code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8
# language: python
# name: python3
# ---
# # Country Economic Conditions for Cargo Carriers (Machine Learning with Classification Models)
# This report is written from the point of view of a data scientist preparing a report to the Head of Analytics for a logistics company. The company needs information on economic and financial conditions is different countries, including data on their international trade, to be aware of any situations that could affect business.
# ## Objective
# The objective of this report is to determine what factors may be driving investment in a country. For our company, it is important to understand the drivers of global growth and investment, as it allows us to plan ahead for where there may be greater demand for cargo services in the future. We want to positions ourselves as the go-to logistics company globally. Positioning ourselves in growing trade hubs will serve the long-term strategic objectives of our global services.
# ## Data Summary
# This dataset is taken from the International Monetary Fund (IMF) data bank. It lists country-level economic and financial statistics from all countries globally. This includes data such as gross domestic product (GDP), inflation, exports and imports, and government borrowing and revenue. The data is given in either US Dollars, or local currency depending on the country and year. Some variables, like inflation and unemployment, are given as percentages.
# ## Data Exploration
#Import required packages
import numpy as np
import pandas as pd
from sklearn import linear_model
from scipy import stats
from numpy import *
from scipy.stats.mstats import normaltest
import math
from sklearn import datasets, linear_model
from sklearn.linear_model import LinearRegression
import statsmodels.api as sm
from sklearn.metrics import r2_score
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import (StandardScaler,
PolynomialFeatures)
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
import warnings
warnings.simplefilter("ignore")
#Import IMF World Economic Outlook Data from GitHub
WEO = pd.read_csv('https://raw.githubusercontent.com/jamiemfraser/machine_learning/main/WEOApr2021all.csv')
WEO=pd.DataFrame(WEO)
WEO.head()
# +
#We are only interested in the most recent year for which data is available, 2019
WEO=WEO.drop(['2000', '2001', '2002', '2003', '2004', '2005', '2006', '2007', '2008', '2009', '2010', '2011', '2012', '2013', '2014', '2015', '2016', '2017', '2018'], axis = 1)
#Reshape the data so each country is one observation
WEO=WEO.pivot_table(index=["Country"], columns='Indicator', values='2019').reset_index()
WEO.dropna(inplace=True)
WEO.head()
# -
WEO.columns = ['Country', 'Current_account', 'Employment', 'Net_borrowing', 'Government_revenue', 'Government_expenditure', 'GDP_percap_constant', 'GDP_percap_current', 'GDP_constant', 'Inflation', 'Investment', 'Unemployment', 'Volume_exports', 'Volume_imports']
WEO.head()
# +
#Generate boolean variable for investment, indicatng whether investment is higher or lower than global mean
WEO.describe()
#Global mean for investment is 23.094944
WEO['InvestMean'] = np.where(WEO['Investment'] >=23.094944, 1, 0)
WEO = WEO.drop(["Country", "Volume_exports", "Volume_imports", "GDP_percap_current", "Government_revenue", "Government_expenditure", "Employment", "Unemployment"], axis=1)
WEO.head()
# -
WEO.dtypes
# +
#Describe the distribution of InvestMean variable
WEO.InvestMean.hist()
#We now have the target variable as a boolean, where 1 = True and 0 = False, for use in classifer models
# -
# Calculate the correlation values
feature_cols = WEO.columns[0:4]
corr_values = WEO[feature_cols].corr()
# Simplify by emptying all the data below the diagonal
tril_index = np.tril_indices_from(corr_values)
# Make the unused values NaNs
for coord in zip(*tril_index):
corr_values.iloc[coord[0], coord[1]] = np.NaN
# Stack the data and convert to a data frame
corr_values = (corr_values
.stack()
.to_frame()
.reset_index()
.rename(columns={'level_0':'feature1',
'level_1':'feature2',
0:'correlation'}))
corr_values['abs_correlation'] = corr_values.correlation.abs()
#Sort the correlation values from largest to smallest
corr_values.sort_values('correlation', ascending=False).query('abs_correlation>=0.0')
# ## Classifier Models
# The three models that I will use for this analysis are
#
# 1. Logistic regression
# 2. K Nearest neighbors
# 3. Decision Tree
# ### Model 1: Logistic regression
WEO.reset_index(inplace=True)
WEO.sort_index()
WEO.head()
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
WEO['InvestMean'] = le.fit_transform(WEO.InvestMean)
WEO['InvestMean'].sample(5)
# +
from sklearn.model_selection import StratifiedShuffleSplit
# Get the split indexes
strat_shuf_split = StratifiedShuffleSplit(n_splits=1, test_size=0.3, random_state=0)
train_idx, test_idx = next(strat_shuf_split.split(WEO[feature_cols], WEO.InvestMean))
# Create the dataframes
X_train = WEO.loc[train_idx, feature_cols]
y_train = WEO.loc[train_idx, 'InvestMean']
X_test = WEO.loc[test_idx, feature_cols]
y_test = WEO.loc[test_idx, 'InvestMean']
y_train.value_counts(normalize=True)
y_test.value_counts(normalize=True)
# -
y_train.value_counts(normalize=True)
y_test.value_counts(normalize=True)
from sklearn.linear_model import LogisticRegression
# Standard logistic regression
lr = LogisticRegression(solver='liblinear').fit(X_train, y_train)
# ### Model 2: K-Nearest Neighbor
df_uniques = pd.DataFrame([[i, len(WEO[i].unique())] for i in WEO.columns], columns=['Variable', 'Unique Values']).set_index('Variable')
df_uniques
from sklearn.preprocessing import LabelBinarizer, LabelEncoder, OrdinalEncoder
lb, le = LabelBinarizer(), LabelEncoder()
WEO['InvestMean'] = lb.fit_transform(WEO['InvestMean'])
from sklearn.preprocessing import MinMaxScaler
mm = MinMaxScaler()
for column in [feature_cols]:
WEO[column] = mm.fit_transform(WEO[column])
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import confusion_matrix, accuracy_score, classification_report, f1_score
y, X = WEO['InvestMean'], WEO.drop(columns='InvestMean')
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=42)
knn = KNeighborsClassifier(n_neighbors=3)
knn = knn.fit(X_train, y_train)
y_pred = knn.predict(X_test)
print(classification_report(y_test, y_pred))
print('Accuracy score: ', round(accuracy_score(y_test, y_pred), 2))
print('F1 Score: ', round(f1_score(y_test, y_pred), 2))
# ### Model 3: Decision Tree
import os, pandas as pd, numpy as np, matplotlib.pyplot as plt, seaborn as sns
from sklearn.tree import DecisionTreeClassifier
dt = DecisionTreeClassifier(random_state=42)
dt = dt.fit(X_train, y_train)
dt.tree_.node_count, dt.tree_.max_depth
y_train.value_counts(normalize=True).sort_index()
y_test.value_counts(normalize=True).sort_index()
from sklearn.metrics import accuracy_score, precision_score, recall_score, f1_score
def measure_error(y_true, y_pred, label):
return pd.Series({'accuracy':accuracy_score(y_true, y_pred),
'precision': precision_score(y_true, y_pred),
'recall': recall_score(y_true, y_pred),
'f1': f1_score(y_true, y_pred)},
name=label)
y_train_pred = dt.predict(X_train)
y_test_pred = dt.predict(X_test)
train_test_full_error = pd.concat([measure_error(y_train, y_train_pred, 'train'),
measure_error(y_test, y_test_pred, 'test')],
axis=1)
train_test_full_error
from sklearn.model_selection import GridSearchCV
param_grid = {'max_depth':range(1, dt.tree_.max_depth+1, 2),
'max_features': range(1, len(dt.feature_importances_)+1)}
GR = GridSearchCV(DecisionTreeClassifier(random_state=42),
param_grid=param_grid,
scoring='accuracy',
n_jobs=-1)
GR = GR.fit(X_train, y_train)
GR.best_estimator_.tree_.node_count, GR.best_estimator_.tree_.max_depth
y_train_pred_gr = GR.predict(X_train)
y_test_pred_gr = GR.predict(X_test)
train_test_gr_error = pd.concat([measure_error(y_train, y_train_pred_gr, 'train'),
measure_error(y_test, y_test_pred_gr, 'test')],
axis=1)
train_test_gr_error
# ### Recommendation
# Based on the results of the analysis above, I would suggest to use the decision tree classifier to achieve the best results.
# ## Key Findings
# The key finding of this analysis is that it suffers from a lack of data. There are too few data points to make a meaningful classification model that could be generalized. However, given the data that we have, I can say that it is clear there are many other factors that determine investment that are not captured by the data we have access to.
# ## Next Steps
# The next steps would be, if possible, to obtain further data that could complement what we already have. If not, then we would have to consider a different class of model to obtain the insights we need.
| Country_Economic_Conditions_Part_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Card and Deck objects
#
# This notebook contains example code from [*Fluent Python*](http://shop.oreilly.com/product/0636920032519.do), by <NAME>.
#
# Code by <NAME>, modified by <NAME>.
#
# MIT License: https://opensource.org/licenses/MIT
# This example demonstrates the Python data model using a simple implementation of playing cards and decks.
#
# `Card` is a namedtuple that represents a playing card.
# +
import collections
Card = collections.namedtuple('Card', ['rank', 'suit'])
# -
# `FrenchDeck` is a class that represents a deck of cards.
class FrenchDeck:
ranks = [str(n) for n in range(2, 11)] + list('JQKA')
suits = 'spades diamonds clubs hearts'.split()
def __init__(self):
self._cards = [Card(rank, suit) for suit in self.suits
for rank in self.ranks]
def __len__(self):
return len(self._cards)
def __getitem__(self, position):
return self._cards[position]
# You can instantiate a `Card` object as if `Card` were a class.
#
# BTW: [beer card](https://en.wikipedia.org/wiki/Beer_card)
beer_card = Card('7', 'diamonds')
beer_card
# You can access the fields of a card by name.
print(beer_card.rank)
print(beer_card.suit)
# Or by index.
beer_card[0], beer_card[1]
# A drawback of using namedtuples is that you can't define methods for them in the usual way.
#
# But you can [monkey-patch](https://en.wikipedia.org/wiki/Monkey_patch) them by defining a function and then making it an attribute of `Card`. For example, here's a function that generates a string representation of a card:
# +
def card_to_str(card):
return '%s of %s' % card
card_to_str(beer_card)
# -
# Here's how we can make that function behave like a method. When we pass a card to `print`, Python invokes the special method `__str__`
Card.__str__ = card_to_str
print(beer_card)
# Now let's instantiate a `FrenchDeck`.
#
# When we call `len`, Python invokes the `__len__` method on the deck.
deck = FrenchDeck()
len(deck)
# When we use the bracket operator, Python invokes the `__getitem__` method:
deck[3]
# And that means that the slice operator works, too:
deck[:3]
# Aside: In this context, we don't get the string generated by `__str__`; we get the one generated by `__repr__` (read about that [here](https://docs.python.org/3/reference/datamodel.html#basic-customization))
#
# Because `FrenchDeck` provides `__len__` and `__getitem__`, it is considered a sequence, which means that the `in` operator works:
Card('Q', 'hearts') in deck
# **Exercise** Make up a card that doesn't exist and confirm that `in` returns `False`.
# Solution goes here
charmander = Card('Charmander', 'Fire' )
charmander in deck
# And the for loop works, too:
for card in deck:
print(card)
# Other methods that work with sequences, like `random.choice`, will work with decks:
from random import choice
choice(deck)
# Sadly, `shuffle` doesn't work because we haven't provided `__setitem__`, so a deck is an immutable sequence:
# +
from random import shuffle
# This should raise a TypeError
shuffle(deck)
# -
# We can use `sorted` to iterate through the cards in the order determined by tuple comparison:
for card in sorted(deck):
print(card)
# If we want an ordering that makes more sense for cards, we can define a function that maps from a card to an integer:
# +
suit_values = dict(spades=3, hearts=2, diamonds=1, clubs=0)
def spades_high_ordering(card):
rank_value = FrenchDeck.ranks.index(card.rank)
return rank_value * len(suit_values) + suit_values[card.suit]
# -
spades_high_ordering(Card('2', 'clubs'))
spades_high_ordering(Card('A', 'spades'))
# And then pass this funcition as a key to `sorted`:
for card in sorted(deck, key=spades_high_ordering):
print(card)
# **Exercise** Define a new ordering that sorts the cards by suit first and then by rank, so all clubs come first, followed by all diamonds, etc.
# +
# Solution goes here
# Aphabetical sort
alpha_suit = sorted(FrenchDeck.suits)
# Our lil function
def alt_high_ordering(card):
rank_value = FrenchDeck.ranks.index(card.rank)
return alpha_suit.index(card.suit) * len(FrenchDeck.ranks) + rank_value
# Time to print
for card in sorted(deck, key=alt_high_ordering):
print(card)
# -
# **Exercise** Write a method called `setcard` that takes a deck, an index, and a card, and assigns the card to the deck at the given position. Then monkey-patch `FrenchDeck` to provide `__setitem__` as a method. Test it by assigning a new card like this:
#
# deck[0] = Card('A', 'spades')
#
# Then shuffle the deck using `random.shuffle`.
# +
# Solution goes here
def set_card(deck, index, card):
deck._cards[index] = card
FrenchDeck.__setitem__ = set_card
# Time to check
deck = FrenchDeck()
deck[0] = Card('A', 'spades')
deck[0]
# -
# We should have two Aces of spades now, which we can confirm by checking the number of unique cards:
len(set(deck))
| 01-data-model/frenchdeck.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++14
// language: C++14
// name: xcpp14
// ---
// [](https://lab.mlpack.org/v2/gh/mlpack/examples/master?urlpath=lab%2Ftree%2Freinforcement_learning_gym%2Fbipedal_walker_sac%2Fbipedal_walker_sac.ipynb)
//
// You can easily run this notebook at https://lab.mlpack.org/
//
// Here, we train a [Soft Actor-Critic](https://arxiv.org/abs/1801.01290) agent to get high scores for the [Bipedal Walker](https://gym.openai.com/envs/BipedalWalker-v2/) environment.
//
// We make the agent train and test on OpenAI Gym toolkit's GUI interface provided through a distributed infrastructure (TCP API). More details can be found [here](https://github.com/zoq/gym_tcp_api).
//
// A video of the trained agent can be seen in the end.
// ## Including necessary libraries and namespaces
#include <mlpack/core.hpp>
#include <mlpack/methods/ann/ffn.hpp>
#include <mlpack/methods/reinforcement_learning/sac.hpp>
#include <mlpack/methods/ann/loss_functions/empty_loss.hpp>
#include <mlpack/methods/ann/init_rules/gaussian_init.hpp>
#include <mlpack/methods/reinforcement_learning/environment/env_type.hpp>
#include <mlpack/methods/reinforcement_learning/training_config.hpp>
// Used to run the agent on gym's environment (provided externally) for testing.
#include <gym/environment.hpp>
// Used to generate and display a video of the trained agent.
#include "xwidgets/ximage.hpp"
#include "xwidgets/xvideo.hpp"
#include "xwidgets/xaudio.hpp"
using namespace mlpack;
using namespace mlpack::ann;
using namespace ens;
using namespace mlpack::rl;
// ## Initializing the agent
// Set up the state and action space.
ContinuousActionEnv::State::dimension = 24;
ContinuousActionEnv::Action::size = 4;
// +
// Set up the actor and critic networks.
FFN<EmptyLoss<>, GaussianInitialization>
policyNetwork(EmptyLoss<>(), GaussianInitialization(0, 0.01));
policyNetwork.Add(new Linear<>(ContinuousActionEnv::State::dimension, 128));
policyNetwork.Add(new ReLULayer<>());
policyNetwork.Add(new Linear<>(128, 128));
policyNetwork.Add(new ReLULayer<>());
policyNetwork.Add(new Linear<>(128, ContinuousActionEnv::Action::size));
policyNetwork.Add(new TanHLayer<>());
policyNetwork.ResetParameters();
FFN<EmptyLoss<>, GaussianInitialization>
qNetwork(EmptyLoss<>(), GaussianInitialization(0, 0.01));
qNetwork.Add(new Linear<>(ContinuousActionEnv::State::dimension + ContinuousActionEnv::Action::size, 128));
qNetwork.Add(new ReLULayer<>());
qNetwork.Add(new Linear<>(128, 128));
qNetwork.Add(new ReLULayer<>());
qNetwork.Add(new Linear<>(128, 1));
qNetwork.ResetParameters();
// -
// Set up the replay method.
RandomReplay<ContinuousActionEnv> replayMethod(32, 10000);
// Set up training configurations.
TrainingConfig config;
config.ExplorationSteps() = 3200;
config.TargetNetworkSyncInterval() = 1;
config.UpdateInterval() = 1;
// In the cell below, we load a pretrained model by manually assigning values to the parameters of the network, after loading the parameters from their respective files `sac_q.txt` and `sac_policy.txt`.
//
// The model was trained for 620 episodes.
arma::mat temp;
data::Load("sac_q.txt", temp);
qNetwork.Parameters() = temp.t();
data::Load("sac_policy.txt", temp);
policyNetwork.Parameters() = temp.t();
// You can train the model from scratch by running the following:
// ```c++
// // Set up Soft actor-critic agent.
// SAC<ContinuousActionEnv, decltype(qNetwork), decltype(policyNetwork), AdamUpdate>
// agent(config, qNetwork, policyNetwork, replayMethod);
//
// const std::string environment = "BipedalWalker-v3";
// const std::string host = "127.0.0.1";
// const std::string port = "4040";
//
// Environment env(host, port, environment);
//
// std::vector<double> returnList;
// size_t episodes = 0;
// bool converged = true;
// size_t consecutiveEpisodesTest = 50;
// while (true)
// {
// double episodeReturn = 0;
// env.reset();
// size_t steps = 0;
// do
// {
// agent.State().Data() = env.observation;
// agent.SelectAction();
// arma::mat action = {agent.Action().action};
//
// env.step(action);
// ContinuousActionEnv::State nextState;
// nextState.Data() = env.observation;
//
// replayMethod.Store(agent.State(), agent.Action(), env.reward, nextState, env.done, 0.99);
// episodeReturn += env.reward;
// agent.TotalSteps()++;
// steps++;
// if (agent.Deterministic() || agent.TotalSteps() < config.ExplorationSteps())
// continue;
// for (size_t i = 0; i < config.UpdateInterval(); i++)
// agent.Update();
// } while (!env.done);
// returnList.push_back(episodeReturn);
// episodes += 1;
//
// if (returnList.size() > consecutiveEpisodesTest)
// returnList.erase(returnList.begin());
//
// double averageReturn = std::accumulate(returnList.begin(),
// returnList.end(), 0.0) /
// returnList.size();
//
// std::cout << "Average return in last " << returnList.size()
// << " consecutive episodes: " << averageReturn
// << " steps: " << steps
// << " Episode return: " << episodeReturn << std::endl;
//
// if (episodes % 10 == 0)
// {
// data::Save("./" + std::to_string(episodes) + "qNetwork.xml", "episode_" + std::to_string(episodes), qNetwork);
// data::Save("./" + std::to_string(episodes) + "policyNetwork.xml", "episode_" + std::to_string(episodes), policyNetwork);
// }
// if (averageReturn > -50)
// break;
// }
//
// ```
// ## Testing the trained agent
//
// It is so amazing to see how just a matrix of numbers, operated in a certain fashion, is able to develop a walking gait.
//
// Thats the beauty of Artificial Neural Networks!
// +
// Set up Soft actor-critic agent.
SAC<ContinuousActionEnv, decltype(qNetwork), decltype(policyNetwork), AdamUpdate>
agent(config, qNetwork, policyNetwork, replayMethod);
agent.Deterministic() = true;
// Creating and setting up the gym environment for testing.
gym::Environment envTest("gym.kurg.org", "4040", "BipedalWalker-v3");
envTest.monitor.start("./dummy/", true, true);
// Resets the environment.
envTest.reset();
envTest.render();
double totalReward = 0;
size_t totalSteps = 0;
// Testing the agent on gym's environment.
while (1)
{
// State from the environment is passed to the agent's internal representation.
agent.State().Data() = envTest.observation;
// With the given state, the agent selects an action according to its defined policy.
agent.SelectAction();
// Action to take, decided by the policy.
arma::mat action = {agent.Action().action};
envTest.step(action);
totalReward += envTest.reward;
totalSteps += 1;
if (envTest.done)
{
std::cout << " Total steps: " << totalSteps << "\t Total reward: "
<< totalReward << std::endl;
break;
}
// Uncomment the following lines to see the reward and action in each step.
// std::cout << " Current step: " << totalSteps << "\t current reward: "
// << totalReward << "\t Action taken: " << action;
}
envTest.close();
std::string url = envTest.url();
std::cout << url;
auto video = xw::video_from_url(url).finalize();
video
| reinforcement_learning_gym/bipedal_walker_sac/bipedal_walker_sac.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import scipy.signal
import matplotlib.pyplot as plt
import json
def load_data(filename):
with open(filename) as json_file:
data = json.load(json_file)
return np.array(data)
data = load_data("../data/круг 2.json")
# +
def data_taking(data):
vectors_accelerometer_x = []
vectors_accelerometer_y = []
vectors_accelerometer_z = []
ox = []
counter = 0
for gesture in data:
counter += 1
if counter != 1: continue
# for time_series in gesture:
for i in range(1, len(gesture)):
vectors_accelerometer_x.append(gesture[i][1])
vectors_accelerometer_y.append(gesture[i][2])
vectors_accelerometer_z.append(gesture[i][3])
ox.append(gesture[i][0])
return ox, vectors_accelerometer_x, vectors_accelerometer_y, vectors_accelerometer_z
ox, vectors_accelerometer_x, vectors_accelerometer_y, vectors_accelerometer_z = data_taking(data)
#print(len(ox))
#print(ox)
#print(vectors_accelerometer_x)
# +
def data_taking_g(data):
vectors_accelerometer_x = []
vectors_accelerometer_y = []
vectors_accelerometer_z = []
ox = []
counter = 0
for gesture in data:
counter += 1
if counter != 1: continue
# for time_series in gesture:
for i in range(1, len(gesture)):
vectors_accelerometer_x.append(gesture[i][4])
vectors_accelerometer_y.append(gesture[i][5])
vectors_accelerometer_z.append(gesture[i][6])
ox.append(gesture[i][0])
return ox, vectors_accelerometer_x, vectors_accelerometer_y, vectors_accelerometer_z
ox, vectors_accelerometer_x, vectors_accelerometer_y, vectors_accelerometer_z = data_taking_g('круг.json')
#print(len(ox))
#print(ox)
#print(vectors_accelerometer_x)
# -
def filter(order, cutoff_freq, sampling_freq, vectors_accelerometer_x, ox):
sampling_duration = int(ox[len(ox) - 1])
number_of_samples = len(vectors_accelerometer_x)
#time = np.linspace(0, sampling_duration, number_of_samples, endpoint=False)
normalized_cutoff_freq = 2 * cutoff_freq / sampling_freq
numerator_coeffs, denominator_coeffs = scipy.signal.butter(order, normalized_cutoff_freq)
filtered_signal = scipy.signal.lfilter(numerator_coeffs, denominator_coeffs, vectors_accelerometer_x)
return filtered_signal
def show_p:
# +
order = 2
sampling_freq = 30
cutoff_freq = 1.5
print(int(ox[len(ox) - 1]))
sampling_duration = int(ox[len(ox) - 1])
number_of_samples = len(vectors_accelerometer_x)
time = np.linspace(0, sampling_duration, number_of_samples, endpoint=False)
normalized_cutoff_freq = 2 * cutoff_freq / sampling_freq
numerator_coeffs, denominator_coeffs = scipy.signal.butter(order, normalized_cutoff_freq)
filtered_signal = scipy.signal.lfilter(numerator_coeffs, denominator_coeffs, vectors_accelerometer_x)
plt.figure(figsize=(20, 16))
plt.style.use('fivethirtyeight')
plt.plot(ox, vectors_accelerometer_x, 'm-', label='сырые данные')
plt.plot(ox, filtered_signal, 'c-', linewidth=4, label='отфильтрованные')
plt.legend(loc=1, prop={'size': 24})
plt.title('Данные с акселерометра по оси ' + 'X жест круг 2')
plt.ylabel('значения в м/с^2')
plt.xlabel('время в миллисекундах')
#plt.legend()
plt.show()
print(len(ox))
print(len(vectors_accelerometer_x))
print(len(filtered_signal))
print(plt.style.available)
# -
def show_p(name_of_pic, number_of_pic, color1, color2, name_ox,
order, cutoff_freq, sampling_freq, vectors_accelerometer_x, ox):
filtered_signal = filter(order, cutoff_freq, sampling_freq, vectors_accelerometer_x, ox)
plt.figure(figsize=(20, 16))
plt.style.use('fivethirtyeight')
plt.plot(ox, vectors_accelerometer_x, 'm-', label='сырые данные')
plt.plot(ox, filtered_signal, 'c-', linewidth=4, label='отфильтрованные')
plt.legend(loc=1, prop={'size': 24})
plt.title('Данные с акселерометра по оси ' + name_ox + ' жест ' + name_of_pic + ' ' + number_of_pic)
plt.ylabel('значения в м/с^2')
plt.xlabel('время в миллисекундах')
plt.show()
# +
filter(1, 2, 30, vectors_accelerometer_x, ox)
filtered_signal = scipy.signal.lfilter(numerator_coeffs, denominator_coeffs, vectors_accelerometer_x)
plt.figure(figsize=(20, 16))
plt.plot(ox, vectors_accelerometer_x, 'b-', label='signal')
plt.plot(ox, filtered_signal, 'g-', linewidth=2, label='filtered signal')
plt.legend()
plt.style.use('seaborn-muted')
plt.show()
# -
# +
order = 5
sampling_freq = 30
cutoff_freq = 2
print(int(ox[len(ox) - 1]))
sampling_duration = int(ox[len(ox) - 1])
number_of_samples = len(vectors_accelerometer_x)
time = np.linspace(0, sampling_duration, number_of_samples, endpoint=False)
#signal = np.sin(2*np.pi*time) + 0.5*np.cos(6*2*np.pi*time) + 1.5*np.sin(9*2*np.pi*time)
normalized_cutoff_freq = 2 * cutoff_freq / sampling_freq
numerator_coeffs, denominator_coeffs = scipy.signal.butter(order, normalized_cutoff_freq)
filtered_signal = scipy.signal.lfilter(numerator_coeffs, denominator_coeffs, vectors_accelerometer_x)
plt.figure(figsize=(20, 16))
#plt.plot(ox, vectors_accelerometer_x, 'b-', label='signal')
plt.plot(time, filtered_signal, 'g-', linewidth=2, label='filtered signal')
plt.legend()
plt.show()
# +
order = 5
sampling_freq = 30
cutoff_freq = 2
print(int(ox[len(ox) - 1]))
sampling_duration = int(ox[len(ox) - 1])
number_of_samples = len(vectors_accelerometer_x)
time = np.linspace(0, sampling_duration, number_of_samples, endpoint=False)
#signal = np.sin(2*np.pi*time) + 0.5*np.cos(6*2*np.pi*time) + 1.5*np.sin(9*2*np.pi*time)
normalized_cutoff_freq = 2 * cutoff_freq / sampling_freq
numerator_coeffs, denominator_coeffs = scipy.signal.butter(order, normalized_cutoff_freq)
filtered_signal = scipy.signal.lfilter(numerator_coeffs, denominator_coeffs, vectors_accelerometer_x)
plt.figure(figsize=(20, 16))
#plt.plot(ox, vectors_accelerometer_x, 'b-', label='signal')
plt.plot(time, filtered_signal, 'g-', linewidth=2, label='filtered signal')
plt.legend()
plt.show()
| old_version2/filter/.ipynb_checkpoints/filter-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib auto
image = cv2.imread('../data/Lena.png', 0).astype(np.float32) / 255
fft = cv2.dft(image, flags=cv2.DFT_COMPLEX_OUTPUT)
print('FFT shape:', fft.shape)
print('FFT data type:', fft.dtype)
shifted = np.fft.fftshift(fft, axes=[0, 1])
magnitude = cv2.magnitude(shifted[:,:,0], shifted[:,:,1])
print(magnitude.shape)
magnitude = np.log(magnitude)
# magnitude -= magnitude.min()
# magnitude /= magnitude.max()
print(magnitude.dtype)
plt.axis('off')
plt.imshow(magnitude, cmap='gray')
plt.tight_layout(True)
plt.show()
cv2.imshow('magnitude', magnitude)
cv2.waitKey()
cv2.destroyAllWindows()
restored = np.fft.ifft2(fft).astype(np.float32)
cv2.imshow('restored', restored)
cv2.waitKey()
cv2.destroyAllWindows()
| Chapter02/14 Going from spatial to frequency domain (and back) using discrete Fourier transform.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="1yQ-INW0vUE5"
'''Visualize all datasets and methods'''
import sys
import os
from data import get_dataset, dummy_clusters, dummy_half_doughnuts, dummy_linear_points, dummy_polynomial_points, prepare_csv
from data_utils import show_random, AddNoise, remove_random, remove_class, combine_datasets
from matplotlib import pyplot as plt
import numpy as np
# %matplotlib inline
# + [markdown] id="cjSZ08rBuCl9"
# ### Get datasets
# + id="j-x1MONUvUE5" outputId="b60f8267-3a84-42a6-876e-a33277945b11"
train, val, test = get_dataset("mnist")
show_random(test, 5)
# + id="ar1KZ2GGvUE6" outputId="bbea705c-bae4-4349-83d1-2c4d57c5bb58"
# normalised, so not viewed properly
train, val, test = get_dataset("cifar10")
show_random(test, 5)
# + id="Awm15ZOfvUE6" outputId="ebd930b0-7625-4850-bcd4-6aaedc7f54a7"
train, val, test = get_dataset("cifar100")
show_random(test, 5)
# + id="wJ0j4spPvUE6" outputId="4f364c79-82fa-4a2a-ffa3-965c7e466165"
train, val, test = get_dataset("fashion-mnist")
show_random(test, 5)
# + id="VZRmW7nQvUE6" outputId="10b333cc-910f-465f-ed54-613a4f2a175a"
train, val, test = get_dataset("csv")
for data, label in train:
print("data", data[5])
print("label", label[5])
break
# + id="IXO9OZvtvUE7" outputId="cae2eba0-e60f-4068-d391-ad34964f0cd9"
(x1, y1), (x2, y2) = dummy_clusters()
plt.scatter(x1, y1)
plt.scatter(x2, y2)
# + id="2QEPMJk2vUE7" outputId="3108d2bf-fb8f-4d55-ea29-bff7ba13fbce"
(x1, y1), (x2, y2) = dummy_half_doughnuts(500, 500)
plt.scatter(x1, y1)
plt.scatter(x2, y2)
# + id="uO7869tFvUE7" outputId="3fa52b0b-ccea-4133-e61f-508eee23270d"
X, Y = dummy_linear_points((100, 1))
fig = plt.figure(figsize=(8, 6))
ax = fig.add_subplot(111, projection='3d')
xs = X[:, 0]
ys = X[:, 1]
zs = Y
ax.scatter(xs, ys, zs, s=50, alpha=0.6, edgecolors='w')
ax.set_zlabel('Y')
# + id="i2WBOZ3QvUE7" outputId="af68ae12-7e5c-416c-b3bd-69076a2c5493"
X, Y, deg = dummy_polynomial_points((100, 1), 3)
fig = plt.figure(figsize=(8, 6))
plt.scatter(X, Y)
#ax = fig.add_subplot(111, projection='3d')
#xs = X[:]
#ys = Y
#ax.scatter(xs, ys, s=50, alpha=0.6, edgecolors='w')
#ax.set_zlabel('Y')
# + [markdown] id="UO1i3HI-uNHD"
# ### Add noise
# + id="rud8RieBvUE8" outputId="fcbcf091-61ca-47fd-ba98-f3842680dd77"
# noise into data
noise = AddNoise(mean=0, std=0.2)
train, val, test = get_dataset("mnist")
for test_images, test_labels in train:
img = test_images[0]
print('\n')
print('--Without noise--')
img = img.reshape(32, 32, -1)
plt.axis('off')
plt.imshow(img)
plt.show()
#------
img = test_images[0]
img = noise.encodes(img)
print('--Gaussian noise--')
img = img.reshape(32, 32, -1)
plt.axis('off')
plt.imshow(img)
plt.show()
#------
img = test_images[0]
noise.noise_type = 'speckle'
noise.mean=0
noise.std=0.003
img = noise.encodes(img)
print('--Speckle noise--')
img = img.reshape(32, 32, -1)
plt.axis('off')
plt.imshow(img)
plt.show()
#-----
img = test_images[0]
noise.noise_type = 's&p'
noise.s_vs_p=0.5
noise.amount=0.5
img = noise.encodes(img)
print('--Salt and Pepper noise--')
img = img.reshape(32, 32, -1)
plt.axis('off')
plt.imshow(img)
plt.show()
#-----
img = test_images[0]
noise.noise_type = 'poisson'
img = noise.encodes(img)
print('--Poisson noise--')
img = img.reshape(32, 32, -1)
plt.axis('off')
plt.imshow(img)
plt.show()
break
# + id="H1M94FWjvUE8" outputId="8a5f0d65-dbbf-4b81-8d03-c45f1f921e8a"
noise = AddNoise(mean=0, std=0.2)
noisy_img = noise.generate()
noisy_img = noisy_img.view(32, 32, -1)
plt.axis('off')
plt.imshow(noisy_img)
plt.show()
# + [markdown] id="e3MsOIequS50"
# ### Separate and combine data
# + id="VBBollGzvUE8" outputId="09201de7-01e0-4eb2-c0b9-bbc91398b0e2"
train, val, test = get_dataset("mnist")
ds = remove_random(train, 5)
print("Initial length: ", len(train))
print("After separating:", len(ds))
show_random(test, 5)
# + id="G6AdfcSBvUE8" outputId="3a655f19-d6f0-4693-fa3c-40f5e2e0c867"
forget, retain = remove_class(train, [1])
for img, lab in forget:
print("lab: ", lab)
print("--FORGET--")
show_random(forget, 5)
print("--RETAIN--")
show_random(retain, 5)
# + id="N7MOX7X1vUE9" outputId="5d2b5eac-3c05-4b9f-a8dc-af9291241056"
noise = AddNoise(mean=0, std=0.2)
train, val, test = get_dataset("mnist")
nval = noise.encode_data(val)
ntest = noise.encode_data(test)
show_random(nval, 5)
# + id="TqzdJaUPttPd" outputId="3624823e-c0ae-499a-b426-a8a806ff862d"
combine = combine_datasets(ntest, nval)
print("ntest: ", len(ntest))
print("nval: ", len(nval))
print("combine: ", len(combine))
| data/data_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Matplotlib Learning
# ## I. Brief
# Here provides the structure of matplotlib
# 
# <br>
# To start with, there are three top APIs contained in matplotlib:
# 
# ## II. pyplot
# 1. process of pyplot
# 
import matplotlib.pyplot as plt
from matplotlib.axes import Axes, Subplot
# 2. Create draw_board:<br>
# 1). plt.figure,主要接收一个元组作为figsize参数设置图形大小,返回一个figure对象用于提供画板;<br>
# 2). plt.axes,接收一个figure或在当前画板上添加一个子图,返回该axes对象,并将其设置为"当前"图,缺省时会在绘图前自动添加;<br>
fig, axes = plt.subplots(2, 2)
# 3). plt.subplot,主要接收3个数字或1个3位数(自动解析成3个数字,要求解析后数值合理)作为子图的行数、列数和当前子图索引,索引从1开始(与MATLAB保存一致),返回一个axes对象用于绘图操作。这里,可以理解成是先隐式执行了plt.figure,然后在创建的figure对象上添加子图,并返回当前子图实例;<br>plt.subplots,主要接收一个行数nrows和列数ncols作为参数(不含第三个数字),创建一个figure对象和相应数量的axes对象,同时返回该figure对象和axes对象嵌套列表,并默认选择最后一个子图作为"当前"图<br>
ax = plt.subplot(221)
ax.plot([2, 1])
ax1 = plt.subplot(224)
plt.plot([1, 2])
# ## III. pylab
# +
import matplotlib.pylab as plb
# 1. create matrics
plb.array([1, 2], dtype=int)
plb.random
plb.plot(range(10))
# -
| .ipynb_checkpoints/matplotlib_brief-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#iris dataset
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.datasets import load_iris
data = load_iris()
data
# +
df = pd.DataFrame(data.data)
df = df.rename(columns={0:'Sepal_length',1:'Sepal_width',2:'Petal_length',3:'Petal_width'})
df.head()
# -
target = pd.Series(data.target)
df['target'] = target
df
df.info()
df['target'].value_counts()
df['target'] = df['target'].astype(str)
df.dtypes
df['target'] = df['target'].replace({'0':'setosa','1':'versicolor','2':'virginicai'}) #['setosa', 'versicolor', 'virginica'])
df
X = df.iloc[:,:-1]
y = df['target']
X,y
# +
fig,(ax,ax1,ax2,ax3) = plt.subplots(4,constrained_layout=True,figsize=(10,10))
_=ax.scatter(X.Sepal_length,y,label='Sepal_length')
_=ax1.scatter(X.Sepal_width,y,label='Sepal_width')
_=ax2.scatter(X.Petal_length,y,label='Petal_length')
_=ax3.scatter(X.Petal_width,y,label='Petal_width')
_=ax.legend()
_=ax1.legend()
_=ax2.legend()
_=ax3.legend()
# -
from sklearn.preprocessing import StandardScaler
SC = StandardScaler()
X_sc = SC.fit_transform(X)
X = pd.DataFrame(X_sc,columns={'Sepal_length','Sepal_width','Petal_length','Petal_width'})
# +
fig,ax = plt.subplots()
_=ax.scatter(X.Sepal_length,y,label='Sepal_length')
_=ax.scatter(X.Sepal_width,y,label='Sepal_width')
_=ax.scatter(X.Petal_length,y,label='Petal_length')
_=ax.scatter(X.Petal_width,y,label='Petal_width')
_=ax.legend()
# -
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size=0.3)
# +
from sklearn.linear_model import LogisticRegression
model_LR = LogisticRegression()
model_LR.fit(X_train,y_train)
model_LR.score(X_train,y_train)
# -
Yp = model_LR.predict(X_test)
Yp
# +
Yp = pd.Series(Yp)
y_test = y_test.replace({'setosa':'0','versicolor':'1','virginicai':'2'})
Yp = Yp.replace({'setosa':'0','versicolor':'1','virginicai':'2'})
from sklearn.metrics import r2_score
r2s = r2_score(y_test,Yp)
r2s
# -
df.head()
# +
#Test= {'Sepal_length':[5.1,4.9],"Petal_width":[3.5,3.0],'Sepal_width':[1.4,1.4],'Petal_length': [0.2,0.2]}
#Test ={'Sepal_length':[2.2,2.4],"Petal_width":[2.1,1.8],'Sepal_width':[0.2,0.11],'Petal_length': [0.2,0.1]}
#Test = pd.DataFrame(Test)
#SS = StandardScaler()
#Test=SS.fit_transform(Test)
model_LR.predict(df.iloc[0:5,:-1])
#df.iloc[0:1,:]
# -
| Iris Dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Getting Started with Distributed Data Parallel
# 분산 데이터 병렬 시작
# DistributedDataParallel (DDP) implements data parallelism at the module level which can run across multiple machines. Applications using DDP should spawn multiple processes and create a single DDP instance per process. DDP uses collective communications in the torch.distributed package to synchronize gradients and buffers. More specifically, DDP registers an autograd hook for each parameter given by model.parameters() and the hook will fire when the corresponding gradient is computed in the backward pass. Then DDP uses that signal to trigger gradient synchronization across processes. Please refer to DDP design note for more details.
#
# The recommended way to use DDP is to spawn one process for each model replica, where a model replica can span multiple devices. DDP processes can be placed on the same machine or across machines, but GPU devices cannot be shared across processes. This tutorial starts from a basic DDP use case and then demonstrates more advanced use cases including checkpointing models and combining DDP with model parallel.
#
# DDP (DistributedDataParallel)는 여러 시스템에서 실행 될 수있는 모듈 수준에서 데이터 병렬 처리를 구현합니다. DDP를 사용하는 응용 프로그램은 여러 프로세스를 생성하고 프로세스 당 단일 DDP 인스턴스를 만들어야합니다. DDP는 torch.distributed 패키지의 집단 통신을 사용하여 그라디언트 및 버퍼를 동기화합니다. 보다 구체적으로, DDP는 model.parameters()에 의해 주어진 각 매개 변수에 대해 autograd 훅을 등록하고 해당 그라디언트가 역방향 패스에서 계산 될 때 훅이 발생합니다. 그런 다음 DDP는 이 신호를 사용하여 프로세스 간 그래디언트 동기화를 트리거합니다. 자세한 내용은 DDP 디자인 노트를 참조하십시오.
#
# DDP를 사용하는 데 권장되는 방법은 모델 복제본이 여러 장치에 걸쳐있을 수 있는 각 모델 복제본에 대해 하나의 프로세스를 생성하는 것 입니다. DDP 프로세스는 동일한 컴퓨터 또는 여러 컴퓨터에 배치 할 수 있지만 GPU 장치는 프로세스 간에 공유 할 수 없습니다. 이 학습서는 기본 DDP 유스 케이스에서 시작하여 체크 포인트 모델 및 DDP와 모델 병렬 결합을 포함하여보다 고급 유스 케이스를 보여줍니다.
#
# Comparison between DataParallel and DistributedDataParallel
#
# Before we dive in, let’s clarify why, despite the added complexity, you would consider using DistributedDataParallel over DataParallel:
#
# First, DataParallel is single-process, multi-thread, and only works on a single machine, while DistributedDataParallel is multi-process and works for both single- and multi- machine training. DataParallel is usually slower than DistributedDataParallel even on a single machine due to GIL contention across threads, per-iteration replicated model, and additional overhead introduced by scattering inputs and gathering outputs.
#
# Recall from the prior tutorial that if your model is too large to fit on a single GPU, you must use model parallel to split it across multiple GPUs. DistributedDataParallel works with model parallel; DataParallel does not at this time. When DDP is combined with model parallel, each DDP process would use model parallel, and all processes collectively would use data parallel.
#
# If your model needs to span multiple machines or if your use case does not fit into data parallelism paradigm, please see the RPC API for more generic distributed training support.
#
# DataParallel과 DistributedDataParallel의 비교
#
# 자세히 알아보기 전에 복잡성이 추가 되었음에도 불구하고 DataParallel 대신 DistributedDataParallel을 사용하는 이유를 명확히 하겠습니다.
#
# 첫째, DataParallel은 단일 프로세스, 다중 스레드이며 단일 시스템에서만 작동하는 반면, DistributedDataParallel은 다중 프로세스이며 단일 및 다중 시스템 훈련에 모두 작동합니다. 스레드 간에 GIL 경합, 반복 복제 모델 및 입력 분산 및 출력 수집으로 인한 추가 오버 헤드로 인해 단일 시스템에서도 DataParallel이 DistributedDataParallel보다 일반적으로 느립니다.
#
# 이전 자습서에서 모델이 너무 커서 단일 GPU에 맞지 않으면 모델을 병렬로 사용하여 여러 GPU로 분할해야 합니다. DistributedDataParallel은 모델 병렬로 작동합니다. 현재 DataParallel은 없습니다. DDP가 모델 병렬과 결합되면 각 DDP 프로세스는 모델 병렬을 사용하고 모든 프로세스는 집합 적으로 데이터 병렬을 사용합니다.
#
# 모델이 여러 시스템에 걸쳐 있거나 사용 사례가 데이터 병렬 패러다임에 맞지 않는 경우 보다 일반적인 분산 교육 지원에 대해서는 RPC API를 참조하십시오.
# Basic Use Case
#
# To create DDP modules, first set up process groups properly. More details can be found in Writing Distributed Applications with PyTorch.
#
# DDP 모듈을 작성하려면 먼저 프로세스 그룹을 올바르게 설정하십시오. 자세한 내용은 PyTorch로 분산 응용 프로그램 작성에서 찾을 수 있습니다.
# +
import os
import tempfile
import torch
import torch.distributed as dist
import torch.nn as nn
import torch.optim as optim
import torch.multiprocessing as mp
from torch.nn.parallel import DistributedDataParallel as DDP
def setup(rank, world_size):
os.environ['MASTER_ADDR'] = 'localhost'
os.environ['MASTER_PORT'] = '12355'
# initialize the process group
dist.init_process_group('gloo', rank=rank, world_size=world_size)
def cleanup():
dist.destroy_process_group()
# -
# Now, let’s create a toy module, wrap it with DDP, and feed it with some dummy input data. Please note, as DDP broadcasts model states from rank 0 process to all other processes in the DDP constructor, you don’t need to worry about different DDP processes start from different model parameter initial values.
#
# 이제 toy 모듈을 만들어 DDP로 감싸서 더미 입력 데이터를 공급해 보겠습니다. DDP는 순위 0 프로세스에서 DDP 생성자에 있는 다른 모든 프로세스로 모델 상태를 브로드 캐스트하므로 다른 모델 매개 변수 초기 값에서 시작하는 다른 DDP 프로세스에 대해 걱정할 필요가 없습니다.
# +
class ToyModel(nn.Module):
def __init__(self):
super(ToyModel, self).__init__()
self.net1 = nn.Linear(10, 10)
self.relu = nn.ReLU()
self.net2 = nn.Linear(10, 5)
def forward(self, x):
return self.net2(self.relu(self.net1(x)))
def demo_basic(rank, world_size):
print(f'Running basic DDP example on rank {rank}.')
setup(rank, world_size)
# create model and move it to GPU with id rank
model = ToyModel().to(rank)
ddp_model = DDP(model, device_ids=[rank])
loss_fn = nn.MSELoss()
optimizer = optim.SGD(ddp_model.parameters(), lr=0.001)
optimizer.zero_grad()
outputs = ddp_model(torch.randn(20, 10))
labels = torch.randn(20, 5).to(rank)
loss_fn(outputs, labels).backward()
optimizer.step()
clearup()
def run_demo(demo_fn, world_size):
mp.spawn(demo_fn, args=(world_size,), nprocs=world_size, join=True)
# -
# As you can see, DDP wraps lower-level distributed communication details and provides a clean API as if it is a local model. Gradient synchronization communications take place during the backward pass and overlap with the backward computation. When the backward() returns, param.grad already contains the synchronized gradient tensor. For basic use cases, DDP only requires a few more LoCs to set up the process group. When applying DDP to more advanced use cases, some caveats require caution.
#
# 보다시피 DDP는 하위 수준의 분산 통신 세부 정보를 래핑하고 마치 로컬 모델인 것 처럼 깨끗한 API를 제공합니다. 그라디언트 동기화 통신은 역방향 패스 동안 발생하며 역방향 계산과 겹칩니다. backward()이 반환되면 param.grad에 이미 동기화 된 그래디언트 텐서가 포함되어 있습니다. 기본 사용 사례의 경우 DDP는 프로세스 그룹을 설정하기 위해 몇 개의 LoC 만 필요합니다. 고급 사용 사례에 DDP를 적용 할 때 주의해야 할 사항이 있습니다.
# Skewed Processing Speeds
#
# In DDP, the constructor, the forward pass, and the backward pass are distributed synchronization points. Different processes are expected to launch the same number of synchronizations and reach these synchronization points in the same order and enter each synchronization point at roughly the same time. Otherwise, fast processes might arrive early and timeout on waiting for stragglers. Hence, users are responsible for balancing workloads distributions across processes. Sometimes, skewed processing speeds are inevitable due to, e.g., network delays, resource contentions, unpredictable workload spikes. To avoid timeouts in these situations, make sure that you pass a sufficiently large timeout value when calling init_process_group.
#
# DDP 에서 생성자, 전달 패스 및 역방향 패스는 분산 동기화 지점입니다. 서로 다른 프로세스는 동일한 수의 동기화를 시작하고 동일한 순서로 이러한 동기화 지점에 도달하고 대략 동시에 각 동기화 지점에 들어갈 것으로 예상됩니다. 그렇지 않으면 빠른 프로세스가 일찍 도착하여 straggler 대기 시간이 초과 될 수 있습니다. 따라서 사용자는 프로세스 간 워크로드 분산의 균형을 유지 해야 합니다. 때로는 네트워크 지연, 리소스 경합, 예측할 수 없는 워크로드 급증으로 인해 처리 속도가 왜곡 될 수 있습니다. 이러한 상황에서 시간 초과를 피하려면 init_process_group을 호출 할 때 시간 초과 값을 충분히 크게 전달하십시오.
# Save and Load Checkpoints
#
# It’s common to use torch.save and torch.load to checkpoint modules during training and recover from checkpoints. See SAVING AND LOADING MODELS for more details. When using DDP, one optimization is to save the model in only one process and then load it to all processes, reducing write overhead. This is correct because all processes start from the same parameters and gradients are synchronized in backward passes, and hence optimizers should keep setting parameters to the same values. If you use this optimization, make sure all processes do not start loading before the saving is finished. Besides, when loading the module, you need to provide an appropriate map_location argument to prevent a process to step into others’ devices. If map_location is missing, torch.load will first load the module to CPU and then copy each parameter to where it was saved, which would result in all processes on the same machine using the same set of devices. For more advanced failure recovery and elasticity support, please refer to TorchElastic.
#
# 훈련 중에 checkpoints 모듈을 검사하고 checkpoints 에서 복구하는 데 torch.save 및 torch.load를 사용하는 것이 일반적입니다. 자세한 내용은 저장 및 로딩 모델을 참조하십시오. DDP를 사용할 때 하나의 최적화는 모델을 하나의 프로세스에만 저장 한 다음 모든 프로세스에 로드하여 쓰기 오버 헤드를 줄이는 것입니다. 모든 프로세스가 동일한 매개 변수에서 시작하고 그라디언트가 역방향 패스로 동기화되므로 최적화 프로그램이 매개 변수를 동일한 값으로 설정해야합니다. 이 최적화를 사용하는 경우 저장이 완료되기 전에 모든 프로세스가 로드를 시작하지 않는지 확인하십시오. 또한 모듈을 로드 할 때 프로세스가 다른 기기로 들어 가지 않도록 적절한 map_location 인수를 제공 해야 합니다. map_location이 없으면 torch.load는 먼저 모듈을 CPU에 로드 한 다음 각 매개 변수를 저장된 위치에 복사하여 동일한 장치에서 동일한 장치 세트를 사용하는 모든 프로세스를 생성합니다. 고급 오류 복구 및 탄력성 지원에 대해서는 TorchElastic을 참조하십시오.
def demo_checkpoint(rank, world_size):
print(f'Running DDP checkpoint example on rank {rank}')
setup(rank, world_size)
model = ToyModel().to(rank)
ddp_model = DDP(model, device_ids=[rank])
loss_fn = nn.MSELoss()
optimizer = optim.SGD(ddp_model.parameters(), lr=0.001)
CHECKPOINT_PATH = tempfile.gettempdir() + '/model.checkpoint'
if rank == 0:
# 모든 프로세스는 모두 동일한 매개 변수로 시작해야합니다.
# 랜덤 파라미터와 그라디언트는 역방향 패스에서 동기화됩니다.
# 따라서 하나의 프로세스로 저장하면 충분합니다.
torch.save(ddp_model.state_dict(), CHECKPOINT_PATH)
# process 0 이 모델을 저장 한 후 process 1 이 모델을 로드하는지 확인하려면
# barrier()를 사용하십시오.
dist.barrier()
# configure map_location properly
map_location = {'cuda:%d' % 0 : 'cuda:%d' % rank}
ddp_model.load_state_dict(
torch.load(CHECKPOINT_PATH, map_location=map_location))
optimizer.zero_grad()
outputs = ddp_model(torch.randn(20, 10))
labels = torch.randn(20, 5).to(rank)
loss_fn = nn.MSELoss()
loss_fn(outputs, labels).backward()
optimizer.step()
# Use a barrier() to make sure that
# all processes have finished reading the checkpoint.
dist.barrier()
if rank == 0:
os.remove(CHECKPOINT_PATH)
clean_up()
# Combine DDP with Model Parallelism
#
# DDP also works with multi-GPU models. DDP wrapping multi-GPU models is especially helpful when training large models with a huge amount of data.
#
#
# DDP는 다중 GPU 모델에서도 작동합니다. 다중 GPU 모델을 래핑하는 DDP는 많은 양의 데이터로 큰 모델을 훈련 할 때 특히 유용합니다.
class ToyMpModel(nn.Module):
def __init__(self, dev0, dev1):
super(ToyMpModel, self).__init__()
self.dev0 = dev0
self.dev1 = dev1
self.net1 = torch.nn.Linear(10, 10).to(dev0)
self.relu = torch.nn.ReLU()
self.net2 = torch.nn.Linear(10, 5).to(dev1)
def forward(self, x):
x = x.to(self.dev0)
x = self.relu(self.net1(x))
x = x.to(self.dev1)
return self.net2(x)
# When passing a multi-GPU model to DDP, device_ids and output_device must NOT be set. Input and output data will be placed in proper devices by either the application or the model forward() method.
#
# 다중 GPU 모델을 DDP에 전달할 때는 device_ids 및 output_device를 설정하지 않아야 합니다. 입력 및 출력 데이터는 응용 프로그램 또는 모델 forward() 메소드에 의해 적절한 장치에 배치됩니다.
# +
def demo_model_parallel(rank, world_size):
print(f'Running DDP with model parallel example on rank {rank}')
setup(rank, world_size)
# setup mp_model and devices for this process
dev0 = rank * 2
dev1 = rank * 2 + 1
mp_model = ToyMpModel(dev0, dev1)
ddp_mp_model = DDP(mp_model)
loss_fn = nn.MSELoss()
optimizer = optim.SGD(ddp_mp_model.parameters(), lr=0.001)
optimizer.zero_grad()
# outputs will be on dev1
outputs = ddp_mp_model(torch.randn(20, 10))
labels = torch.randn(20, 5).to(dev1)
loss_fn(outputs, labels).backward()
optimizer.step()
cleanup()
if __name__ == '__main__':
n_gpus = torch.cuda.device_count()
run_demo(demo_basic, 2)
| ml/deep_learning/pytorch/Distributed Data Parallel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyCharm (Tutorials)
# language: python
# name: pycharm-38c7cf03
# ---
# +
#####################################################################
# This notebook is authored by: <NAME> & <NAME> #
# Date: May 2022 #
# If you use this code or the results from this work please cite: #
# Machine learning the trilinear and light-quark Yukawa couplings #
# from Higgs pair kinematic shapes #
# <NAME>, <NAME>, <NAME>, <NAME> #
# and <NAME> #
# arXiv:2205.XXXXX (https://arxiv.org/abs/2005.XXXXX) #
#####################################################################
import numpy as np
import pandas as pd
import xgboost as xgb
from sklearn import ensemble
import sklearn.model_selection as ms
from sklearn import metrics
import shap
import matplotlib.pyplot as plt
import os
import math as m
import collections
import pickle
from matplotlib.colors import ListedColormap, LinearSegmentedColormap
from colour import Color
from matplotlib import rc
import sys
import time
from joblib import Parallel, delayed
plt.rcParams['text.latex.preamble'] = r"\usepackage{amsmath}"
plt.rcParams['font.family'] = 'monospace'
# To supress warnings from shap
if not sys.warnoptions:
import warnings
warnings.simplefilter("ignore")
N_THREADS = 25 ## Change for reducing load on CPU
os.environ['OMP_NUM_THREADS'] = str(N_THREADS)
seed = 42
colors = ['#3f7f93','#da3b46']
cmp_2 = LinearSegmentedColormap.from_list('my_list', [Color(c1).rgb for c1 in colors], N=len(colors))
colors = ['#3f7f93','#da3b46','#98b83b']
cmp_3 = LinearSegmentedColormap.from_list('my_list', [Color(c1).rgb for c1 in colors], N=len(colors))
colors = ['#3f7f93','#da3b46','#F6AE2D', '#98b83b']
cmp_4 = LinearSegmentedColormap.from_list('my_list', [Color(c1).rgb for c1 in colors], N=len(colors))
colors = ['#3f7f93','#da3b46','#F6AE2D', '#98b83b', '#825FC3']
cmp_5 = LinearSegmentedColormap.from_list('my_list', [Color(c1).rgb for c1 in colors], N=len(colors))
colors = ['#60a0af', '#af6079', '#a8b96f', '#fcb880', '#FDD7D0', '#a5d6e0']
cmp_6 = LinearSegmentedColormap.from_list('my_list', [Color(c1).rgb for c1 in colors], N=len(colors))
colors = ['#60a0af', '#af6079', '#a8b96f', '#fcb880', '#FDD7D0', '#a5d6e0', '#825FC3']
cmp_7 = LinearSegmentedColormap.from_list('my_list', [Color(c1).rgb for c1 in colors], N=len(colors))
# how far in eta the endcap goes
ENDCAP = 4
# +
def fileparser(path, dlist, sclass, sample, L=2, cut=False):
""" The fileparser to read the events from a csv
argument:
path: the path to the file
dlist: the list of variables to be excluded
sample: the number of events that will be the train sample.
L: Luminosity scaling
returns:
df_train: the training dataframe
df_test: the testing dataframe
weight: the weight (related to crosssection)
"""
df = pd.read_csv(path)
n = len(df)
df['class'] = sclass
weight = int(round(np.abs(df['weight'].sum()) * 3. * 1e6 * L)) ## The abs(mean()) is taken to make the weight of ybyt +ve
if cut:
N_cut = df.drop(df[((np.abs(df['etaa1'])>ENDCAP) | (np.abs(df['etaa2'])>ENDCAP) | (np.abs(df['etab1'])>ENDCAP) | (np.abs(df['etab2'])>ENDCAP))].index).shape[0]/df.shape[0]
df = df.drop(df[((np.abs(df['etaa1'])>ENDCAP) | (np.abs(df['etaa2'])>ENDCAP) | (np.abs(df['etab1'])>ENDCAP) | (np.abs(df['etab2'])>ENDCAP))].index).shape[0]
weight /= N_cut
df_train = df.sample(n=sample, random_state=seed)
df_test = df.drop(df_train.index)
df_train.drop(columns=dlist, inplace=True)
df_test.drop(columns=dlist, inplace=True)
return df_train, df_test, weight
def runBDT(df, filename='', rf=False, depth=10, sample=1, seed=seed):
""" The BDT/RF runner
argument:
df: the dataframe with all the events
filename: the name of the pickle file to store the model in
rf: a bolean to toggle between BDT and Random Forest classifiers
sample: The fraction of variables to sample
seed: the seed for the random number generator
returns:
classifier: the classifier
x_test: the features for the test set
y_test: the labels for the test set
shap_values: the SHAP values
X_shap: the feature set with which the shap values have been computed
"""
mshap = True if depth <= 10 else False
df = df.sample(frac=sample)
nchannels = len(df['class'].unique())
X = df.drop(columns=['class', 'weight'])
y = df['class'].values
# Split for training and testing
x_train, x_test, y_train, y_test = ms.train_test_split(X.values, y, test_size=0.2, random_state=seed)
eval_set = [(x_train, y_train), (x_test, y_test)]
# Fit the decision tree
if rf:
classifier = ensemble.RandomForestClassifier(max_depth=depth, n_estimators=1000, criterion='gini', n_jobs=int(N_THREADS/2), random_state=seed)
classifier = classifier.fit(x_train, y_train)
else:
classifier = xgb.XGBClassifier(max_depth=depth, learning_rate=0.1, objective='multi:softprob', num_class=nchannels,
n_jobs=N_THREADS, subsample=1, colsample_bytree=1, n_estimators=5000, random_state=seed)
classifier = classifier.fit(x_train, y_train, early_stopping_rounds=30, eval_set=eval_set,
eval_metric=["merror", "mlogloss"], verbose=False)
# Predictions
y_pred = classifier.predict(x_test)
print('Accuracy Score: {:4.2f}% '.format(100*metrics.accuracy_score(y_test, y_pred)))
if filename != '': pickle.dump(classifier, open(filename, 'wb'))
# Calculate the SHAP scores
if mshap:
X_shap = pd.DataFrame(x_test, columns=df.drop(columns=['class', 'weight']).columns)
explainer = shap.TreeExplainer(classifier)
shap_values = explainer.shap_values(X_shap)
else:
shap_values = []
X_shap = pd.DataFrame()
return classifier, x_test, y_test, shap_values, X_shap
def eval_training(classifier):
""" Evaluate the training
argument:
classifier: the BDT classifier
"""
results = classifier.evals_result()
epochs = len(results['validation_0']['merror'])
x_axis = range(0, epochs)
# plot log loss
plt.figure(figsize=(12,5))
plt.subplot(1, 2, 1)
plt.plot(x_axis, results['validation_0']['mlogloss'], label='train')
plt.plot(x_axis, results['validation_1']['mlogloss'], label='test')
plt.legend()
plt.ylabel('log loss')
plt.title('Classifier log loss')
plt.grid()
# plot classification error
plt.subplot(1, 2, 2)
plt.plot(x_axis, results['validation_0']['merror'], label='train')
plt.plot(x_axis, results['validation_1']['merror'], label='test')
plt.legend()
plt.ylabel('Classification Error')
plt.title('Classification Error')
plt.grid()
plt.show()
def abs_shap(df_shap, df, shap_plot, names, class_names, cmp):
''' A function to plot the bar plot for the mean abs SHAP values
arguments:
df_shap: the dataframe of the SHAP values
df: the dataframe for the feature values for which the SHAP values have been determined
shap_plot: The name of the output file for the plot
names: The names of the variables
class_names: names of the classes
cmp: the colour map
'''
rc('text', usetex=True)
plt.rcParams['text.latex.preamble'] = r"\usepackage{amsmath}"
plt.figure(figsize=(5,5))
shap.summary_plot(df_shap, df, color=cmp, class_names=class_names, class_inds='original', plot_size=(5,5), show=False, feature_names=names)
ax = plt.gca()
handles, labels = ax.get_legend_handles_labels()
ax.legend(reversed(handles), reversed(labels), loc='lower right', fontsize=15)
plt.xlabel(r'$\overline{|S_v|}$', fontsize=15)
ax = plt.gca()
ax.spines["top"].set_visible(True)
ax.spines["right"].set_visible(True)
ax.spines["left"].set_visible(True)
vals = ax.get_xticks()
ax.tick_params(axis='both', which='major', labelsize=15)
for tick in vals:
ax.axvline(x=tick, linestyle='dashed', alpha=0.7, color='#808080', zorder=0, linewidth=0.5)
plt.annotate('HL-LHC', xy=(0.5, 0.01), xycoords='axes fraction', horizontalalignment='center',
verticalalignment='bottom', fontsize=20, fontweight='900', zorder=100, color='#474747')
plt.tight_layout()
plt.savefig(shap_plot, dpi=300)
rc('text', usetex=False)
def get_mclass(i, df_array, weight_array, ps_exp_class, seed=seed):
""" This function is used to create the confusion matrix
arguments:
i: integer corresponding to the class number
df_array: the array of the dataframes of the different classes
weight_array: the array of the weights for the different classes
ps_exp_class: the collection of the pseudo experiment events
seed: the seed for the random number generator
returns:
nevents: the number of events
"""
nchannels = len(df_array)
mclass = []
for j in range(nchannels):
mclass.append(collections.Counter(classifier.predict(df_array[j].iloc[:,:-2].values))[i]/len(df_array[j])*weight_array[j]/weight_array[i])
nevents = np.round(ps_exp_class[i]/np.sum(mclass)*np.array(mclass)).astype(int)
return nevents
def build_confusion(df, weight, clf, filename, keys):
""" Export the confusion matrix to a json
argument:
df: array of dataframes with the signal and background
weight: array of weights for the signal and background
clf: the classifier
filename: the name of the file to export to
keys: the channel names for which the confusion matrix is created
returns:
df: dataframe with the confusion matrix
"""
ps_exp_class = collections.Counter(clf.predict(
pd.concat([df[i].iloc[:,:-2].sample(n=round(weight[i]), random_state=seed, replace=True) for i in range(len(df))])
.values))
confusion = np.column_stack([get_mclass(i, df, weight, ps_exp_class) for i in reversed(range(len(df)))])
df = pd.DataFrame(confusion.T)
df = df[df.columns[::-1]]
df.columns = keys
df.index = keys
df = df.T
df['total'] = df.sum(axis=1)
df = df.T
df['Z'] = 0
for key in keys:
df.loc[key,'Z'] = df.loc[key, key]/np.sqrt(df.loc[key].sum())
df.T.to_json(filename)
return df
# +
dlist = ['ptb2', 'etaaa', 'dphibb', 'etaa1', 'etab1', 'etaa2', 'drba1', 'dphiba1', 'drbamin', 'nbjet', 'etab2']
# k-factors
k_box = 1.98
k_int = 2.15
k_tri = 2.28
k_bbh_yb2 = 1.5
k_bbh_ybyt = 1.9
k_bbh_yt2 = 2.5
k_bbh_zh = 1.3
k_tth = 1.2
k_bbxaa = 1.5
k_ku = 1.29
k_kd = 1.3
# 14 TeV
path = '../simulations/HL-LHC/'
# The bbxaa background
df_bbxaa, df_bbxaa_test, weight_bbxaa = fileparser(path+"bbxaa.tar.gz", dlist, sclass=0, sample=120000)
weight_bbxaa = weight_bbxaa*k_bbxaa
# The tth+bbh background
fact = 10
df_tth, df_tth_test, weight_tth = fileparser(path+"ttH.tar.gz", dlist, sclass=1, sample=4430*fact)
df_yb2, df_yb2_test, weight_yb2 = fileparser(path+"yb2.tar.gz", dlist, sclass=1, sample=663*fact)
df_ybyt, df_ybyt_test, weight_ybyt = fileparser(path+"ybyt.tar.gz", dlist, sclass=1, sample=139*fact)
df_yt2, df_yt2_test, weight_yt2 = fileparser(path+"yt2.tar.gz", dlist, sclass=1, sample=2888*fact)
df_zh, df_zh_test, weight_zh = fileparser(path+"zh.tar.gz", dlist, sclass=1, sample=797*fact)
df_bbh_tth = pd.concat([df_tth, df_yb2, df_ybyt, df_yt2, df_zh])
df_bbh_tth_test = pd.concat([df_tth_test, df_yb2_test, df_ybyt_test, df_yt2_test, df_zh_test])
weight_bbh_tth = int(weight_tth*k_tth + weight_yb2*k_bbh_yb2 - weight_ybyt*k_bbh_ybyt + weight_yt2*k_bbh_yt2 + weight_zh*k_bbh_zh)
# The hhsm signal
df_hhsm_b, df_hhsm_b_test, weight_hhsm_b = fileparser(path+"HH-box.tar.gz", dlist, sclass=2, sample=20000)
df_hhsm_i, df_hhsm_i_test, weight_hhsm_i = fileparser(path+"HH-int.tar.gz", dlist, sclass=3, sample=20000)
df_hhsm_t, df_hhsm_t_test, weight_hhsm_t = fileparser(path+"HH-tri.tar.gz", dlist, sclass=4, sample=20000)
weight_hhsm_b = weight_hhsm_b*k_box
weight_hhsm_i = weight_hhsm_i*k_int
weight_hhsm_t = weight_hhsm_t*k_tri
fact = 10
df_hhsm = pd.concat([df_hhsm_b.sample(n=int(weight_hhsm_b*fact)), df_hhsm_i.sample(n=int(weight_hhsm_i*fact)), df_hhsm_t.sample(n=int(weight_hhsm_t*fact))])
df_hhsm['class'] = 2
df_hhsm_test = pd.concat([df_hhsm_b_test, df_hhsm_i_test, df_hhsm_t_test])
df_hhsm_test['class'] = 2
weight_hhsm = int(weight_hhsm_b - weight_hhsm_i + weight_hhsm_t)
# kappa_u and kappa_d
df_ku, df_ku_test, weight_ku = fileparser(path+"ku-1600.tar.gz", dlist, sclass=5, sample=20000)
df_kd, df_kd_test, weight_kd = fileparser(path+"kd-800.tar.gz", dlist, sclass=5, sample=20000)
weight_ku = weight_ku*k_ku
weight_kd = weight_kd*k_kd
print("No. of kappa_d triangle events: train = {}, test = {}".format(df_kd.shape[0],df_kd_test.shape[0]))
print("No. of kappa_u triangle events: train = {}, test = {}".format(df_ku.shape[0],df_ku_test.shape[0]))
print("No. of hhsm events: train = {}, test = {}".format(df_hhsm.shape[0],df_hhsm_test.shape[0]))
print("No. of hhsm triangle events: train = {}, test = {}".format(df_hhsm_t.shape[0],df_hhsm_t_test.shape[0]))
print("No. of hhsm interference events: train = {}, test = {}".format(df_hhsm_i.shape[0],df_hhsm_i_test.shape[0]))
print("No. of hhsm box events: train = {}, test = {}".format(df_hhsm_b.shape[0],df_hhsm_b_test.shape[0]))
print("No. of bbh+tth events: train = {}, test = {}".format(df_bbh_tth.shape[0],df_bbh_tth_test.shape[0]))
print("No. of bbxaa events: train = {}, test = {}".format(df_bbxaa.shape[0],df_bbxaa_test.shape[0]))
names = [r'$n_{jet}$', r'$p_T^{b_1}$', r'$p_T^{\gamma_1}$', r'$p_T^{\gamma_2}$', r'$p_T^{\gamma\gamma}$', r'$m_{bb}$', r'$m_{\gamma\gamma}$', r'$m_{b_1h}$',
r'$m_{bbh}$', r'$E^{\rm miss}_T$', r'$H_T$']
# -
# ___________________
# # The hh analysis
channels = [df_hhsm_t, df_hhsm_i, df_hhsm_b, df_bbh_tth, df_bbxaa]
df_train = pd.concat(channels, ignore_index=True)
df_train = df_train.sample(frac=1).reset_index(drop=True)
# + pycharm={"is_executing": false, "name": "#%%\n"}
class_names = [r'$bb\gamma\gamma$', r'$Q\bar{Q}h$', r'$hh^{gg\rm F}_{\rm box}$', r'$hh^{ggF}_{\rm int}$', r'$hh^{gg\rm F}_{\rm tri}$']
filename = '../results/models/HL-LHC-BDT/hh-BDT-5class-hhsm.pickle.dat'
shap_plot = '../plots/HL-LHC-shap-bbxaa-bbh-tth-hhsm.pdf'
classifier, x_test, y_test, shap_values_5, X_shap_5 = runBDT(df_train, filename, depth=10)
abs_shap(shap_values_5, X_shap_5, shap_plot, names=names, class_names=class_names, cmp=cmp_5)
# +
hhsm_t_p = df_hhsm_t_test.sample(n=round(weight_hhsm_t), replace=True, random_state=seed).reset_index(drop=True)
hhsm_i_p = df_hhsm_i_test.sample(n=round(weight_hhsm_i), replace=True, random_state=seed).reset_index(drop=True)
hhsm_i_b = df_hhsm_b_test.sample(n=round(weight_hhsm_b), replace=True, random_state=seed).reset_index(drop=True)
bbh_tth_p = df_bbh_tth_test.sample(n=round(weight_bbh_tth), replace=True, random_state=seed).reset_index(drop=True)
bbxaa_p = df_bbxaa_test.sample(n=round(weight_bbxaa), replace=True, random_state=seed).reset_index(drop=True)
print('Accuracy Score for hhsm triangle: {:4.2f}% '.format(100*metrics.accuracy_score(hhsm_t_p['class'].values, classifier.predict(hhsm_t_p.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for hhsm interference: {:4.2f}% '.format(100*metrics.accuracy_score(hhsm_t_p['class'].values, classifier.predict(hhsm_t_p.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for hhsm box: {:4.2f}% '.format(100*metrics.accuracy_score(hhsm_i_b['class'].values, classifier.predict(hhsm_i_b.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for bbh+tth: {:4.2f}% '.format(100*metrics.accuracy_score(bbh_tth_p['class'].values, classifier.predict(bbh_tth_p.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for bbxaa: {:4.2f}% '.format(100*metrics.accuracy_score(bbxaa_p['class'].values, classifier.predict(bbxaa_p.drop(columns=['class', 'weight']).values))))
# +
df_array = [df_bbxaa_test, df_bbh_tth_test, df_hhsm_b_test, df_hhsm_i_test, df_hhsm_t_test]
weight_array = [weight_bbxaa, weight_bbh_tth, weight_hhsm_b, weight_hhsm_i, weight_hhsm_t]
keys = ['tri', 'int', 'box', 'tth+bbh', 'bbxaa']
filename = '../results/confusion/HL-LHC-BDT/hh-BDT-5class-hhsm.confusion.json'
df = build_confusion(df_array, weight_array, classifier, filename, keys)
df
# -
# ### Checks for the confusion matrix
# +
now = time.time()
def func(i):
df = pd.concat([df_bbxaa_test.sample(n=int(weight_bbxaa), replace=True),
df_bbh_tth_test.sample(n=int(weight_bbh_tth)),
df_hhsm_b_test.sample(n=int(weight_hhsm_b)),
df_hhsm_i_test.sample(n=int(weight_hhsm_i)),
df_hhsm_t_test.sample(n=int(weight_hhsm_t))])
true_class = df['class'].values
predicted_class = classifier.predict(df.iloc[:, :-2].values)
conf = metrics.confusion_matrix(y_pred=predicted_class, y_true=true_class)[::-1].T
return conf[4-i][i]/np.sqrt(np.sum(conf[4-i]))
results = Parallel(n_jobs=N_THREADS, backend="loky")(delayed(func)(1) for _ in range(100))
print(time.time() - now, np.mean(results))
# -
# ________________________
# ## The kappa_u & kappa_lambda
channels = [df_ku, df_hhsm_t, df_hhsm_i, df_hhsm_b, df_bbh_tth, df_bbxaa]
df_train = pd.concat(channels, ignore_index=True)
df_train = df_train.sample(frac=1).reset_index(drop=True)
# + pycharm={"is_executing": false, "name": "#%%\n"}
class_names = [r'$bb\gamma\gamma$', r'$Q\bar{Q}h$', r'$hh^{gg\rm F}_{\rm box}$', r'$hh^{gg\rm F}_{\rm int}$', r'$hh^{gg\rm F}_{\rm tri}$', r'$u \bar u hh$']
filename = '../results/models/HL-LHC-BDT/hh-BDT-6class-ku.pickle.dat'
shap_plot = '../plots/HL-LHC-shap-bbxaa-bbh-tth-hhsm-ku.pdf'
classifier, x_test, y_test, shap_values_6u, X_shap_6u = runBDT(df_train, filename, depth=10)
abs_shap(shap_values_6u, X_shap_6u, shap_plot, names=names, class_names=class_names, cmp=cmp_6)
# +
ku_p = df_ku_test.sample(n=round(weight_ku), replace=True, random_state=seed).reset_index(drop=True)
hhsm_t_p = df_hhsm_t_test.sample(n=round(weight_hhsm_t), replace=True, random_state=seed).reset_index(drop=True)
hhsm_i_p = df_hhsm_i_test.sample(n=round(weight_hhsm_i), replace=True, random_state=seed).reset_index(drop=True)
hhsm_i_b = df_hhsm_b_test.sample(n=round(weight_hhsm_b), replace=True, random_state=seed).reset_index(drop=True)
bbh_tth_p = df_bbh_tth_test.sample(n=round(weight_bbh_tth), replace=True, random_state=seed).reset_index(drop=True)
bbxaa_p = df_bbxaa_test.sample(n=round(weight_bbxaa), replace=True, random_state=seed).reset_index(drop=True)
print('Accuracy Score for ku: {:4.2f}% '.format(100*metrics.accuracy_score(ku_p['class'].values, classifier.predict(ku_p.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for hhsm triangle: {:4.2f}% '.format(100*metrics.accuracy_score(hhsm_t_p['class'].values, classifier.predict(hhsm_t_p.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for hhsm interference: {:4.2f}% '.format(100*metrics.accuracy_score(hhsm_t_p['class'].values, classifier.predict(hhsm_t_p.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for hhsm box: {:4.2f}% '.format(100*metrics.accuracy_score(hhsm_i_b['class'].values, classifier.predict(hhsm_i_b.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for bbh+tth: {:4.2f}% '.format(100*metrics.accuracy_score(bbh_tth_p['class'].values, classifier.predict(bbh_tth_p.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for bbxaa: {:4.2f}% '.format(100*metrics.accuracy_score(bbxaa_p['class'].values, classifier.predict(bbxaa_p.drop(columns=['class', 'weight']).values))))
# +
df_array = [df_bbxaa_test, df_bbh_tth_test, df_hhsm_b_test, df_hhsm_i_test, df_hhsm_t_test, df_ku_test]
weight_array = [weight_bbxaa, weight_bbh_tth, weight_hhsm_b, weight_hhsm_i, weight_hhsm_t, weight_ku]
keys = ['ku', 'tri', 'int', 'box', 'tth+bbh', 'bbxaa']
filename = '../results/confusion/HL-LHC-BDT/hh-BDT-6class-ku.confusion.json'
df = build_confusion(df_array, weight_array, classifier, filename, keys)
df
# -
# _________________________________
# ## The kappa_d & kappa_lambda
channels = [df_kd, df_hhsm_t, df_hhsm_i, df_hhsm_b, df_bbh_tth, df_bbxaa]
df_train = pd.concat(channels, ignore_index=True)
df_train = df_train.sample(frac=1).reset_index(drop=True)
# + pycharm={"is_executing": false, "name": "#%%\n"}
class_names = [r'$bb\gamma\gamma$', r'$Q\bar{Q}h$', r'$hh^{gg\rm F}_{\rm box}$', r'$hh^{gg\rm F}_{\rm int}$', r'$hh^{gg\rm F}_{\rm tri}$', r'$d \bar d hh$']
filename = '../results/models/HL-LHC-BDT/hh-BDT-6class-kd.pickle.dat'
shap_plot = '../plots/HL-LHC-shap-bbxaa-bbh-tth-hhsm-kd.pdf'
classifier, x_test, y_test, shap_values_6d, X_shap_6d = runBDT(df_train, filename, depth=10)
abs_shap(shap_values_6d, X_shap_6d, shap_plot, names=names, class_names=class_names, cmp=cmp_6)
# +
kd_p = df_kd_test.sample(n=round(weight_kd), replace=True, random_state=seed).reset_index(drop=True)
hhsm_t_p = df_hhsm_t_test.sample(n=round(weight_hhsm_t), replace=True, random_state=seed).reset_index(drop=True)
hhsm_i_p = df_hhsm_i_test.sample(n=round(weight_hhsm_i), replace=True, random_state=seed).reset_index(drop=True)
hhsm_i_b = df_hhsm_b_test.sample(n=round(weight_hhsm_b), replace=True, random_state=seed).reset_index(drop=True)
bbh_tth_p = df_bbh_tth_test.sample(n=round(weight_bbh_tth), replace=True, random_state=seed).reset_index(drop=True)
bbxaa_p = df_bbxaa_test.sample(n=round(weight_bbxaa), replace=True, random_state=seed).reset_index(drop=True)
print('Accuracy Score for ku: {:4.2f}% '.format(100*metrics.accuracy_score(kd_p['class'].values, classifier.predict(kd_p.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for hhsm triangle: {:4.2f}% '.format(100*metrics.accuracy_score(hhsm_t_p['class'].values, classifier.predict(hhsm_t_p.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for hhsm interference: {:4.2f}% '.format(100*metrics.accuracy_score(hhsm_t_p['class'].values, classifier.predict(hhsm_t_p.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for hhsm box: {:4.2f}% '.format(100*metrics.accuracy_score(hhsm_i_b['class'].values, classifier.predict(hhsm_i_b.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for bbh+tth: {:4.2f}% '.format(100*metrics.accuracy_score(bbh_tth_p['class'].values, classifier.predict(bbh_tth_p.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for bbxaa: {:4.2f}% '.format(100*metrics.accuracy_score(bbxaa_p['class'].values, classifier.predict(bbxaa_p.drop(columns=['class', 'weight']).values))))
# +
df_array = [df_bbxaa_test, df_bbh_tth_test, df_hhsm_b_test, df_hhsm_i_test, df_hhsm_t_test, df_kd_test]
weight_array = [weight_bbxaa, weight_bbh_tth, weight_hhsm_b, weight_hhsm_i, weight_hhsm_t, weight_kd]
keys = ['kd', 'tri', 'int', 'box', 'tth+bbh', 'bbxaa']
filename = '../results/confusion/HL-LHC-BDT/hh-BDT-6class-kd.confusion.json'
df = build_confusion(df_array, weight_array, classifier, filename, keys)
df
# -
# ________________________
# ## The kappa_u, kappa_d and kappa_lambda
df_ku['class'] = 6
df_ku_test['class'] = 6
channels = [df_ku, df_kd, df_hhsm_t, df_hhsm_i, df_hhsm_b, df_bbh_tth, df_bbxaa]
df_train = pd.concat(channels, ignore_index=True)
df_train = df_train.sample(frac=1).reset_index(drop=True)
# + pycharm={"is_executing": false, "name": "#%%\n"}
class_names = [r'$bb\gamma\gamma$', r'$Q\bar{Q}h$', r'$hh^{gg\rm F}_{\rm box}$', r'$hh^{gg\rm F}_{\rm int}$', r'$hh^{gg\rm F}_{\rm tri}$', r'$d \bar d hh$', r'$u \bar u hh$']
filename = '../results/models/HL-LHC-BDT/hh-BDT-7class-ku-kd.pickle.dat'
shap_plot = '../plots/HL-LHC-shap-bbxaa-bbh-tth-hhsm-ku-kd-kl.pdf'
classifier, x_test, y_test, shap_values_7ud, X_shap_7ud = runBDT(df_train, filename, depth=10)
abs_shap(shap_values_7ud, X_shap_7ud, shap_plot, names=names, class_names=class_names, cmp=cmp_7)
# +
ku_p = df_ku_test.sample(n=round(weight_ku), replace=True, random_state=seed).reset_index(drop=True)
kd_p = df_kd_test.sample(n=round(weight_kd), replace=True, random_state=seed).reset_index(drop=True)
hhsm_t_p = df_hhsm_t_test.sample(n=round(weight_hhsm_t), replace=True, random_state=seed).reset_index(drop=True)
hhsm_i_p = df_hhsm_i_test.sample(n=round(weight_hhsm_i), replace=True, random_state=seed).reset_index(drop=True)
hhsm_i_b = df_hhsm_b_test.sample(n=round(weight_hhsm_b), replace=True, random_state=seed).reset_index(drop=True)
bbh_tth_p = df_bbh_tth_test.sample(n=round(weight_bbh_tth), replace=True, random_state=seed).reset_index(drop=True)
bbxaa_p = df_bbxaa_test.sample(n=round(weight_bbxaa), replace=True, random_state=seed).reset_index(drop=True)
print('Accuracy Score for ku: {:4.2f}% '.format(100*metrics.accuracy_score(ku_p['class'].values, classifier.predict(ku_p.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for kd: {:4.2f}% '.format(100*metrics.accuracy_score(kd_p['class'].values, classifier.predict(kd_p.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for hhsm triangle: {:4.2f}% '.format(100*metrics.accuracy_score(hhsm_t_p['class'].values, classifier.predict(hhsm_t_p.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for hhsm interference: {:4.2f}% '.format(100*metrics.accuracy_score(hhsm_t_p['class'].values, classifier.predict(hhsm_t_p.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for hhsm box: {:4.2f}% '.format(100*metrics.accuracy_score(hhsm_i_b['class'].values, classifier.predict(hhsm_i_b.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for bbh+tth: {:4.2f}% '.format(100*metrics.accuracy_score(bbh_tth_p['class'].values, classifier.predict(bbh_tth_p.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for bbxaa: {:4.2f}% '.format(100*metrics.accuracy_score(bbxaa_p['class'].values, classifier.predict(bbxaa_p.drop(columns=['class', 'weight']).values))))
# +
df_array = [df_bbxaa_test, df_bbh_tth_test, df_hhsm_b_test, df_hhsm_i_test, df_hhsm_t_test, df_kd_test, df_ku_test]
weight_array = [weight_bbxaa, weight_bbh_tth, weight_hhsm_b, weight_hhsm_i, weight_hhsm_t, weight_kd, weight_ku]
keys = ['ku', 'kd', 'tri', 'int', 'box', 'tth+bbh', 'bbxaa']
filename = '../results/confusion/HL-LHC-BDT/hh-BDT-7class-ku-kd.confusion.json'
df = build_confusion(df_array, weight_array, classifier, filename, keys)
df
# -
# _____________________________
# ## Standard Model di-Higgs
channels = [df_hhsm, df_bbh_tth, df_bbxaa]
df_train = pd.concat(channels, ignore_index=True)
df_train = df_train.sample(frac=1).reset_index(drop=True)
# + pycharm={"is_executing": false, "name": "#%%\n"}
class_names = [r'$bb\gamma\gamma$', r'$Q\bar{Q}h$', r'$hh^{gg\rm F}$']
filename = '../results/models/HL-LHC-BDT/hh-BDT-3class-hhsm-SM.pickle.dat'
shap_plot = '../plots/HL-LHC-shap-bbxaa-bbh-tth-hhsm-SM.pdf'
classifier, x_test, y_test, shap_values_3, X_shap_3 = runBDT(df_train, filename, depth=10)
abs_shap(shap_values_3, X_shap_3, shap_plot, names=names, class_names=class_names, cmp=cmp_5)
# +
hhsm_p = df_hhsm_test.sample(n=round(weight_hhsm), replace=True, random_state=seed).reset_index(drop=True)
bbh_tth_p = df_bbh_tth_test.sample(n=round(weight_bbh_tth), replace=True, random_state=seed).reset_index(drop=True)
bbxaa_p = df_bbxaa_test.sample(n=round(weight_bbxaa), replace=True, random_state=seed).reset_index(drop=True)
print('Accuracy Score for hhsm: {:4.2f}% '.format(100*metrics.accuracy_score(hhsm_p['class'].values, classifier.predict(hhsm_p.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for bbh+tth: {:4.2f}% '.format(100*metrics.accuracy_score(bbh_tth_p['class'].values, classifier.predict(bbh_tth_p.drop(columns=['class', 'weight']).values))))
print('Accuracy Score for bbxaa: {:4.2f}% '.format(100*metrics.accuracy_score(bbxaa_p['class'].values, classifier.predict(bbxaa_p.drop(columns=['class', 'weight']).values))))
# +
df_array = [df_bbxaa_test, df_bbh_tth_test, df_hhsm_test]
weight_array = [weight_bbxaa, weight_bbh_tth, weight_hhsm]
keys = ['hhsm', 'tth+bbh', 'bbxaa']
filename = '../results/confusion/HL-LHC-BDT/hh-BDT-3class-hhsm-SM.confusion.json'
df = build_confusion(df_array, weight_array, classifier, filename, keys)
df
# -
# ### Checks for the confusion matrix
# +
now = time.time()
def func(i):
df = pd.concat([df_bbxaa_test.sample(n=int(weight_bbxaa), replace=True), df_bbh_tth_test.sample(n=int(weight_bbh_tth)), df_hhsm.sample(n=int(weight_hhsm))])
true_class = df['class'].values
predicted_class = classifier.predict(df.iloc[:, :-2].values)
conf = metrics.confusion_matrix(y_pred=predicted_class, y_true=true_class)[::-1].T
return conf[2][0]/np.sqrt(np.sum(conf[2]))
results = Parallel(n_jobs=N_THREADS, backend="loky")(delayed(func)(1) for _ in range(500))
print(time.time() - now, np.mean(results))
| machine_learning/Mega-BDT-kappa-lambda-HL-LHC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib notebook
# %matplotlib notebook
import numpy as np
from rodigan import *
# -
# ### Static cantilever
# +
# define geometric properties of the rod
geometry = Geometry(length=0.1, radius=5e-4)
# define material properties of the rod
material = Material(geometry=geometry, elastic_modulus=200e9, shear_modulus=85e9)
# initialize the solver for the cantilever problem (fixed displacement and rotation at leftmost end)
cantilever = Cantilever(geometry, material, number_of_elements=100)
# set the Neumann boundary condition at the rightmost end
cantilever.boundary_condition = [0, 1, 0, 0, 0, 0]
# -
# %time cantilever.run_simulation()
cantilever.result.plot_centerline()
cantilever.result.plot_load_step_iterations()
cantilever.result.plot_norms_in_loadstep(load_step=-1)
# ## Sandbox ...
import numba
import matplotlib.pyplot as plt
cantilever.result.centerline[:, -1]
numba.typeof(cantilever.result.centerline)
numba.typeof(cantilever.result.increments_norm_evolution)
numba.typeof(cantilever.result.load_step_iterations)
numba.typeof(cantilever.result.load_steps)
numba.typeof(cantilever.result.residuals_norm_evolution)
numba.typeof(cantilever.load_control_parameters[0])
numba.typeof(cantilever.maximum_iterations_per_loadstep)
numba.typeof(cantilever.boundary_condition)
numba.typeof(cantilever.geometry.length)
| cantilever_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.5.0
# language: julia
# name: julia-0.5
# ---
using JLD
include("Transformation.jl")
include("AbstractSystem.jl")
include("Tree.jl")
include("Evaluation.jl")
include("BackPropogation.jl")
include("Facility.jl")
import Base.push!
# +
type Axiom
tree1 :: Tree
tree2 :: Tree
index :: Dict
end
function Base.show(io :: IO, m :: Axiom)
print(io, "Axiom[")
print(io, m.tree1)
print(io, ", ")
print(io, m.tree2)
print(io, "]")
end
function beautify(m :: Axiom)
string("Axiom:\n", beautify(m.tree1, 1), "\n", beautify(m.tree2, 1), "\n")
end
Axiom(tree1 :: Tree, tree2 :: Tree) = Axiom(tree1, tree2, add!(index(tree1), index(tree2)))
Axiom(skeleton1, skeleton2) = Axiom(toTree(skeleton1), toTree(skeleton2))
function push!(index :: Dict, ops :: Array)
n = length(ops)
for i in 1:n
for t in index[i]
t.op = ops[i]
end
end
end
function push!(index :: Dict, ops :: Dict)
for key in keys(ops)
if haskey(index, key)
ts = index[key]
for t in ts
t.op = ops[key]
end
end
end
end
function push!(axiom :: Axiom, ops)
push!(axiom.index, variables)
end
function init_axiom!(axiom :: Axiom, variables)
push!(axiom.index, variables)
init_tree!(axiom.tree1)
init_tree!(axiom.tree2)
end
init_axioms! = distribute(init_axiom!)
## add some loss function to deal with degenerating problem?
function loss(a, b)
b * (1. - a * b)
end
function train_axiom!(axiom :: Axiom, variables, n = 1, randomize = identity)
push!(axiom.index, variables)
d1 :: Array{Float64, 1} = axiom.tree1.value[:d]
d2 :: Array{Float64, 1} = axiom.tree2.value[:d]
v1 :: Array{Float64, 1} = axiom.tree1.value[:value]
v2 :: Array{Float64, 1} = axiom.tree2.value[:value]
for i in 1:n
randomize(variables)
push!(axiom.index, variables)
eval_tree!(axiom.tree2)
eval_tree!(axiom.tree1)
for j in 1:length(d1)
d1[j] = loss(v1[j], v2[j])
d2[j] = loss(v2[j], v1[j])
end
bp_tree!(axiom.tree1)
bp_tree!(axiom.tree2)
end
end
train_axioms! = distribute(train_axiom!)
## to prevent degeneration problem, we use anti-traing to deal with the problem.
function anti_train_axiom!(axiom :: Axiom, n = 1, randomize = identity)
d1 :: Array{Float64, 1} = axiom.tree1.value[:d]
d2 :: Array{Float64, 1} = axiom.tree2.value[:d]
v1 :: Array{Float64, 1} = axiom.tree1.value[:value]
v2 :: Array{Float64, 1} = axiom.tree2.value[:value]
for i in 1:n
randomize(axiom.tree1)
randomize(axiom.tree2)
eval_tree!(axiom.tree2)
eval_tree!(axiom.tree1)
for j in 1:length(d1)
d1[j] = - loss(v1[j], v2[j])
d2[j] = - loss(v2[j], v1[j])
end
bp_tree!(axiom.tree1)
bp_tree!(axiom.tree2)
end
end
anti_train_axioms! = distribute(anti_train_axiom!)
# +
include("DataStructs.jl")
UChar = Class("Char", 8)
Variable = Class("Variable", 100)
Stream = List(UChar, Variable, [:v, :u, :v1])
Prog = Class("Prog", 100)
Bindingc = Class("Binding", 100)
Binding = Pair(Variable, Prog, Bindingc, [:b, :v, :p])
Framec = Class("Frame", 100)
Frame = List(Bindingc, Framec, [:f1, :b, :f2])
Envc = Class("Env", 200)
Env = List(Framec, Envc, [:e1, :f, :e2])
# +
lookup = DFunction("lookup", [Envc, Variable], Prog)
set = DFunction("set", [Envc, Variable, Prog], Envc)
add_binding = DFunction("add_binding", [Framec, Variable, Prog], Framec)
def = DFunction("def", [Envc, Variable, Prog], Envc)
extend = DFunction("extend", [Envc, Variable, Prog], Envc)
var = DFunction("var", [Variable], Prog)
definition = DFunction("definition", [Variable, Prog], Prog)
assignment = DFunction("assignment", [Variable, Prog], Prog)
procedure = DFunction("procedure", [Variable, Prog], Prog)
func_call = DFunction("func_call", [Prog, Prog], Prog)
Seq = List(Prog, Prog)
f_eval = DFunction("f_eval", [Prog, Envc], Prog)
s_eval = DFunction("eval", [Prog, Envc], Envc)
# -
@load "test.jld"
# +
axiom_lookup1 = Axiom([lookup, [Env.cons, [Frame.cons, [Binding.pair, :v, :p], :f], :e], :v], :p)
axiom_lookup2 = Axiom([lookup, [Env.cons, [Frame.cons, [Binding.pair, :v1, :p1], :f2], :e2], :v2],
[lookup, [Env.cons, :f2, :e2], :v2])
axiom_lookup3 = Axiom([lookup, [Env.cons, Frame.empty, :e], :v], [lookup, :e, :v])
axiom_set1 = Axiom([set, [Env.cons, [Frame.cons, [Binding.pair, :v1, :p1], :f2], :e2], :v1, :p2],
[Env.cons, [Frame.cons, [Binding.pair, :v1, :p2], :f2], :e2])
axiom_set2 = Axiom([set, [Env.cons, [Frame.cons, [Binding.pair, :v1, :p1], :f2], :e2], :v2, :p2],
[Env.cons, [Frame.cons, [Binding.pair, :v1, :p1],
[Env.first, [set, [Env.cons, :f2, :e2], :v2, :p2]]],
[Env.rest, [set, [Env.cons, :f2, :e2], :v2, :p2]]])
axiom_set3 = Axiom([set, [Env.cons, Frame.empty, :e], :v, :p], [Env.cons, Frame.empty, [set, :e, :v, :p]])
axiom_add_binding = Axiom([add_binding, :f, :v, :p], [Frame.cons, [Binding.pair, :v, :p], :f])
axiom_def = Axiom([def, :e, :v, :p], [Env.first!, :e, [add_binding, [Env.first, :e], :v, :p]])
axiom_extend = Axiom([extend, :e, :v, :p], [def, [Env.cons, Frame.empty, :e], :v, :p])
# -
axiom_var_s = Axiom([s_eval, [var, :v], :e], :e)
axiom_var_f = Axiom([f_eval, [var, :v], :e], [lookup, :e, :v])
axiom_definition_s = Axiom([s_eval, [definition, :v, :p], :e], [def, [s_eval, :p, :e], :v, [f_eval, :p, :e]])
axiom_definition_f = Axiom([f_eval, [definition, :v, :p], :e], [f_eval, :p, :e])
axiom_assignment_s = Axiom([s_eval, [assignment, :v, :p], :e], [set, [s_eval, :p, :e], :v, [f_eval, :p, :e]])
axiom_assignment_f = Axiom([f_eval, [assignment, :v, :p], :e], [f_eval, :p, :e])
axiom_proc_s = Axiom([s_eval, [procedure, :v, :p], :e], :e)
axiom_proc_f = Axiom([f_eval, [procedure, :v, :p], :e], [procedure, :v, :p])
axiom_func_s1 = Axiom([s_eval, [func_call, :p1, :p2], :e],
[s_eval, [func_call, [f_eval, :p1, :e], [f_eval, :p2, :e]], :e])
axiom_func_f1 = Axiom([f_eval, [func_call, :p1, :p2], :e],
[f_eval, [func_call, [f_eval, :p1, :e], [f_eval, :p2, :e]], :e])
axiom_func_s2 = Axiom([s_eval, [func_call, [procedure, :v, :p1], :p2], :e],
[Env.rest, [s_eval, :p1, [extend, :e, :v, :p2]]])
axiom_func_f2 = Axiom([f_eval, [func_call, [procedure, :v, :p1], :p2], :e], [f_eval, :p1, [extend, :e, :v, :p2]])
axiom_seq_s = Axiom([s_eval, [Seq.cons, :p1, :p2], :e], [s_eval, :p2, [s_eval, :p1, :e]])
axiom_seq_f = Axiom([f_eval, [Seq.cons, :p1, :p2], :e], [f_eval, :p2, [s_eval, :p1, :e]])
axioms = Dict()
axioms_lookup = [axiom_lookup1, axiom_lookup2, axiom_lookup3]
axioms_set = [axiom_set1, axiom_set2, axiom_set3]
axioms[:base_env] = [Stream.axioms, Binding.axioms, Frame.axioms, Env.axioms]
axioms[:env] = [axioms_lookup, axioms_set, axiom_add_binding, axiom_def, axiom_extend]
axioms_v = [axiom_var_s, axiom_var_f]
axioms_d = [axiom_definition_s, axiom_definition_f]
axioms_a = [axiom_assignment_s, axiom_assignment_f]
axioms_p = [axiom_proc_s, axiom_proc_f]
axioms_f = [axiom_func_s1, axiom_func_f1, axiom_func_s2, axiom_func_f2]
axioms_s = [axiom_seq_s, axiom_seq_f]
axioms[:prog] = [axioms_v, axioms_d, axioms_a, axioms_p, axioms_f, axioms_s];
## The Repl facility, note that for the program structure parsing, we rely on the julia parser;
## and we also need to read the variable name from string.
include("Repl.jl")
r = REPL(Env.empty, Seq.empty)
# +
function object_init(dict :: Dict, name, num :: Int64, class :: Class)
dict[Symbol(name, num)] = Object(string(name, num), class)
end
function object_init(dict :: Dict, name, class :: Class)
dict[Symbol(name)] = Object(string(name), class)
end
function object_init(dict :: Dict, name_dict :: Dict, num :: Int64)
for k in keys(name_dict)
object_init(dict, k, name_dict[k])
for i in 1:num
object_init(dict, k, i, name_dict[k])
end
end
end
function object_init(dict :: Dict, name_dict :: Dict)
for k in keys(name_dict)
object_init(dict, k, name_dict[k])
end
end
# +
type Memory
class :: Class
mclass :: Class
encode :: DFunction
decode :: DFunction
axioms :: Array{Axiom, 1}
end
function Base.show(io :: IO, m :: Memory)
print(io, string("Module.Memory(", m.class.class_name, "->", m.mclass.class_name, ")"))
end
function Memory(class :: Class, mclass :: Class, symbols = [:s, :m])
encode = DFunction("encode", [class], mclass)
decode = DFunction("decode", [mclass], class)
sym = symbols[1]
msym = symbols[2]
axiom1 = Axiom([decode, [encode, sym]], sym)
axiom2 = Axiom([encode, [decode, msym]], msym)
axioms = [axiom1, axiom2]
Memory(class, mclass, encode, decode, axioms)
end
function Memory(class :: Class, type_len :: Int64, symbols = [:s, :m])
memory_name = string("Memory", "_", class.class_name)
mclass = Class(memory_name, type_len)
Memory(class, mclass, symbols)
end
## @time l = List(Sensor, 70)
# +
naive_randomize = distribute(function(o :: Object) randn!(o.value) end)
function another_naive_randomize(tree :: Tree)
foreach(another_naive_randomize, tree.subtrees)
if typeof(tree.op) == Object
randn!(tree.op.value)
end
tree
end
# +
# UChar = Class("Char", 8)
# Variable = Class("Variable", 100)
# Prog = Class("Prog", 100)
# Bindingc = Class("Binding", 100)
# Binding = Pair(Variable, Prog, Bindingc, [:b, :v, :p])
# Framec = Class("Frame", 100)
# Frame = List(Bindingc, Framec, [:f1, :b, :f2])
# Envc = Class("Env", 200)
# Env = List(Framec, Envc, [:e1, :f, :e2])
dict = Dict()
ndict = Dict()
ndict[:u] = UChar
ndict[:v] = Variable
ndict[:p] = Prog
ndict[:b] = Bindingc
ndict[:f] = Framec
ndict[:e] = Envc
object_init(dict, ndict, 2)
# -
@time init_axioms!(axioms, dict)
@time anti_train_axioms!(axioms, 1000, another_naive_randomize)
## init_axioms!(axioms, dict)
## train_axioms!(axioms, dict, 1000)
## @time anti_train_axioms!(axioms, 10000, another_naive_randomize)
@time train_axioms!(axioms, dict, 1000, naive_randomize)
a1 = repl(r, "function(x) x end").value;
a2 = repl(r, "var y = function(x) x end").value;
a3 = repl(r, "y").value;
a4 = repl(r, "function(x) x(x) end").value;
a5 = repl(r, "begin var y = function(x) x end; y end").value;
as = [a1, a2, a3, a4, a5];
pair_computation(function(x, y) sum(abs(as[x] - as[y])) end, 1:5)
# +
Interpreter = Dict()
I_names = Dict()
I_names[:base] = [:UChar, :Variable, :Stream, :Prog, :Bindingc, :Binding, :Framec, :Frame, :Envc, :Env]
I_names[:env] = [:lookup, :set, :add_binding, :def, :extend]
I_names[:prog] = [:var, :definition, :assignment, :procedure, :func_call, :Seq]
I_names[:repl] = [:f_eval, :s_eval]
inject(I_names, Interpreter)
Interpreter
save("test.jld", Interpreter)
| NextGeneration3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Week 8: SVM Classifier
#
# Instructor: <NAME> <br>
# Email: <EMAIL> <br>
#
#
# Citations: <br>
# - Chapter 3: Python Machine Learning 3rd Edition by [<NAME>](https://sebastianraschka.com), Packt Publishing Ltd. 2019
# - https://jakevdp.github.io/PythonDataScienceHandbook/05.07-support-vector-machines.html
# - https://pythonmachinelearning.pro/using-neural-networks-for-regression-radial-basis-function-networks/
# ### Objectives:
# - support vector machines for classification.
# - regularization.
# - breakout room exercise.
# ### Support Vector Machines (SVM):
# - to make classification predictions, SVM rely on few support vectors (suport vectors = observations in the data).
# - as a result, they take up very little memory (!).
# - their relyability on support vectors (observations near the margins) makes them very suitable for high-dimensional data (even when there are more features than observations).
# - finally, SVM is able to adapt to many types of data (linearly separable or not) due to the integration of kernel methods.
#
# ##### The not so good news:
# - the results can change significantly depending on the choice of the regularization parameter C (use cross-validation to choose it).
# - no direct probability interpretation (but easily added using the probability parameter in cross-validation).
# - and a final tought: be aware that the bigger the data the more expensive cross-validation is.
# ### Linear SVM
# - This model performs well on linearly separable classes. Linear logistic regression and linear SVM often yield very similar results.
# - Logistic regression tries to maximize the conditional likelihood of the training data; so this means that it pays equal attention to outliers.
# - SVM avoids this problem, because it cares mostly about the points that are closest to the decission boundary (support vectors).
# ### Nonlinear SVM (kernel SVM)
# - SVM can be easily **kernalized** to solve nonlinear classification problems.
# - The idea is to create nonlinear combinations of the original features, and then project them onto a higher-dimensional space via a mapping function.
# ### Step 1: Import packages
# +
# standard libraries
import pandas as pd
import numpy as np
import os
# data visualization
import matplotlib.pyplot as plt
# %matplotlib inline
from IPython.display import Image
# data preprocessing
from sklearn import datasets
from sklearn.preprocessing import StandardScaler
from sklearn import preprocessing
# prediction models
from sklearn.svm import SVC
# -
# ### Step 2: Define working directories
# ### Step 3: Define classes
# ### Step 4: Define functions
def wine_data():
"""Read the wine dataset from 'https://archive.ics.uci.edu/ml/machine-learning-databases/wine/wine.data'
# param: None
# return df and X,y np.arrays for training and test (cleaning included)
"""
# read data
df = pd.read_csv('https://archive.ics.uci.edu/'
'ml/machine-learning-databases/wine/wine.data',
header=None)
df.columns = ['class_label', 'alcohol', 'malic_acid', 'ash',
'alcalinity_of_ash', 'magnesium', 'total_pphenols',
'flavanoids', 'nonflavanoid_phenols', 'proanthocyanins',
'color_intensity', 'hue', 'OD280/OD315_of_diluted_wines',
'proline']
print('Shape of df wine:', df.shape)
# recode class labels (from 0 to 2)
class_mapping = {label: idx for idx, label in enumerate(np.unique(df.class_label))}
class_mapping
df['class_label'] = df.class_label.map(class_mapping)
# select only labels 0 and 1
df = df[df.class_label !=2]
# select only 2 features
labels = ['class_label']
features = ['alcohol', 'ash']
df = df[labels+features]
print('Class labels:', df['class_label'].unique())
print('Features:', df.columns[1:])
# create X and y arrays
X = np.array(df.iloc[:, 1:])
y = np.array(df.iloc[:, 0])
# standardize features
df['alcohol'] = preprocessing.scale(df['alcohol'])
df['ash'] = preprocessing.scale(df['ash'])
X = preprocessing.scale(X)
return df, X, y
def plot_svc_decision_boundary(model, ax=None, plot_support=True):
"""Plot the decision function for a 2D SVC"""
if ax is None:
ax = plt.gca()
xlim = ax.get_xlim()
ylim = ax.get_ylim()
# create grid to evaluate model
x = np.linspace(xlim[0], xlim[1], 30)
y = np.linspace(ylim[0], ylim[1], 30)
Y, X = np.meshgrid(y, x)
xy = np.vstack([X.ravel(), Y.ravel()]).T
P = model.decision_function(xy).reshape(X.shape)
# plot decision boundary and margins
ax.contour(X, Y, P, colors='k',
levels=[-1, 0, 1], alpha=0.5,
linestyles=['--', '-', '--'])
# plot support vectors
if plot_support:
ax.scatter(model.support_vectors_[:, 0],
model.support_vectors_[:, 1],
s=100, linewidth=1, facecolors='none', edgecolor='black');
ax.set_xlim(xlim)
ax.set_ylim(ylim)
# ---
# ### Step 5: Linear SVM example
# ---
# #### Step 5.1 Read Iris data
# +
s = os.path.join('https://archive.ics.uci.edu', 'ml',
'machine-learning-databases', 'iris','iris.data').replace('\\', '//')
print('URL:', s)
df = pd.read_csv(s,
header=None,
encoding='utf-8')
print('Shape if Iris:', df.shape)
df.head()
# -
# print type of flowers
df[4].unique()
# #### Step 5.2 Data preprocessing
# We will consider only two flower classes (Setosa and Versicolor) for practical reasons.
#
# We will also restrict the analysis to only two feature variables, sepal length and petal length (easier to visualize the decission boundary in a 2D space).
# +
# select setosa and versicolor
y = df.iloc[0:100, 4].values
y = np.where(y == 'Iris-setosa', -1, 1)
# extract sepal length and petal length
X = df.iloc[0:100, [0, 2]].values
# plot head of X
X[:5]
# -
# Let's also standardize the variables for optimal performance.
# +
sc = StandardScaler()
# estimate the sample mean and standard deviation for each feature in X_train
sc.fit(X)
# use the two parameters to standardize both X_train and X_test
X_std = sc.transform(X)
# we don't standardize y but let's rename it to be consistent with notation
y_std = y
# plot head of X_std
X_std[:5]
# -
# Finally, let's visualize the data to understand what we are trying to predict (classify)
# +
xfit = np.linspace(-2, 2.5)
# plot X data for setosa
plt.scatter(X_std[:50, 0], X_std[:50, 1],
color='red', marker='o', label='setosa')
# plot X data for versicolor
plt.scatter(X_std[50:100, 0], X_std[50:100, 1],
color='blue', marker='x', label='versicolor')
plt.xlabel('sepal length [cm]')
plt.ylabel('petal length [cm]')
plt.legend(loc='upper left')
# -
# The linear SVM classifier would attempt to draw a straight line separating the two sets of data to create a model for classification.
#
# But there is a problem: there is more than one possible dividing line (decision boundary) that can perfectly discriminate between the two classes!
# +
xfit = np.linspace(-2, 2.5)
# plot X data for setosa
plt.scatter(X_std[:50, 0], X_std[:50, 1],
color='red', marker='o', label='setosa')
# plot X data for versicolor
plt.scatter(X_std[50:100, 0], X_std[50:100, 1],
color='blue', marker='x', label='versicolor')
# plot decision boundaries (class separating lines)
#for m, b in [(0.2, -0.2)]:
for m, b in [(0.2, -0.2), (0.25, -0.25)]:
#for m, b in [(0.2, -0.2), (0.25, -0.25), (0.28, -0.28)]:
plt.plot(xfit, m * xfit + b, '-k')
plt.xlabel('sepal length [cm]')
plt.ylabel('petal length [cm]')
plt.legend(loc='upper left')
# -
# So how do we choose the optimal line?
# Linear SVM intuition: rather than simply drawing a zero-width line between the two classes, we can draw around each line a margin of some width, up to the nearest point (observation).
#
# The line that maximizes this margin is the one we will choose as the optimal model. Thus, SVM are an example of a maximum margin estimator.
# +
xfit = np.linspace(-2, 2.5)
# plot X data for setosa
plt.scatter(X_std[:50, 0], X_std[:50, 1],
color='red', marker='o', label='setosa')
# plot X data for versicolor
plt.scatter(X_std[50:100, 0], X_std[50:100, 1],
color='blue', marker='x', label='versicolor')
# plot decision boundaries with margins
for m, b, d in [(0.2, -0.2, 0.25), (0.25, -0.25, 0.55)]:
yfit = m * xfit + b
plt.plot(xfit, yfit, '-k')
plt.fill_between(xfit, yfit - d, yfit + d, edgecolor='none',
color='#AAAAAA', alpha=0.4)
plt.xlabel('sepal length [cm]')
plt.ylabel('petal length [cm]')
plt.legend(loc='upper left')
# -
#
# How does it compare with the perceptron model? Well, in the perceptron model, the objective function is to minimize the classification error.
#
# In SVM we want to choose the weights, **w**, to maximize the margin (this is the cost function):
#
# $\mathbf{\frac{w^T(x\_pos - x\_neg)}{||w||}} = \frac{2}{\mathbf{||w||}}$,
#
# where $x\_pos$ are the points to the right of the decision boundary and $x\_neg$ are the points to the left.
#
# #### Step 5.3 Prediction for linear SVM
svm_iris = SVC(kernel='linear', C=1) # C sets regularization
svm_iris.fit(X_std, y_std)
# Let's visualize the optimal decission boundary and it's associated margin width.
# +
# plot X data for setosa
plt.scatter(X_std[:50, 0], X_std[:50, 1],
color='red', marker='o', label='setosa')
# plot X data for versicolor
plt.scatter(X_std[50:100, 0], X_std[50:100, 1],
color='blue', marker='x', label='versicolor')
# add labels
plt.xlabel('sepal length [cm]')
plt.ylabel('petal length [cm]')
plt.legend(loc='upper left')
# plot decision boudary
plot_svc_decision_boundary(svm_iris)
# -
#
# The continous grey line is the dividing line that maximizes the margin between the two sets of points.
#
# Notice that a few of the training points just touch the margin (points highlighted with black circles). These points are the important piece of the SVM solution, and are known as the support vectors. SVM relies mostly on these support vectors to make predictions!
#
# In Scikit-Learn, the identity of these points are stored in the support_vectors_ attribute of the classifier.
svm_iris.support_vectors_
# A key to the SVM's success is that for the fit, only the position of the support vectors matter; any points further from the margin which are on the correct side do not modify the fit!
#
# The reason? these points do not contribute to the loss function used to fit the model, so their position and number do not matter as long as they do not cross the margin.
#
# We can see this, for example, if we plot the model learned from X and the model learned from X where standardized petal length < 1cm (remove the top points):
# +
# from X array keep only if standardized petal length (index 1 in X_std or index 2 in concatenated (y, X_std)) < 1
temp = pd.concat((pd.DataFrame(y),pd.DataFrame(X_std)),axis=1)
temp = temp[temp.iloc[:,2]<1]
X_std2 = np.array(temp.iloc[:,[1,2]])
y_std2 = np.array(temp.iloc[:,0])
X_std2.shape
# -
svm_iris2 = SVC(kernel='linear', C=1) # C sets regularization
svm_iris2.fit(X_std2, y_std2)
# +
## create subplots
models = ['svm_iris', 'svm_iris2']
plt.figure(figsize=(12, 4))
for i in range(len(models)):
# create sublots that are all on the same row
ax = plt.subplot(1, len(models), i+1)
# plot X data for setosa
if i == 0:
plt.scatter(X_std[:50, 0], X_std[:50, 1],
color='red', marker='o', label='setosa')
else:
plt.scatter(X_std2[:50, 0], X_std2[:50, 1],
color='red', marker='o', label='setosa')
# plot X data for versicolor
if i == 0:
plt.scatter(X_std[50:, 0], X_std[50:, 1],
color='blue', marker='x', label='versicolor')
else:
plt.scatter(X_std2[50:, 0], X_std2[50:, 1],
color='blue', marker='x', label='setosa')
# plot decision boundary
if i == 0:
plot_svc_decision_boundary(svm_iris)
else:
plot_svc_decision_boundary(svm_iris2)
# set y limit
plt.ylim(-1.5, 1.5)
plt.title('Model: '+ str(models[i]), size=14)
# -
# In the left panel, we see the model and the support vectors for our initial svm model. In the right panel, we droped the points where standardized petal length < 1cm (remove the top points), but the model has not changed.
#
# The support vectors from the left panel are still the support vectors from the right panel! This insensitivity to distant points is one of the strengths of the SVM model!
# ---
# ### Step 6: Nonlinear (kernel) SVM example
# ---
# The power of SVM becomes more obvious when we combine it with kernels. These kernels are very useful when dealing with nonlinearly separable data.
#
# To give you an understanding of what a kernel is, think about the Linear Regression classifier. This classifier is used to fit linearly separable data, but if we project the data into a higher dimensional space (e.g., by adding polynomials and Gaussian basis functions), we can actually fit a nonlinear relationsip with this linear classifier.
# #### Step 6.1 Read wine data
# Now we will turn our attention to the wine dataset.
#
# Let's focus only on two class labels (cultivar 0 and 1) and two features ('alcohol' and 'ash').
#
# To save time and space , I defined the wine_data() function to import the data, keep only classes and features of interest, and standardizes the features.
df, X_std, y_std = wine_data()
df.head()
# #### Step 6.2 Data vizualization
#
# to understand what we are trying to predict (classify)
plt.scatter(df.loc[df.class_label==0, 'alcohol'], df.loc[df.class_label==0, 'ash'], label='cultivar0')
plt.scatter(df.loc[df.class_label==1, 'alcohol'], df.loc[df.class_label==1, 'ash'], label='cultivar1')
plt.xlabel('alcohol');
plt.ylabel('ash');
plt.legend(loc='upper left');
# #### Step 6.3 Prediction for linear SVM
svm_wine = SVC(kernel='linear', C=1.0)
svm_wine.fit(X_std, y_std)
# Let's visualize the optimal decission boundary and it's associated margin width.
# +
plt.scatter(df.loc[df.class_label==0, 'alcohol'], df.loc[df.class_label==0, 'ash'], label='cultivar0')
plt.scatter(df.loc[df.class_label==1, 'alcohol'], df.loc[df.class_label==1, 'ash'], label='cultivar1')
plt.xlabel('alcohol');
plt.ylabel('ash');
plt.legend(loc='upper left');
plot_svc_decision_boundary(svm_wine, plot_support=True);
# -
# It is clear that no linear discrimination will be able to separate this data.
#
# How we can go around it?
#
# Let's think about how we might project the data into a higher dimensional space (i.e., add more features) such that a linear separator would be sufficient.
#
# For example, one simple projection we could use would be to compute a radial basis function (RBF, also known as Gaussian kernel) **centered on the middle clump**.
# +
def rbf(X, gamma):
""" Code up Gaussian RBF
# param X: np.array containing features of interest
# param gamma: float, a free parameter, a cut-off for the Gaussian sphere (in scikit-learn gamma = 1 / (n_features * X.var()))
# param s: float, standard deviations of the clusters.
"""
return np.exp(-gamma* (np.subtract(X[:,0], X[:,1])**2))
rbf = rbf(X_std, 10) # hint: try changing the value of gamma and see what happens with the separating plane!
print(rbf.min())
print(rbf.max())
print (rbf.shape)
# -
# We can visualize this extra data dimension using a three-dimensional plot
# +
from mpl_toolkits import mplot3d
from ipywidgets import interact, fixed
ax = plt.subplot(projection='3d')
ax.scatter3D(X_std[:, 0], X_std[:, 1], rbf, c=y_std, s=50, cmap='autumn')
ax.view_init(elev=10, azim=30)
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_zlabel('r');
# -
# We can see that with this additional dimension, the data did not really become linearly separable. Linearly separable == we can draw a clearly separating plane at, say, r=0.01.
#
# After projection, we would like our data to look like in the figure below.
Image(filename='ideal_rbf.png', width=350)
# So we can conclude that we did not carefully tune our projection.
#
# We should have searched for the value of gamma that centers our RBF in the right location in order to see such a clea, linearly separable data.
#
# But randomly choosing the best RBF funtion is time consumption, so we would like to find this automatically. The kernalized SVM in scikit-learn does this automatically for us by changing our linear kernel to an RBF, using the kernel model hyperparameter!
# #### Step 6.4 Prediction for nonlinear (kernel) SVM
svm_wine = SVC(kernel='rbf', gamma=0.1, C=1.0)
svm_wine.fit(X_std, y_std)
# Let's look at the plots now
# +
plt.scatter(df.loc[df.class_label==0, 'alcohol'], df.loc[df.class_label==0, 'ash'], label='cultivar0')
plt.scatter(df.loc[df.class_label==1, 'alcohol'], df.loc[df.class_label==1, 'ash'], label='cultivar1')
plt.xlabel('alcohol');
plt.ylabel('ash');
plt.legend(loc='upper left');
plot_svc_decision_boundary(svm_wine, plot_support=True);
# -
# Using this kernelized support vector machine, we learn a suitable nonlinear decision boundary.
#
#
# Question: What are the support vectors used to classify examples? (show them on the graph)
svm_wine.support_vectors_
# #### Step 6.4.1 The role of the gamma parameter
# This parameter can be understood as the cut-off point for the Gaussian sphere.
#
# If we increase the value of gamma we increase the power of training examples (look at the RBF formula above to see the connection), which leads to a tighter and bumpier decision boundary.
#
# If we fit the training data very well (high gamma value) then we would likely end up with a high generalization error on unseen (test) data.
svm_wine = SVC(kernel='rbf', gamma=0.5, C=5) # change gamma from 0.1 to 5
svm_wine.fit(X_std, y_std)
# +
plt.scatter(df.loc[df.class_label==0, 'alcohol'], df.loc[df.class_label==0, 'ash'], label='cultivar0')
plt.scatter(df.loc[df.class_label==1, 'alcohol'], df.loc[df.class_label==1, 'ash'], label='cultivar1')
plt.xlabel('alcohol');
plt.ylabel('ash');
plt.legend(loc='upper left');
plot_svc_decision_boundary(svm_wine, plot_support=True);
# -
# #### Step 6.4.2 The role of the regularization (C) parameter
# A familiar concept? Bias vs. Variance tradeoff
# If a model suffers from overfitting, we also say that the model has high variance (the model is too complex, fits very well the training data).
#
# If a model suffers from underfitting, we also say that the model has high bias (the model is not complex enough to capture the patern in the training data).
# Decreasing the value of C, increases the bias and lowers the variance. So when we decrease the value of C we increase the width of the margin.
# +
C = [20.0, 0.1]
plt.figure(figsize=(12, 4))
for i, c in enumerate(C):
# create sublots that are all on the same row
ax = plt.subplot(1, len(models), i+1)
svm_wine = SVC(kernel='linear', C=c)
svm_wine.fit(X_std, y_std)
plt.scatter(df.loc[df.class_label==0, 'alcohol'], df.loc[df.class_label==0, 'ash'], label='cultivar0')
plt.scatter(df.loc[df.class_label==1, 'alcohol'], df.loc[df.class_label==1, 'ash'], label='cultivar1')
plt.xlabel('alcohol');
plt.ylabel('ash');
plt.legend(loc='upper left');
plot_svc_decision_boundary(svm_wine, plot_support=True)
if i == 0:
plt.title('C = {0:.1f}'.format(c) + str(', i.e. large value of C'), size=14)
else:
plt.title('C = {0:.1f}'.format(c) + str(', i.e. small value of C'), size=14)
# -
# ### Additional exercises
# Using the wine dataset, execute the following tasks:
#
# - [1] split the data into train and test examples:
# - use train_test_split() method from scikit-learn's preprocessing module; set test_size parameter to 0.30;
# - report number of observations in the train and test data;
#
# ``Add answer here:`` train = 91, test = 39
# <br>
#
# - [2] fit a linear SVM on training data and predict on test data:
# - set C hyperparameter to 1.0;
# - report classification error for training and test data;
#
# ``Add answer here:`` 92.3% accuracy
# <br>
#
# - [3] fit a kernel SVM on training data and predict on test data:
# - set gamma and C hyperparameters to 0.20 and 10.0, respectively;
# - report classification error for training and test data;
#
# ``Add answer here:`` 92.3% accuracy
# <br>
#
# - [4] fit a kernel SVM on training data and predict on test data:
# - set gamma and C hyperparameters to 0.20 and 1.0, respectively;
# - report classification error for training and test data;
#
# ``Add answer here:`` 94.9% accuracy
# <br>
# +
# [1]
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X_std, y_std, test_size = 0.3)
print(len(X_train),len(y_train),len(X_test),len(y_test))
# +
# [2]
from sklearn import metrics as mt
svm_wine_2 = SVC(kernel='linear', C=1.0)
svm_wine_2.fit(X_train, y_train)
y_hat_2 = svm_wine_2.predict(X_test) # get test set predictions
acc_2 = mt.accuracy_score(y_test,y_hat_2)
conf_2 = mt.confusion_matrix(y_test,y_hat_2)
print('accuracy:', acc_2 )
print(conf_2)
# +
# [3]
svm_wine_3 = SVC(kernel='rbf', gamma=0.2, C=10.0)
svm_wine_3.fit(X_train, y_train)
y_hat_3 = svm_wine_3.predict(X_test) # get test set predictions
acc_3 = mt.accuracy_score(y_test,y_hat_3)
conf_3 = mt.confusion_matrix(y_test,y_hat_3)
print('accuracy:', acc_3 )
print(conf_3)
# +
# [4]
svm_wine_4 = SVC(kernel='rbf', gamma=0.2, C=1.0)
svm_wine_4.fit(X_train, y_train)
y_hat_4 = svm_wine_4.predict(X_test) # get test set predictions
acc_4 = mt.accuracy_score(y_test,y_hat_4)
conf_4 = mt.confusion_matrix(y_test,y_hat_4)
print('accuracy:', acc_4 )
print(conf_4)
| ds7333_case_study_5/ML1_SVM_classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Image Classification
import cv2
import matplotlib.pyplot as plt
img_path = 'sample_images/000006_04.jpg'
# Load color image
bgr_img = cv2.imread(img_path)
# Convert to grayscale
gray_img = cv2.cvtColor(bgr_img, cv2.COLOR_BGR2GRAY)
# Normalize, rescale entries to lie in [0,1]
gray_img = gray_img.astype("float32")/255
# Plot image
plt.imshow(gray_img, cmap='gray')
plt.show()
import numpy as np
filter_vals = np.array([[1, -1, 1], [1, -1, 1], [1, -1, 1]])
print('Filter shape: ', filter_vals.shape)
# Define four different filters, all of which are linear combinations of the `filter_vals` defined above
filter_1 = filter_vals
filter_2 = -filter_1
filter_3 = filter_1.T
# filter_4 = -filter_3
filters = np.array([filter_1, filter_2, filter_3])
# Print out the values of filter 1 as an example
print('Filter 1: \n', filter_1)
import torch
import torch.nn as nn
import torch.nn.functional as F
# Neural network with one convolutional layer with four filters
class Net(nn.Module):
def __init__(self, weight):
super(Net, self).__init__()
# Initializes the weights of the convolutional layer to be the weights of the 4 defined filters
k_height, k_width = weight.shape[2:]
# Assumes there are 3 grayscale filters
self.conv = nn.Conv2d(1, 3, kernel_size=(k_height, k_width),bias=False)
self.conv.weight = torch.nn.Parameter(weight)
def forward(self, x):
# Calculates the output of a convolutional layer pre- and post-activation
conv_x = self.conv(x)
activated_x = F.relu(conv_x)
# Returns both layers
return conv_x, activated_x
# Instantiate the model and set the weights
weight = torch.from_numpy(filters).unsqueeze(1).type(torch.FloatTensor)
model = Net(weight)
# Print out the layer in the network
print(model)
def viz_layer(layer, n_filters= 3):
fig = plt.figure(figsize=(20, 20))
for i in range(n_filters):
ax = fig.add_subplot(1, n_filters, i+1, xticks=[], yticks=[])
# Grab layer outputs
ax.imshow(np.squeeze(layer[0,i].data.numpy()), cmap='gray')
ax.set_title('Output %s' % str(i+1))
# Plot original image
plt.imshow(gray_img, cmap='gray')
# Visualize all of the filters
fig = plt.figure(figsize=(12, 6))
fig.subplots_adjust(left=0, right=1.5, bottom=0.8, top=1, hspace=0.05, wspace=0.05)
for i in range(3):
ax = fig.add_subplot(1, 3, i+1, xticks=[], yticks=[])
ax.imshow(filters[i], cmap='gray')
ax.set_title('Filter %s' % str(i+1))
# Convert the image into an input tensor
gray_img_tensor = torch.from_numpy(gray_img).unsqueeze(0).unsqueeze(1)
# Get the convolutional layer (pre and post activation)
conv_layer, activated_layer = model(gray_img_tensor)
# Visualize the output of a convolutional layer
viz_layer(conv_layer)
# convert to grayscale for filtering
gray = cv2.cvtColor(bgr_img, cv2.COLOR_RGB2GRAY)
plt.imshow(gray, cmap='gray')
# 3x3 array for edge detection
sobel_y = np.array([[ -1, -2, -1],[ 0, 0, 0], [ 1, 2, 1]])
sobel_x = np.array([[ -1, 0, 1], [ 0, 0, 0],[ 1, 2, 1]])
filtered_image = cv2.filter2D(gray, -1, sobel_y)
plt.imshow(filtered_image, cmap='gray')
viz_layer(activated_layer)
| Filters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="fv1F1HyA9zmX"
# # Capture and organize data in downloaded files
# + [markdown] id="EsXo4Upb9zmb"
# We downloaded these ```.txt``` and ```.pdf``` files in our most recent scrape. We won't scrape them again.
#
# We will read the text files using Python and export data to csv. (are you seeing a pattern yet?)
# + [markdown] id="lyvc6CPd-FUC"
# # Import libraries
#
# + id="m2ru6dyM9zmb"
## in order to export our file to our computer drive, you need this only in Colab:
from google.colab import files
import os ## for file/folder management
from pathlib import Path ## to provide a path to files/folders
## import the glob library for collecting specific files into a list
import glob
import pandas as pd ## to export csv file
# + [markdown] id="AB2_Oy8HlCRW"
# ## where am i?
# + id="CAuO3kqilCRW"
# + id="Ezd1IYJQD95X"
## let's capture the files in a list
## unlike earlier when when we capture the locations,
## these are the actual files with their contents.
# + id="fvax9kwamuMx"
#let's turn each file into readable content
# + [markdown] id="OdcgUwoknrrd"
# ## We can interpret this ```<class '_io.TextIOWrapper'>``` to read the actual contents
# + id="fqN9xmTuD-Ao"
## let's see what the first line of each file contains
# + id="WmZYM2D4D99U"
## let's see what each entire file contains
# + id="oU_37TIvD-Dl"
## let read all the lines and put into a list
## let's see what the first line of file contains
# + id="y72rctb5D-Gu"
## Now let's place clients and decisions into variables called client and decision
# + id="5yuvCxJoD-KT"
## let's remove the word client and the extra line
# + id="opp5s8STjCye"
## We don't want an entire sentence – just what the decision was.
## we just want to know the status of lease in one word renew or terminate
# + id="9hEhNKkXjYHT"
## We want to store in a list to export as CSV file
# + [markdown] id="isxnLpPRlCRa"
# ### Confirm where we are path-wise
# + id="D-Qc3BZ8lCRa"
# + [markdown] id="k9KTw-enlCRb"
# ### Create new results directory (note we come out of the downloaded_files folder first)
# + id="WglOFfLHlCRb"
# + id="otpHcPn4lCRb"
# ### cd into our results folder
# + [markdown] id="oKYRUg24lCRb"
# ### Confirm we are in the results folder
# + id="F-AMX1rIlCRb"
# + id="sIw6MAQNkazG"
## Export as CSV
## use pandas to write to csv file
## we already imported pandas as pd
filename = "lease_decisions.csv" ## what are file name is
df = pd.DataFrame(decisions) ## we turn our list of dicts into a dataframe which we're call df
df
df.to_csv(filename, encoding='utf-8', index=False) ## export to csv as utf-8 coding (it just has to be this)
print(f"{filename} is in your results folder!") ## a print out that tells us the file is ready
| in-class/.ipynb_checkpoints/week-8B-download-and-read_BLANK-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from sklearn import preprocessing
import sys
# The test hyper-parameter, default is argv_list = []
argv_list = ['complex', 'abstract']
#argv_list = []
# +
# load the data from text
data_total = pd.read_csv('../Gene_DATA/sourcePathway.txt')
# extract the info based on regex
Namelist = data_total['TXT'].str.extract(r'^(?P<GenomeType>[a-z]+)\s(?P<GenomeName>[\w\/\-()+]+)$').dropna(axis=0).reset_index(drop=True)
Edgelist = data_total['TXT'].str.extract(r'^(?P<edgeStart>[\w\/\-()+]+)\s(?P<edgeEnd>[\w\/\-()+]+)\s(?P<edgeType>[\w\>\|-]+)$').dropna(axis=0).reset_index(drop=True)
# -
if len(argv_list):
# Here we started to choose to discard some features/node types
le = preprocessing.LabelEncoder()
le.fit(Namelist['GenomeType'])
all_node_class = list(le.classes_)
# Input error check
for elem in argv_list:
if elem not in all_node_class:
raise NameError('The input argument {} is not defined in node class.'.format(elem))
sys.exit()
# Start to exclude the node type
Namelist = Namelist[~Namelist['GenomeType'].isin(argv_list)].reset_index(drop=True)
# +
# The beforehand check
# Here we check the edge that does not showup in node list and drop these edges
Namelist_l = list(Namelist['GenomeName'])
Edgelist_l = list(Edgelist.iloc[:,0].values)
Edgelist_ll = list(Edgelist.iloc[:,1].values)
exclude_list = []
for idx, (elem, elem2) in enumerate(zip(Edgelist_l, Edgelist_ll)):
if ((elem not in Namelist_l) or (elem2 not in Namelist_l)):
exclude_list.append(idx)
Edgelist = Edgelist.drop(exclude_list).reset_index(drop=True)
# +
# Label the index of each edge
# Make Edgelist --> edge_index
# Label edge_index
le = preprocessing.LabelEncoder()
le.fit(Namelist['GenomeName'])
edge_index = le.transform(Edgelist.iloc[:,:2].values.reshape(-1)).reshape(-1,2)
# Label edge_class
le = preprocessing.LabelEncoder()
le.fit(Edgelist['edgeType'])
edge_class = le.transform(Edgelist['edgeType'])
# Label node class
le = preprocessing.LabelEncoder()
le.fit(Namelist['GenomeType'])
node_class = le.transform(Namelist['GenomeType'])
# The combination of node info
node_all = np.concatenate([Namelist['GenomeName'].values.reshape(-1,1), node_class.reshape(-1,1)], axis=1)
# -
node_all
| GenomicData/test_notebook/data_fetcher.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Mini
# language: python
# name: mini
# ---
import cv2
import numpy as np
# +
cap = cv2.VideoCapture(0)
while cap.isOpened():
ret, frame = cap.read()
if not ret:
break
gray = cv2.cvtColor(frame, cv2.COLOR_BGR2GRAY)
canny = cv2.Canny(gray, 50, 200)
canny = np.dstack((canny, canny, canny))
concat = np.hstack((frame, canny))
cv2.imshow('frame', concat)
key = cv2.waitKey(1)
if key == ord('q'):
break
cap.release()
cv2.destroyAllWindows()
| Pratica1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="http://landlab.github.io"><img style="float: left" src="../../../landlab_header.png"></a>
# # Component Overview: `DepthDependentTaylorDiffuser`
# <hr>
# <small>For more Landlab tutorials, click here: <a href="https://landlab.readthedocs.io/en/latest/user_guide/tutorials.html">https://landlab.readthedocs.io/en/latest/user_guide/tutorials.html</a></small>
# <hr>
#
# ## Introduction and background
#
# This tutorial introduces the `DepthDependentTaylorDiffuser` component, which we'll refer to here as "DDTD". The DDTD component models the process of downslope soil creep and its role in modifying topography. It combines the mathematics behind two other components: `DepthDependentDiffuser` and `TaylorNonLinearDiffuser`. The component is described (as one element in the terrainBento package) in Barnhart et al. (2019), which is the appropriate paper to cite for it.
#
# ### Theory
#
# Consider a topographic surface in which the elevation at any time $t$ and horizontal position $(x,y)$ is $\eta (x,y,t)$. The thickness of the mobile soil layer is $H(x,y,t)$. Let $\mathbf{q}_s$ be a 2D vector that represents the rate of soil volume flow per unit slope width (with dimensions of length squared per time; we'll assume that $\mathbf{q}_s$ represents a "bulk" flux that includes pore spaces between soil grains). In the absence of any "local" input sources (such as weathering of rock) or output (such as removal by wash erosion), conservation of mass dictates that:
#
# $$\frac{\partial \eta}{\partial t} = -\nabla \cdot \mathbf{q}_s$$
#
# The DDTD component represents the soil flux as:
#
# $$\mathbf{q}_s = K H_* \mathbf{S} [1 - \exp ( - H / H_*)] [ 1 + (S/S_c)^2 + (S/S_c)^4 + ... + (S/S_c)^2(n-1) ]$$
#
# where $\mathbf{S} = -\nabla \eta$ is the downslope topographic gradient, and $S$ is its magnitude. Parameter $H_*$ is a depth scale that determines how rapidly transport rate decays as the soil thins. Parameter $K$ is a transport coefficient with dimensions of velocity. The effective diffusion-like coefficient is $D=KH_*$. This is the effective diffusivity when the soil is much thicker than $H_*$.
#
# The above can be written slightly more compactly:
#
# $$\mathbf{q}_s = K H_* \mathbf{S} [1 - \exp ( - H / H_*)] \left[ 1 + \sum_{i=1}^N \left(\frac{S}{S_c}\right)^{2i} \right]$$
#
# where $i$ is the number of additional terms desired. If $i=0$, the expression is the same as the depth-dependent, slope-linear transport function implemented by the `DepthDependentDiffuser` component and described, for example, by Johnstone and Hilley (2015).
#
# The use of a truncated Taylor series is meant to approximate the Andrews-Bucknam transport function (e.g., Roering et al., 1999) while avoiding that equation's blow-up at $S=S_c$; the idea of using a truncated Taylor series comes from Ganti et al. (2012).
#
# ### Numerical implementation
#
# The component uses an explicit finite-volume solution method. Soil flux values are calculated from the gradient values on the active links, using the grid method `calc_grad_at_link`. Flux divergence is then calculated using the grid method `calc_flux_div_at_node`. The calculation updates soil thickness, bedrock elevation (using the user-provided values of the `soil_production__rate` field), and total elevation as the sum of the two.
#
# An optional dynamic timestep capability will check the local Courant condition (which can vary in time and space when nonlinear terms are included) and sub-divide the user-specified time step as needed to ensure stability.
# ## Examples
#
# ### Needed imports
#
# Like all Landlab components, DDTD requires a grid object on which to operate, so for this example we'll import `RasterModelGrid` as well as the component itself.
import numpy as np
import matplotlib.pyplot as plt
from landlab import RasterModelGrid
from landlab.components import DepthDependentTaylorDiffuser
# ### Example 1: equilibrium hillslope profile with linear diffusion
#
# For the first example, we'll use a long and skinny grid to effectively create a 1D domain. We'll test the ability of DDTD to reduce to a simple linear, depth-independent diffusive model when $i=0$ and $H \gg H_*$. We'll impose (relative) rock uplift by raising the interior of the domain at a specified rate $U$ relative to the fixed boundary nodes on either side. The expectation is that:
#
# $$\frac{d\eta}{dx} = -\frac{U}{D}x$$
#
# where $x$ is distance from the ridge top (because the ridge top will form in the middle of the domain, $x<0$ on the left and $x>0$ on the right). Integrating this, we get
#
# $$\eta = -\frac{U}{2D} x^2 + C$$
#
# We can evaluate the integration constant by noting that $\eta = 0$ at $x = \pm L$, where $L$ is the distance from base to crest. Therefore,
#
# $$\boxed{\eta = \frac{U}{2D} \left( L^2 - x^2 \right)}$$
#
# We'll test this using a hill that is 100 m long (51 nodes, two of which are fixed boundaries, with 2 m spacing between them; 50 m from base to crest on each side), a soil layer that is much thicker than the characteristic decay depth $H^*$, a transport coefficient of 0.01 m$^2$/y, and an uplift rate of 0.0001 m/y. With these parameters, the predicted ridge height (at $x=0$) is calculated below.
#
# +
# define parameters
L = 50.0 # distance from base to ridgeline, m
dx = 2.0 # node spacing, m
Hstar = 0.1 # characteristic transport depth, m
V0 = 0.1 # transport velocity coefficient, m/y
U = 0.0001 # uplift rate, m/y
H = 100.0 # initial soil thickness, m
num_steps = 20000 # number of time steps
# time step size (calculate using Courant condition for linear diffusion)
D = V0 * Hstar # effective (maximum) diffusivity
dt = 0.1 * dx * dx / D
# prediction
predicted_crest_height = 0.5 * (U / D) * L * L
print("Crest height should be " + str(predicted_crest_height))
# +
# create grid
grid = RasterModelGrid((3, 51), xy_spacing=dx)
grid.set_closed_boundaries_at_grid_edges(False, True, False, True)
# create fields
elev = grid.add_zeros("topographic__elevation", at="node") # this is eta
rock = grid.add_zeros("bedrock__elevation", at="node") # this is eta - H
rock[:] = -H
soil = grid.add_zeros("soil__depth", at="node") # this is H
soil_production_rate = grid.add_zeros("soil_production__rate", at="node")
# instantiate component
ddtd = DepthDependentTaylorDiffuser(
grid, soil_transport_velocity=V0, soil_transport_decay_depth=Hstar, nterms=1
)
# -
# run the model in a time loop with uplift applied
for i in range(num_steps):
elev[grid.core_nodes] += U * dt
rock[grid.core_nodes] += U * dt
ddtd.run_one_step(dt)
midrow = np.arange(51, 102, dtype=int)
plt.plot(grid.x_of_node[midrow], elev[midrow])
plt.xlabel("Distance (m)")
plt.ylabel("Elevation (m)")
print(np.amax(elev))
# ### Example 2: linear, depth-dependent diffusion
#
# In this example we add a rule for soil production that will limit the soil thickness and hence reduce the transport efficiency. The rate of soil production from bedrock will be:
#
# $$P = P_0 \exp ( H / H_0 )$$
#
# where $P_0$ is the maximum production rate and $H_0$ is a characteristic decay depth. In our example, we'll set $P_0$ to twice the uplift rate. At equilibrium, the actual production rate $P = U$, which means that the equilibrium soil thickness can be found from:
#
# $$P = U = 2 U \exp ( -H / H_0 )$$
#
# or
#
# $$H = -H_0 \ln 1/2$$
#
# The effective diffusion coefficient is therefore
#
# $$D_{eff} = D [1 - \exp ( -H / H_* )] = D [1 - \exp ( H_0 \ln 1/2 / H_* )]$$
#
# For the sake of example, we'll assume $H_0 = H_*$, so
#
# $$D_{eff} = D / 2$$
#
# and therefore our hill crest should be twice as high as in the prior case.
# +
# new parameter: maximum soil production rate
P0 = 2 * U # m/yr
# create grid
grid = RasterModelGrid((3, 51), xy_spacing=dx)
grid.set_closed_boundaries_at_grid_edges(False, True, False, True)
# create fields
elev = grid.add_zeros("topographic__elevation", at="node") # this is eta
rock = grid.add_zeros("bedrock__elevation", at="node") # this is eta - H
soil = grid.add_zeros("soil__depth", at="node") # this is H
soil_production_rate = grid.add_zeros("soil_production__rate", at="node")
# instantiate component
ddtd = DepthDependentTaylorDiffuser(
grid, soil_transport_velocity=V0, soil_transport_decay_depth=Hstar, nterms=1
)
# -
# run the model in a time loop with uplift applied
for i in range(num_steps):
elev[grid.core_nodes] += U * dt
rock[grid.core_nodes] += U * dt
soil_production_rate[grid.core_nodes] = P0 * np.exp(-soil[grid.core_nodes] / Hstar)
ddtd.run_one_step(dt)
plt.plot(grid.x_of_node[midrow], elev[midrow])
plt.xlabel("Distance (m)")
plt.ylabel("Elevation (m)")
# Here we haven't quite reached equilibrium yet, but we can see that the hilltop crest is approaching our expected height of 25 m: twice as high as it would be if the soil flux were not limited by soil thickness.
# ### Example 3: Nonlinear behavior
#
# When we include nonlinear terms in the transport law, we expect to see slopes that become more planar in character. We'll test this by setting a critical slope value $S_c = 0.6$ (about 31$^\circ$), and using a higher uplift rate. We'll have two terms, one linear and one cubic. We will also invoke the `dynamic_dt` option, which allows the component to subdivide each "global" timestep if needed for numerical stability: a useful thing to do because now our Courant condition varies according to slope gradient.
U = 0.0005 # uplift rate, m/yr
Sc = 0.6 # critical slope gradient, m/m
H = 1000.0 # plenty of soil
num_steps = 2000 # number of time steps
# +
# create grid
grid = RasterModelGrid((3, 51), xy_spacing=dx)
grid.set_closed_boundaries_at_grid_edges(False, True, False, True)
# create fields
elev = grid.add_zeros("topographic__elevation", at="node") # this is eta
rock = grid.add_zeros("bedrock__elevation", at="node") # this is eta - H
rock[:] = -H
soil = grid.add_zeros("soil__depth", at="node") # this is H
soil_production_rate = grid.add_zeros("soil_production__rate", at="node")
# instantiate component
ddtd = DepthDependentTaylorDiffuser(
grid,
soil_transport_velocity=V0,
soil_transport_decay_depth=Hstar,
slope_crit=Sc,
dynamic_dt=True,
nterms=2,
)
# -
# run the model in a time loop with uplift applied
for i in range(num_steps):
elev[grid.core_nodes] += U * dt
rock[grid.core_nodes] += U * dt
ddtd.run_one_step(dt)
plt.plot(grid.x_of_node[midrow], elev[midrow])
plt.xlabel("Distance (m)")
plt.ylabel("Elevation (m)")
# The resulting hill is taller (due to the higher uplift rate) and no longer has uniform convexity.
#
# How do we know whether it has reached equilibrium? One way is to inspect the soil flux: it should increase linearly with $x$, and be zero at the crest. The values at the base of the slope should equal slope length times uplift rate, or 50 m x 0.0005 m/yr = 0.025 m$^2$/yr.
active_link_midpts = (
grid.x_of_node[grid.node_at_link_tail[grid.active_links]] + 0.5 * dx
)
plt.plot(active_link_midpts, grid.at_link["soil__flux"][grid.active_links])
plt.grid(True)
plt.xlabel("Distance (m)")
plt.ylabel("Soil flux (m2/yr)")
# So it appears as if we are not quite there, but pretty close.
# ### Example 4: Nonlinear, depth dependent, and 2D
#
# In the final example we'll use a proper 2D domain, with both a soil-depth dependence and a nonlinear term in the flux law.
U = 0.0002 # uplift rate, m/yr
# +
# create grid
grid = RasterModelGrid((21, 31), xy_spacing=dx)
# create fields
elev = grid.add_zeros("topographic__elevation", at="node") # this is eta
rock = grid.add_zeros("bedrock__elevation", at="node") # this is eta - H
soil = grid.add_zeros("soil__depth", at="node") # this is H
soil_production_rate = grid.add_zeros("soil_production__rate", at="node")
# instantiate component
ddtd = DepthDependentTaylorDiffuser(
grid,
soil_transport_velocity=V0,
soil_transport_decay_depth=Hstar,
slope_crit=Sc,
dynamic_dt=True,
nterms=2,
)
# -
# run the model in a time loop with uplift applied
for i in range(num_steps):
elev[grid.core_nodes] += U * dt
rock[grid.core_nodes] += U * dt
soil_production_rate[grid.core_nodes] = P0 * np.exp(-soil[grid.core_nodes] / Hstar)
ddtd.run_one_step(dt)
# +
from landlab import imshow_grid
imshow_grid(grid, elev)
# -
# ## References
#
# Barnhart, K., <NAME>., <NAME>., <NAME>. (2019). Terrainbento 1.0: a Python package for multi-model analysis in long-term drainage basin evolution. Geoscientific Model Development 12(4), 1267--1297, [https://dx.doi.org/10.5194/gmd-12-1267-2019](https://dx.doi.org/10.5194/gmd-12-1267-2019).
#
# <NAME>., <NAME>., <NAME>. (2012). A sub-grid scale closure for nonlinear hillslope sediment transport models. Journal of Geophysical Research: Earth Surface, 117(F2), [https://dx.doi.org/10.1029/2011jf002181](https://dx.doi.org/10.1029/2011jf002181).
#
# <NAME>., <NAME>. (2015). Lithologic control on the form of soil-mantled hillslopes. Geology 43(1), 83-86, [https://doi.org/10.1130/G36052.1](https://doi.org/10.1130/G36052.1).
#
# <NAME>., <NAME>., & <NAME>. (1999). Evidence for nonlinear, diffusive sediment transport on hillslopes and implications for landscape morphology. Water Resources Research, 35(3), 853-870.
#
# Congratulations on making it to the end of this tutorial!
#
# ### Click here for more <a href="https://landlab.readthedocs.io/en/latest/user_guide/tutorials.html">Landlab tutorials</a>
| notebooks/tutorials/hillslope_geomorphology/depth_dependent_taylor_diffuser/depth_dependent_taylor_diffuser.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Wrangle Volume Data
#
# We wrangle the traffic volume data into a workable format, and extract just the site and detector we are interested in.
# ## Import data
import pandas as pd
f = pd.read_csv('../data/VSDATA_20150819.csv')
# ## Filter data
#
# Filter to site 2433 (mid-way along segment of Princes freeway monitored by bluetooth detector sites). Detectors 4-6 are in the outbound/westbound lanes.
vols = f[(f["NB_SCATS_SITE"] == 2433) & f["NB_DETECTOR"].between(4,6)]
vols
# ## Date range
#
# Extract date from CSV data
import datetime
start_date = vols["QT_INTERVAL_COUNT"].iloc[0]
start_datetime = datetime.datetime.strptime(start_date, '%Y-%m-%d 00:00:00')
date_range = pd.date_range(start_datetime, periods=96, freq='15T')
date_range[:10] # show first 10 rows
# ## Transform data
#
# Transpose table. Label by time rather than interval. Use detector number as headers.
dets = vols.T
dets.columns = dets.loc["NB_DETECTOR"].values
dets = dets.loc['V00':'V95']
dets.index=date_range
dets.head()
# ## Export data
#
# Extract just detector 6 (the rightmost lane)
d6 = dets[6]
d6.head()
# ## Plots
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
bins = np.linspace(0, max(d6), 51)
plt.hist(d6, bins=bins)
plt.show()
plt.figure(figsize=(16,8))
plt.scatter(np.arange(len(d6)), d6.values)
plt.title("Volume Site 2433 Detector 6 (Outbound along Princes Highway). Wed 19 Aug 2015.")
plt.ylabel("Travel Time (seconds)")
plt.xlabel("Time Leave (15 min offset)")
plt.xlim([0,95])
plt.ylim([0,None])
plt.show()
| jupyter_notebooks/data_vs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.0 64-bit ('3.7')
# name: python3
# ---
# # Build Classification Model
import pandas as pd
cuisines_df = pd.read_csv("../data/cleaned_cuisines.csv")
cuisines_df.head()
cuisines_label_df = cuisines_df['cuisine']
cuisines_label_df.head()
cuisines_feature_df = cuisines_df.drop(['Unnamed: 0', 'cuisine'], axis=1)
cuisines_feature_df.head()
# +
from sklearn.neighbors import KNeighborsClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.metrics import accuracy_score,precision_score,confusion_matrix,classification_report, precision_recall_curve
import numpy as np
X_train, X_test, y_train, y_test = train_test_split(cuisines_feature_df, cuisines_label_df, test_size=0.3)
# +
# SVM - support vector machines
# SVC - support vector clustering, child of above
# C - regularization, and regulates the influence of parameters
# kernel - how to cluster the labels, we set to 'linear' to make sure we get linear SVC
# set probability to true b/c we want to capture that data
# set random state to 0 to shuffle data to get probabilities
C = 10
# create the different classifiers (for others these would be models)
classifiers = {
# 'Linear SVC': SVC(kernel='linear', C=C, probability=True,random_state=0),
# points placed is created and data are gathered around these points so the labels can be predicted
# 'KNN classifier': KNeighborsClassifier(C),
# Support-Vector classifiers: map training samples to points in space to max distance btw 2 categories, later data is mapped onto space to predict category
# 'SVC': SVC(),
'RFST': RandomForestClassifier(n_estimators=100),
'ADA': AdaBoostClassifier(n_estimators=100)
}
n_classifiers = len(classifiers)
for index, (name, classifier) in enumerate(classifiers.items()):
classifier.fit(X_train, np.ravel(y_train))
y_pred = classifier.predict(X_test)
accuracy = accuracy_score(y_test, y_pred)
print("Accuracy (train) for %s: %0.1f%% " % (name, accuracy * 100))
np.set_printoptions(threshold=sys.maxsize)
print(classification_report(y_test,y_pred))
| 4-Classification/3-Classifiers-2/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
import bhc
import numpy as np
import matplotlib.pyplot as plt
from scipy.special import gamma
from scipy.cluster.hierarchy import dendrogram, linkage
import pandas as pd
def Bern_gen(nobs, k, theta, seed):
"""Generate Bernoulli distributed data"""
np.random.seed(seed)
obs_list = []
theta_list = (np.repeat(theta,nobs))
theta_list[:int(nobs/3)] = np.repeat(theta-0.3, int(nobs/3))
theta_list[-int(nobs/3):] = np.repeat(theta+0.3, int(nobs/3))
for i in range(nobs):
X_i = np.random.binomial(1, theta_list[i], k)
obs_list.append(X_i)
return np.matrix(obs_list)
def purity_score(linkage_matrix, y_test, repeats, seed):
"""Compute the expected dendrogram purity.
Sample a leaf uniformly at random. Then sample another leaf from the same
true class uniformly at random. Find their lowest common ancestor in the
tree and compute purity with respect to that class.
return purity_score
"""
np.random.seed(seed)
purity = 0
N = len(y_test)
for i in range(repeats):
class_test = np.random.choice(y_test, 1)[0]
leaf1, leaf2 = np.random.choice(np.arange(N)[np.array(y_test)==class_test], size=2, replace=None)
LL = [[item] for item in range(N)]
for j in range(linkage_matrix.shape[0]):
p, q = int(linkage_matrix[j][0]), int(linkage_matrix[j][1])
LL.append(LL[p]+LL[q])
common_ancestor = [item for item in LL if leaf1 in item and leaf2 in item][0]
predict_label = np.array(y_test)[common_ancestor]
purity += sum(predict_label==y_test[leaf1]) / len(predict_label)
return purity / repeats
# +
BHC_test = np.array(bhc.bhclust_BB(X_test)[0])
single_test = linkage(X_test,method='single')
complete_test = linkage(X_test,method='complete')
average_test = linkage(X_test,method='average')
print("BHC_test:", round(purity_score(BHC_test, y_test, 'A', 5, 12),3), purity_score(BHC_test, y_test, 'B', 5, 12), purity_score(BHC_test, y_test, 'C', 5, 12))
print("Single_linkage:", purity_score(single_test, y_test, 'A', 5, 12), purity_score(single_test, y_test, 'B', 5, 12), purity_score(single_test, y_test, 'C', 5, 12))
print("Complete_linkage:", purity_score(complete_test, y_test, 'A', 5, 12), purity_score(complete_test, y_test, 'B', 5, 12), purity_score(complete_test, y_test, 'C', 5, 12))
print("Average_linkage:", purity_score(average_test, y_test, 'A', 5, 12), purity_score(average_test, y_test, 'B', 5, 12), purity_score(average_test, y_test, 'C', 5, 12))
# -
# ### Comparative Analysis
# ### Test with multivariate data
mdat = np.array([[ 0.93637874, 1.61258974],
[ 1.95192875, 2.84452075],
[ 2.07671748, 3.24442548],
[ 3.122903 , 4.516753 ],
[ 3.56202194, 5.17531994],
[ 3.53211875, 5.75857675],
[ 4.65794237, 6.66995537],
[ 5.83738797, 8.46562797],
[ 6.22595817, 9.28082817],
[ 6.51552067, 9.36110867],
[ 7.24619975, 3.68958775],
[ 6.50554148, 3.69771048],
[ 6.58213752, 4.31283952],
[ 6.02279742, 4.52753342],
[ 5.83280398, 4.85751598],
[ 5.12305078, 4.76874878],
[ 5.0430706 , 5.2911986 ],
[ 2.44081699, 6.35402999]])
mdat_y = list(np.repeat('A',10))
mdat_y.extend(list(np.repeat('B',8)))
# +
Z = bhc.bhclust(mdat, family = "multivariate", alpha = 1, r = 0.001)[0]
Z = np.array(Z)
single_test = linkage(mdat,method='single')
complete_test = linkage(mdat,method='complete')
average_test = linkage(mdat,method='average')
print("BHC_test:", round(purity_score(Z, mdat_y, 5, 12),3))
print("Single_linkage:", round(purity_score(single_test, mdat_y, 5, 12),3))
print("Complete_linkage:", round(purity_score(complete_test, mdat_y, 5, 12),3))
print("Average_linkage:", round(purity_score(average_test, mdat_y, 5, 12),3))
# -
# ### Test with Aggregation dataset and Spiral dataset
#
# **Reference**:
#
# **Aggregation dataset**: *<NAME>, <NAME>, and <NAME>, Clustering aggregation. ACM Transactions on Knowledge Discovery from Data (TKDD), 2007. 1(1): p. 1-30.*
#
# **Spiral dataset**: *<NAME> and <NAME>, Robust path-based spectral clustering. Pattern Recognition, 2008. 41(1): p. 191-203.*
multivariate_test = pd.read_table("/Users/lina/Downloads/Aggregation.txt", names=['X1','X2','class'])
multivariate_test_spiral = pd.read_table("/Users/lina/Downloads/spiral.txt", names=['X1','X2','class'])
mvn_y = multivariate_test.iloc[:,-1]
mvn_X = multivariate_test.ix[:,:2]
mvn_y_spiral = multivariate_test_spiral.iloc[:,-1]
mvn_X_spiral = multivariate_test_spiral.ix[:,:2]
Z = bhc.bhclust(np.array(mvn_X), family = "multivariate", alpha = 1, r = 0.001)[0]
#Z_spiral = bhc.bhclust(np.array(mvn_X_spiral), family = "multivariate", alpha = 1, r = 0.001)[0]
Z_spiral = np.array([[208, 209, 0.25464380105923845, 2],
[312, 210, 0.46200922472289818, 3],
[313, 211, 0.65792654490843439, 4],
[314, 212, 0.85214870893944072, 5],
[315, 213, 1.0474374513171529, 6],
[316, 214, 1.2401314306850928, 7],
[317, 215, 1.4290644232511895, 8],
[318, 216, 1.6167719628504882, 9],
[319, 207, 1.8022719248004808, 10],
[320, 217, 1.9880160854245341, 11],
[321, 218, 2.1718080026801947, 12],
[322, 219, 2.3535387353956598, 13],
[323, 220, 2.5357948210127699, 14],
[324, 221, 2.7183498634980636, 15],
[325, 222, 2.9017376562581312, 16],
[326, 223, 3.0907544545637697, 17],
[327, 224, 3.2835407243264756, 18],
[328, 225, 3.4787545391292647, 19],
[329, 226, 3.6820322935316341, 20],
[330, 227, 3.8956402344230563, 21],
[331, 228, 4.1153078327050796, 22],
[332, 43, 4.3396699231949301, 23],
[333, 44, 4.5486624720656312, 24],
[334, 42, 4.7464192002386101, 25],
[335, 45, 4.9365581738472208, 26],
[336, 41, 5.1199929291513797, 27],
[337, 40, 5.2977606349866289, 28],
[338, 46, 5.4776216603248136, 29],
[339, 39, 5.6549063376429283, 30],
[340, 47, 5.8304496434687749, 31],
[341, 38, 6.0052000241049424, 32],
[342, 37, 6.1827191806557407, 33],
[343, 48, 6.3587118511316882, 34],
[344, 49, 6.5378099770116558, 35],
[345, 36, 6.7185324918892055, 36],
[346, 35, 6.9036638728001432, 37],
[347, 50, 7.0907838886857864, 38],
[348, 229, 7.2801763538373381, 39],
[349, 51, 7.4677772977308505, 40],
[350, 230, 7.6574399478054431, 41],
[351, 52, 7.8473568829173059, 42],
[352, 231, 8.0386787641377513, 43],
[353, 53, 8.2296605046772306, 44],
[354, 232, 8.4222441218560746, 45],
[355, 54, 8.6127260388388311, 46],
[356, 233, 8.8091782774832819, 47],
[357, 55, 9.0042168473094364, 48],
[358, 234, 9.2025562622563744, 49],
[359, 56, 9.3993575041259003, 50],
[360, 235, 9.5972882192286413, 51],
[361, 57, 9.7925316206515998, 52],
[362, 236, 9.9963483747536692, 53],
[363, 58, 10.194643338846785, 54],
[364, 59, 10.403849334855531, 55],
[365, 237, 10.606966238027683, 56],
[366, 60, 10.818012129426682, 57],
[367, 238, 11.022906901719029, 58],
[368, 34, 11.238431919832287, 59],
[369, 61, 11.455127224528686, 60],
[370, 239, 11.666447654540384, 61],
[371, 106, 11.87486178570534, 62],
[372, 240, 12.083548083852753, 63],
[373, 62, 12.286670947503465, 64],
[374, 107, 12.487047629817287, 65],
[375, 241, 12.693687445314367, 66],
[376, 108, 12.895298719628491, 67],
[377, 63, 13.0948951765226, 68],
[378, 109, 13.301127749813435, 69],
[379, 242, 13.50013297263771, 70],
[380, 64, 13.698624125572444, 71],
[381, 243, 13.904419825771392, 72],
[382, 110, 14.10436220748764, 73],
[383, 65, 14.303817818684839, 74],
[384, 111, 14.510206415919098, 75],
[385, 244, 14.711803817291187, 76],
[386, 66, 14.912086865933183, 77],
[387, 184, 15.116000418791536, 78],
[388, 185, 15.318083296125637, 79],
[389, 187, 15.519199656525453, 80],
[390, 186, 15.718407058789058, 81],
[391, 188, 15.915786318715117, 82],
[392, 189, 16.111777858209962, 83],
[393, 183, 16.306838055039179, 84],
[394, 190, 16.500657624436471, 85],
[395, 67, 16.695531436066762, 86],
[396, 191, 16.889371473122157, 87],
[397, 182, 17.082618930413844, 88],
[398, 192, 17.274546172851583, 89],
[399, 181, 17.465607029314985, 90],
[400, 245, 17.65874172242804, 91],
[401, 193, 17.852860792011416, 92],
[402, 194, 18.04556492896484, 93],
[403, 68, 18.237324653586406, 94],
[404, 180, 18.428820995289747, 95],
[405, 179, 18.621090128133329, 96],
[406, 195, 18.812707691326366, 97],
[407, 112, 19.00526495099378, 98],
[408, 196, 19.197344649457349, 99],
[409, 246, 19.38868605556981, 100],
[410, 69, 19.579676522439613, 101],
[411, 113, 19.773221402900422, 102],
[412, 197, 19.966177545304568, 103],
[413, 178, 20.159480686657847, 104],
[414, 70, 20.352871099079472, 105],
[415, 198, 20.545938699533586, 106],
[416, 247, 20.737917909306216, 107],
[417, 114, 20.933474333830262, 108],
[418, 71, 21.128129483817531, 109],
[419, 248, 21.320990525569673, 110],
[420, 199, 21.513335515675845, 111],
[421, 200, 21.708664035004347, 112],
[422, 72, 21.902794639701991, 113],
[423, 177, 22.097313195620178, 114],
[424, 201, 22.293931260843557, 115],
[425, 115, 22.489703191755133, 116],
[426, 249, 22.682138421065023, 117],
[427, 73, 22.87844806818644, 118],
[428, 202, 23.074695353794031, 119],
[429, 176, 23.270679335413632, 120],
[430, 33, 23.468202838845997, 121],
[431, 175, 23.664716556573079, 122],
[432, 203, 23.861430690340345, 123],
[433, 116, 24.059726883673378, 124],
[434, 250, 24.253849795593823, 125],
[435, 74, 24.448553669717572, 126],
[436, 204, 24.643320578970965, 127],
[437, 205, 24.841107548145519, 128],
[438, 174, 25.038993631693437, 129],
[439, 251, 25.237539981043419, 130],
[440, 117, 25.434372015899868, 131],
[441, 75, 25.629974009975534, 132],
[442, 206, 25.825407820894686, 133],
[443, 252, 26.025837308225103, 134],
[444, 76, 26.22411130657791, 135],
[445, 118, 26.423820142184372, 136],
[446, 77, 26.624804788159455, 137],
[447, 253, 26.825100976433188, 138],
[448, 32, 27.029413294965593, 139],
[449, 173, 27.231863915200705, 140],
[450, 78, 27.435912373576297, 141],
[451, 119, 27.639301834192857, 142],
[452, 254, 27.843749041491037, 143],
[453, 79, 28.04876890623898, 144],
[454, 172, 28.254698838791665, 145],
[455, 120, 28.462215434280054, 146],
[456, 80, 28.668550565806434, 147],
[457, 255, 28.873251976708247, 148],
[458, 81, 29.084243575106829, 149],
[459, 121, 29.293657193023151, 150],
[460, 256, 29.499555315834282, 151],
[461, 82, 29.709840100567661, 152],
[462, 171, 29.920206116518838, 153],
[463, 31, 30.130236022217247, 154],
[464, 257, 30.342712960718742, 155],
[465, 122, 30.551278526310849, 156],
[466, 83, 30.760614235670957, 157],
[467, 170, 30.97394087397516, 158],
[468, 258, 31.187073448612239, 159],
[469, 84, 31.39810930479965, 160],
[470, 123, 31.608187581853009, 161],
[471, 259, 31.821204378437194, 162],
[472, 85, 32.032999463374466, 163],
[473, 311, 32.244858629861767, 164],
[474, 310, 32.456648035428216, 165],
[475, 169, 32.667979651988595, 166],
[476, 30, 32.877216235397825, 167],
[477, 309, 33.087066598119783, 168],
[478, 86, 33.296334866674997, 169],
[479, 308, 33.506220396867427, 170],
[480, 87, 33.717009115693848, 171],
[481, 124, 33.926004728779127, 172],
[482, 260, 34.131687862546698, 173],
[483, 307, 34.341429676301622, 174],
[484, 168, 34.550031661046653, 175],
[485, 88, 34.75841162651291, 176],
[486, 306, 34.96694443003247, 177],
[487, 305, 35.175717614553463, 178],
[488, 29, 35.382991317069816, 179],
[489, 167, 35.589483798811209, 180],
[490, 304, 35.795630497772144, 181],
[491, 303, 36.002410706898281, 182],
[492, 89, 36.20884597478468, 183],
[493, 261, 36.413679560955643, 184],
[494, 125, 36.617577360850596, 185],
[495, 90, 36.822155077470342, 186],
[496, 262, 37.026266431662023, 187],
[497, 302, 37.230355717760318, 188],
[498, 91, 37.435007664927845, 189],
[499, 301, 37.63969155305832, 190],
[500, 28, 37.842926961961069, 191],
[501, 166, 38.043661721257003, 192],
[502, 300, 38.245884537773129, 193],
[503, 92, 38.448804390286213, 194],
[504, 299, 38.651182645166479, 195],
[505, 165, 38.853889549393536, 196],
[506, 93, 39.056278370606435, 197],
[507, 263, 39.257675951309302, 198],
[508, 126, 39.456943545411839, 199],
[509, 94, 39.657162775284533, 200],
[510, 95, 39.857214295200905, 201],
[511, 298, 40.056712910678243, 202],
[512, 264, 40.255679068872766, 203],
[513, 96, 40.454349026455674, 204],
[514, 127, 40.652692980440165, 205],
[515, 97, 40.85054749286568, 206],
[516, 265, 41.048104857529459, 207],
[517, 98, 41.24493495196365, 208],
[518, 297, 41.442016669734741, 209],
[519, 99, 41.638527768229039, 210],
[520, 100, 41.834974453255221, 211],
[521, 101, 42.031030636584596, 212],
[522, 296, 42.226799332056068, 213],
[523, 102, 42.422175541388171, 214],
[524, 128, 42.616988634375161, 215],
[525, 266, 42.810087800657335, 216],
[526, 103, 43.003701081458701, 217],
[527, 104, 43.196954505183804, 218],
[528, 105, 43.39018536326202, 219],
[529, 267, 43.583823519102815, 220],
[530, 129, 43.776699989304291, 221],
[531, 295, 43.969825586047527, 222],
[532, 164, 44.162204824221774, 223],
[533, 27, 44.354910583886046, 224],
[534, 294, 44.546706479973089, 225],
[535, 268, 44.739516015186474, 226],
[536, 293, 44.932357063254429, 227],
[537, 163, 45.124500375910586, 228],
[538, 269, 45.317695836200976, 229],
[539, 130, 45.510198390038624, 230],
[540, 292, 45.702581751565084, 231],
[541, 270, 45.894380815650763, 232],
[542, 162, 46.086517407427827, 233],
[543, 26, 46.277921106583918, 234],
[544, 291, 46.468693184072265, 235],
[545, 290, 46.660115028549853, 236],
[546, 271, 46.851242712399461, 237],
[547, 161, 47.042288960561756, 238],
[548, 289, 47.233175296980349, 239],
[549, 25, 47.423684155378766, 240],
[550, 288, 47.613978873870032, 241],
[551, 160, 47.80365108690863, 242],
[552, 272, 47.992845827780371, 243],
[553, 131, 48.181839348784287, 244],
[554, 273, 48.370382206768042, 245],
[555, 287, 48.558455400881861, 246],
[556, 274, 48.746417690467652, 247],
[557, 286, 48.934014474289981, 248],
[558, 275, 49.121212893010608, 249],
[559, 132, 49.308079600236958, 250],
[560, 276, 49.494235053228522, 251],
[561, 285, 49.680141475358589, 252],
[562, 277, 49.865898096414789, 253],
[563, 284, 50.051010903906352, 254],
[564, 159, 50.235532192535473, 255],
[565, 283, 50.419687294787863, 256],
[566, 278, 50.603217545799232, 257],
[567, 282, 50.786361995087354, 258],
[568, 280, 50.968986028803933, 259],
[569, 279, 51.151014330545891, 260],
[570, 281, 51.332584197279814, 261],
[571, 133, 51.513650594986871, 262],
[572, 158, 51.695447058635992, 263],
[573, 24, 51.876798988392814, 264],
[574, 134, 52.05883183935822, 265],
[575, 157, 52.240674452481173, 266],
[576, 156, 52.423192386470177, 267],
[577, 23, 52.604554164984499, 268],
[578, 135, 52.78670822985984, 269],
[579, 155, 52.968823994310441, 270],
[580, 136, 53.151181947673599, 271],
[581, 154, 53.332981843924486, 272],
[582, 22, 53.514271075382979, 273],
[583, 153, 53.695705388191747, 274],
[584, 137, 53.87668769553229, 275],
[585, 152, 54.057427582272808, 276],
[586, 138, 54.237631679908716, 277],
[587, 139, 54.417667541515144, 278],
[588, 151, 54.59703721784976, 279],
[589, 140, 54.775951076160361, 280],
[590, 150, 54.954394694870459, 281],
[591, 149, 55.132110128563099, 282],
[592, 141, 55.308957703757486, 283],
[593, 143, 55.485450704055999, 284],
[594, 142, 55.661006840016022, 285],
[595, 148, 55.835763444513738, 286],
[596, 147, 56.009844122898336, 287],
[597, 144, 56.183147910738306, 288],
[598, 145, 56.355918476527322, 289],
[599, 146, 56.527929600980769, 290],
[600, 21, 56.701145099604936, 291],
[601, 20, 56.874768596683069, 292],
[602, 19, 57.049473911403105, 293],
[603, 18, 57.224426757534133, 294],
[604, 17, 57.399936809324004, 295],
[605, 16, 57.575261094418813, 296],
[606, 15, 57.751331912122289, 297],
[607, 14, 57.92779189099705, 298],
[608, 13, 58.103770505036081, 299],
[609, 12, 58.279802718923897, 300],
[610, 11, 58.455721118208665, 301],
[611, 10, 58.631339581487026, 302],
[612, 9, 58.806495598194736, 303],
[613, 8, 58.981134536182289, 304],
[614, 7, 59.155628417520155, 305],
[615, 6, 59.329478833657468, 306],
[616, 5, 59.502712831224564, 307],
[617, 4, 59.675199527656936, 308],
[618, 3, 59.847384320385984, 309],
[619, 2, 60.018646697963312, 310],
[620, 1, 60.189092488116778, 311],
[621, 0, 60.35935950732884, 312]])
# +
Z = np.array(Z)
single_test = linkage(mvn_X,method='single')
complete_test = linkage(mvn_X,method='complete')
average_test = linkage(mvn_X,method='average')
print("BHC_test:", round(purity_score(Z, mvn_y, 5, 12),3))
print("Single_linkage:", round(purity_score(single_test, mvn_y, 5, 12),3))
print("Complete_linkage:", round(purity_score(complete_test, mvn_y, 5, 12),3))
print("Average_linkage:", round(purity_score(average_test, mvn_y, 5, 12),3))
# +
#Z_spiral = np.array(Z_spiral)
single_test = linkage(mvn_X_spiral,method='single')
complete_test = linkage(mvn_X_spiral,method='complete')
average_test = linkage(mvn_X_spiral,method='average')
print("BHC_test:", round(purity_score(Z_spiral, mvn_y_spiral, 5, 12),3))
print("Single_linkage:", round(purity_score(single_test, mvn_y_spiral, 5, 12),3))
print("Complete_linkage:", round(purity_score(complete_test, mvn_y_spiral, 5, 12),3))
print("Average_linkage:", round(purity_score(average_test, mvn_y_spiral, 5, 12),3))
# -
Z = np.array(Z)
purity_score(linkage_matrix=Z, y_test=mvn_y, class_test=3, repeats=5, seed=16)
Z_spiral = np.array(Z_spiral)
purity_score(linkage_matrix=Z_spiral, y_test=mvn_y_spiral, class_test=3, repeats=5, seed=16)
# ### Test with binary
# +
#SYNTHETIC binary data
X_test = Bern_gen(30, 10, 0.5, 121)
y_test = []
for i in ['A','B','C']:
y_test.extend(np.repeat(i,10))
Zb = bhc.bhclust(X_test, family = "bernoulli", alpha = 0.001)[0]
Zb = np.array(Zb)
single_test = linkage(X_test,method='single')
complete_test = linkage(X_test,method='complete')
average_test = linkage(X_test,method='average')
print("BHC_test:", round(purity_score(Zb, y_test, 5, 12),3))
print("Single_linkage:", round(purity_score(single_test, y_test, 5, 12),3))
print("Complete_linkage:", round(purity_score(complete_test, y_test, 5, 12),3))
print("Average_linkage:", round(purity_score(average_test, y_test, 5, 12),3))
# -
#CEDA data from paper
multivariate_test = pd.read_csv("/Users/lina/Downloads/bindat.csv", header=-1)
bn_y = list(np.repeat('0',40))
bn_y.extend(list(np.repeat('2',40)))
bn_y.extend(list(np.repeat('4',40)))
bn_X = np.array(multivariate_test)
# +
Zb_paper = bhc.bhclust(bn_X, family = "bernoulli", alpha = 0.001)[0]
Zb_paper = np.array(Zb_paper)
#BHC_test = np.array(bhc.bhclust_BB(bn_X)[0])
single_test = linkage(bn_X,method='single')
complete_test = linkage(bn_X,method='complete')
average_test = linkage(bn_X,method='average')
print("BHC_test:", round(purity_score(Zb_paper, bn_y, 5, 12),3))
print("Single_linkage:", round(purity_score(single_test, bn_y, 5, 12),3))
print("Complete_linkage:", round(purity_score(complete_test, bn_y, 5, 12),3))
print("Average_linkage:", round(purity_score(average_test, bn_y, 5, 12),3))
# -
| tests/comparison_tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# ## **Scikit** (**Sci**Py Tool**kit**)-learn, SK-Learn
# dependency
# - NumPy
# - SciPy
# - Matplotlib
# ## Linear Regression Concept
# +
from random import *
from sklearn import linear_model
import matplotlib.pyplot as plt
length = 10
x = [[i] for i in range(length)]
y = [[random() * 10] for i in range(length)]
print("x: \n", x)
print("y: \n", y)
regr = linear_model.LinearRegression()
regr.fit(x, y)
plt.scatter(x, y, color='black')
plt.plot(x, regr.predict(x), color='blue', linewidth=3)
# +
import numpy as np
length = 10
x = np.arange(length).reshape(length, 1)
y = np.array([[random() * 10] for i in range(length)]).reshape(length, 1)
print("x: \n", x)
print("y: \n", y)
regr = linear_model.LinearRegression()
regr.fit(x, y)
plt.scatter(x, y, color='black')
plt.plot(x, regr.predict(x), color='blue', linewidth=3)
# +
from random import *
from sklearn import linear_model
import matplotlib.pyplot as plt
import pandas as pd
xy_data = [[i, random() * 10] for i in range(length)]
df = pd.DataFrame(xy_data, columns=('X', 'Y'))
regr = linear_model.LinearRegression()
regr.fit(df.loc[:, ['X']], df.loc[:, ['Y']])
plt.scatter(df.loc[:, ['X']], df.loc[:, ['Y']], color='black')
plt.plot(df.loc[:, ['X']], regr.predict(df.loc[:, ['X']]), color='blue', linewidth=3)
# -
df.loc[:, ['X']]
# +
from sklearn import datasets, linear_model
diabetes = datasets.load_diabetes()
df = pd.DataFrame(diabetes['data'], columns=diabetes.feature_names)
df.head()
# -
x_df = df.loc[:,['bmi']]
x_train = x_df.iloc[:-20]
x_test = x_df.iloc[-20:]
y_train = diabetes.target[:-20]
y_test = diabetes.target[-20:]
regr.fit(x_train, y_train)
pred = regr.predict(x_test)
print('Mean squared error: %.2f'%np.mean((pred - y_test)**2))
print('Variance score : %.2f'%regr.score(x_test, y_test))
plt.scatter(x_test, y_test, color='black')
plt.plot(x_test, pred, color='blue')
# ## K-Nearest Neighbors (KNN) Classifier
# +
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
n = 15
iris = datasets.load_iris()
x = iris.data[:, :2]
y = iris.target
h = 0.2
# +
df = pd.DataFrame(iris.data, columns=iris.feature_names)
df['Class'] = iris.target
for cls, group in df.groupby('Class'):
plt.scatter(group['sepal length (cm)'], group['sepal width (cm)'], label=cls)
plt.xlabel('sepal length (cm)')
plt.ylabel('sepal width (cm)')
plt.legend()
# +
from sklearn import neighbors
clf = neighbors.KNeighborsClassifier(n, weights='uniform')
clf.fit(x,y)
new_corner = np.array([[3.7, 4.5]])
iris_class = clf.predict(new_corner)
print("the iris_class for new_point : ", iris_class)
# -
# +
## Decision Tree
# -
# !pip install -U scikit-learn
import sklearn
sklearn.__version__
# !top
# ## Neural-Net for MNIST
# +
# %matplotlib inline
import matplotlib.pyplot as plt
from sklearn import datasets
from sklearn.datasets import fetch_mldata
from sklearn.neural_network import MLPClassifier
mnist_data = datasets.fetch_openml('mnist_784', cache=True)
mnist_data
# -
X, y = mnist_data.data / 255., mnist_data.target
n = 60000
X_train, X_test = X[:n], X[n:]
y_train, y_test = y[:n], y[n:]
mlp = MLPClassifier(hidden_layer_sizes=(50,), max_iter=10, alpha=1e-4,solver='sgd',
verbose=10, tol=1e-4, random_state=1, learning_rate_init=.1)
mlp.fit(X_train, y_train)
print('Traning set score:', mlp.score(X_train, y_train))
print('Test set score:', mlp.score(X_test, y_test))
import sys
sys.getrecursionlimit()
sys.maxsize
ulimit -v
# +
data = rawdata
test_data = ddd
class_zero = rawdata[rawdata['Class']]
# 원본 데이터 사용, 테스트 셋을 1:1 추출
data = pd.concat([class_zero, class_one])
train_data = data.iloc[100:-100]
test_data = pd.concat([data.iloc[:100], data.iloc[-100:]])
train_data = train_data.sample(frac=1)
test_data = test_data.sample(frac=1) # 순서에 의한 데이터 연관성 제거. 순서 섞기
# 언더샘플링
model = lm.LogisticRegression()
model = svm.SVC(gamma='scale')
model = nbr.KNeighborsClassifier(n_neighbors=3)
model = tree.DecisionTreeClassifier()
model = nn.MLPClassifier(learning_rate_init=0.01)
model.fit()
| _ipynbs/study-py-sklearn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # APTOS 2019 Blindness Detection
# +
from pathlib import Path
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import cv2
from imgaug import augmenters as iaa
from PIL import Image
# -
# ## White Noise
class WhiteNoise(object):
def __init__(self, prob, scale):
self.prob = prob
self.seq = iaa.Sequential([
iaa.AdditiveGaussianNoise(scale)
])
def __call__(self, img):
rand = np.random.rand()
if rand > self.prob:
arr = np.array(img)
arr = self.seq.augment_image(arr)
img = Image.fromarray(arr)
return img
# ## RandomEraser
class RandomEraser(object):
"""
"""
def __init__(self, prob, size_range=(0.02, 0.4), ratio_range=(0.3, 3)):
self.prob = prob
self.s_range = size_range # TODO error case
self.r_range = ratio_range
def __call__(self, img):
rand = np.random.rand()
if rand > self.prob:
arr = np.array(img)
h, w, _ = arr.shape
mask_area = np.random.randint(h * w * self.s_range[0], h * w * self.s_range[1])
mask_aspect_ratio = np.random.rand() * (self.r_range[1] - self.r_range[0]) + self.r_range[0]
mask_height = int(np.sqrt(mask_area / mask_aspect_ratio))
if mask_height > h - 1:
mask_height = h - 1
mask_width = int(mask_aspect_ratio * mask_height)
if mask_width > w - 1:
mask_width = w - 1
top = np.random.randint(0, h - mask_height)
left = np.random.randint(0, w - mask_width)
bottom = top + mask_height
right = left + mask_width
arr[top:bottom, left:right] = np.random.randint(low=0, high=256, size=(mask_height, mask_width, 3))
img = Image.fromarray(arr)
return img
# ## Cutout
# ## RandomSkewness
class RandomSkewness(object):
"""
"""
def __init__(self, prob, ratio=1.0):
self.prob = prob
self.ratio = ratio
def __call__(self, img):
rand = np.random.rand()
if rand < self.prob:
arr = np.array(img)
h, w, _ = arr.shape
c = (int(w / 2), int(h / 2))
src_pts = np.array([[c[0], c[1]], [c[0] + 100, 0], [0, c[1] + 100]], dtype=np.float32)
dst_pts = np.array([[c[0], c[1]], [c[0] + 100, 0], [0, c[1] + int(self.ratio * 100)]], dtype=np.float32)
mat = cv2.getAffineTransform(src_pts, dst_pts)
print(mat)
img = cv2.warpAffine(img, mat, (w, h))
return img
# ## MixUp
# ## RICAP
# ## Check
train_df = pd.read_csv("../input/train.csv")
def load_image(id):
img_path = Path().absolute().parent / "input" / "train_images" / "{}.png".format(id)
if img_path.exists():
d_level = int(train_df.query("id_code == '{}'".format(id)).iloc[0, 1])
diagnosis_dict = {0: "No DR", 1: "Mild", 2: "Moderate", 3: "Severe", 4: "Proliferative DR"}
else:
img_path = Path().absolute().parent / "input" / "test_images" / "{}.png".format(id)
return cv2.imread(str(img_path))
# augment = RandomEraser(prob=0)
augment = RandomSkewness(prob=1, ratio=1.2)
f_list = ["005b95c28852", "00cb6555d108", "01499815e469", "0167076e7089"]
for file_name in f_list:
img = load_image(file_name)
img = Image.fromarray(img)
plt.figure(figsize=(5, 5))
plt.imshow(img)
img = load_image(file_name)
img = Image.fromarray(img)
img = augment(img)
plt.figure(figsize=(5, 5))
plt.imshow(img)
| notebook/[Trial]_08_Augmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.017271, "end_time": "2021-10-19T19:41:23.917798", "exception": false, "start_time": "2021-10-19T19:41:23.900527", "status": "completed"} tags=[]
# # COVID-19 India Matplotlib Overview
#
# + papermill={"duration": 1.142918, "end_time": "2021-10-19T19:41:25.073007", "exception": false, "start_time": "2021-10-19T19:41:23.930089", "status": "completed"} tags=[]
#hide
import pandas as pd
import numpy as np
import requests
import json
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
import matplotlib as mpl
from IPython.core.display import display,HTML
# %matplotlib inline
# + papermill={"duration": 0.051922, "end_time": "2021-10-19T19:41:25.140369", "exception": false, "start_time": "2021-10-19T19:41:25.088447", "status": "completed"} tags=[]
#hide
dft_cases = pd.read_csv('data/SnapshotCases-28-July.csv')
dft_deaths = pd.read_csv('data/SnapshotDeaths-28-July.csv')
# + papermill={"duration": 0.045397, "end_time": "2021-10-19T19:41:25.198135", "exception": false, "start_time": "2021-10-19T19:41:25.152738", "status": "completed"} tags=[]
#hide
dft_cases.head()
# + papermill={"duration": 0.020369, "end_time": "2021-10-19T19:41:25.232263", "exception": false, "start_time": "2021-10-19T19:41:25.211894", "status": "completed"} tags=[]
#hide
dt_yday = '27-Jul-20'
dt_today = '28-Jul-20'
# + papermill={"duration": 0.02803, "end_time": "2021-10-19T19:41:25.272325", "exception": false, "start_time": "2021-10-19T19:41:25.244295", "status": "completed"} tags=[]
#hide
dfc_cases = dft_cases.groupby('states')[dt_today].sum()
dfc_deaths = dft_deaths.groupby('states')[dt_today].sum()
dfp_cases = dft_cases.groupby('states')[dt_yday].sum()
dfp_deaths = dft_deaths.groupby('states')[dt_yday].sum()
# + papermill={"duration": 0.032463, "end_time": "2021-10-19T19:41:25.320359", "exception": false, "start_time": "2021-10-19T19:41:25.287896", "status": "completed"} tags=[]
#hide
df_table = pd.DataFrame({'states': dfc_cases.index, 'Cases': dfc_cases.values, 'Deaths': dfc_deaths.values, 'PCases': dfp_cases.values, 'PDeaths': dfp_deaths.values}).set_index('states')
df_table = df_table.sort_values(by = ['Cases','Deaths'], ascending = [False, False])
df_table = df_table.reset_index()
df_table.head()
# + papermill={"duration": 0.028491, "end_time": "2021-10-19T19:41:25.362460", "exception": false, "start_time": "2021-10-19T19:41:25.333969", "status": "completed"} tags=[]
#hide
for c in 'Cases, Deaths'.split(', '):
df_table[f'{c} (+)'] = (df_table[c] - df_table[f'P{c}']).clip(0)
df_table['Fatality Rate'] = (100* df_table['Deaths']/ df_table['Cases']).round(2)
# + papermill={"duration": 0.029149, "end_time": "2021-10-19T19:41:25.406507", "exception": false, "start_time": "2021-10-19T19:41:25.377358", "status": "completed"} tags=[]
#hide
df_table.head()
# + papermill={"duration": 0.035902, "end_time": "2021-10-19T19:41:25.456650", "exception": false, "start_time": "2021-10-19T19:41:25.420748", "status": "completed"} tags=[]
#hide
summary = {'updated':'28th July, 2020', 'since':'27th July, 2020'}
list_names = ['Cases', 'PCases', 'Deaths', 'PDeaths', 'Cases (+)', 'Deaths (+)']
for name in list_names:
summary[name] = df_table.sum()[name]
summary
# + papermill={"duration": 0.023749, "end_time": "2021-10-19T19:41:25.494563", "exception": false, "start_time": "2021-10-19T19:41:25.470814", "status": "completed"} tags=[]
#hide
overview = '''
<!-- ####### HTML!! #########-->
<h1 style="color: #5e9ca0; text-align: center;">India</h1>
<p style="text-align: center;">Last update: <strong>{update}</strong></p>
<p style="text-align: center;">Confirmed cases:</p>
<p style="text-align: center;font-size:24px;">{cases} (<span style="color: #ff0000;">+{new}</span>)</p>
<p style="text-align: center;">Confirmed deaths:</p>
<p style="text-align: center;font-size:24px;">{deaths} (<span style="color: #ff0000;">+{dnew}</span>)</p>
'''
# + papermill={"duration": 0.023142, "end_time": "2021-10-19T19:41:25.532267", "exception": false, "start_time": "2021-10-19T19:41:25.509125", "status": "completed"} tags=[]
#hide_input
update = summary['updated']
cases = summary['Cases']
new = summary['Cases (+)']
deaths = summary['Deaths']
dnew = summary['Deaths (+)']
html = HTML(overview.format(update=update, cases=cases,new=new,deaths=deaths,dnew=dnew))
display(html)
# + papermill={"duration": 0.027783, "end_time": "2021-10-19T19:41:25.573582", "exception": false, "start_time": "2021-10-19T19:41:25.545799", "status": "completed"} tags=[]
#hide
dt_cols = list(dft_cases.columns[1:])
dft_ct_new_cases = dft_cases.groupby('states')[dt_cols].sum().diff(axis=1).fillna(0).astype(int)
dft_ct_new_cases.sort_values(by = '28-Jul-20', ascending = False,inplace = True)
# + papermill={"duration": 0.036366, "end_time": "2021-10-19T19:41:25.628225", "exception": false, "start_time": "2021-10-19T19:41:25.591859", "status": "completed"} tags=[]
#hide
dft_ct_new_cases.head()
# + papermill={"duration": 5.523478, "end_time": "2021-10-19T19:41:31.167883", "exception": false, "start_time": "2021-10-19T19:41:25.644405", "status": "completed"} tags=[]
#hide_input
df = dft_ct_new_cases.copy()
df.loc['Total'] = df.sum()
n = 5
ax = []
fig = plt.figure(figsize = (16,20))
gs = fig.add_gridspec(n+2, 3)
# gs = fig.add_gridspec(2, 3)
ax1 = fig.add_subplot(gs[0, :])
ef = df.loc['Total'].rename_axis('date').reset_index()
ef['date'] = ef['date'].astype('datetime64[ns]')
ax1.bar(ef.date,ef.Total,alpha=0.3,color='#007acc')
ax1.plot(ef.date,ef.Total , marker="o", color='#007acc')
ax1.xaxis.set_major_locator(mdates.WeekdayLocator())
ax1.xaxis.set_major_formatter(mdates.DateFormatter('%b %d'))
ax1.text(0.02, 0.5,'India daily case count', transform = ax1.transAxes, fontsize=25);
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
ax2 = fig.add_subplot(gs[1,0])
ef = df.loc['Maharashtra'].rename_axis('date').reset_index()
ef['date'] = ef['date'].astype('datetime64[ns]')
ax2.bar(ef.date, ef.Maharashtra,color = '#007acc',alpha=0.5)
ax2.xaxis.set_major_locator(mdates.WeekdayLocator())
ax2.xaxis.set_major_formatter(mdates.DateFormatter('%b %d'))
ax2.set_xticks(ax2.get_xticks()[::3])
maxyval = ef.Maharashtra.max()
ax2.set_ylim([0,maxyval])
ax2.text(0.05, 0.5,'Maharashtra', transform = ax2.transAxes, fontsize=20);
ax2.spines['right'].set_visible(False)
ax2.spines['top'].set_visible(False)
ax3 = fig.add_subplot(gs[1,1])
ef = df.loc['Tamil Nadu'].rename_axis('date').reset_index()
ef['date'] = ef['date'].astype('datetime64[ns]')
ax3.bar(ef.date, ef['Tamil Nadu'],color = '#007acc',alpha=0.5,)
ax3.xaxis.set_major_locator(mdates.WeekdayLocator())
ax3.xaxis.set_major_formatter(mdates.DateFormatter('%b %d'))
ax3.set_xticks(ax3.get_xticks()[::3])
ax3.text(0.05, 0.5,'Tamil Nadu', transform = ax3.transAxes, fontsize=20);
ax3.spines['right'].set_visible(False)
ax3.spines['top'].set_visible(False)
ax4 = fig.add_subplot(gs[1,2])
ef = df.loc['Delhi'].rename_axis('date').reset_index()
ef['date'] = ef['date'].astype('datetime64[ns]')
ax4.bar(ef.date, ef.Delhi,color = '#007acc',alpha=0.5)
ax4.set_xticks([])
ax4.xaxis.set_major_locator(mdates.WeekdayLocator())
ax4.xaxis.set_major_formatter(mdates.DateFormatter('%b %d'))
ax4.set_xticks(ax4.get_xticks()[::3])
ax4.spines['right'].set_visible(False)
ax4.spines['top'].set_visible(False)
ax4.text(0.05, 0.5,'Delhi', transform = ax4.transAxes, fontsize=20)
for i in range(n):
ax.append(fig.add_subplot(gs[i+2,:]))
ef = df.iloc[i+3].rename_axis('date').reset_index()
ef['date'] = ef['date'].astype('datetime64[ns]')
ax[i].bar(ef.date,ef.iloc[:,-1],color = '#007acc',alpha=0.3)
ax[i].plot(ef.date,ef.iloc[:,-1],marker='o',color='#007acc')
ax[i].text(0.02,0.5,f'{ef.columns.values[-1]}',transform = ax[i].transAxes, fontsize = 20);
ax[i].xaxis.set_major_locator(mdates.WeekdayLocator())
ax[i].xaxis.set_major_formatter(mdates.DateFormatter('%b %d'))
ax[i].set_ylim([0,7000])
ax[i].spines['right'].set_visible(False)
ax[i].spines['top'].set_visible(False)
plt.tight_layout()
# + papermill={"duration": 0.032416, "end_time": "2021-10-19T19:41:31.222310", "exception": false, "start_time": "2021-10-19T19:41:31.189894", "status": "completed"} tags=[]
#hide_input
print(df_table.to_string(index=False))
| _notebooks/2020-07-28-covid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.12 64-bit (''attendance'': conda)'
# language: python
# name: python3
# ---
# +
# pip install dlib
# +
# pip install cmake
# +
# pip install face_recognition
# +
# pip install opencv-python
# -
import face_recognition
import cv2
imgsrc = 'images/'
img = cv2.imread(imgsrc + 'louisrossi.png')
rgb_img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
img_encoding = face_recognition.face_encodings(rgb_img)[0]
img2 = cv2.imread(imgsrc + 'messi.png')
rgb_img2 = cv2.cvtColor(img2, cv2.COLOR_BGR2RGB)
img_encoding2 = face_recognition.face_encodings(rgb_img2)[0]
img3 = cv2.imread(imgsrc + 'ryanreynolds.png')
rgb_img3 = cv2.cvtColor(img3, cv2.COLOR_BGR2RGB)
img_encoding3 = face_recognition.face_encodings(rgb_img3)[0]
img4 = cv2.imread(imgsrc + 'elonmusk.png')
rgb_img4 = cv2.cvtColor(img4, cv2.COLOR_BGR2RGB)
img_encoding4 = face_recognition.face_encodings(rgb_img4)[0]
img5 = cv2.imread(imgsrc + 'jeffbezos.png')
rgb_img5 = cv2.cvtColor(img5, cv2.COLOR_BGR2RGB)
img_encoding5 = face_recognition.face_encodings(rgb_img5)[0]
# comparison louis/messi
result = face_recognition.compare_faces([img_encoding], img_encoding2)
print("Result: ", result)
# comparison jeffbezos/ryanreynolds
result = face_recognition.compare_faces([img_encoding5], img_encoding3)
print("Result: ", result)
messitestimg = cv2.imread('messi2.png')
rgb_messitestimg = cv2.cvtColor(messitestimg, cv2.COLOR_BGR2RGB)
messi_encoding = face_recognition.face_encodings(rgb_messitestimg)[0]
# comparison messi with messi
result = face_recognition.compare_faces([img_encoding2], messi_encoding)
print("Result: ", result)
cv2.imshow('image', img)
cv2.imshow('image2', img2)
cv2.waitKey(0)
| Attendance/facerecognition/nov221115.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Making maps
# 
# + [markdown] slideshow={"slide_type": "fragment"}
# We're going to get some shapefiles from data.gov.au, and then we're going to draw some maps
# + [markdown] slideshow={"slide_type": "slide"}
# You'll need some prequisite libraries:
# ```
# pip install pandas shapely fiona descartes pyproj geopandas
# ```
# You may have some already but pip will be cool with that
# + [markdown] slideshow={"slide_type": "slide"}
# We're going to get a shapefile of suburb boundaries from here:
#
# https://data.gov.au/dataset/ds-dga-91e70237-d9d1-4719-a82f-e71b811154c6/details
# +
import os
import geopandas as gp
import shapely
import fiona
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
shp_file_name = "NSW_LOCALITY_POLYGON_shp/NSW_LOCALITY_POLYGON_shp.shp"
zip_file_name = "nsw_locality_polygon_shp.zip"
web_file_path = ("https://data.gov.au/dataset/91e70237-d9d1-4719-a82f-e71b811154c6/resource/"
"5e295412-357c-49a2-98d5-6caf099c2339/download/nsw_locality_polygon_shp.zip")
# + slideshow={"slide_type": "skip"}
# %matplotlib inline
plt.rcParams['figure.figsize'] = (20, 10)
# + [markdown] slideshow={"slide_type": "slide"}
# To get a load of shape files from data.gov.au, more specifically, [here](https://data.gov.au/dataset/nsw-suburb-locality-boundaries-psma-administrative-boundaries/resource/bf8b5180-fcea-44bd-bd76-af14cc4b0fe0)
# -
def unzip_zipfile(zipped_file_path, put_it_here="."):
import zipfile
zip_of_suburbs = zipfile.ZipFile(zipped_file_path, 'r')
zip_of_suburbs.extractall(put_it_here)
zip_of_suburbs.close()
# +
# Get the data loaded. This is a bit complicated because it's online as a zip file.
# If we don't have it right now, we need to get it and unzip it.
if os.path.isfile(shp_file_name):
print("loading from file")
else:
if os.path.isfile(zip_file_name):
print("unzipping")
unzip_zipfile(zip_file_name)
else:
import requests
print("loading from the internet")
page = requests.get(web_file_path)
with open(zip_file_name, 'wb') as z:
z.write(page.content)
unzip_zipfile(zip_file_name)
print("done")
# + [markdown] slideshow={"slide_type": "slide"}
# #### Shapefiles are the format that a lot of GIS tools use.
#
# #### GIS means Geographic Information System
#
# pronounced: _gee eye ess_
#
# not pronounced: _jizz_
# + slideshow={"slide_type": "slide"}
burbs = gp.GeoDataFrame.from_file(shp_file_name)
burbs.drop(["NSW_LOCA_1", "NSW_LOCA_3", "NSW_LOCA_4", "DT_RETIRE"], axis=1, inplace=True)
burbs.head(2)
# + slideshow={"slide_type": "fragment"}
# burbs.geometry.plot()
# could plot, if you had a really fast computer!
# + slideshow={"slide_type": "slide"}
a = burbs.iloc[0]
print(a)
a.geometry
# + slideshow={"slide_type": "slide"}
def add_centroid(row):
return row.geometry.centroid
burbs["centroid"] = burbs.apply(add_centroid, axis=1)
# + slideshow={"slide_type": "fragment"}
a = burbs.iloc[0]
print(a.centroid)
a.centroid
# + slideshow={"slide_type": "fragment"}
right_here = shapely.geometry.point.Point(151.2299732, -33.9178754)
burbs["distance_from_UNSW"] = burbs.geometry.distance(right_here)
# + slideshow={"slide_type": "slide"}
burbs.distance_from_UNSW.hist(bins=50);
# -
# This gives distance in whole numbers of lat long I think, i.e. degrees!
# + [markdown] slideshow={"slide_type": "slide"}
# Not really sure what to do with that, but let's get rid of everything above 0.2 and see what we can plot:
# -
close_burbs = burbs[burbs.distance_from_UNSW<0.2]
close_burbs.plot();
# + slideshow={"slide_type": "slide"}
close_burbs.geometry.convex_hull.plot();
# + slideshow={"slide_type": "slide"}
close_burbs.geometry.envelope.plot();
# + [markdown] slideshow={"slide_type": "slide"}
# 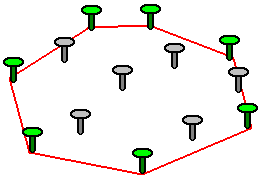
# Convex hulls are a useful aproximation if you want to do fast calculations.
#
# Bounding boxes are even cheaper.
# + [markdown] slideshow={"slide_type": "slide"}
# What about putting labels on the map?
# +
really_close_burbs = burbs[burbs.distance_from_UNSW<0.03]
really_close_burbs.plot()
for idx, row in really_close_burbs.iterrows():
plt.annotate(text=row.NSW_LOCA_2,
xy=tuple(row.centroid.coords)[0],
horizontalalignment='center')
plt.title("Some pretty dodgy suburb names on a map");
# + [markdown] slideshow={"slide_type": "slide"}
# We often want to be able to tell if a point is in a suburb or not. We'd do that with a polygon inclusion test.
#
# 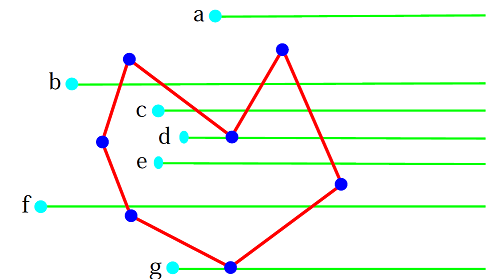
# [How to check if a given point lies inside or outside a polygon?](http://www.geeksforgeeks.org/how-to-check-if-a-given-point-lies-inside-a-polygon/)
# + [markdown] slideshow={"slide_type": "slide"}
# Luckily we've got one built in!
# + slideshow={"slide_type": "-"}
print(right_here)
in_this_burb = None
for _, row in really_close_burbs.iterrows():
if right_here.within(row.geometry):
in_this_burb = row
in_this_burb
# + [markdown] slideshow={"slide_type": "slide"}
# This might actually be a case where a `lambda` is a good idea:
# + slideshow={"slide_type": "-"}
in_this_burb = really_close_burbs[really_close_burbs.apply(lambda x: right_here.within(x.geometry) , axis=1)]
in_this_burb
# + [markdown] slideshow={"slide_type": "slide"}
# How about colouring suburbs according to some kind of scalar value?
#
# [That's called a _chloropleth_](http://geopandas.readthedocs.io/en/latest/mapping.html?highlight=color#chloropleth-maps)
# -
really_close_burbs.plot(column='distance_from_UNSW', cmap='cool');
# I don't think that could be any easier!
#
# You can see the list of [colour scale options here](http://matplotlib.org/users/colormaps.html)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Handy links
#
# * [Geopandas docs](http://geopandas.readthedocs.io)
# * [Geopandas on GitHib](https://github.com/geopandas/geopandas)
# * [Another map making tutorial](http://sensitivecities.com/so-youd-like-to-make-a-map-using-python-EN.html)
# * [MatPlotLib colour scale options](http://matplotlib.org/users/colormaps.html)
#
# I'd love to hear if you come across any other useful things!
# -
| maps/maps.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reading ECMWF data in gribs format
#
# ## Pre-requisites
# - CDS Client is installed via conda and user id and key are valid
# - xarray, eccodes, and cfgrib are installed in conda env
# ## Retrieve data from ECMWF MARS archive
# +
import cdsapi
c = cdsapi.Client()
# +
import cdsapi
c = cdsapi.Client()
c.retrieve(
'reanalysis-era5-single-levels',
{
'product_type': 'reanalysis',
'format': 'grib',
'year': '2019',
'day': [
'01', '02', '03',
'04', '05', '06',
'07', '08', '09',
'10', '11', '12',
'13', '14', '15',
'16', '17', '18',
'19', '20', '21',
'22', '23', '24',
'25', '26', '27',
'28', '29', '30',
'31',
],
'month': [
'05', '06',
],
'time': '13:00',
'variable': '2m_temperature',
},
'download.grib')
# -
# # Read downloaded file
import xarray as xr
import matplotlib.pyplot as plt
ds = xr.open_dataset('download.grib', engine='cfgrib')
ds
ds.t2m.isel(time=0).plot()
# +
# xcube imports
from xcube.core.compute import compute_cube
from xcube.core.maskset import MaskSet
from xcube.core.geom import mask_dataset_by_geometry
from xcube.core.geom import clip_dataset_by_geometry
# -
aoi = dict(type='Polygon', coordinates=[[[0.,30.],[0.,90.],[60.,90],[60.,0.],[0.,30]]] )
masked_ds = mask_dataset_by_geometry(ds,aoi)
masked_ds
masked_ds.to_zarr('t2m_v1.zarr')
| examples/notebooks/read_grib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # My own deep learning framework
# *NB : This notebook is initially a post from my [personal blog](http://colasdroin.eu/DataScienceBlog/).*
# 
# In the context of a course project during my PhD, I had to code my very own deep learning framework in pure Python. We were supposed to keep things simple and implement different modules (e.g. ReLU, Tanh etc.) linked sequentially, each with its own forward and backward pass. But I decided to spice things up a little and consider the whole network as a computational graph, using the [approach already used in TensorFlow](https://www.tensorflow.org/versions/r1.3/get_started/get_started#the_computational_graph). If you're not familiar with the basics of deep learning, before reading what's coming next, I heavily encourage you to have a look at the amazing [3Blue1Brown video tutorials on deep learning](https://www.youtube.com/watch?v=aircAruvnKk&list=PLZHQObOWTQDNU6R1_67000Dx_ZCJB-3pi).
# Before anything, let's import some basic packages for computation and plotting, to make our life easier later.
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('ggplot')
# ## The Node class
# The first thing we're going to do is to define a `Node`, which can be interpret as a layer of the network. It can be either a layer of parameters, either a given transformation (function like SoftMax or Tanh), either simply the input. Parameters nodes are given a value when instancied, while input and functions do not. The value of the input is given when computing the forward pass, and the value returned by each function in the graph is computed iteratively during the forward pass.
class Node:
"""Define a node (input, operation (various types), or parameter) of the
network.
"""
def __init__(self, value = None, type = None, parents = []):
""" Constructor:
-value is a ndarray used to assign a value to
the current node.
-type is a string specifying the type of the node: 'input' as input,
'par' as parameter, and 'add'/'dot' as operations.
-parents is a list of nodes corresponding to the parents of the
current node in the graph.
"""
if type is None:
if value is None:
self.type = 'input'
else:
self.type = 'par'
else:
self.type = type
if type == 'par' and value is None:
raise ValueError('A parameter node must have a value')
if type == 'input' and value is not None:
raise ValueError("An input node can't have a value")
self.value = value
self.parents = parents
self.children = set()
self.dic_op = {'add' : self.compute_add,
'dot' : self.compute_dot,
'relu' : self.compute_relu,
'tanh' : self.compute_tanh}
self.dic_d_op = {'add' : self.compute_d_add,
'dot' : self.compute_d_dot,
'relu' : self.compute_d_relu,
'tanh' : self.compute_d_tanh}
self.grad = 0
self.grad_computed = False
# You can see that each node also come with several other attributes:
# * `parents` is a list containing pointers toward the parent nodes.
# * `children` is a set containing pointers toward the children nodes.
# * `dic_op` is used only for function nodes, in which case it maps the name of the function to the corresponding implementation.
# * `dic_d_op` is exactly the same thing as `dic_op`, except that it is used for the backward pass, therefore using the gradient of the functions.
# * `grad` is a float accumulating the gradient during the backward pass.
# * `grad_computed` is a boolean used to ensure that one doesn't compute twice the gradient of a given node during the backward pass.
#
# We can then define all the common functions used in deep learning networks, as well as their gradient. Note that we don't implement any convolution here (it could be possible, but the gradient computation is a bit trickier).
class Node(Node):
def add(self, node):
"""Function to create a new node whose value is the addition of the two
parents. Self node must be a parameter, 'node' node can be of any
type.
"""
if node.type=='par':
raise ValueError("Self node must be a parameter")
child = Node(type = 'add', parents = [self,node])
self.children.add(child)
node.children.add(child)
return child
def compute_add(self):
"""Function to compute the addition of the values of the two parents.
"""
return np.add(*[parent.value for parent in self.parents])
def compute_d_add(self, grad):
"""Function to compute the derivative of the addition of the values of
the two parents.
"""
return 1*grad
def dot(self, node):
"""Function to create a new node which is the dot product of the values
of the two parents.Self node must be a parameter, 'node' node can be
of any type.
"""
if node.type=='par':
raise ValueError("Self node must be a parameter")
child = Node(type = 'dot', parents = [self,node])
self.children.add(child)
node.children.add(child)
return child
def compute_dot(self):
"""Function to compute the dot product between the parent 1 (w) and
the parent 2 (x).
"""
return np.dot(self.parents[0].value,self.parents[1].value.T).T
def compute_d_dot(self, grad):
"""Function to compute the derivative of the dot product between
parent 1 (w) and parent 2 (x).
"""
return np.dot(self.parents[0].value.T, grad.T ).T
def relu(self):
"""Function to create a new node which is relu of the parent node value.
"""
child = Node(type = 'relu', parents = [self])
self.children.add(child)
return child
def compute_relu(self):
"""Function to compute the value of the current node using relu function
on the parent node.
"""
if len(self.parents)>1:
raise ValueError('reLu takes only a single parent as input')
return self.parents[0].value * (self.parents[0].value > 0)
def compute_d_relu(self, grad):
"""Function to compute the gradient of the current node using relu
function on the parent node.
"""
if len(self.parents)>1:
raise ValueError('reLu takes only a single parent as input')
return (1 * (self.parents[0].value > 0))*grad
def tanh(self):
"""Function to create a new node which is tanh of the parent node value.
"""
child = Node(type = 'tanh', parents = [self])
self.children.add(child)
return child
def compute_tanh(self):
"""Function to compute the value of the current node using tanh function
on the parent node.
"""
if len(self.parents)>1:
raise ValueError('tanh takes only a single parent as input')
return np.tanh(self.parents[0].value )
def compute_d_tanh(self, grad):
"""Function to compute the gradient of the current node using tanh
function on the parent node.
"""
if len(self.parents)>1:
raise ValueError('tanh takes only a single parent as input')
return (1-np.tanh(self.parents[0].value)**2)*grad
# Now that we have all the functions implemented, we can implement the forward pass. Starting from the end node, the principle is simply to compute recurrently the value of all nodes, by using the value of their parent node(s) as input(s) to the function of the current node.
class Node(Node):
def compute_forward(self):
"""Compute the values of the complete graph in a forward manner.
"""
def assert_parents(parents):
for parent in parents:
if parent.value is None:
return False
return True
def compute_nodes_value(current_node):
for parent in current_node.parents:
compute_nodes_value(parent)
if current_node.type!='par' and current_node.type!='input':
if assert_parents(current_node.parents):
current_node.value = current_node.dic_op[current_node.type]()
else:
raise ValueError('A bug occured somewhere in the forward pass')
compute_nodes_value(self)
# We can then do the same thing for the backward pass, taking advantage of the [chain rule](https://en.wikipedia.org/wiki/Chain_rule). Things are getting a little bit more trickier here, but the principle is simply to start from the gradient of the loss computed on the end node, and to backpropagate iteratively to the first node. In order to be sure I don't compute the gradient twice for a given node, I always check that the `grad_computed` is set to `False` before running any computation.
class Node(Node):
def compute_backward(self, d_loss):
"""Compute the gradient of the complete graph in a backward manner.
"""
def assess_children(children):
for child in children:
if not child.grad_computed:
return False
return True
def compute_grad_value(current_node, child = None):
if child is None:
current_node.grad = self.dic_d_op[current_node.type](d_loss)
current_node.grad_computed = True
else:
if child.grad_computed:
if current_node.type=='input':
current_node.grad += child.grad
elif current_node.type=='par':
if child.type == 'add':
current_node.grad += np.sum(child.grad, axis = 0)
if child.type == 'dot':
if len(child.children)==1:
child_children = next(iter(child.children))
current_node.grad += np.dot(child_children.grad.T, child.parents[1].value)
else:
child_grad = np.sum([c.grad for c in child.children], axis = 0)
current_node.grad += np.dot( child_grad.T, child.parents[1].value)
else:
current_node.grad += current_node.dic_d_op[current_node.type](child.grad)
else:
raise ValueError('A bug occured somewhere in the backward pass')
current_node.grad_computed = assess_children(self.children)
if current_node.grad_computed:
for parent in current_node.parents:
compute_grad_value(parent, child = current_node)
compute_grad_value(self)
# We can now create nodes (layers), but we still haven't defined any gradient descent algorithm, nor any loss... This will be done in the `Network` class, that we're going to implement right now.
#
# ## The Network class
# This class contains no argument but the output node of the network one wants to build.
class Network:
"""Contains all the nodes (layers) of the network.
"""
def __init__(self, output_node = Node()):
""" Constructor:
-output_node must correspond to the last node of the previously
built dynamical graph.
"""
self.output_node = output_node
self.input_node = self.find_input_node()
self.s_parameter_nodes = set()
# The input node is then found automatically by browsing the graph in an ascending manner.
class Network(Network):
def find_input_node(self):
""" Function to find the node corresponding to the first layer of the
dynamical graph using recursion.
"""
set_input = set()
def look_parent(current_node):
for parent in current_node.parents:
if parent.type=='input':
set_input.add(parent)
look_parent(parent)
look_parent(self.output_node)
if len(set_input)>1:
raise ValueError("There can only be one input to this graph.")
else:
return set_input.pop()
# The function to compute the foward pass is then nothing more than a wrapper of the same function in the `Node` class.
class Network(Network):
def do_forward(self, input):
""" Do the forward pass:
-input must be a FloatTensor or DoubleTensor. It is with this tensor
that we feed the network.
"""
self.input_node.value = input
self.output_node.compute_forward()
return self.output_node.value
# In preparation for the computation of the backward pass, we can now define the loss and its derivative. For now, we stick to a simple Mean-Squared error loss, but it wouldn't be complicated to do the same thing with, for instance, a cross-entropy loss. If you're interested, have a look at this [really good article](http://rohanvarma.me/Loss-Functions/) about the different types of losses and when to use them.
class Network(Network):
def MSE_loss(self, target):
""" Compute the Mean Squared Error (MSE) loss of the network.
-target must be a FloatTensor or DoubleTensor of the same size as
the ouput layer.
"""
if self.output_node.value is not None:
return 1/len(self.output_node.value)\
*np.sum((self.output_node.value-target)**2)
else:
raise ValueError("The loss can only be computed after having done \
the forward pass.")
def d_MSE_loss(self, target):
""" Compute the derivative of the Mean Squared Error (MSE) loss of the
network.
-target must be a FloatTensor or DoubleTensor of the same size as
the ouput layer.
"""
if self.output_node.value is not None:
return 2*(self.output_node.value-target)
else:
raise ValueError("The loss derivative can only be computed after \
having done the forward pass")
# We can now implement the wrapper for the backward pass, which takes as input both the target vector and the type of the loss.
class Network(Network):
def do_backward(self, target, type_loss):
""" Do the backward pass:
-target must be a FloatTensor or DoubleTensor of the same size as
the ouput layer.
-type_loss sets the type of loss function implemented. Only Mean
Squared Error (MSE) is available for now.
"""
if type_loss == "MSE":
d_loss = self.d_MSE_loss(target)
grad = self.output_node.compute_backward(d_loss)
else:
raise ValueError("Only MSE loss is implemented for now.")
# And as an optimization algorithm, we're going to implement the vanilla stochastic SGD. Since we first need to know which nodes are correspond to layers of parameters before optimizing them, we're also going to make a quick search through the graph before running the SGD itself. Also, we need to ensure every boolean flag telling us if the gradient has already been computed is set to `False` *before* doing the SGD, so we're going to define a function for that. In the end, we get:
class Network(Network):
def find_parameter_nodes(self):
""" Function to find and add in a set the parameters node of the graph.
"""
def find_parameter(current_node):
for parent in current_node.parents:
if parent.type=='par':
self.s_parameter_nodes.add(parent)
else:
find_parameter(parent)
find_parameter(self.output_node)
def reset_flags(self):
""" Function to reset the gradients and the corresponding flags of all
the nodes of the network.
"""
def reset_flag(current_node):
for parent in current_node.parents:
parent.grad_computed = False
parent.grad =0.
reset_flag(parent)
self.output_node.grad_computed = False
self.output_node.grad = 0.
reset_flag(self.output_node)
def do_SGD(self, input, target, type_loss = "MSE", n_iter = 1000, eta = 0.1,
size_batch = 50, verbose = False, plot = False):
""" Do the Stochastic Gradient Descent (SGD) to optimize the parameters
of the graph:
-input must be a FloatTensor or DoubleTensor. It is with this tensor
that we feed the network.
-target must be a FloatTensor or DoubleTensor of the same size as
the ouput layer.
-type_loss sets the type of loss function implemented. Only Mean
Squared Error (MSE) is available for nowself.
-n_iter is an integer specifying the number of SGD iterations.
-eta is a float specifying the the size of the linear expansion used
to compute the gradient.
-verbose is a bool specifying is the MSE loss must be printed at
each iteration.
"""
self.find_parameter_nodes()
l_error = []
n_subiter = int(len(input)/size_batch)
batched_input = [input[k2*size_batch:(k2+1)*size_batch,:] for k2 in range(n_subiter)]
batched_target = [target[k2*size_batch:(k2+1)*size_batch,:] for k2 in range(n_subiter)]
tot_iter = 0
for k in range(n_iter):
for sample, target_sample in zip(batched_input, batched_target):
self.do_forward(sample)
self.do_backward(target_sample, type_loss)
for node in self.s_parameter_nodes:
node.value -= eta * node.grad
self.reset_flags()
l_error.append(float(self.MSE_loss(target_sample)))
tot_iter += 1
if verbose:
print('it ' + str(k)+': '+ str(float(self.MSE_loss(target_sample))))
if plot:
plt.figure(figsize=(8,8))
plt.plot(np.linspace(0,n_iter,tot_iter, endpoint = True), l_error)
plt.xscale('log', basex=10)
plt.xlabel("Iteration")
plt.ylabel('MSE error')
plt.title('Evolution of the error (train)')
plt.show()
plt.close()
# And that's it, our Network class is now over! Let's see how well it works in practice.
# ## Running the network
# +
""" Create train and test sets """
train_set = np.random.rand(1000,2)
train_target = np.array([(0,1) if ( (x1-0.5)**2+(x2-0.5)**2)**0.5>1/(2*np.pi)**0.5 \
else (1,0) for (x1,x2) in train_set])
test_set = np.random.rand(1000,2)
test_target = np.array([(0,1) if ( (x1-0.5)**2+(x2-0.5)**2)**0.5>1/(2*np.pi)**0.5 \
else (1,0) for (x1,x2) in test_set])
nb_classes = train_set.shape[1]
nb_train_samples = train_set.shape[0]
""" Set meta-parameters """
nb_hidden = 25
eta = 1e-1 / nb_train_samples
epsilon = 1e-1
""" Create tensors of parameters for the network """
w1_tensor = np.random.randn(nb_hidden, train_set.shape[1])*epsilon
b1_tensor = np.random.randn(nb_hidden)*epsilon
w2_tensor = np.random.randn(nb_hidden, nb_hidden)*epsilon
b2_tensor = np.random.randn(nb_hidden)*epsilon
w3_tensor = np.random.randn(nb_classes, nb_hidden)*epsilon
b3_tensor = np.random.randn(nb_classes)*epsilon
""" Create computation graph (network) """
x0 = Node()
s1 = Node(b1_tensor).add( Node(w1_tensor).dot(x0) )
x1 = s1.relu()
s2 = Node(b2_tensor).add(Node(w2_tensor).dot(x1))
x2 = s2.relu()
s3 = Node(b3_tensor).add(Node(w3_tensor).dot(x2))
x3 = s3.tanh()
N = Network(x3)
""" Train the network """
print('Training started... It should take less than a minute.')
N.do_SGD(train_set, train_target, type_loss = "MSE", n_iter = 10000, eta = eta,
size_batch = 100, verbose = False, plot = False)
print('Training done')
""" Print final train error """
N.do_forward(train_set)
print('MSE on train set: ', float(N.MSE_loss(train_target)))
""" Print test error """
N.do_forward(test_set)
print('MSE on test set: ', float(N.MSE_loss(test_target)))
""" Plot the result """
l_color = ['r' if np.argmax(r)==1 else 'g' for r,t in zip(N.output_node.value, test_target)]
circle = plt.Circle((0.5, 0.5), 1/(2*np.pi)**0.5, color='b', alpha = 0.1)
fig, ax = plt.subplots(figsize=(8,8))
ax.add_artist(circle)
plt.scatter(np.array(test_set[:,0]), np.array(test_set[:,1]), c = l_color, s = 5)
plt.xlim([0,1])
plt.ylim([0,1])
plt.xlabel('x')
plt.ylabel('y')
plt.title('Classification result')
plt.show()
plt.close()
# -
# That was a little bit of work, but I believe that's the only way to really understand what actually happens in the network. Plus the result is really cool!
| DeepLearningFramework.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import matplotlib
# %matplotlib tk
# %autosave 180
# %load_ext autoreload
# %autoreload 2
import nest_asyncio
# %config Completer.use_jedi = False
#
import matplotlib.pyplot as plt
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
import scipy
import numpy as np
import pandas as pd
import os
import os
os.chdir('..')
from calcium import calcium
from wheel import wheel
from visualize import visualize
from tqdm import trange
from scipy.io import loadmat
import umap
from sklearn.decomposition import PCA
import plotly.express as px
from plotly.subplots import make_subplots
import plotly.graph_objects as go
#
np.set_printoptions(suppress=True)
# -
import h5py
fname = '/media/cat/4TB/donato/steffen/DON-004366/20210228/suite2p/plane0/pos.mat'
with h5py.File(fname, 'r') as f:
print (f.keys())
pos = np.array(f['pos']).squeeze()
print (pos.shape)
# +
#################################################
#################################################
#################################################
from scipy.ndimage import gaussian_filter1d
from matplotlib.gridspec import GridSpec
#
fname = '/media/cat/4TB/donato/steffen/DON-004366/20210228/suite2p/plane0/binarized_traces.npz'
d = np.load(fname,
allow_pickle=True)
#
F_thresholded = d['events_F_threshold_upphase']
oasis_smooth = d['events_Oasis_smooth_upphase']
spikes= d['spikes']
#
print (spikes.shape)
# +
#
y_f = []
y_oasis = []
fig=plt.figure()
imgs=[]
for k in range(3):
if k==0:
data_in = F_thresholded.copy()
label = 'F_thresholded'
elif k==1:
data_in = oasis_smooth.copy()
label = 'Oasis smoothed x F scaled'
elif k==2:
data_in = spikes.copy()
label = 'spikes'
ax = plt.subplot(3,1,k+1)
img = []
width=1
for unit in trange(data_in.shape[0]):
#for unit in np.random.randint(0,data_in.shape[0]):
#unit=57
idx = np.where(data_in[unit]>0)[0]
#print (np.unique(pos[idx]).shape, idx.shape)
if False:
y = np.histogram(pos[idx],bins=np.arange(0,180,width))
else:
y = np.histogram(pos[idx], weights = data_in[unit][idx],
bins=np.arange(0,180,width))
yy=y[0]/(np.max(y[0])+0.0001)
if k == 0:
y_f.append(yy)
elif k==1:
y_oasis.append(yy)
img.append(yy)
#
img=np.vstack(img)
if k==0:
idx2 = np.argsort(np.argmax(img,axis=1))
img=img[idx2]
vmax = np.max(img)*.8
plt.imshow(img,
aspect='auto',
cmap='magma',
vmin=0,
extent=[0,180,0,data_in.shape[0]],
vmax=vmax)
plt.ylabel(label + "\nNeuron Ids (ordered by peak time)")
imgs.append(img)
#
plt.show()
# -
#y_f = []
units = [1,43,54,268,272,416,678,820, 500]
ctr=0
fig=plt.figure()
for unit in units:
ax=plt.subplot(3,3,ctr+1)
labels = ['F','oasis','0.01cm']
#clrs = ['lightblue', 'royalblue','blue','lightcoral','pink','red']
clrs='blue', 'red'
#for k in range(len(y_f)):
t=np.linspace(0, 180, y_f[unit].shape[0])
if False:
plt.plot(t, y_f[unit]/np.max(y_f[unit]), label=labels[0],c=clrs[0])
plt.plot(t, y_oasis[unit]/np.max(y_oasis[unit]),label=labels[1],c=clrs[1])
else:
plt.plot(t, y_f[unit], label=labels[0],c=clrs[0])
plt.plot(t, y_oasis[unit],label=labels[1],c=clrs[1])
plt.legend(fontsize=20)
#plt.title("Spatial footprint as a function of time bin", fontsize=20)
#plt.xlabel("Location on track (cm)", fontsize=20)
plt.xlim(0,180)
plt.ylim(bottom=0)
ctr+=1
plt.show()
# +
fig=plt.figure()
ctr=0
k=0
while True:
#for ctr, k in enumerate(range(20,100,1)):
temp1 = y_f[k]
temp2 = y_oasis[k]
if np.max(temp1)==0 and np.max(temp2)==0:
k+=1
continue
ax=plt.subplot(20,4,ctr+1)
plt.ylabel(str(k),fontsize=6)
plt.plot(temp1/np.max(temp1),label="F_thresh",c='blue')
plt.plot(temp2/np.max(temp2), label = "Oasis x F x rolling window",c='red')
#plt.plot(temp2/np.max(temp2), c='magenta',
# label = "raw spikes")
#plt.plot(imgs[2][k]/100., label = "spikes")
plt.xlim(0,180)
#plt.ylim(bottom=0)
if ctr==0:
plt.legend(fontsize=6)
plt.xticks([])
plt.yticks([])
ctr+=1
k+=1
if ctr==80:
break
plt.show()
# -
| physiology_and_maps/behavior_1D_treadmill_Steffen.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # An introduction to Biogeme
# ## Biogeme Basics: Logit Model
import pandas as pd
import numpy as np
import biogeme.database as db
import biogeme.biogeme as bio
import biogeme.models as models
import biogeme.expressions as exp
import biogeme.tools as tools
import seaborn as sns
import matplotlib.pyplot as plt
# **Import Swissmetro data**
pandas = pd.read_csv("../../Data/8-DiscreteChoiceModels/swissmetro.dat",sep='\t')
database = db.Database("data/swissmetro", pandas)
# ## Let's see what this dataset has
# * dataset consists of survey data collected on the trains between St. Gallen and Geneva, Switzerland, during March 1998
# * It is necessary to obtain data from surveys of hypothetical markets/situations, which include the innovation, to assess the impact.
# * Survey data were collected on rail-based travels, interviewing 470 respondents. Due to data problems, only 441 are used here. A similar method for relevant car trips. A total of 1070 persons filled in the survey completely and were willing to participate in the second SP survey, which was generated using the same approach used for the rail interviews. 750 usable SP surveys were returned, from the license-plate based survey.
# * Nine stated choice situations were generated for each the respondents, offering three alternatives: rail, Swissmetro and car
#
# <NAME>., <NAME>. and <NAME>. (2001), The acceptance of modal innovation: The case of Swissmetro, in ‘Proceedings of the Swiss Transport Research Conference’, Ascona, Switzerland.
# 
# 
plt.figure(figsize=(10,5))
chart = sns.countplot(pandas['PURPOSE'])
chart.set_xticklabels(chart.get_xticklabels(), rotation=30, horizontalalignment='right');
chart.set_xticklabels(['Commuter', 'Shopping', 'Business', 'Leisure', 'Return from work','Return from shopping', 'Return from business','Return from leisure','Other']);
# **Use collumn names as variables**
globals().update(database.variables)
# **Exclude some unwanted entries**
# +
exclude = (( PURPOSE != 1 ) * ( PURPOSE != 3 ) + ( CHOICE == 0 )) > 0
database.remove(exclude)
# -
# **Define some dummy variables**
# +
SM_COST = SM_CO * ( GA == 0 )
TRAIN_COST = TRAIN_CO * ( GA == 0 )
CAR_AV_SP = exp.DefineVariable ('CAR_AV_SP', CAR_AV * ( SP !=0 ), database)
TRAIN_AV_SP = exp.DefineVariable ('TRAIN_AV_SP', TRAIN_AV * ( SP != 0 ), database)
# -
# **Rescale some data**
TRAIN_TT_SCALED = exp.DefineVariable('TRAIN_TT_SCALED', TRAIN_TT / 100.0, database)
TRAIN_COST_SCALED = exp.DefineVariable('TRAIN_COST_SCALED', TRAIN_COST / 100, database)
SM_TT_SCALED = exp.DefineVariable('SM_TT_SCALED', SM_TT / 100.0 , database)
SM_COST_SCALED = exp.DefineVariable('SM_COST_SCALED', SM_COST / 100 , database)
CAR_TT_SCALED = exp.DefineVariable('CAR_TT_SCALED', CAR_TT / 100 , database)
CAR_CO_SCALED = exp.DefineVariable('CAR_CO_SCALED', CAR_CO / 100 , database)
pandas = database.data
plt.figure(figsize=(10,5))
chart = sns.countplot(pandas['PURPOSE'])
chart.set_xticklabels(chart.get_xticklabels(), rotation=30, horizontalalignment='right');
chart.set_xticklabels(['Commuter', 'Business']);
# +
plt.figure(1, figsize=(10,5))
chart = sns.countplot(pandas['TRAIN_AV'])
chart.set_xticklabels(chart.get_xticklabels(), rotation=30, horizontalalignment='right');
chart.set_xticklabels(['Yes']);
chart.set(title='Train Availability', xlabel="Available", ylabel = "Count");
plt.figure(2, figsize=(10,5))
chart = sns.countplot(pandas['CAR_AV'])
chart.set_xticklabels(chart.get_xticklabels(), rotation=30, horizontalalignment='right');
chart.set_xticklabels(['No', 'Yes']);
chart.set(title='Car Availability', xlabel="Available", ylabel = "Count");
plt.figure(3, figsize=(10,5))
chart = sns.countplot(pandas['SM_AV'])
chart.set_xticklabels(chart.get_xticklabels(), rotation=30, horizontalalignment='right');
chart.set_xticklabels(['Yes']);
chart.set(title='Swissmetro Availability', xlabel="Available", ylabel = "Count");
# +
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
pandas.describe()
# -
# **Define the utility functions**
# \begin{align}
# V_1 & = \beta_{Train} + \beta_{time}X_{Train_{TT}} + \beta_{cost}X_{Train_{cost}}\\
# V_2 & = \beta_{SM} + \beta_{time}X_{SM_{TT}} + \beta_{cost}X_{SM_{cost}}\\
# V_3 & = \beta_{Car} + \beta_{time}X_{Car_{TT}} + \beta_{cost}X_{Car_{cost}}\\
# \end{align}
# **Create parameters to be estimated**
#
# `Beta`
# 1. name of parameter
# 2. default value for the parameter
# 3. lower bound
# 4. upper bound
# 5. flag indicating if parameter is to be estimated
ASC_CAR = exp.Beta('ASC_CAR',0,None ,None ,0)
ASC_TRAIN = exp.Beta('ASC_TRAIN',0,None ,None ,0)
ASC_SM = exp.Beta('ASC_SM',0,None ,None ,1)
B_TIME = exp.Beta('B_TIME',0,None ,None ,0)
B_COST = exp.Beta('B_COST',0,None ,None ,0)
# **Define the utility functions**
V1 = ASC_TRAIN + \
B_TIME * TRAIN_TT_SCALED + \
B_COST * TRAIN_COST_SCALED
V2 = ASC_SM + \
B_TIME * SM_TT_SCALED + \
B_COST * SM_COST_SCALED
V3 = ASC_CAR + \
B_TIME * CAR_TT_SCALED + \
B_COST * CAR_CO_SCALED
# **Associate utility functions with alternatives and associate availability of alternatives**
#
# Create a python dictionary with all utility functions
#
# Create a python dictionary with availability of choices
# +
V = {1: V1,
2: V2,
3: V3}
av = {1: TRAIN_AV_SP,
2: SM_AV,
3: CAR_AV_SP}
# -
# **Define the model**
logprob = models.loglogit(V, av, CHOICE)
# **Define the Biogeme object**
#
# * Give the database with all variables
# * Give the log likelihood model
# +
biogeme = bio.BIOGEME(database, logprob)
biogeme.modelName = "swissmetro_logit_basic"
# -
# **Estimate the model**
#
# 1. A `.html` can be generated with a report of the results and can be opened with a browser
# 2. A `.pickle` file can also be generaetd with a snapshot with the results. This file can then be used in other scripts
# +
biogeme.generateHtml = True
biogeme.generatePickle = False
results = biogeme.estimate()
print(f"HTML file: {results.data.htmlFileName}")
print(f"Pickle file: {results.data.pickleFileName }")
# -
# **Print results**
betas = results.getBetaValues()
for k,v in betas.items():
print(f"{k:10}=\t{v:.3g}")
# **Get the variance-covariance matrix**
results.getVarCovar()
# **Model results**
# * Number of estimated parameters($K$)
# * Sample size ($N$)
# * Number of excluded observations
# * Log likelihood of the sample with the default values for the parameters ($\mathcal{L}^i)$)
# * Log likelihood for the final estimated model ($\mathcal{L}^*)$)
# * Likelihood ratio:
# \begin{align}
# -2 (\mathcal{L}^i-\mathcal{L}^*)
# \end{align}
# * Rho-square for the init model
# \begin{align}
# \rho^2 = 1- \frac{\mathcal{L}^*}{\mathcal{L}^i}
# \end{align}
# * Rho-square adjusted for the init model
# \begin{align}
# \rho^2 = 1- \frac{\mathcal{L}^* - K}{\mathcal{L}^i}
# \end{align}
# * Akaike Information Criterion
# \begin{align}
# 2 K - 2 \mathcal{L}^*
# \end{align}
# * Bayesian Information Criterion
# \begin{align}
# K N- 2 \mathcal{L}^*
# \end{align}
# * Final gradient norm
# * Iterations
# * Optimization Time
# +
gs = results.getGeneralStatistics()
for k,v in gs.items():
print("{}= {}".format(k.ljust(45),v[0]))
# -
# **Likelihood ratio test**
# \begin{align}
# -2 (\mathcal{L}^i-\mathcal{L}^*)
# \end{align}
#
# \begin{align}
# Asymptotically \chi^2 Distributed
# \end{align}
# H0 -> Both models are equal
tools.likelihood_ratio_test((gs['Final log likelihood'][0],gs['Number of estimated parameters'][0]),
(gs['Init log likelihood'][0], 0),
significance_level=0.95)
| Code/8.1-MultinomialLogitAndProbitModels/.ipynb_checkpoints/01-biogeme-basics-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Time series basics
# * Trends
# * Seasonality
# * Cyclical
# # Introduction to statsmodels
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import statsmodels.api as sm
# Importing built-in datasets in statsmodels
df = sm.datasets.macrodata.load_pandas().data
df.head()
print(sm.datasets.macrodata.NOTE)
df.head()
df.tail()
# statsmodels.timeseriesanalysis.datetools
index = pd.Index(sm.tsa.datetools.dates_from_range('1959Q1', '2009Q3'))
index
df.index = index
df.head()
df['realgdp'].plot()
# ## Using the Hodrick-Prescott Filter for trend analysis
result = sm.tsa.filters.hpfilter(df['realgdp'])
result
type(result)
type(result[0])
type(result[1])
gdp_cycle, gdp_trend = result
df['trend'] = gdp_trend
df[['realgdp', 'trend']].plot()
# zooming in
df[['realgdp', 'trend']]['2000-03-31':].plot()
# # ETS Theory (Error-Trend-Seasonality)
# * Exponential Smoothing
# * Trend Methods Models
# * ETS Decomposition
# # EWMA Theory
# # (Exponentially Weighted Moving Averages)
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
airline = pd.read_csv('airline_passengers.csv', index_col = 'Month')
airline.head()
# this is a normal index
airline.index
# Get rid of all the missing values in this dataset
airline.dropna(inplace=True)
airline.index = pd.to_datetime(airline.index)
airline.head()
# now its a DatetimeIndex
airline.index
# Recap of making the SMA
airline['6-month-SMA'] = airline['Thousands of Passengers'].rolling(window=6).mean()
airline['12-month-SMA'] = airline['Thousands of Passengers'].rolling(window=12).mean()
airline.plot(figsize=(10,8))
# Weakness of SMA
# * Smaller windows will lead to more noise, rather than signal
# * It will always lag by the size of the window
# * It will never reach to full peak or valley of the data due to the averaging.
# * Does not really inform you about possible future behaviour, all it really does is describe trends in your data.
# * Extreme historical values can skew your SMA significantly
# ### Creating EWMA
airline['EWMA-12'] = airline['Thousands of Passengers'].ewm(span=12).mean()
airline[['Thousands of Passengers', 'EWMA-12']].plot(figsize=(10,8))
# Full reading on mathematics of EWMA
# * http://pandas.pydata.org/pandas-docs/stable/computation.html#exponentially-weighted-windows
# # ETS continued...
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
airline = pd.read_csv('airline_passengers.csv', index_col = 'Month')
airline.head()
airline.plot()
airline.dropna(inplace = True)
airline.index = pd.to_datetime(airline.index)
airline.head()
from statsmodels.tsa.seasonal import seasonal_decompose
# additive/ multiplicative models available
# suspected linear trend = use additive
# suspected non-linear trend = multiplicative model
result = seasonal_decompose(airline['Thousands of Passengers'], model='multiplicative')
result.seasonal
result.seasonal.plot()
result.trend.plot()
result.resid.plot()
fig = result.plot()
# # ARIMA models
# # Auto regressive integrated moving averages
# [https://people.duke.edu/~rnau/411arim3.htm]
# * **Autoregressive integrated moving average (ARIMA)** model is a generalization of an autoregressive moving average (ARMA) model.
# * ARIMA model types
# - **Non-seasonal ARIMA (for non-seasonal data)**
# - **Seasonal ARIMA (for seasonal data)**
# * ARIMA models are applied in some cases where data show evidence of non-stationarity, where an initial differencing step (corresponding to the 'integrated' part of the model) can be applied one or more times to eliminate the non-stationarity.
# * **Non-seasonal ARIMA models are generally denoted as ARIMA(p, d, q)** where parameters p, d and q are non-negative integers.
# * **AR(p): Autoregression component**
# - A regression model that utilizes the dependent relationship between a current observation and observations over a previous period.
# * **I(d): Integrated**
# - Differencing of observations (subtracting an observation from an observation at the previous time step) in order to make the time series stationary.
# * **MA(q): Moving Average**
# - A model that uses the dependency between an observation and a residual error from a moving average model applied to lagged observations
# ### Stationary vs Non-Stationary Data
# * A stationary series has a constant mean and variance over time
# * A stationary dataset will allow our model to predict that the mean and variance will be the same in future periods.
# ___
#
# <img src='stationaryvsnonstationary.png' />
# ___
# * Note above for stationary data (mean and variance both are constant over time)
# * Another aspect to look for is covariance not be a function of time in stationary data
# * If you've determined your data is not stationary (either visually or mathematically), you will then need to transform it to be stationary in order to evaluate it and what type of ARIMA terms you will use.
# * One simple way to do this is through **"differencing"**
# * Original Data
# <table>
# <tr><td>Time1</td><td>10</td></tr>
# <tr><td>Time2</td><td>12</td></tr>
# <tr><td>Time3</td><td>8</td></tr>
# <tr><td>Time4</td><td>14</td></tr>
# <tr><td>Time5</td><td>7</td></tr>
# </table>
#
# * First Difference
# <table>
# <tr><td>Time1</td><td>NA</td></tr>
# <tr><td>Time2</td><td>2</td></tr>
# <tr><td>Time3</td><td>-4</td></tr>
# <tr><td>Time4</td><td>6</td></tr>
# <tr><td>Time5</td><td>-7</td></tr>
# </table>
#
# * Second Difference
# <table>
# <tr><td>Time1</td><td>NA</td></tr>
# <tr><td>Time2</td><td>NA</td></tr>
# <tr><td>Time3</td><td>-6</td></tr>
# <tr><td>Time4</td><td>10</td></tr>
# <tr><td>Time5</td><td>-13</td></tr>
# </table>
# * **For seasonal data,** you can also difference by season. If you had monthly data with yearly seasonality, you could difference by a time unit of 12, instead of just 1.
#
# * Another common techinique with seasonal ARIMA models is to combine both methods, taking the seasonal difference of the first difference.
# # ARIMA models continued... 1
# The general process for ARIMA models is the following:
# * Visualize the Time Series Data
# * Make the time series data stationary
# * Plot the Correlation and AutoCorrelation Charts
# * Construct the ARIMA Model
# * Use the model to make predictions
#
# Let's go through these steps!
# [https://people.duke.edu/~rnau/arimrule.htm]
import numpy as np
import pandas as pd
import statsmodels.api as sm
import matplotlib.pyplot as plt
# %matplotlib inline
df = pd.read_csv('monthly-milk-production-pounds-p.csv')
df.head()
df.columns = ['Month', 'Milk in Pounds per Cow']
df.head()
df.tail()
df.drop(168, axis=0, inplace=True)
df.tail()
df['Month'] = pd.to_datetime(df['Month'])
df.head()
df.set_index('Month', inplace=True)
df.head()
df.index
df.describe()
df.describe().transpose()
# ### Step 1 - Visualize the data
df.plot();
time_series = df['Milk in Pounds per Cow']
type(time_series)
time_series.rolling(12).mean().plot(label='12 SMA')
time_series.rolling(12).std().plot(label='12 STD')
time_series.plot()
plt.legend();
# ### Conclusion: The scale of STD (standard deviation) is always pretty much smaller than the actual scale. If 12 STD does not show crazy behaviour is comparitively flat then its 'workable'
from statsmodels.tsa.seasonal import seasonal_decompose
decomp = seasonal_decompose(time_series)
fig = decomp.plot()
fig.set_size_inches(15,8)
# # ARIMA models continued... 2
# ### Step 2 - Make the time series data stationary (if non-stationary)
# We can use the Augmented [Dickey-Fuller](https://en.wikipedia.org/wiki/Augmented_Dickey%E2%80%93Fuller_test) [unit root test](https://en.wikipedia.org/wiki/Unit_root_test).
#
# In statistics and econometrics, an augmented Dickey–Fuller test (ADF) tests the null hypothesis that a unit root is present in a time series sample. The alternative hypothesis is different depending on which version of the test is used, but is usually stationarity or trend-stationarity.
#
# Basically, we are trying to whether to accept the Null Hypothesis **H0** (that the time series has a unit root, indicating it is non-stationary) or reject **H0** and go with the Alternative Hypothesis (that the time series has no unit root and is stationary).
#
# We end up deciding this based on the p-value return.
#
# * A small p-value (typically ≤ 0.05) indicates strong evidence against the null hypothesis, so you reject the null hypothesis.
#
# * A large p-value (> 0.05) indicates weak evidence against the null hypothesis, so you fail to reject the null hypothesis.
#
# Let's run the Augmented Dickey-Fuller test on our data:
from statsmodels.tsa.stattools import adfuller
result = adfuller(df['Milk in Pounds per Cow'])
result
def adf_check(time_series):
result = adfuller(time_series)
print('Augumented Dicky-Fuller Test')
labels = ['ADF Test Statistic', 'p-value', '# of lags', 'Num of Observations used']
for value, label in zip(result, labels):
print(label + ': ' + str(value))
if result[1] < 0.05:
print('Strong evidence against null hypothesis')
print('Rejecting null hypothesis')
print('Data has no unit root! and is stationary')
else:
print('Weak evidence against null hypothesis')
print('Fail to reject null hypothesis')
print('Data has a unit root, it is non-stationary')
adf_check(df['Milk in Pounds per Cow'])
# * Thus, the ADF test confirms our assumption from visual analysis that definately the data is non-stationary and has a seasionality and trend factor to it.
# Now making the data stationary
df['First Difference'] = df['Milk in Pounds per Cow'] - df['Milk in Pounds per Cow'].shift(1)
df['First Difference'].plot()
# +
# adf_check(df['First Difference']) - THIS RESULTS IN LinAlgError: SVD did not converge ERROR
# -
# Note: we need to drop the first NA value before plotting this
adf_check(df['First Difference'].dropna())
df['Second Difference'] = df['First Difference'] - df['First Difference'].shift(1)
df['Second Difference'].plot();
adf_check(df['Second Difference'].dropna())
# * Since, **p-value('Original') - p-value('First Difference') < p-value('First Difference') - p-value('Second Difference')**, it is the first difference that did most of the elimination of the trend.
# Let's plot seasonal difference
df['Seasonal Difference'] = df['Milk in Pounds per Cow'] - df['Milk in Pounds per Cow'].shift(12)
df['Seasonal Difference'].plot();
adf_check(df['Seasonal Difference'].dropna())
# * Thus, we conclude that seasonal difference *does not make* the data stationary here, **in fact we can observe visually that as we go further in time the variance began to increase**.
# Plotting 'Seasonal first difference'
df['Seasonal First Difference'] = df['First Difference'] - df['First Difference'].shift(12)
df['Seasonal First Difference'].plot();
adf_check(df['Seasonal First Difference'].dropna())
# # ARIMA models continued... 3
# ### Step 3 - Plot the Correlation and Autocorrelation Charts
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
# Plotting the gradual decline autocorrelation
fig_first = plot_acf(df['First Difference'].dropna())
fig_first = plot_acf(df['First Difference'].dropna(), use_vlines=False)
fig_seasonal_first = plot_acf(df['Seasonal First Difference'].dropna(), use_vlines=False)
fig_seasonal_first_pacf = plot_pacf(df['Seasonal First Difference'].dropna(), use_vlines=False)
# #### Plotting the final 'Autocorrelation' and 'Partial autocorrelation'
plot_acf(df['Seasonal First Difference'].dropna());
plot_pacf(df['Seasonal First Difference'].dropna());
# # ARIMA models continued... 4
# ### Step 4 - Construct the ARIMA model
# ARIMA model for non-sesonal data
from statsmodels.tsa.arima_model import ARIMA
# +
# help(ARIMA)
# +
# ARIMA model from seasonal data
# from statsmodels.tsa.statespace import sarimax
# -
# Choosing the **p, d, q values** of the **order** and **seasonal_order** tuple is reading task
#
# More information here...
#
# [https://stackoverflow.com/questions/22770352/auto-arima-equivalent-for-python]
#
# [https://stats.stackexchange.com/questions/44992/what-are-the-values-p-d-q-in-arima]
#
# [https://people.duke.edu/~rnau/arimrule.htm]
model = sm.tsa.statespace.SARIMAX(df['Milk in Pounds per Cow'], order=(0,1,0), seasonal_order=(1,1,1,12))
results = model.fit()
print(results.summary())
# residual errors of prediction on the original training data
results.resid
# plot of residual errors of prediction on the original training data
results.resid.plot();
# KDE plot of residual errors of prediction on the original training data
results.resid.plot(kind='kde');
# Creating a column forecast to house the forecasted values for existing values
df['forecast'] = results.predict(start=150, end=168)
df[['Milk in Pounds per Cow', 'forecast']].plot(figsize=(12,8));
# Forecasting for future data
df.tail()
from pandas.tseries.offsets import DateOffset
future_dates = [df.index[-1] + DateOffset(months=x) for x in range(0,24)]
future_dates
future_df = pd.DataFrame(index=future_dates, columns=df.columns)
future_df.head()
final_df = pd.concat([df, future_df])
final_df.head()
final_df.tail()
final_df['forecast'] = results.predict(start=168, end=192)
final_df.tail()
final_df[['Milk in Pounds per Cow', 'forecast']].plot()
# * **Why ARIMA Models are questioning when it comes to financial forecasting?**
#
# Lot of this stuff assumes that the y-axis value (price) is directly connected to the time (x-axis value) and that the time is really important aspect of the y value.
#
# While that is true for financial series it discounts the **external force** i.e. traders also able to buy and sell securities outside the market and affect its price.
#
# And because of that often you'll hear stock and securities prices are following some sort of **Brownian motion** almost like a random walk.
#
# Because of those aspects of the financial and securities data this sort of forecsting method *doesn't really work with stock*.
| time_series_analysis/time_series_analysis_notes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# Test new TBmodels - kwant interface
# ===================
# + deletable=true editable=true
import kwant
import tbmodels
from matplotlib import pyplot as plt
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
# + [markdown] deletable=true editable=true
# Simple square lattice
# ---------------------
# + deletable=true editable=true
# Test - do simple square lattice
model = tbmodels.Model(
on_site=[1.3],
dim=2,
occ=0,
pos=[[0., 0.]],
uc = [[1, 0], [0, 1]]
)
t1 = 1.0
#for R in ([1,0],[0,1],[0,2],[0,3],[-2,4]): - throws an error
for R in ([1,0],[0,1],[0,2],[0,3],[-2,4]):
model.add_hop(t1, 0, 0, R)
# + deletable=true editable=true
kwant_sys = kwant.Builder()
# + deletable=true editable=true
latt = model.to_kwant_lattice()
# + deletable=true editable=true
def shape(pos):
x, y = pos
return -10 < x < 10 and -5 < y < 5
# + deletable=true editable=true
kwant_sys[latt.shape(shape, (0, 0))] = 0
# + deletable=true editable=true
model.add_hoppings_kwant(kwant_sys)
# + deletable=true editable=true
kwant.plot(kwant_sys)
# -
# Test with wraparound
# -----------------------
import wraparound
import scipy.linalg as la
latt = model.to_kwant_lattice()
sym = kwant.TranslationalSymmetry(
latt.vec((1, 0)),
latt.vec((0, 1))
)
sys = kwant.Builder(sym)
sys[latt.shape(lambda p: True, (0, 0))] = 0
model.add_hoppings_kwant(sys)
sys_wrap = wraparound.wraparound(sys).finalized()
# + deletable=true editable=true
H1 = model.hamilton((0, 0.1))
# -
H2 = sys_wrap.hamiltonian_submatrix((0, 0.2 * np.pi))
H1
H2
# + [markdown] deletable=true editable=true
# Realistic tb model
# --------------------
# + deletable=true editable=true
model2 = tbmodels.Model.from_hr_file('wannier90_hr.dat')
# + deletable=true editable=true
sys2 = kwant.Builder()
latt2 = model2.to_kwant_lattice()
# + deletable=true editable=true
def shape2(pos):
x, y, z = pos
return -10 < x < 10 and -5 < y < 5 and -5 < z < 5
# + deletable=true editable=true
sys2[latt2.shape(shape2, (0, 0, 0))] = 0
# + deletable=true editable=true
model2.add_hoppings_kwant(sys2)
# + deletable=true editable=true
kwant.plot(sys2)
# + [markdown] deletable=true editable=true
# Test with wraparound
# -----------------------
# -
latt3 = model2.to_kwant_lattice()
sym3 = kwant.TranslationalSymmetry(
latt3.vec((1, 0, 0)),
latt3.vec((0, 1, 0)),
latt3.vec((0, 0,1))
)
sys3 = kwant.Builder(sym3)
sys3[latt3.shape(lambda p: True, (0, 0, 0))] = 0
model2.add_hoppings_kwant(sys3)
sys3_wrap = wraparound.wraparound(sys3).finalized()
# + deletable=true editable=true
tbmodels_ham = model2.hamilton((0, 0.1, 0))
# -
kwant_ham = sys3_wrap.hamiltonian_submatrix((0., 0.1 * 2 * np.pi, 0.))
la.eigvalsh(tbmodels_ham)
la.eigvalsh(kwant_ham)
np.isclose(tbmodels_ham, kwant_ham).all()
| playground/to_kwant/kwant_test_v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: nams
# language: python
# name: nams
# ---
# + slideshow={"slide_type": "slide"}
import pandas as pd
import networkx as nx
import os
import numpy as np
import warnings
import numpy as np
import matplotlib.pyplot as plt
from nxviz import CircosPlot
warnings.filterwarnings('ignore')
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
# + [markdown] slideshow={"slide_type": "slide"}
# # Tables to Networks, Networks to Tables
#
# Networks can be represented in a tabular form in two ways: As an adjacency list with edge attributes stored as columnar values, and as a node list with node attributes stored as columnar values.
#
# Storing the network data as a single massive adjacency table, with node attributes repeated on each row, can get unwieldy, especially if the graph is large, or grows to be so. One way to get around this is to store two files: one with node data and node attributes, and one with edge data and edge attributes.
#
# The Divvy bike sharing dataset is one such example of a network data set that has been stored as such.
# + [markdown] slideshow={"slide_type": "slide"}
# # Loading Node Lists and Adjacency Lists
#
# Let's use the Divvy bike sharing data set as a starting point. The Divvy data set is comprised of the following data:
#
# - Stations and metadata (like a node list with attributes saved)
# - Trips and metadata (like an edge list with attributes saved)
#
# The `README.txt` file in the Divvy directory should help orient you around the data.
# + slideshow={"slide_type": "fragment"}
# This block of code checks to make sure that a particular directory is present.
if "divvy_2013" not in os.listdir('datasets/'):
print('Unzip the divvy_2013.zip file in the datasets folder.')
# + slideshow={"slide_type": "subslide"}
stations = pd.read_csv('datasets/divvy_2013/Divvy_Stations_2013.csv', parse_dates=['online date'], encoding='utf-8')
stations.head(10)
# + slideshow={"slide_type": "subslide"}
trips = pd.read_csv('datasets/divvy_2013/Divvy_Trips_2013.csv',
parse_dates=['starttime', 'stoptime'])
trips.head(10)
# + [markdown] slideshow={"slide_type": "subslide"}
# At this point, we have our `stations` and `trips` data loaded into memory.
#
# How we construct the graph depends on the kind of questions we want to answer, which makes the definition of the "unit of consideration" (or the entities for which we are trying to model their relationships) is extremely important.
#
# Let's try to answer the question: "What are the most popular trip paths?" In this case, the bike station is a reasonable "unit of consideration", so we will use the bike stations as the nodes.
#
# To start, let's initialize an directed graph `G`.
# + slideshow={"slide_type": "fragment"}
G = nx.DiGraph()
# + [markdown] slideshow={"slide_type": "subslide"}
# Then, let's iterate over the `stations` DataFrame, and add in the node attributes.
# + slideshow={"slide_type": "fragment"}
for d in stations.to_dict('records'): # each row is a dictionary
node_id = d.pop('id')
G.add_node(node_id, attr_dict=d)
# + [markdown] slideshow={"slide_type": "subslide"}
# In order to answer the question of "which stations are important", we need to specify things a bit more. Perhaps a measure such as **betweenness centrality** or **degree centrality** may be appropriate here.
#
# The naive way would be to iterate over all the rows. Go ahead and try it at your own risk - it may take a long time :-). Alternatively, I would suggest doing a `pandas` `groupby`.
# + slideshow={"slide_type": "subslide"}
# # Run the following code at your own risk :)
# for r, d in trips.iterrows():
# start = d['from_station_id']
# end = d['to_station_id']
# if (start, end) not in G.edges():
# G.add_edge(start, end, count=1)
# else:
# G.edge[start][end]['count'] += 1
# -
counts = trips.groupby(['from_station_id', 'to_station_id'])['trip_id'].count().reset_index()
for d in counts.to_dict('records'):
G.add_edge(d['from_station_id'], d['to_station_id'], count=d['trip_id'])
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Exercise
#
# Flex your memory muscles: can you make a scatter plot of the distribution of the number edges that have a certain number of trips? (3 min.)
#
# The x-value is the number of trips taken between two stations, and the y-vale is be the number of edges that have that number of trips.
# + slideshow={"slide_type": "subslide"}
from collections import Counter
# Count the number of edges that have x trips recorded on them.
trip_count_distr = Counter([d['count'] for _, _, d in G.edges(data=True)])
# Then plot the distribution of these
plt.scatter(list(trip_count_distr.keys()), list(trip_count_distr.values()), alpha=0.1)
plt.yscale('log')
plt.xlabel('num. of trips')
plt.ylabel('num. of edges')
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Exercise
#
# Create a new graph, and filter out the edges such that only those with more than 100 trips taken (i.e. `count >= 100`) are left. (3 min.)
# + slideshow={"slide_type": "subslide"}
# Filter the edges to just those with more than 100 trips.
G_filtered = G.copy()
for u, v, d in G.edges(data=True):
if d['count'] < 100:
G_filtered.remove_edge(u,v)
len(G_filtered.edges())
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's now try drawing the graph.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Exercise
#
# Use `nx.draw(my_graph)` to draw the filtered graph to screen. (1 min.)
# + slideshow={"slide_type": "subslide"}
nx.draw(G_filtered)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Exercise
#
# Try visualizing the graph using a CircosPlot. Order the nodes by their connectivity in the **original** graph, but plot only the **filtered** graph edges. (3 min.)
#
# You may have to first annotate the connectivity of each node, as given by the number of neighbors that any node is connected to.
# + slideshow={"slide_type": "subslide"}
for n in G_filtered.nodes():
G_filtered.node[n]['connectivity'] = len(G.neighbors(n))
c = CircosPlot(G_filtered, node_order='connectivity')
c.draw()
plt.savefig('images/divvy.png', dpi=300)
# + [markdown] slideshow={"slide_type": "subslide"}
# In this visual, nodes are sorted from highest connectivity to lowest connectivity in the **unfiltered** graph.
#
# Edges represent only trips that were taken >100 times between those two nodes.
#
# Some things should be quite evident here. There are lots of trips between the highly connected nodes and other nodes, but there are local "high traffic" connections between stations of low connectivity as well (nodes in the top-right quadrant).
# + [markdown] slideshow={"slide_type": "slide"}
# # Saving NetworkX Graph Files
#
# NetworkX's API offers many formats for storing graphs to disk. If you intend to work exclusively with NetworkX, then pickling the file to disk is probably the easiest way.
#
# To write to disk:
#
# nx.write_gpickle(G, handle)
#
# To load from disk:
#
# G = nx.read_gpickle(handle)
# + slideshow={"slide_type": "subslide"}
nx.write_gpickle(G, 'datasets/divvy_2013/divvy_graph.pkl')
# + slideshow={"slide_type": "fragment"}
G = nx.read_gpickle('datasets/divvy_2013/divvy_graph.pkl')
G.nodes(data=True)[0:2]
# -
| 5-graph-input-output-instructor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
#hide
from nbdev import *
# # Test Cards Demo
#
# > How to play cards!
# This file will become your README and also the index of your documentation.
# ## Install
# `pip install test_cards_demo`
# ## How to use
# Our main classes are for a single card and a deck of cards, let's see them in action below:
from test_cards_demo.card import Card
from test_cards_demo.deck import Deck
d = Deck()
print(f'Number of playing cards in the deck: {len(d.cards)}')
print(d.pop_card())
| index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://cognitiveclass.ai"><img src = "https://ibm.box.com/shared/static/9gegpsmnsoo25ikkbl4qzlvlyjbgxs5x.png" width = 400> </a>
#
# <h1 align=center><font size = 5>From Understanding to Preparation</font></h1>
# ## Introduction
#
# In this lab, we will continue learning about the data science methodology, and focus on the **Data Understanding** and the **Data Preparation** stages.
# ## Table of Contents
#
# <div class="alert alert-block alert-info" style="margin-top: 20px">
# 1. [Recap](#0)<br>
# 2. [Data Understanding](#2)<br>
# 3. [Data Preparation](#4)<br>
# </div>
# <hr>
# # Recap <a id="0"></a>
# In Lab **From Requirements to Collection**, we learned that the data we need to answer the question developed in the business understanding stage, namely *can we automate the process of determining the cuisine of a given recipe?*, is readily available. A researcher named <NAME> scraped tens of thousands of food recipes (cuisines and ingredients) from three different websites, namely:
# <img src = "https://ibm.box.com/shared/static/4fruwan7wmjov3gywiz3swlojw0srv54.png" width=500>
#
# www.allrecipes.com
#
# <img src = "https://ibm.box.com/shared/static/cebfdbr22fjxa47lltp0bs533r103g0z.png" width=500>
#
# www.epicurious.com
#
# <img src = "https://ibm.box.com/shared/static/epk727njg7xrz49pbkpkzd05cm5ywqmu.png" width=500>
#
# www.menupan.com
# For more information on <NAME> and his research, you can read his paper on [Flavor Network and the Principles of Food Pairing](http://yongyeol.com/papers/ahn-flavornet-2011.pdf).
# We also collected the data and placed it on an IBM server for your convenience.
#
# ------------
# # Data Understanding <a id="2"></a>
# <img src="https://ibm.box.com/shared/static/89geb3m0ge1z73s92hl8o8wdcpcrggtz.png" width=500>
# <strong> Important note:</strong> Please note that you are not expected to know how to program in Python. The following code is meant to illustrate the stages of data understanding and data preparation, so it is totally fine if you do not understand the individual lines of code. We have a full course on programming in Python, <a href="http://cocl.us/PY0101EN_DS0103EN_LAB3_PYTHON_Coursera"><strong>Python for Data Science</strong></a>, which is also offered on Coursera. So make sure to complete the Python course if you are interested in learning how to program in Python.
# ### Using this notebook:
#
# To run any of the following cells of code, you can type **Shift + Enter** to excute the code in a cell.
# Get the version of Python installed.
# check Python version
# !python -V
# Download the library and dependencies that we will need to run this lab.
import pandas as pd # import library to read data into dataframe
pd.set_option('display.max_columns', None)
import numpy as np # import numpy library
import re # import library for regular expression
# Download the data from the IBM server and read it into a *pandas* dataframe.
# +
recipes = pd.read_csv("https://ibm.box.com/shared/static/5wah9atr5o1akuuavl2z9tkjzdinr1lv.csv")
print("Data read into dataframe!") # takes about 30 seconds
# -
# Show the first few rows.
recipes.head()
# Get the dimensions of the dataframe.
recipes.shape
# So our dataset consists of 57,691 recipes. Each row represents a recipe, and for each recipe, the corresponding cuisine is documented as well as whether 384 ingredients exist in the recipe or not, beginning with almond and ending with zucchini.
# We know that a basic sushi recipe includes the ingredients:
# * rice
# * soy sauce
# * wasabi
# * some fish/vegetables
# Let's check that these ingredients exist in our dataframe:
# +
ingredients = list(recipes.columns.values)
print([match.group(0) for ingredient in ingredients for match in [(re.compile(".*(rice).*")).search(ingredient)] if match])
print([match.group(0) for ingredient in ingredients for match in [(re.compile(".*(wasabi).*")).search(ingredient)] if match])
print([match.group(0) for ingredient in ingredients for match in [(re.compile(".*(soy).*")).search(ingredient)] if match])
# -
# Yes, they do!
#
# * rice exists as rice.
# * wasabi exists as wasabi.
# * soy exists as soy_sauce.
#
# So maybe if a recipe contains all three ingredients: rice, wasabi, and soy_sauce, then we can confidently say that the recipe is a **Japanese** cuisine! Let's keep this in mind!
#
# ----------------
# # Data Preparation <a id="4"></a>
# <img src="https://ibm.box.com/shared/static/lqc2j3r0ndhokh77mubohwjqybzf8dhk.png" width=500>
# In this section, we will prepare data for the next stage in the data science methodology, which is modeling. This stage involves exploring the data further and making sure that it is in the right format for the machine learning algorithm that we selected in the analytic approach stage, which is decision trees.
# First, look at the data to see if it needs cleaning.
recipes["country"].value_counts() # frequency table
# By looking at the above table, we can make the following observations:
#
# 1. Cuisine column is labeled as Country, which is inaccurate.
# 2. Cuisine names are not consistent as not all of them start with an uppercase first letter.
# 3. Some cuisines are duplicated as variation of the country name, such as Vietnam and Vietnamese.
# 4. Some cuisines have very few recipes.
# #### Let's fixes these problems.
# Fix the name of the column showing the cuisine.
# +
column_names = recipes.columns.values
column_names[0] = "cuisine"
recipes.columns = column_names
recipes
# -
# Make all the cuisine names lowercase.
recipes["cuisine"] = recipes["cuisine"].str.lower()
# Make the cuisine names consistent.
# +
recipes.loc[recipes["cuisine"] == "austria", "cuisine"] = "austrian"
recipes.loc[recipes["cuisine"] == "belgium", "cuisine"] = "belgian"
recipes.loc[recipes["cuisine"] == "china", "cuisine"] = "chinese"
recipes.loc[recipes["cuisine"] == "canada", "cuisine"] = "canadian"
recipes.loc[recipes["cuisine"] == "netherlands", "cuisine"] = "dutch"
recipes.loc[recipes["cuisine"] == "france", "cuisine"] = "french"
recipes.loc[recipes["cuisine"] == "germany", "cuisine"] = "german"
recipes.loc[recipes["cuisine"] == "india", "cuisine"] = "indian"
recipes.loc[recipes["cuisine"] == "indonesia", "cuisine"] = "indonesian"
recipes.loc[recipes["cuisine"] == "iran", "cuisine"] = "iranian"
recipes.loc[recipes["cuisine"] == "italy", "cuisine"] = "italian"
recipes.loc[recipes["cuisine"] == "japan", "cuisine"] = "japanese"
recipes.loc[recipes["cuisine"] == "israel", "cuisine"] = "jewish"
recipes.loc[recipes["cuisine"] == "korea", "cuisine"] = "korean"
recipes.loc[recipes["cuisine"] == "lebanon", "cuisine"] = "lebanese"
recipes.loc[recipes["cuisine"] == "malaysia", "cuisine"] = "malaysian"
recipes.loc[recipes["cuisine"] == "mexico", "cuisine"] = "mexican"
recipes.loc[recipes["cuisine"] == "pakistan", "cuisine"] = "pakistani"
recipes.loc[recipes["cuisine"] == "philippines", "cuisine"] = "philippine"
recipes.loc[recipes["cuisine"] == "scandinavia", "cuisine"] = "scandinavian"
recipes.loc[recipes["cuisine"] == "spain", "cuisine"] = "spanish_portuguese"
recipes.loc[recipes["cuisine"] == "portugal", "cuisine"] = "spanish_portuguese"
recipes.loc[recipes["cuisine"] == "switzerland", "cuisine"] = "swiss"
recipes.loc[recipes["cuisine"] == "thailand", "cuisine"] = "thai"
recipes.loc[recipes["cuisine"] == "turkey", "cuisine"] = "turkish"
recipes.loc[recipes["cuisine"] == "vietnam", "cuisine"] = "vietnamese"
recipes.loc[recipes["cuisine"] == "uk-and-ireland", "cuisine"] = "uk-and-irish"
recipes.loc[recipes["cuisine"] == "irish", "cuisine"] = "uk-and-irish"
recipes
# -
# Remove cuisines with < 50 recipes.
# +
# get list of cuisines to keep
recipes_counts = recipes["cuisine"].value_counts()
cuisines_indices = recipes_counts > 50
cuisines_to_keep = list(np.array(recipes_counts.index.values)[np.array(cuisines_indices)])
# +
rows_before = recipes.shape[0] # number of rows of original dataframe
print("Number of rows of original dataframe is {}.".format(rows_before))
recipes = recipes.loc[recipes['cuisine'].isin(cuisines_to_keep)]
rows_after = recipes.shape[0] # number of rows of processed dataframe
print("Number of rows of processed dataframe is {}.".format(rows_after))
print("{} rows removed!".format(rows_before - rows_after))
# -
# Convert all Yes's to 1's and the No's to 0's
recipes = recipes.replace(to_replace="Yes", value=1)
recipes = recipes.replace(to_replace="No", value=0)
# #### Let's analyze the data a little more in order to learn the data better and note any interesting preliminary observations.
# Run the following cell to get the recipes that contain **rice** *and* **soy** *and* **wasabi** *and* **seaweed**.
recipes.head()
# +
check_recipes = recipes.loc[
(recipes["rice"] == 1) &
(recipes["soy_sauce"] == 1) &
(recipes["wasabi"] == 1) &
(recipes["seaweed"] == 1)
]
check_recipes
# -
# Based on the results of the above code, can we classify all recipes that contain **rice** *and* **soy** *and* **wasabi** *and* **seaweed** as **Japanese** recipes? Why?
# + active=""
# Your Answer:
# -
# Double-click __here__ for the solution.
# <!-- The correct answer is:
# No, because other recipes such as Asian and East_Asian recipes also contain these ingredients.
# -->
# Let's count the ingredients across all recipes.
# sum each column
ing = recipes.iloc[:, 1:].sum(axis=0)
# +
# define each column as a pandas series
ingredient = pd.Series(ing.index.values, index = np.arange(len(ing)))
count = pd.Series(list(ing), index = np.arange(len(ing)))
# create the dataframe
ing_df = pd.DataFrame(dict(ingredient = ingredient, count = count))
ing_df = ing_df[["ingredient", "count"]]
print(ing_df.to_string())
# -
# Now we have a dataframe of ingredients and their total counts across all recipes. Let's sort this dataframe in descending order.
# +
ing_df.sort_values(["count"], ascending=False, inplace=True)
ing_df.reset_index(inplace=True, drop=True)
print(ing_df)
# -
# #### What are the 3 most popular ingredients?
# + active=""
# Your Answer:
# 1.
#
# 2.
#
# 3.
# -
# Double-click __here__ for the solution.
# <!-- The correct answer is:
# // 1. Egg with 21,025 occurrences.
# // 2. Wheat with 20,781 occurrences.
# // 3. Butter with 20,719 occurrences.
# -->
# However, note that there is a problem with the above table. There are ~40,000 American recipes in our dataset, which means that the data is biased towards American ingredients.
# **Therefore**, let's compute a more objective summary of the ingredients by looking at the ingredients per cuisine.
# #### Let's create a *profile* for each cuisine.
#
# In other words, let's try to find out what ingredients Chinese people typically use, and what is **Canadian** food for example.
cuisines = recipes.groupby("cuisine").mean()
cuisines.head()
# As shown above, we have just created a dataframe where each row is a cuisine and each column (except for the first column) is an ingredient, and the row values represent the percentage of each ingredient in the corresponding cuisine.
#
# **For example**:
#
# * *almond* is present across 15.65% of all of the **African** recipes.
# * *butter* is present across 38.11% of all of the **Canadian** recipes.
# Let's print out the profile for each cuisine by displaying the top four ingredients in each cuisine.
# +
num_ingredients = 4 # define number of top ingredients to print
# define a function that prints the top ingredients for each cuisine
def print_top_ingredients(row):
print(row.name.upper())
row_sorted = row.sort_values(ascending=False)*100
top_ingredients = list(row_sorted.index.values)[0:num_ingredients]
row_sorted = list(row_sorted)[0:num_ingredients]
for ind, ingredient in enumerate(top_ingredients):
print("%s (%d%%)" % (ingredient, row_sorted[ind]), end=' ')
print("\n")
# apply function to cuisines dataframe
create_cuisines_profiles = cuisines.apply(print_top_ingredients, axis=1)
# -
# At this point, we feel that we have understood the data well and the data is ready and is in the right format for modeling!
#
# -----------
# ### Thank you for completing this lab!
#
# This notebook was created by [<NAME>](https://www.linkedin.com/in/aklson/). We hope you found this lab session interesting. Feel free to contact us if you have any questions!
# This notebook is part of a course on **Coursera** called *Data Science Methodology*. If you accessed this notebook outside the course, you can take this course, online by clicking [here](https://cocl.us/DS0103EN_Coursera_LAB3).
# <hr>
# Copyright © 2018 [Cognitive Class](https://cognitiveclass.ai/?utm_source=bducopyrightlink&utm_medium=dswb&utm_campaign=bdu). This notebook and its source code are released under the terms of the [MIT License](https://bigdatauniversity.com/mit-license/).
| course_materials/course_3_-_data_science_methodology/02_from-understanding-to-preparation-and-from-modeling-to-evaluation/01_from-understanding-to-preparation/ds0103en-3-3-1-from-understanding-to-preparation-v1.0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: crystal
# language: python
# name: crystal
# ---
# # Tutorial 4: Customizing a crystal
#
# In this tutorial, we show how to customize the domains for a custom crystal so that its PMF approximates a specified target PMF.
# %load_ext autoreload
# %autoreload 2
# +
import numpy as np
import matplotlib.pyplot as plt
from custom_poling.core.target import Target
from custom_poling.core.custom_crystal import CustomCrystal
# +
# Crystal properties
domain_width = 10.0e-6
number_domains = 1000
L = number_domains * domain_width
k0 = np.pi / domain_width
# Numerical integration parameters
k_range = 100/L
dk = k_range/401
k_array = np.arange(k0-k_range/2,k0+k_range/2,dk)
# -
# ### Example 1: Sinc PMF
# Create a custom crystal object
custom_crystal_sinc = CustomCrystal(domain_width,number_domains)
domain_middles_sinc = custom_crystal_sinc.domain_middles
#Define and plot the target function
target_pmf_sinc = lambda k:-(1j*2/(np.pi**2)) * np.sin((k-k0) * L/2)*np.exp(1j * L/2 * (k-k0))/(k-k0)
target_sinc = Target(target_pmf_sinc,k_array)
target_sinc.plot_pmf()
# Compute and plot the target amplitude
target_amplitude_sinc = target_sinc.compute_amplitude(k0,domain_middles_sinc)
target_sinc.plot_amplitude()
# Compute and plot the custom domains
custom_domains_sinc = custom_crystal_sinc.compute_domains(target_amplitude_sinc,k0)
custom_crystal_sinc.plot_domains(n_max=60)
# +
# Compute the PMF for the cystomized crystal
pmf_custom_poled_sinc = custom_crystal_sinc.compute_pmf(k_array)
# Plot pmf
plt.plot(k_array,np.abs(pmf_custom_poled_sinc),label='abs')
plt.plot(k_array,np.real(pmf_custom_poled_sinc),'--',label='real')
plt.plot(k_array,np.imag(pmf_custom_poled_sinc),'--',label='imag')
plt.xlabel(r'$\Delta k$')
plt.ylabel('PMF')
plt.legend()
plt.show()
# -
# Compare the output to the target
plt.plot(k_array,np.abs(target_sinc.pmf))
plt.plot(k_array,np.abs(pmf_custom_poled_sinc))
plt.show()
# ### Example 2: Gaussian PMF
# +
# Create a custom crystal object
custom_crystal_gauss = CustomCrystal(domain_width,number_domains)
domain_middles_gauss = custom_crystal_gauss.domain_middles
#Define and plot the target function
std = 10/L
height = 0.00025
target_pmf_gauss = lambda k:1j*height*np.exp(-(k-k0)**2/(2*std**2))*np.exp(1j * L/2 * k)
# target_pmf_gauss =lambda k:-(1j*2/(np.pi**2)) * np.sin((k-k0) * L/2)*np.exp(1j * L/2 * (k-k0))/(k-k0)
target_gauss = Target(target_pmf_gauss,k_array)
target_gauss.plot_pmf()
# Compute and plot the target amplitude
target_amplitude_gauss = target_gauss.compute_amplitude(k0,domain_middles_gauss)
target_gauss.plot_amplitude()
# Compute and plot the custom domains
custom_domains_gauss = custom_crystal_gauss.compute_domains(target_amplitude_gauss,k0)
custom_crystal_gauss.plot_domains()
# Compute the PMF for the cystomized crystal
pmf_custom_poled_gauss = custom_crystal_gauss.compute_pmf(k_array)
# Plot pmf
plt.plot(k_array,np.abs(pmf_custom_poled_gauss),label='abs')
plt.plot(k_array,np.real(pmf_custom_poled_gauss),'--',label='real')
plt.plot(k_array,np.imag(pmf_custom_poled_gauss),'--',label='imag')
plt.xlabel(r'$\Delta k$')
plt.ylabel('PMF')
plt.legend()
plt.show()
# + tags=[]
# Compare the output PMF to the target PMF
plt.plot(k_array,np.abs(target_gauss.pmf),label='Target PMF')
plt.plot(k_array,np.abs(pmf_custom_poled_gauss),label='Custom PMF')
plt.legend()
plt.show()
# Compute amplitudes
custom_amplitude,z_list= custom_crystal_gauss.compute_amplitude(k0,num_internal_points=1)
# Compare the output amplitudes to the target amplitudes
plt.plot(z_list,np.abs(custom_amplitude),label='Custom amplitude')
plt.plot(custom_crystal_gauss.domain_middles,np.abs(target_gauss.amplitude),label='Target amplitude')
plt.legend()
plt.show()
# -
| Tutorial_4_customizing_a_crystal.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# # !wget https://raw.githubusercontent.com/huseinzol05/Malaya-Dataset/master/dictionary/dialect/kelantan.csv
# +
import pandas as pd
kelantan = pd.read_csv('kelantan.csv')
# +
import malaya
malays = malaya.texts._malay_words._malay_words
# +
import re
from unidecode import unidecode
def cleaning(string):
string = unidecode(string).replace('.', '. ').replace(',', ' , ')
string = re.sub('[^\'"A-Za-z\-/ ]+', ' ', string)
string = re.sub(r'[ ]+', ' ', string.lower()).strip()
return string
# -
words = []
for i in range(kelantan.shape[0]):
try:
words.extend(cleaning(kelantan['0'].iloc[i]).split())
except:
pass
# +
# http://w3hafizm.blogspot.com/2010/11/kamus-kelantan-loghat-kelate.html
additional = """
agah / hagah – sombongan
api stok(api stop)- traffic light
api stop -traffic light
asore bodi – insuran kereta / motora
awe – panggilan manja kepada orang lelaki
awe sejambo lebak - satu watak dalam drama main tteri
basah jjerok – basah kuyup / lecun
bbageh – kaedah pengubatan tradisi / jampib
baloh – berkelahi
bbini – berkawinb
bojeng – guting rambutb
borak – merebak
bedua (sebutan bunyi dihidung) -rasa ingin marah / benci
bekwoh – kenduri (mungkin berasal dari perkataan Inggeris – big work)
belabik – (siapa tahu tolong jelaskan)
belengah – melekit
betak – kenyang
betap wak – lembap
biru kketu – biru yang amat sangat
blebe – berleter
bocah / boceh – buncit
bojeng – fesyen rambut (contoh: bojeng 1-2 = rambut hampir botak)
bok – buku (asal perkataan Iggeris – book)
bokali – barangkali
bokbong – cempelai (spesis musang)
brona – rosak / budak yang menangis tak berhenti
buah spelek – sesuatu (teknik) yg istimewa
buah topoh – buah epal (diambil dr perkataan arab “tuffah” = epal)
buah zabik – kismis
buat nyayo – menyiksa
buje / oghe buje – janda
busuk banga – terlalu busuk
busuk kohong – terlalu busuk
butak – perut buncit
ca – air yang bertakung di bawah atau tepi rumah
cah keting – bentes
cok – cangkul
ccerok – perut berbunyi bila lapar
cceruk – potong rumput untuk haiwan
cebok – cedok (selalunya untuk cecair, mencedok)
cema clote / cpolok – terlalu kotor
cepelak – lancang
cerah craca – terang benderang
cerah perut – cirit birit
cliko – curi tulang / pemalas
cokek makbolok – cucuk pinggang kawan dengan jari telunjuk dgn tujuan bergurau
comel llote – amat cantik
cuwoh – curah
dale so – pusat hiburan di Kota Bharu zaman dulu (asal perkataan Inggeris – show)
ddasing – menari (asal perkataan Inggeris – dancing)
dderak / bederak – suka merayau / berjalan
debek – teruk (asal perkataan Inggeris – “the bad”)
deh? – memohon restu atau persetujuan, lebih kurang “ok?”,
dekpong gak eh – kalau ya pun
dermo basikal – asal perkataan Inggeris -”dynamo”
dok cckoh – duduk bercangkung
dok kene (sebutan English – doc care nay) – ya bukan / “isn’t it”
dok ko? – ya tak?
dok? – lebih kurang “bukan?”, “betul tak?” atau “isn’t it?”
duga / luga – tak sedap perut /loya (sebutan ikut daerah)
gaduh – nak cepat
gak – lebih kurang “habis tu”
gdebe – berani / samsing
gege – bising / riuh rendah biasanya ada bunyi ketawa (asal perkataan Inggeris – giggle) – contohnya: jange ggege gak! = don’t giggle! Please
gelebek – biasanya terjadi pada mata pisau/parang bila kerat benda keras (tumpul gelebek – sangat tumpul).
gelembong boya – kueh daripada beras pulut (species dodol)
gelenyar / gletah – merenyam, getik
gelega – lantai
gemuk ddebok – terlalu gemok
genyeh – tenyeh
geretak – jambatan
getah sokmo – kueh daripada ubi keledek
getek – juga
ggapo – apa
ggatih teksi – kayuh beca
ggocoh – bertumbuk
ggoghi – bagi memulakan sesuatu
ghak – semak
ghohok = sukar, susah, payah
goba = risau
gong – tolol
gonyoh – gosok dgn kuat
griak – kahak
guano – lagu mana / macam mana
hapok kohong – bau hampak
hnja – tendang
ho (sebutan bunyi dihidung) – “ya lah..” atau “yes”
honda samah – honda cub 50cc
hungga – berlari
ike kkhonge – ikan cencaru
istek – ladang (asal perkataan Inggeris – estate)
jamah – pegang / sentuh
jatuh celabok – jatuh berteraburan
jebat – bau yang dikeluar oleh binatang untuk menanda kawasannya (contoh: jebat musang) – kepada orang Melaka… maaf lah ya
jebbeng – berjambang
jebeh – mencebek
jebo – botol kaca
jelira – sedap / kena dgn slera
jellaq – tamak / orang kepalaparan dapat makanan
jemba – berjumpa / sua
jemeleh – sembelih (contoh: tak jemeleh ko? = tak sembelih lembu ke? )
jemore – lantai basah, gelegar bulu atau batang pinang – rumah kampung dulu-dulu
jenera – lena
jerkoh – sergah
jjolor – menjulur
jjughuh = baik (jjughuh budok tu = baik budak itu)
jong kako = tukang angkat hidangan makan / penanggah)
kabil – salah satu teknik mendayung sampan
kaki sbaye – kuku jari kaki yg rosak dan berbau
karya basikal – tempat letak barang di belakang basikal (asal perkataan Inggeris – “carrier”)kasut bok = kasut jenis “boot”
kayae – bisanya digunakan untuk tanda kawasan rumput yang nak disabit
kdolok – lawak
kebek – membuka ruang menjadi lebih besar
kecek – pujuk / goda
kekoh – gigit
kelaring – kotor (asal perkataan Inggeris – colouring)
kelong/belong – tipu muslihat
kelorek – kedekut
kenye bboyah – terlalu kenyang
kepek idung - kueh puteri mandi
kereta plek – kereta sewa / teksi
kerlong – greedy / tenong
kesit – sunyi (lonely)
ketik – gigit kecil
ketik ttunga – kurus / kecil serta pucat
ketok bodi – buat / baiki badan kenderaan
kkecek – bercakap
kketei – kantin
klikpah-klikpah – terpinga pinga
kodi – tidak berkualiti
ko’o – ketawa berdekah-dekah
kota – cukai jalan atau insurans (contoh: kreta kawe tak dok kota )
kuca hanya / kuca lana – berteraboran
kuda – sepak
kuk / kok – sekeh
kuning nnehe / llehe – terlalu kuning
kupik – kedekut
lamoke – nanti kan
lari kecik ppala-ppala – lari terlalu laju / lintang pukang
lecah – payau atau becak (contoh: “toksoh lah awe…lecah” maksudnya ” tak usah lah ya … payau”
leweh – kurang solid
lipotei – tidak tetap duduk / ke sana ke sini
lobey – gila-gila atau bengong
loleh – tak serious
lorong tua (sebutan bunyi dihidung) – kawasan pelacuran di Kota Bharu zaman dulu (dah tak ada lagi sekarang ni)
mahkamah tinggi ayoh kob – mahkamah tinggi high court
main tteri – drama tari pengubatan tradisi
mamba – tok segar / tegar
manih lleting – terlalu manis
mase ppughik – terlalu masam
masin ppeghak – terlalu masin
mek – panggilan manja untuk orang perempuan
merket – pasar (asal perkataan Inggeris – market)
mmeda – buang air besar
mmupo – mandi sungai
mokte – rambutan
ngga – tolol
nghele – menghadiri kenduri
ngidung – sengau
ngusuk – terakhir
nnakut / penakut apah – terlalu penakut
nnate – binatang (kadangkala “simbol” kemesraan antara kawan)
nnawak – bohong
nneja – pengurus (asal perkataan Inggeris – Manager)
nneting – melantun
nngapo – meracaun
nungei nyor – “somersault” – kadang-kadang dibuat perumpamaan frust nungei nyor (frust somersault la tu!!)
nnusuk – sembunyi (main nnusuk – main hide and seek)
nnyaba – tak kuat
nnyaca – terhuyung hayang nak jatuh
nok ssega angin – metodologi pengubatan cara tarditional contohnya main puteri
nyace – kayu atau besi yang dipacak ke tanah..tambak lembu
nyapong – carut
nyayo – kesian
nyior koter – kelapa tua
pah? – lepas itu?
pahit llepe – lerlalu pahit
pakddahak – tanda silang atau pangkah
papok – bapok / pondan
patat siput – kemahiran (skill) contoh: tak dak patat siput – maknanya tak ada kemahiran
pecah peda – kentut
pekong – baling
pelepong lembu – paru-paru lembu
penampa tawa – penampar percuma
pengah – gedik / getik (greedy)
perone – tempat membuang / bakar sampah
perut besar – mengandung
petong – baling
pitu gek – pintu pagar (gate)
plungo – kayu atau besi tajam untuk tambat lembu
pok ko – spesis mengkarung tapi boleh memanjat
pozek – bayar muka (asal perkataan Inggeris – deposit)
ppala bakul – cukai yang dikenakan oleh majlis bandaran ke atas peniaga
ppala bubus – kepala botak
ppatak – paling bawah
ppiyah – ketayap / kopiah
prebet sapu – teksi sapu
prekso – peperiksaan
pungga – baling
putih ssueh – terlalu putih
putung kalong – batang kayu untuk dibaling
ralek – leka
rauk muka – sapu muka
redas – cakap laser / baling
rhoyat – bagitahu / maklumkan
rhukah – panjat
rima – harimau
rizat – keputusan (asal perkataan Inggeris – result)
roba – getah pemadam ( asal perkataan Inggeris – rubber)
sa – satu
sabik – sebab
saing – kawan
saksoba – penyerap hentak (asal perkataan Inggeris – sock absorber)
samah – 50 sen
samah seghia – pendapatan kecil untuk sara anak isteri
saru – serabut
satu sut – berpakaian lengkap
seghia dua - pendapatan kecik untuk sara anak isteri
seh inguh – hembus hingus
selareh – selalu
sele-bele – tak kemas / cuai
sengeleng – sengaja
senyap tipah – tak ada khabar berita
sgeto – kawasan “keras” – ada penunggu
sghia – RM1.00
sia – sembuh dari luka (asal perkataan Inggeris – seal)
sleke – silakan
smaye ssejid – sembahyang di masjid
smeesek – terlalu mudah
smuta – kain lilit kepala
sobek – hias
sokmo – sentiasa
sopeh – serpih
ssong – sesuai (tak ssong – tak sesuai)
ssumba – pewarna untuk makanan/minuman
suku – berasingan (bungkus suku – bungkus asing-asing)
supik – beg plastiksupik gelenyar /supik rhokrhak – beg plastik yang nipis
suwih – swis (asal perkataan Inggeris – switch)
tak cakno – tak hirau
tak mmado – tak peduli / tak padan
tak pok – tak cukup cerdik
tak ppaka – tidak terpakai (tak pakka benda – tak menjadi)
tak rak – tak mampu
tanggong – lebih kurang “ssong” – sesuai
tawar heber – terlalu tawar
tepoh – langgar
tohok – buang
tok bageh – kepala upaca bbageh
tok ggawa – ketua daerah
tok kerani – kerani di pejabat
tok laki – suami
tok mindok – tukang gesek rebab main tteri atau mok yong
tok nebeng – ketua kampung
tok nngulu – ketua mukim
tok peraih – berniaga kecil-kecilan di pasar
tok ppeti – mufti
tok tteri – kepala upacara main puteritonye – ejek dgn memek muka
ttino – betina / perempuan (oghe ttino kawe = isteri saya)
ttino garik – perempuan jalang ( disebut bila dalam keadaan marah)
ttuyup – pepatung
tubik – keluar
tuke ttesen – tukang tulis / taip petisyen
tunja – tendang
turik – berdesing pendengaran – sakit hati, marah
wak nganyi – perli / ejek dgn kata-kata
wok lor – tolol
yak!! – lebih kurang “opocot!!!” atau “oops!!”
"""
# -
for line in additional.split('\n'):
if not len(line):
continue
c = cleaning(unidecode(line).split('-')[0]).split('/')
words.extend([cleaning(i) for i in c])
# +
# https://www.facebook.com/AnakPerantauanKelate/posts/kamus-loghat-kelantanacu-tra-teh-cubaagah-berlagakalik-sebelahambak-kejarambo-sa/1109775215739242/
additional = """
acu tra teh - cuba
agah - berlagak
alik - sebelah
ambak - kejar
ambo - saya
apung - apam balik
abe-abang
apo-apa
baju gumbea - baju melayu
bakpo - kenapa
bbera - berpecah
bberong - crowded
bBobok - meracau
bbol - mentol
beca - silau
bedo oh - melampau
beg duit - dompet
behe - pukul dari sisi
bejebok - limpah
bekeng - garang
bekwoh - kenduri
berat - teruk
bidah - lastik
bim - tidur utk baby
biru dacing - biru bendera umno
blana kokna - sangat banyak
bledo - Agar-agar
bo laa - sudah la
bocong - botol air
boh (boss) - ayah
bokbong - baabon
boktea - dibuat oleh
bolok - selekeh
brona - bermasalah
buah setow - buah manggis
buoh badminton - bulu tangkis
buoh lanah - nenas
buoh pauh - mangga
buoh sawo nilo - ciku
buoh tterea - jagus
busuk banga - sangat busuk
cakkelat - gula-gula
celeng - tabung syiling
.... dale duo lagi – tak pasti lagi
deghak – bersiar-siar
dekcok – main teng-teng
demo – kamu/awak
denu – di sana
do..oh - melampau
dok alik mano? – berada di mana?
dok cace - berdiri
dok chongok – duduk bersila
dok jjerung mano – duduk belah mana?
dok ttinggung - bertinggung
drebar – memandu;pemandu
etek – juga
gaha - berus
gamok - agak
gedebe/ddebe - samseng
gegea glegok – bising sangat
gewe - awek
ggaji – gergaji
ggilo - minat
ggocoh – gaduh
gguling bating – berguling-guling
ghanggoh – makan
gheknge nnapung – sangat ringan
ghoyak - beritahu
ghukah - panjat
glecoh - terseliuh
glenya – mengada-ada
glepar - menggelepar
glewak - menyibuk
gligo - permata
godio – apa dia
golok - parang
gongok – gigi rosak
gonyoh - berus
gostae - undur ke belakang
gouk - reban/kandang
ggorek - pengasah pensel
gu - kawan
guano – bagaimana;tanya keadaan
gumbo air – sedut air
hija pah – hijau bendera PAS
hite legea – terlalu hitam
hoo - ya
huduh sepa – sangat hodoh
igak - tangkap
ikut ah – suka hati la
ipung - kumpul
jambe - tandas
jangok – melaram
jauh jelak – sangat jauh
jebeh – masam muka
jebour – botol
jenero - tertidur
jjerik – menangis
jjolo – terjulur
jok - sejak
jjujuk - tersusun panjang
jo ong - mendung
jolo – betul la tu
kaba-kaba - sedar-sedar/tiba-tiba
kain klubung – kain tudung
kain smaye – kain telekung
kaing sehe – kain basahan
kalea - pensel
katok – puku dari atas
kawe - saya
kecek - bercakap
keheak - ludah
kerek - perigi
keghah kekong - keras yg teramat koho/selo - perlahan/slow
kekoh – gigit
kelik - balik
khapoh - sampah
khepok gote - keropok lekor
khetah - kertas
kheto - kereta
khiput - berkedut
khobek - selongkar
khusi - kerusi
kito – saya(kata ganti nama bg orang muda terhadap org tua)
kkacik - sangat rapat
kkesok - terkalih
kkorea – pengisar kelapa
kkubey - berselerak
kokok – sodok
kokse kokdea – kucar kacir
kona - membelok
koya/agas - perasaan
kube – terabur
kuey kekoh cha – buah melaka
lagu mano - bagaimana
lembik – lemah
li-lah2 – ke sana ke mari
liut – lemau
loghat - nak cepat
lokpak tikea – nama sejenis makanan
lok - biarkan
lugar – loya
liyk - elak
main ceklat – main helah
manggok – bangga diri
manih lleting – terlalu manis
mase purik – terlalu masin
masin perak – terlalu masin
mbek – kambing (panggilan untuk tarik perhatian kanak2)
meroh merea – merah darah
metoo - degil
minyok mati bbunuh - minyak dagu
mmakoh - berhujah
mmapuh - kenakan
mmecok - merajuk
mmeteak – kacau; ganggu
mmolek – buat elok-elok
mmunoh – buat kerosakan
mmutar itik - mencari sesuatu dengan gelabah
moktea - rambutan
monggek - bonceng
mugo – panggilan utk sesuatu yg tidak diketahui namanyo
mung - engkau
nah – menyatakan banyak
namo rima? – apa dia tu?
ndow - buaian
ngaji - belajar
ngaju - merajuk
ngakok – merangkak
ngala – mewarnakan
nganying - mempersenda
ngejah - negotiate
ngekkoh – akikah
ngepek – berleter
nguraa – conteng
nnakak – tertarik perhatian
nnarak kow – berlebih-lebihan
nnawok - menipu
nnebah - menebas
nnepik - menjerit
nnerak - meniarap
nnetea – melentang
nneting - melantun
nnise – gula melaka
nnoney - bergayut
nnonye – mencebik muka
nokpak - melompat
nukih - melukis
ook-aloh – menyatakan keluhan
otokk - tertekan
panje jolo - panjang sangat
parowk - teruk
patak-bawah
penar-silau
pe'ea - perangai
pekdoh - faedah
pekong - baling
peseng – style
peti sejok - peti ais
pitih-duit/wang
pleting - straw
pparowk - selisih
pok daro nitas-polis di depan
pok daro-pakcik
ppena - silau
ppioh kopiah
puah ea lokk – bosan
punoh - rosak
putih sepow – sangat putih
putihstar - purple
putik lingo – pengorek telinga
relaa - tercerai
roba- pemadam
rrobok - almari
rukah – panjat
sedho – kurang baik
segho - terasa
segowk - menanduk
sek – geng;kumpulan
selok - pengsan
seney mesek – sangat senang
senyap ttipah – terlalu senyap
sero – rase, agak2
siga – tangga
singo beng - air limau air
ssika bmx - basikal BMX
smeta - sekejap
skali Arung - Sekaligus
sejuk Ketta – sangat sejuk
Smuta – kain serban
srebea - kebas
Ssakok - tersangkut
ssaloh kaki – terseliuh kaki
ssekong - kaku
ssikal - basikal
ssiko – penyodok
ssoyok – terkoyak
stambung - bertimbun
sugho - bubur asyura
suko selok - ketawa terbahak-bahak
sungguh – kata penguat bagi sifat:comel sungguh
siak dok ggobar - Jangan risau
saksoba - Absorber
tape – rakan suara
tawar hebea – sangat tawar
temo'o – nak sangat
tino lawa-wanita cantik
timu cino - tembikai
tohor - cetek
tohouk - buang
tok bbutir- tak jelas
tok cekak – tak larat
tok kaba - tak sedar
toksoh - jangan / tak usah
trelak – tidur sekejap
triok – nangis
ttala - tertangguh
ttbolah – tidak berhati-hati;cuai, memalukan
ttebeng – ambil risiko
tteke - tertekan
ttumbok – bertumbok
ttuyup - pepatung
tok cakno - tak hirau
tepoh - langgar
tubik - keluar
taik Aye Munea - Tahi Ayam Yang Masih Hangat
ubi stelo - ubi keledek
vi-bagi
wak gapo-buAt apa
woh nyo-buah kelapa
yak boter-ye la tu
"""
# +
for line in additional.split('\n'):
if not len(line):
continue
c = cleaning(unidecode(line).split('-')[0]).split('/')
words.extend([cleaning(i) for i in c])
len(words)
# +
words = [i for i in words if len(i) > 3]
words = set(words) - malays
len(words)
# +
import json
with open('kelantan-words.json', 'w') as fopen:
json.dump(list(words), fopen)
| session/language-detection/kelantan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# -*- coding: utf-8 -*-
# Copyright (c) 2015 <NAME>
import datetime
import json
import os
import matplotlib
import matplotlib.pyplot as plt
import pandas as pd
# -
# ## Load Data for PyLadies and their local Python User Groups
DATA_DIR = "meetup_data"
MEMBER_JSON = "pug_members.json"
GROUP_DIRS = [d for d in os.listdir(DATA_DIR)]
PYLADIES_GROUPS = []
for group in GROUP_DIRS:
if os.path.isdir(os.path.join(DATA_DIR, group)):
PYLADIES_GROUPS.append(group)
def load_group_data(pyladies_group):
pyladies_dir = os.path.join(DATA_DIR, pyladies_group)
members_json = os.path.join(pyladies_dir, MEMBER_JSON)
with open(members_json, "r") as f:
return json.load(f)
# #### Get date for when the PyLadies group was started
def pyladies_created(pyladies_group):
pyladies_dir = os.path.join(DATA_DIR, pyladies_group)
pyladies_file = os.path.join(pyladies_dir, pyladies_group + ".json")
with open(pyladies_file, "r") as f:
data = json.load(f)
created_ms = data.get("created") # in ms after epoch
created_s = created_ms / 1000
created = datetime.datetime.fromtimestamp(created_s)
year = created.strftime('%Y')
month = created.strftime('%m')
day = created.strftime('%d')
return year, month, day
# ## Create some dataframes in Pandas
#
# **WARNING** I do not know how to properly panda. This is my hacky attempt.
# helper function to create a dataframe out of multiple data frames
# one DF per PUG
def _create_dataframe(group, data):
df = pd.read_json(json.dumps(data))
joined = df.loc[:,("id", "joined")]
joined["joined"] = df.joined.apply(lambda x: pd.to_datetime([x], unit="ms"))
joined["mon"] = joined.joined.apply(lambda x: x.month[0])
joined["year"] = joined.joined.apply(lambda x: x.year[0])
agg_joined = joined.groupby(["year", "mon"]).count()
return agg_joined
def collect_dataframes(group_data):
dfs = []
for group in group_data.keys():
data = group_data.get(group)[0]
df = _create_dataframe(group, data)
tmp = {}
tmp[group] = df
dfs.append(tmp)
return dfs
# aggregate dataframes, name columns nicely, etc
def aggregate_dataframes(dfs):
first = dfs.pop(0)
name = first.keys()[0]
_df = first.values()[0]
df = _df.loc[:, ("id", "joined")] # multi indices are hard.
df.rename(columns={"joined": name}, inplace=True)
for d in dfs:
name = d.keys()[0]
_df = d.values()[0]
df[name] = _df["joined"]
df = df.fillna(0)
df.drop('id', axis=1, inplace=True)
return df
# ## Now let's graph
# helper function for x-axis labels
def _get_x_axis(current):
updated = []
for item in current:
_date = item.get_text() # u'(2009, 2)'
if _date == "":
updated.append(_date)
else:
_date = _date.strip("(").strip(")").split(",") # [u'2009', u' 2'] # NOQA
_date = [d.strip() for d in _date] # [2009, 2]
label = "{0}-{1}".format(_date[1], _date[0])
updated.append(label)
return updated
# helper function to plot created date annotation
def _created_xy(df, created):
year, month, _ = created
indexlist = df.index.tolist()
created_index = indexlist.index((int(year), int(month)))
return created_index
# helper function to position the annotated text
def _find_max(df, groups):
maxes = [df[g].max() for g in groups]
return max(maxes)
def create_graph(df, pyladies, created, groups):
created = _created_xy(df, created)
created_yarrow = int(round(_find_max(df, groups) * .80))
created_ylabel = int(round(created_yarrow * .80))
graph = df.plot(figsize=(17, 8), linewidth=4)
graph.set_title(pyladies)
xlabels = _get_x_axis(graph.get_xticklabels())
graph.set_xticklabels(xlabels, rotation=45)
graph.set_xlabel("Month")
graph.set_ylabel("# of members joined")
for i, line in enumerate(graph.get_lines()):
line.set_linewidth(3)
graph.legend() # update legend with line weight changes
graph.axvline(x=created, ymin=0.0, ymax=1.0, linewidth=4)
graph.annotate("PyLadies Created", (created, created_yarrow),
xytext=(created - 8, created_ylabel), xycoords="data",
arrowprops=dict(arrowstyle="->", facecolor='black', linewidth=3))
# needs `%matplotlib inline`
plt.show()
# if you'd rather save the graph as an image locally
# output = os.path.join(self.pyladies_dir, pyladies + ".png")
# plt.savefig(output, dpi=300)
# ## To put it all together now
#
# I'm only going to do a handful, not all the groups. But this should give you an idea.
#
# Also - some shit breaks. Surely I'll fix it in my Copious Amount of Free Time™ :D
# %matplotlib inline
def create_pyladies_graph(pyladies):
group_data = load_group_data(pyladies)
created = pyladies_created(pyladies)
dfs = collect_dataframes(group_data)
df = aggregate_dataframes(dfs)
create_graph(df, pyladies, created, group_data.keys())
create_pyladies_graph(PYLADIES_GROUPS[2])
create_pyladies_graph(PYLADIES_GROUPS[11])
create_pyladies_graph(PYLADIES_GROUPS[14])
create_pyladies_graph(PYLADIES_GROUPS[15])
create_pyladies_graph(PYLADIES_GROUPS[19])
create_pyladies_graph(PYLADIES_GROUPS[22])
create_pyladies_graph(PYLADIES_GROUPS[28])
create_pyladies_graph(PYLADIES_GROUPS[30])
create_pyladies_graph(PYLADIES_GROUPS[39])
| Creating Graphs with Pandas & matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Sequence Data Analysis with Python
# ## Agenda
# 1. Validate Sequence
# 2. Counting Letters / Frequency
# 3. % of Nucleotides
# 4. Calculating `GC` Content
# 5. `GC` Content of Sub-sequence
# 6. Calculating `AT` Content
# 7. `AT` Content of Sub-sequence
# 8. Transcription
# 9. Complement
# 10. Reverse Complement
# 11. DNA Translation
seq = 'GGTCAGAAAAAGCCCTCTCCATGTCTACTCACGATACATCCCTGAAAACCACTGAGGAAGTGGCTTTTCA'
# ## Validate Sequence
# counting letters
seq.count("A")
# length of seq
len(seq)
# % of nucleotide
(seq.count("A") / len(seq)) * 100
def validtae_seq(seq):
"""Checks if DNA sequence is valid. Returns True is sequence is valid, or False otherwise"""
seq = seq.upper()
nt_counts = seq.count("A") + seq.count("T") + seq.count("G") + seq.count("C")
if nt_counts == len(seq):
return True
else:
return False
# function call
validtae_seq(seq)
import screed
def readFASTA(inputfile):
"""Reads a FASTA file and returns with special characters removed!"""
# open file in reading mood
with screed.open(inputfile) as seqfile:
for read in seqfile:
seq = read.sequence
return seq
# data read
seqs = readFASTA("../data/Haemophilus_influenzae.fasta")
validtae_seq(seqs)
# ## Counting Letters / Frequency
seq = 'GGTCAGAAAAAGCCCTCTCCATGTCTACTCACGATACATCCCTGAAAACCACTGAGGAAGTGGCTTTTCA'
base_counts = {'A': 0, 'T': 0, 'G': 0, 'C': 0}
for i in seq:
base_counts[i] += 1
base_counts
base_counts.keys()
base_counts.values()
base_counts.items()
letters = []
values = []
for k, v in base_counts.items():
letters.append(k)
values.append(v)
letters
values
(21/70) * 100
seq = 'GGTCAGAAAAAGCCCTCTCCATGTCTACTCACGATACATCCCTGAAAACCACTGAGGAAGTGGCTTTTCA'
def basecount(seq):
"""Read sequence and returns their frequency."""
# store frequency
base_count = {}
for base in seq:
if base in base_count:
base_count[base] += 1
else:
base_count[base] = 1
return base_count
basecount(seq)
# %%timeit
basecount(seqs)
from collections import Counter
freqs = Counter(seq)
freqs
from collections import Counter
freqs = Counter(seqs)
# %%timeit
freqs
from collections import Counter
def count_fast(seq):
"""Reads sequence and returns their frequency"""
freqse = Counter(seq)
return freqs
# %%timeit
count_fast(seqs)
basecount(seqs[1:500])
basecount(seqs[500:1001])
def pc_freq(seq):
"""Reads sequence and returns their %"""
| old_notebooks/2. Basic Processing of Biological Sequences with Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
mensagem = 'Jupyter Notebook - boa ferramenta!' # var. string no Python
# !echo '$mensagem\n\r$mensagem'
# !dir
# !cd
# + language="html"
# <marquee style='width: 30%; color: blue;'><b>Olá Mundão!</b></marquee>
# + language="html"
# <svg xmlns="http://www.w3.org/2000/svg"
# viewBox="0 0 450 400" width="200" height="200">
# <rect x="80" y="60" width="250" height="250" rx="20"
# style="fill:red; stroke:black; fill-opacity:0.7" />
# <rect x="180" y="110" width="250" height="250" rx="40"
# style="fill:blue; stroke:black; fill-opacity:0.5;" />
# </svg>
# +
import numpy as np
# np.random?
# -
| 01-Sinais/Comandos_do_SO.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import torch
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
from scripts.data import Project
from scripts.util import flatten_with_index, word_list_regex
import nltk
project = Project(model_name='bert-base-uncased', subspaces=['gender'])
subspace = project['gender']
pca = PCA(n_components=2)
base_model = project.get_bert().eval()
debiased_model = torch.load(f'data/{project.model_name}_debiased.pt', map_location=project.device).eval()
kaneko_model = torch.load(f'data/{project.model_name}_kaneko.pt', map_location=project.device).eval()
base_model = project.get_bert().eval()
# +
target_tokenizer = nltk.RegexpTokenizer(word_list_regex([a.word for a in subspace.targets.all]), gaps=True).tokenize
attribute_tokenizer = nltk.RegexpTokenizer(word_list_regex([a.word for a in subspace.attributes.all]), gaps=True).tokenize
def encode(sentence: str, tokenizer):
parts = tokenizer('[CLS] ' + sentence + ' [SEP]')
target = parts[1]
parts = project.tokenizer.batch_encode_plus(parts,
add_special_tokens=False,
return_attention_mask=False,
return_token_type_ids=False)['input_ids']
encoded, indices = flatten_with_index(parts)
return torch.IntTensor(encoded), (indices[1], indices[2]), target
def encode_parts(start_sent, target, end_sent):
parts = project.tokenizer.batch_encode_plus([f'[CLS] {start_sent} ', target, f' {end_sent} [SEP]'],
add_special_tokens=False,
return_attention_mask=False,
return_token_type_ids=False)['input_ids']
encoded, indices = flatten_with_index(parts)
return torch.IntTensor(encoded), (indices[1], indices[2]), target
def process_parts(model, parts):
encoded = [encode_parts(s0, s1, s2) for s0, s1, s2, _ in parts]
padded = max(len(t) for t, _, _ in encoded)
input_ids = torch.zeros(len(encoded), padded, dtype=torch.int32)
attention_mask = torch.zeros_like(input_ids, dtype=torch.int32)
for i, (t, _, _) in enumerate(encoded):
size = len(t)
input_ids[i, :size] = t
attention_mask[i, :size] = 1
# shape: (sents, sent_length, embedding_size)
model_out = model(input_ids=input_ids.to(project.device), attention_mask=attention_mask.to(project.device))['last_hidden_state']
result = []
for i, (_, (j, k), _) in enumerate(encoded):
result.append(model_out[i, j: k].mean(dim=0))
# shape: (sents, 2)
result = pca.fit_transform(torch.stack(result).detach().cpu().numpy())
df = pd.DataFrame(columns=['PC1', 'PC2', 'type'])
for i, (s0, s1, s2, _type) in enumerate(parts):
df = df.append({'PC1': result[i, 0], 'PC2': result[i, 1], 'type': _type}, ignore_index=True)
plt.text(result[i, 0], result[i, 1], f'{s0} {s1.upper()} {s2}')
sns.scatterplot(data=df, x='PC1', y='PC2', hue='type', legend=False)
def process_parts_sim(model, parts):
encoded = [encode_parts(s0, s1, s2) for s0, s1, s2, _ in parts]
padded = max(len(t) for t, _, _ in encoded)
input_ids = torch.zeros(len(encoded), padded, dtype=torch.int32)
attention_mask = torch.zeros_like(input_ids, dtype=torch.int32)
for i, (t, _, _) in enumerate(encoded):
size = len(t)
input_ids[i, :size] = t
attention_mask[i, :size] = 1
# shape: (sents, sent_length, embedding_size)
with torch.no_grad():
model_out = model(input_ids=input_ids.to(project.device), attention_mask=attention_mask.to(project.device))['last_hidden_state']
result = []
for i, (_, (j, k), _) in enumerate(encoded):
result.append(model_out[i, j: k].mean(dim=0))
return torch.stack(result)
def process(model, sentences, tokenizer):
encoded = [encode(s, tokenizer) for s in sentences]
padded = max(len(t) for t, _, _ in encoded)
input_ids = torch.zeros(len(encoded), padded, dtype=torch.int32)
attention_mask = torch.zeros_like(input_ids, dtype=torch.int32)
for i, (t, _, _) in enumerate(encoded):
size = len(t)
input_ids[i, :size] = t
attention_mask[i, :size] = 1
# shape: (sents, sent_length, embedding_size)
model_out = model(input_ids=input_ids.to(project.device), attention_mask=attention_mask.to(project.device))['last_hidden_state']
result = []
for i, (_, (j, k), _) in enumerate(encoded):
result.append(model_out[i, j: k].mean(dim=0))
# shape: (sents, 2)
result = pca.fit_transform(torch.stack(result).detach().cpu().numpy())
df = pd.DataFrame(columns=['PC1', 'PC2', 'type'])
for i, s in enumerate(sentences):
df = df.append({'PC1': result[i, 0], 'PC2': result[i, 1], 'type': i%2}, ignore_index=True)
plt.text(result[i, 0], result[i, 1], s)
sns.scatterplot(data=df, x='PC1', y='PC2', hue='type', legend=False)
def process2(models, names, sentences, tokenizer):
encoded = [encode(s, tokenizer) for s in sentences]
padded = max(len(t) for t, _, _ in encoded)
input_ids = torch.zeros(len(encoded), padded, dtype=torch.int32)
attention_mask = torch.zeros_like(input_ids, dtype=torch.int32)
for i, (t, _, _) in enumerate(encoded):
size = len(t)
input_ids[i, :size] = t
attention_mask[i, :size] = 1
# shape: (sents, sent_length, embedding_size)
df = pd.DataFrame(columns=['PC1', 'PC2', 'type'])
result = []
for m, model in enumerate(models):
model_out = model(input_ids=input_ids.to(project.device), attention_mask=attention_mask.to(project.device))['last_hidden_state']
for i, (_, (j, k), _) in enumerate(encoded):
result.append(model_out[i, j: k].mean(dim=0))
# shape: (sents, 2)
result = pca.fit_transform(torch.stack(result).detach().cpu().numpy())
c = 0
for g in range(len(models)):
for i, s in enumerate(sentences):
df = df.append({'PC1': result[i + c, 0], 'PC2': result[i + c, 1], 'type': names[g]}, ignore_index=True)
plt.text(result[i + c, 0], result[i + c, 1], s)
c += len(sentences)
sns.scatterplot(data=df, x='PC1', y='PC2', hue='type', legend=True)
def process3(model, targets, attributes):
encoded = [encode(s, target_tokenizer) for s in targets] + [encode(s, attribute_tokenizer) for s in attributes]
padded = max(len(t) for t, _, _ in encoded)
input_ids = torch.zeros(len(encoded), padded, dtype=torch.int32)
attention_mask = torch.zeros_like(input_ids, dtype=torch.int32)
for i, (t, _, _) in enumerate(encoded):
size = len(t)
input_ids[i, :size] = t
attention_mask[i, :size] = 1
# shape: (sents, sent_length, embedding_size)
model_out = model(input_ids=input_ids.to(project.device), attention_mask=attention_mask.to(project.device))['last_hidden_state']
result = []
for i, (_, (j, k), _) in enumerate(encoded):
result.append(model_out[i, j: k].mean(dim=0))
# shape: (sents, 2)
result = pca.fit_transform(torch.stack(result).detach().cpu().numpy())
df = pd.DataFrame(columns=['PC1', 'PC2', 'type'])
for i, s in enumerate(targets):
df = df.append({'PC1': result[i, 0], 'PC2': result[i, 1], 'type': 'targets'}, ignore_index=True)
plt.text(result[i, 0], result[i, 1], s)
c = len(targets)
for i, s in enumerate(attributes):
df = df.append({'PC1': result[i + c, 0], 'PC2': result[i + c, 1], 'type':'attributes'}, ignore_index=True)
plt.text(result[i + c, 0], result[i + c, 1], s)
sns.scatterplot(data=df, x='PC1', y='PC2', hue='type', legend=True)
# +
result = process_parts_sim(base_model, [
('He is working in the hospital as a', '[MASK]', '.', 'target'),
('She is working in the hospital as a', '[MASK]', '.', 'target'),
('[MASK] is working in the hospital as a', 'doctor', '.', 'target'),
('[MASK] is working in the hospital as a', 'nurse', '.', 'target')
])
sim = torch.nn.CosineSimilarity(dim=-1)
print(f'He is working in the hospital as a doctor: {round(sim(result[0], result[2]).item(), 3)}')
print(f'She is working in the hospital as a doctor: {round(sim(result[1], result[2]).item(), 3)}')
print()
print(f'He is working in the hospital as a nurse: {round(sim(result[0], result[3]).item(), 3)}')
print(f'She is working in the hospital as a nurse: {round(sim(result[1], result[3]).item(), 3)}')
# -
result = process_parts_sim(base_model, [
('She is working', 'in', 'a hospital.', 'target'),
('She is working', 'at', 'a hospital.', 'target'),
('She is working', 'on', 'a hospital.', 'target'),
('She is working', '[MASK]', 'a hospital.', 'target')
])
print('in', sim(result[0], result[3]).item())
print('at', sim(result[1], result[3]).item())
print('on', sim(result[2], result[3]).item())
# +
A_sents = [
'This is a male.',
'This is a female.',
'This is a man.',
'This is a woman.',
'This is a boy.',
'This is a girl.'
]
T_sents = [
'This is a programmer.',
'This is a homemaker.',
'This is a skipper.',
'This is a nurse.',
'This is a banker.',
'This is a cheerleader.',
'This is a mathematician.',
'This is a librarian'
]
T_A_sents = T_sents[:2]
for s in T_sents[:2]:
T_A_sents.append(s.replace('This', 'He'))
T_A_sents.append(s.replace('This', 'She'))
T_A_sents
# -
process(base_model, A_sents, attribute_tokenizer)
process(debiased_model, A_sents, attribute_tokenizer)
process(kaneko_model, A_sents, attribute_tokenizer)
process(base_model, T_sents, target_tokenizer)
process(debiased_model, T_sents, target_tokenizer)
process(kaneko_model, T_sents, target_tokenizer)
process(base_model, T_A_sents, target_tokenizer)
process(debiased_model, T_A_sents, target_tokenizer)
process(kaneko_model, T_A_sents, target_tokenizer)
homemaker = ['He is the homemaker.', 'She is the homemaker.', 'homemaker.']
process(base_model, homemaker, target_tokenizer)
process(debiased_model, homemaker, target_tokenizer)
process(kaneko_model, homemaker, target_tokenizer)
attr = ['He is here.', 'She is here.', 'The woman is here.', 'The man is here.', 'The boy is here.', 'The girl is here.']
targ = ['She is the homemaker.', 'He is the homemaker.', 'This is the homemaker.']
process3(base_model, targ, attr)
process3(debiased_model, targ, attr)
process3(kaneko_model, targ, attr)
| 7_SEAT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TT-decomposition
# [tt_hse16_slides](https://bayesgroup.github.io/team/arodomanov/tt_hse16_slides.pdf)
# [Tensorising Neural Networks](https://arxiv.org/pdf/1509.06569.pdf) ([NIPS2015](http://papers.nips.cc/paper/5787-tensorizing-neural-networks))
# Unfolding matrices into a tensor $
# A \in \mathbb{R}^{n_0\times \ldots \times n_{d-1}}
# $
#
# $$
# A_k = \bigl(A_{i_{:k}, i_{k:}}\bigr)_{i \in \prod_{j=0}^{d-1} [n_j]}
# \in \mathbb{R}^{
# [n_0 \times \ldots \times n_{k-1}]
# \times [n_k \times \ldots \times n_d]
# }
# \,. $$
#
# where $n_{:k} = (n_j)_{j=0}^{k-1}$ and $n_{k:} = (n_j)_{j=k}^{d-1}$ -- zero-based like numpy.
# TT-format:
#
# $$
# A_{i} = \sum_{\alpha}
# \prod_{j=0}^{d-1} G_{\alpha_j i_j \alpha_{j+1}}
# \,, $$
#
# where $
# G_{\cdot i_j \cdot} \in \mathbb{R}^{r_j \times r_{j+1}}
# $ and $r_0 = r_d = 1$. The rank of the TT-decomposition is $r = \max_{j=0}^d r_j$.
# ## Tensors
# +
import numpy as np
import torch
import torch.nn.functional as F
# %matplotlib inline
import matplotlib.pyplot as plt
# -
# Import Tensor-Train converters
# +
from ttmodule import tensor_to_tt, tt_to_tensor
from ttmodule import matrix_to_tt, tt_to_matrix
# -
# A simple, run-of-the-mill training loop.
# * imports from [`cplxmodule`](https://github.com/ivannz/cplxmodule.git)
# +
import tqdm
from cplxmodule.relevance import penalties
from cplxmodule.utils.stats import sparsity
def train_model(X, y, model, n_steps=20000, threshold=1.0,
klw=1e-3, verbose=False):
model.train()
optim = torch.optim.Adamax(model.parameters(), lr=2e-3)
losses, weights = [], []
with tqdm.tqdm(range(n_steps), disable=not verbose) as bar:
for i in bar:
optim.zero_grad()
y_pred = model(X)
mse = F.mse_loss(y_pred, y)
kl_d = sum(penalties(model))
loss = mse + klw * kl_d
loss.backward()
optim.step()
losses.append(float(loss))
bar.set_postfix_str(f"{float(mse):.3e} {float(kl_d):.3e}")
with torch.no_grad():
weights.append(model.weight.clone())
# end for
# end with
return model.eval(), losses, weights
def test_model(X, y, model, threshold=1.0):
model.eval()
with torch.no_grad():
mse = F.mse_loss(model(X), y)
kl_d = sum(penalties(model))
f_sparsity = sparsity(model, threshold=threshold, hard=True)
print(f"{f_sparsity:.1%} {mse.item():.3e} {float(kl_d):.3e}")
return model
# -
# <br>
# +
from ttmodule import TTLinear
from torch.nn import Linear
from cplxmodule.relevance import LinearARD
from cplxmodule.relevance import LinearL0ARD
# -
# Specify the problem and device
threshold, device_ = 3.0, "cpu"
# Create a simple dataset: $
# (x_i, y_i)_{i=1}^n \in \mathbb{R}^{d}\times\mathbb{R}^{p}
# $ and $y_i = E_{:p} x_i$ with $E_{:p} = (e_j)_{j=1}^p$ the diagonal
# projection matrix onto the first $p$ dimensions. We put $n\leq p$.
# +
import torch.utils.data
n_features, n_output = 250, 50
X = torch.randn(10200, n_features)
y = -X[:, :n_output].clone()
dataset = torch.utils.data.TensorDataset(X.to(device_), y.to(device_))
train, test = dataset[:200], dataset[200:]
# -
# ## A TT-linear layer
models = {}
# A useful way of thinking about the TT-format of tensors is the following.
# If we assume the thelixcographic order of index traversl of the tensor $A$
# (`C`-order, or row-major) then
#
# $$
# A_\mathbf{i}
# = \prod_{k=1}^d G^{(k)}_{i_k}
# = \sum_\mathbf{\alpha}
# \prod_{k=1}^d e_{\alpha_{k-1}}^\top G^{(k)}_{i_k} e_{\alpha_k}
# = \sum_\mathbf{\alpha}
# \prod_{k=1}^d g^k_{\alpha_{k-1} i_k \alpha_k}
# \,, \\
# \mathop{vec} A
# = \sum_\mathbf{\alpha}
# g^1_{\alpha_{0} \alpha_1}
# \otimes g^2_{\alpha_{1} \alpha_2}
# \otimes \cdots
# \otimes g^d_{\alpha_{d-1} \alpha_d}
# \,, $$
#
# with $
# \mathbf{i} = (i_k)_{k=1}^d
# $ running from $1$ to $
# [n_1\times \ldots \times n_d]
# $,
# $\alpha$ running over $\prod_{k=0}^d [r_k]$, $\otimes$ being the Krnoecker product
# and `vec` taken in the lexicographic (row-major) order. The cores are $
# G^{(k)}_{i_k} \in \mathbb{R}^{r_{k-1} \times r_k}
# $
# $i_k \in [n_k]$, and their `vec`-versions -- $
# g^k_{\alpha_{k-1} \alpha_k} \in \mathbb{R}^{n_k}
# $
# for $\alpha_{k-1} \in [n_{k-1}]$ and $\alpha_k \in [n_k]$.
#
# In the case of a matrix TT-decomposition with shapes $(n_k)_{k=1}^d$
# and $(m_k)_{k=1}^d$ we have:
#
# $$
# A = \sum_\mathbf{\alpha}
# B^1_{\alpha_{0} \alpha_1}
# \otimes \cdots
# \otimes B^d_{\alpha_{d-1} \alpha_d}
# \,, $$
#
# with $
# B^k_{\alpha_{k-1} \alpha_k} \in \mathbb{R}^{n_k\times m_k}
# $ and
# $
# B^k_{\alpha_{k-1} \alpha_k p q} = G^{(k)}_{\alpha_{k-1} [p q] \alpha_k}
# $, since each $i_k = [p q]$ is in fact a flattened index of the row-major
# flattened dimension $n_k\times m_k$.
# The matrix dimension factorization determines the block heirarchy of
# the matrix and thus is crucial to the properties and success of a linear
# layer with the weight in TT-format. If the linear layer in upstream,
# i.e. close to the inputs of the network, then the factorization and
# the induced heirarcy has semantic ties to the input features. In the
# mid-stream layers any particular heirarchy has less rationale, albeit
# it seems that the general-to-particular dimension factorization order
# is still preferable.
# #### Detailed deep factorization
# +
models["detailed-deep-lo"] = TTLinear(
[5, 5, 5, 2], [5, 5, 2, 1], rank=1, bias=False, reassemble=True)
models["detailed-deep-hi"] = TTLinear(
[5, 5, 5, 2], [5, 5, 2, 1], rank=5, bias=False, reassemble=True)
# -
# #### Detailed shallow factorization
# +
models["detailed-shallow-lo"] = TTLinear(
[25, 10], [25, 2], rank=1, bias=False, reassemble=True)
models["detailed-shallow-hi"] = TTLinear(
[25, 10], [25, 2], rank=5, bias=False, reassemble=True)
# +
# models["detailed-lo"] = TTLinear(
# [25, 5, 2], [5, 5, 2], rank=1, bias=False, reassemble=True)
# models["detailed-lo"] = TTLinear(
# [5, 5, 5, 1, 2], [5, 5, 2, 1, 1], rank=3, bias=False, reassemble=True)
# -
models["dotted"] = TTLinear(
# [25, 10, 1], [5, 5, 2], rank=1, bias=False, reassemble=True)
[25, 5, 2], [5, 5, 2], rank=1, bias=False, reassemble=True)
# #### Coarse deep factorization
# This one, with inverted hierarchy fails
# +
models["coarse-deep-lo"] = TTLinear(
[2, 5, 5, 5], [1, 2, 5, 5], rank=1, bias=False, reassemble=True)
models["coarse-deep-hi"] = TTLinear(
[2, 5, 5, 5], [1, 2, 5, 5], rank=5, bias=False, reassemble=True)
# -
# #### Coarse shallow factorization
# +
models["coarse-shallow-lo"] = TTLinear(
[10, 25], [5, 10], rank=1, bias=False, reassemble=True)
models["coarse-shallow-hi"] = TTLinear(
[10, 25], [5, 10], rank=5, bias=False, reassemble=True)
# -
models["striped"] = TTLinear(
[5, 25, 2], [5, 5, 2], rank=1, bias=False, reassemble=True)
# +
# model = LinearARD(n_features, n_output, bias=False)
# model = LinearL0ARD(n_features, n_output, bias=False, group=None)
# -
models["blocked"] = TTLinear(
[5, 25, 1, 2], [5, 5, 2, 1], rank=3, bias=False, reassemble=True)
# Train
models["test"] = TTLinear(
[10, 25], [10, 5], rank=1, bias=False, reassemble=True)
models["test"] = TTLinear(
[5, 5, 10], [2, 5, 5], rank=1, bias=False, reassemble=True)
models["test"] = TTLinear(
[25, 10, 1], [2, 5, 5], rank=1, bias=False, reassemble=True)
models["test"] = TTLinear(
[25, 10], [2, 25], rank=1, bias=False, reassemble=True)
models["test"] = TTLinear(
[2, 5, 25], [2, 1, 25], rank=1, bias=False, reassemble=True)
models["test"] = TTLinear(
[2, 5, 25], [2, 5, 5], rank=4, bias=False, reassemble=True)
models["test"] = TTLinear(
[5, 1, 25, 2], [1, 5, 2, 5], rank=3, bias=False, reassemble=True)
models["test"] = TTLinear(
[25, 5, 2], [1, 25, 2], rank=1, bias=False, reassemble=True)
models["test"] = TTLinear(
[5, 25, 1, 2], [5, 5, 2, 1], rank=2, bias=False, reassemble=True)
model, losses, weights = train_model(
*train, models["test"], n_steps=2000,
threshold=threshold, klw=1e0, verbose=True)
# Test the model
test_model(*test, model, threshold=threshold)
for core in model.cores:
plt.imshow(abs(core.detach()).numpy()[0, ..., 0].T,
cmap=plt.cm.bone, interpolation=None)
plt.gca().get_xaxis().set_visible(False)
plt.gca().get_yaxis().set_visible(False)
plt.show()
# <br>
# ## Simple visualization
# ... with not very simple setup
# +
from matplotlib.gridspec import GridSpec
def canvas_setup(figsize, **kwargs):
fig = plt.figure(figsize=figsize)
gs = GridSpec(1, 2, figure=fig, width_ratios=[7, 1])
ax_main = fig.add_subplot(gs[0])
ax_loss = fig.add_subplot(gs[1])
with torch.no_grad():
ax_main.imshow(abs(weights[0]).numpy(), cmap=plt.cm.bone)
ax_loss.semilogy(losses)
plt.tight_layout()
return fig, (ax_main, ax_loss)
# -
def canvas_clear(*axes):
"""Clear axis preserving its aesthetics."""
for ax in axes:
props = ax.properties()
ax.clear()
ax.update({
k: props[k] for k in [
"xticks", "yticks", "xlim", "ylim", "zorder", "alpha"
]
})
return axes
def animate_weight(n_epoch, *axes):
ax_main, ax_loss = canvas_clear(*axes)
artists = []
with torch.no_grad():
artists.append(ax_main.imshow(
abs(weights[n_epoch]).numpy(),
cmap=plt.cm.bone,
interpolation=None
))
artists.append(ax_main.set_title(f"it. {n_epoch}"))
artists.append(
ax_loss.semilogy(losses[:n_epoch + 1], lw=2, color="fuchsia")
)
artists.append(
ax_loss.scatter([n_epoch + 1], [losses[n_epoch]],
s=25, color="cyan")
)
artists.append(
ax_loss.axvline(n_epoch + 1, c='cyan', lw=2, alpha=0.25, zorder=-10)
)
return [
artist_ for artist_ in artists
if hasattr(artist_, "set_animated")
]
# An interactive slider with ipywidgets
# +
from ipywidgets import widgets
def int_slider(value, min, max, step):
return widgets.IntSlider(
value=value, min=min, max=max, step=step, continuous_update=False,
layout=widgets.Layout(min_width='500px', display='flex'))
# +
def plot_weight(n_epoch=0):
fig, axes = canvas_setup(figsize=(16, 3))
animate_weight(n_epoch, *axes)
plt.show()
widgets.interact(plot_weight, n_epoch=int_slider(1000, 0, len(weights)-1, 10));
# -
# <br>
# +
import matplotlib.animation as animation
try:
FFMpegWriter = animation.writers['ffmpeg_file']
class PatchedFFMpegWriter(FFMpegWriter):
def setup(self, fig, outfile, *args, **kwargs):
dpi = kwargs.get("dpi", getattr(self, "dpi", None))
frame_prefix = kwargs.get(
"frame_prefix", getattr(self, "temp_prefix", '_tmp'))
clear_temp = kwargs.get(
"clear_temp", getattr(self, "clear_temp", True))
super().setup(fig, outfile, clear_temp=clear_temp,
frame_prefix=frame_prefix, dpi=dpi)
except:
class PatchedFFMpegWriter(animation.AbstractMovieWriter):
pass
# +
import os
import time
import tempfile
dttm = time.strftime("%Y%m%d-%H%M%S")
fig, axes = canvas_setup(figsize=(16, 3))
fps, n_frames = 15, len(weights)
schedule = [
*range(0, 25, 1)
] + [
*range(25, n_frames, 10)
]
shape_tag = model.extra_repr()
outfile = os.path.join(".", f"weight-{model.__class__.__name__}{shape_tag}-{dttm}.mp4")
# dump the intermediate frames into a temporary dir
with tempfile.TemporaryDirectory() as tmp:
print(f"temp dir at {tmp}", flush=True)
writer = PatchedFFMpegWriter(fps=fps, bitrate=-1, metadata={})
writer.setup(fig, outfile, frame_prefix=os.path.join(tmp, f"_frame_"))
ani = animation.FuncAnimation(
fig, animate_weight, tqdm.tqdm_notebook(schedule, unit="frm"),
interval=1, repeat_delay=None, blit=False, fargs=axes)
ani.save(outfile, writer=writer)
plt.close()
# +
from IPython.display import Video
print(outfile)
Video(data=outfile, embed=True, width=768)
# -
assert False
# ## Matrix-vector product in TT-format
#
# Suppose the TT representation of a matrix $W\in \mathbb{R}^{n\times m}$
# with shapes $(n_k)_{k=1}^d$ and $(m_k)_{k=1}^d$ is given by $ \prod_{k=1}^d
# G^{(k)}_{i_k j_k}$ with $
# G^{(k)}_{i_k j_k} \in \mathbb{R}^{r_{k-1}\times r_k}
# $ with $r_0 = r_d = 1$. Then for index $
# \alpha \in \prod_{k=1}^{d-1} [r_k]
# $ with $\alpha_0 = \alpha_d = 1$ we have:
#
# $$
# y_j = e_j^\top W^\top x
# = \sum_\alpha \sum_i
# \prod_{k=1}^d g_{\alpha_{k-1} i_k j_k \alpha_k} x_i
# = \sum_{\alpha_0, \alpha_{1:}} \sum_{i_{2:}}
# \prod_{k=2}^d g_{\alpha_{k-1} i_k j_k \alpha_k}
# \sum_{i_1} g_{\alpha_0 i_1 j_1 \alpha_1} x_{i_1 i_{2:}}
# = \sum_{\alpha_{1:}} \sum_{i_{2:}}
# \prod_{k=2}^d g_{\alpha_{k-1} i_k j_k \alpha_k}
# \sum_{\alpha_0, i_1} g_{\alpha_0 i_1 j_1 \alpha_1} x_{i_1 i_{2:} \alpha_0}
# \,,\\
# \dots
# = \sum_{\alpha_{1:}} \sum_{i_{2:}}
# \prod_{k=2}^d g_{\alpha_{k-1} i_k j_k \alpha_k} z_{i_{2:} j_1 \alpha_1}
# = \sum_{\alpha_{2:}} \sum_{i_{3:}}
# \prod_{k=3}^d g_{\alpha_{k-1} i_k j_k \alpha_k} z_{i_{3:} j_{:3} \alpha_2}
# \,. $$
# <br>
# ## Tensor Rings
#
# See [Tensor Ring Decomposition](https://arxiv.org/abs/1606.05535). Essentially the same idea but
# with $t_0 = r_d \geq 1$. Tensors in TT-format are a special case of TR-format:
#
# $$
# A_\mathbf{i}
# = \mathop{Tr} \prod_{k=1}^d G^{(k)}_{i_k}
# = \sum_{\mathbf{\alpha}\colon \alpha_0=\alpha_d}
# \prod_{k=1}^d e_{\alpha_{k-1}}^\top G^{(k)}_{i_k} e_{\alpha_k}
# = \sum_{\mathbf{\alpha}\colon \alpha_0=\alpha_d}
# \prod_{k=1}^d g^k_{\alpha_{k-1} i_k \alpha_k}
# \,, $$
#
# where $
# G^{(k)}_{i_k} \in \mathbb{R}^{r_j \times r_{j+1}}
# $ and $r_0 = r_d$ and $
# \alpha \in \prod_{k=0}^d [r_k]
# $.
#
# This might be interesting [The trouble with tensor ring decompositions](https://arxiv.org/abs/1811.03813)
# (minimal-rank recovery).
# +
# ranks = [2, 3, 4, 5, 5]
# shapes = [2, 3, 7, 4, 5], [3, 7, 7, 5, 2]
ranks = [3, 2, 1, 5]
shape = [2, 3, 7, 5], [3, 7, 1, 2]
cores = [torch.randn(r0, n, m, r1, dtype=torch.double)
for r0, n, m, r1 in zip(ranks[-1:] + ranks[:-1], *shape, ranks)]
# -
def tr_to_tensor_zero(*cores):
# chip off the first core and contract the rest
rest = tt_to_tensor(*cores[1:], squeeze=False)
# contract with tensor_dot (reshape + einsum("i...j, j...i->...") was slower)
return torch.tensordot(cores[0], rest, dims=[[0, -1], [-1, 0]])
# +
from ttmodule.tensor import tr_to_tensor
# res = %timeit -o -n 100 -r 25 tr_to_tensor_zero(*cores)
timing = [res]
for k in range(len(cores)):
# res = %timeit -o -n 100 -r 25 tr_to_tensor(*cores, k=k)
timing.append(res)
print(f">>> ({k}) {ranks[k]} {cores[k].shape}")
# -
# $$
# W_{ij} = \mathop{tr}
# \prod_{k=1}^d G^{(k)}_{i_k j_k}
# \,, \\
# y_j = \sum_i W_{ij} x_i
# = \sum_i \mathop{tr} \prod_{k=1}^d G^{(k)}_{i_k j_k} x_i
# = \mathop{tr} \sum_i \prod_{k=1}^d G^{(k)}_{i_k j_k} x_i
# = \mathop{tr} \sum_{i_{1:}} \sum_{i_1} \prod_{k=1}^d G^{(k)}_{i_k j_k} x_i
# \,, \\
# y_j = \mathop{tr} \sum_{i_{1:}}
# \prod_{k=2}^d G^{(k)}_{i_k j_k} \sum_{i_1} G^{(1)}_{i_1 j_1} x_i
# \,. $$
# +
from ttmodule.matrix import tr_to_matrix
weight = tr_to_matrix(shape, *cores, k=2)
# -
# With $\alpha \in \prod_{k=1}^d [r_k]$ and $\alpha_0 = \alpha_d$
# and broadcasting $x_{i \alpha_d} = x_{i}$
# $$
# y = W^\top x
# = \bigl( \sum_i \sum_\alpha
# \prod_{k=1}^d g_{\alpha_{k-1} i_k j_k \alpha_k} x_i \bigr)_j
# = \bigl(\sum_\alpha \sum_{i_{:d}}
# \prod_{k=1}^{d-1} g_{\alpha_{k-1} i_k j_k \alpha_k}
# \sum_{i_d} g_{\alpha_{d-1} i_d j_d \alpha_d} x_i \bigr)_j
# \,, \\
# y = W^\top x
# = \bigl(\sum_{\alpha_d \alpha_1} \sum_{\alpha_{2:d}} \sum_{i_{2:}}
# \prod_{k=2}^d g_{\alpha_{k-1} i_k j_k \alpha_k}
# \sum_{i_1} g_{\alpha_d i_1 j_1 \alpha_1} x_i \bigr)_j
# = \bigl(\sum_{\alpha_d \alpha_1} \sum_{i_{2:}}
# Z_{\alpha_1 i_{2:} j_{2:} \alpha_d}
# \sum_{i_1} g_{\alpha_d i_1 j_1 \alpha_1} x_i \bigr)_j
# \,, \\
# y_j = e_j^\top W^\top x
# = \sum_{\alpha_d} \sum_i Z_{\alpha_d i j \alpha_d} x_i
# \,. $$
def ttmv(shape, input, *cores):
*head, tail = input.shape
data = input.view(-1, *shape[0], 1)
for core in cores:
data = torch.tensordot(data, core, dims=[[1, -1], [1, 0]])
return data.reshape(*head, -1)
#
# +
input = torch.randn(100, np.prod(shape[0])).double()
reference = sum([
ttmv(shape, input, cores[ 0][[a], ...],
*cores[1:-1], cores[-1][..., [a]])
for a in range(ranks[-1])
])
assert torch.allclose(reference, torch.mm(input, weight))
# +
from ttmodule.matrix import invert
def tr_vec(shape, input, *cores, k=0):
k = (len(cores) + k) if k < 0 else k
assert 0 <= k < len(cores)
*head, tail = input.shape
data = input.view(-1, *shape[0])
shuffle = list(range(1, data.dim()))
shuffle = 0, *shuffle[k:], *shuffle[:k]
data = data.permute(shuffle).unsqueeze(-1)
cores, output = cores[k:] + cores[:k], 0
for a in range(cores[0].shape[0]):
cyc = cores[ 0][[a], ...], *cores[1:-1], cores[-1][..., [a]]
interm = data.clone()
for core in cyc:
interm = torch.tensordot(interm, core, dims=[[1, -1], [1, 0]])
output += interm
return output.squeeze(-1).permute(invert(*shuffle)).reshape(*head, -1)
# -
for k in range(len(cores)):
assert torch.allclose(tr_vec(shape, input, *cores, k=k), reference)
# +
*head, tail = input.shape
data = input.view(-1, *shape[0], 1)
for core in cores:
data = torch.tensordot(data, core, dims=[[1, -1], [1, 0]])
data = data.sum(dim=-1).reshape(*head, -1)
assert not torch.allclose(data, reference)
# -
# <br>
# Naive approach: spin the ring to the largest dimension, and reduce it.
# +
def roll(cores, j):
# roll ring so that `j` is at 0
j += len(cores) if j < 0 else 0
if j == 0:
return cores
return cores[j:] + cores[:j]
def ring_dot(*cores):
# n - cores [* x d_i x d_{i+1}] i=0..n-1, d_n = d_0
if len(cores) <= 1:
return cores[0]
# find the largest dim to shrink
dim = [core.shape[-1] for core in cores]
j = max(range(len(dim)), key=dim.__getitem__)
# print(dim, j)
# roll it to origin, cut and shrink link with mat-mul
a, b, *rest = roll(cores, j)
return ring_dot(torch.matmul(a, b), *rest)
# +
x = torch.randn(2, 2, 64)
ranks = [1, 5, 3, 5, 7, 9, 5, 7]
shape = [64] * len(ranks)
cores = torch.nn.ParameterList([
torch.nn.Parameter(torch.rand(r0, r1, n1) - .5)
for r0, n1, r1 in zip(ranks[-1:] + ranks, shape, ranks)
])
# +
mats = [torch.tensordot(x, core, dims=([-1], [-1])) for core in cores]
res = ring_dot(*mats)
# -
res, res.shape
grads = torch.autograd.grad(res.diagonal(dim1=-1, dim2=-2).sum(), cores)
[*map(torch.norm, grads)]
# <br>
# ### Transposed shape for TTLinear
# +
# ranks = [2, 3, 4, 5, 5]
# shapes = [2, 3, 7, 4, 5], [3, 7, 7, 5, 2]
ranks = [1, 3, 2, 5, 1]
shape = [2, 3, 7, 5], [3, 7, 1, 2]
cores = [torch.randn(r0, n, m, r1, dtype=torch.double)
for r0, n, m, r1 in zip(ranks[:-1], *shape, ranks[1:])]
# -
shape_t = shape[1], shape[0]
cores_t = [core.permute(0, 2, 1, 3) for core in cores]
def ttmv_t(shape, input, *cores):
*head, tail = input.shape
data = input.view(-1, *shape[1], 1)
for core in cores:
data = torch.tensordot(data, core, dims=[[1, -1], [2, 0]])
return data.reshape(*head, -1)
assert torch.allclose(ttmv_t(shape_t, input, *cores_t),
ttmv(shape, input, *cores))
assert torch.allclose(tt_to_matrix(shape_t, *cores_t).t(),
tt_to_matrix(shape, *cores))
| notebooks/example_with_animation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.insert(0,'..')
import os
from tollef_conll_converter import LitBankConverter
# +
datadir = "../../../data/COREFERENCE_RESOLUTION/litbank_conll_short"
outdir = "../../coreference_data/litbank_short.coreflite"
litconv = LitBankConverter(datadir, outdir)
# -
litconv.run()
| Dataset Converters/LitBank_to_CorefLite.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Kriging Example1
# - Author: <NAME>
# - Date: Jan 08, 2019
# In this example, Kriging is used to generate a surrogate model for a given data. In this data, sample points are generated using STS class and functional value at sample points are estimated using a model defined in python script ('python_model_function.py).
# Import the necessary libraries. Here we import standard libraries such as numpy and matplotlib, but also need to import the STS, RunModel and Krig class from UQpy.
from UQpy.Surrogates import Kriging
from UQpy.SampleMethods import RectangularStrata, RectangularSTS
from UQpy.RunModel import RunModel
from UQpy.Distributions import Gamma
import numpy as np
import matplotlib.pyplot as plt
# Create a distribution object.
marginals = [Gamma(a= 2., loc=1., scale=3.)]
# Create a strata object.
strata = RectangularStrata(nstrata=[20])
# Run stratified sampling
x = RectangularSTS(dist_object=marginals, strata_object=strata, nsamples_per_stratum=1, random_state=1)
# RunModel is used to evaluate function values at sample points. Model is defined as a function in python file 'python_model_function.py'.
rmodel = RunModel(model_script='python_model_1Dfunction.py', delete_files=True)
rmodel.run(samples=x.samples)
K = Kriging(reg_model='Linear', corr_model='Gaussian', nopt=20, corr_model_params=[1], random_state=2)
K.fit(samples=x.samples, values=rmodel.qoi_list)
print(K.corr_model_params)
# Kriging surrogate is used to compute the response surface and its gradient.
# +
num = 1000
x1 = np.linspace(min(x.samples), max(x.samples), num)
y, y_sd = K.predict(x1.reshape([num, 1]), return_std=True)
y_grad = K.jacobian(x1.reshape([num, 1]))
# -
# Actual model is evaluated at all points to compare it with kriging surrogate.
rmodel.run(samples=x1, append_samples=False)
# This plot shows the input data as blue dot, blue curve is actual function and orange curve represents response curve. This plot also shows the gradient and 95% confidence interval of the kriging surrogate.
# +
fig = plt.figure()
ax = plt.subplot(111)
plt.plot(x1, rmodel.qoi_list, label='Sine')
plt.plot(x1, y, label='Surrogate')
plt.plot(x1, y_grad, label='Gradient')
plt.scatter(K.samples, K.values, label='Data')
plt.fill(np.concatenate([x1, x1[::-1]]), np.concatenate([y - 1.9600 * y_sd,
(y + 1.9600 * y_sd)[::-1]]),
alpha=.5, fc='y', ec='None', label='95% CI')
box = ax.get_position()
ax.set_position([box.x0, box.y0, box.width * 0.8, box.height])
# Put a legend to the right of the current axis
ax.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.show()
| example/Surrogates/Kriging/Kriging_Example1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # single patch conv net
#
# this notebook it an attempt to solve some of the issues being encountered on the full size network by reducing the size of the problem to learn the SSIM score for a single patch of each image (using the upper left most pixels). in principle this task should be identical to what the larger net is trying to do but should give some insight into what is going on.
# #!/usr/bin/env python3
import os
os.environ['TF_CPP_MIN_LOG_LEVEL']='2'
import numpy as np
np.set_printoptions(threshold=np.nan)
import tensorflow as tf
import time
import pandas as pd
import matplotlib.pyplot as plt
import progressbar
# ### network functions
# +
def convolve_inner_layers(x, W, b):
y = tf.nn.conv2d(x, W, strides = [1,1,1,1], padding='VALID')
y = tf.nn.bias_add(y, b)
return tf.nn.tanh(y)
def convolve_ouput_layer(x, W, b):
y = tf.nn.conv2d(x, W, strides = [1,1,1,1], padding='VALID')
y = tf.nn.bias_add(y, b)
return y
def conv_net(x, W, b):
conv1 = convolve_inner_layers(x, W['weights1'], b['bias1'])
conv2 = convolve_inner_layers(conv1, W['weights2'], b['bias2'])
conv3 = convolve_inner_layers(conv2, W['weights3'], b['bias3'])
output_feed = tf.concat([conv1, conv2, conv3],3)
output = convolve_ouput_layer(output_feed, W['weights_out'], b['bias_out'])
return output
# -
# ### data processing functions
# +
def get_variance(training_target):
all_pixels = training_target.flatten()
return all_pixels.var()
def get_epoch(x, y, n):
input_size = x.shape[0]
number_batches = input_size // n
extra_examples = input_size % n
batches = {}
batch_indices = np.arange(input_size)
np.random.shuffle(batch_indices)
for i in range(number_batches):
temp_indices = batch_indices[n*i:n*(i+1)]
temp_x = []
temp_y = []
for j in temp_indices:
temp_x.append(x[j])
temp_y.append(y[j])
batches[i] = [np.asarray(temp_x), np.asarray(temp_y)]
if extra_examples != 0:
extra_indices = batch_indices[input_size-extra_examples:input_size]
temp_x = []
temp_y = []
for k in extra_indices:
temp_x.append(x[k])
temp_y.append(y[k])
batches[i+1] = [np.asarray(temp_x), np.asarray(temp_y)]
return batches
def normalize_input(train_data, test_data):
mean, std_dev = np.mean(train_data, axis=0), np.std(train_data, axis=0)
return (train_data - mean) / std_dev, (test_data - mean) / std_dev
def calculate_ssim(window_orig, window_recon):
k_1 = 0.01
k_2 = 0.03
L = 255
if window_orig.shape != (11,11) or window_recon.shape != (11,11):
raise ValueError('please check window size for SSIM calculation!')
orig_data = window_orig.flatten()
recon_data = window_recon.flatten()
mean_x = np.mean(orig_data)
mean_y = np.mean(recon_data)
var_x = np.var(orig_data)
var_y = np.var(recon_data)
covar = np.cov(orig_data, recon_data)[0][1]
c_1 = (L*k_1)**2
c_2 = (L*k_2)**2
num = (2*mean_x*mean_y+c_1)*(2*covar+c_2)
den = (mean_x**2+mean_y**2+c_1)*(var_x+var_y+c_2)
return num/den
# -
# ### network set up
# network parameters
filter_dim, filter_dim2 = 11, 1
batch_size = 200
image_dim, result_dim = 96, 86
input_layer, first_layer, second_layer, third_layer, output_layer = 4, 17, 9, 4, 1
learning_rate = .01
epochs = 300
# +
# data input
data_path = 'https://raw.githubusercontent.com/michaelneuder/image_quality_analysis/master/data/sample_data/'
# train data --- 500 images, 96x96 pixels
orig_500 = pd.read_csv('{}orig_500.txt'.format(data_path), header=None, delim_whitespace = True)
recon_500 = pd.read_csv('{}recon_500.txt'.format(data_path), header=None, delim_whitespace = True)
# test data --- 140 images, 96x96 pixels
orig_140 = pd.read_csv('{}orig_140.txt'.format(data_path), header=None, delim_whitespace = True)
recon_140 = pd.read_csv('{}recon_140.txt'.format(data_path), header=None, delim_whitespace = True)
# train target --- 500 images, 86x86 pixels (dimension reduction due no zero padding being used)
ssim_500 = pd.read_csv('{}ssim_500_new.csv'.format(data_path), header=None).values
ssim_140 = pd.read_csv('{}ssim_140_new.csv'.format(data_path), header=None).values
# +
# getting 4 input channels for train and test
original_images_train = orig_500.values
original_images_train_sq = orig_500.values**2
reconstructed_images_train = recon_500.values
reconstructed_images_train_sq = recon_500.values**2
original_images_test = orig_140.values
original_images_test_sq = orig_140.values**2
reconstructed_images_test = recon_140.values
reconstructed_images_test_sq = recon_140.values**2
# stack inputs
training_input = np.dstack((original_images_train, reconstructed_images_train, original_images_train_sq, reconstructed_images_train_sq))
testing_input = np.dstack((original_images_test, reconstructed_images_test, original_images_test_sq, reconstructed_images_test_sq))
# normalize inputs
training_input_normalized, testing_input_normalized = normalize_input(training_input, testing_input)
# get size of training and testing set
train_size = original_images_train.shape[0]
test_size = original_images_test.shape[0]
# reshaping
train_data = np.reshape(training_input_normalized, [train_size,image_dim,image_dim,input_layer])
test_data = np.reshape(testing_input_normalized, [test_size,image_dim,image_dim,input_layer])
# reshaping the result data to --- (num pics), 86, 86, 1
train_target = np.reshape(ssim_500, [train_size, result_dim, result_dim, output_layer])
test_target = np.reshape(ssim_140, [test_size, result_dim, result_dim, output_layer])
# -
# ### single patch set up
#
# now that we have all the network data set up, we can just grab the top left corner (11x11 pixels) of each image.
single_train_data, single_test_data = train_data[:,:11,:11,:], test_data[:,:11,:11,:]
# first we will do manual calculations of the ssim score. and compare that to the matlab data (read from file).
# +
ssim_train, ssim_test = [], []
for i in range(train_size):
ssim_train.append(calculate_ssim(single_train_data[i][...,0], single_train_data[i][...,1]))
if i < 140:
ssim_test.append(calculate_ssim(single_test_data[i][...,0], single_test_data[i][...,1]))
ssim_train = np.reshape(np.asarray(ssim_train), (train_size,1))
ssim_test = np.reshape(np.asarray(ssim_test), (test_size,1))
# -
ssim_train.shape, ssim_test.shape
single_train_data.shape
# +
f, axarr = plt.subplots(nrows=3,ncols=4, figsize=(12,9))
for i in range(3):
index = np.random.randint(500)
axarr[i,0].imshow(single_train_data[index,:,:,0], cmap='gray')
axarr[i,1].imshow(single_train_data[index,:,:,1], cmap='gray')
axarr[i,2].imshow(single_train_data[index,:,:,2], cmap='gray')
axarr[i,3].imshow(single_train_data[index,:,:,3], cmap='gray')
axarr[0,0].set_title('original')
axarr[0,1].set_title('reconstructed')
axarr[0,2].set_title('orig squared')
axarr[0,3].set_title('recon squared')
for ax_row in axarr:
for ax in ax_row:
ax.set_xticklabels([])
ax.set_yticklabels([])
f.suptitle('training data sample', size=15)
plt.savefig('single_patch.png')
plt.show()
# -
ssim_train_mat = np.reshape(train_target[:,0,0,0], [train_size,1])
ssim_test_mat = np.reshape(test_target[:,0,0,0], [test_size,1])
ssim_train_mat.shape, ssim_test_mat.shape
x = ssim_test_mat - ssim_test
x.mean()
# now we have the target values set up we can train the network with the python data.
# +
# tensorflow graph set up
tf.reset_default_graph()
# initializing variables --- fan in
scaling_factor = 1.0
initializer = tf.contrib.layers.variance_scaling_initializer(factor=scaling_factor, mode='FAN_IN')
weights = {
'weights1': tf.get_variable('weights1', [filter_dim,filter_dim,input_layer,first_layer], initializer=initializer),
'weights2': tf.get_variable('weights2', [filter_dim2,filter_dim2,first_layer,second_layer], initializer=initializer),
'weights3': tf.get_variable('weights3', [filter_dim2,filter_dim2,second_layer,third_layer], initializer=initializer),
'weights_out': tf.get_variable('weights4', [filter_dim2,filter_dim2,third_layer+second_layer+first_layer,output_layer], initializer=initializer)
}
biases = {
'bias1': tf.get_variable('bias1', [first_layer], initializer=initializer),
'bias2': tf.get_variable('bias2', [second_layer], initializer=initializer),
'bias3': tf.get_variable('bias3', [third_layer], initializer=initializer),
'bias_out': tf.get_variable('bias4', [output_layer], initializer=initializer)
}
# tf Graph input
x = tf.placeholder(tf.float32, [None, filter_dim, filter_dim, input_layer])
y = tf.placeholder(tf.float32, [None, output_layer])
# model
prediction = conv_net(x, weights, biases)
# get variance to normalize error terms during training
variance = get_variance(train_target)
# loss and optimization
cost = tf.reduce_mean(tf.square(tf.subtract(prediction, y)))
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate).minimize(cost)
# -
# session
init = tf.global_variables_initializer()
error_train, error_test = [], []
with tf.Session() as sess:
sess.run(init)
bar = progressbar.ProgressBar()
for epoch_count in bar(range(epochs)):
epoch = get_epoch(single_train_data, ssim_train, batch_size)
for i in epoch:
x_data_train, y_data_train = np.asarray(epoch[i][0]), np.asarray(epoch[i][1])
sess.run(optimizer, feed_dict={x : x_data_train, y : y_data_train})
loss = sess.run(cost, feed_dict={x : x_data_train, y : y_data_train})
percent_error = 100*loss/variance
error_train.append(percent_error)
score = sess.run(cost, feed_dict={x: single_test_data, y: ssim_test})
percent_error = 100*score/variance
error_test.append(percent_error)
fig, ax = plt.subplots(figsize=(14,7))
ax.plot(np.arange(len(error_train)), error_train, label='train', color='k', alpha=0.5)
ax.plot(np.arange(len(error_test)), error_test, label='test', color='b', alpha=0.5)
ax.legend()
ax.set_ylim(0,300)
ax.set_ylabel('error', size=20)
ax.set_xlabel('epoch count', size=20)
ax.set_title('python data : single patch network', size=20)
# plt.savefig('python_single.png')
plt.show()
# this looks great! now lets try it with the matlab data.
# +
# tensorflow graph set up
tf.reset_default_graph()
# initializing variables --- fan in
scaling_factor = 1.0
initializer = tf.contrib.layers.variance_scaling_initializer(factor=scaling_factor, mode='FAN_IN')
weights = {
'weights1': tf.get_variable('weights1', [filter_dim,filter_dim,input_layer,first_layer], initializer=initializer),
'weights2': tf.get_variable('weights2', [filter_dim2,filter_dim2,first_layer,second_layer], initializer=initializer),
'weights3': tf.get_variable('weights3', [filter_dim2,filter_dim2,second_layer,third_layer], initializer=initializer),
'weights_out': tf.get_variable('weights4', [filter_dim2,filter_dim2,third_layer+second_layer+first_layer,output_layer], initializer=initializer)
}
biases = {
'bias1': tf.get_variable('bias1', [first_layer], initializer=initializer),
'bias2': tf.get_variable('bias2', [second_layer], initializer=initializer),
'bias3': tf.get_variable('bias3', [third_layer], initializer=initializer),
'bias_out': tf.get_variable('bias4', [output_layer], initializer=initializer)
}
# tf Graph input
x = tf.placeholder(tf.float32, [None, filter_dim, filter_dim, input_layer])
y = tf.placeholder(tf.float32, [None, output_layer])
# model
prediction = conv_net(x, weights, biases)
# get variance to normalize error terms during training
variance = get_variance(train_target)
# loss and optimization
cost = tf.reduce_mean(tf.square(tf.subtract(prediction, y)))
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate).minimize(cost)
# -
# session
init = tf.global_variables_initializer()
error_train, error_test = [], []
with tf.Session() as sess:
sess.run(init)
bar = progressbar.ProgressBar()
for epoch_count in bar(range(epochs)):
epoch = get_epoch(single_train_data, ssim_train_mat, batch_size)
for i in epoch:
x_data_train, y_data_train = np.asarray(epoch[i][0]), np.asarray(epoch[i][1])
sess.run(optimizer, feed_dict={x : x_data_train, y : y_data_train})
loss = sess.run(cost, feed_dict={x : x_data_train, y : y_data_train})
percent_error = 100*loss/variance
error_train.append(percent_error)
score = sess.run(cost, feed_dict={x: single_test_data, y: ssim_test_mat})
percent_error = 100*score/variance
error_test.append(percent_error)
fig, ax = plt.subplots(figsize=(14,7))
ax.plot(np.arange(len(error_train)), error_train, label='train', color='k', alpha=0.5)
ax.plot(np.arange(len(error_test)), error_test, label='test', color='b', alpha=0.5)
ax.legend()
ax.set_ylim(0,300)
ax.set_ylabel('error', size=20)
ax.set_xlabel('epoch count', size=20)
ax.set_title('matlab data : single patch network', size=20)
plt.savefig('matlab_single.png')
plt.show()
# not so great. this is not good
#
# lets compare the data from matlab and python.
ssim_140_py = pd.read_csv('{}SSIM_140.txt'.format(data_path), header=None, delim_whitespace=True).values
ssim_140_mat_ng = pd.read_csv('{}ssim_140_nogauss.csv'.format(data_path), header=None).values
ssim_140_py = np.reshape(ssim_140_py, (140,96,96))
ssim_140_mat_ng = np.reshape(ssim_140_mat_ng, (140,86,86))
ssim_140_mat_ng.shape, ssim_140_py.shape
# +
f, axarr = plt.subplots(nrows=3,ncols=2, figsize=(6,9))
for i in range(3):
index = np.random.randint(140)
axarr[i,0].imshow(ssim_140_py[index,5:91,5:91], cmap='gray')
axarr[i,1].imshow(ssim_140_mat_ng[index,:,:], cmap='gray')
axarr[0,0].set_title('python')
axarr[0,1].set_title('matlab: no gauss')
for ax_row in axarr:
for ax in ax_row:
ax.set_xticklabels([])
ax.set_yticklabels([])
f.suptitle('training data sample', size=20)
plt.savefig('data_comp.png')
plt.show()
# -
diff = ssim_140_py[index,5:91,5:91]-ssim_140_mat_ng[index,:,:]
diff.mean()
| bin/nets/old/conv_net_single.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Phase 2 - Créer des jeux
# Pour vérifier si votre code Python adhère aux recomendations PEP8 allez sur ce site: http://pep8online.com/
# ## Utiliser le notebook Jupyter
# Les cellules code dans un notebook Jupyter doivent être executable. Si vous voulez simplement montrer un bout de code pour en parler sans l'executer, placez-le entre triple-backquotes.
# ```
# bout de code
# (exemple)
# ```
#
# Toutes les cellules code doivent être executable et produire un résultat :
# * Pour executer une cellule appuyez sur `SHIFT-Return`
# * Avant de créer un PDF, recalculez toutes les cellules code
# * Choisissez le menu `Kernel > Restart & Run all`
# * Ceci renumérote automatiquement toutes les cellules à partir de 1
# * Si une variables ou fonctions ne sont pas définis, vous en êtes avertit
# * Selon la langue du système (Raspberry Pi settings), l'étiquette `In` ou `Entrée` est affichée
# +
from sense_hat import SenseHat
from gamelib import *
from random import randint, choice
from time import time, sleep
import numpy as np
sense = SenseHat()
sense.clear()
# -
# Le module `gamelib` contient les couleurs.
colors = [BLACK, RED, GREEN, YELLOW, BLUE, CYAN, MAGENTA]
# ## Variables gobales
# Nous déclarons quelques variables commes globales. Ces variables seront accessible depuis partout dans Python.
x, y = 0, 0 # cursor position
n = 8 # board size (n x n)
T = np.zeros((n, n), int) # Matrix of the board (table)
colors = (BLACK, RED, GREEN, BLUE) # color list
p = 1 # current player
score = [0, 0] # current score
# ## Le module NumPy
# Pour cette deuxième phase, nous allons remplacer les matrices (listes 2D Python) par des matrices 2D NumPy. Voici les avantages:
# * les calculs sont beaucoup plus rapide
# * la matrice 2D peut être affiché avec `print`
# * il existent beaucoup de fonctions
# *
# Utiliser le module NumPy pour le calcul matriciel.
# C'est standard d'importer le module **NumPy** avec le raccourci `np`.
import numpy as np
# La fonction `np.array()` transforme une liste Python en matrice NumPy.
L = [[0, 1], [1, 1]]
# Regardons le type de cet objet. C'est un `numpy.ndarray` (numerical data array).
type(L)
# Une matrice 2D peut être affiché avec `print`.
print(L)
# ## Créer des matrices
# Il existent des fonctions pour créer des matrices
A = np.zeros((1, 1), int)
A
B = np.ones((1, 1), int)
B
print(L*2)
# Afficher une matrice sur senshat
# https://docs.scipy.org/doc/numpy/reference/generated/numpy.random.randint.html
m = len(colors)
R = np.random.randint(0, m-1, (2, 2))
R
# ## Afficher sur le SenseHAT
# Pour afficher sur les LEDs du SenseHAT nous utilisons une matrice avec des entiers. Cest entiers sont également l'indice pour la liste `colors`
def show_matrix(M):
global colors
n, m = M.shape
for x in range(n):
for y in range(m):
col = colors[M[y, x]] # lines & columns are inversed
sense.set_pixel(x, y, col)
show_matrix(R)
def show_cursor(x, y, p):
"""Show cursor for player p (1, 2)."""
sense.set_pixel(x, y, colors[p])
show_cursor(x, y, p)
# ## Indices
# Dans une matrice 2D nous utilisons les indices i, j. D'habitude on utilise i pour les lignes et j pour les colonnes. Donc les indices sont invertit:
# * i correspond à l'axe y
# * j correspond à l'axe x
T[1, 4] = 3
T[2:4, 2:4] = L * 3
T
# ## Fonctions callback
# Des fonctions peuvent être associé aux 5 directions du joystick. Les fonctions prennent comme argument l'événement `event`.
# * direction_left
# * direction_right
# * direction_up
# * direction_down
# * direction_middle
# * direction_any
#
# Si vous êtes intéressés dans les détails, ces fonctions utilisent un mécanismes qu'on appelle des **threads**. Vous pouvez aller regarder le code source de l'API sur GitHub:
#
# https://github.com/RPi-Distro/python-sense-hat/blob/master/sense_hat/stick.py
# +
# dir(sense.stick)
# +
def right(event):
"""Make display red."""
pass
def left(event):
"""Make display blue."""
pass
def up(event):
"""Make display green."""
pass
def down(event):
"""Make display yellow."""
pass
def middle(event):
"""Clear display."""
pass
sense.stick.direction_left = left
sense.stick.direction_right = right
sense.stick.direction_up = up
sense.stick.direction_down = down
sense.stick.direction_middle = middle
# -
# ### Exercice
# Redéfinir les fonctions `up` et `down` pour incrementer ou décrémenter un nombre qui change de couleur selon l'index dans `colors`.
# +
i = 0
def up(event):
global i, n
pass
def middle(event):
pass
def down(event):
global i, n
pass
# -
# ## Trouver des info sur une fonction callback
# Verifier si une fonction callback et définie.
print(sense.stick.direction_left)
# Afficher le docstring de cette fonction.
print(sense.stick.direction_left.__doc__)
# Afficher le nom de cette fonction.
print(sense.stick.direction_left.__name__)
# +
callbacks = (sense.stick.direction_up, sense.stick.direction_down,
sense.stick.direction_left, sense.stick.direction_right,
sense.stick.direction_middle, sense.stick.direction_any)
for cb in callbacks:
if cb != None:
print(cb.__name__, cb.__doc__, sep='\t')
# -
# ## Déplacer un curseur
# Beaucoup de jeux necessitent le déplacement d'un curseur. Nous utilisons les 4 directions pour déplacer le curseur, et le boutons central pour faire la sélection.
# +
def move_cursor(event):
"""Deplace le curseur (x, y) sur un board (n x n)."""
global x, y, n
if event.direction == 'left':
x = max(x-1, 0)
elif event.direction == 'up':
pass
elif event.direction == 'right':
pass
elif event.direction == 'down':
pass
def play(event):
if event.action == 'pressed':
move_cursor(event)
sense.clear()
show_cursor(x, y, 1)
sense.stick.__init__()
sense.stick.direction_any = play
# -
# En posant $n=3$ nous pouvons restreindre le mouvement du curseur à une région 3x3.
n=3
# ## Afficher le gagnant
# Après chaque match nous affichons le gagnant (1 ou 2) dans sa couleur. Si c'est match nul nous affichons 0 en blanc.
#
# Pour donner d'importance, nous flashons 3 fois.
# +
def show_winner(p, n=3):
"""Show the winner (1, 2) in the player's color."""
col = WHITE if p==0 else colors[p]
for i in range(3):
sense.show_letter(str(p), text_colour=col)
sleep(0.2)
sense.clear()
sleep(0.2)
for i in range(3):
show_winner(i)
# -
# ## Afficher le score
# Après chaque match nous affichons le score des deux jouer, ou le score simple.
# +
def show_score():
"""Display the score as scrolling text."""
text = str(score[0]) + ':' + str(score[1])
sense.show_message(text)
score = [2, 3]
show_score()
# -
# Si le score est entre 0 et 9 on peut l'afficher comme lettre, un après l'autre, dans la couleur respective du joueur.
# +
def show_score2():
"""Display score as two numbers in different colors."""
pass
show_score2()
# -
# Si le score est entre 0 et 8 on peut aussi l'afficher de façon graphique avec une barre.
# +
def show_score3():
"""Display score as two colored bars."""
pass
show_score3()
# -
# ## Cellules conséqutifs identiques
# Dans Morpion et Puissance 4 nous devons tester si une séquence de 3 ou 4 cellules sont identiques. Nous définissons une fonctions qui compare 3 ou 4 cellules à partir d'une position initiale et une direction.
def is_equal(T, x, y, dx, dy, p, n):
"""Check if n cells starting at (x, y),
in direction (dx, dy) are all equal to p"""
for i in range(n):
if T[y, x] != p:
return False
x += dx
y += dy
return True
# Testons notre function dans les 4 directions.
T = np.zeros((8, 8), int)
T[1:5, 3] = 1
T[7, 4:8] = 2
T[2, 0] = T[3, 1] = T[4, 2] = T[5, 3] = 3
T
print(is_equal(T, 3, 1, 0, 1, 1, 4))
print(is_equal(T, 4, 7, 1, 0, 2, 4))
print(is_equal(T, 0, 2, 1, 1, 3, 4))
def in_a_row(T, p, k):
"""Check if player p has k stones in a row."""
n, m = T.shape
# columns
for x in range(m):
for y in range(n-k+1):
if is_equal(T, x, y, 0, 1, p, k):
return True
# lines
for y in range(n):
for x in range(m-k+1):
if is_equal(T, x, y, 1, 0, p, k):
return True
# diagonals
for y in range(n-k+1):
for x in range(m-k+1):
if is_equal(T, x, y, 1, 1, p, k):
return True
if is_equal(T, x+k-1, y, -1, 1, p, k):
return True
return False
T = np.array([[1, 2, 2], [2, 2, 1], [1, 2, 1]])
print(T)
print(in_a_row(T, 2, 3))
# ## Fin du jeu
# Le jeu est terminé quand toutes les places sont occupées. La méthode `all()` est vrai si toutes les éléments sont différent de 0.
def is_finished(T):
"""Returns True if all cells are occupied."""
return T.all()
# On va tester la fonction avec deux cas.
T = np.ones((3, 3), int)
is_finished(T)
T[0, 0] = 0
is_finished(T)
# # Morpion
# Dans le jeu morpion les joueurs dovent placer en alternance une pièce sur un plateau de 3x3 cellules. Celui qui a placé en premier 3 pièces sur une ligne, colonne ou diagonale gagne.
n = 3
T = np.zeros((n, n), int)
print(T)
# La fonction `init` prépare le jeu.
def init(reset=False):
global x, y, p, n, score, colors, T
x, y = 1, 1
n = 3
T = np.zeros((n, n), int)
colors = [BLACK, BLUE, YELLOW]
if reset:
score = [0, 0]
p = 1
show_board()
show_cursor3()
# La fonction `show_board` affiche le tableau.
def show_board():
"""Show a 3x3 matrix on the 8x8 board."""
for x in range(8):
for y in range(8):
if x % 3 == 2 or y % 3 == 2:
sense.set_pixel(x, y, GRAY)
else:
val = T[y//3, x//3]
sense.set_pixel(x, y, colors[val])
def show_cursor3():
"""Scale the cursor from 3x3 matrix to 8x8 board."""
global x, y, p
col = colors[p] if T[y, x] != p else BLACK
sense.set_pixel(3*x, 3*y, col)
# The following implements the `play` function.
# +
def play(event):
"""Place a stone on the board if the position is empty."""
global x, y, p
if event.action == 'pressed':
move_cursor(event)
if event.direction == 'middle':
if T[y, x] == 0:
T[y, x] = p
if in_a_row(T, p, 3):
show_winner(p)
score[p-1] += 1
show_score3()
init()
elif is_finished(T):
show_winner(0)
show_score3()
init()
p = 3 - p
show_board()
show_cursor3()
sense.stick.__init__()
sense.stick.direction_any = play
init(True)
# -
# ## m, n, k game
# Le m, n, k game est joué sur un tableau m x n, par deux joueurs qui placent une pièce en alternance. Le but est de placer k pièces en ligne, colonne ou en diagonale.
#
# https://en.wikipedia.org/wiki/M,n,k-game
def init(reset=False, size=8):
global x, y, p, n, score, T
n = size
T = np.zeros((n, n), int)
x, y = 0, 0
if reset:
score = [0, 0]
p = 1
show_matrix(T)
show_cursor(x, y, p)
# +
def play(event):
"""Use 4 directions keys to move cursor, middle to play."""
global x, y, p
if event.action == 'pressed':
move_cursor(event)
if event.direction == 'middle':
if T[y, x] == 0:
pass
show_matrix(T)
show_cursor(x, y, p)
sense.stick.__init__()
sense.stick.direction_any = play
init(True)
# -
# ## Puissance 4
# Le but du jeu est d'aligner une suite de 4 pions de même couleur sur u
# ne
# grille comptant 7 rangées et 8 colonnes.
#
# Tour à tour les deux joueurs placent un pion dans la colonne de leur choix, le pion coulisse alors jusqu'à la position la plus basse possible dans la dite colonne à la suite de quoi c'est à l'adversaire de jouer.
#
# Le vainqueur est le joueur qui réalise le premier un alignement (horizontal, vertical ou diagonal) consécutif d'au moins quatre pions de sa couleur. Si, alors que toutes les cases de la grille de jeu sont remplies, aucun des deux joueurs n'a réalisé un tel alignement, la partie est déclarée nulle.
def init(reset=False):
"""Initialize global parameters for Connect4."""
global x, y, p, dt, colors, T
x, y = 3, 0
if reset:
score = [0, 0]
p = 1
dt = 0.2
colors = [BLUE, RED, YELLOW, BLACK]
T = np.zeros((8, 8), int)
T[0] = 3
show_matrix(T)
show_cursor(x, y, p)
init()
T
# Ceci sont les fonctions pour le jeu Puissance 4.
# +
def show(event):
if event.action == 'pressed':
show_matrix(T)
show_cursor(x, y, p)
def left(event):
"""Move cursor to the left."""
global x
if event.action == 'pressed':
x = max(x-1, 0)
def right(event):
"""Move cursor to the right."""
global x
if event.action == 'pressed':
x = min(x+1, 7)
def down(event):
"""Move disc downwards."""
global x, y, p
if event.action == 'pressed':
while (y < 7 and T[y+1, x] == 0):
y += 1
show_matrix(T)
show_cursor(x, y, p)
sleep(.1)
T[y, x] = p
if in_a_row(T[1:], p, 4):
show_winner(p)
show_score3()
init()
elif is_finished(T):
show_winner(0)
show_scores3()
init()
y = 0
p = 3 - p
sense.stick.__init__()
sense.stick.direction_left = left
sense.stick.direction_right = right
sense.stick.direction_down = down
sense.stick.direction_up = init
sense.stick.direction_middle = show_score()
sense.stick.direction_any = show
init()
# -
# # Tetris
# Définissons les formes des base: L, O, I.
L = np.array([[1, 0], [1, 1]], int)
print(L)
O = np.ones((2, 2), int)
print(O)
I = np.ones((3, 1), int)
print(I)
# ## Rotation des matrices
np.rot90(L)
np.rot90(L, 2)
np.rot90(L, 3)
np.rot90(I)
# Check if space is empty
def overlap(T, S, x, y):
pass
# Add a shape
def add(T, S, x, y):
"""Add shape S to T at (x, y)."""
n, m = S.shape
T[y:y+n, x:x+m] = S
add(T, L, 0, 0)
add(T, O*2, 5, 0)
add(T, L*3, 6, 0)
print(T)
show_matrix(T)
show_matrix(T)
def rem(T, S, x, y):
"""Remove shape S at (x, y)."""
n, m = S.shape
for i in range(n):
for j in range(m):
T[y+i, x+j] = 0
rem(T, O, 0, 0)
add(T, O, 1, 0)
show_matrix(T)
| games/morpion/Games2-morpion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %run '../00_rh_settings.ipynb'
# %run '../00_rh_load_processed.ipynb'
train_df = df
train_df.head(10)
df.apply(lambda x: x.isna().sum())
#df.apply(lambda x: x.isna().sum())
from src.models.train_model import CatBoostCustomModel
from catboost import CatBoostClassifier
df.drop(columns=['Unnamed: 0',"loanKey"], inplace=True, errors="ignore")
df.dropna(inplace=True)
# df = df.replace(r'\s+', np.nan, regex=True)
# df = df.replace('nan', np.nan)
# +
#objective(X=df.drop(columns="bad_flag"), y=df["bad_flag"], model=model,cat_features=categorical_features_indices, params={"classifier__metadata":{"test_value":"value"}})
# -
train_X = df.drop(columns="bad_flag")
train_y = df["bad_flag"]
# +
cbmft = CatBoostCustomModel(model=
CatBoostClassifier(iterations=250,
learning_rate=0.2,
depth=9,
l2_leaf_reg=5,
#custom_loss = ['Accuracy'],
#loss_function = 'MultiClass',
),
model_params={"metadata":{"test_value":"value"}} ,
)
cbmft.fit(train_X
, train_y
, fit_params={"logging_level": 'Silent', "plot": False}
#, n_folds = 3
)
cbmft.get_features_importance(sorted)
cbmft.plot_features_importance()
#list(cbmft.get("model").get_metadata())
# -
cbmft.get_score()
cbmft.get_crosstab()
from sklearn.externals import joblib
joblib.dump(cbmft, '../../models/scoring_model.joblib')
| notebooks/model/train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/gabrielborja/machine_learning/blob/main/02_regression_models.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="l_LulNCC8z96"
# # 01 - Simple Linear Regression
# + [markdown] id="NtvXTBMb276L"
# Asumptions of Linear Regression Models>
#
#
# 1. Linear relationship.
# 2. Multivariate normality.
# 3. No or little multicollinearity. Independent variables are not correlated with each other.
# 4. No auto-correlation. The residuals are not independent from each other.
# 5. Homoscedasticity meaning “same variance”. The error is the same across all values of the independent variables.
#
#
#
# + [markdown] id="xpXdowrE9DxW"
# ## Importing necessary libraries
# + id="8EkyiHx8G2FH"
#Optional upgrade of matplotlib
# !pip install matplotlib --upgrade
# + id="nhKd4hWx9GFt"
#Importing necessary libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + [markdown] id="6JhpWJi59J1p"
# ## Uploading data
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "Ly8gQ29weXJpZ2h0IDIwMTcgR29vZ2xlIExMQwovLwovLyBMaWNlbnNlZCB1bmRlciB0aGUgQXBhY2hlIExpY2Vuc2UsIFZlcnNpb24gMi4wICh0aGUgIkxpY2Vuc2UiKTsKLy8geW91IG1heSBub3QgdXNlIHRoaXMgZmlsZSBleGNlcHQgaW4gY29tcGxpYW5jZSB3aXRoIHRoZSBMaWNlbnNlLgovLyBZb3UgbWF5IG9idGFpbiBhIGNvcHkgb2YgdGhlIExpY2Vuc2UgYXQKLy8KLy8gICAgICBodHRwOi8vd3d3LmFwYWNoZS5vcmcvbGljZW5zZXMvTElDRU5TRS0yLjAKLy8KLy8gVW5sZXNzIHJlcXVpcmVkIGJ5IGFwcGxpY2FibGUgbGF3IG9yIGFncmVlZCB0byBpbiB3cml0aW5nLCBzb2Z0d2FyZQovLyBkaXN0cmlidXRlZCB1bmRlciB0aGUgTGljZW5zZSBpcyBkaXN0cmlidXRlZCBvbiBhbiAiQVMgSVMiIEJBU0lTLAovLyBXSVRIT1VUIFdBUlJBTlRJRVMgT1IgQ09ORElUSU9OUyBPRiBBTlkgS0lORCwgZWl0aGVyIGV4cHJlc3Mgb3IgaW1wbGllZC4KLy8gU2VlIHRoZSBMaWNlbnNlIGZvciB0aGUgc3BlY2lmaWMgbGFuZ3VhZ2UgZ292ZXJuaW5nIHBlcm1pc3Npb25zIGFuZAovLyBsaW1pdGF0aW9ucyB1bmRlciB0aGUgTGljZW5zZS4KCi8qKgogKiBAZmlsZW92ZXJ2aWV3IEhlbHBlcnMgZm9yIGdvb2dsZS5jb2xhYiBQeXRob24gbW9kdWxlLgogKi8KKGZ1bmN0aW9uKHNjb3BlKSB7CmZ1bmN0aW9uIHNwYW4odGV4dCwgc3R5bGVBdHRyaWJ1dGVzID0ge30pIHsKICBjb25zdCBlbGVtZW50ID0gZG9jdW1lbnQuY3JlYXRlRWxlbWVudCgnc3BhbicpOwogIGVsZW1lbnQudGV4dENvbnRlbnQgPSB0ZXh0OwogIGZvciAoY29uc3Qga2V5IG9mIE9iamVjdC5rZXlzKHN0eWxlQXR0cmlidXRlcykpIHsKICAgIGVsZW1lbnQuc3R5bGVba2V5XSA9IHN0eWxlQXR0cmlidXRlc1trZXldOwogIH0KICByZXR1cm4gZWxlbWVudDsKfQoKLy8gTWF4IG51bWJlciBvZiBieXRlcyB3aGljaCB3aWxsIGJlIHVwbG9hZGVkIGF0IGEgdGltZS4KY29uc3QgTUFYX1BBWUxPQURfU0laRSA9IDEwMCAqIDEwMjQ7CgpmdW5jdGlvbiBfdXBsb2FkRmlsZXMoaW5wdXRJZCwgb3V0cHV0SWQpIHsKICBjb25zdCBzdGVwcyA9IHVwbG9hZEZpbGVzU3RlcChpbnB1dElkLCBvdXRwdXRJZCk7CiAgY29uc3Qgb3V0cHV0RWxlbWVudCA9IGRvY3VtZW50LmdldEVsZW1lbnRCeUlkKG91dHB1dElkKTsKICAvLyBDYWNoZSBzdGVwcyBvbiB0aGUgb3V0cHV0RWxlbWVudCB0byBtYWtlIGl0IGF2YWlsYWJsZSBmb3IgdGhlIG5leHQgY2FsbAogIC8vIHRvIHVwbG9hZEZpbGVzQ29udGludWUgZnJvbSBQeXRob24uCiAgb3V0cHV0RWxlbWVudC5zdGVwcyA9IHN0ZXBzOwoKICByZXR1cm4gX3VwbG9hZEZpbGVzQ29udGludWUob3V0cHV0SWQpOwp9CgovLyBUaGlzIGlzIHJvdWdobHkgYW4gYXN5bmMgZ2VuZXJhdG9yIChub3Qgc3VwcG9ydGVkIGluIHRoZSBicm93c2VyIHlldCksCi8vIHdoZXJlIHRoZXJlIGFyZSBtdWx0aXBsZSBhc3luY2hyb25vdXMgc3RlcHMgYW5kIHRoZSBQeXRob24gc2lkZSBpcyBnb2luZwovLyB0byBwb2xsIGZvciBjb21wbGV0aW9uIG9mIGVhY2ggc3RlcC4KLy8gVGhpcyB1c2VzIGEgUHJvbWlzZSB0byBibG9jayB0aGUgcHl0aG9uIHNpZGUgb24gY29tcGxldGlvbiBvZiBlYWNoIHN0ZXAsCi8vIHRoZW4gcGFzc2VzIHRoZSByZXN1bHQgb2YgdGhlIHByZXZpb3VzIHN0ZXAgYXMgdGhlIGlucHV0IHRvIHRoZSBuZXh0IHN0ZXAuCmZ1bmN0aW9uIF91cGxvYWRGaWxlc0NvbnRpbnVlKG91dHB1dElkKSB7CiAgY29uc3Qgb3V0cHV0RWxlbWVudCA9IGRvY3VtZW50LmdldEVsZW1lbnRCeUlkKG91dHB1dElkKTsKICBjb25zdCBzdGVwcyA9IG91dHB1dEVsZW1lbnQuc3RlcHM7CgogIGNvbnN0IG5leHQgPSBzdGVwcy5uZXh0KG91dHB1dEVsZW1lbnQubGFzdFByb21pc2VWYWx1ZSk7CiAgcmV0dXJuIFByb21pc2UucmVzb2x2ZShuZXh0LnZhbHVlLnByb21pc2UpLnRoZW4oKHZhbHVlKSA9PiB7CiAgICAvLyBDYWNoZSB0aGUgbGFzdCBwcm9taXNlIHZhbHVlIHRvIG1ha2UgaXQgYXZhaWxhYmxlIHRvIHRoZSBuZXh0CiAgICAvLyBzdGVwIG9mIHRoZSBnZW5lcmF0b3IuCiAgICBvdXRwdXRFbGVtZW50Lmxhc3RQcm9taXNlVmFsdWUgPSB2YWx1ZTsKICAgIHJldHVybiBuZXh0LnZhbHVlLnJlc3BvbnNlOwogIH0pOwp9CgovKioKICogR2VuZXJhdG9yIGZ1bmN0aW9uIHdoaWNoIGlzIGNhbGxlZCBiZXR3ZWVuIGVhY2ggYXN5bmMgc3RlcCBvZiB0aGUgdXBsb2FkCiAqIHByb2Nlc3MuCiAqIEBwYXJhbSB7c3RyaW5nfSBpbnB1dElkIEVsZW1lbnQgSUQgb2YgdGhlIGlucHV0IGZpbGUgcGlja2VyIGVsZW1lbnQuCiAqIEBwYXJhbSB7c3RyaW5nfSBvdXRwdXRJZCBFbGVtZW50IElEIG9mIHRoZSBvdXRwdXQgZGlzcGxheS4KICogQHJldHVybiB7IUl0ZXJhYmxlPCFPYmplY3Q+fSBJdGVyYWJsZSBvZiBuZXh0IHN0ZXBzLgogKi8KZnVuY3Rpb24qIHVwbG9hZEZpbGVzU3RlcChpbnB1dElkLCBvdXRwdXRJZCkgewogIGNvbnN0IGlucHV0RWxlbWVudCA9IGRvY3VtZW50LmdldEVsZW1lbnRCeUlkKGlucHV0SWQpOwogIGlucHV0RWxlbWVudC5kaXNhYmxlZCA9IGZhbHNlOwoKICBjb25zdCBvdXRwdXRFbGVtZW50ID0gZG9jdW1lbnQuZ2V0RWxlbWVudEJ5SWQob3V0cHV0SWQpOwogIG91dHB1dEVsZW1lbnQuaW5uZXJIVE1MID0gJyc7CgogIGNvbnN0IHBpY2tlZFByb21pc2UgPSBuZXcgUHJvbWlzZSgocmVzb2x2ZSkgPT4gewogICAgaW5wdXRFbGVtZW50LmFkZEV2ZW50TGlzdGVuZXIoJ2NoYW5nZScsIChlKSA9PiB7CiAgICAgIHJlc29sdmUoZS50YXJnZXQuZmlsZXMpOwogICAgfSk7CiAgfSk7CgogIGNvbnN0IGNhbmNlbCA9IGRvY3VtZW50LmNyZWF0ZUVsZW1lbnQoJ2J1dHRvbicpOwogIGlucHV0RWxlbWVudC5wYXJlbnRFbGVtZW50LmFwcGVuZENoaWxkKGNhbmNlbCk7CiAgY2FuY2VsLnRleHRDb250ZW50ID0gJ0NhbmNlbCB1cGxvYWQnOwogIGNvbnN0IGNhbmNlbFByb21pc2UgPSBuZXcgUHJvbWlzZSgocmVzb2x2ZSkgPT4gewogICAgY2FuY2VsLm9uY2xpY2sgPSAoKSA9PiB7CiAgICAgIHJlc29sdmUobnVsbCk7CiAgICB9OwogIH0pOwoKICAvLyBXYWl0IGZvciB0aGUgdXNlciB0byBwaWNrIHRoZSBmaWxlcy4KICBjb25zdCBmaWxlcyA9IHlpZWxkIHsKICAgIHByb21pc2U6IFByb21pc2UucmFjZShbcGlja2VkUHJvbWlzZSwgY2FuY2VsUHJvbWlzZV0pLAogICAgcmVzcG9uc2U6IHsKICAgICAgYWN0aW9uOiAnc3RhcnRpbmcnLAogICAgfQogIH07CgogIGNhbmNlbC5yZW1vdmUoKTsKCiAgLy8gRGlzYWJsZSB0aGUgaW5wdXQgZWxlbWVudCBzaW5jZSBmdXJ0aGVyIHBpY2tzIGFyZSBub3QgYWxsb3dlZC4KICBpbnB1dEVsZW1lbnQuZGlzYWJsZWQgPSB0cnVlOwoKICBpZiAoIWZpbGVzKSB7CiAgICByZXR1cm4gewogICAgICByZXNwb25zZTogewogICAgICAgIGFjdGlvbjogJ2NvbXBsZXRlJywKICAgICAgfQogICAgfTsKICB9CgogIGZvciAoY29uc3QgZmlsZSBvZiBmaWxlcykgewogICAgY29uc3QgbGkgPSBkb2N1bWVudC5jcmVhdGVFbGVtZW50KCdsaScpOwogICAgbGkuYXBwZW5kKHNwYW4oZmlsZS5uYW1lLCB7Zm9udFdlaWdodDogJ2JvbGQnfSkpOwogICAgbGkuYXBwZW5kKHNwYW4oCiAgICAgICAgYCgke2ZpbGUudHlwZSB8fCAnbi9hJ30pIC0gJHtmaWxlLnNpemV9IGJ5dGVzLCBgICsKICAgICAgICBgbGFzdCBtb2RpZmllZDogJHsKICAgICAgICAgICAgZmlsZS5sYXN0TW9kaWZpZWREYXRlID8gZmlsZS5sYXN0TW9kaWZpZWREYXRlLnRvTG9jYWxlRGF0ZVN0cmluZygpIDoKICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgJ24vYSd9IC0gYCkpOwogICAgY29uc3QgcGVyY2VudCA9IHNwYW4oJzAlIGRvbmUnKTsKICAgIGxpLmFwcGVuZENoaWxkKHBlcmNlbnQpOwoKICAgIG91dHB1dEVsZW1lbnQuYXBwZW5kQ2hpbGQobGkpOwoKICAgIGNvbnN0IGZpbGVEYXRhUHJvbWlzZSA9IG5ldyBQcm9taXNlKChyZXNvbHZlKSA9PiB7CiAgICAgIGNvbnN0IHJlYWRlciA9IG5ldyBGaWxlUmVhZGVyKCk7CiAgICAgIHJlYWRlci5vbmxvYWQgPSAoZSkgPT4gewogICAgICAgIHJlc29sdmUoZS50YXJnZXQucmVzdWx0KTsKICAgICAgfTsKICAgICAgcmVhZGVyLnJlYWRBc0FycmF5QnVmZmVyKGZpbGUpOwogICAgfSk7CiAgICAvLyBXYWl0IGZvciB0aGUgZGF0YSB0byBiZSByZWFkeS4KICAgIGxldCBmaWxlRGF0YSA9IHlpZWxkIHsKICAgICAgcHJvbWlzZTogZmlsZURhdGFQcm9taXNlLAogICAgICByZXNwb25zZTogewogICAgICAgIGFjdGlvbjogJ2NvbnRpbnVlJywKICAgICAgfQogICAgfTsKCiAgICAvLyBVc2UgYSBjaHVua2VkIHNlbmRpbmcgdG8gYXZvaWQgbWVzc2FnZSBzaXplIGxpbWl0cy4gU2VlIGIvNjIxMTU2NjAuCiAgICBsZXQgcG9zaXRpb24gPSAwOwogICAgZG8gewogICAgICBjb25zdCBsZW5ndGggPSBNYXRoLm1pbihmaWxlRGF0YS5ieXRlTGVuZ3RoIC0gcG9zaXRpb24sIE1BWF9QQVlMT0FEX1NJWkUpOwogICAgICBjb25zdCBjaHVuayA9IG5ldyBVaW50OEFycmF5KGZpbGVEYXRhLCBwb3NpdGlvbiwgbGVuZ3RoKTsKICAgICAgcG9zaXRpb24gKz0gbGVuZ3RoOwoKICAgICAgY29uc3QgYmFzZTY0ID0gYnRvYShTdHJpbmcuZnJvbUNoYXJDb2RlLmFwcGx5KG51bGwsIGNodW5rKSk7CiAgICAgIHlpZWxkIHsKICAgICAgICByZXNwb25zZTogewogICAgICAgICAgYWN0aW9uOiAnYXBwZW5kJywKICAgICAgICAgIGZpbGU6IGZpbGUubmFtZSwKICAgICAgICAgIGRhdGE6IGJhc2U2NCwKICAgICAgICB9LAogICAgICB9OwoKICAgICAgbGV0IHBlcmNlbnREb25lID0gZmlsZURhdGEuYnl0ZUxlbmd0aCA9PT0gMCA/CiAgICAgICAgICAxMDAgOgogICAgICAgICAgTWF0aC5yb3VuZCgocG9zaXRpb24gLyBmaWxlRGF0YS5ieXRlTGVuZ3RoKSAqIDEwMCk7CiAgICAgIHBlcmNlbnQudGV4dENvbnRlbnQgPSBgJHtwZXJjZW50RG9uZX0lIGRvbmVgOwoKICAgIH0gd2hpbGUgKHBvc2l0aW9uIDwgZmlsZURhdGEuYnl0ZUxlbmd0aCk7CiAgfQoKICAvLyBBbGwgZG9uZS4KICB5aWVsZCB7CiAgICByZXNwb25zZTogewogICAgICBhY3Rpb246ICdjb21wbGV0ZScsCiAgICB9CiAgfTsKfQoKc2NvcGUuZ29vZ2xlID0gc2NvcGUuZ29vZ2xlIHx8IHt9OwpzY29wZS5nb29nbGUuY29sYWIgPSBzY29wZS5nb29nbGUuY29sYWIgfHwge307CnNjb3BlLmdvb2dsZS5jb2xhYi5fZmlsZXMgPSB7CiAgX3VwbG9hZEZpbGVzLAogIF91cGxvYWRGaWxlc0NvbnRpbnVlLAp9Owp9KShzZWxmKTsK", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 74} id="f8GfYDZ59O18" outputId="6322619a-c68e-411b-c7b5-59a0174bcdb5"
#Uploading dataset from local computer
from google.colab import files
uploaded1 = files.upload()
#Storing dataset in a Pandas Dataframe
import io
df1 = pd.read_csv(io.BytesIO(uploaded1['salary_data.csv']))
# + colab={"base_uri": "https://localhost:8080/"} id="wlJeXatYnHqp" outputId="2a626e9c-b030-4004-c3b8-7089fc8b876f"
#Checking the dataframe info
df1.info()
# + id="CbDV4vXEny3R"
#Extracting the samples matrix and target values
X = df1.iloc[:, :-1].values
y = df1.iloc[:, -1].values #==> Dependent variable vector
# + [markdown] id="AyhQaTwP9RzG"
# ## Splitting data between training and testing
# + id="bxOOauiN9VpC"
#Splitting between train and test sets
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
# + [markdown] id="ZijQwFMQ9itx"
# ## Training the Simple Linear Regression (SLR) model
# + colab={"base_uri": "https://localhost:8080/"} id="B4Aj_8YJ9l7J" outputId="dd6a76cc-e369-45ee-a89f-b439d889a876"
#Training the simple linear regression model
from sklearn.linear_model import LinearRegression
regressor1 = LinearRegression()
regressor1.fit(X_train, y_train)
# + [markdown] id="wa2T1Lq89o5H"
# ## Predicting values using the SLR model
#
#
# + id="Rn_sTJ2o9smm"
#Predicting the test sets results
y_pred = regressor1.predict(X_test)
# + [markdown] id="EUX1Vhsv97ZT"
# ## Visualizing the predicted values
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="Ze9vpBTf-Bol" outputId="53d0e0c5-44e6-49a8-c79a-9149bde44fee"
#Plotting the results from the predictions
plt.scatter(X_train, y_train, color='blue', s=50, label='Train')
plt.scatter(X_test, y_test, color='black', marker=r'$☂$', s=150, label='Test')
plt.plot(X_train, regressor1.predict(X_train), color = 'green', label='Predicted')
plt.title(f'Predicted values: y = {round(regressor1.coef_[0], 1)}*x + {round(regressor1.intercept_, 1)}')
plt.xlabel('X = Years of Experience')
plt.ylabel('y = Salary')
plt.legend()
plt.show()
# + [markdown] id="ad5hyftmwZXw"
# ##Checking the score of the SLR model
# + colab={"base_uri": "https://localhost:8080/"} id="a60qU_d8wb1J" outputId="f5f55bca-77f2-463e-e838-39a0ba869d25"
#Checking the score of the predicted values
round(regressor1.score(X_test, y_test), 3)
# + [markdown] id="YSyJ0d3hivsI"
# ##Making Single Predictions
# + id="ZEMJam_Oiuvg"
#Function to make a single prediction
def slr_predict(experience):
if type(experience)!=int:
return "Invalid input, use numeric value type"
elif experience < 0:
return "Experience must be greater than 0"
else:
return f'Predicted salary for {experience} years of experience is: $ {round(regressor1.predict([[experience]])[0], 2)}'
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="5p8INDk0jSxJ" outputId="afdf6fbd-4cf1-4345-db9f-c05fc2d29caf"
#Predicting numeric values from input
slr_predict(11)
# + [markdown] id="-IW7oONPiUeS"
# # 02 Multiple Linear Regression Model
# + [markdown] id="Y8vyr54sCN-l"
# 5 Methods for building Multiple Regression Models:
#
# 1. Backward Elimination
# 2. Forward selection.
# 3. Bidirectional Elimination.
# 4. Score comparison.
# 5. All-in. 2^N-1.
# + [markdown] id="42DQr9KS15Tz"
# ## Importing necessary libraries
# + id="VCSgQkZX15Tz"
#Importing necessary libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + [markdown] id="vuz-poNL15Tz"
# ## Uploading data
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 74} outputId="31160762-3a9c-429c-ef7a-289920711b08" id="8S3c4Rur15T0"
#Uploading dataset from local computer
from google.colab import files
uploaded2 = files.upload()
#Storing dataset in a Pandas Dataframe
import io
df2 = pd.read_csv(io.BytesIO(uploaded2['50_startups.csv']))
# + colab={"base_uri": "https://localhost:8080/"} outputId="7d1b97d1-a5f2-411a-c5f5-e4bbe0288469" id="QVuo3ePQ15T0"
#Checking the dataframe info
df2.info()
# + colab={"base_uri": "https://localhost:8080/"} id="0utfnCoZ7NyB" outputId="457409a1-06d1-4398-bf6f-2458996d0d4a"
#Checking the categorical values
df2['State'].value_counts()
# + id="pQ1LdRP715T0"
#Extracting the samples matrix and target values
X = df2.iloc[:, :-1].values
y = df2.iloc[:, -1].values #==> Dependent variable vector
# + colab={"base_uri": "https://localhost:8080/"} id="utAQXD3DEXHO" outputId="7ca70cbb-b09b-42f7-b44c-2dd6e18ca87c"
#Checking the array of samples matrix (features)
X[:5]
# + colab={"base_uri": "https://localhost:8080/"} id="ES8gGor7EYzO" outputId="6955125c-43d0-4f5c-f405-2163369e70b7"
#Checking the array of target values
y[:5]
# + [markdown] id="YtrHPYwrEm4G"
# ## Encoding categorical data
# + id="3noUXGzjEpz1"
#Applying One Hot Encoding to categorical features
from sklearn.preprocessing import OneHotEncoder
from sklearn.compose import ColumnTransformer
ct2 = ColumnTransformer(transformers=[('encoder', OneHotEncoder(), [3])], remainder='passthrough')
X = np.array(ct2.fit_transform(X))
# + colab={"base_uri": "https://localhost:8080/"} id="jmdisuoMVrlQ" outputId="b093c0fc-a8e3-4c45-ef95-27f921424ae7"
#Checking the array of samples matrix (features) after the transformation
X[:5]
# + [markdown] id="HoniAn4xWhd4"
# ## Splitting data for training and testing
# + id="zdykokEjWsaM"
#Splitting between train and test sets
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
# + [markdown] id="srcUQzPWX-pG"
# ## Training the Multiple Linear Regression (MLR) model
# + colab={"base_uri": "https://localhost:8080/"} outputId="a6daf6e6-de03-45ec-e913-6785ef19d978" id="16BGNjQgX-pX"
#Training the multiple linear regression model
from sklearn.linear_model import LinearRegression
regressor2 = LinearRegression()
regressor2.fit(X_train, y_train)
# + [markdown] id="rAHmGyhOX-pY"
# ## Predicting values using the MLR model
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 363} id="QzooEZm3cGPf" outputId="f84af0c2-3922-4b73-c52b-9f88d0eb6099"
#Predicting the test sets results
y_pred = regressor2.predict(X_test)
df2_pred = pd.DataFrame(data={'y_test': y_test, 'y_pred': y_pred})
df2_pred = df2_pred.assign(perc = 1-(df2_pred['y_pred']/df2_pred['y_test']))
df2_pred
# + [markdown] id="1CPQ5M_ifdZs"
# ## Visualizing the predicted values
# + colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="90db5be8-eca4-43c1-ce44-f1fe51a06dee" id="Oyx-i5ysfdZt"
#Plotting the results from the predictions
plt.scatter(range(1,11), y_test, color='green', s=50, label='Test')
plt.scatter(range(1,11), y_pred, color='red', s=50, marker=r'$\bigstar$', label='Predicted')
plt.title(f'Test vs. Predicted values - Multiple Linear Regression')
plt.xlabel('Num of tests')
plt.ylabel('Profit')
plt.legend(loc='upper left')
plt.show()
# + [markdown] id="C8Jh8pwzOWZt"
# # 03 Polynomial Linear Regression Model
# + [markdown] id="SMU9KvLGOWZv"
# 5 Methods for building Multiple Regression Models:
#
# 1. Backward Elimination
# 2. Forward selection.
# 3. Bidirectional Elimination.
# 4. Score comparison.
# 5. All-in. 2^N-1.
# + [markdown] id="hIlz_26wOWZw"
# ## Importing necessary libraries
# + id="x23XGwB9OWZw"
#Importing necessary libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + [markdown] id="ZkOrKtysOWZw"
# ## Uploading data
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 74} outputId="b9be0c70-df23-4b00-e2f4-6c7d207772f0" id="Awu1djLiOWZx"
#Uploading dataset from local computer
from google.colab import files
uploaded3 = files.upload()
#Storing dataset in a Pandas Dataframe
import io
df3 = pd.read_csv(io.BytesIO(uploaded3['position_salaries.csv']))
# + colab={"base_uri": "https://localhost:8080/"} outputId="77b6ef73-8b01-44d7-a80f-64b8ec5287bd" id="OSwfVHYjOWZy"
#Checking the dataframe info
df3.info()
# + colab={"base_uri": "https://localhost:8080/"} outputId="16933a22-03d0-4931-f5b4-6c36b1194c16" id="M2jSOTkkOWZy"
#Checking the categorical values
df3['Position'].value_counts()
# + id="u45OL_DHOWZz"
#Extracting the samples matrix and target values
X3 = df3.iloc[:, 1:-1].values
y3 = df3.iloc[:, -1].values #==> Dependent variable vector
# + colab={"base_uri": "https://localhost:8080/"} outputId="6241f06c-7d0c-4ef0-c7e5-7026507dc32c" id="cwQ-JkrnOWZz"
#Checking the array of samples matrix (features)
X3[:5]
# + colab={"base_uri": "https://localhost:8080/"} outputId="2d40e9b1-ece5-410d-ca1c-44c52dcbf05b" id="eElrP2l1OWZ0"
#Checking the array of target values
y3[:5]
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="g4PkDQvfQBdv" outputId="396d3f65-4da6-443d-c5a1-d611ff2b8624"
#Plotting the relationship between Position Level and Salary
plt.scatter(X3, y3, color='green', label='Salary')
plt.title('Position Level vs. Salary')
plt.xlabel('X = Position Level')
plt.ylabel('y = Salary')
plt.legend()
plt.show()
# + [markdown] id="93wozgWoRCfy"
# ## Building a Polynomail Linear Regression (PLR) model function
# + id="Q45u_dHcRH4s"
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import PolynomialFeatures
def polynomial_regression(X, y, deg):
"""
Gets as input an array of X features, array of y target values and number of degrees and
returns a dataframe with polynomial predictions upto the maximum number of degrees.
"""
poly_dict = {}
if (type(deg)!=int) | (deg < 1):
return 'Input must be integer higher than 1'
else:
deg_list = list(range(1,deg+1))
for i in deg_list:
poly_reg = PolynomialFeatures(degree=i)
X_poly = poly_reg.fit_transform(X)
lin_reg = LinearRegression()
lin_reg.fit(X_poly, y)
poly_dict.update({'deg_'+str(i): list(lin_reg.predict(X_poly))})
df = pd.DataFrame(data=poly_dict, index=X.flatten().tolist())
return df
# + [markdown] id="RyCyGoCciHBU"
# ## Training PLR models for different degrees
# + colab={"base_uri": "https://localhost:8080/", "height": 363} id="oeJqbVZbeLju" outputId="46c350de-9d26-4ba9-e260-8e971a6c5e75"
#Building a dataframe of models with different degrees of Polynomials
df3 = polynomial_regression(X=X3, y=y3, deg=5)
df3
# + [markdown] id="dl-ZmetqaOOL"
# ## Visualizing predictions
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="SmCm3xG9aRH1" outputId="75005751-e67f-4d36-c5a8-96d77be3af22"
#Plotting the predictions according to degrees
#plt.scatter(X3, y3, color='green', marker='+', s=100, label='Salary')
df3.plot()
plt.title('Polynomial Predictions: Position Level vs. Salary')
plt.xlabel('X = Position Level')
plt.ylabel('y = Salary')
plt.legend()
plt.show()
# + [markdown] id="HD6Miz_fq8yP"
# # 04 Support Vector Regression (SVR)
# + [markdown] id="-k-gWC2Kq8yQ"
# In most linear regression models, the objective is to minimize the sum of squared errors, for example Ordinary Least Squares (OLS). But what if we are only concerned about reducing error to a certain degree? In Support Vector Regression (SVR) we have the flexibility to define how much error ϵ (Epsilon) is acceptable in our model, in order to find an appropriate hyperplane to fit the data.
# + [markdown] id="Ly0Dr0Cyq8yR"
# ## Importing necessary libraries
# + id="8vVLhVP7q8yR"
# Importing necessary libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + [markdown] id="rr3JQMflq8yS"
# ## Uploading data
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 73} outputId="fefa01fd-593c-4f74-bb69-ff99a061a2b1" id="ThM_6rA3q8yS"
# Uploading dataset from local computer
from google.colab import files
uploaded4 = files.upload()
# Storing dataset in a Pandas Dataframe
import io
df4 = pd.read_csv(io.BytesIO(uploaded4['position_salaries.csv']))
# + colab={"base_uri": "https://localhost:8080/"} outputId="8c527ff1-9f67-42fa-a4b5-15d4d74d48d9" id="v-3amvZXq8yT"
# Checking the dataframe info
df4.info()
# + id="b2QW6TKoq8yU"
# Extracting the samples matrix and target values
X4 = df4.iloc[:, 1:-1].values
y4 = df4.iloc[:, -1].values #==> Dependent variable vector
# + colab={"base_uri": "https://localhost:8080/"} outputId="beef1c75-eea0-44df-e63a-552b34caf4a2" id="j6E_llqjq8yU"
# Checking the array of samples matrix (features)
X4[:5]
# + colab={"base_uri": "https://localhost:8080/"} outputId="4244a7a0-48b3-4a8c-e154-b01d0205e311" id="dKONM6N8q8yV"
# Checking the array of target values
y4[:5]
# + [markdown] id="rWh9-AWOJrzv"
# ## Initial transformations
# + colab={"base_uri": "https://localhost:8080/"} id="B-cl7gYzJuEu" outputId="0d670858-9455-4613-f04f-1c7fc0b19a2b"
# Transform the dependent variable to a 2D array
y4 = y4.reshape(len(y4), 1)
np.shape(y4)
# + [markdown] id="a4llu81CH7Be"
# ## Feature Scaling
# + id="7ibMj6iCH-_u"
# If feature scaling is not applied, the SVR will not work.
from sklearn.preprocessing import StandardScaler
sc4_X = StandardScaler() # => 2 StandardScaler object needed to compute the mean and STD independently.
sc4_y = StandardScaler()
X4 = sc4_X.fit_transform(X4)
y4 = sc4_y.fit_transform(y4)
# + colab={"base_uri": "https://localhost:8080/"} id="wIZ6FFK2MNKF" outputId="37b3e749-750f-4cf2-ebe6-96e54f08bd46"
# Checking the X train after scaling (-3 : +3)
X4
# + colab={"base_uri": "https://localhost:8080/"} id="mfbxseNrMO6l" outputId="0bd0bc2e-0aad-479e-9483-e635d92ec700"
# Checking the y train after scaling (-3 : +3)
y4
# + [markdown] id="ez-ZxnNjJKqo"
# ## Training the SVR model
# + colab={"base_uri": "https://localhost:8080/"} id="XcZlPiCyJKqp" outputId="8d94f8f4-f66b-4599-c6f6-bf0d0d3f38a8"
# Building the SVR model
from sklearn.svm import SVR
regressor_4 = SVR(kernel='rbf')
regressor_4.fit(X4, np.ravel(y4)) #=> Use ravel to change a 2D array to a flattened array
# + [markdown] id="tjYGMo1fJLNo"
# ## Predicting values
# + colab={"base_uri": "https://localhost:8080/"} id="KQc11IzlR2Yt" outputId="08118ef0-6dbb-4a4d-c0fc-dc3157240c6c"
# Predicting new values
sc4_y.inverse_transform([regressor_4.predict(sc4_X.transform([[6.5]]))]) # => Reversing the scaling before predictions
# + [markdown] id="OAtnOAlmJLrH"
# ## Visualizing SVR predictions
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="_0Fmcr8XJLrI" outputId="b7d82e01-dedf-43b7-af43-34ad6234ae19"
# Visualizing SVR predicted values of salary according to position levels
predictions_4 = regressor_4.predict(X4)
plt.plot(sc4_X.inverse_transform(X4), sc4_y.inverse_transform(y4), color='blue', label='Salary data', marker='+')
plt.scatter(sc4_X.inverse_transform(X4), sc4_y.inverse_transform(predictions_4.reshape(len(predictions_4), 1)), color='red',
label='SVR prediction', marker=r'$\bigstar$', s=100)
plt.title('SVR model predictions')
plt.xlabel('Position Level')
plt.ylabel('Salary')
plt.legend()
plt.show()
| 02_regression_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Machine Learning on graphs
# Here we review some simple yet powerful machine learning on graphs
import numpy as np
import networkx as nx
import community as community_louvain
import matplotlib.pyplot as plt
# # Clustering and community detection (unsupervised learning)
# Many real-world networks posesses communities, groups of nodes that are more connected together than with the rest of the network. Detecting these structures is of high importance. It reveals the hierarchy, the organization and interactions between nodes. It helps classifying parts of a network into categories. It can be seen as an equivalent of the unsupervised learning `k-means` clustering algorithm. Instead of computing distances and grouping data points in a high dimensional space, we use the network structure to detect the clusters.
#
# Many methods exists for community detection, see for example [this review](https://arxiv.org/abs/0906.0612). We will see two of them. The first one is available in `networkx`, the second one is the popular Louvain method which allows for fast computation.
# ## Girvan Newman algorithm
# The [GN algorithm](https://en.wikipedia.org/wiki/Girvan%E2%80%93Newman_algorithm) remove edges between communities in an iterative manner. The edges removed are the ones with the highest number of shortest paths passing through them ("bottlenecks" between communities). The idea is clear and intuitive. However, the computation is intensive as all shortest paths have to be computed. Moreover, the number of communities has to be specified as this algorithm do not have a stopping criterium (this can be an advantage or drawback).
# +
# Girvan Newman clustering
G = nx.path_graph(8)
num_clusters = 5 # desired number of clusters
k = num_clusters - 1
comp = nx.algorithms.community.centrality.girvan_newman(G)
comp = list(comp)
for idx in range(k):
print(' {} communities: {}'.format(idx+2, comp[idx]))
# Alternative way using directly the generator
#import itertools
#for idx,communities in enumerate(itertools.islice(comp, k)):
# print(' {} communities: {}'.format(idx+2, tuple(sorted(c) for c in communities)))
# -
# ## Exercise
# * Apply it with a more complex network and visualize the communities using Gephi. (See [graph list in networkx](https://networkx.org/documentation/stable/reference/generators.html), you may test on the "Karate club" graph or "Les Miserables" graph)
# * Try a larger network and experience the limit of scalability. What is a reasonable number of nodes for this method?
# ## Louvain community detection
#
# Community detection with Louvain method. You have to install an external module: `pip install louvain`, see [module Github page](https://github.com/taynaud/python-louvain) for more info or the [paper](https://arxiv.org/abs/0803.0476). This method is much more efficient than the previous one. It is a greedy, non-parametric, algorithm that finds automatically the optimal number of communities.
#Louvain module is called "community"
partition = community_louvain.best_partition(G)
#community_louvain.modularity(partition, G)
# `partition` is a dictionary where each node id is a key and its community is the value.
# re-order partition to have a dictionary of clsuters
clusters = {}
for i, v in partition.items():
clusters[v] = [i] if v not in clusters.keys() else clusters[v] + [i]
print(clusters)
# ## Label propagation (Semi-supervised learning)
# In this approach, the graph structure is combined to values (or feature vectors) associated to the nodes. Missing node values are found by propagating the known values to their neighbors.
L = nx.normalized_laplacian_matrix(G)
# We use label spreading from this [publication](http://citeseer.ist.psu.edu/viewdoc/summary?doi=10.1.1.115.3219) which perform a smoothing of the labels over the graph. We assume 2 classes with one-hot-encoding,i.e. feature vectors on nodes have dimension 2.
# labels
labels = np.zeros((L.shape[0],2))
labels[1,0] = 1 # node 1 has first label
labels[5,1] = 1 # node 5 has second label
print(labels)
def labelSpreading(G, labels, alpha, tol=1e-3):
L = nx.normalized_laplacian_matrix(G)
S = np.identity(L.shape[0]) - L.toarray()
max_iter = 1000
Y = np.zeros(labels.shape)
for i in range(max_iter):
Y_tmp = Y.copy()
Y = alpha * np.dot(S, Y) + (1 - alpha) * labels
if np.linalg.norm(Y-Y_tmp) < tol:
print('Converged after {} iterations.'.format(i))
break
return Y
smooth = labelSpreading(G,labels, 0.9)
print(labels)
print(smooth)
# Let us plot the results
# +
pos = nx.spring_layout(G, iterations=200)
plt.figure(figsize=(14, 6))
ax1 = plt.subplot(1, 2, 1)
ax2 = plt.subplot(1, 2, 2)
#ax3 = plt.subplot(1, 4, 3)
nx.draw(G, pos=pos, ax=ax1, node_color=smooth[:,0], cmap=plt.cm.Blues)
nx.draw(G, pos=pos, ax=ax2, node_color=smooth[:,1], cmap=plt.cm.Blues)
ax1.title.set_text("Smoothing of class 1 over the network")
ax2.title.set_text("Smoothing of class 2 over the network")
plt.show()
# -
propagated_labels = np.argmax(smooth,axis=1)
#label_dic = {k:v for k,v in enumerate(propagated_labels)}
print(propagated_labels)
nx.draw(G, pos=pos, node_color=propagated_labels, cmap=plt.cm.Blues)
plt.title("Final classification")
plt.show()
| 04_ML_on_graphs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from nanopub import Publication, NanopubClient
from rdflib import Graph, URIRef, RDF, FOAF
import pandas as pd
g_schema = Graph()
g_schema.load('../data/rdf/neurodkg.ttl', format="turtle")
qtext= """
PREFIX rdf: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX rdfs: <http://www.w3.org/2000/01/rdf-schema#>
PREFIX biolink: <https://w3id.org/biolink/vocab/>
PREFIX np: <http://www.nanopub.org/nschema#>
PREFIX npx: <http://purl.org/nanopub/x/>
prefix npa: <http://purl.org/nanopub/admin/>
PREFIX dct: <http://purl.org/dc/terms/>
PREFIX orcid: <https://orcid.org/>
PREFIX schema: <https://schema.org/>
PREFIX ndkg: <https://w3id.org/neurodkg/>
PREFIX prov: <http://www.w3.org/ns/prov#>
CONSTRUCT{
?association a schema:MedicalTherapy .
?subject a schema:Drug .
?object a schema:MedicalCondition .
?association schema:drug ?subject .
?object schema:possibleTreatment ?association .
?association ndkg:indicationType ?relation .
?association prov:wasDerivedFrom ?quote .
?quote prov:wasQuotedFrom ?publications .
?quote prov:value ?label .
?association schema:targetPopulation ?target_population .
?target_population a schema:PeopleAudience .
?target_population schema:audienceType ?target_pop_label .
?target_population schema:healthCondition ?phenotype .
?phenotype a schema:MedicalCondition .
?phenotype schema:signOrSymptom ?symptom .
?symptom a schema:MedicalCondition .
?target_population schema:requiredMinAge ?min_age .
?target_population schema:requiredMaxAge ?max_age .
}
WHERE{
?association schema:drug ?subject .
?object schema:possibleTreatment ?association .
?association ndkg:indicationType ?relation .
?association prov:wasDerivedFrom ?quote .
?quote prov:wasQuotedFrom ?publications .
?quote prov:value ?label .
OPTIONAL{
?association schema:targetPopulation ?target_population .
?target_population schema:audienceType ?target_pop_label .
OPTIONAL{
?target_population schema:healthCondition ?phenotype .
OPTIONAL{
?phenotype schema:signOrSymptom ?symptom .
}
}
OPTIONAL{
?target_population schema:requiredMinAge ?min_age .
}
OPTIONAL{
?target_population schema:requiredMaxAge ?max_age .
}
}
}
"""
results = g_schema.query(qtext)
g_biolink = Graph()
for r in results:
#print (r)
g_biolink.add(r)
g_biolink.serialize('../data/rdf/neurodkg_schema_1.ttl', format="turtle")
| notebook/Convert2Biolink.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
#
# Hello dear reader. This book is design for newcomers to Python with little knowledge on basic programming. Instructors can used it as a guide for teaching Python.
# ## Basics
#
# * **Central Processing Unit**: Very very fast but simple calculator. It is always asking what to calculate next?
# * **Main Memory**: Very fast temporary storage that save all instruction and data for CPU.
# * **Input Devices**: Devices that is used to give computer any kind of information, like Keyboards, Mouses, Touch Screens, Microphones, ... .
# * **Output Devices**: Devices that computer use to give information to you with them, like Screens, Speakers, Printers, ... .
# * **Secondary Memory**: If power goes down, main memory erased immediately. That's where secondary memory walks in. It's stores huge data and won't lost them without power. Why we shouldn't use secondary memory instead of main memory? Because it is too slow!
# ## Our Minds Vs. Computer Programs
#
# Read text below quickly:
#
# > Python has a deisgn philozophy which emphasizes code readebility (notably using whitespece indentetion to delimit code blucks rather than curly brackats or keywords), and a syntax which allows programmers to express concepts in fewer lines of code than might be used in langoages such as C++ or Java. The langoage provides constructs intended to enable writing clear progrems on both a small and large scale.
#
# There are many mistakes in the above text. Did you saw them? All of them?! If yes, good for you! If no, that is the most important difference between a computer and a human brain. Humans know how to solve problems automatically, we almost never see many of details, but computers are just fast and studpid calculators!
#
# Remember every thing that computer is doing should be told to it. Computers doesn't have a brain, they are just very advanced calculators.
# ## What is Python?
#
# [Python](https://www.python.org/) is an interpreted, general purposed language that is designed in 1991 by <NAME>. Python can be used for object oriented and functional programming. The language usage areas are tremendous. Some of them are, system programming, graphical user interfaces, game development, developing back-end, machine learning, visualizing data, media processing, scientific programming and many many more.
#
# Python have very readable and simple syntax to write code. Nowadays python frameworks and libraries are every where.
#
# There is only one drawback with python, runtime speed. But even this problem is almost solved by [PyPy](https://pypy.org/) project and many libararies like [numpy](http://www.numpy.org/) and [numby](https://numba.pydata.org/).
# ## What is PyCharm?
#
# As a developer we need an environment to code in it. The best environment that I found is [PyCharm](https://www.jetbrains.com/pycharm/), it has many features like auto-completion, debugging, version controller support, ... .
# ## What is Jupyter?
#
# An integrated development environment like pycharm isn't exactly what need for most of this course. We are dealing with watching effects of each line of code we write. [Jupyter](https://jupyter.org/) is what we use here.
#
# Python is a scripting language. This is an advantage that we will get familiar through the course.
# ## Hello Python
#
# There is a tradition in learning programming languages, called _Hello World_ program. We are going to respect the traditions and do it.
print('Hello Mickey Mouse!')
# ### Exercise
#
# * Make Python write you name
# # Comments
#
# Everything after a sharp will be commented out and will not considered by interpreter.
#
# You can even use strings as a comment.
# +
# A whole line commented out.
"This is a comment too!"
"""This is a comment with a little diference.
This one is a multiline comment.
Remember this strings are not saved in any variable!
"""
print('Hello <NAME>!') # Every thing after it will be commented, not before it.
print('Hello <NAME>!') # Your code should be separated from sharps with two space characters.
# -
# # PEP8
#
# Python enhancement proposal documents are about everything that apply to python. New features, community input on an issue, ... . Probably the most famous and useful one for us, is PEP8.
#
# Development is not single handed anymore, it's a social effort. Groups of many people try to write a code for a same software. PEP8 helps for readability of code for reusing by other people.
#
# Most imporant ones:
#
# * Indentations are 4 spaces.
# * Maximum line length is 80 or 120.
# * Break long line before operators.
#
# ```python
# # Yes: easy to match operators with operands
# income = (gross_wages
# + taxable_interest
# + (dividends - qualified_dividends)
# - ira_deduction
# - student_loan_interest)
# ```
#
# * Surround top-level function and class definitions with two blank lines.
# * Method definitions inside a class are surrounded by a single blank line.
# * Use seperate imports.
#
# ```python
# # This is good
# import os
# import sys
#
# # This is bad
# import os, sys
# ```
#
# * Whitespaces in Expressions and Statements.
#
# ```python
# # This is good
# spam(ham[1], {eggs: 2})
# foo = (0,)
# if x == 4: print x, y; x, y = y, x
#
# # This is bad
# spam( ham[ 1 ], { eggs: 2 } )
# bar = (0, )
# if x == 4 : print x , y ; x , y = y , x
#
# # This is good
# ham[1:9], ham[1:9:3], ham[:9:3], ham[1::3], ham[1:9:]
# ham[lower:upper], ham[lower:upper:], ham[lower::step]
# ham[lower+offset : upper+offset]
# ham[: upper_fn(x) : step_fn(x)], ham[:: step_fn(x)]
# ham[lower + offset : upper + offset]
#
# # This is bad
# ham[lower + offset:upper + offset]
# ham[1: 9], ham[1 :9], ham[1:9 :3]
# ham[lower : : upper]
# ham[ : upper]
# ```
#
# * Naming Conventions:
# * Modules should have short, all-lowercase, underscore (readability) separated names.
# * Packages should have short, all-lowercase names. Underscore are discouraged.
# * Class names follows CapWords convention.
# * Functions should have short, all-lowercase, underscore (readability) separated names.
# * Methods arguments:
# * Always use `self` and `cls`.
# * Group related function by one more blank line between each group.
# * Contants should have all-uppercase, underscore (readability) separated names.
# * Variables should have short, all-lowercase, underscore (readability) separated names.
# # Variables
#
# Variables are **labeled places** that you can store data in them. We use **all lowercase underscore separated** alphabets and numbers for variables. Never start a variable with a number. Python names are **case sensitive**. Variable names can't have spaces.
famous_disney_character = "<NAME>"
print(famous_disney_character)
# # Numbers and Mathematics
#
# Python have **infinite integers**, **booleans**, **floating points**, ... for representing numbers.
print (2 ** 4096)
# ## Arithmetics
#
# These operators are use for arithmetic purposes. Some of them have more priority upon others.
# +
# Arithmetic operators. Upper operators group have higher priority.
# Operators under a group have same priority.
print("2 ** 4: {}".format(2 ** 4)) # Power operator
print("5 * 6: {}".format(5 * 6)) # Multiplication operator
print("4 / 5: {}".format(4 / 5)) # Real division operator
print("10 // 3: {}".format(10 // 3)) # Integer division operator
print("25 % 7: {}".format(25 % 7)) # Reminder operator
print("23 + 45: {}".format(23 + 45)) # Addition operator
print("21 - 8: {}".format(21 - 108)) # Subtraction operator
number = 12 # Assignment operator
print("number: {}".format(number))
# -
# By using parentheses you can change order of operators.
num_1 = 24 + 2 * 12
num_2 = (24 + 2) * 12
print("num_1: {}, num_2: {}".format(num_1, num_2))
# Operators with same priority, get calculated from left to right.
num_3 = 2 * 3 / 4
num_4 = 2 / 3 * 4
print("num_3: {}, num_4: {}".format(num_3, num_4))
# ### Exercise
#
# * What is saved in variable `equation_result`?
#
# ```python
# equation_result = 5 // 3 + 2 ** 3 / 4 * 5 % 7
# ```
# ## Bigger is Better
#
# When you are doing arithmetic operations on numbers, the result is the type that have more precision.
print("35 / 5 = {}".format(35 / 5)) # Result is a floating point number
print("35.0 / 5 = {}".format(35 / 5)) # Result is a floating point number
print("35 / 5.0 = {}".format(35 / 5)) # Result is a floating point number
print("35.0 / 5.0 = {}".format(35 / 5)) # Result is a floating point number
print()
print("23 // 7 = {}".format(23 // 7)) # Result is an integer number
print("23.0 // 7 = {}".format(23.0 // 7)) # Result is a floating point number
print("23 // 7.0 = {}".format(23 // 7.0)) # Result is a floating point number
print("23.0 // 7.0 = {}".format(23.0 // 7.0)) # Result is a floating point number
print()
print("12 * 3 = {}".format(12 * 3)) # Result is an integer number
print("12.0 * 3 = {}".format(12.0 * 3)) # Result is an integer number
print("12 * 3.0 = {}".format(12 * 3.0)) # Result is an integer number
print("12.0 * 3.0 = {}".format(12.0 * 3.0)) # Result is an integer number
# # Names
#
# Each variable need a name.
# ## Reserved Words
#
# They have special meaning for python, so never use reserved words for variable, function, class, ... names!
#
# ```python
# from import as
# True False
# None
# and or not
# try except finally else raise assert
# while for break continue else in
# class
# def lambda return yield
# global nonlocal
# del
# if elif else pass is in
# with
# ```
# ## Readability
#
# There is a technique called **mnemonic**. It's about variables name that you choose. They should be **short**, **simple** and **sensible**. Variables names doesn't have any influence on how python execute your code, but it is very important for who wants to read your code!
#
# These three codes are the same thing to python. Python doesn't care about the meaning of your variables (Remember the fast, stupid calculator). It is just doing what we tell to do.
# +
# VERY VERY bad code!
hqpowiefhpqowi = 35.0
poiwhgpoiqf = 12.5
poiwhzpoiqf = hqpowiefhpqowi * poiwhgpoiqf
print(poiwhzpoiqf)
# Better, but it is not mnemonic.
a = 35.0
b = 12.5
c = a * b
print(c)
# Mnemonic code. You can understand it's purpose.
worked_hours = 35.0
pay_rate = 12.5
pay = worked_hours * pay_rate
print(pay)
# -
# # Strings
#
# String are a sequence of characters that surrounded with single/double quote.
book_author = '<NAME>'
book_title = "Python for You and Me"
# Multiline string are possible with triple single/double quote.
# +
book_description = '''
This book is about Python programming language.
I started this book as a guideline for instructors
and who wants to learn python not as a first programming language.'''
book_license = """
This book is under Creative Common license.
so feel free to contribute to it and read it."""
third_way = 'This is a\
very very long\
string'
print(book_description)
# -
print(book_license)
print(third_way)
# Attaching two strings is easy as sum them up.
# +
greetings = "Hello" + 'Goofy' + "!"
print("Greetings: " + greetings)
firstname = "Felonius"
lastname = "Gru"
fullname = firstname + ' ' + lastname
print('Full Name: {}, greeting: {}'.format(fullname, greetings))
# -
# Each string have a length that is count of characters in the string.
print(len(greetings))
print(len('')) # Empty String
# ## Format
#
# As you have seen, there is a `format` method for string objects that will try to make a new string. This method try to put arguments passed to it in `{}`.
#
# This is somehow like `printf` structure in `C/C++`.
print('You can put numbers: {},\
strings: {} in it to print.'\
.format(125, '#TEST#'))
# As we explain later, any object can get passed to a format method and get a string representation.
print('List: {}, Dictionary: {} and Set {}.'\
.format([110, 'Python'], {'a': 'hi', 'b': 12},\
{1, 2, 3, 2, 5, 1}))
name = 'ali'
print(name[1])
# ### Indexed
print('Second arg: {1}, First arg: {0},\
Second arg again: {1}'.format('FIRST', 'SECOND'))
# ### Named
print('First Name: {firstname}, Last Name: {lastname}'\
.format(firstname='Mike', lastname='Wazowski'))
# ### Template
#
# You can define a variable and put your template string in it and then get a proper string with `format` method.
# +
book_template = '''
{title}
Author: {author}
{description}
Book License: {license}
'''
print(book_template.format(author=book_author,
title=book_title,
description=book_description,
license=book_license))
# -
# ## Escape Sequences
#
# Python interpret `'` as start or end for a strings. What happen if want to use it as a apostrophe? You use an escape sequence. They have a special meaning to Python.
#
# |Escape Sequence|Meaning |
# |:-------------:|:-------------------------|
# |`\\` |Backslash (\\) |
# |`\'` |Single quote (') |
# |`\"` |Double quote (") |
# |`\a` |ASCII Bell (BEL) |
# |`\b` |ASCII Backspace (BS) |
# |`\f` |ASCII Formfeed (FF) |
# |`\n` |ASCII Linefeed (LF) |
# |`\r` |ASCII Carriage Return (CR)|
# |`\t` |ASCII Horizontal Tab (TAB)|
# |`\v` |ASCII Vertical Tab (VT) |
print("Baymax: \"On a scale of one to ten, how would you rate your pain?\"\nHiro Hamada: \"What?!\"")
# +
print('Student Results:\n\nName\tMark\nJodi\t80\nFrank\t30\nBob\t67')
# Above and below have the same output.
# In code below we use backsplash to tell python interpreter to ingnore newlines.
print('Student Results:\n\
\n\
Name\tMark\n\
Jodi\t80\n\
Frank\t30\n\
Bob\t67')
print('''Student Results:
Name\tMark
Jodi\t80
''')
# -
# ## Input from user
#
# For terminal applications, there is a built-in function called `input` that you can prompt user to get data.
string = input() # The output is always a string.
print(string)
name = input("What is your name? ") # You can ask user for special kind of information.
print("Hello {}.".format(name))
# ### Exercise
#
# Ask name and age of user and printing something nice for them.
# ## Indexing
#
# We said strings are created from characters. You can select characters in a string in very customize way.
the_string = 'Hi. This is a simple string that we will use as an example.'
# First of all you can access each character. Remember in Python counting starts from `0`.
print('1st character is: {}\n5th character is: {}'\
.format(the_string[0], the_string[4]))
# You want last character? No problem. Use negative index!
print('Last character is: {}\n5th character from end of the string is: {}'
.format(the_string[-1], the_string[-5]))
# String indexes are like:
#
# | H | e | l | l | o | | W | o | r | l | d | ! |
# |---|---|---|---|---|---|---|---|---|---|---|---|
# | 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10| 11|
# |-12|-11|-10|-9 |-8 |-7 |-6 |-5 |-4 |-3 |-2 |-1 |
# ## Slicing
#
# It's almost like indexing, but instead of selecting a single character, you can select mutiple characters. There is three things you use in here `Start`, `Stop`, `Step` and sperate them by a colon. In any of them missing, Python use their default value.
#
# |Variable|Default Value |
# |:------:|:----------------------|
# |Start |Beginning of the string|
# |Stop |End of the string |
# |Step |Equals to 1 |
print(the_string)
print(the_string[14:20]) # [start stop)
print(the_string[33:])
print(the_string[:27])
print(the_string[12:30:6])
print(the_string[::-1])
# ### Exercise
#
# Use message below and generate word **gun** with slice operator.
#
# ```python
# message = '"Simplicity is about subtracting the obvious and adding the meaningful." - <NAME>'
# ```
# ## `in` Operator
#
# Checks for existence of string in another string.
the_string_contains_hi = 'hi' in the_string
print(the_string_contains_hi)
greeting_contains_hi = 'hi' in 'Hi, how are you?'.lower()
print(greeting_contains_hi)
# # None
#
# This is a special object that represent `Nothing` in Python.
print(None)
# # Boolean
#
# Boolean types are actually intergers `0` and `1`. Boolean keywords are `True`, `False`.
print("What you see when you print\
true: {} and false: {}".format(True, False))
# We will see, any object can have a boolean equal. But for now look at code below.
print("Numbers: bool(814): {}, bool(-38): {}, bool(1.5): {}, bool(0): {}"
.format(bool(814), bool(-38), bool(1.5), bool(0)))
print("Strings: bool('A'): {},\
bool('Python is a wonderful language.'): {},\
bool(''): {}"
.format(bool('A'),\
bool('Python is a wonderful language.'),\
bool('')))
# All empty python datasets are evaluated to false.
print("0: {}, '': {}, []: {}, {}: {}, set(): {}"
.format(bool(0), bool(''), bool([]), '{}', bool({}), bool(set())))
# # Lists
#
# A list is very similar to an array. You can put any data in a list and access them with indexing and slicing.
primes = [2, 3, 5, 7, 11, 13, 17, 19, 23]
print("3rd Prime Number: {}".format(primes[2]))
print("Even indexed Prime Numbers: {}".format(primes[::2]))
print("Last Prime Number: {}".format(primes[-1]))
# Lists can have multiple data types at the same time! A list can have integers, floats, string and even other lists!
stuff = ['Apple', 12, [1, 2, 3], {'a': 'Alice', 'b': 'Bob'}]
# The `len` function return length of our list.
print(len(stuff))
# The difference between normal arrays and Python lists is that Python lists can change!
#
# Add new values with `append` method and remove value with `del` keyword.
primes.append(27)
print(primes)
primes.extend([9, 31, 37])
print(primes)
del primes[9:11]
print(primes)
primes.insert(9, 29)
print(primes)
# With method `pop` you can get the last element of list and remove it.
print(primes.pop())
print(primes)
# `remove`/`index` methods get an object, search for it in the list and remove it/return index of it.
primes.remove(2)
print(primes)
print(primes.index(17))
# Add to lists together and get new one.
positive_nums = [1, 2, 3, 4, 5]
negative_nums = [-1, -2, -3, -4, -5]
integers = negative_nums + [0] + positive_nums
print(integers)
sample_list = [1, 2, 'hi', 2, 'hello', 9, 3.14, 2]
print(sample_list.count(2))
# We have `in` operators for lists too.
[1, 2, 3] in integers
2 in integers
# ## Packing & Unpacking
#
# These two concepts come with sequences. Unpacking means separate each part of a sequence and packing means gather them together in a place.
#
# In Python we have multiple assignments.
name, age, color = 'Bugs Bunny', 77, 'White' # Mixed types. No type limitations.
print(name, age, color)
# You can even put multiple objects in a single variable.
character = 'Spongebob Squarepants', 18, 'Yellow'
print(character) # Character is a Tuple actually.
# If you have a long list and you wanted to get head and tail of it, you can use this syntax.
# +
students = ['Emma', 'Noah', 'Olivia', 'Liam', 'Sophia', 'Mason', 'Ava', 'Jacob', 'William', 'Isabella']
head, *students, tail = students
print('Head: {}, Students: {}, Tail: {}'.format(head, students, tail))
# -
# If you use a star before variable name in a left side of assignment, it means packing variables.
print(students)
print(*students) # Is equal to print('Noah', 'Olivia', ...)
# Packing and unpacking won't happen magically, you have to tell what python should do.
[1, 2, 3] == (1, 2, 3) # No packing or unpacking. No usage here.
# ## Comprehension
#
# There is a faster and better way to define a list.
squared = [num**2 for num in range(100)]
print(squared)
# ### To be or not to be
#
# A little more complex comprehension, let your code decide to add a value or not to add the value there.
squared_cubed = [num for num in squared if round(num ** (1/3)) ** 3 == num]
print(squared_cubed)
# ## Exercise
#
# Consider list below:
#
# ```python
# countries = ['USA', 'Germany', 'France', 'Iran']
# ```
#
# Write piece of code that change upper list to:
#
# ```python
# countries = ['UK', 'USA', 'France', 'Iran', 'Canada']
# ```
# +
countries = ['USA', 'Germany', 'France', 'Iran']
countries.insert(0, 'UK')
# countries = ['UK'] + countries
del countries[2]
# countries.remove('Germany')
countries.append('Canada')
print(countries)
# -
# # Tuples
#
# Tuples are frozen Lists. It means when you define a tuple you can't add to it or remove an object from it. Even you can't change a object at specific index! It is freezed for ever.
alvin_marks = (12, 10, 17, 14)
# Now is you pass alvin marks to any function or method or ... they can only read it and not changing it!
#
# Tuples are much like lists. You can add two tuples and get *new* tuple. You can search for an object in it. You can do anything lists can do with tuples as long as what you want to do won't change it.
woody_marks = (19, 18.5, 20, 17)
alvin_woody_marks = alvin_marks + woody_marks
print(alvin_woody_marks)
alvin_got_12 = 12 in alvin_marks
print("Alving got 12? {}".format(alvin_got_12))
# If you wanted to define a tupe with only one value in it, you should write it like:
a = (12,)
print(a)
# # Dictionaries
#
# They are like lists with a little differences. One of differences is that programmer defines indexes here! Indexes are called keys. A dictionary is like a rope, you can connect a keys to its value.
# +
per2eng_dict = {
'سلام': 'Hello',
'خوبی؟': 'How are you?',
'پایتون': 'Python',
}
print(per2eng_dict['خوبی؟'])
# -
# Adding new value to a dictionary is so simple. Choose your key and value.
#
# Using the same syntax for adding, you can update value of keys that exist.
per2eng_dict['خدانگهدار'] = 'Goodbye'
print(per2eng_dict['خدانگهدار'])
per2eng_dict['خوبی؟'] = 'How is it going?'
print(per2eng_dict['خوبی؟'])
# You can use any immutable datatype as a key for your dictionary.
# +
per2eng_dict[1396] = 2017
per2eng_dict[('شنبه', 'یک شنبه', 'دو شنبه')] = \
('saturday', 'sunday', 'monday')
print(per2eng_dict)
# -
# You can get all keys and values of a dictionary.
print(per2eng_dict.keys())
print(per2eng_dict.values())
# Another example for dictionaries is in simple cryptography algorithm called substitution.
# +
ciphertext = '''QRNE NYVPR,
V NZ N FCL NAQ JBEXVAT SBE HAVGRQ XVATQBZ. NF GUR SVEFG QNL V FGNEG ZL JBEX URER LBH JNF NYJNLF XVAQ GB ZR.
V JVYY ARIRE SBETRG LBHE URYCF.
FVAPRERYL, FNEN.'''
print(list(zip('ABCDEFGHIJKLMNOPQRSTUVWXYZ',
'NOPQRSTUVWXYZABCDEFGHIJKLM')))
# Generate a dictionary with comprehension
subs = {i: j for i, j in zip('ABCDEFGHIJKLMNOPQRSTUVWXYZ',
'NOPQRSTUVWXYZABCDEFGHIJKLM')}
#
# subs = {'J': 'W', 'Y': 'L', 'I': 'V',
# 'A': 'N', 'B': 'O', 'C': 'P',
# 'D': 'Q', 'W': 'J', 'F': 'S',
# 'U': 'H', 'Q': 'D', 'M': 'Z',
# 'P': 'C', 'E': 'R', 'K': 'X',
# 'Z': 'M', 'S': 'F', 'O': 'B',
# 'R': 'E', 'H': 'U', 'X': 'K',
# 'N': 'A', 'L': 'Y', 'G': 'T',
# 'V': 'I', 'T': 'G'}
# Characters that is not encrypted.
subs.update({'.': '.', ' ': ' ',
',': ',', '\n': '\n'})
plaintext = ''.join([subs[char] for char in ciphertext])
print(plaintext)
# -
# ## Comprehensions
#
# There is a better way to define dictionaries.
length_of_names = {name: len(name) for name in students}
print(length_of_names)
# Just like lists, here we can use `if` statement too.
long_names = {name: len(name) for name in students if len(name) >= 6}
print(long_names)
# # Set
#
# You can put unique object in a set.
numbers = {1, 5, 4, 1, 3, 2, 5, 2, 3, 5, 1}
print(numbers)
numbers.add(10)
print(numbers)
numbers.remove(3)
print(numbers)
even_nums = {2, 4, 6, 8, 10}
print(numbers & even_nums) #
print(numbers | even_nums)
print(numbers - even_nums)
# ## Comprehensions
#
# Faster way to define a set.
all_marks = {mark for mark in alvin_woody_marks}
print(all_marks)
# Again, you can use an `if`.
high_marks = {mark*5 for mark in alvin_woody_marks if mark >= 16} # We change marks scale from 0 -> 20 to 0 -> 100
print(high_marks)
# # Conversion
#
# There are function with same name as datatypes that try to convert you data from any datatype, to that you want.
# +
print(int(3.14))
print(int('-45'))
print(float(18))
print(float('3.14'))
print(list({1, 2, 3, 1, 4, 1, 2}))
print(set((10, 1, 5, 2, 9, 10, 4, 2, 1, 5)))
set2str = str({1, 2, 3, 1, 4, 1, 2})
print(type([]))
# -
# ## Mutable Vs. Immutable
#
# Some of datatypes are mutable and some of them are immutable.
#
# If a datatype is mutable, if you change a copy of mutable datatype, you will change original variable.
# +
shop_list = ['Egg', 'Milk']
new_shop_list = shop_list
new_shop_list.append('Sugar')
print(shop_list)
# -
# But if a datatype is immutable, if you change a copy of immutable datatype, you will not change original variable.
# +
book_title = "Python for You and Me"
new_book_title = book_title
new_book_title = "Python 4 U & Me"
print(book_title)
# -
# # Indentation
#
# Until now we had all of our code running sequentially. Now we need to group them, decide when to run what part of our code, or which part is going to be reused.
#
# To define sub-code, we use indentation. Indentation is a set of whitespaces before the actual code begins.
#
# ```python
# password = input('Enter your password: ')
#
# if password == '<PASSWORD>':
# print('Correct.')
# ```
#
# Considering PEP8 it if better to use 4 spaces for each step of indentation.
# # Decision Making
#
# There will be a time that you need to make a decision about age of a user, or correctness of a password, or information on a website and ... .
#
# Decision making in Python is done with this three statements:
# * `if CONDITION:`
# * `elif CONDITION:`
# * `else:`
# +
secret_knock = input('What is the secret knock? ')
if secret_knock == '1 2 1 3':
print('Correct, you can get in.') # This is indented code. This code is sub-code of if statement.
if secret_knock != '1 2 1 3':
print('Incorrect!')
# +
number = int(input('Enter an odd number: '))
if number % 2 == 1:
print('That is correct.')
else:
print('This is an even number!')
# +
age = int(input('How old are you? ')) # input function return datatype is always a string.
where_to_go = ''
if age < 7:
where_to_go = 'kindergarten'
elif age < 18:
where_to_go = 'school'
elif age < 120:
where_to_go = 'college'
else:
where_to_go = 'grave'
print('Then probabely you are going to {}.'.format(where_to_go))
# -
# ## Comparison Operators
#
# In above code, if user input is 5, then all of the conditions evaluate to true, but the first one will be used for printing. When you use `if`, `else` or `if`, `elif`, `else` there is a priority for conditions. If one of upper conditions comes true, none of conditions below the true one will be check or executed.
#
# Python have six comparison operators:
#
# |Operator| Meaning |
# |:------:|:--------------------------------------|
# |`a < b`|`a` is smaller than `b` |
# |`a <= b`|`a` is smaller than **or** equal to `b`|
# |`a > b`|`a` is bigger than `b` |
# |`a >= b`|`a` is bigger than **or** equal to `b` |
# |`a == b`|`a` is equal to `b` |
# |`a != b`|`a` is not equal to `b` |
#
# ## Combining Conditions
#
# You can combine conditions together with `and` and `or` operators.
#
# | Condition |Result|
# |:-----------------:|:----:|
# |True **and** True | True |
# |True **and** False| False|
# |False **and** True | False|
# |False **and** False| False|
# +
ali_age, ali_school, ali_friend = \
17, 'high school', {'mohammad', 'mahdi'}
amir_age, amir_school, amir_friend = \
16, 'high school', {'reza', 'mohammad'}
if ali_school == amir_school and ali_friend & amir_friend:
print('Ali and Amir are friends.')
# -
# | Condition |Result|
# |:----------------:|:----:|
# |True **or** True | True |
# |True **or** False| True |
# |False **or** True | True |
# |False **or** False| False|
if ali_school == amir_school or ali_friend & amir_friend:
print('Ali and Amir are probabely friends.')
# ## Chaining
#
# There is a very useful feature named **Chained comparsion**. In most programming languages like C, C++, Java, PHP, Drupal, ... this is not possible.
if 5 >= 2 <= 3:
print('Correct!')
if -1 <= (ali_age - amir_age) <= 1:
print('Ali and Amir have same age range.')
# ## Condition Operator
#
# It like any operators, it have operands and a result.
# +
age = int(input('How old are you? '))
print('You are {} young.'\
.format('so' if age < 25 else 'not so'))
# -
# ## Special Cases
#
# If you were going to check if a variable is `None` or `True` or `False`, considering PEP8, you have to use `is` operator.
# +
information_about_bermuda_triangle = None
if information_about_bermuda_triangle is None:
print('We don\'t know anything.')
rainy_day = False
if rainy_day is not False:
print('Use an umbrella.')
# -
# There is another operator that you can use in condition.
# +
name = input('What is your name? ') # Mason
if name in students:
print('Your name is on the list.')
else:
print('Your name is not on the list!')
# -
# ## Objects as Condition
#
# Any object is evaluated to `True` or `False`. In Python all empty built-in datatypes are evaluated to `False` and in other situations evaluated to `True`. As we told, `bool` function get the object and return evaluated result.
if [False]:
print('List is true.')
else:
print('List is false.')
# # Loops
#
# When you need to do repetitive tasks you have to use loops.
#
# Python have `for` and `while` loop.
#
# ## For
#
# For loop is almost used for iterating over a sequence.
# +
shop_list = ['Egg', 'Milk', '...']
for item in shop_list:
print('We should buy {}.'.format(item))
print('This is another print function.')
# +
number = int(input('Which factorial number to calculate? '))
total = 1
for i in range(1, number + 1):
total *= i
print('Factorial of {} is {}.'.format(number, total))
# -
# ### Exercise
#
# * Get a number from user and print all prime numbers less than that number.
# +
biggest_num = int(input('Enter a number to get all prime numbers less than that: '))
primes = []
for num in range(2, biggest_num):
is_prime = True
for prime in primes:
if num % prime == 0:
is_prime = False
if is_prime:
primes.append(num)
print(primes)
# +
total = 1
for prime in primes:
total *= prime
print(total)
# -
for key in per2eng_dict:
print('With key {} we get value {}.'\
.format(key, per2eng_dict[key]))
# ## While
#
# This loop is used for situation when we don't know exactly when we should stop.
# +
from random import randint
selected_num = randint(1, 1000)
num = 1
while num != selected_num:
num = int(input('Guess a number: '))
if num < selected_num:
print('Smaller!')
elif num > selected_num:
print('Bigger!')
else:
print('BINGO')
# -
password = input('Enter the password: ')
while password != '<PASSWORD>':
password = input('Password was incorrect!\nPlease enter correct password: ')
# ## Break
#
# Maybe you have complex situation that need to be checked and if they come true, the loop should stop. For this purpose you can use `break` inside a loop.
# +
print('Enter numbers to get total, finish with 0.')
total = 0
while True:
num = int(input('Enter your number: '))
if num == 0:
break
total += num
print('Total: {}'.format(total))
# -
# ## Continue
#
# If you wanted to skip rest of a loop execution, you can use `continue`.
# +
print('Enter number to get total, finish with 0. Negative numbers are ignored.')
total = 0
while True:
num = int(input('Enter your number: '))
if num < 0:
continue
elif num == 0:
break
total += num
print('Total: {}'.format(total))
# -
# ## Comprehensions
#
# To create some of built-in datatypes you need use loops. For example to get students names and put them in a list.
# +
students = []
for i in range(5):
student_name = input('Enter a name: ')
students.append(student_name)
print(students)
# -
students = [input('Enter a name: ') for i in range(5)] # Shorter and faster way to do it.
print(students)
# ### Exercise
#
# Get a number from user and create a list with comprehension that contains square of numbers in range [1, user_number).
# +
user_num = int(input('Enter a number: '))
numbers = [num**2 for num in range(1, user_num + 1)]
print(numbers)
# -
# You can use comprehension with lists, set and dictionary.
num_count = int(input('How many number you have? '))
unique_nums = {int(input('Enter a number: '))\
for i in range(num_count)}
print(unique_nums)
num_of_words = int(input('How many words you want in your dictionary? '))
eng2per = {input('Enter english word: '):\
input('Enter persian translation of your word: ')\
for i in range(num_of_words)}
print(eng2per)
# ### Omiting
#
# Maybe you don't want all of the thing you have, get into your list, set or dictionary, well you can use if statement for that in comprehensions.
words = ["Trip", "restaurant", "GOOD", "reviews", "expectations", "hIGh","service", "SLOW", "full"]
lowercase_words = [word for word in words if word.islower()]
print(lowercase_words)
# ## Nesting
#
# You can nest loops for complex results like multiplication table.
# +
from pprint import pprint
size = int(input('Enter size of multiplication table: '))
mul_tab = []
for row in range(1, size + 1):
mul_tab.append([])
for column in range(1, size + 1):
mul_tab[-1].append(row * column)
pprint(mul_tab)
print()
# -
# You can nest loops with comprehension too!
# +
from pprint import pprint # Better way
size = int(input('Enter size of multiplication table: '))
mul_tab = [[row*col for col in range(1, size + 1)] for row in range(1, size + 1)]
pprint(mul_tab)
# -
# Another way of nesting
mul_tab_nums = {num for row in mul_tab for num in row}
print(mul_tab_nums)
# ## Exercises
#
# * Write a program and generate fibonacci numbers.
# * `1 1 2 3 5 8 13 21 34 55 ...`
# * Find greatest common divisor of two positive numbers.
# * GCD of A and B == GCD of B and A%B (A>=B)
# * Generate a triangle with comprehensions.
#
# ```python
# *
# **
# ***
# ****
# *****
# ******
# ```
# +
fibo_num = [1, 1]
fibo_count = int(input('Enter fibo count: '))
# for i in range(2, fibo_count):
# fibo_num.append(fibo_num[-1] + fibo_num[-2])
nothing = [fibo_num.append(fibo_num[-1] + fibo_num[-2])
for i in range(2, fibo_count)]
print(nothing)
print(fibo_num)
# +
num1, num2 = int(input('Enter first number: ')), int(input('Enter second number: '))
while num1 % num2 != 0:
num1, num2 = num2, num1 % num2
print(num2)
# -
names = ['Jodi', 'Alice', 'Bob']
print(''.join(names))
# +
size = int(input('Size of triangle: '))
for row in range(1, size + 1): # [start, stop)
stars = []
for i in range(row):
stars.append('*')
# print(stars)
print(''.join(stars))
# print('*' * row)
# print('\n'.join(['*' * row
# for row in range(1, size + 1)]))
# -
# # Functions
#
# Some parts of code is going to be reused again and again and again, we can't write them again and again and again! Remember don't repeat yourself (**DRY**). Function are reusable block of code that make coding easy. We can split tasks to function and use them any where. Functions improve readability and hide complexity.
#
# Generally in Python function take zero, one or more input (called arguments) and return exactly one output (called returned value).
# +
def greeting(name):
print('Hello {}'.format(name))
greeting('Alice')
print(greeting('Jack'))
# -
# Each function should have a very specific job and should not alter anything that is not necessary. So we can rewrite above code.
# +
def greeting(name):
return 'Hello {}'.format(name)
print(greeting('Alice'))
print(greeting('Bob'))
# -
# ## Arguments
#
# A function can take multiple arguments.
# +
from pprint import pprint
def multiplication_table(row, column):
return [[i*j for j in range(1, column + 1)] for i in range(1, row + 1)]
pprint(multiplication_table(4, 8))
# -
# ### Pass by Value or Refrence
#
# Mutable and immutable values behave different when passed to a function. Mutable variables is pass by refrence but immutable variables is pass by value.
# +
def add_one(a_list): # Mutable
a_list.append(1)
num_list = [3, 2, 9]
add_one(num_list)
print(num_list)
def add_one(a_string): # Immutable
a_string += 'ONE'
print('a_string is "{}"'.format(a_string))
numbers = 'TWO, '
add_one(numbers)
print(numbers)
# -
# ### Labeled Arguments
#
# You can explicitly pass an argument by using name of that argument.
pprint(multiplication_table(column=10, row=3))
# ### Default Values
#
# Function arguments can have default values. Remember all arguments with default values should be last arguments.
# +
def multiplication_table(row=10, column=10):
return [[i*j for j in range(1, column + 1)] for i in range(1, row + 1)]
pprint(multiplication_table()) # Use default values
# -
pprint(multiplication_table(5, 5)) # Don't use default values
pprint(multiplication_table(3)) # Pass first argument and use second default value
pprint(multiplication_table(column=3)) # Even pass second value and use first default value
# ### List Arguments
#
# Function can get arguments of any count.
# +
def total_of(*numbers):
total = 0
for number in numbers:
total += number
return total
print(total_of(1, 2, 3))
# -
print(total_of(9018, 8712, 2376, 19, 268, 683, 1912875))
print(total_of())
# ### Keyword Arguments
#
# Function arguments even can be labeled.
# +
def keywords(**kwargs):
return(kwargs)
print(keywords(hi='سلام', hello='سلام', goodbye='خداحافظ'))
# -
# Function arguments have order to use, positional args, list args, keyword args.
# +
def function(first, second, third=3, fourth='four', *fifth, **sixth):
print('First: {}'.format(first))
print('Second: {}'.format(second))
print('Third: {}'.format(third))
print('Fourth: {}'.format(fourth))
print('Fifth: {}'.format(fifth))
print('Sixth: {}'.format(sixth))
function(1, 2, 3, 4, 5, 6, test=123)
# -
function(1, 2)
function(1, 2, test='keyword')
# ## Return
#
# Each function always return exactly one data even when you don't return anything!
# +
def no_return():
pass
result = no_return()
print(result)
# -
# With Python function you can simply return multiple values. They will be pack as a single tuple and returned.
# +
def multiple_value_returned():
return '<NAME>', 11, ['Mater', '<NAME>', '<NAME>', '<NAME>']
values = multiple_value_returned() # Single variable to store.
name, age, partners = multiple_value_returned() # Multiple variables to store.
print("Values: {}".format(values))
print("Name: {}, Age: {}, Partners: {}".format(name, age, partners))
# -
# ## Scope
#
# Each function have its one scope of variables.
# +
name = 'Tonto'
if name == 'Tonto':
age = 12
print('Age: {}'.format(age))
while age == 12:
age += 1
shopping_list = ['Milk', 'Egg']
print('Shopping List: {}'.format(shopping_list))
def func():
print('Func -> Age: {}'.format(age))
# print('Func -> Shopping List: {}'.format(shop_list))
var_in_func = 'Function'
func()
print(var_in_func)
# -
# ### Global and Nonlocal
# ## Documentation
#
# Packages, modules, classes and functions all have docstrings. For complete information read docstring conventions, [PEP257](https://www.python.org/dev/peps/pep-0257/).
#
# A triple double quoted string at first line of the function is a docstring.
# +
def multiplication_table(row=10, column=10):
"""Get number of rows and columns of multiplication table then generate and returns a two dimensions list."""
return [[i*j for j in range(1, column + 1)] for i in range(1, row + 1)]
help(multiplication_table)
# -
# ## lambda
#
# We can define nameless functions.
square = lambda x: x**2
print(square(10))
# Everything about function arguments works with lambda functions too.
print((lambda a, b=3, **kwargs: (a, b, kwargs))('Hello', keyword='argument'))
# ## Recursive Functions
#
# If you call a function inside itself, it is called recursive function. Recursive function are useful for solving some complex algorithmic problems.
# +
def factorial(n):
if n == 0:
return 1
return factorial(n - 1) * n
print(factorial(10))
# +
def move_disks3(n, source, intermediate, destination):
"""Print movee needed to move disks in source tower to destination tower.
@param int n: Number of disk in source tower.
@param int source: Name of the source tower.
@param int intermediate: Name of intermediate tower.
@param int destination: Name of destination tower.
@rtype: None
"""
if n > 1:
move_disks3(n - 1, source, destination, intermediate)
move_disks3(1, source, intermediate, destination)
move_disks3(n - 1, intermediate, source, destination)
else:
print("{} -> {}".format(source,destination))
move_disks3(3, 'First', 'Second', 'Third')
# -
# ### Exercise
#
# * GCD recursive function.
# * All prime number between two intervals.
# +
def gcd(num1, num2):
"""This function calculate GCD of two numbers with recursive function."""
return gcd(num2, num1%num2) if num2 else num1
print(gcd(45, 33))
# +
def prime_range(start, stop):
primes = []
for number in range(start, stop):
is_prime = True
for i in range(2, number):
if number % i == 0:
is_prime = False
break
if is_prime:
primes.append(number)
return primes
for prime in prime_range(1000, 1050):
print('{} is a prime number.'.format(prime))
# +
# (stop)
# (start, stop)
# (start, stop, step)
def prime_range(start=None, stop=None, step=None):
"""A function with parameter that is completely like
range function but instead of generate natural numbers
here the function generates prime numbers."""
# Evaluate start, stop and step by what user of this
# function code want it to do.
if start is None:
print('WRONG')
return
elif stop is None:
start, stop, step = 2, start, 1
elif step is None:
step = 1
# A list to store all primes that I found.
primes = []
# Generate all primes from start to stop.
for number in range(start, stop):
is_prime = True
for i in range(2, number):
if number % i == 0:
is_prime = False
break
if is_prime:
primes.append(number)
# Slice primes with step that user wants and return
# it.
return primes[::step]
for prime in prime_range(1000, 1050):
print('{} is a prime number.'.format(prime))
# -
print(prime_range(10))
print(prime_range(5, 15))
print(prime_range(10, 100, 5))
def prime_range(*args):
"""A function with parameter that is completely like
range function but instead of generate natural numbers
here the function generates prime numbers."""
# Evaluate start, stop and step by what user of this
# function code want it to do.
if len(args) == 1:
start, stop, step = 2, args[0], 1
elif len(args) == 2:
start, stop, step = args[0], args[1], 1
elif len(args) == 3:
start, stop, step = args
else:
print('WRONG')
return
# A list to store all primes that I found.
primes = []
# Generate all primes from start to stop.
for number in range(start, stop):
is_prime = True
for i in range(2, number):
if number % i == 0:
is_prime = False
break
if is_prime:
primes.append(number)
# Slice primes with step that user wants and return
# it.
return primes[::step]
# ## Built-in Functions
#
# Python have built-in functions to help easy coding.
# ### help
#
# Print a documentation on object passed to it.
help(help)
# ### print
#
# We work with this one so many times. Print function have 3 labeled arguments:
# * `end`: Specify how to end a printed text.
# * `sep`: Specify separation of each argument passed to `print` function.
# * `file`: Specify a file to print text into that file.
print(1, 2, 3)
print(1, 2, 3, sep='\n')
print(1, 2, 3, sep=', ', end='.')
print('Hello')
# ### len
#
# Get length of an object. You can define length of each object you define. Sequences and collections have length.
# ### bool
#
# Return evaluted boolean value of an object. You can define boolean value of objects you define.
# ### str
#
# Convert object passed to it, to string representaion.
# ### int
#
# Convert object passed to it, to integer values.
# ### set
#
# Convert a sequence passed to it, to a set.
# ### list
#
# Convert a sequence passed to it, to a list.
# ### dict
#
# Defining a dictionary with keyword argument of a function.
marks = dict(Joe='A', Bob='C', Alice='B+')
print(marks)
# ### tuple
#
# Convert a sequence passed to it, to a tuple.
# ### range
#
# Get three arguments, `start`, `stop` and `step` and return an iterable object that count from start (including) to stop (excluding) with specified steps.
print(list(range(10)))
print(list(range(1, 10)))
print(list(range(1, 10, 3)))
# ### zip
#
# Get any number of sequences and work exactly like a zip!
print(list(zip([1, 2, 3], ('one', 'two','three'))))
print(list(zip(range(1, 5), {'this', 'is', 'a', 'set'})))
print(list(zip([1, 2, 3], ['1', '2', '3'], ['one', 'two', 'three'])))
# ### map
#
# You should give one function with `N` arguments, as the first argument of `map` function and then give it `N` sequences with same length. It will calculate function with those and save the output as a new sequence.
print(list(map(lambda x, y: x**y, [1, 2, 3], [10, 20, 30])))
# ### filter
#
# As the name says, it is used to filter some of object out. The first parameter would be a function that if returns `True` the object stays, it returns `false` the object is filtered.
# +
numbers = list(range(1, 20))
print(numbers)
print(list(filter(lambda x: x%2, numbers)))
# -
# ### min/max
#
# Get a sequence and compare each of objects inside the sequence with each other and return minimum/maximum value.
greeting ='Hello World!'
print('Min: "{}", Max: "{}"'.format(min(greeting), max(greeting)))
numbers = [12, 6, 47, -52, 158]
print('Min: {}, Max: {}'.format(min(numbers), max(numbers)))
numbers = {12, 6, 47, -52, 158}
print('Min: {}, Max: {}'.format(min(numbers), max(numbers)))
marks = {'John': 18, 'Sophia': 12, 'Sara': 16.7}
print('Min: {}, Max: {}'.format(min(marks), max(marks)))
# ## Useful Tips
# * Functions should do one thing.
# * Functions should be small.
# * The fewer input parameters, the better.
# * Functions should be consistent in their return values.
# Functions shouldn't have side effects.
# ## Exercise
#
# * Create a function that take a number of digit for precision and calculate $\pi$ with formula below.
#
# $$\frac{\pi}{2} = 1 + \frac{1}{3} + \frac{1 \times 2}{3 \times 5} + \frac{1 \times 2 \times 3}{3 \times 5 \times 7} + \dots$$
#
# Default precision should be 5 digits. You have to write proper documentation for your function.
#
# * Print factors of a number (You will need a prime number generator function).
#
# $$ 12 = 2^2 \times 3^1 $$
#
# ```python
# [(2, 2), (3, 1)]
# ```
# +
def fraction(frac_num):
result = 1.0
for i in range(1, frac_num + 1):
result *= i/(2*i+1)
return result
def pi(precision=5):
total = 0.0
i = 0
while fraction(i) > 10**-precision:
total += fraction(i)
i += 1
return total * 2
print(pi(8))
# +
def factors(number):
result = []
for prime in prime_range(number + 1):
# Find the right superscript for specific prime number.
superscript = 0
while number % prime == 0:
number /= prime
superscript += 1
# If superscript is bigger that zero, we add it to result
if superscript:
result.append((prime, superscript))
return result
print(factors(2017))
# -
print(factors(21000))
# # Files
#
# All data we have is store in files. Even this document you are reading is store in a file. Python have very good API and very very libraries to work with files.
# +
# Simple reading from file.
hello = open('hello')
print(hello.read())
print(hello.read())
hello.close()
# -
# The `open` function first argument is a path to file, second argument is to specify which mode to open the file.
#
# Modes to open a file is:
#
# |Character| Meaning |
# |:-------:|:--------------------------------------------------------------|
# | r |open for reading (default) |
# | w |open for writing, truncating the file first |
# | x |open for exclusive creation, failing if the file already exists|
# | a |open for writing, appending to the end of the file if it exists|
# | b |binary mode |
# | t |text mode (default) |
# | + |open a disk file for updating (reading and writing) |
# | U |universal newlines mode (deprecated) |
# +
def rwfile_test():
rwfile = open('rwfile', 'a+')
rwfile.seek(0, 0)
data = rwfile.read()
line_count = data.count('\n')
print('Line Number {}.'.format(line_count + 1), file=rwfile)
rwfile.close()
open('rwfile', 'w').close() # Clear file.
for i in range(5):
rwfile_test()
print(open('rwfile').read())
# -
# ## Save Objects
#
# You can even save your objects to a file. `pickle` module can do that for you.
# +
# Sample save and load binary object to a file in binary mode.
from pickle import dump, load
shopping_list = ['Milk', 'Egg', 'Bread', 'Sugar']
shopping_list_file = open('shop-list-file', 'wb')
dump(shopping_list, shopping_list_file)
shopping_list_file.close()
shopping_list_file = open('shop-list-file', 'rb')
loaded_shopping_list = load(shopping_list_file)
print(loaded_shopping_list[0])
# -
# ## Easier File Handeling
#
# For each file you should open it to use and then you have to close it. Without closing file, changes that you made to file will be ignored!
#
# It is easier to use `with` statement instead of opening and closing a file yourself.
with open('rwfile') as rwfile:
print(rwfile.read())
# ## Exercise
#
# * Write program that if a file exists, read it and print the list in the file. Then get list of names from user and add names to the file.
# * Write a function that get a filename and return dictionary that keys are words in a file and values are number of occurences of that word.
# * Write a function that get a word and a filename and return number of occurences of the word in the filename.
# +
# Phase 1: Read information and write it to user.
from pathlib import Path
from pickle import load, dump
data = Path("data")
if data.exists():
with open('data', 'rb') as data_file:
print(load(data_file))
# Phase 2: Get information from user and write
# them to file.
num_of_names = int(input('Enter number of names: '))
user_data = [input('Name: ') for i in range(num_of_names)]
with open('data', 'wb') as data_file:
dump(user_data, data_file)
# +
# 1- Open file
# 2- Read all of file
# 3- Convert all file information to a words
# 4- Count each word and create a dictionary
# 5- Return it!
# +
w = "a b c d"
print(w.split())
# +
def frequency_of_words(filename):
with open(filename) as f:
words = f.read().split()
result = {word:words.count(word) for word in set(words) if word.isalpha()}
return result
print(frequency_of_words('todo.md'))
# +
def frequency_of_word(word, fname):
return frequency_of_words(filename=fname)[word]
print(frequency_of_word('section', 'todo.md'))
# -
# # Modules
#
# When your project is getting bigger you have to organize your codes into different files and even different folders. To create new modules just create a file with anyname you want and then import it into your own code when you need it.
# +
import datetime
print(datetime.__file__)
# -
print(datetime.__name__)
# +
from math import pi
print(pi)
# -
import datetime as dt
print(dt.datetime.now())
help(datetime)
# +
from primes import prime_range
print(prime_range(10, 100))
# -
# # Exceptions
#
# Errors happen. User error or logical error are not always avoidable. Standard way of error happening is called exceptions. An exception occures when an error is occured. You have to always be ready for exceptions and try the code for any exception.
# +
# How to try for exceptions.
try:
print('always execute')
print(int('hi'))
print('end')
except TypeError as e:
print('Type Error occured.')
except ValueError as ve:
print('Value Error occured.')
print('Hello world!')
# -
# Different type of exceptions can happen. You can specify different section for each one of them.
# +
# How to trye for different exceptions.
number = input('Enter a number: ') # hi
try:
print('Square of {} is {}.'.format(number, int(number)**2))
print(some_name_that_does_not_exist)
except ValueError as ve:
print(ve)
except NameError as ne:
print(ne)
except SyntaxError as se:
print(se)
print('Running code continues.')
# -
# There are other sections:
# * `finally`: That will executed in any condition. If exception happened or didn't, if we try for it or we didn't and ... .
# * `else`: That will executed if no exception happens.
# +
# How to trye for different exceptions.
number = input('Enter a number: ') # 123
some_name_that_exist = 'No error from this part.'
try:
print('Square of {} is {}.'.format(number, int(number)**2))
print(some_name_that_exist) # Change to some_name_that_does_not_exist
except ValueError as ve:
print(ve)
else:
print('Else Section')
finally:
print('Finally Section')
print('Running code continues.')
# -
# ## Exercise
#
# * Write a program that get two numbers from a user then print result of their division. Remember tow handle divide by zero exception!
# +
num1, num2 = input('num1: '), input('num2: ')
try:
num1, num2 = int(num1), int(num2)
except ValueError as ve:
print('ERROR: You should enter numbers!')
else:
try:
print(num1/num2)
except ZeroDivisionError as zde:
print('ERROR: Second number can not be zero!')
# -
# ## Generate Error
#
# If you wanted to signal an error you can do it by using `Exception` object for now. After OOP section, we learn new and more standard way for defining our exception.
# +
def multiplication_table(row=10, column=10):
if (row <= 0 or column <= 0):
raise Exception('{} argument is {}. It should be bigger than zero!'
.format(*(['Row', row] if row <= 0 else ['Column', column])))
return [[i*j for j in range(1, column + 1)] for i in range(1, row + 1)]
# print(multiplication_table(-1, 5))
try:
print(multiplication_table(2, -3))
except Exception as e:
print(e)
# -
# You can even make this code smaller and more readable by using `assert`. It gets a condition and an exception, if condition is `False`, the exception raised.
# +
def multiplication_table(row=10, column=10):
assert row >= 0 and column >= 0, Exception('{} argument is {}. It should be bigger than zero!'
.format(*(['Row', row] if row <= 0 else ['Column', column])))
return [[i*j for j in range(1, column + 1)] for i in range(1, row + 1)]
try:
multiplication_table(-1, 5)
except Exception as e:
print(e)
# -
# # Object Oriented Programming
#
# Good for easy code designing, reusability and readability.
#
# Concepts:
# * Objects
# * State
# * Behavior
# * Class
# * Attributes
# * Methods
# * Instances
#
# Simplest python class:
# +
# A Class (code blueprints) that represent an Object (real world concepts).
class simple:
pass
# Creating an Instance of the object.
simple_var = simple()
# -
# ## Class Attributes
# +
class Car:
"""This is a simple Car class."""
number_of_wheels = 4
# It's defined withing the structures of our class.
print(Car.number_of_wheels)
# -
# Each instance can see it from the class.
simple_car = Car()
print(simple_car.number_of_wheels)
# Class attributes updates through all instances.
Car.number_of_wheels = 3
not_so_simple_car = Car()
print(not_so_simple_car.number_of_wheels)
print(simple_car.number_of_wheels)
# Class attribute can have shadows for each object instances.
# +
class Car:
number_of_wheels = 4
strange_car = Car()
# Altering class attribute through instances make specific class attribute values for them.
strange_car.number_of_wheels = 18
print(Car.number_of_wheels)
print(strange_car.number_of_wheels)
# -
# But it will break updated through class structure.
Car.number_of_wheels = 3
print(strange_car.number_of_wheels)
# Don't panic, you can bind your instance to your class again.
del strange_car.number_of_wheels
print(strange_car.number_of_wheels)
# ### Exercise
#
# * Write a Circle class
# +
class Circle:
radius = 0
a = Circle()
a.radius = 13
b = Circle()
b.radius = 1.3
# -
# ## Access to Instance
#
# Each method in a class have a default first argument that is not passed by what we code.
# +
class Person:
name = 'Nobody'
def print_name(self, a, b, c):
print(self.name)
print(a, b, c)
david = Person()
# david.name = 'David'
david.print_name(1, 2, 3)
john = Person()
john.name = 'john'
john.print_name('a', 'b', 'c')
# -
# Now that we can access instances, we can define instance attributes.
# +
class Person:
def set(self, name, age):
self.name = name
self.birth_year = 2017 - age
def get(self):
return self.name, self.birth_year
bob = Person()
bob.set(name='Bod', age=25)
print(bob.get())
# -
# ### Exercise
#
# * Write a circle class that have:
# * A class attribute. One thing that is shared between all circles.
# * An instance attribute. What makes two circle different?
# * A method to set radius.
# * A method to calculate area of circle using the class and instance attributes.
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
# +
class Circle:
PI = 3.1415926535
def set(self, radius):
self.radius = radius
def area(self):
return self.PI * self.radius ** 2
c = Circle()
c.set(5)
print("Area of a circle with redius {} is {}".format(c.radius, c.area()))
# -
# ## Special Methods
#
# These methods have specific name and specific job. They have names that starts and ends with `__`
#
# When we are creating an instance, `__init__` is the very first method that get called automatically.
# +
class Person:
def __init__(self, name, age):
self.name, self.age = name, age
def get(self):
return self.name, self.age
alice = Person('Alice', 21)
print(alice.get())
# -
# ### Operator Overloading
#
# Some of these special methods have is use by built-in operator like `len` function. By writing right special methods you can overload those behaviours.
# +
class Person:
def __init__(self, name, age):
self.name, self.age = name, age
def __len__(self):
return self.age
def __bool__(self):
if self.age > 18:
return True
else:
return False
# return True if self.age > 18 else False
# return self.age > 18
def __iter__(self):
return self.name.__iter__()
def __str__(self):
return "{} is {} year{} old.".format(self.name, self.age, 's' if self.age > 1 else '')
me = Person('Amir', 22)
print(len(me))
# -
str(me)
if me:
print('You are allowed to vote.')
else:
print('You are not allowed to vote.')
print(me)
for p in me:
print(p)
# You can find all information on special methods in [Here](http://www.diveintopython3.net/special-method-names.html).
# ## Static Methods
#
# Normal methods need an instance to do operation on it. Static methods on the other hand get called from class name. In this way you can categorize your function under a title.
# +
class String:
@staticmethod
def reverse(string):
return string[::-1]
@staticmethod
def is_upper_lower(string):
"""Checks for strings like ``, `A`, `a`, `Aa`, `aA`, `AaA`, `aAa`, ... ."""
last_upper, last_lower = False, False
for character in string:
if not last_upper and not last_lower:
if character.isupper():
last_upper = True
else:
last_lower = True
else:
if last_upper and not last_lower and character.islower():
last_upper, last_lower = False, True
elif not last_upper and last_lower and character.isupper():
last_upper, last_lower = True, False
elif (last_upper and last_lower) or (not last_upper and not last_lower):
raise Exception("There is bug in the code! last_upper: {} and last_lower: {}"
.format(last_upper, last_lower))
else:
return False
return True
# part1, part2 = string[::2], string[1::2]
# return part1.islower() and part2.isupper() or part1.isupper() and part2.islower()
@staticmethod
def unique_words(string, case_sensitive=False):
return set([s if case_sensitive else s.lower() for s in string.split()])
print(list(map(String.reverse, ['', 'a', 'ab', 'aa', 'aba', 'abc', 'aaa'])))
print(list(map(String.is_upper_lower, ['', 'a', 'A', 'aA', 'Aa', 'aAa', 'AaA', 'ab', 'AB'])))
print(String.unique_words('What you want is unique word, so you should get what you want!', True))
# -
# ## Class Methods
#
# They are special creators. In some ways they are like `__init__`. They create an instance. The first parameter of a class method is always class itself.
# +
class Celsius:
def __init__(self, temperature):
self.temperature = temperature
@classmethod
def from_fahrenheit(cls, fahrenheit):
return cls((fahrenheit - 32) * 5 / 9)
@classmethod
def from_kelvin(cls, kelvin):
return cls(kelvin - 273.15)
def __str__(self):
return "Temperature in celsius is {:0.2f}°C.".format(self.temperature)
print(Celsius(15))
print(Celsius.from_fahrenheit(98.6))
print(Celsius.from_kelvin(300))
# -
# If you have mutiple static methods and use some of in other ones, you have to hardcode the class name. This is not so good, we have so make changes as small as possible when class name is going to change. Another usage of class methods are for this problem.
# +
class String:
@classmethod
def is_palindrome(cls, string, case_insensitive=True):
string = cls._strip_string(string)
# For case insensitive comparison, we lower-case string
if case_insensitive:
string = string.lower()
return cls._is_palindrome(string)
@staticmethod
def _strip_string(string):
return ''.join(c for c in string if c.isalnum())
@staticmethod
def _is_palindrome(string):
for c in range(len(string) // 2):
if string[c] != string[-c -1]:
return False
return True
@staticmethod
def get_unique_words(string):
return set(string.split())
print(String.is_palindrome('A nut for a jar of tuna')) # True
print(String.is_palindrome('A nut for a jar of beans')) # False
# -
# ## Inheritance & Composition
#
# Main purpose of OOP is reusing existing codes. Inheritance (is-a relationship) and composition (has-a relationship) are
# two main ways to reuse code.
# +
class Engine: # Base class, Parent class
def start(self):
return 'Engine started.'
def stop(self):
return 'Engine stopped.'
class Car:
def __init__(self):
self.engine = Engine() # Composition
class MechanicalEngine(Engine): # Inheritance
def __init__(self):
self.max_speed = 120
# Above, Engine is called Parent, Base class and MechanicalEngine is called Child, Derived class.
motorcycle_engine = MechanicalEngine()
print(motorcycle_engine.start()) # MechanicalEngine class inherits attributes and methods of Engine class.
c = Car()
print(c.engine.start())
# -
# ### Overwriting Methods
#
# If you overwrite a method of parent class, the functionality of that method from parent class is gone.
# +
class Book:
def __init__(self, title, author):
self.title, self.author = title, author
def get_info(self):
return "Title: {}, Author: {}".format(self.title, self.author)
class EBook(Book):
def __init__(self, title, author, online_id):
# Not so good way! What if Book init changes? What if we didn't know Book init?
self.title, self.author, self.online_id = title, author, online_id
# Better solution BUT, what Book is not the only one we inherits? What if we
# wanted to change Book to Article?
Book.__init__(self, title, author)
self.online_id = online_id
# Best solution!
super().__init__(title, author)
self.online_id = online_id
def get_info(self):
book_info = super().get_info()
return "{}, Online ID: {}".format(book_info, self.online_id)
alice_in_wonderland = EBook('<NAME> Wonderlang', '<NAME>', 1298)
print(alice_in_wonderland.get_info())
# -
# ### Exercise
#
# * Write Person (coord & walk) and Employee classes.
# * Write Point and Line classes.
# +
class Person:
def __init__(self, name, age, height, x, y):
self.name, self.age, self.height = name, age, height
self.x, self.y = x, y
def walk(self, move_x, move_y):
self.x += move_x
self.y += move_y
def __str__(self):
return '{} at {}, {}'.format(self.name, self.x, self.y)
class Employee(Person):
def __init__(self, name, age, height, x, y, salary):
super().__init__(name, age, height, x, y)
self.salary = salary
def __str__(self):
return "{}, S: {}".format(super().__str__(), self.salary)
ahmad = Person('Ahmad', 16, 180, 4, 18)
print(ahmad)
Sara = Employee('Sara', 25, 180, 10, 5, 100)
print(Sara)
# +
class Point:
def __init__(self, x, y):
self.x, self.y = x, y
def move(self, move_x, move_y):
self.x += move_x
self.y += move_y
class Line:
def __init__(self, point_1, point_2):
self.point_1 = point_1
self.point_2 = point_2
def gradient(self):
x1, y1 = self.point_1.x, self.point_1.y
x2, y2 = self.point_2.x, self.point_2.y
if x1 - x2 == 0:
return float('inf')
return (y1 - y2) / (x1 - x2)
l = Line(Point(2, 3), Point(2, 4))
print(l.gradient())
# -
# ### Multiple Inheritance & Method Resolution Order
#
# Put names, comma seperated, and know that order of them matters (diamond problem)!
#
# As the following code shows, MRO like our class then parents of our class then grandparents of out class and so on.
# +
import math
class Shape:
tag = 'Shape'
def area():
return 0
class Rectangle(Shape):
tag = 'Rectangle'
def __init__(self, a, b):
self.a, self.b = a, b
def area(self):
return self.a * self.b
class Rhombus(Shape):
tag = 'Rhombus'
def __init__(self, a, theta):
self.a, self.theta = a, theta
def area(self):
return self.a * self.a * math.sin(self.theta)
class Square(Rectangle, Rhombus):
def __init__(self, a):
# For Rectangle
super().__init__(a, a)
# For Rhombus
# super().__init__(a, math.pi / 2)
my_square = Square(10)
print("Area of our square is: {} and Tag is: {}".format(my_square.area(), my_square.tag))
# -
# ## Private and Public
#
# Every thing in python is public! And everyone working with it without a single problem. But there is an agreement on it.
#
# If you use a single underscore at the beginning of names you use, means this is used for the inner workings of your code and it is better not to manipulate it.
#
# If you use two or more underscore at the beginning and at most one underscore at end of names you use, means this is more special name that even child classes should not manipulate it.
# +
class A:
def __init__(self):
self._private = 'Private value from A'
self.__sensitive = 'Sensitive value from A'
class B(A):
def __init__(self):
super().__init__()
self._private = 'Private value from B'
self.__sensitive = 'Sensitive value from B'
def how_to_change_private_values(self):
self.__sensitive = 'Use the name defined value with.'
a, b = A(), B()
print('A Private: {}, Sensitive: {}'.format(a._private, a._A__sensitive))
print('B Private: {}, Sensitive: {}'.format(b._private, b._B__sensitive))
print('B also include A sensitive to A works find.')
print('A sensitive value in B: {}'.format(b._A__sensitive))
# -
# ### Name Mangling
#
# When you use two or more underscores at the beginning of a name, the actual name, that is reachable from outside class scope is mangled. Mangling is the process of adding class specific name to beginning of the name you defined. For example `__VARNAME` changes to `_CLASSNAME__VARNAME`.
# ## Useful Tips
#
# * Use `isinstance()` to find a varialble origins.
isinstance(motorcycle_engine, MechanicalEngine)
# * Use `issubclass()` to find examine inheritance relation between to classes.
issubclass(MechanicalEngine, Engine), issubclass(Engine, MechanicalEngine)
# * **Property decorators** are another thing to mention. They are like _setter_ and _getter_.
# +
class Person:
def __init__(self, firstname, lastname):
self.__firstname = firstname
self.__lastname = lastname
@property
def full_name(self):
return '{} {}'.format(self.__firstname, self.__lastname)
@full_name.setter
def full_name(self, full_name):
fname_splited = full_name.split()
self.__firstname = fname_splited[0]
self.__lastname = ' '.join(fname_splited[1:])
p = Person('Nikola', 'Tesla')
print(p.full_name)
p.full_name = '<NAME>'
print(p.full_name)
# -
# # Generators
#
# * `yield`
# * `yeild from`
# # Decorators
#
# Example: Decorator to calculate up execution time.
# ## Decorator Factory
#
# Homeworks:
# * Describing what some codes do.
#
# Notes to talk:
# * Printing structures and customization.
# # PIP
#
# Python have libraries. `pip` is a program that can install and remove your packages. This a fast and safe way to get a python library for our projects.
# ## Virtualenv
#
# Consider situations like 2 projects with same dependencies of different versions. Like `pip`, `virtualenv` helps you manage your packages but in different way.
# ### Installation
#
# ```bash
# pip3 install virtualenv
# ```
# ### Usage
#
# ```bash
# virtualenv .venv # Creating python virtual environment
# source .venv/bin/activate # Activating virtual environment
# deactivate # Deactivating virtual environment
# ```
# # Database
#
# Python has many many libraries for managing databases. Here we use SQLite3 that comes with Python when you installed it.
# +
import sqlite3
import os
try:
os.remove('simple-db')
except FileNotFoundError as e:
pass
db = sqlite3.connect('simple-db')
# Create a new table.
db.execute('create table music ( title text, artist text, rating int)')
db.commit()
# Insert new data to music table.
db.execute('insert into music (title, artist, rating) values ("Immigrant Sont", "Led Zeppelin",8)')
db.execute('insert into music (title, artist, rating) values ("GOT", "<NAME>",9)')
db.execute('insert into music (title, artist, rating) values ("The map of reality", "The New Smiths",9)')
db.commit()
# Grab stored data from our database.
table = db.execute('select * from music')
for row in table:
print(row)
# -
# # Graphical User Interface with Tkinter
#
# We use `tkinter` module for this part of our course. To start we need to import the module first.
# +
# 3 ways to import tkinter module
from tkinter import * # This is very bad way. We import everything in the module to global scope.
# There is chance that we overwrite something imported from or to tkinter module.
import tkinter # This way is not best but works fine. The name is long to type.
import tkinter as tk # This is best way to do this.
# -
# If you have problem with importing `tkinter` you might want to check the installation of it. Read the error completely there should be some clue to how to fix it or search the error online for more clues.
#
# Three components for GUI programming:
# * What to place on screen? -> Widgets
# * Where to place the widgets? -> Geometry Management
# * How do widgets behave? -> Events & Callbacks
#
# Every thing is part of a big tree structure.
# ## Root Window
#
# With `tkinter` there is no need to start from scratch and invent the wheel! The `root window` is what you have to start with. A window that have basic functionality that you need.
#
# The concept of a `mainloop` is like a manager. You are the one who write standards an law, but someone else know better how to execute those.
root = tk.Tk() # root variable is filled with a basic root window object.
root.mainloop()
# ## Widgets
#
# Widgets are building blocks of a GUI application. They work together to serve an application goal. Labels, text inputs, button, sliders, image viewer, tabs and even root window are widgets.
# +
root = tk.Tk()
greetings_label = tk.Label(root, text='Hello User!') # Attaching a label to root with some text.
greetings_label.pack() # This method should be called to define a position for our new widget.
root.mainloop()
# -
# Remember all widgets are objects that is created and returned by a specific functions like `Label`. Then you configure them as you like them to behave (OOP) and then call the manager to do dirty works!
# +
root = tk.Tk()
username_frame = tk.Frame(root)
tk.Label(username_frame, text='Username').pack(side=tk.LEFT)
username_entry = tk.Entry(username_frame)
username_entry.pack(side=tk.LEFT)
username_frame.pack(fill=tk.X)
password_frame = tk.Frame(root)
tk.Label(password_frame, text='Password').pack(side=tk.LEFT)
password_entry = tk.Entry(password_frame, show='*')
password_entry.pack(side=tk.LEFT)
password_frame.pack(fill=tk.X)
def login():
username = username_entry.get()
password = password_entry.get()
if username == 'username' and password == 'password':
print('Correct username and password.')
else:
print('ERROR: Wrong username and password. username: {}, password: {}'.format(username, password))
button_frame = tk.Frame(root)
login_button = tk.Button(button_frame, text='Login', command=login)
login_button.pack(fill=tk.X)
button_frame.pack(fill=tk.X)
root.mainloop()
# -
# ## Geometry Management
#
# Three ways to manage:
# * Pack: Simple for simple layouts.
# * Grid: Most popular one. Manage a window as it is a table.
# * Place: Least popular one. Best control with absolute positioning.
# +
root = tk.Tk()
frame = tk.Frame(root)
tk.Label(frame, text='Pack demo of side and fill.').pack()
tk.Button(frame, text='A').pack(side=tk.LEFT, fill=tk.Y)
tk.Button(frame, text='B').pack(side=tk.TOP, fill=tk.X)
tk.Button(frame, text='C').pack(side=tk.RIGHT, fill=tk.NONE)
tk.Button(frame, text='D').pack(side=tk.TOP, fill=tk.BOTH)
frame.pack()
tk.Label(root, text='Pack demo of expand.').pack()
tk.Button(root, text='I do not expand').pack()
tk.Button(root, text='I do not fill X but I expand').pack(expand=True)
tk.Button(root, text='I fill X and expand.').pack(expand=True, fill=tk.X)
root.mainloop()
# -
# # Examples:
#
# * Count unique words of a file and write a json about it to another file.
# * Prime number function.
# * Prime number generator.
# * Fibonacci number recursive function.
# * Fibonacci number generator.
# * Iterator object.
# * Web crawler. (ircalendarevents)
# # Project
#
# Now learning python syntax and usages is finished. Let us start our big project!
#
# The project is simple client chat. This applicatoin have 3 main parts.
#
# * Graphical User Interface
# * Network Programming
# * Database Management
| python-for-you-and-me.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Desafio 9: Descubra quem fez o ENEM 2016 apenas para treino.
# Importando as Bibliotecas.
import pandas as pd
import numpy as np
from imblearn.over_sampling import SMOTE
from sklearn.decomposition import PCA
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import f1_score
import seaborn as sns
# Lendo os dados de treino e teste.
df_train = pd.read_csv('train.csv')
df_test = pd.read_csv('test.csv')
df_train.head()
df_test.head()
# Selecionando as features dos dados de teste e adicionando a de treineiros.
columns = list(df_test.columns)
columns.append('IN_TREINEIRO')
# Reduzindo o número de features dos dados de treino para o tamanho das do dado de teste.
df_train = df_train[columns]
df_train.columns
df_train.info()
# Selecionando apenas as features numéricas.
df_train_numeric = df_train.select_dtypes(include=['float64','int64'])
df_test_numeric = df_test.select_dtypes(include=['float64','int64'])
# # Dados Desbalanceados
# Vamos agora na feature 'IN_TREINEIRO' observar se os dados estão balanceados ou não.
df_train_numeric.IN_TREINEIRO.value_counts()
# Os dados estão desbalanceados, devemos então balanceá-los. \
# Primeiro vamos tratar os dados, contando a quantidade de valores nulos.
df_train_numeric.isnull().sum()
df_test_numeric.isnull().sum()
# Trocando os valores nulos por 0.
df_train_numeric = df_train_numeric.replace(np.NAN, 0)
df_train_numeric.isnull().sum()
# Fazendo mesma coisa para os dados de teste.
df_test_numeric = df_test_numeric.replace(np.NAN, 0)
df_test_numeric.isnull().sum()
# Contando o número de Zeros nos dados.
(df_train_numeric == 0).sum()
(df_test_numeric == 0).sum()
df_test_numeric
# Selecionando as features que farão parte dos dados de treino e teste.
x_train = df_train_numeric.drop(["IN_TREINEIRO"], axis=1)
y_train = df_train_numeric['IN_TREINEIRO']
x_test = df_test_numeric
# # Aplicando o PCA
# Contando o número de colunas
x_train.head()
# Para visualizar os dados, que estão em $\mathbb{R}^{32}$, utilizamos PCA e os transformamos para $\mathbb{R}^{2}$:
# +
pca = PCA(n_components=2)
pca.fit(x_train)
imbalanced_pca = pca.transform(x_train)
# -
# Visualizando o resultado
sns.scatterplot(imbalanced_pca[:, 0], imbalanced_pca[:, 1], hue=y_train);
# Podemos ver claramente, a maioria dos inscritos no enem como sendo não treineiros(0).
# # Smote
# Utilizaremos o SMOTE (do pacote `imblearn`), que é uma técnica de oversampling para reamostrar os dados e assim balancear o _data set.
# +
# Aplicando a técnica
smote = SMOTE(sampling_strategy="minority")
x_smote, y_smote = smote.fit_resample(x_train, y_train)
imbalanced_pca_smote = pca.transform(x_smote)
# -
# Visualizando a técnica
sns.scatterplot(x=imbalanced_pca_smote[:, 0], y=imbalanced_pca_smote[:, 1], hue=y_smote);
#Conferindo se está balanceado
y_smote.value_counts(dropna=False)
# Com os dados balanceados, vamos agora aplicar modelos de classificação e ver qual deles é mais eficiente.
# # Regressão Logística
# A Regressão Logística é bastante empregada em modelos de classificação binária.
# +
# Implementando o Modelo
logistic = LogisticRegression()
logistic.fit(x_smote, y_smote)
# -
# Prevendo o modelo obtido.
pred_1 = logistic.predict(x_test)
# Com o modelo de Regressão Linear, o score foi de 95,754923 % .
# # Random Forest
# Modelo baseado em Árvores de Decisão, com grande precisão.
# +
# Aplicando o modelo
random_forest = RandomForestClassifier(n_estimators=500)
random_forest.fit(x_smote, y_smote)
# -
# Prevendo usando o modelo
pred_2 = random_forest.predict(x_test)
# Com o modelo de Random Forest, o score foi de 99,628009 % .
# ## $k$ _Nearest Neighbors_
# O método kNN é bastante simples e muito poderoso em resultados. Ao contrário da regressão logística, que é um método paramétrico, o kNN não utiliza nenhuma suposição sobre a distribuição das classes, sendo portanto um método __não__ paramétrico. O kNN atribui a um ponto $X$ de teste a classe majoritária entre os $k$ pontos do conjunto de treinamento mais próximos a $X$. A proximidade aqui é geralmente medida em termos de distância Euclideana, mas outras métricas também são possíveis.
#
# +
# Aplicando o modelo
knn = KNeighborsClassifier(n_neighbors=7)
knn.fit(x_smote, y_smote)
# -
# Prevendo usando o modelo
pred_3 = knn.predict(x_test)
# Utilizando o knn, o score foi de 99,606127 %.
df_resposta = pd.DataFrame()
df_resposta['NU_INSCRICAO'] = df_test['NU_INSCRICAO']
df_resposta
df_resposta['IN_TREINEIRO'] = pred_3
df_resposta
df_resposta.to_csv('answer.csv', index=False, header=True)
| enem-4/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
personal_expenses=pd.read_csv(r"C:\Users\<NAME>\Desktop\data folder\my_own_data.csv")
# +
# Here i created my own expenses data, in order to show you how we can grab insights from your own expenses data.
# What you have to do is like:
# Create your own data by using excel or insert any simple data.
# Import that into your perticular IDE using pandas.
# -
personal_expenses.head(12)
personal_expenses.columns
personal_expenses.info()
# # Lineplot
plt.figure(figsize=(15,5))
plt.subplot(1,2,1)
plt.plot(personal_expenses.Month,personal_expenses.Total_Income,"r--^",markeredgecolor="blue",markersize=10)
plt.title("Total Income(2020)",fontdict={"color":"green","size":20},pad=20)
plt.xlabel("Month", fontdict={'size':20, 'color':'red'}, labelpad=20)
plt.ylabel("Income", fontdict={'size':20, 'color':'red'}, labelpad=2)
plt.ylim(0,41000)
plt.minorticks_off
plt.grid(b=True,which="major",color="yellow",axis="both")
plt.subplot(1,2,2)
plt.plot(personal_expenses.Month,personal_expenses.Total_Investment,"r--^",markeredgecolor="blue",markersize=10)
plt.title("Total Investment(2020)",fontdict={"color":"green","size":20},pad=20)
plt.xlabel("Month", fontdict={'size':20, 'color':'red'}, labelpad=20)
plt.ylabel("Investment", fontdict={'size':20, 'color':'red'}, labelpad=2)
plt.ylim(0,41000)
plt.minorticks_off
plt.grid(b=True,which="major",color="yellow",axis="both")
plt.show()
# # Insights
# * From this above plot-1, you can clearly see that in july my income is huge as compare to other month.
# * From plot-2,i can say that feb and march i did huge investment.
print(plt.style.available)
plt.style.use('classic')
plt.figure(figsize=(15,8))
plt.plot(personal_expenses.Total_Income,personal_expenses.Total_Investment,"ro",markeredgecolor="blue",markersize=10)
plt.title("Comparing investments with income",fontdict={"color":"black","size":20},pad=20)
plt.xlabel("Income", fontdict={'size':20, 'color':'red'}, labelpad=20)
plt.ylabel("Investment", fontdict={'size':20, 'color':'red'}, labelpad=2)
plt.xlim(15000,41000)
plt.ylim(15000,41000)
plt.minorticks_on()
plt.grid(b=True, which= 'minor', color ='blue' ,axis= 'both')
# # Insights
# * From this above picture,i plotted a graph which gonna show me relationship between two veriables(investment,income).
# * Here we observed that, when my income is very high,that time my investment is low.i saved lot of money.
# * some times my investment is huge,not much savings.
# * Here my analysis is that,i need not to do huge investment when my income is less.
plt.style.use('classic')
plt.figure(figsize=(15,8))
plt.plot(personal_expenses.Total_Income,personal_expenses.Total_Saving,"ro",markeredgecolor="blue",markersize= 15)
plt.title("Comparing savings with income",fontdict={"color":"black","size":20},pad=20)
plt.xlabel("Income", fontdict={'size':20, 'color':'red'}, labelpad=20)
plt.ylabel("Total Savings", fontdict={'size':20, 'color':'red'}, labelpad=2)
plt.xlim(0,41000)
plt.ylim(0,41000)
plt.minorticks_on()
plt.grid(b=True, which= 'minor', color ='blue' ,axis= 'both')
# # Insights
# * From this above picture,i plotted a graph which gonna show me relationship between two veriables(savings vs income).
# * when my income is very high,i saved almost around 15k+.
# # Pie Chart
plt.figure(figsize=(15,10))
plt.style.use('seaborn-bright')
plt.subplot(2,2,1)
plt.pie(personal_expenses.Total_Income, labels= personal_expenses.Month)
plt.title("Total Income(2020)")
plt.subplot(2,2,2)
plt.pie(personal_expenses.Total_Investment, labels= personal_expenses.Month)
plt.title("Total Investment(2020)")
plt.subplot(2,2,3)
plt.pie(personal_expenses.Total_Saving, labels= personal_expenses.Month)
plt.title("Total Savings(2020)")
plt.figure(figsize=(5,6))
plt.style.use('Solarize_Light2')
plt.axis('equal')
plt.pie(personal_expenses.Total_Investment, labels= personal_expenses.Total_Income,explode=[0,0.1,0,0,0,0,0,0,0,0,0,0],
radius= 1.5,autopct= '%2.1f%%', pctdistance = 0.8, shadow = True, labeldistance= 1.1, counterclock= True,
wedgeprops={'width': 0.5})
plt.title("Comparing investments with income",pad=40)
plt.show()
plt.figure(figsize=(5,6))
plt.style.use('Solarize_Light2')
plt.axis('equal')
plt.pie(personal_expenses.Total_Saving, labels= personal_expenses.Total_Income,explode=[0,0,0,0,0,0,0.1,0,0,0,0,0],
radius= 1.5,autopct= '%1.1f%%', pctdistance = 0.8, shadow = True, labeldistance= 1.1, counterclock= True,
wedgeprops={'width': 0.5})
plt.title("Comparing Savings with income",pad=40)
plt.show()
# # Insights
# From this above plots,
# * i can clearly say by seeing these images like, in which month what is the ratio of my income,investment and savings.
# # Box plot
plt.figure(figsize=(20,8))
plt.style.use('tableau-colorblind10')
data=[personal_expenses.Total_Income,personal_expenses.Total_Saving,personal_expenses.Total_Investment]
plt.boxplot(data,meanline=False,showmeans=True,showcaps=True,sym='r^',
whis= 1.5, vert= False,manage_ticks=True,widths=0.5,showfliers=True)
plt.xlim(1000,41000)
plt.show()
# # Insights
# From this above plots,
# * A red triangle symbols represents the outliers in my data. Outliers is nothing but things that happen once in a blue moon.its not regular basis.so for analysis we need not to consider this outliers.So this box plot helps you to know is there any presence of outliers or not.
plt.style.use('classic')
plt.figure(figsize=(10,8))
plt.scatter(personal_expenses.Month,personal_expenses.Total_Income,label = 'Income',
color="red",marker="o",s=50,linewidths=2.0)
plt.scatter(personal_expenses.Month,personal_expenses.Total_Investment,label = 'Investment',
color="green",marker="^",linewidths=2.0,s=50)
plt.scatter(personal_expenses.Month,personal_expenses.Total_Saving,label = 'Savings',
color="black",marker="P",linewidths=2.0,s=100)
plt.title("Visualize My Investment",fontdict={"color":"green","size":20},pad=20)
plt.xlabel("Month",fontdict={"color":"green","size":20},labelpad=20)
plt.ylabel("View transaction",fontdict={"color":"green","size":20},labelpad=20)
plt.ylim(0,42000)
plt.legend(loc=4,fontsize=15,framealpha=0.8)
plt.show()
# # Bar plot
plt.style.use('classic')
plt.figure(figsize=(15,7))
plt.subplot(2,2,1)
plt.bar(x=personal_expenses.Month,height=personal_expenses.Total_Income)
plt.title("My Income",fontdict={"color":"green","size":20},pad=20)
plt.xlabel("Month",fontdict={"color":"green","size":20},labelpad=20)
plt.ylabel("Income details",fontdict={"color":"green","size":20},labelpad=20)
plt.ylim(0,41000)
plt.subplot(2,2,2)
plt.bar(x=personal_expenses.Month,height=personal_expenses.Total_Investment,color="red")
plt.title("Investment",fontdict={"color":"green","size":20},pad=20)
plt.xlabel("Month",fontdict={"color":"green","size":20},labelpad=2)
plt.ylabel("Investment details",fontdict={"color":"green","size":20},labelpad=2)
plt.ylim(0,41000)
plt.subplot(2,2,3)
plt.bar(x=personal_expenses.Month,height=personal_expenses.Total_Investment,color="green")
plt.title("My Savings",fontdict={"color":"green","size":20},pad=2)
plt.xlabel("Month",fontdict={"color":"green","size":20},labelpad=2)
plt.ylabel("Savings details",fontdict={"color":"green","size":20},labelpad=2)
plt.ylim(0,41000)
# # Key-Point
# * Create your own data try to plot small small plots and do analysis.
| Data-visualization-Matplotlib(part-2) .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Neural Networks Learning
#
# Stanford CS229 - Machine Learning by Andrew Ng. Programming exercise 4 with solutions.
#
# Please check out [the repository on GitHub](https://github.com/rickwierenga/CS229-Python/). If you spot any mistakes or inconcistencies, please create an issue. For questions you can find me on Twitter: [@rickwierenga](https://twitter.com/rickwierenga). Starring the project on GitHub means a ton to me!
import numpy as np
import pandas as pd
import matplotlib.pylab as plt
from scipy.optimize import minimize
# %matplotlib inline
# ## Neural networks
#
# ---
# In the previous exercise, you implemented feedforward propagation for neural networks and used it to predict handwritten digits with the weights we provided. In this exercise, you will implement the backpropagation algorithm to learn the parameters for the neural network.
#
# Load the data and view some samples in the same way as [ex3](https://github.com/rickwierenga/CS229-Python/tree/master/ex3).
#
# Remember the output of a neural network: $h_\Theta(x) \in \mathbb{R}^K$. We want y to be a 2 dimensional vector in the form that are network should output. For example, we would represent the output 1 as:
#
# $\begin{bmatrix}0\\1\\0\\0\\0\\0\\0\\0\\0\\0\end{bmatrix}$
def remap(y, K):
m = len(y)
out = np.zeros((m, K))
for index in range(m):
out[index][y[index] - 1] = 1
return out
# +
import scipy.io as sio
# Load data
data = sio.loadmat("ex4data1.mat")
X = data["X"]
y = data["y"]
y = y.reshape(len(y))
# Initialize some useful variables
m, n = X.shape
input_layer_size = 400
hidden_layer_size = 25
K = 10 # number of classes / output_layer_size
# remap y
mapped_y = remap(y, K)
# +
import random
from PIL import Image
# get 100 random images from the dataset
num_samples = 100
samples = random.sample(list(X), num_samples)
display_img = Image.new('RGB', (200, 200))
# loop over the images, turn them into a PIL image
i = 0
for col in range(10):
for row in range(10):
array = samples[i]
array = ((array / max(array)) * 255).reshape((20, 20)).transpose() # redistribute values
img = Image.fromarray(array)
display_img.paste(img, (col*20, row*20))
i += 1
# present display_img
plt.title('Examples from the dataset')
plt.imshow(display_img, interpolation='nearest')
# -
# Load the provided weights.
# load the pretrained weights
theta = sio.loadmat("ex4weights.mat")
theta_1 = theta['Theta1']
theta_2 = theta['Theta2']
nn_params = np.concatenate([theta_1.flatten(), theta_2.flatten()])
# ### Feedforward
# These are the functions for doing feedforward as written [ex3](https://github.com/rickwierenga/CS229-Python/tree/master/ex3).
# +
def sigmoid(z):
return 1 / (1 + np.exp(-z))
def add_bias(X):
m = len(X)
bias = np.ones(m)
X = np.vstack((bias, X.T)).T
return X
def forward(theta, X):
return sigmoid(theta @ X)
# -
# ### Cost Function
#
# Remember the following variables from the lectures:
#
# * $L$: Total number of layers in the network
# * $s_l$: number of units (not counting bias unit) in layer $l$.
# * $K$: number of output classes
#
# The cost function for neural networks without regularization:
#
# $$J(\theta) = \frac{1}{m}\displaystyle\sum_{i=1}^{m}\displaystyle\sum_{k=1}^{K}\begin{bmatrix} -y^{(i)}_k \log (h_\theta(x^{(i)}) - (1 - y^{(i)})_k \log(1-(h_\theta(x^{(i)}))_k)\end{bmatrix}$$
#
# And with regularization:
#
# $$J(\theta) = -\frac{1}{m}\displaystyle\sum_{i=1}^{m}\displaystyle\sum_{k=1}^{K}\begin{bmatrix} -y^{(i)}_k \log ((h_\theta(x^{(i)}-(1-y^{(i)})_k) \log(1-(h_\theta(x^{(i)}))_k)\end{bmatrix} + \frac{\lambda}{2m} \displaystyle\sum_{l=1}^{L-1}\displaystyle\sum_{i=1}^{s_l}\displaystyle\sum_{j=1}^{s_l+1}(\Theta^{(l)}_{ji})^2$$
#
# The double sum adds up the costs for each cell in the output layer. The triple sum adds up the squares of all $\Theta$s in the network.
#
# **Exercise**: Implement the cost function for neural networks, `compute_nn_cost`, in Python. There are some structural comments to help you.
# +
def feed_forward(X, theta_1, theta_2):
""" Get predictions from the neural network, given X and Theta (in the form of theta_1, theta_2)"""
m = len(X)
activation_layer_1 = forward(theta_1, add_bias(X).T)
activation_layer_1 = add_bias(activation_layer_1.T).T
predictions = forward(theta_2, activation_layer_1)
predictions = predictions.reshape((m, len(predictions)))
return predictions
def compute_nn_cost(nn_params, X, y, input_layer_size, hidden_layer_size, K, _lambda=0):
m = len(y)
# Extract theta_1 and theta_2 from nn_params
theta_1 = nn_params[:hidden_layer_size * (input_layer_size + 1)].reshape((hidden_layer_size, (input_layer_size + 1)))
theta_2 = nn_params[(hidden_layer_size * (input_layer_size + 1)):].reshape((K, (hidden_layer_size + 1)))
# Feed forward the network to get the predictions
activation_layer_1 = add_bias(forward(theta_1, add_bias(X).T).T)
predictions = forward(theta_2, activation_layer_1.T).T
# Compute the cost of the current prediction
network_cost = (1 / m) * np.sum(-y * np.log(predictions) - (1 - y) * np.log(1 - predictions))
regularization = (_lambda / (2 * m)) * (np.sum(theta_1 ** 2) + np.sum(theta_2 ** 2))
return network_cost + regularization
# +
J = compute_nn_cost(nn_params, X, mapped_y,
input_layer_size=input_layer_size,
hidden_layer_size=hidden_layer_size,
K=10)
_lambda = 1
J_reg = compute_nn_cost(nn_params, X, mapped_y,
input_layer_size=input_layer_size,
hidden_layer_size=hidden_layer_size,
K=10,
_lambda=_lambda)
print('Cost without regularization: {:2}% (0.29 approx)'.format(J))
print('Cost with regularization: {:2}% (0.38 approx)'.format(J_reg))
# -
# ## Backpropogation
#
# ---
#
# ### The Algorithm
# As the name suggests, backpropogation is roughly the opposite of feedforward propogation. Backpropogation is an algorithm that trains neural networks by computing the gradient and thereupon applying it to the neural network.
#
# Backpropogation (Backprop) starts at the end of the network. It finds the difference between the output of the neural network and the desired output. This value gets stored in $\delta_j^{(l)}$, the error/cost for $a_j^{(l)}$. Also, $\frac{\delta}{\delta z_j^{(l)}} = \delta_j^{(l)}$. The process formaly:
#
# $$\delta_j^{(l)} = a_j^{(l)} - y_j$$ for $l = L$
#
# $$\delta_j^{(l)} = (\Theta^{(l)})^T\delta^{(l+1)} .* g'(z^{(l)}) = (\Theta^{(l)})^T\delta^{(l+1)} .* \delta^{(l)}$$ for $L > l > 1$
#
# Also:
#
# $$D^{(l)}_{ij} = \frac{\delta}{\delta \Theta_j^{(l)}}J(\Theta)$$
#
# As you would probably have expected, we don't apply the gradient to the input layer, layer 1, because we don't want to change our input in order to get a better output.
#
# The complete algorithm:
#
# Set $\Delta^{(l)}_{ij} = 0$ for all ($l$, $i$, $j$)
#
# for $i = $ to $m$
# 1. Perform forward propogation to compute $a^{(l)}$ for $l = 2, 3, ..., L$
# 2. Using $y^{(l)}$, compute $\delta^{(L)} = a^{(L)} - y^{(i)}$
# 3. Compute $\delta^{(L-2)}$, ..., $\delta^{(2)}$
# 4. $\Delta^{(l)}_{ij} := \Delta^{(l)}_{ij} + a^{(l)}_j\delta^{(l+1)}_i$
#
# $D^{(l)}_{i0} = \frac{1}{m}\Delta^{(l)}_{i0}$
#
# $D^{(l)}_{ij} = \frac{1}{m}\Delta^{(l)}_{ij} + \lambda\Theta^{(l)}_{ij}$ if y $\neq 0$
#
# ### Gradient Checking
# After you've implemented code to compute the gradient, it's often a good idea to validate your code by comparing the gradient to an approximation of it. The approximiation is defined as:
#
# $$\frac{J(\theta+\epsilon) - J(\theta-\epsilon)}{2\epsilon} \approx D$$
#
# ### Random Initialization
#
# If all values in the neural networks are the same, the neural network will fail to develop advanced patterns and it will not function. This is the reason we use a random value for theta as the initial input (break the symmetry).
#
# ### Neural Network Traning
# Follow these steps when training a neural network:
# 1. Randomly initialize weights (avoid **symmetric ... **)
# 2. Implement forward propogation to get $h_\Theta(x^{(i)})$ from any $x^{(i)}$.
# 3. Implement code to compute cost function $J(\Theta)$.
# 4. Implement backprop to compute partial derrivatives $\frac{\delta}{\delta \Theta^{(l)}_{jk}}J(\Theta)$.
# * Usually with a for loop over the training examples:
# * Perform forward and backward propogation using one example
# * Get activations $a^{(l)}$ and delta terms $d^{(l)}$ for $l = 2, ..., L$
# 5. Use gradient checking to compare $\frac{\delta}{\delta \Theta^{(l)}_{jk}}J(\Theta)$ computed using back propogation vs. using numerical estate of gradient of $J(\Theta)$. Then disable gradient checking code.
# 6. Use gradient descent or advanced optimization method with backpropogation to try to minimize $J(\Theta)$ as a function of parameters $\Theta$.
#
# ---
#
# In this part of the exercise, you will implement the backpropagation algorithm to compute the gradient for the neural network cost function. Once you have computed the gradient, you will be able to train the neural network by minimizing the cost function $J(\theta)$ using an advanced optimization algorithm such as cg.
# ### Sigmoid Gradient
# **Exercise**: Implement the gradient for the sigmoid function.
def sigmoid_gradient(z):
return sigmoid(z) * (1 - sigmoid(z))
# ### Initializing Parameters
# **Exercise**: Initialize random weights.
def initialize_random_weights(L_in, L_out):
epsilon = 0.12
W = np.random.random((L_out, L_in + 1)) * (2 * epsilon) - epsilon
return W
initial_theta_1 = initialize_random_weights(input_layer_size, hidden_layer_size)
initial_theta_2 = initialize_random_weights(hidden_layer_size, K)
initial_nn_parameters = np.concatenate([initial_theta_1.flatten(), initial_theta_2.flatten()])
# ### Implement backpropogation
# **Exercise**: Implement back propogation in Python
def backprop_gradient(nn_params, X, y, input_layer_size, hidden_layer_size, K, _lambda=None):
""" See the backpropogation cell for a detailed explanation about this algorithm """
m = len(y)
# Extract theta_1 and theta_2 from nn_params
theta_1 = nn_params[:hidden_layer_size * (input_layer_size + 1)].reshape((hidden_layer_size, (input_layer_size + 1)))
theta_2 = nn_params[(hidden_layer_size * (input_layer_size + 1)):].reshape((K, (hidden_layer_size + 1)))
# Initialize delta variables
Delta1 = np.zeros(theta_1.shape)
Delta2 = np.zeros(theta_2.shape)
# Loop over every training example
for i in range(m):
# Get predictions
Xi = np.concatenate(([1], X[i]))
activation_layer_2 = np.concatenate(([1], forward(theta_1, Xi))) # include the bias cell
predictions = forward(theta_2, activation_layer_2)
predictions = predictions.reshape((len(predictions), 1))
# Get deltas
delta3 = predictions.reshape(len(predictions)) - y[i]
delta2 = theta_2[:,1:].T @ delta3 * sigmoid_gradient(theta_1 @ Xi) # ignore the first weight because we don't adjust the bias
# Save the partial derrivatives
Delta2 += delta3.reshape((len(delta3), 1)) @ activation_layer_2.reshape((1, len(activation_layer_2)))
Delta1 += delta2.reshape((len(delta2), 1)) @ Xi.reshape((1, len(Xi)))
# get the gradients
if _lambda:
grad1 = (1/m) * Delta1 + (_lambda / m) * theta_1
grad2 = (1/m) * Delta2 + (_lambda / m) * theta_2
else:
grad1 = (1/m) * Delta1
grad2 = (1/m) * Delta2
# Unroll gradients
gradient = np.concatenate((grad1.flatten(), grad2.flatten()))
return gradient
# +
# This cell contains functions for testing the gradient. You do not have to understand them.
def debug_initialize_weights(fan_out, fan_in):
W = np.sin(np.arange(1, (fan_in + 1) * fan_out + 1)) / 10
return W.reshape(fan_out, fan_in + 1)
def compute_numerical_gradient(cost_function, nn_params, X, y, input_layer_size, hidden_layer_size, K, _lambda):
numgrad = np.zeros(nn_params.shape)
perturb = np.zeros(nn_params.shape)
e = 1e-4
for p in range(len(nn_params)):
# Set pertubation vector
perturb[p] = e
loss_1 = cost_function(nn_params-perturb, X, y,
input_layer_size=input_layer_size,
hidden_layer_size=hidden_layer_size,
K=K,
_lambda=_lambda)
loss_2 = cost_function(nn_params+perturb, X, y,
input_layer_size=input_layer_size,
hidden_layer_size=hidden_layer_size,
K=K,
_lambda=_lambda)
# Compute numerical gradient
numgrad[p] = (loss_2 - loss_1) / (2*e)
perturb[p] = 0
return numgrad
def check_gradient(cost_function, gradient_function, _lambda=0):
""" Check the gradient function """
# Initialize test values
input_layer_size = 3
hidden_layer_size = 5
K = 3
m = 5
theta_1 = debug_initialize_weights(hidden_layer_size, input_layer_size)
theta_2 = debug_initialize_weights(K, hidden_layer_size)
X = debug_initialize_weights(m, input_layer_size - 1)
y = 1 + np.mod(np.arange(1, m+1), K)
out = np.zeros((m, K))
for index in range(m):
out[index][y[index] - 1] = 1
y = out
# Unroll parameters
nn_params = np.concatenate([theta_1.flatten(), theta_2.flatten()])
# Compute gradient via backprop
backprop_gradient = gradient_function(nn_params, X, y,
input_layer_size=input_layer_size,
hidden_layer_size=hidden_layer_size,
K=K,
_lambda=_lambda)
# Compute numerical gradient
numerical_gradient = compute_numerical_gradient(cost_function, nn_params, X, y,
input_layer_size=input_layer_size,
hidden_layer_size=hidden_layer_size,
K=K,
_lambda=_lambda)
# Compare the backprop and numerical gradient
gradients = pd.DataFrame({'Backprop': backprop_gradient,
'Numerical': numerical_gradient,
'Difference':np.abs(backprop_gradient - numerical_gradient)})
pd.options.display.max_rows = 5
print(gradients)
# Compute the difference
diff = np.linalg.norm(numerical_gradient - backprop_gradient) / np.linalg.norm(backprop_gradient + numerical_gradient)
print('If the backprop gradient is computed well, the relative diffrence will be no more than 1e-9: {}'.format(diff))
# -
# Test the backpropogation algorithm (with and without regularization)
# +
print('The gradients without regularization: ')
check_gradient(compute_nn_cost, backprop_gradient)
print('\n-------------\n')
print('The gradients with regularization (lambda=3): ')
check_gradient(compute_nn_cost, backprop_gradient, _lambda=3)
# -
# ### Training the neural network
#
# The neural network will now be trained using your functions.
# +
# Get random initial values for theta
initial_theta_1 = initialize_random_weights(input_layer_size, hidden_layer_size)
initial_theta_2 = initialize_random_weights(hidden_layer_size, K)
initial_nn_parameters = np.concatenate([initial_theta_1.flatten(), initial_theta_2.flatten()])
# Set config
_lambda = 1
args = (X, mapped_y, input_layer_size, hidden_layer_size, K, _lambda)
# Train NN
result = minimize(compute_nn_cost, initial_nn_parameters, args=args,
method='CG', jac=backprop_gradient,
options={"maxiter": 50, "disp" : 1})
nn_params = result.x
theta_1 = nn_params[:hidden_layer_size * (input_layer_size + 1)].reshape((hidden_layer_size, (input_layer_size + 1)))
theta_2 = nn_params[(hidden_layer_size * (input_layer_size + 1)):].reshape((K, (hidden_layer_size + 1)))
# -
# ### Visualising the hidden layer
#
# You can now "visualize" what the neural network is learning by displaying the hidden units to see what features they are capturing in the data.
# +
# get 100 random images from the dataset
num_samples = 100
hidden_unit_visual = theta_1[:, 1:]
display_img = Image.new('RGB', (100, 100))
# loop over the images, turn them into a PIL image
i = 0
for col in range(5):
for row in range(5):
array = hidden_unit_visual[i]
array = ((array / max(array)) * 255).reshape((20, 20)).transpose() # redistribute values
img = Image.fromarray(array)
display_img.paste(img, (col*20, row*20))
i += 1
# present display_img
plt.title('Visualisation of hidden layer 1')
plt.imshow(display_img, interpolation='nearest')
# -
# ### Evaluating the model
#
# Get the accuracy on the training set for the trained values of theta. According to the exercise, you should have an accuracy of about 95%. However, this may vary due to the random initalization.
# +
# Make sure to add 1 to the result as `y` is one indexed while the prediction is 0 indexed.
layer2_activation = add_bias(forward(theta_1, add_bias(X).T).T).T
predictions = forward(theta_2, layer2_activation).T
accuracy = np.mean(np.argmax(predictions, axis = 1) + 1 == y) * 100
'Training set accuracy using the a neural network with the trained values for theta: {:2}%'.format(accuracy)
# -
| ex4/PE4 - Learning Neural Networks (Solutions).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Jericohm/daa_2021_1/blob/master/Nivel_de_Hoja_BT.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="Y33udEX2fmPv"
class NodoArbol:
def __init__(self, value, left = None, right = None):
self.data = value
self.left = left
self.right = right
# + id="-VtTAA0FfgoB"
conta = 0
mayor = [0,0]
def test(nodo):
global conta
global mayor
if nodo is None:
return None
if nodo.left is None and nodo.right is None:
if conta > mayor[1]:
mayor = [nodo.data, conta]
return None
else:
conta += 1
test(nodo.left)
test(nodo.right)
conta -= 1
return mayor
# + id="AMBJ7HNvfvln"
arbol_1 = NodoArbol(4, NodoArbol(3, NodoArbol(2, NodoArbol(2, NodoArbol(2, NodoArbol(15))), NodoArbol(7))), NodoArbol(5, NodoArbol(3, NodoArbol(9, NodoArbol(10))), NodoArbol(7, NodoArbol(8))))
# + id="PQEGAM5Mf4j3"
arbol_2 = NodoArbol("Santi",None ,NodoArbol("Jesús", NodoArbol("Pedro", None, NodoArbol("Diana"))))
# + id="nXubqR6_h9dX"
arbol_3 = NodoArbol(1, NodoArbol(2, NodoArbol(3, NodoArbol(4, NodoArbol(5)))))
# + id="eth-xts6iVDq"
arbol_4 = NodoArbol("Raíz", NodoArbol("Hoja_Izq"), NodoArbol("Hoja_Der"))
# + colab={"base_uri": "https://localhost:8080/"} id="9zJe0axRfic_" outputId="12b5420c-77bc-4eb7-8352-59af498d480f"
resultado = test(arbol_1)
print("Hoja con Mayor Nivel")
print("Dato:", resultado[0], " Nivel:", resultado[1])
# + colab={"base_uri": "https://localhost:8080/"} id="iPux3vEtg83F" outputId="3c3455c7-0f7a-4b11-fc26-8c2dd3befe2f"
resultado = test(arbol_2)
print("Hoja con Mayor Nivel")
print("Dato:", resultado[0], " Nivel:", resultado[1])
# + colab={"base_uri": "https://localhost:8080/"} id="OjxrPe_lh6Zg" outputId="22dbb432-cc7c-4048-a0cb-217d709f6566"
resultado = test(arbol_3)
print("Hoja con Mayor Nivel")
print("Dato:", resultado[0], " Nivel:", resultado[1])
# + colab={"base_uri": "https://localhost:8080/"} id="OrLN68X1iTIQ" outputId="4cf3bf49-ac24-4b75-953b-f0a83b04fca7"
resultado = test(arbol_4)
print("Hoja con Mayor Nivel")
print("Dato:", resultado[0], " Nivel:", resultado[1])
| Nivel_de_Hoja_BT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import numpy as np
# +
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense , Dropout , Activation , Flatten , Conv2D , MaxPooling2D
import pickle
X = pickle.load(open("X.pickle_test","rb"))
y = pickle.load(open("y.pickle_test","rb"))
# -
X = X/255.0
hot_vector = [0]*26
print (hot_vector)
new_y = []
for char in y:
h = []
try:
index = (ord(char)-97)
h = hot_vector.copy()
print ("h s ", h)
h[index] = 1
except:
pass
new_y.append(h)
YY = np.array(new_y)
# +
model = Sequential()
model.add( Conv2D(64 , (3,3), input_shape=X.shape[1:]) )
model.add(Activation("relu"))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add( Conv2D(64 , (3,3)) )
model.add(Activation("relu"))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Flatten())
model.add(Dense(64))
model.add(Dense(26))
model.add(Activation("softmax"))
model.compile(loss="categorical_crossentropy",
optimizer="adam",
metrics = ['accuracy'])
# model.compile(loss="hinge",
# optimizer="adadelta",
# metrics = ['accuracy'])
# loss='hinge',
# optimizer='adadelta',
model.fit(X,YY,batch_size=5 ,epochs=100, validation_split=0.22)
# -
# # Loading the Model with Weights
# +
import json
from tensorflow.keras.models import model_from_json
from tensorflow.keras.models import load_model
# load json and create model
json_file = open('model_num_.json', 'r')
loaded_model_json = json_file.read()
json_file.close()
loaded_model = model_from_json(loaded_model_json)
# load weights into new model
loaded_model.load_weights("model_num_.h5")
print("Loaded model from disk")
# model.save('model_num.hdf5')
loaded_model=load_model('model_num_.hdf5')
# -
# # Splitting Areas of img REVERSE avg_block
#
# +
import numpy as np
import cv2
import matplotlib.pyplot as plt
# %matplotlib inline
# image_name = "cts.png"
# img_loc = "res/l_"+image_name
img_loc = "test_img/jump.png"
# skel_img = cv2.imread("res/l_"+image_name+".png",0)
img = cv2.imread(img_loc,0)
res , thresh = cv2.threshold(img,127,255,cv2.THRESH_BINARY_INV)
plt.imshow(thresh)
# avg_block = [204, 280, 345, 421, 516, 648, 748, 837]
# avg_block = [204, 280, 421, 516, 648, 748, 837]
# avg_block = [116, 215, 412, 586, 677, 707, 794, 811, 862]
# -
thresh.shape[0]
avg_block.insert(0,0)
work = thresh[0:240,280:421].copy()
wrk_trans = np.transpose(work)
wrk_trans[0] = [0]*(len(wrk_trans[0]))
wrk_trans[-1] = [0]*(len(wrk_trans[-1]))
work = cv2.dilate(work,None,iterations=2)
plt.imshow(work)
im2,ctrs, hier = cv2.findContours(thresh.copy(), cv2.RETR_EXTERNAL, cv2.CHAIN_APPROX_SIMPLE)
len(ctrs)
x, y, w, h = cv2.boundingRect(ctrs[6])
roi = thresh[y:y+h, x:x+w]
plt.imshow(roi)
print(x,y,w,h)
w_img = cv2.resize(roi,(50,50))
vv = []
vv.append(w_img)
plt.imshow(w_img)
V = np.array(vv).reshape(-1,50,50,1)
clss = loaded_model.predict_classes(V)
print(clss)
chr(97+clss[0])
# +
from tensorflow.keras.models import Model
# model = ... # include here your original model
layer_name = loaded_model.layers[-2].name
intermediate_layer_model = Model(inputs=loaded_model.input,
outputs=loaded_model.get_layer(layer_name).output)
intermediate_output = intermediate_layer_model.predict(V)
# -
new_out = []
for val in intermediate_output[0]:
if(val > 0):
new_out.append(val)
else:
new_out.append(0)
div = sorted(new_out)[-1]
new_out = new_out / div
print ([val for val in new_out if val>.60])
new_out
height = thresh.shape[0]
# +
res = ""
for val in range(len(avg_block)-1):
work = thresh[0:height,avg_block[val]:avg_block[val+1]].copy()
wrk_trans = np.transpose(work)
wrk_trans[0] = [0]*(len(wrk_trans[0]))
wrk_trans[-1] = [0]*(len(wrk_trans[-1]))
work = cv2.dilate(work,None,iterations=2)
print (avg_block[val],avg_block[val+1])
# Find Contours
im2,ctrs, hier = cv2.findContours(work.copy(), cv2.RETR_EXTERNAL, cv2.CHAIN_APPROX_SIMPLE)
if(len(ctrs) == 1):
x, y, w, h = cv2.boundingRect(ctrs[0])
else:
for item in ctrs:
if(item.shape[0]>60):
x, y, w, h = cv2.boundingRect(item)
break
# region of interest
roi = work[y:y+h, x:x+w]
w_img = cv2.resize(roi,(50,50))
vv = []
vv.append(w_img)
plt.imshow(w_img)
V = np.array(vv).reshape(-1,50,50,1)
clss = loaded_model.predict_classes(V)
print(chr(97+clss[0]))
res = res + ( chr(97+clss[0]) )
plt.show()
print (res)
# -
# +
res = ""
for img in ctrs:
try:
w_img = cv2.resize(img,(50,50))
vv = []
vv.append(w_img)
plt.imshow(w_img)
V = np.array(vv).reshape(-1,50,50,1)
clss = loaded_model.predict_classes(V)
print(chr(97+clss[0]))
res = res + ( chr(97+clss[0]) )
plt.show()
except Exception as e:
print (e)
pass
print (res)
# -
from imutils.contours import sort_contours
contours, _ = sort_contours(contours, )
# +
import numpy as np
import cv2
import matplotlib.pyplot as plt
# %matplotlib inline
# image_name = "cts.png"
# img_loc = "res/l_"+image_name
img_loc = "test_img/page_4.png"
# skel_img = cv2.imread("res/l_"+image_name+".png",0)
img = cv2.imread(img_loc,0)
res , thresh = cv2.threshold(img,127,255,cv2.THRESH_BINARY_INV)
thresh = cv2.dilate(thresh,None,iterations=1)
plt.imshow(thresh)
# avg_block = [204, 280, 345, 421, 516, 648, 748, 837]
# avg_block = [204, 280, 421, 516, 648, 748, 837]
# avg_block = [116, 215, 412, 586, 677, 707, 794, 811, 862]
# -
im2,ctrs, hier = cv2.findContours(thresh.copy(), cv2.RETR_EXTERNAL, cv2.CHAIN_APPROX_SIMPLE)
len(ctrs)
contours, _ = sort_contours(ctrs, )
# +
res = ""
for contour in contours:
try:
# w_img = cv2.resize(contour,(50,50))
# vv = []
# vv.append(w_img)
plt.imshow(contour)
# V = np.array(vv).reshape(-1,50,50,1)
# clss = loaded_model.predict_classes(V)
# print(chr(97+clss[0]))
# res = res + ( chr(97+clss[0]) )
except:
pass
plt.show()
break
# -
| Notebooks/prediction during slicing .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PythonData
# language: python
# name: pythondata
# ---
# +
# Goal: Predict if an individual is currently diagnosed with Mental Health disorder based on participant answer.
# The machine learning algorithm use clean_machine_learning.csv as data entries.
# +
import pandas as pd
import warnings
warnings.filterwarnings('ignore')
import numpy as np
# Dependencies for interaction with database:
from sqlalchemy import create_engine
from sqlalchemy.orm import Session
#from config import password
# Machine Learning dependencies:
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from collections import Counter
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
from imblearn.metrics import classification_report_imbalanced
# Dependencies for visualizations:
import matplotlib.pyplot as plt
from matplotlib.pyplot import figure
# -
# Create engine and link to AWS server database:
engine = create_engine('postgresql://postgres:spring01@mht.ciic7sa0kxc0.us-west-2.rds.amazonaws.com:5432/postgres')
connect = engine.connect()
# Create session:
session = Session(engine)
# Import clean_dataset_2016 table:
clean_2016_df = pd.read_sql("SELECT * FROM survey_2016", connect)
# Check:
clean_2016_df.head()
# Import data from survey_2016 table only individual working in a tech-company:
tech_2016_df = pd.read_sql("SELECT * FROM survey_2016 WHERE tech_company = 1", connect)
# Data info:
print(tech_2016_df.shape)
print(tech_2016_df.columns.tolist())
tech_2016_df.head()
# +
# Re-Code work positions:
# Recode 'Executive Leadership':
tech_2016_df['work_position'].replace(to_replace = ['Executive Leadership|Supervisor/Team Lead|Sales','Executive Leadership','DevOps/SysAdmin|Back-end Developer',
'Executive Leadership|Supervisor/Team Lead','Supervisor/Team Lead|Executive Leadership',
'Executive Leadership|DevOps/SysAdmin|Back-end Developer',
'Executive Leadership|Supervisor/Team Lead|Dev Evangelist/Advocate|DevOps/SysAdmin|Back-end Developer|Front-end Developer',
'Executive Leadership|Supervisor/Team Lead|Back-end Developer',
'DevOps/SysAdmin|Dev Evangelist/Advocate|Supervisor/Team Lead|Executive Leadership',
'HR|Supervisor/Team Lead|Executive Leadership','Support|HR|Supervisor/Team Lead|Executive Leadership',
'Executive Leadership|Front-end Developer|Back-end Developer|Sales|Supervisor/Team Lead',
'Executive Leadership|Supervisor/Team Lead|DevOps/SysAdmin|Support|Back-end Developer|Front-end Developer',
'Designer|Front-end Developer|Back-end Developer|Supervisor/Team Lead|Executive Leadership',
'Executive Leadership|Supervisor/Team Lead|Back-end Developer|DevOps/SysAdmin',
'Executive Leadership|Supervisor/Team Lead|DevOps/SysAdmin|Back-end Developer|Front-end Developer|One-person shop',
'Executive Leadership|Supervisor/Team Lead|HR|DevOps/SysAdmin|Support|Sales|Back-end Developer|One-person shop|Designer|Front-end Developer',
'Executive Leadership|Supervisor/Team Lead|Dev Evangelist/Advocate|DevOps/SysAdmin|Support|Back-end Developer|Front-end Developer',
'Executive Leadership|DevOps/SysAdmin|Back-end Developer|Support','Executive Leadership|Dev Evangelist/Advocate',
'Other|Executive Leadership','Front-end Developer|Back-end Developer|Executive Leadership'],
value='Executive Leadership', inplace = True)
# Recode 'Supervisor/Team Lead':
tech_2016_df['work_position'].replace(to_replace = ['Supervisor/Team Lead', 'Supervisor/Team Lead|Back-end Developer','Supervisor/Team Lead|Back-end Developer|Front-end Developer',
'Other|Supervisor/Team Lead','Supervisor/Team Lead|Dev Evangelist/Advocate|DevOps/SysAdmin|Support|Back-end Developer|Designer|Front-end Developer',
'Other|Supervisor/Team Lead|Dev Evangelist/Advocate|DevOps/SysAdmin|Support|Back-end Developer','Supervisor/Team Lead|DevOps/SysAdmin|Back-end Developer',
'Supervisor/Team Lead|Designer','Supervisor/Team Lead|Sales','Supervisor/Team Lead|Front-end Developer',
'Supervisor/Team Lead|Support','Support|DevOps/SysAdmin',
'Other|Supervisor/Team Lead|Dev Evangelist/Advocate|DevOps/SysAdmin|Support|Back-end Developer|Front-end Developer|One-person shop',
'Supervisor/Team Lead|Back-end Developer|Support|DevOps/SysAdmin',
'Supervisor/Team Lead|DevOps/SysAdmin|Back-end Developer|Front-end Developer',
'Supervisor/Team Lead|Front-end Developer|Back-end Developer|Dev Evangelist/Advocate',
'Front-end Developer|Back-end Developer|Dev Evangelist/Advocate|Supervisor/Team Lead',
'Supervisor/Team Lead|DevOps/SysAdmin|Support|Front-end Developer|Designer|One-person shop',
'Supervisor/Team Lead|DevOps/SysAdmin','Supervisor/Team Lead|DevOps/SysAdmin|Support',
'Supervisor/Team Lead|DevOps/SysAdmin|Support|Back-end Developer|Front-end Developer|Designer',
'Other|Supervisor/Team Lead|Dev Evangelist/Advocate|DevOps/SysAdmin|Support|Back-end Developer|Front-end Developer|Designer|One-person shop',
'Supervisor/Team Lead|Support|Front-end Developer|Back-end Developer',
'Front-end Developer|Back-end Developer|DevOps/SysAdmin|Supervisor/Team Lead',
'Back-end Developer|Supervisor/Team Lead','DevOps/SysAdmin|Supervisor/Team Lead',
'Supervisor/Team Lead|Other','Front-end Developer|Supervisor/Team Lead',
'Other|Supervisor/Team Lead|Front-end Developer',
'Front-end Developer|Back-end Developer|Supervisor/Team Lead',
'Back-end Developer|DevOps/SysAdmin|Supervisor/Team Lead','Supervisor/Team Lead|Support|Back-end Developer',
'Other|Supervisor/Team Lead|Back-end Developer|Front-end Developer',
'Supervisor/Team Lead|DevOps/SysAdmin|HR','Designer|Support|Supervisor/Team Lead',
'Supervisor/Team Lead|Back-end Developer|DevOps/SysAdmin',
'Back-end Developer|Dev Evangelist/Advocate|Supervisor/Team Lead',
'Supervisor/Team Lead|Front-end Developer|Back-end Developer',
'Supervisor/Team Lead|Front-end Developer|Back-end Developer|DevOps/SysAdmin',
'Back-end Developer|DevOps/SysAdmin|Dev Evangelist/Advocate|Supervisor/Team Lead',
'Supervisor/Team Lead|Back-end Developer|Front-end Developer|Designer',
'Supervisor/Team Lead|Front-end Developer|Back-end Developer|DevOps/SysAdmin|Dev Evangelist/Advocate',
'Supervisor/Team Lead|Dev Evangelist/Advocate|Back-end Developer|Front-end Developer',
'Supervisor/Team Lead|Designer|Front-end Developer|Support',
'Other|Supervisor/Team Lead|DevOps/SysAdmin|Support|Front-end Developer|Back-end Developer',
'Supervisor/Team Lead|DevOps/SysAdmin|Back-end Developer|Designer',
'Supervisor/Team Lead|DevOps/SysAdmin|Support|Back-end Developer|Front-end Developer',
'Supervisor/Team Lead|Dev Evangelist/Advocate|Back-end Developer',
'Other|Supervisor/Team Lead|DevOps/SysAdmin|Back-end Developer|Support',
'Other|Supervisor/Team Lead|Support|Back-end Developer|Designer',
'Supervisor/Team Lead|DevOps/SysAdmin|Back-end Developer|Designer|Front-end Developer',
'Supervisor/Team Lead|Support|Back-end Developer|Front-end Developer',
'Supervisor/Team Lead|Back-end Developer|One-person shop',
'Supervisor/Team Lead|DevOps/SysAdmin|Support|Back-end Developer',
'Supervisor/Team Lead|Front-end Developer|Back-end Developer|Support|DevOps/SysAdmin',
'Designer|Front-end Developer|Back-end Developer|Supervisor/Team Lead',
'Supervisor/Team Lead|Front-end Developer|Designer',
'Supervisor/Team Lead|Back-end Developer|DevOps/SysAdmin|Dev Evangelist/Advocate',
'Supervisor/Team Lead|Back-end Developer|Designer',
'Supervisor/Team Lead|DevOps/SysAdmin|Front-end Developer|Back-end Developer',
'Other|Back-end Developer|Supervisor/Team Lead'],
value='Supervisor', inplace = True)
# Recode 'Supervisor/Team Lead':
tech_2016_df['work_position'].replace(to_replace = ['DevOps/SysAdmin|Back-end Developer|Front-end Developer','DevOps/SysAdmin|Back-end Developer',
'DevOps/SysAdmin|Designer','Back-end Developer|DevOps/SysAdmin',
'DevOps/SysAdmin|Support|Back-end Developer|Front-end Developer',
'DevOps/SysAdmin|Back-end Developer|Front-end Developer|Designer',
'DevOps/SysAdmin|Support','DevOps/SysAdmin|One-person shop',
'DevOps/SysAdmin|Designer|Front-end Developer|Back-end Developer',
'Front-end Developer|Back-end Developer|DevOps/SysAdmin|Dev Evangelist/Advocate',
'DevOps/SysAdmin|Front-end Developer|Back-end Developer',
'DevOps/SysAdmin|Support|Back-end Developer|Front-end Developer|Designer',
'Back-end Developer|Support|DevOps/SysAdmin',
'Front-end Developer|Back-end Developer|DevOps/SysAdmin',
'DevOps/SysAdmin|Support|Back-end Developer',
'Dev Evangelist/Advocate|DevOps/SysAdmin|Back-end Developer',
'DevOps/SysAdmin|Front-end Developer|Back-end Developer|Support',
'DevOps/SysAdmin|Support|One-person shop',
'Dev Evangelist/Advocate|DevOps/SysAdmin|Support|Back-end Developer|Front-end Developer|One-person shop',
'Other|DevOps/SysAdmin|Back-end Developer',
'Dev Evangelist/Advocate|DevOps/SysAdmin',
'Designer|Front-end Developer|Back-end Developer|DevOps/SysAdmin|Other',
'Other|DevOps/SysAdmin|Support|Back-end Developer',
'Front-end Developer|Back-end Developer|Support|DevOps/SysAdmin',
'DevOps/SysAdmin|Support|Back-end Developer|One-person shop|Front-end Developer',
'Designer|Front-end Developer|Back-end Developer|DevOps/SysAdmin',
'DevOps/SysAdmin|Support|Front-end Developer|Back-end Developer',
'Dev Evangelist/Advocate|Front-end Developer|Back-end Developer|DevOps/SysAdmin',
'Dev Evangelist/Advocate|DevOps/SysAdmin|Designer|Front-end Developer|Back-end Developer'],
value='DevOps/SysAdmin', inplace = True)
# Recode developer roles:
tech_2016_df['work_position'].replace(to_replace = ['Back-end Developer', 'Front-end Developer','Back-end Developer|Front-end Developer',
'Front-end Developer|Back-end Developer','Dev Evangelist/Advocate',
'Front-end Developer|Designer','Designer',
'Back-end Developer|Dev Evangelist/Advocate','Support|Front-end Developer|Back-end Developer',
'Back-end Developer|One-person shop','Support|Designer','Support|Back-end Developer',
'Designer|Front-end Developer','Front-end Developer|Back-end Developer|Other',
'Other|Back-end Developer','Other|Front-end Developer',
'Support|Front-end Developer|Designer','Dev Evangelist/Advocate|Back-end Developer',
'Back-end Developer|Front-end Developer|Designer','Support|Sales|Designer',
'Support|Back-end Developer|Front-end Developer',
'Dev Evangelist/Advocate|Back-end Developer|Front-end Developer',
'Front-end Developer|Back-end Developer|Support',
'Other|Back-end Developer|Front-end Developer|Designer',
'Other|Front-end Developer|Designer|One-person shop',
'Front-end Developer|Back-end Developer|Dev Evangelist/Advocate',
'Dev Evangelist/Advocate|Support|Back-end Developer',
'Support|Back-end Developer|Front-end Developer|Designer',
'Dev Evangelist/Advocate|Back-end Developer|Support',
'Front-end Developer|Back-end Developer|Support|Dev Evangelist/Advocate',
'Other|Dev Evangelist/Advocate|Back-end Developer|Front-end Developer',
'Other|Support|Back-end Developer|Front-end Developer|Designer',
'Other|Dev Evangelist/Advocate|Sales|Back-end Developer|Front-end Developer',
'Other|Front-end Developer|Designer',
'Dev Evangelist/Advocate|Support|Back-end Developer|Front-end Developer',
'Dev Evangelist/Advocate|Back-end Developer|Designer|Front-end Developer',
'One-person shop|Front-end Developer|Back-end Developer|Dev Evangelist/Advocate|Other'],
value='Developer/Designer', inplace = True)
# Recode for Other (including NaN values):
tech_2016_df['work_position'].replace(to_replace = ['Support', 'Other', 'Other|Support', 'One-person shop', 'Support|Other','HR', 'Sales',
'Support|Back-end Developer|One-person shop','HR|Dev Evangelist/Advocate|Sales',
'Other|HR','Dev Evangelist/Advocate|Support','Other|Dev Evangelist/Advocate|Support',
'Other|Dev Evangelist/Advocate',np.NaN],
value='Other', inplace = True)
# -
# Check work_position:
tech_2016_df["work_position"].value_counts()
position = tech_2016_df["work_position"].value_counts()
edgecolor = ["black"]
colors = ["darkgreen","green","lime", "lightgreen", "yellow"]
position. plot.bar(title = "Current Mental Health Disorder:",
xlabel = "Position",
ylabel = "Numbers of answers",
color = colors,
edgecolor = edgecolor)
plt.xticks(rotation = 45)
plt.show()
# Check company_size:
tech_2016_df["company_size"].value_counts()
# +
# Replace values:
tech_2016_df["company_size"].replace(["26-100", "100-500", "25-Jun", "More than 1000", "500-1000", "5-Jan"],
["small","medium","small medium", "large","medium large", "start up"], inplace = True)
tech_2016_df["company_size"].replace(["start up", "small medium", "medium large"],["small", "medium", "large"], inplace = True)
size = tech_2016_df["company_size"].value_counts()
labels = ["Small", "Medium", "Large"]
colors = ["yellow", "orange", "red"]
size.plot.pie(title = "Numbers of company size:",
labels = labels,
colors = colors,
autopct='%1.1f%%')
plt.show()
# -
# Check mh_medical_leave:
tech_2016_df["mh_medical_leave"].value_counts()
# Replace values:
tech_2016_df["mh_medical_leave"].replace(["Very easy", "Neither easy nor difficult","Very difficult"], ["easy", "medium", "difficult"], inplace = True)
tech_2016_df["mh_medical_leave"].replace(["Somewhat easy", "I don't know", "Somewhat difficult"],["easy", "medium", "difficult"], inplace = True)
# Check:
tech_2016_df["mh_hurt_on_career"].value_counts()
# +
# Replace values:
tech_2016_df["mh_hurt_on_career"].replace(["Yes, it has", "No, it has not", "Maybe"], ["yes", "no", "maybe"], inplace = True)
tech_2016_df["mh_hurt_on_career"].replace(["Yes, I think it would", "No, I don't think it would"], ["yes", "no"], inplace = True)
hurt_career = tech_2016_df["mh_hurt_on_career"].value_counts()
edgecolor = ["black"]
colors = ["blue", "yellow", "red"]
hurt_career.plot.bar(title = "Mental Health can hurt employee career:",
xlabel = "Answer",
ylabel = "Numbers of answers",
color = colors,
edgecolor = edgecolor)
plt.xticks(rotation = 360)
plt.show()
# -
# Check mh_dx_current:
tech_2016_df["mh_dx_current"].value_counts()
# +
# Replace values:
tech_2016_df["mh_dx_current"].replace(["Maybe"], ["No"], inplace = True)
current = tech_2016_df["mh_dx_current"].value_counts()
edgecolor = ["black"]
colors = ["blue", "red"]
current.plot.bar(title = "Current Mental Health Disorder:",
xlabel = "Answer",
ylabel = "Numbers of answers",
color = colors,
edgecolor = edgecolor)
plt.xticks(rotation = 360)
plt.show()
#plt.savefig("../images/Current_Mental_Health_Disorder.png")
# -
# Check mh_dx_pro:
tech_2016_df["mh_dx_pro"].value_counts()
# Check gender:
tech_2016_df["gender"].value_counts()
# +
# Replace values:
tech_2016_df["gender"].replace(["nonbinary"], ["female"], inplace = True)
gender = tech_2016_df["gender"].value_counts()
labels = ["Male", "Female"]
colors = ["blue", "red"]
gender.plot.pie(title = "Individual Gender:",
labels = labels,
colors = colors,
autopct='%1.1f%%')
plt.show()
# -
# Check gender:
tech_2016_df["gender"].value_counts()
# Check mh_family_history:
tech_2016_df["mh_family_history"].value_counts()
# Replace values:
tech_2016_df["mh_family_history"].replace(["I don't know"], ["No"], inplace = True)
# Check mh_sought_pro_tx:
tech_2016_df["mh_sought_pro_tx"].value_counts()
# +
# Replace values:
tech_2016_df["mh_sought_pro_tx"].replace([1, 2], ["yes", "no"], inplace = True)
sought_pro = tech_2016_df["mh_sought_pro_tx"].value_counts()
labels = ["Yes", "No"]
colors = ["blue", "red"]
sought_pro.plot.pie(title = "Mental Health Diagnostic in the Past",
labels = labels,
colors = colors,
autopct='%1.1f%%')
plt.show()
# -
# Check mh_dx_past:
tech_2016_df["mh_dx_past"].value_counts()
# +
# Replace values:
tech_2016_df["mh_dx_past"].replace(["Maybe"], ["no"], inplace = True)
tech_2016_df["mh_dx_past"].replace(["No"], ["no"], inplace = True)
dx_past = tech_2016_df["mh_dx_past"].value_counts()
labels = ["Yes", "No"]
colors = ["blue", "red"]
dx_past.plot.pie(title = "Mental Health Diagnostic in the Past",
labels = labels,
colors = colors,
autopct='%1.1f%%')
plt.show()
# -
# Check mh_coverage:
tech_2016_df["mh_coverage"].value_counts()
# +
# Replace values:
tech_2016_df["mh_coverage"].replace(["Not eligible for coverage / N/A"], ["N/A"], inplace = True)
tech_2016_df["mh_coverage"].replace(["I don't know"], ["N/A"], inplace = True)
mh_coverage = tech_2016_df["mh_coverage"].value_counts()
edgecolor = ["black"]
colors = ["blue", "orange", "red"]
mh_coverage.plot.bar(title = "Employer provide mental health coverage:",
xlabel = "Answer",
ylabel = "Numbers of answers",
color = colors,
edgecolor = edgecolor)
plt.xticks(rotation = 360)
plt.show()
# -
# Check prev_mh_benefits:
tech_2016_df["prev_mh_benefits"].value_counts()
# +
# Replace values:
tech_2016_df["prev_mh_benefits"].replace(["Some did"], ["yes"], inplace = True)
tech_2016_df["prev_mh_benefits"].replace(["I don't know"], ["no"], inplace = True)
tech_2016_df["prev_mh_benefits"].replace(["Yes, they all did"], ["yes"], inplace = True)
tech_2016_df["prev_mh_benefits"].replace(["No, none did"], ["no"], inplace = True)
tech_2016_df["prev_mh_benefits"].value_counts()
prev_benefits = tech_2016_df["prev_mh_benefits"].value_counts()
edgecolor = ["black"]
colors = ["blue", "red"]
prev_benefits.plot.bar(title = " Previsous employer provide mental health coverage:",
xlabel = "Answer",
ylabel = "Numbers of answers",
color = colors,
edgecolor = edgecolor)
plt.xticks(rotation = 360)
plt.show()
# -
# Check prev_employers:
tech_2016_df["prev_employers"].value_counts()
# +
tech_2016_df["prev_employers"].replace([1, 0], ["yes", "no"], inplace = True)
tech_2016_df["prev_employers"].value_counts()
prev = tech_2016_df["prev_employers"].value_counts()
labels = ["Yes", "No"]
colors = ["blue", "red"]
prev.plot.pie(title = "Individual who had a previous employer:",
labels = labels,
colors = colors,
autopct='%1.1f%%')
plt.show()
# +
# Check country_live:
tech_2016_df["country_live"].value_counts()
country = tech_2016_df["country_live"].value_counts()
country.plot.pie(title = "Country distribution of survey's responders:", autopct='%1.1f%%')
plt.xticks(rotation = 45)
plt.show()
# +
# Check mh_discussion_coworkers:
print(tech_2016_df["mh_discussion_coworkers"].value_counts())
mh_discussion_coworkers = tech_2016_df["mh_discussion_coworkers"].value_counts()
edgecolor = ["black"]
labels = ["Maybe", "No", "Yes"]
colors = ["blue", "yellow", "red"]
mh_discussion_coworkers.plot.bar(title = " Mental Health discussion with a coworkers:",
xlabel = "Answer",
ylabel = "Numbers of answers",
color = colors,
edgecolor = edgecolor)
plt.xticks(rotation = 360)
plt.show()
# +
# Check mh_discussion_supervisors:
tech_2016_df["mh_discussion_supervisors"].value_counts()
mh_discussion_coworkers = tech_2016_df["mh_discussion_supervisors"].value_counts()
edgecolor = ["black"]
labels = ["Maybe", "No", "Yes"]
colors = ["blue", "yellow", "red"]
mh_discussion_coworkers.plot.bar(title = " Mental Health discussion with a supervisors:",
xlabel = "Answer",
ylabel = "Numbers of answers",
color = colors,
edgecolor = edgecolor)
plt.xticks(rotation = 360)
plt.show()
# +
# Check remote:
tech_2016_df["remote"].value_counts()
size = tech_2016_df["remote"].value_counts()
labels = ["Sometimes", "Always", "Never"]
colors = ["blue", "teal", "green"]
size.plot.pie(title = "Working remotely:",
labels = labels,
colors = colors,
autopct='%1.1f%%')
plt.show()
# -
# Check:
print("-------------Values Counts----------------")
print("mh_discussion_supervisors")
print("------------------------------------------")
print(tech_2016_df["work_position"].value_counts())
print("------------------------------------------")
print("mh_discussion_supervisors")
print("------------------------------------------")
print(tech_2016_df["mh_discussion_supervisors"].value_counts())
print("------------------------------------------")
print("mh_discussion_coworkers")
print("------------------------------------------")
print(tech_2016_df["mh_discussion_coworkers"].value_counts())
print("------------------------------------------")
print("mh_dx_pro")
print("------------------------------------------")
print(tech_2016_df["mh_dx_pro"].value_counts())
print("------------------------------------------")
print("mh_hurt_on_career")
print("------------------------------------------")
print(tech_2016_df["mh_hurt_on_career"].value_counts())
print("------------------------------------------")
print("mh_medical_leave")
print("------------------------------------------")
print(tech_2016_df["mh_medical_leave"].value_counts())
print("------------------------------------------")
print("gender")
print("------------------------------------------")
print(tech_2016_df["gender"].value_counts())
print("------------------------------------------")
print("mh_dx_current")
print("------------------------------------------")
print(tech_2016_df["mh_dx_current"].value_counts())
print("------------------------------------------")
print("mh_family_history")
print("------------------------------------------")
print(tech_2016_df["mh_family_history"].value_counts())
print("------------------------------------------")
print("mh_sought_pro_tx")
print("------------------------------------------")
print(tech_2016_df["mh_sought_pro_tx"].value_counts())
print("------------------------------------------")
print("mh_dx_past")
print("------------------------------------------")
print(tech_2016_df["mh_dx_past"].value_counts())
print("------------------------------------------")
print("mh_coverage")
print("------------------------------------------")
print(tech_2016_df["mh_coverage"].value_counts())
print("------------------------------------------")
print("prev_mh_benefits")
print("------------------------------------------")
print(tech_2016_df["prev_mh_benefits"].value_counts())
print("------------------------------------------")
print("prev_employers")
print("------------------------------------------")
print(tech_2016_df["prev_employers"].value_counts())
print("------------------------------------------")
print("country_live")
print("------------------------------------------")
print(tech_2016_df["country_live"].value_counts())
print("------------------------------------------")
print("mh_discussion_coworkers")
print("------------------------------------------")
print(tech_2016_df["mh_discussion_coworkers"].value_counts())
print("------------------------------------------")
print("mh_discussion_supervisors")
print("------------------------------------------")
print(tech_2016_df["mh_discussion_supervisors"].value_counts())
print("------------------------------------------")
print("remote")
print("------------------------------------------")
print(tech_2016_df["remote"].value_counts())
print("------------------------------------------")
tech_2016_df.to_csv("../resources/clean_machine_learning.csv")
# Log-in database:
#table_name = "clean_machine_learning"
#tech_2016_df.to_sql(
#table_name,
#engine,
#if_exists = "replace")
# Test:
clean_2016_df = pd.read_sql("SELECT * FROM clean_machine_learning", connect)
clean_2016_df.head()
# Check:
#print(tech_2016_df["yes_condition_dx"].value_counts())
print(tech_2016_df["yes_condition_dx"].isnull().count())
print(tech_2016_df["yes_condition_dx"].count())
#print(tech_2016_df["yes_condition_dx"].unique)
# Insight on yes_condition_dx columns:
condition_dx_df = pd.read_sql("SELECT COUNT(new_id), yes_condition_dx FROM clean_machine_learning GROUP BY yes_condition_dx ORDER BY COUNT(new_id) DESC", connect)
print(condition_dx_df.head(20))
condition_dx_df.plot()
plt.show()
condition_dx_df.head()
condition_df = pd.read_sql("Select yes_condition_dx FROM clean_machine_learning", connect)
condition_df.head()
# +
# Cleaning yes_condition_dx column:
condition_list = condition_df["yes_condition_dx"].tolist()
condition_list
disorder = []
no_answer = []
for condition in condition_list:
if condition == 'Anxiety Disorder (Generalized, Social, Phobia, etc)':
disorder.append(condition)
if condition == 'Anxiety Disorder (Generalized, Social, Phobia, etc)|Mood Disorder (Depression, Bipolar Disorder, etc)':
disorder.append(condition)
if condition == 'Anxiety Disorder (Generalized, Social, Phobia, etc)|Substance Use Disorder':
disorder.append(condition)
if condition == 'Mood Disorder (Depression, Bipolar Disorder, etc)':
disorder.append(condition)
if condition == 'Anxiety Disorder (Generalized, Social, Phobia, etc)|Obsessive-Compulsive Disorder':
disorder.append(condition)
if condition == 'Mood Disorder (Depression, Bipolar Disorder, etc)|Attention Deficit Hyperactivity Disorder|Post-traumatic Stress Disorder':
disorder.append(condition)
if condition == 'Anxiety Disorder (Generalized, Social, Phobia, etc)|Attention Deficit Hyperactivity Disorder':
disorder.append(condition)
if condition == 'Mood Disorder (Depression, Bipolar Disorder, etc)|Eating Disorder (Anorexia, Bulimia, etc)|Obsessive-Compulsive Disorder':
disorder.append(condition)
if condition == 'Seasonal Affective Disorder':
disorder.append(condition)
if condition == 'Post-traumatic Stress Disorder|Dissociative Disorder':
disorder.append(condition)
if condition == 'None':
no_answer.append(condition)
# Check unique value:
x = np.array(disorder)
np.unique(x)
# +
# Select the desire columns for machine learning model:
tech_2016_df_current_dx_mh = tech_2016_df[["work_position", "company_size", "mh_medical_leave", "country_work", "mh_family_history",
"mh_dx_pro", "mh_coverage", "mh_dx_current", "gender", "mh_dx_past", "prev_mh_benefits"
,"country_live"
]]
# Check:
print(tech_2016_df_current_dx_mh.shape)
tech_2016_df_current_dx_mh.head()
# +
# Encode dataset:
# Create label encoder instance:
le = LabelEncoder()
# Make a copy of desire data:
encoded_df_dx_mh = tech_2016_df_current_dx_mh.copy()
# Encode all desired columns:
features = encoded_df_dx_mh.columns.tolist()
for feature in features:
encoded_df_dx_mh[feature] = le.fit_transform(encoded_df_dx_mh[feature])
# Check:
print(encoded_df_dx_mh.shape)
encoded_df_dx_mh.head()
# +
# Clean features / add more features / oversample to boost accuracy / change train-test size /
# -
# ### Do you currently have a mental health disorder?
# +
# Create our target:
y = encoded_df_dx_mh["mh_dx_current"]
# Create our features:
X = encoded_df_dx_mh.drop(columns = "mh_dx_current", axis =1)
# -
# Split the data:
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=5, stratify=y, test_size = 0.25)
# ### Balanced Random Forest Classsifier
# +
from sklearn.ensemble import RandomForestClassifier
# Create a random forest classifier:
rf_model = RandomForestClassifier(n_estimators=100, random_state=1)
# Fitting the model:
rf_model = rf_model.fit(X_train, y_train)
# Making predictions using the testing data:
predictions = rf_model.predict(X_test)
# -
# Predict outcomes for test data set:
predictions = rf_model.predict(X_test)
predictions_df = pd.DataFrame({"Actual": y_test, "Prediction": predictions})
predictions_df.head()
# +
import matplotlib.pyplot as plt
from matplotlib.pyplot import figure
import numpy as np
# Plot predictions data frame:
x1 = y_test
y1 = np.arange(0,192,1)
x2 = predictions
y2 = np.arange(0,192,1)
# Plot actual and predictions:
figure(figsize=(10, 6), dpi=80)
plt.plot(x1, y1, label = "Actual", color = "blue")
plt.plot(x2, y2, label = "Predictions", color = "red")
plt.title("Actual and Predictions for current mental health diagnostic target:")
plt.xlabel("MH diagnostic value")
plt.ylabel("Index")
plt.show()
#plt.savefig("../images/random_forrest_current_dx_predictions.png")
# -
y_pred = ([0, 0, 1, 1, 0, 0, 1, 1, 0, 1, 0, 1, 0, 0, 1, 1, 1, 0, 1, 0, 1, 0,
1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 1, 0, 0, 1, 0, 1, 1, 0, 0, 0, 1,
0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 1, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0,
0, 0, 1, 0, 0, 1, 1, 0, 1, 1, 0, 0, 1, 1, 0, 0, 0, 1, 0, 1, 1, 1,
0, 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 1, 1, 0, 0, 1, 0, 1, 1, 1, 0, 0,
1, 0, 1, 1, 1, 0, 1, 1, 0, 1, 1, 1, 0, 0, 0, 0, 1, 1, 1, 0, 0, 1,
0, 1, 0, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 1, 0, 1, 1, 0, 1, 0, 0, 0,
1, 0, 1, 1, 1, 0, 0, 1, 1, 1, 1, 1, 0, 1, 0, 0, 0, 1, 1, 0, 0, 1,
1, 1, 0, 0, 1, 0, 1, 0, 0, 1, 0, 1, 1, 0, 0, 0])
# +
confusion_matrix(y_test, predictions)
# Create a DataFrame from the confusion matrix:
matrix = confusion_matrix(y_test, y_pred)
results = pd.DataFrame(matrix, index = ["Dx_positif", "Dx_negatif"], columns = ["Predicted True", "Predicted False"])
results
# -
x = results["Predicted True"].value_counts()
y = results["Predicted False"].value_counts()
results.plot()
plt.xlabel("Diagnosis")
plt.ylabel("Numbers of Diagnosis")
plt.show()
# +
# Calculated the balanced accuracy score
from sklearn.metrics import accuracy_score
y_pred = predictions
# Print the imbalanced classification report:
from imblearn.metrics import classification_report_imbalanced
print(f"Accuracy score: {accuracy_score(y_test, y_pred)}")
print("------------------------------------------------------------------------------------")
print(f"Classification report: Balanced Random Forest Classifier ")
print("------------------------------------------------------------------------------------")
print(classification_report_imbalanced(y_test, y_pred))
print("------------------------------------------------------------------------------------")
# -
# ### Naive Random Oversampling:
# +
# Resample the training data with the RandomOversampler
from imblearn.over_sampling import RandomOverSampler
from collections import Counter
ros = RandomOverSampler(random_state=1)
X_resampled, y_resampled = ros.fit_resample(X_train, y_train)
Counter(y_resampled)
# +
# Train the Logistic Regression model using the resampled data:
from sklearn.linear_model import LogisticRegression
model = LogisticRegression(solver='lbfgs', random_state=1)
model.fit(X_resampled, y_resampled)
# Predict outcomes for test data set
predictions = model.predict(X_test)
predictions_df = pd.DataFrame({"Actual": y_test, "Prediction": predictions,})
predictions_df.head()
# +
import matplotlib.pyplot as plt
from matplotlib.pyplot import figure
import numpy as np
# Plot predictions data frame:
x1 = y_test
y1 = np.arange(0,192,1)
x2 = predictions
y2 = np.arange(0,192,1)
# Plot actual and predictions:
figure(figsize=(10, 6), dpi=80)
plt.plot(x1, y1, label = "Actual", color = "blue")
plt.plot(x2, y2, label = "Predictions", color = "red")
plt.title("Actual and Predictions for current mental health diagnostic target:")
plt.xlabel("MH diagnostic value")
plt.ylabel("Index")
plt.show()
# +
confusion_matrix(y_test, predictions)
# Create a DataFrame from the confusion matrix:
matrix = confusion_matrix(y_test, y_pred)
results = pd.DataFrame(matrix, index = ["Dx_positif", "Dx_negatif"], columns = ["Predicted True", "Predicted False"])
results
# -
x = results["Predicted True"].value_counts()
y = results["Predicted False"].value_counts()
results.plot()
plt.xlabel("Diagnosis")
plt.ylabel("Numbers of Diagnosis")
plt.show()
# +
# Calculated the balanced accuracy score:
y_pred = model.predict(X_test)
# Print the imbalanced classification report:
print(f"Accuracy score: {accuracy_score(y_test, y_pred)}")
print("------------------------------------------------------------------------------------")
print(f"Classification report: Naive Random Oversampling: ")
print("------------------------------------------------------------------------------------")
print(classification_report_imbalanced(y_test, y_pred))
print("------------------------------------------------------------------------------------")
# -
# ### Model improvement: Add more features into the algorithm.
# +
# Select the desire columns for machine learning model:
tech_2016_df_current_dx_mh2 = tech_2016_df[["work_position", "company_size", "mh_medical_leave", "country_work", "mh_family_history",
"mh_dx_pro", "mh_coverage", "mh_dx_current", "gender", "mh_dx_past", "prev_mh_benefits"
,"country_live", "mh_discussion_coworkers", "mh_discussion_supervisors",
"yes_condition_dx", "remote", "prev_mh_discussion_coworkers", "prev_mh_discussion_supervisors",
"prev_mh_importance_employer", "mh_hurt_on_career"
]]
# Check:
print(tech_2016_df_current_dx_mh2.shape)
tech_2016_df_current_dx_mh2.head()
# +
# Create our target:
y = encoded_df_dx_mh["mh_dx_current"]
# Create our features:
X = encoded_df_dx_mh.drop(columns = "mh_dx_current", axis =1)
# +
# Encode dataset:
# Create label encoder instance:
le = LabelEncoder()
# Make a copy of desire data:
encoded_df_dx_mh = tech_2016_df_current_dx_mh2.copy()
# Encode all desired columns:
features = encoded_df_dx_mh.columns.tolist()
for feature in features:
encoded_df_dx_mh[feature] = le.fit_transform(encoded_df_dx_mh[feature])
# Check:
print(encoded_df_dx_mh.shape)
encoded_df_dx_mh.head()
# Split the data:
X_train1, X_test1, y_train1, y_test1 = train_test_split(X, y, random_state=5, stratify=y, test_size = 0.25)
from sklearn.ensemble import RandomForestClassifier
# Create a random forest classifier:
rf_model = RandomForestClassifier(n_estimators=100, random_state=1)
# Fitting the model:
rf_model = rf_model.fit(X_train1, y_train1)
# Making predictions using the testing data:
predictions = rf_model.predict(X_test1)
# -
# Predict outcomes for test data set:
predictions1 = rf_model.predict(X_test1)
predictions_df = pd.DataFrame({"Actual": y_test1, "Prediction": predictions1})
predictions_df.head()
# +
confusion_matrix(y_test1, predictions)
# Create a DataFrame from the confusion matrix:
matrix = confusion_matrix(y_test1, y_pred)
results = pd.DataFrame(matrix, index = ["Dx_positif", "Dx_negatif"], columns = ["Predicted True", "Predicted False"])
results
# -
x = results["Predicted True"].value_counts()
y = results["Predicted False"].value_counts()
results.plot()
plt.xlabel("Diagnosis")
plt.ylabel("Numbers of Diagnosis")
plt.show()
# +
# Calculated the balanced accuracy score
from sklearn.metrics import accuracy_score
y_pred = predictions1
# Print the imbalanced classification report:
from imblearn.metrics import classification_report_imbalanced
print(f"Accuracy score: {accuracy_score(y_test1, y_pred)}")
print("------------------------------------------------------------------------------------")
print(f"Classification report: Balanced Random Forest Classifier ")
print("------------------------------------------------------------------------------------")
print(classification_report_imbalanced(y_test1, y_pred))
print("------------------------------------------------------------------------------------")
# -
| machine_learning/ML_model_2.0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In this notebook, you will
# - Learn the basic usage of several MRI functions in the `sigpy.mri` submodule
#
# https://sigpy.readthedocs.io/en/latest/mri.html
# %matplotlib notebook
import numpy as np
import sigpy as sp
import sigpy.mri as mr
import sigpy.plot as pl
# # Poisson disc sampling mask
# +
accel = 8
calib = [24, 24]
mask = mr.poisson([256, 256], accel, calib=calib)
pl.ImagePlot(mask)
# -
# # Radial sampling pattern
# +
num_ro = 256
num_tr = 64
img_shape = [256, 256]
golden = False
coord = mr.radial([num_tr, num_ro, 2], img_shape, golden=golden)
pl.ScatterPlot(coord)
# -
# # Simulate Birdcage Coilmaps
# +
mps = mr.birdcage_maps([8, 256, 256])
pl.ImagePlot(mps)
# -
| tutorial/basic/02b-mri_basic_usage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # X3FL dynamic relationships tool
# +
# just run and ignore this block
import numpy as np
import pandas as pd
all_races = ["Teladi", "NMMC", "Goner", "TerraCorp", "Strong Arms", "Argon", "Boron", "Split", "Atreus", "OTAS", "Duke's", "Paranid", "Pirates", "Terran", "Yaki"]
# https://imgur.com/nqI0nbO, https://imgur.com/7BhbUMa, by blazenclaw, forum.egosoft.com
try:
relationship_df = pd.read_csv("relationship.csv", index_col=0)
except:
relationship_df = pd.read_csv("https://raw.githubusercontent.com/mkmark/X3FL-dynamic-relationships/main/relationship.csv", index_col=0)
relationship_df = pd.DataFrame(relationship_df, columns=all_races)
def get_X(races):
R = np.array(relationship_df.loc[races, races])
invR = np.linalg.inv(R)
N = np.ones(len(races))
X = invR@N
return X
def get_X_df(races):
X = get_X(races)
return pd.DataFrame(list(X)+[sum(X)]+[len(races)/sum(X)], index=races+['sum', 'efficiency']).transpose()
# https://stackoverflow.com/questions/26332412/python-recursive-function-to-display-all-subsets-of-given-set
def subs(l):
if l == []:
return [[]]
x = subs(l[1:])
return x + [[l[0]] + y for y in x]
def get_X_optimal_sub(races):
possible_solution_racess = []
possible_solution_Xs = []
possible_solution_workloads = []
for sub_races in subs(races):
X = get_X(sub_races)
R_all = np.array(relationship_df.loc[races, sub_races])
N_all = R_all@X
min_N_all = min(N_all)
if min_N_all>0:
if min_N_all<1:
X = [x*1/min_N_all for x in X]
possible_solution_racess += [sub_races]
possible_solution_Xs += [X]
possible_solution_workloads += [sum(X)]
if len(possible_solution_workloads)>0:
min_index = possible_solution_workloads.index(min(possible_solution_workloads))
optiaml_solution_races = possible_solution_racess[min_index]
optiaml_solution_X = possible_solution_Xs[min_index]
if min(optiaml_solution_X)>0:
return pd.DataFrame(list(optiaml_solution_X)+[sum(optiaml_solution_X)]+[len(races)/sum(optiaml_solution_X)], index=optiaml_solution_races+['sum', 'efficiency']).transpose()
# -
# define target friend races as the following example, i.e. enemy of only Yaki, Terran, and Pirates
races = ["Teladi", "NMMC", "Goner", "TerraCorp", "Strong Arms", "Argon", "Boron", "Split", "Atreus", "OTAS", "Duke's", "Paranid"]
# get result with get_X_optimal_sub(target_friend_races)
get_X_optimal_sub(races)
# The number for each race represents how much effort you will have to put with that race to get 1 notoriety point for each race assuming you're to be friend with all these selected race.
#
# The sum is the total workload of current tactic.
#
# The efficiency is the sum of actual gained notoriety points (equal the number of races selected) divided by total workload.
# ---
# The above shows a very interesting discovery that the best strategy to achieve the current relationship, is not just to improving relationship with every one but the enemies, i.e. Yaki, Terran, and Pirates. It also suggests you that you **do not need to improve relationship with Teladi**, which is quite contrary to intuition.
#
# The logic behind this strategy is that by improving relationship with Teladi's ally, it would already make notoriety points of more than enough to achieve good relationship with Teladi, as verified above.
#
# See more in the analysis report [X3FL-dynamic-relationships.ipynb](https://colab.research.google.com/github/mkmark/X3FL-dynamic-relationships/blob/main/X3FL-dynamic-relationships.ipynb)
| X3FL-dynamic-relationships-tool.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Learning MNIST & Fashion
#
# In this exercise you will design a classifier for the very simple but very popular [MNIST dataset](http://yann.lecun.com/exdb/mnist/), a classic of dataset in computer vision and one of the first real world problems solved by neural networks.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers.core import Dense, Dropout, Activation
from keras.optimizers import SGD, Adam, RMSprop
from keras.utils import to_categorical
# -
# Keras provides access to a few simple datasets for convenience in the `keras.datasets` module. Here we will load MNIST, a standard benchmark dataset for image classification. This will download the dataset if you have run this code before.
# +
(X_train, y_train), (X_test, y_test) = mnist.load_data()
X_train.shape
# -
# MNIST is a simple dataset of grayscale hand-written digits 28x28 pixels big. So there are 10 classes in the dataset corresponding to the digits 0-9. We can get a sense for what this dataset is like (always a good idea) by looking at some random samples for the training data:
plt.imshow(X_train[np.random.randint(len(X_train))], cmap='gray')
# We need to do a little preprocessing of the dataset. Firstly, we will flatten the 28x28 images to a 784 dimensional vector. This is because our first model below does not care about the spatial dimensions, only the pixel values. The images are represented by numpy arrays of integers between 0 and 255. Since this is a fixed range, we should scale the values down to be from 0 to 1. This normalization simplifies things is usually a good idea, especially since weights are usually initialized randomly near zero.
#
# Read the code below and make sure you understand what we are doing to the data.
X_train = X_train.reshape(60000, 784)
X_test = X_test.reshape(10000, 784)
X_train = X_train.astype('float32')
X_test = X_test.astype('float32')
X_train /= 255
X_test /= 255
print(X_train.shape[0], 'train samples')
print(X_test.shape[0], 'test samples')
y_train_cat = to_categorical(y_train, 10)
y_test_cat = to_categorical(y_test, 10)
# ## Exercise 1 - design a fully conncted network for MNIST
#
# Build a fully connected network. It is up to you what the structure of the model will be, but keep in mind that this problem is much higher dimensional than previous problems we have worked on. This is your first chance to design a model on real data! See if you can get 90% accuracy or better.
#
# Here are some of the things you will need to decide about your model:
# * number of layers
# * activation function
# * number of dimensions in each layer
# * batch size
# * number of epochs
# * learning rate
#
# Suggestions:
# * You can pass the argument `verbose=2` to the `model.fit` method to quiet the output a bit, which will speed up the training as well.
# * You already divided the training and test data, but since you will be trying a series of experiments and changing your model, it is good practice to set aside a **validation** dataset for you to use to track your model improvements. You should only use the test data after you believe you have a good model to evaluate the final performance. Keras can create a validation set for you if you pass the `validation_split=0.1` argument to `model.fit` to tell Keras to hold out 10% of the training data to use as validation.
# * You can use the `plot_loss` if you find it useful in setting your learning rate etc. during your experiments.
# * You can refer to previous notebooks and the [documentation](http://keras.io/models/sequential/).
#
# If you want to talk over design decisions, feel free to ask.
def plot_loss(hist):
loss = hist.history['loss']
plt.plot(range(len(loss)), loss)
plt.title('loss')
plt.xlabel('epochs')
# + tags=["solution", "empty"]
model = Sequential()
model.add(Dense(28*28, input_dim=X_train.shape[1], activation='relu'))
model.add(Dense(10, activation='softmax'))
model.compile(loss='categorical_crossentropy',
optimizer=RMSprop(),
metrics=['accuracy'])
model.fit(X_train, y_train_cat, epochs=10, batch_size=1000,
verbose=2, validation_split=0.1)
# -
# Final test evaluation
score = model.evaluate(X_test, y_test_cat, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# ## Exercise 2: Fashion Mnist
# Repeat the classification exercise using the Fashion Mnist dataset from Zalando Research:
#
# https://github.com/zalandoresearch/fashion-mnist
#
# This dataset has the same specs as MNIST but it's designed to be more indicative of a real image classification problem. It contains 10 classes of clothing items:
#
# Label Description
# 0 T-shirt/top
# 1 Trouser
# 2 Pullover
# 3 Dress
# 4 Coat
# 5 Sandal
# 6 Shirt
# 7 Sneaker
# 8 Bag
# 9 Ankle boot
#
# Do you get to similar performance?
# + tags=["solution", "empty"]
from keras.datasets import fashion_mnist
# + tags=["solution"]
(X_train, y_train), (X_test, y_test) = fashion_mnist.load_data()
X_train = X_train.reshape(-1, 784).astype('float32') / 255
X_test = X_test.reshape(-1, 784).astype('float32') / 255
y_train_cat = to_categorical(y_train, 10)
y_test_cat = to_categorical(y_test, 10)
# + tags=["solution"]
model = Sequential()
model.add(Dense(28*28, input_dim=X_train.shape[1], activation='relu'))
model.add(Dense(10, activation='softmax'))
model.compile(loss='categorical_crossentropy',
optimizer=RMSprop(),
metrics=['accuracy'])
model.fit(X_train, y_train_cat, epochs=10, batch_size=1000,
verbose=2, validation_split=0.1)
# + tags=["solution"]
score = model.evaluate(X_test, y_test_cat, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# -
# *Copyright © 2017 CATALIT LLC. All rights reserved.*
| solutions_do_not_open/Lab_13_DL Learning MNIST and Fashion_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Project 3: Astronomical Source Detection
# ## <NAME>
# -------------------------------------------
# # Part 1 : SEP tutorial
#
# ## Start by importing numpy as per usual and, this time we will import SEP which helps us detect sources in an image and perform certain calculations
import numpy as np
import sep
# ## Instead of using fitsio, I will use astropy which means some adjustments will have to
# ## be made.
# ------------
# ## Will import fits from astropy.io
# +
from astropy.io import fits # import fits from astropy.io which will allow us to open fits files
import matplotlib.pyplot as plt # import this to plot whatever we may need to plot
from matplotlib import rcParams # allows us to vary certain parameters that will be used later
# %matplotlib inline
rcParams['figure.figsize'] = [10., 8.] # Set the size: do it once here, and never have to do it again in later in the
#code
# -
# ## Open the fits file, which was downloaded from SEP GitHub account
fname = "image.fits" # already in same folder
hdu_list = fits.open(fname)
hdu_list.info()
# ## Image information typically in PRIMARY block, accessed by indexing hdu_list
image_data = hdu_list["PRIMARY"].data
# ## Now we can close FITS file because stored as a variable
hdu_list.close()
# ## Showing the Data
# show the image
m, s = np.mean(image_data), np.std(image_data)
plt.imshow(image_data, interpolation='nearest', cmap='gray', vmin=m-s, vmax=m+s, origin='lower')
plt.colorbar();
plt.savefig('skyview.png' , bbox_inches="tight",dpi=600)
# ## Background subtraction time, taking away outliers, black is the outlier
bkg = sep.Background(image_data)
print(bkg.globalback)
print(bkg.globalrms)
bkg_image = bkg.back()
# bkg_image = np.array(bkg) # equivalent to above
# ## Show the Background, as well have save the figure
# +
plt.imshow(bkg_image, interpolation='nearest', cmap='gray', origin='lower')
plt.colorbar();
plt.savefig('background.png' , bbox_inches="tight",dpi=600)
# -
# ## Treat background noise as a 2-d array.
# evaluate the background noise as 2-d array, same size as original image
bkg_rms = bkg.rms()
# show the background noise
plt.imshow(bkg_rms, interpolation='nearest', cmap='gray', origin='lower')
plt.colorbar();
plt.savefig('noise.png' , bbox_inches="tight",dpi=600)
# ## Now we subtract the background
# +
# subtract the background
data_sub = image_data - bkg
# -
# ## Now we do object detection
#
# ## note that the number "1.5" refers to a sigma value or threshold value
# +
objects = sep.extract(data_sub, 1.5, err=bkg.globalrms)
len(objects) # how many objects were detected
from matplotlib.patches import Ellipse
# plot background-subtracted image
fig, ax = plt.subplots()
m, s = np.mean(data_sub), np.std(data_sub)
im = ax.imshow(data_sub, interpolation='nearest', cmap='gray',
vmin=m-s, vmax=m+s, origin='lower')
# plot an ellipse for each object
for i in range(len(objects)):
e = Ellipse(xy=(objects['x'][i], objects['y'][i]), # gives us the coordinates
width=6*objects['a'][i],
height=6*objects['b'][i],
angle=objects['theta'][i] * 180. / np.pi)
e.set_facecolor('none')
e.set_edgecolor('red')
ax.add_artist(e)
plt.savefig('tutorialfinal.png' , bbox_inches="tight",dpi=600) # saves the figure
# -
# ## The objects function has several uses/information which can be extracted and displayed
# available fields
objects.dtype.names
# ## Now we will show/calculate the fluxes of 9 objects in the image
# +
flux, fluxerr, flag = sep.sum_circle(data_sub, objects['x'], objects['y'],
3.0, err=bkg.globalrms, gain=1.0)
# show the first 10 objects results:
for i in range(10):
print("object {:d}: flux = {:f} +/- {:f}".format(i, flux[i], fluxerr[i]))
| astr-119-project-3-tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import json
import numpy
import matplotlib.pyplot as plt
from tqdm import tqdm_notebook
path_1 = "/home/yangz2/projects/pcgml_conditional_lstm/data/super_mario_bros"
path_2 = "/home/yangz2/projects/pcgml_conditional_lstm/data/super_mario_bros_2_japan"
annot_txts = [
os.path.join(path_1, fn) for fn in os.listdir(path_1) if fn.split('.')[-1] == 'txt'
] + [
os.path.join(path_2, fn) for fn in os.listdir(path_2) if fn.split('.')[-1] == 'txt'
]
print(len(annot_txts))
def load_txt(txt):
with open(txt, 'r') as txt_f:
return txt_f.readlines()
add_depth = False
with open('./corpusesmario_corpus.txt', 'w+') as txt_f:
for i, fp in enumerate(annot_txts):
infile = load_txt(fp)
lines = []
for line in infile:
lines.append(list(line.rstrip()))
infile_transposed = np.array(lines).T
print(infile_transposed.shape)
counter = 0
for line in infile_transposed: # each line represents an old column
num_chars_to_add = 16 - len(lines)
txt_f.write("".join(['-'] * num_chars_to_add + list(line)))
if add_depth:
if counter % 5 == 0:
txt_f.write("".join(["@"] * int(counter / 5)))
txt_f.write("\n")
counter += 1
if i+1 == len(annot_txts):
txt_f.write(")")
else:
txt_f.write(")\n")
# https://stackoverflow.com/questions/39457744/backpropagation-through-time-in-stateful-rnns
# https://github.com/keras-team/keras/issues/3669
| notebooks/.ipynb_checkpoints/data_preprocessing-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bias in Data
# ### <NAME>
# ### Data512
# ### 10/17/2109
# Imports the required libraries.
import json
import matplotlib.pyplot as plt
import pandas as pd
from pandas.io.json import json_normalize
import requests
import time
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# ### Obtaining the data
# Reads the data located in the raw_data folder.
# %cd ../raw_data
page_data = pd.read_csv("page_data.csv")
wpds_data = pd.read_csv("WPDS_2018_data.csv")
# %cd ../src
# Removes templates from page_data.
page_data = page_data[~page_data['page'].str.contains('template', case=False)]
# +
# Separates countries in different lists depending on region and takes the region data away from the wpds data.
# List of lists order: Africa, Northern America, Latin America, Asia, Europe, Oceania.
country_list = [[],[],[],[],[],[]]
i = -1
for country in wpds_data['Geography']:
if country.isupper():
i += 1
continue
country_list[i].append(country)
# Assign the region name to the list of countries.
africa = country_list[0]
north_america = country_list[1]
latin_america = country_list[2]
asia = country_list[3]
europe = country_list[4]
oceania = country_list[5]
wpds_data = wpds_data[~wpds_data['Geography'].str.isupper()]
# -
wpds_data.head()
page_data.head()
# +
# Function for the api call, edit the headers variable with your own information.
headers = {'User-Agent' : 'https://github.com/ablew', 'From' : '<EMAIL>'}
def get_ores_data(revision_ids, headers=headers):
"""
Uses reivision ids to make api requests to Ores.
:params:
:revision_ids list:
:headers dict:
returns response dict
"""
# Define the endpoint.
endpoint = 'https://ores.wikimedia.org/v3/scores/{project}/?models={model}&revids={revids}'
# Specify the parameters - joining all the revision IDs together separated by | marks.
params = {'project' : 'enwiki',
'model' : 'wp10',
'revids' : '|'.join(str(x) for x in revision_ids)}
api_call = requests.get(endpoint.format(**params))
response = api_call.json()
return response
# +
# obtains the predictions for the data in page_data using its rev_id column.
predictions = []
start_index = 0
while start_index < len(page_data):
# indices for the list of rev_ids to feed to the get_ores_data function.
end_index = start_index + 100
# if end_index is greater than the size of the data, it adjusts it.
if end_index > len(page_data):
end_index = len(page_data)
cropped_rev_id_list = list(page_data['rev_id'][start_index:end_index])
# Measures times so that the calls don't surpass 100 requests per second.
start_time = time.time()
api_call_result = get_ores_data(cropped_rev_id_list)
end_time = time.time()
api_time = end_time - start_time
if api_time <= 1:
time.sleep(1 - api_time)
# Creates list of predictions for the list of rev_ids fed into the get_ores_data function.
# Includes the value 'error' for when an error is encountered in the call.
cropped_predictions = []
for x in cropped_rev_id_list:
if list(api_call_result['enwiki']['scores'][str(x)]['wp10'].keys())[0] == 'error':
cropped_predictions.append('error')
continue
cropped_predictions.append(api_call_result['enwiki']['scores'][str(x)]['wp10']['score']['prediction'])
predictions = predictions + cropped_predictions
start_index += 100
# -
# There were 155 errors encountered in the api calls
predictions.count('error')
# ### Combining the data
# Predictions added to the page_data dataframe.
page_data['prediction'] = predictions
page_data.head()
# Left joins page_data with wpds_data on country
joined_raw_data = pd.merge(page_data, wpds_data, left_on='country', right_on='Geography', how='left')
joined_raw_data.head()
# Creates dataframe for the rows that had no matches when joining and saves it into a .csv file.
wp_wpds_countries_no_match = joined_raw_data[pd.isnull(joined_raw_data).any(axis=1)]
wp_wpds_countries_no_match.head()
wp_wpds_countries_no_match.to_csv('wp_wpds_countries-no_match.csv')
# ### Cleaning the data
# +
# Creates dataframe for the remaining data, reorders and renames the columns.
col_names = ['country', 'page', 'rev_id', 'prediction', 'Population mid-2018 (millions)']
new_col_names = ['country', 'article_name', 'revision_id', 'article_quality', 'population']
wp_wpds_politicians_by_country = joined_raw_data[~pd.isnull(joined_raw_data).any(axis=1)]
wp_wpds_politicians_by_country = wp_wpds_politicians_by_country[col_names]
wp_wpds_politicians_by_country.columns = new_col_names
# Finds articles with errors in their api calls and saves them as a .csv file
wp_wpds_api_call_errors = wp_wpds_politicians_by_country[wp_wpds_politicians_by_country['article_quality'].str.contains('error')]
wp_wpds_api_call_errors.to_csv('wp_wpds_api_call_errors.csv')
# Takes the remainder of the data with no errors
wp_wpds_politicians_by_country = wp_wpds_politicians_by_country[~wp_wpds_politicians_by_country['article_quality'].str.contains('error')]
# Converts population column from str to int.
wp_wpds_politicians_by_country['population'] = wp_wpds_politicians_by_country['population'].str.replace(',', '')
wp_wpds_politicians_by_country['population'] = pd.to_numeric(wp_wpds_politicians_by_country['population'])
wp_wpds_politicians_by_country['population'] = wp_wpds_politicians_by_country['population'] * 1000000
wp_wpds_politicians_by_country.head()
# Exports to .csv file.
wp_wpds_politicians_by_country.to_csv('wp_wpds_politicians_by_country.csv')
# -
# ### Analysis
# Obtains the numbers of articles per country.
count_table = wp_wpds_politicians_by_country.groupby(['country']).count()
country_data = pd.DataFrame(count_table['article_name'])
country_data = country_data.reset_index()
country_data.columns = ['country', 'num_article']
country_data.head()
# +
# Adds population to the country_data dataframe.
# Creates dataframe with country and population.
population_data = wp_wpds_politicians_by_country[['country', 'population']]
population_data = population_data.drop_duplicates()
# Joins population dataframe to country_data.
country_data = pd.merge(country_data, population_data, on='country', how='left')
# Coverage: number of articles divided by population. Multiplied by 100 for the percentage
country_data['coverage'] = (country_data['num_article'] / country_data['population']) * 100
country_data.head()
# +
# Adds a column for each value in article_quality to country_data.
# Creates a dataframe obtained from grouping by country and article quality.
article_quality_by_country = wp_wpds_politicians_by_country.groupby(['country','article_quality']).size()
article_quality_by_country = pd.DataFrame(article_quality_by_country)
article_quality_by_country = article_quality_by_country.reset_index()
article_quality_by_country.columns = ['country', 'article_quality', 'count']
# Pivots data to obtain a column for each value.
article_quality_by_country = article_quality_by_country.pivot(index='country', columns='article_quality', values='count')
article_quality_by_country = article_quality_by_country.fillna(0)
article_quality_by_country = article_quality_by_country.reset_index()
# Joins the dataframe to country_data
country_data = pd.merge(country_data, article_quality_by_country, on='country', how='left')
country_data.head()
# -
# Creates a column for the proportion of high quality articles (rated FA and GA).
country_data['high_quality'] = ((country_data['FA'] + country_data['GA']) / country_data['num_article']) * 100
country_data.head()
# +
# Creates a column for region in the country_data dataframe
region = []
for country in country_data['country']:
if country in africa:
region.append('AFRICA')
elif country in north_america:
region.append('NORTHERN AMERICA')
elif country in latin_america:
region.append('LATIN AMERICA AND THE CARIBBEAN')
elif country in asia:
region.append('ASIA')
elif country in europe:
region.append('EUROPE')
elif country in oceania:
region.append('OCEANIA')
else:
print('error')
break
country_data['region'] = region
country_data.head()
# +
# Creates a dataframe for region by grouping by region and performing a sum.
# Coverage and high_quality have to be corrected.
region_data = country_data.groupby('region').sum()
region_data['coverage'] = (region_data['num_article'] / region_data['population']) * 100
region_data['high_quality'] = ((region_data['FA'] + region_data['GA']) / region_data['num_article']) * 100
region_data = region_data.reset_index()
region_data
# -
# makes a bar plot for high-quality articles by region
plt.bar(list(range(6)), region_data['high_quality'])
plt.xticks(list(range(6)), ['Africa', 'Asia', 'Europe', 'Latin America', 'North America', 'Oceania'], rotation=45)
plt.title('% of High Quality Articles by Region')
plt.xlabel('Regions')
plt.ylabel('% of high quality articles')
plt.savefig('high_quality_by_region.png')
plt.show()
# makes a bar plot for coverage articles by region
plt.bar(list(range(6)), region_data['coverage'])
plt.xticks(list(range(6)), ['Africa', 'Asia', 'Europe', 'Latin America', 'North America', 'Oceania'], rotation=45)
plt.title('Coverage % by Region')
plt.xlabel('Regions')
plt.ylabel('Coverage %')
plt.savefig('coverage_by_region.png')
plt.show()
# Saves country and region data as .csv files.
country_data.to_csv('wp_by_country.csv')
region_data.to_csv('wp_by_region.csv')
#
#
# ### Results
# #### Top 10 countries by coverage
# ##### "10 highest-ranked countries in terms of number of politician articles as a proportion of country population."
# Creates table for the top 10 countries by coverage.
country_data.sort_values(by='coverage', ascending=False).head(10)[['country', 'coverage']].style.hide_index()
# #### Bottom 10 countries by coverage
# ##### "10 lowest-ranked countries in terms of number of politician articles as a proportion of country population."
# Creates table for the bottom 10 countries by coverage.
country_data.sort_values(by='coverage', ascending=True).head(10)[['country', 'coverage']].style.hide_index()
# #### Top 10 countries by relative quality
# ##### "10 highest-ranked countries in terms of the relative proportion of politician articles that are of GA and FA-quality."
# Creates the table for the top 10 countries by relative quality.
country_data.sort_values(by='high_quality', ascending=False).head(10)[['country', 'high_quality']].style.hide_index()
# Checks for row pertaining North Korea for curiosity.
country_data[country_data['country'] == 'Korea, North']
# #### Bottom 10 countries by relative quality
# ##### "10 lowest-ranked countries in terms of the relative proportion of politician articles that are of GA and FA-quality."
# Creates table for the bottom 10 countries by relative quality.
country_data.sort_values(by='high_quality', ascending=True)[['country', 'high_quality']].head(10).style.hide_index()
print("There are {0} countries with 0 GA or FA-quality articles.".format(sum(country_data['high_quality'] == 0)))
# This table shows the number of countries that had 0 GA or FA-quality articles by region.
# Creates table for number of countries with 0 high quality articles by region.
country_data[country_data['high_quality'] == 0][['country', 'region']].groupby('region').count()
# The offending countries are:
# Lists the countries with 0 high quality articles.
zero_high_quality = country_data[country_data['high_quality'] == 0]['country']
zero_high_quality
# Makes a bar plot for number of articles by countries with 0 high quality articles.
plt.bar(list(range(len(zero_high_quality))), country_data[country_data['high_quality'] == 0]['num_article'])
plt.xticks(list(range(len(zero_high_quality))), zero_high_quality, rotation=45)
plt.title('number of articles by countries with 0 high-quality articles')
plt.xlabel('Country')
plt.ylabel('Number of articles')
plt.show()
# This table shows the coverage in countries with 0 high quality articles, per region.
zero_hq_data = country_data[country_data['high_quality'] == 0]
zero_hq_by_region = zero_hq_data.groupby('region').sum()
zero_hq_by_region['coverage'] = zero_hq_by_region['num_article'] / zero_hq_by_region['population'] * 100
zero_hq_by_region[['coverage']]
# We can see that coverage is extremely low in countries with 0 high-quality articles, meaning that the population has little information about politicians in their country, and what little information they have is not high quality.
# #### Top 10 countries by coverage with 0 high quality articles
# ##### "10 highest-ranked countries in terms of number of politician articles as a proportion of country population in countries with 0 high quality articles."
# Creates table for the top 10 countries with 0 high quality articles by coverage.
zero_hq_data.sort_values(by='coverage', ascending=False).head(10)[['country', 'coverage']].style.hide_index()
# I find the table above particularly interesting because these are the countries with the most misinformation.
# #### Bottom 10 countries by coverage with 0 high quality articles
# ##### "10 lowest-ranked countries in terms of number of politician articles as a proportion of country population with 0 high quality articles."
# Creates table for the bottom 10 countries with 0 high quality articles by coverage.
zero_hq_data.sort_values(by='coverage', ascending=True).head(10)[['country', 'coverage']].style.hide_index()
# The table above is also interesting because this would show the countries with least, most of them being from Africa or third-world countries in Asia.
# For those countries that have at least one FA or GA-quality articles, these are the 10 lowest ranked.
# Creates table for bottom 10 countries with at least 1 high quality article by relative quality.
(country_data[country_data['high_quality'] != 0]).sort_values(by='high_quality', ascending=True)[['country', 'high_quality']].head(10).style.hide_index()
# Checks on the row pertaining Belgium.
country_data[country_data['country'] == 'Belgium']
# #### Geographic regions by coverage
# ##### "Ranking of geographic regions (in descending order) in terms of the total count of politician articles from countries in each region as a proportion of total regional population."
# Creates table for the regions by coverage.
region_data.sort_values(by='coverage', ascending=False)[['region', 'coverage']].style.hide_index()
# #### Geographic regions by relative quality
# ##### "Ranking of geographic regions (in descending order) in terms of the relative proportion of politician articles from countries in each region that are of GA and FA-quality."
# Creates table for the regions by relative quality.
region_data.sort_values(by='high_quality', ascending=False)[['region', 'high_quality']].style.hide_index()
#
#
# ### Reflections
# Before starting to work on the data, I expected articles from developing countries, and non-democratic countries to receive a high amount of low scores in the predictions. This is because I suspected that articles from developing countries would be less regulated, partly due to the population's inability to access the internet. In addition, countries like China and North Korea could censor or edit articles in order to promote their political agendas. One example of how my preconceptions were challenged is the fact that North Korea is ranked first in the list of top 10 countries with high-quality articles. It could be explained by the fact that there are a few number of articles related to North Korea, or that other entities or organizations took it upon themselves to write the articles, since it cannot be expected for residents in North Korea to write or edit articles on the English Wikipedia website.
#
# The results suggest that Wikipedia is not that really as a data source when it comes to political articles. This might be because anyone can edit Wikipedia, and there could be several political parties trying to improve their standing on the internet by editing Wikipedia articles. While we assume that the data of the classifications made by people is correct, there is a chance that they could also introduce bias in the predictive model by classifying an article they do not agree with as a low quality article. And lastly, further testing should be done regarding the quality of Wikipedia articles by choosing a different subject, such as science, where the information is more factual. While it was disheartening to see that Wikipedia has a low percentage of high quality articles, the fact that low value is present in articles related to every country and region gives us a warning to always be on the lookout for bias since, as the results show, is everywhere.
#
# For further work, it would be interesting to use another classifier built by different people to see if the predictions are accurate, and if not, could the bias be in the classification of the articles during the training phase? It would also be interesting if the datasets had more features, such as the political teams for each country and for each article and the percentage of their representation in the government. Looking into the correlation between coverage and the percentage of high-quality articles also seems like a promising path for future work.
#
#
# Table subtitles obtained from: https://wiki.communitydata.science/Human_Centered_Data_Science_(Fall_2019)/Assignments#A2:_Bias_in_data
| src/hcds-a2-bias-in-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# 영화 과람객 예측
# 목적: test 데이터의 box_off_num을 예측한다.
# +
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
from sklearn import datasets
from sklearn.model_selection import train_test_split
#from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import RandomForestRegressor
from sklearn import metrics
# -
train = pd.read_csv('../input/movies_train.csv')
test = pd.read_csv('../input/movies_test.csv')
train.tail()
test.tail()
train.columns
x = data[['sepal length', 'sepal width', 'petal length', 'petal width']]
y = data['species']
# TODO 각 필드의 의미;
# TOdo 필드에서 취할 것 정하기
x = train[['genre', 'screening_rat', 'director', 'dir_prev_bfnum', 'dir_prev_num', 'num_actor']]
y = train['box_off_num']
# 테스트 데이터 30%
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.3)
print(len(x_train))
print(len(x_test))
print(len(y_train))
print(len(y_test))
x_train['genre'].unique()
x_train.head(13)
x_train.genre.astype('category').cat.codes # x_train 5: 멜로/로멘스 10: 액션 3: 다큐멘터리 4: 드라마 11: 코메디 1: 공포
x_test.head()
test.genre.astype('category').cat.codes.head()
test.head()
x_test.genre.astype('category').cat.codes # x_train 5: 멜로/로멘스 10: 액션 3: 다큐멘터리 4: 드라마 11: 코메디 1: 공포
# x_test 4: 드라마 11: 코미디 5: 멜로/로맨스 1: 공포
# +
#x_train.loc[:,'genre'] = pd.Categorical(x_train.loc[:,'genre'])
#df['code'] = df.cc.cat.codes
x_train.genre = pd.Categorical(x_train.genre)
x_train['genre_cd'] = x_train.genre.cat.codes
x_train.screening_rat = pd.Categorical(x_train.screening_rat)
x_train['screening_rat_cd'] = x_train.screening_rat.cat.codes
x_train.director = pd.Categorical(x_train.director)
x_train['director_cd'] = x_train.director.cat.codes
# +
x_test.genre = pd.Categorical(x_test.genre)
x_test['genre_cd'] = x_test.genre.cat.codes
x_test.screening_rat = pd.Categorical(x_test.screening_rat)
x_test['screening_rat_cd'] = x_test.screening_rat.cat.codes
x_test.director = pd.Categorical(x_test.director)
x_test['director_cd'] = x_test.director.cat.codes
# -
x_test.columns
x_train_p
x_train_p.fillna(0)
x_test_p.fillna(0)
x_train_p
x_train_p = x_train[[ 'genre_cd', 'screening_rat_cd', 'director_cd', 'dir_prev_num','num_actor']]
x_test_p = x_test[[ 'genre_cd', 'screening_rat_cd', 'director_cd', 'dir_prev_num','num_actor']]
x_train_p
# +
# 학습 진행
forest = RandomForestRegressor(n_estimators=100)
forest.fit(x_train_p, y_train)
# 예측
y_pred = forest.predict(x_test_p)
# 정확도 확인
#print('정확도 :', metrics.accuracy_score(y_test, y_pred))
# -
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
# +
y_pos = np.arange(len(y_test))
audience = y_test.values
plt.bar(y_pos, audience, align='center', alpha=0.5)
#plt.xticks(y_pos, audience)
plt.ylabel('audience')
plt.title('Movie audience')
plt.show()
# +
y_pos = np.arange(len(y_pred))
audience = y_pred
plt.bar(y_pos, audience, align='center', alpha=0.5)
#plt.xticks(y_pos, audience)
plt.ylabel('audience')
plt.title('Movie audience')
plt.show()
# -
#y_test_pd
y_test.values
>>> import seaborn as sns
>>> sns.set(style="whitegrid")
>>> tips = sns.load_dataset("tips")
>>> ax = sns.barplot(x="day", y="total_bill", data=tips)
type(tips)
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.datasets import make_moons
X, y = make_moons(n_samples=100, noise=0.25, random_state=3)
X_train, X_test, y_train, y_test = train_test_split(X, y, stratify=y,
random_state=42)
forest = RandomForestClassifier(n_estimators=5, random_state=2)
forest.fit(X_train, y_train)
# +
import mglearn
fig, axes = plt.subplots(2, 3, figsize=(20, 10))
for i, (ax, tree) in enumerate(zip(axes.ravel(), forest.estimators_)):
ax.set_title("트리 {}".format(i))
mglearn.plots.plot_tree_partition(X, y, tree, ax=ax)
mglearn.plots.plot_2d_separator(forest, X, fill=True, ax=axes[-1, -1], alpha=.4)
axes[-1, -1].set_title("랜덤 포레스트")
mglearn.discrete_scatter(X[:, 0], X[:, 1], y)
# -
# !pip install mglearn
from sklearn.datasets import load_breast_cancer
cancer = load_breast_cancer()
# +
X_train, X_test, y_train, y_test = train_test_split(
cancer.data, cancer.target, random_state=0)
forest = RandomForestClassifier(n_estimators=100, random_state=0)
forest.fit(X_train, y_train)
print("훈련 세트 정확도: {:.3f}".format(forest.score(X_train, y_train)))
print("테스트 세트 정확도: {:.3f}".format(forest.score(X_test, y_test)))
# -
def plot_feature_importances_cancer(model):
n_features = cancer.data.shape[1]
plt.barh(range(n_features), model.feature_importances_, align='center')
plt.yticks(np.arange(n_features), cancer.feature_names)
plt.xlabel("attr importances")
plt.ylabel("attr")
plt.ylim(-1, n_features)
plot_feature_importances_cancer(forest)
| work/expect_audience.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.5.0-dev
# language: julia
# name: julia-0.5
# ---
# # Tel
# ### format event_type.csv
function reformat(data::Array{ASCIIString,2})
data = data[2:end,:]
id_event = Dict{ASCIIString,Set{ASCIIString}}()
for i in 1:size(data,1)
id = data[i,1]
if in(id,keys(id_event))
push!(id_event[id],data[i,1])
else
id_event[id] = Set{ASCIIString}([data[i,2]])
end
end
colname = sort(unique(data[:,2]))
lencol = length(colname)
shape = length(unique(data[:,1])),length(colname)+1
result = Array{ASCIIString,2}(shape)
for (ind,id) in enumerate(keys(id_event))
vec = Array{ASCIIString,1}(lencol)
events = id_event[id]
for (ind,col) in enumerate(colname)
if in(col,events)
vec[ind] = "1"
else
vec[ind] = "0"
end
end
result[ind,:] = vcat(id,vec)
@show result[ind,:]
end
result
end
ev = readcsv("../data/event_type.csv",ASCIIString)
reformat(ev)
unique(ev[:,2])
| notebook/.ipynb_checkpoints/process-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="nCc3XZEyG3XV"
# Lambda School Data Science
#
# *Unit 2, Sprint 3, Module 2*
#
# ---
#
#
# # Permutation & Boosting
#
# You will use your portfolio project dataset for all assignments this sprint.
#
# ## Assignment
#
# Complete these tasks for your project, and document your work.
#
# - [ ] If you haven't completed assignment #1, please do so first.
# - [ ] Continue to clean and explore your data. Make exploratory visualizations.
# - [ ] Fit a model. Does it beat your baseline?
# - [ ] Try xgboost.
# - [ ] Get your model's permutation importances.
#
# You should try to complete an initial model today, because the rest of the week, we're making model interpretation visualizations.
#
# But, if you aren't ready to try xgboost and permutation importances with your dataset today, that's okay. You can practice with another dataset instead. You may choose any dataset you've worked with previously.
#
# The data subdirectory includes the Titanic dataset for classification and the NYC apartments dataset for regression. You may want to choose one of these datasets, because example solutions will be available for each.
#
#
# ## Reading
#
# Top recommendations in _**bold italic:**_
#
# #### Permutation Importances
# - _**[Kaggle / <NAME>: Machine Learning Explainability](https://www.kaggle.com/dansbecker/permutation-importance)**_
# - [<NAME>: Interpretable Machine Learning](https://christophm.github.io/interpretable-ml-book/feature-importance.html)
#
# #### (Default) Feature Importances
# - [<NAME>: Selecting good features, Part 3, Random Forests](https://blog.datadive.net/selecting-good-features-part-iii-random-forests/)
# - [<NAME>, et al: Beware Default Random Forest Importances](https://explained.ai/rf-importance/index.html)
#
# #### Gradient Boosting
# - [A Gentle Introduction to the Gradient Boosting Algorithm for Machine Learning](https://machinelearningmastery.com/gentle-introduction-gradient-boosting-algorithm-machine-learning/)
# - _**[A Kaggle Master Explains Gradient Boosting](http://blog.kaggle.com/2017/01/23/a-kaggle-master-explains-gradient-boosting/)**_
# - [_An Introduction to Statistical Learning_](http://www-bcf.usc.edu/~gareth/ISL/ISLR%20Seventh%20Printing.pdf) Chapter 8
# - [Gradient Boosting Explained](http://arogozhnikov.github.io/2016/06/24/gradient_boosting_explained.html)
# - _**[Boosting](https://www.youtube.com/watch?v=GM3CDQfQ4sw) (2.5 minute video)**_
# -
| module2/assignment_applied_modeling_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Google Traces Parser - clusterdata-2011-2
# This Python notebook parse the google trace called "clusterdata-2011-2" and create the necessary trace files required to run "cluster-scheduler-simulator".
#
# Usefull links:
# <ul>
# <li>
# <a href="https://github.com/google/cluster-data"> GitHub </a>
# </li>
# <li>
# <a href="https://drive.google.com/open?id=0B5g07T_gRDg9Z0lsSTEtTWtpOW8&authuser=0"> Format + Schema Document </a>
# </li>
# <li>
# <a href="https://groups.google.com/forum/#!forum/googleclusterdata-discuss"> Mailing List </a>
# </li>
# </ul>
# Notes:
# <ul>
# <li>
# All resources utilization are normalized as explained in the "Format + Schema Document" in the above link.
# For this reason we will adjust them to real values using the same cell size of the "cluster-scheduler-simulator".
# File "init-cluster-state.log" expect real value of CPU and Memory to calculate the distribution.
# </li>
# </ul>
# +
from pyspark import SparkConf, SparkContext
from pyspark.sql import SQLContext
from pyspark.sql.types import *
import pyspark.sql.functions as func
import os
import errno
from collections import OrderedDict
# +
# Config values
CLUSTER_INFO = {
"mem_machine": 128 * (1024**3),
"cpu_machine": 32
}
SIMULATOR_TRACES_OUTPUT_FOLDER = "google-simulator-traces/"
JOB_TRACES_OUTPUT_FOLDER = "job-distribution-traces/"
# +
def mkdir_p(path):
path = path.replace(" ", "_")
dir_path = os.path.dirname(path)
try:
os.makedirs(dir_path)
except OSError as exc: # Python >2.5
if exc.errno == errno.EEXIST and os.path.isdir(dir_path):
pass
else:
raise
return path
# Create output folder structure
SIMULATOR_TRACES_OUTPUT_FOLDER = mkdir_p(os.path.join(".", SIMULATOR_TRACES_OUTPUT_FOLDER))
JOB_TRACES_OUTPUT_FOLDER = mkdir_p(os.path.join(SIMULATOR_TRACES_OUTPUT_FOLDER, JOB_TRACES_OUTPUT_FOLDER))
# +
# conf = (
# SparkConf()
# .set("spark.master", os.environ["SPARK_MASTER"])
# .set("spark.executor.memory", os.environ["SPARK_EXECUTOR_RAM"])
# )
# sc.stop()
# sc=SparkContext(conf=conf)
sqlContext = SQLContext(sc)
schema = StructType([ \
StructField("file_pattern", StringType(), False), \
StructField("field_number", IntegerType(), False), \
StructField("content", StringType(), False), \
StructField("format", StringType(), False), \
StructField("mandatory", StringType(), False)])
schema_all = sqlContext.read.format("com.databricks.spark.csv").options(header='true').load("clusterdata-2011-2/schema.csv", schema=schema)
# Displays the content of the DataFrame to stdout
schema_all.show(n=schema_all.count(), truncate=False)
format_type = {
"INTEGER": DecimalType(32,0),
"STRING_HASH": StringType(),
"FLOAT": FloatType(),
"BOOLEAN": DecimalType(),
"STRING_HASH_OR_INTEGER": StringType()
}
schemas = {}
for line in schema_all.orderBy("file_pattern", "field_number").select("*").collect():
line = line.asDict()
try:
if line["file_pattern"] not in schemas:
schemas[line["file_pattern"]] = []
schemas[line["file_pattern"]].append(
StructField(
line["content"].replace(' ', '_'),
format_type[line["format"]],
line["mandatory"] != "YES"
)
)
except KeyError as e:
print(e)
dataframes = {}
for file_pattern in schemas:
dataframes[os.path.dirname(file_pattern)] = sqlContext.read.format("com.databricks.spark.csv").load("clusterdata-2011-2/" + file_pattern, schema=StructType(schemas[file_pattern]))
max_time = pow(2,63) - 1
# +
job_events = dataframes["job_events"]
#job_events.show(truncate=False)
task_events = dataframes["task_events"]
#task_events.show(truncate=False)
task_usage = dataframes["task_usage"]
#task_usage.show(truncate=False)
machine_events = dataframes["machine_events"]
#machine_events.show(truncate=False)
# -
jobs_already_running = job_events[(job_events.event_type == 1) & (job_events.time == 0)]\
.select(job_events.job_ID, job_events.scheduling_class)
print("Jobs submitted before the event time window: {}".format(jobs_already_running.count()))
finished_jobs_already_running = {}
for job in job_events[job_events.event_type == 4].join(jobs_already_running, on="job_ID")\
.select(job_events.job_ID, job_events.time).collect():
if job.job_ID not in finished_jobs_already_running:
finished_jobs_already_running[job.job_ID] = job.time
print("Jobs submitted before that endend during the events time window: {}".format(len(finished_jobs_already_running)))
production_jobs_already_running = {}
for job in task_events[(task_events.event_type == 0) & (task_events.priority >= 9)].join(jobs_already_running, on="job_ID")\
.groupBy(task_events.job_ID).count().collect():
if job.job_ID not in production_jobs_already_running:
production_jobs_already_running[job.job_ID] = 1
print("Production jobs submitted before the events time window: {}".format(len(production_jobs_already_running)))
tasks_jobs_already_running = {}
for job in task_events[task_events.event_type == 0].join(jobs_already_running, on="job_ID")\
.groupBy(task_events.job_ID).agg(func.max("task_index").alias("tasks")).collect():
if job.job_ID not in tasks_jobs_already_running:
tasks_jobs_already_running[job.job_ID] = job.tasks + 1 # task_index starts from 0
print("Jobs submitted before the event time window that have at least 1 task: {}".format(len(tasks_jobs_already_running)))
resources_jobs_already_running = {}
for job in task_usage.join(jobs_already_running, on="job_ID")\
.groupBy(task_usage.job_ID, task_usage.task_index)\
.agg(\
func.avg("CPU_rate").alias("cpu"),\
func.avg("canonical_memory_usage").alias("memory")
)\
.groupBy(task_usage.job_ID)\
.agg(\
func.sum("cpu").alias("cpu"),\
func.sum("memory").alias("memory")
)\
.collect():
if job.job_ID not in resources_jobs_already_running:
resources_jobs_already_running[job.job_ID] = {
'cpu': CLUSTER_INFO["cpu_machine"] * job.cpu,
'memory': CLUSTER_INFO["mem_machine"] * job.memory
}
print("Jobs processed: {}".format(len(resources_jobs_already_running)))
job_count = 0
with open(os.path.join(SIMULATOR_TRACES_OUTPUT_FOLDER, "init-cluster-state.log"), "w") as init_cluster_state:
for job in jobs_already_running.collect():
# === Common Columns ===
# Column 0: possible values are 11 or 12
# "11" - (8 column schema) something that was there at the beginning of timewindow
# "12" - (6 column schema) something that was there at beginning of timewindow and ended at [timestamp] (see Column 1)
# Column 1: timestamp
# Column 2: unique job ID
# Column 3: 0 or 1 - prod_job - boolean flag indicating if this job is "production" priority as described in [1]
# Column 4: 0, 1, 2, or 3 - sched_class - see description of "Scheduling Class" in [1]
column_0 = 11
column_1 = 0
column_2 = job.job_ID
column_3 = 0
column_4 = job.scheduling_class
# === 6 column format ===
# Column 5: UNSPECIFIED/UNUSED
#
# === 8 column format ===
# Column 5: number of tasks
# Column 6: aggregate CPU usage of job (in num cores)
# Column 7: aggregate Ram usage of job (in bytes)
column_5 = 0
column_6 = 0
column_7 = 0
if job.job_ID in production_jobs_already_running:
column_3 = 1
if job.job_ID in tasks_jobs_already_running:
column_5 = tasks_jobs_already_running[job.job_ID]
# What's the point in adding a job with 0 tasks to the trace?
# Also, if we do so we will get an error
if column_5 == 0:
continue
if job.job_ID in resources_jobs_already_running:
column_6 = resources_jobs_already_running[job.job_ID]["cpu"]
column_7 = int(resources_jobs_already_running[job.job_ID]["memory"])
init_cluster_state.write("{} {} {} {} {} {} {} {}\n".format(
column_0, column_1, column_2, column_3, column_4, column_5, column_6, column_7))
if job.job_ID in finished_jobs_already_running:
column_0 = 12
column_1 = finished_jobs_already_running[job.job_ID]
init_cluster_state.write("{} {} {} {} {} {}\n".format(
column_0, column_1, column_2, column_3, column_4, column_5))
job_count += 1
print("Jobs processed: {}".format(job_count))
job_that_finished = job_events[((job_events.event_type == 1) | (job_events.event_type == 4)) & ((job_events.time > 0) & (job_events.time < max_time))]\
.orderBy(job_events.time).groupBy(job_events.job_ID).count().where("count = 2")
jobs = OrderedDict({})
for job in job_events[(job_events.event_type == 1)].join(job_that_finished, on="job_ID")\
.orderBy(job_events.time).select(job_events.job_ID, job_events.time, job_events.scheduling_class)\
.collect():
if job.job_ID not in jobs:
jobs[job.job_ID] = {
"production": False,
"tasks": 0,
"start": 0,
"end": 0,
"scheduling_class": None
}
jobs[job.job_ID]["start"] = job.time
jobs[job.job_ID]["scheduling_class"] = job.scheduling_class
print("Jobs submitted during the event time window: {}".format(len(jobs)))
job_count = 0
for job in job_events[(job_events.event_type == 4)].join(job_that_finished, on="job_ID")\
.select(job_events.job_ID, job_events.time).collect():
if job.job_ID in jobs:
jobs[job.job_ID]["end"] = job.time
job_count += 1
print("Jobs processed: {}".format(job_count))
job_count = 0
for job in task_events[task_events.event_type == 0].join(job_that_finished, on="job_ID")\
.groupBy(task_events.job_ID).agg(func.max("task_index").alias("tasks")).collect():
if job.job_ID in jobs:
jobs[job.job_ID]["tasks"] = job.tasks + 1
job_count += 1
print("Jobs processed: {}".format(job_count))
job_count = 0
for job in task_events[(task_events.event_type == 0) & (task_events.priority >= 9)].join(job_that_finished, on="job_ID") \
.groupBy(task_events.job_ID).count().collect():
if job.job_ID in jobs:
jobs[job.job_ID]["production"] = True
job_count += 1
print("Jobs processed: {}".format(job_count))
# +
interarrival_cmb = open(os.path.join(JOB_TRACES_OUTPUT_FOLDER, "interarrival_cmb.log"), "w")
runtimes_cmb = open(os.path.join(JOB_TRACES_OUTPUT_FOLDER, "runtimes_cmb.log"), "w")
csizes_cmb = open(os.path.join(JOB_TRACES_OUTPUT_FOLDER, "csizes_cmb.log"), "w")
previous_arrival = 0
for job in jobs:
job = jobs[job]
# What's the point in adding a job with 0 tasks to the trace?
# Also, if we do so we will get an error
if job["tasks"] == 0:
continue
# === Columns ===
# Column 0: cluster_name
# Column 1: assignment policy ("cmb-new" = "CMB_PBB")
# Column 2: scheduler id, values can be 0 or 1. 0 = batch, service = 1
# Column 3: depending on which trace file:
# interarrival time (seconds since last job arrival)
# OR tasks in job
# OR job runtime (seconds)
column_0 = "test"
column_1 = "cmb-new"
column_2 = 1 if job["production"] and (job["scheduling_class"] != 0 and job["scheduling_class"] != 1) else 0
if job["start"] - previous_arrival < 0:
print("Error. Dataset is not ordered by arrival time!")
break
interarrival_cmb.write("{} {} {} {} \n".format(column_0, column_1, column_2,
(job["start"] - previous_arrival)/1000000))
runtimes_cmb.write("{} {} {} {} \n".format(column_0, column_1, column_2,
(job["end"] - job["start"])/1000000))
csizes_cmb.write("{} {} {} {} \n".format(column_0, column_1, column_2,
job["tasks"]))
previous_arrival = job["start"]
interarrival_cmb.close()
runtimes_cmb.close()
csizes_cmb.close()
| script/Google Traces Parser - clusterdata-2011-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python 網頁爬蟲與資料視覺化應用 Final Project
# ## 成大通識與各系必修撞課分析
# 作者:物理系 施宇庭 C24066096
#
# 1. [問題定義](#問題定義)
# 2. [資料收集](#資料收集)
# 3. [資料前處理](#資料前處理)
# 4. [視覺化呈現](#視覺化呈現)
# 5. [結果分析與討論](#結果分析與討論)
# ## 問題定義
# [回頂部](#成大通識與各系必修撞課分析)
# 選課季節又到了,,但因為和系上必修撞課,很難選到又甜又涼的通識,,,因此,這次期末專題打算來做各系必修與通識的撞課情況分析
#
# 統計各個科系必修的上課時間中有多少通識撞課,
#
# 備註:研究所不需要修通識課,因此這份報告將專注在大學部的部分
#
# 希望藉由這份分析找出,提供各系及通識中心排課參考,以平衡各系學生選擇通識課程的權益
# ## 資料收集
# 資料來源:[NCKU Open Data Platform](http://data.ncku.edu.tw/dataset/8d6462ee513f1c1a601506265cc9f674)
#
# 可從上方連結下載資料,或是直接執行專案目錄下的 `get_data.sh` 來下載資料,預設儲存路徑為 `data/`
# ```bash
# source get_data.sh [儲存路徑]
# ```
#
# **注意:若自行給定下載路徑,則下方程式碼的路徑也要記得修改**
#
# [回頂部](#成大通識與各系必修撞課分析)
# 匯入所需套件
import pandas as pd
# 載入資料並觀察,由於資料的完整性,我們先拿106學年度第2學期的資料進行分析
path = './data/1062.csv'
df = pd.read_csv(path, encoding='cp950')
df.head(3)
# ## 資料前處理
# 1. 查看資料中是否包含所有我們需要的欄位
# 2. 處理缺失值(missing value)
# 3. 處理重複值(duplicate value)
# 3. 最後刪除掉不必要的欄列資料
#
# [回頂部](#成大通識與各系必修撞課分析)
# 先針對資料做更仔細的觀察,以下是重點關注的欄位
#
# 主要分析的欄位有5個:
# - **開課單位**:判斷是否屬於必修或通識
# - **星期**、**開始時間**、**結束時間**:判斷有無撞課
# - **課程類別**:依照不同領域的通識做更細部的分析
#
# 幫助我們篩選資料的欄位有2個:
# - **學制**:篩選出大學部的課程
# - **必選修**:只保留必修部分(各系必修和通識都算必修)
#
# 其餘欄位都不重要,可以捨棄
#
# > 備註:有些研究所會開課給大學部學生,例如大碩合開,因此以學制篩選資料,會看到開課單位還是有研究所
df.columns
# 檢查缺失值,並顯示有缺失值的欄位,這裡我們只在乎上面提到的重要欄位,恰好只有「開始時間」和「結束時間」需要處理
print(df.isnull().sum())
df[df.isnull()['開始時間']]
# 很幸運只有1列資料有缺失值,接著開始清理
print(f'before size: {df.shape}')
df.dropna(subset=['開始時間'], inplace=True)
print(f'after size: {df.shape}')
# 檢查重複值
repeated = df[df.duplicated(keep=False)]
print(f'重複 {len(repeated.index)} 列')
repeated.head(4)
# 清理重複值,重複的列中只保留一列,因此清除掉10列
print(f'before size: {df.shape}')
df.drop_duplicates(keep='first', inplace=True)
print(f'after size: {df.shape}')
# 檢查看看哪些欄位沒有意義可以刪除
for col in df.columns:
print(f'{col}\n{df[col].unique()}\n')
# 刪除不必要的欄位
# +
print(df.shape)
if '學制' in df.columns:
df = df[df['學制'] == '大學部']
if '必選修' in df.columns:
df = df[df['必選修'] == '必修']
df = df[['開課單位', '星期', '開始時間', '結束時間', '課程類別']]
print(df.shape)
df.columns
# -
df['開課單位'].unique()[0:10]
# 資料中存在不是通識中心也非各科系的開課單位,例如:體育室、華語中心、服務學習等
#
# 依據 [國立成功大學通識課程選修要點](http://cge.ncku.edu.tw/p/404-1007-7830.php?Lang=zh-tw#%E5%9C%8B%E7%AB%8B%E6%88%90%E5%8A%9F%E5%A4%A7%E5%AD%B8%E9%80%9A%E8%AD%98%E8%AA%B2%E7%A8%8B%E9%81%B8%E4%BF%AE%E8%A6%81%E9%BB%9E(108%E5%AD%B8%E5%B9%B4%E5%BA%A6%E8%B5%B7%E5%85%A5%E5%AD%B8%E7%94%9F%E9%81%A9%E7%94%A8)),通識課包含「基礎國文」、「外國語言」、「領域通識」及「融合通識」,對應到資料中的開課單位為**基礎國文**、**外國語言**及**通識中心**
#
# 因此其他非系所名稱也非通識的開課單位,都不是我們這次要討論的範圍,刪掉
print(df.shape)
for i in ['體育室', '教育學程', '服務學習(三)', '共同科目英語授課', '華語中心']:
df = df[df['開課單位'] != i]
print(df.shape)
# 為了方便,將上課時間做標籤編碼(label encoding),07:10-08:00 為第一節,08:10-09:00為 第二節,以此類推
# +
for i in range(15):
df.replace([f'{i+7:02d}:10', f'{i+8:02d}:00'], i, inplace=True)
print(df['開始時間'].unique())
print(df['結束時間'].unique())
# -
# 將時間未定的課程刪掉
print(df.shape)
df = df[df['開始時間'] != '未定']
df = df[df['結束時間'] != '未定']
print(df.shape)
# ## 視覺化呈現
# 我們想知道以下幾個問題:
# 1. 通識課的時間分布
# 2. 各系必修的時間分布
# 3. 各系必修課撞了多少通識?
#
# [回頂部](#成大通識與各系必修撞課分析)
# 拆分資料
gener = df[df['開課單位'].isin(['通識中心', '基礎國文', '外國語言'])] # general education
requi = df[~df['開課單位'].isin(['通識中心', '基礎國文', '外國語言'])] # required subject
print(gener.head(3), '\n')
print(requi.head(3))
type(df['開課單位'])
# +
def is_clashed(class1: pd.Series, class2: pd.Series) -> bool:
''' 判斷兩堂課是否衝堂 '''
class1_time = range(class1['開始時間'], class1['結束時間'] + 1)
return class2['開始時間'] in class1_time or class2['結束時間'] in class1_time
def clash(gen, req):
pass
requi.head()
requi.loc[661, :]
# is_clashed(requi[1, :], requi[1, :])
# gener.apply(is_clashed, args=(requi[1, :],))
# -
df['開課單位'].value_counts().head(10)
# ## 結果分析與討論
# [回頂部](#成大通識與各系必修撞課分析)
| main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Louvain Community Detection
#
#
# In this notebook, we will use cuGraph to identify the cluster in a test graph using the Louvain algorithm
#
# Notebook Credits
# * Original Authors: <NAME> and <NAME>
# * Last Edit: 04/30/2019
#
# RAPIDS Versions: 0.7.0
#
# Test Hardware
# * GP100 32G, CUDA 9,2
#
#
#
# ## Introduction
#
# The Louvain method of community detection is a greedy heirarical clsutering algorithm which seeks to optimize Modularity as it progresses. Louvain starts with each vertex in its own clusters and iteratively merges groups using graph contraction.
#
# For a detailed description of the algorithm see: https://en.wikipedia.org/wiki/Louvain_Modularity
#
# It takes as input a cugraph.Graph object and returns as output a
# cudf.Dataframe object with the id and assigned partition for each
# vertex as well as the final modularity score
#
#
# To compute the Louvain cluster in cuGraph use: <br>
#
# **nvLouvain(G)**
# * __G__: A cugraph.Graph object
#
# Returns:
#
# * tupal __lovain dataframe__ and __modularity__
#
#
# * __louvain__: cudf.DataFrame with two names columns:
# * louvain["vertex"]: The vertex id.
# * louvain["partition"]: The assigned partition.
#
# * __modularity__ : the overall modularity of the graph
#
# All vertices with the same partition ID are in the same cluster
#
#
# ## cuGraph 0.7 Notice
# cuGraph version 0.7 has some limitations:
# * Only Int32 Vertex ID are supported
# * Only float (FP32) edge data is supported
# * Vertex numbering is assumed to start at zero
#
# These limitations are being addressed and will be fixed future versions.
# These example notebooks will illustrate how to manipulate the data so that it comforms to the current limitations
#
# A new renumbering feature is being worked and will be reflected in updated notebooks for the next release.
#
# ## References
#
# * <NAME>., <NAME>., <NAME>., and <NAME>. Fast unfolding of communities in large networks. Journal of statistical mechanics: theory and experiment 2008, 10 (2008), P10008.
#
# ### Test Data
# We will be using the Zachary Karate club dataset
# *<NAME>, An information flow model for conflict and fission in small groups, Journal of
# Anthropological Research 33, 452-473 (1977).*
#
#
# 
#
# ### Prep
# Import needed libraries
import cugraph
import cudf
import numpy as np
from collections import OrderedDict
# ## Read data using cuDF
# +
# Test file
datafile='./data//karate-data.csv'
# Read the data file
cols = ["src", "dst"]
dtypes = OrderedDict([
("src", "int32"),
("dst", "int32")
])
gdf = cudf.read_csv(datafile, names=cols, delimiter='\t', dtype=list(dtypes.values()) )
# -
# Louvain is dependent on vertex ID starting at zero
gdf["src_0"] = gdf["src"] - 1
gdf["dst_0"] = gdf["dst"] - 1
# The algorithm also requires that there are vertex weights. Just use 1.0
gdf["data"] = 1.0
# just for fun, let's look at the data types in the dataframe
gdf.dtypes
# create a Graph
G = cugraph.Graph()
G.add_edge_list(gdf["src_0"], gdf["dst_0"], gdf["data"])
# Call Louvain on the graph
df, mod = cugraph.louvain(G)
# Print the modularity score
print('Modularity was {}'.format(mod))
print()
df.dtypes
# How many partitions where found
part_ids = df["partition"].unique()
print(str(len(part_ids)) + " partition detected")
for p in range(len(part_ids)):
part = []
for i in range(len(df)):
if (df['partition'][i] == p):
part.append(df['vertex'][i] +1)
print("Partition " + str(p) + ":")
print(part)
# ___
# Copyright (c) 2019, <NAME>.
#
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
# ___
| cugraph/Louvain.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: algotrading
# language: python
# name: algotrading
# ---
# +
# Instructions here: https://algotrading101.com/learn/alpaca-trading-api-guide/
import os
import pandas as pd
import datetime as dt
from pytz import timezone
import alpaca_trade_api as tradeapi
# authentication and connection details
# store your Alpaca API key and secret in your environment
api_key = os.environ.get('alpaca_api_key')
api_secret = os.environ.get('alpaca_api_secret')
base_url = 'https://api.alpaca.markets'
# for polygon connection
os.environ['APCA_API_KEY_ID']=api_key
os.environ['APCA_API_SECRET_KEY']=api_secret
# instantiate REST API
api = tradeapi.REST(api_key, api_secret, base_url, api_version='v2')
# obtain account information
account = api.get_account()
# +
# helper functions to pull data from polygon
def get_tick_data(symbol='SPY',date='2018-09-19',start='09:30:00',ticks=10000):
full_date = date+" "+start
st = dt.datetime.strptime(full_date, '%Y-%m-%d %H:%M:%S')
st = timezone('US/Eastern').localize(st)
st = int(st.timestamp())*1000
trades = tradeapi.REST().polygon.historic_trades_v2(symbol=symbol, date=date, limit=ticks)
trades.df.reset_index(level=0, inplace=True)
#convert exchange numeric codes to names for readability
exchanges = tradeapi.REST().polygon.exchanges()
ex_lst = [[e.id,e.name,e.type] for e in exchanges]
dfe = pd.DataFrame(ex_lst,columns=['exchange','exch','excode'])
trades.df['exchange'] = trades.df['exchange'].astype(int)
df = pd.merge(trades.df,dfe,how='left',on='exchange')
df = df[df.exchange!=0]
df.drop('exchange', axis=1, inplace=True)
return df
def get_quote_data(symbol='SPY',date='2018-09-19',start='09:30:00',ticks=10000,cond=False):
full_date = date+" "+start
st = dt.datetime.strptime(full_date, '%Y-%m-%d %H:%M:%S')
st = timezone('US/Eastern').localize(st)
st = int(st.timestamp())*1000
trades = tradeapi.REST().polygon.historic_quotes_v2(symbol=symbol, date=date, limit=ticks)
trades.df.reset_index(level=0, inplace=True)
return trades.df
# -
## Pull daily data for AAPL
aapl = api.get_barset('AAPL', 'day')
aapl = aapl.df
aapl.head()
## Pull 15min data for TSLA
tsla = api.get_barset('TSLA', '15Min')
tsla = tsla.df
tsla.head()
## Pull 1min data for MSFT
msft = api.get_barset('MSFT', '1Min')
msft = msft.df
msft.head()
## Pull last 1k bars of 1min data for TTD
ttd = api.get_barset(symbols='TTD', timeframe='1Min', limit=1000)
ttd = ttd.df
ttd.head()
## Pull last 1k bars of 1min data for GOOGL
googl = api.get_barset(symbols='GOOGL', timeframe='1Min', start='2020-01-15T09:30:00-04:00', limit=1000)
googl = googl.df
googl.head()
# get SPY tick data
get_tick_data(date='2018-08-01').head()
# get SPY quote data
get_quote_data(date='2018-08-01').head()
| Example_Notebooks/AlpacaAPI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
#PIDミュレーション Ver 0.1 (暫定版)
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
from tqdm.notebook import tqdm_notebook as tqdm
# +
"""
def rk4(func, t, h, x, *p)
4次のルンゲ・クッタ法を一回分計算する関数
引数リスト
func:導関数
t:現在時刻を表す変数
h:刻み幅
x:出力変数(求めたい値)
*p:引数の数が可変する事に対応する、その他の必要変数
※この関数では時刻は更新されないため、これとは別に時間更新をする必要があります。
導関数の書き方
def func(t, x, *state):
func:自分で好きな関数名をつけられます
t:時刻変数(変数の文字はtで無くても良い)
x:出力変数(変数の文字はxで無くても良い)
*state:その他の必要変数(引数の数は可変可能))
#関数サンプル
def vdot(t, x, *state):
s1=state[0]
s2=state[1]
return t+x+s1+s2
"""
def rk4(func, t, h, x, *p):
#print(x)
k1=h*func(t, x, *p)
k2=h*func(t+0.5*h, x+0.5*k1, *p)
k3=h*func(t+0.5*h, x+0.5*k2, *p)
k4=h*func(t+h, x+k3, *p)
x=x+(k1 + 2*k2 + 2*k3 + k4)/6
return x
# -
#位置型PID制御の実装例
class pid:
def __init__(self, kp, ti, td,limit_upper=100.0,limit_lower=-100.0,limitter=False):
self.Sum=0.0
self.olderr=0.0
self.oldtime=0.0
self.limit_upper= limit_upper
self.limit_lower= limit_lower
self.limitterFlag=True
self.Kp=kp
self.Ti=ti
self.Td=td
self.u=0.0
def controller(self, y, ref, t):
err=ref-y
period=t-self.oldtime
self.Sum=self.Sum+err*period
#print(period)
if period==0.0:
errdot=0.0
else:
errdot=(err-self.olderr)/period
self.u=self.Kp*(err + self.Sum/self.Ti + self.Td*errdot)
#リミッター
if self.limitterFlag==True:
if self.u>self.limit_upper:
self.u=self.limit_upper
elif self.u<self.limit_lower:
self.u=self.limit_lower
self.oldtime=t
self.olderr=err
return self.u
class vpid:
def __init__(self, kp, ti, td,limit_upper=100.0,limit_lower=-100.0,limitter=False):
self.Sum=0.0
self.olderr1=0.0
self.olderr2=0.0
self.oldtime=0.0
self.limit_upper= limit_upper
self.limit_lower= limit_lower
self.limitterFlag=True
self.Kp=kp
self.Ti=ti
self.Td=td
self.u=0.0
def controller(self, y, ref, t):
err=ref-y
period=t-self.oldtime
self.Sum=self.Sum+err*period
#errdot=(err-self.olderr)/period
if period==0.0:
self.u=self.u\
+ self.Kp*(err-self.olderr1 + period*err/self.Ti)
else:
self.u=self.u\
+ self.Kp*(err-self.olderr1 + period*err/self.Ti + self.Td*(err-2*self.olderr1+self.olderr2)/period)
#リミッター
if self.limitterFlag==True:
if self.u>self.limit_upper:
self.u=self.limit_upper
elif self.u<self.limit_lower:
self.u=self.limit_lower
self.oldtime=t
self.olderr2=self.olderr1
self.olderr1=err
return self.u
# +
def xdot(t,x,u):
#
# vdot = -2v -x +u
# xdot = v
#
vdot=-2*x[0]-x[1]+u
xdot=x[0]
return np.array([vdot,xdot])
fintime=3
h=1e-3
Kp= 66.8205
Ti= 0.4000
Td= 0.1383
pidcon=pid(Kp,Ti,Td)
vpidcon=vpid(Kp,Ti,Td)
x1=np.array([0.0, 0.0])
x2=np.array([0.0, 0.0])
r=1
t=0.0
X1=[]
V1=[]
X2=[]
V2=[]
T=[]
for _ in tqdm(range(int(fintime/h))):
V1.append(x1[0])
X1.append(x1[1])
V2.append(x2[0])
X2.append(x2[1])
T.append(t)
u1=pidcon.controller(x1[1],r,t)
u2=vpidcon.controller(x2[1],r,t)
x1=rk4(xdot,t,h,x1,u1)
x2=rk4(xdot,t,h,x2,u2)
t=t+h
V1.append(x1[0])
X1.append(x1[1])
V2.append(x2[0])
X2.append(x2[1])
T.append(t)
plt.figure(figsize=(7,2))
plt.plot(T, X1, lw=2, label='Position PID')
plt.plot(T, X2, lw=2, label='Velocity PID')
plt.grid()
plt.xlabel('Time(s)')
plt.ylabel('Output')
plt.legend()
plt.show()
# -
x[1]
# +
#マルチコプタ角度制御PID制御ゲイン調整(ピッチ角の例)
#https://github.com/kouhei1970/fundamental_of_multicopter_control
import numpy as np
import control.matlab as matlab
import matplotlib.pyplot as plt
#制御対象作成
K=1.0
wn=1.0
zeta=1.0
Plant=matlab.tf([K],[1,2*zeta*wn,wn**2])
print(Plant)
#①位相余裕𝑃𝑀とゲイン交差周波数𝜔_𝑐を決める
PM=60*np.pi/180
omega_c=10
#②位相余裕𝑃𝑀の時の位相 𝜙_𝑚を計算する
phi_m=PM-np.pi
#③制御対象の周波数伝達関数を求める
#pythonではz=1+2*1jの様に複素数が使えます
#角速度制御された角度制御の制御対象の伝達関数
#yaw
#1.86 s^2 + 166.1 s + 1661
#------------------------------------------
#0.0193 s^4 + 2.86 s^3 + 166.1 s^2 + 1661 s
#0.5442 s^2 + 97.17 s + 883.4
#--------------------------------------------
#0.0193 s^4 + 1.544 s^3 + 97.17 s^2 + 883.4 s
def freq_tf(w):
return (1)\
/((1j*w)**2 +2*(1j*w)+ 1.0)
Plant_freq=freq_tf(omega_c)
#④制御対象の周波数伝達関数からの𝜔_𝑐の時の実部𝑢の値と虚部𝑣の値を求める
u=Plant_freq.real
v=Plant_freq.imag
#⑤(f)式から比例ゲインを求める
Kp=(u*np.cos(phi_m)+v*np.sin(phi_m))/(u**2+v**2)
#⑥積分時間を適当に決めて(d)式から微分時間を求める
################################
Tis=np.array([0.4, 0.4, 0.4])
################################
Tds=1/omega_c**2/Tis - (v-u*np.tan(phi_m))/omega_c/(u+v*np.tan(phi_m))
#Tds=np.array([10,20,30])
#⑦求めたゲインで周波数応答、ステップ応答、インパルス応答、外乱からの応答等を見る。良くなければ⑥にもどる。良ければ終了
Kps=np.ones(len(Tis))*Kp
print('積分時間はこの範囲で設定してください')
if (v-u*np.tan(phi_m))/omega_c/(u+v*np.tan(phi_m))>0:
print('0 < Ti < {:f}\n'.format((u+v*np.tan(phi_m))/omega_c/(v-u*np.tan(phi_m))))
else:
print('Ti>0\n')
fig1=plt.figure(figsize=(10,7))
fig2=plt.figure(figsize=(20,3.5))
ax1=fig1.add_subplot(2,1,1)
ax2=fig1.add_subplot(2,1,2)
ax3=fig2.add_subplot(1,3,1)
ax4=fig2.add_subplot(1,3,2)
ax5=fig2.add_subplot(1,3,3)
for Kp,Ti,Td in zip(Kps, Tis, Tds):
Controller=matlab.tf([Kp*Td, Kp, Kp/Ti],[1,0])
Loop=Plant*Controller
closesys=matlab.feedback(Loop)
#位相余裕(ゲイン余裕)
gm, pm, wpc, wgc=matlab.margin(Loop)
print('Kp={:8.4f}'.format(Kp))
print('Ti={:8.4f}'.format(Ti))
print('Td={:8.4f}'.format(Td))
print('Gain crossover frequency={:6.2f}rad/s'.format(wgc))
print('Phase margin={:6.2f}deg\n'.format(pm))
wfreq=np.logspace(-1,4,1000)
#開ループボード線図
magq,phaseq,omegaq = matlab.bode(Loop,wfreq,plot=False)
ax1.semilogx(omegaq,20*np.log10(magq), lw=3,label='Ti={:8.4f}'.format(Ti))
#ax1.semilogx(wgc,20*np.log10(1),marker='o')
ax2.semilogx(omegaq,phaseq*180/np.pi,lw=3,label='Ti={:8.4f}'.format(Ti))
#ax2.semilogx(wgc,pm-180,marker='o')
#ステップ応答
t=np.linspace(0,10,1000)
y,t=matlab.step(closesys, t)
ax3.plot(t,y,lw=2,label='Ti={:5.3f}deg'.format(Ti))
#インパルス応答
#y,t=matlab.impulse(closesys, t)
ax4.plot(t,y,lw=2,label='Ti={:5.3f}deg'.format(Ti))
###外乱応答
t=np.linspace(0,10,1000)
sys_dist=Plant/(1+Plant*Controller)#sysq_pid/(1+cont_ang*sysq_pid)
#外乱ステップ応答
#y,t=matlab.step(sys_dist, t)
ax5.plot(t,y,lw=2,label='Ti={:5.3f}deg'.format(Ti))
#plt.rcParams["font.size"] = 12
bodefontsize=16
timeresfontsize=16
ax1.grid(which="both",ls=':')
ax1.set_yticks(np.arange(-80,81,40))
ax1.set_ylabel('Gain[dB]', fontsize=bodefontsize)
ax1.tick_params(axis='x', labelsize=bodefontsize)
ax1.tick_params(axis='y', labelsize=bodefontsize)
ax2.grid(which="both",ls=':')
#ax2.set_ylim(-200,-70)
ax2.set_yticks([-180-45,-180,-135,-90,-45])
ax2.set_ylabel('Phase[deg]', fontsize=bodefontsize)
ax2.set_xlabel('$\omega$[rad/s]', fontsize=bodefontsize)
ax2.tick_params(axis='x', labelsize=bodefontsize)
ax2.tick_params(axis='y', labelsize=bodefontsize)
ax3.grid()
ax3.tick_params(axis='x', labelsize=timeresfontsize)
ax3.tick_params(axis='y', labelsize=timeresfontsize)
ax3.set_xlabel('Time[s]',fontsize=timeresfontsize)
ax3.set_ylabel('Output',fontsize=timeresfontsize)
ax3.set_title('Step Responce',fontsize=timeresfontsize)
ax4.grid()
ax4.tick_params(axis='x', labelsize=timeresfontsize)
ax4.tick_params(axis='y', labelsize=timeresfontsize)
ax4.set_xlabel('Time[s]',fontsize=timeresfontsize)
ax4.set_title('Impulse Responce',fontsize=timeresfontsize)
ax5.grid()
ax5.tick_params(axis='x', labelsize=timeresfontsize)
ax5.tick_params(axis='y', labelsize=timeresfontsize)
ax5.set_xlabel('Time[s]',fontsize=timeresfontsize)
ax5.set_title('Disturbance Step Responce',fontsize=timeresfontsize)
ax1.legend(bbox_to_anchor=(1.0, 1), loc='upper left')
ax3.legend()
ax4.legend()
ax5.legend()
plt.show()
# -
plt.plot(t,y)
plt.plot(T,X)
plt.grid()
plt.show()
| sandbox/PID_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""import bibliotek"""
from gensim.models import Word2Vec
import numpy as np
import pandas as pd
from sklearn.decomposition import PCA
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib notebook
# -
# ## Dane
# +
#przykałdowe dane
sentences = []
for i in range(10):
sentence = list (range(10))
sentences.append(sentence)
# -
sentences
np.random.randint(10) #ganerator liczb losowych
# +
sentences = []
for i in range(1000000):
start = np.random.randint(0,20)
finish = start + np.random.randint(1,15)
sentence = [str(x) for x in list (range(start,finish))] #rzutowanie na stringa
sentences.append(sentence)
# -
sentences
# ## Model Word2Vec
model = Word2Vec (sentences , size = 10, window = 5, min_count = 1 )
#sentences - zdania,
#size -rozmiar wektora,
#window - szerokosc okno do kontekstu, min wartosc to slowa sa podobne, wieksza wartosc słowa
#min_count ile razy musi byc slowo aby było brane pod uwage
model.wv['1'] # oczyt pierwszego wektora
plt.figure(figsize = (15,8))
sns.heatmap(model.wv [model.wv.vocab],linewidths= 0.5);
"""funckja pomocnicza"""
def plot_heatmap(model):
plt.figure(figsize = (15,8))
sns.heatmap(model.wv [model.wv.vocab],linewidths= 0.5);
plot_heatmap (Word2Vec (sentences , size = 50, window = 5 ) )
# ## Wizualizacja - PCA
X = model.wv[model.wv.vocab]
#X.shape = 10 wymiarow
pca_model = PCA(n_components=2) #redukcja do 2 wymiarów
result = pca_model.fit_transform(X)
result [:,0] #wyciaganie danych
result [:,1] #wyciaganie danych
# +
plt.scatter (result [:,0],result [:,1]);
words = list (model.wv.vocab)#pokazuje wartosci przy pkt
for i, word in enumerate (words):
plt.annotate(word, xy =( result [i, 0],result [i, 1]))
# -
"""funckja pomocnicza"""
def plt_pca(model):
X = model.wv[model.wv.vocab]
pca_model = PCA(n_components=2) #redukcja do 2 wymiarów
result = pca_model.fit_transform(X)
plt.scatter (result [:,0],result [:,1]);
words = list (model.wv.vocab)
for i, word in enumerate (words):
plt.annotate(word, xy =( result [i, 0],result [i, 1]))
plt_pca(Word2Vec (sentences , size = 100, window = 10 ))
# ## Podobne słowa
plt_pca(Word2Vec (sentences , size = 300, window = 3 ))
model.wv.most_similar('20', topn = 3)
# 20 slowo
#topn=3 3 najpbardziiej podobne słowa do 20
#wykryl 19,18 i 23
model.wv.most_similar(positive=['10','8'], negative= ['2'],topn = 3)
| part4/day2/day2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !python -m pip install --user --upgrade pip
# !pip install --user --upgrade pandas matplotlib numpy scikit-learn tensorflow keras
# !pip3 install kfp --upgrade --user
#Check if the install was successful
# !which dsl-compile
# Import Kubeflow SDK
import kfp
import kfp.dsl as dsl
import kfp.components as comp
# +
import os
# where the outputs are stored
out_dir = str(os.getcwd())
model_file = 'modela'
holdout = True
# -
os.getcwd()
def train_tranform(data_path):
import pickle
import sys, subprocess;
subprocess.run([sys.executable, '-m', 'pip', 'install', 'pandas'])
subprocess.run([sys.executable, '-m', 'pip', 'install', 'scikit-learn'])
subprocess.run([sys.executable, '-m', 'pip', 'install', 'ipython'])
import numpy as np
import pandas as pd
from sklearn.preprocessing import PowerTransformer
# Download the dataset and split into training and test data.
data = pd.read_csv("https://raw.githubusercontent.com/Josepholaidepetro/Volve_ML/main/data/train.csv")
# Preprocess
data.drop(['DEPTH', 'BS', 'RD', 'ROP', 'RM', 'DRHO'], axis=1, inplace=True)
# If Nan, drop
data.dropna(inplace=True)
# transform the RT to logarithmic
data['RT'] = np.log10(data['RT'])
# perform a yeo-johnson transform of the train dataset
ptrain = PowerTransformer(method='yeo-johnson')
train_df_yj = ptrain.fit_transform(data.drop('DT', axis=1))
train_df_yj_norm = pd.DataFrame(train_df_yj, columns=data.columns.drop('DT'))
y_train = data['DT']
#Save the train_data as a pickle file to be used by the predict component.
with open(f'{data_path}/train_tranform_data', 'wb') as f:
pickle.dump((train_df_yj_norm, y_train), f)
train_tranform(out_dir)
def test_tranform(data_path):
import pickle
import sys, subprocess;
subprocess.run([sys.executable, '-m', 'pip', 'install', 'pandas'])
subprocess.run([sys.executable, '-m', 'pip', 'install', 'scikit-learn'])
subprocess.run([sys.executable, '-m', 'pip', 'install', 'ipython'])
import numpy as np
import pandas as pd
from sklearn.preprocessing import PowerTransformer
# Download the dataset and split into training and test data.
data = pd.read_csv("https://raw.githubusercontent.com/Josepholaidepetro/Volve_ML/main/data/test.csv")
# Preprocess
data.drop(['DEPTH', 'BS', 'ROP', 'DRHO'], axis=1, inplace=True)
# If Nan, drop
data.dropna(inplace=True)
# transform the RT to logarithmic
data['RT'] = np.log10(data['RT'])
# perform a yeo-johnson transform of the train dataset
ptest = PowerTransformer(method='yeo-johnson')
test_df_yj = ptest.fit_transform(data.drop('DT', axis=1))
test_df_yj_norm = pd.DataFrame(test_df_yj, columns=data.columns.drop('DT'))
y_test = data['DT']
#Save the test_data as a pickle file to be used by the predict component.
with open(f'{data_path}/test_data', 'wb') as f:
pickle.dump((test_df_yj_norm, y_test), f)
test_tranform(out_dir)
import os
print(os.listdir())
def Outlier_removal(data_path):
import pickle
import sys, subprocess;
subprocess.run([sys.executable, '-m', 'pip', 'install', 'pandas'])
subprocess.run([sys.executable, '-m', 'pip', 'install', 'scikit-learn'])
subprocess.run([sys.executable, '-m', 'pip', 'install', 'ipython'])
from sklearn.svm import OneClassSVM
import numpy as np
import pandas as pd
# Load and unpack the train_data
with open(f'{data_path}/train_tranform_data','rb') as f:
train_data = pickle.load(f)
# Separate the train_df_yj_norm from y_train.
train_df_yj_norm, y_train = train_data
# Method 4: One-class SVM
svm = OneClassSVM(nu=0.13)
yhat = svm.fit_predict(train_df_yj_norm)
mask = yhat != -1
train_df_svm = train_df_yj_norm[mask]
y_train_svm = y_train[mask]
# prepare train data for modelling
X_train = train_df_svm.copy().drop(['label'], axis=1)
y_train = y_train_svm.copy()
#Save the train_data as a pickle file to be used by the predict component.
with open(f'{data_path}/Outlier_removal_train_data', 'wb') as f:
pickle.dump((X_train, y_train), f)
#Save the train_data to be used for splitting as a pickle file to be used by the predict component.
with open(f'{data_path}/Outlier_removal_train_valid', 'wb') as f:
pickle.dump((train_df_svm.copy(), y_train_svm.copy()), f)
Outlier_removal(out_dir)
def valid_tranform(data_path):
import pickle
import sys, subprocess;
subprocess.run([sys.executable, '-m', 'pip', 'install', 'pandas'])
subprocess.run([sys.executable, '-m', 'pip', 'install', 'scikit-learn'])
subprocess.run([sys.executable, '-m', 'pip', 'install', 'ipython'])
import numpy as np
import pandas as pd
from sklearn.preprocessing import PowerTransformer
# Load and unpack the full Outlier_removal_train_valid
with open(f'{data_path}/Outlier_removal_train_valid','rb') as f:
X_train, y_train= pickle.load(f)
train_split = pd.concat([X_train.reset_index(drop=True), y_train.reset_index(drop=True)], axis=1)
# Separate the data into X_train and y_train.
X_train2 = train_split[train_split['label'] < 0].drop(['label', 'DT'], axis=1)
y_train2 = train_split[train_split['label'] < 0]['DT']
# Separate the data into X_valid and y_valid.
X_valid = train_split[train_split['label'] > 0].drop(['label', 'DT'], axis=1)
y_valid = train_split[train_split['label'] > 0]['DT']
#Save the valid_data as a pickle file to be used by the predict component.
with open(f'{data_path}/valid_data', 'wb') as f:
pickle.dump((X_valid, y_valid), f)
#Save the train_data as a pickle file to be used by the predict component.
with open(f'{data_path}/valid_tranform_train_data', 'wb') as f:
pickle.dump((X_train2, y_train2), f)
valid_tranform(out_dir)
def model_building(data_path, model_file, holdout):
import pickle
import sys, subprocess;
subprocess.run([sys.executable, '-m', 'pip', 'install', 'pandas'])
subprocess.run([sys.executable, '-m', 'pip', 'install', 'scikit-learn'])
subprocess.run([sys.executable, '-m', 'pip', 'install', 'ipython'])
import numpy as np
import pandas as pd
import tensorflow as tf
from tensorflow import keras
# Load and unpack the full Outlier_removal_train_data
with open(f'{data_path}/Outlier_removal_train_data','rb') as f:
Outlier_removal_train_data = pickle.load(f)
# Separate the X_train from y_train.
X_train, y_train = Outlier_removal_train_data
# Load and unpack the valid_tranform_train_data
with open(f'{data_path}/valid_tranform_train_data','rb') as f:
valid_tranform_train_data = pickle.load(f)
# Separate the X_train from y_train.
X_train_val, y_train_val = valid_tranform_train_data
# Load and unpack the valid_data
with open(f'{data_path}/valid_data','rb') as f:
valid_data = pickle.load(f)
# Separate the train_df_yj_norm from y_train.
X_valid, y_valid = valid_data
# Load and unpack the test_data
with open(f'{data_path}/test_data','rb') as f:
test_data = pickle.load(f)
# Separate the train_df_yj_norm from y_train.
X_test, y_test = test_data
# Define the model using Keras
tf.random.set_seed(1)
model = keras.models.Sequential()
model.add(keras.layers.Dense(units =128, activation='relu', input_dim=X_train.shape[1]))
model.add(keras.layers.Dense(units =128, activation='relu'))
model.add(keras.layers.Dense(units =256, activation='relu'))
model.add(keras.layers.Dense(units =256, activation='relu'))
model.add(keras.layers.Dense(units =1, activation='linear'))
model.summary()
model.compile(optimizer = 'adam', loss='mean_squared_error')
if holdout is False:
# Run a training job
model.fit(X_train, y_train, batch_size=20 , epochs=5 )
else:
# Run a training job
model.fit(X_train_val, y_train_val, batch_size=20 , epochs=20, validation_data=(X_valid, y_valid))
model.save(f'{data_path}/{model_file}')
#Save holdout
with open(f'{data_path}/holdout', 'wb') as f:
pickle.dump((holdout), f)
regressor = model_building(out_dir, model_file, holdout)
def model_evaluate(data_path, model_file, holdout):
import pickle
import sys, subprocess;
subprocess.run([sys.executable, '-m', 'pip', 'install', 'pandas'])
subprocess.run([sys.executable, '-m', 'pip', 'install', 'scikit-learn'])
subprocess.run([sys.executable, '-m', 'pip', 'install', 'ipython'])
import numpy as np
import pandas as pd
import tensorflow as tf
from tensorflow import keras
# Load the saved model
model = keras.models.load_model(f'{data_path}/{model_file}')
# Load and unpack the test_data
with open(f'{data_path}/test_data','rb') as f:
test_data = pickle.load(f)
# Separate the X_test from y_test.
X_test, y_test = test_data
# Load and unpack the valid_data
with open(f'{data_path}/valid_data','rb') as f:
valid_data = pickle.load(f)
# Separate the train_df_yj_norm from y_train.
X_valid, y_valid = valid_data
if holdout is False:
#Evaluate the model and print the results
score = model.evaluate(X_test, y_test, verbose=0)
print("mse of Well 3: {}".format(score))
else:
score = model.evaluate(X_valid,y_valid, verbose=0)
print("mse of Well 3: {}".format(score))
model_evaluate(out_dir, model_file, holdout)
def predict(data_path, model_file):
import pickle
import sys, subprocess;
subprocess.run([sys.executable, '-m', 'pip', 'install', 'pandas'])
subprocess.run([sys.executable, '-m', 'pip', 'install', 'scikit-learn'])
import numpy as np
import pandas as pd
import tensorflow as tf
from tensorflow import keras
# Load the saved model
regressor = keras.models.load_model(f'{data_path}/{model_file}')
# Load and unpack the test_data
with open(f'{data_path}/test_data','rb') as f:
test_data = pickle.load(f)
# Separate the X_test from y_test.
test_df_yj_norm, y_test = test_data
# make predictions.
y_pred = regressor.predict(test_df_yj_norm)
with open(f'{data_path}/result1', 'wb') as f:
pickle.dump(y_pred, f)
with open(f'{data_path}/result.txt', 'w') as result:
result.write(" Prediction: {}, Actual: {} ".format(y_pred,y_test))
print('Prediction has be saved successfully!')
# Create train and predict lightweight components.
train_tranform_op = comp.func_to_container_op(train_tranform , base_image = "tensorflow/tensorflow:latest-gpu-py3")
test_tranform_op = comp.func_to_container_op(test_tranform , base_image = "tensorflow/tensorflow:latest-gpu-py3")
Outlier_removal_op = comp.func_to_container_op(Outlier_removal , base_image = "tensorflow/tensorflow:latest-gpu-py3")
valid_tranform_op = comp.func_to_container_op(valid_tranform , base_image = "tensorflow/tensorflow:latest-gpu-py3")
train_op = comp.func_to_container_op(model_building , base_image = "tensorflow/tensorflow:latest-gpu-py3")
valid_op = comp.func_to_container_op(model_evaluate , base_image = "tensorflow/tensorflow:latest-gpu-py3")
predict_op = comp.func_to_container_op(predict , base_image = "tensorflow/tensorflow:latest-gpu-py3")
# ## Build Kubeflow Pipeline
#Create a client to enable communication with the Pipelines API server.
client = kfp.Client()
# +
# Define the pipeline
@dsl.pipeline(
name='Sonic Log Prediction Pipeline',
description='An ML pipeline that performs Sonic Log Prediction prediction.'
)
# Define parameters to be fed into pipeline
def sonic_container_pipeline(
data_path: str,
holdout: bool,
model_file: str
):
# Define volume to share data between components.
vop = dsl.VolumeOp(
name="create_volume",
resource_name="data-volume",
size="1Gi",
modes=dsl.VOLUME_MODE_RWO) # read write one
# Create deploy component.
train_tranform_container = train_tranform_op(data_path) \
.add_pvolumes({data_path: vop.volume})
# Create data transformation component.
test_tranform_container = test_tranform_op(data_path) \
.add_pvolumes({data_path: train_tranform_container.pvolume})
# Create data transformation component.
Outlier_removal_container = Outlier_removal_op(data_path) \
.add_pvolumes({data_path: test_tranform_container.pvolume})
# Create data transformation component.
valid_tranform_container = valid_tranform_op(data_path) \
.add_pvolumes({data_path: Outlier_removal_container.pvolume})
# Create model training component.
train_container = train_op(data_path, model_file, holdout) \
.add_pvolumes({data_path: valid_tranform_container.pvolume})
# Create model validation component.
valid_container = valid_op(data_path, model_file, holdout) \
.add_pvolumes({data_path: train_container.pvolume})
# Create model training component.
predict_container = predict_op(data_path, model_file) \
.add_pvolumes({data_path: valid_container.pvolume})
# Print the result of the prediction
churn_result_container = dsl.ContainerOp(
name="print_prediction",
image='library/bash:4.4.23', # Image tag for the Docker container to be used.
pvolumes={data_path: predict_container.pvolume}, # the name displayed for the component execution during runtime.
arguments=['cat', f'{data_path}/result.txt']
)
# -
# ## Run the Pipeline
pipeline_func = sonic_container_pipeline
DATA_PATH = out_dir # mount your filesystems or directory
MODEL_PATH = 'model_file'
# +
experiment_name = 'dt_prediction_kubeflow'
run_name = pipeline_func.__name__ + ' run'
arguments = {"data_path":DATA_PATH,
"holdout":holdout,
"model_file":MODEL_PATH}
# Compile pipeline to generate compressed YAML definition of the pipeline.
kfp.compiler.Compiler().compile(pipeline_func, '{}.zip'.format(experiment_name))
# Submit pipeline directly from pipeline function
run_result = client.create_run_from_pipeline_func(pipeline_func,
experiment_name=experiment_name,
run_name=run_name,
arguments=arguments)
# -
# http://localhost:8888/pipeline#/experiments/details/644a33c5-3114-4b3c-b6de-3b0b6467eafa
#
# http://localhost:8888/pipeline#/runs/details/2aadb518-ead4-4035-90ea-011a44bfe16d
| maven/maven_new_kbflow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h2> ======================================================</h2>
# <h1>MA477 - Theory and Applications of Data Science</h1>
# <h1>Lesson 4: Vizualising Data with Matplotlib</h1>
#
# <h4>Dr. <NAME></h4>
# United States Military Academy, West Point
# AY20-2
# <h2>======================================================</h2>
# Matplotlib is one of the (most popular) plotting libraries in Python. If you have experience with MatLab's graphical plotting tools, then you'll realize that Matplotlib is very simmilar to MatLab in many aspects.
#
# Matplotlib is designed in the object oriented spirit; that is, it gives the user tremendous control over every aspect of the figure, and as such one is able to create highly customized graphs.
#
# Matplotlib is compatible with NumPy arrays and Pandas, and as such it is a great tool to have under your belt for some quick exploratory visual analysis of your data.
#
# If you don't already have Matplotlib installed in your computer, then you may install it either from your command line using the `pip` install command or from the anaconda terminal using the `conda` command:
# <center>
#
# ```python
# pip install matplotlib
#
# conda install matplotlib
# ```
# </center>
#
# For a more detailed and complete tutorial of Matplotlib you may visit the official website by clicking on the: <a href='https://matplotlib.org/index.html'>Matplotlib Website</a>
#
# For quick and basic figures we will often use the `pyplot` collection of commands and functions within `matplotlib`. So let's go ahead and import them:
import matplotlib.pyplot as plt
# If the figures do not automatically display within the JupyterNotebook, then go ahead and add the following:
# %matplotlib inline
# You may also include `plt.show()` at the end of each plot for it to display.
#
# Next, let's go ahead and import Numpy as well:
import numpy as np
import pandas as pd
# <h2> Basic Examples</h2>
#
# In Matplotlib, generally speaking, there are two ways of making a plot: a <b>functional</b> and <b> object-oriented</b> way.
#
# The functional way may be preferred if you want to quickly create a figure, however, if you want more control over your figure then the object-oriented approach is the way to go, as it gives you control over essentially every aspect of the graph.
#
# We will briefly discuss both.
#
# First, let's go and create a function that we want to plot:
#
#
#Create a Numpy array with 30 entries of equally separated values between -1 and 3, and define a simple cubic
#function
x=np.linspace(-3,3,30)
y=x**3
x
y
# <h2> Functional Method</h2>
#
# If we want to quickly plot this cubic function then we may use the `.plot()` method:
plt.plot(x,x**3)
plt.show()
# One can further customize this plot, such as changing the color and style of the curve. For example, one can specify the line color, the type of line (solid, dashed, markers etc.) the line width, marker size et. `plot(x, y, color='??', marker='??', linestyle='??',linewidth=??, markersize=??)`
#
#
plt.plot(x, y,color='r', ls='-.',linewidth=5, marker='+',markerfacecolor='black',
markersize=12)
plt.show()
# We can also add `xlabels, ylabels, title` to the plot:
plt.plot(x, y,'g--')
plt.xlabel('x-axis',fontsize=18)
plt.ylabel('y-axis',fontsize=12)
plt.title('Cubic Plot',fontsize=22)
# We can also control the size of the plot:
# +
plt.figure(figsize=(6,4))
plt.plot(x, y,'g--')
plt.xlabel('x-axis',fontsize=12)
plt.ylabel('y-axis',fontsize=12)
plt.title('Cubic Plot',fontsize=14)
# -
# <h3> Plotting multiple plots on the same canvas</h3>
#
# Matplotlib allows us to plot multiple plots on the same canvas via the `.subplot(nrows,ncolumns,index)` method. We will talk about this in a lot more detail shortly once we start discussing the object oriented approach.
#
#
# +
plt.figure(figsize=(12,4))
plt.subplot(2,3,1)
plt.plot(x,y,'g:')
plt.subplot(2,3,2)
plt.plot(x,x**2,'r-.')
plt.xlabel('x-axis')
plt.title("Second Plot")
plt.subplot(2,3,3)
plt.plot(x,np.sin(x),'y--')
plt.subplot(2,3,4)
plt.plot(x,x+2,'b-.')
plt.show()
# -
# <font color='red' size='4'>Take a minute or two and experiment with the different parameters.</font>
# <h2> Matplotlib via Object Oriented API (Application Programming Interface) Method</h2>
#
# In the OO API method, we start by instantiating a figure object and then call methods off it.
#
# We'll demonstrate this step-by step below:
#
# <ul>
# <li>First let us start by creating a figure object via <b>plt.figure()</b>. We may think of this as a blank canvas...it is the enviroment where we will place everything else.</li>
# <br>
# <li>Once the blank canvas is created, we can proceed to adding a set of axis. We need to define four parameters that control the percentage of the canvas you want to use. Namely, where you want to place the left axis, the bottom axis, and what percentage of the widht and height you want to use.</li>
# <br>
# <li> After creating a set of axis we can go ahead and plot in much the same way as we did before</li>
# <br>
# <li> You can customize the plot to your liking. For example, you can set x-labels, y-labels, title, change the color of the line, the styling etc. etc. When adding the labels, just remember that since this is an OO approach, now you need to add the label to the axis you created.</li>
# <br>
# <li> You can also specify the size of the canvas `figsize` as well as dots per inch `dpi` which controlls the quality of the image</li>
# </ul>
# +
fig=plt.figure(figsize=(4,6))
axes=fig.add_axes([0.1,0.1,0.9,0.9])
axes.plot(x,y,'r--')
axes.set_xlabel('x-axis',fontsize=12)
axes.set_ylabel('y-axis',fontsize=12)
axes.set_title('Plot of Cubic Using OO Method',fontsize=14)
# -
# To see the power of the OO approach, let's put two sets of figures in a single canvas and place them wherever we like.
# +
fig=plt.figure(figsize=(6,4))
axes1=fig.add_axes([0.1,0.1,1,1])
axes2=fig.add_axes([0.2,0.62,0.4,0.3])
axes1.plot(x,y,'r--')
axes2.plot(x,np.cos(2*x),'g-.')
axes1.set_title('Cubic Plot')
axes2.set_title('Cosine Plot')
# -
# <font color='red' size='5'>Exercise</font>
# <br>
#
# Create another figure and place it at the lower-right corner of the canvas. Plot any function/graph you want. Set the x and y labels and titles separately for all three figures.
# <h2> Subplots via OO Method</h2>
# +
fig, axes=plt.subplots(nrows=2,ncols=3,figsize=(8,4))
#The method .tight_layout() ensures there is no overlap between the different axes
axes[0,0].plot(x,y)
axes[1,1].plot(x,np.exp(-x))
axes[1,1,].set_title("Plot 1,1")
axes[0,1].set_xlabel('Label for plot 0,1')
# axes[2].plot(x,np.cos(x))
# axes[0].scatter(x,y)
# for ax in axes[0]:
# ax.plot(x,y)
# for ax in axes[1]:
# ax.plot(x,np.cos(3*x))
plt.tight_layout()
# -
# Two things to note:
#
# <ul>
# <li>First, unlike previously, when calling <i>subplots()</i> the axis are automatically created in the canvas. </li>
#
# <li>Second, unlike before, the <i>axes</i> object created when using <i>subplots()</i>, is actually a list of matplotlib axes (similar to what we had to manually create before). That's why we can actually iterate through it and decide what to plot on each individual axis. </li>
# </ul>
#
# Let's check it by calling `axes`:
axes[0,0]
# Instead of iterating through the `axes` we can also index them and plot:
# +
fig, axes=plt.subplots(nrows=2,ncols=3, figsize=(10,6))
#The method .tight_layout() ensures there is no overlap between the different axes
axes[0,0].plot(x,y)
axes[0,0].set_title('Plot 0,0')
axes[1,0].plot(x,0.1*y**2,'r--')
axes[1,1].plot(x,np.cos(2*x),'g-')
axes[1,1].set_title('Plot 1 1')
axes[1,1].set_xlabel('x-axis')
axes[0,1].hist(np.random.randn(500),bins=10)
axes[0,1].set_title('Histogram')
axes[0,2].plot(x,np.exp(-x),'y:')
axes[1,2].scatter(x,np.random.randn(30),s=12,color='r')
axes[1,2].set_title('Scatter Plot')
plt.tight_layout()
# -
# <font color='red' size='5'> Exercise</font>
#
# Create a one by three subplot and graph a few different functions. Set the titles and x and y labels for all of them.
# <h2>Saving Figures</h2>
#
# We can save figures in high quality in any formats `.png, .jpg, .pgf` etc. via the method `.savefig()` method.
# +
fig, axes=plt.subplots(nrows=2,ncols=3, figsize=(10,6))
#The method .tight_layout() ensures there is no overlap between the different axes
axes[0,0].plot(x,y)
axes[0,0].set_title('Plot 0,0')
axes[1,0].plot(x,0.1*y**2,'r--')
axes[1,1].plot(x,np.cos(2*x),'g-')
axes[1,1].set_title('Plot 1 1')
axes[1,1].set_xlabel('x-axis')
axes[0,1].hist(np.random.randn(500),bins=10)
axes[0,1].set_title('Histogram')
axes[0,2].plot(x,np.exp(-x),'y:')
axes[1,2].scatter(x,np.random.randn(30),s=12,color='r')
axes[1,2].set_title('Scatter Plot')
plt.tight_layout()
plt.savefig('My_first_picture.png',dpi=100)
# -
# <h2>Legends</h2>
#
# If we are plotting multiple plots on the same canvas, then it is important to be able to distinguish between the plots. So, it becomes important to add a legend to the plots.
# +
fig=plt.figure(figsize=(8,6))
axes1=fig.add_axes([0.1,0.1,0.9,0.9])
axes2=fig.add_axes([0.15,0.6,0.55,0.33])
axes1.plot(x,x**3,'r-',label='Cubic',lw=3,alpha=0.5,marker='o',markerfacecolor='b',
markersize=12,markeredgecolor='#000000',mew=3)
axes1.plot(x,x**2,'b:',label='Curve 1')
axes1.set_title('Big Plot')
#Setting the Legend for the larger figure
axes1.legend(loc=4)
axes2.plot(x,np.cos(2*x),color='g',linestyle='-.',label='Cos')
axes2.plot(x,np.sin(x),'r--',label='Sin')
axes2.set_title('Small Plot')
#Setting the legend for the smaller figure
axes2.legend(loc='upper left')
# -
# <font color='red' size='5'>Exercise</font>
#
# Create a new plot and experiment with all of the following:
# <ul>
# <li> plot at least two different graphs on the same canvas</li>
# <li>create a legend and experiment with a few different locations you can place it on a canvas</li>
# <li> line styling: line colors, line widths, line shapes/style, transparency etc.</li>
# <li> markers: different shapes, colors, sizes, edgewidths etc.</li>
# </ul>
# <h2> Axes Appearance and Plot Range</h2>
#
# We can set the range in both the x-axis and y-axis that we want to display by using the methods `.xlim(), .ylim()`
#
# Matplotlib also gives us the ability to rename the x-ticks or y-ticks to whatever labels we desire.
#
# Matplotlib has also a built-in method that allows us to plot text, called `.text()`
# +
fig=plt.figure(figsize=(8,5))
ax=fig.add_axes([0,0,1,1])
ax.plot(x,np.sin(x), lw=4,ls='-.')
#Plotting the text 'MA477' at position (-1,0.5)
ax.text(-1,0.5,'MA477',size=18)
#Setting the range for the x-axis
#plt.xlim([-1,1])
#Setting the range for the y-axis
#plt.ylim(-0.1,0.7)
#Setting customized labels along the x-axis at the specified locations
plt.xticks(ticks=[-2,-1,0,1,2],labels=['a','b','c','d','e'],fontsize=16)
#Setting customized labels along the y-axis at the specified locations
#plt.yticks(ticks=[-1,0,1],labels=['Low','Mid','High'],fontsize=18)
plt.show()
# -
# <font color='red' size='5'> Exercise</font>
# <br>
#
# Produce a plot that roughly looks like the one below:
# +
#Enter Code Here
fig=plt.figure(figsize=(8,5))
ax=fig.add_axes([0,0,1,1])
ax.plot(x,np.sin(-x),'r--',lw=4,label='Sin(-x)')
ax.plot(x,np.sin(x),'b-.',lw=3,label='Sin(x)', marker='+', ms=12, markeredgecolor='#000000', mew=3)
for item, letter in zip(x[::3],['MA'+str(i) for i in range(10)]):
ax.text(item+0.2,np.sin(-item)+0.05,letter,fontsize=12)
ax.text(item+0.25,np.sin(item)+0.07, letter[0]+'-'+letter[1]+'-'+letter[-1], fontsize=14)
ax.set_title("Confusing Plot", fontsize=18)
ax.set_ylabel('Y - axis',fontsize=14)
plt.xticks(np.linspace(-3,3,8),['s','i','n','e','w','a','v','e'],fontsize=14)
plt.legend(loc='center right')
# -
# <h2> Special Plots</h2>
#
# Matplotlib has many special plots which are often used for statistical analysis. We will learn later about a new library called Seaborn, which is built on top of Matplotlib, and has many more specialized plots that are often used when trying to visualize our data.
#
# Let's go ahead and create a dataframe with synthetic data:
#
data=[np.random.normal(80,scale=std,size=100) for std in np.random.randint(1,7,4)]
tee_grades=pd.DataFrame({'MA103':data[0],'MA104':data[1],'MA477':data[2],'MA371':data[3]})
tee_grades.head()
# <h2> Scatter Plots</h2>
plt.figure(figsize=(8,6))
plt.scatter(x=tee_grades['MA103'],y=tee_grades['MA104'],c='red',s=72)
# <h2>Histograms and Distribution Plots</h2>
plt.figure(figsize=(8,6))
plt.hist(x=tee_grades['MA477'],density=True,edgecolor='red')
plt.show()
tee_grades.T
# <h2> Box Plots</h2>
#
# A box plot (or formally known as box and whisker plots) displays the five-number summary of a dataset. The five numbers are the minimum, first quartile, median, third quartile, and maximum.
#
#
# In a box plot, one draws a box from the first quartile to the third quartile. A vertical line goes through the box at the median. The whiskers go from each quartile to the minimum or maximum.
#
# We illustrate this with the example below.
plt.figure(figsize=(10,6))
plt.boxplot(tee_grades.T, vert=True, patch_artist=True,showmeans=False)
plt.xlabel('Courses',fontsize=14)
plt.ylabel('TEE Scores',fontsize=14)
plt.xticks([1,2,3,4],labels=['MA103','MA104','MA206','MA477','MA371'],fontsize=14)
plt.title("Grade Distributions by Course",fontsize=18)
plt.show()
# <b>Minimum:</b> The smallest number, represented by the lowest horizontal line, excluding any outliers
# <br>
#
# <b>Maximum:</b> The largest number, represented by the highest (up-top) horizontal line, excluding any outliers
# <br>
#
# <b> Median (Q2/50th percentile):</b> Is the middle value of the dataset
# <br>
#
# <b> First Quartile (Q1/ 25th percentile):</b> is represented by the lower edge of the actual box. It is the middle value between the smallest number(not the minimum) and the median.
# <br>
#
# <b> Third Quartile (Q3/75th percentile):</b> is represented by the upper edge of the actual box. It is the middle value between the largest number (not the maximum) and the median.
#
# <b>Interquartile Range (IQR)</b>:$$IQR=Q3-Q1$$
# <br>
# <b>Outlier:</b> An outlier is considered any value that is 1.5 times the IQR above $Q3$ or below $Q1$, respectively. That is any value that is above $1.5IQR+Q3$ or below $Q1-1.5IQR$.
#
# <b> Lower Whisker:</b> It is drawn at the smallest data value that falls within a distance of $1.5IQR$ from $Q1$.
#
# <b> Upper Whisker:</b> It is drawn at the largest data value that falls within a distance of $1.5IQR$ from $Q3$.
| MA477 - Theory and Applications of Data Science/Lessons/Lesson 4 - Matplotlib/Lesson 4 - Matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Regression Discontinuity Analysis (RDD) in Python
# This tutorial explains and applies RDD by reproduction of the results in [<NAME> Campbell,1960](http://dx.doi.org/10.1037/h0044319). For this purpose, we introduce the python package [rdd](https://github.com/evan-magnusson/rdd).
# ## Scenario
# Thistlethwait and Campell studied the causal effect of Certificates of Merrit (CoM) on short and long term career development. In the USA, a CoM is awarded to those students, who score above a certain threshold score. Now, receiving a CoM is obviously confounded by many variable, e.g. socio-economic status of the family, place of residence, etc.. To get an unbiased estimate of the causal effect, the authors developed RDD.
# RDD exploits the fact, that students become arbitrarily similar in their characteristic variables, when we reduce the set to those who score close to the threshold score. This is because the influence of counfounding variables becomes dominated by variables s.a. day-dependent performance, sickness, family-tragedy, etc.. In practice, this means we can get an unbiased estimate of the causal effect of CoM on e.g. receving a scholarship through fiting a regression model which allows for a discontinuity at threshold score to the datapoint in vicinity of that threshold score. With their analysis, Thistlethwait and Campell could show, that CoM does effect the chances for receiving a scholarship, but is irrelevant for long-term career plans ([Marinescu2018](https://doi.org/10.1038/s41562-018-0466-5)).
# [Marinescu2018](https://doi.org/10.1038/s41562-018-0466-5)
#
#
# ## Application
# Translating into analysis tools, RDD merely requires us to define a bandwidth of data around a threshold. We then need to truncate our data to this bandwidth. After that, we are ready to fit a regression model of outcome onto the running variable on both sides of the threshold. All three steps are implemented in the rdd python package, which we are going to use for our demonstration below.
# $$
# Y_{i} \sim (\alpha_{0} + \beta_0 T_{i}) + (\alpha_{1} + \beta_1 T_{i})\tilde{X_i} + \sum_{k=2}^{K} (\alpha_{k} + \beta_k T_{i}) \phi_{k}(\tilde{X_i}),\:\:\tiny{\begin{aligned}Y&:\text{dependent variable}\\\tilde{X_i} = X_{i} - c&:\text{zero-shifted independent variable}\\c&:\text{treshold value}\\T_{i}\in\{0,1\}&:\text{dummy variable indicating treatment}\end{aligned}}\\
# $$
# ([Keio2016](http://web.mit.edu/teppei/www/teaching/Keio2016/05rd.pdf), [Trochim2006](http://socialresearchmethods.net/kb/statrd.htm))
# The model allows for a disconituity at the value of the theshold in all regression coefficients. Since we ensured, that the subjects in our truncated dataset are similar up to random perturbations in all but the running variable, this discontinuity is accountable to the treatment. The coefficient $\beta_0$, thus, quantifies the causal effect of the treatment in an unbiased fashion. If we have a true causal effect of the treatment, we can fit this coefficient with significance.
# Just as any other analysis method, RDD come with it own specific set of assumption, that the user needs to be aware of. Firstly, The general setting required the threshold to be of binary nature and the running variable to be continuous. Secondly, if the outcome may be controlled precisely, such that we cannot even expect a random distribution of the data around the threshold, we cannot apply RDD. Further, potential confounder may not have a discontinuity at the threshold value, too. Lastly, there must not be any influence on the threshold value by the subjects generating the data.
# ([Marinescu2018](https://doi.org/10.1038/s41562-018-0466-5))
# ### in Neuroscience
# Thresholds are ubiquitous in neuroscience, firing thresholds just being the most prominent ones. The following tables was put together by [Marinescu et al.](https://doi.org/10.1038/s41562-018-0466-5), [Landsdell et al.](https://doi.org/10.1101/253351)).
#
# | Area | Question | Running variable | Threshold | Outcome variable |
# | ----------- |:------------------:| -----------------:| --------- | ---------------- |
# | Neural data science | What are the neural requirements for movement | Neural drive | Firing threshold | Activity of a downstream neuron or muscle |
# | Neural Theory | How much would larger synaptic weight increase reward-seeking behaviour? | Neural drive | Firing Threshold | Behavioural Change |
# | Theoretical Neuroscience | What is the causal contribution of a neuron's activity on the reward signal? | Neural drive | Firing Threshold | Reward |
#
# [Landsdell et al.](https://doi.org/10.1101/253351) provide an example in the theoretical neurosciences (see table, row 3). They build a simple model in which neurons can use RDD to estimate the causal effect of their own activity on a reward signal. The model is based on a intergrate-and-fire model. Neural drive, i.e. the leaky integrated input to the neuron, is the running (dependent) variable. Of course, whenever the input exceeds a threshold, the neuron elicits a spike. In response to the spike, the neuron receives a reward signal, e.g. through feed-back neuromodulator signal. Then, the neuron can compare its marginally subthreshold, i.e. sub-spike, neural drive to its marginally supthreshold neural drive. The difference is its estimate of its on contribution to the reward signal. By means of RDD, a neuron can quite easily estimate this contribution even in the presence of correlated inputs. [Landsdell et al.](https://doi.org/10.1101/253351) show that a two-neuron integrate-fire-model with a RDD-based learning rule can successfully estimate causal effects.
#
# ## Application
# For means of plasticity, we want to reproduce the results by Campbell and Thistlethwait using the original data and an open-source python package.
# Load libraries:
# +
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# RDD
from rdd import rdd
# DATASCIENCE
import numpy as np
import pandas as pd
import statsmodels.api as sm
# PLOTTING
import matplotlib.pyplot as plt
import seaborn as sns
# STYLING
from jupyterthemes import jtplot
jtplot.style(theme='chesterish')
# %matplotlib inline
# -
# ### Load data
# We are going to use the Data available on [github repo](https://github.com/jrnold/datums). You can either download it through your browser and copy it to the folder 'causal_neuroscience/data/raw' or in case you have wget installed simply run the command below.
# %pwd
# !wget http://github.com/jrnold/datums/blob/master/data-raw/CampbellThistlewaite1960.csv
path_2_data = '../data/raw/CampbellThistlewaite1960.csv'
data = pd.read_csv(path_2_data, index_col=None)
data
data.info()
# ### Data preparation
# We are going to use a derivate of the *scholarship* variable as our target (dependent variable) in the regression analysis later on. For unobstructed and clean analysis, we therefore drop the rows containing `NaN` values in the *scholarship* column right away. Further, change the `dtype` of the *test_score* to a numeric type.
data['test_score'].iloc[0] = 0
data['test_score'].iloc[-1] = 21
data['test_score'] = data['test_score'].astype('int64')
data = data.loc[~data['scholarship'].isna()]
data.head(3)
data.tail(3)
data.info()
# We compute the realive number of students in each score intervall that received a scolarship.
data['rel_num_students_w_scholarship'] = data['scholarship'] / data['number_in_sample']
data.head()
# ### Data visualization
# Let's scatter our data to see whether we discern any linear relationships or even thresholds.
# PARAMETERS
predictor = 'test_score' # independent variable, i.e. running variable
target = 'rel_num_students_w_scholarship' # dependent variable
threshold = 10.5 # merrits where awarded for students scoring >10
# +
# VISUALIZATION
# create plot objects
fig = plt.figure(figsize=(8, 8))
ax = fig.add_subplot(111)
# plot data
_ = sns.scatterplot(x=predictor,
y=target,
data=data,
s=62,
ax=ax)
_ = ax.axvline(threshold, color='r', linestyle='dashed', label='merrit score-line') # threshold line
# label plot
_ = ax.set_xlabel('test score')
_ = ax.set_ylabel('percentage of students within interval\nreceiving a scholarship')
_ = ax.set_title('Scatter plot of student test scores over\npercentange of those students receiving merrits', fontsize=16)
_ = ax.legend()
# pretify plot
_ = ax.grid(False)
_ = ax.set_xticks(data[predictor][::2])
_ = ax.set_xticklabels(data[predictor][::2])
# show plot
plt.show()
# -
# The illustration might allow to hypothesize about some linear relation between, e.g. between the theta frequency band data, but certainly not about any thresholds.
# ### Basic linear regression
# As reference we fit a simple linear regression of our predictor *test_score* on the target *rel_num_students_w_scholarship*.
# +
# Create linear regression model
regr = {}
X = data[predictor]
X = sm.add_constant(X) # add column of 1s for slope interception
y = data[target]
regression_model = sm.OLS(y, X, missing='drop').fit()
print(regression_model.summary())
# -
# *Evaluation*
# ### Regression Discontiuity Analysis
# +
# bandwidth_opt = rdd.optimal_bandwidth(data[predictor], data[target], cut=threshold)
# -
# truncate data
# data_rdd = rdd.truncated_data(data,
# xname=predictor,
# bandwidth=bandwidth_opt,
# yname=target,
# cut=threshold)
data_rdd = data
# We perform RDD analysis using the rdd toolbox. The return is an ordinary least squares fit to a linear model. However, more general models can be passed to the function. For details check [rdd tutorial in github](https://github.com/evan-magnusson/rdd/blob/master/tutorial/tutorial.ipynb) and the [statsmodels documentary](https://www.statsmodels.org/dev/generated/statsmodels.regression.linear_model.OLS.html).
# +
# perform rdd analysis
rdd_model = rdd.rdd(data_rdd, xname=predictor, yname=target, cut=threshold)
rdd_model = rdd_model.fit()
# print results
print(rdd_model.summary())
# -
# *evaluation*
# Let's add the *TREATED* variable to our data.
# (Conversion of dtype from boolean to int is necessary for compatibility with `sm.OLS` model.
data['TREATED'] = (~data['merit_scholars'].isna()).astype(int)
print('Head')
data.head(3)
print('Tail')
data.tail(3)
# Let's predict our model and store the results in our original dataframe.
prediction = 'scholarship_prediction'
data[prediction] = rdd_model.predict(data)
# data
# Let's visualize our results.
# +
# VISUALIZATION
# create plot objects
fig = plt.figure(figsize=(8, 8))
ax = fig.add_subplot(111)
# plot data
_ = sns.scatterplot(x=predictor,
y=target,
data=data,
s=62,
ax=ax)
_ = ax.axvline(threshold, color='r', linestyle='dashed', label='merrit score-line') # threshold line
_ = sns.lineplot(x=predictor, y=prediction, data=data, label='prediction')
# label plot
_ = ax.set_xlabel('test score')
_ = ax.set_ylabel('percentage of students within interval\nreceiving a scholarship')
_ = ax.set_title('Scatter plot of student test scores over\npercentange of those students receiving merrits', fontsize=16)
_ = ax.legend()
# pretify plot
_ = ax.grid(False)
_ = ax.set_xticks(data[predictor][::2])
_ = ax.set_xticklabels(data[predictor][::2])
# show plot
plt.show()
# -
| notebooks/rdd_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: QSRL
# language: python
# name: python3
# ---
# + [markdown] id="ef0f3a7e"
# ## Quantitative trading in China A stock market with FinRL
# + [markdown] id="42ac7297"
# ### Import modules
# + id="fluid-taylor" outputId="2549105a-da76-4154-c404-1a5b390d0f93"
import warnings
warnings.filterwarnings("ignore")
import pandas as pd
from IPython import display
display.set_matplotlib_formats("svg")
from finrl_meta import config
from finrl_meta.data_processors.processor_tusharepro import TushareProProcessor, ReturnPlotter
from finrl_meta.env_stock_trading.env_stocktrading_A import StockTradingEnv
from drl_agents.stablebaselines3_models import DRLAgent
pd.options.display.max_columns = None
print("ALL Modules have been imported!")
# + [markdown] id="eb601f4a"
# ### Create folders
# + id="339ab411"
import os
if not os.path.exists("./datasets" ):
os.makedirs("./datasets" )
if not os.path.exists("./trained_models"):
os.makedirs("./trained_models" )
if not os.path.exists("./tensorboard_log"):
os.makedirs("./tensorboard_log" )
if not os.path.exists("./results" ):
os.makedirs("./results" )
# + [markdown] id="74ad0a26"
# ### Download data, cleaning and feature engineering
# + id="transsexual-crack"
ticket_list=['600000.SH', '600009.SH', '600016.SH', '600028.SH', '600030.SH',
'600031.SH', '600036.SH', '600050.SH', '600104.SH', '600196.SH',
'600276.SH', '600309.SH', '600519.SH', '600547.SH', '600570.SH']
train_start_date='2015-01-01'
train_stop_date='2019-08-01'
val_start_date='2019-08-01'
val_stop_date='2021-01-03'
token='<KEY>'
# + id="preceding-selling" outputId="49ad1a8e-50b6-4da5-b731-838cade7673f"
# download and clean
ts_processor = TushareProProcessor("tusharepro", token=token)
ts_processor.download_data(ticket_list, train_start_date, val_stop_date, "1D")
ts_processor.clean_data()
ts_processor.dataframe
# + id="3e40b006" outputId="b6c70823-738e-43c6-8cbf-9955a66cb183"
# add_technical_indicator
ts_processor.add_technical_indicator(config.TECHNICAL_INDICATORS_LIST)
ts_processor.clean_data()
ts_processor.dataframe
# + [markdown] id="25fc2e45"
# ### Split traning dataset
# + id="pending-mother" outputId="e388671a-c877-4b10-d2b8-3ee6c1bb27ba"
train =ts_processor.data_split(ts_processor.dataframe, train_start_date, train_stop_date)
len(train.tic.unique())
# + id="signal-rochester" outputId="ab196937-4f36-4a88-8f36-4548c93a016e"
train.tic.unique()
# + id="future-while" outputId="4b0c4964-259f-4d2a-cd67-471e5485b00e"
train.head()
# + id="72e9bcc2" outputId="c9221982-0c92-46a0-a0a1-2fc571205388"
train.shape
# + id="provincial-wichita" outputId="0ed4e0cd-5883-407a-8aa7-2bc91af39597"
stock_dimension = len(train.tic.unique())
state_space = stock_dimension*(len(config.TECHNICAL_INDICATORS_LIST)+2)+1
print(f"Stock Dimension: {stock_dimension}, State Space: {state_space}")
# + [markdown] id="e90bbf93"
# ### Train
# + id="dcb153fc"
env_kwargs = {
"stock_dim": stock_dimension,
"hmax": 1000,
"initial_amount": 1000000,
"buy_cost_pct":6.87e-5,
"sell_cost_pct":1.0687e-3,
"reward_scaling": 1e-4,
"state_space": state_space,
"action_space": stock_dimension,
"tech_indicator_list": config.TECHNICAL_INDICATORS_LIST,
"print_verbosity": 1,
"initial_buy":True,
"hundred_each_trade":True
}
e_train_gym = StockTradingEnv(df = train, **env_kwargs)
# + id="loaded-modem" outputId="3a2dd72a-ac29-4717-be98-97058db58eda"
env_train, _ = e_train_gym.get_sb_env()
print(type(env_train))
# + id="thick-blackjack" outputId="7bf8c4ae-df5f-415c-8e81-0be4a21174a5"
agent = DRLAgent(env = env_train)
DDPG_PARAMS = {
"batch_size": 256,
"buffer_size": 50000,
"learning_rate": 0.0005,
"action_noise":"normal",
}
POLICY_KWARGS = dict(net_arch=dict(pi=[64, 64], qf=[400, 300]))
model_ddpg = agent.get_model("ddpg", model_kwargs = DDPG_PARAMS, policy_kwargs=POLICY_KWARGS)
# + code_folding=[] id="growing-supplier" outputId="a72243f4-08b0-4882-824c-20ef5fa6e1d8"
trained_ddpg = agent.train_model(model=model_ddpg,
tb_log_name='ddpg',
total_timesteps=1000)
# + [markdown] id="0767b826"
# ### Trade
# + id="responsible-equity"
trade = ts_processor.data_split(df, val_start_date, val_stop_date)
env_kwargs = {
"stock_dim": stock_dimension,
"hmax": 1000,
"initial_amount": 1000000,
"buy_cost_pct":6.87e-5,
"sell_cost_pct":1.0687e-3,
"reward_scaling": 1e-4,
"state_space": state_space,
"action_space": stock_dimension,
"tech_indicator_list": config.TECHNICAL_INDICATORS_LIST,
"print_verbosity": 1,
"initial_buy":False,
"hundred_each_trade":True
}
e_trade_gym = StockTradingEnv(df = trade, **env_kwargs)
# + id="first-hierarchy" outputId="2d6c5731-3bb0-4c48-a298-42e500dde19f"
df_account_value, df_actions = DRLAgent.DRL_prediction(model=trained_ddpg,
environment = e_trade_gym)
# + id="8b9d6c2b" outputId="e14ed292-4149-4630-a59d-5d22204cd6b2"
df_actions.to_csv("action.csv",index=False)
df_actions
# + [markdown] id="6ea8a81c"
# ### Backtest
# + id="727d62e0" outputId="a65f3198-34ac-422b-cb99-8908cdde7e3a"
# %matplotlib inline
plotter = ReturnPlotter(df_account_value, trade, val_start_date, val_stop_date)
plotter.plot_all()
# + id="8813f87d" outputId="7ff9fedc-51cc-4781-a1bc-856bd68ea33b"
# %matplotlib inline
plotter.plot()
# + id="d155bcd5" outputId="81b3990a-2e08-42f7-a665-b63bf7cb5c34"
# %matplotlib inline
# ticket: SSE 50:000016
plotter.plot("000016")
# + [markdown] id="ce724f71"
# #### Use pyfolio
# + id="79c82f77" outputId="882d7be9-9a58-4bdb-a89a-a2b4b2f0c8cd"
# CSI 300
baseline_df = plotter.get_baseline("399300")
# + id="e4ab0438" outputId="774b08b1-dd92-461a-892b-6756b57cbfea"
import pyfolio
from pyfolio import timeseries
daily_return = plotter.get_return(df_account_value)
daily_return_base = plotter.get_return(baseline_df, value_col_name="close")
perf_func = timeseries.perf_stats
perf_stats_all = perf_func(returns=daily_return,
factor_returns=daily_return_base,
positions=None, transactions=None, turnover_denom="AGB")
print("==============DRL Strategy Stats===========")
perf_stats_all
# + id="8215cc99" outputId="0e4bb4f7-d9de-46ae-863a-339f67397a05"
with pyfolio.plotting.plotting_context(font_scale=1.1):
pyfolio.create_full_tear_sheet(returns = daily_return,
benchmark_rets = daily_return_base, set_context=False)
# + [markdown] id="ce11979d"
# ### Authors
# github username: oliverwang15, eitin-infant
| Demo_China_A_share_market.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Training code:
import tensorflow as tf
import numpy as np
from sklearn.model_selection import train_test_split
# Load sample data
X = np.loadtxt("X.csv", delimiter = ",")
y = np.loadtxt("y.csv")
# Convert from indices into categorical form.
# Note: More general approach is:
# y = tf.keras.utils.to_categorical(y)
y = (y == 1)
# Split to training and evaluation:
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2)
# First we initialize the model. "Sequential" means there are no loops.
clf = tf.keras.models.Sequential()
# Add layers one at the time. Each with 100 nodes.
clf.add(tf.keras.layers.Dense(100, input_dim=2, activation = 'sigmoid'))
clf.add(tf.keras.layers.Dense(100, activation = 'sigmoid'))
clf.add(tf.keras.layers.Dense(1, activation = 'sigmoid'))
# The code is compiled to CUDA or C++
clf.compile(loss='binary_crossentropy', optimizer='adam')
clf.fit(X_train, y_train, epochs=20, batch_size=16) # takes a few seconds
# Check accuracy
y_pred = clf.predict(X_test)
# The model outputs probabilities, so let's threshold at 0.5:
y_pred = (y_test > 0.5)
accuracy = np.mean(y_test == y_pred)
print("Accuracy on test data is %.2f %%" % (100 * accuracy))
| code/keras_first_network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.0
# language: julia
# name: julia-1.6
# ---
using DataFrames, TableOperations, Tables, Random
using Recommender: Movielens1M, load_dataset, ratio_split, ImplicitMF, evaluate_u2i, PrecisionAtK, RecallAtK, NDCG, fit!, predict_u2i, make_u2i_dataset
ml1M = Movielens1M()
download(ml1M)
rating, user, movie = load_dataset(ml1M);
rating = rating |> TableOperations.filter(x->Tables.getcolumn(x, :rating) >= 4)
movie2title = Dict()
for row in Tables.rows(movie)
movie2title[row[:movieid]] = row[:title]
end
rating = rating |> TableOperations.transform(Dict(:movieid=>x->movie2title[x]))
# +
Random.seed!(1234);
train_valid_table, test_table = ratio_split(rating, 0.8)
train_table, valid_table = ratio_split(train_valid_table, 0.8)
length(Tables.rows(train_table)), length(Tables.rows(valid_table)), length(Tables.rows(test_table))
# -
prec10 = PrecisionAtK(10)
recall10 = RecallAtK(10)
ndcg10 = NDCG(10)
metrics = [prec10, recall10, ndcg10]
model = ImplicitMF(128, true, 0.005)
evaluate_u2i(model, train_valid_table, test_table, metrics, 10, col_item=:movieid, n_epochs=256, n_negatives=8, learning_rate=0.002, drop_history=true)
| examples/implicit_mf_ml1M.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
from __future__ import print_function, absolute_import, division
import numpy as np
from wgomoku import (
GomokuBoard, GomokuTools as gt, roll_out,
NH9x9, Heuristics, HeuristicGomokuPolicy)
import random
import tensorflow as tf
A,B,C,D,E,F,G,H,I,J,K,L,M,N,O,P,Q,R,S,T,U = \
1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21
BLACK=0
WHITE=1
EDGES=2
heuristics = Heuristics(kappa=3.0)
def new_initial_state(heuristics):
board = GomokuBoard(heuristics, N=15, disp_width=8)
policy = HeuristicGomokuPolicy(bias=.5, topn=5, style=2)
board.set(H,8).set('G',6).set(G,8).set(F,8).set(H,9).set(H,10)
return board, policy
board = new_initial_state(heuristics)[0]
board.display('current')
from wgomoku import transform
stones=board.stones.copy()
transformed = transform(stones, board.N, quarters=0, reflect=True)
GomokuBoard(heuristics, N=15, stones=transformed).display('current')
stones=board.stones.copy()
transformed = transform(stones, board.N, quarters=1)
GomokuBoard(heuristics, N=15, stones=transformed).display('current')
stones=board.stones.copy()
transformed = transform(stones, board.N, quarters=2)
GomokuBoard(heuristics, N=15, stones=transformed).display('current')
stones=board.stones.copy()
transformed = transform(stones, board.N, quarters=3)
GomokuBoard(heuristics, N=15, stones=transformed).display('current')
| GomokuSymmetries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Import Statement
from pyspark.sql import SQLContext
from handyspark import *
from pyspark.sql import functions as sf
from matplotlib import pyplot as plt
from pyspark.sql.functions import col, avg
import pandas as pd
from pyspark.sql.window import Window
log4jLogger = sc._jvm.org.apache.log4j
LOGGER = log4jLogger.LogManager.getLogger(__name__)
LOGGER.error("pyspark script logger initialized")
base_path = "/home/darkmatter/Desktop/smart-meters-in-london/"
sqlcontext = SQLContext(sc)
household_info = sqlcontext.read.csv(base_path+"informations_households.csv",header=True,inferSchema=True)
household_mini = sc.parallelize(household_info.take(15)).toDF()
# household_mini = household_info
household_mini.show()
column_list = []
for i in range(48):
column_list.append("hh_"+str(i))
column_list
new_column_list = []
for i in range(1,49):
if i<20:
new_column_list.append("0"+str(i*0.5))
else:
new_column_list.append(str(i*0.5))
flag = 0
avg_house_data = []
df_file = household_mini.select("file").distinct()
# exprs = {x: "avg" for x in new_column_list}
exprs1 = [avg(x) for x in column_list[0:40]]
exprs2 = [avg(x) for x in column_list[40:48]] #due to the fact large number of column giving error so divide
for row in df_file.rdd.collect():
file = row.file
print(file)
file_path = base_path + "hhblock_dataset/"+ file+".csv"
half_hourly_consumption_data = sqlcontext.read.csv(file_path,header=True,inferSchema=True)
half_hourly_consumption_data.dropna(how='any')
half_hourly_consumption_data2 = half_hourly_consumption_data.groupBy('LCLid').agg(*exprs2)
half_hourly_consumption_data = half_hourly_consumption_data.groupBy('LCLid').agg(*exprs1)
half_hourly_consumption_data = half_hourly_consumption_data.join(half_hourly_consumption_data2,["LCLid"])
half_hourly_consumption_data.dropna(how='any')
half_hourly_consumption_data.printSchema()
if flag == 0:
avg_house_data = sqlcontext.createDataFrame([],half_hourly_consumption_data.schema)
flag = 1
avg_house_data = avg_house_data.union(half_hourly_consumption_data)
half_hourly_consumption_data.show()
flag = 0
avg_house_data = []
block_read = set([])
for row in household_mini.rdd.collect():
house_id = row.LCLid
file = row.file
print(house_id,file)
file_path = base_path + "hhblock_dataset/"+ file+".csv"
if file not in block_read:
# print("hi")
block_read.add(file)
half_hourly_consumption_data = sqlContext.read.csv(file_path,header=True,inferSchema=True)
half_hourly_consumption_data.dropna(how='any')
for c,n in zip(column_list,new_column_list):
half_hourly_consumption_data=half_hourly_consumption_data.withColumnRenamed(c,n)
indiv_house_data = half_hourly_consumption_data.where(col("LCLid") == house_id)
indiv_house_data = indiv_house_data.toHandy()
if indiv_house_data.rdd.isEmpty():
print("Missing Id = {} in file = {}".format(house_id,file))
continue
indiv_house_data = sqlcontext.createDataFrame(indiv_house_data.stratify(['LCLid']).cols[new_column_list].mean().reset_index())
# indiv_house_data.printSchema()
if flag == 0:
avg_house_data = sqlcontext.createDataFrame([],indiv_house_data.schema)
flag = 1
avg_house_data = avg_house_data.union(indiv_house_data)
for c,n in zip(avg_house_data.columns[1:],new_column_list):
avg_house_data=avg_house_data.withColumnRenamed(c,n)
avg_house_data.show()
avg_house_data = avg_house_data.toPandas().set_index("LCLid")
# pd.options.display.max_columns = None
avg_house_data.head(15)
avg_house_data.T.plot(figsize=(13,8), legend=False, color='blue',alpha=0.5)
avg_house_data.to_csv(base_path+"avg.csv")
| Other_Scrap_file/aa/Average_load_profile.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
Make a request from the Forecast.io API for where you were born (or lived, or want to visit!)
import requests
# !pip3 install requests
#new york
response = requests.get("https://api.forecast.io/forecast/94bc3fa3628bfad686b10e7054c67f71/40.7141667, -74.0063889")
data = response.json()
print(data)
# 2) What's the current wind speed? How much warmer does it feel than it actually is?
#
type(data)
data.keys()
print(data['currently'])
print(data['currently']['temperature']-data['currently']['apparentTemperature'])
# 3) The first daily forecast is the forecast for today. For the place you decided on up above, how much of the moon is currently visible?
print(data['daily'])
type(data['daily'])
data['daily'].keys()
print(data['daily']['data'][0])
type(data['daily']['data'])
print(data['daily']['data'][0]['moonPhase'])
# 4) What's the difference between the high and low temperatures for today?
#
weather_today = data['daily']['data'][0]
print(weather_today['temperatureMax']-weather_today['temperatureMin'])
# 5) Loop through the daily forecast, printing out the next week's worth of predictions. I'd like to know the high temperature for each day, and whether it's hot, warm, or cold, based on what temperatures you think are hot, warm or cold.
#
print(data['daily']['data'])
daily_data = data['daily']['data']
weather_next_week = data['daily']['data']
for weather in weather_next_week:
print(weather['temperatureMax'])
if weather['temperatureMax'] > 84:
print("it's a hot day.")
elif weather['temperatureMax'] > 74 and weather['temperatureMax'] < 83:
print("it's a warm day.")
else:
print("it's a cold day.")
# 6) What's the weather looking like for the rest of today in Miami, Florida? I'd like to know the temperature for every hour, and if it's going to have cloud cover of more than 0.5 say "{temperature} and cloudy" instead of just the temperature.
import requests
response = requests.get("https://api.forecast.io/forecast/94bc3fa3628bfad686b10e7054c67f71/25.7738889, -80.1938889")
data = response.json()
print(data['hourly'])
data['hourly'].keys()
data['hourly']['data']
for cloudcover in data['hourly']['data']:
if cloudcover['cloudCover'] > 0.5:
print(cloudcover['temperature'], "and cloudy")
else:
print(cloudcover['temperature'])
# 7) What was the temperature in Central Park on Christmas Day, 1980? How about 1990? 2000?
# Tip: You'll need to use UNIX time, which is the number of seconds since January 1, 1970. Google can help you convert a normal date!
# Tip: You'll want to use Forecast.io's "time machine" API at https://developer.forecast.io/docs/v2
import requests
response = requests.get("https://api.forecast.io/forecast/94bc3fa3628bfad686b10e7054c67f71/40.7141667, -74.0063889,346550400")
data = response.json()
print(data['currently']['temperature'])
response = requests.get("https://api.forecast.io/forecast/94bc3fa3628bfad686b10e7054c67f71/40.7141667, -74.0063889,662083200")
data = response.json()
print(data['currently']['temperature'])
response = requests.get("https://api.forecast.io/forecast/94bc3fa3628bfad686b10e7054c67f71/40.7141667, -74.0063889,977702400")
data = response.json()
print(data['currently']['temperature'])
| homework_6_shengying_zhao.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="3Q2KuaRamHfr"
# # Survival Estimates that Vary with Time
#
# Welcome to the third assignment of Course 2. In this assignment, we'll use Python to build some of the statistical models we learned this past week to analyze surivival estimates for a dataset of lymphoma patients. We'll also evaluate these models and interpret their outputs. Along the way, you will be learning about the following:
#
# - Censored Data
# - Kaplan-Meier Estimates
# - Subgroup Analysis
# -
# ## Outline
#
# - [1. Import Packages](#1)
# - [2. Load the Dataset](#2)
# - [3. Censored Data](#)
# - [Exercise 1](#Ex-1)
# - [4. Survival Estimates](#4)
# - [Exercise 2](#Ex-2)
# - [Exercise 3](#Ex-3)
# - [5. Subgroup Analysis](#5)
# - [5.1 Bonus: Log Rank Test](#5-1)
# + [markdown] colab_type="text" id="UopnLTeLkViX"
# <a name='1'></a>
# ## 1. Import Packages
#
# We'll first import all the packages that we need for this assignment.
#
# - `lifelines` is an open-source library for data analysis.
# - `numpy` is the fundamental package for scientific computing in python.
# - `pandas` is what we'll use to manipulate our data.
# - `matplotlib` is a plotting library.
# + colab={} colab_type="code" id="TZyXoADQmYlt"
import lifelines
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from util import load_data
from lifelines import KaplanMeierFitter as KM
from lifelines.statistics import logrank_test
# + [markdown] colab_type="text" id="5rp2TD1qnGmp"
# <a name='2'></a>
# ## 2. Load the Dataset
#
# + [markdown] colab_type="text" id="WEbu3MtrVsnU"
# Run the next cell to load the lymphoma data set.
# + colab={} colab_type="code" id="e3wHdLrEnSNa"
data = load_data()
# + [markdown] colab_type="text" id="3hrHa0dPqU08"
# As always, you first look over your data.
# + colab={"base_uri": "https://localhost:8080/", "height": 221} colab_type="code" id="QEd504pKqWuc" outputId="7297830a-d316-4623-bb6a-77f8f96b8805"
print("data shape: {}".format(data.shape))
data.head()
# + [markdown] colab_type="text" id="dblUOLQS4UU0"
# The column `Time` states how long the patient lived before they died or were censored.
#
# The column `Event` says whether a death was observed or not. `Event` is 1 if the event is observed (i.e. the patient died) and 0 if data was censored.
#
# Censorship here means that the observation has ended without any observed event.
# For example, let a patient be in a hospital for 100 days at most. If a patient dies after only 44 days, their event will be recorded as `Time = 44` and `Event = 1`. If a patient walks out after 100 days and dies 3 days later (103 days total), this event is not observed in our process and the corresponding row has `Time = 100` and `Event = 0`. If a patient survives for 25 years after being admitted, their data for are still `Time = 100` and `Event = 0`.
# + [markdown] colab_type="text" id="L0d2s2wtn2Pf"
# <a name='3'></a>
# ## 3. Censored Data
#
# We can plot a histogram of the survival times to see in general how long cases survived before censorship or events.
# + colab={"base_uri": "https://localhost:8080/", "height": 285} colab_type="code" id="cFrvXrODZklx" outputId="a260b523-e792-47dd-9833-7677cd4b5741"
data.Time.hist();
plt.xlabel("Observation time before death or censorship (days)");
plt.ylabel("Frequency (number of patients)");
# Note that the semicolon at the end of the plotting line
# silences unnecessary textual output - try removing it
# to observe its effect
# + [markdown] colab_type="text" id="ohXd70ZBaRWJ"
# <a name='Ex-1'></a>
# ### Exercise 1
#
# In the next cell, write a function to compute the fraction ($\in [0, 1]$) of observations which were censored.
# -
# <details>
# <summary>
# <font size="3" color="darkgreen"><b>Hints</b></font>
# </summary>
# <p>
# <ul>
# <li>Summing up the <code>'Event'</code> column will give you the number of observations where censorship has NOT occurred.</li>
#
# </ul>
# </p>
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="9fkHnfJ6bD0f" outputId="21ec0301-5884-48f2-8ed1-10795d567c31"
# UNQ_C1 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def frac_censored(df):
"""
Return percent of observations which were censored.
Args:
df (dataframe): dataframe which contains column 'Event' which is
1 if an event occurred (death)
0 if the event did not occur (censored)
Returns:
frac_censored (float): fraction of cases which were censored.
"""
result = 0.0
### START CODE HERE ###
result = np.sum(df.Event == 0)/df.shape[0]
### END CODE HERE ###
return result
# -
print(frac_censored(data))
# #### Expected Output:
# ```CPP
# 0.325
# ```
# + [markdown] colab_type="text" id="BpzYWXUpbk6x"
# Run the next cell to see the distributions of survival times for censored and uncensored examples.
# + colab={"base_uri": "https://localhost:8080/", "height": 545} colab_type="code" id="1k3qlTQLbulW" outputId="de331041-6612-4df7-953d-41efbc741e49"
df_censored = data[data.Event == 0]
df_uncensored = data[data.Event == 1]
df_censored.Time.hist()
plt.title("Censored")
plt.xlabel("Time (days)")
plt.ylabel("Frequency")
plt.show()
df_uncensored.Time.hist()
plt.title("Uncensored")
plt.xlabel("Time (days)")
plt.ylabel("Frequency")
plt.show()
# + [markdown] colab_type="text" id="WmFDrzYrn-JA"
# <a name='4'></a>
# ## 4. Survival Estimates
#
# We'll now try to estimate the survival function:
#
# $$
# S(t) = P(T > t)
# $$
#
# To illustrate the strengths of <NAME>, we'll start with a naive estimator of the above survival function. To estimate this quantity, we'll divide the number of people who we know lived past time $t$ by the number of people who were not censored before $t$.
#
# Formally, let $i$ = 1, ..., $n$ be the cases, and let $t_i$ be the time when $i$ was censored or an event happened. Let $e_i= 1$ if an event was observed for $i$ and 0 otherwise. Then let $X_t = \{i : T_i > t\}$, and let $M_t = \{i : e_i = 1 \text{ or } T_i > t\}$. The estimator you will compute will be:
#
# $$
# \hat{S}(t) = \frac{|X_t|}{|M_t|}
# $$
#
#
# -
# <a name='Ex-2'></a>
# ### Exercise 2
# Write a function to compute this estimate for arbitrary $t$ in the cell below.
# + colab={"base_uri": "https://localhost:8080/", "height": 306} colab_type="code" id="qUoKpBHJjZM-" outputId="9477925d-ee39-4489-e8be-9af3780f9fc4"
# UNQ_C2 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def naive_estimator(t, df):
"""
Return naive estimate for S(t), the probability
of surviving past time t. Given by number
of cases who survived past time t divided by the
number of cases who weren't censored before time t.
Args:
t (int): query time
df (dataframe): survival data. Has a Time column,
which says how long until that case
experienced an event or was censored,
and an Event column, which is 1 if an event
was observed and 0 otherwise.
Returns:
S_t (float): estimator for survival function evaluated at t.
"""
S_t = 0.0
### START CODE HERE ###
X_t = np.sum(df.Time > t)
M_t = np.sum((df.Event == 1) | (df.Time > t))
S_t = X_t/M_t
### END CODE HERE ###
return S_t
# +
print("Test Cases")
sample_df = pd.DataFrame(columns = ["Time", "Event"])
sample_df.Time = [5, 10, 15]
sample_df.Event = [0, 1, 0]
print("Sample dataframe for testing code:")
print(sample_df)
print("\n")
print("Test Case 1: S(3)")
print("Output: {}, Expected: {}\n".format(naive_estimator(3, sample_df), 1.0))
print("Test Case 2: S(12)")
print("Output: {}, Expected: {}\n".format(naive_estimator(12, sample_df), 0.5))
print("Test Case 3: S(20)")
print("Output: {}, Expected: {}\n".format(naive_estimator(20, sample_df), 0.0))
# Test case 4
sample_df = pd.DataFrame({'Time': [5,5,10],
'Event': [0,1,0]
})
print("Test case 4: S(5)")
print(f"Output: {naive_estimator(5, sample_df)}, Expected: 0.5")
# + [markdown] colab_type="text" id="gKMsSOzfmGwD"
# In the next cell, we will plot the naive estimator using the real data up to the maximum time in the dataset.
# + colab={"base_uri": "https://localhost:8080/", "height": 281} colab_type="code" id="nauOrVd9mNDs" outputId="fc884506-6758-4d22-ab04-6310145f9ed3"
max_time = data.Time.max()
x = range(0, max_time+1)
y = np.zeros(len(x))
for i, t in enumerate(x):
y[i] = naive_estimator(t, data)
plt.plot(x, y)
plt.title("Naive Survival Estimate")
plt.xlabel("Time")
plt.ylabel("Estimated cumulative survival rate")
plt.show()
# + [markdown] colab_type="text" id="jg4VTizxqgKM"
# <a name='Ex-3'></a>
# ### Exercise 3
#
# Next let's compare this with the Kaplan Meier estimate. In the cell below, write a function that computes the Kaplan Meier estimate of $S(t)$ at every distinct time in the dataset.
#
# Recall the Kaplan-Meier estimate:
#
# $$
# S(t) = \prod_{t_i \leq t} (1 - \frac{d_i}{n_i})
# $$
#
# where $t_i$ are the events observed in the dataset and $d_i$ is the number of deaths at time $t_i$ and $n_i$ is the number of people who we know have survived up to time $t_i$.
# -
# <details>
# <summary>
# <font size="3" color="darkgreen"><b>Hints</b></font>
# </summary>
# <p>
# <ul>
# <li>Try sorting by Time.</li>
# <li>Use <a href="https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.unique.html">pandas.Series.unique<a> </li>
# <li>If you get a division by zero error, please double-check how you calculated `n_t`</li>
# </ul>
# </p>
# + colab={"base_uri": "https://localhost:8080/", "height": 527} colab_type="code" id="jnwysrz7CzNG" outputId="a26f0a84-98c2-4fe2-c40e-b18402f8bf20"
# UNQ_C3 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def HomemadeKM(df):
"""
Return KM estimate evaluated at every distinct
time (event or censored) recorded in the dataset.
Event times and probabilities should begin with
time 0 and probability 1.
Example:
input:
Time Censor
0 5 0
1 10 1
2 15 0
correct output:
event_times: [0, 5, 10, 15]
S: [1.0, 1.0, 0.5, 0.5]
Args:
df (dataframe): dataframe which has columns for Time
and Event, defined as usual.
Returns:
event_times (list of ints): array of unique event times
(begins with 0).
S (list of floats): array of survival probabilites, so that
S[i] = P(T > event_times[i]). This
begins with 1.0 (since no one dies at time
0).
"""
# individuals are considered to have survival probability 1
# at time 0
event_times = [0]
p = 1.0
S = [p]
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# get collection of unique observed event times
observed_event_times = np.unique(df.Time)
# sort event times
observed_event_times = sorted(observed_event_times)
# iterate through event times
for t in observed_event_times:
# compute n_t, number of people who survive to time t
n_t = np.sum(df.Time >= t)
# compute d_t, number of people who die at time t
d_t = np.sum((df.Time == t) & (df.Event == 1))
# update p
p *= (1.0 - (d_t/n_t))
# update S and event_times (ADD code below)
# hint: use append
event_times.append(t)
S.append(p)
### END CODE HERE ###
return event_times, S
# +
print("TEST CASES:\n")
print("Test Case 1\n")
print("Test DataFrame:")
sample_df = pd.DataFrame(columns = ["Time", "Event"])
sample_df.Time = [5, 10, 15]
sample_df.Event = [0, 1, 0]
print(sample_df.head())
print("\nOutput:")
x, y = HomemadeKM(sample_df)
print("Event times: {}, Survival Probabilities: {}".format(x, y))
print("\nExpected:")
print("Event times: [0, 5, 10, 15], Survival Probabilities: [1.0, 1.0, 0.5, 0.5]")
print("\nTest Case 2\n")
print("Test DataFrame:")
sample_df = pd.DataFrame(columns = ["Time", "Event"])
sample_df.loc[:, "Time"] = [2, 15, 12, 10, 20]
sample_df.loc[:, "Event"] = [0, 0, 1, 1, 1]
print(sample_df.head())
print("\nOutput:")
x, y = HomemadeKM(sample_df)
print("Event times: {}, Survival Probabilities: {}".format(x, y))
print("\nExpected:")
print("Event times: [0, 2, 10, 12, 15, 20], Survival Probabilities: [1.0, 1.0, 0.75, 0.5, 0.5, 0.0]")
# + [markdown] colab_type="text" id="G7OAiWjS7hLA"
# Now let's plot the two against each other on the data to see the difference.
# + colab={"base_uri": "https://localhost:8080/", "height": 265} colab_type="code" id="JbPlC5717gM_" outputId="06553ed7-9396-4f16-eab3-43a1c6ecddfd"
max_time = data.Time.max()
x = range(0, max_time+1)
y = np.zeros(len(x))
for i, t in enumerate(x):
y[i] = naive_estimator(t, data)
plt.plot(x, y, label="Naive")
x, y = HomemadeKM(data)
plt.step(x, y, label="Kaplan-Meier")
plt.xlabel("Time")
plt.ylabel("Survival probability estimate")
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="iY__6ufG3sDk"
# ### Question
#
# What differences do you observe between the naive estimator and Kaplan-Meier estimator? Do any of our earlier explorations of the dataset help to explain these differences?
# + [markdown] colab_type="text" id="i7tElIKVoQ4R"
# <a name='5'></a>
# ## 5. Subgroup Analysis
#
# We see that along with Time and Censor, we have a column called `Stage_group`.
# - A value of 1 in this column denotes a patient with stage III cancer
# - A value of 2 denotes stage IV.
#
# We want to compare the survival functions of these two groups.
#
# This time we'll use the `KaplanMeierFitter` class from `lifelines`. Run the next cell to fit and plot the Kaplan Meier curves for each group.
# + colab={"base_uri": "https://localhost:8080/", "height": 279} colab_type="code" id="Ge6P3fgVrZLS" outputId="efbf8e54-7623-4d96-e24a-2fbf07ae2aac"
S1 = data[data.Stage_group == 1]
km1 = KM()
km1.fit(S1.loc[:, 'Time'], event_observed = S1.loc[:, 'Event'], label = 'Stage III')
S2 = data[data.Stage_group == 2]
km2 = KM()
km2.fit(S2.loc[:, "Time"], event_observed = S2.loc[:, 'Event'], label = 'Stage IV')
ax = km1.plot(ci_show=False)
km2.plot(ax = ax, ci_show=False)
plt.xlabel('time')
plt.ylabel('Survival probability estimate')
plt.savefig('two_km_curves', dpi=300)
# + [markdown] colab_type="text" id="M4DwaOVEs19Q"
# Let's compare the survival functions at 90, 180, 270, and 360 days
# + colab={} colab_type="code" id="11dhdsUOtEqe"
survivals = pd.DataFrame([90, 180, 270, 360], columns = ['time'])
survivals.loc[:, 'Group 1'] = km1.survival_function_at_times(survivals['time']).values
survivals.loc[:, 'Group 2'] = km2.survival_function_at_times(survivals['time']).values
# + colab={"base_uri": "https://localhost:8080/", "height": 173} colab_type="code" id="-zRlM1SAtYdl" outputId="3642dc3e-01b0-4e96-e91f-8e39c6c0e3e5"
survivals
# + [markdown] colab_type="text" id="RA3amMk__J6e"
# This makes clear the difference in survival between the Stage III and IV cancer groups in the dataset.
# + [markdown] colab_type="text" id="3VoOQREQoXny"
# <a name='5-1'></a>
# ## 5.1 Bonus: Log-Rank Test
#
# To say whether there is a statistical difference between the survival curves we can run the log-rank test. This test tells us the probability that we could observe this data if the two curves were the same. The derivation of the log-rank test is somewhat complicated, but luckily `lifelines` has a simple function to compute it.
#
# Run the next cell to compute a p-value using `lifelines.statistics.logrank_test`.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="_7-QIy8ovsgC" outputId="c7582d94-4c42-4cae-d83d-72a8873fe985"
def logrank_p_value(group_1_data, group_2_data):
result = logrank_test(group_1_data.Time, group_2_data.Time,
group_1_data.Event, group_2_data.Event)
return result.p_value
logrank_p_value(S1, S2)
# + [markdown] colab_type="text" id="nUbv_csdJRSw"
# If everything is correct, you should see a p value of less than `0.05`, which indicates that the difference in the curves is indeed statistically significant.
# -
# # Congratulations!
#
# You've completed the third assignment of Course 2. You've learned about the Kaplan Meier estimator, a fundamental non-parametric estimator in survival analysis. Next week we'll learn how to take into account patient covariates in our survival estimates!
| AI-Prognosis/Week3/C2M3_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1> Logistic Regression using Spark ML </h1>
#
# Set up bucket
BUCKET='data-science-on-gcp-bucket-2' # CHANGE ME
import os
os.environ['BUCKET'] = BUCKET
# +
# Create spark session
from __future__ import print_function
from pyspark.sql import SparkSession
from pyspark import SparkContext
sc = SparkContext('local', 'logistic')
spark = SparkSession \
.builder \
.appName("Logistic regression w/ Spark ML") \
.getOrCreate()
print(spark)
print(sc)
# -
from pyspark.mllib.classification import LogisticRegressionWithLBFGS
from pyspark.mllib.regression import LabeledPoint
# <h2> Read dataset </h2>
traindays = spark.read \
.option("header", "true") \
.csv('gs://{}/flights/trainday.csv'.format(BUCKET))
traindays.createOrReplaceTempView('traindays')
spark.sql("SELECT * from traindays LIMIT 5").show()
# +
from pyspark.sql.types import StringType, FloatType, StructType, StructField
header = 'FL_DATE,UNIQUE_CARRIER,AIRLINE_ID,CARRIER,FL_NUM,ORIGIN_AIRPORT_ID,ORIGIN_AIRPORT_SEQ_ID,ORIGIN_CITY_MARKET_ID,ORIGIN,DEST_AIRPORT_ID,DEST_AIRPORT_SEQ_ID,DEST_CITY_MARKET_ID,DEST,CRS_DEP_TIME,DEP_TIME,DEP_DELAY,TAXI_OUT,WHEELS_OFF,WHEELS_ON,TAXI_IN,CRS_ARR_TIME,ARR_TIME,ARR_DELAY,CANCELLED,CANCELLATION_CODE,DIVERTED,DISTANCE,DEP_AIRPORT_LAT,DEP_AIRPORT_LON,DEP_AIRPORT_TZOFFSET,ARR_AIRPORT_LAT,ARR_AIRPORT_LON,ARR_AIRPORT_TZOFFSET,EVENT,NOTIFY_TIME'
def get_structfield(colname):
if colname in ['ARR_DELAY', 'DEP_DELAY', 'DISTANCE', 'TAXI_OUT']:
return StructField(colname, FloatType(), True)
else:
return StructField(colname, StringType(), True)
schema = StructType([get_structfield(colname) for colname in header.split(',')])
# +
inputs = 'gs://{}/flights/tzcorr/all_flights-00000-*'.format(BUCKET) # 1/30th; you may have to change this to find a shard that has training data
#inputs = 'gs://{}/flights/tzcorr/all_flights-*'.format(BUCKET) # FULL
flights = spark.read\
.schema(schema)\
.csv(inputs)
# this view can now be queried ...
flights.createOrReplaceTempView('flights')
# -
# <h2> Clean up </h2>
trainquery = """
SELECT
f.*
FROM flights f
JOIN traindays t
ON f.FL_DATE == t.FL_DATE
WHERE
t.is_train_day == 'True'
"""
traindata = spark.sql(trainquery)
print(traindata.head(2)) # if this is empty, try changing the shard you are using.
traindata.describe().show()
# Note that the counts for the various columns are all different; We have to remove NULLs in the delay variables (these correspond to canceled or diverted flights).
# <h2> Logistic regression </h2>
trainquery = """
SELECT
DEP_DELAY, TAXI_OUT, ARR_DELAY, DISTANCE
FROM flights f
JOIN traindays t
ON f.FL_DATE == t.FL_DATE
WHERE
t.is_train_day == 'True' AND
f.dep_delay IS NOT NULL AND
f.arr_delay IS NOT NULL
"""
traindata = spark.sql(trainquery)
traindata.describe().show()
trainquery = """
SELECT
DEP_DELAY, TAXI_OUT, ARR_DELAY, DISTANCE
FROM flights f
JOIN traindays t
ON f.FL_DATE == t.FL_DATE
WHERE
t.is_train_day == 'True' AND
f.CANCELLED == '0.00' AND
f.DIVERTED == '0.00'
"""
traindata = spark.sql(trainquery)
traindata.describe().show()
def to_example(fields):
return LabeledPoint(\
float(fields['ARR_DELAY'] < 15), #ontime? \
[ \
fields['DEP_DELAY'], \
fields['TAXI_OUT'], \
fields['DISTANCE'], \
])
examples = traindata.rdd.map(to_example)
lrmodel = LogisticRegressionWithLBFGS.train(examples, intercept=True)
print(lrmodel.weights,lrmodel.intercept)
print(lrmodel.predict([6.0,12.0,594.0]))
print(lrmodel.predict([36.0,12.0,594.0]))
lrmodel.clearThreshold()
print(lrmodel.predict([6.0,12.0,594.0]))
print(lrmodel.predict([36.0,12.0,594.0]))
lrmodel.setThreshold(0.7) # cancel if prob-of-ontime < 0.7
print(lrmodel.predict([6.0,12.0,594.0]))
print(lrmodel.predict([36.0,12.0,594.0]))
# <h2> Predict with the model </h2>
#
# First save the model
# !gsutil -m rm -r gs://$BUCKET/flights/sparkmloutput/model
MODEL_FILE='gs://' + BUCKET + '/flights/sparkmloutput/model'
lrmodel.save(sc, MODEL_FILE)
print('{} saved'.format(MODEL_FILE))
lrmodel = 0
print(lrmodel)
# Now retrieve the model
from pyspark.mllib.classification import LogisticRegressionModel
lrmodel = LogisticRegressionModel.load(sc, MODEL_FILE)
lrmodel.setThreshold(0.7)
print(lrmodel.predict([36.0,12.0,594.0]))
print(lrmodel.predict([8.0,4.0,594.0]))
# <h2> Examine the model behavior </h2>
#
# For dep_delay=20 and taxiout=10, how does the distance affect prediction?
lrmodel.clearThreshold() # to make the model produce probabilities
print(lrmodel.predict([20, 10, 500]))
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
dist = np.arange(10, 2000, 10)
prob = [lrmodel.predict([20, 10, d]) for d in dist]
sns.set_style("whitegrid")
ax = plt.plot(dist, prob)
plt.xlabel('distance (miles)')
plt.ylabel('probability of ontime arrival')
delay = np.arange(-20, 60, 1)
prob = [lrmodel.predict([d, 10, 500]) for d in delay]
ax = plt.plot(delay, prob)
plt.xlabel('departure delay (minutes)')
plt.ylabel('probability of ontime arrival')
# <h2> Evaluate model </h2>
#
# Evaluate on the test data
inputs = 'gs://{}/flights/tzcorr/all_flights-00001-*'.format(BUCKET) # you may have to change this to find a shard that has test data
flights = spark.read\
.schema(schema)\
.csv(inputs)
flights.createOrReplaceTempView('flights')
testquery = trainquery.replace("t.is_train_day == 'True'","t.is_train_day == 'False'")
print(testquery)
testdata = spark.sql(testquery)
examples = testdata.rdd.map(to_example)
testdata.describe().show() # if this is empty, change the shard you are using
# +
def eval(labelpred):
cancel = labelpred.filter(lambda l: l[1] < 0.7)
nocancel = labelpred.filter(lambda l: l[1] >= 0.7)
corr_cancel = cancel.filter(lambda l: l[0] == int(l[1] >= 0.7)).count()
corr_nocancel = nocancel.filter(lambda l: l[0] == int(l[1] >= 0.7)).count()
cancel_denom = cancel.count()
nocancel_denom = nocancel.count()
if cancel_denom == 0:
cancel_denom = 1
if nocancel_denom == 0:
nocancel_denom = 1
return {'total_cancel': cancel.count(), \
'correct_cancel': float(corr_cancel)/cancel_denom, \
'total_noncancel': nocancel.count(), \
'correct_noncancel': float(corr_nocancel)/nocancel_denom \
}
# Evaluate model
lrmodel.clearThreshold() # so it returns probabilities
labelpred = examples.map(lambda p: (p.label, lrmodel.predict(p.features)))
print('All flights:')
print(eval(labelpred))
# keep only those examples near the decision threshold
print('Flights near decision threshold:')
labelpred = labelpred.filter(lambda l: l[1] > 0.65 and l[1] < 0.75)
print(eval(labelpred))
# -
# Copyright 2019 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| 07_sparkml_and_bqml/logistic_regression.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 📝 Exercise M1.01
# Imagine we are interested in predicting penguins species based on two of
# their body measurements: culmen length and culmen depth. First we want to do
# some data exploration to get a feel for the data.
#
# What are the features? What is the target?
# The data is located in `../datasets/penguins_classification.csv`, load it
# with `pandas` into a `DataFrame`.
# +
# Write your code here.
# -
# Show a few samples of the data
#
# How many features are numerical? How many features are categorical?
# +
# Write your code here.
# -
# What are the different penguins species available in the dataset and how many
# samples of each species are there? Hint: select the right column and use
# the [`value_counts`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.value_counts.html) method.
# +
# Write your code here.
# -
# Plot histograms for the numerical features
# +
# Write your code here.
# -
# Show features distribution for each class. Hint: use
# [`seaborn.pairplot`](https://seaborn.pydata.org/generated/seaborn.pairplot.html)
# +
# Write your code here.
# -
# Looking at these distributions, how hard do you think it will be to classify
# the penguins only using "culmen depth" and "culmen length"?
| notebooks/01_tabular_data_exploration_ex_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.3 64-bit (''.venv'': venv)'
# name: python3
# ---
# ## Area level model: empirical best linear unbiased predictor (EBLUP)
# Small area estimation (SAE) are useful techniques when the sample sizes are not sufficient to provide reliable direct domain estimates given the sampling design. In this tutorial, the direct estimates refer to estimates obtained from the design-based approach. It usually consists of applying adjusted design weights to the variable of interest to compute sample parameters as estimates of equivalent population parameters. When auxiliary information is available, we can use model assisted survey methods can be used to estimate population parameters.
#
# In this tutorial, we will go futher and use modeling techniques to produce domains estimates. For the area level model, the modeling is done at the area level using generalized linear mixed models. The sections below shows how to use the *EblupAreaModel* class from the *samplics* package to produce area level estimates.
# ### Milk Expenditure data
#
# To illustrate the EblupAreaModel class, we will use the Milk Expenditure dataset used in Rao and Molina (2015). As mentioned in the book, this dataset was originally used by Arora and Lahiri (1997) and later by You and Chapman (2006). For the R users, this dataset is also used by the R package sae (https://cran.r-project.org/web/packages/sae/index.html).
#
# The Milk Expenditure data contains 43 observations on the average expenditure on fresh milk for the year 1989. The datasets has the following values: major area representing (major_area), small area (small_area), sample size (samp_size), direct survey estimates of average expenditure (direct_est), standard error of the direct estimate (std_error), and coefficient of variation of the direct estimates (coef_variance).
# +
import numpy as np
import pandas as pd
import samplics
from samplics.datasets import load_expenditure_milk
from samplics.sae import EblupAreaModel
# +
# Load Expenditure on Milk sample data
milk_exp_dict = load_expenditure_milk()
milk_exp = milk_exp_dict["data"]
nb_obs = 15
print(f"\nFirst {nb_obs} observations of the Milk Expendure dataset\n")
milk_exp.tail(nb_obs)
# -
# ### EBLUP Predictor
#
# As shown in the milk expenditure datasets, some of the coefficients of variation are not small which indicates unstability of the direct survey estimates. Hence, we can try to reduce the variability of the estimates by smoothing them through modeling. For illustration purpose, we will model the average expenditure on milk using the major areas as auxiliary variables.
#
# First, we use the method *fit()* to estimate the model parameters. The pandas's method *get_dummies()* create a matrix with dummy values (0 and 1) from the categorical variable *major_area*.
# +
area = milk_exp["small_area"]
yhat = milk_exp["direct_est"]
X = pd.get_dummies(milk_exp["major_area"],drop_first=True)
sigma_e = milk_exp["std_error"]
## REML method
fh_model_reml = EblupAreaModel(method="REML")
fh_model_reml.fit(
yhat=yhat, X=X, area=area, error_std=sigma_e, intercept=True, tol=1e-8,
)
print(f"\nThe estimated fixed effects are: {fh_model_reml.fixed_effects}")
print(f"\nThe estimated standard error of the area random effects is: {fh_model_reml.re_std}")
print(f"\nThe convergence statistics are: {fh_model_reml.convergence}")
print(f"\nThe goodness of fit statistics are: {fh_model_reml.goodness}\n")
# -
# Now the the model has been fitted, we can obtain the EBLUP average expenditure on milk by running *predict()* which is a method of *EblupAreaModel* class. This run will produce two main attributes that is *area_est* and *area_mse* which are python dictionaries pairing the small areas to the eblup estimates and the MSE estimates, respectively.
# +
fh_model_reml.predict(
X=X, area=area, intercept=True
)
import pprint
pprint.pprint(fh_model_reml.area_est)
# -
# We can use the utility method *to_dataframe()* to output the estimates as a pandas dataframe. The function provides the area, the estimate and its MSE estimates. We can use *col_names* to customize the name of the columns. For example, using `col_names = ["small_area", "eblup_estimate", "eblup_mse"]`. Otherwise, if col_names is not provided, "_area", "_estimates" and "_mse" are used as defaults.
milk_est_reml = fh_model_reml.to_dataframe(col_names = ["parameter", "small_area", "eblup_estimate", "eblup_mse"])
print(f"\nThe dataframe version of the area level estimates:\n\n {milk_est_reml}")
# We could also fit the model parameters using the maximum likelihood (ML) method which will impact the MSE estimation as well. To estimate the area means using the ML methdo, we only need to set *method="ML"* then run the prediction as follows.
# +
## ML method
fh_model_ml = EblupAreaModel(method="ML")
fh_model_ml.fit(
yhat=yhat, X=X, area=area, error_std=sigma_e, intercept=True, tol=1e-8,
)
milk_est_ml = fh_model_ml.predict(
X=X, area=area, intercept=True
)
milk_est_ml = fh_model_ml.to_dataframe(col_names = ["parameter", "small_area", "eblup_estimate", "eblup_mse"])
print(f"\nThe dataframe version of the ML area level estimates:\n\n {milk_est_ml}")
# -
# Similar, we can use the Fay-Herriot method as follows
# +
## FH method
fh_model_fh = EblupAreaModel(method="FH")
fh_model_fh.fit(
yhat=yhat, X=X, area=area, error_std=sigma_e, intercept=True, tol=1e-8,
)
milk_est_fh = fh_model_fh.predict(
X=X, area=area, intercept=True
)
milk_est_fh = fh_model_fh.to_dataframe(col_names = ["parameter", "small_area", "eblup_estimate", "eblup_mse"])
print(f"\nThe dataframe version of the ML area level estimates:\n\n {milk_est_fh}")
| docs/source/tutorial/eblup_area_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Visualization with Haberman Dataset
# # Haberman's Survival Dataset
# Dataset contains cases from study conducted on the survival of patients who had undergone surgery for breast cancer in University of Chicago's Billings Hospital for the duration of 1958 to 1970
#
# .Number of Instances: 306
# .Number of Attributes: 4 (including the class attribute)
# .Attribute Information:
# a) Age of patient at time of operation (numeric)
# b) Patient's year of operation (year - 1900, numerical)
# c) Number of positive axillary nodes detected (numerical)
# d) Survival status (class attribute)
# 1 = the patient survived 5 years or longer
# 2 = the patient died within 5 year
# Dataset Source : https://www.kaggle.com/gilsousa/habermans-survival-data-set/data
# +
import pandas as pd
import seaborn as sbn
import matplotlib.pyplot as plt
import numpy as np
haberman=pd.read_csv("haberman.csv")
# -
haberman.info()
'''Dataset contains 305 rows and 4 columns; columns name are missing in the dataset.
Add the column name, given in the kaggle column metadata tab.'''
haberman.columns=['Age','Operation Year','Auxillary Node','Survival Status']
print(haberman.columns)
haberman.head()
haberman["Survival Status"].value_counts()
# ### Observation
# 1. Imbalanced dataset.
# 2. Post operation out of the 305 patients; 224 patient survived for more than 5 years and 81 patients died within 5 years.
# ## 2-D Scatter Plot :
haberman.plot(kind='scatter',x='Age',y='Auxillary Node')
plt.show()
# 2-D Scatter plot with color-coding for each survival status type/class.
sbn.set_style("whitegrid");
sbn.FacetGrid(haberman, hue="Survival Status", size=6) \
.map(plt.scatter, "Age", "Auxillary Node") \
.add_legend();
plt.show();
# ### Observation(s):
# 1. A good count of patients have 0 Auxillary Nodes.
# ## Pair Plot :
plt.close();
sbn.set_style("whitegrid")
sbn.pairplot(haberman,hue="Survival Status",size=3)
plt.show()
# ### Observation(s):
# 1. "Age vs Auxillary Node" tells us most of the patients have 0 Auxillary Node post operation.
# 2. Most of the plots exhibits overlapping.
# ## HISTOGRAM, PDF :
sbn.FacetGrid(haberman, hue="Survival Status", size=5) \
.map(sbn.distplot, "Auxillary Node") \
.add_legend();
plt.show();
# ### Observation(s) :
# Patients with (<=3) number of axillary nodes have more chance of survival with maximum of patients having 0 Auxillary node.
sbn.FacetGrid(haberman, hue="Survival Status", size=5) \
.map(sbn.distplot, "Age") \
.add_legend();
plt.show();
# ### Observation(s):
# Patients with age of <= 40 have more chances of survival than others.
#
sbn.FacetGrid(haberman, hue="Survival Status", size=5) \
.map(sbn.distplot, "Operation Year") \
.add_legend();
plt.show();
# ### Observation(s):
# Here the histogram shows the year of operation for each patient and it is completely overlapped for both survived and non survived patients.
# Hence, year of operation hardly depends on the survival of patient.
# But, a lot of patients operated for the year 1965 died.
# ## CDF
SurvivedOp=haberman.loc[haberman["Survival Status"]==1]
NotSurvived=haberman.loc[haberman["Survival Status"]==2]
counts, bin_edges = np.histogram(SurvivedOp['Auxillary Node'], bins=20,
density = True)
##PDF
pdf = counts/(sum(counts))
print(pdf);
print(bin_edges)
##CDF
cdf = np.cumsum(pdf)
plt.plot(bin_edges[1:],pdf)
plt.plot(bin_edges[1:], cdf)
plt.legend(['Pdf Survived',
'Cdf Survived'])
plt.show()
counts, bin_edges = np.histogram(NotSurvived['Auxillary Node'], bins=20,
density = True)
##PDF
pdf = counts/(sum(counts))
print(pdf);
print(bin_edges)
##CDF
cdf = np.cumsum(pdf)
plt.plot(bin_edges[1:],pdf)
plt.plot(bin_edges[1:], cdf)
plt.legend(['Pdf Died',
'Cdf Died'])
plt.show()
# ## Summary Stats :
SurvivedOp.describe()
NotSurvived.describe()
from statsmodels import robust
print("\n Median Absolute Deviation")
print(robust.mad(SurvivedOp['Auxillary Node']))
print(robust.mad(NotSurvived['Auxillary Node']))
# ### Observation(s):
# 1. Here we can observe that taking auxillary nodes into considerations can yield better result as its standard deviation and mean far apart as compared to that of Age and Operation year.
# 2. The people who are not survived tend to have more average number of Auxillary nodes and more spread out the distribution than survived.
# ## Box Plots and Whiskers
sbn.boxplot(x='Survival Status',y='Age', data=haberman)
plt.show()
sbn.boxplot(x='Survival Status',y='Auxillary Node', data=haberman)
plt.show()
sbn.boxplot(x='Survival Status',y='Operation Year', data=haberman)
plt.show()
# ### Observation(s):
# 1. From box plot number of people dead between age of 46-62.
# 2. Number of people survived between age of 42-60.
# ## Violin Plot
sbn.violinplot(x='Survival Status',y='Auxillary Node', data=haberman)
plt.show()
sbn.violinplot(x='Survival Status',y='Age', data=haberman)
plt.show()
sbn.violinplot(x='Survival Status',y='Operation Year', data=haberman)
plt.show()
# ### Observation(s):
# 1. From violin plot number of people dead between age of 59-65
# 2. Number of people survived between age of 60-66
# # Contour Plot
sbn.jointplot(x="Age", y="Operation Year", data=haberman, kind="kde");
plt.show();
# ### Observation(s):
# There are more no of people undergone operation during the year 1959-1964 and between age 42-59.
| HabermanSurvival-BreastCancer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:miniconda3-python-tutorial]
# language: python
# name: conda-env-miniconda3-python-tutorial-py
# ---
# ## Sample Script for the interpolation of variables using ``02_variable_interpolator`` module
#
# Import the module.
exec(open('/glade/u/home/molina/python_scripts/deep-conus/deep-conus/02_variable_interpolator.py').read())
# import project_code from config file (keeping private from public repo)
from config import project_code
# Create the InterpolateVariable object using ``InterpolateVariable`` class.
#
# Example below interpolates the ``TK`` (air temperature) variable on 1, 3, 5, and 7-km above ground level (AGL) height for November 2000 for the current climate period.
test = InterpolateVariable(climate='current',
variable='TK',
month_start=11, month_end=12, year_start=2000, year_end=2000,
destination='/glade/scratch/molina/DL_proj/current_conus_fields/TK/',
start_dask=True,
project_code=project_code,
cluster_min=5, cluster_max=40)
# ``create_the_interp_files()`` generates the TK files on 1, 3, 5, and 7-km AGL and saves them to the destination path provided. Make sure it is for the correct folder!
# +
#test.create_the_interp_files()
# -
#
#
# The maximum vertical velocity data can also be created with ``02_variable_interpolator``.
#
# ``create_the_max_files()`` generates the maximum vertical wind velocity (``w``) data and saves them to the destination path provided.
#
# Make sure it saved in the correct folder!
# +
test = InterpolateVariable(climate='future',
variable='MAXW',
month_start=10, month_end=13, year_start=2000, year_end=2000,
destination='/glade/scratch/molina/DL_proj/future_conus_fields/W_vert/',
start_dask=False)
#test.create_the_max_files()
# -
| notebooks/02_sample_nb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: mozilla_tts
# language: python
# name: mozilla_tts
# ---
# %reload_ext autoreload
# %autoreload 2
# +
import sys
sys.path.append('..')
import torch
from model import Decoder
from model import Attention
# -
attn = Attention(query_dim=256, embedding_dim=256, attention_dim=128)
attn
query = torch.rand(32, 256)
inputs = torch.rand(32, 71, 256)
processed_inputs = torch.rand(32, 71, 128)
mask = torch.zeros([32, 71], dtype=torch.bool)
attn.init_states(inputs)
context = attn(query, inputs, processed_inputs, mask)
context.shape
decoder = Decoder(in_features=256, memory_dim=80, r=7)
decoder
inputs = torch.rand(32, 71, 256) # encoderの出力
memory = torch.rand(32, 231, 80) # decoderへ入力するメルスペクトログラム
mask = torch.zeros([32, 71], dtype=torch.bool)
mel_outputs, alignments, stop_tokens = decoder(inputs, memory, mask)
mel_outputs.shape
alignments.shape
stop_tokens.shape
| notebooks/decoder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Do not delete any row from this dataset
#
# ### Write df = df.fillna(0) instead
# ## Q1. Total number of people vaccinated in Argentina
# ## Q2. Total number of countries in this dataset
# ## Q3. Maximum number of people vaccinated in a country
# ## Q4. Top 5 Countries in terms of number of people vaccinated
| COVID Data Analysis Assignment/COVID-19 Vaccination Analysis - Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Chiraagkv/ASL/blob/main/holdmybeer.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="4F-F2kWc35U3"
import tensorflow_hub as hub
import numpy as np
import pandas as pd
import tensorflow as tf
import os
import random as rd
from PIL import Image
import PIL
# + colab={"base_uri": "https://localhost:8080/"} id="0cky4rmP4hiq" outputId="a13eb685-af9a-499a-ac6a-0daf7b858157"
# !mkdir /root/.kaggle/
# !touch /root/.kaggle/kaggle.json
# !echo '{"username":"kingrohitkumar","key":"734c20a118d1339e94b4192d5fe26313"}' > /root/.kaggle/kaggle.json
# !chmod 600 /root/.kaggle/kaggle.json
# !kaggle datasets download -d grassknoted/asl-alphabet --unzip
# + colab={"base_uri": "https://localhost:8080/"} id="LB-yBHKy4lWw" outputId="881a4c70-407a-406e-c2d0-11fdd0460353"
base_model=hub.KerasLayer(handle='https://tfhub.dev/tensorflow/efficientnet/b7/feature-vector/1', trainable=False, weights=None)
data=tf.keras.preprocessing.image.ImageDataGenerator(rescale=1/255,
samplewise_std_normalization=True,
height_shift_range=0.2,
width_shift_range=0.2,
rotation_range=0.2,
zca_epsilon=1e-6,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
cval=0.0)
data=data.flow_from_directory(directory='/content/sampled_image_path_main',
target_size=(224, 224),
class_mode='categorical',
batch_size=19,
shuffle=False,
interpolation='bilinear')
augmentation=tf.keras.Sequential([
tf.keras.layers.experimental.preprocessing.RandomCrop(2, 2),
tf.keras.layers.experimental.preprocessing.RandomContrast(0.2)
])
# + id="l70dGqw046r3"
def randoms_images (image_path, duplicate_path):
for (roots, dirs, filenames) in os.walk(image_path):
for i in range (len(dirs)):
os.makedirs(duplicate_path+'/'+dirs[i])
cwd=image_path+'/'+dirs[i]
for (root, dir, files) in os.walk(cwd):
for m in range (300):
image_name=files[m]
img_path=Image.open(cwd+'/'+image_name)
du=image_name.split('.')
dup_path=duplicate_path+'/'+dirs[i]+'/'+f"{du[0]}.jpeg"
img_path.save(dup_path)
continue
continue
continue
randoms_images ('/content/asl_alphabet_train/asl_alphabet_train', 'sampled_image_path_main')
# + id="jqYIGuzB5D1C"
inputs=tf.keras.layers.Input(shape=(224, 224, 3))
layer_1=augmentation(inputs)
model=base_model(inputs, training=True)
x2=tf.keras.layers.GlobalAveragePooling2D()(tf.expand_dims(tf.expand_dims(model, axis=0), axis=0))
outputs=tf.keras.layers.Dense(len(os.listdir('asl_alphabet_train')), activation='softmax')(x2)
model=tf.keras.Model(inputs, outputs)
model.compile(loss=tf.keras.losses.categorical_crossentropy,
optimizer=tf.keras.optimizers.Adam(),
metrics='accuracy')
model.fit(data, epochs=1)
# + id="f3CkYSLq5iMP"
| progress_files/holdmybeer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Using SAS DLPy to Create a Neural Network Model for Binary Classification Using Tabular Data
# You can use SAS DLPy to easily create and train a variety of basic task-oriented deep learning models. This notebook example shows how you can use SAS DLPy to build a neural network model for a binary classification task using tabular data.
#
# The clean tabular data used to create the model is the [Scikit-Learn breast cancer data set](https://scikit-learn.org/stable/datasets/toy_dataset.html#breast-cancer-wisconsin-diagnostic-dataset). The breast cancer data is a toy data set, a 569-observation table of 30 cellular attribute measurements from the [UCI ML Breast Cancer Wisconsin (Diagnostic) data sets](https://goo.gl/U2Uwz2). The Wisconsin diagnostic breast cancer data is also available to the public via the University of Wisconsin CS ftp server at `ftp.cs.wisc.edu cd math-prog/cpo-dataset/machine-learn/WDBC/`.
#
# Each anonymized observation in the diagnostic tabular data contains 30 attribute measurements that were computed from digitized images of fine needle aspirates (FNA) of individual breast masses. Each attribute describes characteristics of the cell nuclei present in the image. The two target classes for each observation containing cellular attribute observations are `benign` (target=1) and `malignant` (target=0). More technical information about the composition and use of the Scikit-learn breast cancer data [is available here](https://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_breast_cancer.html#).
#
# The example in this notebook follows these steps:
#
# - Configure SAS DLPy and SAS CAS environments for this task.
# - Upload the toy diagnostic data set to the SAS CAS server.
# - Create train and test data sets in SAS CAS.
# - Use SAS DLPy to build a fully-connected neural network classification model.
# - Train the fully-connected neural network model.
# - Use the trained model to score the test data set.
# - Evaluate model performance using a series of analytic model plots.
#
# This tutorial example assumes that you have SAS DLPy API installed, have an active SAS CAS server, and have installed the common Python utilities used in the code (numpy, sklearn, matplotlib, Pandas).
# ### Table of Contents
# - [Prepare Resources and Configure Environment for Modeling](#getReady)
# - [Import Required Python and SAS DLPy Utilities](#importPythonDLPy)
# - [Configure SAS SWAT and Launch SAS CAS](#launchCAS)
# - [Load the Example Data](#loadData)
# - [Partition into Train and Test Data](#partitionData)
# - [Use SAS DLPy to Create a Neural Network Model](#createModel)
# - [Train the Neural Network Classification Model](#trainModel)
# - [Score the Test Data Using Trained Neural Network Model](#scoreModel)
# - [Examine Neural Network Classification Model Performance Plots](#Plots)
# - [Evaluate the Confusion Matrix](#confusionMatrix)
# - [Calculate the Model's Predictive Accuracy](#predAccuracy)
# - [Assess ROC Curve and Calculate AUC](#rocAUC)
# - [Evaluate Precision-Recall Curve](#precisionRecall)
# - [Evaluate Average Precision and F1 Scores](#f1Average)
# - [Summary](#summary)
#
# <a id = "getReady"></a>
#
# ### Prepare Resources and Configure Computing Environment for Modeling
#
# Use this section to organize all of the resources you need and configure your local computing environment in advance, so you can follow along with the example notebook modeling operations without interruption.
# <a id="importPythonDLPy"></a>
#
# #### Import Required Python and SAS DLPy Utilities
#
# Use this section to import the Python utility libraries and SAS DLPy libraries that will be used for the fully-connected feedforward neural network classification model.
#
# Import the Python and SAS DLPy utilities required for the model task. They include the scikit-Learn utility, the pandas data analysis library, the numPy scientific computing library, and the matplotlib plotting utility.
# +
# Pandas data analysis utility
import pandas as pd
# NumPy scientific computing utility
import numpy as np
# Import scikit-Learn utility
import sklearn
# Matplotlib plotting utility
import matplotlib.pylab as plt
# SAS DLPy model utilities
from dlpy import Model, Sequential
from dlpy.model import Optimizer, AdamSolver
from dlpy.layers import *
from dlpy.splitting import two_way_split
from dlpy.metrics import *
# Display output plots in notebook cells
# %matplotlib inline
# -
# <a id="launchCAS"></a>
#
# ### Configure SAS SWAT and Launch SAS CAS
#
# The following code configures SAS SWAT and launches SAS CAS. SWAT is a Python interface to SAS CAS that enables you to load data into memory and apply CAS actions to the data.
#
# <b>Note:</b> For more information about starting a CAS session with the SWAT package, see https://sassoftware.github.io/python-swat/getting-started.html.
# +
# Import SAS SWAT
from swat import *
# SWAT data message handler
import swat.cas.datamsghandlers as dmh
# Configure CAS session for Analytics
s = CAS('your-host-name.unx.company-name.com', 5570)
# -
# <a id="loadData"></a>
#
# ### Load the Example Data
#
# Use the code block below to import the toy diagnostic data set, create pandas predictor and target dataframes, and then merge the two dataframes. Browse the concatenated pandas table before loading it into SAS CAS.
# +
# Import toy diagnostic breast cancer
# data from the sci-kit Learn library
# (sklearn package version >= 0.24.1)
from sklearn.datasets import load_breast_cancer
# load the breast cancer data into SAS CAS and
# save the CAS table as bc_data.
bc_data = load_breast_cancer()
# Put the bc_data contents into two pandas dataframes:
# One dataframe for the predictors (bc_x_df).
bc_x_df = pd.DataFrame(bc_data['data'], columns=bc_data['feature_names'])
# One dataframe for the target (bc_y_df).
bc_y_df = pd.DataFrame(bc_data['target'], columns=['target_class'])
# Concatenate the predictor and target dataframes
# into one dataframe with 30 feature columns and
# 1 target column.
bc_df = pd.concat([bc_x_df, bc_y_df], axis=1)
# -
# Browse the concatenated pandas diagnostic data.
# Display column headings and 10 rows from the table.
bc_df.loc[:9]
# In the table above, the rightmost column `target_class` represents the observation's classification status: a value of 1 indicates a `benign` tumor, and a value of 0 indicates a `malignant` tumor. What is the distribution of `malignant` and `benign` classes in the diagnostic data table of 569 observations?
#
# Use the `target_class.value_counts()` function to count the two class values in table `bc_df`:
# Show distribution of target values
# for benign and malignant classes
# in the toy diagnostic data.
bc_df.target_class.value_counts()
# The output shows that 357 of the observations in the table `bc_df` indicate benign tumors (target = '1' in the data), and 212 of the observations indicate malignant tumors (target = '0' in the data).
#
# Use the `upload_frame()` function with the concatenated table `bc_df` to load the pandas dataframe to the server. The CAS table `bc_df` is loaded and saved in Python as `big_tbl`.
# Load the concatenated data frame bc_df
# to the CAS server and save as big_tbl.
big_tbl = s.upload_frame(bc_df,
casout=dict(name='bc_df',
replace=True
)
)
# Show CAS session tables
s.table.tableInfo()
# <a id="partitionData"></a>
#
# ### Partition into Train and Test Data
#
# Split the loaded diagnostic table `big_tbl` into train and test partitions. Use the DLPy `two_way_split` function to perform random server-side sampling and partition the data into `train_tbl` and `test_tbl`. Server-side partitioning is preferable to client-side partitioning when the data is large, or when the data is already on the server.
#
# The code below partitions 70% of `big_tbl` into `train_tbl`, and the remaining 30% is partitioned as `test_tbl`. The `seed` value is specified to ensure repeatability of example computation results.
# Partition big_tbl into train and test data sets
train_tbl, test_tbl = two_way_split(big_tbl,
# 30% test partition
test_rate=30,
seed=5309,
# random sampling without stratification
stratify=False,
# using tabular numeric data,
# not an image table
im_table=False)
# Display five rows from the train table.
train_tbl.head()
# Display five rows from the test table.
test_tbl.head()
# <a id="createModel"></a>
#
# ### Use SAS DLPy to Create a Fully-Connected Neural Network Model
#
# Use SAS DLPy to create a fully-connected feed-forward neural network binary classification model with the following architecture:
#
# - one dense layer
# - 20 neurons
# - `relu` activation layer
# - `softmax` output layer activation function
# - `entropy` output layer loss function
# - 2 neurons (classes) in the output layer.
#
# Save the newly created model in Python as `model1`, and save the model table in SAS CAS as `simple_dnn_classifier`.
# +
# Create fully-connected deep neural network 'model1'
model1 = Sequential(s,
model_table=CASTable('simple_dnn_classifier',
replace=True
)
)
# Input layer
model1.add(InputLayer(std='STD'))
# Dense layer
model1.add(Dense(20, act='relu'))
# Output layer
model1.add(OutputLayer(act='softmax',
n=2,
error='entropy'
)
)
# -
# As an additional exercise, uncomment the bottom line of the code below, and run the cell to generate a DAG of `model1`.
# +
# Generate a DAG for the simple
# neural network classification model
#model1.plot_network()
# -
# The generated DAG for the simple neural network classification model should resemble the following:
#
# 
# <a id="trainModel"></a>
#
# ### Train the Neural Network Classification Model
#
# Now use `fit()` with `model1` to train the fully-connected neural network model using a scheduled training approach with the following hyperparameter specifications:
#
# - `AdamSolver` optimizer to show a scheduled training approach
# - `learning_rate_policy='step'` to specify a stepwise learning rate decreasing policy
# - `step_size=5` to multiply the `learning_rate` by factor `gamma=0.9` every 5 epochs
# - `log_level=2` to view the training log
# - `seed=5309` to support model determinism and repeatable results.
#
# Save the trained model table in Python as `result`.
# +
# Specify optimizer settings for scheduled training approach
# These optimizer settings are called in the fit() block that
# follows.
optimizer = Optimizer(algorithm=AdamSolver(learning_rate=0.005,
learning_rate_policy='step',
gamma=0.9,
step_size=5),
mini_batch_size=4,
seed=5309,
max_epochs=50,
log_level=2)
# Train the model model1 using
# data train_tbl, and save the
# trained output table as 'result'
result = model1.fit(train_tbl,
inputs=bc_x_df.columns.tolist(),
nominals=['target_class'],
target='target_class',
optimizer=optimizer
)
# -
# Use the matplotlib function `plot_training_history()` with `model1` to display a graph of the model's training over 50 epochs:
# Plot the training history of 'model1' to
# display the changing model loss and fit
# error values over 50 epochs.
model1.plot_training_history()
# <a id="scoreModel"></a>
#
# ### Score the Test Data Using Trained Neural Network Model
#
# Now use the `predict()` function for `model1` with `test_tbl` to score the test data. Save the scored data in `model1.valid_res_tbl` in Python as `test_result_table`.
# Use model1 to score the test data test_tbl.
test_result = model1.predict(test_tbl)
# Save the scored data table as test_result_table.
test_result_table = model1.valid_res_tbl
# Use the `head()` function to show the first five rows of the scored `test_result_table`. The rightmost column `_DL_PredLevel` contains the scored binary predictions for each observation: 0.0 (malignant) or 1.0 (benign).
# Display the first five rows of
# the scored test data that was
# saved as test_result_table.
test_result_table.head()
# Note that the rightmost column in the scored test table is the predicted binary target classifier, `_DL_PredLevel_`.
# <a id="Plots"></a>
#
# ### Examine Neural Network Classification Model Performance Plots
#
# Now we can use a variety of scored model metrics to assess the performance of the neural network binary classification model `model1`. In this section we use the scored data to calculate the model confusion matrix, the model accuracy score, the ROC curve, the area under ROC curve (AUC), the Precision-Recall curve, and the model's F1 score.
# <a id="confusionMatrix"></a>
#
# #### Evaluate the Confusion Matrix
#
# First, show the confusion matrix for the model1 scored data in test_result_table. The confusion matrix is a performance measurement for deep learning classification models. The confusion matrix for a binary classification problem is a 2 x 2 table that shows a summary of the model's predicted values versus actual ground truth values.
# Show the confusion matrix for the scored table
display(confusion_matrix(test_result_table['target_class'],
test_result_table['I_target_class']
)
)
# The row headings `0.0` and `1.0` represent the target classes `malignant` and `benign`. The confusion matrix shows that out of the scored data, there were 59 true positives, and 3 false positives for target class 0 `malignant`. There were also 2 false negatives, and 107 true negatives for target class 1 `benign`. In other words, out of 171 scored test observations, 166 were correctly classified, and 5 were incorrectly classified.
# <a id="predAccuracy"></a>
#
# #### Calculate the Model's Predictive Accuracy
# Based on the test data set score, what is the model's predictive accuracy? Use the `accuracy_score()` function to create the output table `acc_score`, and use the `print()` function to display it.
#
# In multilabel classification, the accuracy function computes subset accuracy for predicted and ground truth labels. Accuracy scores range from 0 to 1. Model accuracy score values that are very close to 1 indicate higher model accuracy.
# Calculate the accuracy score
acc_score = accuracy_score(test_result_table['target_class'],
test_result_table['I_target_class']
)
print('The accuracy score for the test data is {:.6f}.'.format(acc_score))
# <a id="rocAUC"></a>
#
# #### Assess ROC Curve and Calculate AUC
#
# As another performance measure, use the scored test results in `test_result_table` with the `plot_roc()` function to generate a plot of the ROC curve for the `benign` target class where `target_class=1`. The ROC curve is a graphical plot that illustrates the performance of a binary classification system as its discrimination threshold is varied.
# Plot the ROC curve for target_class = 1.
plot_roc(test_result_table['target_class'],
test_result_table['P_target_class1'],
pos_label=1,
figsize=(6,6),
linewidth=2
)
# The ROC curve rises aggressively towards the ideal predictor's perfect true positive rate of (1,0). This is a strong ROC curve.
#
# Now use the SAS DLPy function `roc_auc_score()` with the scored test table `test_result_table` with `P_target_class1` to calculate the AUC, the area under the ROC curve.
#
# AUC, or the area under the ROC curve, is a performance measurement for the classification model at various threshold settings. The ROC is a probability curve. The AUC represents the degree of separability. AUC indicates how well the model performs in distinguishing between classes. The score is a value between 0.0 and 1.0, with a value of 1.0 being a perfect classifier. The closer to 1 the AUC score is, the better the model performs at predicting 0s as 0s and 1s as 1s.
# Calculate the AUC, or Area under the ROC curve.
# Save the resulting Python table as auc_of_roc.
auc_of_roc = roc_auc_score(test_result_table['target_class'],
test_result_table['P_target_class1'],
pos_label=1
)
print('The AUC area under the ROC curve for the scored test data is {:.6f}.'.format(auc_of_roc))
# <a id="precisionRecall"></a>
#
# #### Evaluate Precision-Recall Curve
#
# Now use the function `plot_precision_recall()` with the scored test table `test_result_table` and `P_target_class1` to generate the Precision-Recall curve for target class 1.
#
# Precision is a metric that quantifies the number of correct positive predictions that were made. It is calculated as the number of true positives divided by the total number of true positives and false positives. The result is a value between 0.0 for no precision and 1.0 for full or perfect precision.
#
# Recall is a metric that describes the number of correct positive predictions made out of all positive predictions that could have been made. It is calculated as the number of true positives divided by the total number of true positives and false negatives. In other words, recall represents the true positive rate. The result is a value between 0.0 for no recall and 1.0 for perfect recall.
#
# A Precision-Recall curve is a plot of the precision (y-axis) and the recall (x-axis) for different probability thresholds.
# Plot the precision recall curve for target_class = 1.
plot_precision_recall(test_result_table['target_class'],
test_result_table['P_target_class1'],
pos_label=1,
figsize=(6,6),
linewidth=2
)
# The plotted precision-recall curve bows heavily towards the perfect skill coordinate at (1,1). A classification model with poor or no skill would be a horizontal line on the plot. This is a good precision-recall curve for the model.
#
# <a id="f1Average"></a>
#
# #### Evaluate Average Precision Score and F1 Score
#
# Now use the function `average_precision_score()` with the scored test table `test_result_table` and `P_target_class1` to calculate the model's average precision score for target class 1. For a binary classification task, average precision summarizes a Precision-Recall curve as the weighted mean of precisions achieved at each threshold. The increase in recall from the previous threshold becomes the weight. Average precision scores range from 0 to 1. The closer the score is to 1, the better the precision of the model.
# Calculate the average precision score
ap = average_precision_score(test_result_table['target_class'],
test_result_table['P_target_class1'],
pos_label=1
)
print('The average precision score for the scored test data is {:.6f}.'.format(ap))
# The average precision score for this binary classification model is 0.995637, indicating strong model precision.
#
# Finally, use the function `f1_score()` with the scored test table `test_result_table` and `I_target_class` to calculate the F1 score for target class 1. The F1 score represents a weighted average of the scoring model's precision and recall. The best score for an F1 value is 1, and the worst score is 0. The influences of precision and recall on the f1 score are equal.
#
# The closer the F1 score is to 1.0, the greater the model performance.
# Calculate the F1 score
f1sc = f1_score(test_result_table['target_class'],
test_result_table['I_target_class'],
pos_label=1
)
print('The F1 score for the scored test data is {:.6f}.'.format(f1sc))
# The F1 score for the neural network binary classification model is 0.977169, very close to 1.0. This is a high f1 score, indicative of a high-performing predictive classification model.
# <a id="summary"></a>
#
# ### Summary
#
# It is relatively easy to create and modify task-centric models using SAS DLPy. This example showed how to use SAS DLPy to create a fully-connected feedforward neural network model for a binary classification task using tabular data. It showed how to use DLPy to score data with a trained model, and it also showed how to use DLPy to generate and view multiple performance metrics (ROC, AUC, Precision-Recall curve, Average Precision score, and F1 score) using scored data table output from the trained neural network model.
| examples/tabular_data_analysis/Two_Class_Classification_with_Fully_Connected_Neural_Network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import scipy
from sklearn import preprocessing
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
from sklearn import model_selection
from sklearn.metrics import classification_report, accuracy_score
from pandas.plotting import scatter_matrix
# ## Load the data
# ### Breast Cancer Wisconsin (Original) Data Set
# The data can be doanloaded from the url: https://archive.ics.uci.edu/ml/machine-learning-databases/breast-cancer-wisconsin/breast-cancer-wisconsin.dat
# +
# loading the data
url = 'https://archive.ics.uci.edu/ml/machine-learning-databases/breast-cancer-wisconsin/breast-cancer-wisconsin.data'
names = ['id', 'clump_thickness', 'uniform_cell_size', 'uniform_cell_shape', 'marginal_adhision',
'single_capitheleial_size', 'bare_nuclei', 'bland_chromatin',
'normal_nucleoli', 'mitosis', 'class']
df =pd.read_csv(url, names=names)
df.info()
# -
# ## Data Exploration
# +
# get the rows with non numeric values
num_filter = df['bare_nuclei'].str.isnumeric().tolist()
count = 0
index = []
for i, val in enumerate(num_filter):
if val == False:
index.append(i)
count = count+1
print('Number of non numeric rows: {}'.format(count))
print('Indices of non numeric rows: {}'.format(index))
# -
# print rows with non muneric values
df.loc[index].head(5)
df['bare_nuclei'].replace('?', np.NaN, inplace=True)
df['bare_nuclei'] = df['bare_nuclei'].astype(float)
df = df.fillna(df.mean())
df.drop(['id'], 1, inplace=True)
print(df.info())
sns.boxenplot(df.mitosis)
# histogram of the dataset
df.hist(figsize=(10, 10))
plt.show()
# creat scatter plot matrix
scatter_matrix(df, figsize=(18, 18))
plt.show()
# Create X and Y datasets
X = np.array(df.drop(['class'], 1))
Y = np.array(df['class'])
X_train, X_test, Y_train, Y_test = model_selection.train_test_split(X, Y, test_size = 0.2)
# Specify testing options
seed = 8
scoring = 'accuracy'
# +
# Define the models to train
models = []
models.append(('KNN', KNeighborsClassifier(n_neighbors=5)))
models.append(('SVM', SVC()))
# Evaluate each model
results = []
names = []
for name, model in models:
print('Method: {}'.format(name))
kfold = model_selection.KFold(n_splits=10, random_state=seed, shuffle=True)
cv_results = model_selection.cross_val_score(model, X_train, Y_train, cv=kfold, scoring=scoring)
names.append(name)
msg = "%s: %f (%f)" % (name, cv_results.mean(), cv_results.std())
print(msg)
# -
print(models)
# +
# Make prediction on the test dataset
for name, model in models:
model.fit(X_train, Y_train)
predictions = model.predict(X_test)
print(name)
print(accuracy_score(Y_test, predictions))
print(classification_report(Y_test, predictions))
# -
# # Test the model on a random example
# +
# last element is mitosis, higher the mitosis higher the likeyhood of cancer
example = np.array([[1, 1, 1, 1, 1, 1, 6, 7, 3]])
# orignal data
#examples = X
# reshape example to make it a column vector
example = example.reshape(len(example), -1)
for name, model in models:
prediction = model.predict(example)
if prediction == 2:
result = 'benign'
else:
result = 'malignant'
print(name)
print('The cancer is ' + result)
# -
| Breast Cancer Detection SVM and KNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: hots
# language: python
# name: hots
# ---
# + active=""
# %pip install git+https://github.com/VainF/pytorch-msssim/
# + active=""
# %pip uninstall -y pytorch_msssim
# +
# Adapted from https://github.com/Po-Hsun-Su/pytorch-ssim/blob/master/max_ssim.py
import torch
from torch.autograd import Variable
from torch import optim
from PIL import Image
import numpy as np
import sys, os
import torch.nn.functional as F
# sys.path.append(os.path.dirname(os.path.dirname(os.path.realpath(__file__))))
from pytorch_msssim import ssim, ms_ssim, SSIM, MS_SSIM
npImg1 = np.array(Image.open("../data/800px-Fox_Hunt_1893_Winslow_Homer.jpg"))
print(npImg1.shape)
# -
#img1 = torch.from_numpy(npImg1).float().unsqueeze(0).unsqueeze(0)/255.0
img1 = torch.from_numpy(npImg1).float().unsqueeze(0)/255.0
img2 = torch.rand(img1.size())
import matplotlib.pyplot as plt
# %matplotlib inline
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(13, 5))
ax1.imshow(img1.squeeze().numpy())
ax2.imshow(img2.squeeze().detach().numpy())
print(img1.min(), img1.max())
if torch.cuda.is_available():
img1 = img1.cuda()
img2 = img2.cuda()
# + active=""
# print(img1.shape)
# img1 = img1.permute(0, 1, 4, 2, 3)
# img2 = img2.permute(0, 1, 4, 2, 3)
# print(img1.shape)
# -
print(img1.shape)
img1 = img1.permute(0, 3, 1, 2)
img2 = img2.permute(0, 3, 1, 2)
print(img1.shape)
img1 = Variable( img1, requires_grad=False)
img2 = Variable( img2, requires_grad=True)
# +
ssim_value = ssim(img1, img2).item()
print("Initial ssim:", ssim_value)
ssim_loss = SSIM(win_size=11, win_sigma=1.5, data_range=1, size_average=True) #, channel=3)
# ms_ssim_loss = 1 - ms_ssim( X, Y, data_range=255, size_average=True )
optimizer = optim.SGD([img2], lr=0.05, momentum=.9, nesterov=True)
optimizer = optim.Adam([img2], lr=0.05)
while ssim_value < 0.9999:
optimizer.zero_grad()
#_ssim_loss = 1 - ms_ssim(img1, img2, data_range=1., size_average=True )
_ssim_loss = 1 - ssim_loss(img1, img2)
_ssim_loss.backward()
optimizer.step()
ssim_value = ssim(img1, img2).item()
print(ssim_value)
#img2_ = (img2 * 255.0).squeeze()
#np_img2 = img2_.detach().cpu().numpy().astype(np.uint8)
#Image.fromarray(np_img2).save('results.png')
# -
print(img2.min(), img2.max())
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(13, 5))
ax1.imshow(img1.squeeze().permute(1, 2, 0).numpy())
ax2.imshow(img2.squeeze().permute(1, 2, 0).detach().numpy());
| dev/2020-11-18 test sssim.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="R3H1Zj5zR94Q"
# # Sorting tables...
# + id="-roMIUk7R04f"
import pandas as pd
parameters_with_filtering = pd.read_csv('/content/FINAL_TABLE_IC.csv')
parameters_without_filtering = pd.read_csv('/content/ORIGINAL_PARAMETERS_IC.csv')
# + colab={"base_uri": "https://localhost:8080/"} id="6MkEajQ5Q2dV" executionInfo={"status": "ok", "timestamp": 1638119752760, "user_tz": 180, "elapsed": 4, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi5zFr3jUWJ1NdrUB6KSy3C4B0uR2RLqUiKRzi0S2M=s64", "userId": "17398030156076443435"}} outputId="65b9d95c-161a-4e29-8606-7114a2ab724f"
parameters_with_filtering.sort_values(by=['e_period', 'chi2']).head()
# + colab={"base_uri": "https://localhost:8080/"} id="5LrPBGcAQ-aH" executionInfo={"status": "ok", "timestamp": 1638119753214, "user_tz": 180, "elapsed": 457, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi5zFr3jUWJ1NdrUB6KSy3C4B0uR2RLqUiKRzi0S2M=s64", "userId": "17398030156076443435"}} outputId="f21ef1be-e87b-4025-a14b-95a3919e1f66"
# parameters_with_filtering.sort_values(by=['e_p', 'chi2']).head()
parameters_with_filtering[parameters_with_filtering['e_p'] != 0].sort_values(by=['e_p', 'chi2']).head()
# + colab={"base_uri": "https://localhost:8080/"} id="b8Smx00BQ-YD" executionInfo={"status": "ok", "timestamp": 1638119753215, "user_tz": 180, "elapsed": 13, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi5zFr3jUWJ1NdrUB6KSy3C4B0uR2RLqUiKRzi0S2M=s64", "userId": "17398030156076443435"}} outputId="e95a50b5-537a-4db3-ea26-a1eabe159075"
parameters_with_filtering.sort_values(by=['e_adivR', 'chi2']).head()
# + colab={"base_uri": "https://localhost:8080/"} id="5qnOuTmBQ-WB" executionInfo={"status": "ok", "timestamp": 1638119753215, "user_tz": 180, "elapsed": 12, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi5zFr3jUWJ1NdrUB6KSy3C4B0uR2RLqUiKRzi0S2M=s64", "userId": "17398030156076443435"}} outputId="b893fb55-f84a-46f5-ec46-c5083e65dbed"
teste = parameters_with_filtering[parameters_with_filtering['e_b'] != 0].sort_values(by=['e_b', 'chi2']); teste.head()
# + id="Tl0G2UIuycxL"
# + [markdown] id="8KS0jjeBshIX"
# # Procedures
#
# 1. Ensure that the parameters are compatible: within a range of +- 3 sigma (99.7% confidence)
# 2. Uncertainty (with filtering) <= Uncertainty (without filtering)
# 3. Compute percentage reduction:
#
# $$
# PR = \frac{\text{uncertantie with filtering} - \text{uncertantie without filtering}}{\text{uncertantie without filtering}} *100
# $$
#
# 4. Sort from largest to smallest for each parameter for each curve by PR
#
# 5. Take the average PR for each curve => Best filtering process
#
# + id="5toMAibYtXL2" cellView="form"
# @title Analyze results function
import pandas as np
import numpy as np
parameters_with_filtering = pd.read_csv('/content/FINAL_TABLE_IC.csv')
parameters_without_filtering = pd.read_csv('/content/ORIGINAL_PARAMETERS_IC.csv')
# Delete 102912369 (e_p original = 0)
parameters_with_filtering = parameters_with_filtering.drop(parameters_with_filtering[parameters_with_filtering['CoRoT_ID']==102912369].index)
parameters_without_filtering = parameters_without_filtering.drop(parameters_without_filtering[parameters_without_filtering['CoRoT_ID']==102912369].index)
def analyze_table(ID: int, parameter: str):
# Selecting lines corresponding to the ID
parameters_of_a_specific_id = parameters_with_filtering[parameters_with_filtering['CoRoT_ID'] == ID]
parameters_without_filtering_of_a_specific_id = parameters_without_filtering[parameters_without_filtering['CoRoT_ID'] == ID]
# How many different uncertainty values do we have?
# print(parameters_of_a_specific_id['e_'+parameter].value_counts())
# Keep values in the range of +- 3*sigma
parameter_referency = parameters_without_filtering_of_a_specific_id[parameter].iloc[0]
e_parameter_referency = parameters_without_filtering_of_a_specific_id['e_'+parameter].iloc[0]
valid_parameters_values_tmp = pd.DataFrame(columns=['CoRoT_ID', parameter, parameter+'_deleuil' , 'e_'+parameter, 'chi2', 'filter_technique', 'filter_order', 'filter_cutoff', 'filter_numNei'])
for index in range(len(parameters_of_a_specific_id)):
actual_parameter = parameters_of_a_specific_id[parameter].iloc[index]
actual_uncertanties = parameters_of_a_specific_id['e_'+parameter].iloc[index]
if (actual_parameter > (parameter_referency - 3*e_parameter_referency)) & (actual_parameter < (parameter_referency + 3*e_parameter_referency)):
valid_parameters_values_tmp = valid_parameters_values_tmp.append([{
'CoRoT_ID': ID,
parameter: actual_parameter,
parameter+'_deleuil': parameters_of_a_specific_id[parameter+'_deleuil'].iloc[index],
'e_'+parameter: parameters_of_a_specific_id['e_'+parameter].iloc[index],
'chi2': parameters_of_a_specific_id['chi2'].iloc[index],
'filter_technique': parameters_of_a_specific_id['filter_technique'].iloc[index],
'filter_order': parameters_of_a_specific_id['filter_order'].iloc[index],
'filter_cutoff': parameters_of_a_specific_id['filter_cutoff'].iloc[index],
'filter_numNei': parameters_of_a_specific_id['filter_numNei'].iloc[index]
}])
range_index = np.arange(0, len(valid_parameters_values_tmp))
valid_parameters_values_tmp = valid_parameters_values_tmp.set_index(range_index)
# Append parameters and uncertanties without filtering
valid_parameters_values_tmp['raw_'+parameter] = parameters_without_filtering_of_a_specific_id[parameter].iloc[0]
valid_parameters_values_tmp['e_raw_'+parameter] = parameters_without_filtering_of_a_specific_id['e_'+parameter].iloc[0]
valid_parameters_values_tmp = valid_parameters_values_tmp.reindex(['CoRoT_ID', parameter, parameter+'_deleuil', 'e_'+parameter, 'raw_'+parameter, 'e_raw_'+parameter, 'chi2', 'filter_technique', 'filter_order', 'filter_cutoff', 'filter_numNei'], axis=1)
# Keep parameters that uncertainties were smaller or equal than the original
count = 0
for index, row in valid_parameters_values_tmp.iterrows():
if row['e_'+parameter] > valid_parameters_values_tmp['e_raw_'+parameter].iloc[0]:
count += 1
valid_parameters_values_tmp.drop([index], inplace=True)
# print(f'Deleted {count} rows')
# Percetage reduction
pr_column = 'pr_'+parameter
valid_parameters_values_tmp[pr_column] = 100*(abs(valid_parameters_values_tmp['e_'+parameter] - valid_parameters_values_tmp['e_raw_'+parameter]))/valid_parameters_values_tmp['e_raw_'+parameter]
valid_parameters_values_tmp = valid_parameters_values_tmp.sort_values(by=pr_column, ascending=False)
valid_parameters_values_tmp = valid_parameters_values_tmp[['CoRoT_ID', parameter+'_deleuil', parameter, 'e_'+parameter, 'e_raw_'+parameter, 'chi2', 'filter_technique', 'filter_order', 'filter_cutoff', 'filter_numNei', pr_column]]
return valid_parameters_values_tmp
# + [markdown] id="D7bZhwQCfqkK"
# ### Parameter: period
# + id="3rrZ0ua1_uZU"
df_period = pd.DataFrame()
for ID in parameters_without_filtering['CoRoT_ID'].values:
analysis_results = analyze_table(ID, 'period').iloc[0]
df_period = df_period.append(analysis_results)
df_period = df_period[['CoRoT_ID', 'period_deleuil', 'period', 'e_period', 'e_raw_period', 'chi2', 'filter_technique', 'filter_order', 'filter_cutoff', 'filter_numNei', 'pr_period']]
df_period = df_period.sort_values(by='pr_period', ascending=False)
df_period.index = np.arange(0, len(df_period))
# df_period.to_csv('table_period.csv')
# + colab={"base_uri": "https://localhost:8080/"} id="173OHEmkVD_d" executionInfo={"status": "ok", "timestamp": 1638120187375, "user_tz": 180, "elapsed": 5, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi5zFr3jUWJ1NdrUB6KSy3C4B0uR2RLqUiKRzi0S2M=s64", "userId": "17398030156076443435"}} outputId="5c163d5a-2a6f-4d41-b0f3-148fd36a0f26"
df_period[df_period['CoRoT_ID']==105833549]
# + id="Jk-Yv112wGOs"
# df_period
# + [markdown] id="2KPflLiSfs0n"
# ### Parameter: p
# + id="iOPSfZ_AFbPm"
df_p = pd.DataFrame()
for ID in parameters_without_filtering['CoRoT_ID'].values:
analysis_results = analyze_table(ID, 'p').iloc[0]
df_p = df_p.append(analysis_results)
df_p = df_p[['CoRoT_ID', 'p_deleuil', 'p', 'e_p', 'e_raw_p', 'chi2', 'filter_technique', 'filter_order', 'filter_cutoff', 'filter_numNei', 'pr_p']]
df_p = df_p.sort_values(by='pr_p', ascending=False)
df_p.index = np.arange(0, len(df_p))
# df_p.to_csv('table_p.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 81} id="WgDEESuzdz_H" executionInfo={"status": "ok", "timestamp": 1638120180134, "user_tz": 180, "elapsed": 8, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi5zFr3jUWJ1NdrUB6KSy3C4B0uR2RLqUiKRzi0S2M=s64", "userId": "17398030156076443435"}} outputId="b1874de5-1b63-4731-d9e8-a1d22c69e6f0"
df_p[df_p['CoRoT_ID']==101086161]
# + id="SD07VaxFwSJ8"
# df_p
# + [markdown] id="85NQMnAufups"
# ### Parameter: adivR
# + id="n4vFvhJBFbNo"
df_adivR = pd.DataFrame()
for ID in parameters_without_filtering['CoRoT_ID'].values:
analysis_results = analyze_table(ID, 'adivR').iloc[0]
df_adivR = df_adivR.append(analysis_results)
df_adivR = df_adivR[['CoRoT_ID', 'adivR_deleuil', 'adivR', 'e_adivR', 'e_raw_adivR', 'chi2', 'filter_technique', 'filter_order', 'filter_cutoff', 'filter_numNei', 'pr_adivR']]
df_adivR = df_adivR.sort_values(by='pr_adivR', ascending=False)
df_adivR.index = np.arange(0, len(df_adivR))
# df_adivR.to_csv('table_adivR.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 81} id="owyiJ_xxYE3V" executionInfo={"status": "ok", "timestamp": 1638120202613, "user_tz": 180, "elapsed": 210, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi5zFr3jUWJ1NdrUB6KSy3C4B0uR2RLqUiKRzi0S2M=s64", "userId": "17398030156076443435"}} outputId="13f0c8a5-9137-4c72-a374-b705b7208f8d"
df_adivR[df_adivR['CoRoT_ID']==101086161]
# + id="nSf-xBLxwaMi"
# df_adivR
# + [markdown] id="JDY2rJlrfmmw"
# ### Parameter: b
# + id="gG3FMsDDFbLW"
############# WARNING #############
## ID: 102890318
## Parameter: b
## e_b = 0 !!!
df_b = pd.DataFrame()
for ID in parameters_without_filtering['CoRoT_ID'].values:
analysis_results = analyze_table(ID, 'b').iloc[0]
df_b = df_b.append(analysis_results)
df_b = df_b[['CoRoT_ID', 'b_deleuil', 'b', 'e_b', 'e_raw_b', 'chi2', 'filter_technique', 'filter_order', 'filter_cutoff', 'filter_numNei', 'pr_b']]
df_b = df_b.sort_values(by='pr_b', ascending=False)
df_b.index = np.arange(0, len(df_b))
# df_b.to_csv('table_b.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 81} id="k1rMniysY_fw" executionInfo={"status": "ok", "timestamp": 1638119827175, "user_tz": 180, "elapsed": 5, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi5zFr3jUWJ1NdrUB6KSy3C4B0uR2RLqUiKRzi0S2M=s64", "userId": "17398030156076443435"}} outputId="f0bff9ac-2761-422e-e2c0-86c3cea937e7"
df_b[df_b['CoRoT_ID']==102764809]
# + id="L-ptOSaaOWRL"
# df_b
# + [markdown] id="VeYd65creGZk"
# # Seeing filtering with details
# + id="ynKyh57neL7B"
# # !pip install /content/imt_lightcurve-2.0-py3-none-any.whl --force-reinstall
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="j6vVj2fyeeUc" executionInfo={"status": "ok", "timestamp": 1638120612717, "user_tz": 180, "elapsed": 1669, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi5zFr3jUWJ1NdrUB6KSy3C4B0uR2RLqUiKRzi0S2M=s64", "userId": "17398030156076443435"}} outputId="e49ad123-de6f-4226-c1b7-978a5bec83b6"
import numpy as np
import pandas as pd
from imt_lightcurve.models.lightcurve import LightCurve
from imt_lightcurve.visualization.data_viz import multi_line_plot
# Chosen a LightCurve to simulation process
CURVE_ID = '105833549'
# Importing lightcurve data from github
data = pd.read_csv('https://raw.githubusercontent.com/Guilherme-SSB/IC-CoRoT_Kepler/main/resampled_files/' + CURVE_ID + '.csv')
time = data.DATE.to_numpy()
flux = data.WHITEFLUX.to_numpy()
# Create the LightCurve object
curve = LightCurve(time=time, flux=flux)
# curve.plot()
# Folded curve
folded_curve = curve.fold(CURVE_ID)
# folded_curve.plot()
filtered = curve.butterworth_lowpass_filter(1, 0.6)
filtered.view_filtering_results()
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="AcupwbQPerIf" executionInfo={"status": "ok", "timestamp": 1638120613129, "user_tz": 180, "elapsed": 418, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gi5zFr3jUWJ1NdrUB6KSy3C4B0uR2RLqUiKRzi0S2M=s64", "userId": "17398030156076443435"}} outputId="5c0859d2-063c-422a-a5b4-a7a7497619ba"
curve_aux = LightCurve(filtered.time, filtered.filtered_flux)
filtered_folded_curve = curve_aux.fold(CURVE_ID)
multi_line_plot(folded_curve.time, folded_curve.flux, filtered_folded_curve.flux, label_y1='Folded original LC', label_y2='Folded filtered LC', title='Folded original LC x Folded filtered LC')
# + id="M9A2WGeohfHt"
| 0X - Analisando tabela final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/maskubica/house_prices/blob/master/HousePrices.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="6XzEiTaidw86" colab_type="code" colab={}
# !echo '{"username":"kamikaze95","key":"<KEY>"}' > /root/.kaggle/kaggle.json
# !kaggle competitions download -c house-prices-advanced-regression-techniques
# !ls
# + id="Attp3G037cD1" colab_type="code" colab={}
import pandas as pd
import matplotlib.pyplot as plt
train = pd.read_csv('train.csv')
test = pd.read_csv('test.csv')
columns_na = train.columns[train.isna().any(axis=0)]
#plt.hist(train[columns_na[0]])
for col in columns_na:
print(f"{col} \t {sum(train[col].isna())/ train[col].shape[0]*100} %")
train['LotFrontage']
# + id="Ts6A0qPS7dgd" colab_type="code" colab={}
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler, OneHotEncoder
from sklearn.compose import ColumnTransformer
numeric_transformer = Pipeline(steps=[
('imputer', SimpleImputer(strategy='median')),
('scaler', StandardScaler())])
categorical_transformer = Pipeline(steps=[
('imputer', SimpleImputer(strategy='constant', fill_value='missing')),
('onehot', OneHotEncoder(handle_unknown='ignore'))])
categorical_features = train.select_dtypes(include=['object']).dropna(axis=1).columns
categorical_features
# + id="cfUt4EZk7fj7" colab_type="code" colab={}
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.model_selection import train_test_split
X = train.dropna(axis=1).drop(['SalePrice', 'Id'], axis=1)
y = train['SalePrice']
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
categorical_transformer = Pipeline(steps=[
('onehot', OneHotEncoder(handle_unknown='ignore'))
])
preprocessor = ColumnTransformer(
transformers=[
('cat', categorical_transformer, categorical_features)]
)
xgb = Pipeline(steps=[('preprocessor', preprocessor),
('classifier', GradientBoostingClassifier())])
xgb.fit(X_train, y_train)
#clf.predict(X_test[:2])
#array([1, 0])
#clf.score(X_test, y_test)
| HousePrices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/iwatake2222/pico-loud_talking_detector/blob/master/01_script/training/train_micro_speech_model_talking_20210529_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="Y52U_k7Sho0w"
# !nvidia-smi
# + [markdown] id="pO4-CY_TCZZS"
# # Train a Simple Audio Recognition Model
# + [markdown] id="BaFfr7DHRmGF"
# This notebook demonstrates how to train a 20 kB [Simple Audio Recognition](https://www.tensorflow.org/tutorials/sequences/audio_recognition) model to recognize keywords in speech.
#
# The model created in this notebook is used in the [micro_speech](https://github.com/tensorflow/tensorflow/tree/master/tensorflow/lite/micro/examples/micro_speech) example for [TensorFlow Lite for MicroControllers](https://www.tensorflow.org/lite/microcontrollers/overview).
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/tensorflow/blob/master/tensorflow/lite/micro/examples/micro_speech/train/train_micro_speech_model.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/tensorflow/blob/master/tensorflow/lite/micro/examples/micro_speech/train/train_micro_speech_model.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
#
# + [markdown] id="8Zai5GNyXji1"
# ## Prepare my dataset (iwatake2222)
# + [markdown] id="xuPZ7HzIE7RO"
# ### Download dataset (iwatake2222)
# + id="Oj1b7Mo08RL5"
import os
import glob
import subprocess
CLIP_DURATION = 10 * 1000
DATASET_DIR = "dataset/"
# !rm -rf {DATASET_DIR} && mkdir -p {DATASET_DIR}
def download(file_id, file_name):
subprocess.run(["curl", "-sc", "/tmp/cookie", f"https://drive.google.com/uc?export=download&id={file_id}"])
cmd = ["awk", "/_warning_/ {print $NF}", "/tmp/cookie"]
code = subprocess.run(cmd, shell=False, stdout=subprocess.PIPE, check=True).stdout.decode("utf-8").replace("\n", "")
subprocess.run(["curl", "-Lb", "/tmp/cookie", f"https://drive.google.com/uc?export=download&confirm={code}&id={file_id}", "-o", f"{file_name}"])
# def extract_dataset(file_id, file_name, dataset_dir):
# download(file_id, file_name)
# subprocess.run(["unzip", "-o", f"{file_name}"])
# data_path = os.path.splitext(os.path.basename(file_name))[0]
# subprocess.run(f"cp -rf {data_path}/* {dataset_dir}/.", shell=True)
def extract_dataset(file_id, file_name, dataset_dir):
if not os.path.exists(file_name):
download(file_id, file_name)
subprocess.run(["tar", "xzvf", file_name, "--strip", "1", "-C", dataset_dir])
''' Download my dataset '''
# AudioSet (use not_talking data only)
extract_dataset("1wLit745TX4rw_KgUJULuhZxXETC58fgd", "balanced_train_segments.tgz", DATASET_DIR)
# extract_dataset("1l5pj8DO0rreT-OimYdZAf2mJdmV_6nYu", "eval_segments.tgz", DATASET_DIR)
# !rm -rf {DATASET_DIR}/ambiguous
# !rm -rf {DATASET_DIR}/talking/*
# My data
extract_dataset("13HC_vZeXwqpz4eKZliLKomvTBuYW1-jt", "music_pops.tgz", DATASET_DIR + "not_talking")
extract_dataset("1tlkCJ2RnhapPmPeIhd09dzTD9xrovyV4", "music.tgz", DATASET_DIR + "not_talking")
extract_dataset("12_z34bHB1bR3QbTM-rzJOq8Fol1G3kpj", "yoshimoto.tgz", DATASET_DIR + "talking")
# !rm -rf temp && mkdir temp
extract_dataset("1MhhJJxqCdd33gxgwBkuTJzABURjOsUNP", "my_jp_talk.tgz", "temp")
# !find temp -name *.wav -exec mv {} temp \;
# !mv temp/*.wav {DATASET_DIR}"/talking/."
# !rm -rf temp
# WANTED_WORDS_LIST = ["talking", "not_talking", "ambiguous"]
WANTED_WORDS_LIST = ["talking", "not_talking"]
# ''' Download background_noise '''
# # !curl -O "https://storage.googleapis.com/download.tensorflow.org/data/speech_commands_v0.02.tar.gz"
# # !tar xzf speech_commands_v0.02.tar.gz ./_background_noise_
# # !cp -r _background_noise_ {DATASET_DIR}/.
# + id="6M7I36qb0i4m"
import os
import glob
import random
''' Adjust the number of dataset '''
def get_file_num(directory):
return len([name for name in os.listdir(directory) if os.path.isfile(directory + "/" + name)])
def choose_random_data(DATASET_DIR, label, data_num):
org_path = DATASET_DIR + "/" + label + "/"
tmp_path = DATASET_DIR + "/temp_" + label + "/"
os.makedirs(tmp_path, exist_ok=True)
subprocess.run(f"mv {org_path}/* {tmp_path}/.", shell=True)
files = [r.split('/')[-1] for r in glob.glob(tmp_path + "/*.wav")]
for i in range(data_num):
chosen_file_name = random.choice(files)
files.remove(chosen_file_name)
chosen_file_path = tmp_path + chosen_file_name
subprocess.run(f"mv {chosen_file_path} {org_path}/.", shell=True)
subprocess.run(f"rm -rf {tmp_path}", shell=True)
# !rm -rf {tmp_path}
# data_num = get_file_num(DATASET_DIR + "talking")
# print(str(get_file_num(DATASET_DIR + "talking")), str(get_file_num(DATASET_DIR + "not_talking")), str(get_file_num(DATASET_DIR + "ambiguous")))
# choose_random_data(DATASET_DIR, "not_talking", data_num)
# choose_random_data(DATASET_DIR, "ambiguous", data_num)
# print(str(get_file_num(DATASET_DIR + "talking")), str(get_file_num(DATASET_DIR + "not_talking")), str(get_file_num(DATASET_DIR + "ambiguous")))
print(str(get_file_num(DATASET_DIR + "talking")), str(get_file_num(DATASET_DIR + "not_talking")))
# + id="pLW7XXU93Wgp"
import os
import glob
import shutil
import time
import random
import librosa
import soundfile
def separate_wav(target_dir, output_dir, sampling_rate=16000, output_duration_time=5,delete_original=False):
output_duration_sample = int(sampling_rate * output_duration_time)
''' Process for selected input files '''
wav_path_list = glob.glob(target_dir + "/*.wav")
for wav_path in wav_path_list:
basename, ext = os.path.splitext(os.path.basename(wav_path))
try:
data, sr = librosa.core.load(wav_path, sr=sampling_rate, mono=True)
except:
continue
duration_sample = len(data)
index = 0
while (index + 0.5) * output_duration_sample <= duration_sample:
if (index + 1) * output_duration_sample <= duration_sample:
data_out = data[index * output_duration_sample : (index + 1) * output_duration_sample]
else:
data_out = data[duration_sample - output_duration_sample : duration_sample]
output_path = output_dir + "/" + basename + "_" + f"{index:02}" + ".wav"
soundfile.write(output_path, data_out, samplerate=sampling_rate, subtype="PCM_16")
index += 1
if delete_original:
os.remove(wav_path)
rest_data_sample = duration_sample - (index * output_duration_sample)
# if rest_data_sample > 0:
# print(f"Warning: data dropped, {basename}, {str(rest_data_sample)}")
separate_wav(DATASET_DIR + "/talking/.", DATASET_DIR + "/talking/.", sampling_rate=16000, output_duration_time=CLIP_DURATION/1000, delete_original=True)
separate_wav(DATASET_DIR + "/not_talking/.", DATASET_DIR + "/not_talking/.", sampling_rate=16000, output_duration_time=CLIP_DURATION/1000, delete_original=True)
print(str(get_file_num(DATASET_DIR + "talking")), str(get_file_num(DATASET_DIR + "not_talking")))
# + [markdown] id="Qv7KX6BvFEiQ"
# ### Add noise and background (iwatake2222)
# + id="9QQG8aa7FC0e"
# !apt install librosa soundfile
# + id="lAvSmGPeFTU_"
import os
import glob
import shutil
import time
import random
import librosa
import soundfile
def clear_last_sep(dir):
dir.replace(os.sep,'/')
if dir[-1] == "/":
dir = dir[:-1]
return dir
def add_noise(target_dir, noise_dir, output_dir, signature_text="", sampling_rate=16000, process_ratio=0.3, original_volume=1.0, noise_volume=1.0):
if signature_text == "":
signature_text = os.path.splitext(os.path.basename(clear_last_sep(noise_dir)))[0]
''' Make sure the shuffling is deterministic for reproduce '''
random.seed(1234)
''' Read noise data as array '''
noise_list = []
noise_wav_path_list = glob.glob(noise_dir + "/*.wav")
for noise_wav_path in noise_wav_path_list:
data, sr = librosa.core.load(noise_wav_path, sr=sampling_rate, mono=True)
noise_list.append(data)
''' Process for selected input files '''
wav_path_list = glob.glob(target_dir + "/*.wav")
random.shuffle(wav_path_list)
wav_path_list = wav_path_list[:int(len(wav_path_list) * process_ratio)]
for wav_path in wav_path_list:
basename, ext = os.path.splitext(os.path.basename(wav_path))
output_path = output_dir + "/" + basename + "_" + signature_text + ".wav"
data, sr = librosa.core.load(wav_path, sr=sampling_rate, mono=True)
duration_sample = len(data)
noise = random.choice(noise_list)
start_sample = int(random.uniform(0, len(noise) - duration_sample - 1))
data = data * original_volume + noise[start_sample:start_sample + duration_sample] * noise_volume
soundfile.write(output_path, data, samplerate=sampling_rate, subtype="PCM_16")
def create_noise(noise_dir, output_dir, signature_text="", sampling_rate=16000, duration_time=10, output_number_of_file=10, noise_volume=1.0):
if signature_text == "":
signature_text = os.path.splitext(os.path.basename(clear_last_sep(noise_dir)))[0]
''' Make sure the shuffling is deterministic for reproduce '''
random.seed(1234)
''' Read noise data as array '''
noise_list = []
noise_wav_path_list = glob.glob(noise_dir + "/*.wav")
for noise_wav_path in noise_wav_path_list:
data, sr = librosa.core.load(noise_wav_path, sr=sampling_rate, mono=True)
noise_list.append(data)
duration_sample = int(duration_time * sampling_rate)
''' Process to create files'''
for i in range(output_number_of_file):
noise = random.choice(noise_list)
start_sample = int(random.uniform(0, len(noise) - duration_sample - 1))
data = noise[start_sample:start_sample + duration_sample] * noise_volume
output_path = output_dir + "/" + signature_text + f"_{i:05}.wav"
soundfile.write(output_path, data, samplerate=sampling_rate, subtype="PCM_16")
# + id="ks77gb92Fcvo"
''' Download noise data '''
BACKGROUND_DIR = "background/"
NOISE_DIR = "noise/"
# !rm -rf {BACKGROUND_DIR} && mkdir -p {BACKGROUND_DIR}
# !rm -rf {NOISE_DIR} && mkdir -p {NOISE_DIR}
extract_dataset("19vFtkVp1d_e8lBBnfnD8K8JohBPcPM_H", "background.tgz", BACKGROUND_DIR)
extract_dataset("12R0jtfXr6cGWYYZaTcV2B0nJvXV8AV-F", "mic_noise.tgz", NOISE_DIR)
''' Add noise files into "not_talking" '''
# Create noise data after separatint test data
# noise_num = int(get_file_num(DATASET_DIR + "not_talking") * 0.2)
# create_noise(BACKGROUND_DIR, DATASET_DIR + "not_talking", duration_time=CLIP_DURATION/1000, output_number_of_file=noise_num, noise_volume=1.0)
# create_noise(NOISE_DIR, DATASET_DIR + "not_talking", duration_time=CLIP_DURATION/1000, output_number_of_file=noise_num, noise_volume=1.0)
''' Mix noise and "talking" '''
# the order is important (mix background, then mix noise)
add_noise(DATASET_DIR + "talking", BACKGROUND_DIR, DATASET_DIR + "talking", process_ratio=0.5, original_volume=1.0, noise_volume=1.0)
add_noise(DATASET_DIR + "talking", NOISE_DIR, DATASET_DIR + "talking", process_ratio=0.5, original_volume=1.0, noise_volume=1.0)
print(str(get_file_num(DATASET_DIR + "talking")), str(get_file_num(DATASET_DIR + "not_talking")))
# + [markdown] id="YWd_qA8kMwe9"
# ### Separate test data
# Manually separate test data from training data. because the training script randomly pick-up training/validation/test data and it causes data leakage.
# Copy wav files which has similar prefix
# + id="puvGiSW9JAg9"
import shutil
DATASET_TEST_DIR = "dataset_test/"
# !rm -rf {DATASET_TEST_DIR} && mkdir -p {DATASET_TEST_DIR} && mkdir -p {DATASET_TEST_DIR}/talking && mkdir -p {DATASET_TEST_DIR}/not_talking
def move_test_data(src_path, dst_path, ratio):
random.seed(984983)
num_to_be_mv = int(len(glob.glob(src_path + "/*.wav")) * ratio)
while num_to_be_mv > 0:
org_file_list = glob.glob(src_path + "/*.wav")
basename, ext = os.path.splitext(os.path.basename(random.choice(org_file_list)))
# Check the first letters to identigy if the files are the same (see longer is filename is date)
num_to_identify_scene = 5
if basename[:8].isdigit():
num_to_identify_scene = 13
target_file_list = glob.glob(src_path + "/" + basename[:num_to_identify_scene] + "*.wav")
for target_file in target_file_list:
shutil.move(target_file, dst_path + "/.")
num_to_be_mv -= 1
move_test_data(DATASET_DIR + "talking", DATASET_TEST_DIR + "talking", 0.1)
move_test_data(DATASET_DIR + "not_talking", DATASET_TEST_DIR + "not_talking", 0.1)
# move_test_data(DATASET_TEST_DIR + "talking", DATASET_DIR + "talking", 1.0) # revert
# move_test_data(DATASET_TEST_DIR + "not_talking", DATASET_DIR + "not_talking", 1.0) # revert
print(str(get_file_num(DATASET_DIR + "talking")), str(get_file_num(DATASET_DIR + "not_talking")))
print(str(get_file_num(DATASET_TEST_DIR + "talking")), str(get_file_num(DATASET_TEST_DIR + "not_talking")))
# + id="2AjjDkt1g7oG"
''' Add noise data to training data (this sould be after separating test data) '''
noise_num = int(get_file_num(DATASET_DIR + "not_talking") * 0.2)
create_noise(BACKGROUND_DIR, DATASET_DIR + "not_talking", duration_time=CLIP_DURATION/1000, output_number_of_file=noise_num, noise_volume=1.0)
create_noise(NOISE_DIR, DATASET_DIR + "not_talking", duration_time=CLIP_DURATION/1000, output_number_of_file=noise_num, noise_volume=1.0)
print(str(get_file_num(DATASET_DIR + "talking")), str(get_file_num(DATASET_DIR + "not_talking")))
print(str(get_file_num(DATASET_TEST_DIR + "talking")), str(get_file_num(DATASET_TEST_DIR + "not_talking")))
# + [markdown] id="XaVtYN4nlCft"
# **Training is much faster using GPU acceleration.** Before you proceed, ensure you are using a GPU runtime by going to **Runtime -> Change runtime type** and set **Hardware accelerator: GPU**. Training 15,000 iterations will take 1.5 - 2 hours on a GPU runtime.
#
# ## Configure Defaults
#
# **MODIFY** the following constants for your specific use case.
# + id="ludfxbNIaegy"
# A comma-delimited list of the words you want to train for.
# The options are: yes,no,up,down,left,right,on,off,stop,go
# All the other words will be used to train an "unknown" label and silent
# audio data with no spoken words will be used to train a "silence" label.
''' Modified by iwatake2222 ---------------------------------------------- '''
# WANTED_WORDS = "yes,no"
WANTED_WORDS = ",".join(WANTED_WORDS_LIST)
# The number of steps and learning rates can be specified as comma-separated
# lists to define the rate at each stage. For example,
# TRAINING_STEPS=12000,3000 and LEARNING_RATE=0.001,0.0001
# will run 12,000 training loops in total, with a rate of 0.001 for the first
# 8,000, and 0.0001 for the final 3,000.
''' Modified by iwatake2222 ---------------------------------------------- '''
TRAINING_STEPS = "12000,3000"
# TRAINING_STEPS = "5000,1000"
LEARNING_RATE = "0.001,0.0001"
# Calculate the total number of steps, which is used to identify the checkpoint
# file name.
TOTAL_STEPS = str(sum(map(lambda string: int(string), TRAINING_STEPS.split(","))))
# Print the configuration to confirm it
print("Training these words: %s" % WANTED_WORDS)
print("Training steps in each stage: %s" % TRAINING_STEPS)
print("Learning rate in each stage: %s" % LEARNING_RATE)
print("Total number of training steps: %s" % TOTAL_STEPS)
# + [markdown] id="gCgeOpvY9pAi"
# **DO NOT MODIFY** the following constants as they include filepaths used in this notebook and data that is shared during training and inference.
# + id="Nd1iM1o2ymvA"
# Calculate the percentage of 'silence' and 'unknown' training samples required
# to ensure that we have equal number of samples for each label.
number_of_labels = WANTED_WORDS.count(',') + 1
number_of_total_labels = number_of_labels + 2 # for 'silence' and 'unknown' label
equal_percentage_of_training_samples = int(100.0/(number_of_total_labels))
''' Modified by iwatake2222 ---------------------------------------------- '''
equal_percentage_of_training_samples = 0
SILENT_PERCENTAGE = equal_percentage_of_training_samples
UNKNOWN_PERCENTAGE = equal_percentage_of_training_samples
# Constants which are shared during training and inference
PREPROCESS = 'micro'
''' Modified by iwatake2222 ---------------------------------------------- '''
# CLIP_DURATION = 10000
WINDOW_SIZE = 30
WINDOW_STRIDE = 20
FEATURE_BIN_COUNT = 40
# MODEL_ARCHITECTURE = 'conv' # Other options include: single_fc, conv,
# low_latency_conv, low_latency_svdf, tiny_embedding_conv
MODEL_ARCHITECTURE = 'tiny_conv' # Other options include: single_fc, conv,
# Constants used during training only
VERBOSITY = 'DEBUG'
''' Modified by iwatake2222 ---------------------------------------------- '''
EVAL_STEP_INTERVAL = '1000'
SAVE_STEP_INTERVAL = '1000'
# EVAL_STEP_INTERVAL = '100'
# SAVE_STEP_INTERVAL = '100'
# Constants for training directories and filepaths
# DATASET_DIR = 'dataset/'
LOGS_DIR = 'logs/'
TRAIN_DIR = 'train/' # for training checkpoints and other files.
# Constants for inference directories and filepaths
import os
MODELS_DIR = 'models'
if not os.path.exists(MODELS_DIR):
os.mkdir(MODELS_DIR)
MODEL_TF = os.path.join(MODELS_DIR, 'model.pb')
MODEL_TFLITE = os.path.join(MODELS_DIR, 'model.tflite')
FLOAT_MODEL_TFLITE = os.path.join(MODELS_DIR, 'float_model.tflite')
MODEL_TFLITE_MICRO = os.path.join(MODELS_DIR, 'model.cc')
SAVED_MODEL = os.path.join(MODELS_DIR, 'saved_model')
QUANT_INPUT_MIN = 0.0
QUANT_INPUT_MAX = 26.0
QUANT_INPUT_RANGE = QUANT_INPUT_MAX - QUANT_INPUT_MIN
# + [markdown] id="6rLYpvtg9P4o"
# ## Setup Environment
#
# Install Dependencies
# + id="ed_XpUrU5DvY"
# %tensorflow_version 1.x
import tensorflow as tf
# + [markdown] id="T9Ty5mR58E4i"
# **DELETE** any old data from previous runs
#
# + id="APGx0fEh7hFF"
''' Modified by iwatake2222 ---------------------------------------------- '''
# # !rm -rf {DATASET_DIR} {LOGS_DIR} {TRAIN_DIR} {MODELS_DIR}
# !rm -rf {LOGS_DIR} {TRAIN_DIR} {MODELS_DIR}
# + [markdown] id="GfEUlfFBizio"
# Clone the TensorFlow Github Repository, which contains the relevant code required to run this tutorial.
# + id="yZArmzT85SLq"
# !git clone -q --depth 1 https://github.com/tensorflow/tensorflow
# + [markdown] id="nS9swHLSi7Bi"
# Load TensorBoard to visualize the accuracy and loss as training proceeds.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 852} id="q4qF1VxP3UE4" outputId="1f58aa19-c75a-41d3-c8b6-0c1fd7b1f3b1"
# %load_ext tensorboard
# %tensorboard --logdir {LOGS_DIR}
# + [markdown] id="x1J96Ron-O4R"
# ## Training
#
# The following script downloads the dataset and begin training.
# + colab={"background_save": true} id="VJsEZx6lynbY"
''' Modified by iwatake2222 ----------------------------------------------- '''
# # !python tensorflow/tensorflow/examples/speech_commands/train.py \
# # --data_dir={DATASET_DIR} \
# # --wanted_words={WANTED_WORDS} \
# # --silence_percentage={SILENT_PERCENTAGE} \
# # --unknown_percentage={UNKNOWN_PERCENTAGE} \
# # --preprocess={PREPROCESS} \
# # --window_stride={WINDOW_STRIDE} \
# # --model_architecture={MODEL_ARCHITECTURE} \
# # --how_many_training_steps={TRAINING_STEPS} \
# # --learning_rate={LEARNING_RATE} \
# # --train_dir={TRAIN_DIR} \
# # --summaries_dir={LOGS_DIR} \
# # --verbosity={VERBOSITY} \
# # --eval_step_interval={EVAL_STEP_INTERVAL} \
# # --save_step_interval={SAVE_STEP_INTERVAL}
# !python tensorflow/tensorflow/examples/speech_commands/train.py \
# --data_url="" \
# --data_dir={DATASET_DIR} \
# --wanted_words={WANTED_WORDS} \
# --silence_percentage={SILENT_PERCENTAGE} \
# --unknown_percentage={UNKNOWN_PERCENTAGE} \
# --preprocess={PREPROCESS} \
# --clip_duration_ms={CLIP_DURATION} \
# --window_size_ms={WINDOW_SIZE} \
# --window_stride={WINDOW_STRIDE} \
# --feature_bin_count={FEATURE_BIN_COUNT} \
# --model_architecture={MODEL_ARCHITECTURE} \
# --how_many_training_steps={TRAINING_STEPS} \
# --learning_rate={LEARNING_RATE} \
# --train_dir={TRAIN_DIR} \
# --summaries_dir={LOGS_DIR} \
# --verbosity={VERBOSITY} \
# --validation_percentage=10 \
# --testing_percentage=0 \
# --eval_step_interval={EVAL_STEP_INTERVAL} \
# --save_step_interval={SAVE_STEP_INTERVAL}
# \
# --start_checkpoint=./train/conv.ckpt-3000
# + [markdown] id="UczQKtqLi7OJ"
# # Skipping the training
#
# If you don't want to spend an hour or two training the model from scratch, you can download pretrained checkpoints by uncommenting the lines below (removing the '#'s at the start of each line) and running them.
# + id="RZw3VNlnla-J"
# #!curl -O "https://storage.googleapis.com/download.tensorflow.org/models/tflite/speech_micro_train_2020_05_10.tgz"
# #!tar xzf speech_micro_train_2020_05_10.tgz
# + [markdown] id="XQUJLrdS-ftl"
# ## Generate a TensorFlow Model for Inference
#
# Combine relevant training results (graph, weights, etc) into a single file for inference. This process is known as freezing a model and the resulting model is known as a frozen model/graph, as it cannot be further re-trained after this process.
# + id="xyc3_eLh9sAg"
''' Modified by iwatake2222 ---------------------------------------------- '''
# # !rm -rf {SAVED_MODEL}
# # !python tensorflow/tensorflow/examples/speech_commands/freeze.py \
# # --wanted_words=$WANTED_WORDS \
# # --window_stride_ms=$WINDOW_STRIDE \
# # --preprocess=$PREPROCESS \
# # --model_architecture=$MODEL_ARCHITECTURE \
# # --start_checkpoint=$TRAIN_DIR$MODEL_ARCHITECTURE'.ckpt-'{TOTAL_STEPS} \
# # --save_format=saved_model \
# # --output_file={SAVED_MODEL}
# !rm -rf {SAVED_MODEL}
# !python tensorflow/tensorflow/examples/speech_commands/freeze.py \
# --wanted_words=$WANTED_WORDS \
# --clip_duration_ms=$CLIP_DURATION \
# --clip_stride_ms=$WINDOW_SIZE \
# --window_size_ms=$WINDOW_SIZE \
# --window_stride_ms=$WINDOW_STRIDE \
# --feature_bin_count=$FEATURE_BIN_COUNT \
# --preprocess=$PREPROCESS \
# --model_architecture=$MODEL_ARCHITECTURE \
# --start_checkpoint=$TRAIN_DIR$MODEL_ARCHITECTURE'.ckpt-'{TOTAL_STEPS} \
# --save_format=saved_model \
# --output_file={SAVED_MODEL}
# + [markdown] id="_DBGDxVI-nKG"
# ## Generate a TensorFlow Lite Model
#
# Convert the frozen graph into a TensorFlow Lite model, which is fully quantized for use with embedded devices.
#
# The following cell will also print the model size, which will be under 20 kilobytes.
# + id="RIitkqvGWmre"
import sys
# We add this path so we can import the speech processing modules.
sys.path.append("/content/tensorflow/tensorflow/examples/speech_commands/")
import input_data
import models
import numpy as np
# + id="kzqECqMxgBh4"
''' Modified by iwatake2222 ---------------------------------------------- '''
SAMPLE_RATE = 16000
# CLIP_DURATION_MS = 1000
# WINDOW_SIZE_MS = 30.0
# FEATURE_BIN_COUNT = 40
# BACKGROUND_FREQUENCY = 0.8
BACKGROUND_FREQUENCY = 0.0
BACKGROUND_VOLUME_RANGE = 0.1
TIME_SHIFT_MS = 100.0
''' Modified by iwatake2222 ---------------------------------------------- '''
# DATA_URL = 'https://storage.googleapis.com/download.tensorflow.org/data/speech_commands_v0.02.tar.gz'
DATA_URL = ''
# VALIDATION_PERCENTAGE = 10
# TESTING_PERCENTAGE = 10
VALIDATION_PERCENTAGE = 0
# use 100% causes error (AudioProcessor needs at least 1 trainign data)
TESTING_PERCENTAGE = 99
# + id="rNQdAplJV1fz"
''' Modified by iwatake2222 ---------------------------------------------- '''
model_settings = models.prepare_model_settings(
len(input_data.prepare_words_list(WANTED_WORDS.split(','))),
SAMPLE_RATE, CLIP_DURATION, WINDOW_SIZE,
WINDOW_STRIDE, FEATURE_BIN_COUNT, PREPROCESS)
# audio_processor = input_data.AudioProcessor(
# DATA_URL, DATASET_DIR,
# SILENT_PERCENTAGE, UNKNOWN_PERCENTAGE,
# WANTED_WORDS.split(','), VALIDATION_PERCENTAGE,
# TESTING_PERCENTAGE, model_settings, LOGS_DIR)
audio_processor = input_data.AudioProcessor(
DATA_URL, DATASET_TEST_DIR,
SILENT_PERCENTAGE, UNKNOWN_PERCENTAGE,
WANTED_WORDS.split(','), VALIDATION_PERCENTAGE,
TESTING_PERCENTAGE, model_settings, LOGS_DIR)
# + id="lBj_AyCh1cC0"
with tf.Session() as sess:
float_converter = tf.lite.TFLiteConverter.from_saved_model(SAVED_MODEL)
float_tflite_model = float_converter.convert()
float_tflite_model_size = open(FLOAT_MODEL_TFLITE, "wb").write(float_tflite_model)
print("Float model is %d bytes" % float_tflite_model_size)
converter = tf.lite.TFLiteConverter.from_saved_model(SAVED_MODEL)
converter.optimizations = [tf.lite.Optimize.DEFAULT]
converter.inference_input_type = tf.lite.constants.INT8
converter.inference_output_type = tf.lite.constants.INT8
def representative_dataset_gen():
for i in range(1000):
data, _ = audio_processor.get_data(1, i*1, model_settings,
BACKGROUND_FREQUENCY,
BACKGROUND_VOLUME_RANGE,
TIME_SHIFT_MS,
'testing',
sess)
''' Modified by iwatake2222 ---------------------------------------------- '''
# flattened_data = np.array(data.flatten(), dtype=np.float32).reshape(1, 1960)
flattened_data = np.array(data.flatten(), dtype=np.float32).reshape(1, int(40 * (CLIP_DURATION / 20 - 1)))
yield [flattened_data]
converter.representative_dataset = representative_dataset_gen
tflite_model = converter.convert()
tflite_model_size = open(MODEL_TFLITE, "wb").write(tflite_model)
print("Quantized model is %d bytes" % tflite_model_size)
# + [markdown] id="EeLiDZTbLkzv"
# ## Testing the TensorFlow Lite model's accuracy
#
# Verify that the model we've exported is still accurate, using the TF Lite Python API and our test set.
# + id="wQsEteKRLryJ"
# Helper function to run inference
def run_tflite_inference(tflite_model_path, model_type="Float"):
# Load test data
np.random.seed(0) # set random seed for reproducible test results.
with tf.Session() as sess:
test_data, test_labels = audio_processor.get_data(
-1, 0, model_settings, BACKGROUND_FREQUENCY, BACKGROUND_VOLUME_RANGE,
TIME_SHIFT_MS, 'testing', sess)
test_data = np.expand_dims(test_data, axis=1).astype(np.float32)
# Initialize the interpreter
interpreter = tf.lite.Interpreter(tflite_model_path)
interpreter.allocate_tensors()
input_details = interpreter.get_input_details()[0]
output_details = interpreter.get_output_details()[0]
# For quantized models, manually quantize the input data from float to integer
if model_type == "Quantized":
input_scale, input_zero_point = input_details["quantization"]
test_data = test_data / input_scale + input_zero_point
test_data = test_data.astype(input_details["dtype"])
correct_predictions = 0
for i in range(len(test_data)):
interpreter.set_tensor(input_details["index"], test_data[i])
''' Modified by iwatake2222 ---------------------------------------------- '''
# To avoid "interpreter.invoke() There is at least 1 reference to internal data" error
try:
interpreter.invoke()
except:
interpreter = tf.lite.Interpreter(tflite_model_path)
interpreter.allocate_tensors()
interpreter.invoke()
output = interpreter.get_tensor(output_details["index"])[0]
top_prediction = output.argmax()
correct_predictions += (top_prediction == test_labels[i])
print('%s model accuracy is %f%% (Number of test samples=%d)' % (
model_type, (correct_predictions * 100) / len(test_data), len(test_data)))
# + colab={"base_uri": "https://localhost:8080/"} id="l-pD52Na6jRa" outputId="f45c70cd-54a5-441a-eca5-22d4076d4077"
# Compute float model accuracy
run_tflite_inference(FLOAT_MODEL_TFLITE)
# Compute quantized model accuracy
run_tflite_inference(MODEL_TFLITE, model_type='Quantized')
# + [markdown] id="dt6Zqbxu-wIi"
# ## Generate a TensorFlow Lite for MicroControllers Model
# Convert the TensorFlow Lite model into a C source file that can be loaded by TensorFlow Lite for Microcontrollers.
# + id="XohZOTjR8ZyE"
# Install xxd if it is not available
# !apt-get update && apt-get -qq install xxd
# Convert to a C source file
# !xxd -i {MODEL_TFLITE} > {MODEL_TFLITE_MICRO}
# Update variable names
REPLACE_TEXT = MODEL_TFLITE.replace('/', '_').replace('.', '_')
# !sed -i 's/'{REPLACE_TEXT}'/g_model/g' {MODEL_TFLITE_MICRO}
# + [markdown] id="2pQnN0i_-0L2"
# ## Deploy to a Microcontroller
#
# Follow the instructions in the [micro_speech](https://github.com/tensorflow/tensorflow/tree/master/tensorflow/lite/micro/examples/micro_speech) README.md for [TensorFlow Lite for MicroControllers](https://www.tensorflow.org/lite/microcontrollers/overview) to deploy this model on a specific microcontroller.
#
# **Reference Model:** If you have not modified this notebook, you can follow the instructions as is, to deploy the model. Refer to the [`micro_speech/train/models`](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/lite/micro/examples/micro_speech/train/models) directory to access the models generated in this notebook.
#
# **New Model:** If you have generated a new model to identify different words: (i) Update `kCategoryCount` and `kCategoryLabels` in [`micro_speech/micro_features/micro_model_settings.h`](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/lite/micro/examples/micro_speech/micro_features/micro_model_settings.h) and (ii) Update the values assigned to the variables defined in [`micro_speech/micro_features/model.cc`](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/lite/micro/examples/micro_speech/micro_features/model.cc) with values displayed after running the following cell.
# + id="eoYyh0VU8pca"
# Print the C source file
# # !cat {MODEL_TFLITE_MICRO}
''' Modified by iwatake2222 ---------------------------------------------- '''
# !tar czvf models.tgz models
| 01_script/training/old/train_micro_speech_model_talking_20210529_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# This notebook uses a simple polynomial regression problem to test out your jupyter notebook setup
import numpy as np
import scipy as sp
from sklearn.preprocessing import PolynomialFeatures
from sklearn import linear_model
import matplotlib.pyplot as plt
# +
N = 500
x1= np.random.normal(loc=17,scale=5,size=N)[:,np.newaxis]
x2= np.random.normal(loc=0,scale=5,size=N)[:,np.newaxis]
y = 3*x1 + np.random.normal(loc=.0, scale=.4, size=N)[:,np.newaxis]
x1 = np.sort(x1)
x1 = np.random.rand(N) * 10 - 5
x1 = np.sort(x1)
x1 = x1[:,np.newaxis]
noise = 0.1
def f(x):
x = x.ravel()
return np.exp(-x ** 2) + 1. * np.exp(-(x - 1) ** 2)
y = f(x1) + np.random.normal(0.0, noise, N)
y = y[:,np.newaxis]
def polynomial_regr(degree=1):
X_tr = x1[:].astype(float)
y_tr = y[:].astype(float)
poly = PolynomialFeatures(degree=degree)
X_tr_ = poly.fit_transform(X_tr)
regr = linear_model.LinearRegression()
regr.fit(X_tr_, y_tr)
y_pred_tr = regr.predict(X_tr_)[:]
plt.plot(X_tr,y_tr,'.b',markersize=6,alpha=.4 );
plt.plot(X_tr,y_pred_tr,'-r',markersize=10,alpha=1 );
# -
from ipywidgets import interact, HTML, FloatSlider
interact(polynomial_regr,degree=(1,25,1));
# Isn't this interactive and fun?
| SETUP/Verify/jupyter_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: py3
# ---
# # Lab 09 - Reinforcement Learning
#
# In reinforcement learning (RL), an "agent" chooses actions given information about the present state of the system, and is given a reward (a measure of how good the action was). The aim is to maximize the cumulative returned reward over many interactions with the system.
#
# The problem is defined as a series of state transitions consisting of: State --> Action --> New State + reward.
#
# Multiple sequences make up an "episode" consisting of N state transitions.
#
# </pre>
# <img src="notebook_images/RL.png" width="600">
# <pre>
#
#
#
# In this example, we'll look at the Deep Deterministic Policy Gradient (DDPG) algorithm (see paper [here](https://arxiv.org/abs/1509.02971)), for two cases:
# 1. The standard **pendulum problem** (balancing a pendulum upright by applying torque):
# - see problem details here https://github.com/openai/gym/wiki/Pendulum-v0
# - see examples of performance of different agents: https://github.com/openai/gym/wiki/Leaderboard#pendulum-v0
#
#
#
#
# 2. **Tuning 3 quadrupoles in an accelerator lattice to minimize beam size** (same setup as Lab01):
# - The actions are the changes in quad settings
# - The state is the present RMS beamsize output and quad settings
#
#
#
# <img src="notebook_images/triplet.png" width="600">
#
#
#
# DDPG is an example of actor-critic RL where a mapping from observed to actions (i.e. a "policy" or "actor") and an estimate of the long-term value of taking actions in a given state (i.e. a "critic") are both learned functions parameterized by neural networks. The critic provides the training signal for the actor, and the returned rewards provide the training signal for the critic.
#
#
#
#
# [Jump to pendulum section here ](#pendulum)
#
# [Jump to beam size section here ](#beamsize)
#
# # Set up environment
import os, sys
sys.path.append(os.getcwd())
# !pip install git+https://github.com/uspas/2021_optimization_and_ml --quiet
# !pip install gym
# +
# %reset -f
import numpy as np
import json
import time
import pickle as pickle
import matplotlib.pyplot as plt
from IPython.display import clear_output
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import torch.autograd as autograd
import random
from collections import deque
import gym
from gym import error, spaces, utils
from gym.utils import seeding
from os import path
#import toy accelerator package
from uspas_ml.accelerator_toy_models import simple_lattices
from uspas_ml.utils import transformer
# %matplotlib inline
# -
# # Take a look at the DDPGAgent class below:
# +
from ddpg.models import Critic, Actor
from common.exp_replay import ExperienceReplayLog
from common.noise import OUNoise
class DDPGAgent:
def __init__(self, env, gamma, tau, buffer_maxlen, critic_learning_rate, actor_learning_rate, max_action = 1):
self.device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
self.env = env
self.obs_dim = env.observation_space.shape[0]
self.action_dim = env.action_space.shape[0]
self.noise = OUNoise(env.action_space)
self.iter = 0.0
self.noisy = False
self.max_action= max_action
#print(self.action_dim)
#print(self.obs_dim)
# RL hyperparameters
self.env = env
self.gamma = gamma
self.tau = tau
# Initialize critic and actor networks
self.critic = Critic(self.obs_dim, self.action_dim).to(self.device)
self.critic_target = Critic(self.obs_dim, self.action_dim).to(self.device)
self.actor = Actor(self.obs_dim, self.action_dim,self.max_action).to(self.device)
self.actor_target = Actor(self.obs_dim, self.action_dim).to(self.device)
# Copy target network paramters for critic
for target_param, param in zip(self.critic_target.parameters(), self.critic.parameters()):
target_param.data.copy_(param.data)
# Set Optimization algorithms
self.critic_optimizer = optim.Adam(self.critic.parameters(), lr=critic_learning_rate)
self.actor_optimizer = optim.Adam(self.actor.parameters(), lr=actor_learning_rate)
self.replay_buffer = ExperienceReplayLog(buffer_maxlen)
def get_action(self, obs):
#print('obs;',obs)
if self.noisy == True:
state = torch.FloatTensor(obs).unsqueeze(0).to(self.device)
action = self.actor.forward(state)
action = action.squeeze(0).cpu().detach().numpy()
action = self.noise.get_action(action,self.iter)
self.iter = self.iter+1
else:
state = torch.FloatTensor(obs).unsqueeze(0).to(self.device)
action = self.actor.forward(state)
action = action.squeeze(0).cpu().detach().numpy()
return action
def update(self, batch_size):
#Batch updates
states, actions, rewards, next_states, _ = self.replay_buffer.sample(batch_size)
state_batch, action_batch, reward_batch, next_state_batch, masks = self.replay_buffer.sample(batch_size)
state_batch = torch.FloatTensor(state_batch).to(self.device)
action_batch = torch.FloatTensor(action_batch).to(self.device)
reward_batch = torch.FloatTensor(reward_batch).to(self.device)
next_state_batch = torch.FloatTensor(next_state_batch).to(self.device)
masks = torch.FloatTensor(masks).to(self.device)
# Q info updates
curr_Q = self.critic.forward(state_batch, action_batch)
next_actions = self.actor_target.forward(next_state_batch)
next_Q = self.critic_target.forward(next_state_batch, next_actions.detach())
expected_Q = reward_batch + self.gamma * next_Q
# Update Critic network
q_loss = F.mse_loss(curr_Q, expected_Q.detach())
self.critic_optimizer.zero_grad()
q_loss.backward()
self.critic_optimizer.step()
# Update Actor network
policy_loss = -self.critic.forward(state_batch, self.actor.forward(state_batch)).mean()
self.actor_optimizer.zero_grad()
policy_loss.backward()
self.actor_optimizer.step()
# Update Actor and Critic target networks
for target_param, param in zip(self.actor_target.parameters(), self.actor.parameters()):
target_param.data.copy_(param.data * self.tau + target_param.data * (1.0 - self.tau))
for target_param, param in zip(self.critic_target.parameters(), self.critic.parameters()):
target_param.data.copy_(param.data * self.tau + target_param.data * (1.0 - self.tau))
# -
# <a id='pendulum'></a>
# # Pendulum
#
# Take a look at how the Pendulum environment is defined in openAI gym below (this is the standard openAI gym implementation, and we've added additional helper functions for plotting)
# +
class PendulumEnv(gym.Env):
metadata = {
'render.modes': ['human', 'rgb_array'],
'video.frames_per_second': 30
}
def __init__(self, g=10.0):
self.max_speed = 8
self.max_torque = 2.
self.dt = .05
self.g = g
self.m = 1.
self.l = 1.
self.viewer = None
#define action and observation space
high = np.array([1., 1., self.max_speed], dtype=np.float32)
self.action_space = spaces.Box(
low=-self.max_torque,
high=self.max_torque, shape=(1,),
dtype=np.float32
)
self.observation_space = spaces.Box(
low=-high,
high=high,
dtype=np.float32
)
self.seed()
#lists for plotting
self.q1s=[]
self.costs=[]
self.reset_ep=[]
self.plot = True
def log_results(self,q11, reward, reset_ep_flag = False):
'''
logs the results of a given iteration, for quad inputs, the reward, and whether the episode was resett
'''
self.costs.append(reward)
self.q1s.append(q11)
if reset_ep_flag == True:
self.reset_ep.append(1)
else:
self.reset_ep.append(0)
def plot_results(self,):
'''
plots the results from the logged values
'''
clear_output(wait=True)
f = plt.figure(figsize=(25,3))
ax = f.add_subplot(141)
ax2 = f.add_subplot(142)
plot_reset = np.where(np.array(self.reset_ep)==1)[0]
for i in range(0, len(plot_reset)):
ax.axvline(x=plot_reset[i],alpha=0.1,color='k')
ax2.axvline(x=plot_reset[i],alpha=0.1,color='k')
ax.plot(self.q1s,'.')
ax.set_ylabel('action',fontsize=12)
ax2.plot(self.costs, 'k.')
ax2.set_ylabel('reward',fontsize=12)
ax.set_xlabel('Iteration',fontsize=12)
ax2.set_xlabel('Iteration',fontsize=12)
plt.show();
def reset_plot(self,):
'''
resets the logs and the plot
'''
self.costs=[]
self.q1s=[]
self.reset_ep=[]
def save_plot(self,name = 'mon_'):
'''
saves results from the logged values to a json file
'''
run_details = {
'q1': self.q1s,
'costs': self.costs,
'reset_ep': self.reset_ep,
}
with open(name + '.json', 'w') as json_file:
json.dump(run_details, json_file, default = to_serializable)
def seed(self, seed=None):
self.np_random, seed = seeding.np_random(seed)
return [seed]
def step(self, u):
th, thdot = self.state # th := theta
g = self.g
m = self.m
l = self.l
dt = self.dt
u = np.clip(u, -self.max_torque, self.max_torque)[0]
self.last_u = u # for rendering
costs = angle_normalize(th) ** 2 + .1 * thdot ** 2 + .001 * (u ** 2)
newthdot = thdot + (-3 * g / (2 * l) * np.sin(th + np.pi) + 3. / (m * l ** 2) * u) * dt
newth = th + newthdot * dt
newthdot = np.clip(newthdot, -self.max_speed, self.max_speed)
self.log_results(u, -costs, reset_ep_flag = False)
if self.plot == True:
self.plot_results()
else:
clear_output()
self.state = np.array([newth, newthdot])
return self._get_obs(), -costs, False, {}
def reset(self):
high = np.array([np.pi, 1])
self.state = self.np_random.uniform(low=-high, high=high)
self.last_u = None
self.log_results(np.nan, np.nan, reset_ep_flag = True)
if self.plot == True:
self.plot_results()
else:
clear_output()
return self._get_obs()
def _get_obs(self):
theta, thetadot = self.state
return np.array([np.cos(theta), np.sin(theta), thetadot])
def render(self, mode='human'):
if self.viewer is None:
from gym.envs.classic_control import rendering
self.viewer = rendering.Viewer(500, 500)
self.viewer.set_bounds(-2.2, 2.2, -2.2, 2.2)
rod = rendering.make_capsule(1, .2)
rod.set_color(.8, .3, .3)
self.pole_transform = rendering.Transform()
rod.add_attr(self.pole_transform)
self.viewer.add_geom(rod)
axle = rendering.make_circle(.05)
axle.set_color(0, 0, 0)
self.viewer.add_geom(axle)
fname = path.join(path.dirname(__file__), "assets/clockwise.png")
self.img = rendering.Image(fname, 1., 1.)
self.imgtrans = rendering.Transform()
self.img.add_attr(self.imgtrans)
self.viewer.add_onetime(self.img)
self.pole_transform.set_rotation(self.state[0] + np.pi / 2)
if self.last_u:
self.imgtrans.scale = (-self.last_u / 2, np.abs(self.last_u) / 2)
return self.viewer.render(return_rgb_array=mode == 'rgb_array')
def close(self):
if self.viewer:
self.viewer.close()
self.viewer = None
def angle_normalize(x):
return (((x+np.pi) % (2*np.pi)) - np.pi)
# -
# #### Initialize the agent and start training
# +
from common.utils import mini_batch_train
from ddpg.ddpg import DDPGAgent
#initialize environment
env = PendulumEnv() # alternative: gym.make("Pendulum-v0")
max_episodes = 10 #total number of episodes to stop training after
max_steps = 100 #max number of steps per episode
batch_size = 300 #batch size for updates
buffer_maxlen = 100000 #max buffer size
# define training hyperparameters
gamma = 0.99 #discount factor
tau = 1e-2 #for updates with target network
critic_lr = 1e-3 #learning rate
actor_lr = 1e-3 #learning rate
#initialize agent
agent = DDPGAgent(env, gamma, tau, buffer_maxlen, critic_lr, actor_lr, max_action = 2) #max_action is set by what the environment expects
# -
#train with mini-batches and plotting for 10 episodes
max_episodes = 5
episode_rewards, env = mini_batch_train(env, agent, max_episodes, max_steps, batch_size)
# #### Question: how does the agent performance look early in training? What would you expect to see?
# Now continue trainining without plotting (plotting slows down execution)
#faster training without plotting for 50 more episodes
env.plot = False
max_episodes = 50
episode_rewards, env = mini_batch_train(env, agent, max_episodes, max_steps, batch_size)
#plot the results of that previous training
env.plot_results()
# #### Question: how does the agent performance look as training progresses? What would you expect to see?
# Now examine the agent's behavior at present for 5 episodes below
#rest plot and see how training is going with live plotting
env.reset_plot()
env.plot = True
max_episodes = 5
episode_rewards, env = mini_batch_train(env, agent, max_episodes, max_steps, batch_size)
#continue training
env.plot = False
max_episodes = 90
episode_rewards, env = mini_batch_train(env, agent, max_episodes, max_steps, batch_size)
env.plot_results()
#rest plot and see how training is going with live plotting
env.reset_plot()
env.plot = True
max_episodes = 5
episode_rewards, env = mini_batch_train(env, agent, max_episodes, max_steps, batch_size)
# #### Question: how does the agent performance look after a lot of training?
# How do the actions it takes and the reward obtained in each episode compare to early on in training?
#
# <a id='beamsize'></a>
| labs/lab_09/lab_09.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# default_exp lightcurve
# %load_ext autoreload
from nbdev.showdoc import show_doc
# !date
# # Light Curve Generation
# > Generate light curves from cells
# A "light curve" is a table with columns
#
# - `t` -- time in MJD
# - `tw` -- the cell width
# - `n` -- number of photons
# - `e` -- exposure, in $10^6 s\ cm^2$ units
# - `fit` -- the fit object, containing info about the flux. This is usually a `PoissonRep` object
#
# It is generated from a set of cells
#
#
# export
import sys
import numpy as np
import pylab as plt
import pandas as pd
from wtlike.config import *
from wtlike.sources import PointSource
from wtlike.loglike import (LogLike, GaussianRep, Gaussian2dRep, PoissonRep, PoissonRepTable, poisson_tolerance)
from wtlike.cell_data import *
# export
class CellFitter(object):
""" Apply likelihood fits to a set of cells
parameters:
- cells : a table with index t, columns tw, n, e, w, S, B
- rep_name : represention to use
Generates a DataFrame with columns n, ep, fit
"""
rep_class = [GaussianRep, Gaussian2dRep, PoissonRep, PoissonRepTable]
rep_list = 'gauss gauss2d poisson poissontable'.split()
def __init__(self, config,
cells,
source_name,
rep_name: 'likelihood rep name'='',
):
"""Apply fits to the likelihoods for a set of cells
"""
self.source_name = source_name
self.config=config
self.cells = cells
# generate a list of LogLike objects for each
cells.loc[:,'loglike'] = cells.apply(LogLike, axis=1)
# analyze using selected rep
rep_name = rep_name or config.likelihood_rep
if rep_name not in self.rep_list:
raise Exception(f'Unrecognized rep: "{rep_name}", must be one of {self.rep_list}')
repcl = self.rep_class[self.rep_list.index(rep_name)]
if config.verbose>1:
print(f'Fitting likelihoods with {rep_name} representation')
# making output with reduced columns
self.ll_fits = cells['t tw n e S'.split()].copy() # ignore ctm?
# add summary of weights, which can be used for fast evaluation if log approx is ok.
self.ll_fits.loc[:,'wsum'] = cells.w.apply(lambda w: np.sum(w))
self.ll_fits.loc[:,'w2sum'] = cells.w.apply(lambda w: np.sum(w**2))
self.ll_fits.loc[:,'wmax'] = cells.w.apply(lambda w: np.max(w))
try:
self.ll_fits.loc[:,'fit'] = cells.loglike.apply(repcl)
except Exception as e:
print(f'Failed a fit: \n{e}', file=sys.stderr)
raise
def __repr__(self):
return f'{self.__class__.__name__}: source "{self.source_name}" fit with {len(self.ll_fits)} cells'
def full_likelihood():
return LogLike(concatenate_cells(self.cells))
@property
def dataframe(self):
return self.ll_fits
# export
def fit_cells(config,
input_cells,
repcl = PoissonRep,
):
"""Apply fits to the likelihoods for a set of cells
return light-curve dataframe
"""
global poisson_tolerance
# select the set of cells
cells = input_cells.copy()
poisson_tolerance = config.poisson_tolerance
# generate a list of LogLike objects for each
cells.loc[:,'loglike'] = cells.apply(LogLike, axis=1)
if config.verbose>0:
print(f'LightCurve: Loaded {len(cells)} / {len(input_cells)} cells for fitting')
# making output with reduced columns
ll_fits = cells['t tw n e'.split()].copy()
ll_fits.loc[:,'fit'] = cells.loglike.apply(repcl)
return ll_fits
# +
# export
def getCellFitter(config, source, cell_query='e>1e-6', key=''):
"""Returns a lightcurve table for the source
- `source` -- a PointSource object
- `cell query` -- query to apply to cell selection
- `bin_edges` -- optional to select other than default described in config
- `key` -- optional cache key. Set to None to disable cache use
"""
def doit():
cell_data = CellData(config, source)
cells = cell_data.df.query(cell_query)
assert len(cells)>0, 'No cells from CellData after query'
lc = CellFitter(config, cells, source).dataframe
return lc
# if bin_edges is None:
# use cache only with default bins
key = f'lightcurve_{source_name}' if key=='' else key
description = f'Light curve with {cell_query} for {source_name}' if config.verbose>0 and key is not None else ''
return config.cache(key, doit, description=description)
# else:
# return doit()
# -
# export
def fit_table(lc, expect=1.0):
"""Generate a summary table from a light curve dataframe"""
fits = lc.fit
flux = fits.apply(lambda f: f.flux)
errors = fits.apply(lambda f: (round(f.errors[0]-f.flux,3), rorebinnedund(f.errors[1]-f.flux ,3) ) )
sigma_dev = fits.apply(lambda f: round(f.poiss.sigma_dev(expect),1) )
df = lc['t tw n e'.split()].copy() # maybe fix warnings?
df.loc[:,'flux'] = flux.values.round(4)
df.loc[:, 'errors'] = errors.values
df.loc[:, 'sigma_dev'] = sigma_dev.values
df.loc[:, 'limit'] = fits.apply(lambda f: f.limit)
return df
# export
def flux_plot(cell_fits,
query='',
ax=None, fignum=1, figsize=(12,4),
log=False,
title=None,
step=False,
tzero:'time offset'=0,
flux_factor=1,
colors=('cornflowerblue','sandybrown', 'blue'),
fmt='', ms=None, error_pixsize=10,
limit_fmt=None,
source_name=None,
legend_loc='upper left',
error_lw=2,
ts_bar_min=4,
zorder=0,
errorbar_args={},
axhline={},
**kwargs):
"""Make a plot of flux vs. time. This is invoked by the `plot` function of `LightCurve`
- cell_fits -- cell fits DataFrame
- query ['']-- DataFrame query to select subset
- ts_bar_min -- threshold for plotting as bar vs limit
- tzero -- time offset, in MJD
- source_name -- draw text in top left
- flux_factor [1]
- ax [None] -- a matplotlib.axes._subplots.AxesSubplot object returned from plt.subplots<br>
if None, create one using subplots with fignum [1] and figsize [(12,4)]
- fmt [''] -- marker symbol -- if not specified, will use '.' if many bins, else 'o'
- ms [None] -- for marker size
- colors -- tuple of colors for signal, limit, step
- step -- add a "step" plot
- zorder -- set to different number to order successive calls with same Axis object
- kwargs -- apply to the `ax` object, like xlim, ylim
returns the Figure instance
"""
def error_size(ax, x,y, ):
topix = ax.transData.transform
frompix = ax.transData.inverted().transform
sizes = []
for a,b in zip(x,y):
sx, sy = topix((a,b))
c,d = frompix((sx, sy-error_pixsize))
sizes.append(b-d)
return np.array(sizes)
import matplotlib.ticker as ticker
label = kwargs.pop('label', None)
step_label = kwargs.pop('step_label', None)
#error_size=kwargs.pop('error_size', 2)
# errorbar_args.update(error_size=kwargs.pop('error_size', 2))
fig, ax = plt.subplots(figsize=figsize, num=fignum) if ax is None else (ax.figure, ax)
# key words for the Axes object
kw=dict(xlabel='MJD'+ f' - {tzero} [{UTC(tzero)[:10]}]' if tzero else 'MJD' ,
ylabel='Relative flux',
yscale='log' if log else 'linear',
)
kw.update(**kwargs)
ax.set(**kw)
ax.set_title(title) # or f'{source_name}, rep {self.rep}')
ax.grid(alpha=0.5)
# potential yaxis formatting
if kw['yscale']=='log' and flux_factor==1:
ax.yaxis.set_major_formatter(ticker.FuncFormatter(
lambda val,pos: { 1.0:'1', 10.0:'10', 100.:'100'}.get(val,'')))
# select data to plot
df = cell_fits.copy()
df.loc[:,'ts'] = df.fit.apply(lambda f: f.ts)
if query:
df = df.query(query)
if fmt=='':
fmt='.' if len(df)>200 else 'o'
limit = df.ts<ts_bar_min
bar = df.loc[~limit,:]
lim = df.loc[limit,:]
allflux= np.select([~limit, limit],
[df.fit.apply(lambda f: f.flux).values,
df.fit.apply(lambda f: f.limit).values],
) * flux_factor
# do the limits first (only for poisson rep)
if len(lim)>0:
t = lim.t-tzero
tw = lim.tw
color = colors[2 if step else 1]
y = allflux[limit]
if limit_fmt is None:
# try to draw an error bar, hard to determine size
yerr = error_size(ax, t, y) #0.2*(1 if kw['yscale']=='linear' else y)#*flux_factor
ax.errorbar(x=t, y=y, xerr=tw/2,
yerr=yerr, color=color ,
uplims=True, ls='',
ms=ms, lw=error_lw, capsize=2*error_lw, capthick=0,
zorder=zorder, label=' 95% limit', **errorbar_args)
else:
# just a symbol, like 'v'
ax.errorbar(x=t,xerr=tw/2, y=y, fmt=limit_fmt, color=color,
zorder=zorder, label=' 95% limit', **errorbar_args)
# then the points with error bars
t = bar.t.values-tzero
tw = bar.tw.values.astype(float)
fluxmeas = allflux[~limit]
upper = bar.fit.apply(lambda f: f.errors[1]).values * flux_factor
lower = bar.fit.apply(lambda f: f.errors[0]).values * flux_factor
error = np.array([upper-fluxmeas, fluxmeas-lower])
# if label is None:
# label = f'{bin_size_name(round(tw.mean(),4))} bins' if np.std(tw)<1e-6 else ''
ax.errorbar(
x=t, xerr=tw/2, ms=ms,
y=fluxmeas, yerr=error, lw=2, fmt=fmt,
color=colors[0],
label=label, zorder=zorder+1,
**errorbar_args)
# finally overlay the step if requested
if step:
t = df.t.values-tzero
xerr = df.tw.values/2;
x = np.append(t-xerr, [t[-1]+xerr[-1]]);
y = np.append(allflux, [allflux[-1]])
ax.step(x, y,
color=colors[2], where='post', lw=2, label=step_label, zorder=zorder+2)
# the legend: if added overplot, make sure it is below
if legend_loc != 'none':
handles, labels = ax.get_legend_handles_labels()
ax.legend(handles[::-1], labels[::-1],loc=legend_loc, fontsize=10, frameon=False)
#ax.legend()
if source_name is not None:
ax.text(0.99, 0.95, source_name, va='top', ha='right', transform=ax.transAxes,)
return fig
# +
#hide
# for testing this
#flux_plot(lc.fits, figsize=(10,5), log=True, tzero=54720, fmt='D',
# xlim=(0,50), ms=8, ylim=(0.05,2), error_pixsize=20);
# source = PointSource('3C 279')
# lc = LightCurve(Config(), source)
# plt.rc('font', size=14)
# ax =flux_plot(lc.fits, figsize=(15,5) ).axes[0];
# yrs = [str(yr) for yr in range(2008,2022,1)]
# mjd_yrs = [MJD(yr) for yr in yrs]
# ax.set(xticks=mjd_yrs, xticklabels=yrs);
# yrs = [MJD(str(yr)) for yr in range(2008,2022)]
# ax.set(xticks=yrs)
# -
show_doc(flux_plot, title_level=3)
# Example plot-- the first 100 days of Fermi data for Geminga
# And forcing upper limits
# +
# export
def fit_beta_free(self):
r"""Make a plot of $\alpha$ and $\beta$ for the given source
Uses the current cells, assuming Gaussian errors, fitting each to $\alpha$ and $\beta$
"""
cells = self.cells
g2rep = cells.apply( lambda cell: Gaussian2dRep(LogLike(cell)).fit, axis=1)
abdf = pd.DataFrame.from_dict(dict(zip(cells.index,g2rep)), orient='index')
return abdf
def plot_beta_free(self, abdf):
cells = self.cells
fig, (ax1,ax2) = plt.subplots(2,1, figsize=(12,5), sharex=True)
plt.subplots_adjust(hspace=0, top=0.92)
t = cells.t; xerr=cells.tw/2
ax1.errorbar(x=t, xerr=xerr, y=abdf.beta, yerr=abdf.sig_beta, fmt='o');
ax1.grid(alpha=0.5)
ax1.set(ylabel=r'$\beta$', xlabel='MJD', title=rf'{self.source_name}: fits with $\beta$ free');
ax2.errorbar(x=t, xerr=xerr, y=abdf.flux-1, yerr=abdf.sig_flux, fmt='o')
ax2.set(ylabel=r'$\alpha$')
ax2.grid(alpha=0.5);
[ax.axhline(0, ls='--',color='grey') for ax in (ax1,ax2)];
# -
# export
class LightCurve(CellData):
"""Full analysis of a source,
Inherits from `CellData`
"""
def __init__(self, *pars, **kwargs):
exp_min = kwargs.pop('exp_min', None) # corresponds to ~2counts
self.n_min = kwargs.pop('n_min', 2)
self.lc_key = kwargs.pop('lc_key', None)
super().__init__(*pars, **kwargs)
self.exp_min = exp_min if exp_min is not None else self.config.exp_min
self.update()
def __repr__(self):
r = super().__repr__()
r += f'\nLightCurve: {len(self.cells)} cells in view={self.time_bins}'
r += f', {len(self.fits)} fits with exposure > {self.exp_min*self.time_bins[2]}'
return r
def update(self):
# invoked by current, again if a copy
# get the cells from superclass
def doit():
# fit the subset that have enought exposure and counts
exp_min = self.exp_min*self.time_bins[2]
query_string = f'e>{exp_min} & n>{self.n_min}'
fit_cells = self.cells.query(query_string).copy()
if self.config.verbose>0:
print(f'LightCurve: select {len(fit_cells)} cells for fitting with {query_string}' )
assert len(fit_cells)>0, 'No cells from CellData after query'
# the local workhorse
return CellFitter(self.config, fit_cells, self.source_name).dataframe
# use cache only with default bins?
key = f'lightcurve_{source_name}' if self.lc_key=='' else self.lc_key
if key is None:
self.fits = doit()
else:
description = f'Light curve with {cell_query} for {source_name}' if self.config.verbose>0 and key is not None else ''
self.fits = self.config.cache(key, doit, description=description)
def check_plot_kwargs(self, kwargs):
tzero = kwargs.get('tzero', None)
xlim = kwargs.get('xlim', None)
if xlim is not None and tzero is None: # scale from start, stop times if less than start
a,b = xlim
if a<self.start: a = self.start+a if a>=0 else self.stop+a
if b<self.start: b = self.start+b if b>0 else self.stop+b
kwargs['xlim'] = (a,b)
log = kwargs.pop('log', None)
if log is not None and log:
kwargs['yscale'] = 'log'
if kwargs.pop('show_flux', False):
kwargs['flux_factor'] = self.src_flux * 1e6
if kwargs.get('ylabel', None) is None:
kwargs['ylabel'] = 'Photon flux [$\mathrm{10^{-6} cm^{-2} s^{-1}}$]'
utc_flag = kwargs.pop('UTC', False)
if utc_flag:
# even years if True else interpret as int
cnt = 2 if type(utc_flag)==bool else utc_flag
yrs = [str(yr) for yr in range(2008,2023, cnt)] #get this from kwarg maybe
yrkw = dict( xticks=[MJD(yr) for yr in yrs], xticklabels=yrs, xlabel='UTC',)
kwargs.update(**yrkw)
@decorate_with(flux_plot)
def plot(self, **kwargs):
"""Make a light curve plot
Invokes flux_plot, after processing kwargs to intercept
- log -- translate to `xscale='log'`
- xlim [None] -- convert to (start, stop) interpreted relative to start, stop if < start.
- show_flux [False]-- convert y scale to photon flux
- UTC [False] -- convert x scale to UTC years
-
"""
source_name = kwargs.pop('source_name', self.source_name)
# special treatment for log and xlim
self.check_plot_kwargs(kwargs)
fig = flux_plot(self.fits, source_name=source_name,
label=f'{self.step_name} bins', **kwargs)
fig.set_facecolor('white')
return fig
def plot_with_exposure(self, **kwargs):
"""Stacked flux plot flux with exposure
"""
fig,(ax1,ax2) = plt.subplots(2,1, figsize=(15,6), sharex=True);
fig.subplots_adjust(hspace=0)
ax1.tick_params(labelbottom=False)
left, bottom, width, height = (0.15, 0.10, 0.75, 0.85)
fraction = 0.7
ax1.set_position([left, bottom+(1-fraction)*height, width, fraction*height])
ax2.set_position([left, bottom, width, (1-fraction)*height])
tzero = kwargs.get('tzero', 0)
self.plot(ax=ax1, **kwargs)
# the step plot
ax2.step(self.cells.t-tzero, self.cells.e, '-', where='mid')
ax2.grid(alpha=0.5)
ax2.set(ylabel='exposure', ylim=(0,None))
return fig
def flux_table(self, lc=None, include_e=True, include_ctm=True):
"""Generate a summary table from the light curve
- lc -- A light curve fits dataframe; use current one if not specified
- include_e -- include the exposure
"""
if lc is None: lc=self.fits
fits = lc.fit
flux = fits.apply(lambda f: f.flux)
errors = fits.apply(lambda f: (round(f.errors[0]-f.flux,3), round(f.errors[1]-f.flux ,3) ) )
# for future
#sigma_dev = fits.apply(lambda f: round(f.poiss.sigma_dev(expect),1) )
df = lc['t tw n'.split()].copy() # maybe fix warnings?
if include_e:
df.loc[:,'e'] = self.fits.e
if include_ctm and 'ctm' in self.fits:
df.loc[:,'ctm'] = self.fits.ctm
df.loc[:, 'ts'] = fits.apply(lambda f: f.ts).round(1)
df.loc[:,'flux'] = flux.values.round(4)
df.loc[:, 'errors'] = errors.values
#df.loc[:, 'sigma_dev'] = sigma_dev.values
df.loc[:, 'limit'] = fits.apply(lambda f: f.limit)
return df
def fit_beta_free(self, show_plot=True):
r"""Fit the cells to $\alpha$ and $\beta$, assuming Gaussian errors.
Make plots if show_plot is set.
return a DF with the fits.
"""
abdf = fit_beta_free(self)
if show_plot:
plot_beta_free(self, abdf)
return abdf
@property
def fluxes(self):
"""A DataFrame table of the flux measurements"""
return self.flux_table()
def __init_subclass__(cls):
if cls.__doc__ is None:
raise AttributeError("No docstring")
import warnings
warnings.filterwarnings('error', category=RuntimeWarning)
config=Config()
if config.valid:
lc = LightCurve('3C 279', bins=(0,0,7), clear=False );
lc.plot(UTC=True);
# fig=lc.plot( show_flux=True, ylim=(2e-2,4),tzero=54720, fmt='o', xlim=(0,50), log=True, ms=8,
# error_pixsize=20)
#hide
from nbdev.export import notebook2script
notebook2script()
# !date
| nbs/09_lightcurve.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: me
# language: python
# name: me
# ---
# ## Model one trade variables
#
# This notebook extracts the selected trade variables in the `indicator_list` from IMF and World Bank (wb) data sources, and writes them to a csv file.
# +
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
# -
warnings.filterwarnings('ignore')
pd.options.display.float_format = '{:20,.2f}'.format
# | variable | origin | source |granularity|countries| description | composition |
# | --------------------------|-------------------|-------------|-----------|---------|-------------------------------------------------------------|-------------------------------------------------------------------|
# | imports | - | wb econ | yearly | 217 | Imports of goods and services (% of GDP) | - |
# | manufacturing value | - | wb econ | yearly | 217 | Manufacturing, value added (% of GDP) | - |
# | consumption per capita | - | wb econ | yearly | 217 | Households and NPISHs Final consumption expenditure per capita (2010 USD)| - |
# | food exports | - | wb econ | yearly | 217 | Food exports (% of merchandise exports) | - |
# | energy imports | - | wb econ | yearly | 217 | Energy imports, net (% of energy use) | - |
# | consumption per capita | - | wb econ | yearly | 217 | Households and NPISHs Final consumption expenditure per capita (constant 2010 USD)| - |
# | GNI per capita | - | wb econ | yearly | 217 | GNI per capita (constant 2010 USD) | - |
# | food imports | - | wb econ | yearly | 217 | Food imports (% of merchandise imports) | - |
# | manufacturing value USD | - | wb econ | yearly | 217 | Manufacturing, value added (constant 2010 USD) | - |
# | exports | - | wb econ | yearly | 217 | Exports of goods and services (% of GDP) | - |
# | trade share of GDP | - | wb econ | yearly | 217 | Trade (% of GDP) | - |
# | high-tech exports | - | wb econ | yearly | 217 | High-technology exports (% of manufactured exports) | - |
# | imports USD | - | wb econ | yearly | 217 | Imports of goods and services (constant 2010 USD) | - |
# | exports USD | - | wb econ | yearly | 217 | Exports of goods and services (constant 2010 USD) | - |
# | services trade | - | wb econ | yearly | 217 | Trade in services (% of GDP) | - |
indicator_list = ['Imports of goods and services (% of GDP)', 'Manufacturing, value added (% of GDP)',
'Households and NPISHs Final consumption expenditure per capita (constant 2010 US$)',
'Food exports (% of merchandise exports)', 'Energy imports, net (% of energy use)',
'GNI per capita (constant 2010 US$)', 'Food imports (% of merchandise imports)',
'Manufacturing, value added (constant 2010 US$)', 'Exports of goods and services (% of GDP)',
'Trade (% of GDP)', 'High-technology exports (% of manufactured exports)',
'Imports of goods and services (constant 2010 US$)', 'Trade in services (% of GDP)',
'Exports of goods and services (constant 2010 US$)']
len(indicator_list)
# ## Load imf monthly data
# + language="bash"
# wc -l imf/*.csv
# -
time_values = [str('%sM%s' % (y, m)) for m in list(range(1, 13)) for y in list(range(1960, 2018))]
imf_columns = ['Country Name', 'Indicator Name'] + time_values
imf_country_aggregates = ['Euro Area']
def load_imf_monthly(file_name, indicators, imf_columns, country_aggregates):
csv_df = pd.read_csv('imf/%s' % file_name).fillna(0)
base_df = csv_df.loc[csv_df['Attribute'] == 'Value'].drop(columns=['Attribute'])
monthly_df = base_df.loc[(base_df['Indicator Name'].isin(indicators))]
imf_df = monthly_df[imf_columns].fillna(0)
df = pd.melt(imf_df, id_vars=['Country Name', 'Indicator Name'], var_name='date', value_name='value')
df['date'] = pd.to_datetime(df['date'], format='%YM%m')
df.columns = ['country', 'indicator', 'date', 'value']
return df.loc[~df['country'].isin(country_aggregates)]
imf_pplt_df = load_imf_monthly('PPLT_11-25-2018 19-25-01-32_timeSeries.csv', indicator_list, imf_columns, imf_country_aggregates)
imf_cpi_df = load_imf_monthly('CPI_11-25-2018 19-14-47-26_timeSeries.csv', indicator_list, imf_columns, imf_country_aggregates)
imf_df = pd.concat([imf_cpi_df, imf_pplt_df], join='outer')
imf_df.size
imf_df.head(15)
len(imf_df['country'].unique())
imf_countries = sorted(list(imf_df['country'].unique()))
# ### Load world bank yearly data
# + language="bash"
# wc -l world_bank/*.csv
# -
wb_country_aggregates = ['nan', 'Lower middle income', 'Post-demographic dividend', 'High income',
'Pre-demographic dividend', 'East Asia & Pacific (IDA & IBRD countries)',
'Europe & Central Asia (excluding high income)', 'Heavily indebted poor countries (HIPC)',
'Caribbean small states', 'Pacific island small states', 'Middle income',
'Late-demographic dividend', 'OECD members', 'IDA & IBRD total', 'Not classified',
'East Asia & Pacific (excluding high income)',
'Latin America & the Caribbean (IDA & IBRD countries)', 'Low income', 'Low & middle income',
'IDA blend', 'IBRD only', 'Sub-Saharan Africa (excluding high income)',
'Fragile and conflict affected situations', 'Europe & Central Asia (IDA & IBRD countries)',
'Euro area', 'Other small states', 'Europe & Central Asia', 'Arab World',
'Latin America & Caribbean (excluding high income)',
'Sub-Saharan Africa (IDA & IBRD countries)', 'Early-demographic dividend', 'IDA only',
'Small states', 'Middle East & North Africa (excluding high income)', 'East Asia & Pacific',
'South Asia', 'European Union', 'Least developed countries: UN classification',
'Middle East & North Africa (IDA & IBRD countries)', 'Upper middle income',
'South Asia (IDA & IBRD)', 'Central Europe and the Baltics', 'Sub-Saharan Africa',
'Latin America & Caribbean', 'Middle East & North Africa', 'IDA total', 'North America',
'Last Updated: 11/14/2018', 'Data from database: World Development Indicators', 'World']
wb_cols = ['Country Name', 'Series Name'] + [str('%s [YR%s]' % (y, y)) for y in list(range(1960, 2018))]
def load_wb_yearly(file_name, indicators, wb_columns, country_aggregates):
csv_df = pd.read_csv('world_bank/%s' % file_name).fillna(0)
base_df = csv_df.loc[(csv_df['Series Name'].isin(indicators))]
wb_df = base_df[wb_columns].fillna(0)
df = pd.melt(wb_df, id_vars=['Country Name', 'Series Name'], var_name='date', value_name='value')
df['date'] = pd.to_datetime(df['date'].map(lambda x: int(x.split(' ')[0])), format='%Y')
df.columns = ['country', 'indicator', 'date', 'value']
return df.loc[~df['country'].isin(country_aggregates)]
wb_econ_df = load_wb_yearly('ECON.csv', indicator_list, wb_cols, wb_country_aggregates)
wb_hnp_df = load_wb_yearly('HNP.csv', indicator_list, wb_cols, wb_country_aggregates)
wb_pop_df = load_wb_yearly('POP.csv', indicator_list, wb_cols, wb_country_aggregates)
wb_df = pd.concat([wb_econ_df, wb_hnp_df, wb_pop_df], join='outer')
wb_df.size
wb_df.head(15)
len(wb_df['country'].unique())
wb_countries = sorted(list(wb_df['country'].unique()))
# ### Combine the two datasets
imf_specific = [country for country in imf_countries if country not in wb_countries]
len(imf_specific)
imf_to_wb_country_map = {
'Afghanistan, Islamic Republic of': 'Afghanistan',
'Armenia, Republic of': 'Armenia',
'Azerbaijan, Republic of': 'Azerbaijan',
'Bahrain, Kingdom of': 'Bahrain',
'China, P.R.: Hong Kong': 'Hong Kong SAR, China',
'China, P.R.: Macao': 'Macao SAR, China',
'China, P.R.: Mainland': 'China',
'Congo, Democratic Republic of': 'Congo, Dem. Rep.',
'Congo, Republic of': 'Congo, Rep.',
'Egypt': 'Egypt, Arab Rep.',
'French Territories: New Caledonia': 'New Caledonia',
'Iran, Islamic Republic of': 'Iran',
'Korea, Republic of': 'Korea, Rep.',
'Kosovo, Republic of': 'Kosovo',
"Lao People's Democratic Republic": 'Lao PDR',
'Serbia, Republic of': 'Serbia',
'Sint Maarten': 'Sint Maarten (Dutch part)',
'Timor-Leste, Dem. Rep. of': 'Timor-Leste',
'Venezuela, Republica Bolivariana de': 'Venezuela, RB',
'Venezuela, República Bolivariana de': 'Venezuela, RB',
'Yemen, Republic of': 'Yemen'
}
imf_df = imf_df.replace({'country': imf_to_wb_country_map})
trade_df = pd.concat([wb_df, imf_df], join='outer')
trade_df.size
trade_df.head(15)
indicators = sorted(list(trade_df['indicator'].unique()))
assert len(indicators) == len(indicator_list), 'The number of retrieved variables (%s) does not match the number of specified variables (%s).\nThe following variables are missing:\n\n %s' % (len(indicators), len(indicator_list), [i for i in indicator_list if i not in indicators])
trade_df.to_csv('model_one/trade.csv', sep=';', index=False)
| dataprep/model_one_trade_variables.ipynb |