code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
s = 'hello'
s.upper()
set1 = set()
print('hi')
import timeit
timeit
singleton = 'hello',
singleton
basket = {'apple', 'orange', 'apple', 'pear', 'orange', 'banana'}
print(type(basket))
print(basket)
chars = {c for c in 'this is something'}
print(chars)
s = 'A man, a plan, a canal: Panama'
s='ab'
s=s.casefold()
chars = list(filter(lambda c:c.isalpha(), s))
low, high=0, len(chars)-1
while(low<high):
if chars[low] != chars[high]:
return False
return True
s='ab'
s = ''.join(filter(lambda c: c.isalnum(), s)).lower()
| code/algorithms/course_udemy_1/.ipynb_checkpoints/Practice-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sos
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SoS
# language: sos
# name: sos
# ---
# + [markdown] kernel="SoS" papermill={"duration": 0.036736, "end_time": "2019-12-23T17:04:37.466053", "exception": false, "start_time": "2019-12-23T17:04:37.429317", "status": "completed"} tags=[]
# # Customized or remote execution of workflows
# + [markdown] kernel="SoS" papermill={"duration": 0.028599, "end_time": "2019-12-23T17:04:37.522218", "exception": false, "start_time": "2019-12-23T17:04:37.493619", "status": "completed"} tags=[]
# * **Difficulty level**: intermediate
# * **Time need to lean**: 30 minutes or less
# * **Key points**:
# * Option `-r host` executes workflow on `host`, optionally through a `workflow_template` specified through host configuration.
# * The remote host could be a regular server, or a cluster system, in which case the workflow could be executed using multiple computing nodes.
# + [markdown] kernel="SoS" papermill={"duration": 0.021278, "end_time": "2019-12-23T17:04:37.564341", "exception": false, "start_time": "2019-12-23T17:04:37.543063", "status": "completed"} tags=[]
# Option `-r host` executes workflow on `host`. Depending on the properties of `host`, this option allows you to
#
# 1. Execute workflows locally, but in a customized environment
# 2. Execute workflows on a remote server directly
# 3. Execute entire workflows on a remote cluster system with option `-r`
# 3. Execute entire workflows on a remote cluster system with job submission with options `-r`, `-q`
#
# Please refer to [host configuration](host_setup.html) for details on host configuration.
# + [markdown] kernel="SoS" papermill={"duration": 0.024947, "end_time": "2019-12-23T17:04:37.612903", "exception": false, "start_time": "2019-12-23T17:04:37.587956", "status": "completed"} tags=[]
# ## Customized local environment for workflow execution
# + [markdown] kernel="SoS" papermill={"duration": 0.025339, "end_time": "2019-12-23T17:04:37.662600", "exception": false, "start_time": "2019-12-23T17:04:37.637261", "status": "completed"} tags=[]
# Assuming a system with two R versions, a system R installation under `/usr/local/bin` and a local installation in a conda environment. The latter version is the default version but if for some reasons the system R is preferred (e.g. if a library is only available there), you can change the local `PATH` of the `R` action using an `env` option (see [SoS actions](sos_actions.html) for details.
# + kernel="SoS" papermill={"duration": 2.454832, "end_time": "2019-12-23T17:04:40.144765", "exception": false, "start_time": "2019-12-23T17:04:37.689933", "status": "completed"} tags=[]
R:
R.Version()$version.string
# + kernel="SoS" papermill={"duration": 1.480646, "end_time": "2019-12-23T17:04:41.663985", "exception": false, "start_time": "2019-12-23T17:04:40.183339", "status": "completed"} tags=[]
import os
R: env={'PATH': f"/usr/local/bin:{os.environ['PATH']}"}
R.Version()$version.string
# + [markdown] kernel="SoS" papermill={"duration": 0.028171, "end_time": "2019-12-23T17:04:41.721396", "exception": false, "start_time": "2019-12-23T17:04:41.693225", "status": "completed"} tags=[]
# This action level `env` configuration is very flexible (e.g. you can use different versions of R in the same workflow) but can be difficult to maintain if you have multiple `R` actions. If your intent to use the same version of R throughout the workflow, it is easier to execute the entire workflow in a customized environment.
#
# To achieve this, you can define a host as follows, which has a default `address` of `localhost`.
# + kernel="SoS" papermill={"duration": 0.038384, "end_time": "2019-12-23T17:04:41.782508", "exception": false, "start_time": "2019-12-23T17:04:41.744124", "status": "completed"} tags=[]
# %save myconfig.yml -f
hosts:
system_R:
workflow_template: |
export PATH=/usr/local/bin:$PATH
{command}
# + [markdown] kernel="SoS" papermill={"duration": 0.02328, "end_time": "2019-12-23T17:04:41.831690", "exception": false, "start_time": "2019-12-23T17:04:41.808410", "status": "completed"} tags=[]
# Then, we will be using the conda version of R by default
# + kernel="SoS" papermill={"duration": 4.976894, "end_time": "2019-12-23T17:04:46.831672", "exception": false, "start_time": "2019-12-23T17:04:41.854778", "status": "completed"} tags=[]
# %run -v1
R:
R.Version()$version.string
# + [markdown] kernel="SoS" papermill={"duration": 0.024947, "end_time": "2019-12-23T17:04:46.879871", "exception": false, "start_time": "2019-12-23T17:04:46.854924", "status": "completed"} tags=[]
# and be using the system R if we execute the workflow in the `system_R` host with option `-r`, despite the fact that `system_R` is just a localhost with a template
# + kernel="SoS" papermill={"duration": 4.532367, "end_time": "2019-12-23T17:04:51.435095", "exception": false, "start_time": "2019-12-23T17:04:46.902728", "status": "completed"} tags=[]
# %run -r system_R -c myconfig.yml -v1
R:
R.Version()$version.string
# + [markdown] kernel="SoS" papermill={"duration": 0.021374, "end_time": "2019-12-23T17:04:51.479065", "exception": false, "start_time": "2019-12-23T17:04:51.457691", "status": "completed"} tags=[]
# As you can imagine, the template can set up a variety of different environment such as conda environments, debug environments, and using `module load` on a cluster system.
# + [markdown] kernel="SoS" papermill={"duration": 0.022024, "end_time": "2019-12-23T17:04:51.522285", "exception": false, "start_time": "2019-12-23T17:04:51.500261", "status": "completed"} tags=[]
# ## Execution of workflow on a remote host
# + [markdown] kernel="SoS" papermill={"duration": 0.020769, "end_time": "2019-12-23T17:04:51.611968", "exception": false, "start_time": "2019-12-23T17:04:51.591199", "status": "completed"} tags=[]
# If the `host` is a real remote host, then
#
# ```bash
# sos run script workflow -r host [other options]
# ```
# would execute the entire workflow on the `host`.
#
# This option is useful if you would like to **write the entire workflow for a remote host and execute the workflow with all input, software, and output files on the remote host**. Typical use cases for this option are when the data is too large to be processed locally, or when the software is only available on the remote host.
#
# For example, with a host definition similar to
#
# ```
# hosts:
# bcb:
# address: myserver.utexas.edu
# paths:
# home: /Users/bpeng1/scratch
# ```
#
# the following cell execute the workflow on `bcb`
# + kernel="SoS" papermill={"duration": 3.587526, "end_time": "2019-12-23T17:04:55.221544", "exception": false, "start_time": "2019-12-23T17:04:51.634018", "status": "completed"} tags=[]
# %run -r bcb
R:
set.seed(1)
x <- 1:100
y <- 0.029*x + rnorm(100)
png("test.png", height=400, width=600)
plot(x, y, pch=19, col=rgb(0.5, 0.5, 0.5, 0.5), cex=1.5)
abline(lm(y ~ x))
dev.off()
# + [markdown] kernel="SoS" papermill={"duration": 0.0319, "end_time": "2019-12-23T17:04:55.284564", "exception": false, "start_time": "2019-12-23T17:04:55.252664", "status": "completed"} tags=[]
# The resulting `test.png` are generated on `bcb` and is unavailable for local preview. You can however preview the file with `-r` option of magic `%preview`
# + kernel="SoS" papermill={"duration": 3.24173, "end_time": "2019-12-23T17:04:58.559507", "exception": false, "start_time": "2019-12-23T17:04:55.317777", "status": "completed"} tags=[]
# %preview -n test.png -r bcb
# + [markdown] kernel="SoS" papermill={"duration": 0.036206, "end_time": "2019-12-23T17:04:58.641017", "exception": false, "start_time": "2019-12-23T17:04:58.604811", "status": "completed"} tags=[]
# In this case we do not define any `workflow_template` for `bcb` so the workflow is executed directly on `bcb`. If a `workflow_template` is defined, the workflow will be executed through the shell script that is expanded from the template.
# -
# Note that local configrations, including the ones specified with option `-c` will be transferred and used on the remote host, with only the `localhost` definition switched to be the remote host. It is therefore safe to use local configurations with option `-r`.
# + [markdown] kernel="SoS" papermill={"duration": 0.057432, "end_time": "2019-12-23T17:05:22.505374", "exception": false, "start_time": "2019-12-23T17:05:22.447942", "status": "completed"} tags=[]
# ## Executing entire workflows on remote cluster systems
# -
# If the remote host specified by option `-r host` is a cluster system, the workflow will be submitted to the cluster as a regular cluster job. The `workflow_template` of `host` will be used, using options specified from command line (`-r host KEY=VALUE KEY=VALUE`.
#
# If `node=1`, sos will be executed with a single node, similar to what will happen if you execute the command locally. If `node=n` is specified, the workflow will be executed in a multi-node mode with `-j` automatically expanded to include remote workers that are assigned to the cluster job. see [option `-j`](verbosity_and_jobs.html) for details.
#
# Because no `-q` option is specified, all tasks will be executed as regular substeps (as long as no `queue=` option is specified) and be executed to the cluster nodes for execution. This method is most efficient for the execution of a large number of small tasks because there is no overhead for the creation and execution of external tasks. It is less efficient for workflows with varying "sizes" of tasks because all computing nodes may have to wait for one computing node to execute one substep. In which case the follow execution method, namely submiting additional tasks from computing nodes could be better.
# +
## Executing entire workflows on remote cluster systems with job submission
# + [markdown] kernel="SoS" papermill={"duration": 0.072575, "end_time": "2019-12-23T17:05:22.639139", "exception": false, "start_time": "2019-12-23T17:05:22.566564", "status": "completed"} tags=[]
# We already discussed the [use of option `-q` to submit tasks to remote hosts or cluster systems](task_statement.html). The key idea is that **the workflow is executed locally**. The main sos process will monitor and wait for the completion of remote tasks, and continue after the completion of the remote tasks. Because tasks are executed separately, it can be safely killed and resumed, which is one of the main advantages of the SoS task execution model.
# + [markdown] kernel="SoS" papermill={"duration": 0.066334, "end_time": "2019-12-23T17:05:22.903036", "exception": false, "start_time": "2019-12-23T17:05:22.836702", "status": "completed"} tags=[]
# It becomes a bit tricky to submit the workflow to a cluster system while allowing the master sos process, now executed on a computing node, to submit additional jobs. The command can be as simple as
#
# ```
# sos run script -r host -q host
# ```
#
# but things can go wrong if you do not understand completely what is happening:
#
# 1. With option `-r`, the entire workflow is executed on `host`. Since `host` has a PBS queue, the workflow will be submitted as a regular cluster job. **The resources needed for the master job needs to be specified from command line and expanded in `workflow_template`**.
# 2. The master `sos` process will be executed with a **transferred local config** on a computing node, with an option `-q host`. Now, because the `localhost` is set to `host` by the `-r` option, the cluster will appear to be a localhost to the `sos`, but with a different IP address etc.
# 3. The `sos` process will try to submit the tasks to the headnode through commands similar to `ssh host qsub job_id`. It is therfore mandatary that the `host` can be accessed with the specified `address` from the computing nodes. This is usually not a problem but it is possible that the headnode has different outfacing and inward addresses, in which case you will have to define a different `host` for option `-q`.
# + [markdown] kernel="SoS" papermill={"duration": 0.06946, "end_time": "2019-12-23T17:05:23.045618", "exception": false, "start_time": "2019-12-23T17:05:22.976158", "status": "completed"} tags=[]
# ```yaml
# hosts:
# htc:
# address: htc_cluster.mdanderson.edu
# description: HTC cluster (PBS)
# queue_type: pbs
# status_check_interval: 60
# submit_cmd: qsub {job_file}
# status_cmd: qstat {job_id}
# kill_cmd: qdel {job_id}
# nodes: 2
# cores: 4
# walltime: 01:00:00
# mem: 4G
# workflow_template: |
# #!/bin/bash
# #PBS -N {job_name}
# #PBS -l nodes={nodes}:ppn={cores}
# #PBS -l walltime={walltime}
# #PBS -l mem={mem}
# #PBS -m n
# module load R
# {command}
# task_template: |
# #!/bin/bash
# #PBS -N {job_name}
# #PBS -l nodes={nodes}:ppn={cores}
# #PBS -l walltime={walltime}
# #PBS -l mem={mem//10**9}GB
# #PBS -o ~/.sos/tasks/{task}.out
# #PBS -e ~/.sos/tasks/{task}.err
# #PBS -m n
# module load R
# {command}
# ```
| src/user_guide/remote_execution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Qbeast Datasource Format
#
#
#
# <p align="center">
# <img src="./images/Qbeast-spark.png" />
# </p>
#
#
#
# ## Content
# - Introduction
# - Table Indexing
# - Sample Pushdown
# - Table Tolerance
# - Analyze and Optimize
#
# ## Predicate PushDown
# **Predicate pushdown** is of great importance when it comes to **optimizing the logical plan** of a query. Among its benefits, one can find reduced usage of computation resources and less I/O from the secondary storage.
#
# Having a **predicate** in a query generally means the subsequent operators will work with fewer data. Without affecting the query output, further down the plan are the filters greater the benefits.
#
# ```sql
# SELECT A.name
# FROM A JOIN B ON A.id = B.id
# WHERE CONDITION_ON_A AND CONDITION_ON_B
#
# ```
#
# Take the query above as an example, a potential physical plan, without predicate pushdown can be as following:
#
#
#
# <p align="center">
# <img src="./images/g1.png" width="400" height="500" />
# </p>
#
#
#
# It starts by reading all the data from both tables **A** and **B**, performs the join on the id column, **A.id = B.id**, proceed to apply the **predicates** specified by the **WHERE** clause, **cond_A**, and **cond_B**, and finally project the target column, **name**.
#
# With the optimization of **predicate pushdown**, both conditions are found at the source and used as filters to select satisfying records, **reducing disk I/O** in this way. The **join** operator now also gets to operate with fewer data.
#
#
#
# <p align="center">
# <img src="./images/g2.png" width="400" height="500" />
# </p>
#
#
#
# As **query complexity** increases, the usage of **optimization rules** ensures moving the predicate is safe from altering the final output. Such rules are present in all major SQL query engines, with **Spark SQL** being no exception.
#
#
# ## Sampling
#
# **Sample** operators are yet another way to achieve the benefits of working with a reduced dataset. Unlike using filters where record selection is done **deterministically**, a (uniform) sample operator constructs a **representative subset** of the original data **randomly and uniformly**. The resulting subset is expected to have a **distribution** that resembles the source, and users generally only need to provide the **fraction** of the source data they desire to work with.
#
# ```python
# df = spark.read.load(source_path)
# df.sample(fraction=0.5)
# ```
#
# Its usage reduces **compute cost and latency** as before, except the result **accuracy** is inevitably compromised in relation to the subset size.
#
# Unlike filters, a sampler operator in Spark SQL can only do its job once all the data is retrieved from the source, for which a complete disk I/O is still required. Apart from that, there's no clear model to understand the **cost and latency vs. accuracy** trade-off when choosing the **fraction** to use, the reason for which there is a general avoidance for using samplers.
#
#
# ## Qbeast Format
#
# To address the above-mentioned issues, we introduce **qbeast datasource format** for Spark, a custom DataSource designed to enable **multidimensional indexing** for datasets together a set of transformation rules we achieve not only to convert the Sample operator into filters so random and uniform record selection can take place at the source, but on top of that, we've also created our own operator, **Table Tolerance**, which given the maximum query **tolerance** it can determine by itself the most cost-effective **fraction** to use, for which the user is no longer left wondering whether the sample they chose to use is accurate for their objectives.
# ## Setup
#
# Download Pyspark version 3.1.1
# + tags=[]
# !conda install pyspark=3.1.1 -y
# -
# Importing dependencies and initialize a Spark session
# +
import os
from pyspark.sql import SparkSession
DATA_ROOT = "/tmp/qbeast-test/data"
parquet_table_path = "s3a://qbeast-public-datasets/store_sales"
qbeast_table_path = os.path.join(DATA_ROOT, "qbeast/qtable")
hadoop_deps = ','.join(map(lambda a: 'org.apache.hadoop:hadoop-' + a + ':3.2.0', ['common','client','aws']))
deps = "io.qbeast:qbeast-spark_2.12:0.2.0,io.delta:delta-core_2.12:1.0.0,com.amazonaws:aws-java-sdk:1.12.20," + hadoop_deps
spark = (SparkSession.builder
.master("local[*]")
.config("spark.sql.extensions", "io.qbeast.spark.internal.QbeastSparkSessionExtension")
.config("fs.s3a.aws.credentials.provider", "org.apache.hadoop.fs.s3a.AnonymousAWSCredentialsProvider")
.config("spark.jars.packages", deps)
.getOrCreate())
spark.sparkContext.setLogLevel('OFF')
# -
# # Table Indexing
# The dataset used here is the **store_sales** table from **TCP-DS**. The format is parquet and its schema is shown below.
# +
parquet_df = spark.read.format("parquet").load(parquet_table_path)
print("Number of rows with na:", parquet_df.count())
# Display the schema
parquet_df.printSchema()
# -
# The table contains 23 columns in total, and the reason why only work with the first 5 is to have a cleaner query plan for later examination.
# +
processed_parquet_df = (
parquet_df
.select(
"ss_sold_time_sk",
"ss_item_sk",
"ss_customer_sk",
"ss_cdemo_sk",
"ss_hdemo_sk") # Selecting only the first 5 columns
.na.drop() # dropping rows with null values
)
print(f"Number of rows in the resulting dataframe: {processed_parquet_df.count()}")
processed_parquet_df.printSchema()
# -
# With the dataset set in place, we can write the table into a **qbeast datasource**, and indexing it using columns **ss_cdemo_sk** and **ss_hdemo_sk**. The choice of columns is trivial, at the moment any numerical column would do the trick. Generally one should choose the columns that they query most frequently on.
(processed_parquet_df
.write
.mode("overwrite")
.format("qbeast") # Saving the dataframe in a qbeast datasource
.option("columnsToIndex", "ss_cdemo_sk,ss_hdemo_sk") # Indexing the table
.option("cubeSize", "300000")
.save(qbeast_table_path)
)
# ## Sampling PushDown
#
# ## Qbeast sample vs Spark vanilla sample
#
# To demonstrate the transformation of the **Sample** operator into **Filters** and the subsequent application of Predicate PushDown, we will examine the query plan of a sample operation on a qbeast table and compare it with its application on a regular parquet table, namely **processed_parquet_df** from above.
# +
# write the processed parquet data to a new folder
# and re-read it so the query plan is simpler to examine.
processed_parquet_dir = os.path.join(DATA_ROOT, "parquet/test_data")
processed_parquet_df.write.mode("overwrite").format("parquet").save(processed_parquet_dir)
processed_parquet_df = spark.read.format("parquet").load(processed_parquet_dir)
# +
qbeast_df = spark.read.format("qbeast").load(qbeast_table_path)
assert qbeast_df.count() == processed_parquet_df.count(), "Both tables should have the same number of rows"
# 2.637.520
# +
print("Query Plan for Sampling on a parquet file\n")
processed_parquet_df.sample(fraction=0.1).explain(True)
# -
# Notice in the query plan for the parquet table that sample is the last operator from the query and it has remained that way for all stages of the query engine execution.
# +
print("\nQuery Plan for Sampling on a qbeast\n")
qbeast_df.sample(fraction=0.1).explain(True)
# -
# On the other hand, the sample operator is no longer present for the table with qbeast source, the optimized logical plan has a Filter that uses **qbeast_hash** to eliminate unnecessary data instead.
#
# Notice that the query plans for both dataframes are the same at the begining and they only started to differ after the application of the optimization rules, which in this case converted the **Sample** operator into **Filters** and applied **Predicate PushDown** rules from Spark query engine.
#
# These filters are pushed down to the level of the data source in the physical plan and are used by Spark as it scans the data from the source relation. The filters applied at the source are shown in the **DataFilters** filed from **FileScan parquet**.
# +
# processed_parquet_df.sample(0.1).collect()
# qbeast_df.sample(fraction=0.1).collect()
# -
# Execute the queries from the previous cell, and check the query plans from **Spark UI**.
#
# The fact that less files are accessed can be seen by comparing the total number of files in the folder and the number of files read from **Query Details**. Also, the **number of output rows** from **Scan parquet** can also indicate whether we are reading all the files.
#
# |Data Source |Total number of files |Number of files read |Number of rows read | Number of output rows
# |-------------------------|:-----------------------:|:-----------------------:|:-----------------------:|:-----------------------:|
# |parquet |16 |16 |2,637,520 | 264,192
# |qbeast |21 |1 |302,715 |262,013
#
# Under the hood, a qbeast table is divided into different partitions according to their states, and each partition is stored in a different parquet file. The filtering at the source is used for partition selection, and the second filtering is the one actually applied to the individual rows.
| docs/sample_pushdown_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Corpus creation
# This IPython notebook documents the creation of the corpora from raw data. The corpora can also be created interactively.
#
# ## Background
#
# Machine Learning task in the field of _Natural Language Processing (NLP)_ often rely on corpora. The ASR-stage in this project is no exception. Raw data is available from manifold sources. For this project, two sources (_ReadyLingua_ and _LibriSpeech_) were considered. However, other sources are conceivable. The final solution should be able to train on data from arbitrary resources. However, since properties and format of the raw data is usually not standardized between sources, some pre-processing is required in order to bring raw data into a format that can be used by the ASR stage.
#
# Since each data source makes its own assumptions about how data should be represented, a separate preprocessing step is required for each data source. The processed data is then stored in _corpora_, which contain the actual data (audio signals and transcripts) as well as metadata (audio segmentation information, audio length, sampling rate, language, speaker gender, etc...). The data from the sources used in this project comes from different distributions (e.g. number of languages, speaker per gender, etc.). Therefore the processed data has been stored in different corpora.
#
# ## Prerequisites
# This project was built using Python 3.6 and Anaconda 3. Please install the packages listed in `requirements.txt`. Additionally, you need the following tools and resources:
#
# * [FFMPEG](http://www.ffmpeg.org/): for the conversion and/or resampling of audio files
# * _ReadyLingua_ raw data: The aligned data from _ReadyLingua_ is not public domain. You need to get permission of the owner and store them on your machine.
#
# All other data is publicly available and will be downloaded as needed by this notebook.
# ### Source directory
# Since data from ReadyLingua and PodClub is not open to the public you must specify the path to the directory where those files are stored in the following cell. You must use an absolute path.
#
# Data from the LibriSpeech is available under the [Creative Commons](https://en.wikipedia.org/wiki/Creative_Commons) license. You can download the files yourself and specify an absolute path to folder where the files are stored. If the directory is empty, LibriSpeech data will automatically be downloaded and extracted there. If the directory is not empty, it is assumed that the data from LibriSpeech was already manually downloaded and extracted in this directory. In this case the directory structure must match the expected structure.
rl_source_root = r'D:\corpus\readylingua-raw' # path to directory where raw ReadyLingua data is stored
ls_source_root = r'D:\corpus\librispeech-raw' # path to directory where LibriSpeech files are or will be downloaded
# ### Target directory
# This notebook will create various corpora that need to be persisted somewhere. Specify the path to a directory that provides enough storage. Approximately 350GB of free storage is required. Note: Final storage use might be lower since some of the memory is only used temporarily.
#
# **Don't forget to execute the cell to apply the changes!**
target_root = r'E:/' # path to the directory where the corpora will be created (must have at least 350GB of free storage)
# ### Imports and helper functions
# Execute the cell below to import modules and helper functions
# +
"""
Imports and some helper functions. You don't need to change anything in here!
"""
import tarfile
import random
from os import listdir, rmdir, remove, makedirs
from random import randint
from shutil import move
import ipywidgets as widgets
import matplotlib.pyplot as plt
import os.path
import requests
from tqdm import tqdm
import create_ls_corpus
import create_rl_corpus
from util.audio_util import *
from util.corpus_util import *
from IPython.display import HTML, Audio
import ipywidgets as widgets
% matplotlib inline
# path to target directory for ReadyLingua corpus files (default value)
rl_target_root = os.path.join(target_root, 'readylingua-corpus')
# path to target directory for LibriSpeech corpus files (default value)
ls_target_root = os.path.join(target_root, 'librispeech-corpus')
def show_corpus_entry(corpus_entry, speech=None, speech_unaligned=None, pause=None):
speech = speech if speech else random.choice(corpus_entry.speech_segments)
speech_unaligned = speech_unaligned if speech_unaligned \
else random.choice(corpus_entry.speech_segments_unaligned) if corpus_entry.speech_segments_unaligned \
else None
pause = pause if pause else random.choice(corpus_entry.pause_segments)
show_audio(corpus_entry)
show_segment(speech)
if speech_unaligned:
show_segment(speech_unaligned)
show_segment(pause)
def show_audio(corpus_entry):
title = HTML(f"""
<h3>Sample corpus entry: {corpus_entry.name}</h3>
<p><strong>Path to raw data</strong>: {corpus_entry.raw_path}</p>
<p>{len(corpus_entry.speech_segments)} speech segments, {len(corpus_entry.pause_segments)} pause segments</p>
""")
audio = Audio(data=corpus_entry.audio, rate=corpus_entry.rate)
transcript = widgets.Accordion(children=[widgets.HTML(f'<pre>{corpus_entry.transcript}</pre>')], selected_index=None)
transcript.set_title(0, 'Transcript')
display(title)
display(audio)
display(transcript)
def show_segment(segment):
title = HTML(f'<strong>Sample {segment.segment_type}</strong> (start_frame={segment.start_frame}, end_frame={segment.end_frame})')
audio = Audio(data=segment.audio, rate=segment.rate)
display(title)
display(audio)
if segment.text:
transcript = HTML(f'<pre>{segment.transcript}</pre>')
display(transcript)
def download_file(url, target_dir):
r = requests.get(url, stream=True)
total_size = int(r.headers.get('content-length', 0));
block_size = 1024
wrote = 0
tmp_file = os.path.join(target_dir, 'download.tmp')
if not exists(target_dir):
makedirs(target_dir)
with open(tmp_file, 'wb') as f:
with tqdm(r.iter_content(32 * block_size), total=total_size, unit='B', unit_divisor=block_size,
unit_scale=True) as pbar:
for data in r.iter_content(32 * 1024):
wrote = wrote + len(data)
f.write(data)
pbar.update(len(data))
if total_size != 0 and wrote != total_size:
print("ERROR, something went wrong")
print('Extracting data...')
tar = tarfile.open(tmp_file, "r:gz")
tar.extractall(target_dir)
tar.close()
remove(tmp_file)
def move_files(src_dir, target_dir):
for filename in listdir(src_dir):
move(os.path.join(src_dir, filename), os.path.join(target_dir, filename))
rmdir(src_dir)
def on_download_ls_button_click(sender):
global ls_source_root
print('Downloading LibriSpeech data... Get lunch or something!')
print('Download 1/2: Audio data')
download_dir = os.path.join(ls_source_root, 'audio')
if exists(download_dir) and listdir(download_dir):
print(f'Directory {download_dir} exists and is not empty. Assuming data was already downloaded there.')
else:
download_file('http://www.openslr.org/resources/12/original-mp3.tar.gz', download_dir)
print('Done! Moving files...')
move_files(os.path.join(download_dir, 'LibriSpeech'), download_dir)
print('Download 2/2: Text data')
download_dir = os.path.join(ls_source_root, 'books')
if exists(download_dir) and listdir(download_dir):
print(f'Directory {download_dir} exists and is not empty. Assuming data was already downloaded there.')
else:
download_file('http://www.openslr.org/resources/12/original-books.tar.gz', download_dir)
move_files(os.path.join(download_dir, 'LibriSpeech'), download_dir)
makedirs(os.path.join(download_dir, 'utf-8'))
move_files(os.path.join(download_dir, 'books', 'utf-8'), os.path.join(download_dir, 'utf-8'))
move_files(os.path.join(download_dir, 'books', 'ascii'), os.path.join(download_dir, 'ascii'))
delete_directory = os.path.join(download_dir, 'books')
print(f'Done! Please delete {delete_directory} manually (not needed)')
print(f'Files downloaded and extracted to: {ls_source_root}')
def on_create_rl_button_click(sender):
global rl_corpus_file
print('Creating ReadyLingua corpus... Get a coffee or something!')
rl_corpus, rl_corpus_file = create_rl_corpus.create_corpus(source_root=rl_source_root, target_root=rl_target_root)
print(f'Done! Corpus with {len(rl_corpus)} entries saved to {rl_corpus_file}')
def on_create_ls_button_click(sender):
global ls_corpus_file
print('Creating LibriSpeech corpus... Go to bed or something!')
ls_corpus, ls_corpus_file = create_ls_corpus.create_corpus(source_root=ls_source_root, target_root=ls_target_root)
print(f'Done! Corpus with {len(ls_corpus)} entries saved to {rl_corpus_file}')
# UI elements
layout = widgets.Layout(width='250px', height='50px')
download_ls_button = widgets.Button(description="Download LibriSpeech Data", button_style='info', layout=layout, icon='download')
download_ls_button.on_click(on_download_ls_button_click)
create_rl_button = widgets.Button(description="Create ReadyLingua Corpus", button_style='warning', layout=layout, icon="book", tooltip='~5 minutes')
create_rl_button.on_click(on_create_rl_button_click)
create_ls_button = widgets.Button(description="Create LibriSpeech Corpus", button_style='warning', layout=layout,icon="book", tooltip='~5 hours')
create_ls_button.on_click(on_create_ls_button_click)
# -
# ## Corpus structure
# The alignment information is extracted from the raw data and stored as a **corpus** containing **corpus entries**. A corpus entry reflects a single instance for training, validation or evaluation. It contains all the information about the audio and its segmentation. It therefore contains information about **segments**, which are either speech segments, pause segments or unaligned speech segments. Unaligned speech segments are parts of the audio signal which are known to contain speech but for which no metadata from manual segmentation is available. They may therefore contain speech or pause segments themselves and could be further subdivided.
#
# The following figure shows an illustration of the most important classes used for corpus creation:
#
# 
#
# ### Preprocessing
# The raw data was integrated as-is applying only the following preprocessing steps:
#
# * **Resampling**: Audio data was resampled to 16kHz (mono) WAV files
# * **Cropping**: Some of the audio files (especially in the LibriSpeech data contained some preliminary information about LibriVox and the book being read before the actual recording. This speech data was not aligned. The audio was therefore cropped at the beginning to the frame where the first alignment information (speech or pause segments) begins. Likewise, the audio is cropped at the end to the frame where the last alignment information ends.
# ### Corpus entries
# In order to allow data from all sources for training, it had to be converted to a common format. Since (to my knowledge) there is not a standardized format for FA, I had to define one myself. Therefore I went for the following structure for a single corpus entry:
#
# ```JSON
# // corpus is iterable over its corpus_entries
# Corpus = {
# 'name': string, // display name
# 'root_path': string, // absolute path to the directory containing the corpus files
# 'corpus_entries': [CorpusEntry] // the entries of the corpus
# }
#
# // corpus_entry is iterableover its segments
# CorpusEntry =
# {
# 'corpus': Corpus, // reference to the corpus
# 'audio_file': string // absolute path to the preprocessed audio file
# 'transcript': string, // transcription of the audio as raw (unaligned) text
# 'segments': [Segment], // speech- and pause-segments of the audio
# 'original_path': string // absolute path to the directory containing the raw files
# 'name': string // display name
# 'id': string // unique identifier
# 'language': string, // 'de'/'fr'/'it'/'en'/'es'/'unknown'
# 'chapter_id': string, // identifier of the chapter of the book if available, else 'unknown'
# 'speaker_id': string, // identifier of the speaker if available, else 'unknown'
# 'original_sampling_rate': string, // sampling rate of the raw audio file
# 'original_channels': string, // number of channels in the raw audio file
# 'subset': string, // membership to a subset ('train'/'dev'/'test'/'unknown')
# 'media_info': dict, // PyDub information about the converted audio file
# 'speech_segments': [Segment], // segments filtered for type=='speech' (at runtime)
# 'pause_segments': [Segment], // segments filtered for type=='pause' (at runtime)
# 'alignment': ([byte], [Segment]), // audio and segmentation information
# 'alignment_cropped': ([byte], [Segment]) // audio and segmentation information with start and end cropped
# }
#
# // definition of a speech or pause segment
# Segment =
# {
# 'corpus_entry': CorpusEntry, // reference to the corresponding CorpusEntry
# 'start_frame': int, // index of the start frame of the segment within the audio
# 'end_frame': int, // index of the end frame of the segment within the audio
# 'start_text': int, // index of first character of the segment in the transcription
# 'end_text': int, // index of the last character of the segment in the transcription
# 'segment_type': string, // 'speech' for a speech segment, 'pause' for a pause segment
# 'audio': [byte], // part of the audio of the corpus entry which belongs to this segment
# 'text': string // part of the transcription of the corpus entry which belongs to this segment
# }
# ```
# ### Create ReadyLingua Corpus
# ReadyLingua (RL) provides alignment data distributed over several files files:
#
# * `*.wav` or `*.mp3`: Audio file containing the speech
# * `*.txt`: UTF-8 encoded (unaligned) transcription
# * `* - Segmentation.xml`: file containing the definition of speech- and pause segments
# ```XML
# <Segmentation>
# <SelectionExtension>0</SelectionExtension>
# <Segments>
# <Segment id="1" start="83790" end="122598" class="Speech" uid="5" />
# ...
# </Segments>
# <Segmenter SegmenterType="SICore.AudioSegmentation.EnergyThresholding">
# <MaxSpeechSegmentExtension>50</MaxSpeechSegmentExtension>
# <Length>-1</Length>
# <Energies>
# <Value id="1" value="0" />
# ...
# </Energies>
# <OriginalSegments>
# <Segment id="1" start="83790" end="100548" class="Speech" uid="2" />
# ...
# </OriginalSegments>
# <EnergyPeak>3569753</EnergyPeak>
# <StepSize>441</StepSize>
# <ITL>146139</ITL>
# <ITU>730695</ITU>
# <LastUid>2048</LastUid>
# <MinPauseDuration>200</MinPauseDuration>
# <MinSpeechDuration>150</MinSpeechDuration>
# <BeginOfSilence>1546255</BeginOfSilence>
# <SilenceLength>100</SilenceLength>
# <ThresholdCorrectionFactor>1</ThresholdCorrectionFactor>
# </Segmenter>
# </Segmentation>
# ```
# * `* - Index.xml`: file containing the actual alignments of text to audio
# ```XML
# <XMLIndexFile>
# <Version>2.0.0</Version>
# <SamplingRate>44100</SamplingRate>
# <NumberOfIndices>91</NumberOfIndices>
# <TextAudioIndex>
# <TextStartPos>0</TextStartPos>
# <TextEndPos>36</TextEndPos>
# <AudioStartPos>952101</AudioStartPos>
# <AudioEndPos>1062000</AudioEndPos>
# <SpeakerKey>-1</SpeakerKey>
# </TextAudioIndex>
# ...
# </XMLIndexFile>
# ```
# * `* - Project.xml`: Project file binding the different files together for a corpus entry (note: this file is optional, i.e. there may be not project file for a corpus entry)
#
# Corpus entries are organized in a folder hierarchy. There is a fileset for each corpus entry. Usually, the files for a specific corpus entry reside in a leaf directory (i.e. a directory without further subdirectories). If there is a project file, this file is used to locate the files needed.
#
# Audio data is provided as Wave-Files with a sampling rate of 44,1 kHz (stereo) or MP3 files. Because most ASR corpora provide their recordings as wave files with a sampling rate of 16 kHz the files were downsampled and the alignment information adjusted. The raw transcription is integrated as-is. The XML files are parsed to extract the alignment data. Alignment-, textual and downsampled audio data are merged into a corpus entry as described above.
# #### Create corpus entries
# We need to extract the alignments from the segmentation information of the raw data. For this, the downloaded data needs to be converted to corpus entries. This process takes a few minutes, so this is a good time to have a coffee break.
display(create_rl_button)
# #### Explore corpus
# Let's load the newly created corpus (needs to be done only once) and print some stats:
rl_corpus = load_corpus(rl_target_root)
rl_corpus.summary()
# You can access each corpus entry either by a numerical index or by its ID (string).
# +
# acces by index
first_entry = rl_corpus[0]
first_entry.summary()
# access by ID
other_entry = rl_corpus['news170524']
other_entry.summary()
# get a list of IDs
rl_corpus.keys
# -
# You can also filter the corpus by language to get only the corpus entry with the specified language(s):
# +
rl_corpus_de = rl_corpus(languages='de')
rl_corpus_de.summary()
rl_corpus_fr = rl_corpus(languages='fr')
rl_corpus_fr.summary()
rl_corpus_de_fr = rl_corpus(languages=['de', 'fr'])
rl_corpus_de_fr.summary()
# -
# To see if everything worked as expected let's check out a sample alignment. You can execute the cell below to show a random alignment from a random corpus entry. You can execute the cell several times to see different samples.
corpus_entry = random.choice(rl_corpus_de)
# corpus_entry = rl_corpus['edznachrichten180201']
show_corpus_entry(corpus_entry)
# ### Create LibriSpeech Corpus
# The _LibriSpeech_ raw data is split into training-, dev- and test-set (`train-*.tar.gz`, `dev-*.tar.gz` and `test-*.tar.gz`). However, those sets only contain the transcript as a set of segments and an audio file for each segment. They do not contain any temporal information which is needed for alignment.
#
# Luckily, there is also the `original-mp3-tar.gz` for download which contains the original LibriVox mp3 files (from which the corpus was created) along with the alignment information. Alignment is made on utterance-level, i.e. the transcript is split up into segments whereas each segment corresponds to an utterance. Segments were derived by allowing splitting on every silence interval longer than 300ms.
#
# The data is organized into subdirectories of the following path format:
#
# ./LibriSpeech/mp3/{speaker_id}/{chapter_id}/
#
# There is one directory per entry containing all the information about a recording. For this project the following files are important:
#
# - **Audio recording** `{chapter_id}.mp3`: The audio file containing the recording. The audio is mono with a bitrate of 128 kB/s and a sampling rate of 44.1 kHz and needs to be converted/resampled to the target format.
# - **Transcription file** `{speaker_id}-{chapter_id}.trans.txt`: Text file containing the transcriptions of the segments (one segment per line). Each line is prefixed with the transcription ID. The transcription is all uppercase and does not contain any punctuation.
# ```
# 14-208_0000 CHAPTER ELEVEN THE MORROW BROUGHT A VERY SOBER LOOKING MORNING THE SUN MAKING ONLY A FEW EFFORTS...
# ```
# - **Segmentation file** `{speaker_id}-{chapter_id}.seg.txt`: Text file containing temporal information about the segments (one segment per line). Each line is prefixed with the ID of the transcription for which the information is valid. The time is indicated in seconds. Example:
# ```
# 14-208_0000 25.16 40.51
# ```
#
# In order to create the corpus, these files had to be parsed and the audio was converted and downsampled to a 16kHz Wave-file.
# Information about the Speakers, Chapters and Books were extracted from the respective files (`SPEAKERS.TXT`, `CHAPTERS.TXT` and `BOOKS.TXT`).
#
# #### Unaligned speech segments
# _Speech segments_ could be derived by exploiting the temporal information from aligned parts of the corpus. Short intervals between speeches were interpreted as _pause segments_. However, since not all passages in the recordings were aligned with text from the underlying book, not all intervals between speech segments correspond to speech pauses. By comparing the transcripts of aligned speech sequences with the underlying book text any time interval could be classified as either _unaligned speech_ or a pause segment:
#
# * if the concatenated transcripts of two subsequent aligned speech segments did not match with any part of the book text, the transcript contains "holes" (i.e. parts of the book were left out during alignment). The interval between the aligned speech segments were then treated as _unaligned speech_ and may contain any number of pause segments at the start, beginning or within the interval.
# * If the concatenated transcripts of two subsequent aligned speech segments roughly matched up with some passage from the book text, the speech segments were deemed consistent (i.e. the recordings of the speech segments were made from subsequent passages of text). The interval between the speech segments were then treated as a _pause segment_.
#
# Since the encoding of the transcripts from the _LibriSpeech_ corpus deviated from the original book text, the comparison between transcripts and book text was made using normalized versions of both. Normalization was acquired through removing non-ASCII characters and punctuation, replacing multiple whitespaces with a single and converting everything to uppercase.
# #### Download raw data
# To create the LibriSpeech corpus you first need to download the raw data. The files are over 80GB and need to be extracted, so this might take a while...
display(download_ls_button)
# #### Create corpus
# We need to extract the alignments from the segmentation information of the raw data. For this, the downloaded data needs to be converted to corpus entries. **This process takes several hours, so you might want to do this just before knocking-off time!**
display(create_ls_button)
# #### Explore corpus
# Again, let's load the newly created corpus:
ls_corpus = load_corpus(ls_target_root)
ls_corpus.summary()
# To see if everything worked as expected let's check out a sample alignment. You can execute the cell below to show a random alignment from a random corpus entry. You can execute the cell several times to see different samples.
corpus_entry = random.choice(ls_corpus)
# corpus_entry = ls_corpus[0]
corpus_entry.summary()
show_corpus_entry(corpus_entry)
# ## Summary
#
# This notebook showed how raw data from _ReadyLingua_ and _LibriSpeech_ was pre-processed and stored in corpora using a common format. The summary of the corpora also showed significant difference between the corpora:
#
# The _ReadyLingua_ corpus contains only 27 hours of transcribed audio in different languages. The amount of recordings is heavily unbalanced, both in number and total lengh. Also, some recordings exhibit audible effects like reverb. Recordings in this corpus partly overlap with the _LibriSpeech_ corpus.
#
# In contrast, the _LibriSpeech_ corpus contains roughly 1400 hours of transcribed audio in English. The raw data for corpus was specifically created for ASR. Because of the sheer amount of data, this corpus is suited much more to train a Neural Network. Also, the recording quality is more homogenous than in the _ReadyLingua_ corpus and free from distortions like echo and reverb. Another advantage of this corpus is that the data has already been split into training-, validation- and test-set. This split was made carefully so that they are disjoint, i.e. the audio of each speaker is assigned to exactly one set. Also, the distribution of male/female speakers is similar between these sets.
| src/01_corpus_creation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="JcjOrJxIo5vp"
# # colab-a11y-util
#
# This library provides the following functions in Google Colab
# - Notification by sound at the time of cell execution
# - One beep sound when executed
# - Two beep sounds when normal completion
# - Two buzzer sounds when abnormal completion
# - Sound notification of progress bar by tqdm
# - Simple audio output function that can be used instead of print
#
# + [markdown] id="FmWA1KWLlOqZ"
# # Install
# + id="3dy25kcpfTg5"
# stable
# !pip install colab-a11y-utils
# latest
# #!pip install -U git+https://github.com/hassaku/colab-a11y-utils.git
# + [markdown] id="1rdqQCaflcRZ"
# # Enable sound notification when cells are executed
# + id="QEy93iWDfaAa"
from colab_a11y_utils import set_sound_notifications
set_sound_notifications()
# + id="J7FHt20JffsX"
import time
print("start")
time.sleep(3)
print("finish")
# + id="CeXV3crrfnpu"
raise
# + [markdown] id="021Jdm2npMLf"
# # Enable sound report in progress bar
# + id="pbflm94fgH_H"
from colab_a11y_utils import tqdm
for _ in tqdm(range(10)):
time.sleep(1)
# + [markdown] id="Mlb1WLUHl-hP"
# # Disable sound notification when cells are executed.
# + id="CKnobZsMf0ho"
from colab_a11y_utils import unset_sound_notifications
unset_sound_notifications()
# + id="OPtonKJdgkdL"
raise
# + [markdown] id="j-FNh9qSzE1N"
# # Speak
# + id="vnJFPZrRon10"
from colab_a11y_utils import speak
speak("Hello")
| colab_a11y_util_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Out-of-bag error and feature selection
# Семинар основан на <a href="https://habr.com/en/company/ods/blog/324402/">материале</a> ODS
import warnings
warnings.simplefilter("ignore")
# +
import numpy as np
import pandas as pd
import seaborn as sns
from scipy.special import binom
from IPython.display import Image
from matplotlib import pyplot as plt
from sklearn.ensemble import BaggingRegressor, BaggingClassifier
from sklearn.ensemble import RandomForestRegressor, RandomForestClassifier
from sklearn.tree import DecisionTreeRegressor, DecisionTreeClassifier
from sklearn.metrics import accuracy_score
from sklearn.datasets import load_digits as load
from sklearn.model_selection import cross_val_score, StratifiedKFold, GridSearchCV, train_test_split
# %matplotlib inline
# -
# ## Out-of-bag error
# ----------
# <img src='img/oob.png' width=700>
# **Задача** Покажите, что примерно 37% примеров остаются вне выборки бутстрэпа и не используются при построении k-го дерева.
# **Решение** Пусть в выборке $l$ объектов. На каждом шаге все объекты попадают в подвыборку с возвращением равновероятно, т.е отдельный объект — с вероятностью $\dfrac{1}{l}$. Вероятность того, что объект НЕ попадет в подвыборку (т.е. его не взяли $l$ раз): $(1-\dfrac{1}{l})^l$
#
#
# $$\lim_{l \rightarrow +\infty} (1-\dfrac{1}{l})^l = \dfrac{1}{e}$$
#
# Тогда вероятность попадания конкретного объекта в подвыборку $1 - \dfrac{1}{e} \approx 63\%$.
# Out-of-Bag оценка — это усредненная оценка базовых алгоритмов на тех ~37% данных, на которых они не обучались.
# ## Отбор признаков с помощью случайного леса
# <img src='img/features.png'>
# Данные можно взять здесь: https://github.com/Yorko/mlcourse_open/blob/master/data/hostel_factors.csv
# +
# #!wget https://raw.githubusercontent.com/Yorko/mlcourse.ai/master/data/hostel_factors.csv -O data/hostel_factors.csv
# -
hostel_data = pd.read_csv("data/hostel_factors.csv")
features = {
"f1":u"Персонал",
"f2":u"Бронирование хостела ",
"f3":u"Заезд в хостел и выезд из хостела",
"f4":u"Состояние комнаты",
"f5":u"Состояние общей кухни",
"f6":u"Состояние общего пространства",
"f7":u"Дополнительные услуги",
"f8":u"Общие условия и удобства",
"f9":u"Цена/качество",
"f10":u"ССЦ",
}
# +
forest = RandomForestRegressor(n_estimators=1000, max_features=10,
random_state=0)
forest.fit(hostel_data.drop(['hostel', 'rating'], axis=1),
hostel_data['rating'])
importances = forest.feature_importances_
indices = np.argsort(importances)[::-1]
# +
# Plot the feature importancies of the forest
num_to_plot = 10
feature_indices = [ind+1 for ind in indices[:num_to_plot]]
# Print the feature ranking
print("Feature ranking:")
for f in range(num_to_plot):
print("%d. %s %f " % (f + 1, features["f"+str(feature_indices[f])], importances[indices[f]]))
plt.figure(figsize=(15, 5))
bars = plt.bar(range(num_to_plot), importances[indices[:num_to_plot]], color=([str(i/float(num_to_plot+1)) for i in range(num_to_plot)]), align="center")
ticks = plt.xticks(range(num_to_plot), feature_indices)
plt.xlim([-1, num_to_plot])
plt.title(u"Важность конструктов")
plt.legend(bars, [u''.join(features["f"+str(i)]) for i in feature_indices])
# -
# <h1 align="center">Выводы</h1>
# **Bagging**:
# - Одна из лучших техник для построения алгоритмов ML
# - Линейно уменьшает разброс и не уменьшает смещение (если не коррелированы ответы базовых алоритмов)
# - Слабое переобучение
# - НО переобучение ЕСТЬ -- от сложности одного алгоритма, лучше все же немного обрезать деревья
#
# ** Random Forest **
#
# Плюсы:
# - имеет высокую точность предсказания, на большинстве задач будет лучше линейных алгоритмов; точность сравнима с точностью бустинга
# - практически не чувствителен к выбросам в данных из-за случайного сэмлирования
# - не чувствителен к масштабированию (и вообще к любым монотонным преобразованиям) значений признаков, связано с выбором случайных подпространств
# - не требует тщательной настройки параметров, хорошо работает «из коробки». С помощью «тюнинга» параметров можно достичь прироста от 0.5 до 3% точности в зависимости от задачи и данных
# - способен эффективно обрабатывать данные с большим числом признаков и классов
# - одинаково хорошо обрабатывет как непрерывные, так и дискретные признаки
# - редко переобучается, на практике добавление деревьев почти всегда только улучшает композицию, но на валидации, после достижения определенного количества деревьев, кривая обучения выходит на асимптоту
# - для случайного леса существуют методы оценивания значимости отдельных признаков в модели
# - хорошо работает с пропущенными данными; сохраняет хорошую точность, если большая часть данных пропущенна
# - предполагает возможность сбалансировать вес каждого класса на всей выборке, либо на подвыборке каждого дерева
# - вычисляет близость между парами объектов, которые могут использоваться при кластеризации, обнаружении выбросов или (путем масштабирования) дают интересные представления данных
# - возможности, описанные выше, могут быть расширены до неразмеченных данных, что приводит к возможности делать кластеризацию и визуализацию данных, обнаруживать выбросы
# - высокая параллелизуемость и масштабируемость.
#
# Минусы:
# - в отличие от одного дерева, результаты случайного леса сложнее интерпретировать
# - нет формальных выводов (p-values), доступных для оценки важности переменных
# - алгоритм работает хуже многих линейных методов, когда в выборке очень много разреженных признаков (тексты, Bag of words)
# - случайный лес не умеет экстраполировать, в отличие от той же линейной регрессии (но это можно считать и плюсом, так как не будет экстремальных значений в случае попадания выброса)
# - алгоритм склонен к переобучению на некоторых задачах, особенно на зашумленных данных
# - для данных, включающих категориальные переменные с различным количеством уровней, случайные леса предвзяты в пользу признаков с большим количеством уровней: когда у признака много уровней, дерево будет сильнее подстраиваться именно под эти признаки, так как на них можно получить более высокое значение оптимизируемого функционала (типа прироста информации)
# - если данные содержат группы коррелированных признаков, имеющих схожую значимость для меток, то предпочтение отдается небольшим группам перед большими
# - больший размер получающихся моделей. Требуется $O(NK)$ памяти для хранения модели, где $K$ — число деревьев.
| week06_rf/week06_extra_oob_and_feature_selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import das Bibliotecas:
import numpy
import sys
import matplotlib.pyplot as plt
from datetime import datetime
import pandas as pd
a = sys.path.append('../modules/') # endereco das funcoes implementadas por voce!
import prism, noise, plot_3D, auxiliars, salve_doc
# ### Etapa 1: Definicão das coordenadas de Observação:
# +
nx = 100 # n de observacoes na direcao x
ny = 100 # n de observacoes na direcao y
size = (nx, ny)
xmin = -10000.0 # metros
xmax = +10000.0 # metros
ymin = -10000.0 # metros
ymax = +10000.0 # metros
z = -100.0 # altura de voo, em metros
# +
dicionario = {'nx': nx,
'ny': ny,
'xmin': xmin,
'xmax': xmax,
'ymin': ymin,
'ymax': ymax,
'z': z,
'color': '.r'}
x, y, X, Y = plot_3D.create_aquisicao(dicionario)
# -
# ### Etapa 2: Definicão das coordenadas dos prismas modelados:
# coordenadas dos vertices (corners) do prisma, em metros:
x1,x2 = (-2000.0, 2000.0)
y1,y2 = (-2500.0, 2500.0)
z1,z2 = (500.0,1000.0) # z eh positivo para baixo!
deltaz = 100.0
deltay = 4000.0
incl = 'positivo'
dic = {'n': 3,
'x': [x1, x2],
'y': [y1, y2],
'z': [z1, z2],
'deltay': deltay,
'deltaz': deltaz,
'incl': 'positivo'}
pointx, pointy, pointz = plot_3D.creat_point(dic)
print(pointx)
print(pointy)
print(pointz)
# +
# #%matplotlib notebook
dic1 = {'x': [pointx[0], pointx[1]],
'y': [pointy[0], pointy[1]],
'z': [pointz[0], pointz[1]]}
dic2 = {'x': [pointx[2], pointx[3]],
'y': [pointy[2], pointy[3]],
'z': [pointz[2], pointz[3]]}
dic3 = {'x': [pointx[4], pointx[5]],
'y': [pointy[4], pointy[5]],
'z': [pointz[4], pointz[5]]}
#----------------------------------------------------------------------------------------------------#
vert1 = plot_3D.vert_point(dic1)
vert2 = plot_3D.vert_point(dic2)
vert3 = plot_3D.vert_point(dic3)
#----------------------------------------------------------------------------------------------------#
color = 'b'
size = [9, 10]
view = [210, 30]
#----------------------------------------------------------------------------------------------------#
prism_1 = plot_3D.plot_prism(vert1, color)
prism_2 = plot_3D.plot_prism(vert2, color)
prism_3 = plot_3D.plot_prism(vert3, color)
#----------------------------------------------------------------------------------------------------#
prisma = {'n': 3,
'prisma': [prism_1, prism_2,prism_3]}
plot_3D.plot_obs_3d(prisma, size, view, x, y, pointz)
# -
# ### Etapa 3: Simulação do campo Principal na região das observações:
# +
I = -30.0 # inclinacao do campo principal em graus
D = -23.0 # declinacao do campo principal em graus
Fi = 40000.0 # Intensidade do campo principal (nT)
# Campo principal variando com as posicao F(X,Y):
F = Fi + 0.013*X + 0.08*Y # nT
# -
# ### Etapa 4: Definição das propriedades das fontes crustais (prismas verticas):
# +
# Propriedades magneticas da fonte crustal:
inc = I # magnetizacao puramente induzida
dec = -10.0
Mi = 10.0 # intensidade da magnetizacao em A/m
Mi2 = 15.0
Mi3 = 7.0
fonte_crustal_mag1 = [pointx[0], pointx[1],
pointy[0], pointy[1],
pointz[0], pointz[1], Mi]
fonte_crustal_mag2 = [pointx[2], pointx[3],
pointy[2], pointy[3],
pointz[2], pointz[3], Mi2]
fonte_crustal_mag3 = [pointx[4], pointx[5],
pointy[4], pointy[5],
pointz[4], pointz[5], Mi3]
# -
# ### Etapa 5: Cálculo das anomalias via function (prism_tf)
# +
tfa1 = prism.prism_tf(Y, X,z, fonte_crustal_mag1, I, D, inc, dec)
tfa2 = prism.prism_tf(Y, X,z, fonte_crustal_mag2, I, D, inc, dec)
tfa3 = prism.prism_tf(Y, X,z, fonte_crustal_mag3, I, D, inc, dec)
tfa_final = tfa1 + tfa2 + tfa3
# -
# ### Etapa 6: Acréscimo de rúido via function (noise_normal_dist)
# +
mi = 0.0
sigma = 0.1
#ACTn = noise.noise_gaussiana(t, mi, sigma, ACT)
tfa_final1 = auxiliars.noise_normal_dist(tfa_final, mi, sigma)
# +
# %matplotlib inline
#xs = [x1, x1, x2, x2, x1]
#ys = [y1, y2, y2, y1, y1]
xs1 = [pointx[0], pointx[0], pointx[5], pointx[5], pointx[0]]
ys1 = [pointy[0], pointy[5], pointy[5], pointy[0], pointy[0]]
#flechax = [[numpy.absolute(pointx[0] + pointx[5])], [pointx[5]]]
#flechay = [[numpy.absolute(pointy[0] + pointy[5])], [pointy[5]]]
origin = [[numpy.absolute(pointx[0] + pointx[5])], [[numpy.absolute(pointy[0] + pointy[5])]]]
ponta = [[pointx[5]], [pointy[5]]]
print(ponta)
# graficos
plt.close('all')
plt.figure(figsize=(10,10))
#******************************************************
plt.contourf(Y, X, tfa_final, 20, cmap = plt.cm.RdBu_r)
plt.title('Anomalia de Campo Total(nT)', fontsize = 12)
plt.xlabel('East (m)', fontsize = 10)
plt.ylabel('North (m)', fontsize = 10)
corpo, = plt.plot(ys1,xs1,'k-*', label = 'Extensão do Dique')
#plt.plot(ys2,xs2,'k-')
#plt.plot(ys3,xs3,'m-')
arrow = plt.arrow(2000.0, 0.0, 4500.0, 0.0, width=250, length_includes_head = True, color = 'k')
first_legend = plt.legend(handles=[corpo], bbox_to_anchor=(1.25, 1), loc='upper left', borderaxespad=0.0, fontsize= 12.0)
plt.legend([arrow, corpo], ['Direção de mergulho', 'Extensão do Dique'], bbox_to_anchor=(1.25, 1), loc='upper left', borderaxespad=0.0, fontsize= 12.0)
plt.colorbar()
#plt.savefig('teste_100_40000_D10.png', format='png')
plt.show()
# +
xs1 = [pointx[0], pointx[0], pointx[1], pointx[1], pointx[0]]
xs2 = [pointx[2], pointx[2], pointx[3], pointx[3], pointx[2]]
xs3 = [pointx[4], pointx[4], pointx[5], pointx[5], pointx[4]]
ys1 = [pointy[0], pointy[1], pointy[1], pointy[0], pointy[0]]
ys2 = [pointy[2], pointy[3], pointy[3], pointy[2], pointy[2]]
ys3 = [pointy[4], pointy[5], pointy[5], pointy[4], pointy[4]]
# graficos
plt.close('all')
plt.figure(figsize=(10,10))
#******************************************************
plt.contourf(Y, X, tfa_final, 20, cmap = plt.cm.RdBu_r)
plt.title('Campo Total (nT)', fontsize = 12)
plt.xlabel('East (m)', fontsize = 10)
plt.ylabel('North (m)', fontsize = 10)
plt.plot(ys1,xs1,'g-')
plt.plot(ys2,xs2,'k-')
plt.plot(ys3,xs3,'m-')
plt.colorbar()
#plt.savefig('teste_100_40000_D10.png', format='png')
plt.show()
# +
dici1 = {'nx': nx,
'ny': ny,
'X': X,
'Y': Y,
'ACTn': tfa_final
}
data_e_hora_atuais = datetime.now()
data_e_hora = data_e_hora_atuais.strftime('%d_%m_%Y_%H_%M')
dicionario = {'Data da Modelagem': data_e_hora,
'Tipo de Modelagem': 'Modelagem de prisma',
'números de corpos': 3,
'Coordenadas do prisma 1 (x1, x2, y1, y2, z1, z2)': [pointx[0], pointx[1], pointy[0], pointy[1], pointz[0], pointz[1]],
'Coordenadas do prisma 2 (x1, x2, y1, y2, z1, z2)': [pointx[2], pointx[3], pointy[2], pointy[3], pointz[2], pointz[3]],
'Coordenadas do prisma 3 (x1, x2, y1, y2, z1, z2)': [pointx[4], pointx[5], pointy[4], pointy[5], pointz[4], pointz[5]],
'inclinação': 'positivo',
'Informação da fonte (Mag, Incl, Decl)': [Mi, inc, dec],
'Informação regional (Camp.Geomag, Incl, Decl)': [Fi, I, D]}
print(dicionario)
# -
Data_f = salve_doc.reshape_matrix(dici1)
Data_f
# +
#salve_doc.create_diretorio(dicionario, Data_f)
# -
| codes/tests/.ipynb_checkpoints/Teste_mag_modelo_escada-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/djswoosh/Uni-Bits/blob/main/Heart_Disease_Dataset_Imputations.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="nOfdSHn49g4y" outputId="8138d96e-17c8-4e31-ac85-dfe9185d0d93"
import pandas as pd
df = pd.read_csv("/content/processed.hungarian.csv",
sep= ",",
names = ["age", "sex", "cp", "treatbps", "chol", "fbs",
"reastecg", "thalach", "exang", "oldpeak",
"slope", "ca", "thal", "num"])
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="5D1tOW-LAjhp" outputId="cd032fb3-1e95-4179-dc07-edfafb7d83f6"
df.info()
#Object Types reflect the fact that
#Pandas treats data types where it caanot infer the data type as objects
#Columns with missing data will have a Question Mark, and treated as Strings
#But the remainder of the data in the colums is a Float Type
# + [markdown] id="W_x2HcjGHJlR"
# The Next thing to do is to replace the Question mark with NAN Values
# + id="ZHSvClpNHSQZ"
import numpy as np
def replace_question_mark(val):
if val == "?":
return np.NaN
elif type(val) ==int:
return val
else:
return float (val)
df2 = df.copy()
for (columnName, _) in df2.iteritems():
df2[columnName] = df2[columnName].apply(replace_question_mark)
# + colab={"base_uri": "https://localhost:8080/"} id="UM9zoLZHI-OM" outputId="5b28aad1-b489-4357-8927-75375a8e33ce"
df2.info() #AfterApplicationofabovefunctionPandas can now read the data types.
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="3kB299heJQqp" outputId="f55a648c-c94a-42aa-abc0-da544ce372b6"
df2.head() #Nan Values are treted as legitimate numerical values in Python
#and legitimate Numpy fucntions can be used on them
# + colab={"base_uri": "https://localhost:8080/"} id="zd0WZytQKRCL" outputId="aa5490de-e9d4-4e15-bf91-dcff9dd2b116"
df2.dropna().shape
# + colab={"base_uri": "https://localhost:8080/", "height": 317} id="sdoIHy0uK1BG" outputId="5796d415-ae59-485e-ca06-df0d7fbdfaf5"
df2.describe() #ThisFunctionGenerates a Quick table of statistics
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="HiRWSlP1K3kF" outputId="6c676849-7565-4b33-a081-d82d88320b70"
#Imputation with mean or median Values
import matplotlib.pyplot as plt
chol = df2["chol"]
plt.hist(chol.apply(lambda x: np.mean(chol) if np.isnan(x) else x), bins=range(0,630,30))
plt.xlabel("cholesterol imputation")
plt.ylabel("count")
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="slXriNdCPZ3-" outputId="eeb90d98-9347-46c8-93f0-867a5031b343"
plt.hist(df2["slope"], bins = 5)
plt.xlabel("slope")
plt.ylabel("count"),
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="DfZPmTtBP3tR" outputId="5a81fa42-3b29-48b1-ed01-76c3f0312af0"
plt.hist(df2["slope"].apply(lambda x: 2 if np.isnan(x) else x), bins = 5)
plt.xlabel("slope mode imputation")
plt.ylabel("count"),
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="s5cw95ecSfu5" outputId="327bf1fa-6518-475c-8568-0b25bdd18ede"
#Standardization
stdChol= np.std(chol)
meanChol = np.mean(chol)
chol2 = chol.apply (lambda x: (x-meanChol)/stdChol)
plt.hist(chol2, bins = range(int (min(chol2)), int(max(chol2))+1,1));
| Heart_Disease_Dataset_Imputations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# language: python
# name: python3
# ---
# Diner Robot Inverse Kinematics calculations
# +
import sympy as sp
import numpy as np
import math
# -
# UR5 angles and link lengths:
# +
q1, q2, q3, q4, q5, q6, q7, = sp.symbols('q1 q2 q3 q4 q5 q6 q7')
d = [0.089159, 0, 0, 0.10915, 0.09465, 0.0823]
a = [0.0, -0.425, -0.39225, 0.0, 0.0, 0]
# -
# Transformation Matrix:
# +
T0 = sp.Matrix([[1, 0, 0, 0],
[0, 1, 0, 0],
[0, 0, 1, 0],
[0, 0, 0, 1]])
T1 = sp.Matrix([[sp.cos(q1), 0, -sp.sin(q1), 0],
[sp.sin(q1), 0, sp.cos(q1), 0],
[0, -1, 0, d[0]],
[0, 0, 0, 1]])
#T1 = T1.subs({q1: math.radians(90), q2: math.radians(0), q3: math.radians(0), q4: math.radians(-90), q5: math.radians(0), q6: math.radians(0), q7: math.radians(0)})
T2 = sp.Matrix([[sp.cos(q2), 0, sp.sin(q2), 0],
[sp.sin(q2), 0, -sp.cos(q2), 0],
[0, 1, 0, d[1]],
[0, 0, 0, 1]])
#T2 = T2.subs({q1: math.radians(90), q2: math.radians(0), q3: math.radians(0), q4: math.radians(-90), q5: math.radians(0), q6: math.radians(0), q7: math.radians(0)})
q3 = 0
T3 = sp.Matrix([[sp.cos(q3), 0, sp.sin(q3), 0],
[sp.sin(q3), 0, -sp.cos(q3), 0],
[0, 1, 0, d[2]],
[0, 0, 0, 1]])
#T3 = T3.subs({q1: math.radians(90), q2: math.radians(0), q3: math.radians(0), q4: math.radians(-90), q5: math.radians(0), q6: math.radians(0), q7: math.radians(0)})
T4 = sp.Matrix([[sp.cos(q4), 0, -sp.sin(q4), 0],
[sp.sin(q4), 0, sp.cos(q4), 0],
[0, -1, 0, d[3]],
[0, 0, 0, 1]])
#T4 = T4.subs({q1: math.radians(90), q2: math.radians(0), q3: math.radians(0), q4: math.radians(-90), q5: math.radians(0), q6: math.radians(0), q7: math.radians(0)})
T5 = sp.Matrix([[sp.cos(q5), 0, -sp.sin(q5), 0],
[sp.sin(q5), 0, sp.cos(q5), 0],
[0, -1, 0, d[4]],
[0, 0, 0, 1]])
#T5 = T5.subs({q1: math.radians(90), q2: math.radians(0), q3: math.radians(0), q4: math.radians(-90), q5: math.radians(0), q6: math.radians(0), q7: math.radians(0)})
T6 = sp.Matrix([[sp.cos(q6), 0, sp.sin(q6), 0],
[sp.sin(q6), 0, -sp.cos(q6), 0],
[0, 1, 0, d[5]],
[0, 0, 0, 1]])
#T6= T6.subs({q1: math.radians(90), q2: math.radians(0), q3: math.radians(0), q4: math.radians(-90), q5: math.radians(0), q6: math.radians(0), q7: math.radians(0)})
# -
# Final Transformation Matrix:
# +
T = (T0*T1*T2*T3*T4*T5*T6)
T
# -
# Transformation at start position:
q = sp.Matrix([ [math.radians(90.001)], [math.radians(0.001)], [math.radians(-90.001)], [math.radians(0.001)], [math.radians(0.001)], [math.radians(0.001)]])
Tstart = T.subs({q1: q[0], q2: q[1], q4: q[2], q5: q[3], q6: q[4], q7: q[5] })
Tstart
# Jacobian calculation:
J = sp.Matrix([[-a[1]*sp.sin(q1)*sp.sin(q2) - d[5]*sp.sin(q1)*sp.sin(q5)*sp.cos(q2 + q3 + q4) - a[2]*sp.sin(q1)*sp.sin(q2 + q3) + d[4]*sp.sin(q1)*sp.sin(q2 + q3 + q4) + d[5]*sp.cos(q1)*sp.cos(q5) + d[3]*sp.cos(q1),
-(d[5]*sp.sin(q5)*sp.sin(q2 + q3 + q4) - a[1]*sp.cos(q2) -
a[2]*sp.cos(q2 + q3) + d[4]*sp.cos(q2 + q3 + q4))*sp.cos(q1),
-(d[5]*sp.sin(q5)*sp.sin(q2 + q3 + q4) - a[2] *
sp.cos(q2 + q3) + d[4]*sp.cos(q2 + q3 + q4))*sp.cos(q1),
-(d[5]*sp.sin(q5)*sp.sin(q2 + q3 + q4) +
d[4]*sp.cos(q2 + q3 + q4))*sp.cos(q1),
- d[5]*sp.sin(q1)*sp.sin(q5) + d[5]*sp.cos(q1) *
sp.cos(q5)*sp.cos(q2 + q3 + q4),
0],
[d[5]*sp.sin(q1)*sp.cos(q5) + d[3]*sp.sin(q1) + a[1]*sp.sin(q2)*sp.cos(q1) + d[5]*sp.sin(q5)*sp.cos(q1)*sp.cos(q2 + q3 + q4) + a[2]*sp.sin(q2 + q3)*sp.cos(q1) - d[4]*sp.sin(q2 + q3 + q4)*sp.cos(q1),
-(d[5]*sp.sin(q5)*sp.sin(q2 + q3 + q4) - a[1]*sp.cos(q2) -
a[2]*sp.cos(q2 + q3) + d[4]*sp.cos(q2 + q3 + q4))*sp.sin(q1),
-(d[5]*sp.sin(q5)*sp.sin(q2 + q3 + q4) - a[2] *
sp.cos(q2 + q3) + d[4]*sp.cos(q2 + q3 + q4))*sp.sin(q1),
-(d[5]*sp.sin(q5)*sp.sin(q2 + q3 + q4) +
d[4]*sp.cos(q2 + q3 + q4))*sp.sin(q1),
d[5]*sp.sin(q1)*sp.cos(q5)*sp.cos(q2 + q3 + q4) +
d[5]*sp.sin(q5)*sp.cos(q1),
0],
[0,
a[1]*sp.sin(q2) + d[5]*sp.sin(q5)*sp.cos(q2 + q3 + q4) +
a[2]*sp.sin(q2 + q3) - d[4]*sp.sin(q2 + q3 + q4),
d[5]*sp.sin(q5)*sp.cos(q2 + q3 + q4) + a[2] *
sp.sin(q2 + q3) - d[4]*sp.sin(q2 + q3 + q4),
d[5]*sp.sin(q5)*sp.cos(q2 + q3 + q4) -
d[4]*sp.sin(q2 + q3 + q4),
d[5]*sp.sin(q2 + q3 + q4)*sp.cos(q5),
0],
[0, sp.sin(q1), sp.sin(q1), sp.sin(q1), -sp.sin(q2 + q3 + q4)*sp.cos(q1),
sp.sin(q1)*sp.cos(q5) + sp.sin(q5)*sp.cos(q1)*sp.cos(q2 + q3 + q4)],
[0, -sp.cos(q1), -sp.cos(q1), -sp.cos(q1), -sp.sin(q1)*sp.sin(q2 + q3 + q4),
sp.sin(q1)*sp.sin(q5)*sp.cos(q2 + q3 + q4) - sp.cos(q1)*sp.cos(q5)],
[1, 0, 0, 0, sp.cos(q2 + q3 + q4), sp.sin(q5)*sp.sin(q2 + q3 + q4)]])
J
q = sp.Matrix([ [math.radians(90.001)], [math.radians(0.001)], [math.radians(-90.001)], [math.radians(0.001)], [math.radians(0.001)], [math.radians(0.001)]])
Jstart = J.subs({q1: q[0], q2: q[1], q4: q[2], q5: q[3], q6: q[4], q7: q[5] })
Jstart
# Waypoint Generation:
# +
qdash = sp.Matrix([ [math.radians(0)], [math.radians(0)], [math.radians(0)], [math.radians(0)], [math.radians(0)], [math.radians(0)]])
theta = np.pi/2
while(theta< ((2*np.pi) + np.pi/2)):
Js = J.subs({q1: q[0], q2: q[1], q4: q[2], q5: q[3], q6: q[4], q7: q[5]})
try:
Jp = Js.inv()
except:
print('Singularity occured')
q = (q + qdash).evalf()
radius = 100
Vx = -radius*np.sin(theta)*((2*np.pi) / 40)
Vz = radius*np.cos(theta)*((2*np.pi) / 40)
theta += ((2*np.pi) / 40)
V = sp.Matrix([ [Vx], [0], [Vz], [0], [0], [0]])
qdash=Jp*V
#q_current= q_pervious + 𝑞̇_current . ∆t
q = (q + qdash).evalf()
Tdash = T.subs({q1: q[0], q2: q[1], q4: q[2], q5: q[3], q6: q[4], q7: q[5] })
x = Tdash[0,3]
y = Tdash[1,3]
z = Tdash[2,3]
print('x: ',x)
print('y: ',y)
print('z: ',z)
| UR5Transformation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: wentao
# language: python
# name: wentao
# ---
# # 第二课 词向量
#
# 第二课学习目标
# - 学习词向量的概念
# - 用skip-gram模型训练词向量
# - 学习使用PyTorch dataset和dataloader
# - 学习定义PyTorch模型
# - 学习torch.nn中常见的Module
# - Embedding
# - 学习常见的PyTorch operations
# - bmm
# - logsigmoid
# - 保存和读取PyTorch模型
#
#
# 第二课使用的训练数据可以从以下链接下载到。
#
# 链接:https://pan.baidu.com/s/1tFeK3mXuVXEy3EMarfeWvg 密码:<PASSWORD>
#
# 在这一份notebook中,我们会(尽可能)尝试复现论文[Distributed Representations of Words and Phrases and their Compositionality](http://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf)中训练词向量的方法. 我们会实现Skip-gram模型,并且使用论文中negative sampling的目标函数。
#
# 这篇论文有很多模型实现的细节,这些细节对于词向量的好坏至关重要。我们虽然无法完全复现论文中的实验结果,主要是由于计算资源等各种细节原因,但是我们还是可以大致展示如何训练词向量。
#
# 以下是一些我们没有实现的细节
# - subsampling:参考论文section 2.3
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.utils.data as tud
from torch.nn.parameter import Parameter
from collections import Counter
import numpy as np
import random
import math
import pandas as pd
import scipy
import sklearn
from sklearn.metrics.pairwise import cosine_similarity
USE_CUDA = torch.cuda.is_available()
# 为了保证实验结果可以复现,我们经常会把各种random seed固定在某一个值
random.seed(53113)
np.random.seed(53113)
torch.manual_seed(53113)
if USE_CUDA:
torch.cuda.manual_seed(53113)
# 设定一些超参数
K = 100 # number of negative samples
C = 3 # nearby words threshold
NUM_EPOCHS = 2 # The number of epochs of training
MAX_VOCAB_SIZE = 30000 # the maximum vocabulary size
BATCH_SIZE = 128 # the batch size
LEARNING_RATE = 0.2 # the initial learning rate
EMBEDDING_SIZE = 100
LOG_FILE = "word-embedding.log"
print(USE_CUDA)
# -
# ### 数据预处理
# - 从文本文件中读取所有的文字,通过这些文本创建一个vocabulary
# - 由于单词数量可能太大,我们只选取最常见的MAX_VOCAB_SIZE个单词
# - 我们添加一个UNK单词表示所有不常见的单词
# - 我们需要记录单词到index的mapping,以及index到单词的mapping,单词的count,单词的(normalized) frequency,以及单词总数。
# +
with open("text8/text8.train.txt", "r") as fin:
text = fin.read()
# tokenize函数,把一篇文本转化成一个个单词
def word_tokenize(text):
return text.split()
text = [w for w in word_tokenize(text.lower())]
vocab = dict(Counter(text).most_common(MAX_VOCAB_SIZE-1))#Counter计算单词频数,count最高取vocab_size-1个,最后一个取unk
vocab["<unk>"] = len(text) - np.sum(list(vocab.values()))#unk频率单独计算
idx_to_word = [word for word in vocab.keys()]
word_to_idx = {word:i for i, word in enumerate(idx_to_word)}#enumerate自带index
word_counts = np.array([count for count in vocab.values()], dtype=np.float32)
word_freqs = word_counts / np.sum(word_counts)
word_freqs = word_freqs ** (3./4.)
word_freqs = word_freqs / np.sum(word_freqs) # normalization
VOCAB_SIZE = len(idx_to_word)
# -
# ### 实现Dataloader
#
# 一个dataloader需要以下内容:
#
# - 把所有text编码成数字,然后用subsampling预处理这些文字。
# - 保存vocabulary,单词count,normalized word frequency
# - 每个iteration sample一个中心词
# - 根据当前的中心词返回context单词
# - 根据中心词sample一些negative单词
# - 返回单词的counts
#
# 这里有一个好的tutorial介绍如何使用[PyTorch dataloader](https://pytorch.org/tutorials/beginner/data_loading_tutorial.html).
# 为了使用dataloader,我们需要定义以下两个function:
#
# - ```__len__``` function需要返回整个数据集中有多少个item
# - ```__get__``` 根据给定的index返回一个item
#
# 有了dataloader之后,我们可以轻松随机打乱整个数据集,拿到一个batch的数据等等。
class WordEmbeddingDataset(tud.Dataset): #继承类,创建specific dataset
def __init__(self, text, word_to_idx, idx_to_word, word_freqs, word_counts):
''' text: a list of words, all text from the training dataset
word_to_idx: mapping word to idx
idx_to_word: list of words
word_freq: the frequency of each word
word_counts: the word counts
'''
super(WordEmbeddingDataset, self).__init__()
self.text_encoded = [word_to_idx.get(t, VOCAB_SIZE-1) for t in text]#找text每个单词的idx
self.text_encoded = torch.Tensor(self.text_encoded).long()#must be integars
self.word_to_idx = word_to_idx
self.idx_to_word = idx_to_word
self.word_freqs = torch.Tensor(word_freqs)
self.word_counts = torch.Tensor(word_counts)
def __len__(self):
''' 返回整个数据集的长度(所有单词的个数)
'''
return len(self.text_encoded)
def __getitem__(self, idx):
''' 这个function返回以下数据用于训练
- 中心词
- 这个单词附近的(positive)单词
- 随机采样的K个单词作为negative sample
- K,C: previously defined hyper parameters
'''
center_word = self.text_encoded[idx]
pos_indices = list(range(idx-C, idx)) + list(range(idx+1, idx+C+1))#omit idx
pos_indices = [i%len(self.text_encoded) for i in pos_indices]#超出范围取余
pos_words = self.text_encoded[pos_indices]
neg_words = torch.multinomial(self.word_freqs, K * pos_words.shape[0], True)#每个pos_word取K个neg_word
return center_word, pos_words, neg_words
# 创建dataset和dataloader
dataset = WordEmbeddingDataset(text, word_to_idx, idx_to_word, word_freqs, word_counts)
dataloader = tud.DataLoader(dataset, batch_size=BATCH_SIZE, shuffle=True, num_workers=4)
dataset.text_encoded[:100]
# ### 定义PyTorch模型
# + tags=[]
class EmbeddingModel(nn.Module):
def __init__(self, vocab_size, embed_size):
''' 初始化输出和输出embedding
'''
super(EmbeddingModel, self).__init__() #初始化
self.vocab_size = vocab_size #存储初始信息
self.embed_size = embed_size
initrange = 0.5 / self.embed_size
self.in_embed = nn.Embedding(self.vocab_size, self.embed_size, sparse=False)
self.in_embed.weight.data.uniform_(-initrange, initrange)
self.out_embed = nn.Embedding(self.vocab_size, self.embed_size, sparse=False)
self.out_embed.weight.data.uniform_(-initrange, initrange)
def forward(self, input_labels, pos_labels, neg_labels):
'''
input_labels: 中心词, [batch_size]
pos_labels: 中心词周围 context window 出现过的单词 [batch_size * (window_size * 2)]
neg_labelss: 中心词周围没有出现过的单词,从 negative sampling 得到 [batch_size, (window_size * 2 * K)]
return: -loss
'''
batch_size = input_labels.size(0)
input_embedding = self.in_embed(input_labels) # batch_size * embed_size
pos_embedding = self.out_embed(pos_labels) # batch_size * (2*C) * embed_size
neg_embedding = self.out_embed(neg_labels) # batch_size * (2*C * K) * embed_size
#batch matrix multiply
log_pos = torch.bmm(pos_embedding, input_embedding.unsqueeze(2)).squeeze() # B * (2*C)
log_neg = torch.bmm(neg_embedding, -input_embedding.unsqueeze(2)).squeeze() # B * (2*C*K)
#unsqueeze(2)在第3个维度加一
#squeeze(2)去掉第3个维度
log_pos = F.logsigmoid(log_pos).sum(1)
log_neg = F.logsigmoid(log_neg).sum(1) # batch_size
# objective function
loss = log_pos + log_neg
return -loss #[batch_size]
#一般取input embedding作为word embedding
def input_embeddings(self):
return self.in_embed.weight.data.cpu().numpy()
# -
# 定义一个模型以及把模型移动到GPU
# + tags=[]
model = EmbeddingModel(VOCAB_SIZE, EMBEDDING_SIZE)
if USE_CUDA:
model = model.cuda()
# -
# 下面是评估模型的代码,以及训练模型的代码
# + tags=[]
def evaluate(filename, embedding_weights):
if filename.endswith(".csv"):
data = pd.read_csv(filename, sep=",")
else:
data = pd.read_csv(filename, sep="\t")
human_similarity = []
model_similarity = []
for i in data.iloc[:, 0:2].index:
word1, word2 = data.iloc[i, 0], data.iloc[i, 1]
if word1 not in word_to_idx or word2 not in word_to_idx:
continue
else:
word1_idx, word2_idx = word_to_idx[word1], word_to_idx[word2]
word1_embed, word2_embed = embedding_weights[[word1_idx]], embedding_weights[[word2_idx]]
model_similarity.append(float(sklearn.metrics.pairwise.cosine_similarity(word1_embed, word2_embed)))
human_similarity.append(float(data.iloc[i, 2]))
return scipy.stats.spearmanr(human_similarity, model_similarity)# , model_similarity
def find_nearest(word):
index = word_to_idx[word]
embedding = embedding_weights[index]
cos_dis = np.array([scipy.spatial.distance.cosine(e, embedding) for e in embedding_weights])
return [idx_to_word[i] for i in cos_dis.argsort()[:10]]
# -
# 训练模型:
# - 模型一般需要训练若干个epoch
# - 每个epoch我们都把所有的数据分成若干个batch
# - 把每个batch的输入和输出都包装成cuda tensor
# - forward pass,通过输入的句子预测每个单词的下一个单词
# - 清空模型当前gradient
# - 用模型的预测和正确的下一个单词计算cross entropy loss
# - backward pass
# - 更新模型参数
# - 每隔一定的iteration输出模型在当前iteration的loss,以及在验证数据集上做模型的评估
optimizer = torch.optim.SGD(model.parameters(), lr=LEARNING_RATE)
# loss function defined in the model class
for e in range(NUM_EPOCHS):
for i, (input_labels, pos_labels, neg_labels) in enumerate(dataloader):
#input_labels = input_labels.long()
#pos_labels = pos_labels.long()
#neg_labels = neg_labels.long()
if USE_CUDA:
input_labels = input_labels.cuda()
pos_labels = pos_labels.cuda()
neg_labels = neg_labels.cuda()
optimizer.zero_grad()
loss = model(input_labels, pos_labels, neg_labels).mean()
loss.backward()
optimizer.step()
if i % 100 == 0:
with open(LOG_FILE, "a") as fout:
fout.write("epoch: {}, iter: {}, loss: {}\n".format(e, i, loss.item()))
#print("epoch: {}, iter: {}, loss: {}".format(e, i, loss.item()))
if i % 2000 == 0:
embedding_weights = model.input_embeddings()
sim_simlex = evaluate("simlex-999.txt", embedding_weights)
sim_men = evaluate("men.txt", embedding_weights)
sim_353 = evaluate("wordsim353.csv", embedding_weights)
with open(LOG_FILE, "a") as fout:
print("epoch: {}, iteration: {}, simlex-999: {}, men: {}, sim353: {}, nearest to monster: {}\n".format(
e, i, sim_simlex, sim_men, sim_353, find_nearest("monster")))
fout.write("epoch: {}, iteration: {}, simlex-999: {}, men: {}, sim353: {}, nearest to monster: {}\n".format(
e, i, sim_simlex, sim_men, sim_353, find_nearest("monster")))
embedding_weights = model.input_embeddings()
np.save("embedding-{}".format(EMBEDDING_SIZE), embedding_weights)
torch.save(model.state_dict(), "embedding-{}.th".format(EMBEDDING_SIZE))
# + jupyter={"outputs_hidden": true}
model.load_state_dict(torch.load("embedding-{}.th".format(EMBEDDING_SIZE)))
# -
# ## 在 MEN 和 Simplex-999 数据集上做评估
embedding_weights = model.input_embeddings()
print("simlex-999", evaluate("simlex-999.txt", embedding_weights))
print("men", evaluate("men.txt", embedding_weights))
print("wordsim353", evaluate("wordsim353.csv", embedding_weights))
# ## 寻找nearest neighbors
for word in ["good", "fresh", "monster", "green", "like", "america", "chicago", "work", "computer", "language"]:
print(word, find_nearest(word))
# ## 单词之间的关系
man_idx = word_to_idx["man"]
king_idx = word_to_idx["king"]
woman_idx = word_to_idx["woman"]
embedding = embedding_weights[woman_idx] - embedding_weights[man_idx] + embedding_weights[king_idx]
cos_dis = np.array([scipy.spatial.distance.cosine(e, embedding) for e in embedding_weights])
for i in cos_dis.argsort()[:20]:
print(idx_to_word[i])
| 2/word-embedding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 8.0
# language: ''
# name: sagemath
# ---
# ## Chudnovski
def F(n):
return ((-1)^n*(factorial(6*n))*(545140134*n+13591409))/(factorial(3*n)*(factorial(n)^3)*(640320^(3*n)))
def pi_chudnovski(digits=100):
k = 0
S = 0
while 1:
S += F(k)
if floor(abs(10^digits*F(k))) == 0:
break
k += 1
return (426880*sqrt(10005).n(digits=digits))/S,k
# ## Algoritmo BBP
# +
def F0(j,n):
S =RR(0.0)
k =0
while k <= n:
r = 8*k+j
S += RR(power_mod(16,n-k,r)/r)-floor(RR(power_mod(16,n-k,r)/r))
k += 1
return RR(S)
def F1(j,n):
S =RR(0.0)
k =n+1
while 1:
r = 8*k+j
nS = S+ RR(16^(n-k)/r)
if S == nS:
break
else:
S = nS
k += 1
return RR(S)
def S(j,n):
return RR(F0(j,n)+F1(j,n))
def cifra_pi(n):
n -= 1
x = (4*RR(S(1,n))-2*RR(S(4,n))-RR(S(5,n))-RR(S(6,n)))
return (x-floor(x)).str(base=16)
# -
# La cadena $C$ contiene, para poder comprobar los resultados obtenidos con el algoritmo, cifras hexadecimales de $\pi$ empezando después del punto.
C = '243F6A8885A308D313198A2E03707344A4093822299F31D0082EFA98EC4E6C89452821E\
638D01377BE5466CF34E90C6CC0AC29B7C97C50DD3F84D5B5B54709179216D5D98979FB1\
BD1310BA698DFB5AC2FFD72DBD01ADFB7B8E1AFED6A267E96BA7C9045F12C7F9924A1994\
7B3916CF70801F2E2858EFC16636920D871574E69A458FEA3F4933D7E0D95748F728EB65\
8718BCD5882154AEE7B54A41DC25A59B59C30D5392AF26013C5D1B023286085F0CA41791\
8B8DB38EF8E79DCB0603A180E6C9E0E8BB01E8A3ED71577C1BD314B2778AF2FDA55605C6\
0E65525F3AA55AB945748986263E8144055CA396A2AAB10B6B4CC5C341141E8CEA15486A\
F7C72E993B3EE1411636FBC2A2BA9C55D741831F6CE5C3E169B87931EAFD6BA336C24CF5\
C7A325381289586773B8F48986B4BB9AFC4BFE81B6628219361D809CCFB21A991487CAC6\
05DEC8032EF845D5DE98575B1DC262302EB651B8823893E81D396ACC50F6D6FF383F4423\
92E0B4482A484200469C8F04A9E1F9B5E21C66842F6E96C9A670C9C61ABD388F06A51A0D\
2D8542F68960FA728AB5133A36EEF0B6C137A3BE4BA3BF0507EFB2A98A1F1651D39AF017\
666CA593E82430E888CEE8619456F9FB47D84A5C33B8B5EBEE06F75D885C12073401A449\
F56C16AA64ED3AA62363F77061BFEDF72429B023D37D0D724D00A1248DB0FEAD349F1C09\
B075372C980991B7B25D479D8F6E8DEF7E3FE501AB6794C3B976CE0BD04C006BAC1A94FB\
6409F60C45E5C9EC2196A246368FB6FAF3E6C53B51339B2EB3B52EC6F6DFC511F9B30952\
CCC814544AF5EBD09BEE3D004DE334AFD660F2807192E4BB3C0CBA85745C8740FD20B5F3\
9B9D3FBDB5579C0BD1A60320AD6A100C6402C7279679F25FEFB1FA3CC8EA5E9F8DB3222F\
83C7516DFFD616B152F501EC8AD0552AB323DB5FAFD23876053317B483E00DF829E5C57B\
BCA6F8CA01A87562EDF1769DBD542A8F6287EFFC3AC6732C68C4F5573695B27B0BBCA58C\
8E1FFA35DB8F011A010FA3D98FD2183B84AFCB56C2DD1D35B9A53E479B6F84565D28E49B\
C4BFB9790E1DDF2DAA4CB7E3362FB1341CEE4C6E8EF20CADA36774C01D07E9EFE2BF11FB\
495DBDA4DAE909198EAAD8E716B93D5A0D08ED1D0AFC725E08E3C5B2F8E7594B78FF6E2F\
BF2122B648888B812900DF01C4FAD5EA0688FC31CD1CFF191B3A8C1AD2F2F2218BE0E177\
7EA752DFE8B021FA1E5A0CC0FB56F74E818ACF3D6CE89E299B4A84FE0FD13E0B77CC43B8\
1D2ADA8D9165FA2668095770593CC7314211A1477E6AD206577B5FA86C75442F5FB9D35C\
FEBCDAF0C7B3E89A0D6411BD3AE1E7E4900250E2D2071B35E226800BB57B8E0AF2464369\
BF009B91E5563911D59DFA6AA78C14389D95A537F207D5BA202E5B9C5832603766295CFA\
911C819684E734A41B3472DCA7B14A94A1B5100529A532915D60F573FBC9BC6E42B60A47\
681E6740008BA6FB5571BE91FF296EC6B2A0DD915B6636521E7B9F9B6FF34052EC585566\
453B02D5DA99F8FA108BA47996E85076A4B7A70E9B5B32944DB75092EC4192623AD6EA6B\
049A7DF7D9CEE60B88FEDB266ECAA8C71699A17FF5664526CC2B19EE1193602A575094C2\
9A0591340E4183A3E3F54989A5B429D656B8FE4D699F73FD6A1D29C07EFE830F54D2D38E\
6F0255DC14CDD20868470EB266382E9C6021ECC5E09686B3F3EBAEFC93C9718146B6A70A\
1687F358452A0E286B79C5305AA5007373E07841C7FDEAE5C8E7D44EC5716F2B8B03ADA3\
7F0500C0DF01C1F040200B3FFAE0CF51A3CB574B225837A58DC0921BDD19113F97CA92FF\
69432477322F547013AE5E58137C2DADCC8B576349AF3DDA7A94461460FD0030EECC8C73\
EA4751E41E238CD993BEA0E2F3280BBA1183EB3314E548B384F6DB9086F420D03F60A04B\
F2CB8129024977C795679B072BCAF89AFDE9A771FD9930810B38BAE12DCCF3F2E5512721\
F2E6B7124501ADDE69F84CD877A5847187408DA17BC9F9ABCE94B7D8CEC7AEC3ADB851DF\
A63094366C464C3D2EF1C18473215D908DD433B3724C2BA1612A14D432A65C4515094000\
2133AE4DD71DFF89E10314E5581AC77D65F11199B043556F1D7A3C76B3C11183B5924A50\
9F28FE6ED97F1FBFA9EBABF2C1E153C6E86E34570EAE96FB1860E5E0A5A3E2AB3771FE71\
C4E3D06FA2965DCB999E71D0F803E89D65266C8252E4CC9789C10B36AC6150EBA94E2EA7\
8A5FC3C531E0A2DF4F2F74EA7361D2B3D1939260F19C279605223A708F71312B6EBADFE6\
EEAC31F66E3BC4595A67BC883B17F37D1018CFF28C332DDEFBE6C5AA56558218568AB980\
2EECEA50FDB2F953B2AEF7DAD5B6E2F841521B62829076170ECDD4775619F151013CCA83\
0EB61BD960334FE1EAA0363CFB5735C904C70A239D59E9E0BCBAADE14EECC86BC60622CA\
79CAB5CABB2F3846E648B1EAF19BDF0CAA02369B9655ABB5040685A323C2AB4B3319EE9D\
5C021B8F79B540B19875FA09995F7997E623D7DA8F837889A97E32D7711ED935F1668128\
10E358829C7E61FD696DEDFA17858BA9957F584A51B2272639B83C3FF1AC24696CDB30AE\
B532E30548FD948E46DBC312858EBF2EF34C6FFEAFE28ED61EE7C3C735D4A14D9E864B7E\
342105D14203E13E045EEE2B6A3AAABEADB6C4F15FACB4FD0C742F442EF6ABBB5654F3B1\
D41CD2105D81E799E86854DC7E44B476A3D816250CF62A1F25B8D2646FC8883A0C1C7B6A\
37F1524C369CB749247848A0B5692B285095BBF00AD19489D1462B17423820E0058428D2\
A0C55F5EA1DADF43E233F70613372F0928D937E41D65FECF16C223BDB7CDE3759CBEE746\
04085F2A7CE77326EA607808419F8509EE8EFD85561D99735A969A7AAC50C06C25A04ABF\
C800BCADC9E447A2EC3453484FDD567050E1E9EC9DB73DBD3105588CD675FDA79E367434\
0C5C43465713E38D83D28F89EF16DFF20153E21E78FB03D4AE6E39F2BDB83ADF7E93D5A6\
8948140F7F64C261C94692934411520F77602D4F7BCF46B2ED4A20068D40824713320F46\
A43B7D4B7500061AF1E39F62E9724454614214F74BF8B88404D95FC1D96B591AF70F4DDD\
366A02F45BFBC09EC03BD97857FAC6DD031CB850496EB27B355FD3941DA2547E6ABCA0A9\
A28507825530429F40A2C86DAE9B66DFB68DC1462D7486900680EC0A427A18DEE4F3FFEA\
2E887AD8CB58CE0067AF4D6B6AACE1E7CD3375FECCE78A399406B2A4220FE9E35D9F385B\
9EE39D7AB3B124E8B1DC9FAF74B6D185626A36631EAE397B23A6EFA74DD5B43326841E7F\
7CA7820FBFB0AF54ED8FEB397454056ACBA48952755533A3A20838D87FE6BA9B7D096954\
B55A867BCA1159A58CCA9296399E1DB33A62A4A563F3125F95EF47E1C9029317CFDF8E80\
204272F7080BB155C05282CE395C11548E4C66D2248C1133FC70F86DC07F9C9EE41041F0\
F404779A45D886E17325F51EBD59BC0D1F2BCC18F41113564257B7834602A9C60DFF8E8A\
31F636C1B0E12B4C202E1329EAF664FD1CAD181156B2395E0333E92E13B240B62EEBEB92\
285B2A20EE6BA0D99DE720C8C2DA2F728D012784595B794FD647D0862E7CCF5F05449A36\
F877D48FAC39DFD27F33E8D1E0A476341992EFF743A6F6EABF4F8FD37A812DC60A1EBDDF\
8991BE14CDB6E6B0DC67B55106D672C372765D43BDCD0E804F1290DC7CC00FFA3B5390F9\
2690FED0B667B9FFBCEDB7D9CA091CF0BD9155EA3BB132F88515BAD247B9479BF763BD6E\
B37392EB3CC1159798026E297F42E312D6842ADA7C66A2B3B12754CCC782EF11C6A12423\
7B79251E706A1BBE64BFB63501A6B101811CAEDFA3D25BDD8E2E1C3C9444216590A12138\
6D90CEC6ED5ABEA2A64AF674EDA86A85FBEBFE98864E4C3FE9DBC8057F0F7C08660787BF\
86003604DD1FD8346F6381FB07745AE04D736FCCC83426B33F01EAB71B08041873C005E5\
F77A057BEBDE8AE2455464299BF582E614E58F48FF2DDFDA2F474EF388789BDC25366F9C\
3C8B38E74B475F25546FCD9B97AEB26618B1DDF84846A0E79915F95E2466E598E20B4577\
08CD55591C902DE4CB90BACE1BB8205D011A862487574A99EB77F19B6E0A9DC09662D09A\
1C4324633E85A1F0209F0BE8C4A99A0251D6EFE101AB93D1D0BA5A4DFA186F20F2868F16\
9DCB7DA83573906FEA1E2CE9B4FCD7F5250115E01A70683FAA002B5C40DE6D0279AF88C2\
7773F8641C3604C0661A806B5F0177A28C0F586E0006058AA30DC7D6211E69ED72338EA6\
353C2DD94C2C21634BBCBEE5690BCB6DEEBFC7DA1CE591D766F05E4094B7C018839720A3\
D7C927C2486E3725F724D9DB91AC15BB4D39EB8FCED54557808FCA5B5D83D7CD34DAD0FC\
41E50EF5EB161E6F8A28514D96C51133C6FD5C7E756E14EC4362ABFCEDDC6C837D79A323\
492638212670EFA8E406000E03A39CE37D3FAF5CFABC277375AC52D1B5CB0679E4FA3374\
2D382274099BC9BBED5118E9DBF0F7315D62D1C7EC700C47BB78C1B6B21A19045B26EB1B\
E6A366EB45748AB2FBC946E79C6A376D26549C2C8530FF8EE468DDE7DD5730A1D4CD04DC\
62939BBDBA9BA4650AC9526E8BE5EE304A1FAD5F06A2D519A63EF8CE29A86EE22C089C2B\
843242EF6A51E03AA9CF2D0A483C061BA9BE96A4D8FE51550BA645BD62826A2F9A73A3AE\
14BA99586EF5562E9C72FEFD3F752F7DA3F046F6977FA0A5980E4A91587B086019B09E6A\
D3B3EE593E990FD5A9E34D7972CF0B7D9022B8B5196D5AC3A017DA67DD1CF3ED67C7D2D2\
81F9F25CFADF2B89B5AD6B4725A88F54CE029AC71E019A5E647B0ACFDED93FA9BE8D3C48\
D283B57CCF8D5662979132E28785F0191ED756055F7960E44E3D35E8C15056DD488F46DB\
A03A161250564F0BDC3EB9E153C9057A297271AECA93A072A1B3F6D9B1E6321F5F59C66F\
B26DCF3197533D928B155FDF5035634828ABA3CBB28517711C20AD9F8ABCC5167CCAD925\
F4DE817513830DC8E379D58629320F991EA7A90C2FB3E7BCE5121CE64774FBE32A8B6E37\
EC3293D4648DE53696413E680A2AE0810DD6DB22469852DFD09072166B39A460A6445C0D\
D586CDECF1C20C8AE5BBEF7DD1B588D40CCD2017F6BB4E3BBDDA26A7E3A59FF453E350A4\
4BCB4CDD572EACEA8FA6484BB8D6612AEBF3C6F47D29BE463542F5D9EAEC2771BF64E637\
0740E0D8DE75B1357F8721671AF537D5D4040CB084EB4E2CC34D2466A0115AF84E1B0042\
895983A1D06B89FB4CE6EA0486F3F3B823520AB82011A1D4B277227F8611560B1E7933FD\
CBB3A792B344525BDA08839E151CE794B2F32C9B7A01FBAC9E01CC87EBCC7D1F6CF0111C\
3A1E8AAC71A908749D44FBD9AD0DADECBD50ADA380339C32AC69136678DF9317CE0B12B4\
FF79E59B743F5BB3AF2D519FF27D9459CBF97222C15E6FC2A0F91FC719B941525FAE5936\
1CEB69CEBC2A8645912BAA8D1B6C1075EE3056A0C10D25065CB03A442E0EC6E0E1698DB3\
B4C98A0BE3278E9649F1F9532E0D392DFD3A0342B8971F21E1B0A74414BA3348CC5BE712\
0C37632D8DF359F8D9B992F2EE60B6F470FE3F11DE54CDA541EDAD891CE6279CFCD3E7E6\
F1618B166FD2C1D05848FD2C5F6FB2299F523F357A632762393A8353156CCCD02ACF0816\
25A75EBB56E16369788D273CCDE96629281B949D04C50901B71C65614E6C6C7BD327A140\
A45E1D006C3F27B9AC9AA53FD62A80F00BB25BFE235BDD2F671126905B2040222B6CBCF7\
CCD769C2B53113EC01640E3D338ABBD602547ADF0BA38209CF746CE7677AFA1C52075606\
085CBFE4E8AE88DD87AAAF9B04CF9AA7E1948C25C02FB8A8C01C36AE4D6EBE1F990D4F86\
9A65CDEA03F09252DC208E69FB74E6132CE77E25B578FDFE33AC372E6'
| 2_Curso/Laboratorio/SAGE-noteb/IPYNB/APROX/72-APROX-chudnovski-bbp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JSJeong-me/YOLOv5_Practitioner_Guide_2/blob/main/0-YOLOv5.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="BkBaKWeOPXep" outputId="494150e6-edf5-4eed-efeb-ff9f327fbc25"
pip install -r https://raw.githubusercontent.com/ultralytics/yolov5/master/requirements.txt
# + id="-joA4toHRJdn"
import torch
import cv2
import numpy as np
from google.colab.patches import cv2_imshow
# + colab={"base_uri": "https://localhost:8080/", "height": 349, "referenced_widgets": ["ead7b5fe70074044bc654b331a72da4c", "a6577ea8dd90433ea622db5c35c094d1", "99f9123286f6413388ffafe2a34177d2", "bb5c929b29a54ce8ae96dfe4339e0815", "89aa61b3c05f42bbbaf5421d0a97606b", "8843b20f73b146459e00d1766232c1da", "<KEY>", "bac5c6939f7f48768d12a086ca7ab02a", "<KEY>", "<KEY>", "<KEY>"]} id="pI8wjBxKRNrx" outputId="6919b691-2c0d-4d76-8343-14d296a331c5"
# Model
model = torch.hub.load('ultralytics/yolov5', 'yolov5s')
# + id="M4Ac2hlhRvgC"
img = cv2.imread('traffic-signal-1.jpg')[..., ::-1]
# + id="l-DrccbUR2Eo"
cv2_imshow(img)
# + id="a4swRrJ2TS8q"
model.classes=[9]
# + id="tyIbPmo9UvGu"
model.conf = 0.8
# + id="jFJmp-kMR72e"
results = model(img, size=920)
# + colab={"base_uri": "https://localhost:8080/"} id="uPChwcUYSlnK" outputId="af3084ac-278c-4bfc-c593-1bcd16f680e7"
results.save()
# + id="cYm3R6lgUYqI"
crops_image = results.pandas().xyxy[0]
# + colab={"base_uri": "https://localhost:8080/", "height": 112} id="mg73yrDWUnQ4" outputId="e3504fa7-b5e0-45a4-f069-39f7320397cd"
crops_image
# + colab={"base_uri": "https://localhost:8080/", "height": 112} id="_zT3mFzWVMfs" outputId="49c67a9c-3bdf-42ec-e769-910566c44d02"
results.pandas().xyxy[0].sort_values('xmin')
# + id="wLUjd26kV5e4"
x1, y1, x2, y2 = int(crops_image.iloc[1,0]), int(crops_image.iloc[1,1]), int(crops_image.iloc[1,2]), int(crops_image.iloc[1,3])
# + id="HBrT7K_ZWaz4"
signals = img[y1:y2, x1:x2]
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="llM_1Pb2Wmi8" outputId="c98b4c63-ffb8-409d-c610-e0be539a7309"
cv2_imshow(signals)
| 0-YOLOv5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Demonstrate Basic Querying of DocumentDB
#
# ### Prerequisite:
# 1. Install DocumentDB Python SDK (pip install pydocumentdb)
# 1. Create DocumentDB account and Document DB database from Azure portal
# 1. Download "DocumentDB Migration tool" from [here](http://www.microsoft.com/downloads/details.aspx?FamilyID=cda7703a-2774-4c07-adcc-ad02ddc1a44d)
# 1. Import JSON data (volcano data) stored on a local file into documentDB with following command parameters to the migration tool. You can also use the GUI tool and enter the source and target location parameters from below.
#
# <code>/s:JsonFile /s.Files:[JSON File Location] /t:DocumentDBBulk /t.ConnectionString:AccountEndpoint=https://[DocDBAccountName].documents.azure.com:443/;AccountKey=[[KEY];Database=volcano /t.Collection:volcano1</code>
#
# Copy of volcano data also be found on a blob: https://cahandson.blob.core.windows.net/samples/volcano.json
#
# Execute rest of the code.
import pydocumentdb.documents as documents
import pydocumentdb.document_client as document_client
import pydocumentdb.errors as errors
#Doc DB access parameters.
# You can find the DocDB Account name and "Key" on Azure Portal.
# ReadyOnly key is adequate if you are not writing new records
masterKey = 'ENTER DOC DB Master KEY'
host = 'https://[ENTER DOCDB ACCOUNT NAME].documents.azure.com:443'
db = u'volcano'
collection = 'volcano1'
# client object is the main object to operate with Doc DB
client = document_client.DocumentClient(host,{'masterKey': masterKey})
# Get the pointer to the database you want
database = next((data for data in client.ReadDatabases() if data['id'] == db))
# Get the link to the collection within the database
coll = next((coll for coll in client.ReadCollections(database['_self']) if coll['id'] == collection))
# +
# Use the Doc DB SQL like query language.
# Cheat sheet for DocDB SQL: https://azure.microsoft.com/en-us/documentation/articles/documentdb-sql-query-cheat-sheet/
# Query tries to get list of volcanoes within 300 kms from a given coordinates (Redmond, WA in this case)
# Uses Geospatial Built-in functions ST_DISTANCE( point1, point2 )
query = u'SELECT * \
FROM volcanoes v \
WHERE ST_DISTANCE(v.Location, { \
"type": "Point", \
"coordinates": [-122.19, 47.36] \
}) < 300 * 1000 \
AND v.Type = "Stratovolcano" \
AND v["Last Known Eruption"] = "Last known eruption from 1800-1899, inclusive"'
# -
query
# Run the query
docs = list(client.QueryDocuments(coll['_self'], {'query': query, 'parameters':[]}))
docs
| Data-Science-Virtual-Machine/Samples/Notebooks/DocumentDBSample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"is_executing": false}
from parkinglot.src.parking import *
lot = Lot.create_lot()
print(f"Can park at regular spot? {lot.can_park(SpotType.regular)}")
ticket = lot.park(SpotType.regular, Car("<NAME>", "Red"))
print(ticket)
fare = lot.exit(ticket)
print(fare)
| parkinglot/src/parking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Passing Numpy Arrays to the Option Classes to Generate Multiple Prices
# %load_ext autoreload
# %autoreload 2
import finoptions as fo
import numpy as np
# ## Calculate GBS call option prices across multiple strike prices
K = np.arange(1,20)
opt = fo.GBSOption(10.0, K, 1.0, 0.02, 0.01, 0.1)
opt.call()
opt.put()
opt.greeks()
# Summary method shortens the array representations of the parameters passed and the results returned to make it neater
opt.summary()
# ## Trying Larger Arrays
S = np.arange(8,12, 0.01)
S.shape
K = S - 2
mopt = fo.GBSOption(S, K, 1.0, 0.02, 0.01, 0.1)
# %%timeit
c = mopt.call()
# %%timeit
p = mopt.put()
# %%timeit
mopt.greeks()
mopt.summary()
mopt
# ## Calculating Multiple Implied Volatilities
K = np.array([8., 10., 12.])
vopt = fo.GBSOption(10, K, 1.0, 0.02, 0.0)
vopt
vopt.volatility(K-5)
obj = getattr(ed,"GBSOption")
obj(10,8,1,0.02,0.01,0.2)
# # Tests
# %load_ext autoreload
# %autoreload 2
import finoptions as fo
import numpy as np
# ## BlackSholesOption
S = 10
K = np.arange(8,12, 0.5)
t = 1
r = 0.02
b = 0.01
sigma = 0.2
opt = fo.BlackScholesOption(S, K, t, r, b, sigma)
opt.call()
opt.put()
opt.summary()
opt.greeks()
# ## Black76Option
S = 10
K = np.arange(8,12, 0.5)
t = 1
r = 0.02
b = 0.01
sigma = 0.2
opt = fo.Black76Option(S, K, t, r, sigma)
opt.call()
opt.put()
opt.summary()
opt.greeks()
opt.volatility(1)
# ## MiltersenSchwartzOption
opt = fo.MiltersenSchwartzOption(
Pt=np.exp(-0.05 / 4),
FT=95,
K=np.arange(79,82),
t=1 / 4,
T=1 / 2,
sigmaS=0.2660,
sigmaE=0.2490,
sigmaF=0.0096,
rhoSE=0.805,
rhoSF=0.0805,
rhoEF=0.1243,
KappaE=1.045,
KappaF=0.200,
)
opt.call()
opt.put()
opt.summary()
opt.greeks()
# ## RollGeskeWhaleyOption
opt = fo.basic_american_options.RollGeskeWhaleyOption(S=80, K=np.arange(70,90), t=1/3, td=1/4, r=0.06, D=4, sigma=0.30)
# ## BAWAmericanApproxOption
opt = fo.basic_american_options.BAWAmericanApproxOption(
S=100, K=np.arange(70,90), t=0.5, r=0.10, b=0, sigma=0.25
)
opt = fo.basic_american_options.BSAmericanApproxOption(
S=100, K=np.arange(70,90), t=0.5, r=0.10, b=0, sigma=0.25
)
| notebooks/A2_Passing_Arrays.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('..')
# %load_ext autoreload
# %autoreload 2
# +
import numpy as np
import math
import matplotlib.pyplot as plt
import random
import time
import dill as pickle
from IPython.display import clear_output
import dali.core as D
import dali
from dali.data import Lines, Process, DiscoverFiles, BatchBenefactor, IdentityReducer
from dali.data.batch import TranslationBatch
from dali.data.translation import TranslationFiles, TranslationMapper, build_vocabs, iterate_examples
from dali.utils.scoring import bleu, ErrorTracker
from dali.utils import (
Vocab,
Solver,
median_smoothing,
subsample,
Throttled,
pickle_globals,
unpickle_globals,
)
from dali import beam_search
# %matplotlib inline
# -
GPU_ID = 0
D.config.default_device = 'gpu'
D.config.default_gpu = GPU_ID
print(D.config.gpu_id_to_name(GPU_ID))
# # (Optional) continue running previous attempt
RELEVANT_VARIABLES = ["model", "vocabs", "solver", "data", "train_error", "validate_error"]
if False:
unpickle_globals("/home/sidor/tmp/translation_overfeat")
params = model.parameters()
# # Data loading
# +
# dataset
TRAIN_PATH = "/home/sidor/datasets/translation/train/fr_en/europarl/"
VALIDATE_PATH = "/home/sidor/datasets/translation/validation/fr_en/europarl/"
FROM_LANG = "en"
TO_LANG = "fr"
FROM_VOCAB_SIZE = 20000
TO_VOCAB_SIZE = 20000
# batching
MINIBATCH = 64
SENTENCE_LENGTH_BOUNDS = (None, 40)
SENTENCES_UNTIL_MINIBATCH = 1000 * MINIBATCH
# network sizes
INPUT_SIZE = 512
HIDDENS = [512,512,512,512]
SOFTMAX_INPUT = 512
# -
# you can press stop at any time if you think enough samples were collected.
vocabs = build_vocabs(TRAIN_PATH, FROM_LANG, TO_LANG, from_max_size=FROM_VOCAB_SIZE, to_max_size=TO_VOCAB_SIZE)
print (FROM_LANG + " vocabulary containts", len(vocabs[0]), "words")
print (TO_LANG + " vocabulary containts", len(vocabs[1]), "words")
p = Process(files=TranslationFiles(VALIDATE_PATH, FROM_LANG, TO_LANG),
mapper=TranslationMapper(sentence_bounds=SENTENCE_LENGTH_BOUNDS),
reducer=IdentityReducer())
validation_pairs = list(p)
# +
def create_dataset_iterator(dataset, sentences_until_minibatch=SENTENCES_UNTIL_MINIBATCH):
return iterate_examples(dataset, FROM_LANG, TO_LANG, vocabs,
minibatch_size=MINIBATCH,
sentence_length_bounds=SENTENCE_LENGTH_BOUNDS,
sentences_until_minibatch=sentences_until_minibatch)
validation_batches = list(create_dataset_iterator(VALIDATE_PATH, MINIBATCH))
# -
# # Model definition
class TranslationModel(object):
def __init__(self, input_size, hiddens,
encoder_vocab_size, decoder_vocab_size,
softmax_input_size=None, dtype=np.float32):
self.input_size = input_size
self.hiddens = hiddens
self.encoder_vocab_size = encoder_vocab_size
self.decoder_vocab_size = decoder_vocab_size
self.softmax_input_size = softmax_input_size
self.dtype = dtype
self.encoder_embedding = D.random.uniform(-0.05, 0.05, (encoder_vocab_size, input_size), dtype=dtype)
self.decoder_embedding = D.random.uniform(-0.05, 0.05, (decoder_vocab_size, input_size), dtype=dtype)
self.encoder_lstm = D.StackedLSTM(input_size, hiddens, memory_feeds_gates=True, dtype=dtype)
self.decoder_lstm = D.StackedLSTM(input_size, hiddens, memory_feeds_gates=True, dtype=dtype)
if self.softmax_input_size is not None:
self.predecoder = D.StackedInputLayer(self.hiddens, self.softmax_input_size)
self.decoder = D.Layer(self.softmax_input_size, decoder_vocab_size, dtype=dtype)
else:
self.decoder = D.Layer(hiddens[-1], decoder_vocab_size, dtype=dtype)
def decode_state(self, state):
if self.softmax_input_size is not None:
decoder_input = self.predecoder.activate([s.hidden for s in state])
else:
decoder_input = state[-1].hidden
return self.decoder.activate(decoder_input)
def error(self, batch):
error = D.Mat(1,1)
state = self.encoder_lstm.initial_states()
for ts in range(batch.timesteps):
inputs = batch.inputs(ts)
targets = batch.targets(ts)
if ts < batch.from_timesteps:
assert targets is None
encoded = self.encoder_embedding[inputs]
state = self.encoder_lstm.activate(encoded, state)
else:
assert inputs is None
decoded = self.decode_state(state)
# mask the error - only for the relevant sentences
tstep_error = batch.masks(ts).T() * D.MatOps.softmax_cross_entropy(decoded, targets)
#tstep_error = D.MatOps.softmax_cross_entropy(decoded, targets)
error = error + tstep_error.sum()
# feedback the predictions
if ts + 1 != batch.timesteps:
# for the last timestep encoding is not necessary
encoded = self.decoder_embedding[targets]
state = self.decoder_lstm.activate(encoded, state)
return error
def predict(self, input_sentence, **kwargs):
with D.NoBackprop():
state = self.encoder_lstm.initial_states()
for word_idx in input_sentence:
encoded = self.encoder_embedding[word_idx]
state = self.encoder_lstm.activate(encoded, state)
def candidate_scores(state):
decoded = self.decode_state(state)
return D.MatOps.softmax(decoded).log()
def make_choice(state, candidate_idx):
encoded = self.decoder_embedding[candidate_idx]
return self.decoder_lstm.activate(encoded, state)
return beam_search(state,
candidate_scores,
make_choice,
**kwargs)
def parameters(self):
ret = ([self.encoder_embedding,
self.decoder_embedding]
+ self.encoder_lstm.parameters()
+ self.decoder_lstm.parameters()
+ self.decoder.parameters())
if self.softmax_input_size is not None:
ret.extend(self.predecoder.parameters())
return ret
def calculate_bleu(model, validation_pairs, beam_width=5, debug=False):
references = []
hypotheses = []
for input_sentence, reference_translation in validation_pairs:
references.append(" ".join(reference_translation))
from_vocab, to_vocab = vocabs
encoded_input = from_vocab.encode(list(reversed(input_sentence)), add_eos=False)
predicted = model.predict(encoded_input,
eos_symbol=to_vocab.eos,
max_sequence_length=SENTENCE_LENGTH_BOUNDS[1] + 1,
blacklist=[], #to_vocab.unk],
beam_width=beam_width)[0].solution
decoded_prediction = " ".join(to_vocab.decode(predicted, strip_eos=True))
hypotheses.append(decoded_prediction)
if debug:
print ("INPUT: ", " ".join(input_sentence))
print ("REFERENCE: ", " ".join(reference_translation))
print ("PREDICTION: ", decoded_prediction)
print ("")
return bleu(references, hypotheses)
calculate_bleu(model, validation_pairs[-50:], 5, debug=True)
def show_reconstructions(model, example_pair):
from_words, to_words = example_pair
from_vocab, to_vocab = vocabs
from_with_unk = ' '.join(from_vocab.decode(from_vocab.encode(from_words)))
to_with_unk = ' '.join(to_vocab.decode(to_vocab.encode(to_words)))
print('TRANSLATING: %s' % from_with_unk)
print('REFERENCE: %s' % to_with_unk)
print('')
for solution, score, _ in model.predict(from_vocab.encode(list(reversed(from_words)), add_eos=False),
eos_symbol=to_vocab.eos,
max_sequence_length=SENTENCE_LENGTH_BOUNDS[1] + 1):
score = math.exp(score.w[0])
# reveal the unks
solution = ' '.join(to_vocab.decode(solution, False))
print(' %f => %s' % (score, to_vocab.decode(solution, True)))
# # Create new experiment
model = TranslationModel(INPUT_SIZE, HIDDENS, len(vocabs[0]), len(vocabs[1]), softmax_input_size=SOFTMAX_INPUT)
params = model.parameters()
solver = Solver("sgd", learning_rate=0.003)
data = []
train_error = ErrorTracker()
validate_error = ErrorTracker()
# # Training
# +
D.config.clear_gpu()
total_time = 0.0
num_words, num_batches = 0, 0
t = Throttled(10)
while True:
total_time = 0.0
num_words, num_batches = 0, 0
#if len(error_evolution) >= 5 and hasattr(solver, 'step_size'):
#solver.step_size = solver.step_size / 1.5
if solver.solver_type == 'adagrad':
solver.reset_caches(params)
for batch in data:
batch_start_time = time.time()
error = model.error(batch)
(error / batch.examples).grad()
D.Graph.backward()
# solver.set_lr_multiplier(model.encoder_embedding, batch.examples)
# solver.set_lr_multiplier(model.decoder_embedding, batch.examples)
solver.step(params)
batch_end_time = time.time()
train_error.append(error / batch.to_tokens)
total_time += batch_end_time - batch_start_time
num_words += batch.from_tokens + batch.to_tokens
num_batches += 1
if num_batches % 10 == 0:
val_batch = random.choice(validation_batches)
with D.NoBackprop():
validate_error.append(model.error(val_batch) / val_batch.to_tokens)
if t.should_i_run() and num_batches > 0 and abs(total_time) > 1e-6:
clear_output()
print('Epochs completed: ', train_error.num_epochs())
print('Error: ', train_error.recent(10))
print('Time per batch: ', total_time / num_batches)
print('Words per second: ', num_words / total_time )
print('Batches processed: ', num_batches)
if hasattr(solver, 'step_size'):
print('Solver step size: ', solver.step_size)
show_reconstructions(model, random.choice(validation_pairs))
sys.stdout.flush()
# free memory as soon as possible
del batch
train_error.finalize_epoch()
validate_error.finalize_epoch()
if train_error.num_epochs() > 0:
pickle_globals("/home/sidor/tmp/lst_hope/epoch_%d" % (train_error.num_epochs(),),
RELEVANT_VARIABLES)
data = create_dataset_iterator(TRAIN_PATH)
# -
pickle_globals("/home/sidor/tmp/translation_overfeat",RELEVANT_VARIABLES)
# +
train_error.__class__ = ErrorTracker
validate_error.__class__ = ErrorTracker
plt.figure(figsize=(20,15))
plt.plot(*train_error.raw(), label="train")
plt.plot(*validate_error.raw(), label="validate")
plt.legend()
# -
x, y = validate_error.raw()
print(train_error.error_evolution)
# +
# canoe trip lr=0.0001
# +
# canoe trip, lr = 0.001
# +
# 0.02 learning rate Jonathan's weights, divide by 1.5 every epoch (starting at fifth)
# +
# 0.3 learning rate Jonathan's weights, divide by 2 every epoch
# +
# 0.03 learning rate Jonathan weights
# +
# 0.03 learning rate Jonathan weights
# +
# 0.3 learning rate Jonathan weights
# +
# 0.3 learning rate, torch weight init
# +
r= epoch_error#median_smoothing(epoch_error, window=500)
plt.plot(range(len(r)), r)
# +
plottable = median_smoothing(train_error.epoch_error, 300)
#plottable = median_smoothing(train_error.epoch_error, 3)
plt.plot(range(len(plottable)), plottable)
# -
solver.__class__ =Solver
show_reconstructions(model, (["Where", "are","my","friends","?"],[]))
np.nan
type(np.arange(1, 10))
| notebooks/Machine Translation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # [ATM 623: Climate Modeling](../index.ipynb)
#
# [<NAME>](http://www.atmos.albany.edu/facstaff/brose/index.html), University at Albany
#
# # Lecture 2: Solving the zero-dimensional energy balance model
# + [markdown] slideshow={"slide_type": "skip"}
# ### About these notes:
#
# This document uses the interactive [`Jupyter notebook`](https://jupyter.org) format. The notes can be accessed in several different ways:
#
# - The interactive notebooks are hosted on `github` at https://github.com/brian-rose/ClimateModeling_courseware
# - The latest versions can be viewed as static web pages [rendered on nbviewer](http://nbviewer.ipython.org/github/brian-rose/ClimateModeling_courseware/blob/master/index.ipynb)
# - A complete snapshot of the notes as of May 2017 (end of spring semester) are [available on Brian's website](http://www.atmos.albany.edu/facstaff/brose/classes/ATM623_Spring2017/Notes/index.html).
#
# [Also here is a legacy version from 2015](http://www.atmos.albany.edu/facstaff/brose/classes/ATM623_Spring2015/Notes/index.html).
#
# Many of these notes make use of the `climlab` package, available at https://github.com/brian-rose/climlab
# -
# Ensure compatibility with Python 2 and 3
from __future__ import print_function, division
# + [markdown] slideshow={"slide_type": "slide"}
# ### From last class:
#
# - we wrote down a budget for the energy content of the global climate system
# - we wrote the OLR in terms of an effective emission temperature $T_e$
# - The equilibrium emission temperature for Earth is about $T_e \approx 255$ K
# - This depends only on energy output from the Sun, and the planetary albedo
# - We assumed that global energy content is proportional to surface temperature $T_s$
# - We thus have a single equation in two unknown temperatures:
#
# $$C \frac{d T_s}{dt} = (1-\alpha) Q - \sigma T_e^4$$
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Parameterizing the dependence of OLR on surface temperature
#
# Later, we will introduce additional physics for column radiative transfer to link $T_s$ and $T_e$.
# For now, we'll make the **simplest assumption** we can:
#
# $$ T_e = \beta T_s$$
#
# where $\beta$ is a dimensionless constant. This is a **parameterization** that we introduce into the model for simplicity. We need a value for our **parameter** $\beta$, which we will get from observations:
# + slideshow={"slide_type": "fragment"}
sigma = 5.67E-8 # Stefan-Boltzmann constant in W/m2/K4
Q = 341.3 # global mean insolation in W/m2
alpha = 101.9 / Q # observed planetary albedo
print(alpha)
Te = ((1-alpha)*Q/sigma)**0.25 # Emission temperature (definition)
print(Te)
Tsbar = 288. # global mean surface temperature in K
beta = Te / Tsbar # Calculate value of beta from observations
print(beta)
# + [markdown] slideshow={"slide_type": "slide"}
# Using this parameterization, we can now write a closed equation for surface temperature:
#
# $$C \frac{d T_s}{dt}=(1-α)Q-σ(\beta T_s )^4$$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Solving the energy balance model
#
# This is a first-order Ordinary Differential Equation (ODE) for $T_s$ as a function of time. It is also our very first climate model.
#
# To solve it (i.e. see how $T_s$ evolves from some specified initial condition) we have two choices:
#
# 1. Solve it analytically
# 2. Solve it numerically
# + [markdown] slideshow={"slide_type": "slide"}
# Option 1 (analytical) will usually not be possible because the equations will typically be too complex and non-linear. This is why computers are our best friends in the world of climate modeling.
#
# HOWEVER it is often useful and instructive to simplify a model down to something that is analytically solvable when possible. Why? Two reasons:
#
# 1. Analysis will often yield a deeper understanding of the behavior of the system
# 2. Gives us a benchmark against which to test the results of our numerical solutions.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Equilibrium solutions
#
# Note that equilibrium surface temperature has been tuned to observations:
#
# $$\bar{T_s} = \frac{1}{β} \bigg( \frac{(1-α)Q}{σ}\bigg)^{\frac{1}{4}} = 288 K $$
# + slideshow={"slide_type": "fragment"}
print( ((1-alpha)*Q/sigma)**0.25 /beta)
# + [markdown] slideshow={"slide_type": "fragment"}
# We are going to linearize the equation for small perturbations away from this equilibrium.
#
# Let $T_s = \bar{T_s} + T_s^\prime$ and restrict our solution to $T_s^\prime << \bar{T_s}$.
# Note this this is not a big restriction! For example, a 10 degree warming or cooling is just $\pm$3.4% of the absolute equilibrium temperature.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Linearizing the governing equation
#
# Now use a first-order Taylor series expansion to write
#
# $$OLR \approx \sigma \big(\beta T_s \big)^4 \approx \sigma \big(\beta \bar{T_s} \big)^4 + \Big(4 \sigma \beta^4 \bar{T_s}^3 \Big) T_s^\prime $$
#
# and the budget for the perturbation temperature thus becomes
#
# $$C \frac{d T_s^\prime}{d t} = -\lambda_0 T_s^\prime$$
#
# where we define
#
# $$\lambda_0 = 4 \sigma \beta^4 \bar{T_s}^3$$
# + [markdown] slideshow={"slide_type": "slide"}
# Putting in our observational values, we get
# + slideshow={"slide_type": "fragment"}
lambda_0 = 4 * sigma * beta**4 * Tsbar**3
# This is an example of formatted text output in Python
print( 'lambda_0 = {:.2f} W m-2 K-1'.format(lambda_0) )
# + [markdown] slideshow={"slide_type": "fragment"}
# This is actually our first estimate of what is often called the **Planck feedback**. It is the tendency for a warm surface to cool by increased longwave radiation to space.
#
# It may also be refered to as the "no-feedback" climate response parameter. As we will see, $\lambda_0$ quantifies the sensitivity of the climate system in the absence of any actual feedback processes.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Solve the linear ODE
#
# Now define
#
# $$ \tau = \frac{C}{\lambda_0} $$
#
# This is a positive constant with dimensions of time (seconds). With these definitions the temperature evolves according to
#
# $$ \frac{d T_s^\prime}{d t} = - \frac{T_s^\prime}{\tau}$$
#
# This is one of the simplest ODEs. Hopefully it looks familiar to most of you. It is the equation for an **exponential decay** process.
# + [markdown] slideshow={"slide_type": "slide"}
# We can easily solve for the temperature evolution by integrating from an initial condition $T_s^\prime(0)$:
#
# $$ \int_{T_s^\prime(0)}^{T_s^\prime(t)} \frac{d T_s^\prime}{T_s^\prime} = -\int_0^t \frac{dt}{\tau}$$
#
# $$\ln \bigg( \frac{T_s^\prime(t)}{T_s^\prime(0)} \bigg) = -\frac{t}{\tau}$$
#
# $$T_s^\prime(t) = T_s^\prime(0) \exp \bigg(-\frac{t}{\tau} \bigg)$$
#
# I hope that the mathematics is straightforward for everyone in this class. If not, go through it carefully and make sure you understand each step.
# + [markdown] slideshow={"slide_type": "slide"}
# ### e-folding time for relaxation of global mean temperature
#
# Our model says that surface temperature will relax toward its equilibrium value over a characteristic time scale $\tau$. This is an **e-folding time** – the time it takes for the perturbation to decay by a factor $1/e = 0.37$
#
# *What should this timescale be for the climate system?*
#
# To estimate $\tau$ we need a value for the effective heat capacity $C$.
# A quick and dirty estimate:
#
# $$C = c_w \rho_w H$$
#
# where
#
# $c_w = 4 \times 10^3$ J kg$^{-1}$ $^\circ$C$^{-1}$ is the specific heat of water,
#
# $\rho_w = 10^3$ kg m$^{-3}$ is the density of water, and
#
# $H$ is an effective depth of water that is heated or cooled.
# + [markdown] slideshow={"slide_type": "slide"}
# #### What is the right choice for water depth $H$?
#
# That turns out to be an interesting and subtle question. It depends very much on the timescale of the problem
#
# - days?
# - years?
# - decades?
# - millenia?
# + [markdown] slideshow={"slide_type": "slide"}
# We will revisit this question later in the course. For now, let’s just assume that $H = 100$ m (a bit deeper than the typical depth of the surface mixed layer in the oceans).
#
# Now calculate the e-folding time for the surface temperature:
# + slideshow={"slide_type": "fragment"}
c_w = 4E3 # Specific heat of water in J/kg/K
rho_w = 1E3 # Density of water in kg/m3
H = 100. # Depth of water in m
C = c_w * rho_w * H # Heat capacity of the model in J/m2/K
tau = C / lambda_0 # Calculated value of relaxation time constant
seconds_per_year = 60.*60.*24.*365.
print( 'The e-folding time is {:1.2e} seconds or about {:1.0f} years.'.format(tau, tau / seconds_per_year))
# + [markdown] slideshow={"slide_type": "fragment"}
# This is a rather fast timescale relative to other processes that can affect the planetary energy budget.
#
# **But notice that the climate feedback parameter $\lambda$ is smaller, the timescale gets longer.** We will come back to this later.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Some take-away messages:
#
# - Earth (or any planet) has a well-defined equilibrium temperature because of the temperature dependence of the outgoing longwave radiation.
# - The system will tend to relax toward its equilibrium temperature on an $e$-folding timescale that depends on (1) radiative feedback processes, and (2) effective heat capacity.
# - In our estimate, this e-folding time is relatively short. In the absence of other processes that can either increase the heat capacity or lower (in absolute value) the feedback parameter, the Earth would never be very far out of energy balance
# - We will quantify this statement more as the term progresses.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Plotting the solution in Python
#
# Here I'm going to show some example code for making simple line plots with Python. I strongly encourage you to try this out on your own.
#
# **Avoid the temptation to copy and paste the code!** You won't learn anything that way. Type the code into your own Python session. Experiment with it!
# + slideshow={"slide_type": "slide"}
# This code uses the `numpy` package to do efficient array operations.
# Before we use the package, we import it into the current Python session.
import numpy as np
t = np.linspace(0, 5*tau) # a time array
print(t)
# + slideshow={"slide_type": "fragment"}
type(t) # this shows that t is numpy.ndarray type
# + slideshow={"slide_type": "fragment"}
t.shape # a tuple showing the dimensions of the array
# + slideshow={"slide_type": "slide"}
Tsprime0 = 6. # initial temperature perturbation
# Here we define the actual solution
Tsprime = Tsbar + Tsprime0 * np.exp(-t/tau)
Tsprime.shape
# got the same size array
# the numpy function np.exp() operated simultaneously
# on all elements of the array
# + slideshow={"slide_type": "fragment"}
# And here is the solution array
print(Tsprime)
# + [markdown] slideshow={"slide_type": "slide"}
# To make a plot, we will use the `matplotlib` library. The plotting commands work a lot like `MATLAB`. But, like any other package, we need to import it before using it.
# + slideshow={"slide_type": "fragment"}
# pyplot is the name of the library of plotting routines within matplotlib
# here we import them and give them a "nickname"
import matplotlib.pyplot as plt
# + slideshow={"slide_type": "fragment"}
# this command allows the plots to appear inline in this notebook
# %matplotlib inline
# + slideshow={"slide_type": "fragment"}
plt.plot(t, Tsprime)
# + slideshow={"slide_type": "slide"}
# use a more convenient unit for time
plt.plot(t / seconds_per_year, Tsprime)
# + slideshow={"slide_type": "slide"}
# Or add some helpful labels
plt.plot(t / seconds_per_year, Tsprime)
plt.xlabel('Years')
plt.ylabel('Global mean temperature (K)')
plt.title('Relaxation to equilibrium temperature')
# + [markdown] slideshow={"slide_type": "slide"}
# ### Solving the ODE numerically
#
# In this case the equation is sufficiently simple that we have an analytical solution. Most models are too mathematically complex for this, and we need numerical solution methods. Because the governing equations for every model are differential in time (and usually in space as well), we need to use some numerical approximations to the governing equations.
# + [markdown] slideshow={"slide_type": "slide"}
# We approximate the time derivative with
#
# $$ \frac{dT}{dt} \approx \frac{∆T}{∆t} = \frac{T_1-T_0}{t_1-t_0} $$
#
# which makes sense as long as the **timestep** $\delta t$ is *sufficiently small*.
#
# What is meant by sufficiently small? In practice, small enough that the numerical solution behaves well! We will not spend much time in this course talking about numerical methods, but there is much we could say about this…
# + [markdown] slideshow={"slide_type": "slide"}
# ### Explicit time discretization
#
# The simplest time discretization is called **Forward Euler** or **Explicit Euler**. Say we know the state of the system at time $t_0$, i.e. we know the temperature $T_0$. Then rearranging the above,
#
# $$T_1 = T_0 + ∆t (dT/dt)$$
#
# So if we can calculate the **tendency** of the system (i.e. the time derivative) at time $t_0$, then we have a formula to predict the next state of the system.
# + [markdown] slideshow={"slide_type": "slide"}
# For our linearized zero-dimensional energy balance model,
#
# $$\frac{dT_s}{dt} = -\frac{1}{\tau} \big(T_s-\bar{T_s} \big)$$
#
# So we can predict the temperature with
#
# $$ T_1 = T_0 - \frac{\Delta t}{\tau} \big( T_0 - \bar{T_s} \big)$$
#
# Let’s implement this formula as a simple function in Python to calculate the next temperature at each timestep
# + slideshow={"slide_type": "fragment"}
def next_temperature(T0, timestep, tau):
Tsbar = 288.
return T0 - timestep/tau * (T0-Tsbar)
# + [markdown] slideshow={"slide_type": "slide"}
# Now let’s construct the full numerical solution by looping in time:
# + slideshow={"slide_type": "fragment"}
Tnumerical = np.zeros_like(t)
print( Tnumerical)
print( Tnumerical.size)
# + slideshow={"slide_type": "slide"}
# Assign the initial condition
Tnumerical[0] = Tsprime0 + Tsbar
print( Tnumerical)
# this shows indexing of the time array. t[0] is the first element
# t[1] is the second element
# in Python we always start counting from zero
# + slideshow={"slide_type": "fragment"}
timestep = t[1] - t[0]
for i in range(Tnumerical.size-1):
# assign the next temperature value to the approprate array element
Tnumerical[i+1] = next_temperature(Tnumerical[i], timestep, tau)
print( Tnumerical)
# + [markdown] slideshow={"slide_type": "slide"}
# Now we are going to plot this alongside the analytical solution.
# + slideshow={"slide_type": "fragment"}
plt.plot(t / seconds_per_year, Tsprime, label='analytical')
plt.plot(t / seconds_per_year, Tnumerical, label='numerical')
plt.xlabel('Years')
plt.ylabel('Global mean temperature (K)')
plt.title('Relaxation to equilibrium temperature')
plt.legend()
# the legend() function uses the labels assigned in the above plot() commands
# + [markdown] slideshow={"slide_type": "slide"}
# So this works quite well; the two solutions look nearly identical.
#
# Now that we have built some confidence in the numerical method, we can use it to study a slightly more complex system for which we don’t have the analytical solution.
#
# E.g. let’s solve the full non-linear energy balance model:
#
# $$C \frac{dT_s}{dt} = (1-\alpha) Q - \sigma \big(\beta T_s \big)^4 $$
#
# We’ll write a new solver function:
# + slideshow={"slide_type": "fragment"}
# absorbed solar is a constant in this model
ASR = (1-alpha)*Q
# but the longwave depends on temperature... define a function for this
def OLR(Ts):
return sigma * (beta*Ts)**4
# Now we put them together to get our simple solver function
def next_temperature_nonlinear(T0, timestep):
return T0 + timestep/C * (ASR-OLR(T0))
# + [markdown] slideshow={"slide_type": "slide"}
# Now solve this nonlinear model using the same procedure as above.
# + slideshow={"slide_type": "fragment"}
Tnonlinear = np.zeros_like(t)
Tnonlinear[0] = Tsprime0 + Tsbar
for i in range(Tnumerical.size-1):
Tnonlinear[i+1] = next_temperature_nonlinear(Tnonlinear[i], timestep)
# + [markdown] slideshow={"slide_type": "slide"}
# And plot the three different solutions together:
# + slideshow={"slide_type": "fragment"}
plt.plot(t / seconds_per_year, Tsprime, label='analytical')
plt.plot(t / seconds_per_year, Tnumerical, label='numerical')
plt.plot(t / seconds_per_year, Tnonlinear, label='nonlinear')
plt.xlabel('Years')
plt.ylabel('Global mean temperature (K)')
plt.title('Relaxation to equilibrium temperature')
plt.legend()
# + [markdown] slideshow={"slide_type": "slide"}
# And we see that the models essentially do the same thing.
#
# Now try some different initial conditions
#
# + slideshow={"slide_type": "fragment"}
T1 = 400. # very hot
for n in range(50):
T1 = next_temperature_nonlinear(T1, timestep)
print( T1)
# + slideshow={"slide_type": "fragment"}
T1 = 200. # very cold
for n in range(50):
T1 = next_temperature_nonlinear(T1, timestep)
print( T1)
# + [markdown] slideshow={"slide_type": "fragment"}
# The system relaxes back to 288 K regardless of its initial condition.
# + [markdown] slideshow={"slide_type": "skip"}
# <div class="alert alert-success">
# [Back to ATM 623 notebook home](../index.ipynb)
# </div>
# + [markdown] slideshow={"slide_type": "skip"}
# ____________
# ## Version information
# ____________
#
# + slideshow={"slide_type": "skip"}
# %load_ext version_information
# %version_information numpy, matplotlib
# + [markdown] slideshow={"slide_type": "slide"}
# ____________
#
# ## Credits
#
# The author of this notebook is [<NAME>](http://www.atmos.albany.edu/facstaff/brose/index.html), University at Albany.
#
# It was developed in support of [ATM 623: Climate Modeling](http://www.atmos.albany.edu/facstaff/brose/classes/ATM623_Spring2015/), a graduate-level course in the [Department of Atmospheric and Envionmental Sciences](http://www.albany.edu/atmos/index.php)
#
# Development of these notes and the [climlab software](https://github.com/brian-rose/climlab) is partially supported by the National Science Foundation under award AGS-1455071 to <NAME>. Any opinions, findings, conclusions or recommendations expressed here are mine and do not necessarily reflect the views of the National Science Foundation.
# ____________
# -
| Lectures/Lecture02 -- Solving the zero-dimensional EBM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python3 (alex_test)
# language: python
# name: alex_test
# ---
# %matplotlib inline
import logging
from datetime import datetime, timedelta
from pprint import pformat
import matplotlib.pyplot as plt
from geophys_point_fetcher import GeophysPointFetcher
logging.getLogger('geophys_point_fetcher').setLevel(logging.DEBUG)
# +
# Setup proxy as required
GA_STAFF_WIFI = False
if GA_STAFF_WIFI:
import os
os.environ['http_proxy'] = 'http://proxy.inno.lan:3128'
os.environ['https_proxy'] = 'http://proxy.inno.lan:3128'
# -
def JW_metadata_filter(metadata_dict):
'''
Example function to filter datasets based on metadata values in metadata_dict
This version applies John Wilford's filter conditions
Returns True for match, False otherwise
'''
try:
# Reject any datasets earlier than 1981
if datetime.strptime(metadata_dict['acquisition_start_date'], '%d/%m/%Y') < datetime.strptime('01/01/1981', '%d/%m/%Y'):
return False
# Only accept GA/BMR/AGSO datasets between 1981 and 1992
if (datetime.strptime(metadata_dict['acquisition_start_date'], '%d/%m/%Y') < datetime.strptime('01/01/1992', '%d/%m/%Y')
and metadata_dict['client'] not in ['Geoscience Australia',
'BMR',
'AGSO',
'GA'
]
):
return False
except ValueError:
logger.warning('WARNING: Unhandled date format: {}'.format(metadata_dict['acquisition_start_date']))
return False
return True
# Instantiate GeophysPointFetcher
gpf = GeophysPointFetcher()
# Set search parameters
bounding_box=(120.0, -29.0, 121, -28) # Bounding box coordinates in form "[<min_xord>,<min_yord>,<max_xord>,<max_yord>]"
keywords = ['geophysics', 'TMI'] # Keywords to match
variable_names = ['mag_awags'] # Variables to read
flight_lines_only=True
# +
# Plot data points
plt.figure(figsize=(30,30))
point_step = 200 # Step between points in plots: 1=plot every point, 2=plot every second point, etc.
# Find all nearest points using above parameters
for dataset_dict in gpf.point_data_generator(bounding_box=bounding_box,
keywords=keywords,
metadata_filter_function=JW_metadata_filter,
variable_names=variable_names,
flight_lines_only=flight_lines_only
):
#print(pformat(dataset_dict))
coordinates = dataset_dict['coordinates']
# Only show one in every point_step points
plt.plot(coordinates[:,0][0:-1:point_step], coordinates[:,1][0:-1:point_step], '.')
# -
| examples/fetch_points_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.7 64-bit (''venv'': venv)'
# language: python
# name: python_defaultSpec_1594411158751
# ---
# # 随机点名
import random
def random_choice_hacker(hackers):
hacker=random.choice(hackers)
return hacker
#加密
def encode_str(hackers=""):
hackers_a=[]
for hacker in hackers.split("\n"):
hacker_a=[]
for h in hacker:
a=ord(h)
hacker_a.append(str(a))
hacker_a=",".join(hacker_a)
hackers_a.append(hacker_a)
hackers_a="\n".join(hackers_a)
return hackers_a
#解密
def dencode_str(hackers_a=""):
hackers=[]
for hacker in hackers_a.split("\n"):
hacker_c=[]
for h in hacker.split(","):
h=chr(int(h))
hacker_c.append(h)
hacker_c="".join(hacker_c)
hackers.append(hacker_c)
return hackers
# + tags=[]
hackers='''38472,21338
26446,24605,21473
26446,21338,38597
21608,24935,33521
35768,36335
21494,26062
26446,26149,31179
37057,26446,33418
26045,36828,23665
28504,36194
34081,38125,20043
38470,28147,39118
21016,29030,38451
26446,33620,26525
24352,26032,28982
26446,38160
32599,21531,20975
19969,19968
29579,22696
29579,23431,26480
26446,25104,26753
21331,20140,28207
29579,23425
26446,19982,20961
39640,19968,26041
38472,28751,40857'''
hacker=random_choice_hacker(dencode_str(hackers))
print(hacker)
# -
# # 入门练习
# ## 重点掌握以下概念及代码练习
# +
#https://www.w3cschool.cn/python3/
# 变量
'''Python中的变量不需要声明。每个变量在使用前都必须赋值,变量赋值以后该变量才会被创建。'''
# 条件控制
'''Python中if语句的一般形式'''
# 循环
'''Python中的循环语句 for 和 while'''
# 函数
'''Python 定义函数使用 def 关键字'''
def color():
print("----")
# 类
'''Python从设计之初就已经是一门面向对象的语言,正因为如此,在Python中创建一个类和对象是很容易的。'''
# -
# # 简单练习
#
# + tags=[]
def color(input_image):
print("配色...",input_image)
main_color=[122,222,88]
return main_color
def layout(image,color):
print("布局...",image)
poster=image+"_"+",".join([str(c) for c in color])
return poster
def imageProcessing(input_image):
print ("图像处理...",input_image)
return input_image
image='test.png'
main_color=color(image)
image=imageProcessing(image)
layout(image,main_color)
# -
# # Design Patterns
#
# Refer to https://refactoring.guru/design-patterns/python
#
# ## 理解单例模式
# 保证一个类仅有一个实例,并提供一个访问它的全局访问点
# + tags=[]
# 类的专有方法:
# __init__ : 构造函数,在生成对象时调用
# __call__: 函数调用
class Singleton(type):
# __init__ : 构造函数,在生成对象时调用
def __init__(self,class_name,class_bases,class_dic):
super(Singleton,self).__init__(class_name,class_bases,class_dic)
self.__instance=None
# __call__: 函数调用
def __call__(self, *args, **kwargs):
if not self.__instance:
obj=object.__new__(self)
self.__init__(obj)
self.__instance=obj
return self.__instance
class MyAIDesign(object,metaclass=Singleton):
def __init__(self):
self.name='darksee.ai'
self.version="0.1"
print("--one --")
def match_color(self,input_color=[0,0,0]):
return [255,255,255]
ai1=MyAIDesign()
ai2=MyAIDesign()
print(ai1 is ai2)
# -
# ## 理解观察者模式
#
# 观察者模式指的是一个对象(Subject)维持一系列依赖于它的对象(Observer),当有关状态发生变更时 Subject 对象则通知一系列 Observer 对象进行更新。
#
# 在观察者模式中,Subject 对象拥有添加、删除和通知一系列 Observer 的方法等等,而 Observer 对象拥有更新方法等等。
# + tags=[]
# 泛化出“观察者”类
class Observer:
def update(self):
pass
class Color(Observer):
def update(self, action):
print ("更新配色规则: %s" % action)
self.run()
def run(self):
print("配色...")
class Layout(Observer):
def update(self, action):
print ("更新布局规则: %s" % action)
self.run()
def run(self):
print("布局...")
class ImageProcessing(Observer):
def update(self, action):
print ("更新图像处理规则: %s" % action)
self.run()
def run(self):
print ("图像处理...")
# Subject 对象拥有添加、删除和通知一系列 Observer 的方法等等
class Subject:
observers=[]
action=""
def add_observer(self,observer):
self.observers.append(observer)
def notify_all(self):
for i,obs in enumerate(self.observers):
print("\nSTEP",i+1)
obs.update(self.action)
class MyAIDesign(Subject):
def set_action(self,action="默认"):
self.action=action
def start_design(self):
return True
#配色
color=Color()
#排版
layout=Layout()
#图像处理,图像裁切、图像的色彩调整……
image_processing=ImageProcessing()
darksee_ai=MyAIDesign()
darksee_ai.add_observer(image_processing)
darksee_ai.add_observer(color)
darksee_ai.add_observer(layout)
darksee_ai.notify_all()
# +
class Singleton(type):
def __init__(self,class_name,class_bases,class_dic):
super(Singleton,self).__init__(class_name,class_bases,class_dic)
self.__instance=None
def __call__(self, *args, **kwargs):
if not self.__instance:
obj=object.__new__(self)
self.__init__(obj)
self.__instance=obj
return self.__instance
# 泛化出“观察者”类
class Observer:
def update(self):
pass
class Color(Observer):
def update(self, action):
print ("触发配色: %s" % action)
self.run()
def run(self):
print("配色...")
class Layout(Observer):
def update(self, action):
print ("触发布局: %s" % action)
self.run()
def run(self):
print("布局...")
class ImageProcessing(Observer):
def update(self, action):
print ("触发图像处理: %s" % action)
self.run()
def run(self):
print ("图像处理...")
# Subject
class Subject:
observers=[]
action=""
def add_observer(self,observer):
self.observers.append(observer)
def notify_all(self):
for i,obs in enumerate(self.observers):
print("\nSTEP",i+1)
obs.update(self.action)
class MyAIDesign(Subject,object,metaclass=Singleton):
def __init__(self):
self.name='darksee.ai'
self.version="0.1"
print("初始化:",self.name,self.version)
def set_action(self,action):
self.action=action
def start_design(self):
return True
def match_color(self,input_color=[0,0,0]):
return [255,255,255]
# + tags=[]
color=Color()
layout=Layout()
image_processing=ImageProcessing()
darksee_ai=MyAIDesign()
darksee_ai.add_observer(image_processing)
darksee_ai.add_observer(color)
darksee_ai.add_observer(layout)
if darksee_ai.start_design():
darksee_ai.set_action("主色不变")
darksee_ai.notify_all()
# + tags=[]
darksee_ai.set_action("人物主题")
darksee_ai.notify_all()
# -
# For basic OOP in Python, refer to https://realpython.com/python3-object-oriented-programming/
# # 课堂演示作业: 自动配色 算法
# + tags=[]
# 根据设定的颜色,改变一张图片的配色
# - 输入
# 一张图片,用户设定主颜色
#step 01 处理图片--》 主色调提取, h :0-360
def extract_main_color_from_image(image="图片"):
print("step 01---提取"+image+"的主色调- 运行中-")
h=100
return h
# hsv
#step 02 用户设定的主颜色 hsv - 图片的主色调 = 差距
def color_xxx(user_color=[10,2,33],main_color=0):
print("用户输入的是",user_color)
print("step 02---计算之间调整的幅度- 运行中-")
k=33
return k
main_color=extract_main_color_from_image("图片1234")
m=color_xxx([20,2,33],main_color)
print('看到 调整的幅度为:',m)
| notebook/Level01-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <p float="center">
# <img src="images/horizontal.png" alt="Coiled logo" width="415" hspace="10"/>
# </p>
#
#
# <p float="center">
# <img src="images/dask_horizontal_no_pad.svg" alt="Dask logo" width="400" hspace="10" />
# </p>
#
# # Scalable Data Science
#
# In this notebook, we'll
#
# * Perform a basic analytics workflow on the NYC taxi dataset using Pandas;
# * Scale up this workflow to a dataset that doesn't fit in RAM using Dask;
# * (Optional) Scale out this workflow to leverage a cluster on the Cloud using Coiled.
#
# The workflow is intentionally boring so that we can see the power of scalable data science immediately: we'll load some data and perform some basic analytics.
#
# In the notebooks that follow, we'll jump into more interesting examples, including machine learning.
#
# *A bit about me:* I'm <NAME>, Head of Data Science Evangelism and Marketing at [Coiled](coiled.io/). We build products that bring the power of scalable data science and machine learning to you, such as single-click hosted clusters on the cloud. We want to take the DevOps out of data science so you can get back to your real job. If you're interested in taking Coiled for a test drive, you can sign up for our [free Beta here](beta.coiled.io/).
#
# Before scaling up, let's look at a common workflow in Pandas.
# ## 1. Pandas: Convert CSV to Parquet and Engineer a Feature
# <img src="images/pandas-logo.svg" alt="pandas logo" style="width: 500px;"/>
#
# In the following, we'll
#
# * use Pandas to load in part of the NYC taxi dataset from a CSV and
# * compute the average tip as a function of the number of passengers.
#
# If you're following along in Binder, you won't be able to execute the code but you can read it.
# ### Download the data from Amazon
# !wget https://s3.amazonaws.com/nyc-tlc/trip+data/yellow_tripdata_2019-{01..12}.csv
# Note: this will take at least several minutes to download the above.
# Check out head of 1st file
# !head data_taxi/yellow_tripdata_2019-01.csv
# ### Investigate data locally with Pandas
#
# %%time
# Import pandas and read in beginning of 1st file
import pandas as pd
df = pd.read_csv(
"data_taxi/yellow_tripdata_2019-01.csv",
#nrows=10000,
)
df
# ## Basic Analytics
# +
# %%time
# Compute average tip as a function of the number of passengers
df.groupby("passenger_count").tip_amount.mean()
# -
# **Recap:** We have
#
# * used Pandas to load in part of the NYC taxi dataset from a CSV and
# * computed the average tip as a function of the number of passengers.
# ### Operate on many files in a for loop?
#
# We could do this, but it's unpleasant
# ```python
# for filename in glob("~/data/nyctaxi/yellow_tripdata_2019-*.parq"):
# df = pd.read_csv(filename)
# ...
# df.to_parquet(...)
# ```
# ## 2. Use Dask locally to process the full dataset
# <img src="images/dask_horizontal_no_pad.svg" alt="Dask logo" style="width: 500px;"/>
#
# The full NYC taxi dataset won't even fit in the RAM of my laptop. Do I need a large or external cluster yet? No. First, I can take advantage of all the cores on my laptop in parallel. This is what we call *scaling up* our computation (out-of-core computing). Later we'll see how to *scale out* computation across a cluster.
#
# One way of doing this is with [Dask](dask.org/). As we're about to see, part of the value of Dask lies in its API being as close as possible to the PyData APIs we know and love, in this case, Pandas.
#
# In [the words of <NAME>](https://coiled.io/blog/history-dask/), core developer and co-maintainer of Dask and CEO of Coiled, there was a social goal of Dask:
# > Invent nothing. We wanted to be as familiar as possible to what users already knew in the PyData stack
#
# Let's do it!
#
# The plan:
#
# * use Dask to load in **all** of the NYC taxi dataset from 10+ CSVs (8+ GBs) and
# * compute the average tip as a function of the number of passengers.
# We'll also dive into the basics of Dask and distributed compute (but we'll execute some code first and dive into this part while it runs!).
# Import Dask parts, spin up a local cluster, and instantiate a Client
from dask.distributed import LocalCluster, Client
cluster = LocalCluster(n_workers=4)
client = Client(cluster)
client
# +
# %%time
import dask.dataframe as dd
# Import the full dataset (note the Dask API!)
df = dd.read_csv(
"data_taxi/yellow_tripdata_2019-*.csv",
parse_dates=["tpep_pickup_datetime", "tpep_dropoff_datetime"],
dtype={'RatecodeID': 'float64',
'VendorID': 'float64',
'passenger_count': 'float64',
'payment_type': 'float64'}
)
df
# +
# %%time
# Prepare to compute the average tip
# as a function of the number of passengers
mean_amount = df.groupby("passenger_count").tip_amount.mean()
# +
# %%time
# compute the average tip as a function of the number of passengers
mean_amount.compute()
# -
# ### 2a Notes on what is happening in Dask and Python
# The above code will take some time to run so let's take this opportunity to see what is going on with Dask, Python, and the distributed computation.
# #### Components of Dask
# Dask contains 3 main components and we have already see two of them above:
# * High-level collections in the form of Dask DataFrames (these set up the steps of your computation);
# * Schedulers (these actually execute the computation, in this case, on a single machine).
#
# Let's get a sense for what these are.
# <img src="images/dask-components.svg" width="400px">
# #### Dask DataFrames
# What exactly is this Dask DataFrame? A schematic is worth a thousand words:
# <img src="images/dask-dataframe.svg" width="400px">
# Essentially, the Dask DataFrame is a large, virtual dataframe divided along the index into multiple Pandas DataFrames.
# #### Dask Schedulers, Workers, and beyond
# <img src="images/dask-cluster.svg" width="400px">
# Work (Python code) is performed on a cluster, which consists of
# * a _scheduler_ (which manages and sends the work / tasks to the workers)
# * _workers_, which compute the tasks.
#
# The _client_ is "the user-facing entry point for cluster users." What this means is that the client lives wherever you are writing your Python code and the client talks to the scheduler, passing it the tasks.
# **Recap:** We have
#
# * used Dask to load in **all** of the NYC taxi dataset from 10+ CSVs (8+ GBs),
# * computed the average tip as a function of the number of passengers, and
# * dived into the basic of Dask and distributed compute and understand the basic concepts.
# ## 3. Optional: Work directly from the cloud with Coiled
#
# <br>
# <img src="images/horizontal.png" alt="Coiled logo" style="width: 500px;"/>
# <br>
#
# Here I'll spin up a cluster on Coiled to show you just how easy it can be. Note that to do so, I've also signed into the [Coiled Beta](beta.coiled.io/), pip installed `coiled`, and authenticated. You can do the same!
#
# The plan:
#
# * use Coiled to load in **all** of the NYC taxi dataset from 10+ CSVs (8+ GBs) on an AWS cluster,
# * massage the data,
# * engineer a feature,
# * compute the average tip as a function of the number of passengers, and
# * save to [Parquet](https://en.wikipedia.org/wiki/Apache_Parquet) (far more efficient than CSV, but not human-readable).
import coiled
from dask.distributed import LocalCluster, Client
# Create a Software Environment
coiled.create_software_environment(
name="my-software-env",
conda="binder/environment.yml",
)
# Control the resources of your cluster by creating a new cluster configuration
coiled.create_cluster_configuration(
name="my-cluster-config",
worker_memory="16 GiB",
worker_cpu=4,
scheduler_memory="8 GiB",
scheduler_cpu=2,
software="my-software-env",
)
# Spin up cluster, instantiate a Client
cluster = coiled.Cluster(n_workers=10, configuration="my-cluster-config")
client = Client(cluster)
client
# +
import dask.dataframe as dd
# Read data into a Dask DataFrame
df = dd.read_csv(
"s3://nyc-tlc/trip data/yellow_tripdata_2019-*.csv",
parse_dates=["tpep_pickup_datetime", "tpep_dropoff_datetime"],
dtype={
'RatecodeID': 'float64',
'VendorID': 'float64',
'passenger_count': 'float64',
'payment_type': 'float64'
},
storage_options={"anon":True}
)
df
# +
# %%time
# Prepare to compute the average tip
# as a function of the number of passengers
mean_amount = df.groupby("passenger_count").tip_amount.mean()
# +
# %%time
# Compute the average tip
# as a function of the number of passengers
mean_amount.compute()
# -
client.shutdown()
# **Recap:** We have
# * used Coiled to load in **all** of the NYC taxi dataset from 10+ CSVs (10 GBs) on an AWS cluster,
# * computed the average tip as a function of the number of passengers, and
# * learned a bunch about using Dask on cloud-based clusters!
| 01-data-analysis-at-scale.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''venv38'': venv)'
# name: python3
# ---
# # Что за хитрые валидации?
# > 🚀 В этой практике нам понадобятся: `numpy==1.21.2, pandas==1.3.3, scikit-learn==0.24.2`
#
# > 🚀 Установить вы их можете с помощью команды: `!pip install numpy==1.21.2, pandas==1.3.3, scikit-learn==0.24.2`
#
# ## Содержание
#
# * [Выборка для валидации (Валидационная выборка - Validation set)](#Выборка-для-валидации-Валидационная-выборка---Validation-set)
# * [Кросс-валидация (Cross-Validation)](#Кросс-валидация-Cross-Validation)
# * [Поиск гиперпараметров](#Поиск-гиперпараметров)
# * [Задание](#Задание)
# * [Вопросы для закрепления](#Вопросы-для-закрепления)
# * [Полезные ссылки](#Полезные-ссылки)
#
# Привет! Сегодня мы узнаем очень важный аспект разработки модели машинного обучения в реальных условиях!
#
# Если раньше мы почти не занимались настройкой моделей (так как параметров почти не было) и нам было достаточно разработать модель на выборке для обучения и дальше проверить работу на "новых данных" (тестовая выборка).
#
# Но на последнем занятии мы столкнулись с кучей разных моделей и, оказывается, в разных моделях бывает много разных параметров! Мы уже задумывались о том, что использовать тестовую выборку для настройки параметров нельзя, потому что так мы проводим "неявное" обучение и настройку гиперпараметров, а значит у нас после этого отсутствуют действительно новые данные, на которых можно снять окончательные метрики.
#
# Что же нам тогда делать? Выход есть и начнём мы с наиболее простого примера!
# + _cell_id="E7NJTs5PfBuCGUgV"
# Настройки для визуализации
# Если используется темная тема - лучше текст сделать белым
import numpy as np
import pandas as pd
import random
import os
pd.set_option('display.max_columns', 50)
# Зафиксируем состояние случайных чисел
RANDOM_SEED = 42
np.random.seed(RANDOM_SEED)
random.seed(RANDOM_SEED)
# -
# ## Выборка для валидации (Валидационная выборка - Validation set)
#
# Валидационной выборкой называется такая выборка, которая используется для настройки *гиперпараметров* модели.
#
# > 🤓 В свою очередь выборка для обучения используется для настройки *параметров* модели.
#
# > ✨ Если вы успели позабыть, что означают эти два термина - обсудите с преподавателем
#
# А как нам эту выборку получить?? Тут тоже нет ничего сложного - просто, после деления на train-test от обучающих данных выделяется ещё одна небольшая выборка (valid). Обычно, если данных очень много, то данные делятся в пропорции 60-20-20 (train-test-valid). Если данных не сильно много (зависит от ситуации и задачи, но в бинарной классификации можно за порог считать порядка 10000 примеров), то лучше больше выделить на тест, чтобы проверить работоспособность модели на совершенно новых невиданных ранее данных - 50-20-30.
# Так вот, давайте сделаем свою функцию, которая производит деление на train-test-valid с указанием процентов соотношений:
#
# > ⚠️ В этом примере мы отключим стратификацию по двум причинам: далее мы будем работать с большими данными без дисбаланса - случайное разделение достаточно, чтобы данные попали во все выборки, и проблема переполнения на Windows с типом по-умолчанию https://github.com/numpy/numpy/issues/9464
#
# > ⚠️ В целом, далее рекомендуется использовать стратификацию, но проверять результат работы!
# + _cell_id="1Dhs84eXmAiwyIhA"
# TODO - создайте функцию train_test_valid_split(), которая принимает
# df - DataFrame
# test_size - пропорция выборки для тестирования
# valid_size - пропорция выборки для валидации
# Стратификацию не используем - причина выше
# Функция должна возвращать три DataFrame в порядке - train, test, valid
# + _cell_id="fux1M1S97McFVHCh"
# TEST
rng = np.random.default_rng(RANDOM_SEED)
_test_size = 1000
_test_df = pd.DataFrame({'col': rng.integers(0, 100, size=_test_size), 'target': rng.choice([0, 1], size=_test_size)})
_test_result_train, _test_result_test, _test_result_valid = train_test_valid_split(_test_df, test_size=0.1, valid_size=0.3)
assert _test_result_train.shape[0] == 600
assert _test_result_test.shape[0] == 100
assert _test_result_valid.shape[0] == 300
_test_expected_vc_train = pd.Series([303, 297], index=[0, 1], name='target')
pd.testing.assert_series_equal(_test_result_train['target'].value_counts(), _test_expected_vc_train)
_test_expected_vc_test = pd.Series([52, 48], index=[1, 0], name='target')
pd.testing.assert_series_equal(_test_result_test['target'].value_counts(), _test_expected_vc_test)
_test_expected_vc_valid = pd.Series([156, 144], index=[0, 1], name='target')
pd.testing.assert_series_equal(_test_result_valid['target'].value_counts(), _test_expected_vc_valid)
print("Well done!")
# -
# Отлично! Молодцы!
#
# Для испытаний новой функции и освоения подходов подбора гиперпараметров воспользуемся датасетом [Covertype Data Set](https://www.kaggle.com/teejmahal20/airline-passenger-satisfaction).
#
# > Выгружайте файлы `train.csv` и `test.csv` и кладите в папку `airline_dataset` рядом с ноутбуком.
# + _cell_id="HuKcBbD6CjWOKdTu"
TRAIN_PATH = os.path.join('airline_dataset', 'train.csv')
TEST_PATH = os.path.join('airline_dataset', 'test.csv')
df_train = pd.read_csv(TRAIN_PATH, sep=',', index_col=0)
df_test = pd.read_csv(TEST_PATH, sep=',', index_col=0)
df = df_train.append(df_test)
df.reset_index(inplace=True, drop=True)
# Определим полезные константы и переменные
TARGET_COLUMN = 'satisfaction'
x_columns = df_train.columns
x_columns = x_columns[x_columns != TARGET_COLUMN]
# -
# Сейчас мы слили файлы воедино, так как наша функция умеет сама разделять с нужной пропорцией на выборки.
#
# > При участии в соревнованиях нельзя использовать тестовую выборку для обучения. Нужно валидацию брать от выборки обучения.
#
# Давайте бегло посмотрим на данные
# + _cell_id="7fKi00vTkkkFxClS"
df.shape
# + _cell_id="OSVFPqZ1SbsEyeA0"
df.head(10)
# + _cell_id="LcKsF3DIqQDjTJYD"
df.isna().sum()
# + _cell_id="MlCfAg8hsU8Xfe97"
df.nunique(dropna=False)
# -
# Как видите, в датасете мы имеем колонку `id`, которая имеет только уникальные числовые значения и колонку `Arrival Delay in Minutes`, которая имеет пропуски.
#
# Так как колонка числовая, то заполнение выполним с помощью SimpleImputer, а колонку `id` просто удалим из датасета.
#
# Также, нам нужно закодировать колонки `Gender`, `Customer Type`, `Type of Travel`, `Class`, так как они имеют строковые значения. Помимо этого нам надо ещё и целевую закондировать!
#
# Уф, столько дел!
#
# Давайте напишем свой класс для предобработки данных!
# + _cell_id="3PnUhkgOOYYluXM1"
df_train, df_test, df_valid = train_test_valid_split(df, test_size=0.2, valid_size=0.2)
# Вот эта проверка очень важна! Вы можете сами убедиться и попробовать разделить датасет стандартной функцией train_test_split() со стратификацией по целевой колонке
# В результате подобная проверка не пройдет
assert len(df) == len(df_train) + len(df_test) + len(df_valid)
df_train.shape, df_test.shape, df_valid.shape
# + _cell_id="29gImr0WpXInP9Af"
from sklearn.preprocessing import OneHotEncoder, LabelEncoder
from sklearn.impute import SimpleImputer
class DataPreprocess:
def __init__(self):
self._imputer = SimpleImputer(strategy='median')
self._impute_cols = ['Arrival Delay in Minutes']
self._encoder = OneHotEncoder(sparse=False)
self._encode_cols = ['Gender', 'Customer Type', 'Type of Travel', 'Class']
self._label_enc = LabelEncoder()
def fit(self, X, y):
self._imputer.fit(X[self._impute_cols])
self._encoder.fit(X[self._encode_cols])
self._label_enc.fit(y)
def transform(self, X, y):
X = X.copy()
X.drop(columns=['id'], inplace=True)
X[self._impute_cols] = self._imputer.transform(X[self._impute_cols])
encoded_data = self._encoder.transform(X[self._encode_cols])
new_col_names = self._encoder.get_feature_names(self._encode_cols)
X_enc = pd.DataFrame(data=encoded_data, columns=new_col_names, index=X.index)
X.drop(columns=self._encode_cols, inplace=True)
X = pd.concat([X, X_enc], axis=1)
y = self._label_enc.transform(y)
return X, y
# -
# Теперь давайте применим и обучим модель случайного леса на наших данных и посмотрим метрики!
# + _cell_id="BG8Al2Kx2Ux9a7Od"
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import classification_report
rf_clf = RandomForestClassifier(max_depth=1, n_estimators=100, random_state=RANDOM_SEED)
data_prep = DataPreprocess()
y_train = df_train[TARGET_COLUMN]
x_train = df_train[x_columns]
y_test = df_test[TARGET_COLUMN]
x_test = df_test[x_columns]
data_prep.fit(x_train, y_train)
x_train_enc, y_train_enc = data_prep.transform(x_train, y_train)
rf_clf.fit(x_train_enc, y_train_enc)
# + _cell_id="ZFUbK8XTF6tgxNIx"
x_test_enc, y_test_enc = data_prep.transform(x_test, y_test)
y_pred = rf_clf.predict(x_test_enc)
report = classification_report(y_test_enc, y_pred)
print(report)
# -
# Отлично, вот мы обучили модель с какими-то параметрами, но что если мы хотим попробовать другие параметры? Нам нужно в таком случае использовать выборку для валидации. Давайте попробуем перебрать пару параметров в цикле:
# + _cell_id="slQer6XYF0xCm93L"
from sklearn.metrics import f1_score
y_train = df_train[TARGET_COLUMN]
x_train = df_train[x_columns]
y_valid = df_valid[TARGET_COLUMN]
x_valid = df_valid[x_columns]
data_prep = DataPreprocess()
data_prep.fit(x_train, y_train)
x_train_enc, y_train_enc = data_prep.transform(x_train, y_train)
x_valid_enc, y_valid_enc = data_prep.transform(x_valid, y_valid)
for max_depth_value in np.arange(3, 6):
for n_estimators_value in np.arange(70, 110, step=10):
print(f'Try max_depth: {max_depth_value} | n_estimators: {n_estimators_value}')
rf_clf = RandomForestClassifier(max_depth=max_depth_value, n_estimators=n_estimators_value, random_state=RANDOM_SEED)
rf_clf.fit(x_train_enc, y_train_enc)
y_val_pred = rf_clf.predict(x_valid_enc)
f1_value = f1_score(y_valid_enc, y_val_pred, average='macro')
print(f'F1 score: {f1_value}')
# -
# Смотрите, мы пробежались по паре параметров и выявили наилучший вариант из возможных заданных! Отлично, таким образом, мы можем взять лучшую пару параметров и улучшить показания на тестовой выборке:
# + _cell_id="4AD4muMrhxUYOHZU"
rf_clf = RandomForestClassifier(max_depth=5, n_estimators=90, random_state=RANDOM_SEED)
rf_clf.fit(x_train_enc, y_train_enc)
y_pred = rf_clf.predict(x_test_enc)
report = classification_report(y_test_enc, y_pred)
print(report)
# -
# Замечательно! Как видите, мы действительно улучшили результат и не наблюдается никакого переобучения из-за **подгона гиперпараметров** под данные. Вот так удобно можно использовать валидационную выборку!
# ## Кросс-валидация (Cross-Validation)
#
# Всё было бы хорошо, но есть одна важная проблема - данных не всегда бывает много!
#
# Иногда, нам приходится обучать модели на данных размеров менее 10000 примеров, что уже накладывает ограничение на разделение. Откусить 30%, чтобы тест был более-менее репрезентативным, по-хорошему на валидацию надо процентов 20, что тогда остаётся - всего пол датасета на обучение? Ну, дела!
#
# Для таких случаев, умные люди придумали [кросс-валидацию](https://scikit-learn.org/stable/modules/cross_validation.html). Это процесс, при котором не требуется отдельно брать валидационную выборку. Мы просто используем часть набора для обучения для валидации.
#
# Но стоп, разве мы не делали именно то же самое? От выборки для обучения откусывали кусочек и фиксировали для оценки того, какие показатели при нынешних гиперпараметрах??
#
# Не всё так просто! Мы не просто откусили и положили, а делаем это по-очереди несколько раз, а именно K раз. Такой процесс зовется **K-fold кросс-валидацией**. Гляньте картинку:
#
# <p align="center"><img src="https://raw.githubusercontent.com/AleksDevEdu/ml_edu/master/assets/32_grid_search.png" width=800/></з>
# По сути, процесс кросс-валидации заключается в следующем:
# - Делаем из выборки для обучения K фолдов, например, 5
# - На первой итерации берём первый фолд как валидационный, а на остальных учим модель (4 фолда)
# - Оцениваем параметры по первому фолду, записываем
# - На второй итерации - второй фолд валидационный, а остальные 4 - обучения
# - Также, оценили, записали
# - После всех K итераций усредняем оценки и получаем результирующую метрику на валидации!
#
# Вуаля!
#
# Что мы получаем в итоге? В K раз больше операций, но зато в ходе кросс-валидации мы всецело провалидировали модель на всем наборе данных для обучения без необходимости отдельно выделять ЕЩЁ данных из и так малого количества!
#
# Давайте посмотрим, как это работает!
# + _cell_id="M9pJOncDcr5BAd12"
# Соединим выборки обучения и валидации так как при кросс-валидации на отдельная выборка не нужна
# Тестовая остается та же, чтобы было корреткное сравнение
df_train_valid = df_train.append(df_valid)
df_train_valid.shape
# + _cell_id="QBjEIWFu4HgOr3Vw"
from sklearn.model_selection import cross_val_score
y_train = df_train_valid[TARGET_COLUMN]
x_train = df_train_valid[x_columns]
data_prep = DataPreprocess()
data_prep.fit(x_train, y_train)
x_train_enc, y_train_enc = data_prep.transform(x_train, y_train)
for max_depth_value in np.arange(3, 6):
for n_estimators_value in np.arange(70, 110, step=10):
print(f'Try max_depth: {max_depth_value} | n_estimators: {n_estimators_value}')
rf_clf = RandomForestClassifier(max_depth=max_depth_value, n_estimators=n_estimators_value, random_state=RANDOM_SEED)
f1_values = cross_val_score(estimator=rf_clf, X=x_train_enc, y=y_train_enc, cv=5, scoring='f1_macro')
print(f'F1 score: {f1_values}, mean: {np.mean(f1_values)}')
# -
# Вот так просто можно получить рабочий вариант валидации на малом датасете! Вообще не сложно, правда?
# ## Поиск гиперпараметров
#
# А теперь представьте, что в модели не два, а куууча параметров. Вообще, если обратиться к странице [RandomForestClassifier](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html), то вы увидите, что параметров там больше, чем два! И что теперь, писать кучу вложенных циклов? Вааау, получится примерно так:
#
# <p align="center"><img src="https://raw.githubusercontent.com/AleksDevEdu/ml_edu/master/assets/indent_meme.jpg" width=800/></p>
# А чего тогда делать?? Есть для вас интересный вариант! Посмотрим на [GridSearchCV](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html).
#
# По сути, это класс, которому мы отдаём нашу модель, список параметров, которые надо перебрать и после этого можно спокойно ожидать результата. Работает это так:
# + _cell_id="oETlLOgzEGL9BBPU"
from sklearn.model_selection import GridSearchCV
y_train = df_train_valid[TARGET_COLUMN]
x_train = df_train_valid[x_columns]
data_prep = DataPreprocess()
data_prep.fit(x_train, y_train)
x_train_enc, y_train_enc = data_prep.transform(x_train, y_train)
rf_clf = RandomForestClassifier(random_state=RANDOM_SEED)
grid_search = GridSearchCV(
estimator=rf_clf,
param_grid={
'max_depth': np.arange(3, 6),
'n_estimators': np.arange(70, 110, step=10),
},
scoring='f1_macro',
cv=5
)
grid_search.fit(x_train_enc, y_train_enc)
# -
# Да, GridSearch работает по такому же интерфейсу, как и любая модель, но важно отметить, что внутри него содержится масса полезных атрибутов! Давайте на них посмотрим!
# + _cell_id="XtpgWLnAnsXABjgB"
# Можно посмотреть сетку, которую перебирал GS
grid_search.cv_results_['params']
# + _cell_id="w8LGFruo6OYkeDop"
# Скор лучшего набора данных
grid_search.best_score_
# + _cell_id="h4IcRtN3MTFqZwO1"
# Лучшие параметры
grid_search.best_params_
# + _cell_id="2CUc9jz4RpJ1kNeT"
# И даже сразу лучшую обученную модель с лучшими параметрами
grid_search.best_estimator_
# -
# По сути, интерфейс GridSearch позволяет выполнять автоматический поиск гиперпараметров методом кросс-валидации, перебирая все возможные комбинации из заданных параметров.
#
# > ⚠️ Существует также и альтернативный способ поиска, который не ограничивается на исключительном переборе, а основывается на вероятностном выборе из распределений. Вы задаете конкретные диапазоны, а метод случайно выбирает значения и испытывает их. Поиск завершается, когда истекло заданное количество итераций, так как при работе с распределениями можно бесконечно делать выборки значений. Вот это метод - [RandomizedSearchCV](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.RandomizedSearchCV.html).
#
# Вот так мы познакомились с полезным инструментом для автоматизированного поиска гиперпараметров модели. Такая возможность позволяет искать гиперпараметры без каких-либо сложностей!
# ## Задание
#
# Сегодня мы научились важному методу поиска лучших гиперпараметров, так как их выбор может сильно сказаться на результатах работы с данными.
#
# Все сегодняшние инструменты позволят вам более гибко и уверенно производить разработку моделей для решения поставленных задач, что делает вас ещё круче, чем вы были раньше. Гиперпараметры моделей теперь вам ни по чём! Круто - поздравляем!
#
# Но не оставлять же вас без веселья? Как видите, у нас на руках датасет с очень внушительным объёмом - попробуйте решить поставленную на нём задачу, найдя наиболее подходящую модель и её гиперпараметры!
#
# * Задачка под зведочкой (подсказка) - если вы проведёте анализ данных и дополнительные обработки, то, вероятно, вам удастся улучшить результаты работы модели ещё сильнее! Дерзайте!
# ## Вопросы для закрепления
#
# А теперь пара вопросов, чтобы закрепить материал!
#
# 1. Почему кросс-валидацию не всегда выгодно использовать? В каких ситуациях?
# 2. Нужно ли задавать отдельную валидационную выборку для кросс-валидации?
# 3. Зачем нужен подбор гиперпараметров модели? Разве это не читерство?
# 4. Выгодно ли с точки зрения временных рамок указывать много разных параметров для GridSearchCV?
# 5. В чём разница между GridSearchCV и RandomizeSearchCV?
# ## Полезные ссылки
# * [Cross Validation от StatQuest](https://www.youtube.com/watch?v=fSytzGwwBVw)
#
| notebooks/32_Validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mrzhbr/mrzhbr.github.io/blob/main/GoogleColabPytorch1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="UdS7Mx6bkE7l"
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torchvision import datasets, transforms
from torch.autograd import Variable
import ssl
import matplotlib
ssl._create_default_https_context = ssl._create_unverified_context
# + colab={"base_uri": "https://localhost:8080/", "height": 290, "referenced_widgets": ["29b7535faa60457295370326ca4ea8b4", "b8bcb6967c06444f823f962eb53e84a3", "931a43444f6740af88450ad8e0e22028", "aadec7b3a4ce49b3ab9d096e6965a6be", "69732160794949c6af8bf895e5d26e95", "c4045a3177354342ad366e4479d0aa65", "3237f6112ad34497be7dba56a864c14c", "4f2e27d123ea4ee3bda627193b78e955"]} id="w1KAQIYekL5r" outputId="77f6affe-e334-4495-d1a8-8326100d6b18"
kwargs = {}
train_data = torch.utils.data.DataLoader(
datasets.EMNIST('data',split='byclass', train=True, download=True,
transform=transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.1307,),(0.3081,))])),
batch_size=256, shuffle=True, **kwargs)
test_data = torch.utils.data.DataLoader(
datasets.EMNIST('data',split='byclass', train=False,
transform=transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.1307,),(0.3081,))])),
batch_size=256, shuffle=True, **kwargs)
# + id="LmADZNGdkecM"
class Netz(nn.Module):
def __init__(self):
super(Netz, self).__init__()
self.conv1 = nn.Conv2d(1, 10, 5)
self.conv2 = nn.Conv2d(10, 20, 5)
self.dropout1 = nn.Dropout2d()
self.fc1 = nn.Linear(320, 128)
self.fc2 = nn.Linear(128, 62)
def forward(self, x):
x = self.conv1(x)
x = F.max_pool2d(x, 2)
x = F.relu(x)
x = self.conv2(x)
x = self.dropout1(x)
x = F.max_pool2d(x, 2)
x = F.relu(x)
x = x.view(-1, 320)
x = F.relu(self.fc1(x))
x = self.fc2(x)
return F.log_softmax(x)
print(x.size())
exit()
# + id="cZ80E01ski12"
model = Netz()
model.cuda()
optimizer = optim.SGD(model.parameters(), lr=0.1, momentum=0.3)
def train(epoch):
model.train()
for batch_id, (data, target) in enumerate(train_data):
data = data.cuda()
target = target.cuda()
data = Variable(data)
target = Variable(target)
optimizer.zero_grad()
out = model(data)
criterion = F.nll_loss
loss = criterion(out, target)
loss.backward()
optimizer.step()
print('Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}'.format
(epoch, batch_id*len(data), len(train_data.dataset),
100. * batch_id / len(train_data), loss.item()))
def test():
model.eval()
loss = 0
correct = 0
for data, target in test_data:
data = Variable(data.cuda(), volatile = True)
target = Variable(target.cuda())
out = model(data)
loss += F.nll_loss(out, target, reduction='sum').item()
prediction = out.data.argmax(dim=1, keepdim=True)
correct += prediction.eq(target.data.view_as(prediction)).sum().item()
loss = loss / len(test_data.dataset)
print('Durchschnittsloss', loss)
print('Accuracy: ', 100.*correct/len(test_data.dataset))
# + colab={"base_uri": "https://localhost:8080/"} id="9Pgu4lzZk1Gz" outputId="88a821ad-c8f6-4e0d-c624-2a376b3c1a60"
for epoch in range(1, 5):
train(epoch)
test()
test()
torch.save(model, 'emnist.pt')
# + colab={"base_uri": "https://localhost:8080/", "height": 528} id="YM0cO1CxvsPd" outputId="3025ecde-65b4-4bc7-c74e-3cdf0c853bd4"
from google.colab import drive
drive.mount('/content/drive')
# + id="Jun_j4aDc8Dl"
model = torch.load('emnist.pt')
# + colab={"base_uri": "https://localhost:8080/"} id="4B_KRB3Nof3v" outputId="3d54af2e-caa6-4aed-e52c-d65eefe6bce3"
test()
| GoogleColabPytorch1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mapping around with Python and Leaflet
#
# Leaflet is an amazing project used by R and Python and elswhere. It lets you create interactive maps with a number of basemap layers and other cool feature. Maps can then be embedded in web-apps or just used in your notbooks...or even screenshot-captured.
#
# In this notebook we explore some of the basic functionality.
# Tech info:
#
# If you are on a windows machine and dont have pip installed, start cmd and run:
#
# ```curl https://bootstrap.pypa.io/get-pip.py | python```
#
#
# Then run
#
# ```pip install ipywidgets ipyleaflet geopy```
#
#
#
# +
# Leaflet controls so called widgets (webdev stuff) and we don't really have to understand that
# What you need to know: Load up these libraries
# Unfortunately documentation for ipyleaflet (it's a rather new library)
# Widgets
from ipywidgets import HTML
# Leaflet
from ipyleaflet import *
# and of cause pandas for data manipulation
import pandas as pd
# -
# Let's start with a map of copenhagen
cph = Map(center=(55.677683, 12.569588), zoom=11)
#cph = Map(center=(55.677683, 12.569588), zoom=10, basemap=basemaps.Hydda.Full) # you can add other layers
cph
# +
# If you want, you can save the map as an html file and open in a browser (your data will be embedded)
from ipywidgets.embed import embed_minimal_html
embed_minimal_html('map.html',views=[z])
# +
# Let's set a marker on our map
# we first define it
marker = Marker(location=(55.692913, 12.599274), draggable=False)
# -
# And then place it on as a new layer
cph.add_layer(marker)
# Wouldn't it be nice to know what is in the spot where we placed the marker?
# Sure thing:
#
# 
# +
# We can create a "popup" where the value of the popup is a a bit of HTML code
# I disect it below
# We then assign the popup to our marker
popup = HTML()
popup.value = "The Little Mermaid <img src='https://upload.wikimedia.org/wikipedia/en/thumb/7/7a/Copenhagen_-_the_little_mermaid_statue_-_2013.jpg/1024px-Copenhagen_-_the_little_mermaid_statue_-_2013.jpg' alt='Mermaid'>"
marker.popup = popup
# -
# HTML: Text, then image, then a short image placeholder text (no rocket science)
#
# ```HTML
# "The Little Mermaid
# <img src='https://upload.wikimedia.org/wikipedia/en/thumb/7/7a/Copenhagen_-_the_little_mermaid_statue_-_2013.jpg/1024px-Copenhagen_-_the_little_mermaid_statue_-_2013.jpg'
# alt='Mermaid'>"
# ```
# Let's bring some more points on the map. This time public toilets in CPH you can get this data [here](https://data.kk.dk/dataset/toiletter)
# +
# Now let's try to plot in some very important data
# import the json library to deal with json data (more on that later)
import json
data = json.load(open('data/cph_toilets.json'))
# -
# Transform the data into a layer for our map
geo_json = GeoJSON(data=data)
# put it on
cph.add_layer(geo_json)
# and remove it again
chp.remove_layer(geo_json)
# +
# We can do better :-)
# Let's load up the WC data as a dataframe
wc = pd.DataFrame([x['properties'] for x in data['features']])
# -
# ```[x['properties'] for x in data['features']]``` is a list comprehension
# Python has these amazingly short loop structures built in
#
# Read:
#
# for each element (here x) in the dictionary "data" under the key "fatures"
# take the element and unpack the nested key "properties"
# put all of these in a list
#
# YES!!! We can pass a list of dictionaries to Pandas and it will still construct a neat dataframe
#
# Inspecting the comlumns
wc.columns
# Don't get scared: The next cell looks a bit tricky but it is not. We will now iterate over all rows of our dataframe and take out the location as well as create popup labels. All of that will be put into an empty list which we define in the very first row.
# In the last step of the loop we are adding our circles to the map
#
circles = []
for x in wc.iterrows():
loc = (x[1]['latitude'], x[1]['longitude'])
message = HTML()
message.value = x[1]['toilet_lokalitet'] + '<br />' + x[1]['adresse']
c = Circle(location=loc, radius=3, color = "#4CB391", fill_opacity = 0.4, popup = message)
circles.append(c)
cph.add_layer(c)
# We can also remove all of the by calling
for c in circles:
m.remove_layer(c)
# ## Geocoding
#
# Geocoding is super important and means simply transforming address data into geocoordinates
# Reverse geocoding is self explanatory
# +
# We will need to install and import geopy
# Geopy supports a large number of services (Google maps, Bing, Baidu etc.)
# We will use the Open Street Maps Nominatim (since that one doesn't require signing in and an API key)
from geopy.geocoders import Nominatim
geolocator = Nominatim(user_agent="SDS_geocode_teaching")
# -
# I created a little dataset of pubs in Aalborg
bodega = pd.read_csv('data/bodega.txt', sep=';')
bodega
# +
# Not th emost elegant approach but it does work and is easy to read
# Simple iteration over the rows and geocoding
lonlist = []
latlist = []
for i in bodega.iterrows():
loc = geolocator.geocode(i[1]['address'])
lonlist.append(loc.longitude)
latlist.append(loc.latitude)
# +
# Enter the values
bodega['lat'] = latlist
bodega['lon'] = lonlist
# -
bodega
# +
# A bit of a different map
aalborg = Map(center=(57.042340, 9.938773), zoom=14, basemap=basemaps.Stamen.Watercolor)
# -
aalborg
# +
# Let's do what we practiced above
circles = []
for x in bodega.iterrows():
loc = (x[1]['lat'], x[1]['lon'])
message = HTML()
message.value = x[1]['bodega'] + '<br />' + x[1]['address']
c = Circle(location=loc, radius=5, color = "#133366", fill_opacity = 0.6, popup = message)
circles.append(c)
aalborg.add_layer(c)
# -
# If you don't like the pubs they can go
for c in circles:
aalborg.remove_layer(c)
| M1_4_mapping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Geo
# language: python
# name: geo
# ---
# ## Imports
import jieba, json, os, re, sys, shutil, time
from datetime import datetime
import numpy as np
import tensorboardX
import torch
import torch.nn as nn
# ## Functions
# +
from fields import Field, Parms, Semantic, Vocab, _make_vocab
from utils import *
from nlp_db import nlp_db
from model_class import NLU_Classify
semantic = Semantic()
args = Parms()
vocab = Vocab(semantic)
args.manual_log = './manuaLog_lstm1.log'
# +
def read_json(file, thresh=np.infty, k=None, func=None):
with open(file, "r", encoding='utf-8') as f:
rzlt = []
cnt = 0
for l in f.readlines():
if k != None and func != None:
rzlt.append(func(json.loads(l)[k]))
elif k != None:
rzlt.append(json.loads(l)[k])
else:
rzlt.append(json.loads(l))
if cnt > thresh:
break
return rzlt
def json_iter(file, batch_size=1000, k=None, func=None):
with open(file, "r", encoding='utf-8') as f:
rzlt = []
for l in f.readlines():
if k != None and func != None:
rzlt.append(func(json.loads(l)[k]))
elif k != None:
rzlt.append(json.loads(l)[k])
else:
rzlt.append(json.loads(l))
if len(rzlt) == batch_size:
yield rzlt
rzlt = []
def restart_iter(batch_size, datafile):
x_iter = json_iter(file = datafile,
batch_size=batch_size,
k='sentence',
func = func_pad
)
y_iter = json_iter(file = datafile,
batch_size = batch_size,
k='label',
func = lambda x: label_rdict[x])
return x_iter, y_iter
def dump_log(log_file):
with open(log_file, 'a') as fp:
json.dump(
{
"epoch": last_epoch,
"loss": last_loss.data.item(),
"train_avg_acc": last_avgac.data.item(),
"dev_avg_acc": dev_acc.data.item()
}, fp)
fp.write('\n')
# with open(log_file, 'r') as f:
# last_lines = f.readlines()[-10:]
# with open(log_file, 'w') as f:
# f.write(last_lines)
# dump_log(args.manual_log)
# -
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
# ## Classification
# ### data file
rel_path = "../nlp_db/tnews_public"
cfiles = [
os.path.join(os.path.abspath(rel_path),
os.listdir(rel_path)[i])
for i in range(len(os.listdir(rel_path)))
]
print(cfiles)
testFile = cfiles[1]
trainFile = cfiles[2]
vocabFile = cfiles[3]
devFile = cfiles[-1]
testFile, trainFile, vocabFile, devFile
# ### Vocab Preprocess
# +
args.vocab_path = vocabFile
os.remove(args.vocab_path)
# # %psource _make_vocab
if not os.path.isfile(args.vocab_path):
_make_vocab(json_file=trainFile,
vocab_path=args.vocab_path,
thres=2,
level='word')
try:
vocab.load(args.vocab_path)
except:
print("Vocab not loaded")
vocab.size, vocab.__getitem__('吃'), vocab.__getitem__(
'<pad>'), vocab.__getitem__('<unk>'), vocab.__getitem__('<sos>')
# -
# ### Data Process => Model Parms Get
# +
labels = read_json(cfiles[0], 100, k='label')
label_rdict = {l:i for i,l in enumerate(labels)}
label_dict = {i:l for i,l in enumerate(labels)}
args.max_sent_len = max([
len(line) for line in read_json(
trainFile, k='sentence', func=lambda x: list(jieba.cut(x)))
])
args.class_num = len(labels)
args.lstm_step_num = 2
args.lstm_hid = 64
args.batch_size = 5000
dirrm(args)
# -
# ## Algorithm Process
# ### Forward, Loss
class NLU_Classify(nn.Module):
def __init__(self, class_num, vocab, args):
super(NLU_Classify, self).__init__()
self.type = 'classifier'
self.batch_size = args.batch_size
self.serial_len = 2
self.emb = nn.Embedding(vocab.size, embedding_dim=128)
self.lstm = nn.LSTM(128,
args.lstm_hid,
args.lstm_step_num,
batch_first=True)
self.fc = nn.Linear(64, class_num)
self.softmax = nn.Softmax(dim=1)
def forward(self, x, sent_lengths):
# # ? not sure serial_len , batch_size is 100% right
embedded_x = self.emb(x)
packed_embedded = nn.utils.rnn.pack_padded_sequence(embedded_x,
sent_lengths,
enforce_sorted=False,
batch_first=True)
h0 = torch.randn(self.serial_len, self.batch_size, args.lstm_hid, device = device)
c0 = torch.randn(self.serial_len, self.batch_size, args.lstm_hid, device = device)
x, (hidden, cn) = self.lstm(packed_embedded, (h0, c0))
hidden = hidden[-1,:,:]
output = self.fc(hidden)
output = self.softmax(output)
result = output
return result
# ### model, loss, optimizer - Definition
# +
model = NLU_Classify(class_num=args.class_num, vocab=vocab, args = args)
model.to(device)
loss_func = torch.nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(), lr=0.01, momentum=0.9)
# -
# ### Dev Eval
# +
dev_max_sent_len = max([
len(line) for line in read_json(
devFile, k='sentence', func=lambda x: list(jieba.cut(x)))
])
args.dev_max_sent_len = dev_max_sent_len
dirrm(args)
# -
# ### Training
# - Evaluation
# - Start from last checkpoint
# - Matrix Capture
# - Stop Rules ?
#
# +
# acc(model(eval_x, eval_sent_lengths), eval_y).data.item() 0.06359999626874924
# -
def acc(y_hat, y_label):
correct = (torch.argmax(y_hat, dim = 1) == y_label).float()
acc_rate = correct.sum() / len(correct)
return acc_rate
# +
def get_last_epoch(model_log):
with open(model_log, 'r') as f:
l = f.readlines()[-1]
last_epoch = json.loads(l.strip())['epoch']
return last_epoch
# get_last_epoch(args.manual_log)
# +
# args.max_sent_len has to be defined to def func_pad
def func_pad(sent, max_sent_len = args.max_sent_len):
return [vocab.__getitem__(token) for token in jieba.cut(sent)
] + [0] * (max_sent_len - len(list(jieba.cut(sent)))) , len(list(jieba.cut(sent)))
def restart_iter(batch_size, datafile):
x_iter = json_iter(
file=datafile,
batch_size=batch_size,
k='sentence',
func=lambda sent : func_pad(sent, max_sent_len=args.max_sent_len))
y_iter = json_iter(file=datafile,
batch_size=batch_size,
k='label',
func=lambda x: label_rdict[x])
return x_iter, y_iter
# +
# Prepare for Eval
eval_x, eval_sent_lengths = list(
zip(*read_json(
devFile,
k='sentence',
thresh=np.infty,
func=lambda sent: func_pad(sent, max_sent_len=args.dev_max_sent_len))))
eval_y = read_json(file=devFile, k='label', func=lambda x: label_rdict[x])
eval_sent_lengths = torch.tensor(eval_sent_lengths)
eval_x = torch.tensor(np.array([np.array(line) for line in eval_x]))
eval_y = torch.tensor(eval_y)
eval_x = eval_x.to(device)
eval_sent_lengths = eval_sent_lengths.to(device)
eval_y = eval_y.to(device)
# -
first_train = False
# +
# Training
if not os.path.isdir('./model_stores'):
os.mkdir('./model_stores')
args.model_path = './model_stores/model_lstm1.pth'
# Load:
if os.path.isfile(args.model_path) and first_train == False:
model.load_state_dict(torch.load(args.model_path))
model.train() # set model to train mode
if first_train:
last_epoch = 1
try:
shutil.rmtree(os.path.abspath('./runs/'))
os.remove(os.path.abspath(args.manual_log))
except:
pass
else:
last_epoch = get_last_epoch(args.manual_log)
writer_train = tensorboardX.SummaryWriter('runs/train_0')
writer_test = tensorboardX.SummaryWriter('runs/test_0')
writer = tensorboardX.SummaryWriter('runs/net_0')
writer.add_graph(model, (eval_x, eval_sent_lengths))
epoch = last_epoch
if not "acc_rates" in locals():
acc_rates = [0] * 10
while True:
# while np.array(acc_rates).sum() / len(acc_rates) < 0.8:
epoch += 1
x_iter, y_iter = restart_iter(args.batch_size, trainFile)
ep_cnt = 0
acc_loss = []
acc_rates = []
for batch_x, batch_y in zip(x_iter, y_iter):
model.train()
batch_x, sent_lengths = list(zip(*batch_x))
batch_x = torch.tensor(np.array([np.array(line) for line in batch_x]))
sent_lengths = torch.tensor(sent_lengths)
batch_y = torch.tensor(batch_y)
batch_x = batch_x.to(device)
sent_lengths = sent_lengths.to(device)
batch_y = batch_y.to(device)
optimizer.zero_grad()
y_hat = model(batch_x, sent_lengths)
loss = loss_func(y_hat, batch_y)
loss.backward()
# loss.backward(retain_graph=True)
optimizer.step()
acc_rate = acc(y_hat, batch_y)
ep_cnt += 1
acc_loss.append(loss)
acc_rates.append(acc_rate)
if ep_cnt % 10 == 0:
# get metrics
idx = epoch + 0.32 * (ep_cnt % 20)
last_loss, last_avgac = np.array(acc_loss).sum() / len(
acc_loss), np.array(acc_rates).sum() / len(acc_rates)
print(epoch, "loss: ", last_loss.data.item(), "Acc: ", last_avgac.data.item())
writer_train.add_scalar('loss', last_loss.data.item(), idx)
writer_train.add_scalar('train_avgAcc:', last_avgac.data.item(), idx)
# writer.add_scalar('loss:', last_loss, epoch + 0.32 * (ep_cnt % 20))
acc_loss = []
acc_rates = []
# Save Model Parameters:
torch.save(model.state_dict(), f=args.model_path)
# Eval
model.eval()
yhat = model(eval_x, eval_sent_lengths)
dev_acc = acc(yhat, eval_y)
print(epoch, "dev_acc: ", dev_acc.data.item())
writer_test.add_scalar('dev_avgAcc', dev_acc.data.item(), idx)
last_epoch = epoch
dump_log(args.manual_log)
# -
| model_train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="4sv1j8lRtwKQ" executionInfo={"status": "ok", "timestamp": 1630069531557, "user_tz": -330, "elapsed": 851, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
import os
project_name = "reco-tut-bok"; branch = "main"; account = "sparsh-ai"
project_path = os.path.join('/content', project_name)
# + id="n8RiXVLstuoO" executionInfo={"status": "ok", "timestamp": 1630069533175, "user_tz": -330, "elapsed": 1630, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="7185e8e7-aa13-4c54-e6ab-e1864aa79575" colab={"base_uri": "https://localhost:8080/"}
if not os.path.exists(project_path):
# !cp /content/drive/MyDrive/mykeys.py /content
import mykeys
# !rm /content/mykeys.py
path = "/content/" + project_name;
# !mkdir "{path}"
# %cd "{path}"
import sys; sys.path.append(path)
# !git config --global user.email "<EMAIL>"
# !git config --global user.name "reco-tut"
# !git init
# !git remote add origin https://"{mykeys.git_token}":x-oauth-basic@github.com/"{account}"/"{project_name}".git
# !git pull origin "{branch}"
# !git checkout main
else:
# %cd "{project_path}"
# + id="drkDUesKtuoT"
# !git status
# + id="pWkwGc0ltuoT"
# !git add . && git commit -m 'commit' && git push origin "{branch}"
# + id="KQoB06RmtuoU"
import glob
files = sorted(glob.glob(f'/content/drive/MyDrive/Colab Notebooks/{project_name}*.ipynb'))
files
# + id="h7le8ERJtuoV"
from shutil import copy
for fpath in files[1:]:
copy(fpath, './notebooks')
# + id="zWGLaL1-tuoV" language="sh"
# mkdir -p ./code/nbs
# for file in ./notebooks/*.ipynb
# do
# jupyter nbconvert --output-dir="./code/nbs" --to python "$file"
# done
# + id="A07k37WmtuoV"
# !sudo apt-get install -qq tree
# + id="Fc0TebaStuoW"
# !tree -L 3 .
# !tree --du -h
# + id="-enykZkUtuoW"
# %%writefile README.md
#
## Project structure
```
.
├── artifacts
```
# + id="lkyg1LhVtuoW"
xproject_name = "reco-nb-stage"; xbranch = "queued"; xaccount = "recohut"
xproject_path = os.path.join('/content', xproject_name)
if not os.path.exists(xproject_path):
# !cp /content/drive/MyDrive/mykeys.py /content
import mykeys
# !rm /content/mykeys.py
path = "/content/" + xproject_name;
# !mkdir "{path}"
# %cd "{path}"
import sys; sys.path.append(path)
# !git config --global user.email "<EMAIL>"
# !git config --global user.name "reco-tut"
# !git init
# !git remote add origin https://"{mykeys.git_token}":x-oauth-basic@github.com/"{xaccount}"/"{xproject_name}".git
# !git pull origin "{xbranch}"
else:
# %cd "{xproject_path}"
# + id="0-b3EHsItuoX"
# !git checkout -b queued
# !cp $project_path/notebooks/*.ipynb ./_notebooks
# !git add . && git commit -m 'commit' && git push origin queued
| _docs/nbs/reco-tut-bok-00-launchpad.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Demonstration of MUSE
#
# This is a demonstration of MUSE analysis on a multi-modality simulated data.
#
# Altschuler & Wu Lab 2020.
#
# Software provided as is under MIT License.
# ## Import packages
# +
import muse_sc as muse
import simulation_tool.multi_modal_simulation as simulation
import phenograph
from sklearn.decomposition import PCA
import numpy as np
from sklearn.metrics.cluster import adjusted_rand_score
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE
import tensorflow as tf
tf.get_logger().setLevel('ERROR')
np.random.seed(0)
# -
# ## Generate simulation data
#
# Simulation parameters
latent_dim = 100
num_cluster = 10
sample_size = 1000
latent_code_dim = 30
observed_data_dim = 500
sigma_1 = 0.1
sigma_2 = 0.1
decay_coef_1 = 0.5
decay_coef_2 = 0.1
merge_prob = 0.7
#
#
# Use simulation tool to generate multi-modality data
data = simulation.multi_modal_simulator(num_cluster, sample_size,
observed_data_dim, observed_data_dim,
latent_code_dim,
sigma_1, sigma_2,
decay_coef_1, decay_coef_2,
merge_prob)
data_a = data['data_a_dropout']
data_b = data['data_b_dropout']
label_a = data['data_a_label']
label_b = data['data_b_label']
label_true = data['true_cluster']
# ## Analyses based on single modality
# Learn features from single modality
view_a_feature = PCA(n_components=latent_dim).fit_transform(data_a)
view_b_feature = PCA(n_components=latent_dim).fit_transform(data_b)
# Perform clustering using PhenoGraph
view_a_label, _, _ = phenograph.cluster(view_a_feature)
view_b_label, _, _ = phenograph.cluster(view_b_feature)
# ## Combined analysis using MUSE
# MUSE learns the joint latent representation
muse_feature, reconstruct_x, reconstruct_y, \
latent_x, latent_y = muse.muse_fit_predict(data_a,
data_b,
view_a_label,
view_b_label,
latent_dim=100,
n_epochs=500,
lambda_regul=5,
lambda_super=5)
# ## Perform clustering
# PhenoGraph clustering
muse_label, _, _ = phenograph.cluster(muse_feature)
# ## Visualization of latent spaces
# Latent spaces of single-modality features or MUSE features were visualized using tSNE, with ground truth cluster labels.
#
# Cluster accuries were quantified using adjusted Rand index (ARI). ARI = 1 indicates perfectly discover true cell identities.
# +
plt.figure(figsize=(17, 5))
plt.subplot(1, 3, 1)
X_embedded = TSNE(n_components=2).fit_transform(view_a_feature)
for i in np.unique(label_true):
idx = np.nonzero(label_true == i)[0]
plt.scatter(X_embedded[idx, 0], X_embedded[idx, 1])
plt.title('Transcript-alone, ARI = %01.3f' % adjusted_rand_score(label_true, view_a_label))
plt.subplot(1, 3, 2)
X_embedded = TSNE(n_components=2).fit_transform(view_b_feature)
for i in np.unique(label_true):
idx = np.nonzero(label_true == i)[0]
plt.scatter(X_embedded[idx, 0], X_embedded[idx, 1])
plt.title('Morphology-alone, ARI = %01.3f' % adjusted_rand_score(label_true, view_b_label))
plt.subplot(1, 3, 3)
X_embedded = TSNE(n_components=2).fit_transform(muse_feature)
for i in np.unique(label_true):
idx = np.nonzero(label_true == i)[0]
plt.scatter(X_embedded[idx, 0], X_embedded[idx, 1])
plt.title('MUSE, ARI = %01.3f' % adjusted_rand_score(label_true, muse_label))
# -
| MUSE_demo/MUSE_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Kvalitetskontroll av budgetdata 2022
import pandas as pd
# %config InlineBackend.figure_format = 'retina'
# ## Detailed party budgets
# +
opposition_parties = ['c', 'kd', 'l', 'm', 'sd', 'v']
data_path = '../data'
frames = []
for party in opposition_parties:
frame = pd.read_csv(f'{data_path}/budgetmotion-2022-{party}.csv', dtype=str)
frame['Parti'] = party.upper()
frames.append(frame)
# -
df = pd.concat(frames, sort=False)
df['Utgiftsområde'] = df['Utgiftsområde'].astype(int)
df['2022'] = df['2022'].astype(int)
df['2023'] = df['2023'].astype(float).astype('Int64')
df['2024'] = df['2024'].astype(float).astype('Int64')
# ## Summary party budgets
sums = pd.read_csv('../data/budgetmotion-2022-summor.csv')
sums = sums[sums['Utgiftsområde'] < 26]
sums = sums.set_index(['Parti', 'Utgiftsområde'])
sums = sums.drop('Namn', axis=1)
# Sums derived from detailed posts
calc_sums = df.groupby(['Parti', 'Utgiftsområde']).sum()
# ## Sanity check
# Convert to MSEK for better overview
sums = sums / 1_000_000
calc_sums = calc_sums / 1_000_000
calc_sums.head()
sums.head()
sums.shape
sums.index.difference(calc_sums.index)
# Drop the diffing rows
sums = sums.drop(sums.index.difference(calc_sums.index))
# Rows that differ for 2022
_ = '2022'
calc_sums[(calc_sums[_].abs() - sums[_].abs() > 2)].merge(sums, left_index=True, right_index=True)[[f'{_}_x', f'{_}_y']]
# Rows that differ for 2023
_ = '2023'
calc_sums[(calc_sums[_].abs() - sums[_].abs() > 2)].merge(sums, left_index=True, right_index=True)[[f'{_}_x', f'{_}_y']]
# Rows that differ for 2024
_ = '2024'
calc_sums[(calc_sums[_].abs() - sums[_].abs() > 2)].merge(sums, left_index=True, right_index=True)[[f'{_}_x', f'{_}_y']]
piv = df.drop(['2023', '2024', 'Namn'], axis=1)
piv = piv[~piv.Anslag.str.startswith('99', na=False)]
piv['2022'] = piv['2022'].astype(float) / 1_000_000
piv = piv.pivot_table(index=['Utgiftsområde', 'Anslag'], columns='Parti', values='2022')
piv = piv.fillna(0)
piv
((piv.apply(lambda x: x == 0, axis=1).sum().sort_values() + 2) / 492 * 100).plot.barh(xlim=(0, 100), grid=True)
df[df.Anslag.str.startswith('99', na=False)].Parti.value_counts()
name_map = df[['Utgiftsområde', 'Anslag', 'Namn']].drop_duplicates(subset=['Utgiftsområde', 'Anslag'])
df = (df
.pivot_table(index=['Anslag', 'Utgiftsområde'],
columns=['Parti'],
values='2022')
.reset_index())
df = df.fillna(0)
gov = pd.read_csv('../data/budgetproposition-2022-regering.csv', dtype=str)
gov[gov.Utgiftsområde.astype(int) < 26].dropna(subset=['Anslag', 'Underområde'], how='any')
gov['Anslag'] = gov['Underområde'] + ':' + gov['Anslag']
gov[~gov.Anslag.isin(df.Anslag)]
gov.Anslag.unique().shape
gov['Kod'] = gov['Utgiftsområde'] + ':' + gov.Anslag
df['Kod'] = df['Utgiftsområde'].astype(str) + ':' + df.Anslag
gov[~gov.Kod.isin(df.Kod)]
gov = gov.rename(columns={'2022': 'Regeringen'})
gov = gov.dropna(subset=['Anslag'])
df = df.merge(gov[['Anslag', 'Utgiftsområde', 'Regeringen', 'Namn']], on=['Utgiftsområde', 'Anslag'], how='left')
df = df[['Utgiftsområde', 'Anslag', 'Namn', 'Regeringen', 'C', 'KD', 'L', 'M', 'SD', 'V']]
df = df.fillna(0)
df.Regeringen = df.Regeringen.astype(float)
df.shape
for col in ['C', 'KD', 'L', 'M', 'SD', 'V']:
if not df[(df[col].abs() > df.Regeringen) & (df[col] < 0)].empty:
print(col)
df[(df['V'].abs() > df.Regeringen) & (df['V'] < 0)]
df.to_csv('../data/budgetmotion-2022-main.csv', index=False)
for col in ['C', 'KD', 'L', 'M', 'SD', 'V']:
df[col] = df[col].astype(int) / 1_000_000
for col in ['Regeringen']:
df[col] = df[col].astype(float) / 1_000_000
for col in ['C', 'KD', 'L', 'M', 'SD', 'V']:
df[f'{col}_prc'] = df[col] / df['Regeringen']
df
p = df[df.Namn == 'Polismyndigheten']
p
p
ax = p[['C', 'KD', 'L', 'M', 'SD', 'V']].T.sort_values(29).plot.barh(legend=False)
ax.axvline(1, c='#000000')
| notebooks/4_kvalitetskoll.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cubeenv
# language: python
# name: cubeenv
# ---
# This is a quick notebook to demonstrate the pyroSAR functionality for importing processed SAR scenes into an Open Data Cube
from pyroSAR.datacube_util import Product, Dataset
from pyroSAR.ancillary import groupby, find_datasets
# +
# define a directory containing processed SAR scenes
dir = '/path/to/some/data'
# define a name for the product YML; this is used for creating a new product in the datacube
yml_product = './product_def.yml'
# define a directory for storing the indexing YMLs; these are used to index the dataset in the datacube
yml_index_outdir = './yml_indexing'
# define a name for the ingestion YML; this is used to ingest the indexed datasets into the datacube
yml_ingest = './ingestion.yml'
# product description
product_name_indexed = 'S1_GRD_index'
product_name_ingested = 'S1_GRD_ingest'
product_type = 'gamma0'
description = 'this is just some test'
# define the units of the dataset measurements (i.e. polarizations)
units = 'backscatter'
# alternatively this could be a dictionary:
# units = {'VV': 'backscatter VV', 'VH': 'backscatter VH'}
ingest_location = './ingest'
# +
# find pyroSAR files by metadata attributes
files = find_datasets(dir, recursive=True, sensor=('S1A', 'S1B'), acquisition_mode='IW')
# group the found files by their file basenames
# files with the same basename are considered to belong to the same dataset
grouped = groupby(files, 'outname_base')
# -
print(len(files))
print(len(grouped))
# In the next step we create a new product, add the grouped datasets to it and create YML files for indexing the datasets in the cube.
# create a new product and add the collected datasets to it
# alternatively, an existing product can be used by providing the corresponding product YML file
with Product(name=product_name_indexed,
product_type=product_type,
description=description) as prod:
for dataset in grouped:
with Dataset(dataset, units=units) as ds:
# add the datasets to the product
# this will generalize the metadata from those datasets to measurement descriptions,
# which define the product definition
prod.add(ds)
# parse datacube indexing YMLs from product and dataset metadata
prod.export_indexing_yml(ds, yml_index_outdir)
# write the product YML
prod.write(yml_product)
# print the product metadata, which is written to the product YML
print(prod)
# Now that we have a YML file for creating a new product and individual YML files for indexing the datasets, we can create a last YML file, which will ingest the indexed datasets into the cube. For this a new product is created and the files are converted to NetCDF, which are optimised for useage in the cube. The location of those NetCDF files also needs to be defined.
with Product(yml_product) as prod:
prod.export_ingestion_yml(yml_ingest, product_name_ingested, ingest_location)
| datacube_prepare.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="_dEaVsqSgNyQ"
# ##### Copyright 2020 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="4FyfuZX-gTKS"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="sT8AyHRMNh41"
# # TensorFlow Recommenders: Quickstart
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/recommenders/quickstart"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/recommenders/blob/main/docs/examples/quickstart.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/recommenders/blob/main/docs/examples/quickstart.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/recommenders/docs/examples/quickstart.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="8f-reQ11gbLB"
# In this tutorial, we build a simple matrix factorization model using the [MovieLens 100K dataset](https://grouplens.org/datasets/movielens/100k/) with TFRS. We can use this model to recommend movies for a given user.
# + [markdown] colab_type="text" id="qA00wBE2Ntdm"
# ### Import TFRS
#
# First, install and import TFRS:
# + colab={} colab_type="code" id="6yzAaM85Z12D"
# !pip install -q tensorflow-recommenders
# !pip install -q --upgrade tensorflow-datasets
# + colab={} colab_type="code" id="n3oYt3R6Nr9l"
from typing import Dict, Text
import numpy as np
import tensorflow as tf
import tensorflow_datasets as tfds
import tensorflow_recommenders as tfrs
# + [markdown] colab_type="text" id="zCxQ1CZcO2wh"
# ### Read the data
# + colab={} colab_type="code" id="M-mxBYjdO5m7"
# Ratings data.
ratings = tfds.load('movie_lens/100k-ratings', split="train")
# Features of all the available movies.
movies = tfds.load('movie_lens/100k-movies', split="train")
# Select the basic features.
ratings = ratings.map(lambda x: {
"movie_title": x["movie_title"],
"user_id": x["user_id"]
})
movies = movies.map(lambda x: x["movie_title"])
# + [markdown] colab_type="text" id="5W0HSfmSNCWm"
# Build vocabularies to convert user ids and movie titles into integer indices for embedding layers:
# + colab={} colab_type="code" id="9I1VTEjHzpfX"
user_ids_vocabulary = tf.keras.layers.experimental.preprocessing.StringLookup()
user_ids_vocabulary.adapt(ratings.map(lambda x: x["user_id"]))
movie_titles_vocabulary = tf.keras.layers.experimental.preprocessing.StringLookup()
movie_titles_vocabulary.adapt(movies)
# + [markdown] colab_type="text" id="Lrch6rVBOB9Q"
# ### Define a model
#
# We can define a TFRS model by inheriting from `tfrs.Model` and implementing the `compute_loss` method:
# + colab={} colab_type="code" id="e5dNbDZwOIHR"
class MovieLensModel(tfrs.Model):
# We derive from a custom base class to help reduce boilerplate. Under the hood,
# these are still plain Keras Models.
def __init__(
self,
user_model: tf.keras.Model,
movie_model: tf.keras.Model,
task: tfrs.tasks.Retrieval):
super().__init__()
# Set up user and movie representations.
self.user_model = user_model
self.movie_model = movie_model
# Set up a retrieval task.
self.task = task
def compute_loss(self, features: Dict[Text, tf.Tensor], training=False) -> tf.Tensor:
# Define how the loss is computed.
user_embeddings = self.user_model(features["user_id"])
movie_embeddings = self.movie_model(features["movie_title"])
return self.task(user_embeddings, movie_embeddings)
# + [markdown] colab_type="text" id="wdwtgUCEOI8y"
# Define the two models and the retrieval task.
# + colab={} colab_type="code" id="EvtnUN6aUY4U"
# Define user and movie models.
user_model = tf.keras.Sequential([
user_ids_vocabulary,
tf.keras.layers.Embedding(user_ids_vocabulary.vocab_size(), 64)
])
movie_model = tf.keras.Sequential([
movie_titles_vocabulary,
tf.keras.layers.Embedding(movie_titles_vocabulary.vocab_size(), 64)
])
# Define your objectives.
task = tfrs.tasks.Retrieval(metrics=tfrs.metrics.FactorizedTopK(
movies.batch(128).map(movie_model)
)
)
# + [markdown] colab_type="text" id="BMV0HpzmJGWk"
#
# ### Fit and evaluate it.
#
# Create the model, train it, and generate predictions:
#
#
# + colab={} colab_type="code" id="H2tQDhqkOKf1"
# Create a retrieval model.
model = MovieLensModel(user_model, movie_model, task)
model.compile(optimizer=tf.keras.optimizers.Adagrad(0.5))
# Train for 3 epochs.
model.fit(ratings.batch(4096), epochs=3)
# Use brute-force search to set up retrieval using the trained representations.
index = tfrs.layers.ann.BruteForce(model.user_model)
index.index(movies.batch(100).map(model.movie_model), movies)
# Get some recommendations.
_, titles = index(np.array(["42"]))
print(f"Top 3 recommendations for user 42: {titles[0, :3]}")
# + colab={} colab_type="code" id="neJAJVwbReNd"
| docs/examples/quickstart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **References:**
# - https://pandas.pydata.org/pandas-docs/stable/advanced.html
# - https://www.somebits.com/~nelson/pandas-multiindex-slice-demo.html
import pandas as pd
import io
data = io.StringIO('''Fruit,Color,Count,Price
Apple,Red,3,$1.29
Apple,Green,9,$0.99
Pear,Red,25,$2.59
Pear,Green,26,$2.79
Lime,Green,99,$0.39
''')
df_unindexed = pd.read_csv(data)
df_unindexed
df = df_unindexed.set_index(['Fruit', 'Color'])
df
df.xs('Apple')
| python/modules/pandas/notebooks/multi_index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# ## {{cookiecutter.project_name}}
#
# {{cookiecutter.description}}
#
# This notebook contains basic statistical analysis and visualization of the data.
#
# ### Data Sources
# - summary : Processed file from notebook 1-Data_Prep
#
# ### Changes
# - {% now 'utc', '%m-%d-%Y' %} : Started project
# +
import pandas as pd
import numpy as np
from numpy import log
from numpy.random import randn
import glob
import datetime as dt
import pickle
import os
from pandas import ExcelWriter
import re
from zipfile import ZipFile
import matplotlib.pyplot as plt
from matplotlib.ticker import PercentFormatter
import seaborn as sns
import datetime
from sklearn import preprocessing
from sklearn.preprocessing import PowerTransformer
import scipy.stats as stats
from scipy.stats import normaltest
from scipy.stats import shapiro
from scipy.stats import anderson
from scipy.stats import boxcox
from scipy.stats import norm
from math import erf, sqrt
# -
# %matplotlib inline
# ### Functions
# +
#80/20 analysis
def pareto(df, rows, columns, sortcol, colmonth1, colmonth2, colmonth3, minvalue):
'''df= dataframe to use
rows= column to use for rows
columns= name of column to use as rows
sortcol = column to sumarrize ej: tickets, cases, persons, etc)
colmonth1:colmonth3 = columns to calculate average for columns
minvalue = value to filter the result, will show recrods with values greater than minvalue
'''
crostab= pd.crosstab(df[rows],df[columns],margins=True)
crostab.sort_values(sortcol, ascending=False,inplace=True)
crostab= crostab.drop(['All'])
print('Total of rows: {}'.format(len(crostab)))
crostab['pc']= 100*crostab[sortcol]/crostab[sortcol].sum()
crostab['cum_pc']=crostab['pc'].cumsum()
crostab['AVG3M']= (crostab[colmonth1] + crostab[colmonth2] + crostab[colmonth3]) /3
print('Total of rows up to 80%: {}'.format(len(crostab[crostab['cum_pc'] < 81])))
print('{} Total of rows below average of {}'.format(len(crostab[crostab['AVG3M'] <= minvalue]), minvalue))
print('to print the table run: crostab2[crostab2["AVG3M"] > 5]')
return crostab
#distribution
def gethrdistribution(df, group1, agg1, titletxt= 'Pie Chart', minpercent=5, filename='figpie.png'):
'''pie distributions per group
consolidate % < 10% in others category
'''
dist1= df.groupby(group1,as_index=False)[agg1].count()
dist1['pc']= 100*dist1[agg1]/dist1[agg1].sum()
dist1[group1]= np.where(dist1['pc']<minpercent,'Others',dist1[group1])
dist1= dist1.groupby(group1,as_index=False)[agg1].sum()
dist1['pc']= 100*dist1[agg1]/dist1[agg1].sum()
dist1= dist1.sort_values('pc', ascending=False)
dist1.reindex(copy=False)
dist1['cum_pc']=dist1['pc'].cumsum()
# Create a list of colors (from iWantHue)
colors = [ '#959a3c', '#55ac69', '#5b86d0', "#E13F29", "#D69A80", "#D63B59",
"#AE5552", "#CB5C3B", "#EB8076", "#96624E" ]
# Create a pie chart
fig, ax = plt.subplots()
plt.pie(
dist1[agg1], # using data agg1
labels=dist1[group1],# with the labels being group1
shadow=False, # with no shadows
colors=colors, # with colors
#explode=(0, 0.15, 0), # with one slide exploded out
# with the start angle at 90%
startangle=90, # with the start angle at 90%
autopct='%1.1f%%', # with the percent listed as a fraction
counterclock= False
)
# View the plot drop above
plt.axis('equal')
plt.title(titletxt)
# View the plot
plt.tight_layout()
plt.show()
figname_file= os.path.join(figures_path,
directory_name + '_' + filename + '{:%m%d%y}.png').format(today)
fig.savefig(figname_file, transparent= True)
dist1= dist1.sort_values('pc', ascending=False)
print(dist1)
def plottickets(df, group1, group2, countfield):
'''plot df grouped by group1 and group2 and counting countfield'''
ts=df.groupby([group1,group2]).agg({countfield: 'count'})
#ts.sort_values(group1, ascending=True,inplace=True)
ts.plot(kind= 'line')
return ts
def weedaysbars(df, group1, agg1, title, xlab, ylab, filename='figbarcharth.png'):
'''function to display bar chart, ej criticality, or weekdays barcharts'''
weekdays= df.groupby(group1,as_index=False)[agg1].count()
fig, ax = plt.subplots()
#plt.bar(weekdays[group1], height= weekdays[agg1], color='#607c8e')
ax.bar(weekdays[group1], height= weekdays[agg1], color='#607c8e')
width = 0.75 # the width of the bars
ax.barh(df.index, df['number'], width)
plt.title(title)
plt.xlabel(xlab)
plt.ylabel(ylab)
plt.grid(axis='y', alpha=0.75)
###
#for i, v in enumerate(weekdays[group1]):
# ax.text(v + 3, i + .0, str(v))
ax.patch.set_visible(False)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['bottom'].set_linewidth(0.5)
ax.spines['left'].set_linewidth(0.5)
###
plt.show()
figname_file= os.path.join(figures_path,
directory_name + '_' + filename + '{:%m%d%y}.png').format(today)
fig.savefig(figname_file, transparent= True)
print(weekdays)
def weedaysbarsh(df, group1, agg1, title, xlab, ylab, filename='figbarcharth.png'):
'''function to display bar chart, ej criticality, or weekdays barcharts'''
weekdays= df.groupby(group1,as_index=False)[agg1].count()
fig, ax = plt.subplots()
width = 0.75 # the width of the bars
ax.barh(weekdays[group1], weekdays[agg1], width)
plt.title(title)
plt.xlabel(xlab)
plt.ylabel(ylab)
plt.grid(axis='y', alpha=0.75)
for i, v in enumerate(weekdays[agg1]):
ax.text(v + 3, i + .0, str(v))
ax.patch.set_visible(False)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['bottom'].set_linewidth(0.5)
ax.spines['left'].set_linewidth(0.5)
plt.show()
figname_file= os.path.join(figures_path,
directory_name + '_' + filename + '{:%m%d%y}.png').format(today)
fig.savefig(figname_file, transparent= True)
print(weekdays)
#cycle_time3
def cycletime3(df, groupby2, groupby3, agg1, agg2, agg3):
'''Caclulate cycletime per vendor just for request and incidents
usage: cycletime2(nuclear0,'yearmonth','Vendor_Closeassign','cycletime','number',
'closed_by_user_name', 'cycletime')
'''
df = df[df.Type.isin(['Requested Item','Incident'])]
#cycle_time and FTE
df2= df.groupby([groupby2,groupby3]).agg({agg1: ['mean','std','max','min'],
agg2: 'count',agg3: 'nunique'})
# Using ravel, and a string join, we can create better names for the columns:
df2.columns = ["_".join(x) for x in df2.columns.ravel()]
agg5= agg3 + '_nunique'
agg6= agg2 + '_count'
agg7= agg1 + '_mean'
# per month
df2= df2.groupby([groupby3]).agg({agg5: ['mean', 'std'], agg6: ['mean','count', 'median','max'],
agg7: ['mean','std', 'median']})
return df2
def barchart(df,x,y,title, x_label, y_label,filename='figbarchart.png'):
'''bar chart tickets per organizatio x_Vendor_Closeassign or vendor'''
field_vendor = 'x_Vendor_Closeassign'
field_vendor = 'x_vendor'
pt_df= df.pivot_table(x, index=[y],
aggfunc='count',
margins=True)#.sort_values(('SSO','All'), ascending=False)
pt_df.index.rename(y_label, inplace= True)
#remove rows with cero count of tickets
pt_df= pt_df[pt_df[x] >0].sort_values(x, ascending=True)
fig, ax = plt.subplots()
width = 0.75 # the width of the bars
ax.barh(pt_df.index, pt_df[x], width)
plt.title(title)
plt.xlabel(x_label)
plt.ylabel(y_label)
for i, v in enumerate(pt_df[x]):
ax.text(v + 3, i + .0, str(v))
ax.patch.set_visible(False)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['bottom'].set_linewidth(0.5)
ax.spines['left'].set_linewidth(0.5)
plt.show()
figname_file= os.path.join(figures_path,
directory_name + '_' + filename + '{:%m%d%y}.png').format(today)
fig.savefig(figname_file, transparent= True)
def histogram(df,x, title, x_label, y_label, filter_in, filename= 'histogram'):
#histogram aging tickets
df_agging=df[(df.x_agingdays > 0) &(df.Type.isin(filter_in))]
df_agging= df_agging[x]
fig, ax = plt.subplots()
ax.hist(df_agging, bins=10)
plt.title(title)
plt.xlabel(x_label)
plt.ylabel(y_label)
ax.patch.set_visible(False)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['bottom'].set_linewidth(0.5)
ax.spines['left'].set_linewidth(0.5)
plt.show()
figname_file= os.path.join(figures_path,
directory_name + '_' + filename + '{:%m%d%y}.png').format(today)
fig.savefig(figname_file, transparent= True)
#df_agging.plot.hist(bins=10, title= 'Aging tickets')
print(df_agging.describe())
df_aggingsum= df[(df.x_agingdays > 0) & (df.Type.isin(filter_in))]
aggingsum= df_aggingsum.groupby(['x_vendor',
'Type']).agg({'x_aging': 'sum',
'number':'count',
'x_agingdays':['mean',
'std','median']}).sort_values('x_vendor',
ascending=False)
aggingsum.rename(columns = {'sum':'Open','count':'Closed',
'std': 'Std Dev',
'mean':'Mean', 'number':'','x_aging':'', 'x_agingdays':''}, inplace= True)
print(aggingsum)
def group_by(df):
''' group by df to report in xls file
'''
#groub by 'yearmonth', 'dayweek', 'hourday', 'cmdb_ci_name','PandL'
grouped= df.groupby(['x_yearmonth', 'x_dayweek', 'x_hourday', 'cmdb_ci_name',
'PandL'],
as_index = False).agg({'closed_by_user_name' :['count', 'nunique'],
'number' : 'count'})
grouped.columns = ["_".join(x) for x in grouped.columns.ravel()]
#groub by 'yearmonth', 'cmdb_ci_name', 'PandL'
grouped1= df.groupby(['x_yearmonth', 'cmdb_ci_name', 'PandL'],
as_index = False).agg({'closed_by_user_name' :['count', 'nunique'],
'number' : 'count'})
grouped1.columns = ["_".join(x) for x in grouped1.columns.ravel()]
#groub by file 'yearmonth', 'PandL'
grouped2= df.groupby(['x_yearmonth', 'PandL'], as_index = False).agg({'number' : 'count'})
return (grouped, grouped1, grouped2)
def verify_normality(df, column):
''' graph distribution for a column, with values > 0
'''
print(df[column].describe())
df2= df[df[column] > 0]
arr = df2[column]
mean=arr.mean()
median=arr.median()
mode=arr.mode()
print('Mean: ',mean,'\nMedian: ',median,'\nMode: ',mode[0])
arr = sorted(arr)
fit = stats.norm.pdf(arr, np.mean(arr), np.std(arr))
#plot both series on the histogram
fig, ax = plt.subplots()
plt.axvline(mean,color='red',label='Mean')
plt.axvline(median,color='yellow',label='Median')
plt.axvline(mode[0],color='green',label='Mode')
plt.plot(arr,fit,'-',linewidth = 2,label="Normal distribution with same mean and var")
plt.hist(arr,density=True,bins = 10,label="Actual distribution")
ax.patch.set_visible(False)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['bottom'].set_linewidth(0.5)
ax.spines['left'].set_linewidth(0.5)
#plt.title('Histogram {}'.format(column))
plt.legend()
plt.show()
fig = plt.figure()
ax1 = fig.add_subplot(211)
prob = stats.probplot(df2[column], dist=stats.norm, plot=ax1)
ax1.set_xlabel('')
ax1.set_title('Probplot against normal distribution')
def transform(df, column, method='power'):
'''Transform a column using log,scale, minmax, boxcox, power, norm
filter out rows with values <=0, takes only positive values.
'''
dfnorm= pd.DataFrame()
df= df[df[column] > 0]
df[column]= df[column].fillna(df[column].mean())
dfnorm['x_original']= df[column]
print(df[column].describe())
x_array = np.array(df[[column]])
if method== 'norm': #Scale transformation
x_scaled = preprocessing.normalize(x_array, norm= 'l2')
dfnorm['x_transformed'] = pd.DataFrame(x_scaled)
if method== 'log': #Log transformation'
dfnorm['x_transformed'] = log(df[column])
#plt.hist(dfnorm['log'])
if method== 'sqt': #Square root transformation
dfnorm['x_transformed'] = np.square(df[column])
if method== 'boxcox': #Box Cox transformatio
xt = stats.boxcox(df[column], lmbda=0)
dfnorm['x_transformed']= xt
if method== 'minmax': #minmax transformation
# Create a minimum and maximum processor object
min_max_scaler = preprocessing.MinMaxScaler()
# Create an object to transform the data to fit minmax processor
x_scaled = min_max_scaler.fit_transform(x_array)
# Run the normalizer on the dataframe
dfnorm['x_transformed'] = pd.DataFrame(x_scaled)
dfnorm['x_transformed']= dfnorm['x_transformed'].fillna(dfnorm['x_transformed'].mean())
if method== 'power' :
pt= preprocessing.PowerTransformer(method= 'box-cox',standardize=False)
dfnorm['x_transformed']= pt.fit_transform(x_array)
if method== 'scale':
x_scaled = preprocessing.scale(x_array)
dfnorm['x_transformed'] = pd.DataFrame(x_scaled)
print(dfnorm['x_transformed'].describe())
fig = plt.figure()
ax1 = fig.add_subplot(321)
ax2 = fig.add_subplot(322)
ax3 = fig.add_subplot(323)
ax4 = fig.add_subplot(324)
ax1.hist(dfnorm['x_original'])
ax1.set_title= ('Histogram before {} transformation for {}'.format(method, column))
ax2.hist(dfnorm['x_transformed'])
ax2.set_title= ('Histogram after {} transformation for {}'.format(method, column))
prob2 = stats.probplot(dfnorm['x_transformed'], dist=stats.norm, plot=ax3)
ax3.set_title('Probplot after {} transformation'.format(method))
ax4.set_title('BoxPlot')
red_square = dict(markerfacecolor='r', marker='s')
ax4.boxplot(dfnorm['x_transformed'], vert=False, flierprops=red_square)
plt.subplots_adjust(top=0.92, bottom=0.08, left=0.10, right=0.95, hspace=0.6, wspace=0.35)
plt.show()
return dfnorm
def nomality_tests(df, column, alpha= 0.05):
'''Test normality using D'Angostino & Pearson, Sahpiro, Anderson-Darling
'''
x= df[column]
stat, p = normaltest(x) #D'Angostino & Pearson test
print(' D Angostino = {:.3f} pvalue = {:.4f}'.format(stat, p))
if p > alpha:
print(' data looks normal (fail to reject H0)')
else:
print(' data does not look normal (reject H0)')
if len(x) < 5000: #Shapiro test is reliable with less than 5K records
stat, p = shapiro(x)
print(' Shapiro = {:.3f} pvalue = {:.4f}'.format(stat, p))
if p > alpha:
print(' data looks normal (fail to reject H0)')
else:
print(' data does not look normal (reject H0)')
stat = anderson(x, dist='norm')
print(' Anderson = {:.3f} '.format(stat.statistic))
for i in range(len(stat.critical_values)):
sl, cv = stat.significance_level[i], stat.critical_values[i]
if stat.statistic < stat.critical_values[i]:
print(' {:.3f}: {:.3f}, data looks normal (fail to reject H0)'.format(sl, cv))
else:
print(' {:.3f}: {:.3f}, data does not look normal (reject H0)'.format(sl, cv))
print(' SL: {} cv = {}'.format(sl, cv))
def outliers_iqr(df, column, output= 'x_outlier'):
'''Interquartile range method to detect outliers
return a df with column for outlier default name x_outlier
'''
quartile_1, quartile_3 = np.percentile(df[column], [25, 75])
iqr = quartile_3 - quartile_1
lower_bound = quartile_1 - (iqr * 1.5)
upper_bound = quartile_3 + (iqr * 1.5)
df[output] = np.where((df[column] > upper_bound) | (df[column] < lower_bound),1,0)
fig = plt.figure()
ax1 = fig.add_subplot(321)
ax2 = fig.add_subplot(322)
red_square = dict(markerfacecolor='r', marker='s')
ax1.boxplot(df[column], vert=False, flierprops=red_square)
ax1.set_title('{} Before'.format(column))
ax2.boxplot(df[output], vert=False, flierprops=red_square)
ax2.set_title('{} After'.format(column))
plt.subplots_adjust(top=0.92, bottom=0.08, left=0.10, right=0.95, hspace=0.6, wspace=0.35)
plt.show()
return df
# -
# ### File Locations
# +
originalpath = (os.getcwd())
print(originalpath)
os.chdir(originalpath)
#os.chdir('..')
path = os.getcwd()
print(path)
today = datetime.datetime.today()
directory_name= '{{cookiecutter.directory_name}}'
report_file= os.path.join(path, 'reports',directory_name + '_report{:%m%d%y}.xlsx').format(today)
figures_path= os.path.join(path, 'reports','figures')
datefile= input('Date of file (MMDDYY: ')
fileoriginaltickets = os.path.join(path, 'data','processed', directory_name + '_tickets' + datefile + '.pkl')
fileoriginalapps = os.path.join(path, 'data','processed', directory_name + '_apps' + datefile + '.pkl')
# -
# ### Read pkl files
# +
#Read PKL files
df2 = pd.read_pickle(fileoriginaltickets,'gzip')
dfreadfile = df2.copy()
df3 = pd.read_pickle(fileoriginalapps,'gzip')
dfreadfileapps = df3.copy()
print('tickets: {}'.format(len(dfreadfile)))
print('Apps: {}'.format(len(dfreadfileapps)))
# -
# ### Perform Data Analysis
# #### Group dataset tickets
grouped, grouped1, grouped2 = group_by(dfreadfile)
# #### 80/20 analysis
#get 80/20 table based in threshold could be cum_pc or AVG3M
threshold = int(input("Enter threshold : [80]") or '81')
basedin = input('Based analysis in [cum_pc] or avg last 3 months [AVG3M] :') or ('cum_pc')
column= input('Column to use [cmdb_ci_name]: ') or ('cmdb_ci_name')
crostab= pareto(dfreadfile, column, 'x_yearmonth', 'All','201812', '201811','201810',threshold)
crostab[crostab[basedin] < threshold]
# +
#pareto graph
ct= crostab[crostab[basedin] < threshold]
fig, ax = plt.subplots()
ax.bar(ct.index, ct.All, color="C0")
plt.xticks(ct.index, rotation='vertical', size=6)
ax2 = ax.twinx()
ax2.plot(ct.index, ct.cum_pc, color="C2", marker=",", ms=5)
ax2.yaxis.set_major_formatter(PercentFormatter())
ax.set_title('Pareto {}'.format(column))
ax.tick_params(axis="y", colors="C0")
ax2.tick_params(axis="y", colors="C2")
#plt.xticks(ct.index, rotation='vertical')
plt.show()
# -
# #### Distribution in the day
#distribution in the day
gethrdistribution(dfreadfile, 'x_bins_day', 'number', 'Distribution in a day',0,'Distday')
# #### Distribution by type of tickets
#types of tickets
gethrdistribution(dfreadfile, 'Type', 'number', 'Types of tickets',10, 'typetks')
# #### Bar chart tickets per vendor
barchart(dfreadfile,'number','x_vendor','Total Tickets', 'Tickets', 'Organization', 'org_tkts_bch')
# #### Aging analysis
filter_in= ['Incident','Requested Item','Change']
histogram(dfreadfile, 'x_agingdays', 'Agging Tickets', 'Aging in Days', 'Tickets', filter_in, 'agingtkts')
# #### Productivity
#productivity
print('Productivity= rate of output (tickets) per unit of input (hrs FTE)')
sumprod= dfreadfile.groupby('x_vendor').agg({'number':'count',
'closed_by_name':'nunique'}).sort_values('number',
ascending=False)
sumprod['Productivity']= sumprod['number'] / (sumprod['closed_by_name'] * 2000)
sumprod['Tickets_per_month']= sumprod['number'] / 12 / sumprod['closed_by_name']
#sumnuc1['Productivity vs effort']= sumnuc1['number'] / sumnuc1['cycletime']
sumprod.rename(columns = {'closed_by_name':'Unique Solvers','number':'Tickets'}, inplace= True)
sumprod = sumprod[sumprod["Tickets"] > 0]
sumprod.index.rename('Org', inplace= True)
sumprod
# #### Type distribution continues variables (cycletime, agging)
verify_normality(dfreadfile, 'x_cycletime')
verify_normality(dfreadfile, 'x_agingdays')
# #### Normality Test
nomality_tests(dfreadfile, 'x_cycletime')
# #### Transform continues variables (cycletime, agging)
dftrans= transform(dfreadfile, 'x_cycletime','power')
nomality_tests(dftrans, 'x_transformed')
# #### Outliers
# +
#if not transformed, run outliers over original df
dftrans= outliers_iqr(dftrans, 'x_transformed')
print('outliers {}'.format(dftrans.x_outlier.sum()))
dftrans= outliers_iqr(dftrans, 'x_original', 'x_outlier2')
print('outliers2 {}'.format(dftrans.x_outlier2.sum()))
# -
#merge outliers in original df, if transformed/normalized
dfreadfile= pd.merge(dfreadfile, dftrans[['x_outlier']], right_index=True, left_index=True)
# #### Area under the curve
# +
mu = dfreadfile.x_cycletime.mean()
sigma = dfreadfile.x_cycletime.std()
x1 = .25 #lower limit 2 hrs
x2 = 8 #upper limit 4 days
# calculate probability
# probability from Z=0 to lower bound
double_prob = erf( (x1-mu) / (sigma*sqrt(2)) )
p_lower = double_prob/2
print('\n Lower Bound: {}'.format(round(p_lower,4)))
# probability from Z=0 to upper bound
double_prob = erf( (x2-mu) / (sigma*sqrt(2)) )
p_upper = double_prob/2
print(' Upper Bound: {}'.format(round(p_upper,4)))
# print the results
Pin = (p_upper) - (p_lower)
print('\n')
print('mean = {} std dev = {} \n'.format(mu, sigma))
print('Calculating the probability of occurring between {} <--> {} days\n'.format(x1, x2))
print('inside interval Pin = {}%'.format(round(Pin*100,1)))
print('outside interval Pout = {}% \n'.format(round((1-Pin)*100,1)))
print('\n')
# calculate the z-transform
z1 = ( x1 - mu ) / sigma
z2 = ( x2 - mu ) / sigma
x = np.arange(z1, z2, 0.001) # range of x in spec
x_all = np.arange(-10, 10, 0.001) # entire range of x, both in and out of spec
# mean = 0, stddev = 1, since Z-transform was calculated
y = norm.pdf(x,0,1)
y2 = norm.pdf(x_all,0,1)
# build the plot
fig, ax = plt.subplots(figsize=(9,6))
plt.style.use('fivethirtyeight')
ax.plot(x_all,y2)
ax.fill_between(x,y,0, alpha=0.3, color='b')
ax.fill_between(x_all,y2,0, alpha=0.1)
ax.set_xlim([-4,4])
ax.set_xlabel('# of Standard Deviations Outside the Mean')
ax.set_yticklabels([])
ax.set_title('Probability to comply')
plt.savefig('normal_curve.png', dpi=72, bbox_inches='tight')
plt.show()
# -
# ### Save Excel file into reports directory
#
# Save an Excel file with intermediate results into the report directory
writer = ExcelWriter(report_file,options={'strings_to_urls': False})
dfreadfile.to_excel(writer, sheet_name='Tickets')
grouped.to_excel(writer, sheet_name='G_by_day_hr_CI')
grouped1.to_excel(writer, sheet_name='G_by_month_CI')
grouped2.to_excel(writer, sheet_name='G_by_month_PL')
dfreadfileapps.to_excel(writer, sheet_name= 'apps')
writer.save()
| {{cookiecutter.directory_name}}/2-EDA_{{cookiecutter.directory_name}}.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Import Resources
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import MinMaxScaler
# %matplotlib inline
train_cnn = pd.read_csv("data/training.csv")
# ### Explore the data
train_cnn.head(1)
train_cnn.shape
# ### Visualize a subset of the data
# +
def string2image(string):
"""Converts a string to a numpy array."""
return np.array([int(item) for item in string.split()]).reshape((96, 96))
def plot_faces(nrows=5, ncols=5):
"""Randomly displays some faces from the training data."""
selection = np.random.choice(train_cnn.index, size=(nrows*ncols), replace=False)
image_strings = train_cnn.loc[selection]['Image']
fig, axes = plt.subplots(figsize=(10, 10), nrows=nrows, ncols=ncols)
for string, ax in zip(image_strings, axes.ravel()):
ax.imshow(string2image(string), cmap='gray')
ax.axis('off')
# -
plot_faces()
# Let's now add to that plot the facial keypoints that were tagged. First, let's do an example :
keypoint_cols = list(train_cnn.columns)[:-1]
xy = train_cnn.iloc[0][keypoint_cols].values.reshape((15, 2))
xy
plt.plot(xy[:, 0], xy[:, 1], 'ro')
plt.imshow(string2image(train_cnn.iloc[0]['Image']), cmap='gray')
# ## Plot faces with keypoints
def plot_faces_with_keypoints(nrows=5, ncols=5):
"""Randomly displays some faces from the training data with their keypoints."""
selection = np.random.choice(train_cnn.index, size=(nrows*ncols), replace=False)
image_strings = train_cnn.loc[selection]['Image']
keypoint_cols = list(train_cnn.columns)[:-1]
keypoints = train_cnn.loc[selection][keypoint_cols]
fig, axes = plt.subplots(figsize=(10, 10), nrows=nrows, ncols=ncols)
for string, (iloc, keypoint), ax in zip(image_strings, keypoints.iterrows(), axes.ravel()):
xy = keypoint.values.reshape((15, 2))
ax.imshow(string2image(string), cmap='gray')
ax.plot(xy[:, 0], xy[:, 1], 'ro')
ax.axis('off')
plot_faces_with_keypoints()
# We can make several observations from this image:
#
# - some images are high resolution, some are low
#
# - some images have all 15 keypoints, while some have only a few
# ## Some stat
train_cnn.describe().loc['count'].plot.bar()
# What this plot tells us is that in this dataset, only 2000 images are "high quality" with all keypoints, while 5000 other images are "low quality" with only 4 keypoints labelled.
# ## Preprocess Data
fully_annotated = train_cnn.dropna()
fully_annotated.shape
X = np.stack([string2image(string) for string in fully_annotated['Image']]).astype(np.float)[:, :, :, np.newaxis]
y = np.vstack(fully_annotated[fully_annotated.columns[:-1]].values)
X.shape, X.dtype
y.shape, y.dtype
X_train = X / 255.
# +
output_pipe = make_pipeline(
MinMaxScaler(feature_range=(-1, 1))
)
y_train = output_pipe.fit_transform(y)
# -
test_cnn = pd.read_csv("data/test.csv")
fully_annotated_test = test_cnn.dropna()
fully_annotated_test.shape
X_test = np.stack([string2image(string) for string in fully_annotated_test['Image']]).astype(np.float)[:, :, :, np.newaxis]
y_test = np.vstack(fully_annotated_test[fully_annotated_test.columns[:-1]].values)
X_test.shape, X_test.dtype
y_test.shape, y_test.dtype
X_test = X_test / 255.
y_test = output_pipe.fit_transform(y_test)
| computer_vision-master/preprocess_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="ezA9vUJudpfE"
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
# %matplotlib inline
# + colab={"base_uri": "https://localhost:8080/"} id="1PzP9-SOshOI" outputId="7097df8f-7ae3-4557-f0a1-f3e05ff3c09b"
# Create images with random rectangles and bounding boxes.
num_imgs = 50000
img_size = 8
min_object_size = 1
max_object_size = 4
num_objects = 1
bboxes = np.zeros((num_imgs, num_objects, 4))
imgs = np.zeros((num_imgs, img_size, img_size)) # set background to 0
for i_img in range(num_imgs):
for i_object in range(num_objects):
w, h = np.random.randint(min_object_size, max_object_size, size=2)
x = np.random.randint(0, img_size - w)
y = np.random.randint(0, img_size - h)
imgs[i_img, x:x+w, y:y+h] = 1. # set rectangle to 1
bboxes[i_img, i_object] = [x, y, w, h]
imgs.shape, bboxes.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 269} id="Do5UwyuWsueY" outputId="cc92f215-ae55-441c-9a10-81c0bb45c863"
i =5
plt.imshow(imgs[i].T, cmap='Greys', interpolation='none', origin='lower', extent=[0, img_size, 0, img_size])
for bbox in bboxes[i]:
plt.gca().add_patch(matplotlib.patches.Rectangle((bbox[0], bbox[1]), bbox[2], bbox[3], ec='r', fc='none'))
# + colab={"base_uri": "https://localhost:8080/"} id="HqiNhxOds-jT" outputId="51c22fd6-cac4-46bc-b23d-02ff056c9727"
# Reshape and normalize the image data to mean 0 and std 1.
X = (imgs.reshape(num_imgs, -1) - np.mean(imgs)) / np.std(imgs)
X.shape, np.mean(X), np.std(X)
# + colab={"base_uri": "https://localhost:8080/"} id="de2SRPKmtFo7" outputId="c985743b-b167-4c8e-af70-6d6e4bd400b7"
# Normalize x, y, w, h by img_size, so that all values are between 0 and 1.
# Important: Do not shift to negative values (e.g. by setting to mean 0), because the IOU calculation needs positive w and h.
y = bboxes.reshape(num_imgs, -1) / img_size
y.shape, np.mean(y), np.std(y)
# + id="6_QB3RnotFeM"
# Split training and test.
i = int(0.8 * num_imgs)
train_X = X[:i]
test_X = X[i:]
train_y = y[:i]
test_y = y[i:]
test_imgs = imgs[i:]
test_bboxes = bboxes[i:]
# + id="NWTMPDCrtcvx"
from keras.models import Sequential
from keras.layers import Dense, Activation, Dropout, Convolution2D, MaxPooling2D, Flatten
from keras.optimizers import SGD
filter_size = 3
pool_size = 2
model = Sequential([
Dense(512, input_dim=X.shape[-1]),
Activation('relu'),
Dense(128, input_dim=X.shape[-1]),
Activation('relu'),
Dropout(0.2),
Dense(y.shape[-1])
])
model.compile('adadelta', 'mse')
# + colab={"base_uri": "https://localhost:8080/"} id="D9amJw0RuMmT" outputId="71bfb868-7064-49bf-b088-6f45a9494c1c"
model.fit(train_X, train_y, epochs=30, validation_data=(test_X, test_y), verbose=2)
# + colab={"base_uri": "https://localhost:8080/"} id="8FjV7Q1cudYm" outputId="0b0c1199-6bd6-41ca-b077-f4f2ee5968c8"
# Predict bounding boxes on the test images.
pred_y = model.predict(test_X)
pred_bboxes = pred_y * img_size
pred_bboxes = pred_bboxes.reshape(len(pred_bboxes), num_objects, -1)
pred_bboxes.shape
# + id="fcZBH-9dunfd"
def IOU(bbox1, bbox2):
'''Calculate overlap between two bounding boxes [x, y, w, h] as the area of intersection over the area of unity'''
x1, y1, w1, h1 = bbox1[0], bbox1[1], bbox1[2], bbox1[3]
x2, y2, w2, h2 = bbox2[0], bbox2[1], bbox2[2], bbox2[3]
w_I = min(x1 + w1, x2 + w2) - max(x1, x2)
h_I = min(y1 + h1, y2 + h2) - max(y1, y2)
if w_I <= 0 or h_I <= 0: # no overlap
return 0.
I = w_I * h_I
U = w1 * h1 + w2 * h2 - I
return(I / U)
# + colab={"base_uri": "https://localhost:8080/", "height": 197} id="BErj61VFvjmB" outputId="74d5a763-cba7-4bfa-f14c-fc7573e825f8"
plt.figure(figsize=(12, 3))
for i_subplot in range(1, 5):
plt.subplot(1, 4, i_subplot)
i = np.random.randint(len(test_imgs))
plt.imshow(test_imgs[i].T, cmap='Greys', interpolation='none', origin='lower', extent=[0, img_size, 0, img_size])
for pred_bbox, exp_bbox in zip(pred_bboxes[i], test_bboxes[i]):
plt.gca().add_patch(matplotlib.patches.Rectangle((pred_bbox[0], pred_bbox[1]), pred_bbox[2], pred_bbox[3], ec='r', fc='none'))
plt.annotate('IOU: {:.2f}'.format(IOU(pred_bbox, exp_bbox)), (pred_bbox[0], pred_bbox[1]+pred_bbox[3]+0.2), color='r')
# + colab={"base_uri": "https://localhost:8080/"} id="wJYdR-W6wI9A" outputId="3b29974b-bb85-4f0b-96a6-423d80bfe5c1"
# Calculate the mean IOU (overlap) between the predicted and expected bounding boxes on the test dataset.
summed_IOU = 0.
for pred_bbox, test_bbox in zip(pred_bboxes.reshape(-1, 4), test_bboxes.reshape(-1, 4)):
summed_IOU += IOU(pred_bbox, test_bbox)
mean_IOU = summed_IOU / len(pred_bboxes)
mean_IOU
| assets/EMSE6575/Object_detection_single_rectangle.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sos
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SoS
# language: sos
# name: sos
# ---
# + [markdown] kernel="SoS"
# # A new approach for EEG feature extraction in P300-based lie detection
#
# <p style="color:rgb(47,79,79);font-size:1.9vh;"> <i><NAME></i><sup>a,b,d</sup>| <i><NAME></i> <sup>b</sup>| <i><NAME></i> <sup>c,d</sup></p>
#
#
# <sup>a</sup> Electrical Engineering Department, Yazd University, Yazd, Iran
# <br>
# <sup>b</sup> Biomedical Engineering Faculty, Amirkabir University of Technology, Tehran, Iran
# <br>
# <sup>c</sup> Islamic Azad University, Mashhad Branch, Iran
# <br>
# <sup>c</sup> Research Center of Intelligent Signal Processing, Tehran, Iran
#
# 
#
# https://doi.org/10.1016/j.cmpb.2008.10.001
#
# ## Abstract
# P300-based Guilty Knowledge Test (GKT) has been suggested as an alternative approach for conventional polygraphy.
#
# The purpose of this study was to extend a previously introduced pattern recognition method for the ERP assessment
# in this application. This extension was done by the further extending the feature set and also employing
# a method for the selection of optimal features.
#
# For the evaluation of the method, several subjects went
# through the designed "Guilty Knowledge Test" paradigm and their respective brain signals were recorded. Next, a P300 detection
# approach based on some features and a statistical classifier was implemented.
# The optimal feature set was
# selected using a genetic algorithm from a primary feature set including some morphological, frequency and
# wavelet features and was used for the classification of the data. The rates of correct detection in guilty
# and innocent subjects were 86%, which was better than other previously used methods.
# + [markdown] kernel="SoS"
# ## Feature Extraction
#
# The following features were described by the original paper, but they were not implemented. I offer an implementation of them in Python, as well as mathematical definitions for each.
#
# #### Morphological Features
# ###### Latency (LAT, $t_{S_{max}}$)
# the ERP’s latency time, i.e. the time
# where the maximum signal value appears:
#
# $ t_{S_{max}} = {t|s(t) = s_{max}} $
#
#
# ###### Amplitude (AMP, $S_{max}$)
# —the maximum signal value:
#
# $ s_{max} = max\{s(t)\} $
#
# ###### Latency/Amplitude ratio (LAR, $t_{S_{max}}/S_{max}$)
#
#
# ###### Absolute amplitude (AAMP, |$S_{max}$|)
#
#
#
# ###### Absolute latency/amplitude ratio (ALAR, |$t_{s_{max}}/s_{max}$|)
#
# ###### Positive area (PAR, $A_{p}$)
# The sum of positive signal values:
#
# $ A_{p} = \sum_{t=400ms}^{800ms} 0.5(s(t) + |s(t)|) $
# ###### Negative area
# The sum of negative signal values:
#
# $ A_{p} = \sum_{t=400ms}^{800ms} 0.5(s(t) - |s(t)|) $
# ###### Total area (TAR, $A_{pn}$)
# $A_{pn} = A_{p} + A_{n}$
# ###### Total absolute area (TAAR, $A_{p|n|}$)
# $A_{pn} = A_{p} + |A_{n}|$
# ###### Average absolute signal slope (AASS, $\dot{s}$)
# $ \dot{s} = \frac{1}{n} \sum_{t=400ms}^{800ms-r} \frac{1}{\tau}(s(t+\tau) - |s(t)|) $
#
# Where the $\tau$ is the sampling interval of the signal, n the number of samples of the digital signal, and s(t) the signal value of the tth sample.
# ###### Peak-to-peak (PP, pp)
# $pp = s_{max} - s_{min}$
# Where s_{max} and s_{min} are the maximum and the minimum signal values, respectively.
# ###### Peak-to-peak time window (PPT, $t_{pp}$)
# $t_{pp} = t_{s_{max}} - t_{s_{min}}$
# ###### Peak-to peak slope (PPS, $\dot{s}_{pp}$)
# $\dot{s}_{pp} = \frac{pp}{t_{pp}}$
# ###### Zero crossings (ZC, $n_{ZC}$)
# the number of times t that s(t)=0, in peak-to-peak time window:
#
# $ n_{ZC} = \sum_{t=t_{s_{min}}}^{t_{s_{max}}} \delta_{s} $
#
# where \delta_{s} = 1 if s(t)=0, 0 otherwise
#
#
# ###### Zero crossings density (ZCD, d_{zc})
# Zero crossings per time unit, in peak-to-peak time window:
#
# $ d_{ZC} = \frac{n_{ZC}}{t_{pp}} $
#
# where n_{ZC} are the zero crossings and t_{pp} is the peak-to-peak time window.
# + kernel="Python 3"
# %use 'Python 3'
import numpy as np
def signal_slope_f(row):
total = 0.0
for idx, val in enumerate(row[:-1]):
total += np.abs(row[idx+1] - val)
total /= len(row)
return total
def latency(row):
return row.argmax()
def positive_area(row):
return row[row>0].sum()
def negative_area(row):
return row[row<0].sum()
def zero_crossing(row):
return np.where(np.diff(np.sign(row[row.argmin():row.argmax()])))[0].size
# + [markdown] kernel="SoS"
# #### Frequency features
#
# ###### Mode frequency:
#
# $f_{mode}$ is the frequency with the most energy content in the
# signal spectrum, so the maximum amplitude in the power
# spectrum density of the signal is at this frequency:
#
# $S(f_{mode}) = Max_f\{S(f)\}$
# S is the power spectral density of signal and f is frequency.
#
# ###### Median frequency:
# Median frequency ($f_{median}$) separates the power spectrum
# into two equal energy areas and is calculated from the
# following equation:
#
# $$ \int_{0}^{f{median}} S(f) df = \int_{f{median}}^{\infty} S(f) df $$
#
# + kernel="Python 3"
# %use 'Python 3'
import pandas as pd
from scipy import signal
from numpy import trapz
def mode_freq_f(row):
test_signal = pd.to_numeric(row)
freqs, psd = signal.welch(test_signal.values, fs = 250)
return 10**(freqs[np.argmax(psd)])
def median_freq_f(row):
test_signal = pd.to_numeric(row)
freqs, psd = signal.welch(test_signal.values, fs = 250)
total_area = trapz(psd,freqs)
half_area = total_area/2
integral_values = np.array([abs(trapz(psd[:i],freqs[:i]) - half_area) for i in range(len(freqs))])
return freqs[np.argmin(integral_values)]
# + [markdown] kernel="SoS"
# ## Learn more
#
# + kernel="Python 3"
# %use 'Python 3'
from IPython.display import YouTubeVideo
YouTubeVideo('N2H4NTAZrhY', width=800, height=600)
# + kernel="Python 3"
# %use 'Python 3'
YouTubeVideo('nZEFgC0oHpA', width=800, height=600)
# + [markdown] kernel="SoS"
# ## Втор колоквиум промени Script of Scripts
#
# Ќе генерираме array од 1000 елементи (семплирани од гаусова дистрибуција) во R, ќе ги земеме во Python и ќе плотнеме со Plotly.
# + kernel="R"
y <- rnorm(1000)
write.csv(y, 'data.csv')
# + kernel="R"
y
# + kernel="Python 3"
# %use 'Python 3'
import scipy
import plotly.figure_factory as ff
import numpy as np
import pandas as pd
x = np.random.randn(200) - 1
y = pd.read_csv('data.csv')['x'].to_numpy()
hist_data = [x, y]
group_labels = ['Python randn numbers', 'R rnorm numbers']
colors = ['#333F44', '#37AA9C']
# Create distplot with curve_type set to 'normal'
fig = ff.create_distplot(hist_data, group_labels, show_hist=False, colors=colors)
# Add title
fig.update_layout(title_text='R and Python arrays')
fig.show()
# + kernel="Python 3"
# %use 'Python 3'
# + kernel="Python 3"
| kolokvium2_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Prep Stuff
drive_path = 'd:/'
import numpy as np
import pandas as pd
import os
import sys
import seaborn
import matplotlib.pyplot as plt
# %matplotlib inline
from scipy.stats import variation
# +
from allensdk.core.brain_observatory_cache import BrainObservatoryCache
from allensdk.brain_observatory.stimulus_analysis import StimulusAnalysis
from allensdk.brain_observatory.findlevel import findlevel
manifest_path = os.path.join(drive_path,'BrainObservatory','manifest.json')
boc = BrainObservatoryCache(manifest_file=manifest_path)
# -
#only get the sessions that contain the natural scenes: 79 sessions
expMetaData = pd.DataFrame(boc.get_ophys_experiments())
SessionThreeMD = expMetaData[expMetaData.session_type=='three_session_B']
SessionThreeMD.head()
for i, expt_id in enumerate(SessionThreeMD['id']):
if i == 0:
df = pd.read_csv("%d_name" % expt_id)
else:
df.append(read_csv...)
# +
#names = FinalDF.columns.tolist()
#names[names[0]] = 'Cell_Specimen_ID'
#FinalDF.columns=names
#FinalDF.head()
| COV_Composite.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (Prod Env)
# language: python
# name: prod
# ---
import petl as etl
from targetprocess.api import TargetProcessAPIClient
from targetprocess.serializers import TargetProcessSerializer
tp = TargetProcessAPIClient(api_url='https://name.tpondemand.com/api/v1/', user='<EMAIL>', password='<PASSWORD>')
request_json = tp.get_collection(collection="Release", take = 1000,include="[Name,StartDate,EndDate,Project]" )
ListJson = TargetProcessSerializer.deserialize(request_json)
Rtable1 = etl.fromdicts(ListJson, header=['Name','StartDate', 'EndDate','Project'])
Rtable2 = etl.addfield(Rtable1, 'Type', lambda row: row['Project']['Name'])
Rtable3 = etl.cutout(Rtable2, 'Project')
Rtable4 = etl.select(Rtable3,lambda rec: rec.Type != 'Flipbox' and rec.Name != 'Priority Backlog')
RList = etl.dicts(Rtable4)
ReleaseList = list(RList)
request_json2 = tp.get_collection(collection="Requests", take = 10000000,include="[Id,Name,RequestType,CreateDate]")
ListJson2 = TargetProcessSerializer.deserialize(request_json2)
Rqtable1 = etl.fromdicts(ListJson2, header=['Id','CreateDate', 'RequestType', 'Name'])
Rqtable2 = etl.addfield(Rqtable1, 'Type', lambda row: row['RequestType']['Name'])
Rqtable3 = etl.cut(Rqtable2, 'Id', 'CreateDate','Name','Type')
Rqtable4 = etl.select(Rqtable3,lambda rec: rec.Type == 'Ошибка')
table5 = etl.addfield(Rqtable4, 'Release', lambda row: [ r['Name'] for r in ReleaseList if (r['StartDate']<row['CreateDate']<r['EndDate'])])
table6 = etl.convert(table5,'Release', lambda v:v[0] if v==v else '' )
etl.toxlsx(table6,'BugListWithRelease.xlsx')
| examples/Targetprocess/TP_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from math import factorial
def combinations(n, k):
return int(factorial(n) / (factorial(k) * factorial(n - k)))
# +
# 1. Даны значения зарплат из выборки выпускников:
# 100, 80, 75, 77, 89, 33, 45, 25, 65, 17, 30, 24, 57, 55, 70, 75, 65, 84, 90, 150.
# Посчитать (желательно без использования статистических методов наподобие std, var, mean) среднее арифметическое,
# среднее квадратичное отклонение, смещенную и несмещенную оценки дисперсий для данной выборки.
salary = [100, 80, 75, 77, 89, 33, 45, 25, 65, 17, 30, 24, 57, 55, 70, 75, 65, 84, 90, 150]
print(f'среднее арифметическое: {sum(salary)/len(salary)}')
print(f'среднее квадратичное отклонение: смещенное = {np.std(salary)}, несмещенное = {np.std(salary, ddof=1)}')
print(f'Оценка дисперсий: смещенная = {np.var(salary)}, несмещенная = {np.var(salary, ddof=1)}')
# +
# 2. В первом ящике находится 8 мячей, из которых 5 - белые.
# Во втором ящике - 12 мячей, из которых 5 белых.
# Из первого ящика вытаскивают случайным образом два мяча, из второго - 4.
# Какова вероятность того, что 3 мяча белые?
P1 = combinations(5, 2)/combinations(8, 2)*(combinations(5, 1)*combinations(7, 3))/combinations(12, 4) # 2 из первой и 1 из второй
P2 = (combinations(5, 1)*combinations(3, 1))/combinations(8, 2)*(combinations(5, 2)*combinations(7, 2))/combinations(12, 4) # 1 из первой и 2 из второй
P3 = combinations(3, 2)/combinations(8, 2)*(combinations(5, 3)*combinations(7, 1))/combinations(12, 4) # 0 из первой и 3 из второй
P1+P2+P3
# +
# 3. На соревновании по биатлону один из трех спортсменов стреляет и попадает в мишень.
# Вероятность попадания для первого спортсмена равна 0.9, для второго — 0.8, для третьего — 0.6.
# Найти вероятность того, что выстрел произведен: a). первым спортсменом б). вторым спортсменом в). третьим спортсменом.
P = 1/3*0.9 + 1/3*0.8 + 1/3*0.6
print(f'первым спортсменом: {(1/3*0.9)/P}')
print(f'вторым спортсменом: {(1/3*0.8)/P}')
print(f'третьим спортсменом: {(1/3*0.6)/P}')
# +
# 4. В университет на факультеты A и B поступило равное количество студентов,
# а на факультет C студентов поступило столько же, сколько на A и B вместе.
# Вероятность того, что студент факультета A сдаст первую сессию, равна 0.8.
# Для студента факультета B эта вероятность равна 0.7, а для студента факультета C - 0.9.
# Студент сдал первую сессию. Какова вероятность, что он учится: a). на факультете A б). на факультете B в). на факультете C?
P = 1/4*0.8+1/4*0.7+1/2*0.9
print(f'на факультете A: {(1/4*0.8)/P}')
print(f'на факультете B: {(1/4*0.7)/P}')
print(f'на факультете C: {(1/2*0.9)/P}')
# +
# 5. Устройство состоит из трех деталей. Для первой детали вероятность выйти из строя в первый месяц равна 0.1,
# для второй - 0.2, для третьей - 0.25. Какова вероятность того, что в первый месяц выйдут из строя:
# а). все детали б). только две детали в). хотя бы одна деталь г). от одной до двух деталей?
print(f'все детали: {0.1*0.2*0.25}')
print(f'только две детали: -')
print(f'хотя бы одна деталь: -')
print(f'от одной до двух деталей: -')
# -
| p_lesson3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
# alpha-mind的data文件夹提供了对于因子数据进行中性化、标准化和去极值化的函数:
# * [neutralize.py](https://github.com/alpha-miner/alpha-mind/blob/master/alphamind/data/neutralize.py):提供了风险中性化函数*neutralize*
# * [standardize.py](https://github.com/alpha-miner/alpha-mind/blob/master/alphamind/data/standardize.py): 提供了标准化函数*standardize*和与sklearn类似的算子*Standardizer*以及*GroupStandardizer*
# * [winsorize.py](https://github.com/alpha-miner/alpha-mind/blob/master/alphamind/data/winsorize.py):提供了去极值化的函数*winsorize_normal*
#
# 标准化和去极值化函数的特别之处在于,他们还可以按组别(对应的实际问题,比如按行业)进行数据处理。
#
# ### 去极值化处理
# #### 全截面处理
#
# alpha-mind提供了*winsorize_normal*这一函数,假设数据服从正太分布,进行去极值化处理。
# +
import numpy as np
import matplotlib.pyplot as plt
from alphamind.data.winsorize import winsorize_normal
# 假设有50只股票,每只股票有1个因子,构成一个矩阵
factors = np.random.rand(50, 1)
# 为了展示方便,取一个标准差为上下界
clean_factors = winsorize_normal(factors, num_stds=1)
# %matplotlib inline
plt.plot(factors)
plt.plot(clean_factors)
# -
# #### 分行业处理
# 有时候需要进行分组去极值化,如按行业来分组。针对这种情况,只需要多传递一个分组参数给*winsorize_normal*。
# +
# 假设所有股票可分为2个组别(如行业),前25个和后25个分属不同类别
industry = np.concatenate([np.array([1.0]*25), np.array([2.0]*25)])
# 此时令前25个因子和后25个因子的标准差不一样
factors = np.concatenate([ 10*np.random.rand(25, 1), np.random.rand(25, 1)])
# 为了展示方便,取一个标准差为上下界
clean_factors = winsorize_normal(factors, num_stds=1, groups=industry)
# %matplotlib inline
plt.plot(factors)
plt.plot(clean_factors)
# -
# ### 标准化处理
# 标准化函数的使用与去极值化类似,故不再重复,下文仅展示分组进行标准化处理的例子。
# +
from alphamind.data.standardize import standardize
# 假设所有股票可分为2个组别(如行业),前25个和后25个分属不同类别
industry = np.concatenate([np.array([1.0]*25), np.array([2.0]*25)])
# 此时令前25个因子和后25个因子的标准差不一样
factors = np.concatenate([ 10*np.random.rand(25, 1), np.random.rand(25, 1)])
# 为了展示方便,取一个标准差为上下界
clean_factors = standardize(factors, groups=industry)
# %matplotlib inline
plt.plot(factors)
plt.plot(clean_factors)
# -
# 另外还提供了类似sklearn风格的算子*Standardizer*和*GroupedStandardizer*,使用*fit,transform*方法可以得到同样的效果。具体请参见sklearn相关的帮助以及本项目代码。
# ### 中性化处理
# #### 风险(风格)中性化
# +
from alphamind.data.neutralize import neutralize
# 假设有30只股票,每只股票有10个因子,构成一个矩阵
raw_factors = np.random.rand(30, 10)
# 假设每只股票对应有4个风险因子,构成一个矩阵
risk_factors = np.random.rand(30, 4)
# 因子的风险中性化,就是原始因子对风险因子求线性回归后的残差
ret_neutralize_1 = neutralize(risk_factors, raw_factors)
# -
# #### 风险以及行业中性化
# 生成环境中一般还需要进行行业中性化,对于*neutralize*来说只需要增加一个入参,把每只股票对应的行业标签以np数组传递进去。
# +
# 假设所有股票可分为3个组别(如行业)
industry = np.random.randint(3, size=30)
# 对因子进行风险和行业中性化
ret_neutralize_2 = neutralize(risk_factors, raw_factors, industry)
# +
# 如果令所有行业都是一样,那么将得到与 ret_neutralize_1一样的结果
industry = np.random.randint(1, size=30)
ret_neutralize_3 = neutralize(risk_factors, raw_factors, industry)
# print(ret_neutralize_1)
# print(ret_neutralize_3)
# -
# #### 仅行业中性
# 如果只想进行行业中性化处理,那么可以令风险因子(*neutralize*的第一个入参)为全部元素为1的矩阵。下面进行一个简单测试。
# +
# 假设有10只股票,每只股票有1个因子
factor = np.array([1.0, 1.0, 1.0, 1.0, 1.0, 2.0, 3.0, 4.0, 5.0, 6.0])
# 假设所有股票可分为2个行业,前5和后5 分属于不同行业
industry = np.array([1.0, 1.0, 1.0, 1.0, 1.0, 2.0, 2.0, 2.0, 2.0, 2.0])
risk_factors = np.ones((10, 1))
ret_neutralize_4 = neutralize(risk_factors, factor, industry)
print(ret_neutralize_4)
# -
# 行业中性化后的因子为原始因子减去行业内平均值,结果与预期一样。
# ### 预处理合成函数
# alpha-mind 还提供了一个预处理函数*factor_processing*, 集成了以上描述函数的所有功能,可以一步进行因子数据的预处理功能。
# - 入参*pre_process*需传递预处理的子函数列表,如去极值化和标准化。如果传递了风险因子的参数,中性化函数会在预处理之后自动触发。
# - 入参*post_process*指的是在中性化处理(如果有的话)或*pre_process*完成之后,还需要进行的处理。同样可以将去极值化和标准化函数作为列表传递进去。
# +
from alphamind.data.processing import factor_processing
from alphamind.data.standardize import standardize
# 假设有30只股票,每只股票有10个因子,构成一个矩阵
raw_factors = np.random.rand(30, 10)
# 假设每只股票对应有4个风险因子,构成一个矩阵
risk_factors = np.random.rand(30, 4)
# 假设所有股票可分为3个组别(如行业)
industry = np.random.randint(3, size=30)
ret_preprocess = factor_processing(raw_factors,
pre_process=[winsorize_normal, standardize],
risk_factors=risk_factors,
groups=industry)
# -
| notebooks/Quick Start 1 - Factor Preprocess.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Pandas III
# + slideshow={"slide_type": "slide"}
# %matplotlib inline
import pandas as pd
import numpy as np
pd.options.display.max_rows = 10
# -
df = pd.read_csv('data/beer_subset.csv.gz', parse_dates=['time'], compression='gzip')
df.head()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Groupby
# + [markdown] slideshow={"slide_type": "fragment"}
# The components of a groupby operation are to
#
# 1. Split a table into groups
# 2. Apply a function to each group
# 3. Combine the results
# + [markdown] slideshow={"slide_type": "fragment"}
# In pandas the first step looks like
#
# ```python
# df.groupby( grouper )
# ```
# + [markdown] slideshow={"slide_type": "fragment"}
# `grouper` can be many things
#
# - Series (or string indicating a column in `df`)
# - function (to be applied on the index)
# - dict : groups by *values*
# - `levels=[ names of levels in a MultiIndex ]`
# + slideshow={"slide_type": "-"}
# -
# + [markdown] slideshow={"slide_type": "fragment"}
# Haven't really done anything yet. Just some book-keeping to figure out which **keys** go with which **rows**. Keys are the things we've grouped by (each `beer_style` in this case).
# + slideshow={"slide_type": "-"}
# + slideshow={"slide_type": "-"}
# + [markdown] slideshow={"slide_type": "subslide"}
# There's a generic aggregation function:
# -
# + [markdown] slideshow={"slide_type": "fragment"}
# Which accepts some common operations as strings:
# + slideshow={"slide_type": "-"}
# + [markdown] slideshow={"slide_type": "fragment"}
# Or functions that can operate on Pandas or Numpy objects:
# + slideshow={"slide_type": "-"}
# + [markdown] slideshow={"slide_type": "fragment"}
# And for many common operations, there are also convenience functions:
# + slideshow={"slide_type": "-"}
# -
# + [markdown] slideshow={"slide_type": "subslide"}
# By default the aggregation functions get applied to all columns, but we can subset:
# -
review_cols = [c for c in df.columns if c[0:6] == 'review']
review_cols
# + [markdown] slideshow={"slide_type": "fragment"}
# `.` attribute lookup works as well.
# -
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example
#
# Find the `beer_styles` with the greatest variance in `abv`:
# -
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Some more complex examples
# + [markdown] slideshow={"slide_type": "fragment"}
# Multiple aggregations on one column
# -
# + [markdown] slideshow={"slide_type": "fragment"}
# Single aggregation on multiple columns
# -
# + [markdown] slideshow={"slide_type": "fragment"}
# Multiple aggregations on multiple columns
# -
# + [markdown] slideshow={"slide_type": "fragment"}
# Hierarchical Indexes in the columns can be awkward to work with, so you can move a level to the Index with `.stack`:
# -
# + [markdown] slideshow={"slide_type": "fragment"}
# You can group by **levels** of a MultiIndex:
# -
# + [markdown] slideshow={"slide_type": "fragment"}
# Group by **multiple** columns
# -
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example
# Find the relationship between `review` length (the text column) and average `review_overall`
# -
# + [markdown] slideshow={"slide_type": "fragment"}
# <div class="alert alert-info">
# <b>Bonus exercise</b>
# </div>
#
# - Try grouping by the number of words
# - Try grouping by the number of sentences
#
# _Hint_: `str.count` accepts a regular expression...
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example
#
# Which **brewer** (`brewer_id`) has the largest gap between the min and max `review_overall` for two of their beers?
# + [markdown] slideshow={"slide_type": "fragment"}
# _Hint_: You'll need to do this in two steps:
# 1. Find the average `review_overall` by `brewer_id` and `beer_name`.
# 2. Find the difference between the max and min by brewer (rembember `.groupby(level=)`)
# -
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Complex Example
#
# Create a more aggregated "kind" of beer, less detailed than `style`
# + slideshow={"slide_type": "fragment"}
kind = df['beer_style'].str.lower()\
.replace({'india pale ale': 'ipa', 'american pale ale': 'apa'})
kind.head()
# + slideshow={"slide_type": "fragment"}
kinds = ['ipa', 'apa', 'amber ale', 'rye', 'scotch', 'stout',
'barleywine', 'porter', 'brown ale', 'lager', 'pilsner',
'tripel', 'bitter', 'farmhouse', 'malt liquour', 'rice']
# -
# + [markdown] slideshow={"slide_type": "fragment"}
# #### Bonus
# We can use [regular expressions](https://docs.python.org/3/library/re.html) to do this in one (convoluted) line...
# -
expr = '|'.join(['(?P<{name}>{pat})'.format(pat=kind, name=kind.replace(' ', '_'))
for kind in kinds])
kind2 = df['beer_style'].str.lower()\
.replace({'india pale ale': 'ipa', 'american pale ale': 'apa'})\
.str.extract(expr).fillna('').sum(1)\
.str.lower().replace('','other')
kind2.head()
# + [markdown] slideshow={"slide_type": "fragment"}
# ### Back to the example...
# -
# Find the highest rate "kind" of beer
# + [markdown] slideshow={"slide_type": "fragment"}
# How are the different kinds of beers rated for each brewer?
# -
# + [markdown] slideshow={"slide_type": "fragment"}
# Find the number of beers of each kind by brewer:
# -
# + [markdown] slideshow={"slide_type": "slide"}
# We've seen a lot of permutations among number of groupers, number of columns to aggregate, and number of aggregators.
#
#
# In fact, the `.agg`, which returns one row per group, is just one kind of way to combine the results. The three ways are
#
# - `agg`: one row per results
# - `transform`: identically shaped output as input
# - `apply`: anything goes
# + [markdown] slideshow={"slide_type": "slide"}
# ### Transform
#
# Combined `Series`/`DataFrame` is the same shape as the input.
# + [markdown] slideshow={"slide_type": "fragment"}
# For example, say you want to standardize the reviews by subtracting the mean.
# -
# + [markdown] slideshow={"slide_type": "fragment"}
# We can do this at the *person* level with `groupby` and `transform`.
# -
# + [markdown] slideshow={"slide_type": "fragment"}
# This uses the *group* means instead of the overall means
# -
df[['profile_name','review_overall','review_overall_demeaned']]\
.sort_values('profile_name').head(10)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Apply
#
# - `.apply()` can return all sorts of things, doesn't have to be the same shape...
# - Lots of uses, too many to go into...
# + slideshow={"slide_type": "fragment"}
def something(x):
return x['review_appearance'].max() - x['review_aroma'].min()
# -
# + [markdown] slideshow={"slide_type": "fragment"}
# Or more succinctly as a `lambda` function:
# -
# + [markdown] slideshow={"slide_type": "slide"}
# ## References
#
# Slide materials inspired by and adapted from [<NAME>](https://github.com/fonnesbeck/statistical-analysis-python-tutorial) and [<NAME>](https://github.com/TomAugspurger/pydata-chi-h2t)
| Lecture 5/Lecture 5 - Pandas III (Template).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# Text and code provided under a Creative Commons Attribution license, CC-BY. (c) <NAME>, <NAME>, 2014. Thanks: NSF for support via CAREER award #1149784.
# -
# ##### Version 0.1 -- February 2014
# # Infinite row of vortices
# The objective of this assignment is to visualize the streamlines around an infinite row of vortices. First, you will consider the case of a finite number of vortices, obtained by simple superposition. By adding more and more vortices, you should be able to see how the flow pattern approaches that of an infinite row of vortices. But there will always be some differences (pay attention to what these may be).
#
# It's possible to derive an analytical expression for the infinite case, and the derivation is provided below. With this analytical expression, you can visualize the streamlines for the infinite case. Observe and think: how are the streamlines different from one case to the other?
#
#
# In this notebook, there is no Python code. Your job is to study the theory (and follow the mathematics on your own handwritten notes), to think how you could implement it in an efficient manner and finally to code it and visualize the results.
# ## Vortex flow (from previous lesson)
# You might not suspect it, but the vortex has a very important role in classical aerodynamic theory. You'll discover some of its uses in this assignment.
#
# First, a little review of the basics. As seen in a previous lesson, a vortex of strength $\Gamma$ has a stream-function:
#
# $$\psi\left(r,\theta\right) = \frac{\Gamma}{2\pi}\ln r$$
#
# and a velocity potential
#
# $$\phi\left(r,\theta\right) = -\frac{\Gamma}{2\pi}\theta$$
#
# We can now derive the velocity components in a polar coordinate system, as follows:
#
# $$u_r\left(r,\theta\right) = 0$$
#
# $$u_\theta\left(r,\theta\right) = -\frac{\Gamma}{2\pi r}$$
# In a Cartesian coordinate system, the velocity components at $\left(x,y\right)$ around a vortex of strength $\Gamma$ located at $\left(x_\text{vortex},y_\text{vortex}\right)$, are given by
#
# $$u\left(x,y\right) = +\frac{\Gamma}{2\pi}\frac{y-y_\text{vortex}}{(x-x_\text{vortex})^2+(y-y_\text{vortex})^2}$$
#
# $$v\left(x,y\right) = -\frac{\Gamma}{2\pi}\frac{x-x_\text{vortex}}{(x-x_\text{vortex})^2+(y-y_\text{vortex})^2}$$
#
# and the stream-function is written as
#
# $$\psi\left(x,y\right) = \frac{\Gamma}{4\pi}\ln\left((x-x_\text{vortex})^2+(y-y_\text{vortex})^2\right)$$
# ## Superposition of many vortices
# In this assignement, we consider a useful example to illustrate the concept of a *vortex sheet*: an infinite row of vortices of equal strength $\Gamma$ (same sign and magnitude) evenly spaced by a distance $a$. But let's start with a finite row of vortices first.
#
#
# The stream-function $\psi_i$ of the $i^{th}$ vortex at a distance $r_i$ is given by:
#
# $$\psi_i = \frac{\Gamma}{2\pi}\ln r_i$$
#
# Applying the principle of superposition, the stream-function of $N$ vortices is, then
#
# $$\psi = \frac{\Gamma}{2\pi} \sum_{i=1}^N \ln r_i$$
#
# And the velocity field (in Cartesian coordinates) of the row of vortices is
#
# $$u\left(x,y\right) = + \frac{\Gamma}{2\pi} \sum_{i=1}^N \frac{y-y_i}{(x-x_i)^2+(y-y_i)^2}$$
#
# $$v\left(x,y\right) = - \frac{\Gamma}{2\pi} \sum_{i=1}^N \frac{x-x_i}{(x-x_i)^2+(y-y_i)^2}$$
#
# where $\left(x_i,y_i\right)$ are the Cartesian coordinates of the $i^{\text{th}}$ vortex.
#
# Here is a diagram of the situation:
#
#
# <center></center>
#
#
# (In the next section, we'll show you how to derive a closed-form expression for the *infinite* sum of vortices. But hang on.)
# ---
# ##### Task:
#
# #### Compute the velocity field and plot the streamlines of a row of vortices.
# You will place $N$ vortices aligned on the horizontal axis and visualize the flow pattern. Do the following:
#
# * Using the equations derived above, compute the velocity components of each vortex on a mesh grid.
#
# * Remember that a finite number of vortices can be represented by a *list* or a NumPy *array*. Think and decide which one to use.
#
# * Define functions to avoid code repetition (and why not classes, if you prefer and are familiar with the concept).
#
# * Once you have all the velocities, apply the principle of superposition and plot the resulting flow pattern.
#
# * Play around with the size of your vortex row and the plotting range. Make your plot of publishable quality!
#
# In the end you should get something similar to this:
#
# <center></center>
# ---
# ## Infinite row of vortices
# This will be a fun mathematical exercise! Follow along, and take your own notes.
#
# To derive the closed-form functions, we use the complex representation:
#
# $$z = x + jy$$
#
# where $j^2=-1$. (We don't use $i$ to not confuse it with our indices!)
#
# The complex potential is defined as $w = \phi + j\psi$, where $\phi$ is the potential velocity and $\psi$ is the stream-function. If we differentiate the complex potential $w$ with respect to the complex variable $z$, we get the complex velocity
#
# $$\frac{dw}{dz} = u-jv$$
#
# where $u$ and $v$ are the Cartesian velocity components.
#
# The complex potential representing a vortex of strength $\Gamma$ located at the origin is
#
# $$w = \frac{j\Gamma}{2\pi}\ln z$$
#
# Why?
#
# Because $z=re^{j\theta}$ and $w$ becomes
#
# $$w = -\frac{\Gamma}{2\pi}\theta + j\frac{\Gamma}{2\pi}\ln r = \phi + j\psi$$
#
# Consider a second vortex located at $\left(a,0\right)$ with the same strength $\Gamma$. Its complex potential is given by
#
# $$w = \frac{j\Gamma}{2\pi}\ln \left(z-a\right)$$
#
# A additional vortex located $a$ further will have a complex potential of the form
#
# $$w = \frac{j\Gamma}{2\pi}\ln \left(z-2a\right)$$
#
# and so on...
#
# Therefore, the complex potential representing an inifinite row of vortices (on the line $y=0$) is given by
#
# $$w = \frac{j\Gamma}{2\pi}\sum_{m=-\infty}^{+\infty} \ln \left( z-ma \right)$$
#
# When we integrate the velocity components, in the previous notebooks, to find the stream-function and the potential, we dropped the constant of integration. Here, we decide to add a constant (not a random one!). Why not, since the differentiation will give the same results.
#
# $$w = \frac{j\Gamma}{2\pi}\sum_{m=-\infty}^{+\infty} \ln \left( z-ma \right) + \text{constant}$$
#
# where
#
# $$\text{constant} = -\frac{j\Gamma}{2\pi} \sum_{m=-\infty, m\neq 0}^{+\infty} \ln \left( -ma \right)$$
#
# so that, the complex potential can be cast in the following form
#
# $$w = \frac{j\Gamma}{2\pi}\sum_{m=-\infty,m\neq 0}^{+\infty} \ln \left( \frac{z-ma}{-ma} \right) + \frac{j\Gamma}{2\pi}\ln z$$
#
# Now, it is time do some mathematics...
#
# $$w = \frac{j\Gamma}{2\pi}\sum_{m=-\infty,m\neq 0}^{+\infty} \ln \left( 1-\frac{z}{ma} \right) + \frac{j\Gamma}{2\pi}\ln z$$
#
# $$w = \frac{j\Gamma}{2\pi}\sum_{m=1}^{+\infty} \left\lbrace \ln\left(1-\frac{z}{ma}\right) + \ln\left(1+\frac{z}{ma}\right) \right\rbrace + \frac{j\Gamma}{2\pi}\ln z$$
#
# $$w = \frac{j\Gamma}{2\pi}\sum_{m=1}^{+\infty} \ln\left(1-\frac{z^2}{m^2a^2}\right) + \frac{j\Gamma}{2\pi}\ln z$$
#
# $$w = \frac{j\Gamma}{2\pi} \ln\left(\prod_{m=1}^{+\infty}\left(1-\frac{z^2}{m^2a^2}\right)\right) + \frac{j\Gamma}{2\pi}\ln z$$
#
# $$w = \frac{j\Gamma}{2\pi} \ln\left(z\prod_{m=1}^{+\infty}\left(1-\frac{z^2}{m^2a^2}\right)\right)$$
#
# $$w = \frac{j\Gamma}{2\pi} \ln\left(z\prod_{m=1}^{+\infty}\left(1-\frac{\left(\frac{z\pi}{a}\right)^2}{m^2\pi^2}\right)\right)$$
#
# The product is actually the formula for the sine and the complex potential becomes
#
# $$w = \frac{j\Gamma}{2\pi}\ln\left(\sin\left(\frac{z\pi}{a}\right)\right)$$
#
# We derive the complex potential with respect to the complex variable to get the complex velocity
#
# $$\frac{dw}{dz} = u-iv = \frac{j\Gamma}{2a}\cot\left(\frac{z\pi}{a}\right)$$
#
# $$u-jv = \frac{j\Gamma}{2a}\frac{\cos\left(\frac{\pi x}{a}+j\frac{\pi y}{a}\right)}{\sin\left(\frac{\pi x}{a}+j\frac{\pi y}{a}\right)}$$
#
# Applying trigonometric identities, we find the following expression
#
# $$u-jv = \frac{j\Gamma}{2a}\frac{\cos\left(\frac{\pi x}{a}\right)\cosh\left(\frac{\pi y}{a}\right) - j\sin\left(\frac{\pi x}{a}\right)\sinh\left(\frac{\pi y}{a}\right)}{\sin\left(\frac{\pi x}{a}\right)\cosh\left(\frac{\pi y}{a}\right) + j\cos\left(\frac{\pi x}{a}\right)\sinh\left(\frac{\pi y}{a}\right)}$$
#
# which can be cast in the form (try it!)
#
# $$u-jv = \frac{\Gamma}{2a}\frac{\sinh\left(\frac{2\pi y}{a}\right)}{\cosh\left(\frac{2\pi y}{a}\right) - \cos\left(\frac{2\pi x}{a}\right)} + j \frac{\Gamma}{2a}\frac{\sin\left(\frac{2\pi x}{a}\right)}{\cosh\left(\frac{2\pi y}{a}\right) - \cos\left(\frac{2\pi x}{a}\right)}$$
#
# Therefore, the Cartesian velocity components of an infinite row of vortices are given by
#
# $$u\left(x,y\right) = +\frac{\Gamma}{2a}\frac{\sinh\left(\frac{2\pi y}{a}\right)}{\cosh\left(\frac{2\pi y}{a}\right)-\cos\left(\frac{2\pi x}{a}\right)}$$
#
# $$v\left(x,y\right) = -\frac{\Gamma}{2a}\frac{\sin\left(\frac{2\pi x}{a}\right)}{\cosh\left(\frac{2\pi y}{a}\right)-\cos\left(\frac{2\pi x}{a}\right)}$$
# ---
# ##### Task:
#
# #### Compute the velocity field and plot the streamlines of an *infinite* row of vortices
# Now that we have derived the functions for the velocity components, implement them in a code cell and plot the streamlines.
#
# *Can you notice the differences with the previous case where the number of vortices was finite?*
#
# Play around with your plotting settings for the *finite* row of vortices, until you can get a plot that looks more like the infinite case. When can you say the finite case is a good approximation of the infinite case?
# #####Think
# Notice that the streamline pattern is parallel to the *vortex sheet*: there is no net normal flow. Therefore, the sheet can model a solid surface in potential flow. We end up with a slip velocity at such an interface: how is this consistent with potential flow?
# ---
# + active=""
# Please ignore the cell below. It just loads our style for the notebook.
# -
from IPython.core.display import HTML
def css_styling():
styles = open('../styles/custom.css', 'r').read()
return HTML(styles)
css_styling()
| lessons/05_Lesson05_InfiniteRowOfVortices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ⊕ [PropBank](http://www.nltk.org/howto/propbank.html)
#
>>> from nltk.corpus import propbank
>>> pb_instances = propbank.instances()
>>> print(pb_instances) # doctest: +NORMALIZE_WHITESPACE
>>> inst = pb_instances[103]
>>> (inst.fileid, inst.sentnum, inst.wordnum)
inst.tagger
inst.inflection
infl = inst.inflection
infl.form, infl.tense, infl.aspect, infl.person, infl.voice
inst.roleset
inst.predicate
inst.arguments
print(inst.predicate.wordnum, inst.predicate.height)
>>> tree = inst.tree
>>> from nltk.corpus import treebank
>>> assert tree == treebank.parsed_sents(inst.fileid)[inst.sentnum]
>>> inst.predicate.select(tree)
>>> for (argloc, argid) in inst.arguments:
... print('%-10s %s' % (argid, argloc.select(tree).pformat(500)[:50]))
>>> treepos = inst.predicate.treepos(tree)
>>> print (treepos, tree[treepos])
| notebook/procs-nltk-propbank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_mxnet_p36
# language: python
# name: conda_mxnet_p36
# ---
# 
# # <a name="0">Machine Learning Accelerator - Tabular Data - Lecture 1</a>
#
#
# ## Exploratory data analysis
#
# In this notebook, we go through basic steps of exploratory data analysis (EDA), performing initial data investigations to discover patterns, spot anomalies, and look for insights to inform later ML modeling choices.
#
# 1. <a href="#1">Read the dataset</a>
# 2. <a href="#2">Overall Statistics</a>
# 3. <a href="#3">Univariate Statistics: Basic Plots</a>
# 4. <a href="#4">Multivariate Statistics: Scatter Plots and Correlations</a>
# 5. <a href="#5">Handling Missing Values</a>
# * <a href="#51">Drop columns with missing values</a>
# * <a href="#52">Drop rows with missing values</a>
# * <a href="#53">Impute (fill-in) missing values with .fillna()</a>
# * <a href="#54">Impute (fill-in) missing values with sklearn's SimpleImputer</a>
#
# __Austin Animal Center Dataset__:
#
# In this exercise, we are working with pet adoption data from __Austin Animal Center__. We have two datasets that cover intake and outcome of animals. Intake data is available from [here](https://data.austintexas.gov/Health-and-Community-Services/Austin-Animal-Center-Intakes/wter-evkm) and outcome is from [here](https://data.austintexas.gov/Health-and-Community-Services/Austin-Animal-Center-Outcomes/9t4d-g238).
#
# In order to work with a single table, we joined the intake and outcome tables using the "Animal ID" column and created a single __review.csv__ file. We also didn't consider animals with multiple entries to the facility to keep our dataset simple. If you want to see the original datasets and the merged data with multiple entries, they are available under data/review folder: Austin_Animal_Center_Intakes.csv, Austin_Animal_Center_Outcomes.csv and Austin_Animal_Center_Intakes_Outcomes.csv.
#
# __Dataset schema:__
# - __Pet ID__ - Unique ID of pet
# - __Outcome Type__ - State of pet at the time of recording the outcome (0 = not placed, 1 = placed). This is the field to predict.
# - __Sex upon Outcome__ - Sex of pet at outcome
# - __Name__ - Name of pet
# - __Found Location__ - Found location of pet before entered the center
# - __Intake Type__ - Circumstances bringing the pet to the center
# - __Intake Condition__ - Health condition of pet when entered the center
# - __Pet Type__ - Type of pet
# - __Sex upon Intake__ - Sex of pet when entered the center
# - __Breed__ - Breed of pet
# - __Color__ - Color of pet
# - __Age upon Intake Days__ - Age of pet when entered the center (days)
# - __Age upon Outcome Days__ - Age of pet at outcome (days)
#
# ## 1. <a name="1">Read the dataset</a>
# (<a href="#0">Go to top</a>)
#
# Let's read the dataset into a dataframe, using Pandas.
# +
import pandas as pd
import warnings
warnings.filterwarnings("ignore")
df = pd.read_csv('../data/review/review_dataset.csv')
print('The shape of the dataset is:', df.shape)
# -
# ## 2. <a name="2">Overall Statistics</a>
# (<a href="#0">Go to top</a>)
#
# We will look at number of rows, columns and some simple statistics of the dataset.
# Print the first five rows
# NaN means missing data
df.head()
# Let's see the data types and non-null values for each column
df.info()
# This prints basic statistics for numerical columns
df.describe()
# Let's separate model features and model target.
print(df.columns)
# +
model_features = df.columns.drop('Outcome Type')
model_target = 'Outcome Type'
print('Model features: ', model_features)
print('Model target: ', model_target)
# -
# We can explore the features set further, figuring out first what features are numerical or categorical. Beware that some integer-valued features could actually be categorical features, and some categorical features could be text features.
# +
import numpy as np
numerical_features_all = df[model_features].select_dtypes(include=np.number).columns
print('Numerical columns:',numerical_features_all)
print('')
categorical_features_all = df[model_features].select_dtypes(include='object').columns
print('Categorical columns:',categorical_features_all)
# -
# ## 3. <a name="3">Basic Plots</a>
# (<a href="#0">Go to top</a>)
#
# In this section, we examine our data with plots. Important note: These plots ignore null (missing) values. We will learn how to deal with missing values in the next section.
#
#
# __Bar plots__: These plots show counts of categorical data fields. __value_counts()__ function yields the counts of each unique value. It is useful for categorical variables.
#
# First, let's look at the distribution of the model target.
df[model_target].value_counts()
# __plot.bar()__ addition to the __value_counts()__ function makes a bar plot of the values.
# +
import matplotlib.pyplot as plt
# %matplotlib inline
df[model_target].value_counts().plot.bar()
plt.show()
# -
# Now onto the categorical features, exploring number of unique values per feature.
for c in categorical_features_all:
print(df[c].value_counts())
# Based on the number of unique values (unique IDs for example won't be very useful to visualize, for example), for some categorical features, let's see some bar plot visualizations. For simplicity and speed, here we only show box plots for those features with less than 50 unique values.
# +
import matplotlib.pyplot as plt
# %matplotlib inline
for c in categorical_features_all:
if len(df[c].value_counts()) < 50:
print(c)
df[c].value_counts().plot.bar()
plt.show()
# -
# __Histograms:__ Histograms show distribution of numeric data. Data is divided into "buckets" or "bins".
# +
import matplotlib.pyplot as plt
# %matplotlib inline
for c in numerical_features_all:
print(c)
df[c].plot.hist(bins=5)
plt.show()
# -
# If for some histograms the values are heavily placed in the first bin, it is good to check for outliers, either checking the min-max values of those particular features and/or explore value ranges.
for c in numerical_features_all:
print(c)
print('min:', df[c].min(), 'max:', df[c].max())
# With __value_counts()__ function, we can increase the number of histogram bins to 10 for more bins for a more refined view of the numerical features.
for c in numerical_features_all:
print(c)
print(df[c].value_counts(bins=10, sort=False))
plt.show()
# If any outliers are identified as very likely wrong values, dropping them could improve the numerical values histograms, and later overall model performance. While a good rule of thumb is that anything not in the range of (Q1 - 1.5 IQR) and (Q3 + 1.5 IQR) is an outlier, other rules for removing 'outliers' should be considered as well. For example, removing any values in the upper 1%.
for c in numerical_features_all:
print(c)
# Drop values below Q1 - 1.5 IQR and beyond Q3 + 1.5 IQR
#Q1 = df[c].quantile(0.25)
#Q3 = df[c].quantile(0.75)
#IQR = Q3 - Q1
#print (Q1 - 1.5*IQR, Q3 + 1.5*IQR)
#dropIndexes = df[df[c] > Q3 + 1.5*IQR].index
#df.drop(dropIndexes , inplace=True)
#dropIndexes = df[df[c] < Q1 - 1.5*IQR].index
#df.drop(dropIndexes , inplace=True)
# Drop values beyond 90% of max()
dropIndexes = df[df[c] > df[c].max()*9/10].index
df.drop(dropIndexes , inplace=True)
for c in numerical_features_all:
print(c)
print(df[c].value_counts(bins=10, sort=False))
plt.show()
# Let's see the histograms again, with more bins for vizibility.
for c in numerical_features_all:
print(c)
df[c].plot.hist(bins=100)
plt.show()
# ## 4. <a name="4">Scatter Plots and Correlation</a>
# (<a href="#0">Go to top</a>)
#
# ### Scatter plot
# Scatter plots are simple 2D plots of two numerical variables that can be used to examine the relationship between two variables.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
fig, axes = plt.subplots(len(numerical_features_all), len(numerical_features_all), figsize=(16, 16), sharex=False, sharey=False)
for i in range(0,len(numerical_features_all)):
for j in range(0,len(numerical_features_all)):
axes[i,j].scatter(x = df[numerical_features_all[i]], y = df[numerical_features_all[j]])
fig.tight_layout()
# -
# ### Scatterplot with Identification
#
# We can also add the target values, 0 or 1, to our scatter plot.
# +
import seaborn as sns
X1 = df[[numerical_features_all[0], numerical_features_all[1]]][df[model_target] == 0]
X2 = df[[numerical_features_all[0], numerical_features_all[1]]][df[model_target] == 1]
plt.scatter(X1.iloc[:,0],
X1.iloc[:,1],
s=50,
c='blue',
marker='o',
label='0')
plt.scatter(X2.iloc[:,0],
X2.iloc[:,1],
s=50,
c='red',
marker='v',
label='1')
plt.xlabel(numerical_features_all[0])
plt.ylabel(numerical_features_all[1])
plt.legend()
plt.grid()
plt.show()
# -
# Scatterplots with identification, can sometimes help identify whether or not we can get good separation between the data points, based on these two numerical features alone.
# ### Correlation Matrix Heatmat
# We plot the correlation matrix. Correlation scores are calculated for numerical fields.
cols=[numerical_features_all[0], numerical_features_all[1]]
#print(df[cols].corr())
df[cols].corr().style.background_gradient(cmap='tab20c')
# Similar to scatterplots, but now the correlation matrix values can more clearly pinpoint relationships between the numerical features. Correlation values of -1 means perfect negative correlation, 1 means perfect positive correlation, and 0 means there is no relationship between the two numerical features.
# ### A fancy example using Seaborn
# +
from string import ascii_letters
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
sns.set(style="white")
# Generate a large random dataset
rs = np.random.RandomState(33)
d = pd.DataFrame(data=rs.normal(size=(100, 26)),
columns=list(ascii_letters[26:]))
# Compute the correlation matrix
corr = d.corr()
# Generate a mask for the upper triangle
mask = np.triu(np.ones_like(corr, dtype=np.bool))
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(11, 9))
# Generate a custom diverging colormap
cmap = sns.diverging_palette(220, 10, as_cmap=True)
# Draw the heatmap with the mask and correct aspect ratio
sns.heatmap(corr, mask=mask, cmap=cmap, vmax=.3, center=0,
square=True, linewidths=.5, cbar_kws={"shrink": .5})
# -
# Also, more exploratory data analysis might reveal other important hidden atributes and/or relationships of the model features considered.
# ## 5. <a name="5">Handling Missing Values</a>
# (<a href="#0">Go to top</a>)
#
# * <a href="#51">Drop columns with missing values</a>
# * <a href="#52">Drop rows with missing values</a>
# * <a href="#53"> Impute (fill-in) missing values with .fillna()</a>
# * <a href="#54"> Impute (fill-in) missing values with sklearn's SimpleImputer</a>
#
# Let's first check the number of missing (nan) values for each column.
df.isna().sum()
# Let's explore a few options dealing with missing values, when there are values missing on many features, both numerical and categorical types.
# ### <a name="51">Drop columns with missing values</a>
# (<a href="#5">Go to Handling Missing Values</a>)
#
# We can drop some feautures/columns if we think there is significant amount of missing data in those features. Here we
# are dropping features having more than 20% missing values.
#
# __Hint:__ You can also use __inplace=True__ parameter to drop features inplace without assignment.
#
# +
threshold = 2/10
print((df.isna().sum()/len(df.index)))
columns_to_drop = df.loc[:,list(((df.isna().sum()/len(df.index))>=threshold))].columns
print(columns_to_drop)
df_columns_dropped = df.drop(columns_to_drop, axis = 1)
df_columns_dropped.head()
# -
df_columns_dropped.isna().sum()
df_columns_dropped.shape
# Note the reduced size of the dataset features. This can sometimes lead to underfitting models -- not having enough features to build a good model able to capture the pattern in the dataset, especially when dropping features that are essential to the task at hand.
# ### <a name="52">Drop rows with missing values</a>
# (<a href="#5">Go to Handling Missing Values</a>)
#
# Here, we simply drop rows that have at least one missing value. There are other drop options to explore, depending on specific problems.
df_missing_dropped = df.dropna()
# Let's check the missing values below.
df_missing_dropped.isna().sum()
df_missing_dropped.shape
# This approach can dramatically reduce the number of data samples. This can sometimes lead to overfitting models -- especially when the number of features is greater or comparable to the number of data samples.
# ### <a name="53">Impute (fill-in) missing values with .fillna()</a>
# (<a href="#5">Go to Handling Missing Values</a>)
#
# Rather than dropping rows (data samples) and/or columns (features), another strategy to deal with missing values would be to actually complete the missing values with new values: imputation of missing values.
#
# __Imputing Numerical Values:__ The easiest way to impute numerical values is to get the __average (mean) value__ for the corresponding column and use that as the new value for each missing record in that column.
# +
# Impute numerical features by using the mean per feature to replace the nans
# Assign our df to a new df
df_imputed = df.copy()
print(df_imputed[numerical_features_all].isna().sum())
# Impute our two numerical features with the means.
df_imputed[numerical_features_all] = df_imputed[numerical_features_all].fillna(df_imputed[numerical_features_all].mean())
print(df_imputed[numerical_features_all].isna().sum())
# -
# __Imputing Categorical Values:__ We can impute categorical values by getting the most common (mode) value for the corresponding column and use that as the new value for each missing record in that column.
# +
# Impute categorical features by using the mode per feature to replace the nans
# Assign our df to a new df
df_imputed_c = df.copy()
print(df_imputed_c[categorical_features_all].isna().sum())
for c in categorical_features_all:
# Find the mode per each feature
mode_impute = df_imputed_c[c].mode()
print(c, mode_impute)
# Impute our categorical features with the mode
# "inplace=True" parameter replaces missing values in place (no need for left handside assignment)
df_imputed_c[c].fillna(False, inplace=True)
print(df_imputed_c[categorical_features_all].isna().sum())
# -
# We can also create a new category, such as "Missing", for alll or elected categorical features.
# +
# Impute categorical features by using a placeholder value
# Assign our df to a new df
df_imputed = df.copy()
print(df_imputed[categorical_features_all].isna().sum())
# Impute our categorical features with a new category named "Missing".
df_imputed[categorical_features_all]= df_imputed[categorical_features_all].fillna("Missing")
print(df_imputed[categorical_features_all].isna().sum())
# -
# ### <a name="54">Impute (fill-in) missing values with sklearn's __SimpleImputer__</a>
# (<a href="#5">Go to Handling Missing Values</a>)
#
# A more elegant way to implement imputation is using sklearn's __SimpleImputer__, a class implementing .fit() and .transform() methods.
#
# +
# Impute numerical columns by using the mean per column to replace the nans
from sklearn.impute import SimpleImputer
# Assign our df to a new df
df_sklearn_imputed = df.copy()
print(df_sklearn_imputed[numerical_features_all].isna().sum())
imputer = SimpleImputer(strategy='mean')
df_sklearn_imputed[numerical_features_all] = imputer.fit_transform(df_sklearn_imputed[numerical_features_all])
print(df_sklearn_imputed[numerical_features_all].isna().sum())
# +
# Impute categorical columns by using the mode per column to replace the nans
# Pick some categorical features you desire to impute with this approach
categoricals_missing_values = df[categorical_features_all].loc[:,list(((df[categorical_features_all].isna().sum()/len(df.index)) > 0.0))].columns
columns_to_impute = categoricals_missing_values[1:3]
print(columns_to_impute)
from sklearn.impute import SimpleImputer
# Assign our df to a new df
df_sklearn_imputer = df.copy()
print(df_sklearn_imputer[columns_to_impute].isna().sum())
imputer = SimpleImputer(strategy='most_frequent')
df_sklearn_imputer[columns_to_impute] = imputer.fit_transform(df_sklearn_imputer[columns_to_impute])
print(df_sklearn_imputer[columns_to_impute].isna().sum())
# +
# Impute categorical columns by using a placeholder "Missing"
# Pick some categorical features you desire to impute with this approach
categoricals_missing_values = df[categorical_features_all].loc[:,list(((df[categorical_features_all].isna().sum()/len(df.index)) > 0.0))].columns
columns_to_impute = categoricals_missing_values[1:3]
print(columns_to_impute)
from sklearn.impute import SimpleImputer
# Assign our df to a new df
df_sklearn_imputer = df.copy()
print(df_sklearn_imputer[columns_to_impute].isna().sum())
imputer = SimpleImputer(strategy='constant', fill_value = "Missing")
df_sklearn_imputer[columns_to_impute] = imputer.fit_transform(df_sklearn_imputer[columns_to_impute])
print(df_sklearn_imputer[columns_to_impute].isna().sum())
| notebooks/MLA-TAB-Lecture1-EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.append('D:\\RCWA\\')
import numpy as np
import matplotlib.pyplot as plt
from numpy.linalg import cond
import cmath;
from scipy import linalg as LA
from numpy.linalg import solve as bslash
import time
from convolution_matrices.convmat1D import *
from RCWA_1D_functions.grating_fft.grating_conv import *
def nonHermitianEigenSorter(eigenvalues):
N = len(eigenvalues);
sorted_indices=[];
sorted_eigs = [];
for i in range(N):
eig = eigenvalues[i];
if(np.real(eig)>0 and np.imag(eig) == 0):
sorted_indices.append(i); sorted_eigs.append(eig);
elif(np.real(eig)==0 and np.imag(eig) > 0):
sorted_indices.append(i); sorted_eigs.append(eig);
elif(np.real(eig)>0 and abs(np.imag(eig)) > 0):
sorted_indices.append(i); sorted_eigs.append(eig);
return sorted_eigs, sorted_indices;
# Moharam et. al Formulation for stable and efficient implementation for RCWA
plt.close("all")
'''
1D TM implementation of PLANAR DIFFRACTiON
STILL NOT WORKING YET
only: sign convention is exp(-ikr) (is the positive propagating wave), so loss is + not -
source for fourier decomps is from the paper: Formulation for stable and efficient implementation of
the rigorous coupled-wave analysis of binary gratings by Moharam et. al
'''
# plt.plot(x, np.real(fourier_reconstruction(x, period, 1000, 1,np.sqrt(12), fill_factor = 0.1)));
# plt.title('check that the analytic fourier series works')
# #'note that the lattice constant tells you the length of the ridge'
# plt.show()
L0 = 1e-6;
e0 = 8.854e-12;
mu0 = 4*np.pi*1e-8;
fill_factor = 0.3; # 50% of the unit cell is the ridge material
num_ord = 3; #INCREASING NUMBER OF ORDERS SEEMS TO CAUSE THIS THING TO FAIL, to many orders induce evanescence...particularly
# when there is a small fill factor
PQ = 2*num_ord+1;
indices = np.arange(-num_ord, num_ord+1)
n_ridge = 3.48; # ridge
n_groove = 3.48; # groove (unit-less)
lattice_constant = 0.7; # SI units
# we need to be careful about what lattice constant means
# in the gaylord paper, lattice constant exactly means (0, L) is one unit cell
d = 0.46; # thickness, SI units
Nx = 2*256;
eps_r = n_groove**2*np.ones((2*Nx, 1)); #put in a lot of points in eps_r
eps_xz = np.zeros((2*Nx,1));
border = int(2*Nx*fill_factor);
eps_r[0:border] = n_ridge**2;
eps_xz[0:border] = 0.5;
#eps_xz[:] =12 ;
eps_zx = eps_xz;
fft_fourier_array = grating_fft(eps_r);
x = np.linspace(-lattice_constant,lattice_constant,1000);
period = lattice_constant;
## simulation parameters
theta = (0)*np.pi/180;
# +
## construct permittivity harmonic components E
#fill factor = 0 is complete dielectric, 1 is air
##construct convolution matrix
Ezz = np.zeros((2 * num_ord + 1, 2 * num_ord + 1)); Ezz = Ezz.astype('complex')
p0 = Nx; #int(Nx/2);
p_index = np.arange(-num_ord, num_ord + 1);
q_index = np.arange(-num_ord, num_ord + 1);
fourier_array = fft_fourier_array;#fourier_array_analytic;
detected_pffts = np.zeros_like(Ezz);
for prow in range(2 * num_ord + 1):
# first term locates z plane, 2nd locates y coumn, prow locates x
row_index = p_index[prow];
for pcol in range(2 * num_ord + 1):
pfft = p_index[prow] - p_index[pcol];
detected_pffts[prow, pcol] = pfft;
Ezz[prow, pcol] = fourier_array[p0 + pfft]; # fill conv matrix from top left to top right
# Exz = np.zeros_like(Ezz);
# Ezx = np.zeros_like(Ezz);
# Exz = 2*np.eye(PQ)
# Ezx = Exz;
# print((Exz.shape, Ezx.shape, Ezz.shape))
## FFT of 1/e;
inv_fft_fourier_array = grating_fft(1/eps_r);
##construct convolution matrix
E_conv_inv = np.zeros((2 * num_ord + 1, 2 * num_ord + 1));
E_conv_inv = E_conv_inv.astype('complex')
p0 = Nx;
p_index = np.arange(-num_ord, num_ord + 1);
for prow in range(2 * num_ord + 1):
# first term locates z plane, 2nd locates y coumn, prow locates x
for pcol in range(2 * num_ord + 1):
pfft = p_index[prow] - p_index[pcol];
E_conv_inv[prow, pcol] = inv_fft_fourier_array[p0 + pfft]; # fill conv matrix from top left to top right
# +
## specialized fourier matrices for exz
exzzxzz = eps_xz*eps_zx/eps_r;
exzzz = eps_xz/eps_r;
ezxzz = eps_zx/eps_r;
be = 1/(eps_r - eps_xz*eps_zx/eps_r);
# plt.figure();
# plt.plot(exzzxzz) #verify anisotropy is localized
# plt.plot(exzzz )
# plt.plot(ezxzz)
# plt.show()
Exzzxzz = np.zeros((2 * num_ord + 1, 2 * num_ord + 1)); Exzzxzz = Exzzxzz.astype('complex')
Exzzz = np.zeros((2 * num_ord + 1, 2 * num_ord + 1)); Exzzz = Exzzz.astype('complex')
bEr = np.zeros((2 * num_ord + 1, 2 * num_ord + 1)); bEr = bEr.astype('complex')
fourier_array_be = grating_fft(be);
fourier_array_xzzx = grating_fft(exzzxzz);
fourier_array_xzzz = grating_fft(exzzz);
for prow in range(2 * num_ord + 1):
# first term locates z plane, 2nd locates y coumn, prow locates x
row_index = p_index[prow];
for pcol in range(2 * num_ord + 1):
pfft = p_index[prow] - p_index[pcol];
detected_pffts[prow, pcol] = pfft;
Exzzxzz[prow, pcol] = fourier_array_xzzx[p0 + pfft]; # fill conv matrix from top left to top right
for prow in range(2 * num_ord + 1):
# first term locates z plane, 2nd locates y coumn, prow locates x
row_index = p_index[prow];
for pcol in range(2 * num_ord + 1):
pfft = p_index[prow] - p_index[pcol];
detected_pffts[prow, pcol] = pfft;
Exzzz[prow, pcol] = fourier_array_xzzx[p0 + pfft]; # fill conv matrix from top left to top right
for prow in range(2 * num_ord + 1):
# first term locates z plane, 2nd locates y coumn, prow locates x
row_index = p_index[prow];
for pcol in range(2 * num_ord + 1):
pfft = p_index[prow] - p_index[pcol];
detected_pffts[prow, pcol] = pfft;
bEr[prow, pcol] = fourier_array_be[p0 + pfft]; # fill conv matrix from top left to top right
Ezxzz = Exzzz
# +
## IMPORTANT TO NOTE: the indices for everything beyond this points are indexed from -num_ord to num_ord+1
## alternate construction of 1D convolution matrix
PQ =2*num_ord+1;
I = np.eye(PQ)
zeros = np.zeros((PQ, PQ))
# E is now the convolution of fourier amplitudes
wavelength_scan = np.linspace(0.5, 4, 300)
spectra = list();
spectra_T = list();
for wvlen in wavelength_scan:
j = cmath.sqrt(-1);
lam0 = wvlen; k0 = 2 * np.pi / lam0; #free space wavelength in SI units
print('wavelength: ' + str(wvlen));
## =====================STRUCTURE======================##
## Region I: reflected region (half space)
n1 = 1;#cmath.sqrt(-1)*1e-12; #apparently small complex perturbations are bad in Region 1, these shouldn't be necessary
## Region 2; transmitted region
n2 = 1;
#from the kx_components given the indices and wvln
kx_array = k0*(n1*np.sin(theta) + indices*(lam0 / lattice_constant)); #0 is one of them, k0*lam0 = 2*pi
k_xi = kx_array;
## IMPLEMENT SCALING: these are the fourier orders of the x-direction decomposition.
KX = np.diag((k_xi/k0)); #singular since we have a n=0, m= 0 order and incidence is normal
## one thing that isn't obvious is that are we doing element by element division or is it matricial
B = (KX@bslash(Ezz, KX) - I);
## SIGN MISMATCH WITH DERIVATION, first term should be positive but in the notes ,it's negative: FIX (3/4/2019)
#bE = np.linalg.inv(E_conv_inv) - Exzzxzz #bslash(Ezz,(Exz@Ezx)); #/Ezz;
bE = np.linalg.inv(bEr);
G = j* Ezxzz @ KX#j*bslash(Ezz,Ezx) @ KX;
#G = j*(Ezx/Ezz)@KX #we should not do pointwise division of these epsilon matrices.
H = -j*KX @Exzzz; #j*KX @bslash(Ezz, Exz);
#print((bE.shape,G.shape, H.shape))
print('conditioning of B and bE: '+str((np.linalg.cond(B), np.linalg.cond(bE))))
print('conditioning of G and H: '+str((np.linalg.cond(G), np.linalg.cond(H))))
bigBlock = np.block([[G, bE],[B,H]]);
print('conditioning of main block: '+str(np.linalg.cond(bigBlock)))
## these matrices aren't poorly conditioned
print('conditioning of eigenvalue prob: '+str((np.linalg.cond(OA), np.linalg.cond(OB))))
## solve eiegenvalues;
beigenvals, bigW = LA.eig(bigBlock); #W contains eigenmodes of the form (lambda x, x)
## AT THIS POINT, we have still extracted TWO times the number of eigenvalues...
#try rounding...
rounded_beigenvals = np.array([round(i,10) for i in beigenvals])
print(rounded_beigenvals)
#quadrant_sort = [1 if abs(np.real(i))>=0 and np.imag(i)>=0 else 0 for i in rounded_beigenvals];
sorted_eigs, sorted_indices = nonHermitianEigenSorter(rounded_beigenvals)
# sorted_indices = np.nonzero(quadrant_sort)[0]
print(len(sorted_indices))
#sorted_indices = np.argsort(np.real(rounded_beigenvals))
sorted_eigenmodes = bigW[:, sorted_indices];
#print(sorted_eigenmodes)
#adding real and imaginary parts seems to work...
sorted_eigenvals = beigenvals[sorted_indices]
print(sorted_eigenvals)
W = sorted_eigenmodes[0:PQ,:]
eigenvals_wp = (sorted_eigenvals[0:PQ]);
# plt.plot(np.real(beigenvals), np.imag(beigenvals), '.', markersize = 20);
# plt.plot(np.real(eigenvals_wp), (np.imag(eigenvals_wp)), '.r', markersize = 10)
# plt.legend(('original', 'sorted'))
# plt.show();
##
Q = np.diag(eigenvals_wp); #eigenvalue problem is for kz, not kz^2
## IS THIS RIGHT? #Q is a diagonal matrix of eigenvalues. W contains modes in columns... we want to column scale
# so Q should be on the RHS of W.
#V = np.linalg.inv(bE)@(W @ Q + H@W);
V = sorted_eigenmodes[PQ:,:]
#enforcing negative sign convention.
X = np.diag(np.exp(-k0*np.diag(Q)*d)); #this is poorly conditioned because exponentiation
## pointwise exponentiation vs exponentiating a matrix
## observation: almost everything beyond this point is worse conditioned
k_I = k0**2*(n1**2 - (k_xi/k0)**2); #k_z in reflected region k_I,zi
k_II = k0**2*(n2**2 - (k_xi/k0)**2); #k_z in transmitted region
## are these formulas correct, as they only pertain teo the reflected and transmitted regions.
k_I = k_I.astype('complex'); k_I = np.sqrt(k_I);
k_II = k_II.astype('complex'); k_II = np.sqrt(k_II);
Z_I = np.diag(k_I / (n1**2 * k0 ));
Z_II = np.diag(k_II /(n2**2 * k0));
delta_i0 = np.zeros((len(kx_array),1));
delta_i0[num_ord] = 1;
n_delta_i0 = delta_i0*j*np.cos(theta)/n1;
O = np.block([
[W, W],
[V,-V]
]); #this is much better conditioned than S..
print('condition of O: '+str(np.linalg.cond(O)))
print((np.linalg.cond(W), np.linalg.cond(V)))
# plt.imshow(abs(O))
# plt.show();
f = I;
g = j * Z_II; #all matrices
fg = np.concatenate((f,g),axis = 0)
ab = np.matmul(np.linalg.inv(O),fg);
a = ab[0:PQ,:];
b = ab[PQ:,:];
term = X @ a @ np.linalg.inv(b) @ X;
f = W @ (I + term);
g = V@(-I+term);
## does this change?, I think it might...
T = np.linalg.inv(np.matmul(j*Z_I, f) + g);
T = np.dot(T, (np.dot(j*Z_I, delta_i0) + n_delta_i0));
R = np.dot(f,T)-delta_i0; #shouldn't change
T = np.dot(np.matmul(np.linalg.inv(b),X),T)
## calculate diffraction efficiencies
#I would expect this number to be real...
DE_ri = R*np.conj(R)*np.real(np.expand_dims(k_I,1))/(k0*n1*np.cos(theta));
DE_ti = T*np.conj(T)*np.real(np.expand_dims(k_II,1)/n2**2)/(k0*np.cos(theta)/n1);
print('R(lam)='+str(np.sum(DE_ri))+' T(lam) = '+str(np.sum(DE_ti)))
spectra.append(np.sum(DE_ri)); #spectra_T.append(T);
spectra_T.append(np.sum(DE_ti))
# +
spectra = np.array(spectra);
spectra_T = np.array(spectra_T)
plt.figure();
plt.plot(wavelength_scan, spectra);
plt.plot(wavelength_scan, spectra_T)
plt.plot(wavelength_scan, spectra+spectra_T)
# plt.legend(['reflection', 'transmission'])
# plt.axhline(((3.48-1)/(3.48+1))**2,xmin=0, xmax = max(wavelength_scan))
# plt.axhline(((3.48-1)/(3.48+1)),xmin=0, xmax = max(wavelength_scan), color='r')
#
plt.ylim([0,2])
plt.show()
plt.figure();
plt.plot(1/wavelength_scan, spectra);
plt.plot(1/wavelength_scan, spectra_T)
plt.plot(1/wavelength_scan, spectra+spectra_T)
plt.show()
# -
sorted_eigenvals.shape
sorted_eigenmodes.shape
| anisotropy_explorations/.ipynb_checkpoints/Longitudinal Anisotropy Full First-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Author: <NAME>
# ## Task 2: Prediction using Unsupervised ML
# ### GRIP(JAN2021): THE SPARKS FOUNDATION
# *From the given dataset, I am gonna predict the optimum number of clusters and represent it visually.*
# Dataset:https://bit.ly/3kXTdox
# *About Iris dataset: The Iris flower dataset or Fisher's Iris dataset is a multivariate dataset introduced by the British statistician, eugenicist, and biologist <NAME> in his 1936 as an example of linear discriminant analysis. The dataset contains a set of 150 records under five attributes - sepal length, sepal width, petal length, petal width and species. Iris data set gives the measurements in centimetres of the variables sepal length and width and petal length and width, respectively, for 50 flowers from each of 3 species of iris. The species are Iris setosa, versicolor, and virginica.*
# 
# 
# **Step 1: Import required libraries**
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import seaborn as sns
# **Step 2: Import dataset**
data=pd.read_csv(r"C:\Users\panch\Desktop\Data\Iris.csv")
print("Data imported successfully!")
data
data.head(10)
# **Step 3: Learn data**
data.info()
# *From above information I can say that there's no null value in the dataset so there's no need of data Cleaning*
#statistical description of data
data.describe()
# **Step 4: Data Visualization**
sns.set_style("whitegrid")
sns.FacetGrid(data, hue ="Species",height = 5).map(plt.scatter,'SepalLengthCm','PetalLengthCm').add_legend()
# *From data visualization, I can see that there's some data points that are to be grouped in some places. And for that next I'm using K-Means algorithm for Cluster Analysis, so from that I can obtain a valid result of the data being clustered at some places.*
# **Step 5: Clustering using K-Means Algorithm**
#importing required library for k-means ML algo
from sklearn.cluster import KMeans
#seleccting data by label
x = data.iloc[:, [0, 1, 2, 3]].values
#calculating distance between each data point and cluster centers
sum_of_squared_distance = []
K = range(1,10)
optimalK = 1
for k in K:
km = KMeans(n_clusters=k)
km = km.fit(x)
sum_of_squared_distance.append(km.inertia_)
if k > 1:
ratio = sum_of_squared_distance[k-1]/sum_of_squared_distance[k-2]
if ratio < 0.55:
optimalK = k
print("Optimal Number of Clusters =",optimalK)
# **Step 6: Visual Representation of Optimal number of clusters using elbow method**
# Plotting Elbow Graph
plt.plot(K, sum_of_squared_distance, 'bx-')
plt.xlabel('Number of Clusters, k')
plt.ylabel('Sum of Squared Distances')
plt.title('Elbow Method for Optimal k')
plt.show()
# *From the above grapgh, I can visualize that Optimal clusters is where the elbow occurs. Thus this is the point when the within cluster sum of squares does not decrease significantly with every iteration.*
#
# Therefore, there are 3 clusters for this dataset.
# **Step 7: Apply K-means to the dataset**
kmeans = KMeans(n_clusters = 3)
y_kmeans = kmeans.fit_predict(x)
kmeans.cluster_centers_
# +
# Visualising the clusters
plt.scatter(x[y_kmeans == 0, 0], x[y_kmeans == 0, 1],
s = 25, c = 'red', label = 'Iris-setosa')
plt.scatter(x[y_kmeans == 1, 0], x[y_kmeans == 1, 1],
s = 25, c = 'blue', label = 'Iris-versicolour')
plt.scatter(x[y_kmeans == 2, 0], x[y_kmeans == 2, 1],
s = 25, c = 'green', label = 'Iris-virginica')
# Plotting the centroids of the clusters
plt.scatter(kmeans.cluster_centers_[:,0], kmeans.cluster_centers_[:,1],s = 100, c = 'purple', label = 'Centroids')
plt.legend()
# -
# ### Conclusion:
# #### I can conclude that the Iris data set is divided into 3 clusters by not only using the k-means algorithm but also visually.
| IrisFlowerClassification_UL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## ChemDataExtractor (CDE) and Paper-parser (PP)
#
# environment setting:
# * use python version 2.7 - 3.6
# * all requirements are inside requirements.txt
# * use pipenv or conda to build virtual env
#
# The scientific abstracts used in this study are available via Elsiver's Scopus and Science Direct API's (https://dev.elsvier.com/), the Springer Nature API (https://dev.springernature.com/),the Royal Chemistry Society API. The list of DOIs used in this study, the pretrained model is built in chemdataextractor package, while extensive parsing rules and searching standards are done by the author.
#
# **Author:** <NAME>
#
#
# ### Extracting a Custom Property
import chemdataextractor
import chemdataextractor.model as model
from chemdataextractor.model import Compound
from chemdataextractor.doc import Document, Heading, Paragraph, Sentence
# ### Example Document
#
# Let's create a simple example document with a single heading followed by a single paragrah:
# +
d = Document(
Heading(u'Synthesis of 2,4,6-trinitrotoluene (3a)'),
Paragraph(u'The procedure was followed to yield a pale yellow solid (b.p. 240 °C)')
)
# display a heading and a paragraph of the given content
d
# -
# ### Default Parsers
#
# By default, chemdataextractor won't extract the boiling point property:
d.records.serialize()
# ### Defining a New Property Model
#
# The first task is to define the schema of a new property, and add it to the `Compound` model:
# +
from chemdataextractor.model import BaseModel, StringType, ListType, ModelType
class BoilingPoint(BaseModel):
value = StringType()
units = StringType()
Compound.boiling_points = ListType(ModelType(BoilingPoint))
# -
# ### Then Writing a New Parser
#
# Next, define parsing rules that define how to interpret text and convert it into the model:
# +
import re
from chemdataextractor.parse import R, I, W, Optional, merge
prefix = (R(u'^b\.?p\.?$', re.I) | I(u'boiling') + I(u'point')).hide()
units = (W(u'°') + Optional(R(u'^[CFK]\.?$')))(u'units').add_action(merge)
value = R(u'^\d+(\.\d+)?$')(u'value')
bp = (prefix + value + units)(u'bp')
# +
from chemdataextractor.parse.base import BaseParser
from chemdataextractor.utils import first
class BpParser(BaseParser):
root = bp
def interpret(self, result, start, end):
compound = Compound(
boiling_points=[
BoilingPoint(
value=first(result.xpath('./value/text()')),
units=first(result.xpath('./units/text()'))
)
]
)
yield compound
# -
Paragraph.parsers = [BpParser()]
# ### Running the New Parser
# +
d = Document(
Heading(u'Synthesis of 2,4,6-trinitrotoluene (3a)'),
Paragraph(u'The procedure was followed to yield a pale yellow solid (b.p. 240 °C)')
)
d.records.serialize()
# -
# ## Example from Documentation
doc = Document('UV-vis spectrum of 5,10,15,20-Tetra(4-carboxyphenyl)porphyrin in Tetrahydrofuran (THF).')
doc
# each individual chemical entity mention (CEM)
doc.cems
doc.abbreviation_definitions
doc.records
doc.records[0].serialize()
doc.records[1].serialize()
# ## Trying on OPV Documents
#
# we are going to extract some basic OPV optoelectronic properties from the documents. Here is the list of some fundamental properties:
#
# * Voc (V)
# * Jsc (mA cm-2)
# * FF (%)
# * PCE (%)
# * Bandgap, energy loss (Eloss), offsets (eV)
# * active area (cm2)
# * exposure time (s, min, hr, d)
# * Molecuar weight (Mw) (kg/mol)
# * hole and electron mobilities (cm2 V-1 s-1)
# * EQE, IQE (unitless)
# * absorption (nm)
#
# Since spectroscopy and other propertis can be automatically extracted, we are not going to focus too much on them.
#
# But we also want to extract AFM and TEM images from documents and their corresponding roughness and dump them into chemical database
#
# **Reading documents can break the doc into either paragraphs, and from paragraphs they can derive sentences or tokens**
# While ChemDataExtractor supports documents in a wide variety of formats, some are better suited for extraction than others. If there is an HTML or XML version available, that is normally the best choice.
#
# Wherever possible, avoid using the PDF version of a paper or patent. At best, the text will be interpretable, but it is extremely difficult to reliably distinguish between headings, captions and main body text. At worst, the document will just consist of a scanned image of each page, and it won't be possible to extract any of the text at all. You can get some idea of what ChemDataExtractor can see in a PDF by looking at the result of copying-and-pasting from the document.
#
# For scientific articles, most publishers offer a HTML version alongside the PDF version. Normally, this will open as a page in your web browser. Just choose "Save As..." and ensure the selected format is "HTML" or "Page Source" to save a copy of the HTML file to your computer.
#
# Most patent offices provide XML versions of their patent documents, but these can be hard to find. Two useful resources are the USPTO Bulk Data Download Service and the EPO Open Patent Services API.
#
# **Reference**: http://www.chemdataextractor.org
f = open('example_doc.pdf', 'rb')
doc = Document.from_file(f)
doc.elements
doc.cems
# the problem is author's name might be mistaken as a chemical name
# cems is returned as a `Span`, which contains the mention text, as well as
# the start and end character offsets within the containing document element.
# element types include Title, Heading, Paragraph, Citation, Table, Figure, Caption and FootNode.
# you can retrieve a specific element by its index within the document
para = doc.elements[3]
para
# you can get the individual sentences of a paragraph
para.sentences
para.tokens
# ### Designing New Parsing Rules for Properties
#
# 1. defining a new property model
# 2. writing a new parser
# 3. running the new parser
#
# we can use functions described in chemdataextractor website
#
# ChemDataExtractor contains a chemistry-aware Part-of-speech tagger. Use the `pos_tagged_tokens` property on a document element to get the tagged tokens:
# use pos_tagged_tokens property on a document element to get the tagged tokens
s = Sentence('1H NMR spectra were recorded on a 300 MHz BRUKER DPX300 spectrometer.')
s.pos_tagged_tokens
# using taggers directly
from chemdataextractor.nlp.pos import ChemCrfPosTagger
cpt = ChemCrfPosTagger()
cpt.tag(['1H', 'NMR', 'spectra', 'were', 'recorded', 'on', 'a', '300', 'MHz', 'BRUKER', 'DPX300', 'spectrometer', '.'])
# +
from chemdataextractor.model import BaseModel, StringType, ListType, ModelType
class Mobility(BaseModel):
value = StringType()
units = StringType()
class Voc(BaseModel):
value = StringType()
units = StringType()
class FF(BaseModel):
value = StringType()
units = StringType()
class PCE(BaseModel):
value = StringType()
units = StringType()
class Area(BaseModel):
value = StringType()
units = StringType()
Compound.Jsc = ListType(ModelType(Jsc))
Compound.Voc = ListType(ModelType(Voc))
Compound.FF = ListType(ModelType(FF))
Compound.PCE = ListType(ModelType(PCE))
Compound.Area = ListType(ModelType(Area))
# +
import re
from chemdataextractor.parse import R, I, W, Optional, merge
prefix = (I(u'mobilities')).hide()
units = (W(u'cm2 V−1 s−1') + Optional(R(u'^[CFK]\.?$')))(u'units').add_action(merge)
value = R(u'^\d+(\.\d+)?$')(u'value')
mu = (prefix + units + value)(u'mu')
# +
from chemdataextractor.parse.base import BaseParser
from chemdataextractor.utils import first
class muParser(BaseParser):
root = mu
def interpret(self, result, start, end):
compound = Compound(
mobility=[
Mobility(
value=first(result.xpath('./value/text()')),
units=first(result.xpath('./units/text()'))
)
]
)
yield compound
Paragraph.parsers = [muParser()]
d = Document(
Paragraph(u'Thiophene-substituted DPP polymers show high hole and electron mobilities above 1 cm2 V−1 s−1 in FETs.16 By copolymerization of the thiophene-substituted DPP with π- conjugated aromatic monomers of different donor strength, such as biphenyl,23 phenyl,8 thiophene,8 and dithienopyrrole,24 the absorption onset can be tuned from 750 nm to above 1000 nm. The PCEs can reach 9.4%.')
)
d.records.serialize()
# -
d = Document(
Paragraph(u'Thiophene-substituted DPP polymers show high hole and electron mobilities above 1 cm2 V−1 s−1 in FETs.16 By copolymerization of the thiophene-substituted DPP with π- conjugated aromatic monomers of different donor strength, such as biphenyl,23 phenyl,8 thiophene,8 and dithienopyrrole,24 the absorption onset can be tuned from 750 nm to above 1000 nm. The PCEs can reach 9.4%.')
)
# we can see cems working fine
d.cems
# but d.records.serialize() doesn't return anything, only []
# ### As shown above and in the extracting_a_custom_property notebook, there is no
| DataExtractor/notebook/Demos/chemdataextractor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Importing CSSEGI data into communities
# [Index](0-index.ipynb)
#
# In this notebook, we assign CSSEGI data to the communities constructed in [1-clustering](1-clustering.ipynb). Data reported in the CSSEGI dataset is assigned to a community based on the latitude and longitude. Data can be accessed [here](https://github.com/CSSEGISandData/COVID-19).
# ## Imports and global variables
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# +
from pathlib import Path
import sys
import numpy as np
import pandas as pd
import geopandas as gpd
import datetime
import json
import matplotlib.pyplot as plt
import matplotlib.colors as mco
import matplotlib.gridspec as mgs
import matplotlib.ticker as ticker
import matplotlib.cm as cm
from matplotlib import animation
plt.rcParams['svg.fonttype'] = 'none'
from IPython.display import HTML
from IPython.display import Image
# -
sys.path.append('../code')
from functions import plot_omega_profile, plot_omega_map
datadir = Path('../data')
if not datadir.is_dir():
raise ValueError("Data dir doesn'nt exist!")
resdir = Path('../results/')
if not resdir.is_dir():
raise ValueError('No results directory!')
# +
complevel=7
complib='zlib'
resfile = resdir / 'safegraph_analysis.hdf5'
# resfile = resdir / 'safegraph_analysis_monthly.hdf5'
with pd.HDFStore(resfile, complevel=complevel, complib=complib) as store:
print(f"File {resfile.stem} has {len(store.keys())} entries.")
cssefile = resdir / 'cssegi_sanddata_analysis.hdf5'
with pd.HDFStore(cssefile, complevel=complevel, complib=complib) as store:
print(f"File {cssefile.stem} has {len(store.keys())} entries.")
# -
tfmt = '%Y-%m-%d'
exts = ['.png', '.svg']
# ## Load clustered CBGs
key = "/clustering/clusters"
with pd.HDFStore(resfile, complevel=complevel, complib=complib) as store:
clusters = store[key]
clusters
clusters_pos = clusters.loc[:, ['X','Y']].rename(columns={'X': 'longitude', 'Y': 'latitude'})
clusters_pos
# ## Load data from geometry
geofile = datadir / 'safegraph_open_census_data' / 'geometry' / 'cbg.geojson'
if not geofile.is_file():
raise ValueError("Geo file doesn't exist!")
geo = gpd.read_file(geofile).astype({'CensusBlockGroup': 'int64'})
geo.set_index('CensusBlockGroup', inplace=True)
XY = geo.representative_point().apply(lambda X: [X.x, X.y]).tolist()
df_xy = pd.DataFrame(data=np.array(XY), index=geo.index, columns=['longitude','latitude'])
df_xy
# ## Load the confirmed cases from CSSEGI data
dfile = datadir / 'time_series_covid19_confirmed_US.csv'
df_confirmed_us = pd.read_csv(dfile)
df_confirmed_us = df_confirmed_us.loc[df_confirmed_us['iso2'] == 'US'] # only keep the US
#.astype({'FIPS': 'int64'})
df_confirmed_us
# Drop the locations with no latitude or longitude information
idx = ( (df_confirmed_us['Lat'] == 0.) | (df_confirmed_us['Long_'] == 0.) )
df_confirmed_us.drop(index=df_confirmed_us.index[idx], inplace=True)
df_confirmed_us
# ## Mapping clusters to CSSEGI FIPS based on (Longitude, Latitude)
clusters_pos.sort_index(inplace=True)
XY_clusters = clusters_pos.loc[:, ['longitude', 'latitude']].to_numpy()
XY_csse = df_confirmed_us.loc[:, ['Long_', 'Lat']].to_numpy()
# iterate over CSSEGI data
cluster_ids = []
for i in range(len(df_confirmed_us.index)):
xy = XY_csse[i]
imin = np.argmin(np.linalg.norm(XY_clusters - xy, axis=1))
cluster_ids.append(clusters_pos.index[imin])
# +
col = 'cluster_id'
if col in df_confirmed_us.columns:
del df_confirmed_us[col]
df_confirmed_us.insert(1, col, cluster_ids)
df_confirmed_us
# -
# ### Retain only the dates and cluster_ID
dates = []
for c in df_confirmed_us.columns:
try:
t = datetime.datetime.strptime(c, '%m/%d/%y')
dates.append(c)
except ValueError:
continue
# +
columns = ['cluster_id'] + dates
df_clusters = df_confirmed_us.loc[:, columns].groupby('cluster_id').apply(sum)
del df_clusters['cluster_id']
df_clusters
# -
clusters_csse = pd.DataFrame(data=np.zeros((len(clusters.index),len(df_clusters.columns)), np.uint), index=clusters.index, columns=df_clusters.columns)
clusters_csse.loc[df_clusters.index] = df_clusters
clusters_csse = clusters_csse.T
clusters_csse.index = [datetime.datetime.strptime(s, '%m/%d/%y') for s in clusters_csse.index]
clusters_csse
# I currently have one empty community.
clusters.loc[1017].to_frame().T
# But the reported number of cases there is zero.
clusters_csse.loc[clusters_csse.index[-1], 1017]
# There is one community where the reported number of cases is larger than the population...
idx = (clusters_csse.loc[clusters_csse.index[-1]] > clusters['population']).to_numpy()
np.sum(idx)
clusters.loc[clusters.index[idx]]
clusters_csse.loc[clusters_csse.index[-1], clusters.index[idx]].to_frame().T
# So I correct this:
# +
for t in clusters_csse.index:
x = np.min(np.array([clusters_csse.loc[t].to_numpy(), clusters['population'].to_numpy()], dtype=np.int_), axis=0)
clusters_csse.loc[t] = x
clusters_csse
# -
idx = (clusters_csse.loc[clusters_csse.index[-1]] > clusters['population']).to_numpy()
np.sum(idx)
total_csse = clusters_csse.sum(axis=1).to_frame()
total_csse.rename(columns={0: 'omega'}, inplace=True)
total_csse
# ### Write
key = '/clustering/cssegi'
with pd.HDFStore(resfile, complevel=complevel, complib=complib) as store:
store[str(key)] = clusters_csse
# ## Plots
figdir = Path('../figures') / '3-import_cssegi'
if not figdir.is_dir():
figdir.mkdir(parents=True, exist_ok=True)
# ### Plot the total evolution
# +
# parameters
figsize = (6,4.5)
dpi = 300
ms=2
lw=1
fig = plt.figure(facecolor='w', figsize=figsize)
ax = fig.gca()
ax.plot(total_csse.index.to_numpy(), total_csse.sum(axis=1).to_numpy(), '-', ms=ms, color='darkblue')
ax.set_yscale('log')
ax.set_xlim(total_csse.index[0],None)
ax.set_ylabel("$T$", fontsize="medium")
plt.xticks(rotation=45)
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.tick_params(left=True, labelleft=True, bottom=True, labelbottom=True)
ax.tick_params(axis='both', length=4)
fig.tight_layout()
fname = 'omega_tot_cssegi'
for ext in exts:
filepath = figdir / (fname + ext)
fig.savefig(filepath, bbox_inches='tight', pad_inches=0, dpi=dpi)
print("Written file: {:s}".format(str(filepath)))
fig.clf()
plt.close('all')
# -
filepath = figdir / (fname + '.png')
Image(filename=filepath, width=4./3*360)
# ### Plot the spatial evolution as a profile
clusters_csse.iloc[-1].max()/1.0e6
# +
# parameters
dpi=150
fps=10
figsize=(6, 4.5)
lw=0.5
ms=4
ylabel="$T_a$"
fileout = figdir / 'dynamic_profile.mp4'
mydir = figdir / 'profiles'
if not mydir.is_dir():
mydir.mkdir(parents=True, exist_ok=True)
# -
plot_omega_profile(np.array([clusters_csse.to_numpy()]), clusters_csse.index.to_list(), colors=['red'], \
fileout=fileout, tpdir=mydir, dpi=dpi, fps=fps, figsize=figsize, ylabel=ylabel, \
lw=lw, ms=ms, styles=['o'], deletetp=False, exts=['.png','.svg'], ymin=1., ymax=1.3e6)
HTML("""
<video height="480" controls>
<source src="{:s}" type="video/mp4">
</video>
""".format(str(fileout)))
# ### Plot the spatial evolution on a map
clusters_csse.iloc[-1].max()/1.0e6
fileout = figdir / 'dynamic_profile.mp4'
mydir = figdir / 'maps'
if not mydir.is_dir():
mydir.mkdir(parents=True, exist_ok=True)
fileout = figdir / 'dynamic_map.mp4'
# +
# parameters
dpi=150
fps=10
figsize=(6, 4.5)
lw=0.5
ms=4
idump=1
fileout = figdir / 'map_T.mp4'
plot_omega_map(clusters_csse.to_numpy(), clusters_csse.index.to_list(), XY=clusters.loc[:, ['X', 'Y']].to_numpy().T, \
fileout=fileout, tpdir=mydir, dpi=dpi, fps=fps, figsize=figsize, idump=idump, \
clabel="$T$", vmin=1., vmax=1.3e6, deletetp=False, exts=['.png','.svg'])
# -
HTML("""
<video height="480" controls>
<source src="{:s}" type="video/mp4">
</video>
""".format(str(fileout)))
| notebooks/3-import_cssegi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# from _future_ import absolute_import, division, print_funciton, unicode_literals
import numpy as np
import tensorflow as tf
import tensorflow_hub as hub
import tensorflow_datasets as tfds
print("Version: ", tf._version_)
print("Eager mode: ", tf.executing_eagerly())
print("Hub version: ", hub._version_)
print("GPU is,""avaliable" if tf.config.experimental.list_physical_devices("GPU") else "NOT AVALIABLE")
# +
train_validation_split = tfds.Split.TRAIN.subsplit([6,4])
(train_data, validation_data), test_data = tfds.load(
name="imdb_reviews",
split=(train_validation_split, tfds.Split.TEST),
as_supervised=True
)
# -
train_examples_batch, train_labels_batch = (next(iter(train_data.batch(10))))
train_examples_batch
train_labels_batch
embedding = 'https://tfhub.dev/google/tf2-preview/gnews-swivel-20dim/1'
hub_layer = hub.KerasLayer(embedding, input_shape=[], dtype=tf.string, trainable=True)
hub_layer(train_examples_batch[:3])
# +
model = tf.keras.Sequential()
model.add(hub_layer)
model.add(tf.keras.layers.Dense(16, activation='relu'))
model.add(tf.keras.layers.Dense(1, activation='sigmoid'))
model.summary()
| wine_data/tipsy_sentiments/imdb_tf_keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] _cell_guid="b7dbedec-9483-4b22-484d-b8a8e04c49ae" _uuid="8807ae46aec34ec427b81344d64a3c0d607b258a"
# ### Hello Kagglers!!
#
# This is a very basic tutorial to Machine Learning for complete Beginners using the Iris Dataset. You can learn how to implement a machine learning to a given dataset by following this notebook. I have explained everything related to the implementation in detail . Hope you find it useful.
#
# For a more advanced notebook that covers some more detailed concepts, have a look at [this notebook](https://www.kaggle.com/ash316/ml-from-scratch-part-2/notebook)
#
# If this notebook to be useful, **Please Upvote!!!**
#
#
# + _cell_guid="c8b7047a-c84c-c0ee-054d-a0c30609cc43" _execution_state="idle" _uuid="43047631b7881a23c63a655f5214b2ebaff946fa"
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import seaborn as sns
import matplotlib.pyplot as plt
# +
# Using MarvinData utility to download file
from marvin_python_toolbox.common.data import MarvinData
# getting the initial data set
file_path = MarvinData.download_file(url="https://s3.amazonaws.com/marvin-engines-data/Iris.csv")
# + _cell_guid="ced2723b-e83e-6aa0-4ffb-9ace2cc4a5e3" _execution_state="idle" _uuid="db8aed63638fcf4833c050541cc0fe5a0b756670"
iris = pd.read_csv(file_path) #load the dataset
# + _cell_guid="ca50ed92-15c7-f9f6-b371-f18393505167" _execution_state="idle" _uuid="9a89db0e5a9d3e5cd7cecf06e16dbf1b6f8d3542"
iris.head(2) #show the first 2 rows from the dataset
# + _cell_guid="4f9370f1-0672-0d8c-4f21-c7eeb694042c" _execution_state="idle" _uuid="46f49d63be72b4eec75da299e355d6dbaa19f65b"
iris.info() #checking if there is any inconsistency in the dataset
#as we see there are no null values in the dataset, so the data can be processed
# + [markdown] _cell_guid="fe064d83-88de-20ee-9ec0-43e52fc49c03" _uuid="eaf309bd3956c549ca5c039a6c62151974ca9481"
# #### Removing the unneeded column
# + _cell_guid="af6dd1be-5c24-27f3-6319-eb6af2c65c27" _execution_state="idle" _uuid="c9b2172cc499bda6d56d2199b8d286217abbd677"
iris.drop('Id',axis=1,inplace=True) #dropping the Id column as it is unecessary, axis=1 specifies that it should be column wise, inplace =1 means the changes should be reflected into the dataframe
# + [markdown] _cell_guid="a2e19920-d24b-7551-10c4-e1088c8a4324" _uuid="f00690de3343002575e680a83ae395ef0060fe1f"
# ## Some Exploratory Data Analysis With Iris
# + _cell_guid="09a16bf0-067b-8da0-3eed-2014dc8cfec7" _execution_state="idle" _uuid="c9fd2c2b0eb51ada3a4f57eb51023cb080a0d308"
fig = iris[iris.Species=='Iris-setosa'].plot(kind='scatter',x='SepalLengthCm',y='SepalWidthCm',color='orange', label='Setosa')
iris[iris.Species=='Iris-versicolor'].plot(kind='scatter',x='SepalLengthCm',y='SepalWidthCm',color='blue', label='versicolor',ax=fig)
iris[iris.Species=='Iris-virginica'].plot(kind='scatter',x='SepalLengthCm',y='SepalWidthCm',color='green', label='virginica', ax=fig)
fig.set_xlabel("Sepal Length")
fig.set_ylabel("Sepal Width")
fig.set_title("Sepal Length VS Width")
fig=plt.gcf()
fig.set_size_inches(10,6)
plt.show()
# + [markdown] _cell_guid="e853b9fa-a1db-cdc8-1f8f-0f5f2f2d1ab6" _uuid="d0793e117fd202f679b6ecb1239e0ab0a8cfd137"
# The above graph shows relationship between the sepal length and width. Now we will check relationship between the petal length and width.
# + _cell_guid="ea5060b8-4067-cf46-99d6-a27be10a7e18" _execution_state="idle" _uuid="16a42e3a6615e48f7b8ed0a6dbb380de74dcd2c8"
fig = iris[iris.Species=='Iris-setosa'].plot.scatter(x='PetalLengthCm',y='PetalWidthCm',color='orange', label='Setosa')
iris[iris.Species=='Iris-versicolor'].plot.scatter(x='PetalLengthCm',y='PetalWidthCm',color='blue', label='versicolor',ax=fig)
iris[iris.Species=='Iris-virginica'].plot.scatter(x='PetalLengthCm',y='PetalWidthCm',color='green', label='virginica', ax=fig)
fig.set_xlabel("Petal Length")
fig.set_ylabel("Petal Width")
fig.set_title(" Petal Length VS Width")
fig=plt.gcf()
fig.set_size_inches(10,6)
plt.show()
# + [markdown] _cell_guid="3068e91a-2455-f7ae-b1d0-a2114b78ea62" _uuid="dd90c7e433f42fea8e5836c91959429710491347"
# As we can see that the Petal Features are giving a better cluster division compared to the Sepal features. This is an indication that the Petals can help in better and accurate Predictions over the Sepal. We will check that later.
# + [markdown] _cell_guid="ac6bb577-9975-39d5-aa20-376e574e703c" _uuid="aaa439a16175b442a42e845060b7fe1bd2d246f0"
# ### Now let us see how are the length and width are distributed
# + _cell_guid="d818068d-5110-c64e-ec6e-92bda44a9723" _execution_state="idle" _uuid="0ca23638e6e067ffd7501552f346674646e3e1d2"
iris.hist(edgecolor='black', linewidth=1.2)
fig=plt.gcf()
fig.set_size_inches(12,6)
plt.show()
# + [markdown] _cell_guid="cdb55848-d7eb-d66c-dec1-79cbbfba2826" _uuid="3e8d0f657f996be6f4020b1850148bade9fdc588"
# ### Now let us see how the length and width vary according to the species
# + _cell_guid="a1d57f07-5c6b-4ab3-ad15-b8b5245c05b9" _execution_state="idle" _uuid="15e65581708a5b67ac8f0c1994fa08ef8158e799"
plt.figure(figsize=(15,10))
plt.subplot(2,2,1)
sns.violinplot(x='Species',y='PetalLengthCm',data=iris)
plt.subplot(2,2,2)
sns.violinplot(x='Species',y='PetalWidthCm',data=iris)
plt.subplot(2,2,3)
sns.violinplot(x='Species',y='SepalLengthCm',data=iris)
plt.subplot(2,2,4)
sns.violinplot(x='Species',y='SepalWidthCm',data=iris)
plt.show()
# + [markdown] _cell_guid="2497705a-9c08-5d8b-6ca0-0edc509e73b9" _uuid="0e907abebd6751fa3597fd6b08bb24eb13081443"
# The violinplot shows density of the length and width in the species. The thinner part denotes that there is less density whereas the fatter part conveys higher density
# + [markdown] _cell_guid="4df5f118-994c-4d7d-2fa1-b8a1ed1a82ec" _uuid="b45c7ca882bf32568268465f041282cfcd0e6fe9"
# ### Now the given problem is a classification problem.. Thus we will be using the classification algorithms to build a model.
# **Classification**: samples belong to two or more classes and we want to learn from already labeled data how to predict the class of unlabeled data
#
# **Regression**: if the desired output consists of one or more continuous variables, then the task is called regression. An example of a regression problem would be the prediction of the length of a salmon as a function of its age and weight.
# + [markdown] _cell_guid="b772ca1b-fd3c-d27a-b787-926c2df4e354" _uuid="a0436d5a5baa07027fdc10ad918d776e7fe948f4"
# Before we start, we need to clear some ML notations.
#
# **attributes**-->An attribute is a property of an instance that may be used to determine its classification. In the following dataset, the attributes are the petal and sepal length and width. It is also known as **Features**.
#
# **Target variable**, in the machine learning context is the variable that is or should be the output. Here the target variables are the 3 flower species.
# + _cell_guid="c27e7e16-6083-5b53-cda4-c43cf4c79a67" _execution_state="idle" _uuid="2b652b2398f95bacb6286eac49694cd422a54e6a"
# importing alll the necessary packages to use the various classification algorithms
from sklearn.linear_model import LogisticRegression # for Logistic Regression algorithm
from sklearn.cross_validation import train_test_split #to split the dataset for training and testing
from sklearn.neighbors import KNeighborsClassifier # for K nearest neighbours
from sklearn import svm #for Support Vector Machine (SVM) Algorithm
from sklearn import metrics #for checking the model accuracy
from sklearn.tree import DecisionTreeClassifier #for using Decision Tree Algoithm
# + _cell_guid="d967de9a-df34-bc84-899b-28b8804f7d58" _execution_state="idle" _uuid="15457e7fa8ff19b86c2858c365f11fa289fca723"
iris.shape #get the shape of the dataset
# + [markdown] _cell_guid="21c81da1-f5f8-8d86-6c23-76928d0ec387" _uuid="a09827c482cacd56f2040afdb162d46270551512"
# Now, when we train any algorithm, the number of features and their correlation plays an important role. If there are features and many of the features are highly correlated, then training an algorithm with all the featues will reduce the accuracy. Thus features selection should be done carefully. This dataset has less featues but still we will see the correlation.
# + _cell_guid="91f4a3b2-6e38-4d41-9ba0-52c7f43a581c" _execution_state="idle" _uuid="86e4d3c42eeec9d2df63d5f60c94e2c30b65ebd7"
plt.figure(figsize=(7,4))
sns.heatmap(iris.corr(),annot=True,cmap='cubehelix_r') #draws heatmap with input as the correlation matrix calculted by(iris.corr())
plt.show()
# + [markdown] _cell_guid="45543c97-83ae-392c-183c-24d118985163" _uuid="a2456971023a8d78416ae87fd1b09173da1e357e"
# **Observation--->**
#
# The Sepal Width and Length are not correlated
# The Petal Width and Length are highly correlated
#
# We will use all the features for training the algorithm and check the accuracy.
#
# Then we will use 1 Petal Feature and 1 Sepal Feature to check the accuracy of the algorithm as we are using only 2 features that are not correlated. Thus we can have a variance in the dataset which may help in better accuracy. We will check it later.
# + [markdown] _cell_guid="74afdd84-2b1f-b6e3-b9e2-6c076e44cb10" _uuid="362675378d1f8d7892597877aa9dbd1caa20c0ff"
# ### Steps To Be followed When Applying an Algorithm
#
# 1. Split the dataset into training and testing dataset. The testing dataset is generally smaller than training one as it will help in training the model better.
# 2. Select any algorithm based on the problem (classification or regression) whatever you feel may be good.
# 3. Then pass the training dataset to the algorithm to train it. We use the **.fit()** method
# 4. Then pass the testing data to the trained algorithm to predict the outcome. We use the **.predict()** method.
# 5. We then check the accuracy by **passing the predicted outcome and the actual output** to the model.
# + [markdown] _cell_guid="2de179a6-ea22-00ff-8756-63d3aef6bd2d" _uuid="ffa8b5a66eb4651f46672b7263774dbcf125ecbf"
# ### Splitting The Data into Training And Testing Dataset
# + _cell_guid="a24c3ab9-8c7d-2a78-113f-d337c5c61f09" _execution_state="idle" _uuid="87bc90a957a8defb5e24b886b641fb0a9aa5fc42"
train, test = train_test_split(iris, test_size = 0.3)# in this our main data is split into train and test
# the attribute test_size=0.3 splits the data into 70% and 30% ratio. train=70% and test=30%
print(train.shape)
print(test.shape)
# + _cell_guid="54dd7ded-5079-fc8c-065b-70bfbfb0b83a" _execution_state="idle" _uuid="7a64738b8be79ad6b0f4899e985d3ab5e475f9e1"
train_X = train[['SepalLengthCm','SepalWidthCm','PetalLengthCm','PetalWidthCm']]# taking the training data features
train_y=train.Species# output of our training data
test_X= test[['SepalLengthCm','SepalWidthCm','PetalLengthCm','PetalWidthCm']] # taking test data features
test_y =test.Species #output value of test data
# + [markdown] _cell_guid="c7d21e9f-8470-9aae-51ef-feea2cec6b1c" _uuid="0b18f7458fd76e93ed4a3932001f6f657cadb647"
# Lets check the Train and Test Dataset
# + _cell_guid="2cf23cff-74d0-b15e-87a9-5ed72f543380" _execution_state="idle" _uuid="8fa3e8f9f4ba0b27796008dd998c41670f1ffd95"
train_X.head(2)
# + _cell_guid="ce373f86-18ae-095f-8629-5bb8307c1273" _execution_state="idle" _uuid="ad5d8e583245a888c9470ef155b7d69a5608d426"
test_X.head(2)
# + _cell_guid="60f7b821-4335-e2e5-b868-3d6f850ea927" _execution_state="idle" _uuid="6e5039b56dec20594cdd457127d39c0009f5572a"
train_y.head() ##output of the training data
# + [markdown] _cell_guid="82010322-d6f3-467d-f1da-4cb3fc82d0bd" _uuid="41536ca3ef126cf761c2b8aca8f02ffa407a5996"
# ### Support Vector Machine (SVM)
# + _cell_guid="be869394-0f6d-f062-dd1f-8d2c68f06104" _execution_state="idle" _uuid="94ed6a0a8a1acc6c42e37db3e50ab6f9380e17f8"
model = svm.SVC() #select the algorithm
model.fit(train_X,train_y) # we train the algorithm with the training data and the training output
prediction=model.predict(test_X) #now we pass the testing data to the trained algorithm
print('The accuracy of the SVM is:',metrics.accuracy_score(prediction,test_y))#now we check the accuracy of the algorithm.
#we pass the predicted output by the model and the actual output
# + [markdown] _cell_guid="78e50a4b-ea63-c546-a8c4-07f4e4084c27" _uuid="9e9fb51ffb0823fb9a7459ad0f164e52a800c15a"
# SVM is giving very good accuracy . We will continue to check the accuracy for different models.
#
# Now we will follow the same steps as above for training various machine learning algorithms.
# + [markdown] _cell_guid="35d98035-f0ae-7d10-fe90-ec13865c5a14" _uuid="a2d76e51ba5e61a1a6f19675295f4405c2633eba"
# ### Logistic Regression
# + _cell_guid="ca772378-d3c4-4c87-ada6-b9d50a383b01" _execution_state="idle" _uuid="2b53cac02465732a986be2463b3f3411ab043716"
model = LogisticRegression()
model.fit(train_X,train_y)
prediction=model.predict(test_X)
print('The accuracy of the Logistic Regression is',metrics.accuracy_score(prediction,test_y))
# + [markdown] _cell_guid="7dfab2de-ebb7-4864-675c-aafef45db7a1" _uuid="756183a2d33bfd269b75b11ce7980f7f22925004"
# ### Decision Tree
# + _cell_guid="f918247c-a76b-4c58-5145-1fb0e8ab70b4" _execution_state="idle" _uuid="3c8158a18da7910b9ba8c3a5169a3009fb27c84f"
model=DecisionTreeClassifier()
model.fit(train_X,train_y)
prediction=model.predict(test_X)
print('The accuracy of the Decision Tree is',metrics.accuracy_score(prediction,test_y))
# + [markdown] _cell_guid="3e4594d9-5fe6-0568-1fc6-e02975278305" _uuid="3410b37d9410b8d9f0ebd5c6509b3250ed7714c1"
# ### K-Nearest Neighbours
# + _cell_guid="bcd55de7-cfab-c81d-25a8-1db8f0deee94" _execution_state="idle" _uuid="dda9e73ae95627994e68db9542d769b897d43bec"
model=KNeighborsClassifier(n_neighbors=3) #this examines 3 neighbours for putting the new data into a class
model.fit(train_X,train_y)
prediction=model.predict(test_X)
print('The accuracy of the KNN is',metrics.accuracy_score(prediction,test_y))
# + [markdown] _cell_guid="e6787513-11b0-5cdb-10ac-457292de9b55" _uuid="9cd754bcd0f4494014b69c4fe11d3b228b5eee3f"
# ### Let's check the accuracy for various values of n for K-Nearest nerighbours
# + _cell_guid="2c79e403-480a-0620-dd84-01ca038c72bd" _execution_state="idle" _uuid="abe2f8b04c14ad0618c13edd1777e7a8c2985781"
a_index=list(range(1,11))
a=pd.Series()
x=[1,2,3,4,5,6,7,8,9,10]
for i in list(range(1,11)):
model=KNeighborsClassifier(n_neighbors=i)
model.fit(train_X,train_y)
prediction=model.predict(test_X)
a=a.append(pd.Series(metrics.accuracy_score(prediction,test_y)))
plt.plot(a_index, a)
plt.xticks(x)
plt.show()
# + [markdown] _cell_guid="de478d66-f9c6-6382-4a89-05c159bf657f" _uuid="0f1e3fad4a5403e758e4a3766c7887c65c9ab24b"
# Above is the graph showing the accuracy for the KNN models using different values of n.
# + [markdown] _cell_guid="cd3deccf-1025-320b-a269-40a99c7dd3f5" _uuid="8ad86c41920a005f238e20d4a84585079d558d16"
# ### We used all the features of iris in above models. Now we will use Petals and Sepals Seperately
# + [markdown] _cell_guid="74a20365-8410-1b88-5f76-522c73c39a09" _uuid="a5b27b547f5c0ad1ec8405aec1117e0e48180633"
# ### Creating Petals And Sepals Training Data
# + _cell_guid="80dc70f3-ad29-fc83-27d7-f378a32f0a46" _execution_state="idle" _uuid="a5c6805e5ea06a4f1c9bf2b4879e609b4951780b"
petal=iris[['PetalLengthCm','PetalWidthCm','Species']]
sepal=iris[['SepalLengthCm','SepalWidthCm','Species']]
# + _cell_guid="7048f50d-5ad5-3c48-3eaa-26a0f5bcb034" _execution_state="idle" _uuid="70b5ef63a17330a3d1f1d918831407bdf535e2bf"
train_p,test_p=train_test_split(petal,test_size=0.3,random_state=0) #petals
train_x_p=train_p[['PetalWidthCm','PetalLengthCm']]
train_y_p=train_p.Species
test_x_p=test_p[['PetalWidthCm','PetalLengthCm']]
test_y_p=test_p.Species
train_s,test_s=train_test_split(sepal,test_size=0.3,random_state=0) #Sepal
train_x_s=train_s[['SepalWidthCm','SepalLengthCm']]
train_y_s=train_s.Species
test_x_s=test_s[['SepalWidthCm','SepalLengthCm']]
test_y_s=test_s.Species
# + [markdown] _cell_guid="c7bc7029-2397-93a9-e063-afb7c129af8d" _uuid="05ed84baa609e7379bcb84bdad50c60650b86e49"
# ### SVM
# + _cell_guid="bf8b63a2-7f74-f2d3-3f42-fb480ae26139" _execution_state="idle" _uuid="2abd62b140e7918b900c05400372ff1b5f1b2e97"
model=svm.SVC()
model.fit(train_x_p,train_y_p)
prediction=model.predict(test_x_p)
print('The accuracy of the SVM using Petals is:',metrics.accuracy_score(prediction,test_y_p))
model=svm.SVC()
model.fit(train_x_s,train_y_s)
prediction=model.predict(test_x_s)
print('The accuracy of the SVM using Sepal is:',metrics.accuracy_score(prediction,test_y_s))
# + [markdown] _cell_guid="4fd9e81b-8aad-1e15-8b88-819112c17071" _uuid="1fea5c6c187d41fe71248597fff47e49621fd83d"
# ### Logistic Regression
# + _cell_guid="8353f86a-d1fe-6f07-7eb5-4f651eba1197" _execution_state="idle" _uuid="f732c0dc0789acd023476ed80107c7a6f4ddeb71"
model = LogisticRegression()
model.fit(train_x_p,train_y_p)
prediction=model.predict(test_x_p)
print('The accuracy of the Logistic Regression using Petals is:',metrics.accuracy_score(prediction,test_y_p))
model.fit(train_x_s,train_y_s)
prediction=model.predict(test_x_s)
print('The accuracy of the Logistic Regression using Sepals is:',metrics.accuracy_score(prediction,test_y_s))
# + [markdown] _cell_guid="d8905552-47ed-3654-6dcc-a9603fe2cd12" _uuid="8fff15cad2855cedd186d19af2e419c79a5e3e4f"
# ### Decision Tree
# + _cell_guid="1383aeef-0009-c16c-3f2c-97d400de6282" _execution_state="idle" _uuid="973393361c06902445a5e8a2a6a1a3c3acdec967"
model=DecisionTreeClassifier()
model.fit(train_x_p,train_y_p)
prediction=model.predict(test_x_p)
print('The accuracy of the Decision Tree using Petals is:',metrics.accuracy_score(prediction,test_y_p))
model.fit(train_x_s,train_y_s)
prediction=model.predict(test_x_s)
print('The accuracy of the Decision Tree using Sepals is:',metrics.accuracy_score(prediction,test_y_s))
# + [markdown] _cell_guid="0d1fcf97-a603-ef57-5085-fab9b931c186" _uuid="52cb931113963aa7790c83c8427f49e03f109a65"
# ### K-Nearest Neighbours
# + _cell_guid="89fc030a-1c30-ee0d-7c62-7495fa476dba" _execution_state="idle" _uuid="122bd22d9a7e63f0e1f86be80a2d2dde82182536"
model=KNeighborsClassifier(n_neighbors=3)
model.fit(train_x_p,train_y_p)
prediction=model.predict(test_x_p)
print('The accuracy of the KNN using Petals is:',metrics.accuracy_score(prediction,test_y_p))
model.fit(train_x_s,train_y_s)
prediction=model.predict(test_x_s)
print('The accuracy of the KNN using Sepals is:',metrics.accuracy_score(prediction,test_y_s))
# + [markdown] _cell_guid="752b273a-43f6-9905-c42b-0a3003ad204c" _uuid="453bb5a7b5ee35d4dcbac52b6ee717d61d578c7f"
# ### Observations:
#
# - Using Petals over Sepal for training the data gives a much better accuracy.
# - This was expected as we saw in the heatmap above that the correlation between the Sepal Width and Length was very low whereas the correlation between Petal Width and Length was very high.
# + [markdown] _cell_guid="6e6e1e3a-07aa-201a-b959-1d3619103325" _uuid="0f45b63a88d023709700b04fd14605ae53c94550"
# Thus we have just implemented some of the common Machine Learning. Since the dataset is small with very few features, I didn't cover some concepts as they would be relevant when we have many features.
#
# I have compiled a notebook covering some advanced ML concepts using a larger dataset. Have a look at that tooo.
# + [markdown] _cell_guid="66bef947-c001-01e6-3064-cec367662f19" _execution_state="idle" _uuid="f561aab5cae104858575057eb7f93e7e8abaa77e"
# I hope the notebook was useful to you to get started with Machine Learning.
#
# If find this notebook, **Please Upvote**.
#
# Thank You!!
# + _cell_guid="14bcdd2b-fc7c-42e2-88c7-ed5d221ecbef" _execution_state="idle" _uuid="41c25d25019131c90859443d37b93d8fbb0fb33f"
| public-engines/iris-species-engine/notebooks/ash316-kaggle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # SQL Parameters
#
# Currently BigQuery SQL does not support parameterization. However, within notebooks, it is quite interesting to be able to use Python variables defined in the notebook as parameter values for SQL.
#
# Cloud Datalab introduces a pattern for declaring and using parameterized queries.
# ## Data Preview
# %%bigquery sample --count 10
SELECT * FROM [cloud-datalab-samples:httplogs.logs_20140615]
# + language="sql"
# SELECT endpoint FROM [cloud-datalab-samples:httplogs.logs_20140615] GROUP BY endpoint
# -
# # Parameterization via SQL Modules
#
# Parameters are declared in SQL modules using a `name = default_value` syntax before the SQL, and then using `$name` within the SQL to reference the parameter.
# + magic_args="--module endpoint_stats" language="sql"
# endpoint = 'Other'
#
# SELECT endpoint, COUNT(latency) As requests, MIN(latency) AS min_latency, MAX(latency) AS max_latency
# FROM [cloud-datalab-samples:httplogs.logs_20140615]
# WHERE endpoint = $endpoint
# GROUP BY endpoint
# -
# This just defined a SQL query with a String `name` parameter named endpoint, which defaults to the value Other, as you'll see when the query is used to sample data without specifying a specific value.
# %%bigquery execute --query endpoint_stats
# ## Declarative Query Execution
# Parameter values can be specified with a `%%bigquery sample` command as follows (parameter values defined in a YAML block):
# %%bigquery execute --query endpoint_stats
endpoint: Recent
# The YAML text can reference values defined in the notebook as well, using again, the `$variable` syntax.
interesting_endpoint = 'Popular'
# %%bigquery execute --query endpoint_stats
endpoint: $interesting_endpoint
# ## Imperative Query Execution
#
# Parameter values can be passed to BigQuery APIs when constructing a `Query` object.
import gcp.bigquery as bq
stats_query = bq.Query(endpoint_stats, endpoint = interesting_endpoint)
print stats_query.sql
# From the SQL above, you can see above the value for the `$endpoint` variable was expanded out. The parameter replacement happens locally, before the resulting SQL is sent to BigQuery.
stats_query.results()
# # Looking Ahead
#
# Parameterization enables one-half of SQL and Python integration - being able to use values in Python code, in the notebook, and passing them in as part of the query when retrieving data from BigQuery.
#
# The next notebook will cover the second-half of SQL and Python integration - retrieving query results into the notebook for use with Python code.
#
# Parameterization is also the building block toward creating complex queries, where whole queries can be used as parameter values.
| docs/tutorials/BigQuery/SQL Parameters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
import os
import warnings
def warn(*args, **kwargs):
pass
warnings.warn = warn
warnings.simplefilter(action='ignore', category=FutureWarning)
import gym
import numpy as np
from tensorforce.agents import Agent
from tensorforce.execution import Runner
from tensorforce.contrib.openai_gym import OpenAIGym
sys.path.append(os.path.dirname(os.path.abspath('')))
from tensortrade.environments import TradingEnvironment
from tensortrade.exchanges.live import CCXTExchange
from tensortrade.actions import DiscreteActionStrategy
from tensortrade.rewards import SimpleProfitStrategy
# +
# Defining the Exchange
coinbase = ccxt.coinbasepro()
exchange = CCXTExchange(exchange=coinbase, base_instrument='USD')
# defining the action strategy
action_strategy = DiscreteActionStrategy()
# Defining the reward strategy
reward_strategy = SimpleProfitStrategy()
# configuring the trading environment
env = TradingEnvironment(exchange=exchange,
action_strategy=action_strategy,
reward_strategy=reward_strategy,
feature_pipeline=None)
# +
# Defining the feature pipeline and also seting it up with the coinbase exchange.
from tensortrade.features import FeaturePipeline
from tensortrade.features.scalers import MinMaxNormalizer
from tensortrade.features.stationarity import FractionalDifference
from tensortrade.features.indicators import SimpleMovingAverage
price_columns = ["open", "high", "low", "close"]
normalize_price = MinMaxNormalizer(price_columns)
moving_averages = SimpleMovingAverage(price_columns)
difference_all = FractionalDifference(difference_order=0.6)
feature_pipeline = FeaturePipeline(steps=[normalize_price,
moving_averages,
difference_all])
# setting the exchange
exchange.feature_pipeline = feature_pipeline
# +
# Defining the agent
agent_config = {
"type": "dqn_agent",
"update_mode": {
"unit": "timesteps",
"batch_size": 64,
"frequency": 4
},
"memory": {
"type": "replay",
"capacity": 10000,
"include_next_states": True
},
"optimizer": {
"type": "clipped_step",
"clipping_value": 0.1,
"optimizer": {
"type": "adam",
"learning_rate": 1e-3
}
},
"discount": 0.999,
"entropy_regularization": None,
"double_q_model": True,
"target_sync_frequency": 1000,
"target_update_weight": 1.0,
"actions_exploration": {
"type": "epsilon_anneal",
"initial_epsilon": 0.5,
"final_epsilon": 0.,
"timesteps": 1000000000
},
"saver": {
"directory": None,
"seconds": 600
},
"summarizer": {
"directory": None,
"labels": ["graph", "total-loss"]
},
"execution": {
"type": "single",
"session_config": None,
"distributed_spec": None
}
}
# +
# Defining the network specifications
network_spec = [
dict(type='dense', size=64),
dict(type='dense', size=32)
]
agent = Agent.from_spec(
spec=agent_config,
kwargs=dict(
states=env.states,
actions=env.actions,
network=network_spec,
)
)
# +
# Create the runner
runner = Runner(agent=agent, environment=env)
# +
# Callback function printing episode statistics
def episode_finished(r):
print("Finished episode {ep} after {ts} timesteps (reward: {reward})".format(ep=r.episode, ts=r.episode_timestep,
reward=r.episode_rewards[-1]))
return True
# +
# Start learning
runner.run(episodes=300, max_episode_timesteps=10000, episode_finished=episode_finished)
runner.close()
# Print statistics
print("Learning finished. Total episodes: {ep}. Average reward of last 100 episodes: {ar}.".format(
ep=runner.episode,
ar=np.mean(runner.episode_rewards))
)
# -
| examples/bitcoin_trading Agent using tensortrade.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _cell_guid="63f97ada-21c8-4755-9fee-0a0cfbff8aa7" _uuid="9853586a0dc75ce39e7c7ffcde1eb4d47c6fb02e"
# # Overview ##
#
# This notebook creates numerous models, eventually settling on a simple yet effective logistic regression model. The models are trained on the seed differences between teams and season average metric differences (e.g., FG%, PPG, Opp. PPG) between teams.
#
# Note that the model is trained entirely on data from 2003-2017 and their known outcomes. The resulting classifier is then used on 2018 data to generate predictions for this year's tournament on March 11.
# + _cell_guid="0c233e05-c63d-4866-96dc-bb38d444bf84" _uuid="5464dc4b196dc4c8dd0323bbd71b75724113e2af"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
from sklearn.linear_model import LogisticRegression
from sklearn.neural_network import MLPClassifier
from sklearn import svm
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
import matplotlib.pyplot as plt
from sklearn.utils import shuffle
from sklearn.model_selection import GridSearchCV
# + [markdown] _cell_guid="9eecd909-c6e5-4a88-8481-bcbca7aef1df" _uuid="819472385a23f3fd5aaf4172b4f8db227cf5271f"
# ## Load the training data ##
# We're keeping it relatively simple & using only a handful of files for this model: the tourney seeds, tourney results, and a detailed results dataset to calculate our other features.
# + _cell_guid="087d26ad-591c-4ff4-bd13-be6aaf436832" _uuid="bf8ee168a0372e883332d6bb0ce5c89c13143650"
data_dir = '../input/'
df_seeds = pd.read_csv(data_dir + 'NCAATourneySeeds.csv')
df_tour = pd.read_csv(data_dir + 'NCAATourneyCompactResults.csv')
# We load detailed season data to calculate season average statistics for each team
df_reg_season_detailed = pd.read_csv(data_dir + 'RegularSeasonDetailedResults.csv')
df_reg_season_detailed.drop(labels=['WFGM3', 'WFGA3', 'WFTM', 'WFTA', 'WDR', 'WAst',
'WStl', 'WBlk', 'WPF', 'LFGM3', 'LFGA3', 'LFTM', 'LFTA', 'LDR',
'LAst', 'LStl', 'LBlk', 'LPF', 'WLoc', 'NumOT', 'WOR', 'LOR'],
inplace=True, axis=1)
df_reg_season_detailed.head()
# -
# ## Create a new data frame with season average metrics ##
# We are creating a new data frame with season average statistics for each team for use as features in our machine learning algorithm.
# +
yearList = range(2003,2019) #2003 is the first year we have detailed data for
teams_pd = pd.read_csv(data_dir + 'Teams.csv')
teamIDs = teams_pd['TeamID'].tolist()
rows = list()
for year in yearList:
for team in teamIDs:
df_curr_season = df_reg_season_detailed[df_reg_season_detailed.Season == year]
df_curr_team_wins = df_curr_season[df_curr_season.WTeamID == team]
df_curr_team_losses = df_curr_season[df_curr_season.LTeamID == team]
# no games played by them this year.. skip (current team didn't win or lose any games)
if df_curr_team_wins.shape[0] == 0 and df_curr_team_losses.shape[0] == 0:
continue;
df_winteam = df_curr_team_wins.rename(columns={'WTeamID':'TeamID', 'WFGM':'FGM',
'WFGA':'FGA', 'WTO':'TO', 'WScore':'Score', 'LScore':'OppScore'})
# drop all columns except the ones we are using
df_winteam = df_winteam[['TeamID', 'FGM', 'FGA', 'TO', 'Score', 'OppScore']]
df_loseteam = df_curr_team_losses.rename(columns={'LTeamID':'TeamID', 'LFGM':'FGM',
'LFGA':'FGA', 'LTO':'TO', 'LScore':'Score', 'WScore':'OppScore'})
# drop all columns except the ones we are using
df_loseteam = df_loseteam[['TeamID', 'FGM', 'FGA', 'TO', 'Score', 'OppScore']]
# dataframe w/ all relevant stats from current year for current team
df_curr_team = pd.concat((df_winteam, df_loseteam))
wins = df_winteam.shape[0]
FGPercent = df_curr_team['FGM'].sum() / df_curr_team['FGA'].sum()
TurnoverAvg = df_curr_team['TO'].sum() / len(df_curr_team['TO'].values)
PPG = df_curr_team['Score'].sum() / len(df_curr_team['Score'].values)
OppPPG = df_curr_team['OppScore'].sum() / len(df_curr_team['OppScore'].values)
# collect all data in rows list first for effeciency
rows.append([year, team, wins, FGPercent, TurnoverAvg, PPG, OppPPG])
df_training_data = pd.DataFrame(rows, columns=['Season', 'TeamID', 'Wins', 'FGPercent',
'TOAvg', 'PPG', 'OppPPG'])
df_training_data.head()
# -
# Here we show the contents of our seeding and tournament results data frames. These, combined with the stats calculated above (df_training_data) will form the final X_train matrix.
# + _cell_guid="e5412069-c89b-4fed-9d2d-4690d6fd71b4" _uuid="9f32e5f9104b7f10d3de7b38d3f292aef045c30f"
df_seeds.head()
# + _cell_guid="3eaeb447-2e68-4790-9bcd-423ccdb7a117" _uuid="dcb3b4cc84f09ea5af4d52da4fd970928e14bfc1"
df_tour.head()
# + [markdown] _cell_guid="ac5dd6af-b871-47c6-b7b0-ef6d85954971" _uuid="42f99f53dd385e23b09378e0de9d3fce5eb1a2e9"
# First, we'll simplify the datasets to remove the columns we won't be using and convert the seedings to the needed format (stripping the regional abbreviation in front of the seed).
# + _cell_guid="4e397ac8-7ac8-4ba7-b92f-571a0e75da18" _uuid="fcb18269a41cfa257bd97c40664e43e701251bed"
def seed_to_int(seed):
#Get just the digits from the seeding. Return as int
s_int = int(seed[1:3])
return s_int
df_seeds['seed_int'] = df_seeds.Seed.apply(seed_to_int)
df_seeds.drop(labels=['Seed'], inplace=True, axis=1) # This is the string label
df_seeds.head()
# + _cell_guid="90db3f9f-7d11-4bac-8d37-3331e4416c6a" _uuid="1f6ecb82fa587f5a95a6833cd224b01407f5c90a"
df_tour.drop(labels=['DayNum', 'WScore', 'LScore', 'WLoc', 'NumOT'], inplace=True, axis=1)
df_tour.head()
# + [markdown] _cell_guid="88e7ab21-0380-42b2-9979-e30fef6e856a" _uuid="3f223cdf4446d6e9c77ab8319237f05393d1a822"
# ## Merge seed for each team ##
# Merge the Seeds with their corresponding TeamIDs in the compact results dataframe.
# + _cell_guid="a22a595b-a6cb-4291-81ae-903a9548cd37" _uuid="53638c1ae27cfb24d47e02007c293d5ee19ebdac"
df_winseeds = df_seeds.rename(columns={'TeamID':'WTeamID', 'seed_int':'WSeed'})
df_lossseeds = df_seeds.rename(columns={'TeamID':'LTeamID', 'seed_int':'LSeed'})
df_dummy = pd.merge(left=df_tour, right=df_winseeds, how='left', on=['Season', 'WTeamID'])
df_concat = pd.merge(left=df_dummy, right=df_lossseeds, on=['Season', 'LTeamID'])
df_concat['SeedDiff'] = df_concat.WSeed - df_concat.LSeed
df_concat.head()
# -
# Now we'll combine our advanced season statistics and merge them into the df_concat data frame.
# +
df_winstats = df_training_data.rename(columns={'TeamID':'WTeamID', 'FGPercent':'WFGPercent',
'TOAvg':'WTOAvg', 'PPG':'WPPG', 'OppPPG':'WOppPPG', 'Wins':'WWins'})
df_lossstats = df_training_data.rename(columns={'TeamID':'LTeamID', 'FGPercent':'LFGPercent',
'TOAvg':'LTOAvg', 'PPG':'LPPG', 'OppPPG':'LOppPPG', 'Wins':'LWins'})
df_dummy = pd.merge(left=df_concat, right=df_winstats, on=['Season', 'WTeamID'])
df_concat = pd.merge(left=df_dummy, right=df_lossstats, on=['Season', 'LTeamID'])
df_concat['FGPercentDiff'] = df_concat.WFGPercent - df_concat.LFGPercent
df_concat['TOAvgDiff'] = df_concat.WTOAvg - df_concat.LTOAvg
df_concat['PPGDiff'] = df_concat.WPPG - df_concat.LPPG
df_concat['OppPPGDiff'] = df_concat.WOppPPG - df_concat.LOppPPG
df_concat['WWinMargin'] = df_concat.WPPG - df_concat.WOppPPG
df_concat['LWinMargin'] = df_concat.LPPG - df_concat.LOppPPG
df_concat['WinMarginDiff'] = df_concat.WWinMargin - df_concat.LWinMargin
df_concat['WinDiff'] = df_concat.WWins - df_concat.LWins
# drop all columns except the ones we are using
df_concat = df_concat[['Season', 'WTeamID', 'LTeamID', 'SeedDiff', 'FGPercentDiff',
'TOAvgDiff', 'PPGDiff', 'OppPPGDiff', 'WinMarginDiff', 'WinDiff']]
# Note: We can have SeedDiff == 0 due to the First Four (68 teams)! Also Final Four onwards!
# Note: Pandas merges tossed out data from before 2003!
df_concat.head()
# + [markdown] _cell_guid="72274b95-581c-4938-88a4-b3e8cb1787d4" _uuid="1c82f60c02545c8c46ab090cb8cefca48e48e434"
# Now we'll create a dataframe that summarizes wins & losses along with their corresponding seed differences, FG% differences, and turnover differences. This is the meat of what we'll be creating our model on.
# + _cell_guid="4279fa78-4700-43d6-a92e-3a372715e0f3" _uuid="1a40000e85c0dd9d2be6850a767acd736bf5f182"
# We create positive and negative versions of the data so the
# supervised learning algorithm has sample data of each class to classify
df_wins = pd.DataFrame()
df_wins['SeedDiff'] = df_concat['SeedDiff']
df_wins['FGPercentDiff'] = df_concat['FGPercentDiff']
df_wins['TOAvgDiff'] = df_concat['TOAvgDiff']
df_wins['PPGDiff'] = df_concat['PPGDiff']
df_wins['OppPPGDiff'] = df_concat['OppPPGDiff']
df_wins['WinMarginDiff'] = df_concat['WinMarginDiff']
df_wins['WinDiff'] = df_concat['WinDiff']
df_wins['Result'] = 1
df_losses = pd.DataFrame()
df_losses['SeedDiff'] = -df_concat['SeedDiff']
df_losses['FGPercentDiff'] = -df_concat['FGPercentDiff']
df_losses['TOAvgDiff'] = -df_concat['TOAvgDiff']
df_losses['PPGDiff'] = -df_concat['PPGDiff']
df_losses['OppPPGDiff'] = -df_concat['OppPPGDiff']
df_losses['WinMarginDiff'] = -df_concat['WinMarginDiff']
df_losses['WinDiff'] = -df_concat['WinDiff']
df_losses['Result'] = 0
df_predictions = pd.concat((df_wins, df_losses))
df_predictions.head()
# + _cell_guid="e6eca4b5-2b22-4abf-9145-7a8d284faab7" _uuid="3cf1b39303c44e73a3fa0f813a9580e91eca6b0b"
X_train = [list(a) for a in zip(df_predictions.SeedDiff.values, df_predictions.FGPercentDiff.values,
df_predictions.TOAvgDiff.values, df_predictions.PPGDiff.values,
df_predictions.OppPPGDiff.values, df_predictions.WinMarginDiff.values,
df_predictions.WinDiff.values)]
X_train = np.array(X_train)
y_train = df_predictions.Result.values
X_train, y_train = shuffle(X_train, y_train)
# + [markdown] _cell_guid="4b0751b0-a04a-4586-b612-6feb201cde7d" _uuid="563937f42bcccd2bbfb8fc1a66a72a9ca1351f43"
# ## Train the model ##
# Train the a variety of models. Tune the hyperparameters for each algorithm and perform cross validation. Logistic regression and SVC perform the best.
# + _cell_guid="34a69207-0b92-4b28-8ad7-5d2cabec878a" _uuid="95f817451eae9b72dc237e734e19c929be136d50"
# Neural Network
params = {'hidden_layer_sizes': [(256,), (512,), (128, 256, 128,)]}
mlp = MLPClassifier(learning_rate='adaptive')
clf = GridSearchCV(mlp, params, scoring='neg_log_loss')
clf.fit(X_train, y_train)
print('Best log_loss Multi Layer Perceptron Classifier: {}'.format(clf.best_score_))
# Gradient Boosted Classifier
GBC = GradientBoostingClassifier()
param_grid_GBC = {
"n_estimators" : [100],
"learning_rate" : [0.1, 0.05, 0.02, 0.01],
"max_depth" : [1,2,3],
"min_samples_leaf" : [1,3,5],
"max_features" : [1.0, 0.3, 0.1]
}
clf = GridSearchCV(GBC, param_grid_GBC, scoring='neg_log_loss')
clf.fit(X_train, y_train)
print('Best log_loss Gradient Boosting Classifier: {}'.format(clf.best_score_))
# Random Forest Classifier
RFC = RandomForestClassifier()
param_grid_RFC = {
'n_estimators': [60, 120, 240],
'max_features': ['auto', 'sqrt', 'log2']
}
clf = GridSearchCV(RFC, param_grid_RFC, scoring='neg_log_loss')
clf.fit(X_train, y_train)
print('Best log_loss Random Forest Classifier: {}'.format(clf.best_score_))
# K Nearest Neighbors Classifier
knn = KNeighborsClassifier()
k = np.arange(80)+1
parameters = {'n_neighbors': k}
clf = GridSearchCV(knn, parameters, scoring='neg_log_loss')
clf.fit(X_train, y_train)
print('Best log_loss K-Nearest Neighbors Classifier: {}'.format(clf.best_score_))
# SVC
SVC = svm.SVC(probability=True)
tuned_parameters = [{'kernel': ['rbf'], 'gamma': [1e-3, 1e-4],
'C': [1, 10, 100, 1000]},
{'kernel': ['linear'], 'C': [1, 10, 100, 1000]}]
tuned_parameters_preselected = [{'kernel': ['linear'], 'C': [10]}]
clf = GridSearchCV(SVC, tuned_parameters_preselected, scoring='neg_log_loss')
clf.fit(X_train, y_train)
print('Best log_loss Support Vector Classification: {}'.format(clf.best_score_))
# Logistic Regression
logreg = LogisticRegression()
params = {'C': np.logspace(start=-15, stop=15, num=31)} # {C: array[1^-15 , 1^-14, ... 1^15] }
clf = GridSearchCV(logreg, params, scoring='neg_log_loss', refit=True) #sklearn model selection
clf.fit(X_train, y_train)
print('Best log_loss Logistic Regression: {}, with best C: {}'.format(clf.best_score_,
clf.best_params_['C']))
# Logistic Regression is typically the top-performer. We compute it last, and use
# this classifier to make future predictions.
# SVC is typically a close second. Comment out Logistic Regression to use
# the SVC classifier instead to make future predictions
# Keep in mind, the provided values are a single representation of our classifier's
# success! Depending on how the data is shuffled, each run of the program may yield
# a slightly different classifier (and thus different predictions/success rate)
# + _cell_guid="2bf46a87-8bbe-4964-935a-89e307f700b9" _uuid="37e01a2b50f69e1f6a0aeaa50c7593d8cae15b1b"
# Create training data with the seeds varying from -10, 10
# All other features are zeroed out so the plot only shows
# the relationship between seed and P(team1 wins)
X1 = np.arange(-10, 10)
X2 = np.zeros(20, dtype=np.int)
X = [list(a) for a in zip(X1, X2, X2, X2, X2, X2, X2)]
X = np.array(X)
preds = clf.predict_proba(X)[:,1]
plt.plot(X1, preds)
plt.xlabel('Team1 seed - Team2 seed')
plt.ylabel('P(Team1 will win)')
# + [markdown] _cell_guid="2e9a55cf-b938-426f-b92c-e92ded0a663f" _uuid="3e8270e8638b6f78317b7f787cc0259af682dba7"
# Plotting validates our intuition, that the probability a team will win decreases as the seed differential to its opponent decreases.
# + _cell_guid="16e4b2a7-91a4-420f-9eee-abaa49ea028b" _uuid="cd5a427eca09adda4e9a42a88208b683020a1f8d"
df_sample_sub = pd.read_csv(data_dir + 'SampleSubmissionStage2.csv')
n_test_games = len(df_sample_sub)
def get_year_t1_t2(ID):
"""Return a tuple with ints `year`, `team1` and `team2`."""
return (int(x) for x in ID.split('_'))
# -
# Now we create our X_test matrix with the expected dimensions for the Kaggle contest and fill it with zeroes. Then we loop over the sample submission, and initialize X_test with the correct features for 2018 teams. This X_test matrix is our test set for our previously trained classifier to make predictions about this year's tournament.
# + _cell_guid="3d842b3e-783b-4ab5-94c5-97cedc9d08c4" _uuid="72d64ebc20c903660108ae9c529be07859396909"
X_test = np.zeros(shape=(n_test_games, 7))
for ii, row in df_sample_sub.iterrows():
year, t1, t2 = get_year_t1_t2(row.ID)
t1_seed = df_seeds[(df_seeds.TeamID == t1) & (df_seeds.Season == year)].seed_int.values[0]
t2_seed = df_seeds[(df_seeds.TeamID == t2) & (df_seeds.Season == year)].seed_int.values[0]
diff_seed = t1_seed - t2_seed
X_test[ii, 0] = diff_seed
t1_FGPercent = df_training_data[(df_training_data.TeamID == t1) &
(df_training_data.Season == year)].FGPercent.values[0]
t2_FGPercent = df_training_data[(df_training_data.TeamID == t2) &
(df_training_data.Season == year)].FGPercent.values[0]
diff_FGPercent = t1_FGPercent - t2_FGPercent
X_test[ii, 1] = diff_FGPercent
t1_TOAvg = df_training_data[(df_training_data.TeamID == t1) &
(df_training_data.Season == year)].TOAvg.values[0]
t2_TOAvg = df_training_data[(df_training_data.TeamID == t2) &
(df_training_data.Season == year)].TOAvg.values[0]
diff_TOAvg = t1_TOAvg - t2_TOAvg
X_test[ii, 2] = diff_TOAvg
t1_PPG = df_training_data[(df_training_data.TeamID == t1) &
(df_training_data.Season == year)].PPG.values[0]
t2_PPG = df_training_data[(df_training_data.TeamID == t2) &
(df_training_data.Season == year)].PPG.values[0]
diff_PPG = t1_PPG - t2_PPG
X_test[ii, 3] = diff_PPG
t1_OppPPG = df_training_data[(df_training_data.TeamID == t1) &
(df_training_data.Season == year)].OppPPG.values[0]
t2_OppPPG = df_training_data[(df_training_data.TeamID == t2) &
(df_training_data.Season == year)].OppPPG.values[0]
diff_OppPPG = t1_OppPPG - t2_OppPPG
X_test[ii, 4] = diff_OppPPG
X_test[ii, 5] = diff_PPG - diff_OppPPG # Win Margin
t1_Wins = df_training_data[(df_training_data.TeamID == t1) &
(df_training_data.Season == year)].Wins.values[0]
t2_Wins = df_training_data[(df_training_data.TeamID == t2) &
(df_training_data.Season == year)].Wins.values[0]
X_test[ii, 6] = t1_Wins - t2_Wins
# + [markdown] _cell_guid="8bacc197-297f-465d-a49c-9b988619c166" _uuid="375748512c55520e00ffd5701c82704856478370"
# ## Make Predictions ##
# Create predictions using the logistic regression model we trained.
# + _cell_guid="b3da5070-e0e3-445a-9af1-ced10032bcc7" _uuid="65dc063a2e9c5e447d800556f7cf67b26b7cbedb"
preds = clf.predict_proba(X_test)[:,1]
clipped_preds = np.clip(preds, 0.05, 0.95)
df_sample_sub.Pred = clipped_preds
df_sample_sub.head()
# + [markdown] _cell_guid="72494058-d8c5-4ab4-99cd-4b50d7d7135c" _uuid="3f4ef6ab893953a811462d240778205c2fdecf97"
# Lastly, create your submission file!
# + _cell_guid="78ded09c-4ee1-49a2-901b-0cbe2ac3f114" _uuid="7c784a9b62d889e83493b70efa17bd233f9abff4"
df_sample_sub.to_csv('predictions.csv', index=False)
| nbs/.ipynb_checkpoints/script-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="-0YeSLuH2aP5"
# # "Poleval 2021 through wav2vec2"
# > "Trying for pronunciation recovery"
#
# - toc: false
# - branch: master
# - comments: true
# - hidden: true
# - categories: [wav2vec2, poleval, colab]
#
# + id="DiMBByvsGeKH"
# %%capture
# !pip install gdown
# + id="geNB2x34hjpU" colab={"base_uri": "https://localhost:8080/"} outputId="2cfd3b3d-1ee9-440c-b49d-442b208391b2"
# !gdown https://drive.google.com/uc?id=1b6MyyqgA9D1U7DX3Vtgda7f9ppkxjCXJ
# + id="BDrFENl1Tx3V"
# %%capture
# !tar zxvf poleval_wav.train.tar.gz && rm poleval_wav.train.tar.gz
# + id="uhVsVOlKwepr"
# %%capture
# !pip install librosa webrtcvad
# + id="a9oo0q7jkyao"
#collapse-hide
# VAD wrapper is taken from PyTorch Speaker Verification:
# https://github.com/HarryVolek/PyTorch_Speaker_Verification
# Copyright (c) 2019, HarryVolek
# License: BSD-3-Clause
# based on https://github.com/wiseman/py-webrtcvad/blob/master/example.py
# Copyright (c) 2016 <NAME>
# License: MIT
import collections
import contextlib
import numpy as np
import sys
import librosa
import wave
import webrtcvad
#from hparam import hparam as hp
sr = 16000
def read_wave(path, sr):
"""Reads a .wav file.
Takes the path, and returns (PCM audio data, sample rate).
Assumes sample width == 2
"""
with contextlib.closing(wave.open(path, 'rb')) as wf:
num_channels = wf.getnchannels()
assert num_channels == 1
sample_width = wf.getsampwidth()
assert sample_width == 2
sample_rate = wf.getframerate()
assert sample_rate in (8000, 16000, 32000, 48000)
pcm_data = wf.readframes(wf.getnframes())
data, _ = librosa.load(path, sr)
assert len(data.shape) == 1
assert sr in (8000, 16000, 32000, 48000)
return data, pcm_data
class Frame(object):
"""Represents a "frame" of audio data."""
def __init__(self, bytes, timestamp, duration):
self.bytes = bytes
self.timestamp = timestamp
self.duration = duration
def frame_generator(frame_duration_ms, audio, sample_rate):
"""Generates audio frames from PCM audio data.
Takes the desired frame duration in milliseconds, the PCM data, and
the sample rate.
Yields Frames of the requested duration.
"""
n = int(sample_rate * (frame_duration_ms / 1000.0) * 2)
offset = 0
timestamp = 0.0
duration = (float(n) / sample_rate) / 2.0
while offset + n < len(audio):
yield Frame(audio[offset:offset + n], timestamp, duration)
timestamp += duration
offset += n
def vad_collector(sample_rate, frame_duration_ms,
padding_duration_ms, vad, frames):
"""Filters out non-voiced audio frames.
Given a webrtcvad.Vad and a source of audio frames, yields only
the voiced audio.
Uses a padded, sliding window algorithm over the audio frames.
When more than 90% of the frames in the window are voiced (as
reported by the VAD), the collector triggers and begins yielding
audio frames. Then the collector waits until 90% of the frames in
the window are unvoiced to detrigger.
The window is padded at the front and back to provide a small
amount of silence or the beginnings/endings of speech around the
voiced frames.
Arguments:
sample_rate - The audio sample rate, in Hz.
frame_duration_ms - The frame duration in milliseconds.
padding_duration_ms - The amount to pad the window, in milliseconds.
vad - An instance of webrtcvad.Vad.
frames - a source of audio frames (sequence or generator).
Returns: A generator that yields PCM audio data.
"""
num_padding_frames = int(padding_duration_ms / frame_duration_ms)
# We use a deque for our sliding window/ring buffer.
ring_buffer = collections.deque(maxlen=num_padding_frames)
# We have two states: TRIGGERED and NOTTRIGGERED. We start in the
# NOTTRIGGERED state.
triggered = False
voiced_frames = []
for frame in frames:
is_speech = vad.is_speech(frame.bytes, sample_rate)
if not triggered:
ring_buffer.append((frame, is_speech))
num_voiced = len([f for f, speech in ring_buffer if speech])
# If we're NOTTRIGGERED and more than 90% of the frames in
# the ring buffer are voiced frames, then enter the
# TRIGGERED state.
if num_voiced > 0.9 * ring_buffer.maxlen:
triggered = True
start = ring_buffer[0][0].timestamp
# We want to yield all the audio we see from now until
# we are NOTTRIGGERED, but we have to start with the
# audio that's already in the ring buffer.
for f, s in ring_buffer:
voiced_frames.append(f)
ring_buffer.clear()
else:
# We're in the TRIGGERED state, so collect the audio data
# and add it to the ring buffer.
voiced_frames.append(frame)
ring_buffer.append((frame, is_speech))
num_unvoiced = len([f for f, speech in ring_buffer if not speech])
# If more than 90% of the frames in the ring buffer are
# unvoiced, then enter NOTTRIGGERED and yield whatever
# audio we've collected.
if num_unvoiced > 0.9 * ring_buffer.maxlen:
triggered = False
yield (start, frame.timestamp + frame.duration)
ring_buffer.clear()
voiced_frames = []
# If we have any leftover voiced audio when we run out of input,
# yield it.
if voiced_frames:
yield (start, frame.timestamp + frame.duration)
def VAD_chunk(aggressiveness, path):
audio, byte_audio = read_wave(path, sr)
vad = webrtcvad.Vad(int(aggressiveness))
frames = frame_generator(20, byte_audio, sr)
frames = list(frames)
times = vad_collector(sr, 20, 200, vad, frames)
speech_times = []
speech_segs = []
for i, time in enumerate(times):
start = np.round(time[0],decimals=2)
end = np.round(time[1],decimals=2)
j = start
while j + .4 < end:
end_j = np.round(j+.4,decimals=2)
speech_times.append((j, end_j))
speech_segs.append(audio[int(j*sr):int(end_j*sr)])
j = end_j
else:
speech_times.append((j, end))
speech_segs.append(audio[int(j*sr):int(end*sr)])
return speech_times, speech_segs
# + id="T<KEY>"
#collapse-hide
# Based on code from PyTorch Speaker Verification:
# https://github.com/HarryVolek/PyTorch_Speaker_Verification
# Copyright (c) 2019, HarryVolek
# Additions Copyright (c) 2021, <NAME>
# License: MIT
import numpy as np
# wav2vec2's max duration is 40 seconds, using 39 by default
# to be a little safer
def vad_concat(times, segs, max_duration=39.0):
"""
Concatenate continuous times and their segments, where the end time
of a segment is the same as the start time of the next
Parameters:
times: list of tuple (start, end)
segs: list of segments (audio frames)
max_duration: maximum duration of the resulting concatenated
segments; the kernel size of wav2vec2 is 40 seconds, so
the default max_duration is 39, to ensure the resulting
list of segments will fit
Returns:
concat_times: list of tuple (start, end)
concat_segs: list of segments (audio frames)
"""
absolute_maximum=40.0
if max_duration > absolute_maximum:
raise Exception('`max_duration` {:.2f} larger than kernel size (40 seconds)'.format(max_duration))
# we take 0.0 to mean "don't concatenate"
do_concat = (max_duration != 0.0)
concat_seg = []
concat_times = []
seg_concat = segs[0]
time_concat = times[0]
for i in range(0, len(times)-1):
can_concat = (times[i+1][1] - time_concat[0]) < max_duration
if time_concat[1] == times[i+1][0] and do_concat and can_concat:
seg_concat = np.concatenate((seg_concat, segs[i+1]))
time_concat = (time_concat[0], times[i+1][1])
else:
concat_seg.append(seg_concat)
seg_concat = segs[i+1]
concat_times.append(time_concat)
time_concat = times[i+1]
else:
concat_seg.append(seg_concat)
concat_times.append(time_concat)
return concat_times, concat_seg
# + id="CDzccsIklFkV"
def make_dataset(concat_times, concat_segs):
starts = [s[0] for s in concat_times]
ends = [s[1] for s in concat_times]
return {'start': starts,
'end': ends,
'speech': concat_segs}
# + id="ZzR4mZBZC3-5"
# %%capture
# !pip install datasets
# + id="lIbDIIkFmQJu"
from datasets import Dataset
def vad_to_dataset(path, max_duration):
t,s = VAD_chunk(3, path)
if max_duration > 0.0:
ct, cs = vad_concat(t, s, max_duration)
dset = make_dataset(ct, cs)
else:
dset = make_dataset(t, s)
return Dataset.from_dict(dset)
# + id="xLuEz02ZlSvC"
# %%capture
# !pip install -q transformers
# + id="Hac9-TrHmyaM"
# %%capture
from transformers import Wav2Vec2Processor, Wav2Vec2ForCTC
# load model and tokenizer
processor = Wav2Vec2Processor.from_pretrained("mbien/wav2vec2-large-xlsr-polish")
model = Wav2Vec2ForCTC.from_pretrained("mbien/wav2vec2-large-xlsr-polish")
model.to("cuda")
# + id="GPoyTvuHm_gC"
def speech_file_to_array_fn(batch):
import torchaudio
speech_array, sampling_rate = torchaudio.load(batch["path"])
batch["speech"] = speech_array[0].numpy()
batch["sampling_rate"] = sampling_rate
batch["target_text"] = batch["sentence"]
return batch
def evaluate(batch):
import torch
inputs = processor(batch["speech"], sampling_rate=16_000, return_tensors="pt", padding=True)
with torch.no_grad():
logits = model(inputs.input_values.to("cuda"), attention_mask=inputs.attention_mask.to("cuda")).logits
pred_ids = torch.argmax(logits, dim=-1)
batch["pred_strings"] = processor.batch_decode(pred_ids)
return batch
# + id="GgjOpU7ynmpT"
import json
def process_wave(filename, duration):
import json
dataset = vad_to_dataset(filename, duration)
result = dataset.map(evaluate, batched=True, batch_size=16)
speechless = result.remove_columns(['speech'])
d=speechless.to_dict()
tlog = list()
for i in range(0, len(d['end']) - 1):
out = dict()
out['start'] = d['start'][i]
out['end'] = d['end'][i]
out['transcript'] = d['pred_strings'][i]
tlog.append(out)
with open('{}.tlog'.format(filename), 'w') as outfile:
json.dump(tlog, outfile)
# + id="uRmfyBqpoZeD"
import glob
for f in glob.glob('/content/poleval_final_dataset_wav/train/*.wav'):
print(f)
process_wave(f, 10.0)
# + id="AeHvI5e37q1Z"
# !find . -name '*tlog'|zip poleval-train.zip -@
| _notebooks/2021-06-16-poleval-through-wav2vec2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ml] *
# language: python
# name: conda-env-ml-py
# ---
# %reload_ext nb_black
# # Day 11
from __future__ import annotations
from typing import List, Dict, NamedTuple
from collections import deque
# ## Part One
def parse(raw: str) -> List[List[str]]:
raw = raw.strip()
seats = raw.split("\n")
seats = [list(row) for row in seats]
return seats
# +
def count_adj(x: int, y: int, seats: List[List[str]]) -> int:
dx = [-1, -1, -1, 0, 0, 1, 1, 1]
dy = [-1, 0, 1, -1, 1, -1, 0, 1]
cnt = 0
for k in range(len(dx)):
nx = x + dx[k]
ny = y + dy[k]
if 0 <= nx < len(seats) and 0 <= ny < len(seats[x]):
if seats[nx][ny] == "#":
cnt += 1
return cnt
def one_round(seats: List[List[str]], limit=4, count_fn=count_adj) -> List[List[str]]:
changed = 0
nseats = [[c for c in row] for row in seats]
for i in range(len(seats)):
for j in range(len(seats[i])):
cnt = count_fn(i, j, seats)
if seats[i][j] == "L" and cnt == 0:
nseats[i][j] = "#"
changed += 1
elif seats[i][j] == "#" and cnt >= limit:
nseats[i][j] = "L"
changed += 1
return nseats, changed
# -
def simulate(seats: List[List[str]], limit=4, count_fn=count_adj) -> int:
changed = -1
while changed:
seats, changed = one_round(seats, limit=limit, count_fn=count_fn)
# import pprint
# pprint.pprint(seats)
# print("\n".join(["".join(row) for row in seats]))
answer = sum(c == "#" for row in seats for c in row)
return answer
RAW = """L.LL.LL.LL
LLLLLLL.LL
L.L.L..L..
LLLL.LL.LL
L.LL.LL.LL
L.LLLLL.LL
..L.L.....
LLLLLLLLLL
L.LLLLLL.L
L.LLLLL.LL"""
seats = parse(RAW)
assert simulate(seats) == 37
with open("../input/day11.txt") as f:
raw = f.read()
seats = parse(raw)
print(simulate(seats))
# ## Part Two
def count_adj2(x: int, y: int, seats: List[List[str]]) -> int:
cnt = 0
dxs = [-1, -1, -1, 0, 0, 1, 1, 1]
dys = [-1, 0, 1, -1, 1, -1, 0, 1]
n, m = len(seats), len(seats[0])
for dx, dy in zip(dxs, dys):
nx, ny = x, y
while True:
nx += dx
ny += dy
if nx < 0 or nx >= n or ny < 0 or ny >= m:
break
if seats[nx][ny] == "#":
cnt += 1
break
elif seats[nx][ny] == "L":
break
return cnt
RAW = """L.LL.LL.LL
LLLLLLL.LL
L.L.L..L..
LLLL.LL.LL
L.LL.LL.LL
L.LLLLL.LL
..L.L.....
LLLLLLLLLL
L.LLLLLL.L
L.LLLLL.LL"""
SEATS = parse(RAW)
assert simulate(SEATS, limit=5, count_fn=count_adj2) == 26
with open("../input/day11.txt") as f:
raw = f.read()
seats = parse(raw)
print(simulate(seats, limit=5, count_fn=count_adj2))
| notebooks/day11.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#1.1.-Python-Keywords" data-toc-modified-id="1.1.-Python-Keywords-1">1.1. Python Keywords</a></span></li><li><span><a href="#1.2.-Identifiers" data-toc-modified-id="1.2.-Identifiers-2">1.2. Identifiers</a></span></li><li><span><a href="#1.3-Python-Comments" data-toc-modified-id="1.3-Python-Comments-3">1.3 Python Comments</a></span></li><li><span><a href="#1.4.-Multi-Line-Comments" data-toc-modified-id="1.4.-Multi-Line-Comments-4">1.4. Multi Line Comments</a></span></li><li><span><a href="#1.5.-DocString-in-python" data-toc-modified-id="1.5.-DocString-in-python-5">1.5. DocString in python</a></span></li><li><span><a href="#1.6.-Python-Indentation" data-toc-modified-id="1.6.-Python-Indentation-6">1.6. Python Indentation</a></span></li><li><span><a href="#1.7.-Python-Statement" data-toc-modified-id="1.7.-Python-Statement-7">1.7. Python Statement</a></span></li><li><span><a href="#1.8.-Multi-Line-Statement" data-toc-modified-id="1.8.-Multi-Line-Statement-8">1.8. Multi-Line Statement</a></span></li><li><span><a href="#1.9.-Variables" data-toc-modified-id="1.9.-Variables-9">1.9. Variables</a></span><ul class="toc-item"><li><span><a href="#1.9.1.-Variable-Assignments" data-toc-modified-id="1.9.1.-Variable-Assignments-9.1">1.9.1. Variable Assignments</a></span></li><li><span><a href="#1.9.2.-Multiple-Assignments" data-toc-modified-id="1.9.2.-Multiple-Assignments-9.2">1.9.2. Multiple Assignments</a></span></li><li><span><a href="#1.9.3-Storage-Locations" data-toc-modified-id="1.9.3-Storage-Locations-9.3">1.9.3 Storage Locations</a></span></li></ul></li></ul></div>
# -
# ## 1.1. Python Keywords
# Keywords are the reserved words in python
#
# We can't use a keyword as variable name, function name or any other identifier
#
# Keywords are case sentive
# +
#Get all keywords in python 3.6
import keyword
print(keyword.kwlist)
print("Total number of keywords ", len(keyword.kwlist))
# -
# ## 1.2. Identifiers
# Identifier is the name given to entities like class, functions, variables etc. in Python. It helps differentiating one entity from another.
# Rules for Writing Identifiers:
#
# 1. Identifiers can be a combination of letters in lowercase (a to z) or uppercase (A to Z) or digits (0 to 9) or an underscore (_).
#
# 2. An identifier cannot start with a digit. 1variable is invalid, but variable1 is perfectly fine.
#
# 3. Keywords cannot be used as identifiers.
abc = 156
abc
global = 1
@ = 1
# !abc = 8
_abc = 'vishnu'
# We cannot use special symbols like !, @, #, $, % etc. in our identifier.
a@ = 10 #can't use special symbols as an identifier
# ## 1.3 Python Comments
# Comments are lines that exist in computer programs that are ignored by compilers and interpreters.
#
# Including comments in programs makes code more readable for humans as it provides some information or explanation about what each part of a program is doing.
#
# In general, it is a good idea to write comments while you are writing or updating a program as it is easy to forget your thought process later on, and comments written later may be less useful in the long term.
# In Python, we use the hash (#) symbol to start writing a comment.
# +
#Print Hello, world to console
print("Hello, world")
# -
# ## 1.4. Multi Line Comments
# If we have comments that extend multiple lines, one way of doing it is to use hash (#) in the beginning of each line.
# +
#This is a long comment
#and it extends
#Multiple lines
# -
# Another way of doing this is to use triple quotes, either ''' or """.
"""This is also a
perfect example of
multi-line comments"""
'''
'''
"""
"""
# ## 1.5. DocString in python
# Docstring is short for documentation string.
#
# It is a string that occurs as the first statement in a module, function, class, or method definition.
# +
def double(num):
"""
function to double the number
"""
return 2 * num
print(double(10))
# -
def adds(num1,num2):
'''
function to add two numbers
'''
return num1+num2
adds(5,48)
print(adds.__doc__)
print(double.__doc__) #Docstring is available to us as the attribute __doc__ of the function
# ## 1.6. Python Indentation
# 1. Most of the programming languages like C, C++, Java use braces { } to define a block of code. Python uses indentation.
#
# 2. A code block (body of a function, loop etc.) starts with indentation and ends with the first unindented line. The amount of indentation is up to you, but it must be consistent throughout that block.
#
# 3. Generally four whitespaces are used for indentation and is preferred over tabs.
for i in range(100):
print(i)
for i in range(10):
print(i)
# Indentation can be ignored in line continuation. But it's a good idea to always indent. It makes the code more readable.
if True:
print("Python")
c = "SEBI"
if True: print("Python"); c = "SEBI"
# ## 1.7. Python Statement
# Instructions that a Python interpreter can execute are called statements.
# Examples:
a = 1 #single statement
# ## 1.8. Multi-Line Statement
# In Python, end of a statement is marked by a newline character. But we can make a statement extend over multiple lines with the line continuation character (\).
# +
a = 1 + 2 + 3 + \
4 + 5 + 6 + \
7 + 8
print(a)
# -
#another way is
a = (1 + 2 + 3 +
4 + 5 + 6 +
7 + 8)
print (a)
a = 10; b = 20; c = 30 #put multiple statements in a single line using ;
# ## 1.9. Variables
# A variable is a location in memory used to store some data (value).
#
# They are given unique names to differentiate between different memory locations. The rules for writing a variable name is same as the rules for writing identifiers in Python.
#
# We don't need to declare a variable before using it. In Python, we simply assign a value to a variable and it will exist. We don't even have to declare the type of the variable. This is handled internally according to the type of value we assign to the variable.
# ### 1.9.1. Variable Assignments
# +
#We use the assignment operator (=) to assign values to a variable
a = 10
b = 5.5
c = "Python"
# -
# ### 1.9.2. Multiple Assignments
a, b, c = 10, 5.5, "Python"
a = b = c = "Python" #assign the same value to multiple variables at once
# ### 1.9.3 Storage Locations
# +
x = 3
print(id(x)) #print address of variable x
# +
y = 3
print(id(y)) #print address of variable y
# -
# Observation:
#
# x and y points to same memory location
y = 2
print(id(y)) #print address of variable y
| python_basics_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## CONCLUSIONS FROM THE DATA
#
# ### 1-Schools with higher budget per students have the lowest passing overall percentages.
# ### 2-Students enrrolled in charter type schools have better performance than students in distric type schools.
# ### 3-Averages for math scores are lower than average reading scores consisntly throughout the entire data.
#
# Importing python libraries
import pandas as pd
import numpy as np
# +
# Loading the data for analysis
Schools_Data_df = "resources/schools_complete.csv"
Students_Data_df= "resources/students_complete.csv"
# +
#read school and students data
schools_df= pd.read_csv(Schools_Data_df)
students_df= pd.read_csv(Students_Data_df)
schools_df.rename(columns = {'name':'school'}, inplace = True)
schools_df.head()
merged_df = students_df.merge(schools_df, how = 'left', on = 'school')
#students_df.head()
# +
#renames for merge
schools_df.rename(columns = {'name':'school'}, inplace = True)
# -
# ## District Summary Table
# +
#merging the two data sets by using school names as common key
# school_complete_data = pd.merge(schools_df,student_df, on=["school_name"], how= "left")
# school_complete_data.head()
# complete_data= schools_df.merge(students_df, on='school', how='left')
# merged_df = pd.merge(schools_df,students_df, on = "name", how="left")
# merged_df.head()
school_data_complete = pd.merge(students_df, schools_df, how="left", on="school")
# -
total_schools = len(schools_df)
total_schools
#total numeber of students
total_students = len(school_data_complete)
total_students
#total number of schools
total_school_budget= schools_df['budget'].sum(numeric_only= False)
total_school_budget
#math score average
math_avg = students_df['math_score'].mean()
np.round(math_avg, decimals=2)
#reading score average
reading_avg = students_df['reading_score'].mean()
np.round(reading_avg, decimals=2)
# +
# #math score passing percent
passing_math = students_df.loc[students_df['math_score'] >= 70]['math_score'].count()
passing_math
# -
#calculatig math passing percentage
percent_passing_math = (passing_math/total_students)*100
np.round(percent_passing_math, decimals=2)
# calculating number of students passing reading
passing_reading = students_df.loc[students_df['reading_score'] >= 70]['reading_score'].count()
passing_reading
#calculating reading passing percentage
percent_passing_reading= np.round((passing_reading/total_students)*100,2)
percent_passing_reading
#calculating_overall_passing_percentage
overall_passing_rate= np.round((percent_passing_reading + percent_passing_math)/2,2)
overall_passing_rate
# +
#constructing the new data_frame using dictionaries
District_df = pd.DataFrame({
"Total Schools" : [total_schools],
'Total Students':[total_students],
"Total Budget" :[total_school_budget],
"Average Math Score" :[math_avg],
"Average Reading Score" :[reading_avg],
"% Passing Math" :[percent_passing_math],
"% Passing Reading" :[percent_passing_reading],
"Overall Passing Rate" :[overall_passing_rate] }, columns = ["Total Students", "Total Schools", "Total Budget", "Average Math Score", "Average Reading Score","% Passing Math", "% Passing Reading", "Overall Passing Rate"])
District_df.style.format({'Average Math Score': '{:.1f}',
'Average Reading Score': '{:.1f}',
'% Passing Math': '{:.0f}%',
'% Passing Reading':'{:.0f}%',
'Overall Passing Rate': '{:.2f}%',
'Total Budget': "${:,}",
'Total Students': "{:,}",})
# +
new_schools_df = schools_df.set_index(["school"])
#Grouping by school to analyzed the data by school
groupby_school = students_df.groupby("school")
# #school type
sch_type = new_schools_df["type"]
# #total students
sch_students= groupby_school['name'].count()
# # Total School Budget
sch_budget= new_schools_df["budget"]
# # #Per Student Budget
per_student_bdgt = schools_df.set_index('school')['budget']/schools_df.set_index('school')['size']
# -
# ## SCHOOL SUMMARY TABLE
# +
# Create the School Summary dataframe by intializing the starting value
School_Summary_DF=pd.DataFrame({"School Name": schools_df["school"],
"School Type":schools_df["type"],
"Budget":schools_df["budget"],
"Total Students":schools_df["size"]})
# Find the count of student who passed math and reading separately in each school
Count_passing_math = school_data_complete.loc[(students_df['math_score'] >= 70)].groupby(["school"]).count()["math_score"]
Count_passing_reading = school_data_complete.loc[(students_df['reading_score'] >= 70)].groupby(["school"]).count()["reading_score"]
Count_passing_overall = school_data_complete.loc[(school_data_complete['math_score'] >=70)
& (school_data_complete['reading_score'] >=70)].groupby(["school"]).count()["Student ID"]
# Create a temporary dataframe to store the calculated values
temp_df=pd.DataFrame({"Avg Math Score": school_data_complete.mean()["math_score"],
"Avg Reading Score": school_data_complete.mean()["reading_score"],
"Count of Passing Math": Count_passing_math,
"Count of Passing Reading":Count_passing_reading,
"Count of Passing Overall":Count_passing_overall})
# Reset the index so school becomes a column
temp_df.reset_index(inplace=True)
# Rename the column school before merging
temp_df = temp_df.rename(columns={"school":"School Name"})
# Merge both the Summary dataframe and the temporary dataframe on School Name
School_Summary_DF=pd.merge(School_Summary_DF,temp_df,how="outer",on="School Name")
# Calculate the remaining values
School_Summary_DF["% Passing Math"]=(School_Summary_DF["Count of Passing Math"]/School_Summary_DF["Total Students"])*100
School_Summary_DF["% Passing Reading"]=(School_Summary_DF["Count of Passing Reading"]/School_Summary_DF["Total Students"])*100
School_Summary_DF["% Passing Overall"]=(School_Summary_DF["Count of Passing Overall"]/School_Summary_DF["Total Students"])*100
# Drop the columns after computation
School_Summary_DF.drop(["Count of Passing Math","Count of Passing Reading","Count of Passing Overall"],axis=1,inplace=True)
# Set the index as School Name
School_Summary_DF.set_index("School Name",inplace=True)
School_Summary_DF.head()
# -
# ## TOP 5 PERFORMING SCHOOLS
# +
#top 5 Peforming Schools
top_5 = School_Summary_DF.sort_values(by="% Passing Overall",ascending=False)
top_5.head()
# -
# ## BOTTOM 5 PERFORMING SCHOOLS
# +
#bottom 5
bottom_5 = School_Summary_DF.sort_values(by = '% Passing Overall', ascending= True)
bottom_5.head()
# -
# ## MATH SCORES BY GRADE
# +
#Math Scores by Grade
#creates grade level average math scores for each school
#creates grade level average math scores for each school
ninth_grade_math = students_df.loc[students_df['grade'] == '9th'].groupby('school')['math_score'].mean()
tenth_grade_math = students_df.loc[students_df['grade']== '10th'].groupby('school')['math_score'].mean()
eleventh_grade_math= students_df.loc[students_df['grade']== '11th'].groupby('school')['math_score'].mean()
twelveth_grade_math= students_df.loc[students_df['grade']=='12th'].groupby('school')['math_score'].mean()
math_scores_by_grade = pd.DataFrame({ '9th': ninth_grade_math,
'10th': tenth_grade_math,
'11th':eleventh_grade_math,
'12th':twelveth_grade_math})
math_scores_by_grade= math_scores_by_grade[['9th', '10th', '11th', '12th']]
math_scores_by_grade.index.name = "School Name"
#format
math_scores_by_grade.style.format({'9th': '{:.1f}',
"10th": '{:.1f}',
"11th": "{:.1f}",
"12th": "{:.1f}"})
# -
# ## READING SCORES BY GRADE
# +
#Reading Scores by Grade
#creates grade level average reading scores for each school
ninth_grade_reading = students_df.loc[students_df['grade'] == '9th'].groupby('school')["reading_score"].mean()
tenth_grade_reading = students_df.loc[students_df['grade']== '10th'].groupby('school')["reading_score"].mean()
eleventh_grade_reading= students_df.loc[students_df['grade']== '11th'].groupby('school')["reading_score"].mean()
twelveth_grade_reading= students_df.loc[students_df['grade']=='12th'].groupby('school')["reading_score"].mean()
reading_scores_by_grade = pd.DataFrame({ '9th': ninth_grade_reading,
'10th': tenth_grade_reading,
'11th':eleventh_grade_reading,
'12th':twelveth_grade_reading})
reading_scores_by_grade= reading_scores_by_grade[['9th', '10th', '11th', '12th']]
reading_scores_by_grade.index.name = "School Name"
#style
reading_scores_by_grade.style.format({'9th': '{:.1f}',
"10th": '{:.1f}',
"11th": "{:.1f}",
"12th": "{:.1f}"})
# -
# ## SCHOOLS BY SPENDING
# +
# School_df= District_df
# School_df["Spending Ranges (per student)"] = pd.cut(per_student_bdgt,
# bins=spending_bins, labels=group_names)
categories = [0, 585, 615, 645, 675]
group_name = ['< $585', "$585 - 615", "$615 - 645", " $645-675"]
merged_df['spending_bins'] = pd.cut(merged_df['budget']/merged_df['size'], categories, labels = group_name)
#group by spending
by_spending = merged_df.groupby('spending_bins')
#calculations
avg_math = by_spending['math_score'].mean()
avg_read = by_spending['reading_score'].mean()
pass_math = merged_df[merged_df['math_score'] >= 70].groupby('spending_bins')['Student ID'].count()/by_spending['Student ID'].count()
pass_read = merged_df[merged_df['reading_score'] >= 70].groupby('spending_bins')['Student ID'].count()/by_spending['Student ID'].count()
overall= (pass_math + pass_read)/2
# Score by spending data frame
scores_by_spend = pd.DataFrame({
"Average Math Score": round(avg_math, 1),
"Average Reading Score": round (avg_read, 1),
'% Passing Math': round(pass_math,2),
'% Passing Reading': round(pass_read,2),
"Overall Passing Rate": round(overall, 2)
})
#Organizing columns
scores_by_spend = scores_by_spend[[
"Average Math Score",
"Average Reading Score",
'% Passing Math',
'% Passing Reading',
"Overall Passing Rate"
]]
scores_by_spend.index.name = "Spending Ranges (Per Student)"
scores_by_spend = scores_by_spend.reindex(group_name)
#style
scores_by_spend.style.format({'Average Math Score': '{:.1f}',
'Average Reading Score': '{:.1f}',
'% Passing Math': '{:.0%}',
'% Passing Reading':'{:.0%}',
'Overall Passing Rate': '{:.0%}'})
# -
# ## SCORES BY SCHOOLS SIZE
# +
# create size bins
bins = [0, 1000, 2000, 5000]
group_name = ["Small (<1000)", "Medium (1000-2000)" , "Large (>2000)"]
merged_df['size_bins'] = pd.cut(merged_df['size'], bins, labels = group_name)
#group by spending
by_size = merged_df.groupby('size_bins')
#calculations
avg_math = by_size['math_score'].mean()
avg_read = by_size['math_score'].mean()
pass_math = merged_df[merged_df['math_score'] >= 70].groupby('size_bins')['Student ID'].count()/by_size['Student ID'].count()
pass_read = merged_df[merged_df['reading_score'] >= 70].groupby('size_bins')['Student ID'].count()/by_size['Student ID'].count()
overall = (pass_math + pass_read)/2
# df build
scores_by_size = pd.DataFrame({
"Average Math Score": avg_math,
"Average Reading Score": avg_read,
'% Passing Math': pass_math,
'% Passing Reading': pass_read,
"Overall Passing Rate": overall
})
#reorder columns
scores_by_size = scores_by_size[[
"Average Math Score",
"Average Reading Score",
'% Passing Math',
'% Passing Reading',
"Overall Passing Rate"
]]
scores_by_size.index.name = "Total Students"
scores_by_size = scores_by_size.reindex(group_name)
#style
scores_by_size.style.format({'Average Math Score': '{:.1f}',
'Average Reading Score': '{:.1f}',
'% Passing Math': '{:.1%}',
'% Passing Reading':'{:.1%}',
'Overall Passing Rate': '{:.1%}'})
# -
# ## SCORES BY SCHOOL TYPE
# +
# group by type of school
by_type = merged_df.groupby("type")
#calculations
avg_math = by_type['math_score'].mean()
avg_read = by_type['math_score'].mean()
pass_math = merged_df[merged_df['math_score'] >= 70].groupby('type')['Student ID'].count()/by_type['Student ID'].count()
pass_read = merged_df[merged_df['reading_score'] >= 70].groupby('type')['Student ID'].count()/by_type['Student ID'].count()
overall= (pass_math + pass_read)/2
# DataFrame
scores_by_type = pd.DataFrame({
"Average Math Score": avg_math,
"Average Reading Score": avg_read,
'% Passing Math': pass_math,
'% Passing Reading': pass_read,
"Overall Passing Rate": overall})
#reorder columns
scores_by_type = scores_by_type[[
"Average Math Score",
"Average Reading Score",
'% Passing Math',
'% Passing Reading',
"Overall Passing Rate"
]]
scores_by_type.index.name = "Type of School"
#style
scores_by_type.style.format({'Average Math Score': '{:.1f}',
'Average Reading Score': '{:.1f}',
'% Passing Math': '{:.1%}',
'% Passing Reading':'{:.1%}',
'Overall Passing Rate': '{:.1%}'})
# -
| PyCitySchools/.ipynb_checkpoints/School Budget Statistical Analysis -checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="_XAKXZG-yBoh"
# #Imports/Permissions to load data
# + colab={"base_uri": "https://localhost:8080/"} id="PpAyxJM7yDw4" outputId="356a40f8-7f46-4596-ca8b-304396611283"
# Python ≥3.5 is required
import sys
assert sys.version_info >= (3, 5)
# Scikit-Learn ≥0.20 is required
import sklearn
assert sklearn.__version__ >= "0.20"
# Common imports
import numpy as np
import os
# to make this notebook's output stable across runs
np.random.seed(42)
# To plot pretty figures
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
#stuff we need
from sklearn.decomposition import PCA, IncrementalPCA
from sklearn.preprocessing import StandardScaler
from sklearn.utils import shuffle
from tensorflow.keras.utils import get_custom_objects
from google.colab import drive
import h5py
import keras
import tensorflow as tf
from keras.models import Sequential
from keras.layers import Dense, Reshape, Dropout, BatchNormalization
from keras.optimizers import SGD
from sklearn.model_selection import train_test_split
import pickle
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "classification"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
drive.mount('/content/drive')
# + [markdown] id="uL4E_3b_ov0h"
# #Data Scaling
# + [markdown] id="fGD_n5k9_YSb"
# We load the training data agin just to redefine the scaling to use on test data
# + colab={"base_uri": "https://localhost:8080/"} id="DrNrGe3jpbUX" outputId="12847fe6-6b76-47d8-f3c4-129550d74c10"
x_val, y_val=np.empty([0, 110006]), np.empty([0, 6])
filepath='./drive/Shareddrives/ML_project/Complete_Spectral_Data/Training_Data/finished_data0'
allfiles, testfiles=np.arange(100), np.arange(0, 100, 10)
trainmask=[False if num in testfiles else True for num in allfiles]
trainfiles=shuffle(allfiles[trainmask], random_state=2)
pickle.dump(trainfiles, open('fileorder', 'wb'))
sc_x, sc_y, sc_y_log=StandardScaler(), StandardScaler(), StandardScaler()
#This loading function is annoying and perhaps overly complicated, but basically, when preparing the data we accidentally left it in order of
#stellar circularity fraction. So now we load 10 random files at a time, combine their entries, and randomly select part of our validation data
#from this.
for i in range(0, 90, 10):
x_chunk, y_chunk=np.empty([0, 110006]), np.empty([0, 6])
for file in trainfiles[i:i+10]:
file_obj=h5py.File(filepath+str(file)+'.hdf5', 'r')
x, y=file_obj['Data']['x_values'], file_obj['Data']['y_values']
x_chunk, y_chunk=np.concatenate((x_chunk, x)), np.concatenate((y_chunk, y))
x_train, xv, y_train, yv=train_test_split(x_chunk, y_chunk, test_size=0.1, random_state=0) #train_test_split also shuffles data
y_train_log, y_val_log=np.log10(y_train), np.log10(y_val)
y_train_log=np.where(y_train_log==np.log10(0), -1, y_train_log)
x_val, y_val=np.concatenate((x_val, xv)), np.concatenate((y_val, yv))
sc_x.partial_fit(x_train)
sc_y.partial_fit(y_train)
sc_y_log.partial_fit(y_train_log)
# + [markdown] id="l2hBtcLGycSx"
# #Load Models
# + [markdown] id="i9T70i4mz7eJ"
# We train our auto-encoder and final model using mini-batches. After each minibatch, we save the autoencoder. This allows us to resume training manually after disconnects. When we finally train our model we simply load the autoencoder.
# + id="fMfM33n9yiYj"
#LeakyReLU definition and model initialization
def leaky_relu(z,name=None):
return tf.maximum(0.01*z, z, name=name)
get_custom_objects().update({'leaky_relu':leaky_relu})
log_ae_nn=keras.models.load_model('./drive/Shareddrives/ML_project/log_10f_70e/log_ae_nn.h5')
ae_nn=keras.models.load_model('./drive/Shareddrives/ML_project/ae_nn.h5')
# + [markdown] id="ltUMryfO3PXJ"
# #Load and Prepare Test Data
# + id="-Y6iUhjL3Uo1"
filepath='./drive/Shareddrives/ML_project/Complete_Spectral_Data/Test_Data/finished_data0'
x_values, y_values=np.empty([0, 110006]), np.empty([0, 6])
for i in range(0, 100, 10):
file_obj=h5py.File(filepath+str(i)+'.hdf5', 'r')
x, y=file_obj['Data']['x_values'], file_obj['Data']['y_values']
x_values, y_values=np.concatenate((x_values, x)), np.concatenate((y_values, y))
x_test, y_test=shuffle(x_values, y_values)
# + colab={"base_uri": "https://localhost:8080/"} id="y9cfG1QIE1js" outputId="f0e0a290-06b9-4142-f6c7-90500e021166"
y_test_log=np.log10(y_test)
y_test_log=np.where(y_test_log==np.log10(0), -1, y_test_log)
x_test_scaled, y_test_scaled, y_test_scaled_log=sc_x.transform(x_test), sc_y.transform(y_test), sc_y_log.transform(y_test_log)
# + [markdown] id="7J0htNFM6yey"
# #Testing the Models
# + id="wGxoU5-V61v1" colab={"base_uri": "https://localhost:8080/"} outputId="70e3df30-8961-4056-e110-ea0f8e3aabee"
results = ae_nn.evaluate(x_test_scaled, y_test_scaled, batch_size=32)
results_log=log_ae_nn.evaluate(x_test_scaled, y_test_scaled_log)
print("test loss for non log model:", results)
print('\n test loss for log model:', results_log)
ae_nn_test_sc = sc_y.inverse_transform(ae_nn.predict(x_test_scaled))
ae_nn_test_sc_log=sc_y_log.inverse_transform(log_ae_nn.predict(x_test_scaled))
# + colab={"base_uri": "https://localhost:8080/"} id="nn8TNrG9ONVX" outputId="897b64eb-12a8-4b79-b469-d0054c8c7738"
max(y_val[:, -2]), max(ae_nn_test_sc[:, -2])
# + id="dNUIsylN72PP" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="66126515-d639-490c-eee6-bd54cece5a6e"
from matplotlib.pyplot import figure
CDs=['H', 'C', 'N', 'O', 'Fe', 'Neutral Hydrogen']
for i in range(6):
f, (ax0, ax1)=plt.subplots(1, 2, figsize=(15, 15))
#non-log plots
axis_limits=[min(min(y_test[:, i]), min(ae_nn_test_sc[:, i])), max(max(y_test[:, i]), max(ae_nn_test_sc[:, i]))]
ax0.set_xlim(*axis_limits)
ax0.set_ylim(*axis_limits)
ax0.set_title('nonlog plot of '+CDs[i])
ax0.set_aspect(aspect=1)
ax0.scatter(y_test[:,i], ae_nn_test_sc[:,i], s=0.7)
ax0.plot(np.linspace(*axis_limits), np.linspace(*axis_limits))
#log plots
axis_limits=[min(min(y_test_log[:, i]), min(ae_nn_test_sc_log[:, i])), max(max(y_test_log[:, i]), max(ae_nn_test_sc_log[:, i]))]
ax1.set_xlim(*axis_limits)
ax1.set_ylim(*axis_limits)
ax1.set_title('log plot of '+CDs[i])
ax1.set_aspect(aspect=1)
ax1.scatter(y_test_log[:,i], ae_nn_test_sc_log[:,i], s=0.7)
ax1.plot(np.linspace(*axis_limits), np.linspace(*axis_limits))
plt.show()
plt.clf()
# + colab={"base_uri": "https://localhost:8080/"} id="hrI0obHFKzPA" outputId="40bb629a-aecb-4cd3-c1be-910cca74fa56"
y_test_scaled_log.shape
| students_final_projects/group-c/Project_Final_Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Scraping data relating to different breeds of dogs from American Kennel Club website
from urllib.request import Request, urlopen
import requests
from bs4 import BeautifulSoup
import csv
# ### Getting url of individual pages for each breed
mainsite = "http://www.akc.org/content/news/articles/most-popular-dog-breeds-full-ranking-list/?button"
hdr = {'User-Agent': 'Mozilla/5.0'}
page = requests.get(mainsite)
mainsoup = BeautifulSoup(page.text, "html.parser")
#print(mainsoup.prettify())
allurl = mainsoup.find_all('td')
#allurl
type(allurl)
allurl[0].a['href']
allurl[5].a['href']
len(allurl)
allurl[940].a['href']
940/5
for i in range(0, 5):
#for i in range(0, 189):
print(allurl[5*i].text.strip())
for i in range(0, 5):
#for i in range(0, 189):
print(allurl[5*i].a['href'])
name_db = []
url_db = []
for i in range(0, 189):
name = allurl[5*i].text.strip()
url = allurl[5*i].a['href']
name_db.append(name)
url_db.append(url)
name_db
# ### Scraping data of a particular breed
site = "http://www.akc.org/dog-breeds/havanese/"
hdr = {'User-Agent': 'Mozilla/5.0'}
bookpage = requests.get(site)
soup = BeautifulSoup(bookpage.text, "html.parser")
#print(soup.prettify())
soup.find_all('span', class_ = "item")
soup.find_all('span', class_ = "item")[1].get_text()
soup.find('span', class_ = "title").get_text()
soup.find('span', class_ = "energy_levels").get_text().strip()
soup.find('span', class_ = "size").get_text().strip()
soup.find('div', class_ = "bigrank").get_text().strip()
j = soup.find('div', class_ = "breed-details__main")
j
j("li")[2].get_text().strip()
j("li")[3].get_text().strip()
j("li")[10].get_text().strip()
site = "http://www.akc.org/dog-breeds/labrador-retriever/"
hdr = {'User-Agent': 'Mozilla/5.0'}
bookpage = requests.get(site)
soup = BeautifulSoup(bookpage.text, "html.parser")
j = soup.find('div', class_ = "breed-details__main")
j
len(j)
# ### Putting all together
import time
data = []
##testing ten pages
for count in range(0,11):
##comment the above and uncomment the below to run all pages
#for count in range(0,189):
hdr = {'User-Agent': 'Mozilla/5.0'}
page = requests.get(url_db[count])
soup = BeautifulSoup(page.text, "html.parser")
#print(soup.prettify())
name = name_db[count]
try:
grp = soup.find('span', class_ = "title").get_text()
except:
grp = "NA"
try:
trait1 = soup.find_all('span', class_ = "item")[0].get_text()
except:
trait1 = "NA"
try:
trait2 = soup.find_all('span', class_ = "item")[1].get_text()
except:
trait2 = "NA"
try:
trait3 = soup.find_all('span', class_ = "item")[2].get_text()
except:
trait3 = "NA"
try:
energy = soup.find('span', class_ = "energy_levels").get_text().strip()
except:
energy = "NA"
try:
size = soup.find('span', class_ = "size").get_text().strip()
except:
size = "NA"
try:
rank = soup.find('div', class_ = "bigrank").get_text().strip()
except:
rank = "NA"
j = soup.find('div', class_ = "breed-details__main")
try:
children = j("li")[2].get_text().strip()
except:
children = "NA"
try:
otherdogs = j("li")[3].get_text().strip()
except:
otherdogs = "NA"
try:
shedding = j("li")[4].get_text().strip()
except:
shedding = "NA"
try:
grooming = j("li")[5].get_text().strip()
except:
grooming = "NA"
try:
trainability = j("li")[6].get_text().strip()
except:
trainability = "NA"
try:
height = j("li")[7].get_text().strip()
except:
height = "NA"
try:
weight = j("li")[8].get_text().strip()
except:
weight = "NA"
try:
life = j("li")[9].get_text().strip()
except:
life = "NA"
try:
barking = j("li")[10].get_text().strip()
except:
barking = "NA"
if(barking == "NA"):
j = str(j)
else:
j = ""
data.append((name,grp,trait1,trait2,trait3,energy,size,rank,children,otherdogs,shedding,grooming,trainability,
height,weight,life,barking,j))
print(count)
time.sleep(1)
# +
#data
# -
import pandas as pd
import numpy as np
df = pd.DataFrame(np.array(data))
df.columns = ['name','grp','trait1','trait2','trait3','energy','size','rank','children','otherdogs','shedding','grooming','trainability',
'height','weight','life','barking','remarks']
df.head()
# ### Saving output as csv
df.to_csv('akcdogs.csv')
| notebooks/AKCDogs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:seychelles]
# language: python
# name: conda-env-seychelles-py
# ---
import tensorflow as tf
print(tf.__version__)
# +
# %matplotlib inline
import numpy as np
from matplotlib import pyplot as plt
#from tensorflow.nn.rnn import *
from tensorflow.python.ops import *
# %load_ext autoreload
# %autoreload 2
# +
import numpy as np
import pandas as pd
import tensorflow as tf
from tensorflow.python.framework import dtypes
import seaborn as sns
# -
# cd /
# cd Users/Seychelles/Desktop/GoogleTensorflow/Data_Collector/CSV_file_from_2007_to_2017/
DataKarpos = pd.read_csv('Target.csv') # Only Karpos Location is used here . Other locations are zero
DataKarpos[1:2]
TargetKarpos = DataKarpos[['PM10']].copy(deep=True) # copy PM10 data out from data sets
# # Analysis Of Misiing Values
Target = DataKarpos[['PM10']].copy(deep=True)
Target.shape
null_data_PM10 = Target[Target.isnull().any(axis=1)] # extract data sets with null values
null_data_PM10.to_csv("PM10_NUll.txt")# save to file
Target.to_csv("R_target.csv")# save to file
null_data_PM10.head()
def getPctMissing(series):
'''
Returns percentage of Missing data in a data set.
Input : Pandas series
Output: Percentage of missing data in data set
'''
num = series.isnull().sum()
Total_data = len(series)
return 100*(num/Total_data)
print("Number of null values in PM10 : ");Target.isnull().sum()
print("Number of non null values in PM10 : ");Target.count()
print("Percentage of Missing Value: "); getPctMissing(Target)
# # Fill Missing Values With Nan
pwd
Target.fillna(method='ffill',inplace=True)
Target.fillna(Target.median,inplace=True) # Fill Missing value with 0. Default model
Target.describe()
Target_list = Target.values.tolist() # create a list of values for use in data model
# # Prepare Data
counter=0
def dataGetter(datalist):
"""
Returns a single PM10 value.
Input: PM10 Data list named Target_list above
Output: single PM10 value
"""
global counter;
assert type(datalist) == list;
value = datalist[counter]
#print(counter)
counter = counter +1
return np.array([value])
# Define window size or learning
sliding_window = []
lag=25 # 24 hours time step from 0 to 24 plus the window_step_move (25) which is to be predicted
window_move_step = 1
for i in range(lag - window_move_step):
sliding_window.append(dataGetter(Target_list))
len(sliding_window)
def get_pair(Target):
"""
Returns an (current, target)
Target is lag steps 24 ahead of current
"""
global sliding_window
sliding_window.append(dataGetter(Target))
#print(len(sliding_window))
input_value = sliding_window[0:24]
input_value=np.array(input_value)
#print(input_value.shape)
input_value=np.reshape(input_value,(1,24))
##print(input_value)
output_value = sliding_window[-1]
##print(output_value)
sliding_window = sliding_window[1:]
#print(len(sliding_window))
return input_value, output_value
len(sliding_window)
# #Input Params
# input_dim = 1
#
# #To maintain state
# last_value = np.array([0 for i in range(input_dim)])
# last_derivative = np.array([0 for i in range(input_dim)])
#
# def get_total_input_output(Target):
# """
# Get Total feed vector - input and output
# """
# global last_value, last_derivative
# raw_i, raw_o = get_pair(Target)
# raw_i = raw_i[0]
# l1 = list(raw_i)
# derivative = raw_i - last_value
# l2 = list(derivative)
# last_value = raw_i
# l3 = list(derivative - last_derivative)
# last_derivative = derivative
# return np.array([l1 + l2 + l3]), raw_o
# # Build Model
#Imports
import tensorflow as tf
#from tensorflow.nn.rnn import *
from tensorflow.python.ops import *
#Input Params
with tf.name_scope("input_target_placeholders"):
input_dim = 1
##The Input Layer as a Placeholder
#Since we will provide data sequentially, the 'batch size'
#is 1.
input_layer = tf.placeholder(tf.float32, [1, input_dim*24],name="input_data")
correct_output = tf.placeholder(tf.float32, [1, input_dim],name="target_data")
# +
###inistate = tf.Variable(lstm_cell_with_dropout.zero_state(BATCH_SIZE, tf.float32), trainable=False)
# -
with tf.name_scope("lstmLayer"):
lstm_layer1 = rnn_cell.BasicLSTMCell(input_dim*24,state_is_tuple=False)
#The LSTM state as a Variable initialized to zeroes
lstm_state1 = tf.Variable(tf.zeros([1, lstm_layer1.state_size]),trainable=False,name="initial_state")
#lstm_state1 = tf.Variable(lstm_layer1.zero_state(1,lstm_layer1.state_size[-1] ), trainable=False)
#Connect the input layer and initial LSTM state to the LSTM cell
lstm_output1, lstm_state_output1 = lstm_layer1(input_layer, lstm_state1)
#The LSTM state will get updated
lstm_update_op1 = lstm_state1.assign(lstm_state_output1)
lstm_state_output1.get_shape()# verify shape
with tf.name_scope("weight_Bias_learning_rate"):
global_step = tf.Variable(0, trainable=False,name="global_step")
starter_learning_rate = 0.09
learning_rate = tf.train.exponential_decay(starter_learning_rate, global_step,
1000, 0.8, staircase=False,name ="Exponential_decay")
##The Regression-Output Layer
#The Weights and Biases matrices first
output_W1 = tf.Variable(tf.truncated_normal([input_dim*48, input_dim]),name="weight")
output_b1 = tf.Variable(tf.zeros([input_dim]),name="bias")
# lambda_l2_reg=0.5
# l2 = lambda_l2_reg * sum(
# tf.nn.l2_loss(tf_var)
# for tf_var in tf.trainable_variables()
# if not ("noreg" in tf_var.name or "bias" in tf_var.name)
# )
# #loss += l2
with tf.name_scope("prediction"):
#Compute the output
final_output = tf.matmul(lstm_state_output1, output_W1) + output_b1
final_output.get_shape() # verify output shape
output_W1 # verify weight shape
with tf.name_scope("RMS_error"):
##Calculate the Sum-of-Squares Error
error = tf.pow(tf.sub(final_output, correct_output), 2)+l2
# +
#error = tf.reshape(error,(1,))
# -
with tf.name_scope("optimizer"):
##The Optimizer
#Adam works best
train_step = tf.train.AdamOptimizer(learning_rate).minimize(error)
# +
# Create a summary to monitor MSE
mse=tf.summary.tensor_summary("errors_Summary",error)
# Create a summary to monitor predictions
prediction=tf.summary.tensor_summary("predictions_Summmary", final_output)
# Create a summary to monitor bias
bias_vec=tf.summary.tensor_summary("bias", output_b1)
# create sumary
#rate_vec=tf.summary.scalar("rate", learning_rate)
#histogram plot
error_stats=tf.histogram_summary("errors_Histogram",error)
weight_stats=tf.histogram_summary("weights_Histogram",output_W1)
bias_stats=tf.histogram_summary("biases_Histogram",output_b1)
#learning_stats=tf.histogram_summary("biases_Histogram",learning_rate)
#merged_summary_op = tf.merge_all_summaries()
merged_summary_op = tf.merge_summary([mse,prediction,bias_vec,error_stats,weight_stats,bias_stats])
# -
##Session
sess = tf.Session()
#Initialize all Variables
sess.run(tf.initialize_all_variables())
len(Target_list)
# # Evaluation Data set
logs_path = '/Users/Seychelles/Desktop/GoogleTensorflow/finals/tensorboardData/sliding_window/'
##Training Parameters
n_iter = 80000
inner_iter = 8000
actual_output1 = []
network_output1 = []
validation_prediction=[];
validation_target =[];
lower_bound=0;
upper_bound=800;
prediction_window=800
import copy
#new_list = copy.deepcopy(old_list)
Test_eval=copy.deepcopy(Target_list[80001:])
len(Test_eval)
len(Target_list)-80000
# We have 800000 iterations
# Between itertions and every 10000 steps we make all initial_state = 0 and perform cross-validate
# We have 80000 validation set
# We test with the remainder
Remainder = len(Test_eval) - (n_iter/inner_iter*prediction_window)
test_part =48;
Remainder= Remainder-test_part
Remainder = int(Remainder)
Remainder
#
# error_=[]
# with tf.Session() as sess:
#
#
#
# writer = tf.train.SummaryWriter(logs_path, graph= tf.get_default_graph())
#
# counter =0
# assert counter==0
#
# for i in range(n_iter):
# input_v, output_v = get_pair(Target_list)
# _, _, network_output,errors,summary = sess.run([lstm_update_op1,
# train_step,
# final_output,error,merged_summary_op],
# feed_dict = {
# input_layer: input_v,
# correct_output: output_v})
# writer.add_summary(summary)
# error_.append(errors)
# if i%inner_iter==0 & i!=0:
# assert i!=0;
# sess.run(lstm_state1.assign(tf.zeros([1, lstm_layer1.state_size])))
#
# for j in range(len(Test_eval[lower_bound:upper_bound])):
# input_val, output_val = get_pair(Test2)
# _, network_output = sess.run([lstm_update_op1,
# final_output],
# feed_dict = {
# input_layer: input_val,
# correct_output: output_val})
# lower_bound= lower_bound+prediction_window;
# upper_bound = upper_bound + prediction_window;
# validation_target.append(output_val)
# validation_prediction.append(network_output1)
# #sess.run(lstm_state1.assign(tf.zeros([1, lstm_layer1.state_size])))
#
# actual_output1.append(output_v)
# #actual_output2.append(output_v[0][1])
# network_output1.append(network_output)
# #network_output2.append(network_output[0][1])
# #x_axis.append(i)
#
#
#
#
# +
error_=[]
writer = tf.train.SummaryWriter(logs_path, graph= tf.get_default_graph())
counter =0
assert counter==0
for i in range(n_iter):
input_v, output_v = get_pair(Target_list)
_, _, network_output,errors,summary = sess.run([lstm_update_op1,
train_step,
final_output,error,merged_summary_op],
feed_dict = {
input_layer: input_v,
correct_output: output_v})
writer.add_summary(summary)
error_.append(errors)
if i%inner_iter==0 & i!=0:
assert i!=0;
sess.run(lstm_state1.assign(tf.zeros([1, lstm_layer1.state_size])))
for j in range(len(Test_eval[lower_bound:upper_bound])):
input_val, output_val = get_pair(Test2)
_, network_output = sess.run([lstm_update_op1,
final_output],
feed_dict = {
input_layer: input_val,
correct_output: output_val})
lower_bound= lower_bound+prediction_window;
upper_bound = upper_bound + prediction_window;
validation_target.append(output_val)
validation_prediction.append(network_output1)
#sess.run(lstm_state1.assign(tf.zeros([1, lstm_layer1.state_size])))
actual_output1.append(output_v)
#actual_output2.append(output_v[0][1])
network_output1.append(network_output)
#network_output2.append(network_output[0][1])
#x_axis.append(i)
# -
# # Errors Plot
errorplot = np.array(error_) # make errors into arrays
errorplot = errorplot.reshape(80000,1)
# +
import matplotlib.mlab as mlab
fig, ax = plt.subplots()
n, bins, patches=plt.hist(errorplot,100,normed=1,facecolor='deepskyblue',alpha=0.9,label='Histogram')
mu= np.mean(errorplot);sigma=np.std(errorplot);
y = mlab.normpdf( bins,mu,sigma)
#y = mlab.normpdf( bins, mu, sigma)
plt.plot(bins, y, 'r--', linewidth=5,label="pdf plot Over Histogram")
#ax.set_yscale('log')
#ax.set_xscale('log')
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.title("Histogram plot for error",fontweight='bold')
plt.ylabel("Error",fontweight='bold')
plt.xlabel("Training example",fontweight='bold')
plt.show()
# +
import matplotlib.mlab as mlab
fig, ax = plt.subplots()
plt.hist(errorplot,bins=200,normed=1,facecolor='deepskyblue',label='Error Histogram plot (log scale)')
ax.set_yscale('log')
#ax.set_xscale('log')
plt.title("Error plot",fontweight='bold')
plt.ylabel("Error",fontweight='bold')
plt.xlabel("Training Samples",fontweight='bold')
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.show()
# -
# # Extract Date in the series
from matplotlib.dates import date2num
Data = pd.read_csv('ItemListWithDummy.csv')
pd.to_datetime(Data.date);
x_axis = Data['date']
x_axis = pd.to_datetime(x_axis)
# +
#80000-len(network_output1)
# -
network = np.array(network_output1)
actual= np.array(actual_output1)
actual.shape
# +
import matplotlib
plt.rcParams["figure.figsize"] = (8,5)
fig = plt.figure(figsize=(100, 100))
fig, ax = plt.subplots()
ax.plot(x_axis[0:48], network.reshape(80000,1)[0:48], 'r-',color="deepskyblue")
ax.plot_date(x_axis[0:48], actual.reshape(80000,1)[0:48], 'b-',color='goldenrod')
#ax.xaxis.set_minor_locator(dates.MonthLocator())
#ax.xaxis.set_minor_formatter(dates.DateFormatter('%H:%M:%S'))
#ax.xaxis.set_minor_formatter(dates.DateFormatter('%d\n%a'))
#ax.xaxis.grid(True, which="minor")
#ax.yaxis.grid()
#ax.xaxis.set_major_locator(dates.DayLocator())
#ax.xaxis.set_major_formatter(dates.DateFormatter('\n\n%a\%b\%Y'))
ax.xaxis.set_major_formatter( matplotlib.dates.DateFormatter('%Y-%b-%a %H:%M:%S'))
ax.xaxis.grid(True, which="minor")
#ax.xaxis.set_minor_formatter(dates.DateFormatter('\n\n%a\%b\%Y'))
plt.xticks( rotation=25 )
plt.tight_layout()
plt.ylabel("PM10 Values",fontweight='bold')
plt.title("PM10 Prediction for Small Time interval During Training",fontweight='bold')
plt.xlabel('Date',fontweight='bold')
plt.show()
# -
# # Plots predictions during Training
# +
#fig = plt.figure(figsize=(30, 2))
plt.rcParams["figure.figsize"] = (8,5)
fig, ax = plt.subplots()
ax.plot(x_axis[0:80000], network.reshape(80000,1)[0:80000], 'r-',color = 'deepskyblue',label="Traninning Prediction")
ax.plot_date(x_axis[0:80000], actual.reshape(80000,1)[0:80000], 'b-',color='goldenrod',label='Actual Prediction')
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.,fontsize='x-large')
#ax.xaxis.set_minor_locator(dates.MonthLocator())
#ax.xaxis.set_minor_formatter(dates.DateFormatter('%H:%M:%S'))
#ax.xaxis.set_minor_formatter(dates.DateFormatter('%d\n%a'))
#ax.xaxis.grid(True, which="minor")
#ax.yaxis.grid()
#ax.xaxis.set_major_locator(dates.DayLocator())
#ax.xaxis.set_major_formatter(dates.DateFormatter('\n\n%a\%b\%Y'))
ax.xaxis.set_major_formatter( matplotlib.dates.DateFormatter('%Y-%b-%a %H:%M:%S'))
ax.xaxis.grid(True, which="minor")
#ax.xaxis.set_minor_formatter(dates.DateFormatter('\n\n%a\%b\%Y'))
plt.xticks( rotation=25 )
plt.tight_layout()
plt.ylabel("PM10 values",fontweight='bold')
plt.xlabel("Date",fontweight='bold')
plt.title("PM10 Training prediction and Actual Value Plots",fontweight='bold')
plt.show()
# -
# # Model Testing
# +
# tf.get_default_session()
# -
validation_set = Test_eval[-Remainder:-48]
#Reset counter
counter = 0
# +
##Testing
#Flush LSTM state
sess.run(lstm_state1.assign(tf.zeros([1, lstm_layer1.state_size])))
# +
actual_output_test = []
network_output_test = []
for i in range(len(validation_set)):
input_v, output_v = get_pair(validation_set)
_, network_output = sess.run([lstm_update_op1,
final_output],
feed_dict = {
input_layer: input_v,
correct_output: output_v})
actual_output_test.append(output_v)
network_output_test.append(network_output)
# -
actual_output_test[-48:];
# +
import matplotlib.pyplot
fig = plt.figure(figsize=(30, 2))
fig, ax = plt.subplots()
ax.plot( np.array(network_output_test[-48:]).reshape(48,1), 'r-',color = 'deepskyblue',label='Predicted Values for 48 hrs Time Frame')
ax.plot( np.array(actual_output_test[-48:]).reshape(48,1), 'b-',color = 'goldenrod',label='Actual Values 48 hours Time Frame')
#ax.xaxis.set_minor_locator(dates.MonthLocator())
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.,fontsize='x-large')
#ax.xaxis.set_minor_formatter(dates.DateFormatter('%H:%M:%S'))
#ax.xaxis.set_minor_formatter(dates.DateFormatter('%d\n%a'))
#ax.xaxis.grid(True, which="minor")
#ax.yaxis.grid()
#ax.xaxis.set_major_locator(dates.DayLocator())
#ax.xaxis.set_major_formatter(dates.DateFormatter('\n\n%a\%b\%Y'))
#ax.xaxis.set_major_formatter(dates.DateFormatter('%Y-%b-%a %H:%M:%S'))
#ax.xaxis.grid(True, which="minor")
plt.title("PM10 Validation prediction and Actual Value Plots",fontweight='bold')
#ax.xaxis.set_minor_formatter(dates.DateFormatter('\n\n%a\%b\%Y'))
plt.ylabel("PM10 values",fontweight='bold')
plt.xlabel("Time/hr",fontweight='bold')
#plt.xticks( rotation=25 )
#plt.tight_layout()
plt.show()
# -
# +
import matplotlib.pyplot
fig = plt.figure(figsize=(30, 2))
fig, ax = plt.subplots()
ax.plot( np.array(network_output_test).reshape(len(network_output_test),1), 'r-',color = 'deepskyblue',label="Validation Prediction")
ax.plot( np.array(actual_output_test).reshape(len(actual_output_test),1), 'b-',color = 'goldenrod',label='Actual Values ')
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.,fontsize='x-large')
#ax.xaxis.set_minor_locator(dates.MonthLocator())
#ax.xaxis.set_minor_formatter(dates.DateFormatter('%H:%M:%S'))
#ax.xaxis.set_minor_formatter(dates.DateFormatter('%d\n%a'))
#ax.xaxis.grid(True, which="minor")
#ax.yaxis.grid()
#ax.xaxis.set_major_locator(dates.DayLocator())
#ax.xaxis.set_major_formatter(dates.DateFormatter('\n\n%a\%b\%Y'))
#ax.xaxis.set_major_formatter(dates.DateFormatter('%Y-%b-%a %H:%M:%S'))
#ax.xaxis.grid(True, which="minor")
plt.title("PM10 Validation prediction and Actual Value Plots",fontweight='bold')
#ax.xaxis.set_minor_formatter(dates.DateFormatter('\n\n%a\%b\%Y'))
plt.ylabel("PM10 values",fontweight='bold')
plt.xlabel("Time/hr for Validation Sample",fontweight='bold')
#ax.xaxis.set_minor_formatter(dates.DateFormatter('\n\n%a\%b\%Y'))
#plt.xticks( rotation=25 )
#plt.tight_layout()
plt.show()
# -
from scipy.interpolate import interp1d
# +
xn_ax = np.linspace(0,48,48*2)
# -
x_ax = np.linspace(0,50,48)
y_cor=np.array(network_output_test[-48:]).reshape(48,1)
y_cor=np.reshape(y_cor,(48))
# +
# new x values
xn_ax = np.linspace(1,48,48*10)
# new y values
yn_cor = interp1d(x_ax,y_cor, kind='cubic')
# -
plt.plot(xn_ax, yn_cor(xn_ax),color='deepskyblue',label='Smothened Validation set')
plt.xticks(x_ax, x_axis[-48:], size='small', rotation=70)
plt.tight_layout()
plt.xlabel('date',fontweight='bold')
plt.ylabel('PM Values', fontweight='bold')
plt.title('Smothened PM Value Prediction',fontweight='bold')
plt.show
# ### Testing
n_list = 48;
Feature_eval=copy.deepcopy(Test_eval[-test_part:])
def get_pair_Test(Target):
"""
Returns an (current, target)
Target is lag steps 24 ahead of current
"""
global sliding_window_test
#sliding_window_test.append(dataGetter(Target))
#print(len(sliding_window))
#sliding_window_featureset.append(dataGetter_target(Target2))
#input_value = sliding_window_test
input_value=np.array(sliding_window_test)
#print(input_value.shape)
#print(input_value.shape)
input_value=np.reshape(input_value,(1,len(sliding_window_test)))
mx = np.ma.masked_invalid(input_value)
##print(input_value)
#output_value = sliding_window[-1]
#my = np.ma.masked_invalid(output_value)
####remove last value from sliding_window
#_ = sliding_window_featureset[-1]
##print(output_value)
# sliding_window = sliding_window[1:]
#sliding_window_featureset = sliding_window_featureset[1:]
#print(len(sliding_window))
# Process output
mask_x = ~mx.mask # let mask return true for those non-nan values
## y output
#mask_y =~my.mask
#mask_yfloat = mask_y
return input_value
counter_test=0;
def dataGetter_test(datalist):
"""
Returns a single PM10 value.
Input: PM10 Data list named Target_list above
Output: single PM10 value
"""
global counter_test;
assert type(datalist) == list;
value = datalist[counter_test]
#print(counter)
counter_test = counter_test +1
return value
sliding_window_test = []
lag=25 # 24 hours time step from 0 to 24 plus the window_step_move (25) which is to be predicted
window_move_step = 1
for i in range(lag - window_move_step):
temp=dataGetter_test(Feature_eval)
sliding_window_test.append(temp);
previous_sliding_window_length=24;
time_step_shift = 24;
len(sliding_window_test)
# +
##Testing
#Flush LSTM state
sess.run(lstm_state1.assign(tf.zeros([1, lstm_layer1.state_size])));
# +
actual_output_test = []
network_output_test = []
for i in range(previous_sliding_window_length):
input_v = get_pair_Test(sliding_window_test)
_, network_output = sess.run([lstm_update_op1,
final_output],
feed_dict = {
input_layer: input_v})
array_store=np.array(Feature_eval[time_step_shift])
#print(array_store)
array_store[[0]] = network_output # replace real PM10 values with predicted values
#print(array_store.shape)
list_store = array_store.tolist() # convert to list
#print(len(list_store))
sliding_window_test.append( list_store) # append new values for prediction
#print(len(sliding_window_test))
sliding_window_test.pop(0) # remove the first value of the list
#print(len(sliding_window_test))
time_step_shift = time_step_shift + 1; # increae the time step
network_output_test.append(network_output)
# -
# +
import matplotlib.pyplot
fig = plt.figure(figsize=(30, 2))
fig, ax = plt.subplots()
ax.plot( np.array(network_output_test).reshape(len(network_output_test),1), 'r-',color = 'deepskyblue',label=" Prediction")
ax.plot(np.array(Feature_eval[-24:]).reshape(len(Feature_eval[-24:]),1), 'b-',color = 'goldenrod',label="Actual Value")
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.,fontsize='x-large')
#ax.xaxis.set_minor_locator(dates.MonthLocator())
#ax.xaxis.set_minor_formatter(dates.DateFormatter('%H:%M:%S'))
#ax.xaxis.set_minor_formatter(dates.DateFormatter('%d\n%a'))
#ax.xaxis.grid(True, which="minor")
#ax.yaxis.grid()
#ax.xaxis.set_major_locator(dates.DayLocator())
#ax.xaxis.set_major_formatter(dates.DateFormatter('\n\n%a\%b\%Y'))
#ax.xaxis.set_major_formatter(dates.DateFormatter('%Y-%b-%a %H:%M:%S'))
#ax.xaxis.grid(True, which="minor")
plt.title("PM10 Test prediction and Actual Value Plots (24hr Time Frame)",fontweight='bold')
#ax.xaxis.set_minor_formatter(dates.DateFormatter('\n\n%a\%b\%Y'))
plt.ylabel("PM10 values",fontweight='bold')
plt.xlabel("Time/hr for Test Sample",fontweight='bold')
#ax.xaxis.set_minor_formatter(dates.DateFormatter('\n\n%a\%b\%Y'))
#plt.xticks( rotation=25 )
#plt.tight_layout()
plt.show()
# -
from sklearn.metrics import mean_squared_error
mean_squared_error(np.array(network_output_test).reshape(len(Target_list[-24:]),1),np.array(Target_list[-24:]).reshape(len(network_output_test),1) )**0.5
import scipy
def rsquared(x, y):
""" Return R^2 where x and y are array-like."""
slope, intercept, r_value, p_value, std_err = scipy.stats.linregress(x, y)
return r_value**2
rsquared(np.array(network_output_test).reshape(len(Target_list[-24:]),),
np.array(Target_list[-24:]).reshape(len(network_output_test),) )
rsquared(np.array(network_output_test).reshape(len(Target_list[-24:]),),
np.array(Target_list[-24:]).reshape(len(network_output_test),) )
# +
import scipy.stats as stats
stats.f_oneway(np.array(network_output_test).reshape(len(Target_list[-24:]),),
np.array(Target_list[-24:]).reshape(len(network_output_test),))
# -
| LSTM_RNN/2_prediction_using_only_PM10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:dl-minicourse] *
# language: python
# name: conda-env-dl-minicourse-py
# ---
# # Signal echoing
#
# Echoing signal `n` steps is an example of synchronized many-to-many task.
# +
from res.sequential_tasks import EchoData
import torch
import torch.nn as nn
# import torch.nn.functional as F
import torch.optim as optim
torch.manual_seed(1);
# +
batch_size = 5
echo_step = 3
series_length = 20_000
BPTT_T = 20
train_data = EchoData(
echo_step=echo_step,
batch_size=batch_size,
series_length=series_length,
truncated_length=BPTT_T,
)
train_size = len(train_data)
test_data = EchoData(
echo_step=echo_step,
batch_size=batch_size,
series_length=series_length,
truncated_length=BPTT_T,
)
test_size = len(test_data)
# -
# Let's print first 20 timesteps of the first sequences to see the echo data:
print('(1st input sequence) x:', *train_data.x_batch[0, :20], '... ')
print('(1st target sequence) y:', *train_data.y_batch[0, :20], '... ')
# batch_size different sequences are created:
print('x_batch:', *(str(d)[1:-1] + ' ...' for d in train_data.x_batch[:, :20]), sep='\n')
print('x_batch size:', train_data.x_batch.shape)
print()
print('y_batch:', *(str(d)[1:-1] + ' ...' for d in train_data.y_batch[:, :20]), sep='\n')
print('y_batch size:', train_data.y_batch.shape)
# In order to use RNNs data is organized into temporal
# chunks of size [batch_size, T, feature_dim]
print('x_chunk:', *train_data.x_chunks[0].squeeze(), sep='\n')
print('1st x_chunk size:', train_data.x_chunks[0].shape)
print()
print('y_chunk:', *train_data.y_chunks[0].squeeze(), sep='\n')
print('1st y_chunk size:', train_data.y_chunks[0].shape)
class SimpleRNN(nn.Module):
def __init__(self, input_size, rnn_hidden_size, output_size):
super().__init__()
self.rnn_hidden_size = rnn_hidden_size
self.rnn = torch.nn.RNN(
input_size=input_size,
hidden_size=rnn_hidden_size,
num_layers=1,
nonlinearity='relu',
batch_first=True
)
self.linear = torch.nn.Linear(
in_features=rnn_hidden_size,
out_features=1
)
def forward(self, x, hidden):
x, hidden = self.rnn(x, hidden)
x = self.linear(x)
return x, hidden
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
def train(hidden):
model.train()
correct = 0
for batch_idx in range(train_size):
data, target = train_data[batch_idx]
data, target = torch.from_numpy(data).float().to(device), torch.from_numpy(target).float().to(device)
optimizer.zero_grad()
if hidden is not None: hidden.detach_()
logits, hidden = model(data, hidden)
loss = criterion(logits, target)
loss.backward()
optimizer.step()
pred = (torch.sigmoid(logits) > 0.5)
correct += (pred == target.byte()).int().sum().item()
return correct, loss.item(), hidden
def test(hidden):
model.eval()
correct = 0
with torch.no_grad():
for batch_idx in range(test_size):
data, target = test_data[batch_idx]
data, target = torch.from_numpy(data).float().to(device), torch.from_numpy(target).float().to(device)
logits, hidden = model(data, hidden)
pred = (torch.sigmoid(logits) > 0.5)
correct += (pred == target.byte()).int().sum().item()
return correct
# +
feature_dim = 1 #since we have a scalar series
h_units = 4
model = SimpleRNN(
input_size=1,
rnn_hidden_size=h_units,
output_size=feature_dim
).to(device)
hidden = None
criterion = torch.nn.BCEWithLogitsLoss()
optimizer = torch.optim.RMSprop(model.parameters(), lr=0.001)
# +
n_epochs = 5
epoch = 0
while epoch < n_epochs:
correct, loss, hidden = train(hidden)
epoch += 1
train_accuracy = float(correct) / train_size
print(f'Train Epoch: {epoch}/{n_epochs}, loss: {loss:.3f}, accuracy {train_accuracy:.1f}%')
#test
correct = test(hidden)
test_accuracy = float(correct) / test_size
print(f'Test accuracy: {test_accuracy:.1f}%')
# -
# Let's try some echoing
my_input = torch.empty(1, 100, 1).random_(2).to(device)
hidden = None
my_out, _ = model(my_input, hidden)
print(my_input.view(1, -1).byte(), (my_out > 0).view(1, -1), sep='\n')
| PyTorch_examples/pytorch-Deep-Learning-master/09-echo_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
from papyri.crosslink import IngestedBlobs
from papyri.config import ingest_dir
from papyri.crosslink import TreeReplacer
from papyri.graphstore import GraphStore
from json import loads, dumps
from papyri.take2 import RefInfo
import json
gs = GraphStore(ingest_dir)
key = ('scipy', '1.5.0', 'module', 'scipy.signal.filter_design.cheb2ord')
key = 'numpy/1.20.0/api/numpy.polyfit'.replace('api','module').split('/')
data = gs.get(key)
data = json.loads(data.decode())
gbr_data = gs.get_backref(key)
data['backrefs'] = [RefInfo(*x).to_json() for x in gbr_data]
blob = IngestedBlobs.from_json(data)
def cgraph(gs, blob, key):
nodes_names = [b.path for b in blob.backrefs+blob.refs]+[key[3]]
#nodes_names = [n for n in nodes_names if n.startswith('numpy')]
raw_edges = []
for k in blob.backrefs+blob.refs:
orig = [x[3] for x in gs.get_backref(tuple(k))]
for o in orig:
raw_edges.append((k.path, o))
data = {'nodes':[], 'links':[]}
nums_ = set()
edges = list(raw_edges)
nodes = list(set(nodes_names))
for a,b in edges:
if (a not in nodes) or (b not in nodes):
continue
nums_.add(a)
nums_.add(b)
nums = {x:i for i,x in enumerate(nodes, start=1)}
for i,(from_,to) in enumerate(edges[:N]):
if (from_ not in nodes):
continue
if (to not in nodes):
continue
if key[3] in (to, from_):
continue
data['links'].append({"source":nums[from_],"target":nums[to], "id":i})
x = nums.keys()
for node in nodes:
diam = 8
if node == key[3]:
diam = 18
data['nodes'].append({"id":nums[node],"val":diam, "label":node, "mod":'.'.join(node.split('.')[0:1])})
return data
data = cgraph(gs, blob, key)
with open('d3.json', 'w') as f:
f.write(dumps(data))
len(data['nodes']), len(data['links'])
# -
# cat d3.json
pwd
raw_edges
sa = blob.see_also[2]
sa.ref
'numpy.ma.core.asanyarray' in nodes, nodes
nid = [n['id'] for n in data['nodes']]
tid = [e['target'] for e in data['edges']]
sid = [e['source'] for e in data['edges']]
set(tid) - set(nid)
edges[:N]
len(data['edges']),len(data['nodes'])
with open('JavaScript-Network-Graphs/assets/data/user_network.json') as f:
dd = json.loads(f.read())
dd
# !cat npg.json
| GraphTest1-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Cutting Cloud Costs with Infrastructure Automation (Part I - AWS Infrastructure Automation)
# ## Review
# 2 posts ago, we discussed the concept of cloud and how cheap cloud could be by looking at a very specific use case of the compute-optimized p2.xlarge EC2 instance. I determined that, at a spot-price cost of \$0.17 and an incompetency of having to run the instance for 5 hours to train my model, I'd still be spending less than a dollar. I started thinking about how I could continue cutting costs to _**ensure**_ that I wouldn't be spending more than a few cents every time I need to train a model. If I'm looking at a dollar, that could still quickly add up if I somehow end up having to spin up an instance every day. At a dollar a day, that's still \$365 that could go towards something else... That could buy me 365 items from the dollar store!
#
# [](https://www.youtube.com/watch?v=-qtrAMK7_Qk)
# I don't mean to sound cheap or anything (in real life I'm actually really cheap, but I don't mean to sound cheap here), but it's just good business practice to think through a solution and find any points of weakness that might exist. This conversation might start with a dollar or two, but we have to consider the end-to-end process of exactly how this will work because, as soon as we spin up that EC2, we're on the clock. Thinking through the process, there is a lot we have to do to even start training the model...
# 1. Create a VPC with an internet gateway and default route so our application is accessible via public internet
# 2. Create default subnet so our EC2 can be assigned a public IPv4 address
# 3. Set default ACL and Security Group settings
# 4. Spin up EC2
# 5. Set up credentials
# 6. Install python, CUDA
# 7. Install common python libraries: pandas, matplotlib, numpy
# 8. Install domain-specific python libraries: tflearn, tensorflow, sklearn
# 9. Install additional tools: jupyter
# 10. Configure tensorflow to work with GPU
# 11. Import data into EC2
# 12. Write all code to ETL data
# 13. Write all code to train model
# 14. Save model / results
#
# Man... that's a lot of steps right there. Is there a chance we can possibly automate some of this? The short answer from some previous experiences I've had and materials I've read is yes and yes.
#
# Steps 1 - 5 essentially have to do with AWS, steps 6 - 10 have to do with setting up the virtual machine itself, and steps 11 - 14 are executing the code itself and training the model. There will be different ways to automate each batch of steps here, so let's go through each of these one by one over the next 3 posts. In this post, I will discuss AWS infrastructure automation (steps 1 - 5).
#
# ## AWS Infrastructure Automation - Terraform
# Terraform is a way of capturing an entire AWS infrastructure in code. I was pretty blown away by the fact that this exists, because you can literally spin up an entire infrastructure in seconds. It almost feels like a cheat code, especially with some of the work experience I've had in the past working for the internet service provider.
#
# To implement a route, you had to speak to multiple teams and open up various tickets and maybe wait up to 2-3 weeks for someone to work through a queue and get to your request. To add storage capacity to a server cluster, we might have had to work with the vendor to procure an additional hardware (where we'd have a vendor manager who negotiated a bulk rate), wait a few weeks for the hardware to arrive, have an entire lab team rack the server, cable it up to connect to the network, possibly have an implementation consultant come in and configure the server, and if the server ever breaks the consultant would have to come back and fix it or we'd order a new one and go through this entire process again. Accurate depiction of daily life below:
#
# <img src="https://us.123rf.com/450wm/flairmicro/flairmicro1503/flairmicro150300669/38023860-frustrated-technician-working-in-server-room-of-data-center.jpg" style="width: 400px;"/>
#
# This is obviously an oversimplification of the topic, and there are reasons why the service provider would not be able to use AWS, but maybe it's more of a commentary on the stuff I like to do that gets me going. There are folks who specialize in hardware and there are good reasons why you'd want your own dedicated hardware... For a service provider, they are literally the ones building out this infrastructure across the country so people across the country can have internet access, so they would never be building off AWS' network because, well, they _**are**_ the network for that region.
#
# As a data scientist, our objective is to _**solve a business problem**_. Although this is melding quite a bit with technology, your traditional data scientist probably would like to stay as close to their objective and have as least distractions as possible. If we can automate the build of our infrastructure and networking, _**why wouldn't we**_?
#
# Back to Terraform now... lol. Yes, Terraform lets us define our network and infrastructure in code, and run the code to spin up everything at once. You define your AWS infrastructure in a _**.tf**_ file using the _**HashiCorp Configuration Language (HCL)**_. A nice feature of Terraform is that changes are made incrementally, and this can be illustrated pretty simply by an example: Let's say we have a base infrastructure set up with a VPC, internet gateway, ACLs and Security Groups and we spin that up with one EC2... at some point, we'd probably want to add another EC2 right? We probably wouldn't want to tear the entire thing down and spin everything back up. The first EC2 could already be in production! If we run the same HCL script with the additional EC2 added, Terraform has the right context to say "everything in this configuration file already exists except for this second EC2, so I'm only going to act on this EC2 and leave everything else as it is".
#
# So we talked a lot about all the different AWS components, but we haven't really tried to visualize it. The fact of the matter is, each EC2 has to live within a security group, which has to live within a VPC, which has to have a gateway, default routes, and ACLs. That's just all the default networking components we actually need to even get an EC2 online, so let's draw an extremely simple diagram as to what's happening here.
#
# <img src="https://s3.ca-central-1.amazonaws.com/2017edmfasatb/chi_lars_face_detection/images/32_aws_arch_diagram_1.png" style="width: 800px;"/>
#
# Pretty simple, right? My laptop goes through the TELUS cloud and the Amazon cloud to eventually hit the internet gateway within my VPC. Then, within my VPC, I have ACLs and Security Groups which both (for now until I need more security) let any traffic in and out from any host and port to any other host and port. Since the EC2 is open to the public internet, I should be able to reach it via a public IPv4 address or DNS host name.
#
# The Terraform HCL code that is used to generate this infrastructure can be found under the _**infrastructure**_ directory in this project's root. Let's tackle the sections of Terraform code:
#
# ### Initial Variables
# ~~~~
# variable "myRegion" {
# type = "string"
# default = "us-east-1"
# }
#
# variable "myCidrBlock" {
# type = "string"
# default = "10.0.0.0/24"
# }
#
# variable "myKeyPair" {
# type = "string"
# default = "ec2-user"
# }
# ~~~~
#
# It's useful to set some initial variables within Terraform, and it's quite nice that Terraform actually allows us to define variables to aid in the set up of our infrastructure. These variables can be used in the main code to define parameters of our AWS components. Here, I've only set 3 variables:
# - The region we want to set up the infrastructure in (us-east-1 has the most services available to it, and we don't really care about region anyways because we're not dealing with any sensitive data per se, so why not go with AWS' flagship region for now)
# - A /24 group of IPv4 addresses of the format 10.0.0.X, which will act as our subnet within our VPC
# - The SSH key pair to use for our EC2, something that is already defined under AWS EC2 key pairs
#
# ### AWS Configuration
# ~~~~
# provider "aws" {
# region = "${var.myRegion}"
# }
# ~~~~
#
# All we do here is set our region. Easy enough. I have already been working in us-east-1 through some previous projects I've done on AWS, so going forward, anything I do should be in us-east-1:
#
# <img src="https://s3.ca-central-1.amazonaws.com/2017edmfasatb/chi_lars_face_detection/images/33_aws_region.png" alt="Drawing" style="width: 300px;"/>
#
# ### VPC
# ~~~~
# resource "aws_vpc" "main_vpc" {
# cidr_block = "${var.myCidrBlock}"
# instance_tenancy = "default"
# enable_dns_hostnames = true
#
# tags {
# Name = "main_vpc"
# }
# }
# ~~~~
#
# The first line of this code defines an AWS _**resource**_ of the type <em>**aws_vpc**</em> named <em>**main_vpc**</em>. This pattern is followed throughout for other resources as well.
#
# The important parts here is that we're assigning our _**/24**_ block of IP addresses (256 IPs available) to the VPC and we're allowing elements in this VPC to have a public DNS hostname (Amazon will assign a random name to it).
#
# After running the code, our VPC will be defined by these parameters, with other parameters being filled in with default values (for a project of this size and scope, we really don't need to get into too many details):
#
# <img src="https://s3.ca-central-1.amazonaws.com/2017edmfasatb/chi_lars_face_detection/images/34_aws_vpc.png" style="width: 1000px;"/>
#
# ### Internet Gateway & Default Route Table
# ~~~~
# resource "aws_internet_gateway" "main_vpc_igw" {
# vpc_id = "${aws_vpc.main_vpc.id}"
#
# tags {
# Name = "main_vpc_igw"
# }
# }
#
# resource "aws_default_route_table" "main_vpc_default_route_table" {
# default_route_table_id = "${aws_vpc.main_vpc.default_route_table_id}"
#
# route {
# cidr_block = "0.0.0.0/0"
# gateway_id = "${aws_internet_gateway.main_vpc_igw.id}"
# }
#
# tags {
# Name = "main_vpc_default_route_table"
# }
# }
#
# ~~~~
#
# This block is saying we want to assign an internet gateway to our VPC so we can send and receive traffic to the public internet, and add a route to the default route table saying traffic _**to and from any host**_ will be, by default, sent to the internet gateway to figure out and route. Simple!
#
# <img src="https://s3.ca-central-1.amazonaws.com/2017edmfasatb/chi_lars_face_detection/images/35_aws_igw.png" style="width: 1000px;"/>
#
# -----------
#
# <img src="https://s3.ca-central-1.amazonaws.com/2017edmfasatb/chi_lars_face_detection/images/36_aws_default_route.png" style="width: 1000px;"/>
#
# ### Subnet
# ~~~~
# resource "aws_subnet" "main_vpc_subnet" {
# vpc_id = "${aws_vpc.main_vpc.id}"
# cidr_block = "${var.myCidrBlock}"
# map_public_ip_on_launch = true
#
# tags {
# Name = "main_vpc_subnet"
# }
# }
# ~~~~
#
# In our subnet, we define the same IP block, but it's within the subnet that we define we want instances within this subnet to be assigned a _**public IP**_ as well. This will not be the 10.0.0.X IP that we assign because that's the _**private**_ IP referenced from within our VPC LAN, but the public IP will provide a routable IP address public hosts can reach as well.
#
# In the screenshot below, AWS shows that our /24 block of IPs has 251 IPs. I was expecting 256 addresses, but perhaps AWS reserves some of these for standard processes. I'm not quite sure. Regardless, we won't be needing anywhere close to even 10 IPs, so we should be okay.
#
# <img src="https://s3.ca-central-1.amazonaws.com/2017edmfasatb/chi_lars_face_detection/images/37_aws_subnet.png" style="width: 1000px;"/>
#
# ### ACL & Security Groups
# ~~~~
# resource "aws_default_network_acl" "main_vpc_nacl" {
# default_network_acl_id = "${aws_vpc.main_vpc.default_network_acl_id}"
# subnet_ids = ["${aws_subnet.main_vpc_subnet.id}"]
#
# ingress {
# protocol = -1
# rule_no = 1
# action = "allow"
# // cidr_block = "${var.myIp}"
# cidr_block = "0.0.0.0/0"
# from_port = 0
# to_port = 0
# }
#
# egress {
# protocol = -1
# rule_no = 2
# action = "allow"
# cidr_block = "0.0.0.0/0"
# from_port = 0
# to_port = 0
# }
#
# tags {
# Name = "main_vpc_nacl"
# }
# }
#
# resource "aws_default_security_group" "main_vpc_security_group" {
# vpc_id = "${aws_vpc.main_vpc.id}"
#
# ingress {
# protocol = "-1"
# // cidr_blocks = ["${var.myIp}"]
# cidr_blocks = ["0.0.0.0/0"]
# from_port = 0
# to_port = 0
# }
#
# egress {
# protocol = "-1"
# cidr_blocks = ["0.0.0.0/0"]
# from_port = 0
# to_port = 0
# }
#
# tags {
# Name = "main_vpc_security_group"
# }
# }
# ~~~~
#
# I'm going to group these two together because they are basically both security measures that are saying the same thing. Keep in mind that the ACL is actually be implemented on the subnet within the VPC, and the Security Group is a set of rules one level deeper than the VPC, but for the purposes of our project, we're keeping security as a very low priority and we're just allowing any traffic from anywhere on any port.
#
# Some details to note:
# - A protocol of "-1" means "any protocol"
# - An IP of 0.0.0.0/0 means "any IP"
#
# <img src="https://s3.ca-central-1.amazonaws.com/2017edmfasatb/chi_lars_face_detection/images/38_aws_acl.png" style="width: 1000px;"/>
#
# ----------
#
# <img src="https://s3.ca-central-1.amazonaws.com/2017edmfasatb/chi_lars_face_detection/images/39_aws_security_group.png" style="width: 1000px;"/>
#
# ### EC2 Spot Instance Request
# ~~~~
# resource "aws_spot_instance_request" "aws_deep_learning_custom_spot" {
# ami = "ami-45fdf753"
# spot_price = "0.20"
# instance_type = "p2.xlarge"
# security_groups = ["${aws_default_security_group.main_vpc_security_group.id}"]
# subnet_id = "${aws_subnet.main_vpc_subnet.id}"
# key_name = "${var.myKeyPair}"
#
# tags {
# Name = "aws_deep_learning_custom_spot"
# }
# }
# ~~~~
#
# Our final piece. The EC2 spot instance! Simple enough here as well as Terraform makes the code pretty easily understandable. This spot request is referencing a specific _**AMI**_ (which I will go through in the next section) at a spot price of _**$0.20 / hr**_ with the instance type _**p2.xlarge**_ that we've already explored for GPU compute optimization. It will belong in the security group and subnets defined before and use the EC2 key-pair tied with the user _**ec2-user**_.
#
# <img src="https://s3.ca-central-1.amazonaws.com/2017edmfasatb/chi_lars_face_detection/images/40_aws_spot_request.png" style="width: 1000px;"/>
#
# ----------
#
# <img src="https://s3.ca-central-1.amazonaws.com/2017edmfasatb/chi_lars_face_detection/images/41_aws_instance.png" style="width: 1000px;"/>
#
# ## Summary
# At this point, we should have an up and running EC2 instance that is publically accessible by internet, but perhaps more importantly, we've built a infrastructure skeleton that we can now modify to easily implement various security measures as well! In the next post, I'll actually be diving into this EC2 to see how can set the instance up with all the tools we need.
| notebook/6 - Cutting Cloud Costs with Infrastructure Automation (Part I - AWS Infrastructure Automation).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Demo for the DoWhy causal API
# We show a simple example of adding a causal extension to any dataframe.
# +
import dowhy.datasets
import dowhy.api
import numpy as np
import pandas as pd
from statsmodels.api import OLS
# +
data = dowhy.datasets.linear_dataset(beta=5,
num_common_causes=1,
num_instruments = 0,
num_samples=1000,
treatment_is_binary=True)
df = data['df']
df['y'] = df['y'] + np.random.normal(size=len(df)) # Adding noise to data. Without noise, the variance in Y|X, Z is zero, and mcmc fails.
#data['dot_graph'] = 'digraph { v ->y;X0-> v;X0-> y;}'
treatment= data["treatment_name"][0]
outcome = data["outcome_name"][0]
common_cause = data["common_causes_names"][0]
df
# -
# data['df'] is just a regular pandas.DataFrame
df.causal.do(x=treatment,
variable_types={treatment: 'b', outcome: 'c', common_cause: 'c'},
outcome=outcome,
common_causes=[common_cause],
proceed_when_unidentifiable=True).groupby(treatment).mean().plot(y=outcome, kind='bar')
df.causal.do(x={treatment: 1},
variable_types={treatment:'b', outcome: 'c', common_cause: 'c'},
outcome=outcome,
method='weighting',
common_causes=[common_cause],
proceed_when_unidentifiable=True).groupby(treatment).mean().plot(y=outcome, kind='bar')
# +
cdf_1 = df.causal.do(x={treatment: 1},
variable_types={treatment: 'b', outcome: 'c', common_cause: 'c'},
outcome=outcome,
dot_graph=data['dot_graph'],
proceed_when_unidentifiable=True)
cdf_0 = df.causal.do(x={treatment: 0},
variable_types={treatment: 'b', outcome: 'c', common_cause: 'c'},
outcome=outcome,
dot_graph=data['dot_graph'],
proceed_when_unidentifiable=True)
# -
cdf_0
cdf_1
# ## Comparing the estimate to Linear Regression
# First, estimating the effect using the causal data frame, and the 95% confidence interval.
(cdf_1['y'] - cdf_0['y']).mean()
1.96*(cdf_1['y'] - cdf_0['y']).std() / np.sqrt(len(df))
# Comparing to the estimate from OLS.
model = OLS(np.asarray(df[outcome]), np.asarray(df[[common_cause, treatment]], dtype=np.float64))
result = model.fit()
result.summary()
| docs/source/example_notebooks/dowhy_causal_api.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
pd.options.display.float_format = '{:.3f}'.format
avocado = pd.read_csv("dataset/avocado_2015_to_2020.csv")
avocado.head()
avocado = avocado.rename(columns={"4046": "Small/Medium Hass", "4225": "Large Hass", "4770": "Extra Large Hass"})
avocado = avocado[avocado["Region"] != "TotalUS"]
avocado.head()
month_order = avocado["Month"].unique()
year_order = avocado["Year"].unique()
average_total_per_year_month_region = avocado.groupby(["Year", "Month", "Region"])\
.agg({"AveragePrice": "mean", "Total Volume": "sum"})\
.reindex(month_order, level=1)
average_total_per_year_month_region["Total Value (Million $)"] = average_total_per_year_month_region["AveragePrice"] *\
average_total_per_year_month_region["Total Volume"] /\
1000000
average_total_per_year_month_region = average_total_per_year_month_region.reset_index()
average_total_per_year_month_region.head()
plt.figure(figsize=(15, 7))
sns.pointplot(x= "Month", y= "AveragePrice",hue= "Year", data = average_total_per_year_month_region,
palette="Set1", ci= False)
plt.show()
plt.figure(figsize=(15, 7))
sns.lineplot(x= "Year", y= "Total Volume", data = average_total_per_year_month_region, palette="Set1")
plt.show()
plt.figure(figsize=(18, 7))
sns.barplot(x= "Region", y= "AveragePrice", data = average_total_per_year_month_region, ci= False,
palette=sns.color_palette("Paired", n_colors=53, desat=.5),
order= average_total_per_year_month_region.groupby("Region")["AveragePrice"].mean().sort_values(ascending= False).index)
plt.xticks(rotation = 90)
plt.show()
avocado["Region"].nunique()
average_total_per_year_month_region.groupby("Region")["AveragePrice"].mean().sort_values(ascending= False)
| avocado_data_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''base'': conda)'
# language: python
# name: python3
# ---
# +
def load(fname):
rules = []
with open(fname) as f:
template = f.readline().strip()
for ll in f:
if ll.strip() == "":
continue
rule = [ x.strip() for x in ll.split("->") ]
rules.append(rule)
rules = dict(rules)
return template, rules
template, rules = load("../dat/day14.txt")
def pairwise(iterable): # itertools.pairwise is not available until version 3.10+
iterator = iter(iterable)
try:
first = next(iterator)
second = next(iterator)
while second != None:
yield first, second
first = second
second = next(iterator)
except StopIteration:
pass
# +
def iterate(template, steps):
"""
Brute force method that will run out of memory if 'steps' becomes large.
"""
for ss in range(steps):
template = list(template)
for ii in range(len(template)-1, 0, -1):
pair = template[ii-1] + template[ii]
insertion = rules.get(pair)
if insertion:
template.insert(ii, insertion)
return "".join(template)
result = iterate(template, 10)
ans = {}
for xx in result:
ans[xx] = ans.get(xx, 0) + 1
min = (None, None)
max = (None, None)
for kk, vv in ans.items():
if min[1] is None or vv < min[1]:
min = (kk, vv)
if max[1] is None or vv > max[1]:
max = (kk, vv)
print(max[1] - min[1])
# +
def xiterate(template, steps):
"""
"""
# convert template into pairs of letters
_template = {}
for pair in pairwise(template):
pair = "".join(pair)
_template[pair] = _template.get(pair, 0) + 1
for ss in range(steps):
for tt, cc in list(_template.items()):
insertion = rules.get(tt)
if insertion:
_template[tt[0] + insertion] = _template.get(tt[0] + insertion, 0) + cc
_template[insertion + tt[1]] = _template.get(insertion + tt[1], 0) + cc
_template[tt] -= cc
return _template
result = xiterate(template, 40)
print(result)
ans = {}
for kk, vv in result.items():
ans[kk[0]] = ans.get(kk[0], 0) + vv
ans[template[-1]] = ans.get(template[-1], 0) + 1
print(ans)
min = (None, None)
max = (None, None)
for kk, vv in ans.items():
if min[1] is None or vv < min[1]:
min = (kk, vv)
if max[1] is None or vv > max[1]:
max = (kk, vv)
print(max[1] - min[1])
| python/day14.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ur_env
# language: python
# name: ur_env
# ---
# # The Royal game of Ur, you vs TD-Ur
#
# ## Instructions
#
# - To play a move, click the square you want to move
# - To make TD-Ur play its move, click its name, or to make it do this automatically, click auto-play on the bottom left
# - When you cannot make any moves, pass by clicking 'you'
from td_ur import interactive
game = interactive.InteractiveGame()
game.play()
| play_Ur_local.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # HP 8722D pyVNA tutorial
# Import the things we need - pyVNA is the package of interest.
# You might need to add its location to path manually, but not in this case.
import sys,time,struct,math,numpy as np,matplotlib.pyplot as plt
from timeit import default_timer as timer
#sys.path.append('/home/pi/Desktop/ADRNX/VNA')
from pyVNA import core,constants
# Initialize vna object, overrding some defaults. Debug flag turns on/off print statements.
vna = core.VNA(dev='/dev/ttyUSB2',debug=False)
# Set up gpib adapter comms
vna.init_and_setup()
# Preset VNA to factory defaults - command blocks until this completes or timeout of 5s
vna.preset(tm=5)
# ** Changing measurement type and mode **
# A special class, ``constants``, is provided for convenient enumeration of possible mode and measurement values
# For instance: '``constants.VNA_MODE.POLAR``'
# First check if desired mode or measurement type is on(true)
# Only VNA_MODE and VNA_MEAS constants are valid arguments
print(vna.check_if_true(constants.VNA_MODE.POLAR),vna.check_if_true(constants.VNA_MEAS.S21))
# Set things to what we want
vna.set_meas_mode(constants.VNA_MEAS.S21)
time.sleep(0.1)
vna.set_plot_mode(constants.VNA_MODE.POLAR)
# Verify that changes did go through (wait a bit for things to switch)
print(vna.check_if_true(constants.VNA_MODE.POLAR),vna.check_if_true(constants.VNA_MEAS.S21))
# ** Changing sweep parameters **
# +
# Set setting to what we want, printing state after each change
vna.print_current_state()
vna.set_ifbw(30)
vna.print_current_state()
vna.set_num_points(1601)
vna.print_current_state()
fmin, fmax = 2.275E3, 2.276E3
print(fmin, fmax)
vna.set_freq_range(fmin, fmax,units="MHz")
vna.print_current_state()
vna.set_power(-10)
vna.print_current_state()
# -
# ** Doing a sweep **
# After setting desired parameters, several method will use those to give useful sweep properties
# For example, we can get a rough local estimate of sweep time (quickly, for algorithm scheduling purposes)
print(vna.estimate_sweep_time())
# Or query VNA for actual value (which is by default the lowest possible sweep time for current settings)
swt = vna.get_actual_sweep_time()
print(swt)
# And also get number of bytes we expect from VNA
# Note how result of form 2 (binary transfer) is so much smaller than ASCII form 4
# (don't worry - parsing of these is handled internally, so this is just for your memory consumption calcs)
print(vna.estimate_sweep_size(form=2),vna.estimate_sweep_size(form=4))
# Now, lets start the sweep - this command returns immediately, and you can do other things while vna sweeps
# Note how sweep params dictionary is generated at this point
sweep_parameters = vna.start_sweep()
# But here, we will block to await completion with a timeout (returns -1 if timeout occurs)
timetaken = vna.await_completion(swt+10.0)
print(timetaken)
# ** Reading out data **
# Now, we need to explicitly read out the data.
# This can take awhile if run immediately after sweep, since vna is formatting things internally.
start = timer()
data2 = vna.retrieve_sweep_data(pause=0,form=2)
print(timer()-start)
# You can read out same data many times. Note how ASCII readout takes longer and is generally discouraged.
start = timer()
data4 = vna.retrieve_sweep_data(pause=0,form=4)
print(timer()-start)
# Finally, we can at a later timer parse data blob + sweep parameters to obtain results dictionary
results = vna.parse_vna_data(sweep_parameters, data2, form=2)
results.keys()
# Let's plot it!
[il,ql] = [results["I"],results["Q"]]
datatoplot = [10.0*math.log10((il[i]**2+ql[i]**2)/(50.0*.001)) for i in range(0,len(il))]
plt.figure()
plt.plot(results["F"],datatoplot,'-',lw=2)
plt.show()
# ** Saving data **
# Assuming you want to mimic the standard output, utilities are provided to create text files from results dictionary
# You can either specify folder (which exists) (and use autonaming 'vnasweep-%Y-%m-%d-%H-%M-%S.txt')
vna.save_results_to_file(results,folder='/home/pi/Desktop/DataNX/vna/tests/')
# Or, specify a full path
vna.save_results_to_file(results,fullpath='/home/pi/Desktop/DataNX/vna/tests/test.blaa')
# ** But what if I am lazy and want 1 liners? **
# +
results = vna.do_full_measurement({"ifbw": 30, "numpoints": 1601, "freqs": [500,700,"MHz"], "power": -8,\
"mode": constants.VNA_MODE.POLAR, "meas": constants.VNA_MEAS.S21, "preset": True})
# its two lines...close enough!
# -
# Let's plot it (again)!
[il,ql] = [results["I"],results["Q"]]
datatoplot = [10.0*math.log10((il[i]**2+ql[i]**2)/(50.0*.001)) for i in range(0,len(il))]
plt.figure()
plt.plot(results["F"],datatoplot,'-')
plt.show()
| VNANX_take_sweep_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python Environment
# We show here some examples of how to run Python on a Pynq platform. Python 3.6
# is running exclusively on the ARM processor.
#
# In the first example, which is based on calculating the factors and primes
# of integer numbers, give us a sense of the performance available when running
# on an ARM processor running Linux.
#
# In the second set of examples, we leverage Python's `numpy` package and `asyncio`
# module to demonstrate how Python can communicate
# with programmable logic.
#
#
# ## Factors and Primes Example
# Code is provided in the cell below for a function to calculate factors and
# primes. It contains some sample functions to calculate the factors and primes
# of integers. We will use three functions from the `factors_and_primes` module
# to demonstrate Python programming.
# +
"""Factors-and-primes functions.
Find factors or primes of integers, int ranges and int lists
and sets of integers with most factors in a given integer interval
"""
def factorize(n):
"""Calculate all factors of integer n.
"""
factors = []
if isinstance(n, int) and n > 0:
if n == 1:
factors.append(n)
return factors
else:
for x in range(1, int(n**0.5)+1):
if n % x == 0:
factors.append(x)
factors.append(n//x)
return sorted(set(factors))
else:
print('factorize ONLY computes with one integer argument > 0')
def primes_between(interval_min, interval_max):
"""Find all primes in the interval.
"""
primes = []
if (isinstance(interval_min, int) and interval_min > 0 and
isinstance(interval_max, int) and interval_max > interval_min):
if interval_min == 1:
primes = [1]
for i in range(interval_min, interval_max):
if len(factorize(i)) == 2:
primes.append(i)
return sorted(primes)
else:
print('primes_between ONLY computes over the specified range.')
def primes_in(integer_list):
"""Calculate all unique prime numbers.
"""
primes = []
try:
for i in (integer_list):
if len(factorize(i)) == 2:
primes.append(i)
return sorted(set(primes))
except TypeError:
print('primes_in ONLY computes over lists of integers.')
def get_ints_with_most_factors(interval_min, interval_max):
"""Finds the integers with the most factors.
"""
max_no_of_factors = 1
all_ints_with_most_factors = []
# Find the lowest number with most factors between i_min and i_max
if interval_check(interval_min, interval_max):
for i in range(interval_min, interval_max):
factors_of_i = factorize(i)
no_of_factors = len(factors_of_i)
if no_of_factors > max_no_of_factors:
max_no_of_factors = no_of_factors
results = (i, max_no_of_factors, factors_of_i,\
primes_in(factors_of_i))
all_ints_with_most_factors.append(results)
# Find any larger numbers with an equal number of factors
for i in range(all_ints_with_most_factors[0][0]+1, interval_max):
factors_of_i = factorize(i)
no_of_factors = len(factors_of_i)
if no_of_factors == max_no_of_factors:
results = (i, max_no_of_factors, factors_of_i, \
primes_in(factors_of_i))
all_ints_with_most_factors.append(results)
return all_ints_with_most_factors
else:
print_error_msg()
def interval_check(interval_min, interval_max):
"""Check type and range of integer interval.
"""
if (isinstance(interval_min, int) and interval_min > 0 and
isinstance(interval_max, int) and interval_max > interval_min):
return True
else:
return False
def print_error_msg():
"""Print invalid integer interval error message.
"""
print('ints_with_most_factors ONLY computes over integer intervals where'
' interval_min <= int_with_most_factors < interval_max and'
' interval_min >= 1')
# -
# Next we will call the factorize() function to calculate the factors of an integer.
factorize(1066)
# The primes_between() function can tell us how many prime numbers there are in an
# integer range. Let’s try it for the interval 1 through 1066. We can also use one
# of Python’s built-in methods len() to count them all.
len(primes_between(1, 1066))
# Additionally, we can combine len() with another built-in method, sum(), to calculate
# the average of the 180 prime numbers.
primes_1066 = primes_between(1, 1066)
primes_1066_average = sum(primes_1066) / len(primes_1066)
primes_1066_average
# This result makes sense intuitively because prime numbers are known to become less
# frequent for larger number intervals. These examples demonstrate how Python treats
# functions as first-class objects so that functions may be passed as parameters to
# other functions. This is a key property of functional programming and demonstrates
# the power of Python.
#
# In the next code snippet, we can use list comprehensions (a ‘Pythonic’ form of the
# map-filter-reduce template) to ‘mine’ the factors of 1066 to find those factors that
# end in the digit ‘3’.
primes_1066_ends3 = [x for x in primes_between(1, 1066)
if str(x).endswith('3')]
print('{}'.format(primes_1066_ends3))
# This code tells Python to first convert each prime between 1 and 1066 to a string and
# then to return those numbers whose string representation end with the number ‘3’. It
# uses the built-in str() and endswith() methods to test each prime for inclusion in the list.
#
# And because we really want to know what fraction of the 180 primes of 1066 end in a
# ‘3’, we can calculate ...
len(primes_1066_ends3) / len(primes_1066)
# These examples demonstrate how Python is a modern, multi-paradigmatic language. More
# simply, it continually integrates the best features of other leading languages, including
# functional programming constructs. Consider how many lines of code you would need to
# implement the list comprehension above in C and you get an appreciation of the power
# of productivity-layer languages. Higher levels of programming abstraction really do
# result in higher programmer productivity!
# ## Numpy Data Movement
#
# Code in the cells below show a very simple data movement code snippet that can be used
# to share data with programmable logic. We leverage the Python numpy package to
# manipulate the buffer on the ARM processors and can then send a buffer pointer to
# programmable logic for sharing data.
#
# We do not assume what programmable logic design is loaded, so here we only allocate
# the needed memory space and show that it can manipulated as a numpy array and contains
# a buffer pointer attribute. That pointer can then can be passed to programmable
# logic hardware.
# +
import numpy as np
import pynq
def get_pynq_buffer(shape, dtype):
""" Simple function to call PYNQ's memory allocator with numpy attributes
"""
return pynq.allocate(shape, dtype)
# -
# With the simple wrapper above, we can get access to memory that can be shared by both
# numpy methods and programmable logic.
buffer = get_pynq_buffer(shape=(4,4), dtype=np.uint32)
buffer
# To double-check we show that the buffer is indeed a numpy array.
isinstance(buffer,np.ndarray)
# To send the buffer pointer to programmable logic, we use its physical address which
# is what programmable logic would need to communicate using this shared buffer.
pl_buffer_address = hex(buffer.physical_address)
pl_buffer_address
# In this short example, we showed a simple allocation of a numpy array that is now ready
# to be shared with programmable logic devices. With numpy arrays that are accessible to programmable logic, we can quickly manipulate and move data across software and hardware.
# ## Asyncio Integration
#
# PYNQ also leverages the Python asyncio module for communicating with programmable logic
# devices through events (namely interrupts).
#
# A Python program running on PYNQ can use the asyncio library to manage multiple IO-bound
# tasks asynchronously, thereby avoiding any blocking caused by waiting for responses from
# slower IO subsystems. Instead, the program can continue to execute other tasks that are
# ready to run. When the previously-busy tasks are ready to resume, they will be executed
# in turn, and the cycle is repeated.
#
# Again, since we won't assume what interrupt enabled devices are loaded on programmable
# logic, we will show an example here a software-only asyncio example that uses asyncio's
# sleep method.
# +
import asyncio
import random
import time
# Coroutine
async def wake_up(delay):
'''A function that will yield to asyncio.sleep() for a few seconds
and then resume, having preserved its state while suspended
'''
start_time = time.time()
print(f'The time is: {time.strftime("%I:%M:%S")}')
print(f"Suspending coroutine 'wake_up' at 'await` statement\n")
await asyncio.sleep(delay)
print(f"Resuming coroutine 'wake_up' from 'await` statement")
end_time = time.time()
sleep_time = end_time - start_time
print(f"'wake-up' was suspended for precisely: {sleep_time} seconds")
# -
# With the wake_up function defined, we then can add a new task to the event loop.
# +
delay = random.randint(1,5)
my_event_loop = asyncio.get_event_loop()
try:
print("Creating task for coroutine 'wake_up'\n")
wake_up_task = my_event_loop.create_task(wake_up(delay))
my_event_loop.run_until_complete(wake_up_task)
except RuntimeError as err:
print (f'{err}' +
' - restart the Jupyter kernel to re-run the event loop')
finally:
my_event_loop.close()
# -
# All the above examples show standard Python 3.6 running on the PYNQ platform. This entire notebook can be run on the PYNQ board - see the getting_started folder on the Jupyter landing page to rerun this notebook.
| jupyter_notebooks/getting_started/2_python_environment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Web-based Tools for Teaching and Research: Jupyter Notebooks and GitHub
# A workshop of the Academy of Data Sciences at the College of Science
# ## Jupyter Notebooks & Python
#
# <NAME>
#
# ---
#
# ## Lesson 5: Crash Course on Python Statements: Loops, Decisions and Functions
#
# This lesson is contains selected material of the carpentry tutorial "Programming with Python":
# https://software-carpentry.org/lessons/
#
# ---
# ## Importance of Indent for Statements
#
# Unlike many other languages, there is no command to signify the end of statement bodies like decisions, loops or function ; what is **indented** after the statement (loop, decision or function) belongs to it:
#
# ```python
# this_statement(parameter): # decision/loop/function
# indented content 1
# indented content 2
# indented content 3
#
# next_statement
# ```
# ---
#
# ## Decisions in Python
#
# How can we use Python to automatically recognize the different features we saw, and take a different action for each?
#
# We can ask Python to take different actions, depending on a condition, with an `if` statement:
num = 37
if num > 100:
print('greater 100')
else:
print('not greater 100')
print('done')
# #### Comparing in Python
# Along with the > and == operators we have already used for comparing values in our conditionals, there are a few more options to know about:
#
# >: greater than
# <: less than
# ==: equal to
# !=: does not equal
# >=: greater than or equal to
# <=: less than or equal to
# ---
# ## Loops in Python
#
# We have a dozen data sets right now, though, and more on the way. We want to create plots for all of our data sets with a single statement. To do that, we’ll have to teach the computer how to repeat things.
#
# In Python we can use for- and while-loops.
#
# ### for-loop
#
# A for loop is used for iterating over a sequence (that is either a list, a tuple, a dictionary, a set, or a string).
#
# This is less like the for keyword in other programming languages, and works more like an iterator method as found in other object-orientated programming languages.
#
# With the for loop we can execute a set of statements, once for each item in a list, tuple, set etc.
# prints out every second number between 2 and 8
for num in range(2,9,2):
print(num)
# ---
# ### while-loop
#
# With the while loop we can execute a set of statements as long as a condition is true.
# +
# Prints out 0,1,2,3,4
count = 0
while count < 5:
print(count)
count += 1 # This is the same as count = count + 1
# -
# ---
#
# ## Functions in Python
#
# We’d like a way to package our code so that it is easier to reuse, and Python provides for this by letting us define things called ‘functions’ — a shorthand way of re-executing longer pieces of code. Let’s start by defining a function `fahr_to_celsius` that converts temperatures from Fahrenheit to Celsius.
#
# See the anatomy of such a function displayed below:
#
# <img src="img/python-function.svg" alt="Function" title="Function" width="500" />
#
#
# The function definition opens with the keyword def followed by the name of the function (`fahr_to_celsius`) and a parenthesized list of parameter names (`temp`). The body of the function — the statements that are executed when it runs — is indented below the definition line. The body concludes with a return keyword followed by the `return` value.
#
# When we call the function, the values we pass to it are assigned to those variables so that we can use them inside the function. Inside the function, we use a return statement to send a result back to whoever asked for it.
#
# Now, let's define the function in a code cell and use it:
def fahr_to_celsius(temp):
celcius = ((temp - 32) * (5/9))
return celcius
print('freezing point of water:', fahr_to_celsius(32), 'C')
print('boiling point of water:', fahr_to_celsius(212), 'C')
print('body temperature:', fahr_to_celsius(98.6), 'C')
# ---
# ## Exercise
#
# Write a function of name `greater100`, which allows you to perform the following decision for any number.
#
num = 37
if num > 100:
print('greater')
else:
print('not greater')
| L5_CrashCourseStatements.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: newenv
# language: python
# name: newenv
# ---
# +
# Fun some setup for the project
# Silence annoying pytorch deprecated warnings
import warnings
warnings.filterwarnings("ignore")
from train import *
from create_graphs import *
from evaluate import *
# %matplotlib inline
# for auto-reloading extenrnal modules
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
print ('CUDA Available:', torch.cuda.is_available())
# +
### Generate the model datasets (i.e. the dataset used to train
# the model - normal data)
args_ladder_t, train_ladder_t, val_ladder_t, test_ladder_t = get_graph_data("ladder_tree_10", isModelDataset=True)
# Save the max_previous node to allow for model
# compatability on future datasets
max_prev_node = args_ladder_t.max_prev_node
# Create dataset and dataloader object for the out of distribution ladder dataset
# Note that instead of passing in args.max_prev_node, we pass in the saved max_prev_node
# saved specifically for the enzyme dataset - hack to allow for GraphRNN to work on any dataset
train_dataset = Graph_sequence_sampler_pytorch_rand(train_ladder_t,max_prev_node=max_prev_node,max_num_node=args_ladder_t.max_num_node)
train_loader = torch.utils.data.DataLoader(train_dataset, batch_size=1, num_workers=args_ladder_t.num_workers)
test_dataset = Graph_sequence_sampler_pytorch_rand(test_ladder_t, max_prev_node=max_prev_node,max_num_node=args_ladder_t.max_num_node)
test_loader = torch.utils.data.DataLoader(test_dataset, batch_size=1, num_workers=args_ladder_t.num_workers)
# Model initialization
# Using GraphRNN
rnn = GRU_plain(input_size=args_ladder_t.max_prev_node, embedding_size=args_ladder_t.embedding_size_rnn,
hidden_size=args_ladder_t.hidden_size_rnn, num_layers=args_ladder_t.num_layers, has_input=True,
has_output=True, output_size=args_ladder_t.hidden_size_rnn_output).to(device)
output = GRU_plain(input_size=1, embedding_size=args_ladder_t.embedding_size_rnn_output,
hidden_size=args_ladder_t.hidden_size_rnn_output, num_layers=args_ladder_t.num_layers, has_input=True,
has_output=True, output_size=1).to(device)
# Let's see how the nlls of the ladder graphs compare to
# the trained on enzymes.
# Note we pass in the args_enzy because the model we are
# using are trained in the enzyme dataset
train_nlls, train_avg_nlls = calc_nll(args_ladder_t, train_loader, rnn, output, max_iter=10, load_epoch=3000, log=1)
train_avg_nlls_iter = np.array(train_avg_nlls)
train_avg_nlls_iter = train_avg_nlls_iter.reshape((10, len(train_loader))) #todo ordering
train_avg_nlls_iter = np.mean(train_avg_nlls_iter, axis=0)
# Analysis of the test data set nlls.
# We really gotta train over more data!
test_nlls, test_avg_nlls = calc_nll(args_ladder_t, test_loader, rnn, output, load_epoch=3000, max_iter=10, log=1)
test_avg_nlls_iter = np.array(test_avg_nlls)
test_avg_nlls_iter = test_avg_nlls_iter.reshape((10, len(test_loader)))
test_avg_nlls_iter = np.mean(test_avg_nlls_iter, axis=0)
# -
def plot_nlls(nlls, iterations, num_graphs):
# Calculate also the nlls averaged over the iterations for each batch
nlls_iter = np.mean(np.array(nlls).reshape(iterations, num_graphs), axis=0)
fig, ax = plt.subplots(1, 2, figsize=(12, 5))
# Plot the two distributions side by side
sns.distplot(nlls, ax=ax[0], kde=True)
ax[0].set_title("NLL Distribution across many permutations")
ax[0].set_xlabel("Negative Log Likelihood")
sns.distplot(nlls_iter, ax=ax[1], kde=True)
ax[1].set_title("NLL Averaged Across Permutations")
ax[1].set_xlabel("Negative Log Likelihood")
return ax
def compare_dist(nlls, labels=['Nomarl', 'Anomalous'], title="Normal vs. Anomalous NLL Distributions"):
fig, ax = plt.subplots()
for i in range(len(nlls)):
sns.distplot(nlls[i], ax=ax, kde=True, label=labels[i])
ax.legend()
ax.set_xlabel("Negative Log Likelihood")
ax.set_title(title)
return fig, ax
fig, ax_compare = compare_dist([train_avg_nlls_iter, test_avg_nlls_iter],
['Train (normal)', 'Test (normal)'],
'Train vs. Test Distributions')
fig.savefig("NLL_Figures/Train_Test_Type_3")
# +
# Now let's try using a different dataset on the same GraphRNN
# trained on the enzyemes small dataset. By plotting the distribution
# over the nlls of this test set from a different family, hopefully
# we can see the success of the generative model
args_rand, graphs_rand = get_graph_data("random_6_60", isModelDataset=False)
# Create dataset and dataloader object for the out of distribution ladder dataset
# Note that instead of passing in args.max_prev_node, we pass in the saved max_prev_node
# saved specifically for the enzyme dataset - hack to allow for GraphRNN to work on any dataset
rand_dataset = Graph_sequence_sampler_pytorch_rand(graphs_rand,max_prev_node=max_prev_node,max_num_node=args_rand.max_num_node)
rand_loader = torch.utils.data.DataLoader(rand_dataset, batch_size=1, num_workers=args_rand.num_workers)
# Let's see how the nlls of the ladder graphs compare to
# the trained on enzymes.
# Note we pass in the args_enzy because the model we are
# using are trained in the enzyme dataset
rand_nlls, rand_avg_nlls = calc_nll(args_ladder_t, rand_loader, rnn, output, max_iter=10, train_dataset="ladder_tree_10", load_epoch=300, log=1)
rand_avg_nlls_iter = np.array(rand_avg_nlls)
rand_avg_nlls_iter = rand_avg_nlls_iter.reshape((10, len(rand_loader)))
rand_avg_nlls_iter = np.mean(rand_avg_nlls_iter, axis=0)
# -
fig, ax_compare = compare_dist([train_avg_nlls_iter, rand_avg_nlls_iter],
['Type 3', '6-Random'])
fig.savefig("Type_3-VS-6_Random")
# +
# Starting with enzyme with label 2
args_ladder_e_full_circ, graphs_ladder_e_full_circ = get_graph_data("ladder_extra_full_circular", isModelDataset=False)
# Create dataset and dataloader object for the out of distribution ladder dataset
# Note that instead of passing in args.max_prev_node, we pass in the saved max_prev_node
# saved specifically for the enzyme dataset - hack to allow for GraphRNN to work on any dataset
ladder_e_full_circ_dataset = Graph_sequence_sampler_pytorch_rand(graphs_ladder_e_full_circ,max_prev_node=max_prev_node,max_num_node=args_ladder_e_full_circ.max_num_node)
ladder_e_full_circ_loader = torch.utils.data.DataLoader(ladder_e_full_circ_dataset, batch_size=1, num_workers=args_ladder_e_full_circ.num_workers)
# Let's see how the nlls of the ladder graphs compare to
# the trained on enzymes.
# Note we pass in the args_enzy because the model we are
# using are trained in the enzyme dataset
ladder_e_full_circ_nlls, ladder_e_full_circ_avg_nlls = calc_nll(args_ladder_t,
ladder_e_full_circ_loader,
rnn, output, max_iter=10,
load_epoch=3000, train_dataset="ladder_tree_10", log=1)
ladder_e_full_circ_avg_nlls_iter = np.array(ladder_e_full_circ_avg_nlls)
ladder_e_full_circ_avg_nlls_iter = ladder_e_full_circ_avg_nlls_iter.reshape((10, len(ladder_e_full_circ_loader)))
ladder_e_full_circ_avg_nlls_iter = np.mean(ladder_e_full_circ_avg_nlls_iter, axis=0)
# +
# Let us look at the differences between the graphs for train and test at a very basic level
metrics = compare_graph_list(train_ladder_t, graphs_ladder_e_full_circ)
print (metrics['avg_degree'])
print (metrics['avg_clust'])
# Plot the connectivity distributions
# Plot the clust distributions
fig1, ax1 = plt.subplots()
labels = ['ladder_tree', 'ladder_extra_Full_circular']
connect = metrics['connect_dist']
for i in range(len(connect)):
sns.distplot(connect[i], ax=ax1, kde=False, label=labels[i])
ax1.legend()
ax1.set_title("Comparision of avg degree connectivy")
fig1.savefig("Degree_Conn-Ladder_Tree-Vs-Ladder_Extra_Full_Circular")
# Plot the clust distributions
fig2, ax2 = plt.subplots()
labels = ['ladder_tree', 'ladder_extra_full_circular']
clust = metrics['clust_dist']
for i in range(len(clust)):
sns.distplot(clust[i], ax=ax2, kde=False, label=labels[i])
ax2.legend()
ax2.set_title("Comparision of avg clusering coefficients")
fig1.savefig("Clustering-Ladder_Tree-Vs-Ladder_Extra_Full_Circular")
# -
fig, ax_compare = compare_dist([train_avg_nlls_iter,
ladder_e_full_circ_avg_nlls_iter],
['Type 3 (normal)', 'Type 2c'],
'Type 3 vs. Type 2c')
fig.savefig('NLL_Figures/Type_3-VS-Type_2c')
# +
# Starting with enzyme with label 2
args_ladder_e_circ, graphs_ladder_e_circ = get_graph_data("ladder_extra_circular", isModelDataset=False)
# Create dataset and dataloader object for the out of distribution ladder dataset
# Note that instead of passing in args.max_prev_node, we pass in the saved max_prev_node
# saved specifically for the enzyme dataset - hack to allow for GraphRNN to work on any dataset
ladder_e_circ_dataset = Graph_sequence_sampler_pytorch_rand(graphs_ladder_e_circ,max_prev_node=max_prev_node,max_num_node=args_ladder_e_circ.max_num_node)
ladder_e_circ_loader = torch.utils.data.DataLoader(ladder_e_circ_dataset, batch_size=1, num_workers=args_ladder_e_circ.num_workers)
# Let's see how the nlls of the ladder graphs compare to
# the trained on enzymes.
# Note we pass in the args_enzy because the model we are
# using are trained in the enzyme dataset
ladder_e_circ_nlls, ladder_e_circ_avg_nlls = calc_nll(args_ladder_t,
ladder_e_circ_loader,
rnn, output, max_iter=10,
load_epoch=3000, train_dataset="ladder_tree_10", log=1)
ladder_e_circ_avg_nlls_iter = np.array(ladder_e_circ_avg_nlls)
ladder_e_circ_avg_nlls_iter = ladder_e_circ_avg_nlls_iter.reshape((10, len(ladder_e_circ_loader)))
ladder_e_circ_avg_nlls_iter = np.mean(ladder_e_circ_avg_nlls_iter, axis=0)
# -
fig, ax_compare = compare_dist([train_avg_nlls_iter,
ladder_e_circ_avg_nlls_iter],
['Type 3 (normal)', 'Type 2b'],
'Type 3 vs. Type 2b')
fig.savefig('NLL_Figures/Type_3-VS-Type_2b')
# # Now lets go the other way. Where the train distribution is Ladder Extra Full Circular Graphs which have more rigid structure.
# +
# Generate the model datasets (i.e. the dataset used to train
# the model - normal data)
args_ladder_e_full_circ, train_ladder_e_full_circ, _, test_ladder_e_full_circ = get_graph_data("ladder_extra_full_circular", isModelDataset=True)
# Save the max_previous node to allow for model
# compatability on future datasets
max_prev_node2 = args_ladder_e_full_circ.max_prev_node
# Create dataset and dataloader object for the out of distribution ladder dataset
# Note that instead of passing in args.max_prev_node, we pass in the saved max_prev_node
# saved specifically for the enzyme dataset - hack to allow for GraphRNN to work on any dataset
train_dataset2 = Graph_sequence_sampler_pytorch_rand(train_ladder_e_full_circ,max_prev_node=max_prev_node2,max_num_node=args_ladder_e_full_circ.max_num_node)
train_loader2 = torch.utils.data.DataLoader(train_dataset2, batch_size=1, num_workers=args_ladder_e_full_circ.num_workers)
test_dataset2 = Graph_sequence_sampler_pytorch_rand(test_ladder_e_full_circ, max_prev_node=max_prev_node2,max_num_node=args_ladder_e_full_circ.max_num_node)
test_loader2 = torch.utils.data.DataLoader(test_dataset2, batch_size=1, num_workers=args_ladder_e_full_circ.num_workers)
# Model initialization
# Using GraphRNN
rnn2 = GRU_plain(input_size=max_prev_node2, embedding_size=args_ladder_e_full_circ.embedding_size_rnn,
hidden_size=args_ladder_e_full_circ.hidden_size_rnn, num_layers=args_ladder_e_full_circ.num_layers, has_input=True,
has_output=True, output_size=args_ladder_e_full_circ.hidden_size_rnn_output).to(device)
output2 = GRU_plain(input_size=1, embedding_size=args_ladder_e_full_circ.embedding_size_rnn_output,
hidden_size=args_ladder_e_full_circ.hidden_size_rnn_output, num_layers=args_ladder_e_full_circ.num_layers, has_input=True,
has_output=True, output_size=1).to(device)
# Let's see how the nlls of the ladder graphs compare to
# the trained on enzymes.
# Note we pass in the args_enzy because the model we are
# using are trained in the enzyme dataset
train_nlls2, train_avg_nlls2 = calc_nll(args_ladder_e_full_circ, train_loader2, rnn2, output2, max_iter=10, load_epoch=3000, log=1)
train_avg_nlls_iter2 = np.array(train_avg_nlls2)
train_avg_nlls_iter2 = train_avg_nlls_iter2.reshape((10, len(train_loader2)))
train_avg_nlls_iter2 = np.mean(train_avg_nlls_iter2, axis=0)
# Analysis of the test data set nlls.
# We really gotta train over more data!
test_nlls2, test_avg_nlls2 = calc_nll(args_ladder_e_full_circ, test_loader2, rnn2, output2, load_epoch=3000, max_iter=10, log=1)
test_avg_nlls_iter2 = np.array(test_avg_nlls2)
test_avg_nlls_iter2 = test_avg_nlls_iter2.reshape((10, len(test_loader2)))
test_avg_nlls_iter2 = np.mean(test_avg_nlls_iter2, axis=0)
# -
fig, ax_compare = compare_dist([train_avg_nlls_iter2, test_avg_nlls_iter2],
['Train (normal)', 'Test (normal)'],
'Train vs. Test Distributions')
fig.savefig("NLL_Figures/Train_Test_Type_2c")
# +
# Starting with enzyme with label 2
args_ladder_t, graphs_ladder_t = get_graph_data("ladder_tree_10", isModelDataset=False)
# Create dataset and dataloader object for the out of distribution ladder dataset
# Note that instead of passing in args.max_prev_node, we pass in the saved max_prev_node
# saved specifically for the enzyme dataset - hack to allow for GraphRNN to work on any dataset
ladder_t_dataset = Graph_sequence_sampler_pytorch_rand(graphs_ladder_t,max_prev_node=max_prev_node2,max_num_node=args_ladder_t.max_num_node)
ladder_t_loader = torch.utils.data.DataLoader(ladder_t_dataset, batch_size=1, num_workers=args_ladder_t.num_workers)
# Let's see how the nlls of the ladder graphs compare to
# the trained on enzymes.
# Note we pass in the args_enzy because the model we are
# using are trained in the enzyme dataset
ladder_t_nlls, ladder_t_avg_nlls = calc_nll(args_ladder_e_full_circ, ladder_t_loader, rnn2, output2, max_iter=10, load_epoch=3000, train_dataset="ladder_extra_full_circular", log=1)
ladder_t_avg_nlls_iter = np.array(ladder_t_avg_nlls)
ladder_t_avg_nlls_iter = ladder_t_avg_nlls_iter.reshape((10, len(ladder_t_loader)))
ladder_t_avg_nlls_iter = np.mean(ladder_t_avg_nlls_iter, axis=0)
# -
fig, ax_compare = compare_dist([train_avg_nlls_iter2,
ladder_t_avg_nlls_iter],
['Type_2c (normal)', 'Type_3'],
'Type 2c vs. Type 3')
fig.savefig('NLL_Figures/Type_2c-VS-Type_3')
# # We should also now give a score to how anomalous each is by looking at the ratio of nodes in ladder tree that correctly connect to the node bellow them
def anomalous_score(graphs, height):
"""
Given Ladder Tree graphs, calculate their anomalous
score in comparrison to ladder extre full ciruclar graphs
by seeing how often the constraint is broken where a node
does not connect to the node directly bellow it in the lader
"""
anom_ratios = []
for G in graphs:
anom_nodes = 0
for node in G.nodes():
# Check if the node connects to
# the node directly bellow it in the ladder
row = node[0]
indx = node[1]
if not G.has_edge(node, ((row + 1) % height, indx)):
anom_nodes += 1
anom_ratios.append(anom_nodes / len(G.nodes()))
return anom_ratios
def anomalous_score2(graphs, height):
"""
Given Ladder Tree graphs, calculate their anomalous
score in comparrison to ladder extre full ciruclar graphs
by seeing how often the constraint is broken where a node
does not connect to the node directly bellow and above it in the lader
"""
anom_ratios = []
for G in graphs:
anom_nodes = 0
for node in G.nodes():
# Check if the node connects to
# the node directly bellow it in the ladder
row = node[0]
indx = node[1]
if not G.has_edge(node, ((row + 1) % height, indx)) \
and not G.has_edge(node, ((row - 1) % height, indx)):
anom_nodes += 1
anom_ratios.append(anom_nodes / len(G.nodes()))
return anom_ratios
# +
anom_ratios = anomalous_score(graphs_ladder_t, 10)
fig_scat, ax_scat = plt.subplots()
scatter = ax_scat.scatter(anom_ratios, ladder_t_avg_nlls_iter)
ax_scat.set_title("Anomalous Ratio vs. NLL in Type 3 Graphs")
ax_scat.set_xlabel("Anomalous Ratio")
ax_scat.set_ylabel("NLL")
fig_scat.savefig('NLL_Figures/Scatter_Anomalous_Ratio_Type_3')
# +
anom_ratios = anomalous_score2(graphs_ladder_t, 10)
fig_scat, ax_scat = plt.subplots()
scatter = plt.scatter(anom_ratios, ladder_t_avg_nlls_iter)
ax_scat.set_title("Strict Anomalous Ratio vs. NLL in Ladder Tree Graphs")
ax_scat.set_xlabel("Anomalous Ratio")
ax_scat.set_ylabel("NLL")
fig_scat.savefig('NLL_Figures/Scatter_Strict_Anomalous_Ratio_Ladder_Tree')
# -
def draw_ladder(graph, height, highlight_grid=True):
fig, ax = plt.subplots()
# Plot the edge
for edge in graph.edges():
node1 = edge[0]
node2 = edge[1]
# Flip the coordinates so the ladder
# Goes downward
y = [height - node1[0] - 1, height - node2[0] - 1]
x = [node1[1], node2[1]]
# Don't include circular connections from the final layer
if highlight_grid:
if (y[0] != height -1 or y[1] != 0) \
and (y[0] != 0 or y[1] != height -1):
# See if it defines the grid structure
if (x[0] == x[1]) or y[0] == y[1]:
ax.plot(x, y, 'r-')
else:
ax.plot(x, y, 'g-')
else:
ax.plot(x, y, 'g-')
# Plot the nodes
for node in graph.nodes():
ax.plot(node[1], height - node[0] - 1, 'bo')
return fig, ax
# +
# Plt the graphs with heighest and lowest nll
sort_indx = np.argsort(ladder_t_avg_nlls_iter)
graph_sorted_nll = [graphs_ladder_t[i] for i in sort_indx]
fig_low, ax_low = draw_ladder(graph_sorted_nll[0], 10)
ax_low.set_title("Lowest NLL Graph")
fig_high, ax_high = draw_ladder(graph_sorted_nll[len(graph_sorted_nll) - 1], 10)
ax_high.set_title("Highest NLL Graph")
# +
# Plt the graphs with heighest and lowest nll
anom_ratios = np.array(anomalous_score(graphs_ladder_t, 10))
sort_indx = np.argsort(anom_ratios)
nlls_sorted_ratio = [ladder_t_avg_nlls_iter[i] for i in sort_indx]
graph_sorted_nll = [graphs_ladder_t[i] for i in sort_indx]
fig_low, ax_low = draw_ladder(graph_sorted_nll[0], 10)
ax_low.set_title("Lowest Ratio Graph - NLL = {}".format(nlls_sorted_ratio[0]))
fig_high, ax_high = draw_ladder(graph_sorted_nll[len(graph_sorted_nll) - 1], 10)
ax_high.set_title("Highest NLL Graph - NLL = {}".format(nlls_sorted_ratio[len(graph_sorted_nll) - 1]))
# -
def edge_distance(graphs, height):
"""
Given Ladder Tree graphs, calculate the average
seperation between the two edges connecting each
node to the next layer.
"""
avg_seperations = []
for G in graphs:
distances = 0
for node in G.nodes():
# Get the edges for that node
edges = G.edges(node)
# Find the edges that connect to the next row
# and get there column coordinates
connecting_points = []
for edge in edges:
if edge[0][0] == (node[0] + 1) % height:
connecting_points.append(edge[0][1])
elif edge[1][0] == (node[0] + 1) % height:
connecting_points.append(edge[1][1])
distances += abs(connecting_points[0] - connecting_points[1])
avg_seperations.append(distances / len(G.nodes()))
return avg_seperations
# +
avg_seperations = edge_distance(graphs_ladder_t, 10)
fig_scat, ax_scat = plt.subplots()
scatter = ax_scat.scatter(avg_seperations, ladder_t_avg_nlls_iter)
ax_scat.set_title("Avg Seperation vs. NLL in Ladder Tree Graphs")
ax_scat.set_xlabel("Avg Seperation")
ax_scat.set_ylabel("NLL")
#fig_scat.savefig('NLL_Figures/Scatter_Anomalous_Ratio_Ladder_Tree')
# -
def avg_edge_distance(graphs, height):
"""
Given Ladder Tree graphs, calculate the average
seperation between the two edges connecting each
node to the next layer.
"""
avg_seperations = []
for G in graphs:
distances = 0
for node in G.nodes():
# Get the edges for that node
edges = G.edges(node)
# Find the edges that connect to the next row
# and get there column coordinates
row = node[0]
col = node[1]
for edge in edges:
if edge[0][0] == (row + 1) % height:
distances += abs(edge[0][1] - col)
elif edge[1][0] == (row + 1) % height:
distances += abs(edge[1][1] - col)
avg_seperations.append(distances / len(G.nodes()))
return avg_seperations
# +
avg_seperations = avg_edge_distance(graphs_ladder_t, 10)
fig_scat, ax_scat = plt.subplots()
scatter = ax_scat.scatter(avg_seperations, ladder_t_avg_nlls_iter)
ax_scat.set_title("Avg Seperation vs. NLL in Ladder Tree Graphs")
ax_scat.set_xlabel("Avg Seperation")
ax_scat.set_ylabel("NLL")
#fig_scat.savefig('NLL_Figures/Scatter_Anomalous_Ratio_Ladder_Tree')
# -
def node_degree_variance(graphs):
variances = []
for G in graphs:
degrees = [degree for _, degree in G.degree().items()]
var = np.var(degrees)
variances.append(var)
return variances
# solve for a and b
def best_fit(X, Y):
xbar = sum(X)/len(X)
ybar = sum(Y)/len(Y)
n = len(X) # or len(Y)
numer = sum([xi*yi for xi,yi in zip(X, Y)]) - n * xbar * ybar
denum = sum([xi**2 for xi in X]) - n * xbar**2
b = numer / denum
a = ybar - b * xbar
print('best fit line:\ny = {:.2f} + {:.2f}x'.format(a, b))
return a, b
# +
variances = node_degree_variance(graphs_ladder_t)
fig_scat, ax_scat = plt.subplots()
scatter = ax_scat.scatter(variances, ladder_t_avg_nlls_iter)
# Add best fit line
a, b = best_fit(variances, ladder_t_avg_nlls_iter)
yfit = [a + b * xi for xi in variances]
plt.plot(variances, yfit, 'r', label='r = 0.635')
ax_scat.set_title("Variance vs. NLL in Type 3 Graphs")
ax_scat.set_xlabel("Variance")
ax_scat.set_ylabel("NLL")
ax_scat.legend()
fig_scat.savefig('NLL_Figures/Scatter_Variance_Type_3')
print ("r = {}".format(np.corrcoef(variances, ladder_t_avg_nlls_iter)[0][1]))
# +
# Plt the graphs with heighest and lowest nll
variances = np.array(node_degree_variance(graphs_ladder_t))
sort_indx = np.argsort(variances)
nlls_sorted_ratio = [ladder_t_avg_nlls_iter[i] for i in sort_indx]
graph_sorted_variance = [graphs_ladder_t[i] for i in sort_indx]
fig_low, ax_low = draw_ladder(graph_sorted_nll[0], 10)
ax_low.set_title("Lowest Var Graph - NLL = {}".format(nlls_sorted_ratio[0]))
fig_high, ax_high = draw_ladder(graph_sorted_nll[len(graph_sorted_nll) - 1], 10)
ax_high.set_title("Highest Var Graph - NLL = {}".format(nlls_sorted_ratio[len(graph_sorted_nll) - 1]))
# -
def number_squares(graphs):
"""
Calculate the number of closed squares in a list of graphs
with the formula:
tr(A^4) - sum(degree ^ 2)
"""
squares = []
for G in graphs:
adj = np.asarray(nx.to_numpy_matrix(G))
trace = np.trace(np.linalg.matrix_power(adj, 4))
degree_sum = sum([degree ** 2 for _, degree in G.degree().items()])
squares.append(trace - degree_sum)
return squares
# +
# Plt the graphs with heighest and lowest nll
squares = np.array(number_squares(graphs_ladder_t))
sort_indx = np.argsort(squares)
nlls_sorted_ratio = [ladder_t_avg_nlls_iter[i] for i in sort_indx]
graph_sorted_variance = [graphs_ladder_t[i] for i in sort_indx]
fig_low, ax_low = draw_ladder(graph_sorted_nll[0], 10)
ax_low.set_title("Lowest squares Graph - NLL = {}".format(nlls_sorted_ratio[0]))
fig_high, ax_high = draw_ladder(graph_sorted_nll[len(graph_sorted_nll) - 1], 10)
ax_high.set_title("Highest squares Graph - NLL = {}".format(nlls_sorted_ratio[len(graph_sorted_nll) - 1]))
# +
squares = number_squares(graphs_ladder_t)
fig_scat, ax_scat = plt.subplots()
scatter = ax_scat.scatter(squares, ladder_t_avg_nlls_iter)
# Add best fit line
a, b = best_fit(squares, ladder_t_avg_nlls_iter)
yfit = [a + b * xi for xi in squares]
plt.plot(squares, yfit, 'r', label='r = 0.635')
ax_scat.set_title("Number Squares vs. NLL in Type 3 Graphs")
ax_scat.set_xlabel("Squares")
ax_scat.set_ylabel("NLL")
ax_scat.legend()
#fig_scat.savefig('NLL_Figures/Scatter_Variance_Ladder_Tree')
print ("r = {}".format(np.corrcoef(squares, ladder_t_avg_nlls_iter)[0][1]))
# -
def layers_connectedness(graphs, width, height):
mapping_scores = []
for G in graphs:
# For each row we want to basically give a score
# for how well it has a one to one mapping
# to the next row
mapping = 0
for row in range(height + 1):
# Now we want to try the different permutations
# where we rotate the row by permute
max_row = 0
for permute in range(width):
row_score = 0
for col in range(width):
# Check now if the current (row, col)
# connects to the (row + 1, col + permute)
# To make the one to one mapping.
if G.has_edge((row % height, col), ((row + 1)%height, (col + permute) % width)):
row_score += 1
max_row = max(row_score, max_row)
#print (max_row)
mapping += max_row
#break
mapping_scores.append(mapping)
return mapping_scores
# +
squares = layers_connectedness(graphs_ladder_t, width=6, height=10)
fig_scat, ax_scat = plt.subplots()
scatter = ax_scat.scatter(squares, ladder_t_avg_nlls_iter)
ax_scat.set_title("One-to-One Mapping Score vs. NLL in Ladder Tree Graphs")
ax_scat.set_xlabel("One-to-One Mapping Score")
ax_scat.set_ylabel("NLL")
#fig_scat.savefig('NLL_Figures/Scatter_Variance_Ladder_Tree')
# -
def node_degree_score(graphs, threshold=4):
scores = []
for G in graphs:
# Count the number of nodes with in degree bellow/=
# to the provided threshold
num_anom_nodes = sum([1 for _, degree in G.degree().items() if degree <= threshold])
# Maybe we should get percent this?
# does not matter for now!
scores.append(num_anom_nodes/len(G.nodes()))
return scores
# +
node_d_scores = node_degree_score(graphs_ladder_t, threshold=4)
fig_scat, ax_scat = plt.subplots()
scatter = ax_scat.scatter(node_d_scores, ladder_t_avg_nlls_iter)
# Add best fit line
a, b = best_fit(node_d_scores, ladder_t_avg_nlls_iter)
yfit = [a + b * xi for xi in node_d_scores]
plt.plot(node_d_scores, yfit, 'r', label='r = 0.608')
ax_scat.set_title("One-to-One Mapping Score vs. NLL in Type 3 Graphs")
ax_scat.set_xlabel("One-to-One Mapping Score")
ax_scat.set_ylabel("NLL")
ax_scat.legend()
fig_scat.savefig('NLL_Figures/Scatter_One-to-One_Type_3')
print ("r = {}".format(np.corrcoef(node_d_scores, ladder_t_avg_nlls_iter)[0][1]))
# -
| Fine_Grained_Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# Copyright 2021 NVIDIA Corporation. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
# -
# # 1.Overview
#
# In this notebook, we want to provide an tutorial how to train a wdl model using HugeCTR High-level python API with original Criteo dataset as training data. And get prediction result based on different type of local database
#
# 1. Overview
# 2. Dataset Preprocessing
# 3. WDL Model Training
# 4. Save the Model Files
# 5. Create prediction scripts
# 6. Prediction
# # 2. Dataset Preprocessing
# ## 2.1 Generate training and validation data folders
# +
# define some data folder to store the original and preprocessed data
# Standard Libraries
import os
from time import time
import re
import shutil
import glob
import warnings
BASE_DIR = "/wdl_train"
train_path = os.path.join(BASE_DIR, "train")
val_path = os.path.join(BASE_DIR, "val")
CUDA_VISIBLE_DEVICES = os.environ.get("CUDA_VISIBLE_DEVICES", "0")
n_workers = len(CUDA_VISIBLE_DEVICES.split(","))
frac_size = 0.15
allow_multi_gpu = False
use_rmm_pool = False
max_day = None # (Optional) -- Limit the dataset to day 0-max_day for debugging
if os.path.isdir(train_path):
shutil.rmtree(train_path)
os.makedirs(train_path)
if os.path.isdir(val_path):
shutil.rmtree(val_path)
os.makedirs(val_path)
# -
# ls -l $train_path
# ## 2.2 Download the Original Criteo Dataset
# !apt-get install wget
# !wget -P $train_path http://azuremlsampleexperiments.blob.core.windows.net/criteo/day_0.gz
#Download the split data set to training and validation
# !gzip -d -c $train_path/day_0.gz > day_0
# !head -n 45840617 day_0 > $train_path/train.txt
# !tail -n 2000000 day_0 > $val_path/test.txt
# ## 2.3 Preprocessing by NVTabular
# + tags=[]
# %%writefile /wdl_train/preprocess.py
import os
import sys
import argparse
import glob
import time
from cudf.io.parquet import ParquetWriter
import numpy as np
import pandas as pd
import concurrent.futures as cf
from concurrent.futures import as_completed
import shutil
import dask_cudf
from dask_cuda import LocalCUDACluster
from dask.distributed import Client
from dask.utils import parse_bytes
from dask.delayed import delayed
import cudf
import rmm
import nvtabular as nvt
from nvtabular.io import Shuffle
from nvtabular.utils import device_mem_size
from nvtabular.ops import Categorify, Clip, FillMissing, HashBucket, LambdaOp, Normalize, Rename, Operator, get_embedding_sizes
# #%load_ext memory_profiler
import logging
logging.basicConfig(format='%(asctime)s %(message)s')
logging.root.setLevel(logging.NOTSET)
logging.getLogger('numba').setLevel(logging.WARNING)
logging.getLogger('asyncio').setLevel(logging.WARNING)
# define dataset schema
CATEGORICAL_COLUMNS=["C" + str(x) for x in range(1, 27)]
CONTINUOUS_COLUMNS=["I" + str(x) for x in range(1, 14)]
LABEL_COLUMNS = ['label']
COLUMNS = LABEL_COLUMNS + CONTINUOUS_COLUMNS + CATEGORICAL_COLUMNS
#/samples/criteo mode doesn't have dense features
criteo_COLUMN=LABEL_COLUMNS + CATEGORICAL_COLUMNS
#For new feature cross columns
CROSS_COLUMNS = []
NUM_INTEGER_COLUMNS = 13
NUM_CATEGORICAL_COLUMNS = 26
NUM_TOTAL_COLUMNS = 1 + NUM_INTEGER_COLUMNS + NUM_CATEGORICAL_COLUMNS
# Initialize RMM pool on ALL workers
def setup_rmm_pool(client, pool_size):
client.run(rmm.reinitialize, pool_allocator=True, initial_pool_size=pool_size)
return None
#compute the partition size with GB
def bytesto(bytes, to, bsize=1024):
a = {'k' : 1, 'm': 2, 'g' : 3, 't' : 4, 'p' : 5, 'e' : 6 }
r = float(bytes)
return bytes / (bsize ** a[to])
class FeatureCross(Operator):
def __init__(self, dependency):
self.dependency = dependency
def transform(self, columns, gdf):
new_df = type(gdf)()
for col in columns.names:
new_df[col] = gdf[col] + gdf[self.dependency]
return new_df
def dependencies(self):
return [self.dependency]
#process the data with NVTabular
def process_NVT(args):
if args.feature_cross_list:
feature_pairs = [pair.split("_") for pair in args.feature_cross_list.split(",")]
for pair in feature_pairs:
CROSS_COLUMNS.append(pair[0]+'_'+pair[1])
logging.info('NVTabular processing')
train_input = os.path.join(args.data_path, "train/train.txt")
val_input = os.path.join(args.data_path, "val/test.txt")
PREPROCESS_DIR_temp_train = os.path.join(args.out_path, 'train/temp-parquet-after-conversion')
PREPROCESS_DIR_temp_val = os.path.join(args.out_path, 'val/temp-parquet-after-conversion')
PREPROCESS_DIR_temp = [PREPROCESS_DIR_temp_train, PREPROCESS_DIR_temp_val]
train_output = os.path.join(args.out_path, "train")
val_output = os.path.join(args.out_path, "val")
# Make sure we have a clean parquet space for cudf conversion
for one_path in PREPROCESS_DIR_temp:
if os.path.exists(one_path):
shutil.rmtree(one_path)
os.mkdir(one_path)
## Get Dask Client
# Deploy a Single-Machine Multi-GPU Cluster
device_size = device_mem_size(kind="total")
cluster = None
if args.protocol == "ucx":
UCX_TLS = os.environ.get("UCX_TLS", "tcp,cuda_copy,cuda_ipc,sockcm")
os.environ["UCX_TLS"] = UCX_TLS
cluster = LocalCUDACluster(
protocol = args.protocol,
CUDA_VISIBLE_DEVICES = args.devices,
n_workers = len(args.devices.split(",")),
enable_nvlink=True,
device_memory_limit = int(device_size * args.device_limit_frac),
dashboard_address=":" + args.dashboard_port
)
else:
cluster = LocalCUDACluster(
protocol = args.protocol,
n_workers = len(args.devices.split(",")),
CUDA_VISIBLE_DEVICES = args.devices,
device_memory_limit = int(device_size * args.device_limit_frac),
dashboard_address=":" + args.dashboard_port
)
# Create the distributed client
client = Client(cluster)
if args.device_pool_frac > 0.01:
setup_rmm_pool(client, int(args.device_pool_frac*device_size))
#calculate the total processing time
runtime = time.time()
#test dataset without the label feature
if args.dataset_type == 'test':
global LABEL_COLUMNS
LABEL_COLUMNS = []
##-----------------------------------##
# Dask rapids converts txt to parquet
# Dask cudf dataframe = ddf
## train/valid txt to parquet
train_valid_paths = [(train_input,PREPROCESS_DIR_temp_train),(val_input,PREPROCESS_DIR_temp_val)]
for input, temp_output in train_valid_paths:
ddf = dask_cudf.read_csv(input,sep='\t',names=LABEL_COLUMNS + CONTINUOUS_COLUMNS + CATEGORICAL_COLUMNS)
## Convert label col to FP32
if args.parquet_format and args.dataset_type == 'train':
ddf["label"] = ddf['label'].astype('float32')
# Save it as parquet format for better memory usage
ddf.to_parquet(temp_output,header=True)
##-----------------------------------##
COLUMNS = LABEL_COLUMNS + CONTINUOUS_COLUMNS + CROSS_COLUMNS + CATEGORICAL_COLUMNS
train_paths = glob.glob(os.path.join(PREPROCESS_DIR_temp_train, "*.parquet"))
valid_paths = glob.glob(os.path.join(PREPROCESS_DIR_temp_val, "*.parquet"))
categorify_op = Categorify(freq_threshold=args.freq_limit)
cat_features = CATEGORICAL_COLUMNS >> categorify_op
cont_features = CONTINUOUS_COLUMNS >> FillMissing() >> Clip(min_value=0) >> Normalize()
cross_cat_op = Categorify(freq_threshold=args.freq_limit)
features = LABEL_COLUMNS
if args.criteo_mode == 0:
features += cont_features
if args.feature_cross_list:
feature_pairs = [pair.split("_") for pair in args.feature_cross_list.split(",")]
for pair in feature_pairs:
col0 = pair[0]
col1 = pair[1]
features += col0 >> FeatureCross(col1) >> Rename(postfix="_"+col1) >> cross_cat_op
features += cat_features
workflow = nvt.Workflow(features, client=client)
logging.info("Preprocessing")
output_format = 'hugectr'
if args.parquet_format:
output_format = 'parquet'
# just for /samples/criteo model
train_ds_iterator = nvt.Dataset(train_paths, engine='parquet', part_size=int(args.part_mem_frac * device_size))
valid_ds_iterator = nvt.Dataset(valid_paths, engine='parquet', part_size=int(args.part_mem_frac * device_size))
shuffle = None
if args.shuffle == "PER_WORKER":
shuffle = nvt.io.Shuffle.PER_WORKER
elif args.shuffle == "PER_PARTITION":
shuffle = nvt.io.Shuffle.PER_PARTITION
logging.info('Train Datasets Preprocessing.....')
dict_dtypes = {}
for col in CATEGORICAL_COLUMNS:
dict_dtypes[col] = np.int64
if not args.criteo_mode:
for col in CONTINUOUS_COLUMNS:
dict_dtypes[col] = np.float32
for col in CROSS_COLUMNS:
dict_dtypes[col] = np.int64
for col in LABEL_COLUMNS:
dict_dtypes[col] = np.float32
conts = CONTINUOUS_COLUMNS if not args.criteo_mode else []
workflow.fit(train_ds_iterator)
if output_format == 'hugectr':
workflow.transform(train_ds_iterator).to_hugectr(
cats=CATEGORICAL_COLUMNS + CROSS_COLUMNS,
conts=conts,
labels=LABEL_COLUMNS,
output_path=train_output,
shuffle=shuffle,
out_files_per_proc=args.out_files_per_proc,
num_threads=args.num_io_threads)
else:
workflow.transform(train_ds_iterator).to_parquet(
output_path=train_output,
dtypes=dict_dtypes,
cats=CATEGORICAL_COLUMNS + CROSS_COLUMNS,
conts=conts,
labels=LABEL_COLUMNS,
shuffle=shuffle,
out_files_per_proc=args.out_files_per_proc,
num_threads=args.num_io_threads)
###Getting slot size###
#--------------------##
embeddings_dict_cat = categorify_op.get_embedding_sizes(CATEGORICAL_COLUMNS)
embeddings_dict_cross = cross_cat_op.get_embedding_sizes(CROSS_COLUMNS)
embeddings = [embeddings_dict_cat[c][0] for c in CATEGORICAL_COLUMNS] + [embeddings_dict_cross[c][0] for c in CROSS_COLUMNS]
print(embeddings)
##--------------------##
logging.info('Valid Datasets Preprocessing.....')
if output_format == 'hugectr':
workflow.transform(valid_ds_iterator).to_hugectr(
cats=CATEGORICAL_COLUMNS + CROSS_COLUMNS,
conts=conts,
labels=LABEL_COLUMNS,
output_path=val_output,
shuffle=shuffle,
out_files_per_proc=args.out_files_per_proc,
num_threads=args.num_io_threads)
else:
workflow.transform(valid_ds_iterator).to_parquet(
output_path=val_output,
dtypes=dict_dtypes,
cats=CATEGORICAL_COLUMNS + CROSS_COLUMNS,
conts=conts,
labels=LABEL_COLUMNS,
shuffle=shuffle,
out_files_per_proc=args.out_files_per_proc,
num_threads=args.num_io_threads)
embeddings_dict_cat = categorify_op.get_embedding_sizes(CATEGORICAL_COLUMNS)
embeddings_dict_cross = cross_cat_op.get_embedding_sizes(CROSS_COLUMNS)
embeddings = [embeddings_dict_cat[c][0] for c in CATEGORICAL_COLUMNS] + [embeddings_dict_cross[c][0] for c in CROSS_COLUMNS]
print(embeddings)
##--------------------##
## Shutdown clusters
client.close()
logging.info('NVTabular processing done')
runtime = time.time() - runtime
print("\nDask-NVTabular Criteo Preprocessing")
print("--------------------------------------")
print(f"data_path | {args.data_path}")
print(f"output_path | {args.out_path}")
print(f"partition size | {'%.2f GB'%bytesto(int(args.part_mem_frac * device_size),'g')}")
print(f"protocol | {args.protocol}")
print(f"device(s) | {args.devices}")
print(f"rmm-pool-frac | {(args.device_pool_frac)}")
print(f"out-files-per-proc | {args.out_files_per_proc}")
print(f"num_io_threads | {args.num_io_threads}")
print(f"shuffle | {args.shuffle}")
print("======================================")
print(f"Runtime[s] | {runtime}")
print("======================================\n")
def parse_args():
parser = argparse.ArgumentParser(description=("Multi-GPU Criteo Preprocessing"))
#
# System Options
#
parser.add_argument("--data_path", type=str, help="Input dataset path (Required)")
parser.add_argument("--out_path", type=str, help="Directory path to write output (Required)")
parser.add_argument(
"-d",
"--devices",
default=os.environ.get("CUDA_VISIBLE_DEVICES", "0"),
type=str,
help='Comma-separated list of visible devices (e.g. "0,1,2,3"). '
)
parser.add_argument(
"-p",
"--protocol",
choices=["tcp", "ucx"],
default="tcp",
type=str,
help="Communication protocol to use (Default 'tcp')",
)
parser.add_argument(
"--device_limit_frac",
default=0.5,
type=float,
help="Worker device-memory limit as a fraction of GPU capacity (Default 0.8). "
)
parser.add_argument(
"--device_pool_frac",
default=0.9,
type=float,
help="RMM pool size for each worker as a fraction of GPU capacity (Default 0.9). "
"The RMM pool frac is the same for all GPUs, make sure each one has enough memory size",
)
parser.add_argument(
"--num_io_threads",
default=0,
type=int,
help="Number of threads to use when writing output data (Default 0). "
"If 0 is specified, multi-threading will not be used for IO.",
)
#
# Data-Decomposition Parameters
#
parser.add_argument(
"--part_mem_frac",
default=0.125,
type=float,
help="Maximum size desired for dataset partitions as a fraction "
"of GPU capacity (Default 0.125)",
)
parser.add_argument(
"--out_files_per_proc",
default=1,
type=int,
help="Number of output files to write on each worker (Default 1)",
)
#
# Preprocessing Options
#
parser.add_argument(
"-f",
"--freq_limit",
default=0,
type=int,
help="Frequency limit for categorical encoding (Default 0)",
)
parser.add_argument(
"-s",
"--shuffle",
choices=["PER_WORKER", "PER_PARTITION", "NONE"],
default="PER_PARTITION",
help="Shuffle algorithm to use when writing output data to disk (Default PER_PARTITION)",
)
parser.add_argument(
"--feature_cross_list", default=None, type=str, help="List of feature crossing cols (e.g. C1_C2, C3_C4)"
)
#
# Diagnostics Options
#
parser.add_argument(
"--profile",
metavar="PATH",
default=None,
type=str,
help="Specify a file path to export a Dask profile report (E.g. dask-report.html)."
"If this option is excluded from the command, not profile will be exported",
)
parser.add_argument(
"--dashboard_port",
default="8787",
type=str,
help="Specify the desired port of Dask's diagnostics-dashboard (Default `3787`). "
"The dashboard will be hosted at http://<IP>:<PORT>/status",
)
#
# Format
#
parser.add_argument('--criteo_mode', type=int, default=0)
parser.add_argument('--parquet_format', type=int, default=1)
parser.add_argument('--dataset_type', type=str, default='train')
args = parser.parse_args()
args.n_workers = len(args.devices.split(","))
return args
if __name__ == '__main__':
args = parse_args()
process_NVT(args)
# -
import pandas as pd
# !python3 /wdl_train/preprocess.py --data_path wdl_train/ --out_path wdl_train/ --freq_limit 6 --feature_cross_list C1_C2,C3_C4 --device_pool_frac 0.5 --devices "0" --num_io_threads 2
# ### 2.4 Checke the preprocessed training data
# !ls -ll /wdl_train/train
import pandas as pd
df = pd.read_parquet("/wdl_train/train/0.8870d61b8a1f4deca0f911acfb072999.parquet")
df.head(2)
# ## 3. WDL Model Training
# + tags=[]
# %%writefile './model.py'
import hugectr
#from mpi4py import MPI
solver = hugectr.CreateSolver(max_eval_batches = 4000,
batchsize_eval = 2720,
batchsize = 2720,
lr = 0.001,
vvgpu = [[2]],
repeat_dataset = True,
i64_input_key = True)
reader = hugectr.DataReaderParams(data_reader_type = hugectr.DataReaderType_t.Parquet,
source = ["./train/_file_list.txt"],
eval_source = "./val/_file_list.txt",
check_type = hugectr.Check_t.Non,
slot_size_array = [249058, 19561, 14212, 6890, 18592, 4, 6356, 1254, 52, 226170, 80508, 72308, 11, 2169, 7597, 61, 4, 923, 15, 249619, 168974, 243480, 68212, 9169, 75, 34, 278018, 415262])
optimizer = hugectr.CreateOptimizer(optimizer_type = hugectr.Optimizer_t.Adam,
update_type = hugectr.Update_t.Global,
beta1 = 0.9,
beta2 = 0.999,
epsilon = 0.0000001)
model = hugectr.Model(solver, reader, optimizer)
model.add(hugectr.Input(label_dim = 1, label_name = "label",
dense_dim = 13, dense_name = "dense",
data_reader_sparse_param_array =
[hugectr.DataReaderSparseParam("wide_data", 1, True, 2),
hugectr.DataReaderSparseParam("deep_data", 2, False, 26)]))
model.add(hugectr.SparseEmbedding(embedding_type = hugectr.Embedding_t.DistributedSlotSparseEmbeddingHash,
workspace_size_per_gpu_in_mb = 24,
embedding_vec_size = 1,
combiner = "sum",
sparse_embedding_name = "sparse_embedding2",
bottom_name = "wide_data",
optimizer = optimizer))
model.add(hugectr.SparseEmbedding(embedding_type = hugectr.Embedding_t.DistributedSlotSparseEmbeddingHash,
workspace_size_per_gpu_in_mb = 405,
embedding_vec_size = 16,
combiner = "sum",
sparse_embedding_name = "sparse_embedding1",
bottom_name = "deep_data",
optimizer = optimizer))
model.add(hugectr.DenseLayer(layer_type = hugectr.Layer_t.Reshape,
bottom_names = ["sparse_embedding1"],
top_names = ["reshape1"],
leading_dim=416))
model.add(hugectr.DenseLayer(layer_type = hugectr.Layer_t.Reshape,
bottom_names = ["sparse_embedding2"],
top_names = ["reshape2"],
leading_dim=2))
model.add(hugectr.DenseLayer(layer_type = hugectr.Layer_t.ReduceSum,
bottom_names = ["reshape2"],
top_names = ["wide_redn"],
axis = 1))
model.add(hugectr.DenseLayer(layer_type = hugectr.Layer_t.Concat,
bottom_names = ["reshape1", "dense"],
top_names = ["concat1"]))
model.add(hugectr.DenseLayer(layer_type = hugectr.Layer_t.InnerProduct,
bottom_names = ["concat1"],
top_names = ["fc1"],
num_output=1024))
model.add(hugectr.DenseLayer(layer_type = hugectr.Layer_t.ReLU,
bottom_names = ["fc1"],
top_names = ["relu1"]))
model.add(hugectr.DenseLayer(layer_type = hugectr.Layer_t.Dropout,
bottom_names = ["relu1"],
top_names = ["dropout1"],
dropout_rate=0.5))
model.add(hugectr.DenseLayer(layer_type = hugectr.Layer_t.InnerProduct,
bottom_names = ["dropout1"],
top_names = ["fc2"],
num_output=1024))
model.add(hugectr.DenseLayer(layer_type = hugectr.Layer_t.ReLU,
bottom_names = ["fc2"],
top_names = ["relu2"]))
model.add(hugectr.DenseLayer(layer_type = hugectr.Layer_t.Dropout,
bottom_names = ["relu2"],
top_names = ["dropout2"],
dropout_rate=0.5))
model.add(hugectr.DenseLayer(layer_type = hugectr.Layer_t.InnerProduct,
bottom_names = ["dropout2"],
top_names = ["fc3"],
num_output=1))
model.add(hugectr.DenseLayer(layer_type = hugectr.Layer_t.Add,
bottom_names = ["fc3", "wide_redn"],
top_names = ["add1"]))
model.add(hugectr.DenseLayer(layer_type = hugectr.Layer_t.BinaryCrossEntropyLoss,
bottom_names = ["add1", "label"],
top_names = ["loss"]))
model.compile()
model.summary()
model.fit(max_iter = 21000, display = 1000, eval_interval = 4000, snapshot = 20000, snapshot_prefix = "wdl")
model.graph_to_json(graph_config_file = "wdl.json")
# -
# !python ./model.py
# !ls -ll
# !python /wdl_infer/wdl_python_infer.py "wdl" "/wdl_infer/model/wdl/1/wdl.json" "/wdl_infer/model/wdl/1/wdl_dense_20000.model" "/wdl_infer/model/wdl/1/wdl0_sparse_20000.model/,/wdl_infer/model/wdl/1/wdl1_sparse_20000.model" "/wdl_infer/first_ten.csv"
# # 4. Prepare Inference Request
# !ls -l /wdl_train/val
import pandas as pd
df = pd.read_parquet("/wdl_train/val/0.110d099942694a5cbf1b71eb73e10f27.parquet")
df.head()
df.head(10).to_csv('/wdl_train/infer_test.csv', sep=',', index=False,header=True)
# # 5 Create prediction scripts
# +
# %%writefile '/wdl_train/wdl_predict.py'
from hugectr.inference import InferenceParams, CreateInferenceSession
import hugectr
import pandas as pd
import numpy as np
import sys
from mpi4py import MPI
def wdl_inference(model_name, network_file, dense_file, embedding_file_list, data_file,enable_cache,dbtype=hugectr.Database_t.Local,rocksdb_path=""):
CATEGORICAL_COLUMNS=["C" + str(x) for x in range(1, 27)]+["C1_C2","C3_C4"]
CONTINUOUS_COLUMNS=["I" + str(x) for x in range(1, 14)]
LABEL_COLUMNS = ['label']
emb_size = [249058, 19561, 14212, 6890, 18592, 4, 6356, 1254, 52, 226170, 80508, 72308, 11, 2169, 7597, 61, 4, 923, 15, 249619, 168974, 243480, 68212, 9169, 75, 34, 278018, 415262]
shift = np.insert(np.cumsum(emb_size), 0, 0)[:-1]
test_df=pd.read_csv(data_file,sep=',')
config_file = network_file
row_ptrs = list(range(0,21))+list(range(0,261))
dense_features = list(test_df[CONTINUOUS_COLUMNS].values.flatten())
test_df[CATEGORICAL_COLUMNS].astype(np.int64)
embedding_columns = list((test_df[CATEGORICAL_COLUMNS]+shift).values.flatten())
# create parameter server, embedding cache and inference session
inference_params = InferenceParams(model_name = model_name,
max_batchsize = 64,
hit_rate_threshold = 0.5,
dense_model_file = dense_file,
sparse_model_files = embedding_file_list,
device_id = 2,
use_gpu_embedding_cache = enable_cache,
cache_size_percentage = 0.9,
i64_input_key = True,
use_mixed_precision = False,
db_type = dbtype,
rocksdb_path=rocksdb_path,
cache_size_percentage_redis=0.5)
inference_session = CreateInferenceSession(config_file, inference_params)
output = inference_session.predict(dense_features, embedding_columns, row_ptrs)
print("WDL multi-embedding table inference result is {}".format(output))
if __name__ == "__main__":
model_name = sys.argv[1]
print("{} multi-embedding table prediction".format(model_name))
network_file = sys.argv[2]
print("{} multi-embedding table prediction network is {}".format(model_name,network_file))
dense_file = sys.argv[3]
print("{} multi-embedding table prediction dense file is {}".format(model_name,dense_file))
embedding_file_list = str(sys.argv[4]).split(',')
print("{} multi-embedding table prediction sparse files are {}".format(model_name,embedding_file_list))
data_file = sys.argv[5]
print("{} multi-embedding table prediction input data path is {}".format(model_name,data_file))
input_dbtype = sys.argv[6]
print("{} multi-embedding table prediction input dbtype path is {}".format(model_name,input_dbtype))
if input_dbtype=="local":
wdl_inference(model_name, network_file, dense_file, embedding_file_list, data_file, True, hugectr.Database_t.Local)
if input_dbtype=="rocksdb":
rocksdb_path = sys.argv[7]
print("{} multi-embedding table prediction rocksdb_path path is {}".format(model_name,rocksdb_path))
wdl_inference(model_name, network_file, dense_file, embedding_file_list, data_file, True, hugectr.Database_t.RocksDB,rocksdb_path)
# -
# # 6. Prediction
# Use different types of databases as local Parameter Server to get the wdl model prediction results.
# ## 6.1 Load model embedding tables into local memory as Parameter Server
# !python /wdl_train/wdl_predict.py "wdl" "/wdl_infer/model/wdl/1/wdl.json" \
# "/wdl_infer/model/wdl/1/wdl_dense_20000.model" \
# "/wdl_infer/model/wdl/1/wdl0_sparse_20000.model/,/wdl_infer/model/wdl/1/wdl1_sparse_20000.model" \
# "/wdl_train/infer_test.csv" \
# "local"
# ## 6.2 Load model embedding tables into local RocksDB as Parameter Server
# Create a RocksDB directory with read and write permissions for storing model embedded tables
# !mkdir -p -m 700 /wdl_train/rocksdb
# !python /wdl_train/wdl_predict.py "wdl" "/wdl_infer/model/wdl/1/wdl.json" \
# "/wdl_infer/model/wdl/1/wdl_dense_20000.model" \
# "/wdl_infer/model/wdl/1/wdl0_sparse_20000.model/,/wdl_infer/model/wdl/1/wdl1_sparse_20000.model" \
# "/wdl_train/infer_test.csv" \
# "rocksdb" "/wdl_train/rocksdb"
| notebooks/hugectr_wdl_prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Classical ML Algorithm Legends
# ### Table of Contents
#
# * [Decision Trees](#DecisionTrees)
# * [Decision tree learning](#Learning)
# * [Decision tree pruning](#Pruning)
# * [Boosted trees!](#Algorithms)
# * [Indexing](#Indexing)
# * [Other useful links](#Finally...)
# # DecisionTrees
#
# **prerequisites**
#
#
# * Information Gain:
# In decision tree learning, Information gain ratio is a ratio of information gain to the intrinsic information. It was proposed to reduce a bias towards multi-valued attributes by taking the number and size of branches into account when choosing an attribute.
# Check [this tutorial](https://www.youtube.com/watch?v=FuTRucXB9rA) for more information on this topic.
#
# * [Decision Trees](https://en.wikipedia.org/wiki/Decision_tree)
# 
# Generally there are two types of decision trees:
# 1. [**Classification tree** ](https://www.solver.com/classification-tree)
# : A Classification tree labels, records, and assigns variables to discrete classes.
# 
# 2. [**Regression tree**](https://www.solver.com/regression-trees): A regression tree is built through a process known as binary recursive partitioning, which is an iterative process that splits the data into partitions or branches, and then continues splitting each partition into smaller groups as the method moves up each branch.
# 
# ## Learning
#
# Decision Tree Learning is a type of Supervised Machine Learning where the data is continuously split according to a certain parameter.
#
# you can check out [this](https://en.wikipedia.org/wiki/Decision_tree_learning) wikipedia page for more thorough details.
#
# ### Algorithms
# 1. [Iterative Dichotomiser 3 (ID3)](https://en.wikipedia.org/wiki/ID3_algorithm) - ([sample implementation](https://sefiks.com/2017/11/20/a-step-by-step-id3-decision-tree-example/))
# 2. [C4.5](https://en.wikipedia.org/wiki/C4.5_algorithm) (Successor of ID3) - ([sample implementation](https://sefiks.com/2018/05/13/a-step-by-step-c4-5-decision-tree-example/))
# 3. Classification And Regression Tree - ([useful article](https://sefiks.com/2018/08/27/a-step-by-step-cart-decision-tree-example/))
# 4. Chi-square automatic interaction detection - ([useful article](https://sefiks.com/2020/03/18/a-step-by-step-chaid-decision-tree-example/))
# 5. Multivariate adaptive regression spline ([useful tutorial](https://www.youtube.com/watch?v=9COLjUxSzx8))
# ## Avoid Overfitting
# Decision trees are prone to overfitting, especially when a tree is particularly deep. This is due to the amount of specificity we look at leading to smaller sample of events that meet the previous assumptions. This small sample could lead to unsound conclusions.
# In decision trees, pruning is a process which is applied to control or limit the depth (size) of the trees. By default, decision tree model hyperparameters were created to grow the tree into its full depth. These trees are called fully-grown trees which are always overfitting.
#
# ### Pruning
# Pruning reduces the size of decision trees by removing parts of the tree that do not provide power to classify instances. Check [this Wikipedia link](https://en.wikipedia.org/wiki/Decision_tree_pruning) for full explanation.
#
# **Pre-pruning**: As the names suggest, pre-pruning or early stopping involves stopping the tree before it has completed classifying the training set and post-pruning refers to pruning the tree after it has finished.
#
# **Post-pruning**: Post-pruning a decision tree implies that we begin by generating the (complete) tree and then adjust it with the aim of improving the accuracy on unseen instances.
#
# Also see:
#
# * [pre-pruning and post-pruning](https://www.displayr.com/machine-learning-pruning-decision-trees/)
#
# * [Bottom-up pruning](https://en.wikipedia.org/wiki/Decision_tree_pruning)
#
# * [Top-down pruning](https://en.wikipedia.org/wiki/Decision_tree_pruning)
# <br>
#
# **Algorithms**
#
# * [Reduced error pruning](https://www.cs.auckland.ac.nz/~pat/706_98/ln/node90.html): In this algorithm, by starting at the leaves, each node is replaced with its most popular class. If the prediction accuracy is not affected then the change is kept. While somewhat naive, reduced error pruning has the advantage of simplicity and speed.
# * [Cost complexity pruning](http://mlwiki.org/index.php/Cost-Complexity_Pruning): This generates a series of trees and at each step a tree is made from the previous one by subtracting a subtree from it and replacing it with a leaf node with value chosen as in the tree building algorithm
#
#
#
# Some techniques, often called **ensemble methods**, construct more than one decision tree:
# <ol>
# <li>Boosted trees</li>
# <li>Rotation forest</li>
# <li>Bootstrap aggregated</li>
# </ol>
#
# **ensemble methods**: In statistics and machine learning, ensemble methods use multiple learning algorithms to obtain better predictive performance than could be obtained from any of the constituent learning algorithms alone.
# * **Boosted trees**: Incrementally building an ensemble by training each new instance to emphasize the training instances previously mis-modeled. These can be used for regression-type and classification-type problems.
#
# * **Rotation forest**: in which every decision tree is trained by first applying principal component analysis (PCA) on a random subset of the input features.
#
# * **Bootstrap aggregated**: bagged decision trees, an early ensemble method, builds multiple decision trees by repeatedly resampling training data with replacement, and voting the trees for a consensus prediction.
# [Bagging](https://en.wikipedia.org/wiki/Bootstrap_aggregating), also known as bootstrap aggregation, is the ensemble learning method that is commonly used to reduce variance within a noisy dataset.
# [Here](https://www.cs.cornell.edu/courses/cs4780/2018fa/lectures/lecturenote18.html) is the link of bagging topic in ML course at Cornell and [this](https://www.youtube.com/watch?v=2Mg8QD0F1dQ) is a simple video explaining Bootstrap aggregating bagging.
# The term ['Boosting'](https://en.wikipedia.org/wiki/Boosting_(machine_learning)) refers to a family of algorithms which converts weak learner to strong learners.
# [This video](https://www.youtube.com/watch?v=MIPkK5ZAsms) is *A Short Introduction to Boosting*. You can read [this Medium article](https://medium.com/greyatom/a-quick-guide-to-boosting-in-ml-acf7c1585cb5) for a better perspective on the topic.
# There are many boosting algorithms. The original ones are as followed:
# <ol>
# <li>a recursive majority gate formulation</li>
# <li>boost by majority</li>
# </ol>
#
# check [this paper](https://web.archive.org/web/20121010030839/http://www.cs.princeton.edu/~schapire/papers/strengthofweak.pdf) for more information on this topic.
#
# But these algorithms were not adaptive and could not take full advantage of the weak learners. Later, AdaBoost was developed, which was an adaptive boosting algorithm.
# [AdaBoost]((https://en.wikipedia.org/wiki/AdaBoost)), short for Adaptive Boosting, is a machine learning meta-algorithm. It can be used in conjunction with many other types of learning algorithms to improve performance. [This video](https://www.youtube.com/watch?v=LsK-xG1cLYA) explains the algorithm clearly.
# # Indexing
# [This link](https://cilvr.cs.nyu.edu/diglib/lsml/lecture12_indexing.pdf) explains indexing in machine learning. You can also use [this](https://chartio.com/learn/databases/how-does-indexing-work/) tutorial to see how it works and what it does. Also check [this paper](http://learningsys.org/nips17/assets/papers/paper_22.pdf) out for more information.
# ## Finally...
# Check these links out for more information on topics covered in this notebook.
#
# * [How To Implement The Decision Tree Algorithm From Scratch In Python](https://machinelearningmastery.com/implement-decision-tree-algorithm-scratch-python/)
#
# * [Let’s Solve Overfitting! Quick Guide to Cost Complexity Pruning of Decision Trees](https://www.analyticsvidhya.com/blog/2020/10/cost-complexity-pruning-decision-trees/)
#
# * [Minimax Algorithm with Alpha-beta pruning](https://www.hackerearth.com/blog/developers/minimax-algorithm-alpha-beta-pruning/)
#
# * [Post pruning decision trees with cost complexity pruning](https://scikit-learn.org/stable/auto_examples/tree/plot_cost_complexity_pruning.html)
#
# * [Information Gain and Mutual Information for Machine Learning](https://machinelearningmastery.com/information-gain-and-mutual-information/)
#
# * [How to Develop a Bagging Ensemble with Python](https://machinelearningmastery.com/bagging-ensemble-with-python/)
#
# * [Boosting in Machine Learning and the Implementation of XGBoost in Python](https://towardsdatascience.com/boosting-in-machine-learning-and-the-implementation-of-xgboost-in-python-fb5365e9f2a0)
#
# * [Implementing the AdaBoost Algorithm From Scratch](https://www.kdnuggets.com/2020/12/implementing-adaboost-algorithm-from-scratch.html)
#
# * [Last, but not least!](https://www.youtube.com/watch?v=dQw4w9WgXcQ)
| notebooks/decision_trees/index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Euler Problem 22
# ================
#
# Using [names.txt](files/p022_names.txt) (right click and 'Save Link/Target As...'), a 46K text file containing over five-thousand first names, begin by sorting it into alphabetical order. Then working out the alphabetical value for each name, multiply this value by its alphabetical position in the list to obtain a name score.
#
# For example, when the list is sorted into alphabetical order, COLIN, which is worth 3 + 15 + 12 + 9 + 14 = 53, is the 938th name in the list. So, COLIN would obtain a score of 938 × 53 = 49714.
#
# What is the total of all the name scores in the file?
with open("data/p022_names.txt", 'r') as f:
names = sorted(f.readline().replace('"','').split(','))
print(sum((i+1)*sum(ord(c)-64 for c in name) for i, name in enumerate(names)))
| Euler 022 - Names score.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_py371)
# language: python
# name: conda_py371
# ---
# %load_ext autoreload
# %autoreload
exec(open("plot_utils.py").read())
def fix_logliks(dir_name,country_name):
file_name = dir_name+country_name+'.npz'
data = load_data(file_name, country_name,0)
tau_prior = get_τ_prior(start_date, ndays, country_name, τ_model)
priors = [log_prior(s,tau_prior,τ_model) for s in sample]
logliks = lnprobability.reshape(-1)-priors
npz = np.load(file_name)
dic = {k:npz[k] for k in npz.files}
dic['logliks'] = logliks
np.savez_compressed(file_name,**dic)
# %%time
import os,fnmatch
dir_name = '../{}/{}/inference/'.format('output','2020-04-30-prior-walkers-model2-normal')
countries = [a[:-4] for a in fnmatch.filter(os.listdir(dir_name), '*.npz')]
for country_name in countries:
print(country_name)
fix_logliks(dir_name, country_name)
| src/fix_logliks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/CharlesPoletowin/YCBS-273/blob/master/Lecture4_conv_pool.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="5nE33IvZIEl9" colab_type="code" colab={}
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from scipy.ndimage.filters import convolve
from scipy.signal import convolve2d
from scipy import misc
# + [markdown] id="vABujkpx9u7c" colab_type="text"
# # Convolution
# + [markdown] id="BB4lkWP_5ItZ" colab_type="text"
# ## 1-d convolution
#
# The mathematical definition of convolution of two 1-dimensional signals is given by
#
# $$
# (i * f)(T) = \sum_{t=0}^{T}i(t)f(T-t)
# $$
#
# Let $i$ be the input signal and $f$ be called the filter. If we had to overly simplify, given an appropriate filter the result of the convolution indicates the changes in input signal.
# + id="Pgd9H8Q53AI3" colab_type="code" colab={}
i = np.array([0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 0, 0, 0, 0, 0], dtype='float32')
f = np.array([-1,1], dtype='float32')
c = np.convolve(i, f)
# + [markdown] id="GiosjoTb6xVT" colab_type="text"
# The spikes in result of the convolution, $c$, indicates the change in the input signal, $i$.
# + id="KxQ66I-v3ALj" colab_type="code" outputId="5b618e51-d074-4a5d-a7ac-a896b5ef4c62" colab={"base_uri": "https://localhost:8080/", "height": 287}
plt.subplot(311)
plt.plot(i, 'o-')
plt.subplot(312)
plt.plot(f, 'o')
plt.subplot(313)
plt.plot(c, 'o-')
# + [markdown] id="SC41DzlX7YT1" colab_type="text"
# ## 2-d convolution
#
# While the mathematical equation for 2-d convolution is slightly more complicated, it is similar to 1-d convolution in spirit. The result of the convulution of a filter with an input signal indicates changes in the same. The most popular 2-d signal is an image. A stark in the pixel values of an image indiacte an edge. Below example of filteres which approximately detect horizontal/vertical edges.
# + id="s3qBFpDzyI9Q" colab_type="code" outputId="881975f7-ca0f-4017-f619-669ea3a26d5c" colab={"base_uri": "https://localhost:8080/", "height": 304}
img = misc.ascent()
print('Image shape: ', img.shape)
plt.imshow(img, cmap='gray')
# + [markdown] id="PBpijoA0816K" colab_type="text"
# ### Horizontal edge filter
# + id="6OHXF6BcyJAZ" colab_type="code" outputId="4670fdb8-3884-488d-a971-67d5e3ef5e3e" colab={"base_uri": "https://localhost:8080/", "height": 286}
# horizontal lines where top is white and bottom is black
h_filter = np.array([[ 2, 2, 2],
[ 0, 0, 0],
[-2, -2, -2]])
plt.imshow(h_filter, cmap='gray')
# + id="VHhidf87yJD1" colab_type="code" outputId="259cb212-bee9-4471-9ea8-f12cad11f0c5" colab={"base_uri": "https://localhost:8080/", "height": 286}
res = convolve2d(img, h_filter)
# enhanced some parts and suppressed other parts
plt.imshow(res, cmap='gray')
# + [markdown] colab_type="text" id="9sTJeSzH8_en"
# ### Vertical edge filter
# + id="YEt7HdZiyJKL" colab_type="code" outputId="7259bfa0-a755-4249-a4d4-1996632ed3f9" colab={"base_uri": "https://localhost:8080/", "height": 286}
# vertical lines where top is white and bottom is black
v_filter = np.array([[ 2, 0, -2],
[ 2, 0, -2],
[2, 0, -2]])
plt.imshow(v_filter, cmap='gray')
# + id="SYxhbqweyiMQ" colab_type="code" outputId="5a60f7cc-9237-4139-dfd2-a31bcb6083eb" colab={"base_uri": "https://localhost:8080/", "height": 286}
res = convolve2d(img, v_filter)
# enhanced some parts and suppressed other parts
plt.imshow(res, cmap='gray')
# + [markdown] id="OIv7PqakBUwY" colab_type="text"
# ## Convolutional layer in PyTorch
# + id="rn9DLULrBZtQ" colab_type="code" colab={}
import torch
from torch import nn
class ConvLayer(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(in_channels=1, out_channels=1, kernel_size=(3, 3), stride=1, padding=0)
def forward(self, xb):
return self.conv1(xb)
conv_layer = ConvLayer()
# + id="RVTfAKADTq6_" colab_type="code" outputId="01ec2a0a-7a98-4719-ec83-e66258d5acfa" colab={"base_uri": "https://localhost:8080/", "height": 87}
conv_layer.conv1.weight
# + id="JiOSyMV9Dob1" colab_type="code" outputId="26381f88-6ab9-4446-a16e-1869e711e1a2" colab={"base_uri": "https://localhost:8080/", "height": 339}
img = torch.tensor(img).view(1, 1, 512, 512)
conv_layer.conv1.weight = nn.Parameter(torch.FloatTensor([[[[ 2, 0, -2],
[ 2, 0, -2],
[2, 0, -2]]]]))
out = conv_layer(img.float())
plt.imshow(out.view(510, 510).detach().numpy(), cmap='gray')
# + id="5tYRhWpgFPBG" colab_type="code" outputId="1cedd317-f424-4af1-adef-163611112f67" colab={"base_uri": "https://localhost:8080/", "height": 286}
plt.imshow(conv_layer.conv1.weight.view(3, 3).detach().numpy(), cmap='gray')
# + [markdown] id="ujEoeH_Z-l9j" colab_type="text"
# # Pooling
#
# Pooling is useful to induce spacial invariance in our model. It also helps to reduce the parameters of our model by reducing the dimension of input.
# + [markdown] id="_cQUrIzPCES5" colab_type="text"
# ## Pooling layers in PyTorch
# + [markdown] id="l-RHny3uHA5a" colab_type="text"
# ### Max pooling
# + id="sZtMZUuZCCqT" colab_type="code" colab={}
import torch
from torch import nn
class MaxPoolLayer(nn.Module):
def __init__(self):
super().__init__()
self.pool1 = nn.MaxPool2d(kernel_size=(4, 4), padding=0)
def forward(self, xb):
return self.pool1(xb)
max_pool_layer = MaxPoolLayer()
# + id="EcYB_xwKHG4y" colab_type="code" outputId="559f5290-1991-43a6-c8a7-988b30449ff1" colab={"base_uri": "https://localhost:8080/", "height": 339}
img = torch.tensor(img).view(1, 1, 512, 512)
out = max_pool_layer(img.float())
plt.imshow(out.view(128, 128).detach().numpy(), cmap='gray')
# + [markdown] id="C4CMws21HDbi" colab_type="text"
# ### Average pooling
# + id="96LEPXTxHXl9" colab_type="code" colab={}
import torch
from torch import nn
class AvgPoolLayer(nn.Module):
def __init__(self):
super().__init__()
self.pool1 = nn.AvgPool2d(kernel_size=(4, 4), padding=0)
def forward(self, xb):
return self.pool1(xb)
avg_pool_layer = AvgPoolLayer()
# + id="jW5wz6QBHdYQ" colab_type="code" outputId="089d1aeb-8f92-43a8-e9d5-3011375ca561" colab={"base_uri": "https://localhost:8080/", "height": 339}
img = torch.tensor(img).view(1, 1, 512, 512)
out = avg_pool_layer(img.float())
plt.imshow(out.view(128, 128).detach().numpy(), cmap='gray')
| Lecture4_conv_pool.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="y_bBSTWDFhYX"
# # Import and read
# + id="7lrrr5BfBuPy"
import pandas as pd
# + colab={"base_uri": "https://localhost:8080/", "height": 332} id="0b8t_G7SFULI" outputId="73b85cd3-43c8-4836-b26a-f27354f6e3d1"
df = pd.read_csv('https://raw.githubusercontent.com/dayanandv/Data-Science/main/dataset/property%20data.csv')
df
# + colab={"base_uri": "https://localhost:8080/"} id="DxEPJZ21lB4v" outputId="7e508ee5-91c6-4e1e-f44d-2dcda1772744"
df.dtypes
# + [markdown] id="D5T6R7PNFm_d"
# # Check for null values
# + id="hF8EDKp6Fcl8" colab={"base_uri": "https://localhost:8080/"} outputId="de7590a7-a9ce-4b13-e00b-4c03d1d3b0ff"
df['SQ_FT'].isnull()
# + colab={"base_uri": "https://localhost:8080/", "height": 332} id="pqKXWM1vmp5L" outputId="2a0563dc-64fb-4f46-a10f-85e3b651adae"
missing_values = ['na', '--', 'n/a', 'NA', '-']
df = pd.read_csv('https://raw.githubusercontent.com/dayanandv/Data-Science/main/dataset/property%20data.csv', na_values=missing_values)
df
# + [markdown] id="N-B_mqBkGEY9"
# # Replace wrong category of data in columns
# + id="C_W36NLDGI2x" colab={"base_uri": "https://localhost:8080/", "height": 332} outputId="51eb39fb-39fe-413d-d7eb-3fbbec62d77b"
import numpy as np
# Using regular expressions to match and replace numbers/strings in columns where they are not expected
df['OWN_OCCUPIED'] = df['OWN_OCCUPIED'].replace(r'[0-9]+', np.nan, regex = True)
df['NUM_BATH'] = df['NUM_BATH'].replace(r'[A-Za-z]+', np.nan, regex = True)
df
# + [markdown] id="8X-LPZ0pGbYh"
# # Summary stats of missing data
# + [markdown] id="-CT0xPmtEJqP"
# Number of null values per column
# + id="qLWiOW1BGfKT" colab={"base_uri": "https://localhost:8080/"} outputId="de4162a3-89d7-4953-c9dc-8ceb1eae3b45"
df.isnull().sum()
# + [markdown] id="_vxgnV8AEN4k"
# Total number of null values in the dataframe
# + colab={"base_uri": "https://localhost:8080/"} id="uErOcekNES8B" outputId="4a9b9446-9540-4291-dc7c-cc600a212fd2"
df.isnull().sum().sum()
# + [markdown] id="ayVGdByaGsFR"
# # Replacing missing values in the whole dataframe
# + [markdown] id="2tQqD2xuFxuT"
# Backfill
# + id="_AJ5FSxbGyRv" colab={"base_uri": "https://localhost:8080/", "height": 332} outputId="4051730c-5c54-416b-8104-f0f8c2e3198c"
# Backfill
df.fillna(method = 'bfill')
# + [markdown] id="UAiVjWBGFujt"
# Fill with a custom value
# + colab={"base_uri": "https://localhost:8080/", "height": 332} id="RVaNKTjWFCUN" outputId="37108e66-bd59-409c-82a5-7cb0eec627cc"
#df.replace(to_replace=np.nan, value=-99)
df.fillna('-99')
# + [markdown] id="v3ZnojIeG4pr"
# # Dropping NaN values
# + colab={"base_uri": "https://localhost:8080/", "height": 81} id="ii3cEKj4Gowc" outputId="d5bd4201-e951-4af3-be7c-8076dde10f36"
df.dropna()
# + id="O5rrYi8RHbSQ" colab={"base_uri": "https://localhost:8080/", "height": 332} outputId="21f28391-5795-4234-a1e5-eba06425162b"
df.dropna(how ='all') # Drop only if all values of a row are nans
# + [markdown] id="EDTFwbC4Hbqn"
# # Imputation using sklearn SimpleImputer
# Strategy argument can take the values – ‘mean'(default), ‘median’, ‘most_frequent’ and ‘constant’
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="iNVDjNMRHe1I" outputId="3dc832b7-1c73-43c6-e305-5bffa1d1df78"
from sklearn.impute import SimpleImputer
dataset = pd.read_csv('https://raw.githubusercontent.com/dayanandv/Data-Science/main/dataset/pima-indians-diabetes.csv', skiprows=1, header=None)
dataset
# + colab={"base_uri": "https://localhost:8080/"} id="aF2zO5VtIcuI" outputId="0e7fb5ec-d743-4c8c-9cba-47a890ae748f"
dataset.isnull().sum().sum()
# + [markdown] id="8GYrM0ZvIztn"
# Replace all '0's in the first five columns with nan
# + id="EF57wZAGIF7B" colab={"base_uri": "https://localhost:8080/", "height": 424} outputId="c23ebb20-60c4-41c8-8ec9-98bffc8d16bc"
dataset[[1,2,3,4,5]] = dataset[[1,2,3,4,5]].replace(0, np.nan)
dataset
#values = dataset.values
#values
# + [markdown] id="e5IzT0lLI55t"
# Replace all nan with column's mean
# + id="_BSkyQ_CI9wf" colab={"base_uri": "https://localhost:8080/"} outputId="e4f74569-5eeb-451e-d652-27cff28bd121"
imputer = SimpleImputer(missing_values=np.nan, strategy='mean')
transformed_values = imputer.fit_transform(values)
transformed_values
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="mk0yJUMzIScd" outputId="bcb6a0bf-317e-4a4b-c6c9-8bc567376279"
dataset_transformed = pd.DataFrame(transformed_values)
dataset_transformed
# + colab={"base_uri": "https://localhost:8080/"} id="I6xW7qVOIsXl" outputId="b91e0319-d954-462d-a4ae-05ea5eb066ac"
dataset_transformed.isnull().sum().sum()
| Missing_Values.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# ---
# description: Create your Continual Learning Benchmark and Start Prototyping
# ---
#
# # Benchmarks
#
# Welcome to the "_benchmarks_" tutorial of the "_From Zero to Hero_" series. In this part we will present the functionalities offered by the `Benchmarks` module.
# + pycharm={"name": "#%%\n"}
# !pip install git+https://github.com/ContinualAI/avalanche.git
# + [markdown] pycharm={"name": "#%% md\n"}
# ## 🎯 Nomenclature
#
# First off, let's clarify a bit the nomenclature we are going to use, introducing the following terms: `Datasets`, `Scenarios`, `Benchmarks` and `Generators`.
#
# * By `Dataset` we mean a **collection of examples** that can be used for training or testing purposes but not already organized to be processed as a stream of batches or tasks. Since Avalanche is based on Pytorch, our Datasets are [torch.utils.Datasets](https://pytorch.org/docs/stable/_modules/torch/utils/data/dataset.html#Dataset) objects.
# * By `Scenario` we mean a **particular setting**, i.e. specificities about the continual stream of data, a continual learning algorithm will face.
# * By `Benchmark` we mean a well-defined and carefully thought **combination of a scenario with one or multiple datasets** that we can use to asses our continual learning algorithms.
# * By `Generator` we mean a function that **given a specific scenario and a dataset can generate a Benchmark**.
#
# ## 📚 The Benchmarks Module
#
# The `bechmarks` module offers 3 types of utils:
#
# * **Datasets**: all the Pytorch datasets plus additional ones prepared by our community and particularly interesting for continual learning.
# * **Classic Benchmarks**: classic benchmarks used in CL litterature ready to be used with great flexibility.
# * **Benchmarks Generators**: a set of functions you can use to create your own benchmark starting from any kind of data and scenario. In particular, we distinguish two type of generators: `Specific` and `Generic`. The first ones will let you create a benchmark based on a clear scenarios and Pytorch dataset\(s\); the latters, instead, are more generic and flexible, both in terms of scenario definition then in terms of type of data they can manage.
# * _Specific_:
# * **nc\_benchmark**: given one or multiple datasets it creates a benchmark instance based on scenarios where _New Classes_ \(NC\) are encountered over time. Notable scenarios that can be created using this utility include _Class-Incremental_, _Task-Incremental_ and _Task-Agnostic_ scenarios.
# * **ni\_benchmark**: it creates a benchmark instance based on scenarios where _New Instances_ \(NI\), i.e. new examples of the same classes are encountered over time. Notable scenarios that can be created using this utility include _Domain-Incremental_ scenarios.
# * _Generic_:
# * **filelist\_benchmark**: It creates a benchmark instance given a list of filelists.
# * **paths\_benchmark**: It creates a benchmark instance given a list of file paths and class labels.
# * **tensors\_benchmark**: It creates a benchmark instance given a list of tensors.
# * **dataset\_benchmark**: It creates a benchmark instance given a list of pytorch datasets.
#
# But let's see how we can use this module in practice!
#
# ## 🖼️ Datasets
#
# Let's start with the `Datasets`. As we previously hinted, in _Avalanche_ you'll find all the standard Pytorch Datasets available in the torchvision package as well as a few others that are useful for continual learning but not already officially available within the Pytorch ecosystem.
# + pycharm={"name": "#%%\n"}
import torch
import torchvision
from avalanche.benchmarks.datasets import MNIST, FashionMNIST, KMNIST, EMNIST, \
QMNIST, FakeData, CocoCaptions, CocoDetection, LSUN, ImageNet, CIFAR10, \
CIFAR100, STL10, SVHN, PhotoTour, SBU, Flickr8k, Flickr30k, VOCDetection, \
VOCSegmentation, Cityscapes, SBDataset, USPS, Kinetics400, HMDB51, UCF101, \
CelebA, CORe50Dataset, TinyImagenet, CUB200, OpenLORIS
# As we would simply do with any Pytorch dataset we can create the train and
# test sets from it. We could use any of the above imported Datasets, but let's
# just try to use the standard MNIST.
train_MNIST = MNIST(
'./data/mnist', train=True, download=True, transform=torchvision.transforms.ToTensor()
)
test_MNIST = MNIST(
'./data/mnist', train=False, download=True, transform=torchvision.transforms.ToTensor()
)
# Given these two sets we can simply iterate them to get the examples one by one
for i, example in enumerate(train_MNIST):
pass
print("Num. examples processed: {}".format(i))
# or use a Pytorch DataLoader
train_loader = torch.utils.data.DataLoader(
train_MNIST, batch_size=32, shuffle=True
)
for i, (x, y) in enumerate(train_loader):
pass
print("Num. mini-batch processed: {}".format(i))
# -
# Of course also the basic utilities `ImageFolder` and `DatasetFolder` can be used. These are two classes that you can use to create a Pytorch Dataset directly from your files \(following a particular structure\). You can read more about these in the Pytorch official documentation [here](https://pytorch.org/vision/stable/datasets.html#torchvision.datasets.ImageFolder).
#
# We also provide an additional `FilelistDataset` and `AvalancheDataset` classes. The former to construct a dataset from a filelist [\(caffe style\)](https://ceciliavision.wordpress.com/2016/03/08/caffedata-layer/) pointing to files anywhere on the disk. The latter to augment the basic Pytorch Dataset functionalities with an extention to better deal with a stack of transformations to be used during train and test.
# + pycharm={"name": "#%%\n"}
from avalanche.benchmarks.utils import ImageFolder, DatasetFolder, FilelistDataset, AvalancheDataset
# + [markdown] pycharm={"name": "#%%\n"}
# ## 🛠️ Benchmarks Basics
#
# The _Avalanche_ benchmarks \(instances of the _Scenario_ class\), contains several attributes that characterize the benchmark. However, the most important ones are the `train` and `test streams`.
#
# In _Avalanche_ we often suppose to have access to these **two parallel stream of data** \(even though some benchmarks may not provide such feature, but contain just a unique test set\).
#
# Each of these `streams` are _iterable_, _indexable_ and _sliceable_ objects that are composed of unique **experiences**. Experiences are batch of data \(or "_tasks_"\) that can be provided with or without a specific task label.
#
# #### Efficiency
#
# It is worth mentioning that all the data belonging to a _stream_ are not loaded into the RAM beforehand. Avalanche actually loads the data when a specific _mini-batches_ are requested at training/test time based on the policy defined by each `Dataset` implementation.
#
# This means that memory requirements are very low, while the speed is guaranteed by a multi-processing data loading system based on the one defined in Pytorch.
#
# #### Scenarios
#
# So, as we have seen, each `scenario` object in _Avalanche_ has several useful attributes that characterizes the benchmark, including the two important `train` and `test streams`. Let's check what you can get from a scenario object more in details:
# + pycharm={"name": "#%%\n"}
from avalanche.benchmarks.classic import SplitMNIST
split_mnist = SplitMNIST(n_experiences=5, seed=1)
# Original train/test sets
print('--- Original datasets:')
print(split_mnist.original_train_dataset)
print(split_mnist.original_test_dataset)
# A list describing which training patterns are assigned to each experience.
# Patterns are identified by their id w.r.t. the dataset found in the
# original_train_dataset field.
print('--- Train patterns assignment:')
print(split_mnist.train_exps_patterns_assignment)
# A list describing which test patterns are assigned to each experience.
# Patterns are identified by their id w.r.t. the dataset found in the
# original_test_dataset field
print('--- Test patterns assignment:')
print(split_mnist.test_exps_patterns_assignment)
# the task label of each experience.
print('--- Task labels:')
print(split_mnist.task_labels)
# train and test streams
print('--- Streams:')
print(split_mnist.train_stream)
print(split_mnist.test_stream)
# A list that, for each experience (identified by its index/ID),
# stores a set of the (optionally remapped) IDs of classes of patterns
# assigned to that experience.
print('--- Classes in each experience:')
split_mnist.classes_in_experience
# -
# #### Train and Test Streams
#
# The _train_ and _test streams_ can be used for training and testing purposes, respectively. This is what you can do with these streams:
# + pycharm={"name": "#%%\n"}
# each stream has a name: "train" or "test"
train_stream = split_mnist.train_stream
print(train_stream.name)
# we have access to the scenario from which the stream was taken
train_stream.benchmark
# we can slice and reorder the stream as we like!
substream = train_stream[0]
substream = train_stream[0:2]
substream = train_stream[0,2,1]
len(substream)
# -
# #### Experiences
#
# Each stream can in turn be treated as an iterator that produces a unique `experience`, containing all the useful data regarding a _batch_ or _task_ in the continual stream our algorithms will face. Check out how can you use these experiences below:
# + pycharm={"name": "#%%\n"}
# we get the first experience
experience = train_stream[0]
# task label and dataset are the main attributes
t_label = experience.task_label
dataset = experience.dataset
# but you can recover additional info
experience.current_experience
experience.classes_in_this_experience
experience.classes_seen_so_far
experience.previous_classes
experience.future_classes
experience.origin_stream
experience.benchmark
# As always, we can iterate over it normally or with a pytorch
# data loader.
# For instance, we can use tqdm to add a progress bar.
from tqdm import tqdm
for i, data in enumerate(tqdm(dataset)):
pass
print("\nNumber of examples:", i + 1)
print("Task Label:", t_label)
# -
# ## 🏛️ Classic Benchmarks
#
# Now that we know how our benchmarks work in general through scenarios, streams and experiences objects, in this section we are going to explore **common benchmarks** already available for you with one line of code yet flexible enough to allow proper tuning based on your needs:
# + pycharm={"name": "#%%\n"}
from avalanche.benchmarks.classic import CORe50, SplitTinyImageNet, \
SplitCIFAR10, SplitCIFAR100, SplitCIFAR110, SplitMNIST, RotatedMNIST, \
PermutedMNIST, SplitCUB200, SplitImageNet
# creating PermutedMNIST (Task-Incremental)
perm_mnist = PermutedMNIST(
n_experiences=2,
seed=1234,
)
# -
# Many of the classic benchmarks will download the original datasets they are based on automatically and put it under the `"~/.avalanche/data"` directory.
#
# ### How to Use the Benchmarks
#
# Let's see now how we can use the classic benchmark or the ones that you can create through the generators \(see next section\). For example, let's try out the classic `PermutedMNIST` benchmark \(_Task-Incremental_ scenario\).
# + pycharm={"name": "#%%\n"}
# creating the benchmark instance (scenario object)
perm_mnist = PermutedMNIST(
n_experiences=3,
seed=1234,
)
# recovering the train and test streams
train_stream = perm_mnist.train_stream
test_stream = perm_mnist.test_stream
# iterating over the train stream
for experience in train_stream:
print("Start of task ", experience.task_label)
print('Classes in this task:', experience.classes_in_this_experience)
# The current Pytorch training set can be easily recovered through the
# experience
current_training_set = experience.dataset
# ...as well as the task_label
print('Task {}'.format(experience.task_label))
print('This task contains', len(current_training_set), 'training examples')
# we can recover the corresponding test experience in the test stream
current_test_set = test_stream[experience.current_experience].dataset
print('This task contains', len(current_test_set), 'test examples')
# -
# ## 🐣 Benchmarks Generators
#
# What if we want to create a new benchmark that is not present in the "_Classic_" ones? Well, in that case _Avalanche_ offer a number of utilites that you can use to create your own benchmark with maximum flexibility: the **benchmarks generators**!
#
# ### Specific Generators
#
# The _specific_ scenario generators are useful when starting from one or multiple Pytorch datasets you want to create a "**New Instances**" or "**New Classes**" benchmark: i.e. it supports the easy and flexible creation of a _Domain-Incremental_, _Class-Incremental or Task-Incremental_ scenarios among others.
#
# For the **New Classes** scenario you can use the following function:
#
# * `nc_benchmark`
#
# for the **New Instances**:
#
# * `ni_benchmark`
# + pycharm={"name": "#%%\n"}
from avalanche.benchmarks.generators import nc_benchmark, ni_benchmark
# -
# Let's start by creating the MNIST dataset object as we would normally do in Pytorch:
# + pycharm={"name": "#%%\n"}
from torchvision.transforms import Compose, ToTensor, Normalize, RandomCrop
train_transform = Compose([
RandomCrop(28, padding=4),
ToTensor(),
Normalize((0.1307,), (0.3081,))
])
test_transform = Compose([
ToTensor(),
Normalize((0.1307,), (0.3081,))
])
mnist_train = MNIST(
'./data/mnist', train=True, download=True, transform=train_transform
)
mnist_test = MNIST(
'./data/mnist', train=False, download=True, transform=test_transform
)
# -
# Then we can, for example, create a new benchmark based on MNIST and the classic _Domain-Incremental_ scenario:
# + pycharm={"name": "#%%\n"}
scenario = ni_benchmark(
mnist_train, mnist_test, n_experiences=10, shuffle=True, seed=1234,
balance_experiences=True
)
train_stream = scenario.train_stream
for experience in train_stream:
t = experience.task_label
exp_id = experience.current_experience
training_dataset = experience.dataset
print('Task {} batch {} -> train'.format(t, exp_id))
print('This batch contains', len(training_dataset), 'patterns')
# -
# Or, we can create a benchmark based on MNIST and the _Class-Incremental_ \(what's commonly referred to as "_Split-MNIST_" benchmark\):
# + pycharm={"name": "#%%\n"}
scenario = nc_benchmark(
mnist_train, mnist_test, n_experiences=10, shuffle=True, seed=1234,
task_labels=False
)
train_stream = scenario.train_stream
for experience in train_stream:
t = experience.task_label
exp_id = experience.current_experience
training_dataset = experience.dataset
print('Task {} batch {} -> train'.format(t, exp_id))
print('This batch contains', len(training_dataset), 'patterns')
# -
# ### Generic Generators
#
# Finally, if you cannot create your ideal benchmark since it does not fit well in the aforementioned _new classes_ or _new instances_ scenarios, you can always use our **generic generators**:
#
# * **filelist\_benchmark**
# * **paths\_benchmark**
# * **dataset\_benchmark**
# * **tensors\_benchmark**
# + pycharm={"name": "#%%\n"}
from avalanche.benchmarks.generators import filelist_benchmark, dataset_benchmark, \
tensors_benchmark, paths_benchmark
# -
# Let's start with the `filelist_benchmark` utility. This function is particularly useful when it is important to preserve a particular order of the patterns to be processed \(for example if they are frames of a video\), or in general if we have data scattered around our drive and we want to create a sequence of batches/tasks providing only a txt file containing the list of their paths.
#
# For _Avalanche_ we follow the same format of the _Caffe_ filelists \("_path_ _class\_label_"\):
#
# /path/to/a/file.jpg 0
# /path/to/another/file.jpg 0
# ...
# /path/to/another/file.jpg M
# /path/to/another/file.jpg M
# ...
# /path/to/another/file.jpg N
# /path/to/another/file.jpg N
#
#
# So let's download the classic "_Cats vs Dogs_" dataset as an example:
# !wget -N --no-check-certificate \
# https://storage.googleapis.com/mledu-datasets/cats_and_dogs_filtered.zip
# !unzip -q -o cats_and_dogs_filtered.zip
# You can now see in the `content` directory on colab the image we downloaded. We are now going to create the filelists and then use the `filelist_benchmark` function to create our benchmark:
# + pycharm={"name": "#%%\n"}
import os
# let's create the filelists since we don't have it
dirpath = "cats_and_dogs_filtered/train"
for filelist, rel_dir, t_label in zip(
["train_filelist_00.txt", "train_filelist_01.txt"],
["cats", "dogs"],
[0, 1]):
# First, obtain the list of files
filenames_list = os.listdir(os.path.join(dirpath, rel_dir))
# Create the text file containing the filelist
# Filelists must be in Caffe-style, which means
# that they must define path in the format:
#
# relative_path_img1 class_label_first_img
# relative_path_img2 class_label_second_img
# ...
#
# For instance:
# cat/cat_0.png 1
# dog/dog_54.png 0
# cat/cat_3.png 1
# ...
#
# Paths are relative to a root path
# (specified when calling filelist_benchmark)
with open(filelist, "w") as wf:
for name in filenames_list:
wf.write(
"{} {}\n".format(os.path.join(rel_dir, name), t_label)
)
# Here we create a GenericCLScenario ready to be iterated
generic_scenario = filelist_benchmark(
dirpath,
["train_filelist_00.txt", "train_filelist_01.txt"],
["train_filelist_00.txt"],
task_labels=[0, 0],
complete_test_set_only=True,
train_transform=ToTensor(),
eval_transform=ToTensor()
)
# -
# In the previous cell we created a benchmark instance starting from file lists. However, `paths_benchmark` is a better choice if you already have the list of paths directly loaded in memory:
# +
train_experiences = []
for rel_dir, label in zip(
["cats", "dogs"],
[0, 1]):
# First, obtain the list of files
filenames_list = os.listdir(os.path.join(dirpath, rel_dir))
# Don't create a file list: instead, we create a list of
# paths + class labels
experience_paths = []
for name in filenames_list:
instance_tuple = (os.path.join(dirpath, rel_dir, name), label)
experience_paths.append(instance_tuple)
train_experiences.append(experience_paths)
# Here we create a GenericCLScenario ready to be iterated
generic_scenario = paths_benchmark(
train_experiences,
[train_experiences[0]], # Single test set
task_labels=[0, 0],
complete_test_set_only=True,
train_transform=ToTensor(),
eval_transform=ToTensor()
)
# -
# Let us see how we can use the `dataset_benchmark` utility, where we can use several PyTorch datasets as different batches or tasks. This utility expectes a list of datasets for the train, test (and other custom) streams. Each dataset will be used to create an experience:
# + pycharm={"name": "#%%\n"}
train_cifar10 = CIFAR10(
'./data/cifar10', train=True, download=True
)
test_cifar10 = CIFAR10(
'./data/cifar10', train=False, download=True
)
generic_scenario = dataset_benchmark(
[train_MNIST, train_cifar10],
[test_MNIST, test_cifar10]
)
# -
# Adding task labels can be achieved by wrapping each datasets using `AvalancheDataset`. Apart from task labels, `AvalancheDataset` allows for more control over transformations and offers an ever growing set of utilities (check the documentation for more details).
# +
# Alternatively, task labels can also be a list (or tensor)
# containing the task label of each pattern
train_MNIST_task0 = AvalancheDataset(train_cifar10, task_labels=0)
test_MNIST_task0 = AvalancheDataset(test_cifar10, task_labels=0)
train_cifar10_task1 = AvalancheDataset(train_cifar10, task_labels=1)
test_cifar10_task1 = AvalancheDataset(test_cifar10, task_labels=1)
scenario_custom_task_labels = dataset_benchmark(
[train_MNIST_task0, train_cifar10_task1],
[test_MNIST_task0, test_cifar10_task1]
)
print('Without custom task labels:',
generic_scenario.train_stream[1].task_label)
print('With custom task labels:',
scenario_custom_task_labels.train_stream[1].task_label)
# -
# And finally, the `tensors_benchmark` generator:
# + pycharm={"name": "#%%\n"}
pattern_shape = (3, 32, 32)
# Definition of training experiences
# Experience 1
experience_1_x = torch.zeros(100, *pattern_shape)
experience_1_y = torch.zeros(100, dtype=torch.long)
# Experience 2
experience_2_x = torch.zeros(80, *pattern_shape)
experience_2_y = torch.ones(80, dtype=torch.long)
# Test experience
# For this example we define a single test experience,
# but "tensors_benchmark" allows you to define even more than one!
test_x = torch.zeros(50, *pattern_shape)
test_y = torch.zeros(50, dtype=torch.long)
generic_scenario = tensors_benchmark(
train_tensors=[(experience_1_x, experience_1_y), (experience_2_x, experience_2_y)],
test_tensors=[(test_x, test_y)],
task_labels=[0, 0], # Task label of each train exp
complete_test_set_only=True
)
# + [markdown] pycharm={"name": "#%% md\n"}
# This completes the "_Benchmark_" tutorial for the "_From Zero to Hero_" series. We hope you enjoyed it!
#
# ## 🤝 Run it on Google Colab
#
# You can run _this chapter_ and play with it on Google Colaboratory: [](https://colab.research.google.com/github/ContinualAI/avalanche/blob/master/notebooks/from-zero-to-hero-tutorial/03_benchmarks.ipynb)
| notebooks/from-zero-to-hero-tutorial/03_benchmarks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Resources
# - https://www.cs.ubc.ca/~schmidtm/Courses/340-F15/L31.pdf
# - https://en.wikipedia.org/wiki/Spectral_clustering
# - https://en.wikipedia.org/wiki/Laplacian_matrix
# - https://towardsdatascience.com/spectral-clustering-for-beginners-d08b7d25b4d8
# - http://blog.shriphani.com/2015/04/06/the-smallest-eigenvalues-of-a-graph-laplacian/
# # Import
import numpy as np
import matplotlib.pyplot as plt
from scipy.spatial import KDTree
from scipy.stats import norm, multivariate_normal
from SimPEG.utils import mkvc
from sklearn import datasets
from sklearn.cluster import KMeans
import scipy.sparse as sp
# example dataset
n_samples = 1500
noisy_circles = datasets.make_circles(n_samples=n_samples, factor=.5,
noise=.05)
plt.scatter(noisy_circles[0][:,0],noisy_circles[0][:,1],c=noisy_circles[1],cmap='bwr')
plt.gca().set_title('TRUE CLASSIFICATION')
# # Build the Matrices
# ## Adjacency Matrix
data_subset_for_test = noisy_circles[0]
tree = KDTree(data_subset_for_test)
# find k-neighbors of each point
kneighbors = 10
knearest, kindx = tree.query(data_subset_for_test,k=kneighbors+1)
kindx = kindx[:,1:]
kindx
# build sparse adjacency matrix
A = sp.lil_matrix((data_subset_for_test.shape[0],data_subset_for_test.shape[0]))
A[:,kindx] = 1.
plt.spy(A,markersize=0.1)
# build sparse Laplacian matrix
L = sp.diags(kneighbors * np.ones(data_subset_for_test.shape[0])) - A
plt.spy(L,markersize=0.1)
# find the 2 biggest eigenvectors of the adjacency matrix
keig = 2
eigenvalues,eigenvectors = sp.linalg.eigs(A,k=keig)
eigenvalues,eigenvectors = eigenvalues.real,eigenvectors.real
plt.scatter(eigenvectors[:,0],eigenvectors[:,1])
# +
# find the 2 smallest eigenvectors of the adjacency matrix
keig = 2
eigenvalues,eigenvectors = sp.linalg.eigs(L,k=keig,which='SM')
eigenvalues,eigenvectors = eigenvalues.real,eigenvectors.real
plt.scatter(eigenvectors[:,0],eigenvectors[:,1])
# -
kmeans = KMeans(n_clusters=2, random_state=0).fit(eigenvectors)
plt.scatter(noisy_circles[0][:,0],noisy_circles[0][:,1],c=kmeans.labels_,cmap='bwr')
plt.gca().set_title('SPECTRAL CLUSTERING CLASSIFICATION PREDICTION')
# # Summarize in function
def spectral_clustering_adjacency(data,kneighbors,keigenvalues,n_clusters,tree_opts={}):
tree = KDTree(data)
# find kn-neighbors of each point
print('Building the KDTree')
knearest, kindx = tree.query(data,k=kneighbors+1,**tree_opts)
kindx = kindx[:,1:]
# build sparse adjacency matrix
print('Building the Adjacency matrix')
A = sp.lil_matrix((data.shape[0],data.shape[0]))
A[:,kindx] = 1.
# find the ke-biggest eigenvectors of the adjacency matrix
print('Computing the eigenvectors')
eigenvalues,eigenvectors = sp.linalg.eigs(A,k=keigenvalues, which='LR')
eigenvalues,eigenvectors = eigenvalues.real,eigenvectors.real
#
print('Running KMeans')
kmeans = KMeans(n_clusters=n_clusters, random_state=0).fit(eigenvectors)
return kmeans, eigenvectors
# Laplacian approach might be easier for eigenvalue thanks to the dominating diagonal
def spectral_clustering_laplacian(data,kneighbors,keigenvalues,n_clusters,tree_opts={}):
tree = KDTree(data)
# find kn-neighbors of each point
print('Building the KDTree')
knearest, kindx = tree.query(data,k=kneighbors+1,**tree_opts)
kindx = kindx[:,1:]
# build sparse adjacency matrix
print('Building the Adjacency matrix')
A = sp.lil_matrix((data.shape[0],data.shape[0]))
A[:,kindx] = 1.
L = sp.diags(kneighbors * np.ones(data.shape[0])) - A
# find the ke-biggest eigenvectors of the adjacency matrix
print('Computing the eigenvectors')
eigenvalues,eigenvectors = sp.linalg.eigs(L,k=keigenvalues, which='SR')
eigenvalues,eigenvectors = eigenvalues.real,eigenvectors.real
#
print('Running KMeans')
kmeans = KMeans(n_clusters=n_clusters, random_state=0).fit(eigenvectors)
return kmeans, eigenvectors
# ### test
spectral_cir, _ = spectral_clustering_adjacency(
noisy_circles[0],
kneighbors=10,
keigenvalues=2,
n_clusters=2,
tree_opts={'p':1}
)
plt.scatter(noisy_circles[0][:,0],noisy_circles[0][:,1],c=spectral_cir.labels_,cmap='bwr')
plt.gca().set_title('SPECTRAL CLUSTERING CLASSIFICATION PREDICTION: ADJACENT')
spectral_cir, _ = spectral_clustering_laplacian(
noisy_circles[0],
kneighbors=10,
keigenvalues=2,
n_clusters=2,
tree_opts={'p':1}
)
plt.scatter(noisy_circles[0][:,0],noisy_circles[0][:,1],c=spectral_cir.labels_,cmap='bwr')
plt.gca().set_title('SPECTRAL CLUSTERING CLASSIFICATION PREDICTION: LAPLACIAN')
# # Spectral clustering for Image Segmentation
from skimage import data as data_import
from skimage.transform import rescale
from scipy.stats import boxcox, yeojohnson
astronaut = data_import.astronaut()
# downsample for speed of tutorial
astronaut = rescale(astronaut, 1.0 / 4.0, anti_aliasing=True,multichannel=True,mode='reflect')
plt.imshow(astronaut)
# +
# built the data to train the tree
# including the coords for continuous classification is a bad idea
#x = np.linspace(0,1.,astronaut.shape[0])
#y = np.linspace(0,1.,astronaut.shape[1])
#X,Y = np.meshgrid(x,y)
#normalize the data
rgb = astronaut.reshape(-1,3)/255.
r = yeojohnson(rgb[:,0])[0]
g = yeojohnson(rgb[:,1])[0]
b = yeojohnson(rgb[:,2])[0]
image_data = np.c_[
#yeojohnson(X.flatten())[0].reshape(-1,1),
#yeojohnson(Y.flatten())[0].reshape(-1,1),
r,g,b
]
# -
spectral_ast, eigv_ast = spectral_clustering_laplacian(
rgb,
kneighbors=128,
keigenvalues=3, # dimensionality reduction
n_clusters=3,
tree_opts={'p':2}
)
# compare original space, yeojohnson space, and eigenvector space
fig, ax = plt.subplots(1,3,figsize=(15,5))
ax[0].scatter(rgb[:,0],rgb[:,1],c=spectral_ast.labels_)
ax[1].scatter(rgb[:,0],rgb[:,2],c=spectral_ast.labels_)
ax[2].scatter(rgb[:,1],rgb[:,2],c=spectral_ast.labels_)
fig.suptitle('Original space and spectral clustering result')
# compare original space, yeojohnson space, and eigenvector space
fig, ax = plt.subplots(1,3,figsize=(15,5))
ax[0].scatter(r,g,c=spectral_ast.labels_)
ax[1].scatter(r,b,c=spectral_ast.labels_)
ax[2].scatter(g,b,c=spectral_ast.labels_)
fig.suptitle('Yeojohnson space and spectral clustering result')
# compare original space, yeojohnson space, and eigenvector space
fig, ax = plt.subplots(1,3,figsize=(15,5))
ax[0].scatter(eigv_ast[:,0],eigv_ast[:,1],c=spectral_ast.labels_)
ax[1].scatter(eigv_ast[:,0],eigv_ast[:,2],c=spectral_ast.labels_)
ax[2].scatter(eigv_ast[:,1],eigv_ast[:,2],c=spectral_ast.labels_)
fig.suptitle('Eigenvector space and spectral clustering result')
# visualize the location of clusters
plt.imshow(spectral_ast.labels_.reshape(astronaut.shape[:2]),cmap='bwr')
# visualize the resulting segmented image
fig, ax = plt.subplots(1,3,figsize=(30,10))
ax = ax.flatten()
clusters_label = np.unique(spectral_ast.labels_)
for i, cls in enumerate(clusters_label):
cluster = np.zeros_like(astronaut)
indx = spectral_ast.labels_.reshape(astronaut.shape[:2]) == cls
cluster[indx] = astronaut[indx]
ax[i].imshow(cluster)
| Machine Learning - jupyter notebooks/Spectral Clustering from scratch or almost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 課程目標:
#
# 了解MLP 在神經網路上的應用
# # 範例重點:
# 以Keras 自帶的 手寫辨識的數據集來說明 MLP 建構的網路模型
# + [markdown] colab_type="text" id="zfI5zCjRPlnc"
# # 資料預處理
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 20601, "status": "ok", "timestamp": 1533441753568, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-ANnwjGu3IBk/AAAAAAAAAAI/AAAAAAAAAAc/qXXg6Jek9xw/s50-c-k-no/photo.jpg", "userId": "112554017642991017343"}, "user_tz": -480} id="GyIQ7tpGPlnf" outputId="f3642b99-3ef2-43fe-ee42-e17b1ef50cbb"
from keras.utils import np_utils
import numpy as np
np.random.seed(10)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 5937, "status": "ok", "timestamp": 1533441759521, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-ANnwjGu3IBk/AAAAAAAAAAI/AAAAAAAAAAc/qXXg6Jek9xw/s50-c-k-no/photo.jpg", "userId": "112554017642991017343"}, "user_tz": -480} id="OLzoKlJRPlno" outputId="c75f10a4-951d-4b3b-ad5f-c8fc4dff0e96"
#載入手寫辨識的資料集
from keras.datasets import mnist
(x_train_image,y_train_label),\
(x_test_image,y_test_label)= mnist.load_data()
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="QEuQwa4nPlnu"
#指定測試集與訓練資料集
x_Train =x_train_image.reshape(60000, 784).astype('float32')
x_Test = x_test_image.reshape(10000, 784).astype('float32')
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="YisRq-SxPln0"
# normalize inputs from 0-255 to 0-1
x_Train_normalize = x_Train / 255
x_Test_normalize = x_Test / 255
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="AHtfmzpfPln4"
#把LABEL轉成NUMERICAL Categorical
y_Train_OneHot = np_utils.to_categorical(y_train_label)
y_Test_OneHot = np_utils.to_categorical(y_test_label)
# + [markdown] colab_type="text" id="0s8PjvqJPln7"
# # 建立模型
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="N10v1eX2Pln8"
from keras.models import Sequential
from keras.layers import Dense
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="jmaxrsXwPln-"
#宣告採用序列模型
model = Sequential()
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="aZzGH92NPloB"
#建構輸入層
model.add(Dense(units=256,
input_dim=784,
kernel_initializer='normal',
activation='relu'))
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="NNSYvuNhPloD"
#建構輸出層
model.add(Dense(units=10,
kernel_initializer='normal',
activation='softmax'))
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 221} colab_type="code" executionInfo={"elapsed": 1500, "status": "ok", "timestamp": 1533441769991, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-ANnwjGu3IBk/AAAAAAAAAAI/AAAAAAAAAAc/qXXg6Jek9xw/s50-c-k-no/photo.jpg", "userId": "112554017642991017343"}, "user_tz": -480} id="Li6CzpyNPloF" outputId="7a9e22a4-93dc-49b0-8c8b-690e330aee88"
print(model.summary())
# + [markdown] colab_type="text" id="jdoAZgv1PloK"
# # 訓練模型
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="5d5wrEL3PloK"
model.compile(loss='categorical_crossentropy',
optimizer='adam', metrics=['accuracy'])
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 374} colab_type="code" executionInfo={"elapsed": 15426, "status": "ok", "timestamp": 1533441786624, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-ANnwjGu3IBk/AAAAAAAAAAI/AAAAAAAAAAc/qXXg6Jek9xw/s50-c-k-no/photo.jpg", "userId": "112554017642991017343"}, "user_tz": -480} id="VlpjoqQ9PloM" outputId="b91454c9-af9a-40fe-9736-b4a1c97462cd"
train_history =model.fit(x=x_Train_normalize,
y=y_Train_OneHot,validation_split=0.2,
epochs=10, batch_size=32,verbose=1)
# + [markdown] colab_type="text" id="pV1Ftdb0PloP"
# # 以圖形顯示訓練過程
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="EEmuoa1dPloQ"
import matplotlib.pyplot as plt
def show_train_history(train_history,train,validation):
plt.plot(train_history.history[train])
plt.plot(train_history.history[validation])
plt.title('Train History')
plt.ylabel(train)
plt.xlabel('Epoch')
plt.legend(['train', 'validation'], loc='upper left')
plt.show()
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 376} colab_type="code" executionInfo={"elapsed": 1451, "status": "ok", "timestamp": 1533441789030, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-ANnwjGu3IBk/AAAAAAAAAAI/AAAAAAAAAAc/qXXg6Jek9xw/s50-c-k-no/photo.jpg", "userId": "112554017642991017343"}, "user_tz": -480} id="fvE2WsejPloT" outputId="bb56e170-8850-409e-bbb2-eb6b5b89e528"
show_train_history(train_history,'acc','val_acc')
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 376} colab_type="code" executionInfo={"elapsed": 1937, "status": "ok", "timestamp": 1533441791049, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-ANnwjGu3IBk/AAAAAAAAAAI/AAAAAAAAAAc/qXXg6Jek9xw/s50-c-k-no/photo.jpg", "userId": "112554017642991017343"}, "user_tz": -480} id="zIYyrnFWPloW" outputId="4bdbd1fa-6f25-49af-88d0-8d35f6e2ecaa"
show_train_history(train_history,'loss','val_loss')
# + [markdown] colab_type="text" id="h1z2y0g-PloZ"
# # 評估模型準確率
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 68} colab_type="code" executionInfo={"elapsed": 1425, "status": "ok", "timestamp": 1533441792584, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-ANnwjGu3IBk/AAAAAAAAAAI/AAAAAAAAAAc/qXXg6Jek9xw/s50-c-k-no/photo.jpg", "userId": "112554017642991017343"}, "user_tz": -480} id="vMawurJqPloZ" outputId="31465f8d-cfad-49b8-dc9e-90d643e6b4a8"
scores = model.evaluate(x_Test_normalize, y_Test_OneHot)
print()
print('accuracy=',scores[1])
# -
| homeworks/D070/Day70-Keras_Mnist_MLP_Sample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Bruno-Messias/data_science_visagio/blob/main/Aprendizado_Nao_Supervisionado.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="2-ELSZ2gbffT"
# # <center>Métodos de Aprendizado não Supervisionado</center>
# + [markdown] id="VuX0sQnnbffX"
# <a id="recap"></a>
# ## 1. Introdução
# + [markdown] id="dD__jDwnbffY"
# Nos capítulos anteriores, exploramos conceitos iniciais de ML, numpy/pandas e uma introdução a algumas das técnicas de EDA. Neste capítulo, vamos nos concentrar em uma abordagem diferente de ML: a **Aprendizagem não Supervisionada**. Mais especificamente, vamos nos aprofundar nas principais técnicas e algoritmos utilizados para abordar este tópico, explorando as armadilhas mais comuns que esse tipo de problema traz, como implementar esse tipo de algoritmo usando Python e como avaliar e selecionar o melhor modelo para seu problema.
#
# > **Recapitulação da definição**: a principal diferença entre os tipos não supervisionado e supervisionado é que o **Aprendizado supervisionado** é feito utilizando um conhecimento prévio da variável resposta, ou em outras palavras, temos conhecimento prévio de quais devem ser os valores de saída de nossos modelos. Portanto, o objetivo da aprendizagem supervisionada é aprender uma função que, dada uma amostra de dados e saídas desejadas, melhor as correlacione. A **Aprendizagem não Supervisionada**, por outro lado, não possui saídas rotuladas, então seu objetivo é inferir a estrutura natural presente dentro de um conjunto de dados. As tarefas mais comuns na aprendizagem não supervisionada são a clusterização e a análise de associação. O aprendizado não supervisionado também é muito utilizado na análise exploratória já que é capaz de identificar agrupamentos ou similaridade entre as instâncias analisadas.
#
# Para apresentar as técnicas de **Aprendizagem não Supervisionada** mais comuns, esta aula será dividida em 3 seções. A primeira apresenta a metodologia de Análise de Associação, útil para descobrir correleações ocultas em grandes conjuntos de dados. A segunda seção apresenta a Análise de Cluster, um grupo de técnicas que o ajudará a descobrir semelhanças entre instâncias. Por fim, teremos uma última seção que tratara sobre a técnica de Soft clustering.
#
# + [markdown] id="x-aoAT3qbffZ"
# <a id="association_analysis"></a>
# ## 2. Análise de Associação
# + [markdown] id="4D0zaRqJbffZ"
# <a id="problem_definition_association"></a>
# ### 2.1. Definição do Problema
#
# Imagine a seguinte situação hipotética: você possui uma loja de varejo que vende produtos ao público em quantidades relativamente pequenas e percebeu que quase todos os clientes que compram fraldas também compram cervejas. Naturalmente, você se pergunta: _ "Nossa, que padrão estranho! Será que devo colocar os dois produtos lado a lado na prateleira ?" _. Bem, é um tipo de correlação estranha, mas imagine que você pudesse identificar padrões comuns em todos os itens vendidos por sua loja. Não seria interessante ?!
#
# Infelizmente, esta história é provavelmente uma lenda urbana de dados. No entanto, é um exemplo ilustrativo (e divertido) dos insights que podem ser obtidos pela **Análise de associação**, que tenta encontrar padrões comuns sobre itens em grandes conjuntos de dados. Esta aplicação específica é frequentemente chamada de análise de cesta de compras (mais especificamente, este é o caso do "cerveja e fraldas"), mas também pode ser aplicada a outras situações, pedido de peças de reposição e mecanismos de recomendação online - apenas para citar um pouco.
#
# Para apresentá-lo ao aprendizado de regras de associação, vamos examinar o Dataset chamado **Online Retail Data Set**, que contém todas as transações ocorridas entre 01/12/2010 e 09/12/2011 para um e-commerce.
# + colab={"base_uri": "https://localhost:8080/"} id="lAZ-nExnd3Qf" outputId="a3ad2a12-9830-4c93-89f9-ed9cb8dbdd29"
# Usando outro caminho para obter os csv do google drive
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="8-pzFsHHeGA_" outputId="c1e949c3-9f2c-47a1-87af-fff1e64b9eff"
# !ln -s /content/drive/MyDrive/Data\ Science/Entrega3 /mydrive
# !ls /mydrive
# + id="CBQYKn9jbffa" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="5474c43a-bad7-4a4b-deb9-401b985dab0d"
# Leitura do Dataset
import pandas as pd
df = pd.read_excel('/mydrive/dados/Online_Retail.xlsx')
df.head()
# + id="7EZgByxXbffe"
# É necessária uma primeira etapa de tratamento dos dados.
# Primeiramente, algumas das descrições possuem espaços que precisam ser removidos.
# Além disso, também iremos remover linhas sem informação
df['Description'] = df['Description'].str.strip()
df.dropna(axis=0, subset=['InvoiceNo'], inplace=True)
df['InvoiceNo'] = df['InvoiceNo'].astype('str')
df = df[~df['InvoiceNo'].str.contains('C')]
# + [markdown] id="xrM8cET-bffi"
# <a id="initial_analysis_association"></a>
# ### 2.2. Análises Iniciais
#
# A Análise de Associação é relativamente simples em termos matemáticos. Esta técnica é um bom começo para certos casos de exploração de dados e pode apontar o caminho para uma análise mais profunda nos dados utilizando outras abordagens.
#
# Mas antes de começarmos a modelar nosso problema, há alguns termos usados na análise de associação que são fundamentais para sua compreenção: ***Itemset, Suporte, Confiança e Lift***. Nas próximas subseções, explicamos esses termos em detalhes, com base nas seguintes transações (Exemplificadas na imagem abaixo):
#
# <img src="https://annalyzin.files.wordpress.com/2016/04/association-rule-support-table.png?w=376&h=334" width="350">
#
#
# #### 2.2.1 Itemset
# A Análise de Associação tenta identificar associações frequentes "se-então" chamadas regras de associação, que consistem em um antecedente (se) e um consequente (então). Para uma determinada regra, chamamos ***Itemset*** a lista de todos os itens no antecedente e no consequente. Por exemplo: “Se maçã e cerveja, arroz” (“Se maçã e cerveja forem comprados, então há uma grande chance de que arroz também seja comprado pelo cliente”). Nesse caso, maçã e cerveja são o antecedente e o arroz é o consequente.
#
# #### 2.2.2 Suporte
#
# Isso mostra a popularidade de um conjunto de itens, medido pela proporção de transações nas quais um conjunto de itens aparece. Na tabela mostrada acima, o suporte de {Maçã} é 4 de 8, ou 50%. Os conjuntos de itens também podem conter vários itens. Por exemplo, o suporte de {maçã, cerveja, arroz} é 2 de 8, ou 25%.
#
# 
#
# Se você descobrir que as vendas de itens além de uma determinada proporção tendem a ter um impacto significativo em seus lucros, você pode considerar usar essa proporção como seu limite de suporte.
#
# #### 2.2.3 Confiança
#
# Isso indica a probabilidade de compra do item Y quando o item X é comprado, expressa como {X -> Y}. Isso é medido pela proporção de transações com o item X, em que o item Y também aparece. Na Tabela 1, a confiança de {Maçã -> Cerveja} é 3 de 4, ou 75%.
#
# 
#
# Uma desvantagem da medida de confiança é que ela pode representar mal a importância de uma associação. Isso ocorre porque ela só explica a popularidade das maçãs, mas não das cervejas. Se as cervejas também forem muito populares em geral, haverá uma chance maior de que uma transação contendo maçãs também contenha cervejas, aumentando assim a medida de confiança.
#
# #### 2.2.4 Lift
#
# Isso diz a probabilidade de o item Y ser comprado quando o item X é comprado, enquanto controla a popularidade do item Y. Na tabela mostrada acima, o Lift de {maçã -> cerveja} é 1, o que não implica nenhuma associação entre os itens. Um valor de Lift maior que 1 significa que o item Y provavelmente será comprado se o item X for comprado, enquanto um valor inferior a 1 significa que o item Y provavelmente não será comprado se o item X for comprado.
#
# 
#
# <a id="modeling_association"></a>
# ### 2.3. Modelagem
#
# #### 2.3.1 Apriori
#
# Apriori é um algoritmo popular para extrair conjuntos de itens frequentes com aplicações no aprendizado de regras de associação. Para grandes conjuntos de dados, pode haver centenas de itens em centenas de milhares de transações. O algoritmo a priori tenta extrair regras para cada combinação possível de itens. Por exemplo, o Lift pode ser calculado para o item 1 e item 2, item 1 e item 3, item 1 e item 4 e, em seguida, item 2 e item 3, item 2 e item 4 e, em seguida, combinações de itens, por exemplo item 1, item 2 e item 3; da mesma forma, item 1, item2 e item 4 e assim por diante.
#
# O algoritmo a priori foi projetado para operar em bancos de dados contendo transações, como compras de clientes de uma loja. Um conjunto de itens é considerado "frequente" se atender a um limite de suporte especificado pelo usuário. Por exemplo, se o limite de suporte for definido como 0,5 (50%), um conjunto de itens frequente é definido como um conjunto de itens que ocorrem juntos em pelo menos 50% de todas as transações no banco de dados.
#
# Para aplicar o algoritmo Apriori, utilizaremos a implementação python em [MLxtend](http://rasbt.github.io/mlxtend/user_guide/frequent_patterns/apriori/).
# + id="U2tHWDtCbffk"
# Caso não possua, você deve instalar a biblioteca mlxtend
from mlxtend.frequent_patterns import apriori
from mlxtend.frequent_patterns import association_rules
# + [markdown] id="Cue4Blllbffq"
# A função implementada de MLxtend espera dados em um DataFrame pandas codificado no formato one-hot encoding. Isso significa que os itens de dados devem ser consolidados em uma transação por linha. Isso pode ser feito manualmente como ilustrado abaixo.
# + id="2ONzwfRhbffr" colab={"base_uri": "https://localhost:8080/", "height": 541} outputId="c3e7e5c8-6ecf-4e7c-830d-17c866b1b564"
# Consolide os itens em 1 transação por linha.
# Para manter o conjunto de dados pequeno, analisaremos apenas as vendas para a França.
# obs.: uma outra maneira de fazer isto é usando o método pivot_table()
basket = (df[df['Country'] =="France"]
.groupby(['InvoiceNo', 'Description'])['Quantity']
.sum().unstack().reset_index().fillna(0)
.set_index('InvoiceNo'))
basket.head(10)
# + [markdown] id="keBg14Ssbffu"
# Além disso, o algoritmo apriori só aceita números inteiros. Precisamos substituir todos os valores ≥1 por 1 e <1 por 0.
# + id="k_cez62ubffu" colab={"base_uri": "https://localhost:8080/", "height": 541} outputId="fb58ff1a-7aeb-4920-eb21-6fe3618c4b9e"
# Certifique-se de que todos os valores positivos sejam convertidos em 1 e qualquer valor menor que 0 seja definido como 0
def encode_units(x):
if x <= 0:
return 0
if x >= 1:
return 1
basket_sets = basket.applymap(encode_units)
basket_sets.drop('POSTAGE', inplace=True, axis=1)
basket_sets.head(10)
# + [markdown] id="BMOJcyVGbffy"
# Agora que os dados estão estruturados corretamente, podemos gerar conjuntos de itens frequentes que têm um suporte de pelo menos 7% (esse número foi escolhido arbitrariamente).
# + id="8Gg0Aq03bff2"
frequent_itemsets = apriori(basket_sets, min_support=0.07, use_colnames=True)
# + [markdown] id="Pp7ZZNb2bff7"
# Finalmente, podemos gerar as regras com seu suporte, confiança e lift correspondentes:
# + id="bVFWCQ-Obff8" colab={"base_uri": "https://localhost:8080/", "height": 855} outputId="5897fba1-14fb-4390-c126-d76594890fdb"
rules = association_rules(frequent_itemsets, metric="lift", min_threshold=1)
rules
# + [markdown] id="mNn07Kk_bfgB"
# Bem, isso é tudo que há para fazer! Acabamos de construir os itens frequentes usando apriori e, em seguida, construir as regras com association_rules. Mas agora, a parte complicada é descobrir o que isso nos diz. Por exemplo, podemos ver que existem algumas regras com um alto Lift, o que significa que ocorre com mais frequência do que seria esperado, dado o número de combinações de transações e produtos.
#
# ### Exercicio 1
#
# Use a célula abaixo para verificar as regras com aumento acima de 6 e confiança acima de 0,6. Que conclusões você consegue obter? Discuta
# + id="O_IePJPubfgB" colab={"base_uri": "https://localhost:8080/", "height": 762} outputId="ec268645-4279-4eca-df3d-0d07da6b8b57"
rules[(rules.confidence > 0.6)]
# Analizando os valores obtidos podmemos ver que objetos de papelaria costuman ser compradas muitas vezes em conjunto variando apenas as cores e os temas
# + [markdown] id="T4kWt5gjbfgG"
# Além disso, não seria interessante ver como as combinações variam de acordo com o país de compra? Use a célula abaixo para verificar algumas combinações populares na Alemanha. Você consegue obter algum insight ?
# + id="UUUTdcOQbfgH" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="8f85ada7-9300-47b3-f2ec-a33bb6c5b9d6"
basket = (df[df['Country'] =="Germany"]
.groupby(['InvoiceNo', 'Description'])['Quantity']
.sum().unstack().reset_index().fillna(0)
.set_index('InvoiceNo'))
basket_sets = basket.applymap(encode_units)
basket_sets.drop('POSTAGE', inplace=True, axis=1)
frequent_itemsets = apriori(basket_sets, min_support=0.07, use_colnames=True)
rules = association_rules(frequent_itemsets, metric="lift", min_threshold=1)
rules
#Analisando o obtido vemos que os suporte médio dos produtos na Alemanha parecem serem bem baixos, indicando que os clientes nesse paises diversificam bastante a compra de produtos.
# + [markdown] id="ZeIMpS85bfgL"
# <a id="clustering_analysis"></a>
# ## 3. Análise de Clusterização
# + [markdown] id="WVwo7o4dbfgM"
# Na seção anterior, apresentamos a metodologia de Análise de Associação, que é um dos métodos de Aprendizagem não Supervisionados mais comuns. Agora vamos apresentar a você outra técnica extremamente usada: a **Análise de Clusterização**.
# + [markdown] id="erCN2_ngbfgM"
# <a id="problem_definition_clustering"></a>
# ### 3.1. Definição do Problema
#
# Suponha que você esteja agora na seguinte situação hipotética: você ainda possui uma loja de varejo e depois de identificar padrões comuns de itens vendidos por sua loja, agora você gostaria de atingir grupos específicos de clientes com campanhas publicitárias específicas.
#
# Para fazer isso, você se pergunta: _"Existe alguma maneira de identificar quais são os diferentes tipos de clientes que compram em minha loja, por exemplo, considerando as características de meus clientes, como histórico de compras, interesses ou monitoramento de atividades padrões?"_. Bem, neste caso, a Analise de Clusterização poderia definitivamente ajudá-lo a responder a esta pergunta.
#
# 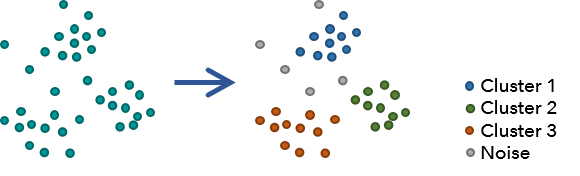
#
# De maneira simplificada, o objetivo da Clusterização é encontrar grupos diferentes dentro dos dados. Para fazer isso, os algoritmos de clusterização encontram a estrutura nos dados de forma que os elementos do mesmo cluster (ou grupo) sejam mais semelhantes uns aos outros do que aos de diferentes clusters.
#
# Dado um conjunto de pontos de dados, podemos usar um algoritmo de agrupamento para classificar cada ponto de dados em um grupo específico. Em teoria, os pontos de dados que estão no mesmo grupo devem ter propriedades e/ou features semelhantes, enquanto os pontos de dados em grupos diferentes devem ter propriedades e/ou features consideravelmente diferentes.
#
# Nas próximas subseções, discutiremos e implementaremos alguns algoritmos de agrupamento. No entanto, diferentemente do que fizemos na parte de Análise de associação, não trabalharemos em um único conjunto de dados. Em vez disso, para cada algoritmo apresentado, trabalharemos em um conjunto de dados específico.
# + [markdown] id="cy8Si84obfgN"
# <a id="initial_analysis_clustering"></a>
# ### 3.2. Análises Iniciais
#
# A Clusterização é utilizada para determinar o agrupamento intrínseco entre os dados não rotulados presentes. No entanto, "não há critérios" claros para analisar um bom agrupamento. Naturalmente, pode-se utilizar critérios (como veremos a frente) relacionados a inércia dos clusters, porém, isso não avalia se, de fato, conseguimos fazer separações que façam sentido do ponto de vista prático ou de negócios. Portanto, cabe ao usuário determinar quais são os critérios que ele pode usar para atender às suas necessidades.
# Por exemplo, podemos estar interessados em encontrar representantes para grupos homogêneos (data reduction), em encontrar "clusters naturais" e descrever suas propriedades desconhecidas, em encontrar agrupamentos úteis e adequados ou na localização de amostras incomuns (detecção de outlier). Este algoritmo deve fazer algumas suposições que constituem a similaridade de pontos e cada suposição faz clusters diferentes e igualmente válidos.
#
# Antes de começarmos a modelar, há dois conceitos importantes que precisamos abordar. Estamos falando sobre ***Determinar o número de clusters*** (que deve ser feito antes de rodar o algoritmo) e ***Feature Selection*** (ou seleção de features/variáveis).
#
# #### 3.2.1 Determinando o número de clusters
#
# Determinar o número ideal de clusters em um conjunto de dados é uma questão fundamental no processo. Infelizmente, não há uma resposta definitiva para essa pergunta. O número ideal de clusters é de alguma forma subjetiva e depende do método usado para medir semelhanças e dos parâmetros usados para particionamento. Por exemplo, se você deseja segmentar clientes que têm maior probabilidade de comprar cervejas e clientes que provavelmente não compram cervejas, pode definir um número de dois grupos. Ou, ainda, o número de clusters pode ser definido anteriormente através de regras ou restrições de negócio.
#
# No entanto, se você não tem ideia de quantos clusters precisa, pode usar alguns métodos para determinar o número ótimo.
#
# #### 3.2.2 Feature Selection
#
# Consiste em criar um subconjunto de uma lista de features/variáveis úteis entre todo o conjunto de variáveis à nossa disposição. Esta etapa pode parecer contra-intuitiva, uma vez que estamos excluindo informações que nosso modelo futuro poderia aprender, mas, se feito da maneira certa, a seleção de features pode até ser capaz de melhorar o desempenho do modelo.
#
# Um dos métodos estatísticos mais comuns utilizados para lidar com feature selection é o que chamamos de ***Análise de Componentes Principais (PCA)***. Imagine que a dimensionalidade do conjunto de features é maior do que apenas dois ou três. Usando o PCA, podemos agora identificar quais são as dimensões mais importantes e apenas manter algumas delas para explicar a maior parte da variação que vemos em nossos dados.
#
# Além disso, o PCA pode ser realmente útil para visualização e compressão de dados. Os dados nem sempre vêm com dimensionalidade igual ou menor a 3 (ou seja, 3 variáveis / features). Portanto, não podemos conceber uma visualização do gráfico de dispersão de nossos dados, uma vez que estamos limitados a apenas 3 dimensões. Isso torna impossível para nós ver sua distribuição conjunta neste espaço N-dimensional. Mas, usando o PCA, podemos contornar esse problema retendo apenas as dimensões mais úteis (ou seja, aquelas que explicam a maior parte da variação que vemos em nossos dados). No entanto, essas dimensões não correspondem às nossas originais. O PCA tenta encontrar um sistema de coordenadas neste espaço N-dimensional que maximize a variância ao longo de cada eixo.
#
# Não entraremos em mais detalhes, mas o PCA também é uma técnica de aprendizado não supervisionado. Fica a sugestão de pesquisa!
# + [markdown] id="_I5SS9PObfgN"
# <a id="modeling_clustering"></a>
# ### 3.3. Modelagem
# + [markdown] id="A48emC9bbfgO"
# ### Kmeans
# + [markdown] id="WuAH3bP3bfgO"
# O algoritmo K-means foi proposto como uma forma de **agrupar pontos de dados semelhantes em clusters**. Como veremos a frente, o algoritmo k-means é extremamente fácil de implementar e também é computacionalmente muito eficiente em comparação com outros algoritmos de agrupamento, o que pode explicar sua popularidade.
#
# Este algoritmo pertence à categoria de **prototype-based clustering**. Isso significa que cada cluster é representado por um protótipo, que pode ser o **centróide (média)** de pontos semelhantes. Embora k-means seja muito bom para identificar grupos de forma esférica, uma das desvantagens deste algoritmo de agrupamento é que temos que especificar o número de clusters k a priori.
#
# Uma escolha inadequada para k pode resultar em agrupamento de mal desempenho. Além disso, discutiremos o **método do cotovelo e a silhueta**, que são técnicas úteis para avaliar a qualidade de um agrupamento para nos ajudar determinar o número ideal de clusters k.
#
# + [markdown] id="Zut4Be4ZbfgP"
# #### Algoritmo
# + [markdown] id="_THljbN2bfgQ"
# O algoritmo k-means pode ser resumido pelas quatro etapas a seguir:
#
# 1. Escolha aleatoriamente k centróides dos pontos da amostra como centros iniciais do cluster.
# 2. Atribua cada amostra ao centroide mais próximo
# 3. Mova os centróides para o centro das amostras que foram atribuídas a ele.
# 4. Repita as etapas 2 e 3 até que as atribuições do cluster não mudem ou uma tolerância definida pelo usuário ou um número máximo de iterações seja alcançado.
#
# 
# + [markdown] id="8pXGD8tabfgR"
# Podemos definir semelhança como o oposto de distância. Uma fórmula comumente utilizada para avaliar distância em agrupamento de
# amostras com features contínuas é a distância euclidiana entre dois pontos x e y em um espaço m-dimensional:
# + [markdown] id="8cZHjLm-bfgS"
# \begin{equation*}
# d(u,v) = \sqrt{\sum_{j=1}^{m}(u_{j} - v_{j})^2} = \left \|u_{j} - v_{j} \right \|
# \end{equation*}
# + [markdown] id="yX6Je1pbbfgT"
# Observe que, na equação anterior, o índice j se refere à j-ésima dimensão (coluna de característica) dos pontos de amostra u e v.
# + [markdown] id="hEGSOQ7dbfgV"
# Com base nesta métrica de distância euclidiana, podemos descrever o algoritmo de k-means como um problema de otimização simples: uma abordagem iterativa para minimizar **a soma dos erros quadráticos (SSE) dentro do cluster**, que às vezes também é chamada de **inércia do cluster**:
# + [markdown] id="R9l1ACg5bfgW"
# \begin{equation*}
# SSE = \sum_{i=1}^{n} \sum_{j=1}^{k} w_{(i,j)} \left \|x_{i} - \mu_{j} \right \|
# \end{equation*}
# + [markdown] id="WqAsG4ulbfgX"
# Aqui, $\mu_ {j}$ é o ponto representativo (centróide) para o cluster j, <br>
# $w_ {i, j}=1$ se a amostra $x_ {i}$ está no cluster j, $w_ {i, j} = 0$ caso contrário
# + [markdown] id="taUxf-9VbfgZ"
# ##### Exemplo
# + [markdown] id="IfXQjdaebfga"
# Por questão de simplicidade, vamos criar um conjunto de dados de cluster bem definido, usando o método ```blob``` da biblioteca sklearn.
# + id="aCRUA6tSbfgc" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="75f4bd0c-6f53-44b4-914e-8d3de28ce092"
# %matplotlib inline
# Importando make blobs e matplotlib
from sklearn.datasets import make_blobs
import matplotlib.pyplot as plt
# Criando blobs (dados aleatórios em torno de centros definidos)
X,y = make_blobs(n_samples=150,
n_features=2,
centers=3,
cluster_std=0.4,
shuffle=True,
random_state=0)
# Plotando os blobs
plt.scatter(X[:,0],
X[:,1],
c='black',
marker='o',
s=50)
plt.grid()
plt.show()
# + [markdown] id="Vf5is50Gbfgf"
# Podemos usar KMeans do sklearn para realizar clusterização entre nosso conjunto de dados construído
# + id="8vI2QmCDbfgf"
# Importando o KMeans
from sklearn.cluster import KMeans
# Criando o objeto KMeans
# Observe que já sabemos a priori quantos clusters precisaremos
num_clusters = 3
km = KMeans(n_clusters=num_clusters)
# Performando clusterização K-means
cluster_km = km.fit_predict(X)
# + id="v5qRWpwpbfgj" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="dd657e35-8172-451e-ea69-823752ce3cf2"
# Plotando os dados com cores para o cluster
for cluster in range(num_clusters):
plt.scatter(X[cluster_km==cluster,0],
X[cluster_km==cluster,1],
s=50,
cmap='Pastel1',
label='cluster {}'.format(cluster))
# Plotando o centro dos CLusters
plt.scatter(km.cluster_centers_[:,0],
km.cluster_centers_[:,1],
s=250,
c='black',
marker='*',
label='centroid')
plt.legend()
plt.grid()
plt.show()
# + [markdown] id="RqWZkRvdbfgq"
# Embora k-means funcione bem neste conjunto de dados, precisamos ressaltar alguns dos
# principais desafios do k-means. Uma das desvantagens do k-means é que temos que
# especificar o número de clusters k a priori, o que pode nem sempre ser tão óbvio em
# aplicações do mundo real, especialmente se estivermos trabalhando com uma dimensão mais elevada do
# conjunto de dados que não pode ser visualizado. As outras propriedades do k-means são que os clusters
# não se sobrepõem e não são hierárquicos, e também assumimos que há pelo menos
# um item em cada cluster.
# + [markdown] id="yhI8Fgqobfgr"
# #### Métodos de Validação - Curva do cotovelo
# + [markdown] id="tFungrSmbfgr"
# A fim de quantificar a **qualidade da clusterização**, precisamos usar métricas intrínsecas, como o SSE dentro do cluster (distorção) que discutimos anteriormente neste capítulo - para comparar o desempenho de diferentes agrupamentos k-means.
# Convenientemente, não precisamos calcular o SSE dentro do cluster explicitamente, pois é já acessível através do atributo ``inertia`` após fazer o fit de um modelo KMeans:
# + id="wkjgxe2Dbfgs" colab={"base_uri": "https://localhost:8080/"} outputId="638450d5-90ee-49d4-ee3d-990c0bebaf4f"
print('Distorção: %.2f' % km.inertia_)
# + [markdown] id="celpx3CZbfgv"
# Com base no SSE dentro do cluster, podemos usar uma ferramenta gráfica, o chamado método do cotovelo, para estimar o número ótimo de clusters k para uma determinada tarefa. Intuitivamente,
# podemos dizer que, se k aumentar, a distorção diminuirá. Isso ocorre porque as amostras estarão mais próximas dos centróides aos quais estão atribuídas. A ideia por trás do
# método do cotovelo é identificar o valor de k onde a distorção começa a aumentar
# mais rapidamente, o que ficará mais claro se traçarmos a distorção para diferentes
# valores de k:
# + id="QfnZjRqHbfgw" colab={"base_uri": "https://localhost:8080/", "height": 334} outputId="c06258f6-f7f7-4e50-8de8-fc2312462b59"
# Criando lista vazia
distortions = []
# Cálculo da distorção para uma série de valores de k
for i in range(1, 11):
km = KMeans(n_clusters=i)
km.fit(X)
distortions.append(km.inertia_)
# Gráfico da Distorção
plt.figure(figsize=(8,5))
plt.plot(range(1,11), distortions, marker='o')
plt.xticks(range(1,11))
plt.xlabel('Número de Clusters')
plt.ylabel('Distorção')
plt.show()
# + [markdown] id="4RWgAe2vbfgy"
# Como podemos ver no gráfico a seguir, o cotovelo está localizado em k = 3, o que fornece
# evidência de que k = 3 é de fato uma boa escolha para este conjunto de dados.
# + [markdown] id="JYHj_dhPbfgz"
# #### Coeficiente de Silhueta
# + [markdown] id="wASoTC9Dbfgz"
# A análise da silhueta pode ser usada para medir a coesão do cluster entre os pontos de dados e o centróide. Para o cálculo, tem-se o passo a passo:
# 1. Calcule a coesão do cluster $a_ {i}$ como a distância média entre uma amostra $x_ {i}$ e todos os outros pontos no mesmo cluster.
# 2. Calcule a separação de cluster $b_ {i}$ do próximo cluster mais próximo como a distância média entre a amostra $x_ {i}$ e todas as amostras no cluster mais próximo.
# 3. Calcule a silhueta $s_ {i}$ como a diferença entre a coesão e a separação do cluster dividida pelo maior dos dois, conforme mostrado aqui:
#
# \begin{equation*}
# s_{i} = \frac{b_{i} - a_{i}}{max\left \{ b_{i},a_{i} \right \}}
# \end{equation*}
# + [markdown] id="vx10R2wwbfg0"
# O coeficiente de silhueta é limitado no intervalo de -1 a 1. Com base na fórmula anterior, podemos ver que o coeficiente de silhueta é 0 se a separação e coesão do cluster forem iguais ($ b_ {i} = a_ {i} $). Além disso, chegamos perto de um coeficiente de silhueta ideal de 1 se ($ b_ {i} >> a_ {i} $), uma vez que $ b_ {i} $ quantifica quão diferente é uma amostra de outros clusters, e $ a_ {i } $ nos diz o quão semelhante são as outras amostras em seu próprio cluster.
# + [markdown] id="bQv8ZB5Cbfg0"
# O coeficiente de silhueta está disponível como ```silhouette_score``` no módulo ```sklearn.metrics```. Isso calcula o coeficiente de silhueta médio em todas as amostras, que é equivalente a numpy.mean (silhouette_samples (…)).
# + [markdown] id="9AnhF5-Fbfg2"
# Para melhor ilustrar esta importante métrica, você pode executar as 2 células a seguir e alterar a dispersão dos dados no widget ```cluster_cohesion```. Não se preocupe se você não entender parte do código que usamos aqui para fazer o plot.
# + id="fCOlpyR1bfg4"
# Importando Python widgets para construit o plot interativo
# Execute o comando comentado abaixo, caso tenha problema no import
# # !jupyter nbextension enable --py widgetsnbextension
import ipywidgets as widgets
from ipywidgets import interact, interact_manual
# + id="xC6l52O0bfg8" colab={"base_uri": "https://localhost:8080/", "height": 401, "referenced_widgets": ["684c5157f79c4aad920d8fd77cf574ea", "dc4ed43068b14532bff8cae115518deb", "ddb67caf2c1d41ee9a043e309a8b2899", "130d8c105104436293108a799fc1672f", "1cf1baebbe9a4d8ca3e3c7c27567a0ef", "8450a0e0bbb44f8a88986634d505e195", "afeeb9ace93c4f89a77686c0fa7f8b72", "037826ccabdb4b8f91faea0bbc1636d6", "df6977f2796b4044a972fbf9ff1e6e6b", "c976eb6b42e5498b8cfd2b7615591c59"]} outputId="0b3de81b-5de6-40dd-b0f6-059317630b73"
# Importando silhouette_score
from sklearn.metrics import silhouette_score
# Declarando esta função para ser iteraiva
@interact
def calculate_kmeans(Dispersao=(0.1,0.8,0.1), n_cluster=(2,5,1)):
# Criando os blobs
X,y = make_blobs(n_samples=500,
n_features=2,
centers=3,
cluster_std=Dispersao,
shuffle=True,
random_state=0)
km = KMeans(n_clusters=n_cluster)
# Clusterização K-Means
cluster_km = km.fit_predict(X)
score = silhouette_score(X,
km.labels_,
metric='euclidean')
print("SSE: {}, Coeficiente de Silhueta: {}".format(km.inertia_,score))
# Plotando os CLusters
plt.figure(figsize = (8,5))
for cluster in range(n_cluster):
plt.scatter(X[cluster_km==cluster,0],
X[cluster_km==cluster,1],
s=50,
cmap='Pastel1',
marker='s',
label='cluster {}'.format(cluster))
# Plotando o centro dos clusters
plt.scatter(km.cluster_centers_[:,0],
km.cluster_centers_[:,1],
s=250,
c='black',
marker='*',
label='centroid')
plt.legend()
plt.grid()
plt.show()
# + [markdown] id="WGiEOX2Kbfg_"
# Observe que, quando a dispersão dos dados é pequena (clusters são mais coesos), a pontuação da silhueta fica mais próxima de 1. <br>
# E, mesmo se a dispersão for 0.1, mas o número do cluster não for 3, a pontuação sai de 1. <br>
#
# Tente variar o número de clusters para cima e para baixo no widget e veja o que ocorre (não há a necessidade de alterar o código).
# <br>
# <br>
# Portanto, este experimento mostra a importância de equilibrar o número correto de clusters usando algumas métricas de avaliação e como essa escolha afeta o desempenho do modelo.
# + [markdown] id="jqhf7_YHbfg_"
# ### Exercicio 2
#
# Agora que você aprendeu os conceitos de agrupamento Kmeans, vamos tentar usá-lo com nosso conjunto de dados de varejo para agrupar nossos clientes. Primeiro, precisamos tratar um pouco os dados e criar algumas features relacionadas ao cliente para que possamos utilizá-las com o algoritmo. Como não é o foco desta lição, não comentaremos muito sobre esta primeira parte.
# + id="3Urc6Nn3bfhA"
# Criando preço por produto
df['Price'] = df.Quantity * df.UnitPrice
# Consolidando os itens em 1 cliente por linha.
product_features = (df.groupby(['CustomerID', 'Description'])['Quantity']
.sum().unstack().reset_index().fillna(0)
.set_index('CustomerID'))
# Certifique-se de que todos os valores positivos sejam convertidos em 1 e qualquer valor menor que 0 seja definido como 0
product_presence = product_features.applymap(encode_units)
# Criando uma coluna para quantidade total e número de produtos
product_features["total"] = product_features.sum(axis = 1, skipna = True)
product_presence["total"] = product_presence.sum(axis = 1, skipna = True)
# + id="9pH5YzaGbfhF" colab={"base_uri": "https://localhost:8080/", "height": 444} outputId="f10b7240-1c98-4bbb-8cb0-04f4363eb283"
# Obtendo apenas os produtos mais comuns para reduzir o número de features
number_of_products = 10
most_common_products = df.Description.value_counts(sort=True)[0:number_of_products].index.tolist()
most_common_products.append("total")
product_features_filtered = product_features[most_common_products].add_suffix("_quantidade")
product_presence_filtered = product_presence[most_common_products].add_suffix("_numero")
product_features_filtered.head(10)
# + id="kDc1gr7nbfhL" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="2d94b6dc-40e5-4166-80a2-fa913036abb8"
# Calculando o numero de invoices por cliente
num_invoices = df.groupby('CustomerID').InvoiceNo.nunique().to_frame()
num_invoices.head()
# + id="gCupHnVdbfhP" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="eb69ffd9-a926-4502-927c-e756dede68ae"
# Calculando o tempo desde o ultimo invoice
import numpy as np
last_invoice = df.groupby('CustomerID').InvoiceDate.max().to_frame()
last_invoice['time_from_last'] = (pd.to_datetime('today') - last_invoice['InvoiceDate']) / np.timedelta64(1,'D')
last_invoice.head()
# + id="nxQtBKeSbfhY" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="602bdd9c-d5f4-4879-ba78-75c0a649c6c1"
# Calculando a média de preços
invoicePrice = df.groupby(['CustomerID', 'InvoiceNo']).Price.sum().to_frame()
averagePrice = invoicePrice.groupby('CustomerID').Price.mean().to_frame()
averagePrice.head()
# + id="srnphzxDbfhb" colab={"base_uri": "https://localhost:8080/", "height": 306} outputId="eb47eebd-18c2-4199-9501-668f0f533abb"
# Consolidando variaveis
final_df = pd.merge(product_features_filtered, product_presence_filtered, on = 'CustomerID')
final_df = pd.merge(final_df, num_invoices, on = 'CustomerID')
final_df = pd.merge(final_df, last_invoice.drop(columns=['InvoiceDate']), on = 'CustomerID')
final_df = pd.merge(final_df, averagePrice, on = 'CustomerID')
final_df.head()
# + id="mxpZF3Dxbfhe"
from scipy import stats
# Remoção de Outliers
final_df_no_outliers = final_df[(np.abs(stats.zscore(final_df)) < 3).all(axis=1)]
# + id="ICHgVtu9bfhj" colab={"base_uri": "https://localhost:8080/", "height": 306} outputId="7c061aa2-7b5a-437e-a220-1f906c2ab3a8"
# Scale para normalização
from sklearn.preprocessing import MinMaxScaler
# Iniciando scaler
scaler = MinMaxScaler()
# Aplicando scaler
final_df_no_outliers_scaled = pd.DataFrame(scaler.fit_transform(final_df_no_outliers))
# Mudando nome das colunas
final_df_no_outliers_scaled.columns = final_df_no_outliers.columns
final_df_no_outliers_scaled['CustomerID'] = final_df_no_outliers.index.tolist()
final_df_no_outliers_scaled.set_index('CustomerID', inplace = True)
final_df_no_outliers_scaled.head()
# + id="6X4jGjxYbfhn"
# Criação do Dataset final
customers_df = final_df_no_outliers_scaled.copy()
# + [markdown] id="disoDiKCbfhs"
# Agora que temos nosso conjunto de dados final ```customers_df``` criado, podemos plotar a distorção pelo número de clusters para escolher o melhor k.
# + id="IuVoulmpbfht" colab={"base_uri": "https://localhost:8080/", "height": 334} outputId="8c2cedfb-30e2-4471-d542-d6349e3d3e6d"
distortions_customers = []
max_clusters = 50
for i in range(1, max_clusters):
km_customers = KMeans(n_clusters=i)
km_customers.fit(customers_df)
distortions_customers.append(km_customers.inertia_)
plt.figure(figsize=(8,5))
plt.plot(range(1,max_clusters), distortions_customers, marker='o')
plt.xticks(range(1,max_clusters,max_clusters//10))
plt.xlabel('Number of clusters')
plt.ylabel('Distortion')
plt.show()
# + [markdown] id="ak9t8kQAbfhw"
# Agora, usando o gráfico da célula anterior, escolha o melhor número de clusters para usar em nosso modelo final abaixo. Como agora temos um problema real, tente usar não apenas a regra do cotovelo, mas também algum conhecimento de negócios para decidir o número de clusters. Por exemplo, devemos ter 50 clusters diferentes? Isso ajudaria em nossa operação?
# + id="INka-slTbfhx" colab={"base_uri": "https://localhost:8080/", "height": 351} outputId="2d30fdc6-0abf-4642-a127-9f1fd00115f3"
from sklearn.cluster import KMeans
#Criaçao do Objeto
num_clusters = ___
km_customers = ___
#Realizando Clusterização
cluster_km_customers = km_customers.fit_predict(customers_df)
#Plotando distorção
print('Distortion: %.2f' % km_customers.inertia_)
# + [markdown] id="ELCFIjbzbfh1"
# Como temos muitos dados com muitas dimensões (features), vamos apenas traçar um histograma do número de clientes por cluster.
# + id="zleWrlpcbfh2" colab={"base_uri": "https://localhost:8080/", "height": 198} outputId="1843ada9-c2cb-4824-9796-4f5b42cdcdee"
# Quantidade de Clientes por Cluster
plt.hist(x=cluster_km_customers, bins=num_clusters)
plt.show()
# + [markdown] id="G5lKHsMfaUv6"
# Agora que os clusters já estão montados, tente interpreta-los:
# * Quais são as peculiaridades de cada cluster ?
# * Qual é a caracteristica mais forte de cada um ?
#
# Sinta-se a vontade para revisitar a aula de EDA caso necessite de ajuda para montar as análises
# + [markdown] id="0ev_w8HIbfh7"
# <a id="modeling_clustering"></a>
# ### 3.4. Clusterização Hierárquica
# + [markdown] id="xNKc31sFbfh8"
# Vamos dar uma olhada em uma abordagem alternativa para clusterização: **Clusterização hierárquica**.
#
# Esta técnica é uma altenativa a anterior já que possui um mecanismo diferente para a montagem dos clusters. Esta técnica baseia-se na união das amostras para a montagem do cluster, ou na divisão sequencial do conjunto de todas as amostras para a formação dos clusters (explicaremos melhor abaixo).
#
# Uma das vantagens desta técnica é que nos permite traçar dendrogramas (visualizações de um agrupamento hierárquico binário), o que pode ajudar na interpretação dos resultados já que nos permite entender o processo de formação/divisão dos clusters.
# Outra vantagem útil dessa abordagem hierárquica é que não precisamos especificar o número de clusters antecipadamente.
# + [markdown] id="21qOBFMEbfh9"
# As duas principais abordagens para agrupamento hierárquico são o **aglomerativo e o divisivo**:
#
# * **Divisivo**: começa com um cluster que abrange todas as nossas amostras e divide iterativamente o cluster em clusters menores até que cada um contenha apenas uma amostra.
#
# * **Aglomerativo**: adota a abordagem oposta, começando com cada amostra como um cluster individual e mesclando os pares mais próximos de clusters até que apenas um cluster permaneça.
#
# Nesta seção, vamos nos concentrar no agrupamento aglomerativo, pois é mais comum e útil para obter insights.
# + [markdown] id="VLvEAsjKbfh9"
# Os dois algoritmos padrão para agrupamento hierárquico aglomerativo são os de **Simple Linkage** e **Complete Linkage**.
#
# * **Simple Linkage**: calculamos as distâncias entre os membros mais semelhantes para cada par de clusters e fundimos os dois clusters para os quais a distância entre os membros mais semelhantes é a menor.
#
# * **Complete Linkage**: é semelhante à Simple Linkage, mas, em vez de comparar os membros mais semelhantes em cada par de clusters, comparamos os membros mais diferentes para realizar a fusão.
# + [markdown] id="-86AOeDDbfh-"
# Outros algoritmos comumente usados para agrupamento hierárquico aglomerativo incluem **average linkage** e **Ward's linkage**. No average linkage, mesclamos os pares do cluster com base nas distâncias médias mínimas entre todos os membros do grupo nos dois clusters. No Ward's linkage, os dois clusters que levam ao aumento mínimo do SSE total dentro do cluster são mesclados.
# + [markdown] id="xq_fi62Vbfh_"
# **Método Aglomerativo:**
#
# Este é um procedimento iterativo que pode ser resumido pelo
# seguintes passos:
#
# 1. Calcule a matriz de distância de todas as amostras.
# 2. Represente cada ponto de dados como um cluster singleton.
# 3. Mescle os dois clusters mais próximos com base na distância dos membros mais diferentes (distantes).
# 4. Atualize a matriz de distância.
# 5. Repita as etapas 2 a 4 até que um único cluster permaneça.
#
# 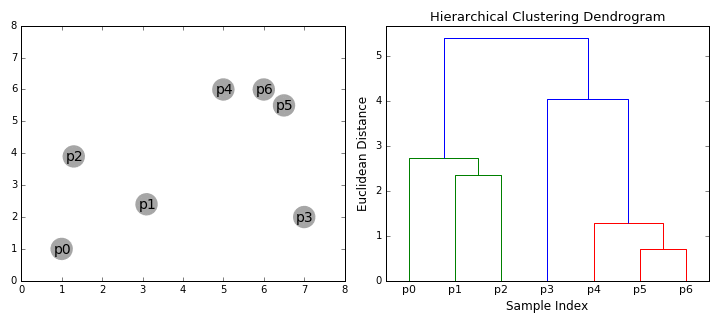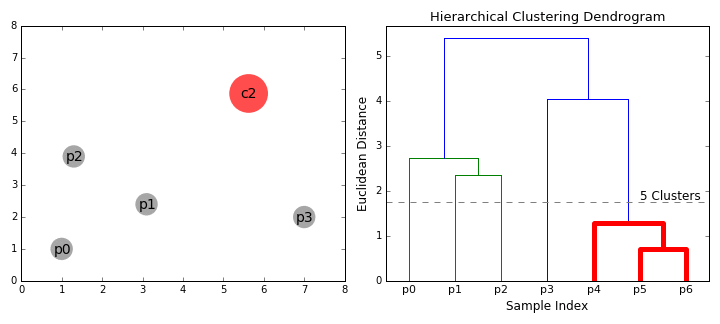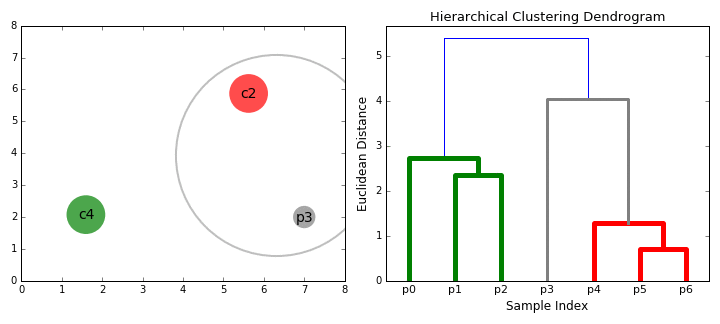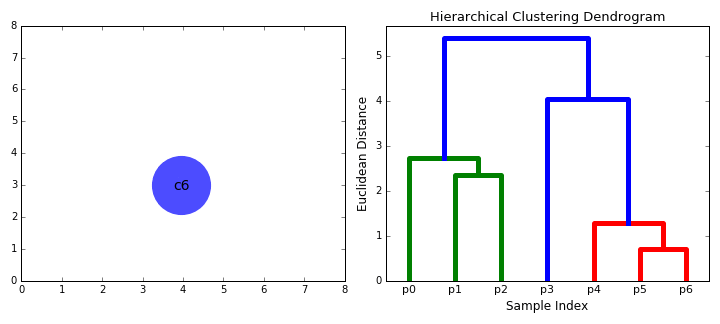
# + [markdown] id="-tg1zJ55bfiA"
# Vamos usar a biblioteca ```scipy``` para traçar um dendograma e visualizar o número de clusters que podem se ajustar melhor aos nossos dados e mostrar```AgglomerativeClustering``` de ``` sklearn``` para computar facilmente os clusters.
# + id="gYahfbETbfiA" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="b94a260a-04d7-4247-c869-7b9b2f4a58a6"
#Gerando amostra com 5 clusters
X,y = make_blobs(n_samples=500,
n_features=2,
centers=5,
cluster_std=0.5,
shuffle=True,
random_state=0)
# Gráfico
plt.scatter(X[:,0],
X[:,1],
c="black",
marker='o',
s=50)
# + id="Vz2DZYJzbfiD" colab={"base_uri": "https://localhost:8080/", "height": 428} outputId="8278bad7-ba9f-4279-b55c-bd58560facc5"
# Importando libs
from scipy.cluster.hierarchy import dendrogram, linkage
from matplotlib import pyplot as plt
# Criação do cluster hierarquico
linked = linkage(X, method='complete', metric='euclidean', optimal_ordering=True)
# Dendograma
plt.figure(figsize=(8, 7))
dendrogram(linked,
orientation='top',
distance_sort='descending',
show_leaf_counts=True)
h_line = 3
plt.axhline(y=h_line, c='k')
plt.show()
# + [markdown] id="vwNw_53nbfiO"
# Este dendrograma nos mostra que temos uma grande diminuição na distância global ao quebrar de 1 para 2 clusters, e também de 2 para 3, e assim por diante até chegarmos de 4 para 5 clusters (aqueles acima da linha horizontal quando ```hline = 3```). Então, a partir de 6 clusters, as distâncias quando adicionamos mais um cluster são muito semelhantes. Isso nos diz, semelhante à regra do cotovelo, que o número de clusters a serem usados aqui é 5.
#
# Agora podemos usar ``AgglomerativeClustering`` para separar nossos dados.
# + id="Ko7UqOkKbfiP"
# Importando AgglomerativeClustering e StandardScaler
from sklearn.cluster import AgglomerativeClustering
from sklearn.preprocessing import StandardScaler
# Criação do objeto Agglomerative Clustering
ac = AgglomerativeClustering(affinity='euclidean',linkage='ward', n_clusters=5)
# Fitting e predicting
cluster_ac = ac.fit_predict(X)
# + id="eoNxegYhbfiT" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="f8e897e9-8ea6-4153-f2e1-9529a6d4ba16"
# Clusters com cores
for cluster in range(ac.n_clusters):
plt.scatter(X[cluster_ac==cluster,0],
X[cluster_ac==cluster,1],
s=50,
cmap='Pastel1',
marker='s',
label='cluster {}'.format(cluster))
plt.legend()
plt.grid()
plt.show()
# + [markdown] id="0Ny9N_f0bfij"
#
# ### Exercicio 3
#
# Vamos tentar usar o clustering hierárquico para nosso conjunto de dados de varejo.
# + id="ixRmqr5nbfik"
# Criação do Cluster Hierarquico
linked_customers = linkage(customers_df, method='complete', metric='euclidean', optimal_ordering=True)
# + id="549SMRo4bfi4" colab={"base_uri": "https://localhost:8080/", "height": 435} outputId="db5896fb-fce1-480a-e6ed-c423c9e411fe"
# Dendograma
plt.figure(figsize=(14, 7))
dendrogram(linked_customers,
truncate_mode='lastp', # usamos o modo truncado para que possamos definir um número máximo p de clusters para mostrar
p=100, # definindo p como 100 para tornar o gráfico mais fácil de ler
orientation='top',
distance_sort='descending',
show_leaf_counts=True)
plt.show()
# + [markdown] id="fs4aW4G_bfi-"
# #### Parte 1
# Assim como no KMeans, utilize o gráfico anterior para definir o número de clusters. Em seguida, aplique o ``AgglomerativeClustering`` e plote o histograma dos clientes em cada cluster.
# + id="I1o4Wv5Ybfi_"
# Criando o objeto AgglomerativeClustering
ac_customers = AgglomerativeClustering(affinity='euclidean',linkage='ward', n_clusters=5)
# Fitting e predicting
cluster_ac_customers = ac_customers.fit_predict(X)
# + id="d0bklBoHbfjG" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="6d941231-29d8-4c3d-b0f0-5005c5835bff"
# Mostrando a quantidade de clientes por clusters
for cluster in range(ac_customers.n_clusters):
plt.scatter(X[cluster_ac_customers==cluster,0],
X[cluster_ac_customers==cluster,1],
s=50,
cmap='Pastel1',
marker='s',
label='cluster {}'.format(cluster))
plt.legend()
plt.grid()
plt.show()
# + [markdown] id="BRUtnFw3aUwx"
# #### Parte 2
# Agora possuimos novos clusters! Analise e discuta as principais diferenças dos clusters obtidos com o método hierárquico e com o método anterior.
# + [markdown] id="Di4Iei5mbfjK"
# <a id="soft_clustering_analysis"></a>
# ## 4. Soft Clustering
# + [markdown] id="DfOE9WjsbfjO"
# Nesta seção, apresentaremos a análise de Soft Clustering. Para isso, utilizaremos a base de dados 2018_medias_jogadores.xlsx. Ela é baseada no Cartola FC, que é um jogo que deixa os torcedores muito mais próximos da função de técnico e de diretor de um clube da Série A do Campeonato Brasileiro. O cartoleiro tem como missão escalar seu time a cada rodada do Campeonato Brasileiro, considerando que alguns atletas podem estar lesionados, suspensos ou em condições incertas para entrar em campo na próxima partida. Além disso, cada atleta apresenta diferentes atributos que podem ajudar o cartoleiro a escalar o time da melhor maneira possível, sabendo que cada jogador tem seu preço (medido em cartoletas, moeda oficial do jogo).
# + [markdown] id="cPTkyZiIbfjR"
# <a id="problem_definition_soft_clustering"></a>
# ### 4.1. Definição do Problema
#
# Suponha que você seja um jogador do Cartola FC e você não escolheu o time da sua semana. Então imagine o seguinte cenário: você está ficando sem dinheiro e por isso não consegue escolher o jogador que está acostumado. Então você se pergunta: _"Que jogador devo escolher?"_.
#
# Este é um exemplo de problema que pode ser resolvido usando uma técnica de análise de Clusterização. Por exemplo, poderíamos agrupar os jogadores com as mesmas características e, entre aqueles que são semelhantes ao avatar a que está habituado, poderíamos escolher outro com preço inferior.
# + id="cv5NxrfYbfjS" colab={"base_uri": "https://localhost:8080/", "height": 360} outputId="5c634b29-fdf3-4d14-f152-f72674e88da9"
# Importando as Bibliotecas e as base de dados
import pandas as pd
import numpy as np
from sklearn.utils import shuffle
df_orig = pd.read_excel('/mydrive/dados/2018_medias_jogadores.xlsx')
df_orig = shuffle(df_orig).reset_index(drop = True)
df_orig.head()
# + [markdown] id="kZHXMeXgbfjY"
# <a id="initial_analysis_soft_clustering"></a>
# ### 4.2. Análises Iniciais
#
# A principal diferença entre os métodos tradicionais de clustering (o que chamamos de hard clustering) e o soft clustering são os componentes de cada cluster. No Hard Clustering, cada ponto no conjunto de dados pertence a apenas um cluster, enquanto no Soft Clustering cada ponto tem uma probabilidade de estar em um determinado cluster. Em outras palavras, o agrupamento é flexível a ponto de permitir que um item possa existir em "vários clusters".
#
# No nosso caso, imagine que estamos acostumados a escolher o jogador <NAME>, meio-campista do Palmeiras, que vale 8,22 cartoletas. Infelizmente esta semana não podemos pagar este valor. Suponha que usamos um método de agrupamento para descobrir quem são os jogadores semelhantes ao Lucas Lima e seus preços. Podemos ver algumas boas opções mais baratas que Lucas Lima no cluster, então podemos inferir que se pegarmos Everton Ribeiro, por exemplo, há uma grande chance de termos um score semelhante pagando menos cartoletas.
#
# 
#
# Se este problema fosse resolvido usando um método de hard clustering e analisássemos o cluster de Everton Ribeiro, o cluster seria exatamente igual ao de Lucas Lima (mostrado acima). Por outro lado, se escolhermos um método de Soft Clustering, para cada avatar haveria um cluster específico com os mais semelhantes entre eles. Dê uma olhada em um possível soft cluster para Everton Ribeiro e observe que o cluster não é exatamente o mesmo que o de Lucas Lima.
#
# 
#
# O soft clustering nos traz um cluster mais customizado para cada jogador analisado, já que cada jogador tem uma probabilidade de ser semelhante a todos os outros jogadores no jogo de fantasia.
# + [markdown] id="Lx7hF_2SbfjZ"
# <a id="modeling_soft_clustering"></a>
# ### 4.3. Modelagem
# + [markdown] id="sIaLiximbfja"
# Não existe apenas uma implementação da técnica Soft Clustering. Nesta seção, apresentamos uma (das muitas possíveis) implementação baseada no modelo Random Forest. Portanto, para implementá-lo, primeiro precisamos importar todos os pacotes necessários do ```scikit-learn```.
# + id="K92GkMmSbfja"
# Importando Bibliotecas
from sklearn.model_selection import cross_val_score, GroupKFold
from sklearn.ensemble import RandomForestRegressor
from sklearn.utils import shuffle
from sklearn.metrics import make_scorer
from sklearn.preprocessing import OneHotEncoder
from sklearn.neighbors import KNeighborsRegressor
# + [markdown] id="RQk5Zchvbfjj"
# Além disso, precisamos separar nossas variáveis X e y. Mas, para fazer isso, existem algumas features no conjunto de dados que não devem ser considerados na clusterização. Por exemplo, algumas características categóricas e outras que podem ser extremamente correlacionadas ao preço dos jogadores (relação de causa e efeito).
# + id="QgiB5a3-bfjk"
# Eliminando Variaveis Desnecessárias
cols_to_drop = ['player_position','score_no_cleansheets_mean']
df = df_orig.drop(cols_to_drop, axis=1)
df.head(5)
# Removendo possíveis variáveis de causa e efeito e removendo outras categóricas
cols_possible_cause_effect = ['score_mean','diff_home_away_s','score_mean_home','score_mean_away']
X = df.copy()
X.drop(cols_possible_cause_effect, axis=1, inplace=True)
X = X.loc[:, 'position_id' : 'DP_mean'].fillna(0)
# Obtendo a váriavel Y
y = df['price_cartoletas']
# + [markdown] id="4dKVikRmbfjp"
# Agora que os dados estão estruturados corretamente, podemos modelar nosso problema usando a implementação ``scikit-learn`` Random Forest.
# + id="KMsPYdDjbfjq" colab={"base_uri": "https://localhost:8080/"} outputId="f320e7d0-b09b-4f9a-ddcd-320686247d1f"
# Gerando o Modelo de Random Forest
rfr = RandomForestRegressor(n_estimators=500, criterion='mse', min_samples_leaf=5)
# Fit
rfr.fit(X, y)
# Obtendo a importância das variaveis
importances = pd.Series(index=X.columns, data=rfr.feature_importances_)
importances.sort_values(ascending=False, inplace=True)
print('Variable importances:\n',importances)
# + [markdown] id="SlFwlpcQbfjy"
# A implementação parte da ideia de que, para cada jogador, obtemos uma lista de outros jogadores semelhantes ao que foi analisado. Isso pode ser alcançado criando uma matriz dissimilar - esta matriz fornece uma estimativa grosseira da distância entre as amostras com base na proporção de vezes que as amostras terminam no mesmo nó de folha na floresta aleatória (não se preocupe com estes detalhes mais técnicos, explicaremos melhor nas próximas aulas).
# + id="k6JYtBP_bfj2" colab={"base_uri": "https://localhost:8080/"} outputId="76afc6b6-a0b1-453a-aa17-d4479e9faa10"
# Obtendo Folhas
leaves = rfr.apply(X)
print('\nNº Folhas:\n', leaves, '\n\nDimensão das Folhas:', leaves.shape)
# Construindo a matriz de dissimilaridade
M = leaves.copy()
M = OneHotEncoder().fit_transform(M)
M = (M * M.transpose()).todense()
M = 1 - M / M.max()
print('\nMatriz de Dissimilaridade:\n', M, '\n\nDimensão da Matriz:', M.shape)
# + [markdown] id="4rHtixZvbfj8"
# Observe que, na matriz de dissimilaridade, cada linha *i* e cada coluna *j* representa um jogador. Portanto, o valor (*i*, *j*) é dado pela frequência com que o jogador *i* e o jogador *j* terminaram no mesmo nó folha no Modelo Random Forest (o que significa que eles são similares). Agora, estamos quase prontos para responder ao nosso problema. Mas primeiro, vamos construir nossa estrutura de cluster!
# + id="cQ9EGzBFbfj-"
# Construindo Clusters
size_of_cluster = 5
neighboors = []
distances = []
for i in range(len(leaves)):
s = pd.Series(np.array(M[i])[0])
s.drop(i, inplace=True)
# Ordenando os Jogadores
s.sort_values(ascending=True, inplace=True)
neighboors.append([i] + list(s[:size_of_cluster].index))
distances.append([0] + list(s[:size_of_cluster].values))
# Salvando Cluster na estrutura
clusters = {}
for i in range(len(neighboors)):
L = []
for j in range(len(neighboors[i])):
L.append([neighboors[i][j], y[neighboors[i][j]]])
clusters['C' + str(i)] = L
# Função usada para responder à pergunta: "Quais são os jogadores semelhantes ao Lucas Lima?"
def getCluster(df, clusters, search_variable, key=None, index=None):
if index == None:
index = df[df[search_variable] == key].index[0]
return df.iloc[[e[0] for e in clusters['C' + str(index)]]]
# + [markdown] id="wvA0GpebbfkC"
# Agora está tudo pronto! Use a célula abaixo para verificar os clusters de cada um dos possíveis jogadores!
# + id="yi1c_NxmbfkF" colab={"base_uri": "https://localhost:8080/", "height": 459} outputId="82d2a029-7a19-470e-dcff-34ce8e596823"
getCluster(df=df, clusters=clusters, search_variable='player_slug', key='lucas-lima')
# + [markdown] id="F3gaH65pT3Tw"
# Parabéns! Você conseguiu obter uma lista de jogadores similares a <NAME>, o que lhe permite escolher um que tenha um custo menor.
# + [markdown] id="3wIUlVqWgId1"
# <a id="pan"></a>
# ## Fim da Aula!
# Com a conclusão desta aula esperamos que você esteja mais familiarizado com os conceitos de Aprendizagem não Supervisionada!
# Na próxima aula iremos abordar o outro lado da moeda, ou seja, o Aprendizado Supervisionado! Iremos elucidar alguns conceitos e técnicas associados aos problemas de Classificação!
#
# Até a próxima aula!!
| Aprendizado_Nao_Supervisionado.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="qM_XHcPVzUei"
# # UMAP embeddings for the monthly data comments from subreddits
# + [markdown] id="4FFq3DIYzgak"
# # IMPORT MODULES
# + executionInfo={"elapsed": 1914, "status": "ok", "timestamp": 1617608526887, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06886034211416817939"}, "user_tz": -120} id="J3vC9YwjzTi4"
#import json
import os
#from google.colab import drive
from tqdm.notebook import tqdm
import pickle
from collections import Counter
from datetime import datetime, timedelta
import pandas as pd
import numpy as np
from scipy import spatial
import umap
import matplotlib.pyplot as plt
# import torch
# from sentence_transformers import SentenceTransformer, util
#from sklearn.metrics.pairwise import cosine_similarity
# from sklearn.decomposition import PCA
# from sklearn.manifold import TSNE
# from sklearn.cluster import KMeans
# from sklearn.cluster import OPTICS
# import seaborn as sns
# + [markdown] id="3ZT3Z71WzpsJ"
# # TECHNICAL FUNCTIONS
# + executionInfo={"elapsed": 854, "status": "ok", "timestamp": 1617608526888, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06886034211416817939"}, "user_tz": -120} id="lQnpzkhQzTlF"
def get_date_range(month_start, year_start, month_end, year_end):
from itertools import cycle
month_range = list(range(1,13))
cycle_month_range = cycle(month_range)
while True:
current_month = next(cycle_month_range)
if current_month == month_start:
break
date_tuples = []
year = year_start
while True:
date_tuples.append((current_month, year))
if year == year_end and current_month == month_end:
break
current_month = next(cycle_month_range)
if current_month == 1:
year += 1
return date_tuples
# + [markdown] id="Loyy5mT1zxWx"
# # UPLOAD DATA
# + executionInfo={"elapsed": 335, "status": "ok", "timestamp": 1617608527092, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06886034211416817939"}, "user_tz": -120} id="LYD2LqiwzTnQ"
# google_drive_path = "./"
comptech_opinion_analizer_path = "./"
# + colab={"base_uri": "https://localhost:8080/", "height": 66, "referenced_widgets": ["363e44aafcdf44a682124d8a01a27a46", "2b41d2b3a9f94eeca0b0e3ad0cbc557d", "98261b6660e042699989c1ac169455ab", "61d761e57dac42988b9e5a9f4f38ecc5", "251bb3c61c2746cc8639de8ea167506a", "487255908c35447e9220909ae32b3990", "<KEY>", "<KEY>"]} executionInfo={"elapsed": 33792, "status": "ok", "timestamp": 1617608560922, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06886034211416817939"}, "user_tz": -120} id="hHQtO4_dzTpg" outputId="a1ed8402-e665-4533-8541-6b4a3515592a"
# UPLOAD THE DATA
data_dir = os.path.join(comptech_opinion_analizer_path, "embeddings_bert/")
data_files = [f for f in os.listdir(data_dir) if "pickle" in f]
entity = "Trump"
entity_data_files = sorted([f for f in data_files if entity in f])
df_vecs = pd.DataFrame()
for f in tqdm(entity_data_files):
data_path = os.path.join(data_dir, f)
df_vecs = df_vecs.append(pickle.load(open(data_path, "rb")))
# + [markdown] id="nwWI5u0oz7I2"
# # Show the timeline of comment counts
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 159} executionInfo={"elapsed": 33287, "status": "ok", "timestamp": 1617608561419, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06886034211416817939"}, "user_tz": -120} id="R2ogWUfpzTrb" outputId="f3d00ad3-9044-49e3-b14b-3dd52ae60d07"
created_list = sorted(df_vecs.created_utc.to_list())
b_width = 3600*24*3 # weekly
bins = np.arange(min(created_list), max(created_list) + 1, b_width)
hist, bins = np.histogram(created_list, bins = bins)
dt_bins = [datetime.fromtimestamp(t) for t in bins[:-1]]
plt.figure(figsize=(15,1.5))
plt.title(f"/r/{entity} :: Number of comments per week")
plt.plot(dt_bins, hist, marker = "x")
plt.xlabel("Time")
plt.ylabel("Count")
plt.show()
# + [markdown] id="aq1AxNmb0Dg2"
# # TSNE EMBEDDING OF COMMENTS
# + executionInfo={"elapsed": 915, "status": "ok", "timestamp": 1617608562340, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06886034211416817939"}, "user_tz": -120} id="AWdnoZWQ0FtF"
# ADD FOLDER
# colab_notebooks_path = os.path.join(google_drive_path, "Colab Notebooks/opinion_analyzer/")
umap_embedding_dir = os.path.join(comptech_opinion_analizer_path, "umap_embeddings")
os.makedirs(umap_embedding_dir, exist_ok = True)
# + colab={"base_uri": "https://localhost:8080/", "height": 66, "referenced_widgets": ["356e4993ab144abe8cfc278907f348d4", "cb4ef2239edc479ca9c74c1589718a37", "<KEY>", "<KEY>", "afe3e4cf09ee40a0b4f74485819026b2", "f94ba187485e47b28676319c821d7904", "4b24f2eecae94cf6867811a9de6be66e", "84c371ee88be44e08f5a3a9f35d74ddd"]} id="mk3x6vfszTtq" outputId="44f4f365-e231-4688-8202-b69cd668b7d5"
# DIMENSIONALITY REDUCTION FOR ALL MONTHLY DATA
date_range = get_date_range(1, 2020, 1, 2021)
min_dist = 0.001
for my_start, my_end in tqdm(list(zip(date_range, date_range[1:]))):
# PREPARATIONS
dt_start = datetime(my_start[1], my_start[0], 1)
dt_end = datetime(my_end[1], my_end[0], 1)
month_str = dt_start.strftime("%b %Y")
t_start, t_end = dt_start.timestamp(), dt_end.timestamp()
month_vecs_df = df_vecs[(t_start < df_vecs.created_utc ) & (df_vecs.created_utc < t_end)]
month_embeddings = month_vecs_df.embedding.to_list()
month_labels = month_vecs_df.body.to_list()
month_ids = month_vecs_df.link_id.to_list()
print(f"Month labels {len(month_labels)}")
# TSNE
embedder = umap.UMAP(min_dist = min_dist, metric = "cosine")
month_embeddings_2d = embedder.fit_transform(month_embeddings)
# OUTPUT
out_file = f"umap_embedding_2d_{entity}_{my_start[0]}_{my_start[1]}_min_dist_{round(min_dist, 2)}.pickle"
out_path = os.path.join(umap_embedding_dir, out_file)
out_pack = (month_ids, month_labels, month_embeddings_2d)
pickle.dump(out_pack, open(out_path, "wb"))
# + [markdown] id="jSo7b_9Q1U4B"
# # Visualisation of comments each month
# + id="K_r3pzNhzTvo"
target_month = 1
dt_start = datetime(2020, target_month, 1)
dt_end = datetime(2020, target_month+1, 1)
dt_str = dt_start.strftime("%b %Y")
dt_month = int(dt_start.strftime("%m"))
dt_year = int(dt_start.strftime("%Y"))
t_start, t_end = dt_start.timestamp(), dt_end.timestamp()
month_vecs_df = df_vecs[(t_start < df_vecs.created_utc ) & (df_vecs.created_utc < t_end)]
# -
min_dist = 0.1
embedding_file = f"umap_embedding_2d_{entity}_{dt_month}_{dt_year}_min_dist_{round(min_dist, 2)}.pickle"
embedding_path = os.path.join(umap_embedding_dir, embedding_file)
(month_ids, month_labels, month_embeddings_2d) = pickle.load(open(embedding_path, "rb"))
# + id="Jv4ryTEG1XUZ"
month_labels_short = [s[:60]+"..." if len(s)>60 else s for s in month_labels]
len(month_labels_short)
# + id="37on4pv31YU0"
# VISUALISATION
import plotly.graph_objects as go
marker_style = dict(color='lightblue', size=6, line=dict(color='black', width = 0.5))
X, Y = zip(*month_embeddings_2d)
scatter_gl = go.Scattergl(x = X, y = Y, hovertext = month_labels_short, mode='markers', marker= marker_style)
fig = go.Figure(data = scatter_gl)
fig.update_layout(width=1000, height=700, plot_bgcolor = "white", margin=dict(l=10, r=10, t=30, b=10),
title=f"UMAP comments /r/{entity} :: period {dt_str}")
fig.show()
# -
| data_2d_projection/umap embeddings monthly - Trump.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7
# language: python
# name: python3
# ---
# + [markdown] nbgrader={"grade": false, "locked": true, "solution": false} editable=false deletable=false
# # Ridge Regression and Bayesian Methods
#
#
# **_Author: <NAME>, W.P.G.Peterson_**
#
# **_Reviewer: <NAME>_**
#
# **Expected time = 2.5 hours**
#
# **Total points = 80 points**
#
#
#
# ## Assignment Overview
#
# This assignment will test your ability to code your own version of ridge-regularized regression in `Python`. Ridge Regression offers a way to mitigate some of the weaknesses of Least Squares Linear Regression to build more robust models. This assignment draws upon and presupposes the knowledge found in the lectures for Module 13.
#
# The assignment also builds upon the work performed in the module 13 assignment "*Linear Regression - Least Squares*". The data used will be the same. Though the last assignment tested your ability to read data into `Pandas` from a `.csv`. Those fundamental processes will not be directly tested here.
#
#
# This assignment is designed to build your familiarity and comfort coding in Python while also helping you review key topics from each module. As you progress through the assignment, answers will get increasingly complex. It is important that you adopt a data scientist's mindset when completing this assignment. **Remember to run your code from each cell before submitting your assignment.** Running your code beforehand will notify you of errors and give you a chance to fix your errors before submitting. You should view your Vocareum submission as if you are delivering a final project to your manager or client.
#
# ***Vocareum Tips***
# - Do not add arguments or options to functions unless you are specifically asked to. This will cause an error in Vocareum.
# - Do not use a library unless you are expicitly asked to in the question.
# - You can download the Grading Report after submitting the assignment. This will include feedback and hints on incorrect questions.
#
#
# ### Learning Objectives
#
# - Test both Python and mathematical competencies in ridge regression
# - Calculate ridge regression weights using linear algebra
# - Understand how to standardize data and its working
# - Process data for regularized methods
# - Implement ridge regression from scratch
# - Use Naïve grid search to optimize regularization and set hyperparameters
# - Familiarize with the concept of hyperparameter tuning
# -
# ## Index:
#
# #### Ridge Regression and Bayesian Methods
#
# - [Question 1](#q1)
# - [Question 2](#q2)
# - [Question 3](#q3)
# - [Question 4](#q4)
# - [Question 5](#q5)
# - [Question 6](#q6)
# - [Question 7](#q7)
# - [Question 8](#q8)
# - [Question 9](#q9)
# - [Question 10](#q10)
# - [Question 11](#q11)
# ## Ridge Regression and Bayesian Methods
#
#
# For this assignment, we will use a regression model on a housing price dataset to predict the price of a house based on its living area above the ground. More information about this dataset can be found [here](https://www.kaggle.com/c/house-prices-advanced-regression-techniques/data).
#
# Before coding an algorithm, we will take a look at our data using Python's `pandas`. For visualizations, we'll use `matplotlib`.
# Let's import the necessary libraries and load the datasets by using the pandas pd.read_csv() function.
#
#
# + nbgrader={"grade": false, "locked": true, "solution": false} editable=false deletable=false
### Import the necessary modules
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = (20.0, 10.0)
### Read in the data
tr_path = 'data/train.csv'
data = pd.read_csv(tr_path)
# -
# We begin by performing some basic exploratory data analysis by using the function head() and the attribute columns.
#
data.head()
# + [markdown] nbgrader={"grade": false, "locked": true, "solution": false} editable=false deletable=false
# [Back to top](#Index:)
# <a id='q1'></a>
#
# ### Question 1:
#
# *5 points*
#
# Which column has the most "null" values? Assign name of the column as string to `ans1a`.
# How many nulls are in that column? Assign number as int to `ans1b`.
#
# **NOTE:** To find the column that has the higher number of null values, use the `idxmin()` function.
# +
### GRADED
### YOUR SOLUTION HERE
ans1a = None
ans1b = None
###
### YOUR CODE HERE
###
# + nbgrader={"grade": true, "grade_id": "Question 01", "locked": true, "points": "5", "solution": false} editable=false deletable=false
###
### AUTOGRADER TEST - DO NOT REMOVE
###
# -
# Next, let's plot the relationship between our variables of interest: the price for each house and the above ground living area in square feet.
#
# We can do so by creating a scatter plot using matplotlib.
# + nbgrader={"grade": false, "locked": true, "solution": false} editable=false deletable=false
data.plot('GrLivArea', 'SalePrice', kind = 'scatter', marker = 'x');
# + [markdown] nbgrader={"grade": false, "locked": true, "solution": false} editable=false deletable=false
# [Back to top](#Index:)
# <a id='q2'></a>
#
# ### Question 2:
#
# *5 points*
#
# Create a subset of our dataframe below containing only the "Street" and "Alley" columns from the `data`.
# Assign the new dataframe to 'ans2'
# +
### GRADED
### YOUR SOLUTION HERE
ans2 = None
###
### YOUR CODE HERE
###
# + nbgrader={"grade": true, "grade_id": "Question 02", "locked": true, "points": "5", "solution": false} editable=false deletable=false
###
### AUTOGRADER TEST - DO NOT REMOVE
###
# + [markdown] nbgrader={"grade": false, "locked": true, "solution": false} editable=false deletable=false
# <a id = "code"></a>
# ### Coding Ridge Regression
#
# #### Preprocessing
# Before implementing ridge regression, it is important to mean-center our target variable and mean-center and standardize observations. We will do this by using the following formulas:
# #### Mean Center Target
# $$y_{cent} = y_0 - \bar{y}$$
#
# #### Standardize Observations
# $$X_{std} = \frac{X_0-\bar{X}}{s_{X}}$$
#
# Where $\bar{X}$ is the sample mean of X and $s_{X}$ is the sample standard deviation of X.
#
# NOTE: The sample standard deviation should be calculated with 0 "Delta Degrees of Freedom"
#
# -
# [Back to top](#Index:)
# <a id='q3'></a>
#
# ### Question 3:
#
# *5 points*
#
# Why are the centering/standardization transformations described above important for ridge regression?
# - a) Regression works best when values are unitless
# - b) The transformations makes the regression more interpretable
# - c) Ridge penalizes large coefficients; the transformations make the coefficients of similar scales
# - d) It isn't important
#
# Assign character associated with your choice as a string to `ans3`.
# +
### GRADED
### YOUR ANSWER BELOW
ans3 = None
###
### YOUR CODE HERE
###
# + nbgrader={"grade": true, "grade_id": "Question 03", "locked": true, "points": "5", "solution": false} editable=false deletable=false
###
### AUTOGRADER TEST - DO NOT REMOVE
###
# + [markdown] nbgrader={"grade": false, "locked": true, "solution": false} editable=false deletable=false
# [Back to top](#Index:)
# <a id='q4'></a>
#
# ### Question 4:
#
# *10 points*
#
# Standardized values can be calculated using the following formula:
#
# $$X_{std} = \frac{X_0-\bar{X}}{s_{X}}.$$
#
# Define a function "standardize" that accepts, as input a list of numbers and returns a list where those values have been standardized. Please use the NumPy function `std()` for calculating standard deviation
#
# NOTE: The sample standard deviation should be calculated with 0 "Delta Degrees of Freedom".
# If your answer does not match the example answer, check the default degrees of freedom in your standard deviation function.
# +
### GRADED
### YOUR ANSWER BELOW
def standardize( num_list):
"""
Standardize the given list of numbers
Positional arguments:
num_list -- a list of numbers
Example:
num_list = [1,2,3,3,4,4,5,5,5,5,5]
nl_std = standardize(num_list)
print(np.round(nl_std,2))
#--> np.array([-2.11, -1.36, -0.61, -0.61,
0.14, 0.14, 0.88, 0.88, 0.88,
0.88, 0.88])
NOTE: the sample standard deviation should be calculated with 0 "Delta Degrees of Freedom"
"""
return
###
### YOUR CODE HERE
###
# + nbgrader={"grade": true, "grade_id": "Question 04", "locked": true, "points": "10", "solution": false} editable=false deletable=false
###
### AUTOGRADER TEST - DO NOT REMOVE
###
# + [markdown] nbgrader={"grade": false, "locked": true, "solution": false} editable=false deletable=false
# Below we will create a function which will preprocess our data by performing:
# * mean subtraction from $y$,
# * dimension standardization for $x$.
#
# The formulas to Mean Center Target and Standardize Observations are given above.
#
# NOTE: The sample standard deviation should be calculated with 0 "Delta Degrees of Freedom"
# If your answer does not match the example answer, check the default degrees of freedom in your standard deviation function.
#
# [Back to top](#Index:)
# <a id='q5'></a>
#
# ### Question 5:
#
# *10 points*
#
# Code a function called "preprocess_for_regularization" that accepts, as input, the DataFrame, a `y_column_name` input and an `x_column_names` input input
# Your function should preprocess our data by performing:
# - mean subtraction from $y$,
# - dimension standardization for $x$
# +
### GRADED
### YOUR ANSWER BELOW
def preprocess_for_regularization(data, y_column_name, x_column_names):
"""
Perform mean subtraction and dimension standardization on data
Positional argument:
data -- a pandas dataframe of the data to pre-process
y_column_name -- the name (string) of the column that contains
the target of the training data.
x_column_names -- a *list* of the names of columns that contain the
observations to be standardized
Returns:
Return a DataFrame consisting only of the columns included
in `y_column_name` and `x_column_names`.
Where the y_column has been mean-centered, and the
x_columns have been mean-centered/standardized.
Example:
data = pd.read_csv(tr_path).head()
prepro_data = preprocess_for_regularization(data,'SalePrice', ['GrLivArea','YearBuilt'])
print(prepro_data) #-->
GrLivArea YearBuilt SalePrice
0 -0.082772 0.716753 7800.0
1 -1.590161 -0.089594 -19200.0
2 0.172946 0.657024 22800.0
3 -0.059219 -1.911342 -60700.0
4 1.559205 0.627159 49300.0
NOTE: The sample standard deviation should be calculated with 0 "Delta Degrees of Freedom"
If your answer does not match the example answer,
check the default degrees of freedom in your standard deviation function.
"""
return
###
### YOUR CODE HERE
###
# + nbgrader={"grade": true, "grade_id": "Question 05", "locked": true, "points": "10", "solution": false} editable=false deletable=false
###
### AUTOGRADER TEST - DO NOT REMOVE
###
# + [markdown] nbgrader={"grade": false, "locked": true, "solution": false} editable=false deletable=false
# Next, you'll implement the equation for ridge regression using the closed form equation:
#
# $$w_{RR}=(\lambda+X^TX)^{-1}X^Ty$$
#
# The function will be very similar to the function you wrote for Least Squares Regression with a slightly different matrix to invert.
#
# NB: Many `numpy` matrix functions will be useful. e.g. `np.matmul`, `np.linalg.inv`, `np.ones`, `np.transpose`, and `np.identity`.
#
# The main change from Least Squares Regression is that $\lambda$ is a parameter *we* must set. This is different from the $w$ parameters that we calculate from either closed form or approximation algorithms.
#
# We will address tuning parameters such as $\lambda$ in the next section.
#
# [Back to top](#Index:)
# <a id='q6'></a>
#
# ### Question 6:
#
# *10 points*
#
# Code a function called "ridge_regression_weights" that takes, as input, three inputs: two matricies corresponding to the x inputs and y target and a number (int or float) for the lambda parameter
#
# Your function should return a numpy array of regression weights
#
# The following steps must be accomplished:
#
# Ensure the number of rows of each the X matrix is greater than the number of columns.
# If not, transpose the matrix.
# Ultimately, the y input will have length n. Thus the x input should be in the shape n-by-p
#
# *Prepend* an n-by-1 column of ones to the input_x matrix
#
# Use the above equation to calculate the least squares weights. This will involve creating the lambda matrix - a p+1-by-p+1 matrix with the "lambda_param" on the diagonal
# +
### GRADED
### YOUR ANSWER BELOW
def ridge_regression_weights(input_x, output_y, lambda_param):
"""Calculate ridge regression least squares weights.
Positional arguments:
input_x -- 2-d matrix of input data
output_y -- 1-d numpy array of target values
lambda_param -- lambda parameter that controls how heavily
to penalize large weight values
Example:
training_y = np.array([208500, 181500, 223500,
140000, 250000, 143000,
307000, 200000, 129900,
118000])
training_x = np.array([[1710, 1262, 1786,
1717, 2198, 1362,
1694, 2090, 1774,
1077],
[2003, 1976, 2001,
1915, 2000, 1993,
2004, 1973, 1931,
1939]])
lambda_param = 10
rrw = ridge_regression_weights(training_x, training_y, lambda_param)
print(rrw) #--> np.array([-576.67947107, 77.45913349, 31.50189177])
print(rrw[2]) #--> 31.50189177
Assumptions:
-- output_y is a vector whose length is the same as the
number of observations in input_x
-- lambda_param has a value greater than 0
"""
weights = np.array([])
return weights
###
### YOUR CODE HERE
###
# + nbgrader={"grade": true, "grade_id": "Question 06", "locked": true, "points": "10", "solution": false} editable=false deletable=false
###
### AUTOGRADER TEST - DO NOT REMOVE
###
# + [markdown] nbgrader={"grade": false, "locked": true, "solution": false} editable=false deletable=false
# ### Selecting the $\lambda$ parameter
#
# For our final function before looking at the `sklearn` implementation of ridge regression, we will create a hyperparameter tuning algorithm.
#
# In ridge regression, we must pick a value for $\lambda$. We have some intuition about $\lambda$ from the equations that define it: small values tend to emulate the results from Least Squares, while large values will reduce the dimensionality of the problem. But the choice of $\lambda$ can be motivated with a more precise quantitative treatment.
#
# Eventually, we will look to choose the value of $\lambda$ that minimizes the validation error, which we will determine using $k$-fold cross-validation.
#
# For this example, we will solve a simpler problem on finding a value that minimizes the list returned by the function.
# + nbgrader={"grade": false, "locked": true, "solution": false} editable=false deletable=false
### Example of hiden function below:
### `hidden` takes a single number as a parameter (int or float) and returns a list of 1000 numbers
### the input must be between 0 and 50 exclusive
def hidden(hp):
if (hp<=0) or (hp >= 50):
print("input out of bounds")
nums = np.logspace(0,5,num = 1000)
vals = nums** 43.123985172351235134687934
user_vals = nums** hp
return vals-user_vals
# -
hidden(10)
# + [markdown] nbgrader={"grade": false, "locked": true, "solution": false} editable=false deletable=false
# Run the above cell and test out the functionality of `hidden`. Remember, it takes a single number between 0 and 50 as an argument.
# [Back to top](#Index:)
# <a id='q7'></a>
#
# ### Question 7:
#
# *10 points*
#
# Code a function called "minimize" that takes, as input, a function.
#
# That function will be similar to `hidden` created above and available for your exploration. Like 'hidden', the passed function will take a single argument, a number between 0 and 50 exclusive and then, the function will return a numpy array of 1000 numbers.
#
# Your function should return the value that makes the mean of the array returned by 'passed_func' as close to 0 as possible
#
# Note, you will almost certainly NOT be able to find the number that makes the mean exactly 0
# +
### GRADED
### YOUR ANSWER BELOW
def minimize( passed_func):
"""
Find the numeric value that makes the mean of the
output array returned from 'passed_func' as close to 0 as possible.
Positional Argument:
passed_func -- a function that takes a single number (between 0 and 50 exclusive)
as input, and returns a list of 1000 floats.
Example:
passed_func = hidden
min_hidden = minimize(passed_func)
print(round(min_hidden,4))
#--> 43.1204 (answers will vary slightly, must be close to 43.123985172351)
"""
# Create values to test
test_vals = ...
# Find mean of returned array from function
ret_vals = ...
# Find smallest mean
min_mean = ...
# Return the test value that creates the smallest mean
return ...
###
### YOUR CODE HERE
###
# + nbgrader={"grade": true, "grade_id": "Question 07", "locked": true, "points": "10", "solution": false} editable=false deletable=false
###
### AUTOGRADER TEST - DO NOT REMOVE
###
# + [markdown] nbgrader={"grade": false, "locked": true, "solution": false} editable=false deletable=false
# The above simulates hyperparameter tuning.
#
# In the case of ridge regression, you would be searching lambda parameters to minimize the validation error.
#
# The `hidden` function would be analogous to the model building, the returned list would be analogous to the residuals, and the mean of that list would be analogous to the validation error.
#
# See below for an example of using the functions built above that automatically perform hyperparameter tuning using mean absolute deviation.
# + nbgrader={"grade": false, "locked": true, "solution": false} editable=false deletable=false
def lambda_search_func(lambda_param):
# Define X and y
# with preprocessing
df = preprocess_for_regularization(data.head(50),'SalePrice', ['GrLivArea','YearBuilt'])
y_true = df['SalePrice'].values
X = df[['GrLivArea','YearBuilt']].values
# Calculate Weights then use for predictions
weights = ridge_regression_weights(X, y_true, lambda_param )
y_pred = weights[0] + np.matmul(X,weights[1:])
# Calculate Residuals
resid = y_true - y_pred
# take absolute value to tune on mean-absolute-deviation
# Alternatively, could use:
# return resid **2-S
# for tuning on mean-squared-error
return abs(resid)
minimize(lambda_search_func) # --> about 2.9414414414414414
# + [markdown] nbgrader={"grade": false, "locked": true, "solution": false} editable=false deletable=false
# Implementing a k-folds cross-validation strategy will come in later assignments.
#
# [Back to top](#Index:)
# <a id='q8'></a>
#
# ### Question 8:
#
# *5 points*
#
# Why is cross-validation useful?
# - a) to minimize the liklihood of overfitting
# - b) Cross-validation allows us to fit on all our data
# - c) cross-validation standardizes outputs
# - d) cross-validation is not useful
#
# Assing the character associated with your choice as a string to `ans1`
# +
### GRADED
### YOUR ANSWER BELOW
ans1 = None
###
### YOUR CODE HERE
###
# + nbgrader={"grade": true, "grade_id": "Question 08", "locked": true, "points": "5", "solution": false} editable=false deletable=false
###
### AUTOGRADER TEST - DO NOT REMOVE
###
# + [markdown] nbgrader={"grade": false, "locked": true, "solution": false} editable=false deletable=false
# ### Ridge Regression in `sklearn`
#
# In the next question, we will ask you to implement Ridge regression in `sklearn`.
#
# [Back to top](#Index:)
# <a id='q9'></a>
#
# ### Question 9:
#
# *10 points*
#
# Use the function `LinearRegression` from `sklearn` to instantiate the classifier `lr`.
# Use the function `Ridge` from `sklear` to instantiate the classifier `reg`. For this classifier, set the parameter `alpha=100000`. Use the `Ridge` function to instantiate another classifier, `reg0`, but, this time, set `alpha=0`.
#
#
# Define a for loop with two indices, `m` and `name`. `m` will run over the three classifiers just defined and `name` will run over the list `["LeastSquares","Ridge alpha = 100000","Ridge, alpha = 0"]`. In each iteration of your fol loop, you should fit the "X" ('GrLivArea' and 'YearBuilt') and the "y" ('SalePrice') from `data`.
#
# Complete your for loop with the following print statement: `print(name, "Intercept:", m.intercept_, "Coefs:",m.coef_,"\n")`
#
#
# **NOTE: Note, the "alpha" parameter defines regularization strength. Lambda is a reserved word in `Python` -- Thus "alpha" instead**
# +
### GRADED
from sklearn.linear_model import Ridge, LinearRegression
### YOUR ANSWER BELOW
lr = None
reg = None
reg0 = None
###
### YOUR CODE HERE
###
# + nbgrader={"grade": true, "grade_id": "Question 09", "locked": true, "points": "10", "solution": false} editable=false deletable=false
###
### AUTOGRADER TEST - DO NOT REMOVE
###
# + [markdown] nbgrader={"grade": false, "locked": true, "solution": false} editable=false deletable=false
# Note in the above example, an alpha of 100,000 is set for the ridge regularization. The reason an alpha value this high is required is because standardization/mean centering of our inputs did not occur, and instead of working with inputs on the order of [-4,4] we are on the interval of [0,2000].
#
# [Back to top](#Index:)
# <a id='q10'></a>
#
# ### Question 10:
#
# *5 points*
#
# Above, the coefficent around 95/96 corresponds with:
# - a) Living Area
# - b) Year Built
# - c) Sale Price
# Assign character associated with your choice as string to `ans2`.
# +
### GRADED
### YOUR ANSWER BELOW
ans2 = None
###
### YOUR CODE HERE
###
# + nbgrader={"grade": true, "grade_id": "Question 10", "locked": true, "points": "5", "solution": false} editable=false deletable=false
###
### AUTOGRADER TEST - DO NOT REMOVE
###
# + [markdown] nbgrader={"grade": false, "locked": true, "solution": false} editable=false deletable=false
# [Back to top](#Index:)
# <a id='q11'></a>
#
# ### Question 11:
#
# *5 points*
#
# True or False:
#
# A larger "alpha" corresponds to a greater amount of regularization
#
# Assign boolean choice to `ans3`
# +
### GRADED
### YOUR SOLUTION HERE
ans3 = None
###
### YOUR CODE HERE
###
# + nbgrader={"grade": true, "grade_id": "Question 11", "locked": true, "points": "5", "solution": false} editable=false deletable=false
###
### AUTOGRADER TEST - DO NOT REMOVE
###
| AML_02_Ridge_Regression_Naive_Bayes/Ridge_Regression_Naive_Bayes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Getting started with DeBaCl
# ## 1. Create some data
# Our first step is to create some data using the scikit-learn `make_blobs` and `make_circles` utility. To make this a hard (but not impossible) clustering problem, we set the random state of the blob so that it's always outside the two concentric circles.
# +
import numpy as np
from sklearn.datasets import make_circles, make_blobs
import matplotlib.pyplot as plt
# %matplotlib inline
circles = make_circles(500, factor=0.5, noise=0.06, random_state=23)
blob = make_blobs(100, centers=1, center_box=(-1.7, 1.7), cluster_std=0.1,
random_state=19)
X = np.vstack((circles[0], blob[0]))
print "Dataset shape:", X.shape
# -
with plt.style.context('ggplot'):
fig, ax = plt.subplots(figsize=(6, 4.5))
ax.scatter(X[:, 0], X[:, 1], c='black', s=50, alpha=0.5)
fig.show()
# ## 2. Estimate the level set tree
# The Level Set Tree (LST) can be constructed directly from our tabular dataset. The most important choice is **the bandwidth parameter `k`, which controls the complexity** of the level set tree.
#
# Small values of `k` allow for complex trees with many leaves, which can good for discovering small clusters, with the caveat that these clusters may not be true features of the underlying data-generating process. Large values of `k` lead to very simple trees with few branches, but these trees are likely to be very stable across repeated samples from the same probability distribution.
#
# Choosing the bandwidth parameter in a principled way remains an open area of research, and a future tutorial will illustrate some helpful heuristics for finding a good value. For now, we use the value `k=20`, which was chosen through trial and error.
# +
import debacl as dcl
tree = dcl.construct_tree(X, k=20)
print tree
# -
# The summary of an LST is a table where each row corresponds to a cluster in the tree. Each cluster has an **ID number**, start and end **density levels**, start and end **mass levels**, a **parent** cluster, **child** clusters, and a list of the data points that belong to the cluster, represented in the table by the **size** of this list.
#
# The LST is constructed by finding connected components at successively higher levels in the estimated probability density function. Think of the density function as the water level rising around a set of islands representing the data points. When there is no water, all of the land masses are connected; as the water rises, land masses split into islands and vanish when the water gets high enough.
#
# The **start density level** is the density value where a cluster first appears by splitting from its parent cluster. The **end density level** is the density value where the cluster disappears, either by splitting into child clusters or by vanishing when all of its points have insufficiently high density.
#
# At each of the start and end density levels, a certain fraction of the points have been removed to create the *upper level sets* of the density function; these fractions are the **start and end mass levels**, respectively.
# ## 3. Plot the tree to see the cluster hierarchy
plot = tree.plot()
plot[0].show()
# Clusters are represented by the vertical line segments in the dendrogram. The default is to plot the dendrogram on the **mass** scale, so that the lower endpoint of a cluster's branch is at its starting mass level and the upper endpoint is at its end mass level. The mass from of the LST dendrogram is typically more useful than plotting on the **density** scale because the mass scale always starts at 0 and ends at 1, while estimated density values are often extremely large or small.
#
# In this example, the dendrogram shows that there are two very well-separated clusters in our dataset, indicated by two distinct trunk clusters. One of the trunks has no splits at all, indicating a very simple uni-modal distribution (this is not surprising, given that one of our synthetic clusters is a Gaussian ball). The other trunk splits repeatedly, indicating a complex hierarchical and multi-modal cluster structure. Within this structure there are two clear primary sub-clusters.
#
# The width of a cluster's branch indicates how many points belong to the cluster *when it first appears*. The dendrogram for this example shows (correctly) that the Gaussian blob has far less mass than the concentric circles.
# ## 4. Prune the tree
pruned_tree = tree.prune(60)
pruned_tree.plot()[0].show()
# Our LST is not as useful as it should be because there are many very small leaves; some have only a single point. **Pruning the tree** recursively merges clusters from the leaves downward, until every cluster has at least some minimum size. The `prune` method is computationally cheap and returns a new tree, so this can be done repeatedly until the tree is most useful for you. In this example we set the minimum cluster size to be 60 points, which results in a much simpler tree.
#
# We could have also pruned immediately by setting the `prune_threshold` parameter in our original `construct_tree` call. A good strategy is to set the `prune_threshold` to be the same as `k` in that constructor, then increase the prune threshold later if there are too many high-density clusters for your task.
# ## 5. Get clusters from the tree
# +
cluster_labels = pruned_tree.get_clusters()
print "Cluster labels shape:", cluster_labels.shape
# -
# There are many ways to use the LST to assign data points to clusters. **The default method** of the `get_clusters` method is to **return the points in the leaves** of the tree, which are the highest-density modal regions of the dataset. The big advantage of this method is that the user doesn't need to know the right number of clusters beforehand, or even to choose a single density level.
#
# All clustering methods return a numpy matrix with two columns. The first column contains indices of data points and the second contains the integer label of a given point's cluster.
#
# In general, the clusters obtained from the LST **exclude low-density data points** (the specific pattern of which points to exclude depends on the clustering strategy). In our example above, the cluster labels include only 493 of the original 600 data points.
# ## 6. Plot the clusters in feature space
# +
upper_level_idx = cluster_labels[:, 0]
upper_level_set = X[upper_level_idx, :]
with plt.style.context('ggplot'):
fig, ax = plt.subplots(figsize=(6, 4.5))
ax.scatter(upper_level_set[:, 0], upper_level_set[:, 1],
c=cluster_labels[:, 1], s=70, alpha=0.9)
fig.show()
| examples/getting_started.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Project: Charity ML
# In the code block below we load all necessary packages and modules for this project. In addition, the data is loaded, the categorical features are converted to the 'category' data-type, and the top 5 observations of data are displayed to get a sense of what the data looks like.
#
#
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as pl
import matplotlib.patches as mpatches
import warnings
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler, OrdinalEncoder
from xgboost import XGBClassifier
from sklearn.base import clone
from time import time
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier
from sklearn.metrics import fbeta_score, accuracy_score, make_scorer, f1_score
from sklearn.tree import DecisionTreeClassifier
from sklearn.svm import SVC
from sklearn.model_selection import GridSearchCV
from sklearn.exceptions import DataConversionWarning
#warningTypes = [DataConversionWarning,UserWarning,FutureWarning,UndefinedMetricWarning]
warnings.filterwarnings(action='ignore', category=DataConversionWarning)
warnings.filterwarnings(action='ignore', category=UserWarning)
warnings.filterwarnings(action='ignore', category=FutureWarning)
data = pd.read_csv('/Users/jaredthacker/Udacity/IntroMchnLrng/Project1/census.csv',header=0)
categoricalFeatures = ['workclass','education_level','marital-status','occupation','relationship',
'race','sex','native-country','income']
for i in categoricalFeatures:
data[i] = data[i].astype('category')
data.head()
# -
data.describe()
# ### The following cell is for displaying each of the data-types for each of the features and the 'income' variable. In addition, I give the max and min value for each column/variable.
# +
variables = ['age','workclass','education_level','education-num','marital-status',
'occupation','relationship','race','sex','capital-gain','capital-loss',
'hours-per-week','native-country','income']
for i in variables:
print('The data-type for {}'.format(i),'is: {}'.format(data[i].dtype.name))
print('--------------------------------------')
print('The minimum of all the census data is:',data.min(numeric_only=True))
print('--------------------------------------')
print('The maximum of all the census data is:',data.max(numeric_only=True))
# -
# ### In the following 3 code-cells, the histograms for the numerical features are given so that we can get a sense of the distribution of each of these variables.
histo = data.hist(column=['age','capital-gain','capital-loss','education-num','hours-per-week'])
capGainHist = data['capital-gain'].hist(bins=15)
capLossHist = data['capital-loss'].hist(bins=15)
# ### In the code block below we answer all questions in the "Exploring the Data" section
print('The total number of records is:',data['income'].shape[0])
gt50K = len(data[data['income'] =='>50K'])
ltet50K = len(data[data['income'] == '<=50K'])
print('The number of individuals with an income of greater than $50k is:',gt50K)
print('The number of individulas with an income of $50k or less is:',ltet50K)
percGt50K = gt50K/(ltet50K + gt50K)
print('The percentage of individuals with an income of greater than $50k is:',percGt50K)
# ## In the following code-cell, the census dataset is split into the features and the response and then the 'capital-gain' and 'capital-loss' features are log-tranformed.
# +
X_raw = data.iloc[:,:-1]
y = data.iloc[:,-1]
skewed = ['capital-gain','capital-loss']
X_log_transformed = pd.DataFrame(data=X_raw)
X_log_transformed[skewed] = X_raw[skewed].apply(lambda x: np.log(x+1))
logTransCG = X_log_transformed['capital-gain'].hist()
logTransCL = X_log_transformed['capital-loss'].hist()
y.head()
# +
scaler = MinMaxScaler()
numerical = ['age', 'education-num', 'capital-gain', 'capital-loss', 'hours-per-week']
X_log_minmax_transform = pd.DataFrame(data = X_log_transformed)
X_log_minmax_transform[numerical] = scaler.fit_transform(X_log_transformed[numerical])
display(X_log_minmax_transform.head(n = 5))
print(X_log_minmax_transform.min(numeric_only=True))
print(X_log_minmax_transform.max(numeric_only=True))
# -
# In the code block below we apply one-hot encoding to the <i>census</i> data.
categoricalFeatures = ['workclass','education_level','marital-status','occupation','relationship',
'race','sex','native-country']
features = pd.get_dummies(X_log_minmax_transform,columns=categoricalFeatures)
y = OrdinalEncoder().fit_transform(np.asarray(y).reshape(-1,1))
y = pd.DataFrame(y)
y.columns = ['income']
features.head()
# +
X_train, X_test, y_train, y_test = train_test_split(features,
y,
test_size = 0.2,
random_state = 42)
print("Training set has {} samples.".format(X_train.shape[0]))
print("Testing set has {} samples.".format(X_test.shape[0]))
# -
# # Question 1:
# ## Below we calculate the performance of the baseline Naive Bayes predictor
# +
y_1Hot = pd.get_dummies(data['income'])
y_1Hot.head()
TP = y.sum()[0]
FN = 0
FP = y.shape[0] - TP
accuracy = TP/(TP + FP)
recall = TP/(TP + FN)
precision = TP/(TP + FP)
F_Score = (1 + (0.5)**2) * precision * recall/((((0.5)**2) * precision) + recall)
print('The accuracy of the baseline Naive Bayes predictor is {}'.format(accuracy))
print('The recall of the baseline Naive Bayes predictor is {}'.format(recall))
print('The precision of the baseline Naive Bayes predictor is {}'.format(precision))
print('The F_Score of the baseline Naive Bayes predictor is {}'.format(F_Score))
# -
# # Question 2:
#
# I chose to use decision trees, adaptive boosting and naive bayes. The answers for each model is given below:
#
# ## Decision Trees:
# Decision trees: A supervised-learning model that can be applied to any regression or classification problem. The applications are endless, but one application could be predicting success given certain information/features about each student.
#
# ### Strengths:
# Decision trees are computationally inexpensive and are thus preferred when processing power is limited. In addition, decision trees are very interpretable which makes them useful when an inference is needed. In fact, it has been argued that decision trees mirror human decision making.
#
# Decision trees can be visually displayed, and are thus, even more easy to interpret than say, linear regression.
#
# Decision trees can easily handle qualitative features without the need to create dummy variables.
#
# ### Weaknesses:
# Decision trees have a high degree of variance and are thuse sensitive outliers and even slight changes to the data.
#
# Decision trees do not have the same predictive power that other supervised methods have.
#
# ### Candidate potential:
# Decision trees are a good baseline model to attempt. As long as measures are taken to avoid overfitting, decisions tree make for a good initial fit to the dataset.
#
# ## Adaptive Boosting (with decision tree as the base estimator):
#
# ### Strengths:
# Adaptive boosting improves the predictive power of decision trees.
#
# Boosting tends to fit to the data slowly, which makes boosting overfit much less than decision trees.
#
# ### weaknesses:
# Adaptive boosting is not interpretable, and cannot be used for inference.
#
# ### Candidate potential:
# Boosting algorithms have been the gold standard for a while now. As long as we are not interested inference and are more interested in predictive power boosting should usually at least be tried once.
#
# ## Naive Bayes:
#
# ### Strengths:
# Ideal for predicting classes for both binary- and multi-class problems.
#
# In the case where the independent assumption among features assumption holds, the method can be quite accurate.
#
# ### Weaknesses:
# Bad predictive power for regression problems.
#
# The independent features assumption is usually not practical in real life.
#
# ### Candidate potential:
# Since the proble we are studying for charityML is a binary-classification problem, naive bayes would at least make a good bench-mark model.
# +
clf_A = AdaBoostClassifier()
clf_A.fit(X_train,y_train)
y_pred = clf_A.predict(X_test)
accuracy_score(y_test,y_pred)
fbeta_score(y_test,y_pred,beta=0.5)
# -
print(y_train.shape)
print(X_train.shape)
# +
clf_B = DecisionTreeClassifier()
clf_B.fit(X_train,y_train)
y_pred = clf_B.predict(X_test)
accuracy_score(y_test,y_pred)
fbeta_score(y_test,y_pred,beta=0.5)
# +
clf_C = GaussianNB()
clf_C.fit(X_train,y_train)
y_pred = clf_C.predict(X_test)
fbeta_score(y_test,y_pred,beta=0.5)
# -
def train_predict_pipeline(learner,sample_size,X_train,y_train,X_test,y_test):
results = {}
start = time()
learner.fit(X_train.iloc[:sample_size,:],y_train.iloc[:sample_size])
end = time()
results['train_time'] = end - start
start = time()
y_pred = learner.predict(X_test)
y_fit = learner.predict(X_train)
end = time()
results['pred_time'] = end - start
results['acc_train'] = accuracy_score(y_train[:300],y_fit[:300])
results['acc_test'] = accuracy_score(y_test,y_pred)
results['f_train'] = fbeta_score(y_train[:300],y_fit[:300],beta=0.5)
results['f_test'] = fbeta_score(y_test,y_pred,beta=0.5)
print("{} trained on {} samples.".format(learner.__class__.__name__, sample_size))
return results
# +
samples_100 = round(y_train.shape[0] * 1.0)
samples_10 = round(y_train.shape[0] * 0.10)
samples_1 = round(y_train.shape[0] * 0.010)
results = {}
for clf in [clf_A, clf_B, clf_C]:
clf_name = clf.__class__.__name__
results[clf_name] = {}
for i, samples in enumerate([samples_1, samples_10, samples_100]):
results[clf_name][i] = \
train_predict_pipeline(clf, samples, X_train, y_train, X_test, y_test)
train_predict_pipeline(clf,samples,X_train,y_train,X_test,y_test)
# -
def evaluate(results, accuracy, f1):
"""
Visualization code to display results of various learners.
inputs:
- learners: a list of supervised learners
- stats: a list of dictionaries of the statistic results from 'train_predict()'
- accuracy: The score for the naive predictor
- f1: The score for the naive predictor
"""
# Create figure
fig, ax = pl.subplots(2, 3, figsize = (11,7))
# Constants
bar_width = 0.3
colors = ['#A00000','#00A0A0','#00A000']
# Super loop to plot four panels of data
for k, learner in enumerate(results.keys()):
for j, metric in enumerate(['train_time', 'acc_train', 'f_train', 'pred_time', 'acc_test', 'f_test']):
for i in np.arange(3):
# Creative plot code
ax[j//3, j%3].bar(i+k*bar_width, results[learner][i][metric], width = bar_width, color = colors[k])
ax[j//3, j%3].set_xticks([0.45, 1.45, 2.45])
ax[j//3, j%3].set_xticklabels(["1%", "10%", "100%"])
ax[j//3, j%3].set_xlabel("Training Set Size")
ax[j//3, j%3].set_xlim((-0.1, 3.0))
# Add unique y-labels
ax[0, 0].set_ylabel("Time (in seconds)")
ax[0, 1].set_ylabel("Accuracy Score")
ax[0, 2].set_ylabel("F-score")
ax[1, 0].set_ylabel("Time (in seconds)")
ax[1, 1].set_ylabel("Accuracy Score")
ax[1, 2].set_ylabel("F-score")
# Add titles
ax[0, 0].set_title("Model Training")
ax[0, 1].set_title("Accuracy Score on Training Subset")
ax[0, 2].set_title("F-score on Training Subset")
ax[1, 0].set_title("Model Predicting")
ax[1, 1].set_title("Accuracy Score on Testing Set")
ax[1, 2].set_title("F-score on Testing Set")
# Add horizontal lines for naive predictors
ax[0, 1].axhline(y = accuracy, xmin = -0.1, xmax = 3.0, linewidth = 1, color = 'k', linestyle = 'dashed')
ax[1, 1].axhline(y = accuracy, xmin = -0.1, xmax = 3.0, linewidth = 1, color = 'k', linestyle = 'dashed')
ax[0, 2].axhline(y = f1, xmin = -0.1, xmax = 3.0, linewidth = 1, color = 'k', linestyle = 'dashed')
ax[1, 2].axhline(y = f1, xmin = -0.1, xmax = 3.0, linewidth = 1, color = 'k', linestyle = 'dashed')
# Set y-limits for score panels
ax[0, 1].set_ylim((0, 1))
ax[0, 2].set_ylim((0, 1))
ax[1, 1].set_ylim((0, 1))
ax[1, 2].set_ylim((0, 1))
# Create patches for the legend
patches = []
for i, learner in enumerate(results.keys()):
patches.append(mpatches.Patch(color = colors[i], label = learner))
pl.legend(handles = patches, bbox_to_anchor = (-.80, 2.53), \
loc = 'upper center', borderaxespad = 0., ncol = 3, fontsize = 'x-large')
# Aesthetics
pl.suptitle("Performance Metrics for Three Supervised Learning Models", fontsize = 16, y = 1.10)
pl.tight_layout()
pl.show()
# # Question 3
# The Adaptive Boosting model produces the highest F-scores and Accuracies for among all 3 models that were used. One aspect that the Ada Boost model did not perform well on is the time it took to fit to the training set. However, this should not be an issue since the data set is pretty small. Since this is a binary-classification problem where many of the features are categorical , Ada Boost with a decision tree base estimator should be fine.
evaluate(results,accuracy,F_Score)
# # Question 4:
# Ada Boost works by fitting a single "weak" base-model, i.e, a decision tree to the training data. After this initial base-model is fit, we then fit a second base-model that prioritizes correcting the prediction error of the first base-model. We repeat this process a fixed number of times and combine the outputs of all of these individual base models to make future predictions on unknown data.
# +
clf = AdaBoostClassifier()
parameters = {'n_estimators':[100,200,400],'learning_rate':[0.1,0.01,0.001]}
scorer= make_scorer(fbeta_score,beta=0.5)
grid_obj = GridSearchCV(clf, parameters, scoring = scorer)
grid_fit = grid_obj.fit(X_train,y_train)
best_clf = grid_fit.best_estimator_
clf.fit(X_train,y_train)
predictions = clf.predict(X_test)
best_predictions = best_clf.predict(X_test)
print("Unoptimized model\n------")
print("Accuracy score on testing data: {:.4f}".format(accuracy_score(y_test, predictions)))
print("F-score on testing data: {:.4f}".format(fbeta_score(y_test, predictions, beta = 0.5)))
print("\nOptimized Model\n------")
print("Final accuracy score on the testing data: {:.4f}".format(accuracy_score(y_test, best_predictions)))
print("Final F-score on the testing data: {:.4f}".format(fbeta_score(y_test, best_predictions, beta = 0.5)))
# -
# # Question 5
#
# #### Results:
#
# | Metric | Unoptimized Model | Optimized Model |
# | :------------: | :---------------: | :-------------: |
# | Accuracy Score | 0.8607 | 0.8598 |
# | F-score | 0.7491 | 0.7495 |
#
# The metrics for the optimized and unoptimized model are approximately the same. This can be fixed by using grid search over a larger portion of the parameter space, this will be done as an exercise to post on my github at a future date.
#
# The F-score for the Ada-Boost model was roughly 0.8598 whereas the F-score for the benchmark Naive-Bayes was approximately 0.2920. Ada-Boost provides a dramatic increase in performance over the benchmark method.
# # Question 6:
# If we go off just simply using intuition, then my guess for the top predicitive features would be:
#
# 1) Age (Generally people make more money as they age and receive promotions etc)
#
# 2) Education Level (Job-wise income is usually correlated with education level)
#
# 3) Number of hours-per-week you work (Gives a good indication of a person's work ethic)
#
# 4) Working Class (People in the private sector usually make more than the government sector)
#
# 5) Marital-Status
importances = clf_B.feature_importances_
def feature_plot(importances, X_train, y_train):
# Display the five most important features
indices = np.argsort(importances)[::-1]
columns = X_train.columns.values[indices[:5]]
values = importances[indices][:5]
# Creat the plot
fig = pl.figure(figsize = (9,5))
pl.title("Normalized Weights for First Five Most Predictive Features", fontsize = 16)
pl.bar(np.arange(5), values, width = 0.6, align="center", color = '#00A000', \
label = "Feature Weight")
pl.bar(np.arange(5) - 0.3, np.cumsum(values), width = 0.2, align = "center", color = '#00A0A0', \
label = "Cumulative Feature Weight")
pl.xticks(np.arange(5), columns)
pl.xlim((-0.5, 4.5))
pl.ylabel("Weight", fontsize = 12)
pl.xlabel("Feature", fontsize = 12)
pl.legend(loc = 'upper center')
pl.tight_layout()
pl.show()
feature_plot(importances, X_train, y_train)
# # Question 7:
# I correctly predicted 3/5 of the most important features, although they were not in the correct order. In the future, I would like to use more rigorous methods like the Chi-square test to determine if categorical features are correlated with the response and also use scatter matrices. The visualization confirms my thoughts by showing that marial-status, age, and hours-per-week are the features with the largest weights when predicting the response, income, in the decision tree.
# +
from sklearn.base import clone
X_train_reduced = X_train[X_train.columns.values[(np.argsort(importances)[::-1])[:5]]]
X_test_reduced = X_test[X_test.columns.values[(np.argsort(importances)[::-1])[:5]]]
clf = (clone(best_clf)).fit(X_train_reduced, y_train)
reduced_predictions = clf.predict(X_test_reduced)
print("Final Model trained on full data\n------")
print("Accuracy on testing data: {:.4f}".format(accuracy_score(y_test, best_predictions)))
print("F-score on testing data: {:.4f}".format(fbeta_score(y_test, best_predictions, beta = 0.5)))
print("\nFinal Model trained on reduced data\n------")
print("Accuracy on testing data: {:.4f}".format(accuracy_score(y_test, reduced_predictions)))
print("F-score on testing data: {:.4f}".format(fbeta_score(y_test, reduced_predictions, beta = 0.5)))
# -
# # Question 8:
# The final model for the full vs the reduced data set is approximately the same. If the charityML dataset were huge, I would absolutely use the reduced set with on the 5 most important features.
| project_1/.ipynb_checkpoints/project1-JaredThacker-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="MW70Bu_eKZxY" colab_type="text"
# #Scaling and Centering data
# + id="ToUGecYHKfni" colab_type="code" colab={}
from sklearn.preprocessing import StandardScaler
steps= [('scaler', StandardScaler()),
('knn', KNeighborsClassifier())]
pipeline= Pipeline(steps)
parameters= {knn_n_neighbors= np.arrange(1,50)}
X_train, X_test, y_train, y_test= train_test_split(X, y, test_size=0.2, random_state=21)
cv= GridSearchCV(pipeline, param_grid=parameters)
# + [markdown] id="85W-gl-xN9Nx" colab_type="text"
# #Exercise 1
# + id="AqfrvVONODuM" colab_type="code" colab={}
# Import scale
from sklearn.preprocessing import scale
# Scale the features: X_scaled
X_scaled = scale(X)
# Print the mean and standard deviation of the unscaled features
print("Mean of Unscaled Features: {}".format(X.mean()))
print("Standard Deviation of Unscaled Features: {}".format(X.std()))
# Print the mean and standard deviation of the scaled features
print("Mean of Scaled Features: {}".format(X_scaled.mean()))
print("Standard Deviation of Scaled Features: {}".format(X_scaled.std()))
# + [markdown] id="fB7aNFGgOOIW" colab_type="text"
# #Exercise 2
# + id="WjxBbBJ2OQC5" colab_type="code" colab={}
# Import the necessary modules
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
# Setup the pipeline steps: steps
steps = [('scaler', StandardScaler()),
('knn', KNeighborsClassifier())]
# Create the pipeline: pipeline
pipeline = Pipeline(steps)
# Create train and test sets
X_train, X_test, y_train, y_test = train_test_split(X,
y, test_size=0.3, random_state=42)
# Fit the pipeline to the training set: knn_scaled
knn_scaled = pipeline.fit(X_train, y_train)
# Instantiate and fit a k-NN classifier to the unscaled data
knn_unscaled = KNeighborsClassifier().fit(X_train, y_train)
# Compute and print metrics
print('Accuracy with Scaling: {}'.format(knn_scaled.score(X_test, y_test)))
print('Accuracy without Scaling: {}'.format(knn_unscaled.score(X_test,y_test)))
| DataCamp-Scikit-Learn/Scaling and Centering data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <font color=darkblue>Week 9 Assignment </font>
# ## Full name:
# ## R#:
# ## Title of the notebook:
# ## Date:
#  <br>
#
# ### Exercise 1: From a normally distributed population, we randolmy took a sample of 200 dogs with a mean weight of 70 pounds. Suppose the standard deviation of the population is 20: <br>
# ### What is the estimated true population mean for the 95% confidence interva? <br>
# ### How about 90% confidence interval? <br>
# ### How about 99% confidence interval? <br>
# ### Exercise 1: Amazon is considering changing the color of their logo. The smile will be green instead of orange!
#  <br>
#
# #### Let us assume out of 5000 users, they have directed 2500 to site A with the previous logo, and the rest to site B with the new logo. In the first group, 1863 users made a purchase. In the second group, 1904 users made a purchase. Is this a statistically significant result? Should Amazon change their logo in order to make more sells?
# 
| 2-Homework/Spring2021/ES-9/Week 9 Assignment_Dev.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''tf'': conda)'
# metadata:
# interpreter:
# hash: 31d56fa8489417c6958bc95e73b8d3766816132fea40e62e4bcabc0084854ff7
# name: python3
# ---
# # Toy Datasets
#
# Spravime si generatory dat, nad ktorymi budeme vypracovavat demonstracie.
#
# ## Circle dataset
#
# Vygenerujeme vzorky s dvoma hodnotami (nx=2), ktore budeme klasifikovat do 2 tried, podla toho, ci ich vzdialenost od stredu je vacsia ako hranicny polomer r.
#
import numpy as np
import matplotlib.pyplot as plt
def dataset_Circles(m=10, radius=0.7, noise=0.0, verbose=False):
# Hodnoty X budu v intervale <-1; 1>
X = (np.random.rand(2, m) * 2.0) - 1.0
if (verbose): print('X: \n', X, '\n')
# Element-wise nasobenie nahodnym sumom
N = (np.random.rand(2, m)-0.5) * noise
if (verbose): print('N: \n', N, '\n')
Xnoise = X + N
if (verbose): print('Xnoise: \n', Xnoise, '\n')
# Spocitame polomer
# Element-wise druha mocnina
XSquare = Xnoise ** 2
if (verbose): print('XSquare: \n', XSquare, '\n')
# Spocitame podla prvej osi. Ziskame (1, m) array.
RSquare = np.sum(XSquare, axis=0, keepdims=True)
if (verbose): print('RSquare: \n', RSquare, '\n')
R = np.sqrt(RSquare)
if (verbose): print('R: \n', R, '\n')
# Y bude 1, ak je polomer vacsi ako argument radius
Y = (R > radius).astype(float)
if (verbose): print('Y: \n', Y, '\n')
# Vratime X, Y
return X, Y
# +
# Spravime ukazkovy dataset
X, Y = dataset_Circles(m=5, radius=0.5, noise=0.0, verbose=True)
# Skusime z X a Y ziskat vlastnosti datasetu
nx, m = X.shape
ny, _ = Y.shape
print('Vlastnosti datasetu: ')
print(' nx = ', nx)
print(' ny = ', ny)
print(' m = ', m)
print('')
# -
# ## Vizualizacia datasetu
#
# Je dobre si vediet vizualizovat data, s ktorymi pracujeme. Pouzijeme PyPlot kniznicu a spravime SCATTER graf.
def draw_Scatter(X, Y):
# 9 x 9 palcovy obrazok s linearnou mierkou
plt.figure(figsize=(9, 9))
plt.xscale('linear')
plt.yscale('linear')
# Nakreslime scatter graf
plt.scatter(X[0,:], X[1,:], c=Y, cmap=plt.cm.RdBu)
plt.show()
plt.close()
# +
# Spravime ukazkovy dataset a vizualizujeme ho
X, Y = dataset_Circles(m=10000, radius=0.8, noise=0.0, verbose=False)
draw_Scatter(X, Y)
# -
# ## Kvetinkovy vzor (Flower pattern)
#
# Druhy hrackarsky dataset bude kreslit vzor pripominajuci lupene kvetu.
#
def dataset_Flower(m=10, noise=0.0):
# Inicializujeme matice
X = np.zeros((m, 2), dtype='float')
Y = np.zeros((m, 1), dtype='float')
a = 1.0
pi = 3.141592654
M = int(m/2)
for j in range(2):
ix = range(M*j, M*(j+1))
t = np.linspace(j*pi, (j+1)*pi, M) + np.random.randn(M)*noise
r = a*np.sin(4*t) + np.random.randn(M)*noise
X[ix] = np.c_[r*np.sin(t), r*np.cos(t)]
Y[ix] = j
X = X.T
Y = Y.T
return X, Y
# +
# Ukazka Flower datasetu
X, Y = dataset_Flower(m=5000, noise=0.0)
draw_Scatter(X, Y)
# -
# ## Batch vs. MiniBatch
#
# Spravme si este jednu pomocnu funkciu, ktorou budeme vediet ovladat velkost davok, po ktorych sa data budu spracovavat. Touto funkciou budeme vediet dosiahnut vsetky 3 moznosti:
# - spracovat cely dataset v jednom kroku
# - spracovavat dataset po mensich davkach
# - spracovavat dataset jednotlivo po vzorkach
def MakeBatches(dataset, batchSize):
# Set obsahuje 2 mnoziny - X, Y
X, Y = dataset
# Zistime celkovy pocet vzoriek
nx, m = X.shape
ny, _ = Y.shape
# Vysledny zoznam
result = []
# Ak je batchSize = 0, berieme celu mnozinu
if (batchSize <= 0):
batchSize = m
# Celkovy pocet davok sa zaokruhluje nahor
steps = int(np.ceil(m / batchSize))
for i in range(steps):
# Spocitame hranice rezu
mStart = i * batchSize
mEnd = min(mStart + batchSize, m)
# Vyberame data pre aktualny rez - chceme dodrzat rank
minibatchX = X[:,mStart:mEnd]
minibatchY = Y[:,mStart:mEnd]
assert(len(minibatchX.shape) == 2)
assert(len(minibatchY.shape) == 2)
# Pridame novu dvojicu do vysledneho zoznamu
result.append((minibatchX, minibatchY))
return result
# +
# Vyskusajme, ci MakeBatches funguje spravne
trainSet = dataset_Circles(m=8)
# Spravime tri verzie z datasetu
setComplete = MakeBatches(trainSet, 0)
setThree = MakeBatches(trainSet, 3)
setOne = MakeBatches(trainSet, 1)
print('Complete set: \n', setComplete, '\n')
print('Batches with size 3: \n', setThree, '\n')
print('Batches with size 1: \n', setOne, '\n')
# -
# ## Vizualizacia hranice rozhodovania (decision boundary)
#
# Filled Contour graf, ktory vizualizuje vysledok funkcie (modelu, neuronovej siete) pre vsetky kombinacie, ktore mozu nadobudat vstupne data X1, X2.
def draw_DecisionBoundary(X, Y, model):
# Najdeme hranice, pre ktore budeme skumat predikciu
pad = 0.5
x1_Min, x1_Max = X[0,:].min()-pad, X[0,:].max()+pad
x2_Min, x2_Max = X[1,:].min()-pad, X[1,:].max()+pad
# Spravime mriezku dvojic - vzorkujeme cely interval <MIN; MAX> s granularitou h
h = 0.01
x1_Grid, x2_Grid = np.meshgrid(
np.arange(x1_Min, x1_Max, h),
np.arange(x2_Min, x2_Max, h)
)
# Usporiadame si mriezku hodnot do rovnakeho tvaru ako ma X
XX = np.c_[x1_Grid.ravel(), x2_Grid.ravel()].T
# Vypocitame predikciu pomocou modelu na vsetky hodnoty mriezky
YHat = model(XX)
# A usporiadame si vysledok tak, aby sme ho mohli podhodit PyPlotu
YHat = YHat.reshape(x1_Grid.shape)
# Najskor nakreslime contour graf - vysledky skumania pre mriezku
plt.figure(figsize=(9, 9))
plt.xscale('linear')
plt.yscale('linear')
plt.contourf(x1_Grid, x2_Grid, YHat, cmap=plt.cm.RdYlBu)
# Potom este pridame scatter graf pre X, Y
plt.scatter(X[0,:], X[1,:], c=Y, cmap=plt.cm.RdBu)
plt.show()
plt.close()
# +
def testModel(X):
# Vstup je tvaru (2, m)
# Vystup bude tvaru (1, m)
x1 = X[0, None]
x2 = X[1, None]
# Zatial iba jednoducha ukazkova funkcia
yhat = (x1 > x2).astype(float)
return yhat
# Spravime ukazkovy dataset a vizualizujeme ho
X, Y = dataset_Circles(m=500, radius=0.8, noise=0.0, verbose=False)
# Kreslime decision boundary
draw_DecisionBoundary(X, Y, testModel)
# -
| lecture_2/helpers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework: Paint Estimator
#
#
# ## The Problem
#
#
# The big-box hardware store *House Depot* has contracted you to write an App to estimate the amount of paint required to paint a room. Given that 1 gallon of paint covers 400 square feet, you must output the number of 1 gallon paint cans required to paint a rectangular room.
#
# The program should input:
#
# - the length of the room in feet,
# - the width of the room in feet,
# - the height of the room in feet,
# - number of coats of paint (how many times will you go over the walls? 1, 2, 3, etc...)
#
# The program should output:
#
# - total area to be painted (sum of each of the 4 walls, multiplied by the height of the room)
# - the number of gallons required to paint the room with the number of coats of paint.
#
# Example 1:
#
# ```
# Enter length of room: 12
# Enter width of room: 14
# Enter height of room: 8
# Enter number of coats: 2
# Total area to be painted: 416.00 sqft
# 2 Coats requires: 832.00 sqft
# 1 Can of paint covers 400 sqft
# Total gallons of paint requried for 2 coats is : 3 cans
# ```
#
# Example 2:
#
# ```
# Enter length of room: 2
# Enter width of room: 4
# Enter height of room: 10
# Enter number of coats: 3
# Total area to be painted: 120.00 sqft
# 3 Coats requires: 360.00 sqft
# 1 Can of paint covers 400 sqft
# Total gallons of paint requried for 3 coats is : 1 cans
# ```
#
# HINTS:
#
# - Follow the same problem solving approaches from small group: Understanding the problem by example.
# - Round up to the nearest whole can of paint.
# - Code exactly one of the examples then change to accept inputs.
#
#
# + [markdown] label="problem_analysis_cell"
# ## Part 1: Problem Analysis
#
# Inputs:
#
# ```
# TODO: Inputs
# ```
#
# Outputs:
#
# ```
# TODO: Outputs
# ```
#
# Algorithm (Steps in Program):
#
# ```
# TODO:Steps Here
#
# ```
# -
# ## Part 2: Code Solution
#
# You may write your code in several cells, but place the complete, final working copy of your code solution within this single cell below. Only the within this cell will be considered your solution.
# + label="code_solution_cell"
# Step 2: Write code here
# + [markdown] label="homework_questions_cell"
# ## Part 3: Questions
#
# 1. Why does the program still run when you enter a negative number for length?
#
# `--== Double-Click and Write Your Answer Below This Line ==--`
#
#
# 2. Does the output make sense when you enter a negative length? What type of error is this?
#
#
# `--== Double-Click and Write Your Answer Below This Line ==--`
#
#
# 3. Why do we use `math.ceil()` in this program? In other words are you allowed to buy 3.75 gallons of paint?
#
# `--== Double-Click and Write Your Answer Below This Line ==--`
#
#
#
# + [markdown] label="reflection_cell"
# ## Part 4: Reflection
#
# Reflect upon your experience completing this assignment. This should be a personal narrative, in your own voice, and cite specifics relevant to the activity as to help the grader understand how you arrived at the code you submitted. Things to consider touching upon: Elaborate on the process itself. Did your original problem analysis work as designed? How many iterations did you go through before you arrived at the solution? Where did you struggle along the way and how did you overcome it? What did you learn from completing the assignment? What do you need to work on to get better? What was most valuable and least valuable about this exercise? Do you have any suggestions for improvements?
#
# To make a good reflection, you should journal your thoughts, questions and comments while you complete the exercise.
#
# Keep your response to between 100 and 250 words.
#
# `--== Double-Click and Write Your Reflection Below Here ==--`
#
# -
# run this code to turn in your work!
from coursetools.submission import Submission
Submission().submit()
| lessons/02-Variables/HW-Variables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
import scipy
import sklearn
import sys
data = pd.read_csv('CC.csv')
data
data.columns
data.shape
data.describe()
data = data.sample(frac = 0.1, random_state = 1)
data.shape
data.hist(figsize =(20,20))
plt.show()
# +
Fraud = data[data['Class'] == 1]
Valid =data[data['Class'] == 0]
outlier_fraction = len(Fraud)/len(Valid)
outlier_fraction
# -
print('Fraud Cases: {}'.format(len(Fraud)))
print('Valid Cases: {}'.format(len(Valid)))
# +
corrmat = data.corr()
fig = plt.figure(figsize = (12, 9))
sns.heatmap(corrmat, vmax = 0.8, square = True)
plt.show()
# +
columns = data.columns.tolist()
columns = [c for c in columns if c not in ["Class"]]
target = 'Class'
X = data[columns]
Y = data[target]
print(X.shape)
print(Y.shape)
# +
from sklearn.metrics import classification_report,accuracy_score
from sklearn.ensemble import IsolationForest
from sklearn.neighbors import LocalOutlierFactor
state = 1
classifiers = {
"Isolation Forest": IsolationForest(max_samples=len(X),
contamination=outlier_fraction,
random_state=state),
"Local Outlier Factor": LocalOutlierFactor(
n_neighbors=20,
contamination=outlier_fraction)}
# +
plt.figure(figsize=(9, 7))
n_outliers = len(Fraud)
for i, (clf_name, clf) in enumerate(classifiers.items()):
if clf_name == "Local Outlier Factor":
y_pred = clf.fit_predict(X)
scores_pred = clf.negative_outlier_factor_
else:
clf.fit(X)
scores_pred = clf.decision_function(X)
y_pred = clf.predict(X)
y_pred[y_pred == 1] = 0
y_pred[y_pred == -1] = 1
n_errors = (y_pred != Y).sum()
print('{}: {}'.format(clf_name, n_errors))
print(accuracy_score(Y, y_pred))
print(classification_report(Y, y_pred))
# -
| Credit Card Fraud Detection/Credit-Card-Fraud-Detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# > Texto fornecido sob a Creative Commons Attribution license, CC-BY. Todo o código está disponível sob a FSF-approved BSD-3 license.<br>
# > (c) Original por <NAME>, <NAME> em 2017, traduzido por <NAME> em 2020.<br>
# > [@LorenaABarba](https://twitter.com/LorenaABarba) - [@fschuch](https://twitter.com/fschuch)
# 12 passos para Navier-Stokes
# ======
# ***
# Você experimentou escolher diferentes parêmetros numéricos nos passos [1](./01_Passo_1.ipynb) e [2](./02_Passo_2.ipynb)? Se a resposta é sim, você provavelmente encontrou algum comportamento inesperado. Alguma vez sua resolução explodiu? (Na minha experiência, estudantes de CFD *amam* fazer as coisas explodirem).
#
# Você provavelmente está se perguntando porque mudar os parâmetros de discretização afetam a solução de maneira tão drástica. Esse notebook é um complemento para nosso [curso CFD interativo](https://github.com/fschuch/CFDPython-BR), onde discutimos a condição CFL.
# Convergência e Condição CFL
# ----
# ***
# Para alguns dos primeiros passos, nós temos usado a mesma condição inicial e mesmas condições de contorno. Com os parâmetros que sugerimos inicialmente, a malha com 41 pontos e um passo de tempo de 0,025 segundos. Agora, vamos experimentar incrementar o tamanho da nossa malha. O código abaixo é idêntico ao que usamos no [Passo 1](./01_Passo_1.ipynb), mas aqui ele foi empacotado dentro de uma função, então podemos facilmente verificar o que acontece se ajustamos uma variável: **o tamanho da malha**.
# + jupyter={"outputs_hidden": false}
import numpy #Aqui carregamos numpy
from matplotlib import pyplot #Aqui carregamos matplotlib
# %matplotlib inline
def linearconv(nx):
nt = 20 #Número de passos de tempo que queremos calcular
dt = .025 #Tamanho de cada passo de tempo
c = 1 #Velocidade de propagação da onda
x = numpy.linspace(0., 2., num = nx)
nx = x.size
dx = x[1] - x[0]
u = numpy.ones_like(x) #Função ones_like do numpy
u[(0.5<=x) & (x<=1)] = 2 #Define u = 2 entre 0,5 e 1,
#de acordo com nossa CI
un = numpy.ones_like(u) #Inicializar o arranjo temporário
for n in range(nt): #Laço temporal
un = u.copy() ##Cópia dos valores de u para un
for i in range(1, nx): ##Laço espacial
u[i] = un[i] - c * dt / dx * (un[i] - un[i-1])
pyplot.plot(x, u);
# -
# Vamos examinar os resultados do nosso problema de convecção linear à medida que refinamos a malha.
# + jupyter={"outputs_hidden": false}
linearconv(41) #Convecção usando 41 pontos na malha
# -
# O obtemos o mesmo resultado que calculamos no Passo 1, reproduzido aqui for referência.
# + jupyter={"outputs_hidden": false}
linearconv(61)
# -
# Aqui, ainda temos difusão numérica presente, embora menos severa.
# + jupyter={"outputs_hidden": false}
linearconv(71)
# -
# Aqui o mesmo padrão está presente: a onda é mais quadrada que nos testes anteriores.
# + jupyter={"outputs_hidden": false}
linearconv(85)
# -
# Isso não se parece em nada com a nossa função chapéu original.
# ### O que aconteceu?
# Para responder essa questão, temos que pensar um pouco sobre o que realmente implementamos no nosso código.
#
# Em cada interação do loop temporal, nós usamos os dados existentes sobre a nossa onda para estimar a velocidade da onda no passo de tempo subsequente. Inicialmente, o incremento no número de pontos retorna uma solução mais precisa. Existia menos difusão numérica e a onda quadrada foi parecendo muito mais como uma onda quadrada do que no exemplo inicial.
#
# Cada iteração no laço temporal cobre o comprimento do passo de tempo $\Delta t$, que acabamos definindo como 0,025.
#
# Durante essa iteração, calculamos a velocidade da onda para cada ponto que criamos ao longo de $x$. No último gráfico, alguma coisa claramente deu errado.
#
# O que aconteceu é que durante o último período $\Delta t$, a onda viajou uma distância que é maior que `dx`. O comprimento `dx` de cada fração da malha está relacionado ao número total de pontos `nx`, então pode-se fazer cumprir a estabilidade se o passo de tempo $\Delta t$ é calculado com respeito à resolução da malha `dx`:
#
# $$\sigma = \frac{u \Delta t}{\Delta x} \leq \sigma_{\max}$$
#
# onde $u$ é a velocidade da onda, $sigma$ é denominado como [número de Courant](https://en.wikipedia.org/wiki/Courant%E2%80%93Friedrichs%E2%80%93Lewy_condition) e o valor de $\sigma_{\max}$ que vai garantir a estabilidade depende do tipo de discretização utilizada.
#
# Em uma nova versão do nosso código, vamos usar o número CFL para calcular o passo de tempo `dt` apropriado, em função do comprimento `dx`.
# + jupyter={"outputs_hidden": false}
import numpy
from matplotlib import pyplot
def linearconv(nx):
nt = 20 #Número de passos de tempo que queremos calcular
c = 1 #Velocidade de propagação da onda
sigma = .5
x = numpy.linspace(0., 2., num = nx)
nx = x.size
dx = x[1] - x[0]
dt = sigma * dx
u = numpy.ones_like(x)
u[(0.5<=x) & (x<=1)] = 2
un = numpy.ones_like(u)
for n in range(nt): #iterate through time
un = u.copy() ##copy the existing values of u into un
for i in range(1, nx):
u[i] = un[i] - c * dt / dx * (un[i] - un[i-1])
pyplot.plot(x, u)
# + jupyter={"outputs_hidden": false}
linearconv(41)
# + jupyter={"outputs_hidden": false}
linearconv(61)
# + jupyter={"outputs_hidden": false}
linearconv(81)
# + jupyter={"outputs_hidden": false}
linearconv(101)
# + jupyter={"outputs_hidden": false}
linearconv(121)
# -
# Note que conforme o número de pontos `nx` aumenta, a onda viaja por uma distância cada vez menor. O número de iterações temporais pelas quais a solução avança foi mantido constante em `nt = 20`. Entretanto, dependendo do valor de `nx` e o correspondente valor de `dx` e `dt`, uma janela temporal cada vez menor tem sido examinada, afinal de contas.
# Leia o material complementar, ou avance para o [Passo 3](./04_Passo_3.ipynb).
# Material Complementar
# -----
# ***
# É possível fazer uma análise rigososa sobre a estabilidade do esquema numérico, em alguns casos. Assista a apresentação da Prof. Barba sobre esse assunto em **Video Lecture 9**, no Youtube.
# + jupyter={"outputs_hidden": false}
from IPython.display import YouTubeVideo
YouTubeVideo('Yw1YPBupZxU')
# + jupyter={"outputs_hidden": false}
from IPython.core.display import HTML
def css_styling():
styles = open("../styles/custom.css", "r").read()
return HTML(styles)
css_styling()
# -
# > A célula acima executa o estilo para esse notebook. Nós modificamos o estilo encontrado no GitHub de [CamDavidsonPilon](https://github.com/CamDavidsonPilon), [@Cmrn_DP](https://twitter.com/cmrn_dp).
| tarefas/03_Condicao_CFL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# <div class="contentcontainer med left" style="margin-left: -50px;">
# <dl class="dl-horizontal">
# <dt>Title</dt> <dd> Scatter3D Element</dd>
# <dt>Dependencies</dt> <dd>Matplotlib</dd>
# <dt>Backends</dt> <dd><a href='../matplotlib/Image.ipynb'>Matplotlib</a></dd>
#
# </dl>
# </div>
import numpy as np
import holoviews as hv
hv.extension('matplotlib')
# ``Scatter3D`` represents three-dimensional coordinates which may be colormapped or scaled in size according to a value. They are therefore very similar to [``Points``](Points.ipynb) and [``Scatter``](Scatter.ipynb) types but have one additional coordinate dimension. Like other 3D elements the camera angle can be controlled using ``azimuth``, ``elevation`` and ``distance`` plot options:
# %%opts Scatter3D [azimuth=40 elevation=20 color_index=2] (s=50 cmap='fire')
y,x = np.mgrid[-5:5, -5:5] * 0.1
heights = np.sin(x**2+y**2)
hv.Scatter3D(zip(x.flat,y.flat,heights.flat))
# Just like all regular 2D elements, ``Scatter3D`` types can be overlaid and will follow the default color cycle:
#
# %%opts Scatter3D [size_index='Size' scaling_factor=5] (marker='^')
hv.Scatter3D(np.random.randn(100,4), vdims='Size') * hv.Scatter3D(np.random.randn(100,4)+2, vdims='Size')
# For full documentation and the available style and plot options, use ``hv.help(hv.Scatter3D).``
| examples/reference/elements/matplotlib/Scatter3D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This code displays a plot of the ion velocity from the Omni and Artemis data before filtering out invalid values. This data is not usable in its current state. 2019-08-01T00:08:17.788Z 2019-08-31T23:58:45.303Z
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
fig = plt.figure()
fig.suptitle("Omni and Artemis velocity data")
om = pd.read_csv("OnmiForArtemis.csv")
ar = pd.read_csv("ArtemisSolarWind.csv")
ar['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'] = pd.to_datetime(ar['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'])
om['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'] = pd.to_datetime(om['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'])
om = om.drop(['HELIOGRAPHIC_LATITUDE_deg', 'HELIOGRAPHIC_LONGITUDE_deg', 'ION_DENSITY_N_cm3'], axis=1)
om = om.loc[:, ~om.columns.str.contains('^Unnamed')]
ar = ar.drop(['Ion_Density_n_cc'], axis=1)
ar.rename(columns={'Ion_Velocity_km_s':'Artemis Velocity kms'}, inplace=True)
om.rename(columns={'BULK_FLOW_SPEED_km_s':'OMNI Velocity kms'}, inplace=True)
ar.head()
ax = plt.gca()
ar.plot(kind='line', x='EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ', y='Artemis Velocity kms', color='red', ax=ax)
om.plot(kind='line', x='EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ', y='OMNI Velocity kms', ax=ax)
ax.set_xlim([datetime.date(2019, 8, 1), datetime.date(2019, 9, 1)])
plt.show()
# -
# Here is the data after filtering out the invalid values. As you can see this data looks much more meaningful, but there are still some inconsistencies with the Artemis data.
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import datetime
fig = plt.figure()
fig.suptitle("Omni and Artemis velocity data")
om = pd.read_csv("OnmiForArtemis.csv")
ar = pd.read_csv("ArtemisSolarWind.csv")
ar['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'] = pd.to_datetime(ar['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'])
om['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'] = pd.to_datetime(om['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'])
om = om[om.ION_DENSITY_N_cm3 != -1.0E+31]
om = om.drop(['HELIOGRAPHIC_LATITUDE_deg', 'HELIOGRAPHIC_LONGITUDE_deg', 'ION_DENSITY_N_cm3'], axis=1)
om = om.loc[:, ~om.columns.str.contains('^Unnamed')]
ar = ar[ar.Ion_Density_n_cc != -1.0E+31]
ar = ar.drop(['Ion_Density_n_cc'], axis=1)
ar.rename(columns={'Ion_Velocity_km_s':'Artemis Velocity kms'}, inplace=True)
om.rename(columns={'BULK_FLOW_SPEED_km_s':'OMNI Velocity kms'}, inplace=True)
ar.head()
ax = plt.gca()
ar.plot(kind='line', x='EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ', y='Artemis Velocity kms', color='red', ax=ax)
om.plot(kind='line', x='EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ', y='OMNI Velocity kms', ax=ax)
ax.set_xlim([datetime.date(2019, 8, 2), datetime.date(2019, 8, 3)])
plt.show()
# -
# Here is the code and plot for resampling the Artemis data based on hourly averages. This means that the individual data points can be matched up with the data points from the OMNI dataset.
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
fig = plt.figure()
fig.suptitle("Omni and Artemis velocity data")
om = pd.read_csv("OnmiForArtemis.csv")
ar = pd.read_csv("ArtemisSolarWind.csv")
ar['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'] = pd.to_datetime(ar['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'])
om['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'] = pd.to_datetime(om['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'])
om = om[om.ION_DENSITY_N_cm3 != -1.0E+31]
om = om.drop(['HELIOGRAPHIC_LATITUDE_deg', 'HELIOGRAPHIC_LONGITUDE_deg', 'ION_DENSITY_N_cm3'], axis=1)
om = om.loc[:, ~om.columns.str.contains('^Unnamed')]
ar = ar[ar.Ion_Density_n_cc != -1.0E+31]
ar = ar.drop(['Ion_Density_n_cc'], axis=1)
ar.rename(columns={'Ion_Velocity_km_s':'Artemis Velocity kms'}, inplace=True)
om.rename(columns={'BULK_FLOW_SPEED_km_s':'OMNI Velocity kms'}, inplace=True)
ar = ar.resample('H', on = 'EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ').mean().reset_index()
ax = plt.gca()
ar.plot(kind='line', x='EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ', y='Artemis Velocity kms', color='red', ax=ax)
om.plot(kind='line', x='EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ', y='OMNI Velocity kms', ax=ax)
ax.set_xlim([datetime.date(2019, 8, 1), datetime.date(2019, 9, 1)])
plt.show()
# -
# This data does not match up exactly, but it's possible that the position of the satelite in comparison to Artemis could explain some of these inconsistencies. Here is a narrow slice of the data, which should help validate our naive approach for the time shift.
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
fig = plt.figure()
fig.suptitle("Omni and Artemis velocity data")
om = pd.read_csv("OnmiForArtemis.csv")
ar = pd.read_csv("ArtemisSolarWind.csv")
ar['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'] = pd.to_datetime(ar['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'])
om['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'] = pd.to_datetime(om['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'])
om = om[om.ION_DENSITY_N_cm3 != -1.0E+31]
om = om.drop(['HELIOGRAPHIC_LATITUDE_deg', 'HELIOGRAPHIC_LONGITUDE_deg', 'ION_DENSITY_N_cm3'], axis=1)
om = om.loc[:, ~om.columns.str.contains('^Unnamed')]
ar = ar[ar.Ion_Density_n_cc != -1.0E+31]
ar = ar.drop(['Ion_Density_n_cc'], axis=1)
ar.rename(columns={'Ion_Velocity_km_s':'Artemis Velocity kms'}, inplace=True)
om.rename(columns={'BULK_FLOW_SPEED_km_s':'OMNI Velocity kms'}, inplace=True)
ar = ar.resample('H', on = 'EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ').mean().reset_index()
ax = plt.gca()
ar.plot(kind='line', x='EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ', y='Artemis Velocity kms', color='red', ax=ax)
om.plot(kind='line', x='EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ', y='OMNI Velocity kms', ax=ax)
ax.set_xlim([datetime.date(2019, 8, 1), datetime.date(2019, 8, 7)])
plt.show()
# -
# There are still some inconsistencies with the data, but the majority of these are occuring at a rate of 1 per day. This could be indicitive of the moon passing behind the Earth and being shielded from the solar wind somewhat.
# Here are some plots generated from the data from Maven from 08/01/2015 to 09/01/2015. This data does not look as good, which may be because the velocity had to be recomposed from 3 seperate vectors. We also attempted to use the quality data provided for the Maven velocity components, but the result was the same.
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import datetime
fig = plt.figure()
fig.suptitle("Omni and Maven velocity data")
om = pd.read_csv("OmniForMaven.csv")
ma = pd.read_csv("MavenVelocityWithQuality.csv")
om['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'] = pd.to_datetime(om['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'])
om = om[om.ION_DENSITY_N_CC != -1.0E+31]
om = om.drop(['HELIOGRAPHIC_LATITUDE_deg', 'HELIOGRAPHIC_LONGITUDE_deg', 'ION_DENSITY_N_CC'], axis=1)
om = om.loc[:, ~om.columns.str.contains('^Unnamed')]
ma['EPOCH__yyyy-mm-ddThh:mm:ss.sssZ'] = pd.to_datetime(ma['EPOCH__yyyy-mm-ddThh:mm:ss.sssZ'])
ma = ma[ma.Xquality == 1]
ma = ma[ma.Xquality != -1.0E+31]
ma['BULK_FLOW_VELOCITY_km_s'] = np.linalg.norm(ma[['Xvelocity','Yvelocity','Zvelocity']].values,axis=1)
ma = ma[ma.BULK_FLOW_VELOCITY_km_s < 800]
ma = ma.drop(['Xvelocity', 'Yvelocity', 'Zvelocity', 'Xquality', 'Yquality', 'Zquality'], axis=1)
ma = ma.resample('H', on = 'EPOCH__yyyy-mm-ddThh:mm:ss.sssZ').mean().reset_index()
ma.rename(columns={'BULK_FLOW_VELOCITY_km_s':'Maven Velocity kms'}, inplace=True)
om.rename(columns={'BULK_FLOW_SPEED_km_s':'OMNI Velocity kms'}, inplace=True)
ax = plt.gca()
ma.plot(kind='line', x='EPOCH__yyyy-mm-ddThh:mm:ss.sssZ', y='Maven Velocity kms', color='red', ax=ax)
om.plot(kind='line', x='EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ', y='OMNI Velocity kms', ax=ax)
ax.set_xlim([datetime.date(2015, 8, 1), datetime.date(2015, 9, 1)])
plt.show()
ma.describe()
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
fig = plt.figure()
fig.suptitle("Artemis Merged data")
ar = pd.read_csv("ArtemisMerged.csv")
ar['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'] = pd.to_datetime(ar['EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ'])
ar = ar[ar.Ion_Density_n_cc != -1.0E+31]
ar = ar.resample('H', on = 'EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ').mean().reset_index()
ar.plot(kind='line', x='EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ', y='Ion_Velocity_km_s', color='red')
ar.plot(kind='line', x='EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ', y='Ion_Density_n_cc', color='red')
ar.plot(kind='line', x='EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ', y='Distance_From_Sun_AU', color='red')
ar.plot(kind='line', x='EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ', y='HGI_LAT_deg', color='red')
ar.plot(kind='line', x='EPOCH_TIME_yyyy-mm-ddThh:mm:ss.sssZ', y='HGI_LON_deg', color='red')
plt.show()
# -
| Data Processing/Plotting Stuff.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Removes bot accounts from the originally annotated dataset
# +
import tweepy
from tweepy import OAuthHandler
import pandas as pd
def get_api(consumer_key, consumer_secret, access_token, access_secret):
auth = OAuthHandler(consumer_key, consumer_secret)
auth.set_access_token(access_token, access_secret)
return tweepy.API(auth, wait_on_rate_limit=True, wait_on_rate_limit_notify=True, compression=True)
# +
key = ['<KEY>',
'<KEY>',
'<KEY>',
'<KEY>']
api = get_api(key[0], key[1], key[2], key[3])
# -
df = pd.read_csv("BotClassificationResults.csv")
df.head()
df['bot'].value_counts()
screen_names = []
for i in df.itertuples(index=True, name='Pandas'):
if i.bot == 1:
user = api.get_user(i.id)
screen_names.append(user.screen_name)
df = pd.read_csv('DownsampledCoreData.csv')
df.head()
for row in screen_names:
df = df[df.screenname != row]
df['migration_relevance'].value_counts()
df.to_csv('DownsampledCoreDataWithoutBots.csv')
| bot_removal.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Cifar10 Drift Detection
#
# In this example we will deploy an image classification model along with a drift detector trained on the same dataset. For in depth details on creating a drift detection model for your own dataset see the [alibi-detect project](https://github.com/SeldonIO/alibi-detect) and associated [documentation](https://docs.seldon.io/projects/alibi-detect/en/latest/). You can find details for this [CIFAR10 example in their documentation](https://docs.seldon.io/projects/alibi-detect/en/latest/examples/cd_ks_cifar10.html) as well.
#
#
# Prequisites:
#
# * Running cluster with
# * [kfserving installed](https://github.com/kubeflow/kfserving/blob/master/README.md)
# * [Knative eventing installed](https://knative.dev/docs/install/)
#
# !pip install -r requirements_notebook.txt
# ## Setup Resources
# Enabled eventing on default namespace. This will activate a default Knative Broker.
# !kubectl label namespace default knative-eventing-injection=enabled
# Create a Knative service to log events it receives. This will be the example final sink for outlier events.
# !pygmentize message-dumper.yaml
# !kubectl apply -f message-dumper.yaml
# Create the Kfserving image classification model for Cifar10. We add in a `logger` for requests - the default destination is the namespace Knative Broker.
# !pygmentize cifar10.yaml
# !kubectl apply -f cifar10.yaml
# Create the pretrained Drift Detector. We forward replies to the message-dumper we started. Notice the `drift_batch_size`. The drift detector will wait until `drify_batch_size` number of requests are received before making a drift prediction.
# !pygmentize cifar10cd.yaml
# !kubectl apply -f cifar10cd.yaml
# Create a Knative trigger to forward logging events to our Outlier Detector.
# !pygmentize trigger.yaml
# !kubectl apply -f trigger.yaml
# Get the IP address of the Istio Ingress Gateway. This assumes you have installed istio with a LoadBalancer.
CLUSTER_IPS=!(kubectl -n istio-system get service istio-ingressgateway -o jsonpath='{.status.loadBalancer.ingress[0].ip}')
CLUSTER_IP=CLUSTER_IPS[0]
print(CLUSTER_IP)
SERVICE_HOSTNAMES=!(kubectl get inferenceservice tfserving-cifar10 -o jsonpath='{.status.url}' | cut -d "/" -f 3)
SERVICE_HOSTNAME_CIFAR10=SERVICE_HOSTNAMES[0]
print(SERVICE_HOSTNAME_CIFAR10)
SERVICE_HOSTNAMES=!(kubectl get ksvc drift-detector -o jsonpath='{.status.url}' | cut -d "/" -f 3)
SERVICE_HOSTNAME_VAEOD=SERVICE_HOSTNAMES[0]
print(SERVICE_HOSTNAME_VAEOD)
# +
import matplotlib.pyplot as plt
import numpy as np
import requests
import json
import tensorflow as tf
tf.keras.backend.clear_session()
train, test = tf.keras.datasets.cifar10.load_data()
X_train, y_train = train
X_test, y_test = test
X_train = X_train.astype('float32') / 255
X_test = X_test.astype('float32') / 255
print(X_train.shape, y_train.shape, X_test.shape, y_test.shape)
classes = ('plane', 'car', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
def show(X):
plt.imshow(X.reshape(32, 32, 3))
plt.axis('off')
plt.show()
def predict(X):
formData = {
'instances': X.tolist()
}
headers = {}
headers["Host"] = SERVICE_HOSTNAME_CIFAR10
res = requests.post('http://'+CLUSTER_IP+'/v1/models/tfserving-cifar10:predict', json=formData, headers=headers)
if res.status_code == 200:
j = res.json()
if len(j["predictions"]) == 1:
return classes[np.array(j["predictions"])[0].argmax()]
else:
print("Failed with ",res.status_code)
return []
def drift(X):
formData = {
'instances': X.tolist()
}
headers = {}
headers["Host"] = SERVICE_HOSTNAME_VAEOD
res = requests.post('http://'+CLUSTER_IP+'/', json=formData, headers=headers)
if res.status_code == 200:
od = res.json()
return od
else:
print("Failed with ",res.status_code)
return []
# -
# ## Normal Prediction
idx = 1
X = X_train[idx:idx+1]
show(X)
predict(X)
# ## Test Drift
# We need to accumulate a large enough batch size so no drift will be tested as yet.
# !kubectl logs $(kubectl get pod -l serving.knative.dev/configuration=message-dumper -o jsonpath='{.items[0].metadata.name}') user-container
# We will now send 5000 requests to the model in batches. The drift detector will run at the end of this as we set the `drift_batch_size` to 5000 in our yaml above.
from tqdm.notebook import tqdm
for i in tqdm(range(0,5000,100)):
X = X_train[i:i+100]
predict(X)
# Let's check the message dumper and extract the first drift result.
# res=!kubectl logs $(kubectl get pod -l serving.knative.dev/configuration=message-dumper -o jsonpath='{.items[0].metadata.name}') user-container
data= []
for i in range(0,len(res)):
if res[i] == 'Data,':
data.append(res[i+1])
j = json.loads(json.loads(data[0]))
print("Drift",j["data"]["is_drift"]==1)
# Now, let's create some CIFAR10 examples with motion blur.
from alibi_detect.datasets import fetch_cifar10c, corruption_types_cifar10c
corruption = ['motion_blur']
X_corr, y_corr = fetch_cifar10c(corruption=corruption, severity=5, return_X_y=True)
X_corr = X_corr.astype('float32') / 255
show(X_corr[0])
show(X_corr[1])
show(X_corr[2])
# Send these examples to the predictor.
for i in tqdm(range(0,5000,100)):
X = X_corr[i:i+100]
predict(X)
# Now when we check the message dump we should find a new drift response.
# res=!kubectl logs $(kubectl get pod -l serving.knative.dev/configuration=message-dumper -o jsonpath='{.items[0].metadata.name}') user-container
data= []
for i in range(0,len(res)):
if res[i] == 'Data,':
data.append(res[i+1])
j = json.loads(json.loads(data[1]))
print("Drift",j["data"]["is_drift"]==1)
# ## Tear Down
# !kubectl delete -f cifar10.yaml
# !kubectl delete -f cifar10cd.yaml
# !kubectl delete -f trigger.yaml
# !kubectl delete -f message-dumper.yaml
| docs/samples/drift-detection/alibi-detect/cifar10/cifar10_drift.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow_p27
# language: python
# name: conda_tensorflow_p27
# ---
# # Using Amazon Elastic Inference with a pre-trained TensorFlow Serving model on SageMaker
#
# This notebook demonstrates how to enable and use Amazon Elastic Inference with our predefined SageMaker TensorFlow Serving containers.
#
# Amazon Elastic Inference (EI) is a resource you can attach to your Amazon EC2 instances to accelerate your deep learning (DL) inference workloads. EI allows you to add inference acceleration to an Amazon SageMaker hosted endpoint or Jupyter notebook for a fraction of the cost of using a full GPU instance. For more information please visit: https://docs.aws.amazon.com/sagemaker/latest/dg/ei.html
#
# This notebook's main objective is to show how to create an endpoint, backed by an Elastic Inference, to serve our pre-trained TensorFlow Serving model for predictions. With a more efficient cost per performance, Amazon Elastic Inference can prove to be useful for those looking to use GPUs for higher inference performance at a lower cost.
#
# 1. [The model](#The-model)
# 1. [Setup role for SageMaker](#Setup-role-for-SageMaker)
# 1. [Load the TensorFlow Serving Model on Amazon SageMaker using Python SDK](#Load-the-TensorFlow-Serving-Model-on-Amazon-SageMaker-using-Python-SDK)
# 1. [Deploy the trained Model to an Endpoint with EI](#Deploy-the-trained-Model-to-an-Endpoint-with-EI)
# 1. [Using EI with a SageMaker notebook instance](#Using-EI-with-a-SageMaker-notebook-instance)
# 1. [Invoke the Endpoint to get inferences](#Invoke-the-Endpoint-to-get-inferences)
# 1. [Delete the Endpoint](#Delete-the-Endpoint)
#
# If you are familiar with SageMaker and already have a trained model, skip ahead to the [Deploy the trained Model to an Endpoint with an attached EI accelerator](#Deploy-the-trained-Model-to-an-Endpoint-with-an-attached-EI-accelerator)
#
# For this example, we will use the SageMaker Python SDK, which helps deploy your models to train and host in SageMaker. In this particular example, we will be interested in only the hosting portion of the SDK.
#
# 1. Set up our pre-trained model for consumption in SageMaker
# 2. Host the model in an endpoint with EI
# 3. Make a sample inference request to the model
# 4. Delete our endpoint after we're done using it
#
# ## The model
#
# The pre-trained model we will be using for this example is a NCHW ResNet-50 model from the [official Tensorflow model Github repository](https://github.com/tensorflow/models/tree/master/official/resnet#pre-trained-model). For more information in regards to deep residual networks, please check [here](https://github.com/tensorflow/models/tree/master/official/resnet). It isn't a requirement to train our model on SageMaker to use SageMaker for serving our model.
#
# SageMaker expects our models to be compressed in a tar.gz format in S3. Thankfully, our model already comes in that format. The predefined TensorFlow Serving containers use REST API for handling inferences, for more informationm, please see [Deploying to TensorFlow Serving Endpoints](https://github.com/aws/sagemaker-python-sdk/blob/master/src/sagemaker/tensorflow/deploying_tensorflow_serving.rst#making-predictions-against-a-sagemaker-endpoint).
#
# To host our model for inferences in SageMaker, we need to first upload the SavedModel to S3. This can be done through the AWS console or AWS command line.
#
# For this example, the SavedModel object will already be hosted in a public S3 bucket owned by SageMaker.
# +
# %%time
import boto3
# use the region-specific saved model object
region = boto3.Session().region_name
saved_model = (
"s3://sagemaker-sample-data-{}/tensorflow/model/resnet/resnet_50_v2_fp32_NCHW.tar.gz".format(
region
)
)
# -
# ## Setup role for SageMaker
#
# Let's start by creating a SageMaker session and specifying the IAM role arn used to give hosting access to your model. See the [documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/sagemaker-roles.html) for how to create these. Note, if more than one role is required for notebook instances, training, and/or hosting, please replace the `sagemaker.get_execution_role()` with a the appropriate full IAM role arn string(s).
# + isConfigCell=true
import sagemaker
role = sagemaker.get_execution_role()
# -
# ## Load the TensorFlow Serving Model on Amazon SageMaker using Python SDK
#
# We can use the SageMaker Python SDK to load our pre-trained TensorFlow Serving model for hosting in SageMaker for predictions.
#
# There are a few parameters that our TensorFlow Serving Model is expecting.
# 1. `model_data` - The S3 location of a model tar.gz file to load in SageMaker
# 2. `role` - An IAM role name or ARN for SageMaker to access AWS resources on your behalf.
# 3. `framework_version` - TensorFlow Serving version you want to use for handling your inference request .
#
# +
from sagemaker.tensorflow.serving import Model
tensorflow_model = Model(model_data=saved_model, role=role, framework_version="1.14")
# -
# ## Deploy the trained Model to an Endpoint with an attached EI accelerator
#
# The `deploy()` method creates an endpoint which serves prediction requests in real-time.
#
# The only change required for utilizing EI with our SageMaker TensorFlow Serving containers only requires providing an `accelerator_type` parameter, which determines which type of EI accelerator to attach to your endpoint. The supported types of accelerators can be found here: https://aws.amazon.com/sagemaker/pricing/instance-types/
#
# %%time
predictor = tensorflow_model.deploy(
initial_instance_count=1, instance_type="ml.m4.xlarge", accelerator_type="ml.eia1.medium"
)
# ## Using EI with a SageMaker notebook instance
#
# There is also the ability to utilize an EI accelerator attached to your local SageMaker notebook instance. For more information, please reference: https://docs.aws.amazon.com/sagemaker/latest/dg/ei-notebook-instance.html
# ## Invoke the Endpoint to get inferences
#
# Invoking prediction:
# +
# %%time
import numpy as np
random_input = np.random.rand(1, 1, 3, 3)
prediction = predictor.predict({"inputs": random_input.tolist()})
print(prediction)
# -
# ## Delete the Endpoint
#
# After you have finished with this example, remember to delete the prediction endpoint to release the instance(s) associated with it.
print(predictor.endpoint)
# +
import sagemaker
predictor.delete_endpoint()
| sagemaker-python-sdk/tensorflow_serving_using_elastic_inference_with_your_own_model/tensorflow_serving_pretrained_model_elastic_inference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:mnc]
# language: python
# name: conda-env-mnc-py
# ---
# # Starting and testing OVRO-LWA
# ### Setup
#
# 1. Use dashboard to confirm that services are up for etcd, bifrost pipelines, and data capture.
# 2. Identify server names for snap2, gpu, and data capture.
# 3. Set list of server names in notebook.
# 4. Execute cells
#
#
# ### F-eng
# * Repo https://github.com/realtimeradio/caltech-lwa
# * python install "control_sw"
#
# ### X-eng
# * Repo https://github.com/realtimeradio/caltech-bifrost-dsp
# * python install "pipeline-control"
#
# ### mnc
# * Repo https://github.com/ovro-lwa/mnc_python
# * python install "mnc_python"
snap2names = ['snap01'] # f-eng
feng_config = '/home/ubuntu/proj/caltech-lwa/control_sw/config/lwa_corr_config.yaml' # f-eng and x-eng
xhosts = ['lxdlwagpu02'] # x-eng
npipeline = 4 # x-eng
drips = ['10.41.0.25', '10.41.0.41'] # data recorder (two ips for lxdlwagpu09)
dest_ip = list(sorted(drips*2)) # correlator -> data recorder (destination IPs)
dest_port = [10001+i//npipeline for i in range(npipeline*len(xhosts))] # correlator -> data recorder (destination ports)
drroot = [f'drvs{ip.split(".")[-1]}' for ip in drips] # data recorder root names
drids = [f'{drr}{dp-10000:02}' for dp in set(dest_port) for drr in drroot] # data recorder ids
print(snap2names, xhosts, drips, dest_ip, dest_port, drids)
from lwa_f import snap2_fengine
from lwa352_pipeline_control import Lwa352CorrelatorControl
from mnc import mcs, common, ezdr
for snap2name in snap2names:
lwa_f = snap2_fengine.Snap2Fengine(snap2name)
lwa_f.cold_start_from_config(feng_config)
lwa_corr = Lwa352CorrelatorControl(xhosts, npipeline_per_host=npipeline)
lwa_corr.stop_pipelines() # maybe not needed?
lwa_corr.start_pipelines()
lwa_corr.configure_corr(dest_ip=dest_ip, dest_port=dest_port)
rs = ezdr.Lwa352RecorderControl('slow')
rs.start()
# ### Read data and visualize/test
# ## Stop
rs.stop()
lwa_corr.stop_pipelines()
| notebooks/LWA_startup_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from collections import Counter
# +
train_data = pd.read_csv('./combined-swell-classification-eda-train-dataset.csv')
test_data = pd.read_csv('./combined-swell-classification-eda-test-dataset.csv')
val_data = pd.read_csv('./combined-swell-classification-eda-validation-dataset.csv')
train_X = train_data
test_X = test_data
val_X = val_data
# train_Y, test_Y, val_Y
train_Y = train_X["condition"]
test_Y = test_X["condition"]
val_Y = val_X["condition"]
# +
# train_X
del train_X["NasaTLX class"]
del train_X["Condition Label"]
del train_X["NasaTLX Label"]
del train_X["condition"]
del train_X["subject_id"]
# test_X
del test_X["NasaTLX class"]
del test_X["Condition Label"]
del test_X["NasaTLX Label"]
del test_X["condition"]
del test_X["subject_id"]
# val_X
del val_X["NasaTLX class"]
del val_X["Condition Label"]
del val_X["NasaTLX Label"]
del val_X["condition"]
del val_X["subject_id"]
# -
class Node:
def __init__(self):
self.split_feature = None
self.split_point = None
self.result = None
self.childs = None
class DecisionTree:
def __init__(self):
self.root = None
def DT_print(self, cur_node = None, cnt = 0):
if cnt == 0:
cur_node = self.root
print(' ' * cnt, "Level ", cnt," :: ", cur_node.split_feature, cur_node.split_point, cur_node.result)
if cur_node.childs is None:
return
for child in cur_node.childs:
self.DT_print(child, cnt + 1)
def Gini_Impurity(self, data_Y): # input label dataset of a group
impurity = 1
label_counts = Counter(data_Y)
for label in label_counts:
p_of_label = label_counts[label] / len(data_Y)
impurity -= p_of_label ** 2
return impurity
def Information_Gain(self, unsplited_data_Y, splited_data_Y):
gain = self.Gini_Impurity(unsplited_data_Y)
# print(gain)
# print(splited_data_Y)
for subset in splited_data_Y:
# print("-", Gini_Impurity(subset), " X ( ", len(subset), " / ", len(unsplited_data_Y), " )")
gain -= self.Gini_Impurity(subset) * (len(subset)/ len(unsplited_data_Y))
return gain
def Split(self, data_X, data_Y, column):
data_X_subsets = [] # 분할 후의 data_X 그룹을 저장하는 배열
data_Y_subsets = [] # 분할 후의 data_Y 그룹을 저장하는 배열
split_point = 0.0 # 최적 분할 기준 값
split_point_gain = 0.0 # 최적 분할 지점에서의 Information Gain
'''
print("=-=- Before reset Index =-=-=-")
print("**** data_X ****")
print(data_X[column])
print("**** data_Y ****")
print(data_Y)
'''
data_X = data_X.sort_values(by=column)
data_Y = data_Y[data_X.index]
data_X = data_X.reset_index(drop=True)
data_Y = data_Y.reset_index(drop=True)
'''
print("=-=- After reset Index =-=-=-")
print(data_X.index)
print(data_X[column])
print(data_Y)
print(data_X[column].iloc[5], data_Y[5])
'''
for i in range(1, len(data_Y)):
candidate_splited_data_X = []
candidate_splited_data_Y = []
if data_Y[i-1] != data_Y[i]:
# print(i, data_Y[i-1], data_Y[i])
candidate_point = (data_X[column].iloc[i-1] + data_X[column].iloc[i]) / 2
candidate_splited_data_Y.append(data_Y[:i])
candidate_splited_data_Y.append(data_Y[i:])
gain = self.Information_Gain(data_Y, candidate_splited_data_Y)
if gain > split_point_gain:
candidate_splited_data_X.append(data_X[:i])
candidate_splited_data_X.append(data_X[i:])
split_point = candidate_point
split_point_gain = gain
data_X_subsets = candidate_splited_data_X
data_Y_subsets = candidate_splited_data_Y
# print("== Updated :: ", split_point_gain, split_point)
return split_point_gain, split_point, data_X_subsets, data_Y_subsets
def Find_Best_Split(self, data_X, data_Y):
# print("=-=-New Group=-=-")
best_feature = '' # 데이터를 분할 할 feature
best_gain = 0.0 # 데이터를 특정 feature로 분할했을 때 가장 높게 측정된 Information Gain
best_split_point = 0.0
for column in data_X.columns: # RF에서 Bagging Features 적용 필요.
# print("check column :: ", column)
gain, split_point = self.Split(data_X, data_Y, column)[0:2]
if gain > best_gain:
best_gain = gain
best_feature = column
best_split_point = split_point
return best_feature, best_gain, best_split_point
def fit(self, data_X, data_Y, cnt=0):
root = Node()
data_X = data_X.reset_index(drop=True)
data_Y = data_Y.reset_index(drop=True)
best_feature, best_gain, best_split_point = self.Find_Best_Split(data_X, data_Y)
if best_gain == 0:
root.result = data_Y[0]
# print(' ' * cnt, "== No Split ", cnt," :: ", root.result)
return root
data_X_subsets, data_Y_subsets = self.Split(data_X, data_Y, best_feature)[2:]
# print(' ' * cnt, "== Split ", cnt," :: ", best_feature, best_gain)
childs = []
for i in range(len(data_X_subsets)):
childs.append(self.fit(data_X_subsets[i], data_Y_subsets[i], cnt+1))
root.split_feature = best_feature
root.split_point = best_split_point
root.childs = childs
if cnt == 0:
self.root = root
return root
def predict(self, dataset):
result = []
for i in dataset.index:
cur_node = self.root
while cur_node.result is None:
value = dataset.loc[i, cur_node.split_feature]
if value < cur_node.split_point:
cur_node = cur_node.childs[0]
else :
cur_node = cur_node.childs[1]
result.append(cur_node.result)
return result
# +
newDT = DecisionTree()
print(newDT)
newDT.fit(test_X.head(100), test_Y.head(100))
# -
newDT.DT_print()
dataset = test_X.loc[0:10]
newDT.predict(dataset)
test_Y[0:10]
| Decision Tree Implementation From Scratch in Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 13.1 Interfacing Between pandas and Model Code
# The point of contact between pandas and other analysis libraries is usually NumPy arrays.<mark> To turn a DataFrame into a NumPy array, use the `.values` property.</mark>
# +
import pandas as pd
import numpy as np
data = pd.DataFrame({
'x0': [1,2,3,4,5],
'x1': [0.01, 0.02, 0.03, 0.04, 0.05],
'x2': [-1.5, 0., 3.6, 1.3, -2.]
})
data
# -
data.columns
data.values
# To convert back to DataFrame
df2 = pd.DataFrame(data.values, columns=['one', 'two', 'three'])
df2
# The `.values` attribute is intended to be used when you data is homogeneous-for example, all numeric type. If you have heterogeneous data, the result will be an ndarray of Python objects.
df3 = data.copy()
df3['strings'] = ['a', 'b', 'c', 'd', 'e']
df3
df3.values
# To use subset of columns/rows; use `.loc`.
model_cold = ['x0', 'x1']
data.loc[:, model_cold].values
# Suppose we had a nonumeric column in our dataset.
data['category'] = pd.Categorical(['a', 'b', 'a', 'a', 'b'], categories=['a', 'b'])
data
# +
# We might need to add dummy category varaible
dummies = pd.get_dummies(data.category, prefix='category')
data_with_dummies = data.drop('category', axis=1).join(dummies)
data_with_dummies
# -
# # 13.2 Creating Model Description with Patsy
# Patsy is a Python library for describing statistical models (especially linear models) with a small string-based "formula syntax"
# ### `patsy.dmatrices`
# Tkaes a formula string along with a dataset (which can be a DataFrame or a dict) and produces design matrices for a linear model.
# +
data = pd.DataFrame({
'x0': [1, 2, 3, 4, 5],
'x1': [0.01, -0.01, 0.25, -4.1, 0.],
'y': [-1.5, 0., 3.6, 1.3, -2.]
})
data
# +
import patsy
y, X = patsy.dmatrices('y ~ x0 + x1', data)
y
# -
X
# These Patsy `DesignMatrix` instances are NumPy ndarrays with additional metadata:
np.asarray(y)
np.asarray(X)
# `Intercept` term is a convension for linear models like Ordinary least squares(OLS) regression. You can suppress this by adding a `0` to the model.
patsy.dmatrices('y ~ x0 + x1 + 0', data)[1]
# Patsy objects can be passed directly into algorithm like `numpy.linalg.lstsq`, which performs an ordinary least squares regression.
coef, resid, _, _ = np.linalg.lstsq(X, y, rcond=None)
coef
# The <mark>model metadata is retained in</mark> `design_info` attribute, so you can reattach the model column names to the fitted coefficients to obatain a Series.
coef = pd.Series(coef.squeeze(), index=X.design_info.column_names)
coef
# ## Data Transformation in Pasty Formulas
# You can mix Python code into your Patsy Formulas;
y, X = patsy.dmatrices('y ~ x0 + np.log(np.abs(x1) + 1)', data)
X
# Patsy has built-in functions for some common used varaible transformations like **standardizing** <mark>(to mean 0 and variance 1)</mark> and **centering** <mark>(substracting the mean)</mark>.
y, X = patsy.dmatrices('y ~ standardize(x0) + center(x1)', data)
X
# The `patsy.build_design_matrices` function can <mark>apply transformations to new *out-of-sample* data usng the saved information from the original *in-sample* dataset.</mark>
# +
new_data = pd.DataFrame({
'x0': [6, 7, 8, 9],
'x1': [4.1, -0.5, 0., 2.3],
'y': [1, 2, 3, 4]
})
new_X = patsy.build_design_matrices([X.design_info], new_data)
new_X
# -
# Because the plus symbol in the context of Patsy Formula does not mean addition when you want to add columns from a dataset by name, <mark>you must wrap them in the special $I$ function.</mark>
y, X = patsy.dmatrices('y ~ I(x0 + x1)', data)
X
# Patsy has several other built-in transforms in the `patsy.builtins` module. See the online documentation.
# ## Categorical Data and Patsy
# When you use non-numeric terms in a Patsy formula,<mark> they are converted to dummy varaibles by default.</mark> If there is an intercept, one of the levels will be left out to avoid collinearity.
# +
data = pd.DataFrame({
'key1': ['a', 'a', 'b', 'b', 'a', 'b', 'a', 'b'],
'key2': [0, 1, 0, 1, 0, 1, 0, 0],
'v2': [-1, 0, 2.5, -0.5, 4.0, -1.2, 0.2, -1.7]
})
y, X = patsy.dmatrices('v2 ~ key1', data)
X
# -
# <mark>If you omit the intercept</mark> from the model, <mark>then columns for each category value will be included</mark> in the model design matrix:
y, X = patsy.dmatrices('v2 ~ key1 + 0', data)
X
# <mark>Numeric columns</mark> can be <mark>interpreted as categorical</mark> with `C` function:
y, X = patsy.dmatrices('v2 ~ C(key2)', data)
X
# You can <mark>include interaction terms of the form`key1:key2`</mark>, which can be used in say, <mark>analysis of variance (ANOVA)</mark> models:
data['key2'] = data['key2'].map({0: 'zero', 1: 'one'})
data
y, X = patsy.dmatrices('v2 ~ key1 + key2', data)
X
y, X = patsy.dmatrices('v2 ~ key1 + key2 + key1:key2', data)
X
# # 13.2 Introduction to statsmodels
# statsmodels is a Python library for fitting many kinds of statistical models, performing statistical test, and data exploration and visualisation.
#
# Next up, we will use a few basic tools in statsmodels and explore how to use the modeling interfaces with Patsy formulas and pandas DataFrame objects.
# ### Estimating Linear Models
# There are several kinds of linear models in statsmodels, and have <mark>two different main interface</mark>:
# 1. **Array-based**
# 2. **Formula-based**
import statsmodels.api as sm
import statsmodels.formula.api as smf
# To show how to use these, we generate a linear model from some random data:
#
# **Note:**
# 1. Inside a **function header**
# - `*` collects all the poisitional arguments in a tuple.
#
# - `**` collects all the keyword arguments in a dictionary.
# - `>>> def functionA(*a, **kw):
# print(a)
# print(kw)`
#
# `>>> functionA(1, 2, 3, a=2, b=3, c=5)
# (1, 2, 3)
# {'a': 2, 'b': 3, 'c': 5}`
# 2. Inside a **function call**
# - `*` unpacks a list or tuple into position arguments.
# - `**` unpacks a dictionary into keyword arguments.
# - `>>> lis = [1, 2, 3]`
#
# `>>> dic = {'a': 10, 'b': 20}`
#
# `>>> functionA(*lis, **dic)
# (1, 2, 3)
# {'a': 10, 'b': 20}`
#
# +
# "true" model with known parameters beta
def dnorm(mean, variance, size=1):
if isinstance(size, int):
size=size,
return mean + np.sqrt(variance) * np.random.randn(*size)
# For reproducibility
np.random.seed(1234)
N = 100
X = np.c_[dnorm(0, 0.4, size=N),
dnorm(0, 0.6, size=N),
dnorm(0, 0.2, size=N)]
eps = dnorm(0, 0.1, size=N)
beta = [0.1, 0.3, 0.5]
y = np.dot(X, beta) + eps
# -
# In this case,`dnorm` is a helper function for <mark>generating normally distributed data with particular mean and variance.</mark>
X[:5]
# A linear model is generally fitted with an intercept term as we saw before with Patsy.
# ### `sm.add_constant`
# Function can <mark>add an intercept column to an existing matrix.</mark>
X_model = sm.add_constant(X)
X_model[:5]
# The `sm.OLS` class can <mark>fit an ordinary least squares linear regression</mark>
model = sm.OLS(y, X)
# The model's `fit` method returns a regression results object containing estimated model parameters and other diagnostics:
results = model.fit()
results.params
# The `summary` method on `results` can <mark>print a model detailing diagnostic output of the model</mark>:
print(results.summary())
# The parameter here have been given the generic names. What if instead that all of the model parameter are in a DataFrame?
data = pd.DataFrame(X, columns=['col0', 'col1', 'col2'])
data['y'] = y
data[:5]
# Now we can use the statsmodels formula API and Patsy formula strings:
results = smf.ols('y ~ col0 + col1 + col2', data=data).fit()
results.params
results.tvalues
# Observe how statsmodels has returned results as Series with DataFrame names attached. We calso don't need to use `add_constant` when using formulas and pandas objects.
#
# Given new out-of-sample data, you can compute predicted values given the estimated model parameters.
results.predict(data[:5])
# ## Estimating Time Series Processes
# let's simulate some time series data with an autoregressive structure and noise:
# +
import random
init_x = 4
values = [init_x, init_x]
N = 1000
b0 = 0.8
b1 = -0.4
noise = dnorm(0, 0.1, N)
for i in range(N):
new_x = values[-1] * b0 + values[-2] * b1 + noise[i]
values.append(new_x)
# -
# This data has an <mark>AR(2) structure (two *lags*)</mark> with <mark>parameter 0.8 and 0.4</mark>. When you fit an AR model, you may not know the number of lagged terms to include, so you can fit the model with some larger number of flags.
MAXFLAGS = 5
model = sm.tsa.AR(values)
results = model.fit(MAXFLAGS)
results.params
| pydata/Ch 13 Introduction to Modeling Libraries in Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # The VESIcal web app
#
# Here we use Jupyter Notebook and voila to render a webapp for running VESIcal calculations on an input file describing a dataset.
import VESIcal as v
import io
import pandas as pd
import ipywidgets
from IPython.display import display, FileLink, FileLinks
# # Upload your file
# +
import ipywidgets as widgets
# ipywidgets file uploader button
uploader = widgets.FileUpload()
display(uploader)
# set custom layout for widgets
layout = widgets.Layout(width='auto', height='40px') #set width and height
# allow too long of descriptions
style = {'description_width': 'initial'}
# -
# # Calculate Saturation Pressures
# +
# create widgets so the user can choose model options
# Model selection
model_selector_satP = widgets.Dropdown(
options=[('MagmaSat', 'MagmaSat'),
('Dixon', 'Dixon'),
('Shishkina', 'ShishkinaIdealMixing'),
('IaconoMarziano', 'IaconoMarziano'),
('Liu', 'Liu'),
('MooreWater', 'MooreWater'),
('AllisonCarbon (Vesuvius)', 'AllisonCarbon_vesuvius'),
('AllisonCarbon (Sunset)', 'AllisonCarbon_sunset'),
('AllisonCarbon (SFVF)', 'AllisonCarbon_sfvf'),
('AllisonCarbon (Erebus)', 'AllisonCarbon_erebus'),
('AllisonCarbon (Stromboli)', 'AllisonCarbon_stromboli')
],
value='MagmaSat',
description='Model:',
)
# Temperature selection
temperature_selector_satP = widgets.Text(
value='',
placeholder='Temperature (degC)',
description='Temperature:',
disabled=False,
style=style
)
# Temperature type check box if user wants to pass temp from file
temperature_selection_checkbox_satP = widgets.Checkbox(value=False,
description='Use temperature from file (if checked, put column name in Temperature box)',
disabled=False,
indent=False,
layout=layout)
display(model_selector_satP,
temperature_selector_satP,
temperature_selection_checkbox_satP)
# Create a button that, when clicked, gets name of file uploaded and
# creates a BatchFile object and runs the model calculation
button_satP = widgets.Button(
description='Calculate Saturation Pressures',
disabled=False,
button_style='', # 'success', 'info', 'warning', 'danger' or ''
tooltip='Calculate Saturation Pressures',
icon='check', # (FontAwesome names without the `fa-` prefix)
layout=layout
)
button_satP.style.button_color = 'lightgreen'
output = widgets.Output()
display(button_satP, output)
def on_button_clicked_satP(b):
with output:
# print successful click
print("Button clicked.")
# get filename, content of file and create BatchFile object
uploaded_filename = next(iter(uploader.value))
content = uploader.value[uploaded_filename]['content']
with open('myfile', 'wb') as f:
f.write(content)
myfile = v.BatchFile(content)
print(model_selector_satP.value)
# sanitize inputs
if temperature_selection_checkbox_satP.value is True:
temperature = str(temperature_selector_satP.value)
else:
temperature = float(temperature_selector_satP.value)
# run model calculation
result = myfile.calculate_saturation_pressure(temperature=temperature,
model=model_selector_satP.value)
myfile.save_excel("VESIcal_Output.xlsx", calculations=[result], sheet_names=["SatPs"])
local_file = FileLink('./VESIcal_Output.xlsx', result_html_prefix="Click here to download: ")
display(local_file)
button_satP.on_click(on_button_clicked_satP)
# -
# # Calculate Dissolved Volatile Concentrations
# +
# create widgets so the user can choose model options
# Model selection
model_selector_diss = widgets.Dropdown(
options=[('MagmaSat', 'MagmaSat'),
('Dixon', 'Dixon'),
('Shishkina', 'ShishkinaIdealMixing'),
('IaconoMarziano', 'IaconoMarziano'),
('Liu', 'Liu'),
('MooreWater', 'MooreWater'),
('AllisonCarbon (Vesuvius)', 'AllisonCarbon_vesuvius'),
('AllisonCarbon (Sunset)', 'AllisonCarbon_sunset'),
('AllisonCarbon (SFVF)', 'AllisonCarbon_sfvf'),
('AllisonCarbon (Erebus)', 'AllisonCarbon_erebus'),
('AllisonCarbon (Stromboli)', 'AllisonCarbon_stromboli')
],
value='MagmaSat',
description='Model:',
)
# Temperature selection
temperature_selector_diss = widgets.Text(
placeholder='Temperature (degC)',
description='Temperature:',
disabled=False,
style=style
)
# Pressure selection
pressure_selector_diss = widgets.Text(
value='',
placeholder='Pressure (bars)',
description='Pressure:',
disabled=False,
style=style
)
# X_fluid selection
Xfluid_selector_diss = widgets.Text(
value='',
placeholder='XH2O fluid',
description='XH2O fluid:',
disabled=False,
style=style
)
# give user option to use args in file
temp_checkbox_diss = widgets.Checkbox(
value=False,
description='Use temperature from file (if checked, enter column name in Temperature box)',
disabled=False,
indent=False,
layout=layout,
style=style
)
# give user option to use args in file
press_checkbox_diss = widgets.Checkbox(
value=False,
description='Use pressure from file (if checked, enter column name in Pressure box)',
disabled=False,
indent=False,
layout=layout,
style=style
)
# give user option to use args in file
XH2O_checkbox_diss = widgets.Checkbox(
value=False,
description='Use XH2Ofluid from file (if checked, enter column name in XH2O box)',
disabled=False,
indent=False,
layout=layout,
style=style
)
display(model_selector_diss,
temperature_selector_diss,
temp_checkbox_diss,
pressure_selector_diss,
press_checkbox_diss,
Xfluid_selector_diss,
XH2O_checkbox_diss)
# Create a button that, when clicked, gets name of file uploaded and
# creates a BatchFile object and runs the model calculation
button_diss = widgets.Button(
description='Calculate Dissolved Volatile Concentrations',
disabled=False,
button_style='', # 'success', 'info', 'warning', 'danger' or ''
tooltip='Calculate Dissolved Volatile Concentrations',
icon='check', # (FontAwesome names without the `fa-` prefix)
layout=layout
)
button_diss.style.button_color = 'lightgreen'
output = widgets.Output()
display(button_diss, output)
def on_button_clicked_diss(b):
with output:
# print successful click
print("Button clicked.")
# get filename, content of file and create BatchFile object
uploaded_filename = next(iter(uploader.value))
content = uploader.value[uploaded_filename]['content']
with open('myfile', 'wb') as f:
f.write(content)
myfile = v.BatchFile(content)
print(model_selector_diss.value)
# sanitize inputs
if temp_checkbox_diss.value == False:
temperature = float(temperature_selector_diss.value)
else:
temperature = str(temperature_selector_diss.value)
if press_checkbox_diss.value == False:
pressure = float(pressure_selector_diss.value)
else:
pressure = str(pressure_selector_diss.value)
if XH2O_checkbox_diss.value == False:
X_fluid = float(Xfluid_selector_diss.value)
else:
X_fluid = str(Xfluid_selector_diss.value)
# run model calculation
result = myfile.calculate_dissolved_volatiles(temperature=temperature,
pressure=pressure,
X_fluid=X_fluid,
model=model_selector_diss.value)
myfile.save_excel("VESIcal_Output.xlsx", calculations=[result], sheet_names=["Dissolved_volatiles"])
local_file = FileLink('./VESIcal_Output.xlsx', result_html_prefix="Click here to download: ")
display(local_file)
button_diss.on_click(on_button_clicked_diss)
# -
# # Calculate Equilibrium Fluid
# +
# create widgets so the user can choose model options
# Model selection
model_selector_eq = widgets.Dropdown(
options=[('MagmaSat', 'MagmaSat'),
('Dixon', 'Dixon'),
('Shishkina', 'ShishkinaIdealMixing'),
('IaconoMarziano', 'IaconoMarziano'),
('Liu', 'Liu'),
('MooreWater', 'MooreWater'),
('AllisonCarbon (Vesuvius)', 'AllisonCarbon_vesuvius'),
('AllisonCarbon (Sunset)', 'AllisonCarbon_sunset'),
('AllisonCarbon (SFVF)', 'AllisonCarbon_sfvf'),
('AllisonCarbon (Erebus)', 'AllisonCarbon_erebus'),
('AllisonCarbon (Stromboli)', 'AllisonCarbon_stromboli')
],
value='MagmaSat',
description='Model:',
)
# Temperature selection
temperature_selector_eq = widgets.Text(
value='',
placeholder='Temperature (degC)',
description='Temperature:',
disabled=False,
style=style
)
# Temperature type check box if user wants to pass temp from file
temperature_selection_checkbox_eq = widgets.Checkbox(value=False,
description='Use temperature from file (if checked, put column name in Temperature box)',
disabled=False,
indent=False,
layout=layout)
# Pressure selection
pressure_selector_eq = widgets.Text(
value='',
placeholder='Pressure (bars)',
description='Pressure:',
disabled=False,
style=style
)
# Temperature type check box if user wants to pass temp from file
pressure_selection_checkbox_eq = widgets.Checkbox(value=False,
description='Use pressure from file (if checked, put column name in Pressure box)',
disabled=False,
indent=False,
layout=layout)
display(model_selector_eq,
temperature_selector_eq,
temperature_selection_checkbox_eq,
pressure_selector_eq,
pressure_selection_checkbox_eq)
# Create a button that, when clicked, gets name of file uploaded and
# creates a BatchFile object and runs the model calculation
button_eq = widgets.Button(
description='Calculate Equilibrium Fluid Compositions',
disabled=False,
button_style='', # 'success', 'info', 'warning', 'danger' or ''
tooltip='Calculate Equilibrium Fluid Compositions',
icon='check', # (FontAwesome names without the `fa-` prefix)
layout=layout
)
button_eq.style.button_color = 'lightgreen'
output = widgets.Output()
display(button_eq, output)
def on_button_clicked_eq(b):
with output:
# print successful click
print("Button clicked.")
# get filename, content of file and create BatchFile object
uploaded_filename = next(iter(uploader.value))
content = uploader.value[uploaded_filename]['content']
with open('myfile', 'wb') as f:
f.write(content)
myfile = v.BatchFile(content)
# sanitize inputs
if temperature_selection_checkbox_eq.value is True:
temperature = str(temperature_selector_eq.value)
else:
temperature = float(temperature_selector_eq.value)
if pressure_selection_checkbox_eq.value is True:
pressure = str(pressure_selector_eq.value)
else:
pressure = float(pressure_selector_eq.value)
# run model calculation
result = myfile.calculate_equilibrium_fluid_comp(temperature=temperature,
pressure=pressure,
model=model_selector_eq.value)
myfile.save_excel("VESIcal_Output.xlsx", calculations=[result], sheet_names=["EQ_Fluids"])
local_file = FileLink('./VESIcal_Output.xlsx', result_html_prefix="Click here to download: ")
display(local_file)
button_eq.on_click(on_button_clicked_eq)
# -
| webapp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Consume native Keras model served by TF-Serving
#
# This notebook shows client code needed to consume a native Keras model served by Tensorflow serving. The Tensorflow serving model needs to be started using the following command:
#
# bazel-bin/tensorflow_serving/model_servers/tensorflow_model_server \
# --port=9000 --model_name=keras-mnist-fcn \
# --model_base_path=/home/sujit/Projects/polydlot/data/tf-export/keras-mnist-fcn
from __future__ import division, print_function
from google.protobuf import json_format
from grpc.beta import implementations
from sklearn.preprocessing import OneHotEncoder
from tensorflow_serving.apis import predict_pb2
from tensorflow_serving.apis import prediction_service_pb2
from sklearn.metrics import accuracy_score, confusion_matrix
import json
import os
import sys
import threading
import time
import numpy as np
import tensorflow as tf
# +
SERVER_HOST = "localhost"
SERVER_PORT = 9000
DATA_DIR = "../../data"
TEST_FILE = os.path.join(DATA_DIR, "mnist_test.csv")
IMG_SIZE = 28
MODEL_NAME = "keras-mnist-fcn"
# -
# ## Load Test Data
# +
def parse_file(filename):
xdata, ydata = [], []
fin = open(filename, "rb")
i = 0
for line in fin:
if i % 10000 == 0:
print("{:s}: {:d} lines read".format(os.path.basename(filename), i))
cols = line.strip().split(",")
ydata.append(int(cols[0]))
xdata.append(np.reshape(np.array([float(x) / 255. for x in cols[1:]]),
(IMG_SIZE * IMG_SIZE, )))
i += 1
fin.close()
print("{:s}: {:d} lines read".format(os.path.basename(filename), i))
X = np.array(xdata, dtype="float32")
y = np.array(ydata, dtype="int32")
return X, y
Xtest, ytest = parse_file(TEST_FILE)
print(Xtest.shape, ytest.shape)
# -
# ## Make Predictions
channel = implementations.insecure_channel(SERVER_HOST, SERVER_PORT)
stub = prediction_service_pb2.beta_create_PredictionService_stub(channel)
labels, predictions = [], []
for i in range(Xtest.shape[0]):
request = predict_pb2.PredictRequest()
request.model_spec.name = MODEL_NAME
request.model_spec.signature_name = "predict"
Xbatch, ybatch = Xtest[i], ytest[i]
request.inputs["images"].CopyFrom(
tf.contrib.util.make_tensor_proto(Xbatch, shape=[1, Xbatch.size]))
result = stub.Predict(request, 10.0)
result_json = json.loads(json_format.MessageToJson(result))
y_ = np.array(result_json["outputs"]["scores"]["floatVal"], dtype="float32")
labels.append(ybatch)
predictions.append(np.argmax(y_))
print("Test accuracy: {:.3f}".format(accuracy_score(labels, predictions)))
print("Confusion Matrix")
print(confusion_matrix(labels, predictions))
| src/tf-serving/03b-consume-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''AI_ITU'': conda)'
# name: python3
# ---
# + [markdown] _uuid="5dbc8f61ce01988614b8bea0bc878ccc603c32b9" id="1WV0mfSRo1NC"
# ## Implement and apply a Two layer neural network classifier with Keras
#
# ### MNIST and Fashion-MNIST dataset
# The original [MNIST dataset](http://yann.lecun.com/exdb/mnist/) contains a lot of handwritten digits. Members of the AI/ML/Data Science community love this dataset and use it as a benchmark to validate their algorithms. In fact, MNIST is often the first dataset researchers try. *"If it doesn't work on **MNIST**, it won't work at all"*, they said. *"Well, if it does work on MNIST, it may still fail on others."*
# Here are some good reasons for using [Fashion-MNIST dataset](https://github.com/zalandoresearch/fashion-mnist):
# - MNIST is too easy. Convolutional nets can achieve 99.7% on MNIST. Classic machine learning algorithms can also achieve 97% easily. Check out our side-by-side benchmark for Fashion-MNIST vs. MNIST, and read "Most pairs of MNIST digits can be distinguished pretty well by just one pixel."
# - MNIST is overused. In this April 2017 Twitter thread, Google Brain research scientist and deep learning expert Ian Goodfellow calls for people to move away from MNIST.
# - MNIST can not represent modern CV tasks, as noted in this April 2017 Twitter thread, deep learning expert/Keras author <NAME>.
#
#
# # Task 1:
# Find Fashion MNIST on the internet and save it as `./input`
# + [markdown] _uuid="aba7736343f2285046a25c48c0fd5bdd59dc9785" id="oaNNIwIto1NE"
# ## Import packages
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" id="wVrJyvJEo1NE" outputId="222ea8c8-0e32-42db-9744-a4d7b09f1ba8" colab={"base_uri": "https://localhost:8080/"}
# #!/usr/bin/env python3
##########################################################
# Copyright (c) <NAME> <<EMAIL>> #
# Created on 4 Aug 2021 #
# Version: 0.0.1 #
# What: #
##########################################################
# ploting and EDA
import matplotlib
import matplotlib.pyplot as plt
import seaborn as sns
# packages
import pandas as pd
import numpy as np
import os
import sklearn
import tensorflow as tf
# import keras
from keras.datasets import fashion_mnist
from sklearn.metrics import accuracy_score, confusion_matrix
print(f"Matplotlib Version: {matplotlib.__version__}")
print(f"Seaborn Version: {sns.__version__}")
print(f"Pandas Version: {pd.__version__}")
print(f"Numpy Version: {np.__version__}")
print(f"Scikit-learn Version: {sklearn.__version__}")
print(f"Tensorflow Version: {tf.__version__}")
# print(f"Keras Version: {keras.__version__}")
np.set_printoptions(precision=4)
# %matplotlib inline
#plt.rcParams["figure.figsize"] = (10, 8)
# + [markdown] _uuid="9cc237434fdbd04f77d6a4a9434cf33ded6862cf" id="TWNlZIcpo1NF"
# ## Load the datasets
# We haven't use the CSV file because we want to load the dataset using Numpy.
# **Notice:** If you use the `mnist_reader.py` codes from [GitHub repository](https://github.com/zalandoresearch/fashion-mnist/blob/master/utils/mnist_reader.py), it might doesn't work. Because the `.gz` file has been unpacked.
# + [markdown] id="LNNXs2DnwMsB"
# ### skipping dataload with numpy
# + [markdown] id="Ggr1PTe4wMsB"
# We split our training data into train and validate datasets, in order to train our model and validate it using the validation data set to avoid overfitting before testing the model on the test datasets which is as real-world data for our model.
# + id="jAYaYe54wMsC"
(X_train_full, y_train_full), (X_test, y_test) = fashion_mnist.load_data()
# + _uuid="e5dc172684d46547d8f0f37d3bf7a6bd4ef9dbcb" id="GhpTMUpto1NG" outputId="fcf5a350-a27a-4f45-8c8d-3cd825eee389" colab={"base_uri": "https://localhost:8080/"}
print(f'X_train_full shape: {str(X_train_full.shape)}')
print(f'y_train_full shape: {str(y_train_full.shape)}')
print(f'X_test shape: {str(X_test.shape)}')
print(f'y_test shape: {str(y_test.shape)}')
print(f'Number of training examples: {str(y_train_full.shape[0])}')
print(f'Number of testing examples:{str(y_test.shape[0])}')
# + [markdown] id="yL579DjfwMsC"
# Note that the dataset is already split into a training set and a test set, but there is no validation set, so we’ll create one now
# + [markdown] id="YKbBZr61wMsD"
# ## Further split into Validation and Preproces
# Additionally, since we are going to train the neural network using Gradient Descent, we must scale the input features. For simplicity, we’ll scale the pixel intensities down to the 0–1 range by dividing them by 255.0 (this also converts them to floats):
# + id="2cmJr7ETwMsD" outputId="41d7cfc4-dacf-4910-9a7c-84f59d988e70" colab={"base_uri": "https://localhost:8080/"}
# Split the data into train / validation / test
X_valid, X_train = X_train_full[:5000] / 255.0, X_train_full[5000:] / 255.0
y_valid, y_train = y_train_full[:5000], y_train_full[5000:]
print (f"Number of validation examples: X_valid {str(X_valid.shape)}")
print (f"Number of training examples: X_train {str(X_train.shape)}")
print (f"Number of testing examples: X_test {str(X_test.shape)}")
# + [markdown] id="ooF2MNZpwMsD"
# When loading MNIST or Fashion MNIST using Keras rather than Scikit-Learn, one important difference is that every image is represented as a 28 × 28 array rather than a 1D array of size 784. Moreover, the pixel intensities are represented as integers (from 0 to 255) rather than floats (from 0.0 to 255.0). Let’s take a look at the shape and data type of the training set:
# + id="Q590HntgwMsE" outputId="f23f9066-a974-4dd7-ea59-0989f5a255aa" colab={"base_uri": "https://localhost:8080/"}
print(X_train_full.shape)
print(X_train_full.dtype)
# + _uuid="efeff328c6964df7b6099c9e65f4533ab115dd52" id="X5f5nH3lo1NH" outputId="6ed0d1ac-8683-4377-a5a0-3d50bae945a2" colab={"base_uri": "https://localhost:8080/", "height": 297}
# random check with nine training examples
np.random.seed(0);
indices = list(np.random.randint(y_train.shape[0],size=9))
for i in range(9):
plt.subplot(3,3,i+1)
plt.imshow(X_train[indices[i]].reshape(28,28), cmap='gray', interpolation='none')
plt.title(f"Index {indices[i]} Class {y_train[indices[i]]}")
plt.tight_layout()
# + [markdown] _uuid="4457d2213139ebf65c0eff199d283da772a20245" id="nkXgY8uvo1NH"
# | 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 |
# | ----------- | ------- | -------- | ----- | ---- | ----- | ----- | ------- | ---- | ---------- |
# | T-shirt/top | Trouser | Pullover | Dress | Coat | Sanda | Shirt | Sneaker | Bag | Ankle boot |
#
# + _uuid="e72f4e27e1c03fabcdb1ac046c24ef3647b9b044" id="cYn1enbwo1NI" outputId="a67086cc-97e3-44e3-aab4-150985892a56" colab={"base_uri": "https://localhost:8080/"}
print("Feature Values: \nFrom " + str(np.min(X_train)) + " to " + str(np.max(X_train)))
print("Data type: " + str(X_train.dtype))
print("\nLabel Values: ")
print(set(list(y_train)))
print("Data type: " + str(y_train.dtype))
# + [markdown] id="M-t8iT27BlS7"
# ## Add callback function to stop the training at when a accuracy is reached
# fit() method accepts a callbacks argument that lets you specify a list of objects that Keras will call at the start and end of training, each epoch and before and after processing each batch.
# + id="XvM9MyJEwMsF"
class myCallback(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs={}):
if(logs.get('accuracy')>0.96):
print("\nReached 96% accuracy so cancelling training!")
self.model.stop_training = True
# + [markdown] _uuid="7c4802b4a024d30f949f7859511c93d5af383d95" id="HVZzxruCo1NI"
# ## Initialize the neural network with Keras
# + _uuid="8cf77fab4c742286fad45ab1e91baa7550616261" id="0U8tWvj0o1NJ"
# Simple classification MLP with two hidden layers:
callbacks = myCallback()
model = tf.keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="relu"),
keras.layers.Dense(100, activation="relu"),
keras.layers.Dense(10, activation="softmax")
])
# + [markdown] _uuid="d5bd190cb03a1b59a1b06a7b3410f0058c36da2d" id="SytJ_P3Do1NL"
# ## Train your network
# + _uuid="aaebb656111f9806e88f8663f09d282f5548b55d" id="OPprpt-4o1NM"
model.compile(loss="sparse_categorical_crossentropy",
optimizer="adam",
metrics=["accuracy"])
# + _uuid="b59c706ae21525d11ad5722d06607bc359100ede" id="Io9YQejPo1NM" outputId="04a3df2b-fe3c-4293-feee-73b90b26b91a" colab={"base_uri": "https://localhost:8080/"}
history = model.fit(X_train, y_train, epochs=50, callbacks=[callbacks], validation_data=(X_valid, y_valid))
# + id="CWs2IaK3EeXn" outputId="d8d1fdbd-e2f7-430a-e3b5-33718cfbf805" colab={"base_uri": "https://localhost:8080/"}
model.summary()
# + [markdown] _uuid="d9816eff48579c48546ea94ba0659c9ddcc3f766" id="uYY4T5Veo1NM"
# ## Print the training vs validation curves
#
#
# + _uuid="e1a9bcbe3b680acab4e3c9158a85109331f48580" id="WZOkoVw4o1NN" outputId="870bdbe4-3e82-4e7a-f500-039b08416a9a" colab={"base_uri": "https://localhost:8080/", "height": 324}
pd.DataFrame(history.history).plot(figsize=(8, 5))
plt.grid(True)
plt.gca().set_ylim(0, 1) # set the vertical range to [0-1]
plt.show()
# + [markdown] id="S69kmNvcuF3J"
# ## Caclulate the Accuracy and F1 metric
# + id="NbTatClq6xz9" outputId="b4a67737-b3f1-4806-eca6-2293a5772537" colab={"base_uri": "https://localhost:8080/"}
from sklearn.metrics import accuracy_score
y_pred_nn = np.argmax(model.predict(X_test), axis=-1)
perf = accuracy_score(y_test, y_pred_nn)
print(f"Accuracy score: {perf}")
# + id="6VLF0bFt8igD" outputId="39b9ab79-d3d5-4b12-9e86-a56c3b57fd37" colab={"base_uri": "https://localhost:8080/"}
evaluation = model.evaluate(X_test, y_test)
print(f'Test Accuracy : {evaluation[1]:.3f}')
# + [markdown] id="VFOtAyGU9aNN"
# With the model trained, you can use it to make predictions about some images.
# + id="oADFzIwj659I" outputId="9b1615b9-09f6-4a8a-a525-c2a9828f676a" colab={"base_uri": "https://localhost:8080/", "height": 609}
# get the predictions for the test data and pass to confusion_matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred_nn)
plt.figure(figsize = (14,10))
sns.heatmap(cm, annot=True)
# Sum the diagonal element to get the total true correct values
# + _uuid="c225c8302419b1c46ac972a63c0119a93ad68de3" id="c5Ggu_mUo1NN" outputId="d7832f88-4413-4d77-82cd-98d99ec23172" colab={"base_uri": "https://localhost:8080/"}
from sklearn.metrics import classification_report
num_classes = 10
target_names = [f"Class {i}" for i in range(num_classes)]
print(classification_report(y_test, predicted_classes, target_names = target_names))
| day12 - Hand-ins/03_fashion_mnist_with_keras_neural_networks.ipynb |