)
# + id="s56zQjnRLxik" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="712664db-0211-420f-998a-a33b50573f72"
list1.pop(0)
# + id="xERI1E1qL2Nr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="68e8b7b5-512c-4aa7-af8b-22891be79810"
print(list1)
# + id="BGHCJWEuNX5H" colab_type="code" colab={}
list1.insert(1, 2)
# + id="z52JF3mkNmfm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a3523cf4-b571-432a-e36e-e9c1182c66fe"
print(list1)
# + [markdown] id="sjYTo_pIMFPE" colab_type="text"
# #### Pilha
# + id="wTolJMsMMJJf" colab_type="code" colab={}
list1.append(5)
# + id="iLMec8h-Mfni" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="0f013780-2cf8-4865-ea5b-908a89250543"
print(list1)
# + id="629llV-vMudN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="57315ce3-ef69-486d-9794-eb0b732ea2dd"
list1.pop()
# + [markdown] id="kA6vVklJM6qB" colab_type="text"
# #### Fila
# + id="-kQTunlcM8_t" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="0b03395e-a0fd-4bf4-fa31-166009925a06"
list1.append(6)
print(list1)
# + id="b50S2zOgNFum" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="8d2c9584-17c8-4c44-ed14-05a63f4c5974"
list1.pop(0)
# + [markdown] id="vjv0_AyfASkz" colab_type="text"
# ### Deque
# + id="ZlLAYaz8NvA9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="3cdd676f-0ddd-4cef-9a69-f7e17b69a6f8"
from collections import deque
queue = deque([1, 2, 3, 4])
print(queue)
# + id="hEZJCw0rOCA4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="7ae3b0fc-d0e8-48eb-bf1c-60b76671f519"
queue.append(5)
print(queue)
# + id="gw2NSShfOHx6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="602d0fb1-3522-4a36-f827-c168ac22e8c9"
queue.popleft()
# + [markdown] id="Fjq9G6XBAaaR" colab_type="text"
# ### Array
# + id="c66d5tmKOZ8p" colab_type="code" colab={}
from array import array
array1 = array('l', [1, 2, 3, 4])
# + id="tPpltNb5Ou3i" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="9ce83200-f097-402d-af20-5803cb191508"
print(array1)
# + [markdown] id="-xSVkv5u6BSF" colab_type="text"
# ## Estruturas Associativas
# + [markdown] id="eQB0bk6oAnJ2" colab_type="text"
# ### Set
# + id="SSfrYQVxPfFK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="072879b7-770a-4766-deb3-56fbc8915e93"
set3 = set([1, 2, 3, 4])
print(set3)
# + id="QzKY4nN_O5oA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="1629802f-953b-43df-fd1a-af26a65c9479"
set1 = {1, 2, 3, 4}
print(set1)
# + id="wmtKmgrAO-0b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="d00530bd-d910-4044-83e9-d64ed3239a7b"
set2 = {1, 3, 4, 5}
print(set2)
# + id="cuAe0plnPEdM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="1191f56a-65d5-4d44-f8bc-46df3aa1b544"
set1 | set2
# + id="eALW8m8BPRTh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="2a8342cf-93fa-4570-838a-6a27654dbea9"
set1 ^ set2
# + id="y3YWx_FwPXRM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="39b370fe-0681-47ee-8920-ee0bf53d6b6a"
set1 & set2
# + [markdown] id="sXWtQJM2Ar4M" colab_type="text"
# ### Dict
# + id="fC95WTcsPoo-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="7e58851f-1243-4953-a270-02da4550e7fa"
dict1 = {'first': 1, 'last': 2}
print(dict1)
# + id="K-WTYNE9Pyow" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="621cd2bb-7e66-4859-9d24-71ee9637128a"
dict2 = dict({'one': 1, 'two': 2})
print(dict2)
# + id="pYJuk2rPQFbP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="8104207d-ffcd-41e8-a0a8-ff322f6b9c3f"
dict1.values()
# + id="TBPu5eqKQLjb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="121238a9-9720-4a7e-9e43-bb5f05f34831"
dict2.items()
# + id="rPITL2GKQRQt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 88} outputId="00eda386-9fa0-4974-93f5-9609ac212661"
for k, v in dict2.items():
print('Key: {}'.format(k))
print('Values: {}'.format(v))
# + [markdown] id="XuFY0Thp6Em9" colab_type="text"
# # Referências
# + [markdown] id="qV5tLVrHBJO7" colab_type="text"
# Wiki Unicamp
#
# * [Princípios de estruturas de dados](http://calhau.dca.fee.unicamp.br/wiki/images/5/5b/Cap2.pdf)
#
# Documentação Oficial do Python
#
# * [Tutorial Estrutura de Dados](https://docs.python.org/pt-br/3/tutorial/datastructures.html#looping-techniques)
# * [Tipos de dados - collections](https://docs.python.org/pt-br/3/library/collections.html)
# * [Tipos de dados - array](https://docs.python.org/pt-br/3/library/array.html)
#
# Palestra de <NAME> sobre conjuntos
#
# * [Python Set Practice](https://speakerdeck.com/ramalho/python-set-practice-at-pycon)
#
| estrutura_de_dados.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Sklearn
# ## Bike Sharing Demand
# Задача на kaggle: https://www.kaggle.com/c/bike-sharing-demand
#
# По историческим данным о прокате велосипедов и погодным условиям необходимо оценить спрос на прокат велосипедов.
#
# В исходной постановке задачи доступно 11 признаков: https://www.kaggle.com/c/prudential-life-insurance-assessment/data
#
# В наборе признаков присутсвуют вещественные, категориальные, и бинарные данные.
#
# Для демонстрации используется обучающая выборка из исходных данных train.csv, файлы для работы прилагаются.
# ### Библиотеки
# +
from sklearn import cross_validation, grid_search, linear_model, metrics
import numpy as np
import pandas as pd
# -
# %pylab inline
# ### Загрузка данных
raw_data = pd.read_csv('bike_sharing_demand.csv', header = 0, sep = ',')
raw_data.head()
# ***datetime*** - hourly date + timestamp
#
# ***season*** - 1 = spring, 2 = summer, 3 = fall, 4 = winter
#
# ***holiday*** - whether the day is considered a holiday
#
# ***workingday*** - whether the day is neither a weekend nor holiday
#
# ***weather*** - 1: Clear, Few clouds, Partly cloudy, Partly cloudy
# 2: Mist + Cloudy, Mist + Broken clouds, Mist + Few clouds, Mist
# 3: Light Snow, Light Rain + Thunderstorm + Scattered clouds, Light Rain + Scattered clouds
# 4: Heavy Rain + Ice Pallets + Thunderstorm + Mist, Snow + Fog
#
# ***temp*** - temperature in Celsius
#
# ***atemp*** - "feels like" temperature in Celsius
#
# ***humidity*** - relative humidity
#
# ***windspeed*** - wind speed
#
# ***casual*** - number of non-registered user rentals initiated
#
# ***registered*** - number of registered user rentals initiated
#
# ***count*** - number of total rentals
print raw_data.shape
raw_data.isnull().values.any()
# ### Предобработка данных
# #### Типы признаков
raw_data.info()
raw_data.datetime = raw_data.datetime.apply(pd.to_datetime)
raw_data['month'] = raw_data.datetime.apply(lambda x : x.month)
raw_data['hour'] = raw_data.datetime.apply(lambda x : x.hour)
raw_data.head()
# #### Обучение и отложенный тест
train_data = raw_data.iloc[:-1000, :]
hold_out_test_data = raw_data.iloc[-1000:, :]
print raw_data.shape, train_data.shape, hold_out_test_data.shape
print 'train period from {} to {}'.format(train_data.datetime.min(), train_data.datetime.max())
print 'evaluation period from {} to {}'.format(hold_out_test_data.datetime.min(), hold_out_test_data.datetime.max())
# #### Данные и целевая функция
#обучение
train_labels = train_data['count'].values
train_data = train_data.drop(['datetime', 'count'], axis = 1)
#тест
test_labels = hold_out_test_data['count'].values
test_data = hold_out_test_data.drop(['datetime', 'count'], axis = 1)
# #### Целевая функция на обучающей выборке и на отложенном тесте
# +
pylab.figure(figsize = (16, 6))
pylab.subplot(1,2,1)
pylab.hist(train_labels)
pylab.title('train data')
pylab.subplot(1,2,2)
pylab.hist(test_labels)
pylab.title('test data')
# -
# #### Числовые признаки
numeric_columns = ['temp', 'atemp', 'humidity', 'windspeed', 'casual', 'registered', 'month', 'hour']
train_data = train_data[numeric_columns]
test_data = test_data[numeric_columns]
train_data.head()
test_data.head()
# ### Модель
regressor = linear_model.SGDRegressor(random_state = 0)
regressor.fit(train_data, train_labels)
metrics.mean_absolute_error(test_labels, regressor.predict(test_data))
print test_labels[:10]
print regressor.predict(test_data)[:10]
regressor.coef_
# ### Scaling
from sklearn.preprocessing import StandardScaler
#создаем стандартный scaler
scaler = StandardScaler()
scaler.fit(train_data, train_labels)
scaled_train_data = scaler.transform(train_data)
scaled_test_data = scaler.transform(test_data)
regressor.fit(scaled_train_data, train_labels)
metrics.mean_absolute_error(test_labels, regressor.predict(scaled_test_data))
print test_labels[:10]
print regressor.predict(scaled_test_data)[:10]
# ### Подозрительно хорошо?
print regressor.coef_
print map(lambda x : round(x, 2), regressor.coef_)
train_data.head()
train_labels[:10]
np.all(train_data.registered + train_data.casual == train_labels)
train_data.drop(['casual', 'registered'], axis = 1, inplace = True)
test_data.drop(['casual', 'registered'], axis = 1, inplace = True)
scaler.fit(train_data, train_labels)
scaled_train_data = scaler.transform(train_data)
scaled_test_data = scaler.transform(test_data)
regressor.fit(scaled_train_data, train_labels)
metrics.mean_absolute_error(test_labels, regressor.predict(scaled_test_data))
print map(lambda x : round(x, 2), regressor.coef_)
# ### Pipeline
from sklearn.pipeline import Pipeline
#создаем pipeline из двух шагов: scaling и классификация
pipeline = Pipeline(steps = [('scaling', scaler), ('regression', regressor)])
pipeline.fit(train_data, train_labels)
metrics.mean_absolute_error(test_labels, pipeline.predict(test_data))
# ### Подбор параметров
pipeline.get_params().keys()
parameters_grid = {
'regression__loss' : ['huber', 'epsilon_insensitive', 'squared_loss', ],
'regression__n_iter' : [3, 5, 10, 50],
'regression__penalty' : ['l1', 'l2', 'none'],
'regression__alpha' : [0.0001, 0.01],
'scaling__with_mean' : [0., 0.5],
}
grid_cv = grid_search.GridSearchCV(pipeline, parameters_grid, scoring = 'mean_absolute_error', cv = 4)
# %%time
grid_cv.fit(train_data, train_labels)
print grid_cv.best_score_
print grid_cv.best_params_
# ### Оценка по отложенному тесту
metrics.mean_absolute_error(test_labels, grid_cv.best_estimator_.predict(test_data))
np.mean(test_labels)
test_predictions = grid_cv.best_estimator_.predict(test_data)
print test_labels[:10]
print test_predictions[:10]
# +
pylab.figure(figsize=(16, 6))
pylab.subplot(1,2,1)
pylab.grid(True)
pylab.scatter(train_labels, pipeline.predict(train_data), alpha=0.5, color = 'red')
pylab.scatter(test_labels, pipeline.predict(test_data), alpha=0.5, color = 'blue')
pylab.title('no parameters setting')
pylab.xlim(-100,1100)
pylab.ylim(-100,1100)
pylab.subplot(1,2,2)
pylab.grid(True)
pylab.scatter(train_labels, grid_cv.best_estimator_.predict(train_data), alpha=0.5, color = 'red')
pylab.scatter(test_labels, grid_cv.best_estimator_.predict(test_data), alpha=0.5, color = 'blue')
pylab.title('grid search')
pylab.xlim(-100,1100)
pylab.ylim(-100,1100)
| 2 Supervised learning/Lectures notebooks/7 bike sharing demand part 1/sklearn.case_part1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ysAK7mqy8f6Q"
# # Rodent Inspection
# + [markdown] id="tu_Geg2M8joY"
# ## Cargamos las librerías
# + id="GYE0Kp-2z3VT"
from sklearn.metrics import accuracy_score
import pandas as pd
from sklearn.model_selection import train_test_split
import xgboost as xgb
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
import pickle
from sklearn.preprocessing import LabelEncoder
# + [markdown] id="Zf6iCFhE8nLA"
# ## Cargamos los datos
# + id="KxGRznIVwppw"
#data = pd.read_csv("/src/utils/Rodent_Inspection.csv")
data = pd.read_csv("/content/drive/MyDrive/Rodent_inspection/data/Rodent_Inspection.csv")
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="-nUM9nVHz-vE" outputId="ef822224-2062-405b-a4da-6112c4d35002"
data.head()
# + colab={"base_uri": "https://localhost:8080/"} id="j7HEELw60qDa" outputId="c103667b-b1a9-49d8-e119-c3f9869dbe67"
data.columns
# + [markdown] id="2VKqCZEr3f4p"
# ## Entrenamiento
#
# Dividimos en train y test
# + id="EgV5fkaM3P7E"
X = data.drop(['result'], axis=1)
y = data['result']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.10, random_state=12345)
# + [markdown] id="MTamojgC2og_"
# ## Función para obtener los pkl de los mejores modelos
# + id="2ASxVJrywog5"
def complete_model(data):
columns_to_drop = ["job_ticket_or_work_order_id","inspection_date"]
data = data.drop(columns_to_drop,axis=1)
data = data.drop(data[data.result == "Bait applied"].index)
data.drop_duplicates()
def conditions(s):
if (s['result'] == "Passed") or (s['result'] == "Monitoring visit"):
return 0
else:
return 1
data['result'] = data.apply(conditions, axis=1)
Insp = pd.get_dummies(data['inspection_type'])
Insp=Insp.join(data.job_id)
data = pd.merge(data.drop(['inspection_type'], axis = 1),Insp, on="job_id")
le = LabelEncoder()
data['boro_code'] = le.fit_transform(data['boro_code'])
data['result'] = le.fit_transform(data['result'])
data['job_id'] = le.fit_transform(data['job_id'].astype(str))
X = data.drop(['result'], axis=1)
y = data['result']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=12345)
# xgboost
model_xgb = xgb.XGBClassifier(
n_estimators=10,
max_depth=7,
learning_rate=0.4,
colsample_bytree=0.6,
missing=-999,
random_state=66)
model_xgb.fit(X_train, y_train)
pickle.dump(model_xgb, open('/content/drive/MyDrive/Rodent_inspection/data/entrenamiento_xgb.pkl', 'wb'))
# logistic regression
model_lr = LogisticRegression(
penalty = 'l2',
C= 1,
solver='lbfgs')
model_lr.fit(X_train, y_train)
pickle.dump(model_lr, open('/content/drive/MyDrive/Rodent_inspection/data/entrenamiento_lr.pkl', 'wb'))
# KNN
model_knn=KNeighborsClassifier(
n_neighbors=10,
weights='uniform',
algorithm='auto')
model_knn.fit(X_train, y_train)
pickle.dump(model_knn, open('/content/drive/MyDrive/Rodent_inspection/data/entrenamiento_knn.pkl', 'wb'))
acc_xgb = accuracy_score(y_test, pickle.load(open('/content/drive/MyDrive/Rodent_inspection/data/entrenamiento_xgb.pkl', 'rb')).predict(X_test))
acc_lr = accuracy_score(y_test, pickle.load(open('/content/drive/MyDrive/Rodent_inspection/data/entrenamiento_lr.pkl', 'rb')).predict(X_test))
acc_knn = accuracy_score(y_test, pickle.load(open('/content/drive/MyDrive/Rodent_inspection/data/entrenamiento_knn.pkl', 'rb')).predict(X_test))
acc_diccionario = {"XGB": acc_xgb, "LR": acc_lr, "KNN": acc_knn}
print("####### Las precisiones de los modelos son: ", acc_diccionario)
mejor_modelo=max(acc_diccionario, key=acc_diccionario.get)
acc_mejor_modelo=max(acc_diccionario.values())
print("####### Mejor modelo: ", mejor_modelo)
print("####### El accuracy del mejor modelo es: ", acc_mejor_modelo)
# + [markdown] id="mXw3M5UA5UBv"
# ## Desempeño
# + colab={"base_uri": "https://localhost:8080/"} id="_NEaBmhI0CjK" outputId="90515bf0-29b2-4e80-c18e-40e8f6916fe1"
complete_model(pd.concat([X_train, y_train], axis=1))
# + [markdown] id="dqkCjPXR5Zni"
# ## Re-entrenamiento
#
# Con el 10% de la muestra restante
# + colab={"base_uri": "https://localhost:8080/"} id="XyDPsKVV5fu7" outputId="862d5994-75be-4062-d171-ab440d795d81"
complete_model(pd.concat([X_test, y_test], axis=1))
# + [markdown] id="Q_C1ZySS5j4g"
# Muy parecido
# + colab={"base_uri": "https://localhost:8080/"} id="yJ3A8tR31tXk" outputId="7091523c-794e-4558-9d0b-078cf25c2452"
# Aquí se muestra el uso del pkl con la base completa
# Load from file
pkl_filename = "/content/drive/MyDrive/Rodent_inspection/data/entrenamiento_lr.pkl"
with open(pkl_filename, 'rb') as file:
pickle_model = pickle.load(file)
#Limpiamos
columns_to_drop = ["job_ticket_or_work_order_id","inspection_date"]
data1 = data.drop(columns_to_drop,axis=1)
data1 = data1.drop(data1[data1.result == "Bait applied"].index)
data1.drop_duplicates()
def conditions(s):
if (s['result'] == "Passed") or (s['result'] == "Monitoring visit"):
return 0
else:
return 1
data1['result'] = data1.apply(conditions, axis=1)
Insp = pd.get_dummies(data1['inspection_type'])
Insp=Insp.join(data1.job_id)
data1 = pd.merge(data1.drop(['inspection_type'], axis = 1),Insp, on="job_id")
le = LabelEncoder()
data1['boro_code'] = le.fit_transform(data1['boro_code'])
data1['result'] = le.fit_transform(data1['result'])
data1['job_id'] = le.fit_transform(data1['job_id'].astype(str))
X = data1.drop(['result'], axis=1)
y = data1['result']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=1)
# Calculate the accuracy score and predict target values
score = pickle_model.score(X_test, y_test)
print("Test score: {0:.2f} %".format(100 * score))
Ypredict = pickle_model.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="6-7AuLZf6uz4" outputId="beaf83af-df29-4a40-915e-21427970e2a6"
Ypredict
# + [markdown] id="ziDOmLM66y9z"
# ## Ejemplo con un dato nuevo
# + id="ABo4m7nU64jG"
d = {'job_id': [55874], 'boro_code': [3], 'zip_code': [12345], 'latitude': [40.825241], 'longitude': [-73.988733], 'BAIT': [1], 'CLEAN_UPS': [0], 'Compliance': [0], 'Initial': [0], 'STOPPAGE': [0]}
df = pd.DataFrame(d)
# + colab={"base_uri": "https://localhost:8080/", "height": 81} id="KetRbfwR71-l" outputId="d891fc2a-66df-4aeb-ffe8-7fc3a1a586cf"
df
# + colab={"base_uri": "https://localhost:8080/"} id="yv11uoCg8OY_" outputId="edf0968b-ab70-4000-d097-a8b650282ac2"
pickle_model.predict(df)
| notebooks/Model_rodent.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:base-data-science]
# language: python
# name: conda-env-base-data-science-py
# ---
# +
from xgboost import XGBClassifier
import anndata as an
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import sys
sys.path.append('../src')
from models.lib.lightning_train import DataModule
import xgboost as xgb
# +
module = DataModule(
datafiles=['../data/retina/retina_T.h5ad'],
labelfiles=['../data/retina/retina_labels_numeric.csv'],
class_label='class_label',
index_col='cell',
batch_size=16,
num_workers=0,
shuffle=True,
drop_last=False,
normalize=False,
)
module.setup()
# -
loader = module.trainloader
sample = next(iter(loader))
X = sample[0].numpy()
y = sample[1].numpy()
# +
from sklearn.metrics import accuracy_score
model = XGBClassifier()
train = module.trainloader
test = module.valloader
# Train on one minibatch to get started
sample = next(iter(loader))
X = sample[0].numpy()
y = sample[1].numpy()
model = model.fit(X, y)
# -
for i, (trainsample, valsample) in enumerate(zip(train, test)):
if i == 5:
break
X_train, y_train = trainsample
X_test, y_test = valsample
model = model.fit(X_train, y_train, xgb_model=model.get_booster())
y_pred = model.predict(X_test)
accuracy = accuracy_score(y_test, y_pred)
print(accuracy)
| notebooks/xgboost_model_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
# +
# line plot
themes=plt.style.available
print(themes)
# -
plt.style.use("seaborn")
x=np.array([1,2,3,4,5,6,7,8,9,10])
y1=2*x+1
y2=x**2
print(y1)
print(y2)
plt.plot(x,y1,color="orange",label="Apple",linestyle="dashed")
plt.plot(x,y2,color="green",label="banana",marker="o")
plt.xlabel("time")
plt.ylabel("frequency")
plt.title("time vs frequency")
plt.legend()
plt.show()
plt.plot(x,y2,'bo')
# scatter plot
plt.scatter(x,y1)
plt.show()
plt.scatter(x,y2)
plt.show()
plt.figure(figsize=(5,5))
plt.scatter(x,y1,color="orange",label="Apple",linestyle="dashed")
plt.scatter(x,y2,color="green",label="banana",marker="^")
plt.xlabel("time")
plt.ylabel("frequency")
plt.title("scatter plot :time vs frequency")
plt.legend()
plt.show()
# bar graph
xcoordinates=np.array([1,2,3])*2
plt.bar(xcoordinates-0.25,[10,20,45],width=0.5,label="previous year",tick_label=["gold","silver","diamond"],color="cyan")
plt.bar(xcoordinates+0.25,[20,15,25],width=0.5,label="current year",color="blue")
plt.title("Metal price comparison")
plt.xlabel("Metal")
plt.ylabel("price")
plt.ylim(0,90)
plt.legend()
plt.show()
# # pie chart
plt.style.use("seaborn")
subjects="English","maths","science","hindi","sst"
weightage=[10,40,20,20,10]
plt.pie(weightage,labels=subjects,explode=(0,0,0,0.1,0.1),autopct='%2.1f%%',shadow=True)
plt.show()
# ##
# # Histogram
xsn=np.random.randn(100)
sigma=8
u=70
x=np.round(xsn*sigma+u)
x1=np.round(xsn*4+45)
print(x)
# +
plt.style.use("seaborn")
plt.hist(x1,alpha=0.8)
plt.hist(x,alpha=0.8)
plt.xlabel("marks of student")
plt.ylabel("frequency of marks")
plt.show()
# -
# # Normal distribution
u=2
sigma=1
vals=u+sigma*np.random.randn(1000)
plt.hist(vals,50)
plt.show()
| MachineLearning/Data-visualisation/datavisualisation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Preprocessing of the data to replicate baseline.
import pandas as pd
import numpy as np
import os
#from tqdm import tqdm
# ### Set data paths
data_path = "/home/shivam/Documents/wikipedia-biography-dataset/wikipedia-biography-dataset/train"
data_files_paths = {
"table_content": os.path.join(data_path, "train.box"),
"nb_sentences" : os.path.join(data_path, "train.nb"),
"train_sentences": os.path.join(data_path, "train.sent")
}
# ### Read data and split into pairs(field and content)
print("Reading from the train.box file ...")
with open(data_files_paths["table_content"]) as t_file:
# read all the lines from the file:
table_contents = t_file.readlines()
table_contents = map(lambda x: x.strip().split('\t'), table_contents)
print(type(table_contents))
print("splitting the samples into field_names and content_words ...")
# convert this list of string pairs into list of lists of tuples
table_contents = map(lambda y: map(lambda x: tuple(x.split(":")), y), table_contents)
# ### Subsetting data
x = [list(data) for data in table_contents]
limited_data = x[:5]
#print(limited_data)
#len(x)
# ## Remove _1, _2 characters trailing in field names
# +
def remove_integer(array):
new_array = []
for item_tuple in array:
item_array =list(item_tuple)
if len(item_array[0].split("_")) > 1:
if item_array[0].split("_")[-1].isdigit():
item_array[0] ="_".join(item_array[0].split("_")[:-1])
new_array.append(item_array)
return (new_array)
edited_items = list(map(remove_integer, x))
# edited_items = [item for subitem in editem_items for item in subitem]
# list(map(remove_integer, limited_data))
# -
# Created a dictionary earlier, but that is of no use as of now
'''
dict_data = []
for x in edited_items:
new_item_dict ={}
for y in x:
if y[1].lower() != "<none>":
new_item_dict[y[0].lower()] = y[1].lower()
dict_data.append(new_item_dict)
print(dict_data)
'''
# ### Remove fields having None in their content
processed_data = []
for x in edited_items:
new_item_li =[]
for y in x:
if y[1].lower() != "<none>":
new_item_li.append(y)
processed_data.append(new_item_li)
#print(processed_data)
#print(new_item_li)
# #### Generate field names list
field_list = []
for li in processed_data:
for x in li:
field_list.append(x[0])
#print(field_list)
len(set(field_list)) #number of unique field names in dataset
# ### Count number of occurence of each field name
def count_field_names(lst):
elements = {}
for elem in lst:
if elem in elements.keys():
elements[elem] += 1
else:
elements[elem] = 1
return elements
count_dict = count_field_names(field_list)
# ### Remove field names occuring less than 100 times in dataset
final_list = []
for key, value in count_dict.items():
if value > 100:
final_list.append(key)
print(len(final_list)) #baseline vocubulary replicated
field_list[:10]
# ### Generate List of list of tuples as baseline vocubulary replica
pre_processed_li = []
for x in processed_data:
one_iter = []
for y,z in zip(x, field_list):
if y[0] == z:
one_iter.append((z ,y[1]))
pre_processed_li.append(one_iter)
len(pre_processed_li)
pre_processed_li[:5]
| TensorFlow_implementation/pre_processing_op.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="OxYEo003HLAI"
# # **Importing External Packages**
# + colab={"base_uri": "https://localhost:8080/"} id="HUD9ot2RGVb9" outputId="37c928f9-514a-4942-c903-937a1de7b116"
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import numpy as np
print("Libraries Imported.");
# + [markdown] id="-fTAUdfcHUDj"
# # **Reading Data Into The DataFrame**
# + colab={"base_uri": "https://localhost:8080/"} id="muDtI05ONyA7" outputId="8d051145-eebd-49d9-ad50-d3ec013e5248"
home_ownership = pd.read_excel('/content/Home Ownership Rate.xlsx')
final_dataset = pd.read_excel('/content/House Final.xlsx')
permit_df = pd.read_excel('/content/House Permit.xlsx')
house_price_index = pd.read_excel('/content/House Price Index.xlsx')
supply_ratio = pd.read_excel('/content/House Supply Ratio.xlsx')
house_vacancy = pd.read_excel('/content/House Vacancy.xlsx')
demand_df = pd.read_excel('/content/Housing Demand.xlsx')
usa_housing = pd.read_csv('/content/USA_Housing.csv')
new_houses_supply = pd.read_excel('/content/new houses supply.xlsx')
cred_index = pd.read_excel('/content/Home Cred Index.xlsx')
print("We are all set now!!!")
# + colab={"base_uri": "https://localhost:8080/", "height": 444} id="vONgcu0J5okZ" outputId="2f8651e1-91fb-4fca-d6c0-54b448fee951"
final_dataset
# + id="J4X3sEhuaJI4"
final_dataset.dropna(inplace= True)
# + colab={"base_uri": "https://localhost:8080/", "height": 444} id="Tfb3DPxQaTW_" outputId="569fd061-c6d7-4a10-87c3-2d5f4cb43d0e"
final_dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 320} id="HeZXcQ_Rabuo" outputId="240fe92e-5587-4cac-ea47-0a93d2f90cd7"
final_dataset.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="095RZqqzbVGM" outputId="c290af35-dbda-44ea-b208-e004ef720457"
final_dataset.info()
# + id="9lAGuG0adcfA"
final_dataset.to_csv('homes_final__.csv', index = False)
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="CAbSAy7Y-7_Y" outputId="24ea7c1c-853c-43c0-e23a-eb2b954a7f71"
cred_index
# + colab={"base_uri": "https://localhost:8080/", "height": 735} id="d9r1uInqmJUv" outputId="bfbd3409-0326-4e5f-9cea-d64523a1bade"
sns.pairplot(final_dataset);
# + [markdown] id="Y1Xd6fqWi26j"
# # **How Permits Influence Supply And Demand of Homes**
# + colab={"base_uri": "https://localhost:8080/", "height": 402} id="SQaBjEJUcDkL" outputId="66c6fb9b-59ff-4c0f-e611-b439db8ec291"
plt.figure(figsize=(12,6))
sns.scatterplot(x = final_dataset['PERMIT'], y= final_dataset['supply_ratio'], data = final_dataset)
plt.title('Relationship between Permit and Supply Ratio');
# + colab={"base_uri": "https://localhost:8080/", "height": 397} id="jcFhZixRmznC" outputId="ce48001b-5286-4eef-883f-73a0cd2ac61f"
plt.figure(figsize=(12,6))
sns.scatterplot(x = final_dataset['PERMIT'], y= final_dataset['new_houses'], data = final_dataset)
plt.title('Relationship between Permit and New Houses');
# + id="Ur9UG7yU05OP" colab={"base_uri": "https://localhost:8080/", "height": 401} outputId="d571c717-0999-4870-e7e5-6f7e13cd78fa"
plt.figure(figsize=(12,6))
sns.scatterplot(x = final_dataset['PERMIT'], y= final_dataset['new_sales'], data = final_dataset)
plt.title('Relationship between Permit and New Homes Sales');
# + colab={"base_uri": "https://localhost:8080/", "height": 401} id="nFrR_YgO180h" outputId="5472fa7b-53d6-49d4-cce8-0f7f5a7f744e"
plt.figure(figsize=(12,6))
sns.scatterplot(x = final_dataset['PERMIT'], y= final_dataset['house_price_index'], data = final_dataset)
plt.title('Relationship between Permit and House Price Index');
# + colab={"base_uri": "https://localhost:8080/", "height": 404} id="2FgmF2cF2L6O" outputId="56914b32-45c3-41f5-adbb-5b15c8fc636f"
plt.figure(figsize=(12,6))
sns.scatterplot(x = final_dataset['PERMIT'], y= final_dataset['placements_US'], data = final_dataset)
plt.title('Relationship between Permit and Supply');
# + colab={"base_uri": "https://localhost:8080/", "height": 404} id="_UUPUgyz7UjS" outputId="1a559d27-8033-4fff-f080-9cfcdf7bec3f"
plt.figure(figsize=(12,6))
sns.scatterplot(x = final_dataset['PERMIT'], y= final_dataset['home_ownership'], data = final_dataset)
plt.title('Relationship between Permit and Home Ownership');
# + id="WIq-ALL-7iU7"
final_dataset['DATE'] = pd.to_datetime(final_dataset.DATE)
# + id="rh0KtFcZ-czu"
final_dataset['year'] = pd.DatetimeIndex(final_dataset.DATE).year
# + colab={"base_uri": "https://localhost:8080/", "height": 444} id="cbMeOTQ4AvND" outputId="87c4ead9-729e-4098-bbc5-3ff6e18e1130"
final_dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="OqieEKHADQuB" outputId="6cbbc5eb-6dd6-4283-9150-c59164b00e57"
cred_index
# + id="4aDJ8mNLIeOn"
cred_index.rename(columns={"Year":"year"}, inplace= True)
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="jPCL3KrbIen3" outputId="c48df342-888b-4220-e543-3261e49855e3"
cred_index
# + [markdown] id="0VLZkx0X-09U"
# # **How Credit (Mortgage) Risk Has Influenced Supply and Demands Of Residential Homes**
# + colab={"base_uri": "https://localhost:8080/", "height": 404} id="_3d45ur4DevA" outputId="7b1d3752-2dea-488b-b25a-b32187838e19"
plt.figure(figsize=(12,6))
sns.barplot(x = cred_index['year'], y= cred_index['Total_risk'], data = cred_index)
plt.title('Cred Risk Barplot');
# + colab={"base_uri": "https://localhost:8080/", "height": 641} id="fiLdXKjlGMuV" outputId="afe44543-28aa-473e-82d6-898b24d1b318"
plt.figure(figsize= (25, 10))
plt.subplot(1,2,1)
plt.title('Credit Risk')
sns.barplot(x = cred_index['year'], y = cred_index['Total_risk'], data = cred_index )
plt.subplot(1,2,2)
plt.title('New Sales')
sns.barplot(x = final_dataset['year'], y = final_dataset['new_sales'], data = final_dataset);
# + id="XjQiFHzFJde1" colab={"base_uri": "https://localhost:8080/", "height": 317} outputId="cf32240b-e8e9-45c3-b6d8-d0caccbf527d"
plt.figure(figsize= (25, 10))
plt.subplot(1,2,1)
plt.title('Credit Risk')
sns.barplot(x = cred_index['year'], y = cred_index['Total_risk'], data = cred_index )
plt.subplot(1,2,2)
plt.title('House Price Index')
sns.barplot(x = final_dataset['year'], y = final_dataset['house_price_index'], data = final_dataset);
# + colab={"base_uri": "https://localhost:8080/", "height": 641} id="lqHENJCStyWI" outputId="5bfb88b3-94f1-4f15-aa89-733399a672c9"
plt.figure(figsize= (25, 10))
plt.subplot(1,2,1)
plt.title('Credit Risk')
sns.barplot(x = cred_index['year'], y = cred_index['Total_risk'], data = cred_index )
plt.subplot(1,2,2)
plt.title('Supply Ratio')
sns.barplot(x = final_dataset['year'], y = final_dataset['supply_ratio'], data = final_dataset);
# + colab={"base_uri": "https://localhost:8080/", "height": 412} id="a6xr6fzqdhas" outputId="18c6a33c-5262-4266-82c5-dee0872441d8"
with plt.xkcd():
plt.figure(figsize=(12,6))
sns.scatterplot(x = cred_index['Total_risk'], y= final_dataset['placements_US'], data = final_dataset)
plt.title('Relationship between Credit Risk and Vacant Home Placemets');
# + colab={"base_uri": "https://localhost:8080/", "height": 412} id="scgu4hAJeT3n" outputId="9a7158e6-4bac-48b5-a8ad-093b3bd73c5b"
with plt.xkcd():
plt.figure(figsize=(12,6))
sns.scatterplot(x = cred_index['Total_risk'], y= final_dataset['new_sales'], data = final_dataset)
plt.title('Relationship between Credit Risk and Demand');
# + colab={"base_uri": "https://localhost:8080/", "height": 412} id="Tg9iosMgeugs" outputId="1378a0ee-7fc8-47b7-b206-465cc3956860"
with plt.xkcd():
plt.figure(figsize=(12,6))
sns.scatterplot(x = cred_index['Total_risk'], y= final_dataset['supply_ratio'], data = final_dataset)
plt.title('Relationship between Credit Risk and Supply Ratio');
# + [markdown] id="XvEzsUd9Kv31"
# # **How Prices affect Supply and Demand**
# + colab={"base_uri": "https://localhost:8080/", "height": 621} id="YBYrt8FPK3T3" outputId="44ae2a4d-f39e-4494-8e7a-80a5a2681525"
plt.figure(figsize= (25, 10))
plt.subplot(1,2,1)
plt.title('House Price Index vs Demand')
sns.scatterplot(x = final_dataset['new_sales'], y = final_dataset['house_price_index'], data = final_dataset )
plt.xlabel('Demand')
plt.ylabel('House Price Index')
plt.subplot(1,2,2)
plt.title('House Price Index Vs Supply')
sns.scatterplot(x = final_dataset['placements_US'], y = final_dataset['house_price_index'], data = final_dataset)
plt.xlabel('Supply')
plt.ylabel('House Price Index');
# + colab={"base_uri": "https://localhost:8080/", "height": 621} id="vvfxVGiVQHAi" outputId="5fbb6423-0658-4ca1-bf9a-1f60a03aaf33"
plt.figure(figsize= (25, 10))
plt.subplot(1,2,1)
plt.title('Average Home Prices vs Demand')
sns.scatterplot(x = final_dataset['new_sales'], y = final_dataset['AvgPrice_US'], data = final_dataset )
plt.xlabel('Demand')
plt.ylabel('Average Home Prices')
plt.subplot(1,2,2)
plt.title('Average Home Prices VS Supply')
sns.scatterplot(x = final_dataset['placements_US'], y = final_dataset['AvgPrice_US'], data = final_dataset)
plt.xlabel('Supply')
plt.ylabel('Average Home Prices');
# + [markdown] id="aH_Zye90_ML_"
# # **Further EDA**
# + colab={"base_uri": "https://localhost:8080/", "height": 316} id="c-759XFquJ5I" outputId="5aae4b85-d7be-40f4-819b-3e83136dd4d1"
plt.figure(figsize= (25, 10))
plt.subplot(1,2,1)
plt.title('Vacant Homes Placements')
sns.barplot(x = final_dataset['year'], y = final_dataset['placements_US'], data = final_dataset);
plt.subplot(1,2,2)
plt.title('New Homes Sales')
sns.barplot(x = final_dataset['year'], y = final_dataset['new_sales'], data = final_dataset);
# + colab={"base_uri": "https://localhost:8080/", "height": 403} id="2zJz98pbvIPu" outputId="8f387547-5ead-4253-f5fa-32fd605f5449"
with plt.xkcd():
plt.figure(figsize=(12,6))
sns.scatterplot(x = final_dataset['placements_US'], y= final_dataset['new_sales'], data = final_dataset)
plt.title('Relationship between Vacant Homes Placements to New Sales');
# + id="pLGaI034vUWm" colab={"base_uri": "https://localhost:8080/", "height": 412} outputId="28b226d7-a84c-4535-b748-ae891eecbace"
with plt.xkcd():
plt.figure(figsize=(12,6))
sns.scatterplot(x = final_dataset['year'], y= final_dataset['AvgPrice_US'], data = final_dataset)
plt.title('Average House Prices in the US over the last 20+ years');
# + colab={"base_uri": "https://localhost:8080/", "height": 621} id="5kscDUiYb1dd" outputId="68bc7592-6f29-453d-97ec-32951848c0b2"
plt.figure(figsize= (25, 10))
plt.subplot(1,2,1)
plt.title('Supply In the last 20 years')
sns.barplot(x = final_dataset['year'], y = final_dataset['placements_US'], data = final_dataset);
# + [markdown] id="Ssx6MdOk_UOf"
# # **DATA INSIGHTS**
# + [markdown] id="KtTxpPpw_cfJ"
#
#
# * To view or download a fully detailed report on google docs kindly [Click Here](https://docs.google.com/document/d/1liRLWiRHOK5Czjnx_l-she4iBBAcYPeo0ohtVOVRij0/edit?usp=sharing)
# * For more factors that could influence prices of US. Homes visit my [Github Repository](https://github.com/ihechiluru/House-Price-Analysis-USA-) on House Prices in the US.
#
#
| Comprehensive_Analysis_US_Homes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:nlp_course]
# language: python
# name: conda-env-nlp_course-py
# ---
# ## Tokenization Basics
# Walkthrough for the blog:
# ### Loading in spaCy library and the target Language object
import spacy
from spacy import displacy
# Loading in the spaCy library
nlp = spacy.load('en_core_web_sm')
# Viewing the contents within our **nlp pipeline**
nlp.pipeline
# Creating a document object
doc = nlp(u"Here is our new fancy document. It's not very complex, but it will get the job done.")
# Viewing each token within the document object
for token in doc:
print(token.text)
# Viewing the length of the document object
len(doc)
# Now we will create a function that prints out each token's text, part of speech, and syntactic dependency.
def doc_breakdown(doc):
for token in doc:
print(f"Actual text: {token.text:{10}} "
f"Part of Speech: {token.pos_:{10}} "
f"Syntactic dependency: {token.dep_:{10}}")
doc_breakdown(doc)
# Document objects are technically lists, but they do not support item reassignment.
doc[0]
try:
doc[0] = 'Hear'
except TypeError as e:
print(e)
# If we are not sure what a tag means, we can use the spacy.explain() method to give a definition of the tag or label.
print(f"Part of Speech: \n{doc[0].pos_} = {spacy.explain(doc[0].pos_)}\n\n"
f"Syn. Dependency: \n{doc[0].dep_} = {spacy.explain(doc[0].dep_)}")
# We can also directly pass in a string of the tag or label we want to be explained.
spacy.explain('advmod')
# ### Understanding Named Entities in text
# Named Entity objects take tokens to the next level. If we check the contents of the nlp pipeline, we see that it contains an **'NER'** object. This object is a named entity recognizer. The nlp pipeline can recognize that certain words are organizational names, locations, monetary values, dates, etc. Named entities are accessible through the **.ents** property of a Doc object.
# Creating a new document
doc2 = nlp(u"Tesla Company will pay $750,000 and build a solar roof to settle dozens of \
air-quality violations at its Fremont factory.")
print(doc2)
# For each named entity found in doc2, we will print out the text of the named entity, the tag/label the pipeline predicts it to be, and an explanation of the tag/label.
for entity in doc2.ents:
print(f"Entity: {entity}\nLabel: {entity.label_}\nLabel Explanation: {spacy.explain(entity.label_)}\n")
# ### Using displacy With the Experimental Jupyter Parameter
# One final thing we will touch on before ending this blog is the **displacy module** within the spaCy library. **Displacy** is a built-in dependency visualizer that lets us check our model's predictions. We can pass in one or more Document objects and start a web server, export HTML files, or even view the visualization directly from a Jupyter Notebook. Since we are using a Jupyter Notebook for this blog, we will be viewing our visualizations directly from our notebook.
displacy.render(doc2, style='ent', jupyter=True)
# +
colors = {"MONEY": "lightgreen",
"CARDINAL": "linear-gradient(180deg, yellow, orange)"
}
options = {"colors": colors,
"ents": ["MONEY", "CARDINAL"]}
displacy.render(doc2, style='ent', jupyter=True, options=options)
# +
colors = {"ORG": "linear-gradient(90deg, #aa9cfc, #fc9ce7)",
"MONEY": "linear-gradient(45deg, lightgreen, white)",
"CARDINAL": "linear-gradient(180deg, yellow, orange)",
"GPE": "lightblue"}
options = {"ents": ["ORG", "MONEY", "CARDINAL", "GPE"], "colors": colors}
displacy.render(doc2, style='ent', jupyter=True, options=options)
# -
# ### The Dependency Style
doc3 = nlp(u"SpaCy Basics: The Importance of Tokens in Natural Language Processing")
displacy.render(doc3, style='dep', jupyter=True)
options = {
'bg': 'linear-gradient(180deg, orange, #FEE715FF)',
'color': 'black',
'font': 'Verdana'
}
displacy.render(doc3, style='dep', options=options, jupyter=True)
# ### Extra Stuff
# Create a Doc object with a unicode string (u-string)
doc4 = nlp(u"SpaCy is a library for advanced Natural Language Processing in Python \
and Cython. It's built on the very latest research, and was designed from day \
one to be used in real products. SpaCy comes with pretrained pipelines and currently \
supports tokenization and training for 60+ languages. It features state-of-the-art \
speed and neural network models for tagging, parsing, named entity recognition, \
text classification and more, multi-task learning with pretrained transformers \
like BERT, as well as a production-ready training system and easy model packaging, \
deployment and workflow management. SpaCy is commercial open-source software, released \
under the MIT license.")
# Understand that tokens are the basic building blocks of a doc object. Everything that helps us comprehend the meaning of text is derived from a token object and the relationship between tokens.
# spaCy is also able to detect and separate sentences in a Doc object.
for i, sentence in enumerate(doc4.sents):
print(f"{i+1}. {sentence}")
for chunk in doc4.noun_chunks:
print(chunk)
options = {"compact": True,
"bg": "#09a3d5",
"color": "white",
"distance": 250}
displacy.render(doc2, jupyter=True, options=options)
# ### Using Lemmatization on Tokens
doc5 = nlp(u"I love to hike, especially on the weekends. I went hiking yesterday with my hiker friends.")
for token in doc3:
print(f"{token.text:{10}} {token.pos_:{10}} {token.lemma:{20}} {token.lemma_:>{10}}")
# ### Coarse Versus Fine grained Tokens
# Using fine grained tokens let's us take advantage of things like present and past tense within the tokens.
doc5 = nlp(u"I love to hike, especially on the weekends. I went hiking yesterday with my hiker friends.")
for token in doc5:
print(f"{token.text:{10}} {token.pos_:{6}} {token.tag:<{25}} {token.tag_:{10}} {spacy.explain(token.tag_)}")
# Creating a dictionary to view the type of part-of-speech and count within doc4
doc4_dict = doc4.count_by(spacy.attrs.POS)
# Viewing coarse-grained part-of-speech
for k,v in doc4_dict.items():
print(f"{spacy.explain(doc4.vocab[k].text)}: {v:}")
# Viewing fine-grained tags
doc4_dict = doc4.count_by(spacy.attrs.TAG)
for k,v in doc4_dict.items():
print(f"{spacy.explain(doc4.vocab[k].text)}: {v:}")
| spacy_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.core.display import HTML
HTML("""
<style>
div.cell { /* Tunes the space between cells */
margin-top:1em;
margin-bottom:1em;
}
div.text_cell_render h1 { /* Main titles bigger, centered */
font-size: 2.2em;
line-height:1.4em;
text-align:center;
}
div.text_cell_render h2 { /* Parts names nearer from text */
margin-bottom: -0.4em;
}
div.text_cell_render { /* Customize text cells */
font-family: 'Times New Roman';
font-size:1.2em;
line-height:1.4em;
padding-left:3em;
padding-right:3em;
}
</style>
""")
# <h1 align="center"> All about Parts of Speech (POS) tags</h1>
#
# <h3 align="center"> Understanding of POS tags and build a POS tagger from scratch </h3>
# <img src="https://raw.githubusercontent.com/jalajthanaki/POS-tag-workshop/master/imgs/Image_1_1.gif"
# align="middle" alt="Image_1_1" data-canonical-src="" style="width:70%;height:70%">
# <h2 align="center"> There are main three sections of this workshop </h2>
# ```
#
# Section 1. Introduction to Parts of Speech
#
#
# Section 2. Generate Parts of Speech tags using various python libraries
#
#
# Section 3. Build our own statistical POS tagger form scratch
#
#
# ```
# <h1 align="center"> Introduction to Parts of Speech </h1>
#
# ```
# 1.1 What is Parts of Speech?
#
# 1.2 What is Parts of Speech tagging?
#
# 1.3 What is Part of Speech tagger?
#
# 1.4 What are the various types of the Part of Speech tags?
#
# 1.5 Which applications are using POS tagging?
#
# ```
# **What is Parts of Speech?**
# **Parts of Speech (POS):** POS helps us to get an idea __how a particular word or a phrase used in a given sentence in order to convey the logical meaning__ of the sentence.
#
# * It is important to know the __category of the words__ that helps us to form logical meaning of the sentence for any given language.
# <img src="https://raw.githubusercontent.com/jalajthanaki/POS-tag-workshop/master/imgs/Image_1_2.jpg"
# align="middle" alt="Image_1_2" data-canonical-src="" style="width:60%;height:60%">
# **What is Part of Speech tagging?**
# **Part of Speech tagging (POS tagging):** POS tagging is defined as the process of marking words in the corpus corresponding to their suitable/particular Parts of Speech.
#
#
# * The __set of tags__ is called the __tag set.__
#
#
# * Standard tag set is : __Penn Treebank__ (This is for English language.)
#
#
# * The POS tags of the word is __dependent__ on both __its definition and its context.__
#
#
# * POS tags of the words are dependent on their relationship with adjacent and related words in the given phrase,
# sentence and paragraph
#
#
# * It is also called grammatical tagging or word-category disambiguation.
#
#
#
#
#
#
# **Example**
#
# Sentence: **John likes the blue house at the end of the street.**
#
#
# | Words | Word Category |
# |---|---|
# | John | Noun |
# | likes | Verb |
# | the | Determiner |
# | blue | Adjective |
# | house | Noun |
# | at | Preposition |
# | the| Determiner |
# | end | Noun |
# | of | Preposition |
# | the | Determiner |
# | street | Noun |
# | . | . |
#
# __Example__
#
# * She saw a __bear.__ : Here, __bear__ is __"Noun"__
#
# * Your efforts will __bear__ fruit.: Here, __bear__ is __"Verb"__
#
# * Muje __khaana(1) khaana(2)__ hai. (Hindi Language) : Here, __khaana(1) is "Noun"__ and __khaana(2) is "Verb".__
# **What is Part of Speech tagger?**
# **Part of Speech tagger (POS tagger):** POS tagger is the tool that is used to assign POS tags for the given sentence or dataset/corpus
# **What are the various types of the Part of Speech tags?**
# <img src="https://raw.githubusercontent.com/jalajthanaki/POS-tag-workshop/master/imgs/Image_1_3.png"
# align="middle" alt="Image_1_3" data-canonical-src="" style="width:60%;height:60%">
# <img src="https://raw.githubusercontent.com/jalajthanaki/POS-tag-workshop/master/imgs/Image_1_4.png"
# align="middle" alt="Image_1_4" data-canonical-src="" style="width:40%;height:40%">
# <p style="text-align: center;"> You can see the list of tags from <a href ="https://github.com/jalajthanaki/NLPython/blob/master/ch5/POStagdemo/POS_tags.txt" target="_blank"> here </a></p>
#
#
# <p style="text-align: center;"> You can see the tags with example from <a href ="https://www.sketchengine.eu/penn-treebank-tagset/" target="_blank"> here </a></p>
#
#
# **Which applications are using POS tagging? **
# * Word Sense Disambiguation
#
#
# * Grammar correction system
#
#
# * Question-Answering system
#
#
# * Machine Translation
#
#
# * Sentiment Analysis
#
#
# * Detection of Multi word Expression
#
#
| POS-tag-workshop-master/Introduction_to_POS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#import pandas
import pandas as pd
#little tutorial for help
#https://medium.com/@ageitgey/quick-tip-the-easiest-way-to-grab-data-out-of-a-web-page-in-python-7153cecfca58
#scrap the first dataset and get the Hofstede cultural dimension data
df = pd.read_html("http://www.clearlycultural.com/geert-hofstede-cultural-dimensions/masculinity/", header=0)
#save first table on page as df1
df1 = (df[1])
#Remove column 0 from table df1 and save cleaned table to df2
df2 = df1[1:]
#print the head of the table df2
print (df2.head(10))
#save df2 table as hofstede.csv
df2.to_csv('hofstede.csv')
# +
#scrap the second dataset and get the gender gap wage (ggw) data from wikipedia
dfggw = pd.read_html("https://en.wikipedia.org/wiki/Gender_pay_gap", header=0)
#save desired table of the page as dfggw1
dfggw1 = (dfggw[1])
#Select the desired data from table dfggw1 and save cleaned table as dfggw2
dfggw2 = dfggw1[1:]
#print the head of the table dfggw2
print (dfggw2.head(10))
#save dfgwg2 table as gender_gap_wage.csv
dfggw2.to_csv('gender_gap_wage.csv')
# +
#opens dataset hofstede.csv
hos = pd.read_csv('hofstede.csv', index_col=0)
#print the head of hofstede.csv
print(hos.head(5))
#prints dataset hofstede.csv
ggw = pd.read_csv('gender_gap_wage.csv', index_col=0)
print (ggw.head(5))
#merges two datasets: gender gap wage and hofstede.csv on 'Country'
dataset = pd.merge(hos, ggw, on=['Country'])
print(dataset.head(10))
#Saves dataset as data.csv
dataset.to_csv('data.csv')
# -
# ls
| Hofstede+Cultural+Dimensions+Project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: myriam
# language: python
# name: myriam
# ---
import json
def create_config(num_classes, samples_per_class, target_samples_per_class, seed, model_name, num_features,
num_stages, total_epochs, save_results, connectivity_matrix_path):
data = {
"train_seed": seed, "val_seed": seed,
"load_into_memory": False,
# Loading data.
"image_height": 360,
"image_width": 1,
"image_channels": 1,
"num_dataprovider_workers": 2,
"dataset_name": "IBC",
"dataset_path": "IBC",
"indexes_of_folders_indicating_class": [-3, -2],
"sets_are_pre_split": True,
"num_of_gpus": 1,
"batch_size": 8,
"num_target_samples": target_samples_per_class,
"num_samples_per_class": samples_per_class,
"num_classes_per_set": num_classes,
# Meta-training hyperparameters.
## Inner loop.
"task_learning_rate": 0.01,
"number_of_training_steps_per_iter": 5,
"learnable_per_layer_per_step_inner_loop_learning_rate": True,
## Outer loop.
"meta_learning_rate": 0.005,
"total_epochs": total_epochs,
"min_learning_rate": 0.001,
"second_order": True,
"first_order_to_second_order_epoch": -1,
"use_multi_step_loss_optimization": True,
"multi_step_loss_num_epochs": 10,
# Meta-testing hyperparameters.
"evaluate_on_test_set_only": False,
"number_of_evaluation_steps_per_iter": 5,
"num_evaluation_tasks": 496,
# Architecture
"model_name": model_name, #'MLP', 'GNN' or 'Conv1d'
"num_features": num_features,
"num_stages": num_stages,
# Save results
"experiment_name":f"IBC_{num_classes}_way_{samples_per_class}_shot_{seed}_seed_{model_name}_model_{num_stages}_depth_{num_features}_features",
"max_models_to_save": 1,
"continue_from_epoch": "from_scratch",
"total_epochs_before_pause": 150,
"total_iter_per_epoch": 50,
"save_results": save_results,
"connectivity_matrix_path": connectivity_matrix_path,
}
return data
# +
### TO UPDATE ###
save_results = '/home/brain/Myriam/fMRI_transfer/git/fewshot_neuroimaging_classification/MAML_plus_plus/results'
connectivity_matrix_path = '/home/brain/Myriam/fMRI_transfer/git/fewshot_neuroimaging_classification/dataset/SC_avg56.mat'
# Examples of experimental settings.
num_classes = 5
target_samples_per_class = 15 ## Be careful: samples_per_class + target_samples_per_class <= 21.
seed = 0
total_epochs = 20
for model_name in ['GNN', 'MLP', 'Conv1d']:
for samples_per_class in [1, 5]:
# num_features can be in [64, 128, 256, 360, 512, 1024] for the 'MLP', 'GNN'. It represents the number of features.
# num_features can be in [64, 128, 256, 360, 512, 1024] for the 'Conv1d'. It represents the number of feature maps.
# in practice, you can use other numbers but you need to change the configuration of the files storing the results in utils.store_results.
for num_features in [64, 128, 256, 360, 512, 1024]:
for num_stages in [1, 2,]:
data = create_config(num_classes, samples_per_class, target_samples_per_class, seed, model_name, num_features, num_stages, total_epochs, save_results, connectivity_matrix_path)
with open(f"IBC_{num_classes}_way_{samples_per_class}_shot_{seed}_seed_{model_name}_model_{num_stages}_depth_{num_features}_features.json", 'w') as file:
json.dump(data, file)
| MAML_plus_plus/experiment_config/Create config file.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/XavierCarrera/Tutorial-Machine-Learning-Regresion-Lineal/blob/main/2_Regresion_Linear_Algebra.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="lpS9IhRJsmEM"
# # Introducción
#
# En esta ocasión, usaremos algebra lineal para entender que tanto se separa nuestros data points. Esta es una métrica muy importante para saber que tan bien se desempeña un modelo de regresión lineal.
#
# En este caso utilizaremos la librería **Scikit-Learn** (que sirvce para mahcine learning) con los siguientes modulos:
#
# * **linear_model**: con el cual entrenamos un modelo de machine learning de regresión lineal.
# * **train_test_split**: que nos sirve separar datos de entrenamiento y pruebas.
# * **metrics**: donde podemos encontrar formas de medir el desempeño de nuestro modelo.
# + id="_R4yrqxvqz3K"
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error, r2_score
# %matplotlib inline
plt.rcParams['figure.figsize'] = (20, 10)
plt.style.use('ggplot')
import warnings
warnings.filterwarnings('ignore')
# + [markdown] id="XdQcu1kudrOS"
# Seguimos usando el mismo dataset de pingüinos del notebook anterior.
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="G38GEea2r_N6" outputId="4c436a7c-e15a-434e-c71c-a2439faa58ba"
df = sns.load_dataset("penguins")
df
# + [markdown] id="6Kke9L_8eKgM"
# El dataset viene con datos nulos, que no nos sirven para entrenar un modelo de machine learning. Por ende, los contaremos y luego los eliminaremos.
# + colab={"base_uri": "https://localhost:8080/"} id="217LQBfZtsHg" outputId="6fc1da77-d96d-4de5-edf2-0eb3068d4e3d"
df.isnull().sum()
# + id="Pli5XaQEt1Ny"
df.dropna(inplace=True)
# + [markdown] id="G_7z0jjeea-0"
# Para este ejercicio, crearemos un gráfico de correlación linear como el que hicimos anteriormente. En este caso suponemos que existe una relación entre la masa corporal de los pingüinos y el tamaño de sus alas.
# + colab={"base_uri": "https://localhost:8080/", "height": 625} id="K6jWtgpTsE3q" outputId="7af6876b-1659-43eb-f318-211aa5f8571c"
sns.regplot(x=df["body_mass_g"], y=df["flipper_length_mm"])
# + [markdown] id="u83QqoA5sqF8"
# # Entrenamiento del Modelo
#
# A partir de la correlación linear anterior, entrenaremos un modelo simple de regresión linear.
#
# Lo primero es crear dos vectores. En este caso usamos la masa corporal como nuestro vector X (predictor) y el tamaño de las alas como el vector y (valor por predecirse).
#
# La función *train_test_split* es fundamental en machine learning, ya que nos divide nuestro data sets en datos de entrenamiento y prueba. Los datos de prueba nos servirán para medir el desempeño de nuestro modelo.
#
# Las funciones *values* y *reshape* nos permiten crear un arreglo artificial en 2D que nos permitirá entrenar nuestro modelo.
#
# Para entrenar al modelo, solo tenemos que invocar a *Linear Regression* y luego entrenar al modelo con la función *fit*.
# + id="geqh7kcjzAVN"
X = df["body_mass_g"].values.reshape(-1,1)
y = df["flipper_length_mm"]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.20, random_state = 42)
# + colab={"base_uri": "https://localhost:8080/"} id="KMrZah8Tzcd8" outputId="80b66d50-14fd-456c-df70-54ac3d94fc4a"
reg = LinearRegression()
reg.fit(X_train, y_train)
# + [markdown] id="jCdwCPWd0TtP"
# # Suma de Errores Cuadráticos
#
# En este modelo tenemos ya un modelo entrenado y listo para ser medido.
#
# *Scikit Learn* no tiene un modulo para calcular la summa de errores cuadráticos por lo que lo haremos manualemente.
#
# Lo que tenemos es una vector llamado *y_test* (datos reales) que utiliza a *X_test* para generar una predicción. Lo que hacemos con una sola línea de código es sumar todos los vectores al cuadrado.
# + colab={"base_uri": "https://localhost:8080/"} id="ZhEYwh6h0R6o" outputId="2dd8ee9b-0bbb-4b27-f6b6-6b3f9815d5c2"
sec = np.sum((y_test - reg.predict(X_test))**2)
sec
# + [markdown] id="znhuZj_P1MDh"
# # Otras Métricas de Desempeño
#
# En este caso usamos dos métricas adicionales que vienen preconstruidas en *Scikit Learn*.
#
# Por un lado tenemos al promedio de los errores cuadráticos (mean_squared_error). Como su nombre lo indica, hace un promedio de los errores cuadráticos en vez de sumarlos.
#
# Una métrica que se usa popularmente en regresión linear son los mínimos cuadrados ordinarios (r²). Este genera un rango de 0 a 1 para medir la varianza capturada por el modelo.
#
# + colab={"base_uri": "https://localhost:8080/"} id="XfEhnt6QNRJt" outputId="3816b7a2-ea87-49f1-8988-72ef08868c9e"
y_test_predict = reg.predict(X_test)
rmse = mean_squared_error(y_test, y_test_predict)
r2 = r2_score(y_test, y_test_predict)
print(rmse)
print(r2)
| 2_Regresion_Linear_Algebra.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.insert(1, '../pyKinectAzure/')
import numpy as np
from pyKinectAzure import pyKinectAzure, _k4a
import cv2
import open3d as o3d
from open3d import geometry, visualization
#modulePath = r'/usr/lib/x86_64-linux-gnu/libk4a.so'
modulePath = 'C:\\Program Files\\Azure Kinect SDK v1.4.1\\sdk\\windows-desktop\\amd64\\release\\bin\\k4a.dll'
# -
pyK4A = pyKinectAzure(modulePath)
pyK4A.device_open()
device_config = pyK4A.config
device_config.color_resolution = _k4a.K4A_COLOR_RESOLUTION_1080P
device_config.depth_mode = _k4a.K4A_DEPTH_MODE_WFOV_2X2BINNED
#print(device_config)
# +
pyK4A = pyKinectAzure(modulePath)
pyK4A.device_open()
device_config = pyK4A.config
device_config.color_resolution = _k4a.K4A_COLOR_RESOLUTION_1080P
device_config.depth_mode = _k4a.K4A_DEPTH_MODE_WFOV_2X2BINNED
#print(device_config)
pyK4A.device_start_cameras(device_config)
k = 0
while (cv2.waitKey(1) & 0xFF != ord('q')):
# Get capture
pyK4A.device_get_capture()
# Get the color image from the capture
color_image_handle = pyK4A.capture_get_color_image()
# Check the image has been read correctly
if color_image_handle:
# Read and convert the image data to numpy array:
color_image = pyK4A.image_convert_to_numpy(color_image_handle)
# Plot the image
cv2.namedWindow('Color Image',cv2.WINDOW_NORMAL)
cv2.imshow("Color Image",color_image)
k = cv2.waitKey(10)
# Release the image
pyK4A.image_release(color_image_handle)
pyK4A.capture_release()
if k==27 or k==113: # Esc key to stop
break
pyK4A.device_stop_cameras()
pyK4A.device_close()
# -
pyK4A.device_stop_cameras()
pyK4A.device_close()
| examples/depthToNumpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Time series example
#
# *This worksheet was originally designed by [<NAME>](https://www.math.ucla.edu/~egeo/) (Department of Mathematics, UCLA). It has been subsequently revised by later TAs and instructors.*
#
# Warning: the model we get is pretty bad! We used [this dataset](https://www.kaggle.com/felsal/ibovespa-stocks).
import pandas as pd
import numpy as np
import tensorflow as tf
df = pd.read_csv('b3_stocks_1994_2020.csv')
df1 = df.pivot(index='ticker', columns='datetime', values='close')
df2 = df1.loc[:,'2020-12-1':]
df3 = df2.dropna()
df3.head()
x = df3.drop(columns=['2020-12-30'])
y = df3['2020-12-30']
# +
from sklearn.model_selection import train_test_split
x_trv, x_test, y_trv, y_test = train_test_split(x, y, random_state=209)
x_train, x_val, y_train, y_val = train_test_split(x_trv, y_trv)
display(x_train.head(1))
x_train.shape
# +
import tensorflow.keras as keras
model = keras.models.Sequential([
keras.layers.Reshape((x.shape[1], 1), input_shape=(x.shape[1],)),
keras.layers.Conv1D(10, 10, activation = 'sigmoid'),
keras.layers.Flatten(),
keras.layers.Dense(units = 30, activation = 'sigmoid'),
keras.layers.Dense(units = 1),
])
model.summary()
# +
model.compile(optimizer='adam',
loss='mse')
history = model.fit(x_train,
y_train,
epochs=5,
validation_data=(x_val, y_val))
# +
train_mse = model.evaluate(x_train, y_train)
train_var = y_train.var()
val_mse = model.evaluate(x_val, y_val)
val_var = y_val.var()
print(f'Train RMSE: {np.sqrt(train_mse)}')
print(f'Train r^2: {(train_var - train_mse)/train_var}')
print(f'Validation RMSE: {np.sqrt(val_mse)}')
print(f'Validation r^2: {(val_var - val_mse)/val_var}')
| discussion/time_series_ex.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Prepare the dataset
# +
import pandas as pd
df=pd.read_csv('AB_test_data.csv')
df
# -
a=df.groupby('Variant')['purchase_TF'].value_counts()
a_df = pd.DataFrame(a)
a_df
a_df.index
# # 1. Conduct an AB Test
A_False=a_df.loc[('A',False), 'purchase_TF']
print(A_False)
A_True=a_df.loc[('A',True), 'purchase_TF']
print(A_True)
B_True=a_df.loc[('B',True), 'purchase_TF']
print(B_True)
B_False=a_df.loc[('B',False), 'purchase_TF']
print(B_False)
p=A_True/(A_False+A_True)
p
p_hat=B_True/(B_True+B_False)
p_hat
import math
z=(p_hat-p)/math.sqrt(p_hat*(1-p_hat)/5000)
z
if math.fabs(z) >=1.64:
print('z score is %f. Reject the null hypotheis and conclude that Alternative B improved conversion rates over alternative A.' %(z))
else:
print('z score is %f. Fail to reject the null hypothesis and conclude that Alternative B did not improve conversion rates over alternative A.'%(z))
# # 2. Calculate the optimal sample size
#
# Optimal sample size balances committing Type I and Type 2 error.
# For proportions, assuming large enough data to use normal approximation to binomial distribution, variance = p(1-p) such that: 
#
# where p_bar is the average of the sample
# proportions.
#
# According to the calculation above, p0=0.149616, p1=0.1766, by searching the t-distribution table, t(0.025)=1.96, t(0.2)=1.28
p0 = 0.1507
p1 = 0.149616
p2 = 0.1766
p_average=0.1631
delta = p2-p1
import scipy.stats as st
t_0025=st.norm.ppf(0.975)
t_0025
t_02=st.norm.ppf(0.8)
t_02
a = t_0025 * math.sqrt(2*p_average*(1-p_average))
b = t_02 * math.sqrt(p1*(1-p1)+p2*(1-p2))
delta = p2-p1
sample_size = ((a+b) * (a+b)) / ((delta)*(delta))
sample_size
# Therefore, the optimal sample size is 2942.
#
# # Conduct the test 10 times
# +
import pandas as pd
sample1 = df.sample(n=2942)
sample2 = df.sample(n=2942)
sample3 = df.sample(n=2942)
sample4 = df.sample(n=2942)
sample5 = df.sample(n=2942)
sample6 = df.sample(n=2942)
sample7 = df.sample(n=2942)
sample8 = df.sample(n=2942)
sample9 = df.sample(n=2942)
sample10 = df.sample(n=2942)
sample1.head()
sample2.head()
sample3.head()
sample4.head()
sample5.head()
sample6.head()
sample7.head()
sample8.head()
sample9.head()
sample10.head()
# -
list = [sample1,sample2,sample3,sample4,sample5,sample6,sample7,sample8,sample9,sample10]
i = 0
while i < 10:
df = list[i]
print(df)
a=df.groupby('Variant')['purchase_TF'].value_counts()
a_df = pd.DataFrame(a)
A_False=a_df.loc[('A',False), 'purchase_TF']
A_True=a_df.loc[('A',True), 'purchase_TF']
B_True=a_df.loc[('B',True), 'purchase_TF']
B_False=a_df.loc[('B',False), 'purchase_TF']
p=A_True/(A_False+A_True)
p_hat=B_True/(B_True+B_False)
z=(p_hat-p)/math.sqrt(p_hat*(1-p_hat)/5000)
if math.fabs(z) >=1.64:
print('z score is %f.Reject the null hypotheis and conclude that Alternative B improved conversion rates over alternative A.' %(z))
else:
print('z score is %f. Fail to reject the null hypothesis and conclude that Alternative B did not improve conversion rates over alternative A.'%(z))
i += 1
# # results:
# the optimal sample size is 2942 and in the 10 samples generated, only 3 samples conclude that Alternative B did not improve conversion rates over alternative A; 7 samples conclude that Alternative B improved conversion rates over alternative A.
# # 3 Sequential Testing
alpha = 0.05
ln_A=math.log(1/alpha)
ln_A
beta = 0.2
ln_B=math.log(beta)
ln_B
PA=p
PB=p_hat
from random import sample
ln_0_xi = math.log((1-PA)/(1-PB))
ln_1_xi = math.log(PA/PB)
def SPRT():
ln_lamba = 0
i=0
B_sample = sample(list(df['purchase_TF'].values),2942)
for record in B_sample:
if ln_lamba<ln_A and ln_lamba>ln_B:
if record==0:
ln_lamba=ln_lamba+ln_0_xi
else:
ln_lamba=ln_lamba+ln_1_xi
i+=1
elif ln_lamba>=ln_A:
return(i,'Reject H0')
break
else:
return(i,'Fail to reject H0')
break
result_3 = [SPRT() for i in range(10)]
result_3
total = 0
for item in result_3:
total += item[0]
print('The average number of iterations required to stop the test is:',total/10)
# Based on the optimal sample size in question 2, we conduct the sequential test for 10 times. In all of the SPRT tests, we stop the test prior to using full samples, about 587.2 iterations to stop the test on average.
| Marketing Analytics HW1_Final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="I08sFJYCxR0Z"
# 
# + [markdown] id="FwJ-P56kq6FU"
# [](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/Certification_Trainings/Healthcare/4.2.Clinical_Deidentification_in_Spanish.ipynb)
# + [markdown] id="Z7X1prqVxdB2"
# # Clinical Deidentification in Spanish
#
# **Protected Health Information**:
#
# Individual’s past, present, or future physical or mental health or condition
# provision of health care to the individual
# past, present, or future payment for the health care
# Protected health information includes many common identifiers (e.g., name, address, birth date, Social Security Number) when they can be associated with the health information.
# + id="RWh6i1PtvE77"
import json
import os
from google.colab import files
if 'spark_jsl.json' not in os.listdir():
license_keys = files.upload()
os.rename(list(license_keys.keys())[0], 'spark_jsl.json')
with open('spark_jsl.json') as f:
license_keys = json.load(f)
# Defining license key-value pairs as local variables
locals().update(license_keys)
os.environ.update(license_keys)
# + id="MfOGbhC2wTyp" colab={"base_uri": "https://localhost:8080/"} outputId="7a5ab257-57e7-4ff3-ed39-e7f6ce798dec"
# Installing pyspark and spark-nlp
# ! pip install --upgrade -q pyspark==3.1.2
# Installing Spark NLP Healthcare
# ! pip install --upgrade -q spark-nlp-jsl==$JSL_VERSION --extra-index-url https://pypi.johnsnowlabs.com/$SECRET
# + id="nou6cgDm35Vq" colab={"base_uri": "https://localhost:8080/", "height": 222} outputId="5780f74a-b743-4e61-dbad-4470d7da40d8"
from pyspark.ml import Pipeline, PipelineModel
from pyspark.sql import functions as F
from pyspark.sql import SparkSession
from sparknlp.base import *
from sparknlp.annotator import *
from sparknlp.pretrained import ResourceDownloader
from sparknlp.util import *
from sparknlp_jsl.annotator import *
import sys
import os
import json
import pandas as pd
import string
import numpy as np
import sparknlp
import sparknlp_jsl
params = {"spark.driver.memory":"16G",
"spark.kryoserializer.buffer.max":"2000M",
"spark.driver.maxResultSize":"2000M"}
spark = sparknlp_jsl.start(SECRET, params=params)
spark
# + colab={"base_uri": "https://localhost:8080/"} id="fwMiy7-qmUF-" outputId="15581e56-562a-4369-989e-4c54527cafc7"
print ("Spark NLP Version :", sparknlp.version())
print ("Spark NLP_JSL Version :", sparknlp_jsl.version())
# + [markdown] id="VAyEoiHVuhbp"
# # 1. Spanish NER Deidentification Models
# We have eight different models you can use:
# * `ner_deid_generic`, detects 7 entities, uses SciWiki 300d embeddings.
# * `ner_deid_generic_roberta`, same as previous, but uses Roberta Clinical Embeddings.
# * `ner_deid_generic_augmented`, detects 8 entities (now includes 'SEX' entity), uses SciWiki 300d embeddings and has been trained with more data
# * `ner_deid_generic_roberta_augmented`, same as previous, but uses Roberta Clinical Embeddings.
# * `ner_deid_subentity`, detects 13 entities, uses SciWiki 300d embeddings.
# * `ner_deid_subentity_roberta`, same as previous, but uses Roberta Clinical Embeddings.
# * `ner_deid_subentity_augmented`, detects 17 entities, uses SciWiki 300d embeddings and has been trained with more data.
# * `ner_deid_subentity_roberta_augmented`, same as previous, but uses Roberta Clinical Embeddings.
#
# Since `augmented` models improve their results compared to the non augmented ones, we are going to show case them in this notebook
# + [markdown] id="V1eksdJQoF4e"
# ### Creating pipeline for Sciwiki 300d-based augmented model
# + colab={"base_uri": "https://localhost:8080/"} id="snKDCdXwoNy4" outputId="e140635b-3e36-43dc-adf3-385753711cd5"
# Annotator that transforms a text column from dataframe into an Annotation ready for NLP
documentAssembler = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
sentencerDL = SentenceDetectorDLModel.pretrained("sentence_detector_dl", "xx") \
.setInputCols(["document"])\
.setOutputCol("sentence")
# Tokenizer splits words in a relevant format for NLP
tokenizer = Tokenizer()\
.setInputCols(["sentence"])\
.setOutputCol("token")
word_embeddings = WordEmbeddingsModel.pretrained("embeddings_sciwiki_300d","es","clinical/models")\
.setInputCols(["document","token"])\
.setOutputCol("embeddings")
# + [markdown] id="3ZuJrSX6tT9m"
# ## 1.1. NER Deid Generic (Augmented)
#
# **`ner_deid_generic_augmented`** extracts:
# - Name
# - Profession
# - Age
# - Date
# - Contact (Telephone numbers, FAX numbers, Email addresses)
# - Location (Address, City, Postal code, Hospital Name, Employment information)
# - Id (Social Security numbers, Medical record numbers, Internet protocol addresses)
# - Sex
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="C-qUIvqpsjV0" outputId="f513a5b9-ba83-4cce-82f3-1d4b4050f6be"
ner_generic = MedicalNerModel.pretrained("ner_deid_generic_augmented", "es", "clinical/models")\
.setInputCols(["sentence","token","embeddings"])\
.setOutputCol("ner_deid_generic")
ner_converter_generic = NerConverter()\
.setInputCols(["sentence","token","ner_deid_generic"])\
.setOutputCol("ner_chunk_generic")
# + colab={"base_uri": "https://localhost:8080/"} id="j7Xy9E_hxN5y" outputId="2994165a-1894-431b-f2a3-ea62ff33b570"
ner_generic.getClasses()
# + [markdown] id="3NjkLF70tYdZ"
# ## 1.2. NER Deid Subentity
#
# **`ner_deid_subentity`** extracts:
#
# - Patient
# - Doctor
# - Hospital
# - Date
# - Organization
# - City
# - Street
# - User Name
# - Profession
# - Phone
# - Country
# - Age
# - Sex
# - Email
# - ZIP
# - ID
# - Medical Record
# + colab={"base_uri": "https://localhost:8080/"} id="C8n-h6D9tJXx" outputId="5593e6d4-fe86-47d7-fa5f-9fde3586de94"
ner_subentity = MedicalNerModel.pretrained("ner_deid_subentity_augmented", "es", "clinical/models")\
.setInputCols(["sentence","token","embeddings"])\
.setOutputCol("ner_deid_subentity")
ner_converter_subentity = NerConverter()\
.setInputCols(["sentence", "token", "ner_deid_subentity"])\
.setOutputCol("ner_chunk_subentity")
# + colab={"base_uri": "https://localhost:8080/"} id="Oda9sjHDxRyi" outputId="2d4d9cb8-b12d-40ae-fbb0-a8cb1889da89"
ner_subentity.getClasses()
# + [markdown] id="Eqg29dPuvl5f"
# ## 1.4. Pipeline
# + id="P0HZXLF6ueWi"
nlpPipeline = Pipeline(stages=[
documentAssembler,
sentencerDL,
tokenizer,
word_embeddings,
ner_generic,
ner_converter_generic,
ner_subentity,
ner_converter_subentity,
])
empty_data = spark.createDataFrame([[""]]).toDF("text")
model = nlpPipeline.fit(empty_data)
# + id="fG4Vc36EwhFk"
text = "<NAME>, un varón de 35 años de edad, de profesión auxiliar de enfermería y nacido en Cadiz, España. Aún no estaba vacunado, se infectó con Covid-19 el dia 14/03/2022 y tuvo que ir al Hospital. Fue tratado con anticuerpos monoclonales en la Clinica San Carlos."
text_df = spark.createDataFrame([[text]]).toDF("text")
result = model.transform(text_df)
# + [markdown] id="y-CMPYTHz-L2"
# ### Results for `ner_subentity`
# + colab={"base_uri": "https://localhost:8080/"} id="vEtBcyIjzLA3" outputId="5980b321-894c-4223-d192-d1b81fc8d796"
result.select(F.explode(F.arrays_zip('ner_chunk_subentity.result', 'ner_chunk_subentity.metadata')).alias("cols")) \
.select(F.expr("cols['0']").alias("chunk"),
F.expr("cols['1']['entity']").alias("ner_label")).show(truncate=False)
# + [markdown] id="MNUKLQMi0GjT"
# ### Results for `ner_generic`
# + colab={"base_uri": "https://localhost:8080/"} id="K2wDmdiFzwDb" outputId="de7ca06c-6a96-45b4-c59b-280f30439e8b"
result.select(F.explode(F.arrays_zip('ner_chunk_generic.result', 'ner_chunk_generic.metadata')).alias("cols")) \
.select(F.expr("cols['0']").alias("chunk"),
F.expr("cols['1']['entity']").alias("ner_label")).show(truncate=False)
# + [markdown] id="u-OTUyBK6yrt"
# ## Deidentification
# + id="pbJisU_u7Kpl"
# Downloading faker entity list.
# ! wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Healthcare/data/obfuscate_es.txt
# + id="CBo2T-sZ64IJ"
deid_masked_entity = DeIdentification()\
.setInputCols(["sentence", "token", "ner_chunk_subentity"])\
.setOutputCol("masked_with_entity")\
.setMode("mask")\
.setMaskingPolicy("entity_labels")
deid_masked_char = DeIdentification()\
.setInputCols(["sentence", "token", "ner_chunk_subentity"])\
.setOutputCol("masked_with_chars")\
.setMode("mask")\
.setMaskingPolicy("same_length_chars")
deid_masked_fixed_char = DeIdentification()\
.setInputCols(["sentence", "token", "ner_chunk_subentity"])\
.setOutputCol("masked_fixed_length_chars")\
.setMode("mask")\
.setMaskingPolicy("fixed_length_chars")\
.setFixedMaskLength(4)
deid_obfuscated = DeIdentification()\
.setInputCols(["sentence", "token", "ner_chunk_subentity"]) \
.setOutputCol("obfuscated") \
.setMode("obfuscate")\
.setObfuscateDate(True)\
.setObfuscateRefSource('faker')\
.setObfuscateRefFile('obfuscate_es.txt')\
.setObfuscateRefSource("file")\
# + id="h9pmXn0f75ST"
nlpPipeline = Pipeline(stages=[
documentAssembler,
sentencerDL,
tokenizer,
word_embeddings,
ner_subentity,
ner_converter_subentity,
deid_masked_entity,
deid_masked_char,
deid_masked_fixed_char,
deid_obfuscated
])
empty_data = spark.createDataFrame([[""]]).toDF("text")
model = nlpPipeline.fit(empty_data)
# + id="oVOL3bwr8J18"
deid_lp = LightPipeline(model)
# + id="Maco1EiD8TK4"
text = "<NAME>, un varón de 35 años de edad, de profesión auxiliar de enfermería y nacido en Cadiz, España. Aún no estaba vacunado, se infectó con Covid-19 el dia 14/03/2022 y tuvo que ir al Hospital. Fue tratado con anticuerpos monoclonales en la Clinica San Carlos."
# + colab={"base_uri": "https://localhost:8080/"} id="CXEmE1i78PX4" outputId="1f0855e1-4724-4b4e-e981-80ed3be1440f"
result = deid_lp.annotate(text)
print("\n".join(result['masked_with_entity']))
print("\n")
print("\n".join(result['masked_with_chars']))
print("\n")
print("\n".join(result['masked_fixed_length_chars']))
print("\n")
print("\n".join(result['obfuscated']))
# + [markdown] id="RRuq9NeemSaf"
# # 2. Pretrained Spanish Deidentification Pipeline
#
# - We developed a clinical deidentification pretrained pipeline that can be used to deidentify PHI information from German medical texts. The PHI information will be masked and obfuscated in the resulting text.
# - The pipeline can mask and obfuscate:
# - Patient
# - Doctor
# - Hospital
# - Date
# - Organization
# - City
# - Street
# - Country
# - User name
# - Profession
# - Phone
# - Age
# - Contact
# - ID
# - Phone
# - ZIP
# - Account
# - SSN
# - Driver's License Number
# - Plate Number
# - Sex
# + colab={"base_uri": "https://localhost:8080/"} id="_vBWjdVTFGHD" outputId="867ca617-bebb-4852-9c36-43103b9d2858"
from sparknlp.pretrained import PretrainedPipeline
deid_pipeline = PretrainedPipeline("clinical_deidentification_augmented", "es", "clinical/models")
# + colab={"base_uri": "https://localhost:8080/"} id="Esy1Yis8Hn1X" outputId="6fa4769c-48f6-4463-bdae-0cb07b7b4970"
text = """Datos del paciente.
Nombre: Ernesto.
Apellidos: <NAME>.
NHC: 368503.
NASS: 26 63514095.
Domicilio: <NAME> 90.
Localidad/ Provincia: Madrid.
CP: 28016.
Datos asistenciales.
Fecha de nacimiento: 03/03/1946.
País: España.
Edad: 70 años Sexo: H.
Fecha de Ingreso: 12/12/2016.
Médico: <NAME> NºCol: 28 28 70973.
Informe clínico del paciente: Paciente de 70 años de edad, minero jubilado, sin alergias medicamentosas conocidas, que presenta como antecedentes personales: accidente laboral antiguo con fracturas vertebrales y costales; intervenido de enfermedad de Dupuytren en mano derecha y by-pass iliofemoral izquierdo; Diabetes Mellitus tipo II, hipercolesterolemia e hiperuricemia; enolismo activo, fumador de 20 cigarrillos / día.
Es derivado desde Atención Primaria por presentar hematuria macroscópica postmiccional en una ocasión y microhematuria persistente posteriormente, con micciones normales.
En la exploración física presenta un buen estado general, con abdomen y genitales normales; tacto rectal compatible con adenoma de próstata grado I/IV.
En la analítica de orina destaca la existencia de 4 hematíes/ campo y 0-5 leucocitos/campo; resto de sedimento normal.
Hemograma normal; en la bioquímica destaca una glucemia de 169 mg/dl y triglicéridos de 456 mg/dl; función hepática y renal normal. PSA de 1.16 ng/ml.
Las citologías de orina son repetidamente sospechosas de malignidad.
En la placa simple de abdomen se valoran cambios degenerativos en columna lumbar y calcificaciones vasculares en ambos hipocondrios y en pelvis.
La ecografía urológica pone de manifiesto la existencia de quistes corticales simples en riñón derecho, vejiga sin alteraciones con buena capacidad y próstata con un peso de 30 g.
En la UIV se observa normofuncionalismo renal bilateral, calcificaciones sobre silueta renal derecha y uréteres arrosariados con imágenes de adición en el tercio superior de ambos uréteres, en relación a pseudodiverticulosis ureteral. El cistograma demuestra una vejiga con buena capacidad, pero paredes trabeculadas en relación a vejiga de esfuerzo. La TC abdominal es normal.
La cistoscopia descubre la existencia de pequeñas tumoraciones vesicales, realizándose resección transuretral con el resultado anatomopatológico de carcinoma urotelial superficial de vejiga.
Remitido por: <NAME> c/ del Abedul 5-7, 2º dcha 28036 Madrid, España E-mail: <EMAIL>.
"""
result = deid_pipeline.annotate(text)
print("\n".join(result['masked_with_chars']))
print("\n")
print("\n".join(result['masked']))
print("\n")
print("\n".join(result['masked_fixed_length_chars']))
print("\n")
print("\n".join(result['obfuscated']))
# + [markdown] id="cshjoPRXVTkP"
# # Other NER versions: Using Roberta Clinical Embeddings based NER
# You can use also Roberta Clinical Embeddings and `_roberta` , instead of Sciwi for NER models (not for Pretrained Pipeline, that comes only with `Sciwi`). This is an example of how to use the
# + colab={"base_uri": "https://localhost:8080/"} id="jubtKtrOVQoX" outputId="d4376122-cbd0-425f-8351-d7600b883726"
documentAssembler = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
sentenceDetector = SentenceDetectorDLModel.pretrained("sentence_detector_dl","xx")\
.setInputCols(["document"])\
.setOutputCol("sentence")
tokenizer = Tokenizer()\
.setInputCols(["sentence"])\
.setOutputCol("token")
roberta_embeddings = RoBertaEmbeddings.pretrained("roberta_base_biomedical", "es")\
.setInputCols(["sentence", "token"])\
.setOutputCol("embeddings")
clinical_ner = MedicalNerModel.pretrained("ner_deid_subentity_roberta_augmented", "es", "clinical/models")\
.setInputCols(["sentence","token","embeddings"])\
.setOutputCol("ner")
ner_converter = NerConverter()\
.setInputCols(["sentence","token","ner"])\
.setOutputCol("ner_chunk")
nlpPipeline = Pipeline(stages=[
documentAssembler,
sentenceDetector,
tokenizer,
roberta_embeddings,
clinical_ner,
ner_converter])
empty_data = spark.createDataFrame([[""]]).toDF("text")
model = nlpPipeline.fit(empty_data)
# + id="3G5v1GOxWUlJ"
text = ['''
<NAME>, varón de de 35 años de edad, de profesión auxiliar de enfermería y nacido en Cadiz, España. Aún no estaba vacunado, se infectó con Covid-19 el dia 14 de Marzo y tuvo que ir al Hospital
Fue tratado con anticuerpos monoclonales en la Clinica San Carlos.
''']
# + id="C0YoiNe4W-En"
result = model.transform(spark.createDataFrame([text]).toDF("text"))
# + colab={"base_uri": "https://localhost:8080/"} id="2arYlCoQWrco" outputId="1270abce-a6d8-4026-b365-e67b8c3b3cfd"
result.select(F.explode(F.arrays_zip('ner_chunk.result', 'ner_chunk.metadata')).alias("cols")) \
.select(F.expr("cols['0']").alias("chunk"),
F.expr("cols['1']['entity']").alias("ner_label")).show(truncate=False)
# + [markdown] id="91h16KhvXNT_"
# # About non-augmented models
# You can use any of the previous models without the `_augmented` suffix. However, those models were trained with less data and have less entities, so we highly recommend to use the `augmented` versions.
| tutorials/Certification_Trainings/Healthcare/4.2.Clinical_Deidentification_in_Spanish.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-1:081325390199:image/datascience-1.0
# ---
# # Feature Transformation with Scikit-Learn In This Notebook
# ## Saving Features into the SageMaker Feature Store
#
# In this notebook, we convert raw text into BERT embeddings. This will allow us to perform natural language processing tasks such as text classification. We save the features into the SageMaker Feature Store.
#
# 
# +
import sagemaker
import boto3
sess = sagemaker.Session()
bucket = sess.default_bucket()
role = sagemaker.get_execution_role()
region = boto3.Session().region_name
sm = boto3.Session().client(service_name="sagemaker", region_name=region)
s3 = boto3.Session().client(service_name="s3", region_name=region)
# -
# 
# # Convert Raw Text to BERT Features using Hugging Face and TensorFlow
# +
import tensorflow as tf
import collections
import json
import os
import pandas as pd
import csv
from transformers import DistilBertTokenizer
tokenizer = DistilBertTokenizer.from_pretrained("distilbert-base-uncased")
REVIEW_BODY_COLUMN = "review_body"
REVIEW_ID_COLUMN = "review_id"
# DATE_COLUMN = 'date'
LABEL_COLUMN = "star_rating"
LABEL_VALUES = [1, 2, 3, 4, 5]
label_map = {}
for (i, label) in enumerate(LABEL_VALUES):
label_map[label] = i
class InputFeatures(object):
"""BERT feature vectors."""
def __init__(self, input_ids, input_mask, segment_ids, label_id, review_id, date, label):
self.input_ids = input_ids
self.input_mask = input_mask
self.segment_ids = segment_ids
self.label_id = label_id
self.review_id = review_id
self.date = date
self.label = label
class Input(object):
"""A single training/test input for sequence classification."""
def __init__(self, text, review_id, date, label=None):
"""Constructs an Input.
Args:
text: string. The untokenized text of the first sequence. For single
sequence tasks, only this sequence must be specified.
label: (Optional) string. The label of the example. This should be
specified for train and dev examples, but not for test examples.
"""
self.text = text
self.review_id = review_id
self.date = date
self.label = label
def convert_input(the_input, max_seq_length):
# First, we need to preprocess our data so that it matches the data BERT was trained on:
# 1. Lowercase our text (if we're using a BERT lowercase model)
# 2. Tokenize it (i.e. "sally says hi" -> ["sally", "says", "hi"])
# 3. Break words into WordPieces (i.e. "calling" -> ["call", "##ing"])
#
# Fortunately, the Transformers tokenizer does this for us!
tokens = tokenizer.tokenize(the_input.text)
tokens.insert(0, '[CLS]')
tokens.append('[SEP]')
print("**{} tokens**\n{}\n".format(len(tokens), tokens))
encode_plus_tokens = tokenizer.encode_plus(
the_input.text,
pad_to_max_length=True,
max_length=max_seq_length,
truncation=True
)
# The id from the pre-trained BERT vocabulary that represents the token. (Padding of 0 will be used if the # of tokens is less than `max_seq_length`)
input_ids = encode_plus_tokens["input_ids"]
# Specifies which tokens BERT should pay attention to (0 or 1). Padded `input_ids` will have 0 in each of these vector elements.
input_mask = encode_plus_tokens["attention_mask"]
# Segment ids are always 0 for single-sequence tasks such as text classification. 1 is used for two-sequence tasks such as question/answer and next sentence prediction.
segment_ids = [0] * max_seq_length
# Label for each training row (`star_rating` 1 through 5)
label_id = label_map[the_input.label]
features = InputFeatures(
input_ids=input_ids,
input_mask=input_mask,
segment_ids=segment_ids,
label_id=label_id,
review_id=the_input.review_id,
date=the_input.date,
label=the_input.label,
)
print("**input_ids**\n{}\n".format(features.input_ids))
print("**input_mask**\n{}\n".format(features.input_mask))
print("**segment_ids**\n{}\n".format(features.segment_ids))
print("**label_id**\n{}\n".format(features.label_id))
print("**review_id**\n{}\n".format(features.review_id))
print("**date**\n{}\n".format(features.date))
print("**label**\n{}\n".format(features.label))
return features
# We'll need to transform our data into a format that BERT understands.
# - `text` is the text we want to classify, which in this case, is the `Request` field in our Dataframe.
# - `label` is the star_rating label (1, 2, 3, 4, 5) for our training input data
def transform_inputs_to_tfrecord(inputs, output_file, max_seq_length):
records = []
tf_record_writer = tf.io.TFRecordWriter(output_file)
for (input_idx, the_input) in enumerate(inputs):
if input_idx % 10000 == 0:
print("Writing input {} of {}\n".format(input_idx, len(inputs)))
features = convert_input(the_input, max_seq_length)
all_features = collections.OrderedDict()
# Create TFRecord With input_ids, input_mask, segment_ids, and label_ids
all_features["input_ids"] = tf.train.Feature(int64_list=tf.train.Int64List(value=features.input_ids))
all_features["input_mask"] = tf.train.Feature(int64_list=tf.train.Int64List(value=features.input_mask))
all_features["segment_ids"] = tf.train.Feature(int64_list=tf.train.Int64List(value=features.segment_ids))
all_features["label_ids"] = tf.train.Feature(int64_list=tf.train.Int64List(value=[features.label_id]))
tf_record = tf.train.Example(features=tf.train.Features(feature=all_features))
tf_record_writer.write(tf_record.SerializeToString())
# Create Record For Feature Store With All Features
records.append(
{ #'tf_record': tf_record.SerializeToString(),
"input_ids": features.input_ids,
"input_mask": features.input_mask,
"segment_ids": features.segment_ids,
"label_id": features.label_id,
"review_id": the_input.review_id,
"date": the_input.date,
"label": features.label,
# 'review_body': features.review_body
}
)
tf_record_writer.close()
return records
# -
# Three(3) feature vectors are created from each raw review (`review_body`) during the feature engineering phase to prepare for BERT processing:
#
# * **`input_ids`**: The id from the pre-trained BERT vocabulary that represents the token. (Padding of 0 will be used if the # of tokens is less than `max_seq_length`)
#
# * **`input_mask`**: Specifies which tokens BERT should pay attention to (0 or 1). Padded `input_ids` will have 0 in each of these vector elements.
#
# * **`segment_ids`**: Segment ids are always 0 for single-sequence tasks such as text classification. 1 is used for two-sequence tasks such as question/answer and next sentence prediction.
#
# And one(1) label is created from each raw review (`star_rating`) :
#
# * **`label_id`**: Label for each training row (`star_rating` 1 through 5)
# # Demonstrate the BERT-specific Feature Engineering Step
# While we are demonstrating this code with a small amount of data here in the notebook, we will soon scale this to much more data on a powerful SageMaker cluster.
# ## Feature Store requires an Event Time feature
#
# We need a record identifier name and an event time feature name. This will match the column of the corresponding features in our data.
#
# Note: Event time date feature type provided Integral. Event time type should be either Fractional(Unix timestamp in seconds) or String (ISO-8601 format) type.
# +
from datetime import datetime
from time import strftime
# timestamp = datetime.now().replace(microsecond=0).isoformat()
timestamp = datetime.now().strftime("%Y-%m-%dT%H:%M:%SZ")
print(timestamp)
# +
import pandas as pd
data = [
[
5,
"ABCD12345",
"""I needed an "antivirus" application and know the quality of Norton products. This was a no brainer for me and I am glad it was so simple to get.""",
],
[
3,
"EFGH12345",
"""The problem with ElephantDrive is that it requires the use of Java. Since Java is notorious for security problems I haveit removed from all of my computers. What files I do have stored are photos.""",
],
[
1,
"IJKL2345",
"""Terrible, none of my codes worked, and I can't uninstall it. I think this product IS malware and viruses""",
],
]
df = pd.DataFrame(data, columns=["star_rating", "review_id", "review_body"])
# Use the InputExample class from BERT's run_classifier code to create examples from the data
inputs = df.apply(
lambda x: Input(label=x[LABEL_COLUMN], text=x[REVIEW_BODY_COLUMN], review_id=x[REVIEW_ID_COLUMN], date=timestamp),
axis=1,
)
# -
# Make sure the date is in the correct ISO-8601 format for Feature Store
print(inputs[0].date)
# ## Save TFRecords
#
# The three(3) features vectors and one(1) label are converted into a list of `TFRecord` instances (1 per each row of training data):
# * **`tf_records`**: Binary representation of each row of training data (3 features + 1 label)
#
# These `TFRecord`s are the engineered features that we will use throughout the rest of the pipeline.
output_file = "./data-tfrecord-featurestore/data.tfrecord"
# # Add Features to SageMaker Feature Store
# ## SageMaker Feature Store Runtime
#
# A low-level client representing Amazon SageMaker Feature Store Runtime
#
# Contains all data plane API operations and data types for the Amazon SageMaker Feature Store. Use this API to put, delete, and retrieve (get) features from a feature store.
featurestore_runtime = boto3.Session().client(service_name="sagemaker-featurestore-runtime", region_name=region)
# ## Create FeatureGroup
#
# A feature group is a logical grouping of features, defined in the Feature Store, to describe records. A feature group definition is composed of a list of feature definitions, a record identifier name, and configurations for its online and offline store.
#
# Create feature group, describe feature group, update feature groups, delete feature group and list feature groups APIs can be used to manage feature groups.
#
# +
from time import gmtime, strftime, sleep
feature_group_name = "reviews-feature-group-" + strftime("%d-%H-%M-%S", gmtime())
print(feature_group_name)
# +
from sagemaker.feature_store.feature_definition import (
FeatureDefinition,
FeatureTypeEnum,
)
feature_definitions = [
FeatureDefinition(feature_name="input_ids", feature_type=FeatureTypeEnum.STRING),
FeatureDefinition(feature_name="input_mask", feature_type=FeatureTypeEnum.STRING),
FeatureDefinition(feature_name="segment_ids", feature_type=FeatureTypeEnum.STRING),
FeatureDefinition(feature_name="label_id", feature_type=FeatureTypeEnum.INTEGRAL),
FeatureDefinition(feature_name="review_id", feature_type=FeatureTypeEnum.STRING),
FeatureDefinition(feature_name="date", feature_type=FeatureTypeEnum.STRING),
FeatureDefinition(feature_name="label", feature_type=FeatureTypeEnum.INTEGRAL),
# FeatureDefinition(feature_name='review_body', feature_type=FeatureTypeEnum.STRING),
FeatureDefinition(feature_name="split_type", feature_type=FeatureTypeEnum.STRING),
]
# +
from sagemaker.feature_store.feature_group import FeatureGroup
feature_group = FeatureGroup(name=feature_group_name, feature_definitions=feature_definitions, sagemaker_session=sess)
print(feature_group)
# -
# ## Specify `record identifier` and `event time` features
record_identifier_feature_name = "review_id"
event_time_feature_name = "date"
# ## Set S3 Prefix for Offline Feature Store
prefix = "reviews-feature-store-" + timestamp
print(prefix)
# ## Create Feature Group
#
# The last step for creating the feature group is to use the `create` function. The online store is not created by default, so we must set this as `True` if we want to enable it. The `s3_uri` is the location of our offline store.
feature_group.create(
s3_uri=f"s3://{bucket}/{prefix}",
record_identifier_name=record_identifier_feature_name,
event_time_feature_name=event_time_feature_name,
role_arn=role,
enable_online_store=False,
)
# ## Describe the Feature Group
feature_group.describe()
# ## List All Feature Groups
#
# We use the boto3 SageMaker client to list all FeatureGroups.
# +
# sm.list_feature_groups()
# -
# ## Wait For The Feature Group Creation Complete
#
# Creating a feature group takes time as the data is loaded. We will need to wait until it is created before you can use it. You can check status using the following method.
# +
import time
def wait_for_feature_group_creation_complete(feature_group):
status = feature_group.describe().get("FeatureGroupStatus")
while status == "Creating":
print("Waiting for Feature Group Creation")
time.sleep(5)
status = feature_group.describe().get("FeatureGroupStatus")
if status != "Created":
raise RuntimeError(f"Failed to create feature group {feature_group.name}")
print(f"FeatureGroup {feature_group.name} successfully created.")
# -
wait_for_feature_group_creation_complete(feature_group=feature_group)
# ## Review The Records To Ingest Into Feature Store
max_seq_length = 64
records = transform_inputs_to_tfrecord(inputs, output_file, max_seq_length)
# # Ingest Records into Feature Store
#
# After the FeatureGroups have been created, we can put data into the FeatureGroups by using the `PutRecord` API.
#
# This API can handle high TPS and is designed to be called by different streams. The data from all of these Put requests is buffered and written to S3 in chunks.
#
# The files will be written to the offline store within a few minutes of ingestion. To accelerate the ingestion process, we can specify multiple workers to do the job simultaneously.
#
# Use `put_record(...)` to put a single record in the FeatureGroup.
#
# Use `ingest(...)` to ingest the content of a pandas DataFrame to Feature Store. You can set the `max_worker` to the number of threads to be created to work on different partitions of the `data_frame` in parallel.
# +
import pandas as pd
df_records = pd.DataFrame.from_dict(records)
df_records["split_type"] = "train"
df_records
# -
# # Cast DataFrame `Object` to Supported Feature Store Data Type `String`
def cast_object_to_string(data_frame):
for label in data_frame.columns:
if data_frame.dtypes[label] == "object":
data_frame[label] = data_frame[label].astype("str").astype("string")
cast_object_to_string(df_records)
df_records
feature_group.ingest(data_frame=df_records, max_workers=3, wait=True)
# ## Wait For Data In Offline Feature Store To Become Available
#
# Creating a feature group takes time as the data is loaded. We will need to wait until it is created before we can use it.
# +
offline_store_contents = None
while offline_store_contents is None:
objects_in_bucket = s3.list_objects(Bucket=bucket, Prefix=prefix)
if "Contents" in objects_in_bucket and len(objects_in_bucket["Contents"]) > 1:
offline_store_contents = objects_in_bucket["Contents"]
else:
print("Waiting for data in offline store...\n")
sleep(60)
print("Data available.")
# -
# ## _Wait For The Cell Above To Complete and show `Data available`._
# ## Get Record From Online Feature Store
#
# Use for OnlineStore serving from a FeatureStore. Only the latest records stored in the OnlineStore can be retrieved. If no Record with `RecordIdentifierValue` is found, then an empty result is returned.
# +
# record_identifier_value = "IJKL2345"
# featurestore_runtime.get_record(
# FeatureGroupName=feature_group_name, RecordIdentifierValueAsString=record_identifier_value
# )
# -
# # Build Training Dataset
#
# SageMaker FeatureStore automatically builds the Glue Data Catalog for FeatureGroups (we can optionally turn it on/off while creating the FeatureGroup). We can create a training dataset by querying the data in the feature store. This is done by utilizing the auto-built Catalog and run an Athena query.
# # Create An Athena Query
feature_store_query = feature_group.athena_query()
# # Get The Feature Group Table Name
feature_store_table = feature_store_query.table_name
# # Build an Athena SQL Query
#
# Show Hive DDL commands to define or change structure of tables or databases in Hive. The schema of the table is generated based on the feature definitions. Columns are named after feature name and data-type are inferred based on feature type.
#
# Integral feature type is mapped to INT data-type. Fractional feature type is mapped to FLOAT data-type. String feature type is mapped to STRING data-type.
print(feature_group.as_hive_ddl())
# +
query_string = """
SELECT input_ids, input_mask, segment_ids, label_id, split_type FROM "{}" WHERE split_type='train' LIMIT 5
""".format(
feature_store_table
)
print("Running " + query_string)
# -
# ## Run Athena Query
# The query results are stored in a S3 bucket.
# +
feature_store_query.run(query_string=query_string, output_location="s3://" + bucket + "/" + prefix + "/query_results/")
feature_store_query.wait()
# -
# ## View Query Results
#
# Load query results in a Pandas DataFrame.
# +
dataset = pd.DataFrame()
dataset = feature_store_query.as_dataframe()
dataset
# -
# # Review the Feature Store
#
# 
# # Release Resources
# + language="html"
#
# <p><b>Shutting down your kernel for this notebook to release resources.</b></p>
# <button class="sm-command-button" data-commandlinker-command="kernelmenu:shutdown" style="display:none;">Shutdown Kernel</button>
#
# <script>
# try {
# els = document.getElementsByClassName("sm-command-button");
# els[0].click();
# }
# catch(err) {
# // NoOp
# }
# </script>
# + language="javascript"
#
# try {
# Jupyter.notebook.save_checkpoint();
# Jupyter.notebook.session.delete();
# }
# catch(err) {
# // NoOp
# }
| 06_prepare/01_Prepare_Dataset_BERT_Scikit_AdHoc_FeatureStore.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 [dev-base]
# language: python
# name: anaconda-project-dev-base-python3
# ---
namenode_url = 'http://ec2-3-93-61-21.compute-1.amazonaws.com:50070'
from livy_submit.hdfs_api import get_client
client = get_client(namenode_url)
# !echo anaconda | kinit edill
client.list('')
| dev-notebooks/check-webhdfs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/GitMarco27/TMML/blob/main/Notebooks/004_Visualize_Model.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="eZqrzUILLSQ6"
# # 3 Minutes Machine Learning
# ## Episode 4: Visualize a Model
#
# #### <NAME>, 2021
# ---
# Welcome to 3 minutes Machine Learning!
# + id="cRD3PvNVB_cW"
import tensorflow as tf
from tensorflow import keras
from keras.layers import Dense, Input, Conv2D, MaxPool2D, Flatten
from keras.models import Model
import matplotlib.pyplot as plt
# + id="7LRdURNnLbgo"
def loadThumb(path):
# Let's import this video thumbnail!
myThumb = plt.imread(path)
fig, ax = plt.subplots(figsize=(15, 10))
plt.axis('off')
ax.imshow(myThumb)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 502} id="OGQ7JXI4LdgN" outputId="be90df99-c0c4-4e6e-fb8c-28de082a425d"
# loadThumb('/tmp/yt_thumb_004.png')
# + [markdown] id="qGMAbZBjLimI"
# #### Video Topics
# > 1. Create a dummy tensor
# > 2. Create a model with the keras API
# > 3. Plot our model and evaluate with dummy data
# > 4. See you on next video!
# + [markdown] id="6KXkxr1PLsbk"
# # Create dummy a dummy tensor
# ---
# + id="tLKyQBTJNqs4"
# Create a dummy image with the shape: (B, N, M, 3) where
# B: Batch Size
# N: Dummy image Length
# M: Dummy image Heigth
# 3: Channels
# + id="sY-vDgqsCDxJ"
data = tf.random.uniform(shape=[1, 1920, 1080, 3], dtype=tf.dtypes.float32, name='data')
# + [markdown] id="74nL7T1sLwz9"
# # Create a model with keras api
# ---
# + id="0UNRObD1OGJw"
# Let's crate the model that we want to visualize with the tensorflow.keras Functional APIa
# + id="lXeOferSCuxZ"
input = Input(shape=(1920, 1080, 3), name='input')
conv_1 = Conv2D(16, (3, 3), activation=tf.nn.relu)(input)
max_p_1 = MaxPool2D((2, 2))(conv_1)
conv_2 = Conv2D(32, (3, 3), activation=tf.nn.relu)(max_p_1)
max_p_2 = MaxPool2D((2, 2))(conv_2)
conv_3 = Conv2D(64, (3, 3), activation=tf.nn.relu)(max_p_2)
max_p_3 = MaxPool2D((2, 2))(conv_3)
flat = Flatten()(max_p_3)
dense_1 = Dense(1, activation=tf.nn.sigmoid, name='output')(flat)
# + colab={"base_uri": "https://localhost:8080/"} id="ROSSnqd1EIyu" outputId="878c1966-908a-4043-ece7-2eead320f6f3"
model = Model(inputs=input, outputs=dense_1)
model.compile(optimizer='SGD')
model.summary()
# + [markdown] id="5X5LmM97L2ij"
# # Plot and test the model
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 976} id="js3cQpOLE63R" outputId="241946c2-195e-4a41-da7c-ccd81d3f04ea"
tf.keras.utils.plot_model(model, to_file="my_model.png", show_shapes=True)
# + colab={"base_uri": "https://localhost:8080/"} id="qMgoqyPzHwCe" outputId="2ca441cf-3a6e-4c35-ff00-ac0300ab7661"
model.predict(data, batch_size=1)
# + [markdown] id="cA-z4jszL9sl"
# # Greetings
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="3dNMiD3oMAMN" outputId="e521f0b3-c786-4b94-b62d-b7f73aeeb2a3"
# !pip install art
# + colab={"base_uri": "https://localhost:8080/"} id="4pN6CKU1MF26" outputId="53b75554-73f5-4791-e568-ad57fcc1a065"
from art import tprint, aprint
tprint('See you on next videos!')
def subscribe():
"""
Attractive subscription form
"""
aprint("giveme", number=5)
print(f'\n\tLike and subscribe to support this work!\n')
aprint("giveme", number=5)
subscribe()
| Notebooks/004_Visualize_Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="Rwq8g-CTl_z0" colab_type="code" colab={}
import mxnet as mx
# + id="3rXpSs8Den1i" colab_type="code" outputId="e5e8747f-e267-4ec3-90b8-81cf4bf94f1a" colab={"base_uri": "https://localhost:8080/", "height": 180}
# !pip install mxnet-cu80
# + id="UGmnvKCldJEa" colab_type="code" colab={}
import numpy as np
x_input = mx.nd.empty((1, 5), mx.cpu())
x_input[:] = np.array([[1, 2, 3, 4, 5]], np.float32)
y_input = mx.nd.empty((1, 5), mx.cpu())
y_input[:] = np.array([[10, 15, 20, 22.5, 25]], np.float32)
# + id="80lFO21JeX4I" colab_type="code" colab={}
x_input
w_input = x_input
z_input = x_input.copyto(mx.cpu())
x_input += 1
w_input /= 2
z_input *= 2
# + id="gBwV6S14gFr-" colab_type="code" outputId="6062b578-1b6c-4312-c126-db3000c16f31" colab={"base_uri": "https://localhost:8080/", "height": 71}
print("x_input: ", x_input.asnumpy())
print("w_input: ", w_input.asnumpy())
print("z_input: ", z_input.asnumpy())
# + id="B1ewMnI8gZSj" colab_type="code" colab={}
batch_size = 1
train_iter = mx.io.NDArrayIter(x_input, y_input, batch_size,
shuffle = True, data_name = 'input',
label_name = 'target')
# + id="QTqw251Sg3aZ" colab_type="code" colab={}
X = mx.sym.Variable('input')
Y = mx.symbol.Variable('target')
fc1 = mx.sym.FullyConnected(data = X, name = 'fc1', num_hidden = 5)
lin_reg = mx.sym.LinearRegressionOutput(data = fc1, label = Y,
name = "lin_reg")
# + id="4QkuWyt-hUVP" colab_type="code" colab={}
model = mx.mod.Module(
symbol = lin_reg,
data_names = ['input'],
label_names = ['target']
)
# + id="Ch5szLFzhd4y" colab_type="code" colab={}
model.fit(train_iter,
optimizer_params = {'learning_rate': 0.01, 'momentum': 0.9},
num_epoch = 100,
batch_end_callback = mx.callback.Speedometer(batch_size, 2))
# + id="IBJoB-iIh5xp" colab_type="code" outputId="d66a241d-66de-4bd7-b24a-91bc6c771239" colab={"base_uri": "https://localhost:8080/", "height": 53}
model.predict(train_iter).asnumpy()
| Chapter 1/5_Building efficient models with MXNet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.5
# language: python
# name: python3
# ---
# # SIT742: Modern Data Science
# **(Week 07: Big Data Platform (II))**
#
# ---
# - Materials in this module include resources collected from various open-source online repositories.
# - You are free to use, change and distribute this package.
# - If you found any issue/bug for this document, please submit an issue at [tulip-lab/sit742](https://github.com/tulip-lab/sit742/issues)
#
# Prepared by **SIT742 Teaching Team**
#
# ---
#
#
#
# ## Session 7B - Spark MLlib (1): Data Types
#
#
# The purpose of this session is to demonstrate different coefficient and linear regression.
#
#
# ### Content
#
# ### Part 1 Vectors
#
# 1.1 [Dense and Sparse Vectors](#dsvec)
#
# 1.2 [Labeled Points](#lpoints)
#
#
# ### Part 2 Matrix
#
# Local Matrix
# Row Matrix
#
#
# 2.1 [Local Matrix](#lm)
#
# 2.2 [Row Matrix](#rm)
#
# 2.3 [Indexed Row Matrix](#irm)
#
# 2.4 [Coordinate Matrix](#cm)
#
# 2.5 [Block Matrix](#bm)
#
# ### Part 3 Matrix Conversions
#
# 3.1 [Indexed Row Matrix Conversions](#irmc)
#
# 3.2 [Coordinate Matrix Conversions](#cmc)
#
# 3.3 [Block Matrix Conversions](#bmc)
#
# ---
# ## <span style="color:#0b486b">1. Vectors</span>
#
# <a id = "dsvec"></a>
# ### <span style="color:#0b486b">1.1 Dense and Sparse Vectors</span>
#
# Spark has many libraries, namely under MLlib (Machine Learning Library)! Spark allows for quick and easy scalability of practical machine learning!
#
# In this lab exercise, you will learn about the basic Data Types that are used in Spark MLlib. This lab will help you develop the building blocks required to continue developing knowledge in machine learning with Spark.
#
# Import the following libraries: <br>
# <ul>
# <li> numpy as np </li>
# <li> scipy.sparse as sps </li>
# <li> Vectors from pyspark.mllib.linalg </li>
# </ul>
# +
import numpy as np
import scipy.sparse as sps
from pyspark.mllib.linalg import Vectors
import time
# -
# First, we will be dealing with <b>Dense Vectors</b>. There are 2 types of <b>dense vectors</b> that we can create.<br>
# The dense vectors will be modeled having the values: <b>8.0, 312.0, -9.0, 1.3</b>
# The first <b>dense vector</b> we will create is as easy as creating a <b>numpy array</b>. <br>
# Using the np.array function, create a <b>dense vector</b> called <b>dense_vector1</b> <br> <br>
# Note: numpy's array function takes an array as input
dense_vector1 = np.array([8.0, 312.0, -9.0, 1.3])
# Print <b>dense_vector1</b> and its <b>type</b>
print (dense_vector1)
type(dense_vector1)
# The second <b>dense vector</b> is easier than the first, and is made by creating an <b>array</b>. <br>
# Create a <b>dense vector</b> called <b>dense_vector2</b>
dense_vector2 = [8.0, 312.0, -9.0, 1.3]
# Print <b>dense_vector2</b> and its <b>type</b>
print (dense_vector2)
type (dense_vector2)
# Next, we will be dealing with <b>sparse vectors</b>. There are 2 types of <b>sparse vectors</b> we can create. <br>
# The sparse vectors we will be creating will follow these values: <b> 7.0, 0.0, 0.0, 2.0, 0.0, 1.0, 0.0, 0.0, 0.0, 6.5 </b>
# First, create a <b>sparse vector</b> called <b>sparse_vector1</b> using Vector's <b>sparse</b> function. <br>
# Inputs to Vector.sparse: <br>
# <ul>
# <li>1st: Size of the sparse vector</li>
# <li>2nd: Indicies of array</li>
# <li>3rd: Values placed where the indices are</li>
# </ul>
sparse_vector1 = Vectors.sparse(10, [0, 3, 5, 9], [7.0, 2.0, 1.0, 6.5])
# Print <b>sparse_vector1</b> and its <b>type</b>
print(sparse_vector1)
type(sparse_vector1)
# Next we will create a <b>sparse vector</b> called <b>sparse_vector2</b> using a single-column SciPy <b>csc_matrix</b> <br> <br>
# The inputs to sps.csc_matrix are: <br>
# <ul>
# <li>1st: A tuple consisting of the three inputs:</li>
# <ul>
# <li>1st: Data Values (in a numpy array) (values placed at the specified indices)</li>
# <li>2nd: Indicies of the array (in a numpy array) (where the values will be placed)</li>
# <li>3rd: Index pointer of the array (in a numpy array)</li>
# </ul>
# <li>2nd: Shape of the array (#rows, #columns) Use 10 rows and 1 column</li>
# <ul>
# <li>shape = (\_,\_)</li>
# </ul>
# </ul> <br>
# Note: You may get a deprecation warning. Please Ignore it.
sparse_vector2 = sps.csc_matrix((np.array([7.0, 2.0, 1.0, 6.5]), np.array([0, 3, 5, 9]), np.array([0, 4])), shape = (10, 1))
# Print <b>sparse_vector2</b> and its <b>type</b>
print (sparse_vector2)
print (type(sparse_vector2))
# <a id = "lpoints"></a>
# ### <span style="color:#0b486b">1.2 Labeled Points</span>
#
# So the next data type will be Labeled points. Remember that this data type is mainly used for classification algorithms in supervised learning.<br>
#
# Start by importing the following libraries: <br>
# <ul>
# <li>SparseVector from pyspark.mllib.linalg</li>
# <li>LabeledPoint from pyspark.mllib.regression</li>
# </ul>
from pyspark.mllib.linalg import SparseVector
from pyspark.mllib.regression import LabeledPoint
# Remember that with a lableled point, we can create binary or multiclass classification. In this lab, we will deal with binary classification for ease. <br> <br>
# The <b>LabeledPoint</b> function takes in 2 inputs:
# <ul>
# <li>1st: Label of the Point. In this case (for binary classification), we will be using <font color="green">1.0</font> for <font color="green">positive</font> and <font color="red">0.0</font> for <font color="red">negative</font></li>
# <li>2nd: Vector of features for the point (We will input a Dense or Sparse Vector using any of the methods defined in the <b>Dense and Sparse Vectors</b> section of this lab.</b>
# </ul>
# Using the LabelPoint class, create a <b>dense</b> feature vector with a <b>positive</b> label called <b>pos_class</b> with the values: <b>5.0, 2.0, 1.0, 9.0</b>
pos_class = LabeledPoint(1.0, [5.0, 2.0, 1.0, 9.0])
# Print <i>pos_class</i> and its <i>type</i>
print(pos_class)
type(pos_class)
# Next we will create a <b>sparse</b> feature vector with a <b>negative</b> label called <b>neg_class</b> with the values: <b>1.0, 0.0, 0.0, 4.0, 0.0, 2.0</b>
neg_class = LabeledPoint(0.0, SparseVector(6, [0, 3, 5], [1.0, 4.0, 2.0]))
# Print <b>neg_class</b> and its <b>type</b>
print(neg_class)
type(neg_class)
# ---
# ## <span style="color:#0b486b">2. Matrix Data Types</span>
#
#
# In this next section, we will be dealing creating the following matrices:
# <ul>
# <li>Local Matrix</li>
# <li>Row Matrix</li>
# <li>Indexed Row Matrix</li>
# <li>Coordinate Matrix</li>
# <li>Block Matrix</li>
# </ul>
#
# Throughout this section, we will be modelling the following matricies: <br>
#
# <center>For a Dense Matrix:</center> <br>
#
# $$
# \begin{pmatrix}
# 1.00 & 6.00 & 3.00 & 0.00 \\
# 3.00 & 2.00 & 5.00 & 1.00 \\
# 9.00 & 4.00 & 0.00 & 3.00
# \end{pmatrix}
# $$
#
# <center>For a Sparse Matrix:</center> <br>
#
# $$
# \begin{pmatrix}
# 1.00 & 0.00 & 3.00 & 0.00 \\
# 3.00 & 0.00 & 0.00 & 1.00 \\
# 0.00 & 4.00 & 0.00 & 0.00
# \end{pmatrix}
# $$
# <a id = "lm"></a>
# ### <span style="color:#0b486b">2.1 Local Matrix</span>
#
# Import the following Library:
# <ul>
# <li>pyspark.mllib.linalg as laMat</li>
# </ul>
import pyspark.mllib.linalg as laMat
# Create a dense local matrix called <b>dense_LM</b> <br>
# The inputs into the <b>laMat.Matrices.dense</b> function are:
# <ul>
# <li>1st: Number of Rows</li>
# <li>2nd: Number of Columns</li>
# <li>3rd: Values in an array format (Read as Column-Major)</li>
# </ul>
dense_LM = laMat.Matrices.dense(3,4, [1.0, 3.0, 9.0, 6.0, 2.0, 4.0, 3.0, 5.0, 0.0, 0.0, 1.0, 3.0])
# Print <b>dense_LM</b> and its <b>type</b>
print(dense_LM)
type(dense_LM)
# Next we will do the same thing with a sparse matrix, calling the output <b>sparse_LM</b>
# The inputs into the <b>laMat.Matrices.sparse</b> function are:
# <ul>
# <li>1st: Number of Rows</li>
# <li>2nd: Number of Columns</li>
# <li>3rd: Column Pointers (in a list)</li>
# <li>4th: Row Indices (in a list)</li>
# <li>5th: Values of the Matrix (in a list)</li>
# </ul> <br>
# <b>Note</b>: Remember that this is <b>column-major</b> so all arrays should be read as columns first (top down, left to right)
sparse_LM = laMat.Matrices.sparse(3, 4, [0, 2, 3, 4, 5], [0, 1, 2, 1, 1], [1.0, 3.0, 4.0, 3.0, 1.0])
# Print <b>sparse_LM</b> and its <b>type</b>
print(sparse_LM)
type(sparse_LM)
# Make sure the output of <b>sparse_LM</b> matches the original matrix.
# <a id = "rm"></a>
# ### <span style="color:#0b486b">2.2 Row Matrix</span>
#
# A RowMatrix is a Row-oriented distributed matrix that doesn't have meaningful row indices.
#
# Import the following library:
# <ul>
# <li>RowMatrix from pyspark.mllib.linalg.distributed</li>
# </ul>
from pyspark.mllib.linalg.distributed import RowMatrix
# Now, let's create a RDD of vectors called <b>rowVecs</b>, using the SparkContext's parallelize function on the <b>Dense Matrix</b>.<br>
# The input into <b>sc.parallelize</b> is:
# <ul>
# <li>A list (The list we will be creating will be a list of the row values (each row is a list))</li>
# </ul> <br>
# <b>Note</b>: And RDD is a fault-tolerated collection of elements that can be operated on in parallel. <br>
rowVecs = sc.parallelize([[1.0, 6.0, 3.0, 0.0],
[3.0, 2.0, 5.0, 1.0],
[9.0, 4.0, 0.0, 3.0]])
# Next, create a variable called <b>rowMat</b> by using the <b>RowMatrix</b> function and passing in the RDD.
rowMat = RowMatrix(rowVecs)
# Now we will retrieve the <font color="green">row numbers</font> (save it as <font color="green">m</font>) and <font color="blue">column numbers</font> (save it as <font color="blue">n</font>) from the RowMatrix.
# <ul>
# <li>To get the number of rows, use <i>numRows()</i> on rowMat</li>
# <li>To get the number of columns, use <i>numCols()</i> on rowMat</li>
# </ul>
m = rowMat.numRows()
n = rowMat.numCols()
# Print out <b>m</b> and <b>n</b>. The results should be:
# <ul>
# <li>Number of Rows: 3</li>
# <li>Number of Columns: 4</li>
# </ul>
# +
print(m)
print(n)
# -
# <a id = "irm"></a>
# ### <span style="color:#0b486b">2.3 Indexed Row Matrix</span>
#
# Since we just created a RowMatrix, which had no meaningful row indicies, let's create an <b>IndexedRowMatrix</b> which has meaningful row indices!
#
# Import the following Library:
# <ul>
# <li> IndexedRow, IndexedRowMatrix from pyspark.mllib.linalg.distributed</li>
# </ul>
from pyspark.mllib.linalg.distributed import IndexedRow, IndexedRowMatrix
# Now, create a RDD called <b>indRows</b> by using the SparkContext's parallelize function on the <b>Dense Matrix</b>. <br>
# There are two different inputs you can use to create the RDD:
# <ul>
# <li>Method 1: A list containing multiple IndexedRow inputs</li>
# <ul>
# <li>Input into IndexedRow:</li>
# <ul>
# <li>1. Index for the given row (row number)</li>
# <li>2. row in the matrix for the given index</li>
# </ul>
# <li>ex. sc.parallelize([IndexedRow(0,[1, 2, 3]), ...])</li>
# </ul> <br>
# <li>Method 2: A list containing multiple tuples</li>
# <ul>
# <li>Values in the tuple:</li>
# <ul>
# <li>1. Index for the given row (row number) (type:long)</li>
# <li>2. List containing the values in the row for the given index (type:vector)</li>
# </ul>
# <li>ex. sc.parallelize([(0, [1, 2, 3]), ...])</li>
# </ul>
# </ul>
# +
# Method 1: Using IndexedRow class
indRows = sc.parallelize([IndexedRow(0, [1.0, 6.0, 3.0, 0.0]),
IndexedRow(1, [3.0, 2.0, 5.0, 1.0]),
IndexedRow(2, [9.0, 4.0, 0.0, 3.0])])
# Method 2: Using (long, vector) tuples
indRows = sc.parallelize([(0, [1.0, 6.0, 3.0, 0.0]),
(1, [3.0, 2.0, 5.0, 1.0]),
(2, [9.0, 4.0, 0.0, 3.0])])
# -
# Now, create the <b>IndexedRowMatrix</b> called <b>indRowMat</b> by using the IndexedRowMatrix function and passing in the <b>indRows</b> RDD
indRowMat = IndexedRowMatrix(indRows)
# Now we will retrieve the <font color="green">row numbers</font> (save it as <font color="green">m2</font>) and <font color="blue">column numbers</font> (save it as <font color="blue">n2</font>) from the IndexedRowMatrix.
# <ul>
# <li>To get the number of rows, use <i>numRows()</i> on indRowMat</li>
# <li>To get the number of columns, use <i>numCols()</i> on indRowMat</li>
# </ul>
m2 = indRowMat.numRows()
n2 = indRowMat.numCols()
# Print out <b>m2</b> and <b>n2</b>. The results should be:
# <ul>
# <li>Number of Rows: 3</li>
# <li>Number of Columns: 4</li>
# </ul>
# +
print(m2)
print(n2)
# -
# <a id = "cm"></a>
# ### <span style="color:#0b486b">2.3 Coordinate Matrix</span>
#
#
# Now it's time to create a different type of matrix, whos use should be when both the dimensions of the matrix is very large, and the data in the matrix is sparse. <br>
# <b>Note</b>: In this case, we will be using the small, sparse matrix above, just to get the idea of how to initialize a CoordinateMatrix
#
# Import the following libraries:
# <ul>
# <li>CoordinateMatrix, MatrixEntry from pyspark.mllib.linalg.distributed</li>
# </ul>
from pyspark.mllib.linalg.distributed import CoordinateMatrix, MatrixEntry
# Now, create a RDD called <b>coordRows</b> by using the SparkContext's parallelize function on the <b>Sparse Matrix</b>. There are two different inputs you can use to create the RDD:
# <ul>
# <li>Method 1: A list containing multiple MatrixEntry inputs</li>
# <ul>
# <li>Input into MatrixEntry:</li>
# <ul>
# <li>1. Row index of the matrix (row number) (type: long)</li>
# <li>2. Column index of the matrix (column number) (type: long)</li>
# <li>3. Value at the (Row Index, Column Index) entry of the matrix (type: float)</li>
# </ul>
# <li>ex. sc.parallelize([MatrixEntry(0, 0, 1,), ...])</li>
# </ul> <br>
# <li>Method 2: A list containing multiple tuples</li>
# <ul>
# <li>Values in the tuple:</li>
# <ul>
# <li>1. Row index of the matrix (row number) (type: long)</li>
# <li>2. Column index of the matrix (column number) (type: long)</li>
# <li>3. Value at the (Row Index, Column Index) entry of the matrix (type: float)</li>
# </ul>
# <li>ex. sc.parallelize([(0, 0, 1), ...])</li>
# </ul>
# </ul>
# +
# Method 1. Using MatrixEntry class
coordRows = sc.parallelize([MatrixEntry(0, 0, 1.0),
MatrixEntry(0, 2, 3.0),
MatrixEntry(1, 0, 3.0),
MatrixEntry(1, 3, 1.0),
MatrixEntry(2, 2, 4.0)])
# Method 2. Using (long, long, float) tuples
coordRows = sc.parallelize([(0, 0, 1.0),
(0, 2, 3.0),
(1, 1, 3.0),
(1, 3, 1.0),
(2, 2, 4.0)])
# -
# Now, create the <b>CoordinateMatrix</b> called <b>coordMat</b> by using the CoordinateMatrix function and passing in the <b>coordRows</b> RDD
coordMat = CoordinateMatrix(coordRows)
# Now we will retrieve the <font color="green">row numbers</font> (save it as <font color="green">m3</font>) and <font color="blue">column numbers</font> (save it as <font color="blue">n3</font>) from the CoordinateMatrix.
# <ul>
# <li>To get the number of rows, use <i>numRows()</i> on coordMat</li>
# <li>To get the number of columns, use <i>numCols()</i> on coordMat</li>
# </ul>
m3 = coordMat.numRows()
n3 = coordMat.numCols()
# Print out <b>m3</b> and <b>n3</b>. The results should be:
# <ul>
# <li>Number of Rows: 3</li>
# <li>Number of Columns: 4</li>
# </ul>
# +
print(m3)
print(n3)
# -
# Now, we can get the <b>entries</b> of coordMat by calling the entries method on it. Store this in a variable called coordEnt.
coordEnt = coordMat.entries
# Check out the <i>type</i> of coordEnt.
type(coordEnt)
# It should be a <b>PipelinedRDD</b> type, which has many methods that are associated with it. One of them is <b>first()</b>, which will get the first element in the RDD. <br> <br>
#
# Run coordEnt.first()
coordEnt.first()
# <a id = "bm"></a>
# ### <span style="color:#0b486b">2.4 Block Matrix</span>
#
# A BlockMatrix is essentially a matrix consisting of elements which are partitions of the matrix that is being created.
#
# Import the following libraries:
# <ul>
# <li>Matrices from pyspark.mllib.linalg</li>
# <li>BlockMatrix from pyspark.mllib.linalg.distributed</li>
# </ul>
from pyspark.mllib.linalg import Matrices
from pyspark.mllib.linalg.distributed import BlockMatrix
# Now create a <b>RDD</b> of <b>sub-matrix blocks</b>. <br>
# This will be done using SparkContext's parallelize function. <br>
#
# The input into <b>sc.parallelize</b> requires a <b>list of tuples</b>. The tuples are the sub-matrices, which consist of two inputs:
# <ul>
# <li>1st: A tuple containing the row index and column index (row, column), denoting where the sub-matrix will start</li>
# <li>2nd: The sub-matrix, which will come from <b>Matrices.dense</b>. The sub-matrix requires 3 inputs:</li>
# <ul>
# <li>1st: Number of rows</li>
# <li>2nd: Number of columns</li>
# <li>3rd: A list containing the elements of the sub-matrix. These values are read into the sub-matrix column-major fashion</li>
# </ul>
# </ul> <br>
# (ex. ((51, 2), Matrices.dense(2, 2, [61.0, 43.0, 1.0, 74.0])) would be one row (one tuple)).
# The matrix we will be modelling is <b>Dense Matrix</b> from above. Create the following sub-matrices:
# <ul>
# <li>Row: 0, Column: 0, Values: 1.0, 3.0, 6.0, 2.0, with 2 Rows and 2 Columns </li>
# <li>Row: 2, Column: 0, Values: 9.0, 4.0, with 1 Row and 2 Columns</li>
# <li>Row: 0, Column: 2, Values: 3.0, 5.0, 0.0, 0.0, 1.0, 3.0, with 3 Rows and 2 Columns</li>
# </ul>
blocks = sc.parallelize([((0, 0), Matrices.dense(2, 2, [1.0, 3.0, 6.0, 2.0])),
((2, 0), Matrices.dense(1, 2, [9.0, 4.0])),
((0, 2), Matrices.dense(3, 2, [3.0, 5.0, 0.0, 0.0, 1.0, 3.0]))])
# Now that we have the RDD, it's time to create the BlockMatrix called <b>blockMat</b> using the BlockMatrix class. The <b>BlockMatrix</b> class requires 3 inputs:
# <ul>
# <li>1st: The RDD of sub-matricies</li>
# <li>2nd: The rows per block. Keep this value at 1</li>
# <li>3rd: The columns per block. Keep this value at 1</li>
# </ul>
blockMat = BlockMatrix(blocks, 1, 1)
# Now we will retrieve the <font color="green">row numbers</font> (save it as <font color="green">m4</font>) and <font color="blue">column numbers</font> (save it as <font color="blue">n4</font>) from the BlockMatrix.
# <ul>
# <li>To get the number of rows, use <i>numRows()</i> on blockMat</li>
# <li>To get the number of columns, use <i>numCols()</i> on blockMat</li>
# </ul>
m4 = blockMat.numRows()
n4 = blockMat.numCols()
# Print out <b>m4</b> and <b>n4</b>. The results should be:
# <ul>
# <li>Number of Rows: 3</li>
# <li>Number of Columns: 4</li>
# </ul>
# +
print(m4)
print(n4)
# -
# Now, we need to check if our matrix is correct. We can do this by first converting <b>blockMat</b> into a LocalMatrix, by using the <b>.toLocalMatrix()</b> function on our matrix. Store the result into a variable called <b>locBMat</b>
locBMat = blockMat.toLocalMatrix()
# Now print out <b>locBMat</b> and its <b>type</b>. The result should model the original <b>Dense Matrix</b> and the type should be a DenseMatrix.
print(locBMat)
print(type(locBMat))
# ---
# ## <span style="color:#0b486b">3. Matrix Conversions</span>
#
#
# In this bonus section, we will talk about a relationship between the different types of matrices. You can convert between these matrices that we discussed with the following functions. <br>
# <ul>
# <li>.toRowMatrix() converts the matrix to a RowMatrix</li>
# <li>.toIndexedRowMatrix() converts the matrix to an IndexedRowMatrix</li>
# <li>.toCoordinateMatrix() converts the matrix to a CoordinateMatrix</li>
# <li>.toBlockMatrix() converts the matrix to a BlockMatrix</li>
# </ul>
# <a id = "irmc"></a>
# ### <span style="color:#0b486b">3.1 Indexed Row Matrix Conversions</span>
#
# The following conversions are supported for an IndexedRowMatrix:
# <ul>
# <li>IndexedRowMatrix -> RowMatrix</li>
# <li>IndexedRowMatrix -> CoordinateMatrix</li>
# <li>IndexedRowMatrix -> BlockMatrix</li>
# </ul>
# +
# Convert to a RowMatrix
rMat = indRowMat.toRowMatrix()
print(type(rMat))
# Convert to a CoordinateMatrix
cMat = indRowMat.toCoordinateMatrix()
print(type(cMat))
# Convert to a BlockMatrix
bMat = indRowMat.toBlockMatrix()
print(type(bMat))
# -
# <a id = "cmc"></a>
# ### <span style="color:#0b486b">3.2 Coordinate Matrix Conversions</span>
#
# The following conversions are supported for an CoordinateMatrix:
# <ul>
# <li>CoordinateMatrix -> RowMatrix</li>
# <li>CoordinateMatrix -> IndexedRowMatrix</li>
# <li>CoordinateMatrix -> BlockMatrix</li>
# </ul>
# +
# Convert to a RowMatrix
rMat2 = coordMat.toRowMatrix()
print(type(rMat2))
# Convert to an IndexedRowMatrix
iRMat = coordMat.toIndexedRowMatrix()
print(type(iRMat))
# Convert to a BlockMatrix
bMat2 = coordMat.toBlockMatrix()
print(type(bMat2))
# -
# <a id = "bmc"></a>
# ### <span style="color:#0b486b">3.3 Block Matrix Conversions</span>
#
#
# The following conversions are supported for an BlockMatrix:
# <ul>
# <li>BlockMatrix -> LocalMatrix (Can display the Matrix)</li>
# <li>BlockMatrix -> IndexedRowMatrix</li>
# <li>BlockMatrix -> CoordinateMatrix</li>
# </ul>
# +
# Convert to a LocalMatrix
lMat = blockMat.toLocalMatrix()
print(type(lMat))
# Convert to an IndexedRowMatrix
iRMat2 = blockMat.toIndexedRowMatrix()
print(type(iRMat2))
# Convert to a CoordinateMatrix
cMat2 = blockMat.toCoordinateMatrix()
print(type(cMat2))
| Jupyter/SIT742P07B-MLlib-DataType.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Lab 3 Tutorial: Model Selection in scikit-learn
# + hide_input=false slideshow={"slide_type": "skip"}
# General imports
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import openml as oml
from matplotlib import cm
# We can ignore ConvergenceWarnings for illustration purposes
import warnings
warnings.simplefilter(action="ignore", category=UserWarning)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Evaluation procedures
# ### Holdout
# The simplest procedure is [train_test_split](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html), which splits arrays or matrices into random train and test subsets.
# + hide_input=true slideshow={"slide_type": "-"}
from sklearn.datasets import make_blobs
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
# create a synthetic dataset
X, y = make_blobs(centers=2, random_state=0)
# split data and labels into a training and a test set
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
# Instantiate a model and fit it to the training set
model = LogisticRegression().fit(X_train, y_train)
# evaluate the model on the test set
print("Test set score: {:.2f}".format(model.score(X_test, y_test)))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Cross-validation
# - [cross_val_score](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_val_score.html?highlight=cross%20val%20score#sklearn.model_selection.cross_val_score)
# - `cv` parameter defines the kind of cross-validation splits, default is 5-fold CV
# - `scoring` defines the scoring metric. Also see below.
# - Returns list of all scores. Models are built internally, but not returned
# - [cross_validate](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_validate.html?highlight=cross%20validate#sklearn.model_selection.cross_validate)
# - Similar, but also returns the fit and test times, and allows multiple scoring metrics.
# + hide_input=true
from sklearn.model_selection import cross_val_score
from sklearn.datasets import load_iris
from sklearn.linear_model import LogisticRegression
iris = load_iris()
logreg = LogisticRegression()
scores = cross_val_score(logreg, iris.data, iris.target, cv=5)
print("Cross-validation scores: {}".format(scores))
print("Average cross-validation score: {:.2f}".format(scores.mean()))
print("Variance in cross-validation score: {:.4f}".format(np.var(scores)))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Custom CV splits
# - You can build folds manually with [KFold](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.KFold.html?highlight=kfold#sklearn.model_selection.KFold) or [StratifiedKFold](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.StratifiedKFold.html#sklearn.model_selection.StratifiedKFold)
# - randomizable (`shuffle` parameter)
# - [LeaveOneOut](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.LeaveOneOut.html?highlight=leave%20one%20out#sklearn.model_selection.LeaveOneOut) does leave-one-out cross-validation
# + hide_input=true
from sklearn.model_selection import KFold, StratifiedKFold
kfold = KFold(n_splits=5)
print("Cross-validation scores KFold(n_splits=5):\n{}".format(
cross_val_score(logreg, iris.data, iris.target, cv=kfold)))
skfold = StratifiedKFold(n_splits=5, shuffle=True)
print("Cross-validation scores StratifiedKFold(n_splits=5, shuffle=True):\n{}".format(
cross_val_score(logreg, iris.data, iris.target, cv=skfold)))
# -
from sklearn.model_selection import LeaveOneOut
loo = LeaveOneOut()
scores = cross_val_score(logreg, iris.data, iris.target, cv=loo)
print("Number of cv iterations: ", len(scores))
print("Mean accuracy: {:.2f}".format(scores.mean()))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Shuffle-split
# These shuffle the data before splitting it.
# - `ShuffleSplit` and `StratifiedShuffleSplit` (recommended for classification)
# - `train_size` and `test_size` can be absolute numbers or a percentage of the total dataset
# -
from sklearn.model_selection import ShuffleSplit, StratifiedShuffleSplit
shuffle_split = StratifiedShuffleSplit(test_size=.5, train_size=.5, n_splits=10)
scores = cross_val_score(logreg, iris.data, iris.target, cv=shuffle_split)
print("Cross-validation scores:\n{}".format(scores))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Grouped cross-validation
# - Add an array with group membership to `cross_val_scores`
# - Use `GroupKFold` with the number of groups as CV procedure
# -
from sklearn.model_selection import GroupKFold
# create synthetic dataset
X, y = make_blobs(n_samples=12, random_state=0)
# the first three samples belong to the same group, etc.
groups = [0, 0, 0, 1, 1, 1, 2, 2, 2, 3, 3, 3]
scores = cross_val_score(logreg, X, y, groups=groups, cv=GroupKFold(n_splits=4))
print("Cross-validation scores :\n{}".format(scores))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Evaluation Metrics
# + [markdown] slideshow={"slide_type": "slide"}
# ### Binary classification
# - [confusion_matrix](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.confusion_matrix.html?highlight=confusion%20matrix#sklearn.metrics.confusion_matrix) returns a matrix counting how many test examples are predicted correctly or 'confused' with other metrics.
# - [sklearn.metrics](https://scikit-learn.org/stable/modules/classes.html?highlight=metrics#module-sklearn.metrics) contains implementations many of the metrics discussed in class
# - They are all implemented so that 'higher is better'.
# - [accuracy_score](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.accuracy_score.html#sklearn.metrics.accuracy_score) computes accuracy explictly
# - [classification_report](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.classification_report.html) returns a table of binary measures, per class, and aggregated according to different aggregation functions.
# + hide_input=false
from sklearn.metrics import accuracy_score, confusion_matrix, classification_report, f1_score
from sklearn.model_selection import train_test_split
from sklearn.datasets import load_breast_cancer
from sklearn.linear_model import LogisticRegression
data = load_breast_cancer()
X_train, X_test, y_train, y_test = train_test_split(
data.data, data.target, stratify=data.target, random_state=0)
lr = LogisticRegression().fit(X_train, y_train)
y_pred = lr.predict(X_test)
print("confusion_matrix(y_test, y_pred): \n", confusion_matrix(y_test, y_pred))
print("accuracy_score(y_test, y_pred): ", accuracy_score(y_test, y_pred))
print("model.score(X_test, y_test): ", lr.score(X_test, y_test))
# + hide_input=false slideshow={"slide_type": "-"}
plt.rcParams['figure.dpi'] = 100
print(classification_report(y_test, lr.predict(X_test)))
# -
# You can explictly define the averaging function for class-level metrics
pred = lr.predict(X_test)
print("Micro average f1 score: {:.3f}".format(f1_score(y_test, pred, average="micro")))
print("Weighted average f1 score: {:.3f}".format(f1_score(y_test, pred, average="weighted")))
print("Macro average f1 score: {:.3f}".format(f1_score(y_test, pred, average="macro")))
# ### Probabilistic predictions
# To retrieve the uncertainty in the prediction, scikit-learn offers 2 functions. Often, both are available for every learner, but not always.
#
# - decision_function: returns floating point (-Inf,Inf) value for each prediction
# - predict_proba: returns probability [0,1] for each prediction
# + [markdown] slideshow={"slide_type": "slide"}
# You can also use these to compute any metric with non-standard thresholds
# + hide_input=false
print("Threshold -0.8")
y_pred_lower_threshold = lr.decision_function(X_test) > -.8
print(classification_report(y_test, y_pred_lower_threshold))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Uncertainty in multi-class classification
#
# - `decision_function` and `predict_proba` also work in the multiclass setting
# - always have shape (n_samples, n_classes)
# - Example on the Iris dataset, which has 3 classes:
# +
from sklearn.datasets import load_iris
iris = load_iris()
X_train2, X_test2, y_train2, y_test2 = train_test_split(
iris.data, iris.target, random_state=42)
lr2 = LogisticRegression()
lr2 = lr2.fit(X_train2, y_train2)
print("Decision function:\n{}".format(lr2.decision_function(X_test2)[:6, :]))
# show the first few entries of predict_proba
print("Predicted probabilities:\n{}".format(lr2.predict_proba(X_test2)[:6]))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Precision-Recall and ROC curves
#
# - [precision_recall_curve](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.precision_recall_curve.html?highlight=precision_recall_curve) returns all precision and recall values for all possible thresholds
# - [roc_curve](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.roc_curve.html?highlight=roc%20curve#sklearn.metrics.roc_curve) does the same for TPR and FPR.
# - The average precision score is returned by the `average_precision_score` measure
# - The area under the ROC curve is returned by the `roc_auc_score` measure
# - Don't use `auc` (this uses a less accurate trapezoidal rule)
# - Require a decision function or predict_proba.
#
# -
from sklearn.metrics import precision_recall_curve
precision, recall, thresholds = precision_recall_curve(
y_test, lr.decision_function(X_test))
# + hide_input=true
from sklearn.metrics import average_precision_score
ap_pp = average_precision_score(y_test, lr.predict_proba(X_test)[:, 1])
ap_df = average_precision_score(y_test, lr.decision_function(X_test))
print("Average precision of logreg: {:.3f}".format(ap_df))
# -
from sklearn.metrics import roc_auc_score
rf_auc = roc_auc_score(y_test, lr.predict_proba(X_test)[:, 1])
svc_auc = roc_auc_score(y_test, lr.decision_function(X_test))
print("AUC for Random Forest: {:.3f}".format(rf_auc))
print("AUC for SVC: {:.3f}".format(svc_auc))
# ### Multi-class prediction
# * Build C models, one for every class vs all others
# * Use micro-, macro-, or weighted averaging
print("Micro average f1 score: {:.3f}".format(f1_score(y_test, pred, average="micro")))
print("Weighted average f1 score: {:.3f}".format(f1_score(y_test, pred, average="weighted")))
print("Macro average f1 score: {:.3f}".format(f1_score(y_test, pred, average="macro")))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Using evaluation metrics in model selection
#
# - You typically want to use AUC or other relevant measures in `cross_val_score` and `GridSearchCV` instead of the default accuracy.
# - scikit-learn makes this easy through the `scoring` argument
# - But, you need to need to look the [mapping between the scorer and the metric](http://scikit-learn.org/stable/modules/model_evaluation.html#model-evaluation)
# + [markdown] slideshow={"slide_type": "slide"}
# 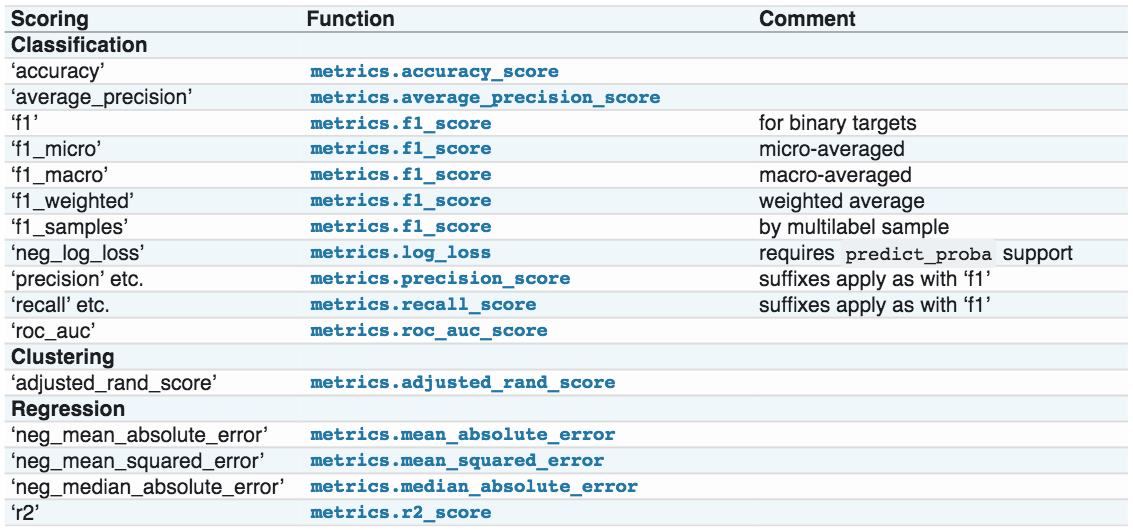
# + [markdown] slideshow={"slide_type": "slide"}
# Or simply look up like this:
# -
from sklearn.metrics import SCORERS
print("Available scorers:\n{}".format(sorted(SCORERS.keys())))
# + [markdown] slideshow={"slide_type": "slide"}
# Cross-validation with AUC
# +
from sklearn.model_selection import cross_val_score, GridSearchCV
from sklearn .svm import SVC
from sklearn.datasets import load_digits
digits = load_digits()
# default scoring for classification is accuracy
print("Default scoring: {}".format(
cross_val_score(SVC(), digits.data, digits.target == 9)))
# providing scoring="accuracy" doesn't change the results
explicit_accuracy = cross_val_score(SVC(), digits.data, digits.target == 9,
scoring="accuracy")
print("Explicit accuracy scoring: {}".format(explicit_accuracy))
roc_auc = cross_val_score(SVC(), digits.data, digits.target == 9,
scoring="roc_auc")
print("AUC scoring: {}".format(roc_auc))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Hyperparameter tuning
# Now that we know how to evaluate models, we can improve them by tuning their hyperparameters
# + [markdown] slideshow={"slide_type": "slide"}
# ### Grid search
# - Create a parameter grid as a dictionary
# - Keys are parameter names
# - Values are lists of hyperparameter values
# -
param_grid = {'C': [0.001, 0.01, 0.1, 1, 10, 100],
'gamma': [0.001, 0.01, 0.1, 1, 10, 100]}
print("Parameter grid:\n{}".format(param_grid))
# + [markdown] slideshow={"slide_type": "slide"}
# - `GridSearchCV`: like a classifier that uses CV to automatically optimize its hyperparameters internally
# - Input: (untrained) model, parameter grid, CV procedure
# - Output: optimized model on given training data
# - Should only have access to training data
# -
from sklearn.model_selection import GridSearchCV
from sklearn.svm import SVC
grid_search = GridSearchCV(SVC(), param_grid, cv=5)
X_train, X_test, y_train, y_test = train_test_split(
iris.data, iris.target, random_state=0)
grid_search.fit(X_train, y_train)
# + [markdown] slideshow={"slide_type": "slide"}
# The optimized test score and hyperparameters can easily be retrieved:
# -
print("Test set score: {:.2f}".format(grid_search.score(X_test, y_test)))
print("Best parameters: {}".format(grid_search.best_params_))
print("Best cross-validation score: {:.2f}".format(grid_search.best_score_))
print("Best estimator:\n{}".format(grid_search.best_estimator_))
# + [markdown] slideshow={"slide_type": "slide"}
# When hyperparameters depend on other parameters, we can use lists of dictionaries to define the hyperparameter space
# -
param_grid = [{'kernel': ['rbf'],
'C': [0.001, 0.01, 0.1, 1, 10, 100],
'gamma': [0.001, 0.01, 0.1, 1, 10, 100]},
{'kernel': ['linear'],
'C': [0.001, 0.01, 0.1, 1, 10, 100]}]
print("List of grids:\n{}".format(param_grid))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Nested cross-validation
#
# - Nested cross-validation:
# - Outer loop: split data in training and test sets
# - Inner loop: run grid search, splitting the training data into train and validation sets
# - Result is a just a list of scores
# - There will be multiple optimized models and hyperparameter settings (not returned)
# - To apply on future data, we need to train `GridSearchCV` on all data again
# + slideshow={"slide_type": "-"}
scores = cross_val_score(GridSearchCV(SVC(), param_grid, cv=5),
iris.data, iris.target, cv=5)
print("Cross-validation scores: ", scores)
print("Mean cross-validation score: ", scores.mean())
# + [markdown] slideshow={"slide_type": "slide"}
# ### Parallelizing cross-validation and grid-search
# - On a practical note, it is easy to parallellize CV and grid search
# - `cross_val_score` and `GridSearchCV` have a `n_jobs` parameter defining the number of cores it can use.
# - set it to `n_jobs=-1` to use all available cores.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Random Search
# - `RandomizedSearchCV` works like `GridSearchCV`
# - Has `n_iter` parameter for the number of iterations
# - Search grid can use distributions instead of fixed lists
# +
from sklearn.model_selection import RandomizedSearchCV
from scipy.stats import expon
param_grid = {'C': expon(scale=100),
'gamma': expon(scale=.1)}
random_search = RandomizedSearchCV(SVC(), param_distributions=param_grid,
n_iter=20)
X_train, X_test, y_train, y_test = train_test_split(
iris.data, iris.target, random_state=0)
random_search.fit(X_train, y_train)
| labs/Lab 3 - Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/DJCordhose/buch-machine-learning-notebooks/blob/master/kap7-iris.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="v4eRosoGpl8_" colab_type="text"
# # Kapitel 7: Neuronale Netzwerke - Grundlagen
# + id="KYbHMTMjpl8_" colab_type="code" colab={}
import warnings
warnings.filterwarnings('ignore')
# + id="83Ttu0Pnpl9D" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="85822c65-8551-41f3-9bf1-ea0dfc9291f6"
# %matplotlib inline
# %pylab inline
# + id="rJ1DvBlApl9G" colab_type="code" colab={}
import matplotlib.pylab as plt
import numpy as np
# + id="g0rvxIp4pl9J" colab_type="code" colab={}
colors = 'bwr'#['b','y','r']
CMAP = colors#plt.cm.rainbow
# + id="n5AGdJTFpl9N" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0b1b66a4-ee5b-4b89-c789-5ca2e27f31f4"
import sklearn
print(sklearn.__version__)
# + id="f5bwPx-ipl9P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="49d8eea1-8c56-439a-f5bc-a82f32e6b6aa"
import tensorflow as tf
print(tf.__version__)
# + id="ssBE6lbypl9S" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="92738183-eed5-40d3-ccf6-f0d3e4e810af"
import keras
print(keras.__version__)
# + id="_9-yW3Skpl9V" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="19437069-f13b-4c88-f89b-e4635850f0e2"
import pandas as pd
print(pd.__version__)
# + [markdown] id="lKGPNR5Vpl9Y" colab_type="text"
# ## Iris mit Neuronalen Netzwerken
# + id="5h7LeQWFpl9Y" colab_type="code" colab={}
from sklearn.datasets import load_iris
iris = load_iris()
# + id="FtNqwr6Gpl9a" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="0caf111b-fe2f-4113-987b-ef3bdaf81934"
print(iris.DESCR)
# + id="LEmm7Weipl9d" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="9fe728e2-2ea0-486c-e520-a3e958b7b3d4"
iris_df = pd.DataFrame(iris.data, columns=iris.feature_names)
pd.plotting.scatter_matrix(iris_df, c=iris.target, cmap=CMAP, edgecolor='black', figsize=(20, 20))
# + id="oa3iYStkpl9f" colab_type="code" colab={}
# plt.savefig('ML_0701.png', bbox_inches='tight')
# + [markdown] id="c-exG806pl9g" colab_type="text"
# ## Das künstliche Neuron
# + id="PrYfSutGpl9h" colab_type="code" colab={}
w0 = 3
w1 = -4
w2 = 2
def neuron_no_activation(x1, x2):
sum = w0 + x1 * w1 + x2 * w2
return sum
# + id="Yhw0DMUWpl9j" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="dcc8b20c-be1b-43b6-c60c-2c130cfd3c5b"
iris.data[0]
# + id="rVqDyYWKpl9l" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4208fc66-5604-4799-e353-65e52fea9e98"
neuron_no_activation(5.1, 3.5)
# + [markdown] id="TpcxmzDspl9o" colab_type="text"
# ### Activation Functions
# + id="wP80A7hkpl9o" colab_type="code" colab={}
def centerAxis(uses_negative=False):
# http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.plot
ax = plt.gca()
ax.spines['left'].set_position('center')
if uses_negative:
ax.spines['bottom'].set_position('center')
ax.spines['right'].set_color('none')
ax.spines['top'].set_color('none')
ax.xaxis.set_ticks_position('bottom')
ax.yaxis.set_ticks_position('left')
# + [markdown] id="yNq7ZiU0pl9r" colab_type="text"
# #### Step Function: abrupter, nicht stetig differenzierbarer Übergang zwischen 0 und 1
# + id="ZqUtjXQ1pl9r" colab_type="code" colab={}
def np_step(X):
return 0.5 * (np.sign(X) + 1)
# + id="BIBsOI4epl9t" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="f6676a2a-0eda-4b1f-b4ee-a9aab2298c2d"
x = np.arange(-10,10,0.01)
y = np_step(x)
centerAxis()
plt.plot(x, y, lw=3)
# + [markdown] id="trtAvxTmpl9w" colab_type="text"
# #### Sigmoid Function: Fließender Übergang zwischen 0 und 1
# + id="BN8Y7fXPpl9x" colab_type="code" colab={}
def np_sigmoid(X):
return 1 / (1 + np.exp(X * -1))
# + id="HyC7OoVnpl9z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="bc6f637c-c76a-4afd-89bc-2f27b6622948"
x = np.arange(-10,10,0.01)
y = np_sigmoid(x)
centerAxis()
plt.plot(x,y,lw=3)
# + [markdown] id="WAsTI6Chpl91" colab_type="text"
# #### Tangens Hyperbolicus Function: Fließender Übergang zwischen -1 und 1
# + id="_WDP-00Lpl92" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="b9014b7e-a164-4e50-bec4-76d1a0071941"
x = np.arange(-10,10,0.01)
y = np.tanh(x)
centerAxis()
plt.plot(x,y,lw=3)
# + [markdown] id="JWZW6pkrpl95" colab_type="text"
# #### Relu: Einfach zu berechnen, setzt kompletten negativen Wertebereich auf 0
# + id="6LTGgI99pl96" colab_type="code" colab={}
def np_relu(x):
return np.maximum(0, x)
# + id="mqtXbFQvpl98" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="b560795b-7df1-412f-a8b3-726a99006aff"
x = np.arange(-10,10,0.01)
y = np_relu(x)
centerAxis()
plt.plot(x,y,lw=3)
# + id="pDcTe_YHpl9-" colab_type="code" colab={}
# https://docs.python.org/3/library/math.html
import math as math
def sigmoid(x):
return 1 / (1 + math.exp(x * -1))
w0 = 3
w1 = -4
w2 = 2
def neuron(x1, x2):
sum = w0 + x1 * w1 + x2 * w2
return sigmoid(sum)
# + id="8jQTVmi6pl-A" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e7ce10f8-c0a0-43c5-971c-6dd870ee5649"
neuron(5.1, 3.5)
# + id="-Ci-Qwh3pl-C" colab_type="code" colab={}
# Version that takes as many values as you like
weights_with_bias = np.array([3, -4, 2])
def np_neuron(X):
inputs_with_1_for_bias = np.concatenate((np.array([1]), X))
return np_sigmoid(np.sum(inputs_with_1_for_bias*weights_with_bias));
# + id="_kIgFmAhpl-F" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="11c1ab58-7f3b-4ce0-818c-b4a5b069ffc8"
np_neuron(np.array([5.1, 3.5]))
# + [markdown] id="fcynLtuepl-H" colab_type="text"
# ## Unser erste Neuronales Netz mit Keras
# + id="Uv0NsPfhpl-H" colab_type="code" colab={}
from keras.layers import Input
inputs = Input(shape=(4, ))
# + id="8id5zq2Bpl-K" colab_type="code" colab={}
from keras.layers import Dense
fc = Dense(3)(inputs)
# + id="RiFhyYi3pl-L" colab_type="code" colab={}
from keras.models import Model
model = Model(inputs=inputs, outputs=fc)
# + id="wKyUqPAupl-N" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="29c0ce17-5d53-4174-cd6b-863383e59ae7"
model.summary()
# + id="lhrAh8GApl-P" colab_type="code" colab={}
model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
# + id="-WcVU5Rgpl-R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="42dd925d-cc8c-4594-cffb-ed5843f9f511"
model.predict(np.array([[ 5.1, 3.5, 1.4, 0.2]]))
# + id="i1OwT7TWpl-T" colab_type="code" colab={}
inputs = Input(shape=(4, ))
fc = Dense(3)(inputs)
predictions = Dense(3, activation='softmax')(fc)
model = Model(inputs=inputs, outputs=predictions)
# + id="KTxUA8jOpl-X" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 255} outputId="f404a16b-f8f0-417b-f768-87548a4328d1"
model.summary()
# + id="mGszqEMDpl-a" colab_type="code" colab={}
model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
# + id="MI_BdRKbpl-c" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a1b7dcd4-05ac-46bc-b813-bc3ea1f2f1d4"
model.predict(np.array([[ 5.1, 3.5, 1.4, 0.2]]))
# + [markdown] id="sdoL_Z7bpl-e" colab_type="text"
# # Training
# + id="fQ1oRcd-pl-e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e2597cc0-5451-48da-f469-42d0fe6884e3"
X = np.array(iris.data)
y = np.array(iris.target)
X.shape, y.shape
# + id="c4LEAqRlpl-h" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="db3f7986-2b35-45d7-b081-30475080d3a8"
y[100]
# + id="-2gMaXFGpl-m" colab_type="code" colab={}
from keras.utils.np_utils import to_categorical
num_categories = 3
y = to_categorical(y, num_categories)
# + id="_GdCZmogpl-o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9b966b62-bff3-4274-bb14-12b4dab702e7"
y[100]
# + id="TJL9v7tQpl-r" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20, random_state=42, stratify=y)
# + id="2W10jFK_pl-t" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="cf0f8104-5b4c-410d-fe9b-bfafde834311"
X_train.shape, X_test.shape, y_train.shape, y_test.shape
# + id="JNNf0b4kpl-w" colab_type="code" colab={}
# # !rm -r tf_log
# https://keras.io/callbacks/#tensorboard
# tb_callback = keras.callbacks.TensorBoard(log_dir='./tf_log')
# To start tensorboard
# tensorboard --logdir=/mnt/c/Users/olive/Development/ml/tf_log
# open http://localhost:6006
# + id="Px5unIxQpl-0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="aeaeb4c7-e800-4fa2-b98b-3bcaa0881dd8"
# # %time model.fit(X_train, y_train, epochs=500, validation_split=0.3, callbacks=[tb_callback])
# %time model.fit(X_train, y_train, epochs=500, validation_split=0.3)
# + [markdown] id="pmsuPaqYpl-2" colab_type="text"
# # Bewertung
# + id="iCCbIyrjpl-3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8279ee31-3dfc-4575-eb27-b0a83a909bb4"
model.predict(np.array([[ 5.1, 3.5, 1.4, 0.2]]))
# + id="jQaEAtyVpl-4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4bdab368-1515-4a61-e77b-399c5741bc2e"
X[0], y[0]
# + id="wrzGbtchpl-7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="5c1af142-3b53-4bd4-8bd0-7467ef8bf6d8"
train_loss, train_accuracy = model.evaluate(X_train, y_train)
train_loss, train_accuracy
# + id="hnXEggalpl-8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="50a0eed0-1d06-4a5b-9894-23c55f073178"
test_loss, test_accuracy = model.evaluate(X_test, y_test)
test_loss, test_accuracy
| kap7-iris.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Ruby 2.3.1
# language: ruby
# name: ruby
# ---
require 'daru/view'
Daru::View.plotting_library = :highcharts
# +
# pie-basic chart : pie basic
opts = {
chart: {
plotBackgroundColor: nil,
plotBorderWidth: nil,
plotShadow: false,
type: 'pie'
},
title: {
text: 'Browser market shares January, 2015 to May, 2015'
},
tooltip: {
pointFormat: '{series.name}: <b>{point.percentage:.1f}%</b>'
},
plotOptions: {
pie: {
allowPointSelect: true,
cursor: 'pointer',
dataLabels: {
enabled: true,
format: '<b>{point.name}</b>: {point.percentage:.1f} %',
style: {
color: "(Highcharts.theme && Highcharts.theme.contrastTextColor) || 'black'".js_code
}
}
}
},
}
series_dt = [
{
name: 'Brands',
colorByPoint: true,
data: [{
name: 'Microsoft Internet Explorer',
y: 56.33
}, {
name: 'Chrome',
y: 24.03,
sliced: true,
selected: true
}, {
name: 'Firefox',
y: 10.38
}, {
name: 'Safari',
y: 4.77
}, {
name: 'Opera',
y: 0.91
}, {
name: 'Proprietary or Undetectable',
y: 0.2
}]
}
]
pie_basic = Daru::View::Plot.new
pie_basic.chart.options = opts;
pie_basic.chart.series_data = series_dt
pie_basic.show_in_iruby
# +
# pie chart : pie legend
opts = {
chart: {
plotBackgroundColor: nil,
plotBorderWidth: nil,
plotShadow: false,
type: 'pie'
},
title: {
text: 'Browser market shares January, 2015 to May, 2015'
},
tooltip: {
pointFormat: '{series.name}: <b>{point.percentage:.1f}%</b>'
},
plotOptions: {
pie: {
allowPointSelect: true,
cursor: 'pointer',
dataLabels: {
enabled: false
},
showInLegend: true
}
},
}
series_dt = [
{
name: 'Brands',
colorByPoint: true,
data: [{
name: 'Microsoft Internet Explorer',
y: 56.33
}, {
name: 'Chrome',
y: 24.03,
sliced: true,
selected: true
}, {
name: 'Firefox',
y: 10.38
}, {
name: 'Safari',
y: 4.77
}, {
name: 'Opera',
y: 0.91
}, {
name: 'Proprietary or Undetectable',
y: 0.2
}]
}
]
pie_legend = Daru::View::Plot.new
pie_legend.chart.options = opts;
pie_legend.chart.series_data = series_dt
pie_legend.show_in_iruby
# -
#Todo: pie-basic chart : pie-donut/
# a lot of stuff. will do latter
# +
# pie chart : semi-circle/
opts = {
chart: {
plotBackgroundColor: nil,
plotBorderWidth: 0,
plotShadow: false
},
title: {
text: 'Browser<br>shares<br>2015',
align: 'center',
verticalAlign: 'middle',
y: 40
},
tooltip: {
pointFormat: '{series.name}: <b>{point.percentage:.1f}%</b>'
},
plotOptions: {
pie: {
dataLabels: {
enabled: true,
distance: -50,
style: {
fontWeight: 'bold',
color: 'white'
}
},
startAngle: -90,
endAngle: 90,
center: ['50%', '75%']
}
},
}
series_dt = [
{
type: 'pie',
name: 'Browser share',
innerSize: '50%',
data: [
['Firefox', 10.38],
['IE', 56.33],
['Chrome', 24.03],
['Safari', 4.77],
['Opera', 0.91],
{
name: 'Proprietary or Undetectable',
y: 0.2,
dataLabels: {
enabled: false
}
}
]
}
]
pie_semi_circle = Daru::View::Plot.new
pie_semi_circle.chart.options = opts;
pie_semi_circle.chart.series_data = series_dt
pie_semi_circle.show_in_iruby
# +
# note : drilldown.js is needed for it.
# pie-basic chart : pie drilldown
opts = {
chart: {
type: 'pie'
},
title: {
text: 'Browser market shares. January, 2015 to May, 2015'
},
subtitle: {
text: 'Click the slices to view versions. Source: netmarketshare.com.'
},
plotOptions: {
series: {
dataLabels: {
enabled: true,
format: '{point.name}: {point.y:.1f}%'
}
}
},
tooltip: {
headerFormat: '<span style="font-size:11px">{series.name}</span><br>',
pointFormat: '<span style="color:{point.color}">{point.name}</span>: <b>{point.y:.2f}%</b> of total<br/>'
},
drilldown: {
series: [{
name: 'Microsoft Internet Explorer',
id: 'Microsoft Internet Explorer',
data: [
['v11.0', 24.13],
['v8.0', 17.2],
['v9.0', 8.11],
['v10.0', 5.33],
['v6.0', 1.06],
['v7.0', 0.5]
]
}, {
name: 'Chrome',
id: 'Chrome',
data: [
['v40.0', 5],
['v41.0', 4.32],
['v42.0', 3.68],
['v39.0', 2.96],
['v36.0', 2.53],
['v43.0', 1.45],
['v31.0', 1.24],
['v35.0', 0.85],
['v38.0', 0.6],
['v32.0', 0.55],
['v37.0', 0.38],
['v33.0', 0.19],
['v34.0', 0.14],
['v30.0', 0.14]
]
}, {
name: 'Firefox',
id: 'Firefox',
data: [
['v35', 2.76],
['v36', 2.32],
['v37', 2.31],
['v34', 1.27],
['v38', 1.02],
['v31', 0.33],
['v33', 0.22],
['v32', 0.15]
]
}, {
name: 'Safari',
id: 'Safari',
data: [
['v8.0', 2.56],
['v7.1', 0.77],
['v5.1', 0.42],
['v5.0', 0.3],
['v6.1', 0.29],
['v7.0', 0.26],
['v6.2', 0.17]
]
}, {
name: 'Opera',
id: 'Opera',
data: [
['v12.x', 0.34],
['v28', 0.24],
['v27', 0.17],
['v29', 0.16]
]
}]
}
}
series_dt = [
{
name: 'Brands',
colorByPoint: true,
data: [{
name: 'Microsoft Internet Explorer',
y: 56.33,
drilldown: 'Microsoft Internet Explorer'
}, {
name: 'Chrome',
y: 24.03,
drilldown: 'Chrome'
}, {
name: 'Firefox',
y: 10.38,
drilldown: 'Firefox'
}, {
name: 'Safari',
y: 4.77,
drilldown: 'Safari'
}, {
name: 'Opera',
y: 0.91,
drilldown: 'Opera'
}, {
name: 'Proprietary or Undetectable',
y: 0.2,
drilldown: nil
}]
}
]
pie_drill = Daru::View::Plot.new
pie_drill.chart.options = opts;
pie_drill.chart.series_data = series_dt
pie_drill.show_in_iruby
# +
# pie chart : pie gradient
# script for color is not added.
opts = {
chart: {
plotBackgroundColor: nil,
plotBorderWidth: nil,
plotShadow: false,
type: 'pie'
},
title: {
text: 'Browser market shares. January, 2015 to May, 2015'
},
tooltip: {
pointFormat: '{series.name}: <b>{point.percentage:.1f}%</b>'
},
plotOptions: {
pie: {
allowPointSelect: true,
cursor: 'pointer',
dataLabels: {
enabled: true,
format: '<b>{point.name}</b>: {point.percentage:.1f} %',
style: {
color: "(Highcharts.theme && Highcharts.theme.contrastTextColor) || 'black'".js_code
},
connectorColor: 'silver'
}
}
},
exporting: {
sourceWidth: 400,
sourceHeight: 200,
scale: 2,
chartOptions: {
subtitle: nil
}
}
}
series_dt = [
{
name: 'Brands',
data: [
{ name: 'Microsoft Internet Explorer', y: 56.33 },
{
name: 'Chrome',
y: 24.03,
sliced: true,
selected: true
},
{ name: 'Firefox', y: 10.38 },
{ name: 'Safari', y: 4.77 }, { name: 'Opera', y: 0.91 },
{ name: 'Proprietary or Undetectable', y: 0.2 }
]
}
]
pie_gradient = Daru::View::Plot.new
pie_gradient.chart.options = opts;
pie_gradient.chart.series_data = series_dt
pie_gradient.show_in_iruby
# -
| spec/dummy_iruby/HighCharts- Pie charts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [**Blueprints for Text Analysis Using Python**](https://github.com/blueprints-for-text-analytics-python/blueprints-text)
# <NAME>, <NAME>, <NAME>
#
# **If you like the book or the code examples here, please leave a friendly comment on [Amazon.com](https://www.amazon.com/Blueprints-Text-Analytics-Using-Python/dp/149207408X)!**
# <img src="../rating.png" width="100"/>
#
# # Chapter 5:<div class='tocSkip'/>
# # Feature Engineering and Syntactic Similarity
# ## Remark<div class='tocSkip'/>
#
# The code in this notebook differs slightly from the printed book.
#
# Several layout and formatting commands, like `figsize` to control figure size or subplot commands are removed in the book.
#
# All of this is done to simplify the code in the book and put the focus on the important parts instead of formatting.
# ## Setup<div class='tocSkip'/>
#
# Set directory locations. If working on Google Colab: copy files and install required libraries.
# +
import sys, os
ON_COLAB = 'google.colab' in sys.modules
if ON_COLAB:
GIT_ROOT = 'https://github.com/blueprints-for-text-analytics-python/blueprints-text/raw/master'
os.system(f'wget {GIT_ROOT}/ch05/setup.py')
# %run -i setup.py
# -
# ## Load Python Settings<div class="tocSkip"/>
#
# Common imports, defaults for formatting in Matplotlib, Pandas etc.
# +
# %run "$BASE_DIR/settings.py"
# %reload_ext autoreload
# %autoreload 2
# %config InlineBackend.figure_format = 'png'
# -
# # Data preparation
# +
sentences = ["It was the best of times",
"it was the worst of times",
"it was the age of wisdom",
"it was the age of foolishness"]
tokenized_sentences = [[t for t in sentence.split()] for sentence in sentences]
vocabulary = set([w for s in tokenized_sentences for w in s])
import pandas as pd
[[w, i] for i,w in enumerate(vocabulary)]
# -
# # One-hot by hand
# +
def onehot_encode(tokenized_sentence):
return [1 if w in tokenized_sentence else 0 for w in vocabulary]
onehot = [onehot_encode(tokenized_sentence) for tokenized_sentence in tokenized_sentences]
for (sentence, oh) in zip(sentences, onehot):
print("%s: %s" % (oh, sentence))
# -
pd.DataFrame(onehot, columns=vocabulary)
sim = [onehot[0][i] & onehot[1][i] for i in range(0, len(vocabulary))]
sum(sim)
import numpy as np
np.dot(onehot[0], onehot[1])
np.dot(onehot, onehot[1])
# ## Out of vocabulary
onehot_encode("the age of wisdom is the best of times".split())
onehot_encode("John likes to watch movies. Mary likes movies too.".split())
# ## document term matrix
onehot
# ## similarities
import numpy as np
np.dot(onehot, np.transpose(onehot))
# # scikit learn one-hot vectorization
from sklearn.preprocessing import MultiLabelBinarizer
lb = MultiLabelBinarizer()
lb.fit([vocabulary])
lb.transform(tokenized_sentences)
# # CountVectorizer
from sklearn.feature_extraction.text import CountVectorizer
cv = CountVectorizer()
more_sentences = sentences + ["John likes to watch movies. Mary likes movies too.",
"Mary also likes to watch football games."]
pd.DataFrame(more_sentences)
cv.fit(more_sentences)
print(cv.get_feature_names())
dt = cv.transform(more_sentences)
dt
pd.DataFrame(dt.toarray(), columns=cv.get_feature_names())
from sklearn.metrics.pairwise import cosine_similarity
cosine_similarity(dt[0], dt[1])
len(more_sentences)
pd.DataFrame(cosine_similarity(dt, dt))
# # TF/IDF
from sklearn.feature_extraction.text import TfidfTransformer
tfidf = TfidfTransformer()
tfidf_dt = tfidf.fit_transform(dt)
pd.DataFrame(tfidf_dt.toarray(), columns=cv.get_feature_names())
pd.DataFrame(cosine_similarity(tfidf_dt, tfidf_dt))
headlines = pd.read_csv(ABCNEWS_FILE, parse_dates=["publish_date"])
headlines.head()
from sklearn.feature_extraction.text import TfidfVectorizer
tfidf = TfidfVectorizer()
dt = tfidf.fit_transform(headlines["headline_text"])
dt
dt.data.nbytes
# %%time
cosine_similarity(dt[0:10000], dt[0:10000])
# ## Stopwords
from spacy.lang.en.stop_words import STOP_WORDS as stopwords
print(len(stopwords))
tfidf = TfidfVectorizer(stop_words=stopwords)
dt = tfidf.fit_transform(headlines["headline_text"])
dt
# ## min_df
tfidf = TfidfVectorizer(stop_words=stopwords, min_df=2)
dt = tfidf.fit_transform(headlines["headline_text"])
dt
tfidf = TfidfVectorizer(stop_words=stopwords, min_df=.0001)
dt = tfidf.fit_transform(headlines["headline_text"])
dt
# ## max_df
tfidf = TfidfVectorizer(stop_words=stopwords, max_df=0.1)
dt = tfidf.fit_transform(headlines["headline_text"])
dt
tfidf = TfidfVectorizer(max_df=0.1)
dt = tfidf.fit_transform(headlines["headline_text"])
dt
# ## n-grams
tfidf = TfidfVectorizer(stop_words=stopwords, ngram_range=(1,2), min_df=2)
dt = tfidf.fit_transform(headlines["headline_text"])
print(dt.shape)
print(dt.data.nbytes)
tfidf = TfidfVectorizer(stop_words=stopwords, ngram_range=(1,3), min_df=2)
dt = tfidf.fit_transform(headlines["headline_text"])
print(dt.shape)
print(dt.data.nbytes)
# ## Lemmas
from tqdm.auto import tqdm
import spacy
nlp = spacy.load("en")
nouns_adjectives_verbs = ["NOUN", "PROPN", "ADJ", "ADV", "VERB"]
for i, row in tqdm(headlines.iterrows(), total=len(headlines)):
doc = nlp(str(row["headline_text"]))
headlines.at[i, "lemmas"] = " ".join([token.lemma_ for token in doc])
headlines.at[i, "nav"] = " ".join([token.lemma_ for token in doc if token.pos_ in nouns_adjectives_verbs])
headlines.head()
tfidf = TfidfVectorizer(stop_words=stopwords)
dt = tfidf.fit_transform(headlines["lemmas"].map(str))
dt
tfidf = TfidfVectorizer(stop_words=stopwords)
dt = tfidf.fit_transform(headlines["nav"].map(str))
dt
# ## remove top 10,000
top_10000 = pd.read_csv("https://raw.githubusercontent.com/first20hours/google-10000-english/master/google-10000-english.txt", header=None)
tfidf = TfidfVectorizer(stop_words=set(top_10000.iloc[:,0].values))
dt = tfidf.fit_transform(headlines["nav"].map(str))
dt
tfidf = TfidfVectorizer(ngram_range=(1,2), stop_words=set(top_10000.iloc[:,0].values), min_df=2)
dt = tfidf.fit_transform(headlines["nav"].map(str))
dt
# ## Finding document most similar to made-up document
tfidf = TfidfVectorizer(stop_words=stopwords, min_df=2)
dt = tfidf.fit_transform(headlines["lemmas"].map(str))
dt
made_up = tfidf.transform(["australia and new zealand discuss optimal apple size"])
sim = cosine_similarity(made_up, dt)
sim[0]
headlines.iloc[np.argsort(sim[0])[::-1][0:5]][["publish_date", "lemmas"]]
# # Finding the most similar documents
# there are "test" headlines in the corpus
stopwords.add("test")
tfidf = TfidfVectorizer(stop_words=stopwords, ngram_range=(1,2), min_df=2, norm='l2')
dt = tfidf.fit_transform(headlines["headline_text"])
# ### Timing Cosine Similarity
# %%time
cosine_similarity(dt[0:10000], dt[0:10000], dense_output=False)
# %%time
r = cosine_similarity(dt[0:10000], dt[0:10000])
r[r > 0.9999] = 0
print(np.argmax(r))
# %%time
r = cosine_similarity(dt[0:10000], dt[0:10000], dense_output=False)
r[r > 0.9999] = 0
print(np.argmax(r))
# ### Timing Dot-Product
# %%time
r = np.dot(dt[0:10000], np.transpose(dt[0:10000]))
r[r > 0.9999] = 0
print(np.argmax(r))
# ## Batch
# %%time
batch = 10000
max_sim = 0.0
max_a = None
max_b = None
for a in range(0, dt.shape[0], batch):
for b in range(0, a+batch, batch):
print(a, b)
#r = np.dot(dt[a:a+batch], np.transpose(dt[b:b+batch]))
r = cosine_similarity(dt[a:a+batch], dt[b:b+batch], dense_output=False)
# eliminate identical vectors
# by setting their similarity to np.nan which gets sorted out
r[r > 0.9999] = 0
sim = r.max()
if sim > max_sim:
# argmax returns a single value which we have to
# map to the two dimensions
(max_a, max_b) = np.unravel_index(np.argmax(r), r.shape)
# adjust offsets in corpus (this is a submatrix)
max_a += a
max_b += b
max_sim = sim
print(max_a, max_b)
print(max_sim)
pd.set_option('max_colwidth', -1)
headlines.iloc[[max_a, max_b]][["publish_date", "headline_text"]]
# # Finding most related words
tfidf_word = TfidfVectorizer(stop_words=stopwords, min_df=1000)
dt_word = tfidf_word.fit_transform(headlines["headline_text"])
r = cosine_similarity(dt_word.T, dt_word.T)
np.fill_diagonal(r, 0)
voc = tfidf_word.get_feature_names()
size = r.shape[0] # quadratic
for index in np.argsort(r.flatten())[::-1][0:40]:
a = int(index/size)
b = index%size
if a > b: # avoid repetitions
print('"%s" related to "%s"' % (voc[a], voc[b]))
| ch05/Feature_Engineering_Similarity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: dev
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Update sklearn to prevent version mismatches
# #!pip install sklearn --upgrade
# +
# install joblib. This will be used to save your model.
# Restart your kernel after installing
# #!pip install joblib
# -
import pandas as pd
# # Read the CSV and Perform Basic Data Cleaning
df = pd.read_csv("exoplanet_data.csv")
# Drop the null columns where all values are null
df = df.dropna(axis='columns', how='all')
# Drop the null rows
df = df.dropna()
df
# # Select your features (columns)
df.columns
# Set features. This will also be used as your x values.
X = df.drop("koi_disposition", axis=1)
X.head()
# # Create a Train Test Split
#
# Use `koi_disposition` for the y values
# +
from sklearn.preprocessing import LabelEncoder
label_encoder = LabelEncoder()
label_encoder.fit(df["koi_disposition"])
label_encoder.classes_
y = label_encoder.transform(df["koi_disposition"])
y
# -
# Split data into training and testing
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,y)
X_train.head()
# # Pre-processing
#
# Scale the data using the MinMaxScaler and perform some feature selection
# # Train the Model
#
#
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier(random_state=1)
rf.fit(X_train,y_train)
import numpy as np
np.array(y_test)
# +
predictions = rf.predict(X_test)
for x in range(0,5,1):
print(f"Predictions: {predictions[x]}, Actual: {y_test[x]}")
# -
print(f"Training Data Score: {rf.score(X_train, y_train)}")
print(f"Testing Data Score: {rf.score(X_test, y_test)}")
# # Hyperparameter Tuning
#
# Use `GridSearchCV` to tune the model's parameters
# +
# Create the GridSearchCV model
from sklearn.model_selection import GridSearchCV
param_grid = {
'n_estimators': [200,500],
'max_features': ['auto', 'sqrt', 'log2'],
'max_depth' : [5,8,10],
'criterion' :['gini', 'entropy']
}
grid_rf = GridSearchCV(rf, param_grid, verbose=3)
# + tags=[]
# Train the model with GridSearch
grid_rf.fit(X_train, y_train)
# -
print(grid_rf.best_params_)
print(grid_rf.best_score_)
# Make predictions with the hypertuned model
predictions = grid_rf.predict(X_test)
predictions
# Calculate classification report
from sklearn.metrics import classification_report
print(classification_report(y_test, predictions,
target_names =["CONFIRMED", "FALSE POSITIVE", "CANDIDATE"]))
# # Save the Model
# save your model by updating "your_name" with your name
# and "your_model" with your model variable
# be sure to turn this in to BCS
# if joblib fails to import, try running the command to install in terminal/git-bash
import joblib
filename = 'Models/randomForestClassifier.sav'
joblib.dump(grid_rf, filename)
| starter_code/.ipynb_checkpoints/Random Forest-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pyspark DataFrame API
# ## Contents
# 1 - DataFrame.show()
# 2 - StructType & StructField & Schema
# 3 - Column Operators
#
# 1 - alias(*alias, *kwargs)
# 2 - when(condition, value) / otherwise(value)
# 3 - between(lowerBound, upperBound)
# 4 - contains()
# 5 - startswith() / endswith()
# 6 - isNotNull() / isNull()
# 7 - isin(*cols)
# 8 - like()
# 9 - asc() & desc() – Sort()
# 10 - cast() & astype()
# 11 - between()
# 12 - contains()
# 13 - startswith() & endswith()
# 14 - isNull & isNotNull()
# 15 - like() & rlike()
# 16 - when() & otherwise()
# 17 - isin()
# 18 - Where() - Filter()
# 19 - na.drop()
# 20 - Collect()
# 21 - select()
# 22 - describe()
# 23 - withColumn()
# 24 - lit()
# 25 - withColumnRenamed()
# 26 - drop()
# 27 - distinct() / dropDuplicates()
# 28 - orderBy() and sort()
# 29 - groupBy()
# 1-count()
# 2-mean()
# 3-max()
# 4-min()
# 5-sum()
# 6-avg()
# 30 - agg()
# 5 - Join Types | Join Two DataFrames
# 6 - Union()
# # Import Libraries
from pyspark.sql import SparkSession
from pyspark.sql import functions as f
from pyspark.sql.functions import *
# # Build SparkSession
spark = SparkSession.builder \
.master("local[*]") \
.appName("ml") \
.config("spark.executor.memory","8g") \
.config("spark.driver.memory","8g") \
.getOrCreate()
# # Create DataFrame
adult_train_df = spark.read \
.option("header", True) \
.option("inferSchema", True) \
.option("sep", ",") \
.csv("/home/jovyan/work/archive/adult/adult.data")
# ## .show()
# PySpark DataFrame show() is used to display the contents of the DataFrame in a Table Row & Column Format. By default, it shows only 20 Rows, and the column values are truncated at 20 characters.
adult_train_df.show(4)
# ## .toPandas() & .limit()
# PySpark DataFrame provides a method toPandas() to convert it Python Pandas DataFrame.
#
# toPandas() results in the collection of all records in the PySpark DataFrame to the driver program and should be done on a small subset of the data. running on larger dataset’s results in memory error and crashes the application.
adult_train_df.limit(4).toPandas()
# ## .columns
# To get dataframe columns's names in a list
adult_train_df.columns
# ## .printSchema()
# To look at dataframe's schema
adult_train_df.printSchema()
# ## Define dataframe's schema
schema="age INT, workclass STRING, fnlwgt DOUBLE, education STRING, education_num DOUBLE, marital_status STRING, occupation STRING, relationship STRING, race STRING, sex STRING, capital_gain DOUBLE, capital_loss DOUBLE, hours_per_week DOUBLE, native_country STRING, output STRING"
#schema="polarity FLOAT, id LONG, date_time STRING, query STRING, user STRING, text STRING"
adult_test_df = spark.read.csv(
path="/home/jovyan/work/archive/adult/adult.test",
sep=",",
header=True,
quote='"',
schema=schema)
# ## .select()
# We use the .select() function to select the column we want.
#from pyspark.sql.functions import col
#df.select(col("gender")).show()
#df.select(df["relationship"]).show()
adult_test_df.select(adult_test_df.relationship).show(5)
adult_test_df.select(adult_test_df.columns[2:5]).show(3)
# ## .describe()
# Descriptive Statistic
adult_test_df.describe(["age", "fnlwgt", "education_num", "capital_gain",
"capital_loss", "hours_per_week"]).toPandas()
# ## Column Operators
#df.select(df.col1 + df.col2).show()
#df.select(df.col1 * df.col2).show()
#df.select(df.col1 / df.col2).show()
#df.select(df.col1 % df.col2).show()
#df.select(df.col2 > df.col3).show()
#df.select(df.col2 < df.col3).show()
#df.select(df.col2 == df.col3).show()
adult_test_df.select( adult_test_df.hours_per_week - adult_test_df.education_num).show(5)
# ## .sort() & orderBy()
# Sort the DataFrame columns by Ascending or Descending order.
# adult_test_df.select("age").orderBy(adult_test_df.age.desc()).show(5)
adult_test_df.select("age").sort(adult_test_df.age.desc()).show(5)
# ## .cast() & astype()
# To convert the data Type
adult_test_df.select(adult_test_df.capital_gain.cast("int")).printSchema()
# ## .union()
# Dataframe union() – union() method of the DataFrame is used to combine two DataFrame’s of the same structure/schema. If schemas are not the same it returns an error.
adult_whole_df = adult_train_df.union(adult_test_df)
# ## withColumn()
# withColumn() is a transformation function of DataFrame which is used to change the value, convert the datatype of an existing column, create a new column, and many more.
adult_whole_df.withColumn("capital_loss_multiply",col("capital_loss")*100).show(4)
# ## .trim()
# PySpark you can remove whitespaces or trim by using pyspark.sql.functions.trim() SQL functions.
adult_whole_df = adult_whole_df \
.withColumn("workclass", f.trim(f.col("workclass"))) \
.withColumn("education", f.trim(f.col("education"))) \
.withColumn("marital_status", f.trim(f.col("marital_status"))) \
.withColumn("occupation", f.trim(f.col("occupation"))) \
.withColumn("relationship", f.trim(f.col("relationship"))) \
.withColumn("race", f.trim(f.col("race"))) \
.withColumn("sex", f.trim(f.col("sex"))) \
.withColumn("native_country", f.trim(f.col("native_country"))) \
.withColumn("output", f.trim(f.col("output")))
# ## Where & Filter
# filter() function is used to filter the rows from DataFrame based on the given condition or SQL expression, you can also use where() clause instead of the filter() if you are coming from an SQL background, both these functions operate exactly the same.
adult_whole_df.filter(adult_whole_df.race != "White") \
.select("race") \
.show(5,truncate=False)
adult_whole_df.filter(~((adult_whole_df.marital_status == "Never-married")
& (adult_whole_df.sex == "Male")
& (adult_whole_df.relationship == "Not-in-family")
& (adult_whole_df.hours_per_week >= 50.0)
| (adult_whole_df.age <= 40))) \
.limit(5).toPandas()
# ## .like()
adult_whole_df.select(adult_whole_df.workclass) \
.filter(adult_whole_df.workclass.like("%va%")).show(5)
# ## .isNull & .isNotNull()
# Checks if the DataFrame column has NULL or non NULL values.
#adult_test_df.filter(adult_test_df.occupation.isNull()).show()
adult_whole_df.filter(adult_whole_df.occupation.isNotNull()).limit(5).toPandas()
# ## .contains()
# Checks if a DataFrame column value contains a a value specified in this function.
adult_whole_df.select("marital_status").filter(adult_whole_df.marital_status.contains("Divorced")).show(5)
# ## .between()
# Returns a Boolean expression when a column values in between lower and upper bound.
adult_whole_df.select("age").filter(adult_test_df.age.between(60,65)).show(5)
# ## .isin()
# Check if value presents in a List.
other=["Other","Amer-Indian-Eskimo", "Asian-Pac-Islander"]
adult_whole_df.select(adult_whole_df.race) \
.filter(adult_whole_df.race.isin(other)) \
.show(5)
# ## . groupBy()
# Similar to SQL GROUP BY clause,
# groupBy() function is used to collect the identical data into groups on DataFrame
# and perform aggregate functions on the grouped data.
adult_whole_df.groupBy(f.col("marital_status")).agg({"*":"count"}) \
.toPandas()
adult_whole_df.groupBy(f.col("workclass")).agg({"*":"count"}) \
.toPandas()
train_df[["Sex", "Survived"]].groupby(["Sex"], as_index=False).mean().sort_values(by = "Survived", ascending=False)
train_df[["SibSp", "Survived"]].groupby(["SibSp"], as_index=False).mean().sort_values(by = "Survived", ascending=False)
adult_whole_df.filter(adult_whole_df.race != "White").count()
def get_columns_which_have_sought_value(dataframe:dataframe.DataFrame,
sought_value:object):
list_of_cols = []
for col in dataframe.columns:
if dataframe.filter(f.col(col).contains(sought_value)).count() > 0:
list_of_cols.append(col)
return list_of_cols
cols = show_columns_which_have_question_values(adult_whole_df)
cols
def change_values_in_columns(dataframe:dataframe.DataFrame,
columns:list[str],
value_to_change:object,
value_to_assign=None):
new_dataframe = dataframe
for col in columns:
new_dataframe = new_dataframe \
.withColumn(col, f.regexp_replace(f.col(col), value_to_change, value_to_assign))
return new_dataframe
for col in cols:
print(col)
df = change_values_in_columns(adult_whole_df, cols, '?')
# +
#adult_whole_df11 = adult_whole_df.withColumn(col, regexp_replace(f.col(col), '?', "None"))
# -
adult_whole_df13 = adult_whole_df \
.withColumn('workclass', f.regexp_replace(f.col('workclass'), '?', None))
adult_whole_df14.filter(f.col('occupation').contains("?")).count()
adult_whole_df.filter(f.col('workclass').contains("?")).count()
adult_whole_df.groupBy("workclass").sum("hours_per_week").show(truncate=False)
adult_whole_df.groupBy("workclass","marital_status") \
.avg("hours_per_week","age").show(10)
adult_whole_df.groupBy("race") \
.agg(sum("capital_gain").alias("sum_capital_gain"), \
avg("hours_per_week").alias("avg_hours"), \
min("education_num").alias("min_education_num"), \
max("age").alias("max_age"), \
avg("age").alias("avg_age"),
stddev("capital_gain")
).where(col("avg_age")<=38) \
.toPandas()
adult_whole_df.groupBy("race").count().show()
# ## Correlation Analysis
adult_whole_df.corr("age", "fnlwgt")
for col in adult_whole_df.columns:
adult_whole_df.corr()
# ## .when() & otherwise()
# It is similar to SQL Case When, executes sequence of expressions until it matches the condition and returns a value when match.
# AND (&) OR(|) NOT(!)
adult_whole_df.select(adult_whole_df.sex, when(adult_whole_df.sex=="Male", 1) \
.when(adult_whole_df.sex=="Female", 0) \
.otherwise(adult_whole_df.sex).alias("new_gender") \
).show(10)
adult_whole_df = adult_whole_df \
.withColumn("education_merged",when(f.col("education") \
.isin("1st-4th","5th-6th","7th-8th"),"Elementary-School") \
.when(f.col("education").isin("9th","10th","11th", "12th"),"High-School") \
.when(f.col("education").isin("Masters","Doctorate"),"High-Education") \
.when(f.col("education").isin("Bachelors","Some-college"),"Under-Education") \
.otherwise(col("education")))
adult_whole_df.select("education_merged", "education").show(5)
# +
adult_whole_df = adult_whole_df.withColumn("marital_status_merged",
when((col("marital_status") == "Separated")
| (col("marital_status") == "Widowed")
| (col("marital_status") == "Divorced")
| (col("marital_status") == "Never-married"), "Single")
.otherwise("Married"))
adult_whole_df.select("marital_status_merged").show(5)
# -
adult_whole_df.groupBy("marital_status_merged").agg({"*":"count"}).show()
# ## .drop()
# To drop dataframe's columns
adult_whole_df = adult_whole_df.drop("education")
# ## .na.drop()
# To drop null values
adult_whole_df = adult_whole_df.na.drop()
# ## withColumnRenamed()
# Rename Column Name withColumnRenamed() funcsion
adult_whole_df = adult_whole_df.withColumnRenamed("sex", "gender") \
.withColumnRenamed("output", "label")
adult_whole_df.limit(5).toPandas()
# ## .distinct()
adult_whole_df.select("race").distinct().count()
adult_whole_df.select("race").distinct().show()
# ## Detect Null Values
# #### We need to write a function to look up the number of null values in the columns
def show_columns_which_have_null_values(dataframe):
index = 0
for col in dataframe.columns:
if dataframe.filter(f.col(col).isNull()).count() > 0:
print(index, ".", col, " : There ara Null values")
index += 1
show_columns_which_have_null_values(adult_whole_df) # we don't have columns with missing values
def get_null_values_as_pandas_dataframe(dataframe):
data = dict()
for sutun in adult_whole_df.columns:
data.update({f"{sutun}" : adult_whole_df.filter(f.col(sutun).isNull()).count()})
s1 = pd.Series(list(data.keys()))
s2 = pd.Series(list(data.values()))
df = pd.concat([s1, s2], axis=1)
df = df.rename(columns={0:"Columns", 1:"Null_values"})
return df
df = show_null_values_as_pandas_dataframe(adult_whole_df)
df
# ## Aggregate Functions
#print("avg: " + str(adult_whole_df.select(avg("hours_per_week")).collect()[0][0]))
#print("count: "+str(adult_whole_df.select(count("hours_per_week")).collect()[0]))
#print(adult_whole_df.select(skewness("hours_per_week")).show())
#print(adult_whole_df.select(stddev("hours_per_week")).show())
#print(adult_whole_df.select(sum("hours_per_week")).show())
#print(adult_whole_df.select(max("hours_per_week")).show())
#print(adult_whole_df.select(min("hours_per_week")).show())
print(adult_whole_df.select(mean("hours_per_week")).show())
# ## fillna() & fill()
# DataFrame.fillna() or DataFrameNaFunctions.fill() is used to replace NULL/None values on all or selected multiple DataFrame columns with either zero(0), empty string, space, or any constant literal values.
#Replace Replace 0 for null on only population column
df.na.fill(value=0,subset=["population"]).show()
# ## sample()
# PySpark sampling (pyspark.sql.DataFrame.sample()) is a mechanism to get random sample records from the dataset, this is helpful when you have a larger dataset and wanted to analyze/test a subset of the data for example 10% of the original file.
adult_random_sample_df = adult_whole_df.sample(fraction=0.2, seed=1234)
print("sample count : ",adult_random_sample_df.count())
print("original count : ", adult_whole_df.count())
| Spark-DataFrame Api.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] cell_style="center" slideshow={"slide_type": "slide"}
# # Objects
# + [markdown] slideshow={"slide_type": "-"} tags=["remove-cell"]
# **CS1302 Introduction to Computer Programming**
# ___
# + slideshow={"slide_type": "fragment"} tags=["remove-cell"]
from manim import *
# %reload_ext mytutor
# + [markdown] slideshow={"slide_type": "fragment"}
# **Why object-oriented programming?**
# -
# %%manim -ql --progress_bar=none --disable_caching --flush_cache -v ERROR HelloWorld
class HelloWorld(Scene):
def construct(self):
self.play(Write(Text("Hello, World!")))
# The above code defines
# - `HelloWorld` as a `Scene`
# - `construct`ed by
# - `play`ing an animation that `Write`
# - the `Text` message `'Hello, World!'`.
# + [markdown] slideshow={"slide_type": "fragment"}
# **Exercise** Try changing
# - Mobjects: `Text('Hello, World!')` to `MathTex(r'\int tan(x)\,dx = -\ln(\cos(x))')` or `Circle()` or `Square()`.
# - Animation objects: `Write` to `FadeIn` or `GrowFromCenter`.
#
# See the [documentation](https://docs.manim.community/) and [tutorial](https://talkingphysics.wordpress.com/2019/01/08/getting-started-animating-with-manim-and-python-3-7/) for other choices.
# + [markdown] slideshow={"slide_type": "fragment"}
# More complicated behavior can be achieved by using different objects.
# + code_folding=[0] slideshow={"slide_type": "-"} language="html"
# <iframe width="800" height="450" src="https://www.youtube.com/embed/ENMyFGmq5OA" frameborder="0" allow="accelerometer; autoplay; clipboard-write; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>
# + [markdown] slideshow={"slide_type": "subslide"}
# **What is an object?**
# + [markdown] slideshow={"slide_type": "fragment"}
# Almost everything is an [`object`](https://docs.python.org/3/library/functions.html?highlight=object#object) in Python.
# + slideshow={"slide_type": "-"}
# isinstance?
isinstance(1, object), isinstance(1.0, object), isinstance("1", object)
# + [markdown] slideshow={"slide_type": "fragment"}
# A function is also a [first-class](https://en.wikipedia.org/wiki/First-class_function) object object.
# + slideshow={"slide_type": "-"}
isinstance(print, object), isinstance("".isdigit, object)
# + [markdown] slideshow={"slide_type": "fragment"}
# A data type is also an object.
# + slideshow={"slide_type": "-"}
# chicken and egg relationship
isinstance(type, object), isinstance(object, type), isinstance(object, object)
# + [markdown] slideshow={"slide_type": "subslide"}
# Python is a [*class-based* object-oriented programming](https://en.wikipedia.org/wiki/Object-oriented_programming#Class-based_vs_prototype-based) language:
# - Each object is an instance of a *class* (also called type in Python).
# - An object is a collection of *members/attributes*, each of which is an object.
# + slideshow={"slide_type": "-"}
# hasattr?
hasattr(str, "isdigit")
# + [markdown] slideshow={"slide_type": "fragment"}
# Different objects of a class
# - have the same set of attributes as that of the class, but
# - the attribute values can be different.
# + slideshow={"slide_type": "-"}
# dir?
dir(1) == dir(int), complex(1, 2).imag != complex(1, 1).imag
# + [markdown] slideshow={"slide_type": "subslide"}
# **How to operate on an object?**
# + [markdown] slideshow={"slide_type": "fragment"}
# - A class can define a function as an attribute for all its instances.
# - Such a function is called a *method* or *member function*.
# + slideshow={"slide_type": "fragment"}
complex.conjugate(complex(1, 2)), type(complex.conjugate)
# + [markdown] slideshow={"slide_type": "fragment"}
# A [method](https://docs.python.org/3/tutorial/classes.html#method-objects) can be accessed by objects of the class:
# + slideshow={"slide_type": "-"}
complex(1, 2).conjugate(), type(complex(1, 2).conjugate)
# + [markdown] slideshow={"slide_type": "fragment"}
# `complex(1,2).conjugate` is a *callable* object:
# - Its attribute `__self__` is assigned to `complex(1,2)`.
# - When called, it passes `__self__` as the first argument to `complex.conjugate`.
# + slideshow={"slide_type": "-"}
callable(complex(1, 2).conjugate), complex(1, 2).conjugate.__self__
# + [markdown] slideshow={"slide_type": "slide"}
# ## File Objects
# + [markdown] slideshow={"slide_type": "subslide"}
# **How to read a text file?**
# + [markdown] slideshow={"slide_type": "fragment"}
# Consider reading a csv (comma separated value) file:
# + slideshow={"slide_type": "-"}
# !more 'contact.csv'
# + [markdown] slideshow={"slide_type": "fragment"}
# To read the file by a Python program:
# + slideshow={"slide_type": "-"}
f = open("contact.csv") # create a file object for reading
print(f.read()) # return the entire content
f.close() # close the file
# + [markdown] slideshow={"slide_type": "fragment"}
# 1. [`open`](https://docs.python.org/3/library/functions.html?highlight=open#open) is a function that creates a file object and assigns it to `f`.
# 1. Associated with the file object:
# - [`read`](https://docs.python.org/3/library/io.html#io.TextIOBase.read) returns the entire content of the file as a string.
# - [`close`](https://docs.python.org/3/library/io.html#io.IOBase.close) flushes and closes the file.
# + [markdown] slideshow={"slide_type": "subslide"}
# **Why close a file?**
# + [markdown] slideshow={"slide_type": "fragment"}
# If not, depending on the operating system,
# - other programs may not be able to access the file, and
# - changes may not be written to the file.
# + [markdown] slideshow={"slide_type": "subslide"}
# To ensure a file is closed properly, we can use the [`with` statement](https://docs.python.org/3/reference/compound_stmts.html#with):
# + slideshow={"slide_type": "fragment"}
with open("contact.csv") as f:
print(f.read())
# + [markdown] slideshow={"slide_type": "subslide"}
# The `with` statement applies to any [context manager](https://docs.python.org/3/reference/datamodel.html#context-managers) that provides the methods
# - `__enter__` for initialization, and
# - `__exit__` for finalization.
# + slideshow={"slide_type": "fragment"}
with open("contact.csv") as f:
print(f, hasattr(f, "__enter__"), hasattr(f, "__exit__"), sep="\n")
# + [markdown] slideshow={"slide_type": "fragment"}
# - `f.__enter__` is called after the file object is successfully created and assigned to `f`, and
# - `f.__exit__` is called at the end, which closes the file.
# - `f.closed` indicates whether the file is closed.
# + slideshow={"slide_type": "fragment"}
f.closed
# + [markdown] slideshow={"slide_type": "fragment"}
# We can iterate a file object in a for loop,
# which implicitly call the method `__iter__` to read a file line by line.
# + slideshow={"slide_type": "fragment"}
with open("contact.csv") as f:
for line in f:
print(line, end="")
hasattr(f, "__iter__")
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercise** Print only the first 5 lines of the file `contact.csv`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "484ee06e7a39a307fd057628f40d2113", "grade": false, "grade_id": "read-head", "locked": false, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
with open("contact.csv") as f:
# YOUR CODE HERE
raise NotImplementedError()
# + [markdown] slideshow={"slide_type": "subslide"}
# **How to write to a text file?**
# + [markdown] slideshow={"slide_type": "fragment"}
# Consider backing up `contact.csv` to a new file:
# + slideshow={"slide_type": "-"}
destination = "private/new_contact.csv"
# + [markdown] slideshow={"slide_type": "fragment"}
# The directory has to be created first if it does not exist:
# + slideshow={"slide_type": "-"}
import os
os.makedirs(os.path.dirname(destination), exist_ok=True)
# + slideshow={"slide_type": "-"}
# os.makedirs?
# !ls
# + [markdown] slideshow={"slide_type": "fragment"}
# To write to the destination file:
# + code_folding=[] slideshow={"slide_type": "-"}
with open("contact.csv") as source_file:
with open(destination, "w") as destination_file:
destination_file.write(source_file.read())
# + slideshow={"slide_type": "-"}
# destination_file.write?
# !more {destination}
# + [markdown] slideshow={"slide_type": "fragment"}
# - The argument `'w'` for `open` sets the file object to write mode.
# - The method `write` writes the input strings to the file.
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercise** We can also use `a` mode to *append* new content to a file.
# Complete the following code to append `new_data` to the file `destination`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "57ff4870e8750ee5331f28704e69da5b", "grade": false, "grade_id": "append", "locked": false, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
new_data = "<NAME>,<EMAIL>, (888) 311-9512"
with open(destination, "a") as f:
# YOUR CODE HERE
raise NotImplementedError()
# !more {destination}
# + [markdown] slideshow={"slide_type": "subslide"}
# **How to delete a file?**
# + [markdown] slideshow={"slide_type": "fragment"}
# Note that the file object does not provide any method to delete the file.
# Instead, we should use the function `remove` of the `os` module.
# + slideshow={"slide_type": "fragment"}
if os.path.exists(destination):
os.remove(destination)
# !ls {destination}
# + [markdown] slideshow={"slide_type": "slide"}
# ## String Objects
# + [markdown] slideshow={"slide_type": "subslide"}
# **How to search for a substring in a string?**
# + [markdown] slideshow={"slide_type": "fragment"}
# A string object has the method `find` to search for a substring.
# E.g., to find the contact information of <NAME>:
# + slideshow={"slide_type": "fragment"}
# str.find?
with open("contact.csv") as f:
for line in f:
if line.find("<NAME>ing") != -1:
record = line
print(record)
break
# + [markdown] slideshow={"slide_type": "subslide"}
# **How to split and join strings?**
# + [markdown] slideshow={"slide_type": "fragment"}
# A string can be split according to a delimiter using the `split` method.
# + slideshow={"slide_type": "-"}
record.split(",")
# + [markdown] slideshow={"slide_type": "fragment"}
# The list of substrings can be joined back together using the `join` methods.
# + slideshow={"slide_type": "-"}
print("\n".join(record.split(",")))
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercise** Print only the phone number (last item) in `record`. Use the method `rstrip` or `strip` to remove unnecessary white spaces at the end.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "58a4fa1d2dc8687eede742a0ac3cc60b", "grade": false, "grade_id": "strip", "locked": false, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
# str.rstrip?
# YOUR CODE HERE
raise NotImplementedError()
# + [markdown] slideshow={"slide_type": "fragment"}
# **Exercise** Print only the name (first item) in `record` but with
# - surname printed first with all letters in upper case
# - followed by a comma, a space, and
# - the first name as it is in `record`.
#
# E.g., `<NAME> Chan` should be printed as `CHAN, T<NAME>`.
#
# *Hint*: Use the methods `upper` and `rsplit` (with the parameter `maxsplit=1`).
# + deletable=false nbgrader={"cell_type": "code", "checksum": "0d20ad9121f6e12c225da56bba6233a0", "grade": false, "grade_id": "process-name", "locked": false, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
# str.rsplit?
# YOUR CODE HERE
raise NotImplementedError()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Operator Overloading
# + [markdown] slideshow={"slide_type": "subslide"}
# ### What is overloading?
# + [markdown] slideshow={"slide_type": "fragment"}
# Recall that the addition operation `+` behaves differently for different types.
# + slideshow={"slide_type": "fragment"}
for x, y in (1, 1), ("1", "1"), (1, "1"):
print(f"{x!r:^5} + {y!r:^5} = {x+y!r}")
# -
# - Having an operator perform differently based on its argument types is called [operator *overloading*](https://en.wikipedia.org/wiki/Operator_overloading).
# - `+` is called a *generic* operator.
# - We can also have function overloading to create generic functions.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### How to dispatch on type?
# + [markdown] slideshow={"slide_type": "fragment"}
# The strategy of checking the type for the appropriate implementation is called *dispatching on type*.
# + [markdown] slideshow={"slide_type": "fragment"}
# A naive idea is to put all different implementations together:
# + slideshow={"slide_type": "-"}
def add_case_by_case(x, y):
if isinstance(x, int) and isinstance(y, int):
print("Do integer summation...")
elif isinstance(x, str) and isinstance(y, str):
print("Do string concatenation...")
else:
print("Return a TypeError...")
return x + y # replaced by internal implementations
for x, y in (1, 1), ("1", "1"), (1, "1"):
print(f"{x!r:^10} + {y!r:^10} = {add_case_by_case(x,y)!r}")
# + [markdown] slideshow={"slide_type": "subslide"}
# It can get quite messy with all possible types and combinations.
# + slideshow={"slide_type": "-"}
for x, y in ((1, 1.1), (1, complex(1, 2)), ((1, 2), (1, 2))):
print(f"{x!r:^10} + {y!r:^10} = {x+y!r}")
# + [markdown] slideshow={"slide_type": "subslide"}
# **What about new data types?**
# + slideshow={"slide_type": "-"}
from fractions import Fraction # non-built-in type for fractions
for x, y in ((Fraction(1, 2), 1), (1, Fraction(1, 2))):
print(f"{x} + {y} = {x+y}")
# + [markdown] slideshow={"slide_type": "fragment"}
# Weaknesses of the naive approach:
# 1. New data types require rewriting the addition operation.
# 1. A programmer may not know all other types and combinations to rewrite the code properly.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### How to have data-directed programming?
# + [markdown] slideshow={"slide_type": "fragment"}
# The idea is to treat an implementation as a datum that can be returned by the operand types.
# + [markdown] slideshow={"slide_type": "fragment"}
# - `x + y` is a [*syntactic sugar*](https://en.wikipedia.org/wiki/Syntactic_sugar) that
# - invokes the method `type(x).__add__(x,y)` of `type(x)` to do the addition.
# + slideshow={"slide_type": "subslide"}
for x, y in (Fraction(1, 2), 1), (1, Fraction(1, 2)):
print(f"{x} + {y} = {type(x).__add__(x,y)}") # instead of x + y
# + [markdown] slideshow={"slide_type": "fragment"}
# - The first case calls `Fraction.__add__`, which provides a way to add `int` to `Fraction`.
# - The second case calls `int.__add__`, which cannot provide any way of adding `Fraction` to `int`. (Why not?)
# + [markdown] slideshow={"slide_type": "fragment"}
# **Why does python return a [`NotImplemented` object](https://docs.python.org/3.6/library/constants.html#NotImplemented) instead of raising an error/exception?**
# + [markdown] slideshow={"slide_type": "fragment"}
# - This allows `+` to continue to handle the addition by
# - dispatching on `Fraction` to call its reverse addition method [`__radd__`](https://docs.python.org/3.6/library/numbers.html#implementing-the-arithmetic-operations).
# + code_folding=[] slideshow={"slide_type": "fragment"}
# %%mytutor -h 500
from fractions import Fraction
def add(x, y):
"""Simulate the + operator."""
sum = x.__add__(y)
if sum is NotImplemented:
sum = y.__radd__(x)
return sum
for x, y in (Fraction(1, 2), 1), (1, Fraction(1, 2)):
print(f"{x} + {y} = {add(x,y)}")
# + [markdown] slideshow={"slide_type": "subslide"}
# The object-oriented programming techniques involved are formally called:
# - [*Polymorphism*](https://en.wikipedia.org/wiki/Polymorphism_(computer_science)): Different types can have different implementations of the `__add__` method.
# - [*Single dispatch*](https://en.wikipedia.org/wiki/Dynamic_dispatch): The implementation is chosen based on one single type at a time.
# + [markdown] slideshow={"slide_type": "fragment"}
# Remarks:
# - A method with starting and trailing double underscores in its name is called a [*dunder method*](https://dbader.org/blog/meaning-of-underscores-in-python).
# - Dunder methods are not intended to be called directly. E.g., we normally use `+` instead of `__add__`.
# - [Other operators](https://docs.python.org/3/library/operator.html?highlight=operator) have their corresponding dunder methods that overloads the operator.
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Object Aliasing
# + [markdown] slideshow={"slide_type": "subslide"}
# **When are two objects identical?**
# + [markdown] slideshow={"slide_type": "fragment"}
# The keyword `is` checks whether two objects are the same object:
# + slideshow={"slide_type": "-"}
sum = 1 .__add__(1.0)
sum is NotImplemented, sum is None
# + [markdown] slideshow={"slide_type": "fragment"}
# **Is `is` the same as `==`?**
# + [markdown] slideshow={"slide_type": "fragment"}
# `is` is faster.
# + slideshow={"slide_type": "-"}
# %%timeit
sum == NotImplemented
# + slideshow={"slide_type": "-"}
# %%timeit
sum is NotImplemented
# + [markdown] slideshow={"slide_type": "fragment"}
# - `is` checks whether two objects occupy the same memory but
# - `==` calls the method `__eq__`.
# + slideshow={"slide_type": "-"}
1 is 1, 1 is 1.0, 1 == 1.0
# + [markdown] slideshow={"slide_type": "fragment"}
# To see this, we can use the function `id` which returns an id number for an object based on its memory location.
# + slideshow={"slide_type": "fragment"}
# %%mytutor -h 400
x, y = complex(1, 2), complex(1, 2)
z = x
for expr in ("id(x)", "id(y)", "id(z)",
"x == y == z", "x is y", "x is z"):
print(expr, eval(expr))
# + [markdown] slideshow={"slide_type": "fragment"}
# As the box-pointer diagram shows:
# - `x` is not `y` because they point to objects at different memory locations,
# even though the objects have the same type and value.
# - `x` is `z` because the assignment `z = x` binds `z` to the same memory location `x` points to.
# `z` is said to be an *alias* (another name) of `x`.
# + [markdown] slideshow={"slide_type": "subslide"}
# **Can we use `is` instead of `==` to compare integers/strings?**
# + slideshow={"slide_type": "-"}
10 ** 10 is 10 ** 10, 10 ** 100 is 10 ** 100
# + slideshow={"slide_type": "-"}
x = y = "abc"
y = "abc"
x is y, y is "abc", x + y is x + "abc"
# + [markdown] slideshow={"slide_type": "fragment"}
# The behavior is not entirely predictable because:
# - it is possible to avoid storing the same integer/string at different locations by [*interning*](https://www.codesansar.com/python-programming/integer-interning.htm), but
# - it is impractical to always avoid it.
#
# + [markdown] slideshow={"slide_type": "fragment"}
# **When should we use `is`?**
# + [markdown] slideshow={"slide_type": "fragment"}
# `is` can be used for [built-in constants](https://docs.python.org/3/library/constants.html#built-in-constants) such as `None` and `NotImplemented`
# because there can only be one instance of each of them.
| Lecture5/Objects.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Linking the functional and stimulus datasets
#
# Now that we have a functional dataset and a mappings from stimulus to labels, we can link the two together and finalize our dataset pipeline.
from packages import *
# %matplotlib inline
# run = False
run = True
# save = False
save = True
load = False
# load = True
# check = False
check = True
subject = 3
def load_events_files(subject):
'''
loads the events files associated with a subject in a dictionary keyed by session and run
'''
data = {}
sub_dir = os.path.join(DATA_DIR, get_subject_dir(subject))
for session in SESSIONS:
data[session] = {}
ses_dir = os.path.join(sub_dir, get_session_dir(session), 'events/')
for f in os.listdir(ses_dir):
if 'localizer' in f:
continue
run = extract_run(f)
f_path = os.path.join(ses_dir, f)
df = load_file(f_path)[0]
# we can get rid of irrelevant columns in data
data[session][run] = df[['ImgName']]
return data
if run:
data = load_events_files(subject)
data[1][1].head(10)
# Next up we will transform our ses:run dictionary into ses:stim, which is the format of our fmri dataset.
if run:
dataset = {}
sessions = data.keys()
sessions.sort()
for ses in sessions:
combined = []
runs = data[ses].keys()
runs.sort()
for r in runs:
for i in xrange(len(data[ses][r])):
combined.append(data[ses][r].iloc[i].ImgName)
dataset[ses] = combined
len(dataset[1])
if save:
save_pickle(dataset, 'func_stim_link', subdir='sub-{}'.format(subject))
| src/b3_link_functional_stim.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 align=center> Homework 5 - Support Vector Machines</h1>
# <br>
# $$
# \textbf{Team G} \\
# \text{<NAME> 2159}\\
# \text{<NAME> 2146}\\
# \text{<NAME> 2209}\\
# $$
# # $\triangleright$ Exercise 1
# ## Question a :
# ### Load and visualize the data
# +
#Libraries
import numpy as np
import matplotlib.pyplot as plt
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report,confusion_matrix
from scipy.sparse import csc_matrix
from math import *
from sklearn import preprocessing
import scipy.io as sio
import pandas as pd
from sklearn import svm
def plot_data(data,labels):
#Split the data based on labels
positive = []
negative = []
for i in range(0,len(labels)):
if(labels[i] == 1):
positive.append(data[i])
else:
negative.append(data[i])
positive =np.array(positive)
negative =np.array(negative)
#Plot the data
plt.scatter(positive[:,0],positive[:,1])
plt.scatter(negative[:,0],negative[:,1])
return
#Split the data from the file
def load_twofeature(filepath):
#Initialite the regular expression
regexp = r"([-+]?\d+) 1:([0-9.]+) 2:([0-9.]+)"
#Parse the regular expression
output = np.fromregex(filepath,regexp,[('label',np.int32),('1',np.float64),('2',np.float64)])
#Parse the ouput of the regex expressio
labels = []
features = []
for i in range(0,len(output['label'])):
labels.append([output['label'][i]]);
features.append([output['1'][i],output['2'][i]])
return np.array(features),np.array(labels)
#Separate the data in labels,features
data, labels = load_twofeature('ex1Data/twofeature.txt')
#Plot data
plot_data(data,labels)
# -
# From a first perspective we can infer that the two classes are linearly separable, although the leftmost blue point seems likely to be misclassified .
# ### Learn the SVM
# +
#Split the data
trainX, testX, trainY, testY = train_test_split(data, labels, test_size=0.30)
def linearSVMForDifferentC(trainX,trainY,testX,testY,C):
classifier = SVC(kernel='linear', C=C).fit(trainX,trainY)
SVs = classifier.support_vectors_ #support vectors
sv_coef = classifier.coef_ #weights
b = classifier.intercept_ #bias
# Visualize the learned model
plot_data(trainX,trainY)
dbDim1 = np.arange(min(trainX[:,0]),max(trainX[:,0]),0.01)
dbDim2 = -(b + sv_coef[0][0]*dbDim1)/sv_coef[0][1]
plt.plot(SVs[:,0],SVs[:,1],'r*')
plt.plot(dbDim1,dbDim2,'k-')
plt.xlabel('Dimension 1')
plt.ylabel('Dimension 2')
# highlight support vectors
plt.title('SVM Linear Classifier with C = %d' %C)
plt.show()
return(b,sv_coef, SVs)
trainY = trainY.reshape((trainY.shape[0],))
#Test SVM for different C
C = 1
b1,sv_coef1,SVs_1 = linearSVMForDifferentC(trainX,trainY,testX,testY,C)
C = 10
b2,sv_coef2,SVs_2 = linearSVMForDifferentC(trainX,trainY,testX,testY,C)
C = 100
b3,sv_coef3,SVs_3 = linearSVMForDifferentC(trainX,trainY,testX,testY,C)
# -
# ##### Do you observe any differences in the learned hyperplane for different values of C? In the evolution of the support vectors?
#
# C is a regularization factor of the SVM algorithm. We can see that for C=1 and progressively speaking for lower Cs, the margin is bigger and the supported vectors are more sparse, although for bigger Cs the margin tends to be smaller and the supported vectors less and closer to the decision boundary. With smaller Cs we make the algorithm look for a large margin even though it might have some misclassifications, but it creates a better general decision boundary. As we raise the C value, we are telling the algorithm that we cannot afford to misclassify examples, but the final result is a smaller margin besides the decision boundary.
# ### Linear kernel
def linear_kernel( xi, xj ):
K = np.inner(xi,xj)
return (K)
# ### Estimate the decision boundary
# +
def plot_db(testX,testY,C,b,sv_coef,SVs):
plot_data(testX,testY)
dbDim1 = np.arange(min(testX[:,0]),max(testX[:,0]),0.01)
dbDim2 = -(b + sv_coef[0][0]*dbDim1)/sv_coef[0][1]
plt.plot(SVs[:,0],SVs[:,1],'r*')
plt.plot(dbDim1,dbDim2,'k-')
plt.xlabel('Dimension 1')
plt.ylabel('Dimension 2')
# highlight support vectors
plt.title('SVM Linear Classifier with C = %d' %C)
plt.show()
C=1
plot_db(testX, testY ,C,b1,sv_coef1,SVs_1)
C=10
plot_db(testX, testY ,C,b2,sv_coef2,SVs_2)
C=100
plot_db(testX, testY ,C,b3,sv_coef3,SVs_3)
# -
# ## Question b :
# ### Load the data
#Read the data and transform it from csc to matrices
def load_emails(numTrainDocs):
numTestDocs = 260
numTokens = 2500
#Prepare the file paths
features_path_train ="ex1Data/emails/train-features-"+str(numTrainDocs)+".txt"
features_path_test ="ex1Data/emails/test-features.txt"
labels_path_train ="ex1Data/emails/train-labels-"+str(numTrainDocs)+".txt"
labels_path_test ="ex1Data/emails/test-labels.txt"
#Get features and labels for training
M = np.loadtxt(features_path_train,delimiter=' ')
#Fix the array to start from zero
for i in range(0,len(M)):
M[i] = [M[i][0]-1,M[i][1]-1,M[i][2]]
rows = np.array(M[:,0])
cols = np.array(M[:,1])
data = np.array(M[:,2])
#Transform the array to compressed column sparse
features_train = csc_matrix((data, (rows, cols)), shape=(numTrainDocs,numTokens)).toarray()
labels_train = np.loadtxt(labels_path_train,delimiter=" ")
#Get features and labels for testing
M = np.loadtxt(features_path_test,delimiter=" ")
#Fix the array to start from zero
for i in range(0,len(M)):
M[i] = [M[i][0]-1,M[i][1]-1,M[i][2]]
rows = np.array(M[:,0])
cols = np.array(M[:,1])
data = np.array(M[:,2])
features_test = csc_matrix((data, (rows, cols)), shape=(numTestDocs,numTokens)).toarray()
labels_test = np.loadtxt(labels_path_test,delimiter=" ")
return features_train,features_test,labels_train,labels_test
# ### Learn and test SVM models - Compute the accuracy
# +
def learn_and_test(numTrainDocs):
#Get the data and split it to test,train
[trainX,testX,trainY,testY] =load_emails(numTrainDocs)
#Fit the model and train it
C = 1
svc = svm.SVC(kernel='linear', C=C).fit(trainX,trainY)
print()
print("For C : ",C," and numTrainDocs: ",numTrainDocs)
print("Weights: ")
print(svc.coef_[0])
plt.plot(svc.coef_[0])
plt.show()
print("Bias:")
print(svc.intercept_)
#Calculate the accuracy
print("Accuracy: {}%".format(svc.score(testX, testY) * 100 ))
#Print out some metrics
yPred = svc.predict(testX)
print(classification_report(testY, yPred))
#learn and test for different files
learn_and_test(50)
learn_and_test(100)
learn_and_test(400)
learn_and_test(700)
# -
# #### As number of training Docs rises we notice that:
# The weight graph is starting to get a certain shape so the weights gradually converge to a certain value. <br>
# Obviously, as with every Machine Learning algorithm we have seen so far, with a large training size we have the ability to train the model better, make it adapt to more occasions of input data and make more accurate predictions. <br>
# Also,we said earlier that a low C (like C=1) is affording some misclassification (especially with low train size) but provides a better general solution. Thus, we can see from the very start that as the problem's size raises (becoming more general) the misclassifications are way less in comparison, because the problem is solved in a better and more general way.
# # $\triangleright$ Exercise 2
# ## Question a:
# ### Load and visualize data.
# Load and plot data.
data,labels = load_twofeature('ex2Data/ex2a.txt')
plot_data(data,labels)
# ### Learn the SVM model
# +
# Set the gamma parameter
gamma = 100
# set the C value
C = 1
#Training the model
X_train, X_test, y_train, y_test = train_test_split(data, labels, test_size = 0.30)
y_train = y_train.reshape((y_train.shape[0],)) # added for warning avoidance
classifier = SVC(kernel="rbf",C=C,gamma=gamma).fit(X_train,y_train)
# Visualize the learned model
plot_data(X_train,y_train)
# -
# ### RBF Kernel
def rbf_kernel( xi, xj, gamma ):
K = exp(-gamma * np.linalg.norm(xi-xj,ord=2)**2 )
return(K)
# ### Visualize the decision cost
# +
def rbf_scoring_function(x,SVs,yi_ai,b,gamma):
score = 0
for i in range(len(yi_ai[0])):
score += yi_ai[0][i]*rbf_kernel(x,SVs[i],gamma)
return(score + b)
# Plot the image and the contours of scoring function
def plot_data_contour(X_train,y_train,classifier,gamma):
step = 0.01
[X,Y] = np.mgrid[0:1:step,0.4:1:step]
X=X.T
Y=Y.T
Z = np.zeros(X.shape)
SVs = classifier.support_vectors_ #support vectors
yi_ai = classifier.dual_coef_
b = classifier.intercept_ #bias
for i in range(len(X)):
for j in range(len(X[0])):
Z[i,j] = rbf_scoring_function([X[i,j],Y[i,j]],SVs,yi_ai,b,gamma)
positive = []
negative = []
for i in range(0,len(y_train)):
if(y_train[i] == 1):
positive.append(X_train[i])
else:
negative.append(X_train[i])
positive =np.array(positive)
negative =np.array(negative)
plt.plot(SVs[:,0],SVs[:,1],"rX",markersize = 4)
plt.scatter(positive[np.where(positive[:,1]>=0.4),0],positive[np.where(positive[:,1]>=0.4),1])
plt.scatter(negative[np.where(negative[:,1]>=0.4),0],negative[np.where(negative[:,1]>=0.4),1])
c=plt.contour(X,Y,Z,levels=[-0.5,0,0.5],color='k')
plt.show()
return(len(SVs))
len_SVS = plot_data_contour(X_train,y_train,classifier,gamma)
# -
# ## Question b:
# ### Load and visualize data
# +
#Split the data from the file(Different from first --> Negative zeros inside)
def load_twofeatures_with_negatives(filepath):
#Initialite the regular expression
regexp = r"([-+]?\d+) 1:([-+]?[0-9.]+) 2:([-+]?[0-9.]+)"
#Parse the regular expression
output = np.fromregex(filepath,regexp,[('label',np.int32),('1',np.float64),('2',np.float64)])
#Parse the ouput of the regex expression
labels = []
features = []
for i in range(0,len(output['label'])):
labels.append([output['label'][i]]);
features.append([output['1'][i],output['2'][i]])
return np.array(features),np.array(labels)
data, labels = load_twofeatures_with_negatives('ex2Data/ex2b.txt')
plot_data(data,labels)
# -
# From an eye perspective the classes are not strictly separable. <br>But a high accuracy general decision boundary can be found, so that only a few misclassifications exist.
# ### Learn SVM models for different hyperparameter values
# +
# Set the parameters
C = np.array([1,1000])
gamma = np.array([1,10,100,1000])
X_train, X_test, y_train, y_test = train_test_split(data, labels, test_size = 0.30)
for i in range(len(C)):
for j in range(len(gamma)):
classifier = SVC(kernel="rbf",C=C[i],gamma=gamma[j])
y_train = y_train.reshape((y_train.shape[0],))
classifier.fit(X_train, y_train)
SVs = classifier.support_vectors_ #support vectors
print("For C : ",C[i]," Gamma: ",gamma[j])
print("Number of Support Vectors: %d" %len(SVs))
print("Accuracy: {}%".format(classifier.score(X_test, y_test) * 100 ))
print('\n')
# -
# ## Question c :
# ### Load the data - Cross validation
# +
#Separate the data in labels,features
[features,labels]=load_twofeatures_with_negatives('ex2Data/ex2b.txt')
#Shuffle the data
R = np.random.randint(0,211,211)
features=features[R,:]
labels = labels[R]
#Build 3 sets for cross-validation
n=70
V1 = features[0:n,:]
labelsV1 = labels[0:n,:]
V2 = features[n:2*n,:]
labelsV2 = labels[n:2*n,:]
V3 = features[2*n:len(features),:]
labelsV3 = labels[2*n:len(features),:];
#Array with 3 sets
V = np.array([V1,V2,V3])
L = np.array([labelsV1,labelsV2,labelsV3])
# -
# ### Perform cross-validation
# +
#Create a meshgrid of (C,gamma) parameters
[X,Y] = np.meshgrid(np.logspace(-1,3,5),np.logspace(-1,3,5))
#Accuracy array
A = np.zeros(shape=X.shape)
n=3
#Check the accuracy for every C,gamma that we generated
for i in range(0,len(X)):
for j in range(0,len(X[0])):
#Get the parameters
C = X[i][j]
gamma = Y[i][j]
acc = np.zeros(n)
#For the 3 folds run the SVM with 2 of 3 as training set and 1 as test
for k in range(0,n):
#Prepare the data
testFeatures = V[k]
testLabels = L[k].flatten()
trainFeatures = np.concatenate((V[(k-1)% n],V[(k+1)% n]),axis=0)
trainLabels = np.concatenate((L[(k-1)% n],L[(k+1)% n]),axis=0)
#Train the model
svc = svm.SVC(kernel='rbf', C=C,gamma=gamma).fit(trainFeatures,trainLabels.flatten())
#Test the model
predictedLabels = svc.predict(testFeatures)
#Calculate the accuracy
acc[k] = svc.score(testFeatures, testLabels)
#Calculate the mean accuracy for all the folds
A[i][j] = np.mean(acc)
#Best combination of parameters
[max_i,max_j] = np.where(A == np.amax(A))
#Retrieve the parameters
C_max = X[max_i[0]][max_j[0]]
gamma_max = Y[max_i[0]][max_j[0]]
A_max = A[max_i[0]][max_j[0]]
print("Best parameters are C: ",C_max," gamma: ",gamma, " Score: ",A_max)
# -
# The results for gamma and C values we get from cross-validating were expected to an extent. <br>
# The C is mid-low, because we can allow some misclassification as the classes not separable, and a high C value would lead to a very complex line with very small margins.A lower C of say 1 would also lead to a lot more misclassifications than this because of the big margin (we can see that the two classes don't have much distance between them). <br>
# Also, to solve the problem we used an RBF kernel to project the points into a higher dimension. Gamma value controls the peak of the RBF "bell". Since we needed a high bias- low variance solution, we got to use a high gamma value, which is 1000. That way, and in a simple manner of speaking, the points that are considered similar are the points that are closer with each other, and closer to the support vectors.
| SVM - HW5/SVM_hw5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## TODO
# * now using app ImplicitSession, because aiovk can't auth from user side. Probably need to auth throw vk-api and then use aiovk.TokenSession
# ### Experiment to asynchonously get users friends using aiovk lib
# +
from typing import List, Optional
import aiovk
import asyncio
import aiohttp
import vk_api
import requests
import json
# -
login = input()
password = input()
proxy_address = input()
proxy_port = input()
# + jupyter={"outputs_hidden": true} tags=[]
s = requests.Session()
user_agent = 'Mozilla/5.0 (Windows NT 6.1; rv:52.0) Gecko/20100101 Firefox/52.0'
s.headers.update({'User-agent': user_agent})
for proxy_protocol in ["http", "https"]:
s.proxies.update({proxy_protocol: f"{proxy_address}:{proxy_port}"})
vk_api_session = vk_api.VkApi(login, password, session=s)
vk_api_session.auth()
# -
access_token = vk_api_session.token["access_token"]
proxy = aiovk.drivers.ProxyDriver(proxy_address, proxy_port)
session = aiovk.TokenSession(access_token, driver=proxy)
async_vk_api = aiovk.API(session)
await async_vk_api.users.get(user_ids=1)
async def get_friends(user_id, async_api):
return await async_api.friends.get(user_id=user_id)
await get_friends(491221043, async_vk_api)
# +
with open("../resources/checkpoints/data_checkpoint.json") as f:
user_data = json.load(f)
groups_to_get = user_data["68076353"]["groups"]
del user_data
async def get_group_wall(group_id, async_api):
return await async_api.wall.get(owner_id=-1 * group_id, extended=False) # negative value means group, positive - user
# -
# ### Asynchronous way
class TokenSessionWithProxy:
"""wrapper for AsyncVkExecuteRequestPool from aiovk because it doesn't support passing
Driver to session"""
def __init__(self, proxy_address, proxy_port):
self.proxy_address, self.proxy_port = proxy_address, proxy_port
def __call__(self, token):
proxy_driver = aiovk.drivers.ProxyDriver(self.proxy_address, self.proxy_port)
return aiovk.TokenSession(token, driver=proxy_driver)
# +
from aiovk.pools import AsyncVkExecuteRequestPool
async def get_groups_walls(groups_ids: List[int], access_token, pool_size=25, req_params: Optional[dict]=None):
# !!!: TODO: should not await for one pool execute, need to make all executes() simultaneously and wait for results
if req_params is None:
req_params = {}
groups_ids = groups_ids.copy()
responses = []
pool_executes = []
while groups_ids:
pool = AsyncVkExecuteRequestPool(token_session_class=TokenSessionWithProxy(proxy_address, proxy_port))
for _ in range(pool_size):
if not groups_ids:
break
group_id = groups_ids.pop(0)
resp = pool.add_call("wall.get", access_token, {"owner_id": -1 * group_id, **req_params})
responses.append(resp)
pool_executes.append(pool.execute())
await asyncio.gather(*pool_executes)
return responses
# +
from time import time
start_time = time()
groups_walls = await get_groups_walls(groups_to_get, access_token=session.access_token, req_params={"extended": False})
print("time needed", time() - start_time)
# -
groups_walls[0].result
# ### synchronous way
pool = vk_api.requests_pool.VkRequestsPool(vk_api_session)
# +
# %%time
responses = []
for idx, group in enumerate(groups_to_get):
resp_raw = pool.method("wall.get", values={"owner_id": -1 * group, "extended": False})
responses.append(resp_raw)
if idx % 25 == 24:
pool.execute()
# -
| notebooks/aiovk.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:miniconda3-metabolic]
# language: python
# name: conda-env-miniconda3-metabolic-py
# ---
# # The Metabolic Index
#
# The Metabolic Index {cite}`Deutsch-Ferrel-etal-2015` is defined as the ratio of oxygen supply to the oxygen demand necessary to sustain respiratory metabolism. Ignoring dependence on body size (mass), the Metabolic Index ($\Phi$) is defined as follows.
#
#
# $$
# \Phi = A_o \frac{P_{\mathrm{O}_2}}
# {\mathrm{exp}\left[
# \frac{-E_o}{k_B}\left(
# \frac{1}{T} - \frac{1}{T_{ref}}
# \right)
# \right]
# }
# $$
#
# Oxygen supply depends on the availability of oxygen in the environment, quantified by the partial pressure of oxygen ($P_{\mathrm{O}_2}$), as well as the physiological acquisition and transport capacities of organisms. The aggegration of these capacities yields a "hypoxic tolerance" trait that varies across marine organisms and is represented by the parameter $A_o$.
#
# Oxygen demand scales as a function of temperature-dependent metabolic rate, represented by the [Arrhenius Equation](https://en.wikipedia.org/wiki/Arrhenius_equation). The temperature sensitivity of metabolic rate is specified by the parameter $E_o$, which is a trait that varies across species.
#
#
# Where the $\Phi >= 1$, an organism can meet its resting metabolic demand. Notably, however, this precludes activity necessary for reproduction and feeding; thus organisms require $\Phi >= \Phi_{crit}$, where $\Phi_{crit}$ is the minimum value of $\Phi$ sufficient to sustain ecologically-relevant metabolic rates.
#
# Here we provide a illustration of how $\Phi$ enables quanitification of habitability in the context of constraints imposed by the requirements of aerobic metabolism.
# + tags=["hide-input"]
# %load_ext autoreload
# %autoreload 2
import cmocean
import constants
import matplotlib.pyplot as plt
import metabolic as mi
import numpy as np
import util
import xarray as xr
from scipy import stats as scistats
# -
# ## Load traits database
#
# Load a subset of the trait data from {cite:t}`Deutsch-Penn-etal-2020`, including only the marine organisms for which temerature-dependent hypoxia metabolic traits have been determined.
#
# The `open_traits_df` function is defined in the [metabolic](https://github.com/matt-long/aerobic-safety-margins/blob/main/notebooks/metabolic.py) module and makes the trait data available via a [pandas](https://pandas.pydata.org/) `DataFrame`.
df = mi.open_traits_df()
df
# Pull out some median traits for illustration purposes. [This notebook](./trait-space-joint-pdf.ipynb) presents a full exploration of trait PDFs.
Ac_med = mi.trait_pdf(df, 'Ac', 30).median()
print(f'Median Ac = {Ac_med:0.3f} 1/kPa')
Ao_med = mi.trait_pdf(df, 'Ao', 30).median()
print(f'Median Ao = {Ao_med:0.3f} 1/kPa')
Eo_med = mi.trait_pdf(df, 'Eo', 30).median()
print(f'Median Eo = {Eo_med:0.3f} eV')
# ## Explaining the Metabolic Index
#
# In the cell below, we define a function that plots curves of constant $\Phi$ in in \PO2{}-temperature space (click "+" at right to see the code).
# + tags=["hide-input"]
def plot_MI_illustration(which):
fig, ax = plt.subplots()
PO2_atm = constants.XiO2 * constants.kPa_per_atm
T = np.arange(0, 32, 0.1)
pO2_at_Phi_crit = mi.pO2_at_Phi_one(T, Ac_med, Eo_med * 2)
pO2_at_Phi_one = mi.pO2_at_Phi_one(T, Ao_med, Eo_med * 2)
ATmax_crit = mi.compute_ATmax(PO2_atm, Ac_med, Eo_med * 2)
ATmax_one = mi.compute_ATmax(PO2_atm, Ao_med, Eo_med * 2)
color_rest = 'tab:blue'
color_active = 'tab:red'
# active
if 'active' in which:
ax.plot(T, pO2_at_Phi_crit, '-', linewidth=2, color=color_active)
ax.fill_between(
T,
pO2_at_Phi_crit,
constants.XiO2 * constants.kPa_per_atm,
where=pO2_at_Phi_crit <= PO2_atm,
color='tab:green',
alpha=0.5,
)
ax.plot(
ATmax_crit,
PO2_atm,
'o',
color=color_active,
)
ax.text(
10,
mi.pO2_at_Phi_one(10, Ac_med, Eo_med * 2) - 1.5,
r'$\Phi = \Phi_{crit}$',
color=color_active,
fontsize=16,
rotation=35,
)
# resting
ax.plot(T, pO2_at_Phi_one, '-', linewidth=2, color=color_rest)
ax.plot(
ATmax_one,
PO2_atm,
'o',
color=color_rest,
)
ax.text(
25,
mi.pO2_at_Phi_one(25, Ao_med, Eo_med * 2) - 1.5,
r'$\Phi = 1$',
color=color_rest,
fontsize=16,
rotation=48,
)
if 'resting' in which:
ax.fill_between(
T,
pO2_at_Phi_one,
constants.XiO2 * constants.kPa_per_atm,
where=pO2_at_Phi_one <= PO2_atm,
color='tab:green',
alpha=0.5,
)
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.spines['left'].set_linewidth(2)
ax.spines['bottom'].set_linewidth(2)
ax.set_xticks([])
ax.set_yticks([])
ax.set_xlim([T.min(), T.max()])
ax.set_xlabel('Temperature [°C]')
ax.set_ylabel(r'$P_{\mathrm{O}_2}$ [kPa]')
xlm = ax.get_xlim()
ylm = (-2.6251270379913803, 73.89798889088694) # ax.get_ylim()
ax.set_ylim([ylm[0], PO2_atm + 0.05 * np.diff(ylm)])
ax.axhline(PO2_atm, linewidth=1, color='k', linestyle='--', zorder=-100)
ax.text(
xlm[0] + 0.05 * np.diff(xlm), PO2_atm + 0.01 * np.diff(ylm), r'$P_{\mathrm{O}_2}^{atm}$'
)
if 'base' in which:
ax.plot(
[15.0, 15.0],
[ylm[0], mi.pO2_at_Phi_one(15, Ao_med, Eo_med * 2)],
'--',
color=color_rest,
)
ax.plot(
[xlm[0], 15.0],
[mi.pO2_at_Phi_one(15, Ao_med, Eo_med * 2), mi.pO2_at_Phi_one(15, Ao_med, Eo_med * 2)],
'--',
color=color_rest,
)
ax.text(15 - 0.25, ylm[0] + 0.01 * np.diff(ylm), 'T$_{ref}$', ha='right', color=color_rest)
ax.text(
15 / 2,
mi.pO2_at_Phi_one(15, Ao_med, Eo_med * 2) + 0.01 * np.diff(ylm),
'$1/A_o$ = Hypoxic tolerance',
ha='center',
color=color_rest,
)
spc = ' ' * 23
ax.text(
18,
mi.pO2_at_Phi_one(18, Ao_med, Eo_med * 2) - 0.06 * np.diff(ylm),
f'slope $\\propto$ $E_o$ = Temperature\n{spc}sensitivity of\n{spc}metabolism',
ha='left',
color=color_rest,
)
if 'resting' in which:
ax.text(7, PO2_atm / 1.5, 'Habitable', color='tab:green', fontsize=16, fontweight='bold')
ax.text(18, PO2_atm / 12, 'Not\nhabitable', color='tab:red', fontsize=16, fontweight='bold')
if 'resting' in which or 'active' in which:
ax.plot([ATmax_one, ATmax_one], [ylm[0], PO2_atm], '--', color=color_rest)
ax.text(
ATmax_one - 0.25,
ylm[0] + 0.01 * np.diff(ylm),
'Resting\nAT$_{max}$',
ha='right',
color=color_rest,
)
if 'active' in which:
ax.plot([ATmax_crit, ATmax_crit], [ylm[0], PO2_atm], '--', color=color_active)
ax.text(
ATmax_crit - 0.25,
ylm[0] + 0.01 * np.diff(ylm),
'Active\nAT$_{max}$',
ha='right',
color=color_active,
)
plt.savefig(f'figures/misc/phi-explain-{which}.png', dpi=300)
# -
# Using this function, we can produce a figure similar to Fig 1 of {cite:t}`Deutsch-Penn-etal-2020`, showing curves of constant $\Phi$ in $P_{\mathrm{O}_2}$-temperature space.
#
# This plot illustrates how the paramters control the shape of the curve.
# + tags=[]
plot_MI_illustration('base')
# -
# The $\Phi = 1$ line delineates the region in $P_{\mathrm{O}_2}$-temperature space that is habitable from that that is too warm with insufficient oxygen. The intersection of this line with $P_{\mathrm{O}_2}^{atm}$ defines the maximum temperature at which metabolism can be sustained.
# + tags=[]
plot_MI_illustration('resting')
# -
# Ecological requirements to sustain metabolic rates above those of resting metabolism impose further restrictions on habiability.
#
# The $\Phi = \Phi_{crit}$ line inscribes a smaller region of habitability in $P_{\mathrm{O}_2}$-temperature space than that corresponding to resting metabolism ($\Phi = 1$).
# + tags=[]
plot_MI_illustration('active')
| notebooks/metabolic-index-defining.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import seaborn as sns
from matplotlib import pyplot as plt
# %matplotlib inline
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
from sklearn.decomposition import PCA
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from imblearn.over_sampling import SMOTE
import xgboost as xgb
from sklearn import metrics
from sklearn import preprocessing
from sklearn import svm
df_original = pd.read_csv('online_shoppers_intention.csv')
# ### Opis skupa:
df_original.sample(10)
df_original.describe()
# ### Atributi:
for c in df_original.columns:
print(c)
# ### Vizualizacija raspodele podataka
pd.DataFrame.hist(df_original, column = df_original.columns[:12], figsize = [15,15])
print(f'Broj instanci: {df_original.shape[0]}\nBroj atributa: {df_original.shape[1]}')
# ### Ciljna promenljiva biće Revenue (prihod)
all_revenue = df_original['Revenue']
revenue = df_original['Revenue'].unique()
n_classes = revenue.shape[0]
# ### Klase:
for c in revenue:
print(c)
print('False: {}\nTrue: {}'.format(np.sum(all_revenue == False), np.sum(all_revenue == True)))
changes = dict( zip(revenue, range(n_classes)))
binary_revenue = all_revenue.replace(changes)
# +
plt.title('Podaci')
plt.xticks([0,1])
plt.hist(binary_revenue)
#plt.savefig('klase.png')
# -
print('Nedostajuce vrednosti u podacima:\n')
print(df_original.isnull().sum())
# ### Koje atribute zadržavamo?
# +
plt.figure(figsize = (20,15))
correlation_matrix = df_original.corr()
sns.heatmap(correlation_matrix, annot = True, cmap = plt.cm.Blues)
#plt.savefig('korelacija.png')
# -
# ### Možemo izbaciti kolone koje su visoko korelirane ( >= 0.85 )
df_original = df_original.drop(['ExitRates', 'ProductRelated_Duration'], axis = 1)
# ### Na osnovu onoga što nam je vratio XGBoost izbacujemo nerelevantne atribute ( eksperimentalno )
# +
# Pod komentarom je jer je korišćeno samo eksperimentalno
#df_original = df_original.drop(['Informational_Duration', 'Weekend', 'Browser', 'SpecialDay'], axis = 1)
# -
# ### Enkodiranje kategoričkih atributa
# +
df = pd.get_dummies(df_original, columns = ['OperatingSystems', 'Browser', 'Region', 'TrafficType', 'VisitorType', 'Month', 'Weekend'])
# Ako izbacimo atribute na osnovu XGBoost algoritma, enkodiramo sledeće atribute:
#df = pd.get_dummies(df_original, columns = ['OperatingSystems', 'Region', 'TrafficType', 'VisitorType', 'Month'])
# -
df = df.replace(changes)
print(df.columns)
# Novi izgled skupa:
df.head()
# ### Podela na skup atributa i specijalni atribut klase
X = df.drop('Revenue', axis = 1)
Y = df['Revenue']
print(f'Atributi koji ce biti korisceni:')
for c in X.columns:
print(c)
# ### Podela na trening i test skup i standardizacija podataka
# +
x_train, x_test, y_train, y_test = train_test_split(X, Y, test_size = 0.3, stratify = Y, random_state = 7)
x_train_before = pd.DataFrame(x_train, columns = X.columns)
scaler = preprocessing.StandardScaler()
scaler.fit(x_train)
x_train_scaled = scaler.transform(x_train)
x_test_scaled = scaler.transform(x_test)
x_train_scaled = pd.DataFrame(x_train_scaled, columns = X.columns)
x_test_scaled = pd.DataFrame(x_test_scaled, columns = X.columns)
# +
fig, (ax1, ax2) = plt.subplots(ncols = 2, figsize = (15, 5))
ax1.set_title('Pre standardizacije')
sns.kdeplot(x_train_before['Administrative'], ax = ax1)
sns.kdeplot(x_train_before['Informational'], ax = ax1)
sns.kdeplot(x_train_before['BounceRates'], ax = ax1)
ax2.set_title('Posle standardizacije')
sns.kdeplot(x_train_scaled['Administrative'], ax = ax2)
sns.kdeplot(x_train_scaled['Informational'], ax = ax2)
sns.kdeplot(x_train_scaled['BounceRates'], ax = ax2)
#plt.savefig('standardizacija.png')
# -
# ### Balansiranje klasa
# +
print('Broj instanci u klasi "1" pre balansiranja: {}'.format(sum(y_train == 1)))
print('Broj instanci u klasi "0" pre balansiranja: {}'.format(sum(y_train == 0)))
sm = SMOTE(sampling_strategy = 'minority', random_state = 2)
x_train_balanced, y_train = sm.fit_sample(x_train_scaled, y_train.ravel())
print()
print('Broj instanci u klasi "1" nakon balansiranja: {}'.format(sum(y_train == 1)))
print('Broj instanci u klasi "0" pre balansiranja: {}'.format(sum(y_train == 0)))
x_train_balanced = pd.DataFrame(x_train_balanced, columns = X.columns)
# -
# ### Skup podataka nakon preprocesiranja ( bez PCA jer on neće biti korišćen svuda )
# +
X_new = pd.concat([x_train_balanced, x_test_scaled])
df1 = pd.DataFrame(y_train)
df1.columns = ['Revenue']
df2 = pd.DataFrame(y_test)
Y_new = pd.concat([df1, df2])
X_new = X_new.loc[~X_new.index.duplicated(keep = 'first')]
Y_new = Y_new.loc[~Y_new.index.duplicated(keep = 'first')]
final = pd.concat([X_new, Y_new], axis = 1)
#final.to_csv(r'preprocesiran_skup.csv')
# -
# ### Algoritam XGBoost - korišćen za odabir atributa
xg_clf = xgb.XGBClassifier()
xg_clf.fit(x_train_balanced, y_train)
# +
xgb.plot_importance(xg_clf, ylabel = 'Atributi', title = 'Značaj atributa', xlabel = 'F mera')
# plt.savefig('atr.png')
# -
# Na osnovu onoga što nam je vratio XGBoost možemo odrediti bitne atribute
# ### Primena algoritma PCA za smanjenje dimenzionalnosti
# +
pca = PCA(0.95)
pca.fit(x_train_balanced)
x_train_pca = pca.transform(x_train_balanced)
x_test_pca = pca.transform(x_test_scaled)
# -
# ## Učenje modela
# ### Logistička regresija
# +
parameters = {
'C': np.linspace(1, 10, 10),
}
lr = LogisticRegression()
clf = GridSearchCV(lr, parameters, verbose = 8, cv = 5, n_jobs = 3)
clf.fit(x_train_balanced, y_train)
clf.best_params_
# -
lr = LogisticRegression(C = 1.0, penalty = 'l2', solver = 'lbfgs', max_iter = 200)
lr.fit(x_train_balanced, y_train)
# #### Evaluacija
def evaluate(model, x_tr, x_tst):
y_test_predicted = model.predict(x_tst)
test_score = metrics.accuracy_score(y_test, y_test_predicted)
y_train_predicted = model.predict(x_tr)
train_score = metrics.accuracy_score(y_train, y_train_predicted)
print("Train score: {train}\nTest score: {test}".format(train = train_score, test = test_score))
print()
print(metrics.classification_report(y_test, y_test_predicted))
return y_test_predicted
def draw_matrix(y_tst_predicted, cmap):
confusion_matrix = metrics.confusion_matrix(y_test, y_tst_predicted)
fig, ax = plt.subplots()
img = ax.imshow(confusion_matrix, interpolation = 'nearest', cmap = cmap)
ax.figure.colorbar(img, ax = ax)
ax.set(xticks = np.arange(confusion_matrix.shape[1]),
yticks = np.arange(confusion_matrix.shape[0]),
xticklabels = ['False', 'True'], yticklabels = ['False', 'True'],
title = 'Matrica konfuzije',
ylabel = 'Prava klasa',
xlabel = 'Predviđena klasa')
fmt = '.2f'
thresh = confusion_matrix.max() / 2.
for i in range(confusion_matrix.shape[0]):
for j in range(confusion_matrix.shape[1]):
ax.text(j, i, format(confusion_matrix[i, j], fmt),
ha = "center", va = "center",
color = "white" if confusion_matrix[i, j] > thresh else "black")
fig.tight_layout()
#plt.savefig('konf3.png')
y_test_predicted = evaluate(lr, x_train_balanced, x_test_scaled)
draw_matrix(y_test_predicted, plt.cm.Reds)
# ### Stabla odlučivanja
# +
# params = {
# 'max_depth' : [3, 5, 7, 10, 15, 20],
# 'criterion' : ['gini', 'entropy']
# }
# +
# dt = DecisionTreeClassifier()
# grid = GridSearchCV(dt, params, cv = 5, verbose = 8)
# grid.fit(x_train_balanced, y_train)
# grid.best_params_
# -
dt = DecisionTreeClassifier(criterion = 'entropy', max_depth = 7)
dt.fit(x_train_balanced, y_train)
y_test_predicted = evaluate(dt, x_train_balanced, x_test_scaled)
draw_matrix(y_test_predicted, plt.cm.Greens)
# ### SVM
# #### SVM bez kernela
# +
# params = {
# 'C' : range(1, 3, 2),
# }
# +
# lin = svm.LinearSVC()
# grid = GridSearchCV(lin, params, cv = 5, verbose = 8)
# grid.fit(x_train_balanced, y_train)
# grid.best_params_
# -
lin_svm = svm.LinearSVC(loss = 'hinge', C = 1.0)
lin_svm.fit(x_train_balanced, y_train)
y_test_predicted = evaluate(lin_svm, x_train_balanced, x_test_scaled)
draw_matrix(y_test_predicted, plt.cm.Blues)
# #### SVM sa kernelom
# +
# params = {
# 'kernel' : ['rbf', 'poly', 'sigmoid'],
# 'C' : [1.0, 2.0]
# }
# +
# lin_ker = svm.SVC(kernel = 'rbf')
# grid = GridSearchCV(lin_ker, params, cv = 5, verbose = 8)
# grid.fit(x_train_balanced, y_train)
# grid.best_params_
# -
kernelized_svm = svm.SVC(kernel = 'rbf', C = 2.0)
kernelized_svm.fit(x_train_balanced, y_train)
y_test_predicted = evaluate(kernelized_svm, x_train_balanced, x_test_scaled)
draw_matrix(y_test_predicted, plt.cm.Purples)
# ### Random decision forest
# +
# params = {
# 'n_estimators' : range(5, 20, 5),
# 'criterion' : ['gini', 'entropy']
# }
# +
# rfc = RandomForestClassifier()
# grid = GridSearchCV(rfc, params, cv = 5, verbose = 8)
# grid.fit(x_train_balanced, y_train)
# grid.best_params_
# -
rfc = RandomForestClassifier(n_estimators = 15, criterion = 'entropy')
rfc.fit(x_train_balanced, y_train)
y_test_predicted = evaluate(rfc, x_train_balanced, x_test_scaled)
draw_matrix(y_test_predicted, plt.cm.Oranges)
# ### Konačni model
# +
X = pd.concat([x_train_balanced, x_test_scaled])
y_train = pd.DataFrame(y_train, columns = ['Revenue'])
y_test = pd.DataFrame(y_test, columns = ['Revenue'])
Y = pd.concat([y_train, y_test])
# -
rfc.fit(X, Y.values.ravel())
| kodovi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1.2.1.Add_Background
import os
import csv
import numpy as np
from PIL import Image
# making tripo maps
import cv2
# +
label_ids = ['10_espalda', '10_frente',
'20_espalda', '20_frente',
'50_espalda', '50_frente',
'100_espalda', '100_frente',
'200_espalda', '200_frente'
]
width = 224
height = 224
#sizes = [0.116, 0.07, 0.05]
sizes = [0.116]
count_per_size = 1
# +
count_background = 0
for label_id in label_ids:
label = label_id
prefix_filename = label + "_"
path_images_labels = 'data_augmentation/banknotes/' + label + '/'
path_images_backgrounds = 'data_augmentation/background_' + str(width) + '/'
path_images_labeled = 'data_' + str(width) + '/train/' + label + '/'
bkg_images = [f for f in os.listdir(path_images_backgrounds) if not f.startswith(".")]
obj_images = [f for f in os.listdir(path_images_labels) if not f.startswith(".")]
coordinates = [] # store coordinate here
print("Labeling images for label: ", label)
print("...")
# Helper functions
def get_obj_positions(obj, bkg, count=1):
obj_w, obj_h = [], []
x_positions, y_positions = [], []
bkg_w, bkg_h = bkg.size
# Rescale our obj to have a couple different sizes
obj_sizes = [tuple([int(s*x) for x in obj.size]) for s in sizes]
for w, h in obj_sizes:
obj_w.extend([w]*count)
obj_h.extend([h]*count)
max_x, max_y = bkg_w-w, bkg_h-h
if max_x < w:
x_positions.extend([0])
else:
x_positions.extend(list(np.random.randint(0, max_x, count)))
if max_y < h:
y_positions.extend([0])
else:
y_positions.extend(list(np.random.randint(0, max_y, count)))
return obj_h, obj_w, x_positions, y_positions
# Creando los bounding boxes
n = 0
index_selected = 0
#for bkg in bkg_images:
for img_path in obj_images:
# validate is count_background is mayor than len background
if count_background >= len(bkg_images):
count_background = 0
# Load the background image
bkg_path = path_images_backgrounds + bkg_images[count_background]
bkg_img = Image.open(bkg_path)
bkg_x, bkg_y = bkg_img.size
count_background += 1
# Do single objs first
#if index_selected >= len(obj_images):
# index_selected = 0
#i = obj_images[index_selected] # seleccionando una imagen
i = img_path # seleccionando una imagen
index_selected += 1 # Aumentando en uno para la siguiente imagen
# Load the single obj
i_path = path_images_labels + i
obj_img = Image.open(i_path)
# Get an array of random obj positions (from top-left corner)
obj_h, obj_w, x_center_pos, y_center_pos = get_obj_positions(obj=obj_img, bkg=bkg_img, count=count_per_size)
# Create synthetic images based on positions
for h, w, x_center, y_center in zip(obj_h, obj_w, x_center_pos, y_center_pos):
# Copy background
bkg_w_obj = bkg_img.copy()
# Adjust obj size
new_obj = obj_img.resize(size=(w, h))
# Paste on the obj
bkg_w_obj.paste(new_obj, (x_center, y_center), new_obj)
filename = prefix_filename + str(n)
output_fp = path_images_labeled + label + filename + '.png'
# Save the image
bkg_w_obj.save(fp=output_fp, format="png")
n += 1
print("finished")
# -
| 1.2.1.Add_Background.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import pymongo
import json
import warnings
warnings.filterwarnings('ignore')
# Function was developed to load two csvfiles into MongoDB
def import_content(filepath, collection_name):
client = pymongo.MongoClient('localhost', 27017)
db = client['gene_db'] # Database name was given here
collection_name = collection_name
db_cm = db[collection_name]
data = pd.read_csv(filepath)
data_json = json.loads(data.to_json(orient='records'))
db_cm.remove()
db_cm.insert(data_json)
# +
# csvfile was loaded to database in collection "gene_variants"
filepath = 'gene_variants.csv'
collection_name = 'gene_variants'
import_content(filepath,collection_name)
# -
# csvfile was loaded to database in collection "gene_variants"
filepath = 'geneData.csv'
collection_name = 'geneData'
import_content(filepath,collection_name)
# csvfile was loaded to database in collection "gene_variants"
filepath = 'somatic.csv'
collection_name = 'somatic'
import_content(filepath,collection_name)
# +
client = pymongo.MongoClient('localhost', 27017)
db = client.gene_db
# -
db.collection_names()
db.gene_variants.find_one()
db.geneData.find_one()
db.gene_variants.count()
| ETL/dataload/mongodb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import pandas as pd
import numpy as np
from laplace_equation.utils.result_generation import build_input_matrix
# -
data = pd.read_csv("laplace_equation/data/result.csv")
data
# +
input_data = []
for i, record in data.iterrows():
input_temperature = build_input_matrix(record["nominal_temperature"], record["boundary_conditions"])
input_data.append(input_temperature)
input_data = np.asarray(input_data)
output_data = np.load("laplace_equation/data/output_data.npy")
# -
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(input_data, output_data.reshape(-1, 100), test_size=0.2, random_state=42)
x_train = x_train[..., tf.newaxis].astype("float32")
x_test = x_test[..., tf.newaxis].astype("float32")
train_dataset = tf.data.Dataset.from_tensor_slices((x_train, y_train)).shuffle(10000).batch(32)
test_dataset = tf.data.Dataset.from_tensor_slices((x_test, y_test)).batch(32)
# +
from tensorflow.keras.layers import Dense, Flatten, Conv2D
from tensorflow.keras import Model
class MyModel(Model):
def __init__(self):
super(MyModel, self).__init__()
self.conv1 = Conv2D(32, 3, activation='relu')
# self.conv2 = Conv2D(64, 3, activation='relu')
self.flatten = Flatten()
self.fc1 = Dense(128, activation='relu')
self.fc2 = Dense(100)
def call(self, x):
x = self.conv1(x)
# x = self.conv2(x)
x = self.flatten(x)
x = self.fc1(x)
return self.fc2(x)
# Create an instance of the model
model = MyModel()
loss_fn = tf.keras.losses.MeanSquaredError()
optimizer = tf.keras.optimizers.Adam()
train_loss = tf.keras.metrics.Mean(name='train_loss')
train_accuracy = tf.keras.metrics.Accuracy(name='train_accuracy')
test_loss = tf.keras.metrics.Mean(name='test_loss')
test_accuracy = tf.keras.metrics.Accuracy(name='test_accuracy')
# -
@tf.function
def train_step(input_data, output_data):
with tf.GradientTape() as tape:
predictions = model(input_data, training=True)
loss = loss_fn(output_data, predictions)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
train_loss(loss)
train_accuracy(tf.round(output_data), tf.round(predictions))
@tf.function
def test_step(input_data, output_data):
predictions = model(input_data, training=False)
t_loss = loss_fn(output_data, predictions)
test_loss(t_loss)
test_accuracy(tf.round(output_data), tf.round(predictions))
# +
nb_epoch = 50
for epoch in range(nb_epoch):
# Reset the metrics at the start of the next epoch
train_loss.reset_states()
train_accuracy.reset_states()
test_loss.reset_states()
test_accuracy.reset_states()
for input_temperature, output_temperature in train_dataset:
train_step(input_temperature, output_temperature)
for input_temperature, output_temperature in test_dataset:
test_step(input_temperature, output_temperature)
template = 'Epoch {}, Loss: {}, Accuracy: {}, Test Loss: {}, Test Accuracy: {}'
print(template.format(epoch + 1,
train_loss.result(),
train_accuracy.result() * 100,
test_loss.result(),
test_accuracy.result() * 100))
# -
| Conv2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import pandas as pd
np.random.seed(45)
import matplotlib
import matplotlib.pyplot as plt
plt.rcParams['axes.labelsize'] = 14
plt.rcParams['xtick.labelsize'] = 12
plt.rcParams['ytick.labelsize'] = 12
indicator = pd.read_csv('indicator1.csv')
indicator.head()
# Drop the code column and year column
indicator.drop(indicator.columns[[0, 2]], axis=1, inplace=True)
indicator.head()
indicator.info()
# Use describe() method to show the summary statistics of numeric attributes.
indicator.describe()
# The count, mean, min and max rows are self-explanatory. The std shows standard deviation. The 25%, 50% and 75% rows show the corresponding percentiles.
#
# To get a feel of what type of the data we are dealing with, we plot a histogram for each numeric attribute.
indicator.hist(bins=50, figsize=(20, 15))
plt.savefig('numeric_attributes.png')
plt.show()
# Observations:
#
# These attributes have very different scales, we will need to apply feature scaling.
#
# Many histogram are right skewed. This may make it harder for some machine learning algorithms to detect patterns. We will need to transform them to more normal distributions.
# check for correlation between attributes.
# +
from pandas.plotting import scatter_matrix
attributes = ["GDP_per_capita", "Hours_do_tax", "Days_reg_bus", "Cost_start_Bus",
"Bus_tax_rate", "Ease_Bus"]
scatter_matrix(indicator[attributes], figsize=(12, 8))
plt.savefig("scatter_matrix_plot.png")
plt.show()
# -
# It seems GDP per Capita has a negative correlation with Ease of doing business. The other attributes all have a positive correlation with Ease of doing business. Let's find the most promising attribute to predict the Ease of doing business.
from sklearn.linear_model import LinearRegression
X = indicator.drop(['country', 'Ease_Bus'], axis=1)
regressor = LinearRegression()
regressor.fit(X, indicator.Ease_Bus)
print('Estimated intercept coefficient:', regressor.intercept_)
print('Number of coefficients:', len(regressor.coef_))
pd.DataFrame(list(zip(X.columns, regressor.coef_)), columns = ['features', 'est_coef'])
# The most promising attribute to predict the "ease of doing business" is the "days spent to register a business", so let’s zoom in on their correlation scatterplot.
indicator.plot(kind="scatter", x="Days_reg_bus", y="Ease_Bus",
alpha=0.8)
plt.savefig('scatter_plot.png')
# The correlation is indeed very strong; you can clearly see the upward trend and the points are not too dispersed.
# Split the data into training and test
# +
from sklearn.cross_validation import train_test_split
y = indicator.Ease_Bus
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
# -
# Build a Linear Regression model
from sklearn.linear_model import LinearRegression
regressor = LinearRegression()
regressor.fit(X_train, y_train)
y_pred = regressor.predict(X_test)
regressor.score(X_test, y_test)
# So in our model, 60.7% of the variability in Y can be explained using X.
# +
from sklearn.metrics import mean_squared_error
regressor_mse = mean_squared_error(y_pred, y_test)
import math
math.sqrt(regressor_mse)
# -
# So we are an average of 33.59 away from the ground true score on "ease of doing business" when making predictions on our test set.
#
# The median score of "Ease of doing business" is 95, so a typical prediction error of 33.59 is not very satisfying. This is an example of a model underfitting the training data. When this happens it can mean that the features do not provide enough information to make good predictions, or that the model is not powerful enough. The main ways to fix underfitting are to select more features from Wordbank indicators(e.g., "getting credit", 'registering property" and so on).
regressor.predict([[41096.157300, 5.0, 3, 58.7, 161.0]])
indicator.loc[indicator['country'] == 'Belgium']
regressor.predict([[42157.927990, 0.4, 2, 21.0, 131.0]])
indicator.loc[indicator['country'] == 'Canada']
plt.scatter(regressor.predict(X_train), regressor.predict(X_train)-y_train, c='indianred', s=40)
plt.scatter(regressor.predict(X_test), regressor.predict(X_test)-y_test, c='b', s=40)
plt.hlines(y=0, xmin=0, xmax=200)
plt.title('Residual plot using training(red) and test(blue) data')
plt.ylabel('Residual')
plt.savefig('residual_plot.png')
| notebook/general/Ease_of_Business.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] Collapsed="false"
# # Create climatology and persistence forecasts
#
# In this note book we will create the most basic baselines: persistence and climatology forecasts.
# + Collapsed="false"
# %load_ext autoreload
# %autoreload 2
# + Collapsed="false"
import numpy as np
import xarray as xr
import matplotlib.pyplot as plt
import seaborn as sns
from src.score import *
# + Collapsed="false"
sns.set_style('darkgrid')
sns.set_context('notebook')
# + [markdown] Collapsed="false"
# ## Load data
#
# First, we need to specify the directories and load the data.
# + Collapsed="false"
res = '5.625'
DATADIR = f'/data/weather-benchmark/{res}deg/'
PREDDIR = '/data/weather-benchmark/predictions/'
# + Collapsed="false"
# Load the entire dataset
z500 = xr.open_mfdataset(f'{DATADIR}geopotential_500/*.nc', combine='by_coords').z
t850 = xr.open_mfdataset(f'{DATADIR}temperature_850/*.nc', combine='by_coords').t.drop('level')
data = xr.merge([z500, t850])
# + Collapsed="false"
# Load the validation subset of the data: 2017 and 2018
z500_valid = load_test_data(f'{DATADIR}geopotential_500', 'z')
t850_valid = load_test_data(f'{DATADIR}temperature_850', 't').drop('level')
valid_data = xr.merge([z500_valid, t850_valid])
# + [markdown] Collapsed="false"
# ## Persistence
#
# Persistence simply means: Tomorrow's weather is today's weather.
# + Collapsed="false"
def create_persistence_forecast(ds, lead_time_h):
assert lead_time_h > 0, 'Lead time must be greater than 0'
ds_fc = ds.isel(time=slice(0, -lead_time_h))
return ds_fc
# + Collapsed="false"
lead_times = xr.DataArray(
np.arange(6, 126, 6), dims=['lead_time'], coords={'lead_time': np.arange(6, 126, 6)}, name='lead_time')
# + Collapsed="false"
persistence = []
for l in lead_times:
persistence.append(create_persistence_forecast(valid_data, int(l)))
persistence = xr.concat(persistence, dim=lead_times)
# + Collapsed="false"
persistence
# + [markdown] Collapsed="false"
# The forecast files have dimensions `[init_time, lead_time, lat, lon]`. Let's now save these files so we can evaluate them later.
# + Collapsed="false"
# Save the predictions
persistence.to_netcdf(f'{PREDDIR}persistence_{res}.nc')
# + [markdown] Collapsed="false"
# ## Climatology
#
# First let's create a single climatology from the entire training dataset (meaning everything before 2017).
# + Collapsed="false"
def create_climatology_forecast(ds_train):
return ds_train.mean('time')
# + Collapsed="false"
train_data = data.sel(time=slice(None, '2016'))
# + Collapsed="false"
climatology = create_climatology_forecast(train_data)
# + Collapsed="false"
climatology
# + Collapsed="false"
climatology.z.plot();
# + Collapsed="false"
climatology.to_netcdf(f'{PREDDIR}climatology_{res}.nc')
# + [markdown] Collapsed="false"
# ## Climatology by week
#
# We can create amuch better climatology by taking the seasonal cycle into account. Here we will do this by creating a separate climatology for every week.
# + Collapsed="false"
def create_weekly_climatology_forecast(ds_train, valid_time):
ds_train['week'] = ds_train['time.week']
weekly_averages = ds_train.groupby('week').mean('time')
valid_time['week'] = valid_time['time.week']
fc_list = []
for t in valid_time:
fc_list.append(weekly_averages.sel(week=t.week))
return xr.concat(fc_list, dim=valid_time)
# + Collapsed="false"
weekly_climatology = create_weekly_climatology_forecast(train_data, valid_data.time)
# + Collapsed="false"
weekly_climatology
# + Collapsed="false"
weekly_climatology.to_netcdf(f'{PREDDIR}weekly_climatology_{res}.nc')
# + [markdown] Collapsed="false"
# # The same for higher resolutions
# + Collapsed="false"
for res in ['2.8125','1.40625']:
DATADIR = f'/media/rasp/Elements/weather-benchmark/{res}deg/'
# Load the entire dataset
z500 = xr.open_mfdataset(f'{DATADIR}geopotential/*.nc', combine='by_coords').z.sel(level=500)
t850 = xr.open_mfdataset(f'{DATADIR}temperature/*.nc', combine='by_coords').t.sel(level=850)
data = xr.merge([z500.drop('level'), t850.drop('level')])
# Load the validation subset of the data: 2017 and 2018
z500_valid = load_test_data(f'{DATADIR}geopotential', 'z')
t850_valid = load_test_data(f'{DATADIR}temperature', 't')
valid_data = xr.merge([z500_valid, t850_valid])
# Persistence forecast
persistence = []
for l in lead_times:
persistence.append(create_persistence_forecast(valid_data, int(l)))
persistence = xr.concat(persistence, dim=lead_times)
print(persistence)
persistence.to_netcdf(f'{PREDDIR}persistence_{res}.nc')
# Climatology
train_data = data.sel(time=slice(None, '2016'))
climatology = create_climatology_forecast(train_data)
print(climatology)
climatology.to_netcdf(f'{PREDDIR}climatology_{res}.nc')
# Weekly climatology
weekly_climatology = create_weekly_climatology_forecast(train_data, valid_data.time)
print(weekly_climatology)
weekly_climatology.to_netcdf(f'{PREDDIR}weekly_climatology_{res}.nc')
# + [markdown] Collapsed="false"
# # The End
| notebooks/1-climatology-persistence.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import ipyvuetify as v
from traitlets import (Any, Bool, Dict, Int, Unicode, List)
import demjson
# -
# %%HTML
# For showing mdi icons in notebook
<link href="https://cdn.jsdelivr.net/npm/@mdi/font@4.x/css/materialdesignicons.min.css" rel="stylesheet">
# # Sample components
js_headers = '''[
{
text: 'Dessert (100g serving)',
align: 'left',
sortable: false,
value: 'name',
},
{ text: 'Calories', value: 'calories' },
{ text: 'Fat (g)', value: 'fat' },
{ text: 'Carbs (g)', value: 'carbs' },
{ text: 'Protein (g)', value: 'protein' },
{ text: 'Iron (%)', value: 'iron' },
]'''
js_desserts = '''[
{
name: 'Frozen Yogurt',
calories: 159,
fat: 6.0,
carbs: 24,
protein: 4.0,
iron: '1%',
},
{
name: 'Ice cream sandwich',
calories: 237,
fat: 9.0,
carbs: 37,
protein: 4.3,
iron: '1%',
},
{
name: 'Eclair',
calories: 262,
fat: 16.0,
carbs: 23,
protein: 6.0,
iron: '7%',
},
{
name: 'Cupcake',
calories: 305,
fat: 3.7,
carbs: 67,
protein: 4.3,
iron: '8%',
},
{
name: 'Gingerbread',
calories: 356,
fat: 16.0,
carbs: 49,
protein: 3.9,
iron: '16%',
},
{
name: 'Jelly bean',
calories: 375,
fat: 0.0,
carbs: 94,
protein: 0.0,
iron: '0%',
},
{
name: 'Lollipop',
calories: 392,
fat: 0.2,
carbs: 98,
protein: 0,
iron: '2%',
},
{
name: 'Honeycomb',
calories: 408,
fat: 3.2,
carbs: 87,
protein: 6.5,
iron: '45%',
},
{
name: 'Donut',
calories: 452,
fat: 25.0,
carbs: 51,
protein: 4.9,
iron: '22%',
},
{
name: 'KitKat',
calories: 518,
fat: 26.0,
carbs: 65,
protein: 7,
iron: '6%',
},
]'''
# +
py_desserts = demjson.decode(js_desserts)
py_headers = demjson.decode(js_headers)
class MyDataTable(v.VuetifyTemplate):
headers = List(py_headers).tag(sync=True)
desserts = List(py_desserts).tag(sync=True)
template = Unicode('''
<v-layout>
<v-data-table
:headers="headers"
:items="desserts"
:items-per-page="5"
class="elevation-1"
></v-data-table>
</v-layout>''').tag(sync=True)
dataTable = MyDataTable()
dataTable
# -
# ## TreeView
js_items = '''[
{
id: 1,
name: 'Applications :',
children: [
{ id: 2, name: 'Calendar : app' },
{ id: 3, name: 'Chrome : app' },
{ id: 4, name: 'Webstorm : app' },
],
},
{
id: 5,
name: 'Documents :',
children: [
{
id: 6,
name: 'vuetify :',
children: [
{
id: 7,
name: 'src :',
children: [
{ id: 8, name: 'index : ts' },
{ id: 9, name: 'bootstrap : ts' },
],
},
],
},
{
id: 10,
name: 'material2 :',
children: [
{
id: 11,
name: 'src :',
children: [
{ id: 12, name: 'v-btn : ts' },
{ id: 13, name: 'v-card : ts' },
{ id: 14, name: 'v-window : ts' },
],
},
],
},
],
},
{
id: 15,
name: 'Downloads :',
children: [
{ id: 16, name: 'October : pdf' },
{ id: 17, name: 'November : pdf' },
{ id: 18, name: 'Tutorial : html' },
],
},
{
id: 19,
name: 'Videos :',
children: [
{
id: 20,
name: 'Tutorials :',
children: [
{ id: 21, name: 'Basic layouts : mp4' },
{ id: 22, name: 'Advanced techniques : mp4' },
{ id: 23, name: 'All about app : dir' },
],
},
{ id: 24, name: 'Intro : mov' },
{ id: 25, name: 'Conference introduction : avi' },
],
},
]'''
# +
py_items = demjson.decode(js_items)
class MyTreeView(v.VuetifyTemplate):
items = List(py_items).tag(sync=True)
template = Unicode('''
<v-layout>
<v-treeview :items="items"></v-treeview>
</v-layout>''').tag(sync=True)
treeview = MyTreeView()
treeview
# -
# ## Timeline
# +
class MyTimeline(v.VuetifyTemplate):
template = Unicode('''
<v-timeline>
<v-timeline-item>timeline item</v-timeline-item>
<v-timeline-item class="text-right">timeline item</v-timeline-item>
<v-timeline-item>timeline item</v-timeline-item>
</v-timeline>
''').tag(sync=True)
timeline = MyTimeline()
timeline
# -
# ## Navigation drawer
js_items = '''[
{ title: 'Dashboard', icon: 'mdi-view-dashboard' },
{ title: 'Photos', icon: 'mdi-image' },
{ title: 'About', icon: 'mdi-help-box' },
]
'''
# +
py_items = demjson.decode(js_items)
class MyNavigationDrawer(v.VuetifyTemplate):
items = List(py_items).tag(sync=True)
template = Unicode('''
<v-card
height="400"
width="256"
class="mx-auto"
>
<v-navigation-drawer permanent>
<v-list-item>
<v-list-item-content>
<v-list-item-title class="title">
Application
</v-list-item-title>
<v-list-item-subtitle>
subtext
</v-list-item-subtitle>
</v-list-item-content>
</v-list-item>
<v-divider></v-divider>
<v-list
dense
nav
>
<v-list-item
v-for="item in items"
:key="item.title"
link
>
<v-list-item-icon>
<v-icon>{{ item.icon }}</v-icon>
</v-list-item-icon>
<v-list-item-content>
<v-list-item-title>{{ item.title }}</v-list-item-title>
</v-list-item-content>
</v-list-item>
</v-list>
</v-navigation-drawer>
</v-card>
''').tag(sync=True)
navigation_drawer = MyNavigationDrawer()
navigation_drawer
# -
class MyIcons(v.VuetifyTemplate):
items = List(py_items).tag(sync=True)
template = Unicode('''
<v-row justify="space-around">
<v-icon>mdi-anchor</v-icon>
<v-icon>mdi-xbox-controller</v-icon>
<v-icon>mdi-watch</v-icon>
<v-icon>mdi-tilde</v-icon>
<v-icon>mdi-tennis</v-icon>
<v-icon>mdi-mouse</v-icon>
</v-row>
''').tag(sync=True)
icons = MyIcons()
icons
# ## Footer
# +
js_icons = '''[
'mdi-home',
'mdi-email',
'mdi-calendar',
'mdi-delete',
]'''
py_icons = demjson.decode(js_icons)
# -
class MyFooter(v.VuetifyTemplate):
icons = List(py_icons).tag(sync=True)
padless = Bool(False).tag(sync=True)
template = Unicode('''
<v-footer
:padless="padless"
>
<v-card
flat
tile
width="100%"
class="red lighten-1 text-center"
>
<v-card-text>
<v-btn
v-for="icon in icons"
:key="icon"
class="mx-4"
icon
>
<v-icon size="24px">{{ icon }}</v-icon>
</v-btn>
</v-card-text>
<v-divider></v-divider>
<v-card-text class="white--text">
{{ new Date().getFullYear() }} — <strong>Vuetify</strong>
</v-card-text>
</v-card>
</v-footer>
''').tag(sync=True)
footer = MyFooter()
footer
# ## Expansion panels
# +
class MyPanel(v.VuetifyTemplate):
panel = List([]).tag(sync=True)
items = Int(5).tag(sync=True)
template = Unicode('''
<v-app id="inspire">
<div>
<div class="text-center d-flex pb-4">
<v-btn @click="all">all</v-btn>
<div>{{ panel }}</div>
<v-btn @click="none">none</v-btn>
</div>
<v-expansion-panels
v-model="panel"
multiple
>
<v-expansion-panel
v-for="(item,i) in items"
:key="i"
>
<v-expansion-panel-header>Header {{ item }}</v-expansion-panel-header>
<v-expansion-panel-content>
Lorem ipsum dolor sit amet, consectetur adipiscing elit, sed do eiusmod tempor incididunt ut labore et dolore magna aliqua. Ut enim ad minim veniam, quis nostrud exercitation ullamco laboris nisi ut aliquip ex ea commodo consequat.
</v-expansion-panel-content>
</v-expansion-panel>
</v-expansion-panels>
</div>
</v-app>
''').tag(sync=True)
def vue_all(self, data):
self.panel = list(range(self.items))
def vue_none(self, data):
self.panel = []
panel_simple = MyPanel()
panel_simple
# -
trip = {
'name': '',
'location': None,
'start': None,
'end': None,
}
# +
# Not synchronizing dict entries
class MyPanelComplex(v.VuetifyTemplate):
date = Any(None, allow_none=True).tag(sync=True)
trip = Dict(trip).tag(sync=True)
locations= List(['Australia', 'Barbados', 'Chile', 'Denmark', 'Equador', 'France']).tag(sync=True)
template = Unicode('''
<v-expansion-panels>
<v-expansion-panel>
<v-expansion-panel-header>
<template v-slot:default="{ open }">
<v-row no-gutters>
<v-col cols="4">Trip name</v-col>
<v-col
cols="8"
class="text--secondary"
>
<v-fade-transition leave-absolute>
<span
v-if="open"
key="0"
>
Enter a name for the trip
</span>
<span
v-else
key="1"
>
{{ trip.name }}
</span>
</v-fade-transition>
</v-col>
</v-row>
</template>
</v-expansion-panel-header>
<v-expansion-panel-content>
<v-text-field
v-model="trip.name"
placeholder="Caribbean Cruise"
></v-text-field>
</v-expansion-panel-content>
</v-expansion-panel>
<v-expansion-panel>
<v-expansion-panel-header v-slot="{ open }">
<v-row no-gutters>
<v-col cols="4">Location</v-col>
<v-col
cols="8"
class="text--secondary"
>
<v-fade-transition leave-absolute>
<span
v-if="open"
key="0"
>
Select trip destination
</span>
<span
v-else
key="1"
>
{{ trip.location }}
</span>
</v-fade-transition>
</v-col>
</v-row>
</v-expansion-panel-header>
<v-expansion-panel-content>
<v-row no-gutters>
<div class="flex-grow-1"></div>
<v-col cols="5">
<v-select
v-model="trip.location"
:items="locations"
chips
flat
solo
></v-select>
</v-col>
<v-divider
vertical
class="mx-4"
></v-divider>
<v-col cols="3">
Select your destination of choice
<br>
<a href="javascript:void(0)">Learn more</a>
</v-col>
</v-row>
<v-card-actions>
<div class="flex-grow-1"></div>
<v-btn
text
color="secondary"
>
Cancel
</v-btn>
<v-btn
text
color="primary"
>
Save
</v-btn>
</v-card-actions>
</v-expansion-panel-content>
</v-expansion-panel>
<v-expansion-panel>
<v-expansion-panel-header v-slot="{ open }">
<v-row no-gutters>
<v-col cols="4">Start and end dates</v-col>
<v-col
cols="8"
class="text--secondary"
>
<v-fade-transition leave-absolute>
<span v-if="open">When do you want to travel?</span>
<v-row
v-else
no-gutters
style="width: 100%"
>
<v-col cols="6">Start date: {{ trip.start || 'Not set' }}</v-col>
<v-col cols="6">End date: {{ trip.end || 'Not set' }}</v-col>
</v-row>
</v-fade-transition>
</v-col>
</v-row>
</v-expansion-panel-header>
<v-expansion-panel-content>
<v-row
justify="space-around"
no-gutters
>
<v-col cols="3">
<v-menu
ref="startMenu"
:close-on-content-click="false"
:return-value.sync="trip.start"
offset-y
min-width="290px"
>
<template v-slot:activator="{ on }">
<v-text-field
v-model="trip.start"
label="Start date"
prepend-icon="event"
readonly
v-on="on"
></v-text-field>
</template>
<v-date-picker
v-model="date"
no-title
scrollable
>
<div class="flex-grow-1"></div>
<v-btn
text
color="primary"
@click="$refs.startMenu.isActive = false"
>Cancel</v-btn>
<v-btn
text
color="primary"
@click="$refs.startMenu.save(date)"
>OK</v-btn>
</v-date-picker>
</v-menu>
</v-col>
<v-col cols="3">
<v-menu
ref="endMenu"
:close-on-content-click="false"
:return-value.sync="trip.end"
offset-y
min-width="290px"
>
<template v-slot:activator="{ on }">
<v-text-field
v-model="trip.end"
label="End date"
prepend-icon="event"
readonly
v-on="on"
></v-text-field>
</template>
<v-date-picker
v-model="date"
no-title
scrollable
>
<div class="flex-grow-1"></div>
<v-btn
text
color="primary"
@click="$refs.endMenu.isActive = false"
>
Cancel
</v-btn>
<v-btn
text
color="primary"
@click="$refs.endMenu.save(date)"
>
OK
</v-btn>
</v-date-picker>
</v-menu>
</v-col>
</v-row>
</v-expansion-panel-content>
</v-expansion-panel>
</v-expansion-panels>
''').tag(sync=True)
panel_complex = MyPanelComplex()
panel_complex
# -
# ## Carousels
# +
class MyCarousel(v.VuetifyTemplate):
colors = List( ['primary',
'secondary',
'yellow darken-2',
'red',
'orange']).tag(sync=True)
template = Unicode('''
<v-carousel>
<v-carousel-item
v-for="(color, i) in colors"
:key="color"
>
<v-sheet
:color="color"
height="100%"
tile
>
<v-row
class="fill-height"
align="center"
justify="center"
>
<div class="display-3">Slide {{ i + 1 }}</div>
</v-row>
</v-sheet>
</v-carousel-item>
</v-carousel>
''').tag(sync=True)
carousel = MyCarousel()
carousel
# -
# ## Avatar
# +
class MyAvatarCard(v.VuetifyTemplate):
template = Unicode('''
<v-card
class="mx-auto"
max-width="434"
tile
>
<v-img
height="100%"
src="https://cdn.vuetifyjs.com/images/cards/server-room.jpg"
>
<v-row
align="end"
class="fill-height"
>
<v-col
align-self="start"
class="pa-0"
cols="12"
>
<v-avatar
class="profile"
color="grey"
size="164"
tile
>
<v-img src="https://cdn.vuetifyjs.com/images/profiles/marcus.jpg"></v-img>
</v-avatar>
</v-col>
<v-col class="py-0">
<v-list-item
color="rgba(0, 0, 0, .4)"
dark
>
<v-list-item-content>
<v-list-item-title class="title"><NAME></v-list-item-title>
<v-list-item-subtitle>Network Engineer</v-list-item-subtitle>
</v-list-item-content>
</v-list-item>
</v-col>
</v-row>
</v-img>
</v-card>
''').tag(sync=True)
MyAvatarCard()
# -
# ## Avatar - complex
js_messages = '''[
{
avatar: 'https://avatars0.githubusercontent.com/u/9064066?v=4&s=460',
name: '<NAME>',
title: 'Welcome to Vuetify.js!',
excerpt: 'Thank you for joining our community...',
},
{
color: 'red',
icon: 'people',
name: 'Social',
new: 1,
total: 3,
title: 'Twitter',
},
{
color: 'teal',
icon: 'local_offer',
name: 'Promos',
new: 2,
total: 4,
title: 'Shop your way',
exceprt: 'New deals available, Join Today',
},
]'''
# +
py_messages = demjson.decode(js_messages)
class MyAvatarComplex(v.VuetifyTemplate):
messages = List(py_messages).tag(sync=True)
lorem = Unicode('''
'Lorem ipsum dolor sit amet, at aliquam vivendum vel,
everti delicatissimi cu eos. Dico iuvaret debitis mel an,
et cum zril menandri. Eum in consul legimus accusam.
Ea dico abhorreant duo, quo illum minimum incorrupte no,
nostro voluptaria sea eu. Suas eligendi ius at, at nemore
equidem est. Sed in error hendrerit, in consul constituam cum.'
''').tag(sync=True)
template = Unicode('''
<v-container fluid>
<v-row justify="center">
<v-subheader>Today</v-subheader>
<v-expansion-panels popout>
<v-expansion-panel
v-for="(message, i) in messages"
:key="i"
hide-actions
>
<v-expansion-panel-header>
<v-row
align="center"
class="spacer"
no-gutters
>
<v-col
cols="4"
sm="2"
md="1"
>
<v-avatar
size="36px"
>
<img
v-if="message.avatar"
alt="Avatar"
src="https://avatars0.githubusercontent.com/u/9064066?v=4&s=460"
>
<v-icon
v-else
:color="message.color"
v-text="message.icon"
></v-icon>
</v-avatar>
</v-col>
<v-col
class="hidden-xs-only"
sm="5"
md="3"
>
<strong v-html="message.name"></strong>
<span
v-if="message.total"
class="grey--text"
>
({{ message.total }})
</span>
</v-col>
<v-col
class="text-no-wrap"
cols="5"
sm="3"
>
<v-chip
v-if="message.new"
:color="`${message.color} lighten-4`"
class="ml-0"
label
small
>
{{ message.new }} new
</v-chip>
<strong v-html="message.title"></strong>
</v-col>
<v-col
v-if="message.excerpt"
class="grey--text text-truncate hidden-sm-and-down"
>
—
{{ message.excerpt }}
</v-col>
</v-row>
</v-expansion-panel-header>
<v-expansion-panel-content>
<v-divider></v-divider>
<v-card-text v-text="lorem"></v-card-text>
</v-expansion-panel-content>
</v-expansion-panel>
</v-expansion-panels>
</v-row>
</v-container>
''').tag(sync=True)
avatar = MyAvatarComplex()
avatar
# -
# ## Banner
# +
banner = v.Banner(single_line=True,
v_slots=[{
'name': 'icon',
'children': v.Icon(children=['thumb_up'])
}, {
'name': 'actions',
'children': v.Btn(text=True, color='deep-purple accent-4', children=['Action'])
}],
children=['One line message text string with two actions on tablet / Desktop'])
banner
# -
# # Content component
container_main = v.Container(
_metadata={'mount_id': 'content-main'},
children=[
dataTable
]
)
container_main
# # Navigation component
# +
components_map = {
'DataTable': dataTable,
'Timeline': timeline,
'Treeview': treeview,
'Navigation Drawer': navigation_drawer,
'Icons': icons,
'Footer': footer,
'Expansion Panel': panel_simple,
'Expansion Panel Complex': panel_complex,
'Carousel': carousel,
'Avatar': avatar,
'Banner': banner
}
py_items = [
{ 'title': 'DataTable', 'icon': 'mdi-view-dashboard'},
{ 'title': 'Timeline', 'icon': 'mdi-image'},
{ 'title': 'Treeview', 'icon': 'mdi-help-box'},
{ 'title': 'Navigation Drawer', 'icon': 'mdi-view-dashboard'},
{ 'title': 'Icons', 'icon': 'mdi-image'},
{ 'title': 'Footer', 'icon': 'mdi-help-box'},
{ 'title': 'Expansion Panel', 'icon': 'mdi-view-dashboard'},
{ 'title': 'Expansion Panel Complex', 'icon': 'mdi-image'},
{ 'title': 'Carousel', 'icon': 'mdi-help-box'},
{ 'title': 'Avatar', 'icon': 'mdi-view-dashboard'},
{ 'title': 'Banner', 'icon': 'mdi-image'},
]
class MyNavigation(v.VuetifyTemplate):
items = List(py_items).tag(sync=True)
template = Unicode('''
<v-list
dense
nav
>
<v-list-item
v-for="item in items"
:key="item.title"
link
@click="item_click(item)"
>
<v-list-item-icon>
<v-icon>{{ item.icon }}</v-icon>
</v-list-item-icon>
<v-list-item-content>
<v-list-item-title>{{ item.title }}</v-list-item-title>
</v-list-item-content>
</v-list-item>
</v-list>
''').tag(sync=True)
def vue_item_click(self, data):
component = components_map[data['title']]
container_main.children = [component]
navigation = MyNavigation()
container_nav = v.Container(
_metadata={'mount_id': 'content-nav'},
children=[
navigation
]
)
container_nav
# -
# Check if it is working
container_main
| voila-vuetify-examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Two-Level: Gaussian Pulse 1.8π
mb_solve_json = """
{
"atom": {
"fields": [
{
"coupled_levels": [[0, 1]],
"rabi_freq_t_args": {
"n_pi": 1.8,
"centre": 0.0,
"fwhm": 1.0
},
"rabi_freq_t_func": "gaussian"
}
],
"num_states": 2
},
"t_min": -2.0,
"t_max": 10.0,
"t_steps": 120,
"z_min": -0.5,
"z_max": 1.5,
"z_steps": 100,
"interaction_strengths": [
10.0
],
"savefile": "mbs-two-gaussian-1.8pi"
}
"""
from maxwellbloch import mb_solve
mbs = mb_solve.MBSolve().from_json_str(mb_solve_json)
import numpy as np
# Check the input pulse area is correct
print('The input pulse area is {0:.3f}'.format(
np.trapz(mbs.Omegas_zt[0,0,:].real, mbs.tlist)/np.pi))
# ## Solve the Problem
Omegas_zt, states_zt = mbs.mbsolve(recalc=False)
# ## Plot Output
# +
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
sns.set_style("darkgrid")
fig = plt.figure(1, figsize=(16, 6))
ax = fig.add_subplot(111)
cmap_range = np.linspace(0.0, 1.0, 11)
cf = ax.contourf(mbs.tlist, mbs.zlist,
np.abs(mbs.Omegas_zt[0]/(2*np.pi)),
cmap_range, cmap=plt.cm.Blues)
ax.set_title('Rabi Frequency ($\Gamma / 2\pi $)')
ax.set_xlabel('Time ($1/\Gamma$)')
ax.set_ylabel('Distance ($L$)')
for y in [0.0, 1.0]:
ax.axhline(y, c='grey', lw=1.0, ls='dotted')
plt.colorbar(cf);
# -
fig, ax = plt.subplots(figsize=(16, 4))
ax.plot(mbs.zlist, mbs.fields_area()[0]/np.pi, clip_on=False)
ax.set_ylim([0.0, 8.0])
ax.set_xlabel('Distance ($L$)')
ax.set_ylabel('Pulse Area ($\pi$)');
# ## Movie
# +
# FNAME = "mb-solve-two-gaussian-0.8pi"
# +
# z_steps was 200 for movie
# +
# C = 0.1 # speed of light
# Y_MIN = 0.0 # Y-axis min
# Y_MAX = 4.0 # y-axis max
# ZOOM = 2 # level of linear interpolation
# FPS = 30 # frames per second
# ATOMS_ALPHA = 0.2 # Atom indicator transparency
# +
# FNAME_IMG = "images/" + FNAME
# FNAME_JSON = FNAME_IMG + '.json'
# with open(FNAME_JSON, "w") as f:
# f.write(mb_solve_json)
# +
# # !make-mp4-fixed-frame.py -f $FNAME_JSON -c $C --fps $FPS --y-min $Y_MIN --y-max $Y_MAX \
# # --zoom $ZOOM --atoms-alpha $ATOMS_ALPHA #--peak-line --c-line
# +
# FNAME_MP4 = FNAME_IMG + '.mp4'
# # !make-gif-ffmpeg.sh -f $FNAME_MP4 --in-fps $FPS
# +
# from IPython.display import Image
# Image(url=FNAME_MP4+'.gif', format='gif')
| docs/examples/mbs-two-gaussian-1.8pi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Classes and Objects I
#
# - [Download the lecture notes](https://philchodrow.github.io/PIC16A/content/object_oriented_programming/class_and_objects_I.ipynb).
#
#
# Like C++, Python is an **object-oriented** programming language. While the idea of object-orientation is somewhat difficult to define, a fairly general rule of thumb is that the right solution to complex problems in Python often involves creating one or more **objects**, which you can think of as bundles of related data and behaviors.
#
# A **class** defines an abstract set of possible objects sharing certain characteristics. For example, "dog" would be a good candidate for a class. There are many dogs, all of which have the same species. On the other hand, "my dog" refers to a single dog, who could be an **instance** in this class.
#
# For additional optional reading, here is a [nice, concise explanation](https://realpython.com/python3-object-oriented-programming/) (with excellently chosen examples) of object-oriented programming in Python.
#
# <br>
# <br>
# <br>
# <br>
# <br>
# <br>
# <br>
# <br>
# <br>
# <br>
# <br>
# <br>
# <br>
# <br>
# <br>
# <br>
# ## Example: Totoro
#
# A **Totoro** is a friendly forest spirit native to charming rural villages in southern Japan. For our first example, we'll make a simple `Totoro` class that models some of the Totoro's typical properties and behaviors.
# <figure class="image" style="width:50%">
# <img src="https://i0.wp.com/thespool.net/wp-content/uploads/2019/04/totoro.0.jpg?w=1200&ssl=1" alt="Totoro and two smaller Totoros alongside two happy girls, in a tree.">
# <figcaption><i>Three Totoros in their natural habitat</i></figcaption>
# </figure>
# As we can see from the picture, there are multiple kinds of Totoros: large, medium, and small. So, the first thing we should do is define a class that models all three.
class Totoro:
pass # nothing happens
# Great! Now we have a class, and can create instances of this class by calling the class with `()` parentheses.
my_neighbor = Totoro()
type(my_neighbor)
# We observe that the type of the object `my_neighbor` is the class which we have defined. But this is pretty boring -- there's not much that we can **do** with our new class. We need to add *variables* (data) and *methods* (behaviors).
#
# **Class variables** are shared between all instances of the class. For example, all Totoros have the scientific binomial nomenclature *Totoro miyazakiensis*. These variables should be assigned directly within the class definition.
#
# The `self` prefix is required to refer to local data and functions -- that is, data and functions that are only available within the object. These include class variables.
#
# Additionally, all methods need to take `self` as their first argument.
class Totoro:
# class variables: shared across all instances of the Totoro class
genus = "Totoro"
species = "miyazakiensis"
def binomial_nomenclature(self):
return(self.genus + " " + self.species)
# We can now initialize a new `Totoro` and call the `binomial_nomenclature()` method to print its biological genus and species.
my_neighbor = Totoro()
my_neighbor.genus, my_neighbor.binomial_nomenclature()
my_neighbor_2 = Totoro()
my_neighbor_2.genus
# ## `__init__`
#
# Classes can have a special `__init__()` method, which allows one to pass additional data when initializing an object. Data passed to the object this way should be **instance variables,** which may differ between different instances of the same class. For example, all `Totoro`s have a size, color, and weight, but these attributes may differ between `Totoro`s. These variables should therefore be assigned in the `__init__()` method.
#
# We'll also write a `yell` method that depends on the size of the `Totoro`.
# <figure class="image" style="width:50%">
# <img src="https://entropymag.org/wp-content/uploads/2015/01/Tonari.no_.Totoro.full_.279470.jpg" alt="Totoro flying through the air, carrying two young girls.">
# </figure>
#
# *Despite its size, the Totoro is surprisingly light*.
#
#
class Totoro:
genus = "Totoro"
species = "miyazakiensis"
def __init__(self, size, color, weight):
self.size = size
self.color = color
self.weight = weight
def yell(self):
if self.size == "large":
return("AAAAAHHHHHHHHHH!!!!!!!")
elif self.size == "medium":
return("AAAAAHHHHH!")
else:
return("aaahhhhh")
# <figure class="image" style="width:50%">
# <img src="https://data.whicdn.com/images/67713812/original.png" alt="Three Totoros, two girls, and the Catbus yelling.">
# </figure>
#
# *Illustration of the `yell()` method of the `Totoro` class*.
my_neighbor = Totoro("medium", "grey", 1)
my_neighbor.size
my_neighbor.yell()
# You should always add docstrings to both your classes and your functions.
class Totoro:
'''
A friendly forest spirit! Has size, color, and weight specified
by the user, as well as a yell method.
'''
genus = "Totoro"
species = "miyazakiensis"
def __init__(self, size, color, weight):
self.size = size
self.color = color
self.weight = weight
def yell(self):
'''
Return a yell (as a string) depending on the size of the Totoro.
Larger Totoros have louder yells.
'''
if self.size == "large":
return("AAAAAHHHHHHHHHH!!!!!!!")
elif self.size == "medium":
return("AAAAAHHHHH!")
else:
return("aaahhhhh")
# We can now get help on both the overall class and the inidividual methods:
# ?Totoro
# ?Totoro.yell
# ---
# ## Getters and Setters?
#
# In many other languages, it's recommended to use *getters* and *setters* in order to access and modify instance variables. For example, we might write functions like `Totoro.get_size()` and `Totoro.set_size()` in order to modify the `size` variable after a `Totoro` object has been created.
#
# In Python, however, this is generally unnecessary. The reason is that data encapsulation in other object-oriented languages requires the use of private instance variables. However, the broadly-used practice in Python is to use public instance variables, in which case direct access is no problem.
my_neighbor.size
my_neighbor.size = "small"
my_neighbor.size
# This [page on getters, setters, and the `@property` decorator](https://www.python-course.eu/python3_properties.php) gives a useful overview of these topics. While private instance variables do have their uses, especially in production code, we won't discuss them further in this course.
| content/object_oriented_programming/class_and_objects_I.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/hihunjin/Code-snippet-for-everything/blob/main/pandas/pandas_add_column.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="8dWIb4aM7hNK"
import pandas as pd
# + id="8YqHREY978Bo" colab={"base_uri": "https://localhost:8080/", "height": 175} outputId="1b7606ac-7798-40b4-811d-d3842b43b84a"
data = {'Name': ['Jai', 'Princi', 'Gaurav', 'Anuj'],
'Height': [5.1, 6.2, 5.1, 5.2],
'Qualification': ['Msc', 'MA', 'Msc', 'Msc']}
df = pd.DataFrame(data)
df
# + colab={"base_uri": "https://localhost:8080/", "height": 175} id="NKA46UeQ7jGV" outputId="946fd021-bc65-4393-c6da-ef242218dc66"
df['Address'] = [""]*len(df)
df
# + colab={"base_uri": "https://localhost:8080/", "height": 175} id="y1dgobeb7nsv" outputId="aa6c6f97-7808-4f88-909d-90cb8174e811"
address = ['Delhi', 'Bangalore', 'Chennai', 'Patna']
df['Address'] = address
df
# + id="mWY43GE1_Ksi"
| pandas/pandas_add_column.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/StanleyLiangYork/NLP_deepLearning/blob/master/text_predict.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="E56_uWzJ0l8u" colab_type="code" colab={}
import tensorflow as tf
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.layers import Embedding, LSTM, Dense, Bidirectional
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.models import Sequential
from tensorflow.keras.optimizers import Adam
import numpy as np
# + id="uzzgE6B50wE_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="93305bf8-ebf8-4baf-b354-86fa2c950d5c"
tokenizer = Tokenizer()
data="In the town of Athy one <NAME> \n Battered away til he hadnt a pound. \nHis father died and made him a man again \n Left him a farm and ten acres of ground. \nHe gave a grand party for friends and relations \nWho didnt forget him when come to the wall, \nAnd if youll but listen Ill make your eyes glisten \nOf the rows and the ructions of Lanigans Ball. \nMyself to be sure got free invitation, \nFor all the nice girls and boys I might ask, \nAnd just in a minute both friends and relations \nWere dancing round merry as bees round a cask. \n<NAME>, that nice little milliner, \nShe tipped me a wink for to give her a call, \nAnd I soon arrived with Peggy McGilligan \nJust in time for Lanigans Ball. \nThere were lashings of punch and wine for the ladies, \nPotatoes and cakes; there was bacon and tea, \nThere were the Nolans, Dolans, OGradys \nCourting the girls and dancing away. \nSongs they went round as plenty as water, \nThe harp that once sounded in Taras old hall,\nSweet Nelly Gray and The Rat Catchers Daughter,\nAll singing together at Lanigans Ball. \nThey were doing all kinds of nonsensical polkas \nAll round the room in a whirligig. \nJulia and I, we banished their nonsense \nAnd tipped them the twist of a reel and a jig. \nAch mavrone, how the girls got all mad at me \nDanced til youd think the ceiling would fall. \nFor I spent three weeks at Brooks Academy \nLearning new steps for Lanigans Ball. \nThree long weeks I spent up in Dublin, \nThree long weeks to learn nothing at all,\n Three long weeks I spent up in Dublin, \nLearning new steps for Lanigans Ball. \nShe stepped out and I stepped in again, \nI stepped out and she stepped in again, \nShe stepped out and I stepped in again, \nLearning new steps for Lanigans Ball. \nBoys were all merry and the girls they were hearty \nAnd danced all around in couples and groups, \nTil an accident happened, young <NAME> \nPut his right leg through miss Finnertys hoops. \nPoor creature fainted and cried Meelia murther, \nCalled for her brothers and gathered them all. \nCarmody swore that hed go no further \nTil he had satisfaction at Lanigans Ball. \nIn the midst of the row miss Kerrigan fainted, \nHer cheeks at the same time as red as a rose. \nSome of the lads declared she was painted, \nShe took a small drop too much, I suppose. \nHer sweetheart, <NAME>, so powerful and able, \nWhen he saw his fair colleen stretched out by the wall, \nTore the left leg from under the table \nAnd smashed all the Chaneys at Lanigans Ball. \nBoys, oh boys, twas then there were runctions. \nMyself got a lick from big Phelim McHugh. \nI soon replied to his introduction \nAnd kicked up a terrible hullabaloo. \nOld Casey, the piper, was near being strangled. \nThey squeezed up his pipes, bellows, chanters and all. \nThe girls, in their ribbons, they got all entangled \nAnd that put an end to Lanigans Ball."
corpus = data.lower().split("\n")
tokenizer.fit_on_texts(corpus)
total_words = len(tokenizer.word_index) + 1
print(tokenizer.word_index)
print(total_words)
# + id="wW5QFtBZ05Jx" colab_type="code" colab={}
input_sequences = []
for line in corpus:
token_list = tokenizer.texts_to_sequences([line])[0]
for i in range(1, len(token_list)):
n_gram_sequence = token_list[:i+1]
input_sequences.append(n_gram_sequence)
# pad sequences
max_sequence_len = max([len(x) for x in input_sequences])
input_sequences = np.array(pad_sequences(input_sequences, maxlen=max_sequence_len, padding='pre'))
# create predictors and label
xs, labels = input_sequences[:,:-1],input_sequences[:,-1]
ys = tf.keras.utils.to_categorical(labels, num_classes=total_words)
# + id="gZC3yM8W1cgU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 159} outputId="1558ba72-42d9-4ba4-a69c-d0f5cc875848"
print(tokenizer.word_index['in'])
print(tokenizer.word_index['the'])
print(tokenizer.word_index['town'])
print(tokenizer.word_index['of'])
print(tokenizer.word_index['athy'])
print(tokenizer.word_index['one'])
print(tokenizer.word_index['jeremy'])
print(tokenizer.word_index['lanigan'])
# + id="RBGR3uP91kb_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 230} outputId="7d0425b9-cefa-4439-e7ff-64ebb9cdfc01"
print(xs[6])
print(ys[6])
# + id="A4uPJVed13CF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 230} outputId="f8efadfd-e66e-440e-f893-624f4faec545"
print(xs[5])
print(ys[5])
# + id="lMpucTQ62ORO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="2c7a69e6-cc45-4f66-e148-909a8d3096aa"
model = Sequential()
model.add(Embedding(total_words, 64, input_length=max_sequence_len-1))
model.add(Bidirectional(LSTM(20)))
model.add(Dense(total_words, activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
history = model.fit(xs, ys, epochs=500, verbose=1)
# + id="kHYQmqPO5cps" colab_type="code" colab={}
import matplotlib.pyplot as plt
def plot_graphs(history, string):
plt.plot(history.history[string])
plt.xlabel("Epochs")
plt.ylabel(string)
plt.show()
# + id="5vC2OVcp5gb5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="f4a91aac-2cab-4484-ff52-2d54e86d43c9"
plot_graphs(history, 'accuracy')
# + id="w9O-z9MM5kBj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 55} outputId="a9572a7a-1b4b-4585-b89e-2b06a498b165"
seed_text = "<NAME>"
next_words = 100
for _ in range(next_words):
token_list = tokenizer.texts_to_sequences([seed_text])
token_list = pad_sequences(token_list, maxlen=max_sequence_len-1, padding='pre')
predicted = model.predict_classes(token_list, verbose=0)
output_word = ""
for word, index in tokenizer.word_index.items():
if index == predicted:
output_word = word
break
seed_text += " " + output_word
print(seed_text)
# + [markdown] id="fMjMdb93OB8R" colab_type="text"
# Predict irish poem
# + id="Laa9N3ikJ_wH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 212} outputId="cf284f1d-922b-4b96-a32e-78a3937b6f4c"
# !wget --no-check-certificate \
# https://storage.googleapis.com/laurencemoroney-blog.appspot.com/irish-lyrics-eof.txt \
# -O /tmp/irish-lyrics-eof.txt
# + id="acm14iuNJ_9E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="4f44718e-ff2b-4a75-dc9c-03a91622b3c3"
tokenizer = Tokenizer()
data = open('/tmp/irish-lyrics-eof.txt').read()
corpus = data.lower().split("\n")
tokenizer.fit_on_texts(corpus)
total_words = len(tokenizer.word_index) + 1
print(tokenizer.word_index)
print(total_words)
# + id="EJaRBfVUOlaK" colab_type="code" colab={}
input_sequences = []
for line in corpus:
token_list = tokenizer.texts_to_sequences([line])[0]
for i in range(1, len(token_list)):
n_gram_sequence = token_list[:i+1]
input_sequences.append(n_gram_sequence)
# pad sequences
max_sequence_len = max([len(x) for x in input_sequences])
input_sequences = np.array(pad_sequences(input_sequences, maxlen=max_sequence_len, padding='pre'))
# create predictors and label
xs, labels = input_sequences[:,:-1],input_sequences[:,-1]
ys = tf.keras.utils.to_categorical(labels, num_classes=total_words)
# + id="5bMAXKo6PP94" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 159} outputId="46b790a5-b220-4b26-8d7b-88102c5b2781"
print(tokenizer.word_index['in'])
print(tokenizer.word_index['the'])
print(tokenizer.word_index['town'])
print(tokenizer.word_index['of'])
print(tokenizer.word_index['athy'])
print(tokenizer.word_index['one'])
print(tokenizer.word_index['jeremy'])
print(tokenizer.word_index['lanigan'])
# + id="1NsYdOGIPbhv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 88} outputId="f800f5ac-1923-4e06-a834-d7bc52a4351f"
print(xs[7])
print(ys[7])
print(xs[8])
print(ys[8])
# + id="L0teT9PAP65h" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 55} outputId="81a7e379-abfb-497e-8320-45130b397eaa"
print(tokenizer.word_index)
# + id="YaQZWo9AQDa6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="b94ee009-654c-4e88-f594-f9d57e4b80d7"
model = Sequential()
model.add(Embedding(total_words, 120, input_length=max_sequence_len-1))
model.add(Bidirectional(LSTM(180)))
model.add(Dense(total_words, activation='softmax'))
adam = Adam(lr=0.008)
model.compile(loss='categorical_crossentropy', optimizer=adam, metrics=['accuracy'])
#earlystop = EarlyStopping(monitor='val_loss', min_delta=0, patience=5, verbose=0, mode='auto')
history = model.fit(xs, ys, epochs=100, verbose=1)
# + id="ehwoBsO2ZL7Y" colab_type="code" colab={}
import matplotlib.pyplot as plt
def plot_graphs(history, string):
plt.plot(history.history[string])
plt.xlabel("Epochs")
plt.ylabel(string)
plt.show()
# + id="qfTOHOMlZPWc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="e65b6834-813c-439d-9596-7a116a61479b"
plot_graphs(history, 'accuracy')
# + [markdown] id="zXkKS2zFuq8X" colab_type="text"
# Character based RNN sequence generation
# + id="oxfSFa06uze1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="8aae0608-e8e8-4ecb-8c11-64d425a635e9"
seed_text = "I've got a bad feeling about this"
next_words = 120
for _ in range(next_words):
token_list = tokenizer.texts_to_sequences([seed_text])
token_list = pad_sequences(token_list, maxlen=max_sequence_len-1, padding='pre')
predicted = model.predict_classes(token_list, verbose=0)
output_word = ""
for word, index in tokenizer.word_index.items():
if index == predicted:
output_word = word
break
seed_text += " " + output_word
print(seed_text)
# + id="RxuV2TGbup9s" colab_type="code" colab={}
import os
import time
# + id="LmHfaLKlvzL8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="43ef6565-9517-4216-8d18-9922246bbea9"
path_to_file = tf.keras.utils.get_file('/content/shakespeare.txt', 'https://storage.googleapis.com/download.tensorflow.org/data/shakespeare.txt')
# + id="GC7Pz-95v7hH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="85cac436-0a0c-4721-e4ff-39304093626a"
# Read, then decode for py2 compat.
text = open(path_to_file, 'rb').read().decode(encoding='utf-8')
# length of text is the number of characters in it
print ('Length of text: {} characters'.format(len(text)))
# + id="fZGEeHLnwfcR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="7d4d915f-70ac-4f6b-f341-2b96d670abf1"
# Take a look at the first 20 characters in text
print(text[:20])
# + id="Cwx6OWwowqLr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="5019b8ef-de09-42b5-bfba-fdd3e76d38b7"
# The unique characters in the file
vocab = sorted(set(text))
print ('{} unique characters'.format(len(vocab)))
# + [markdown] id="hWRbRZlYw_y3" colab_type="text"
# Vectorize the text -two lookup tables
#
# * mapping characters to numbers
# * mapping numbers to characters
#
#
# + id="C801tNLrw2eT" colab_type="code" colab={}
# Creating a mapping from unique characters to indices
# dict for character to number index
char2idx = {u:i for i, u in enumerate(vocab)}
# numpy version of characters
idx2char = np.array(vocab)
# mapping each character to a number index
text_as_int = np.array([char2idx[c] for c in text])
# + id="rXgk7TxB-cmW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="4620ea64-5ab6-4b81-8ec6-ff44214d6505"
print(text[:10])
print(text_as_int[:10]) # one character --> one number
# + id="DND6VSIR_Y3y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 426} outputId="3c1cbefa-304e-4804-bd0d-f212a77ee694"
print('{')
for char,_ in zip(char2idx, range(20)):
print(' {:4s}: {:3d},'.format(repr(char), char2idx[char]))
print(' ...\n}')
# + id="h9czclgM_8uy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="9dfdaf97-bbcd-4390-8124-91d65968778b"
# Show how the first 13 characters from the text are mapped to integers
print ('{} ---- characters mapped to int ---- > {}'.format(text[:13], text_as_int[:13]))
# + [markdown] id="ntQ7j_UzCiwo" colab_type="text"
# For each input sequence, the corresponding targets contain the same length of text, except shifted one character to the right.
# e.g. Hell --> ello <p>
# So we break the text into chunks of seq_length+1 (holds X --> Y) <P>
# Use the tf.data.Dataset.from_tensor_slices function to convert the text vector into a stream of character indices
# + id="VILZWhWLALSl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 106} outputId="a024ffc9-d9ef-4fb2-883f-6828af1a2987"
# The maximum length sentence we want for a single input in characters
seq_length = 100
examples_per_epoch = len(text)//(seq_length+1)
# Create training examples / targets
char_dataset = tf.data.Dataset.from_tensor_slices(text_as_int)
for i in char_dataset.take(5):
print('{} ----> {}'.format(i, idx2char[i]))
# + [markdown] id="dCo7K67eDO63" colab_type="text"
# The batch method lets us easily convert these individual characters to sequences of the desired size.
# + id="s49u0f5JCUx8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 266} outputId="fff507f3-d8e6-4d1c-962b-3bf5db5c767a"
print(text[:200])
# + id="ib50wSSHEFsv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 106} outputId="2fc09f38-2a49-421f-d6af-81ed4f2ab77b"
# devide the whole char sequence into chunks with pre-set batch size
sequences = char_dataset.batch(seq_length+1, drop_remainder=True)
for item in sequences.take(5):
print(repr(''.join(idx2char[item])))
# + id="I11wrPSHExMk" colab_type="code" colab={}
# get the input sequence and the label sequence
def split_input_target(chunk):
input_text = chunk[:-1]
target_text = chunk[1:]
return input_text, target_text
dataset = sequences.map(split_input_target)
# + id="1AjeNEVZHAcg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 88} outputId="567571d3-83c5-41a8-9c39-b7668c996ea6"
# show the first two input sequence and target
for input_example, target_example in dataset.take(2):
print ('Input data: ', repr(''.join(idx2char[input_example.numpy()])))
print ('Target data:', repr(''.join(idx2char[target_example.numpy()])))
# + [markdown] id="24wjtmPFH553" colab_type="text"
# Each index of these vectors are processed as one time step. For the input at time step 0, the model receives the index for "F" and trys to predict the index for "i" as the next character. At the next timestep, it does the same thing but the RNN considers the previous step context in addition to the current input character.
# + id="DpF8g3NvHsGZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 284} outputId="a1b0fc02-1eb1-4cca-9457-c93f9121905b"
for i, (input_idx, target_idx) in enumerate(zip(input_example[:5], target_example[:5])):
print("Step {:4d}".format(i))
print(" input: {} ({:s})".format(input_idx, repr(idx2char[input_idx])))
print(" expected output: {} ({:s})".format(target_idx, repr(idx2char[target_idx])))
# + [markdown] id="z-a42ln-UgRj" colab_type="text"
# Create training batches
# + id="Z9kJzlaWUbJw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="2b36be1c-044b-40fb-9bce-122ef0e0e4ec"
# Batch size
BATCH_SIZE = 64
# Buffer size to shuffle the dataset
# (TF data is designed to work with possibly infinite sequences,
# so it doesn't attempt to shuffle the entire sequence in memory. Instead,
# it maintains a buffer in which it shuffles elements).
BUFFER_SIZE = 10000
dataset = dataset.shuffle(BUFFER_SIZE).batch(BATCH_SIZE, drop_remainder=True)
dataset
# + [markdown] id="nogtnXC8U2iB" colab_type="text"
# Build the RNN model
# + id="tDMs72vWUskS" colab_type="code" colab={}
# Length of the vocabulary in chars
vocab_size = len(vocab) # 65 unique chars
# The embedding dimension
embedding_dim = 256
# Number of RNN units
rnn_units = 1024
# + id="mC_NkH77U5kb" colab_type="code" colab={}
def build_model(vocab_size, embedding_dim, rnn_units, batch_size):
model = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size, embedding_dim,
batch_input_shape=[batch_size, None]),
tf.keras.layers.GRU(rnn_units,
return_sequences=True,
stateful=True,
recurrent_initializer='glorot_uniform'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(vocab_size)
])
return model
# + id="Jzs0Ys9Wk4L_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 301} outputId="000cc9fc-3cea-4d1c-9418-430201ebd852"
model = build_model(
vocab_size = len(vocab),
embedding_dim=embedding_dim,
rnn_units=rnn_units,
batch_size=BATCH_SIZE)
model.summary()
# + id="sMh3UTPDrOl7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="bac8e67b-4873-4dde-c7b5-36f33a1cb8ac"
for input_example_batch, target_example_batch in dataset.take(1):
example_batch_predictions = model(input_example_batch)
print(f'input batch shape: {input_example_batch.shape}, vocab-char size: {len(vocab)}')
print(example_batch_predictions.shape, "= ( batch_size, sequence_length, vocab_size)")
# + id="G9nJ_-m7uh0n" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 657} outputId="481c69a9-2838-4e96-9939-30353c1abea5"
# get the predictioin of 1 data point, one batch has 64 points
sampled_indices = tf.random.categorical(example_batch_predictions[0], num_samples=1)
print(f'the input seq:\n {np.array(input_example_batch[0])}')
print(f'the output predict:\n {example_batch_predictions[0]}, \n total predict: {len(example_batch_predictions[0])}, \n equal to the length of the input sequence')
# reduce to vector
sampled_indices = tf.squeeze(sampled_indices,axis=-1).numpy()
print(f'the index with highest logit output: {(sampled_indices)}')
print("Input: \n",f'{"".join(idx2char[input_example_batch[0]])}')
print("\n")
print("Next Char Predictions: \n", repr("".join(idx2char[sampled_indices])))
# + id="aqXtKJI1k47h" colab_type="code" colab={}
def loss(labels, logits):
return tf.keras.losses.sparse_categorical_crossentropy(labels, logits, from_logits=True)
# + id="o1SwevCyPHQA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="e32030d4-510c-44e1-82d2-637e63e126ee"
print(f'Y(batch, sequence):{target_example_batch.shape}, Y-hat(batch, sequence, vocab):{example_batch_predictions.shape}')
example_batch_loss = loss(target_example_batch, example_batch_predictions)
print("scalar_loss: ", example_batch_loss.numpy().mean())
# + id="LNk47D6Tloe1" colab_type="code" colab={}
model.compile(optimizer='adam', loss=loss)
# + id="8Eh2FyRQl2KP" colab_type="code" colab={}
# Directory where the checkpoints will be saved
checkpoint_dir = './training_checkpoints'
# Name of the checkpoint files
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt_{epoch}")
checkpoint_callback=tf.keras.callbacks.ModelCheckpoint(
filepath=checkpoint_prefix,
save_weights_only=True)
# + id="Uq85AiFDl3CS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 728} outputId="5cbbc80e-cea1-4547-9edd-fba6c6eda935"
EPOCHS = 20
history = model.fit(dataset, epochs=EPOCHS, callbacks=[checkpoint_callback])
# + id="IkY0paXxnPcV" colab_type="code" colab={}
import matplotlib.pyplot as plt
def plot_graphs(history, string):
plt.plot(history.history[string])
plt.xlabel("Epochs")
plt.ylabel(string)
plt.show()
# + id="Qbsh7TutnVDV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="a4c42b88-3af9-4f6b-9971-ce562c4919a4"
plot_graphs(history, 'loss')
# + id="eMLCaHnNmE_U" colab_type="code" colab={}
model_predict = build_model(vocab_size, embedding_dim, rnn_units, batch_size=1)
model_predict.load_weights(tf.train.latest_checkpoint(checkpoint_dir))
model_predict.build(tf.TensorShape([1, None]))
# + id="B74K5n8NWtQ1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="4018272a-b370-49c7-c9bc-30f00e6c0d98"
num_generate = 2000
start_string = u"ROMEO: " # 7 chars
# char to sequence
input_eval = [char2idx[s] for s in start_string]
input_eval = tf.expand_dims(input_eval, 0)
print(input_eval.numpy())
text_gen = []
# higer temp more rush text, lower temp more predictable text
temp = 0.8
model_predict.reset_states()
for i in range(num_generate):
predictions = model_predict(input_eval)
predictions = tf.squeeze(predictions, 0)
predictions = predictions / temp
predicted_id = tf.random.categorical(predictions, num_samples=1)[-1,0].numpy()
input_eval = tf.expand_dims([predicted_id], 0)
text_gen.append(idx2char[predicted_id])
output_text = start_string+"".join(text_gen)
print(output_text)
# + id="-N85KvSame2A" colab_type="code" colab={}
def generate_text(model, start_string):
# Evaluation step (generating text using the learned model)
# Number of characters to generate
num_generate = 1000
# Converting our start string to numbers (vectorizing)
input_eval = [char2idx[s] for s in start_string]
input_eval = tf.expand_dims(input_eval, 0)
# Empty string to store our results
text_generated = []
# Low temperatures results in more predictable text.
# Higher temperatures results in more surprising text.
# Experiment to find the best setting.
temperature = 1.0
# Here batch size == 1
model.reset_states()
for i in range(num_generate):
predictions = model(input_eval)
# remove the batch dimension
predictions = tf.squeeze(predictions, 0)
# using a categorical distribution to predict the character returned by the model
predictions = predictions / temperature
predicted_id = tf.random.categorical(predictions, num_samples=1)[-1,0].numpy()
# We pass the predicted character as the next input to the model
# along with the previous hidden state
input_eval = tf.expand_dims([predicted_id], 0)
text_generated.append(idx2char[predicted_id])
return (start_string + ''.join(text_generated))
# + id="8Qnv2J9KmfwN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 604} outputId="a6cb8d90-0589-4363-c0ff-a9119e1410c8"
print(generate_text(model, start_string=u"ROMEO: "))
# + [markdown] id="Ilh1pY0vgAcD" colab_type="text"
# Customize the training loss
# + id="F34rhAkwgCzB" colab_type="code" colab={}
model_custom = build_model(
vocab_size = len(vocab),
embedding_dim=embedding_dim,
rnn_units=rnn_units,
batch_size=BATCH_SIZE)
# + id="kB_0lXGTgtvI" colab_type="code" colab={}
optimizer = tf.keras.optimizers.Adam(learning_rate=0.01)
# + [markdown] id="hanYv585g474" colab_type="text"
# Build the optimized loss function
# + id="wlS3YCcGg136" colab_type="code" colab={}
@tf.function
def train_step(inp, target):
with tf.GradientTape() as tape:
predictions = model_custom(inp)
loss = tf.reduce_mean(
tf.keras.losses.sparse_categorical_crossentropy(
target, predictions, from_logits=True))
grads = tape.gradient(loss, model_custom.trainable_variables) # get the gradients
# pipeline the gradients to the optimizer
optimizer.apply_gradients(zip(grads, model_custom.trainable_variables))
return loss
# + id="sMscUFeivocQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="92562f90-e7cc-4482-83f0-ca3e5729fb0f"
import time
# Training step
EPOCHS = 20
record = []
for epoch in range(EPOCHS):
start = time.time()
# initializing the hidden state at the start of every epoch
# initally hidden is None
hidden = model_custom.reset_states()
for (batch_n, (inp, target)) in enumerate(dataset):
loss = train_step(inp, target)
if batch_n % 100 == 0:
template = 'Epoch {} Batch {} Loss {}'
print(template.format(epoch+1, batch_n, loss))
# saving (checkpoint) the model every 5 epochs
if (epoch + 1) % 5 == 0:
model_custom.save_weights(checkpoint_prefix.format(epoch=epoch))
print ('Epoch {} Loss {:.4f}'.format(epoch+1, loss))
print ('Time taken for 1 epoch {:.2f} sec\n'.format(time.time() - start))
record.append(loss)
model_custom.save_weights(checkpoint_prefix.format(epoch=epoch))
history = {'loss':record}
# + id="U-uS5-IKpR7u" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="d7760312-8f21-4c9b-d3ef-93d5df88f21f"
import matplotlib.pyplot as plt
plt.plot(history['loss'])
plt.xlabel("Epochs")
plt.ylabel('loss')
plt.show()
# + id="CBkTuSCFpbOw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 464} outputId="b948ad3e-b32a-4e07-aedb-a81d75a8f289"
model_predict = build_model(vocab_size, embedding_dim, rnn_units, batch_size=1)
model_predict.load_weights(tf.train.latest_checkpoint(checkpoint_dir))
model_predict.build(tf.TensorShape([1, None]))
num_generate = 1000
start_string = u"ROMEO: " # 7 chars
# char to sequence
input_eval = [char2idx[s] for s in start_string]
input_eval = tf.expand_dims(input_eval, 0)
print(input_eval.numpy())
text_gen = []
# higer temp more rush text, lower temp more predictable text
temp = 1.0
model_predict.reset_states()
for i in range(num_generate):
predictions = model_predict(input_eval)
predictions = tf.squeeze(predictions, 0)
predictions = predictions / temp
predicted_id = tf.random.categorical(predictions, num_samples=1)[-1,0].numpy()
input_eval = tf.expand_dims([predicted_id], 0)
text_gen.append(idx2char[predicted_id])
output_text = start_string+"".join(text_gen)
print(output_text)
# + [markdown] id="oWXEEoMjE2sT" colab_type="text"
# Shakespeare problem
# + id="3l4xH36qE4UA" colab_type="code" colab={}
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.layers import Embedding, LSTM, Dense, Dropout, Bidirectional
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.models import Sequential
from tensorflow.keras.optimizers import Adam
from tensorflow.keras import regularizers
import tensorflow.keras.utils as ku
import numpy as np
# + id="e8h6gKWTFBRx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 212} outputId="71d889e6-d525-4bd9-ea05-dd6288694175"
tokenizer = Tokenizer()
# !wget --no-check-certificate \
# https://storage.googleapis.com/laurencemoroney-blog.appspot.com/sonnets.txt \
# -O /content/sonnets.txt
data = open('/content/sonnets.txt').read()
corpus = data.lower().split("\n")
tokenizer.fit_on_texts(corpus)
total_words = len(tokenizer.word_index) + 1
# create input sequences using list of tokens
input_sequences = []
for line in corpus:
token_list = tokenizer.texts_to_sequences([line])[0]
for i in range(1, len(token_list)):
n_gram_sequence = token_list[:i+1]
input_sequences.append(n_gram_sequence)
# pad sequences
max_sequence_len = max([len(x) for x in input_sequences])
input_sequences = np.array(pad_sequences(input_sequences, maxlen=max_sequence_len, padding='pre'))
# create predictors and label
predictors, label = input_sequences[:,:-1],input_sequences[:,-1]
label = ku.to_categorical(label, num_classes=total_words)
# + id="kN28sJFHFOrw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="fa55e654-c9f2-4e2e-a837-6aa4d4135cad"
model = Sequential()
model.add(Embedding(total_words, 100, input_length=max_sequence_len-1))
model.add(Bidirectional(LSTM(256, return_sequences = True)))
model.add(Dropout(0.2))
model.add(LSTM(128))
model.add(Dense(total_words/2, activation='relu', kernel_regularizer=regularizers.l2(0.01)))
model.add(Dense(total_words, activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
print(model.summary())
# + id="3vmBJjpJFZvt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="e852c6a1-0ae5-449e-a623-c435827b7615"
history = model.fit(predictors, label, epochs=200, verbose=1)
# + id="NqBw39FSI3s1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 607} outputId="cf44a7af-e695-408d-8174-f97c8a455ab7"
import matplotlib.pyplot as plt
acc = history.history['accuracy']
loss = history.history['loss']
epochs = range(len(acc))
plt.figure(figsize=(10,10))
plt.subplot(1,2,1)
plt.plot(epochs, acc, 'b', label='Training accuracy')
plt.title('Training accuracy')
plt.legend()
plt.subplot(1,2,2)
plt.plot(epochs, loss, 'b', label='Training Loss')
plt.title('Training loss')
plt.legend()
plt.show()
# + id="nDnUmW1Cb3dM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 55} outputId="28e5cf53-ab19-4616-e1fa-daf9f5fe87cc"
seed_text = "Help me <NAME>, you're my only hope"
next_words = 100
for _ in range(next_words):
token_list = tokenizer.texts_to_sequences([seed_text])
token_list = pad_sequences(token_list, maxlen=max_sequence_len-1, padding='pre')
predicted = model.predict_classes(token_list, verbose=0)
output_word = ""
for word, index in tokenizer.word_index.items():
if index == predicted:
output_word = word
break
seed_text += " " + output_word
print(seed_text)
| text_predict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example Capacity Test using pvcaptest
#
# The captest module of the Captest package contains the `CapData` class and a few top level functions. `CapData` objects hold simulated data from PVsyst (or other simulation) or measured data from a DAS or SCADA system and provide methods for loading, filtering, visualizing and regressing.
#
# This example goes through typical steps of performing a capacity test following the ASTM E2848 standard using the Captest package.
#
# ## Imports
# +
# %matplotlib inline
import pandas as pd
# import captest as pvc
from captest import capdata as pvc
from bokeh.io import output_notebook, show
# uncomment below two lines to use cptest.scatter_hv in notebook
import holoviews as hv
hv.extension('bokeh')
#if working offline with the CapData.plot() method may fail
#run 'export BOKEH_RESOURCES=inline' at the command line before
#running the jupyter notebook
output_notebook()
# -
# ## Load and Plot Measured Data
#
# We begin by instantiating a `CapData` object, which we will use to load and store the measured data. In this example we will calculate reporting conditions from the measured data, so we load and filter the measured data first.
das = pvc.CapData('das')
# The `load_data` method by default will look for and attempt to load all files ending with '.csv' in a 'data' folder. In this case we have a single file and provide the filename, so only the file specified is loaded.
das.load_data(fname='example_meas_data.csv', source='AlsoEnergy')
# The `load_data` method loads the data into a pandas DataFrame, which it assigns to the `data` attribute of the `CapData` object. Here we use the pandas DataFrame `head` method to return the first three rows.
das.data.head(3)
# In addition to loading data, by default the `load_data` method calls the `group_columns` method, which attempts to infer the type of measurement recorded in each column of the data. For each inferred measurement type, `group_columns` creates an abbreviated name and a list of columns that contain measurements of that type. This information is stored in a python [dictionary](https://docs.python.org/3/tutorial/datastructures.html#dictionaries) where the abbreviated names are the keys and the corresponding values are the lists of columns. The python dictionary created by `group_columns` is stored in the `column_groups` attribute.
#
# The `review_column_groups` method prints the `group_columns` dictionary in an easy to read format to facilitate checking the grouping and identifying which key is linked to which group.
das.review_column_groups()
# The `view` method uses the dictionary stored in the `column_groups` attribute to allow easy access to columns of data of a certain type without renaming columns or typing long column names. The `column_groups` dictionary also enables much of the functionality of `CapData` methods to perform common capacity testing tasks, like generating scatter plots, filtering data, and performing regressions, with minimal user input.
das.view('irr-poa-').iloc[100:103, :]
# pvcaptest does not attempt to determine which columns of data or groups of columns are the data to be used in the regressions. The link between regression variables and the imported data is made by a dictionary stored in the `regression_cols` attribute. pvcaptest provides the convience method `set_regression_cols` for this purpose. `regression_cols` should be set immediately after loading data as many other `CapData` methods rely on this attribute.
das.set_regression_cols(power='-mtr-', poa='irr-poa-', t_amb='temp-amb-', w_vel='wind--')
# Once the regression columns are set, the `rview` method, similar to the `view` method, will return the data for each type of sensor identified in the `column_groups` attribute. The difference is that you pass `rview` one of the following:
# - any of 'power', 'poa', 't_amb', 'w_vel'
# - a list of some subset of any of the previous four strings
# - 'all' to return data for all four
#
# Here we are again accessing the same POA irradiance data as above with `view`.
das.rview('poa').iloc[100:103, :]
# For datasets that have multiple measurements of the same value, like the two POA irradiance measurements in this sample data, these values must be aggregated prior to filtering or regressing the data. The `agg_sensors` method provides a convient way to do this for all the groups of measurements in `column_groups` in one step.
#
# The desired aggregations are specified by passing a dictionary to the `agg_map` argument where the keys are groups from `column_groups` and the values are aggregation functions. Here we are using string functions that are recognized by pandas. Most of the common aggregation functions (mean, median, max, sum, min, etc.) are available as string functions. If you would like to apply a different aggregation function, please refer to the pandas documentation for `DataFrame.agg`. By default, the `agg_sensors` method adds a new column to the dataframe in the `data` attribute for the results of each aggregation and copies over the `data_filtered` attribute with the new dataframe.
#
# There is a also a method, `filter_sensors`, for filtering data on comparisons between measurements of the same value described below.
das.agg_sensors(agg_map={'-inv-':'sum', 'irr-poa-':'mean', 'temp-amb-':'mean', 'wind--':'mean'},
inv_sum_vs_power=False)
# The `plot` method creates a group of time series plots that are useful for visually inspecting the imported data.
#
# `plot` uses the structure of the `column_group` attribute to create a layout of plots. A single plot is generated for each measurement type and each column with measurements of that type are plotted as a separate line on the plot. In this example there are two different weather stations, which each have pyranometers measuring plane of array and global horizontal irradiance. This arrangement of sensors results in two plots which each have two lines.
das.plot(marker='line', width=900, height=250, ncols=1)
# ## Filtering Measured Data
# The `CapData` class provides a number of convience methods to apply filtering steps as defined in ASTM E2848. The following section demonstrates the use of the more commonly used filtering steps to remove measured data points.
# Uncomment and run to copy over the filtered dataset with the unfiltered data.
das.reset_filter()
# A common first step is to review the scatter plot of the POA irradiance against the power production.
#
# If you have the optional dependency Holoviews installed, `scatter_hv` will return an interactive scatter plot. Additionally, `scatter_hv` includes an option to return a timeseries plot of power that is linked to the scatter plot, so points selected in the scatter plot will be highlighted in the time series.
# Uncomment the below line to use scatter_hv with linked time series
das.scatter_hv(timeseries=True)
# In this example, we have multiple measurements of the same value from different sensors. In this case a common first step is to compare measurements from the different sensors and remove data for timestamps where the measurements differ above some acceptable threshold. The `filter_sensors` method provides a convient method to accomplish this taks for the groups of measurements identified as regression values.
das.filter_sensors()
# The `get_summary` method will return a dataframe summarizing the filtering steps that have been applied, the agruments passed to them, the number of points prior to filtering, and the number of points after filtering.
das.get_summary()
# The `filter_custom` method provides a way to use your own filtering method within captest and update the summary data. The `filter_custom` method allows passing any function or method that takes a DataFrame as the first argument and returns a filtered dataframe with rows removed. Passed methods can be user-defined functions or Pandas DataFrame methods.
#
# Below, we use the `filter_custom` method with the pandas DataFrame `dropna` method to removing missing data and update the summary data.
das.filter_custom(pd.DataFrame.dropna)
# The `filter_irr` method provides a convient way to remove remove data based on the irradiance measurments. Here we use it to remove periods of low irradiance. Values greater than 2000 W/m<sup>2</sup> will also be removed, if present.
das.get_summary()
das.filter_irr(200, 2000)
# We can re-run the `scatter` method to see the results of the filtering steps.
das.scatter_hv()
# The `filter_outliers` method uses scikit-learn's elliptic envelope to remove outlier points. A future release will include a way to interactively select points to be removed.
das.filter_outliers()
das.scatter_hv()
# The `fit_regression` method performs a regression on the data stored in `data_filtered` using the regression equation specified by the standard. The regression equation is stored in the `regression_formula` attribute as shown below. Regressions are performed using the statsmodels package.
#
# Below, we set the filter argument of the `fit_regression` method to `True` to remove time periods when the residual exceeds two standard deviations of the mean residual.
das.regression_formula
das.fit_regression(filter=True, summary=False)
das.get_summary()
# ____
# #### Calculation of Reporting Conditions
#
# The `rep_cond` method provide a variety of ways to calculate reporting conditions. Using `rep_cond` the reporting conditions are always calculated from the data store in the df_flt attribute. Refer to the example notebook "Reporting Conditions Examples" for a thourough explanation of the `rep_cond` functionality. By default the reporting conditions are calcualted following the guidance of ASTM E2939-13.
das.rep_cond()
# ----
# Previously we used the irradiance filter to filter out data below 200 W/m<sup>2</sup>. The irradiance filter can also be used to filter irradiance based on a percentage band around a reference value. This approach is shown here to remove data where the irradiance is outside of +/- 50% of the reporting irradiance.
das.filter_irr(0.5, 1.5, ref_val=das.rc['poa'][0])
das.scatter_hv()
# The `fit_regression` method is used again with the default arguments, which result in fitting the regression, printing and storing the results, but not filtering. The result of the regression is a statsmodels `RegressionResultsWrapper` object containing the regression coefficients and other information generated when performing the regression. This object is stored in the CapData `regression_results` attribute.
das.fit_regression()
# The regression coefficients and p-values for each term are attributes available in the `regression_results`.
das.regression_results.params
das.regression_results.pvalues
# ## Load and Filter PVsyst Data
#
# To load and filter the modeled data, often from PVsyst, we simply create a new CapData object, load the PVsyst data, and apply the filtering methods as appropriate.
sim = pvc.CapData('sim')
# To load pvsyst data we use the `load_data` method with the `load_pvsyst` option set to True. By default the `load_data` method will search for a csv file that includes `pvsyst` in the filename in a `data` directory in the same directory as this file. If you have saved the pvsyst file in a different location, you can use the `path` and `fname` arguments to load it.
sim.load_data(load_pvsyst=True)
sim.column_groups
sim.set_regression_cols(power='real_pwr--', poa='irr-poa-', t_amb='temp-amb-', w_vel='wind--')
# +
# sim.plot()
# -
# Write over cptest.flt_sim dataframe with a copy of the original unfiltered dataframe
sim.reset_filter()
# As a first step we use the `filter_time` method to select a 60 day period of data centered around the measured data.
sim.filter_time(test_date='10/11/1990', days=60)
sim.scatter_hv()
sim.filter_irr(200, 930)
sim.scatter_hv()
sim.get_summary()
# The `filter_pvsyt` method removes data for times when shade is present or the `IL Pmin`, `IL Vmin`, `IL Pmax`, `IL Vmax` output values are greater than 0.
sim.filter_pvsyst()
sim.filter_irr(0.5, 1.5, ref_val=das.rc['poa'][0])
sim.fit_regression()
# ## Results
#
# The `get_summary` and `captest_results_check_pvalues` functions display the results of filtering on simulated and measured data and the final capacity test results comparing measured capacity to expected capacity, respectively.
pvc.get_summary(das, sim)
pvc.captest_results_check_pvalues(sim, das, 6000, '+/- 7', print_res=True)
# Uncomment and run the below lines to produce a scatter plot overlaying the final measured and PVsyst data.
# %%opts Scatter (alpha=0.3)
# %%opts Scatter [width=600]
das.scatter_hv().relabel('Measured') * sim.scatter_hv().relabel('PVsyst')
| docs/examples/complete_capacity_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
##====================================================
## このセルを最初に実行せよ---Run this cell initially.
##====================================================
import sys
if 'google.colab' in sys.modules:
# !wget -P . https://raw.githubusercontent.com/UTokyo-IPP/utpython/master/appendix/argsprint.py
# !wget -P . https://raw.githubusercontent.com/UTokyo-IPP/utpython/master/appendix/B1S.xml
# !wget -P . https://raw.githubusercontent.com/UTokyo-IPP/utpython/master/appendix/cos_sim.py
# !wget -P . https://raw.githubusercontent.com/UTokyo-IPP/utpython/master/appendix/sample.py
# !wget -P . https://raw.githubusercontent.com/UTokyo-IPP/utpython/master/appendix/small.csv
# !wget -P . https://raw.githubusercontent.com/UTokyo-IPP/utpython/master/appendix/text-sample.txt
# !wget -P . https://raw.githubusercontent.com/UTokyo-IPP/utpython/master/appendix/tokyo-july-temps.csv
# !wget -P . https://raw.githubusercontent.com/UTokyo-IPP/utpython/master/appendix/tokyo-temps.csv
# # ▲再帰
# **再帰**について説明します。
#
# 関数の**再帰呼び出し**とは、定義しようとしている関数を、その定義の中で呼び出すことです。
# 定義の中で直接呼び出す場合に限らず、他の関数を経由して間接的に呼び出す場合も、再帰呼び出しに含まれます。
# 再帰呼び出しを行う関数を、**再帰関数**といいます。
#
# 再帰関数は、**分割統治**アルゴリズムの記述に適しています。
# 分割統治とは、問題を容易に解ける小さな粒度まで分割していき、
# 個々の小さな問題を解いて、その部分解を合成することで問題全体を解くような方法を指します。
# 分割統治の考え方は、関数型プログラミングにおいてもよく用いられます。
# 再帰関数による分割統治の典型的な形は、次の通りです。
#
# ```Python
# def recursive_function(...):
# if 問題粒度の判定:
# 再帰呼び出しを含まない基本処理
# else:
# 再帰呼び出しを含む処理(問題の分割や部分解の合成を行う)
# ```
# 以下で、再帰関数を使った処理の例をいくつか見ていきましょう。
# ## 再帰関数の例:接頭辞リストと接尾辞リスト
# +
# 入力の文字列の接頭辞リストを返す関数prefixes
def prefixes(s):
if s == '':
return []
else:
return [s] + prefixes(s[:-1])
prefixes('aabcc')
# +
# 入力の文字列の接尾辞リストを返す関数suffixes
def suffixes(s):
if s == '':
return []
else:
return [s] + suffixes(s[1:])
suffixes('aabcc')
# -
# ## 再帰関数の例:べき乗の計算
# +
# 入力の底baseと冪指数exptからべき乗を計算する関数power
def power(base, expt):
if expt == 0:
# exptが0ならば1を返す
return 1
else:
# exptを1つずつ減らしながらpowerに渡し、再帰的にべき乗を計算
# (2*(2*(2*....*1)))
return base * power(base, expt - 1)
power(2,10)
# -
# 一般に、再帰処理は、繰り返し処理としても書くことができます。
# +
# べき乗の計算を繰り返し処理で行った例
def power(base, expt):
e = 1
for i in range(expt):
e *= base
return e
power(2,10)
# -
# 単純な処理においては、繰り返しの方が効率的に計算できることが多いですが、
# 特に複雑な処理になってくると、再帰的に定義した方が読みやすいコードで効率的なアルゴリズムを記述できることもあります。
# たとえば、次に示すべき乗計算は、上記よりも高速なアルゴリズムですが、計算の見通しは明快です。
# +
# べき乗を計算する高速なアルゴリズム
def power(base, expt):
if expt == 0:
return 1
elif expt % 2 == 0:
return power(base * base, expt // 2) # x**(2m) == (x*x)**m
else:
return base * power(base, expt - 1)
power(2,10)
# -
# ## 再帰関数の例:マージソート
#
# マージソートは、典型的な分割統治アルゴリズムで、以下のように再帰関数で実装することができます。
# +
# マージソートを行い、比較回数 n を返す
def merge_sort_rec(data, l, r, work):
n = 0
if r - l <= 1:
return n
m = l + (r - l) // 2
n1 = merge_sort_rec(data, l, m, work)
n2 = merge_sort_rec(data, m, r, work)
i1 = l
i2 = m
for i in range(l, r):
from1 = False
if i2 >= r:
from1 = True
elif i1 < m:
n = n + 1
if data[i1] <= data[i2]:
from1 = True
if from1:
work[i] = data[i1]
i1 = i1 + 1
else:
work[i] = data[i2]
i2 = i2 + 1
for i in range(l, r):
data[i] = work[i]
return n1 + n2 + n
def merge_sort(data):
return merge_sort_rec(data, 0, len(data), [0]*len(data))
# -
# `merge_sort` は、与えられた配列をインプレースでソートするとともに、比較の回数を返します。
# `merge_sort` は、再帰関数 `merge_sort_rec` を呼び出します。
#
# `merge_sort_rec(data, l, r, work)` は、配列 `data` のインデックスが `l` 以上で `r` より小さいところをソートします。
#
# - 要素が1つかないときは何もしません。
# - そうでなければ、`l` から `r` までの要素を半分にしてそれぞれを再帰的にソートします。
# - その結果を作業用の配列 `work` に順序を保ちながらコピーします。この操作はマージ(併合)と呼ばれます。
# - 最後に、`work` から `data` に要素を戻します。
#
# `merge_sort_rec` は自分自身を2回呼び出していますので、繰り返しでは容易には実装できません。
import random
a = [random.randint(1,10000) for i in range(100)]
merge_sort(a)
a
| appendix/3-recursion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np;
import math;
import csv;
class Simplex:
initialMatrix = [];
initialDirectives = [];
solutionMatrix = [];
def __init__(self, filename):
print("Init. Filename = " + filename);
self.initialDirectives, self.initialMatrix = Simplex.readFile(filename);
print(self.initialDirectives);
Simplex.printArray(self.initialMatrix);
def solve(self):
print("Solving using simplex...");
t_matrix, t_directives = self.checkObjective();
t_matrix, t_directives = self.correctSigns(t_matrix, t_directives);
self.solutionMatrix = self.correctSurplus( t_matrix, t_directives );
self.solutionMatrix = self.maximize(self.solutionMatrix);
print("="*25 + "Final Result" + "="*25 + "\n");
Simplex.printArray(self.solutionMatrix);
print("="*62);
def showResultValues(self):
solutionArray = [];
if(self.initialDirectives[-1] == "max"):
col = 0;
while (col < len(self.solutionMatrix[0])-1):
isNotZero, row = self.hasOne(self.solutionMatrix[:, col]);
if(isNotZero):
solutionArray.append(self.solutionMatrix[row][col] * self.solutionMatrix[row][-1]);
else:
solutionArray.append(0);
col += 1;
elif(self.initialDirectives[-1] == "min"):
i = 0;
for el in self.solutionMatrix[-1, :-1]:
i+= 1;
if(i < len(self.initialMatrix)):
solutionArray.append(el);
else:
solutionArray.insert(i - len(self.initialMatrix),el);
else:
print("Objective was not defined!");
return;
i = 0;
print("Solution is:");
for el in solutionArray:
i += 1;
if(i < len(self.initialMatrix[0])):
print("x" + str(i) + ": " + str(el));
else:
print("s" + str(i - ( len(self.initialMatrix[0]) -1 ) ) + ": " + str(el) );
def hasOne(self, array):
index = -1;
count = 0;
i = 0;
for el in array:
if(el != 0):
count += 1;
index = i;
if(count > 1):
return False, index;
i += 1;
return True, index;
def checkObjective(self):
matrix = np.copy(self.initialMatrix);
directives = np.copy(self.initialDirectives);
if directives[-1] == "min":
for i in range(matrix.shape[0]):
if directives[i] == "leq":
matrix[i] = np.multiply(matrix[i], -1.);
temp = [];
matrix = np.transpose(matrix);
for i in range(1,matrix.shape[0]):
temp.append("leq");
temp.append("max");
directives = temp;
return matrix, directives;
def correctSigns(self, matrix, directives):
matrix[-1] = np.multiply(matrix[-1], -1.);
for i in range(matrix.shape[0]-1):
if matrix[i, -1] < 0.:
if directives[i] == "geq":
directives[i] = "leq";
elif directives[i] == "leq":
directives[i] = "geq";
matrix[i] = np.multiply(matrix, -1.);
return matrix, directives;
def correctSurplus(self, matrix, directives):
s = np.zeros((matrix.shape[0], matrix.shape[0]-1), dtype=np.float32);
for i in range(matrix.shape[0]-1):
if directives[i] == "leq":
s[i, i] = 1;
else:
s[i,i] = -1;
matrix = np.insert(matrix, [matrix.shape[1]-1], s, axis=1);
return matrix;
@staticmethod
def divideArrays(array1, array2): #divides array1 by array2. If value in array2 <= 0 -> result = infinity
temp = np.zeros(len(array1));
i = 0;
for x in array2:
if( x <= 0 ):
temp[i] = math.inf;
else:
temp[i] = array1[i]/array2[i];
i += 1;
return temp;
def maximize(self, matrix):
while(min(matrix[-1]) < 0):
print("Starting a new iteration with the following matrix: ");
Simplex.printArray(matrix);
entryColumn = np.argmin(matrix[-1]);
temp = Simplex.divideArrays(matrix[0:-1, -1] , matrix[0:-1, entryColumn]);
entryRow = np.argmin(temp);
print("Pivot point is at: [" + str(entryRow) + ";" + str(entryColumn) + "]");
matrix = Simplex.performElimiation(matrix, entryRow, entryColumn);
print();
return matrix;
@staticmethod
def printArray(matrix):
for row in matrix:
line = " [";
for el in row:
line = line + "%7.1f " % (el);
line = line + "]";
print(line);
print();
@staticmethod
def readFile(filepath):
with open(filepath) as file:
reader = csv.reader(file, delimiter=",");
matrix = [];
obj_fun = [];
objective = None;
directives = [];
for row in reader:
line = [];
for el in row[1:]:
line.append(float(el));
if(row[0] == "min" or row[0] == "max"):
obj_fun = line;
objective = row[0];
else:
directives.append(row[0]);
matrix.append(line);
matrix.append(obj_fun);
directives.append(objective);
matrix = np.asarray(matrix);
return directives, matrix;
@staticmethod
def performElimiation(matrix, entryRow, entryColumn):
matrix[entryRow] = matrix[entryRow]/matrix[entryRow][entryColumn];
i = 0;
while i < len(matrix):
if(i != entryRow):
matrix[i] = matrix[i] - matrix[i][entryColumn]*matrix[entryRow]
i += 1;
return matrix;
v15min = Simplex("15min.txt");
v15max = Simplex("15max.txt");
v15min.solve();
v15min.showResultValues();
v15max.solve();
v15max.showResultValues();
mixMax = Simplex("mixMax.csv");
mixMax.solve();
mixMax.showResultValues();
| Lab3/.ipynb_checkpoints/Optimization_v5-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problem 001
#
# If we list all the natural numbers below 10 that are multiples of 3 or 5, we get 3, 5, 6 and 9. The sum of these multiples is 23.
#
# Find the sum of all the multiples of 3 or 5 below 1000.
# # Solution
# Note that a much easier to compute solution can be found using the fact that
#
# $$
# 1 + 2 + \dots + n = \frac{n(n+1)}{2}
# $$
#
# along with the inclusion-exclusion principle. However, this is quick enough to compute by brute-force.
sum([n for n in range(1000 + 1) if n % 3 == 0 or n % 5 == 0])
| python/001.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from xml.dom import minidom
import numpy as np
import geopy.distance
import pandas as pd
import xml.etree.ElementTree as ET
import uuid
import matplotlib.pyplot as plt
COLUMNS = ["type", "from", "to", "start_tick", "duration", "slowdown"]
TRIPS_COMPLETE_PATH = "trips_pesquisa_od.xml"
tree = ET.parse(TRIPS_COMPLETE_PATH)
trips = tree.getroot()
# -
selected_trips = []
edges = {}
df = pd.read_csv('../scenario_2_sao_paulo/inputs/events.csv', sep=';', names=COLUMNS)
forbidden_nodes = []
forbidden_nodes.append(df['from'].unique())
forbidden_nodes.append(df['to'].unique())
forbidden_nodes = np.array(forbidden_nodes).flatten()
# +
def load_nodes(path):
nodes = []
events_xml = minidom.parse(path)
map_nodes_tags = events_xml.getElementsByTagName('node')
for n in map_nodes_tags:
nodes.append(int(n.getAttribute('id')))
return nodes
def load_edges(path):
map_xml = minidom.parse(path)
map_links_tags = map_xml.getElementsByTagName('link')
for n in map_links_tags:
edges[int(n.getAttribute('id'))] = [
int(n.getAttribute('from')),
int(n.getAttribute('to'))
]
# -
nodes = load_nodes('../scenario_2_sao_paulo/inputs/map.xml')
load_edges('../scenario_2_sao_paulo/inputs/map.xml')
def find_start_link(origin, destination):
for k, e in edges.items():
if (e[0] == origin):
return k
return -1
for trip in trips.iter('trip'):
origin = int(trip.attrib['origin'])
destination = int(trip.attrib['destination'])
if (origin in nodes and destination in nodes):
selected_trips.append(trip)
df = pd.DataFrame(columns=["name", "origin", "destination", "link_origin", "count", "start", "mode", "uuid"])
counter = 0
for s in selected_trips:
origin = int(s.attrib['origin'])
destination = int(s.attrib['destination'])
link_origin = int(find_start_link(origin, destination))
if (link_origin == -1):
continue
if ((origin in forbidden_nodes) or (destination in forbidden_nodes)):
continue
skip_nodes = [2051762297, 3316840685, 303561733, 2006965204, 2465920967, 2397982708, 2397319383, 5629463536, 4483150089]
if ((origin in skip_nodes) or (destination in skip_nodes)):
continue
name = s.attrib['name']
old_count = int(s.attrib['count'])
start = int(s.attrib['start'])
mode = s.attrib['mode']
uuid_trip = 'a'
df.loc[counter] = [name, origin, destination, link_origin, old_count, start, mode, uuid_trip]
counter += 1
df = df[(df.start >= 28800) & (df.start <= 30000)]
df.start = df.start - 28800
final_df = pd.DataFrame(columns=["name", "origin", "destination", "link_origin", "count", "start", "mode", "uuid"])
df.describe()
counter = 0
for row in df.iterrows():
access = row[1]
repeat = access['count']
for v in range(1, repeat):
final_df.loc[counter] = [
access['name'],
access['origin'],
access['destination'],
access['link_origin'],
1,
access['start'],
access['mode'],
uuid.uuid4()
]
counter += 1
final_df.to_csv('../scenario_2_sao_paulo/inputs/trips-with-uuid.csv', header=False, sep=';', index=False)
final_df.groupby(['origin', 'destination']).count()
final_df.destination.unique()
| utils/filter_trips.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="-zMKQx6DkKwt"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" id="J307vsiDkMMW"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="vCMYwDIE9dTT"
# # Keras functional API в TensorFlow
# + [markdown] id="lAJfkZ-K9flj"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/guide/keras/functional"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />Смотрите на TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ru/guide/keras/functional.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Запустите в Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ru/guide/keras/functional.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />Изучайте код на GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ru/guide/keras/functional.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Скачайте ноутбук</a>
# </td>
# </table>
# + [markdown] id="fj66ZXAzrJC2"
# Note: Вся информация в этом разделе переведена с помощью русскоговорящего Tensorflow сообщества на общественных началах. Поскольку этот перевод не является официальным, мы не гарантируем что он на 100% аккуратен и соответствует [официальной документации на английском языке](https://www.tensorflow.org/?hl=en). Если у вас есть предложение как исправить этот перевод, мы будем очень рады увидеть pull request в [tensorflow/docs](https://github.com/tensorflow/docs) репозиторий GitHub. Если вы хотите помочь сделать документацию по Tensorflow лучше (сделать сам перевод или проверить перевод подготовленный кем-то другим), напишите нам на [<EMAIL> list](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-ru).
# + [markdown] id="ITh3wzORxgpw"
# ## Setup
# + id="HFbM9dcfxh4l"
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# # %tensorflow_version существует только Colab.
# %tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
tf.keras.backend.clear_session() # Для простого сброса состояния ноутбука.
# + [markdown] id="ZI47-lpfkZ5c"
# ## Введение
#
# Вы уже знакомы с использованием `keras.Sequential()` для создания моделей.
# Functional API позволяет создавать модели более гибко чем `Sequential`:
# он может обрабатывать модели с нелинейной топологией, модели с общими слоями,
# и модели с несколькими входами или выходами.
#
# Функциональный подход основан на том, что модель глубокого обучения
# обычно представляет собой ориентированный ациклический граф (DAG) слоев.
# Functional API - это набор инструментов для **построения графа слоев**.
#
# Рассмотрим следующую модель:
#
# ```
# (вход: 784-мерный вектор)
# ↧
# [Плотный слой (64 элемента, активация relu)]
# ↧
# [Плотный слой (64 элемента, активация relu)]
# ↧
# [Плотный слой (10 элементов, активация softmax)]
# ↧
# (выход: вероятностное распределение на 10 классов)
# ```
#
# Это простой граф из 3 слоев.
#
# Для построения этой модели с помощью Functional API,
# вам надо начать с создания входного узла:
# + id="Yxi0LaSHkDT-"
from tensorflow import keras
inputs = keras.Input(shape=(784,))
# + [markdown] id="Mr3Z_Pxcnf-H"
# Здесь мы просто указываем размерность наших данных: 784-мерных векторов.
# Обратите внимание, что количество данных всегда опускается, мы указываем только размерность каждого элемента.
# Для ввода предназначенного для изображений размеров `(32, 32, 3)`, мы бы использовали:
# + id="0-2Q2nJNneIO"
img_inputs = keras.Input(shape=(32, 32, 3))
# + [markdown] id="HoMFNu-pnkgF"
# То, что возвращает `inputs`, содержит информацию о размерах и типе данных
# которые вы планируете передать в вашу модель:
# + id="ddIr9LPJnibj"
inputs.shape
# + id="lZkLJeQonmTe"
inputs.dtype
# + [markdown] id="kZnhhndTnrzC"
# Вы создаете новый узел в графе слоев, вызывая слой на этом объекте `inputs`:
# + id="sMyyMTqDnpYV"
from tensorflow.keras import layers
dense = layers.Dense(64, activation='relu')
x = dense(inputs)
# + [markdown] id="besm-lgFnveV"
# "Вызов слоя" аналогичен рисованию стрелки из "входных данных" в созданный нами слой.
# Мы "передаем" входные данные в `dense` слой, и мы получаем `x`.
#
# Давайте добавим еще несколько слоев в наш граф слоев:
# + id="DbF-MIO2ntf7"
x = layers.Dense(64, activation='relu')(x)
outputs = layers.Dense(10, activation='softmax')(x)
# + [markdown] id="B38UlEIlnz_8"
# Сейчас мы можем создать `Model` указав его входы и выходы в графе слоев:
# + id="MrSfwvl-nx9s"
model = keras.Model(inputs=inputs, outputs=outputs)
# + [markdown] id="5EeeV1xJn3jW"
# Напомним полный процесс определения модели:
# + id="xkz7oqj2n1-q"
inputs = keras.Input(shape=(784,), name='img')
x = layers.Dense(64, activation='relu')(inputs)
x = layers.Dense(64, activation='relu')(x)
outputs = layers.Dense(10, activation='softmax')(x)
model = keras.Model(inputs=inputs, outputs=outputs, name='mnist_model')
# + [markdown] id="jJzocCbdn6qj"
# Давайте посмотрим как выглядит сводка модели:
# + id="GirC9odQn5Ep"
model.summary()
# + [markdown] id="mbNqYAlOn-vA"
# Мы также можем начертить модель в виде графа:
# + id="JYh2wLain8Oi"
keras.utils.plot_model(model, 'my_first_model.png')
# + [markdown] id="QtgX2RoGoDZo"
# И опционально выведем размерности входа и выхода каждого слоя на построенном графе:
# + id="7FGesSSuoAG5"
keras.utils.plot_model(model, 'my_first_model_with_shape_info.png', show_shapes=True)
# + [markdown] id="PBZ9XE6LoWvi"
# Это изображение и код который мы написали идентичны. В версии кода,
# связывающие стрелки просто заменены операциями вызова.
#
# "Граф слоев" это очень интуитивный ментальный образ для модели глубокого обучения,
# а functional API это способ создания моделей которые близко отражают этот ментальный образ.
# + [markdown] id="WUUHMaKLoZDn"
# ## Обучение, оценка и вывод
#
# Обучение, оценка и вывод работают для моделей построенных
# с использованием Functional API точно так же как и в Sequential моделях.
#
# Вот быстрая демонстрация.
#
# Тут мы загружаем датасет изображений MNIST, преобразуем его в векторы,
# обучаем модель на данных (мониторя при этом качество работы на проверочной выборке),
# и наконец мы оцениваем нашу модель на тестовых данных:
# + id="DnHvkD22oFEY"
(x_train, y_train), (x_test, y_test) = keras.datasets.mnist.load_data()
x_train = x_train.reshape(60000, 784).astype('float32') / 255
x_test = x_test.reshape(10000, 784).astype('float32') / 255
model.compile(loss='sparse_categorical_crossentropy',
optimizer=keras.optimizers.RMSprop(),
metrics=['accuracy'])
history = model.fit(x_train, y_train,
batch_size=64,
epochs=5,
validation_split=0.2)
test_scores = model.evaluate(x_test, y_test, verbose=2)
print('Test loss:', test_scores[0])
print('Test accuracy:', test_scores[1])
# + [markdown] id="c3nq2fjiLCkE"
# Полное руководство посвященное обучению и оценки моделей, см. по ссылке [руководство обучения и оценки](./train_and_evaluate.ipynb).
# + [markdown] id="XOsL56zDorLh"
# ## Сохранение и сериализация
#
# Сохранение и сериализация для моделей построенных с использованием
# Functional API работает точно так же как и для Sequential моделей.
#
# Стандартным способом сохранения Functional модели является вызов `model.save()` позволяющий сохранить всю модель в один файл.
# Позже вы можете восстановить ту же модель из этого файла, даже если у вас больше нет доступа к коду
# создавшему модель.
#
# Этот файл включает:
# - Архитектуру модели
# - Значения весов модели (которые были получены во время обучения)
# - Конфигурация обучения модели (то что вы передавали в `compile`)
# - Оптимизатор и его состояние, если оно было (это позволяет возобновить обучение с того места, где вы остановились)
# + id="kN-AO7xvobtr"
model.save('path_to_my_model.h5')
del model
# Восстановить в точности ту же модель исключительно из файла:
model = keras.models.load_model('path_to_my_model.h5')
# + [markdown] id="u0J0tFPHK4pb"
# Полное руководство по сохранению моделей см. в [Руководство по сохранению и сериализации моделей](./save_and_serialize.ipynb).
# + [markdown] id="lKz1WWr2LUzF"
# ## Использование одного и того же графа слоев для определения нескольких моделей
#
#
# В functional API, модели создаются путем указания входных
# и выходных данных в графе слоев. Это значит что один граф слоев
# может быть использован для генерации нескольких моделей.
#
# В приведенном ниже примере мы используем один и тот же стек слоев для создания двух моделей:
# модель `кодировщика (encoder)` которая преобразует входные изображения в 16-мерные вектора,
# и сквозную модель `автокодировщика (autoencoder)` для обучения.
#
# + id="WItZQr6LuVbF"
encoder_input = keras.Input(shape=(28, 28, 1), name='img')
x = layers.Conv2D(16, 3, activation='relu')(encoder_input)
x = layers.Conv2D(32, 3, activation='relu')(x)
x = layers.MaxPooling2D(3)(x)
x = layers.Conv2D(32, 3, activation='relu')(x)
x = layers.Conv2D(16, 3, activation='relu')(x)
encoder_output = layers.GlobalMaxPooling2D()(x)
encoder = keras.Model(encoder_input, encoder_output, name='encoder')
encoder.summary()
x = layers.Reshape((4, 4, 1))(encoder_output)
x = layers.Conv2DTranspose(16, 3, activation='relu')(x)
x = layers.Conv2DTranspose(32, 3, activation='relu')(x)
x = layers.UpSampling2D(3)(x)
x = layers.Conv2DTranspose(16, 3, activation='relu')(x)
decoder_output = layers.Conv2DTranspose(1, 3, activation='relu')(x)
autoencoder = keras.Model(encoder_input, decoder_output, name='autoencoder')
autoencoder.summary()
# + [markdown] id="oNeg3WWFuYZK"
# Обратите внимание, что мы делаем архитектуру декодирования строго симметричной архитектуре кодирования,
# таким образом мы получим размерность выходных данных такую же как и входных данных `(28, 28, 1)`.
# Обратным к слою `Conv2D` является слой `Conv2DTranspose`, а обратным к слою `MaxPooling2D`
# будет слой `UpSampling2D`.
# + [markdown] id="h1FVW4j-uc6Y"
# ## Модели можно вызывать как слои
#
# Вы можете использовать любую модель так, как если бы это был слой передавая ее в `Input` или на выход другого слоя.
# Обратите внимание, что вызывая модель, вы не только переиспользуете ее архитектуру, вы также повторно используете ее веса.
#
# Давайте посмотрим на это в действии. Вот другой взгляд на пример автокодировщика, когда создается модель кодировщика, модель декодировщика, и они связываются в два вызова для получения модели автокодировщика:
# + id="Ld7KdsQ_uZbr"
encoder_input = keras.Input(shape=(28, 28, 1), name='original_img')
x = layers.Conv2D(16, 3, activation='relu')(encoder_input)
x = layers.Conv2D(32, 3, activation='relu')(x)
x = layers.MaxPooling2D(3)(x)
x = layers.Conv2D(32, 3, activation='relu')(x)
x = layers.Conv2D(16, 3, activation='relu')(x)
encoder_output = layers.GlobalMaxPooling2D()(x)
encoder = keras.Model(encoder_input, encoder_output, name='encoder')
encoder.summary()
decoder_input = keras.Input(shape=(16,), name='encoded_img')
x = layers.Reshape((4, 4, 1))(decoder_input)
x = layers.Conv2DTranspose(16, 3, activation='relu')(x)
x = layers.Conv2DTranspose(32, 3, activation='relu')(x)
x = layers.UpSampling2D(3)(x)
x = layers.Conv2DTranspose(16, 3, activation='relu')(x)
decoder_output = layers.Conv2DTranspose(1, 3, activation='relu')(x)
decoder = keras.Model(decoder_input, decoder_output, name='decoder')
decoder.summary()
autoencoder_input = keras.Input(shape=(28, 28, 1), name='img')
encoded_img = encoder(autoencoder_input)
decoded_img = decoder(encoded_img)
autoencoder = keras.Model(autoencoder_input, decoded_img, name='autoencoder')
autoencoder.summary()
# + [markdown] id="icQFny_huiXC"
# Как вы видите, модель может быть вложена: модель может содержать подмодель (поскольку модель можно рассматривать как слой).
#
# Распространенным вариантом использования вложения моделей является *ensembling*.
# Вот пример того, как можно объединить набор моделей в одну модель которая усредняет их прогнозы:
# + id="ZBlZbRn5uk-9"
def get_model():
inputs = keras.Input(shape=(128,))
outputs = layers.Dense(1, activation='sigmoid')(inputs)
return keras.Model(inputs, outputs)
model1 = get_model()
model2 = get_model()
model3 = get_model()
inputs = keras.Input(shape=(128,))
y1 = model1(inputs)
y2 = model2(inputs)
y3 = model3(inputs)
outputs = layers.average([y1, y2, y3])
ensemble_model = keras.Model(inputs=inputs, outputs=outputs)
# + [markdown] id="e1za1TZxuoId"
# ## Манипулирование сложными топологиями графов
#
#
# ### Модели с несколькими входами и выходами
#
# Functional API упрощает манипуляции с несколькими входами и выходами.
# Это не может быть сделано с использованием Sequential API.
#
# Вот простой пример.
#
# Допустим, вы создаете систему для ранжирования клиентских заявок по приоритетам и направления их в нужный отдел.
#
# У вашей модели будет 3 входа:
#
# - Заголовок заявки (текстовые входные данные)
# - Текстовое содержание заявки (текстовые входные данные)
# - Любые теги добавленные пользователем (категорийные входные данные)
#
# У модели будет 2 выхода:
#
# - Оценка приоритета между 0 и 1 (скаляр, результат сигмоидного выхода)
# - Отдел который должен обработать заявку (softmax выход относительно множества отделов)
#
# Давайте построим модель в несколько строк с помощью Functional API.
# + id="Gt91OtzbutJy"
num_tags = 12 # Количество различных тегов проблем
num_words = 10000 # Размер словаря полученный в результате предобработки текстовых данных
num_departments = 4 # Количество отделов для предсказаний
title_input = keras.Input(shape=(None,), name='title') # Последовательность целых чисел переменной длины
body_input = keras.Input(shape=(None,), name='body') # Последовательность целых чисел переменной длины
tags_input = keras.Input(shape=(num_tags,), name='tags') # Бинарный вектор размера `num_tags`
# Вложим каждое слово заголовка в 64-мерный вектор
title_features = layers.Embedding(num_words, 64)(title_input)
# Вложим каждое слово текста в 64-мерный вектор
body_features = layers.Embedding(num_words, 64)(body_input)
# Сокращаем последовательность вложенных слов заголовка до одного 128-мерного вектора
title_features = layers.LSTM(128)(title_features)
# Сокращаем последовательность вложенных слов содержимого до одного 32-мерного вектора
body_features = layers.LSTM(32)(body_features)
# Объединим все признаки в один вектор с помощью конкатенации
x = layers.concatenate([title_features, body_features, tags_input])
# Добавим логистическую регрессию для прогнозирования приоритета по признакам
priority_pred = layers.Dense(1, activation='sigmoid', name='priority')(x)
# Добавим классификатор отделов прогнозирующий на признаках
department_pred = layers.Dense(num_departments, activation='softmax', name='department')(x)
# Создание сквозной модели, прогнозирующей приоритет и отдел
model = keras.Model(inputs=[title_input, body_input, tags_input],
outputs=[priority_pred, department_pred])
# + [markdown] id="KIS7lqW0uwh-"
# Давайте начертим граф модели:
# + id="IMij4gzhuzYV"
keras.utils.plot_model(model, 'multi_input_and_output_model.png', show_shapes=True)
# + [markdown] id="oOyuig2Hu00p"
# При компиляции этой модели, мы можем присвоить различные функции потерь каждому выходу.
# Вы даже можете присвоить разные веса каждой функции потерь, чтобы варьировать их
# вклад в общую функцию потерь обучения.
# + id="Crtdpi5Uu2cX"
model.compile(optimizer=keras.optimizers.RMSprop(1e-3),
loss=['binary_crossentropy', 'categorical_crossentropy'],
loss_weights=[1., 0.2])
# + [markdown] id="t42Jrn0Yu5jL"
# Так как мы дали имена нашим выходным слоям, мы можем также указать именованные функции потерь:
# + id="dPM0EwW_u6mV"
model.compile(optimizer=keras.optimizers.RMSprop(1e-3),
loss={'priority': 'binary_crossentropy',
'department': 'categorical_crossentropy'},
loss_weights=[1., 0.2])
# + [markdown] id="bpTx2sXnu3-W"
# Мы можем обучить модель передавая входные данные(списки массивов Numpy) и метки:
# + id="nB-upOoGu_k4"
import numpy as np
# Учебные входные данные
title_data = np.random.randint(num_words, size=(1280, 10))
body_data = np.random.randint(num_words, size=(1280, 100))
tags_data = np.random.randint(2, size=(1280, num_tags)).astype('float32')
# Учебные целевые данные
priority_targets = np.random.random(size=(1280, 1))
dept_targets = np.random.randint(2, size=(1280, num_departments))
history = model.fit({'title': title_data, 'body': body_data, 'tags': tags_data},
{'priority': priority_targets, 'department': dept_targets},
epochs=2,
batch_size=32)
# + [markdown] id="qNguhBWuvCtz"
# При вызове fit с объектом `Dataset`, должны возвращаться либо
# кортеж списков, таких как `([title_data, body_data, tags_data], [priority_targets, dept_targets])`
# либо кортеж словарей
# `({'title': title_data, 'body': body_data, 'tags': tags_data}, {'priority': priority_targets, 'department': dept_targets})`.
#
# Для более подробного объяснения обратитесь к полному руководству [руководство по обучению и оценке](./train_and_evaluate.ipynb).
# + [markdown] id="tR0X5tTOvPyg"
# ### Учебная ResNet модель
#
# В дополнение к моделям с несколькими входами и выходами,
# Functional API упрощает манипулирование топологиями с нелинейной связностью,
# то есть моделями, в которых слои не связаны последовательно.
# Это также не может быть реализовано с помощью Sequential API (это видно из названия).
#
# Распространенный пример использования этого - residual connections.
#
# Давайте построим учебную ResNet модель для CIFAR10 чтобы продемонстрировать это.
# + id="VzMoYrMNvXrm"
inputs = keras.Input(shape=(32, 32, 3), name='img')
x = layers.Conv2D(32, 3, activation='relu')(inputs)
x = layers.Conv2D(64, 3, activation='relu')(x)
block_1_output = layers.MaxPooling2D(3)(x)
x = layers.Conv2D(64, 3, activation='relu', padding='same')(block_1_output)
x = layers.Conv2D(64, 3, activation='relu', padding='same')(x)
block_2_output = layers.add([x, block_1_output])
x = layers.Conv2D(64, 3, activation='relu', padding='same')(block_2_output)
x = layers.Conv2D(64, 3, activation='relu', padding='same')(x)
block_3_output = layers.add([x, block_2_output])
x = layers.Conv2D(64, 3, activation='relu')(block_3_output)
x = layers.GlobalAveragePooling2D()(x)
x = layers.Dense(256, activation='relu')(x)
x = layers.Dropout(0.5)(x)
outputs = layers.Dense(10, activation='softmax')(x)
model = keras.Model(inputs, outputs, name='toy_resnet')
model.summary()
# + [markdown] id="ISQX32bgrkis"
# Давайте начертим модель:
# + id="pNFVkAd3rlCM"
keras.utils.plot_model(model, 'mini_resnet.png', show_shapes=True)
# + [markdown] id="ECcG87yZrxp5"
# Давайте обучим ее:
# + id="_iXGz5XEryou"
(x_train, y_train), (x_test, y_test) = keras.datasets.cifar10.load_data()
x_train = x_train.astype('float32') / 255.
x_test = x_test.astype('float32') / 255.
y_train = keras.utils.to_categorical(y_train, 10)
y_test = keras.utils.to_categorical(y_test, 10)
model.compile(optimizer=keras.optimizers.RMSprop(1e-3),
loss='categorical_crossentropy',
metrics=['acc'])
model.fit(x_train, y_train,
batch_size=64,
epochs=1,
validation_split=0.2)
# + [markdown] id="XQfg0JUkr7SH"
# ## Совместное использование слоев
#
# Другим хорошим использованием functional API являются модели, использующие общие слои. Общие слои - это экземпляры слоев, которые переиспользуются в одной и той же модели: они изучают признаки, которые относятся к нескольким путям в графе слоев.
#
# Общие слои часто используются для кодирования входных данных, которые поступают из одинаковых пространств (скажем, из двух разных фрагментов текста, имеющих одинаковый словарь), поскольку они обеспечивают обмен информацией между этими различными данными, что позволяет обучать такие модели на меньшем количестве данных. Если определенное слово появилось на одном из входов, это будет способствовать его обработке на всех входах, которые проходят через общий уровень.
#
# Чтобы совместно использовать слой в Functional API, просто вызовите тот же экземпляр слоя несколько раз. Например, здесь слой `Embedding` используется совместно на двух текстовых входах:
# + id="R9pAPQCnKuMR"
# Вложения для 1000 различных слов в 128-мерные вектора
shared_embedding = layers.Embedding(1000, 128)
# Целочисленные последовательности переменной длины
text_input_a = keras.Input(shape=(None,), dtype='int32')
# Целочисленные последовательности переменной длины
text_input_b = keras.Input(shape=(None,), dtype='int32')
# Мы переиспользуем тот же слой для кодирования на обоих входах
encoded_input_a = shared_embedding(text_input_a)
encoded_input_b = shared_embedding(text_input_b)
# + [markdown] id="xNEKvfUpr-Kf"
# ## Извлечение и повторное использование узлов в графе слоев
# + [markdown] id="JHVGI6bEr-ze"
# Поскольку граф слоев, которыми вы манипулируете в Functional API, является статической структурой данных, к ней можно получить доступ и проверить ее. Именно так мы строим Functional модели, например, в виде изображений.
#
# Это также означает, что мы можем получить доступ к активациям промежуточных слоев ("узлов" в графе) и использовать их в других местах. Это чрезвычайно полезно для извлечения признаков, например!
#
# Давайте посмотрим пример. Это модель VGG19 с весами предобученными на ImageNet:
# + id="c-gl3xHBH-oX"
from tensorflow.keras.applications import VGG19
vgg19 = VGG19()
# + [markdown] id="AKefin_xIGBP"
# И это промежуточные активации модели, полученные путем запроса к структуре данных графа:
# + id="1_Ap05fgIRgE"
features_list = [layer.output for layer in vgg19.layers]
# + [markdown] id="H1zx5qM7IYu4"
# Мы можем использовать эти признаки для создания новой модели извлечения признаков, которая возвращает значения активаций промежуточного уровня - и мы можем сделать все это в 3 строчки
# + id="NrU82Pa8Igwo"
feat_extraction_model = keras.Model(inputs=vgg19.input, outputs=features_list)
img = np.random.random((1, 224, 224, 3)).astype('float32')
extracted_features = feat_extraction_model(img)
# + [markdown] id="G-e2-jNCLIqy"
# Это удобно когда [реализуется neural style transfer](https://medium.com/tensorflow/neural-style-transfer-creating-art-with-deep-learning-using-tf-keras-and-eager-execution-7d541ac31398), как и в других случаях.
# + [markdown] id="t9M2Uvi3sBy0"
# ## Расширение API при помощи написания кастомных слоев
#
# tf.keras обладает широким набором встроенных слоев. Вот несколько примеров:
#
# - Сверточные слои: `Conv1D`, `Conv2D`, `Conv3D`, `Conv2DTranspose`, и т.д.
# - Слои пулинга: `MaxPooling1D`, `MaxPooling2D`, `MaxPooling3D`, `AveragePooling1D`, и т.д.
# - Слои RNN: `GRU`, `LSTM`, `ConvLSTM2D`, и т.д.
# - `BatchNormalization`, `Dropout`, `Embedding`, и т.д.
#
# Если вы не нашли то, что вам нужно, легко расширить API создав собственный слой.
#
# Все слои наследуют класс `Layer` и реализуют:
# - Метод `call`, определяющий вычисления выполняемые слоем.
# - Метод `build`, создающий веса слоя (заметим что это всего лишь стилевое соглашение; вы можете также создать веса в `__init__`).
#
# Чтобы узать больше о создании слоев с нуля, проверьте руководство [Руководство по написанию слоев и моделей с нуля](./custom_layers_and_models.ipynb).
#
# Вот простая реализация `Dense` слоя:
# + id="ztAmarbgNV6V"
class CustomDense(layers.Layer):
def __init__(self, units=32):
super(CustomDense, self).__init__()
self.units = units
def build(self, input_shape):
self.w = self.add_weight(shape=(input_shape[-1], self.units),
initializer='random_normal',
trainable=True)
self.b = self.add_weight(shape=(self.units,),
initializer='random_normal',
trainable=True)
def call(self, inputs):
return tf.matmul(inputs, self.w) + self.b
inputs = keras.Input((4,))
outputs = CustomDense(10)(inputs)
model = keras.Model(inputs, outputs)
# + [markdown] id="NXxp_32bNWTy"
# Если вы хотите, чтобы ваш пользовательский слой поддерживал сериализацию, вы также должны определить метод` get_config`,возвращающий аргументы конструктора экземпляра слоя:
# + id="K3OQ4XxzNfAZ"
class CustomDense(layers.Layer):
def __init__(self, units=32):
super(CustomDense, self).__init__()
self.units = units
def build(self, input_shape):
self.w = self.add_weight(shape=(input_shape[-1], self.units),
initializer='random_normal',
trainable=True)
self.b = self.add_weight(shape=(self.units,),
initializer='random_normal',
trainable=True)
def call(self, inputs):
return tf.matmul(inputs, self.w) + self.b
def get_config(self):
return {'units': self.units}
inputs = keras.Input((4,))
outputs = CustomDense(10)(inputs)
model = keras.Model(inputs, outputs)
config = model.get_config()
new_model = keras.Model.from_config(
config, custom_objects={'CustomDense': CustomDense})
# + [markdown] id="kXg6hZN_NfN8"
# Опционально, вы также можете реализовать метод класса `from_config (cls, config)`, который отвечает за пересоздание экземпляра слоя, учитывая его словарь конфигурации. Реализация по умолчанию` from_config` выглядит так:
#
# ```python
# def from_config(cls, config):
# return cls(**config)
# ```
# + [markdown] id="ifOVqn84sCNU"
# ## Когда использовать Functional API
#
# Как определить когда лучше использовать Functional API для создания новой модели, или просто наследовать класс `Model` напрямую?
#
# В целом, Functional API более высокоуровневый и простой в использовании, он имеет ряд функций, которые не поддерживаются наследуемыми `Model`.
#
# Однако, наследование `Model` дает вам большую гибкость при создании моделей, которые не описываются легко в виде направленного ациклического графа слоев (например, вы не сможете реализовать Tree-RNN с Functional API, вам нужно напрямую наследоваться от `Model`).
#
#
# ### Cильные стороны Functional API:
#
# Свойства перечисленные ниже являются верными и для Sequential моделей (которые также являются структурами данных), но не верны для наследуемых моделей (которые представляют собой код Python, а не структуры данных).
#
#
# #### С Functional API получается более короткий код.
#
# Нет `super(MyClass, self).__init__(...)`, нет `def call(self, ...):`, и т.д.
#
# Сравните:
#
# ```python
# inputs = keras.Input(shape=(32,))
# x = layers.Dense(64, activation='relu')(inputs)
# outputs = layers.Dense(10)(x)
# mlp = keras.Model(inputs, outputs)
# ```
#
# С наследуемой версией:
#
# ```python
# class MLP(keras.Model):
#
# def __init__(self, **kwargs):
# super(MLP, self).__init__(**kwargs)
# self.dense_1 = layers.Dense(64, activation='relu')
# self.dense_2 = layers.Dense(10)
#
# def call(self, inputs):
# x = self.dense_1(inputs)
# return self.dense_2(x)
#
# # Создадим экземпляр модели.
# mlp = MLP()
# # Необходимо создать состояние модели.
# # У модели нет состояния пока она не была вызвана хотя бы раз.
# _ = mlp(tf.zeros((1, 32)))
# ```
#
#
# #### Он валидирует вашу модель пока вы ее определяете.
#
# В Functional API входные спецификации (shape и dtype) создаются заранее (через `Input`), и каждый раз, когда вы вызываете слой, слой проверяет, что спецификации переданные ему соответствует его предположениям, если это не так то вы получите полезное сообщение об ошибке.
#
# Это гарантирует, что любая модель которую вы построите с Functional API запустится. Вся отладка (не относящаяся к отладке сходимости) будет происходить статично во время конструирования модели, а не во время выполнения. Это аналогично проверке типа в компиляторе.
#
#
# #### Вашу Functional модель можно представить графически, а также она проверяема.
#
# Вы можете начертить модель в виде графа, и вы легко можете получить доступ к промежуточным узлам графа, например, чтобы извлечь и переиспользовать активации промежуточных слоев, как мы видели в предыдущем примере:
#
# ```python
# features_list = [layer.output for layer in vgg19.layers]
# feat_extraction_model = keras.Model(inputs=vgg19.input, outputs=features_list)
# ```
#
#
# #### Ваша Functional модель может быть сериализована или клонирована.
#
# Поскольку Functional модель это скорее структура данных чем кусок кода, она безопасно сериализуется и может быть сохранена в виде одного файла который позволяет вам воссоздать в точности ту же модель без доступа к исходному коду. Смотрите наше [руководство по сохранению и сериализации](./save_and_serialize.ipynb) для лучшего понимания.
#
#
# ### Слабые стороны Functional API:
#
#
# #### Он не поддерживает динамичные архитектуры.
#
# Functional API обрабатывает модели как группы DAG слоев. Это справедливо для большинства архитектур глубокого обучения, но не для всех: например, рекурсивные сети или Tree RNN не соответствуют этому предположению и не могут быть реализованы в Functional API.
#
#
# #### Иногда вам просто нужно написать все с нуля.
#
# При написании продвинутых архитектур вы можете захотеть сделать то, что выходит за рамки "определения DAG слоев": например, вы можете использовать несколько пользовательских методов обучения и вывода на экземпляре вашей модели. Это требует наследования `Model`.
#
#
# ---
#
#
# Чтобы лучше понять разницу между Functional API и наследованием `Model`, советуем прочитать [Что такое Symbolic и Imperative API в TensorFlow 2.0?](https://medium.com/tensorflow/what-are-symbolic-and-imperative-apis-in-tensorflow-2-0-dfccecb01021).
# + [markdown] id="Ym1jrCqusGvj"
# ## Сочетание различных стилей API
#
# Важно отметить, что выбор между Functional API или наследованием `Model` не является бинарным решением, которое ограничивает вас одной категорией моделей. Все модели в API tf.keras могут взаимодействовать друг с другом, будь то Sequential модели, Functional модели или наследуемые Models/Layers, написанные с нуля.
#
# Вы всегда можете использовать Functional модель или Sequential модель как часть наследуемой Model/Layer:
# + id="9zF5YTLy_vGZ"
units = 32
timesteps = 10
input_dim = 5
# Define a Functional model
inputs = keras.Input((None, units))
x = layers.GlobalAveragePooling1D()(inputs)
outputs = layers.Dense(1, activation='sigmoid')(x)
model = keras.Model(inputs, outputs)
class CustomRNN(layers.Layer):
def __init__(self):
super(CustomRNN, self).__init__()
self.units = units
self.projection_1 = layers.Dense(units=units, activation='tanh')
self.projection_2 = layers.Dense(units=units, activation='tanh')
# Our previously-defined Functional model
self.classifier = model
def call(self, inputs):
outputs = []
state = tf.zeros(shape=(inputs.shape[0], self.units))
for t in range(inputs.shape[1]):
x = inputs[:, t, :]
h = self.projection_1(x)
y = h + self.projection_2(state)
state = y
outputs.append(y)
features = tf.stack(outputs, axis=1)
print(features.shape)
return self.classifier(features)
rnn_model = CustomRNN()
_ = rnn_model(tf.zeros((1, timesteps, input_dim)))
# + [markdown] id="oxW1d0a8_ufg"
# Вы можете использовать любой Layer или Model, наследующий keras.Model в Functional API в том случае если реализован метод `call` который соответствует одному из следующих паттернов:
#
# - `call(self, inputs, **kwargs)` где `inputs` это тензор или вложенная струтура тензоров (напр. список тензоров), и где `**kwargs` это нетензорные аргументы (не входные данные).
# - `call(self, inputs, training=None, **kwargs)` где `training` это булево значение показывающее в каком режиме должен вести себя слой - обучения или вывода.
# - `call(self, inputs, mask=None, **kwargs)` где `mask` это тензор булевой маски (полезно для RNN, например).
# - `call(self, inputs, training=None, mask=None, **kwargs)` -- конечно вы можете иметь одновременно все параметры, определяющие поведение слоя.
#
# В дополнение, если вы реализуете метод `get_config` на вашем пользовательском Layer или Model, Functional модели которые вы создадите с ним, будут сериализуемы и клонируемы.
#
# Далее приведем небольшой пример где мы используем кастомный RNN написанный с нуля в стиле Functional API:
# + id="TmTEZ6F3ArJR"
units = 32
timesteps = 10
input_dim = 5
batch_size = 16
class CustomRNN(layers.Layer):
def __init__(self):
super(CustomRNN, self).__init__()
self.units = units
self.projection_1 = layers.Dense(units=units, activation='tanh')
self.projection_2 = layers.Dense(units=units, activation='tanh')
self.classifier = layers.Dense(1, activation='sigmoid')
def call(self, inputs):
outputs = []
state = tf.zeros(shape=(inputs.shape[0], self.units))
for t in range(inputs.shape[1]):
x = inputs[:, t, :]
h = self.projection_1(x)
y = h + self.projection_2(state)
state = y
outputs.append(y)
features = tf.stack(outputs, axis=1)
return self.classifier(features)
# Заметьте, что мы задаем статичный размер пакета для входных данных с
# аргументом `batch_shape`, потому что внутренние вычисления `CustomRNN` требуют фиксированного размера пакета
# (когда мы создает нулевые тензоры `state`).
inputs = keras.Input(batch_shape=(batch_size, timesteps, input_dim))
x = layers.Conv1D(32, 3)(inputs)
outputs = CustomRNN()(x)
model = keras.Model(inputs, outputs)
rnn_model = CustomRNN()
_ = rnn_model(tf.zeros((1, 10, 5)))
# + [markdown] id="6VxcYb4qArlb"
# Это завершает наше руководство по Functional API!
#
# Теперь у вас под рукой мощный набор инструментов для построения моделей глубокого обучения.
| site/ru/guide/keras/functional.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Week 1
#
# ## Overview
#
# As explained in the [*Before week 1* notebook](http://nbviewer.jupyter.org/github/suneman/socialgraphs2019/blob/master/lectures/How_To_Take_This_Class.ipynb), each week of this class is an IPython notebook like this one. **_In order to follow the class, you simply start reading from the top_**, following the instructions.
#
# **Hint**: And you can ask me - or any of the friendly Teaching Assistants - for help at any point if you get stuck!
# ## Today
#
# This first lecture will go over a few different topics to get you started
#
# * First, I'll help you get your system in shape for all the Python programming we'll do later on.
# * Second, we'll make sure that you remember how to code in Python.
# * Third, we talk a bit about APIs and how they work.
# * Fourth, and finally, we'll be getting our hands dirty getting to know the awesome Network Analysis package `NetworkX`.
# ## Part 1: Installing Python
#
# Now it's time to install Python.
#
# * We recommend you use the _Anaconda distribution_ of Python. You can download it [**here**](https://www.anaconda.com/download/). We will be using Python 3!
# * You start up the notebook system by typing "`jupyter notebook`" and your terminal, and the system should be ready to use in your favorite browser.
# * Be sure to check the keyboards shortcuts under the heading of "Help" where you will find for instance shortcut to code-completion (Tab) and tooltip with documentation (Shift-Tab) which will save you a ton of time.
#
# Part 3 will teach you how to use the IPython Notebook. Note that if you want to use another Python distribution, that's fine, but we cannot promise to help you with anything other than Anaconda.
# ## Part 2: Simple Python exercises
#
# > **_Video lecture_**: You get started on this part by watching the "IPython Notebook" video below.
from IPython.display import YouTubeVideo
YouTubeVideo("yC754EgHpck",width=600, height=337.5)
# Oh, and I forgot a few important things when I made this video, so here are some additional tips & tricks on IPython.
#
# (*Note*: both of these videos were set up using Python 2.7, so there are a couple of differences now that we're using Python 3, e.g. how print works, but I leave that as an exercise for you to figure out the right syntax.)
YouTubeVideo("NDZyU_NlX0I",width=600, height=337.5)
# > _Exercises_
# >
# > * Download the IPython file that I've prepared for you and save it somewhere where you can find it again. The link is [**here**](https://raw.githubusercontent.com/suneman/socialgraphs2019/master/files/Training_notebook.ipynb). (**Hint**: Be careful not to save this in _.txt_ format - make sure the extension is _.ipynb_.)
# > * Work through exercise 1-9 in the file, solving the simple Python exercises in there. Also use this as a chance to become familiar with how the IPython notebook works. (And a little bit of `json`).
# ### STOP: Super important notice
#
# Now that you've completed working through the IPython notebook, it's time for the moment of truth! If you had great difficulty with the Python coding itself, you're going to be in trouble. Everything we do going forward in this class will depend on you being comfortable with Python. There is simply **no way** that you will be able to do well, if you're also struggling with Python on top of everything else you'll be learning.
#
# **So if you're not 100% comfortable with Python, I recommend you stop right now, and follow a tutorial to teach you Python, for example [this one](https://www.learnpython.org), before proceeding**. This might seem tough, but the ability to program is a prerequisite for this class, and if you know how to program, you should be able to handle the Python questions above.`
# ## Part 3: What is an API?
#
# Ok, so you're now on top of Python, so let's get started with a quick overview of APIs.
# >
# > **_Video lecture_**: Click on the image below to watch it on YouTube.
# >
YouTubeVideo("9l5zOfh0CRo",width=600, height=337.5)
# It's time for you to get to work. Take a look at the two texts below - just to get a sense of a more technical description of how APIs wor.
#
# Again, this is a Python 2 video, so small changes may apply. This video will be updated soon. Hint: **[Here](https://raw.githubusercontent.com/suneman/socialgraphs2019/master/files/API_check.ipynb)** is an Python3 version of the notebook used in the video.
#
#
# > _Reading_ (just skim): [Wikipedia page on APIs](https://en.wikipedia.org/wiki/Web_API)
# > _Reading_ (just skim): [Wikipedia page on REST for web services](https://en.wikipedia.org/wiki/Representational_state_transfer#Applied_to_web_services)
# >
# > *Exercise*:
# > * Explain in your own words: What is the the difference between the html page and the wiki-source?
# > * What are the various parameters you can set for a query of the wikipedia api?
# > * Write your own little `notebook` to download wikipedia pages based on the video above. Download the source for your 4 favorite wikipedia pages.
# # Part 4: The awesome `NetworkX` library
#
# In case you didn't know it, **this class is about analyzing networks**. And it wouldn't be right to start the first lecture without playing a little bit with network analysis (there will be much more on this in the following lectures). So here goes...
#
# `NetworkX` should already be installed as part of your _Anaconda_ Python distribution. But you don't know how to use it yet. The best way to get familiar is to work through a tutorial. That's what the next exercise is about
#
# > _Exercises_:
# >
# > * Go to the `NetworkX` project's [tutorial page](https://networkx.github.io/documentation/stable/tutorial.html). The goal of this exercise is to create your own `notebook` that contains the entire tutorial. You're free to add your own (e.g. shorter) comments in place of the ones in the official tutorial - and change the code to make it your own where ever it makes sense.
#
# There will be much more on NetworkX next time.
| lectures/Week1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# orphan: true
# ---
# # Running Tune experiments with BOHB
#
# In this tutorial we introduce BOHB, while running a simple Ray Tune experiment.
# Tune’s Search Algorithms integrate with BOHB and, as a result,
# allow you to seamlessly scale up a BOHB optimization
# process - without sacrificing performance.
#
# Bayesian Optimization HyperBand (BOHB) combines the benefits of Bayesian optimization
# together with Bandit-based methods (e.g. HyperBand). BOHB does not rely on
# the gradient of the objective function,
# but instead, learns from samples of the search space.
# It is suitable for optimizing functions that are non-differentiable,
# with many local minima, or even unknown but only testable.
# Therefore, this approach belongs to the domain of
# "derivative-free optimization" and "black-box optimization".
#
# In this example we minimize a simple objective to briefly demonstrate the usage of
# BOHB with Ray Tune via `BOHBSearch`. It's useful to keep in mind that despite
# the emphasis on machine learning experiments, Ray Tune optimizes any implicit
# or explicit objective. Here we assume `ConfigSpace==0.4.18` and `hpbandster==0.7.4`
# libraries are installed. To learn more, please refer to the
# [BOHB website](https://github.com/automl/HpBandSter).
# + tags=["remove-cell"]
# # !pip install ray[tune]
# !pip install ConfigSpace==0.4.18
# !pip install hpbandster==0.7.4
# -
# Click below to see all the imports we need for this example.
# You can also launch directly into a Binder instance to run this notebook yourself.
# Just click on the rocket symbol at the top of the navigation.
# + tags=["hide-input"]
import time
import ray
from ray import tune
from ray.tune.suggest import ConcurrencyLimiter
from ray.tune.schedulers.hb_bohb import HyperBandForBOHB
from ray.tune.suggest.bohb import TuneBOHB
import ConfigSpace as CS
# -
# Let's start by defining a simple evaluation function.
# We artificially sleep for a bit (`0.1` seconds) to simulate a long-running ML experiment.
# This setup assumes that we're running multiple `step`s of an experiment and try to tune
# two hyperparameters, namely `width` and `height`, and `activation`.
def evaluate(step, width, height, activation):
time.sleep(0.1)
activation_boost = 10 if activation=="relu" else 1
return (0.1 + width * step / 100) ** (-1) + height * 0.1 + activation_boost
# Next, our `objective` function takes a Tune `config`, evaluates the `score` of your
# experiment in a training loop, and uses `tune.report` to report the `score` back to Tune.
def objective(config):
for step in range(config["steps"]):
score = evaluate(step, config["width"], config["height"], config["activation"])
tune.report(iterations=step, mean_loss=score)
# + tags=["remove-cell"]
ray.init(configure_logging=False)
# -
# Next we define a search space. The critical assumption is that the optimal
# hyperparameters live within this space. Yet, if the space is very large,
# then those hyperparameters may be difficult to find in a short amount of time.
search_space = {
"steps": 100,
"width": tune.uniform(0, 20),
"height": tune.uniform(-100, 100),
"activation": tune.choice(["relu", "tanh"]),
}
# Next we define the search algorithm built from `TuneBOHB`, constrained
# to a maximum of `4` concurrent trials with a `ConcurrencyLimiter`.
# Below `algo` will take care of the BO (Bayesian optimization) part of BOHB,
# while scheduler will take care the HB (HyperBand) part.
algo = TuneBOHB()
algo = tune.suggest.ConcurrencyLimiter(algo, max_concurrent=4)
scheduler = HyperBandForBOHB(
time_attr="training_iteration",
max_t=100,
reduction_factor=4,
stop_last_trials=False,
)
# The number of samples is the number of hyperparameter combinations
# that will be tried out. This Tune run is set to `1000` samples.
# (you can decrease this if it takes too long on your machine).
num_samples = 1000
# + tags=["remove-cell"] pycharm={"name": "#%%\n"}
num_samples = 10
# -
# Finally, we run the experiment to `min`imize the "mean_loss" of the `objective`
# by searching within `"steps": 100` via `algo`, `num_samples` times. This previous
# sentence is fully characterizes the search problem we aim to solve.
# With this in mind, notice how efficient it is to execute `tune.run()`.
analysis = tune.run(
objective,
search_alg=algo,
scheduler=scheduler,
metric="mean_loss",
mode="min",
name="bohb_exp",
num_samples=num_samples,
config=search_space
)
# Here are the hyperparameters found to minimize the mean loss of the defined objective.
print("Best hyperparameters found were: ", analysis.best_config)
# ## Optional: Passing the search space via the TuneBOHB algorithm
#
# We can define the hyperparameter search space using `ConfigSpace`,
# which is the format accepted by BOHB.
config_space = CS.ConfigurationSpace()
config_space.add_hyperparameter(
CS.UniformFloatHyperparameter("width", lower=0, upper=20)
)
config_space.add_hyperparameter(
CS.UniformFloatHyperparameter("height", lower=-100, upper=100)
)
config_space.add_hyperparameter(
CS.CategoricalHyperparameter(
"activation", choices=["relu", "tanh"]
)
)
algo = TuneBOHB(
space=config_space,
metric="episode_reward_mean",
mode="max",
)
algo = tune.suggest.ConcurrencyLimiter(algo, max_concurrent=4)
scheduler = HyperBandForBOHB(
time_attr="training_iteration",
max_t=100,
reduction_factor=4,
stop_last_trials=False,
)
# + pycharm={"name": "#%%\n"}
analysis = tune.run(
objective,
config=config_space,
scheduler=bohb_hyperband,
search_alg=algo,
num_samples=num_samples,
name="bohb_exp_2",
stop={"training_iteration": 100},
)
# -
# Here again are the hyperparameters found to minimize the mean loss of the
# defined objective.
print("Best hyperparameters found were: ", analysis.best_config)
# + tags=["remove-cell"]
ray.shutdown()
| doc/source/tune/examples/bohb_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="wJcYs_ERTnnI"
# ##### Copyright 2021 The TensorFlow Authors.
# + cellView="form" id="HMUDt0CiUJk9"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="77z2OchJTk0l"
# # Migrate metrics and optimizers
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/guide/migrate/metrics_optimizers">
# <img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
# View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/migrate/metrics_optimizers.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
# Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/migrate/metrics_optimizers.ipynb">
# <img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
# View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/migrate/metrics_optimizers.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="meUTrR4I6m1C"
# In TF1, `tf.metrics` is the API namespace for all the metric functions. Each of the metrics is a function that takes `label` and `prediction` as input parameters and returns the corresponding metrics tensor as result. In TF2, `tf.keras.metrics` contains all the metric functions and objects. The `Metric` object can be used with `tf.keras.Model` and `tf.keras.layers.layer` to calculate metric values.
# + [markdown] id="YdZSoIXEbhg-"
# ## Setup
#
# Let's start with a couple of necessary TensorFlow imports,
# + id="iE0vSfMXumKI"
import tensorflow as tf
import tensorflow.compat.v1 as tf1
# + [markdown] id="Jsm9Rxx7s1OZ"
# and prepare some simple data for demonstration:
# + id="m7rnGxsXtDkV"
features = [[1., 1.5], [2., 2.5], [3., 3.5]]
labels = [0, 0, 1]
eval_features = [[4., 4.5], [5., 5.5], [6., 6.5]]
eval_labels = [0, 1, 1]
# + [markdown] id="xswk0d4xrFaQ"
# ## TF1: tf.compat.v1.metrics with Estimator
#
# In TF1, the metrics can be added to `EstimatorSpec` as the `eval_metric_ops`, and the op is generated via all the metrics functions defined in `tf.metrics`. You can follow the example to see how to use `tf.metrics.accuracy`.
# + id="lqe9obf7suIj"
def _input_fn():
return tf1.data.Dataset.from_tensor_slices((features, labels)).batch(1)
def _eval_input_fn():
return tf1.data.Dataset.from_tensor_slices(
(eval_features, eval_labels)).batch(1)
def _model_fn(features, labels, mode):
logits = tf1.layers.Dense(2)(features)
predictions = tf.argmax(input=logits, axis=1)
loss = tf1.nn.sparse_softmax_cross_entropy_with_logits(labels=labels, logits=logits)
optimizer = tf1.train.AdagradOptimizer(0.05)
train_op = optimizer.minimize(loss, global_step=tf1.train.get_global_step())
accuracy = tf1.metrics.accuracy(labels=labels, predictions=predictions)
return tf1.estimator.EstimatorSpec(mode,
predictions=predictions,
loss=loss,
train_op=train_op,
eval_metric_ops={'accuracy': accuracy})
estimator = tf1.estimator.Estimator(model_fn=_model_fn)
estimator.train(_input_fn)
# + id="HsOpjW5plH9Q"
estimator.evaluate(_eval_input_fn)
# + [markdown] id="Wk4C6qA_OaQx"
# Also, metrics could be added to estimator directly via `tf.estimator.add_metrics()`.
# + id="B2lpLOh9Owma"
def mean_squared_error(labels, predictions):
labels = tf.cast(labels, predictions.dtype)
return {"mean_squared_error":
tf1.metrics.mean_squared_error(labels=labels, predictions=predictions)}
estimator = tf1.estimator.add_metrics(estimator, mean_squared_error)
estimator.evaluate(_eval_input_fn)
# + [markdown] id="KEmzBjfnsxwT"
# ## TF2: Keras Metrics API with tf.keras.Model
#
# In TF2, `tf.keras.metrics` contains all the metrics classes and functions. They are designed in a OOP style and integrate closely with other `tf.keras` API. All the metrics can be found in `tf.keras.metrics` namespace, and there is usually a direct mapping between `tf.compat.v1.metrics` with `tf.keras.metrics`.
#
# In the following example, the metrics are added in `model.compile()` method. Users only need to create the metric instance, without specifying the label and prediction tensor. The Keras model will route the model output and label to the metrics object.
# + id="atVciNgPs0fw"
dataset = tf.data.Dataset.from_tensor_slices((features, labels)).batch(1)
eval_dataset = tf.data.Dataset.from_tensor_slices(
(eval_features, eval_labels)).batch(1)
inputs = tf.keras.Input((2,))
logits = tf.keras.layers.Dense(2)(inputs)
predictions = tf.argmax(input=logits, axis=1)
model = tf.keras.models.Model(inputs, predictions)
optimizer = tf.keras.optimizers.Adagrad(learning_rate=0.05)
model.compile(optimizer, loss='mse', metrics=[tf.keras.metrics.Accuracy()])
# + id="Kip65sYBlKiu"
model.evaluate(eval_dataset, return_dict=True)
# + [markdown] id="_mcGoCm_X1V0"
# With eager execution enabled, `tf.keras.metrics.Metric` instances can be directly used to evaluate numpy data or eager tensors. `tf.keras.metrics.Metric` objects are stateful containers. The metric value can be updated via `metric.update_state(y_true, y_pred)`, and the result can be retrieved by `metrics.result()`.
#
# + id="TVGn5_IhYhtG"
accuracy = tf.keras.metrics.Accuracy()
accuracy.update_state(y_true=[0, 0, 1, 1], y_pred=[0, 0, 0, 1])
accuracy.result().numpy()
# + id="wQEV2hHtY_su"
accuracy.update_state(y_true=[0, 0, 1, 1], y_pred=[0, 0, 0, 0])
accuracy.update_state(y_true=[0, 0, 1, 1], y_pred=[1, 1, 0, 0])
accuracy.result().numpy()
# + [markdown] id="E3F3ElcyadW-"
# For more details about `tf.keras.metrics.Metric`, please take a look for the API documentation at `tf.keras.metrics.Metric`, as well as the [migration guide](https://www.tensorflow.org/guide/effective_tf2#new-style_metrics_and_losses).
# + [markdown] id="eXKY9HEulxQC"
# ## Migrate TF1.x optimizers to Keras optimizers
#
# The optimizers in `tf.compat.v1.train`, such as the
# [Adam optimizer](https://www.tensorflow.org/api_docs/python/tf/compat/v1/train/AdamOptimizer)
# and the
# [gradient descent optimizer](https://www.tensorflow.org/api_docs/python/tf/compat/v1/train/GradientDescentOptimizer),
# have equivalents in `tf.keras.optimizers`.
#
# The table below summarizes how you can convert these legacy optimizers to their
# Keras equivalents. You can directly replace the TF1.x version with the TF2
# version unless additional steps (such as
# [updating the default learning rate](../../guide/effective_tf2.ipynb#optimizer_defaults))
# are required.
#
# Note that converting your optimizers
# [may make old checkpoints incompatible](./migrating_checkpoints.ipynb).
#
# <table>
# <tr>
# <th>TF1.x</th>
# <th>TF2</th>
# <th>Additional steps</th>
# </tr>
# <tr>
# <td>`tf.v1.train.GradientDescentOptimizer`</td>
# <td>`tf.keras.optimizers.SGD`</td>
# <td>None</td>
# </tr>
# <tr>
# <td>`tf.v1.train.MomentumOptimizer`</td>
# <td>`tf.keras.optimizers.SGD`</td>
# <td>Include the `momentum` argument</td>
# </tr>
# <tr>
# <td>`tf.v1.train.AdamOptimizer`</td>
# <td>`tf.keras.optimizers.Adam`</td>
# <td>Rename `beta1` and `beta2` arguments to `beta_1` and `beta_2`</td>
# </tr>
# <tr>
# <td>`tf.v1.train.RMSPropOptimizer`</td>
# <td>`tf.keras.optimizers.RMSprop`</td>
# <td>Rename the `decay` argument to `rho`</td>
# </tr>
# <tr>
# <td>`tf.v1.train.AdadeltaOptimizer`</td>
# <td>`tf.keras.optimizers.Adadelta`</td>
# <td>None</td>
# </tr>
# <tr>
# <td>`tf.v1.train.AdagradOptimizer`</td>
# <td>`tf.keras.optimizers.Adagrad`</td>
# <td>None</td>
# </tr>
# <tr>
# <td>`tf.v1.train.FtrlOptimizer`</td>
# <td>`tf.keras.optimizers.Ftrl`</td>
# <td>Remove the `accum_name` and `linear_name` arguments</td>
# </tr>
# <tr>
# <td>`tf.contrib.AdamaxOptimizer`</td>
# <td>`tf.keras.optimizers.Adamax`</td>
# <td>Rename the `beta1`, and `beta2` arguments to `beta_1` and `beta_2`</td>
# </tr>
# <tr>
# <td>`tf.contrib.Nadam`</td>
# <td>`tf.keras.optimizers.Nadam`</td>
# <td>Rename the `beta1`, and `beta2` arguments to `beta_1` and `beta_2`</td>
# </tr>
# </table>
#
# Note: In TF2, all epsilons (numerical stability constants) now default to `1e-7`
# instead of `1e-8`. This difference is negligible in most use cases.
| site/en/guide/migrate/metrics_optimizers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Setting up imports
# +
# %load_ext autoreload
# %autoreload 2
# %pdb off
from __future__ import print_function
from __future__ import division
from __future__ import unicode_literals
__author__ = "<NAME>"
__copyright__ = "Copyright 2016, Stanford University"
__license__ = "LGPL"
import os
import unittest
import tempfile
import shutil
import numpy as np
import numpy.random
from deepchem import metrics
from deepchem.datasets import Dataset
from deepchem.featurizers.featurize import DataFeaturizer
from deepchem.featurizers.featurize import FeaturizedSamples
from deepchem.hyperparameters import HyperparamOpt
from deepchem.metrics import Metric
from deepchem.models import Model
from deepchem.models.sklearn_models import SklearnModel
from deepchem.transformers import NormalizationTransformer
from deepchem.utils.evaluate import Evaluator
from sklearn.ensemble import RandomForestRegressor
from sklearn.kernel_ridge import KernelRidge
# -
# Creating temporary directories
feature_dir = tempfile.mkdtemp()
samples_dir = tempfile.mkdtemp()
train_dir = tempfile.mkdtemp()
valid_dir = tempfile.mkdtemp()
test_dir = tempfile.mkdtemp()
model_dir = tempfile.mkdtemp()
# Setting up model variables
from deepchem.featurizers.coulomb_matrices import CoulombMatrixEig
compound_featurizers = [CoulombMatrixEig(23, remove_hydrogens=False)]
complex_featurizers = []
tasks = ["atomization_energy"]
task_type = "regression"
task_types = {task: task_type for task in tasks}
input_file = "../datasets/gdb1k.sdf"
smiles_field = "smiles"
mol_field = "mol"
# Load featurized data
featurizers = compound_featurizers + complex_featurizers
featurizer = DataFeaturizer(tasks=tasks,
smiles_field=smiles_field,
mol_field=mol_field,
compound_featurizers=compound_featurizers,
complex_featurizers=complex_featurizers, verbosity="high")
featurized_samples = featurizer.featurize(input_file, feature_dir, samples_dir)
# Perform Train, Validation, and Testing Split
from deepchem.splits import RandomSplitter
random_splitter = RandomSplitter()
train_samples, valid_samples, test_samples = random_splitter.train_valid_test_split(featurized_samples,
train_dir, valid_dir, test_dir)
# Creating datasets
train_dataset = Dataset(data_dir=train_dir, samples=train_samples,
featurizers=featurizers, tasks=tasks)
valid_dataset = Dataset(data_dir=valid_dir, samples=valid_samples,
featurizers=featurizers, tasks=tasks)
test_dataset = Dataset(data_dir=test_dir, samples=test_samples,
featurizers=featurizers, tasks=tasks)
# Transforming datasets
input_transformers = [NormalizationTransformer(transform_X=True, dataset=train_dataset)]
output_transformers = [NormalizationTransformer(transform_y=True, dataset=train_dataset)]
transformers = input_transformers + output_transformers
for transformer in transformers:
transformer.transform(train_dataset)
for transformer in transformers:
transformer.transform(valid_dataset)
for transformer in transformers:
transformer.transform(test_dataset)
# Fit Random Forest with hyperparameter search
# +
def rf_model_builder(tasks, task_types, params_dict, model_dir, verbosity=None):
"""Builds random forests given hyperparameters.
"""
n_estimators = params_dict["n_estimators"]
max_features = params_dict["max_features"]
return SklearnModel(
tasks, task_types, params_dict, model_dir,
mode="regression",
model_instance=RandomForestRegressor(n_estimators=n_estimators,
max_features=max_features))
params_dict = {
"n_estimators": [10, 100],
"data_shape": [train_dataset.get_data_shape()],
"max_features": ["auto"],
}
metric = Metric(metrics.mean_absolute_error)
optimizer = HyperparamOpt(rf_model_builder, tasks, task_types, verbosity="low")
best_model, best_hyperparams, all_results = optimizer.hyperparam_search(
params_dict, train_dataset, valid_dataset, output_transformers,
metric, use_max="False", logdir=None)
# +
def kr_model_builder(tasks, task_types, params_dict, model_dir, verbosity=None):
"""Builds random forests given hyperparameters.
"""
kernel = params_dict["kernel"]
alpha = params_dict["alpha"]
gamma = params_dict["gamma"]
return SklearnModel(
tasks, task_types, params_dict, model_dir,
mode="regression",
model_instance=KernelRidge(alpha=alpha,kernel=kernel,gamma=gamma))
params_dict = {
"kernel": ["laplacian"],
"alpha": [0.0001],
"gamma": [0.0001]
}
metric = Metric(metrics.mean_absolute_error)
optimizer = HyperparamOpt(kr_model_builder, tasks, task_types, verbosity="low")
best_model, best_hyperparams, all_results = optimizer.hyperparam_search(
params_dict, train_dataset, valid_dataset, output_transformers,
metric, use_max="False", logdir=None)
# -
| examples/broken/quantum_machine_gdb1k.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Basic of Python
#
# In this video we will discuss about
#
# - Various Datatypes in Python
# - Variable
# - Variables Assigment
# - Print Formatting
#
# ## Various DataTypes in Python
#
# ### Numbers
# # Python Tutorial
#
# ## Welcome to Python Tutorial
#
#
# - Datatype
1+1
4*5
2*10
10/2
10%2
10**2
# +
##Check the Data types
# -
type(True)
type("Hello")
# ### Strings
"Hello"
'<NAME>'
type("<NAME>")
# ### Variable Assigment
# +
# syntax
# var_name=values
a=10
# -
type(a)
a='<NAME>'
type(a)
# +
## Mathematical operation with Variable Assigment
a=10
b=20
# -
print(a*b)
print(a/b)
print(a%b)
print((a*b)+(a/b)) ##BODMAS
# +
## Various Ways of Printing
print("Hello")
# -
first_name='Krish'
last_name='Naik'
print("My first name is {} and last name is {}".format(first_name,last_name))
print("My First name is {first} and last name is {last}".format(last=last_name,first=first_name))
len('Krish')
type(['1',2,3,4,5])
| Basics of Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
physio_diagnosis_train = pd.read_csv('data/physio_diagnosis_train.csv')
physio_appts_train = pd.read_csv('data/physio_appts_train.csv')
physio_diagnosis_train.info()
physio_appts_train.info()
physio_appts_train['appt_id'] = physio_appts_train['appt_id'].astype(str)
physio_diagnosis_train['Appt_id'] = physio_diagnosis_train['Appt_id'].astype(str)
md = pd.merge(physio_appts_train, physio_diagnosis_train, how='left', left_on=['appt_id', 'pat_id'], right_on=['Appt_id','patientId'])
md.info()
physio_diagnosis_train = physio_diagnosis_train.rename(columns={'Appt_id': 'appt_id'})
md.head()
md.tail()
pat_ids = [id for id in md[['patientId', 'pat_id']] if id['patiendId'] == id['patientId']]
len(pat_ids)
pat_ids = md[['patientId', 'pat_id']]
md_pats = [row['patientId'], row['pat_id'] for id,row in pat_ids.iterrows() if row['patientId'] == row['patientId'] and row['pat_id'] == row['pat_id']]
ls_patds = list()
for id, row in pat_ids.iterrows():
if row['pat_id'] != row['patientId']:
ls_patds.append((row['pat_id'], row['patientId']))
print(ls_patds)
len(ls_patds)
ls_patds
| 03_18_portea_challenge/merging_appts_diagnosis_appt_id_train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 4.3.3 グラフの種類と出力方法
import matplotlib.pyplot as plt
import matplotlib.style
matplotlib.style.use('ggplot')
# ## 折れ線グラフ
# +
fig, ax = plt.subplots()
x = [1, 2, 3]
y1 = [1, 2, 3]
y2 = [3, 1, 2]
ax.plot(x, y1) # 折れ線グラフを描画
ax.plot(x, y2)
plt.show()
# +
import numpy as np
x = np.arange(0.0, 15.0, 0.1)
y1 = np.sin(x)
y2 = np.cos(x)
fig, ax = plt.subplots()
ax.plot(x, y1, label='sin')
ax.plot(x, y2, label='cos')
ax.legend()
plt.show()
# -
# ## 棒グラフ
# +
fig, ax = plt.subplots()
x = [1, 2, 3]
y = [10, 2, 3]
ax.bar(x, y) # 棒グラフを描画
plt.show()
# +
fig, ax = plt.subplots()
x = [1, 2, 3]
y = [10, 2, 3]
labels = ['spam', 'ham', 'egg']
ax.bar(x, y, tick_label=labels) # ラベルを指定
plt.show()
# +
fig, ax = plt.subplots()
x = [1, 2, 3]
y = [10, 2, 3]
labels = ['spam', 'ham', 'egg']
ax.barh(x, y, tick_label=labels) # 横向きの棒グラフを描画
plt.show()
# +
fig, ax = plt.subplots()
x = [1, 2, 3]
y1 = [10, 2, 3]
y2 = [5, 3, 6]
labels = ['spam', 'ham', 'egg']
width = 0.4 # 棒グラフの幅を0.4にする
ax.bar(x, y1, width=width, tick_label=labels,
label='y1') # 幅を指定して描画
# 幅分ずらして棒グラフを描画
x2 = [num + width for num in x]
ax.bar(x2, y2, width=width, label='y2')
ax.legend()
plt.show()
# +
fig, ax = plt.subplots()
x = [1, 2, 3]
y1 = [10, 2, 3]
y2 = [5, 3, 6]
labels = ['spam', 'ham', 'egg']
# y1とy2を足した値を格納
y_total = [num1 + num2 for num1, num2 in zip(y1, y2)]
# y1とy2を足した高さの棒グラフを描画
ax.bar(x, y_total, tick_label=labels, label='y1')
ax.bar(x, y2, label='y2')
ax.legend()
plt.show()
# -
# ## 散布図
# +
fig, ax = plt.subplots()
# ランダムに50個の要素を生成
np.random.seed(123)
x = np.random.rand(50)
y = np.random.rand(50)
ax.scatter(x, y) # 散布図を描画
plt.show()
# +
fig, ax = plt.subplots()
# ランダムに50個の要素を生成
np.random.seed(123)
x = np.random.rand(50)
y = np.random.rand(50)
ax.scatter(x[0:10], y[0:10], marker='v',
label='traiangle down') # 下向き三角
ax.scatter(x[10:20], y[10:20], marker='^',
label='traiangle up') # 上向き三角
ax.scatter(x[20:30], y[20:30], marker='s',
label='square') # 正方形
ax.scatter(x[30:40], y[30:40], marker='*',
label='star') # 星形
ax.scatter(x[40:50], y[40:50], marker='x',
label='x') # X
ax.legend()
plt.show()
# -
# ## ヒストグラム
# +
# データを生成
np.random.seed(123)
mu = 100 # 平均値
sigma = 15 # 標準偏差
x = np.random.normal(mu, sigma, 1000)
fig, ax = plt.subplots()
# ヒストグラムを描画
n, bins, patches = ax.hist(x)
plt.show()
# -
# 度数分布表を出力
for i, num in enumerate(n):
print('{:.2f} - {:.2f}: {}'.format(bins[i], bins[i+1], num))
# +
fig, ax = plt.subplots()
ax.hist(x, bins=25) # ビンの数を指定して描画
plt.show()
# +
fig, ax = plt.subplots()
# 横向きのヒストグラムを描画
ax.hist(x, orientation='horizontal')
plt.show()
# +
# データを生成
np.random.seed(123)
mu = 100 # 平均値
x0 = np.random.normal(mu, 20, 1000)
# 異なる標準偏差でデータを生成
x1 = np.random.normal(mu, 15, 1000)
x2 = np.random.normal(mu, 10, 1000)
fig, ax = plt.subplots()
labels = ['x0', 'x1', 'x2']
# 3つのデータのヒストグラムを描画
ax.hist((x0, x1, x2), label=labels)
ax.legend()
plt.show()
# +
fig, ax = plt.subplots()
labels= ['x0', 'x1', 'x2']
# 積み上げたヒストグラムを描画
ax.hist((x0, x1, x2), label=labels, stacked=True)
ax.legend()
plt.show()
# -
# ## 箱ひげ図
# +
# 異なる標準偏差でデータを生成
np.random.seed(123)
x0 = np.random.normal(0, 10, 500)
x1 = np.random.normal(0, 15, 500)
x2 = np.random.normal(0, 20, 500)
fig, ax = plt.subplots()
labels = ['x0', 'x1', 'x2']
ax.boxplot((x0, x1, x2), labels=labels) # 箱ひげ図を描画
plt.show()
# +
fig, ax = plt.subplots()
labels = ['x0', 'x1', 'x2']
# 横向きの箱ひげ図を描画
ax.boxplot((x0, x1, x2), labels=labels, vert=False)
plt.show()
# -
# ## 円グラフ
# +
labels = ['spam', 'ham', 'egg']
x = [10, 3, 1]
fig, ax = plt.subplots()
ax.pie(x, labels=labels) # 円グラフを描画
plt.show()
# +
fig, ax = plt.subplots()
ax.pie(x, labels=labels)
ax.axis('equal') # アスペクト比を保持して描画する
plt.show()
# +
fig, ax = plt.subplots()
ax.pie(x, labels=labels, startangle=90,
counterclock=False) # 上から時計回り
ax.axis('equal')
plt.show()
# +
fig, ax = plt.subplots()
ax.pie(x, labels=labels, startangle=90,
counterclock=False,
shadow=True, autopct='%1.2f%%') # 影と%表記を追加
ax.axis('equal')
plt.show()
# +
explode = [0, 0.2, 0] # 1番目の要素 (ham) を切り出す
fig, ax = plt.subplots()
ax.pie(x, labels=labels, startangle=90,
counterclock=False,
shadow=True, autopct='%1.2f%%',
explode=explode) # explodeを指定する
ax.axis('equal')
plt.show()
# -
# ## 複数のグラフを組み合わせる
# +
fig, ax = plt.subplots()
x1 = [1, 2, 3]
y1 = [5, 2, 3]
x2 = [1, 2, 3, 4]
y2 = [9, 5, 4, 6]
ax.bar(x1, y1, label='y1') # 棒グラフを描画
ax.plot(x2, y2, label='y2') # 折れ線グラフを描画
ax.legend()
plt.show()
# +
np.random.seed(123)
x = np.random.randn(1000)
fig, ax = plt.subplots()
# ヒストグラムを描画
counts, edges, patches = ax.hist(x, bins=25)
# 近似曲線に用いる点を求める(ヒストグラムのビンの中点)
x_fit = (edges[:-1] + edges[1:]) / 2
# 近似曲線をプロット
y = 1000 * np.diff(edges) * np.exp(-x_fit**2 / 2) / np.sqrt(2 * np.pi)
ax.plot(x_fit, y)
plt.show()
| ds-newtextbook-python/notebooks/4-3-3-Matplotlib-graphs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + slideshow={"slide_type": "slide"}
# # %load ../standard_import.txt
from IPython.display import Image
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import statsmodels.api as sm
import statsmodels.formula.api as smf
# %matplotlib inline
# + [markdown] slideshow={"slide_type": "slide"}
# ### Logistic Regrssion
#
# * The linear regression model discussed assumes response is quantitative
# * But in many situations, the response variable is instead qualitative
#
# * Logistic Regression for the outcome to be qualitative (or categorical)
#
# * The main difference between Linear versus logistic regression is in the type of outcomes predicted
#
# * Email: SPAM vs. HAM
# * An individual will get a disease or not
# * Etc.
#
# * Logistic Regression is one among many solutions to classification
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Classification
#
# * Classification takes a set of inputs and assigns them to a qualitative outcome
# * A qualitative outcome is one that takes values in a set of possible categories
#
# * Often, we are interested in predicting the probability that an input belongs to each category
# * We can then take the assignment with the highest probability
# * I.e., more relevant to say a computer is infected with 0.99 probability or 0.52 probability than just infected
# + [markdown] slideshow={"slide_type": "slide"}
# ### Classes of Logistic Regression
#
# * There are two types of logistic regression into:
# * Simple or binary logistic regression
# * Multiclass logistic regression
# * We focus on (binary) logistic regression
# * Ease to generalize to more than two classes
# + [markdown] slideshow={"slide_type": "slide"}
# ### Dataset
#
# * People who default on their credit card paymment (brown marks) versus those who don't (blue circles)
#
# <img src="images/classification_1.png" alt="drawing" style="width:1000px;"/>
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Justification for logistic regression
#
# - Can we use linear regression to determine whether a client defaults on credit card payment? How?
# - One possible way is to assign given output to classes (say 0 and 1) and use linear regression to generate value that span the 0,1 range
#
# $$
# \left\{
# \begin{array}{ll}
# 1 & \mbox{if default} \\
# 0 & \mbox{otherwise}
# \end{array}
# \right.
# $$
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Using Linear in Classification
#
# <img src="images/regression_1.png" alt="drawing" style="width:500px;"/>
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Using Linear in Classification - Cont'd
#
#
# * Remember that linear regression predicts continuous values
# * We can say then that:
# * Any value that our linear regression predicts as above a certain threshold represents class 1
# * Any value that our linear regression predicts below a certain threshold represents class 0
# * 0.5 is a reasonable threshold as it sits exactly in the middle classes 0 and 1
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Using Linear in Classification - Model
#
# <img src="images/regression_2.png" alt="drawing" style="width:500px;"/>
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Using Linear in Classification - Prediction
#
# <img src="images/regression_3.png" alt="drawing" style="width:800px;"/>
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Using Linear in Classification - Prediction Regions
#
# <img src="images/regression_4.png" alt="drawing" style="width:800px;"/>
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Inconvenients of Using Linear Regression
#
# * Problems with "hacking" linear regression for classification
#
#
# * The regression line is sensitive to the data
# * outliers can drastically change the intersection point of the threshold on the line
#
#
# * Predicted value can be much smaller than 0 or much larger than 1
# * Impossible to interpret as a probability
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="images/regression_5.png" alt="drawing" style="width:500px;"/>
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Logistic Regression
#
# * A very simple yet popular method for classification
#
# * Constrains the predicted value to be between 0 and 1
# * Can be interpreted as a probability for a class
#
# Use the logistic, or sigmoid function, to model the separation between the classes
#
#
# $$ g(x) = \frac{1}{1+e^-(\beta_0 + \beta_1~x)}$$
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Logistic Regression
#
# * Assuming $\beta_0=0$ and $\beta_1=1$
#
# <img src="images/logistic_regression.png" alt="drawing" style="width:700px;"/>
#
# * The logistic function g(x) provides $prob(y=1~|~x, \beta_1, \beta_2)$
# * The probability that a value x is 1.
#
# + slideshow={"slide_type": "slide"}
x_axis = np.arange(-10, 10, 0.05)
y_axis = 1 / (1 + np.e**-x_axis)
plt.figure(figsize=(12,5))
plt.plot(x_axis, y_axis)
plt.axhline(0.5, linestyle="--", color='r', alpha=0.2)
plt.axvline(0, linestyle="--", color='k', alpha=0.2)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Parameters of the Logstic
# - As with the linear regression, we assumed that the params of the model $\beta_0$ and $\beta_1$ can be anything
# + slideshow={"slide_type": "slide"}
np.exp(-(beta_0 + beta_1*x))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Changing $\beta_0$
#
# <img src="images/changing_beta0.png" alt="drawing" style="width:1500px;"/>
#
# + slideshow={"slide_type": "slide"}
plt.figure(figsize=(20,5))
plt.subplot(1, 2, 1)
plt.axhline(0.5, linestyle="--", color='r', alpha=0.2)
plt.axvline(0, linestyle="--", color='k', alpha=0.2)
x = np.arange(-10, 10, 0.05)
beta_0, beta_1 = (0, 1)
y = 1 / (1 + np.e **-(beta_0 + beta_1*x))
plt.plot(x, y, label = r'$\beta_0 = {}, \beta_1 = {}$'.format(beta_0, beta_1))
beta_0, beta_1 = (-5, 1)
y = 1 / (1 + np.e **-(beta_0 + beta_1*x))
plt.plot(x, y, label = r'$\beta_0 = {}, \beta_1 = {}$'.format(beta_0, beta_1))
beta_0, beta_1 = (5, 1)
y = 1 / (1 + np.e **-(beta_0 + beta_1*x))
plt.plot(x, y, label = r'$\beta_0 = {}, \beta_1 = {}$'.format(beta_0, beta_1))
plt.legend()
plt.subplot(1, 2, 2)
beta_0, beta_1 = (0, -1)
y = 1 / (1 + np.e **-(beta_0 + beta_1*x))
plt.plot(x, y, label = r'$\beta_0 = {}, \beta_1 = {}$'.format(beta_0, beta_1))
beta_0, beta_1 = (-5, -1)
y = 1 / (1 + np.e **-(beta_0 + beta_1*x))
plt.plot(x, y, label = r'$\beta_0 = {}, \beta_1 = {}$'.format(beta_0, beta_1))
beta_0, beta_1 = (5, -1)
y = 1 / (1 + np.e **-(beta_0 + beta_1*x))
plt.plot(x, y, label = r'$\beta_0 = {}, \beta_1 = {}$'.format(beta_0, beta_1))
plt.legend()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Changing $\beta_1$
#
# <img src="images/logistic_beta_1.png" alt="drawing" style="width:1500px;"/>
#
# + slideshow={"slide_type": "slide"}
plt.figure(figsize=(20,5))
plt.subplot(1, 2, 1)
plt.axhline(0.5, linestyle="--", color='r', alpha=0.2)
plt.axvline(0, linestyle="--", color='k', alpha=0.2)
x = np.arange(-10, 10, 0.05)
beta_0, beta_1 = (0, 0.6)
y = 1 / (1 + np.e **-(beta_0 + beta_1*x))
plt.plot(x, y, label = r'$\beta_0 = {}, \beta_1 = {}$'.format(beta_0, beta_1))
beta_0, beta_1 = (0, 1)
y = 1 / (1 + np.e **-(beta_0 + beta_1*x))
plt.plot(x, y, label = r'$\beta_0 = {}, \beta_1 = {}$'.format(beta_0, beta_1))
beta_0, beta_1 = (0, 20)
y = 1 / (1 + np.e **-(beta_0 + beta_1*x))
plt.plot(x, y, label = r'$\beta_0 = {}, \beta_1 = {}$'.format(beta_0, beta_1))
plt.legend()
plt.subplot(1, 2, 2)
beta_0, beta_1 = (0, -0.6)
y = 1 / (1 + np.e **-(beta_0 + beta_1*x))
plt.plot(x, y, label = r'$\beta_0 = {}, \beta_1 = {}$'.format(beta_0, beta_1))
beta_0, beta_1 = (0, -1)
y = 1 / (1 + np.e **-(beta_0 + beta_1*x))
plt.plot(x, y, label = r'$\beta_0 = {}, \beta_1 = {}$'.format(beta_0, beta_1))
beta_0, beta_1 = (0, -20)
y = 1 / (1 + np.e **-(beta_0 + beta_1*x))
plt.plot(x, y, label = r'$\beta_0 = {}, \beta_1 = {}$'.format(beta_0, beta_1))
plt.legend()
# + slideshow={"slide_type": "slide"}
x_axis = np.arange(-10, 10, 0.05)
y_axis = 1 / (1 + np.e**-x_axis)
plt.figure(figsize=(12,6))
plt.plot([2.5], [1 / (1 + np.e**-2.5)], 'or', markersize=12)
plt.plot(x_axis, y_axis)
plt.axhline(0.5, linestyle="--", color='k', alpha=0.2)
plt.axvline(0, linestyle="--", color='k', alpha=0.2)
plt.axhline(1 / (1 + np.e**-2.5), linestyle="--", color='r', alpha=0.2)
plt.axvline(2.5, linestyle="--", color='r', alpha=0.2)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Fitting the Model: Linear Regression Parallel
#
# * For the Linear regression, we used the data to parameterize the model
# * We trained the model to find the line that minimizes the RSS
#
# * I.e, among all possible lines, the one selected has the smallest sum of squared residuals
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Fitting the Model
#
# * In binary logistic regression, the outcomes are 0 and 1, and the predictions are also either 0 or 1
#
# * One trivial approach would consist in identifying the sigmoid function that minimizes the mispredictions
# * Using the same method as in linear regression, we could describe the function as:
#
# $$
# \frac{1}{n} \sum_1^n (g(x) - y)
# $$
#
# * Unfortunately, this function is not convex
#
# * Does guarantee that the function will converge to the global minimum
# + [markdown] slideshow={"slide_type": "slide"}
# ### Fitting the Model- Cont'd
#
#
# - Instead, given some tuple (beta_0 and beta_1), the following convex function is commonly used to compute the cost
#
# $$
# cost(g(x),y) = \left\{
# \begin{array}{ll}
# -log(g(x)) & \mbox{ if } y = 1 \\
# -log(1 - g(x)) & \mbox{ if } y = 0 \\
# \end{array}
# \right.
# $$
#
# * Remember that $g(x)$ is simply the probability that the class is 1
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Cost Function Explained - $y=1$
#
# $$
# -log(g(x)) \mbox{ if } y = 1
# $$
#
# * For $y = 1$
# - If the prediction is also 1, then we incur no cost
# - If the prediction is 0, then we are incurring an infinite cost
# + slideshow={"slide_type": "slide"}
x_axis = np.arange(0.000001, 1.1, 0.05)
y_axis = - np.log(x_axis)
plt.plot(x_axis, y_axis)
plt.xlim(0,1)
plt.ylim(0,3)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Cost Function Explained - $y=1$
#
# $$
# -log(1 - g_(x)) \mbox{ if } y = 0 \\
# $$
# - for $y = 0$
# - If the prediction is also 0, then we incur no cost
# - If the prediction is 1, then we are incurring an infinite cost
#
# + slideshow={"slide_type": "slide"}
x_axis = np.arange(0, 0.9999999, 0.05)
y_axis = - np.log(1 - x_axis)
plt.plot(x_axis, y_axis)
plt.xlim(0,1)
plt.ylim(0,3)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Training
#
# - Therefore, by averaging the cost over all instances in the data, we end up with the parameters ($\beta$s) that best fit the data
#
# - Thus for a dataset of size $n$, we pick the tuple $\beta$ of params which minimizes:
#
# $$
# \sum_{\forall x}cost(g(x)_{\beta},y)~~/~~n
# $$
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Prediction
#
# * Use the parameters $\beta$ identified in the training to predict the probabality of true using
# $$
# p(y=1| x, \beta_0,\beta_1) = g(x) = \frac{1}{1+e^{\beta_0+\beta_1~x}}
# $$
# * Probabolty of False is merely $1- p(y=1| x, \beta_0,\beta_1)$
#
# + slideshow={"slide_type": "slide"}
data = pd.read_excel("data/Default.xlsx")
data.head()
# + slideshow={"slide_type": "slide"}
data["encoded_default"] = data["default"].map({"No":0, "Yes":1})
data.head()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Logistic Regression with `sklearn`
#
# * Using statsmodels, we can infer the values of \beta_0 and \beta_1 such that
#
# $$
# default = \frac{1}{1 + e^{-(\beta_o + \beta_1 ~ balance})} + \epsilon_x
# $$
#
# * Similar to the regression, $\epsilon$ is the noise component
#
# + slideshow={"slide_type": "slide"}
data.loc[:, "balance"]
# + slideshow={"slide_type": "slide"}
Y.ravel()
# + slideshow={"slide_type": "slide"}
from sklearn.linear_model import LogisticRegression
logreg = LogisticRegression(C=1e5, solver='lbfgs')
X = data["balance"].values.reshape(-1,1)
Y = data["encoded_default"].values
logreg.fit(X, Y)
# + slideshow={"slide_type": "slide"}
logreg.coef_, logreg.intercept_
# + [markdown] slideshow={"slide_type": "slide"}
# ### Model Interpretation - Balance
#
#
# $$
# P(default=1| x, \beta_0, \beta_1) \approx \frac{1}{1 + e^{10 - 0.0005 ~ balance}}
# $$
#
# - Note that $\beta_1$ reflects the units, which in this case is dollar
#
# - Therefore, an increase of a balnce of $1, increases the probability of defaulting by
#
# $$
# P(default=1| x=1, \beta_0=-10.65, \beta_1=0.0055) \approx \frac{1}{1 + e^{10 - 0.0005 * 1}} = 0.000045
# $$
#
# - An increase of a balnce of $2,000, increases the probability of defaulting by
#
# $$
# P(default=1| x=2,000, \beta_0=-10.65, \beta_1=0.0055) \approx \frac{1}{1 + e^{10 - 0.0005 * 2,000}} \approx 0.5
# $$
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Multiple Logistic Regression
# + slideshow={"slide_type": "slide"}
data["encoded_student"] = data["student"].map({"No":0, "Yes":1})
data.head()
# + slideshow={"slide_type": "slide"}
X = data["encoded_student"].values.reshape(-1,1)
y = data["encoded_default"].values
logreg.fit(X, Y)
logreg.coef_, logreg.intercept_
# + [markdown] slideshow={"slide_type": "slide"}
# ### Model Interpretation - Student Status
#
#
# $$
# P(default=1| x, \beta_0, \beta_1) \approx \frac{1}{1 + e^{-3.5 - 0.4 ~ student}}
# $$
#
# - the probability of a student defaulting is
#
# $$
# P(default=1| x=1, \beta_0=-3.5, \beta_1=0.4) \approx \frac{1}{1 + e^{3.5 - 0.4 ~ 1}} \approx 0.04
# $$
#
#
# - the probability of a non student defaulting is
#
# $$
# P(default=1 | x=0, \beta_0=-3.5, \beta_1=0.4) \approx \frac{1}{1 + e^{-3.5 - 0.4 ~ student}} \approx 0.02
# $$
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Multiple Logistic Regression
#
# * As we added more relevant variables, the logistic regression model becomes more discriminative
#
#
# * Think of the FICO score for instance
#
# * We can combine into a single regression the student status, total debt, age, income, number of recent credit applications, etc.
# + slideshow={"slide_type": "slide"}
X = data[['balance', 'encoded_student']]
y = data["encoded_default"].values
logreg.fit(X, Y)
logreg.coef_, logreg.intercept_
# + [markdown] slideshow={"slide_type": "slide"}
# $$
# P(default=1| x, \beta_0, \beta_1) \approx \frac{1}{1 + e^{10 - 0.0005 balance + 0.71 ~ student}}
# $$
# + slideshow={"slide_type": "slide"}
import matplotlib as mpl
import seaborn as sns
fig = plt.figure(figsize=(12,5))
gs = mpl.gridspec.GridSpec(1, 4)
ax1 = plt.subplot(gs[0,:-2])
ax2 = plt.subplot(gs[0,-2])
ax3 = plt.subplot(gs[0,-1])
# Take a fraction of the samples where target value (default) is 'no'
df_no = data[data["encoded_default"] == 0].sample(frac=0.15)
# Take all samples where target value is 'yes'
df_yes = data[data["encoded_default"] == 1]
df_ = df_no.append(df_yes)
ax1.scatter(df_[df_.default == 'Yes'].balance, df_[df_.default == 'Yes'].income, s=40, c='orange', marker='+',
linewidths=1)
ax1.scatter(df_[df_.default == 'No'].balance, df_[df_.default == 'No'].income, s=40, marker='o', linewidths='1',
edgecolors='lightblue', facecolors='white', alpha=.6)
ax1.set_ylim(ymin=0)
ax1.set_ylabel('Income')
ax1.set_xlim(xmin=-100)
ax1.set_xlabel('Balance')
c_palette = {'No':'lightblue', 'Yes':'orange'}
sns.boxplot('default', 'balance', data=data, orient='v', ax=ax2, palette=c_palette)
sns.boxplot('default', 'income', data=data, orient='v', ax=ax3, palette=c_palette)
gs.tight_layout(plt.gcf())
# + [markdown] slideshow={"slide_type": "slide"}
# ### Inspecting the Contribution of Income
#
# - The plot above did not indicate that income is useful for separating the data
#
# - What does the logistic regression model say about the contribution of the variable income?
# + slideshow={"slide_type": "slide"}
X = sm.add_constant(data[['balance', 'income', 'encoded_student']])
y = data["encoded_default"].values
logreg.fit(X, Y)
logreg.coef_, logreg.intercept_
# + [markdown] slideshow={"slide_type": "slide"}
# ### Categortical Variables as Predictors
#
# * A Categorical variable (sometimes called a nominal variable) is one that has two or more categories
#
# * categorical variables have no intrinsic ordering to the categories
#
# * For instance, car types can be encoded as electric (0), sedan (1), compact (2), SUV (3), etc...
#
# * The resulting numerical encoding does not have the same meaning as regular numerical values
#
# * Ex.: we cannot use the encoding to compute Euclidean distance or to compute averages on features
#
# * Data values for electric 0 and sedan 1 are not more similar than those with the values 0 and 2 ('electric' and 'compact')
#
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### One-Hot-Encoding
#
#
# * Categorical variables are commonly encoded using attributes for each category
#
# * The value is 1 for the attribute if the instance is in that category or the value is 0 otherwise
#
# * Ex.:
# * if a data point is of type electric vehicle 0, then its encoding would be [1,0,0,0]
# * if a data point is in SUV 4, then its encoding would be [0,0,0,1]
#
# * etc...
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Relationship Between Logistic and Linear Model
#
# We have:
#
# $$p(x) = \frac{1}{1+e^-(\beta_0 + \beta_1~x)}$$
#
# or:
#
# $$p(x) = \frac{e^{(\beta_0 + \beta_1~x)}}{1+e^{e(\beta_0 + \beta_1~x)}}$$
#
# This can be written as
#
# $$\frac{p(x)}{1-p(x)} = e^{(\beta_0 + \beta_1~x)}$$
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Relationship Between Logistic and Linear Model - Cont'd
#
#
# * The ratio $\frac{p(x)}{1-p(x)}$ is the odd ratio
# * Takes values between 0 and $\inf$
#
#
# * Say the probavility of defaulting is p(x) = 0.2, then $\frac{p(x)}{1-p(x)} = 1/5$
# * So, 1 in 5 people will defautl
# + [markdown] slideshow={"slide_type": "slide"}
# ### Relationship Between Logistic and Linear Model - Cont'd
#
# We have that:
#
# $$\frac{p(x)}{1-p(x)} = e^{(\beta_0 + \beta_1~x)}$$
#
# By taking log on both side, we have:
#
# This can be written as
#
# $$log(\frac{p(x)}{1-p(x)}) =(\beta_0 + \beta_1~x)$$
#
# * Left hand side if called log-odds or logit
#
# * Therefore, the linear model has a logit that is linear in $X$
#
# Unline in linear model, $\beta_1$ does not correspond to the change associate in one-unit increase in X.
#
# -
| morea/ML_intro/resources/.ipynb_checkpoints/logistic_regression-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Conversion Script for Criteo Dataset (CSV-to-Parquet)
#
# __Step 1__: Import libraries
# +
import os
import rmm
import cudf
from cudf.io.parquet import ParquetWriter
from fsspec.core import get_fs_token_paths
import numpy as np
import pyarrow.parquet as pq
from dask.dataframe.io.parquet.utils import _analyze_paths
from dask.base import tokenize
from dask.utils import natural_sort_key
from dask.highlevelgraph import HighLevelGraph
from dask.delayed import Delayed
from dask.distributed import Client
from dask_cuda import LocalCUDACluster
import nvtabular as nvt
from nvtabular.io import device_mem_size
# -
# __Step 2__: Specify options
#
# Specify the input and output paths, unless the `INPUT_DATA_DIR` and `OUTPUT_DATA_DIR` environment variables are already set. In order to utilize a multi-GPU system, be sure to specify `allow_multi_gpu=True` (and check the setting of your `CUDA_VISIBLE_DEVICES` environment variable).
INPUT_PATH = os.environ.get('INPUT_DATA_DIR', '/datasets/criteo/crit_orig')
OUTPUT_PATH = os.environ.get('OUTPUT_DATA_DIR', '/raid/criteo/tests/demo_out')
CUDA_VISIBLE_DEVICES = os.environ.get("CUDA_VISIBLE_DEVICES", "0")
n_workers = len(CUDA_VISIBLE_DEVICES.split(","))
frac_size = 0.15
allow_multi_gpu = False
use_rmm_pool = False
max_day = None # (Optional) -- Limit the dataset to day 0-max_day for debugging
# __Step 3__: Define helper/task functions
# +
def _pool(frac=0.8):
rmm.reinitialize(
pool_allocator=True,
initial_pool_size=frac * device_mem_size(),
)
def _convert_file(path, name, out_dir, frac_size, fs, cols, dtypes):
fn = f"{name}.parquet"
out_path = fs.sep.join([out_dir, f"{name}.parquet"])
writer = ParquetWriter(out_path, compression=None)
for gdf in nvt.Dataset(
path,
engine="csv",
names=cols,
part_memory_fraction=frac_size,
sep='\t',
dtypes=dtypes,
).to_iter():
writer.write_table(gdf)
del gdf
md = writer.close(metadata_file_path=fn)
return md
def _write_metadata(md_list, fs, path):
rg_sizes = []
if md_list:
metadata_path = fs.sep.join([path, "_metadata"])
_meta = (
cudf.io.merge_parquet_filemetadata(md_list)
if len(md_list) > 1
else md_list[0]
)
with fs.open(metadata_path, "wb") as fil:
_meta.tofile(fil)
return True
# -
# __Step 4__: (Optionally) Start a Dask cluster
# Start up cluster if we have multiple devices
# (and `allow_multi_gpu == True`)
client = None
if n_workers > 1 and allow_multi_gpu:
cluster = LocalCUDACluster(
n_workers=n_workers,
CUDA_VISIBLE_DEVICES=CUDA_VISIBLE_DEVICES,
)
client = Client(cluster)
if use_rmm_pool:
client.run(_pool)
elif use_rmm_pool:
_pool()
# __Step 5__: Main conversion script (build Dask task graph)
# +
fs = get_fs_token_paths(INPUT_PATH, mode="rb")[0]
file_list = [
x for x in fs.glob(fs.sep.join([INPUT_PATH, "day_*"]))
if not x.endswith("parquet")
]
file_list = sorted(file_list, key=natural_sort_key)
file_list = file_list[:max_day] if max_day else file_list
name_list = _analyze_paths(file_list, fs)[1]
cont_names = ["I" + str(x) for x in range(1, 14)]
cat_names = ["C" + str(x) for x in range(1, 27)]
cols = ["label"] + cont_names + cat_names
dtypes = {}
dtypes["label"] = np.int32
for x in cont_names:
dtypes[x] = np.int32
for x in cat_names:
dtypes[x] = "hex"
dsk = {}
token = tokenize(file_list, name_list, OUTPUT_PATH, frac_size, fs, cols, dtypes)
convert_file_name = "convert_file-" + token
for i, (path, name) in enumerate(zip(file_list, name_list)):
key = (convert_file_name, i)
dsk[key] = (_convert_file, path, name, OUTPUT_PATH, frac_size, fs, cols, dtypes)
write_meta_name = "write-metadata-" + token
dsk[write_meta_name] = (
_write_metadata,
[(convert_file_name, i) for i in range(len(file_list))],
fs,
OUTPUT_PATH,
)
graph = HighLevelGraph.from_collections(write_meta_name, dsk, dependencies=[])
conversion_delayed = Delayed(write_meta_name, graph)
# -
# **__Step 6__**: Execute conversion
# %%time
if client:
conversion_delayed.compute()
else:
conversion_delayed.compute(scheduler="synchronous")
| examples/optimize_criteo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://www.bigdatauniversity.com"><img src="https://ibm.box.com/shared/static/qo20b88v1hbjztubt06609ovs85q8fau.png" width="400px" align="center"></a>
#
# <h1 align="center"><font size="5">AUTOENCODERS</font></h1>
# <div class="alert alert-block alert-info" style="margin-top: 20px">
# Welcome to this notebook about autoencoders.
# <font size="3"><strong>In this notebook you will find an explanation of what is an autoencoder, how it works, and see an implementation of an autoencoder in TensorFlow.</strong></font>
# <br>
# <br>
# <h2>Table of Contents</h2>
# <ol>
# <li><a href="#ref1">Introduction</a></li>
# <li><a href="#ref2">Feature Extraction and Dimensionality Reduction</a></li>
# <li><a href="#ref3">Autoencoder Structure</a></li>
# <li><a href="#ref4">Performance</a></li>
# <li><a href="#ref5">Training: Loss Function</a></li>
# <li><a href="#ref6">Code</a></li>
# </ol>
# </div>
# <br>
# By the end of this notebook, you should be able to create simple autoencoders and how to apply them to problems that involves unsupervised learning.
# <br>
# <p></p>
# <hr>
# <a id="ref1"></a>
# <h2>Introduction</h2>
# An autoencoder, also known as autoassociator or Diabolo networks, is an artificial neural network employed to recreate the given input.
# It takes a set of <b>unlabeled</b> inputs, encodes them and then tries to extract the most valuable information from them.
# They are used for feature extraction, learning generative models of data, dimensionality reduction and can be used for compression.
#
# A 2006 paper named <b><a href="https://www.cs.toronto.edu/~hinton/science.pdf">Reducing the Dimensionality of Data with Neural Networks</a>, done by <NAME> and <NAME></b>, showed better results than years of refining other types of network, and was a breakthrough in the field of Neural Networks, a field that was "stagnant" for 10 years.
#
# Now, autoencoders, based on Restricted Boltzmann Machines, are employed in some of the largest deep learning applications. They are the building blocks of Deep Belief Networks (DBN).
#
# <center><img src="https://ibm.box.com/shared/static/xlkv9v7xzxhjww681dq3h1pydxcm4ktp.png" style="width: 350px;"></center>
# <hr>
# <a id="ref2"></a>
# <h2>Feature Extraction and Dimensionality Reduction</h2>
#
# An example given by <NAME> in KdNuggets (<a href="http://www.kdnuggets.com/2015/03/deep-learning-curse-dimensionality-autoencoders.html">link</a>) which gave an excellent explanation of the utility of this type of Neural Network.
#
# Say that you want to extract what emotion the person in a photography is feeling. Using the following 256x256 pixel grayscale picture as an example:
#
# <img src="https://ibm.box.com/shared/static/r5knpow4bk2farlvxia71e9jp2f2u126.png">
#
# But when use this picture we start running into a bottleneck! Because this image being 256x256 pixels in size correspond with an input vector of 65536 dimensions! If we used an image produced with conventional cellphone cameras, that generates images of 4000 x 3000 pixels, we would have 12 million dimensions to analyze.
#
#
# This bottleneck is further problematized as the difficulty of a machine learning problem is increased as more dimensions are involved. According to a 1982 study by <NAME> (<a href="http://www-personal.umich.edu/~jizhu/jizhu/wuke/Stone-AoS82.pdf">link</a>), the time to fit a model, is optimal if:
#
# <br><br>
# <div class="alert alert-block alert-info" style="margin-top: 20px">
# <h3><strong>$$m^{-p/(2p+d)}$$</strong></h3>
# <br>
# Where:
# <br>
# m: Number of data points
# <br>
# d: Dimensionality of the data
# <br>
# p: Parameter that depends on the model
# </div>
#
# As you can see, it increases exponentially!
# Returning to our example, we don't need to use all of the 65,536 dimensions to classify an emotion. A human identify emotions according to some specific facial expression, some <b>key features</b>, like the shape of the mouth and eyebrows.
#
# <center><img src="https://ibm.box.com/shared/static/m8urvuqujkt2vt1ru1fnslzh24pv7hn4.png" height="256" width="256"></center>
# <hr>
# <a id="ref3"></a>
# <h2>Autoencoder Structure</h2>
#
# <img src="https://ibm.box.com/shared/static/no7omt2jhqvv7uuls7ihnzikyl9ysnfp.png" style="width: 400px;">
#
# An autoencoder can be divided in two parts, the <b>encoder</b> and the <b>decoder</b>.
#
# The encoder needs to compress the representation of an input. In this case we are going to reduce the dimension the face of our actor, from 2000 dimensions to only 30 dimensions, by running the data through layers of our encoder.
#
# The decoder works like encoder network in reverse. It works to recreate the input, as closely as possible. This plays an important role during training, because it forces the autoencoder to select the most important features in the compressed representation.
#
# <hr>
# <a id="ref4"></a>
# <h2>Performance</h2>
#
# After the training has been done, you can use the encoded data as a reliable dimensionally-reduced data, applying it to any problems where dimensionality reduction seems appropriate.
#
# <img src="https://ibm.box.com/shared/static/yt3xyon4g2jyw1w9qup1mvx7cgh28l64.png">
#
# This image was extracted from the <NAME> and <NAME>'s <a href="https://www.cs.toronto.edu/~hinton/science.pdf">paper</a>, on the two-dimensional reduction for 500 digits of the MNIST, with PCA on the left and autoencoder on the right. We can see that the autoencoder provided us with a better separation of data.
# <hr>
# <a id="ref5"></a>
# <h2>Training: Loss function</h2>
#
# An autoencoder uses the Loss function to properly train the network. The Loss function will calculate the differences between our output and the expected results. After that, we can minimize this error with gradient descent. There are more than one type of Loss function, it depends on the type of data.
# <h3>Binary Values:</h3>
# $$l(f(x)) = - \sum_{k} (x_k log(\hat{x}_k) + (1 - x_k) \log (1 - \hat{x}_k) \ )$$
# For binary values, we can use an equation based on the sum of Bernoulli's cross-entropy.
#
# $x_k$ is one of our inputs and $\hat{x}_k$ is the respective output.
#
# We use this function so that if $x_k$ equals to one, we want to push $\hat{x}_k$ as close as possible to one. The same if $x_k$ equals to zero.
#
# If the value is one, we just need to calculate the first part of the formula, that is, $- x_k log(\hat{x}_k)$. Which, turns out to just calculate $- log(\hat{x}_k)$.
#
# And if the value is zero, we need to calculate just the second part, $(1 - x_k) \log (1 - \hat{x}_k) \ )$ - which turns out to be $log (1 - \hat{x}_k) $.
#
#
# <h3>Real values:</h3>
# $$l(f(x)) = - \frac{1}{2}\sum_{k} (\hat{x}_k- x_k \ )^2$$
# As the above function would behave badly with inputs that are not 0 or 1, we can use the sum of squared differences for our Loss function. If you use this loss function, it's necessary that you use a linear activation function for the output layer.
#
# As it was with the above example, $x_k$ is one of our inputs and $\hat{x}_k$ is the respective output, and we want to make our output as similar as possible to our input.
# <h3>Loss Gradient:</h3>
#
# $$\nabla_{\hat{a}(x^{(t)})} \ l( \ f(x^{(t)})) = \hat{x}^{(t)} - x^{(t)} $$
# We use the gradient descent to reach the local minimum of our function $l( \ f(x^{(t)})$, taking steps towards the negative of the gradient of the function in the current point.
#
# Our function about the gradient $(\nabla_{\hat{a}(x^{(t)})})$ of the loss of $l( \ f(x^{(t)})$ in the preactivation of the output layer.
#
# It's actually a simple formula, it is done by calculating the difference between our output $\hat{x}^{(t)}$ and our input $x^{(t)}$.
#
# Then our network backpropagates our gradient $\nabla_{\hat{a}(x^{(t)})} \ l( \ f(x^{(t)}))$ through the network using <b>backpropagation</b>.
# <hr>
# <a id="ref6"></a>
# <h2>Code</h2>
#
# For this part, we walk through a lot of Python 2.7.11 code. We are going to use the MNIST dataset for our example.
# The following code was created by <NAME>. You can find some of his code in <a href="https://github.com/aymericdamien">here</a>. We made some modifications for us to import the datasets to Jupyter Notebooks.
# Let's call our imports and make the MNIST data available to use.
# +
#from __future__ import division, print_function, absolute_import
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# Import MINST data
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("/tmp/data/", one_hot=True)
# -
# Now, let's give the parameters that are going to be used by our NN.
# +
learning_rate = 0.01
training_epochs = 20
batch_size = 256
display_step = 1
examples_to_show = 10
# Network Parameters
n_hidden_1 = 256 # 1st layer num features
n_hidden_2 = 128 # 2nd layer num features
n_input = 784 # MNIST data input (img shape: 28*28)
# tf Graph input (only pictures)
X = tf.placeholder("float", [None, n_input])
weights = {
'encoder_h1': tf.Variable(tf.random_normal([n_input, n_hidden_1])),
'encoder_h2': tf.Variable(tf.random_normal([n_hidden_1, n_hidden_2])),
'decoder_h1': tf.Variable(tf.random_normal([n_hidden_2, n_hidden_1])),
'decoder_h2': tf.Variable(tf.random_normal([n_hidden_1, n_input])),
}
biases = {
'encoder_b1': tf.Variable(tf.random_normal([n_hidden_1])),
'encoder_b2': tf.Variable(tf.random_normal([n_hidden_2])),
'decoder_b1': tf.Variable(tf.random_normal([n_hidden_1])),
'decoder_b2': tf.Variable(tf.random_normal([n_input])),
}
# -
# Now we need to create our encoder. For this, we are going to use sigmoidal functions. Sigmoidal functions delivers great results with this type of network. This is due to having a good derivative that is well-suited to backpropagation. We can create our encoder using the sigmoidal function like this:
# Building the encoder
def encoder(x):
# Encoder first layer with sigmoid activation #1
layer_1 = tf.nn.sigmoid(tf.add(tf.matmul(x, weights['encoder_h1']), biases['encoder_b1']))
# Encoder second layer with sigmoid activation #2
layer_2 = tf.nn.sigmoid(tf.add(tf.matmul(layer_1, weights['encoder_h2']), biases['encoder_b2']))
return layer_2
# And the decoder:
#
# You can see that the layer_1 in the encoder is the layer_2 in the decoder and vice-versa.
# Building the decoder
def decoder(x):
# Decoder first layer with sigmoid activation #1
layer_1 = tf.nn.sigmoid(tf.add(tf.matmul(x, weights['decoder_h1']),biases['decoder_b1']))
# Decoder second layer with sigmoid activation #2
layer_2 = tf.nn.sigmoid(tf.add(tf.matmul(layer_1, weights['decoder_h2']), biases['decoder_b2']))
return layer_2
# Let's construct our model.
# In the variable <code>cost</code> we have the loss function and in the <code>optimizer</code> variable we have our gradient used for backpropagation.
# +
# Construct model
encoder_op = encoder(X)
decoder_op = decoder(encoder_op)
# Reconstructed Images
y_pred = decoder_op
# Targets (Labels) are the input data.
y_true = X
# Define loss and optimizer, minimize the squared error
cost = tf.reduce_mean(tf.pow(y_true - y_pred, 2))
optimizer = tf.train.RMSPropOptimizer(learning_rate).minimize(cost)
# Initializing the variables
init = tf.global_variables_initializer()
# -
# For training we will run for 20 epochs.
# +
# Launch the graph
# Using InteractiveSession (more convenient while using Notebooks)
sess = tf.InteractiveSession()
sess.run(init)
total_batch = int(mnist.train.num_examples / batch_size)
# Training cycle
for epoch in range(training_epochs):
# Loop over all batches
for i in range(total_batch):
batch_xs, batch_ys = mnist.train.next_batch(batch_size)
# Run optimization op (backprop) and cost op (to get loss value)
_, c = sess.run([optimizer, cost], feed_dict={X: batch_xs})
# Display logs per epoch step
if epoch % display_step == 0:
print("Epoch:", '%04d' % (epoch+1),
"cost=", "{:.9f}".format(c))
print("Optimization Finished!")
# -
# Now, let's apply encoder and decoder for our tests.
# Applying encode and decode over test set
encode_decode = sess.run(
y_pred, feed_dict={X: mnist.test.images[:examples_to_show]})
# Let's simply visualize our graphs!
# Compare original images with their reconstructions
f, a = plt.subplots(2, 10, figsize=(10, 2))
for i in range(examples_to_show):
a[0][i].imshow(np.reshape(mnist.test.images[i], (28, 28)))
a[1][i].imshow(np.reshape(encode_decode[i], (28, 28)))
# As you can see, the reconstructions were successful. It can be seen that some noise were added to the image.
# <hr>
# ## Want to learn more?
#
# Running deep learning programs usually needs a high performance platform. __PowerAI__ speeds up deep learning and AI. Built on IBM’s Power Systems, __PowerAI__ is a scalable software platform that accelerates deep learning and AI with blazing performance for individual users or enterprises. The __PowerAI__ platform supports popular machine learning libraries and dependencies including TensorFlow, Caffe, Torch, and Theano. You can use [PowerAI on IMB Cloud](https://cocl.us/ML0120EN_PAI).
#
# Also, you can use __Watson Studio__ to run these notebooks faster with bigger datasets.__Watson Studio__ is IBM’s leading cloud solution for data scientists, built by data scientists. With Jupyter notebooks, RStudio, Apache Spark and popular libraries pre-packaged in the cloud, __Watson Studio__ enables data scientists to collaborate on their projects without having to install anything. Join the fast-growing community of __Watson Studio__ users today with a free account at [Watson Studio](https://cocl.us/ML0120EN_DSX).This is the end of this lesson. Thank you for reading this notebook, and good luck on your studies.
# ### Thanks for completing this lesson!
# Created by <a href="https://www.linkedin.com/in/franciscomagioli"><NAME></a>, <a href="https://ca.linkedin.com/in/erich-natsubori-sato"><NAME></a>, <a href="https://ca.linkedin.com/in/saeedaghabozorgi"><NAME>i</a>
# ### References:
# - https://en.wikipedia.org/wiki/Autoencoder
# - http://ufldl.stanford.edu/tutorial/unsupervised/Autoencoders/
# - http://www.slideshare.net/billlangjun/simple-introduction-to-autoencoder
# - http://www.slideshare.net/danieljohnlewis/piotr-mirowski-review-autoencoders-deep-learning-ciuuk14
# - https://cs.stanford.edu/~quocle/tutorial2.pdf
# - https://gist.github.com/hussius/1534135a419bb0b957b9
# - http://www.deeplearningbook.org/contents/autoencoders.html
# - http://www.kdnuggets.com/2015/03/deep-learning-curse-dimensionality-autoencoders.html/
# - https://www.youtube.com/watch?v=xTU79Zs4XKY
# - http://www-personal.umich.edu/~jizhu/jizhu/wuke/Stone-AoS82.pdf
# <hr>
#
# Copyright © 2018 [Cognitive Class](https://cocl.us/DX0108EN_CC). This notebook and its source code are released under the terms of the [MIT License](https://bigdatauniversity.com/mit-license/).
| IBM_AI/5_TensorFlow/ML0120EN-5.1-Review-Autoencoders.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:web]
# language: python
# name: conda-env-web-py
# ---
# # Chapter 2. Getting Started
#
# ## 2.1 Installation steps
#
# ### 2.1.1 Install Python and pip
#
# (1) Ubuntu has Python pre-installed.
#
# (2) Starting with Python 3.4, `pip` comes pre-installed.
#
# ### 2.1.2 Install virtualenv and virtualenvwrapper
#
# ```bash
# $ sudo pip install virtualenv
# $ sudo pip install virtualenvwrapper
# ```
#
# Add the following lines in `~/.bashrc`.
#
# ```
# # virtualenv
# export WORKON_HOME=~/.virtualenvs
# # Work around for https://stackoverflow.com/questions/33216679/usr-bin-python3-error-while-finding-spec-for-virtualenvwrapper-hook-loader
# export VIRTUALENVWRAPPER_PYTHON=/usr/bin/python
# source /usr/local/bin/virtualenvwrapper.sh
# ```
#
# ### 2.1.3 Create project environment
#
# ```bash
# $ mkvirtualenv HelloWorld
# $ setvirtualenvproject
# ```
#
# or
#
# ```bash
# $ mkproject HelloWorld
# ```
#
# To enter a virtual environment,
#
# ```bash
# $ workon HelloWorld
# ```
#
# To leave a virtual environment,
#
# ```bash
# $ deactivate
# ```
#
# ### 2.1.4 Install Flask and create the first project
#
# ```bash
# $ pip install flask
# ```
#
# ## 2.2 Creating our first Flask project
# +
# SAVE AS helloworld.py
from flask import Flask
# Get the flask application object
app = Flask(__name__)
# A view function
@app.route('/index')
def index():
return 'Hello World!'
if __name__ == '__main__':
app.run()
# -
# ## 2.3 Flask routing
#
# ### 2.3.1 Routing path
#
# http://localhost:5000/index ==> Flask ==> `@app.route('/index')` ==> `def index()`
#
# ### 2.3.2 Check the mapping rule
#
# Besides our rule of mapping "index", there is always a static rule.
from helloworld import app
app.url_map
# ## 2.4 Model, template, view
#
# ### 2.4.1 Model
#
# * Holds data
# * Usually represents rows in a database table
# * Flask leaves it to you
# * sqlite3, SQLAlchemy
#
# ### 2.4.2 Template (called "View" in MVC)
#
# * Used to generate HTML
# * Flask includes Jinja2
#
# ### 2.4.3 View (called "Controller" in MVC)
#
# * A function that generates a HTTP response for a HTTP request
# * Mapped to one or more URLs
#
# ### 2.4.4 Flask request handling
#
# request ==> http://localhost:5000/index ==> Flask ==> @app.route('/index') ==> "View": def index() ==> "Model": data from database; and "Template": html ==> response
| python/pluralsight-intro2flask/ch02-GettingStarted.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mapping out our data
#
# We have some data that is geo-aware. This means that we can place it on a map.
#
# To run this notebook you will need `geopandas`, `contextily`, `mapclassify` in addition to the fairly standard `pandas` and `matplotlib.pyplot`. The resulting plots have been saved, and are available to view directly.
import pandas as pd
import geopandas as gpd
import matplotlib.pyplot as plt
import contextily as ctx
# Load our data:
all_coords = pd.read_csv('s3://geotermaldata/S3FluidInclusionGasAnalysisData/COSO Field/COSO_wells_coord_gen.csv',
skipinitialspace=True,
delimiter=r'\s*,', # this removes some annoying spaces before the comma.
engine='python',
)
fname = './data/cleaned_types.csv'
df = pd.read_csv(fname)
df
# There are some columns that contain no data in the coordinates DataFrame, so we will drop those.
all_coords.dropna(axis='columns', inplace=True)
all_coords
all_coords.columns
gdf = gpd.GeoDataFrame(df)
# Although we have created `gdf` as a GeoDataFrame it does not have a proper geometry column yet.
#
# To create this column we need to have each point as a geometry. The easiest approach is to use `.apply` to combine the `Long83` and `Lat83` columns into a proper `Point`. First we need to add the information from `all_coords` to `gdf`, which we can do with the `merge` method:
gdf = gdf.merge(all_coords, left_on='Well ID', right_on='WellNumber')
gdf
# Notice that we have more columns than we started with, which are the ones from `all_coords`. We have also lost some rows, which are those from our fluid inclusion data which do not have a matching well in the coordinates DataFrame.
# Now we can use `.apply` to make a valid shapely `Point` based on the `Long83` and `Lat83` columns. This will give us a `geometry` column which will get used when plotting.
# +
from shapely.geometry import Point
def make_point(row):
#print(row)
return Point(float(row['Long83']), float(row['Lat83']))
# -
gdf['geometry'] = gdf.apply(make_point, axis=1)
gdf = gdf.set_geometry('geometry', crs="EPSG:4326")
gdf
# We can also make a GeoDataFrame of all of the wells in much the same way. These have no fluid inclusion data associated with them though.
all_wells = gpd.GeoDataFrame(all_coords) # turn the wells into a GeoDataFrame
all_wells['geometry'] = all_wells.apply(make_point, axis=1)
all_wells.set_crs(epsg=4326, inplace=True)
all_wells
# ## Plotting Things!
#
# Our wells with fluid inclusions are shown in orange, with the remainder in blue.
fig, ax = plt.subplots()
all_wells.plot(ax=ax)
gdf.plot(ax=ax)
# Since we are plotting maps, we need to have a single value per point to plot. We can use the maximum depth as something useful for now.
# +
wells = gpd.GeoDataFrame()
for group in gdf.groupby('Well ID'):
wells = wells.append(group[1].loc[group[1]['Depth (ft)'] == group[1]['Depth (ft)'].max()])
wells
# -
# Notice that this has some wells that are duplicated, because there are more than one in the `all_coords` DataFrame (one is active and one is inactive. We can either ignore these or drop them later. I am just ignoring this for now, because the data is the same.
for idx, well in enumerate(wells['Well ID']):
try:
if wells.iloc[idx-1]['Well ID'] == well:
print(f'{idx-1} and {idx} are both well {well}.')
except IndexError:
continue
# Adding some basemaps to our plots.
# +
tiles = ctx.providers.Stamen.Terrain
image = ctx.providers.Esri.WorldImagery
fig, ax = plt.subplots(figsize=(12,12))
all_wells.plot(ax=ax, c='white', ec='k', markersize=40)
wells.plot(ax=ax, ec='k', markersize=50, column='WellType', legend=True)
ctx.add_basemap(ax=ax, source=tiles, crs=4326, attribution='')
ctx.add_basemap(ax=ax, source=image, crs=4326, alpha=0.6, attribution='')
ctx.add_attribution(ax=ax, text=f'{tiles.attribution}\n{image.attribution}')
# Add labels for our wells of interest.
for well, lat, lon in zip(wells['Well ID'], wells['Lat83'], wells['Long83']):
geom = (float(lon)+0.001, float(lat)-0.001)
ax.annotate(well, geom)
plt.tight_layout()
plt.savefig('./img/all_wells.png', dpi=300)
# +
fig, ax = plt.subplots(figsize=(12,12))
wells.plot(ax=ax, ec='k', markersize=50, column='WellType', legend=True)
ctx.add_basemap(ax=ax, source=tiles, crs=4326, attribution='')
ctx.add_basemap(ax=ax, source=image, crs=4326, alpha=0.6, attribution='')
ctx.add_attribution(ax=ax, text=f'{tiles.attribution}\n{image.attribution}')
for well, lat, lon in zip(wells['Well ID'], wells['Lat83'], wells['Long83']):
geom = (float(lon)+0.001, float(lat)-0.001)
ax.annotate(well, geom)
plt.tight_layout()
plt.savefig('./img/fi_wells.png', dpi=300)
# -
# ### Plotting maximum depths
#
# The depths are in feet, so I am converting them to metres.
wells['Depth (ft)'].describe()
wells['Depth_m'] = wells['Depth (ft)'] * 0.3048
wells['Depth_m']
# Now we can plot them easily:
# +
fig, ax = plt.subplots(figsize=(12,12))
wells.plot(ax=ax, ec='k', markersize=50,
column='Depth_m',
scheme='percentiles',
legend=True,
cmap=plt.cm.plasma,
)
ctx.add_basemap(ax=ax, source=tiles, crs=4326, attribution='')
ctx.add_basemap(ax=ax, source=image, crs=4326, alpha=0.6, attribution='')
ctx.add_attribution(ax=ax, text=f'{tiles.attribution}\n{image.attribution}')
for well, lat, lon in zip(wells['Well ID'], wells['Lat83'], wells['Long83']):
geom = (float(lon)+0.001, float(lat)-0.001)
#print(geom)
ax.annotate(well, geom)
plt.tight_layout()
plt.savefig('./img/fi_wells_depth.png', dpi=300)
# -
gdf.to_file('./data/fi_wells.gpkg', driver='GPKG')
all_wells.to_file('./data/well_locations.gpkg', driver='GPKG')
| mapping_the_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import curve_fit
# %matplotlib notebook
# +
k=0.98 #N/m
mu0 = 4*np.pi*1e-7
def B2coil_der(z,I):
return ((mu0*I*N*R**2)/2)*((-(3/2)*((2*z-R)/((z-(R/2)**2+R**2)**(5.0/2))))+(-(3/2)*((2*z-R)/((z+(R/2)**2+R**2)**(5.0/2)))))
def mu(z,s):
return (4*np.pi*k*s*z**3)/(2*mu0)
# +
r = 0.07
I = 3
N = 168
mnofp = 10**(-7)
z = np.asarray([0.008,0.018,0.028,0.038,0.048,0.058,0.068,0.078,0.088,0.098,0.108,0.118])
#nz = np.asarray([0.038,0.044,0.048,0.055,0.060,0.069,0.079,0.093,0.106])
displacement = np.array([0.005,0.008,0.01,0.012,0.013,0.014, 0.015,0.017,0.018])
amp = np.array([0.5,0.75,1,1.25,1.5,1.75,2,2.25,2.5])
for i in range(len(z)):
z[i] = z[i] - 0.035
#nz[i] = nz[i] - 0.035
B = mnofp*(2*np.pi*(r**2)*I*N)/(((r**2)+(z**2))**(3/2))
print(B)
# +
plt.figure()
plt.scatter(z,B)
def line(x,m,n,b):
return m*(x**n)+b
def line2(x,m,b):
return m*x+b
linefit,lcov= np.polyfit(z,B,4,cov=True)
exfit = np.polyfit(z,B,4)
y = np.poly1d(linefit)
xfit = np.linspace(-0.03,0.1,1000)
y2=y(xfit)
error = np.diagonal(lcov)
error = np.sqrt(error)
print(error)
plt.plot(xfit,y2,color='red')
plt.ylim(0.0001,0.00535)
plt.xlabel('Axial Displacement from Center (m)')
plt.ylabel('Magnetic Field Strength (T)')
plt.title('Field Strength vs. Axial Displacement')
plt.plot(xfit,line(xfit,*exfit),color='green',linestyle='--')
# +
np.mean(displacement)
#print(amp)
#plt.scatter(displacement, amp)
# +
fig,ax = plt.subplots()
moment_fit,mcov = curve_fit(line2,amp,displacement,p0=[0.0075,0])
print(moment_fit)
xmom = np.linspace(0.25,2.75)
ymom = line2(xmom,*moment_fit)
ax.scatter(amp,displacement)
ax.plot(xmom,ymom,color='orange')
plt.minorticks_on()
ax.set_ylim(0.003,0.022)
ax.set_xlabel('I [A]')
ax.set_ylabel(r'$\Delta$x [m]')
# -
R = 0.07
mag_moment = (-k*(displacement))/B2coil_der((0.07-displacement),amp)
mag_moment
print(np.mean(mag_moment))
np.mean(((displacement**3)*0.0005*3))
np.mean(1/((2*k*(0.07-displacement)**3)/(0.07**2*mu0)))
B
| physics/em/CompLab/MAGNETOFORCEEEE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.11 64-bit (''ml'': conda)'
# language: python
# name: python3
# ---
# # Backpropagation on DAGs
# 
# In this notebook, we will look at **backpropagation** (BP) on **directed acyclic computational graphs** (DAG). Our main result is that a single training step for a single data point (consisting of both a forward and a backward pass) has a time complexity that is linear in the number of edges of the network. In the last section, we take a closer look at the implementation of `.backward` in PyTorch.
#
# **Readings**
# * [Evaluating $\nabla f(x)$ is as fast as $f(x)$](https://timvieira.github.io/blog/post/2016/09/25/evaluating-fx-is-as-fast-as-fx/)
# * [Back-propagation, an introduction](http://www.offconvex.org/2016/12/20/backprop/)
# ## Gradient descent on the loss surface
# ```{margin}
# **Constructing the loss surface**. The loss function $\ell$ acts as an almost-everywhere differentiable surrogate to the true objective. The empirical loss surface will generally vary for different samples drawn. But we except these surfaces to be very similar, assuming the samples are drawn from the same distribution.
# ```
#
# For every data point $(\mathbf x, y)$, the loss function $\ell$ assigns a nonnegative number $\ell(y, f_{\mathbf w}(\mathbf x))$ that approaches zero whenever the predictions $f_{\mathbf w}(\mathbf x)$ approach the target values $y$. Given the current parameters $\mathbf w \in \mathbb R^d$ of a neural network $f$, we can imagine the network to be at a certain point $(\mathbf w, \mathcal L_{\mathcal X}(\mathbf w))$ on a surface in $\mathbb R^d \times \mathbb R$ where $\mathcal L_{\mathcal X}(\mathbf w)$ is the average loss over the dataset:
#
# $$
# \mathcal L_{\mathcal X}(\mathbf w) = \frac{1}{|\mathcal X|} \sum_{(\mathbf x, y) \in \mathcal X} \ell(y, f_{\mathbf w}(\mathbf x)).
# $$
#
# So training a neural network is equivalent to finding the minimum of this surface. In practice, we use variants of gradient descent, characterized by the update rule $\mathbf w \leftarrow \mathbf w - \varepsilon \nabla_{\mathbf w} \mathcal L_{\mathcal X}$, to find a local minimum. Here $-\nabla_{\mathbf w} \mathcal L_{\mathcal X}$ is the direction of steepest descent at $\mathbf w$ and the learning rate $\varepsilon > 0$ is a constant that controls the step size.
#
#
# ```{figure} ../../img/loss_surface_resnet.png
# ---
# name: loss-surface-resnet
# width: 35em
# ---
# Much of deep learning research is dedicated to studying the geometry of loss surfaces and its effect on optimization. **Source**: Visualizing the Loss Landscape of Neural Nets
# [[arxiv.org/abs/1712.09913](https://arxiv.org/abs/1712.09913)]
# ```
#
#
# ```{margin}
# **Derivatives of comp. graphs**
# ```
#
# In principle, we can perturb the current state of the network (obtained during forward pass) by perturbing the network weights / parameters. This results in perturbations flowing up to the final loss node (assuming each computation is differentiable). So it's not a mystery that we can compute derivatives of computational graphs which may appear, at first glance, as "discrete" objects. Another perspective is that a computational DAG essentially models a a sequence of function compositions which can be easily differentiated using chain rule. However, looking at the network structure allows us to easily code the computation into a computer, exploit modularity, and efficiently compute the flow of derivatives at each layer. This is further discussed below.
#
#
# ```{margin}
# **The need for efficient BP**
# ```
#
# Observe that $\nabla_\mathbf w \mathcal L_{\mathcal X}$ consists of partial derivatives for each weight in the network. This can easily number in millions. So this backward pass operation can be huge. To compute these values efficiently, we will perform both forward and backward passes in a dynamic programming fashion to avoid recomputing any known value. As an aside, this improvement in time complexity turns out to be insufficient for pratical uses, and is supplemented with sophisticated hardware for parallel computation (GPUs / TPUs) which can reduce training time by some factor, e.g. from days to hours.
# ## Backpropagation on Computational Graphs
# ```{margin}
# **Forward pass**
# ```
#
# A neural network can be modelled as a **directed acyclic graph** (DAG) of compute and parameter nodes that implements a function $f$ and can be extended to implement the calculation of the loss value for each training example and parameter values. In computing $f(\mathbf x)$, the input $\mathbf x$ is passed to the first layer and propagated forward through the network, computing the output value of each node. Every value in the nodes is stored to preserve the current state for backward pass, as well as to avoid recomputation for the nodes in the next layer. Assuming a node with $n$ inputs require $n$ operations, then one forward pass takes $O(E)$ calculations were $E$ is the number of edges of the graph.
# ```{figure} ../../img/backprop-compgraph2.png
# ---
# width: 35em
# name: backprop-compgraph2
# ---
# Backpropagation through a single layer neural network with weights $w_0$ and $w_1$, and input-output pair $(x, y).$ Shown here is the gradient flowing from the loss node $\mathcal L$ to the weight $w_0.$ [[source]](https://drive.google.com/file/d/1JCWTApGieKZmFW4RjCANZM8J6igcsdYg/view)
# ```
# ```{margin}
# **Backward pass**
# ```
#
# During backward pass, we divide gradients into two groups: **local gradients** ${\frac{\partial{\mathcal u}}{\partial w}}$ between connected nodes $u$ and $w,$ and **backpropagated gradients** ${\frac{\partial{\mathcal L}}{\partial u}}$ for each node ${u}.$ Our goal is to calculate the backpropagated gradient of the loss with respect to parameter nodes. Note that parameter nodes have zero fan-in ({numref}`backprop-compgraph2`). BP proceeds inductively. First, $\frac{\partial{\mathcal L}}{\partial \mathcal L} = 1$ is stored as gradient of the node which computes the loss value. If the backpropagated gradient ${\frac{\partial{\mathcal L}}{\partial u}}$ is stored for each compute node $u$ in the upper layer, then after computing local gradients ${\frac{\partial{u}}{\partial w}}$, the backpropagated gradient ${\frac{\partial{\mathcal L}}{\partial w}}$ for compute node $w$ can be calculated via the chain rule:
#
# ```{math}
# :label: backprop
# {\frac{\partial\mathcal L}{\partial w} } = \sum_{ {u} }\left( {{\frac{\partial\mathcal L}{\partial u}}} \right)\left( {{\frac{\partial{u}}{\partial w}}} \right).
# ```
# ```{margin}
# BP is a useful tool for understanding how derivatives flow through a model. This can be extremely helpful in reasoning about why some models are difficult to optimize. Classic examples are vanishing or exploding gradients as we go into deeper layers of the network.
# ```
#
# Thus, continuing the "flow" of gradients to the current layer. The process ends on nodes with zero fan-in. Note that the partial derivatives are evaluated on the current network state — these values are stored during forward pass which precedes backward pass. Analogously, all backpropagated gradients are stored in each compute node for use by the next layer. On the other hand, there is no need to store local gradients; these are computed as needed. Hence, it suffices to compute all gradients with respect to compute nodes to get all gradients with respect to the weights of the network.
#
#
# ```{figure} ../../img/backprop-compgraph.png
# ---
# width: 30em
# name: backprop-compgraph
# ---
# BP on a generic comp. graph with fan out > 1 on node <code>y</code>. Each backpropagated gradient computation is stored in the corresponding node. For node <code>y</code> to calculate the backpropagated gradient we have to sum over the two incoming gradients which can be implemented using matrix multiplication of the gradient vectors. [[source]](https://drive.google.com/file/d/1JCWTApGieKZmFW4RjCANZM8J6igcsdYg/view)
# ```
#
# **Backpropagation algorithm.** Now that we know how to compute each backpropagated gradient implemented as `u.backward()` for node `u` which sends its gradient $\frac{\partial \mathcal L}{\partial u}$ to all its parent nodes (nodes on the lower layer connected to `u`). The complete recursive algorithm with SGD is implemented below. Note that this abstracts away autodifferentiation.
#
# ```python
# def Forward():
# for c in compute:
# c.forward()
#
# def Backward(loss):
# for c in compute: c.grad = 0
# for c in params: c.grad = 0
# for c in inputs: c.grad = 0
# loss.grad = 1
#
# for c in compute[::-1]:
# c.backward()
#
# def SGD(eta):
# for w in params:
# w.value -= eta * w.grad
# ```
# <br>
# Two important properties of the algorithm which makes it the practical choice for training huge neural networks are as follows:
# * **Modularity.** The dependence only on nodes belonging to the upper layer suggests a modularity in the computation, e.g. we can connect DAG subnetworks with possibly distinct network architectures by only connecting nodes that are exposed between layers.
#
# <br>
#
# * **Efficiency.** From the backpropagation equation {eq}`backprop`, the backpropagated gradient $\frac{\partial \mathcal L}{\partial w}$ for node $w$ is computed by a sum that is indexed by $u$ for every node connected to $w$. Iterating over all nodes $w$ in the network, we cover all the edges in the network with no edge counted twice. Assuming computing local gradients take constant time, then backward pass requires $O(E)$ computations.
# $\phantom{3}$
# ```{admonition} BP equations for MLPs
#
# Consider an MLP which can be modelled as a computational DAG with edges between preactivation and activation values, as well as edges from weights and input values that fan into preactivations ({numref}`backprop-compgraph2`). Let ${z_j}^{[t]} = \sum_k {w_{jk}}^{[t]}{a_k}^{[t-1]}$ and ${\mathbf a}^{[t]} = \phi^{[t]}({\mathbf z}^{[t]})$ be the values of compute nodes at the $t$-th layer of the network. The backpropagated gradients for the compute nodes of the current layer are given by
#
# $$\begin{aligned}
# \dfrac{\partial \mathcal L}{\partial {a_j}^{[t]}}
# &= \sum_{k}\dfrac{\partial \mathcal L}{\partial {z_k}^{[t+1]}} \dfrac{\partial {z_k}^{[t+1]}}{\partial {a_j}^{[t]}} = \sum_{k}\dfrac{\partial \mathcal L}{\partial {z_k}^{[t+1]}} {w_{kj}}^{[t+1]}
# \end{aligned}$$
#
# and
#
# $$\begin{aligned}
# \dfrac{\partial \mathcal L}{\partial {z_j}^{[t]}}
# &= \sum_{l}\dfrac{\partial \mathcal L}{\partial {a_l}^{[t]}} \dfrac{\partial {a_l}^{[t]}}{\partial {z_j}^{[t]}}.
# \end{aligned}$$
#
# This sum typically reduces to a single term for activations such as ReLU, but not for activations which depend on multiple preactivations such as softmax. Similarly, the backpropagated gradients for the parameter nodes (weights and biases) are given by
#
# $$\begin{aligned}
# \dfrac{\partial \mathcal L}{\partial {w_{jk}}^{[t]}}
# &= \dfrac{\partial \mathcal L}{\partial {z_j}^{[t]}} \dfrac{\partial {z_j}^{[t]}}{\partial {w_{jk}}^{[t]}} = \dfrac{\partial \mathcal L}{\partial {z_j}^{[t]}} {a^{[t-1]}_k} \\
# \text{and}\qquad\dfrac{\partial \mathcal L}{\partial {b_{j}}^{[t]}}
# &= \dfrac{\partial \mathcal L}{\partial {z_j}^{[t]}} \dfrac{\partial {z_j}^{[t]}}{\partial {b_{j}}^{[t]}} = \dfrac{\partial \mathcal L}{\partial {z_j}^{[t]}}.
# \end{aligned}$$
#
# Backpropagated gradients for compute nodes are stored until the weights are updated, e.g. $\frac{\partial \mathcal L}{\partial {z_k}^{[t+1]}}$ are retrieved in the compute nodes of the $t+1$-layer to compute gradients in the $t$-layer. On the other hand, the local gradients $\frac{\partial {a_k}^{[t]}}{\partial {z_j}^{[t]}}$ are computed directly using autodifferentiation and evaluated with the current network state obtained during forward pass.
# ```
# ## Autodifferentiation with PyTorch `autograd`
#
# The `autograd` package allows automatic differentiation by building computational graphs on the fly every time we pass data through our model. Autograd tracks which data combined through which operations to produce the output. This allows us to take derivatives over ordinary imperative code. This functionality is consistent with the memory and time requirements outlined in above for BP.
#
# <br>
#
# **Backward for scalars.** Let $y = \mathbf x^\top \mathbf x = \sum_i {x_i}^2.$ In this example, we initialize a tensor `x` which initially has no gradient. Calling backward on `y` results in gradients being stored on the leaf tensor `x`.
# +
x = torch.arange(4, dtype=torch.float, requires_grad=True)
y = x.T @ x
y.backward()
(x.grad == 2*x).all()
# -
# **Backward for vectors.** Let $\mathbf y = g(\mathbf x)$ and let $\mathbf v$ be a vector having the same length as $\mathbf y.$ Then `y.backward(v)` implements
#
# $$\sum_i v_i \left(\frac{\partial y_i}{\partial x_j}\right)$$
#
# resulting in a vector of same length as `x` that is stored in `x.grad`. Note that the terms on the right are the local gradients in backprop. Hence, if `v` contains backpropagated gradients of nodes that depend on `y`, then this operation gives us the backpropagated gradients with respect to `x`, i.e. setting $v_i = \frac{\partial \mathcal{L} }{\partial y_i}$ gives us the vector $\frac{\partial \mathcal{L} }{\partial x_j}.$
# +
x = torch.rand(size=(4,), dtype=torch.float, requires_grad=True)
v = torch.rand(size=(2,), dtype=torch.float)
y = x[:2]
# Computing the Jacobian by hand
J = torch.tensor(
[[1, 0, 0, 0],
[0, 1, 0, 0]], dtype=torch.float
)
# Confirming the above formula
y.backward(v)
(x.grad == v @ J).all()
# -
# **Locally disabling gradient tracking.** Disabling gradient computation is useful when computing values, e.g. accuracy, whose gradients will not be backpropagated into the network. To stop PyTorch from building computational graphs, we can put the code inside a `torch.no_grad()` context or inside a function with a `@torch.no_grad()` decorator.
#
# Another technique is to use the `.detach()` method which returns a new tensor detached from the current graph but shares the same storage with the original one. In-place modifications on either of them will be seen, and may trigger errors in correctness checks.
| docs/notebooks/fundamentals/backpropagation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from cpmpy import *
import numpy as np
# +
model = Model()
gr,bl,og,ye,gy = boolvar(shape=5)
model += (12*gr + 2*bl + 1*og + 4*ye + 1*gy <= 15)
model.maximize(4*gr + 2*bl + 1*og + 10*ye + 2*gy)
model.solve()
# -
print(gr.value(), bl.value(), og.value(), ye.value(), gy.value())
v = boolvar()
print(v)
v = boolvar(shape=5)
print(v)
v = intvar(1,9, shape=5)
print(v)
m = intvar(1,9, shape=(3,3))
print(m)
t = boolvar(shape=(2,3,4))
print(t)
# +
puzzle_start = np.array([
[0,3,6],
[2,4,8],
[1,7,5]])
(dim,dim2) = puzzle_start.shape
assert (dim == dim2), "puzzle needs square shape"
n = dim*dim2 - 1 # e.g. an 8-puzzle
# State of puzzle at every step
K = 20
x = intvar(0,n, shape=(K,dim,dim), name="x")
print(x)
# -
x,y,z = intvar(1,9, shape=3)
print( x + y )
print( x * y )
print( abs(x - y) )
a = intvar(1,9, shape=5, name="a")
print( sum(a) )
c = ( abs(sum(a) - (x+y)) == z )
print( c )
type(c)
c.name
c.args[1]
type(c.args[0])
c.args[0].name
x = boolvar(shape=(4,4), name="x")
print(x)
print(x[0,:])
print(x[:,0])
print(x[:,1:-1])
x = boolvar(shape=4, name="x")
print(x)
sel = np.array([True, False, True, False])
print(x[sel])
print(x[np.arange(4) % 2 == 0])
idx = [1,3]
print(x[idx])
x = intvar(1,9, shape=3, name="x")
y = intvar(1,9, shape=3, name="y")
print(x + y)
print(x == [1,2,3])
print(x == 1)
# +
import numpy as np
from cpmpy import *
e = 0 # value for empty cells
given = np.array([
[e, e, e, 2, e, 5, e, e, e],
[e, 9, e, e, e, e, 7, 3, e],
[e, e, 2, e, e, 9, e, 6, e],
[2, e, e, e, e, e, 4, e, 9],
[e, e, e, e, 7, e, e, e, e],
[6, e, 9, e, e, e, e, e, 1],
[e, 8, e, 4, e, e, 1, e, e],
[e, 6, 3, e, e, e, e, 8, e],
[e, e, e, 6, e, 8, e, e, e]])
# Variables
puzzle = intvar(1,9, shape=given.shape, name="puzzle")
# +
model = Model()
n = given.shape[0]
# Constraints on rows and columns
for i in range(n):
model += AllDifferent([puzzle[i,j] for j in range(n)])
model += AllDifferent([puzzle[j,i] for j in range(n)])
# Constraints on blocks
for i in range(0,9, 3):
for j in range(0,9, 3):
model += AllDifferent([puzzle[r,c]
for r in range(i,i+3)
for c in range(j,j+3)])
# Constraints on values (cells that are not empty)
for r in range(n):
for c in range(n):
if given[r,c] != e:
model += puzzle[r,c] == given[r,c]
model.solve()
# +
model = Model()
# Constraints on rows and columns
model += [AllDifferent(row) for row in puzzle]
model += [AllDifferent(col) for col in puzzle.T]
# Constraints on blocks
for i in range(0,9, 3):
for j in range(0,9, 3):
model += AllDifferent(puzzle[i:i+3, j:j+3])
# Constraints on values (cells that are not empty)
model += [puzzle[given!=e] == given[given!=e]]
model.solve()
# -
# +
jobs_data = cpm_array([ # (job, machine) = duration
[3,2,2], # job 0
[2,1,4], # job 1
[0,4,3], # job 2 (duration 0 = not used)
])
max_dur = sum(jobs_data.flat)
n_jobs, n_machines = jobs_data.shape
all_jobs = range(n_jobs)
all_machines = range(n_machines)
# Variables
start_time = intvar(0, max_dur, shape=(n_machines,n_jobs), name="start")
end_time = intvar(0, max_dur, shape=(n_machines,n_jobs), name="stop")
# -
from itertools import combinations
# +
model = Model()
# end = start + dur
for j in all_jobs:
for m in all_machines:
model += (end_time[m,j] == start_time[m,j] + jobs_data[j,m])
# Precedence constraint per job
for j in all_jobs:
for m1,m2 in combinations(all_machines,2): # [0,1,2]->[(0,1),(0,2),(1,2)]
model += (end_time[m1,j] <= start_time[m2,j])
# No overlap constraint: one starts before other one ends
for m in all_machines:
for j1,j2 in combinations(all_jobs, 2):
model += (start_time[m,j1] >= end_time[m,j2]) | \
(start_time[m,j2] >= end_time[m,j1])
# Objective: makespan
makespan = Maximum([end_time[m,j] for m in all_machines for j in all_jobs])
model.minimize(makespan)
model.solve()
# +
model = Model()
# end = start + dur
model += (end_time == start_time + jobs_data.T)
# Precedence constraint per job
for m1,m2 in combinations(all_machines,2):
model += (end_time[m1,:] <= start_time[m2,:])
# No overlap constraint: one starts before other one ends
for j1,j2 in combinations(all_jobs, 2):
model += (start_time[:,j1] >= end_time[:,j2]) | \
(start_time[:,j2] >= end_time[:,j1])
# Objective: makespan
makespan = max(end_time)
model.minimize(makespan)
model.solve()
# +
model = Model()
# FOR THE EXPERTS: NOT A CONSTRAINT!
end_time = start_time + jobs_data.T
# Precedence constraint per job
for m1,m2 in combinations(all_machines,2): # [(0,1), (0,2), (1,2)]
model += (end_time[m1,:] <= start_time[m2,:])
# No overlap constraint: one starts before other one ends
for j1,j2 in combinations(all_jobs, 2):
model += (start_time[:,j1] >= end_time[:,j2]) | \
(start_time[:,j2] >= end_time[:,j1])
# Objective: makespan, NOT A CP VARIABLE!
makespan = max(end_time)
model.minimize(makespan)
model.solve()
# +
print("Makespan:",makespan.value())
print("Schedule:")
grid = -8*np.ones((n_machines, makespan.value()), dtype=int)
for j in all_jobs:
for m in all_machines:
grid[m,start_time[m,j].value():end_time[m,j].value()] = j
print(grid)
# +
from PIL import Image, ImageDraw, ImageFont
# based on <NAME>'s https://github.com/Alexander-Schiendorfer/cp-examples
def visualize_scheduling(start, end):
nMachines, nJobs = start.shape
makespan = max(end.value())
# Draw solution
# Define start location image & unit sizes
start_x, start_y = 30, 40
pixel_unit = 50
pixel_task_height = 100
vert_pad = 10
imwidth, imheight = makespan * pixel_unit + 2 * start_x, start_y + start_x + nMachines * (pixel_task_height + vert_pad)
# Create new Image object
img = Image.new("RGB", (imwidth, imheight), (255, 255,255))
# Create rectangle image
img1 = ImageDraw.Draw(img)
# Get a font
try:
myFont = ImageFont.truetype("arialbd.ttf", 20)
except:
myFont = ImageFont.load_default()
# Draw makespan label
center_x, center_y = imwidth / 2, start_y / 2
msg = f"Makespan: {makespan}"
w, h = img1.textsize(msg, font=myFont)
img1.text((center_x - w / 2, center_y - h / 2), msg, fill="black", font=myFont)
task_cs = ["#4bacc6", "#f79646", "#9bbb59"]
task_border_cs = ["#357d91", "#b66d31", "#71893f"]
# Draw three rectangles for machines
machine_upper_lefts = []
for i in range(nMachines):
start_m_x, start_m_y = start_x, start_y + i * (pixel_task_height + vert_pad)
end_m_x, end_m_y = start_m_x + makespan * pixel_unit, start_m_y + pixel_task_height
machine_upper_lefts += [(start_m_x, start_m_y)]
shape = [(start_m_x, start_m_y), (end_m_x, end_m_y)]
img1.rectangle(shape, fill ="#d3d3d3")
# Draw tasks for each job
inner_sep = 5
for j in range(nJobs):
job_name = str(j)
for m in range(nMachines):
if start[m,j].value() == end[m,j].value():
continue # skip
start_m_x, start_m_y = machine_upper_lefts[m]
start_rect_x, start_rect_y = start_m_x + start[m,j].value() * pixel_unit, start_m_y + inner_sep
end_rect_x, end_rect_y = start_m_x + end[m,j].value() * pixel_unit, start_m_y + pixel_task_height - inner_sep
shape = [(start_rect_x, start_rect_y), (end_rect_x, end_rect_y)]
img1.rectangle(shape, fill=task_cs[j], outline=task_border_cs[j])
# Write a label for each task of each job
msg = f"{job_name}"
text_w, text_h = img1.textsize(msg, font=myFont)
center_x, center_y = (start_rect_x + end_rect_x) / 2, (start_rect_y + end_rect_y) / 2
img1.text((center_x - text_w / 2, center_y - text_h / 2), msg, fill="white", font=myFont)
img.show()
visualize_scheduling(start_time, end_time)
# -
# +
# '0' is empty spot
puzzle_start = np.array([
[3,7,5],
[1,6,4],
[8,2,0]])
puzzle_end = np.array([
[1,2,3],
[4,5,6],
[7,8,0]])
def n_puzzle(puzzle_start, puzzle_end, K):
print("Max steps:", K)
m = Model()
(dim,dim2) = puzzle_start.shape
assert (dim == dim2), "puzzle needs square shape"
n = dim*dim2 - 1 # e.g. an 8-puzzle
# State of puzzle at every step
x = intvar(0,n, shape=(K,dim,dim), name="x")
# Start state constraint
m += (x[0] == puzzle_start)
# End state constraint
m += (x[-1] == puzzle_end)
# define neighbors = allowed moves for the '0'
def neigh(i,j):
# same, left,right, down,up, if within bounds
for (rr, cc) in [(0,0),(-1,0),(1,0),(0,-1),(0,1)]:
if 0 <= i+rr and i+rr < dim and 0 <= j+cc and j+cc < dim:
yield (i+rr,j+cc)
# Transition: define next (t) based on prev (t-1) + invariants
for t in range(1, K):
# Invariant: in each step, all cells are different
m += AllDifferent(x[t])
# Invariant: only the '0' position can move
m += ((x[t-1] == x[t]) | (x[t-1] == 0) | (x[t] == 0))
# for each position, determine reachability of the '0' position
for i in range(dim):
for j in range(dim):
m += (x[t,i,j] == 0).implies(any(x[t-1,r,c] == 0 for r,c in neigh(i,j)))
return (m,x)
# -
(m,x) = n_puzzle(puzzle_start, puzzle_end, 10)
m.solve()
print(m.status())
(m,x) = n_puzzle(puzzle_start, puzzle_end, 100)
m.solve()
print(m.status())
# +
K0 = 5
step = 3
(m,x) = n_puzzle(puzzle_start, puzzle_end, K0)
while not m.solve():
print(m.status())
K0 = K0 + step
(m,x) = n_puzzle(puzzle_start, puzzle_end, K0)
print(m.status())
# -
(m,x) = n_puzzle(puzzle_start, puzzle_end, 14)
m.solve()
m.status()
# +
(m,x) = n_puzzle(puzzle_start, puzzle_end, 20)
m.solve()
base_runtime = m.status().runtime
print("Runtime with default params", base_runtime)
from cpmpy.solvers import CPM_ortools, param_combinations
all_params = {'cp_model_probing_level': [0,1,2,3],
'linearization_level': [0,1,2],
'symmetry_level': [0,1,2]}
configs = [] # (runtime, param)
for params in param_combinations(all_params):
s = CPM_ortools(m)
print("Running", params, end='\r')
s.solve(time_limit=base_runtime*1.05, **params) # timeout of 105% of base_runtime
configs.append( (s.status().runtime, params) )
best = sorted(configs)[0]
print("\nFastest in", round(best[0],4), "seconds, config:", best[1])
# -
(m,x) = n_puzzle(puzzle_start, puzzle_end, 100)
m = CPM_ortools(m)
m.solve(**best[1])
print(m.status())
| examples/tutorial/part1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # OpenEO Connection to EURAC Backend
import openeo
import logging
logging.basicConfig(level=logging.INFO)
# +
# Define constants
# Connection
EURAC_DRIVER_URL = "http://saocompute.eurac.edu/openEO_0_3_0/openeo"
OUTPUT_FILE = "/tmp/openeo-wcps.nc"
OUTFORMAT = "netcdf"
user = "group1"
password = "<PASSWORD>"
# Data
PRODUCT_ID = "S2_L2A_T32TPS_20M"
DATE_START = "2016-01-01"
DATE_END = "2016-03-10"
IMAGE_WEST = 652000
IMAGE_EAST = 672000
IMAGE_NORTH = 5161000
IMAGE_SOUTH = 5181000
IMAGE_SRS = "EPSG:32632"
# Processes
NDVI_RED = "B04"
NDVI_NIR = "B8A"
# -
# Connect with EURAC backend
connection = openeo.connect(EURAC_DRIVER_URL, auth_options={"username": user, "password": password})
connection
# Get available processes from the back end.
processes = connection.list_processes()
processes
# +
# Retrieve the list of available collections
collections = connection.list_collections()
list(collections)[:2]
# -
# Get detailed information about a collection
process = connection.describe_collection(PRODUCT_ID)
process
# +
# Select collection product
datacube = connection.imagecollection(PRODUCT_ID)
print(datacube.to_json())
# +
# Specifying the date range and the bounding box
datacube = datacube.filter_bbox(west=IMAGE_WEST, east=IMAGE_EAST, north=IMAGE_NORTH,
south=IMAGE_SOUTH, crs=IMAGE_SRS)
datacube = datacube.filter_daterange(extent=[DATE_START, DATE_END])
print(datacube.to_json())
# -
# Sending the job to the backend
job = datacube.create_job()
job.start_job()
job
# Describe Job
job.describe_job()
# +
# Download job result
resp = datacube.download(OUTPUT_FILE, OUTFORMAT)
#job.download_results(OUTPUT_FILE)
resp
# +
import matplotlib.pyplot as plt
import netCDF4
import numpy as np
# open a local NetCDF file or remote OPeNDAP URL
url = '/tmp/openeo-wcps.nc'
nc = netCDF4.Dataset(url)
# examine the variables
print (nc.variables.keys())
print (nc.variables['B8A'])
# sample every 10th point of the 'B8A' variable
topo = nc.variables['B8A'][::1,::1]
topo = np.flipud(np.rot90(topo))
# make image
plt.figure(figsize=(100,100))
plt.imshow(topo,origin='lower')
plt.title('EURAC Plot')
plt.savefig('/tmp/image.png', bbox_inches=0)
# -
| examples/notebooks/EODC_Forum_2019/EURAC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Goodreads: Science Fiction Books by Female Authors (Scraping to a CSV)
# Scrape the fields below, and save as a CSV file.
#
# - Field Example
# - Rank 1
# - Title The Handmaid's Tale
# - Author <NAME>
# - Score score: 30,733
# - Votes 314 people voted
# - Rating 4.09 avg rating — 1,101,120 ratings
import re
import pandas as pd
import requests
from bs4 import BeautifulSoup
response = requests.get('https://www.goodreads.com/list/show/6934.Science_Fiction_Books_by_Female_Authors')
doc = BeautifulSoup(response.text)
books = doc.find_all('tr')
rows = []
for book in books:
row = {}
row['Rank'] = book.find(class_="number").text.strip()
row['Title'] = book.find(class_="bookTitle").text.strip()
row['Author'] = book.find(class_="authorName").text.strip()
row['Score'] = book.find(href="#").text.strip()
row['Votes'] = book.find(href="#").find_next_sibling('a').text
row['Rating'] = book.find(class_="minirating").text.strip()
rows.append(row)
rows
# ## Cleaning Up
# +
import re
rows = []
for book in books:
row = {}
row['Rank'] = book.find(class_="number").text.strip()
row['Title'] = book.find(class_="bookTitle").text.strip()
row['Author'] = book.find(class_="authorName").text.strip()
row['Number in Series'] = book.find(class_="bookTitle").text.strip()
row['Series'] = book.find(class_="bookTitle").text.strip()
row['Score'] = book.find(href="#").text.strip()
row['Votes'] = book.find(href="#").find_next_sibling('a').text
row['Rating'] = book.find(class_="minirating").text.strip()
row['Number of Ratings'] = book.find(class_="minirating").text.strip()
rows.append(row)
df = pd.DataFrame(rows)
df.head()
# -
import re
title = "( [(].*[)])"
series_number = "(.* #)"
series = "(.*[(])"
score = "(.* )"
rating = "( .*$)"
num_ratings = "(.* — )"
votes = "( .*)$"
# +
#Title Regex
df['Title'] = df['Title'].str.replace(title, '', regex = True)
# Series Regex
df['Series'] = df['Series'].str.replace(series, '', regex = True)
df['Series'] = df['Series'].str.replace('( #[0-9]*[)]$)', '', regex = True)
df['Series'] = df['Series'].str.replace('(,$)', '', regex = True)
#Number in Series Regex
df['Number in Series'] = df['Number in Series'].str.replace(series_number, '', regex = True)
df['Number in Series'] = df['Number in Series'].str.replace('([)]$)', '', regex = True)
df['Number in Series'] = df['Number in Series'].str.replace("([a-zA-z'])", '', regex = True)
#Num of Ratings Regex
df['Number of Ratings'] = df['Number of Ratings'].str.replace(num_ratings, '', regex = True)
df['Number of Ratings'] = df['Number of Ratings'].str.replace('( .*$)', '', regex = True)
#Rating Regex
df['Rating'] = df['Rating'].str.replace(rating, '', regex = True)
#Score Regex
df['Score'] = df['Score'].str.replace(score, '', regex = True)
#Votes
df['Votes'] = df['Votes'].str.replace(votes, '', regex = True)
# -
df.head()
| 09-homework/.ipynb_checkpoints/Goodreads - Science Fiction Books-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # IRIS DATASET
# +
#Importing essential libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings('ignore')
# +
#Loading the dataset
from sklearn.datasets import load_iris
iris = load_iris()
# +
# Creating a dataframe
df = pd.DataFrame(iris.data , columns = iris.feature_names)
df.head()
# -
iris.target_names
# +
#adding a target column species
df['Species'] = iris.target
df.head()
# +
# describing the dataset
df.describe()
# -
df.info()
# Checking null values
df.isnull().any()
# +
#Scaling the features
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
df.columns
# +
# Performing MinMaxScaling
df[['sepal length (cm)', 'sepal width (cm)', 'petal length (cm)','petal width (cm)']] = scaler.fit_transform(df[['sepal length (cm)', 'sepal width (cm)', 'petal length (cm)','petal width (cm)']])
df.head()
# +
# Visualizing the data
sns.scatterplot(x = df['sepal length (cm)'] , y = df['sepal width (cm)'] ,hue = df.Species)
plt.title("Sepal Length vs Sepal Width")
# +
# Visualizing the data
sns.scatterplot(x = df['sepal length (cm)'] , y = df['petal length (cm)'] ,hue = df.Species)
plt.title("Sepal Length vs Petal Length")
# +
# Checking Correlations
plt.figure(figsize = (10,8))
sns.heatmap(df.corr(),annot= True , cmap ='PuBu')
# +
# Creating X & y
X = df.drop('Species',1)
y = df['Species']
# +
# creating X_train, X_test ,y_train, y_test
from sklearn.model_selection import train_test_split
X_train, X_test ,y_train, y_test = train_test_split(X ,y , train_size = 0.7 , random_state = 101 )
X_train, X_test ,y_train, y_test
# +
# Using RandomForestClassifier
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier(max_depth = 3 , random_state = 42 , n_jobs = -1 , n_estimators = 8 , min_samples_split= 30)
rf.fit(X_train,y_train)
rf.score(X_test,y_test)
# +
# Using Decision Trees
from sklearn.tree import DecisionTreeClassifier
dt = DecisionTreeClassifier(max_depth = 4 , random_state = 0 )
dt.fit(X_train,y_train)
dt.score(X_train,y_train)
# +
# making predictions
y_pred = dt.predict(X_test)
y_pred
# -
# Test accuracy
print("\n Test accuracy = {}".format(dt.score(X_test, y_test)))
# +
# Using XGBoost
from xgboost import XGBClassifier
xgb = XGBClassifier()
xgb.fit(X_train,y_train)
print('\n Training accuracy = {}'.format(xgb.score(X_train,y_train)))
print('\n Test accuracy = {}'.format(xgb.score(X_test,y_test)))
# +
# using logistic regression
from sklearn.linear_model import LogisticRegression
lr = LogisticRegression(n_jobs= -1 , random_state= 42 , )
lr.fit(X_train,y_train)
print('\n Training accuracy = {}'.format(lr.score(X_train,y_train)))
print('\n Test accuracy = {}'.format(lr.score(X_test,y_test)))
# -
| IRIS DataSet using Logistic Regression , XGBoost , Random Forest ,& Decision Trees.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Loading in and retrieving tip/branch point objects
# +
from imageio import imread, imsave
from matplotlib.pylab import plt
import numpy as np
from plantcv import plantcv as pcv
# Set global debug behavior to None (default), "print" (to file), or "plot" (Jupyter Notebooks or X11)
pcv.params.debug = "plot"
# +
########################################
## Segmentation Steps Here
########################################
# We skip to reading in the previously created image mask
mask, path, filename = pcv.readimage("18RGBcustom_thresh_mask.png")
#mask = pcv.rotate(mask, 90, True)
# manually crop the mask until I figure out how to reproducible split the branches
cropped_mask = mask[500:700, 300:400]
cropped_mask = pcv.rotate(cropped_mask, 90, True)
# -
#skeletonize
skeleton = pcv.morphology.skeletonize(mask=cropped_mask)
fig = plt.figure(figsize=(5,5), dpi=100)
plt.imshow(skeleton)
# +
# Adjust line thickness with the global line thickness parameter (default = 5),
# and provide binary mask of the plant for debugging.
pcv.params.line_thickness = 5
# Prune the skeleton
pruned, seg_img, edge_objects = pcv.morphology.prune(skel_img=skeleton, size=0, mask=cropped_mask)
fig = plt.figure(figsize=(5,5), dpi=100)
plt.imshow(pruned)
# -
# Identify branch points
branch_pts_mask = pcv.morphology.find_branch_pts(skel_img=skeleton, mask=cropped_mask, label="default")
# Identify tip points
tip_pts_mask = pcv.morphology.find_tips(skel_img=skeleton, mask=None, label="default")
# Sort segments into branch objects and stem objects
branch_obj, stem_obj = pcv.morphology.segment_sort(skel_img=skeleton,
objects=edge_objects,
mask=cropped_mask)
# Identify segments
segmented_img, labeled_img = pcv.morphology.segment_id(skel_img=skeleton,
objects=branch_obj,
mask=cropped_mask)
# # Branch angle identifier
def indexer(array):
to_process = np.where(array == 255)
ys = to_process[0]
xs = to_process[1]
coords = []
for i in range(len(xs)):
coord = [xs[i],ys[i]]
coords.append(coord)
return coords
# +
## find base
def find_base(tips):
y_coordinates = []
for i in range(len(indices)):
y_coordinates.append(indices[i][1])
max_index = np.argmax(y_coordinates)
to_return = tips[max_index]
return to_return
# -
## find branchpoint with smallest euclidian distance to base
import math
def find_branch_point(array,base):
to_process = np.where(array == 255)
ys = to_process[0]
xs = to_process[1]
coords = []
for i in range(len(xs)):
coord = [xs[i],ys[i]]
coords.append(coord)
coords_to_use = []
for i in range(len(coords)):
if coords[i] != base:
coords_to_use.append(coords[i])
distances = []
for i in range(len(coords_to_use)):
distance = math.sqrt((coords_to_use[i][0] - base[0])**2 + (coords_to_use[i][1] - base[1])**2)
distances.append(distance)
branch_point_index = np.argmin(distances)
branch_point_coord = coords_to_use[branch_point_index]
return branch_point_coord
# +
# I've got the relevant branch point + the relevant base. Now, find the two tips farthest from the branch point.
def find_branch_tips(array,branch_point,base):
to_process = np.where(array == 255)
ys = to_process[0]
xs = to_process[1]
coords = []
for i in range(len(xs)):
coord = [xs[i],ys[i]]
coords.append(coord)
coords_to_use = []
for i in range(len(coords)):
if coords[i] != base:
coords_to_use.append(coords[i])
# calculate distances of remaining tips to the branch point
distances = []
for i in range(len(coords_to_use)):
distance = math.sqrt((coords_to_use[i][0] - branch_point[0])**2 + (coords_to_use[i][1] - branch_point[1])**2)
distances.append(distance)
original_distances = distances
distances.sort()
first = distances[0]
second = distances[1]
top_index = original_distances.index(first)
second_index = original_distances.index(second)
branch_tips = [coords_to_use[top_index],coords_to_use[second_index]]
return branch_tips
# +
# now we've got the base, our branch point, and our two things to compare against. Now
# (1) branchpoint to branch tip 1, and branchpoint to branch tip 2 vectors
# (2) Figure out the angle between them
def branch_angle(branch_tips,branch_point):
first_branch_tip = branch_tips[0]
second_branch_tip = branch_tips[1]
branch_vector_one = [first_branch_tip[0] - branch_point[0],first_branch_tip[1] - branch_point[1]]
branch_vector_two = [second_branch_tip[0] - branch_point[0],second_branch_tip[1] - branch_point[1]]
unit_vector_1 = branch_vector_one / np.linalg.norm(branch_vector_one)
unit_vector_2 = branch_vector_two / np.linalg.norm(branch_vector_two)
dot_product = np.dot(unit_vector_1, unit_vector_2)
angle_in_radians = np.arccos(dot_product)
angle_in_degrees = np.degrees(angle_in_radians)
return angle_in_degrees
# +
## Put everything together into one function
def calculate_branch_angle(tip_pts_mask,branch_pts_mask):
indices = indexer(tip_pts_mask)
base = find_base(indices)
branch_point = find_branch_point(branch_pts_mask,base)
branch_tips = find_branch_tips(tip_pts_mask,branch_point,base)
degrees = branch_angle(branch_tips,branch_point)
return degrees
# -
result = calculate_branch_angle(tip_pts_mask,branch_pts_mask)
result
| 2021-4-16_2D_Branch_Angle_Estimation_JK.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="XV_D73H7KDZL"
from scipy.stats import norm
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import cm
import seaborn as sns
from numba import njit, prange # just-in-time compiler for parallel computation
# + [markdown] id="c7Rle9db3BtO"
# Let's consider 2 populations that are in competition; $y_1(t),\ y_2(t)$ - sizes of these populations.
#
# Assuming that in small time interval $\Delta t$:
# * the probability of a birth for the first population is $b_1y_1\Delta t = \frac{5}{6}y_1\Delta t$
# * the probability of a birth for the second population is $b_2y_2\Delta t = \frac{9}{10}y_2\Delta t$
# * the probability of a death for the first population is $d_1y_1\Delta t = (\frac{2}{5} + \frac{1}{100}y_1+\frac{1}{45}y_2)y_1 \Delta t$
# * the probability of a death for the second population is $d_2y_2\Delta t = (\frac{3}{4} + \frac{1}{150}y_2+\frac{1}{200}y_1)y_2 \Delta t$
# * the initial population sizes are $y_1(0) = y_2(0) = 15$
#
# The corresponding system of SDEs:
# $$
# \begin{cases}
# dy_i = (b_i - d_i)y_idt + B_idW_i,\ i = 1, 2\\[5pt]
# B_i = ((b_i + d_i)y_i + w)/c,\ i = 1, 2\\[5pt]
# w = \sqrt{(b_1+d_1)(b_2+d_2)y_1y_2} \\[5pt]
# c = \sqrt{(b_1+d_1)y_1 + (b_2+d_2)y_2 + 2w}
# \end{cases}
# $$
#
# Below there are several functions for modelling this system with different return parameters and simulation ending condition based on what parameter is being studied. All of them are otherwise the same and use Monte Carlo simulation and Euler's method.
#
# + [markdown] id="ViGXEBVdTRbK"
#
# # 1. Extinction probabilities and extinction time distribution
# + id="uUAf0OqRTnB7"
def Modelling(N):
step = 1 / 10
number_of_samples = N
y1_0 = 15
y2_0 = 15
@njit
def iteration():
y1, y2 = y1_0, y2_0
t = 0
y1_hist = [y1_0]
y2_hist = [y2_0]
while y1 > 0 and y2 > 0: # while both populations are not extinct
# birth and death coefficients
b1 = 5/6
b2 = 9/10
d1 = 2/5 + 1/100 * y1 + 1/45 * y2
d2 = 3/4 + 1/150 * y2 + 1/200 * y1
# drift coefficients
mu1 = (-d1 + b1) * y1
mu2 = (-d2 + b2) * y2
# diffusion coefficients
a = (d1 + b1) * y1
c = (d2 + b2) * y2
w = np.sqrt(a * c)
d = np.sqrt(a + c + 2 * w)
B1 = (a + w) / d
B2 = (c + w) / d
W1 = np.random.normal(0, step)
W2 = np.random.normal(0, step)
y1 += mu1 * step + B1 * W1
y2 += mu2 * step + B2 * W2
t += step
return y1, y2, t
ext1 = []
ext2 = []
for i in prange(number_of_samples):
y1, y2, t = iteration()
if y1 <= 0:
ext1.append(t)
else:
ext2.append(t)
return ext1, ext2
# + id="jAJCKVPHUJ5L"
np.random.seed(0)
nsamp = 100000
ext1, ext2 = Modelling(nsamp)
# + colab={"base_uri": "https://localhost:8080/"} id="vL7wAGr1FRj-" outputId="f2a1a65c-970a-405a-ce12-6206bd2bb322"
p2ext = len(ext2) / nsamp
p1ext = 1 - p2ext
print('Probability of extinction for the first population:', round(p1ext, 5))
print('Probability of extinction for the second population:', round(p2ext, 5))
# + colab={"base_uri": "https://localhost:8080/", "height": 304} id="KFLouQqUUU1K" outputId="a3da3467-c6bc-472d-927a-a0d304984477"
from scipy.stats import gaussian_kde
plt.rc('axes', titlesize=15) # fontsize of the axes title
plt.rc('axes', labelsize=14) # fontsize of the x and y labels
plt.rc('legend', fontsize=14.5) # legend fontsize
plt.rc('font', size=13) # fontsize of the tick labels
t = np.linspace(0, 300)
kd1 = gaussian_kde(ext1)
kd2 = gaussian_kde(ext2)
plt.plot(t, kd1(t), 'g', label='First population')
plt.plot(t, kd2(t), 'b', label='Second population')
plt.grid()
plt.title('Extinction time probability density');
plt.xlabel('Time')
plt.legend();
# + [markdown] id="gKB_JxzJKDZo"
# # Average and median of the trajectories
# + [markdown] id="Bk5AhugoCH2r"
# Simulating the trajectories:
# + id="vKqLjIP_HrLl"
def Modelling(N, T=200):
step = 1 / 10
number_of_samples = N
y1_0 = 15
y2_0 = 15
@njit
def iteration():
y1, y2 = y1_0, y2_0
t = 0
y1_hist = [y1_0]
y2_hist = [y2_0]
y1_dead = False
y2_dead = False
while t < T: # modelling over a fixed time interval
# birth and death coefficients
b1 = 5/6
b2 = 9/10
d1 = 2/5 + 1/100 * y1 + 1/45 * y2
d2 = 3/4 + 1/150 * y2 + 1/200 * y1
# drift coefficients
mu1 = (-d1 + b1) * y1
mu2 = (-d2 + b2) * y2
# diffusion coefficients
a = (d1 + b1) * y1
c = (d2 + b2) * y2
w = np.sqrt(a * c)
d = np.sqrt(a + c + 2 * w)
B1 = (a + w) / d
B2 = (c + w) / d
W1 = np.random.normal(0, step)
W2 = np.random.normal(0, step)
y1 = y1 + mu1 * step + B1 * W1 if not y1_dead else 0
y2 = y2 + mu2 * step + B2 * W2 if not y2_dead else 0
if y1 < 0:
y1_dead = True
y1 = 0
if y2 < 0:
y2_dead = True
y2 = 0
t += step
y1_hist.append(y1)
y2_hist.append(y2)
return y1_hist, y2_hist
p1 = []
p2 = []
for i in range(number_of_samples):
y1, y2 = iteration()
p1.append(y1)
p2.append(y2)
return np.array(p1), np.array(p2)
np.random.seed(0)
T = 200
pp1, pp2 = Modelling(10000, T=T)
# + [markdown] id="--CKRq5XCPya"
# Solving the determenistic system:
# + id="mOOC6zWDn-EQ"
def system(y, t):
y1, y2 = y
b1 = 5/6
b2 = 9/10
d1 = 2/5 + 1/100 * y1 + 1/45 * y2
d2 = 3/4 + 1/150 * y2 + 1/200 * y1
dydt = [(b1 - d1) * y1, (b2 - d2) * y2]
return dydt
y0 = [15, 15]
t = np.linspace(0, 200, 2002)
from scipy.integrate import odeint
sol = odeint(system, y0, t)
# + [markdown] id="Tr4sGxMvCW0s"
# Plotting the trajectories:
# + colab={"base_uri": "https://localhost:8080/", "height": 376} id="g4EK1Oe1o6SQ" outputId="24f97271-cb39-4884-b39a-dd16dde8dbbb"
from scipy.stats import gaussian_kde # used for density approximation
fig, ax = plt.subplots(1, 2, figsize=(14,5))
plt.rc('axes', titlesize=15) # fontsize of the axes title
plt.rc('axes', labelsize=14) # fontsize of the x and y labels
plt.rc('legend', fontsize=14.5) # legend fontsize
plt.rc('font', size=13) # fontsize of the tick labels
ax[0].plot(t, sol[:, 0], 'g', label='$y_1^{det}(t)$')
ax[0].plot(t, np.mean(pp1, axis=0), color='green', linestyle='--', label='$Ey_1(t)$')
ax[0].plot(t, sol[:, 1], 'b', label='$y_2^{det}(t)$')
ax[0].plot(t, np.mean(pp2, axis=0), color='blue', linestyle='--', label='$Ey_2(t)$')
ax[0].legend(loc='best')
ax[0].set_title('Determenistic solution and \naverage of 10000 trajectories')
ax[0].set_xlabel('Time')
ax[0].set_ylabel('Population size')
ax[0].grid()
kd1 = gaussian_kde(ext1)
kd2 = gaussian_kde(ext2)
ax[1].plot(t, sol[:, 0], 'g', label='$y_1^{det}(t)$')
ax[1].plot(t, np.median(pp1, axis=0), color='green', linestyle='--', label='$Me\ y_1(t)$')
ax[1].plot(t, sol[:, 1], 'b', label='$y_2^{det}(t)$')
ax[1].plot(t, np.median(pp2, axis=0), color='blue', linestyle='--', label='$Me\ y_2(t)$')
ax[1].grid()
ax[1].set_title('Determenistic solution and \nmedian of 10000 trajectories');
ax[1].set_xlabel('Time');
ax[1].set_ylabel('Population size')
plt.legend();
# + [markdown] id="h-8vRNVyuqkS"
# # Estimation of the population size probabilty density at time t
# + id="oyJHsAKYuo6s"
def Modelling(N, population=1, T=200):
step = 1 / 10
number_of_samples = N
y1_0 = 15
y2_0 = 15
@njit
def iteration():
y1, y2 = y1_0, y2_0
t = 0
y1_hist = [y1_0]
y2_hist = [y2_0]
y1_dead = False
y2_dead = False
while t < T:
# birth and death coefficients
b1 = 5/6
b2 = 9/10
d1 = 2/5 + 1/100 * y1 + 1/45 * y2
d2 = 3/4 + 1/150 * y2 + 1/200 * y1
# drift coefficients
mu1 = (-d1 + b1) * y1
mu2 = (-d2 + b2) * y2
# diffusion coefficients
a = (d1 + b1) * y1
c = (d2 + b2) * y2
w = np.sqrt(a * c)
d = np.sqrt(a + c + 2 * w)
B1 = (a + w) / d
B2 = (c + w) / d
W1 = np.random.normal(0, step)
W2 = np.random.normal(0, step)
y1 = y1 + mu1 * step + B1 * W1 # if not y1_dead else 0
y2 = y2 + mu2 * step + B2 * W2 # if not y2_dead else 0
if y1 < 0:
y1_dead = True
y1 = 0
if y2 < 0:
y2_dead = True
y2 = 0
t += step
y1_hist.append(y1)
y2_hist.append(y2)
return y1_hist, y2_hist
p1 = []
p2 = []
i = 0
for i in range(number_of_samples):
y1, y2 = iteration()
p1.append(y1)
p2.append(y2)
return np.array(p1), np.array(p2)
np.random.seed(0)
T = 200
nsamp = 3000
pp1, pp2 = Modelling(nsamp, T=T)
# + [markdown] id="rZo6FCgGC_iC"
# Filtering by which population went extinct:
# + id="RC-ocB_Z8WqY"
pp1not_dead = []
for traj in pp1:
if traj[-1] != 0:
pp1not_dead.append(traj)
pp1not_dead = np.array(pp1not_dead)
pp2not_dead = []
for traj in pp2:
if traj[-1] != 0:
pp2not_dead.append(traj)
pp2not_dead = np.array(pp2not_dead)
# + [markdown] id="qJipodZGDXVq"
# Approximating density:
# + id="yQaK3v-Fv03x"
from scipy.stats import gaussian_kde
starting_row = 15
t = np.linspace(0, T, len(pp2not_dead[0]))[starting_row:]
X = np.linspace(0, 50, 1000)
dens1 = []
dens2 = []
tmp1 = pp1not_dead.transpose()
tmp2 = pp2not_dead.transpose()
for i in range(starting_row, len(tmp1)):
dens1.append(gaussian_kde(tmp1[i])(X))
dens2.append(gaussian_kde(tmp2[i])(X))
dens1 = np.array(dens1)
dens2 = np.array(dens2)
X, t = np.meshgrid(X, t)
# -
# Plotting approximated density:
# + colab={"base_uri": "https://localhost:8080/", "height": 684} id="lc9NsQZRv7pR" outputId="5007572c-2833-4bb2-8dff-84ad8ef6167a"
fig = plt.figure(figsize=(14, 10))
plt.rc('figure', titlesize=18) # fontsize of the figure title
plt.rc('axes', titlesize=15) # fontsize of the axes title
plt.rc('axes', labelsize=14) # fontsize of the x and y labels
plt.rc('legend', fontsize=13) # legend fontsize
plt.rc('font', size=13) # fontsize of the tick labels
ax = fig.add_subplot(2, 2, 1, projection='3d')
ax.set_title('The first population\n(survival probability$\\approx$0.68)')
surf = ax.plot_surface(t, X, dens1, linewidth=0, cmap=plt.get_cmap('coolwarm'))
fig.colorbar(surf, shrink=0.5, aspect=5);
ax2 = fig.add_subplot(2, 2, 2, projection='3d')
ax2.set_title('The second population\n(survival probability$\\approx$0.32)')
surf2 = ax2.plot_surface(t, X, dens2, linewidth=0, cmap=plt.get_cmap('coolwarm'))
fig.colorbar(surf2, shrink=0.5, aspect=5);
fig.suptitle('Conditional probability density (if survived)')
for a in (ax, ax2):
a.set_xlabel('Time')
a.set_ylabel('Population size')
a.set_zlabel('')
a.view_init(30, 140)
ax3 = fig.add_subplot(2, 2, 3)
im = ax3.pcolormesh(X, t, dens1, cmap=plt.get_cmap('Spectral'))
fig.colorbar(im, ax=ax3)
ax4 = fig.add_subplot(2, 2, 4)
im = ax4.pcolormesh(X, t, dens2, cmap=plt.get_cmap('Spectral'))
fig.colorbar(im, ax=ax4);
for a in ax3, ax4:
a.set_xlabel('Population size')
a.set_ylabel('Time')
| Modelling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
N_FOLDS = 5
oof_df = pd.read_parquet('cache/new_oof_ensemble.parquet')
print(oof_df.shape)
oof_df.head()
# -
gt_df = pd.read_csv("../train_folds.csv")
print(gt_df.shape)
gt_df.head()
# +
oof_df = oof_df.drop("kfold", axis=1).merge(gt_df[["id", "kfold"]].drop_duplicates(), on="id", how="left")
oof_df["kfold"].isnull().mean()
# +
from utils import score_feedback_comp
score_feedback_comp(oof_df[oof_df["f_7"] > 0.9999], gt_df, return_class_scores=True)
# +
oof_df["begin"] = oof_df["predictionstring"].apply(lambda x: int(str(x).split()[0]))
#oof_df["end"] = oof_df["predictionstring"].apply(lambda x: int(str(x).split()[-1]))
#oof_df["len"] = oof_df["end"] - oof_df["begin"] + 1
#oof_df["loc"] = (oof_df["begin"] + oof_df["end"]) / 2
oof_df = oof_df.sort_values(["id", "begin"]).reset_index(drop=True)
oof_df.head(20)
# -
oof_df["target"] = oof_df["overlap"] >= 0.5
oof_df["target"].mean()
discourse_types = gt_df["discourse_type"].unique()
# +
#for dtype in discourse_types:
# oof_df["x"] = oof_df["avg_score"]*(oof_df["class"] == dtype)
# oof_df[f"sum_{dtype}"] = oof_df.groupby("id")["x"].transform("sum")
#oof_df.drop("x", axis=1, inplace=True)
#oof_df.head()
# +
from tqdm import tqdm
import os
def read_texts(directory):
names, texts = [], []
for f in tqdm(list(os.listdir(directory))):
names.append(f.replace('.txt', ''))
texts.append(open(directory + f, 'r').read())
df = pd.DataFrame({'id': names, 'full_text': texts})
return df
#text_df = read_texts("data/train/")
#print(text_df.shape)
#text_df.head()
#oof_df = oof_df.merge(text_df, on="id", how="left")
#oof_df["loc_rate"] = oof_df["loc"] / oof_df["full_text"].apply(lambda x: len(x.split()))
# +
features_dict = {'Lead': [f"f_{i}" for i in range(34)],
'Position': [f"f_{i}" for i in range(34)],
'Evidence': [f"f_{i}" for i in range(20)],
'Claim': [f"f_{i}" for i in range(20)],
'Concluding Statement': [f"f_{i}" for i in range(34)],
'Counterclaim': [f"f_{i}" for i in range(17)] + [f"f_{i}" for i in range(27, 34)],
'Rebuttal': [f"f_{i}" for i in range(17)]}
target = "target"
# +
import xgboost as xgb
from sklearn.metrics import roc_auc_score, f1_score
param = {'objective': 'reg:logistic',
'eval_metric': "auc",
'learning_rate': 0.05,
'max_depth': 4,
"min_child_weight": 200,
"colsample_bynode": 0.8,
"subsample": 0.5,
"tree_method": 'gpu_hist', "gpu_id": 1
}
y_oof = np.zeros(oof_df.shape[0])
res = dict()
best_th = dict()
lvl1_stacking_df = []
for dtype in discourse_types:
features = features_dict[dtype]
all_indices = np.where(oof_df["class"] == dtype)[0]
discourse_df = oof_df[oof_df["class"] == dtype].reset_index(drop=True)
tm = discourse_df["target"].mean()
param["scale_pos_weight"] = (1 - tm)/tm
best_its = []
print(dtype, len(features))
for f in range(N_FOLDS):
val_ind = all_indices[np.where(discourse_df["kfold"] == f)[0]]
train_df, val_df = discourse_df[discourse_df["kfold"] != f], discourse_df[discourse_df["kfold"] == f]
d_train = xgb.DMatrix(train_df[features], train_df[target])
d_val = xgb.DMatrix(val_df[features], val_df[target])
model = xgb.train(param, d_train, evals=[(d_val, "val")], num_boost_round=2000, verbose_eval=50,
early_stopping_rounds=50)
model.save_model(f'gbm_models/xgb_{dtype}_{f}.json')
y_oof[val_ind] = model.predict(d_val)
best_its.append(model.best_iteration)
print("...")
discourse_df["prob"] = y_oof[all_indices]
discourse_df = discourse_df.sort_values(["id", "prob"], ascending=False).reset_index(drop=True)
print("removing overlaps...")
pred_df = []
prev_id = -1
overlap_array = np.zeros(4096)
for id, ps, cls, prob, overlap in zip(discourse_df["id"].values, discourse_df["predictionstring"].values,
discourse_df["class"].values, discourse_df["prob"].values,
discourse_df["overlap"].values):
if id != prev_id:
prev_id = id
overlap_array = np.zeros(4096)
ps_list = ps.split()
begin, end = int(ps_list[0]), int(ps_list[-1]) + 1
intersect = np.sum(overlap_array[begin:end])
total = end - begin
condition = intersect <= 1
if dtype in {"Counterclaim", "Rebuttal"}:
condition = intersect/total <= 0.2
if condition:
pred_df.append({"id": id, "class": cls, "prob": prob, "predictionstring": ps, "overlap": overlap})
overlap_array[begin:end] = 1
pred_df = pd.DataFrame(pred_df)
lvl1_stacking_df.append(pred_df)
print("tuning...")
thresholds = np.arange(10, 90, 1)/100
all_gt = gt_df[gt_df["discourse_type"] == dtype].shape[0]
print(pred_df.shape, all_gt)
f1s = []
for t in thresholds:
tp = (pred_df[pred_df["prob"] > t]["overlap"] >= 0.5).sum()
fp = (pred_df[pred_df["prob"] > t]["overlap"] < 0.5).sum()
fn = all_gt - tp
f1 = tp / (tp + (fp + fn)/2)
#print(t, tp, fp, fn)
f1s.append(f1)
best_ind = np.argmax(f1s)
print()
print(thresholds[best_ind], f1s[best_ind])
print()
best_th[dtype] = thresholds[best_ind]
res[dtype] = f1s
#d_train = xgb.DMatrix(discourse_df[features], discourse_df[target])
#model = xgb.train(param, d_train, num_boost_round=int(np.mean(best_its)*1.2))
#model.save_model(f'models/xgb_{dtype}.json')
# -
best_th
{k: np.mean(v) for k, v in res.items()}
lvl1_stacking_df = pd.concat(lvl1_stacking_df)
lvl1_stacking_df.shape
score_feedback_comp(lvl1_stacking_df[lvl1_stacking_df["prob"] > lvl1_stacking_df["class"].map(best_th)],
gt_df, return_class_scores=True)
xgb.plot_importance(model, importance_type="gain")
len(oof_df["id"].unique())
len(gt_df["id"].unique())
lvl1_stacking_df.to_parquet("cache_oof2/level1_oof.parquet", index=False)
lvl1_stacking_df.sort_values("prob")
# +
th_df = pd.DataFrame()
for k, v in res.items():
th_df[k] = v
th_df["threshold"] = np.arange(10, 90, 1)/100
th_df.plot(figsize=(12, 12), x="threshold")
# -
| Stacking/Stacking_shujun_72958_4096.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('../python_packages_static/')
import pandas as pd
import numpy as np
import os
import pyemu
import matplotlib as mpl
import matplotlib.pyplot as plt
# # read in the PST file and the Morris output (in the MSN file)
# ### NOTE: Make sure `run_ensemble` is set appropriately - If `run_ensemble` is set to `True`, local runs are performed. If `run_ensemble` set to `False`results from the journal article are used.
run_ensemble=False
if run_ensemble==True:
resdir = '../run_data'
else:
resdir = '../output/parallel'
pstfile = 'prior_mc_wide_sens.pst'
pst = pyemu.Pst(os.path.join(resdir, pstfile))
morris_sum_file = os.path.join(resdir, pstfile.replace('.pst','.msn'))
morris_sum_df = pd.read_csv(morris_sum_file, index_col=0)
# ## Set up the parameter names
pardata = pst.parameter_data
morris_sum_df['pargp']=pardata.loc[pardata.index.isin(morris_sum_df.index)].pargp
morris_sum_df.set_index(morris_sum_df.pargp, drop=True, inplace=True)
morris_sum_df
renames = {'k_pp_:0':'Kh PPs: Layer 1',
'k_pp_:1':'Kh PPs: Layer 2',
'k_pp_:2':'Kh PPs: Layer 3',
'k_pp_:3':'Kh PPs: Layer 4',
'k33_pp_:0':'Kv PPs: Layer 1',
'k33_pp_:1':'Kv PPs: Layer 2',
'k33_pp_:2':'Kv: Layer 3',
'k33_pp_:3':'Kv PPs: Layer 4' ,
'rch_pp_:0':'Recharge PPs',
'pp_rch__multiplier':'Mean Recharge',
'sfrk':'SFR K by reach',
'wel':'Well Pumping',
'chd': 'Constant Head',
'zn_k__multiplier':'Kh by Zone',
'zn_k33__multiplier':'Kv by Zone'}
# ## Just make a barchart of the renamed parameter groups to show the relative sensitivity. We plot the log because the elemetary effects span orders of magnitude
if ' sen_mean_abs' in morris_sum_df.columns:
morris_sum_df = morris_sum_df.rename(columns={' sen_mean_abs':'sen_mean_abs'}) # remove space in df column name
# +
from matplotlib.ticker import ScalarFormatter
morris_sum_df.index = [renames[i] for i in morris_sum_df.index]
mpl.rcParams.update({'font.size':16})
ax = morris_sum_df.sort_values(by='sen_mean_abs',ascending=False).sen_mean_abs.apply(np.log10).plot.bar(figsize=(7,7))
#ax.set_yscale('log')
ax.set_ylabel('Log Mean absolute value of elementary effects ($\mu*$)')
ax.set_xlabel('Parameter Group')
ax.yaxis.set_major_formatter(ScalarFormatter())
ax.ticklabel_format(useOffset=False,style='plain', axis='y', )
ax.grid()
plt.tight_layout()
plt.title('Method of Morris Global Sensitivity Results')
plt.savefig('../figures/morris_plot.pdf')
# -
| notebooks_workflow_blank/2.1_Sensitivity_Results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.12 ('visionrobots')
# language: python
# name: python3
# ---
# # Features and feature detection
# ---
#
# Most of you will have played the jigsaw puzzle games. You get a lot of small pieces of an image, where you need to assemble them correctly to form a big real image. The question is, how you do it?
#
# 
#
# But it all depends on the most basic question: How do you play jigsaw puzzles? How do you arrange lots of scrambled image pieces into a big single image? How can you stitch a lot of natural images to a single image?
#
# The answer is, we are looking for specific patterns or specific features which are unique, can be easily tracked and can be easily compared. If we go for a definition of such a feature, we may find it difficult to express it in words, but we know what they are. If someone asks you to point out one good feature which can be compared across several images, you can point out one. That is why even small children can simply play these games. We search for these features in an image, find them, look for the same features in other images and align them. That's it. (In jigsaw puzzle, we look more into continuity of different images).
#
# So our one basic question expands to more in number, but becomes more specific. What are these features?.
#
# 
#
# In computer vision and image processing, a feature is a piece of information about the content of an image; typically about whether a certain region of the image has certain properties. Features may be specific structures in the image such as points, edges or objects. Features may also be the result of a general neighborhood operation or feature detection applied to the image. Other examples of features are related to motion in image sequences, or to shapes defined in terms of curves or boundaries between different image regions.
#
# 
#
# The image is very simple. At the top of image, six small image patches are given. Question for you is to find the exact location of these patches in the original image. How many correct results can you find?
#
# A and B are flat surfaces and they are spread over a lot of area. It is difficult to find the exact location of these patches.
#
# C and D are much more simple. They are edges of the building. You can find an approximate location, but exact location is still difficult. This is because the pattern is same everywhere along the edge. At the edge, however, it is different. An edge is therefore better feature compared to flat area, but not good enough (It is good in jigsaw puzzle for comparing continuity of edges).
#
# Finally, E and F are some corners of the building. And they can be easily found. Because at the corners, wherever you move this patch, it will look different. So they can be considered as good features. So now we move into simpler (and widely used image) for better understanding.
#
#
# # Harris Corner Detection
# ---
#
# One early attempt to find corners was done by <NAME> & <NAME> in their paper A Combined Corner and Edge Detector in 1988, so now it is called the Harris Corner Detector.
#
# A corner is a point whose local neighborhood stands in two dominant and different edge directions. In other words, a corner can be interpreted as the junction of two edges, where an edge is a sudden change in image brightness. Corners are the important features in the image, and they are generally termed as interest points which are invariant to translation, rotation, and illumination.
#
# 
#
# OpenCV has the function cv2.cornerHarris(). Its arguments are:
#
# - img: Input image. It should be grayscale and float32 type.
# - blockSize: It is the size of neighbourhood considered for corner detection
# - ksize: Aperture parameter of the Sobel derivative used.
# - k: Harris detector free parameter in the equation.
import cv2
import matplotlib.pyplot as plt
import numpy as np
# +
image = cv2.imread('images/chess.jpg')
gray = cv2.cvtColor(image,cv2.COLOR_BGR2GRAY)
plt.imshow(gray, cmap='gray')
# -
# ### Corner detection
# +
# Detect corners using Harris Corner detector
# - img: Input image. It should be grayscale and float32 type.
# - blockSize: It is the size of neighbourhood considered for corner detection
# - ksize: Aperture parameter of the Sobel derivative used.
# - k: Harris detector free parameter in the equation.
dst = cv2.cornerHarris(gray,2,3,0.04)
# Dilate corners to enhance center points
kernel = np.ones((5,5), np.uint8)
dst = cv2.dilate(dst,kernel, iterations=3)
plt.imshow(dst)
# -
# ### Detect "strong" corners
# +
# This value vary depending on the image and how many corners you want to detect
# Try changing this free parameter, 0.1, to be larger or smaller and see what happens
thresh = 0.1*dst.max()
# Create an image copy to draw corners on
corner_image = np.copy(image)
# Change all the pixels of the corners to red
corner_image[dst>thresh]=[255,0,0]
plt.imshow(corner_image)
# +
image_shear = cv2.imread('images/chess_transform.jpg')
gray_shear = cv2.cvtColor(image_shear,cv2.COLOR_BGR2GRAY)
plt.imshow(gray_shear, cmap='gray')
# +
# Detect corners
dst_shear = cv2.cornerHarris(gray_shear,2,3,0.04)
# Dilate corners to enhance center points
kernel = np.ones((5,5), np.uint8)
dst_shear = cv2.dilate(dst_shear,kernel, iterations=2)
# This value vary depending on the image and how many corners you want to detect
thresh_shear = 0.01*dst_shear.max()
# Create an image copy to draw corners on
corner_image_shear = np.copy(image_shear)
# Change all the pixels of the corners to red
corner_image_shear[dst_shear>thresh_shear]=[255,0,0]
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(20,10))
ax1.imshow(dst_shear)
ax2.imshow(corner_image_shear)
# -
# # Shi-Tomasi Corner Detector & Good Features to Track
# ---
#
# In 1994, <NAME> and <NAME> made a small modification to the Harris corner detector in their paper Good Features to Track which shows better results when compared to the original algorithm.
#
# OpenCV has a function, cv.goodFeaturesToTrack(). It finds N strongest corners in the image. As usual, image should be a grayscale image. Then you specify number of corners you want to find. Then you specify the quality level, which is a value between 0-1, which denotes the minimum quality of corner below which everyone is rejected. Then we provide the minimum euclidean distance between corners detected.
#
# With all this information, the function finds corners in the image. All corners below quality level are rejected. Then it sorts the remaining corners based on quality in the descending order. Then function takes first strongest corner, throws away all the nearby corners in the range of minimum distance and returns N strongest corners.
# +
image = cv2.imread('images/blox.jpg')
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
corners_shi = np.copy(image)
# Shi and Tomasi corner detector
# image: grayscale image
# N number of corners
# Quality of the corner, between 0 and 1
# Euclidian distance bewteen corners
corners = cv2.goodFeaturesToTrack(gray,30,0.1,10)
corners = np.int0(corners)
for i in corners:
x,y = i.ravel()
cv2.circle(corners_shi,(x,y),3,(255,0,0),-1)
# Detect corners with Harris
dst = cv2.cornerHarris(gray,2,3,0.04)
kernel = np.ones((3,3), np.uint8)
dst = cv2.dilate(dst,kernel, iterations=1)
thresh = 0.01*dst.max()
# Create an image copy to draw corners on
corner_image = np.copy(image)
# Change all the pixels of the corners to red
corner_image[dst>thresh]=[255,0,0]
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(20,10))
ax1.imshow(corners_shi)
ax2.imshow(corner_image)
# -
| Notebooks/12. CV. Feature detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Building WaveWatch3(TM) ERDDAP Datasets
#
# This notebook documents the process of creating XML fragments
# for SalishSeaCast rolling forecast WaveWatch3(TM) run results files
# for inclusion in `/results/erddap-datasets/datasets.xml`
# which is symlinked to `/opt/tomcat/content/erddap/datasets.xml`
# on the `skookum` ERDDAP server instance.
#
# The contents are a combination of:
#
# * instructions for using the
# `GenerateDatasetsXml.sh` and `DasDds.sh` tools found in the
# `/opt/tomcat/webapps/erddap/WEB-INF/` directory
# * instructions for forcing the server to update the datasets collection
# via the `/results/erddap/flags/` directory
# * code and metadata to transform the output of `GenerateDatasetsXml.sh`
# into XML fragments that are ready for inclusion in `/results/erddap-datasets/datasets.xml`
# +
from collections import OrderedDict
from copy import copy
from lxml import etree
# -
# **NOTE**
#
# The next cell mounts the `/results` filesystem on `skookum` locally.
# It is intended for use if when this notebook is run on a laptop
# or other non-Waterhole machine that has `sshfs` installed
# and a mount point for `/results` available in its root filesystem.
#
# Don't execute the cell if that doesn't describe your situation.
# # !sshfs skookum:/results /results
# # Metadata for All Datasets
#
# The `metadata` dictionary below contains information for dataset
# attribute tags whose values need to be changed,
# or that need to be added for all datasets.
#
# The keys are the dataset attribute names.
#
# The values are dicts containing a required `text` item
# and perhaps an optional `after` item.
#
# The value associated with the `text` key is the text content
# for the attribute tag.
#
# When present,
# the value associated with the `after` key is the name
# of the dataset attribute after which a new attribute tag
# containing the `text` value is to be inserted.
metadata = OrderedDict([
('infoUrl', {
'text':
'https://salishsea-meopar-docs.readthedocs.io/en/latest/results_server/index.html#salish-sea-model-results',
}),
('institution', {
'text': 'UBC EOAS',
}),
('institution_fullname', {
'text': 'Earth, Ocean & Atmospheric Sciences, University of British Columbia',
'after': 'institution',
}),
('license', {
'text': '''The Salish Sea MEOPAR NEMO model results are copyright
by the Salish Sea MEOPAR Project Contributors and The University of British Columbia.
They are licensed under the Apache License, Version 2.0. https://www.apache.org/licenses/LICENSE-2.0''',
}),
('project', {
'text':'Salish Sea MEOPAR NEMO Model',
'after': 'title',
}),
('creator_name', {
'text': 'Salish Sea MEOPAR Project Contributors',
'after': 'project',
}),
('creator_email', {
'text': '<EMAIL>',
'after': 'creator_name',
}),
('creator_url', {
'text': 'https://salishsea-meopar-docs.readthedocs.io/',
'after': 'creator_email',
}),
('acknowledgement', {
'text': 'MEOPAR, ONC, Compute Canada',
'after': 'creator_url',
}),
('drawLandMask', {
'text': 'over',
'after': 'acknowledgement',
}),
])
# # Dataset Attributes
#
# The `datasets` dictionary below provides the content
# for the dataset `title` and `summary` attributes.
#
# The `title` attribute content appears in the the datasets list table
# (among other places).
# It should be `<`80 characters long,
# and note that only the 1st 40 characters will appear in the table.
#
# The `summary` attribute content appears
# (among other places)
# when a user hovers the cursor over the `?` icon beside the `title`
# content in the datasets list table.
# The text that is inserted into the `summary` attribute tag
# by code later in this notebook is the
# `title` content followed by the `summary` content,
# separated by a blank line.
#
# The keys of the `datasets` dict are the `datasetID` strings that
# are used in many places by the ERDDAP server.
# They are structured as follows:
#
# * `ubc` to indicate that the dataset was produced at UBC
# * `SS` to indicate that the dataset is a product of the Salish Sea NEMO model
# * a few letters to indicate the model runs that produce the dataset:
#
# * `n` to indicate that the dataset is from a nowcast run,
# * `f` for rolling forecast composed of the more recent 5 days of nowcast run results and the most recent forecast or forecast2 run,
# * `g` for nowcast-green,
# * `a` for atmospheric forcing,
# * a description of the dataset variables; e.g. `PointAtkinsonSSH` or `3DuVelocity`
# * the time interval of values in the dataset; e.g. `15m`, `1h`, `1d`
# * the dataset version; e.g. `V16-10`, or `V1`
#
# Versioning was changed to a [CalVer](http://calver.org/) type scheme in Oct-2016.
# Thereafter versions are of the form `Vyy-mm` and indicate the year and month when the dataset entered production.
#
# So:
#
# * `ubcSSnPointAtkinsonSSH15mV1` is the version 1 dataset of 15 minute averaged sea surface height values at Point Atkinson from `PointAtkinson.nc` output files
#
# * `ubcSSn3DwVelocity1hV2` is the version 2 dataset of 1 hr averaged vertical (w) velocity values over the entire domain from `SalishSea_1h_*_grid_W.nc` output files
#
# * `ubcSSnSurfaceTracers1dV1` is the version 1 dataset of daily averaged surface tracer values over the entire domain from `SalishSea_1d_*_grid_T.nc` output files
#
# * `ubcSSnBathymetry2V16-07` is the version 16-07 dataset of longitude, latitude, and bathymetry of the Salish Sea NEMO model grid that came into use in Jul-2016.
# The corresponding NEMO-generated mesh mask variables are in the `ubcSSn2DMeshMaskDbo2V16-07` (y, x variables),
# and the `ubcSSn3DMeshMaskDbo2V16-07` (z, y, x variables) datasets.
#
# The dataset version part of the `datasetID` is used to indicate changes in the variables
# contained in the dataset.
# For example,
# the transition from the `ubcSSn3DwVelocity1hV1` to the `ubcSSn3DwVelocity1hV2` dataset
# occurred on 24-Jan-2016 when we started to output vertical eddy viscosity and diffusivity
# values at the `w` grid points.
#
# All dataset ids end with their version identifier and their `summary` ends with a notation about the variables
# that they contain; e.g.
# ```
# v1: wVelocity variable
# ```
# When the a dataset version is incremented a line describing the change is added
# to the end of its `summary`; e.g.
# ```
# v1: wVelocity variable
# v2: Added eddy viscosity & diffusivity variables ve_eddy_visc & ve_eddy_diff
# ```
datasets = {
'ubcSSf2DWaveFields30mV17-02': {
'type': '2d fields',
'title': 'Forecast, Salish Sea, 2d Wave Fields, 30min, v17-02',
'keywords': '''atmosphere,
Atmosphere > Atmospheric Winds > Surface Winds,
atmospheric, breaking wave height, circulation, currents, direction, drift,
eastward_sea_water_velocity, eastward_surface_stokes_drift, eastward_wave_to_ocean_stress, eastward_wind,
energy, flux, foc, frequency, height, latitude, length, local, longitude, mean, mean_wave_length, moment,
northward, northward_sea_water_velocity, northward_surface_stokes_drift, northward_wave_to_ocean_stress, northward_wind, ocean, oceans,
Oceans > Ocean Circulation > Ocean Currents,
Oceans > Ocean Waves > Significant Wave Height,
Oceans > Ocean Waves > Wave Frequency,
Oceans > Ocean Waves > Wave Period,
Oceans > Ocean Waves > Wave Spectra,
Oceans > Ocean Waves > Wave Speed/Direction,
Oceans > Ocean Waves > Wind Waves,
peak, period, sea, sea_surface_wave_from_direction, sea_surface_wave_peak_direction, sea_surface_wave_peak_frequency,
sea_surface_wave_significant_height, sea_surface_wind_wave_mean_period_from_variance_spectral_density_second_frequency_moment,
seawater, second, significant, significant_breaking_wave_height, source, spectra, spectral, speed, stokes, stress, surface,
swell, t02, time, ucur, utwo, uuss, uwnd, variance, vcur, velocity, vtwo, vuss, vwnd, water, wave, wave_to_ocean_energy_flux,
waves, wcc, wch, whitecap coverage, whitecap_coverage, wind, winds''',
'summary': '''2d wave field values calculated at 30 minute intervals
from the most recent Strait of Georgia WaveWatch3(TM) model forecast runs.
The values are calculated for a model grid that covers the Strait of Georgia
on the coast of British Columbia. The time values are UTC.
The Strait of Georgia WaveWatch3(TM) model grid and configuration were developed
by <NAME> at the University of Victoria. The WaveWatch3(TM) model
is forced with currents from the Salish Sea NEMO model and the same ECCC HRDPS
GEM 2.5km resolution winds that are used to force the NEMO model.
This dataset is updated daily to move it forward 1 day in time.
It starts at 00:00:00 UTC 5 days prior to the most recently completed forecast run,
and extends to 11:30:00 UTC on the 2nd day after the forecast run date.
So, for example, after completion of the 10-Nov-2017 forecast run,
this dataset included data from 2017-11-05 00:00:00 UTC to 2017-11-12 11:30:00 UTC.
v17-02: WaveWatch3(TM)-5.16; NEMO-3.6; ubcSSnBathymetryV17-02 bathymetry; see infoUrl link for full details.
''',
'fileNameRegex': '.*SoG_ww3_fields_\d{8}_\d{8}\.nc$',
}
}
# # Convenience Functions
#
# A few convenient functions to reduce code repetition:
def print_tree(root):
"""Display an XML tree fragment with indentation.
"""
print(etree.tostring(root, pretty_print=True).decode('ascii'))
def find_att(root, att):
"""Return the dataset attribute element named att
or raise a ValueError exception if it cannot be found.
"""
e = root.find('.//att[@name="{}"]'.format(att))
if e is None:
raise ValueError('{} attribute element not found'.format(att))
return e
# +
def update_xml(root, datasetID, metadata, datasets):
root.attrib['datasetID'] = datasetID
root.find('.//fileNameRegex').text = datasets[datasetID]['fileNameRegex']
title = datasets[datasetID]['title']
if 'keywords' in datasets[datasetID]:
keywords = find_att(root, 'keywords')
keywords.text = datasets[datasetID]['keywords']
summary = find_att(root, 'summary')
summary.text = f'{title}\n\n{datasets[datasetID]["summary"]}'
e = etree.Element('att', name='title')
e.text = title
summary.addnext(e)
for att, info in metadata.items():
e = etree.Element('att', name=att)
e.text = info['text']
try:
root.find(f'''.//att[@name="{info['after']}"]'''.format()).addnext(e)
except KeyError:
find_att(root, att).text = info['text']
attrs = root.find('addAttributes')
etree.SubElement(attrs, 'att', name='NCO').text = 'null'
if not 'Bathymetry' in datasetID:
etree.SubElement(attrs, 'att', name='history').text = 'null'
etree.SubElement(attrs, 'att', name='name').text = 'null'
for axis_name in root.findall('.//axisVariable/destinationName'):
attrs = axis_name.getparent().find('addAttributes')
etree.SubElement(attrs, 'att', name='coverage_content_type').text = 'modelResult'
if axis_name.text == 'time':
etree.SubElement(attrs, 'att', name='comment').text = ('time values are UTC')
# for var_name in root.findall('.//dataVariable/destinationName'):
# if var_name.text in dataset_vars:
# var_name.text = dataset_vars[var_name.text]['destinationName']
# if var_name.text in var_colour_ranges:
# for att_name in ('colorBarMinimum', 'colorBarMaximum'):
# cb_att = var_name.getparent().find(f'addAttributes/att[@name="{att_name}"]')
# if cb_att is not None:
# cb_att.text = var_colour_ranges[var_name.text][att_name]
# else:
# attrs = var_name.getparent().find('addAttributes')
# etree.SubElement(attrs, 'att', name=att_name, type='double').text = (
# var_colour_ranges[var_name.text][att_name])
# attrs = var_name.getparent().find('addAttributes')
# etree.SubElement(attrs, 'att', name='coverage_content_type').text = 'modelResult'
# etree.SubElement(attrs, 'att', name='cell_measures').text = 'null'
# etree.SubElement(attrs, 'att', name='cell_methods').text = 'null'
# etree.SubElement(attrs, 'att', name='interval_operation').text = 'null'
# etree.SubElement(attrs, 'att', name='interval_write').text = 'null'
# etree.SubElement(attrs, 'att', name='online_operation').text = 'null'
# if var_name.text in ioos_categories:
# etree.SubElement(attrs, 'att', name='ioos_category').text = ioos_categories[var_name.text]
# -
# # Generate Initial Dataset XML Fragment
# Now we're ready to produce a dataset!!!
#
# Use the `/opt/tomcat/webapps/erddap/WEB-INF/GenerateDatasetsXml.sh` script
# generate the initial version of an XML fragment for a dataset:
# ```
# $ cd /opt/tomcat/webapps/erddap/WEB-INF/
# $ bash GenerateDatasetsXml.sh EDDGridFromNcFiles /results/SalishSea/rolling-forecasts/ ".*SoG_ww3_fields_\d{8}_\d{8}\.nc$" "" 10080
# ```
# The `EDDGridFromNcFiles`,
# `/results/SalishSea/nowcast/`,
# `".*SalishSea_1h_\d{8}_\d{8}_grid_U\.nc$"`,
# `""`,
# and `10080` arguments
# tell the script:
#
# * which `EDDType`
# * what parent directory to use
# * what file name regex to use
# * `""` to concatenate the parent directory and the file name regex to find a sample file
# * to reload the dataset every 10080 minutes
#
# avoiding having to type those in answer to prompts.
#
# The output is written to `/results/erddap/logs/GenerateDatasetsXml.out`
#
# Dataset ids and file name regexs from datasets dict:
for dataset in sorted(datasets):
print(dataset, datasets[dataset]['fileNameRegex'])
# # Finalize Dataset XML Fragment
#
# Now, we:
#
# * set the `datasetID` we want to use
# * parse the output of `GenerateDatasetsXml.sh` into an XML tree data structure
# * set the `datasetID` dataset attribute value
# * re-set the `fileNameRegex` dataset attribute value because it looses its `\` characters during parsing(?)
# * edit and add dataset attributes from the `metadata` dict
# * set the `title` and `summary` dataset attributes from the `datasets` dict
# +
parser = etree.XMLParser(remove_blank_text=True)
tree = etree.parse('/results/erddap/logs/GenerateDatasetsXml.out', parser)
root = tree.getroot()
datasetID = 'ubcSSf2DWaveFields30mV17-02'
update_xml(root, datasetID, metadata, datasets)
# -
# Inspect the resulting dataset XML fragment below and edit the dicts and
# code cell above until it is what is required for the dataset:
print_tree(root)
# Store the XML fragment for the dataset:
with open('/results/erddap-datasets/fragments/{}.xml'.format(datasetID), 'wb') as f:
f.write(etree.tostring(root, pretty_print=True))
# Edit `/results/erddap-datasets/datasets.xml` to include the
# XML fragment for the dataset that was stored by the above cell.
#
# That file is symlinked to `/opt/tomcat/content/erddap/datasets.xml`.
#
# Create a flag file to signal the ERDDAP server process to load the dataset:
# ```
# $ cd /results/erddap/flag/
# $ touch <datasetID>
# ```
#
# If the dataset does not appear on https://salishsea.eos.ubc.ca/erddap/info/,
# check `/results/erddap/logs/log.txt` for error messages from the dataset load process
# (they may not be at the end of the file because ERDDAP is pretty chatty).
#
# Once the dataset has been successfully loaded and you are happy with the metadata
# that ERDDAP is providing for it,
# commit the changes in `/results/erddap-datasets/` and push them to Bitbucket.
| WWatch3_datasets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import re
import matplotlib.pyplot as plt
from math import log10
regex = re.compile(r"(\d+):\d+:\s+\d+\.\d+:(\d+\.?(\d+)?)")
# +
def read_from_txt(txt):
x_lst = []
y_lst = []
budget_acc_lst = []
step = 0
with open(txt) as fp:
for line in fp:
line = line.rstrip('\n')
r = re.search(regex, line)
if r is not None:
x_lst.append(log10(float(r.group(1))))
y_lst.append(float(r.group(2)))
return x_lst, y_lst
theta_lst = [0.2,0.25,0.3,0.35,0.4,0.45,0.5]
x_obs_lst, x_ref_lst, y_obs_lst, y_ref_lst = [], [], [], []
for theta in theta_lst:
x_obs, y_obs = read_from_txt('monte_carlo_sampling/MCCS_{}_obs.txt'.format(theta))
x_ref, y_ref = read_from_txt('monte_carlo_sampling/MCCS_{}_ref.txt'.format(theta))
x_obs_lst.append(x_obs)
x_ref_lst.append(x_ref)
y_obs_lst.append(y_obs)
y_ref_lst.append(y_ref)
for i in range(len(theta_lst)):
x_obs, x_ref, y_obs, y_ref = x_obs_lst[i], x_ref_lst[i], y_obs_lst[i], y_ref_lst[i]
theta = theta_lst[i]
plt.plot(x_obs, y_obs, marker='.', markersize=8, label=r'$\theta={}$'.format(theta))
#plt.plot(x_ref[:20], y_ref[:20], 'bo-', label=r'$\theta={}$'.format(theta))
plt.legend(loc='best', ncol=len(theta_lst), shadow=True,
facecolor='white', framealpha=1,prop={'size': 5.5}, borderaxespad=0, frameon=False)
plt.xlabel(r'$log_{10}(N)$')
plt.ylabel(r'$MCCS(I_{\mathcal{R}_{obs}},G)$')
plt.savefig('MCCS.pdf', bbox_inches='tight', dpi=800, pad_inches=0)
plt.show()
plt.close()
# +
import numpy as np
import operator
cmap = plt.get_cmap('gnuplot')
colors = [cmap(i) for i in np.linspace(0, 1, len(theta_lst))]
for i in range(len(theta_lst)):
x_obs, x_ref, y_obs, y_ref = x_obs_lst[i], x_ref_lst[i], y_obs_lst[i], y_ref_lst[i]
theta = theta_lst[i]
y_gap = list(map(operator.sub, y_obs, y_ref[:20]))
plt.plot(x_obs, y_gap, label=r'$\theta={}$'.format(theta), marker='.', markersize=8)
plt.legend(loc='best', ncol=len(theta_lst), shadow=True,
facecolor='white', framealpha=1,prop={'size': 5.5}, borderaxespad=0, frameon=False)
plt.xlabel(r'$log_{10}(N)$')
plt.ylabel(r'$MCCS(I_{\mathcal{R}_{obs}},G)-MCCS(I_{\mathcal{R}_{ref}},G)$')
plt.savefig('MCCS_GAP.pdf', bbox_inches='tight', dpi=800, pad_inches=0)
plt.show()
plt.close()
| stylegan/sampling/obsolete/plot_MCCS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="VHc1eQ87cTXC"
# <a href="https://colab.research.google.com/github/adasegroup/ML2021_seminars/blob/master/seminar7/seminar_GB.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="8SBTW7jZcTXF"
# # Seminar: Gradient Boosting
# Course: Machine Learning by professor <NAME>
# <br>
# Author: <NAME>
# + [markdown] colab_type="text" id="duezzdlyhUqS"
# ### The problem statement
#
# + [markdown] colab_type="text" id="_rZVv6TzdvZ4"
# The solution is found in the form of sum over random trees $h_m(x)$,
# $$F(x) = \sum_{m=1}^{M} h_m(x).$$
#
# The additive model is built in a greedy fashion:
# $$f_m(x) = f_{m-1}(x) + h_m(x).$$
#
# Having loss function $L(y, f)$, we find every new tree from the optimization
# $$h_m = \arg\min_{h} \sum_{i=1}^{n} L(y_i, f_{m-1}(x_i) + h(x_i)).$$
#
#
#
#
#
#
#
#
# + [markdown] colab_type="text" id="PblcHpTGhbqq"
# ### How the problem is solved
#
# + [markdown] colab_type="text" id="bJY6lpLQiGLX"
# Linear approximation of loss function $L(y, f)$ and the gradient descent method:
#
# $$\gamma_m = \arg\min_{\gamma} \sum_{i=1}^{n} L(y_i, f_{m-1}(x_i)
# - \gamma \frac{\partial L(y_i, f_{m-1}(x_i))}{\partial f_{m-1}(x_i)}).$$
#
# A random tree $h(x)$ is fit to targets that are the gradients $\quad -\frac{\partial L(y_i, f_{m-1}(x_i))}{\partial f_{m-1}(x_i)}.$
#
# A new tree is added to the approximation with optimal $\gamma_m$ and additional shrinkage $\nu$:
#
# $$f_m(x) = f_{m-1}(x) + \nu \gamma_m h_m(x).$$
#
# The initial model $f_0(x)$ is problem specific, for least-squares regression one usually chooses the mean of the target values.
#
#
# + colab={} colab_type="code" id="J-hogeRtcTXH"
# !pip install catboost
# !pip install xgboost
# !pip install -U scikit-learn
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.tree import DecisionTreeRegressor
# + [markdown] colab_type="text" id="dq6D_NWqcTXL"
# ## Example 1: Gradient Boosting for regression
# + colab={} colab_type="code" id="MW_HdsyncTXM"
def get_dataset_1d():
# prepare dataset
n = 1 # number of features
N = 100**n # number of samples
np.random.seed(0)
X = np.random.random((N, n))*3
coeffs = 1 + 2 * np.random.random((n, 1))
y = np.sin(np.matmul(X*X, coeffs)) + np.random.random((N, 1))/3
y = y.ravel()
return (X, y)
def plot_results(X, y, y_pred, title=''):
plt.plot(X, y, '*b')
plt.plot(X, y_pred, '.r')
plt.title(title)
plt.xlabel('x1')
plt.ylabel('y')
plt.show()
# + [markdown] colab_type="text" id="mdF0Vb9vcTXP"
#
# ---
#
# ### Question 1.
# In the following example a Gradient Boosting regression is performed by only 1 tree and very small shrinkage `nu=1e-10`. The solution looks like a constant.
#
# 1.1. Why it looks like a constant?
# <br>
# 1.2. Try to find that constant value taking into account that `loss='ls'` and draw it on the same plot.
# + colab={"base_uri": "https://localhost:8080/", "height": 279} colab_type="code" id="iUQSSADRcTXQ" outputId="10d4c746-9ef2-4ef4-c3d3-a5bfbbb4cf65"
X, y = get_dataset_1d()
clf = GradientBoostingRegressor(loss='lad', max_depth=1, learning_rate=1e-10, n_estimators=1) # learning rate = nu
clf.fit(X, y)
y_pred = clf.predict(X)
plot_results(X, y, y_pred)
# + [markdown] colab_type="text" id="4z6MXN9TcTXT"
#
# ---
#
# ### Question 2.
# Solve above question for `loss='lad'`.
# + [markdown] colab_type="text" id="H0x2u10TcTXV"
#
# ---
#
# ### Question 3.
# Some managers of industrial Data Science projects said me that each tree in Gradient Boosting is fit to the targets that are simply the differences $y_i-f_{m-1}(x_i)$ between the values $y_i$ and the current approximation $f_{m-1}(x_i)$ found on the previous step $m-1$. When is it correct?
# + [markdown] colab_type="text" id="SdD2dJaLcTXX"
#
# ---
#
# ### Question 4*.
# Actually, the managers from the question above said not only to what the trees are fit but also that the current solution is updated by simply adding each new tree multiplied by a shrinkage parameter and ignore any sophisticated math! :[]
#
# The following code shows 2 ways of using the Gradient Boosting for regression:
# 1. Using the class `GradientBoostingRegressor()` - as it should be done if you use scikit-learn
# 2. Our own implementation by adding a tree step by step with shrinkage
#
# Ensure that the pictures are the same no matter how you change the hyperparameters `max_depth`, `n_estimators` and `nu`!
#
# In our implementation we fit each tree to the targets $y_i-f_{m-1}(x_i)$ (if you answered on the above question you know that it is correct here) multiply by shrinkage $\nu$ and add to the current model. But it seems that something is missing in our code. No matter how exactly the Gradient Boosting is implemented in scikit-learn, there are many slightly different variants. Assume that it is as described here https://scikit-learn.org/stable/modules/ensemble.html#mathematical-formulation, where the steepest descent chooses the optimal step length
# $$
# \gamma_m = \arg\min_{\gamma} \sum_{i=1}^{n}
# L
# \left(
# y_i, f_{m-1}(x_i)
# - \gamma \nabla_F L(y_i, f_{m-1}(x_i))
# \right),
# $$
# which is used for the model update
# $$
# f_m(x) = f_{m-1}(x) - \gamma_m \sum_{i=1}^{n} \nabla_F L(y_i, f_{m-1}(x_i)).
# $$
# But we did not implement it! Justify the correctness of our easy implementation of GB.
# + colab={"base_uri": "https://localhost:8080/", "height": 573} colab_type="code" id="zWi2oLtQcTXY" outputId="b72cb3b6-17ea-46dd-9b2d-b8890afeca05"
X, y = get_dataset_1d()
max_depth = 1
n_estimators = 50
nu = 1 # 0.8 # shrinkage
# usual Gradient Boosting call
clf = GradientBoostingRegressor(loss='ls', max_depth=max_depth, learning_rate=nu, n_estimators=n_estimators)
clf.fit(X, y)
f = clf.predict(X)
plot_results(X, y, f, 'Gradient Boosting from scikit-learn')
# my Gradient Boosting implementation
clf = DecisionTreeRegressor(max_depth=max_depth)
f = np.mean(y) # initialization
for m in range(1, n_estimators+1):
f = f + clf.fit(X, y - f).predict(X) * nu # fit to the difference and shrink
plot_results(X, y, f, 'My simple Gradient Boosting, the same as above!')
# + [markdown] colab_type="text" id="JJK1f9XDcTXb"
# ## Example 2: Gradient Boosting for classification, CatBoost.
#
# Lets consider Titanic data! Below is yet another example of easy feature engineering, data preprocessing and the application of Gradient Boosting
# + colab={} colab_type="code" id="Bu59fC0hcTXb"
## load data and some easy preprocessing, even Feature Engineering
# we use only training dataset because the Kaggle's testing one does not have class labels
# and so we can not measure the model quality
X_train = pd.read_csv('https://raw.githubusercontent.com/adasegroup/ML2021_seminars/master/seminar7/data/train.csv')
y_train = X_train['Survived']
X_train = X_train.drop(['PassengerId', 'Survived', 'Name', 'Ticket'], axis=1)
# take only the first letter from the Cabin number, which is maybe the ship level...
def keep_only_level_of_cabin(X):
idx = X['Cabin'].notnull()
X.loc[idx, 'Cabin'] = [s.strip()[0] for s in X['Cabin'][idx].values]
return X
# CatBoost can not process categorical features with NaN, we set them to the string 'MISS'
def change_NaN_to_str(X, cat_features_cols):
for col in cat_features_cols:
idx = X[col].isnull()
X.loc[idx, col] = 'MISS'
return X
X_train = keep_only_level_of_cabin(X_train)
X_train = change_NaN_to_str(X_train, ['Sex', 'Cabin', 'Embarked'])
X_train.head()
# + [markdown] colab_type="text" id="EK6KVUekcTXg"
# ### Note:
#
# 1. We use categorical features and CatBoost processes it ok.
#
#
# 2. Below you can see the tuning of CatBoost with cross-validation, you can play with the hyperparameter ranges.
#
#
# 3. You can "unnatural disable" cross-validation and ensure that the best score increases (although it is impractical because of overfitting).
#
#
# 4. CatBoost processes contineous feature 'Age' despite there are NaNs. It is easy for trees - we can, say, just ignore these values.
#
#
# 5. We preprocess the 'Cabin' feature so that it has only the level ('C', 'E', ...).
# This is so that there would not be a large number of unique noninformative values. And here we process NaNs as special 'MISS' value because it may mean not the incompletness in the data, but that no any proper value can be for some samples (it is highly correlated with 'Pclass' feature).
#
#
# 6. You can play with feature importances: drop the most important feature and check how the accuracy and the roc_auc reduce. Do the same with the least important feature and feel who affects more.
# + colab={"base_uri": "https://localhost:8080/", "height": 977} colab_type="code" id="KH9MlZSjcTXh" outputId="90ef074f-b000-4800-af50-e5ad2d2c3cd7"
from catboost import CatBoostClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import roc_curve
clf = CatBoostClassifier(loss_function = 'Logloss',
# categorical features processing:
cat_features = np.where(X_train.columns.isin(['Sex', 'Cabin', 'Embarked']))[0],
verbose = 0, thread_count=1, random_state=0)
clf = GridSearchCV(clf,
# you can play with tuning, up to your CPU performance:
{'max_depth': [1, 3, 5], 'n_estimators': [100], 'learning_rate': [0.1, 1]},
# cross-validation prevents overfitting! you can disable it by setting
# cv = [(np.array(range(len(y_train))), np.array(range(len(y_train))))]
cv = 3, scoring='roc_auc', n_jobs=-1)
clf.fit(X_train, y_train)
print('best params:', clf.best_params_)
print('best score:', clf.best_score_) # validation score
# feature importances
fi = pd.Series(clf.best_estimator_.feature_importances_, index=X_train.columns)
fi.sort_values(ascending=False).plot(kind='bar')
plt.title('feature importances')
plt.show()
# now let's draw different ROC curves
plt.figure(figsize=[9, 6])
fpr, tpr, _ = roc_curve(y_train, clf.predict_proba(X_train)[:, 1])
plt.plot(fpr, tpr, 'r', label='train')
fpr, tpr, _ = roc_curve(y_train, clf.predict(X_train))
plt.plot(fpr, tpr, '--o', label='binary output, train')
plt.legend(bbox_to_anchor=(0.999, 1))
plt.title('ROC curves')
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
plt.show()
# -
# ## Example 3: Gradient Boosting Libraries: CatBoost (above), XGBoost.
# +
from sklearn.preprocessing import OneHotEncoder
X_train = pd.read_csv('https://raw.githubusercontent.com/adasegroup/ML2021_seminars/master/seminar7/data/train.csv')
y_train = X_train['Survived']
X_train = X_train.drop(['PassengerId', 'Survived', 'Name', 'Ticket'], axis=1)
X_train = keep_only_level_of_cabin(X_train)
# there are also NaN values!
print('Sex:', X_train['Sex'].unique(), ', Cabin:', X_train['Cabin'].unique(), ', Embarked:', X_train['Embarked'].unique())
X_train['Sex'] = X_train['Sex'] == 'male' # binary feature - its easy!
# use OneHotEncoder to work with categorial features:
ohe = OneHotEncoder(sparse=False, handle_unknown='ignore', categories=[['C', 'E', 'G', 'D', 'A', 'B', 'F', 'T'], ['S', 'C', 'Q']])
new_features = pd.DataFrame(ohe.fit_transform(X_train[['Cabin', 'Embarked']]),
columns = ohe.get_feature_names(['Cabin', 'Embarked']), index=X_train.index)
X_train = pd.concat((X_train, new_features), axis=1, sort=False).drop(['Cabin', 'Embarked'], axis=1)
X_train.head()
# +
import xgboost.sklearn as xgb
clf = GridSearchCV(xgb.XGBClassifier(loss = 'binary:logistic', nthread=1, random_state=0),
# you can play with tuning, up to your CPU performance:
{'max_depth': [1, 3, 5], 'n_estimators': [100], 'learning_rate': [0.1, 1]},
# cross-validation prevents overfitting! you can disable it by setting
# cv = [(np.array(range(len(y_train))), np.array(range(len(y_train))))]
cv = 3, scoring='roc_auc', n_jobs=-1)
clf.fit(X_train, y_train)
print('best params:', clf.best_params_)
print('best score:', clf.best_score_) # validation score
# feature importances
fi = pd.Series(clf.best_estimator_.feature_importances_, index=X_train.columns)
fi.sort_values(ascending=False).plot(kind='bar')
plt.title('feature importances')
plt.show()
# now let's draw different ROC curves
plt.figure(figsize=[9, 6])
fpr, tpr, _ = roc_curve(y_train, clf.predict_proba(X_train)[:, 1])
plt.plot(fpr, tpr, 'r')
plt.title('ROC curve')
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
plt.show()
# -
| seminar7/seminar_GB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
from run_bnn import run_bnn, bnn_experiment
from run_sbi import run_snpe, sbi_experiment
import numpy as np
import torch
# # Run ABC-SMC (Reproducibility)
# run the script run_abc.py to generate all the necessary data from the ABC-SMC sampler. Alternatively one can directly discover and tune the notebook: abc_smc.ipynb.
#
# Will compute and store the following files
# * smcabc_post_10gen - posterior samples
# * smcabc_trails_10gen.npy - number of simulated proposals
#
# Obs ABC-SMC requre many proposals, therefore this takes some time
#obs to stop the script, interupt the kernel several times...
# %run run_abc.py
# ## Run SNPE-C (Reproducibility)
# run the SNPE-C (sbi package) script.
#
# ### Generating the following files
# * data/sbi_data_post.npy - posterior samples
# * data/sbi_data_time.npy - elapsed times
sbi_experiment()
# # Custom SNPE (Testing)
# For testing the script with fewer runs and other parameters. Obs that these results might not work for all plots in plotting.ipynb
ID = 'data'
sbi_post, sbi_time, sbi_post_object = run_snpe(total_runs=1, num_generation=6, seed=2, nde='maf')
#Obs need "data" folder
np.save(f'{ID}/sbi_{ID}_post', sbi_post)
np.save(f'{ID}/sbi_{ID}_time', sbi_time)
#Take DirectPosterior to get the NN
test = sbi_post_object[0]
#count the number of trainable weights/parameters
def count_parameters(model):
total_params = 0
for name, parameter in model.named_parameters():
if not parameter.requires_grad: continue
param = parameter.numel()
total_params+=param
print(name,':', param)
print(f"Total Trainable Params: {total_params}")
count_parameters(test.net)
# # Run BNN (Reproducibility)
#
#
# ### The following files are produced
# posterior samples from 4 bins per parameter, repeated 10 times, a 6 step iteration, with 0.05 threshold. With the time elapsed.
# * data/bcnn_data_post.npy' - posterior samples
# * data/bcnn_data_time.npy' - time elapsed
#
# posterior samples from 3 bins per parameter, repeated 10 times, a 6 step iteration, with 0.05 threshold.
# * data/bcnn_data_bins3_post.npy - posterior samples
#
# posterior samples from 5 bins, repeated 10 times, a 6 step iteration, with 0.05 threshold.
# * data/bcnn_data_bins5_post.npy - posterior samples
#
# posterior samples from 4 bins, repeated 10 times, a 6 step iteration, with no threshold.
# * data/bcnn_data_no_thresh_post.npy - posterior samples
#
# posterior samples from 4 bins, repeated 10 times, a 6 step iteration, with exponential decreasing threshold.
# * data/bcnn_data_exp_thresh_post.npy - posterior samples
# +
import tensorflow as tf
#in case CUDA is causing problems...
tf.config.set_visible_devices([], 'GPU')
# -
bnn_experiment()
# # Custom BNN (Testing)
# For testing the script with fewer runs and other parameters.
# Obs that these results might not work for all plots in plotting.ipynb
ID = 'data'
bcnn_post, bcnn_proposals, bcnn_time = run_bnn(total_runs=1, num_rounds=6, seed=3,
ID=ID)
# without correction
ID = 'data'
bcnn_post_nocorr, bcnn_proposals_nocorr, bcnn_time_nocorr = run_bnn(total_runs=1, num_rounds=6, seed=3,
ID=ID, correction=False)
# +
import matplotlib.backends.backend_pdf
import matplotlib.pyplot as plt
from tensorflow_probability import distributions as tfd
import scipy
#get true posterior from mcmc run
subset_exact_samples = np.load('exact_mcmc_post.npy')
# -
def plot_conv(proposals):
for theta_small in proposals:
f = plt.figure(figsize=(15, 10), constrained_layout=True)
gs = f.add_gridspec(3, 5)
#BCNN KDE
def multivar(grid, m, var, xlabel='', ylabel=''):
ax = f.add_subplot(grid)
x, y = np.mgrid[-2:2:.01, -1:1:.01]
pos = np.dstack((x, y))
rv = tfd.MultivariateNormalFullCovariance(loc=m,
covariance_matrix=var)
ax.contourf(x, y, rv.prob(pos))
ax.set_xlim(-2,2)
ax.set_ylim(-1,1)
ax.set_yticks([])
ax.set_xticks([])
return ax
#BCNN
mcmc_mean = subset_exact_samples.mean(axis=0)
c=0
c1 = 0
for i, res in enumerate(theta_small):
m = res[0]
var = res[1]
if i < 4:
ax = multivar(gs[0,i+1],m, var, xlabel=f'round {i+1}')
ax.scatter(mcmc_mean[0], mcmc_mean[1], color='C3')
elif i < 9:
ax = multivar(gs[1,c],m, var, xlabel=f'round {i+1}')
ax.scatter(mcmc_mean[0], mcmc_mean[1], color='C3')
c+=1
elif i < 15:
ax = multivar(gs[2,c1],m, var, xlabel=f'round {i+1}')
ax.scatter(mcmc_mean[0], mcmc_mean[1], color='C3')
c1+=1
#MCMC Gaussian approx
cov = np.cov(subset_exact_samples, rowvar=0)
mean = subset_exact_samples.mean(axis=0)
x, y = np.mgrid[-2:2:.01, -1:1:.01]
pos = np.dstack((x, y))
rv = scipy.stats.multivariate_normal(mean, cov)
ax = f.add_subplot(gs[0, 0])
ax.contourf(x, y, rv.pdf(pos))
#ax2.scatter(target_theta[:,0],target_theta[:,1], color="red", label="true")
ax.set_xlim(-2,2)
ax.set_ylim(-1,1)
ax.set_yticks([])
ax.set_xticks([])
ax.set_xlabel('Gaussian fit')
ax.set_ylabel('MCMC (true)')
pdf.savefig(f)
pdf.close()
pdf = matplotlib.backends.backend_pdf.PdfPages("regression_withcorrection.pdf")
plot_conv(bcnn_proposals)
pdf = matplotlib.backends.backend_pdf.PdfPages("regression_nocorrection.pdf")
plot_conv(bcnn_proposals_nocorr)
#Obs need "data" folder
np.save(f'{ID}/bcnn_{ID}_post', bcnn_post)
np.save(f'{ID}/bcnn_{ID}_time', bcnn_time)
| MA2/experiments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 파이썬개발자[6] 데이터엔지니어[7] 데이터사이언티스트[10] 머신러닝엔지니어[15] 빅데이터엔지니어[16]
category = [6, 7, 10, 15, 16]
# +
import sys
mod = sys.modules[__name__]
for category_num in range(18):
data = open('../vocabulary_all_1211/over_del_tokens__/tokens{}__.txt'.format(category_num), 'r', encoding='utf-8')
token = data.readlines()
print("#", category_num, "번째 category (tokens):", len(token))
for i in range(len(token)):
token[i] = token[i].replace('\n', '')
setattr(mod, 'tokens{}'.format(category_num), token)
#voca로도 저장해줍니다, 리스트 형태로 바꿔서 저장해줍시다
voca = list(set(token))
setattr(mod, 'voca{}'.format(category_num), voca)
print("# voca size:", len(voca), '\n')
# +
import nltk
for category_num in category:
tokens = globals()['tokens{}'.format(category_num)]
print('category{} :'.format(category_num), nltk.FreqDist(tokens).most_common(5))
setattr(mod, 'freqdist{}'.format(category_num), nltk.FreqDist(tokens).most_common(15))
# -
for category_num in category:
print('# token 빈도수\n')
FD = globals()['freqdist{}'.format(category_num)]
for i in FD:
print(i[0], ' : ', i[1])
print('\n')
| analysis/FreqDist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Вопросы для повторения**:
#
# 1. для чего вводили рэнжи?
# 2. какие билиотеки рэнжей мы смотрели?
# 3. чем view отличается от action?
# 4. какая здесь проблема?
#
# ```c++
# auto make_some_event_numbers()
# {
# std::vector<int> v = /* some numbers here */;
# return v | rv::filter(is_even);
# }
# ```
#
# 5. что делает функция? как называется фича?
#
# ```c++
# struct Person
# {
# std::string surname;
# std::string name;
# };
#
# void f(std::vector<Person>& people)
# {
# rs::sort(people, rg::less{}, &Person::name);
# }
# ```
#
# 6. что делает этот код и в чём его проблема? как чинить?
#
# ```c++
# for (const auto& person : people_ids | rv::transform(get_person_from_database_by_id)
# | rv::filter(is_student))
# {
# std::cout << person.name << ", stay home" << std::endl;
# }
# ```
#
# 7. Какие плюсы и минусы ренжей, когда их использовать скорее следует?
#
# 8. Работает ли код? Какие ошибки?
#
# ```c++
# using Task = std::function<void>;
#
# class TasksExecutor
# {
# private:
# bool is_stopped = false;
# std::queue<Task> tasks;
#
# public:
# TasksExecutor()
# {
# std::thread([this](){
# while (!is_stopped) { /* process tasks from queue */ }
# }).detach();
# }
#
# ~TasksExecutor() {
# is_stopped = true;
# }
#
# void stop() { is_stopped = true; }
#
# void push_task(Task&& task) { /* push task to queue */ }
# };
# ```
#
# <details>
# <summary>подсказка</summary>
# <p>
#
# * race condition по `is_stopped`
# * захват `this` + `detach` - хождение по разрушенному объекту после отработки `~TasksExecutor()`
#
# </p>
# </details>
| 2020/sem2/lecture_8_modern_cpp_misc_features/repetition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
import nltk
import re
import numpy as np
import tensorflow as tf
from prepro import convert_idx
with open('/home/soham/data/squad/sample1k-HCVerifyAll.json', 'r') as f:
foo = json.load(f)
meh = []
train_pairs = np.load('final_train_pairs.npy').tolist()
test_pairs = np.load('final_test_pairs.npy').tolist()
adv_count = lambda x: sum(map(lambda y: 1 if y[-2] == 'adv' else 0, x))
print('Train:', '%d,' % len(train_pairs), adv_count(train_pairs), 'of which are adversarial')
print('Test:', '%d,' % len(test_pairs), adv_count(test_pairs), 'of which are adversarial')
import spacy
nlp = spacy.blank("en")
def word_tokenize(sent):
doc = nlp(sent)
return [token.text for token in doc]
# +
def _get_word(word):
for each in (word, word.lower(), word.capitalize(), word.upper()):
if each in word2idx_dict:
return word2idx_dict[each]
return 1
def _get_char(char):
if char in char2idx_dict:
return char2idx_dict[char]
return 1
# +
with open('data/word2idx.json', 'r') as fh:
word2idx_dict = json.load(fh)
with open('data/char2idx.json', 'r') as fh:
char2idx_dict = json.load(fh)
# -
para_limit = 1000
ques_limit = 100
char_limit = 16
import random
random.shuffle(train_pairs)
random.shuffle(test_pairs)
# %rm data/badptr_t*.tf
# +
writer = tf.python_io.TFRecordWriter('data/badptr_train.tf')
for id_, (id_but_ignore, context, ques, ans, start_end, tag, title) in enumerate(train_pairs):
context = context.replace("''", '" ').replace("``", '" ')
context_tokens = word_tokenize(context)
context_chars = [list(token) for token in context_tokens]
ques = ques.replace("''", '" ').replace("``", '" ')
ques_tokens = word_tokenize(ques)
ques_chars = [list(token) for token in ques_tokens]
context_idxs = np.zeros([para_limit], dtype=np.int32)
context_char_idxs = np.zeros([para_limit, char_limit], dtype=np.int32)
ques_idxs = np.zeros([ques_limit], dtype=np.int32)
ques_char_idxs = np.zeros([ques_limit, char_limit], dtype=np.int32)
for i, token in enumerate(context_tokens):
context_idxs[i] = _get_word(token)
for i, token in enumerate(ques_tokens):
ques_idxs[i] = _get_word(token)
for i, token in enumerate(context_chars):
for j, char in enumerate(token):
if j == char_limit:
break
context_char_idxs[i, j] = _get_char(char)
for i, token in enumerate(ques_chars):
for j, char in enumerate(token):
if j == char_limit:
break
ques_char_idxs[i, j] = _get_char(char)
### NEW PROC GOES HERE
y1 = np.zeros([para_limit], dtype=np.float32)
y2 = np.zeros([para_limit], dtype=np.float32)
if start_end is None:
y1[0] = 1
y2[0] = 1
else:
spans = convert_idx(context, context_tokens)
start, end = start_end
answer_span = []
for idx, span in enumerate(spans):
if not (end <= span[0] or start >= span[1]):
answer_span.append(idx)
y1[answer_span[0]] = 1
y2[answer_span[-1]] = 1
record = tf.train.Example(features=tf.train.Features(feature={
"context_idxs": tf.train.Feature(bytes_list=tf.train.BytesList(value=[context_idxs.tostring()])),
"ques_idxs": tf.train.Feature(bytes_list=tf.train.BytesList(value=[ques_idxs.tostring()])),
"context_char_idxs": tf.train.Feature(bytes_list=tf.train.BytesList(value=[context_char_idxs.tostring()])),
"ques_char_idxs": tf.train.Feature(bytes_list=tf.train.BytesList(value=[ques_char_idxs.tostring()])),
"id": tf.train.Feature(int64_list=tf.train.Int64List(value=[id_])),
"y1": tf.train.Feature(bytes_list=tf.train.BytesList(value=[y1.tostring()])),
"y2": tf.train.Feature(bytes_list=tf.train.BytesList(value=[y2.tostring()])),
}))
writer.write(record.SerializeToString())
writer.close()
# -
from prepro import save
# +
writer = tf.python_io.TFRecordWriter('data/badptr_test.tf')
classic_meta_info = {}
meta_info = {}
for id_, (id_but_ignore, context, ques, ans, start_end, tag, title) in enumerate(test_pairs):
context = context.replace("''", '" ').replace("``", '" ')
context_tokens = word_tokenize(context)
context_chars = [list(token) for token in context_tokens]
ques = ques.replace("''", '" ').replace("``", '" ')
ques_tokens = word_tokenize(ques)
ques_chars = [list(token) for token in ques_tokens]
context_idxs = np.zeros([para_limit], dtype=np.int32)
context_char_idxs = np.zeros([para_limit, char_limit], dtype=np.int32)
ques_idxs = np.zeros([ques_limit], dtype=np.int32)
ques_char_idxs = np.zeros([ques_limit, char_limit], dtype=np.int32)
for i, token in enumerate(context_tokens):
context_idxs[i] = _get_word(token)
for i, token in enumerate(ques_tokens):
ques_idxs[i] = _get_word(token)
for i, token in enumerate(context_chars):
for j, char in enumerate(token):
if j == char_limit:
break
context_char_idxs[i, j] = _get_char(char)
for i, token in enumerate(ques_chars):
for j, char in enumerate(token):
if j == char_limit:
break
ques_char_idxs[i, j] = _get_char(char)
### NEW PROC GOES HERE
y1 = np.zeros([para_limit], dtype=np.float32)
y2 = np.zeros([para_limit], dtype=np.float32)
spans = convert_idx(context, context_tokens)
if start_end is None:
y1[0] = 1
y2[0] = 1
else:
start, end = start_end
answer_span = []
for idx, span in enumerate(spans):
if not (end <= span[0] or start >= span[1]):
answer_span.append(idx)
y1[answer_span[0]] = 1
y2[answer_span[-1]] = 1
if start_end is None:
new_answer_ = context[spans[0][0]:spans[0][1]]
else:
new_answer_ = context[spans[answer_span[0]][0]:spans[answer_span[-1]][1]]
meta_info[id_] = {'answers': [new_answer_], 'spans': spans, 'context': context, 'uuid': id_but_ignore}
classic_meta_info[id_] = {'answers': ans, 'spans': spans, 'context': context, 'ques': ques, 'tag': tag, 'title': title, 'id': id_}
record = tf.train.Example(features=tf.train.Features(feature={
"context_idxs": tf.train.Feature(bytes_list=tf.train.BytesList(value=[context_idxs.tostring()])),
"ques_idxs": tf.train.Feature(bytes_list=tf.train.BytesList(value=[ques_idxs.tostring()])),
"context_char_idxs": tf.train.Feature(bytes_list=tf.train.BytesList(value=[context_char_idxs.tostring()])),
"ques_char_idxs": tf.train.Feature(bytes_list=tf.train.BytesList(value=[ques_char_idxs.tostring()])),
"id": tf.train.Feature(int64_list=tf.train.Int64List(value=[id_])),
"y1": tf.train.Feature(bytes_list=tf.train.BytesList(value=[y1.tostring()])),
"y2": tf.train.Feature(bytes_list=tf.train.BytesList(value=[y2.tostring()])),
}))
writer.write(record.SerializeToString())
writer.close()
save('data/badptr_test_meta.json', meta_info, message='meta_info')
np.save('data/badptr_test_meta.npy', classic_meta_info)
# -
len(set([a for a in map(lambda x: x['uuid'], meta_info.values())]))
| 3 Data Processing (Bad Pointers).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Summary
#
# Use [`python-fitbit`](http://python-fitbit.readthedocs.io/en/latest/#) to interact with the Fitbit API and download sleep data. Convert data to pandas dataframe and write to disk.
# +
import fitbit
import os
import pandas as pd
from tqdm import tqdm
from dotenv import load_dotenv, find_dotenv
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
sns.set_context('poster')
# +
# find .env automagically by walking up directories until it's found
dotenv_path = find_dotenv()
# load up the entries as environment variables
load_dotenv(dotenv_path)
client_id = os.environ.get("CLIENT_ID")
client_secret = os.environ.get("CLIENT_SECRET")
access_token = os.environ.get("ACCESS_TOKEN")
refresh_token = os.environ.get("REFRESH_TOKEN")
expires_at = float(os.environ.get("EXPIRES_AT"))
# -
authd_client = fitbit.Fitbit(client_id, client_secret, access_token=access_token,
refresh_token=refresh_token, expires_at=expires_at)
# +
sleep_logs = []
rate_limit = 145
datetimes = pd.date_range('2016-11-09', '2017-07-31', freq='1D')
for datetime in tqdm(datetimes[:rate_limit]):
sleep_log = authd_client.get_sleep(datetime)
sleep_logs.append(sleep_log)
# -
for datetime in tqdm(datetimes[rate_limit:]):
sleep_log = authd_client.get_sleep(datetime)
sleep_logs.append(sleep_log)
len(sleep_logs)
sleep_logs[-1]
sleep_keys = sleep_logs[-1]['sleep'][0].keys()
sleep_keys
trimmed_list = []
for sleep_log in sleep_logs:
for sleep_event in sleep_log['sleep']:
trim_dict = {}
key_list = sleep_event.keys()
for this_key in key_list:
if this_key != 'minuteData':
trim_dict[this_key] = sleep_event[this_key]
trimmed_list.append(trim_dict)
df_sleep = pd.DataFrame(trimmed_list)
df_sleep.head()
df_sleep.columns
df_sleep['dateOfSleep'].head()
start_sleep = pd.to_datetime(df_sleep['startTime'])
df_sleep['start_datetime'] = start_sleep
# df_sleep = df_sleep.set_index('start_datetime')
# Sum from 6pm to 6pm instead of midnight to midnight.
df_sleep['shifted_datetime'] = df_sleep['start_datetime'] - pd.Timedelta(hours=6)
df_sleep = df_sleep.set_index('shifted_datetime')
plt.plot(df_sleep['minutesAsleep'].resample('1D').sum()/60., '.')
hours_sleep = df_sleep['minutesAsleep'].resample('1D').sum()/60.
hours_sleep.mean()
hours_sleep.std()
os.getcwd()
data_path = os.path.join(os.getcwd(), os.pardir, 'data', 'interim', 'sleep_data.csv')
df_sleep.to_csv(data_path, index_label='shifted_datetime')
| notebooks/sbussmann_data-sleep.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Tutorial about the LocData class
# +
import numpy as np
import pandas as pd
import locan as lc
# -
lc.show_versions(system=False, dependencies=False, verbose=False)
# ## Sample data
# A localization has certain properties such as 'position_x'. A list of localizations can be assembled into a dataframe:
df = pd.DataFrame(
{
'position_x': np.arange(0,10),
'position_y': np.random.random(10),
'frame': np.arange(0,10),
})
# ## Instantiate LocData from a dataframe
# A LocData object carries localization data together with metadata and aggregated properties for the whole set of localizations.
#
# We first instantiate a LocData object from the dataframe:
locdata = lc.LocData.from_dataframe(dataframe=df)
attributes = [x for x in dir(locdata) if not x.startswith('_')]
attributes
# ## LocData attributes
# The class attribute Locdata.count represents the number of all current LocData instantiations.
print('LocData count: ', lc.LocData.count)
# The localization dataset is provided by the data attribute:
print(locdata.data.head())
# Aggregated properties are provided by the attribute properties. E.g. the property `position_x` represents the mean of the `position_x` for all localizations. We keep the name, since the aggregated dataset can be treated as just a single locdata event with `position_x`. This is used when dealing with data clusters.
locdata.properties
# Since spatial coordinates are quite important one can check on *coordinate_labels* and dimension:
locdata.coordinate_labels
locdata.dimension
# A numpy array of spatial coordinates is returned by:
locdata.coordinates
# ## Metadata
# For detailed information see the `Tutorial about metadata`.
# Metadata is provided by the attribute meta and can be printed as
locdata.print_meta()
# A summary of the most important metadata fields is printed as:
locdata.print_summary()
# Metadata fields can be printed and changed individually:
print(locdata.meta.comment)
locdata.meta.comment = 'user comment'
print(locdata.meta.comment)
# LocData.meta.map represents a dictionary structure that can be filled by the user. Both key and value have to be strings, if not a TypeError is thrown.
print(locdata.meta.map)
locdata.meta.map['user field'] = 'more information'
print(locdata.meta.map)
# Metadata can also be added at Instantiation:
locdata_2 = lc.LocData.from_dataframe(dataframe=df, meta={'identifier': 'myID_1',
'comment': 'my own user comment'})
locdata_2.print_summary()
# ## Instantiate locdata from selection
# A LocData object can also be instantiated from a selection of localizations. In this case the LocData object keeps a reference to the original locdata together with a list of indices (or a slice object)). The new dataset is assembled on request of the data attribute.
# *Typically a selection is derived using a selection method such that using LocData.from_selection() is not often necessary.*
# +
locdata_2 = lc.LocData.from_selection(locdata, indices=[1,2,3,4])
locdata_3 = lc.LocData.from_selection(locdata, indices=[5,6,7,8])
print('count: ', lc.LocData.count)
print('')
print(locdata_2.data)
# -
locdata_2.print_summary()
# The reference is kept in a private attribute as are the indices.
print(locdata_2.references)
print(locdata_2.indices)
# The reference is the same for both selections.
print(locdata_2.references is locdata_3.references)
# ## Instantiate locdata from collection
# A LocDat object can further be instantiated from a collection of other LocData objects.
# +
del(locdata_2, locdata_3)
locdata_1 = lc.LocData.from_selection(locdata, indices=[0,1,2])
locdata_2 = lc.LocData.from_selection(locdata, indices=[3,4,5])
locdata_3 = lc.LocData.from_selection(locdata, indices=[6,7,8])
locdata_c = lc.LocData.from_collection(locdatas=[locdata_1, locdata_2, locdata_3], meta={'identifier': 'my_collection'})
print('count: ', lc.LocData.count, '\n')
print(locdata_c.data, '\n')
print(locdata_c.properties, '\n')
locdata_c.print_summary()
# -
# In this case the reference are also kept in case the original localizations from the collected LocData object are requested.
print(locdata_c.references)
# In case the collected LocData objects are not needed anymore and should be free for garbage collection the references can be deleted by a dedicated Locdata method
locdata_c.reduce()
print(locdata_c.references)
# ## Concatenating LocData objects
# Lets have a second dataset with localization data:
# +
del(locdata_2)
df_2 = pd.DataFrame(
{
'position_x': np.arange(0,10),
'position_y': np.random.random(10),
'frame': np.arange(0,10),
})
locdata_2 = lc.LocData.from_dataframe(dataframe=df_2)
print('First locdata:')
print(locdata.data.head())
print('')
print('Second locdata:')
print(locdata_2.data.head())
# -
# In order to combine two sets of localization data from two LocData objects into a single LocData object use the class method *LocData.concat*:
locdata_new = lc.LocData.concat([locdata, locdata_2])
print('Number of localizations in locdata_new: ', len(locdata_new))
locdata_new.data.head()
# ## Modifying data in place
# In case localization data has been modified in place, i.e. the dataset attribute is changed, all properties and hulls must be recomputed. This is best done by re-instantiating the LocData object using `LocData.from_dataframe()`; but it can also be done using the `LocData.reset()` function.
# +
del(df, locdata)
df = pd.DataFrame(
{
'position_x': np.arange(0,10),
'position_y': np.random.random(10),
'frame': np.arange(0,10),
})
locdata = lc.LocData.from_dataframe(dataframe=df)
print(locdata.data.head())
# -
locdata.centroid
# Now if localization data is changed in place (which you should not do unless you have a good reason), properties and bounding box are not automatically adjusted.
# +
locdata.dataframe = pd.DataFrame(
{
'position_x': np.arange(0,8),
'position_y': np.random.random(8),
'frame': np.arange(0,8),
})
print(locdata.data.head())
# -
locdata.centroid # so this returns incorrect values here
# Update them by re-instantiating a new LocData object:
locdata_new = lc.LocData.from_dataframe(dataframe=locdata.data)
locdata_new.centroid
locdata_new.meta
# Alternatively you can use `reset()`. In this case, however, metadata is not updated and will provide wrong information.
locdata.reset()
locdata.centroid
locdata.meta
# ## Copy LocData
# Shallow and deep copies can be made from LocData instances. In either case the class variable count and the metadata is not just copied but adjusted accordingly.
print('count: ', lc.LocData.count)
print('')
print(locdata_2.meta)
# +
from copy import copy, deepcopy
print('count before: ', lc.LocData.count)
locdata_copy = copy(locdata_2)
locdata_deepcopy = deepcopy(locdata_2)
print('count after: ', lc.LocData.count)
# -
print(locdata_copy.meta)
print(locdata_deepcopy.meta)
# ## Adding a property
# Any property that is created for a set of localizations (and represented as a python dictionary) can be added to the Locdata object. As an example, we compute the maximum distance between any two localizations and add that `max_distance` as new property to `locdata`.
max_distance = lc.max_distance(locdata)
max_distance
# + tags=[]
locdata.properties.update(max_distance)
locdata.properties
# -
# ## Adding a property to each localization in LocData.data
# In case you have processed your data and come up with a new property for each localization in the LocData object, this property can be added to data. As an example, we compute the nearest neighbor distance for each localization and add `nn_distance` as new property.
locdata.data
nn = lc.NearestNeighborDistances().compute(locdata)
nn.results
# To add `nn_distance` as new property to each localization in LocData object, use the `pandas.assign` function on the `locdata.dataframe`.
locdata.dataframe = locdata.dataframe.assign(nn_distance=nn.results['nn_distance'])
locdata.data
# ### Adding nn_distance as new property to each localization in LocData object with dataframe=None
# In case the LocData object was created with LocData.from_selection() the LocData.dataframe attribute is None and LocData.data is generated from the referenced locdata and the index list.
#
# In this case LocData.dataframe can still be filled with additional data that is merged upon returning LocData.data.
locdata_selection = lc.LocData.from_selection(locdata, indices=[1, 3, 4, 5])
locdata_selection.data
locdata_selection.dataframe
nn_selection = lc.NearestNeighborDistances().compute(locdata_selection)
nn_selection.results
# Make sure the indices in nn.results match those in dat_selection.data:
locdata_selection.data.index
nn_selection.results.index = locdata_selection.data.index
nn_selection.results
# Then assign the corresponding result to dataframe:
locdata_selection.dataframe = locdata_selection.dataframe.assign(nn_distance= nn_selection.results['nn_distance'])
locdata_selection.dataframe
# Calling `data` will return the complete dataset.
locdata_selection.data
| docs/tutorials/notebooks/LocData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bayesian experimental design for Kepler-36
#
# Suppose we would like to make a follow-up measurement on Kepler-36c (the more observable of the two planets in the system). How would we know the optimal time(s) to observe to decreaes uncertainties on the dynamical masses in the system? Because the sampling on TTV curves is often sparse and non-uniform, it can be difficult to quantify from first principles how much information you gain on the dynamical masses at any given epoch.
#
# Here, we use Bayesian experimental design to solve this problem. Largely following Goldberg et al. (2018), we can *simulate* what the posteriors look like after a follow-up measurement is taken by rejection sampling our original posteriors. We can then quantify the difference between the old and new posteriors using the Kullback-Leibler Divergence, and use this as a metric to assess the expected information gain at each epoch.
#
# ttvnest has a Bayesian experimental design module called 'followup' that does just that. Let's use it!
# +
# %matplotlib inline
import ttvnest
import numpy as np
kepler36 = ttvnest.io_utils.load_results('kepler36.p')
# -
# Let's check out the posterior summary and TTV plots again:
kepler36.posterior_summary()
ttvnest.plot_utils.plot_results(kepler36, uncertainty_curves = 100,
sim_length = 365.25*10)
# Now, let's do some Bayesian experimental design!
# +
measurement_uncertainty = 4./1440 #about as good as Kepler
measured_planet = 2 #one-indexed
info = ttvnest.followup.calculate_information_timeseries(
kepler36, measurement_uncertainty, measured_planet)
# -
ttvnest.plot_utils.plot_information_timeseries(info)
| examples/kepler-36/kepler-36_bayesian_experimental_design.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Click-prediction with XDeepFM
#
# (PROPOSAL: Start with what is the problem we are addressing and why the user should care)
#
# In this notebook we are going to analyze an example of collaborative filtering using the Microsoft Research algorithm xDeepFM ([Paper](https://arxiv.org/abs/1803.05170)). For it, we are going to use the [dataset CRITEO](https://www.kaggle.com/c/criteo-display-ad-challenge/data), which contains:
#
# - Label - Target variable that indicates if an ad was clicked (1) or not (0).
# - I1-I13 - A total of 13 columns of integer features (mostly count features).
# - C1-C26 - A total of 26 columns of categorical features. The values of these features have been hashed onto 32 bits for anonymization purposes.
#
# The training set consists of a portion of Criteo's traffic over a period of 7 days. Each row corresponds to a display ad served by Criteo. Positive (clicked) and negatives (non-clicked) examples have both been subsampled at different rates in order to reduce the dataset size. The examples are chronologically ordered. Label - Target variable that indicates if an ad was clicked (1) or not (0).
#
# An algorithm like xDeepFM can be beneficial for problems of click optimization, where the objective is to maximize the CTR (TODO: link to CTR info). The evaluation metrics that we are going to use are regression metrics like RMSE, AUC or logloss.
# ## xDeepFM
#
# Combinatorial features are essential for the success of many com- mercial models. Manually crafting these features usually
# comes with high cost due to the variety, volume and velocity of raw data in web-scale systems. Factorization based models,
# which measure interactions in terms of vector product, can learn patterns of com- binatorial features automatically and
# generalize to unseen features as well. With the great success of deep neural works (DNNs) in various fields, recently
# researchers have proposed several DNN- based factorization model to learn both low- and high-order feature interactions.
# Despite the powerful ability of learning an arbitrary function from data, plain DNNs generate feature interactions im-
# plicitly and at the bit-wise level. In this paper, we propose a novel Compressed Interaction Network (CIN), which aims
# to generate feature interactions in an explicit fashion and at the vector-wise level. We show that the CIN share some
# functionalities with con- volutional neural networks (CNNs) and recurrent neural networks (RNNs). We further combine a
# CIN and a classical DNN into one unified model, and named this new model eXtreme Deep Factor- ization Machine (xDeepFM).
# On one hand, the xDeepFM is able to learn certain bounded-degree feature interactions explicitly; on the other hand, it
# can learn arbitrary low- and high-order feature interactions implicitly. We conduct comprehensive experiments on three
# real-world datasets. Our results demonstrate that xDeepFM outperforms state-of-the-art models.
# %load_ext blackcellmagic
# + tags=["parameters"]
max_rows = 1000000
data_url = "https://s3-eu-west-1.amazonaws.com/kaggle-display-advertising-challenge-dataset/dac.tar.gz"
# +
import os
import sys
import collections
import csv
import math
import random
import time
from collections import defaultdict
from machine_utils import (
get_gpu_name,
get_number_processors,
get_gpu_memory,
get_cuda_version,
)
import numpy as np
import pandas as pd
from urllib.request import urlretrieve
import tarfile
import tensorflow as tf
sys.path.append("..")
sys.path.append("xDeepFM/exdeepfm")
import config_utils
import utils.util as util
import utils.metric as metric
import train
from train import cache_data, run_eval, run_infer, create_train_model
from utils.log import Log
from src.exDeepFM import ExtremeDeepFMModel
import utilities
print("OS: ", sys.platform)
print("Python: ", sys.version)
print("Numpy: ", np.__version__)
print("Number of CPU processors: ", get_number_processors())
# breaks built on CPU/mac
# print("GPU: ", get_gpu_name())
# print("GPU memory: ", get_gpu_memory())
# print("CUDA: ", get_cuda_version())
# runtime checks
util.check_tensorflow_version()
util.check_and_mkdir()
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# +
# Parameters
# TODO
# T=4 # cut-off for minimum counts
# nrows=10000 # limit the data to reduce runtime
# supplied files come without header
fieldnames = ['Label', \
'I1', 'I2', 'I3', 'I4', 'I5', 'I6', 'I7', 'I8', 'I9', 'I10', 'I11', 'I12', 'I13', 'C1', 'C2', 'C3', 'C4', \
'C5', 'C6', 'C7', 'C8', 'C9', 'C10', 'C11', 'C12', 'C13', 'C14', 'C15', 'C16', 'C17', 'C18', \
'C19', 'C20', 'C21', 'C22', 'C23', 'C24', 'C25', 'C26']
# -
# ### Dataset retrieval
#
# See [Criteo](https://www.kaggle.com/c/criteo-display-ad-challenge/data) for licencing.
# +
# Download dataset
urlretrieve(
data_url,
"dac.tar.gz",
lambda count, blockSize, totalSize: sys.stdout.write(
"\rDownloading...%d%%" % int(count * blockSize * 100 / totalSize)
)
)
print("\n\nExtracing data")
with tarfile.open("dac.tar.gz", "r:gz") as tar:
tar.extractall("data")
train_head = pd.read_csv("data/train.txt", names=fieldnames, sep="\t", nrows=5)
train_head
# -
# ### Data preparation
# +
# create data staging directory
utilities.mkdir_safe("data_prep")
full_data = "data_prep/full.txt"
full_data_ffm = "data_prep/full.ffm"
utilities.split_files("data/train.txt", [full_data], [1], max_rows=max_rows)
feat_cnt = defaultdict(lambda: 0)
for row in csv.DictReader(open(full_data), fieldnames=fieldnames, delimiter="\t"):
for key, val in row.items():
if "C" in key:
if val == "":
feat_cnt[str(key) + "#" + "absence"] += 1
else:
feat_cnt[str(key) + "#" + str(val)] += 1
print("Found %d features" % len(feat_cnt))
# -
# ### Feature engineering
#
# * Handle missing values
# * Integers > 2: logarithmic transform (discriminate the small values [details](https://www.csie.ntu.edu.tw/~r01922136/kaggle-2014-criteo.pdf))
# * Integers $\le$ 2: convert to categorical
# * Categoricals below minimum threshold (=T): replace with categorical floor feature (**column name** # **feature count**)
#
def get_feature(key, val):
if "I" in key and key != "Id":
if val == "":
# handle missing values
return str(key) + "#" + "absence"
else:
val = int(val)
if val > 2:
# log transform + ^2 to discriminate small values
val = int(math.log(float(val)) ** 2)
else:
# convert to categorical
val = "SP" + str(val)
return str(key) + "#" + str(val)
if "C" in key:
if val == "":
# handle missing values
return str(key) + "#" + "absence"
else:
return str(key) + "#" + str(val)
if feat_cnt[feat] <= T:
# group values with small frequencies
return str(key) + "#" + str(feat_cnt[feat])
raise ValueError("Unsupported key: '%s'" % key)
# ### Discover all categorical features
# +
featSet = set()
label_cnt = defaultdict(lambda: 0)
for row in csv.DictReader(open(full_data), fieldnames=fieldnames, delimiter="\t"):
for key, val in row.items():
if key == "Label":
label_cnt[str(val)] += 1
continue
feat = get_feature(key, val)
featSet.add(feat)
rows = sum(label_cnt.values())
print(
"%25s 0: %8d (%.1f%%) 1: %8d (%.1f%%)"
% (
full_data,
label_cnt["0"],
label_cnt["0"] * 100 / rows,
label_cnt["1"],
label_cnt["1"] * 100 / rows,
)
)
# -
# ### Calculate feature and column statistics
# +
featIndex = dict()
for index, feat in enumerate(featSet, start=1):
featIndex[feat] = index
print("Categorical features count:", len(featIndex))
fieldIndex = dict()
fieldList = fieldnames[1:]
for index, field in enumerate(fieldList, start=1):
fieldIndex[field] = index
print("Field count:", len(fieldIndex))
# -
# ### Convert to [ffm format](https://github.com/guestwalk/libffm)
with open(full_data_ffm, "w") as out:
for row in csv.DictReader(open(full_data), fieldnames=fieldnames, delimiter="\t"):
feats = []
for key, val in row.items():
if key == "Label":
feats.append(val)
continue
feat = get_feature(key, val)
# lookup field index + lookup feature index
feats.append(str(fieldIndex[key]) + ":" + str(featIndex[feat]) + ":1")
out.write(" ".join(feats) + "\n")
# #### FFM Format
# One example entry "1:552:1" can be split into
#
# * field: 1
# * feature index: 552
# * feature value: 1
train_ffm_head = pd.read_csv(full_data_ffm, names=fieldnames, sep=' ', nrows=5)
train_ffm_head
# ### Split data into train, test and eval
# +
train_file_ffm = "data_prep/train.ffm"
eval_file_ffm = "data_prep/eval.ffm"
test_file_ffm = "data_prep/test.ffm"
utilities.split_files(
full_data_ffm, [train_file_ffm, test_file_ffm, eval_file_ffm], [0.8, 0.1, 0.1]
)
# -
# ### Update configuration
# +
# load network hyper-parameter
config = config_utils.load_yaml("config/exDeepFM.yaml")
# patch config to reflect the current data set
config["data"]["FEATURE_COUNT"] = len(featIndex)
config["data"]["FIELD_COUNT"] = len(fieldIndex)
config["data"]["train_file"] = train_file_ffm
config["data"]["eval_file"] = eval_file_ffm
config["data"]["test_file"] = test_file_ffm
del config["data"]["infer_file"]
# setup hparams
hparams = config_utils.create_hparams(config)
log = Log(hparams)
hparams.logger = log.logger
# -
hparams
# ### Create extreme deep FM network
# +
cache_data(hparams, hparams.train_file, flag='train')
cache_data(hparams, hparams.eval_file, flag='eval')
cache_data(hparams, hparams.test_file, flag='test')
train_model = create_train_model(ExtremeDeepFMModel, hparams)
gpuconfig = tf.ConfigProto()
gpuconfig.gpu_options.allow_growth = True
tf.set_random_seed(1234)
train_sess = tf.Session(target='', graph=train_model.graph, config=gpuconfig)
train_sess.run(train_model.model.init_op)
print('Epochs: %d' % hparams.epochs)
print('total_loss = data_loss+regularization_loss, data_loss = logloss\n')
with tf.summary.FileWriter(util.SUMMARIES_DIR, train_sess.graph) as writer:
last_eval = 0
for epoch in range(hparams.epochs):
print('Epoch %d' % epoch)
step = 0
train_sess.run(train_model.iterator.initializer, feed_dict={train_model.filenames: [hparams.train_file_cache]})
epoch_loss = 0
# TODO: collect timing infomration
while True:
try:
# TODO: collect timing information
(_, step_loss, step_data_loss, summary) = train_model.model.train(train_sess)
writer.add_summary(summary, step)
epoch_loss += step_loss
step += 1
if step % hparams.show_step == 0:
print('Step {0:d}: total_loss: {1:.4f} data_loss: {2:.4f}' \
.format(step, step_loss, step_data_loss))
except tf.errors.OutOfRangeError as e:
break
# TODO: do we need model saving in between?
if epoch % hparams.save_epoch == 0:
checkpoint_path = train_model.model.saver.save(
sess=train_sess,
save_path=util.MODEL_DIR + 'epoch_' + str(epoch))
eval_res = run_eval(train_model, train_sess, hparams.eval_file_cache, util.EVAL_NUM, hparams, flag='eval')
test_res = run_eval(train_model, train_sess, hparams.test_file_cache, util.TEST_NUM, hparams, flag='test')
print ('Train loss: %1.3f; Test loss: %1.3f; Eval loss: %1.3f auc: %0.3f\n'
% (epoch_loss / step, test_res['logloss'], eval_res['logloss'], eval_res['auc']))
# early stopping
if eval_res["auc"] - last_eval < - 0.003:
break
if eval_res["auc"] > last_eval:
last_eval = eval_res["auc"]
| examples/notebooks/xDeepFM/criteo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Awari - Data Science
#
# ## Projeto - Python com SQLite
#
# ## 1. Considerações iniciais
#
# Para que você assimile melhor os comandos do SQL, preparamos este projeto de Python com SQLite. Neste notebook Jupyter, você irá reforçando alguns conceitos da unidade atual e deverá executar algumas tarefas pelo caminho. Todo o projeto deve ser executado neste Jupyter Notebook. Por fim, se desejar, revise as respostas com o seu mentor.
#
# ### 1.1. Por que SQLite?
# O SQLite é uma pequena biblioteca em C que possui um banco de dados SQL embutido. Sem o apoio de um poderoso SGDB (Sistema de Gerenciamento de Banco de Dados) como nas soluções do mercado, o SQLite é ideal para fins de aprendizagem, pois sua instalação é fácil e não requer nenhuma configuração inicial. Deste modo, podemos focar nos comandos do SQL.
#
# ### 1.2. Prepare seu ambiente
# Baixe e instale o SQLite antes de começar. Resumidamente, você irá acessar a página de [Downloads](https://www.sqlite.org/download.html) do SQLite e baixar os executáveis para a mesma pasta deste notebook. Caso você encontre dificuldades:
#
# - Assista a este [vídeo](https://www.youtube.com/watch?v=VcKKt6OTlJc) para "instalar" o SQLite no Windows ou;
# - Ou siga este [manual](http://www.devfuria.com.br/linux/instalando-sqlite/) para as plataformas Linux.
#
# ### 1.3. Conjunto de dados
#
# O conjunto de dados que iremos utilizar é derivado do [Titanic](https://www.kaggle.com/c/titanic/data). Na verdade, dividimos o conjunto em dois arquivos CSV - [passageiros](passageiros.csv) e [passagens](passagens.csv). Cada um destes arquivo serão inseridos dentro do SQLite como tabelas, onde você executará suas consultas SQL.
#
# #### 1.3.1. Descrição do dados
#
# No arquivo [passageiros](passageiros.csv), você encontrará:
# - PassengerId: Id do passageiro
# - Survived: Se o passageiro sobreviveu ou não. Se 0, então não sobreviveu. Se 1, sobreviveu.
# - Pclass: Classe em que o passageiro estava viajando. Se 1, então estava na primeira classe e assim por diante.
# - Name: Nome do passageiro.
# - Sex: Gênero do passageiro. Masculino ou feminino.
# - Age: Idade.
# - Sibsp: número de irmãos / esposas a bordo do navio.
# - Parch: número de pais / filhos a bordo do navio.
#
# E no arquivo [passagens](passagens.csv):
# - PassengerId: Id do passageiro
# - Ticket: número do ticket de embarque
# - Fare: valor pago pelo ticket.
# - Cabin: cabine do passageiro.
# - Embarked: porto de embarque do passageiro. C = Cherbourg, Q = Queenstown e S = Southampton
#
# ## 2. Procedimentos
#
# ### 2.1. Leitura dos arquivos CSV
# Antes de importarmos os dados dentro das tabelas, precisamos começar lendo os arquivos CSV. O pequeno código abaixo importa os dados do CSV em uma lista de dicionários em Python.
# +
import csv
def importar_csv(arquivo):
dados = []
with open(arquivo) as arq:
leitor = csv.DictReader(arq)
for l in leitor:
dados.append(dict(l))
print('O conjunto de dados tem %s registros.' % len(dados))
return dados
# -
# ### TAREFA 01
#
# 1. Importe o arquivo passagens.csv usando o mesmo código anterior.
#
# **DICA**: Não reinvente a roda, reutilize a função anterior.
# Insira seu código aqui
passagens = importar_csv('passagens.csv')
passageiros = importar_csv('passageiros.csv')
# ### TAREFA 02
# 1. Cria a tabela passageiros.
# 2. PassenderId deve ser a chave primária.
# Insira seu código aqui
passageiros_sql = 'CREATE TABLE passageiros(\
PassengerId INTEGER PRIMARY KEY,\
Survived INTEGER,\
Pclass INTEGER,\
Name TEXT,\
Sex TEXT,\
Age INTEGER,\
Sibsp INTEGER,\
Parch INTEGER\
)'
executa(passageiros_sql)
# ### 2.2. Criando o banco de dados
# Para manipular um banco SQLite com Python, temos que utilizar uma biblioteca específica.
import sqlite3
conn = sqlite3.connect('titanic.db')
# O comando acima cria o arquivo [titanic.db](titanic.db), que armazará nossos dados.
# ### 2.3. Criando as tabelas
# Neste momento, temos do dados e o banco SQLlite. Então é hora de começar à usar comandos SQL para criar as tabelas que armazenarão nossos dados dentro do banco.
#
# No SQLlite, precisamos primeiro criar uma conexão e um cursor para executar nossos comandos no banco. Para facilicar, o código abaixo facilita este processo. Você só precisa passar seu comando SQL para a função.
def executa(comando):
cursor = conn.cursor()
cursor.execute(comando)
cursor.close()
# Não execute o comando abaixo, pois ele dá um erro. É só um exemplo
# +
# Exemplo
#comando = '<SEU_COMANDO_AQUI>'
#executa(comando)
# -
# ### TAREFA 03
# 1. Cria a tabela passagens.
# 2. PassenderId deve ser a chave primária.
# Insira seu código aqui
passagens_sql = 'CREATE TABLE passagens(\
PassengerId INTEGER PRIMARY KEY,\
Ticket TEXT,\
Fare REAL,\
Cabin TEXT,\
Embarked TEXT\
)'
executa(passagens_sql)
# ### TAREFA 04
# 1. Cria a tabela passagens.
# 2. PassenderId deve ser a chave primária.
# Insira seu código aqui
passagens_sql_2 = 'CREATE TABLE passagens(\
PassengerId INTEGER PRIMARY KEY,\
Ticket TEXT,\
Fare REAL,\
Cabin TEXT,\
Embarked TEXT\
)'
executa(passagens_sql_2)
# **Atenção:** Caso você tente executar os comandos de criação de tabelas mais de uma vez, perceberá um erro de operação informando que a tabela já existe.
#
# Vamos verificar se as tabelas foram realmente criadas? Abra o terminal e na pasta atual digite:
# ```
# $ sqlite3.exe titanic.db ".tables"
# ```
# O terminal deve retornar com as tabelas existentes em seu banco [titanic.db](titanic.db). Neste casos, *passageiros* e *passagens*. Lembre-se que neste momento, não temos nenhum dado inseridos nas tabelas.
#
# Caso queira verifcar mais algumas informações das tabelas criadas, pode executar esta outra instrução:
# ```
# $ sqlite3.exe titanic.db "PRAGMA table_info(passageiros)"
# ```
# ### TAREFA 05
# 1. Pelo terminal, verifique mais informações da tabela passagens.
# ### 2.4. Inserindo dados nas tabelas
# Usando o comando INSERT INTO, vamos popular as nossas tabelas. O processo é simples, percorremos as listas de dicionários, montamos nosso comando INSERT INT e executamos - uma a uma. Para a tabela passageiros:
for p in passageiros:
p['Name'] = p['Name'].replace('"', "'")
inserir_passageiro = ("""
INSERT INTO passageiros(PassengerId, Survived, Pclass, Name, Sex, Age, Sibsp, Parch)
VALUES (%s, "%s", "%s", "%s", "%s", "%s", "%s", "%s")""") % (p['PassengerId'], p['Survived'], p['Pclass'],
p['Name'], p['Sex'], p['Age'], p['SibSp'],
p['Parch'])
conn.commit()
executa(inserir_passageiro)
# ### TAREFA 06
# 1. Insira os dados de passagens dentro da tabela passagens
# Insira seu código aqui
for psg in passagens:
psg['Ticket'] = psg['Ticket'].replace('"', "'")
psg['Cabin'] = psg['Cabin'].replace('"', "'")
psg['Embarked'] = psg['Embarked'].replace('"', "'")
inserir_passagens = ("""
INSERT INTO passagens(PassengerId, Ticket, Fare, Cabin, Embarked)
VALUES (%s, "%s", %s, "%s", "%s")""") % (psg['PassengerId'], psg['Ticket'], psg['Fare'],
psg['Cabin'], psg['Embarked'])
conn.commit()
executa(inserir_passagens)
# ### 2.5. Executando consultas
# Agora que os dados estão inseridos no arquivo banco SQLite, vamos executar alguns consultas. Vamos começar fazendo uma seleção de todas as colunas das tabelas.
#
# O comando SELECT retorna dados, deste modo, precisamos fazer alguma alteração na função executa. utilize o *executa_consulta()*.
def executa_consulta(consulta):
cursor = conn.cursor()
cursor.execute(consulta)
for linha in cursor.fetchall():
print(linha)
cursor.close()
# Testando nosso código. Repare que o comando LIMIT foi utilizado para limitar a quantidade de linhas.
# Insira seu código aqui
# ### TAREFA 07
# 1. Verifique quantos tipos de classe (PClass) existem na tabela passageiros.
# Insira seu código aqui
consulta_1 = 'SELECT p.Pclass FROM passageiros AS p GROUP BY p.Pclass'
executa_consulta(consulta_1)
# +
# Resposta: Há 3 tipos de classes.
# -
# ### TAREFA 08
# 1. Selenione os passageiros do sexo masculino (Sex) e que estejam na segunda classe (PClass).
#
# **DICA**: Use a cláusula WHERE e uma condicional (AND ou OR).
# Insira seu código aqui
consulta_2 = 'SELECT * FROM passageiros as p \
WHERE p.Sex = "male" and p.Pclass = 2 '
executa_consulta(consulta_2)
# ### TAREFA 09
# 1. Conte quantos passageiros existem por sexo.
#
# **DICA**: Use GROUP BY e COUNT().
# Insira seu código aqui
consulta_3 = 'SELECT p.Sex, count()\
FROM passageiros as p\
GROUP BY p.Sex'
executa_consulta(consulta_3)
# ### TAREFA 10
# 1. Verifique o valor médio das passagens em cada porto de embarque
#
# **DICA**: Use GROUP BY e AVG(). A presença de valores missing pode apresentar alguns resultados estranhos.
# Insira seu código aqui
consulta_4 = "SELECT psg.Embarked, avg(psg.Fare)\
FROM passagens as psg \
GROUP BY psg.Embarked"
executa_consulta(consulta_4)
# ### TAREFA 11
# 1. Selecione o nome (Name), sexo(Sex) e tarifa(Fare) paga de 5 passageiros
# 2. Utilize as tabelas passageiros e passagens.
#
# **DICA**: Use JOIN.
consulta_5 = 'SELECT p.Name, p.Sex, psg.Fare \
FROM passageiros AS p\
JOIN passagens AS psg ON p.PassengerID = psg.PassengerId\
LIMIT 5'
# Insira seu código aqui
executa_consulta(consulta_5)
# ### TAREFA 12
# 1. Selecione o valor máximo (Fare) das passagens por classe.
# Insira seu código aqui
consulta_6 = 'SELECT p.Pclass, max(psg.Fare) \
FROM passageiros as p \
JOIN passagens as psg ON p.PassengerID = psg.PassengerId \
GROUP BY p.Pclass'
executa_consulta(consulta_6)
# ### TAREFA 13
# 1. Descubra quantas pessoas embarcaram por porto
# 2. Agrupando por sexo.
# Insira seu código aqui
consulta_7 = 'SELECT psg.Embarked, p.Sex, count()\
FROM passageiros AS p, passagens as psg\
WHERE p.PassengerId = psg.PassengerId \
GROUP BY psg.Embarked, p.Sex'
executa_consulta(consulta_7)
# ## 3. Conclusão
#
# A quantidade de consultas e a lista de comandos SQL é enorme. A interação foi somente relembrar alguns comandos básicos para que você se sinta confortável para explorar qualquer banco de dados SQL que você encontrar.
#
# ### Awari - <a href="https://awari.com.br/"> awari.com.br</a>
| Unidade7-Banco-de-Dados-SQL/Projeto - Python com SQLite.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TEXT DOCUMENT CLUSTERING WITH TF-IDF, KMEANS, AND FP-GROWTH
# +
import re
import nltk
import json
import numpy as np
import pandas as pd
from html import unescape
from sklearn.cluster import KMeans
from sklearn.feature_extraction.text import TfidfVectorizer
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer
import pandas as pd
from mlxtend.preprocessing import TransactionEncoder
from mlxtend.frequent_patterns import apriori
from kneed import KneeLocator
# -
# ## PREPROCESSING FUNCTIONS
# +
def tokenize_and_stem(text):
lemmatizer = WordNetLemmatizer()
tokens = [word for sent in nltk.sent_tokenize(text) for word in nltk.word_tokenize(sent)]
filtered_tokens = []
for token in tokens:
if re.search('[a-zA-Z]', token):
filtered_tokens.append(token.lower())
stems = []
for t in filtered_tokens:
stems.append(lemmatizer.lemmatize(t))
return ' '.join(stems)
def punc(docs):
toreturn = []
for word in re.findall(r"[\w\-_#]+", docs):
if word not in stopwords.words('english') and not word.isnumeric():
toreturn.append(word)
return ' '.join(toreturn)
# -
# ## LOADING DATASET
with open('mixed.json') as f:
linklist = json.load(f)
df = pd.DataFrame(linklist['results'])
df
df.drop(['a', 'c'], axis=1, inplace=True)
df['d'] = df['t'] + ' ' + df['d']
df
# ## PREPROCESSING - TOKENIZING, LEMMATIZING TOKENS, AND REMOVING PUNCTUTATION
df['d'] = df['d'].apply(tokenize_and_stem)
df['d'] = df['d'].apply(punc)
df
# ## CREATING A TFIDF VECTORIZER
vectorizer = TfidfVectorizer(max_features=20, min_df=0.2)
X = vectorizer.fit_transform(df['d'])
print(vectorizer)
# ## USING ELBOW METHOD TO FIND OPTIMAL K FOR K-MEANS
# +
# FIND OPTIMAL K
inertias = []
fitted = []
for cluster_count in range(1,20):
km = KMeans(n_clusters=cluster_count)
km.fit(X)
inertias.append(km.inertia_)
fitted.append(km)
kn = KneeLocator(range(1, len(inertias)+1), inertias, curve='convex', direction='decreasing')
optimal_k = kn.knee
print(f'OPTIMAL K = {optimal_k}')
# -
from matplotlib import pyplot as plt
plt.scatter(range(1, len(inertias)+1), inertias)
plt.show()
# # CHOOSING MODEL FITTED WITH OPTIMAL K FOR ASSIGNING CLUSTERS
# +
best_fit_index = optimal_k-1
km = fitted[best_fit_index]
terms = vectorizer.get_feature_names()
order_centroids = km.cluster_centers_.argsort()[:, ::-1]
summaries = [[] for _ in range(optimal_k)]
titles = [[] for _ in range(optimal_k)]
for i in range(optimal_k):
for index, cluster in enumerate(km.labels_.tolist()):
if cluster == i:
summaries[i].append(df['d'][index])
titles[i].append(df['t'][index])
# -
# ## USING FP-GROWTH TO GET FREQUENTLY APPEARING SETS OF WORDS FROM DATA FOR TITLES
# +
# APRIORI AS ALTERNATIVE TO FPGROWTH (SLOWER)
# te = TransactionEncoder()
# from time import time
# def get_frequent_itemsets(dataset):
# # PASSED DATASET MUST CONTAIN SPLIT WORDS APPEARING IN THE SAME ORDER AS THEIR SENTENCES
# te_ary = te.fit_transform(dataset)
# dataset = pd.DataFrame(te_ary, columns=te.columns_)
# freq_itemsets = apriori(dataset, min_support=0.2, use_colnames=True)
# freq_itemsets = freq_itemsets.sort_values('support', ascending=False)['itemsets']
# freq_itemsets = list(filter(lambda x:len(x)>1, freq_itemsets.values.tolist()))
# return freq_itemsets
# for index, cluster in enumerate(summaries):
# d = list(map(lambda x: x.split(), cluster))
# probable_titles = get_frequent_itemsets(d)
# cluster_title = ' | '.join(list(map(lambda x: ' '.join(list(x)[::-1]), probable_titles))[:5])
# print(f"CLUSTER {index+1}")
# print(f"TITLE: {cluster_title}")
# print(' '+'\n '.join(titles[index]))
# print('\n\n')
# +
from mlxtend.frequent_patterns import fpgrowth
te = TransactionEncoder()
from time import time
def get_frequent_itemsets(dataset):
# PASSED DATASET MUST CONTAIN SPLIT WORDS APPEARING IN THE SAME ORDER AS THEIR SENTENCES
te_ary = te.fit_transform(dataset)
dataset = pd.DataFrame(te_ary, columns=te.columns_)
freq_itemsets = fpgrowth(dataset, min_support=0.2, use_colnames=True)
freq_itemsets = freq_itemsets.sort_values('support', ascending=False)['itemsets']
freq_itemsets = list(filter(lambda x:len(x)>1, freq_itemsets.values.tolist()))
return freq_itemsets
for index, cluster in enumerate(summaries):
d = list(map(lambda x: x.split(), cluster))
probable_titles = get_frequent_itemsets(d)
cluster_title = ' | '.join(list(map(lambda x: ' '.join(list(x)[::-1]), probable_titles))[:5])
print(f"CLUSTER {index+1}")
print(f"TITLE: {cluster_title}")
print(' '+'\n '.join(titles[index]))
print('\n\n')
| clustering_new.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1 概述
#
# 再来回忆一下我们的终极目标,我们的终极目标是搭建一个人脸识别的模型,经过之前的努力,我们已经训练出来了一个人脸检测模型,然后我们就可以利用检测到的人脸,将其输入人脸识别模型中,来达到人脸识别的目的。
#
# 现在有一个问题,之前的人脸检测模型是通过darknet平台实现的,但是接下来的人脸识别模型我们不会使用darknet网络了,而会使用keras+tensorflow,为了整个模型的灵活性,也为了更好的管理人脸识别模型,我们需要将darknet移植出来,用Python来实现。当然并不会抑制整个darknet平台,仅仅移植和YOLO相关的部分,当然,我也不会自己动手去移植,因为已经有人已经帮助我们做好了这些事情!
#
# 所以接下来,我们将使用[YAD2K](https://github.com/allanzelener/YAD2K)这个开源工程,其实就是darknet的Python版本,但是支持的模型仅仅是类YOLO的模型(tiny-yolo等)。
#
# 所以darknet在本项目中就是为了训练YOLO人脸检测模型的权重,然后我们利用YAD2K来搭建YOLO人脸检测模型,加载权重。最后再搭建人脸识别模型。
# # 2 YAD2K
#
# 本项目就是基于[YAD2K](https://github.com/allanzelener/YAD2K)上进行开发的,所以你没必要再去克隆。再声明一点,本notebook就位于YAD2K内。
#
# 接下来我们就来走一遍YAD2K的流程,看看他到底做了哪些事情。然后我们再提取我们需要的部分来搭建YOLO模型!
# ## 2.1 YAD2K运行环境搭建
#
# 首先根据YAD2K的使用说明来搭建环境,输入如下命令
#
# git clone https://github.com/allanzelener/yad2k.git
# cd yad2k
#
# ## 为了方便,作者把yad2k的环境信息都放在了environment.yml文件里面,运行下面的命令创建环境
# conda env create -f environment.yml
# source activate yad2k
#
# # 安装一些常用的库
# pip install numpy h5py pillow
# # 安装GPU版本的tensorflow,如果没有GPU的话,安装CPU版本的也可以,就是CPU版本的预测的速度要慢很多
# pip install tensorflow-gpu # CPU-only: conda install -c conda-forge tensorflow
# pip install keras # Possibly older release: conda install keras
# ## 2.2 YOLO模型配置文件以及权重
#
# 就像之前[第二个notebook](https://github.com/rikichou/yolo/blob/master/YOLO_face_detection_and_recognition_2--train_yolo_on_CelebA_datasheet.ipynb)说的那样,darknet需要一个模型的配置文件和权重才能把模型搭建起来,同理,对于YAD2K也需要这两样东西。
#
# 1,YOLO模型配置文件,本git已经自带,位于cfg/yolov2-celeba.cfg
# 2,权重文件是我们在CelebA数据集上训练的权重,有大概200M,我放在了另一个仓库里面,记得克隆前要安装git LFS
# git lfs clone https://github.com/rikizhou/yolo_face_weights.git
# 下载完了把它拷贝到weights目录下就可以了
# ## 2.3 转换darknet模型
#
# 在上一步中,我们获得darknet下载YOLO模型的配置文件和权重,这两样东西包含了搭建模型需要的全部东西。现在我们就根据这两样东西,将darknet模型转换为keras模型。
#
# 注意注意!各位同学!!!这里需要更改源码的一个BUG,在yad2k.py,在第 83 行,将 buffer=weights_file.read(16) 改为 buffer=weights_file.read(20),否则会出现权重错位,导致模型完全错误!当然,在本项目中,已经更改好了!
# ./yad2k.py cfg/yolov2-celeba.cfg weights/yolov2-celeba_5000.weights model_data/yolo_face_detectoin.h5
#
# 运行了之后,会在model_data目录下生成yolo_face_detectoin_anchors.txt文件
# ## 2.4 预测测试图片
#
# YAD2K是利用test_yolo.py这个文件来实现预测的,所以我们应该查看其源代码,分析该脚本需要提供什么参数?
#
# model_path:keras模型的路径,model_data/yolo_face_detectoin.h5
# --anchors_path:我们需要指定anchors文件的路径,model_data/yolo_face_detectoin_anchors.txt
# --classes_path:类别文件的路径,里面指定了每个类别的名字,我们就只有一个face,model_data/celeba_classes.txt
# --test_path:输入的测试图片的路径,默认是在images目录下,我们就用默认的就好了
# --output_path:输出的测试图片的路径,默认在images/out,我们就用默认
# --score_threshold:将每个anchors的pc乘以该anchors的类别概率就等于scores,对于YOLO模型的预测输出,如果某个anchors的score小于score_threshold,那么该anchors的预测将直接会被删除,不会参与IOU的流程。默认为0.3,我们暂时使用0.3,然后根据效果来调节该阈值。
# --iou_threshold:IOU的阈值,在执行非最大值抑制(NMS)的时候,如果两个同类预测的bbox的IOU>iou_threshold,那么就认为该对象被重复检测了,需要删除几率小的那个Bbox。默认为0.5,我们暂时使用0.5,然后根据效果来调节该阈值。
#
# 如果我上面说的这些大家不懂的话,那么可能需要看关于YOLO的视频了,这里推荐吴恩达教授Coursera上的DeepLearning.ai系列课程,在CNN部分讲解了YOLO的详细流程。
# 接下来我们就要预测一下images里面的图片,输出的结果在images/out文件夹里面(照片我可能没有上传到github,所以你可以预测你自己的图片)
#
# ./test_yolo.py model_data/yolo_face_detectoin.h5 -a model_data/yolo_face_detectoin_anchors.txt -c model_data/celeba_classes.txt
# +
import matplotlib.pyplot as plt
import cv2
import numpy as np
# %matplotlib inline
img_path = "images/out/me.png"
plt.figure(figsize=(6,6))
img = cv2.imread(img_path)
img = cv2.cvtColor(img,cv2.COLOR_BGR2RGB)
plt.imshow(img)
plt.axis("off")
plt.show()
# -
# # 3 我们需要怎样的接口
#
# 经过上面的对图片的预测,我们可以基本判定我们需要什么样的接口了,一个接口用于根据传入的配置创建模型,另一个接口用于预测输入的图片,输出图片的所有的bounding box以及所有的bounding box属于脸的概率。
#
# 所以,我们可以创建一个类,如下代码:
# +
import configparser
import io
import os
from collections import defaultdict
import colorsys
import imghdr
import random
import numpy as np
from keras import backend as K
K.set_image_data_format('channels_last')
from PIL import Image, ImageDraw, ImageFont
from keras.layers import (Conv2D, GlobalAveragePooling2D, Input, Lambda,
MaxPooling2D)
from keras.layers.advanced_activations import LeakyReLU
from keras.layers.merge import concatenate
from keras.layers.normalization import BatchNormalization
from keras.models import Model
from keras.regularizers import l2
from keras.utils.vis_utils import plot_model as plot
from yad2k.models.keras_yolo import yolo_eval, yolo_head
from yad2k.models.keras_yolo import (space_to_depth_x2,
space_to_depth_x2_output_shape)
default_config_path = "cfg/yolov2-celeba.cfg"
default_weights_path = "weights/yolov2-celeba_5000.weights"
default_anchors_path = "model_data/yolo_face_detectoin_anchors.txt"
default_classes_path = "model_data/celeba_classes.txt"
default_score_threshold = 0.3
default_iou_threshold = 0.5
class Detector(object):
def __init__(self, config_path=default_config_path, weights_path=default_weights_path, anchors_path=default_anchors_path, \
classes_path = default_classes_path, score_threshold=default_score_threshold, iou_threshold=default_iou_threshold):
self.config_path = config_path
self.weights_path = weights_path
self.anchors_path = anchors_path
self.classes_path = classes_path
self.sess = K.get_session()
self.iou_threshold = iou_threshold
self.score_threshold = score_threshold
output_root = "model_data"
print('Loading weights.')
# Load weights and config.
weights_file = open(weights_path, 'rb')
weights_header = np.ndarray(
shape=(4, ), dtype='int32', buffer=weights_file.read(20))
unique_config_file = self.unique_config_sections(self.config_path)
cfg_parser = configparser.ConfigParser()
cfg_parser.read_file(unique_config_file)
image_height = int(cfg_parser['net_0']['height'])
image_width = int(cfg_parser['net_0']['width'])
self.image_height = image_height
self.image_width = image_width
prev_layer = Input(shape=(image_height, image_width, 3))
all_layers = [prev_layer]
weight_decay = float(cfg_parser['net_0']['decay']
) if 'net_0' in cfg_parser.sections() else 5e-4
count = 0
for section in cfg_parser.sections():
print('Parsing section {}'.format(section))
if section.startswith('convolutional'):
filters = int(cfg_parser[section]['filters'])
size = int(cfg_parser[section]['size'])
stride = int(cfg_parser[section]['stride'])
pad = int(cfg_parser[section]['pad'])
activation = cfg_parser[section]['activation']
batch_normalize = 'batch_normalize' in cfg_parser[section]
# padding='same' is equivalent to Darknet pad=1
padding = 'same' if pad == 1 else 'valid'
# Setting weights.
# Darknet serializes convolutional weights as:
# [bias/beta, [gamma, mean, variance], conv_weights]
prev_layer_shape = K.int_shape(prev_layer)
# TODO: This assumes channel last dim_ordering.
weights_shape = (size, size, prev_layer_shape[-1], filters)
darknet_w_shape = (filters, weights_shape[2], size, size)
weights_size = np.product(weights_shape)
print('conv2d', 'bn'
if batch_normalize else ' ', activation, weights_shape)
conv_bias = np.ndarray(
shape=(filters, ),
dtype='float32',
buffer=weights_file.read(filters * 4))
count += filters
if batch_normalize:
bn_weights = np.ndarray(
shape=(3, filters),
dtype='float32',
buffer=weights_file.read(filters * 12))
count += 3 * filters
# TODO: Keras BatchNormalization mistakenly refers to var
# as std.
bn_weight_list = [
bn_weights[0], # scale gamma
conv_bias, # shift beta
bn_weights[1], # running mean
bn_weights[2] # running var
]
conv_weights = np.ndarray(
shape=darknet_w_shape,
dtype='float32',
buffer=weights_file.read(weights_size * 4))
count += weights_size
# DarkNet conv_weights are serialized Caffe-style:
# (out_dim, in_dim, height, width)
# We would like to set these to Tensorflow order:
# (height, width, in_dim, out_dim)
# TODO: Add check for Theano dim ordering.
conv_weights = np.transpose(conv_weights, [2, 3, 1, 0])
conv_weights = [conv_weights] if batch_normalize else [
conv_weights, conv_bias
]
# Handle activation.
act_fn = None
if activation == 'leaky':
pass # Add advanced activation later.
elif activation != 'linear':
raise ValueError(
'Unknown activation function `{}` in section {}'.format(
activation, section))
# Create Conv2D layer
conv_layer = (Conv2D(
filters, (size, size),
strides=(stride, stride),
kernel_regularizer=l2(weight_decay),
use_bias=not batch_normalize,
weights=conv_weights,
activation=act_fn,
padding=padding))(prev_layer)
if batch_normalize:
conv_layer = (BatchNormalization(
weights=bn_weight_list))(conv_layer)
prev_layer = conv_layer
if activation == 'linear':
all_layers.append(prev_layer)
elif activation == 'leaky':
act_layer = LeakyReLU(alpha=0.1)(prev_layer)
prev_layer = act_layer
all_layers.append(act_layer)
elif section.startswith('maxpool'):
size = int(cfg_parser[section]['size'])
stride = int(cfg_parser[section]['stride'])
all_layers.append(
MaxPooling2D(
padding='same',
pool_size=(size, size),
strides=(stride, stride))(prev_layer))
prev_layer = all_layers[-1]
elif section.startswith('avgpool'):
if cfg_parser.items(section) != []:
raise ValueError('{} with params unsupported.'.format(section))
all_layers.append(GlobalAveragePooling2D()(prev_layer))
prev_layer = all_layers[-1]
elif section.startswith('route'):
ids = [int(i) for i in cfg_parser[section]['layers'].split(',')]
layers = [all_layers[i] for i in ids]
if len(layers) > 1:
print('Concatenating route layers:', layers)
concatenate_layer = concatenate(layers)
all_layers.append(concatenate_layer)
prev_layer = concatenate_layer
else:
skip_layer = layers[0] # only one layer to route
all_layers.append(skip_layer)
prev_layer = skip_layer
elif section.startswith('reorg'):
block_size = int(cfg_parser[section]['stride'])
assert block_size == 2, 'Only reorg with stride 2 supported.'
all_layers.append(
Lambda(
space_to_depth_x2,
output_shape=space_to_depth_x2_output_shape,
name='space_to_depth_x2')(prev_layer))
prev_layer = all_layers[-1]
elif section.startswith('region'):
with open('{}_anchors.txt'.format(output_root), 'w') as f:
print(cfg_parser[section]['anchors'], file=f)
elif (section.startswith('net') or section.startswith('cost') or
section.startswith('softmax')):
pass # Configs not currently handled during model definition.
else:
raise ValueError(
'Unsupported section header type: {}'.format(section))
self.model = Model(inputs=all_layers[0], outputs=all_layers[-1])
print(self.model.summary())
remaining_weights = len(weights_file.read()) / 4
weights_file.close()
print('Read {} of {} from Darknet weights.'.format(count, count +
remaining_weights))
if remaining_weights > 0:
print('Warning: {} unused weights'.format(remaining_weights))
"""
=========================================================================
"""
with open(self.classes_path) as f:
class_names = f.readlines()
self.class_names = [c.strip() for c in class_names]
with open(self.anchors_path) as f:
anchors = f.readline()
anchors = [float(x) for x in anchors.split(',')]
self.anchors = np.array(anchors).reshape(-1, 2)
# Verify model, anchors, and classes are compatible
num_classes = len(self.class_names)
num_anchors = len(self.anchors)
# TODO: Assumes dim ordering is channel last
model_output_channels = self.model.layers[-1].output_shape[-1]
assert model_output_channels == num_anchors * (num_classes + 5), \
'Mismatch between model and given anchor and class sizes. ' \
'Specify matching anchors and classes with --anchors_path and ' \
'--classes_path flags.'
##### check if need
# Check if model is fully convolutional, assuming channel last order.
self.model_image_size = self.model.layers[0].input_shape[1:3]
is_fixed_size = self.model_image_size != (None, None)
# Generate colors for drawing bounding boxes.
hsv_tuples = [(x / len(self.class_names), 1., 1.)
for x in range(len(self.class_names))]
colors = list(map(lambda x: colorsys.hsv_to_rgb(*x), hsv_tuples))
colors = list(
map(lambda x: (int(x[0] * 255), int(x[1] * 255), int(x[2] * 255)),
colors))
random.seed(10101) # Fixed seed for consistent colors across runs.
random.shuffle(colors) # Shuffle colors to decorrelate adjacent classes.
random.seed(None) # Reset seed to default.
##### check if need
# Generate output tensor targets for filtered bounding boxes.
# TODO: Wrap these backend operations with Keras layers.
yolo_outputs = yolo_head(self.model.output, self.anchors, len(self.class_names))
self.input_image_shape = K.placeholder(shape=(2, ))
self.boxes, self.scores, self.classes = yolo_eval(
yolo_outputs,
self.input_image_shape,
score_threshold=self.score_threshold,
iou_threshold=self.iou_threshold)
def predict(self, image):
"""
input : np array, h,w,c
return :
out_boxes:all bounding boxes
out_scores:all bounding box's score
out_classes:all bounding box's classes
"""
h, w, c = image.shape
image_data = np.array(image, np.float32)
image_data = cv2.resize(image_data, (self.image_height, self.image_width))
image_data /= 255.
image_data = np.expand_dims(image_data, 0) # Add batch dimension.
out_boxes, out_scores, out_classes = self.sess.run(
[self.boxes, self.scores, self.classes],
feed_dict={
self.model.input: image_data,
self.input_image_shape: [h, w],
K.learning_phase(): 0
})
#print('Found {} boxes for image'.format(len(out_boxes)))
return out_boxes, out_scores, out_classes
"""
crop_imgs = []
for i, c in reversed(list(enumerate(out_classes))):
predicted_class = self.class_names[c]
box = out_boxes[i]
score = out_scores[i]
label = '{} {:.2f}'.format(predicted_class, score)
top, left, bottom, right = box
print (label + "top:{} left:{} bottom:{} right:{}".format(top, left, bottom, right))
item = image[int(top):int(bottom), int(left):int(right), :]
crop_imgs.append(item)
return crop_imgs
"""
def unique_config_sections(self, config_file):
"""Convert all config sections to have unique names.
Adds unique suffixes to config sections for compability with configparser.
"""
section_counters = defaultdict(int)
output_stream = io.StringIO()
with open(config_file) as fin:
for line in fin:
if line.startswith('['):
section = line.strip().strip('[]')
_section = section + '_' + str(section_counters[section])
section_counters[section] += 1
line = line.replace(section, _section)
output_stream.write(line)
output_stream.seek(0)
return output_stream
# -
a = Detector()
# +
import cv2
import matplotlib.pyplot as plt
img_path = "images/party.png"
img = cv2.imread(img_path)
img = cv2.cvtColor(img,cv2.COLOR_BGR2RGB)
out_boxes, out_scores, out_classes = a.predict(img)
print (out_boxes)
#plt.imshow(crop_imgs[0])
#plt.axis("off")
#plt.show()
# -
# # 4 人脸识别
#
# 可以看到,上面的代码可以将图片中所有的人脸检测出来,现在我们就可以根据这些人脸来构建人脸的识别部分!
# ## 4.1 一个简短的教程
#
# 在人脸识别领域,通常有两个概念,一个是人脸验证,一个是人脸识别。
# 人脸验证:输入一张图片和人名/ID,输出的结果是该图片是否和人名/ID匹配。
#
# 人脸识别:输入一个图片,和有K个人的数据库进行匹配,输出和该输入匹配的ID或者无法识别。该问题要难于人脸验证,对于一个人脸验证的应用,有99%的正确率是可以接受的。但是如果要把该人脸验证用于人脸识别(100张人脸的数据库)的话,99%的验证正确率只能有37%左右的识别正确率,所以对于人脸识别来说,需要99.99%以及更高的验证成功率
# ### 4.1.1 one-shot 学习问题
#
# 也就是说在大多数人脸识别应用中,你需要仅仅通过一张图片,就能识别这个人,但是对于监督学习来说,只有一个训练样本的话,训练效果并不好。
#
# 例如:
# 比如我们训练了一个模型,输入一张图片,输出101个softmax的结果,看看输入的这个人属于/不属于数据库中某个人的概率,但是实际上这样效果并不好。因为训练样本太少了。而且当有新人加入数据库的时候,此时你的模型的softmax的输出单元就变成了102,难道要重新训练整个卷积网络?
#
# 所以我们需要的是学习一个差异性的函数,通过输入两张图片来输出一个差异值,如果差异值小于某个阈值τ,那么就是同一个人。
#
# distance = d(img1, img2)
#
# if distance < threshold:
# same person
# else:
# different person
# 这样的话,当要识别一个新人,只需要把他的图片加入到数据库中就行了。
# ### 4.1.2 Siamese网络
#
# 上一节了解到我们的d(img1, img2)函数的作用输入两张图片,输出这两张图片的差异值。实现这个功能的一个好的方式就是采用[Siamese](https://www.cs.toronto.edu/~ranzato/publications/taigman_cvpr14.pdf)网络了.
#
# 在我们平常看到的CNN中,一般是由若干卷积层加全连接层加softmax输出来构成的,现在我们不关注softmax层,将其去掉,那么就剩下了一个输出维度为比如128的全连接层,那么此时就相当于输入了一张图片,我们的CNN网络将其转换为了一个128维的向量,标记为f(x1)。这相当于图片x1的编码,将一张输入片表示成了一个128维的向量。如果此时有里那个一张输入的图片x2,那么这张图片对应的输出向量就是f(x1)。所以现在我们定义d(x1, x2)就相当于定义两个向量之间的差异!自然,在数学上有很多方式可以来表示两个向量的差异,我们这里使用两个向量之差的二阶范数的平方来表示。
#
# d(x1, x2) = L2 norm(f(x1) - f(x2))
#
# 这种通过运行同一个神经网络,来得到两个输入的两个输出向量,然后在比较这个输出向量的网络叫做Siamese网络。这里提到的思想都可以在[DeepFace: Closing the Gap to Human-Level Performance in Face Verification](https://www.cs.toronto.edu/~ranzato/publications/taigman_cvpr14.pdf)中找到。
#
# 所以我们现在学习的目的就是:
#
# 如果x1,x2是同一个人的话,我们就要使得d(x1, x2)尽量小。反之,d(x1, x2)尽量大
# ### 4.1.3 Triplet loss
#
# 上面了解了学习目的,现在来介绍一下人脸识别的学习策略:我们会同时比较三张图片,这三张图片分别为anchor,positive,negative,anchor为基准图片,positive是和anchor同一个人的不同图片,negative是和anchor不同人的不同图片,所以现在的学习目标就是distance(anchor, positive) 远远小于 distance(anchor, negative):
#
# d(A, P) - d(A, N) + margin <= 0
#
# 所以我们定义loss函数为:
#
# L(A,N,P) = max(d(A, P) - d(A, N) + margin, 0)
#
# 所以我们的训练数据集就是A,P,N这样的三元组,所以在训练集中,我们需要同一个人的多张照片组成A,P,然后再选择另一个人的照片N,构成一个三元组。而且对于三元组的选择也是有要求的,如果说随机地选择AP和N,那么这样会导致loss的条件很容易达成,模型将学不会足够有用的信息,我们应该选择P和AN很像,但是又不是同一个人的图片来构成三元组。也就是说d(A, N)很接近d(A, P),这样的话算法会竭尽全力使得d(A,P)小于d(A,N)。上述的观点很多都来自于[FaceNet: A Unified Embedding for Face Recognition and Clustering](https://arxiv.org/abs/1503.03832)。
# ### 4.1.4 另一种人脸验证的模型--二分类模型
#
# 上面说了Siamese网络+triplet loss实现人脸验证功能。现在来介绍另一个模型,该模型同样采用Siamese网络,只是loss function采用的sigmoid。
#
# 同样是网络输入两张图片,分别得到两个128维的向量,然后将这两个向量相减,向量的结果的绝对值作为sigmoid函数的输入,然后输出为0--1之间,表示这两张图片是同一个人的概率。这里提到的思想都可以在[DeepFace: Closing the Gap to Human-Level Performance in Face Verification](https://www.cs.toronto.edu/~ranzato/publications/taigman_cvpr14.pdf)中找到。
# ## 4.2 just coding
#
# 首先我们需要搭建一个模型,该模型输入一张图片,输出一个128维的向量。接下来的代码借鉴了facenet的开源实现[openface](https://github.com/iwantooxxoox/Keras-OpenFace),感兴趣的朋友可以去看看。
# ### 4.2.1 加载模型
# facenet的模型采用了GoogleNet的inception block,相应的实现在inception_blocks.py里面也可以找到,感兴趣的可以看看。对于模型比较重要的几件事情就是
#
# 1. 该网络采用了96*96维度的RGB图像作为输入,而且该模型采用channel first的方式,所以输入的tensor维度为(m, 3, 96, 96)
# 2. 该网络的输出维度为(m, 128),也就是将输入图片编码为128维的向量
# +
from keras.models import Sequential
from keras.layers import Conv2D, ZeroPadding2D, Activation, Input, concatenate
from keras.models import Model
from keras.layers.normalization import BatchNormalization
from keras.layers.pooling import MaxPooling2D, AveragePooling2D
from keras.layers.merge import Concatenate
from keras.layers.core import Lambda, Flatten, Dense
from keras.initializers import glorot_uniform
from keras.engine.topology import Layer
from keras import backend as K
K.set_image_data_format('channels_first')
import cv2
import os
import numpy as np
from numpy import genfromtxt
import pandas as pd
import tensorflow as tf
from openface.fr_utils import *
from openface.inception_blocks_v2 import *
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
np.set_printoptions(threshold=np.nan)
FRmodel = faceRecoModel(input_shape=(3, 96, 96))
# -
FRmodel.summary()
# ### 4.2.2 Triplet Loss
#
# 在模型输出了特征向量之后,我们需要根据4.1节介绍的Triplet Loss来计算损失。其中参数alpha代表损失函数的margin
def triplet_loss(y_true, y_pred, alpha = 0.2):
anchor, positive, negative = y_pred[0], y_pred[1], y_pred[2]
pos_dist = tf.reduce_sum(tf.square(anchor-positive), axis=-1)
neg_dist = tf.reduce_sum(tf.square(anchor-negative), axis=-1)
basic_loss = (pos_dist - neg_dist + alpha)
loss = tf.reduce_sum(tf.maximum(basic_loss, 0))
return loss
# ### 4.2.3 加载预训练权重
#
# 因为facenet是在大量的数据集中利用triplet loss训练出来的,需要极大的计算量,所以我不打算重头开始训练,这里我将直接加载预训练权重。
# +
FRmodel.compile(optimizer = 'adam', loss = triplet_loss, metrics = ['accuracy'])
load_weights_from_FaceNet(FRmodel)
# -
# ### 4.2.4 人脸验证代码(员工打卡加刷脸上班)
#
# 在前几节中已经说过了什么是人脸验证,这里在通俗地解释一下,就是你拿着一张员工卡,到公司准备打卡过闸机,打完卡之后,闸机在数据库内找到了员工卡里描述的那个人的头像信息,然后闸机需要扫描一下你的脸,来把你的脸和数据库中根据员工卡找到的那张脸进行比对,然后再决定是否让你过闸机。这就是人脸验证!
def verify(image_path, identity, database, model):
"""
image_path:image of input face
identity:your identity
database:company's data face
model:FRmodel
"""
encoding = img_to_encoding(image_path, model)
dist = np.linalg.norm(encoding-database[identity])
if dist < 0.7:
print("It's " + str(identity) + ", Welcom!")
else:
print("It's not " + str(identity) + ", please go away")
return dist
# 那么在运行这个小的程序之前,我们先要定义一个简单公司的数据库,里面存放了员工面部编码,也就是模型输出的128维向量。
# +
database = {}
database["jay1"] = img_to_encoding("openface/images/jay1.png", FRmodel)
database["jay2"] = img_to_encoding("openface/images/jay2.png", FRmodel)
# -
verify('openface/images/jay1.png', "jay2", database, FRmodel)
# ### 4.2.4 人脸识别(员工刷脸上班)
#
# 之前的那种人脸验证的闸机很麻烦(但是它也有优点,就是准确率比较高),我们可不可以不需要员工卡,只刷脸就行?当然可以了,但是现在的扫描到的人脸需要和数据库的所有人脸进行比较,求出输入和数据库中每一张图片的distance,如果所有distance均大于0.7,则表明数据库中没有改对象。否则选择distance最小的那一个作为人脸识别的结果
def who_is_it(image_path, database, model):
"""
image_path:image of input face
database:company's data face
model:FRmodel
"""
encoding = img_to_encoding(image_path, model)
min_dist = 100
for (name, db_enc) in database.items():
# Compute L2 distance between the target "encoding" and the current "emb" from the database.
dist = np.linalg.norm(encoding-db_enc)
# If this distance is less than the min_dist, then set min_dist to dist, and identity to name.
if dist < min_dist:
min_dist = dist
identity = name
### END CODE HERE ###
if min_dist > 0.7:
print("Not in the database.")
else:
print ("it's " + str(identity) + ", the distance is " + str(min_dist))
return min_dist, identity
who_is_it("openface/images/jay3.png", database, FRmodel)
# # 5 最终网络
#
# 我们已经有了构建一个人脸识别系统的所有组件,人脸检测和人脸识别,现在我们需要把这两个组件组合起来使用,输入一张图片,这张图片包含一个单个或者多个人脸(但是一般闸机一次性只通过一人,但是无所谓,我们可以在人脸检测的时候设置非最大值抑制的参数,设置输出最大数量为1即可,这样就保留了图片中最可能是人脸的图像),将这些检测到的人脸输入到我们的识别模型来获取该头像属于哪个员工?
# ## 5.1 图片模型
#
# 这是一个输入图片输出图片的人脸识别流程
# ### 5.1.1 构建数据库
#
# 数据库里面存放的是员工的图片,为了得到比较好的效果,有以下两点建议:
#
# 1. 数据库里面员工的头像图片尽量只包含脸,少一些背景干扰
# 2. 可以为同一个员工存放多张不同角度,不同表情的照片,这样也能够增加匹配的概率
#
# 你也可以更改数据库,这样就可以预测你自己的图片
# +
database = {}
database["jay1"] = img_to_encoding("openface/images/jay1.png", FRmodel)
database["jay2"] = img_to_encoding("openface/images/jay2.png", FRmodel)
database["jay3"] = img_to_encoding("openface/images/jay3.png", FRmodel)
database["jay4"] = img_to_encoding("openface/images/jay4.png", FRmodel)
# -
# ### 5.1.2 构建最终识别流程
# +
import cv2
import matplotlib.pyplot as plt
from openface.fr_utils import *
input_img_path = "images/jay.jpg"
output_img_path = "images/out/jay.jpg"
img_origin = cv2.imread(input_img_path)
img = cv2.cvtColor(img_origin,cv2.COLOR_BGR2RGB)
# predict faces for input image
out_boxes, out_scores, out_classes = a.predict(img)
# recognize faces of previous output(out_classes is unuseless, we have only one class)
for box,score in zip(out_boxes, out_scores):
""" 1. crop face according to the box """
top, left, bottom, right = box
top = int(top)
left = int(left)
bottom = int(bottom)
right = int(right)
crop_img = img[top:bottom, left:right, :]
""" 2. recognize """
encoding = img_array_to_encoding(crop_img, FRmodel)
min_dist = 100
for (name, db_enc) in database.items():
# Compute L2 distance between the target "encoding" and the current "emb" from the database.
dist = np.linalg.norm(encoding-db_enc)
# If this distance is less than the min_dist, then set min_dist to dist, and identity to name.
if dist < min_dist:
min_dist = dist
identity = name
if min_dist > 0.7:
dis_text = identity
text_color = (0xEE, 0x12, 0x89)
else:
dis_text = "Not in database"
text_color = (0x0, 0xff, 0x0)
""" 3, draw bounding box """
cv2.rectangle(img_origin, (left,top), (right,bottom), text_color,4)
cv2.putText(img_origin, dis_text, (left, top-10), cv2.FONT_HERSHEY_SIMPLEX, 1.2, text_color, 2)
cv2.imwrite(output_img_path, img_origin)
# -
# ## 5.2 视频模型
#
# 输入一个视频,输出一个视频
# ### 5.1.1 构建数据库
#
# 数据库里面存放的是员工的图片,为了得到比较好的效果,有以下两点建议:
#
# 1. 数据库里面员工的头像图片尽量只包含脸,少一些背景干扰
# 2. 可以为同一个员工存放多张不同角度,不同表情的照片,这样也能够增加匹配的概率
#
# 你也可以更改数据库,这样就可以预测你自己的图片
# +
database = []
database.append(("jay", img_to_encoding("openface/images/jay1.png", FRmodel)))
database.append(("jay", img_to_encoding("openface/images/jay2.png", FRmodel)))
database.append(("jay", img_to_encoding("openface/images/jay3.png", FRmodel)))
database.append(("jay", img_to_encoding("openface/images/jay4.png", FRmodel)))
database.append(("jay", img_to_encoding("openface/images/jay5.png", FRmodel)))
database.append(("jay", img_to_encoding("openface/images/jay6.png", FRmodel)))
database.append(("jay", img_to_encoding("openface/images/jay7.png", FRmodel)))
database.append(("jay", img_to_encoding("openface/images/jay8.png", FRmodel)))
database.append(("jay", img_to_encoding("openface/images/jay9.png", FRmodel)))
database.append(("jay", img_to_encoding("openface/images/jay10.png", FRmodel)))
database.append(("jay", img_to_encoding("openface/images/jay11.png", FRmodel)))
database.append(("jay", img_to_encoding("openface/images/jay12.png", FRmodel)))
database.append(("jay", img_to_encoding("openface/images/jay13.png", FRmodel)))
database.append(("jay", img_to_encoding("openface/images/jay14.png", FRmodel)))
database.append(("jay", img_to_encoding("openface/images/jay15.png", FRmodel)))
database.append(("jay", img_to_encoding("openface/images/jay16.png", FRmodel)))
database.append(("jay", img_to_encoding("openface/images/jay17.png", FRmodel)))
database.append(("riki", img_to_encoding("openface/images/riki1.png", FRmodel)))
database.append(("riki", img_to_encoding("openface/images/riki2.png", FRmodel)))
database.append(("riki", img_to_encoding("openface/images/riki3.png", FRmodel)))
database.append(("riki", img_to_encoding("openface/images/riki4.png", FRmodel)))
database.append(("riki", img_to_encoding("openface/images/riki5.png", FRmodel)))
# -
# ### 5.1.2 构建最终识别流程
# +
import cv2
import numpy as np
import time
import matplotlib.pyplot as plt
# %matplotlib inline
def face_reco(video_name):
input_video_path = "video/" + video_name
output_video_path = "video/out/" + video_name
# read video
cap = cv2.VideoCapture(input_video_path)
if(cap.isOpened() == False):
print ("Failed to open " + input_video_path)
assert(False)
video_width = cap.get(cv2.CAP_PROP_FRAME_WIDTH)
video_high = cap.get(cv2.CAP_PROP_FRAME_HEIGHT)
video_fps = cap.get(cv2.CAP_PROP_FPS)
# create video writer
fourcc = cv2.VideoWriter_fourcc(*'XVID')
writer = cv2.VideoWriter(output_video_path, fourcc, video_fps, (int(video_width), int(video_high)))
while True:
# get fram
ret, frame = cap.read()
# check if the video is over
if(ret != True):
print ("Complete!")
break
img_origin = frame
img = cv2.cvtColor(img_origin,cv2.COLOR_BGR2RGB)
print (img.shape)
# predict faces for input image
out_boxes, out_scores, out_classes = a.predict(img)
# recognize faces of previous output(out_classes is unuseless, we have only one class)
for box,score in zip(out_boxes, out_scores):
""" 1. crop face according to the box """
top, left, bottom, right = box
top = max(int(top), 0)
left = max(int(left), 0)
bottom = int(bottom)
right = int(right)
crop_img = img[top:bottom, left:right, :]
""" 2. recognize """
print(top, bottom, left, right)
encoding = img_array_to_encoding(crop_img, FRmodel)
min_dist = 100
for (name, db_enc) in database:
# Compute L2 distance between the target "encoding" and the current "emb" from the database.
dist = np.linalg.norm(encoding-db_enc)
# If this distance is less than the min_dist, then set min_dist to dist, and identity to name.
if dist < min_dist:
min_dist = dist
identity = name
if min_dist <= 0.7:
dis_text = identity
text_color = (0xEE, 0x12, 0x89)
else:
dis_text = "Not in database"
text_color = (0x0, 0xff, 0x0)
""" 3, draw bounding box """
cv2.rectangle(img_origin, (left,top), (right,bottom), text_color,4)
cv2.putText(img_origin, dis_text, (left, top-10), cv2.FONT_HERSHEY_SIMPLEX, 1.2, text_color, 2)
# generate video
writer.write(img_origin)
if cv2.waitKey(1) & 0xFF == ord('q'):
break
# clean
cap.release()
cv2.destroyAllWindows()
# +
video_path = "zhou1.mp4"
face_reco(video_path)
# -
| YOLO_face_detection_and_recognition_3--YAD2K_and_facenet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# http://www.inf.u-szeged.hu/~szepet/python/tip.py
#
# https://scikit-fuzzy.readthedocs.io/en/latest/auto_examples/plot_control_system_advanced.html
#
# https://pythonhosted.org/scikit-fuzzy/api/skfuzzy.membership.html
#
# https://github.com/scikit-fuzzy/scikit-fuzzy/blob/master/skfuzzy/control/tests/test_controlsystem.py
# ## Attempt-1 uisng `mf=3`
import skfuzzy as fuzz
from skfuzzy import control as ctrl
import numpy as np
import matplotlib.pyplot as plt
# +
w1 = ctrl.Antecedent(np.arange(0, 100, 10), 'B_leaf')
w1.automf(3) # Available options: 'poor'; 'average', or 'good'
w2 = ctrl.Antecedent(np.arange(0, 100, 10), 'Peels')
w2.automf(3)
w3 = ctrl.Antecedent(np.arange(0, 100, 10), 'Bones')
w3.automf(3)
speed = ctrl.Consequent(np.arange(0, 1400), 'speed')
# speed.automf(3) # Available options: 'poor'; 'mediocre'; 'average'; 'decent', or 'good'
temp = ctrl.Consequent(np.arange(0, 2000), 'temp')
# temp.automf(3)
# +
# mf for
# speed['lowest'] = fuzz.trimf(speed.universe, [0, 150, 300])
speed['low'] = fuzz.trapmf(speed.universe, [0, 200, 400, 600])
speed['medium'] = fuzz.trapmf(speed.universe, [400, 600, 800, 1000])
speed['high'] = fuzz.trapmf(speed.universe, [800, 1000, 1200, 1400])
# speed['highest'] = fuzz.trimf(speed.universe, [800, 950, 1100])
# mf for temperature
# temp['lowest'] = fuzz.trimf(temp.universe, [0, 300, 600])
temp['low'] = fuzz.trimf(temp.universe, [0, 0, 600])
temp['medium'] = fuzz.trimf(temp.universe, [250, 800, 1300])
temp['high'] = fuzz.trimf(temp.universe, [900, 1450, 2000])
# temp['highest'] = fuzz.trimf(temp.universe, [1600, 1900, 2200])
# -
# temp['low'] = fuzz.trimf(temp.universe, [0, 400, 800])
# temp['medium'] = fuzz.trimf(temp.universe, [500, 900, 1300])
# temp['high'] = fuzz.trimf(temp.universe, [1000, 1400, 1800])
# +
# Antecedent[w1, w2, w3] --> Available options: 'poor'; 'average', or 'good'
# Condequent[speed, temp] --> Available options: 'low', 'medium', 'high'
r1 = ctrl.Rule(w1['good'] | w2['good'] | w3['poor'], speed['high'])
r2 = ctrl.Rule(w3['average'] | w3['good'], speed['low'])
r3 = ctrl.Rule(w1['good'] & w2['average'] | w3['poor'], speed['high'])
r4 = ctrl.Rule(w2['average'] & w3['average'], speed['medium'])
r5 = ctrl.Rule(w1['good'] | w2['good'] | w3['poor'], temp['high'])
r6 = ctrl.Rule(w1['poor'] | w2['poor'] & w3['good'], temp['low'])
r7 = ctrl.Rule(w1['poor'] | w2['average'] | w3['average'], temp['medium'])
r8 = ctrl.Rule(w1['poor'] | w2['poor'] & w3['average'], temp['low'])
# -
velocity_ctrl = ctrl.ControlSystem([r1, r2, r3, r4])
velocity = ctrl.ControlSystemSimulation(velocity_ctrl)
velocity.input['B_leaf'] = 40
velocity.input['Peels'] = 20
velocity.input['Bones'] = 40
velocity.compute()
velocity.output['speed']
temperature_ctrl = ctrl.ControlSystem([r5, r6, r7, r8])
temperature = ctrl.ControlSystemSimulation(temperature_ctrl)
temperature.input['B_leaf'] = 40
temperature.input['Peels'] = 20
temperature.input['Bones'] = 40
temperature.compute()
temperature.output['temp']
speed.view()
temp.view()
import warnings
warnings.filterwarnings('ignore')
speed.view(sim=velocity)
print("Speed output by fuzzy: {: .2f} RPM".format(velocity.output['speed']), end=' -- ')
# speed.view()
temp.view(sim=temperature)
print(("Temperature by fuzzy: {: .2f}" + u" \N{DEGREE SIGN}" +"C").format(temperature.output['temp']))
temp.view(sim=temperature)
print(("{:.2f}" + u" \N{DEGREE SIGN}" +"C").format(temperature.output['temp']))
# temp.view()
'''
* dismal
* poor
* mediocre
* average (always middle)
* decent
* good
* excellent
and for ``'quant'`` as::
* lowest
* lower
* low
* average (always middle)
* high
* higher
* highest
'''
# +
import numpy as np
from skfuzzy.control.controlsystem import (
Antecedent, Consequent, ControlSystem, ControlSystemSimulation,
CrispValueCalculator, Rule)
def test_crisp_value_calculator_1():
x1 = Antecedent(np.linspace(0, 10, 11), "x1")
x1.automf(3) # term labels: poor, average, good
x2 = Antecedent(np.linspace(0, 10, 11), "x2")
x2.automf(3)
y1 = Consequent(np.linspace(0, 10, 11), "y1")
y1.automf(3)
y2 = Consequent(np.linspace(0, 10, 11), "y2")
y2.automf(3)
r1 = Rule(x1["poor"], y1["good"])
r2 = Rule(x2["poor"], y2["good"])
sys = ControlSystem([r1, r2])
sim = ControlSystemSimulation(sys)
cvc = CrispValueCalculator(x1, sim)
cvc.fuzz(0)
values = {label: term.membership_value[sim]
for label, term in x1.terms.items()}
assert values == {
"poor": 1,
"average": 0,
"good": 0,
}
cvc.fuzz(2.5)
values = {label: term.membership_value[sim]
for label, term in x1.terms.items()}
assert values == {
"poor": .5,
"average": .5,
"good": 0,
}
cvc.fuzz(5)
values = {label: term.membership_value[sim]
for label, term in x1.terms.items()}
assert values == {
"poor": 0,
"average": 1,
"good": 0,
}
# -
test_crisp_value_calculator_1()
# +
# time for heating ouput
import numpy as np
import skfuzzy as fuzz
from skfuzzy import control as ctrl
temprature = ctrl.Antecedent(np.arange(0, 2001, 200), 'temperature')
moisture = ctrl.Antecedent(np.arange(0, 11, 1), 'moisture')
minutes = ctrl.Consequent(np.arange(0,26, 1), 'minutes')
temprature.automf(3)
moisture.automf(3)
minutes['low'] = fuzz.trimf(minutes.universe, [0, 0, 13])
minutes['medium'] = fuzz.trimf(minutes.universe, [0, 13, 25])
minutes['high'] = fuzz.trimf(minutes.universe, [13, 25, 25])
rule1 = ctrl.Rule(temprature['poor'] | moisture['poor'] , minutes['low'])
rule2 = ctrl.Rule(temprature['average'] | moisture['average'], minutes['medium'])
rule3 = ctrl.Rule(temprature['good'] | moisture['good'], minutes['high'])
tap_ctrl = ctrl.ControlSystem([rule1, rule2, rule3])
tap_control_system = ctrl.ControlSystemSimulation(tap_ctrl)
tap_control_system.input['temperature'] = 1500
tap_control_system.input['moisture'] = 1
tap_control_system.compute()
print(tap_control_system.output['minutes'], minutes.view(sim=tap_control_system))
# -
# ## Looping over inputs to cal time required for heating
# +
import numpy as np
import skfuzzy as fuzz
from skfuzzy import control as ctrl
import random, time
def init():
quality = ctrl.Antecedent(np.arange(0, 11, 1), 'quality')
service = ctrl.Antecedent(np.arange(0, 11, 1), 'service')
tip = ctrl.Consequent(np.arange(0, 26, 1), 'tip')
quality.automf(3)
service.automf(3)
tip['low'] = fuzz.trimf(tip.universe, [0, 0, 13])
tip['medium'] = fuzz.trimf(tip.universe, [0, 13, 25])
tip['high'] = fuzz.trimf(tip.universe, [13, 25, 25])
rule1 = ctrl.Rule(quality['poor'] | service['poor'], tip['low'])
rule2 = ctrl.Rule(service['average'], tip['medium'])
rule3 = ctrl.Rule(service['good'] | quality['good'], tip['high'])
#quality.automf(7) ##If I put new membership function here, I will get an error
'''
#I have to define the rules again
'''
global tipping_ctrl
tipping_ctrl = ctrl.ControlSystem([rule1, rule2, rule3])
def calc(quality,service):
#I would like to change the control system( membership function) here, but how?
tipping = ctrl.ControlSystemSimulation(tipping_ctrl)
tipping.input['quality'] = quality
tipping.input['service'] = service
tipping.compute()
print(tipping.output['tip'])
if __name__ == '__main__':
init()
count = 0
while count <=10:
quality = float(random.randint(0,11))
service = float(random.randint(0,11))
calc(quality,service)
count += 1
time.sleep(1)
# -
# ## 3D graph output
# +
from skfuzzy import control as ctrl
from mpl_toolkits.mplot3d import Axes3D # Required for 3D plotting
# New Antecedent/Consequent objects hold universe variables and membership
# functions
quality = ctrl.Antecedent(np.arange(0, 10, 0.1), 'quality')
service = ctrl.Antecedent(np.arange(0, 10, 0.1), 'service')
tip = ctrl.Consequent(np.arange(0, 25, 0.1), 'tip')
quality['poor'] = fuzz.zmf(quality.universe, 0,5)
quality['average'] = fuzz.gaussmf(quality.universe,5,1)
quality['good'] = fuzz.smf(quality.universe,5,10)
service['poor'] = fuzz.zmf(service.universe, 0,5)
service['average'] = fuzz.gaussmf(service.universe,5,1)
service['good'] = fuzz.smf(service.universe,5,10)
tip['low'] = fuzz.trimf(tip.universe, [0, 0, 13])
tip['medium'] = fuzz.trimf(tip.universe, [0, 13, 25])
tip['high'] = fuzz.trimf(tip.universe, [13, 25, 25])
rule1 = ctrl.Rule(quality['poor'] | service['poor'], tip['low'])
rule2 = ctrl.Rule(service['average'], tip['medium'])
rule3 = ctrl.Rule(service['good'] | quality['good'], tip['high'])
tipping_ctrl = ctrl.ControlSystem([rule1, rule2, rule3])
tipping = ctrl.ControlSystemSimulation(tipping_ctrl)
# We can simulate at higher resolution with full accuracy
upsampled = np.linspace(0, 10, 10)
x, y = np.meshgrid(upsampled, upsampled)
z = np.zeros_like(x)
# Loop through the system 10*10 times to collect the control surface
for i in range(10):
for j in range(10):
tipping.input['quality'] = x[i, j]
tipping.input['service'] = y[i, j]
tipping.compute()
z[i, j] = tipping.output['tip']
# Plot the result in pretty 3D with alpha blending
fig = plt.figure(figsize=(8, 8))
ax = fig.add_subplot(111, projection='3d')
surf = ax.plot_surface(x, y, z, rstride=1, cstride=1, cmap='viridis',
linewidth=0.4, antialiased=True)
cset = ax.contourf(x, y, z, zdir='z', offset=-2.5, cmap='viridis', alpha=0.5)
cset = ax.contourf(x, y, z, zdir='x', offset=13, cmap='viridis', alpha=0.5)
cset = ax.contourf(x, y, z, zdir='y', offset=13, cmap='viridis', alpha=0.5)
ax.view_init(30, 200)
plt.show()
plt.ioff()
# -
# ## Attempt-2 with `mf=5`
# +
w1 = ctrl.Antecedent(np.arange(0, 100, 10), 'B_leaf')
w1.automf(3) # Available options: 'poor'; 'average', or 'good'
w2 = ctrl.Antecedent(np.arange(0, 100, 10), 'Peels')
w2.automf(3)
w3 = ctrl.Antecedent(np.arange(0, 100, 10), 'Bones')
w3.automf(3)
speed = ctrl.Consequent(np.arange(0, 1400), 'speed')
# speed.automf(5) # Available options: 'poor'; 'mediocre'; 'average'; 'decent', or 'good'
temp = ctrl.Consequent(np.arange(0, 2000), 'temp')
# temp.automf(5)
# +
# mf for
speed['lowest'] = fuzz.trapmf(speed.universe, [0, 0, 100, 300])
speed['low'] = fuzz.trapmf(speed.universe, [200, 400, 600, 750])
speed['medium'] = fuzz.trapmf(speed.universe, [500, 700, 900, 1050])
speed['high'] = fuzz.trapmf(speed.universe, [850, 1000, 1200, 1350])
speed['highest'] = fuzz.trapmf(speed.universe, [1050, 1300, 1500, 1650])
# mf for temperature
temp['lowest'] = fuzz.trimf(temp.universe, [0, 300, 600])
temp['low'] = fuzz.trimf(temp.universe, [400, 700, 1000])
temp['medium'] = fuzz.trimf(temp.universe, [800, 1100, 1400])
temp['high'] = fuzz.trimf(temp.universe, [1200, 1500, 1800])
temp['highest'] = fuzz.trimf(temp.universe, [1600, 1900, 2200])
# +
# Antecedent[w1, w2, w3] --> Available options: 'poor'; 'average', or 'good'
# Condequent[speed, temp] --> Available options: 'low', 'medium', 'high'
r1 = ctrl.Rule(w1['good'] | w2['good'] | w3['poor'], speed['highest'])
r2 = ctrl.Rule(w3['average'] | w3['good'], speed['lowest'])
r3 = ctrl.Rule(w1['good'] & w2['average'] | w3['poor'], speed['high'])
r4 = ctrl.Rule(w2['average'] & w3['average'], speed['medium'])
r41 = ctrl.Rule(w1['poor'] | w2['poor'] & w3['average'], speed['low'])
r5 = ctrl.Rule(w1['good'] | w2['good'] | w3['poor'], temp['high'])
r6 = ctrl.Rule(w1['poor'] | w2['poor'] & w3['good'], temp['low'])
r7 = ctrl.Rule(w1['poor'] | w2['average'] | w3['average'], temp['medium'])
r8 = ctrl.Rule(w1['poor'] | w2['poor'] & w3['average'], temp['low'])
# -
velocity_ctrl = ctrl.ControlSystem([r1, r2, r3, r4])
velocity = ctrl.ControlSystemSimulation(velocity_ctrl)
velocity.input['B_leaf'] = 20
velocity.input['Peels'] = 20
velocity.input['Bones'] = 60
velocity.compute()
velocity.output['speed']
temperature_ctrl = ctrl.ControlSystem([r5, r6, r7, r8])
temperature = ctrl.ControlSystemSimulation(temperature_ctrl)
temperature.input['B_leaf'] = 60
temperature.input['Peels'] = 0
temperature.input['Bones'] = 40
temperature.compute()
temperature.output['temp']
speed.view(sim=velocity)
# plt.ion()
# plt.show()
# plt.ioff()
# speed.view()
temp.view(sim=temperature)
# temp.view()
| Fuzz_simulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# ## Map Mass2Motifs on mass spectral molecular networks (feature based workflow)
# download MolNetEnhancer package
devtools::install_github("madeleineernst/RMolNetEnhancer", force = T)
# download rCSCS package, for direct import of GNPS data (https://www.biorxiv.org/content/10.1101/546150v1)
devtools::install_github("askerdb/rCSCS", force = T)
# load libraries
library('igraph')
# <font color='blue'>**specify GNPS job ID**</font>
gnps_id = "b817262cb6114e7295fee4f73b22a3ad"
folder_name = "AGP_Example"
# download GNPS data
rCSCS::download_GNPS(gnps_id, folder_name)
edges <- read.csv(paste(folder_name,gnps_id,'edges_file.txt',sep='/'), sep = '\t')
head(edges)
# <font color='blue'>**specify MS2LDA job ID**</font>
ms2lda_id = '907'
motifs <- read.csv(paste('http://ms2lda.org/basicviz/get_gnps_summary',ms2lda_id, sep = "/"))
head(motifs)
# create network data with mapped motifs
motif_network <- MolNetEnhancer::Mass2Motif_2_Network(edges,motifs,prob = 0.01,overlap = 0.3, top = 5)
head(motif_network$edges)
head(motif_network$nodes)
# write network data with mapped motifs to files: <br>
# The <i>edges</i> file can be importet as network into Cytoscape, whereas the <i>nodes</i> file can be imported as table. Select column 'CLUSTERID1' as Source Node, column 'interact' as Interaction Type and 'CLUSTERID2' as Target Node.
write.table(motif_network$edges,"Mass2Motifs_Edges_FeatureBased.tsv",quote=F,row.names = F,sep="\t")
write.table(motif_network$nodes,"Mass2Motifs_Nodes_FeatureBased.tsv",quote=F,row.names = F,sep="\t")
# create graphML file containing motif information
MG <- MolNetEnhancer::make_motif_graphml(motif_network$nodes,motif_network$edges)
# write graphML file containing motif information
write_graph(MG, 'Motif_Network_FeatureBased.graphml', format = "graphml")
# ## map chemical class information
# read in chemical class information (to create this file follow descriptions in ChemicalClasses_2_Network.ipynb)
final <- read.table('ClassyFireResults_Network.txt', sep = "\t", header = T, check.names = F, stringsAsFactors = F)
head(final)
# create graphML file containing motif and chemical class information
graphML_classy <- MolNetEnhancer::make_classyfire_graphml(MG, final)
# write graphML file containing motif and chemical class information
write_graph(graphML_classy, 'Motif_ChemicalClass_Network_FeatureBased.graphml', format = "graphml")
| Example_notebooks/Mass2Motifs_2_Network_FeatureBased.ipynb |