code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Containerization with Docker
# ## What is containerization? What are Containers?
#
# Containerization is the process of defining the software components required to run a *specific application*. In the context of a Docker-based containerization process this definition is written into a *Dockerfile*. A Dockerfile is then used to create a *Docker Image*. That Docker Image can then be used as the basis for one or more separate *Docker Containers*, or as the starting point for defining a new Dockerfile. Docker containers are immutable (i.e. changes to the container are **do not** update the source image, and are lost when the container is stopped *and* removed). A modified container can be used to create a new image through the *commit* command.
#
# 
#
# This process of building a new image from a base image plus changes defined in a dockerfile, or changes made to a container based on that image and committed to a new image yields a model in which individual Docker images are made up of stacks of images that have been used to build previous base images.
#
# 
#
#
# ### How is it different from virtualization (i.e. virtual machines)?
#
# You may have encountered references to *virtual machines* (or VMs) as a model for running virtual computer environments within/on computer hardware. A common question that arises when discussing containerization is how it differs from virtualization. The primary difference between the two is that the machine images and corresponding virtual machines that they enable are fully self-contained virtual systems that provide all of the core capabilities (in virtual form) of a dedicated computer, including a full operating system (such as Windows or Liunx [Mac OS is infrequently used as an OS for VMs]) and set of applications running within that operating system. Containerized apps, on the other hand share a common container engine that provides core operating system capabilities that allow the application code within the container to run *as if* it were operating within a fully functional OS and hardware environment.
#
# 
# ## Why would I use containers in my work?
#
# * Portability of **applications**
# * Comparatively lightweight
# * Consistent behavior across systems
# * Broad support
# * Cloud providers
# * [Google Cloud Platform](https://cloud.google.com/containers/)
# * [Amazon Elastic Container Service](https://aws.amazon.com/ecs/)
# * [Microsoft Azure Container Service](https://azure.microsoft.com/en-us/services/container-service/)
# * Desktop operating systems (Docker Engine & additional tools - CE)
# * [Mac OS](https://docs.docker.com/docker-for-mac/install/)
# * [Windows](https://docs.docker.com/docker-for-windows/install/)
# * Servers (CE)
# * Linux
# * [Ubuntu](https://docs.docker.com/install/linux/docker-ce/ubuntu/)
# * [Debian](https://docs.docker.com/install/linux/docker-ce/debian/)
# * [CentOS](https://docs.docker.com/install/linux/docker-ce/centos/)
# * [Fedora](https://docs.docker.com/install/linux/docker-ce/fedora/)
# * You don't need to reinvent the wheel - Many pre-configured containers have been developed and maintained by the community
# * There are numerous container images and documentation maintained in [Docker Hub](https://hub.docker.com) and [Docker Store](https://store.docker.com/search?q=&source=community&type=image) - You can share your images through Docker Hub as sell
# * Many, many Dockerfiles maintained and accessible through [GitHub](https://github.com/search?utf8=✓&q=dockerfile&type=)
# * Containers can provide a foundation for a reproducable environment & workflow
# * Application containers are *imutable* and you can store data in data containers or through mounted directories on the host operating system
# * Multiple containers can be "composed" into combined services
# ## Why would I *not* use containers?
#
# * Your workflows and applications are primarily Windows or Mac OS application based
# * While you can run the Docker platform (i.e. Docker Engine) on a wide variety of operating systems, the applications that you can run in Docker containers are typically Linux based.
# * You typically work with applications for which there are robust installers for your particular operating system.
# ## Sample Use Scenarios
# ### Run a Debian Linux environment
# + language="bash"
# # Pull the "Official" Debian docker image from Dockerhub
# docker pull debian
# -
# Karls-MacBook-Pro-2:~ kbene$ docker run -it debian
# root@e3d7e52268a5:/# cd
# root@e3d7e52268a5:~# touch demo.txt
# root@e3d7e52268a5:~# ls
# demo.txt
# root@e3d7e52268a5:~# exit
# exit
# Karls-MacBook-Pro-2:~ kbene$
#
# Karls-MacBook-Pro-2:~ kbene$ docker run -it debian
# root@5f4a7f5c490f:/# cd
# root@5f4a7f5c490f:~# ls
# root@5f4a7f5c490f:~# exit
# exit
# Karls-MacBook-Pro-2:~ kbene$
# ### Setup and configure a web application
# + language="bash"
# cd httpd
# docker build -t kbene/httpd-local .
# docker run -d -p 8081:80 --name httpd-local kbene/httpd-local
# + language="bash"
# docker ps
# -
# [http://localhost:8081](http://localhost:8081)
# + language="bash"
# docker stop httpd-local
# docker rm httpd-local
# docker ps
# -
# ### Database server
#
# [MySQL Official Image](https://hub.docker.com/_/mysql/)
# + language="bash"
# # Run the MySql server container from the official image in Dockerhub
# docker pull mysql
# docker run --name some-mysql -p 3306:3306 -e MYSQL_USER=demouser -e MYSQL_PASSWORD=<PASSWORD> -e MYSQL_ROOT_PASSWORD=<PASSWORD> -e MYSQL_DATABASE=testdb -d mysql
# + language="bash"
# docker ps
# -
#
# docker exec -it some-mysql bash
#
# root@9098f791534c:/# mysql -u demouser -p
# Enter password:
# Welcome to the MySQL monitor. Commands end with ; or \g.
# Your MySQL connection id is 3
# Server version: 5.7.21 MySQL Community Server (GPL)
#
# Copyright (c) 2000, 2018, Oracle and/or its affiliates. All rights reserved.
#
# Oracle is a registered trademark of Oracle Corporation and/or its
# affiliates. Other names may be trademarks of their respective
# owners.
#
# Type 'help;' or '\h' for help. Type '\c' to clear the current input statement.
#
# mysql> SHOW DATABASES;
# +--------------------+
# | Database |
# +--------------------+
# | information_schema |
# | testdb |
# +--------------------+
# 2 rows in set (0.00 sec)
#
# mysql> QUIT;
# Bye
# root@9098f791534c:/#
# + language="bash"
# docker stop some-mysql
# docker rm some-mysql
# docker ps
# -
# ### Desktop analysis tool - Demo
# ## Docker Resources
#
# * [Documentation Home Page](https://docs.docker.com)
# * [Getting Started with Docker](https://docs.docker.com/get-started/)
# * [Docker Overview](https://docs.docker.com/engine/docker-overview/)
# * [Command Reference](https://docs.docker.com/reference/)
# * [Command Line Interface](https://docs.docker.com/engine/reference/commandline/cli/)
# * [Dockerfile Reference](https://docs.docker.com/engine/reference/builder/)
| 01-Docker Overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + nbsphinx="hidden"
# Delete this cell to re-enable tracebacks
import sys
ipython = get_ipython()
def hide_traceback(exc_tuple=None, filename=None, tb_offset=None,
exception_only=False, running_compiled_code=False):
etype, value, tb = sys.exc_info()
value.__cause__ = None # suppress chained exceptions
return ipython._showtraceback(etype, value, ipython.InteractiveTB.get_exception_only(etype, value))
ipython.showtraceback = hide_traceback
# + nbsphinx="hidden"
# JSON output syntax highlighting
from __future__ import print_function
from pygments import highlight
from pygments.lexers import JsonLexer, TextLexer
from pygments.formatters import HtmlFormatter
from IPython.display import display, HTML
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
def json_print(inpt):
string = str(inpt)
formatter = HtmlFormatter()
if string[0] == '{':
lexer = JsonLexer()
else:
lexer = TextLexer()
return HTML('<style type="text/css">{}</style>{}'.format(
formatter.get_style_defs('.highlight'),
highlight(string, lexer, formatter)))
globals()['print'] = json_print
# -
# ## Using Environments
#
# An [Environment](../api/stix2.environment.rst#stix2.environment.Environment) object makes it easier to use STIX 2 content as part of a larger application or ecosystem. It allows you to abstract away the nasty details of sending and receiving STIX data, and to create STIX objects with default values for common properties.
#
# ### Storing and Retrieving STIX Content
#
# An [Environment](../api/stix2.environment.rst#stix2.environment.Environment) can be set up with a [DataStore](../api/stix2.datastore.rst#stix2.datastore.DataStoreMixin) if you want to store and retrieve STIX content from the same place.
# +
from stix2 import Environment, MemoryStore
env = Environment(store=MemoryStore())
# -
# If desired, you can instead set up an [Environment](../api/stix2.environment.rst#stix2.environment.Environment) with different data sources and sinks. In the following example we set up an environment that retrieves objects from [memory](../api/datastore/stix2.datastore.memory.rst) and a directory on the [filesystem](../api/datastore/stix2.datastore.filesystem.rst), and stores objects in a different directory on the filesystem.
# +
from stix2 import CompositeDataSource, FileSystemSink, FileSystemSource, MemorySource
src = CompositeDataSource()
src.add_data_sources([MemorySource(), FileSystemSource("/tmp/stix2_source")])
env2 = Environment(source=src,
sink=FileSystemSink("/tmp/stix2_sink"))
# -
# Once you have an [Environment](../api/stix2.environment.rst#stix2.environment.Environment) you can store some STIX content in its [DataSinks](../api/stix2.datastore.rst#stix2.datastore.DataSink) with [add()](../api/stix2.environment.rst#stix2.environment.Environment.add):
# +
from stix2 import Indicator
indicator = Indicator(id="indicator--a740531e-63ff-4e49-a9e1-a0a3eed0e3e7",
pattern_type="stix",
pattern="[file:hashes.md5 = 'd41d8cd98f00b204e9800998ecf8427e']")
env.add(indicator)
# -
# You can retrieve STIX objects from the [DataSources](../api/stix2.datastore.rst#stix2.datastore.DataSource) in the [Environment](../api/stix2.environment.rst#stix2.environment.Environment) with [get()](../api/stix2.environment.rst#stix2.environment.Environment.get), [query()](../api/stix2.environment.rst#stix2.environment.Environment.query), [all_versions()](../api/stix2.environment.rst#stix2.environment.Environment.all_versions), [creator_of()](../api/stix2.datastore.rst#stix2.datastore.DataSource.creator_of), [related_to()](../api/stix2.datastore.rst#stix2.datastore.DataSource.related_to), and [relationships()](../api/stix2.datastore.rst#stix2.datastore.DataSource.relationships) just as you would for a [DataSource](../api/stix2.datastore.rst#stix2.datastore.DataSource).
print(env.get("indicator--a740531e-63ff-4e49-a9e1-a0a3eed0e3e7").serialize(pretty=True))
# ### Creating STIX Objects With Defaults
#
# To create STIX objects with default values for certain properties, use an [ObjectFactory](../api/stix2.environment.rst#stix2.environment.ObjectFactory). For instance, say we want all objects we create to have a ``created_by_ref`` property pointing to the ``Identity`` object representing our organization.
# +
from stix2 import Indicator, ObjectFactory
factory = ObjectFactory(created_by_ref="identity--311b2d2d-f010-4473-83ec-1edf84858f4c")
# -
# Once you've set up the [ObjectFactory](../api/stix2.environment.rst#stix2.environment.ObjectFactory), use its [create()](../api/stix2.environment.rst#stix2.environment.ObjectFactory.create) method, passing in the class for the type of object you wish to create, followed by the other properties and their values for the object.
ind = factory.create(Indicator,
pattern_type="stix",
pattern="[file:hashes.md5 = 'd41d8cd98f00b204e9800998ecf8427e']")
print(ind.serialize(pretty=True))
# All objects we create with that [ObjectFactory](../api/stix2.environment.rst#stix2.environment.ObjectFactory) will automatically get the default value for ``created_by_ref``. These are the properties for which defaults can be set:
#
# - ``created_by_ref``
# - ``created``
# - ``external_references``
# - ``object_marking_refs``
#
# These defaults can be bypassed. For example, say you have an [Environment](../api/stix2.environment.rst#stix2.environment.Environment) with multiple default values but want to create an object with a different value for ``created_by_ref``, or none at all.
# +
factory2 = ObjectFactory(created_by_ref="identity--311b2d2d-f010-4473-83ec-1edf84858f4c",
created="2017-09-25T18:07:46.255472Z")
env2 = Environment(factory=factory2)
ind2 = env2.create(Indicator,
created_by_ref=None,
pattern_type="stix",
pattern="[file:hashes.md5 = 'd41d8cd98f00b204e9800998ecf8427e']")
print(ind2.serialize(pretty=True))
# -
ind3 = env2.create(Indicator,
created_by_ref="identity--962cabe5-f7f3-438a-9169-585a8c971d12",
pattern_type="stix",
pattern="[file:hashes.md5 = 'd41d8cd98f00b204e9800998ecf8427e']")
print(ind3.serialize(pretty=True))
# For the full power of the Environment layer, create an [Environment](../api/stix2.environment.rst#stix2.environment.Environment) with both a [DataStore](../api/stix2.datastore.rst#stix2.datastore.DataStoreMixin)/[Source](../api/stix2.datastore.rst#stix2.datastore.DataSource)/[Sink](../api/stix2.datastore.rst#stix2.datastore.DataSink) and an [ObjectFactory](../api/stix2.environment.rst#stix2.environment.ObjectFactory):
# +
environ = Environment(ObjectFactory(created_by_ref="identity--311b2d2d-f010-4473-83ec-1edf84858f4c"),
MemoryStore())
i = environ.create(Indicator,
pattern_type="stix",
pattern="[file:hashes.md5 = 'd41d8cd98f00b204e9800998ecf8427e']")
environ.add(i)
print(environ.get(i.id).serialize(pretty=True))
| docs/guide/environment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="PUB0cvWV2d9q" colab_type="text"
# <div>
# <center>
# <img src="https://drive.google.com/uc?id=1IHXf1W23kjHNsPR4pt61pWa0-43R7BL1" width="400"/>
# </center>
# </div>
#
# **<NAME>, <NAME>**. [Predicting mRNA abundance directly from genomic sequence using deep convolutional neural networks](https://doi.org/10.1101/416685). 2020. *Cell Reports*. [*Github*](https://github.com/vagarwal87/Xpresso). [*Website*](https://xpresso.gs.washington.edu).
# + [markdown] id="ycvRpM20OO0e" colab_type="text"
# # Setup Xpresso dependencies, datasets, and imports
#
# + id="1cLMEYVUOWQU" colab_type="code" outputId="b822a1d8-ba5e-4459-a57f-c781259c6b4b" colab={"base_uri": "https://localhost:8080/", "height": 1000}
#Run with Python3/GPU-enabled runtime
#Train/Valid/Test set for predicting median gene expression levels in the human
# !wget -r -np -nH --reject "index.html*" --cut-dirs 6 \
# https://krishna.gs.washington.edu/content/members/vagar/Xpresso/data/datasets/pM10Kb_1KTest/
#Train/Valid/Test set for predicting median gene expression levels in the mouse
# !wget -r -np -nH --reject "index.html*" --cut-dirs 6 \
# https://krishna.gs.washington.edu/content/members/vagar/Xpresso/data/datasets/pM10Kb_1KTest_Mouse/
#Prepare set of input sequences to generate predictions
# !wget https://xpresso.gs.washington.edu/data/Xpresso-predict.zip
# !unzip Xpresso-predict.zip
#set up dependencies
# !pip install biopython
# !pip install hyperopt
# %tensorflow_version 1.x
#set up imports
import tensorflow as tf
import sys, gzip, h5py, pickle, os
import numpy as np
import pandas as pd
from mimetypes import guess_type
from Bio import SeqIO
from functools import partial
from scipy import stats
from IPython.display import Image
from tensorflow import keras
from keras.models import Model, load_model
from keras.utils.vis_utils import plot_model
from keras.optimizers import Adam, SGD
from keras.layers import *
from keras.metrics import *
from keras.callbacks import Callback, ModelCheckpoint, EarlyStopping
from hyperopt import hp, STATUS_OK
print("TF version", tf.__version__)
print("Keras version", keras.__version__)
device_name = tf.test.gpu_device_name()
if device_name != '/device:GPU:0':
raise SystemError('GPU device not found')
print('Found GPU at: {}'.format(device_name))
# + [markdown] id="mgTWlzY7Nin4" colab_type="text"
# # Train Xpresso model for human data
# + id="Kkxgr6A-OL-8" colab_type="code" outputId="1556bf03-3440-4756-9db3-91214302dece" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# Results presented in the paper are the best of 10 independent trials, choosing the one that minimizes
# validation mean squared error loss.
# These results are not exactly those shown in paper due to variability in performance
global X_trainhalflife, X_trainpromoter, y_train, X_validhalflife, X_validpromoter, y_valid, X_testhalflife, X_testpromoter, y_test, geneName_test, params, best_file
def main(datadir):
global X_trainhalflife, X_trainpromoter, y_train, X_validhalflife, X_validpromoter, y_valid, X_testhalflife, X_testpromoter, y_test, geneName_test, params
params['datadir'] = datadir
trainfile = h5py.File(os.path.join(datadir, 'train.h5'), 'r')
X_trainhalflife, X_trainpromoter, y_train, geneName_train = trainfile['data'], trainfile['promoter'], trainfile['label'], trainfile['geneName']
validfile = h5py.File(os.path.join(datadir, 'valid.h5'), 'r')
X_validhalflife, X_validpromoter, y_valid, geneName_valid = validfile['data'], validfile['promoter'], validfile['label'], validfile['geneName']
testfile = h5py.File(os.path.join(datadir, 'test.h5'), 'r')
X_testhalflife, X_testpromoter, y_test, geneName_test = testfile['data'], testfile['promoter'], testfile['label'], testfile['geneName']
#best hyperparams learned
params = { 'datadir' : datadir, 'batchsize' : 2**7, 'leftpos' : 3000, 'rightpos' : 13500, 'activationFxn' : 'relu', 'numFiltersConv1' : 2**7, 'filterLenConv1' : 6, 'dilRate1' : 1,
'maxPool1' : 30, 'numconvlayers' : { 'numFiltersConv2' : 2**5, 'filterLenConv2' : 9, 'dilRate2' : 1, 'maxPool2' : 10, 'numconvlayers1' : { 'numconvlayers2' : 'two' } },
'dense1' : 2**6, 'dropout1' : 0.00099, 'numdenselayers' : { 'layers' : 'two', 'dense2' : 2, 'dropout2' : 0.01546 } }
print("Using best identified hyperparameters from architecture search, these are:")
print(params)
results = objective(params)
print("Best Validation MSE = %.3f" % results['loss'])
params = {
'tuneMode' : 1,
'batchsize' : 2**hp.quniform('batchsize', 5, 7, 1),
'leftpos' : hp.quniform('leftpos', 0, 10000, 500),
'rightpos' : hp.quniform('rightpos', 10000, 20000, 500),
'activationFxn' : 'relu',
'numFiltersConv1' : 2**hp.quniform('numFiltersConv1', 4, 7, 1),
'filterLenConv1' : hp.quniform('filterLenConv1', 1, 10, 1),
'dilRate1' : hp.quniform('dilRate1', 1, 4, 1),
'maxPool1' : hp.quniform('maxPool1', 5, 100, 5), #5, 100, 5),
'numconvlayers' : hp.choice('numconvlayers', [
{
'numconvlayers1' : 'one'
},
{
'numFiltersConv2' : 2**hp.quniform('numFiltersConv2', 4, 7, 1),
'filterLenConv2' : hp.quniform('filterLenConv2', 1, 10, 1),
'dilRate2' : hp.quniform('dilRate2', 1, 4, 1),
'maxPool2' : hp.quniform('maxPool2', 5, 100, 5),
'numconvlayers1' : hp.choice('numconvlayers1', [
{
'numconvlayers2' : 'two'
},
{
'numFiltersConv3' : 2**hp.quniform('numFiltersConv3', 4, 7, 1),
'filterLenConv3' : hp.quniform('filterLenConv3', 1, 10, 1),
'dilRate3' : hp.quniform('dilRate3', 1, 4, 1),
'maxPool3' : hp.quniform('maxPool3', 5, 100, 5),
'numconvlayers2' : hp.choice('numconvlayers2', [
{
'numconvlayers3' : 'three'
},
{
'numFiltersConv4' : 2**hp.quniform('numFiltersConv4', 4, 7, 1),
'filterLenConv4' : hp.quniform('filterLenConv4', 1, 10, 1),
'dilRate4' : hp.quniform('dilRate4', 1, 4, 1),
'maxPool4' : hp.quniform('maxPool4', 5, 100, 5),
'numconvlayers3' : 'four'
}])
}])
}]),
'dense1' : 2**hp.quniform('dense1', 1, 8, 1),
'dropout1' : hp.uniform('dropout1', 0, 1),
'numdenselayers' : hp.choice('numdenselayers', [
{
'layers' : 'one'
},
{
'layers' : 'two' ,
'dense2' : 2**hp.quniform('dense2', 1, 8, 1),
'dropout2' : hp.uniform('dropout2', 0, 1)
}
])
}
def objective(params):
global best_file
leftpos = int(params['leftpos'])
rightpos = int(params['rightpos'])
activationFxn = params['activationFxn']
global X_trainhalflife, y_train
X_trainpromoterSubseq = X_trainpromoter[:,leftpos:rightpos,:]
X_validpromoterSubseq = X_validpromoter[:,leftpos:rightpos,:]
halflifedata = Input(shape=(X_trainhalflife.shape[1:]), name='halflife')
input_promoter = Input(shape=X_trainpromoterSubseq.shape[1:], name='promoter')
mse = 1
# defined architecture with best hyperparameters
x = Conv1D(int(params['numFiltersConv1']), int(params['filterLenConv1']), dilation_rate=int(params['dilRate1']), padding='same', kernel_initializer='glorot_normal', input_shape=X_trainpromoterSubseq.shape[1:],activation=activationFxn)(input_promoter)
x = MaxPooling1D(int(params['maxPool1']))(x)
if params['numconvlayers']['numconvlayers1'] != 'one':
maxPool2 = int(params['numconvlayers']['maxPool2'])
x = Conv1D(int(params['numconvlayers']['numFiltersConv2']), int(params['numconvlayers']['filterLenConv2']), dilation_rate=int(params['numconvlayers']['dilRate2']), padding='same', kernel_initializer='glorot_normal',activation=activationFxn)(x) #[2, 3, 4, 5, 6, 7, 8, 9, 10]
x = MaxPooling1D(maxPool2)(x)
if params['numconvlayers']['numconvlayers1']['numconvlayers2'] != 'two':
maxPool3 = int(params['numconvlayers']['numconvlayers1']['maxPool3'])
x = Conv1D(int(params['numconvlayers']['numconvlayers1']['numFiltersConv3']), int(params['numconvlayers']['numconvlayers1']['filterLenConv3']), dilation_rate=int(params['numconvlayers']['numconvlayers1']['dilRate3']), padding='same', kernel_initializer='glorot_normal',activation=activationFxn)(x) #[2, 3, 4, 5]
x = MaxPooling1D(maxPool3)(x)
if params['numconvlayers']['numconvlayers1']['numconvlayers2']['numconvlayers3'] != 'three':
maxPool4 = int(params['numconvlayers']['numconvlayers1']['numconvlayers2']['maxPool4'])
x = Conv1D(int(params['numconvlayers']['numconvlayers1']['numconvlayers2']['numFiltersConv4']), int(params['numconvlayers']['numconvlayers1']['numconvlayers2']['filterLenConv4']), dilation_rate=int(params['numconvlayers']['numconvlayers1']['numconvlayers2']['dilRate4']), padding='same', kernel_initializer='glorot_normal',activation=activationFxn)(x) #[2, 3, 4, 5]
x = MaxPooling1D(maxPool4)(x)
x = Flatten()(x)
x = Concatenate()([x, halflifedata])
x = Dense(int(params['dense1']))(x)
x = Activation(activationFxn)(x)
x = Dropout(params['dropout1'])(x)
if params['numdenselayers']['layers'] == 'two':
x = Dense(int(params['numdenselayers']['dense2']))(x)
x = Activation(activationFxn)(x)
x = Dropout(params['numdenselayers']['dropout2'])(x)
main_output = Dense(1)(x)
model = Model(inputs=[input_promoter, halflifedata], outputs=[main_output])
model.compile(SGD(lr=0.0005, momentum=0.9),'mean_squared_error', metrics=['mean_squared_error'])
# model.compile(Adam(lr=0.0005, beta_1=0.9, beta_2=0.90, epsilon=1e-08, decay=0.0),'mean_squared_error', metrics=['mean_squared_error'])
print(model.summary())
modelfile = os.path.join(params['datadir'], 'plotted_model.png')
plot_model(model, show_shapes=True, show_layer_names=True, to_file=modelfile)
#train model on training set and eval on 1K validation set
check_cb = ModelCheckpoint(os.path.join(params['datadir'], 'bestparams.h5'), monitor='val_loss', verbose=1, save_best_only=True, mode='min')
earlystop_cb = EarlyStopping(monitor='val_loss', patience=7, verbose=1, mode='min')
result = model.fit([X_trainpromoterSubseq, X_trainhalflife], y_train, batch_size=int(params['batchsize']), shuffle="batch", epochs=100,
validation_data=[[X_validpromoterSubseq, X_validhalflife], y_valid], callbacks=[earlystop_cb, check_cb])
mse_history = result.history['val_mean_squared_error']
mse = min(mse_history)
#evaluate performance on test set using best learned model
best_file = os.path.join(params['datadir'], 'bestparams.h5')
model = load_model(best_file)
print('Loaded results from:', best_file)
X_testpromoterSubseq = X_testpromoter[:,leftpos:rightpos,:]
predictions_test = model.predict([X_testpromoterSubseq, X_testhalflife], batch_size=64).flatten()
slope, intercept, r_value, p_value, std_err = stats.linregress(predictions_test, y_test)
print('Test R^2 = %.3f' % r_value**2)
df = pd.DataFrame(np.column_stack((geneName_test, predictions_test, y_test)), columns=['Gene','Pred','Actual'])
print('Rows & Cols:', df.shape)
df.to_csv(os.path.join(params['datadir'], 'predictions.txt'), index=False, header=True, sep='\t')
return {'loss': mse, 'status': STATUS_OK }
datadir="pM10Kb_1KTest"
main(datadir=datadir)
#Matches FigS2A
Image(retina=True, filename=os.path.join(datadir, 'plotted_model.png'))
# + [markdown] id="4F0TJSN1NnZ8" colab_type="text"
# # Generate predictions on a tiled genomic locus or other group of DNA sequences
# + id="2cfmd9lfOMg8" colab_type="code" outputId="30cff592-0a3f-49d0-d284-daa6f2ab1261" colab={"base_uri": "https://localhost:8080/", "height": 353}
def one_hot(seq):
num_seqs = len(seq)
seq_len = len(seq[0])
seqindex = {'A':0, 'C':1, 'G':2, 'T':3, 'a':0, 'c':1, 'g':2, 't':3}
seq_vec = np.zeros((num_seqs,seq_len,4), dtype='bool')
for i in range(num_seqs):
thisseq = seq[i]
for j in range(seq_len):
try:
seq_vec[i,j,seqindex[thisseq[j]]] = 1
except:
pass
return seq_vec
def generate_predictions(model_file, input_file, output_file):
model = load_model(best_file) #or use one of several pre-trained models
encoding = guess_type(input_file)[1] # uses file extension to guess zipped or unzipped
if encoding is None:
_open = open
elif encoding == 'gzip':
_open = partial(gzip.open, mode='rt')
else:
raise ValueError('Unknown file encoding: "{}"'.format(encoding))
i, bs, names, predictions, sequences = 0, 32, [], [], []
hlfeatures=8
halflifedata = np.zeros((bs,hlfeatures), dtype='float32')
with _open(input_file) as f:
for fasta in SeqIO.parse(f, 'fasta'):
name, sequence = fasta.id, str(fasta.seq)
sequences.append(sequence)
names.append(name)
i += 1
if (len(sequence) != 10500):
sys.exit( "Error in sequence %s, length is not equal to the required 10,500 nts. Please fix or pad with Ns if necessary." % name )
if i % bs == 0:
seq = one_hot(sequences)
predictions.extend( model.predict([seq, halflifedata], batch_size=bs).flatten().tolist() )
sequences = []
remain = i % bs
if remain > 0:
halflifedata = np.zeros((remain,hlfeatures), dtype='float32')
seq = one_hot(sequences)
predictions.extend( model.predict([seq, halflifedata], batch_size=remain).flatten().tolist() )
df = pd.DataFrame(np.column_stack((names, predictions)), columns=['ID','SCORE'])
print(df[1:10]) #print first 10 entries
df.to_csv(output_file, index=False, header=True, sep='\t')
generate_predictions(model_file="pM10Kb_1KTest/bestparams.h5",
input_file="input_fasta/testinput.fa.gz",
output_file="test_predictions.txt")
generate_predictions(model_file="pM10Kb_1KTest/bestparams.h5",
input_file="input_fasta/human_promoters.fa.gz",
output_file="human_promoter_predictions.txt")
# + [markdown] id="c1X21GN57TY_" colab_type="text"
# # Train Xpresso model for mouse data and predict promoters
#
# + id="VeAQTtrQ7pVJ" colab_type="code" outputId="4d06d48f-323c-432b-c300-9b541d14d835" colab={"base_uri": "https://localhost:8080/", "height": 1000}
main(datadir="pM10Kb_1KTest_Mouse")
generate_predictions(model_file="pM10Kb_1KTest_Mouse/bestparams.h5",
input_file="input_fasta/mouse_promoters.fa.gz",
output_file="mouse_promoter_predictions.txt")
| Xpresso.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/CindyMG/Core-II-Wk-4/blob/main/Core_II_W4_KNN.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="YoC9kcuG4TPz"
# #Importing libraries
# + id="ZSdAbinF3m5Y"
#importing our libraries
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# + [markdown] id="tQJucjb_Ff-4"
# #Loading our dataset
#
#
# + id="F3L29JgJ7FtE"
#for this project, we will be using the Titanic dataset: http://bit.ly/TitanicTrainDataset
#The columns and their descriptions are as shown below:
# Survived - Survival (0 = No; 1 = Yes)
# Pclass - Passenger Class (1 = 1st; 2 = 2nd; 3 = 3rd)
# Name - Name
# Sex - Sex
# Age - Age
# SibSp - Number of Siblings/Spouses Aboard
# Parch - Number of Parents/Children Aboard
# Ticket - Ticket Number
# Fare - Passenger Fare (British pound)
# Cabin - Cabin
# Embarked - Port of Embarkation (C = Cherbourg; Q = Queenstown; S = Southampton)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="9UeRLICahjvR" outputId="37e183c8-3462-457a-ee09-49e73ba28fce"
#let us load the dataset
train = pd.read_csv('/content/train.csv')
#preview the first five records
train.head()
# + colab={"base_uri": "https://localhost:8080/"} id="Pien6uD3Bhcu" outputId="44c943fe-4389-4c30-824f-049cd35a5347"
#checking the number or rows and columns in our dataset
train.shape
# 891 rows and 12 columns!
# + colab={"base_uri": "https://localhost:8080/"} id="XqpIxXDYAD7a" outputId="5d9159c0-c207-43f5-db55-62ddd27c283c"
#let us get some more details concerning our dataset - shape, data types etc.
train.info()
# + [markdown] id="PNYLxVC4D6ZF"
# #DATA CLEANING & EXPLORATORY DATA ANALYSIS
# + [markdown] id="1uLVxY1QQP3Q"
# ##Removing null values
# + colab={"base_uri": "https://localhost:8080/"} id="PWK3gkL-BkjZ" outputId="0255e1e9-1331-4fc9-eaf5-c41929f54c7b"
#finding and removing null or missing values
#finding out the sum of null values in each column
train.isnull().sum()
#all the columns have no missing values except 'Age' and 'Cabin'
# 'Cabin' has way too many null values and it will not be very relevant for our analysis so we can just drop the whole column
# The 'Age' column however, seems to be an important variable so we will have to find a different approach to get rid of the null values
# 'Embarked' has only 2 null values so when the time comes to drop them,
#we can comfortably drop them without worrying about compromising the accuracy of our dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="g0ZYt90GHmhX" outputId="ba4b595a-e044-4741-d0b7-c67d7b411473"
#as we are dropping the 'cabin' column, we might as well drop other columns that may not be of much help to our study
titanic = train.drop(['PassengerId','Name','Ticket','Cabin'], 1)
titanic.head()
# + colab={"base_uri": "https://localhost:8080/"} id="PjLt9g3uLkpC" outputId="502181c7-551e-430e-af9d-e5934efd77ff"
#checking and dropping any duplicates in our dataset
titanic.drop_duplicates(subset=None, keep='first', inplace=True)
#let us check how many records we have left after removing duplicates
titanic.shape
#only a slight difference in rows, we can move on!
# + [markdown] id="c92GhaVuVy3w"
# ##Collinearity
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="ewQ6Fn0dU-Fy" outputId="24672131-1738-4d4f-b0fc-33e6f9bf6cbf"
#checking the correlation between the variables
sns.heatmap(titanic.corr())
# we can see how some variables have strong correlations - such as 'Fare' and 'Pclass' (the higher the class, the more the fare)
# others are 'Age' and 'Pclass' , 'Survived' and 'Pclass' etc.
# + [markdown] id="kq4Boe6vQII0"
# ##Bivariate analysis/ data cleaning
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="oyrBuymZM6SH" outputId="45ab917d-009f-474d-8d16-52d6d8eabafb"
# now, we want to handle the null values in the 'Age' column
# dropping the entire column is unwise because 'Age' seems like an important variable and dropping 177 null values is completely out of question!
# so, how do we go about this?
# we are going to use the variable 'Pclass' to help us out a little bit
# Let's look at how passenger age is related to their class as a passenger on the boat.
#
sns.boxplot(x='Pclass', y='Age', data=titanic, palette='hls')
# although it is a weak correlation, we can see that Pclass and Age are negatively correlated;
# younger passengers appear to be in the 3rd class while older passengers seem to be in the former classes
# let us try checking the average ages of all 3 classes: 1st class - 38, 2nd class - 30, 3rd class - 25
# + id="KNnMNnpvJtRl"
# Write a function that finds each null value in the 'Age' variable,
#For each null value, it checks the value of the Pclass and assigns an age value according to the average age of passengers in that class.
#
def age_approx(cols):
Age = cols[0]
Pclass = cols[1]
if pd.isnull(Age):
if Pclass == 1:
return 38
elif Pclass == 2:
return 30
else:
return 25
else:
return Age
# + [markdown] id="G_nc8WVhQUzf"
# ##Removing the remaining null values
# + colab={"base_uri": "https://localhost:8080/"} id="GYwHpbOYPD0p" outputId="d8a748be-5edc-4a92-bb62-30bdbc89503d"
# Call the function.
#
titanic['Age'] = titanic[['Age', 'Pclass']].apply(age_approx, axis=1)
# now, check whether all the null values in 'Age' have been replaced
titanic.isnull().sum()
#Great! Now let us drop those 2 null values in 'Embarked'
titanic.dropna(inplace=True)
titanic.isnull().sum()
#No null values left!
# + [markdown] id="cSlcsnQyQdsf"
# ##Univariate analysis
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="skVerU6hQpDk" outputId="38095932-4d20-45a4-af94-15ac81d4c9cd"
# let us pick one variable to analyze, 'Survived'
# we are going to use a countplot to compare the number of those who survived(1) and those who perished(0)
sns.countplot(x='Survived',data=titanic, palette='hls')
# clearly from the visualization above, we can see that the number of passengers who perished is higher than those who survived
# + [markdown] id="v-iZDozYSHwG"
# ##Encoding categorical variables using dummies
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="XblFoIIbRZnu" outputId="bd25ed5f-c213-465e-884e-09267b7ca570"
# We need to convert the 'Sex' and 'Embarked' variables into numerical variables.
# =
sex = pd.get_dummies(titanic['Sex'],drop_first=True)
sex.head()
# male - 1, female - 0
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="y6Dehv97ShUW" outputId="1475e9a0-a0f3-4974-f22f-748a2cbca21d"
# just for a bit of context, Embarked - Port of Embarkation (C = Cherbourg; Q = Queenstown; S = Southampton)
embarked = pd.get_dummies(titanic['Embarked'],drop_first=True)
embarked.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="PSom_8mvTtdf" outputId="83c51831-e34e-4731-a357-d606a055eeea"
#let us review the original dataset
titanic.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="zu3tpd4STrQH" outputId="ebbef361-f74c-4dbc-a257-acec13884a1a"
# now, we will drop the 'sex' and 'embarked' columns and concatenate the new columns we made after creating the dummies
titanic.drop(['Sex', 'Embarked'],axis=1,inplace=True)
titanic.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="GWUCTRqOUYbk" outputId="41bc920a-c3d7-4d81-de42-d5f120b666f0"
# assign a new variable for the dataset after concatenation
titanic1 = pd.concat([titanic,sex,embarked],axis=1)
titanic1.head()
# Now the categorical data variables have been converted into numerical format!
# + [markdown] id="Qhrcq668WuOc"
# #K-Nearest Neighbour Classifier
# + [markdown] id="fQxIZNIEa--N"
# ## Using an 80% - 20% ratio
# + id="4xOJ8smWXlqB"
#let us define our X and y variables
X = titanic1.drop("Survived",axis=1)
y = titanic1["Survived"]
#import the train test split libraries
from sklearn.model_selection import train_test_split
#split the data into 80% training and 20% test data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20)
# + id="bgE9hK9_Xxk_"
# Feature Scaling
#
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="lRY-9USsYHnU" outputId="4940c32e-65be-4442-eb92-f9cdae55b345"
# Training and predictions
#
# import the KNN classifier libraries
from sklearn.neighbors import KNeighborsClassifier
# now, we will choose n_neighbours, or basically the value of K - 5 is the most commonly used one
classifier = KNeighborsClassifier(n_neighbors=5)
classifier.fit(X_train, y_train)
# + id="tmfg5ICTYQFS"
# The final step is to make predictions on our test data
y_pred = classifier.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="b6IzDOmgYWis" outputId="849e5ead-eca8-47a6-afb1-76b404171275"
# Evaluation
# For evaluating an algorithm, confusion matrix, precision, recall and f1 score are the most commonly used metrics.
#importing the confusion matrix libraries
from sklearn.metrics import classification_report, confusion_matrix
#displaying the confusion matrix and classification report
print(confusion_matrix(y_test, y_pred))
print(classification_report(y_test, y_pred))
# + [markdown] id="wSPWEcfbYoqd"
# **Confusion matrix:** Our ***true negative***(number of people who died in the crash) is 78; only seventy eight perished cases out of the total were correctly predicted.
#
# Our ***true positive***(number of survivors) is 46; only 46 survived cases out of the total were correctly predicted.
#
# We can see that the precision score for 0(perished) is 78% while that of 1(survived) is 80%. The general ***accuracy score*** of this model is **79%** - not bad, but it shows that this model is not that good.
# + [markdown] id="WWECmTw1bJSs"
# ## Using a 70% - 30% ratio
# + id="62uDl05qaV7J"
# we had initially split the data into 80% training and 20% test data
# what if we change the ratio?
# let us try creating a new model but now splitting the data into 70% training data and 30% test data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30)
# + id="jbPV3FTIaxUC"
# Feature Scaling
#
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="zKBgNQb9a0qV" outputId="e978ae79-8595-4990-e4d3-4438f866aca4"
# Training and predictions
#
# import the KNN classifier libraries
from sklearn.neighbors import KNeighborsClassifier
# now, we will choose n_neighbours, or basically the value of K - 5 is the most commonly used one
classifier = KNeighborsClassifier(n_neighbors=5)
classifier.fit(X_train, y_train)
# + id="xFnO5zUpa3gX"
# The final step is to make predictions on our test data
y_pred = classifier.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="x7HVBF1Ia9dY" outputId="3e0c81fa-247a-4f8c-b20f-a7489268c94b"
# Evaluation
# For evaluating an algorithm, confusion matrix, precision, recall and f1 score are the most commonly used metrics.
#importing the confusion matrix libraries
from sklearn.metrics import classification_report, confusion_matrix
#displaying the confusion matrix and classification report
print(confusion_matrix(y_test, y_pred))
print(classification_report(y_test, y_pred))
# + [markdown] id="NdZJ1WJdbSpe"
# **Confusion matrix:** Our ***true negative***(number of people who died in the crash) is 107; one hundred and seven perished cases out of the total were correctly predicted.
#
# Our ***true positive***(number of survivors) is 76; only 76 survived cases out of the total were correctly predicted.
#
# We can see that the precision score for 0(perished) is 77% - a slight decrease from the previous model; while that of 1(survived) is 80% - a slight decrease from the previous model. The general ***accuracy score*** of this model is **78%** - a slightly lower score than the previous model.
#
# Not much has changed, right? Let us try a different approach.
# + [markdown] id="NUxRHFLtb-xp"
# ## Changing the n-neighbours
# + colab={"base_uri": "https://localhost:8080/"} id="Q7oYMScbcE4I" outputId="a377deda-597a-48e3-89f1-66dd2402e920"
# now, we will choose n_neighbours, or basically the value of K - 3
classifier = KNeighborsClassifier(n_neighbors=3)
classifier.fit(X_train, y_train)
# + id="10_7DLP4axmo"
# The final step is to make predictions on our test data
y_pred = classifier.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="DObh3UNma2QT" outputId="77018405-f7d1-4531-cc62-b15065a08cbd"
# Evaluation
# For evaluating an algorithm, confusion matrix, precision, recall and f1 score are the most commonly used metrics.
#importing the confusion matrix libraries
from sklearn.metrics import classification_report, confusion_matrix
#displaying the confusion matrix and classification report
print(confusion_matrix(y_test, y_pred))
print(classification_report(y_test, y_pred))
# + [markdown] id="mDRshTAna583"
# **Confusion matrix:** Our ***true negative***(number of people who died in the crash) is 106; one hundred and six perished cases out of the total were correctly predicted.
#
# Our ***true positive***(number of survivors) is 73; only 73 survived cases out of the total were correctly predicted.
#
# We can see that the precision score for 0(perished) is 75% - a slight decrease from the previous models; while that of 1(survived) is 78% - similar to the previous model. The general ***accuracy score*** of this model is **76%** - a slightly lower score than the previous models.
#
# Changing the n-neighbours from 5 to 3, seems to have lowered
| Core_II_W4_KNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
#
#
#
# # Simulateur SVOM
# Dans le cadre de la mission SVOM (Space Variable Objects Monitor) de détection de sursauts gamma plusieurs détecteurs, caméras et téléscopes ont été choisis pour des observations sur terre. Grâce à ce simulateur et ce tutoriel vous avez accès à des simulations de ce que peuvent nous donner ces différents instruments lors de phases de test (champ sombre, champ plat, ...) ou lors d'observations (données collectées grâce à différentes mesures faites sur les téléscopes).
# ## Sommaire :
# 1. Fonctionnement
# 1. Fichiers de configuration
# 2. Explication du code principal
# 2. Simulation d'une image
# 3. Simulation d'une rampe
# 1. Simulation
# 2. Test des différents effets simulés
# 4. Simulation d'un bruit CDS
# A réaliser au départ absolument :
# %reload_ext autoreload
# %autoreload 2
# %matplotlib notebook
import os
import numpy as np
import matplotlib.pyplot as plt
from ImSimpy.ImSimpy import ImageSimulator
# # 1. Fonctionnement
# ### A. Fichiers de configuration
# Dans le dossier *ImSimpy/ImSimpy/configFiles* et *pyETC/pyETC/telescope_database* se trouve des fichiers *.hjson* qui sont des fichiers de configuration. Vous pourrez par la suite choisir de simuler une rampe ou un bruit CDS (cf. 3ème partie ou 4ème partie respectivement).
# Dans le dossier *ImSimpy/ImSimpy/configFiles* vous pouvez choisir de simuler des champs sombre (*Dark.hjson*), des champs plat (*Flat.hjson*), des images réalisées sur un temps très court pour mesurer le bruit de lecture (Texp=0s, *Bias.hjson*). Vous pouvez aussi décider de créer votre propre fichier .hjson en remplissant vous même le fichier *Make_one_image_is.hjson* (is pour image simulator). Dans ce fichier vous pouvez choisir le temps d'exposition, des paramètres de calcul, choisir des conditions locale comme la température ou l'élévation de la lune et des paramètres du téléscope comme le seeing ou des paramètres de caméras. Mais c'est surtout à l'intérieur de ce fichier que vous pouvez choisir d'observer le ciel et l'endroit d'observation.
#
# Le fichier pris par défault est un Dark : [Dark.hjson](../ImSimpy/configFiles/Dark.hjson)
#
# *Remarque :* C'est à l'intérieur de ces fichiers que l'on choisit l'emplacement des fichiers .fits de configuration : carte des facteurs de conversion, carte des coefficients dépendant du temps du bruit thermique que produit les instruments devant le détecteur, carte des pixels morts et chauds dans le cas d'une simulation d'image simple et des facteurs de non linéarité pour une rampe. Un fichier fits pour le vignettage et pour l'offset.
# Dans le dossier *pyETC/pyETC/telescope_database* vous pouvez choisir le téléscope (associé à sa/ses caméra/s). Par exemple, pour simuler des images du détecteur Teledyne il faut choisir *colibri_teledyne.hjson* et pour simuler des images faites par la caméra de Sofradir choisissez *colibri.hjson*.
#
# Le téléscope choisi par défault est : [colibri_teledyne.hjson](../../pyETC/pyETC/telescope_database/colibri_teledyne.hjson)
#
# *Remarque :* C'est à l'intérieur de ces fichiers que l'on doit placer certains coefficients importants comme la taille des pixels correspondant au détecteur, la largeur des bandes de pixels de référence ou les neufs coefficients de diaphonie interpixel.
# Dans la cellule ci-dessous vous pouvez donc choisir votre type de mesure et votre téléscope en remplaçant `Dark.hjson` et `'colibri_teledyne'` par ce que vous souhaitez après avoir mis les coefficients correspondant à votre simulation.
Simulation=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'/ImSimpy/configFiles/Test_des_differents_effets/01.Sources.hjson',
name_telescope='colibri_teledyne')
# L'objet est stocké dans la variable "Simulation". Cela peut être n'importe quoi d'autre.
#
# Si vous n'avez pas le répertoire standard, vous devez spécifier le chemin où se trouve le fichier de configuration. Voir l'exemple dans la partie 2.
# ### B. Explication du code principal
# *Remarque sur le fonctionnement du code* : Comme dit plus haut, c'est dans les fichiers de configuration que l'on choisit quel type d'image on veut simuler. C'est dans ces fichiers que l'on choisit la partie du ciel à oberserver, mais c'est aussi à l'aide de ces fichiers que l'on peut choisir de voir qu'une partie des instruments. Il suffit de mettre à ce moment là la bonne carte en entrée du bruit intrinsèque de l'instrument et d'afficher ou non les sources, les rayons cosmique et autres signal externe ou interne au détecteur ou au télescope.
#
# Le **coeur du code** se trouve dans le programme *ImSimpy* (qui se trouve dans *ImSimpy\ImSimpy\ImSimpy.py*) car les différents objets appelés (`Simulation`, `Test`, `AcquisitionBruitCDS`) font parti de la classe `ImageSimulator` définie au début de ce programme.
#
# Le code est divisé en trois parties :
# * La première contient des **fonctions de configurations** (dans l’ordre : l’initialisation, la recherche des paramètres dans les fichiers de configuration, la création des en-têtes fits puis la lecture et la génération des sources et des PSF (étalement)). Cette partie s’étend donc de la fonction `__init__` à la fonction `generateFinemaps`).
# * La deuxième contient toutes les fonctions qui **simulent** les différents effets (ajout des sources, non linéarité, discrétisation, …).
# * La dernière partie est constitué de deux fonctions, Simulate et Rampsimulation. Comme leur nom l’indique c’est elles qui appellent les différentes fonctions de configuration puis les différentes fonctions produisant les effets dans l’ordre convenu.
#
#
# # 2. Simulation d'une image
# L'image est enregistrée dans le dossier *ImSimpy/images* et vous devez placez le nom ci-dessous :
# +
#File name
Simulation.nom = 'test'
#Nom du dossier dans lequel il sera enregistrée (le dossier doit être déjà existant)
Simulation.output_dir= 'Simulation d_une image'
#Read the configfile
Simulation.acquisition='simple'
Simulation.readConfigs()
#execute it:
Simulation.simulate()
# -
# *NB: Parmi les fichiers fits que vous appliquez à l'image l'offset, le gain de conversion et la cosmétique sont appliqués aux pixels de références, tandis que le bruit thermique de l'instrument et le vignetting ne leur sont pas appliqués.*
#
# Pour récupérer le nom de l'image simulée :
fits_filename=os.getenv('ImSimpy_DIR')+'/ImSimpy/images/'+Simulation.information['output']
print (fits_filename)
# # 3. Simulation d'une rampe
# ### A. Simulation
# Pour simuler une rampe choisissez dans un premier temps un type de mesure et un téléscope (cf 1ère partie). Remplacez ci-dessous dans la variable `Simulation` le type et le téléscope choisis.
#
# Ensuite, dans la variable `bands` placez un tableau avec les bandes spectrales que vous souhaitez du système photométrique. Voici un tableau comportant toutes les bandes (par défault seule la bande J est choisie) :
#
# bands= ['g','r','i','z','y','J','H']
# Dans la variable `output_dir` placez le nom du dossier dans lequel la rampe sera enregistrée dans le dossier *images* de ImSimpy (vous devez créer le dossier auparavant).
#
# 
#
# Choisissez enfin les paramètres à l'aide du schéma ci-dessus et placez les dans les variables suivantes :
# * nombre de reset : `nbReset`
# * nombre de read : `nbRead`
# * nombre de drop : `nbDrop`
# * nombre de groupes : `nbGroup`
# *NB : La fonction diginoise est supprimée, son effet est déjà simulé grâce à la fonction * `intscale` * qui discretise, elle convertit les entiers en flotants. Le shotnoise non plus, la statistique est ajoutée au fur et à mesure sur chaque fonction. Et pour le moment les rayons cosmique et la persistance ne sont pas simulés car ils sont encore en recherche (il manque des mesures faites à l'aide de téléscope pour simuler les rayons cosmique).*
# +
from ImSimpy.ImSimpy import ImageSimulator
Simulation=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'/ImSimpy/configFiles/Dark.hjson',
name_telescope='colibri_teledyne')
Simulation.bands = ['J']
Simulation.output_dir='H2RGSim'
Simulation.nbReset=5
Simulation.nbRead=1
Simulation.nbDrop=9
Simulation.nbGroup=20
Simulation.acquisition='ramp'
Simulation.readConfigs()
Simulation.config['verbose'] = 'False'
for band in Simulation.bands:
# Select appropriate channel for the selected filter band
if band in ['g','r','i']:
Simulation.config['channel']='DDRAGO-B'
elif band in ['z','y']:
Simulation.config['channel']='DDRAGO-R'
elif band in['J','H']:
Simulation.config['channel']='CAGIRE'
# Set filter band
Simulation.config['filter_band']=band
# Simulation
Simulation.Rampsimulation('data')
# -
# ### B. Test des différents effets
# Si vous souhaitez tester le fonctionnement d'un des effets appliqué, le code ci-dessous créé ces images dans le dossier *ImSimpy\ImSimpy\images\Test_des_différents_effets* :
# * Image 1 : Seulement les sources
# * Image 2 : Seulement le fond du ciel
# * Image 3 : Sources + fond du ciel
# * Image 4 : Seulement les rayons cosmiques
# * Image 5 : Sources + fond du ciel + rayons cosmiques
# * etc...
#
# L'offset n'est pas pris en compte car il empêche une bonne visibilité des effets. Le diginoise non plus car il n'a pas sa place ici, son effet est déjà simulé grâce à la fonction `intscale` qui converti les entiers en flotants.
#
# *NB : Pour le moment les rayons cosmique et la persistance ne sont pas simulés car c'est encore en recherche !*
# +
# Première image, seulement les sources :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\01.Sources.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '01.Sources'
Simulation.acquisition='ramp'
Test.readConfigs()
Test.simulate()
# Seulement le fond du ciel:
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\02.Sky Background.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '02.Sky Background'
Test.readConfigs()
Test.simulate()
# Sources et fond du ciel :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\03.Sources+SkyB.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '03.Sources+SkyB'
Test.readConfigs()
Test.simulate()
# Seulement les rayons cosmiques :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\04.CosmicRay.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '04.CosmicRay'
Test.readConfigs()
Test.simulate()
# Sources+SkyB+CosmicR :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\05.Sources+SkyB+CosmicR.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '05.Sources+SkyB+CosmicR'
Test.readConfigs()
Test.simulate()
# Seulement la persistance :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\06.Persistance.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '06.Persistance'
Test.readConfigs()
Test.simulate()
# Sources+SkyB+CosmicR+Persis :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\07.Sources+SkyB+CosmicR+Persis.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '07.Sources+SkyB+CosmicR+Persis'
Test.readConfigs()
Test.simulate()
# Seulement le vignetting :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\08.Vignetting.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '08.Vignetting'
Test.readConfigs()
Test.simulate()
# Sources+SkyB+CosmicR+Persis+Vignet :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\09.Sources+SkyB+CosmicR+Persis+Vignet.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '09.Sources+SkyB+CosmicR+Persis+Vignet'
Test.readConfigs()
Test.simulate()
# Seulement le bruit thermique de l'instrument :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\10.Intrument intrinsic noise.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '10.Intrument intrinsic noise'
Test.readConfigs()
Test.simulate()
# Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\11.Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '11.Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise'
Test.readConfigs()
Test.simulate()
# Seulement le dark current :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\12.Dark Current.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '12.Dark Current'
Test.readConfigs()
Test.simulate()
# Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\13.Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '13.Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC'
Test.readConfigs()
Test.simulate()
# Seulement la diaphonie interpixel :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\14.Interpixel CrossTalk.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '14.Interpixel CrossTalk'
Test.readConfigs()
Test.simulate()
# Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC+ICT :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\15.Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC+ICT.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '15.Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC+ICT'
Test.readConfigs()
Test.simulate()
# Seulement la statistique, le shot noise :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\16.Shot Noise.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '16.Shot Noise'
Test.readConfigs()
Test.simulate()
# Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC+ICT+ShotNoise :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\17.Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC+ICT+ShotNoise.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '17.Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC+ICT+ShotNoise'
Test.readConfigs()
Test.simulate()
# Seulement la cosmétique :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\18.Cosmetique.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '18.Cosmetique'
Test.readConfigs()
Test.simulate()
# Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC+ICT+ShotNoise+Cosmetic :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\19.Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC+ICT+ShotNoise+Cosmetic.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '19.Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC+ICT+ShotNoise+Cosmetic'
Test.readConfigs()
Test.simulate()
# Seulement le bleeding :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\20.Bleeding.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '20.Bleeding'
Test.readConfigs()
Test.simulate()
# Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC+ICT+ShotNoise+Cosmetic+Bleeding :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\21.Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC+ICT+ShotNoise+Cosmetic+Bleeding.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '21.Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC+ICT+ShotNoise+Cosmetic+Bleeding'
Test.readConfigs()
Test.simulate()
# Seulement le bruit de lecture :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\22.ReadOutNoise.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '22.ReadOutNoise'
Test.readConfigs()
Test.simulate()
# Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC+ICT+ShotNoise+Cosmetic+Bleeding+RON :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\23.Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC+ICT+ShotNoise+Cosmetic+Bleeding+RON.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '23.Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC+ICT+ShotNoise+Cosmetic+Bleeding+RON'
Test.readConfigs()
Test.simulate()
# Seulement le gain de conversion :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\24.Convert.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '24.Convert'
Test.readConfigs()
Test.simulate()
# Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC+ICT+ShotNoise+Cosmetic+Bleeding+RON+Convert :
Test=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'\\ImSimpy\\configFiles\\Test_des_differents_effets\\25.Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC+ICT+ShotNoise+Cosmetic+Bleeding+RON+Convert.hjson',
name_telescope='colibri_teledyne')
Test.output_dir='Test_des_differents_effets'
Test.nom = '25.Sources+SkyB+CosmicR+Persis+Vignet+InstrumNoise+DC+ICT+ShotNoise+Cosmetic+Bleeding+RON+Convert'
Test.readConfigs()
Test.simulate()
# -
# # 4. Bruit CDS
# L'acquisition de bruit CDS (Correlated double sampling ou échantillonnage double corrélé) est un mode où on peut mettre en avant le bruit. Comme dans la figure ci-dessous, quatre images sont acquises suivant deux rampes et une dernière image est créée où seulement le bruit apparaît.
#
# 
# Pour simuler cette dernière on soustrait deux images consécutives pour en trouver l’écart-type. Or, avec des CMOS cet écart-type n’est pas représentatif de l’erreur, le gain et l’offset varient par pixel. C’est pourquoi, on compare 4 images, on calcul le l’écart-type deux à deux. Un coefficient de racine de 2 apparaît :
#
# 
# 
#
#
# Cette dernière image subira aussi des traitements des pixels de références expliqués plus loins.
#
#
# De la même manière que dans la partie trois, choisissez le type de téléscope, le type d'image à simuler, la fenêtre atmosphérique et le dossier dans lequel votre acquisition sera enregistrée.
# +
from ImSimpy.ImSimpy import ImageSimulator
AcquisitionBruitCDS=ImageSimulator(configFile=os.getenv('ImSimpy_DIR')+'/ImSimpy/configFiles/Dark.hjson',
name_telescope='colibri_teledyne')
AcquisitionBruitCDS.bands = ['J']
AcquisitionBruitCDS.output_dir='AcquisitionBruitCDS'
# Acquisition CDS (ne plus rien modifier) :
AcquisitionBruitCDS.nbReset=5
AcquisitionBruitCDS.nbRead=2
AcquisitionBruitCDS.nbDrop=0
AcquisitionBruitCDS.nbGroup=1
AcquisitionBruitCDS.acquisition='CDS'
AcquisitionBruitCDS.readConfigs()
AcquisitionBruitCDS.config['verbose'] = 'False'
for band in AcquisitionBruitCDS.bands:
# Select appropriate channel for the selected filter band
if band in ['g','r','i']:
AcquisitionBruitCDS.config['channel']='DDRAGO-B'
elif band in ['z','y']:
AcquisitionBruitCDS.config['channel']='DDRAGO-R'
elif band in['J','H']:
AcquisitionBruitCDS.config['channel']='CAGIRE'
# Set filter band
AcquisitionBruitCDS.config['filter_band']=band
# Simulation des quatres images
AcquisitionBruitCDS.number=1
AcquisitionBruitCDS.Rampsimulation('data')
AcquisitionBruitCDS.number=2
AcquisitionBruitCDS.Rampsimulation('data')
AcquisitionBruitCDS.AcquisitionBruitCDS('data')
| notebooks/Simulateur SVOM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def extraction(csv_entree, csv_sortie_ambig, csv_sortie_inc, stats_txt):
"""
Fonction permettant, depuis un CSV extrait avec les colonnes
- ID (numéro de token)
- TOKEN (mot_forme)
- LEMME (lemme ou liste des lemmes, séparés par des '|'
- POS (pos ou liste des pos, séparés par des '|',
1 - d'extraire les tokens où subsiste une ambiguïté sur le POS
dans un nouveau CSV avec le contexte (3 tokens à gauche,
3 tokens à droite, ainsi que leurs POS respectifs)
2 - de calculer :
- le total de tokens dans le fichier,
- le pourcentage de lemmes résolus,
- le pourcentage de pos résolus,
- le nombre de lemmes inconnus,
- le pourcentage de tokens extraits car toujours ambigus.
:param csv_entree: Le chemin interne du fichier CSV dont on
souhaite extraire les tokens et calculer les pourcentages.
:param csv_sortie: Le chemin interne du fichier CSV où seront
extraits les tokens encore ambigus.
"""
import csv
# Dictionnaires servant à l'écriture du nouveau CSV avec tokens ambigus.
dico_mots = {}
dico_local = {}
dico_ambig = {}
dico_inc = {}
# Compteur pour les identifiants de dico_mots.
identifiant = 1
# Séparateur de valeurs pour les colonnes contenant le POS.
intervale = " - "
# Colonnes du futur fichier CSV avec tokens ambigus.
colonnes = ['POS g', 'G1', 'G2', 'G3', 'TOKEN', 'D1', 'D2', 'D3', 'POS d']
# Compteurs servant à l'établissement des pourcentages.
nb_total_tokens = 0
pos_ambigus = 0
lemmes_uniques = 0
pos_uniques = 0
lemmes_inc = 0
# Lecture du fichier CSV d'entrée et stockage de l'information dans csv_lu.
with open(csv_entree) as csv_a_lire:
csv_lu = csv.DictReader(csv_a_lire, delimiter=";") #
for row in csv_lu:
# Établissement du dictionnaire servant à la création du nouveau CSV.
dico_mots[int(row['ID'])] = row
# Calcul du nombre de lemmes et pos résolus, ainsi que du total de tokens.
if "|" not in row['LEMMES']:
lemmes_uniques += 1
if "|" not in row['POS']:
pos_uniques += 1
if row['LEMMES'] == 'INC':
lemmes_inc += 1
nb_total_tokens += 1
# On boucle sur le dictionnaire des tokens pour créer le concordancier.
for identifiant in dico_mots.keys():
# Création des identifiants des contextes les plus éloignés.
g1 = identifiant - 3
d3 = identifiant + 3
# Conditions : on ne créera ces entrées de dictionnaire que si le POS est ambigu,
# et s'il y a bien trois tokens de contexte à inclure de chaque côté
# (ce qui sera le cas à partir du token n°4 et jusqu'au token n°-4).
if g1 in dico_mots.keys() and d3 in dico_mots.keys() and "|" in dico_mots[identifiant]['POS']:
# Si les conditions sont réunies, on peut ajouter 1 au compteur de POS ambigus.
pos_ambigus += 1
# On crée les identifiants des autres tokens de contexte.
g2 = identifiant - 2
g3 = identifiant - 1
d1 = identifiant + 1
d2 = identifiant + 2
# On crée une liste des POS pour le contexte gauche et droit, séparément.
posg = [dico_mots[g1]['POS'], dico_mots[g2]['POS'], dico_mots[g3]['POS']]
posd = [dico_mots[d1]['POS'], dico_mots[d2]['POS'], dico_mots[d3]['POS']]
# On ajoute au dictionnaire local les valeurs construites
# d'après le token et son contexte.
dico_local['posG'] = intervale.join(posg)
dico_local['G1'] = dico_mots[g1]['TOKEN']
dico_local['G2'] = dico_mots[g2]['TOKEN']
dico_local['G3'] = dico_mots[g3]['TOKEN']
dico_local['token'] = dico_mots[identifiant]['TOKEN'] + " - " + str(identifiant)
dico_local['D1'] = dico_mots[d1]['TOKEN']
dico_local['D2'] = dico_mots[d2]['TOKEN']
dico_local['D3'] = dico_mots[d3]['TOKEN']
dico_local['posD'] = intervale.join(posd)
# On ajoute le dictionnaire pour le token en cours
# dans le dictionnaire final, comme valeur,
# avec pour identifiant celui du token central.
dico_ambig[identifiant] = dico_local
identifiant += 1
dico_local = {}
elif g1 in dico_mots.keys() and d3 in dico_mots.keys() and dico_mots[identifiant]['POS'] == "Inconnu":
# On crée les identifiants des autres tokens de contexte.
g2 = identifiant - 2
g3 = identifiant - 1
d1 = identifiant + 1
d2 = identifiant + 2
# On crée une liste des POS pour le contexte gauche et droit, séparément.
posg = [dico_mots[g1]['POS'], dico_mots[g2]['POS'], dico_mots[g3]['POS']]
posd = [dico_mots[d1]['POS'], dico_mots[d2]['POS'], dico_mots[d3]['POS']]
# On ajoute au dictionnaire local les valeurs construites
# d'après le token et son contexte.
dico_local['posG'] = intervale.join(posg)
dico_local['G1'] = dico_mots[g1]['TOKEN']
dico_local['G2'] = dico_mots[g2]['TOKEN']
dico_local['G3'] = dico_mots[g3]['TOKEN']
dico_local['token'] = dico_mots[identifiant]['TOKEN'] + " - " + str(identifiant)
dico_local['D1'] = dico_mots[d1]['TOKEN']
dico_local['D2'] = dico_mots[d2]['TOKEN']
dico_local['D3'] = dico_mots[d3]['TOKEN']
dico_local['posD'] = intervale.join(posd)
# On ajoute le dictionnaire pour le token en cours
# dans le dictionnaire final, comme valeur,
# avec pour identifiant celui du token central.
dico_inc[identifiant] = dico_local
identifiant += 1
dico_local = {}
# On ouvre le CSV de sortie en mode écriture et on écrit les noms des colonnes.
with open(csv_sortie_ambig, 'w') as csv_a_ecrire:
a_ecrire = csv.DictWriter(csv_a_ecrire, fieldnames = colonnes)
# Écrire les noms des colonnes.
a_ecrire.writeheader()
# On boucle sur les clés du dictionnaire final.
for identifiant in dico_ambig.keys():
a_ecrire.writerow({
'POS g' : dico_ambig[identifiant]['posG'],
'G1' : dico_ambig[identifiant]['G1'],
'G2' : dico_ambig[identifiant]['G2'],
'G3' : dico_ambig[identifiant]['G3'],
'TOKEN' : dico_ambig[identifiant]['token'],
'D1' : dico_ambig[identifiant]['D1'],
'D2' : dico_ambig[identifiant]['D2'],
'D3' : dico_ambig[identifiant]['D3'],
'POS d' : dico_ambig[identifiant]['posD']
})
with open(csv_sortie_inc, 'w') as csv_a_ecrire:
a_ecrire = csv.DictWriter(csv_a_ecrire, fieldnames = colonnes)
# Écrire les noms des colonnes.
a_ecrire.writeheader()
# On boucle sur les clés du dictionnaire final.
for identifiant in dico_inc.keys():
a_ecrire.writerow({
'POS g' : dico_inc[identifiant]['posG'],
'G1' : dico_inc[identifiant]['G1'],
'G2' : dico_inc[identifiant]['G2'],
'G3' : dico_inc[identifiant]['G3'],
'TOKEN' : dico_inc[identifiant]['token'],
'D1' : dico_inc[identifiant]['D1'],
'D2' : dico_inc[identifiant]['D2'],
'D3' : dico_inc[identifiant]['D3'],
'POS d' : dico_inc[identifiant]['posD']
})
# L'écriture du CSV étant terminée, on calcule les pourcentages.
pourcentage_lemmes = lemmes_uniques * 100 / nb_total_tokens
pourcentage_pos = pos_uniques * 100 / nb_total_tokens
pourcentage_lemmes_inc = lemmes_inc * 100 / nb_total_tokens
pourcentage_extraits = pos_ambigus * 100 / (nb_total_tokens-4)
# On donne le résultat des calculs de pourcentages.
with open(stats_txt, 'w') as txttobe:
txttobe.write("À partir du fichier '" + csv_entree.split("/")[-1] + "' :\n\n")
txttobe.write(str(round(pourcentage_lemmes,2)) + "% de lemmes uniques.\n")
txttobe.write(str(round(pourcentage_pos,2)) + "% de POS uniques.\n")
txttobe.write(str(lemmes_inc) + " lemmes inconnus, soit " + str(round(pourcentage_lemmes_inc,2)) + "%.\n")
txttobe.write(str(pos_ambigus) + "(" + str(round(pourcentage_extraits,2)) + "%) POS ambigus extraits dans '" + csv_sortie_ambig.split("/")[-1] + "'.\n")
# +
extraction('/home/erminea/Documents/CONDE/Encodage/basnage_tokens.csv',
'/home/erminea/Documents/CONDE/Encodage/basnage_tokens_ambig.csv',
'/home/erminea/Documents/CONDE/Encodage/basnage_tokens_inc.csv',
'/home/erminea/Documents/CONDE/Encodage/basnage_stats.txt')
extraction('/home/erminea/Documents/CONDE/Encodage/berault_tokens.csv',
'/home/erminea/Documents/CONDE/Encodage/berault_tokens_ambig.csv',
'/home/erminea/Documents/CONDE/Encodage/berault_tokens_inc.csv',
'/home/erminea/Documents/CONDE/Encodage/berault_stats.txt')
extraction('/home/erminea/Documents/CONDE/Encodage/instructions_tokens.csv',
'/home/erminea/Documents/CONDE/Encodage/instructions_tokens_ambig.csv',
'/home/erminea/Documents/CONDE/Encodage/instructions_tokens_inc.csv',
'/home/erminea/Documents/CONDE/Encodage/instructions_stats.txt')
extraction('/home/erminea/Documents/CONDE/Encodage/merville_tokens.csv',
'/home/erminea/Documents/CONDE/Encodage/merville_tokens_ambig.csv',
'/home/erminea/Documents/CONDE/Encodage/merville_tokens_inc.csv',
'/home/erminea/Documents/CONDE/Encodage/merville_stats.txt')
extraction('/home/erminea/Documents/CONDE/Encodage/pesnelle_tokens.csv',
'/home/erminea/Documents/CONDE/Encodage/pesnelle_tokens_ambig.csv',
'/home/erminea/Documents/CONDE/Encodage/pesnelle_tokens_inc.csv',
'/home/erminea/Documents/CONDE/Encodage/pesnelle_stats.txt')
extraction('/home/erminea/Documents/CONDE/Encodage/rouille_tokens.csv',
'/home/erminea/Documents/CONDE/Encodage/rouille_tokens_ambig.csv',
'/home/erminea/Documents/CONDE/Encodage/rouille_tokens_inc.csv',
'/home/erminea/Documents/CONDE/Encodage/rouille_stats.txt')
extraction('/home/erminea/Documents/CONDE/Encodage/ruines_tokens.csv',
'/home/erminea/Documents/CONDE/Encodage/ruines_tokens_ambig.csv',
'/home/erminea/Documents/CONDE/Encodage/ruines_tokens_inc.csv',
'/home/erminea/Documents/CONDE/Encodage/ruines_stats.txt')
extraction('/home/erminea/Documents/CONDE/Encodage/tac_tokens.csv',
'/home/erminea/Documents/CONDE/Encodage/tac_tokens_ambig.csv',
'/home/erminea/Documents/CONDE/Encodage/tac_tokens_inc.csv',
'/home/erminea/Documents/CONDE/Encodage/tac_stats.txt')
extraction('/home/erminea/Documents/CONDE/Encodage/terrien_tokens.csv',
'/home/erminea/Documents/CONDE/Encodage/terrien_tokens_ambig.csv',
'/home/erminea/Documents/CONDE/Encodage/terrien_tokens_inc.csv',
'/home/erminea/Documents/CONDE/Encodage/terrien_stats.txt')
| corpus-construction/disambiguate-lemmatization-in-corrected-file/REV_3_extract_derniers_ambigus_et_inconnus_avec_ctxt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import tensorflow_probability as tfp
import numpy as np
from timeit import default_timer
from collections import namedtuple
import pylab as plt
import os
import sys
import gpflow as gp
import astropy.coordinates as ac
import astropy.time as at
import astropy.units as au
from scipy.cluster.vq import kmeans2
from bayes_tec.frames import ENU
from bayes_tec.datapack import DataPack
from bayes_tec.utils.data_utils import make_coord_array, calculate_weights
plt.style.use('ggplot')
float_type = tf.float64
jitter = 1e-6
# +
from gpflow.transforms import Identity
class Parameter(object):
def __init__(self, name, value, transform=Identity(), prior=None,
trainable=True, dtype=float_type, unconstrained_tensor=None):
self.name = name
self.prior = prior # pylint: disable=W0201
self.transform = transform # pylint: disable=W0201
if unconstrained_tensor is None:
self._initial_value_tensor = tf.convert_to_tensor(value,dtype=dtype)
self._unconstrained_tensor_ref = tf.get_variable(name, dtype=dtype,
initializer=self.transform.backward_tensor(self._initial_value_tensor),
trainable=trainable,
use_resource=True)
self._unconstrained_tensor = tf.identity(self.unconstrained_tensor_ref)
else:
self._unconstrained_tensor_ref = None
self._unconstrained_tensor = unconstrained_tensor
self._initial_value_tensor = self.transform.forward_tensor(unconstrained_tensor)
self._constrained_tensor = self.transform.forward_tensor(self.unconstrained_tensor)
self._prior_tensor = self._build_prior(self.unconstrained_tensor, self.constrained_tensor)
@property
def unconstrained_tensor_ref(self):
return self._unconstrained_tensor_ref
@property
def unconstrained_tensor(self):
return self._unconstrained_tensor
@property
def constrained_tensor(self):
return self._constrained_tensor
@property
def prior_tensor(self):
"""log P(constrained_param) + log |det transform(unconstrained_param)|"""
return self._prior_tensor
@property
def initializer(self):
if self.unconstrained_tensor_ref is None:
raise ValueError("No variable referenced")
return self.unconstrained_tensor_ref.initializer
def assign_op(self, value):
if self.unconstrained_tensor_ref is None:
raise ValueError("No variable referenced")
return tf.assign(self._unconstrained_tensor_ref,self.transform.backward_tensor(value))
def _build_prior(self, unconstrained_tensor, constrained_tensor):
prior_name = '{}_logp'.format(self.name)
if self.prior is None:
logp_param = tf.constant(0.0, float_type)
else:
logp_param = self.prior(constrained_tensor)
log_jacobian = self.transform.log_jacobian_tensor(unconstrained_tensor)
return tf.squeeze(tf.add(logp_param, log_jacobian, name=prior_name))
class Kernel(object):
def __init__(self, time_lengthscale, dir_lengthscale, ant_lengthscale, dot_var, dot_offset,
active_dims_time=None,active_dims_dir=None,active_dims_ant=None):
self.dir_lengthscale = dir_lengthscale
self.time_lengthscale = time_lengthscale
self.ant_lengthscale = ant_lengthscale
self.dot_var = dot_var
self.dot_offset = dot_offset
self.active_dims_time = active_dims_time or slice(0,1,1)
self.active_dims_dir = active_dims_dir or slice(1,3,1)
self.active_dims_ant = active_dims_ant or slice(3,5,1)
def scaled_square_dist_batched(self,X, X2, lengthscale):
"""
X: tensor B, N, D
X2: tensor B, M, D (or 1, M, D) and will be broadcast to B, M ,D
Return:
tensor B, N, M
"""
# Clipping around the (single) float precision which is ~1e-45.
X = X / lengthscale
Xs = tf.reduce_sum(tf.square(X), axis=2)#B,N
if X2 is None:
dist = -2.*tf.matmul(X,X,transpose_b=True)
dist += Xs[:,:,None] + Xs[:,None,:]
return tf.maximum(dist, 1e-40)
# B (1), M, D
X2 = X2 / lengthscale
X2s = tf.reduce_sum(tf.square(X2), axis=2)# B (1), M
dist = -2 * tf.matmul(X, X2, transpose_b=True)
dist += Xs[:,:,None] + X2s[:,None,:]
return dist
def scaled_square_dist(self,X, X2, lengthscale):
"""
X: tensor N, D
X2: tensor M, D
Return:
tensor N, M
"""
# Clipping around the (single) float precision which is ~1e-45.
X = X / lengthscale
Xs = tf.reduce_sum(tf.square(X), axis=1)#N
if X2 is None:
dist = -2.*tf.matmul(X,X,transpose_b=True)
dist += Xs[:,None] + Xs[None,:]
return tf.maximum(dist, 1e-40)
# M, D
X2 = X2 / lengthscale
X2s = tf.reduce_sum(tf.square(X2), axis=1)# M
dist = -2 * tf.matmul(X, X2, transpose_b=True)
dist += Xs[:,None] + X2s[None,:]
return dist
def _clipped_sqrt(self, r2):
# Clipping around the (single) float precision which is ~1e-45.
return tf.sqrt(tf.maximum(r2, 1e-40))
def K(self, X, X2=None):
"""Returns the covariance at X and X2.
(dot_offset + dot_var*X.X2) * M52(time) * RBF(dir) * M12(ant)
Args:
:param X: float Tensor [N, ndims]
:param X2: float Tensor [M, ndims]
Returns:
float Tensor [N,M]
"""
Xt = X[:,self.active_dims_time]
Xd = X[:,self.active_dims_dir]
Xa = X[:,self.active_dims_ant]
if X2 is None:
X2t = None
X2d = None
X2a = None
else:
X2t = X2[:,self.active_dims_time]
X2d = X2[:,self.active_dims_dir]
X2a = X2[:,self.active_dims_ant]
r2t = self.scaled_square_dist(Xt, X2t, self.time_lengthscale)
rt = self._clipped_sqrt(r2t)
r2d = self.scaled_square_dist(Xd, X2d, self.dir_lengthscale)
ra = self._clipped_sqrt(self.scaled_square_dist(Xa, X2a, self.ant_lengthscale))
combined_exp = tf.accumulate_n([np.sqrt(5.)*rt, 0.5*r2d, ra])
combined_exp = tf.exp(-combined_exp)
dot_kern = self.dot_offset + self.dot_var * tf.matmul(Xd, Xd if X2d is None else X2d, transpose_b=True)
time_m52 = (1. + np.sqrt(5.) * rt + (5./3.) * r2t)
return combined_exp*dot_kern*time_m52
def Kdiag(self, X):
"""Returns the diag of the covariance at X.
Args:
:param X: float Tensor [N, ndims]
Returns:
float Tensor [N]
"""
return self.dot_var*tf.linalg.norm(X,axis=-1,keepdims=False) + self.dot_offset
def make_solsets(datapack,output_solset, screen_res=15, extend = 0., solset='sol000'):
screen_solset = "screen_{}".format(output_solset)
datapack.switch_solset(solset)
datapack.select(ant=None,time=None, dir=None, freq=None, pol=slice(0,1,1))
axes = datapack.__getattr__("axes_{}".format('phase'))
antenna_labels, antennas = datapack.get_antennas(axes['ant'])
patch_names, directions = datapack.get_sources(axes['dir'])
timestamps, times = datapack.get_times(axes['time'])
freq_labels, freqs = datapack.get_freqs(axes['freq'])
pol_labels, pols = datapack.get_pols(axes['pol'])
Npol, Nd, Na, Nf, Nt = len(pols), len(directions), len(antennas), len(freqs), len(times)
# screen_directions = dialated_faceted(directions.ra.rad.mean(), directions.dec.rad.mean(),
# N=screen_res)
screen_ra = np.linspace(np.min(directions.ra.rad) - extend*np.pi/180.,
np.max(directions.ra.rad) + extend*np.pi/180., screen_res)
screen_dec = np.linspace(max(-90.*np.pi/180.,np.min(directions.dec.rad) - extend*np.pi/180.),
min(90.*np.pi/180.,np.max(directions.dec.rad) + extend*np.pi/180.), screen_res)
screen_directions = np.stack([m.flatten() \
for m in np.meshgrid(screen_ra, screen_dec, indexing='ij')], axis=1)
screen_directions = ac.SkyCoord(screen_directions[:,0]*au.rad,screen_directions[:,1]*au.rad,frame='icrs')
Nd_screen = screen_res**2
datapack.switch_solset(output_solset,
array_file=DataPack.lofar_array,
directions = np.stack([directions.ra.rad,directions.dec.rad],axis=1), patch_names=patch_names)
datapack.add_freq_indep_tab('tec', times.mjd*86400., pols = pol_labels)
datapack.add_freq_dep_tab('amplitude', times.mjd*86400., pols = pol_labels,freqs=freqs)
datapack.add_freq_dep_tab('phase', times.mjd*86400., pols = pol_labels,freqs=freqs)
datapack.switch_solset(screen_solset,
array_file = DataPack.lofar_array,
directions = np.stack([screen_directions.ra.rad,screen_directions.dec.rad],axis=1))
datapack.add_freq_indep_tab('tec', times.mjd*86400., pols = pol_labels)
datapack.add_freq_dep_tab('amplitude', times.mjd*86400., pols = pol_labels,freqs=freqs)
datapack.add_freq_dep_tab('phase', times.mjd*86400., pols = pol_labels,freqs=freqs)
datapack.switch_solset(solset)
def get_solset_coords(datapack,solset):
datapack.switch_solset(solset)
axes = datapack.axes_phase
antenna_labels, antennas = datapack.get_antennas(axes['ant'])
patch_names, directions = datapack.get_sources(axes['dir'])
timestamps, times = datapack.get_times(axes['time'])
pol_labels, pols = datapack.get_pols(axes['pol'])
antennas_enu = antennas.transform_to(ENU(obstime=times[0],location=datapack.array_center))
X_a = np.array([antennas_enu.east.value,
antennas_enu.north.value]).T/1000.
X_d = np.array([directions.ra.deg - directions.ra.deg.mean(), directions.dec.deg - directions.dec.deg.mean()]).T
X_t = (times.mjd*86400 - times[0].mjd*86400.)[:,None]
return X_t, X_d, X_a
# +
UpdateResult = namedtuple('UpdateResult',['x_samples','z_samples','log_prob', 'acceptance','step_size'])
class TargetDistribution(object):
def __init__(self,
kerns,
z_tm1,
X_t,
X_tm1,
Y_t,
freqs,
L_tm1,
num_chains,
max_lik=True,
step_size=0.01,
Y_sigma=1.,
approximate_posterior = 'mfsg',
prior_opts = {}):
"""
The target distribution of the Bayes filter.
Args:
:param z_tm1: float Tensor [S, num_chains, M, Np]
:param X: float Tensor [N,K]
:param last_X: float Tensor [Np, K]
:param Y: float Tensor [D, N, Nf]
:param Y_std: float Tensor [D, N, Nf]
:param freqs: float Tensor [Nf]
:param L11: float Tensor [M, Np, Np]
"""
self.M = tf.shape(z_tm1)[2]
self.S = tf.shape(z_tm1)[0]
self.num_chains = num_chains
self.N = tf.shape(Y_t)[1]
self.Np = tf.shape(z_tm1)[-1]
#M, N, N
self.Kx_t = kerns[0](X_t)
#M, N, N
self.Kh_t = kerns[1](X_t)
#M, Np, Np
self.Kx_tm1 = kerns[0](X_tm1)
#M, Np, Np
self.Kh_tm1 = kerns[1](X_tm1)
self.jitter = tf.convert_to_tensor(jitter,dtype=float_type,name='jitter')
self.offset_t = self.jitter*tf.eye(self.N,dtype=float_type)
self.offset_tm1 = self.jitter*tf.eye(self.Np,dtype=float_type)
#M, N, N
self.Lx_t = tf.cholesky(self.Kx_t + self.offset_t)
#M, N, N
self.Lh_t = tf.cholesky(self.Kh_t + self.offset_t)
#M, Np, Np
self.Lx_tm1 = tf.cholesky(self.Kx_tm1 + self.offset_tm1)
#M, Np, Np
self.Lh_tm1 = tf.cholesky(self.Kh_tm1 + self.offset_tm1)
#M, Np, N
self.Kx_tm1t = kern(X_tm1, X_t)
#S, num_chains, M, N
self.z_tm1 = z_tm1
#D, N, Nf
self.Y = Y
#Nf
self.freqs = freqs
self.step_size = tf.get_variable(
name='step_size',
initializer=lambda: tf.constant(step_size,dtype=tf.float64),
use_resource=True,
dtype=tf.float64,
trainable=False)
self.Y_sigma = Y_sigma
self.max_lik = max_lik
self.approximate_posterior = approximate_posterior
self.prior_opts = prior_opts
def likelihood(self, x_t):
"""
Calculate the likelihood of Y given hidden_state.
I.e.
sum_i log[P(Y_j(X_i) | X_i)]
If Y_j in C^Nf and assuming independence between
real, imag, and components we get,
sum_i sum_j log[P(Re[Y_j(X_i)] | X_i)]
+ log[P(imag[Y_j(X_i)] | X_i)]
Args:
:param x_t: float Tensor [num_chains, M, N+H]
Returns:
float Tensor [num_chains]
"""
#num_chains, N
x_t = x_t[:, 0, :self.N]
#Nf
tec_conv = tf.div(tf.cast(-8.448e6,tf.float64),self.freqs,name='tec_conv')
#num_chains, N, Nf
phi = tec_conv*x_t[:,:,None]
g_real = tf.cos(phi)
g_imag = tf.sin(phi)
#D, num_chains, N, Nf
g = tf.stack([g_real, g_imag],axis=0,name='g')
L = tfp.distributions.MultivariateNormalDiag(loc=g, scale_identity_multiplier = self.sigma_amp,#scale_diag=self.sigma_amp*self.Y_std[:, None,:,:]
name='data_likelihood')
#D,num_chains, N
logp = L.log_prob(self.Y[:,None,:,:])
#num_chains
return tf.reduce_sum(logp,axis=[0, 2])
def _mfsg_logp(self,x_t,**prior_opts):
'''
Evaluates log probability of the predict step assuming
a the mean field single Gaussian (MF) representing the
resulting mixture of Gaussians as a single Gaussian.
It is less correct than the (MF) approximation
but has complexity O(SM^2 + M^3).
Args:
:param x_t: float Tensor [num_chains, M, N]
Returns:
float Tensor [num_chains]
'''
pass
def _mf_logp(self,x_t, num_sigma_points = 10,**prior_opts):
'''
Evaluates log probability of the predict step assuming
a the mean field approximation (MF). It is the most correct
approximation but also the highest complexity O(SM^3).
Args:
:param x_t: float Tensor [num_chains, M, N]
Returns:
float Tensor [num_chains]
'''
num_sigma_points = tf.convert_to_tensor(num_sigma_points, dtype=tf.int32)
s = tf.minimum(num_sigma_points, tf.shape(self.z_tm1)[0])
shuffle_index = tf.random_shuffle(tf.range(tf.shape(self.z_tm1)[0],dtype=tf.int32))[:s]
#s, num_chains, M, Np
z_tm1 = tf.gather(self.z_tm1, shuffle_index, axis=0,name='z_tm1')
x_tm1 = tf.einsum('mij,snmj->snmi',self.L11, z_tm1)
# log[P(Z_i | Z_i-1,s)] = log[N[m_i, C]] + log
#M, Np, N
A = tf.matrix_triangular_solve(self.L11,self.K10)
# #s, num_chains, M, Np, N
# A_expand = tf.tile(A[None, None, :, :, :], [s, self.num_chains,1,1,1])
# #s, num_chains, M, N
# m = tf.matmul(A_expand, z_tm1[:,:,:,:,None],transpose_a=True)[:,:,:,:,0]
#s, num_chains, M, N
m = tf.einsum('mij,snmi->snmj',A,x_tm1)
#M, N, N
C = self.K00 - tf.matmul(A, A, transpose_a=True)
#M, N, N
L = tf.cholesky(C + tf.cast(jitter,tf.float64)*tf.eye(tf.shape(C)[2],dtype=tf.float64))
P = tfp.distributions.MultivariateNormalTriL(loc=m, scale_tril=L[None, None,:,:,:])
#s, num_chains, M
log_prob = P.log_prob(x_t) - tf.reduce_sum(tf.log(tf.matrix_diag_part(self.L00)),axis=1)
#s, num_chains
log_prob = tf.reduce_sum(log_prob, axis=2)
#num_chains
log_prob = tf.reduce_logsumexp(log_prob,axis=0) - tf.log(tf.cast(s,tf.float64))
log_prob.set_shape(tf.TensorShape([self.num_chains]))
return log_prob
def _gpp_logp(self,x_t,**prior_opts):
'''
Evaluates log probability of the predict step assuming
a Gaussian previous posterior (GPP) and conditional
independence of the hyperparameters. In this case,
marginalisation is analytic.
Args:
:param x_t: float Tensor [num_chains, M, N]
Returns:
float Tensor [num_chains]
'''
#S, num_chains, M, Np,1
x_tm1 = tf.einsum('mij,snmj->snmi',self.L11, self.z_tm1)[..., None]
#num_chains, M, Np,1
m_tm1 = tf.reduce_mean(x_tm1,axis=0)
#num_chains, M, Np,Np
m2 = tf.matmul(m_tm1,m_tm1, transpose_b=True)
#num_chains, M, Np, Np
C_tm1 = tf.reduce_mean(tf.matmul(x_tm1, x_tm1,transpose_b=True), axis=0) - m2
def prior_logp(self, x_t):
"""
Calculate the predict step, i.e.
log[P(X_i | Y_i-1)] = log E_i-1[P(X_i | X_i-1)]
= log sum_s P(X_i | X_i-1,s) - log S
= logsumexp_s log[P(X_i | X_i-1,s)] - log S
If we transform the variables through,
X = L.Z + m => log P(X) = log P(Z) - log det L
log[P(X_i | X_i-1,s)] = log[P(Z_i | Z_i-1,s)] - log det L_i
Assumes hidden state is a GP marginal.
Args:
:param x_t: float Tensor [num_chains, M, N]
Returns:
[num_chains]
"""
if self.approximate_posterior == 'mfsg':
log_prob = self._mfsg_logp(x_t, **self.prior_opts)
elif self.approximate_posterior == 'mf':
log_prob = self._mf_logp(x_t,**self.prior_opts)
elif self.approximate_posterior == 'gpp':
log_prob = self._gpp_logp(x_t,**self.prior_opts)
log_prob.set_shape(tf.TensorShape([self.num_chains]))
return log_prob
def unnormalized_logp(self,z_t):
"""
Returns the unnormalized probability density of the Bayes filter posterior.
log P(y_t | z_t) + log (1/S) sum_s P(z_t | z^s_t-1)
Args:
:param z_t: float Tensor [num_chains, M*(N+H)]
Returns:
[num_chains]
"""
#num_chains, M, N+H
z_t = tf.cast(tf.reshape(z_t,[self.num_chains, self.M, -1]),tf.float64)
#num_chains, M, N+H
x_t = tf.einsum('mij,nmj->nmi', self.L00, z_t)
# #num_chains, M, N, N
# L00_expand = tf.tile(self.L00[None, :, :self.N, :self.N], [self.num_chains, 1,1,1])
# #num_chains, N
# x_t = tf.matmul(L00_expand, z_t[:, :, :self.N, None])[:, 0, :, 0]
#num_chains, N
x_t = x_t[:, 0, :self.N]
max_lik_logp = self.likelihood(x_t)
full_post_logp = max_lik_logp + self.prior_logp(x_t)
logp = tf.cond(self.max_lik,
lambda: max_lik_logp,
lambda: full_post_logp)
return logp#self.likelihood(x_t) + self.prior_logp(x_t)
def sample(self,num_samples=10, step_size = 1., num_leapfrog_steps=2, target_rate=0.75):
hmc = tfp.mcmc.HamiltonianMonteCarlo(
target_log_prob_fn=self.unnormalized_logp,
num_leapfrog_steps=num_leapfrog_steps,#tf.random_shuffle(tf.range(3,60,dtype=tf.int64))[0],
step_size=self.step_size,
step_size_update_fn=tfp.mcmc.make_simple_step_size_update_policy(target_rate=target_rate))
# step_size_update_fn=lambda v, _: v)
#num_chains, M, Np
q0 = tf.reduce_mean(self.z_tm1,axis=0)
q0 = tf.reshape(q0,(self.num_chains, -1))
# q0.set_shape(tf.TensorShape([self.num_chains, None]))
# Run the chain (with burn-in).
z_samples, kernel_results = tfp.mcmc.sample_chain(
num_results=num_samples,
num_burnin_steps=0,
current_state=q0,
kernel=hmc)
avg_acceptance_ratio = tf.reduce_mean(tf.exp(tf.minimum(kernel_results.log_accept_ratio, 0.)),name='avg_acc_ratio')
posterior_log_prob = tf.reduce_sum(kernel_results.accepted_results.target_log_prob,name='marginal_log_likelihood')
z_samples = tf.reshape(z_samples, tf.concat([tf.shape(z_samples)[:2], [self.M], [-1]],axis=0))
x_samples = tf.einsum("mij,snmj->snmi",self.L00,z_samples)
res = UpdateResult(x_samples, z_samples, posterior_log_prob, avg_acceptance_ratio, kernel_results.extra.step_size_assign)
return res
# -
# +
a = np.random.uniform(size=10)
A = np.random.uniform(size=10)**2
p = np.random.uniform(size=10)
p /= p.sum()
m = np.sum(p*a)
M = np.sum(p*(A + a*a - m*m))
x = np.linspace(-5,5,1000)
y = np.sum([pi*np.exp(-0.5*(x - ai)**2/Ai)/np.sqrt(2*np.pi*Ai) for (pi,ai,Ai) in zip(p,a,A)], axis=0)
print(y.sum())
# y /= y.sum()
Y = np.exp(-0.5*(x-m)**2 / M)/np.sqrt(2*np.pi*M)
print(Y.sum())
# Y /= Y.sum()
print(m,M)
plt.plot(x,Y)
plt.plot(x,y)
plt.show()
# -
| notebooks/devel/path_integration_approach.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Perturbing histograms using the Laplace mechanism
# Acknowledgement: The core functions defined in this notebook are derived from source code developed at Data61 by <NAME> and <NAME>.
# +
import numpy as np
from pandas import DataFrame, Index, MultiIndex, RangeIndex, read_csv
from typing import cast, Mapping, NewType, Sequence, Text, Tuple
# -
Record = NewType('Record', Tuple[int, ...])
Dataset = NewType('Dataset', Mapping[Record, float])
# ## Utility functions
def encode_values_as_integers(raw_data: DataFrame, include_all_in_range: bool = False) -> DataFrame:
data = raw_data.copy()
for column_name in data.columns:
if include_all_in_range:
sorted_values = list(range(min(data[column_name]), max(data[column_name]) + 1))
else:
sorted_values = sorted({value for value in data[column_name]})
value_dict = {value: i for i, value in enumerate(sorted_values)}
data[column_name] = raw_data[column_name].map(value_dict)
return data
def get_dataset_from_data(data: DataFrame) -> Dataset:
maxima = data.agg('max')
column_names = [column for column in data.columns]
if len(column_names) > 1:
index = MultiIndex.from_product([RangeIndex.from_range(range(maxima[column_name] + 1))
for column_name in column_names], names=column_names)
else:
index = Index(range(maxima[column_names[0]] + 1), name=column_names[0])
dataset_as_series = data.groupby(column_names).size().reindex(index=index, fill_value=0)
d = dataset_as_series.to_dict()
if len(column_names) == 1:
d = {(k,): v for k, v in d.items()}
dataset = cast(Dataset, d)
return dataset
def as_data_frame(dataset: Dataset, feature_names: Sequence[Text], output_index_name: Text) -> DataFrame:
index = MultiIndex.from_tuples(dataset.keys(), names=feature_names)
return DataFrame.from_dict(cast(dict, dataset),
'index',
columns=[output_index_name]).reindex(index, fill_value=0)
# ## Read in the raw data
data_path = '1_PIF/our-synthetic.csv'
feature_names = ['AGE', 'eye_color', 'countryofresidence', 'POSTCODE'] # Currently only numeric features are supported, but this is easily fixed.
raw_data = read_csv(data_path)[feature_names]
raw_data.head(10)
# ## Encode the raw data
data_encoded = encode_values_as_integers(raw_data=raw_data, include_all_in_range=False)
data_encoded.head(10)
# ## Represent the raw data as a histogram
dataset = get_dataset_from_data(data_encoded)
as_data_frame(dataset, feature_names, 'count').head(10)
# ## The Laplace mechanism
def apply_laplace_mechanism(dataset: Dataset, epsilon: float, sensitivity: float = 1.0):
return cast(Dataset, dict((possible_record, np.random.laplace(loc=true_count, scale=sensitivity / epsilon))
for possible_record, true_count in dataset.items()))
# ## Perturb the histogram with a 'low' value of epsilon (= 0.05)
perturbed_dataset_low_epsilon = apply_laplace_mechanism(dataset=dataset, epsilon=0.05, sensitivity=1)
as_data_frame(perturbed_dataset_low_epsilon, feature_names, 'count').head(10)
# ## Perturb the histogram with a 'medium' value of epsilon (= 0.1)
perturbed_dataset_medium_epsilon = apply_laplace_mechanism(dataset=dataset, epsilon=0.1, sensitivity=1)
as_data_frame(perturbed_dataset_medium_epsilon, feature_names, 'count').head(10)
# ## Perturb the histogram with a 'high' value of epsilon (= 1.0)
perturbed_dataset_high_epsilon = apply_laplace_mechanism(dataset=dataset, epsilon=1.0, sensitivity=1)
as_data_frame(perturbed_dataset_high_epsilon, feature_names, 'count').head(10)
# ## Perturb the histogram with a 'very high' value of epsilon (= 50.0)
perturbed_dataset_very_high_epsilon = apply_laplace_mechanism(dataset=dataset, epsilon=50.0, sensitivity=1)
as_data_frame(perturbed_dataset_very_high_epsilon, feature_names, 'count').head(10)
| Scripts/Round 3/Baysically Measure Zero/differential_privacy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Describe
# Describe and visualizes dataset statistics
# ### Set-up
from sklearn.datasets import load_boston
import pandas as pd
import matplotlib.pyplot as plt
import mlrun
import os
mlrun.set_environment(project = "function-marketplace")
context = mlrun.get_or_create_ctx("function-marketplace")
# ### Loading dataset
# We will use boston hose prices dataset from sklearn
boston_dataset = load_boston()
print(boston_dataset["DESCR"])
boston_df = pd.DataFrame(data = boston_dataset["data"],columns = boston_dataset["feature_names"])
boston_df["TARGET"] = boston_dataset["target"]
# taking first 100 rows - so calculating all the histograms will be fast.
boston_df = boston_df[:100]
os.mkdir('artifacts')
boston_df.to_parquet("artifacts/boston_housing.parquet")
# ### Importing and running the function
describe = mlrun.import_function("hub://describe").apply(mlrun.auto_mount())
describe_task = mlrun.new_task(name = 'describe_boston',
params = {'key' : 'summary',
'label_column' : 'TARGET',
'plot_hist' : True,
'plot_dest' : os.getcwd()},
inputs = {"table" : "artifacts/boston_housing.parquet"},
artifact_path = os.getcwd()+'/artifacts',
in_path = os.getcwd())
describe.run(describe_task)
# ### Plotting results
from IPython.display import IFrame
IFrame("artifacts/plots/violin.html", width=1300, height=600)
from seaborn import heatmap
plt.figure(figsize = (15,10))
my_heatmap = pd.read_csv("artifacts/plots/correlation-matrix.csv").set_index("Unnamed: 0").drop(["CHAS"],axis = 1).drop(["CHAS"],axis=0)
heatmap(my_heatmap,annot=True)
| describe/describe.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://github.com/awslabs/aws-data-wrangler)
#
# # 17 - Partition Projection
#
# https://docs.aws.amazon.com/athena/latest/ug/partition-projection.html
import awswrangler as wr
import pandas as pd
from datetime import datetime
import getpass
# ## Enter your bucket name:
bucket = getpass.getpass()
# ## Integer projection
# +
df = pd.DataFrame({
"value": [1, 2, 3],
"year": [2019, 2020, 2021],
"month": [10, 11, 12],
"day": [25, 26, 27]
})
df
# -
wr.s3.to_parquet(
df=df,
path=f"s3://{bucket}/table_integer/",
dataset=True,
partition_cols=["year", "month", "day"],
database="default",
table="table_integer",
projection_enabled=True,
projection_types={
"year": "integer",
"month": "integer",
"day": "integer"
},
projection_ranges={
"year": "2000,2025",
"month": "1,12",
"day": "1,31"
},
);
wr.athena.read_sql_query(f"SELECT * FROM table_integer", database="default")
# ## Enum projection
# +
df = pd.DataFrame({
"value": [1, 2, 3],
"city": ["São Paulo", "Tokio", "Seattle"],
})
df
# -
wr.s3.to_parquet(
df=df,
path=f"s3://{bucket}/table_enum/",
dataset=True,
partition_cols=["city"],
database="default",
table="table_enum",
projection_enabled=True,
projection_types={
"city": "enum",
},
projection_values={
"city": "São Paulo,Tokio,Seattle"
},
);
wr.athena.read_sql_query(f"SELECT * FROM table_enum", database="default")
# ## Date projection
# +
ts = lambda x: datetime.strptime(x, "%Y-%m-%d %H:%M:%S")
dt = lambda x: datetime.strptime(x, "%Y-%m-%d").date()
df = pd.DataFrame({
"value": [1, 2, 3],
"dt": [dt("2020-01-01"), dt("2020-01-02"), dt("2020-01-03")],
"ts": [ts("2020-01-01 00:00:00"), ts("2020-01-01 00:00:01"), ts("2020-01-01 00:00:02")],
})
df
# -
wr.s3.to_parquet(
df=df,
path=f"s3://{bucket}/table_date/",
dataset=True,
partition_cols=["dt", "ts"],
database="default",
table="table_date",
projection_enabled=True,
projection_types={
"dt": "date",
"ts": "date",
},
projection_ranges={
"dt": "2020-01-01,2020-01-03",
"ts": "2020-01-01 00:00:00,2020-01-01 00:00:02"
},
);
wr.athena.read_sql_query(f"SELECT * FROM table_date", database="default")
# ## Injected projection
# +
df = pd.DataFrame({
"value": [1, 2, 3],
"uuid": ["761e2488-a078-11ea-bb37-0242ac130002", "b89ed095-8179-4635-9537-88592c0f6bc3", "87adc586-ce88-4f0a-b1c8-bf8e00d32249"],
})
df
# -
wr.s3.to_parquet(
df=df,
path=f"s3://{bucket}/table_injected/",
dataset=True,
partition_cols=["uuid"],
database="default",
table="table_injected",
projection_enabled=True,
projection_types={
"uuid": "injected",
}
);
wr.athena.read_sql_query(
sql=f"SELECT * FROM table_injected WHERE uuid='b89ed095-8179-4635-9537-88592c0f6bc3'",
database="default"
)
# ## Cleaning Up
wr.s3.delete_objects(f"s3://{bucket}/table_integer/")
wr.s3.delete_objects(f"s3://{bucket}/table_enum/")
wr.s3.delete_objects(f"s3://{bucket}/table_date/")
wr.s3.delete_objects(f"s3://{bucket}/table_injected/")
wr.catalog.delete_table_if_exists(table="table_integer", database="default")
wr.catalog.delete_table_if_exists(table="table_enum", database="default")
wr.catalog.delete_table_if_exists(table="table_date", database="default")
wr.catalog.delete_table_if_exists(table="table_injected", database="default");
| tutorials/017 - Partition Projection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PY36]
# language: python
# name: conda-env-PY36-py
# ---
# <div align="right">Creation Env: <font color=blue>Python [conda env:PY36]</font></div>
# # Playing with Python 2.7 Comment and String Syntax in Python 3.6
# This stuff should work in 3.5 as well but has only been tested in 3.6 on Anaconda / IPython
'''this is a quote with a newline.\nIt then continues with more text.
And even more text than that.'''
myVar = '''this is a quote with a newline.\nIt then continues with more text.
And even more text than that.'''
print(myVar)
print('''this is a quote with a newline.\nIt then continues with more text.''')
# +
# Note: note the strange behavior of period in these examples (when it shows versus when it does not)
''' ... and then he said ... something or other'''
# -
'''This is a test.
... This is only a test.'''
print('''This is a test.
... This is only a test.''')
# +
# using formatters on strings, not just inside print()
'Insert %s, ten: %d' % ('string', 10)
# -
'My name is %s and I am %d years old' % ('Mike', 25)
# want to lose the quotes around the string? put in a variable and call with print() or just put in print():
myVar2 = 'My name is %s and I am %d years old' % ('Mike', 25)
print('My name is %s and I am %d years old' % ('Mike', 25))
print(myVar2)
| PY_Basics/str_arr_lst_prnt/TMWP_Comments_and_MultiLine_Strings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# Import the pathlib and csv library
from pathlib import Path
import csv
# Set the file path
import pandas as pd
df = pd.read_csv(r'C:\Users\yfkok\OneDrive\Desktop\Monash Bootcamp\python-homework\PyBank\Resources\budget_data.csv')
#export 1st column into a list
matrix1 = df[df.columns[0]].to_numpy()
date = matrix1.tolist()
#export 2nd column into a list
matrix2 = df[df.columns[1]].to_numpy()
budget = matrix2.tolist()
#merging both lists into a dictionary
budgetdict = dict(zip(date, budget))
#creating a list of numbers which consists the change in profit between two roles of pnl
pnlchange = [budget[i+1]-budget[i] for i in range(len(budget)-1)]
# +
# Initialize metric variables
max_pnl = 0
min_pnl = 0
pnl_average = 0
total_pnl = 0
count_pnl = 0
# Calculate the max, mean, and average of the list of profits/losses
for pnl in budget:
# Sum the total and count variables
total_pnl += pnl
count_pnl += 1
# Logic to determine min and max salaries
if min_pnl == 0:
min_pnl = pnl
elif pnl > max_pnl:
max_pnl = pnl
elif pnl < min_pnl:
min_pnl = pnl
#function to determine which month corresponds to its respective profits/losses
def return_key(val):
for key, value in budgetdict.items():
if value==val:
return key
return('ERROR in CELL')
# alling out dates from dictionary BASED on their pnl metrics
max_month = return_key(max_pnl)
min_month = return_key(min_pnl)
max_pnl_change = max(pnlchange)
min_pnl_change = min(pnlchange)
print(min_pnl_change)
# +
# Calculate the average salary, round to the nearest 2 decimal places
pnl_average = round(sum(pnlchange)/len(pnlchange),2)
# function to print the metrics
# {max_pnl:,} - to include , after every 1000
def print_financial_result():
print("Financial Analysis")
print("-------------------------------------")
print(f"Total Months: {count_pnl:,}")
print(f"Total: {total_pnl:,}")
print(f"Average Change: {pnl_average:,}")
print(f"Greatest Increase in Profits: ({max_month}) {max_pnl_change}")
print(f"Greatest Decrease in Profits: ({min_month}) {min_pnl_change}")
print_financial_result()
# +
# Generate print output in a text file
f = open("output.txt", "w")
print("Financial Analysis", file=f)
print("-------------------------------------", file=f)
print(f"Total Months: {count_pnl:,}", file=f)
print(f"Total: {total_pnl:,}", file=f)
print(f"Average Change: {pnl_average:,}", file=f)
print(f"Greatest Increase in Profits: {return_key(max_pnl)} > {max_pnl_change:,}", file=f)
print(f"Greatest Decrease in Profits: {return_key(min_pnl)} > {min_pnl_change:,}", file=f)
f.close()
# -
| PyBank/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## PYTHON INTERMEDIO - LECCIÓN NRO. 1 (SEGUNDO BIMESTRE)
#
# - Nombre y Apellido: <NAME>
# - Fecha: 2021/01/12
#
# ## Problema 1
#
# Empleando expresiones regulares escriba un programa que devuelva todas las fechas(dia-mes-año) del siguiente texto: 'Amit 34-3456 12-05-2007, XYZ 56-4532 11-11-2011, ABC 67-8945 12-01-2009'. De las fechas obtenidas utilice compresion de listas para imprimir solo el valor del día.
#
# Resultado: ['12', '11', '12']
#
#
#
# +
import re
text = 'Amit 34-3456 12-05-2007, XYZ 56-4532 11-11-2011, ABC 67-8945 12-01-2009'
date_expression = re.compile(r'\d\d-\d\d-\d\d\d\d')
dates = date_expression.findall(text)
print('Dates: ')
print(dates)
only_days = [date[:2] for date in dates]
print('\nOnly days: ')
print(only_days)
| Lecciones/SegundoBimestre/LeccionNro1/LeccionNro1-SB-V2_A.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
import numpy as np
import pandas as pd
data = pd.read_csv('./data/top_5_employment_ratio_1990-2015.csv')
data
# +
import matplotlib.pyplot as plt
% matplotlib inline
# -
plt.figure();
data.plot(x='Ano', **{'fontsize':12}).legend(loc='center left', bbox_to_anchor=(1, 0.5))
| top_5_Employment_Ratio_analisys.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# 21 June 2017 - WNixalo - Practical Deep Learning I - Lesson 6 CodeAlong
# [Notebook](https://github.com/fastai/courses/blob/master/deeplearning1/nbs/lesson6.ipynb)[Lecture](https://www.youtube.com/watch?v=ll9y1U0SoVY)
import theano
# %matplotlib inline
import sys, os
sys.path.insert(1, os.path.join('../utils'))
import utils; reload(utils)
from utils import *
from __future__ import division, print_function
# ## Setup
#
# We're going to download the collected works of Nietzsche to use as our data for this class.
path = get_file('nietzsche.txt', origin="https://s3.amazonaws.com/text-datasets/nietzsche.txt")
text = open(path).read()
print('corpus length:', len(text))
chars = sorted(list(set(text)))
vocab_size = len(chars) + 1
print('total chars:', vocab_size)
# Sometimes it's useful to have a zero value in the dataset, eg. for padding
chars.insert(0, "\0")
''.join(chars[1:-6])
# Map from chars to indices and back again
char_indices = dict((c, i) for i, c in enumerate(chars))
indices_char = dict((i, c) for i, c in enumerate(chars))
# *idx* will be the data we use from now on -- it simply converts all the characters to their index (based on the mapping above)
idx = [char_indices[c] for c in text]
# the 1st 10 characters:
idx[:10]
''.join(indices_char[i] for i in idx[:70])
# ## 3 Char Model
#
# ### Create Inputs
#
# Create a list of every 4th character, starting at the 0th, 1st, 2nd, then 3rd characters.
#
# *We're going to build a model that attempts to predict the 4th character from the previous 3. To do that we're going ot go through our whole list of indexes from 0 to the end minus 3, and we'll create a whole list of the 0th, 4th, 8th, 12th, etc characters; the 1st, 5th, 9th, etc; and 2nd, 6th, 10th, & so forth..*
cs = 3
c1_dat = [idx[i] for i in xrange(0, len(idx)-1-cs, cs)]
c2_dat = [idx[i+1] for i in xrange(0, len(idx)-1-cs, cs)]
c3_dat = [idx[i+2] for i in xrange(0, len(idx)-1-cs, cs)]
c4_dat = [idx[i+3] for i in xrange(0, len(idx)-1-cs, cs)] # <-- gonna predict this
# Our inputs
# we can turn these into Numpy arrays just by stacking them up together
x1 = np.stack(c1_dat[:-2]) # 1st chars
x2 = np.stack(c2_dat[:-2]) # 2nd chars
x3 = np.stack(c3_dat[:-2]) # 3rd chars
# for every 4 character peice of this - collected works
# Our output
# labels will just be the 4th characters
y = np.stack(c4_dat[:-2])
# The first 4 inputs and ouputs
# 1st, 2nd, 3rd chars of text
x1[:4], x2[:4], x3[:4]
# 4th char of text
y[:3]
# Will try to predict `30` from `40, 42, 29`, `29` from `30, 25, 27`, & etc. That's our data format.
x1.shape, y.shape
# The number of latent factors to create (ie. the size of our 3 character inputs)
# we're going to turn these into embeddings
n_fac = 42
# by creating an embedding matrix
def embedding_input(name, n_in, n_out):
inp = Input(shape=(1,), dtype='int64', name=name)
emb = Embedding(n_in, n_out, input_length=1)(inp)
return inp, Flatten()(emb)
c1_in, c1 = embedding_input('c1', vocab_size, n_fac)
c2_in, c2 = embedding_input('c2', vocab_size, n_fac)
c3_in, c3 = embedding_input('c3', vocab_size, n_fac)
# c1, c2, c3 represent result of putting each char through the embedding &
# getting out 42 latent vectors. <-- those are input to greenarrow.
# ### Create and train model
#
# Pick a size for our hidden state
n_hidden = 256
# This is the 'green arrow' from our diagram - the layer operation from input to hidden.
dense_in = Dense(n_hidden, activation='relu')
# Our first hidden activation is simpmly this function applied to the result of the embedding of the first character.
c1_hidden = dense_in(c1)
# This is the 'orange arrow' from our diagram - the layer operation from hidden to hidden.
dense_hidden = Dense(n_hidden, activation='tanh')
# Our second and third hidden activations sum up the previous hidden state (after applying dense_hidden) to the new input state.
c2_dense = dense_in(c2) # char-2 embedding thru greenarrow
hidden_2 = dense_hidden(c1_hidden) # output of char-1's hidden state thru orangearrow
c2_hidden = merge([c2_dense, hidden_2]) # merge the two together (default: sum)
c3_dense = dense_in(c3)
hidden_3 = dense_hidden(c2_hidden)
c3_hidden = merge([c3_dense, hidden_3])
# This is the 'blue arrow' from our diagram - the layer operation from hidden to hidden.
dense_out = Dense(vocab_size, activation='softmax') #output size: 86 <-- vocab_size
# The third hidden state is the input to our output layer.
c4_out = dense_out(c3_hidden)
# passing in our 3 inputs & 1 output
model = Model([c1_in, c2_in, c3_in], c4_out)
model.summary()
model.compile(loss='sparse_categorical_crossentropy', optimizer=Adam())
model.optimizer.lr=0.001
model.fit([x1, x2, x3], y, batch_size=64, nb_epoch=10)
# ### Test model
#
# We test it by creating a function that we pass 3 letters into. Turn those letters into character indices (by looking them up in char_indices); turn each of those into a Numpy array; call model.predict on those 3 arrays -- that gives us 86 outputs; We then do an argmax to find which index of those 86 is the highest: and that's the character number we want to return.
#
# So basically: we give it 3 letters, it gives us back the letter it thinks is most likely next.
def get_next(inp):
idxs = [char_indices[c] for c in inp]
arrs = [np.array(i)[np.newaxis] for i in idxs]
p = model.predict(arrs)
i = np.argmax(p)
return chars[i]
get_next('phi')
get_next(' th')
get_next(' an')
# ## Our first RNN:
#
# ### Create inputs
#
# This is the size of our unrolled RNN.
cs = 8 # use 8 characters to predict the 9th
# For each 0 thru 7, create a list of every 8th character with that starting point. These will be the 8 inputs to our model.
c_in_dat = [[idx[i+n] for i in xrange(0, len(idx)-1-cs, cs)] for n in range(cs)]
# ^ create an array with 8 elements; ea. elem contains a list of the 0th,8th,16th,24th char, the 1st,9th,17th,25th char, etc just as before. A sequence of inputs where ea. one is offset by 1 from the previous one.
#
# Then create a list of the next character in each of these series. This will be the labels for our model. -- *so our output will be exactly the same thing, except we're going to look at the indexed across by cs, so: 8. So this'll be the 8th thing in each sequence, predicted by the previous ones.*
c_out_dat = [idx[i+cs] for i in xrange(0, len(idx)-1-cs,cs)]
# go thru every one of those input lists and turn into Numpy array:
xs = [np.stack(c[:-2]) for c in c_in_dat]
len(xs), xs[0].shape
y = np.stack(c_out_dat[:-2])
# So each column below is one series of 8 characters from the text.
# visualizing xs:
[xs[n][:cs] for n in range(cs)]
# The first column in each row is the 1st 8 characters of our test.
#
# ...and this is the next character after each sequence:
y[:cs]
# NOTE: it's almost the same as the 1-7th characters in the first row of xs. The final character in ea. sequence is the same as the first character of this sequence. It's almost the same as our previous data, just done in a more flexible way.
# ## Create and train Model
n_fac = 42
def embedding_input(name, n_in, n_out):
inp = Input(shape=(1,), dtype='int64', name=name+'_in')
emb = Embedding(n_in, n_out, input_length=1, name=name+'_emb')(inp)
return inp, Flatten()(emb)
c_ins = [embedding_input('c'+str(n), vocab_size, n_fac) for n in range(cs)]
n_hidden = 256
dense_in = Dense(n_hidden, activation='relu')
dense_hidden = Dense(n_hidden, activation='relu', init='identity')
dense_out = Dense(vocab_size, activation='softmax')
# The first character of each sequence goes through ```dense_in()```, to create our first hidden activations.
hidden = dense_in(c_ins[0][1])
# Then for each successive layer we combine the output of `dense_in()` on the ext character with the output of `dense_hidden()` on the current hidden state, to create the new hidden state.
for i in range(1,cs):
c_dense = dense_in(c_ins[i][1]) #green arrow
hidden = dense_hidden(hidden) #orange arrow
hidden = merge([c_dense, hidden]) #merge the two together
# Putting the final hidden state through `dense_out()` gives us our output:
c_out = dense_out(hidden)
# So now we can create our model
model = Model([c[0] for c in c_ins], c_out)
model.compile(loss='sparse_categorical_crossentropy', optimizer=Adam())
model.fit(xs, y, batch_size=64, nb_epoch=12)
# With 8 pieces of context instead of 3, we'd expect it to do better; and we see a loss of ~1.8 instead of ~2.0
#
# ## Test model:
def get_next(inp):
idxs = [np.array(char_indices[c])[np.newaxis] for c in inp]
p = model.predict(idxs)
return chars[np.argmax(p)]
get_next('for thos')
get_next('part of ')
get_next('queens a')
# ## Returning Sequences
#
# ### Create Inputs
#
# Here, `c_out_dat` is identical to `c_in_dat`, but moved across 1 character.
#
# *So now, in ea sequece, the 1st char will be used to predict the 2nd, the 1st & 2nd to predict the 3rd, and so on. A lot more predictions going on --> a lot more opportunity for the model to learn.*
# c_in_dat = [[idx[i+n] for i in xrange(0, len(idx)-1-cs, cs)] for n in range(cs)]
c_out_dat = [[idx[i+n] for i in xrange(1, len(idx)-cs, cs)] for n in range(cs)]
ys = [np.stack(c[:-2]) for c in c_out_dat]
[xs[n][:cs] for n in range(cs)]
[ys[n][:cs] for n in range(cs)]
# Now our y dataset looks exactly like our x dataset did before, but everything's shifted over by 1 character.
#
# ### Create and train the model:
dense_in = Dense(n_hidden, activation='relu')
dense_out = Dense(vocab_size, activation='softmax', name='output')
# We're going to pass a vector of all zeros as our starting point - here's our input layers for that:
# our char1 input is moved within the diagram's loop-box; so now need
# initialized input (zeros)
inp1 = Input(shape=(n_fac,), name='zeros')
hidden = dense_in(inp1)
# +
outs = []
for i in range(cs):
c_dense = dense_in(c_ins[i][1])
hidden = dense_hidden(hidden)
hidden = merge([c_dense, hidden], mode='sum')
# every layer now has an output
outs.append(dense_out(hidden))
# our loop is identical to before, except at the end of every loop,
# we're going to append this output; so now we're going to have
# 8 outputs for every sequence instead of just 1.
# -
# model now has vector of 0s: [inp1], and array of outputs: outs
model = Model([inp1] + [c[0] for c in c_ins], outs)
model.compile(loss='sparse_categorical_crossentropy', optimizer=Adam())
zeros = np.tile(np.zeros(n_fac), (len(xs[0]),1))
zeros.shape
# Now when we fit, we add the array of zeros to the start of our inputs; our ouputs are going to be those lists of 8, offset by 1. We get 8 losses instead of 1 bc ea. one of those 8 outputs has its own loss. You'll see the model's ability to predict the 1st character from a bunch of zeros is very limited and flattens out; but predicting the 8th char with the context of 7 is much better and keeps improving.
model.fit([zeros]+xs, ys, batch_size=64, nb_epoch=12)
# This is what a sequence model looks like. We pass in a sequence and after every character, it returns a guess.
#
# ## Test Model:
def get_nexts(inp):
idxs = [char_indices[c] for c in inp]
arrs = [np.array(i)[np.newaxis] for i in idxs]
p = model.predict([np.zeros(n_fac)[np.newaxis,:]] + arrs)
print(list(inp))
return [chars[np.argmax(o)] for o in p]
get_nexts(' this is')
get_nexts(' part of')
# ## Sequence model with Keras
#
# `return_sequences=True` says: rather than put the triangle outside the loop, put it inside the recurrent loop; ie: return an output every time you go to another time-step, intead of just a single output at the end.
n_hidden, n_fac, cs, vocab_size
# To convert our previous ekras model into a sequence model, simply add the `return_sequences=True` parameter, and `TimeDistributed()` around our dense layer.
model = Sequential([
Embedding(vocab_size, n_fac, input_length=cs),
SimpleRNN(n_hidden, return_sequences=True, activation='relu', inner_init='identity'),
TimeDistributed(Dense(vocab_size, activation='softmax')),
])
model.summary()
# Note 8 outputs. What `TimeDistributed` does is create 8 copies of the weight matrix for each output.
#
# NOTE: in Keras anytime you specfy `return_sequences=True`, any dense layers after that must have `TimeDistributed` wrapped around them. - Bc, in this case, we want to create not 1 dense layer, but 8.
model.compile(loss='sparse_categorical_crossentropy', optimizer=Adam())
# just some dimensionality changes required; otherwise same
x_rnn = np.stack(np.squeeze(xs), axis=1)
y_rnn = np.stack(ys, axis=1)
x_rnn.shape, y_rnn.shape
model.fit(x_rnn, y_rnn, batch_size=64, nb_epoch=8)
def get_nexts_keras(inp):
idxs = [char_indices[c] for c in inp]
arr = np.array(idxs)[np.newaxis,:]
p = model.predict(arr)[0]
print(list(inp))
return [chars[np.argmax(o)] for o in p]
get_nexts_keras(' this is')
# ## Stateful model with Keras
#
# A stateful model is easy to create (just add `stateful=True`) but harder to train. We had to add batchnorm and use LSTM to get reasonable results.
#
# When using stateful in Keras, you have to also add `batch_input_shape` to the first layer, and fix the batch size there.
#
# ----------
# Need `shuffle=False` and `stateful=True` in order to have memory for LSTMs. Having stateful True, tells Keras not to not to reset the hidden activations to zero, but leave them as they are -- allowing the model to build up as much state as it wants. If this done, then shuffle must be False, so it'll pass in the 1st 8 chars, then 2nd 8, and so on, in order, leaving the hidden state untouched in between each one.
#
# Training these stateful models is a lot harder than other models due to exploding gradients (exploding activations). These Long-term Dependency Models were thought impossible until the '90s when researchers invented the LSTM model.
#
# In the LSTM model, the recurrent weight-activations-matrix loop is replaced with a loop with a Neural Network inside of it that decides how much of this state matrix to keep, and to use at each activation. Therefore the model can learn how to avoid gradient explosions. It can actually learn how to create an effective sequence.
#
# Below an LSTM & BatchNormed inputs are used bc J.H. had no luck with pure RNNs and ReLUs.
bs = 64
model = Sequential([
Embedding(vocab_size, n_fac, input_length=cs, batch_input_shape=(bs,8)),
BatchNormalization(),
LSTM(n_hidden, return_sequences=True, stateful=True),
TimeDistributed(Dense(vocab_size, activation='softmax')),
])
# dont forget to compile (accidetnly hit `M` in JNB)
model.compile(loss='sparse_categorical_crossentropy', optimizer=Adam())
# Since we're using a fixed batch shape, we have to ensure our inputs and outputs are an even multiple of the batch size.
mx = len(x_rnn)//bs*bs
# The LSTM model takes much longer to run than the regular RNN because it isn't in parallel: each operation has to be run in order.
model.fit(x_rnn[:mx], y_rnn[:mx], batch_size=bs, nb_epoch=4, shuffle=False)
model.optimizer.lr=1e-4
model.fit(x_rnn[:mx], y_rnn[:mx], batch_size=bs, nb_epoch=4, shuffle=False)
# ## One-Hot Sequence Model with Keras
#
# Necessary to use onehot encoding in order to build an RNN straight in Theano, next up.
#
# This is the keras version of the theano model we're about to create.
model = Sequential([
SimpleRNN(n_hidden, return_sequences=True, input_shape=(cs, vocab_size),
activation='relu', inner_init='identity'),
TimeDistributed(Dense(vocab_size, activation='softmax')),
])
model.compile(loss='categorical_crossentropy', optimizer=Adam())
# no embedding layer, so inputs must be onhotted too.
# +
oh_ys = [to_categorical(o, vocab_size) for o in ys]
oh_y_rnn = np.stack(oh_ys, axis=1)
oh_xs = [to_categorical(o, vocab_size) for o in xs]
oh_x_rnn = np.stack(oh_xs, axis=1)
oh_x_rnn.shape, oh_y_rnn.shape
# -
# The `86` is the onehotted dimension; classes of characters
model.fit(oh_x_rnn, oh_y_rnn, batch_size=64, nb_epoch=8)
def get_nexts_oh(inp):
idxs = np.array([char_indices[c] for c in inp])
arr = to_categorical(idxs, vocab_size)
p = model.predict(arr[np.newaxis,:])[0]
print(list(inp))
return [chars[np.argmax(o)] for o in p]
get_nexts_oh(' this is')
# ## Theano RNN
#
# *Sometimes you just need more control over an artificial mind.*
n_input = vocab_size
n_output = vocab_size
# Using raw theano, we have to create our weight matrices and bias vectors outselves - here are the functions we'll use to do so (using glorot initialization).
#
# The return values are wrapped in `shard()`, which is how we tell theano that it can manage this data (copying it to and from the GPU as necessary).
def init_wgts(rows, cols):
scale = math.sqrt(2/rows) # 1st calc Glorot number to scale weights
return shared(normal(scale=scale, size=(rows, cols)).astype(np.float32))
def init_bias(rows):
return shared(np.zeros(rows, dtype=np.float32))
# We return the weights and biases together as a tuple. For the hidden weights, we'll use an identity initialization (as recommended by [Hinton](https://arxiv.org/abs/1504.00941).)
def wgts_and_bias(n_in, n_out):
return init_wgts(n_in, n_out), init_bias(n_out)
def id_and_bias(n):
return shared(np.eye(n, dtype=np.float32)), init_bias(n)
# Different than Python; Theano requires us to build up a computation graph first. `shared(..)` basically tells Theano to keep track of something to send to the GPU later. Once you wrap smth in `shared` it basically belongs to Theano now.
#
# ----
# Theano doesn't actually do any computations until we explicitly compile and evaluate the function (at which point it'll be turned into CUDA code and sent off to the GPU). So our job is to describe the computations that we'll want theano to do - the first step is to tell theano what inputs we'll be providing to our computation:
# +
# Theano variables
t_inp = T.matrix('inp')
t_outp = T.matrix('outp')
t_h0 = T.vector('h0')
lr = T.scalar('lr')
all_args = [t_h0, t_inp, t_outp, lr]
# -
# Now we're ready to create our initial weight matrices.
W_h = id_and_bias(n_hidden)
W_x = wgts_and_bias(n_input, n_hidden)
W_y = wgts_and_bias(n_hidden, n_output)
w_all = list(chain.from_iterable([W_h, W_x, W_y]))
# We now need to tell Theano what happens each time we take a single step of this RNN.
#
# ----
# Theano handles looping by using the [GPU scan](http://http.developer.nvidia.com/GPUGems3/gpugems3_ch39.html) operation. We have to tell theano what to do at each step through the scan - this is the function we'll use, which does a single forward pass for one character.
def step(x, h, W_h, b_h, W_x, b_x, W_y, b_y):
# Calculate the hidden activations
h = nnet.relu(T.dot(x, W_x) + b_x + T.dot(h, W_h) + b_h)
# Calculate the output activations
y = nnet.softmax(T.dot(h, W_y) + b_y)
# Return both (the 'Flatten()' is to work around a theano bug)
return h, T.flatten(y, 1)
# Now we can provide everything necessary for the scan operation, so we can set that up - we have to pass in the function to call at each step, the sequence to step through, the initial values of the outputs, and any other arguments to pass to the step function.
[v_h, v_y], _ = theano.scan(step, sequences=t_inp,
outputs_info=[t_h0, None], non_sequences=w_all)
# You get this error if you accidently define `step` as:
#
# ```
# def step(x, h, W_h, W_x, b_h, b_x, W_y, b_y):
# ```
[v_h, v_y], _ = theano.scan(step, sequences=t_inp,
outputs_info=[t_h0, None], non_sequences=w_all)
# We can now calculate our loss function, and *all* of our gradients, with just a couple lines of code!
error = nnet.categorical_crossentropy(v_y, t_outp).sum()
g_all = T.grad(error, w_all)
# We even have to show Theano how to do SGD - so we set up this dictionary of updates to complete after every forward pass, which apply the standard SGD update rule to every weight.
def upd_dict(wgts, grads, lr):
return OrderedDict({w: w-g*lr for (w,g) in zip(wgts,grads)})
# +
upd = upd_dict(w_all, g_all, lr)
# we're finally ready to compile the function!:
fn = theano.function(all_args, error, updates=upd, allow_input_downcast=True)
# -
X = oh_x_rnn
Y = oh_y_rnn
X.shape, Y.shape
# To use it, we simply loop through our input data, calling the function compiled above, and printing our progress from time to time.
#
# ---
# We have to manually define our loop because Theano doesn't have it built-in.
err=0.0; l_rate=0.01
for i in xrange(len(X)):
err += fn(np.zeros(n_hidden), X[i], Y[i], l_rate)
if i % 1000 == 999:
print ("Error:{:.3f}".format(err/1000))
err=0.0
f_y = theano.function([t_h0, t_inp], v_y, allow_input_downcast=True)
pred = np.argmax(f_y(np.zeros(n_hidden), X[6]), axis=1)
act = np.argmax(X[6], axis=1)
[indices_char[o] for o in act]
[indices_char[o] for o in pred]
# looking at how to use Python debugger
import numpy as np
import pdb
err=0.; lrate=0.01
for i in range(len(np.zeros(10))):
err += np.sin(lrate+np.e**i)
pdb.set_trace()
| FAI_old/Lesson6/Lesson6_CodeAlong.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/anabelcarol/Machine-Learning/blob/master/Projects/PyTorch_Keras/Classification/Keras_Classification.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="GFTgoxhb26C4" colab_type="text"
# # PyTorch_Keras/Classification setting:
# Main focus on how to identify and deal with overfitting through __Early Stopping Callbacks__ and __Dropout Layers__
#
# ** __Early Stopping__ Callbacks: keras automatically stop training based on a __loss__ condition on the validation data passed during the model.fit() call.
#
# ** __Dropout Layers__: are layers that can be added to turn off neurons during training to prevent overfitting. Each dropout layer will 'drop' a user-defined percentage of neuron units in the previous layer every batch.
#
# + [markdown] id="V2yov6RR22HW" colab_type="text"
# ## Keras TF 2.0 - Code Along Classification Project
#
# Let's explore a classification task with Keras API for TF 2.0
#
# ### The Data
#
# #### Breast cancer wisconsin (diagnostic) dataset
# --------------------------------------------
#
# **Data Set Characteristics:**
#
# :Number of Instances: 569
#
# :Number of Attributes: 30 numeric, predictive attributes and the class
#
# :Attribute Information:
# - radius (mean of distances from center to points on the perimeter)
# - texture (standard deviation of gray-scale values)
# - perimeter
# - area
# - smoothness (local variation in radius lengths)
# - compactness (perimeter^2 / area - 1.0)
# - concavity (severity of concave portions of the contour)
# - concave points (number of concave portions of the contour)
# - symmetry
# - fractal dimension ("coastline approximation" - 1)
#
# The mean, standard error, and "worst" or largest (mean of the three
# largest values) of these features were computed for each image,
# resulting in 30 features. For instance, field 3 is Mean Radius, field
# 13 is Radius SE, field 23 is Worst Radius.
#
# - class:
# - WDBC-Malignant
# - WDBC-Benign
#
# :Summary Statistics:
#
# ===================================== ====== ======
# Min Max
# ===================================== ====== ======
# radius (mean): 6.981 28.11
# texture (mean): 9.71 39.28
# perimeter (mean): 43.79 188.5
# area (mean): 143.5 2501.0
# smoothness (mean): 0.053 0.163
# compactness (mean): 0.019 0.345
# concavity (mean): 0.0 0.427
# concave points (mean): 0.0 0.201
# symmetry (mean): 0.106 0.304
# fractal dimension (mean): 0.05 0.097
# radius (standard error): 0.112 2.873
# texture (standard error): 0.36 4.885
# perimeter (standard error): 0.757 21.98
# area (standard error): 6.802 542.2
# smoothness (standard error): 0.002 0.031
# compactness (standard error): 0.002 0.135
# concavity (standard error): 0.0 0.396
# concave points (standard error): 0.0 0.053
# symmetry (standard error): 0.008 0.079
# fractal dimension (standard error): 0.001 0.03
# radius (worst): 7.93 36.04
# texture (worst): 12.02 49.54
# perimeter (worst): 50.41 251.2
# area (worst): 185.2 4254.0
# smoothness (worst): 0.071 0.223
# compactness (worst): 0.027 1.058
# concavity (worst): 0.0 1.252
# concave points (worst): 0.0 0.291
# symmetry (worst): 0.156 0.664
# fractal dimension (worst): 0.055 0.208
# ===================================== ====== ======
#
# :Missing Attribute Values: None
#
# :Class Distribution: 212 - Malignant, 357 - Benign
#
# :Creator: Dr. <NAME>, <NAME>, <NAME>
#
# :Donor: <NAME>
#
# :Date: November, 1995
#
# This is a copy of UCI ML Breast Cancer Wisconsin (Diagnostic) datasets.
# https://goo.gl/U2Uwz2
#
# Features are computed from a digitized image of a fine needle
# aspirate (FNA) of a breast mass. They describe
# characteristics of the cell nuclei present in the image.
#
# Separating plane described above was obtained using
# Multisurface Method-Tree (MSM-T) [<NAME>, "Decision Tree
# Construction Via Linear Programming." Proceedings of the 4th
# Midwest Artificial Intelligence and Cognitive Science Society,
# pp. 97-101, 1992], a classification method which uses linear
# programming to construct a decision tree. Relevant features
# were selected using an exhaustive search in the space of 1-4
# features and 1-3 separating planes.
#
# The actual linear program used to obtain the separating plane
# in the 3-dimensional space is that described in:
# [<NAME> and <NAME>: "Robust Linear
# Programming Discrimination of Two Linearly Inseparable Sets",
# Optimization Methods and Software 1, 1992, 23-34].
#
# This database is also available through the UW CS ftp server:
#
# ftp ftp.cs.wisc.edu
# # # cd math-prog/cpo-dataset/machine-learn/WDBC/
#
# .. topic:: References
#
# - W.<NAME>, <NAME> and <NAME>. Nuclear feature extraction
# for breast tumor diagnosis. IS&T/SPIE 1993 International Symposium on
# Electronic Imaging: Science and Technology, volume 1905, pages 861-870,
# San Jose, CA, 1993.
# - <NAME>, W.N. Street and <NAME>. Breast cancer diagnosis and
# prognosis via linear programming. Operations Research, 43(4), pages 570-577,
# July-August 1995.
# - <NAME>, W.N. Street, and <NAME>ian. Machine learning techniques
# to diagnose breast cancer from fine-needle aspirates. Cancer Letters 77 (1994)
# 163-171.
# + id="TkPwbq0h22HZ" colab_type="code" colab={}
import pandas as pd
import numpy as np
# + id="7LyFlEAZ5NwQ" colab_type="code" outputId="87bec84a-7456-4c3e-c60f-58114bcd62da" colab={"base_uri": "https://localhost:8080/", "height": 71}
import matplotlib.pyplot as plt
import seaborn as sns
# + id="SY5KNrYx5eTV" colab_type="code" outputId="b2359db4-33e8-4036-9cc3-bb59cfd901bc" colab={"base_uri": "https://localhost:8080/", "height": 122}
from google.colab import drive
drive.mount('/content/drive')
# + id="nDDyRtjH22Hg" colab_type="code" colab={}
df = pd.read_csv('/content/drive/My Drive/Colab Notebooks/DeepLearning_TensorFlow/TensorFlow_FILES/DATA/cancer_classification.csv')
# + id="6jopDMP322Hm" colab_type="code" outputId="4fd61830-3c29-47b1-8c22-5cfd712ea2ba" colab={"base_uri": "https://localhost:8080/", "height": 663}
df.info()
# + id="Ok-FsOLU22Hq" colab_type="code" outputId="0799a347-0d6d-4fc1-fc1b-92e620aacccd" colab={"base_uri": "https://localhost:8080/", "height": 1000}
df.describe().transpose()
# + [markdown] id="zhYdag4H22Ht" colab_type="text"
# ## EDA
# + [markdown] id="HXTyAGTv63_2" colab_type="text"
# For classification settings, it is usually useful to plot the target variable to check if the dataset is balanced or not
# + id="vgHGDzmW22Hx" colab_type="code" outputId="6bb8cdba-6e19-4ed5-d452-740bc9b390b0" colab={"base_uri": "https://localhost:8080/", "height": 297}
sns.countplot(x='benign_0__mal_1',data=df)
# + id="u2RQqZK_7N4n" colab_type="code" outputId="34bc886e-ea9e-4e23-e611-438bf02919b7" colab={"base_uri": "https://localhost:8080/", "height": 391}
# correlations with respect to the target variable 'benign or maling'
df.corr()['benign_0__mal_1'][:-1].sort_values().plot(kind='bar')
# + id="hEipnUh_22H1" colab_type="code" outputId="f41481ed-1351-40da-d497-b405c4e07c1e" colab={"base_uri": "https://localhost:8080/", "height": 511}
plt.figure(figsize=(12,8))
sns.heatmap(df.corr())
# + [markdown] id="gVHkftrX22IB" colab_type="text"
# ## Train Test Split
# + id="sf8Vb2wv22IB" colab_type="code" colab={}
X = df.drop('benign_0__mal_1',axis=1).values
y = df['benign_0__mal_1'].values
# + id="2-Wah0EJ22IG" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
# + id="wYWNXXcE22II" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.25,random_state=101)
# + [markdown] id="yPigcnNq22IL" colab_type="text"
#
# ## Scaling Data
# + id="dW5ltSYB22IM" colab_type="code" colab={}
from sklearn.preprocessing import MinMaxScaler
# + id="JDHBpBMc22IN" colab_type="code" colab={}
scaler = MinMaxScaler()
# + id="J_yeQyU322IP" colab_type="code" outputId="38091111-16da-439d-d6e5-08c273bdc7d1" colab={"base_uri": "https://localhost:8080/", "height": 34}
scaler.fit(X_train)
# + id="XUbo78nK22IR" colab_type="code" colab={}
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
# + [markdown] id="3eqOavrN22IS" colab_type="text"
# ## Creating the Model
#
# # For a binary classification problem
# model.compile(optimizer='rmsprop',
# loss='binary_crossentropy',
# metrics=['accuracy'])
#
#
# + id="PfvI5NYm22IS" colab_type="code" colab={}
# import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation,Dropout
# + id="8PSUG-rW22IU" colab_type="code" outputId="d99da4c6-5d84-495b-e6cc-838df374ca88" colab={"base_uri": "https://localhost:8080/", "height": 34}
X_train.shape
# + id="anNgRRMo22IW" colab_type="code" colab={}
model = Sequential()
# https://stats.stackexchange.com/questions/181/how-to-choose-the-number-of-hidden-layers-and-nodes-in-a-feedforward-neural-netw
model.add(Dense(units=30,activation='relu'))
model.add(Dense(units=15,activation='relu'))
# for binary_classification problem
model.add(Dense(units=1,activation='sigmoid'))
# For a binary classification problem
model.compile(loss='binary_crossentropy', optimizer='adam')
# + [markdown] id="gBoRCEl722IY" colab_type="text"
# ## Training the Model
#
# ### Example One: Choosing too many epochs and overfitting!
# + id="87xzqpSB22IY" colab_type="code" outputId="b06f158a-d0c8-4205-e525-17a10b4c6ca5" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# https://stats.stackexchange.com/questions/164876/tradeoff-batch-size-vs-number-of-iterations-to-train-a-neural-network
# https://datascience.stackexchange.com/questions/18414/are-there-any-rules-for-choosing-the-size-of-a-mini-batch
model.fit(x=X_train,
y=y_train,
epochs=600,
validation_data=(X_test, y_test), verbose=1
)
# + id="WJP_bNZU22Ia" colab_type="code" colab={}
# model.history.history
# + id="ueDv31jq22Ib" colab_type="code" colab={}
losses = pd.DataFrame(model.history.history)
# + id="D_2YkC_U22Ie" colab_type="code" outputId="2383c32a-f0c9-452d-a1b8-5992a0a0547f" colab={"base_uri": "https://localhost:8080/", "height": 282}
losses.plot()
# + [markdown] id="FLevAUMG-Jpg" colab_type="text"
# _This is a clear example of overfitting_
# + [markdown] id="MlkvgXAb22If" colab_type="text"
# ## Example Two: Early Stopping
#
# We obviously trained too much! Let's use early stopping to track the val_loss and stop training once it begins increasing too much!
# + id="aToZOcGA22Ig" colab_type="code" colab={}
model = Sequential()
model.add(Dense(units=30,activation='relu'))
model.add(Dense(units=15,activation='relu'))
model.add(Dense(units=1,activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer='adam')
# + id="gfXas2pF22Ih" colab_type="code" colab={}
from tensorflow.keras.callbacks import EarlyStopping
# + [markdown] id="n5bJqh9922Ii" colab_type="text"
# Stop training when a monitored quantity has stopped improving.
#
# Arguments:
# monitor: Quantity to be monitored.
# min_delta: Minimum change in the monitored quantity
# to qualify as an improvement, i.e. an absolute
# change of less than min_delta, will count as no
# improvement.
# patience: Number of epochs with no improvement
# after which training will be stopped.
# verbose: verbosity mode.
# mode: One of `{"auto", "min", "max"}`. In `min` mode,
# training will stop when the quantity
# monitored has stopped decreasing; in `max`
# mode it will stop when the quantity
# monitored has stopped increasing; in `auto`
# mode, the direction is automatically inferred
# from the name of the monitored quantity.
# + id="xq4At2O522Ii" colab_type="code" colab={}
early_stop = EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=25)
# verbose = 1 --> just a small report
# patience --> we wait 25 epochs even after we have kind of detecting a stoping point because of noise that could occur
# + id="szmTzPGB22Ij" colab_type="code" outputId="8f0c03ba-4bae-4e2e-ea11-8a9a80551d7a" colab={"base_uri": "https://localhost:8080/", "height": 1000}
model.fit(x=X_train,
y=y_train,
epochs=600,
validation_data=(X_test, y_test), verbose=1,
callbacks=[early_stop]
)
# it is going to attempt to run on 600 epochs unless the early stop is triggered.
# in this case it is triggered. the training stopped
# + id="bPUUfQwJ22Ik" colab_type="code" outputId="888dd60e-7f60-4cf8-f046-c6c36894910a" colab={"base_uri": "https://localhost:8080/", "height": 282}
# the fit stopped at 97 epochs.
model_loss = pd.DataFrame(model.history.history)
model_loss.plot()
# + [markdown] id="f7jc8KHOAsCJ" colab_type="text"
# flattening is ok for val_loss
# + [markdown] id="D4vLqw6522Im" colab_type="text"
# ## Example Three: Adding in DropOut Layers
# + id="hegGRuk122Im" colab_type="code" colab={}
from tensorflow.keras.layers import Dropout
# + id="wGyxEyH322In" colab_type="code" colab={}
model = Sequential()
model.add(Dense(units=30,activation='relu'))
model.add(Dropout(0.5)) # rate --> the portion of neurons that we turn off
# rate = 1 --> 100% of neurons will be turn off
# rate = 0 --> 0% neurons will be turn off
model.add(Dense(units=15,activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(units=1,activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer='adam')
# + id="J1fQMsGd22Ip" colab_type="code" outputId="7c3970d8-53da-4e0f-b1f2-90a822c373cb" colab={"base_uri": "https://localhost:8080/", "height": 1000}
model.fit(x=X_train,
y=y_train,
epochs=600,
validation_data=(X_test, y_test), verbose=1,
callbacks=[early_stop]
)
# + id="2-BDH2Nx22Iq" colab_type="code" outputId="a2f7e427-3909-49a5-9ddd-fa81218e2fb8" colab={"base_uri": "https://localhost:8080/", "height": 282}
# the model stopped on the 131 epochs that is better because it continued to learn
model_loss = pd.DataFrame(model.history.history)
model_loss.plot()
# + [markdown] id="vGTKoj-DBy1-" colab_type="text"
# Train_Loss and Val_loss both are flatenning at the same time
# + [markdown] id="rIA-etU022Ir" colab_type="text"
# # Model Evaluation
# + [markdown] id="n8h5C6_kB9Fp" colab_type="text"
# for classsification setting!!!!!
# + id="0nhppkUb22Is" colab_type="code" outputId="346f2a67-f0bb-4ca1-9fdc-0c0f219dcc7b" colab={"base_uri": "https://localhost:8080/", "height": 88}
predictions = model.predict_classes(X_test)
# + [markdown] id="vt-hLBiSC56a" colab_type="text"
# WARNING:tensorflow:From <ipython-input-38-bc83193b8b59>:1: Sequential.predict_classes (from tensorflow.python.keras.engine.sequential) is deprecated and will be removed after 2021-01-01.
# Instructions for updating:
# Please use instead:* `np.argmax(model.predict(x), axis=-1)`, if your model does multi-class classification (e.g. if it uses a `softmax` last-layer activation).* `(model.predict(x) > 0.5).astype("int32")`, if your model does binary classification (e.g. if it uses a `sigmoid` last-layer activation).
# + id="e9Z61mbZCrq9" colab_type="code" colab={}
# pred = model.predict((X_test) > 0.5).astype("int32")
# + id="lygJs5eo22It" colab_type="code" colab={}
from sklearn.metrics import classification_report,confusion_matrix
# + id="KacJosjj22Iu" colab_type="code" outputId="dd978345-ef5f-4db4-d4b2-2fc656a2e391" colab={"base_uri": "https://localhost:8080/", "height": 170}
# https://en.wikipedia.org/wiki/Precision_and_recall
print(classification_report(y_test,predictions))
# + id="0pZKKlvE22Iv" colab_type="code" outputId="970d5a68-7912-41ec-87f0-017351f20343" colab={"base_uri": "https://localhost:8080/", "height": 51}
print(confusion_matrix(y_test,predictions))
| Projects/TensorFlow_Keras/Classification/theory.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.3
# language: julia
# name: julia-1.5
# ---
# # Bayesian brain model of an attentional phenomenon
# <NAME>, [@inferenceLab](https://twitter.com/inferencelab)
#
# _This notebook uses the same methods as used in Vincent (2015a). It is faster to compute, but required some maths and thinking to do. Please see the other version of the attention notebook for a version which is much easier - in that we construct a forward model of the task and rely on the computer to do the hard work_
#
# This notebook moves on from the linear regression example where the model _describes_ the data, and from the discounting model which potentially _explains_ the data, to look at another explanatory model. This kind of model is often referred to as a 'Bayesian brain' model in that our model is that participants themselves are conducting Bayesian inference about the state of the world. Another way of saying this is that we are graduating from us as scientists being Bayesian, to considering whether people are Bayesian.
#
# We will not fully explore this here, but we will replicate the Bayesian optimal observer predictions made in Vincent (2011), and later discussed in a broader review of Bayesian explanations of attentional phenomena (Vincent, 2015a).
#
# ## The task
# This is a covert attentional task where participants look at a computer screen:
# - They fixate the centre of the screen so that stimuli which later appear are all equidistant from the fovea.
# - A cue appears uniformly randomly in one of $N$ locations before disappearing.
# - The participants have been informed about the validity of the cue in advance. For example, if the cue is 80% valid, then there is an 80% chance that a target object will appear at the cued location and a 20% of being in one of the other $N-1$ locations.
# - Distractors appear in all other locations.
# - The stimulus items are chosen to be visually similar and to appear for short durations of time, so that there is remaining uncertainty about the target's location.
#
# <img src="img/cued localisation.png" width="600">
#
# ## The theoretical question asked in Vincent (2011)
# The main aim was to see if participants acted in a Bayesian manner by incorporating prior information (from the cue) and the likelihood (sensory information consistent with hypotheses about the world).
#
# Vincent (2011) examined how people's performance in a cued localisation task varied depending on how valid the cue was. People's performance was worst when the cue was uninformative - in this case with 4 locations this was when the cue was valid 1/4 of the time. Performance improved when the cue was more predictive of the target location, suggesting that participants did combine their prior beliefs (from the cue) with the likelihood (from the stimuli). However, it was also the case that performance improved when the cue signalled where a target would be _less_ likely to appear. For example, when it was 0% valid the target was never at the cued location. Participants were able to use the information from the cue so as to update their beliefs that the targets were more likely to occur in the remaining locations, which lead to enhanced performance.
#
# <img src="img/vincent2011.png" width="400">
#
# There were some interesting peculiarities about the participant's performance which indicated that while they combined the cue-derived prior beliefs and the stimulus-based likelihood, they seemed to have biases in their beliefs akin to those seen in Prospect Theory. In this notebook, we will ignore this fascinating probability bias issue and focus on replicating the optimal observer predictions as shown in Figure 3 of Vincent (2011) and expanded upon in Figure 8 of Vincent (2015a). Further information about Bayesian optimal observer models of covert visual attention can be found in Vincent (2015a). Further useful background into Bayesian modelling of perception can be found in Vincent (2015b).
#
# **Note:** The original Matlab code associated with Vincent (2011) is in GitHub at [drbenvincent/BayesCovertAttention](https://github.com/drbenvincent/BayesCovertAttention). Although beware, I was a less seasoned programmer at that point, so apologies if the code is not as clear as it could be.
# ## Let's start coding
using Plots, StatsBase, Distributions
# Write a function to simulate if an observer correctly infers the true location of a target amongst $N-1$ distractors.
# +
function simulated_response(N::Int64, v::Float64, σ::Float64)
# Step 1: simulate events in a trial ======================================
# Cue has equal chance of appearing in location 1:N
cue_location = sample(1:N, Weights(fill(1/N, N)))
# Prior over cue location
# v at cued location
# (1-v)/(N-1) at uncued location
prior = [n==cue_location ? v : (1-v)/(N-1) for n in 1:N]
target_location = sample(1:N, Weights(prior))
# Noisy sensory observations
x = rand.([Normal(n==target_location, σ) for n in 1:N])
# Step 2: inference ======================================================
# Evaluate likelihood target is in location L AND that distractors are
# in remaining locations, for each of 1:N possible locations
log_likelihood = [log_likelihood_location(x, L, N, σ) for L in 1:N]
# Calculate log posterior probability over locations 1:N
log_posterior = log.(prior) + log_likelihood
# Step 3: Decision =======================================================
response = argmax(log_posterior)
correct = response==target_location ? 1 : 0
return correct
end;
# -
# Evaluate log likelihood that a target is in location $L$ of $N$ total locations. For example, if the hypothesis is that the target is in location $L=2$ of $N=4$ locations, then we sum up:
# - distractor in location 1 = `logpdf(Normal(0, σ), x[1])`
# - target in location 2 = `logpdf(Normal(1, σ), x[2])`
# - distractor in location 3 = `logpdf(Normal(0, σ), x[3])`
# - distractor in location 4 = `logpdf(Normal(0, σ), x[4])`
function log_likelihood_location(x::Array{Float64,1}, L::Int64, N::Int64, σ::Float64)
sum([logpdf(Normal(i==L ? 1 : 0, σ), x[i]) for i in 1:N])
end;
# Create a `simulate` function which runs many simulated experiments, repeated with the various cue validities we are interested in.
function simulate(N::Int64, cue_validities::LinRange{Float64}, σ::Float64, trials::Int64)
pc = zeros(length(cue_validities))
for (i, v) in enumerate(cue_validities)
pc[i] = sum([simulated_response(N, v, σ) for t in 1:trials]) / trials
end
return pc
end;
function make_figure(N, cue_validities, dprime_list, trials, subplot)
# set up plot
plot!(xlabel="cue validity, v",
ylabel="percent correct localisations",
ylim=[0, 1],
legend=:bottomright,
title="set size = $N",
subplot=subplot)
# run simulations for each of the dprime values and plot
for dprime in dprime_list
σ = 1 / dprime
pc = simulate(N, cue_validities, σ, trials)
plot!(cue_validities, pc, lw=5, label="d' = $dprime", subplot=subplot)
end
hline!([1/N], color=:black, label="chance performance", subplot=subplot)
vline!([1/N], color=:black, label="cue uninformative", linestyle=:dash, subplot=subplot)
end;
# Now let's replicate the optimal observer predictions for a set sizes of $N=2$ and $N=4$.
# +
cue_validities = LinRange(0.01, 0.99, 50)
dprime_list = [0.5, 1.0, 2.0]
trials = 100_000
plot(layout=(1, 2), size=(800, 400))
make_figure(2, cue_validities, dprime_list, trials, 1)
make_figure(4, cue_validities, dprime_list, trials, 2)
# -
# It worked! We now have pretty concise code to replicate the Bayesian optimal observer predictions in Vincent (2011), and more specifically, Figure 8 in Vincent (2015a).
#
# <img src="img/vincent_2015_figure_8.png" width="800">
# ## Going further: Estimating the $d'$ parameter based on behavioural data
# While this is very useful, the next step would be to build on this and to estimate the $d'$ of an observer based upon behavioural data. I will leave this as an exercise for the reader, but the general approach might be:
# - think about the behavioural data we would have and whether the analysis runs at the trial level or the participant level
# - think about an appropriate likelihood, which depends upon whether we are modelling trial level correct/incorrect responses, or participant level number or proportion correct data
# - use (or adapt) the existing `simulated_response()` function and embed it within a Bayesian model using Turing.jl
# - construct appropriate priors
# - sample from the posterior to estimate something like $P(d' | N, \text{response data}, \text{cue validities})$
# # References
# - <NAME>. (2011). Covert visual search: Prior beliefs are optimally combined with sensory evidence. Journal of Vision, 11(13), 25-25.
# - <NAME>. (2015a). Bayesian accounts of covert selective attention: a tutorial review. Attention, Perception, & Psychophysics, 77(4), 1013-1032.
# - <NAME>. (2015b). A tutorial on Bayesian models of perception. Journal of Mathematical Psychology, 66, 103-114.
| covert_attention_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Solution to exercise 1.3.1
#
# - Write a function that takes two arguments and returns their mean.
# - Give your function a meaningful name, and a good documentation.
# - Call your function multiple times with different values, and once using the keyword arguments with their associated values.
# - Print the result of these different function calls.
def mean_of_two_values(value1, value2):
"""
Returns the mean of the two arguments.
value1 --- first value
value2 --- second value
"""
print('calculating the mean of', value1, 'and', value2)
return (value1 + value2)/2
mean_of_variables = mean_of_two_values(2, 5)
print(mean_of_variables)
mean_of_variables = mean_of_two_values(-6, 5)
print(mean_of_variables)
mean_of_variables = mean_of_two_values('4', 5)
print(mean_of_variables)
mean_of_variables = mean_of_two_values(value2=2, value1=5)
print(mean_of_variables)
results = mean_and_median([2, 5])
print(results)
# **BEWARE** Do not use [Python built-in names](https://docs.python.org/3/library/functions.html#built-in-funcs) for your variables and functions otherwise you will change their behaviour.
| solutions/ex_1_3_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import os
raw_data_path = os.path.join(os.pardir, 'data', 'raw')
train_file_path = os.path.join(raw_data_path, "train.csv")
test_file_path = os.path.join(raw_data_path, "test.csv")
train_df = pd.read_csv(train_file_path, index_col='PassengerId')
test_df = pd.read_csv(test_file_path, index_col='PassengerId')
type(train_df)
train_df.info()
test_df.info()
test_df['Survived'] = -888
test_df.info()
df = pd.concat((train_df, test_df))
df.info()
df.head()
df.tail()
df.Survived
df.Name[3]
df.loc[5:10:2,]
df.loc[5:10:2,['Age','Sex']]
df.loc[5:10:2,'Age':'Sex']
df.iloc[5:10:2,1:3]
male_passengers = df.loc[(df.Sex=='male') & (df.Survived == 1)]
print("There are {} male passengers that survived.".format(len(male_passengers)))
type(male_passengers)
male_passengers.loc[:,['Pclass','Sex']].describe()
df.info()
df.describe()
print("Mean fare is {}".format(df.Fare.mean()))
print("Median fare is {}".format(df.Fare.median()))
print("StdDev fare is {}".format(df.Fare.std()))
print("75th percentile is {}".format(df.Fare.quantile(0.75)))
# %matplotlib inline
df.Age.plot(kind='box')
df.describe(include='all')
df.Sex.value_counts()
df.Sex.value_counts(normalize=True)
df.Survived[df.Survived != -888].value_counts(normalize=True)
df.Pclass.value_counts()
df.Pclass.value_counts().plot(kind='bar')
df.Pclass.value_counts().plot(kind='bar', rot=0, title="Class-wise passenger count");
df.Age.value_counts()
histTitle = 'Median Age is {}. Mean is {:.2f}'.format(df.Age.median(), df.Age.mean())
df.Age.plot(kind='hist', bins=20, color='r', title=histTitle)
df.Age.plot(kind='kde')
df.plot.scatter(x = 'Age', y = 'Fare', title = 'Age/Fare Correlation')
df.plot.scatter(x = 'Age', y = 'Fare', title = 'Age/Fare Correlation', alpha=0.1)
df.plot.scatter(x = 'Pclass', y = 'Fare', title = 'Passenger Class/Fare Correlation', alpha=0.15, color='c')
df.groupby('Sex').Age.mean()
df[df.Survived != -888].groupby(['Sex','Pclass']).mean()
pd.crosstab(df.Sex, df.Pclass)
pd.crosstab(df.Sex, df.Pclass).plot(kind='bar');
df.pivot_table(index='Sex', columns='Pclass', aggfunc='mean', values='Age')
df.groupby(['Sex', 'Pclass']).Age.mean()
df.groupby(['Sex', 'Pclass']).Age.mean().unstack()
| notebooks/Exploring the Titanic Dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.7 64-bit (''cryptoalgowheel'': conda)'
# name: python_defaultSpec_1598751479385
# ---
import requests
import numpy as np
import pandas as pd
from datetime import datetime
from dateutil.tz import tzutc
# #### required output columns info: <br>
# close float64 <br>
# high float64 <br>
# low float64 <br>
# open float64 <br>
# volume float64 <br>
# baseVolume float64 <br>
# datetime object <br>
# dtype: object
def histohour_main(fsym='BTC', tsym='USDT', start_time='2017-04-01', end_time='2020-04-01', e='binance'):
#create UTC Unix timestamps from start & end date strings passed in
req_start_epoch = int(datetime.strptime(start_time, '%Y-%m-%d').replace(tzinfo=tzutc()).timestamp())
req_end_epoch = int(datetime.strptime(end_time, '%Y-%m-%d').replace(tzinfo=tzutc()).timestamp())
result_df = pd.DataFrame()
next_endtime = req_end_epoch #initialize "end time"
while True:
r = requests.get("https://min-api.cryptocompare.com/data/v2/histohour?", params={'fsym':fsym, 'tsym': tsym, 'limit': 2000, 'toTs': str(next_endtime), 'e': e})
response = r.json() #the API query response actually can be decoded as a JSON (list of dicts) object!
current_df = pd.DataFrame(response['Data']['Data']).drop(['conversionType', 'conversionSymbol'], axis=1)
result_df = pd.concat([current_df, result_df], axis=0, ignore_index=True) #stack/add the current df to result df along the y-axis
current_starttime = result_df.time.min() #check the smallest value in the 'time' column of df as the start time of the current iteration
next_endtime = current_starttime - 3600 #the end time for the next round is 'one hour'(3600 Unix timestamp units) before the start time of the current iteration
if current_starttime <= req_start_epoch:
break
#result df Data Cleaning
result_df.sort_values(by="time", inplace=True) #(just in case)
result_df = result_df[result_df['time'] >= req_start_epoch] #delete superfluous data earlier than the required start datetime
result_df = result_df.rename(columns={'time':'datetime', 'volumefrom':'volume', 'volumeto':'baseVolume'})
result_df.datetime = result_df.datetime.apply(lambda d: datetime.utcfromtimestamp(d).strftime("%Y/%m/%d %H:%M")) #(*)change the UTC Unix timestamps display format to be string
result_df = result_df[["close", "high", "low", "open", "volume", "baseVolume", "datetime"]]
#write the final result dataframe to csv file
result_df.to_csv("/Users/baixiao/Desktop/S1T1_output.csv", index=False)
histohour_main()
| section1/task1/[Submission]Main_Code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ec333_env
# language: python
# name: ec333_env
# ---
# # Panel Data Models
# ---
# +
import pandas as pd
import numpy as np
import statsmodels.api as sm
from linearmodels import PooledOLS
from linearmodels import RandomEffects
from linearmodels import PanelOLS
from linearmodels import FirstDifferenceOLS
from stargazer.stargazer import Stargazer
# -
# Import crime data from North Carolina between 1981 to 1987
data = pd.read_csv("data/crime.csv", index_col=0)
data
# +
# Best practice: check dtypes
# There are two columns with object dtypes that should be further investigated
# BUT since we will not be using those columns in this notebook,
# we can safely ignore them
data.dtypes
# -
data.columns
# # 1. Random Effects, Fixed Effects, and First Differences
# ---
# ## Comments:
# - In notebook 1_introduction, I wrote the code for both non-formula and formula APIs to running regressions in statsmodels.
# - However, for simplicity and comparability with R code, I will only use the formula API from this notebook onwards.
# - The main library from this notebook onwards is linearmodels.
# - Linearmodels is a library that extends statsmodels with econometric models.
# +
# Important! You must set a multi-index (entity, time) for panel data methods in linearmodels to work
data = data.set_index(['county','year'])
data
# -
# ## 1.1. Pooled OLS
# +
# Naive OLS assuming i.i.d. data
mod_OLS = PooledOLS.from_formula("crmrte ~ 1 + density + taxpc + wcon + pctmin", data=data)
res_OLS = mod_OLS.fit()
res_OLS.summary
# -
# ## 1.2. Random Effects
mod_randeff = RandomEffects.from_formula("crmrte ~ 1 + density + taxpc + wcon + pctmin", data=data)
res_randeff = mod_randeff.fit()
res_randeff.summary
# ## 1.3. Fixed Effects
# +
# Entity fixed effects
entity_fe_formula = "crmrte ~ density + taxpc + wcon + pctmin + EntityEffects"
mod_entity_fe = PanelOLS.from_formula(entity_fe_formula, data=data, drop_absorbed=True)
res_entity_fe = mod_entity_fe.fit()
res_entity_fe.summary
# +
# Entity and time period fixed effects
twoways_fe_formula = "crmrte ~ density + taxpc + wcon + pctmin + TimeEffects + EntityEffects"
mod_twoways_fe = PanelOLS.from_formula(twoways_fe_formula, data=data, drop_absorbed=True)
res_twoways_fe = mod_twoways_fe.fit()
res_twoways_fe.summary
# -
# ## 1.4. First Differences
mod_fd = FirstDifferenceOLS.from_formula("crmrte ~ density + taxpc + wcon", data=data)
res_fd = mod_fd.fit()
res_fd.summary
# # 2. Standard Errors
# ---
# ### Note:
# - Three covariance estiminators are supported in linearmodels:
# 1. White's (1980) robust standard errors
# 2. Clustered standard errors (entity, time, two-way)
# 3. Driscoll-Kraay HAC standard errors
#
# Reference: https://bashtage.github.io/linearmodels/doc/panel/models.html
# +
# warning: this is probably bad
# White's (1980) robust standard errors
res_entity_fe_robust = mod_entity_fe.fit(cov_type="robust")
res_entity_fe_robust.summary
# +
# Clustered by entity (fixed effects)
res_fe_clustered_entity = mod_entity_fe.fit(cov_type="clustered", cluster_entity=True)
res_fe_clustered_entity.summary
# +
# Clustered by entity (first difference)
res_fd_clustered_entity = mod_fd.fit(cov_type="clustered", cluster_entity=True)
res_fd_clustered_entity.summary
# +
# Clustered by time (fixed effects)
res_fe_clustered_time = mod_entity_fe.fit(cov_type="clustered", cluster_time=True)
res_fe_clustered_time.summary
# +
# Clustered by time (first difference)
res_fd_clustered_time = mod_fd.fit(cov_type="clustered", cluster_time=True)
res_fd_clustered_time.summary
# +
# Newey-West
# WARNING: linearmodels currently does not implement Newey-West SEs
# But statsmodels does!
# The following code implements a entity-dummies fixed effects regression
# Note: you must include "-1" into the formula to remove the intercept
data_noindex = data.reset_index()
mod_fe_NW = sm.formula.ols("crmrte ~ density + taxpc + wcon + C(county) -1", data=data_noindex)
res_fe_NW = mod_fe_NW.fit(cov_type="HAC", cov_kwds={"maxlags":1})
res_fe_NW.summary()
# +
# Driscoll-Kraay HAC
res_fe_HAC = mod_entity_fe.fit(cov_type="kernel", kernel="andrews")
res_fe_HAC.summary
| notebooks/2_panel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# +
import pandas as pd
import psycopg2
import sqlalchemy
import matplotlib as plt
import getpass
from IPython.display import display
from ipywidgets import widgets, Label
# %matplotlib inline
# +
# Postgres username, password, and database name
POSTGRES_ADDRESS = widgets.Text(description="HOST:", width=1000)
display(POSTGRES_ADDRESS)
POSTGRES_PORT = widgets.Text(description="PORT:", width=1000)
display(POSTGRES_PORT)
POSTGRES_USERNAME = widgets.Text(description="USER:", width=1000)
display(POSTGRES_USERNAME)
POSTGRES_DBNAME = widgets.Text(description="DB:", width=1000)
display(POSTGRES_DBNAME)
POSTGRES_PASSWORD = widgets.Password(description="Password:", width=1000)
display(POSTGRES_PASSWORD)
# +
from sqlalchemy import create_engine
postgres_str = ('postgresql://{username}:{password}@{ipaddress}:{port}/{dbname}'
.format(username=POSTGRES_USERNAME.value,
password=<PASSWORD>.value,
ipaddress=POSTGRES_ADDRESS.value,
port=POSTGRES_PORT.value,
dbname=POSTGRES_DBNAME.value))
# Create the connection
cnx = create_engine(postgres_str)
# -
pd.read_sql_query('''select * from pg_awr_snapshots_cust order by snap_id;''', cnx)
# +
sql_dist = pd.read_sql_query('''with get_sql_id as (
select sample_start_time,a.snap_id,dbid,userid,queryid,round(avg(calls)) AS calls,round(avg(total_time)) AS total_time,round(avg(rows)) AS rows,
round(avg(shared_blks_hit)) AS shared_blks_hit,round(avg(shared_blks_read)) AS shared_blks_read, round(avg(shared_blks_dirtied)) AS shared_blks_dirtied,
round(avg(shared_blks_written)) AS shared_blks_written, round(avg(local_blks_hit)) AS local_blks_hit, round(avg(local_blks_read)) AS local_blks_read,
round(avg(local_blks_dirtied)) As local_blks_dirtied, round(avg(local_blks_written)) AS local_blks_written, round(avg(temp_blks_read)) AS temp_blks_read,
round(avg(temp_blks_written)) AS temp_blks_written, round(avg(blk_read_time)) AS blk_read_time, round(avg(blk_write_time)) AS blk_write_time
from pg_stat_statements_history a, pg_awr_snapshots_cust b where a.snap_id = b.snap_id
and a.snap_id between %(begin_snap_id)s and %(end_snap_id)s
group by sample_start_time,a.snap_id,dbid,userid,queryid
order by dbid,userid,queryid,a.snap_id
),
get_lag_data as (
select sample_start_time,dbid,userid,queryid,snap_id,calls,
case WHEN (calls-lag(calls::numeric,1,0::numeric) OVER (partition by dbid, userid, queryid ORDER BY snap_id) ) = calls then null
else (calls-lag(calls::numeric,1,0::numeric) OVER (partition by dbid, userid, queryid ORDER BY snap_id) ) END AS delta_calls,
case WHEN (total_time-lag(total_time::numeric,1,0::numeric) OVER (partition by dbid, userid, queryid ORDER BY snap_id) ) = total_time then null
else (total_time-lag(total_time::numeric,1,0::numeric) OVER (partition by dbid, userid, queryid ORDER BY snap_id) ) END as Delta_total_time,
case WHEN (rows-lag(rows::numeric,1,0::numeric) OVER (partition by dbid, userid, queryid ORDER BY snap_id) ) = rows then null
else (rows-lag(rows::numeric,1,0::numeric) OVER (partition by dbid, userid, queryid ORDER BY snap_id) ) END as Delta_rows,
case WHEN (shared_blks_hit-lag(shared_blks_hit::numeric,1,0::numeric) OVER (partition by dbid, userid, queryid ORDER BY snap_id) ) = shared_blks_hit then null
else (shared_blks_hit-lag(shared_blks_hit::numeric,1,0::numeric) OVER (partition by dbid, userid, queryid ORDER BY snap_id) ) END AS Delta_shared_blks_hit,
case WHEN (shared_blks_read-lag(shared_blks_read::numeric,1,0::numeric) OVER (partition by dbid, userid, queryid ORDER BY snap_id) ) = shared_blks_read then null
else (shared_blks_read-lag(shared_blks_read::numeric,1,0::numeric) OVER (partition by dbid, userid, queryid ORDER BY snap_id) ) END AS DELTA_shared_blks_read,
case WHEN (shared_blks_dirtied-lag(shared_blks_dirtied::numeric,1,0::numeric) OVER (partition by dbid, userid, queryid ORDER BY snap_id) ) = shared_blks_dirtied then null
else (shared_blks_dirtied-lag(shared_blks_dirtied::numeric,1,0::numeric) OVER (partition by dbid, userid, queryid ORDER BY snap_id) ) END AS DELTA_shared_blks_dirtied,
case WHEN (shared_blks_written-lag(shared_blks_written::numeric,1,0::numeric) OVER (partition by dbid, userid, queryid ORDER BY snap_id) ) = shared_blks_written then null
else (shared_blks_written-lag(shared_blks_written::numeric,1,0::numeric) OVER (partition by dbid, userid, queryid ORDER BY snap_id) ) END AS DELTA_shared_blks_written,
case WHEN (temp_blks_written-lag(temp_blks_written::numeric,1,0::numeric) OVER (partition by dbid, userid, queryid ORDER BY snap_id) ) = temp_blks_written then null
else (temp_blks_written-lag(temp_blks_written::numeric,1,0::numeric) OVER (partition by dbid, userid, queryid ORDER BY snap_id) ) END AS DELTA_temp_blks_written,
case WHEN (blk_read_time-lag(blk_read_time::numeric,1,0::numeric) OVER (partition by dbid, userid, queryid ORDER BY snap_id) ) = blk_read_time then null
else (blk_read_time-lag(blk_read_time::numeric,1,0::numeric) OVER (partition by dbid, userid, queryid ORDER BY snap_id) ) END AS DELTA_blk_read_time
from get_sql_id
where snap_id between %(begin_snap_id)s-1 and %(end_snap_id)s
)
,time_partitioned_data as (
select dbid,userid,queryid,sum(delta_calls) AS calls,
round((sum(Delta_total_time)/sum(delta_calls))::numeric,2) AS AVG_ELAPSED_TIME,
round((sum(Delta_shared_blks_hit)/sum(delta_calls))::numeric) AS avg_shared_blks_hit,
round((sum(Delta_rows)/sum(delta_calls))::numeric) AS avg_rows,
round((sum(DELTA_shared_blks_dirtied)/sum(delta_calls))::numeric) AS avg_shared_blks_dirtied,
round((sum(DELTA_shared_blks_read)/sum(delta_calls))::numeric) AS avg_shared_blks_read,
round((sum(DELTA_shared_blks_written)/sum(delta_calls))::numeric) AS avg_shared_blks_written,
round((sum(DELTA_temp_blks_written)/sum(delta_calls))::numeric) AS avg_temp_blks_written,
round((sum(DELTA_blk_read_time)/sum(delta_calls))::numeric,2) AS avg_blk_read_time
from get_lag_data
where Delta_total_time is not null
and (delta_calls is not null )
group by dbid,userid,queryid
having sum(delta_calls)>0
)
select calls,avg_elapsed_time AS avg_elapsed_time_msec from time_partitioned_data
order by calls desc;''',cnx,params={'begin_snap_id' : 30, 'end_snap_id' : 180})
sql_dist.head()
# -
sql_dist.plot.scatter(x='calls', y='avg_elapsed_time_msec',title='SQL elpased Time vs # of Executions')
# +
index_util = pd.read_sql_query('''with pg_stat_database_vw as (
select a.snap_id,sample_start_time,datname,numbackends,
xact_commit ,
xact_rollback ,
blks_read as blks_read ,
blks_hit as blks_hit ,
tup_returned as tup_returned ,
tup_fetched as tup_fetched ,
tup_inserted as tup_inserted ,
tup_updated as tup_updated ,
tup_deleted as tup_deleted ,
conflicts ,
temp_files ,
temp_bytes as temp_bytes ,
deadlocks ,
blk_read_time ,
blk_write_time ,
stats_reset
from pg_stat_database_history a, pg_awr_snapshots_cust b where a.snap_id = b.snap_id
and a.snap_id between %(begin_snap_id)s and %(end_snap_id)s
and datname not in ('rdsadmin','template0','template1')
)
,get_lag_data as (select snap_id,sample_start_time,datname,
(lead(xact_commit::numeric,0) over (partition by datname order by snap_id)-lead(xact_commit::numeric,-1) over (partition by datname order by snap_id))/60 as delta_xact_commit,
(lead(xact_rollback::numeric,0) over (partition by datname order by snap_id)-lead(xact_rollback::numeric,-1) over (partition by datname order by snap_id))/60 as delta_xact_rollback,
(lead(blks_read::numeric,0) over (partition by datname order by snap_id)-lead(blks_read::numeric,-1) over (partition by datname order by snap_id))/60 as delta_blks_read,
(lead(blks_hit::numeric,0) over (partition by datname order by snap_id)-lead(blks_hit::numeric,-1) over (partition by datname order by snap_id))/60 as delta_blks_hit,
(lead(tup_returned::numeric,0) over (partition by datname order by snap_id)-lead(tup_returned::numeric,-1) over (partition by datname order by snap_id))/60 as delta_tup_returned,
(lead(tup_fetched::numeric,0) over (partition by datname order by snap_id)-lead(tup_fetched::numeric,-1) over (partition by datname order by snap_id))/60 as delta_tup_fetched,
(lead(tup_inserted::numeric,0) over (partition by datname order by snap_id)-lead(tup_inserted::numeric,-1) over (partition by datname order by snap_id))/60 as delta_tup_inserted,
(lead(tup_updated::numeric,0) over (partition by datname order by snap_id)-lead(tup_updated::numeric,-1) over (partition by datname order by snap_id))/60 as delta_tup_updated,
(lead(tup_deleted::numeric,0) over (partition by datname order by snap_id)-lead(tup_deleted::numeric,-1) over (partition by datname order by snap_id))/60 as delta_tup_deleted,
(lead(conflicts::numeric,0) over (partition by datname order by snap_id)-lead(conflicts::numeric,-1) over (partition by datname order by snap_id))/60 as delta_conflicts,
(lead(temp_files::numeric,0) over (partition by datname order by snap_id)-lead(temp_files::numeric,-1) over (partition by datname order by snap_id))/60 as delta_temp_files,
(lead(temp_bytes::numeric,0) over (partition by datname order by snap_id)-lead(temp_bytes::numeric,-1) over (partition by datname order by snap_id))/60 as delta_temp_bytes,
(lead(deadlocks::numeric,0) over (partition by datname order by snap_id)-lead(deadlocks::numeric,-1) over (partition by datname order by snap_id))/60 as delta_deadlocks,
(lead(blk_read_time::numeric,0) over (partition by datname order by snap_id)-lead(blk_read_time::numeric,-1) over (partition by datname order by snap_id))/60 as delta_blk_read_time,
(lead(blk_write_time::numeric,0) over (partition by datname order by snap_id)-lead(blk_write_time::numeric,-1) over (partition by datname order by snap_id))/60 as delta_blk_write_time,
stats_reset
from pg_stat_database_vw)
select snap_id,delta_tup_returned,delta_tup_fetched
from get_lag_data
where datname=%(db_name)s;''',cnx,params={'begin_snap_id' : 30, 'end_snap_id' : 180, 'db_name' : 'pg-ces-4c664f1d-e9b0-4dbf-a2dd-3a4b77b5161e'})
index_util.head()
# -
ax = index_util.plot(x='snap_id',title='rows fetched vs. rows returned',figsize=(20,5))
ax.set_ylabel("# of tuples")
# +
dml_stat = pd.read_sql_query('''with pg_stat_database_vw as (
select a.snap_id,sample_start_time,datname,numbackends,
xact_commit ,
xact_rollback ,
blks_read as blks_read ,
blks_hit as blks_hit ,
tup_returned as tup_returned ,
tup_fetched as tup_fetched ,
tup_inserted as tup_inserted ,
tup_updated as tup_updated ,
tup_deleted as tup_deleted ,
conflicts ,
temp_files ,
temp_bytes as temp_bytes ,
deadlocks ,
blk_read_time ,
blk_write_time ,
stats_reset
from pg_stat_database_history a, pg_awr_snapshots_cust b where a.snap_id = b.snap_id
and a.snap_id between %(begin_snap_id)s and %(end_snap_id)s
and datname not in ('rdsadmin','template0','template1')
)
,get_lag_data as (select snap_id,sample_start_time,datname,
(lead(xact_commit::numeric,0) over (partition by datname order by snap_id)-lead(xact_commit::numeric,-1) over (partition by datname order by snap_id))/60 as delta_xact_commit,
(lead(xact_rollback::numeric,0) over (partition by datname order by snap_id)-lead(xact_rollback::numeric,-1) over (partition by datname order by snap_id))/60 as delta_xact_rollback,
(lead(blks_read::numeric,0) over (partition by datname order by snap_id)-lead(blks_read::numeric,-1) over (partition by datname order by snap_id))/60 as delta_blks_read,
(lead(blks_hit::numeric,0) over (partition by datname order by snap_id)-lead(blks_hit::numeric,-1) over (partition by datname order by snap_id))/60 as delta_blks_hit,
(lead(tup_returned::numeric,0) over (partition by datname order by snap_id)-lead(tup_returned::numeric,-1) over (partition by datname order by snap_id))/60 as delta_tup_returned,
(lead(tup_fetched::numeric,0) over (partition by datname order by snap_id)-lead(tup_fetched::numeric,-1) over (partition by datname order by snap_id))/60 as delta_tup_fetched,
(lead(tup_inserted::numeric,0) over (partition by datname order by snap_id)-lead(tup_inserted::numeric,-1) over (partition by datname order by snap_id))/60 as delta_tup_inserted,
(lead(tup_updated::numeric,0) over (partition by datname order by snap_id)-lead(tup_updated::numeric,-1) over (partition by datname order by snap_id))/60 as delta_tup_updated,
(lead(tup_deleted::numeric,0) over (partition by datname order by snap_id)-lead(tup_deleted::numeric,-1) over (partition by datname order by snap_id))/60 as delta_tup_deleted,
(lead(conflicts::numeric,0) over (partition by datname order by snap_id)-lead(conflicts::numeric,-1) over (partition by datname order by snap_id))/60 as delta_conflicts,
(lead(temp_files::numeric,0) over (partition by datname order by snap_id)-lead(temp_files::numeric,-1) over (partition by datname order by snap_id))/60 as delta_temp_files,
(lead(temp_bytes::numeric,0) over (partition by datname order by snap_id)-lead(temp_bytes::numeric,-1) over (partition by datname order by snap_id))/60 as delta_temp_bytes,
(lead(deadlocks::numeric,0) over (partition by datname order by snap_id)-lead(deadlocks::numeric,-1) over (partition by datname order by snap_id))/60 as delta_deadlocks,
(lead(blk_read_time::numeric,0) over (partition by datname order by snap_id)-lead(blk_read_time::numeric,-1) over (partition by datname order by snap_id))/60 as delta_blk_read_time,
(lead(blk_write_time::numeric,0) over (partition by datname order by snap_id)-lead(blk_write_time::numeric,-1) over (partition by datname order by snap_id))/60 as delta_blk_write_time,
stats_reset
from pg_stat_database_vw)
select snap_id,delta_tup_inserted,delta_tup_updated,delta_tup_deleted
from get_lag_data
where datname=%(db_name)s;''',cnx,params={'begin_snap_id' : 30, 'end_snap_id' : 180, 'db_name' : 'pg-ces-4c664f1d-e9b0-4dbf-a2dd-3a4b77b5161e'})
dml_stat.head()
# -
ax = dml_stat.plot(x='snap_id',title='Tuples inserted / updated / deleted',figsize=(20,5))
ax.set_ylabel("# of tuples")
# +
scan_stat = pd.read_sql_query('''with pg_stat_all_tables_vw as (
select
a.snap_id ,
sample_start_time ,
relid ,
schemaname ,
relname ,
seq_scan ,
seq_tup_read ,
idx_scan ,
idx_tup_fetch ,
n_tup_ins ,
n_tup_upd ,
n_tup_del ,
n_tup_hot_upd ,
n_live_tup ,
n_dead_tup ,
autovacuum_count
from pg_stat_all_tables_history a, pg_awr_snapshots_cust b where a.snap_id = b.snap_id
and a.snap_id between %(begin_snap_id)s and %(end_snap_id)s
and schemaname not in ('pg_catalog')),
get_delta_data as (select snap_id,relid,sample_start_time,schemaname,relname,
case WHEN (seq_scan-lag(seq_scan::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) = seq_scan then null
else (seq_scan-lag(seq_scan::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) END AS delta_seq_scan,
case WHEN (seq_tup_read-lag(seq_tup_read::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) = seq_tup_read then null
else (seq_tup_read-lag(seq_tup_read::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) END AS delta_seq_tup_read,
case WHEN (idx_scan-lag(idx_scan::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) = idx_scan then null
else (idx_scan-lag(idx_scan::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) END AS delta_idx_scan,
case WHEN (idx_tup_fetch-lag(idx_tup_fetch::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) = idx_tup_fetch then null
else (idx_tup_fetch-lag(idx_tup_fetch::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) END AS delta_idx_tup_fetch,
case WHEN (n_tup_ins-lag(n_tup_ins::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) = n_tup_ins then null
else (n_tup_ins-lag(n_tup_ins::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) END AS delta_n_tup_ins,
case WHEN (n_tup_upd-lag(n_tup_upd::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) = n_tup_upd then null
else (n_tup_upd-lag(n_tup_upd::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) END AS delta_n_tup_upd,
case WHEN (n_tup_del-lag(n_tup_del::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) = n_tup_del then null
else (n_tup_del-lag(n_tup_del::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) END AS delta_n_tup_del,
case WHEN (n_tup_hot_upd-lag(n_tup_hot_upd::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) = n_tup_hot_upd then null
else (n_tup_hot_upd-lag(n_tup_hot_upd::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) END AS delta_n_tup_hot_upd,
case WHEN (n_live_tup-lag(n_live_tup::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) = n_live_tup then null
else (n_live_tup-lag(n_live_tup::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) END AS delta_n_live_tup,
case WHEN (n_dead_tup-lag(n_dead_tup::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) = n_dead_tup then null
else (n_dead_tup-lag(n_dead_tup::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) END AS delta_n_dead_tup,
case WHEN (autovacuum_count-lag(autovacuum_count::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) = autovacuum_count then null
else (autovacuum_count-lag(autovacuum_count::bigint,1,0::bigint) OVER (partition by schemaname,relname ORDER BY snap_id) ) END AS delta_autovacuum_count
from pg_stat_all_tables_vw)
select snap_id, sum(delta_seq_scan) AS total_full_table_scans,
sum(delta_idx_scan) AS total_index_scans
from get_delta_data
group by snap_id
order by snap_id;''',cnx,params={'begin_snap_id' : 30, 'end_snap_id' : 180})
scan_stat.head()
# -
ax = scan_stat.plot(x='snap_id',title='Sequential Scans vs Index Scans initiated on tables',figsize=(20,5))
ax.set_ylabel("# of Scans")
# +
session_stat = pd.read_sql_query('''select snap_id,case when state is NULL then 'Undefined' ELSE state END as session_state,count(*) total_cnt from pg_stat_activity_history
where datname not in ('template0','template1','rdsadmin')
and snap_id between %(begin_snap_id)s and %(end_snap_id)s
group by snap_id,state
order by 1,2;''',cnx,params={'begin_snap_id' : 30, 'end_snap_id' : 180})
session_stat.head()
# -
session_stat1=session_stat.pivot(index='snap_id', columns='session_state', values='total_cnt')
ax = session_stat1.plot(title='Session State',figsize=(20,10))
ax.set_ylabel("# of sessions")
| Code/PGPerfStatsSnapper/Juypter/snapper_exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
print(math.factorial(5))
print(math.factorial(0))
# +
# print(math.factorial(1.5))
# ValueError: factorial() only accepts integral values
# +
# print(math.factorial(-1))
# ValueError: factorial() not defined for negative values
# -
def permutations_count(n, r):
return math.factorial(n) // math.factorial(n - r)
print(permutations_count(4, 2))
print(permutations_count(4, 4))
def combinations_count(n, r):
return math.factorial(n) // (math.factorial(n - r) * math.factorial(r))
print(combinations_count(4, 2))
def combinations_with_replacement_count(n, r):
return combinations_count(n + r - 1, r)
print(combinations_with_replacement_count(4, 2))
| notebook/math_factorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
__name__ = "k1lib.callbacks"
#export
import k1lib, torch; from k1lib.cli import empty, shape
from .callbacks import Callback, Callbacks, Cbs
from typing import Tuple, List
__all__ = ["Recorder"]
#export
@k1lib.patch(Cbs)
class Recorder(Callback):
"""Records xb, yb and y from a short run. No training involved.
Example::
l = k1lib.Learner.sample()
l.cbs.add(Cbs.Recorder())
xbs, ybs, ys = l.Recorder.record(1, 2)
xbs # list of x batches passed in
ybs # list of y batches passed in, "the correct label"
ys # list of network's output
If you have extra metadata in your dataloader, then the recorder will return
(xb, yb, metab, ys) instead::
# creating a new dataloader that yields (xb, yb, metadata)
x = torch.linspace(-5, 5, 1000); meta = torch.tensor(range(1000))
dl = [x, x+2, meta] | transpose() | randomize(None) | repeatFrom() | batched()\
| (transpose() | (toTensor() + toTensor() + toTensor())).all() | stagger(50)
l = k1lib.Learner.sample(); l.data = [dl, []]
l.cbs.add(Cbs.Recorder())
xbs, ybs, metabs, ys = l.Recorder.record(1, 2)
"""
def __init__(self):
super().__init__(); self.order = 20; self.suspended = True
def startRun(self):
self.xbs = []; self.ybs = []; self.metabs = []; self.ys = []
def startBatch(self):
self.xbs.append(self.l.xb.detach())
self.ybs.append(self.l.yb.detach())
self.metabs.append(self.l.metab)
def endRun(self):
n = min(len(self.xbs), len(self.ybs), len(self.metabs), len(self.ys))
self.xbs = self.xbs[:n]; self.ybs = self.ybs[:n]
self.metabs = self.metabs[:n]; self.ys = self.ys[:n]
def endPass(self):
self.ys.append(self.l.y.detach())
@property
def values(self):
hasMeta = self.metabs | ~empty() | shape(0) > 0
if hasMeta: return self.xbs, self.ybs, self.metabs, self.ys
else: return self.xbs, self.ybs, self.ys
def record(self, epochs:int=1, batches:int=None) -> Tuple[List[torch.Tensor], List[torch.Tensor], List[torch.Tensor]]:
"""Returns recorded xBatch, yBatch and answer y"""
self.suspended = False
try:
with self.cbs.context(), self.cbs.suspendEval():
self.cbs.add(Cbs.DontTrain()).add(Cbs.TimeLimit(5))
self.l.run(epochs, batches)
finally: self.suspended = True
return self.values
def __repr__(self):
return f"""{self._reprHead}, can...
- r.record(epoch[, batches]): runs for a while, and records x and y batches, and the output
{self._reprCan}"""
l = k1lib.Learner.sample()
l.cbs.add(Cbs.Recorder())
xb, yb, y = l.Recorder.record(1, 2)
assert len(xb) == 2; assert len(yb) == 2; assert len(y) == 2
assert xb[0].shape == torch.Size([32])
# +
from k1lib.cli import *
# creating a new dataloader that yields (xb, yb, metadata)
x = torch.linspace(-5, 5, 1000); meta = torch.tensor(range(1000))
dl = [x, x+2, meta] | transpose() | randomize(None) | repeatFrom() | batched()\
| (transpose() | (toTensor() + toTensor() + toTensor())).all() | stagger(50)
l = k1lib.Learner.sample(); l.data = [dl, []]
l.cbs.add(Cbs.Recorder())
xbs, ybs, metabs, ys = l.Recorder.record(1, 2)
# -
# !../../export.py callbacks/recorder
| k1lib/callbacks/recorder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using R with pyABC
# This example illustrates how to use models, summary statistics and
# distance functions defined in R. We're assuming you're already
# familiar with the basic workings of pyABC. If not, consult the
# other tutorial examples.
# + raw_mimetype="text/restructuredtext" active=""
# Download this notebook :download:`Using R with pyABC <using_R.ipynb>`.
# + raw_mimetype="text/restructuredtext" active=""
# In this example, we're introducing the new class
# :class:`R <pyabc.external.R>` which is our interface with R.
# We use this class to to load an external R script.
# +
# %matplotlib inline
from pyabc.external import R
r = R("myRModel.R")
# + raw_mimetype="text/restructuredtext" active=""
# .. note::
#
# ``R("myRModel.R")`` does, amongst other things, the equivalent
# to R's ``source("myRModel.R")``. That is, the entire script is
# executed with all the possible side effects this might have.
#
#
# You can download the file here: :download:`myRModel.R <myRModel.R>`.
# But now, let's have a look at it.
# -
r.display_source_ipython()
# + [markdown] raw_mimetype="text/restructuredtext"
# We see that four relevant objects are defined in the file.
#
# * myModel
# * mySummaryStatistics (optional)
# * myDistance
# * mySumStatData
#
# The names of these do not matter. The ``mySummaryStatistics`` is actually optional and can be omitted
# in case the model calculates the summary statistics directly. We load the defined functions using the ``r`` object:
# -
model = r.model("myModel")
distance = r.distance("myDistance")
sum_stat = r.summary_statistics("mySummaryStatistics")
# + [markdown] raw_mimetype="text/restructuredtext"
# From there on, we can use them (almost) as if they were ordinary Python functions.
# +
from pyabc import Distribution, RV, ABCSMC
prior = Distribution(meanX=RV("uniform", 0, 10),
meanY=RV("uniform", 0, 10))
abc = ABCSMC(model, prior, distance,
summary_statistics=sum_stat)
# + [markdown] raw_mimetype="text/restructuredtext"
# We also load the observation with ``r.observation`` and pass it to a new ABC run.
# +
import os
from tempfile import gettempdir
db = "sqlite:///" + os.path.join(gettempdir(), "test.db")
abc.new(db, r.observation("mySumStatData"))
# + [markdown] raw_mimetype="text/restructuredtext"
# We start a run which terminates as soon as an acceptance threshold of 0.9 or less is reached
# or the maximum number of 4 populations is sampled.
# -
history = abc.run(minimum_epsilon=0.9, max_nr_populations=4)
# + [markdown] raw_mimetype="text/restructuredtext"
# Lastly, we plot the results and observe how the generations contract slowly around the oserved value.
# (Note, that the contraction around the observed value is a particular property of the chosen example and not always the case.)
# +
from pyabc.visualization import plot_kde_2d
for t in range(history.n_populations):
df, w = abc.history.get_distribution(0, t)
ax = plot_kde_2d(df, w, "meanX", "meanY",
xmin=0, xmax=10,
ymin=0, ymax=10,
numx=100, numy=100)
ax.scatter([4], [8],
edgecolor="black",
facecolor="white",
label="Observation");
ax.legend();
ax.set_title("PDF t={}".format(t))
# + [markdown] raw_mimetype="text/restructuredtext"
# And we can also retrieve summary statistics such as a stored
# DataFrame, although the DataFrame was acutally defined in R.
# -
history.get_weighted_sum_stats_for_model(m=0, t=1)[1][0]["cars"].head()
# + raw_mimetype="text/restructuredtext" active=""
# Dumping the results to a file format R can read
# -----------------------------------------------
#
# Although you could query pyABC's database directly from R since the database
# is just a SQL database (e.g. SQLite), pyABC ships with a utility for facilitate
# export of the database.
# Use the ``abc-dump`` utility provided by pyABC to dump results to
# file formats such as csv, feather, html, json and others.
# These can be easiliy read in by R. See `Exporting pyABC's database <../export_db.rst>`_ for how to use this
# utility.
#
# Assume you dumped to the feather format::
#
# abc-export --db results.db --out exported.feather --format feather
#
# You could read the results in with the following R snippet
#
# .. code:: R
#
#
# install.packages("feather")
# install.packages("jsonlite")
#
# library("feather")
# library("jsonlite")
#
# loadedDf <- data.frame(feather("exported.feather"))
#
# jsonStr <- loadedDf$sumstat_ss_df[1]
#
# sumStatDf <- fromJSON(jsonStr)
#
# If you prefer CSV over the feather format you can also do that.
| doc/examples/using_R.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import os
from datetime import date
cols = ["SERIAL","EQUIPMENTKEY","CUSTOMATTR15","SUMMARY","LASTOCCURRENCE","CLEARTIMESTAMP","ALARMDETAILS"]
td = date.today()
pt = os.getcwd() + "\\" + "RRU_" + td.strftime('%Y-%m-%d') + ".csv"
TS2 = lambda y : ('NA' if (y is None) else y)
single = os.getcwd() + "\\" + "DWRRU.csv"
df = pd.read_csv(single)
df2 = df[cols] #pick customized column using list
df2['ABC'] = df2.apply(lambda x : TS2(df2['CUSTOMATTR15'].value_counts()),axis=1)
#code= [df2['CUSTOMATTR15'].value_counts(dropna=False)]
#ndf = pd.DataFrame(code).T #list to dataframe
#ndf.to_csv(pt)
# -
| Z_ALL_FILE/Jy1/lamda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
from omegaconf import OmegaConf
# Encoder settings
encoder_config = OmegaConf.create({
"embedding": {
"name": "FeatureEmbedding",
},
"backbone": {
"name": "FTTransformerBackbone",
}
})
# model settings (learning rate, scheduler...)
model_config = OmegaConf.create({
"name": "MLPHeadModel"
})
# training settings (epoch, gpu...): not necessary
trainer_config = OmegaConf.create({
"max_epochs": 1,
})
# +
import os,sys; sys.path.append(os.path.abspath(".."))
from deep_table.data.data_module import TabularDatamodule
from deep_table.data.datasets import Adult
adult_dataset = Adult(root="../data")
adult_dataframes = adult_dataset.processed_dataframes()
datamodule = TabularDatamodule(
train=adult_dataframes["train"],
val=adult_dataframes["val"],
test=adult_dataframes["test"],
task=adult_dataset.task,
dim_out=adult_dataset.dim_out,
categorical_columns=adult_dataset.categorical_columns,
continuous_columns=adult_dataset.continuous_columns,
target=adult_dataset.target_columns,
num_categories=adult_dataset.num_categories(),
)
# +
from deep_table.estimators.base import Estimator
from deep_table.utils import get_scores
estimator = Estimator(
encoder_config,
model_config,
trainer_config
)
estimator.fit(datamodule)
# -
predict = estimator.predict(datamodule.dataloader(split="test"))
get_scores(predict, target=datamodule.dataloader(split="test"), task="binary")
# +
pretrain_model_config = OmegaConf.create({
"name": "SAINTPretrainModel"
})
pretrain_model = Estimator(
encoder_config,
pretrain_model_config,
trainer_config
)
pretrain_model.fit(datamodule)
estimator = Estimator(
encoder_config, model_config,
trainer_config)
estimator.fit(datamodule, from_pretrained=pretrain_model)
# -
predict = estimator.predict(datamodule.dataloader(split="test"))
get_scores(predict, target=datamodule.dataloader(split="test"), task="binary")
| notebooks/train_adult.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fast Python3 For Beginners
# ___
# +
import logging
import logging.handlers
import os
import os.path
global g_log_inst
class Logger(object):
_inst = None
_level_dict = {
'CRITICAL': logging.CRITICAL,
'ERROR': logging.ERROR,
'WARNING': logging.WARNING,
'INFO': logging.INFO,
'DEBUG': logging.DEBUG,
'NOTSET': logging.NOTSET,
}
@classmethod
def start(cls, log_path, name=None, level=None):
if cls._inst is not None:
return cls._inst
fpath = '/'.join(log_path.split('/')[0 : -1])
if os.path.exists(fpath) == False:
os.mkdir(fpath)
fmt = '[%(levelname)s] %(asctime)s, pid=%(process)d, src=%(filename)s:%(lineno)d, %(message)s'
datefmt = '%Y-%m-%d %H:%M:%S'
cls._inst = logging.getLogger(name)
log_level = Logger._level_dict[level] if level else 'DEBUG'
cls._inst.setLevel(log_level)
# 1 << 20 即 2**20 or 2^20
handler = logging.handlers.RotatingFileHandler(log_path, maxBytes=500 * (1<<20), backupCount=8)
fmtter = logging.Formatter(fmt, datefmt)
handler.setFormatter(fmtter)
cls._inst.addHandler(handler)
@classmethod
def get(cls):
return cls._inst
g_log_inst = Logger
# -
# ### Usage
# +
from log import g_log_inst as logger
logger.get().debug()
logger.get().info()
# -
'Done!\N{Cat}'
| 03 Exception Handling/02_A_Logger_Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # knn
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
dfx = pd.read_csv('./xdata.csv')
dfy = pd.read_csv('./ydata.csv')
# +
x = dfx.values
y = dfy.values
x=x[:,1:]
y=y[:,1:].reshape((-1,))
print(x)
print(x.shape)
print(y.shape)
# -
plt.scatter(x[:,0],x[:,1],c=y)
plt.show()
query_x = np.array([2,3])
plt.scatter(x[:,0],x[:,1],c=y)
plt.scatter(query_x[0],query_x[1],color='red')
plt.show()
# +
def dist(x1,x2):
return np.sqrt(sum((x1-x2)**2))
def knn(x,y,queryPoint,k=5):
vals = []
m = x.shape[0]
for i in range(m):
d = dist(queryPoint,x[i])
vals.append((d,y[i]))
vals = sorted(vals)
vals = vals[:k]
vals = np.array(vals)
# print(vals)
new_vals = np.unique(vals[:,1],return_counts = True)
# print(new_vals)
index = new_vals[1].argmax()
pred = new_vals[0][index]
return pred
# -
xk = knn(x,y,[2.9,1])
print(xk)
| knn/knn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Arrays: Left Rotation
# [Challenge Link](https://www.hackerrank.com/challenges/ctci-array-left-rotation/problem)
# Check out the resources on the page's right side to learn more about arrays. The video tutorial is by <NAME>, author of the best-selling interview book Cracking the Coding Interview.
#
# A left rotation operation on an array shifts each of the array's elements 1 unit to the left. For example, if 2 left rotations are performed on array `[1, 2, 3, 4, 5]`, then the array would become `[3, 4, 5, 1, 2]`.
#
# Given an array `a` of `n` integers and a number, `d`, perform `d` left rotations on the array. Return the updated array to be printed as a single line of space-separated integers.
# ### Objective
# Complete the function rotLeft in the editor below. It should return the resulting array of integers.
#
# rotLeft has the following parameter(s):
#
# An array of integers a.
# An integer d, the number of rotations.
# ### Input
from io import StringIO
sample_input = lambda: StringIO('''5 4
1 2 3 4 5''')
# #### Solution (Naive)
def rot_left(array, d):
new_array = list(array)
for index, element in enumerate(array):
new_index = (index - d) % len(array)
new_array[new_index] = element
return new_array
# #### Output
def solve_array_left_rotation(stream):
len, d = [int(x) for x in stream.readline().split()]
array = [int(x) for x in stream.readline().split()]
print(' '.join(
[str(x) for x in rot_left(array, d)]
))
solve_array_left_rotation(sample_input())
# ### Analysis
# The time and space complexity of rot_left are currently dependent on array input length, thus `O(N)`.
# Apart from looking at the Code in order to figure out complexity, we can show this by constructing large test cases and looking at the mean execution time.
import random
list_50k = [random.randint(0, 65535) for _ in range(50000)]
list_100k = [random.randint(0, 65535) for _ in range(100000)]
# %timeit rot_left(list_50k, random.randint(0, 1000))
# %timeit rot_left(list_100k, random.randint(0, 1000))
# ### Improvement
# Can we do any better?
# #### Inplace Rotation
# We can rotate the array inplace, by first determining which parts of it will overflow to the right, then copying that part, deleting the beginning of the list, and appending it to the end. Depending on the Array Implementation, this makes the algorithm be O(d) -- rotation is soley dependant on the rotation argument. In Python, this isn't the case because list slice deletion is O(n) unfortunately. In any case, this approach is orders of magnitudes faster, because we don't have to copy anything!
def rot_left_inplace(array, d):
overflow_left = array[0:d]
del array[0:d]
array.extend(overflow_left)
return array
# %timeit rot_left_inplace(list_50k, 1000)
# %timeit rot_left_inplace(list_100k, 1000)
# #### Juggling Algorithm
# Another smart approach is to compute `gcd(d, len(list))`, break the array into sections depending on that, and do everything inplace. Geek2Geek has a good explanation on this: https://www.geeksforgeeks.org/array-rotation/ The original Algorithm is from the Book Programming Pearls by <NAME>. In Python, we won't do better than the last Solution though in terms of speed, because the array is copied one by one, which is expensive. However, our space requirements are consistent no matter the input size with this!
import math
def rot_left_inplace_juggling(array, d):
n = len(array)
for i in range(math.gcd(d, n)):
temp = array[i]
j = i
while 1:
k = j + d
if k >= n:
k = k - n
if k == i:
break
array[j] = array[k]
j = k
array[j] = temp
# %timeit rot_left_inplace_juggling(list_50k, 1000)
# %timeit rot_left_inplace_juggling(list_100k, 1000)
| cracking_the_coding_interview/arrays_left_rotation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="9fUs4rmB9ZN7" colab_type="code" outputId="e8d2c3cd-9f79-4a09-f6b0-8cfbb780e048" executionInfo={"status": "ok", "timestamp": 1584022975857, "user_tz": -180, "elapsed": 2617, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjf7mL8J_VkRjlogmAa7HYBCvaLMJSd7yIl5KJK=s64", "userId": "00418095213724128690"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# %tensorflow_version 2.x
import tensorflow as tf
# layers
from tensorflow.keras.layers import Input, SimpleRNN, Dense, Flatten
# model
from tensorflow.keras.models import Model
# optimizers
from tensorflow.keras.optimizers import Adam, SGD
# additional imports
import numpy as np # linear algebra
import pandas as pd # data manipulation
import matplotlib.pyplot as plt # data visualisation
# + id="5fPQfPum_DSZ" colab_type="code" outputId="a35f3c22-3b8a-4fe0-b741-36b1af1843af" executionInfo={"status": "ok", "timestamp": 1584022975858, "user_tz": -180, "elapsed": 2607, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjf7mL8J_VkRjlogmAa7HYBCvaLMJSd7yIl5KJK=s64", "userId": "00418095213724128690"}} colab={"base_uri": "https://localhost:8080/", "height": 265}
# make the data # adding noise
series = np.sin(0.1*np.arange(200)) + np.random.randn(200)*0.1
# plot the data
plt.plot(series)
plt.show()
# + id="n1B_PH7F_bDp" colab_type="code" outputId="a5b36ce4-953e-477e-aa16-71127504e0ef" executionInfo={"status": "ok", "timestamp": 1584023266041, "user_tz": -180, "elapsed": 746, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjf7mL8J_VkRjlogmAa7HYBCvaLMJSd7yIl5KJK=s64", "userId": "00418095213724128690"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# build the dataset
T = 10
D = 1
X = []
Y = []
for t in range(len(series) -T):
x = series[t:t+T]
X.append(x)
y = series[t+T]
Y.append(y)
X = np.array(X).reshape(-1, T, 1) # Now the data should be NxTxD to be passed into RNN
Y = np.array(Y)
N = len(X)
print('X.shape', X.shape, 'Y.shape', Y.shape)
# + id="yjZ0ISjO_4WM" colab_type="code" outputId="81b5cc92-74ad-4215-d4b2-62ef3caa654c" executionInfo={"status": "ok", "timestamp": 1584023387093, "user_tz": -180, "elapsed": 5386, "user": {"displayName": "<NAME>", "photoUrl": "<KEY>", "userId": "00418095213724128690"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
# Now try autoregressive RNN model
i = Input(shape=(T, 1))
# SimpleRNN has the default activation function as tanh
# but you can specify something else like ReLU
# Also you can pass None as activation, this'll reduce RNN into linear model.
# Also the data we generated is linear hence we shall pass None as activation function. Without activation function, RNN will reduce to linear model
# and it will work like the model in: https://colab.research.google.com/drive/1pD2k_5gIoLNTYoh_dLcQOKSiMrtpQ17_
x = SimpleRNN(5)(i)
x = Dense(1)(x)
model = Model(i,x)
model.compile(loss = 'mse', optimizer = Adam(lr = 0.1))
# Train the RNN
r = model.fit(
X[:-N//2], Y[:-N//2], # Using first half as a training data
validation_data = (X[-N//2:], Y[-N//2:]), # using last half as validation
epochs = 100
)
# + id="HWibwBhKA118" colab_type="code" outputId="781cdd6b-caed-418d-b420-4f51e3fbaf91" executionInfo={"status": "ok", "timestamp": 1584023387094, "user_tz": -180, "elapsed": 5231, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjf7mL8J_VkRjlogmAa7HYBCvaLMJSd7yIl5KJK=s64", "userId": "00418095213724128690"}} colab={"base_uri": "https://localhost:8080/", "height": 265}
# now plot the results
plt.plot(r.history['loss'], label = 'loss')
plt.plot(r.history['val_loss'], label = 'val_loss')
plt.legend()
plt.show()
# + id="DxM7NDMlCNOi" colab_type="code" outputId="a26955ee-161a-43ab-e986-d214d1293dfd" executionInfo={"status": "ok", "timestamp": 1584023389706, "user_tz": -180, "elapsed": 7701, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjf7mL8J_VkRjlogmAa7HYBCvaLMJSd7yIl5KJK=s64", "userId": "00418095213724128690"}} colab={"base_uri": "https://localhost:8080/", "height": 54}
# 'wrong' forecast using true targets
validation_target = Y[-N//2:]
validation_predictions = []
i = -N//2
while len(validation_predictions) < len(validation_target):
p = model.predict(X[i].reshape(1, -1, 1))[0,0] # 1x1 array -> scalar
i += 1
validation_predictions.append(p)
'''
This kind of forecasting is wrong because it uses every X on the
dataset. So this isn't the correct way to do forecasting.
In the correct way,
forecasting starts by last X value and continues to predict next value
using its own PREDICTIONS. Not every X like this approach
'''
# + id="Rr2WLVOfDOvt" colab_type="code" outputId="e2a73c54-b9ca-4b32-b4ef-00113d28961d" executionInfo={"status": "ok", "timestamp": 1584023389707, "user_tz": -180, "elapsed": 7572, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjf7mL8J_VkRjlogmAa7HYBCvaLMJSd7yIl5KJK=s64", "userId": "00418095213724128690"}} colab={"base_uri": "https://localhost:8080/", "height": 265}
# plot the loss
plt.plot(validation_target, label = 'forecast target')
plt.plot(validation_predictions, label = 'forecast predictions')
plt.legend()
plt.show()
# + id="x1HDrEduDW3a" colab_type="code" colab={}
# correct way of doing forecasting which uses only self-predictions for making future predictions
validation_target = Y[-N//2:]
validation_predictions = []
# last train input
last_x = X[-N//2] # 1-D array of length T
while len(validation_predictions) < len(validation_target):
p = model.predict(last_x.reshape(1, -1, 1))[0,0] # 1x1 array -> scalar
# update the predictions list
validation_predictions.append(p)
# make the new input
last_x = np.roll(last_x, -1)
last_x[-1] = p
# + id="nNFRhy-MGZi1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="1d9a15d2-47bd-467f-b714-66c8f1d23524" executionInfo={"status": "ok", "timestamp": 1584023392492, "user_tz": -180, "elapsed": 10076, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjf7mL8J_VkRjlogmAa7HYBCvaLMJSd7yIl5KJK=s64", "userId": "00418095213724128690"}}
# plot the loss
plt.plot(validation_target, label = 'forecast target')
plt.plot(validation_predictions, label = 'forecast preds')
plt.legend()
plt.show()
# + id="uNBdInGbGhET" colab_type="code" colab={}
# So we can say that RNN doesn't works well on noisy data
# I guess what we've created isn't even a model
# it just takes last value to predict the next one
# so we can say that RNN is not optimal or needs to be tuned
# in order to work efficiently on this problem
| Tensorflow 2/RNN/SimpleRNN to forecast Sine TF2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Which of the following **must be true** for the following code to execute without error?
#
# ```python
# for x in C:
# print(x)
# ```
#
# If one of the choices **must be true**, then explain. If not, write a (tiny!) class + iterator such that the code above runs. Note: the same class + iterator might serve as a counterexample for multiple parts, in which case you can just say so.
#
# First alphabetical by first name is the **Recorder**. Second is the **Facilitator**. Third is the **Reporter**.
# +
# (A): C has a __next__() method.
# Explanation or counterexample here.
class my_class:
def __iter__(self):
return my_iterator()
class my_iterator:
def __next__(self):
return 0
# ----------------
class my_class:
def __iter__(self):
return self
def __next__(self):
return 0
C = my_class()
# for x in C:
# print(x)
# +
#### (B): C has an __iter__() method.
# Explanation or counterexample here.
# in order for this function call to do anything iter(C)
# must have __iter__() magic method
# +
# (C): C is a container.
# Explanation or counterexample here.
# +
# (D): The code len(C) executes without error.
# Explanation or counterexample here.
# __len__() needed for len(C) to work
len(C)
# +
# (E): The output of iter(C) has a __next__() method.
# Explanation or counterexample here.
# need to be able to next() on iter(C)
# next(iter(C))
# +
# (F): C possesses an instance variable i
# which represents the current state of iteration.
# Explanation or counterexample here.
# not required, only needed for certain ways to stop iteration
| live_lectures/live-lecture-8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="wDlWLbfkJtvu" colab_type="code" cellView="form" colab={}
#@title Copyright 2020 Google LLC. Double-click here for license information.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="TL5y5fY9Jy_x" colab_type="text"
# # Linear Regression with a Real Dataset
#
# This Colab uses a real dataset to predict the prices of houses in California.
#
#
#
#
#
# + [markdown] id="h8wtceyJj2uX" colab_type="text"
# ## Learning Objectives:
#
# After doing this Colab, you'll know how to do the following:
#
# * Read a .csv file into a [pandas](https://developers.google.com/machine-learning/glossary/#pandas) DataFrame.
# * Examine a [dataset](https://developers.google.com/machine-learning/glossary/#data_set).
# * Experiment with different [features](https://developers.google.com/machine-learning/glossary/#feature) in building a model.
# * Tune the model's [hyperparameters](https://developers.google.com/machine-learning/glossary/#hyperparameter).
# + [markdown] id="JJZEgJQSjyK4" colab_type="text"
# ## The Dataset
#
# The [dataset for this exercise](https://developers.google.com/machine-learning/crash-course/california-housing-data-description) is based on 1990 census data from California. The dataset is old but still provides a great opportunity to learn about machine learning programming.
# + [markdown] id="tX_umRMMsa3z" colab_type="text"
# ## Use the right version of TensorFlow
#
# The following hidden code cell ensures that the Colab will run on TensorFlow 2.X.
# + id="lM75uNH-sTv2" colab_type="code" cellView="form" colab={}
#@title Run on TensorFlow 2.x
# %tensorflow_version 2.x
# + [markdown] id="xchnxAsaKKqO" colab_type="text"
# ## Import relevant modules
#
# The following hidden code cell imports the necessary code to run the code in the rest of this Colaboratory.
# + id="9n9_cTveKmse" colab_type="code" cellView="both" colab={}
#@title Import relevant modules
import pandas as pd
import tensorflow as tf
from matplotlib import pyplot as plt
# The following lines adjust the granularity of reporting.
pd.options.display.max_rows = 10
pd.options.display.float_format = "{:.1f}".format
# + [markdown] id="X_TaJhU4KcuY" colab_type="text"
# ## The dataset
#
# Datasets are often stored on disk or at a URL in [.csv format](https://wikipedia.org/wiki/Comma-separated_values).
#
# A well-formed .csv file contains column names in the first row, followed by many rows of data. A comma divides each value in each row. For example, here are the first five rows of the .csv file file holding the California Housing Dataset:
#
# ```
# "longitude","latitude","housing_median_age","total_rooms","total_bedrooms","population","households","median_income","median_house_value"
# -114.310000,34.190000,15.000000,5612.000000,1283.000000,1015.000000,472.000000,1.493600,66900.000000
# -114.470000,34.400000,19.000000,7650.000000,1901.000000,1129.000000,463.000000,1.820000,80100.000000
# -114.560000,33.690000,17.000000,720.000000,174.000000,333.000000,117.000000,1.650900,85700.000000
# -114.570000,33.640000,14.000000,1501.000000,337.000000,515.000000,226.000000,3.191700,73400.000000
# ```
#
#
# + [markdown] id="sSFQkzNlj-l6" colab_type="text"
# ### Load the .csv file into a pandas DataFrame
#
# This Colab, like many machine learning programs, gathers the .csv file and stores the data in memory as a pandas Dataframe. pandas is an open source Python library. The primary datatype in pandas is a DataFrame. You can imagine a pandas DataFrame as a spreadsheet in which each row is identified by a number and each column by a name. pandas is itself built on another open source Python library called NumPy. If you aren't familiar with these technologies, please view these two quick tutorials:
#
# * [NumPy](https://colab.research.google.com/github/google/eng-edu/blob/master/ml/cc/exercises/numpy_ultraquick_tutorial.ipynb?utm_source=linearregressionreal-colab&utm_medium=colab&utm_campaign=colab-external&utm_content=numpy_tf2-colab&hl=en)
# * [Pandas DataFrames](https://colab.research.google.com/github/google/eng-edu/blob/master/ml/cc/exercises/pandas_dataframe_ultraquick_tutorial.ipynb?utm_source=linearregressionreal-colab&utm_medium=colab&utm_campaign=colab-external&utm_content=pandas_tf2-colab&hl=en)
#
# The following code cell imports the .csv file into a pandas DataFrame and scales the values in the label (`median_house_value`):
# + id="JZlvdpyYKx7V" colab_type="code" colab={}
# Import the dataset.
training_df = pd.read_csv(filepath_or_buffer="https://download.mlcc.google.com/mledu-datasets/california_housing_train.csv")
# Scale the label.
training_df["median_house_value"] /= 1000.0
# Print the first rows of the pandas DataFrame.
training_df.head()
# + [markdown] id="5inxx49n4U9u" colab_type="text"
# Scaling `median_house_value` puts the value of each house in units of thousands. Scaling will keep loss values and learning rates in a friendlier range.
#
# Although scaling a label is usually *not* essential, scaling features in a multi-feature model usually *is* essential.
# + [markdown] id="yMysi6-3IAbu" colab_type="text"
# ## Examine the dataset
#
# A large part of most machine learning projects is getting to know your data. The pandas API provides a `describe` function that outputs the following statistics about every column in the DataFrame:
#
# * `count`, which is the number of rows in that column. Ideally, `count` contains the same value for every column.
#
# * `mean` and `std`, which contain the mean and standard deviation of the values in each column.
#
# * `min` and `max`, which contain the lowest and highest values in each column.
#
# * `25%`, `50%`, `75%`, which contain various [quantiles](https://developers.google.com/machine-learning/glossary/#quantile).
# + id="rnUSYKw4LUuh" colab_type="code" colab={}
# Get statistics on the dataset.
training_df.describe()
# + [markdown] id="f9pcW_Yjtoo8" colab_type="text"
# ### Task 1: Identify anomalies in the dataset
#
# Do you see any anomalies (strange values) in the data?
# + id="UoS7NWRXEs1H" colab_type="code" cellView="form" colab={}
#@title Double-click to view a possible answer.
# The maximum value (max) of several columns seems very
# high compared to the other quantiles. For example,
# example the total_rooms column. Given the quantile
# values (25%, 50%, and 75%), you might expect the
# max value of total_rooms to be approximately
# 5,000 or possibly 10,000. However, the max value
# is actually 37,937.
# When you see anomalies in a column, become more careful
# about using that column as a feature. That said,
# anomalies in potential features sometimes mirror
# anomalies in the label, which could make the column
# be (or seem to be) a powerful feature.
# Also, as you will see later in the course, you
# might be able to represent (pre-process) raw data
# in order to make columns into useful features.
# + [markdown] id="3014ezH3C7jT" colab_type="text"
# ## Define functions that build and train a model
#
# The following code defines two functions:
#
# * `build_model(my_learning_rate)`, which builds a randomly-initialized model.
# * `train_model(model, feature, label, epochs)`, which trains the model from the examples (feature and label) you pass.
#
# Since you don't need to understand model building code right now, we've hidden this code cell. You may optionally double-click the following headline to see the code that builds and trains a model.
# + id="pedD5GhlDC-y" colab_type="code" cellView="form" colab={}
#@title Define the functions that build and train a model
def build_model(my_learning_rate):
"""Create and compile a simple linear regression model."""
# Most simple tf.keras models are sequential.
model = tf.keras.models.Sequential()
# Describe the topography of the model.
# The topography of a simple linear regression model
# is a single node in a single layer.
model.add(tf.keras.layers.Dense(units=1,
input_shape=(1,)))
# Compile the model topography into code that TensorFlow can efficiently
# execute. Configure training to minimize the model's mean squared error.
model.compile(optimizer=tf.keras.optimizers.RMSprop(lr=my_learning_rate),
loss="mean_squared_error",
metrics=[tf.keras.metrics.RootMeanSquaredError()])
return model
def train_model(model, df, feature, label, epochs, batch_size):
"""Train the model by feeding it data."""
# Feed the model the feature and the label.
# The model will train for the specified number of epochs.
history = model.fit(x=df[feature],
y=df[label],
batch_size=batch_size,
epochs=epochs)
# Gather the trained model's weight and bias.
trained_weight = model.get_weights()[0]
trained_bias = model.get_weights()[1]
# The list of epochs is stored separately from the rest of history.
epochs = history.epoch
# Isolate the error for each epoch.
hist = pd.DataFrame(history.history)
# To track the progression of training, we're going to take a snapshot
# of the model's root mean squared error at each epoch.
rmse = hist["root_mean_squared_error"]
return trained_weight, trained_bias, epochs, rmse
print("Defined the create_model and traing_model functions.")
# + [markdown] id="Ak_TMAzGOIFq" colab_type="text"
# ## Define plotting functions
#
# The following [matplotlib](https://developers.google.com/machine-learning/glossary/#matplotlib) functions create the following plots:
#
# * a scatter plot of the feature vs. the label, and a line showing the output of the trained model
# * a loss curve
#
# You may optionally double-click the headline to see the matplotlib code, but note that writing matplotlib code is not an important part of learning ML programming.
# + id="QF0BFRXTOeR3" colab_type="code" cellView="form" colab={}
#@title Define the plotting functions
def plot_the_model(trained_weight, trained_bias, feature, label):
"""Plot the trained model against 200 random training examples."""
# Label the axes.
plt.xlabel(feature)
plt.ylabel(label)
# Create a scatter plot from 200 random points of the dataset.
random_examples = training_df.sample(n=200)
plt.scatter(random_examples[feature], random_examples[label])
# Create a red line representing the model. The red line starts
# at coordinates (x0, y0) and ends at coordinates (x1, y1).
x0 = 0
y0 = trained_bias
x1 = 10000
y1 = trained_bias + (trained_weight * x1)
plt.plot([x0, x1], [y0, y1], c='r')
# Render the scatter plot and the red line.
plt.show()
def plot_the_loss_curve(epochs, rmse):
"""Plot a curve of loss vs. epoch."""
plt.figure()
plt.xlabel("Epoch")
plt.ylabel("Root Mean Squared Error")
plt.plot(epochs, rmse, label="Loss")
plt.legend()
plt.ylim([rmse.min()*0.97, rmse.max()])
plt.show()
print("Defined the plot_the_model and plot_the_loss_curve functions.")
# + [markdown] id="D-IXYVfvM4gD" colab_type="text"
# ## Call the model functions
#
# An important part of machine learning is determining which [features](https://developers.google.com/machine-learning/glossary/#feature) correlate with the [label](https://developers.google.com/machine-learning/glossary/#label). For example, real-life home-value prediction models typically rely on hundreds of features and synthetic features. However, this model relies on only one feature. For now, you'll arbitrarily use `total_rooms` as that feature.
#
# + id="nj3v5EKQFY8s" colab_type="code" cellView="both" colab={}
# The following variables are the hyperparameters.
learning_rate = 0.01
epochs = 30
batch_size = 30
# Specify the feature and the label.
my_feature = "total_rooms" # the total number of rooms on a specific city block.
my_label="median_house_value" # the median value of a house on a specific city block.
# That is, you're going to create a model that predicts house value based
# solely on total_rooms.
# Discard any pre-existing version of the model.
my_model = None
# Invoke the functions.
my_model = build_model(learning_rate)
weight, bias, epochs, rmse = train_model(my_model, training_df,
my_feature, my_label,
epochs, batch_size)
print("\nThe learned weight for your model is %.4f" % weight)
print("The learned bias for your model is %.4f\n" % bias )
plot_the_model(weight, bias, my_feature, my_label)
plot_the_loss_curve(epochs, rmse)
# + [markdown] id="Btp8zUNbYOcd" colab_type="text"
# A certain amount of randomness plays into training a model. Consequently, you'll get different results each time you train the model. That said, given the dataset and the hyperparameters, the trained model will generally do a poor job describing the feature's relation to the label.
# + [markdown] id="1xNqWWos_zyk" colab_type="text"
# ## Use the model to make predictions
#
# You can use the trained model to make predictions. In practice, [you should make predictions on examples that are not used in training](https://developers.google.com/machine-learning/crash-course/training-and-test-sets/splitting-data). However, for this exercise, you'll just work with a subset of the same training dataset. A later Colab exercise will explore ways to make predictions on examples not used in training.
#
# First, run the following code to define the house prediction function:
# + id="nH63BmncAcab" colab_type="code" colab={}
def predict_house_values(n, feature, label):
"""Predict house values based on a feature."""
batch = training_df[feature][10000:10000 + n]
predicted_values = my_model.predict_on_batch(x=batch)
print("feature label predicted")
print(" value value value")
print(" in thousand$ in thousand$")
print("--------------------------------------")
for i in range(n):
print ("%5.0f %6.0f %15.0f" % (training_df[feature][i],
training_df[label][i],
predicted_values[i][0] ))
# + [markdown] id="NbBNQujU5WjK" colab_type="text"
# Now, invoke the house prediction function on 10 examples:
# + id="Y_0DGBt0Kz_N" colab_type="code" colab={}
predict_house_values(10, my_feature, my_label)
# + [markdown] id="-gGaqArcpqY3" colab_type="text"
# ### Task 2: Judge the predictive power of the model
#
# Look at the preceding table. How close is the predicted value to the label value? In other words, does your model accurately predict house values?
# + id="yVpjhUFm9uID" colab_type="code" cellView="form" colab={}
#@title Double-click to view the answer.
# Most of the predicted values differ significantly
# from the label value, so the trained model probably
# doesn't have much predictive power. However, the
# first 10 examples might not be representative of
# the rest of the examples.
# + [markdown] id="wLoqis3IUPSd" colab_type="text"
# ## Task 3: Try a different feature
#
# The `total_rooms` feature had only a little predictive power. Would a different feature have greater predictive power? Try using `population` as the feature instead of `total_rooms`.
#
# Note: When you change features, you might also need to change the hyperparameters.
# + id="H0ab6HD4ZO75" colab_type="code" colab={}
my_feature = "population" # Replace the ? with population or possibly
# a different column name.
# Experiment with the hyperparameters.
learning_rate = 0.05
epochs = 18
batch_size = 3
# Don't change anything below this line.
my_model = build_model(learning_rate)
weight, bias, epochs, rmse = train_model(my_model, training_df,
my_feature, my_label,
epochs, batch_size)
plot_the_model(weight, bias, my_feature, my_label)
plot_the_loss_curve(epochs, rmse)
predict_house_values(15, my_feature, my_label)
# + id="107mDkW7U6mg" colab_type="code" cellView="both" colab={}
#@title Double-click to view a possible solution.
my_feature = "population" # Pick a feature other than "total_rooms"
# Possibly, experiment with the hyperparameters.
learning_rate = 0.05
epochs = 18
batch_size = 3
# Don't change anything below.
my_model = build_model(learning_rate)
weight, bias, epochs, rmse = train_model(my_model, training_df,
my_feature, my_label,
epochs, batch_size)
plot_the_model(weight, bias, my_feature, my_label)
plot_the_loss_curve(epochs, rmse)
predict_house_values(10, my_feature, my_label)
# + [markdown] id="Nd_rHJ59AUtk" colab_type="text"
# Did `population` produce better predictions than `total_rooms`?
# + id="F0tPEtzcC-vK" colab_type="code" cellView="both" colab={}
#@title Double-click to view the answer.
# Training is not entirely deterministic, but population
# typically converges at a slightly higher RMSE than
# total_rooms. So, population appears to be about
# the same or slightly worse at making predictions
# than total_rooms.
# + [markdown] id="C8uYpyGacsIg" colab_type="text"
# ## Task 4: Define a synthetic feature
#
# You have determined that `total_rooms` and `population` were not useful features. That is, neither the total number of rooms in a neighborhood nor the neighborhood's population successfully predicted the median house price of that neighborhood. Perhaps though, the *ratio* of `total_rooms` to `population` might have some predictive power. That is, perhaps block density relates to median house value.
#
# To explore this hypothesis, do the following:
#
# 1. Create a [synthetic feature](https://developers.google.com/machine-learning/glossary/#synthetic_feature) that's a ratio of `total_rooms` to `population`. (If you are new to pandas DataFrames, please study the [Pandas DataFrame Ultraquick Tutorial](https://colab.research.google.com/github/google/eng-edu/blob/master/ml/cc/exercises/pandas_dataframe_ultraquick_tutorial.ipynb?utm_source=linearregressionreal-colab&utm_medium=colab&utm_campaign=colab-external&utm_content=pandas_tf2-colab&hl=en).)
# 2. Tune the three hyperparameters.
# 3. Determine whether this synthetic feature produces
# a lower loss value than any of the single features you
# tried earlier in this exercise.
# + id="4Kx2xHSgdcpg" colab_type="code" colab={}
# Define a synthetic feature named rooms_per_person
training_df["rooms_per_person"] = training_df['total_rooms'] / training_df['population'] # write your code here.
# Don't change the next line.
my_feature = "rooms_per_person"
# Assign values to these three hyperparameters.
learning_rate = 0.06
epochs = 24
batch_size = 30
# Don't change anything below this line.
my_model = build_model(learning_rate)
weight, bias, epochs, rmse = train_model(my_model, training_df,
my_feature, my_label,
epochs, batch_size)
plot_the_loss_curve(epochs, rmse)
predict_house_values(15, my_feature, my_label)
# + id="xRfxp_3yofe3" colab_type="code" cellView="both" colab={}
#@title Double-click to view a possible solution to Task 4.
# Define a synthetic feature
training_df["rooms_per_person"] = training_df["total_rooms"] / training_df["population"]
my_feature = "rooms_per_person"
# Tune the hyperparameters.
learning_rate = 0.06
epochs = 24
batch_size = 30
# Don't change anything below this line.
my_model = build_model(learning_rate)
weight, bias, epochs, mae = train_model(my_model, training_df,
my_feature, my_label,
epochs, batch_size)
plot_the_loss_curve(epochs, mae)
predict_house_values(15, my_feature, my_label)
# + [markdown] id="HBiDWursB1Wi" colab_type="text"
# Based on the loss values, this synthetic feature produces a better model than the individual features you tried in Task 2 and Task 3. However, the model still isn't creating great predictions.
#
# + [markdown] id="XEG_9oU9O54u" colab_type="text"
# ## Task 5. Find feature(s) whose raw values correlate with the label
#
# So far, we've relied on trial-and-error to identify possible features for the model. Let's rely on statistics instead.
#
# A **correlation matrix** indicates how each attribute's raw values relate to the other attributes' raw values. Correlation values have the following meanings:
#
# * `1.0`: perfect positive correlation; that is, when one attribute rises, the other attribute rises.
# * `-1.0`: perfect negative correlation; that is, when one attribute rises, the other attribute falls.
# * `0.0`: no correlation; the two column's [are not linearly related](https://en.wikipedia.org/wiki/Correlation_and_dependence#/media/File:Correlation_examples2.svg).
#
# In general, the higher the absolute value of a correlation value, the greater its predictive power. For example, a correlation value of -0.8 implies far more predictive power than a correlation of -0.2.
#
# The following code cell generates the correlation matrix for attributes of the California Housing Dataset:
# + id="zFGKL45LO8Tt" colab_type="code" colab={}
# Generate a correlation matrix.
training_df.corr()
# + [markdown] id="hp0r3NAVPEdt" colab_type="text"
# The correlation matrix shows nine potential features (including a synthetic
# feature) and one label (`median_house_value`). A strong negative correlation or strong positive correlation with the label suggests a potentially good feature.
#
# **Your Task:** Determine which of the nine potential features appears to be the best candidate for a feature?
# + id="RomQTd1OPVd0" colab_type="code" cellView="both" colab={}
#@title Double-click here for the solution to Task 5
# The `median_income` correlates 0.7 with the label
# (median_house_value), so median_income` might be a
# good feature. The other seven potential features
# all have a correlation relatively close to 0.
# If time permits, try median_income as the feature
# and see whether the model improves.
# + [markdown] id="8RqvEbaVSlRt" colab_type="text"
# Correlation matrices don't tell the entire story. In later exercises, you'll find additional ways to unlock predictive power from potential features.
#
# **Note:** Using `median_income` as a feature may raise some ethical and fairness
# issues. Towards the end of the course, we'll explore ethical and fairness issues.
| Copy_of_Linear_Regression_with_a_Real_Dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Convolutional Variational Autoencoder example using MNIST dataset
# ## Import the required libraries
# +
import math
import tempfile
import os
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
import fastestimator as fe
from fastestimator.estimator.trace import ModelSaver
from fastestimator.network.loss import Loss
from fastestimator.network.model import FEModel, ModelOp
from fastestimator.util.op import TensorOp
# -
# ## Introduction to CVAE
# **Introduction**
#
# CVAE are Convolutional Variational Autoencoder. They are composed by two models using convolutions: an Encoder to represent the input in a latent dimension, and a Decoder that will generate data from the latent dimension to the input space. The figure below illustrates the main idea of CVAE. In this example, we will use a CVAE to generate data similar to MNIST dataset.
#
#
#
# **Goal**
#
# The main goal of VAEs is to optimize the likelihood of the real data according to the generative model. In other words, maximize $\mathbb{E}_{p_D} \log p_\theta(x)$ where $p_D$ is the real distribution, $x$ real data (observation) and $p_\theta$ corresponds to the generated distribution. We will denote z the latent representation of x.
#
#
# **Encoder**
#
# The encoder is responsible for the posterior distribution ($q(z|x)$). It takes as input an observation of the real data and outputs parameters (mu and logvar, log is used for stability) of a Gaussian distribution (the conditional distribution of the latent representation), which will be used to sample noisy representations (thanks to reparametrization) of the input in the latent space. We set a Gaussian prior on p(z).
#
#
# **Decoder**
#
# The decoder, a generative model, will take the latent representation as input and outputs the parameters for a conditional distribution of the observation ($p(x|z)$) , from which we'll obtain an image that should resemble our initial input.
# 
#
# [Source: Machine Learning Explained - An Intuitive Explanation of Variational Autoencoders (VAEs Part 1)]
# ## Step 1 - Data and pipeline preparation
# In this step, we will load MNIST training and validation datasets and prepare FastEstimator's pipeline.
# ### Dataset
# +
# Load dataset
(x_train, _), (x_eval, _) = tf.keras.datasets.mnist.load_data()
x_train = x_train.reshape(x_train.shape[0], 28, 28, 1).astype('float32')
x_eval = x_eval.reshape(x_eval.shape[0], 28, 28, 1).astype('float32')
plt.imshow(x_train[0,:,:,0], cmap='gray')
plt.title('Image example')
plt.show()
# -
# ### Preprocessing
# +
# Define custom preprocessing functions for the Pipeline
# We create a rescaling Tensor Operation.
class Myrescale(TensorOp):
"""Normalize pixel values from uint8 to float32 between 0 and 1."""
def forward(self, data, state):
data = tf.cast(data, tf.float32)
data = data / 255
return data
# We create a Tensor Operation to binarize the input.
class Mybinarize(TensorOp):
"""Pixel values assigned to either 0 or 1 """
def forward(self, data, state):
data = tf.where(data >= 0.5, 1., 0.)
return data
# -
# ### Pipeline
# Now, we can compile the whole data preparation process into FE pipeline, indicating the batch size, the data and preprocessing ops.
# +
# Prepare pipeline for a specific batch size
batch_size=100
# We indicate in a dictionary data names for each dataset and the corresponding array.
data = {"train": {"x": x_train}, "eval": {"x": x_eval}}
# Pipeline compiles batch size, datasets and preprocessing
pipeline = fe.Pipeline(batch_size=batch_size,
data=data,
ops=[Myrescale(inputs="x", outputs="x"), Mybinarize(inputs="x", outputs="x")])
# -
# ## Step 2 - Models definition
# In this step, we will create and compile the two models building our CVAE : an Encoder and a Decoder. The Encoder is a Convolutional Neural Network and the Decoder uses Deconvolutional layers. The user needs to define the latent dimension of the hidden space.
# Parameters
LATENT_DIM = 50
# ### Encoder
# The Encoder outputs mean and log(variance) (log for stability) of the data distribution in the latent dimension. We create a simple CNN with two convolutional layers and a fully connected layer at the end.
# As the two parameters are concatenated by the fully connected layer, we define a Tensor Operation to split the Encoder output in two Tensors.
# +
# Encoder network
def encoder_net():
encoder_model = tf.keras.Sequential()
encoder_model.add(tf.keras.layers.InputLayer(input_shape=(28, 28, 1)))
encoder_model.add(tf.keras.layers.Conv2D(filters=32, kernel_size=3, strides=(2, 2), activation='relu'))
encoder_model.add(tf.keras.layers.Conv2D(filters=64, kernel_size=3, strides=(2, 2), activation='relu'))
encoder_model.add(tf.keras.layers.Flatten())
encoder_model.add(tf.keras.layers.Dense(LATENT_DIM + LATENT_DIM))
return encoder_model
# The encoder network outputs a flatten array concatenating mean and logvar,we define SplitOp to split them.
class SplitOp(TensorOp):
"""To split the infer net output into two """
def forward(self, data, state):
mean, logvar = tf.split(data, num_or_size_splits=2, axis=1)
return mean, logvar
# -
# ### Decoder
#
# The Decoder is the generative part of the model. It takes as input the representation of the input in the latent space and outputs a reconstructed image. The Encoder takes as input the latent representation of the image and mirrors the Encoder. Hence, the first layer is a fully connected layer of size 7x7x32. It is then composed of two Conv2DTranspose layers (also called Deconvolutional layers).
# Decoder model
def generative_net():
generative_model = tf.keras.Sequential()
generative_model.add(tf.keras.layers.InputLayer(input_shape=(LATENT_DIM, )))
generative_model.add(tf.keras.layers.Dense(units=7 * 7 * 32, activation=tf.nn.relu))
generative_model.add(tf.keras.layers.Reshape(target_shape=(7, 7, 32)))
generative_model.add(
tf.keras.layers.Conv2DTranspose(filters=64, kernel_size=3, strides=(2, 2), padding="SAME", activation='relu'))
generative_model.add(
tf.keras.layers.Conv2DTranspose(filters=32, kernel_size=3, strides=(2, 2), padding="SAME", activation='relu'))
generative_model.add(tf.keras.layers.Conv2DTranspose(filters=1, kernel_size=3, strides=(1, 1), padding="SAME"))
return generative_model
# ### FEModels
# We will finally compile the models thanks to FEModel and use an Adam optimizer with a learning rate of 1e-4.
# +
# Prepare/Compile models
enc_model = FEModel(model_def=encoder_net,
model_name="encoder",
loss_name="loss",
optimizer=tf.optimizers.Adam(1e-4))
gen_model = FEModel(model_def=generative_net,
model_name="decoder",
loss_name="loss",
optimizer=tf.optimizers.Adam(1e-4))
# -
# ## Step 3 - Network definition
# In this step, we build the network, summarizing all operations of our model that are pictured in the graph above. We will first need to define the reparametrization and loss.
# ### Reparametrization to sample from encoder's output
# Reparametrization is the trick used to sample from our parameters with operations allowing backpropagation. We will generate a white noise vector epsilon (Gaussian distribution N(0,1)) whose length is the latent dimension, multiply it with standard deviation and add the mean.
# (See Auto-Encoding Variational Bayes by Kingma and Welling)
# +
# We use reparametrization to sample from mean and logvar.
class ReparameterizepOp(TensorOp):
"""Reparameterization trick. Ensures gradients pass through the sample to the encoder parameters"""
def forward(self, data, state):
mean, logvar = data
eps = tf.random.normal(shape=mean.shape)
return eps * tf.exp(logvar * .5) + mean
# -
# ### CVAE loss
# To train a VAE, we have to maximize the evidence lower bound (ELBO) on the marginal log-likelihood $log p(x)$.
# In this example, we approximate the CVAE loss ELBO (evidence lower bound) with the Monte Carlo estimate:
# $log p(x|z) + log p(z) - log q(z|x)$ with $z$ sampled from $q(z|x)$.
# We obtain a loss by changing the sign.
# +
# The log normal pdf function will be used in the loss.
def _log_normal_pdf(sample, mean, logvar, raxis=1):
log2pi = tf.math.log(2. * tf.constant(math.pi))
return tf.reduce_sum(-.5 * ((sample - mean)**2. * tf.exp(-logvar) + logvar + log2pi), axis=raxis)
class CVAELoss(Loss):
"""Convolutional variational auto-encoder loss"""
def forward(self, data, state):
x, mean, logvar, z, x_logit = data
cross_ent = tf.nn.sigmoid_cross_entropy_with_logits(logits=x_logit, labels=x)
logpx_z = -tf.reduce_sum(cross_ent, axis=[1, 2, 3])
logpz = _log_normal_pdf(z, 0., 0.)
logqz_x = _log_normal_pdf(z, mean, logvar)
# Changing the sign to obtain a loss
return -(logpx_z + logpz - logqz_x)
# -
# ### Network
# In Network, we summarize all operations and the loss (which has by definition outputs = "loss").
# +
# Defining FE network
network = fe.Network(ops=[
ModelOp(inputs="x", model=enc_model, outputs="meanlogvar", mode=None),
SplitOp(inputs="meanlogvar", outputs=("mean", "logvar"), mode=None),
ReparameterizepOp(inputs=("mean", "logvar"), outputs="z", mode=None),
ModelOp(inputs="z", model=gen_model, outputs="x_logit"),
CVAELoss(inputs=("x", "mean", "logvar", "z", "x_logit"), mode=None)
])
# -
# ## Step 4 - Estimator definition and training
# In this step, we define the estimator to compile the network and pipeline and indicate in traces that we want to save the best models.
# +
# Parameters
epochs=100
model_dir=tempfile.mkdtemp()
# Estimator
estimator = fe.Estimator(network=network,
pipeline=pipeline,
epochs=epochs,
traces=[ModelSaver(model_name="decoder", save_dir=model_dir, save_best=True),
ModelSaver(model_name="encoder", save_dir=model_dir, save_best=True)])
# -
# We simply call the method fit to launch the training. Logs display for each step the loss and number of examples per second to evaluate the training speed.
# Training
estimator.fit()
# ## Step 5 - Inferencing
#
# After training, the model is saved to a temporary folder. we can load the model from file and do inferencing on a sample image.
# You can also access the model via estimator.network.model['encoder'] .
# ### Load the models
model_path = os.path.join(model_dir, 'decoder_best_loss.h5')
trained_decoder = tf.keras.models.load_model(model_path, compile=False)
model_path = os.path.join(model_dir, 'encoder_best_loss.h5')
trained_encoder = tf.keras.models.load_model(model_path, compile=False)
# ### Select an image
# +
# Select a random image from the validation dataset
selected_idx = np.random.randint(len(x_eval))
# Normalize and reshape the image
img = x_eval[selected_idx] / 255
img = np.where(img>= 0.5, 1., 0.)
img = np.expand_dims(img, 0)
print("test image idx {}".format(selected_idx))
# -
# ### Generate an image
# +
# Generate an image following the network steps
mulogvar = trained_encoder(img)
mean, logvar = np.split(mulogvar,2, axis=1)
eps = tf.random.normal(shape=mean.shape)
reparametrized_img = eps * tf.exp(logvar * .5) + mean
gen_img = trained_decoder(reparametrized_img)
bin_gen_img=np.where(gen_img>= 0.5, 1., 0.)
# -
# ### Plotting results
# +
# Plot the results
fig, ax = plt.subplots(2,2, figsize=(12,12))
ax[0,0].imshow(x_eval[selected_idx, :, :, 0], cmap='gray')
ax[0,0].set_title('Normal image')
ax[1,0].imshow(img[0, :, :, 0], cmap='gray')
ax[1,0].set_title('Binarized image')
ax[0,1].imshow(gen_img[0,:,:,0], cmap='gray')
ax[0,1].set_title('Normal generated image')
ax[1,1].imshow(bin_gen_img[0,:,:,0], cmap='gray')
ax[1,1].set_title('Binarized generated image')
plt.show()
# -
| apphub/image_generation/cvae_mnist/cvae_mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Earnings This Week
# ### Includes: Every Company Reporting Earnings this Week
import numpy as np
import pandas as pd
import datetime as dt
import yahoo_fin.stock_info as si
# +
### EARNINGS FOR THIS WEEK INFORMATION
def get_earnings_fordates(startdate, enddate):
# Earnings Data in DataFrame Format
earnings_this_week = si.get_earnings_in_date_range(startdate, enddate)
earnings_thiswk = pd.DataFrame(earnings_this_week)
# Retreives Week Number
today = dt.date.today()
weeknumber = today.isocalendar()[1]
# Write to CSV file in 'CSVfiles' folder
dropcols = ['startdatetimetype', 'timeZoneShortName', 'gmtOffsetMilliSeconds' ,'quoteType','epsactual','epssurprisepct']
editearningsdf = earnings_thiswk.drop(columns=dropcols)
return editearningsdf
# Calculates the input dates for the week (release every sunday)
def this_weeks_earnings():
today = dt.date.today()
one_week = dt.timedelta(days=7)
one_week_ahead = today + one_week
today = str(today)
one_week = str(one_week_ahead)
# Call earnings for week function
earningsDF = get_earnings_fordates(today, one_week)
return earningsDF
this_weeks_earnings()
# -
| ThisWeekEarnings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Export and Import from/to Brightway2 example
# ### Import the `DisclosureExporter`
from lca_disclosures.brightway2.disclosure import Bw2Disclosure as DisclosureExporter
# Give it a project name and database name to export
bw_project_name = "Example Model"
bw_db_name = "Example Model_Database"
de = DisclosureExporter(bw_project_name, bw_db_name, folder_path="exporter_testing")
# Write the file and store the path
disclosure_file = de.write_json()
# ### Import brightway and select the project to import into
from brightway2 import *
projects.set_current("CoTDisclosure")
# ### Import the `DisclosureImporter`
from lca_disclosures.brightway2.importer import DisclosureImporter
# Give it a path and (optionally) a name to give the database
di = DisclosureImporter(disclosure_file, bw_db_name)
# Apply strategies to resolve links
di.apply_strategies()
# If everything is ok, write the database
if di.statistics()[2] == 0:
di.write_database()
# Do an LCA to check it worked
fu = {Database(di.db_name).search('tea')[0]: 1}
m = ('IPCC 2013', 'climate change', 'GWP 100a')
lca = LCA(fu, m)
lca.lci()
lca.lcia()
print(lca.score)
| examples/Example usage with Brightway2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # (7) Signal (CPD) Search and Characterization
# In Part (6), we learned that any CPDs of interest in the SR 4 disk should be point sources (i.e., their size is $\ll$ the resolution) and could be pretty faint, perhaps comparable to the residuals from the circumstellar disk model. We need to develop a means of quantifying what level of emission we are able to robustly detect. One intuitive way for doing that is to inject a fake signal into the data, perform the same fitting/post-processing analysis we used on the real observations to get a residual dataset, and then try to recover the fake signal.
#
# For the last part, *recovery*, there is not a widely agreed-upon metric. We'll have to develop our own approach and demonstrate that it works in practice. Your goal in this part of the project is to establish an **automated** way to search for and quantify a point source in the SR 4 disk gap. This machinery should (1) quantify the *significance* of any such feature (i.e., its signal-to-noise ratio, or the ratio of the peak to the "local" standard deviation); and (2) measure its flux; and (3) measure its position in the SR 4 disk gap (given how narrow the gap is, the radius should be just about 0.08", but the *azimuth* is unknown a priori). There are no right or wrong answers here: your job is to experiment and see what might work. Start simple...if we can make it work with a straightforward search in a (r,az)-map like before, let's do that!
#
# I know this is sort of backwards, worrying about the recovery part before the injection part. But I think it makes more sense in terms of the work that needs to be done. The *injection* doesn't involve a lot of activity from a research perspective (its more of just a code machinery in a big loop). I will show you how that works once your machinery is tested. To help you develop, I've posted two example residual images online (see below) with mock CPDs already injected.
# +
# your code here
# -
# Sometime later this week I'll point you to some different models where the mock CPDs should be much harder to find and characterize. Your method will need to handle such cases also!
| notebooks/Part7_SignalDetection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example of Liftingline Analysis
# +
# numpy and matplotlib imports
import numpy as np
from matplotlib import pyplot as plt
# import of wingstructure submodels
from wingstructure import data, aero
# -
# ## Creating wing defintion
# +
# create wing object
wing = data.Wing()
# add sections to wing
# leading edge position, chord length, twist
wing.append((0.0, 0.0, 0.0), 1.0, 0.0)
wing.append((0.05, 4.25, 0.0), 0.7, 0.0)
wing.append((0.1, 7.75, 0.0), 0.35, 0.0)
# define spoiler position
wing.add_controlsurface('BK', 1.5, 2.9, 0.5, 0.5, 'airbrake')
# define control-surfaces
wing.add_controlsurface('flap', 1, 2.8, 0.3, 0.3, 'aileron')
wing.add_controlsurface('flap2', 4.25, 7, 0.3, 0.2, 'aileron')
wing.plot()
# -
# ## Lift calculation using LiftAnalysis object
#
# The LiftAnalysis object calculates base lift distributions (e.q. for aerodynamical twist, control surfaces and so on) and only superposes those, when calculations are invoked.
# +
liftana = aero.LiftAnalysis.generate(wing)
span_pos = liftana.ys
α, distribution, C_Dib, C_Mxb = liftana.calculate(C_L=0.8, all_results=True)
α_qr, distribution_q, C_Dia, C_Mxa = liftana.calculate(C_L=0.8,
controls={'flap2': [5, -5]}, all_results=True)
α_ab, distribution_ab, C_Di, C_Mx = liftana.calculate(C_L=0.8, airbrake=True,
all_results=True)
plt.figure(figsize=(8,5))
plt.plot(span_pos, distribution, label='clean')
plt.plot(span_pos, distribution_ab, '--', label='airbrakes')
plt.plot(span_pos, distribution_q, '-.', label='flaps')
plt.xlabel('wing span in m')
plt.ylabel('local lift coefficient $c_l$')
plt.title('Lift distribution for $C_L = 0,8$')
plt.grid()
plt.legend()
plt.savefig('Liftdistribution.png')
plt.savefig('Liftdistribution.pdf')
# -
# ## Lift calculation using calculate function
#
# The calculate function does only calculate those distributions needed and does not cache results. Furhtermore it allows for calculation of moment coefficent regarding x axis (flight direction). This coefficient is defined as follows:
#
# $$ C_\mathrm{Mx} = \frac{M_\mathrm{x}}{q S b}.$$
#
# $q$ - dynamic pressure
#
# $S$ - wing surface
#
# $b$ - wing span
#
aero.calculate(wing, target=1.0, controls={'flap':(5,-5)}, calc_cmx=True)
| examples/02_Analysis_Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/abbaasalif/YOLO_custom/blob/main/custom_YOLO_training_in_Colab.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="sNgMH7RX9dNG"
# !mount --bind /content/drive/My\ Drive /content/MyDrive
# + id="SjSh4W65EWcp" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="068adaf7-fcc5-471d-b97a-36d0d529ddb0"
# %cd MyDrive/
# + id="uQK3aq-4E-bX" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="74b18d9b-c8a7-48c0-e2ea-31246a1b1b9e"
# %cd darknet/
# + id="W89wdtNOFDaq" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0e306dc4-79ef-4edc-cfb3-9aac5c5f60d6"
# !make
# + id="detMc7vQF5NM" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="b63ad924-0a1a-47a5-9255-dd22c4d9c97e"
# !./darknet detector train cfg/ambulance.data cfg/ambulance-yolo.cfg backup/ambulance-yolo.backup
# + id="QqTr5USNJDja"
# !nvidia-smi
# + id="7Y-RngrOJpuy" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="098ba5c5-8407-48e8-f8f2-4b4f5cab291f"
# !chmod -u +x ./darknet
# + id="0xOlBgSGKGxJ"
# !chmod 755 ./darknet
# + id="mvA-UDHyKK_M"
| custom_YOLO_training_in_Colab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
import numpy as np
import tensorflow as tf
from tensorflow.keras.applications.xception import Xception, preprocess_input
from sklearn import metrics
from sklearn.metrics import confusion_matrix, classification_report, roc_curve
from sklearn.preprocessing import LabelBinarizer
from sklearn.utils import class_weight
import keras
from keras import backend as K
from keras.models import Sequential, Model
from keras.layers import Input,Dense, Dropout, Activation, Flatten, Convolution2D, Conv2D, MaxPooling2D, GlobalAveragePooling2D
from keras.layers.core import Dropout
from keras.layers.normalization import BatchNormalization
from keras.preprocessing.image import ImageDataGenerator
from keras.optimizers import Adam
from keras.utils import np_utils
from keras.regularizers import l2
from keras.callbacks import EarlyStopping, ModelCheckpoint
import matplotlib.pyplot as plt
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3), activation='relu', input_shape=(288, 288, 3)))
model.add(Conv2D(64, kernel_size=(3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(64, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(5, activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
print(model.summary())
early_stopping = EarlyStopping(monitor='val_loss', patience=50, verbose=1)
callbacks = [early_stopping]
train_datagen = ImageDataGenerator(preprocessing_function=preprocess_input,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.1,
fill_mode='nearest',
horizontal_flip = False)
test_datagen = ImageDataGenerator(preprocessing_function=preprocess_input)
training_set = train_datagen.flow_from_directory('data/19_class2/train',
target_size = (288, 288),
batch_size = 16,
class_mode = 'categorical', shuffle=True)
test_set = test_datagen.flow_from_directory('data/19_class2/test',
target_size = (288, 288),
batch_size = 16,
class_mode = 'categorical')
val_set = test_datagen.flow_from_directory('data/19_class2/val',
target_size = (288, 288),
batch_size = 16,
class_mode = 'categorical')
class_weights = class_weight.compute_class_weight(
'balanced',
np.unique(training_set.classes),
training_set.classes)
model.fit_generator(training_set, epochs=10, class_weight=class_weights, validation_data = test_set, verbose=True, callbacks=callbacks, shuffle=True)
model.save('MK_19class2.h5')
Y_pred = model.predict_generator(val_set)
y_pred = np.argmax(Y_pred, axis=1)
print(classification_report(val_set.classes, y_pred))
score = model.evaluate_generator(training_set, verbose=0)
print("Training Accuracy: ", score[1])
score = model.evaluate_generator(test_set, verbose=0)
print("Testing Accuracy: ", score[1])
score = model.evaluate_generator(val_set, verbose=0)
print("Validating Accuracy: ", score[1])
| Sound-based-bird-species-detection-master/MK_neural_network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext ipyext.writeandexecute
# +
# %%writeandexecute -i alkylsilane alkylsilane.py
import mbuild as mb
from mbuild.examples.alkane.alkane import Alkane
from mbuild.lib.moieties import Silane
class AlkylSilane(mb.Compound):
"""A silane functionalized alkane chain with one Port. """
def __init__(self, chain_length):
super(AlkylSilane, self).__init__()
alkane = Alkane(chain_length, cap_end=False)
self.add(alkane, 'alkane')
silane = Silane()
self.add(silane, 'silane')
mb.force_overlap(self['alkane'], self['alkane']['down'], self['silane']['up'])
# Hoist silane port to AlkylSilane level.
self.add(silane['down'], 'down', containment=False)
# -
alkyl_silane = AlkylSilane(10)
print(alkyl_silane)
alkyl_silane.visualize()
| mbuild/examples/alkane_monolayer/alkylsilane.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.9 64-bit (''research_track-0UGwfk25'': venv)'
# language: python
# name: python3
# ---
# ## Cache Plots
# +
import matplotlib.pyplot as plt
from matplotlib import rcParams
import pandas as pd
import seaborn as sns
def plot(p_data, p_yId, p_xId, p_hueId, p_styleId, p_logScale=False, p_notiling_marker=False, p_notiling_value=0, p_export_filename=None, p_xLabel=None, p_yLabel=None):
rcParams['figure.figsize'] = 12,8
rcParams['font.size'] = 12
rcParams['svg.fonttype'] = 'none'
plot = sns.lineplot(x=p_xId,
y=p_yId,
hue=p_hueId,
style=p_styleId,
data=p_data)
if p_logScale == True:
plot.set_yscale('log')
plot.set_xscale('log')
if p_xLabel != None:
plot.set(xlabel=p_xLabel)
else:
plot.set(xlabel=p_xId)
if p_yLabel != None:
plot.set(ylabel=p_yLabel)
else:
plot.set(ylabel=p_yId)
plt.grid(color='gainsboro')
plt.grid(True,which='minor', linestyle='--', linewidth=0.5, color='gainsboro')
if(p_notiling_marker == True):
plt.axhline(p_notiling_value, linestyle='--', color='red', label='\Verb{notiling}')
plt.legend(title='TS_xz')
if(p_export_filename != None):
plt.savefig(p_export_filename)
plt.show()
# -
# ### Gauss3
# ### Runtime as a function of tiling
# +
import pandas as pd
data_frame = pd.read_csv('./e_runtime_tiling.csv')
data_frame = data_frame[data_frame.region_id == 'apply']
data_frame = data_frame[data_frame.impl_id == 'linear_stencil_noedgecases_tiling']
data_frame = data_frame[data_frame.tiling_cols < 65536]
data_frame = data_frame[data_frame.tiling_rows < 1024]
data_frame['tiling_rows'] = data_frame['tiling_rows'].map(str)
plot(p_data=data_frame,
p_yId='runtime',
p_xId='tiling_cols',
p_hueId='tiling_rows',
p_styleId=None,
p_logScale=True,
p_notiling_marker=True,
p_notiling_value=4.27924,
p_export_filename='runtime_tiling.svg',
p_xLabel="TS_y",
p_yLabel="Runtime [s]")
| src_optimization/08_openmp_stencil_03_noedgecases_02/e_plot_tiling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
#
# # Creating MNE objects from data arrays
#
#
# In this simple example, the creation of MNE objects from
# numpy arrays is demonstrated. In the last example case, a
# NEO file format is used as a source for the data.
#
#
# +
# Author: <NAME> <<EMAIL>>
#
# License: BSD (3-clause)
import numpy as np
import neo
import mne
print(__doc__)
# -
# Create arbitrary data
#
#
# +
sfreq = 1000 # Sampling frequency
times = np.arange(0, 10, 0.001) # Use 10000 samples (10s)
sin = np.sin(times * 10) # Multiplied by 10 for shorter cycles
cos = np.cos(times * 10)
sinX2 = sin * 2
cosX2 = cos * 2
# Numpy array of size 4 X 10000.
data = np.array([sin, cos, sinX2, cosX2])
# Definition of channel types and names.
ch_types = ['mag', 'mag', 'grad', 'grad']
ch_names = ['sin', 'cos', 'sinX2', 'cosX2']
# -
# Create an :class:`info <mne.Info>` object.
#
#
# It is also possible to use info from another raw object.
info = mne.create_info(ch_names=ch_names, sfreq=sfreq, ch_types=ch_types)
# Create a dummy :class:`mne.io.RawArray` object
#
#
# +
raw = mne.io.RawArray(data, info)
# Scaling of the figure.
# For actual EEG/MEG data different scaling factors should be used.
scalings = {'mag': 2, 'grad': 2}
raw.plot(n_channels=4, scalings=scalings, title='Data from arrays',
show=True, block=True)
# It is also possible to auto-compute scalings
scalings = 'auto' # Could also pass a dictionary with some value == 'auto'
raw.plot(n_channels=4, scalings=scalings, title='Auto-scaled Data from arrays',
show=True, block=True)
# -
# EpochsArray
#
#
# +
event_id = 1 # This is used to identify the events.
# First column is for the sample number.
events = np.array([[200, 0, event_id],
[1200, 0, event_id],
[2000, 0, event_id]]) # List of three arbitrary events
# Here a data set of 700 ms epochs from 2 channels is
# created from sin and cos data.
# Any data in shape (n_epochs, n_channels, n_times) can be used.
epochs_data = np.array([[sin[:700], cos[:700]],
[sin[1000:1700], cos[1000:1700]],
[sin[1800:2500], cos[1800:2500]]])
ch_names = ['sin', 'cos']
ch_types = ['mag', 'mag']
info = mne.create_info(ch_names=ch_names, sfreq=sfreq, ch_types=ch_types)
epochs = mne.EpochsArray(epochs_data, info=info, events=events,
event_id={'arbitrary': 1})
picks = mne.pick_types(info, meg=True, eeg=False, misc=False)
epochs.plot(picks=picks, scalings='auto', show=True, block=True)
# -
# EvokedArray
#
#
# +
nave = len(epochs_data) # Number of averaged epochs
evoked_data = np.mean(epochs_data, axis=0)
evokeds = mne.EvokedArray(evoked_data, info=info, tmin=-0.2,
comment='Arbitrary', nave=nave)
evokeds.plot(picks=picks, show=True, units={'mag': '-'},
titles={'mag': 'sin and cos averaged'})
# -
# Create epochs by windowing the raw data.
#
#
# +
# The events are spaced evenly every 1 second.
duration = 1.
# create a fixed size events array
# start=0 and stop=None by default
events = mne.make_fixed_length_events(raw, event_id, duration=duration)
print(events)
# for fixed size events no start time before and after event
tmin = 0.
tmax = 0.99 # inclusive tmax, 1 second epochs
# create :class:`Epochs <mne.Epochs>` object
epochs = mne.Epochs(raw, events=events, event_id=event_id, tmin=tmin,
tmax=tmax, baseline=None, verbose=True)
epochs.plot(scalings='auto', block=True)
# -
# Create overlapping epochs using :func:`mne.make_fixed_length_events` (50 %
# overlap). This also roughly doubles the amount of events compared to the
# previous event list.
#
#
duration = 0.5
events = mne.make_fixed_length_events(raw, event_id, duration=duration)
print(events)
epochs = mne.Epochs(raw, events=events, tmin=tmin, tmax=tmax, baseline=None,
verbose=True)
epochs.plot(scalings='auto', block=True)
# Extracting data from NEO file
#
#
# +
# The example here uses the ExampleIO object for creating fake data.
# For actual data and different file formats, consult the NEO documentation.
reader = neo.io.ExampleIO('fakedata.nof')
bl = reader.read(cascade=True, lazy=False)[0]
# Get data from first (and only) segment
seg = bl.segments[0]
title = seg.file_origin
ch_names = list()
data = list()
for ai, asig in enumerate(seg.analogsignals):
# Since the data does not contain channel names, channel indices are used.
ch_names.append('Neo %02d' % (ai + 1,))
# We need the ravel() here because Neo < 0.5 gave 1D, Neo 0.5 gives
# 2D (but still a single channel).
data.append(asig.rescale('V').magnitude.ravel())
data = np.array(data, float)
sfreq = int(seg.analogsignals[0].sampling_rate.magnitude)
# By default, the channel types are assumed to be 'misc'.
info = mne.create_info(ch_names=ch_names, sfreq=sfreq)
raw = mne.io.RawArray(data, info)
raw.plot(n_channels=4, scalings={'misc': 1}, title='Data from NEO',
show=True, block=True, clipping='clamp')
| 0.15/_downloads/plot_objects_from_arrays.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="1G_TVh1ybQkl"
# # eICU Collaborative Research Database
#
# # Notebook 4: Summary statistics
#
# This notebook shows how summary statistics can be computed for a patient cohort using the `tableone` package. Usage instructions for tableone are at: https://pypi.org/project/tableone/
#
# + [markdown] colab_type="text" id="L9XF77F2bnee"
# ## Load libraries and connect to the database
# + colab_type="code" id="wXiSE558bn_w" colab={}
# Import libraries
import numpy as np
import os
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import matplotlib.path as path
# Make pandas dataframes prettier
from IPython.display import display, HTML
# Access data using Google BigQuery.
from google.colab import auth
from google.cloud import bigquery
# + colab_type="code" id="pLGnLAy-bsKb" colab={}
# authenticate
auth.authenticate_user()
# + colab_type="code" id="PUjFDFdobszs" colab={}
# Set up environment variables
project_id='aarhus-critical-2019-team'
os.environ["GOOGLE_CLOUD_PROJECT"]=project_id
# + colab_type="code" id="bkJUF8HBbvWe" colab={}
# Helper function to read data from BigQuery into a DataFrame.
def run_query(query):
return pd.io.gbq.read_gbq(query, project_id=project_id, dialect="standard")
# + [markdown] colab_type="text" id="iWDUCA5Nb5BK"
# ## Install and load the `tableone` package
#
# The tableone package can be used to compute summary statistics for a patient cohort. Unlike the previous packages, it is not installed by default in Colab, so will need to install it first.
# + colab_type="code" id="F9doCgtscOJd" colab={}
# !pip install tableone
# + colab_type="code" id="SDI_Q7W0b4Le" colab={}
# Import the tableone class
from tableone import TableOne
# + [markdown] colab_type="text" id="14TU4lcrdD7I"
# ## Load the patient cohort
#
# In this example, we will load all data from the patient data, and link it to APACHE data to provide richer summary information.
# + colab_type="code" id="HF5WF5EObwfw" colab={}
# Link the patient and apachepatientresult tables on patientunitstayid
# using an inner join.
query = """
SELECT p.unitadmitsource, p.gender, p.age, p.ethnicity, p.admissionweight,
p.unittype, p.unitstaytype, a.acutephysiologyscore,
a.apachescore, a.actualiculos, a.actualhospitalmortality,
a.unabridgedunitlos, a.unabridgedhosplos
FROM `physionet-data.eicu_crd_demo.patient` p
INNER JOIN `physionet-data.eicu_crd_demo.apachepatientresult` a
ON p.patientunitstayid = a.patientunitstayid
WHERE apacheversion LIKE 'IVa'
"""
cohort = run_query(query)
# + colab_type="code" id="k3hURHFihHNA" colab={}
cohort.head()
# + [markdown] colab_type="text" id="qnG8dVb2iHSn"
# ## Calculate summary statistics
#
# Before summarizing the data, we will need to convert the ages to numerical values.
# + colab_type="code" id="oKHpqwAPkx6U" colab={}
cohort['agenum'] = pd.to_numeric(cohort['age'], errors='coerce')
# + colab_type="code" id="FQT-u8EXhXRG" colab={}
columns = ['unitadmitsource', 'gender', 'agenum', 'ethnicity',
'admissionweight','unittype','unitstaytype',
'acutephysiologyscore','apachescore','actualiculos',
'unabridgedunitlos','unabridgedhosplos']
# + colab_type="code" id="3ETr3NCzielL" colab={}
TableOne(cohort, columns=columns, labels={'agenum': 'age'},
groupby='actualhospitalmortality',
label_suffix=True, limit=4)
# + [markdown] colab_type="text" id="LCBcpJ9bZpDp"
# ## Questions
#
# - Are the severity of illness measures higher in the survival or non-survival group?
# - What issues suggest that some of the summary statistics might be misleading?
# - How might you address these issues?
# + [markdown] colab_type="text" id="2_8z1CIVahWg"
# ## Visualizing the data
#
# Plotting the distribution of each variable by group level via histograms, kernel density estimates and boxplots is a crucial component to data analysis pipelines. Vizualisation is often is the only way to detect problematic variables in many real-life scenarios. We'll review a couple of the variables.
# + colab_type="code" id="81yp2bSUigzh" colab={}
# Plot distributions to review possible multimodality
cohort[['acutephysiologyscore','agenum']].dropna().plot.kde(figsize=[12,8])
plt.legend(['APS Score', 'Age (years)'])
plt.xlim([-30,250])
# + [markdown] colab_type="text" id="kZDUZB5sdhhU"
# ## Questions
#
# - Do the plots change your view on how these variable should be reported?
| tutorials/eicu/04-summary-statistics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="xazrh9eIcgTO" colab_type="code" outputId="eed0e41e-8fad-4940-bab3-8e0a147e0871" colab={"base_uri": "https://localhost:8080/", "height": 383}
# !pip install torch
# !pip install torchtext
# !python -m spacy download en
# K80 gpu for 12 hours
import torch
from torch import nn, optim
from torchtext import data, datasets
print('GPU:', torch.cuda.is_available())
torch.manual_seed(123)
# + id="sPOkbQz1dfMS" colab_type="code" colab={}
TEXT = data.Field(tokenize='spacy')
LABEL = data.LabelField(dtype=torch.float)
train_data, test_data = datasets.IMDB.splits(TEXT, LABEL)
# + id="LodNOFuEeRuv" colab_type="code" outputId="742cd4b9-10a2-4de8-f0e3-2c525b3d5364" colab={"base_uri": "https://localhost:8080/", "height": 52}
print('len of train data:', len(train_data))
print('len of test data:', len(test_data))
# + id="gnQaJuCLee2o" colab_type="code" outputId="fb9424f5-2604-4680-c472-2557c0988817" colab={"base_uri": "https://localhost:8080/", "height": 72}
print(train_data.examples[15].text)
print(train_data.examples[15].label)
# + id="u3R5sgSme-Tt" colab_type="code" colab={}
# word2vec, glove
TEXT.build_vocab(train_data, max_size=10000, vectors='glove.6B.100d')
LABEL.build_vocab(train_data)
batchsz = 30
device = torch.device('cuda')
train_iterator, test_iterator = data.BucketIterator.splits(
(train_data, test_data),
batch_size = batchsz,
device=device
)
# + id="PBKKxxFBgRTM" colab_type="code" colab={}
class RNN(nn.Module):
def __init__(self, vocab_size, embedding_dim, hidden_dim):
"""
"""
super(RNN, self).__init__()
# [0-10001] => [100]
self.embedding = nn.Embedding(vocab_size, embedding_dim)
# [100] => [256]
self.rnn = nn.LSTM(embedding_dim, hidden_dim, num_layers=2,
bidirectional=True, dropout=0.5)
# [256*2] => [1]
self.fc = nn.Linear(hidden_dim*2, 1)
self.dropout = nn.Dropout(0.5)
def forward(self, x):
"""
x: [seq_len, b] vs [b, 3, 28, 28]
"""
# [seq, b, 1] => [seq, b, 100]
embedding = self.dropout(self.embedding(x))
# output: [seq, b, hid_dim*2]
# hidden/h: [num_layers*2, b, hid_dim]
# cell/c: [num_layers*2, b, hid_di]
output, (hidden, cell) = self.rnn(embedding)
# [num_layers*2, b, hid_dim] => 2 of [b, hid_dim] => [b, hid_dim*2]
hidden = torch.cat([hidden[-2], hidden[-1]], dim=1)
# [b, hid_dim*2] => [b, 1]
hidden = self.dropout(hidden)
out = self.fc(hidden)
return out
# + id="cxq70oc9lK-4" colab_type="code" outputId="b7bb2f08-0c99-4ac1-9594-e5e98168c19d" colab={"base_uri": "https://localhost:8080/", "height": 155}
rnn = RNN(len(TEXT.vocab), 100, 256)
pretrained_embedding = TEXT.vocab.vectors
print('pretrained_embedding:', pretrained_embedding.shape)
rnn.embedding.weight.data.copy_(pretrained_embedding)
print('embedding layer inited.')
optimizer = optim.Adam(rnn.parameters(), lr=1e-3)
criteon = nn.BCEWithLogitsLoss().to(device)
rnn.to(device)
# + id="_Rw_PZsZnBuJ" colab_type="code" colab={}
import numpy as np
def binary_acc(preds, y):
"""
get accuracy
"""
preds = torch.round(torch.sigmoid(preds))
correct = torch.eq(preds, y).float()
acc = correct.sum() / len(correct)
return acc
def train(rnn, iterator, optimizer, criteon):
avg_acc = []
rnn.train()
for i, batch in enumerate(iterator):
# [seq, b] => [b, 1] => [b]
pred = rnn(batch.text).squeeze(1)
#
loss = criteon(pred, batch.label)
acc = binary_acc(pred, batch.label).item()
avg_acc.append(acc)
optimizer.zero_grad()
loss.backward()
optimizer.step()
if i%10 == 0:
print(i, acc)
avg_acc = np.array(avg_acc).mean()
print('avg acc:', avg_acc)
def eval(rnn, iterator, criteon):
avg_acc = []
rnn.eval()
with torch.no_grad():
for batch in iterator:
# [b, 1] => [b]
pred = rnn(batch.text).squeeze(1)
#
loss = criteon(pred, batch.label)
acc = binary_acc(pred, batch.label).item()
avg_acc.append(acc)
avg_acc = np.array(avg_acc).mean()
print('>>test:', avg_acc)
# + id="lrjzCiiao4Qw" colab_type="code" outputId="ddc45f41-982d-4afc-e1c1-36e96a0a6e76" colab={"base_uri": "https://localhost:8080/", "height": 14878}
for epoch in range(10):
eval(rnn, test_iterator, criteon)
train(rnn, train_iterator, optimizer, criteon)
| DeepLearning/Deep-Learning-with-PyTorch-Tutorials/lesson53-情感分类实战/lstm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:aneeb]
# language: python
# name: conda-env-aneeb-py
# ---
import matplotlib.pyplot as plt
import pandas as pd
import hashlib
# %matplotlib inline
# +
data_path = 'https://raw.githubusercontent.com/vohcolab/ANEEB-2021-Python-Workshop/main/Pandas/Time%20Series/Intro%20to%20Time%20series/data/covid19Portugal.csv'
# data_path = 'data/covid19Portugal.csv' # also works if you are running locally
data_raw = pd.read_csv(data_path)
data_raw.head(5)
# -
# # Exercise 1 : Index and datetime
#
# **Objectives:**
# - Turn the date column to datetime
# - set it as index
# - sort the index (always do this)
# +
# We expect the solution to be a dataframe
data = data_raw.copy() # copy the contents of data_raw into a new variable
# YOUR CODE HERE
# ...
# -
data.head(3)
expected_hash = '8aed642bf5118b9d3c859bd4be35ecac75b6e873cce34e7b6f554b06f75550d7'
assert hashlib.sha256(str(data.iloc[0].deaths).encode()).hexdigest() == expected_hash
assert hashlib.sha256(str(data.index.dtype).encode()).hexdigest() == '261738f2e43a1c47a16f043b46deb993943d61f4a2bbe5ef4b03c3fb1af362b5'
# # Exercise 2: Working with timestamps
# #### Exercise 2.1) Day with most deaths
# +
# hint: the answer should be a timestamp
#worst_day =
# YOUR CODE HERE
#raise NotImplementedError()
# -
worst_day
expected_hash = '7c3185b857e1103d2e9aed349c3797c03510902dea53857bbb05e0ede17441d1'
assert hashlib.sha256(str(worst_day).encode()).hexdigest() == expected_hash
# #### 2.2) Tuesday with most confirmed cases
# +
# worst_tuesday_cases = ...
# -
worst_tuesday_cases
# +
expected_hash = 'ed0cfdeed24c0a3cde2783625c6af8e952f2be2989d74dfc0762d844d4727cd0'
assert hashlib.sha256(str(worst_tuesday_cases).encode()).hexdigest() == expected_hash
# -
# # Exercise 3: Time series methods
# #### 3.1) A new high since the summer started
#
# It's The first week of October. Most students are back to having classes, and Summer vacations are officially over. Your friend comes up to you and says "this week has been the worst week with most deaths since the 1st of July. I believe we need to start being more careful now".
#
# To confirm if what your friend is saying is true, compute the deadliest week of the covid data from the 1st of July up until the first week of october (including), and confirm if it actually turns out to be the first week of October.
#
# The answer should be a timestamp
# +
# assume the first week of october ends on the 4th. (For grading purposes)
# weekly_high = ...
# -
weekly_high
# +
expected_hash = '290519bb74966f3ebeab68bcb3b51550f6954f65918ee48c805aea3a8e4a4422'
assert hashlib.sha256(str(weekly_high).encode()).hexdigest() == expected_hash
# -
# #### 3.2) Is it getting better?
#
# Summer is reaching the half point now and rumors says that the number of daily new confirmed cases has been decreasing for a week now. Can you confirm that?
#
# It's currently the 29th of August, and you are asked to return the variation of daily confirmed cases of the past 7 days, excluding today. In other words from the 22nd up to the 28th.
#
# The answer should be a series with the dates in the index and the variations from the previous date in the values
#
# _hint_: be wary of the order of operations that you are going to use if you don't want any NaN values in your final result
#
#
# +
# pastweek = ...
# -
pastweek
# +
expected_hash = '5f646c309cc217ddbb46b4b46cf94b54a2dd9c44c77036b82ad36dd1ff3385be'
assert hashlib.sha256(str(pastweek).encode()).hexdigest() == expected_hash
# -
# #### 3.3) Let's get the trend
#
#
# It's currently October the 20th and you want to understand the current trend on how the death cases are evolving, without being distracted by noise. You decide to use data from the beginning of October up to today (excluding because today's data hasn't arrived yet).
#
# Use a window of 7 days and compute the number of deaths' trend of October. You must not have any NaN value in your final result
# +
# trend_deaths_october_so_far = ...
# -
trend_deaths_october_so_far
# +
expected_hash = 'e4a1f1a7670518a033b529883223f1549d1b2b127e718fba5283392a39daa9d0'
assert hashlib.sha256(str(trend_deaths_october_so_far).encode()).hexdigest() == expected_hash
# -
# Here's a basic plot of your data
trend_deaths_october_so_far.plot()
| Pandas/Time Series/Intro to Time series/Exercise Notebook - Intro to Time Series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: manim7
# language: python
# name: manim7
# ---
# # Forced Pendulum with Damping
#
# The following programs take a modified version of code from this [paper.](https://arxiv.org/pdf/1306.0949.pdf) Then the pendulum's motion is animated in manim. The first cell should be ran first as it will import every package needed for the page. The second cell should be run afterwards to initialize the physical paramaters for the pendulum, as well as to produce an array with the angles after each time step. Note any changes made on the mybinder page will not be saved unless you save the notebook locally.
from math import *
from scipy import *
import numpy as np
import matplotlib.pyplot as plt
from manim import * #Imports the animation package
# +
#From Simulation and Visualization of Chaos in a Driven Nonlinear Pendulum
#– An Aid to Introducing Chaotic Systems in Physics by <NAME> and <NAME>
#With modifications to constants and updates to compile without error
#Initial theta values
theta0 = (180*2*pi)/360
omega0 = 5*2*pi/360
#Constants
length_of_string = 9.8
gravity = 9.8
drive_frequency = 1/3
damping_force = 0.05
#Defining the driving force - controls the chaos
FD = -0.1
#Assigning the number of data points to be considered
data_points = 400
#Preallocating space for time, theta and omega
time = np.zeros(data_points)
theta = np.zeros(data_points)
omega = np.zeros(data_points)
#Initializing theta and omega
theta[0] = theta0
omega[0] = omega0
#Defining time step size
dt = 0.05
for i in range(0,data_points - 1):
time[i+1] = time[i] + dt
#Calculating for FD = 0, 0.1... in omegas
omega[i+1] = omega[i] - (gravity/length_of_string)*np.sin(theta[i])*dt - (damping_force*omega[i]*dt +FD*np.sin(drive_frequency*time[i])*dt)
theta[i+1] = theta[i] + omega[i+1]*dt
plt.plot(time,theta)
plt.ylabel("theta")
plt.xlabel("time")
plt.show()
plt.plot(time,omega)
plt.ylabel("omega")
plt.xlabel("time")
plt.show()
# -
# ## Manim
# To learn more about manim visit [here.](https://www.manim.community/)
# ### Basics for Quick Use
# The -qh in the first line refers to the quality of the video. With -ql for low quality, -qm for medium quality, and -qh for high quality. Note as the quality increases so does the run time. There more options for quality available in the documentation.
#
# The last part of the first line must have the same name as the class with the ```def construct(self)```. From there the actual animation begins.
#
# ### Example Without Physics
# This is a toy example of what will become the pendulum animation.
# The first four lines form static shapes. self.add() places those shapes on the screen. If we stopped at that point all that would produced is a still image. The self.play() line allows there to be movement. That line specifies that one dot will rotate one revolution about the other in three seconds. The rod.add_updater() line makes sure that the two dots have a line between them even as they are moving. This prevents us from having to specify an animation for the rod.
# %%manim -qh -v WARNING rotation
class rotation(Scene):
def construct(self):
sdot = Dot().shift(UP) #Stationary dot that pendulum rotates about
mdot = Dot().shift(DOWN) #Non-stationary dot
c = Circle(radius = 2).shift(UP) #Circle to demonstrate it is rotating correctly
rod = Line(sdot.get_center(),mdot.get_center())
self.add(c,sdot,mdot,rod)
rod.add_updater(lambda m: m.become(Line(sdot.get_center(),mdot.get_center())))
self.play(Rotating(mdot,radians=2*PI,about_point=sdot.get_center()),run_time=3)
# ## With Physics
# The toy example is altered to contain the physics of the second cell.
#
# Instead of having the dot rotate one revolution, for each time step the dot rotates by the amount the angle changed. This loops until there is a complete animation of the pendulum's motion over time.
#
# ### Note
# The following two animations are identical except the second one has a polar axis in the background.
#
# The timing of the animation is incorrect when using -ql or -qm please use -qh
# %%manim -qh -v WARNING fpend
class fpend(Scene):
def construct(self):
#Initializing animation
sdot = Dot().shift(UP) #Stationary dot that pendulum rotates about
mdot = Dot().shift(DOWN).rotate(theta0,about_point=sdot.get_center()) #Non-stationary dot
c = Circle(radius = 2).shift(UP) #Circle to demonstrate it is rotating correctly
rod = Line(sdot.get_center(),mdot.get_center())
self.add(sdot,mdot,rod)
rod.add_updater(lambda m: m.become(Line(sdot.get_center(),mdot.get_center())))
#Implementing the forced pendulum
for i in range(len(theta)-1):
angle_change = theta[i+1] - theta[i] #Change in angle
self.play(Rotating(mdot,radians=angle_change,about_point=sdot.get_center()),run_time=dt)
# %%manim -qh -v WARNING fpend
class fpend(Scene):
def construct(self):
#Initializing animation
sdot = Dot().shift(UP).set_color(RED) #Stationary dot that pendulum rotates about
mdot = Dot().shift(DOWN).set_color(RED).rotate(theta0,about_point=sdot.get_center()) #Non-stationary dot
c = Circle(radius = 2).shift(UP) #Circle to demonstrate it is rotating correctly
rod = Line(sdot.get_center(),mdot.get_center()).set_color(RED)
p = PolarPlane(radius_max=2,azimuth_offset=3*PI/2).add_coordinates()
p.shift(UP)
self.add(p,rod,sdot,mdot)
rod.add_updater(lambda m: m.become(Line(sdot.get_center(),mdot.get_center()).set_color(RED)))
#Implementing the forced pendulum
for i in range(len(theta)-1):
angle_change = theta[i+1] - theta[i] #Change in angle
self.play(Rotating(mdot,radians=angle_change,about_point=sdot.get_center()),run_time=dt)
# ## Comparison to the Simple Pendulum V.1
# The following implements an undamped/undriven simple pendulum (small angle approximation). This is overlayed with the previous pendulum. Both pendulums have the same initial angle but the simple pendulum is released from rest. Try changing the initial conditions in the second cell.
#
# The two pendulums should overlap when:
# * The initial angular velocity is 0
# * There is no damping or driving forces
# * The initial angle is small
#
# The <span style="color:blue"> blue pendulum </span> is the <span style="color:blue">simple pendulum</span>
#
# The <span style="color:red"> red </span> one is the same as the previous examples.
# %%manim -qh -v WARNING fpend
class fpend(Scene):
def construct(self):
#Initializing animation for the forced pendulum
sdot = Dot().shift(UP).set_color(PURPLE) #Stationary dot that pendulums rotates about
mdot = Dot().shift(DOWN).rotate(theta0,about_point=sdot.get_center()).set_color(RED) #Non-stationary dot
c = Circle(radius = 2).shift(UP) #Circle to demonstrate it is rotating correctly
rod = Line(sdot.get_center(),mdot.get_center()).set_color(RED)
self.add(mdot,rod)
rod.add_updater(lambda m: m.become(Line(sdot.get_center(),mdot.get_center()).set_color(RED)))
#Initializing animation for the simple pendulum
mdot_simple = Dot().shift(DOWN).rotate(theta0,about_point=sdot.get_center()).set_color(BLUE) #Non-stationary dot
rod_simple = Line(sdot.get_center(),mdot_simple.get_center()).set_color(BLUE)
self.add(mdot_simple,rod_simple)
rod_simple.add_updater(lambda m: m.become(Line(sdot.get_center(),mdot_simple.get_center()).set_color(BLUE)))
self.add(sdot)
#Implementing the forced pendulum
def angle_simple(time,theta_0 = theta0,length =length_of_string, acceleration = gravity):
theta = theta_0 * np.cos(np.sqrt(length/acceleration) * time)
return theta
for i in range(len(theta)-1):
angle_change_forced = theta[i+1] - theta[i] #Change in angle
angle_change_simple = angle_simple(dt * (i +1)) - angle_simple(dt * i)
self.play(Rotating(mdot,radians=angle_change_forced,about_point=sdot.get_center()),Rotating(mdot_simple,radians=angle_change_simple,about_point=sdot.get_center()),run_time=dt)
# ## Comparison to the Simple Pendulum V.2
# The following implements an undamped/undriven simple pendulum (small angle approximation). This is overlayed with the previous pendulum. Both pendulums have the same conditions from the second cell. The following cell uses that information and the same numerical process as the second cell except $\sin{(\theta)} \approx \theta$. Try changing the initial conditions in the second cell.
#
# The two pendulums should overlap when:
# * The initial angle is small (try $10$ degrees = $10 \cdot 2\pi / 360$ radians for example)
#
#
# The <span style="color:blue"> blue pendulum </span> is the <span style="color:blue">simple pendulum</span>
#
# The <span style="color:red"> red </span> one is the same as the previous examples.
# +
#Simple pendulum numerical implementation
time_simple = np.zeros(data_points)
theta_simple = np.zeros(data_points)
omega_simple = np.zeros(data_points)
theta_simple[0] = theta0
omega_simple[0] = omega0
for i in range(0,data_points - 1):
time_simple[i+1] = time_simple[i] + dt
#Calculating for FD = 0, 0.1... in omegas
omega_simple[i+1] = omega_simple[i] - (gravity/length_of_string)*theta_simple[i]*dt - (damping_force*omega_simple[i]*dt +FD*np.sin(drive_frequency*time_simple[i])*dt)
theta_simple[i+1] = theta_simple[i] + omega_simple[i+1]*dt
plt.plot(time_simple,theta_simple)
plt.ylabel("theta")
plt.xlabel("time")
plt.show()
plt.plot(time_simple,omega_simple)
plt.ylabel("omega")
plt.xlabel("time")
plt.show()
# -
# %%manim -qh -v WARNING fpend
class fpend(Scene):
def construct(self):
#Initializing animation for the forced pendulum
sdot = Dot().shift(UP).set_color(PURPLE) #Stationary dot that pendulums rotates about
mdot = Dot().shift(DOWN).rotate(theta0,about_point=sdot.get_center()).set_color(RED) #Non-stationary dot
c = Circle(radius = 2).shift(UP) #Circle to demonstrate it is rotating correctly
rod = Line(sdot.get_center(),mdot.get_center()).set_color(RED)
self.add(mdot,rod)
rod.add_updater(lambda m: m.become(Line(sdot.get_center(),mdot.get_center()).set_color(RED)))
#Initializing animation for the simple pendulum
mdot_simple = Dot().shift(DOWN).rotate(theta0,about_point=sdot.get_center()).set_color(BLUE) #Non-stationary dot
rod_simple = Line(sdot.get_center(),mdot_simple.get_center()).set_color(BLUE)
self.add(mdot_simple,rod_simple)
rod_simple.add_updater(lambda m: m.become(Line(sdot.get_center(),mdot_simple.get_center()).set_color(BLUE)))
self.add(sdot)
for i in range(len(theta)-1):
angle_change_forced = theta[i+1] - theta[i] #Change in angle
angle_change_simple = theta_simple[i+1] - theta_simple[i]
self.play(Rotating(mdot,radians=angle_change_forced,about_point=sdot.get_center()),Rotating(mdot_simple,radians=angle_change_simple,about_point=sdot.get_center()),run_time=dt)
# ## Comparison to the (Less) Simple Pendulum
# The following implements an undamped/undriven simple pendulum (small angle approximation). This is overlayed with the previous pendulum. Both pendulums have the same conditions from the second cell. The following cell uses that information and the same numerical process as the second cell except $\sin{(\theta)} \approx \theta - \frac{\theta^3}{3!}$. Try changing the initial conditions in the second cell.
#
# The two pendulums should overlap when:
# * The initial angle is small but for larger amounts than the previous example.
#
# Note there is an overflow error for particulary high angles.
#
# The <span style="color:blue"> blue pendulum </span> is the <span style="color:blue"> (less) simple pendulum</span>
#
# The <span style="color:red"> red </span> one is the same as the previous examples.
# +
#Less Simple pendulum numerical implementation
time_simple = np.zeros(data_points)
theta_simple = np.zeros(data_points)
omega_simple = np.zeros(data_points)
theta_simple[0] = theta0
omega_simple[0] = omega0
for i in range(0,data_points - 1):
time_simple[i+1] = time_simple[i] + dt
#Calculating for FD = 0, 0.1... in omegas
omega_simple[i+1] = omega_simple[i] - (gravity/length_of_string)*(theta_simple[i] - (theta_simple[i] **3 / 6))*dt - (damping_force*omega_simple[i]*dt +FD*np.sin(drive_frequency*time_simple[i])*dt)
theta_simple[i+1] = theta_simple[i] + omega_simple[i+1]*dt
plt.plot(time_simple,theta_simple)
plt.ylabel("theta")
plt.xlabel("time")
plt.show()
plt.plot(time_simple,omega_simple)
plt.ylabel("omega")
plt.xlabel("time")
plt.show()
# -
# %%manim -qh -v WARNING fpend
class fpend(Scene):
def construct(self):
#Initializing animation for the forced pendulum
sdot = Dot().shift(UP).set_color(PURPLE) #Stationary dot that pendulums rotates about
mdot = Dot().shift(DOWN).rotate(theta0,about_point=sdot.get_center()).set_color(RED) #Non-stationary dot
c = Circle(radius = 2).shift(UP) #Circle to demonstrate it is rotating correctly
rod = Line(sdot.get_center(),mdot.get_center()).set_color(RED)
self.add(mdot,rod)
rod.add_updater(lambda m: m.become(Line(sdot.get_center(),mdot.get_center()).set_color(RED)))
#Initializing animation for the simple pendulum
mdot_simple = Dot().shift(DOWN).rotate(theta0,about_point=sdot.get_center()).set_color(BLUE) #Non-stationary dot
rod_simple = Line(sdot.get_center(),mdot_simple.get_center()).set_color(BLUE)
self.add(mdot_simple,rod_simple)
rod_simple.add_updater(lambda m: m.become(Line(sdot.get_center(),mdot_simple.get_center()).set_color(BLUE)))
self.add(sdot)
for i in range(len(theta_simple)-1):
angle_change_forced = theta[i+1] - theta[i] #Change in angle
angle_change_simple = theta_simple[i+1] - theta_simple[i]
self.play(Rotating(mdot,radians=angle_change_forced,about_point=sdot.get_center()),Rotating(mdot_simple,radians=angle_change_simple,about_point=sdot.get_center()),run_time=dt)
# ## Comparison to the (Arbitarily Less) Simple Pendulum
# The following implements an undamped/undriven simple pendulum (small angle approximation). This is overlayed with the previous pendulum. Both pendulums have the same conditions from the second cell. The following cell uses that information and the same numerical process as the second cell except $\sin{(\theta)} \approx \sum \frac{(-1)^n \theta^{2n +1}}{(2n+1)!}$ from $n = 0$ to $n =$ the number of terms wanted. Try changing the initial conditions in the second cell.
#
# The two pendulums should overlap when:
# * The initial angle is smaller or when the number of terms in the sine are larger.
#
# Try adjusting the number of terms.
#
# Note there is an overflow error for particulary high angles and for certain numbers of terms.
#
# The <span style="color:blue"> blue pendulum </span> is the <span style="color:blue"> (arbitarily less) simple pendulum</span>
#
# The <span style="color:red"> red </span> one is the same as the previous examples.
#
# ### Note
# Interestingly the overflow error occurs for when the sine is approximated by an even number of terms less than 19 when using an initial angle of 180 degrees.
# +
def sine_approx(x,term_num):
term = x
series = x
for n in range(term_num-1):
if term_num != 1:
term = term * -(x**2 / ((2*n + 3) * (2*n+2)))
series += term
else:
return series
return series
# +
#Less Simple pendulum numerical implementation
time_simple = np.zeros(data_points)
theta_simple = np.zeros(data_points)
omega_simple = np.zeros(data_points)
theta_simple[0] = theta0
omega_simple[0] = omega0
for i in range(0,data_points - 1):
time_simple[i+1] = time_simple[i] + dt
#Calculating for FD = 0, 0.1... in omegas
omega_simple[i+1] = omega_simple[i] - (gravity/length_of_string)*(sine_approx(theta_simple[i],40))*dt - (damping_force*omega_simple[i]*dt +FD*np.sin(drive_frequency*time_simple[i])*dt)
theta_simple[i+1] = theta_simple[i] + omega_simple[i+1]*dt
plt.plot(time_simple,theta_simple)
plt.ylabel("theta")
plt.xlabel("time")
plt.show()
plt.plot(time_simple,omega_simple)
plt.ylabel("omega")
plt.xlabel("time")
plt.show()
# -
# %%manim -qh -v WARNING fpend
class fpend(Scene):
def construct(self):
#Initializing animation for the forced pendulum
sdot = Dot().shift(UP).set_color(PURPLE) #Stationary dot that pendulums rotates about
mdot = Dot().shift(DOWN).rotate(theta0,about_point=sdot.get_center()).set_color(RED) #Non-stationary dot
c = Circle(radius = 2).shift(UP) #Circle to demonstrate it is rotating correctly
rod = Line(sdot.get_center(),mdot.get_center()).set_color(RED)
self.add(mdot,rod)
rod.add_updater(lambda m: m.become(Line(sdot.get_center(),mdot.get_center()).set_color(RED)))
#Initializing animation for the simple pendulum
mdot_simple = Dot().shift(DOWN).rotate(theta0,about_point=sdot.get_center()).set_color(BLUE) #Non-stationary dot
rod_simple = Line(sdot.get_center(),mdot_simple.get_center()).set_color(BLUE)
self.add(mdot_simple,rod_simple)
rod_simple.add_updater(lambda m: m.become(Line(sdot.get_center(),mdot_simple.get_center()).set_color(BLUE)))
self.add(sdot)
for i in range(len(theta_simple)-1):
angle_change_forced = theta[i+1] - theta[i] #Change in angle
angle_change_simple = theta_simple[i+1] - theta_simple[i]
self.play(Rotating(mdot,radians=angle_change_forced,about_point=sdot.get_center()),Rotating(mdot_simple,radians=angle_change_simple,about_point=sdot.get_center()),run_time=dt)
| Forced_Pendulum.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import platform
import numpy as np
import pandas as pd
import matplotlib
import notebook
import cv2
import tensorflow as tf
import keras
import tqdm
import imageio
print()
# !head -2 /etc/os-release
print()
# !nvidia-smi -L
print()
print('%-16s %s' % ('Python', platform.python_version()))
print('%-16s %s' % ('Notebook', notebook.__version__))
print('%-16s %s' % ('NumPy', np.__version__))
print('%-16s %s' % ('Pandas', pd.__version__))
print('%-16s %s' % ('Matplotlib', matplotlib.__version__))
print('%-16s %s' % ('OpenCV', cv2.__version__))
print('%-16s %s' % ('TensorFlow', tf.__version__))
print('%-16s %s' % ('Keras', keras.__version__))
print('%-16s %s' % ('tqdm', tqdm.__version__))
print('%-16s %s' % ('imageio', imageio.__version__))
# -
| misc/environment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook was prepared by [<NAME>](http://donnemartin.com). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# # Challenge Notebook
# ## Problem: Implement fibonacci recursively, dynamically, and iteratively.
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# * [Solution Notebook](#Solution-Notebook)
# ## Constraints
#
# * Does the sequence start at 0 or 1?
# * 0
# * Can we assume the inputs are valid non-negative ints?
# * Yes
# * Are you looking for a recursive or iterative solution?
# * Implement both
# * Can we assume this fits memory?
# * Yes
# ## Test Cases
#
# * n = 0 -> 0
# * n = 1 -> 1
# * n = 6 -> 8
# * Fib sequence: 0, 1, 1, 2, 3, 5, 8, 13, 21, 34...
# ## Algorithm
#
# Refer to the [Solution Notebook](http://nbviewer.ipython.org/github/donnemartin/interactive-coding-challenges/blob/master/recursion_dynamic/fibonacci/fibonacci_solution.ipynb). If you are stuck and need a hint, the solution notebook's algorithm discussion might be a good place to start.
# ## Code
class Math(object):
def fib_iterative(self, n):
# TODO: Implement me
pass
def fib_recursive(self, n):
# TODO: Implement me
pass
def fib_dynamic(self, n):
# TODO: Implement me
pass
# ## Unit Test
#
#
#
# **The following unit test is expected to fail until you solve the challenge.**
# +
# # %load test_fibonacci.py
import unittest
class TestFib(unittest.TestCase):
def test_fib(self, func):
result = []
expected = [0, 1, 1, 2, 3, 5, 8, 13, 21, 34]
for i in range(len(expected)):
result.append(func(i))
self.assertEqual(result, expected)
print('Success: test_fib')
def main():
test = TestFib()
math = Math()
test.test_fib(math.fib_recursive)
test.test_fib(math.fib_dynamic)
test.test_fib(math.fib_iterative)
if __name__ == '__main__':
main()
# -
# ## Solution Notebook
#
# Review the [Solution Notebook](http://nbviewer.ipython.org/github/donnemartin/interactive-coding-challenges/blob/master/recursion_dynamic/fibonacci/fibonacci_solution.ipynb) for a discussion on algorithms and code solutions.
| recursion_dynamic/fibonacci/fibonacci_challenge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''py3.9'': conda)'
# name: python3
# ---
# +
import cranet
from cranet import nn, optim
from cranet.nn import functional as F
from cranet.data import Dataset, DataLoader
import numpy as np
from matplotlib import pyplot as plt
from scipy.io import loadmat
cranet.__version__
# -
train_mat = loadmat('train_32x32.mat')
class SvhnDataset(Dataset):
def __init__(self, mat, transform=None, transform_target=None) -> None:
super().__init__()
self.mat = mat
self.transform = transform
self.transform_target = transform_target
def __len__(self):
return self.mat['X'].shape[3]
def __getitem__(self, idx):
img = self.mat['X'][:, :, :, idx]
lab = self.mat['y'][idx, :]
if self.transform:
img = self.transform(img)
if self.transform_target:
lab = self.transform_target(lab)
return img, lab
# +
def transform(img: np.ndarray):
img = img.transpose((2, 0, 1)).astype(np.float32)
return cranet.Tensor(img)
def transform_target(lab: np.ndarray):
lab = lab.squeeze().astype(np.int64)
return cranet.Tensor([lab])
train_ds = SvhnDataset(train_mat, transform=transform,
transform_target=transform_target)
# +
def batch_fn(p):
rx = cranet.concat([i[0].reshape(1, 3, 32, 32) for i in p], dim=0)
ry = cranet.concat([i[1].reshape(1) for i in p], dim=0)
return rx, ry
train_ld = DataLoader(train_ds, batch_size=64,
batch_fn=batch_fn, shuffle=True)
# -
sample_img, sample_lab = train_ld[0]
sample_img.shape
sample_lab.shape
class Model(nn.Module):
def __init__(self):
super().__init__()
self.conv0 = nn.Conv2d(3, 32, 3, padding=1)
self.conv1 = nn.Conv2d(32, 32, 3, padding=1)
self.dropout0 = nn.Dropout(0.25)
self.conv2 = nn.Conv2d(32, 64, 3, padding=1)
self.conv3 = nn.Conv2d(64, 64, 3, padding=1)
self.dropout1 = nn.Dropout(0.25)
self.linear0 = nn.Linear(64*8*8, 512)
self.dropout2 = nn.Dropout(0.5)
self.linear1 = nn.Linear(512, 10)
def forward(self, x):
x = self.conv0(x)
x = F.relu(x)
x = self.conv1(x)
x = F.relu(x)
x = F.max_pool2d(x, 2)
x = self.dropout0(x)
x = self.conv2(x)
x = F.relu(x)
x = self.conv3(x)
x = F.relu(x)
x = F.max_pool2d(x, 2)
x = self.dropout1(x)
x = F.flatten(x, start_dim=1)
x = self.linear0(x)
x = F.relu(x)
x = self.dropout2(x)
x = self.linear1(x)
out = F.log_softmax(x, dim=1)
return out
model = Model()
optm = optim.SGD(model.parameters(), 0.1)
train_loss = []
def train(epoch: int):
for i, (inp, lab) in enumerate(train_ld):
pre = model(inp)
loss = F.nll_loss(pre, lab)
optm.zero_grad()
loss.backward()
optm.step()
loss_v = loss.numpy()
train_loss.append(loss_v)
print(f"Epoch:{epoch+1}\t:Step:{i+1}\tLoss:{loss_v}")
epochs = 10
# + pycharm={"name": "#%%\n"}
for i in range(epochs):
train(i)
| examples/svhn/svhn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# Autograd: automatic differentiation
# ===================================
#
# Central to all neural networks in PyTorch is the ``autograd`` package.
# Let’s first briefly visit this, and we will then go to training our
# first neural network.
#
#
# The ``autograd`` package provides automatic differentiation for all operations
# on Tensors. It is a define-by-run framework, which means that your backprop is
# defined by how your code is run, and that every single iteration can be
# different.
#
# Let us see this in more simple terms with some examples.
#
# Tensor
# --------
#
# ``torch.Tensor`` is the central class of the package. If you set its attribute
# ``.requires_grad`` as ``True``, it starts to track all operations on it. When
# you finish your computation you can call ``.backward()`` and have all the
# gradients computed automatically. The gradient for this tensor will be
# accumulated into ``.grad`` attribute.
#
# To stop a tensor from tracking history, you can call ``.detach()`` to detach
# it from the computation history, and to prevent future computation from being
# tracked.
#
# To prevent tracking history (and using memory), you can also wrap the code block
# in ``with torch.no_grad():``. This can be particularly helpful when evaluating a
# model because the model may have trainable parameters with `requires_grad=True`,
# but we don't need the gradients.
#
# There’s one more class which is very important for autograd
# implementation - a ``Function``.
#
# ``Tensor`` and ``Function`` are interconnected and build up an acyclic
# graph, that encodes a complete history of computation. Each variable has
# a ``.grad_fn`` attribute that references a ``Function`` that has created
# the ``Tensor`` (except for Tensors created by the user - their
# ``grad_fn is None``).
#
# If you want to compute the derivatives, you can call ``.backward()`` on
# a ``Tensor``. If ``Tensor`` is a scalar (i.e. it holds a one element
# data), you don’t need to specify any arguments to ``backward()``,
# however if it has more elements, you need to specify a ``gradient``
# argument that is a tensor of matching shape.
#
#
import torch
# Create a tensor and set requires_grad=True to track computation with it
#
#
x = torch.ones(2, 2, requires_grad=True)
print(x)
# Do an operation of tensor:
#
#
y = x + 2
print(y)
# ``y`` was created as a result of an operation, so it has a ``grad_fn``.
#
#
print(y.grad_fn)
# Do more operations on y
#
#
z = y * y * 3
out = z.mean()
print(z, out)
# ``.requires_grad_( ... )`` changes an existing Tensor's ``requires_grad``
# flag in-place. The input flag defaults to ``True`` if not given.
#
#
# Gradients
# ---------
# Let's backprop now
# Because ``out`` contains a single scalar, ``out.backward()`` is
# equivalent to ``out.backward(torch.tensor(1))``.
#
#
out.backward()
# print gradients d(out)/dx
#
#
#
print(x.grad)
print(y.grad)
# You should have got a matrix of ``4.5``. Let’s call the ``out``
# *Tensor* “$o$”.
# We have that $o = \frac{1}{4}\sum_i z_i$,
# $z_i = 3(x_i+2)^2$ and $z_i\bigr\rvert_{x_i=1} = 27$.
# Therefore,
# $\frac{\partial o}{\partial x_i} = \frac{3}{2}(x_i+2)$, hence
# $\frac{\partial o}{\partial x_i}\bigr\rvert_{x_i=1} = \frac{9}{2} = 4.5$.
#
#
# **You should read this explanation, it explains how autograd works really well:**
#
# https://towardsdatascience.com/pytorch-autograd-understanding-the-heart-of-pytorchs-magic-2686cd94ec95
#
# In essence it can be explained like this: when you call .backward() function on a tensor (let's call it z), it goes to the computation graph of that tensor, and constructs the inverse computation graph up to the leaves (leaves are tensors that are initialized by user or data) of the computation graph which have requires_grad=True. After this construction, it computes the gradient of z with respect to these leaf tensors and **ADDS** the computed gradient to the .grad field of leaf tensors.
#
# This can be demonstrated better by an example:
#
# Let's say we have x=1, y=2 and z=x\*y and we want to take the gradient of z with respect to x. Here, x and y are the **leaves** of the computation graph:
#
# 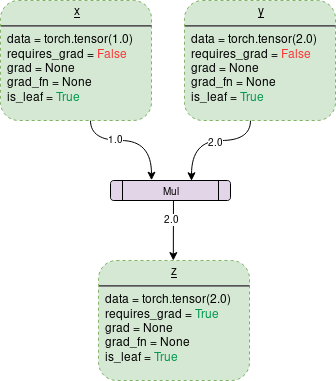
#
# Now let's examine each tensor's properties:
#
# +
import torch
# Creating the graph
x = torch.tensor(1.0, requires_grad = True)
y = torch.tensor(2.0)
z = x * y
# Displaying
for i, name in zip([x, y, z], "xyz"):
print(f"{name}\ndata: {i.data}\nrequires_grad: {i.requires_grad}\n\
grad: {i.grad}\ngrad_fn: {i.grad_fn}\nis_leaf: {i.is_leaf}\n")
# -
# Notice how grad of x and y are None before we call z.backward()
# +
z.backward()
# Displaying
for i, name in zip([x, y, z], "xyz"):
print(f"{name}\ndata: {i.data}\nrequires_grad: {i.requires_grad}\n\
grad: {i.grad}\ngrad_fn: {i.grad_fn}\nis_leaf: {i.is_leaf}\n")
# -
# Now notice that x.grad is 2 and y.grad is still None. That's because we didn't keep track of y's operations by keeping y.requires_grad=False.
#
# Pay attention to what happens when we introduce another variable w and call w.backward():
# +
w=x*y
w.backward()
# Displaying
for i, name in zip([x, y,z, w], "xyzw"):
print(f"{name}\ndata: {i.data}\nrequires_grad: {i.requires_grad}\n\
grad: {i.grad}\ngrad_fn: {i.grad_fn}\nis_leaf: {i.is_leaf}\n")
# -
# x.grad=4 because the gradient $\frac{\partial w}{\partial x}$ = 2 was **ADDED** to the existing value of x.grad which resulted in x.grad=4. In order to get the actual value of $\frac{\partial w}{\partial x} $ we first need to set x.grad to 0, and then call w.backward():
# +
x.grad.zero_()
w=x*y
w.backward()
# Displaying
for i, name in zip([x, y,z, w], "xyzw"):
print(f"{name}\ndata: {i.data}\nrequires_grad: {i.requires_grad}\n\
grad: {i.grad}\ngrad_fn: {i.grad_fn}\nis_leaf: {i.is_leaf}\n")
# -
# We can also do the same thing for tensors:
# +
# Creating the graph
x = torch.tensor([0.0, 2.0, 8.0], requires_grad = True)
y = torch.tensor([5.0 , 1.0 , 7.0])
z = x + y
# Displaying
for i, name in zip([x, y, z], "xyz"):
print(f"{name}\ndata: {i.data}\nrequires_grad: {i.requires_grad}\n\
grad: {i.grad}\ngrad_fn: {i.grad_fn}\nis_leaf: {i.is_leaf}\n")
# +
z.backward(torch.tensor([1.0, 1.0, 1.0]))
# Displaying
for i, name in zip([x, y, z], "xyz"):
print(f"{name}\ndata: {i.data}\nrequires_grad: {i.requires_grad}\n\
grad: {i.grad}\ngrad_fn: {i.grad_fn}\nis_leaf: {i.is_leaf}\n")
# -
# You can do many crazy things with autograd!
#
#
# +
x = torch.randn(3, requires_grad=True)
y = x * 2
print(y)
# +
gradients = torch.tensor([0.1, 1.0, 0.0001], dtype=torch.float)
y.backward(gradients)
print(x.grad)
# -
# You can also stops autograd from tracking history on Tensors
# with requires_grad=True by wrapping the code block in
# ``with torch.no_grad():``
#
#
# +
print(x.requires_grad)
print((x ** 2).requires_grad)
with torch.no_grad():
print((x ** 2).requires_grad)
print(x.requires_grad)
# -
# **Read Later:**
#
# Documentation of ``autograd`` and ``Function`` is at
# http://pytorch.org/docs/autograd
#
#
| jupyter/ECE283 tutorials/PyTorch tutorial/2-autograd_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="N5EAvbdlH-BG"
# # Instrumental Noise in _Kepler_ and _K2_ #3: Seasonal and Detector Effects
# ---
# + [markdown] colab_type="text" id="L57_f2QscO85"
# ## Learning Goals
#
# By the end of this tutorial, you will:
#
# - Be able to identify changes in data collection and processing caused by the _Kepler_ mission's seasonal cycles.
# - Understand instrumental noise caused by physical features of the _Kepler_ telescope.
# - Be able to compare the impact of different seasonal and detector-based noise sources.
#
# + [markdown] colab_type="text" id="D0ixWhW-cTfm"
# ## Introduction
#
# This notebook is the third part of a series on identifying instrumental and systematic sources of noise in _Kepler_ and _K2_ data. The first two tutorials in this series are suggested (but not necessary) reading for working through this one. Assumed knowledge for this tutorial is a working familiarity with _Kepler_ light curve files, target pixel files, and their associated metadata, as well as a basic understanding of the transit method for detecting exoplants.
# + [markdown] colab_type="text" id="f0yfyjrD2ERM"
# ## Imports
#
# We'll use **[Lightkurve](https://docs.lightkurve.org/)** for downloading and handling _Kepler_ data throughout this tutorial. We'll also use **[NumPy](https://numpy.org/)** to handle arrays for aperture masks, and **[Matplotlib](https://matplotlib.org/)** to help with some plotting.
# + colab={} colab_type="code" id="LQ6s2Mlwc4l9"
import lightkurve as lk
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.gridspec as gs
# %matplotlib inline
# + [markdown] colab_type="text" id="NDHPQkWhhgxv"
# ---
# + [markdown] colab_type="text" id="pG12XB_YIlu4"
# ## 1. Background
#
# The _Kepler_ telescope observed the same area of sky for four years, rotating every three months and dividing the mission into 18 observing quarters. This meant that the same star would fall on four different modules — and different pixels — throughout the mission.
#
# In this tutorial, we'll explore three types of systematic effects caused by features of the detector, including those which are influenced by seasonal rotation. We will not cover any effects of the onboard electronics, nor the electronic response to external events, such as cosmic rays. These will be covered in the fourth and final tutorial in this series on instrumental noise.
# + [markdown] colab_type="text" id="4GohdQohI2py"
# ## 2. Differential Velocity Aberration and Image Motion
#
# In this section, we'll look at the changes in a star's position on two different scales: between quarters, and within one quarter.
#
# Throughout the four-year nominal _Kepler_ mission, differential velocity aberration (DVA) was the main cause of changes in stellar positions ([_Kepler_ Instrument Handbook](https://archive.stsci.edu/files/live/sites/mast/files/home/missions-and-data/kepler/_documents/KSCI-19033-002-instrument-hb.pdf), Section 3.9). To understand DVA, first imagine the _Kepler_ telescope travelling through space. We can draw a "velocity vector" in the direction of the telescope's motion. We can draw similar vectors from every point on the focal plane, which represent lines of sight. The angle between the velocity vector and any given line of sight varies across the detector. Because the velocity vector changed each quarter during the _Kepler_ mission, the effective diameter of _Kepler's_ field of view changed by 6 arcseconds every year of the mission. This effect is on order 100 times greater than the effects of pointing instability; it leads to low frequency noise, which can be removed by light curve corrections. Because it's dependent on the line of sight, the effects of DVA are more pronounced towards the edges of the _Kepler_ field of view; some targets can move as much as half a pixel over the course of a quarter.
#
# We can observe the effects of DVA by looking at how one star's position changes across four quarters. The following code plots Quarters 8–11 for KIC 2437394, with the right ascension and declination overlayed using the [World Coordinate System](https://docs.astropy.org/en/stable/wcs/index.html) (WCS), which is stored in the target pixel files (TPFs).
# + colab={"base_uri": "https://localhost:8080/", "height": 632} colab_type="code" executionInfo={"elapsed": 72352, "status": "ok", "timestamp": 1600731753320, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="A76jlOQGJcyH" outputId="bf25f632-3673-42ad-9b02-7cd712ab8b21"
fig = plt.figure(figsize=(12, 9))
grid = gs.GridSpec(2,2)
plt.rcParams.update({'font.size': 12})
i = 0
for q in np.arange(8,12):
x,y = np.unravel_index(i, (2,2))
tpf = lk.search_targetpixelfile('KIC 2437394', quarter=q).download()
ax = plt.subplot(grid[x,y], projection=tpf.wcs)
tpf.plot(ax=ax, title=f'Quarter {q}', xlabel=' ', ylabel=' ')
i+=1
fig.text(0, 0.5, 'Dec', ha='center', rotation='vertical', size=20)
fig.text(0.5, 0, 'RA', ha='center', size=20)
# + [markdown] colab_type="text" id="oSCfcu-8uz2R"
# In the above plots, you can see the changing pixel response function (PRF), (see [_Kepler_ Instrument Handbook](https://archive.stsci.edu/files/live/sites/mast/files/home/missions-and-data/kepler/_documents/KSCI-19033-002-instrument-hb.pdf), Section 3.5) as the star changes modules; we'll look at the result of this in Section 3 of this tutorial. Also notice the very slight changes in position, on the scale of seconds, between quarters. The star's coordinates themselves are not changing, but rather the angle between this part of the detector's line of sight and the telescope's velocity vector is changing.
#
# DVA is one of the effects that contributed to image motion within quarters during the _Kepler_ mission ([_Kepler_ Data Characteristics Handbook](https://archive.stsci.edu/files/live/sites/mast/files/home/missions-and-data/kepler/_documents/Data_Characteristics.pdf), Section 5.1). Additionally, target position was influenced by focus changes, as outlined in the second tutorial in this series, and other small pointing errors. These cumulative effects led to target motion on the detector by several 100ths of a pixel per quarter.
#
# Image motion is measured onboard, and included in light curve and target pixel file metadata in the columns `pos_corr1` and `pos_corr2`. We can use this data to visualize the image motion across one quarter:
# + colab={"base_uri": "https://localhost:8080/", "height": 406} colab_type="code" executionInfo={"elapsed": 72345, "status": "ok", "timestamp": 1600731753322, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/<KEY>4", "userId": "05704237875861987058"}, "user_tz": 420} id="OXz9LIarJVNz" outputId="dab64924-4eb8-4403-e368-86d08b58f7d5"
example_tpf = lk.search_targetpixelfile('KIC 2437394', quarter=8).download()
fig = plt.figure()
plt.rcParams.update({'font.size': 20})
plt.scatter(example_tpf.pos_corr1, example_tpf.pos_corr2, c=example_tpf.time.value, s=2)
plt.xlabel('column motion (pixels)')
plt.ylabel('row motion (pixels)')
cbar = plt.colorbar()
cbar.set_label('time')
fig.set_size_inches(9,6)
# + [markdown] colab_type="text" id="G24UMNHfJE8F"
# ## 3. Quarter Boundary Discontinuities
#
# When a star was observed by new pixels in a new quarter, it was also subject to a different pixel response function (PRF). Because of this, the data processing pipeline also changed its optimal aperture, used to select pixels for aperture photometry.
#
# For most stars, these changes were not drastic. Additionally, incorrect optimal apertures in the early mission were corrected in later data releases ([_Kepler_ Data Characteristics Handbook](https://archive.stsci.edu/files/live/sites/mast/files/home/missions-and-data/kepler/_documents/Data_Characteristics.pdf), Section 5.7, and [Smith et al., 2016](https://iopscience.iop.org/article/10.1088/1538-3873/128/970/124501/pdf)), reducing many disparities between quarters. However, some stars retain a degree of variation across their optimal apertures, particularly those in crowded fields.
#
# To observe this, let's take another look at our example star from before, KIC 2437394, and see how the optimal aperture changes between quarters:
# + colab={"base_uri": "https://localhost:8080/", "height": 617} colab_type="code" executionInfo={"elapsed": 8927, "status": "ok", "timestamp": 1600732001627, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="knmpcU69HmYw" outputId="2371ceb7-dc3b-4c26-ac05-fec2db80bd20"
fig = plt.figure(figsize=(12, 9))
grid = gs.GridSpec(2, 2)
for idx, quarter in enumerate(range(8, 12)):
x, y = np.unravel_index(idx, (2, 2))
tpf = lk.search_targetpixelfile('KIC 2437394', quarter=quarter).download()
ax = plt.subplot(grid[x, y])
tpf.plot(aperture_mask='pipeline', ax=ax, title=f'Quarter {quarter}')
plt.tight_layout();
# + [markdown] colab_type="text" id="JNAuVZi8IQRR"
# KIC 2437394 is in the field of open cluster NGC 6791, meaning that crowding is a major factor in determining the optimal aperture, alongside the PRF. The changing optimal aperture leads to each quarter contributing different amounts of photon flux to the aperture photometry process, which causes discontinuities at quarter boundaries. We can see this effect by looking at the full light curve for KIC 2437394.
#
# The following code downloads all 17 available quarters for KIC 2437394 and stitches them together, plotting them with dashed lines to indicate quarter boundaries:
# + colab={"base_uri": "https://localhost:8080/", "height": 387} colab_type="code" executionInfo={"elapsed": 20669, "status": "ok", "timestamp": 1600732069828, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="xRAAd_TkJgCI" outputId="45e48d00-c7f8-48f7-92f3-9844fbf5257d"
lc = lk.search_lightcurve('KIC 2437394', cadence='long').download_all().stitch().remove_outliers()
ax = lc.plot()
quarterlines = [120, 131, 169, 259, 351, 442, 538, 629, 734, 808, 906, 1000, 1098, 1182, 1273, 1372, 1471, 1558]
for i, q in enumerate(quarterlines):
ax.axvline(q, c='r', ls='--', alpha=0.5)
# + [markdown] colab_type="text" id="YNzoqyDjKd1e"
# Notice how the structure of the light curve appears to rise and fall: Quarters 3, 7, and 11 (as pictured above) all have larger apertures. A larger aperture collects more photons, meaning their contribution to noise levels is also greater, and the light curve shows a larger amount of scatter in those quarters.
#
# You can confirm this for yourself by running the code below, which prints out the standard deviation of the normalized flux in each quarter:
# + colab={"base_uri": "https://localhost:8080/", "height": 306} colab_type="code" executionInfo={"elapsed": 30779, "status": "ok", "timestamp": 1600732157709, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="RJiiWZIYNq_W" outputId="36bdc0f4-fb94-4ea8-c08b-e6767fa36695"
for q in range(1, 18):
lc = lk.search_lightcurve('KIC 2437394', cadence='long', quarter=q).download().normalize().remove_outliers()
print(f'Quarter {q} standard deviation: {np.std(lc.flux.value):.1e}')
# + [markdown] colab_type="text" id="dTHHCCyWghr8"
# In SAP and PDCSAP data, quarter boundary discontinuities are mostly present in stars in crowded fields. However, when performing custom aperture photometry, it's useful to be aware of this phenomenon and factor it into your choice of aperture and/or light curve corrections procedure.
# + [markdown] colab_type="text" id="As3_k9okJJ82"
# ## 4. Ghosts and Scattered Light
#
# Moving away from seasonal effects now, the final phenomenon we'll look at in this tutorial is a natural consequence of the construction of the _Kepler_ telescope, as documented in the [_Kepler_ Instrument Handbook](https://archive.stsci.edu/files/live/sites/mast/files/home/missions-and-data/kepler/_documents/KSCI-19033-002-instrument-hb.pdf), Section 3.10. Scattered light on the detector results from bright stars which lie near the aluminium masking at the edges of modules, or between charge-coupled device (CCD) chips. As this scattering occurs in the vicinity of these bright stars, it is less likely to affect photometry beyond regular contamination, and the more diffuse it is, the less impact it has.
#
# Ghost images are the result of reflections on the CCDs. One well-documented _Kepler_ ghost is the Schmidt Corrector Ghost, which causes stars to be reflected around the telescope's optical axis. However, these reflections are low magnitude. The type of ghost you're more likely to encounter is reflected light from very bright stars, bouncing off the field flattener lens and returning diffusely to the CCDs. The Schmidt Corrector and the field flattener are described in Sections 3.2.2 and 3.2.3 of the [_Kepler_ Instrument Handbook](https://archive.stsci.edu/files/live/sites/mast/files/home/missions-and-data/kepler/_documents/KSCI-19033-002-instrument-hb.pdf).
#
# These ghosts can be a particular problem when dealing with exoplanet false positives; it is not immediately evident when one star is being contaminated by one that's far away from it.
#
# KIC 11911580 was flagged as a _Kepler_ Object of Interest — KOI-3900 — after Quarter 12 of the mission. Let's have a look at a periodogram of its time series, and a light curve folded on the highest amplitude peak:
# + colab={"base_uri": "https://localhost:8080/", "height": 422} colab_type="code" executionInfo={"elapsed": 20078, "status": "ok", "timestamp": 1600732209601, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="t_y_E7jjJq9F" outputId="35e7b732-a48a-4efb-837c-a764f947b33d"
koi = lk.search_lightcurve('KIC 11911580', cadence='long').download_all().stitch()
koi_pg = koi.remove_nans().to_periodogram()
ax = koi_pg.plot()
ax.set_xlim(0, 2.2);
# + colab={"base_uri": "https://localhost:8080/", "height": 405} colab_type="code" executionInfo={"elapsed": 3730, "status": "ok", "timestamp": 1600732219445, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="y6im423V2i9i" outputId="461859a7-7055-4743-aec4-b43b13202b9b"
koi.fold(period=koi_pg.period_at_max_power).bin(time_bin_size=0.1).plot();
# + [markdown] colab_type="text" id="TbJUwF-7rZ3K"
# This does appear to be the signal of a transit, but the high noise levels in the periodogram suggest it's more likely to be the diluted signal of a nearby eclipsing binary. In the case of normal contamination, we'd use Lightkurve's `plot_pixels()` function to identify a nearby binary:
# + colab={"base_uri": "https://localhost:8080/", "height": 548} colab_type="code" executionInfo={"elapsed": 6941, "status": "ok", "timestamp": 1600732245060, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="Tpf6E86vsfD3" outputId="7f3bab46-7b02-43a5-f1a5-5b87c740d2e7"
koi_tpf = lk.search_targetpixelfile('KIC 11911580', cadence='long', quarter=10).download()
koi_tpf.plot_pixels(aperture_mask='pipeline', corrector_func=lambda x: x.flatten().remove_outliers());
# + [markdown] colab_type="text" id="iFJiqD8BtZFV"
# There's nothing obvious here, which indicates this is a more complicated case of contamination. In fact, KOI-3900 is contaminated by KIC 3644542, an eclipsing binary 18 rows and 44 columns away from KOI-3900 on the detector. This is an example of a diffuse ghost on the detector causing a spurious variable signal to contaminate far away pixels.
#
# KOI-3900 was determined to be a false positive due to a ghost shortly after it was flagged as a KOI, by [Coughlin et al. 2014](https://iopscience.iop.org/article/10.1088/0004-6256/147/5/119/pdf), who used ephemeris matching to compare its period to that of known eclipsing binaries. In later work, [Thompson et al. (2018)](https://iopscience.iop.org/article/10.3847/1538-4365/aab4f9/pdf) flagged false positives due to ghosts by measuring the relative strength of the signal in the optimal aperture and in background pixels.
#
# Below, we show the periodogram and phased light curve of KIC 3644542 for comparison:
# + colab={"base_uri": "https://localhost:8080/", "height": 401} colab_type="code" executionInfo={"elapsed": 20873, "status": "ok", "timestamp": 1600732298572, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="AXvNTkJrJsU1" outputId="90cabd7e-202d-4b3c-d59c-03c88e4d8e12"
ghost = lk.search_lightcurve('KIC 3644542', cadence='long').download_all().stitch()
ghost_pg = ghost.remove_nans().to_periodogram()
ax = ghost_pg.plot()
ax.set_xlim(0, 2.2);
# + colab={"base_uri": "https://localhost:8080/", "height": 387} colab_type="code" executionInfo={"elapsed": 3504, "status": "ok", "timestamp": 1600732302101, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="_BjQY-7U4NEf" outputId="8ea2aa89-1699-4cb4-c6bc-1187b2c820c1"
ghost.fold(period=ghost_pg.period_at_max_power).bin(time_bin_size=0.1).plot();
# + [markdown] colab_type="text" id="vbge5j28DVeN"
# ## About this Notebook
#
# **Author:** [<NAME>](http://orcid.org/0000-0001-8196-516X) (`<EMAIL>`)
#
# **Updated on:** 2020-09-29
# + [markdown] colab_type="text" id="RODabQGSoQHX"
# # Citing Lightkurve and Astropy
#
# If you use `lightkurve` or `astropy` for published research, please cite the authors. Click the buttons below to copy BibTeX entries to your clipboard.
# + colab={"base_uri": "https://localhost:8080/", "height": 144} colab_type="code" executionInfo={"elapsed": 799, "status": "ok", "timestamp": 1600732302966, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="7vUtrWVjnlY7" outputId="ce51a516-1596-40f7-de2d-4cf25b128960"
lk.show_citation_instructions()
# + [markdown] colab_type="text" id="y1HbVfHwDVeN"
# <img style="float: right;" src="https://raw.githubusercontent.com/spacetelescope/notebooks/master/assets/stsci_pri_combo_mark_horizonal_white_bkgd.png" alt="Space Telescope Logo" width="200px"/>
| notebooks/MAST/Kepler/kepler_instrumental_noise_3_seasonal_and_detector_effects/kepler_instrumental_noise_3_seasonal_and_detector_effects.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Link prediction with Heterogeneous GraphSAGE (HinSAGE)
# + [markdown] nbsphinx="hidden" tags=["CloudRunner"]
# <table><tr><td>Run the latest release of this notebook:</td><td><a href="https://mybinder.org/v2/gh/stellargraph/stellargraph/master?urlpath=lab/tree/demos/link-prediction/hinsage-link-prediction.ipynb" alt="Open In Binder" target="_parent"><img src="https://mybinder.org/badge_logo.svg"/></a></td><td><a href="https://colab.research.google.com/github/stellargraph/stellargraph/blob/master/demos/link-prediction/hinsage-link-prediction.ipynb" alt="Open In Colab" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg"/></a></td></tr></table>
# -
# In this example, we use our generalisation of the [GraphSAGE](http://snap.stanford.edu/graphsage/) algorithm to heterogeneous graphs (which we call HinSAGE) to build a model that predicts user-movie ratings in the MovieLens dataset (see below). The problem is treated as a supervised link attribute inference problem on a user-movie network with nodes of two types (users and movies, both attributed) and links corresponding to user-movie ratings, with integer `rating` attributes from 1 to 5 (note that if a user hasn't rated a movie, the corresponding user-movie link does not exist in the network).
#
# To address this problem, we build a model with the following architecture: a two-layer HinSAGE model that takes labeled `(user, movie)` node pairs corresponding to user-movie ratings, and outputs a pair of node embeddings for the `user` and `movie` nodes of the pair. These embeddings are then fed into a link regression layer, which applies a binary operator to those node embeddings (e.g., concatenating them) to construct the link embedding. Thus obtained link embeddings are passed through the link regression layer to obtain predicted user-movie ratings. The entire model is trained end-to-end by minimizing the loss function of choice (e.g., root mean square error between predicted and true ratings) using stochastic gradient descent (SGD) updates of the model parameters, with minibatches of user-movie training links fed into the model.
# + nbsphinx="hidden" tags=["CloudRunner"]
# + nbsphinx="hidden" tags=["VersionCheck"]
# verify that we're using the correct version of StellarGraph for this notebook
import stellargraph as sg
try:
sg.utils.validate_notebook_version("1.2.1")
except AttributeError:
raise ValueError(
f"This notebook requires StellarGraph version 1.2.1, but a different version {sg.__version__} is installed. Please see <https://github.com/stellargraph/stellargraph/issues/1172>."
) from None
# +
import json
import pandas as pd
import numpy as np
from sklearn import preprocessing, feature_extraction, model_selection
from sklearn.metrics import mean_absolute_error, mean_squared_error
import stellargraph as sg
from stellargraph.mapper import HinSAGELinkGenerator
from stellargraph.layer import HinSAGE, link_regression
from tensorflow.keras import Model, optimizers, losses, metrics
import multiprocessing
from stellargraph import datasets
from IPython.display import display, HTML
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# Specify the minibatch size (number of user-movie links per minibatch) and the number of epochs for training the ML model:
# + tags=["parameters"]
batch_size = 200
epochs = 20
# Use 70% of edges for training, the rest for testing:
train_size = 0.7
test_size = 0.3
# -
# ## Load the dataset
# + [markdown] tags=["DataLoadingLinks"]
# (See [the "Loading from Pandas" demo](../basics/loading-pandas.ipynb) for details on how data can be loaded.)
# + tags=["DataLoading"]
dataset = datasets.MovieLens()
display(HTML(dataset.description))
G, edges_with_ratings = dataset.load()
# -
edges_with_ratings
print(G.info())
# Split the edges into train and test sets for model training/evaluation:
# +
edges_train, edges_test = model_selection.train_test_split(
edges_with_ratings, train_size=train_size, test_size=test_size
)
edgelist_train = list(edges_train[["user_id", "movie_id"]].itertuples(index=False))
edgelist_test = list(edges_test[["user_id", "movie_id"]].itertuples(index=False))
labels_train = edges_train["rating"]
labels_test = edges_test["rating"]
# -
edges_train
labels_train
labels_test
# Our machine learning task of learning user-movie ratings can be framed as a supervised Link Attribute Inference: given a graph of user-movie ratings, we train a model for rating prediction using the ratings edges_train, and evaluate it using the test ratings edges_test. The model also requires the user-movie graph structure, to do the neighbour sampling required by the HinSAGE algorithm.
# We create the link mappers for sampling and streaming training and testing data to the model. The link mappers essentially "map" user-movie links to the input of HinSAGE: they take minibatches of user-movie links, sample 2-hop subgraphs of G with `(user, movie)` head nodes extracted from those user-movie links, and feed them, together with the corresponding user-movie ratings, to the input layer of the HinSAGE model, for SGD updates of the model parameters.
# Specify the sizes of 1- and 2-hop neighbour samples for HinSAGE:
#
# Note that the length of `num_samples` list defines the number of layers/iterations in the HinSAGE model.
num_samples = [8, 4]
# Create the generators to feed data from the graph to the Keras model. We need to specify the nodes types for the user-movie pairs that we will feed to the model. The `shuffle=True` argument is given to the `flow` method to improve training.
generator = HinSAGELinkGenerator(
G, batch_size, num_samples, head_node_types=["user", "movie"]
)
train_gen = generator.flow(edgelist_train, labels_train, shuffle=True)
test_gen = generator.flow(edgelist_test, labels_test)
# Build the model by stacking a two-layer HinSAGE model and a link regression layer on top.
#
# First, we define the HinSAGE part of the model, with hidden layer sizes of 32 for both HinSAGE layers, a bias term, and no dropout. (Dropout can be switched on by specifying a positive `dropout` rate, `0 < dropout < 1`)
#
# Note that the length of `layer_sizes` list must be equal to the length of `num_samples`, as `len(num_samples)` defines the number of hops (layers) in the HinSAGE model.
generator.schema.type_adjacency_list(generator.head_node_types, len(num_samples))
generator.schema.schema
# +
hinsage_layer_sizes = [32, 32]
assert len(hinsage_layer_sizes) == len(num_samples)
hinsage = HinSAGE(
layer_sizes=hinsage_layer_sizes, generator=generator, bias=True, dropout=0.0
)
# -
# Expose input and output sockets of hinsage:
x_inp, x_out = hinsage.in_out_tensors()
# Add the final estimator layer for predicting the ratings. The edge_embedding_method argument specifies the way in which node representations (node embeddings) are combined into link representations (recall that links represent user-movie ratings, and are thus pairs of (user, movie) nodes). In this example, we will use `concat`, i.e., node embeddings are concatenated to get link embeddings.
# So basically we are saying that:
#
# An EDGE embedding is equal to CONCATENATION of the source and target NODE embedding. OK!
# Final estimator layer
score_prediction = link_regression(edge_embedding_method="concat")(x_out)
# Create the Keras model, and compile it by specifying the optimizer, loss function to optimise, and metrics for diagnostics:
# +
import tensorflow.keras.backend as K
def root_mean_square_error(s_true, s_pred):
return K.sqrt(K.mean(K.pow(s_true - s_pred, 2)))
model = Model(inputs=x_inp, outputs=score_prediction)
model.compile(
optimizer=optimizers.Adam(lr=1e-2),
loss=losses.mean_squared_error,
metrics=[root_mean_square_error, metrics.mae],
)
# -
# Summary of the model:
model.summary()
# Specify the number of workers to use for model training
num_workers = 8
# Evaluate the fresh (untrained) model on the test set (for reference):
# +
test_metrics = model.evaluate(
test_gen, verbose=1, use_multiprocessing=False, workers=num_workers
)
print("Untrained model's Test Evaluation:")
for name, val in zip(model.metrics_names, test_metrics):
print("\t{}: {:0.4f}".format(name, val))
# -
# Train the model by feeding the data from the graph in minibatches, using mapper_train, and get validation metrics after each epoch:
history = model.fit(
train_gen,
validation_data=test_gen,
epochs=epochs,
verbose=1,
shuffle=False,
use_multiprocessing=False,
workers=num_workers,
)
# Plot the training history:
sg.utils.plot_history(history)
# Evaluate the trained model on test user-movie rankings:
# +
test_metrics = model.evaluate(
test_gen, use_multiprocessing=False, workers=num_workers, verbose=1
)
print("Test Evaluation:")
for name, val in zip(model.metrics_names, test_metrics):
print("\t{}: {:0.4f}".format(name, val))
# -
# Compare the predicted test rankings with "mean baseline" rankings, to see how much better our model does compared to this (very simplistic) baseline:
# +
y_true = labels_test
# Predict the rankings using the model:
y_pred = model.predict(test_gen)
# Mean baseline rankings = mean movie ranking:
y_pred_baseline = np.full_like(y_pred, np.mean(y_true))
rmse = np.sqrt(mean_squared_error(y_true, y_pred_baseline))
mae = mean_absolute_error(y_true, y_pred_baseline)
print("Mean Baseline Test set metrics:")
print("\troot_mean_square_error = ", rmse)
print("\tmean_absolute_error = ", mae)
rmse = np.sqrt(mean_squared_error(y_true, y_pred))
mae = mean_absolute_error(y_true, y_pred)
print("\nModel Test set metrics:")
print("\troot_mean_square_error = ", rmse)
print("\tmean_absolute_error = ", mae)
# -
# Compare the distributions of predicted and true rankings for the test set:
h_true = plt.hist(y_true, bins=30, facecolor="green", alpha=0.5)
h_pred = plt.hist(y_pred, bins=30, facecolor="blue", alpha=0.5)
plt.xlabel("ranking")
plt.ylabel("count")
plt.legend(("True", "Predicted"))
plt.show()
# We see that our model beats the "mean baseline" by a significant margin. To further improve the model, you can try increasing the number of training epochs, change the dropout rate, change the sample sizes for subgraph sampling `num_samples`, hidden layer sizes `layer_sizes` of the HinSAGE part of the model, or try increasing the number of HinSAGE layers.
#
# However, note that the distribution of predicted scores is still very narrow, and rarely gives 1, 2 or 5 as a score.
# This model uses a bipartite user-movie graph to learn to predict movie ratings. It can be further enhanced by using additional relations, e.g., friendships between users, if they become available. And the best part is: the underlying algorithm of the model does not need to change at all to take these extra relations into account - all that changes is the graph that it learns from!
# + [markdown] nbsphinx="hidden" tags=["CloudRunner"]
# <table><tr><td>Run the latest release of this notebook:</td><td><a href="https://mybinder.org/v2/gh/stellargraph/stellargraph/master?urlpath=lab/tree/demos/link-prediction/hinsage-link-prediction.ipynb" alt="Open In Binder" target="_parent"><img src="https://mybinder.org/badge_logo.svg"/></a></td><td><a href="https://colab.research.google.com/github/stellargraph/stellargraph/blob/master/demos/link-prediction/hinsage-link-prediction.ipynb" alt="Open In Colab" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg"/></a></td></tr></table>
| notebooks/hinsage-link-prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: text-as-data
# language: python
# name: text-as-dada
# ---
# # In a few paragraphs, explain when and why dictionaries successfully measure that which they attempt to measure, when they do not, and the possible risks of analyzing text with dictionaries.
#
# Dictionaries successfully classify a document through the mathematical process of applying measurable relationships between words and their associated classes. Both approaches, let it be using per-existing relationships (ie weights) or generating new ones, take what's known about a group of documents (classified documents) and apply that knowledge to a set of new documents (unclassified documents). Thousands, if not tens of thousands, of relationships are applied to each new document individually - all that knowledge, applied to each new document - is what makes this form of classification powerful.
#
# Dictionaries successfully measure things because of the way in which they measure individual words, in particular, the same word in different sentences. A word holds its value relative to itself and its neighboring words. For example in the sentences below,
#
# "Carl likes to go mountain biking."
# "Carl likes to go road biking."
# From a mathametcal standpoint the word 'Carl' has two, subtle, different values. These values are not only determeined by the word itself but also the sentence it resides in. I feel it's safe to say the above sentences' Carl-values are close to eachother.
# Conversely, let's consider a third sentence,
#
# "When Carl was a wee lad he hated using training wheels, but they were a necessary evil in the pursuit to two-wheeled glory."
#
# In the above sentence the value of the word 'Carl' is significatnly different than the value of first two. Storing and accounting for this diversity is, in part, why dictionaries are able to successfully meaure things, because they are able to account for differences in word-useage. This makes me wonder, if there's a way to observe the 'strength' a dictionary has to analyze a word, and in turn, lack-there-of.
#
#
#
# ## Problems with Dictionaries.
#
# As discussed in the lecture @15:37, "dictionaries are context invariant". Thus begs the questions, what determines one context different from another and, a bit more interesting, to what extent can a dictionary be used outside of its original context? How do we know when it cannot be used?
#
# As I was thinking about these questions I recalled a youtube video I watched from a couple days ago on another COVID-resultant youtube binge of mine to kill some time: https://www.youtube.com/watch?v=oAbQEVmvm8Y
#
# In the video the speaker showcases how data scientists are using datasets from different domains, together, to solve one specific problem. That got me wondering, are there absolute bounds to which a dictionary can be used in solving a problem? The starry eyed <NAME> in me says no, no way (wave arms magically and say, "there's infinite possibilities") but the practical side of me says yes. As there's no such thing as infinate gpu memory.
#
# In the example used in the lecture ~@18:00 surrounding the word 'Hopkins'. Agreed, this dictionary should not be used to explore sentiment-relationships between Hopkins and, let's say, the public. So then what should we use? Thinking off the cuff here, after living in Baltimore for 6 years, downtown, I can say for 99% certainty there are communities in Baltimore that do not like Hopkins. Period. On the other hand, its students, well, we like Hopkins quite a bit. For every one that dislikes/likes Hopkins there’s 2 that are indifferent.
#
# So, in my opinion, we’d need to find a balance between the two extremes if we wanted to generate a dictionary surrounding sentiment and Hopkins. This balance would also need to be representative of the population to which the impending results would be applied to. I suppose there's seemingly an infinite amount of things one could do to create the 'perfect' dictionary. (It would be kind of fun to generate a dictionary such as this, one that's evenly distributed, to use to monitor FB/Insta/Twitter feeds.) From a development standpoint it would be interesting to see how generating a new model, then adding more training data to it, would work in a production environment.
# # 2. Using a dictionary in qdapDictionaries, conduct sentiment analysis on your corpus. Write up the results in a pdf and interpret them.
# # Setup
# +
import os as os
import sys as sys
import numpy as np
import pandas as pd
import seaborn as sns
from matplotlib import rc
from pylab import rcParams
import matplotlib.pyplot as plt
from collections import defaultdict
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, classification_report
#torch goodness
import torch
from torch import nn, optim
import torch.nn.functional as F
from torch.utils.data import Dataset, DataLoader
#huggingface goodness
import transformers
from transformers import BertModel, BertTokenizer, AdamW, get_linear_schedule_with_warmup
#set graph config
# %matplotlib inline
# %config InlineBackend.figure_format='retina'
sns.set(style='whitegrid', palette='muted', font_scale=1.2)
HAPPY_COLORS_PALETTE = ["#01BEFE", "#FFDD00", "#FF7D00", "#FF006D", "#ADFF02", "#8F00FF"]
sns.set_palette(sns.color_palette(HAPPY_COLORS_PALETTE))
rcParams['figure.figsize'] = 12, 8
#set seed
RANDOM_SEED = 42
np.random.seed(RANDOM_SEED)
torch.manual_seed(RANDOM_SEED)
#check gpu
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
device
# -
# # Load training datasets
# +
# https://www.kaggle.com/lava18/google-play-store-apps
# google play app reviews ~16k
df_google = pd.read_csv("google_reviews.csv")
# imdb from http://ai.stanford.edu/~amaas/data/sentiment/
# "We provide a set of 25,000 highly polar movie reviews for training, and 25,000 for testing."
# this file was larger than 100mb (github's file size limit)
df_imdb = pd.read_csv('imdb_reviews.csv')
# -
# # Preprocess training datasets
# ### imdb
# first look, includes .head() shape and missing values
def first_look(df):
# gives basic info about the df
lst =[df.head(0), df.shape, df.info()]
for item in lst:
print(item)
print('-------------------------------------------------')
# first look
first_look(df_imdb)
# +
# let's look at the scores
# 0 = negative
# 1 = neutral
# 2 = postitive
#review changes
class_names = ['negative', 'neutral', 'positive']
ax = sns.countplot(df_imdb.sentiment)
plt.xlabel('review sentiment')
sns.countplot(df_imdb.sentiment)
ax.set_xticklabels(class_names);
# -
# ### google
# first look
first_look(df_google)
# Let's look at the scores
sns.countplot(df_google.score)
plt.xlabel('review score');
# +
# dimension reduction
def to_sentiment(rating):
#restruture review scores to emulate imdb
rating = int(rating)
if rating <= 2:
return 0
elif rating == 3:
return 1
else:
return 2
#apply above function to dataset
df_google['sentiment'] = df_google.score.apply(to_sentiment)
# -
#review changes
class_names = ['negative', 'neutral', 'positive']
ax = sns.countplot(df_google.sentiment)
plt.xlabel('review sentiment')
sns.countplot(df_google.sentiment)
ax.set_xticklabels(class_names);
# +
# merge google and imdb datasets and rename to 'df'
#subset google for relavent info
df_google_subset = df_google[['content', 'sentiment']]
frames = [df_google_subset, df_imdb] #I omitted df_imdb, due to limited gpu memory
df = pd.concat(frames)
# -
# # Tokenize training dataset
# load pre-trained bert model
PRE_TRAINED_MODEL_NAME = 'bert-base-cased'
tokenizer = BertTokenizer.from_pretrained(PRE_TRAINED_MODEL_NAME)
# +
def tokenize_inspect(df):
#BERT works with fixed-length sequences. Let's inspect the length of each review from our training dataset and tokenize it
token_lens = []
for txt in df.content:
tokens = tokenizer.encode(txt, max_length=512)
token_lens.append(len(tokens))
#let's plot the distribution to get a sense of what we're working with
sns.distplot(token_lens)
plt.xlim([0, 600]);
plt.xlabel('Tokens');
# pass training dataset into tokenizer/inspection
tokenize_inspect(df)
# -
# select max len of tokens you want to train with
MAX_LEN = 512
#class that formatts df into pytorch format
class GPReviewDataset(Dataset):
def __init__(self, reviews, targets, tokenizer, max_len):
self.reviews = reviews
self.targets = targets
self.tokenizer = tokenizer
self.max_len = max_len
def __len__(self):
return len(self.reviews)
def __getitem__(self, item):
review = str(self.reviews[item])
target = self.targets[item]
encoding = self.tokenizer.encode_plus(
review,
add_special_tokens=True,
max_length=self.max_len,
return_token_type_ids=False,
pad_to_max_length=True,
return_attention_mask=True,
return_tensors='pt',
)
return {
'review_text': review,
'input_ids': encoding['input_ids'].flatten(),
'attention_mask': encoding['attention_mask'].flatten(),
'targets': torch.tensor(target, dtype=torch.long)
}
#text train split data
df_train, df_test = train_test_split(
df,
test_size=0.1,
random_state=RANDOM_SEED
)
df_val, df_test = train_test_split(
df_test,
test_size=0.5,
random_state=RANDOM_SEED
)
df_train.shape, df_val.shape, df_test.shape
# +
#We also need to create a couple of data loaders. Here’s a helper function to do it:
def create_data_loader(df, tokenizer, max_len, batch_size):
ds = GPReviewDataset(
reviews=df.content.to_numpy(),
targets=df.sentiment.to_numpy(),
tokenizer=tokenizer,
max_len=max_len
)
return DataLoader(
ds,
batch_size=batch_size,
num_workers=0
)
#set batch size and load data
BATCH_SIZE = 6
train_data_loader = create_data_loader(df_train, tokenizer, MAX_LEN, BATCH_SIZE)
val_data_loader = create_data_loader(df_val, tokenizer, MAX_LEN, BATCH_SIZE)
test_data_loader = create_data_loader(df_test, tokenizer, MAX_LEN, BATCH_SIZE)
# -
#let's look at an example batch from out trining data loader:
data = next(iter(train_data_loader))
data.keys()
print(data['input_ids'].shape)
print(data['attention_mask'].shape)
print(data['targets'].shape)
# # Sentiment Classification with BERT and Hugging Face
#load classification model
bert_model = BertModel.from_pretrained(PRE_TRAINED_MODEL_NAME)
#BERT sentiment classifier
class SentimentClassifier(nn.Module):
def __init__(self, n_classes):
super(SentimentClassifier, self).__init__()
self.bert = BertModel.from_pretrained(PRE_TRAINED_MODEL_NAME)
self.drop = nn.Dropout(p=0.3)
self.out = nn.Linear(self.bert.config.hidden_size, n_classes)
def forward(self, input_ids, attention_mask):
_, pooled_output = self.bert(
input_ids=input_ids,
attention_mask=attention_mask
)
output = self.drop(pooled_output)
return self.out(output)
# create instance and send to gpu
model = SentimentClassifier(len(class_names))
model = model.to(device)
# ### Training
# +
#To reproduce the training procedure from the BERT paper,
#we’ll use the AdamW optimizer provided by Hugging Face.
#It corrects weight decay, so it’s similar to the original paper.
#We’ll also use a linear scheduler with no warmup steps:
EPOCHS = 8
optimizer = AdamW(model.parameters(), lr=2e-5, correct_bias=False)
total_steps = len(train_data_loader) * EPOCHS
scheduler = get_linear_schedule_with_warmup(
optimizer,
num_warmup_steps=0,
num_training_steps=total_steps
)
loss_fn = nn.CrossEntropyLoss().to(device)
# -
#Let’s continue with writing a helper function for training our model for one epoch:
def train_epoch(
model,
data_loader,
loss_fn,
optimizer,
device,
scheduler,
n_examples
):
model = model.train()
losses = []
correct_predictions = 0
for d in data_loader:
input_ids = d["input_ids"].to(device)
attention_mask = d["attention_mask"].to(device)
targets = d["targets"].to(device)
outputs = model(
input_ids=input_ids,
attention_mask=attention_mask
)
_, preds = torch.max(outputs, dim=1)
loss = loss_fn(outputs, targets)
correct_predictions += torch.sum(preds == targets)
losses.append(loss.item())
loss.backward()
nn.utils.clip_grad_norm_(model.parameters(), max_norm=1.0)
optimizer.step()
scheduler.step()
optimizer.zero_grad()
return correct_predictions.double() / n_examples, np.mean(losses)
# Let’s write another one that helps us evaluate the model on a given data loader:
def eval_model(model, data_loader, loss_fn, device, n_examples):
model = model.eval()
losses = []
correct_predictions = 0
with torch.no_grad():
for d in data_loader:
input_ids = d["input_ids"].to(device)
attention_mask = d["attention_mask"].to(device)
targets = d["targets"].to(device)
outputs = model(
input_ids=input_ids,
attention_mask=attention_mask
)
_, preds = torch.max(outputs, dim=1)
loss = loss_fn(outputs, targets)
correct_predictions += torch.sum(preds == targets)
losses.append(loss.item())
return correct_predictions.double() / n_examples, np.mean(losses)
# Using those two, we can write our training loop. We’ll also store the training history:
history = defaultdict(list)
best_accuracy = 0
for epoch in range(EPOCHS):
print(f'Epoch {epoch + 1}/{EPOCHS}')
print('-' * 10)
train_acc, train_loss = train_epoch(
model,
train_data_loader,
loss_fn,
optimizer,
device,
scheduler,
len(df_train)
)
print(f'Train loss {train_loss} accuracy {train_acc}')
val_acc, val_loss = eval_model(
model,
val_data_loader,
loss_fn,
device,
len(df_val)
)
print(f'Val loss {val_loss} accuracy {val_acc}')
print()
history['train_acc'].append(train_acc)
history['train_loss'].append(train_loss)
history['val_acc'].append(val_acc)
history['val_loss'].append(val_loss)
if val_acc > best_accuracy: #fail safe
torch.save(model.state_dict(), 'best_model_state.bin')
best_accuracy = val_acc
# +
#started at 8:18 pm
#ended at 11:08 am
# +
#note that my validation accuracy went down as a result of adding the new training data
plt.plot(history['train_acc'], label='train accuracy')
plt.plot(history['val_acc'], label='validation accuracy')
plt.title('Training history')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend()
plt.ylim([0, 1]);
# -
# # Predicting Raw Text
# load raw text
df_game_reviews = pd.read_csv("game_reviews.csv")
# inspect
first_look(df_game_reviews)
# subset df to just english reviews
df_english = df_game_reviews[df_game_reviews["language"] == 'english']
# set reviews to list
title_lst = df_english['review'].tolist()
# +
title_lst_final = []
for item in title_lst:
temp = str(item)
title_lst_final.append(temp)
# +
# pass each review through sentiment model
review_text_lst = []
prediction_lst = []
for item in title_lst_final:
encoded_review = tokenizer.encode_plus(item, max_length=MAX_LEN,
add_special_tokens=True,
return_token_type_ids=False,
pad_to_max_length=True,
return_attention_mask=True,
return_tensors='pt',)
input_ids = encoded_review['input_ids'].to(device)
attention_mask = encoded_review['attention_mask'].to(device)
output = model(input_ids, attention_mask)
_, prediction = torch.max(output, dim=1)
review_text_lst.append(item)
prediction_lst.append(class_names[prediction])
# +
#count up your glorious sentiment results
pos_count = 0
neg_count = 0
neu_count = 0
for item in prediction_lst:
if item == 'positive':
pos_count += 1
elif item == 'neutral':
neu_count += 1
else:
neg_count += 1
# -
print('Positive articles: ', pos_count)
print('Negative articles: ', neg_count)
print('Neutral articles: ', neu_count)
df = pd.DataFrame(prediction_lst)
df_english.to_csv('english_game_reviews.csv')
df.to_csv('prediction_lst.csv')
# In this week's model, I 'attempted' to add an additional training dataset in hopes that it would increase my Validation Accuracy - wrong. The question I have though is my training dataset more-representative, now, of my Steam Reviews and in-turn more 'correct'?
#
# (more questions)
# Should have I analyzed my Steam Reviews first and then found a training dataset, that's structurally (len(), etc, etc, other parameters?) the same to use? I really didn't give my training dataset(s) much thought, other than more is better. Also, regarding the max_len of 512, for argument's sake let us say I have a corpus containing docs that are ONLY +512 in len. What then? Would I just split the sentences in half and analyze per usual? Maybe use one dictionary for < 512 and another for +512?
#
# Unrelated, but notable, I ran into GPU limitations (again) but reduced my batch size from 16 -> 4, took 18 hours of training time.
#
# Future work will involve analyzing my Game Reviews dataset to try and find a training dataset that looks like it? Also, trying to find literature on sentiment analysis BEYOND just pos/neg/neutral? Any suggestions?
#
# What I've found: http://www.crystalfeel.socialanalyticsplus.net/ and its manual: http://172.16.31.10/crystalfeel/[CrystalFeel]_User_Manual.pdf
# Also, potential app thingy? https://medium.com/@mirzamujtaba10/sentiment-analysis-642b935ab6f9
#
# And finally, I might be running into trouble with gathering Sales for the video game ('Squad').
# (things to explore)
# Crystal Feel http://www.crystalfeel.socialanalyticsplus.net/
# Crystal Feel Manual http://172.16.31.10/crystalfeel/[CrystalFeel]_User_Manual.pdf
# Super Cool App Thingy https://medium.com/@mirzamujtaba10/sentiment-analysis-642b935ab6f9
# NLP survey @17:00 https://www.youtube.com/watch?v=G5lmya6eKtc&t=1075s
# Cool NLP data size paper: https://arxiv.org/abs/2001.08361![http---172.16.31.10-crystalfeel-[CrystalFeel]_User_Manual.pdf.url](attachment:http---172.16.31.10-crystalfeel-[CrystalFeel]_User_Manual.pdf.url)
| notebooks/TAD_Week_5_Broker_Carl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# initialize the %%d3 notebook magic
import nbvis.magics
# +
# %%d3
var bannerName = "callysto-notebook-bottom-banner";
var url = "https://raw.githubusercontent.com/callysto/notebook-templates/master/" + bannerName + ".svg";
// get banner svg data, then proceed with modifications
d3.svg(url)
.then(function(svg) {
// append banner to cell output
var bannerDisplay = d3.select(element[0])
.append("g")
.html(svg.querySelector("svg")
.outerHTML);
// specify website links
var links = [
["#callysto-logo", "https://www.callysto.ca/"],
["#callysto-url", "https://www.callysto.ca/"],
["#callysto-twitter", "https://twitter.com/callysto_canada/"],
["#cybera-logo", "https://www.cybera.ca/"],
["#cybera-url", "https://www.cybera.ca/"],
["#cybera-twitter", "https://twitter.com/cybera/"],
["#pims-logo", "https://www.pims.math.ca/"],
["#pims-url", "https://www.pims.math.ca/"],
["#pims-twitter", "https://twitter.com/pimsmath/"],
["#canada-logo", "https://www.canada.ca/en/innovation-science-economic-development/programs/science-technology-partnerships/cancode.html"],
["#cc-license", "https://creativecommons.org/licenses/by/4.0/"],
["#open-source", "https://opensource.org/"],
["#mit-license", "https://opensource.org/licenses/MIT/"]
];
// link each identifier to its corresponding website
for (var i=0; i < links.length; i++) addLink(...links[i]);
// serialize the modified banner svg data to a string
var serializer = new XMLSerializer();
var bannerSource = serializer.serializeToString(element[0].querySelector("svg"));
// create a Python variable `banner_source` with the modified banner svg data
Jupyter.notebook.kernel.execute("banner_name = '" + bannerName + "'");
Jupyter.notebook.kernel.execute("banner_source = '" + bannerSource + "'");
})
function addLink(identifier, url) {
// get the identified svg element's bounding box
var groupRect = d3.select(identifier).node().getBBox();
d3.select(identifier)
// link to its url
.append("a")
.attr("xlink:href", url)
// add an invisible clickable rectangle that is sensitive to mouseover behaviour
.append("rect")
.attr("x", groupRect.x)
.attr("y", groupRect.y)
.attr("height", groupRect.height)
.attr("width", groupRect.width)
.style("opacity", 0)
.attr("onmouseover", "this.parentNode.parentNode.style.opacity=0.67;")
.attr("onmouseout", "this.parentNode.parentNode.style.opacity=1;");
}
# +
from IPython.display import SVG
# write modified banner svg data to file
with open(banner_name + '-interactive' + '.svg', 'w') as banner:
banner.write(banner_source)
# preview modified banner svg data
display(SVG(banner_source))
| banners/generate_interactive_banners.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# -*- coding: utf-8 -*-
# %matplotlib inline
import pandas as pd
import numpy as np
import math
import matplotlib
import matplotlib.pyplot as plt
from dateutil import parser
from datetime import datetime
plt.style.use('fivethirtyeight')
# -
df = pd.read_csv("calif_stations-reworked.csv")
df
fig, (ax1,ax2,ax3) = plt.subplots(nrows=1, ncols=3, figsize=(16,12), sharex=True, sharey=True)
df.plot(kind="line",x="FYear",y="Diego", figsize=(16,9), ax=ax1)
df.plot(kind="line",x="FYear",y="LA", figsize=(16,9), ax=ax2)
df.plot(kind="line",x="FYear",y="Beach", figsize=(16,9), ax=ax3)
fig, (ax1,ax2,ax3) = plt.subplots(nrows=1, ncols=3, figsize=(16,12), sharex=True, sharey=False)
df.plot(kind="line",x="FYear",y="Diego", figsize=(15,6), ax=ax1)
df.plot(kind="line",x="FYear",y="LA", figsize=(15,6), ax=ax2)
df.plot(kind="line",x="FYear",y="Beach", figsize=(15,6), ax=ax3)
fig, (ax1,ax2) = plt.subplots(nrows=1, ncols=2, figsize=(16,12), sharex=True, sharey=True)
df.plot(kind="line",x="FYear",y="Diego", figsize=(15,6), ax=ax1)
df.plot(kind="line",x="FYear",y="Beach", figsize=(15,6), ax=ax2)
df_percent = pd.read_csv("calif_stations-perc.csv")
df_percent
fig, (ax1,ax2,ax3) = plt.subplots(nrows=1, ncols=3, figsize=(16,12), sharex=True, sharey=True)
df_percent.plot(kind="bar",x="FYear",y="Diego", figsize=(15,4), ax=ax1)
df_percent.plot(kind="bar",x="FYear",y="LA", figsize=(15,4), ax=ax2)
df_percent.plot(kind="bar",x="FYear",y="Beach", figsize=(15,4), ax=ax3)
| 2017/01/va-onboard-staff-calif-20170126/data/graphic_exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # On-Axis Field Due to a Current Loop
# *This simple formula uses the [Law of Biot Savart](../basics/biotsavart.html), integrated over a circular current loop to obtain the magnetic field at any point along the axis of the loop.*
# 
# $B = \frac {\mu_o i r^2}{2(r^2 + x^2)^{\frac 3 2}}$
#
# **B** is the magnetic field, in teslas, at any point on the axis of the current loop. The direction of the field is perpendicular to the plane of the loop.
#
# $\mathbf \mu_o$ is the permeability constant (1.26x10<sup>-6</sup> Hm<sup>-1</sup>)
#
# **i** is the current in the wire, in amperes.
#
# **r** is the radius of the current loop, in meters.
#
# **x** is the distance, on axis, from the center of the current loop to the location where the magnetic field is calculated, in meters.
# ## Special Case: *x* = 0
# $B = \frac {\mu_o i}{2 r}$
# ## Special Case: *x* >> 0
# $B = \frac {\mu_o i r^2}{2 x^3}$
#
# Note that this is equivalent to the expression for on-axis magnetic field due to a magnetic dipole:
#
# $B = \frac {\mu_o i A}{2 \pi x^3}$
#
# where **A** is the area of the current loop, or $\pi r^2$.
# ## Code Example
#
# The following IPython code illustrates how to compute the on-axis field due to a simple current loop.
# +
# %matplotlib inline
from scipy.special import ellipk, ellipe, ellipkm1
from numpy import pi, sqrt, linspace
from pylab import plot, xlabel, ylabel, suptitle, legend, show
uo = 4E-7*pi # Permeability constant - units of H/m
# On-Axis field = f(current and radius of loop, x of measurement point)
def Baxial(i, a, x, u=uo):
if a == 0:
if x == 0:
return NaN
else:
return 0.0
else:
return (u*i*a**2)/2.0/(a**2 + x**2)**(1.5)
# -
# Use the `Baxial` function to compute the central field of a unit loop (1 meter radius, 1 ampere of current), in teslas:
print("{:.3} T".format(Baxial(1, 1, 0)))
# You can try selecting your own current (a), radius (m) and axial position (m) combination to see what the resulting field is:
# +
from ipywidgets import interactive
from IPython.display import display
def B(i, a, x):
return "{:.3} T".format(Baxial(i,a,x))
v = interactive(B, i=(0.0, 20.0), a=(0.0, 10.0), x=(0.0, 10.0))
display(v)
# -
# Now plot the field intensity, as a fraction of the central field, at various positions along the axis (measured as multiples of the coil radius):
axiallimit = 5.0 # meters from center
radius = 1.0 # loop radius in meters
X = linspace(0,axiallimit)
Bcenter = Baxial(1,1,0)
plot(X, [Baxial(1,1,x)/Bcenter for x in X])
xlabel("Axial Position (multiples of radius)")
ylabel("Axial B field / Bo (unitless)")
suptitle("Axial B field of simple loop")
show()
# ---
# [Magnet Formulas](../index.html), © 2018 by <NAME>. Source code and License on [Github](https://github.com/tiggerntatie/emagnet.py)
| solenoids/current_loop.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="udDs_biH0n5U" colab_type="text"
# #### Copyright 2020 Google LLC.
# + id="WPY-OyyM0pSs" colab_type="code" colab={}
# Licensed under the Apache License, Version 2.0 (the "License")
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
https://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="psnUF-8c02o_" colab_type="text"
# # Reformer: Text Generation [](https://colab.research.google.com/github/google/trax/blob/master/trax/models/reformer/text_generation.ipynb)
# + [markdown] id="1lnRd_IoERdk" colab_type="text"
# This notebook was designed to run on TPU.
#
# To use TPUs in Colab, click "Runtime" on the main menu bar and select Change runtime type. Set "TPU" as the hardware accelerator.
# + id="8PluCmWbZIpJ" colab_type="code" colab={}
# Install JAX.
# !gsutil cp gs://trax-ml/reformer/jaxlib-0.1.39-cp36-none-manylinux2010_x86_64.whl .
# !gsutil cp gs://trax-ml/reformer/jax-0.1.59-cp36-none-manylinux2010_x86_64.whl .
# !pip install --upgrade -q ./jaxlib-0.1.39-cp36-none-manylinux2010_x86_64.whl
# !pip install --upgrade -q ./jax-0.1.59-cp36-none-manylinux2010_x86_64.whl
# Make sure the Colab Runtime is set to Accelerator: TPU.
import requests
import os
if 'TPU_DRIVER_MODE' not in globals():
url = 'http://' + os.environ['COLAB_TPU_ADDR'].split(':')[0] + ':8475/requestversion/tpu_driver0.1-dev20191206'
resp = requests.post(url)
TPU_DRIVER_MODE = 1
# The following is required to use TPU Driver as JAX's backend.
from jax.config import config
config.FLAGS.jax_xla_backend = "tpu_driver"
config.FLAGS.jax_backend_target = "grpc://" + os.environ['COLAB_TPU_ADDR']
print(config.FLAGS.jax_backend_target)
# + id="yiPdBenoZwH6" colab_type="code" colab={}
# !pip install --upgrade -q sentencepiece
# !pip install --upgrade -q gin git+https://github.com/google/trax.git@v1.2.3
from tensorflow.compat.v1.io.gfile import GFile
import gin
import os
import jax
import trax
from trax.models.beam_search import Search
from trax.supervised import inputs
import numpy as onp
import jax.numpy as np
from scipy.special import softmax
from sentencepiece import SentencePieceProcessor
# + [markdown] colab_type="text" id="FQ89jHCYfhpg"
# ## Setting up data and model
# + [markdown] id="9_OCIqghSyfs" colab_type="text"
# In this notebook, we'll be pushing the limits of just how many tokens we can fit on a single TPU device. The TPUs available in Colab have 8GB of memory per core, and 8 cores. We will set up a Reformer model that can fit a copy of "Crime and Punishment" on *each* of the 8 TPU cores (over 500,000 tokens per 8GB of memory).
# + id="tYSOVGR47LVL" colab_type="code" colab={}
# Import a copy of "Crime and Punishment", by <NAME>
with GFile('gs://trax-ml/reformer/crime-and-punishment-2554.txt') as f:
text = f.read()
# The file read above includes metadata and licensing information.
# For training our language model, we will only use the actual novel text.
start = text.find('CRIME AND PUNISHMENT') # skip header
start = text.find('CRIME AND PUNISHMENT', start + 1) # skip header
start = text.find('CRIME AND PUNISHMENT', start + 1) # skip translator preface
end = text.rfind('End of Project') # skip extra text at the end
text = text[start:end].strip()
# + id="mMntV3H-6OR0" colab_type="code" colab={}
# Load a BPE vocabulaary with 320 types. This mostly consists of single letters
# and pairs of letters, but it has some common words and word pieces, too.
# !gsutil cp gs://trax-ml/reformer/cp.320.* .
TOKENIZER = SentencePieceProcessor()
TOKENIZER.load('cp.320.model')
# + id="HnJzxSi_77zP" colab_type="code" outputId="ec510c06-5a49-42aa-ebde-585e487348b7" colab={"base_uri": "https://localhost:8080/", "height": 35}
# Tokenize
IDS = TOKENIZER.EncodeAsIds(text)
IDS = onp.asarray(IDS, dtype=onp.int32)
PAD_AMOUNT = 512 * 1024 - len(IDS)
print("Number of tokens:", IDS.shape[0])
# + [markdown] id="bzQ7G9uGSga5" colab_type="text"
# As we see above, "Crime and Punishment" has just over half a million tokens with the BPE vocabulary we have selected.
#
# Normally we would have a dataset with many examples, but for this demonstration we fit a language model on the single novel only. We don't want the model to just memorize the dataset by encoding the words in its position embeddings, so at each training iteration we will randomly select how much padding to put before the text vs. after it.
#
# We have 8 TPU cores, so we will separately randomize the amount of padding for each core.
# + id="PdAwmpS220ub" colab_type="code" outputId="ff1e17a9-f63d-4c02-ac19-877737a5673c" colab={"base_uri": "https://localhost:8080/", "height": 35}
# Set up the data pipeline.
def my_inputs(n_devices):
while True:
inputs = []
mask = []
pad_amounts = onp.random.choice(PAD_AMOUNT, n_devices)
for i in range(n_devices):
inputs.append(onp.pad(IDS, (pad_amounts[i], PAD_AMOUNT - pad_amounts[i]),
mode='constant'))
mask.append(onp.pad(onp.ones_like(IDS, dtype=onp.float32),
(pad_amounts[i], PAD_AMOUNT - pad_amounts[i]),
mode='constant'))
inputs = onp.stack(inputs)
mask = onp.stack(mask)
yield (inputs, inputs, mask)
print("(device count, tokens per device) = ",
next(my_inputs(trax.math.device_count()))[0].shape)
# + id="Ei90LdK024r_" colab_type="code" colab={}
# Configure hyperparameters.
gin.parse_config("""
import trax.layers
import trax.models
import trax.optimizers
import trax.supervised.inputs
import trax.supervised.trainer_lib
# Parameters that will vary between experiments:
# ==============================================================================
train.model = @trax.models.ReformerLM
# Our model will have 6 layers, alternating between the LSH attention proposed
# in the Reformer paper and local attention within a certain context window.
n_layers = 6
attn_type = [
@SelfAttention,
@LSHSelfAttention,
@SelfAttention,
@LSHSelfAttention,
@SelfAttention,
@LSHSelfAttention,
]
share_qk = False # LSH attention ignores this flag and always shares q & k
n_heads = 2
attn_kv = 64
dropout = 0.05
n_tokens = 524288
# Parameters for MultifactorSchedule:
# ==============================================================================
MultifactorSchedule.constant = 0.01
MultifactorSchedule.factors = 'constant * linear_warmup * cosine_decay'
MultifactorSchedule.warmup_steps = 100
MultifactorSchedule.steps_per_cycle = 900
# Parameters for Adam:
# ==============================================================================
Adam.weight_decay_rate=0.0
Adam.b1 = 0.86
Adam.b2 = 0.92
Adam.eps = 1e-9
# Parameters for SelfAttention:
# ==============================================================================
SelfAttention.attention_dropout = 0.05
SelfAttention.chunk_len = 64
SelfAttention.n_chunks_before = 1
SelfAttention.n_parallel_heads = 1
# Parameters for LSHSelfAttention:
# ==============================================================================
LSHSelfAttention.attention_dropout = 0.0
LSHSelfAttention.chunk_len = 64
LSHSelfAttention.n_buckets = [64, 128]
LSHSelfAttention.n_chunks_after = 0
LSHSelfAttention.n_chunks_before = 1
LSHSelfAttention.n_hashes = 1
LSHSelfAttention.n_parallel_heads = 1
LSHSelfAttention.predict_drop_len = 128
LSHSelfAttention.predict_mem_len = 1024
# Parameters for ReformerLM:
# ==============================================================================
ReformerLM.attention_type = %attn_type
ReformerLM.d_attention_key = %attn_kv
ReformerLM.d_attention_value = %attn_kv
ReformerLM.d_model = 256
ReformerLM.d_ff = 512
ReformerLM.dropout = %dropout
ReformerLM.ff_activation = @trax.layers.Relu
ReformerLM.max_len = %n_tokens
ReformerLM.mode = 'train'
ReformerLM.n_heads = %n_heads
ReformerLM.n_layers = %n_layers
ReformerLM.vocab_size = 320
ReformerLM.share_qk = %share_qk
ReformerLM.axial_pos_shape = (512, 1024)
ReformerLM.d_axial_pos_embs= (64, 192)
""")
# + id="RGGt0WaT3a-h" colab_type="code" colab={}
# Set up a Trainer.
output_dir = os.path.expanduser('~/train_dir/')
# !rm -f ~/train_dir/model.pkl # Remove old model
trainer = trax.supervised.Trainer(
model=trax.models.ReformerLM,
loss_fn=trax.layers.CrossEntropyLoss,
optimizer=trax.optimizers.Adam,
lr_schedule=trax.lr.MultifactorSchedule,
inputs=trax.supervised.inputs.Inputs(my_inputs),
output_dir=output_dir,
has_weights=True)
# + id="y6VQkmKO3a1L" colab_type="code" outputId="d5519372-44e9-4311-f84b-931b12e85232" colab={"base_uri": "https://localhost:8080/", "height": 269}
# Run one training step, to make sure the model fits in memory.
# The first time trainer.train_epoch is called, it will JIT the entire network
# architecture, which takes around 2 minutes. The JIT-compiled model is saved
# so subsequent runs will be much faster than the first.
trainer.train_epoch(n_steps=1, n_eval_steps=1)
# + id="EFnX4G6z3asD" colab_type="code" colab={}
# Train for 600 steps total
# The first ~20 steps are slow to run, but after that it reaches steady-state
# speed. This will take at least 30 minutes to run to completion, but can safely
# be interrupted by selecting "Runtime > Interrupt Execution" from the menu.
# The language model won't be exceptionally good when trained for just a few
# steps and with minimal regularization. However, we can still sample from it to
# see what it learns.
trainer.train_epoch(n_steps=9, n_eval_steps=1)
for _ in range(59):
trainer.train_epoch(n_steps=10, n_eval_steps=1)
# + [markdown] id="zY3hpgnI5Rgn" colab_type="text"
# ## Sample from the model
# + id="ffeLSbJk35pv" colab_type="code" colab={}
# As we report in the Reformer paper, increasing the number of hashing rounds
# helps with quality. We can even increase the number of hashing rounds at
# evaluation time only.
gin.parse_config("""LSHCausalAttention.n_hashes = 4""")
# + id="Eq45QGXKG3UG" colab_type="code" colab={}
# Construct the decoder instance.
# Unfortunately this code ends up leaking some memory, so we can only set up the
# decoder once before the memory leak prevents us from running the model and we
# have to restart the notebook.
sampling_decoder = Search(
trax.models.ReformerLM,
trainer.model_weights,
temperature=1.0,
max_decode_len=128,
)
# + id="dfeXilrHHJ6P" colab_type="code" colab={}
# Sample from the Reformer language model.
seqs, scores = sampling_decoder.decode(batch_size=1)
sample = seqs[0, -1]
TOKENIZER.DecodeIds(sample.tolist())
# + id="o31Wtxuu5Ehf" colab_type="code" colab={}
| trax/models/reformer/text_generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/HKang42/DS-Unit-2-Linear-Models/blob/master/module2-regression-2/Harrison_Kang_LS_DS_212_assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="3IJe6Aa1zA5n" colab_type="text"
# Lambda School Data Science
#
# *Unit 2, Sprint 1, Module 2*
#
# ---
# + [markdown] colab_type="text" id="7IXUfiQ2UKj6"
# # Regression 2
#
# ## Assignment
#
# You'll continue to **predict how much it costs to rent an apartment in NYC,** using the dataset from renthop.com.
#
# - [ ] Do train/test split. Use data from April & May 2016 to train. Use data from June 2016 to test.
# - [ ] Engineer at least two new features. (See below for explanation & ideas.)
# - [ ] Fit a linear regression model with at least two features.
# - [ ] Get the model's coefficients and intercept.
# - [ ] Get regression metrics RMSE, MAE, and $R^2$, for both the train and test data.
# - [ ] What's the best test MAE you can get? Share your score and features used with your cohort on Slack!
# - [ ] As always, commit your notebook to your fork of the GitHub repo.
#
#
# #### [Feature Engineering](https://en.wikipedia.org/wiki/Feature_engineering)
#
# > "Some machine learning projects succeed and some fail. What makes the difference? Easily the most important factor is the features used." — <NAME>, ["A Few Useful Things to Know about Machine Learning"](https://homes.cs.washington.edu/~pedrod/papers/cacm12.pdf)
#
# > "Coming up with features is difficult, time-consuming, requires expert knowledge. 'Applied machine learning' is basically feature engineering." — <NAME>, [Machine Learning and AI via Brain simulations](https://forum.stanford.edu/events/2011/2011slides/plenary/2011plenaryNg.pdf)
#
# > Feature engineering is the process of using domain knowledge of the data to create features that make machine learning algorithms work.
#
# #### Feature Ideas
# - Does the apartment have a description?
# - How long is the description?
# - How many total perks does each apartment have?
# - Are cats _or_ dogs allowed?
# - Are cats _and_ dogs allowed?
# - Total number of rooms (beds + baths)
# - Ratio of beds to baths
# - What's the neighborhood, based on address or latitude & longitude?
#
# ## Stretch Goals
# - [ ] If you want more math, skim [_An Introduction to Statistical Learning_](http://faculty.marshall.usc.edu/gareth-james/ISL/ISLR%20Seventh%20Printing.pdf), Chapter 3.1, Simple Linear Regression, & Chapter 3.2, Multiple Linear Regression
# - [ ] If you want more introduction, watch [<NAME>, Statistics 101: Simple Linear Regression](https://www.youtube.com/watch?v=ZkjP5RJLQF4)
# (20 minutes, over 1 million views)
# - [ ] Add your own stretch goal(s) !
# + colab_type="code" id="o9eSnDYhUGD7" colab={}
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# Ignore this Numpy warning when using Plotly Express:
# FutureWarning: Method .ptp is deprecated and will be removed in a future version. Use numpy.ptp instead.
import warnings
warnings.filterwarnings(action='ignore', category=FutureWarning, module='numpy')
# + colab_type="code" id="cvrw-T3bZOuW" colab={}
import numpy as np
import pandas as pd
# Read New York City apartment rental listing data
df = pd.read_csv(DATA_PATH+'apartments/renthop-nyc.csv')
assert df.shape == (49352, 34)
# Remove the most extreme 1% prices,
# the most extreme .1% latitudes, &
# the most extreme .1% longitudes
df = df[(df['price'] >= np.percentile(df['price'], 0.5)) &
(df['price'] <= np.percentile(df['price'], 99.5)) &
(df['latitude'] >= np.percentile(df['latitude'], 0.05)) &
(df['latitude'] < np.percentile(df['latitude'], 99.95)) &
(df['longitude'] >= np.percentile(df['longitude'], 0.05)) &
(df['longitude'] <= np.percentile(df['longitude'], 99.95))]
# + [markdown] id="U86NC8ID40bO" colab_type="text"
# # Engineer at least two new features.
#
# - I don't want to split first. Otherwise the features I engineer won't show up in the split data frames
# + id="tKKjm96qgbHT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 513} outputId="2104e797-c78b-41e3-9071-f375afe4794b"
df.head()
# + id="EdsfvTQw4194" colab_type="code" colab={}
# Let's create a category for any apartment that allows a cat OR a dog
df['catordog'] = ((df['cats_allowed'] == 1 ) | (df['dogs_allowed'] == 1) )
# + id="0_6clNcENujZ" colab_type="code" colab={}
# convert catordog to a numerical column
df['catordog'] = df['catordog'].replace({True:1, False:0})
# + id="aBQC6Rk_O8IT" colab_type="code" colab={}
# let's make a feature for bathrooms per bedroom
# first let's set any 0 values to 0.5
df['bath'] = df['bathrooms'].astype(float).replace( { 0 : 0.5 } )
df['bed'] = df['bedrooms'].astype(float).replace( { 0 : 0.5 } )
# + id="HNqbVm1_cV0B" colab_type="code" colab={}
df[ 'bathperbed' ] = df['bath'] / df['bed']
# + [markdown] id="ay30nWua17Mw" colab_type="text"
# # Do train/test split. Use data from April & May 2016 to train. Use data from June 2016 to test.
# + id="apSV9_0O2X_O" colab_type="code" colab={}
train = df [ (df['created'] > '2016-04-01') & (df['created'] < '2016-06-01') ]
test = df [ (df['created'] < '2016-04-01') | (df['created'] > '2016-06-01') ]
# + [markdown] id="q-7ZhUGlZfBR" colab_type="text"
# # Fit a linear regression model with at least two features.
# + id="7v8sBW8cZXOA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4fe1d864-7b47-4b74-e746-41499936ae3c"
from sklearn.linear_model import LinearRegression
model = LinearRegression()
x = ['catordog' , 'bathperbed']
y = ['price']
model.fit( train[x], train[y] )
# + [markdown] id="8NQ2lnP-dt08" colab_type="text"
# # Get the model's coefficients and intercept.
# + id="1P-qunEKdrF3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="6f617e43-1915-40a8-ab17-5730f030365e"
catdog_coeff = model.coef_[0][0]
bathbed_coeff = model.coef_[0][1]
intercept = model.intercept_[0]
print('Cat or Dog coefficient is:', catdog_coeff)
print('Bath per bed coefficient is:', bathbed_coeff)
print('The y intercept is:', intercept)
# + [markdown] id="gOPf2skTd3_i" colab_type="text"
# # Get regression metrics RMSE, MAE, and R2 , for both the train and test data.
# + id="irqPIVkCgsl4" colab_type="code" colab={}
# now let's make our predictions for whatever we set our features equal to
predict = model.predict( train[x] )
# + id="oEiA3L8QhSMO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="45ee88d6-cff3-4e02-b6ad-656cfeaf21ad"
from sklearn.metrics import mean_absolute_error, mean_squared_error,r2_score
mae = mean_absolute_error(train['price'], predict)
mse = mean_squared_error(train['price'], predict)
rmse = np.sqrt(mse)
r2 = r2_score(train['price'], predict)
print('For our training data:')
print('Our mean absolute error is: $' + str(mae.round(2)))
print('Our root mean square error is: $' + str(rmse.round(2)))
print('Our r^2 value is: ' + str(r2.round(4)))
# + id="XFVL8sd8jhj_" colab_type="code" colab={}
# now let's see how our model looks for the test data
predict = model.predict( test[x] )
# + id="8kgQHp4hjnUa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="165257e8-b565-4995-a361-29476da0e2c1"
mae = mean_absolute_error(test['price'], predict)
mse = mean_squared_error(test['price'], predict)
rmse = np.sqrt(mse)
r2 = r2_score(test['price'], predict)
print('For our test data:')
print('Our mean absolute error is: $' + str(mae.round(2)))
print('Our root mean square error is: $' + str(rmse.round(2)))
print('Our r^2 value is: ' + str(r2.round(4)))
# + [markdown] id="R5HGTukioLMp" colab_type="text"
# I looks like the training and testing error is roughly the same which is a decent result. Let's get descriptive statistics for the price and see how big the error is.
# + id="ts3-QGSjoUyw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="003af2c6-234f-43dd-f63b-1dfbfe74c985"
df['price'].describe()
# + [markdown] id="yk3XIPVRo7Z0" colab_type="text"
# Given that the mean price is about $3,600, our errors for both data sets are pretty big. Let's try again but with different features.
# + id="sJYgsnPPqBvv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="522fc173-74a2-4911-ab4e-6ac9c98acf74"
from sklearn.linear_model import LinearRegression
model = LinearRegression()
x = ['bedrooms' , 'balcony']
y = ['price']
model.fit( train[x], train[y] )
# + id="PGUJRX3OsiBC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="9f5fa840-80e1-4075-84b4-f464c96b5afc"
bedroom_coeff = model.coef_[0][0]
balcony_coeff = model.coef_[0][1]
intercept = model.intercept_[0]
print('Cat or Dog coefficient is:', bedroom_coeff)
print('Bath per bed coefficient is:', balcony_coeff)
print('The y intercept is:', intercept)
# + id="_uxTKWsotsr_" colab_type="code" colab={}
predict = model.predict( train[x] )
# + id="PPOXDeC7sm4Y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="8916a5d3-fd5e-484a-88f0-071dddbdcd20"
from sklearn.metrics import mean_absolute_error, mean_squared_error,r2_score
mae = mean_absolute_error(train['price'], predict)
mse = mean_squared_error(train['price'], predict)
rmse = np.sqrt(mse)
r2 = r2_score(train['price'], predict)
print('For our training data:')
print('Our mean absolute error is: $' + str(mae.round(2)))
print('Our root mean square error is: $' + str(rmse.round(2)))
print('Our r^2 value is: ' + str(r2.round(4)))
# + id="9SIZoh38sqjZ" colab_type="code" colab={}
# now let's see how our model looks for the test data
predict = model.predict( test[x] )
# + id="yV_uQgP2stDp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="11a23809-feb5-406b-b1b2-611531782f61"
mae = mean_absolute_error(test['price'], predict)
mse = mean_squared_error(test['price'], predict)
rmse = np.sqrt(mse)
r2 = r2_score(test['price'], predict)
print('For our test data:')
print('Our mean absolute error is: $' + str(mae.round(2)))
print('Our root mean square error is: $' + str(rmse.round(2)))
print('Our r^2 value is: ' + str(r2.round(4)))
# + [markdown] id="HiZvVmsqt5km" colab_type="text"
# It looks like bedrooms and balconies are a slightly better pair of features based on the error values.
| module2-regression-2/Harrison_Kang_LS_DS_212_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.12 64-bit (''DataCamp'': conda)'
# language: python
# name: python3
# ---
# # Predicting Customer Churn in python: Data Preprocessing
# ## The Dataset
# In this project, we will learn how to build a churn model from beginning to end. The data we will be using comes from a Cellular Usage dataset that consists of records of actual Cell Phone customers, and features that include:
# * voice mail
# * international calling
# * cost for the service
# * customer usage
# * customer churn
#
import pandas as pd
telco_df=pd.read_csv('./Churn.csv')
telco_df
# ## Identifying features to convert
# It is preferable to have features like 'Churn' encoded as 0 and 1 instead of no and yes, so that we can then feed it into machine learning algorithms that only accept numeric values.
# Besides 'Churn', other features that are of type object can be converted into 0s and 1s. In the following, we will explore the different data types of telco in the IPython Shell and identify the ones that are of type object.
print(telco_df.dtypes)
# ## Encoding binary features
# Recasting data types is an important part of data preprocessing. In this exercise you will assign the values 1 to 'yes' and 0 to 'no' to the 'Vmail_Plan' and 'Churn' features, respectively.
from sklearn.preprocessing import LabelEncoder
le=LabelEncoder()
telco_df['Intl_Plan']=le.fit_transform(telco_df['Intl_Plan'])
telco_df['Intl_Plan'].head()
telco_df['Vmail_Plan']=le.fit_transform(telco_df['Vmail_Plan'])
telco_df['Vmail_Plan'].head()
telco_df['Churn']=le.fit_transform(telco_df['Churn'])
telco_df['Churn'].head()
# ## One hot encoding
# +
# Perform one hot encoding on 'State'
telco_state = pd.get_dummies(telco_df['State'])
# Print the head of telco_state
telco_state.head()
# -
telco_df.drop(columns=['State'],inplace=True)
telco_df=pd.concat([telco_df,telco_state],axis=1)
telco_df.head()
# ## Feature Scaling
# Let's investigate the different scales of the 'Intl_Calls' and 'Night_Mins'.
telco_df['Intl_Calls'].describe()
telco_df['Night_Mins'].describe()
# Here we will re-scale them using StandardScaler.
# +
# Import StandardScaler
from sklearn.preprocessing import StandardScaler
# Scale telco using StandardScaler
telco_scaled = StandardScaler().fit_transform(telco_df[["Intl_Calls", "Night_Mins"]])
# Add column names back for readability
telco_scaled_df = pd.DataFrame(telco_scaled, columns=["Intl_Calls", "Night_Mins"])
# Print summary statistics
print(telco_scaled_df.describe())
# +
#telco_df[["Intl_Calls", "Night_Mins"]]=telco_scaled_df
#telco_df.head()
# -
# ## Dropping unnecessary features
# +
# Drop the unnecessary features
telco_df = telco_df.drop(telco_df[['Area_Code','Phone']], axis=1)
# Verify dropped features
print(telco_df.columns)
# -
# ## Engineering a new column
# Leveraging domain knowledge to engineer new features is an essential part of modeling.
#
# This quote from <NAME> summarizes the importance of feature engineering:
#
# _Coming up with features is difficult, time-consuming, requires expert knowledge. "Applied machine learning" is basically feature engineering._
#
# Here, we will create a new feature that contains information about the average length of night calls made by customers.
# +
# Create the new feature
telco_df['Avg_Night_Calls'] = telco_df['Night_Mins']/telco_df['Night_Calls']
# Print the first five rows of 'Avg_Night_Calls'
print(telco_df['Avg_Night_Calls'].head())
# -
telco_df.to_csv('./telco_preprocessed.csv',index=False)
| 02-Data Preprocessing/Data Preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
import json
# +
json_path = "/home/chen/workshop/thesis/human-pose-estimation.pytorch/data/crowdpose/annotations/crowdpose_train.json"
j = json.load(open(json_path))
num_vis = 0
num_unvis = 0
num_all = 0
count_list = [[0, 0, 0] for i in range(14)]
for item in j['annotations']:
kps = item['keypoints']
for idx, start in enumerate (range(0,14*3,3)):
i, j, v = kps[start:start+3]
num_all += 1
count_list[idx][2] += 1
if v == 1:
count_list[idx][1] += 1
num_unvis += 1
elif v == 2:
count_list[idx][0] += 1
num_vis += 1
num_vis / num_unvis
# -
for v, uv, a in count_list:
print(v / uv)
num_vis / num_all
num_unvis / num_all
| scripts/stat_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ''
# language: python
# name: python3
# ---
# +
from copy import deepcopy
import pandas as pd
from BitBoard import osero
from osero_learn import learn
PLAY_WAY = deepcopy(osero.PLAY_WAY)
del PLAY_WAY["human"]
PLAY_WAY = PLAY_WAY.values()
# -
eva = [[
1.0, -0.6, 0.6, 0.4, 0.4, 0.6, -0.6, 1.0,
-0.6, -0.8, 0.0, 0.0, 0.0, 0.0, -0.8, -0.6,
0.6, 0.0, 0.8, 0.6, 0.6, 0.8, 0.0, 0.6,
0.4, 0.0, 0.6, 0.0, 0.0, 0.6, 0.0, 0.4,
0.4, 0.0, 0.6, 0.0, 0.0, 0.6, 0.0, 0.4,
0.6, 0.0, 0.8, 0.6, 0.6, 0.8, 0.0, 0.6,
-0.6, -0.8, 0.0, 0.0, 0.0, 0.0, -0.8, -0.6,
1.0, -0.6, 0.6, 0.4, 0.4, 0.6, -0.6, 1.0
] for i in range(2)]
# +
df = pd.DataFrame({})
check_point = [i for i in range(5, 64, 5)]
for i in range(10):
print("\r[" + "#" * (i+1) + " " * (10-i+1) + "]", end="")
for black in PLAY_WAY:
for white in PLAY_WAY:
run = learn(\
black,
white,
check_point=check_point,
seed_num=i,
eva=eva
)
data = run.play()
df = df.append(data, ignore_index=True)
print("\r[" + "#" * 10 + "]")
# +
import matplotlib.pyplot as plt
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_absolute_error
from sklearn.linear_model import Ridge
x_data = df.drop(["turn", "last_score"], axis=1)
y_data = df[["turn_num", "last_score"]]
turn_vari = df["turn_num"].unique()
drop_vari = ["custom_score and score", "custom_score", "score", "None"]
# +
train_score = []
test_score = []
train_MAE = []
test_MAE = []
for turn_num in turn_vari:
train_score.append([])
test_score.append([])
train_MAE.append([])
test_MAE.append([])
for drop_list in [["custom_score", "score"], ["custom_score"], ["score"], []]:
drop_list.append("turn_num")
x_train, x_test, y_train, y_test = train_test_split(\
x_data.query("turn_num==%d" % turn_num).drop(drop_list, axis=1),
y_data.query("turn_num==%d" % turn_num).drop("turn_num", axis=1),
test_size=0.3,
random_state=0
)
model = Ridge(random_state=0)
model.fit(x_train, y_train)
train_score[-1].append(model.score(x_train, y_train))
test_score[-1].append(model.score(x_test, y_test))
train_predict = model.predict(x_train)
train_MAE[-1].append(mean_absolute_error(train_predict, y_train))
test_predict = model.predict(x_test)
test_MAE[-1].append(mean_absolute_error(test_predict, y_test))
# +
width = 0.3
x_axis = np.array([i + 1 for i in range(len(drop_vari))])
for i in range(len(turn_vari)):
fig_name = "score of each dropping (number of turn is %d)" % turn_vari[i]
fig = plt.figure(figsize=(10, 10))
plt.bar(x_axis, train_score[i], label="train score", width=width)
plt.bar(x_axis + width, test_score[i], label="test score", width=width)
plt.xticks(x_axis + width/2, labels=drop_vari, rotation=15)
plt.legend()
plt.title(fig_name)
plt.xlabel("dropped column")
plt.ylabel("score")
plt.savefig("fig/" + fig_name)
# plt.show()
plt.clf()
plt.close()
fig_name = "MAE of each dropping (number of turn is %d)" % turn_vari[i]
fig = plt.figure(figsize=(10, 10))
plt.bar(x_axis, train_MAE[i], label="train MAE", width=width)
plt.bar(x_axis + width, test_MAE[i], label="test MAE", width=width)
plt.xticks(x_axis + width/2, labels=drop_vari, rotation=15)
plt.legend()
plt.title(fig_name)
plt.xlabel("dropped column")
plt.ylabel("mean absolute error")
plt.savefig("fig/" + fig_name)
# plt.show()
plt.clf()
plt.close()
# +
x_axis = np.array([i + 1 for i in range(len(turn_vari))])
x_axis_name = [str(i) for i in turn_vari]
train_score_T = np.array(train_score).T
test_score_T = np.array(test_score).T
train_MAE_T = np.array(train_MAE).T
test_MAE_T = np.array(test_MAE).T
for i in range(len(drop_vari)):
fig_name = "score of each turn number (dropped column is %s)" % drop_vari[i]
fig = plt.figure(figsize=(10, 10))
plt.plot(x_axis_name, train_score_T[i], label="train score")
plt.plot(x_axis_name, test_score_T[i], label="test score")
plt.legend()
plt.title(fig_name)
plt.xlabel("turn number")
plt.ylabel("score")
plt.savefig("fig/" + fig_name)
# plt.show()
plt.clf()
plt.close()
fig_name = "MAE of each turn number (dropped column is %s)" % drop_vari[i]
fig = plt.figure(figsize=(10, 10))
plt.plot(x_axis_name, train_MAE_T[i], label="train MAE")
plt.plot(x_axis_name, test_MAE_T[i], label="test MAE")
plt.legend()
plt.title(fig_name)
plt.xlabel("turn number")
plt.ylabel("mean absolute error")
plt.savefig("fig/" + fig_name)
# plt.show()
plt.clf()
plt.close()
| mecha_learn/28/02/run.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Title:** 22 minutes to 2nd place in a Kaggle competition with deep learning and the Microsoft Azure cloud
# **Author:** <NAME>, [PyImageSearch.com](https://www.pyimagesearch.com/)
#
# The code in this tutorial was executed on a [Microsoft Data Science Virtual Machine](https://azure.microsoft.com/en-us/services/virtual-machines/data-science-virtual-machines/). Results were gathered for Microsoft's blog.
#
# For an in-depth treatment of deep learning and Convolutional Neural Networks, please refer to [*Deep Learning for Computer Vision with Python*](https://www.pyimagesearch.com/deep-learning-computer-vision-python-book/).
# import the necessary packages
from keras.applications import ResNet50
from keras.applications import imagenet_utils
from keras.preprocessing.image import img_to_array
from keras.preprocessing.image import load_img
from sklearn.preprocessing import LabelEncoder
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import classification_report
from sklearn.metrics import accuracy_score
from imutils import paths
import numpy as np
import progressbar
import random
import os
# +
# since we are not using command line arguments (like we typically
# would inside Deep Learning for Computer Vision with Python, let's
# "pretend" we are by using an `args` dictionary -- this will enable
# us to easily reuse and swap out code depending if we are using the
# command line or Jupyter Notebook
args = {
"dataset": "../datasets/kaggle_dogs_vs_cats/train",
"batch_size": 32,
}
# store the batch size in a convenience variable
bs = args["batch_size"]
# -
# grab the list of images in the Kaggle Dogs vs. Cats download and
# shuffle them to allow for easy training and testing splits via
# array slicing during training time
imagePaths = list(paths.list_images(args["dataset"]))
random.shuffle(imagePaths)
print(len(imagePaths))
# extract the class labels from the image paths then encode the
# labels
labels = [p.split(os.path.sep)[-1].split(".")[0] for p in imagePaths]
le = LabelEncoder()
labels = le.fit_transform(labels)
# load the ResNet50 network (i.e., the network we'll be using for
# feature extraction)
model = ResNet50(weights="imagenet", include_top=False)
# initialize the progress bar
widgets = ["Extracting Features: ", progressbar.Percentage(), " ",
progressbar.Bar(), " ", progressbar.ETA()]
pbar = progressbar.ProgressBar(maxval=len(imagePaths),
widgets=widgets).start()
# +
# %%time
# initialize our data matrix (where we will store our extracted
# features)
data = None
# loop over the images in batches
for i in np.arange(0, len(imagePaths), bs):
# extract the batch of images and labels, then initialize the
# list of actual images that will be passed through the network
# for feature extraction
batchPaths = imagePaths[i:i + bs]
batchLabels = labels[i:i + bs]
batchImages = []
# loop over the images and labels in the current batch
for (j, imagePath) in enumerate(batchPaths):
# load the input image using the Keras helper utility
# while ensuring the image is resized to 224x224 pixels
image = load_img(imagePath, target_size=(224, 224))
image = img_to_array(image)
# preprocess the image by (1) expanding the dimensions and
# (2) subtracting the mean RGB pixel intensity from the
# ImageNet dataset
image = np.expand_dims(image, axis=0)
image = imagenet_utils.preprocess_input(image)
# add the image to the batch
batchImages.append(image)
# pass the images through the network and use the outputs as
# our actual features
batchImages = np.vstack(batchImages)
features = model.predict(batchImages, batch_size=bs)
# reshape the features so that each image is represented by
# a flattened feature vector of the `MaxPooling2D` outputs
features = features.reshape((features.shape[0], 2048))
# if our data matrix is None, initialize it
if data is None:
data = features
# otherwise, stack the data and features together
else:
data = np.vstack([data, features])
# update the progress bar
pbar.update(i)
# finish up the progress bar
pbar.finish()
# -
# show the data matrix shape and amount of memory it consumes
print(data.shape)
print(data.nbytes)
# +
# %%time
# determine the index of the training and testing split (75% for
# training and 25% for testing)
i = int(data.shape[0] * 0.75)
# define the set of parameters that we want to tune then start a
# grid search where we evaluate our model for each value of C
print("[INFO] tuning hyperparameters...")
params = {"C": [0.0001, 0.001, 0.01, 0.1, 1.0]}
clf = GridSearchCV(LogisticRegression(), params, cv=3, n_jobs=-1)
clf.fit(data[:i], labels[:i])
print("[INFO] best hyperparameters: {}".format(clf.best_params_))
# +
# generate a classification report for the model
print("[INFO] evaluating...")
preds = clf.predict(data[i:])
print(classification_report(labels[i:], preds, target_names=le.classes_))
# compute the raw accuracy with extra precision
acc = accuracy_score(labels[i:], preds)
print("[INFO] score: {}".format(acc))
# -
| pyimagesearch-22-minutes-to-2nd-place.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/adventurousAyan/AyanRepo/blob/master/Model_LGBM.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="jg0fHQeByBwr"
import gc
import os
import random
import lightgbm as lgb
import numpy as np
import pandas as pd
import seaborn as sns
from matplotlib import pyplot as plt
from sklearn.metrics import mean_squared_error
from sklearn.preprocessing import LabelEncoder
path_data = "/content/drive/MyDrive/Colab Notebooks/datasets/"
path_train = path_data + "train_data.csv"
path_test = path_data + "test_data.csv"
path_building = path_data + "farm_data.csv"
path_weather_train = path_data + "train_weather.csv"
path_weather_test = path_data + "test_weather.csv"
plt.style.use("seaborn")
sns.set(font_scale=1)
myfavouritenumber = 0
seed = myfavouritenumber
random.seed(seed)
# + id="oCse_pzYyHq5"
df_train = pd.read_csv(path_train)
building = pd.read_csv(path_building)
le = LabelEncoder()
building.farming_company = le.fit_transform(building.farming_company)
weather_train = pd.read_csv(path_weather_train)
# + id="L5mSubAo1bAu"
## Memory optimization
# Original code from https://www.kaggle.com/gemartin/load-data-reduce-memory-usage by @gemartin
# Modified to support timestamp type, categorical type
# Modified to add option to use float16
from pandas.api.types import is_datetime64_any_dtype as is_datetime
from pandas.api.types import is_categorical_dtype
def reduce_mem_usage(df, use_float16=False):
"""
Iterate through all the columns of a dataframe and modify the data type to reduce memory usage.
"""
start_mem = df.memory_usage().sum() / 1024**2
print("Memory usage of dataframe is {:.2f} MB".format(start_mem))
for col in df.columns:
if is_datetime(df[col]) or is_categorical_dtype(df[col]):
continue
col_type = df[col].dtype
if col_type != object:
c_min = df[col].min()
c_max = df[col].max()
if str(col_type)[:3] == "int":
if c_min > np.iinfo(np.int8).min and c_max < np.iinfo(np.int8).max:
df[col] = df[col].astype(np.int8)
elif c_min > np.iinfo(np.int16).min and c_max < np.iinfo(np.int16).max:
df[col] = df[col].astype(np.int16)
elif c_min > np.iinfo(np.int32).min and c_max < np.iinfo(np.int32).max:
df[col] = df[col].astype(np.int32)
elif c_min > np.iinfo(np.int64).min and c_max < np.iinfo(np.int64).max:
df[col] = df[col].astype(np.int64)
else:
if use_float16 and c_min > np.finfo(np.float16).min and c_max < np.finfo(np.float16).max:
df[col] = df[col].astype(np.float16)
elif c_min > np.finfo(np.float32).min and c_max < np.finfo(np.float32).max:
df[col] = df[col].astype(np.float32)
else:
df[col] = df[col].astype(np.float64)
else:
df[col] = df[col].astype("category")
end_mem = df.memory_usage().sum() / 1024**2
print("Memory usage after optimization is: {:.2f} MB".format(end_mem))
print("Decreased by {:.1f}%".format(100 * (start_mem - end_mem) / start_mem))
return df
# + colab={"base_uri": "https://localhost:8080/"} id="bJ2CL02T1weG" outputId="c23159a2-ee6e-4652-9e1c-f1766ffb74a9"
df_train = reduce_mem_usage(df_train, use_float16=True)
building = reduce_mem_usage(building, use_float16=True)
weather_train = reduce_mem_usage(weather_train, use_float16=True)
# + id="kKOebaX819YY"
def prepare_data(X, building_data, weather_data, test=False):
"""
Preparing final dataset with all features.
"""
X = X.merge(building_data, on="farm_id", how="left")
X = X.merge(weather_data, on=["deidentified_location", "timestamp"], how="left")
X.timestamp = pd.to_datetime(X.timestamp, format="%Y-%m-%d %H:%M:%S")
X.farm_area = np.log1p(X.farm_area)
if not test:
X.sort_values("timestamp", inplace=True)
X.reset_index(drop=True, inplace=True)
gc.collect()
holidays = ["2016-01-01", "2016-01-18", "2016-02-15", "2016-05-30", "2016-07-04",
"2016-09-05", "2016-10-10", "2016-11-11", "2016-11-24", "2016-12-26",
"2017-01-01", "2017-01-16", "2017-02-20", "2017-05-29", "2017-07-04",
"2017-09-04", "2017-10-09", "2017-11-10", "2017-11-23", "2017-12-25",
"2018-01-01", "2018-01-15", "2018-02-19", "2018-05-28", "2018-07-04",
"2018-09-03", "2018-10-08", "2018-11-12", "2018-11-22", "2018-12-25",
"2019-01-01"]
X["hour"] = X.timestamp.dt.hour
X["weekday"] = X.timestamp.dt.weekday
#X["is_holiday"] = (X.timestamp.dt.date.astype("str").isin(holidays)).astype(int)
drop_features = ["timestamp", "cloudiness", "dew_temp"]
X.drop(drop_features, axis=1, inplace=True)
if test:
row_ids = X.id
X.drop("id", axis=1, inplace=True)
return X, row_ids
else:
y = np.log1p(X['yield'])
X.drop("yield", axis=1, inplace=True)
return X, y
# + colab={"base_uri": "https://localhost:8080/"} id="Z2QqwabV26HS" outputId="0b338771-5c55-445f-f130-9c016f4b8fe2"
df_train.columns
# + id="WWN3LTQv27w-"
df_train.columns = ['timestamp', 'farm_id', 'ingredient_type', 'yield']
# + colab={"base_uri": "https://localhost:8080/"} id="QxFAqwbb3GMg" outputId="6cda8e2f-f6f5-4130-9c67-ee16b26942b6"
X_train, y_train = prepare_data(df_train, building, weather_train)
del df_train, weather_train
gc.collect()
# + colab={"base_uri": "https://localhost:8080/"} id="788NlMbp3KYA" outputId="d26c7e36-094d-4ddf-d1b3-4bc1932cc6b0"
X_half_1 = X_train[:int(X_train.shape[0] / 2)]
X_half_2 = X_train[int(X_train.shape[0] / 2):]
y_half_1 = y_train[:int(X_train.shape[0] / 2)]
y_half_2 = y_train[int(X_train.shape[0] / 2):]
categorical_features = ["farm_id", "deidentified_location", "ingredient_type", "farming_company", "hour", "weekday"]
d_half_1 = lgb.Dataset(X_half_1, label=y_half_1, categorical_feature=categorical_features, free_raw_data=False)
d_half_2 = lgb.Dataset(X_half_2, label=y_half_2, categorical_feature=categorical_features, free_raw_data=False)
watchlist_1 = [d_half_1, d_half_2]
watchlist_2 = [d_half_2, d_half_1]
params = {
"objective": "regression",
"boosting": "gbdt",
"num_leaves": 40,
"learning_rate": 0.05,
"feature_fraction": 0.85,
"reg_lambda": 2,
"metric": "rmse"
}
print("Building model with first half and validating on second half:")
model_half_1 = lgb.train(params, train_set=d_half_1, num_boost_round=1000, valid_sets=watchlist_1, verbose_eval=200, early_stopping_rounds=200)
print("Building model with second half and validating on first half:")
model_half_2 = lgb.train(params, train_set=d_half_2, num_boost_round=1000, valid_sets=watchlist_2, verbose_eval=200, early_stopping_rounds=200)
# + colab={"base_uri": "https://localhost:8080/", "height": 509} id="rEZH8Eaw3_B3" outputId="6fd6bd8b-2404-47f2-e3be-44edd4288836"
df_fimp_1 = pd.DataFrame()
df_fimp_1["feature"] = X_train.columns.values
df_fimp_1["importance"] = model_half_1.feature_importance()
df_fimp_1["half"] = 1
df_fimp_2 = pd.DataFrame()
df_fimp_2["feature"] = X_train.columns.values
df_fimp_2["importance"] = model_half_2.feature_importance()
df_fimp_2["half"] = 2
df_fimp = pd.concat([df_fimp_1, df_fimp_2], axis=0)
plt.figure(figsize=(14, 7))
sns.barplot(x="importance", y="feature", data=df_fimp.sort_values(by="importance", ascending=False))
plt.title("LightGBM Feature Importance")
plt.tight_layout()
# + colab={"base_uri": "https://localhost:8080/"} id="68qAJ4_M_H5u" outputId="98b93d66-864b-43b9-8467-ebff827384c9"
df_test.columns
# + id="DCOWtHEe_Eu3"
#
# + colab={"base_uri": "https://localhost:8080/"} id="RdTvdZs8-hSN" outputId="d0fd1823-cfc4-4e3a-e0b7-319a3419ee23"
df_test = pd.read_csv(path_test)
weather_test = pd.read_csv(path_weather_test)
df_test = reduce_mem_usage(df_test)
df_test.columns = ['timestamp', 'farm_id', 'ingredient_type', 'id']
weather_test = reduce_mem_usage(weather_test)
X_test, row_ids = prepare_data(df_test, building, weather_test, test=True)
# + id="IDueJdbL-qDp"
pred = np.expm1(model_half_1.predict(X_test, num_iteration=model_half_1.best_iteration)) / 2
del model_half_1
#gc.collect()
pred += np.expm1(model_half_2.predict(X_test, num_iteration=model_half_2.best_iteration)) / 2
del model_half_2
#gc.collect()
# + id="51zKRrhp_5X3"
submission = pd.DataFrame({"id": row_ids, "yield": np.clip(pred, 0, a_max=None)})
submission.to_csv("/content/drive/MyDrive/Colab Notebooks/datasets/Model_LGBM.csv", index=False)
# + colab={"base_uri": "https://localhost:8080/"} id="Rma8Wt_rIiE7" outputId="46486501-13b7-4122-af06-557887c4d874"
submission.shape
# + colab={"base_uri": "https://localhost:8080/"} id="ugLkAZUIJBH4" outputId="79a0e6fc-1644-4e5d-9853-307454fbc014"
submission.id.nunique()
# + id="rLs-9jrXJLQo"
| Model_LGBM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow import keras
import seaborn as sns
from os.path import join
plt.style.use(["seaborn", "thesis"])
# -
plt.rc("figure", figsize=(8,4))
model_path = "../../thesis/models/DescriptorHomo/CC/"
# # Molecules
# +
from SCFInitialGuess.utilities.dataset import ScreenedData
target = "P"
basis = "6-311++g**"
data = ScreenedData(r_max=10)
data.include(data_path = "../../thesis/dataset/MethanT/", postfix = "MethanT", target=target)
# +
from SCFInitialGuess.utilities.dataset import ScreenedData
target = "P"
data = ScreenedData(r_max=10)
data.include(data_path = "../../thesis/dataset/MethanT/", postfix = "MethanT", target=target)
data.include(data_path = "../../thesis/dataset/MethanT2/", postfix = "MethanT2", target=target)
data.include(data_path = "../../thesis/dataset/MethanT3/", postfix = "MethanT3", target=target)
data.include(data_path = "../../thesis/dataset/MethanT4/", postfix = "MethanT4", target=target)
data.include(data_path = "../../thesis/dataset/EthanT/", postfix = "EthanT", target=target)
data.include(data_path = "../../thesis/dataset/EthanT2/", postfix = "EthanT2", target=target)
data.include(data_path = "../../thesis/dataset/EthanT3/", postfix = "EthanT3", target=target)
data.include(data_path = "../../thesis/dataset/EthanT4/", postfix = "EthanT4", target=target)
data.include(data_path = "../../thesis/dataset/EthanT5/", postfix = "EthanT5", target=target)
data.include(data_path = "../../thesis/dataset/EthanT6/", postfix = "EthanT6", target=target)
data.include(data_path = "../../thesis/dataset/EthenT/", postfix = "EthenT", target=target)
data.include(data_path = "../../thesis/dataset/EthenT2/", postfix = "EthenT2", target=target)
data.include(data_path = "../../thesis/dataset/EthenT3/", postfix = "EthenT3", target=target)
data.include(data_path = "../../thesis/dataset/EthenT4/", postfix = "EthenT4", target=target)
data.include(data_path = "../../thesis/dataset/EthenT5/", postfix = "EthenT5", target=target)
data.include(data_path = "../../thesis/dataset/EthenT6/", postfix = "EthenT6", target=target)
data.include(data_path = "../../thesis/dataset/EthinT/", postfix = "EthinT", target=target)
data.include(data_path = "../../thesis/dataset/EthinT2/", postfix = "EthinT2", target=target)
data.include(data_path = "../../thesis/dataset/EthinT3/", postfix = "EthinT3", target=target)
#data.include(data_path = "../../dataset/QM9/", postfix = "QM9-300")
# -
# # Descriptor
# +
from SCFInitialGuess.descriptors.high_level import AtomicNumberWeighted
from SCFInitialGuess.descriptors.cutoffs import BehlerCutoff1
from SCFInitialGuess.descriptors.models import RADIAL_GAUSSIAN_MODELS, make_uniform
from SCFInitialGuess.descriptors.coordinate_descriptors import \
Gaussians, SPHAngularDescriptor
import pickle
model = make_uniform(25, 5, eta_max=60, eta_min=20)
descriptor = AtomicNumberWeighted(
Gaussians(*model),
SPHAngularDescriptor(4),
BehlerCutoff1(5)
)
pickle.dump(descriptor, open(model_path + "descriptor.dump", "wb"))
descriptor.radial_descriptor.number_of_descriptors, descriptor.angular_descriptor.number_of_descriptors, descriptor.number_of_descriptors
# -
# # Package Dataset
# +
from SCFInitialGuess.utilities.dataset import make_block_dataset, extract_HOMO_block_dataset_pairs
dataset = make_block_dataset(
descriptor,
data.molecules,
data.T,
"C",
extract_HOMO_block_dataset_pairs
)
np.save(model_path + "normalisation.npy", (dataset.x_mean, dataset.x_std))
# -
len(dataset.training[0]), len(dataset.validation[0]), len(dataset.testing[0]),
# +
from SCFInitialGuess.utilities.constants import number_of_basis_functions as N_BASIS
species = "C"
dim = N_BASIS[basis][species]
dim_triu = dim * (dim + 1) // 2
# -
# # NN Utils
# +
#keras.backend.clear_session()
#activation = "elu"
#learning_rate = 1e-5
intializer = keras.initializers.TruncatedNormal(mean=0.0, stddev=0.01)
def make_model(
structure,
input_dim,
output_dim,
activation="elu",
learning_rate=1e-3
):
model = keras.Sequential()
# input layer
model.add(keras.layers.Dense(
structure[0],
activation=activation,
input_dim=input_dim,
kernel_initializer=intializer
))
for layer in structure[1:]:
model.add(keras.layers.Dense(
layer,
activation=activation,
kernel_initializer=intializer,
#bias_initializer='zeros',
kernel_regularizer=keras.regularizers.l2(5e-3)
))
#output
model.add(keras.layers.Dense(output_dim))
model.compile(
optimizer=keras.optimizers.Adam(learning_rate),
loss='MSE',
metrics=['mse']
)
return model
# +
early_stopping = keras.callbacks.EarlyStopping(
monitor="val_mean_squared_error",
min_delta=1e-8,
patience=20,
verbose=1
)
reduce_lr = keras.callbacks.ReduceLROnPlateau(
monitor='val_mean_squared_error',
factor=0.1,
patience=3,
verbose=1,
mode='auto',
min_delta=1e-6,
cooldown=2,
min_lr=1e-10
)
epochs = 1000
def train_model(model, dataset, filepath=None, learning_rate=1e-4, log_dir=None):
if not log_dir is None:
tensorboard = keras.callbacks.TensorBoard(
log_dir=log_dir,
histogram_freq=0,
batch_size=32,
#update_freq='epoch'
)
if not filepath is None:
checkpoint = keras.callbacks.ModelCheckpoint(
filepath,
monitor='val_mean_squared_error',
verbose=1,
save_best_only=True,
save_weights_only=False,
mode='auto',
period=1
)
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
train, validation = [], []
while True:
keras.backend.set_value(model.optimizer.lr, learning_rate)
history = model.fit(
x = dataset.training[0],
y = dataset.training[1],
epochs=epochs,
shuffle=True,
validation_data=dataset.validation,
verbose=1,
callbacks=[
early_stopping,
reduce_lr,
checkpoint,
#tensorboard
]
)
#error.append(model.evaluate(
# dataset.testing[0],
# dataset.testing[1],
# verbose=1
#)[1])
return error
# -
# # Training
dataset.training[0].shape, dataset.training[1].shape
descriptor.number_of_descriptors, dim**2
structure = [100, 70]
keras.backend.clear_session()
# +
model = make_model(
structure=structure,
input_dim=descriptor.number_of_descriptors * 2,
output_dim=dim**2,
)
model.summary()
# -
#i+=1
train_model(
model,
dataset,
model_path + "model.h5",
learning_rate=1e-3,
#log_dir=None#"./logs/H/" + name + "_" + "x".join(list(map(str, structure))) + "_" + str(i)
)
| notebooks/ApplicationCarbohydrates/Traning_HOMO_CC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import csv
# +
#file name
fname="test.tsv"
#assay date & name
asname="21xxxx_dna-dye"
#the information about what you want to measure (dna or protein)
#absorbance max
mol_wl=260
#extinction coefficient
mol_coef=230364
#correction factor for tag
#if not needed, please enter '0'
mol_cf=0
#the information about what you want to measure together with other molecule (dna or protein)
#absorbance max
tag_wl=500
#extinction coefficient
tag_coef=90000
#correction factor for tag
#if not needed, please enter '0'
tag_cf=0.22
#pathlength of nanodrop
pl=0.1
# +
test=[]
with open(fname) as file:
tsv_file=csv.reader(file,delimiter='\t')
for line in tsv_file:
test.append(line)
num=int((test.count([])/2)+1)
if num==1:
line_num=len(test)
else:
line_num=test.index([])
data=[]
for i in range(num):
data.append(test[i*(line_num+2):line_num+i*(line_num+2)])
dfs=[]
for i in range(num):
df=pd.DataFrame(data[i])
df.columns=[data[i][2][0],data[i][0][0]]
df=df.drop(df.index[0:3])
df=df.astype({data[i][2][0]:int,
data[i][0][0]:np.float})
dfs.append(df)
if i>0:
dfs[i].drop(data[i][2][0],axis='columns',inplace=True)
result=pd.concat(dfs,axis=1)
result=result.reset_index(drop=True)
# +
fig, ax=plt.subplots(figsize=(5,5))
x=result.loc[:,['Wavelength (nm)']]
yval=result.drop(['Wavelength (nm)'],axis=1)
for i in yval.columns:
y=yval.loc[:,[i]]
ax.plot(x,y,label=i, alpha=0.5, linewidth=2)
#line
plt.axvline(mol_wl, color='black', linestyle='--', linewidth=2, alpha=0.3)
plt.axvline(tag_wl, color='black', linestyle='--', linewidth=2, alpha=0.3)
plt.ylim (-0.05,1)
plt.xlim (190,840)
plt.xlabel('Wavelength (nm)')
plt.ylabel('1 mm Absorbance')
plt.legend()
plt.grid(True)
plt.savefig(asname+'.png')
plt.show()
# +
mol_data=[]
for l in yval.columns:
mol_data.append(yval.loc[mol_wl-x.loc[0][0],[l]].values)
if tag_wl>0:
tag_data=[]
for j in yval.columns:
tag_data.append(yval.loc[tag_wl-x.loc[0][0],[j]].values)
def cal(a,b,c,d,e):
return (a-(b*c))/d/e*1000000
mol_conc=[]
if tag_wl>0:
for m in range(num):
mol_conc.append(cal(mol_data[m],tag_data[m],tag_cf,mol_coef,pl))
else:
for m in range(num):
mol_conc.append(cal(mol_data[m],0,tag_cf,mol_coef,pl))
if tag_wl>0:
tag_conc=[]
for k in range(num):
tag_conc.append(cal(tag_data[k],mol_data[k],mol_cf,tag_coef,pl))
def eff(a,b):
return (a/b*100)
if tag_wl>0:
label_eff=[]
for n in range(num):
label_eff.append(eff(tag_conc[n],mol_conc[n]))
final_mol=pd.DataFrame(mol_conc,index=yval.columns.values,columns=['Molecule conc.(µM)'])
if tag_wl>0:
final_tag=pd.DataFrame(tag_conc,index=yval.columns.values,columns=['tag conc.(µM)'])
final_eff=pd.DataFrame(label_eff,index=yval.columns.values,columns=['Labeling efficiency(%)'])
if tag_wl>0:
final=final_mol.merge(final_tag,left_index=True,
right_index=True).merge(final_eff,
left_index=True,right_index=True)
else:
final=final_mol
final.to_csv(asname+'_final.csv',encoding='utf-8')
final
| notebooks/concentration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
from tqdm import tqdm_notebook as tqdm
from time import sleep
from IPython import embed
# %reload_ext autoreload
# %autoreload 2
# -
with open("../data/mydata/train_list_full.txt", 'r') as f:
img_files = f.readlines()
len(img_files)
# +
import random
random.shuffle(img_files)
s = 150
train_list, test_list = img_files[:s], img_files[s:]
len(train_list), len(test_list)
# -
with open("../data/mydata/train_list.txt", 'w') as f:
for item in train_list:
f.write("%s" % item)
with open("../data/mydata/test_list.txt", 'w') as f:
for item in test_list:
f.write("%s" % item)
| ml/dish_detector/notebooks/split_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Registration transform example
# This example illustrates how to use a RegistrationTransform to temporally align the frames of an EOPatch using different algorithms
# ### Create cloudless timelapse
# Imports from `sentinelhub` and `eolearn` to set up workflow that creates a timelapse
# +
import datetime
import numpy as np
from sentinelhub import BBox, CRS, MimeType, CustomUrlParam, DataCollection
from eolearn.mask import AddCloudMaskTask, get_s2_pixel_cloud_detector
from eolearn.core import EOPatch, FeatureType, LinearWorkflow
from eolearn.features import SimpleFilterTask
from eolearn.io import S2L1CWCSInput
from eolearn.io.processing_api import SentinelHubInputTask
# -
# Set up BBox of ROI and time interval
# +
INSTANCE_ID = None
roi_bbox = BBox(bbox=[31.112895,29.957240,31.154222,29.987687], crs=CRS.WGS84)
# roi_bbox = BBox(bbox=[-6.57257, 37.2732, -5.728, 36.8549], crs=CRS.WGS84)
time_interval = ('2018-01-01', '2020-06-01')
# -
# This predicate function filters the images with a cloud coverage larger than a threshold to ensure images do not contain cloudy pixels
class MaxCCPredicate:
def __init__(self, maxcc):
self.maxcc = maxcc
def __call__(self, img_cm):
w, h, _ = img_cm.shape
cc = np.sum(img_cm) / (w * h)
return cc <= self.maxcc
# Tasks of the workflow:
# * download S2 images (only 3 bands needed for true color visualitzation)
# * Download cloud mask (CLM) provided by Sentinel Hub
# * filter out images with cloud coverage larger than a given threshold (e.g. 0.05)
download_task = SentinelHubInputTask(data_collection=DataCollection.SENTINEL2_L1C,
bands_feature=(FeatureType.DATA, 'BANDS'),
resolution=10,
maxcc=0.5,
bands=['B02', 'B03', 'B04'],
time_difference=datetime.timedelta(hours=2),
additional_data=[(FeatureType.MASK, 'dataMask', 'IS_DATA'), (FeatureType.MASK, 'CLM')]
)
filter_clouds = SimpleFilterTask((FeatureType.MASK, 'CLM'), MaxCCPredicate(maxcc=0.05))
# Build and execute timelapse as chain of transforms
# +
timelapse = LinearWorkflow(download_task, filter_clouds)
result = timelapse.execute({
download_task: {
'bbox': roi_bbox,
'time_interval': time_interval
}
})
# -
# Get result as an eopatch
eopatch_clean = [result[key] for key in result.keys()][0]
eopatch_clean
# #### Help function to create GIFs
# +
import imageio, os
def make_gif(eopatch, project_dir, filename, fps):
"""
Generates a GIF animation from an EOPatch.
"""
with imageio.get_writer(os.path.join(project_dir, filename), mode='I', fps=fps) as writer:
for image in eopatch:
writer.append_data(np.array(np.clip(2.8*image[..., [2, 1, 0]], 0, 255), dtype=np.uint8))
# -
# Write clean EOPatch to GIF
make_gif(eopatch=eopatch_clean.data['BANDS'] * 255, project_dir='.', filename='eopatch_clean.gif', fps=3)
# ## Run registrations
# Import registrations
from eolearn.coregistration import ECCRegistration, ThunderRegistration, PointBasedRegistration
# The constructor of the Registration objects takes the attribute type, field name and index of the channel to be used for registration, a dictionary specifying the parameters of the registration, and the interpolation method to be applied to the images. The interpolation methods are (NEAREST, LINEAR and CUBIC). Default is CUBIC. A nearest neighbour interpolation is used on ground-truth data to avoid creation of new labels.
# ### Thunder registration
# This algorithm computes translations only between pairs of images, using correlation on the Fourier transforms of the images
# +
coregister_thunder = ThunderRegistration((FeatureType.DATA, 'BANDS'), channel=2)
eopatch_thunder = coregister_thunder(eopatch_clean)
# -
# Write result to GIF
make_gif(eopatch=eopatch_thunder.data['BANDS']*255, project_dir='.', filename='eopatch_thunder.gif', fps=3)
# ### Enhanced Cross-Correlation in OpenCV
# This algorithm uses intensity values to maximise cross-correlation between pair of images. It uses an Euler transformation (x,y translation plus rotation).
# +
params = {'MaxIters': 200}
coregister_ecc = ECCRegistration((FeatureType.DATA, 'BANDS'), channel=2, params=params)
eopatch_ecc = coregister_ecc(eopatch_clean)
# -
make_gif(eopatch=eopatch_ecc.data['BANDS']*255, project_dir='.', filename='eopatch_ecc.gif', fps=3)
# ### Point-Based Registration in OpenCV
# Three transformation models are supported for point-based registration, i.e. Euler, PartialAffine and Homography. These methods compute feature descriptors (i.e. SIFT or SURF) of the pair of images to be registered, and estimate a robust transformation using RANSAC to align the matching points. These methods perform poorly compared to the other methods due to the inaccuracies of the feature extraction, point-matching and model fitting. If unplausible transformations are estimated, a warning is issued and an identity matrix is employed instead of the estimated transform. Default parameters are (Model=Euler, Descriptor=SIFT, RANSACThreshold=7.0, MaxIters=1000).
#
# Note: In case the following cell will raise an error
#
# ```Python
# AttributeError: module 'cv2.cv2' has no attribute 'xfeatures2d'
# ```
#
# uninstall and reinstall Python package `opencv-contrib-python`
# +
params = {
'Model': 'Euler',
'Descriptor': 'SURF',
'RANSACThreshold': 7.0,
'MaxIters': 1000
}
coregister_pbased = PointBasedRegistration((FeatureType.DATA, 'BANDS'), channel=2, params=params)
eopatch_pbased = coregister_pbased(eopatch_clean)
# -
make_gif(eopatch=eopatch_pbased.data['BANDS']*255, project_dir='.', filename='eopatch_pbased.gif', fps=3)
| examples/coregistration/Coregistration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
import scipy.signal as sig
import scipy.stats as stat
import matplotlib.pyplot as plt
import seaborn as sns
import os
import h5py
import pandas as pd
from pandas import DataFrame,Series,read_table
# +
## To run just one bird:
birdToRun = "Bird 4"
birdToRunID = int(birdToRun[5]) - 1
# -
# General info
# +
savePlots = True # whether or not to save plots
saveData = True # whether or not to save csv files
saveAsPath = './Fig 07/'
if not os.path.exists(saveAsPath):
os.mkdir(saveAsPath)
saveAsName = 'Fig7'
# +
#path = '/Users/svcanavan/Dropbox/Coding in progress/00_BudgieSleep/Data_copies/'
birdPaths = ['../data_copies/01_PreprocessedData/01_BudgieFemale_green1/00_Baseline_night/',
'../data_copies/01_PreprocessedData/02_BudgieMale_yellow1/00_Baseline_night/',
'../data_copies/01_PreprocessedData/03_BudgieFemale_white1/00_Baseline_night/',
'../data_copies/01_PreprocessedData/04_BudgieMale_yellow2/00_Baseline_night/',
'../data_copies/01_PreprocessedData/05_BudgieFemale_green2/00_Baseline_night/']
arfFilePaths = ['EEG 2 scored/',
'EEG 3 scored/',
'EEG 3 scored/',
'EEG 4 scored/',
'EEG 4 scored/']
### load BEST EEG channels - as determined during manual scoring ####
channelsToLoadEEG_best = [['5 LEEGf-LEEGp', '6 LEEGm-LEEGp'],
['5 LEEGf-LEEGm', '4 LEEGf-Fgr'],
['4LEEGf-LEEGp', '9REEGm-REEGp'],
['9REEGf-REEGp', '6LEEGm-LEEGf'],
['4LEEGf-LEEGp','7REEGf-REEGp']]
### load ALL of EEG channels ####
channelsToLoadEEG = [['1 LEOG-Fgr', '2 REOG-Fgr', '4 LEEGf-Fgr', '5 LEEGf-LEEGp', '6 LEEGm-LEEGp', '7 LEEGp-Fgr', '8 REEGp-Fgr','9 REEGp-LEEGp'],
['2 LEOG-Fgr', '3 REOG-Fgr', '4 LEEGf-Fgr','5 LEEGf-LEEGm', '6 LEEGm-LEEGp', '7 REEGf-Fgr', '8 REEGm-Fgr', '9 REEGf-REEGm'],
['2LEOG-Fgr', '3REOG-Fgr', '4LEEGf-LEEGp', '5LEEGf-LEEGm', '6LEEGm-LEEGp', '7REEGf-REEGp', '8REEGf-REEGm', '9REEGm-REEGp'],
['2LEOG-Fgr', '3REOG-Fgr', '4LEEGf-LEEGp', '5LEEGm-LEEGp', '6LEEGm-LEEGf', '7REEGf-Fgr', '8REEGf-REEGm','9REEGf-REEGp',],
['2LEOG-Fgr', '3REOG-Fgr', '4LEEGf-LEEGp', '5LEEGf-LEEGm', '6LEEGm-LEEGp', '7REEGf-REEGp', '8REEGf-REEGm', '9REEGm-REEGp']]
channelsToLoadEOG = [['1 LEOG-Fgr', '2 REOG-Fgr'],
['2 LEOG-Fgr', '3 REOG-Fgr'],
['2LEOG-Fgr', '3REOG-Fgr'],
['2LEOG-Fgr', '3REOG-Fgr'],
['2LEOG-Fgr', '3REOG-Fgr']]
birds_LL = [1,2,3]
nBirds_LL = len(birds_LL)
birdPaths_LL = ['../data_copies/01_PreprocessedData/02_BudgieMale_yellow1/01_Constant_light/',
'../data_copies/01_PreprocessedData/03_BudgieFemale_white1/01_Constant_light/',
'../data_copies/01_PreprocessedData/04_BudgieMale_yellow2/01_Constant_light/',]
arfFilePaths_LL = ['EEG 2 preprocessed/',
'EEG 2 preprocessed/',
'EEG 2 preprocessed/']
lightsOffSec = np.array([7947, 9675, 9861 + 8*3600, 9873, 13467]) # lights off times in seconds from beginning of file
lightsOnSec = np.array([46449, 48168, 48375+ 8*3600, 48381, 52005]) # Bird 3 gets 8 hours added b/c file starts at 8:00 instead of 16:00
epochLength = 3
sr = 200
scalingFactor = (2**15)*0.195 # scaling/conversion factor from amplitude to uV (when recording arf from jrecord)
stages = ['w','d','u','i','s','r'] # wake, drowsy, unihem sleep, intermediate sleep, SWS, REM
stagesSleep = ['u','i','s','r']
stagesVideo = ['m','q','d', 'l', 'r', 's','u'] # moving wake, quiet wake, drowsy, left uni, right uni, sleep, unclear
## Path to scores formatted as CSVs
formatted_scores_path = '../formatted_scores/'
## Path to detect SW ands EM events
events_path = '../data_copies/SWs_and_EMs/'
# -
# ##### TEST DATA - THIS NOTEBOOK ONLY ######
#
# birdPaths = ['','','/Users/Sofija/Dropbox/Coding in progress/00_BudgieSleep/data_copies/']
# arfFilePaths = ['','','EEG/']
#
# saveAsPath = '/Users/Sofija/Desktop/'
# +
colors = sns.color_palette(np.array([[234,103,99],
[218,142,60],
[174,174,62],
[97,188,101],
[140,133,232],
[225,113,190]])
/255)
sns.palplot(colors)
# colorpalette from iWantHue
# -
# Plot-specific info
# +
sns.set_context("notebook", font_scale=1)
sns.set_style("white")
# Markers for legends of EEG scoring colors
legendMarkersEEG = []
for stage in range(len(stages)):
legendMarkersEEG.append(plt.Line2D([0],[0], color=colors[stage], marker='o', linestyle='', alpha=0.7))
# -
# Calculate general variables
# +
lightsOffEp = lightsOffSec / epochLength
lightsOnEp = lightsOnSec / epochLength
nBirds = len(birdPaths)
epochLengthPts = epochLength*sr
nStages = len(stagesSleep)
birds = np.arange(1,6)
# -
# ## Read in files
# Read in EEG traces
# +
## To only load 1 bird: ##
b = birdToRunID
arf_path = birdPaths[b] + arfFilePaths[b]
EEGdataAll = {}
for channel in channelsToLoadEEG[b]:
all_data_array = np.array([])
for file in np.sort(os.listdir(arf_path)):
if file.endswith('.arf'):
arffile = h5py.File(arf_path+file, 'r')
data_array = arffile['.'][channel].value
data_array = np.ndarray.flatten(data_array)
# Pad the end with NaN's to make it divisible by epoch length
nanPadding = np.zeros(epochLengthPts - np.mod(len(data_array), epochLengthPts))
nanPadding.fill(np.nan)
data_array = np.append(data_array,nanPadding)
all_data_array = np.append(all_data_array,data_array)
# Do not reshape
# Save in dict under bird number and channel
data_name = 'Bird ' + str(b+1) + ': ' + channel
EEGdataAll[data_name] = scalingFactor * all_data_array
EEGchannels = np.sort(list(EEGdataAll.keys()))
# +
# Create time index for EEG
all_time_array = np.array([], dtype='datetime64')
for file in np.sort(os.listdir(arf_path)):
if file.endswith('.arf'):
arffile = h5py.File(arf_path+file, 'r')
date = file.split('_')[2]
if b == 0:
hours = '17'
minutes = '32'
else:
time = file.split('_')[3]
hours = time.split('-')[0]
minutes = time.split('-')[1]
datetime_start = np.datetime64(date + 'T' + hours + ':' + minutes + ':06') # assume 6-s delay in starting recording
# time index in datetime format
length_s = len(arffile['.'][channel].value)/sr
length_ms = np.timedelta64(int(1000 * length_s), 'ms')
datetime_end = datetime_start + length_ms
time_array = np.arange(datetime_start, datetime_end, np.timedelta64(int(1000/sr),'ms'))
# Add to end of whole-night time index
all_time_array = np.append(all_time_array, time_array)
TimeIndexEEG = {}
data_name = 'Bird ' + str(b+1)
TimeIndexEEG[data_name] = all_time_array
# -
# # Load scores from CSV
AllScores = {}
for b in birds:
# Load from file
scores_file = 'All_scores_Bird {}.csv'.format(b)
tmp_scores = pd.read_table(formatted_scores_path + scores_file, sep=',', index_col=0)
# save to dict
AllScores['Bird ' + str(b)] = tmp_scores
# # for each channel, mark as artifact epochs w/ data that crosses an amplitude threshold
# +
# Set thresholds
artifact_threshold_uV = 2000
artifact_threshold_SD = 8 # of SDs away from mean
# Make a scores array for each channel so it has independent artifact removal
ChannelScores = {}
for ch in EEGchannels:
data = EEGdataAll[ch]
artifact_threshold_SD_uV = np.abs(data[~np.isnan(data)]).mean() + artifact_threshold_SD*np.abs(data[~np.isnan(data)]).std()
print(ch + ' : ' + str(artifact_threshold_SD_uV))
b_name = ch[0:6]
bird_scores = AllScores[b_name]['Label (#)'].values # get scores as an array of numbers
nEpochs = len(bird_scores)
for ep in range(nEpochs):
start_pts = ep * epochLengthPts
stop_pts = (ep+1) * epochLengthPts
ep_data = data[start_pts:stop_pts]
if any(np.abs(ep_data) > artifact_threshold_SD_uV):
bird_scores[ep] = -2
# Save to dataframe
ChannelScores[ch] = bird_scores
# -
# ## Calculate lights off in Zeitgeber time (s and hrs)
# Lights on is 0
lightsOffZeit_s = lightsOffSec - lightsOnSec
lightsOffZeit_hr = lightsOffZeit_s / 3600
# ## Detect slow waves: define function
def detectSlowWaves(rawdata, data_scores, freqLowCut=0.5, freqHighCut=4, pos_or_neg_waves='neg', halfwaveAmpCrit_uV=75/2,
halfwaveArtifactCrit_uV=600/2, peak_within_halfwaveAmpCrit_uV=300/2, artifact_pts=np.array([])):
# if detecting positive waves, flip data before continuing with analysis
if 'pos' in pos_or_neg_waves:
rawdata = -rawdata
# Calculate wavelength criteria for half-waves
halfwaveMinLength = (1/freqHighCut)*sr/2
halfwaveMaxLength = (1/freqLowCut)*sr/2
# Calculate resolution & Nyquist frequency
res = 1/sr
fN = sr/2
# Filter data in delta range
filtOrder = 2
filtB, filtA = sig.butter(filtOrder, [freqLowCut/fN, freqHighCut/fN], 'bandpass', output='ba')
data = sig.filtfilt(filtB, filtA, rawdata)
# Find upwards and downwards zero-crossings
zeroCrossingsDown = np.where(np.diff(np.sign(data)) < 0)[0]
zeroCrossingsUp = np.where(np.diff(np.sign(data)) > 0)[0]
# To select for only negative half waves, remove first UZC if it is earlier than DZC
if zeroCrossingsUp[0] < zeroCrossingsDown[0]:
zeroCrossingsUp = zeroCrossingsUp[1::]
# If last DZC doesn't have a corresponding UZC following it, remove
if zeroCrossingsDown[-1] > zeroCrossingsUp[-1]:
zeroCrossingsDown = zeroCrossingsDown[0:-1]
# Get wavelengths, keep those in sleep epochs & within delta frequency
halfwaveLengths = zeroCrossingsUp - zeroCrossingsDown
# Too short
BadZC = np.where(halfwaveLengths < halfwaveMinLength)[0]
# Too long
BadZC = np.append(BadZC, np.where(halfwaveLengths > halfwaveMaxLength)[0])
# Remove bad DZC/UZC pairs
BadZC = np.sort(np.unique(BadZC))
DZC = np.delete(zeroCrossingsDown,BadZC)
UZC = np.delete(zeroCrossingsUp, BadZC)
# e. Loop through each wave, test for criteria
# Initialize
swCount = 0
swStartInds = []
swPeakInds = []
swStopInds = []
swPeakAmps = []
swNegSlopes = []
swPosSlopes = []
swLengths = []
swFreqs = []
for n in range(len(DZC)):
half_waveform = data[DZC[n]:UZC[n]]
# Not within sleep epochs
data_scores_starts_pts = (data_scores['Time (s)'] - start)*sr
data_scores_stops_pts = (data_scores['Time (s)'] + epochLength - start)*sr
epochs_spanned = data_scores.loc[(DZC[n]<=data_scores_stops_pts.values) & (UZC[n]>=data_scores_starts_pts.values)]
if any(epochs_spanned['Label (#)'] < 2):
isSlowWave = False
else:
# Test for overlap with pre defined list of artifact points
overlap_thres = .5 # threshold for proportion of wave that can overlap artifact
start_pts = DZC[n]
stop_pts = UZC[n]
length_pts = stop_pts - start_pts
wave_range_pts = np.arange(start_pts, stop_pts)
# repeat_counts: the number of times an index appears in both the slow wave and artifacts ranges
unique, repeat_counts = np.unique(np.append(wave_range_pts, artifact_pts), return_counts=True)
overlap_pts = sum(repeat_counts > 1) # count overlapping points
if overlap_pts >=overlap_thres*length_pts: # compare to length of slow wave
isSlowWave = False
else:
# Test amplitude peak: larger than amplitude criteria, smaller than artifact threshold
peakAmp = np.abs(np.min(half_waveform ))
if (peakAmp>=halfwaveAmpCrit_uV) & (peakAmp<halfwaveArtifactCrit_uV):
# Get the negative & positive slopes
minNegSlope = np.min(np.diff(half_waveform )/res)
maxPosSlope = np.max(np.diff(half_waveform )/res)
# Get amplitude of local upward peaks within the half wave
peaks_within_halfwave_ind = sig.argrelmax(half_waveform)
if len(peaks_within_halfwave_ind) > 0:
peaks_within_halfwave_amp = half_waveform[peaks_within_halfwave_ind]
if any(peaks_within_halfwave_amp >= -peak_within_halfwaveAmpCrit_uV):
isSlowWave = False
else:
isSlowWave = True
else:
isSlowWave = True
if isSlowWave:
# It's a slow wave - add characteristics to arrays
swCount = swCount + 1
swStartInds.append(DZC[n])
swPeakInds.append(DZC[n] + np.argmin(half_waveform))
swStopInds.append(UZC[n])
swPeakAmps.append(peakAmp)
swNegSlopes.append(minNegSlope)
swPosSlopes.append(maxPosSlope)
swLengths.append((UZC[n] - DZC[n])/sr)
swFreqs.append((1/((UZC[n] - DZC[n])/sr))/2)
swStart_s = start + np.array(swStartInds)/sr
swStop_s = start + np.array(swStopInds)/sr
# if had been detecting positive waves, flip the sign of data, amplitudes
if 'pos' in pos_or_neg_waves:
swPeakAmps = list(np.array(swPeakAmps))
data = -data
rawdata = -rawdata
else:
halfwaveAmpCrit_uV = -halfwaveAmpCrit_uV
halfwaveArtifactCrit_uV = -halfwaveArtifactCrit_uV
peak_within_halfwaveAmpCrit_uV = -peak_within_halfwaveAmpCrit_uV
waves_detected = {'data':data,'zeroCrossingsDown':zeroCrossingsDown,'zeroCrossingsUp':zeroCrossingsUp,'DZC':DZC,'UZC':UZC,
'swCount':swCount,'swStartInds':swStartInds,'swPeakInds':swPeakInds,'swStopInds':swStopInds,
'swPeakAmps':swPeakAmps,'swNegSlopes':swNegSlopes,'swPosSlopes':swPosSlopes, 'swLengths':swLengths,
'swFreqs':swFreqs,'swStart_s':swStart_s,'swStop_s':swStop_s}
return(waves_detected)
# ## Detect eye movements in EOG: define function
def detectEyeMovements(rawdata, data_scores, freqLowCut=0.2, freqHighCut=60, pos_or_neg_waves='neg', halfwaveAmpCrit_uV=5000,
halfwaveArtifactCrit_uV=500000, negSlopeThres=75):
# if detecting positive waves, flip data before continuing with analysis
if 'pos' in pos_or_neg_waves:
rawdata = -rawdata
# Calculate wavelength criteria for half-waves
halfwaveMinLength = (1/freqHighCut)*sr/2
halfwaveMaxLength = (1/freqLowCut)*sr/2
# Calculate resolution & Nyquist frequency
res = 1/sr
fN = sr/2
# No filtering
#filtOrder = 2
#filtB, filtA = sig.butter(filtOrder, [freqLowCut/fN, freqHighCut/fN], 'bandpass', output='ba')
#data = sig.filtfilt(filtB, filtA, rawdata)
data = rawdata
# Find upwards and downwards zero-crossings
zeroCrossingsDown = np.where(np.diff(np.sign(data)) < 0)[0]
zeroCrossingsUp = np.where(np.diff(np.sign(data)) > 0)[0]
# To select for only negative half waves, remove first UZC if it is earlier than DZC
if zeroCrossingsUp[0] < zeroCrossingsDown[0]:
zeroCrossingsUp = zeroCrossingsUp[1::]
# If last DZC doesn't have a corresponding UZC following it, remove
if zeroCrossingsDown[-1] > zeroCrossingsUp[-1]:
zeroCrossingsDown = zeroCrossingsDown[0:-1]
# Get wavelengths, keep those in sleep epochs & within delta frequency
halfwaveLengths = zeroCrossingsUp - zeroCrossingsDown
# Too short
BadZC = np.where(halfwaveLengths < halfwaveMinLength)[0]
# Too long
BadZC = np.append(BadZC, np.where(halfwaveLengths > halfwaveMaxLength)[0])
# Remove bad DZC/UZC pairs
BadZC = np.sort(np.unique(BadZC))
DZC = np.delete(zeroCrossingsDown,BadZC)
UZC = np.delete(zeroCrossingsUp, BadZC)
# e. Loop through each wave, test for criteria
# Initialize
swCount = 0
swStartInds = []
swPeakInds = []
swStopInds = []
swPeakAmps = []
swNegSlopes = []
swPosSlopes = []
swLengths = []
swFreqs = []
for n in range(len(DZC)):
half_waveform = data[DZC[n]:UZC[n]]
# Not within sleep epochs
data_scores_starts_pts = (data_scores['Time (s)'] - start)*sr
data_scores_stops_pts = (data_scores['Time (s)'] + epochLength - start)*sr
epochs_spanned = data_scores.loc[(DZC[n]<=data_scores_stops_pts.values) & (UZC[n]>=data_scores_starts_pts.values)]
if any(epochs_spanned['Label (#)'] < 2):
isSlowWave = False
else:
# Test amplitude peak: larger than amplitude criteria, smaller than artifact threshold
peakAmp = np.abs(np.min(half_waveform ))
if (peakAmp>=halfwaveAmpCrit_uV) & (peakAmp<halfwaveArtifactCrit_uV):
# Get the negative & positive slopes
minNegSlope = np.min(np.diff(half_waveform )/res)
maxPosSlope = np.max(np.diff(half_waveform )/res)
# Test if the fastest negative slope
if minNegSlope > -negSlopeThres:
isSlowWave = False
else:
isSlowWave = True
if isSlowWave:
# It's a slow wave - add characteristics to arrays
swCount = swCount + 1
swStartInds.append(DZC[n])
swPeakInds.append(DZC[n] + np.argmin(half_waveform))
swStopInds.append(UZC[n])
swPeakAmps.append(peakAmp)
swNegSlopes.append(minNegSlope)
swPosSlopes.append(maxPosSlope)
swLengths.append((UZC[n] - DZC[n])/sr)
swFreqs.append((1/((UZC[n] - DZC[n])/sr))/2)
swStart_s = start + np.array(swStartInds)/sr
swStop_s = start + np.array(swStopInds)/sr
# if had been detecting positive waves, flip the sign of data, amplitudes
if 'pos' in pos_or_neg_waves:
swPeakAmps = list(-np.array(swPeakAmps))
data = -data
rawdata = -rawdata
else:
halfwaveAmpCrit_uV = -halfwaveAmpCrit_uV
halfwaveArtifactCrit_uV = -halfwaveArtifactCrit_uV
waves_detected = {'data':data,'zeroCrossingsDown':zeroCrossingsDown,'zeroCrossingsUp':zeroCrossingsUp,'DZC':DZC,'UZC':UZC,
'swCount':swCount,'swStartInds':swStartInds,'swPeakInds':swPeakInds,'swStopInds':swStopInds,
'swPeakAmps':swPeakAmps,'swNegSlopes':swNegSlopes,'swPosSlopes':swPosSlopes, 'swLengths':swLengths,
'swFreqs':swFreqs,'swStart_s':swStart_s,'swStop_s':swStop_s}
return(waves_detected)
# ## Define function to plot manual scores
def PlotScoresBar(b, startTime, dT=20, scoreBarWidth=10, sr=sr, colors=colors, stages=stages):
'''Plot Figure 1: sleep score, EEG & EOG
b = bird name
startTime = where to start plotting, in seconds
dT = number of seconds to plot
ylimAmtEOG / EEG = set range of y axis above & below 0
yCalibBarEOG / EEG = how big to make the calibration bar for uV
xCalibBarEOG / EEG = how big to make the calibration bar for sec
sr = sampling rate
colors = list of colors to use for plotting sleep stages
stages = list of sleep/wake stages
EOGchannels = dictionary of EOG channels to use (for all birds)
EEGchannels = dictionary of EEG channels to use (for all birds)
'''
# Bird number from 0-4:
birdID = int(b[5])-1
# Get datetime index
time_index = TimeIndexEEG[b]
start_datetime_rec = time_index[0]
# calc start and stop datetimes
start_timedelta = np.timedelta64(startTime, 's')
dt_timedelta = np.timedelta64(dT, 's')
start_datetime = start_datetime_rec + start_timedelta
stop_datetime = start_datetime + dt_timedelta
dP = dT*sr
offset = 0/15 # fraction of plot size to leave blank on either side
###########################################################################
# Plotting parameters
width = scoreBarWidth
scoreLoc = 0
# Get scores to plot
scoresToPlot = AllScores[b][str(start_datetime).replace('T', ' '):str(stop_datetime).replace('T', ' ')]['Label'].values
start_sec = pd.to_datetime(start_datetime).second
firstEpOffset = (start_sec%epochLength)*sr # how much of first epoch is cut off at beginning, in pts
nEpochs = len(scoresToPlot)
# replace 'l' or 'g' in "Scores to Plot" with 'u' for unihem
unihem_inds = [x for x in range(nEpochs) if ('l' in scoresToPlot[x])|('g' in scoresToPlot[x])]
scoresToPlot[unihem_inds] = 'u'
# 1. Plot first epoch (which might be cut off at beginning):
# determine color based on sleep stage
scoreNum = [x for x in range(len(stages)) if stages[x] in scoresToPlot[0]][0]
scoreColor = colors[scoreNum]
# determine where to draw the bar
start = 0
stop = epochLengthPts - firstEpOffset
# draw the bar
plt.hlines(scoreLoc, start, stop, color=scoreColor, linewidth=width)
# 2. Plot middle epochs
for ep in np.arange(1,nEpochs-1):
# determine color based on sleep stage
scoreNum = [x for x in range(len(stages)) if stages[x] in scoresToPlot[ep]][0]
scoreColor = colors[scoreNum]
# determine where to draw the bar
start = ep*epochLengthPts - firstEpOffset
stop = ep*epochLengthPts - firstEpOffset + epochLengthPts
# draw the bar
plt.hlines(scoreLoc, start, stop, color=scoreColor, linewidth=width)
# 3. Plot last epoch (which might be cut off at end)
lastEp = nEpochs-1
# determine color based on sleep stage
scoreNum = [x for x in range(len(stages)) if stages[x] in scoresToPlot[lastEp]][0]
scoreColor = colors[scoreNum]
# determine where to draw the bar
start = lastEp*epochLengthPts - firstEpOffset
stop = dP
# draw the bar
plt.hlines(scoreLoc, start, stop, color=scoreColor, linewidth=width)
# Get rid of axes
plt.yticks([])
plt.xticks([])
sns.despine(left=True, bottom=True)
plt.xlim(plt.xlim(-dP*offset, dP + dP*offset))
# # Run on data
# ## Pick data
b = birdToRunID
bird_time = TimeIndexEEG[birdToRun]
# +
# Pick start and stop times
start = (11*3600) + (40*60)
end = start + (5*60)
dT = end - start
start_pts = start*sr
end_pts = end*sr
freqLowCut=0.5
freqHighCut=4
fN=sr/2
# -
# Get sleep scores
scores = AllScores[birdToRun]
data_scores = scores[(scores['Time (s)']>=start) & (scores['Time (s)']<=end)]
# ## Detect eye movements
# +
EOGchannels = [x for x in EEGchannels if 'EOG' in x]
data_LEOG = EEGdataAll[EOGchannels[0]][start_pts:end_pts]
data_REOG = EEGdataAll[EOGchannels[1]][start_pts:end_pts]
EOG_product = data_LEOG * data_REOG
rems_detected = detectEyeMovements(EOG_product, data_scores, pos_or_neg_waves='neg')
# -
# # Plot
chToPlot = EEGchannels[4]
# +
figsize = (9,10)
axis_label_fontsize = 24
# EEG plot formatting
linewidth = 1
linealpha = 1
color = [0,0,0]
eog_color = [.7, .7, .7]
product_color = [.3, .3, .3]
dzc_color = colors[4]
uzc_color = (0.4490196078431373, 0.42156862745098043, 0.80980392156862744)
em_color = colors[5]
artifact_color = colors[0]
# -
# ### FIGURE 07A: Plot eye movements
xlim_min = 3*60 + 21
xlim_max = xlim_min + 15
# +
nPlots = 6
plt.figure(figsize=(8,2*nPlots))
time = np.arange(0,dT,1/sr)
row=1
plt.subplot(nPlots, 1, row)
PlotScoresBar(birdToRun, start+xlim_min, dT=xlim_max-xlim_min)
row += 1
for ch in EOGchannels:
plt.subplot(nPlots, 1, row)
rawdata = EEGdataAll[ch][start_pts:end_pts]
plt.plot(time,rawdata, lw=linewidth, alpha=linealpha, color=eog_color)
#plt.ylabel(ch[9::])
plt.xlim(xlim_min,xlim_max)
plt.ylim(-200,200)
plt.xticks([])
if row==2:
ax1=plt.gca()
sns.despine(bottom=True, ax=ax1)
else:
ax2=plt.gca()
sns.despine(bottom=False, ax=ax2)
plt.yticks([])
row+=1
plt.subplot(nPlots, 1, row)
plt.plot(time, EOG_product, lw=linewidth, alpha=linealpha, color=product_color)
ax3=plt.gca()
sns.despine(ax=ax3)
plt.xlim(xlim_min,xlim_max)
plt.ylim(-60000,1000)
plt.yticks([])
plt.xticks([])
#plt.ylabel('EOG product')
row+=1
plt.subplot(nPlots, 1, row)
plt.plot(time, EOG_product, lw=linewidth, alpha=linealpha, color=product_color)
plt.axhline(-5000, color=em_color, alpha=linealpha/2)
# scatterplot of peaks: if peaks exceed y axis, plot dot on edge of plot
peaks = -1*np.array(rems_detected['swPeakAmps'])
peaks[peaks<=-60000] = -59900
plt.scatter(np.array(rems_detected['swPeakInds'])/sr, peaks, color=em_color)
ax4=plt.gca()
sns.despine(ax=ax4)
plt.xlim(xlim_min,xlim_max)
plt.ylim(-60000,1000)
plt.yticks([])
plt.xticks([])
#plt.ylabel('EOG product')
row+=1
plt.subplot(nPlots, 1, row)
plt.hlines(y=np.repeat(0.5, rems_detected['swCount']), xmin=rems_detected['swStart_s']-start, xmax=rems_detected['swStop_s']-start, linewidth=1000,alpha=.2, color=em_color)
offset = 0
for ch in EOGchannels:
rawdata = EEGdataAll[ch][start_pts:end_pts]
plt.plot(time, rawdata + offset*400, lw=linewidth, alpha=linealpha, color=eog_color)
offset-=1
ax5=plt.gca()
sns.despine(ax=ax5)
plt.xlim(xlim_min,xlim_max)
plt.ylim(10+offset*300, 200)
plt.yticks([])
#plt.ylabel('eye movements')
if savePlots:
#plt.savefig(saveAsPath + saveAsName + birdToRun + str(start) + 's_em_detection.pdf')
plt.savefig(saveAsPath + saveAsName + 'a_' + birdToRun + str(start) + 's_em_detection.tiff', dpi=300)
# -
# ## Detect eye movement artifacts
# +
Artifacts_EM_pos = {}
Artifacts_EM_neg = {}
ampCrit_artifacts = 10000
for ch in EEGchannels:
if 'EEG' in ch:
if 'LEEG' in ch:
eog_ch = EOGchannels[0]
elif 'REEG' in ch:
eog_ch = EOGchannels[1]
data_EEG = EEGdataAll[ch][start*sr:end*sr]
data_EOG = EEGdataAll[eog_ch][start*sr:end*sr]
EEG_EOG_product = data_EEG * data_EOG # Take product of EEG channel x ipsilateral EOG
artifacts_detected = detectEyeMovements(EEG_EOG_product, data_scores, pos_or_neg_waves='pos',halfwaveAmpCrit_uV=ampCrit_artifacts,
halfwaveArtifactCrit_uV=ampCrit_artifacts*10000, negSlopeThres=10)
Artifacts_EM_pos[ch] = artifacts_detected
artifacts_detected = detectEyeMovements(EEG_EOG_product, data_scores, pos_or_neg_waves='neg',halfwaveAmpCrit_uV=ampCrit_artifacts,
halfwaveArtifactCrit_uV=ampCrit_artifacts*10000, negSlopeThres=10)
Artifacts_EM_neg[ch] = artifacts_detected
# -
# ### Define list of points encompassed by artifacts
# +
All_Artifacts_points = {}
for ch in EEGchannels:
if 'EEG' in ch:
artifact_points_ch = np.array([]) # init
# Pos artifacts
artifacts = Artifacts_EM_pos[ch]
for artifact in range(artifacts['swCount']):
start_pts = artifacts['swStartInds'][artifact]
stop_pts = artifacts['swStopInds'][artifact]
range_pts = np.arange(start_pts, stop_pts)
artifact_points_ch = np.append(artifact_points_ch, range_pts)
# Neg artifacts
artifacts = Artifacts_EM_neg[ch]
for artifact in range(artifacts['swCount']):
start_pts = artifacts['swStartInds'][artifact]
stop_pts = artifacts['swStopInds'][artifact]
range_pts = np.arange(start_pts, stop_pts)
artifact_points_ch = np.append(artifact_points_ch, range_pts)
All_Artifacts_points[ch] = artifact_points_ch
# -
# ## Detect slow waves, excluding eye movement artifacts
# +
All_Waves_neg = {}
All_Waves_pos = {}
for ch in EEGchannels:
if 'EEG' in ch:
rawdata = EEGdataAll[ch][start*sr:end*sr]
waves_detected_neg = detectSlowWaves(rawdata,
data_scores, artifact_pts=All_Artifacts_points[ch],
pos_or_neg_waves='neg')
waves_detected_pos = detectSlowWaves(rawdata,
data_scores, artifact_pts=All_Artifacts_points[ch],
pos_or_neg_waves='pos')
All_Waves_neg[ch] = waves_detected_neg
All_Waves_pos[ch] = waves_detected_pos
# -
birdToRun
# ### FIGURE 07D: Plot slow wave detection
# #### used Bird 3 for this particular figure in the paper?
chToPlot=EEGchannels[4]
xlim_min = 60*2 + 30
xlim_max = xlim_min+15
# +
nPlots = 7
plt.figure(figsize=(8,2*nPlots))
time = np.arange(0,dT,1/sr)
# Scores bar
row=1
plt.subplot(nPlots, 1, row)
PlotScoresBar(birdToRun, start+xlim_min, dT=xlim_max-xlim_min)
row += 1
# Pick channel
ch = chToPlot
neg_waves = All_Waves_neg[ch]
pos_waves = All_Waves_pos[ch]
# raw data
plt.subplot(nPlots, 1, row)
rawdata = EEGdataAll[ch][start*sr:end*sr]
plt.plot(time,rawdata, lw=linewidth, alpha=linealpha, color=color)
plt.ylabel(ch[9::])
plt.xlim(xlim_min,xlim_max)
plt.ylim(-150,150)
plt.yticks([])
plt.xticks([])
row+=1
# filtered data
plt.subplot(nPlots, 1, row)
filtOrder = 2
filtB, filtA = sig.butter(filtOrder, [freqLowCut/fN, freqHighCut/fN], 'bandpass', output='ba')
filt_data = sig.filtfilt(filtB, filtA, rawdata)
plt.plot(time, filt_data, lw=linewidth, alpha=linealpha, color=color)
plt.ylabel('filtered')
plt.xlim(xlim_min,xlim_max)
plt.ylim(-150,150)
plt.yticks([])
plt.xticks([])
row+=1
# filtered, initial UZC and DZC
plt.subplot(nPlots, 1, row)
zeroCrossingsDown = np.where(np.diff(np.sign(filt_data)) < 0)[0]
zeroCrossingsUp = np.where(np.diff(np.sign(filt_data)) > 0)[0]
plt.plot(time, filt_data, lw=linewidth, alpha=linealpha, color=color)
plt.scatter(zeroCrossingsDown/sr, np.zeros_like(zeroCrossingsDown), color=dzc_color)
plt.scatter(zeroCrossingsUp/sr, np.zeros_like(zeroCrossingsUp), color=uzc_color)
# thresholds
plt.axhline(-75/2, alpha=linealpha/2, color=dzc_color)
plt.axhline(75/2, alpha=linealpha/2, color=uzc_color)
plt.ylabel('zero crossings')
plt.xlim(xlim_min,xlim_max)
plt.ylim(-150,150)
plt.yticks([])
plt.xticks([])
row+=1
# final amp peaks
plt.subplot(nPlots, 1, row)
plt.plot(time, filt_data, lw=linewidth, alpha=linealpha, color=color)
plt.scatter(np.array(neg_waves['swPeakInds'])/sr, -1*np.array(neg_waves['swPeakAmps']), color=dzc_color)
plt.scatter(np.array(pos_waves['swPeakInds'])/sr, np.array(pos_waves['swPeakAmps']), color=uzc_color)
# thresholds
plt.axhline(-75/2, alpha=linealpha/2, color=dzc_color)
plt.axhline(75/2, alpha=linealpha/2, color=uzc_color)
plt.ylabel('slow wave peaks')
plt.xlim(xlim_min,xlim_max)
plt.ylim(-150,150)
plt.yticks([])
plt.xticks([])
row+=1
# slow waves detected
plt.subplot(nPlots, 1, row)
plt.hlines(y=np.repeat(0.5, neg_waves['swCount']), xmin=neg_waves['swStart_s']-start, xmax=neg_waves['swStop_s']-start,linewidth=140,alpha=.2, color=dzc_color)
plt.hlines(y=np.repeat(0.5, pos_waves['swCount']), xmin=pos_waves['swStart_s']-start, xmax=pos_waves['swStop_s']-start,linewidth=140,alpha=.2, color=uzc_color)
plt.plot(time,rawdata, lw=linewidth, alpha=linealpha, color=color)
plt.ylabel('slow waves')
plt.xlim(xlim_min,xlim_max)
plt.ylim(-150,150)
row+=1
#if savePlots:
#plt.savefig(saveAsPath + saveAsName + birdToRun + '_' + ch[0:9] + str(start) + 's_sw_detection.pdf')
#plt.savefig(saveAsPath + saveAsName + 'd_' + birdToRun + '_' + ch[0:9] + str(start) + 's_sw_detection.tiff', dpi=300)
# -
# ### Plot eye movement artifacts
# +
nPlots = 6
plt.figure(figsize=(8,2*nPlots))
time = np.arange(0,dT,1/sr)
# Pick channel
ch = chToPlot
neg_waves = All_Waves_neg[ch]
pos_waves = All_Waves_pos[ch]
row=1
plt.subplot(nPlots, 1, row)
PlotScoresBar(birdToRun, start+xlim_min, dT=xlim_max-xlim_min)
row += 1
# plot EOGs
plt.subplot(nPlots, 1, row)
plt.hlines(y=np.repeat(0.5, rems_detected['swCount']), xmin=rems_detected['swStart_s']-start, xmax=rems_detected['swStop_s']-start, linewidth=130,alpha=.2, color=em_color)
offset = 0
if 'LEEG' in ch:
eog_ch = EOGchannels[0]
elif 'REEG' in ch:
eog_ch = EOGchannels[1]
rawdata = EEGdataAll[eog_ch][start*sr:end*sr]
plt.plot(time, rawdata, lw=linewidth, alpha=linealpha, color=eog_color)
plt.xlim(xlim_min,xlim_max)
plt.ylim(-150,150)
plt.ylabel('eye movements')
plt.yticks([]);
row+=1
# plot EEG channel
plt.subplot(nPlots, 1, row)
rawdata = EEGdataAll[ch][start*sr:end*sr]
plt.hlines(y=np.repeat(0.5, neg_waves['swCount']), xmin=neg_waves['swStart_s']-start, xmax=neg_waves['swStop_s']-start,linewidth=140,alpha=.2, color=dzc_color)
plt.hlines(y=np.repeat(0.5, pos_waves['swCount']), xmin=pos_waves['swStart_s']-start, xmax=pos_waves['swStop_s']-start,linewidth=140,alpha=.2, color=uzc_color)
plt.plot(time,rawdata, lw=linewidth, alpha=linealpha, color=color)
plt.ylabel('slow waves')
plt.yticks([])
plt.xticks([])
plt.xlim(xlim_min,xlim_max)
plt.ylim(-150,150)
row+=1
plt.subplot(nPlots, 1, row)
plt.plot(time, filt_data, lw=linewidth, alpha=linealpha, color=color)
plt.scatter(np.array(neg_waves['swPeakInds'])/sr, -1*np.array(neg_waves['swPeakAmps']), color=dzc_color)
plt.scatter(np.array(pos_waves['swPeakInds'])/sr, np.array(pos_waves['swPeakAmps']), color=uzc_color)
# thresholds
plt.axhline(-75/2, alpha=linealpha/2, color=dzc_color)
plt.axhline(75/2, alpha=linealpha/2, color=uzc_color)
plt.ylabel('slow wave peaks')
plt.xlim(xlim_min,xlim_max)
plt.ylim(-150,150)
row+=1
# Ipsilateral EOG
plt.subplot(nPlots, 1, row)
if 'LEEG' in ch:
eog_ch = EOGchannels[0]
elif 'REEG' in ch:
eog_ch = EOGchannels[1]
data_EOG = EEGdataAll[eog_ch][start*sr:end*sr]
# Product of EEG x ipsEOG
plt.subplot(nPlots, 1, row)
plt.plot(time, rawdata*data_EOG, lw=linewidth, alpha=linealpha, color=product_color)
# thresholds
plt.axhline(-ampCrit_artifacts, color=artifact_color, alpha=linealpha/2)
plt.axhline(ampCrit_artifacts, color=artifact_color, alpha=linealpha/2)
# artifact peaks
neg_artifacts = Artifacts_EM_neg[ch]
pos_artifacts = Artifacts_EM_pos[ch]
plt.scatter(np.array(neg_artifacts['swPeakInds'])/sr, -1*np.array(neg_artifacts['swPeakAmps']), color=artifact_color)
plt.scatter(np.array(pos_artifacts['swPeakInds'])/sr, -1*np.array(pos_artifacts['swPeakAmps']), color=artifact_color)
plt.xlim(xlim_min,xlim_max)
plt.ylim(-20000,20000)
plt.yticks([])
plt.ylabel('EEG-EOG product')
row+=1
# EEG with slow waves & artifacts
plt.subplot(nPlots, 1, row)
rawdata = EEGdataAll[ch][start*sr:end*sr]
plt.hlines(y=np.repeat(0.5, neg_waves['swCount']), xmin=neg_waves['swStart_s']-start, xmax=neg_waves['swStop_s']-start,linewidth=140,alpha=.2, color=dzc_color)
plt.hlines(y=np.repeat(0.5, pos_waves['swCount']), xmin=pos_waves['swStart_s']-start, xmax=pos_waves['swStop_s']-start,linewidth=140,alpha=.2, color=uzc_color)
plt.hlines(y=np.repeat(0.5, neg_artifacts['swCount']), xmin=neg_artifacts['swStart_s']-start, xmax=neg_artifacts['swStop_s']-start,linewidth=140,alpha=.5, color=artifact_color)
plt.hlines(y=np.repeat(0.5, pos_artifacts['swCount']), xmin=pos_artifacts['swStart_s']-start, xmax=pos_artifacts['swStop_s']-start,linewidth=140,alpha=.5, color=artifact_color)
plt.plot(time,rawdata, lw=linewidth, alpha=linealpha, color=color)
plt.ylabel('artifacts vs SWs')
plt.yticks([])
plt.xticks([])
plt.xlim(xlim_min,xlim_max)
plt.ylim(-150,150)
row+=1
#if savePlots:
#plt.savefig(saveAsPath + saveAsName + birdToRun + str(start) + 's_artifact_detection.pdf')
#plt.savefig(saveAsPath + saveAsName + birdToRun + str(start) + 's_artifact_detection.tiff', dpi=300)
# -
# ## Plot slow waves in all channels
# +
nPlots = len(EEGchannels)
plt.figure(figsize=(16,1*nPlots+1))
time = np.arange(0,dT,1/sr)
row=1
plt.subplot(nPlots, 1, row)
PlotScoresBar(birdToRun,start,dT)
row+=1
# plot EOGs
plt.subplot(nPlots, 1, row)
plt.hlines(y=np.repeat(0.5, rems_detected['swCount']), xmin=rems_detected['swStart_s']-start, xmax=rems_detected['swStop_s']-start, linewidth=130,alpha=.2, color=em_color)
offset = 0
for eog_ch in EOGchannels:
rawdata = EEGdataAll[eog_ch][start*sr:end*sr]
plt.plot(time, rawdata + offset*300, lw=linewidth, alpha=linealpha, color=eog_color)
offset+=1
plt.xlim(0,dT)
#plt.ylim(-150,150)
plt.ylabel('eye movements')
plt.yticks([]);
row+=1
for ch in EEGchannels:
if 'EEG' in ch:
plt.subplot(nPlots, 1, row)
rawdata = EEGdataAll[ch][start*sr:end*sr]
plt.plot(time,rawdata, lw=linewidth, alpha=linealpha, color=color)
neg_waves = All_Waves_neg[ch]
plt.hlines(y=np.repeat(-100, neg_waves['swCount']), xmin=neg_waves['swStart_s']-start, xmax=neg_waves['swStop_s']-start,linewidth=25,color=dzc_color,alpha=.8)
pos_waves = All_Waves_pos[ch]
plt.hlines(y=np.repeat(100, pos_waves['swCount']), xmin=pos_waves['swStart_s']-start, xmax=pos_waves['swStop_s']-start,linewidth=25,color=uzc_color,alpha=.8)
plt.ylabel(ch[9::],rotation=0)
plt.yticks([])
plt.xticks([])
plt.ylim(-150,150)
plt.xlim(0, dT)
row+=1
#if savePlots:
#plt.savefig(saveAsPath + saveAsName + birdToRun + str(start) + 's_detected_waves.pdf')
#plt.savefig(saveAsPath + saveAsName + birdToRun + str(start) + 's_detected_waves.tiff')
# -
| Fig07d - Eye movement detecting demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="3YLyiTwPfVCT"
# <a href="https://colab.research.google.com/github/magenta/ddsp/blob/master/ddsp/colab/demos/timbre_transfer.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
#
# ##### Copyright 2020 Google LLC.
#
# Licensed under the Apache License, Version 2.0 (the "License");
#
#
#
#
# + colab={} colab_type="code" id="Bvp6GWqtfVCW"
# Copyright 2020 Google LLC. All Rights Reserved.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
# + [markdown] colab_type="text" id="JndnmDMp66FL"
# # DDSP Timbre Transfer Demo
#
# This notebook is a demo of timbre transfer using DDSP (Differentiable Digital Signal Processing).
# The model here is trained to generate audio conditioned on a time series of fundamental frequency and loudness.
#
# * [DDSP ICLR paper](https://openreview.net/forum?id=B1x1ma4tDr)
# * [Audio Examples](http://goo.gl/magenta/ddsp/blob/master/ddsp/colab-examples)
#
# By default, the notebook will download pre-trained models for Violin and Flute. You can train a model on your own sounds by using the [Train Autoencoder Colab](https://github.com/magenta/ddsp/blob/master/ddsp/colab/demos/train_autoencoder.ipynb).
#
# <img src="https://storage.googleapis.com/ddsp/additive_diagram/ddsp_autoencoder.png" alt="DDSP Autoencoder figure" width="700">
#
#
# # Environment Setup
#
#
# This notebook extracts these features from input audio (either uploaded files, or recorded from the microphone) and resynthesizes with the model.
#
# Have fun! And please feel free to hack this notebook to make your own creative interactions.
#
# ### Instructions for running:
#
# * Make sure to use a GPU runtime, click: __Runtime >> Change Runtime Type >> GPU__
# * Press the ▶️button on the left of each of the cells
# * View the code: Double-click any of the cells
# * Hide the code: Double click the right side of the cell
#
# + cellView="form" colab={} colab_type="code" id="6wZde6CBya9k"
#@title #Install and Import
#@markdown Install ddsp, define some helper functions, and download the model. This transfers a lot of data and _should take a minute or two_.
# %tensorflow_version 2.x
print('Installing from pip package...')
# !pip install -qU ddsp
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
# Ignore a bunch of deprecation warnings
import warnings
warnings.filterwarnings("ignore")
import copy
import os
import time
import crepe
import ddsp
import ddsp.training
from ddsp.colab.colab_utils import (download, play, record, specplot, upload,
DEFAULT_SAMPLE_RATE)
import gin
from google.colab import files
import librosa
import matplotlib.pyplot as plt
import numpy as np
import tensorflow.compat.v2 as tf
import tensorflow_datasets as tfds
# Helper Functions
sample_rate = DEFAULT_SAMPLE_RATE # 16000
print('Done!')
# + cellView="form" colab={} colab_type="code" id="Go36QW9AS_CD"
#@title Record or Upload Audio
#@markdown * Either record audio from microphone or upload audio from file (.mp3 or .wav)
#@markdown * Audio should be monophonic (single instrument / voice)
#@markdown * Extracts fundmanetal frequency (f0) and loudness features.
record_or_upload = "Record" #@param ["Record", "Upload (.mp3 or .wav)"]
record_seconds = 5 #@param {type:"number", min:1, max:10, step:1}
if record_or_upload == "Record":
audio = record(seconds=record_seconds)[np.newaxis, :].astype(np.float32)
else:
# Load audio sample here (.mp3 or .wav3 file)
# Just use the first file.
filenames, audios = upload()
audio = audios[0:1].astype(np.float32)
print('\nExtracting audio features...')
# Plot.
specplot(audio)
play(audio)
# Setup the session.
ddsp.spectral_ops.reset_crepe()
# Compute features.
start_time = time.time()
audio_features = ddsp.training.eval_util.compute_audio_features(audio)
audio_features['loudness_db'] = audio_features['loudness_db'].astype(np.float32)
audio_features_mod = None
print('Audio features took %.1f seconds' % (time.time() - start_time))
# Plot Features.
fig, ax = plt.subplots(nrows=3,
ncols=1,
sharex=True,
figsize=(6, 8))
ax[0].plot(audio_features['loudness_db'])
ax[0].set_ylabel('loudness_db')
ax[1].plot(librosa.hz_to_midi(audio_features['f0_hz']))
ax[1].set_ylabel('f0 [midi]')
ax[2].plot(audio_features['f0_confidence'])
ax[2].set_ylabel('f0 confidence')
_ = ax[2].set_xlabel('Time step [frame]')
# + cellView="form" colab={} colab_type="code" id="wmSGDWM5yyjm"
#@title Choose a model
model = 'Violin' #@param ['Violin', 'Flute', 'Flute2', 'Upload your own (checkpoint folder as .zip)']
MODEL = model
GCS_CKPT_DIR = 'gs://ddsp/models/tf2'
def find_model_dir(dir_name):
# Iterate through directories until model directory is found
for root, dirs, filenames in os.walk(dir_name):
for filename in filenames:
if filename.endswith(".gin") and not filename.startswith("."):
model_dir = root
break
return model_dir
if model in ('Violin', 'Flute', 'Flute2'):
# Pretrained models.
PRETRAINED_DIR = '/content/pretrained'
# Copy over from gs:// for faster loading.
# !rm -r $PRETRAINED_DIR &> /dev/null
# !mkdir $PRETRAINED_DIR &> /dev/null
model_dir = os.path.join(GCS_CKPT_DIR, 'solo_%s_ckpt' % model.lower())
# !gsutil cp $model_dir/* $PRETRAINED_DIR &> /dev/null
model_dir = PRETRAINED_DIR
gin_file = os.path.join(model_dir, 'operative_config-0.gin')
else:
# User models.
UPLOAD_DIR = '/content/uploaded'
# !mkdir $UPLOAD_DIR
uploaded_files = files.upload()
for fnames in uploaded_files.keys():
print("Unzipping... {}".format(fnames))
# !unzip -o "/content/$fnames" -d $UPLOAD_DIR &> /dev/null
model_dir = find_model_dir(UPLOAD_DIR)
gin_file = os.path.join(model_dir, 'operative_config-0.gin')
# Parse gin config,
with gin.unlock_config():
gin.parse_config_file(gin_file, skip_unknown=True)
# Assumes only one checkpoint in the folder, 'ckpt-[iter]`.
ckpt_files = [f for f in tf.io.gfile.listdir(model_dir) if 'ckpt' in f]
ckpt_name = ckpt_files[0].split('.')[0]
ckpt = os.path.join(model_dir, ckpt_name)
# Ensure dimensions and sampling rates are equal
time_steps_train = gin.query_parameter('DefaultPreprocessor.time_steps')
n_samples_train = gin.query_parameter('Additive.n_samples')
hop_size = int(n_samples_train / time_steps_train)
time_steps = int(audio.shape[1] / hop_size)
n_samples = time_steps * hop_size
# print("===Trained model===")
# print("Time Steps", time_steps_train)
# print("Samples", n_samples_train)
# print("Hop Size", hop_size)
# print("\n===Resynthesis===")
# print("Time Steps", time_steps)
# print("Samples", n_samples)
# print('')
gin_params = [
'Additive.n_samples = {}'.format(n_samples),
'FilteredNoise.n_samples = {}'.format(n_samples),
'DefaultPreprocessor.time_steps = {}'.format(time_steps),
]
with gin.unlock_config():
gin.parse_config(gin_params)
# Trim all input vectors to correct lengths
for key in ['f0_hz', 'f0_confidence', 'loudness_db']:
audio_features[key] = audio_features[key][:time_steps]
audio_features['audio'] = audio_features['audio'][:, :n_samples]
# Set up the model just to predict audio given new conditioning
model = ddsp.training.models.Autoencoder()
model.restore(ckpt)
# Build model by running a batch through it.
start_time = time.time()
_ = model(audio_features, training=False)
print('Restoring model took %.1f seconds' % (time.time() - start_time))
# + cellView="form" colab={} colab_type="code" id="uQFUlIJ_5r36"
#@title Modify conditioning
#@markdown These models were not explicitly trained to perform timbre transfer, so they may sound unnatural if the incoming loudness and frequencies are very different then the training data (which will always be somewhat true).
#@markdown This button will at least adjusts the average loudness and pitch to be similar to the training data (although not for user trained models).
auto_adjust = True #@param{type:"boolean"}
#@markdown You can also make additional manual adjustments:
#@markdown * Shift the fundmental frequency to a more natural register.
#@markdown * Silence audio below a threshold on f0_confidence.
#@markdown * Adjsut the overall loudness level.
f0_octave_shift = 0 #@param {type:"slider", min:-2, max:2, step:1}
f0_confidence_threshold = 0 #@param {type:"slider", min:0.0, max:1.0, step:0.05}
loudness_db_shift = 0 #@param {type:"slider", min:-20, max:20, step:1}
#@markdown You might get more realistic sounds by shifting a few dB, or try going extreme and see what weird sounds you can make...
audio_features_mod = {k: v.copy() for k, v in audio_features.items()}
## Helper functions.
def shift_ld(audio_features, ld_shift=0.0):
"""Shift loudness by a number of ocatves."""
audio_features['loudness_db'] += ld_shift
return audio_features
def shift_f0(audio_features, f0_octave_shift=0.0):
"""Shift f0 by a number of ocatves."""
audio_features['f0_hz'] *= 2.0 ** (f0_octave_shift)
audio_features['f0_hz'] = np.clip(audio_features['f0_hz'],
0.0,
librosa.midi_to_hz(110.0))
return audio_features
def mask_by_confidence(audio_features, confidence_level=0.1):
"""For the violin model, the masking causes fast dips in loudness.
This quick transient is interpreted by the model as the "plunk" sound.
"""
mask_idx = audio_features['f0_confidence'] < confidence_level
audio_features['f0_hz'][mask_idx] = 0.0
# audio_features['loudness_db'][mask_idx] = -ddsp.spectral_ops.LD_RANGE
return audio_features
def smooth_loudness(audio_features, filter_size=3):
"""Smooth loudness with a box filter."""
smoothing_filter = np.ones([filter_size]) / float(filter_size)
audio_features['loudness_db'] = np.convolve(audio_features['loudness_db'],
smoothing_filter,
mode='same')
return audio_features
if auto_adjust:
if MODEL in ['Violin', 'Flute', 'Flute2']:
# Adjust the peak loudness.
l = audio_features['loudness_db']
model_ld_avg_max = {
'Violin': -34.0,
'Flute': -45.0,
'Flute2': -44.0,
}[MODEL]
ld_max = np.max(audio_features['loudness_db'])
ld_diff_max = model_ld_avg_max - ld_max
audio_features_mod = shift_ld(audio_features_mod, ld_diff_max)
# Further adjust the average loudness above a threshold.
l = audio_features_mod['loudness_db']
model_ld_mean = {
'Violin': -44.0,
'Flute': -51.0,
'Flute2': -53.0,
}[MODEL]
ld_thresh = -50.0
ld_mean = np.mean(l[l > ld_thresh])
ld_diff_mean = model_ld_mean - ld_mean
audio_features_mod = shift_ld(audio_features_mod, ld_diff_mean)
# Shift the pitch register.
model_p_mean = {
'Violin': 73.0,
'Flute': 81.0,
'Flute2': 74.0,
}[MODEL]
p = librosa.hz_to_midi(audio_features['f0_hz'])
p[p == -np.inf] = 0.0
p_mean = p[l > ld_thresh].mean()
p_diff = model_p_mean - p_mean
p_diff_octave = p_diff / 12.0
round_fn = np.floor if p_diff_octave > 1.5 else np.ceil
p_diff_octave = round_fn(p_diff_octave)
audio_features_mod = shift_f0(audio_features_mod, p_diff_octave)
else:
print('\nUser uploaded model: disabling auto-adjust.')
audio_features_mod = shift_ld(audio_features_mod, loudness_db_shift)
audio_features_mod = shift_f0(audio_features_mod, f0_octave_shift)
audio_features_mod = mask_by_confidence(audio_features_mod, f0_confidence_threshold)
# Plot Features.
fig, ax = plt.subplots(nrows=3,
ncols=1,
sharex=True,
figsize=(6, 8))
ax[0].plot(audio_features['loudness_db'])
ax[0].plot(audio_features_mod['loudness_db'])
ax[0].set_ylabel('loudness_db')
ax[1].plot(librosa.hz_to_midi(audio_features['f0_hz']))
ax[1].plot(librosa.hz_to_midi(audio_features_mod['f0_hz']))
ax[1].set_ylabel('f0 [midi]')
ax[2].plot(audio_features_mod['f0_confidence'])
ax[2].plot(np.ones_like(audio_features_mod['f0_confidence']) * f0_confidence_threshold)
ax[2].set_ylabel('f0 confidence')
_ = ax[2].set_xlabel('Time step [frame]')
# + cellView="form" colab={} colab_type="code" id="SLwg1WkHCXQO"
#@title #Resynthesize Audio
af = audio_features if audio_features_mod is None else audio_features_mod
# Run a batch of predictions.
start_time = time.time()
audio_gen = model(af, training=False)
print('Prediction took %.1f seconds' % (time.time() - start_time))
# Plot
print('Original')
play(audio)
print('Resynthesis')
play(audio_gen)
specplot(audio)
plt.title("Original")
specplot(audio_gen)
_ = plt.title("Resynthesis")
| ddsp/colab/demos/timbre_transfer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("./mnist/data/", one_hot=True)
X = tf.placeholder(tf.float32, [None, 784])
Y = tf.placeholder(tf.float32, [None, 10])
dropProb = tf.placeholder(tf.float32)
phase = tf.placeholder(tf.bool, name='phase')
W1 = tf.Variable(tf.random_normal([784, 256], stddev=0.01))
# L1 = tf.layers.batch_normalization(W1, training=phase)
L1 = tf.nn.relu(tf.matmul(X, W1))
L1 = tf.nn.dropout(L1, dropProb)
W2 = tf.Variable(tf.random_normal([256, 256], stddev=0.01))
# L2 = tf.layers.batch_normalization(W2, training=phase)
L2 = tf.nn.relu(tf.matmul(L1, W2))
L2 = tf.nn.dropout(L2, dropProb)
W3 = tf.Variable(tf.random_normal([256, 10], stddev=0.01))
model = tf.matmul(L2, W3)
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits_v2(logits=model, labels=Y))
optimizer = tf.train.AdamOptimizer(0.001).minimize(cost)
init = tf.global_variables_initializer()
sess = tf.Session()
sess.run(init)
batch_size = 100
total_batch = int(mnist.train.num_examples / batch_size)
for epoch in range(30):
total_cost = 0
for i in range(total_batch):
x_batch, y_batch = mnist.train.next_batch(batch_size)
# _, cost_val = sess.run([optimizer ,cost], feed_dict={X: x_batch, Y: y_batch, dropProb: 0.8, phase: True})
_, cost_val = sess.run([optimizer ,cost], feed_dict={X: x_batch, Y: y_batch, dropProb: 0.8})
# _, cost_val = sess.run([optimizer ,cost], feed_dict={X: x_batch, Y: y_batch, phase: True})
total_cost += cost_val
print("epoch : %04d" % (epoch + 1), "Avg. cost = %.3f" % (total_cost/total_batch))
is_correct = tf.equal(tf.argmax(model, 1), tf.argmax(Y, 1))
accuracy = tf.reduce_mean(tf.cast(is_correct, tf.float32))
print("Accuracy : " , (sess.run(accuracy, feed_dict = {X: mnist.test.images, Y: mnist.test.labels, dropProb: 1., phase: False})))
print("Accuracy : " , (sess.run(accuracy, feed_dict = {X: mnist.test.images, Y: mnist.test.labels, dropProb: 1.})))
# print("Accuracy : " , (sess.run(accuracy, feed_dict = {X: mnist.test.images, Y: mnist.test.labels, phase: False})))
| 3minTensorflow/06. MNIST/06-3. MNIST_Result_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from scale import minmax_scale
import matplotlib.pyplot as plt
import numpy as np
from scipy import stats
import sklearn.preprocessing
from sklearn.model_selection import train_test_split
import pandas as pd
from wrangle import split_telco
import wrangle
from wrangle import clean_telco
df = wrangle.acquire_telco()
df = clean_telco(df)
train, validate, test = split_telco(df)
train_scaled, validate_scaled, test_scaled = minmax_scale(train, validate, test)
train_scaled.head()
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # FP Growth Approach to determine Frequent Item Sets
# + active=""
# Student Name : <NAME>
# Student ID : 2040973
#
#
# Steps :
#
# Step1: Read the csv file
# Step2: Input Min support
# Step3: Find the unique items and their support count
# Step4: Sort the list
# Step5: Build FP Tree from the obtained support count table
# Step6: Mine the FP Tree to find frequent sets
# Step7: Print the output and write it to a file
# -
#open file to write the intermediate updates
output_file = open("Z:\Big Data and Data Mining\Assignments\CourseWork1\FPGrowth.txt",'w')
#Import libararies
from timeit import default_timer as timer
import os
os.environ['KMP_DUPLICATE_LIB_OK']='True'
#Read minimum support
freq_itemset = []
min_support = input("Enter the value of minimum support\n")
#Class FP Tree Node
class FPTreeNode:
def __init__(self, name,count,parent_node):
self.name = name
self.count = count
self.link = None
self.parent = parent_node
self.children = {}
def __str__(self, level=0):
ret = "\t"*level+repr(self.name)+"\n"
for child in self.children:
ret += (self.children[child]).__str__(level+1)
return ret
def __repr__(self):
return '<fpnode>'
def increment_counter(self, count):
self.count += count
#Add fp node to a specific position
def add_fp_nodes(item, fptree, header, count):
if item[0] in fptree.children:
fptree.children[item[0]].increment_counter(count)
else:
fptree.children[item[0]] = FPTreeNode(item[0], count, fptree)
if header[item[0]][1] == None:
header[item[0]][1] = fptree.children[item[0]]
else:
while (header[item[0]][1].link != None):
header[item[0]][1] = header[item[0]][1].link
header[item[0]][1].link = fptree.children[item[0]]
if len(item) > 1:
add_fp_nodes(item[1::], fptree.children[item[0]], header, count)
#Traversal
def traverse(leaf, path):
if leaf.parent != None:
path.append(leaf.name)
traverse(leaf.parent,path)
#Function to read a file, and build fptree based on the data provided
def read_and_createTree(set_dictionary,read_from_file):
if(read_from_file == 1):
file_content = list()
with open('Z:\Big Data and Data Mining\Assignments\CourseWork1\GroceryStore.csv') as f:
rows = f.readlines()
for i in rows:
line = i.strip().strip(",")
file_content.append(line.split(','))
set_dictionary = {}
for j in file_content:
if frozenset(j) in set_dictionary.keys():
set_dictionary[frozenset(j)] += 1
else:
set_dictionary[frozenset(j)] = 1
header = {}
for k in set_dictionary:
for key in k:
header[key] = header.get(key,0) + set_dictionary[k]
for l in list(header):
if(header[l] < int(min_support)):
del(header[l])
freq_items = set(header.keys())
for m in header:
header[m] = [header[m], None]
Tree = FPTreeNode('Null',1,None)
for item_set_fptree,count in set_dictionary.items():
tx = {}
for item in item_set_fptree:
if item in freq_items:
tx[item] = header[item][0]
if len(tx) > 0:
ordered_itemset = [v[0] for v in sorted(tx.items(), key=lambda p: p[1], reverse=True)]
#the nodes are updated into tree
add_fp_nodes(ordered_itemset, Tree, header, count)
return Tree, header
#Function to mine the obtained fptree adn generate frequent item sets
def mining(fptree, header, preset, freq_itemset):
FP = [x[0] for x in sorted(header.items(),key=lambda p: p[1][0])]
for base in FP:
frequent_item_set = preset.copy()
frequent_item_set.add(base)
freq_itemset.append(frequent_item_set)
pattern_base = {}
while header[base][1] != None:
path = []
traverse(header[base][1],path)
if len(path) > 1:
pattern_base[frozenset(path[1:])] = header[base][1].count
header[base][1] = header[base][1].link
conditional_Tree, conditional_header = read_and_createTree(pattern_base,0)
if(conditional_header != None):
mining(conditional_Tree, conditional_header,frequent_item_set, freq_itemset)
start = timer()
fp_Dict = {}
fptree,header = read_and_createTree(fp_Dict,1)
preset = set([])
mining(fptree,header,preset,freq_itemset)
end = timer()
# +
fptree_str =str(fptree)
freq_item_str = str(freq_itemset)
total_time = end-start
output_file.write("1.minimum support :" + min_support)
print("1.minimum support:"+min_support)
output_file.write("\n\n2.FP Growth Approach:")
print("\n\n2 FP Growth Approach:")
output_file.write("\n\n\n3.FP_tree:\n"+fptree_str)
print("\n\n\n3.FP_tree:\n"+fptree_str)
output_file.write("\n\n\n4.Frequent_sets:\n"+str(freq_itemset))
print("\n\n\n4.Frequent_sets:\n"+str(freq_itemset))
output_file.write("\n\n5.Time taken for execution using bruteforce approach:"+str(total_time))
print("\n\n5.Time taken for execution using FP Growth approach:" + str(total_time))
# -
#Close output file after writing
output_file.close()
| FPGrowth.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
transactions = pd.read_csv('csv/train_transaction.csv')
transactions.head()
data = transactions[['TransactionAmt','ProductCD','isFraud']]
data = data.loc[data['isFraud'] == 1]
data = data.loc[data['TransactionAmt'] <= 500]
data
sns.set(rc={"figure.dpi":200,'figure.figsize':(15.2,6.6)})
sns.histplot(data=data, x="TransactionAmt", hue="ProductCD",palette='tab10')
ax.set_title("Datos estadisticos sobre las cantidades pagadas segun el tipo de IP",fontsize=15)
ax.set_ylabel("Cantidad de $USD",fontsize=14)
ax.set_xlabel("Tipo de IP",fontsize=14)
| histograma.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # Dog Breed Classification
#
# This example is based on a very popular [Udacity project](https://github.com/udacity/dog-project), upgraded to use TensorFlow `2.3.0` with GPU accelaration. The goal is to classify images of dogs according to their breed.
#
# In this notebook, we take the first steps towards developing an algorithm that could be used as part of a mobile or web app. At the end of this project, our code will accept any user-supplied image as input. If a dog is detected in the image, it will provide an estimate of the dog's breed. In this real-world setting, we will piece together a series of state-of-the-art computer vision models to perform different tasks (Dog detection -> Breed classification).
# + [markdown] tags=[]
# ## Table of contents
#
# We break the notebook into separate steps. Feel free to use the links below to navigate the notebook.
#
# * [Step 0](#step0): Download Datasets and Install Dependencies
# * [Step 1](#step1): Import Datasets
# * [Step 2](#step2): Detect Dogs
# * [Step 3](#step3): Create a CNN to Classify Dog Breeds (from Scratch)
# * [Step 4](#step4): Create a CNN (VGG16) to Classify Dog Breeds (using Transfer Learning)
# * [Step 5](#step5): Create a CNN (ResNet-50) to Classify Dog Breeds (using Transfer Learning)
# * [Step 6](#step6): Write your Own Dog Classifier
# * [Step 7](#step7): Test Your Classifier
# + [markdown] colab_type="text" id="VvAyLIR-R0af" tags=[]
# <a id='step0'></a>
# ## Step 0: Download Datasets and Install Dependencies
#
# For this task we use TensorFlow `2.3.0`, as well as a few helper libraries like `Pillow`. Also, we need to download and extract the dataset we will use to train our classifier.
# + [markdown] colab_type="text" id="Fm1cOPKZSJQB" tags=[]
# ### Download the dataset
#
# The dataset is available [here](https://s3-us-west-1.amazonaws.com/udacity-aind/dog-project/dogImages.zip). The following cell downloads it as a zip file, extracts it and moves it to the corresponding folder. Finally, it removes the `zip` file.
# + colab={} colab_type="code" id="5MOpDJZCQGRb" tags=["skip"]
# !wget https://s3-us-west-1.amazonaws.com/udacity-aind/dog-project/dogImages.zip
# !unzip -qo dogImages.zip
# !rm dogImages.zip
# + [markdown] colab_type="text" id="q3iHERkNSNY-" tags=[]
# ### Install dependencies
#
# The task requires `Pillow` the friendly PIL fork by [<NAME> and Contributors](https://github.com/python-pillow/Pillow/graphs/contributors). PIL is the Python Imaging Library by <NAME> and Contributors. Below, we install the `Pillow` package using `pip`.
# + colab={} colab_type="code" id="WU7JZLVSRyAT" tags=["skip"]
# !pip3 install --user -r requirements/requirements-v2.txt
# + [markdown] colab_type="text" id="67p9fU72SQE9" tags=[]
# ### Import the necessary libraries
#
# We use Tensorflow `2.3.0` to build and train our dog breed classifier. We also need `Pillow` to load the images in memory, which we specifically instruct to load any truncated images also.
# + colab={} colab_type="code" id="7X11SD82SUhb" tags=["imports"]
import os
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from glob import glob
from PIL import Image
from PIL import ImageFile
ImageFile.LOAD_TRUNCATED_IMAGES = True
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="DKq8rHg5SWXq" outputId="7056f7a7-d327-4622-a59a-32b2108977f4" tags=["skip"]
print(f"Version of TensorFlow in use: {tf.__version__}")
print(f"Using GPU device: {tf.config.list_physical_devices('GPU')}")
# + [markdown] colab_type="text" id="ac4gM-_KSkV4" tags=[]
# <a id='step1'></a>
# ## Step 1: Import Datasets
#
# First, let us define the pipeline-parameters cell. We use it to define the hyperparametes we would like to tune later. These variables will be converted to KFP pipeline parameters, so we should make sure they are used as global variables throughout the notebook.
# + colab={} colab_type="code" id="ifMA-P1ySYK2" tags=["pipeline-parameters"]
LR = 6e-4
BATCH_SIZE = 32
NUMBER_OF_NODES = 256
EPOCHS = 4
IMG_SIZE = 224
# + [markdown] colab_type="text" id="EsWsiKbhTN4E" tags=[]
# ### Processing the Dataset
#
# We use TensorFlow native generators to load and transform the data. Pay attention to the `train_datagen` which also includes several transformations to augment our dataset (e.g. width and height shift, brightness alterations and horizontal flip). These transformations are taking place in memory, leaving the original data untouched.
# + tags=["block:data_processing"]
def get_train_generator():
data_datagen = ImageDataGenerator(
rescale=1./255,
width_shift_range=.2,
height_shift_range=.2,
brightness_range=[0.5,1.5],
horizontal_flip=True
)
return data_datagen.flow_from_directory(
"dogImages/train/",
target_size=(IMG_SIZE, IMG_SIZE),
batch_size=BATCH_SIZE,
)
def get_valid_generator():
data_datagen = ImageDataGenerator(rescale=1./255)
return data_datagen.flow_from_directory(
"dogImages/valid/",
target_size=(IMG_SIZE, IMG_SIZE),
batch_size=BATCH_SIZE
)
def get_test_generator():
data_datagen = ImageDataGenerator(rescale=1./255)
return data_datagen.flow_from_directory(
"dogImages/test/",
target_size=(IMG_SIZE, IMG_SIZE),
batch_size=BATCH_SIZE
)
# + [markdown] colab_type="text" id="r021ns1ZUzi4" tags=[]
# <a id='step2'></a>
# ## Step 2: Detect Dogs
#
# In this section, we use a pre-trained [ResNet V2](https://link.springer.com/chapter/10.1007/978-3-319-46493-0_38) model to detect dogs in images. First, we download a pretrained ResNet-50 model on [ImageNet](http://www.image-net.org/), a very large, very popular dataset used for image classification and other computer vision tasks. ImageNet contains over 10 million URLs, each linking to an image containing an object from one of 1000 categories. Given an image, this pre-trained ResNet-50 model returns a prediction (derived from the available categories in ImageNet) for the object that is contained in the image.
# + colab={} colab_type="code" id="PgWSAhN-TFNf" tags=["block:dog_detector", "prev:data_processing", "limit:nvidia.com/gpu:1"]
dog_classifier = tf.keras.applications.ResNet50V2(
weights="imagenet",
input_shape=(IMG_SIZE, IMG_SIZE, 3)
)
# + [markdown] tags=[]
# The categories corresponding to dogs appear in an uninterrupted sequence referring to dictionary keys 151-268, inclusive, to include all categories from 'Chihuahua' to 'Mexican hairless'. Thus, in order to check if an image is predicted to contain a dog by the pre-trained ResNet model, we need only check if the function below returns a value between 151 and 268 (inclusive).
#
# We use these ideas to complete the `is_dog` function below, which returns True if a dog is detected in an image.
# + colab={} colab_type="code" id="lguOZx_PWE73" tags=[]
def is_dog(data):
probs = dog_classifier.predict(data)
preds = tf.argmax(probs, axis=1)
return ((preds >= 151) & (preds <= 268))
# + [markdown] tags=[]
# To test the classifier we request a batch from our training data generator and feed it through the network. The accuracy, as expected, is really high.
# + colab={} colab_type="code" id="7AkKSHeaYIcg" tags=[]
train_generator = get_train_generator()
batch = train_generator.next()
predictions = is_dog(batch)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="wGeUt1jidjat" outputId="19f96954-ed31-4ab5-917a-53178f696e10" tags=[]
n_dog = np.sum(predictions)
dog_percentage = n_dog/BATCH_SIZE
print('{:.0%} of the files have a detected dog'.format(dog_percentage))
# + [markdown] colab_type="text" id="8WJ6MCYrf4sR" tags=[]
# <a id='step3'></a>
# ## Step 3: Create a CNN to Classify Dog Breeds (from Scratch)
#
# Now that we have a function for detecting dogs in images, we need a way to predict the dog breed from images. In this step, you will create a CNN that classifies dog breeds. We should be careful with adding too many trainable layers though. More parameters means longer training and you may also fall in the trap of overfitting. Thankfully, `tf.keras` provides a handy estimate of the time that each epoch is likely to take; you can extrapolate this estimate to figure out how long it will take for your algorithm to train.
#
# We mention that the task of assigning breed to dogs from images is considered exceptionally challenging. To see why, consider that *even a human* would have great difficulty in distinguishing between a Brittany and a Welsh Springer Spaniel.
#
# Brittany | Welsh Springer Spaniel
# - | -
# <img src="images/Brittany_02625.jpg" width="100"> | <img src="images/Welsh_springer_spaniel_08203.jpg" width="200">
#
# It is not difficult to find other dog breed pairs with minimal inter-class variation (for instance, Curly-Coated Retrievers and American Water Spaniels).
#
# Curly-Coated Retriever | American Water Spaniel
# - | -
# <img src="images/Curly-coated_retriever_03896.jpg" width="200"> | <img src="images/American_water_spaniel_00648.jpg" width="200">
#
# Likewise, recall that labradors come in yellow, chocolate, and black. Your vision-based algorithm will have to conquer this high intra-class variation to determine how to classify all of these different shades as the same breed.
#
# Yellow Labrador | Chocolate Labrador | Black Labrador
# - | - | -
# <img src="images/Labrador_retriever_06457.jpg" width="150"> | <img src="images/Labrador_retriever_06455.jpg" width="240"> | <img src="images/Labrador_retriever_06449.jpg" width="220">
#
# We also mention that random chance presents an exceptionally low bar: setting aside the fact that the classes are slightly imabalanced, a random guess will provide a correct answer roughly 1 in 133 times, which corresponds to an accuracy of less than 1%.
# + [markdown] tags=[]
# ### Model architecture
#
# Create a CNN to classify dog breed. At the end of your code cell block, summarize the layers of your model by executing `model.summary()`.
# + colab={} colab_type="code" id="cX4dZkEjfe6u" tags=["block:custom_classifier", "prev:data_processing", "limit:nvidia.com/gpu:1"]
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(16, 3, activation="relu", input_shape=(IMG_SIZE, IMG_SIZE, 3)),
tf.keras.layers.MaxPool2D(),
tf.keras.layers.Conv2D(32, 3, activation="relu"),
tf.keras.layers.MaxPool2D(),
tf.keras.layers.Conv2D(64, 3, activation="relu"),
tf.keras.layers.MaxPool2D(),
tf.keras.layers.GlobalAveragePooling2D(),
tf.keras.layers.Dense(NUMBER_OF_NODES, activation="relu"),
tf.keras.layers.Dense(133, activation="softmax")
])
# + colab={"base_uri": "https://localhost:8080/", "height": 459} colab_type="code" id="iMmibLNahFRp" outputId="31a84afd-576e-4262-efea-978fd98d0255" tags=[]
model.summary()
# + [markdown] tags=[]
# The next step is to compile the model. For this, we need to pass an optimizer and a loss function. We can also pass a list of metrics we want. In this example, we pass the _accuracy_ metric.
# + colab={} colab_type="code" id="9uRX3_UyhN90" tags=[]
model.compile(
optimizer=tf.optimizers.Adam(learning_rate=LR),
loss=tf.losses.categorical_crossentropy,
metrics=["accuracy"]
)
# + [markdown] tags=[]
# Finally, we can train the model using the `fit` method. This runs on batches yielded by the data generator and prints out the _loss_ and _accuracy_ both for train and validation sets.
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="Vj67Ok9dhbOC" outputId="2e64d6cf-0c21-4fac-c51f-d88661d85923" tags=[]
train_generator = get_train_generator()
valid_generator = get_valid_generator()
tb_callback = tf.keras.callbacks.TensorBoard(log_dir="custom_classifier_logs")
model.fit(train_generator, epochs=2,
validation_data=valid_generator,
callbacks=[tb_callback]
)
# + [markdown] tags=[]
# ### Evaluation
#
# To evaluate the final model we feed it with the test dataset and call the `evaluate` method.
# + tags=["block:custome_classifier_eval", "prev:custom_classifier", "limit:nvidia.com/gpu:1"]
test_generator = get_test_generator()
test_loss_custom, test_accuracy_custom = model.evaluate(test_generator)
print(f"The accuracy in the test set is {test_accuracy_custom:.3f}.")
# + [markdown] colab_type="text" id="Q-kl1HXWo0mP" tags=[]
# <a id='step4'></a>
# ## Step 4: Create a CNN (VGG16) to Classify Dog Breeds (using Transfer Learning)
#
# To reduce training time without sacrificing accuracy, we train a CNN using Transfer Learning. Transfer Learning is the fine-tuning of a network that was pre-trained on some big dataset with new classification layers. The idea behind is that we want to keep all the good features learned in the lower levels of the network (because there's a high probability the new images will also have those features) and just learn a new classifier on top of those. This tends to work well, especially with small datasets that don't allow for a full training of the network from scratch (it's also much faster than a full training).
#
# One way of doing Transfer Learning is by loading a pretrained model up to a point, usually chopping off the final dense part of the model and adding a fully connected layer with the output that we want (e.g. an 133-node classifier). Then, we freeze the first part of the model (i.e. the body) and train only the final layer we added.
# + colab={} colab_type="code" id="j6fsA2R8nVBa" tags=["block:vgg16_classifier", "prev:data_processing", "limit:nvidia.com/gpu:1"]
vgg_body = tf.keras.applications.VGG16(
weights="imagenet",
include_top=False,
input_shape=(IMG_SIZE, IMG_SIZE, 3)
)
# + colab={} colab_type="code" id="szJl_qQ9sZuz" tags=[]
vgg_body.trainable = False
# + colab={} colab_type="code" id="YdbFhBtIp-Jx" tags=[]
inputs = tf.keras.layers.Input(shape=(IMG_SIZE, IMG_SIZE, 3))
# We make sure that the vgg_body is running in inference mode here,
# by passing `training=False`. This is important for fine-tuning, as you will
# learn in a few paragraphs.
x = vgg_body(inputs, training=False)
# Convert features of shape `vgg_body.output_shape[1:]` to vectors
x = tf.keras.layers.GlobalAveragePooling2D()(x)
# A Dense classifier (categorical classification)
outputs = tf.keras.layers.Dense(133, activation="softmax")(x)
vgg_model = tf.keras.Model(inputs, outputs)
# + colab={"base_uri": "https://localhost:8080/", "height": 289} colab_type="code" id="9I2yPjo_qMTL" outputId="25b81b5f-e3d3-411c-d011-a20e18d38a11" tags=[]
vgg_model.summary()
# + colab={} colab_type="code" id="kTMJ5joCqpTE" tags=[]
vgg_model.compile(
optimizer=tf.optimizers.Adam(learning_rate=LR),
loss=tf.losses.categorical_crossentropy,
metrics=["accuracy"]
)
# + colab={"base_uri": "https://localhost:8080/", "height": 459} colab_type="code" id="8IoRWrQnq5c2" outputId="3f69f065-815e-4c9c-b665-ecc2a10337b6" tags=[]
train_generator = get_train_generator()
valid_generator = get_valid_generator()
vgg_model.fit(train_generator, epochs=2,
validation_data=valid_generator
)
# + [markdown] tags=[]
# ### Evaluation
#
# To evaluate the model on the test set we call the same `evaluate` method.
# + tags=["block:vgg16_classifier_eval", "prev:vgg16_classifier"]
test_generator = get_test_generator()
test_loss_vgg, test_accuracy_vgg = vgg_model.evaluate(test_generator)
print(f"The accuracy in the test set is {test_accuracy_vgg:.3f}.")
# + [markdown] tags=[]
# <a id='step5'></a>
# ## Step 5: Create a CNN (ResNet-50) to Classify Dog Breeds (using Transfer Learning)
#
# In this section, we will use the same procedure but with a pretrained ResNet-50 model.
# + tags=["block:resnet50_classifier", "prev:data_processing", "limit:nvidia.com/gpu:1"]
resnet_body = tf.keras.applications.ResNet50V2(
weights="imagenet",
include_top=False,
input_shape=(IMG_SIZE, IMG_SIZE, 3)
)
# + tags=[]
resnet_body.trainable = False
# + tags=[]
inputs = tf.keras.layers.Input(shape=(IMG_SIZE, IMG_SIZE, 3))
# We make sure that the vgg_body is running in inference mode here,
# by passing `training=False`. This is important for fine-tuning, as you will
# learn in a few paragraphs.
x = resnet_body(inputs, training=False)
# Convert features of shape `vgg_body.output_shape[1:]` to vectors
x = tf.keras.layers.Flatten()(x)
# A Dense classifier (categorical classification)
outputs = tf.keras.layers.Dense(133, activation="softmax")(x)
resnet_model = tf.keras.Model(inputs, outputs)
# + tags=[]
resnet_model.compile(
optimizer=tf.optimizers.Adam(learning_rate=LR),
loss=tf.losses.categorical_crossentropy,
metrics=["accuracy"]
)
# + tags=[]
train_generator = get_train_generator()
valid_generator = get_valid_generator()
resnet_model.fit(train_generator, epochs=EPOCHS,
validation_data=valid_generator
)
# + [markdown] tags=[]
# ### Evaluation
# + tags=["block:resnet50_classifier_eval", "prev:resnet50_classifier", "limit:nvidia.com/gpu:1"]
test_generator = get_test_generator()
test_loss_resnet, test_accuracy_resnet = resnet_model.evaluate(test_generator)
print(f"The accuracy in the test set is {test_accuracy_resnet:.3f}.")
# + [markdown] tags=[]
# <a id='step6'></a>
# ## Step 6: Write your Own Dog Classifier
#
# To create our own classifier we need a class to predict if there is a dog in the image and if that's true, return the breed. For the first part we use the `dog_classifier` method and then predict the breed using the `predict_breed` method.
# + tags=["skip"]
idx_to_class = {value: key for key, value in train_generator.class_indices.items()}
def predict_breed(images):
probs = resnet_model.predict(images)
pred = tf.argmax(probs, axis=1)
label = idx_to_class[pred.numpy()[0]]
return label.split(".")[-1]
# + tags=["skip"]
def predict_dog(image):
image = image[None,...]
if is_dog(image):
pred = predict_breed(image)
print(f"This photo looks like a(n) {pred}.")
return
print("No dog detected")
image = train_generator.next()[0][0]
plt.imshow(image)
plt.show()
pred = predict_dog(image)
# + [markdown] tags=[]
# <a id='step7'></a>
# ## Step 7: Test Your Classifier
#
# In the last section, we take your new algorithm for a spin; if you have a dog, does it predict your dog's breed accurately? If you have a cat, does it mistakenly think that your cat is a dog?
# + tags=["skip"]
for img_path in sorted(glob("check_images/*")):
print(img_path)
img = Image.open(img_path)
img = img.resize((224, 224))
plt.imshow(img)
plt.show()
img = np.array(img)
predict_dog(img)
# + [markdown] tags=[]
# ### Pipeline metrics
#
# This is the pipeline-metrics cell. Use it to define the pipeline metrics that KFP will produce for every pipeline run. Kale will associate each one of these metrics to the steps that produced them. Also, you will have to choose one these metrics as the Katib search objective metric.
# + tags=["pipeline-metrics"]
print(test_accuracy_resnet)
| examples/dog-breed-classification/dog-breed-v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import screed # A Python library for reading FASTA and FASQ file format.
from Bio import SeqIO
class ReadSeq:
"""Class for Reading Sequence Data"""
def readText(self, inputfile):
# opne data
with open(inputfile, "r") as seqfile:
# read data
seq = seqfile.read()
# remove special characters \n and \t
seq = seq.replace("\n", "")
seq = seq.replace("\t", "")
return seq
def readFASTA(self, inputfile):
# open file
with open(inputfile, "r") as f:
# remove name line / info line
seq = f.readline()
# read data
seq = f.read()
# remove special character
seq = seq.replace("\n", "")
seq = seq.replace("\t", "")
return seq
def readFastq(self, filename):
sequences = []
qualities = []
with open(filename) as fh:
while True:
fh.readline() # skip name line
seq = fh.readline().rstrip() # read base sequence
fh.readline() # skip placeholder line
qual = fh.readline().rstrip() #base quality line
if len(seq) == 0:
break
sequences.append(seq)
qualities.append(qual)
return sequences, qualities
def readFastaWithScreed(self, inputfile):
"""
Reads and returns file as FASTA format with special characters removed.
"""
with screed.open(inputfile) as seqfile:
for read in seqfile:
seq = read.sequence
return seq
def readFastqWithScreed(self, inputfile):
"""
Reads and returns file as FASTA format with special characters removed.
"""
with screed.open(inputfile) as seqfile:
for read in seqfile:
seq = read.sequence
return seq
def readFastqWithBiopython(self, inputfile):
"""
Reads and returns file as FASTA format with special characters removed.
"""
with open(inputfile) as seqfile:
for record in SeqIO.parse(seqfile, "fasta"):
seq = record.seq
return seq
# -
# create an object of ReadSeq class
data = ReadSeq()
seq1 = data.readFASTA("../data/Haemophilus_influenzae.fasta")
seq2 = data.readFastq("../data/SRR835775_1.first1000.fastq")
seq3 = data.readFastq("../data/dna.txt")
seq4 = data.readFastaWithScreed("../data/Haemophilus_influenzae.fasta")
seq5 = data.readFastqWithBiopython("../data/Haemophilus_influenzae.fasta")
| old_notebooks/Class for Reading Data .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Domain quantization of video frames
# +
import torch
from torch import nn
import torch.optim as optim
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from scipy.stats import norm
def normal_distribution(n, mean, var=0.05):
x = norm.pdf(np.arange(0, 1, 1.0 / n), mean, var)
x = x / np.sum(x)
#return torch.tensor(x).float()
return x
# +
import pandas as pd
class DomainQuantizationLayer(nn.Module):
def __init__(self, input_features, bin_count=10, init_low_bound=0.0, init_high_bound=1.0):
assert bin_count > 1
super().__init__()
self.input_features = input_features
self.bin_count = bin_count
self.bins = [[] for _ in range(input_features)]
self.bins_initialized = False
self.values_used = 0
def forward(self, input):
assert input.shape[0] == self.input_features, f"self.input_features is {self.input_features}, input is {input.shape}"
# print('input shape = ', input.shape)
if self.training or not self.bins_initialized:
self.update_bins(input)
return self.map_values(input)
def extra_repr(self):
return f"input_features = {self.input_features}, resolution = {self.resolution}"
def qcut_bins(self, values):
assert len(values.shape) == 1, "Need 1D numpy array of values"
_, bins = pd.qcut(values, q=self.bin_count-1, retbins=True, duplicates='drop', labels=False)
return bins
def map_values(self, values):
output = torch.stack([self.map_feature_values(feature_values, feature_index) for feature_index, feature_values in enumerate(values)])
return output
def map_feature_values(self, values, feature_index):
assert len(values.shape) == 1
bins = self.bins[feature_index]
digits = np.digitize(values, bins)
output = digits.astype(float)
for index, digit in enumerate(digits):
if digit > 0 and digit < len(bins):
val_left = bins[digit-1]
val_right = bins[digit]
output[index] += (values[index] - val_left) / (val_right - val_left)
output -= 1
output.clip(0, len(bins) - 1, out=output)
#output = [normal_distribution(self.bin_count, item * 1.0 / self.bin_count) for item in output]
return torch.Tensor(output)
def update_bins(self, values):
alpha = self.values_used / (self.values_used + values.size)
for feature_index, feature_values in enumerate(values):
self.update_feature_bins(feature_values, feature_index, alpha)
self.bins_initialized = True
self.values_used += values.shape[-1]
if self.values_used > 10000:
self.values_used = 10000
def update_feature_bins(self, values, feature_index, alpha):
bins = self.qcut_bins(values)
if not self.bins_initialized or self.bins[feature_index].shape[0] < bins.shape[0]:
self.bins[feature_index] = bins
else:
if self.bins[feature_index].shape[0] == bins.shape[0]:
self.bins[feature_index] = self.bins[feature_index] * alpha + bins * (1 - alpha)
else:
pass # ignore smaller size bins, we never want to reduce resolution
# +
from scipy.ndimage.filters import gaussian_filter
def generate_frames():
width = 20
height = 20
gutter = 10
frames = []
for y in range(gutter-5, gutter+height-5):
for x in range(width+2*gutter):
frame = np.zeros((width + 2 * gutter, height + 2 * gutter))
frame[y:10+y, x:x+3] = 1
frame=gaussian_filter(frame, 0.5)
clipped = frame[gutter:-gutter, gutter:-gutter]
frames.append(clipped)
return np.asarray(frames)
X = generate_frames()
# -
frame_count, width, height = X.shape
X = X.reshape(frame_count, width * height)
print(X.shape)
feature_count = width * height
dq = DomainQuantizationLayer(feature_count, bin_count=200)
# +
batch_start_index = 0
for batch in range(1):
batch_size = frame_count
pixel_data = X[batch_start_index:batch_start_index+batch_size,:].transpose()
output = dq(pixel_data)
# dq = DomainQuantizationLayer(X.shape[-1], bin_count=20)
# output = dq(X.transpose())
output_sample = output[:, 20].numpy().reshape(height, width)
fig, ax1 = plt.subplots(figsize=(15, 5))
ax1.imshow(output_sample)
plt.show()
input_sample = X[batch_start_index+20].reshape(height, width)
fig, ax1 = plt.subplots(figsize=(15, 5))
ax1.imshow(input_sample)
plt.show()
# fig, ax1 = plt.subplots(figsize=(15, 5))
# ax1.imshow(output_sample - input_sample)
# plt.show()
batch_start_index += batch_size
# -
dq.bins
# +
image = X[260].reshape(height, width).transpose().reshape(height * width)
dq.eval()
output = dq(image[:, None])
fig, ax1 = plt.subplots(figsize=(15, 5))
ax1.imshow(output.reshape(height, width))
plt.show()
output = dq(X[260][:, None])
fig, ax1 = plt.subplots(figsize=(15, 5))
ax1.imshow(output.reshape(height, width))
plt.show()
# -
| Domain Quantization/06 - Video domain quantization.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
# # 建立中央氣象局測站列表資料表
#
# ## 目標
#
# 將中央氣象局網頁上的氣象測站點位資料轉換成結構化的資料表,並轉成具有空間資訊的資料表及檔案格式。這樣有什麼好處呢?之後處理資料比較快,而且應用層面也比較廣
#
# ## 必須具備的背景知識/技巧
#
# 雖然說應該要有一些背景知識,但可以照著步驟仿效,再自己慢慢找資料學。底下兩項是基礎的技能
#
# * 資料庫(知道什麼是 table, constraints, etc.)
# * 為了防止意外,我們使用 [transaction](https://www.postgresql.org/docs/9.5/static/sql-begin.html) 來處理 SQL command,也就是
#
# ```
# 1 BEGIN;
# 2 SQL
# 3 ...
# 4 COMMIT;
# ```
#
# * shell programming (case switch, EOF(end-of-file, stream text), i/o, sed/awk 這類的 stream editor),但都不會太難,照範例做改一下就行了。
# * ```#``` 開頭的為 shell script 的註解
# * ```--``` 開頭的為 SQL 的註解
# * EOF 的使用,用 ```cat > filename << _EOF ... _EOF```,中間包起來的文字會存進 ```filename``` 這個檔案中
# ```
# 1 cat > filename << _EOF
# 2
# 3 abc
# 4
# 5 _EOF
# ```
# 例如上面的源碼中,我們把三行(空白行、abc、空白行)插入到 ```filename``` 中
#
# ## 工具
#
# * [PostgreSQL](http://postgresql.org) / [PostGIS extension](http://postgis.net)
# 要先把 PostgreSQL 架好至少可以用
# * GNU utilities, such as sed, awk
# 在 windows 下面使用會有些痛苦,建議你可以用 [cygwin](https://www.cygwin.com)
#
#
# ## 0. 看一下資料長什麼樣子
#
# 網頁資料:
# http://e-service.cwb.gov.tw/wdps/obs/state.htm#%B2%7B%A6s%B4%FA%AF%B8
#
# Big5 編碼,用表格處理起來有點麻煩,所以乾脆貼到 LibreOffice 轉成 csv 檔(| 分隔)
#
# 現存測站資料
head ../data/cwb_current_station_list.csv
# 撤站的資料
head ../data/cwb_revoked_station_list.csv
# ## 1. 清理資料
#
# 把不要的半型空白和全型空白都去除,另外把日期整理成 ISO-8601 格式 (YYYY-mm-dd)。
#
# 1.1 用 ```case``` 來做,```$1``` 代表輸入時的第一個參數,```$2``` 為第二個,以此類推。例如:
# ```sh
# command a b
# ```
# 其中```$1``` 為 ```a```,而 ```$2``` 為 ```b```
#
# 1.2 用 sed 來取代字串,那些全形半形的空白很惱人,所以先把它們都除掉
# +
cat > ./cleandata.sh << __EOF
#!/usr/bin/env sh
case "\$1" in
"-c")
# 取代半型空白
sed -i '' 's/ //g' \$2
# 取代全形空白
sed -i '' 's/ //g' \$2
echo "clean up done!"
;;
"-d")
# 把 YYYY/mm/dd 改成 ISO8601 格式(YYYY-mm-dd)
sed -i '' 's/\//-/g' \$2
echo "substitution done!"
;;
*)
echo "Usage: cleandata.sh {-c|-d} filename"
echo "清理資料,把全/半形空白取代並且整理日期為 ISO 8601 格式(YYYY-mm-dd)"
;;
esac
__EOF
# -
./cleandata.sh -c ../data/cwb_current_station_list.csv
./cleandata.sh -d ../data/cwb_revoked_station_list.csv
# ## 2. 建立 postgresql Table
# +
# 設定變數,資料庫名稱為 nvdimp
export DB="nvdimp"
# 用資料定義語言(Data definition language, DDL)來建立新的資料表(table),資料表名稱為
cat > create_station_list_table.sql << _EOF
-- 如果 cwb_station_list 資料表存在的話,則刪除之
DROP TABLE IF EXISTS cwb_station_list;
CREATE TABLE public.cwb_station_list
(
station_id character varying NOT NULL, -- 站號
station character varying, -- 站名
county character varying, -- 縣市
township character varying, -- 鄉鎮
longitude double precision, -- 經度
latitude double precision, -- 緯度
elevation double precision, -- 海拔高度
address character varying, -- 地址
establish_date date, -- 資料起始日期
revoke_date date, -- 撤站日期
note character varying, -- 備註
twd97x double precision, -- TWD97X 座標
twd97y double precision, -- TWD97Y 座標
geom_twd97 geometry(Point,3826), -- twd97 geometry column
geom_wgs84 geometry(Point,4326), -- wgs84 geometry column
station_type character varying, -- 測站類型
CONSTRAINT cwb_stations_pk PRIMARY KEY (station_id)
)
WITH (
OIDS=FALSE
);
_EOF
# 建立資料表架構
# nvdimp 為資料庫名稱
psql -d ${DB} -f create_station_list_table.sql
# -
# ## 3. 餵資料進去 Table
#
# 因為欄位順序不太一樣,所以先把測站的代碼(```station_id```)丟進去
# 先處理現存測站
echo "BEGIN;" > insert_station_id_data.sql
awk -v q="'" -F'|' 'NR>1 { print "INSERT INTO cwb_station_list (station_id) VALUES (" q$1q ");"}' \
../data/cwb_current_station_list.csv >> insert_station_id_data.sql
echo "COMMIT;" >> insert_station_id_data.sql
# execute sql
psql -q -d ${DB} -f insert_station_id_data.sql
# 餵其他資料進去,這邊使用
# ```SQL
# UPDATE _table_ SET _column_name_ = _value_;
# ```
# 來將資料更新進去
# +
# 欄位的位置編號對照
# 站號|站名|海拔高度(M)|經度|緯度|城市|地址|資料起始日期|撤站日期|備註|原站號|新站號
# 1 2 3 4 5 6 7 8 9 10 11 12
# awk 中可以用 $number (number 為正整數) 代表欄位
echo "BEGIN;" > update_station_data.sql
awk -v q="'" -F '|' \
'NR>1 { print "UPDATE cwb_station_list SET station=" q$2q \
", elevation = " $3 \
", longitude = " $4 \
", latitude = " $5 \
", county = " q$6q \
", address = " q$7q \
", establish_date = " q$8q \
", note = "q$10q " WHERE station_id=" q$1q ";"}' \
../data/cwb_current_station_list.csv >> update_station_data.sql
echo "COMMIT;" >> update_station_data.sql
# 執行
psql -q -d ${DB} -f update_station_data.sql
# -
# ## 4. 建立空間屬性的欄位
#
# 接下來我們要把測站的經緯度建立具有空間屬性(spatial geometry)的資料表欄位,在上面 DDL 建立資料表時,已經建立了兩個欄位,```geom_wgs84``` 及 ```geom_twd97``` 兩個空間屬性欄位,分別是 WGS84 經緯度座標([EPSG:4326](http://spatialreference.org/ref/epsg/wgs-84/))以及臺灣使用的橫麥卡托投影二度分帶座標系統(即 TWD97 TM2 Zone 121, [EPSG:3826](http://spatialreference.org/ref/epsg/twd97-tm2-zone-121/),以下使用 TWD97 來縮寫)。原始資料中提供的只有經緯度,為了方便之後的使用,所以我也同時轉換成 TWD97 的座標系統。
#
# 使用的 PostGIS function:
#
# * [ST_SetSRID](http://postgis.net/docs/ST_SetSRID.html): 設定 SRID (spatial reference ID)
# * [ST_MakePoint](http://postgis.net/docs/ST_MakePoint.html): 將 x, y 座標建立具空間屬性的點位
# * [ST_Transform](http://postgis.net/docs/ST_Transform.html): 轉換不同的座標系統
#
#
# 同樣的我們使用```UPDATE```加上 subquery 來更新資料:
#
# **<font style="color:red">注意:測站資料若是在外島,像是金門、馬祖要另外處理成 TWD97 TM2 zone 119 的投影座標系統,
# 在此先略過不提,但請記得這件事情</font> **
# +
# 更新 geometry,先處理 WGS84
cat > set_srid.sql << _EOF
BEGIN;
UPDATE
cwb_station_list
SET geom_wgs84 = s.wgs84
FROM
(SELECT station_id, st_setsrid(st_makepoint(longitude,latitude), 4326) wgs84
FROM cwb_station_list) as s
WHERE
cwb_station_list.station_id = s.station_id;
COMMIT;
-- 轉換座標系統
BEGIN;
UPDATE
cwb_station_list
SET
geom_twd97 = st_transform(geom_wgs84, 3826);
COMMIT;
_EOF
# 執行上面的 set_srid.sql
psql -d ${DB} -f set_srid.sql
# -
# 接下來你可以用 QGIS 來看顯示的座標對不對(略)
# ## 5. 轉成不同格式的檔案
#
# 我們可以把資料 dump 成 SQL,或是轉成 ESRI shapefile, json 等格式
# pg_dump
pg_dump -d ${DB} -O -t cwb_station_list > ../data/cwb_station_list.sql
# +
# 轉成 ESRI Shapefile,分別轉成 wgs84 和 twd97 tm2 zone 121
# 先建立 shp 資料夾,這樣才不會看起來一堆檔案亂七八糟
if [ ! -d ../data/shp ]; then
mkdir ../data/shp
fi
pgsql2shp -f ../data/shp/cwb_station_list_wgs84 -g geom_wgs84 nvdimp public.cwb_station_list
pgsql2shp -f ../data/shp/cwb_station_list_twd97 -g geom_twd97 nvdimp public.cwb_station_list
# -
# ## 參考資料:
# [PostGIS manual](http://postgis.net/docs)
| src/建立中央氣象局測站列表.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Feature extraction: Aspect Ratio, Echogenicity, Composition
# #### Micro-calicification is extracted by U-Net
# +
import numpy as np
from matplotlib import pyplot as plt
import cv2
from skimage import draw
from skimage.morphology import erosion
from feature_extractor import ThyroidFeature
# +
image = cv2.imread('./data/P_TT-2271.png', cv2.IMREAD_GRAYSCALE)
mask = cv2.imread('./data/P_TT-2271_mask.png', cv2.IMREAD_GRAYSCALE)
thyroid_feature = ThyroidFeature('P_TT-2271', image, mask)
# +
def plot_lesion_axis(image, mask, major_axis, minor_axis):
plt.figure(figsize=(8,8))
image = cv2.cvtColor(image, cv2.COLOR_GRAY2BGR)
# add major/minor axis
rr, cc, val = draw.line_aa(major_axis[1][0][0], major_axis[1][0][1], major_axis[1][1][0], major_axis[1][1][1])
image[cc, rr, :] = [0, 0, 255]
rr, cc, val = draw.line_aa(minor_axis[1][0][0], minor_axis[1][0][1], minor_axis[1][1][0], minor_axis[1][1][1])
image[cc, rr, :] = [255, 0, 0]
# Add contour
contour = mask - erosion(mask)
contour_coordinate = np.where(contour != 0)
image[contour_coordinate] = [0, 255, 0]
plt.imshow(image)
return image
major_axis, minor_axis = thyroid_feature.get_lesion_axis()
img_aspect_ratio = plot_lesion_axis(thyroid_feature.image, thyroid_feature.mask, major_axis, minor_axis)
print('aspect ratio: ', thyroid_feature.compute_aspect_ratio())
# +
def plot_echo_image(echo_image, composition):
plt.imshow(echo_image, cmap='gray')
plt.title(str(composition))
echo_image, composition = thyroid_feature.compute_echo()
plot_echo_image(echo_image, composition)
# +
def plot_composition(echo_image, mask):
composition = np.zeros([echo_image.shape[0], echo_image.shape[1],3])
composition[np.where(echo_image > 1)] = [120,0,120]
# Add contour
contour = mask - erosion(mask)
contour_coordinate = np.where(contour != 0)
composition[contour_coordinate] = [0, 255, 0]
plt.imshow(composition)
return composition
echo_image, _ = thyroid_feature.compute_echo()
img_composition = plot_composition(echo_image, thyroid_feature.roi_mask)
print('Quality: ', thyroid_feature.analyze_quality())
# -
# + active=""
# fig = plt.figure(figsize=(20,8))
#
# ax = fig.add_subplot(131)
# ax.set_title('Aspect Ratio')
# ax.imshow(img_aspect_ratio)
#
# ax = fig.add_subplot(132)
# ax.set_title('Echogenicity')
# ax.imshow(echo_image, cmap='gray')
#
# ax = fig.add_subplot(133)
# ax.set_title('Composition')
# ax.imshow(img_composition)
| ai_thyroid_clinical_feature/feature_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tidal Evolution of the Earth-Moon System
#
# ### <NAME>
# +
# The following packages will be used:
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
import math
# %matplotlib inline
plt.style.use('dark_background')
from scipy.integrate import ode
# -
# ## 1.
#
# Here, we pick our unit system.
#
# Because the majority of the quanitities presented in this set are given in cgs, we'll use that as our primary unit system. The other quantities will then be converted, accordingly.
#
# Gravitational constant: $G$ = $ 6.67 {\times} 10^{-8} g^{-1} cm^{3} s^{-2}$
#
#
# Solar mass: $M_{\odot}$ = $1.98 {\times} 10^{33} g$
#
# Earth Mass: $M_{\oplus}$ = $5.97 {\times} 10^{27} g$
#
# Lunar mass: $m_{moon}$ = $7.349 {\times} 10^{25} g$
#
# Earth's radius: $R_{\oplus}$ = $6,371 km$ = $637,100,000 cm$
#
#
# Present day Lunar semimajor axis: $a_{moon}(0)$ = $384,000 km$ = $3.84 {\times} 10^{10} cm$
#
#
# Semimajor axis of Earth's orbit: $a_{\oplus}$ = $1.49 {\times} 10^{8} km$ = $1.49 {\times} 10^{13} cm$
#
# Love number of Earth: $k_{2}$ = $0.298$ (dimensionless)
#
# Tidal quality factor: $Q_{moon} $ = $11.5$ (dimensionless)
#
#
#
# Because we're working with an initial value problem, we need to set our intiial conditions. Thus, we'll proceed by calculating $L_{\oplus}$, $S_{\oplus}$, and $L_{moon}$.
#
#
# Those quantities are given by the following equations:
#
# $L_{\oplus}$ = $M_{\oplus} \sqrt{G(M_{\odot} + M_{\oplus})a_{\oplus}}$
#
# $S_{\oplus}$ = $I{\Omega}_{\oplus}$
#
# where $I$ = $0.3299 M_{\oplus}R^{2}_{\oplus}$
#
# and ${\Omega}_{\oplus}$ = $\frac{2 \pi}{lod}$
#
# where $lod$ = $86164 s$
#
# $L_{moon}$ = $m_{moon} \sqrt{G(M_{\oplus} + m_{moon})a_{moon}}$
# Then performing the calculations:
#
# $L_{\oplus}$ = $M_{\oplus} \sqrt{G(M_{\odot} + M_{\oplus})a_{\oplus}}$
#
# $L_{\oplus}$ = $(5.97 {\times} 10^{27} g) \sqrt{(6.67 {\times} 10^{-8} g^{-1} cm^{3} s^{-2})(1.98 {\times} 10^{33} g + (5.97 {\times} 10^{27} g))(1.49 {\times} 10^{13} cm)}$
#
# $L_{\oplus}$ = $2.648 {\times} 10^{47} g cm^{2} s^{-1}$
#
# Then the next:
#
# $S_{\oplus}$ = $I{\Omega}_{\oplus}$
#
# $S_{\oplus}$ = $0.3299 M_{\oplus}R^{2}_{\oplus} \frac{2 \pi}{lod}$
#
# $S_{\oplus}$ = $0.3299(5.97 {\times} 10^{27} g)(637,100,000 cm)^{2} \frac{2 \pi}{86164 s}$
#
# $S_{\oplus}$ = $5.829 {\times} 10^{40} gcm^{2}s^{-1}$
#
#
# And the final computation:
#
# $L_{moon}$ = $m_{moon} \sqrt{G(M_{\oplus} + m_{moon})a_{moon}}$
#
# $L_{moon}$ = $(7.349 {\times} 10^{25} g) \sqrt{( 6.67 {\times} 10^{-8} g^{-1} cm^{3} s^{-2})((5.97 {\times} 10^{27} g) + (7.349 {\times} 10^{25} g))(3.84 {\times} 10^{10} cm)}$
#
# $L_{moon}$ = $2.891 {\times} 10^{41} g cm^{2} s^{-1}$
# +
# Define variables
G = 6.67e-8
M_Earth = 5.97e27
M_Sun = 1.98e33
m_moon = 7.349e25
R_Earth = 6.371e8
a_moon0 = 3.84e10
a_Earth = 1.49e13
lod = 86164
I = 0.3299*M_Earth*R_Earth**2
# +
# Earth Orbital Angular Momentum
L_Earth0 = (M_Earth)*((G)*((M_Sun) + (M_Earth))*(a_Earth))**0.5
L_Earth0
# +
# Earth Angular Momentum
S_Earth0 = I*(2*np.pi/lod)
S_Earth0
# +
# Lunar Orbital Angular Momentum
L_moon0 = (m_moon)*((G)*((M_Earth) + (m_moon))*(a_moon0))**0.5
L_moon0
# -
# ## 2.
#
# Here, we will give the present day values of $T_{moon}$ and $T_{\odot}$.
#
#
# There is a simple tidal model which computes the Lundar tidal torque:
#
#
# $T_{moon}$ = $\frac{3}{2} \frac{G m^{2}_{moon}}{a_{moon}}(\frac{R_{\oplus}}{a_{moon}})^{5} \frac{k_{2}}{Q_{moon}}$
k_2 = 0.298
Q_moon = 11.5
# +
T_moon0 = (3/2)*(G*m_moon**2/a_moon0)*(R_Earth/a_moon0)**5*(k_2/Q_moon)
T_moon0
# -
# $T_{moon}$ and $T_{\odot}$ are related through the ratio $\beta$
#
# $\frac{T_{\odot}}{T_{moon}}$ = $\beta$
#
# So that $T_{\odot}$ = $\beta T_{moon}$
# +
B = 1/4.7
T_Sun0 = B*T_moon0
T_Sun0
# -
# Where both values have units of $\frac{cm^{4}}{gs^{2}}$
# ## 3.
#
# Here, we'll calculate the three timescales associated with equations (1) through (3) in the project outline.
#
# $\tau_{L_{\oplus}}$ = $\frac{L_{\oplus}}{T_{\odot}}$
#
#
# $\tau_{S_{\oplus}}$ = $\frac{S_{\oplus}}{T_{\odot} + T_{moon}}$
#
# $\tau_{L_{moon}}$ = $\frac{L_{moon}}{T_{moon}}$
#
# The timescale $\tau_{L_{\oplus}}$ in years:
# +
# First in seconds
tau_L_S = L_Earth0/T_Sun0
# Then converted to years
year_tau_L_S = tau_L_S / 60 / 60 / 24 / 365.25
year_tau_L_S
# -
# Then the timescale $\tau_{S_{\oplus}}$ in years:
# +
# First in seconds
tau_S_S = S_Earth0/(T_Sun0 + T_moon0)
# Then converted to years
year_tau_S_S = tau_S_S / 60 / 60 / 24 / 365.25
year_tau_S_S
# -
# And last, the timescale $\tau_{L_{moon}}$:
# +
tau_L_m = L_moon0/T_moon0
# Then converted to years
year_tau_L_m = tau_L_m / 60 / 60 / 24 / 365.25
year_tau_L_m
# -
# ## 4.
#
#
# In this problem, we are tasked with writing a function to evaluate the right hand side of the differential equations we've been provided.
#
# Prior to that, we need to define auxiliary functions for the lunar and solar torque.
# +
# bringing back the constants from earlier
G = 6.67e-8
M_Earth = 5.97e27
M_Sun = 1.98e33
m_moon = 7.349e25
R_Earth = 6.371e8
a_moon0 = 3.84e10
a_Earth = 1.49e13
lod = 86164
I = 0.3299
k_2 = 0.298
Q_moon = 11.5
L_Earth0 = (M_Earth)*((G)*((M_Sun) + (M_Earth))*(a_Earth))**0.5
S_Earth0 = I*M_Earth*R_Earth**2*(2*np.pi/lod)
L_moon0 = (m_moon)*((G)*((M_Earth) + (m_moon))*(a_moon0))**0.5
# First defining the functions we need to feed in our evolution equation
# Lunar torque
def T_moon(a_moon):
T_m = (3/2) * ((G*m_moon**2)/a_moon) * (R_Earth/a_moon)**5 * (k_2/Q_moon)
return T_m
# Solar torque
def T_Sun(a_moon):
T_S = (1/4.7) * (a_moon/a_moon0)**6 * T_moon(a_moon)
return T_S
# Write out the system of equations
def evolution(t, X):
L_Earth, S_Earth, L_moon = X
a_moon = (L_moon/m_moon)**2 / (G*(M_Earth+m_moon))
f = [T_Sun(a_moon), -T_Sun(a_moon) - T_moon(a_moon) , T_moon(a_moon)]
return f
# -
# ## 5.
#
# From the step above, we're now going to integrate and store our solution.
#
# We're integrating backwards in time, until the Moon hits Earth.
#
#
# +
# Choose solver
solver = ode(evolution)
solver.set_integrator('dopri5')
# Set the initial conditions
t0 = 0.0
X0 = [L_Earth0, S_Earth0, L_moon0]
# These intial conditions were provided in the assignment and calculated earlier
solver.set_initial_value(X0, t0)
# Integrating back in time
t1 = -5.0e16 # seconds
# Step size (larger values presented issues)
N = 250
t = np.linspace(t0, t1, N)
sol = np.empty((N, 3))
# Create an empty array to store the solution
sol[0] = X0
# Setting up our counter
i = 1
while solver.successful() and solver.t > t1:
solver.integrate(t[i])
sol[i] = solver.y
i += 1
# Defining the semimajor axis equation in kilometers
a_moon_km = (sol[:,2]/m_moon)**2 / (G*(M_Earth + m_moon)) / 100000
# Then to find when the Earth and Moon are in contact
np.where(a_moon_km == 0)[0][0]
# +
# First converting to billions of years
t_years = t / 60 / 60 / 24 / 365.25 / 1e9
# Then finding the time when the Moon and Earth are in contact
# using the index we found
t_years[243]
# -
# Thus, the Moon formed about 1.546 billion years ago, according to this model.
# ## 6.
#
# Here, we're asked to plot the function of the Moon's semimajor axis over time. The solution from above is used.
# +
# Plotting our solution
plt.plot(t_years, a_moon_km , label='Semimajor axis', color = 'limegreen')
plt.grid(b=True, color='DarkTurquoise', alpha=0.2, linestyle=':', linewidth=2)
plt.xlabel('Age [Billions of Years]')
plt.ylabel('Semimajor Axis [km]')
plt.title('Evolution of the Moons Semimajor Axis from Collision')
plt.xlim([t_years[243], 0]) # Limiting from the time of contact and the present
plt.savefig("Q6.pdf", dpi=300, bbox_inches='tight')
plt.show()
# -
# ## 7.
#
# Here, we plot the length of a day versus age.
#
# In order to obtain the equation used, we rearrange equation 7 and contrast present and past values.
# +
# Plotting our solution
lod_then = (86164 / 60 / 60)*S_Earth0/(sol[:,1])
plt.plot(t_years, lod_then, label='Semimajor axis', color = 'tomato')
plt.grid(b=True, color='DarkTurquoise', alpha=0.2, linestyle=':', linewidth=2)
plt.xlabel('Age [Billions of Years]')
plt.ylabel('Length of Day [Hours]')
plt.title('Change in the Length of Day')
plt.xlim([t_years[243], 0])
plt.ylim([0,30])
plt.savefig("Q7.pdf", dpi=300, bbox_inches='tight')
plt.show()
# -
# ## 8.
#
# Assuming the moon is a rigid object, it is reported to have a Roche limit of 9496km.
#
# Value obtained from: https://www.cs.mcgill.ca/~rwest/wikispeedia/wpcd/wp/r/Roche_limit.htm
#
# We want to find the length of the day at the time of the Moon's formation if it formed from this radius.
#
# In order to do this, we need to find the index at which we have this radius.
np.where(a_moon_km <= 9496)[0][0]
lod_then[242]
# Thus, the length of day was slighly more than 4 hours at the Roche radius.
# ## 9.
# The age of the moon has been reported to be approximately 4.425 billion years (via: https://www.cnn.com/2020/07/16/world/moon-85-million-years-younger-scn-trnd/index.html) whereas the Earth is believed to be approximately 4.54 billion years old (via: https://www.nationalgeographic.org/topics/resource-library-age-earth/?q=&page=1&per_page=25)
#
#
# These values are vastly different than those dictated by the tidal equations in the project.
# ## 10.
#
# Here, we've been asked to discuss possible errors in our model.
#
#
# Issues surrounding this model arise from the incorrect age we get for our system. I suspect that there may be issues with how our model predicts the evolution of this system, and possibly doesn't account for the conditions at impact.
#
# Firstly, I do not believe that this model allows for the moon to go through the process of mass accretion. This process would change the dynamics of the system over time as the gravitational effects change.
#
# Second, I believe that this model assumes that the Earth-Moon system has been tidally locked from impact. This is an extension of the point made above; the dynamics of this system age the Earth-Moon system based on the tidal evolution. However, if the moon spent a period of time in asynchronous rotation, this model doesn't capture this.
#
# I believe that this model interprets the collision as formation, which isn't necessarily true. The moon would have began it's formation at some distance away (i.e possibly the Roche radius) versus at impact.
#
# Additional inconsistencies may be caused by changes in the eccentricity and alignment of the moons orbit, as well as external gravitational influences. Stabilization and tidal-locking may also result in changes in the energy in this system, which could have influenced the evolution of our system.
#
# These reasons are listed in the order of what I assume to be the most reasonable assumption. My primary concern is that the initial dynamics of the system differ than those observed today, and they are not accounted for in the scope of this project.
| Final Project/CTA200 Final Project (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7
# language: python
# name: python3
# ---
# + [markdown] id="3c649d447b514f1e92bbdc7c50752de6"
# # Train a movie recommendation engine with Watson Machine Learning Accelerator
# + [markdown] id="67d8bbffa4864c65896e95dbef1cd0e4"
# ### Notebook created by <NAME>, <NAME> in Jan 2021
#
# ### In this notebook, you will learn how to use the Watson Machine Learning Accelerator (WML-A) API and accelerate the training of a movie recommendation model on GPU with Watson Machine Learning Accelerator.
#
# This notebook uses Tensorflow to build a movie recommendation engine. The model will be trained both on CPU and GPU to demonstrate that training models on GPU hardware deliver faster result times.
#
#
# This notebook covers the following sections:
#
# 1. [Setup movie recommendation model using Tensorflow](#rbm-model)<br>
#
# 1. [Training the model on CPU](#cpu)<br>
#
# 1. [Training the model on GPU with Watson Machine Learning Accelerator](#gpu)<br>
# + [markdown] id="5b848a28b78641549e59ad6abbc2b8e8"
# <a id = "rbm-model"></a>
# ## Step 1 : Setup movie recommendation model using Tensorflow
#
# + [markdown] id="103349384f9842bf87b80713aa103950"
# #### Prepare directory and file for writing movie recommendation engine.
# + id="9cf161dd-d7bc-438c-a5db-3a0591c23f19"
from pathlib import Path
model_dir = f'/project_data/data_asset/samaya'
model_main = f'main_2.py'
Path(model_dir).mkdir(exist_ok=True)
# + [markdown] id="a4d2671bc4044dea886919c664afff84"
# #### Tensorflow code to build and train a restricted Boltzmann machine using collaborative filtering. The details of how the model is built can be found in this [tutorial](https://developer.ibm.com/technologies/deep-learning/tutorials/build-a-recommendation-engine-with-a-restricted-boltzmann-machine-using-tensorflow/).
# + id="00f369b2-569f-46e2-a077-d456bcd94c3f"
# %%writefile {model_dir}/{model_main}
import sys
import subprocess
import os
import datetime
import numpy as np
import pandas as pd
import pyarrow as pa
print('pyarrow version :', pa.__version__)
path = os.path.abspath( pa.__file__)
print("pyarrow path : ", path)
import tensorflow as tf
print('Tensorflow version : ',tf.__version__)
import argparse
import os
from pathlib import PurePath
from tensorflow.python.keras import backend as K
from tensorflow.python.client import device_lib
print('List of available devices : ',device_lib.list_local_devices())
def _get_available_devices():
return [x.name for x in K.get_session().list_devices()]
def _normalize_device_name(name):
name = '/' + ':'.join(name.lower().replace('/', '').split(':')[-2:])
return name
def download_data(use_cuda):
import wget, os
from zipfile import ZipFile,ZipInfo
zip_file = 'ml-latest-small.zip'
url = 'https://github.com/IBM/wmla-assets/raw/master/dli-learning-path/movie-recommendation-use-case/dataset/'+zip_file
CPU_PATH='/project_data/data_asset/'
GPU_PATH='/gpfs/mydatafs/'
if(use_cuda):
DOWNLOAD_PATH = GPU_PATH
else :
DOWNLOAD_PATH = CPU_PATH
wget.download(url, out=DOWNLOAD_PATH)
path_to_zip_file = os.path.join(DOWNLOAD_PATH, zip_file)
with ZipFile(path_to_zip_file, 'r') as zip_ref:
zip_ref.extractall(DOWNLOAD_PATH)
os.remove(path_to_zip_file)
size = len(path_to_zip_file)
return path_to_zip_file[:size - 4]
def load_data(use_cuda):
DATA_PATH = download_data(use_cuda)
print('data path :', DATA_PATH)
MOVIE_PATH = DATA_PATH +'/movies.csv'
RATINGS_PATH = DATA_PATH +'/ratings.csv'
print('movie path :', MOVIE_PATH)
print('ratings path :', RATINGS_PATH )
if use_cuda:
print('load_data GPU')
# support multiple gpu
available_devices = _get_available_devices()
available_devices = [_normalize_device_name(name)
for name in available_devices]
print('available devices : ',available_devices)
gpu_names = [x for x in available_devices if '/gpu:' in x]
num_gpus = len(gpu_names)
print('gpu names = ',gpu_names)
print("Let's use gpus: " + str(gpu_names))
if num_gpus <= 0:
raise ValueError('Unable to find any gpu device ')
import cudf
print('cudf version : ',cudf.__version__)
#using gpu get path
movies_df = cudf.read_csv(MOVIE_PATH)
ratings_df = cudf.read_csv(RATINGS_PATH)
else :
print('load_data CPU')
movies_df = pd.read_csv(MOVIE_PATH)
ratings_df = pd.read_csv(RATINGS_PATH)
return movies_df, ratings_df
def preprocess_data(movies_df,ratings_df):
movies_df.columns = ['MovieID', 'Title', 'Genres']
movies_df = movies_df.drop('Genres',axis=1)
movies_df.head()
print('shape of movies data frame : ',movies_df.shape)
ratings_df.columns = ['UserID', 'MovieID', 'Rating', 'Timestamp']
ratings_df = ratings_df.drop('Timestamp',axis=1)
ratings_df.head()
print('shape of ratings data frame : ',ratings_df.shape)
ratings_df = ratings_df.head(200000)
# create pivot of ratings dataframe
user_rating_df = ratings_df.pivot(index='UserID', columns='MovieID')
#Remove NaNs and normalize
user_rating_df = user_rating_df.fillna(0)
norm_user_rating_df = user_rating_df / 5.0
if use_cuda:
norm_user_rating_df_pd = norm_user_rating_df.to_pandas()
trX = norm_user_rating_df_pd.values
else :
trX = norm_user_rating_df.values
return trX,len(user_rating_df.columns)
class RBM_Model(tf.Module):
def __init__(self, visibleUnits,hiddenUnits):
print('init')
self.vb = tf.Variable(tf.zeros([visibleUnits]), tf.float32) # Initialze bias to 0 for visible units(i.e. number of unique movies)
self.hb = tf.Variable(tf.zeros([hiddenUnits]), tf.float32) # Initialze bias to 0 for hidden units(i.e. numer of features we're going to learn )
self.W = tf.Variable(tf.zeros([visibleUnits, hiddenUnits]), tf.float32)
def hidden_layer(self,v0_state, W, hb):
h0_prob = tf.nn.sigmoid(tf.matmul([v0_state], W) + hb) #probabilities of the hidden units
h0_state = tf.nn.relu(tf.sign(h0_prob - tf.random.uniform(tf.shape(h0_prob)))) #sample_h_given_X
return h0_state
def reconstructed_output(self,h0_state, W, vb):
v1_prob = tf.nn.sigmoid(tf.matmul(h0_state, tf.transpose(W)) + vb)
v1_state = tf.nn.relu(tf.sign(v1_prob - tf.random.uniform(tf.shape(v1_prob)))) #sample_v_given_h
return v1_state[0]
def error(self,v0_state, v1_state):
return tf.reduce_mean(tf.square(v0_state - v1_state))
def train(self,v0_state,v1_state,h0_state,h1_state):
delta_W = tf.matmul(tf.transpose([v0_state]), h0_state) - tf.matmul(tf.transpose([v1_state]), h1_state)
self.W = self.W + alpha * delta_W
self.vb = self.vb + alpha * tf.reduce_mean(v0_state - v1_state, 0)
self.hb = self.hb + alpha * tf.reduce_mean(h0_state - h1_state, 0)
if __name__ == '__main__':
parser = argparse.ArgumentParser(description='Tensorflow Movie Recommender Example')
parser.add_argument('--batch-size', type=int, default=128, metavar='N',
help='input batch size for training (default: 128)')
parser.add_argument('--epochs', type=int, default=2, metavar='N',
help='number of epochs to train (default: 10)')
parser.add_argument('--lr', type=float, default=0.01, metavar='LR',
help='learning rate (default: 0.01)')
parser.add_argument('--no-cuda', action='store_true', default=False,
help='disables CUDA training')
args = parser.parse_args()
print(args)
use_cuda = not args.no_cuda
print('using cuda : ', use_cuda)
#### load data ####
movies_df,ratings_df = load_data(use_cuda)
#### preprocess data ####
trX,visibleUnits = preprocess_data(movies_df,ratings_df)
hiddenUnits = 20
epochs = args.epochs
batchsize = args.batch_size
errors = []
weights = []
K=1
alpha = args.lr
train_ds = tf.data.Dataset.from_tensor_slices(np.float32(trX)).batch(batchsize)
rbm = RBM_Model(visibleUnits,hiddenUnits)
_train_starttime = datetime.datetime.now()
for epoch in range(epochs):
batch_number = 0
for batch_x in train_ds:
for i_sample in range(len(batch_x)):
v0_state = batch_x[i_sample]
for k in range(K):
h0_state = rbm.hidden_layer(v0_state, rbm.W, rbm.hb)
v1_state = rbm.reconstructed_output(h0_state, rbm.W, rbm.vb)
h1_state = rbm.hidden_layer(v1_state, rbm.W, rbm.hb)
rbm.train(v0_state,v1_state,h0_state,h1_state)
v0_state = v1_state
if i_sample == len(batch_x):
err = rbm.error(batch_x[i_sample], v1_state)
errors.append(err)
weights.append(rbm.W)
print ( 'Epoch: %d' % (epoch + 1),
"batch #: %i " % batch_number, "of %i" % (len(trX)/batchsize),
"sample #: %i" % i_sample,
'reconstruction error: %f' % err)
batch_number += 1
_train_endtime = datetime.datetime.now()
print("\n exclusive Training cost: ", (_train_endtime - _train_starttime).seconds, " seconds.")
# + [markdown] id="c40e16ba6c004a0e8678737e192af2eb"
#
# ## Step 2 : Training the model on CPU
#
# #### Training was run from a Cloud Pak for Data Notebook utilizing a CPU kernel.
#
#
# In the custom environment that was created with **16vCPU** and **32GB**, it took **277 seconds** (or approximately **5 minutes**) to complete 5 EPOCH training.
# + id="b31b7adc93ea4484b56776c8c442a547"
import datetime
starttime = datetime.datetime.now()
# ! python /project_data/data_asset/samaya/main_2.py --no-cuda --epochs 5 --batch-size 10000
endtime = datetime.datetime.now()
print("Training cost: ", (endtime - starttime).seconds, " seconds.")
# + [markdown] id="103f6e2de64a468c95b36feacb60c5aa"
# <a id = "gpu"></a>
# ## Step 3 : Training the model on GPU with Watson Machine Learning Accelerator
#
# #### Prepare the model files for running on GPU:
# + id="46dcdb10044d41a88a1d007855ea159f"
import requests
from requests.packages.urllib3.exceptions import InsecureRequestWarning
requests.packages.urllib3.disable_warnings(InsecureRequestWarning)
from matplotlib import pyplot as plt
# %pylab inline
import base64
import json
import time
import urllib
# + [markdown] id="cb98cb991d8f4b8bbf2a3067e5739882"
#
# #### Configuring your environment and project details
# + [markdown] id="f116e642da6346309ce9dbcee125bcb6"
# To set up your project details, provide your credentials in this cell. You must include your cluster URL, username, and password.
# + id="23f13b0c84954e6d901d04c29ea7e284"
hostname='wmla-console-wmla.apps.cpd35-beta.cpolab.ibm.com' # please enter Watson Machine Learning Accelerator host name
login='username:password' # please enter the login and password
es = base64.b64encode(login.encode('utf-8')).decode("utf-8")
commonHeaders={'Authorization': 'Basic '+es}
req = requests.Session()
auth_url = 'https://{}/auth/v1/logon'.format(hostname)
print(auth_url)
a=requests.get(auth_url,headers=commonHeaders, verify=False)
access_token=a.json()['accessToken']
# + id="b8119a4a969c4fa98b46cceb3e9715a3"
dl_rest_url = 'https://{}/platform/rest/deeplearning/v1'.format(hostname)
commonHeaders={'accept': 'application/json', 'X-Auth-Token': access_token}
req = requests.Session()
# + id="586dd0b63f594f47ab73a212d5554e5c"
# Health check
confUrl = 'https://{}/platform/rest/deeplearning/v1/conf'.format(hostname)
r = req.get(confUrl, headers=commonHeaders, verify=False)
# + id="f6a3e859ea5949919ea8d196c49fa20b"
import tarfile
import tempfile
import os
import json
import pprint
import pandas as pd
from IPython.display import clear_output
def query_job_status(job_id,refresh_rate=3) :
execURL = dl_rest_url +'/execs/'+ job_id['id']
pp = pprint.PrettyPrinter(indent=2)
keep_running=True
res=None
while(keep_running):
res = req.get(execURL, headers=commonHeaders, verify=False)
monitoring = pd.DataFrame(res.json(), index=[0])
pd.set_option('max_colwidth', 120)
clear_output()
print("Refreshing every {} seconds".format(refresh_rate))
display(monitoring)
pp.pprint(res.json())
if(res.json()['state'] not in ['PENDING_CRD_SCHEDULER', 'SUBMITTED','RUNNING']) :
keep_running=False
time.sleep(refresh_rate)
return res
files = {'file': open("/project_data/data_asset/samaya/main_2.py", 'rb')}
args = '--exec-start tensorflow --cs-datastore-meta type=fs \
--workerDeviceNum 1 \
--conda-env-name rapids-0.18-movie-recommendation \
--model-main main_2.py --epochs 5 --batch-size 10000'
# -
# In the conda environment that was created, it took 5 seconds to complete 5 EPOCH training.
# + id="3bf3183bbfee434980bb0c22dea80088"
starttime = datetime.datetime.now()
r = requests.post(dl_rest_url+'/execs?args='+args, files=files,
headers=commonHeaders, verify=False)
if not r.ok:
print('submit job failed: code=%s, %s'%(r.status_code, r.content))
job_status = query_job_status(r.json(),refresh_rate=5)
endtime = datetime.datetime.now()
print("\nTraining cost: ", (endtime - starttime).seconds, " seconds.")
# + [markdown] id="de52ede930434b3280ab2c65cc276aaa"
# #####
| dli-learning-path/movie-recommendation-use-case/notebook/Movie-Recommendation-Engine-CPUvsGPU-with-results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import csv
import matplotlib.pyplot as plt
# %matplotlib inline
x=[]
y=[]
df = pd.read_csv('/home/rimali2009/FITTNUSS-forward_training_DNN/jk_e_Rw.csv', delimiter=',')
df.head()
plt.plot(df["Ranges of roi"],df["jk_e"])
plt.xlabel('Ranges of ROI')
plt.ylabel('JK_error')
plt.legend(loc="upper right")
plt.savefig('/home/rimali2009/FITTNUSS-forward_training_DNN/jk_e_Rw.eps')
plt.savefig('/home/rimali2009/FITTNUSS-forward_training_DNN/jk_e_Rw.png')
plt.show()
import numpy as np
import pandas as pd
import csv
import matplotlib.pyplot as plt
# %matplotlib inline
x=[]
y=[]
df = pd.read_csv('/home/rimali2009/FITTNUSS-forward_training_DNN/jk_e_U.csv', delimiter=',')
df.head()
plt.plot(df["Ranges of roi"],df["jk_e"])
plt.xlabel('Ranges of ROI')
plt.ylabel('JK_error')
plt.legend(loc="upper right")
plt.savefig('/home/rimali2009/FITTNUSS-forward_training_DNN/jk_e_U.eps')
plt.savefig('/home/rimali2009/FITTNUSS-forward_training_DNN/jk_e_U.png')
plt.show()
import numpy as np
import pandas as pd
import csv
import matplotlib.pyplot as plt
# %matplotlib inline
x=[]
y=[]
df = pd.read_csv('/home/rimali2009/FITTNUSS-forward_training_DNN/jk_e_H.csv', delimiter=',')
df.head()
plt.plot(df["Ranges of roi"],df["jk_e"])
plt.xlabel('Ranges of ROI')
plt.ylabel('JK_error')
plt.legend(loc="upper right")
plt.savefig('/home/rimali2009/FITTNUSS-forward_training_DNN/jk_e_H.eps')
plt.savefig('/home/rimali2009/FITTNUSS-forward_training_DNN/jk_e_H.png')
plt.show()
import numpy as np
import pandas as pd
import csv
import matplotlib.pyplot as plt
# %matplotlib inline
x=[]
y=[]
df = pd.read_csv('/home/rimali2009/FITTNUSS-forward_training_DNN/jk_e_C1.csv', delimiter=',')
df.head()
plt.plot(df["Ranges of roi"],df["jk_e"])
plt.xlabel('Ranges of ROI')
plt.ylabel('JK_error')
plt.legend(loc="upper right")
plt.savefig('/home/rimali2009/FITTNUSS-forward_training_DNN/jk_e_C1.eps')
plt.savefig('/home/rimali2009/FITTNUSS-forward_training_DNN/jk_e_C1.png')
import numpy as np
import pandas as pd
import csv
import matplotlib.pyplot as plt
# %matplotlib inline
x=[]
y=[]
df = pd.read_csv('/home/rimali2009/FITTNUSS-forward_training_DNN/jk_e_C6.csv', delimiter=',')
df.head()
plt.plot(df["Ranges of roi"],df["jk_e"])
plt.xlabel('Ranges of ROI')
plt.ylabel('JK_error')
plt.legend(loc="upper right")
plt.savefig('/home/rimali2009/FITTNUSS-forward_training_DNN/jk_e_C6.eps')
plt.savefig('/home/rimali2009/FITTNUSS-forward_training_DNN/jk_e_C6.png')
import numpy as np
import pandas as pd
import csv
import matplotlib.pyplot as plt
# %matplotlib inline
x=[]
y=[]
df = pd.read_csv('/home/rimali2009/FITTNUSS-forward_training_DNN/roi_2000_TC.csv', delimiter=',')
df.head()
plt.plot(df["Training data"],df["Loss"],label='Final loss value')
plt.xlabel('Number of training data')
plt.ylabel('Loss of Validation')
plt.legend(loc="upper right")
plt.savefig('/home/rimali2009/FITTNUSS-forward_training_DNN/roi2000_TC_case2.eps')
plt.savefig('/home/rimali2009/FITTNUSS-forward_training_DNN/roi2000_TC_case2.png')
plt.show()
| jupyter_notebook_files/DNN_plots_JK_Error_training_loss_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modeling
# Import modules
import os
import sys
import pickle
import seaborn as sns
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV, train_test_split, cross_val_score
from sklearn.metrics import accuracy_score
import warnings
# Unpickle clean df and target and store as X and y
# +
with open('../data/processed/df_clean.pickle', 'rb') as file:
X = pickle.load(file)
with open('../data/processed/target.pickle', 'rb') as file:
y = pickle.load(file)
# -
# Print shape of X and y
print('Predictor shape:', X.shape, '\n',
'Target shape:', y.shape, '\n')
# Split data into X and y train and test sets stratifying by the target
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=0, stratify=y)
# Create scaler object and fit to both train and test X sets
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
# Create scikit-learn K-Nearest Neighbors object and Random Forest object
knn = KNeighborsClassifier()
rf = RandomForestClassifier(n_estimators=100)
# Fit the models to the training data
knn.fit(X_train, y_train)
rf.fit(X_train, y_train)
# Use models to make predictions on training data
y_hat_knn = knn.predict(X_train)
y_hat_rf = rf.predict(X_train)
# Evaluate performance of models on training predictions and print accuracy scores
score_knn = accuracy_score(y_train, y_hat_knn)
score_rf = accuracy_score(y_train, y_hat_rf)
print('KNN test score:', round(score_knn, 3))
print('RF test score:', round(score_rf, 5))
# Create pipeline to scale then call model for knn and random forest
knn_pipe = Pipeline([('scale', StandardScaler()),
('knn', KNeighborsClassifier())
])
rf_pipe = Pipeline([('scale', StandardScaler()),
('rf', RandomForestClassifier())
])
# Cross validate training data using knn and print accuracy score
scores_knn = cross_val_score(knn_pipe, X_train, y_train, cv=5, scoring='accuracy')
print('KNN scores:', scores_knn, '\n')
# Cross validate training data using Random Forest and print accuracy score
scores_rf = cross_val_score(rf_pipe, X_train, y_train, cv=5, scoring='accuracy')
print('Random forest scores:', scores_rf)
warnings.filterwarnings('ignore')
# Print mean accuracy score from knn and random forest
print('knn cv mean:', scores_knn.mean(),'\n',
'rf cv mean:', scores_rf.mean())
# Inspect features importance
features = X.columns
importances = rf.feature_importances_
indices = np.argsort(importances)
plt.figure(figsize=[20,15])
plt.barh(range(len(indices)), importances[indices], color='lightpink')
plt.yticks(range(len(indices)), [features[i] for i in indices])
plt.xlabel('Relative Importance')
plt.title('Feature Importances')
plt.style.use('seaborn')
plt.show()
# Create dictionary of parameters to search
param_grid = {
'n_estimators': [10, 100, 250],
'max_depth': [2, 3, 5, 7],
'oob_score': [True],
'max_features': [None]
}
# Do a gridsearch on the parameter grid to find best params with cross-validation
# +
search = GridSearchCV(rf, param_grid, scoring='accuracy',
n_jobs=-1, iid=False, cv=5)
search.fit(X_train, y_train)
y_hat_search = search.predict(X_train)
score = accuracy_score(y_train, y_hat_search)
# -
# View best parameters from the gridsearch
search.best_params_
# Make predictions using test data
y_hat_search_test = search.predict(X_test)
# Print test score using rf model with best params from gridsearch
score_search = accuracy_score(y_test, y_hat_search_test)
print('RF Search test score:', round(score_search, 5))
| notebooks/2-al-modeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Inference of parameters with latent variables (SIR model)
# In this notebook, we consider the SIR model with symptomatically and asymptomatically infected. We are trying to infer the epidemiological parameters as well as control parameters from the $I_\mathrm{s}$ trajectory.
#
# For the pre-intervention period, we infer the following parameters as well as initial conditions:
# * $\alpha$ (fraction of asymptomatic infectives),
# * $\beta$ (probability of infection on contact),
# * $\gamma_{I_a}$ (rate of recovery for asymptomatic infected individuals), and
# * $\gamma_{I_s}$ (rate of recovery for symptomatic infected individuals)
#
# For the post-intervention period, we infer the following control parameters:
# * $a_\mathrm{W}$ fraction of work contact,
# * $a_\mathrm{S}$ fraction of social contact,
# * $a_\mathrm{O}$ fraction of other contacts
# %%capture
## compile PyRoss for this notebook
import os
owd = os.getcwd()
os.chdir('../../')
# %run setup.py install
os.chdir(owd)
# %matplotlib inline
import numpy as np
import pyross
import pandas as pd
import time
import matplotlib.pyplot as plt
from scipy.io import loadmat
# ## 1) Generate a trajectory
# We generate a test trajectory on a population with four ages groups using UK contact matrix
# +
## population and age classes
M=4 ## number of age classes
my_data = np.genfromtxt('../data/age_structures/UK.csv', delimiter=',', skip_header=1)
aM, aF = my_data[:, 1], my_data[:, 2]
Ni0=aM+aF; Ni=np.zeros((M))
# scale the population down to a more manageble level
Ni[0] = (np.sum(Ni0[0:4])).astype('int')
Ni[1] = (np.sum(Ni0[4:8])).astype('int')
Ni[2] = (np.sum(Ni0[8:12])).astype('int')
Ni[3] = (np.sum(Ni0[12:16])).astype('int')
N=np.sum(Ni)
fi = Ni/N
# Get individual contact matrices
CH0, CW0, CS0, CO0 = pyross.contactMatrix.UK()
CH = np.zeros((M, M))
CW = np.zeros((M, M))
CS = np.zeros((M, M))
CO = np.zeros((M, M))
for i in range(16):
CH0[i,:] = CH0[i,:]*Ni0[i]
CW0[i,:] = CW0[i,:]*Ni0[i]
CS0[i,:] = CS0[i,:]*Ni0[i]
CO0[i,:] = CO0[i,:]*Ni0[i]
for i in range(M):
for j in range(M):
i1, j1 = i*4, j*4
CH[i,j] = np.sum( CH0[i1:i1+4, j1:j1+4] )/Ni[i]
CW[i,j] = np.sum( CW0[i1:i1+4, j1:j1+4] )/Ni[i]
CS[i,j] = np.sum( CS0[i1:i1+4, j1:j1+4] )/Ni[i]
CO[i,j] = np.sum( CO0[i1:i1+4, j1:j1+4] )/Ni[i]
Ni = (Ni/5e2).astype('int') # Scale the numbers to avoid numerical problems
N = np.sum(Ni)
# +
# Generate class with contact matrix for SIR model with UK contact structure
generator = pyross.contactMatrix.SIR(CH, CW, CS, CO)
T_intervention = 20
times= [T_intervention] # temporal boundaries between different contact-behaviour
aW, aS, aO = 0.5, 0.5, 0.4
# prefactors for CW, CS, CO:
interventions = [[1.0,1.0,1.0], # before first time
[aW, aS, aO], # after first time
]
# generate corresponding contact matrix function
C = generator.interventions_temporal(times=times,interventions=interventions)
# +
beta = 0.04 # contact rate parameter
gIa = 1./7 # recovery rate of asymptomatic infectives
gIs = 1./7 # recovery rate of symptomatic infectives
alpha = 0.2 # asymptomatic fraction
fsa = 0.8 # suppresion of contact by symptomatics
# initial conditions
Is0 = np.ones(M)*10
Ia0 = np.ones((M))*10
R0 = np.zeros((M))
S0 = Ni - (Ia0 + Is0 + R0)
parameters = {'alpha':alpha,'beta':beta, 'gIa':gIa,'gIs':gIs,'fsa':fsa}
model = pyross.stochastic.SIR(parameters, M, Ni)
# start simulation
Tf=100; Nf=Tf+1
data=model.simulate(S0, Ia0, Is0, C, Tf, Nf)
np.save('cal_lat_SIR_traj.npy', data['X'])
# -
# plot the number of infected people
IC = np.zeros((Nf))
for i in range(M):
IC += data['X'][:,2*M+i]
t = data['t']
plt.plot(t, IC)
plt.axvspan(0, T_intervention,
label='Pre intervention',
alpha=0.3, color='dodgerblue')
plt.xlim([0, Tf])
plt.show()
# ## 2) Infer the parameters and initial conditions for the pre-intervention trajectory
# +
Tf_initial = T_intervention # truncate to only getting the pre-intervention trajectory
Nf_initial = Tf_initial+1
x = (np.load('cal_lat_SIR_traj.npy')).astype('float')
x = (x/N)[:Nf_initial]
steps = 101 # number internal integration steps taken
fltr=np.repeat(np.array([False, False, True]), M)
obs=x[:, fltr]
# First, check the deterministic simulation against stochstic simulations with the same parameters and initial conditons
# They are likely to be different due to the inherent stochasticity of the model
contactMatrix = generator.constant_contactMatrix()
det_model = pyross.deterministic.SIR(parameters, int(M), fi)
estimator = pyross.inference.SIR(parameters, M, fi, int(N), steps)
xm = estimator.integrate(x[0], 0, Tf_initial, Nf_initial, det_model, contactMatrix)
t = np.linspace(0, Tf_initial, Nf_initial)
plt.plot(t, np.sum(x[:,2*M:3*M], axis=1))
plt.plot(t, np.sum(xm[:,2*M:3*M], axis=1))
plt.show()
# +
# Initialise the estimator
estimator = pyross.inference.SIR(parameters, M, fi, int(N), steps)
# Compute -log_p for the original (correct) parameters
# This gives an estimate of the final -logp we expect from the optimisation scheme
start_time = time.time()
parameters = {'alpha':alpha, 'beta':beta, 'gIa':gIa, 'gIs':gIs,'fsa':fsa}
logp = estimator.minus_logp_red(parameters, x[0], obs[1:], fltr, Tf_initial, Nf_initial, contactMatrix)
end_time = time.time()
print(logp)
print(end_time - start_time)
# +
# make parameter guesses and set up bounds for each parameter
eps = 1e-3
alpha_g = 0.15
alpha_std = 0.1
alpha_bounds = (eps, 1-2*eps)
# the bound for alpha must be 1-2*eps to avoid alpha>1 in hessian calculation performed by optimizer
beta_g = 0.1
beta_std = 0.1
beta_bounds = (eps, 1)
gIa_g = 0.13
gIa_std = 0.05 # small stds for rate of recovery (can obtain from clinical data)
gIa_bounds = (0.1, 0.3)
gIs_g = 0.13
gIs_std = 0.05 # small stds for rate of recovery (can obtain from clinical data)
gIs_bounds = (0.1, 0.3)
# not inferred
fsa_g = 0.8
Ia0_g = (Ia0-3)/N
Ia_std = Ia0_g*2
bounds_for_Ia = np.tile([0.1/N, 100/N], M).reshape(M, 2)
Is0_g = (Is0-3)/N
Is_std = Is0_g*0.5
bounds_for_Is = np.tile([0.1/N, 100/N], M).reshape(M, 2)
S0_g = (S0+10)/N
S_std = Ia_std*2
bounds_for_S = np.tile([0.1/N, 1], M).reshape(M, 2)
# Optimisation parameters
ftol = 1e-5 # the relative tol in (-logp)
# Set up bounds, guess and stds for the rest of the params
bounds = np.array([alpha_bounds, beta_bounds, gIa_bounds, gIs_bounds,
*bounds_for_S, *bounds_for_Ia, *bounds_for_Is])
guess = np.array([alpha_g, beta_g, gIa_g, gIs_g, *S0_g, *Ia0_g, *Is0_g])
stds = np.array([alpha_std, beta_std, gIa_std, gIs_std, *S_std, *Ia_std, *Is_std])
# Run the optimiser
start_time = time.time()
params = estimator.latent_inference(guess, stds, obs, fltr, Tf_initial, Nf_initial,
generator.constant_CM, bounds,
global_max_iter=15, global_ftol_factor=1e3,
verbose=True, ftol=ftol)
end_time = time.time()
print(end_time - start_time)
# -
# save the parameters
np.save('SIR_cal_lat_param.npy', params)
# +
# print the correct params and best estimates
print('True parameters')
parameters = {'alpha':alpha, 'beta':beta, 'gIa':gIa, 'gIs':gIs,'fsa':fsa}
print(parameters)
params = np.load('SIR_cal_lat_param.npy')
best_estimates = estimator.make_params_dict(params)
print('\n Best estimates')
print(best_estimates)
print('\n True initial conditions: ')
print(S0, Ia0, Is0)
print('\n Inferred initial conditons: ')
print((params[4:]*N).astype('int'))
# -
# ### Check the inferred trajectory against the true trajectory for pre-intervention
# +
x = np.load('cal_lat_SIR_traj.npy')/N
x0 = params[4:]
# set params for estimate
estimator.set_params(best_estimates)
# make det_model and contactMatrix
det_model = pyross.deterministic.SIR(best_estimates, M, fi)
contactMatrix = generator.constant_contactMatrix()
# generate the deterministic trajectory
x_det = estimator.integrate(x0, 0, Tf_initial, Nf_initial, det_model, contactMatrix)
plt.plot(np.sum(x[:Nf_initial,M:2*M], axis=1), label='True Ia')
plt.plot(np.sum(x_det[:, M:2*M], axis=1), label='Inferred Ia')
plt.plot(np.sum(x[:Nf_initial,2*M:3*M], axis=1), label='True Is')
plt.plot(np.sum(x_det[:, 2*M:3*M], axis=1), label='Inferred Is')
plt.legend()
plt.show()
# -
# Quite good at inferring Is (which it can see), not so good at inferring Ia (which it cannot see)
# ## 3) Infer the control parameters
# We infer the control parameters, assuming that the previously inferred initial conditions and epidemiological parameters
# +
# Test the -logp for the correct control parameters
# get the trajectory
x = np.load('cal_lat_SIR_traj.npy')/N
x = x[Nf_initial:]
obs = x[:, fltr]
Nf_control = x.shape[0]
Tf_control = Nf_control-1
# Set the initial condition for post-intervention
x0_control = np.array(x_det[-1])
# get the contact matrix for post-intervention
times = [Tf+1]
interventions = [[aW, aS, aO]]
contactMatrix = generator.interventions_temporal(times=times,interventions=interventions)
# calculate minus_log_p for the correct control parameters
logp = estimator.minus_logp_red(best_estimates, x0, obs[1:], fltr, Tf_control, Nf_control, contactMatrix)
print(logp)
# +
# set up initial guess and bounds
aW_g = 0.5
aW_std = 0.1
aW_bounds = [0.2, 0.8]
aS_g = 0.55
aS_std = 0.1
aS_bounds = [0.2, 0.8]
aO_g = 0.45
aO_std = 0.1
aO_bounds = [0.2, 0.8]
guess = np.array([aW_g, aS_g, aO_g])
stds = np.array([aW_std, aS_std, aO_std])
bounds = np.array([aW_bounds, aS_bounds, aO_bounds])
ftol = 1e-4
start_time = time.time()
estimator.set_params(best_estimates)
control_params = estimator.latent_infer_control(guess, stds, x0_control, obs, fltr, Tf_control, Nf_control,
generator, bounds,
global_max_iter=5, global_ftol_factor=1e4,
verbose=True,
ftol=ftol)
end_time = time.time()
print(control_params) # best guess
print(end_time - start_time)
# +
times = [T_intervention]
interventions = [[1.0,1.0,1.0], # before first time
control_params, # after first time
]
# compare the true trajectory with the
contactMatrix = generator.interventions_temporal(times=times,interventions=interventions)
det_model = pyross.deterministic.SIR(parameters, M, fi)
x0 = params[len(parameters)-1:]
x_det_control = estimator.integrate(x0, 0, Tf, Nf, det_model, contactMatrix)
x = np.load('cal_lat_SIR_traj.npy')/N
plt.plot(np.sum(x[:,M:2*M], axis=1), label='Ia')
plt.plot(np.sum(x_det_control[:, M:2*M], axis=1), label='Inferred Ia')
plt.plot(np.sum(x[:,2*M:3*M], axis=1), label='Is')
plt.plot(np.sum(x_det_control[:, 2*M:3*M], axis=1), label='Inferred Is')
plt.axvspan(0, T_intervention,
label='Pre intervention',
alpha=0.3, color='dodgerblue')
plt.xlim([0, Tf])
plt.legend()
plt.show()
| examples/inference/ex13_calibration_latent_SIR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# https://github.com/shafaua/visionary
#
# https://cloud.google.com/vision/
#
# https://cloud.google.com/vision/docs/auth
import json
from visionary import GoogleCloudVision, LabelDetection, LogoDetection
def load_api_secrets(filename):
"""Convenience to load the API key from a json file."""
try:
with open(filename, 'r') as fp:
api_params = json.load(fp)
except Exception as e:
print('Failed to load API secrets key: {}'.format(e))
api_params = None
return api_params['key']
API_key_filepath = '/Users/robincole/Desktop/Google_API_key.json'
API_key = load_api_secrets(API_key_filepath)
#API_key
client = GoogleCloudVision(API_key)
# ls
response = client.annotate(open("apple.jpg"), LogoDetection())
response = client.annotate("http://google.com/dummy.jpg")
| Google Vision/resources/Visionary 25-10-2017.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Q#
# language: qsharp
# name: iqsharp
# ---
# # Distinguish Unitaries Kata Workbook
#
# **What is this workbook?**
# A workbook is a collection of problems, accompanied by solutions to them.
# The explanations focus on the logical steps required to solve a problem; they illustrate the concepts that need to be applied to come up with a solution to the problem, explaining the mathematical steps required.
#
# Note that a workbook should not be the primary source of knowledge on the subject matter; it assumes that you've already read a tutorial or a textbook and that you are now seeking to improve your problem-solving skills. You should attempt solving the tasks of the respective kata first, and turn to the workbook only if stuck. While a textbook emphasizes knowledge acquisition, a workbook emphasizes skill acquisition.
#
# This workbook describes the solutions to the problems offered in the [Distinguish Unitaries kata](./DistinguishUnitaries.ipynb). Since the tasks are offered as programming problems, the explanations also cover some elements of Q# that might be non-obvious for a first-time user.
#
# **What you should know for this workbook**
#
# You should be familiar with the following concepts before tackling the Distinguish Unitaries kata (and this workbook):
#
# 1. [Basic linear algebra](./../tutorials/LinearAlgebra/LinearAlgebra.ipynb)).
# 2. [The concept of qubit](./../tutorials/Qubit/Qubit.ipynb) and [multi-qubit systems](./../tutorials/MultiQubitSystems/MultiQubitSystems.ipynb).
# 3. [Single-qubit](./../tutorials/SingleQubitGates/SingleQubitGates.ipynb) and [multi-qubit quantum gates](./../tutorials/MultiQubitGates/MultiQubitGates.ipynb) and using them to manipulate the state of the system.
# 4. Measurements and using them to distinguish quantum states.
# To begin, first prepare this notebook for execution (if you skip the first step, you'll get "Syntax does not match any known patterns" error when you try to execute Q# code in the next cells; if you skip the second step, you'll get "Invalid test name" error):
%package Microsoft.Quantum.Katas::0.12.20072031
%workspace reload
# ## Part I. Single-Qubit Gates
# ### Task 1.1. Identity or Pauli X?
#
# **Input:** An operation that implements a single-qubit unitary transformation:
# either the identity (**I** gate)
# or the Pauli X gate (**X** gate).
# The operation will have Adjoint and Controlled variants defined.
#
# **Output:** 0 if the given operation is the **I** gate, 1 if the given operation is the **X** gate.
#
# You are allowed to apply the given operation and its adjoint/controlled variants exactly **once**.
# ### Solution
#
# The only way to extract information out of a quantum system is measurement.
# Measurements give us information about the states of a system, so to get information about the gate, we need to find a way to convert it into information about a state.
# If we want to distinguish two gates, we need to figure out to prepare a state and perform a measurement on it that will give us a result that we can interpret.
# To do this, we’ll need to find a qubit state that, by applying to it I gate or X gate, will be transformed into states that can be distinguished using measurement, i.e., orthogonal states.
# Let's find such state.
#
# > As a reminder, here are the matrices that correspond to the given gates:
# > $$I = \begin{bmatrix} 1 & 0 \\ 0 & 1 \end{bmatrix}, X = \begin{bmatrix} 0 & 1 \\ 1 & 0 \end{bmatrix}$$
#
# Consider the effects of these gates on the basis states:
#
# $$I|0\rangle = |0\rangle, I|1\rangle = |1\rangle$$
# $$X|0\rangle = |1\rangle, X|1\rangle = |0\rangle$$
#
# We see that the **I** gate leaves the $|0\rangle$ state unchanged, and the **X** gate transforms it into the $|1\rangle$ state.
# So the easiest thing to do is to prepare a $|0\rangle$ state, apply the given unitary to it and measure the resulting state in the computational basis:
# * If the measurement result is `Zero`, the measured state was $|0\rangle$, and we know the unitary applied to it was the **I** gate.
# * If the measurement result is `One`, the measured state was $|1\rangle$, and we know the unitary applied to it was the **X** gate.
#
# > In Q#, the freshly allocated qubits start in the $|0\rangle$ state, so you don't need to do anything to prepare the necessary state before applying the unitary to it.
# > You also have to return the qubits you allocated to the $|0\rangle$ state before releasing them.
# > You can do that by measuring the qubit using the `M` operation and applying the **X** gate if it was measured in the $|1\rangle$ state, or you can use [`MResetZ`](https://docs.microsoft.com/en-us/qsharp/api/qsharp/microsoft.quantum.measurement.mresetz) operation that wraps this measurement and fixup into one operation.
# +
%kata T11_DistinguishIfromX_Test
open Microsoft.Quantum.Measurement;
operation DistinguishIfromX (unitary : (Qubit => Unit is Adj+Ctl)) : Int {
using (q = Qubit()) {
unitary(q);
return MResetZ(q) == Zero ? 0 | 1;
}
}
# -
# [Return to task 1.1 of the Distinguish Unitaries kata.](./DistinguishUnitaries.ipynb#Task-1.1.-Identity-or-Pauli-X?)
# ### Task 1.2. Identity or Pauli Z?
#
# **Input:** An operation that implements a single-qubit unitary transformation:
# either the identity (**I** gate)
# or the Pauli Z gate (**Z** gate).
# The operation will have Adjoint and Controlled variants defined.
#
# **Output:** 0 if the given operation is the **I** gate, 1 if the given operation is the **Z** gate.
#
# You are allowed to apply the given operation and its adjoint/controlled variants exactly **once**.
# ### Solution
#
# > As a reminder, $Z = \begin{bmatrix} 1 & 0 \\ 0 & -1 \end{bmatrix}$
#
# We won't be able to distinguish **I** from **Z** by applying them to the basis states, since they both leave the $|0\rangle$ state unchanged and add a phase to the $|1\rangle$ state:
#
# $$I|0\rangle = |0\rangle, I|1\rangle = |1\rangle$$
# $$Z|0\rangle = |0\rangle, Z|1\rangle = -|1\rangle$$
#
# However, if we try applying these gates to a superposition of basis states, we'll start seeing a difference between the resulting states:
#
# $$I \big(\frac{1}{\sqrt2}(|0\rangle + |1\rangle)\big) = \frac{1}{\sqrt2}(|0\rangle + |1\rangle)$$
# $$Z \big(\frac{1}{\sqrt2}(|0\rangle + |1\rangle)\big) = \frac{1}{\sqrt2}(|0\rangle \color{blue}{-} |1\rangle)$$
#
# These two states are orthogonal and can be distinguished by measuring them in the $\{ |+\rangle, |-\rangle\}$ basis using [`MResetX`](https://docs.microsoft.com/en-us/qsharp/api/qsharp/microsoft.quantum.measurement.mresetx) operation (which is equivalent to applying an **H** gate and measuring in the computational basis).
#
# > The task of distinguishing these two states is covered in more detail in the [Measurements kata](./..//Measurements/Measurements.ipynb#Task-1.3.-$|+\rangle$-or-$|-\rangle$?) and the corresponding workbook.
# +
%kata T12_DistinguishIfromZ_Test
open Microsoft.Quantum.Measurement;
operation DistinguishIfromZ (unitary : (Qubit => Unit is Adj+Ctl)) : Int {
using (q = Qubit()) {
H(q);
unitary(q);
return MResetX(q) == Zero ? 0 | 1;
}
}
# -
# [Return to task 1.2 of the Distinguish Unitaries kata.](./DistinguishUnitaries.ipynb#Task-1.2.-Identity-or-Pauli-Z?)
# ### Task 1.3. Z or S?
#
# **Input:** An operation that implements a single-qubit unitary transformation:
# either the **Z** gate
# or the **S** gate.
# The operation will have Adjoint and Controlled variants defined.
#
# **Output:** 0 if the given operation is the **Z** gate, 1 if the given operation is the **S** gate.
#
# You are allowed to apply the given operation and its adjoint/controlled variants at most **twice**.
# ### Solution
#
# > As a reminder, $S = \begin{bmatrix} 1 & 0 \\ 0 & i \end{bmatrix}$
#
# This task differs from the previous two in that it allows you to apply the given unitary **twice**.
# Let's treat this as a hint that it is, and check how the given gates looks when applied twice.
# If you square the corresponding matrices (which is quite simple to do for diagonal matrices), you'll get
#
# $$Z^2 = \begin{bmatrix} 1 & 0 \\ 0 & 1 \end{bmatrix} = I, S^2 = \begin{bmatrix} 1 & 0 \\ 0 & -1 \end{bmatrix} = Z$$
#
# This means that the task of identifying the *square* of the given unitary transformation is the same as distinguishing **I** from **Z** gates - and that's exactly the [task 1.2](#Task-1.2.-Identity-or-Pauli-Z?)!
# +
%kata T13_DistinguishZfromS_Test
open Microsoft.Quantum.Measurement;
operation DistinguishZfromS (unitary : (Qubit => Unit is Adj+Ctl)) : Int {
using (q = Qubit()) {
H(q);
unitary(q);
unitary(q);
return MResetX(q) == Zero ? 0 | 1;
}
}
# -
# [Return to task 1.3 of the Distinguish Unitaries kata.](./DistinguishUnitaries.ipynb#Task-1.3.-Z-or-S?)
# ### Task 1.4. Z or $-$Z?
#
# **Input:** An operation that implements a single-qubit unitary transformation:
# either the **Z** gate
# or the minus **Z** gate (i.e., the gate $- |0\rangle\langle0| + |1\rangle\langle1|$).
# The operation will have Adjoint and Controlled variants defined.
#
# **Output:** 0 if the given operation is the **Z** gate, 1 if the given operation is the $-$**Z** gate.
#
# You are allowed to apply the given operation and its adjoint/controlled variants exactly **once**.
# ### Solution
#
# This task is more interesting: the given gates differ by a global phase they introduce (i.e., each of them is a multiple of the other), and the results of applying them to any single-qubit state are going to be indistinguishable by any measurement you can devise.
#
# Fortunately, we are given not just the unitary itself, but also its controlled variant, i.e., the gate which applies the given unitary if the control qubit or qubits are in the $|1\rangle$ state, and does nothing if they are in the $|0\rangle$ state.
# This allows us to use so called "phase kickback" trick, in which applying a controlled version of a gate allows us to observe the phase introduced by this gate on the control qubit. Indeed,
#
# | State | Controlled Z | Controlled $-$Z |
# |-------|---------------|------|
# | $|00\rangle$ | $|00\rangle$ | $|00\rangle$ |
# | $|01\rangle$ | $|01\rangle$ | $|01\rangle$ |
# | $|10\rangle$ | $\color{blue}{|10\rangle}$ | $\color{blue}{-|10\rangle}$ |
# | $|11\rangle$ | $\color{blue}{-|11\rangle}$ | $\color{blue}{|11\rangle}$ |
#
# We see that both controlled gates don't modify the states with the control qubit in the $|0\rangle$ state, but if the control qubit is in the $|1\rangle$ state, they introduce a $-1$ phase to different basis states.
# We can take advantage of this if we apply the controlled gate to a state in which the *control qubit* is in superposition, such as $\frac{1}{\sqrt2}(|0\rangle + |1\rangle) \otimes |0\rangle$:
#
# $$\text{Controlled Z}\frac{1}{\sqrt2}(|0\rangle + |1\rangle) \otimes |0\rangle = \frac{1}{\sqrt2}(|0\rangle + |1\rangle) \otimes |0\rangle$$
# $$\text{Controlled }-\text{Z}\frac{1}{\sqrt2}(|0\rangle + |1\rangle) \otimes |0\rangle = \frac{1}{\sqrt2}(|0\rangle - |1\rangle) \otimes |0\rangle$$
#
# After this we can measure the first qubit to distinguish $\frac{1}{\sqrt2}(|0\rangle + |1\rangle)$ from $\frac{1}{\sqrt2}(|0\rangle - |1\rangle)$, like we did in [task 1.2](#Task-1.2.-Identity-or-Pauli-Z?).
#
# > In Q# we can express controlled version of a gate using [Controlled functor](https://docs.microsoft.com/en-us/quantum/user-guide/using-qsharp/operations-functions#controlled-functor): the first argument of the resulting gate will be an array of control qubits, and the second one - the arguments of the original gate (in this case just the target qubit).
# +
%kata T14_DistinguishZfromMinusZ_Test
open Microsoft.Quantum.Measurement;
operation DistinguishZfromMinusZ (unitary : (Qubit => Unit is Adj+Ctl)) : Int {
using (qs = Qubit[2]) {
// prep (|0⟩ + |1⟩) ⊗ |0⟩
H(qs[0]);
Controlled unitary(qs[0..0], qs[1]);
return MResetX(qs[0]) == Zero ? 0 | 1;
}
}
# -
# [Return to task 1.4 of the Distinguish Unitaries kata.](./DistinguishUnitaries.ipynb#Task-1.4.-Z-or--Z?)
# ## Part II. Multi-Qubit Gates
# ### Task 2.1. $I \otimes X$ or CNOT?
#
# **Input:** An operation that implements a single-qubit unitary transformation:
# either the $I \otimes X$ (the **X** gate applied to the second qubit)
# or the **CNOT** gate with the first qubit as control and the second qubit as target.
# * The operation will accept an array of qubits as input, but it will fail if the array is empty or has one or more than two qubits.
# * The operation will have Adjoint and Controlled variants defined.
#
# **Output:** 0 if the given operation is $I \otimes X$, 1 if the given operation is the **CNOT** gate.
#
# You are allowed to apply the given operation and its adjoint/controlled variants exactly **once**.
# ### Solution
#
# Let's consider the effect of these gates on the basis states:
#
# | State | $I \otimes X$ | **CNOT** |
# |-------|---------------|------|
# | $|00\rangle$ | $|01\rangle$ | $|00\rangle$ |
# | $|01\rangle$ | $|00\rangle$ | $|01\rangle$ |
# | $|10\rangle$ | $|11\rangle$ | $|11\rangle$ |
# | $|11\rangle$ | $|10\rangle$ | $|10\rangle$ |
#
# We can see that applying these two gates to states with the first qubit in the $|1\rangle$ state yields identical results, but applying them to states with the first qubit in the $|0\rangle$ state produces states that differ in the second qubuit.
# This makes sense, since the **CNOT** gate is defined as "apply **X** gate to the target qubit if the control qubit is in the $|1\rangle$ state, and do nothing if it is in the $|0\rangle$ state".
#
# Thus, the easiest solution is: allocate two qubits in the $|00\rangle$ state and apply the unitary to them, then measure the second qubit; if it is `One`, the gate is $I \otimes X$, otherwise it's **CNOT**.
# +
%kata T21_DistinguishIXfromCNOT_Test
open Microsoft.Quantum.Measurement;
operation DistinguishIXfromCNOT (unitary : (Qubit[] => Unit is Adj+Ctl)) : Int {
using (qs = Qubit[2]) {
unitary(qs);
return MResetZ(qs[1]) == One ? 0 | 1;
}
}
# -
# [Return to task 2.1 of the Distinguish Unitaries kata.](./DistinguishUnitaries.ipynb#Task-2.1.-$I-\otimes-X$-or-CNOT?)
# *To be continued...*
| DistinguishUnitaries/Workbook_DistinguishUnitaries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/SpectralPoet/Amazon_Vine_Analysis/blob/main/Vine_Review_Analysis.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="cOce8klccffk" outputId="98f528e1-962c-45f1-d521-fa31da776349"
import os
# Find the latest version of spark 3.0 from http://www.apache.org/dist/spark/ and enter as the spark version
# For example:
# spark_version = 'spark-3.0.3'
spark_version = 'spark-3.1.2'
os.environ['SPARK_VERSION']=spark_version
# Install Spark and Java
# !apt-get update
# !apt-get install openjdk-11-jdk-headless -qq > /dev/null
# !wget -q http://www.apache.org/dist/spark/$SPARK_VERSION/$SPARK_VERSION-bin-hadoop2.7.tgz
# !tar xf $SPARK_VERSION-bin-hadoop2.7.tgz
# !pip install -q findspark
# Set Environment Variables
import os
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-11-openjdk-amd64"
os.environ["SPARK_HOME"] = f"/content/{spark_version}-bin-hadoop2.7"
# Start a SparkSession
import findspark
findspark.init()
# + colab={"base_uri": "https://localhost:8080/"} id="A_POzswcSgNT" outputId="bb4f7d6c-42aa-4d66-bf7c-2c4fa542b7ab"
# !wget https://jdbc.postgresql.org/download/postgresql-42.2.16.jar
# + id="eHSW979BSlRt"
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName("CloudETL").config("spark.driver.extraClassPath","/content/postgresql-42.2.16.jar").getOrCreate()
# + id="Zc6hx-7kW2e-"
# Read in data from S3 Buckets
from pyspark import SparkFiles
url ="https://spectralpoet-bucket.s3.us-east-2.amazonaws.com/amazon_reviews_us_Video_Games_v1_00.tsv"
spark.sparkContext.addFile(url)
user_data_df = spark.read.csv(SparkFiles.get("amazon_reviews_us_Video_Games_v1_00.tsv"), sep="\t", header=True, inferSchema=True)
# + colab={"base_uri": "https://localhost:8080/"} id="Zy7hYssJbn6Z" outputId="fc313515-b76f-47b1-9304-15c155c16c70"
user_data_df.show()
# + id="3T096oyixcu8"
# Filter the data and create a new DataFrame or table to retrieve all the rows where the total_votes count is equal to or greater than 20
# to pick reviews that are more likely to be helpful and to avoid having division by zero errors later on.
over_20_votes_df = user_data_df.filter(user_data_df.total_votes >= 20)
# + colab={"base_uri": "https://localhost:8080/"} id="vViAl7Rc04Eo" outputId="15adfd88-cb13-4067-fe33-6c7d239cc8c6"
over_20_votes_df.show()
# + id="xaRDb_dEy6cW"
# Filter the new DataFrame or table created in Step 1 and create a new DataFrame or table to retrieve all the rows where the number of helpful_votes divided by total_votes is equal to or greater than 50%.
helpful_votes_df = over_20_votes_df.filter((over_20_votes_df.helpful_votes/over_20_votes_df.total_votes) >= 0.5)
# + colab={"base_uri": "https://localhost:8080/"} id="FkJMsjpZ2nGR" outputId="cffd6e11-5a55-4520-ad9c-a1270692f9d1"
helpful_votes_df.show()
# + id="Uul86TDl37BL"
# Filter the DataFrame or table created in Step 2, and create a new DataFrame or table that retrieves all the rows where a review was written as part of the Vine program (paid), vine == 'Y'.
vines_df = helpful_votes_df.filter(helpful_votes_df.vine == 'Y')
# + colab={"base_uri": "https://localhost:8080/"} id="7wDJDPBY6AXy" outputId="9c0c28f7-fb3c-48de-cd73-b92091256386"
vines_df.show()
# + id="EIwgDKHx-FwS"
# Repeat Step 3, but this time retrieve all the rows where the review was not part of the Vine program (unpaid), vine == 'N'.
n_vines_df = helpful_votes_df.filter(helpful_votes_df.vine == 'N')
# + colab={"base_uri": "https://localhost:8080/"} id="V5NxG8Gk_Wbi" outputId="a8a611e3-50a4-4323-80a2-445dfa07322e"
n_vines_df.show()
# + id="o_aY2D6i-qE3"
# Determine the total number of reviews, the number of 5-star reviews, and the percentage of 5-star reviews for the two types of review (paid vs unpaid).
# NOTE: This is being done on the data already filtered to include only games with over 50% helpful voted reviews
helpful_votes_df.agg({'total_votes': 'count'}).show() #total votes = 2201075
5_star_df = helpful_votes_df.filter(helpful_votes_df.star_rating == 5)
5_star_df.agg({'total_votes': 'sum'}).show() # total 5 star votes = 898963
# Paid 5-star ratio 0.41131284916201116/1
vines_df.agg({'total_votes': 'sum'}).show() # paid votes = 5728
vine_y_5_star_df = vines_df.filter(vines_df.star_rating == 5)
vine_y_5_star_df.agg({'total_votes': 'sum'}).show() # paid 5 star votes = 2536
# Unpaid 5-star ratio 0.40833043705619204/1
n_vines_df.agg({'total_votes': 'sum'}).show() # unpaid votes = 2195347
vine_n_5_star_df = n_vines_df.filter(n_vines_df.star_rating == 5)
vine_n_5_star_df.agg({'total_votes': 'sum'}).show() # unpaid 5 star votes = 896427
# + id="vSe965rMAdif"
# Export your Vine_Review_Analysis Google Colab Notebook as an ipynb file, and save it to your Amazon_Vine_Analysis GitHub repository.
| Vine_Review_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python-in-the-lab: introduction to the classes I
import os
import numpy as np
import scipy.integrate as integrate
import matplotlib.pylab as plt
# %matplotlib inline
# We will reuse files already analysed, so we can concentrate on the logic!
mainDir = "/home/gf/src/Python/Python-in-the-lab/Bk"
filename = "F64ac_0.01_S.dat"
filename = os.path.join(mainDir, filename)
s, ps = np.loadtxt(filename, comments="#", unpack=True)
plt.loglog(s, ps, 'o'); # Nothing new...
# ### We would like to work a little with the data
#
# The idea is the following:
#
# Can we construction something general and sufficiently abstract from which we can extract the values more easily?
# +
# Your first class
class Sdist: # Note the capital letter
def __init__(self, filename): # Self???
self.size, self.ps = np.loadtxt(filename, comments="#", unpack=True)
# # Done?
THIS_IS_A_CONSTANT = 131231.23131
filename = "F64ac_0.01_S.dat"
filename = os.path.join(mainDir, filename)
s001 = Sdist(filename)
# What is "self" ???
# -
# *self* is not a reserved word, but it is universality used a (self)-reference to the class.
# It simply indentifies the class itself.
#
# s001 is a call to the class, named a *instance* of the class
s001.ps[s001.ps != 0]
# s001.<tab> ?
# s001.__???
# Let's redifine the class
# as I want to get rid of the zeros of ps
class Sdist:
def __init__(self, filename):
self.size, self.ps = np.loadtxt(filename, comments="#", unpack=True)
s_len_ori = len(self.size)
self.size, self.ps = self.avoid_zeros()
print("%i lines deleted" % (s_len_ori - len(self.size)))
def avoid_zeros(self):
is_not_zero = self.ps != 0
s = self.size[is_not_zero]
ps = self.ps[is_not_zero]
return s, ps
s001 = Sdist(filename)
plt.loglog(s001.size, s001.ps, 'o');
s001.ps[-10:] # good!
# ## Improvements
# * Why don't we give just the frequency, so it takes care of all the rest?
# * Can we leave the possibility to avoid the zeros as a choice?
# Let's redifine the class again
class Sdist:
def __init__(self, freq, mainDir, is_avoid_zeros=True):
filename = "F64ac_%s_S.dat" % freq
filename = os.path.join(mainDir, filename)
self.size, self.ps = np.loadtxt(filename, comments="#", unpack=True)
s_len_ori = len(self.size)
self._filename = filename
if is_avoid_zeros:
self.size, self.ps = self.avoid_zeros()
print("%i lines deleted" % (s_len_ori - len(self.size)))
def avoid_zeros(self):
is_not_zero = self.ps != 0
s = self.size[is_not_zero]
ps = self.ps[is_not_zero]
return s, ps
# Is it better to pass a string or a float?
s001 = Sdist(0.01, mainDir, is_avoid_zeros=False)
s002 = Sdist(0.02, mainDir, is_avoid_zeros=False)
s003 = Sdist(0.03, mainDir, is_avoid_zeros=False)
plt.loglog(s001.size, s001.ps, 'o')
plt.loglog(s002.size, s002.ps, 'o')
plt.loglog(s003.size, s003.ps, 'o')
s001._filename
# s001.ps, s001.size: Nothing else?
s001.__dict__.keys()
# ### You seem to be able to do it for all the frequencies...
# +
# Can we now do it for the 3 frequencies?
freqs = np.arange(1,4) / 100
# Can I make a dictionary?
s_distributions = dict()
for freq in freqs:
s_distributions[freq] = Sdist(freq, mainDir)
s_distributions
# -
s_distributions[0.03].ps[:10]
# Let's plot it
for freq in freqs:
sd = s_distributions[freq]
label = "%.2f Hz" % freq
plt.loglog(sd.size, sd.ps, 'o', label=label)
plt.legend(numpoints=1);
# ### Ok, ok, but this not so general enough!
# #### I do not remember the frequencies, and I am interested in the files ending with S, T, u, v. Can we do something more general?
# +
# It would be nice to have something like...
# d_T = Dist("Duration", mainDir)
# d_S = Dist("Size", mainDir)
# d_E = Dist("Energy", mainDir)
# -
# ### It is better to stop for a second. This starts to be pretty complex. I cannot put everything here
#
# Let's make a separated script for the class. We will learn how to use it...
#
# [Work on cplot_distributions.py]
#
# Done?
import cplot_distributions as d1
mainDir = "/home/gf/src/Python/Python-in-the-lab/Bk"
d_size = d1.Dist('Size', mainDir)
d_size.plot()
# ### Classes can redefine operations
# +
class Rectangle:
"""
This class defines operations on Rectangles
"""
def __init__(self, base, height):
self.base = base
self.height = height
self.area = base * height
self.p = self._get_perim()
self._diagonal = (base*base + height * height)**0.5
def _get_perim(self, half=False):
"""
Get the perimeter of the Rectange
Parameters:
----------
half: Bool.
If True, get the semiperimeter
"""
p = self.base + self.height
if half:
return p
else:
return 2 * p
def print_perimeter(self):
return "The perimeter is %f" % self.p
def __add__(self, other):
base = self.base + other.base
height = self.height + other.height
return Rectangle(base,height)
def __repr__(self):
return "Rectangle of {0} by {1}".format(self.base, self.height)
p0 = Rectangle(3,2)
p1 = Rectangle(1,1)
p2 = Rectangle(2,0.5)
# -
print p0.base, p0.height, p0.p, p0.area
p0.print_perimeter()
# Construct a larger Rectange
p3 = p0 + p1
print p3.base, p3.height, p3.p, p3.area
p3
| Py_lectures_04_Classes_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Gn1RVu2xkMdA" colab_type="text"
# # Tutorial Part 7: Uncertainty in Deep Learning
#
# A common criticism of deep learning models is that they tend to act as black boxes. A model produces outputs, but doesn't given enough context to interpret them properly. How reliable are the model's predictions? Are some predictions more reliable than others? If a model predicts a value of 5.372 for some quantity, should you assume the true value is between 5.371 and 5.373? Or that it's between 2 and 8? In some fields this situation might be good enough, but not in science. For every value predicted by a model, we also want an estimate of the uncertainty in that value so we can know what conclusions to draw based on it.
#
# DeepChem makes it very easy to estimate the uncertainty of predicted outputs (at least for the models that support it—not all of them do). Let's start by seeing an example of how to generate uncertainty estimates. We load a dataset, create a model, train it on the training set, predict the output on the test set, and then derive some uncertainty estimates.
#
# ## Colab
#
# This tutorial and the rest in this sequence are designed to be done in Google colab. If you'd like to open this notebook in colab, you can use the following link.
#
# [](https://colab.research.google.com/github/deepchem/deepchem/blob/master/examples/tutorials/07_Uncertainty_In_Deep_Learning.ipynb)
#
# ## Setup
#
# To run DeepChem within Colab, you'll need to run the following cell of installation commands. This will take about 5 minutes to run to completion and install your environment.
# + id="p0MdAUAvkMdD" colab_type="code" colab={}
# %%capture
# %tensorflow_version 1.x
# !wget -c https://repo.anaconda.com/miniconda/Miniconda3-latest-Linux-x86_64.sh
# !chmod +x Miniconda3-latest-Linux-x86_64.sh
# !bash ./Miniconda3-latest-Linux-x86_64.sh -b -f -p /usr/local
# !conda install -y -c deepchem -c rdkit -c conda-forge -c omnia deepchem-gpu=2.3.0
import sys
sys.path.append('/usr/local/lib/python3.7/site-packages/')
# + [markdown] id="BUFgitSSkMdG" colab_type="text"
# We'll use the SAMPL dataset from the MoleculeNet suite to run our experiments in this tutorial. Let's load up our dataset for our experiments, and then make some uncertainty predictions.
# + id="4mHPuoOPkMdH" colab_type="code" outputId="5f0fcd4a-64fa-4d14-ebaf-7f9669ddcb2c" colab={"base_uri": "https://localhost:8080/", "height": 984}
import deepchem as dc
import numpy as np
import matplotlib.pyplot as plot
tasks, datasets, transformers = dc.molnet.load_sampl(reload=False)
train_dataset, valid_dataset, test_dataset = datasets
model = dc.models.MultitaskRegressor(len(tasks), 1024, uncertainty=True)
model.fit(train_dataset, nb_epoch=200)
y_pred, y_std = model.predict_uncertainty(test_dataset)
# + [markdown] id="_DlPZsaekMdL" colab_type="text"
# All of this looks exactly like any other example, with just two differences. First, we add the option `uncertainty=True` when creating the model. This instructs it to add features to the model that are needed for estimating uncertainty. Second, we call `predict_uncertainty()` instead of `predict()` to produce the output. `y_pred` is the predicted outputs. `y_std` is another array of the same shape, where each element is an estimate of the uncertainty (standard deviation) of the corresponding element in `y_pred`. And that's all there is to it! Simple, right?
#
# Of course, it isn't really that simple at all. DeepChem is doing a lot of work to come up with those uncertainties. So now let's pull back the curtain and see what is really happening. (For the full mathematical details of calculating uncertainty, see https://arxiv.org/abs/1703.04977)
#
# To begin with, what does "uncertainty" mean? Intuitively, it is a measure of how much we can trust the predictions. More formally, we expect that the true value of whatever we are trying to predict should usually be within a few standard deviations of the predicted value. But uncertainty comes from many sources, ranging from noisy training data to bad modelling choices, and different sources behave in different ways. It turns out there are two fundamental types of uncertainty we need to take into account.
#
# ### Aleatoric Uncertainty
#
# Consider the following graph. It shows the best fit linear regression to a set of ten data points.
# + id="iLgia0GVkMdM" colab_type="code" outputId="3cd342eb-1bea-476a-8005-94db738b5829" colab={"base_uri": "https://localhost:8080/", "height": 265}
# Generate some fake data and plot a regression line.
x = np.linspace(0, 5, 10)
y = 0.15*x + np.random.random(10)
plot.scatter(x, y)
fit = np.polyfit(x, y, 1)
line_x = np.linspace(-1, 6, 2)
plot.plot(line_x, np.poly1d(fit)(line_x))
plot.show()
# + [markdown] id="7fTPkHSakMdP" colab_type="text"
# The line clearly does not do a great job of fitting the data. There are many possible reasons for this. Perhaps the measuring device used to capture the data was not very accurate. Perhaps `y` depends on some other factor in addition to `x`, and if we knew the value of that factor for each data point we could predict `y` more accurately. Maybe the relationship between `x` and `y` simply isn't linear, and we need a more complicated model to capture it. Regardless of the cause, the model clearly does a poor job of predicting the training data, and we need to keep that in mind. We cannot expect it to be any more accurate on test data than on training data. This is known as *aleatoric uncertainty*.
#
# How can we estimate the size of this uncertainty? By training a model to do it, of course! At the same time it is learning to predict the outputs, it is also learning to predict how accurately each output matches the training data. For every output of the model, we add a second output that produces the corresponding uncertainty. Then we modify the loss function to make it learn both outputs at the same time.
#
# ### Epistemic Uncertainty
#
# Now consider these three curves. They are fit to the same data points as before, but this time we are using 10th degree polynomials.
# + id="hVoRaGn6kMdQ" colab_type="code" outputId="c1e58e58-becf-41d4-afc6-88ad42b943fa" colab={"base_uri": "https://localhost:8080/", "height": 215}
plot.figure(figsize=(12, 3))
line_x = np.linspace(0, 5, 50)
for i in range(3):
plot.subplot(1, 3, i+1)
plot.scatter(x, y)
fit = np.polyfit(np.concatenate([x, [3]]), np.concatenate([y, [i]]), 10)
plot.plot(line_x, np.poly1d(fit)(line_x))
plot.show()
# + [markdown] id="P_1Ag-VPkMdT" colab_type="text"
# Each of them perfectly interpolates the data points, yet they clearly are different models. (In fact, there are infinitely many 10th degree polynomials that exactly interpolate any ten data points.) They make identical predictions for the data we fit them to, but for any other value of `x` they produce different predictions. This is called *epistemic uncertainty*. It means the data does not fully constrain the model. Given the training data, there are many different models we could have found, and those models make different predictions.
#
# The ideal way to measure epistemic uncertainty is to train many different models, each time using a different random seed and possibly varying hyperparameters. Then use all of them for each input and see how much the predictions vary. This is very expensive to do, since it involves repeating the whole training process many times. Fortunately, we can approximate the same effect in a less expensive way: by using dropout.
#
# Recall that when you train a model with dropout, you are effectively training a huge ensemble of different models all at once. Each training sample is evaluated with a different dropout mask, corresponding to a different random subset of the connections in the full model. Usually we only perform dropout during training and use a single averaged mask for prediction. But instead, let's use dropout for prediction too. We can compute the output for lots of different dropout masks, then see how much the predictions vary. This turns out to give a reasonable estimate of the epistemic uncertainty in the outputs.
#
# ### Uncertain Uncertainty?
#
# Now we can combine the two types of uncertainty to compute an overall estimate of the error in each output:
#
# $$\sigma_\text{total} = \sqrt{\sigma_\text{aleatoric}^2 + \sigma_\text{epistemic}^2}$$
#
# This is the value DeepChem reports. But how much can you trust it? Remember how I started this tutorial: deep learning models should not be used as black boxes. We want to know how reliable the outputs are. Adding uncertainty estimates does not completely eliminate the problem; it just adds a layer of indirection. Now we have estimates of how reliable the outputs are, but no guarantees that those estimates are themselves reliable.
#
# Let's go back to the example we started with. We trained a model on the SAMPL training set, then generated predictions and uncertainties for the test set. Since we know the correct outputs for all the test samples, we can evaluate how well we did. Here is a plot of the absolute error in the predicted output versus the predicted uncertainty.
# + id="r3jD4V4rkMdU" colab_type="code" outputId="7ea95f5c-a141-412a-e483-6a719c53a7bc" colab={"base_uri": "https://localhost:8080/", "height": 279}
abs_error = np.abs(y_pred.flatten()-test_dataset.y.flatten())
plot.scatter(y_std.flatten(), abs_error)
plot.xlabel('Standard Deviation')
plot.ylabel('Absolute Error')
plot.show()
# + [markdown] id="rdGOqq_DkMdX" colab_type="text"
# The first thing we notice is that the axes have similar ranges. The model clearly has learned the overall magnitude of errors in the predictions. There also is clearly a correlation between the axes. Values with larger uncertainties tend on average to have larger errors.
#
# Now let's see how well the values satisfy the expected distribution. If the standard deviations are correct, and if the errors are normally distributed (which is certainly not guaranteed to be true!), we expect 95% of the values to be within two standard deviations, and 99% to be within three standard deviations. Here is a histogram of errors as measured in standard deviations.
# + id="IrD6swafkMdY" colab_type="code" outputId="166c3b20-2b8e-45bb-a619-bbd237985d23" colab={"base_uri": "https://localhost:8080/", "height": 265}
plot.hist(abs_error/y_std.flatten(), 20)
plot.show()
# + [markdown] id="bucmsdGSkMda" colab_type="text"
# Most of the values are in the expected range, but there are a handful of outliers at much larger values. Perhaps this indicates the errors are not normally distributed, but it may also mean a few of the uncertainties are too low. This is an important reminder: the uncertainties are just estimates, not rigorous measurements. Most of them are pretty good, but you should not put too much confidence in any single value.
# + [markdown] id="4NwKVrwCkMdb" colab_type="text"
# # Congratulations! Time to join the Community!
#
# Congratulations on completing this tutorial notebook! If you enjoyed working through the tutorial, and want to continue working with DeepChem, we encourage you to finish the rest of the tutorials in this series. You can also help the DeepChem community in the following ways:
#
# ## Star DeepChem on GitHub
# Starring DeepChem on GitHub helps build awareness of the DeepChem project and the tools for open source drug discovery that we're trying to build.
#
# ## Join the DeepChem Gitter
# The DeepChem [Gitter](https://gitter.im/deepchem/Lobby) hosts a number of scientists, developers, and enthusiasts interested in deep learning for the life sciences. Join the conversation!
| examples/tutorials/07_Uncertainty_In_Deep_Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.2 64-bit
# metadata:
# interpreter:
# hash: 767d51c1340bd893661ea55ea3124f6de3c7a262a8b4abca0554b478b1e2ff90
# name: python3
# ---
# from pathlib import Path
import pathlib
# file_path = __file__
# file_path = '/home/vas/Documents/GitHub/vscode_extensions-and-solutions/omnisharp-tracking-sln-for-csproj_for-linux/_03_Operators-and-Expressions/.vscode/'
file_path = '/home/vas/Documents/GitHub/vscode_extensions-and-solutions/omnisharp-tracking-sln-for-csproj_for-linux/_03_Operators-and-Expressions/'
parent_dir = pathlib.PurePath(file_path).parent[0]
| omnisharp-tracking-sln-for-csproj_for-linux/_03_Operators-and-Expressions/.vscode/script-as-jupiterNB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Performance Overview
#
# Here, we will example the performance of FNGS as a function of time on several datasets. These investigations were performed on a 4 core machine (4 threads) with a 4.0 GhZ processor. These investigations were performed on the version of FNGS in ndmg/eric-dev-gkiar-fmri on 03/27.
#
# + magic_args="false" language="script"
#
# ## disklog.sh
# #!/bin/bash -e
# # run this in the background with nohup ./disklog.sh > disk.txt &
# #
# while true; do
# echo "$(du -s $1 | awk '{print $1}')"
# sleep 30
# done
#
#
# ##cpulog.sh
# import psutil
# import time
# import argparse
#
# def cpulog(outfile):
# with open(outfile, 'w') as outf:
# while(True):
# cores = psutil.cpu_percent(percpu=True)
# corestr = ",".join([str(core) for core in cores])
# outf.write(corestr + '\n')
# outf.flush()
# time.sleep(1) # delay for 1 second
#
# def main():
# parser = argparse.ArgumentParser()
# parser.add_argument('outfile', help='the file to write core usage to.')
# args = parser.parse_args()
# cpulog(args.outfile)
#
# if __name__ == "__main__":
# main()
#
#
# ## memlog.sh
# #!/bin/bash -e
# # run this in the background with nohup ./memlog.sh > mem.txt &
# #
# while true; do
# echo "$(free -m | grep buffers/cache | awk '{print $3}')"
# sleep 1
# done
#
#
# ## runonesub.sh
# # A function for generating memory and cpu summaries for fngs pipeline.
# #
# # Usage: ./generate_statistics.sh /path/to/rest /path/to/anat /path/to/output
#
# rm -rf $3
# mkdir $3
#
# ./memlog.sh > ${3}/mem.txt &
# memkey=$!
# python cpulog.py ${3}/cpu.txt &
# cpukey=$!
# ./disklog.sh $3 > ${3}/disk.txt &
# diskkey=$!
#
# res=2mm
# atlas='/FNGS_server/atlases/atlas/MNI152_T1-${res}.nii.gz'
# atlas_brain='/FNGS_server/atlases/atlas/MNI152_T1-${res}_brain.nii.gz'
# atlas_mask='/FNGS_server/atlases/mask/MNI152_T1-${res}_brain_mask.nii.gz'
# lv_mask='/FNGS_server/atlases/mask/HarvOx_lv_thr25-${res}.nii.gz'
# label='/FNGS_server/atlases/label/desikan-${res}.nii.gz'
#
# exec 4<$1
# exec 5<$2
#
# fngs_pipeline $1 $2 $atlas $atlas_brain $atlas_mask $lv_mask $3 none $label --fmt graphml
#
# kill $memkey $cpukey $diskkey
# +
# %matplotlib inline
import numpy as np
import re
import matplotlib.pyplot as plt
from IPython.display import Image, display
def memory_function(infile, dataset):
with open(infile, 'r') as mem:
lines = mem.readlines()
testar = np.asarray([line.strip() for line in lines]).astype(float)/1000
fig=plt.figure()
ax = fig.add_subplot(111)
ax.plot(range(0, testar.shape[0]), testar - min(testar))
ax.set_ylabel('memory usage in GB')
ax.set_xlabel('Time (s)')
ax.set_title(dataset + ' Memory Usage; max = %.2f GB; mean = %.2f GB' % (max(testar), np.mean(testar)))
return fig
def cpu_function(infile, dataset):
with open(infile, 'r') as cpuf:
lines = cpuf.readlines()
testar = [re.split(',',line.strip()) for line in lines][0:-1]
corear = np.zeros((len(testar), len(testar[0])))
for i in range(0, len(testar)):
corear[i,:] = np.array([float(cpu) for cpu in testar[i]])
fig=plt.figure()
ax = fig.add_subplot(111)
lines = [ax.plot(corear[:,i], '--', label='cpu '+ str(i),
alpha=0.5)[0] for i in range(0, corear.shape[1])]
total = corear.sum(axis=1)
lines.append(ax.plot(total, label='all cores')[0])
labels = [h.get_label() for h in lines]
fig.legend(handles=lines, labels=labels, loc='lower right', prop={'size':6})
ax.set_ylabel('CPU usage (%)')
ax.set_ylim([0, max(total)+10])
ax.set_xlabel('Time (s)')
ax.set_title(dataset + ' Processor Usage; max = %.1f per; mean = %.1f per' % (max(total), np.mean(total)))
return fig
def disk_function(infile, dataset):
with open(infile, 'r') as disk:
lines = disk.readlines()
testar = np.asarray([line.strip() for line in lines]).astype(float)/1000000
fig=plt.figure()
ax = fig.add_subplot(111)
ax.plot(range(0, testar.shape[0]), testar - min(testar))
ax.set_ylabel('Disk usage GB')
ax.set_xlabel('Time (30 s)')
ax.set_title(dataset + ' Disk Usage; max = %.2f GB; mean = %.2f GB' % (max(testar), np.mean(testar)))
return fig
# -
# # BNU 1
# +
memfig = memory_function('/data/BNU_sub/BNU_single/mem.txt', 'BNU 1 single')
diskfig = disk_function('/data/BNU_sub/BNU_single/disk.txt', 'BNU 1 single')
cpufig = cpu_function('/data/BNU_sub/BNU_single/cpu.txt', 'BNU 1 single')
memfig.show()
diskfig.show()
cpufig.show()
# -
# # HNU Dataset
memfig = memory_function('/data/HNU_sub/HNU_single/mem.txt', 'HNU 1 single')
diskfig = disk_function('/data/HNU_sub/HNU_single/disk.txt', 'HNU 1 single')
cpufig = cpu_function('/data/HNU_sub/HNU_single/cpu.txt', 'HNU 1 single')
memfig.show()
diskfig.show()
cpufig.show()
# # DC1 Dataset
memfig = memory_function('/data/DC_sub/DC_single/mem.txt', 'DC 1 single')
diskfig = disk_function('/data/DC_sub/DC_single/disk.txt', 'DC 1 single')
cpufig = cpu_function('/data/DC_sub/DC_single/cpu.txt', 'DC 1 single')
memfig.show()
diskfig.show()
cpufig.show()
# # NKI 1
memfig = memory_function('/data/NKI_sub/NKI_single/mem.txt', 'NKI 1 single')
diskfig = disk_function('/data/NKI_sub/NKI_single/disk.txt', 'NKI 1 single')
cpufig = cpu_function('/data/NKI_sub/NKI_single/cpu.txt', 'NKI 1 single')
memfig.show()
diskfig.show()
cpufig.show()
#
| docs/ebridge2/fngs_reg/week_0327/specs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# import pandas as pd
# import numpy as np
dietDF = pd.read_csv("sampleFromQiitaDownload.txt",sep='\t',low_memory=False)
dietDF
# These are the ones from the CleaningVioscreen notebook that do not have really any valid answers
reallyNoAnswerColumns=['alcohol_types',
'allergic_to',
'coprophage',
'diet',
'food_source','food_source_human_food','food_source_pet_store_food','food_source_unspecified','food_source_wild_food',
'food_special',
'food_special_grain_free',
'food_special_organic',
'food_type',
'host',
'hours_outside',
'humans_free_text',
'living_status',
'mental_illness_type',
'name',
'non_food_allergies',
'other_animals_free_text',
'pets_other_freetext',
'setting',
'specialized_diet',
'surf_board_type',
'surf_frequency',
'surf_loal_break_frequency',
'surf_local_break',
'surf_shower_frequency',
'surf_stance',
'surf_sunscreen',
'surf_sunscreen_frequency',
'surf_travel_distance',
'surf_travel_frequency',
'surf_wax',
'surf_weetsuit',
'toilet_water_access',
'weight_cat',
'fermented_consumed',
'fermented_consumed_beer',
'fermented_consumed_chicha',
'fermented_consumed_cider',
'fermented_consumed_cottage_cheese',
'fermented_consumed_fermented_beansmisonatto',
'fermented_consumed_fermented_breadsourdoughinjera',
'fermented_consumed_fermented_fish',
'fermented_consumed_fermented_tofu',
'fermented_consumed_fish_sauce',
'fermented_consumed_kefir_milk',
'fermented_consumed_kefir_water',
'fermented_consumed_kimchi',
'fermented_consumed_kombucha',
'fermented_consumed_mead',
'fermented_consumed_other',
'fermented_consumed_pickled_vegetables',
'fermented_consumed_sauerkraut',
'fermented_consumed_sour_creamcreme_fraiche',
'fermented_consumed_tempeh',
'fermented_consumed_unspecified',
'fermented_consumed_wine',
'fermented_consumed_yogurtlassi',
'fermented_frequency',
'fermented_increased',
'fermented_other',
'fermented_produce_commercial',
'fermented_produce_commercial_beer',
'fermented_produce_commercial_chicha',
'fermented_produce_commercial_cider',
'fermented_produce_commercial_cottage_cheese',
'fermented_produce_commercial_fermented_beansmisonatto',
'fermented_produce_commercial_fermented_breadsourdoughinjera',
'fermented_produce_commercial_fermented_fish',
'fermented_produce_commercial_fermented_tofu',
'fermented_produce_commercial_fish_sauce',
'fermented_produce_commercial_kefir_milk',
'fermented_produce_commercial_kefir_water',
'fermented_produce_commercial_kimchi',
'fermented_produce_commercial_kombucha',
'fermented_produce_commercial_mead',
'fermented_produce_commercial_other',
'fermented_produce_commercial_pickled_vegetables',
'fermented_produce_commercial_sauerkraut',
'fermented_produce_commercial_sour_creamcreme_fraiche',
'fermented_produce_commercial_tempeh',
'fermented_produce_commercial_unspecified',
'fermented_produce_commercial_wine',
'fermented_produce_commercial_yogurtlassi',
'fermented_produce_personal',
'fermented_produce_personal_beer',
'fermented_produce_personal_chicha',
'fermented_produce_personal_cider',
'fermented_produce_personal_cottage_cheese',
'fermented_produce_personal_fermented_beansmisonatto',
'fermented_produce_personal_fermented_breadsourdoughinjera',
'fermented_produce_personal_fermented_fish',
'fermented_produce_personal_fermented_tofu',
'fermented_produce_personal_fish_sauce',
'fermented_produce_personal_kefir_milk',
'fermented_produce_personal_kefir_water',
'fermented_produce_personal_kimchi',
'fermented_produce_personal_kombucha',
'fermented_produce_personal_mead',
'fermented_produce_personal_other',
'fermented_produce_personal_pickled_vegetables',
'fermented_produce_personal_sauerkraut',
'fermented_produce_personal_sour_creamcreme_fraiche',
'fermented_produce_personal_tempeh',
'fermented_produce_personal_unspecified',
'fermented_produce_personal_wine',
'fermented_produce_personal_yogurtlassi',
'body_habitat', #dont want
'body_product', #dont want
'body_site', #dont want
'country_of_birth', #We will just use country
'country_residence', #We will just use country
'depth', #dont want
'economic_region', #We will use country
'height_units', #all are in cm
'last_move', #dont want
'qiita_empo_1', #dont want
'qiita_empo_2', #dont want
'qiita_empo_3', #dont want
'qiita_study_id', #dont want
'state', #We will use country for now
'census_region', #We will use country for now
'altitude' #This has basically no answers
]
# These are all of the boolean columns for the metadata (besides vioscreen)
#
# The only boolean vioscreen ones are:
#
# vioscreen_multivitamin,
# vioscreen_calcium
allBoolsSurvey=["acne_medication",
"acne_medication_otc",
"alcohol_consumption",
"alcohol_types_beercider",
"alcohol_types_red_wine",
"alcohol_types_sour_beers",
"alcohol_types_spiritshard_alcohol",
"alcohol_types_unspecified",
"alcohol_types_white_wine",
"allergic_to_i_have_no_food_allergies_that_i_know_of",
"allergic_to_other",
"allergic_to_peanuts",
"allergic_to_shellfish",
"allergic_to_tree_nuts",
"allergic_to_unspecified",
"appendix_removed",
"assigned_from_geo",
"cat",
"csection",
"dna_extracted",
"dog",
"has_physical_specimen",
"lactose",
"lowgrain_diet_type",
"mental_illness",
"mental_illness_type_anorexia_nervosa",
"mental_illness_type_bipolar_disorder",
"mental_illness_type_bulimia_nervosa",
"mental_illness_type_depression",
"mental_illness_type_ptsd_posttraumatic_stress_disorder",
"mental_illness_type_schizophrenia",
"mental_illness_type_substance_abuse",
"mental_illness_type_unspecified",
"multivitamin",
"nail_biter",
"non_food_allergies_beestings",
"non_food_allergies_drug_eg_penicillin",
"non_food_allergies_pet_dander",
"non_food_allergies_poison_ivyoak",
"non_food_allergies_sun",
"non_food_allergies_unspecified",
"other_supplement_frequency",
"pets_other",
"physical_specimen_remaining",
"public",
"roommates_in_study",
"seasonal_allergies",
"softener",
"specialized_diet_exclude_dairy",
"specialized_diet_exclude_nightshades",
"specialized_diet_exclude_refined_sugars",
"specialized_diet_fodmap",
"specialized_diet_halaal",
"specialized_diet_i_do_not_eat_a_specialized_diet",
"specialized_diet_kosher",
"specialized_diet_modified_paleo_diet",
"specialized_diet_other_restrictions_not_described_here",
"specialized_diet_paleodiet_or_primal_diet",
"specialized_diet_raw_food_diet",
"specialized_diet_unspecified",
"specialized_diet_westenprice_or_other_lowgrain_low_processed_fo",
"subset_age",
"subset_antibiotic_history",
"subset_bmi",
"subset_diabetes",
"subset_healthy",
"subset_ibd",
"tonsils_removed",
"pregnant",
"chickenpox",
"consume_animal_products_abx"]
# These are internal booleans not needed to be processed by this analysis
internalBools=[#"alcohol_consumption_unspecified",
"assigned_from_geo",
"dna_extracted",
"has_physical_specimen",
"mental_illness_type_unspecified",
"non_food_allergies_unspecified",
"physical_specimen_remaining",
"public",
"specialized_diet_i_do_not_eat_a_specialized_diet",
"specialized_diet_other_restrictions_not_described_here",
"specialized_diet_unspecified",
"subset_age",
"subset_bmi",
"subset_diabetes",
"subset_healthy",
"subset_ibd",
"roommates_in_study"]
#Remove the internalBools from the allBools
for rem in internalBools:
allBoolsSurvey.remove(rem)
allBoolsSurvey
# These are booleans we do not care about since we can process them with other means or we dont want them
nonBoolsDontCare=["age_cat",
"age_years",
"anonymized_name",
"birth_year",
"bmi",
"bmi_cat",
"collection_date",
"collection_month",
"collection_season",
"collection_time",
"collection_timestamp",
"description",
"env_biome",
"env_feature",
"env_material",
"env_package",
"exercise_location",
"geo_loc_name",
"host_taxid",
"host_common_name",
#"host_subject_id", Lets just keep this ID, but MUST REMEMBER to not include in analysis
"latitude",
"livingwith",
"longitude",
"physical_specimen_location",
"sample_type",
"scientific_name",
"survey_id",
"taxon_id",
"title",
"weight_units",
"weight_change",
"animal_age",
"animal_free_text",
"animal_gender",
"animal_origin",
"animal_type"]
# These are all the vioscreen columns
vioscreenCols=['vioscreen_a_bev',
'vioscreen_a_cal',
'vioscreen_acesupot',
'vioscreen_activity_level',
'vioscreen_add_sug',
'vioscreen_addsugar',
'vioscreen_adsugtot',
'vioscreen_age',
'vioscreen_alanine',
'vioscreen_alcohol',
'vioscreen_alcohol_servings',
'vioscreen_alphacar',
'vioscreen_alphtoce',
'vioscreen_alphtoco',
'vioscreen_arginine',
'vioscreen_ash',
'vioscreen_aspartam',
'vioscreen_aspartic',
'vioscreen_avcarb',
'vioscreen_bcodeid',
'vioscreen_betacar',
'vioscreen_betacryp',
'vioscreen_betaine',
'vioscreen_betatoco',
'vioscreen_biochana',
'vioscreen_bmi',
'vioscreen_caffeine',
'vioscreen_calcium',
'vioscreen_calcium_avg',
'vioscreen_calcium_dose',
'vioscreen_calcium_freq',
'vioscreen_calcium_from_dairy_servings',
'vioscreen_calcium_servings',
'vioscreen_calories',
'vioscreen_carbo',
'vioscreen_cholest',
'vioscreen_choline',
'vioscreen_clac9t11',
'vioscreen_clat10c12',
'vioscreen_copper',
'vioscreen_coumest',
'vioscreen_cystine',
'vioscreen_d_cheese',
'vioscreen_d_milk',
'vioscreen_d_tot_soym',
'vioscreen_d_total',
'vioscreen_d_yogurt',
'vioscreen_daidzein',
'vioscreen_database',
'vioscreen_delttoco',
'vioscreen_discfat_oil',
'vioscreen_discfat_sol',
'vioscreen_dob',
'vioscreen_eer',
'vioscreen_email',
'vioscreen_erythr',
'vioscreen_f_citmlb',
'vioscreen_f_nj_citmlb',
'vioscreen_f_nj_other',
'vioscreen_f_nj_total',
'vioscreen_f_other',
'vioscreen_f_total',
'vioscreen_fat',
'vioscreen_fiber',
'vioscreen_fibh2o',
'vioscreen_fibinso',
'vioscreen_finished',
'vioscreen_fish_servings',
'vioscreen_fol_deqv',
'vioscreen_fol_nat',
'vioscreen_fol_syn',
'vioscreen_formontn',
'vioscreen_fried_fish_servings',
'vioscreen_fried_food_servings',
'vioscreen_frt5_day',
'vioscreen_frtsumm',
'vioscreen_fructose',
'vioscreen_fruit_servings',
'vioscreen_g_nwhl',
'vioscreen_g_total',
'vioscreen_g_whl',
'vioscreen_galactos',
'vioscreen_gammtoco',
'vioscreen_gender',
'vioscreen_genistn',
'vioscreen_glac',
'vioscreen_gltc',
'vioscreen_glucose',
'vioscreen_glutamic',
'vioscreen_glycine',
'vioscreen_glycitn',
'vioscreen_grams',
'vioscreen_hei2010_dairy',
'vioscreen_hei2010_empty_calories',
'vioscreen_hei2010_fatty_acids',
'vioscreen_hei2010_fruit',
'vioscreen_hei2010_greens_beans',
'vioscreen_hei2010_protien_foods',
'vioscreen_hei2010_refined_grains',
'vioscreen_hei2010_score',
'vioscreen_hei2010_sea_foods_plant_protiens',
'vioscreen_hei2010_sodium',
'vioscreen_hei2010_veg',
'vioscreen_hei2010_whole_fruit',
'vioscreen_hei2010_whole_grains',
'vioscreen_hei_drk_g_org_veg_leg',
'vioscreen_hei_fruit',
'vioscreen_hei_grains',
'vioscreen_hei_meat_beans',
'vioscreen_hei_milk',
'vioscreen_hei_non_juice_frt',
'vioscreen_hei_oils',
'vioscreen_hei_sat_fat',
'vioscreen_hei_score',
'vioscreen_hei_sodium',
'vioscreen_hei_sol_fat_alc_add_sug',
'vioscreen_hei_veg',
'vioscreen_hei_whl_grains',
'vioscreen_height',
'vioscreen_histidin',
'vioscreen_inositol',
'vioscreen_iron',
'vioscreen_isoleuc',
'vioscreen_isomalt',
'vioscreen_joules',
'vioscreen_juice_servings',
'vioscreen_lactitol',
'vioscreen_lactose',
'vioscreen_legumes',
'vioscreen_leucine',
'vioscreen_line_gi',
'vioscreen_low_fat_dairy_serving',
'vioscreen_lutzeax',
'vioscreen_lycopene',
'vioscreen_lysine',
'vioscreen_m_egg',
'vioscreen_m_fish_hi',
'vioscreen_m_fish_lo',
'vioscreen_m_frank',
'vioscreen_m_meat',
'vioscreen_m_mpf',
'vioscreen_m_nutsd',
'vioscreen_m_organ',
'vioscreen_m_poult',
'vioscreen_m_soy',
'vioscreen_magnes',
'vioscreen_maltitol',
'vioscreen_maltose',
'vioscreen_mangan',
'vioscreen_mannitol',
'vioscreen_methhis3',
'vioscreen_methion',
'vioscreen_mfa141',
'vioscreen_mfa161',
'vioscreen_mfa181',
'vioscreen_mfa201',
'vioscreen_mfa221',
'vioscreen_mfatot',
'vioscreen_multi_calcium_avg',
'vioscreen_multi_calcium_dose',
'vioscreen_multivitamin',
'vioscreen_multivitamin_freq',
'vioscreen_natoco',
'vioscreen_nccglbr',
'vioscreen_nccglgr',
'vioscreen_niacin',
'vioscreen_niacineq',
'vioscreen_nitrogen',
'vioscreen_non_fried_fish_servings',
'vioscreen_nutrient_recommendation',
'vioscreen_omega3',
'vioscreen_oxalic',
'vioscreen_oxalicm',
'vioscreen_pantothe',
'vioscreen_pectins',
'vioscreen_pfa182',
'vioscreen_pfa183',
'vioscreen_pfa184',
'vioscreen_pfa204',
'vioscreen_pfa205',
'vioscreen_pfa225',
'vioscreen_pfa226',
'vioscreen_pfatot',
'vioscreen_phenylal',
'vioscreen_phosphor',
'vioscreen_phytic',
'vioscreen_pinitol',
'vioscreen_potass',
'vioscreen_procdate',
'vioscreen_proline',
'vioscreen_protanim',
'vioscreen_protein',
'vioscreen_protocol',
'vioscreen_protveg',
'vioscreen_questionnaire',
'vioscreen_recno',
'vioscreen_retinol',
'vioscreen_rgrain',
'vioscreen_ribofla',
'vioscreen_sacchar',
'vioscreen_salad_vegetable_servings',
'vioscreen_satoco',
'vioscreen_scf',
'vioscreen_scfv',
'vioscreen_selenium',
'vioscreen_serine',
'vioscreen_sfa100',
'vioscreen_sfa120',
'vioscreen_sfa140',
'vioscreen_sfa160',
'vioscreen_sfa170',
'vioscreen_sfa180',
'vioscreen_sfa200',
'vioscreen_sfa220',
'vioscreen_sfa40',
'vioscreen_sfa60',
'vioscreen_sfa80',
'vioscreen_sfatot',
'vioscreen_sodium',
'vioscreen_sorbitol',
'vioscreen_srvid',
'vioscreen_starch',
'vioscreen_started',
'vioscreen_subject_id',
'vioscreen_sucpoly',
'vioscreen_sucrlose',
'vioscreen_sucrose',
'vioscreen_sweet_servings',
'vioscreen_tagatose',
'vioscreen_tfa161t',
'vioscreen_tfa181t',
'vioscreen_tfa182t',
'vioscreen_tgrain',
'vioscreen_thiamin',
'vioscreen_threonin',
'vioscreen_time',
'vioscreen_totaltfa',
'vioscreen_totcla',
'vioscreen_totfolat',
'vioscreen_totsugar',
'vioscreen_tryptoph',
'vioscreen_tyrosine',
'vioscreen_user_id',
'vioscreen_v_drkgr',
'vioscreen_v_orange',
'vioscreen_v_other',
'vioscreen_v_potato',
'vioscreen_v_starcy',
'vioscreen_v_tomato',
'vioscreen_v_total',
'vioscreen_valine',
'vioscreen_veg5_day',
'vioscreen_vegetable_servings',
'vioscreen_vegsumm',
'vioscreen_visit',
'vioscreen_vita_iu',
'vioscreen_vita_rae',
'vioscreen_vita_re',
'vioscreen_vitb12',
'vioscreen_vitb6',
'vioscreen_vitc',
'vioscreen_vitd',
'vioscreen_vitd2',
'vioscreen_vitd3',
'vioscreen_vitd_iu',
'vioscreen_vite_iu',
'vioscreen_vitk',
'vioscreen_water',
'vioscreen_weight',
'vioscreen_wgrain',
'vioscreen_whole_grain_servings',
'vioscreen_xylitol',
'vioscreen_zinc']
#Items that have frequency
freqAnsArray=['alcohol_frequency',
'cosmetics_frequency',
'exercise_frequency',
'fermented_plant_frequency',
'flossing_frequency',
'frozen_dessert_frequency',
'fruit_frequency',
'high_fat_red_meat_frequency',
'homecooked_meals_frequency',
'meat_eggs_frequency',
'milk_cheese_frequency',
'milk_substitute_frequency',
'one_liter_of_water_a_day_frequency',
'pool_frequency',
'poultry_frequency',
'prepared_meals_frequency',
'probiotic_frequency',
'ready_to_eat_meals_frequency',
'red_meat_frequency',
'salted_snacks_frequency',
'seafood_frequency',
'smoking_frequency',
'sugar_sweetened_drink_frequency',
'sugary_sweets_frequency',
'teethbrushing_frequency',
'vegetable_frequency',
'vitamin_b_supplement_frequency',
'vitamin_d_supplement_frequency',
'whole_grain_frequency',
'vivid_dreams',
'artificial_sweeteners',
'olive_oil',
'whole_eggs']
# +
#Unique column situations
#'bowel_movement_frequency'
#This has answers
bowelQualityAns=['One','Not provided','Two','Less than one','Three','Unspecified','Four','Five or more']
# -
# Lets start cleaning!
dietDF = pd.read_csv("sampleFromQiitaDownload.txt",sep='\t',low_memory=False)
dietDF=dietDF.loc[dietDF['host_common_name'] == "human"]
dietDF=dietDF.loc[dietDF['env_material'] == "feces"]
print(dietDF.shape)
dietDF=dietDF.drop(columns=reallyNoAnswerColumns)
print(dietDF.shape)
dietDF=dietDF.drop(columns=internalBools)
print(dietDF.shape)
dietDF=dietDF.drop(columns=nonBoolsDontCare)
print(dietDF.shape)
#Lets avoid looking at vioscreen right now in this group
dietDF=dietDF.drop(columns=vioscreenCols)
print(dietDF.shape)
#Lets set the sample_name and host_subject_id as the index
dietDF=dietDF.set_index(["sample_name","host_subject_id"])
# +
#Collect all the bad answers if needing to view!
#and Collect array with frequency in name
allColCountsBad={}
allColsDietDF=dietDF.columns
freqArry=[]
for curCol in allColsDietDF:
if 'frequency' in curCol:
freqArry.append(curCol)
allColCountsBad[curCol]=dietDF[curCol].value_counts().to_dict()
# -
# # IMPUTATION METHODS
# After reading alot about imputation methods (search item nonresponse on Google), it is clear to me we need to run some type of algorithm to help fill in all of the missing data.
#
# https://medium.com/ibm-data-science-experience/missing-data-conundrum-exploration-and-imputation-techniques-9f40abe0fd87
# At the bottom described what I was thinking, using machine learning to find the best values...
# "Machine learning algorithms like eXtreme Gradient Boosting (xgboost) automatically learn the best imputation value for the missing data based on the training loss reduction."
#
# I think trying MICE with fancyimpute as well as xgboost research may be a great approach
#
# https://github.com/iskandr/fancyimpute
#
# https://xgboost.readthedocs.io/en/latest/
# # TEST IMPUTATION
# At first, to test out imputation methods, we need to gather a sample of the data that has NO missing values, first we need to clean up the discrete and boolean values, and then grab the no missings. THEN we need to randomly select throughout the matrix a proportional amount of NaN replacement, so we can then test the mean squared error of the imputation methods.
dietDF
# +
#dietDF
#Lets start making functions to clean up things
#Lets clean the unique bowel quality
def fixBowelQual(item):
bowelQualityDict={np.nan: np.nan, 'One': 1,'Not provided':np.nan,'Two':2,'Less than one':1,'Three':3,'Unspecified':np.nan,'Four':4,'Five or more':5}
result=bowelQualityDict[item]
return result
dietDF['bowel_movement_frequency']=dietDF['bowel_movement_frequency'].map(fixBowelQual)
# +
#Lets clean up all the booleans that are normal
#**IMPORTANT: We assume here that not applicable means false/No**
def fixNormBools(item):
boolAnswerPossible={np.nan:np.nan,'Unspecified':np.nan,'Not applicable':0,'not applicable':0,'Not provided':np.nan,
'Yes':1,'yes':1,'No':0,'no':0,'true':1,'True':1,'false':0,'False':0,
'Not sure': np.nan, 'not sure': np.nan, 'TRUE': 1, 'FALSE': 0}
result=boolAnswerPossible[item]
return result
for bCol in allBoolsSurvey:
dietDF[bCol]=dietDF[bCol].map(fixNormBools)
# +
#Lets clean up all the booleans that are like medical conditions
#**IMPORTANT: We assume here that not applicable means false/No**
#**Also, marking Self-diagnosed as np.nan, could change to alter results?**
#**Marking alternative med as a true/1**
medFeatures=["acid_reflux","add_adhd","alzheimers","asd","autoimmune","cancer","cardiovascular_disease","cdiff","clinical_condition",
"depression_bipolar_schizophrenia","diabetes","epilepsy_or_seizure_disorder","fungal_overgrowth","ibd","ibs",
"kidney_disease","liver_disease","lung_disease","migraine","pku","sibo","skin_condition","thyroid"]
def fixMedicalBools(item):
boolMedPossible={np.nan:np.nan,'Unspecified':np.nan,'Not applicable': 0,'not applicable': 0,'Not provided':np.nan,
"I do not have this condition": 0, "Diagnosed by a medical professional (doctor, physician assistant)": 1,
"Self-diagnosed": np.nan, "Diagnosed by an alternative medicine practitioner": 0}
result=boolMedPossible[item]
return result
for bCol in medFeatures:
dietDF[bCol]=dietDF[bCol].map(fixMedicalBools)
# +
#Lets work on the items that have frequency!
def fixfrequencyFeatures(item):
boolFreqPossible={np.nan:np.nan,'Unspecified':np.nan,'Not provided':np.nan, "Daily": 4, "Regularly (3-5 times/week)": 3,
"Occasionally (1-2 times/week)":2, "Rarely (less than once/week)": 1, "Never": 0,
"Rarely (a few times/month)":1}
result=boolFreqPossible[item]
return result
for bCol in freqAnsArray:
print(bCol)
dietDF[bCol]=dietDF[bCol].map(fixfrequencyFeatures)
# +
#Used to view what I was missing
#allAlteredArr=freqAnsArray+medFeatures+allBoolsSurvey+['bowel_movement_frequency']
#print(len(allAlteredArr))
#print(len(dietDF.columns))
#notWorkedFeats=list(dietDF.columns)
#for featCol in allAlteredArr:
# notWorkedFeats.remove(featCol)
#for jj in notWorkedFeats:
# print(dietDF[jj].value_counts())
# print(jj)
#Assume they all have nans are part of their counts btw
# +
types_of_plantsAns={'11 to 20':2,
'Not provided':np.nan,
'6 to 10':1,
'21 to 30':3,
'More than 30':4,
'Less than 5':0,
'Unspecified':np.nan,
np.nan:np.nan}
sleep_durationAns={'7-8 hours':3,
'6-7 hours':2,
'8 or more hours':4,
'5-6 hours':1,
'Not provided':np.nan,
'Less than 5 hours':0,
'Unspecified':np.nan,
np.nan:np.nan}
roommatesAns={'None':0,
'Not provided':np.nan,
'One':1,
'Two':2,
'Three':3,
'More than three':4,
'Unspecified':np.nan,
np.nan:np.nan}
sexAns={'female':2,
'male':1,
'Not provided':np.nan,
'unspecified':np.nan,
'other':np.nan,
np.nan:np.nan}
raceAns={'Caucasian':1,
'Asian or Pacific Islander':2,
'Other':0,
'Hispanic':3,
'Not provided':np.nan,
'African American':4,
'Unspecified':np.nan,
np.nan:np.nan}
level_of_educationAns={'Graduate or Professional degree':6,
'Not provided':np.nan,
'Bachelor\'s degree':4,
'Some college or technical school':2,
'Some graduate school or professional':5,
'High School or GED equilivant':1,
'Did not complete high school':0,
'Unspecified':np.nan,
'Associate\'s degree':3,
np.nan:np.nan}
last_travelAns={'I have not been outside of my country of residence in the past year.':0,
'1 year':4,
'3 months':2,
'6 months':3,
'Month':1,
'Not provided':np.nan,
'Unspecified':np.nan,
np.nan:np.nan}
antibiotic_historyAns={'I have not taken antibiotics in the past year.':0,
'Year':4,
'6 months':3,
'Month':2,
'Not provided':np.nan,
'Week':1,
'Unspecified':np.nan,
np.nan:np.nan}
cancer_treatmentAns={'Not provided':np.nan,
'Unspecified':np.nan,
'Surgery only':1,
'Chemotherapy':3,
'Radiation therapy':2,
'No treatment':0,
np.nan:np.nan}
breastmilk_formula_ensureAns={'false':0,
'No':0,
'Not provided':np.nan,
'Unspecified':np.nan,
'I eat both solid food and formula/breast milk':2,
'true':1,
'Yes':1,
np.nan:np.nan}
bowel_movement_qualityAns={'I tend to have normal formed stool - Type 3 and 4':0,
'Not provided':np.nan,
'I tend to have diarrhea (watery stool) - Type 5, 6 and 7':1,
'I tend to have normal formed stool':0,
'I tend to be constipated (have difficulty passing stool) - Type 1 and 2':2,
'I tend to have diarrhea (watery stool)':1,
'I tend to be constipated (have difficulty passing stool)':2,
'I don\'t know, I do not have a point of reference':np.nan,
'Unspecified':np.nan,
np.nan:np.nan}
contraceptiveAns={'No':0,
'Not provided':np.nan,
'Yes, I am taking the "pill"':1,
'Yes, I use a hormonal IUD (Mirena)':2,
'Unspecified':np.nan,
'Yes, I use the NuvaRing':3,
'Yes, I use an injected contraceptive (DMPA)':4,
'Yes, I use a contraceptive patch (Ortho-Evra)':5,
np.nan:np.nan}
countryAns={'USA':56,
'United Kingdom':55,
'Australia':54,
'Canada':53,
'Unspecified':np.nan,
'Philippines':52,
'Switzerland':51,
'Germany':50,
'Ireland':49,
'Belgium':48,
'France':47,
'Not provided':np.nan,
'Sweden':46,
'Netherlands':45,
'New Zealand':44,
'Norway':43,
'Italy':42,
'Spain':41,
'Japan':40,
'Czech Republic':39,
'Morocco':38,
'Singapore':37,
'Austria':36,
'Denmark':35,
'Thailand':34,
'China':33,
'Guernsey':32,
'United Arab Emirates':31,
'India':30,
'Slovakia':29,
'Brazil':28,
'Jersey':27,
'Portugal':26,
'Serbia':25,
'Oman':24,
'Isle of Man':23,
'Hong Kong':22,
'Poland':21,
'Finland':20,
'South Korea':19,
'Argentina':18,
'Colombia':17,
'United States Minor Outlying Islands':16,
'Croatia':15,
'Mexico':14,
'Greece':13,
'Israel':12,
'Puerto Rico':11,
'Peru':10,
'Latvia':9,
'Estonia':8,
'Slovenia':7,
'Lebanon':6,
'Cyprus':5,
'Romania':4,
'Georgia':3,
'Paraguay':2,
'Malta':1,
np.nan:np.nan}
deodorant_useAns={'I use deodorant':2,
'I do not use deodorant or an antiperspirant':0,
'I use an antiperspirant':1,
'Not sure, but I use some form of deodorant/antiperspirant':1,
'Not provided':np.nan,
'Unspecified':np.nan,
np.nan:np.nan}
diabetes_typeAns={'Not provided':0,
'Unspecified':0,
'Type II diabetes':3,
'Type I diabetes':2,
'Gestational diabetes':1,
np.nan:np.nan}
diet_typeAns={'Omnivore':1,
'Omnivore but do not eat red meat':2,
'Vegetarian but eat seafood':3,
'Vegetarian':4,
'Vegan':5,
'Not provided':np.nan,
'Unspecified':np.nan,
np.nan:np.nan}
dominant_handAns={'I am right handed':3,
'I am left handed':2,
'Not provided':np.nan,
'I am ambidextrous':1,
'Unspecified':np.nan,
np.nan:np.nan}
drinking_water_sourceAns={'City':1,
'Filtered':2,
'Bottled':4,
'Well':3,
'Not provided':np.nan,
'Not sure':np.nan,
'Unspecified':np.nan,
np.nan:np.nan}
drinks_per_sessionAns={'Not provided':np.nan,
'1-2':2,
'I don\'t drink':0,
'1':1,
'2-3':3,
'Unspecified':np.nan,
'3-4':4,
'4+':6,
'2-Jan':np.nan,
'3-Feb':np.nan,
'4-Mar':np.nan,
np.nan:np.nan}
fed_as_infantAns={'Primarily breast milk':3,
'Not provided':np.nan,
'Primarily infant formula':2,
'A mixture of breast milk and formula':1,
'Not sure':np.nan,
'Unspecified':np.nan,
np.nan:np.nan}
flu_vaccine_dateAns={'I have not gotten the flu vaccine in the past year.':0,
'Year':0.25,
'6 months':0.5,
'Month':1,
'Not provided':np.nan,
'Week':1,
'Unspecified':np.nan,
np.nan:np.nan}
glutenAns={'No':0,
'I do not eat gluten because it makes me feel bad':1,
'Not provided':np.nan,
'I was diagnosed with gluten allergy (anti-gluten IgG), but not celiac disease':2,
'Unspecified':np.nan,
'I was diagnosed with celiac disease':3,
np.nan:np.nan}
ibd_diagnosisAns={'Not provided':0,
'Unspecified':0,
'Crohn\'s disease':1,
'Ulcerative colitis':2,
np.nan:np.nan}
ibd_diagnosis_refinedAns={'Not provided':0,
'Unspecified':0,
'Colonic Crohn\'s Disease':1,
'Ulcerative colitis':2,
'Ileal Crohn\'s Disease':3,
'Microcolitis':4,
'Ileal and Colonic Crohn\'s Disease':5,
np.nan:np.nan}
# -
toughFeatureDict={'types_of_plants':types_of_plantsAns,'sleep_duration':sleep_durationAns,'roommates':roommatesAns,'sex':sexAns,
'race':raceAns,'level_of_education':level_of_educationAns,'last_travel':last_travelAns,'antibiotic_history':antibiotic_historyAns,
'cancer_treatment':cancer_treatmentAns, 'breastmilk_formula_ensure':breastmilk_formula_ensureAns,
'bowel_movement_quality':bowel_movement_qualityAns,'contraceptive':contraceptiveAns,'country':countryAns,
'deodorant_use':deodorant_useAns,'diabetes_type':diabetes_typeAns,'diet_type':diet_typeAns,'dominant_hand':dominant_handAns,
'drinking_water_source':drinking_water_sourceAns,'drinks_per_session':drinks_per_sessionAns,'fed_as_infant':fed_as_infantAns,
'flu_vaccine_date':flu_vaccine_dateAns,'gluten':glutenAns,'ibd_diagnosis':ibd_diagnosisAns,'ibd_diagnosis_refined':ibd_diagnosis_refinedAns}
#Lets convert the final harder features into numbered categories
for tf in toughFeatureDict:
print(tf)
tfPossibles=toughFeatureDict[tf]
dietDF[tf]=dietDF[tf].map(lambda x: tfPossibles[x])
# +
#Lets fix the final few!
def finalfeatFixMethod(item):
if (item == np.nan) or (item=='Unspecified') or (item=='Not provided'):
return np.nan
else:
return item
finalFixingFeats=['bmi_corrected','age_corrected','elevation','height_cm','weight_kg']
for ffeet in finalFixingFeats:
dietDF[ffeet]=dietDF[ffeet].map(finalfeatFixMethod)
# -
dietDF.to_csv(path_or_buf='cleanedUpMetadata_noVio_AGP_humfece.csv')
# # DONE CLEANING NOW NEED IMPUTATION
# +
#See notebook: metadataImputationAnalysisAGP.ipynb
| VioAndMetadata_Cleaning/CleanUpMetadata_NoVioUse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="sOmDhenwAp0b"
# # !pip install imblearn
# To enable plotting graphs in Jupyter notebook
# %matplotlib inline
# + colab={} colab_type="code" id="LEhc_mvZAp0p"
import pandas as pd
from sklearn.linear_model import LogisticRegression
# importing ploting libraries
import matplotlib.pyplot as plt
#importing seaborn for statistical plots
import seaborn as sns
#Let us break the X and y dataframes into training set and test set. For this we will use
#Sklearn package's data splitting function which is based on random function
from sklearn.model_selection import train_test_split
import numpy as np
# calculate accuracy measures and confusion matrix
from sklearn import metrics
# + colab={} colab_type="code" id="-jyxMF0YAp1B"
# Since it is a data file with no header, we will supply the column names which have been obtained from the above URL
# Create a python list of column names called "names"
colnames = ['preg', 'plas', 'pres', 'skin', 'test', 'mass', 'pedi', 'age', 'class']
#Load the file from local directory using pd.read_csv which is a special form of read_table
#while reading the data, supply the "colnames" list
pima_df = pd.read_csv("pima-indians-diabetes.data", names= colnames)
# + colab={} colab_type="code" id="qvUWbqauAp2U" outputId="6b64a203-e04e-4d46-9840-aa04f2f862e3"
pima_df.head(50)
# + colab={} colab_type="code" id="eVVZ2neGAp21" outputId="576e94b5-3775-44c3-e63e-0abc1bcaf026"
# Let us check whether any of the columns has any value other than numeric i.e. data is not corrupted such as a "?" instead of
# a number.
# we use np.isreal a numpy function which checks each column for each row and returns a bool array,
# where True if input element is real.
# applymap is pandas dataframe function that applies the np.isreal function columnwise
# Following line selects those rows which have some non-numeric value in any of the columns hence the ~ symbol
pima_df[~pima_df.applymap(np.isreal).all(1)]
# + colab={} colab_type="code" id="xaaqMkXuAp3H"
# replace the missing values in pima_df with median value :Note, we do not need to specify the column names
# every column's missing value is replaced with that column's median respectively
#pima_df = pima_df.fillna(pima_df.median())
#pima_df
# + colab={} colab_type="code" id="ztkviwlpAp3j"
#Lets analysze the distribution of the various attributes
pima_df.describe().transpose()
# + colab={} colab_type="code" id="ZPBj-vRdAp4C" outputId="bea55c7a-8e17-4977-a6c5-aa57dc4f290a"
# Let us look at the target column which is 'class' to understand how the data is distributed amongst the various values
pima_df.groupby(["class"]).count()
# Most are not diabetic. The ratio is almost 1:2 in favor or class 0. The model's ability to predict class 0 will
# be better than predicting class 1.
# + colab={} colab_type="code" id="C6yNuY4XCtY5"
# Pairplot using sns
sns.pairplot(pima_df , hue='class' , diag_kind = 'kde')
# + colab={} colab_type="code" id="w30eoOQWMetl"
#data for all the attributes are skewed, especially for the variable "test"
#The mean for test is 80(rounded) while the median is 30.5 which clearly indicates an extreme long tail on the right
# + colab={} colab_type="code" id="Ti87GrOVMmUb"
# Attributes which look normally distributed (plas, pres, skin, and mass).
# Some of the attributes look like they may have an exponential distribution (preg, test, pedi, age).
# Age should probably have a normal distribution, the constraints on the data collection may have skewed the distribution.
# There is no obvious relationship between age and onset of diabetes.
# There is no obvious relationship between pedi function and onset of diabetes.
# + colab={} colab_type="code" id="-jB72BJqCuwi" outputId="01801023-1805-435c-df36-b85f494e0fab"
array = pima_df.values
X = array[:,0:7] # select all rows and first 8 columns which are the attributes
Y = array[:,8] # select all rows and the 8th column which is the classification "Yes", "No" for diabeties
test_size = 0.30 # taking 70:30 training and test set
seed = 7 # Random numbmer seeding for reapeatability of the code
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size=test_size, random_state=seed)
type(X_train)
# + colab={} colab_type="code" id="CxMHHw95Ap7W" outputId="e92183bc-9ee0-4a79-abfa-bdfc5c9453be"
# Fit the model on original data i.e. before upsampling
model = LogisticRegression()
model.fit(X_train, y_train)
y_predict = model.predict(X_test)
model_score = model.score(X_test, y_test)
print(model_score)
print(metrics.confusion_matrix(y_test, y_predict))
print(metrics.classification_report(y_test, y_predict))
# + colab={} colab_type="code" id="pF1EEd9uAp7g" outputId="9cc1a59e-c264-41f0-f1cf-954120e49e15"
cm = metrics.confusion_matrix(y_test, y_predict)
plt.clf()
plt.imshow(cm, interpolation='nearest', cmap=plt.cm.Wistia)
classNames = ['NonDiabetic', 'Diabetic']
plt.title('Confusion Matrix - Test Data')
plt.ylabel('Actual label')
plt.xlabel('Predicted label')
tick_marks = np.arange(len(classNames))
plt.xticks(tick_marks, classNames, rotation=45)
plt.yticks(tick_marks, classNames)
s = [['G1', 'G2'], ['G1','G2']]
for i in range(2):
for j in range(2):
plt.text(j,i, str(s[i][j])+" = "+str(cm[i][j]))
plt.show()
# -
| M5 Pridictive Modeling/M5 W2 Logistics Regression/Logistic_Pima_Indians-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tic-Tac-Toe Game
#
# This is the first milestone project in the course "Complete Python Bootcamp" in Udemy.
# The script below can serve as detailed tutorial on how to create a simple Tic-Tac-Toe game executed in Jupyter Notebook. <br>
# <sub> **Important Note:** The current version of the game is developed by me with the help of the Walkthrough Steps Workbook provided in the course materials so more efficient solutions could exist. Link to course: https://www.udemy.com/complete-python-bootcamp/ <sub/>
# **Step 1: Write a function that can print out a board. Set up your board as a list, where each index 1-9 corresponds with a number on a number pad, so you get a 3 by 3 board representation.**
# +
from IPython.display import clear_output
def display_board(board):
print('---------------------')
print('|' + ' ' + '|' + ' ' + '|' + ' '+ '|' )
print('|'+ ' ' + board[7]+ ' ' + ' ' + board[8]+ ' ' + ' ' + board[9]+ ' ' )
print('|' + ' ' + '|' + ' ' + '|' + ' '+ '|' )
print('---------------------')
print('|' + ' ' + '|' + ' ' + '|' + ' '+ '|' )
print('|'+ ' ' + board[4]+ ' ' + ' ' + board[5]+ ' ' + ' ' + board[6]+ ' ' )
print('|' + ' ' + '|' + ' ' + '|' + ' '+ '|' )
print('---------------------')
print('|' + ' ' + '|' + ' ' + '|' + ' '+ '|' )
print('|'+ ' ' + board[1]+ ' ' + ' ' + board[2]+ ' ' + ' ' + board[3]+ ' ' )
print('|' + ' ' + '|' + ' ' + '|' + ' '+ '|' )
print('---------------------')
# -
# **Step 2: Write a function that can take in a player input and assign their marker as 'X' or 'O'. Think about using *while* loops to continually ask until you get a correct answer.**
def player_input():
answer1= input("Which player do you choose- X or O? ")
while answer1.upper() not in ["X","O"]:
answer1 = input("Hmm! It looks like there is a typo in your previous answer. Please, confirm which player do you choose- X or O? ")
else:
print (f"You have chosen to be player: {answer1.upper()}")
if answer1.upper() == "X":
answer2 = "O"
else:
answer2 = "X"
# **Step 3: Write a function that takes in the board list object, a marker ('X' or 'O'), and a desired position (number 1-9) and assigns it to the board.**
test_board = ['#','X',' ',' ',' ',' ',' ',' ',' ',' '] # first, create an empty test board
def place_marker(board, marker, position):
if position in range(1,10):
board[position] = marker
# **Step 4: Write a function that takes in a board and a mark (X or O) and then checks to see if that mark has won. **
def win_check(board, mark):
combinationslist = [''.join(board[1:4]), ''.join(board[4:7]), ''.join(board[7:10]), ''.join(board[1:10:3]),
''.join(board[2:10:3]), ''.join(board[3:10:3]), ''.join(board[1:10:4]), ''.join(board[3:8:2])]
check = []
for item in combinationslist:
check.append(mark*3 in item)
return sum(check) >0
# **Step 5: Write a function that uses the random module to randomly decide which player goes first. You may want to lookup random.randint() Return a string of which player went first.**
# +
import random
def choose_first():
first = random.randint(1,1000000)
if first in range(1,500000):
print("Player X is first")
else:
print("Player O is first")
return first
# -
# **Step 6: Write a function that returns a boolean indicating whether a space on the board is freely available.**
def space_check(board, position):
return board[position] == ' '
# **Step 7: Write a function that checks if the board is full and returns a boolean value. True if full, False otherwise.**
def full_board_check(board):
return ' ' not in board
# **Step 8: Write a function that asks for a player's next position (as a number 1-9) and then uses the function from step 6 to check if it's a free position. If it is, then return the position for later use. Handle potential error if a string is provided in this field.**
def player_choice(board):
chosenpos = 0
while True:
try:
chosenpos = int(input("Please, choose a position (provide an integer value from 1 to 9): "))
except:
print("Looks like you did not enter an integer!")
continue
else:
break
while chosenpos not in [1,2,3,4,5,6,7,8,9] or not space_check(board,chosenpos):
chosenpos = int(input("Oops, it looks like this position has already been taken! Please, choose another one (provide an integer value from 1 to 9): "))
return chosenpos
# **Step 9: Write a function that asks the player if they want to play and returns a boolean True if they do want.**
def replay():
yesorno = input("Do you want to play (Yes/No)?")
while yesorno.lower() not in ['yes','no']:
yesorno = input("Hmm! It looks like there is a typo in your previous answer. Please, confirm if you want to play (Yes/No)?")
else:
return yesorno.lower() == "yes"
# **Step 10: Use while loops and the functions you've made to run the game!**
print('Welcome to Tic Tac Toe!')
answer = replay()
while answer:
# Set the game up here
test_board = ['#',' ',' ',' ',' ',' ',' ',' ',' ',' ']
# Choose player
player_input()
# Choose who will be first
firstplayer = choose_first()
if firstplayer in range(1,500000):
player1 = 'X'
player2 = 'O'
else:
player1 = 'O'
player2 = 'X'
playgame = input("Are you ready to play? Yes or No ")
while playgame.lower() not in ['yes','no']:
playgame = input("Are you ready to play? Yes or No ")
if playgame.lower() == "no":
break
while True:
if (full_board_check(test_board) == True) or (win_check(test_board, 'X') == True) or (win_check(test_board, 'O') == True):
break
clear_output()
display_board(test_board)
chosenpos = player_choice(test_board)
place_marker(test_board,player1, chosenpos)
clear_output()
if win_check(test_board, player1):
print (f"Congratulations! {player1} has won!")
print("------Game over-----")
display_board(test_board)
if full_board_check(test_board):
print ("------Game over-----")
print("Nobody has won.")
display_board(test_board)
if (full_board_check(test_board) == True) or (win_check(test_board, 'X') == True) or (win_check(test_board, 'O') == True):
break
# Player2's turn.
clear_output()
display_board(test_board)
chosenpos = player_choice(test_board)
place_marker(test_board,player2, chosenpos)
if win_check(test_board, player2):
print (f"Congratulations! {player2} has won!")
print("------Game over-----")
display_board(test_board)
if full_board_check(test_board):
print("------Game over-----")
print("Nobody has won.")
display_board(test_board)
answer = input("Game is over :) Do you want to play again?")
while answer not in ['yes','no']:
answer = input("Game is over :) Do you want to play again (Yes/No)?")
else:
pass
if answer.lower() == 'no':
break
else:
clear_output()
answer = True
| Tic-tac-toe-game.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Switching Linear Dynamical System Demo
# _Switching Linear Dynamical Systems_ (SLDS) provide a natural way of combining Linear Dynamical Systems with Hidden Markov Models. They allow us to approximate a system that has globally non-linear dynamics by a series linear systems. A good reference for these types of systems is ["Variational Inference for State Space models"](https://www.cs.toronto.edu/~hinton/absps/switch.pdf) by Ghahramani and Hinton.
#
# An LDS comprises $K$ discrete hidden states, which evolve according to a Markov chain. We'll call the hidden state $z$, and use the notation $z_t = k$ to mean that the system is in state $k$ at time $t$. The Markov chain for the hidden state is specified by a state-transition matrix $Q$, where $Q_{ij} = P(z_t = j \mid z_{t-1} = i)$.
#
# ### Generative Model for SLDS
# The generative model for an SLDS combines an HMM with a set of linear dynamical systems as follows. In addition to the discrete state, we have a continuous latent state $x_t \in \mathbb{R}^D$ and an observation $y_t \in \mathbb{R}^N$. Each discrete state $\{1,\ldots,K
# \}$ is associated with a different dynamics matrix $A_k$ and a different measurement matrix $C_k$. Formally, we generate data from an SLDS as follows:
#
# 1. **Discrete State Update**. At each time step, sample a new discrete state $z_t \mid z_{t-1}$ with probabilities given by a Markov chain.
#
# 2. **Continuous State Update**. Update the state using the dynamics matrix corresponding to the new discrete state:
# $$
# x_t = A_k x_{t-1} + V_k u_{t} + b_k + w_t
# $$
# $A_k$ is the dynamics matrix corresponding to discrete state $k$. $u_t$ is the input vector (specified by the user, not inferred by SSM) and $V_k$ is the corresponding control matrix. The vector $b$ is an offset vector, which can drive the dynamics in a particular direction.
# The terms $w_t$ is a noise terms, which perturbs the dynamics.
# Most commonly these are modeled as zero-mean multivariate Gaussians,
# but one nice feature of SSM is that it supports many distributions for these noise terms. See the Linear Dynamical Systems notebook for a list of supported dynamics models.
#
# 3. **Emission**. We now make an observation of the state, according to the specified observation model. In the general case, the state controlls the observation via a Generalized Linear Model:
# $$
# y_t \sim \mathcal{P}(\eta(C_k x_t + d_k + F_k u_t + v_t))
# $$
# $\mathcal{P}$ is a probabibility distribution. The inner arguments form an affine measurement of the state, which is then passed through the inverse link function $\eta(\cdot)$.
# In this case, $C_k$ is the measurement matrix corresponding to discrete state $k$, $d_k$ is an offset or bias term corresponding to discrete state $k$, $F_k$ is called the feedthrough matrix or passthrough matrix (it passes the input directly to the emission). In the Gaussian case, the emission can simply be written as $y_t = C_k x_t + d_k + F_k u_t + v_t$ where $v_t$ is a Gaussian r.v. See the Linear Dynamical System notebook for a list of the observation models supported by SSM.
#
#
#
# ## 1. Setup
# We import SSM as well as a few other utilities for plotting.
# +
import autograd.numpy as np
import autograd.numpy.random as npr
npr.seed(0)
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
# %matplotlib inline
import seaborn as sns
sns.set_style("white")
sns.set_context("talk")
color_names = ["windows blue",
"red",
"amber",
"faded green",
"dusty purple",
"orange",
"clay",
"pink",
"greyish",
"mint",
"cyan",
"steel blue",
"forest green",
"pastel purple",
"salmon",
"dark brown"]
colors = sns.xkcd_palette(color_names)
cmap = ListedColormap(colors)
import ssm
from ssm.util import random_rotation, find_permutation
from ssm.plots import plot_dynamics_2d
save_figures = False
# -
# ## 2. Creating an SLDS and Sampling
# Below, we set some parameters for our SLDS: 5 discrete states, latent state of dimension 2, emissions of dimensions 10. We'll be sampling for 100 time bins for the purpose of visualizing the output of our SLDS.
#
# We then create an SLDS object:
# ```python
# true_slds = ssm.SLDS(emissions_dim,
# n_disc_states,
# latent_dim,
# emissions="gaussian_orthog")
# ```
# We specify the emissions model as `"gaussian_orthog"` which ensures that each measurement matrix $C_k$ will be orthogonal. Because an orthogonal matrix is full-rank, this means that our system is fully observable. In other words, the emissions model does not "losing" information about the state.
#
# The syntax for sampling from an SLDS is the same as for an LDS:
# ```python
# states_z, states_x, emissions = true_lds.sample(time_bins)
# ```
# The sample function for SLDS returns a tuple of (discrete states, continuous states, observations).
#
# Set the parameters of the SLDS
time_bins = 100 # number of time bins
n_disc_states = 5 # number of discrete states
latent_dim = 2 # number of latent dimensions
emissions_dim = 10 # number of observed dimensions
# +
# Make an SLDS with the true parameters
true_slds = ssm.SLDS(emissions_dim,
n_disc_states,
latent_dim,
emissions="gaussian_orthog")
for k in range(n_disc_states):
true_slds.dynamics.As[k] = .95 * random_rotation(latent_dim, theta=(k+1) * np.pi/20)
states_z, states_x, emissions = true_slds.sample(time_bins)
# -
# ### 2.1 Visualize the Latent States
# Below, we visualize the 2-dimensional trajectory of the continuous latent state $x_t$. The different colors correspond to different values of the discrete state variable $z_t$. We can see how the different colors correspond to different dynamics on the latent state.
# +
for k in range(n_disc_states):
curr_states = states_x[states_z == k]
plt.plot(curr_states[:,0],
curr_states[:,1],
'-',
color=colors[k],
lw=3,
label="$z=%i$" % k)
# Draw lines connecting the latent state between discrete state transitions,
# so they don't show up as broken lines.
next_states = states_x[states_z == k+1]
if len(next_states) > 0 and len(curr_states) > 0:
plt.plot((curr_states[-1,0], next_states[0,0]),
(curr_states[-1,1], next_states[0,1]),
'-',
color='gray',
lw=1)
plt.xlabel("$x_1$")
plt.ylabel("$x_2$")
plt.title("Simulated Latent States")
plt.legend(bbox_to_anchor=(1.0,1.0))
plt.show()
plt.figure(figsize=(10,2))
cmap_limited = ListedColormap(colors[0:n_disc_states])
plt.imshow(states_z[None,:], aspect="auto", cmap=cmap_limited)
plt.title("Simulated Discrete Latent States")
plt.yticks([])
plt.xlabel("Time")
plt.show()
# -
# ### 2.1 Visualize the Emissions
# Below, we visualize the 10-dimensional emissions from our SLDS.
# +
plt.figure(figsize=(10, 6))
gs = plt.GridSpec(2, 1, height_ratios=(1, emissions_dim/latent_dim))
# Plot the continuous latent states
lim = abs(states_x).max()
plt.subplot(gs[0])
for d in range(latent_dim):
plt.plot(states_x[:, d] + lim * d, '-k')
plt.yticks(np.arange(latent_dim) * lim, ["$x_{}$".format(d+1) for d in range(latent_dim)])
plt.xticks([])
plt.xlim(0, time_bins)
plt.title("Simulated Latent States")
lim = abs(emissions).max()
plt.subplot(gs[1])
for n in range(emissions_dim):
plt.plot(emissions[:, n] - lim * n, '-')
plt.yticks(-np.arange(emissions_dim) * lim, ["$y_{{ {} }}$".format(n+1) for n in range(emissions_dim)])
plt.xlabel("time")
plt.xlim(0, time_bins)
plt.title("Simulated emissions")
plt.tight_layout()
if save_figures:
plt.savefig("lds_2.pdf")
# -
# ## 3. Fit an SLDS From Data
# SSM provides the capability to learn the parameters of an SLDS from data. In the above cells, we sampled from 100 time-steps in order to visualize the state trajectory.
#
# In order to learn an SLDS, we'll need more data, so we start by sampling for a longer period. In the following cells, we'll treat our observations as a dataset, and demonstrate how to learn an SLDS using SSM.
# Sample again, for more time-bins
time_bins = 1000
states_z, states_x, emissions = true_slds.sample(time_bins)
data = emissions
# ### 3.1 Compare Fitting Methods
#
# **Important Note:**
# <span style="font-size:larger;">
# Understanding the following section is not necessary to use SSM! _For practical purposes, it is almost always best to use the Laplace-EM method, which is the default._
# </span>
#
# **Parameter Learning for SLDS**
# Parameter learning in an SLDS requires approximate methods. SSM provides two approximate inference algorithms: Stochastic Variational Inference (`"svi"`), Laplace-Approximate EM (`"laplace_em"`). We don't have the space to describe these methods in detail here, but Stochastic Variational Inference was described in ["Stochastic Variational Inference"](http://www.columbia.edu/~jwp2128/Papers/HoffmanBleiWangPaisley2013.pdf) by Hoffamn et al. The Laplace Approximation is described in several sources, but a good reference for the context of state-space models is ["Estimating State and Parameters in state-space models of Spike Trains,"](https://pdfs.semanticscholar.org/a71e/bf112cabd47cc67284dc8c12ab7644195d60.pdf) a book chapter by Macke et al.
#
#
#
# **Approximate Posterior Distributions**
# When using approximate methods, we must choose the form of the distribution we use to approximate the posterior. Here, SSM provides three options:
# 1. `variational_posterior="meanfield"`
# The mean-field approximation uses a factorized distribution as the approximating posterior. Compatible with the SVI method.
#
# 2. `variational_posterior="tridiag"`
# This approximates the posterior using a Gaussian with a block tridiagonal covariance matrix, which can be thought of as approximating the SLDS posterior with the posterior from an LDS. Compatible with the SVI method.
#
# 3. `variational_posterior="structured_meanfield"`
# This assumes a posterior where the join distribution over the continuous and discrete latent states factors as follows. If $q(z,x \mid y)$ is the joint posterior of the discrete and continuous states given the data, we use the approximation $q(z,x \mid y) \approx q(z \mid y)q(x \mid y)$, where $q(z \mid y)$ is the posterior for a Markov chain. Compatible with the SVI and Laplace-EM methods.
#
# **Calling the Fit function in SSM**
# All models in SSM share the same general syntax for fitting a model from data. Below, we call the fit function using three different methods and compare convergence. The syntax is as follows:
# ```python
# elbos, posterior = slds.fit(data, method= "...",
# variational_posterior="...",
# num_iters= ...)
# ```
# In the the call to `fit`, method should be one of {`"svi"`, `"laplace_em"`}.
# The `variational_posterior` argument should be one of {`"mf"`, `"structured_meanfield"`}. However, when using Laplace-EM _only_ structured mean field is supported.
# Below, we fit using four methods, and compare convergence.
# **Fit using BBVI and Mean-Field Posterior**
# +
print("Fitting SLDS with BBVI and Mean-Field Posterior")
# Create the model and initialize its parameters
slds = ssm.SLDS(emissions_dim, n_disc_states, latent_dim, emissions="gaussian_orthog")
# Fit the model using BBVI with a mean field variational posterior
q_mf_elbos, q_mf = slds.fit(data, method="bbvi",
variational_posterior="mf",
num_iters=1000)
# Get the posterior mean of the continuous states
q_mf_x = q_mf.mean[0]
# Find the permutation that matches the true and inferred states
slds.permute(find_permutation(states_z, slds.most_likely_states(q_mf_x, data)))
q_mf_z = slds.most_likely_states(q_mf_x, data)
# Smooth the data under the variational posterior
q_mf_y = slds.smooth(q_mf_x, data)
# -
# **Fit using BBVI and Structured Variational Posterior**
# +
print("Fitting SLDS with BBVI using structured variational posterior")
slds = ssm.SLDS(emissions_dim, n_disc_states, latent_dim, emissions="gaussian_orthog")
# Fit the model using SVI with a structured variational posterior
q_struct_elbos, q_struct = slds.fit(data, method="bbvi",
variational_posterior="tridiag",
num_iters = 1000)
# Get the posterior mean of the continuous states
q_struct_x = q_struct.mean[0]
# Find the permutation that matches the true and inferred states
slds.permute(find_permutation(states_z, slds.most_likely_states(q_struct_x, data)))
q_struct_z = slds.most_likely_states(q_struct_x, data)
# Smooth the data under the variational posterior
q_struct_y = slds.smooth(q_struct_x, data)
# -
# **Fit using Laplace-EM**
# +
print("Fitting SLDS with Laplace-EM")
# Create the model and initialize its parameters
slds = ssm.SLDS(emissions_dim, n_disc_states, latent_dim, emissions="gaussian_orthog")
# Fit the model using Laplace-EM with a structured variational posterior
q_lem_elbos, q_lem = slds.fit(data, method="laplace_em",
variational_posterior="structured_meanfield",
num_iters=100, alpha=0.0)
# Get the posterior mean of the continuous states
q_lem_x = q_lem.mean_continuous_states[0]
# Find the permutation that matches the true and inferred states
slds.permute(find_permutation(states_z, slds.most_likely_states(q_lem_x, data)))
q_lem_z = slds.most_likely_states(q_lem_x, data)
# Smooth the data under the variational posterior
q_lem_y = slds.smooth(q_lem_x, data)
# -
# Plot the ELBOs
plt.plot(q_mf_elbos, label="SVI: Mean-Field Posterior")
plt.plot(q_struct_elbos, label="SVI: Block-Tridiagonal Structured Posterior")
plt.plot(q_lem_elbos, label="Laplace-EM: Structured Mean-Field Posterior")
plt.xlabel("Iteration")
plt.ylabel("ELBO")
plt.legend(bbox_to_anchor=(1.0,1.0))
plt.title("Convergence for learning an SLDS")
plt.show()
# ### 3.2 Exercise: The Evidence Lower Bound (ELBO)
# In the SLDS model (and even in the LDS case with non-Gaussian observations), we can't optimize the log-likelihood directly. Instead, we optimize a lower bound on the log likelihood called the Evidence Lower Bound (ELBO).
#
# We denote the parameters of the model as $\Theta$, which are considered fixed for the purposes of this exercise. Concretely, we need to find a lower bound on $\log(P(Y \mid \Theta))$ where $Y=[y_1,\ldots,y_T]$. Can you use Jensen's inequality to derive a lower bound on this likelihood?
# ## 4. Visualize True and Inferred Latent States
# Below, we compare how well each fitting algorithm recovers the discrete latent states. We then inspect the true vs. inferred continuos latent states.
# +
# Plot the true and inferred states
titles = ["True", "Laplace-EM", "SVI with Structured MF", "SVI with MF"]
states_list = [states_z, q_lem_z, q_struct_z, q_mf_z]
fig, axs = plt.subplots(4,1, figsize=(8,6))
for (i, ax, states) in zip(range(len(axs)), axs, states_list):
ax.imshow(states[None,:], aspect="auto", cmap=cmap_limited)
ax.set_yticks([])
ax.set_title(titles[i])
if i < (len(axs) - 1):
ax.set_xticks([])
plt.suptitle("True and Inferred States for Different Fitting Methods", va="baseline")
plt.tight_layout()
# +
title_str = ["$x_1$", "$x_2$"]
fig, axs = plt.subplots(2,1, figsize=(14,4))
for (d, ax) in enumerate(axs):
ax.plot(states_x[:,d] + 4 * d, '-', color=colors[0], label="True" if d==0 else None)
ax.plot(q_lem_x[:,d] + 4 * d, '-', color=colors[2], label="Laplace-EM" if d==0 else None)
ax.set_yticks([])
ax.set_title(title_str[d], loc="left", y=0.5, x=-0.03)
axs[0].set_xticks([])
axs[0].legend(loc="upper right")
plt.suptitle("True and Inferred Continuous States", va="bottom")
plt.tight_layout()
# -
# ### 4.2 Exercise: Fitting with fewer datapoints
# From the above plots, it seems we were able to match the discrete states quite well using our learned model. Try reducing the number of time-bins used for fitting from 1000 to 500 or 100. At what point do we begin to fit badly?
# ## 5. Inference on unseen data
# After learning a model from data, a common use-case is to compute the distribution over latent states given some new observations. For example, in the case of a simple LDS, we could use the Kalman Smoother to estimate the latent state trajectory given a set of observations.
#
# In the case of an SLDS (or Recurrent SLDS), the posterior over latent states can't be computed exactly. Instead, we need to live with a variational approximation to the true posterior. SSM allows us to compute this approximation using the `SLDS.approximate_posterior()` method.
#
# In the below example, we generate some new data from the true model. We then use the `approximate_posterior()` function to estimate the continuous and discrete states.
# +
# Generate data which was not used for fitting
time_bins = 100
data_z, data_x, data = true_slds.sample(time_bins)
# Compute the approximate posterior over latent and continuous
# states for the new data under the current model parameters.
elbos, posterior = slds.approximate_posterior(data,
method="laplace_em",
variational_posterior="structured_meanfield",
num_iters=50)
# Verify that the ELBO increases during fitting. We don't expect a substantial increase:
# we are updating the estimate of the latent states but we are not changing model params.
plt.plot(elbos)
plt.xlabel("Iteration")
plt.ylabel("ELBO")
plt.show()
# -
# **Estimating Latent States**
#
# `posterior` is now an `ssm.variational.SLDSStructuredMeanFieldVariationalPosterior` object. Using this object, we can estimate the continuous and discrete states just like we did after calling the fit function.
#
# In the below cell, we get the estimated continuous states as follows:
# ```python
# posterior_x = posterior.mean_continuous_states[0]
# ```
# This line uses the `mean_continuous_states` property of the posterior object, which returns a list, where each entry of the list corresponds to a single trial of data. Since we have only passed in a single trial the list will have length 1, and we take the first entry.
#
# We then permute the discrete and continuous states to best match the ground truth. This is for aesthetic purposes when plotting. The following lines compute the best permutation which match the predicted states (`most_likely`) to the ground truth discrete states (`data_z`). We then permute the states of the SLDS accordingly:
# ```python
#
# most_likely = slds.most_likely_states(posterior_x, data)
# perm = find_permutation(data_z, most_likely)
# slds.permute(perm)
# z_est = slds.most_likely_states(posterior_x, data)
#
# ```
#
#
# +
# Get the posterior mean of the continuous states
posterior_x = posterior.mean_continuous_states[0]
# Find the permutation that matches the true and inferred states
most_likely = slds.most_likely_states(posterior_x, data)
perm = find_permutation(data_z, most_likely)
slds.permute(perm)
z_est = slds.most_likely_states(posterior_x, data)
# +
# Plot the true and inferred states
titles = ["True", "Estimated"]
states_list = [data_z, z_est]
fig, axs = plt.subplots(2,1, figsize=(6,4))
for (i, ax, states) in zip(range(len(axs)), axs, states_list):
ax.imshow(states[None,:], aspect="auto", cmap=cmap_limited)
ax.set_yticks([])
ax.set_title(titles[i])
if i < (len(axs) - 1):
ax.set_xticks([])
plt.suptitle("True and Inferred States using Structured Meanfield Posterior", va="baseline")
plt.tight_layout()
# -
x_est = posterior.mean_continuous_states[0]
# +
title_str = ["$x_1$", "$x_2$"]
fig, axs = plt.subplots(2,1, figsize=(14,4))
for (d, ax) in enumerate(axs):
ax.plot(data_x[:,d] + 4 * d, '-', color=colors[0], label="True" if d==0 else None)
ax.plot(x_est[:,d] + 4 * d, '-', color=colors[2], label="Laplace-EM" if d==0 else None)
ax.set_yticks([])
ax.set_title(title_str[d], loc="left", y=0.5, x=-0.03)
axs[0].set_xticks([])
axs[0].legend(loc="upper right")
plt.suptitle("True and Estimated Continuous States", va="bottom")
plt.tight_layout()
| notebooks/3 Switching Linear Dynamical System.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def sockMerchant(n, ar):
count = {}
sum = 0
for i in ar:
count.setdefault(i, 0)
count[i] += 1
for i in count.values():
sum += (i//2)
return sum
| hacker-rank/Algorithms/Implementation/Sock Merchant.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.12 ('aiffel_3.8')
# language: python
# name: python3
# ---
# # 12. 사이킷런으로 구현해 보는 머신러닝
#
# **머신러닝의 다양한 알고리즘에 대해 알아보고 사이킷런 라이브러리 사용법을 익힙니다. 사이킷런에서 제공하는 모듈을 이해하고, 머신러닝에 적용해 봅니다.**
# ## 12-1. 들어가며
# ## 12-2. 머신러닝 알고리즘
# ## 12-3. 사이킷런에서 가이드하는 머신러닝 알고리즘
# ## 12-4. Hello Scikit-learn
# ```bash
# $ pip install scikit-learn
# ```
import sklearn
print(sklearn.__version__)
# ## 12-5. 사이킷런의 주요 모듈 (1) 데이터 표현법
# ## 12-6. 사이킷런의 주요 모듈 (2) 회귀 모델 실습
import numpy as np
import matplotlib.pyplot as plt
r = np.random.RandomState(10)
x = 10 * r.rand(100)
y = 2 * x - 3 * r.rand(100)
plt.scatter(x,y)
x.shape
y.shape
from sklearn.linear_model import LinearRegression
model = LinearRegression()
model
# ! 에러 발생
model.fit(x, y)
X = x.reshape(100,1)
model.fit(X,y)
x_new = np.linspace(-1, 11, 100)
X_new = x_new.reshape(100,1)
y_new = model.predict(X_new)
X_ = x_new.reshape(-1,1)
X_.shape
# +
from sklearn.metrics import mean_squared_error
# 직접 구현해 보세요
error = np.sqrt(mean_squared_error(y,y_new))
print(error)
# -
# ```python
# # 정답 코드
#
# error = np.sqrt(mean_squared_error(y,y_new))
# ```
plt.scatter(x, y, label='input data')
plt.plot(X_new, y_new, color='red', label='regression line')
# ## 12-7. 사이킷런의 주요 모듈 (3) datasets 모듈
from sklearn.datasets import load_wine
data = load_wine()
type(data)
print(data)
data.keys()
data.data
data.data.shape
data.data.ndim
data.target
data.target.shape
data.feature_names
len(data.feature_names)
data.target_names
print(data.DESCR)
# ## 12-8. 사이킷런의 주요 모듈 (4) 사이킷런 데이터셋을 이용한 분류 문제 실습
# +
import pandas as pd
pd.DataFrame(data.data, columns=data.feature_names)
# -
X = data.data
y = data.target
from sklearn.ensemble import RandomForestClassifier
model = RandomForestClassifier()
model.fit(X, y)
y_pred = model.predict(X)
# +
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
#타겟 벡터 즉 라벨인 변수명 y와 예측값 y_pred을 각각 인자로 넣습니다.
print(classification_report(y, y_pred))
#정확도를 출력합니다.
print("accuracy = ", accuracy_score(y, y_pred))
# -
# ## 12-9. 사이킷런의 주요 모듈 (5) Estimator
# ## 12-10. 훈련 데이터와 테스트 데이터 분리하기
from sklearn.datasets import load_wine
data = load_wine()
print(data.data.shape)
print(data.target.shape)
X_train = data.data[:142]
X_test = data.data[142:]
print(X_train.shape, X_test.shape)
y_train = data.target[:142]
y_test = data.target[142:]
print(y_train.shape, y_test.shape)
from sklearn.ensemble import RandomForestClassifier
model = RandomForestClassifier()
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
# +
from sklearn.metrics import accuracy_score
print("정답률=", accuracy_score(y_test, y_pred))
# +
from sklearn.model_selection import train_test_split
result = train_test_split(X, y, test_size=0.2, random_state=42)
# -
print(type(result))
print(len(result))
result[0].shape
result[1].shape
result[2].shape
result[3].shape
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
# +
# 데이터셋 로드하기
# [[your code]
from sklearn.datasets import load_wine
from sklearn.ensemble import RandomForestClassifier
data = load_wine()
# 훈련용 데이터셋 나누기
# [[your code]
X = data.data
y = data.target
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
# 훈련하기
# [[your code]
model = RandomForestClassifier()
model.fit(X_train, y_train)
# 예측하기
# [[your code]
y_pred = model.predict(X_test)
print(classification_report(y_test, y_pred))
# 정답률 출력하기
# [[your code]
print("accuracy = ", accuracy_score(y_test, y_pred))
# -
# ```python
# # 정답 코드
#
# # 데이터셋 로드하기
# data = load_wine()
# # 훈련용 데이터셋 나누기
# X_train, X_test, y_train, y_test = train_test_split(data.data, data.target, test_size=0.2)
# # 훈련하기
# model = RandomForestClassifier()
# model.fit(X_train, y_train)
# # 예측하기
# y_pred = model.predict(X_test)
# # 정답률 출력하기
# print("정답률=", accuracy_score(y_test, y_pred))
# ```
# ## 12-11. 마무리
| FUNDAMENTALS/Node_12/[F-12] Only_LMS_Code_Blocks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import warnings
from sklearn.exceptions import ConvergenceWarning
warnings.filterwarnings(action='ignore', category=ConvergenceWarning)
from datetime import datetime
from sklearn.svm import SVC
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
from sklearn import datasets, svm, metrics
from sklearn.metrics import f1_score
from sklearn.multiclass import OneVsRestClassifier
# ## Read MNIST data
# +
import os
import struct
import numpy as np
import matplotlib.pyplot as pyplot
"""
Loosely inspired by http://abel.ee.ucla.edu/cvxopt/_downloads/mnist.py
which is GPL licensed.
"""
def read(dataset = "training", path = "."):
"""
Python function for importing the MNIST data set. It returns an iterator
of 2-tuples with the first element being the label and the second element
being a numpy.uint8 2D array of pixel data for the given image.
"""
if dataset is "training":
fname_img = os.path.join(path, 'train-images.idx3-ubyte')
fname_lbl = os.path.join(path, 'train-labels.idx1-ubyte')
elif dataset is "testing":
fname_img = os.path.join(path, 't10k-images.idx3-ubyte')
fname_lbl = os.path.join(path, 't10k-labels.idx1-ubyte')
else:
raise Exception("dataset must be 'testing' or 'training'")
# Load everything in some numpy arrays
with open(fname_lbl, 'rb') as flbl:
magic, num = struct.unpack(">II", flbl.read(8))
lbl = np.fromfile(flbl, dtype=np.int8)
with open(fname_img, 'rb') as fimg:
magic, num, rows, cols = struct.unpack(">IIII", fimg.read(16))
img = np.fromfile(fimg, dtype=np.uint8).reshape(len(lbl), rows, cols)
return img, lbl
# -
path = 'MNIST/'
X_train, y_train = read("testing", path)
X_test, y_test = read("training", path)
X_train = X_train.reshape(X_train.shape[0], -1)
X_test = X_test.reshape(X_test.shape[0], -1)
print(X_train.shape)
print(y_train.shape)
print(X_test.shape)
print(y_test.shape)
# ## Preprocessing with StandardScaler
# +
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler(copy=True, with_mean=True, with_std=True)
StandardScaler(copy=True, with_mean=True, with_std=True)
scaler.fit(X_train)
X_train_scaled = scaler.transform(X_train)
X_test_scaled = scaler.transform(X_test)
# -
# ## SVM training
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import warnings
from sklearn.exceptions import ConvergenceWarning
warnings.filterwarnings(action='ignore', category=ConvergenceWarning)
from datetime import datetime
from sklearn.svm import SVC
from sklearn.model_selection import cross_val_score
from sklearn import datasets, svm, metrics
from sklearn.metrics import f1_score
from sklearn.preprocessing import StandardScaler
# Preprocessing with StandardScaler
scaler = StandardScaler(copy=True, with_mean=True, with_std=True)
StandardScaler(copy=True, with_mean=True, with_std=True)
scaler.fit(X_train)
X_train_scaled = scaler.transform(X_train)
X_test_scaled = scaler.transform(X_test)
# hyper_parameter
C = [0.001, 0.01, 0.1, 1.0, 10.0, 100.0, 1000.0]
df = pd.DataFrame(columns=['C value', 'f1 score', 'learning time'])
bookmark = 0
for c in C:
# Model Instance
classifier = svm.SVC(kernel = 'linear',
C = c,
decision_function_shape= 'ovr',
random_state=108)
# fit, prediction
clf = classifier.fit(X_train_scaled, y_train)
y_pred = clf.predict(X_test_scaled)
score = cross_val_score(classifier, X_train_scaled, y_train, cv=5, scoring='f1_macro')
df.loc[bookmark] = [c, score, end_time - begin_time]
print(df.loc[bookmark])
df.to_csv("SVC(linear)_ovr.csv", mode='w')
bookmark = bookmark + 1
print('=======================================')
df.to_csv("SVC(linear)_ovr.csv", mode='w')
# -
# ## Plot
# +
f1_score = [0.11, 0.65, 0.88, 0.93, 0.94, 0.94]
c = [0.001, 0.01, 0.1, 1.0, 10, 1000]
plt.title("relation between C and f1_score")
plt.plot(c, f1_score)
plt.grid()
plt.xscale('log')
plt.ylabel('f1_score')
plt.xlabel('Parameter C')
plt.ylim(0, 1.1)
plt.tight_layout()
plt.show()
# +
f1_score = [0.11, 0.65, 0.88, 0.93, 0.94, 0.94]
c = [0.001, 0.01, 0.1, 1.0, 10, 1000]
plt.title("relation between C and f1_score")
plt.plot(c, f1_score)
plt.grid()
plt.xscale('log')
plt.ylabel('f1_score')
plt.xlabel('Parameter C')
plt.ylim(0, 1.1)
plt.tight_layout()
plt.show()
| study/SVM_SVC/SVC(Linear kernel).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 20: Introduction to Machine Learning
#
# CSCI 1360E: Foundations for Informatics and Analytics
# + [markdown] slideshow={"slide_type": "slide"}
# ## Overview and Objectives
# + [markdown] slideshow={"slide_type": "-"}
# We've covered statistics, probability, linear algebra, data munging, and the basics of Python. You're ready. By the end of this lecture, you should be able to
#
# - Define machine learning, optimization, and artificial intelligence, and how these subfields interact.
# - Understand when to use supervised versus unsupervised learning.
# - Create a basic classifier.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Part 1: Machine Learning
# + [markdown] slideshow={"slide_type": "-"}
# What, exactly, is machine learning?
# + [markdown] slideshow={"slide_type": "-"}
# From <NAME> in 1959: Machine Learning is a
#
# > "...Field of study that gives computers the ability to learn without being explicitly programmed."
#
# Probably the most succinct definition.
# + [markdown] slideshow={"slide_type": "slide"}
# Imagine how you first learned what a "cat" was when you were young. The process probably went something like this:
# + [markdown] slideshow={"slide_type": "-"}
# - You pointed at a cat and asked "What is that?"
# + [markdown] slideshow={"slide_type": "-"}
# - Somebody (probably a parent) said "That's a cat."
# + [markdown] slideshow={"slide_type": "slide"}
# You internalized this experience. Perhaps later, you pointed at a dog and asked "Cat?"
# + [markdown] slideshow={"slide_type": "-"}
# - Somebody (again, probably a parent) corrected you, saying "No, that's a dog."
# + [markdown] slideshow={"slide_type": "-"}
# - With some more back and forth, you were able to reliably determine on your own whether or not something in the real world was a cat.
# + [markdown] slideshow={"slide_type": "slide"}
# The first process, in which you asked for and received feedback, is the *learning* step ("what is that?").
# + [markdown] slideshow={"slide_type": "-"}
# The second process, in which you took your experience and identified cats without any feedback, is the *generalization* step ("I think that's a cat").
# + [markdown] slideshow={"slide_type": "fragment"}
# **These two steps define a machine learning algorithm.**
# + [markdown] slideshow={"slide_type": "slide"}
# Some other ways of describing machine learning include
# + [markdown] slideshow={"slide_type": "-"}
# - Inferring knowledge from data
# - Generalizing to unseen or unobserved data
# - Emphasizing the computational challenges (e.g., "big" data)
# + [markdown] slideshow={"slide_type": "-"}
# There's a whole ecosystem of buzzwords around machine learning; we'll only explore a very small subset in this lecture.
#
# 
# + [markdown] slideshow={"slide_type": "slide"}
# Most machine learning algorithms involve the development of a *rule* to describe the data.
# + [markdown] slideshow={"slide_type": "fragment"}
# This rule is sometimes referred to as a *hypothesis* or a *decision boundary*.
#
# 
# + [markdown] slideshow={"slide_type": "-"}
# The way in which this rule is formed, and the resulting shape of the decision boundary, is largely what distinguishes one machine learning algorithm from another.
# + [markdown] slideshow={"slide_type": "slide"}
# Choosing what algorithm to use is important.
# + [markdown] slideshow={"slide_type": "-"}
# - Different algorithms embody and quantify different assumptions about the underlying data.
# + [markdown] slideshow={"slide_type": "-"}
# - A prevalent heuristic throughout machine learning is to use the simplest possible model or algorithm that adequately explains the data.
# + [markdown] slideshow={"slide_type": "-"}
# - Using extremely complex algorithms will almost always give you lower error rates, but this can potentially lead to *overfitting*: a situation where your algorithm so closely matches the data that it can't generalize to new data.
# + [markdown] slideshow={"slide_type": "slide"}
# **Overfitting** is a problem in machine learning where your model performs exceptionally well on your training data, but fits so tightly to it that it cannot generalize well to new data.
# -
# Imagine if, in learning what a "cat" was, you only observed a single species, like Siamese. You might think, therefore, that any non-Siamese cat you saw actually wasn't a cat at all, because your internal representation was *overfit* to Siamese.
# + [markdown] slideshow={"slide_type": "fragment"}
# Typically, when you build a machine learning pipeline, you separate your data into [at least] two subsets:
# + [markdown] slideshow={"slide_type": "-"}
# - *Training data*, which you use to construct your machine learning model. If you are building a classifier, this will also contain the data to which you know what "class" it belongs (e.g. if you're building a spam detector, your training data will include emails for which you know whether or not they are spam)
# + [markdown] slideshow={"slide_type": "-"}
# - *Testing data*, which you will use to test the model constructed in the training step. This data may or may not also have "class" labels; if so, it serves to simulate how your model would perform in the "real world."
# + [markdown] slideshow={"slide_type": "slide"}
# **Evaluating your model is crucial to good model design.** Performing this training-testing split is one step to avoid the problems of overfitting, whereby your model tunes itself so tightly to training data that it cannot generalize to any new information.
# + [markdown] slideshow={"slide_type": "fragment"}
# This can be mitigated through the use of *k-fold cross-validation*.
#
# 
# + [markdown] slideshow={"slide_type": "-"}
# Cross-validation is a process through which the data you use to train your model is split into two sets, or *folds*: *training* and *testing*. The model is trained using the training set, and then is tested for generalization accuracy with the testing set.
# + [markdown] slideshow={"slide_type": "-"}
# This process is then repeated with new training and testing sets. The number of times this process is repeated is the number $k$ in **k**-fold.
# + [markdown] slideshow={"slide_type": "slide"}
# Another important problem to keep in mind when designing your algorithms is the concept of the **bias-variance tradeoff**.
# + [markdown] slideshow={"slide_type": "-"}
# - **Variance** is a concept we've seen before; it's related directly to the standard deviation. This quantifies how much a distribution varies from its mean, or average. Small variance means a distribution is concentrated around its mean; large variance means the distribution is very spread out.
# + [markdown] slideshow={"slide_type": "-"}
# - **Bias** is a term we've all heard as a colloquialism, but which has a very specific statistical definition: this quantifies the difference between the expected mean of a random variable, and the true mean of that random variable.
# + [markdown] slideshow={"slide_type": "-"}
# In statistical terms, neither of these quantities is inherently "bad"; this is especially true with bias, as in daily language use the term tends to be a pejorative. However, it is critical to understand the effects these quantities can have on any modeling strategy you employ in machine learning.
# + [markdown] slideshow={"slide_type": "slide"}
# This picture perfectly describes the effects of the two quantities:
# + [markdown] slideshow={"slide_type": "-"}
# 
# + [markdown] slideshow={"slide_type": "slide"}
# In an ideal world, we'd all build machine learning models with **low variance and low bias** (upper left corner).
# + [markdown] slideshow={"slide_type": "-"}
# Unfortunately, this is rarely, if ever, possible. In fact, we can explicitly decompose the error rates of virtually any machine learning algorithm into three distinct components: the bias, the variance, and the irreducible error (noise in the problem itself).
# + [markdown] slideshow={"slide_type": "-"}
# The **tradeoff** comes from the fact that, as we try to *decrease* the error of one of the terms, we often simultaneously *increase* the error from the other term.
# + [markdown] slideshow={"slide_type": "-"}
# Different machine learning algorithms and models will make different assumptions and therefore give different preferences to bias versus variance. The specifics of the problem on which you are working will dictate whether your model can tolerate high-bias or high-variance.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Part 2: Supervised and Unsupervised Learning
# + [markdown] slideshow={"slide_type": "-"}
# Learning algorithms can, at a coarse level, be divided into two distinct groups: *supervised* and *unsupervised* learning.
# + [markdown] slideshow={"slide_type": "fragment"}
# - **Supervised learning** involves training a model using labeled data, or data for which you know the answer. For example, if you wanted to design an automatic spam filter, you might start with a training dataset containing sample emails that are labeled as either spam or not spam. This is the equivalent of your parent figure providing the correct answer regarding your inquiries about what is a "cat."
# + [markdown] slideshow={"slide_type": "-"}
# - **Unsupervised learning** is the process of building a model using data for which you have no ground truth and no explicit set of labels for your data. Usually this implies *clustering*: the process of grouping similar things together.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Supervised Learning
# + [markdown] slideshow={"slide_type": "-"}
# The basic idea behind supervised learning is that you start with a ground-truth, labeled dataset of $N$ points with the form
# + [markdown] slideshow={"slide_type": "-"}
# $\{(\vec{x}_1, y_1), (\vec{x}_2, y_2), ..., (\vec{x}_N, y_N)\}$
# + [markdown] slideshow={"slide_type": "-"}
# where $\vec{x}_i$ is the $i^{th}$ data point, and $y_i$ is the ground-truth label (e.g. "spam" or "not spam" for email, or "cat" or "not a cat" for learning what a cat is) of that data point.
# + [markdown] slideshow={"slide_type": "fragment"}
# The goal, then, is to use this training data to learn a function $f$ that most accurately maps the training data $X$ to the correct labels $Y$. In mathematical parlance, this looks like
# + [markdown] slideshow={"slide_type": "-"}
# $f : X \rightarrow Y$
# + [markdown] slideshow={"slide_type": "slide"}
# Within the branch of supervised learning, there are two distinct strategies: *classification* and *regression*.
# + [markdown] slideshow={"slide_type": "fragment"}
# - **Classification** is the process of mapping an input data point $\vec{x}$ to a discrete label. Spam filters would fall in thsi category: we are explicitly trying to identify an incoming email as either "spam" or "not spam." In mathematical terms: we're mapping continuous input to one of a handful of discrete outputs.
# + [markdown] slideshow={"slide_type": "-"}
# - **Regression** is the process of mapping an input data point $\vec{x}$ to a real-valued label. Linear regression, in which the goal is to find a best-fit line to a cloud of data points, is an example. Rather than a handful of possible discrete outputs, your output is instead a continuous floating-point value. An example of regression might be predicting the stock value of a company.
# + [markdown] slideshow={"slide_type": "-"}
# In both instances of supervised learning, we're still concerned with mapping inputs to certain correct outputs. The difference is the form of the output: **discrete (one of a possible handful) for classification tasks**, and **continuous (real-valued; floating-point) for regression tasks**.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Unsupervised Learning
# + [markdown] slideshow={"slide_type": "-"}
# Unlike supervised learning, tasks in unsupervised learning deal strictly with unlabeled data of $N$ points of the form
# + [markdown] slideshow={"slide_type": "-"}
# $\{\vec{x}_1, \vec{x}_2, ..., \vec{x}_N\}$
# + [markdown] slideshow={"slide_type": "-"}
# Since all we have are the data, the goal of unsupervised learning is to find a model which best explains the underlying process that gave rise to the data.
# + [markdown] slideshow={"slide_type": "slide"}
# One example of unsupervised learning is the identification of communities in social networks.
# + [markdown] slideshow={"slide_type": "-"}
# A social network is well-represented using the graph structure of nodes connected by edges, where each user (you and me) is a node in the graph, and any users who are "friends" are connected by an edge.
# + [markdown] slideshow={"slide_type": "-"}
# Community-finding is decidedly an unsupervised process: you have no idea how many communities there are ahead of time, nor do you know who is part of what community. All you have is your database of users and their connections.
# + [markdown] slideshow={"slide_type": "slide"}
# We can use these inter-person connections to identify groups that are highly interconnected with each other, therefore representing a "community." A research paper in 2010 identified such networks of bloggers on different sides of the political spectrum:
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# + [markdown] slideshow={"slide_type": "slide"}
# Other ways to refer to unsupervised learning include
# + [markdown] slideshow={"slide_type": "fragment"}
# - **Clustering**, since we are effectively "clustering" similar data together
# + [markdown] slideshow={"slide_type": "-"}
# - **Anomaly detection**, sort of an antonym for clustering: rather than having the goal of grouping similar things together, we use that as a stepping stone to then try and identify things that *don't* fit
# + [markdown] slideshow={"slide_type": "-"}
# - **Data mining**, though this colloquialism has largely been co-opted by the popular media and may therefore not necessarily equate to unsupervised learning in certain contexts
# + [markdown] slideshow={"slide_type": "-"}
# In all cases, the common thread is that you don't have any kind of ground-truth labeling scheme.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Part 3: Building a Classifier
# + [markdown] slideshow={"slide_type": "-"}
# Let's illustrate some of these concepts by building a classifier for a well-known dataset: the **Iris dataset.**
# + [markdown] slideshow={"slide_type": "-"}
# This dataset was one of the first available for statistical classification, dating back to the 1930s.
# + [markdown] slideshow={"slide_type": "fragment"}
# 
#
# It involves recognizing [Iris flower species](https://en.wikipedia.org/wiki/Iris_(plant)) based on their physical characteristics. Today, these species would be identified using DNA, but in the 1930s DNA had not even been discovered.
# + [markdown] slideshow={"slide_type": "slide"}
# The Iris dataset is comprised of 150 samples of Iris flowers over 3 species, or *classes* (Setosa, Versicolor, and Virginica).
# + [markdown] slideshow={"slide_type": "fragment"}
# Each sample has four attributes associated with it:
# + [markdown] slideshow={"slide_type": "-"}
# - Sepal length
# - Sepal width
# - Petal length
# - Petal width
# + [markdown] slideshow={"slide_type": "-"}
# These attributes are our **features** to describe each sample, and what we'll use in order to **classify** each sample into one of the previous three categories.
# + [markdown] slideshow={"slide_type": "-"}
# Our classification challenge: **If we saw a new flower out in a field and measured its sepal and petal length and width, could we make a good prediction about its species?**
# + [markdown] slideshow={"slide_type": "slide"}
# One reason the dataset is so popular is because it's small (150 data points) and easily visualized (4 dimensions), so we can easily take all possible 2-D combinations of the four features and plot them out to see about building some intuition for the data.
# + slideshow={"slide_type": "fragment"}
# %matplotlib inline
import itertools
import matplotlib.pyplot as plt
import numpy as np
import sklearn.datasets as data
iris = data.load_iris() # The dataset comes back as a dictionary.
features = iris['data']
feature_names = iris['feature_names']
classes = iris['target']
feature_combos = itertools.combinations([0, 1, 2, 3], 2)
for i, (x1, x2) in enumerate(feature_combos):
fig = plt.subplot(2, 3, i + 1)
fig.set_xticks([])
fig.set_yticks([])
for t, marker, c in zip(range(3), ">ox", "rgb"):
plt.scatter(features[classes == t, x1],
features[classes == t, x2],
marker = marker, c = c)
plt.xlabel(feature_names[x1])
plt.ylabel(feature_names[x2])
# + [markdown] slideshow={"slide_type": "slide"}
# This gives us some good intuition! For instance--
# + [markdown] slideshow={"slide_type": "fragment"}
# - The first plot, sepal width vs sepal length, gives us a really good separation of the Setosa (red triangles) from the other two, but a poor separation between Versicolor and Virginica
# + [markdown] slideshow={"slide_type": "-"}
# - In fact, this is a common theme across most of the subplots--we can easily pick two dimensions and get a good separation of Setosa from the others, but separating Versicolor and Virginica may be more difficult
# + [markdown] slideshow={"slide_type": "-"}
# - The best pairings for separating Versicolor and Virginica may be either petal length vs sepal width, or petal width vs sepal width.
# + [markdown] slideshow={"slide_type": "slide"}
# Still, for any given pair of features, we can't get a *perfect* classification rule.
# + [markdown] slideshow={"slide_type": "fragment"}
# **...but that's OK!**
# + [markdown] slideshow={"slide_type": "-"}
# Remember overfitting? Designing a classifier that obtains 100% accuracy on the data we have available may sound great, but what happens when *new* data arrive?
# + [markdown] slideshow={"slide_type": "slide"}
# So! Using these pictures, can we design a very simple classifier?
# -
# We don't need anything sophisticated. Just look at the plots and pick some feature values that seem to separate out the classes pretty well.
# + [markdown] slideshow={"slide_type": "slide"}
# For example, the following code says: if `petal_length` is less than 1.9, it's definitely a Setosa. Otherwise, it performs another conditional: if `petal_width` is less than or equal to 1.6, it predicts Versicolor; otherwise, it predicts Virginica.
#
# And that's it!
# + slideshow={"slide_type": "fragment"}
sepal_length = 0
sepal_width = 1
petal_length = 2
petal_width = 3
setosa = 0
versicolor = 1
virginica = 2
def classifier1(X):
global sepal_length, sepal_width, petal_length, petal_width
global setosa, versicolor, virginica
y_predicted = np.zeros(shape = X.shape[0])
for i, x in enumerate(X):
if x[petal_length] < 1.9: # Definitely a setosa
y_predicted[i] = setosa
else: # Either a versicolor or a virginica
if x[petal_width] <= 1.6:
y_predicted[i] = versicolor
else:
y_predicted[i] = virginica
return y_predicted
# + slideshow={"slide_type": "fragment"}
y_pred = classifier1(features)
correct = (y_pred == classes).sum()
print("{} out of {}, or {:.2f}% accuracy.".format(correct, y_pred.shape[0], (correct / y_pred.shape[0]) * 100))
# + [markdown] slideshow={"slide_type": "fragment"}
# **Congratulations, we've just designed our first classifier!**
# + [markdown] slideshow={"slide_type": "slide"}
# The classifier we designed may seem a little *ad-hoc*, but in fact it's not far off from a well-recognized classification strategy known as **Decision Trees**.
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# + [markdown] slideshow={"slide_type": "slide"}
# Decision trees use a series of "splits" to ultimately perform classification on any new data. They are an extremely powerful classification algorithm...
# + [markdown] slideshow={"slide_type": "-"}
# ...that can be **very prone to overfitting if you're not careful.**
# + [markdown] slideshow={"slide_type": "-"}
# A common strategy used to avoid overfitting when building decision trees is to *prune*: that is, clip the depth of the trees at a certain [admittedly arbitrary] point.
# + [markdown] slideshow={"slide_type": "slide"}
# Some other common classification strategies you may or may not have heard of:
# + [markdown] slideshow={"slide_type": "fragment"}
# - **K-Nearest Neighbors**. This algorithm classifies an unknown data point into the majority class of its K-nearest neighbors (hence the name). If a new, unknown data point appeared, and its 5-nearest neighbors consisted of 4 spam emails and 1 non-spam email, we would classify this new point as *spam*.
# + [markdown] slideshow={"slide_type": "-"}
# - **Naive Bayes**. This is a variant of Bayesian learning, and involves the concept of "conditional independence" in order to vastly simplify the problem.
# + [markdown] slideshow={"slide_type": "-"}
# - **Neural Networks** (or *deep learning*). Neural networks are not a new classification strategy, but they have become extremely popular in the last half decade. They are very powerful, capable of learning almost any concept given sufficient training data. They consist of multiple "layers" of stacked neurons that augment certain signals in the data while squelching others.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Review Questions
#
# Some questions to discuss and consider:
# + [markdown] slideshow={"slide_type": "-"}
# 1: There is such a concept as **semi-supervised learning**. It usually involves using a small quantity of labeled data and a much larger quantity of unlabeled data. Based on the name, and what you learned from this lecture on supervised and unsupervised learning, can you speculate as to how the unlabeled and labeled data might be used in this context?
# + [markdown] slideshow={"slide_type": "-"}
# 2: In many machine learning tasks, it is common to have data that is very high-dimensional; in fact, it is common for the number of dimensions of the data to exceed the number of data points you have. This is referred to as the "curse of dimensionality" and has very real implications for training models that generalize well. Imagine again that you're trying to design a spam filter for email. If words in an email consist of that email's dimensions, explain how this task suffers from the "curse of dimensionality".
# + [markdown] slideshow={"slide_type": "-"}
# 3: I'm interested in training a classifier to perform some supervised task. I have built the model and now want to test it; unfortunately, I have very little labeled data available, only 100 data points. When performing $k$-fold cross-validation to test my model, what $k$ should I choose (large, medium, small) and why?
# + [markdown] slideshow={"slide_type": "-"}
# 4: Does overfitting negatively impact the *learning* phase, or the *generalization* phase? Which phase is more important? Explain.
# + [markdown] slideshow={"slide_type": "-"}
# 5: I've designed a model that achieves zero error due to variance, but therefore has high bias. Is my model *underfitting* or *overfitting*? What if I instead design a model that achieves zero error due to bias, but has high variance? Explain.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Course Administrivia
# + [markdown] slideshow={"slide_type": "-"}
# - **How are Assignments 9 and 10 going?** The former is due **tomorrow evening**, the latter due **Monday evening.** Post questions to the Slack `#questions` channel!
# + [markdown] slideshow={"slide_type": "-"}
# - **Final exam review session Wednesday, July 26 at 10am until 12pm!** The link to the Google Hangouts will be posted in the Slack chat when the review begins. Come if you have any questions!
# + [markdown] slideshow={"slide_type": "-"}
# - **The final exam itself will be Friday, July 28.** It will follow the same format as the midterm--it will be available on JupyterHub all day (midnight to midnight), only this time you can check it out for a duration of 3 hours.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Additional Resources
#
# 1. <NAME> and <NAME>. *Building Machine Learning Systems with Python*. 2013. ISBN-13: 978-1782161400
| lectures/L20.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Scientific Programming II
#
# ### <NAME>
# In this second lesson, we'll apply the core elements of programming languages to a common scientific task: reading a data table, performing analysis on its contents, and saving the results. As we go through these different stages of analysis, we'll note how each task relates to one of the elements from the last lesson (we'll note these as `E1`, for example, for element 1, "a thing").
#
# For this lesson, we'll use two important scientific Python packages. `numpy` is the main package providing numerical analysis functions, and `pandas` is designed to make it easy to work with tabular data.
# Import pandas and set up inline plotting
import pandas as pd
# %matplotlib inline
# ## 1. Reading and examining a data table
#
# Unless you're generating your own data by simulation (as in our previous logistic growth function), most scientific analyses begin with loading an external data set.
#
# For this lesson, we'll use data from the North American Breeding Bird Survey. As part of this survey, volunteers have driven cars along fixed routes once a year for the past forty years, stopping periodically along the way and counting all of the birds that they see when they do. The particular data tables that we'll work with today summarize the number of birds of many different species that were counted along routes in the state of California. The large table contains forty years of data for all sighted species, while the small table is a subset of the large table.
#
# You can download and play with this data yourself at:
#
# <NAME>., <NAME>., <NAME>. 2015. North American Breeding Bird Survey Dataset 1966 - 2014, version 2014.0. U.S. Geological Survey, Patuxent Wildlife Research Center http://www.pwrc.usgs.gov/BBS/RawData/.
#
# __Tip:__ It's often a good idea to take a large data set and extract a small portion of it to use while building and testing your analysis code. Small data sets can be analyzed faster and allow you to see, visually, what the "right answer" should be when you write code to perform analysis. Determining whether your function gives the right answer on a small data set is the core idea behind unit testing, which we'll discuss later.
# +
# You can use the exclamation point symbol (the "bang") to run a shell command
# Let's use cat to see the contents of the small data table
# +
# Read the small table using pandas
# The DataFrame function (E2) in pandas creates a thing (E1) called a DataFrame
# +
# Now let's look at the contents of our data frame "thing" (E1)
# +
# Like other "things" in Python, a data frame is an object
# The object contains methods that operate on it (E2)
# -
# A data frame can be conceptualized as a kind of "thing", like we have above, that we can move around and perform operations on. However, it also shares some characteristics in common with a collection of things (E3) because we can use indexes and slicing to pull out subsets of the data table.
#
# There are two main ways that we can select rows and columns from our table: using the labels for the rows and columns or using numeric indexes for the row and column locations. Below we'll focus on label names - check out the `pandas` help for the method `iloc` to learn about using numeric indexes.
# +
# Look at the table again and think about it as a collection (E3)
# +
# Use the loc method to pull out rows and columns by name
# Like a matrix, the row goes first, then the column
# +
# You can use ranges of names, similar to what we saw before for lists
# +
# You can also use lists of names
# -
# ## 2. Perform analysis
#
# Once we have our data table read in, we generally want to perform some sort of analysis with it. Let's presume that we want to get a count of the mean number of individuals sighted per species in each year. However, we only want the average over the species that were actually sighted in the state that year, ignoring species with counts of zero (this is a fairly common analysis in ecology).
#
# Conceptually, one way to approach this problem is to imagine looping through (E4) all of the years, that is the columns of the data frame, one by one. For each year, we want to count the number of species present, sum their counts, and divide the sum of the counts by the number of species seen. We should record this information in some other sort of collection (E3) - we'll use another data frame.
# +
# First, let's set up a new data frame to hold the result of our calculation
# We'll get the column names from the bird table, then use DataFrame to make a new df
# +
# Next, let's figure out how we would do our analysis for one year, say 2010
# +
# Our final calculation code could look like this
# -
# ### Exercise 1 - Calculating mean counts for every year
#
# 1. Put the above calculation code into a `for` loop (E4) that loops over all years, calculating the mean count of birds per species present each year, and stores the result in a new empty data frame.
# 2. Put all of the code that you just wrote into a new function (E6) that takes, as an argument, a data frame of bird counts (like `bird_sm`) and returns the result data frame. Test it with `bird_sm` to make sure that it works.
#
# __Bonus:__
#
# 1. Using label-based indexing, create a new data frame that has the same years but only includes these three species: Spotted Owl, Barred Owl, Great Gray Owl. Try running your function using this new smaller data frame and look at the results. Do you see a result that you may not want?
# 2. Add an `if-else` statement (E5) that checks for the problem that you just uncovered and takes some reasonable action when it occurs.
# ## 3. Save the results
#
# Now that we've managed to generate some useful results, we want to save them somewhere on our computer for later use. There are two broad types of outputs that we might want to save, tables and plots, and we'll use the built-in methods for data frames to do both.
#
# Getting a plot to look just right can take a very long time. Here we'll just use the pandas default styles. For more help on plotting, have a look at the extra lesson on `matplotlib`.
# +
# First we make sure that we've saved our results table
# +
# Data frames have a method to save themselves as a csv file - easy!
# +
# Data frames also have a method to plot their contents
# There's one trick though - by default they put the rows on the x axis and columns on the y
# We want the reverse, so we need to transpose our data frame before plotting it
# +
# With a few extra steps, we can save the plot
# This code looks strange, since we haven't talked about the details of matplotlib
# At this stage, it's best to just use it as a recipe
# -
# ### Exercise 2 - A complete analysis
#
# 1. Using all of the code that we wrote above, put the following lines of code in the cell below (this will form a complete analysis that would run without the rest of this notebook):
# - Import the pandas package
# - Read the `birds_sm.csv` table
# - Define a function to perform the analysis (just copy the one you wrote in Exercise 1)
# - Use that function to make a results dataframe for the `birds_sm.csv` data
# - Saves the resulting table as `birds_results.csv`
# - Saves a plot of the result as `birds_results.pdf`
# 2. To test that your cell works on its own, go to the Jupyter menu bar, under Kernel, and choose "Restart Kernel". This will restart your notebook, so that everything that you've run so far (all the variables stored in memory, in particular) is erased. Run the cell below, and make sure it works correctly.
# 3. Instead of `bird_sm.csv`, make your cell use `bird_lg.csv` and see what the saved results look like. If necessary, modify your code and variable names so that all you have to do is change two letters (`sm` to `lg`) in one place in the code to make this change.
| lessons/python/python2-student.ipynb |