code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''base'': conda)'
# language: python
# name: python397jvsc74a57bd0f16b4f9c118091e521975b08f7e11c780453654917dc239e484e68cca92583fe
# ---
from splinter import Browser
from bs4 import BeautifulSoup
import pandas as pd
from webdriver_manager.chrome import ChromeDriverManager
# Setup splinter
executable_path = {'executable_path': ChromeDriverManager().install()}
browser = Browser('chrome', **executable_path, headless=False)
url = 'https://redplanetscience.com'
browser.visit(url)
html = browser.html
soup = BeautifulSoup(html, 'html.parser')
news_title = soup.find('div', class_='content_title').text
news_p = soup.find('div', class_='article_teaser_body').text
print(news_title)
print(news_p)
url = 'https://spaceimages-mars.com/'
browser.visit(url)
html = browser.html
soup = BeautifulSoup(html, 'html.parser')
relative_image_url = soup.find('img', class_='headerimage')["src"]
featured_image_url = url + relative_image_url
print(featured_image_url)
mars_table = pd.read_html('https://galaxyfacts-mars.com', header=0)
mars_df = mars_table[0]
mars_df.head()
mars_html = mars_df.to_html()
print(mars_html)
url = 'https://marshemispheres.com/'
browser.visit(url)
# +
html = browser.html
soup = BeautifulSoup(html, 'html.parser')
hemisphere_image_urls = []
image_infos = soup.find_all('div', class_='item')
for image in image_infos:
title = image.h3.text
rel_image_url = image.find('img', class_='thumb')['src']
image_url = url + rel_image_url
hemisphere_dict = {"title": title, "img_url": image_url}
hemisphere_image_urls.append(hemisphere_dict)
hemisphere_image_urls
# -
browser.quit()
| .ipynb_checkpoints/mission_to_mars-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="inqwDoOzFX3D" executionInfo={"status": "ok", "timestamp": 1609843087250, "user_tz": -60, "elapsed": 38151, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjPmJNhCoYmd7uFIXkkB5FOlzcQOBV3of5hL8tOBQ=s64", "userId": "05263850157645353544"}} outputId="c57b79b6-c1cb-47ec-809d-a69aa5df672c"
from google.colab import drive
drive.mount('/content/gdrive')
# + colab={"base_uri": "https://localhost:8080/"} id="tuOgSlgSyuYz" executionInfo={"status": "ok", "timestamp": 1609843107671, "user_tz": -60, "elapsed": 20412, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjPmJNhCoYmd7uFIXkkB5FOlzcQOBV3of5hL8tOBQ=s64", "userId": "05263850157645353544"}} outputId="47562af8-1f05-483e-9994-7c12accd917e"
# %cd /content/gdrive/MyDrive
# !unzip BrixIA_small.zip -d /content/
# %cd /content/gdrive/MyDrive/Colab\ Notebooks
# + colab={"base_uri": "https://localhost:8080/"} id="4VIbHT9MzA1V" executionInfo={"status": "ok", "timestamp": 1609843119608, "user_tz": -60, "elapsed": 8715, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjPmJNhCoYmd7uFIXkkB5FOlzcQOBV3of5hL8tOBQ=s64", "userId": "05263850157645353544"}} outputId="6d7cbc62-f795-4cd6-ddc7-1adbbbebe617"
# !pip install tensorboardX
# + colab={"base_uri": "https://localhost:8080/"} id="AnXRLG4EynqI" executionInfo={"status": "ok", "timestamp": 1609843129890, "user_tz": -60, "elapsed": 13617, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjPmJNhCoYmd7uFIXkkB5FOlzcQOBV3of5hL8tOBQ=s64", "userId": "05263850157645353544"}} outputId="687dff9a-a77d-4232-90d4-6a44912a005c"
import model as M
# + colab={"base_uri": "https://localhost:8080/"} id="J2lJwXb3SUCe" outputId="1e1bb71c-11d1-4113-82ba-9e4b736f4491"
# you will need to customize PATH_TO_IMAGES to where you have uncompressed
# BrixIA images
PATH_TO_IMAGES = "/content/BrixIAsmall"
WEIGHT_DECAY = 1e-4
LEARNING_RATE = 0.001
M.train_cnn(PATH_TO_IMAGES, LEARNING_RATE, WEIGHT_DECAY, fine_tune=True, freeze=True, initial_model_path='/content/checkpoint_best')
# + id="251xt2LWR4Cp"
| model/notebooks/fine_tune_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
from tensorflow.examples.tutorials.mnist import input_data # MNIST 데이터를 위한 유틸리티
# Mac OS Error: https://github.com/dmlc/xgboost/issues/1715
import os
os.environ['KMP_DUPLICATE_LIB_OK']='True'
# -
DATA_DIR = './tmp/data'
NUM_STEPS = 1000
MINIBATCH_SIZE = 100
data = input_data.read_data_sets(DATA_DIR, one_hot=True) # 로컬에 저장하여 사용
# +
x = tf.placeholder(tf.float32, [None, 784]) # Placeholder: 연산 그래프가 실행될 때 제공되어야 하는 값
W = tf.Variable(tf.zeros([784, 10])) # Variable: 연산 과정에서 조작되는 값
y_true = tf.placeholder(tf.float32, [None, 10]) # None: 한 번에 얼마나 많은 이미지를 사용할지는 이 시점에서는 정하지 않음
y_pred = tf.matmul(x, W)
# +
cross_entropy = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits_v2(
logits=y_pred, labels=y_true))
gd_step = tf.train.GradientDescentOptimizer(0.5).minimize(cross_entropy)
correct_mask = tf.equal(tf.argmax(y_pred, 1), tf.argmax(y_true, 1))
accuracy = tf.reduce_mean(tf.cast(correct_mask, tf.float32))
# +
with tf.Session() as sess:
# Train
sess.run(tf.global_variables_initializer()) # 모든 변수 초기화
for _ in range(NUM_STEPS):
batch_xs, batch_ys = data.train.next_batch(MINIBATCH_SIZE)
# print(batch_xs)
# print(batch_ys)
# break
sess.run(gd_step, feed_dict={x: batch_xs, y_true: batch_ys}) # Placeholder에 값을 feed 함
# Test
ans = sess.run(accuracy, feed_dict={x: data.test.images, y_true: data.test.labels})
print(x.name)
print(W.name)
print("Accuracy: {:.4}%".format(ans*100))
# -
| 02__up_and_running/Softmax.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="_ALAeBKo7Got"
# #Creation of Arrays#
# + id="SHyh2rlT8Cp7"
import numpy as np
def print_np_details(arr, name):
print('array ', name)
print(arr)
print('python type = ', arr.astype)
print("numpy data type = ", arr.dtype)
print("shape = ", arr.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="jE2Dkjg920H5" outputId="c750d5ca-5b50-4d95-cee3-a23fc9c16162"
#import numpy as np
a1D = np.array([1, 2, 3, 4]) # this creates a numpy.ndarray object from a Python list
print_np_details(a1D, "a1D")
# + colab={"base_uri": "https://localhost:8080/"} id="h_znn5TJ8rQd" outputId="cbcb40be-ca8f-483f-a4a2-a432112a4331"
# this creates a 2d array
a2D = np.array([[1, 2, 3, 4], [5, 6, 7, 8]])
print_np_details(a2D, "a2D")
# + colab={"base_uri": "https://localhost:8080/"} id="8cKErYlw7YNk" outputId="f50194cb-351f-4196-8fa9-bbc196669194"
#specify the type of the array
a = np.array([127, 128, 129], dtype=np.int8)
# 8-bit integer represents value from -128 to 127
print_np_details(a, "a")
# + colab={"base_uri": "https://localhost:8080/"} id="wad5Kg_k91WM" outputId="25a19ff1-1632-4db5-bb95-3283b13ba6ad"
#setting the data type to unsigned int
a = np.array([127, 128, 129, 256], dtype=np.uint8)
# 8-bit unsigned integer represents value from 0 to 255
print_np_details(a, "a")
# + colab={"base_uri": "https://localhost:8080/"} id="dKXafIh7_PXP" outputId="141b7060-d791-4915-ca30-8c53a44e7672"
#setting the data type to 16-bit int
a = np.array([127, 128, 129, 255], dtype=np.int16)
print_np_details(a, "a")
#setting the data type to 32-bit float
b = np.array([127, 128, 129, 255], dtype=np.float32)
print_np_details(b, "b")
# + colab={"base_uri": "https://localhost:8080/"} id="Bn4a1KrUAyia" outputId="15deafc5-91a1-4745-8daf-565c240a6bc2"
#zero matrix
a = np.zeros((2, 3))
print_np_details(a, "a")
b = np.zeros((2, 3), dtype=np.int16)
print_np_details(b, "b")
# + colab={"base_uri": "https://localhost:8080/"} id="_z4j92ewBPF7" outputId="d8c3d899-44ad-4aeb-f997-667860cb188c"
# one matrix
a = np.ones((3, 5))
print(a)
# identity matrix
a = np.eye(4)
print(a)
# + [markdown] id="mC00GNxoFSH0"
# #Slicing Arrays#
# + colab={"base_uri": "https://localhost:8080/"} id="_kIwUlNGFQdD" outputId="86e64df8-67b1-4798-ce1b-640ad6872abf"
a = np.array([[1, 2, 3], [3, 4, 6.7], [5, 9.0, 5]])
print(a)
print("selecting the first row")
print(a[0, :]) # zero-based indexing
print("selecting the second column")
print(a[:, 1])
print("selecting the second and the third columns")
print(a[:, 1:3])
print("selecting the second and the third rows and the 3rd column")
print(a[1:3, 2])
print("selecting the entry a_{2, 3}")
print(a[1, 2])
# + colab={"base_uri": "https://localhost:8080/"} id="7zJ_oBTkoh_A" outputId="c34ea86c-a4b8-4037-e938-227ffe193dce"
a = np.array([[1, 2, 3], [3, 4, 6.7], [5, 9.0, 5]])
print(a)
print("assigning values to the second and the third rows and the 3rd column")
a[1:3, 2] = np.array([0.1, 0.2])
print(a)
print("assigning values to the first row")
a[0, :] = np.array([100, 200, 300])
print(a)
# + colab={"base_uri": "https://localhost:8080/"} id="TF9JPOLEnPJ2" outputId="38fb96bd-268c-4d31-d121-3ad29eec6089"
a = np.array([[1, 2, 3], [3, 4, 6.7], [5, 9.0, 5]])
print('selecting the diagonal')
print(np.diagonal(a))
print('selecting the diagonal from the first and second row')
print(np.diagonal(a[0:2, :]))
print('assign a new diagonal to a')
np.fill_diagonal(a, np.array([-4, -5, -6]))
print(a)
# + [markdown] id="kE88JzE_F-4T"
# # Element-wise Operations
# + colab={"base_uri": "https://localhost:8080/"} id="PeQ1L7DoGCJm" outputId="8ba2b3ea-d948-4d21-9f66-b5574ff30e20"
a = np.array([1, 2, 3, 4])
b = np.array([2, 6, 9, 12])
print("element-wise addition")
print(a+0.2)
print("element-wise multiplication")
print(a*2)
print("element-wise division")
print(a/3)
print("element-wise addition")
print(a+b)
print("element-wise division")
print(a/b)
# + [markdown] id="MtExvSsrbWJ2"
# # Broadcasting
# + colab={"base_uri": "https://localhost:8080/"} id="XnzG5yWobViH" outputId="eacb4142-ab83-4402-d519-c2d6726120c8"
A = np.array([1, 2, 3])
B = np.array([[2, 3, 4], [5, 6, 7], [9, 10, 11]])
print(A.shape)
print(B.shape)
A+B
# + [markdown] id="oOnLT6iSGl0I"
# #Linear Algrebra Operations#
# + colab={"base_uri": "https://localhost:8080/"} id="aUHbUYsEGkoZ" outputId="1acbe62b-a165-4991-81fc-edb5970e67ef"
# inner product
a = np.array([1, 2, 3, 4])
b = np.array([1/2, 1/16, 9, 12])
print('inner product')
print(a.dot(b))
# matrix-vector multiplication
A = np.array([[1, 1/2, 1/3], [3, 0.4, 6.7], [5, 9.0, 5]])
x = np.array([2, 1.3, 4.8])
print('matrix-vector product')
print(A.dot(x))
# linear combination of columns
y = x[0] * A[:, 0] + x[1] * A[:, 1] + x[2] * A[:, 2]
print('linear combination of columns')
print(y)
# + colab={"base_uri": "https://localhost:8080/", "height": 506} id="_NvO4SSnsQiI" outputId="3068c8fb-a131-4227-b083-fcf8ff5cddda"
from numpy.linalg import inv
import timeit
from matplotlib import pyplot as plt
dim = [10, 100, 200, 500, 750, 1000]
lst = []
for i in range(len(dim)):
print(str(dim[i])+ ' dimensions')
stmt = 'import numpy; A = numpy.random.rand(' + str(dim[i]) + ',' + str(dim[i])+ ')'
t = timeit.timeit('numpy.linalg.inv(A)', setup=stmt, number=100)
print(t)
lst.append(t)
plt.plot(dim, lst, 'bo-')
# + [markdown] id="aXWSh-4SxRty"
# From 750 to 1000 dimensions, n grows by 33% but the computational time doubles!
# + id="t0gzrqWuxn-Y"
# + [markdown] id="GCrp9t6K0f2b"
# ## SVD Decomposition
# + colab={"base_uri": "https://localhost:8080/"} id="5z8-b4fm05sK" outputId="0a01730e-58b0-4dbf-ee4b-898a3f198c8b"
A = np.array([[1, 2, 3, 6], [-10, 3, 4, 6.7], [5, 9.0, 11, 5]])
U,S,V = np.linalg.svd(A)
print('U')
print(U)
print('S')
print(S)
print('V^T')
print(V)
# + colab={"base_uri": "https://localhost:8080/"} id="MsbB7ovS1oie" outputId="6489031e-78d2-404a-d376-9085dbe80f83"
print('verify that U is orthogonal')
print(U.T.dot(U))
print('verify that V is orthogonal')
print(V.T.dot(V))
# + colab={"base_uri": "https://localhost:8080/"} id="VVlGuXly1xHX" outputId="87a4b5f5-312c-44a4-cc35-bee9bd948919"
import math
#what happens when we remove the largest singular value?
s1 = S.copy()
s1[0]=0 # set the largest singular value to zero
S1 = np.diag(s1)
S1 = np.hstack([S1, np.zeros((3, 1))]) # make a 3-by-4 diagonal matrix, with the last column being 0
print(S1)
A1 = U.dot(S1).dot(V)
error = pow(np.linalg.norm(A), 2) - pow(np.linalg.norm(A1), 2) # np.linalg.norm(A) is known as the Frobenius norm of A
error = math.sqrt(error)
print(error) # error in matrix Frobenius norm
s1 = S.copy()
s1[2]=0 # set the smallest singular value to zero
S1 = np.diag(s1)
S1 = np.hstack([S1, np.zeros((3, 1))]) # make a 3-by-4 diagonal matrix, with the last column being 0
print(S1)
A1 = U.dot(S1).dot(V)
error = pow(np.linalg.norm(A), 2) - pow(np.linalg.norm(A1), 2) # np.linalg.norm(A) is known as the Frobenius norm of A
error = math.sqrt(error)
print(error) # error in matrix Frobenius norm
# + [markdown] id="9A-AGsV78ADv"
# Is this a coincidence? Of course not!
#
# The Frobenius norm of matrix $A$ is defined as $\|A\|_{F} = \sqrt{\sum_i \sum_j A_{ij}^2}$
#
# Denote the singular values by $\sigma_1, \sigma_2,\ldots, \sigma_k$, we have $\sum \sigma_i^2 = \|A\|_{F}^2$. That is, the sum of squared singular values equals the squared Frobenius norm.
#
# Thus, removing a singular value has the effect of reducing the Frobenius norm by that amount.
# + [markdown] id="N8bKhtahZaAv"
# ## Eigendecomposition
# + colab={"base_uri": "https://localhost:8080/"} id="rS10sAiPZnAN" outputId="7dc0ccd3-7135-470a-ff40-648d9a5bf597"
# create an orthogonal matrix
from scipy.stats import ortho_group
U = ortho_group.rvs(dim=5)
U.dot(U.T)
# + colab={"base_uri": "https://localhost:8080/"} id="UYfD6kIfafmU" outputId="58617a4d-46bc-405f-c4af-9a9e3b28c63e"
# create a symmetric positive definite matrix
S = np.diag([1.0, 0.5, 0.72, 1.22, 0.93])
A = U.dot(S).dot(U.T)
A
# + colab={"base_uri": "https://localhost:8080/"} id="9tNG7hI5an7f" outputId="93ddec2c-65cf-48fb-d9d2-e4e96cca6c6d"
# verify positive definiteness
for i in range(10):
v = np.random.rand(5)
print(v.T.dot(A).dot(v))
# + colab={"base_uri": "https://localhost:8080/"} id="sE9mXLwZa4XS" outputId="3a2f32ef-41c7-4c7d-84d8-ee12ca06248e"
# perform eigendecomposition
np.linalg.eig(A)
| PythonExamples/Numpy_Usage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import yfinance
import talib
from matplotlib import pyplot as plt
from mplfinance.original_flavor import candlestick_ohlc
from matplotlib.pylab import date2num
data = yfinance.download('NFLX','2016-1-1','2020-1-1')
rsi = talib.RSI(data["Close"])
fig = plt.figure()
fig.set_size_inches((25, 18))
ax_rsi = fig.add_axes((0, 0.24, 1, 0.2))
ax_rsi.plot(data.index, [70] * len(data.index), label="overbought")
ax_rsi.plot(data.index, [30] * len(data.index), label="oversold")
ax_rsi.plot(data.index, rsi, label="rsi")
ax_rsi.plot(data["Close"])
ax_rsi.legend()
# +
import numpy as np
section = None
sections = []
for i in range(len(rsi)):
if rsi[i] < 30:
section = 'oversold'
elif rsi[i] > 70:
section = 'overbought'
else:
section = None
sections.append(section)
patience = np.linspace(1,15,15)
profits = []
for pat in range(1,1000):
trades = []
for i in range(1,len(sections)):
trade = None
if sections[i-pat] == 'oversold' and sections[i] == None:
trade = True
if sections[i-pat] == 'overbought' and sections[i] == None:
trade = False
trades.append(trade)
acp = data['Close'][len(data['Close'])-len(trades):].values
profit = 0
logs = []
qty = 10
for i in range(len(acp)-1):
true_trade = None
if acp[i] < acp[i+1]:
true_trade = True
elif acp[i] > acp[i+1]:
true_trade = False
if trades[i] != None:
if trades[i] == true_trade:
profit += abs(acp[i+1] - acp[i]) * qty
logs.append(abs(acp[i+1] - acp[i]) * qty)
elif trades[i] != true_trade:
profit += -abs(acp[i+1] - acp[i]) * qty
logs.append(-abs(acp[i+1] - acp[i]) * qty)
profits.append(profit)
| RSI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
tupleE = "1", "3", "5"
tupleE
# -
tupleE[1] = "5"
1, 3, 5 = tupleE
# +
print(tupleE[0])
print(tupleE[1])
| Chapter01/Exercise 1.11/Exercise 1.11.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
from PIL import Image
import shutil
from multiprocessing import Pool
from resize_image import resize_image224
# +
IMAGES = os.path.join('data', 'images')
PROCESSED_IMAGES = os.path.join('data', 'processed_images')
if os.path.exists(PROCESSED_IMAGES):
shutil.rmtree(PROCESSED_IMAGES)
os.mkdir(PROCESSED_IMAGES)
pool = Pool(processes=16)
images = os.listdir(IMAGES)
images = [(os.path.join(IMAGES, file), os.path.join(PROCESSED_IMAGES, file),) for file in images]
_ = pool.map(resize_image224, images)
| nhs-chest-xray/preprocess.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''wfst_tutorial'': conda)'
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="Qq1Hz6CKWdwl" outputId="3d8f5bd6-f10e-431d-9039-eb88164fbb95"
### WARNING: This notebook will not work in a Colab environment.
BRANCH= "r1.5.0"
# !git clone -b $BRANCH https://github.com/NVIDIA/NeMo
# %cd NeMo
# !./reinstall.sh
# +
import pynini
import nemo_text_processing
from pynini.lib import pynutil
# +
from nemo_text_processing.text_normalization.en.graph_utils import GraphFst, NEMO_DIGIT, delete_space, NEMO_SIGMA, NEMO_NOT_QUOTE, delete_extra_space, NEMO_NON_BREAKING_SPACE
from nemo_text_processing.text_normalization.normalize import Normalizer
from nemo_text_processing.inverse_text_normalization.fr.taggers.cardinal import CardinalFst
from nemo_text_processing.inverse_text_normalization.fr.taggers.decimal import DecimalFst
from nemo_text_processing.inverse_text_normalization.fr.taggers.money import MoneyFst
from nemo_text_processing.inverse_text_normalization.fr.taggers.ordinal import OrdinalFst
from nemo_text_processing.inverse_text_normalization.fr.taggers.punctuation import PunctuationFst
from nemo_text_processing.inverse_text_normalization.fr.taggers.time import TimeFst
from nemo_text_processing.inverse_text_normalization.fr.taggers.whitelist import WhiteListFst
from nemo_text_processing.inverse_text_normalization.fr.taggers.word import WordFst
from nemo_text_processing.inverse_text_normalization.fr.verbalizers.cardinal import CardinalFst
from nemo_text_processing.inverse_text_normalization.fr.verbalizers.decimal import DecimalFst
from nemo_text_processing.inverse_text_normalization.fr.verbalizers.money import MoneyFst
from nemo_text_processing.inverse_text_normalization.fr.verbalizers.ordinal import OrdinalFst
from nemo_text_processing.inverse_text_normalization.fr.verbalizers.time import TimeFst
from nemo_text_processing.inverse_text_normalization.fr.verbalizers.whitelist import WhiteListFst
from nemo_text_processing.inverse_text_normalization.fr.verbalizers.word import WordFst
# + [markdown] id="T0JxcvuPHvn9"
# NeMo's Text Processing module uses Weighted Finite State Transducers (WFST) to deploy grammars for both efficient text normalization (TN) and inverse text normalization (ITN). In this tutorial, you will learn to build a normalization grammar from the ground up to use in your own text processing tasks.
# -
# # Table of Contents
# - [WFSTs](#wfsts)
# - [NeMo Text Processing](#nemo-text-processing)
# - [Getting Started](#getting-started)
# - [Cardinal WFST](#cardinal-wfst)
# - [Ordinal WFST](#ordinal-wfst)
# - [Decimal WFST](#decimal-wfst)
# - [Money WFST](#money-wfst)
# - [Time WFST](#time-wfst)
# - [WhiteList WFST](#whitelist-wfst)
# - [Word and Punctuation WFST](#word-and-punctuation-wfst)
# - [Other Classes](#other-classes)
# - [Tokenize and Classify](#tokenize-and-classify)
# - [Verbalize and Verbalize Final](#verbalize-and-verbalize-final)
# - [Deployment](#deployment)
# + [markdown] id="lMUovcMsfXyI"
# # WFSTs <a id="wfsts"></a>
# + [markdown] id="Y1ejNMLbH1jM"
# WFSTs are a form of [Finite State Machines](https://en.wikipedia.org/wiki/Finite-state_machine) used to graph relations between regular languages (or [regular expressions](https://en.wikipedia.org/wiki/Regular_expression)). For our purposes, they can be defined by two major properties:
#
# 1. Mappings between accepted input and output expressions for text substitution
# 2. Path weighting to direct graph traversal
# + [markdown] id="nNg45ZuaP_A8"
# For example, consider a simple normalization task of mapping the word "cent" (French for "one hundred") to the numerical representation `100`. We would begin with a Finite State representation of the regex `/cent/`:
# + [markdown] id="uxo7gUkW_XKT"
# 
# + [markdown] id="fahsjMVFlbCa"
# And then create a mapping to the text string `100`:
# + [markdown] id="IMJ-fNSk_jXC"
# 
# + [markdown] id="bPKW0I4yAGUb"
# *Note: Null characters are expressed as `ε` by convention*
# + [markdown] id="_0NK3aW5nG9C"
# This would give us a WFST with universal path weights. (By default, `pynini` uses [tropical semirings](https://en.wikipedia.org/wiki/Tropical_semiring) for arcs, giving each arc a default weight of `0`.)
# + [markdown] id="CzBc9D3qTGJ-"
# Now, let us consider expanding our model. To indicate values between `100` and `200`, French uses the number scheme of `cent + digit`. For example, `120` would be pronounced as "cent-vingt". To create the appropriate output string, we would now want to map "cent" to `1` and the remaining aspect of our string to the appropriate digit representation.
# + [markdown] id="GRrKNQRjFDoL"
# 
# + [markdown] id="jLpm4mufAfUz"
# However this would make our graph [non-deterministic](https://en.wikipedia.org/wiki/Nondeterministic_algorithm) - it will have multiple possibilities for termination. Now an input of "cent-vingt" could have the outcome of `100` or `10020` when only one is correct.
# -
# 
# + [markdown] id="c-GJTpgIAf7S"
# To correct this, we may add a new end state and a weight to the path that accepts the input without `s`:
# + [markdown] id="6GJcsdttGg_S"
# 
# + [markdown] id="mHft1gzsAipc"
# Now, we can guarantee an ideal mapping by relying on a shortest-path (smallest-weight) heuristic: traversal of the graph will prioritize longer inputs, only converting "cent" to `100` when a larger input isn't available. As such, we've now removed the undesired output `10020` while preserving our desired coverage in string mapping.
#
# This use of weights to ensure predictable behavior allows WFSTs to exploit the efficiency of standard graph traversal algorithms while also maintaining versatility.
# + [markdown] id="8Ik4PBXafSSB"
# # NeMo Text Processing <a id="nemo-text-processing"></a>
# + [markdown] id="b2fcWKhqYVF5"
# Following [Google's Kestrel](https://www.researchgate.net/publication/277932107_The_Kestrel_TTS_text_normalization_system) framework, NeMo deploys two composite WFSTs for text normalization. They are as follows:
# 1. A *classifier* (or tagger) to label potential tokens by 'semiotic class' (e.g. currency, ordinal number, street address)
# 2. A *verbalizer* to render a tagged token in conventional written form
#
# For example, consider the sentence: <<le premier juillet il a mangé trente-cinq pommes>>
#
# For an ITN task, a tokenizer would identify the following tokens:
#
# `["le" ,"premier", "juillet", "il", "a", "mangé", "trente-cinq", "pommes"]`
#
# and provide each a class token:
#
# - `tokens { name: "le" }`
# - `tokens { date { day: "1" month: "juillet" } } `
# - `tokens { name: "il" }`
# - `tokens { name: "a" }`
# - `tokens { name: "mangé" }`
# - `tokens { cardinal { integer: "35" } }`
# - `tokens { name: "pommes" }`
#
# These tokens are then passed to a 'verbalizer' WFST, which renders each token in a conventional written form:
#
# - `tokens { name: "le" }` -> `le`
# - `tokens { date { day: "1" month: "juillet" } } ` -> `1ᵉʳ`
# - `tokens { name: "il" }` -> `juillet`
# - `tokens { name: "il" }` -> `il`
# - `tokens { name: "a" }` -> `a`
# - `tokens { name: "mangé" }` -> `mangé`
# - `tokens { cardinal { integer: "35" } }` -> `35`
# - `tokens { name: "pommes" }` -> `pommes`
#
# and merged into a normalized string:
#
# `le 1ᵉʳ juillet il a mangé 35 pommes`
#
# With the equivalent TN task being the reverse process.
# + [markdown] id="_n-5JExAbvwr"
# A few things to note:
# - Each class token has a unique set of field names that must be parsed by the classifier. The default field names for NeMo are chosen to mirror the syntax in [Sparrowhawk](https://github.com/google/sparrowhawk) to enable deployment. If these fields are not exact, you will not be able to use Sparrowhawk.
# - NeMo assumes no punctuation (unless explicitly provided in the grammar) and all lower casing to ease integration with upstream ASR.
# - The `name` class token is default for any token that does not require processing. It will be left 'as is.'
# - You may note how the tokenizer performed the conversion of `premier` to `1` while the verbalizer normalized `1` -> `1ᵉʳ`. Such decisions are implementation dependent and will vary depending on preference and language. (That is, normalization from `premier` -> `1ᵉʳ` could have been a tokenization step.)
# - By default, NeMo will create several permutations of key values in a token to ease normalization. That is, given the token `tokens { date { day: "1" month: "juillet" } }`, it will also produce paths for `tokens { date { month: "juillet" day: "1" } }`. To prevent this and avoid ambiguity in verbalizer input, tokens can be assigned a `preserve_order` attribute to prevent permutation. (e.g. `tokens { date { day: "1" month: "juillet" preserve_order: true } }`) (We will discuss this [later in the tutorial](#verbalizer).)
# -
# ## WFST Classes
# NeMo Text Processing's base languages currently support only the following semiotic classes to permit integration with Sparrowhawk deployment.
#
# - CARDINAL
# - ORDINAL
# - DECIMAL
# - FRACTION
# - MEASURE
# - MONEY
# - TIME
# - DATE
# - ELECTRONIC
# - TELEPHONE
# - WHITELIST
# - WORD
# - PUNCTUATION
#
# For this tutorial, we will be focusing on the following classes:
# - CARDINAL
# - ORDINAL
# - DECIMAL
# - MONEY
# - TIME
# - WHITELIST
# - WORD
# - PUNCTUATION
#
# While not comprehensive, these classes will provide enough foundation and exposure to edge cases that you will feel comfortable constructing for other cases.
#
# **NOTE**: *If you intend to only develop for personal use with NeMo, you may rename these classes as desired. However, Sparrowhawk integration
# REQUIRES use of only these tags and their assigned attributes. For list of Sparrowhawk tokens and attributes, [consult the Sparrowhawk repository](https://github.com/yzhang123/sparrowhawk/blob/test/src/proto/semiotic_classes.proto)*
# ## Futher Reading
# If you wish to learn more about NeMo Text Processing, you may wish to consult the following:
# - [<NAME>, <NAME>, <NAME>, and <NAME>, "NeMo Inverse Text Normalization: From Development To Production"](https://arxiv.org/pdf/2104.05055.pdf)
# - [NeMo's Text Normalization Documentation](https://docs.nvidia.com/deeplearning/nemo/user-guide/docs/en/main/nemo_text_processing/intro.html)
# - [NeMo's Text Normalization Deployment Documentation](https://docs.nvidia.com/deeplearning/nemo/user-guide/docs/en/main/tools/text_processing_deployment.html)
# - NeMo's [Text Normalization Tutorial](https://colab.research.google.com/github/NVIDIA/NeMo/blob/stable/tutorials/text_processing/Text_Normalization.ipynb) or [Inverse Text Normalization](https://colab.research.google.com/github/NVIDIA/NeMo/blob/stable/tutorials/text_processing/Inverse_Text_Normalization.ipynb) tutorials
# - [Sparrowhawk Documentation](https://github.com/google/sparrowhawk)
# For further information regarding WFSTs, please see:
# - [<NAME> and <NAME>, *Natural Language Processing*, Ch. 2](https://web.stanford.edu/~jurafsky/slp3/2.pdf)
# - [<NAME> and <NAME>, *Finite-State Text Processing*](http://www.morganclaypoolpublishers.com/catalog_Orig/product_info.php?products_id=1636)
# + [markdown] id="XFdXRcnUfI25"
# # Getting Started <a id="getting-started"></a>
#
# + [markdown] id="K3Zl3VwqdYqL"
# To begin tokenizer development, make sure you have [installed NeMo from source](https://github.com/NVIDIA/NeMo).
# + [markdown] id="rGg7Bf13FXgc"
# For this tutorial, we will focus on developing an Inverse Text Normalization system, such as one you may encounter in downstream ASR processing. As such, we will navigate to
# `nemo_text_processing/inverse_text_normalization` and create a directory for our target language (French) and subdirectories
# for `taggers` and `verbalizers`. You may also wish to create a `data` subdirectory to ease navigation.
#
# (Note, for text normalization, the suggested directory structure would be the same within the `nemo_text_processing/text_normalization` folder. In fact, many of NeMo's grammars actively share between.)
# + id="T58E4pU4FN3A"
LANGUAGE= "MY_LANGUAGE" # Change this to your desired language
# + colab={"base_uri": "https://localhost:8080/"} id="_PAyEPhaFpCD" outputId="23d034d2-8c93-4e8b-e3ce-5ba9e870f82d"
# %cd nemo_text_processing/inverse_text_normalization/
# !mkdir {LANGUAGE}
# !mkdir "{LANGUAGE}/taggers"
# !mkdir "{LANGUAGE}/verbalizers"
# !mkdir "{LANGUAGE}/data"
# %cd {LANGUAGE}
# !pwd && ls
# -
# ## Dependencies
# + [markdown] id="O1vfz-bUFpwz"
# All WFSTs deployed in NeMo inherit from the `GraphFst` class.
# While in most cases you can simply import from a pre-existing `graph_utils.py`, you may occasionally find it helpful for deployment to keep a copy
# in your working directory for language specific edits. (For our purposes, we will be utilizing `nemo_text_processing.text_normalization.en.graph_utils`, which serves as default for NeMo's grammars.)
#
# You may also wish to keep a copy of `utils.py` (found in each language system's directory)
# in your working directory to assist with pathing. (Make sure to adjust the imports towards your language.)
# + colab={"base_uri": "https://localhost:8080/"} id="3OME84EmOQ4h" outputId="6eea17f9-aae9-4176-ae35-3d1f0e94b4ea"
# !cp ../../text_normalization/en/graph_utils.py .
# !cp ../../text_normalization/en/utils.py .
# ! cd ../../..
# -
# For development, we utilize `nemo_text_processing` and `pynini` (a Python library for efficient WFST construction and traversal, installed with `NeMo-toolkit` by default).
# While this tutorial will attempt to make use of `pynini` tools transparent, it does assume some familiarity with its syntax. For a more in-depth guide, the following will provide a function overview:
#
# - [<NAME>, Pynini: A Python library for weighted finite-state grammar compilation](https://aclanthology.org/W16-2409.pdf)
# - [<NAME>, Pynini Tutorial](http://wellformedness.com/courses/pynini/)
# - [Pynini Documentation](https://www.openfst.org/twiki/bin/view/GRM/PyniniDocs)
# We will also import the `pynutil` module for access to some extra functionality, along with writing a simple helper function for printing `pynini` graphs through the previously discussed 'shortest-path' heuristic.
# + id="sz18Ui8-8Kf4"
from pynini.lib import pynutil
def apply_fst(text, fst):
""" Given a string input, returns the output string
produced by traversing the path with lowest weight.
If no valid path accepts input string, returns an
error.
"""
try:
print(pynini.shortestpath(text @ fst).string())
except pynini.FstOpError:
print(f"Error: No valid output with given input: '{text}'")
# -
# # Cardinal WFST <a id="cardinal-wfst"></a>
# + [markdown] id="rOyLZb9DgLoh"
# The vast majority of ITN tasks require the ability to recognize and denormalize numbers. As such, we will begin with developing a Classifier and Verbalizer for Cardinal (integer) numbers. (e.g. `-3,-2,-1,0,1,2,3,4,5....99,100,101...`)
# + [markdown] id="9GZQkH1V89kh"
# ## Grammar
# -
# We will begin by first constructing a Cardinal WFST, using French as an example language. While your target language will obviously differ greatly from our example, you will likely find some several similarities, such as:
# - Use of a (semi) regular decimal (base-10) counting system. (A common - but not universal - feature of natural languages.)
# - Incorporation of several irregularities requiring contingencies in our WFST construction. (e.g. a pseudo vigesimal (base-20) series.)
# - Use of gender and number agreement in enumeration.
# ### Digits
# + [markdown] id="NzJ2DIwc_TT3"
# We shall begin with the first decimal place. As these numbers serve as the building blocks for the rest of our WFST, we shall begin by explicitly calling their WFST mappings with `pynini.string_map`:
# + id="u0H4qg4BjYfB"
zero = pynini.string_map([("zéro","0")]) # French only pronounces zeroes as stand alone
digits = pynini.string_map([ # pynini function that creates explicit input-output mappings for a WFST
("un","1"),
("une","1"),
("deux","2"),
("trois","3"),
("quatre","4"),
("cinq","5"),
("six","6"),
("sept","7"),
("huit","8"),
("neuf","9")
])
# + [markdown] id="0nHjY-NNjdWQ"
# We may also simply write a `tsv` file in a separate data folder
# -
# - zéro 0
# - un 1
# - une 1
# - deux 2
# - trois 3
# - quatre 4
# - cinq 5
# - six 6
# - sept 7
# - huit 8
# - neuf 9
# + [markdown] id="xicKcZLEzQTg"
# and import with `string_file`
# -
# `digits = pynini.string_file("data/digits.tsv")`
#
# If utils.py is in working directory you may also use `get_abs_path`, which will always call paths relative to your {LANGUAGE} directory:
#
# `from nemo_text_processing.inverse_normalization.{LANGUAGE}.utils import get_abs_path`
#
# `digits = pynini.string_file(get_abs_path("data/digits.tsv"))`
# + [markdown] id="yPccmicQkYAB"
# While we will use `string_map` throughout this tutorial, please note that NeMo employs the later option for maintainability and recommends its use instead.
# -
# ### Teens
# + [markdown] id="FQJiJcVMrNmC"
# Let us consider our next set of numbers:
# - 10 - dix
# - 11 - onze
# - 12 - douze
# - 13 - treize
# - 14 - quatorze
# - 15 - quinze
# - 16 - seize
# - 17 - dix-sept
# - 18 - dix-huit
# - 19 - dix-neuf
#
# Like before, we can simply use `string_map` to compose a WFST for them. But note how there is some redundancy in the number set: `17`, `18`, and `19` are all of the form `dix + digit`. It would be more efficient simply to reuse our prior WFST in these cases than simply creating new arcs, states, and weights.
#
# We can achieve this using pynini's string concatenation function to extend the accepted input strings. First we will create an WFST for `11-16`.
# + id="orSgBwyXsfY5"
teens = pynini.string_map([
("onze","11"),
("douze","12"),
("treize","13"),
("quatorze","14"),
("quinze","15"),
("seize","16"),
])
# + [markdown] id="s1yIgigdtriQ"
# Now, we will create a `tens` WFST that is responsible for mapping all instances of "dix" and concatenate (accomplished with the overloaded `+` operator) with the prior `digits` WFST. (Deleting any possible hyphens in-between with a build in `delete_hyphen`.)
# + id="CzwZrFCkt87W"
tens = pynini.string_map([("dix", "1")])
delete_hyphen = pynini.closure(pynutil.delete("-"), 0, 1) # Applies a closure from 0-1 of operation. Equivalent to regex /?/
graph_tens = tens + delete_hyphen + digits
# + [markdown] id="2knCwybmuTDn"
# We now can combine the `teens` and `graph_tens` WFST together through the union operation (done with the overloaded `|` operator), allowing our choice of either graph.
# + id="WIRJ4PE7uRrl"
graph_tens_and_teens = graph_tens | teens
# + [markdown] id="TGkzKoeuxbeA"
# Let's see if it works through the string function:
# + colab={"base_uri": "https://localhost:8080/"} id="v2iD0_HnxdUV" outputId="1d8f434f-ff8a-4c85-b8d0-1127e4587ddf"
apply_fst("dix-huit", graph_tens_and_teens)
apply_fst("seize", graph_tens_and_teens)
apply_fst("dix", graph_tens_and_teens)
# + [markdown] id="Yh2f-3rux8_2"
# The first two worked, but why did we get an error with "dix"? If you look back, you'll notice that while our graph has a mapping from "dix" to `1` - the concatenation with `digits` makes the assumption that some input from those strings will follow. That is, we left no opportunity for an *omission* of `digits`.
#
#
# + [markdown] id="OM_eJYlV1UVp"
# 
# + [markdown] id="M4xCMKRA1Wzw"
# You may also note that this issue would hold also if we wanted to normalize only digits - our graph would error out since it's expecting a `tens` or input first.
#
# + [markdown] id="XJHnlJCm1dPv"
# We can fix both of these problems by allowing an option to simply insert a zero without any extra input. (Much like our "cent" example.)
# + [markdown] id="9_vvJ9Bl1dYQ"
# 
# + [markdown] id="hJq3uoMN2OcC"
# This may be accomplished through use of the `pynutil.insert` function:
# + id="7h9xuNfA081P"
graph_digits = digits | pynutil.insert("0") # inserts zero if no digit follows
# + [markdown] id="fA_L_6Ky2SHm"
# And for `graph_tens`:
# + id="jelVA81o2RXu"
tens = tens | pynutil.insert("0") | tens + delete_hyphen
graph_tens = tens + graph_digits
# + [markdown] id="Gb5uhpGr3I4X"
# Bringing everything together:
# + id="bLkDddkA3Stu"
graph_teens_and_tens = graph_tens | teens
graph_all = graph_teens_and_tens | zero
# + [markdown] id="DESDKScv3r3P"
# Let us now check our tests:
# + colab={"base_uri": "https://localhost:8080/"} id="7wrDNXuD3oh9" outputId="661d2526-5aa0-4640-9285-bca15cd56c75"
apply_fst("dix-huit", graph_all)
apply_fst("seize" , graph_all)
apply_fst("dix" , graph_all)
apply_fst("une" , graph_all)
apply_fst("trois" , graph_all)
apply_fst("quatre" , graph_all)
apply_fst("zéro" , graph_all)
# + [markdown] id="Tz_k3NoB66Bv"
# Now we have no more error - albeit at the cost of leading zeroes. (We will take care of this later in the section.)
# -
# ### Tens
# + [markdown] id="2dJZAhE57an3"
# Now that we've taken care of the teens, we can proceed with the rest of the tens. Like many languages, French employs a (fairly) regular schema of: `tens_digit + ones_digit` for 20-100. Indeed, we can summarize 20-69 in the following template:
#
# - 20 - vingt
# - 21 - vingt-et-un
# - 22 - vingt-deux
# - 23 - vingt-trois
# - 24 - vingt-quatre
# - 25 - vingt-cinq
# - 26 - vingt-six
# - 27 - vingt-sept
# - 28 - vingt-huit
# - 29 - vingt-neuf
# - 30 - trente
# - 31 - trente-et-un
# - 32 - trente-deux
# - 33 - trente-trois
# ...
# - 40 - quarante
# ...
# - 50 - cinquante
# ...
# - 60 - soixante
# ...
# + [markdown] id="BuaxVG35UKcs"
# Expanding `tens` is fairly easy to accommodate this template: we simply extend our earlier `string_map` for the new terms in the 'tens place.' From there, we once again concatenate the `digits` WFST (along with a simple WFST to delete the occurence of the "-et-" term that occasionally occurs.)
# + id="qAnXlRkR32wt"
tens = pynini.string_map([
("dix", "1"),
("vingt","2"),
("trente","3"),
("quarante","4"),
("cinquante","5"),
("soixante","6"),
])
graph_et = pynutil.delete("-et-")
tens = tens | pynutil.insert("0") | tens + pynutil.delete("-") | tens + graph_et
graph_tens = tens + graph_digits
graph_teens_and_tens = graph_tens | teens
graph_all = graph_teens_and_tens | zero
# + [markdown] id="-hJwqPDx8I2R"
# #### Special Cases: 70-99
# + [markdown] id="zvBLvJdY9XPA"
# However, things get tricky once we go beyond the 60s. Here, standard French possesses a notorious psuedo-vigecimal (base-20) system. For numbers 70-99:
#
# - 70 - soixante-dix <- Literally in English: "sixty-ten"
# - 71 - soixante-et-onze <- Literally in English: "sixty-and-eleven"
# - 72 - soixante-douze
# - 73 - soixante-treize
# - 74 - soixante-quatorze
# - 75 - soixante-quinze
# - 76 - soixante-seize
# - 77 - soixante-dix-sept
# - 78 - soixante-dix-huit
# - 79 - soixante-dix-neuf
# - 80 - quatre-vingts <- Literally in English: "four-twenties"
# - 81 - quatre-vingt-un
# - 82 - quatre-vingt-deux
# - 83 - quatre-vingt-trois
# - 84 - quatre-vingt-quatre
# - 85 - quatre-vingt-cinq
# - 86 - quatre-vingt-six
# - 87 - quatre-vingt-sept
# - 88 - quatre-vingt-huit
# - 89 - quatre-vingt-nuef
# - 90 - quatre-vingt-dix <- Literally in English: "four-twenties-ten"
# - 91 - quatre-vingt-onze
# - 92 - quatre-vingt-douze
# - 93 - quatre-vingt-treize
# - 94 - quatre-vingt-quatorze
# - 95 - quatre-vingt-quinze
# - 96 - quatre-vingt-seize
# - 97 - quatre-vingt-dix-sept
# - 98 - quatre-vingt-dix-huit
# - 99 - quatre-vingt-dix-neuf
# + [markdown] id="HQNiwFDyVV_3"
# As before, we want to take advantage of as much redundancy as we can without creating additional ambiguities that will impede graph traversal.
#
# We first note that - despite repeating prior words - "quatre-vingt" can be mapped to `8` without introducing ambiguity. This is because, despite "quatre" and "vingt" being present in our prior graphs, our WFST has no pathing for them in this exact order. As such, we can simply add it to `tens` and immediately improve our coverage for 81-89.
# + id="AvJqaHhE9Wbd"
tens = pynini.string_map([
("dix", "1"),
("vingt","2"),
("trente","3"),
("quarante","4"),
("cinquante","5"),
("soixante","6"),
("quatre-vingt", "8")
])
tens = tens | pynutil.insert("0") | tens + delete_hyphen | tens + graph_et
graph_tens = tens + graph_digits
graph_teens_and_tens = graph_tens | teens
graph_all = graph_teens_and_tens | zero
# + [markdown] id="0_DtcpZxZTzX"
# Of course, now we permit the occurence of:
# + colab={"base_uri": "https://localhost:8080/"} id="V2leANlDhCvj" outputId="db8d5d02-c848-4e50-df23-d8499538281c"
apply_fst("quatre-vingt", graph_all)
# + [markdown] id="_X_ef3sihCHH"
# which is invalid (French uses the plural "quatre-vingt**s**" here.)
# + [markdown] id="vgKT903Y6rIQ"
# Should we alter the grammar because of this? Such a decision will largely be dependent on your intended implementation and design aims. If you see the question of 'legal' tokens as a responsibility of your upstream model, then there is no need for any alteration: "quatre-vingt" as a standalone token will simply not occur, so there is no input to be concerned with.
#
# However, if your ITN grammars are developed for an environment with low-fidelity ASR and/or where mistaken transcriptions incur heavy loss (e.g. ASR for driving directions, telephone-numbers, banking) then you may wish to err on the side of caution.
# + [markdown] id="Hf_FghLT7jdY"
# If we wanted to go for the latter, we would want to mark that "quatre-vingts" maps **only** to `80`.
# + id="JliFTF3mZSsJ"
quatre_vingt_plural = pynini.string_map([
("quatre-vingts", "80")
])
# + [markdown] id="81_b3XPbicT1"
# And that "quatre vingt" can only accompany non-zero digits:
# + id="E4_dmg6uin2j"
quatre_vingt_singular = pynini.string_map([
("quatre-vingt-", "8") # Note that the hyphen can be assumed now
])
graph_digits_without_zero = pynini.string_map([
("un","1"),
("une","1"),
("deux","2"),
("trois","3"),
("quatre","4"),
("cinq","5"),
("six","6"),
("sept","7"),
("huit","8"),
("neuf","9")
])
graph_eighties = (quatre_vingt_singular + graph_digits_without_zero) | quatre_vingt_plural
# + [markdown] id="mL7jpekV8VgP"
# For the `70`'s and `90`'s, we would likewise need to form exclusive configurations for their number series, rewriting digits to recognize "onze", "douze", "treize"... as `1,2,3....` (Note, we'll have to separate `71` and `91` to manage "soixante-**et**-onze" vs. "quatre-vingt-onze".)
# + id="y3dYkwK29zCX"
seventy_and_ninety = pynini.string_map([
("soixante-dix", "70"),
("quatre-vingt-dix", "90"),
])
seventy_and_ninety_tens = pynini.string_map([
("soixante-", "7"),
("quatre-vingt-", "9"),
])
seventy_and_ninety_one = pynini.string_map([
("soixante-et-onze", "71"),
("quatre-vingt-onze", "91"),
])
seventy_and_ninety_digits = digits = pynini.string_map([
("douze","2"),
("treize","3"),
("quatorze","4"),
("quinze","5"),
("seize","6"),
("dix-sept","7"), # For 97-99, digits are used as normal.
("dix-huit","8"),
("dix-neuf","9")
])
graph_seventies_and_nineties = (seventy_and_ninety_tens + seventy_and_ninety_digits) | seventy_and_ninety | seventy_and_ninety_one
# + [markdown] id="4NCrCwEH9HVg"
# Now we union them with our original `tens` series:
# + id="psGCgxaH-btn"
tens = pynini.string_map([
("dix", "1"),
("vingt","2"),
("trente","3"),
("quarante","4"),
("cinquante","5"),
("soixante","6"),
])
tens = tens | pynutil.insert("0") | tens + delete_hyphen | tens + graph_et
graph_tens = tens + graph_digits
graph_tens_with_special_cases = graph_tens | graph_seventies_and_nineties | graph_eighties
graph_teens_and_tens = graph_tens_with_special_cases | teens
graph_all = graph_teens_and_tens | zero
# + [markdown] id="xWjSAGRX_s0H"
# Making sure test cases work:
# + colab={"base_uri": "https://localhost:8080/"} id="kapWmgos-xcn" outputId="5e9c6f5c-1450-495f-cadf-2945355b651c"
apply_fst("quatre-vingt-treize" , graph_all)
apply_fst("quatre-vingts", graph_all)
apply_fst("quatre-vingt-deux", graph_all)
# + [markdown] id="hNUepfKZ_vS_"
# And the other cases fail as expected:
# + colab={"base_uri": "https://localhost:8080/"} id="wo2pCOXGAgYn" outputId="0bbe2792-8bc9-40f7-dd28-4745bd1390e3"
apply_fst("quatre-vingt", graph_all)
# + [markdown] id="4VPuCTTtigh-"
# Of course, there are other ways we could have reconfigured the grammar: we could simply make specific graphs for multiples of ten (`10,20,30..`) and all cases where "-et-" occurs (`21,31,41,51...91`).
#
# But this ignores a more important question: was any of this necessary in the first place? All these extra grammars did was simply expand coverage for thirty additional cardinals. And they still didn't exclude all faulty inputs! Note the following cases:
# + colab={"base_uri": "https://localhost:8080/"} id="KICvpeewCFyH" outputId="174dd910-7329-4a5f-a5b0-5e796a174217"
apply_fst("dix-une", graph_all) # supposed to be "onze"
apply_fst("dix-deux", graph_all) # supposed to be "douze"
apply_fst("vingt-un", graph_all) # supposed to be "vingt-et-un"
apply_fst("trente-un", graph_all) # supposed to be "trente-et-un"
# + [markdown] id="0D130jIVCLp2"
# We *still* need to address possible edge cases!
#
# All of this is to say that knowing your input domain before construction is imperative, as small decisions can easily determine your output range later down the line.
#
# Indeed, if you're particularly concerned with limiting input possibilities, it may be valid simply to write all unique options within a `string_map`. While a tad inelegant, it certainly assists in controlling your outputs.
# + id="RSp9w5ayA9ii"
graph_tens_special = pynini.string_map([
("soixante-dix", "70"),
("soixante-et-onze","71"),
("soixante-douze","72"),
("soixante-treize","73"),
("soizante-quatorze","74"),
("soixante-quinze","75"),
("soixante-seize","76"),
("soixante-dix-sept","77"),
("soixante-dix-huit","78"),
("soixante-dix-neuf","79"),
("quatre-vingts", "80"),
("quatre-vingt-un", "81"),
("quatre-vingt-une", "81"),
("quatre-vingt-deux","82"),
("quatre-vingt-trois","83"),
("quatre-vingt-quatre","84"),
("quatre-vingt-cinq","85"),
("quatre-vingt-six","86"),
("quatre-vingt-sept","87"),
("quatre-vingt-huit","88"),
("quatre-vingt-neuf","89"),
("quatre-vingt-dix","90"),
("quatre-vingt-onze","91"),
("quatre-vingt-douze","92"),
("quatre-vingt-treize","93"),
("quatre-vingt-quatorze","94"),
("quatre-vingt-quinze","95"),
("quatre-vingt-sieze","96"),
("quatre-vingt-dix-sept","97"),
("quatre-vingt-dix-huit","98"),
("quatre-vingt-dix-neuf","99"),])
# + [markdown] id="NUPs1qOUg-hE"
# Which is more efficient? Once again, it is dependent on your language and implementation. If we simply vizualize each graph and their number of states:
# + colab={"base_uri": "https://localhost:8080/"} id="sQ9GsIkNzxsU" outputId="d70ca927-9c43-4f49-846c-c181e725e011"
constructed_version = (graph_seventies_and_nineties | graph_eighties)
constructed_version.num_states()
# + colab={"base_uri": "https://localhost:8080/"} id="Xsgdu5TYx09_" outputId="5812912f-883b-42e8-afbf-3ec4a0170345"
string_map_version = graph_tens_special
string_map_version.num_states()
# + [markdown] id="9jzn_U7s0Sit"
# We see that their number of states (graph vertexes) are almost equal. Yet, if we use `pynini.optimize` - a method that calls a suite of WFST minimization algorithms:
# + colab={"base_uri": "https://localhost:8080/"} id="7YtqhOY90iF0" outputId="26f0f51b-b00d-4f5a-9b2f-330c9812666a"
constructed_version.optimize()
constructed_version.num_states()
# + colab={"base_uri": "https://localhost:8080/"} id="y93SqnOf0qa8" outputId="74efcbfa-a272-4fc6-e36e-f1e31c6df221"
string_map_version.optimize()
string_map_version.num_states()
# + [markdown] id="2cTdQj9L0xhl"
# We see the latter possessing a significantly larger amount of graph vertices.
#
# So the decision will be dependent on your ITN needs, language, concern with efficiency, and design philosophy. Further, even decisions of language dialect will have an influence.
# (e.g. Belgian, Canadian, and Swiss dialects of French will dispense with elements of the vigecimal system for the decimal schema.)
#
# **N.B.** *For reference: while `nemo_text_processing` grammars aim to minimize invalid productions, they assume input tokens are valid strings for a target language. (e.g. The mapping of "quatre-vingt" to `80` is permitted since it is not likely to occur in a valid French string.)*
# + [markdown] id="V1djCnvY3CjW"
# For more information on optimization algorithms for WFSTs, please see:
#
# - [<NAME>,"Generic epsilon-removal and input epsilon-normalization algorithms for weighted transducers"](https://cs.nyu.edu/~mohri/pub/ijfcs.pdf)
# - [<NAME>, "Weighted automata algorithms"](https://cs.nyu.edu/~mohri/pub/hwa.pdf)
# - [<NAME>, "Programming techniques: regular expression search algorithm"](http://www.oilshell.org/archive/Thompson-1968.pdf)
#
# -
# ### Hundreds
#
# + [markdown] id="dqPUdVBbi6gU"
#
# Moving on to the case of three digit cardinals ("hundreds"), it is likely that your chosen language becomes more regular in its schema. For instance, practically all French numbers `>100` obey the following:
#
# - `digit_from_1_to_9 + word_for_hundred + digit_from_1_to_99`
#
# For example:
# - `203` - "deux-cent-trois"
# - `530` - "cinq-cent-trente"
# - `880` - "huit-cent-quatre-vingt"
#
# As such, we can write a simple `hundreds` WFST as:
# + id="lOt-gc-FiF-X"
hundreds = graph_digits + delete_hyphen + pynutil.delete("cent") + delete_hyphen + graph_all
# + colab={"base_uri": "https://localhost:8080/"} id="Fyn1uL_NoEiz" outputId="d491680b-1b3e-4762-8470-497833b82b0e"
apply_fst("deux-cent-trois", hundreds)
apply_fst("huit-cent-quatre-vingts", hundreds)
apply_fst("cinq-cent-trente" , hundreds)
# + [markdown] id="qDjq_KfnoD5C"
# Indeed, the use of French only presents two complications:
# - French uses *only* the word "cent" for `100`. (Instead of "un cent".)
# - 'Pure' multiples of a hundred (`200,300,400....`) use the plural "cents".
#
# The second one is the easier of the two so let's start there. There are actually two options open to us. First, we could treat "cents" the same way as we did "cent" in the base case and simply delete it. From there, the lack of any following inputs will allow the WFST to insert the trailing zeroes as appropriate.
# + id="m2F-sumbxqLE"
cents = pynini.accep("cent") | pynini.accep("cents") # Creates a Finite State (Accep)tor, mapping inputs back to themselves
hundreds = graph_digits + delete_hyphen + pynutil.delete(cents) + delete_hyphen + graph_all
# + [markdown] id="VisQu_Etx-QB"
# Or we can use it as a cue to 'shortcut' the WFST to immediately insert zeroes.
# + id="VspiTN5Vxxjl"
graph_cents = pynini.cross("cents", "00") # Creates a single input-output mapping
hundreds = graph_digits + delete_hyphen + ((pynutil.delete("cent") + delete_hyphen + graph_all) | graph_cents)
# + [markdown] id="meVn5BiyyX5v"
# For the case of solitary "cent", we need to make sure our output is `1` only in the case that no digit preceeds the occurence. Here we need to be confident in the structure of our WFST and that any possible ambiguity has been dealt with by this point. (Something to keep in mind as we move to the thousands.)
# + id="277Z-zLWyWAf"
graph_cent = pynini.cross("cent", "1")
graph_hundreds_first_digit = (graph_digits + delete_hyphen + pynutil.delete(cents)) | graph_cent
graph_hundreds = graph_hundreds_first_digit + delete_hyphen + graph_all
# + colab={"base_uri": "https://localhost:8080/"} id="FNZlJsvS_Yvt" outputId="e85ae561-e7a1-4b6a-e394-f0194fdb89e7"
apply_fst("trois-cents", graph_hundreds)
apply_fst("cent", graph_hundreds)
apply_fst("cent-trois", graph_hundreds)
# -
# ### Thousands
# + [markdown] id="e7Dy5slLzp-K"
# For quite a few languages, managing the WFST for the thousands place is the last aspect to figure out, as the higher powers of ten reuse the same schema. (For those working with counting systems that reserve special terms for "ten-thousand" (e.g. Chinese derived counting systems), you may need to extend unique coverage to the next power of ten.)
#
# For French, the question of thousands is rather simple: `digits_from_1_to_999 + mille + digits_from_1_to_999`
#
# With only the exception that any expression of one thousand drops a leading digit.
# - `1,000` -> "mille"
# - `1,001` -> "mille-un"
# + id="AvsnAAiPzlu_"
graph_one_thousand = pynini.cross("mille", "1")
graph_many_thousand = graph_hundreds + delete_hyphen + pynutil.delete("mille")
graph_thousands = (graph_one_thousand | graph_many_thousand) + delete_hyphen + graph_hundreds
# + colab={"base_uri": "https://localhost:8080/"} id="i3m9TG7Y4tkl" outputId="d3f1f81d-c463-4934-9df7-3b8f2b67798f"
apply_fst("cent-mille-deux-cents", graph_thousands)
apply_fst("deux-cent-mille-deux-cents", graph_thousands)
# + [markdown] id="NoevSTZGGT17"
# ### Weighting
# + [markdown] id="A2gcVIZM0-iv"
# Question: will this cover all our grammar so far? (Hint: what assumptions were made about "cent"/"cents"?)
#
# + colab={"base_uri": "https://localhost:8080/"} id="cCFtPhr1BjAc" outputId="048e0d93-a4a8-4f4e-d461-bfd70e911aff"
apply_fst("deux-mille-un", graph_thousands)
# + [markdown] id="Ne-7L9Cd4t-8"
# Once again, we need to introduce the possibility of the prior power of ten not occuring in the string. There must be an option for simply inserting a string of `0` in place of the omitted "cent".
# + [markdown] id="iockqXdn-aG4"
# Further, we want to be careful with how cavalier we have been with insertions. Consider the following:
# + colab={"base_uri": "https://localhost:8080/"} id="bxJlSnj2-Xw3" outputId="6722e5ef-8a7f-43e1-84fe-b3f5f18307e1"
apply_fst("mille-cent-un", graph_thousands) # Should be 1101
apply_fst("mille-cent", graph_thousands) # 1100
# + [markdown] id="fq5zEayA-kOx"
# It appears that our WFST has developed a tendency to simply 'ignore' some of these higher powers. Let us return to our code for `graph_hundreds` and `graph_thousands`.
# + id="S2aV1KQ4-1iP"
graph_cents = pynini.cross("cents", "00")
graph_cent = pynini.cross("cent", "1")
graph_hundreds_first_digit = (graph_digits + delete_hyphen + pynutil.delete(cents)) | graph_cent
graph_hundreds = (graph_hundreds_first_digit + delete_hyphen | pynutil.insert("0")) + graph_all
graph_one_thousand = pynini.cross("mille", "1")
graph_many_thousand = graph_hundreds + delete_hyphen + pynutil.delete("mille")
graph_thousands = (graph_one_thousand | graph_many_thousand) + delete_hyphen + graph_hundreds
# + [markdown] id="9avwOIkk-9qt"
# Recall that throughout we have provided options for simply inserting zeroes in the case of omitted numbers? That tendency has finally caught up with us. The use of our previous `graph_hundreds` in `graph_many_thousands` now allows our graph to insert a string of `0`'s without penalty.
#
# You may note that this is very similar to the "cents" example brought up at the beginning, presenting a similar solution. We can control this output by making it too costly to traverse unless absolutely necessary for the graph. This can be accomplished simply by appending a weight to the insertion for hundreds:
# + id="MQG3j0U8CUAQ"
graph_hundreds = (graph_hundreds_first_digit + delete_hyphen | pynutil.insert("0", weight=.1)) + graph_all
graph_one_thousand = pynini.cross("mille", "1")
graph_many_thousand = graph_hundreds + delete_hyphen + pynutil.delete("mille")
graph_thousands = (graph_one_thousand | graph_many_thousand) + delete_hyphen + graph_hundreds
# + colab={"base_uri": "https://localhost:8080/"} id="KNHhrYZ7Ca58" outputId="a7d07372-733d-4837-c1e9-1dc58ba2b87c"
apply_fst("mille-cent-un", graph_thousands)
apply_fst("mille-cent", graph_thousands)
# + [markdown] id="51yPEaf2EkbD"
# Why choose a weight of `.1`? Quite simply: it's arbitrary. As mentioned earlier, the default graph in `pynini` is a tropical semiring, which uses the `min` function to select among two arcs for path traversal. Since all our paths so far are weight `0`, any positive value will ensure that it is a last option among path traversal. (Note, this conversely entails any negative weight path will be prioritized.)
#
# That we chose this number as a small value comes from a place of caution: the tropical semiring uses an additive function to calculate the total weight of an entire path to traverse a WFST. As our grammars can easily become massive, this means that small weights can have major impact down the line. Further, by constraining path weights to small values, we can have general certainty towards the maximum weight of any individual graph, allowing us to add constraints regarding maximum token length and token hierarchy. (As explained in [later sections](#classifyweights).) As such, when using weights in a localized setting, it is best to use small values to avoid unforseen escalation.
# + [markdown] id="iScKgvRxGt-B"
# ### Higher Powers
#
# + [markdown] id="rtHEd6OE2WSg"
# At this point, we can propose a general heuristic with escalating to higher powers of ten: they always need a way for their absence to be accomodated in the WFST. Further, they require some weighting to prevent this absence from developing into a string of omitted values. To aviod further bumps, we'll take care of this now with `graph_thousands`.
# + id="iZMN7wcE2lH5"
graph_one_thousand = pynini.cross("mille", "1")
graph_many_thousand = graph_hundreds + delete_hyphen + pynutil.delete("mille")
graph_thousands = (graph_one_thousand | graph_many_thousand | pynutil.insert("000", weight=.001)) + delete_hyphen + graph_hundreds
# + [markdown] id="Fkc3LIH824P7"
#
# For the rest of French (and many other languages), the rest of the work is simply repeating the prior pattern for the thousands element:
# `hundreds + word_for_higher_power + hundreds.....` Of course there will be some variation in this schema, but the recursion should be regular. (It is rather rare that languages appropriate unique forms for these higher counts.)
# + [markdown] id="qGnK4ARX4Nay"
# To finish French, we can list off the following equivalent for higher powers of ten:
# - `million` - "million/millions"
# - `billion` - "milliard/milliards"
# - `trillion` - "billion/billions"
#
# Like the "cent/cents" rule, these values alternate with a plural form in the case of multiples of the value. Writing them out:
# + id="sBu7-dub4vxz"
millions = pynini.accep("million") | pynini.accep("millions")
graph_millions = ((graph_hundreds + delete_hyphen + pynutil.delete(millions) + delete_hyphen) | pynutil.insert("000", weight=.1) # We need three zeroes now
) + graph_thousands
# + id="LmMeCHXr5Bb5"
billions = pynini.accep("milliards") | pynini.accep("milliard")
graph_billions = ((graph_hundreds + delete_hyphen + pynutil.delete(billions) + delete_hyphen)| pynutil.insert("000",weight=.1) # We need three zeroes now
) + graph_millions
# + id="CIRIeQEg5B0J"
trillions = pynini.accep("billion") | pynini.accep("billions")
graph_trillions = ((graph_hundreds + delete_hyphen + pynutil.delete(trillions) + delete_hyphen) | pynutil.insert("000",weight=.1) # We need three zeroes now
) + graph_billions
# + [markdown] id="sRNUPx-15J1v"
# Bringing all together:
# + id="0dLOWm_B5SwQ"
graph = graph_trillions | zero
# + [markdown] id="nBFE3BrN6IPR"
# Let's try it out:
# + colab={"base_uri": "https://localhost:8080/"} id="6lWwtR1S6LI4" outputId="3a6740ee-9e92-4500-c2c8-965131167e58"
example = "deux-cent-milliard-quatre-million-deux-cent-quatre-vingt-onze"
apply_fst(example, graph)
# -
# ### Finishing Touches
# + [markdown] id="-w3KgX6C6mff"
# Now that we have our cardinal in place, we can take care of that stylistic issue of the leading zeroes. For this, we want to develop a 'filter' that deletes all zeroes preceeding the first non-zero in the string, and leave the rest 'as is.'
#
# First let us create the filter by calling on `NEMO_DIGIT`- a `graph_util` WFST that only permits digits as input. With it, we'll create a WFST that will delete all leading zeroes in a sting. We then compose this (using `@`) onto our original graph, creating a new graph that accepts inputs from our original by outputs only outpus of `clean_cardinal`.
# + colab={"base_uri": "https://localhost:8080/", "height": 290} id="EA4VnRe6FO-2" outputId="59e412b3-a445-4172-ee64-b0f80281a167"
delete_leading_zeroes = pynutil.delete(pynini.closure("0")) # will delete all zeroes under closure. Equivalent to regex * operator
stop_at_non_zero = pynini.difference(NEMO_DIGIT, "0") # creates a graph that accepts all input-outputs from NEMO_DIGIT except 0
rest_of_cardinal = pynini.closure(NEMO_DIGIT) # accepts all digits that may follow
clean_cardinal = delete_leading_zeroes + stop_at_non_zero + rest_of_cardinal
clean_cardinal = clean_cardinal | "0" # We don't want to ignore the occurence of zero
graph = graph @ clean_cardinal
# + [markdown] id="piP9nqQkHpo3"
# Now our WFST will output our numbers as normal:
# + id="dnQ9odSpIAB7"
apply_fst(example, graph)
# -
# ### Final Notes
#
# + [markdown] id="p7zt8lVsK2rY"
# We have finally formulated a grammar that will process French cardinals into numeric representation. Of course, not every grammar you write will be for French. But several of the principles we've worked through will be invaluable in your own development. Before moving on, here's a quick summary of (almost) universal points to take away for WFST construction.
# - Decide at the beginning of construction the level of constraint you wish for your grammar. Is it necessary to have a specific domain or can you rely on upstream models to narrow your input possibilities for you?
# - Work iteratively upwards from the smallest place value of your numeric system. This will assist you in forming building blocks for larger values.
# - Always allow for the possibility of omission of previous place values. (Not every number in the thousands will contain mention of the hundreds place.)
# - For each place value, consider how the sub-grammar will affect the preceeding and following place values. Are there exceptions that you've built into the grammar that may become problematic later on?
# - Utilize weights for default insertions to limit path traversal to only final options. When doing so, use small values to avoid escalating problems in your larger grammar.
# + [markdown] id="nvyHg1bQIIHD"
# With that handled, we can move on to converting this grammar into a Classifier.
# + [markdown] id="gJ1YJUvhIZwm"
# ## Classifier
# + [markdown] id="q2L2x0crIeXQ"
# Now that we have a grammar that will convert individual tokens into number strings, we now want to focus on building it into a classifier to properly tag candidate tokens. This requires a couple of properties:
# - It recognizes any valid token and permits traversal through the WFST graph
# - Conversely, it does not allow invalid tokens to traverse the WFST graph
# - It properly disambiguates overlap among ambiguous cases
# - It attributes the proper attributes to a classified token
#
# While this seems like a lot, in practice this just means that your grammar will need a few more tweaks to improve exclusivity.
# + [markdown] id="ArEYn7RWKcYI"
# NeMo ITN performs token classification through a series of `GraphFst` classes and assumes deployment of your grammars through an object that inherits from this class. As such, you will need to instantiate your grammar as a `CardinalFST`
# + colab={"base_uri": "https://localhost:8080/", "height": 368} id="GWgMSybqLqiS" outputId="597c00ae-0f62-417f-888c-88c81c24a3fc"
class CardinalFst(GraphFst):
def __init__(self):
super().__init__(name="cardinal", kind="classify")
# Rest of the grammar here
# .......
#.........
# + [markdown] id="SIE8dNQlL52G"
# While the naming convention may vary, the `name` and `kind` properties must be set accordingly to permit Sparrowhawk integration.
#
# Further, the reulting graph must produce the classified token within the following format:
# `token { cardinal { integer: "DIGIT_STRING" } }`
#
# This is accomplished by a series of string insertions:
# + id="aC_c64KSNTCg"
class CardinalFst(GraphFst):
def __init__(self):
super().__init__(name="cardinal", kind="classify")
# Rest of the grammar here
# .......
#.........
self.fst = pynutil.insert("integer: \"") + graph + pynutil.insert("\"")
# + [markdown] id="AGLQxOSzOK1F"
# Followed by a call of the parent `GraphFst.add_tokens()` method:
# + id="Jz-UXFipORps"
class CardinalFst(GraphFst):
def __init__(self):
super().__init__(name="cardinal", kind="classify")
# Rest of the grammar here
# .......
#.........
self.fst = pynutil.insert("integer: \"") + graph + pynutil.insert("\"")
final_graph = self.add_tokens(graph)
# + [markdown] id="gh23S7BHOY0r"
# Which will insert the appropriate formatting. Note that this formatting must be exact: a single space must follow each field name and each value must be within escaped double quotes.
#
# In the event that you also wish for `CardinalFst` to indicate negative values, the optional `negative: ` property may be used.
#
# For instance, French indicates negative values by prefacing the quantity with "moins." As such:
# + id="3JbTn35cOx0k"
optional_minus_graph = pynini.closure(
pynutil.insert("negative: ") + pynini.cross("moins", "\"-\"") + " ", 0, 1 # Note the extra space to separate the value from the integer field
)
final_graph = optional_minus_graph + pynutil.insert("integer: \"") + graph + pynutil.insert("\"")
# + [markdown] id="DCs1048v6N0K"
# All together, your `CardinalFst` ultimately serves as a wrapper for your grammar, save with the addition of a few insertions to assist processing:
#
#
# + id="eo6uEz1s5TJY"
class CardinalFst(GraphFst):
def __init__(self):
super().__init__(name="cardinal", kind="classify")
### Cardinal Grammar....
### .....
graph = graph_trillions | zero
### Formatting grammar....
### .....
graph = graph @ clean_cardinal
### Token insertion
optional_minus_graph = pynini.closure(
pynutil.insert("negative: ") + pynini.cross("moins", "\"-\"") + " ", 0, 1
)
final_graph = optional_minus_graph + pynutil.insert("integer: \"") + graph + pynutil.insert("\"")
final_graph = self.add_tokens(final_graph) # inserts the cardinal tag
self.fst = final_graph
# + [markdown] id="MFIMdLCoZzLK"
# Let's see a demonstration.
# + id="4CF6Iz9NZ7R_"
cardinal = CardinalFst().fst
example = "moins deux-cent-quatre"
apply_fst(example, cardinal)
# -
# ## Verbalizer
# + [markdown] id="uvUqpC_Q8FSt"
# The verbalizer can be both the most crucial and simplest part of building each grammar. On one hand, it is the component that finalizes all of your previous work. If it is unable to properly normalize your text, everything has been for naught.
#
# On the other hand, your previous work has vastly limited the unpredictability of your input. Recall from our initial demonstration of the classifier-verbalizer system that and input like <<le premier juillet il a mangé trente-cinq pommes>> becomes:
#
# - `tokens { name: "le" }`
# - `tokens { date { day: "1" month: "juillet" }`
# - `tokens { name: "il" }`
# - `tokens { name: "a" }`
# - `tokens { name: "mangé" }`
# - `tokens { cardinal { integer: "35" } }`
# - `tokens { name: "pommes" }`
#
# Part of the purpose of the two stage set-up is that the input space for each verbalizer is obvious: it's simply the name of its semiotic class. As such, we only need to write our grammar to recognize its class, remove tokens accordingly, and then manage the attributes of each semiotic token.
# + [markdown] id="et1GgmBuAWzY"
# We will begin as we did with our classifier and create a class to inherit from the `GraphFST` utility class:
# + id="NNKpgWtkAgEW"
class CardinalFst(GraphFst):
def __init__(self):
super().__init__(name="cardinal", kind="verbalize")
# + [markdown] id="OyAV39NsAqSN"
# One of the useful aspects of the `GraphFst` utility is that it already posseses a built in graph that will recognize and remove semiotic tokens: `delete_tokens`. As such we need only concern ouselves with managing the properties of the Cardinal class:
# - `integers`
# - `negative`
#
# Here, the desired written format of your chosen language will dictate how you proceed. For French, we have the following rules for Cardinal numbers:
# - A negative sign is written before the numeral.
# - Cardinal numbers representing quantities (e.g. "mille euros"/ "one thousand dollars") are written with spaces in-between every three digits. (e.g. `1 000`)
# - Cardinal numbers representing place in a sequence or addresses ("page mille"/"page one thousand") are written without spacing. (`1000`)
#
# The first property seems easy enough to handle: write a grammar that simply removes the `negative` formatting, leaving only `-`. (Recall that our Classifier only inserted the string if it was present.)
#
# For the final two, we may note that our intention to develop WFSTs for the Decimal, Measure, and Money classes already will cover most desired quantities. As such, we can leave the issue of spacing to those instances and let the Cardinal WFST default to the non-spacing case. (Note that this will be helpful with Time, Date, Telephone, Electronic, and Ordinal classes as they will not use the spacing format either. It is usually better to reserve specific formatting rules to other classes and let the Cardinal serve as a default.)
#
# As such, we just need our WFST to remove the `integer` property and `negative` property (if it occurs). These can be managed through the `pynutil.delete` function, as seen in the following:
#
# + colab={"base_uri": "https://localhost:8080/", "height": 368} id="6MF2I6SLU7nf" outputId="0437c4af-5c96-4122-8af0-ca37723c7228"
class CardinalFst(GraphFst):
def __init__(self):
super().__init__(name="cardinal", kind="verbalize")
# Removes the negative attribute and leaves the sign if occurs
optional_sign = pynini.closure(
pynutil.delete("negative:")
+ delete_space
+ pynutil.delete("\"")
+ pynini.accep("-")
+ pynutil.delete("\"")
+ delete_space,
0,
1,
)
# removes integer aspect
graph = (
pynutil.delete("integer:")
+ delete_space
+ pynutil.delete("\"")
+ pynini.closure(NEMO_DIGIT, 1) # Accepts at least one digit
+ pynutil.delete("\"")
)
graph = optional_sign + graph # concatenates two properties
delete_tokens = self.delete_tokens(graph) # removes semiotic class tag
self.fst = delete_tokens.optimize()
# + [markdown] id="QSX2KlZJbRAA"
# Let's see if it will properly render a given token:
# + id="JxaLm2k0bYIJ"
cardinal = CardinalFst().fst
example = 'cardinal { negative: "-" integer: "204" }'
apply_fst(example, cardinal)
# + [markdown] id="Bc0-QCBHWg-8"
# That's it! We've now completed all aspects of our `CardinalFst` from grammar writing to Verbalization. While we still have quite a few semiotic classes left, you will find that they build off the `CardinalFst` quite easily, making progression much simpler and straightforward.
#
# Before proceeding, there are two things to note:
# - `delete_tokens` is called on the completed graph, despite the token class occuring first in the tokenized string. This is because the function intersects with an initial WFST that deletes the tags. As such, the function must be passed a completed graph.
# - In our initial example, all tokens were enclosed within a `token` category. Insertion and deletion of this category is managed by the main [Classifier](#tokenize-and-classify) and [Verbalizer](#verbalize-and-verbalize-final) respectively and is not a concern during individual class grammar development.
# - Earlier in the tutorial we noted that NeMo ITN permutates all WFSTs unless the `preserve_order` tag is passed as part of the Classifier. This allows you to ignore possible variation in designing the verbalizer and focus on whatever form of processing is easiest for the grammar. That is, the decision to process the `negative` property before the `integer` property is not chosen because of a consequence of the French language but instead because it is easier to write out with `pynini`.
# - Conversely, if your language is completely invariant in this regard, it may be more efficient to pass `preserve_order` through the Classifier and manage the property here in the Verbalizer. This allows NeMo ITN to avoid building states and arcs for each permutation, reducing graph size and compiling time.
# + [markdown] id="aFUrbSdJ8Wk7"
# # Ordinal WFST <a id="ordinal-wfst"></a>
# + [markdown] id="w1b0Z7f5Z9Ar"
# Ordinals is the class of numbers used for enumerating order or placement of entities in a series. In some languages, they are simply derivations of cardinal numbers. For instance, English enumerates order as `first, second, third, fourth, fifth....` After the third ordinal, they become a regular pattern of `cardinal + 'th'`.
#
# Meanwhile, other languages may reserve specific counting systems for ordinals. For example, while Korean uses a Chinese derived counting system for several Cardinal related tasks, it uses derivations from a native counting system for ordering:
#
# **Cardinal**/**Ordinal** = **English**
# - il/cheot-jae = "First"
# - i/dul-jae = "Second"
# - sam/set-jae = "Third"
# - sa/net-jae = "Fourth"
# - o/daseot-jae = "Fifth"
#
# If your language is of the latter variety, you will likely need to begin development of Ordinal WFST by repeating Cardinal WFST development before proceeding. (Or make it part of your previous Cardinal WFST and combining with a `union` operation.) While you can extend coverage to the level of Cardinal WFST, you will find most Ordinals to be sufficiently covered by only enumerating to a few hundreds. (e.g. Is it common in your language to speak of the "one millionth" in an order and/or write out `1,000,000th`?)
#
# For this portion of the tutorial, we will focus on the first type of ordinals - those that primarily derived by altering Cardinals.
# + [markdown] id="oq_xA8NPiANw"
# ## Grammar
# + [markdown] id="lhjcQS6oiD_w"
# Continuing with our example language, we first begin by laying out our expected inputs and pinpointing a regular pattern to guide our WFSTs. We note the following examples:
#
# **English = French**
# - "first" = "premier/première"
# - "second" = "second/seconde/deuxième"
# - "third" = "troisième"
# - "fourth" = "quatrième"
# - "fith" = "cinquième"
# - "sixth" = "sixième"
# - "seventh" = "septième"
#
# From our examples inputs, it appears that spelling of French Ordinals follows a general format of: `cardinal + ième`. The only exceptions appear to be in the case of the first and second Ordinals - for which completely different roots appear - and the fourth and the fith Ordinals - where the former drops the "e" at the end of the root (`quatre -> quatr`) and the latter appends a "u" (`cinq -> cinqu`).
#
# For the expected outputs, we observe the following examples:
# - "premier/première" -> `1ᵉʳ/1ʳᵉ`
# - "second/seconde" -> `2ᵈ/2ᵈᵉ`
# - "deuxième" -> `2ᵉ`
# - "troisième" -> `3ᵉ`
# - "quatrième" -> `4ᵉ`
# - "cinquième" -> `5ᵉ`
# - "sixième" -> `6ᵉ`
# - "septième" -> `7ᵉ`
#
# It appears that the output is simply the cardinal number of the root with an associated superscript. Since we have already constructed the Cardinal WFST, this means that the job of constructing an Ordinal WFST is simply a case of recognizing the cardinal root for the input and then utilizing a preconstructed Cardinal grammar to render the proper form alongside an associated superscript. That is, our tasks are to:
# - Identify the proper superscript for the ordinal
# - Change the ordinal back into a cardinal
# - Use the Cardinal WFST to transform the cardinal into normalized form
# - Properly render the ordinal using the normalized cardinal and proper superscript
#
# As information regarding the superscript will need to be conveyed through development of the Classifier, we will begin with creating the grammar necessary for rendering the ordinal as its cardinal root.
#
#
# + [markdown] id="AOUVZhiwT7hE"
# ### Stripping Suffixes
# + [markdown] id="5nw0_lOTsEik"
# Since French forms Ordinals by appending a suffix to Cardinals, we should start by creating a WFST to remove the suffix. Assuming that our grammar processes one token at a time, this means that we just need an WFST that will accept all tokens that end with "ième" and then delete the suffix from that token:
# + id="Rk89LhsxsHTO"
strip_morpheme = pynutil.delete("ième") # deletes suffix
graph_strip_morpheme = NEMO_SIGMA + strip_morpheme # accepts all strings until passed suffix, then deletes suffix
# + [markdown] id="pLg-PzdntV4N"
# Now we can create a graph that permits all characters in a word token and deletes the ordinal suffix. (Note that this also means that the graph won't accept tokens without the suffix, helping us avoid false inputs.)
#
# We can now intersect this graph with our Cardinal WFST to now strip the suffixes from ordinals and treat them as cardinals. However, recall that our `CardinalFst` also inserted its own class tag. Obviously, we do not want to do this here as it will disrupt the formatting of the token. Instead, we should create a new subgraph *within* the `CardinalFst` class that will only produce the cardinals without tokens.
# -
class CardinalFst(GraphFst):
def __init__(self):
super().__init__(name="cardinal", kind="classify")
### Cardinal Grammar....
### .....
graph = graph_trillions | zero
### Formatting grammar....
### .....
graph = graph @ clean_cardinal
### NEW GRAPH
self.just_cardinals = graph # will produce cardinals without formatting
### Token insertion
optional_minus_graph = pynini.closure(
pynutil.insert("negative: ") + pynini.cross("moins", "\"-\"") + " ", 0, 1
)
final_graph = optional_minus_graph + pynutil.insert("integer: \"") + graph + pynutil.insert("\"")
final_graph = self.add_tokens(final_graph)
self.fst = final_graph
# Now we call it for our graph:
# + id="vxDgBa4_t1nD"
graph_cardinal = CardinalFst().just_cardinals
graph_ordinal_regular_suffix = graph_strip_morpheme @ graph_cardinal
# + [markdown] id="hSpk5M7BuXRz"
# Let's see if it works and gives us the desired cardinal:
# + id="7cJ7fieouY2r"
example = "sixième" # dervied from six/6
apply_fst(example, graph_ordinal_regular_suffix)
# + [markdown] id="GtEuV7sOuxek"
# Now we can consider the edge cases. Beyond the first and second ordinals, French exhibits irregular behavior in the following cases:
# - If the cardinal root ends with an "e", the "e" is dropped before adding the suffix (e.g. "quatrième").
# - Cardinals ending with "cinq", "neuf", and "dix" change their endings to "cinqu", "neuv" , and "diz" before appending the suffix, respectively.
#
# We could start by proposing a WFST that replaces the suffix "ième" with "e" and then compose this onto the Cardinal WFST. If it is a legitimate cardinal, then there will be a path through CardinalFST and the integer will be rendered as normal.
#
# Meanwhile, the case of "dix", "cinq", and "neuf" would each require a distinct WFST as they are each a consequence of different rules of orthography and phonology. Like the case with "e", we could change each back to its root and then see if the CardinalWFST will permit a path with the new input.
#
# It is at this point that we can do a cost-benefit analysis and realize that all these cases can be managed by an explicit `string_map/string_file`:
# + id="_9KTNQeIw4sq"
graph_root_change = pynini.string_map([("quatrième", "quatre"),
("cinquième", "cinq"),
("neuvième", "neuf"),
("onzième", "onze"),
("douzième", "douze"),
("treizième", "treize"),
("quatorzième", "quatorze"),
("quinzième", "quinze"),
("seizième", "seize"),
("trentième", "trente"),
("quarantième", "quarante"),
("cinquantième", "cinquante"),
("soixantième", "soixante"),
("millième", "mille"),
])
# + [markdown] id="eo2_keFVqaY4"
# We could then concatenate these with a WFST that accepts all tokens with these endings and then change the endings as desired. These will provide the cardinal roots just as effectively.
# + [markdown] id="O7I29ezmxylx"
# The same can be said for "premier/première" and "second/seconde":
# + id="3JZoz51VyGS6"
graph_firsts = pynini.string_map([("premier", "un"),("première", "un")])
graph_seconds = pynini.string_map([("second", "deux"),("seconde", "deux")])
# + [markdown] id="NJ9BGGAwyTQ5"
# *Note: We graph separately to manage their different superscripts later on.*
#
# Depending on your language of focus, the choice of implicitly reversing the root token or explicitly mapping back to root will be the most efficient, but it is worth considering both options if only to check your understanding of the language.
# + [markdown] id="8PgVwDRRq9gr"
# Putting our grammar together, we have:
# + id="ko2kAeKwrRSH"
strip_morpheme = pynutil.delete("ième") # deletes suffix
graph_root_change = pynini.string_map([("quatrième", "quatre"),
("cinquième", "cinq"),
("neuvième", "neuf"),
("onzième", "onze"),
("douzième", "douze"),
("treizième", "treize"),
("quatorzième", "quatorze"),
("quinzième", "quinze"),
("seizième", "seize"),
("trentième", "trente"),
("quarantième", "quarante"),
("cinquantième", "cinquante"),
("soixantième", "soixante"),
("millième", "mille"),
])
# Component will accept all tokens that end with desired strings
graph_get_cardinal = NEMO_SIGMA + (strip_morpheme | graph_root_change)
graph_firsts = pynini.string_map([("premier", "un"),("première", "un")])
graph_seconds = pynini.string_map([("second", "deux"),("seconde", "deux")])
graph_get_cardinal = pynini.union(graph_firsts, graph_seconds, graph_get_cardinal)
graph_cardinal = CardinalFst().just_cardinals
graph_ordinal = graph_get_cardinal @ graph_cardinal
# + id="ESxY3LsCdE8q"
apply_fst("sixième", graph_ordinal)
apply_fst("première", graph_ordinal)
apply_fst("seconde", graph_ordinal)
# + [markdown] id="qo_g8UdoUFJB"
# ## Classifier
# + [markdown] id="kemhdKAjzEIa"
# Now that we've found a way to pass the work of the Ordinal grammar back onto the Cardinal grammar, we can move onto the Classifier. Like before, we need to inherit from `GraphFst` to properly insert token formatting and required attributes. As well, we will again use the `integer` property to tag our digit string.
#
# Indeed, the only major difference between the Ordinal Classifier and the Cardinal Classifier is the replacement of optional `negative` attribute with the `morphosyntactic_feature` attribute to indicate the superscript function.
# + [markdown] id="EHM4Y3TW2nXT"
# Since we are relying on the `CardinalFst` class in our grammar, we want to consider how to instantiate an instance of it. Since our ultimate goal is to build a Classifier that unites all semiotic classes, it makes sense to simply use the `CardinalFst` that we will need to call for our ITN and pass it as an argument to our new class.
# + colab={"base_uri": "https://localhost:8080/", "height": 273} id="KsmPhWSa3LF_" outputId="9e881ca9-a926-4249-dda8-9c52175569b5"
def __init__(self, cardinal: GraphFst):
super().__init__(name="ordinal", kind="classify")
# + [markdown] id="CtBQ-udB3S5Q"
# To clear up the namespace, we will now be importing from the NeMo implementation of `CardinalFst` for French.
# + id="L-JAcidf4QQg"
from nemo_text_processing.inverse_text_normalization.fr.taggers.cardinal import CardinalFst
class OrdinalFst(GraphFst):
def __init__(self, cardinal: GraphFst):
super().__init__(name="ordinal", kind="classify")
graph_cardinal = cardinal.graph_no_exception # NeMo equivalent to self.just_cardinals
# + [markdown] id="FQfkAqZavCAB"
# We now add in our grammar:
# + id="uUQ4BLuivGut"
class OrdinalFst(GraphFst):
def __init__(self, cardinal: GraphFst):
super().__init__(name="ordinal", kind="classify")
graph_cardinal = cardinal.graph_no_exception # may replace
strip_morpheme = pynutil.delete("ième") # deletes suffix
graph_root_change = pynini.string_map([("quatrième", "quatre"),
("cinquième", "cinq"),
("neuvième", "neuf"),
("onzième", "onze"),
("douzième", "douze"),
("treizième", "treize"),
("quatorzième", "quatorze"),
("quinzième", "quinze"),
("seizième", "seize"),
("trentième", "trente"),
("quarantième", "quarante"),
("cinquantième", "cinquante"),
("soixantième", "soixante"),
("millième", "mille"),
])
# Component will accept all tokens that end with desired strings
graph_get_cardinal = NEMO_SIGMA + (strip_morpheme | graph_root_change)
graph_firsts = pynini.string_map([("premier", "un"),("première", "un")])
graph_seconds = pynini.string_map([("second", "deux"),("seconde", "deux")])
graph_get_cardinal = pynini.union(graph_firsts, graph_seconds, graph_get_cardinal)
graph_ordinal = graph_get_cardinal @ graph_cardinal
# + [markdown] id="F_6EXPRMvnp2"
# Now we come to the `morphosyntactic_features` property - a linguistic term for aspects of a word related to grammar. If intending to deploy your WFST through Sparrowhawk, this is the only ordinal property that is permitted (outside of the universal properties like `preserve_order`) and thus must carry all information regarding how to properly normalize the ordinal. (If Sparrowhawk deployment is not necessary, you may add additional properties to the tag.)
#
# How should we convey this information? Since the Verbalizer will be the main interface for our tags, it really does not matter - so long as we can reliably process the features. For the purposes of French, we just need `morphosyntactic_features` to decide the following:
# - Insert the specific superscripts for "premier/première" or "second/seconde"
# - Insert "ᵉ" otherwise
#
# We will also introduce another aspect of French Ordinals: they can be either plural or singular, identified by the suffix "s" on input and superscript "ˢ" on output. As such, our `morphosyntactic_features` should also decide the additional property:
# - Insert the plural superscript
# + [markdown] id="atctz6p-2GtV"
# Since the default superscript is near universal, we will just specify this in our WFST and focus on the second and first ordinals as specific cases. We will create a `graph_morpheme` component that inserts the default superscript - indicated with a standard "e" to avoid possible encoding issues. We will then append a WFST that will graph any possible plural marker - "s" - as part the `morphosyntactic_features`.
# + id="ui99osyP2UuQ"
graph_morpheme = pynutil.insert("e") # Insert e superscript
graph_plural = pynini.closure(pynini.accep("s"), 0, 1) # We create an acceptor since we must process the possible "s"
graph_morpheme_component = graph_morpheme + graph_plural
graph_morphosyntactic_features = (pynutil.insert(" morphosyntactic_features: \"")
+ graph_morpheme_component
)
# + [markdown] id="QAlqubA25gq0"
# Introducing the `integer` feature:
# + id="rs2TyIBc5la6"
graph_reg_ordinals = graph_get_cardinal @ graph_cardinal # Rewriting ordinals to remove the first and second ordinal.
graph_ordinal = pynutil.insert("integer: \"") + graph_reg_ordinals + pynutil.insert("\"")
graph_ordinal += graph_morphosyntactic_features
# + [markdown] id="xoqk20Pi2gT8"
# For the first and second ordinals, we can explicitly state their mappings, as these occurences are invariable. (First and second ordinals do not need to accomodate being the endings of other terms.) As such, we can just have mappings from the token to the superscripts.
# + id="54aqdH_P63Ea"
firsts = pynini.string_map([("premier", "er"), ("première","re")])
firsts += graph_plural # Still accepts plural marker in superscript
seconds = pynini.string_map([("second", "d"),("seconde", "de")])
seconds += graph_plural
graph_firsts = pynutil.insert("integer: \"1\" morphosyntactic_features: \"") + firsts
graph_seconds = pynutil.insert("integer: \"2\" morphosyntactic_features: \"") + seconds
# + [markdown] id="D2vQ4m7o7p84"
# Placing them in our class:
# + id="w_JKT8JMf-Mz"
class OrdinalFst(GraphFst):
def __init__(self, cardinal: GraphFst):
super().__init__(name="ordinal", kind="classify")
graph_cardinal = cardinal.graph_no_exception # may replace
strip_morpheme = pynutil.delete("ième") # deletes suffix
graph_root_change = pynini.string_map([("quatrième", "quatre"),
("cinquième", "cinq"),
("neuvième", "neuf"),
("onzième", "onze"),
("douzième", "douze"),
("treizième", "treize"),
("quatorzième", "quatorze"),
("quinzième", "quinze"),
("seizième", "seize"),
("trentième", "trente"),
("quarantième", "quarante"),
("cinquantième", "cinquante"),
("soixantième", "soixante"),
("millième", "mille"),
])
# Component will accept all tokens that end with desired strings
graph_get_cardinal = NEMO_SIGMA + (strip_morpheme | graph_root_change)
# Graph will map ordinals beyond second ordinal to their cardinals
graph_reg_ordinals = graph_get_cardinal @ graph_cardinal
# Graphing morphosyntactic_features
graph_morpheme = pynutil.insert("e") # Insert e superscript
graph_plural = pynini.accep("s").ques # ques is equivalent to pynini.closure(, 0, 1)
graph_morpheme_component = graph_morpheme + graph_plural
graph_morphosyntactic_features = (pynutil.insert(" morphosyntactic_features: \"")
+ graph_morpheme_component
)
# Adding in the `integer` property:
graph_ordinal = pynutil.insert("integer: \"") + graph_reg_ordinals + pynutil.insert("\"")
graph_ordinal += graph_morphosyntactic_features
# Case of first and second ordinals
firsts = pynini.string_map([("premier", "er"), ("première","re")])
firsts += graph_plural # Still accepts plural marker in superscript
seconds = pynini.string_map([("second", "d"),("seconde", "de")])
seconds += graph_plural
graph_firsts = pynutil.insert("integer: \"1\" morphosyntactic_features: \"") + firsts
graph_seconds = pynutil.insert("integer: \"2\" morphosyntactic_features: \"") + seconds
# All together
graph_ordinal = pynini.union(graph_ordinal, graph_firsts, graph_seconds)
self.fst = graph_ordinal.optimize()
# + [markdown] id="CpGHVg6chmA0"
# Trying out on some examples:
# + id="b5DL3PZRhpc8"
cardinal = CardinalFst()
ordinal = OrdinalFst(cardinal).fst
apply_fst("premier", ordinal)
apply_fst("premiers", ordinal)
apply_fst("seconde", ordinal)
apply_fst("douzièmes", ordinal)
apply_fst("cent-cinquièmes", ordinal)
# + [markdown] id="MNQVgiv-UK29"
# ### Special Tokens
# + [markdown] id="UdiNAHGh71O9"
# If you are particularly astute, you may have noticed that we have not closed the quotations around the `morphosyntactic_features` throughout, despite doing so for `integer`. This is not a typo, as there is one more aspect of the Classifier that must be addressed: special cases.
#
# For your language, you may notice that there are occasional exceptions to writing rules that are signaled by a specific vocabulary token in a string. As this must be communciated to our Verbalizer, it is important that we signal this vocabulary through our Classifier.
#
# For French, this can occur in the normalization of centuries. When using Ordinals to indicate centuries, French commonly writes with Roman numerals. For example:
# - "Fifth century" -> "cinquième siècle" -> `Vᵉ siècle`
# - "Twentieth century" -> "vintième siècle" -> `XXᵉ siècle`
#
# As such, we must allow our Classifier to pass on the information that "siècle" follows an ordinal to our Verbalizer, so it may normalize with Roman numerals. We accomplish this by appending a WFST that accepts special tokens that follow our Ordinals, adding them to our `morphosyntactic_features` attribute with a forward slach to delineate.
# + id="MsWnT4BfQKcC"
special_tokens = pynini.accep("siècle")
graph_special_tokens = delete_space + pynutil.insert("/") + special_tokens # We need to delete the space in between this token and the following one.
graph_special_tokens = pynini.closure(graph_special_tokens, 0, 1)
graph_ordinal += graph_special_tokens + pynutil.insert("\"")
# + [markdown] id="698_n5SFQ_jP"
# *Once again, it is advised to retain a tsv file in `data` to quickly append these key-words.*
#
# Having taken care of the special case, we may now call `add_tokens` and complete the graph (fully written out below).
# + id="nZ1dkft0Riou"
class OrdinalFst(GraphFst):
def __init__(self, cardinal: GraphFst):
super().__init__(name="ordinal", kind="classify")
graph_cardinal = cardinal.graph_no_exception # may replace
strip_morpheme = pynutil.delete("ième") # deletes suffix
graph_root_change = pynini.string_map([("quatrième", "quatre"),
("cinquième", "cinq"),
("neuvième", "neuf"),
("onzième", "onze"),
("douzième", "douze"),
("treizième", "treize"),
("quatorzième", "quatorze"),
("quinzième", "quinze"),
("seizième", "seize"),
("trentième", "trente"),
("quarantième", "quarante"),
("cinquantième", "cinquante"),
("soixantième", "soixante"),
("millième", "mille"),
])
# Component will accept all tokens that end with desired strings
graph_get_cardinal = NEMO_SIGMA + (strip_morpheme | graph_root_change)
# Graph will map ordinals beyond second ordinal to their cardinals
graph_reg_ordinals = graph_get_cardinal @ graph_cardinal
# Graphing morphosyntactic_features
graph_morpheme = pynutil.insert("e") # Insert e superscript
graph_plural = pynini.accep("s").ques # We create an acceptor since we must process the possible "s"
graph_morpheme_component = graph_morpheme + graph_plural
graph_morphosyntactic_features = (pynutil.insert(" morphosyntactic_features: \"")
+ graph_morpheme_component
)
# Adding in the `integer` property:
graph_ordinal = pynutil.insert("integer: \"") + graph_reg_ordinals + pynutil.insert("\"")
graph_ordinal += graph_morphosyntactic_features
# Case of first and second ordinals
firsts = pynini.string_map([("premier", "er"), ("première","re")])
firsts += graph_plural # Still accepts plural marker in superscript
seconds = pynini.string_map([("second", "d"),("seconde", "de")])
seconds += graph_plural
graph_firsts = pynutil.insert("integer: \"1\" morphosyntactic_features: \"") + firsts
graph_seconds = pynutil.insert("integer: \"2\" morphosyntactic_features: \"") + seconds
# Special tokens
special_tokens = pynini.accep("siècle")
graph_special_tokens = delete_space + pynutil.insert("/") + special_tokens # We need to delete the space in between this token and the following one.
graph_special_tokens = pynini.closure(graph_special_tokens, 0, 1)
graph_ordinal += graph_special_tokens + pynutil.insert("\"")
# Finishing
graph_ordinal = self.add_tokens(graph_ordinal)
self.fst = graph_ordinal.optimize()
# + [markdown] id="7a4zBo-YS1QD"
# ## Verbalizer
# + [markdown] id="zYbrcGyGS2rW"
# The initial part of the Ordinal Verbalizer is similar to the Cardinal WFST: we simply need to build a Verbalizer that inherits from `GraphFST` and removes the `integer` property tag.
# + id="KUv99A_rYjb9"
class OrdinalFst(GraphFst):
def __init__(self):
super().__init__(name="ordinal", kind="verbalize")
graph_integer = (
pynutil.delete("integer:")
+ delete_space
+ pynutil.delete("\"")
+ pynini.closure(NEMO_DIGIT, 1)
+ pynutil.delete("\"")
)
# + [markdown] id="zKCt_EapZXGW"
# Now we need to manage the `morphosyntactic_features` component. The first steps seem simple enough: delete the property tag and replace the superscript indicators with the actual superscripts.
# + id="yoa_mXMLabrU"
# Create mappings for all superscripts
superscript = pynini.union(
pynini.cross("e", "ᵉ"), # only delete first quote since there may be more features
pynini.cross("d", "ᵈ"),
pynini.cross("r", "ʳ"),
pynini.cross("s", "ˢ"),
)
# Append to deletion of feature property. Note that we use plus closure for multiple superscripts.
graph_morphosyntactic_features = pynutil.delete(" morphosyntactic_features: \"") + superscript.plus
# + [markdown] id="xOA7_MsUrSJS"
# ### Romanization
# + [markdown] id="K_SaG0DUa2t7"
# Now we come to the possible Romanization component. Since we need to graph the superscript components as following the number, we want to design our graph so that `morphosyntactic_features` is the last component of the graph. However, we do not know that we need Romanization until we see the `morphosyntactic_features` component. As such, we need to design our graph such that two options are available initially for an input, but only one allows full traversal.
# + [markdown] id="7dalc-tablG-"
# 
# + [markdown] id="mPTNCddNcEEE"
# In cases where your WFST decisions are dependent on latter parts of an input string, permitting the union of two separate paths when only one is valid usually assists, as a standard pathing heuristic will only choose the valid path.
#
# In the case of French, this would require us to separate our Verbalizer into two parts: one for Arabic numerals and one for Roman numerals. For the Arabic WFST, we simply conclude the graph.
# + id="0YSy1PYOcuyD"
graph_integer = (
pynutil.delete("integer:")
+ delete_space
+ pynutil.delete("\"")
+ pynini.closure(NEMO_DIGIT, 1)
+ pynutil.delete("\"")
)
graph_arabic = graph_integer + graph_morphosyntactic_features + pynutil.delete("\"")
# + [markdown] id="nnXjUU5Pf7Sh"
# For the Roman graph, things get a bit trickier. Ideally, we would want to build a WFST that maps each digit of `graph_arabic` to a Roman equivalent. However, consider the following examples:
# - 1 -> I
# - 10 -> X
# - 11 -> XI
# - 100 -> C
# - 101 -> CI
# - 110 -> CX
# - 111 -> CXI
#
# Since Roman numerals do not preserve powers of ten through digit placement, we will need to design separate FSTs for each digit position and apply them accordingly. As this can quickly become intensive, we will only work to enumerate the Ordinals from 1 to 100. (Note: We are doing this to accomodate centuries; there is little likelihood that any century beyond the 99th will be used in regular strings.)
# + [markdown] id="3-fQHMc2iQrz"
# First we design our graphs for converting from Arabic to Roman numerals:
# + id="d6PDySykiXTh"
digits = pynini.string_map([("1", "I"),
("2", "II"),
("3", "III"),
("4", "IV"),
("5", "V"),
("6", "VI"),
("7", "VII"),
("8", "VIII"),
("9", "IX"),
])
tens = pynini.string_map([("1", "X"),
("2", "XX"),
("3", "XXX"),
("4", "XL"),
("5", "L"),
("6", "LX"),
("7", "LXX"),
("8", "LXXX"),
("9", "XC"),
])
zero = pynutil.delete("0") # No Roman representation for zero.
# + [markdown] id="wb-LmwJdk59m"
# Now we build two separate filters: one will accept only single digit arabic numerals and the other will accept two digit arabic numerals. For this we can use `NEMO_DIGIT`:
# + id="DW3oD7Hbli2X"
map_one_digit = NEMO_DIGIT
map_two_digits = NEMO_DIGIT ** 2 # pynini overloads the exponent function to allow self-concatenation.
# + [markdown] id="xtYKLy9AmJZS"
# We now build mappings between two digit Arabic numerals and Roman numerals, composing them onto the filters:
# + id="dUy7uEUXmT_g"
graph_one_digit_romans = NEMO_DIGIT @ digits
graph_two_digit_romans = tens + (digits | zero)
graph_two_digit_romans = map_two_digits @ graph_two_digit_romans
graph_romans = graph_one_digit_romans | graph_two_digit_romans
# + [markdown] id="JEinyAMdm7RJ"
# We now take care of the occurence of "siècle" before composing onto `graph_integer`:
# + id="ERO19BbynPNX"
graph_romans = (graph_integer @ graph_romans) + graph_morphosyntactic_features
graph_romans += pynini.cross("/", " ") + "siècle" + pynutil.delete("\"")
# + [markdown] id="zN-fwrCGoToQ"
# We finalize with a union and calling `delete_tokens`, the complete Verbalizer now being::
# + id="kr2wcToAofWB"
class OrdinalFst(GraphFst):
def __init__(self):
super().__init__(name="ordinal", kind="verbalize")
# Maps integer and removes attribute
graph_integer = (
pynutil.delete("integer:")
+ delete_space
+ pynutil.delete("\"")
+ pynini.closure(NEMO_DIGIT, 1)
+ pynutil.delete("\"")
)
# Create mappings for all superscripts
superscript = pynini.union(
pynini.cross("e", "ᵉ"), # only delete first quote since there may be more features
pynini.cross("d", "ᵈ"),
pynini.cross("r", "ʳ"),
pynini.cross("s", "ˢ"),
)
# Append to deletion of feature property. Note that we use plus closure for multiple superscripts.
graph_morphosyntactic_features = pynutil.delete(" morphosyntactic_features: \"") + superscript.plus
# Writing WFST for arabic
graph_arabic = graph_integer + graph_morphosyntactic_features + pynutil.delete("\"")
# Mapping Roman numerals
digits = pynini.string_map([("1", "I"),
("2", "II"),
("3", "III"),
("4", "IV"),
("5", "V"),
("6", "VI"),
("7", "VII"),
("8", "VIII"),
("9", "IX"),
])
tens = pynini.string_map([("1", "X"),
("2", "XX"),
("3", "XXX"),
("4", "XL"),
("5", "L"),
("6", "LX"),
("7", "LXX"),
("8", "LXXX"),
("9", "XC"),
])
zero = pynutil.delete("0") # No Roman representation for zero.
# filters for Roman digits
map_one_digit = NEMO_DIGIT
map_two_digits = NEMO_DIGIT ** 2 # pynini overloads the exponent function to allow self-concatenation.
# Composing onto roman digits
graph_one_digit_romans = NEMO_DIGIT @ digits
graph_two_digit_romans = tens + (digits | zero)
graph_two_digit_romans = map_two_digits @ graph_two_digit_romans
graph_romans = graph_one_digit_romans | graph_two_digit_romans
# Writing WFST for Roman
graph_romans = (graph_integer @ graph_romans) + graph_morphosyntactic_features
graph_romans += pynini.cross("/", " ") + "siècle" + pynutil.delete("\"")
# Final composition
graph = (graph_romans | graph_arabic)
delete_tokens = self.delete_tokens(graph)
self.fst = delete_tokens.optimize()
# -
# Trying out our examples:
# +
example_regular = 'ordinal { integer: "12" morphosyntactic_features: "es" }'
example_roman = 'ordinal { integer: "12" morphosyntactic_features: "es/siècle" }'
fst = OrdinalFst().fst
apply_fst(example_regular, fst)
apply_fst(example_roman, fst)
# + [markdown] id="yBgLhTq9pWZe"
# We have now completed an Ordinal WFST from the ground up, allowing a separate numbering system for special cases.
# + [markdown] id="-W1-BMVJUXXk"
# ## Final notes
# + [markdown] id="kR7E64P4pPU_"
# Before moving on, there are some key takeaways that you may find useful for most (if not all) languages:
# - Many ordinal systems rely on alteration of Cardinals. Even in the example of Korean, it is using a pre-existing counting system and adding a suffix to indicate ordering. As such, your Ordinal WFST will likely follow this tutorial's structure of changing the Ordinal to its original root and then relying on your Cardinal WFST for the majority of processing.
# - The `morphosyntactic_features` property will carry the vast majority of information necessary for normalization through your Verbalizer.
# - While not all writing systems have the same quirk as using Roman numerals in reference to centuries, you will likely find cases in your language when a specific token indicates unique rules for a semiotic class. Carrying this information to the Verbalizer is usually the simplest means of preserving the token while also facilitating normalization.
# + [markdown] id="Rx8-LuJOUaa5"
# # Decimal WFST <a id="decimal-wfst"></a>
# + [markdown] id="D2MRXYxz8TGA"
#
# If the Cardinal WFST is the most crucial element of a normalization grammar, the construction of the Decimal WFST is a close second. Much like in the case of constructing Ordinals from Cardinal grammars, many aspects of the Decimal WFST will be reused throughout your other semiotic classes.
#
# To get started, you should study the numerical conventions in your language. In particular, you should take note of the following:
# - How is the decimal component of a number pronounced in your language of focus. (e.g. The English number `1.33` can be verbalized as "one point three three" or "one and thirty three hundredths.")
# - What is the punctuation mark used for decimal demarcation? (In North America, several writing systems use `.` while European nations will use `,`.)
# - Are there general rules regarding pronunciation/formatting of numbers past the decimal demarcation? (e.g. Does your language pronounce each digit or pronounce as a series of three digit numbers?)
#
# Such questions will likely require some deep familiarity with the language, and it may benefit to ask a native speaker for some input. Of course, the level of depth is dependent on your needs, but researching these questions will help your normalization system appear more organic.
# + [markdown] id="UsK78ib4N-gb"
# ## Grammar
# + [markdown] id="p4CLOOA9OAwZ"
# In the case of French, we have the following guidelines:
# - French uses the comma ( `,` ) for decimal delineation. It is articulated as "virgule".
# - Decimals can be read as a series of digits or grouped as Cardinal numbers arbitrarily. (e.g. "`.333` can be "virgule trois trois trois" or "virgule trois-cent-trente-trois".)
#
# As such, our grammar needs to accomodate the following pattern:
#
# `cardinal + "virgule" + string_of_cardinals`
#
# Given our experience with our previous WFSTs, this seems simple enough. We assume we have an instance of CardinalFST availble and create a subcomponent to map the integer portion of a decimal:
# + id="XSp9FTzhf0XZ"
cardinal = CardinalFst().graph_no_exception # NeMo equivalent of just_cardinals
# place cardinal under closure to permit values <=1
graph_integer = pynini.closure(cardinal, 0, 1)
# + [markdown] id="bk3_3iawgAZE"
# Compose it on a subcomponent that detects the delineator "virgule":
# + id="UMzfAKkngH6z"
delete_virgule = pynutil.delete("virgule")
graph_decimal = graph_integer + delete_space + delete_virgule
# + [markdown] id="GXjbtbLYgn17"
# And permit the occurence of several strings of cardinals to follow:
# + id="LMMNBJz8gtTA"
graph_string_of_cardinals = delete_space + graph_cardinal
graph_string_of_cardinals = pynini.closure(graph_string_of_cardinals, 1)
graph_decimal += graph_string_of_cardinals
# + [markdown] id="jTgnRLddhGdE"
# Let us try an example:
# + id="D4rjDh0ShJAp"
example = "trois virgule trois cinquante-cinq"
apply_fst(example, graph_decimal) # Should output only the cardinals in the string
# + [markdown] id="RfD1d9JOioyl"
# ### Ambiguity?
# + [markdown] id="3IaI1mCIe_6i"
# Note that our decision to include multiple strings of cardinals after the decimal marker has introduced some ambiguity into our WFST. Consider if a decimal number was followed by an integer series (e.g. `2.5, 5, 6`). Now what should be an application of one DecimalFST and two applications of a CardinalFST can be interpreted as a single DecimalFST application (e.g. `2.556`). What can be done?
#
# While we will address this in greater depth later (see [Tokenize and Classify](#tokenize-and-classify)), the short answer is that cases such as these must be calibrated according to use and linguistic intuition. As this is an inherent ambiguity in the language and its writing system, we can never truly remove this possibility without restricting our ability to model the language. However, we can rely on a few logical assumptions to guide our decision making:
# - Unless the grammar is deployed in a restrictive setting (e.g. a Financial or environment where strings of numbers are often read in series) it's not likely for a valid string to exhibit this level of ambiguity. Speakers typically try to reduce possible ambiguity in their language production and would likely rephrase to avoid issues such as these. [See Grice's maxims](https://en.wikipedia.org/wiki/Cooperative_principle).
# - While a language may allow a specific string by *rule*, speakers may typically avoid them *in practice* due to conventions or difficulty. In our case, while it may be possible to read `2,100 05` as "deux virgule dix-mille-cinq" ("two point ten-thousand and five"), it's dubious that a speaker would find such easier to read than "deux virgule une zéro zéro zéro cinq". (The place value of large strings tend to take longer to recognize.)
#
# While hardly satisfying, these two points will allow us to dismiss *some* worry. With the former observation being outside our grammar's ability to manage, we accomodate the latter point by using an alternate WFST from our CardinalFST: `numbers_up_to_million`. (To utilize in your own language, create a WFST in the Cardinal class right before building up to `graph_millions`. Again, calling `optimize` is advised.)
#
# + id="piNe1AWspa4J"
cardinal = CardinalFst().numbers_up_to_million
# place cardinal under closure to permit values <=1
graph_integer = pynini.closure(cardinal, 0, 1)
delete_virgule = pynutil.delete("virgule")
graph_decimal = graph_integer + delete_space + delete_virgule
graph_string_of_cardinals = delete_space + cardinal
graph_string_of_cardinals = pynini.closure(graph_string_of_cardinals, 1)
graph_decimal += graph_string_of_cardinals
# + [markdown] id="B1gglt0tfM5V"
# ## Classifier
# + [markdown] id="fVkOWkncgOZc"
# Like with our previous WFSTs, the main duty for the classifier is inserting the necessary properties for the semiotic token. For the `decimal` tag, the following properties are used:
# - `integer_part` - indicates value before decimal marker
# - `fractional_part` - indicates values after the decimal marker
# - `negative` - indicates if value is positive or negative (Optional)
# - `quantity` - designates if decimal is in regards to a specific quantity. (See Quantities.)
#
# We can begin by inserting the `integer_part` around our `cardinal` subcomponent and the `fractional_part` around our `graph_string_of_cardinals`
# + id="_zw_cDszh-fB"
graph_integer = pynutil.insert("integer_part: \"") + cardinal + pynutil.insert("\" ")
graph_fractional = pynutil.insert("fractional_part: \"") + graph_string_of_cardinals + pynutil.insert("\"")
# + [markdown] id="bxlnn_7tiQMn"
# We then concatenate them together with a component that recognizes and removes the decimal separator.
# + id="BxNS9_AwiWHf"
graph_integer_or_none = graph_integer | pynutil.insert("integer_part: \"0\" ", weight=.1) # In cases we don't always have an integer preceeding
graph_decimal_no_sign = graph_integer_or_none + delete_space + pynutil.delete("virgule") + graph_fractional
# + [markdown] id="b7uGfsi4i5UI"
# *Note that we allow insertion of 0 if there is no integer to accomodate reading of only decimal values*
#
# Now we allow the possibility of negative values. (Recall French uses "moins" to indicate the negative.)
# + id="VsP79naojQZR"
graph_negative = pynini.cross("moins", "negative: \"-\" ") + delete_space
graph_decimal = graph_negative + graph_decimal_no_sign
# + id="QTcvq5HqllqW"
example = "moins deux virgule cent-quatre"
apply_fst(example, graph_decimal)
# + [markdown] id="FVKuGj_9mZ75"
# Placing within a `DecimalFst` class, we have:
# + id="tXwr32ermesp"
class DecimalFst(GraphFst):
def __init__(self, cardinal: GraphFst):
super().__init__(name="decimal", kind="classify")
cardinal = cardinal.numbers_up_to_million
delete_virgule = pynutil.delete("virgule")
graph_integer = pynutil.insert("integer_part: \"") + cardinal + pynutil.insert("\" ") + delete_space
graph_integer_or_none = graph_integer | pynutil.insert("integer_part: \"0\" ", weight=.001) # In cases we don't always have an integer preceeding
graph_string_of_cardinals = delete_space + cardinal
graph_string_of_cardinals = pynini.closure(graph_string_of_cardinals, 1)
graph_fractional = pynutil.insert("fractional_part: \"") + graph_string_of_cardinals + pynutil.insert("\"")
graph_decimal_no_sign = graph_integer_or_none + pynutil.delete("virgule") + graph_fractional
graph_negative = pynini.cross("moins", "negative: \"-\" ") + delete_space
graph_negative = pynini.closure(graph_negative, 0, 1)
graph_decimal = graph_negative + graph_decimal_no_sign
graph = self.add_tokens(graph_decimal)
self.fst = graph.optimize()
# + [markdown] id="gjxI5mEKfHLo"
# ### Quantities
# + [markdown] id="3WuwWPf3py7G"
# Recalling our earlier remarks regarding convention in language use, you may find a need to adjust the DecimalFst when processing specific values. For instance, consider the following equivalencies from English:
# - `1,500,000` = "one million five hundred thousand" = "one point five million" = `1.5 million`
# - `2,750,000` = "two million seven hundred and fifty thousand" = "two point seven five million" = `2.75 million`
#
# For large numbers, there is a tendency to use the decimal system as though one is describing a quantity. Notably, there is a minimum value for which this is comfortable. (A speaker of English may say "three point five trillion" but "three point five hundred" comes off as odd.)
#
# This behavior can occur in other languages. For example, the amount of `$1,500,000` may be read in French as "une virgule cinq million de dollars" ("one point five million dollars").
# + [markdown] id="RgMBIKlYdsGz"
# Our Classifier can be made to accomodate this behavior: we simply need to repeat what we did for `OrdinalFst` and set aside several key terms to trigger our model. For French, we will choose all terms added for values greater than a million. (Chosen empirically.)
# + id="vEcsUXw5fUEe"
suffix = pynini.union(
"million",
"millions",
"milliard",
"milliards",
"billion",
"billions",
"billiard",
"billiards",
"trillion",
"trillions",
"trilliard",
"trilliards",
)
# + [markdown] id="wIIUAsR-fgQA"
# We will then need to use a WFST to graph any numbers the preceed these amounts. Note, unlike for our `DecimalFst`, we need to permit cardinals as well as decimals. This is because we want to be able to normalize a phrase like "three million" to `3 million` as this will be less obtrusive than `3,000,000`.
#
# As such, we will call a `CardinalFst` and a `DecimalFst` in for `graph_quantities`. Since these are both utilized for our `DecimalFst`, it would be more efficient to just pass them along as function/class variables.
# + id="yern-idtycWg"
def get_quantity(decimal, cardinal_up_to_thousand):
key_values = pynini.union(
"million",
"millions",
"milliard",
"milliards",
"billion",
"billions",
"billiard",
"billiards",
"trillion",
"trillions",
"trilliard",
"trilliards",
)
# The French WFST that this borrows from has not removed leading zeroes yet.
numbers = cardinal_up_to_thousand @ (
pynutil.delete(pynini.closure("0")) + pynini.difference(NEMO_DIGIT, "0") + pynini.closure(NEMO_DIGIT)
)
res = (
pynutil.insert("integer_part: \"")
+ numbers
+ pynutil.insert("\"")
+ (
pynini.union(delete_hyphen, delete_extra_space)
) # Can be written either as 'deux-millions' or 'deux millions' depending on whether it registers as a noun or part of cardinal.
+ pynutil.insert(" quantity: \"")
+ suffix
+ pynutil.insert("\"")
)
# Union with decimal to permit either a cardinal or decimal representation.
res |= decimal + delete_extra_space + pynutil.insert(" quantity: \"") + suffix + pynutil.insert("\"")
return res
# + [markdown] id="uT4LMo8ADBAq"
# We can now insert this into our Classifier, producing the following:
# + id="d2KrCuyGDLwh"
class DecimalFst(GraphFst):
def __init__(self, cardinal: GraphFst):
super().__init__(name="decimal", kind="classify")
quantities_cardinal = cardinal.graph_hundreds_component_at_least_one_none_zero_digit
cardinal = cardinal.graph_no_exception
delete_virgule = pynutil.delete("virgule")
graph_integer = pynutil.insert("integer_part: \"") + cardinal + pynutil.insert("\" ") + delete_space
graph_integer_or_none = graph_integer | pynutil.insert("integer_part: \"0\" ", weight=.001) # In cases we don't always have an integer preceeding
graph_string_of_cardinals = delete_space + cardinal
graph_string_of_cardinals = pynini.closure(graph_string_of_cardinals, 1)
graph_fractional = pynutil.insert("fractional_part: \"") + graph_string_of_cardinals + pynutil.insert("\"")
graph_decimal_no_sign = graph_integer_or_none + delete_virgule + graph_fractional
graph_negative = pynini.cross("moins", "negative: \"-\" ") + delete_space
graph_negative = pynini.closure(graph_negative, 0, 1)
graph_decimal = graph_negative + graph_decimal_no_sign
# Union default decimal with version that accepts quantities
graph_decimal |= graph_negative + get_quantity(
graph_decimal_no_sign, quantities_cardinal
)
final_graph = self.add_tokens(graph_decimal)
self.fst = final_graph.optimize()
# + id="cD-eKqO6qTyh"
cardinal = CardinalFst()
decimal = DecimalFst(cardinal).fst
example = "trois virgule cent-quatre billion"
apply_fst(example, decimal)
# + [markdown] id="HiSLKF3RfRZA"
# ## Verbalizer
# + [markdown] id="QnkOV5FlteQA"
# As before, the Verbalizer is responsible for removing the formatting and rendering a given token in conventional form. As the process remains similar to Ordinals and Cardinals (deleting strings in a regular matter) we will instead focus on a unique concern for `DecimalFst`: numeral spacing.
#
# For some writing systems, decimal numbers and other strings are typically not written as a single string, instead using punctuation to group numbers for clarity. For example, in the United States, integer digits greater than a thousand are separated by commas for every three digits:
# - `12345.678` -> `12,345.678`
#
# A similar rule occurs in French, save it employs spaces on each side of the decimal marker:
# - `12345,6789` -> `12 345,678 9`
# + [markdown] id="2h4WQZ1a4Cpc"
# While simple enough, this rule poses a slight complication: it works from the left and right of the decimal separator, whereas WFSTs process linearly from the beginning (or end) of strings. As such we will need to break the formatting rule into two components: one for the integer component and one for the decimal component.
# + [markdown] id="ViOFNdZw4-qu"
# Starting with the integer component, we need our subcomponent to recognize every three digits and insert a space before. We can achieve this with some `graph_utils` helper objects - `NEMO_DIGIT` and `NEMO_NON_BREAKING_SPACE`, which accept all digits and non-breaking spaces, respectively.
# + id="Z36be2Vo5VbR"
every_three_digits = NEMO_DIGIT ** 3 # accepts a string of three digits
space_every_three_integer = pynini.closure(NEMO_NON_BREAKING_SPACE + every_three_digits) # inserts space before every three digits.
# + [markdown] id="RSB2gGH-5vwi"
# However, we cannot let the component insert spaces when there are *only* three digits (e.g. `100`.) As such, we need to make sure the insertion only begins starting from the beginning of a string (e.g. when there is a string between one and three digits.)
# + id="wfWp3ghH6mDQ"
space_every_three_integer = pynini.closure(NEMO_DIGIT, 1, 3) + space_every_three_integer
# + [markdown] id="NJrQYSfA6vyu"
# For the case of the decimal spacing, we simply reverse the logic:
# + id="vBP6ncTp6yXX"
space_every_three_decimal = pynini.closure(NEMO_NON_BREAKING_SPACE + every_three_digits)
space_every_three_decimal = space_every_three_decimal + pynini.closure(NEMO_DIGIT, 1, 3)
# + [markdown] id="WRXPN_gk69VV"
# Placed into our Verbalizer, we would see the following:
# + id="h49eztvs7BXH"
class DecimalFst(GraphFst):
"""
Finite state transducer for verbalizing decimal, e.g.
decimal { negative: "true" integer_part: "12" fractional_part: "5006" quantity: "billion" } -> -12.5006 billion
"""
def __init__(self):
super().__init__(name="decimal", kind="verbalize")
# Need parser to group digits by threes
exactly_three_digits = NEMO_DIGIT ** 3
at_most_three_digits = pynini.closure(NEMO_DIGIT, 1, 3)
space_every_three_integer = (
at_most_three_digits + (pynutil.insert(NEMO_NON_BREAKING_SPACE) + exactly_three_digits).closure()
)
space_every_three_decimal = (
pynini.accep(",")
+ (exactly_three_digits + pynutil.insert(NEMO_NON_BREAKING_SPACE)).closure()
+ at_most_three_digits
)
group_by_threes = space_every_three_integer | space_every_three_decimal
self.group_by_threes = group_by_threes
optional_sign = pynini.closure(pynini.cross("negative: \"true\"", "-") + delete_space, 0, 1)
integer = (
pynutil.delete("integer_part:")
+ delete_space
+ pynutil.delete("\"")
+ pynini.closure(NEMO_NOT_QUOTE, 1)
+ pynutil.delete("\"")
)
integer = integer @ group_by_threes
optional_integer = pynini.closure(integer + delete_space, 0, 1)
fractional = (
pynutil.insert(",")
+ pynutil.delete("fractional_part:")
+ delete_space
+ pynutil.delete("\"")
+ pynini.closure(NEMO_NOT_QUOTE, 1)
+ pynutil.delete("\"")
)
fractional = fractional @ group_by_threes
optional_fractional = pynini.closure(fractional + delete_space, 0, 1)
quantity = (
pynutil.delete("quantity:")
+ delete_space
+ pynutil.delete("\"")
+ pynini.closure(NEMO_NOT_QUOTE, 1)
+ pynutil.delete("\"")
)
optional_quantity = pynini.closure(pynutil.insert(" ") + quantity + delete_space, 0, 1)
graph = (optional_integer + optional_fractional + optional_quantity).optimize()
self.numbers = graph # Saving just the part of the graph used for numbers
graph = optional_sign + graph
delete_tokens = self.delete_tokens(graph)
self.fst = delete_tokens.optimize()
# -
# Trying out some examples:
# +
fst = DecimalFst().fst
example1 = 'decimal { integer_part: "3" fractional_part: "10453" quantity: "billion" }'
example2 = 'decimal { integer_part: "22323" fractional_part: "104553" }'
apply_fst(example1, fst)
apply_fst(example2, fst)
# + [markdown] id="CZbshZCW8clI"
# # Money WFST <a id="money-wfst"></a>
# + [markdown] id="xuiv8HMz7yjm"
# Now that we've handled some of the foundational classes, it's time to see how they build up to permit more concrete ones. Let's see how the previous WFSTs assist in building a WFST for normalizing currency: the `MoneyFst`.
# + [markdown] id="wTU2c7MtUpqF"
# ## Grammar
# + [markdown] id="qqyRm8Ru8TDf"
# While the exact phrasing will vary, a valid string for currency will possess the following qualities:
# - A major and/or minor denomination of currency
# - A numeric quantity of the denomination
#
# As our `CardinalFst` and `OrdinalFst` already allow us to normalize the quantity, the only issue for `MoneyFst` is to graph the amounts and build a vocabulary to recognize the denominations.
#
# For French, we will use the following examples to build upon:
# - "une euros" -> `1 €`
# - "deux euros" -> `2 €`
# - "deux euros cinq" -> `2,5 €`
# - "cinq centimes" -> `0,5 €`
# - "deux billions de euros" -> `2 billions de euros`
# + [markdown] id="FMqUir9n9_cA"
# These suggest the following requirements of our grammar:
# - There must be a mapping between "euro" and "centime" and `€` in our vocabulary
# - This mapping must allow both singular and plural forms
# - The currency denomination is phrased between major and minor denominations ("une euro cinq" and not "une cinq euro")
# - Large quantities of currency are left 'as is' instead of normalized
#
# We may deal with the vocabulary in the typical fashion:
# + id="XN9nbNhB-vEV"
major_currency = pynini.string_map([("euro", "€")])
minor_currency = pynini.string_map([("centime", "€")])
graph_plural = pynutil.delete("s").ques
major_currency += graph_plural
minor_currency += graph_plural
# + [markdown] id="3aHrm1qPAc-f"
# Moving to the numbers, note that we need to append a leading zero to the value of fractional currency amounts ("five cents" -> `$0.05`). We bring back the subgraph from `CardinalFst` that maps tokens to numbers without tokenization to assist with this:
# + id="jwi-yQW1AjvG"
from nemo_text_processing.inverse_text_normalization.fr.taggers import cardinal
cardinal_graph = cardinal.CardinalFst()
graph_cardinal = cardinal_graph.graph_no_exception # graphs cardinals w/o tokenization
add_leading_zero_to_double_digit = (NEMO_DIGIT + NEMO_DIGIT) | (pynutil.insert("0") + NEMO_DIGIT)
graph_fractional_values = graph_cardinal @ add_leading_zero_to_double_digit
# -
# Now, let us consider how to manage arge quantities of currency. In our example ("deux billions de euros" -> `2 billions de euros`) we see that its behavior mirrors that of our `get_quantity` portion of `DecimalFst`. As such, it would be useful if there was a subcomponent of that graph that we could use in here. Like in the case of `CardinalFst`, let us go back and create a subgraph for later use. Since all our quantities are positive, this would be best accomplished right before incorporating the `negative` property, creating a `self.final_graph_wo_negative`:
class DecimalFst(GraphFst):
def __init__(self, cardinal: GraphFst):
super().__init__(name="decimal", kind="classify")
quantities_cardinal = cardinal.graph_hundreds_component_at_least_one_none_zero_digit
cardinal = cardinal.graph_no_exception
delete_virgule = pynutil.delete("virgule")
graph_integer = pynutil.insert("integer_part: \"") + cardinal + pynutil.insert("\" ") + delete_space
graph_integer_or_none = graph_integer | pynutil.insert("integer_part: \"0\" ", weight=.001) # In cases we don't always have an integer preceeding
graph_string_of_cardinals = delete_space + cardinal
graph_string_of_cardinals = pynini.closure(graph_string_of_cardinals, 1)
graph_fractional = pynutil.insert("fractional_part: \"") + graph_string_of_cardinals + pynutil.insert("\"")
graph_decimal_no_sign = graph_integer_or_none + delete_virgule + graph_fractional
### NEW GRAPH HERE
self.final_graph_wo_negative = graph_decimal_no_sign | get_quantity(
final_graph_wo_sign, cardinal.graph_hundreds_component_at_least_one_none_zero_digit
)
graph_negative = pynini.cross("moins", "negative: \"-\" ") + delete_space
graph_negative = pynini.closure(graph_negative, 0, 1)
graph_decimal = graph_negative + graph_decimal_no_sign
# Union default decimal with version that accepts quantities
graph_decimal |= graph_negative + get_quantity(
graph_decimal_no_sign, quantities_cardinal
)
final_graph = self.add_tokens(graph_decimal)
self.fst = final_graph.optimize()
# Allowing us to change our grammar to:
# +
from nemo_text_processing.inverse_text_normalization.fr.taggers import cardinal, decimal
cardinal_graph = cardinal.CardinalFst()
decimal_graph = decimal.DecimalFst(cardinal_graph)
graph_cardinal = cardinal_graph.graph_no_exception # graphs cardinals w/o tokenization
graph_decimal = decimal_graph.final_graph_wo_negative # graphs positive decimals w/o tokenization
add_leading_zero_to_double_digit = (NEMO_DIGIT + NEMO_DIGIT) | (pynutil.insert("0") + NEMO_DIGIT)
graph_fractional_values = graph_cardinal @ add_leading_zero_to_double_digit
# + [markdown] id="L1RHoW-TLzIz"
# Note that by doing this, we're also incorporating the formatting from the `decimal` class up to this point. Since these overlap with the `money` class (see next section), we have saved ourselves some work.
#
# Since we already made `graph_quantity` part of our `DecimalFst`, we can avoid dealing with large quantities now. However, this does mean we still need a way to leave currencies 'as is' without normalization. We can do this by using the `project` method, which will create a WFST that excepts either all valid inputs or all valid outputs of another WFST (depending on argument).
# + id="7l_TLtJkMluU"
major_currency_no_normalize = major_currency.project("input")
apply_fst("euro", major_currency_no_normalize)
# + [markdown] id="raBdHc_WXEpG"
# We then append this WFST with a WFST that recognizes prepositions commonly used before large values of currency ("d'", "des")
# + id="CEuxiVgDXRBf"
graph_preposition = pynini.union("des ", "d'") # Used for large amounts (billions de euros)
major_currency_no_normalize = pynini.closure(graph_preposition, 0, 1) + major_currency.project("input")
# + [markdown] id="FlXmf8Fq_Rm1"
# ## Classifier
# + [markdown] id="T5BBuQRzLuXS"
# For the Money semiotic class, we have available the following properties for tokenization:
# - `integer_part`
# - `fractional_part`
# - `currency`
#
# Laying the initial groundwork seems simple enough. We first instantiate our `MoneyFst` classifier with our initial grammars:
# + id="EZaCeHcFWVP3"
class MoneyFst(GraphFst):
def __init__(self, cardinal: GraphFst, decimal: GraphFst):
super().__init__(name="money", kind="classify")
major_currency = pynini.string_map([("euro", "€")])
minor_currency = pynini.string_map([("centime", "€")])
graph_plural = pynutil.delete("s").ques
major_currency += graph_plural
minor_currency += graph_plural
major_currency_no_normalize = major_currency.project("input")
graph_preposition = pynini.union("des ", "d'") # Used for large amounts (billions de euros)
major_currency_no_normalize = graph_preposition + major_currency.project("input")
graph_cardinal = cardinal.graph_no_exception
graph_decimal = decimal.final_graph_wo_negative
add_leading_zero_to_double_digit = (NEMO_DIGIT + NEMO_DIGIT) | (pynutil.insert("0") + NEMO_DIGIT)
graph_fractional_values = graph_cardinal @ add_leading_zero_to_double_digit
# + [markdown] id="_bpkXroLWaBo"
# Let us now manage the `currency` property. We have the following scenarios to consider:
# - Major denomination only
# - Minor denomination only
# - Major denomination and implicit minor denomination ("cinq euro trois")
# - Major denomination and explicit minor denomination ("cinq euros et trois centimes")
# - Large quantities of euros ("cinq billion des euros")
#
# Note how across cases the use of `graph_cardinal` and `graph_decimal` will be applied differently. Further, we may have varying orders in which tags are assigned proper values. For instance, if we have only minor denomination we would assign `fractional_part` before `currency`. Meanwhile, major denomination and implicit minor denomination would be the order of `integer_part`, `currency`, `fractional_part`. While we could try and figure out a way to preserve order, recall that the use of permutations in NeMo ITN makes that unnecessary: we can assume the desired order of tags reach our Verbalizer without make overt efforts in our Classifier!
#
# For example, let's say we need to process "five dollars" as `$5.00`. Processed linearly, we could get a token sequence along the lines of: `{ integer_part: "5" currency: "$" }`. If we passed this token array straight to a Verbalizer, we would need to configure a graph that effectively reverses the order so we could parse the `currency` field prior to the `integer_part` field, perhaps something along the lines of:
#
# `pynutil.insert("$") + delete_space + pynutil.delete('integer_part: \"') +.... + pynutil.delete('currency: "$"')`
#
# But since NeMo creates permutations of our Classifier outputs, this is unnecessary. We can simply assume whatever would be the most convenient order for us (e.g. `{ currency: "$" integer_part: "5" }`) and build our Verbalizer around that:
#
# `pynutil.delete('currency: \"') + NEMO_SIGMA + pynutil.delete('\" integer_part: \"') + NEMO_DIGIT +...`
#
# Along with helping to keep our script simpler (we can focus simply on tokenization and not worry about what input order our Verbalizers will accept), this also allows us to overcome structural constraints of WFSTs, namely that they are [limited in reordering text strings](https://en.wikipedia.org/wiki/Pushdown_automaton).
# + [markdown] id="fMZ13D2Dh9ZF"
# Keeping this in mind, let's begin mapping the proper tags. Since they're relatively simple, we can start with only major and minor denominations:
# + id="EtwWLp7VbbjM"
graph_integer_component = pynutil.insert("integer_part: \"") + graph_cardinal + pynutil.insert("\"")
graph_fractional_component = pynutil.insert("fractional_part: \"") + graph_fractional_values + pynutil.insert("\"")
graph_major_currency = pynutil.insert(" currency: \"") + major_currency + pynutil.insert("\"")
graph_minor_currency = pynutil.insert(" currency: \"") + minor_currency + pynutil.insert("\"")
graph_only_major_money = graph_integer_component + delete_space + graph_major_currency
graph_only_minor_money = graph_fractional_component + delete_space + graph_minor_currency
# + [markdown] id="XTmxrK4DmS39"
# Now we may append the case of an implicit `fractional_part` to `graph_only_major_money`
# + id="Zvzn3pQinkT0"
implicit_fractional_part = delete_space + pynutil.insert("fractional_part: \"") + graph_fractional_values + pynutil.insert("\"")
implicit_fractional_part = pynini.closure(implicit_fractional_part, 0, 1)
# + [markdown] id="tKFZkCVmn1OX"
# And the explicit fractional portion:
# + id="d_h0pTlMn3jz"
delete_et = pynutil.delete("et ") # Sometimes prefaces the minor currency
delete_et = pynini.closure(delete_et, 0 , 1)
delete_minor = pynutil.delete(minor_currency.project("input")) # to remove the minor currency
explicit_fractional_part = pynutil.insert("fractional_part: \"") + graph_fractional_values + pynutil.insert("\"")
explicit_fractional_part = delete_space + delete_et + explicit_fractional_part + delete_space + delete_minor
explicit_fractional_part = pynini.closure(explicit_fractional_part, 0, 1)
# + [markdown] id="rvnpAudgo-o3"
# We join them together:
# + id="qYzlIRWTpD8e"
graph_major_money = graph_only_major_money + (implicit_fractional_part | explicit_fractional_part)
graph_standard_money = graph_major_money | graph_only_minor_money
# + [markdown] id="TzeaKXVzpYs8"
# Finishing with the case the the large quantities of money, we need to use `graph_decimal` so we can exploit its ability to map quantities. Note that since we are using a pre-existing WFST, we can ignore inserting the tags ourselves, since this is already done by the Decimal WFST. As long as we remember to process this aspect with our Verbalizer, we can spare ourselves the extra step.
# + id="LnqX9mGFpmJm"
graph_large_money = pynutil.insert(" currency: \"") + major_currency_no_normalize + pynutil.insert("\"")
graph_large_money = graph_decimal + delete_space + graph_large_money
# + [markdown] id="24TUZnJKqgPA"
# Alltogether, this would give the following Classifier:
# + id="B7-muCO2qizg"
class MoneyFst(GraphFst):
def __init__(self, cardinal: GraphFst, decimal: GraphFst):
super().__init__(name="money", kind="classify")
major_currency = pynini.string_map([("euro", "€")])
minor_currency = pynini.string_map([("centime", "€")])
graph_plural = pynutil.delete("s").ques
major_currency += graph_plural
minor_currency += graph_plural
major_currency_no_normalize = major_currency.project("input")
graph_preposition = pynini.union("des ", "d'") # Used for large amounts (billions de euros)
major_currency_no_normalize = graph_preposition + major_currency.project("input")
graph_cardinal = cardinal.graph_no_exception
graph_decimal = decimal.final_graph_wo_negative
add_leading_zero_to_double_digit = (NEMO_DIGIT + NEMO_DIGIT) | (pynutil.insert("0") + NEMO_DIGIT)
graph_fractional_values = graph_cardinal @ add_leading_zero_to_double_digit
graph_integer_component = pynutil.insert("integer_part: \"") + graph_cardinal + pynutil.insert("\"")
graph_fractional_component = pynutil.insert("fractional_part: \"") + graph_fractional_values + pynutil.insert("\"")
graph_major_currency = pynutil.insert(" currency: \"") + major_currency + pynutil.insert("\"")
graph_minor_currency = pynutil.insert(" currency: \"") + minor_currency + pynutil.insert("\"")
graph_only_major_money = graph_integer_component + delete_space + graph_major_currency
graph_only_minor_money = graph_fractional_component + delete_space + graph_minor_currency
implicit_fractional_part = delete_space + pynutil.insert("fractional_part: \"") + graph_fractional_values + pynutil.insert("\"")
implicit_fractional_part = pynini.closure(implicit_fractional_part, 0, 1)
delete_et = pynutil.delete("et ") # Sometimes prefaces the minor currency
delete_et = pynini.closure(delete_et, 0 , 1)
delete_minor = pynutil.delete(minor_currency.project("input")) # to remove the minor currency
explicit_fractional_part = pynutil.insert("fractional_part: \"") + graph_fractional_values + pynutil.insert("\"")
explicit_fractional_part = delete_space + delete_et + explicit_fractional_part + delete_space + delete_minor
explicit_fractional_part = pynini.closure(explicit_fractional_part, 0, 1)
graph_major_money = graph_only_major_money + (implicit_fractional_part | explicit_fractional_part)
graph_large_money = pynutil.insert(" currency: \"") + major_currency_no_normalize + pynutil.insert("\"")
graph_large_money = graph_decimal + delete_space + graph_large_money
final_graph = graph_large_money | graph_major_money | graph_only_minor_money
final_graph = self.add_tokens(final_graph)
self.fst = final_graph.optimize()
# -
# Let's see the results:
# +
from nemo_text_processing.inverse_text_normalization.fr.taggers import decimal, cardinal
cardFst = cardinal.CardinalFst()
decFst = decimal.DecimalFst(cardFst)
moneyFst = MoneyFst(cardFst, decFst).fst
example = "douze virgule cinq billions d'euros"
apply_fst(example, moneyFst)
# + [markdown] id="gxdcyuLmAZZa"
# ## Verbalizer
# + [markdown] id="ZZFDWNwY6sOG"
# By this point, the creation of the Verbalizer should be rather straight-forward - delete the expected tokens and perform any specific formatting that was not caught by the Classifier.
#
# In fact, it is so straight-forward that much of the work does not even need to be explicitly managed by the Verbalizer. As mentioned previously, two of the properties we inserted in our Classifier where already referenced in our `DecimalFst` - `integer_part` and `fractional_part`. We even went so far to directly call a component of `DecimalFst` in our Classifier. As such, outside of the `currency` property - there is little in our Money token that is different from a standard Decimal token. Indeed, even the normalized forms are similar (`200,5` vs. `200,5 €`.)
# + [markdown] id="T7sgH0t79tmU"
# Given these similarities, it seems that we can save ourselves some work and simply use the Decimal Verbalizer to manage much of the normalization. Let's look at the basic format of our `MoneyFst` verbalizer, writing it so it accepts a `DecimalFst` as input:
# + id="BEu8nITP9mSG"
class MoneyFst(GraphFst):
def __init__(self, decimal: GraphFst):
super().__init__(name="money", kind="verbalize")
# + [markdown] id="JYVLou5N-Dk8"
# We manage the issue of deleting the `currency` property:
# + id="LO35tJ7G-H6N"
class MoneyFst(GraphFst):
def __init__(self, decimal: GraphFst):
super().__init__(name="money", kind="verbalize")
unit = (
pynutil.delete("currency:")
+ delete_extra_space
+ pynutil.delete("\"")
+ pynini.closure(NEMO_NOT_QUOTE, 1)
+ pynutil.delete("\"")
)
# + [markdown] id="bDS8XSII-Dpd"
# Now consider, we need to normalize an integer component, a fractional component, and a decimal to separate them. Since NeMo will automatically permutate all tags, we can assume whatever order we want. As such, we can assume we get the exact order that is accepted by our `DecimalFst`.
# + id="VtGfpjVA-r3u"
def __init__(self, decimal: GraphFst):
super().__init__(name="money", kind="verbalize")
unit = (
pynutil.delete("currency:")
+ delete_extra_space
+ pynutil.delete("\"")
+ pynini.closure(NEMO_NOT_QUOTE, 1)
+ pynutil.delete("\"")
)
graph = decimal.numbers + delete_space + unit
delete_tokens = self.delete_tokens(graph)
self.fst = delete_tokens.optimize()
# + [markdown] id="ZefxZLIU-uRU"
# It is as simple and compact as appending the `unit` component to the prexisting `decimal.numbers`.
#
# This feature is worth keeping in mind as you build up to more concrete classes: the combination of guaranteed tag permutations and prebuilt Verbalizers make the addition of semiotic classes progressively simpler despite the building complexity of your entire grammar.
# + [markdown] id="WydC7Cn28l5Y"
# # Time WFST <a id="time-wfst"></a>
# + [markdown] id="VelunbumCJJe"
# Our next composite graph will be for the Time WFST. Here, you may see more variation between your language and our example than with our previous classes. This is for a number of reasons, among them being that while there may be some standard cross linguistic patterns regarding time (e.g. `quantity_of_hours + quantity_of_minutes`), the use of various equivalent phrases can make an exhaustive grammar incredibly specific (e.g. consider managing "twelve fifteen", "twelve and a quarter", "quarter past twelve", "quarter after twelve", and "forty five until one" all together). You may find yourself drawing upon WFSTs that accomodate Cardinals, Fractions, and some basic subtraction.
#
# As such, we are going to focus on those aspects of the Time WFST that are necessary for a functional normalization of time related phrases, saving a more exhaustive grammar for your own specific languages and use cases.
# + [markdown] id="8wqb28wzATOR"
# ## Grammar
# + [markdown] id="AVntDM3AEz0v"
# For our Time WFST, we will focus on the following aspects:
# - Use of 24 or 12 hour base
# - Use of fraction terminology (e.g. "quarter" = `15`)
# - Accomodation of key-words ("noon", "midnight")
# - Counting backwards from the hour ("ten to five", "five to three")
# + [markdown] id="seU9hTbgFgu7"
# We'll start with the basic system.
#
# For French, time operates on a twenty-four hour system, with the zeroth hour being midnight. Time is given in the following format:
#
# `cardinal + heure(s) + (cardinal)`
#
# This is normalized as:
#
# `cardinal h (cardinal)`
#
# For instance, for `3:03`, we would have:
# - input: "trois heures trois"
# - output: `3 h 03`
#
# As such, our grammar needs to utilize a Cardinal WFST and have a means to accept "heures" from the input. Taking care of the latter case is simple enough:
# + id="HTSVxf4fI_ND"
graph_heures = pynini.accep("heure") + pynini.accep("s").ques
graph_heures = pynutil.delete(graph_heures)
# + [markdown] id="6LW7pXaXJSZa"
# For the cardinals, we could pass an instance of `CardinalFST` to our graph. But do we really need that level of coverage? We only really need to cover the numbers 0 - 60, which we could simply write a new WFST for. Further, it may be beneficial to allow our graph to separate possible ambiguity. While we will not cover it in our tutorial, you may in the future find it necessary to build a WFST for Measurements, of which quantities of time may play a part. Would it not be helpful for you WFST to know that "thirty hours" could only ever be a measurement instead of a possible time of day?
#
# Given the little amount of effort necessary and the quick benefit, we choose to make our hours and minutes explicit in the Time WFST.
# + id="R4aa06ZPLKIR"
hours = pynini.string_map([
("zéro","0"),
("une","1"),
("deux","2"),
("trois","3"),
("quatre","4"),
("cinq","5"),
("six","6"),
("sept","7"),
("huit","8"),
("neuf","9"),
("dix","10"),
("onze","11"),
("douze","12"),
("treize","13"),
("quatorze","14"),
("quinze","15"),
("seize","16"),
("dix-sept","17"),
("dix-huit","18"),
("dix-neuf","19"),
("vingt","20"),
("vingt-et-une","21"),
("vingt et une","21"),
("vingt-deux","22"),
("vingt-trois","23"),
("vingt-quatre","24"),
])
minutes = pynini.string_map([
("une", "01"),
("deux", "02"),
("trois", "03"),
("quatre", "04"),
("cinq", "05"),
("six", "06"),
("sept", "07"),
("huit", "08"),
("neuf", "09"),
("dix", "10"),
("onze", "11"),
("douze", "12"),
("treize", "13"),
("quatorze", "14"),
("quinze", "15"),
("seize", "16"),
("dix-sept", "17"),
("dix-huit", "18"),
("dix-neuf", "19"),
("vingt", "20"),
("vingt-et-une", "21"),
("vingt et une", "21"),
("vingt-deux", "22"),
("vingt-trois", "23"),
("vingt-quatre", "27"),
("vingt-cinq", "25"),
("vingt-six", "26"),
("vingt-sept", "27"),
("vingt-huit", "28"),
("vingt-neuf", "29"),
("trente", "30"),
("trente-et-une", "31"),
("trente et une", "31"),
("trente-deux", "32"),
("trente-trois", "33"),
("trente-quatre", "34"),
("trente-cinq", "35"),
("trente-six", "36"),
("trente-sept", "37"),
("trente-huit", "38"),
("trente-neuf", "39"),
("quarante", "40"),
("quarante-et-une", "41"),
("quarante et une", "41"),
("quarante-deux", "42"),
("quarante-trois", "43"),
("quarante-quatre", "44"),
("quarante-cinq", "45"),
("quarante-six", "46"),
("quarante-sept", "47"),
("quarante-huit", "48"),
("quarante-neuf", "49"),
("cinquante", "50"),
("cinquante-et-une", "51"),
("cinquante et une", "51"),
("cinquante-deux", "52"),
("cinquante-trois", "53"),
("cinquante-quatre", "54"),
("cinquante-cinq", "55"),
("cinquante-six", "56"),
("cinquante-sept", "57"),
("cinquante-huit", "58"),
("cinquante-neuf", "59"),
])
# + [markdown] id="4SmNsNKLM9cC"
# Now that we've managed the basic graph, we can address some of the more niche rules of French timekeeping.
#
# To start, French employs some colliquialisms that will be familiar to English speakers: minutes that are multiples of fifteen are referred to as fractions of a clock. In particular:
# - `5 h 15` -> "cinq heures **et quart**"
# - `5 h 30` -> "cinq heures **et demie**"
# - `5 h 45` -> "cinq eures **et trois quarts**"
#
# We thus need a means of rendering these as their numerical equivalents:
# + id="xHe3nfrpSlrE"
# Mapping 'et demi' and 'et qart'
graph_et = pynutil.delete("et") + delete_space
graph_demi = pynini.accep("demi")
graph_demi += pynini.accep("e").ques # people vary on feminine or masculine form
graph_demi = pynini.cross(graph_demi, "30")
graph_quart = pynini.accep('quart')
graph_quart = pynini.cross(graph_quart, '15')
graph_trois_quart = pynini.cross("trois quarts", "45")
graph_fractions = graph_demi | graph_quart | graph_trois_quart
graph_fractions = graph_et + graph_fractions
# + [markdown] id="HD2wobIQS3fX"
# Also like English, French will use key words to designate a specific timeslot. Noon and midnight are "midi" and "minuit" respectively.
# + id="ahbkiZFuTN2t"
# Midi and minuit
graph_midi = pynini.cross("midi", "12")
graph_minuit = pynini.cross("minuit", "0")
# + [markdown] id="6OyMoqfZTX1U"
# Now it's time to throw a wrench into things: counting backwards from the hour. How are we to get what is essentially a graph to do the subtraction necessarily for "ten to twelve" to become `11:50`?
#
# Easy: we build the subtraction into the graph itself. That is, we map the hours and minutes produced by our graph onto another graph that produces their amount shifted back a value.
#
# Let's take our "ten to twelve" example. Normally "ten" would map to `10` and "twelve" to `12`. But with these new graphs, the detection of the pattern `minute + to + hour` would signal that `10` should now become `50` and `12` become `11`.
# + [markdown] id="uMWifbm1VQjP"
# Let us do this for our French example. Luckily enough, the indication that a French string is regular: counting backwards from the hour is by use of the pattern `cardinal + heures + moins + minutes`
# + id="c4bV3T1pViCH"
hours_to = pynini.string_map([
("1","0"),
("2","1"),
("3","2"),
("4","3"),
("5","4"),
("6","5"),
("7","6"),
("8","7"),
("9","8"),
("10","9"),
("11","10"),
("12","11"),
("13","12"),
("14","13"),
("15","14"),
("16","15"),
("17","16"),
("18","17"),
("19","18"),
("20","19"),
("21","20"),
("22","21"),
("23","22"),
("24","23"),
("0","23"),
])
minutes_to = pynini.string_map([
("59", "01"),
("58", "02"),
("57", "03"),
("56", "04"),
("55", "05"),
("54", "06"),
("53", "07"),
("52", "08"),
("51", "09"),
("50", "10"),
("49", "11"),
("48", "12"),
("47", "13"),
("46", "14"),
("45", "15"),
("44", "16"),
("43", "17"),
("42", "18"),
("41", "19"),
("40", "20"),
("39", "21"),
("38", "22"),
("37", "23"),
("36", "24"),
("35", "25"),
("34", "26"),
("33", "27"),
("32", "28"),
("31", "29"),
("30", "30"),
("29", "31"),
("28", "32"),
("27", "33"),
("26", "34"),
("25", "35"),
("24", "36"),
("23", "37"),
("22", "38"),
("21", "39"),
("20", "40"),
("19", "41"),
("18", "42"),
("17", "43"),
("16", "44"),
("15", "45"),
("14", "46"),
("13", "47"),
("12", "48"),
("11", "49"),
("10", "50"),
("09", "51"),
("08", "52"),
("07", "53"),
("06", "54"),
("05", "55"),
("04", "56"),
("03", "57"),
("02", "58"),
("01", "59"),
])
graph_moins = pynutil.delete("moins")
# + [markdown] id="XOKETkIYZy5M"
# Why graph the digits instead of the tokens themselves? Along with avoiding some minor repetition and making editing more apparent, it allows this subgraph to be ported to other languages - if so desired.
#
# Further, it helps us illustrate a helpful idea within this tutorial: as long as a pattern is regular and/or finite, it is no major issue to accomodate it in our graph, regardless of mathematic or logic system it employs.
# + [markdown] id="DJbFiD2fAUc5"
# ## Classifier
# + [markdown] id="cK0SGXntaDkI"
# Once again we place the grammar within the proper child class of `GraphFst`. We also insert the proper tags for the `Time` class, which are:
# - `hours`
# - `minutes`
# - `suffix` (explained within this section)
# + id="9Eq5r-_VbBIg"
graph_hours_component = pynini.union(hours, graph_midi, graph_minuit)
graph_hours_component = pynutil.insert("hours: \"") + graph_hours_component + pynutil.insert("\"")
graph_minutes_component = (
pynutil.insert(" minutes: \"") + pynini.union(minutes, graph_fractions) + pynutil.insert("\"")
)
graph_minutes_component = delete_space + graph_minutes_component
graph_time_standard = (graph_hours_component + delete_space + graph_heures
+ pynini.closure(graph_minutes_component, 0, 1))
# + [markdown] id="2avfS3IacSiC"
# We now setup the alternate graph that allows backwards counting. Note, this is triggered by the occurence of "moins" between the hour and minute component.
# + id="TmpwisOVcn0T"
graph_hours_to_component = hours | graph_midi | graph_minuit
graph_hours_to_component @= hours_to
graph_hours_to_component = pynutil.insert("hours: \"") + graph_hours_to_component + pynutil.insert("\"")
graph_hours_to_component = graph_hours_to_component + delete_space + graph_heures
graph_minutes_to_component = (minutes | graph_demi | # No 'et' in fractions
(pynutil.delete("le ") + graph_quart) | graph_trois_quart)
graph_minutes_to_component @= minutes_to
graph_minutes_to_component = pynutil.insert(" minutes: \"") + graph_minutes_to_component + pynutil.insert("\"")
graph_time_to = graph_hours_to_component + delete_space + graph_moins + delete_space + graph_minutes_to_component
# + [markdown] id="FkO4tRRfdQT4"
# We now join it with our main component, allowing us to graph all times:
# + id="0O0vUVizdU8c"
graph_time = graph_time_standard | graph_time_to
# + [markdown] id="jbX4JV-LdY3Y"
# Once again we throw a wrench into things with the `suffix` feature. As in the case of Ordinals and Decimals, key-words can play into our Time WFST. For French, this occurs with the words "du matin", "de l'après-midi", and "du soir". (Respectively: "in the morning", "in the afternoon", and "in the evening".) Much like in English, these phrases alter how we write down the time. But instead of indicating `a.m.` or `p.m.`, these indicate *what hour system is used*. For example:
# - "deux heures du matin" -> `2 h` = `2:00 a.m.`
# - "deux heures de l'après-midi" -> `14 h` = `2:00 p.m.`
#
# Only a twelve hour system is used when these suffixes accompany the time. As such, our Classifier will need to either adjust the times like in the case of counting backwards or must pass the information to the Verbalizer so it can adjust.
#
# Since our Classifier is long enough as is, we will simply store this information in the `suffix` property and allow the Verbalizer to manage.
# + id="OqVa78zRgJw9"
graph_suffix_am = pynini.cross("du matin", "am")
graph_suffix_pm = pynini.string_map([("de l'après-midi", "pm"),("du soir", "pm")])
graph_suffix = pynini.cross(graph_suffix_am, "am") | pynini.cross(graph_suffix_pm, "pm")
graph_suffix_component = pynutil.insert(" suffix: \"") + graph_suffix + pynutil.insert("\"")
graph_suffix_component = delete_space + graph_suffix_component
graph_suffix_component = pynini.closure(graph_suffix_component, 0, 1)
# + [markdown] id="-LaJMIjUf1XR"
# And we append to our graph:
# + id="76myCFiggX3E"
class TimeFst(GraphFst):
def __init__(self):
super().__init__(name="time", kind="classify")
"""grammar omitted for length
....
....
....
"""
graph_hours_component = pynini.union(hours, graph_midi, graph_minuit)
graph_hours_component = pynutil.insert("hours: \"") + graph_hours_component + pynutil.insert("\"")
graph_minutes_component = (
pynutil.insert(" minutes: \"") + pynini.union(minutes, graph_fractions) + pynutil.insert("\"")
)
graph_minutes_component = delete_space + graph_minutes_component
graph_time_standard = (graph_hours_component + delete_space + graph_heures
+ pynini.closure(graph_minutes_component, 0, 1))
graph_hours_to_component = hours | graph_midi | graph_minuit
graph_hours_to_component @= hours_to
graph_hours_to_component = pynutil.insert("hours: \"") + graph_hours_to_component + pynutil.insert("\"")
graph_hours_to_component = graph_hours_to_component + delete_space + graph_heures
graph_minutes_to_component = (minutes | graph_demi | # No 'et' in fractions
(pynutil.delete("le ") + graph_quart) | graph_trois_quart)
graph_minutes_to_component @= minutes_to
graph_minutes_to_component = pynutil.insert(" minutes: \"") + graph_minutes_to_component + pynutil.insert("\"")
graph_time_to = graph_hours_to_component + delete_space + graph_moins + delete_space + graph_minutes_to_component
graph_time_no_suffix = graph_time_standard | graph_time_to
graph_suffix_am = pynini.cross("du matin", "am")
graph_suffix_pm = pynini.string_map([("de l'après-midi", "pm"),("du soir", "pm")])
graph_suffix = pynini.cross(graph_suffix_am, "am") | pynini.cross(graph_suffix_pm, "pm")
graph_suffix_component = pynutil.insert(" suffix: \"") + graph_suffix + pynutil.insert("\"")
graph_suffix_component = delete_space + graph_suffix_component
graph_suffix_component = pynini.closure(graph_suffix_component, 0, 1)
final_graph = graph_time_no_suffix + graph_suffix_component
final_graph = self.add_tokens(final_graph)
self.fst = final_graph.optimize()
# -
# Let's see how we did:
time = TimeFst().fst
example = "quatre heures moins cinq"
apply_fst(example, time)
# + [markdown] id="lPlJ1qyeAWOL"
# ## Verbalizer
# + [markdown] id="CrO-xtJ87PEl"
# The initial part of the Verbalizer should appear familiar. We delete the property tags `hours` and `minutes`, making sure they preserve the actual values for formatting.
# + id="fCzZKR7ek0Mz"
hour = (
pynutil.delete("hours:")
+ delete_space
+ pynutil.delete("\"")
+ pynini.closure(NEMO_DIGIT, 1, 2)
+ pynutil.delete("\"")
)
minute = (
pynutil.delete("minutes:")
+ delete_extra_space
+ pynutil.delete("\"")
+ pynini.closure(NEMO_DIGIT, 1, 2)
+ pynutil.delete("\"")
)
graph = hour + delete_extra_space + pynutil.insert("h") + minute.ques
# + [markdown] id="WnVV9GUKk-b7"
# We then deal with the case of `suffix`. We first note that if the suffix is for a morning time (before noon), then there is no further conversion that is needed. We may simply delete the property and its value.
# + id="haOEiSbglc6s"
day_suffixes = pynutil.delete("suffix: \"am\"")
graph = hours + delete_extra_space + pynutil.insert("h") + minute.ques + delete_space + day_suffixes.ques
# + [markdown] id="wL0FNg6Xlhb-"
# Meanwhile, the post-noon suffixes would require us shifting the hours value by twelve. Much like in the case of counting backwards from the hour, we can simply create a WFST to do this addition work for us.
# + id="YLrabUNplwG7"
hour_to_night = pynini.string_map([
("1", "13"),
("2", "14"),
("3", "15"),
("4", "16"),
("5", "17"),
("6", "18"),
("7", "19"),
("8", "20"),
("9", "21"),
("10", "22"),
("11", "23"), # Note that 12 and 24 would be phrased "midi" and "minuit" respectively
])
# + [markdown] id="X0-z-qJAmIiI"
# We then create an alternate graph where this conversion is mapped onto the hours function - given a post-noon suffix - and create a union with our earlier graph:
# + id="8CdEmo9NmN7u"
night_suffixes = pynutil.delete("suffix: \"pm\"")
graph |= (
hour @ hour_to_night
+ delete_extra_space
+ pynutil.insert("h")
+ minute.ques
+ delete_space
+ night_suffixes
)
# + [markdown] id="YnoIkZBqmaTo"
# Giving us a final Verbalizer of:
# + id="ZfXimvFBmdDD"
class TimeFst(GraphFst):
def __init__(self):
super().__init__(name="time", kind="verbalize")
hour_to_night = pynini.string_map([
("1", "13"),
("2", "14"),
("3", "15"),
("4", "16"),
("5", "17"),
("6", "18"),
("7", "19"),
("8", "20"),
("9", "21"),
("10", "22"),
("11", "23"),
])
day_suffixes = pynutil.delete("suffix: \"am\"")
night_suffixes = pynutil.delete("suffix: \"pm\"")
hour = (
pynutil.delete("hours:")
+ delete_space
+ pynutil.delete("\"")
+ pynini.closure(NEMO_DIGIT, 1, 2)
+ pynutil.delete("\"")
)
minute = (
pynutil.delete("minutes:")
+ delete_extra_space
+ pynutil.delete("\"")
+ pynini.closure(NEMO_DIGIT, 1, 2)
+ pynutil.delete("\"")
)
graph = hour + delete_extra_space + pynutil.insert("h") + minute.ques + delete_space + day_suffixes.ques
graph |= (
hour @ hour_to_night
+ delete_extra_space
+ pynutil.insert("h")
+ minute.ques
+ delete_space
+ night_suffixes
)
delete_tokens = self.delete_tokens(graph)
self.fst = delete_tokens.optimize()
# + [markdown] id="e5tPcCaSYuhY"
# If you've noticed, the Verbalizer process has become simpler as we've progressed through our WFSTs. Commonly, you will seldom need to even provide the amount of overhead we've seen in `TimeFst`, `MoneyFst`, and `OrdinalFst`, and the majority of this component is simply removing tokens as an intermediary step, as we'll see for our Name class.
# + [markdown] id="iHmRe3UIhyIH"
# # WhiteList WFST <a id="whitelist-wfst"></a>
# + [markdown] id="8kMn2qB9bVFy"
#
# While developing your grammars, you may encounter tokens that refuse standard categorization and yet still require normalization. For example, you may need to render "<NAME>" as `<NAME>` or "<NAME>" as `<NAME>`. As these cases are rather specific, they lack a regular pattern for a specific classifier. (What about "mister" as a token requires tokenization as opposed to "Brown".) Instead, we need to explicitly list their input-output mappings (i.e. a whitelist).
#
# For NeMo, this is performed through the `WhiteListFst`:
#
#
# + [markdown] id="6B4oPXYcccWs"
# ## Grammar
# + [markdown] id="RThTLUCRceOO"
# `WhitelistFst` is essentially just a wrapper for a `string_map` or `string_file` mapping with the appropriate formatting for deployment. Per our example, we can make a graph with the following:
# + id="eIOOb_wJdMMx"
graph = pynini.string_map([
("mister", "mr."),
("h m s", "h.m.s"),
("doctor", "dr.")
])
# + [markdown] id="O5kTXwmPZ9Tt"
# As previously mentioned, here is where the use of `string_file` will make maintnance much easier. Discovering whitelist mappings is an iterative process and you will more than likely need to return to your list throughout development. For instance, it may be obvious that tokens such as "madame", "miss", "esquire", but would you think of providing abreviations for "the right honorable" or "tennessee valley authority"? Keeping a tsv file available for quick insertions greatly assists here.
# + [markdown] id="RC5Cf-Z8dYVk"
# ## Classifier
# + [markdown] id="144nvAHEdfBJ"
# Unlike for our other WFSTs, There is no specific semiotic class for `WhiteListFst`. It instead falls under the default Name class to designate there is no need for further processing beyond obligatory tokenization. Indeed, we can simply insert the token ourselves instead of calling `add_tokens`.
# + id="oPkrmg2gdznd"
class WhiteListFst(GraphFst):
def __init__(self):
super().__init__(name="whitelist", kind="classify")
whitelist = pynini.string_map([
("mister", "mr."),
("h m s", "h.m.s"),
("doctor", "dr.")])
graph = pynutil.insert("name: \"") + convert_space(whitelist) + pynutil.insert("\"")
self.fst = graph.optimize()
# + [markdown] id="B05kdSIdd2dv"
# ## Verbalizer
# -
# Since the whitelisted token has already been rendered in the desired normalized form, all that is necessary is to strip the `name` token and render the string 'as is'. This can be done by through the following:
# + id="gaq3voIYiUCA"
class WhiteListFst(GraphFst):
def __init__(self):
super().__init__(name="whitelist", kind="verbalize")
graph = (
pynutil.delete("name:")
+ delete_space
+ pynutil.delete("\"")
+ pynini.closure(NEMO_CHAR - " ", 1)
+ pynutil.delete("\"")
)
graph = graph @ pynini.cdrewrite(pynini.cross(u"\u00A0", " "), "", "", NEMO_SIGMA) # Removes possible null token
self.fst = graph.optimize()
# + [markdown] id="cUE7Gg35bWKb"
# While the graph is largely self-explanatory, take note that the default implementation assumes a character string without spacing. If you intend to include additional formatting in your normalization (e.g. `H. M. S.` instead of `H.M.S.`), you may need to adjust the graph to expand coverage.
# + [markdown] id="_o_a15Fg7niv"
# # Word and Punctuation WFST <a id="word-and-punctuation-wfst"></a>
# + [markdown] id="Zi6lP7mTmnUV"
# Continuing with the Name class, we will conclude with the Word and Punctuation WFSTs. These are among the simplest and most crucial classes of the entire ITN system, as they classify all tokens that are not caught by other semiotic classes. Since these other tokens make up the majority of all strings your normalization system will encounter, they are essential for general functionality.
#
# However, they escape discussion as their function is self-evident: since they function as default classes, tokens only reach Word WFST and Punctuation WFST if they have not been accepted by the other WFSTs. As such, we can simply accept the tokens as they are, providing them a `name` tag.
# + [markdown] id="9zCqczLqp5NW"
# ## Classifier
# + [markdown] id="eUWum5U0p99c"
# For instance, consider the entire `WordFst` Classifier in its entirety:
# + id="CCZSTeDHofDl"
class WordFst(GraphFst):
def __init__(self):
super().__init__(name="word", kind="classify")
word = pynutil.insert("name: \"") + pynini.closure(NEMO_NOT_SPACE, 1) + pynutil.insert("\"")
self.fst = word.optimize()
# + [markdown] id="9ys2VpjjoiEC"
# It just processes the entire token string with the `NEMO_NOT_SPACE` utility WFST (which accepts any string that is not a space). For your language, you may simply use one of the prexisting `WordFst`.
#
# Depending on language, the `PunctuationFst` may require some (minimal) adjustment. Note the following:
# + id="Mnnd3PVMpF4t"
class PunctuationFst(GraphFst):
def __init__(self):
super().__init__(name="punctuation", kind="classify")
s = "!#$%&\'()*+,-./:;<=>?@^_`{|}~"
punct = pynini.union(*s)
graph = pynutil.insert("name: \"") + punct + pynutil.insert("\"")
self.fst = graph.optimize()
# + [markdown] id="_afW02LXpLtz"
# If your language uses other punctuation than that in the `s` string (or reserves some of the punctuation as characters), you may simply edit `s` to accomodate.
#
# For instance, French has a unique quotation style that utilizes guillemets "« »". We may add their Unicode codepoints (to avoid encoding issues) to `s`:
# + id="mgfZIKzVplVm"
class PunctuationFst(GraphFst):
def __init__(self):
super().__init__(name="punctuation", kind="classify")
s = "!#$%&\'()*+,-./:;<=>?@^_`{|}~"
guillemets = "\u00AB" + "\u00BB" # quotation marks in French.
s += guillemets
punct = pynini.union(*s)
graph = pynutil.insert("name: \"") + punct + pynutil.insert("\"")
self.fst = graph.optimize()
# + [markdown] id="6Upb5-wcp_7H"
# ## Verbalizer
# + [markdown] id="ufWT1T6GqCCT"
# Note that both `PunctuationFst` and `WordFst` both encode with the `name` property. This leaves no differentiation between the two for a Verbalizer. This makes sense as there are no particular formatting rules for them, they simply need a placeholder tag to avoid alteration between the Classifier and Verbalizer step. Once passed to the verbalizer, they are rendered as normal by simply removing the tag (this is practically identical to the WhiteListFST):
# + id="LqyhqQKZqcph"
class WordFst(GraphFst):
def __init__(self):
super().__init__(name="word", kind="verbalize")
chars = pynini.closure(NEMO_CHAR - " ", 1)
char = pynutil.delete("name:") + delete_space + pynutil.delete("\"") + chars + pynutil.delete("\"")
graph = char @ pynini.cdrewrite(pynini.cross(u"\u00A0", " "), "", "", NEMO_SIGMA) # Cleans up possible null character
self.fst = graph.optimize()
# + [markdown] id="lGbrUkcpapyi"
# For many languages, the writing of your `WordFst` and `PunctuationFst` (both Classifiers and Verbalizers) will require no more than duplicating the prexisting grammars found in NeMo Text Processing.
# + [markdown] id="5y9jhkhQ7p4W"
# # Other Classes <a id="other-classes"></a>
# + [markdown] id="j1mgnISmiu-g"
# While the preceeding discussion should be suitable for development of the remaining classes, some helpful notes may be of use before continuing:
# - Fraction WFST: This is the last of the 'fundamental' classes and should take priority after completion of the Decimal WFST. It operates very similarly to the Ordinal WFST in that you wish to recover the Cardinal roots for the numerator and denominator prior to tagging. Its properties are: `negative`, `integer_part`, `numerator`, and `denominator`.
# - Measure WFST: Like the Money WFST, this will require management of several 'parent' WFSTS (Fraction, Cardinal, Decimal) to be suitably comprehensive. As well, you may find it more productive to find ways to compose new measurement units instead of simply listing all (e.g. micrometers, petameters, miles per hour, feet per second). Its properties are: `negative`, `units` and it allows subgraphs of the `cardinal`, `decimal`, and `fraction` classes. (This is, it allows tokenization within the tokenization.)
# - Date WFST: Depending on writing conventions, this may vary in complexity. For instance, English speakers may write dates as `01/01/2021/` or `Jan. 1 2021`. Are there specific use cases where one is preferred or should you simply decide on a format? Further, you may wish to take advantage of the `preserve order` property to avoid possible unwanted verbalizations (some implementations will permit both `Jan. 1` and `1 Jan.` if not careful.) Its properties are: `month`, `day`, and `year`.
# - Telephone WFST: These will be heavily dependent not only on writing conventions but even regional preference. For instance, the U.S. commonly uses a ten digit system broken into the following sequence: `###-###-####`. Meanwhile, mainland France breaks a ten digit sequence into groups of two: `##-##-##-##-##`. Take careful note of how your language's target region verbalizes these figures and leave room for some variation in development. The `telephone` class has only one property: `number_part`.
# - Electronic WFST: For normalizing email addresses or urls, you will need to develop for the `electronic` class. The main concerns will be managing alphanumeric strings and parsing the reserved symbols used for protocols and domains. (How does your target language pronounce "https://"? www? '.' or '@'?") Depending on whether you are normalizing a url or email, the following properties will be needed:
# - email: `username`, `domain`
# - url: `protocol` (Sparrowhawk allows further detail here but NeMo passes the entire url through the `protocol` property)
# + [markdown] id="-i25X8mK90n3"
# # Tokenize and Classify <a id="tokenize-and-classify"></a>
# + [markdown] id="v4bcigU6b9ss"
# We are now ready to build a general Classifier for our entire language. Upon completion of your grammars, the next step is to unite them together in a general Classifier WFST - located within a `tokenize_and_classify.py` file, preferably. This WFST will be responsible for determining the appropriate semiotic class for each token in your string and processing the necessary properties for normalization.
#
# For this section, we will focus on the following: grammar composition, assignment of weights, and importing/exporting as a FAR file. Since we will need to work with some instantiated graphs, let's preload them before proceeding. (Note the compilement time.)
# +
from nemo_text_processing.inverse_text_normalization.fr.taggers.cardinal import CardinalFst
from nemo_text_processing.inverse_text_normalization.fr.taggers.decimal import DecimalFst
from nemo_text_processing.inverse_text_normalization.fr.taggers.money import MoneyFst
from nemo_text_processing.inverse_text_normalization.fr.taggers.ordinal import OrdinalFst
from nemo_text_processing.inverse_text_normalization.fr.taggers.punctuation import PunctuationFst
from nemo_text_processing.inverse_text_normalization.fr.taggers.time import TimeFst
from nemo_text_processing.inverse_text_normalization.fr.taggers.whitelist import WhiteListFst
from nemo_text_processing.inverse_text_normalization.fr.taggers.word import WordFst
cardinal = CardinalFst()
cardinal_graph = cardinal.fst
ordinal = OrdinalFst(cardinal)
ordinal_graph = ordinal.fst
decimal = DecimalFst(cardinal)
decimal_graph = decimal.fst
whitelist_graph = WhiteListFst().fst
word_graph = WordFst().fst
time_graph = TimeFst().fst
money_graph = MoneyFst(cardinal, decimal).fst
punct_graph = PunctuationFst().fst
# + [markdown] id="MIv58eSocOV1"
# ## Grammar
# + [markdown] id="k_RPlnfVdG5E"
# As for all previous grammars, the `tokenize_and_classify` grammar inherits from a `GraphFst` as an individual class: `ClassifyFst`.
# + id="WHKG4c2WdW0G"
class ClassifyFst(GraphFst):
def __init__(self):
super().__init__(name="tokenize_and_classify", kind="classify")
# + [markdown] id="j9_I6DJmdcOG"
# This class is responsible for instantiating all subgraphs and passing necessary dependencies:
# + id="4YtmcxLOdlas"
class ClassifyFst(GraphFst):
def __init__(self):
super().__init__(name="tokenize_and_classify", kind="classify")
cardinal = CardinalFst()
cardinal_graph = cardinal.fst
ordinal = OrdinalFst(cardinal)
ordinal_graph = ordinal.fst
decimal = DecimalFst(cardinal)
decimal_graph = decimal.fst
whitelist_graph = WhiteList().fst
word_graph = WordFst().fst
time_graph = TimeFst().fst
money_graph = MoneyFst(cardinal, decimal).fst
punct_graph = PunctuationFst().fst
# + [markdown] id="y5vGvv3HeAY9"
# We then join all the grammars together so `ClassifyFst` can apply them. Rather uncermoniously, this is accomplished by performing a union across all grammars (excluding `PunctuationFst`, to assist tokenization). We then follow this union by inserting the `tokens` class around the resulting formatting (required for processing):
# + id="oocgPQ5geZJO"
class ClassifyFst(GraphFst):
def __init__(self):
super().__init__(name="tokenize_and_classify", kind="classify")
cardinal = CardinalFst()
cardinal_graph = cardinal.fst
ordinal = OrdinalFst(cardinal)
ordinal_graph = ordinal.fst
decimal = DecimalFst(cardinal)
decimal_graph = decimal.fst
whitelist_graph = WhiteListFst().fst
word_graph = WordFst().fst
time_graph = TimeFst().fst
money_graph = MoneyFst(cardinal, decimal).fst
punct_graph = PunctuationFst().fst
classify = (
time_graph
| whitelist_graph
| decimal_graph
| cardinal_graph
| ordinal_graph
| money_graph
| word_graph
)
token = pynutil.insert("tokens { ") + classify + pynutil.insert(" }")
# + [markdown] id="ASWDXWQjfLEU"
# Our graph is now able to process an individual token. But what about a string? Here you will need to be mindful of the tokenization behavior for your language and decide on your desired treatment of punctuation (hence exclusion from the main graph).
#
# For our purposes, we will assume the convention of space and punctuation serving as token seperators. We graph punctuation as individual tokens
# + id="r6WztK2jwhFt"
punct_graph = PunctuationFst().fst
punct = pynutil.insert("tokens { ") + punct_graph + pynutil.insert(" }")
# + [markdown] id="9T2rT89jw3T1"
# and join the `punct` graph with our `tokens` graph (inserting spaces between tokens for formatting)
# + id="rGtVOK-txKOP"
token = "PLACEHOLDER"
token_plus_punct = (
pynini.closure(punct + pynutil.insert(" ")) + token + pynini.closure(pynutil.insert(" ") + punct)
) # Note the use of closure incase there are multiple punctuations
graph = token_plus_punct + pynini.closure(delete_extra_space + token_plus_punct)
# + [markdown] id="_gixfQ69xWPe"
# then address space between tokens:
#
# `graph = delete_space + graph + delete_space`
# + [markdown] id="DWnmazWecyUG"
# ## Weighting <a id="classifyweights"></a>
# + [markdown] id="egHbwIbMx-hT"
# Were we to leave our `ClassifyFst` like this, we would undoubtedly encounter a mountain of erros. What will stop our graph from treating punctuation that is part of a previous grammar as a token separator (e.g. "vingt-et-un")? How do we ensure that a currency string isn't treated as solely a decimal string with a `name` token following?
#
# As in previous cases, the solution lies in our choice of weights for the grammar.
# + [markdown] id="y3U7_M8CyxZ1"
# Let us return to the main graph:
# + id="9VXe1dfsy3Be"
classify = (
time_graph
| whitelist_graph
| decimal_graph
| cardinal_graph
| ordinal_graph
| money_graph
| word_graph
)
punct = pynutil.insert("tokens { ") + punct_graph + pynutil.insert(" }")
# + [markdown] id="aY4vOFqxy5ua"
# Beyond the path weights that we explicitly added, these graphs are currently weightless. Since we want the graphs themselves to be the general determiners of a path, let us use some default weights an order of magnitude beyond our path weights (we use `pynutil.add_weight`):
# + id="bthyt_Le2rsA"
classify = (
pynutil.add_weight(time_graph, 1)
| pynutil.add_weight(whitelist_graph, 1)
| pynutil.add_weight(decimal_graph, 1)
| pynutil.add_weight(cardinal_graph, 1)
| pynutil.add_weight(ordinal_graph, 1)
| pynutil.add_weight(money_graph, 1)
| pynutil.add_weight(word_graph, 1)
)
punct = pynutil.insert("tokens { ") + pynutil.add_weight(punct_graph, 1) + pynutil.insert(" }")
# + [markdown] id="xMNIJbzj3MMP"
# Let's see what logical adjustements should be made. First off, we know that we want each class token to span the largest string possible. (e.g. We don't want "quatre-vingt" to be rendered as two `cardinal` classes with a hyphen in between.) As such, we want to penalize our graph for using more than one tokens. We can do so by establishing the following constraint: the sum of two or more tokens cannot be less than the weight of a single token. Or, for any pair of tokens `w_1` and `w_2`, their sum must always be greater than any other individual token (including themselves):
#
# `w_1 + w_2 > k >= w`
#
# To keep things simple, let us make the upper limit `2`. This means we should increase all the weights to keep our constraint:
#
# -
classify = (
pynutil.add_weight(time_graph, 1.1)
| pynutil.add_weight(whitelist_graph, 1.1)
| pynutil.add_weight(decimal_graph, 1.1)
| pynutil.add_weight(cardinal_graph, 1.1)
| pynutil.add_weight(ordinal_graph, 1.1)
| pynutil.add_weight(money_graph, 1.1)
| pynutil.add_weight(word_graph, 1.1)
)
punct = pynutil.insert("tokens { ") + pynutil.add_weight(punct_graph, 1.1) + pynutil.insert(" }")
# Do we want this constraint to include all tokens? Imagine if we had a string of multiple semiotic tokens in a row. Since this string's combined weight would be larger than any single class token, a grammar that served as a universal acceptor (i.e. `word_graph`) would be preferred over these individual classes. This would be obviously incorrect. As such, we want to make sure that `word_graph` would only be traversed when there is truly no other option:
# + id="qc_CU2ro63eg"
classify = (
pynutil.add_weight(time_graph, 1.1)
| pynutil.add_weight(whitelist_graph, 1.1)
| pynutil.add_weight(decimal_graph, 1.1)
| pynutil.add_weight(cardinal_graph, 1.1)
| pynutil.add_weight(ordinal_graph, 1.1)
| pynutil.add_weight(money_graph, 1.1)
| pynutil.add_weight(word_graph, 100)
)
punct = pynutil.insert("tokens { ") + pynutil.add_weight(punct_graph, 1.1) + pynutil.insert(" }")
# -
# Now, even with a string of fifty different class tokens, `word_graph` would still not be considered as a path to traverse.
# + [markdown] id="fW8C3vD-7Dbl"
# Next, let us consider our foundational graph: `cardinal_graph`. As Cardinals occur in practically all our WFSTs, it's possible for `cardinal_graph` to apply in almost all cases. Yet, we've specifically invoked `CardinalFST` when it was required in any of the other classes, so it will never be needed in any of those cases. This means that we want all those graphs to have *priority* over `cardinal_graph`. As such, we will increase its weight so it takes second lowest precedence (while stillpaying attention to the combined weight constraint).
# + id="97UwGaEn8pj7"
classify = (
pynutil.add_weight(time_graph, 1.1)
| pynutil.add_weight(whitelist_graph, 1.1)
| pynutil.add_weight(decimal_graph, 1.1)
| pynutil.add_weight(cardinal_graph, 1.2)
| pynutil.add_weight(ordinal_graph, 1.1)
| pynutil.add_weight(money_graph, 1.1)
| pynutil.add_weight(word_graph, 100)
)
punct = pynutil.insert("tokens { ") + pynutil.add_weight(punct_graph, 1.1) + pynutil.insert(" }")
# + [markdown] id="0d9Lw4Ot88_B"
# This form of thinking can be applied to all the 'foundational' graphs you may develop: the dependent graphs should take higher precedence than the graphs they borrow from. For instance, since `money_graph` utilizes `decimal_graph`, we know it should take precedence. However, since `decimal_graph` borrows from `cardinal_graph`, its weight must still be less than `1.2`. As such:
# + id="-wF8cgLK9tpU"
classify = (
pynutil.add_weight(time_graph, 1)
| pynutil.add_weight(whitelist_graph, 1)
| pynutil.add_weight(decimal_graph, 1.1)
| pynutil.add_weight(cardinal_graph, 1.2)
| pynutil.add_weight(ordinal_graph, 1)
| pynutil.add_weight(money_graph, 1.09)
| pynutil.add_weight(word_graph, 100)
)
punct = pynutil.insert("tokens { ") + pynutil.add_weight(punct_graph, 1) + pynutil.insert(" }")
# + [markdown] id="huMzDoZ2-FD2"
# For those classes that don't seem affected, we can set their weights as the same as those below their 'foundation' graphs, simply to prevent prioritization when not required
#
# Meanwhile, `whitelist_graph` should take precedence over all others, as it may contain unique normalizations that may get accidentally caught by the other graphs.
# + id="gWG6ttyd-bbD"
classify = (
pynutil.add_weight(time_graph, 1.1)
| pynutil.add_weight(whitelist_graph, 1.07)
| pynutil.add_weight(decimal_graph, 1.1)
| pynutil.add_weight(cardinal_graph, 1.2)
| pynutil.add_weight(ordinal_graph, 1.1)
| pynutil.add_weight(money_graph, 1.08)
| pynutil.add_weight(word_graph, 100)
)
punct = pynutil.insert("tokens { ") + pynutil.add_weight(punct_graph, 1.1) + pynutil.insert(" }")
# + [markdown] id="1TH08f8O-fWx"
# Keep in mind that building weights in this manner is hardly a rule for grammar development and is instead intended as a means to initialize weights for empirical development. You will find that actual strings will cause unexpected behavior that require fine tuning.
#
# For instance, the Classifier for French in NeMo ITN benefits from having varying precedence for some weights, as seen in the following excerpt:
# + id="gKdkyDK3_r46"
class ClassifyFst(GraphFst):
"""
Final class that composes all other classification grammars. This class can process an entire sentence, that is lower cased.
For deployment, this grammar will be compiled and exported to OpenFst Finate State Archiv (FAR) File.
More details to deployment at NeMo/tools/text_processing_deployment.
Args:
cache_dir: path to a dir with .far grammar file. Set to None to avoid using cache.
overwrite_cache: set to True to overwrite .far files
"""
def __init__(self, cache_dir: str = None, overwrite_cache: bool = False):
super().__init__(name="tokenize_and_classify", kind="classify")
far_file = None
if cache_dir is not None and cache_dir != "None":
os.makedirs(cache_dir, exist_ok=True)
far_file = os.path.join(cache_dir, "_fr_itn.far")
if not overwrite_cache and far_file and os.path.exists(far_file):
self.fst = pynini.Far(far_file, mode="r")["tokenize_and_classify"]
logging.info(f"ClassifyFst.fst was restored from {far_file}.")
else:
logging.info(f"Creating ClassifyFst grammars.")
cardinal = CardinalFst()
cardinal_graph = cardinal.fst
fraction = FractionFst(cardinal)
fraction_graph = fraction.fst
ordinal = OrdinalFst(cardinal)
ordinal_graph = ordinal.fst
decimal = DecimalFst(cardinal)
decimal_graph = decimal.fst
measure_graph = MeasureFst(cardinal=cardinal, decimal=decimal, fraction=fraction).fst
date_graph = DateFst(cardinal).fst
word_graph = WordFst().fst
time_graph = TimeFst().fst
money_graph = MoneyFst(cardinal, decimal).fst
whitelist_graph = WhiteListFst().fst
punct_graph = PunctuationFst().fst
electronic_graph = ElectronicFst().fst
telephone_graph = TelephoneFst().fst
classify = (
pynutil.add_weight(whitelist_graph, 1.01)
| pynutil.add_weight(time_graph, 1.05)
| pynutil.add_weight(date_graph, 1.09)
| pynutil.add_weight(decimal_graph, 1.08)
| pynutil.add_weight(measure_graph, 1.1)
| pynutil.add_weight(cardinal_graph, 1.1)
| pynutil.add_weight(ordinal_graph, 1.1)
| pynutil.add_weight(fraction_graph, 1.09)
| pynutil.add_weight(money_graph, 1.07)
| pynutil.add_weight(telephone_graph, 1.1)
| pynutil.add_weight(electronic_graph, 1.1)
| pynutil.add_weight(word_graph, 100)
)
punct = pynutil.insert("tokens { ") + pynutil.add_weight(punct_graph, weight=1.1) + pynutil.insert(" }")
token = pynutil.insert("tokens { ") + classify + pynutil.insert(" }")
token_plus_punct = (
pynini.closure(punct + pynutil.insert(" ")) + token + pynini.closure(pynutil.insert(" ") + punct)
)
graph = token_plus_punct + pynini.closure(delete_extra_space + token_plus_punct)
graph = delete_space + graph + delete_space
self.fst = graph.optimize()
# + [markdown] id="qc4B_0rNcQZu"
# ## FAR import/export
# + [markdown] id="0nRRPvy-AYsA"
# While working through these code excerpts, you may have noticed some latency with each instantiation of our WFSTs (notably whereever `CardinalFst` was involved). This is because the `pynini.optimize` that we call with each graph's instantiation is computationally expensive. For our ultimate purpose of deployment, it seems a waste of resources to recreate stable graphs for each use.
#
# To address this, NeMo ITN supports WFST caching through use of `pynini.Far`, storing and recovering Classify grammars as FAR (Fst ARchives).
#
# Let us update our `ClassifyFst` to permit passing a cache and allowing overwriting (for development):
# + id="5XgWevUzD1AE"
class ClassifyFst(GraphFst):
def __init__(self, cache_dir: str = None, overwrite_cache: bool = False):
super().__init__(name="tokenize_and_classify", kind="classify")
# + [markdown] id="l28GMR70ESz0"
# For storing our graphs as FARs, we can use `graph_utils.generator_main`, which saves our WFSTs by type for easier management. For arguments it takes a string name and a dict mapping of WFST type to graph:
# + id="AzTkcmAWFLYm"
import os
class ClassifyFst(GraphFst):
def __init__(self, cache_dir: str = None, overwrite_cache: bool = False):
super().__init__(name="tokenize_and_classify", kind="classify")
# Grammar here
# ....
if cache_dir is not None and cache_dir != "None":
os.makedirs(cache_dir, exist_ok=True)
far_file = os.path.join(cache_dir, "_fr_itn.far")
generator_main(far_file, {"tokenize_and_classify": self.fst})
# + [markdown] id="Wz8wjCQSD6eJ"
# We pair this with the ability to load from cache (note the `"tokenize_and_classify"` key being passed):
# + id="FRFYgMmuD_53"
import os
class ClassifyFst(GraphFst):
def __init__(self, cache_dir: str = None, overwrite_cache: bool = False):
super().__init__(name="tokenize_and_classify", kind="classify")
if not overwrite_cache and far_file and os.path.exists(far_file):
self.fst = pynini.Far(far_file, mode="r")["tokenize_and_classify"]
else:
# Grammar here
# ....
if cache_dir is not None and cache_dir != "None":
os.makedirs(cache_dir, exist_ok=True)
far_file = os.path.join(cache_dir, "_fr_itn.far")
generator_main(far_file, {"tokenize_and_classify": self.fst})
# + [markdown] id="ib9nggZxF38s"
# Producing our `ClassifyFst` as:
# + id="d2BZyx6sGGg2"
class ClassifyFst(GraphFst):
def __init__(self, cache_dir: str = None, overwrite_cache: bool = False):
super().__init__(name="tokenize_and_classify", kind="classify")
far_file = None
if cache_dir is not None and cache_dir != "None":
os.makedirs(cache_dir, exist_ok=True)
far_file = os.path.join(cache_dir, "_fr_itn.far")
if not overwrite_cache and far_file and os.path.exists(far_file):
self.fst = pynini.Far(far_file, mode="r")["tokenize_and_classify"]
else:
cardinal = CardinalFst()
cardinal_graph = cardinal.fst
ordinal = OrdinalFst(cardinal)
ordinal_graph = ordinal.fst
decimal = DecimalFst(cardinal)
decimal_graph = decimal.fst
whitelist_graph = WhiteList().fst
word_graph = WordFst().fst
time_graph = TimeFst().fst
money_graph = MoneyFst(cardinal, decimal).fst
whitelist_graph = WhiteListFst().fst
punct_graph = PunctuationFst().fst
classify = (
pynutil.add_weight(time_graph, 1.1)
| pynutil.add_weight(whitelist_graph, 1.01)
| pynutil.add_weight(decimal_graph, 1.09)
| pynutil.add_weight(cardinal_graph, 1.1)
| pynutil.add_weight(ordinal_graph, 1.09)
| pynutil.add_weight(money_graph, 1.08)
| pynutil.add_weight(word_graph, 100)
)
punct = pynutil.insert("tokens { ") + pynutil.add_weight(punct_graph, weight=1.1) + pynutil.insert(" }")
token = pynutil.insert("tokens { ") + classify + pynutil.insert(" }")
token_plus_punct = (
pynini.closure(punct + pynutil.insert(" ")) + token + pynini.closure(pynutil.insert(" ") + punct)
)
graph = token_plus_punct + pynini.closure(delete_extra_space + token_plus_punct)
graph = delete_space + graph + delete_space
self.fst = graph.optimize()
if far_file:
generator_main(far_file, {"tokenize_and_classify": self.fst})
# + [markdown] id="nEhY6wKKtfhn"
# You should find the caching to vastly speed up compilement time.
# + [markdown] id="rTtCnC5w95CI"
# # Verbalize and Verbalize Final <a id="verbalize-and-verbalize-final"></a>
# + [markdown] id="H9y5yuk1HaGj"
# Our last step is to create a universal Verbalizer for all classes. This is very similar to development of `ClassifierFst`, except that the Verbalizer breaks its normalization task into two components:
# - `VerbalizeFst`, which removes formatting for each token
# - `VerbalizeFinalFst`, which extends `VerbalizeFst` across all tokens in a string
# Why two componenets when `tokenize_and_classify` was one? Because Sparrowhawk performs all the functionality of `VerbalizeFinalFst`, so its inclusion would break deployment. However, without it, your NeMo grammar would be unable to function at base. So we separate the two to allow the best of both world.
# + [markdown] id="vUawTJVuH8iR"
# ## VerbalizeFst
# + [markdown] id="xghiBV06IIWU"
# Much like `ClassifyFst`, `VerbalizeFst` instanatiates all its subgraphs and then joins them together under a union operation. However, it does not need to employ weighting. Why? Because `ClassifyFst` has assigned each token a specific class. As each class is unique, there is no possibility that a subgraph will be employed for the wrong token.
#
# As such, our `VerbalizeFst` is formed by a simple union operation across all previous Verbalizer graphs:
# + id="uMVCqCvsIt2v"
from nemo_text_processing.inverse_text_normalization.fr.verbalizers.cardinal import CardinalFst
from nemo_text_processing.inverse_text_normalization.fr.verbalizers.decimal import DecimalFst
from nemo_text_processing.inverse_text_normalization.fr.verbalizers.money import MoneyFst
from nemo_text_processing.inverse_text_normalization.fr.verbalizers.ordinal import OrdinalFst
from nemo_text_processing.inverse_text_normalization.fr.verbalizers.time import TimeFst
from nemo_text_processing.inverse_text_normalization.fr.verbalizers.whitelist import WhiteListFst
from nemo_text_processing.inverse_text_normalization.fr.verbalizers.word import WordFst
class VerbalizeFst(GraphFst):
def __init__(self):
super().__init__(name="verbalize", kind="verbalize")
cardinal = CardinalFst()
cardinal_graph = cardinal.fst
ordinal_graph = OrdinalFst().fst
decimal = DecimalFst()
decimal_graph = decimal.fst
whitelist_graph = WhiteListFst().fst
money_graph = MoneyFst(decimal=decimal).fst
time_graph = TimeFst().fst
graph = (
time_graph
| whitelist_graph
| money_graph
| ordinal_graph
| decimal_graph
| cardinal_graph
)
self.fst = graph
# + [markdown] id="Wap-LU6EI2Iu"
# ## Verbalize Final
# + [markdown] id="TYaEt_0tI47t"
# With `VerbalizeFst` complete, we now extend our graph to cover any series of tokens. All this requires is deletion of the `tokens` formatting (note the absence of such in our previous graph) and use of closure for any series of one or more tokens.
#
# This provides the following graph:
# + id="L-9lJNE6JPCW"
class VerbalizeFinalFst(GraphFst):
def __init__(self):
super().__init__(name="verbalize_final", kind="verbalize")
verbalize = VerbalizeFst().fst
word = WordFst().fst
types = verbalize | word
graph = (
pynutil.delete("tokens")
+ delete_space
+ pynutil.delete("{")
+ delete_space
+ types
+ delete_space
+ pynutil.delete("}")
)
graph = delete_space + pynini.closure(graph + delete_extra_space) + graph + delete_space
self.fst = graph
# + [markdown] id="WwMKFw-QJVgm"
# Unlike `ClassifyFst`, NeMo ITN does not cache `VerbalizeFst` or `VerbalizeFinalFst`. (While you are welcome to provide such functionality in your own development, keep in mind that the limited complexity of our Verbalizers makes compilement times less significant.)
# + [markdown] id="7U21AZearZMK"
# # Deployment <a id="deployment"></a>
# + [markdown] id="VrSccoh9K6JK"
# Now that we have done all the groundwork, we can finally move to deployment. This final section will just cover the minor code alterations required to call your language through NeMo ITN and deploy through Sparrowhawk. For further information on using NeMo ITN, please see [this tutorial](https://colab.research.google.com/github/NVIDIA/NeMo/blob/stable/tutorials/text_processing/Inverse_Text_Normalization.ipynb).
# + [markdown] id="0Le2aJvFIAKd"
# ## InverseNormalize
# + [markdown] id="r2R3TUCDLi5-"
# NeMo calls upon the `InverseNormalizer` class for all ITN tasks. Given a string and language, it will instantiate both the `ClassifierFst` and `VerbalizeFst` respective for the given language. (Note: we do not use `VerbalizeFinal` as its functions are managed by Sparrowhawk.) To make your language deployable in the general NeMo ITN system, you must designate the availbility of these classes for instantiation. (For more information, see the [source code](https://github.com/NVIDIA/NeMo/blob/main/nemo_text_processing/inverse_text_normalization/inverse_normalize.py).)
#
# To do so requires only two changes. The first is providing a string to identify your language as an option for `parse_args` ([ISO codes are advised](https://en.wikipedia.org/wiki/List_of_ISO_639-1_codes)):
# + id="tfv4Ee3ML-Fg"
def parse_args():
parser = ArgumentParser()
parser.add_argument("input_string", help="input string", type=str)
parser.add_argument("--language", help="language", choices=['en', 'de', 'es', 'ru', 'fr', 'MY_LANGUAGE'], default="en", type=str)
parser.add_argument("--verbose", help="print info for debugging", action='store_true')
parser.add_argument("--overwrite_cache", help="set to True to re-create .far grammar files", action="store_true")
parser.add_argument(
"--cache_dir",
help="path to a dir with .far grammar file. Set to None to avoid using cache",
default=None,
type=str,
)
return parser.parse_args()
# + [markdown] id="awVl5nAsMUTl"
# The next is to call your `ClassifyFst` and `VerbalizeFst` from `__init__`:
# -
class InverseNormalizer(Normalizer):
def __init__(self, lang: str = 'en', cache_dir: str = None, overwrite_cache: bool = False):
if lang == 'en':
from nemo_text_processing.inverse_text_normalization.en.taggers.tokenize_and_classify import ClassifyFst
from nemo_text_processing.inverse_text_normalization.en.verbalizers.verbalize_final import (
VerbalizeFinalFst,
)
# Other languages
# ....
elif lang == 'MY_LANGUAGE':
from nemo_text_processing.inverse_text_normalization.MY_LANGUAGE.taggers.tokenize_and_classify import ClassifyFst
from nemo_text_processing.inverse_text_normalization.MY_LANGUAGE.verbalizers.verbalize_final import (
VerbalizeFst,
)
# + [markdown] id="TI1PuejLMxdI"
# And you're done! NeMo will handle the rest.
# + [markdown] id="xrksINQoICfj"
# ## Sparrowhawk
# + [markdown] id="rP9-dmMJSg3h"
# Sparrowhawk is an open-source implementation of Google's Kestrel Text Normalization system. Functionally it operates similar to NeMo ITN (the two-step Classify and Verbalize functions stem from [intentional NeMo integration](https://https://arxiv.org/pdf/2104.05055.pdf) but is better optimized for backend deployment.
#
# Like the preceeding section, this portion of the tutorial will highlight a few necessary edits so you may deploy your normalization system.
# + [markdown] id="u1eGMGxkVZmM"
# ### Grammar Export
# + [markdown] id="v9dr0E-uVgoT"
# The first step in deploying your grammar is by exporting both `ClassifyFst` and `VerbalizeFst` WFST as FAR files. This is done through `pynini_export.py`, found in `NeMo/tools/text_processing_deployment`. To allow export of your grammar, we must make the similar edits as wed did for `inverse_normalize.py`
# + [markdown] id="qtek2bMMWbMj"
# First append your language to the list of accepted strings in `parse_args`
# + id="5pTGX9YAWiTZ"
def parse_args():
parser = ArgumentParser()
parser.add_argument("--output_dir", help="output directory for grammars", required=True, type=str)
parser.add_argument("--language", help="language", choices=["en", "de", "es", "ru", 'fr', 'MY_LANGUAGE'], type=str, default='en')
parser.add_argument(
"--grammars", help="grammars to be exported", choices=["tn_grammars", "itn_grammars"], type=str, required=True
)
parser.add_argument(
"--input_case", help="input capitalization", choices=["lower_cased", "cased"], default="cased", type=str
)
parser.add_argument("--overwrite_cache", help="set to True to re-create .far grammar files", action="store_true")
parser.add_argument(
"--cache_dir",
help="path to a dir with .far grammar file. Set to None to avoid using cache",
default=None,
type=str,
)
return parser.parse_args()
# + [markdown] id="Fm3CTmdLWlUt"
# And then call `ClassifyFst` and `VerbalizeFinalFst` in `main`
# +
LANG="FOO"
if LANG == 'en':
from nemo_text_processing.inverse_text_normalization.en.taggers.tokenize_and_classify import (
ClassifyFst as ITNClassifyFst,
)
from nemo_text_processing.inverse_text_normalization.en.verbalizers.verbalize import (
VerbalizeFst as ITNVerbalizeFst,
)
# Other languages
# ...
elif LANG == 'MY_LANGUAGE':
from nemo_text_processing.inverse_text_normalization.MY_LANGUAGE.taggers.tokenize_and_classify import (
ClassifyFst as ITNClassifyFst,
)
from nemo_text_processing.inverse_text_normalization.MY_LANGUAGE.verbalizers.verbalize import (
VerbalizeFst as ITNVerbalizeFst,
)
# + [markdown] id="JFgGhCMMW3UQ"
# ### Deployment
# + [markdown] id="V8RH0aGbW41U"
# By default, NeMo ITN is structured to allow deployment through a Docker based backend. This involved building a container from file, exporting your grammars to the container and then deploying Sparrowhawk for processing.
#
# NeMo automates this entire process through `export_grammars.sh`, which will automatically compile your grammars for deployment (assuming you edited `pynini_export` appropriately) and mount them in a container for you. For our purposes, `export_grammar` only requires the following arguments:
# - `LANGUAGE` - the string you have used throughout to indicate your language
# - `GRAMMARS` - only accepts `itn_grammars`(Inverse Text Normalization) or `tn_grammars` (Text Normalization)
#
# For instance, we would call our French ITN with:
# + [markdown] id="KYdbawAfZIco"
# `bash export_grammar.sh --GRAMMARS=itn_grammars --LANGUAGE={LANGUAGE}`
# + [markdown] id="UXVr2twdZMO2"
# Which will return the Docker prompt for further normalization.
# + [markdown] id="TDoVUxCE-Dax"
# # Final Notes
# + [markdown] id="Fw-9mU7ql8iY"
# Congratulations, you have now constructed an entire ITN system from the ground up! While your experience will vary with each language, you will find several commonalities that will assist you in further development.
#
# If you are interested in working further with your language WFSTs, you may wish to construct a TN system. Broadly, this is accomplished by inverting your previous graphs (`pynini.invert` may assist here) and changing your outputs to avoid indeterminancy (i.e. decide on one canonical output for your grammar for each class). But outside of such grammar specific edits, you repeat many of the steps exhibited here, such as:
# - Use of a two step classifier-verbalizer system
# - Same semiotic classes for tagging
# - Inheritance of `GraphFst`
# - Minor import edits to `pynini_export` and `export_grammar`
| tutorials/text_processing/WFST_Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
from google.cloud import firestore
import gcsfs
import pickle5 as pickle
import textblob
import nltk
# %env GOOGLE_APPLICATION_CREDENTIALS="/Users/marcelcanhisares/Development/each/covid-news/keys/covid-news-a9d0938c837d.json"
DB = firestore.Client.from_service_account_json("/Users/marcelcanhisares/Development/each/covid-news/keys/covid-news-a9d0938c837d.json")
# +
KEYWORDS = ["corona", "covid", "pandemia", "SARS", "covid-19"]
STOP_WORDS = [ 'a', 'à', 'agora', 'ainda', 'alguém', 'algum', 'alguma', 'algumas', 'alguns', 'ampla', 'amplas', 'amplo', 'amplos', 'ante', 'antes', 'ao', 'aos', 'após', 'aquela', 'aquelas', 'aquele', 'aqueles', 'aquilo', 'as', 'às', 'até', 'através', 'cada', 'coisa', 'coisas', 'com', 'como', 'contra', 'contudo', 'da', 'daquele', 'daqueles', 'das', 'de', 'dela', 'delas', 'dele', 'deles', 'depois', 'dessa', 'dessas', 'desse', 'desses', 'desta', 'destas', 'deste', 'destes', 'deve', 'devem', 'devendo', 'dever', 'deverá', 'deverão', 'deveria', 'deveriam', 'devia', 'deviam', 'disse', 'disso', 'disto', 'dito', 'diz', 'dizem', 'do', 'dos','e','é','ela','elas','ele','eles','em','entre','era','eram','éramos','essa','essas','esse','esses','esta','está','estamos','estão','estas','estava','estavam','estávamos','este','esteja','estejam','estejamos','estes','esteve','estive','estivemos','estiver','estivera','estiveram','estivéramos','estiverem','estivermos','estivesse','estivessem','estivéssemos','estou','eu','foi','fomos','for','fora','foram','fôramos','forem','formos','fosse','fossem','fôssemos','fui','há','haja','hajam','hajamos','hão','havemos','havia','hei','houve','houvemos','houver','houvera','houverá','houveram','houvéramos','houverão','houverei','houverem','houveremos','houveria','houveriam','houveríamos','houvermos','houvesse','houvessem','houvéssemos','isso','isto','já','lhe','lhes','mais','mas','me','mesmo','meu','meus','minha','minhas','muito','na','não','nas','nem','no','nos','nós','nossa','nossas','nosso','nossos','num','numa','o','os','ou','para','pela','pelas','pelo','pelos','por','qual','quando','que','quem','são','se','seja','sejam','sejamos','sem','ser','será','serão','serei','seremos','seria','seriam','seríamos','seu','seus','só','sobre','somos','sou','sua','suas','também','te','tem','têm','temos','tenha','tenham','tenhamos','tenho','ter','terá','terão','terei','teremos','teria','teriam','teríamos','teu','teus','teve','tinha','tinham','tínhamos','tive','tivemos','tiver','tivera','tiveram','tivéramos','tiverem','tivermos','tivesse','tivessem','tivéssemos','tu','tua','tuas','um','uma','você','vocês','vos']
# -
fs = gcsfs.GCSFileSystem(token="/Users/marcelcanhisares/Development/each/covid-news/keys/covid-news-a9d0938c837d.json", project="covid-news-291320")
def getClassifier():
directory = "covid-news-291320.appspot.com"
filename = "Classificador.pickle"
with fs.open(directory + '/' + filename, 'rb') as handle:
return pickle.load(handle)
document = DB.collection('articles').document('0006126928ebb6f907e325da8a67b3944a6d1f07540639d03d1268f0edaca0f3')
document.get().to_dict()
collection = DB.collection('articles').stream()
articlesDocs = []
for document in collection:
articlesDocs.append(document.to_dict())
def removeArticleSource(text:str):
index = text.rfind('-')
return text[:index - 1] if index > 1 else text
def removeStopWords(text: str):
splitedResults = []
splited = text.split()
for word in splited:
if word not in STOP_WORDS and word not in KEYWORDS:
splitedResults.append(word)
return splitedResults
def stemWords(words):
stemmedWords = []
stemmer = nltk.stem.RSLPStemmer()
for word in words:
stemmedWords.append(stemmer.stem(word))
return stemmedWords
def classifyText(classifier, text):
words = removeStopWords(text)
stemmedWords = stemWords(words)
textToClassify = " ".join(stemmedWords)
return classifier.classify(textToClassify)
classifier = getClassifier()
def updateDocument(item_id, field_updates):
doc = DB.collection('articles').document(item_id)
doc.update(field_updates)
for article in articlesDocs:
if article["category"] is None:
field_updates = {}
field_updates["title"] = removeArticleSource(article["title"])
field_updates["category"] = classifyText(classifier, article["title"])
updateDocument(article["articleId"], field_updates)
articlesDocs[1]
classifyText(classifier, articlesDocs[1]["title"])
| classifica_artigos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
import numpy as np
# %load_ext autoreload
# %autoreload 2
import pinot
import math
import os
os.environ['KMP_DUPLICATE_LIB_OK']='True'
# +
# define the target
f = lambda x: torch.sin(x)
x_tr = torch.tensor(np.concatenate(
[
np.linspace(-3 * math.pi, -math.pi, 50),
np.linspace(math.pi, 3 * math.pi, 50)
]),
dtype=torch.float32)[:, None]
x_te = torch.tensor(np.concatenate(
[
np.linspace(-7 * math.pi, -3 * math.pi, 100),
np.linspace(-math.pi, math.pi, 50),
np.linspace(3 * math.pi, 7 * math.pi, 100)
]),
dtype=torch.float32)[:, None]
x = torch.tensor(np.linspace(-7 * math.pi, 7 * math.pi, 375),
dtype=torch.float32)[:, None]
y_tr = f(x_tr)
y_te = f(x_te)
y = f(x)
# -
x.shape
# +
base_kernel=pinot.inference.gp.kernels.rbf.RBF()
kernel = pinot.inference.gp.kernels.deep_kernel.DeepKernel(
lambda x:x,
base_kernel=base_kernel)
gpr = pinot.inference.gp.gpr.sparse_variational_gpr.SVGPR(
kernel,
log_sigma=-5.0,
n_inducing_points=100,
initializer_std=0.1,
in_features=50,
kl_loss_scaling=0.0,
grid_boundary=30)
# -
gpr.kernel.representation(x).shape
opt = torch.optim.Adam(gpr.parameters(), 1e-3)
for _ in range(1000):
opt.zero_grad()
loss = gpr.loss(x_tr, y_tr).sum()
loss.backward()
print(loss)
opt.step()
gpr.condition(x)
y_hat_distribution = gpr.condition(x)
y_hat_distribution.sample()
from matplotlib import pyplot as plt
plt.plot(y_hat_distribution.variance.detach())
def plot():
from pinot.inference.utils import confidence_interval
from matplotlib import pyplot as plt
plt.figure(figsize=(10, 6))
# y_hat_distribution = pinot.inference.utils.condition_mixture(net, x, sampler=opt, n_samples=1000)
nll = y_hat_distribution.log_prob(y).mean() / float(y.shape[0])
plt.plot(x.flatten(), y_hat_distribution.mean.detach().numpy().flatten(), c='k', linewidth=5)
low, high = confidence_interval(y_hat_distribution, 0.95, n_samples=10)
plt.fill_between(x.flatten(), low.flatten().detach(), high.flatten().detach(), color='k', alpha=0.1)
low, high = confidence_interval(y_hat_distribution, 0.75, n_samples=10)
plt.fill_between(x.flatten(), low.flatten().detach(), high.flatten().detach(), color='k', alpha=0.2)
low, high = confidence_interval(y_hat_distribution, 0.50, n_samples=10)
plt.fill_between(x.flatten(), low.flatten().detach(), high.flatten().detach(), color='k', alpha=0.3)
low, high = confidence_interval(y_hat_distribution, 0.25, n_samples=10)
plt.fill_between(x.flatten(), low.flatten().detach(), high.flatten().detach(), color='k', alpha=0.4)
plt.scatter(x_te, y_te, label='te')
plt.scatter(x_tr, y_tr, label='tr')
plt.legend()
plt.ylim(-2, 2)
plt.title('NLL=%s' % nll.detach().numpy().round(3))
plot()
| scripts/gp/gp_variational_playground.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
import codecademylib
from matplotlib import pyplot as plt
months = ["Jan", "Feb", "Mar", "Apr", "May", "Jun", "Jul", "Aug", "Sep", "Oct", "Nov", "Dec"]
visits_per_month = [9695, 7909, 10831, 12942, 12495, 16794, 14161, 12762, 12777, 12439, 10309, 8724]
# numbers of limes of different species sold each month
key_limes_per_month = [92.0, 109.0, 124.0, 70.0, 101.0, 79.0, 106.0, 101.0, 103.0, 90.0, 102.0, 106.0]
persian_limes_per_month = [67.0, 51.0, 57.0, 54.0, 83.0, 90.0, 52.0, 63.0, 51.0, 44.0, 64.0, 78.0]
blood_limes_per_month = [75.0, 75.0, 76.0, 71.0, 74.0, 77.0, 69.0, 80.0, 63.0, 69.0, 73.0, 82.0]
# create your figure here
plt.figure(figsize=(12, 8))
ax1 = plt.subplot(1, 2, 1)
x_values = range(len(months))
plt.plot(x_values, visits_per_month, marker='s')
plt.xlabel('Months')
plt.ylabel('Number of Visits')
plt.title('Number of Site Visits Per Month')
ax1.set_xticks(x_values)
ax1.set_xticklabels(months)
ax2 = plt.subplot(1, 2, 2)
plt.plot(x_values, key_limes_per_month, color='red', label='Key Limes', marker='o')
plt.plot(x_values, persian_limes_per_month, color='green', label='Persian Limes', marker='o')
plt.plot(x_values, blood_limes_per_month, color='purple', label='Blood Limes', marker='o')
plt.xlabel('Months')
plt.ylabel('Number Sold')
plt.title('Limes Sold Per Month')
ax2.set_xticks(x_values)
ax2.set_xticklabels(months)
plt.legend()
plt.subplots_adjust(wspace=.25)
plt.show()
plt.savefig('Sublime Limes 2020.png')
| Sublime Limes Line Graphs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/StephenSpicer/Unit_02_Clone/blob/master/Stephen_Lupsha_DS23_Sprint_Challenge_22.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] deletable=false editable=false id="M3XH_XLsy_Bn" nbgrader={"cell_type": "markdown", "checksum": "3cda62270676bf56b34fe29465f1add1", "grade": false, "grade_id": "cell-7b7a3d8bc00f7c5d", "locked": true, "schema_version": 3, "solution": false, "task": false}
# _Lambda School Data Science, Unit 2_
#
# ---
# + [markdown] id="2zUhd6ww1ajT"
# 👇 **Do not change the code in this cell.** If you're working in Google Colab, you can run this cell to install `category_encoders`.
# + deletable=false editable=false id="-a7Lo3XiOUrH" nbgrader={"cell_type": "code", "checksum": "2cf805202878a43b6677e1d45ac4da6d", "grade": false, "grade_id": "cell-fd3a0dbfc4b9265f", "locked": true, "schema_version": 3, "solution": false, "task": false}
# %%capture
import sys
if 'google.colab' in sys.modules:
# Install packages in Colab
# !pip install category_encoders
# + [markdown] deletable=false editable=false id="VGLNhZO_OUrM" nbgrader={"cell_type": "markdown", "checksum": "88ba1fa9ea78f0afb94fcb9aa53f7435", "grade": false, "grade_id": "cell-d6eff2274645bd61", "locked": true, "schema_version": 3, "solution": false, "task": false}
#
# # Sprint Challenge: Predict <NAME>'s Shots 🏀
#
# For your Sprint Challenge, you'll use a dataset with all <NAME>ry's NBA field goal attempts from October 2009 through June 2019 (regular season and playoffs). The dataset was collected with the [nba_api](https://github.com/swar/nba_api) Python library.
#
# You'll create a model to predict whether Curry will make a shot based on his past perfomance.
# + [markdown] deletable=false editable=false id="K3xt9YTSOUrN" nbgrader={"cell_type": "markdown", "checksum": "746679bdefe8a36452807b584846628e", "grade": false, "grade_id": "cell-9aed6a1cff735430", "locked": true, "schema_version": 3, "solution": false, "task": false}
# # Directions
#
# This notebook contains 10 tasks, which cover the material we've learned in this sprint. Here's a summary:
#
# - **Task 1:** Importing data.
# - **Task 2:** Feature engineering.
# - **Task 3:** Splitting data into a feature matrix and target vector.
# - **Task 4:** Splitting data into training, validation, and test sets.
# - **Task 5:** Establishing baseline accuracy.
# - **Task 6:** Building a model with a transformer and a tree-based predictor.
# - **Task 7:** Calculating training and validation accuracy.
# - **Task 8 (`stretch goal`):** Tuning model hyperparameters.
# - **Task 9:** Calculating precision and recall from a confusion matrix.
# - **Task 10 (`stretch goal`):** Plotting a confusion matrix.
#
# For each task you should do the following:
#
# - Read the task instructions.
# - Write your code in the cell below the task. Delete the `raise
# NotImplementedError` before your start.
# - Run the testing cell below the task. If you get an error, read the error message and re-evaluate your code.
#
# **You should limit your code to the following libraries:**
#
# - `category_encoders`
# - `numpy`
# - `matplotlib`
# - `pandas`
# - `sklearn`
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="zhnJVWNl1lDK" outputId="2de6b6ec-34eb-4a1e-c120-fd53bfbf30e9"
# PREAMBLE - we hold these packages to be self evident.
# thank god for those who came before us, we stand on their shoulders.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# import seaborn as sns
# that takes care of the basics now for the sci-kit
import category_encoders as ce
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline
from category_encoders import OneHotEncoder, OrdinalEncoder
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import FunctionTransformer
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import GridSearchCV, RandomizedSearchCV
from sklearn.metrics import accuracy_score
from sklearn.model_selection import cross_val_score
from sklearn.tree import plot_tree
# + [markdown] deletable=false editable=false id="8CakIhmwOUrP" nbgrader={"cell_type": "markdown", "checksum": "eea5568a328ffc86470fc220567bfcd4", "grade": false, "grade_id": "cell-09a1516a86d168cc", "locked": true, "schema_version": 3, "solution": false, "task": false}
# # I. Wrangle Data
#
# **Task 1:** Change the code below to import your dataset. Be sure to examine the columns carefully and determine if one of them should be set at the index.
# + id="dIklN7u34R9D"
def wrangle(df):
# We know it's steph curry, I think we can drop that.
df = df.copy()
df.drop(columns=['player_name'], inplace=True)
df['seconds_remaining_inperiod'] = df['minutes_remaining'] + df['seconds_remaining']
df['seconds_remaining_ingame'] = df['seconds_remaining_inperiod'] + df['period']
df['homecourt_adv'] = df['htm'] == 'GSW'
return df
# + deletable=false id="NWVOhicLOUrQ" nbgrader={"cell_type": "code", "checksum": "3a6e0dfee7d760919c2fd0182e764b5f", "grade": false, "grade_id": "225SC_01a_q", "locked": false, "schema_version": 3, "solution": true, "task": false}
'''T1. Import DataFrame `df`'''
url = 'https://drive.google.com/uc?export=download&id=1fL7KPyxgGYfQDsuJoBWHIWwCAf-HTFpX'
df = pd.read_csv(url,
parse_dates=['game_date'],
index_col='game_date')
df = wrangle(df)
# + colab={"base_uri": "https://localhost:8080/", "height": 340} id="MF6Ddyou3dJX" outputId="ec2aef00-17c4-4471-cedf-1b1ae8b17c66"
df.head()
# + id="fSDKA3Os1ajV" colab={"base_uri": "https://localhost:8080/"} outputId="1558fcc7-1cb7-4357-daab-8cb63e5770f7"
type(df.index)
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "9d175018548e99ccb464c05bcd0a8aec", "grade": false, "grade_id": "cell-7738d198f39f8908", "locked": true, "schema_version": 3, "solution": false, "task": false} id="Thc3dfzq1ajV"
# **Task 1 Testing**
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "5b221a1569fb4bb63422ef4860840fb6", "grade": true, "grade_id": "cell-6071eb9ae89cfaca", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="XLVlVL5b1ajW"
'''T1 Testing'''
assert isinstance(df, pd.DataFrame), 'Have you created a DataFrame named `df`?'
assert len(df) == 13958, 'Is `df` the correct length?'
# + [markdown] deletable=false editable=false id="1YaoMGLTOUrS" nbgrader={"cell_type": "markdown", "checksum": "a3b8a435c3baf7116b611f2e8de888e8", "grade": false, "grade_id": "cell-8490d284c49fed2c", "locked": true, "schema_version": 3, "solution": false, "task": false}
# **Task 2a:** Engineer at least 1 new feature. You can use something from the list below or your own idea.
#
# - **Homecourt Advantage**: Is the home team (`htm`) the Golden State Warriors (`GSW`) ?
# - **Opponent**: Who is the other team playing the Golden State Warriors?
# - **Seconds remaining in the period**: Combine minutes remaining with seconds remaining, to get the total number of seconds remaining in the period.
# - **Seconds remaining in the game**: Combine period, and seconds remaining in the period, to get the total number of seconds remaining in the game. A basketball game has 4 periods, each 12 minutes long.
# - **Made previous shot**: Was <NAME>'s previous shot successful?
#
# **Task 2b (`stretch goal — optional`):** Create a total of 3 new features, using your own ideas or the list above. Make sure you're not just duplicating features that are already in the feature matrix.
# + deletable=false id="X_rdysSxOUrS" nbgrader={"cell_type": "code", "checksum": "b2678ff5b2c193d4af565689966550e8", "grade": false, "grade_id": "225SC_01b_q", "locked": false, "schema_version": 3, "solution": true, "task": false} colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="5aac7f77-4858-427c-f206-3287ed8e0251"
"""T2. Create new feature"""
# df['seconds_remaining_inperiod'] = df['minutes_remaining'] + df['seconds_remaining']
# df['seconds_remaining_ingame'] = df['seconds_remaining_inperiod'] + df['period']
# df['homecourt_adv'] = df['htm'] == 'GSW'
# + [markdown] deletable=false editable=false id="9JivGpcEOUrV" nbgrader={"cell_type": "markdown", "checksum": "18dbccc05e878201fa3b667e5937b755", "grade": false, "grade_id": "cell-58c684eafc846d48", "locked": true, "schema_version": 3, "solution": false, "task": false}
# **Task 2 Test**
# + deletable=false editable=false id="700-jf3nOUrW" nbgrader={"cell_type": "code", "checksum": "188d989e0e75d8376d5dce4ca8ae3fd6", "grade": true, "grade_id": "225SC_01_a", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
'''T2 Testing'''
assert df.shape[1] >= 20, '`df` does not appear to have new features.'
# + [markdown] deletable=false editable=false id="Nh3whBpUOUrZ" nbgrader={"cell_type": "markdown", "checksum": "d4c27458bcc532324a3a506a1cf41a2c", "grade": false, "grade_id": "cell-965e44063af534b5", "locked": true, "schema_version": 3, "solution": false, "task": false}
# # II. Split Data
#
# **Task 3:** Divide the DataFrame `df` into the feature matrix `X` and the target vector `y`. Your target is `'shot_made_flag'`.
# + deletable=false id="1S9T0AfMOUrZ" nbgrader={"cell_type": "code", "checksum": "0f30252ff9e8f3519a29d0364bb2f53b", "grade": false, "grade_id": "225SC_02a_q", "locked": false, "schema_version": 3, "solution": true, "task": false} colab={"base_uri": "https://localhost:8080/"} outputId="e213ee0c-1fcd-448b-c007-13c5f12d6931"
'''T3. Create `X` and `y`.'''
# target
target = 'shot_made_flag'
#feature matrix
X = df.drop(columns=target)
# target vector
y = df[target]
print('features matrix shape: ', X.shape)
print()
print( 'target vector shape: ', y.shape)
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "4e82cdb48c341d83735b201986c66813", "grade": false, "grade_id": "cell-6ad9cd44f4e272c6", "locked": true, "schema_version": 3, "solution": false, "task": false} id="jhn4Ts4B1ajX"
# **Task 3 Test**
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "a0c8ebd72a7b107af8a8cb73e3a1f530", "grade": true, "grade_id": "cell-1a89d8c845a9424c", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="-4R_IiDN1ajX"
'''T3 Tests'''
assert y.shape == (13958,), '`y` either has the wrong number of rows, or is two-dimentional.'
assert len(X) == 13958, '`X` has the wrong number of rows.'
assert X.shape[1] > 1, '`X` has the wrong number of columns'
assert 'shot_made_flag' not in X.columns, 'Target is still part of feature matrix.'
# + [markdown] deletable=false editable=false id="YE1j6w2lOUrc" nbgrader={"cell_type": "markdown", "checksum": "efdf9f11506650d3ae868cf4811f8292", "grade": false, "grade_id": "cell-5f93454935747581", "locked": true, "schema_version": 3, "solution": false, "task": false}
# **Task 4:** Divide your dataset into training, validation, and test sets.
#
# - Your training set (`X_train`, `y_train`) should contain games from the 2009-10 season through the end of the 2016-17 season.
# - Your validation set (`X_val`, `y_val`) should contain games from the 2017-18 season.
# - Your test set (`X_test`, `y_test`) should contain games from the 2018-2019 season.
# - **Tip:** The NBA season begins in October and ends in June.
# + deletable=false id="9BvdaqRoOUrc" nbgrader={"cell_type": "code", "checksum": "a786d072437c2876b466722a64ff5085", "grade": false, "grade_id": "225SC_02b_q", "locked": false, "schema_version": 3, "solution": true, "task": false} colab={"base_uri": "https://localhost:8080/"} outputId="24262154-5ce6-4301-842f-1e8018772a5e"
'''T4. Create training and val'''
cutoff = '2017-06-18'
cutoff_val = '2018-06-18'
mask_train = X.index < cutoff
mask_val = (X.index > cutoff) & (X.index < cutoff_val)
mask_test = X.index > cutoff_val
X_train, y_train = X.loc[mask_train], y.loc[mask_train]
X_val, y_val = X.loc[mask_val], y.loc[mask_val]
X_test, y_test = X.loc[mask_test], y.loc[mask_test]
#let's get some shapes
print('X_Train Shape:', X_train.shape,'y_train Shape:', y_train.shape)
print()
print('X_val Shape:', X_val.shape, 'y_val Shape:', y_val.shape)
print()
print('X_test shape: ', X_test.shape, 'y_test shape: ', y_test.shape)
# + [markdown] deletable=false editable=false id="IfMncrzIOUre" nbgrader={"cell_type": "markdown", "checksum": "d7a7ca9cdb31cdddad5c680c31a0d947", "grade": false, "grade_id": "cell-025dae513375f87c", "locked": true, "schema_version": 3, "solution": false, "task": false}
# **Test 4**
# + deletable=false editable=false id="MjBRCehWOUrf" nbgrader={"cell_type": "code", "checksum": "542e32df51f2c93a1f6676c076bbcaa6", "grade": true, "grade_id": "225SC_02_a", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
'''T4 Tests'''
assert len(X_train) == len(y_train) == 11081
assert len(X_val) == len(y_val) == 1168
assert len(X_test) == len(y_test) == 1709
# + [markdown] deletable=false editable=false id="8NYOGp4MOUrh" nbgrader={"cell_type": "markdown", "checksum": "416ca1511e492007ecf7e71ae9634237", "grade": false, "grade_id": "cell-28f236308828133c", "locked": true, "schema_version": 3, "solution": false, "task": false}
# # III. Establish Baseline
#
# **Task 5:** Establish the baseline accuracy score for this classification problem using your training set. Save the score to the variable `baseline_acc`.
# + deletable=false id="6eZiCkSDOUri" nbgrader={"cell_type": "code", "checksum": "c28391427425a03d66b5e324e586f299", "grade": false, "grade_id": "225SC_03_q", "locked": false, "schema_version": 3, "solution": true, "task": false} colab={"base_uri": "https://localhost:8080/"} outputId="79ea4ab2-bd38-498c-90b2-d6ac90bad4ce"
'''T5. Calculate baseline accuracy `baseline_acc`.'''
baseline_acc = df['shot_made_flag'].value_counts(normalize=True).max()
print('Baseline Accuracy:', baseline_acc)
# + [markdown] deletable=false editable=false id="seuVCNLfOUrk" nbgrader={"cell_type": "markdown", "checksum": "a0ef09195e90b4901f5d80da3babd310", "grade": false, "grade_id": "cell-a7dd0a3aa55b90e0", "locked": true, "schema_version": 3, "solution": false, "task": false}
# **Task 5 Testing**
# + deletable=false editable=false id="qNO8Cc0oOUrl" nbgrader={"cell_type": "code", "checksum": "4b10c894698890e348e6d02c786d1e96", "grade": true, "grade_id": "225SC_03_a", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
'''T5 Testing'''
assert isinstance(baseline_acc, float)
assert 0.0 <= baseline_acc <= 1.0
# + [markdown] deletable=false editable=false id="ZhvHv71JOUrn" nbgrader={"cell_type": "markdown", "checksum": "0373e6c6d748ecfb29c1638f8c93f33a", "grade": false, "grade_id": "cell-36a0288a6c00e795", "locked": true, "schema_version": 3, "solution": false, "task": false}
# # IV. Build Model
#
# **Task 6:** Build a model that includes (1) a transformer for categorical features and (2) a tree-based predictor. You should combine these two components (and any other pieces you think are necessary) in a pipeline named `model`. Be sure to fit your model to your training data.
# + deletable=false id="UidpqqMwOUro" nbgrader={"cell_type": "code", "checksum": "3fc6b9e2872bf60a7b829741ab8d8774", "grade": false, "grade_id": "225SC_04_q", "locked": false, "schema_version": 3, "solution": true, "task": false} colab={"base_uri": "https://localhost:8080/"} outputId="66e945a1-d025-4cd2-f398-2e913e6f9540"
'''T6. Build a pipeline `model` with encoder and tree-based predictor.'''
model = make_pipeline(
#FunctionTransformer(wrangle, validate=False),
ce.OrdinalEncoder(),
SimpleImputer(strategy='mean'),
RandomForestClassifier(n_estimators=50, random_state=42, n_jobs=-1)
)
# Fit on train, score on val
model.fit(X_train, y_train)
y_pred = model.predict(X_val)
print('Validation Accuracy', accuracy_score(y_val, y_pred))
# + [markdown] deletable=false editable=false id="BdBRcK50OUrq" nbgrader={"cell_type": "markdown", "checksum": "51f76ac7ea3fe7974c0b09203e95ea67", "grade": false, "grade_id": "cell-ab9483c995e69b66", "locked": true, "schema_version": 3, "solution": false, "task": false}
# **Task 6 Testing**
# + deletable=false editable=false id="qw0oHcuQOUrr" nbgrader={"cell_type": "code", "checksum": "120cc78af9936eedd151799c30bb6f2f", "grade": true, "grade_id": "225SC_04_a", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
'''T6 Testing'''
from sklearn.pipeline import Pipeline
assert isinstance(model, Pipeline), '`model` should be type `Pipeline`.'
# Does `model` Pipeline have multiple components?
assert len(model.named_steps.keys()) > 1, '`model` should have more than one component.'
# Has `model` been trained?
assert hasattr(model, 'classes_'), 'Have you trained `model`?'
# + [markdown] deletable=false editable=false id="k_A786FUOUrt" nbgrader={"cell_type": "markdown", "checksum": "b830a678806f0afb1fae00d3aae4019c", "grade": false, "grade_id": "cell-c4ae57b31cfead21", "locked": true, "schema_version": 3, "solution": false, "task": false}
# # V. Check Metrics
#
# **Task 7:** Check the training and validation accuracy of your model, and assign the scores to `train_acc` and `val_acc` respectively.
# + deletable=false id="gHSZCmBhOUru" nbgrader={"cell_type": "code", "checksum": "57204172c2fe58d81276cfcb3fe2b88e", "grade": false, "grade_id": "225SC_05_q", "locked": false, "schema_version": 3, "solution": true, "task": false} colab={"base_uri": "https://localhost:8080/"} outputId="9b60ab4f-f549-4d89-8dbd-c2001b89f5f7"
'''T7. Calculate train and test accuracy.'''
train_acc = model.score(X_train, y_train)
val_acc = model.score(X_val, y_val)
print('Training Accuracy Score:', train_acc)
print('Validation Accuracy Score:', val_acc)
# + [markdown] deletable=false editable=false id="cejPfZyvOUrw" nbgrader={"cell_type": "markdown", "checksum": "5f26eacfafe6f5b12083dfe93033add4", "grade": false, "grade_id": "cell-1c8ee42427653f95", "locked": true, "schema_version": 3, "solution": false, "task": false}
# **Task 7 Test**
# + deletable=false editable=false id="6irQ3xW5OUrw" nbgrader={"cell_type": "code", "checksum": "c5f81eb42f3976b48fd0561541addb87", "grade": true, "grade_id": "225SC_05_a", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
'''T7 Testing'''
# Is `training_acc` a float between 0.0 and 1.0?
assert isinstance(train_acc, float)
assert 0.0 <= train_acc <= 1.0
# Is `validation_acc` a float between 0.0 and 1.0?
assert isinstance(val_acc, float)
assert 0.0 <= val_acc <= 1.0
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "26c6585ca7e1d35df0b4f509ad79ac41", "grade": false, "grade_id": "cell-40a154aa7afa5ce0", "locked": true, "schema_version": 3, "solution": false, "task": false} id="8dDYHKAG1aja"
# # VI. Tune Model
#
# **Task 8 (`stretch goal — optional`):** Using your training and validation sets as a guide, tune the hyperparameters of your model to see if you can improve its accuracy. You can perform your tuning "by hand" or using [`RandomizedSearchCV`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.RandomizedSearchCV.html).
#
# - Important hyperparameters for all tree-based models: `max_depth`.
# - Important hyperparameters for random forest models: `n_estimators`, `max_samples`.
# - Important hyperparametes for gradient boosting tree models: `n_estimators`, `learning_rate`.
# - **TIP:** If you use `RandomizedSearchCV`, set `n_iter` to `3`. Any larger and the auto-grader will stop working.
# - **TIP:** Since you already created a validation set above, there is no need to do k-fold cross-validation here. Set `cv` to `None`.
#
# Once you're satisfied with your model's performance on the validation set (if you can get above `0.6`, you're doing good), calculate its accuracy with your test set. Assign the result to `test_acc`.
# + colab={"base_uri": "https://localhost:8080/"} id="GWnQcYmSQkZn" outputId="e9d116e3-ce47-4095-fbe9-5e274af225b9"
X_test.shape
# + deletable=false nbgrader={"cell_type": "code", "checksum": "b5cca689a3aaf250e4f52c3e910bce29", "grade": false, "grade_id": "cell-573c39cfe6281f06", "locked": false, "schema_version": 3, "solution": true, "task": false} id="VWgRahfv1aja" colab={"base_uri": "https://localhost:8080/"} outputId="13152efa-7c89-40ca-c42f-10f9bd6ccc1a"
'''T8. Tune model.'''
paras = {'randomforestclassifier__n_estimators': np.arange(2, 20, 2),
'randomforestclassifier__max_depth': np.arange(10, 21, 2),
'randomforestclassifier__max_samples': np.arange(0.1, .8, 0.1),
'randomforestclassifier__max_features': np.arange(12, 24, 2)}
rf_rs = RandomizedSearchCV(model, param_distributions=paras, n_iter=3, cv=None, n_jobs=-1, verbose=1)
rf_rs.fit(X_train, y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="YUA0Iq1VQ7-G" outputId="93a8b489-d580-4768-f630-e499914f81c5"
rf_rs.best_params_
# + colab={"base_uri": "https://localhost:8080/"} id="Y296MTq5WZQ-" outputId="6b758ad6-f157-4ad6-94c9-63ef396d4f1e"
rf_rs.score(X_train, y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="7ps1dVlFWcrp" outputId="067e6b7a-c9ca-454c-d5d0-7ba57b9e8c03"
rf_rs.score(X_val, y_val)
# + colab={"base_uri": "https://localhost:8080/"} id="seWTkk6oQ1MY" outputId="62dbd7a1-ff1e-4bab-9f01-da039fa280fb"
test_acc = rf_rs.score(X_test, y_test)
print(test_acc)
# + [markdown] id="yxpxGLhv1aja"
# **Task 8 Testing**
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "aaad1200c6f3f83d3f97435d6731e405", "grade": true, "grade_id": "cell-6028e9cba0fc28d3", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="mwyPCdAL1aja"
'''T8 Testing'''
assert isinstance(test_acc, float)
assert 0.0 <= test_acc <= 1.0
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "8db3430fd99f6657dd83938a0d536c1e", "grade": false, "grade_id": "cell-0e008b7c42ac3b70", "locked": true, "schema_version": 3, "solution": false, "task": false} id="UtKNp2bo1aja"
# # VII. Communication
#
# **Task 9:** Below is a confusion matrix for the model that your instructor made for this challenge (based on the **test data**). Calculate the precision and recall of this model, naming them `instructor_precision` and `instructor_recall`, respectively.
#
# 
# + deletable=false nbgrader={"cell_type": "code", "checksum": "b8ae36a67a4ab9283a7f843d9697c57d", "grade": false, "grade_id": "cell-83509e08a73e81b5", "locked": false, "schema_version": 3, "solution": true, "task": false} id="H8BfqBcv1aje" colab={"base_uri": "https://localhost:8080/"} outputId="5a3dcb02-19e3-4dd7-cbf6-9732db9a52af"
'''T9. Calculate precision and recall'''
instructor_precision = 538/(538+387)
instructor_recall = 538/(538+259)
print('Instuctor model precision', instructor_precision)
print('Instuctor model recall', instructor_recall)
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "48c102d0b0ca01e1a1a84f2b31437358", "grade": false, "grade_id": "cell-f312715b3057666e", "locked": true, "schema_version": 3, "solution": false, "task": false} id="H9h8ECsq1ajh"
# **Task 9 Testing**
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "c303f6e8a1ad2f81e090a7f304d6a474", "grade": true, "grade_id": "cell-ec6f47b7dca9966b", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="HWlwE-Jo1ajh"
'''T9 Testing'''
assert isinstance(instructor_precision, float)
assert isinstance(instructor_recall, float)
assert 0 <= instructor_precision <= 1
assert 0 <= instructor_recall <= 1
# + [markdown] deletable=false editable=false id="gREDVKaGOUrz" nbgrader={"cell_type": "markdown", "checksum": "1bd2e7b3005ac29a4d27641b947dd63a", "grade": false, "grade_id": "cell-235fe09bb4f2bda6", "locked": true, "schema_version": 3, "solution": false, "task": false}
# **Task 10 (`stretch goal — optional`):** Plot confusion matrix for your model using your **test data**. Does your model have higher recall or higher precision? How does your model's metrics compare to the metrics you calculated above?
# + deletable=false id="W9_4caTROUrz" nbgrader={"cell_type": "code", "checksum": "43e2179a73f1d179803ed2ec2a3c97a8", "grade": false, "grade_id": "225SC_06_a", "locked": false, "schema_version": 3, "solution": true, "task": false} colab={"base_uri": "https://localhost:8080/", "height": 405} outputId="f136fe1a-db76-48a6-85d1-4b7a94a1eebe"
'''T10. Plot ROC curve.'''
from sklearn.metrics import plot_confusion_matrix
plot_confusion_matrix(model, X_val, y_val, values_format='.0f')
# + colab={"base_uri": "https://localhost:8080/"} id="o_Ho0NnvUP4A" outputId="c22676e6-f921-42df-a818-b4070bb7a077"
# if i ever write a book about learning modeling...
# ...I'm going to title it 'The Confusion Matrix'.
my_precision = 285/(285+192)
my_recall = 285/(285+280)
print(' model precision', my_precision)
print(' model recall', my_recall)
# + [markdown] id="NODeUf0MVZO9"
# # I have higher precision than recall.
#
# ## I feel like I could have done much better on this, but I spent way too much time reading about how to reference datetime formats - hyphens not dots - and how to set up > < - turns out parenthesis are our friends, as usual.
#
# anyways, moving on to the questions...
| Stephen_Lupsha_DS23_Sprint_Challenge_22.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Създаване на речник
empty_dict = dict()
empty_dict
empty_dict = {}
empty_dict
# Достъп до стойност по ключ
numbers = {'one': 'uno', 'two': 'dos'}
numbers['one']
numbers['two']
# Добавянен на двойка ключ/стойност
code = {}
code['a'] = 1
code
# Задаване на стойност за даден ключ
numbers['one'] = 'eins'
# Извличане на всички ключове
numbers.keys()
# Извличане на всички стойности
numbers.values()
# Проверка дали даден ключ се съдържа в речник
'one' in numbers
| lectures/Dictionary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
# for cleaning
import regex as re
import nltk
nltk.download('stopwords')
nltk.download('punkt')
from wordcloud import WordCloud
# for tokenizing
from nltk.tokenize import word_tokenize
import matplotlib.pyplot as plt
import seaborn as sns
import time
# reading file form data directory
import os
# for clustering
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.cluster import KMeans
# get names of all the files to be clustered
files = os.listdir('/mnt/c/_programming/medius-intern-challenge/ClusterDocs/data')
# ignore the .DS_STORE file
files.remove(files[0])
# store the contents into a list
docsList = []
path = '/mnt/c/_programming/medius-intern-challenge/ClusterDocs/data/'
for file in files:
p = path+file
# to handle different encodings
f = open(p, encoding="ascii", errors="surrogateescape")
contents = f.read()
docsList.append(contents)
docsList[101]
# process the data by removing all punctuations and stop words
cleanDocs = []
p1 = re.compile(r'^\W+|\W+$')
p2 = re.compile(r'\s')
p3 = re.compile(r'[^a-zA-Z0-9]')
for doc in docsList:
tokens = word_tokenize(doc)
newToken = ""
for word in tokens:
if not p1.match(word) and not p2.match(word) and not p3.match(word) and word not in stopwords.words('english'):
newToken = newToken + " " + word.lower()
cleanDocs.append(newToken)
print(cleanDocs[101])
# +
long_string = ','.join(list(cleanDocs))
# Create a WordCloud object
wordcloud = WordCloud(background_color="white", max_words=5000, contour_width=3, contour_color='steelblue')
# Generate a word cloud
wordcloud.generate(long_string)
# Visualize the word cloud
wordcloud.to_image()
# -
| .ipynb_checkpoints/clusterDocs-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import time
Name = input("What is your name ?",)
print("Hello", Name , "! It is time for a hangman game!")
time.sleep(1)
print("Start guessing...")
time.sleep(0.2)
word = "<PASSWORD>"
guesses = ''
turns = 13
while turns > 0:
failed = 0
for char in word:
if char in guesses:
print(char),
else:
print('_')
failed += 1
if failed == 0:
print('You won !!!')
break
print
guess = input('Please give in a character,')
guesses += guess
if guess not in word:
turns -= 1
print('Wrong!')
print('You have', turns, 'turns')
if turns == 0:
print('YOU LOSE!!!!!!')
# + active=""
#
#
| Python scripts/.ipynb_checkpoints/Python Hangman game withoutdrawing-checkpoint-DESKTOP-MEBM21O.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 ('base')
# language: python
# name: python3
# ---
from pyfracman.data import read_ors
is_data_fname = "C:\\Users\\scott.mckean\\Desktop\\Data Exports\\InducedEvents.ors"
is_data = read_ors(is_data_fname)
| examples/point_pattern_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Ruby 2.5.1
# language: ruby
# name: ruby
# ---
# # IRuby input widget demo
#
# For comparison with ipywidgets. Run this notebook with Cell -> Run All, and note in particular that the inputs block execution until the user submits the form.
#
# Read more about IRuby widgets in [this notebook](https://github.com/SciRuby/sciruby-notebooks/blob/master/IRuby%20Examples/input.ipynb).
# ## Single input
name = IRuby.input 'Enter your name:'
puts "Hello, #{name}!"
# ## Inline form
data = IRuby.form do
input :name, label: 'Name:'
select :animal, 'cat', 'dog', 'bird', label: 'Animal:'
button
end
puts "#{data[:name]} has a pet #{data[:animal]}"
# ## Pop-up form
data = IRuby.popup do
input :name, label: 'Name:'
select :animal, 'cat', 'dog', 'bird', label: 'Animal:'
button
end
puts "#{data[:name]} has a pet #{data[:animal]}"
| ruby/iruby_widgets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="TgdzjVyyQMrN" colab_type="code" colab={}
# <NAME> Cenário 2
import pandas as pd
import numpy as np
from numpy.linalg import matrix_power
import numpy.linalg as lin
import matplotlib.pyplot as plt
# + id="X_1dAstyQMrS" colab_type="code" colab={}
def get_state_id(array, size):
return tuple(array + [0]*(size - len(array)))
# + id="sPhCNeIyQMrV" colab_type="code" colab={}
# Gera os estados adjacentes
def get_next_states(f, λ1, λ2, µ1, µ2, size):
r = {}
f = [i for i in f if i > 0]
# Chegada de um cliente do tipo 1
if λ1 > 0:
r[get_state_id(f + [1], size)] = λ1
# Chegada de um cliente do tipo 2
if λ2 > 0:
r[get_state_id(f + [2], size)] = λ2
# Saida de um cliente do tipo 1
if µ1 > 0 and len(f) > 0 and f[0] == 1:
r[get_state_id(f[1:], size)] = µ1
# Saida de um cliente do tipo 2
if µ1 > 0 and len(f) > 0 and f[0] == 2:
r[get_state_id(f[1:], size)] = µ2
return r
# + id="Hq78tv7TQMrX" colab_type="code" outputId="ceaf9ebd-6c85-4966-c1cc-73dc2ff43f47" executionInfo={"status": "ok", "timestamp": 1577119920305, "user_tz": 180, "elapsed": 993, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17736361451805105123"}} colab={"base_uri": "https://localhost:8080/", "height": 85}
print( get_next_states([], 0.1, 0.2, 0.3, 0.4, 3) )
print( get_next_states([], 0.0, 0.2, 0.3, 0.4, 3) )
print( get_next_states([], 0.1, 0.0, 0.3, 0.4, 3) )
print( get_next_states([1], 0.1, 0.2, 0.3, 0.4, 3) )
# + id="oncSls1XQMra" colab_type="code" colab={}
# Cria os estados da Cadeia de Markov
def create_ctmc_states(size, λ1, λ2, µ1, µ2):
states = {}
queue = [tuple([0]*size)]
while len(queue) > 0:
f = queue.pop(0)
n = get_next_states(f, λ1, λ2, µ1, µ2, size)
queue.extend([list(i) for i in n.keys() if len(i) <= size and i not in states])
states[tuple(f)] = {k:v for (k,v) in n.items() if len(k) <= size}
return states
# + id="J1M4I3iTQMrc" colab_type="code" outputId="a60c78a4-0926-4d9d-a839-54e3ecf4fbae" executionInfo={"status": "ok", "timestamp": 1577119920307, "user_tz": 180, "elapsed": 987, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17736361451805105123"}} colab={"base_uri": "https://localhost:8080/", "height": 272}
create_ctmc_states(3, 0.6, 0.2, 1, 0.5)
# + id="BdBfo773QMre" colab_type="code" colab={}
# Gera a matriz referente a Cadeia de Markov
def create_ctmc_matrix(size, λ1, λ2, µ1, µ2):
states = create_ctmc_states(size, λ1, λ2, µ1, µ2)
df = pd.DataFrame(states)
df = df.reindex(sorted(df.columns), axis=1)
df = df.sort_index()
df = df.transpose()
df = df[df.index]
columns = df.columns
np.fill_diagonal(df.values, -df.sum(axis=1))
df = df.fillna(0)
return df.to_numpy(), columns
# + id="BYYnlnqdQMrg" colab_type="code" outputId="378e6474-0240-447c-d745-2a4aba362fd6" executionInfo={"status": "ok", "timestamp": 1577119920895, "user_tz": 180, "elapsed": 1567, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17736361451805105123"}} colab={"base_uri": "https://localhost:8080/", "height": 286}
Q, columns = create_ctmc_matrix(3, 0.1, 0.2, 0.3, 0.4)
plt.imshow(Q)
plt.colorbar()
# + id="Agmgi2QAQMri" colab_type="code" colab={}
def get_ctmc_steady_state(Q):
sz = Q.shape[0]
Qt = Q.transpose()
Qt[sz-1,:] = 1
b = np.zeros((sz,1))
b[sz-1] = 1
x = lin.solve(Qt, b)
return x
# + id="If1tQW0IQMrk" colab_type="code" colab={}
def get_pi_N(steady_state, states):
df_steady_state = pd.DataFrame(steady_state, index=states)
get_N_from_state = lambda ix: sum(x > 0 for x in ix)
df_steady_state_N = df_steady_state.set_index(df_steady_state.index.map(get_N_from_state))
df_steady_state_N = df_steady_state_N.groupby(df_steady_state_N.index).agg(sum)
return df_steady_state_N[0].reindex(range(df_steady_state_N[0].index.max() + 1)).ffill(0).to_numpy()
# + id="zkZDbSt-QMrm" colab_type="code" outputId="6219832e-e75b-4069-e117-2a27af69edb6" executionInfo={"status": "ok", "timestamp": 1577119920896, "user_tz": 180, "elapsed": 1554, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17736361451805105123"}} colab={"base_uri": "https://localhost:8080/", "height": 272}
π = get_ctmc_steady_state(Q)
π
# + id="AqlUpfLCQMro" colab_type="code" outputId="7978dfa7-1b2f-4aae-b4a7-394846b453ad" executionInfo={"status": "ok", "timestamp": 1577119920897, "user_tz": 180, "elapsed": 1548, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17736361451805105123"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
get_pi_N(π, columns)
# + id="zxBFHePdQMrq" colab_type="code" colab={}
def calculaCadeiaMarkovCenario2(λ1, sz):
print("create_ctmc_matrix")
λ2 = 0.2
µ1 = 1
µ2 = 0.5
Q, columns = create_ctmc_matrix(sz, λ1, λ2, µ1, µ2)
print("get_ctmc_stationary_state")
π = get_ctmc_steady_state(Q)
print("get_pi_N")
πk = get_pi_N(π, columns)
return πk
# + id="nVv5ck3wQMrs" colab_type="code" outputId="f897fd3d-87a3-4f53-8038-18c7865c829c" executionInfo={"status": "ok", "timestamp": 1577119920898, "user_tz": 180, "elapsed": 1539, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17736361451805105123"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
rho = 0.6/1 + 0.2/0.5
rho
# + id="sEZbMHKHQMrt" colab_type="code" outputId="b6e65fa1-733c-4073-ff34-b12029dcd684" executionInfo={"status": "ok", "timestamp": 1577119921910, "user_tz": 180, "elapsed": 2545, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17736361451805105123"}} colab={"base_uri": "https://localhost:8080/", "height": 68}
πk = calculaCadeiaMarkovCenario2(0.6, 10)
# + id="wbXlZKXJQMrv" colab_type="code" outputId="60b06abb-8704-4cd9-b186-930bc9ffc6f4" executionInfo={"status": "ok", "timestamp": 1577119922188, "user_tz": 180, "elapsed": 2817, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17736361451805105123"}} colab={"base_uri": "https://localhost:8080/", "height": 333}
plt.plot(πk)
#plt.xlim(right=10)
r = np.array([0] + list(range(len(πk) - 1)))
print("πk =", πk)
print("r =", r)
πk.dot(r)
# + id="BEuYXdsbQMrx" colab_type="code" colab={}
def CadeiaMarkovCenario2():
Nqs = []
W = []
Nq = 0
for i in np.linspace(0.05, 0.6, 12):
print("λ1:", i)
λ1 = i
λ2 = 0.2
πk = calculaCadeiaMarkovCenario2(λ1, 10)
# Faz os cálculos dos Nq e W
r = np.array([0] + list(range(len(πk) - 1)))
Nq = πk.dot(r)
Nqs.append(Nq)
W.append(Nq/(λ1+λ2))
return Nqs, W
# + id="8TSq9GPPQMrz" colab_type="code" outputId="ca8eb771-cc4f-41b1-85db-add497de1bed" executionInfo={"status": "ok", "timestamp": 1577119934602, "user_tz": 180, "elapsed": 15223, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17736361451805105123"}} colab={"base_uri": "https://localhost:8080/", "height": 887}
Nqs, W = CadeiaMarkovCenario2()
print(Nqs)
print(W)
# + id="1FXT-_qNQMr1" colab_type="code" colab={}
| Trabalho_AD/Q3_-_Cenario_2_-_Cadeia_de_Markov_truncada.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.1
# language: julia
# name: julia-1.6
# ---
using JuMP
using GLPK
using ECOS
using MAT
using SparseArrays
using LinearAlgebra
using MathOptInterface
const MOI = MathOptInterface
using DelimitedFiles
using COSMO
using CSV
using DataFrames
using DelimitedFiles
# +
vars = matread("R4T3.mat")
S = vars["S"]
U = vars["U"]
L = vars["L"]
lambda = vars["lambda"]
n,d = size(S)
print(size(S))
print(size(L))
n,d = size(S) # n=10 d=100
d,k = size(L) # k = 5
print("done")
# +
X = zeros(size(L)[1],size(L)[2]);
n,d = size(S)
for j in 1:size(X)[2]
model = Model(ECOS.Optimizer)
@variable(model, L[i,j] <= x[i = 1:d] <= U[i,j],start=X[i,j])
@variable(model, y_abs[i = 1:n])
@constraint(model,y_abs .>= S*x)
@constraint(model,y_abs .>= -S*x)
@variable(model, x_abs[1:d])
@constraint(model, x_abs .>= x)
@constraint(model, x_abs .>= -x)
@objective(model, Min, (1/lambda)*sum(x_abs[i] for i in 1:d) + sum(y_abs[i] for i in 1:n));
set_silent(model)
optimize!(model)
x = value.(x);
X[:,j] = x;
print(j)
print(" ,")
end
# -
writedlm("output.txt", Matrix(reshape(value.(X),(d,k))), ',')
| R4-base.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import pandas as pd
import numpy as np
from sklearn.linear_model import LinearRegression
import matplotlib.pyplot as plt
import matplotlib
directory = '../execution/'#'<path-to>/results'
#filename = 'exp1002_uc3_75000_1_totallag.csv'
filename = 'exp1_uc1_10000_2_totallag.csv'#'exp1002_uc3_50000_2_totallag.csv'
warmup_sec = 60
threshold = 2000 #slope
# +
df = pd.read_csv(os.path.join(directory, filename))
input = df.iloc[::3]
#print(input)
input['sec_start'] = input.loc[0:, 'timestamp'] - input.iloc[0]['timestamp']
#print(input)
#print(input.iloc[0, 'timestamp'])
regress = input.loc[input['sec_start'] >= warmup_sec] # Warm-Up
#regress = input
#input.plot(kind='line',x='timestamp',y='value',color='red')
#plt.show()
X = regress.iloc[:, 4].values.reshape(-1, 1) # values converts it into a numpy array
Y = regress.iloc[:, 3].values.reshape(-1, 1) # -1 means that calculate the dimension of rows, but have 1 column
linear_regressor = LinearRegression() # create object for the class
linear_regressor.fit(X, Y) # perform linear regression
Y_pred = linear_regressor.predict(X) # make predictions
# -
print(linear_regressor.coef_)
# +
plt.style.use('ggplot')
plt.rcParams['axes.facecolor']='w'
plt.rcParams['axes.edgecolor']='555555'
#plt.rcParams['ytick.color']='black'
plt.rcParams['grid.color']='dddddd'
plt.rcParams['axes.spines.top']='false'
plt.rcParams['axes.spines.right']='false'
plt.rcParams['legend.frameon']='true'
plt.rcParams['legend.framealpha']='1'
plt.rcParams['legend.edgecolor']='1'
plt.rcParams['legend.borderpad']='1'
#filename = f"exp{exp_id}_{benchmark}_{dim_value}_{instances}"
t_warmup = input.loc[input['sec_start'] <= warmup_sec].iloc[:, 4].values
y_warmup = input.loc[input['sec_start'] <= warmup_sec].iloc[:, 3].values
plt.figure()
#plt.figure(figsize=(4, 3))
plt.plot(X, Y, c="#348ABD", label="observed")
#plt.plot(t_warmup, y_warmup)
plt.plot(X, Y_pred, c="#E24A33", label="trend") # color='red')
#348ABD, 7A68A6, A60628, 467821, CF4457, 188487, E24A33
plt.gca().yaxis.set_major_formatter(matplotlib.ticker.FuncFormatter(lambda x, pos: '%1.0fK' % (x * 1e-3)))
plt.ylabel('queued messages')
plt.xlabel('seconds since start')
plt.legend()
#ax.set_ylim(ymin=0)
#ax.set_xlim(xmin=0)
plt.savefig("plot.pdf", bbox_inches='tight')
| analysis/lag-trend-graph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ASSIGNMENT 3
#
# #### Machine Learning in Korea University
# #### COSE362, Fall 2018
# #### Due : 12/18 (THU) 11:59 PM
#
# #### In this assignment, you will implement and train a Recommender System. Also, you will learn how to utilize scikit-learn to analyze data using clustering.
# * Implemented using Anaconda 5.3 with python 3.7. Please use <b>python 3</b>
# * Use given dataset. Please do not change data split.
# * Use numpy, scikit-learn, and matplotlib library
# * You don't have to use all imported packages below. (some are optional). <br>
# * <b>*DO NOT MODIFY OTHER PARTS OF CODES EXCEPT "Your Code Here"*</b>
#
# ### 1. Recommender System
#
# In the problem 1, you will implement a simple recommender system.
# * See your lecture note (Lecture 15. Recommender Systems - Collaborative filtering).
# * Do not implement matrix factorization and Do not import any other packages and libraries. <b>You should use only numpy</b>.
# * Analyze train and validation error. <br>
# ### Dataset description : MovieLens Dataset
#
#
# Here are brief descriptions of the data.
#
# ><b>u.data</b> -- The full u data set, 100000 ratings by 943 users on 1682 items.
# Each user has rated at least 20 movies. Users and items are
# numbered consecutively from 1. The data is randomly
# ordered. This is a tab separated list of
# user id | item id | rating | timestamp.
# The time stamps are unix seconds since 1/1/1970 UTC
#
# ><b>u1.base</b> -- Subset of u.data. You should use u1.base at training time. <br>
# ><b>u1.test</b> -- Subset of u.data. You should use u1.test at testing.
# +
import numpy as np
import matplotlib.pyplot as plt
np.random.seed(1000)
# +
with open('./data/u.info') as f:
info = f.readlines()
num_users = int(info[0].split()[0])
num_movies = int(info[1].split()[0])
def read_data(file_name):
with open('./data/' + file_name) as f:
dataset = []
cnt = 0
while True:
line = f.readline()
if not line: break
d = list(map(int, line.split()[:3]))
d[0] -= 1
d[1] -= 1
dataset.append(d)
return dataset
###########################################
train_dataset = read_data('u1.base')
valid_dataset = read_data('u1.test')
# -
# ### 1 - (1) Design your recommender system model.
#
# ##### Instructions <br>
# * <b>\__init\__</b> : configure model. <b>DO NOT MODIFY</b>.
# * <b>compute_loss</b> : compute loss between user i and movie j.
# * <b>update</b> : update the parameter of user i and movie j with gradient descent.
# * <b>run_epoch</b> : train one iteration of collaborative filtering. <b>If "trainable=False", this function doesn't update parameters.</b> <br>
# <t>The variable "trainable" is false when the model is on validation or test.
#
#
# <br>* <b>loss</b> --> see your lecture note.
# <br>* <b>rmse</b> indicates Root Mean Square Error that is widely used for rating prediction evaluation.
# +
class RecommenderSystem():
def __init__(self, num_users, num_movies, user_size, movie_size, learning_rate, reg_coef):
self.user_mat = np.random.normal(0, 1, (num_users, user_size))
self.movie_mat = np.random.normal(0, 1, (num_movies, movie_size))
self.learning_rate = learning_rate
self.reg_coef = reg_coef
self.loss = 0.0
def compute_loss(self, i, j, rating):
# Your Code Here
# End Your Code
return target, loss
def update(self, target, i, j, rating):
# Your Code Here
# End Your Code
##
def run_epoch(self, dataset, trainable=False):
loss_sum = rmse_sum = target_sum = 0
np.random.shuffle(dataset)
for s_idx, sample in enumerate(dataset):
# Your Code Here
# End Your Code
return loss_sum / len(dataset), rmse
# +
def main(config):
#####
# optimal : (int) the epoch where validation loss is minimum
# eps : (list) a list of training epochs
# loss_tr : (list) a list of training losses
# loss_va : (list) a list of validation losses
# rmse_tr : (list) a list of training rmse(root mean square error)
# rmse_va : (list) a list of validation rmse(root mean square error)
model = RecommenderSystem(num_users, num_movies,
config['user_size'],
config['movie_size'],
config['learning_rate'],
config['reg_coef'])
min_loss = optimal = 99999
eps, loss_tr, loss_va, rmse_tr, rmse_va = [], [], [], [], []
for epoch in range(config['max_epoch']):
# ls_tr : mean of total losses in an epoch
# e_tr : mean of total root mean square errors in an epoch
ls_tr, e_tr = model.run_epoch(train_dataset, trainable=True)
ls_va, e_va = model.run_epoch(valid_dataset, trainable=False)
# Your Code Here
# End Your Code
return optimal, eps, loss_tr, loss_va, rmse_tr, rmse_va, model
###################################################################
config = {'user_size': 10,
'movie_size': 10,
'learning_rate': 0.01,
'reg_coef': 0.001,
'max_epoch': 50,
'eval_step': 5}
optimal, eps, loss_tr, loss_va, rmse_tr, rmse_va, model = main(config)
print ("\[Exp have been finished !]\nOptimal : {}, Train loss : {:2.3f}, Valid loss : {:2.3f}, RMSE : {:3.2f}"
.format(optimal, loss_tr[optimal], loss_va[optimal], rmse_va[optimal]))
# -
# ### 1 - (2) Plot the training and validation loss against epochs and Analyze.
#
#
# Plot your train error and validation error by number of iterations.
plt.plot(eps, loss_tr, eps, loss_va, 'r-')
plt.title("error graph")
plt.legend(["Training", "Validation"])
plt.show()
# ##### Analyze the result.
# +
# Write description here
# -
# ## 2. Clustering
#
# In the problem 2, you would learn how to analyze data with unsupervised learning algorithm.
# * Implement <b>k-means clustering</b> algorithm using <b>scikit-learn packages</b>. <br>
# * Visualize your result and analyze. <br>
# * Implement <b>PCA(principle component analysis)</b> and visualize your data onto 2-dimensional domain, and visualize your data by class-labels.
# +
from sklearn.cluster import KMeans
from sklearn.datasets import make_blobs
from sklearn import cluster
from sklearn.datasets import load_digits
from sklearn.decomposition import PCA
from sklearn.preprocessing import scale
n_samples = 3000
random_state = 1182
X, y = make_blobs(n_samples=n_samples, random_state=random_state)
# -
# ### 2 - (1) Implement K-means clustering algorithm and visualize data with class labels.
# For given data, find the best number of clusters (each cluster is well-divided). <br>
# Visualize your results using <b>scatter plot</b>.
# +
# Use sklearn.cluster.KMeans for k-means clustering
# Use plt.scatter for visualization
# Your Code Here
# End Your Code
# -
# ### 2 - (2) Implement PCA and visualize data with class labels.
# Conduct K-means clustering on given data. <br>
# Implement <b>PCA(principle component analysis)</b> to convert high-dimensional vectors into 2-dimensional vectors. <br>
# Compare plots by K-means result and class labels by visualization. <br>
# * Dataset : Handwritten digit dataset (Class : digit, Data : digit image)
# * Visualize <b>two scatter plots</b>. (One for class label and one for k-means clustering)
# +
# use sklearn.decomposition.PCA
digits = load_digits()
data = scale(digits.data)
labels = digits.target
# Your Code Here
# End Your Code
| HW3/HW3_20XXXXXXXX_name.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: my_py
# language: python
# name: my_py
# ---
# +
import pandas as pd
import os
import datetime
import pickle
import re
import time
from collections import Counter
import numpy as np
import nltk
nltk.data.path
# -
from nltk.tokenize import sent_tokenize, word_tokenize
#nltk.download('stopwords')
from nltk.corpus import stopwords
import spacy
nlp = spacy.load('en_core_web_sm', disable=['parser', 'ner'])
from ast import literal_eval
from collections import Counter
# ## 1. Import Regulatory Sections and Noun Chunks with Areas
# Noun chunks with areas
nounchunks_area=pd.read_csv('/home/ec2-user/SageMaker/New Uncertainty/DictionaryOfRegulatoryNounChunks.csv')
print(nounchunks_area.info())
nounchunks_area.head()
# Convert to dictionary
nounchunks_area=nounchunks_area[nounchunks_area['area_no']>0].set_index('noun_chunks')
nounchunks_area_dict=nounchunks_area.to_dict()['area']
print(len(nounchunks_area_dict))
# Expanded reg sentences with matched noun chunks
df_regSentsExpand=pd.read_pickle('/home/ec2-user/SageMaker/New Uncertainty/Reg Relevance/RegSentsExpand_NounChunks3.pkl')
print(df_regSentsExpand.info())
# Remove duplicated articles
IDs_nodup=pd.read_csv('/home/ec2-user/SageMaker/New Uncertainty/IDs_no_duplicates.csv')
print(IDs_nodup.info())
df_regSentsExpand['ID']=df_regSentsExpand['ID'].astype('int64')
df_regSentsExpand=IDs_nodup.merge(df_regSentsExpand,on='ID',how='left').reset_index(drop=True)
print(df_regSentsExpand.info())
# Refine to reg relevant articles
df_regSentsExpandRelevant=df_regSentsExpand[df_regSentsExpand['NounChunkMatchFiltered']>0].reset_index(drop=True)
print(df_regSentsExpandRelevant.info())
# ## 2. Link Expanded Reg Sentences to Areas
# ### Approch 1 - dominant noun chunk area (dnca): use all unique areas from dominant noun chunks
df_regSentsExpandRelevant['DominantNounChunk']=''
df_regSentsExpandRelevant['DominantNounChunkArea']=''
for i in range(0, len(df_regSentsExpandRelevant)):
nc_list=df_regSentsExpandRelevant['NounChunkMatchWordsFiltered'][i]
dominant_nc=[j for j in Counter(nc_list).keys() if nc_list.count(j)==max(Counter(nc_list).values())]
dominant_nc_area=set()
for nc in dominant_nc:
if nc in nounchunks_area_dict:
area=literal_eval(nounchunks_area_dict[nc])
dominant_nc_area.update(area)
df_regSentsExpandRelevant['DominantNounChunk'][i]=dominant_nc
df_regSentsExpandRelevant['DominantNounChunkArea'][i]=dominant_nc_area
print(df_regSentsExpandRelevant.info())
# ### Approach 2 - dominant area (da): use the dominant areas from all noun chunks
# +
df_regSentsExpandRelevant['AllAreas']=''
for i in range(0, len(df_regSentsExpandRelevant)):
nounchunks=df_regSentsExpandRelevant['NounChunkMatchWordsFiltered'][i]
area_list=[]
for nc in nounchunks:
if nc in nounchunks_area_dict:
area=sorted(literal_eval(nounchunks_area_dict[nc]))
area_list=area_list+area
df_regSentsExpandRelevant['AllAreas'][i]=area_list
print(df_regSentsExpandRelevant.info())
# -
df_regSentsExpandRelevant['AreaCount']=''
df_regSentsExpandRelevant['DominantArea']=''
for i in range(0, len(df_regSentsExpandRelevant)):
area_list=df_regSentsExpandRelevant['AllAreas'][i]
area_count=Counter(area_list).most_common()
dominant_area=[j for j in Counter(area_list).keys() if area_list.count(j)==max(Counter(area_list).values())]
df_regSentsExpandRelevant['AreaCount'][i]=area_count
df_regSentsExpandRelevant['DominantArea'][i]=dominant_area
print(df_regSentsExpandRelevant.info())
# ### Approach 3 - unique distinct area (uda): use all unique areas from area-specific noun chunks
df_regSentsExpandRelevant['AllDistinctAreas']=''
df_regSentsExpandRelevant['UniqueDistinctAreas']=''
for i in range(0, len(df_regSentsExpandRelevant)):
nounchunks=df_regSentsExpandRelevant['NounChunkMatchWordsFiltered'][i]
area_list=[]
area_set=set()
for nc in nounchunks:
if nc in nounchunks_area_dict:
area=sorted(literal_eval(nounchunks_area_dict[nc]))
if len(area)==1:
area_list=area_list+area
area_set=set(area_list)
df_regSentsExpandRelevant['AllDistinctAreas'][i]=area_list
df_regSentsExpandRelevant['UniqueDistinctAreas'][i]=area_set
print(df_regSentsExpandRelevant.info())
# ### Approach 4: dominant distinct area (dda): use the dominant areas from area-specific noun chunks
df_regSentsExpandRelevant['DistinctAreaCount']=''
df_regSentsExpandRelevant['DominantDistinctArea']=''
for i in range(0, len(df_regSentsExpandRelevant)):
area_list=df_regSentsExpandRelevant['AllDistinctAreas'][i]
area_count=Counter(area_list).most_common()
dominant_area=[j for j in Counter(area_list).keys() if area_list.count(j)==max(Counter(area_list).values())]
df_regSentsExpandRelevant['DistinctAreaCount'][i]=area_count
df_regSentsExpandRelevant['DominantDistinctArea'][i]=dominant_area
print(df_regSentsExpandRelevant.info())
print(df_regSentsExpandRelevant.head())
df_regSentsExpandRelevant.to_pickle('/home/ec2-user/SageMaker/New Uncertainty/Categorical Index/RegSentsExpand_NounChunks_Area.pkl')
# ## 3. Filtered Noun Chunk Occurences by Area
# Reg relevant articles
df_regSentsExpandRelevant=pd.read_pickle('/home/ec2-user/SageMaker/New Uncertainty/Categorical Index/RegSentsExpand_NounChunks_Area.pkl')
print(df_regSentsExpandRelevant.info())
# Filtered noun chunk occurences across regulation-related articles
df_nounchunk_occurences=pd.read_csv('/home/ec2-user/SageMaker/New Uncertainty/Reg Relevance/RegSentsExpand_FilteredNounChunkOccurences.csv')
print(df_nounchunk_occurences.info())
# Dummies by dominant distinct area (dda)
for i in range(1,15):
var='DominantDistinctArea'+str(i)
df_regSentsExpandRelevant[var]=0
for j in range(0, len(df_regSentsExpandRelevant)):
if i in df_regSentsExpandRelevant['DominantDistinctArea'][j]:
df_regSentsExpandRelevant[var][j]=1
# Filtered noun chunks across regulation-related articles by area
for i in range(1,15):
allMatchWords=[]
for list in df_regSentsExpandRelevant[df_regSentsExpandRelevant['DominantDistinctArea'+str(i)]==1]['NounChunkMatchWordsFiltered']:
allMatchWords=allMatchWords+list
allMatchWordsCount=Counter(allMatchWords)
var_name='Occurences_dda'+str(i)
df_MatchWords = pd.DataFrame(allMatchWordsCount.items(),columns = ['Noun Chunks',var_name])
df_nounchunk_occurences=df_nounchunk_occurences.merge(df_MatchWords,on='Noun Chunks',how='outer')
print(df_nounchunk_occurences)
# +
#df_nounchunk_occurences.to_csv('/home/ec2-user/SageMaker/New Uncertainty/Categorical Index/RegArea_FilteredNounChunkOccurences.csv',index=False)
# +
# df_nounchunk_occurences=pd.read_csv('/home/ec2-user/SageMaker/New Uncertainty/Categorical Index/RegArea_FilteredNounChunkOccurences.csv')
# print(df_nounchunk_occurences.info())
# +
# # Combine Areas 8 & 9
# df_regSentsExpandRelevant['DominantDistinctArea8_9']=0
# df_regSentsExpandRelevant.loc[(df_regSentsExpandRelevant['DominantDistinctArea8']==1) | (df_regSentsExpandRelevant['DominantDistinctArea9']==1),
# 'DominantDistinctArea8_9']=1
# print(df_regSentsExpandRelevant[['DominantDistinctArea8_9']])
# +
# # Filtered noun chunks across regulation-related articles for area8_9
# allMatchWords=[]
# for list in df_regSentsExpandRelevant[df_regSentsExpandRelevant['DominantDistinctArea8_9']==1]['NounChunkMatchWordsFiltered']:
# allMatchWords=allMatchWords+list
# allMatchWordsCount=Counter(allMatchWords)
# var_name='Occurences_dda8_9'
# df_MatchWords = pd.DataFrame(allMatchWordsCount.items(),columns = ['Noun Chunks',var_name])
# df_nounchunk_occurences=df_nounchunk_occurences.merge(df_MatchWords,on='Noun Chunks',how='outer')
# print(df_nounchunk_occurences)
# -
df_nounchunk_occurences.to_csv('/home/ec2-user/SageMaker/New Uncertainty/Categorical Index/RegArea_FilteredNounChunkOccurences.csv',index=False)
# ## A1. Examine the Results
# Investigate the results
df=pd.read_pickle('/home/ec2-user/SageMaker/New Uncertainty/Categorical Index/RegSentsExpand_NounChunks_Area.pkl')
print(df.info())
for i in range(50000,55000):
if len(df['AllDistinctAreas'][i])>1 and len(df['DominantDistinctArea'][i])==1:
print(df['ID'][i], df['RegSentsExpand'][i],df['NounChunkMatchWordsFiltered'][i],df['AllDistinctAreas'][i], df['DominantDistinctArea'][i])
print('\n')
print(df[df['ID']=='294286922']['NounChunkMatchWordsFiltered'].values)
print(df[df['ID']=='294286922'][['AllAreas','DominantArea']])
for i in range(200000,205000):
if len(df['AllDistinctAreas'][i])>1 and len(df['DominantDistinctArea'][i])==1 and (len(df['RegSentsExpand'][i])<600):
print(df['ID'][i], df['RegSentsExpand'][i],df['NounChunkMatchWordsFiltered'][i],df['AllDistinctAreas'][i], df['DominantDistinctArea'][i],'\n')
print(df[df['ID']=='282489466']['NounChunkMatchWordsFiltered'].values)
| Code/9_Categorize_Articles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Data visualisation respirometer
# %matplotlib inline
import os
import sys
import numpy as np
import matplotlib.pyplot as plt
import matplotlib as mpl
import seaborn as sns
from matplotlib.ticker import LinearLocator, MaxNLocator
sns.set_style('whitegrid')
mpl.rcParams['font.size'] = 16
mpl.rcParams['axes.labelsize'] = 18
mpl.rcParams['xtick.labelsize'] = 16
mpl.rcParams['ytick.labelsize'] = 16
# ## Example data Cierkens
observations = pd.read_csv("./bayesian_story/respirometer_data.txt", sep="\t", index_col=0,
names=["DO", "OURex"], skiprows=2)
observations.index = observations.index/(60*24) #everything to days
observations.index.name='Time (min)'
observations.head()
# +
fig, axs = plt.subplots(2, 1, figsize=(12, 8), sharex=True)
fig.subplots_adjust(hspace=0.1)
axs[0].plot(observations.index*24*60, observations['DO'],'-', color ='0.1')
axs[0].set_yticks([6., 7., 8., 9.])
axs[0].set_ylabel('$\mathrm{S}_{O}$ ($mg\ l^{-1}$)')
axs[1].plot(observations.index*24*60, observations['OURex'],'-', color ='0.3')
axs[1].set_yticks([-0.2, 0., .2, .4, .6])
axs[1].set_xticks([0, 10, 20, 30, 40])
axs[1].set_ylabel('$\mathrm{OUR}_{\mathrm{ex}}$ ($mg\ l^{-1} min^{-1}$)')
axs[1].set_xlabel('Time (min)')
labelx = -0.05
for ax in axs:
#ax.yaxis.set_major_locator(MaxNLocator(5))
ax.yaxis.set_label_coords(labelx, 0.5)
#remove spines
#ax.spines['bottom'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
# set grid
ax.grid(which='major', axis='both', color='0.7',
linestyle='--', linewidth=0.8)
plt.savefig('respiro_data1.png', dpi=300)
plt.savefig('respiro_data1.pdf', dpi=300)
# -
# ## Example DATA DECUBBER
cubber_ref = pd.read_csv("./thesis_decubber/0508Aall_pieken.csv")[['NewTime','DO', 'OUR filt']]
cubber_ref = cubber_ref.set_index('NewTime')
cubber_ref.head()
ac_dosing = np.array([17.25, 34.5, 8.625, 4.3125, 69])
ac_moments = np.array([0., 1540, 3650, 5510, 7775])
shots = pd.DataFrame({'dosing': ac_dosing, 'moment': ac_moments})
shots = shots.set_index('moment')
ac_dosing*2
# +
import matplotlib.gridspec as gridspec
#fig, axs = plt.subplots(2, 1, figsize=(12, 8), sharex=True)
fig = plt.figure(figsize=(12, 8))
gs = gridspec.GridSpec(2, 1, height_ratios=[1, 3], hspace=0.05)
ax2 = plt.subplot(gs[0])
ax2.vlines(ac_moments/60., [0.]*5, ac_dosing, linewidth=4, colors='0.4')
ax2.set_xticks([0., 40, 80, 120, 160])
ax2.set_yticks([30, 60])
ax2.set_ylabel('$\mathrm{S}_{A}$ ($mg\ l^{-1}$)')
ax1 = plt.subplot(gs[1], sharex=ax2)
fig.subplots_adjust(hspace=0.1)
ax1.plot(cubber_ref.index/60., cubber_ref['DO'],'-', color ='0.1')
ax1.set_yticks([6., 7., 8., 9.])
ax1.set_ylabel('$\mathrm{S}_{O}$ ($mg\ l^{-1}$)')
ax1.set_xlim([-5, 180])
ax1.set_xlabel('Time (min)')
#ax1.set_xlim([-50., 11000])
#axs[1].plot(cubber_ref.index/60, cubber_ref['OUR filt'],'-', color ='0.3')
#axs[1].set_yticks([-3, 0., 3])
#axs[1].set_xticks([0, 40, 80, 120, 160])
#axs[1].set_ylabel('$\mathrm{OUR}_{\mathrm{ex}}$ ($mg\ l^{-1} min^{-1}$)')
#axs[1].set_xlabel('Time (min)')
labelx = -0.05
for ax in [ax1, ax2]:
#ax.yaxis.set_major_locator(MaxNLocator(5))
ax.yaxis.set_label_coords(labelx, 0.5)
#remove spines
#ax.spines['bottom'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
# set grid
ax.grid(which='major', axis='both', color='0.7',
linestyle='--', linewidth=0.8)
plt.setp(ax2.get_xticklabels(), visible=False)
plt.savefig('respiro_data2.png', dpi=300)
plt.savefig('respiro_data2.pdf', dpi=300)
# + active=""
# 0508A
# 1.75187374811
# 3.84249794621
# -
| respiro_datashowcase.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:75% !important; }</style>"))
import os
import numpy as np
import torch
import time
import pandas as pd
from carle.env import CARLE
from carle.mcl import RND2D, AE2D, SpeedDetector, PufferDetector, CornerBonus
from game_of_carle.agents.grnn import ConvGRNN
from game_of_carle.agents.carla import CARLA
from game_of_carle.agents.harli import HARLI
from game_of_carle.algos.cma import CMAPopulation
import bokeh
import bokeh.io as bio
from bokeh.io import output_notebook, show
from bokeh.plotting import figure
from bokeh.layouts import column, row
from bokeh.models import TextInput, Button, Paragraph
from bokeh.models import ColumnDataSource
from bokeh.events import DoubleTap, Tap
import matplotlib
import matplotlib.pyplot as plt
matplotlib.rc("font", size=28)
cmap_name = "magma"
my_cmap = plt.get_cmap(cmap_name)
output_notebook()
# +
# manual designs
life_glider = torch.zeros(1,1, 64, 64)
life_glider[:,:,33,33] = 1.0
life_glider[:,:,34,32:34] = 1.0
life_glider[:,:,35,32:35:2] = 1.0
morley_puffer = torch.zeros(1, 1, 64, 64)
morley_puffer[:,:, 33, 35] = 1.0
morley_puffer[:,:, 34, 33:37] = 1.0
morley_puffer[:,:, 35, 32] = 1.0
morley_puffer[:,:, 35, 35] = 1.0
morley_puffer[:,:, 35, 37] = 1.0
morley_puffer[:,:, 36, 32] = 1.0
morley_puffer[:,:, 36, 35] = 1.0
morley_puffer[:,:, 36, 37] = 1.0
morley_puffer[:,:, 37, 33:37] = 1.0
morley_puffer[:,:, 38, 35] = 1.0
morley_glider = torch.zeros(1,1, 64, 64)
morley_glider[:, :, 42, 47:50] = 1.0
morley_glider[:, :, 43, 48:50] = 1.0
morley_glider[:, :, 44, 49:50] = 1.0
morley_glider[:, :, 42, 55:58] = 1.0
morley_glider[:, :, 43, 56:58] = 1.0
morley_glider[:, :, 44, 57:58] = 1.0
seed_pattern = torch.zeros(1,1, 64,64)
seed_pattern[:,:, 32, 30:33] = 1.0
seed_pattern[:,:, 33:35, 32] = 1.0
seed_pattern = torch.zeros(1,1, 64,64)
seed_pattern[:,:, 0, 4:7] = 1.0
seed_pattern[:,:, 1, 6] = 1.0
seed_pattern[:,:, 2, 4:7] = 1.0
seed_pattern[:,:, 9, 1:4] = 1.0
seed_pattern[:,:, 10, 3] = 1.0
seed_pattern[:,:, 11, 1:4] = 1.0
plt.figure(figsize=(16,16))
plt.subplot(221)
plt.imshow(life_glider.squeeze()[30:46,30:46].numpy(), cmap=cmap_name)
plt.title("Life Glider", fontweight="bold")
plt.subplot(222)
plt.imshow(morley_puffer.squeeze()[30:46,30:46].numpy(), cmap=cmap_name)
plt.title("Common Morley Puffer", fontweight="bold")
plt.subplot(223)
plt.imshow(seed_pattern.squeeze()[0:16, 0:16].numpy(), cmap=cmap_name)
plt.title("Life Without Death Seed", fontweight="bold")
plt.subplot(224)
plt.imshow(morley_glider.squeeze()[35:51,45:61].numpy(), cmap=cmap_name)
plt.title("Morley Gliders", fontweight="bold")
plt.tight_layout()
plt.show()
# +
# Glider (speed) Reward
env = CARLE(device="cpu", instances=1, height=128, width=128)
env.rules_from_string("B3/S23")
env = SpeedDetector(env)
obs = env.reset()
rewards = [0]
action = life_glider
my_image = obs + env.inner_env.action_padding(1.0*action)
obs, reward, done, info = env.step(action)
for step in range(1,236):
if (step % 17 == 0):
my_image += obs.numpy() * step
rewards.append(reward.item())
obs, reward, done, info = env.step(action*0)
my_image += obs.numpy() * step
rewards.append(reward.item())
fig, ax = plt.subplots(2, 1, figsize=(14,32))
rect = plt.Rectangle(((env.height-64) // 2, (env.width-64) // 2), 64, 64,
facecolor="blue", alpha=0.3)
ax[0].add_patch(rect)
disp_image = 1.0 * my_image.squeeze()
disp_image[disp_image > 0 ] += my_image.max()/2
#plt.subplot(121)
ax[0].imshow(disp_image, cmap=cmap_name)
ax[0].add_patch(rect)
ax[0].set_title("Game of Life Glider Progression", fontsize=32, fontweight="bold")
for ii in range(len(rewards)-1):
ax[1].plot([ii, ii+1], rewards[ii:ii+2], '-',ms=10,lw=6, \
color=my_cmap((len(rewards)/2 + ii/2) / len(rewards)))
ax[1].set_title("Game of Life SpeedDetector Bonus", fontsize=32, fontweight="bold")
plt.ylabel("reward",fontweight="bold")
plt.xlabel("step", fontweight="bold")
plt.show()
# +
# Corner Bonus Reward
env = CARLE(device="cpu", instances=1, height=196, width=196)
env.rules_from_string("B3/S012345678")
env = CornerBonus(env)
obs = env.reset()
my_image = (obs + env.inner_env.action_padding(1.0*action)).numpy()
rewards = [0]
action = seed_pattern
obs, reward, done, info = env.step(action)
for step in range(1,512):
if (step % 17 == 0):
my_image[my_image == 0] = (obs.numpy() * step)[my_image == 0]
rewards.append(reward.item())
obs, reward, done, info = env.step(action*0)
my_image[my_image == 0] = (obs.numpy() * step)[my_image == 0]
rewards.append(reward.item())
fig, ax = plt.subplots(2, 1, figsize=(14,32))
action_rect = plt.Rectangle(((env.height-64) // 2, (env.width-64) // 2), 64, 64,
facecolor="blue", alpha=0.3)
punish_rect_0 = plt.Rectangle(((env.width-64),0), 64, 64,
facecolor="red", alpha=0.3)
punish_rect_1 = plt.Rectangle(((env.width-64),(env.width-64)), 64, 64,
facecolor="red", alpha=0.3)
inferno_cmap = plt.get_cmap("inferno")
reward_rect_0 = plt.Rectangle((0,0), 16, 16,
facecolor=inferno_cmap(64), alpha=0.3)
disp_image = 1.0 * my_image.squeeze()
disp_image[disp_image > 0 ] += my_image.max()/2
#plt.subplot(121)
ax[0].imshow(disp_image, cmap=cmap_name)
ax[0].add_patch(action_rect)
ax[0].add_patch(punish_rect_0)
ax[0].add_patch(punish_rect_1)
ax[0].add_patch(reward_rect_0)
for jj in range(14,96):
reward_rect = plt.Rectangle((jj,jj), 4, 4, facecolor=inferno_cmap(64), alpha=0.3/2)
ax[0].add_patch(reward_rect)
ax[0].add_patch(reward_rect_0)
ax[0].set_title("Life Without Death Growth Pattern", fontsize=28, fontweight="bold")
for ii in range(len(rewards)-1):
ax[1].plot([ii, ii+1], rewards[ii:ii+2], 'o-',ms=10,lw=6, \
color=my_cmap((len(rewards)/2 + ii/2) / len(rewards)))
ax[1].set_title("Life Without Death CornerBonus Bonus", fontweight="bold")
plt.ylabel("reward", fontweight="bold")
plt.xlabel("step", fontweight="bold")
plt.show()
# +
# "Puffer Detection" growth Bonus Reward
env = CARLE(device="cpu", instances=1, height=128, width=128)
env.rules_from_string("B368/S245")
env = PufferDetector(env)
obs = env.reset()
my_image = (obs + env.inner_env.action_padding(1.0*action)).numpy()
rewards = [0]
action = morley_puffer
obs, reward, done, info = env.step(action)
for step in range(1, 601):
if (step == 300):
my_image2 = (obs.numpy() * step)
rewards.append(reward.item())
obs, reward, done, info = env.step(action*0)
my_image3 = (obs.numpy() * step)
rewards.append(reward.item())
fig, ax = plt.subplots(1, 2, figsize=(16, 8))
fig.suptitle("Common Morley Puffer Progression", fontsize=32, fontweight="bold")
steps = [0, 300, 600]
for hh, img in enumerate([my_image, my_image2]):
disp_image = 1.0 * img.squeeze()/2
disp_image[disp_image > 0 ] += my_image3.max()/2
#plt.subplot(1,3,hh+1)
reward_rect_0 = plt.Rectangle((0,0), 16, 16,
facecolor="blue", alpha=0.3)
ax[hh].set_xticklabels("")
ax[hh].set_yticklabels("")
action_rect = plt.Rectangle(((env.height-64) // 2, (env.width-64) // 2), 64, 64,
facecolor="blue", alpha=0.5)
ax[hh].add_patch(action_rect)
ax[hh].imshow(disp_image, cmap=cmap_name, vmin = 0, vmax = my_image3.max())
ax[hh].text(0.365*128, 120, f"Step {steps[hh]}",
color=[1,1,1] ) #my_cmap((max_steps/2 + snapshot_steps[kk-1]/2)/max_steps))
plt.show()
disp_image = 1.0 * my_image3.squeeze()/2
disp_image[disp_image > 0 ] += my_image3.max()/2
fig, ax = plt.subplots(2, 1, figsize=(16,32))
action_rect = plt.Rectangle(((env.height-64) // 2, (env.width-64) // 2), 64, 64,
facecolor="blue", alpha=0.5)
punish_rect_0 = plt.Rectangle(((env.width-64),0), 64, 64,
facecolor="red", alpha=0.3)
punish_rect_1 = plt.Rectangle(((env.width-64),(env.width-64)), 64, 64,
facecolor="red", alpha=0.3)
reward_rect_0 = plt.Rectangle((0,0), 16, 16,
facecolor="blue", alpha=0.3)
#plt.subplot(121)
ax[0].imshow(disp_image, cmap=cmap_name)
ax[0].add_patch(action_rect)
ax[0].text(0.4*128, 120, f"Step {steps[-1]}",
color=[1,1,1] ) #my_cmap((max_steps/2 + snapshot_steps[kk-1]/2)/max_steps))
#ax[0].set_title("Common Morley Puffer Progression", fontsize=32, fontweight="bold")
for ii in range(len(rewards)-1):
ax[1].plot([ii, ii+1], rewards[ii:ii+2], 'o-',ms=10,\
lw=6, color=my_cmap((len(rewards)/2 + ii/2) / len(rewards)))
ax[1].set_title("PufferDetector Bonus", fontweight="bold")
plt.ylabel("reward")
plt.xlabel("step")
plt.show()
# +
# Autoencoder Exploration Bonus Reward
#random seeds
np.random.seed(42)
torch.random.manual_seed(42)
env = CARLE(device="cpu", instances=1, height=128, width=128)
env.rules_from_string("B368/S245")
env = AE2D(env)
obs = env.reset()
my_image = (obs + env.inner_env.action_padding(1.0*action)).numpy()
rewards = [0]
action = morley_puffer # + morley_glider
obs, reward, done, info = env.step(action)
count = 0
max_steps = 2001
snapshot_steps = []
for step in range(1, max_steps):
if step % ((max_steps) // 4) == 0 or step == max_steps-1:
snapshot_steps.append(step)
my_image[:,:, count*32:count*32+32, :] = step * obs.numpy()[:,:,48:80,:]
count += 1
rewards.append(reward.item())
obs, reward, done, info = env.step(action*0)
rewards.append(reward.item())
fig, ax = plt.subplots(2, 1, figsize=(14,32))
ax[0].imshow(my_image.squeeze(), cmap=cmap_name)
for jj in range(4):
time_rect = plt.Rectangle((0, jj*32), 128, 64, \
facecolor=my_cmap(32+32*jj), alpha=0.2)
ax[0].add_patch(time_rect)
ax[0].set_title("Common Morley Puffer Progression", fontsize=32, fontweight="bold")
for ii in range(len(rewards)-1):
ax[1].plot([ii, ii+1], rewards[ii:ii+2], 'o-',ms=10,lw=6, color=my_cmap((ii+1.)/len(rewards)))
ax[1].set_title("Autoencoder Loss Exploration Bonus", fontsize=32, fontweight="bold")
plt.ylabel("reward", fontweight="bold")
plt.xlabel("step", fontweight="bold")
plt.show()
rewards_ae = rewards
# +
# random network distillation
#random seeds
np.random.seed(42)
torch.random.manual_seed(42)
env = CARLE(device="cpu", instances=1, height=128, width=128)
env.rules_from_string("B368/S245")
env = RND2D(env)
obs = env.reset()
my_image = (obs + env.inner_env.action_padding(1.0*action)).numpy()
rewards = [0]
action = morley_puffer #+ morley_glider
obs, reward, done, info = env.step(action)
count = 0
snapshot_steps = []
for step in range(1, max_steps):
if step % ((max_steps) // 4) == 0 or step == max_steps-1:
print(step)
snapshot_steps.append(step)
my_image[:,:, count*32:count*32+32, :] = step * obs.numpy()[:,:,48:80,:]
count += 1
rewards.append(reward.item())
obs, reward, done, info = env.step(action*0)
rewards.append(reward.item())
fig, ax = plt.subplots(2, 1, figsize=(14,28))
#plt.subplot(121)
ax[0].imshow(my_image.squeeze(), cmap=cmap_name)
for jj in range(4):
time_rect = plt.Rectangle((0, jj*32), 128, 64, \
facecolor=my_cmap(32+32*jj), alpha=0.2)
ax[0].add_patch(time_rect)
ax[0].set_title("Common Morley Puffer Progression", fontsize=32, fontweight="bold")
for ii in range(len(rewards)-1):
ax[1].plot([ii, ii+1], rewards[ii:ii+2], 'o-',ms=10,lw=6, color=my_cmap((ii+1.)/len(rewards)))
ax[1].set_title("Random Network Distillation Exploration Bonus", fontsize=32, fontweight="bold")
plt.ylabel("reward", fontweight="bold")
plt.xlabel("step", fontweight="bold")
plt.show()
rewards_rnd = rewards
# +
disp_image = 1.0 * my_image.squeeze()
disp_image[disp_image > 0 ] += my_image.max()/2
disp_image[0,:] = disp_image.max()
disp_image[-1, :] = disp_image.max()
disp_image[:,0] = disp_image.max()
disp_image[:,-1] = disp_image.max()
for jj in range(5):
disp_image[min(jj*32,127),:] = disp_image.max()
# +
fig, ax = plt.subplots(2, 1, figsize=(14,28))
ax[0].imshow(disp_image, cmap=cmap_name)
ax[0].set_title("Common Morley Puffer Progression", fontsize=32, fontweight="bold")
ax2 = ax[1].twinx()
ii = 0
ax[1].plot([ii, ii+1], rewards_rnd[ii:ii+2], '-', \
lw=10, color=my_cmap((max_steps//2 + ii/2) / max_steps), label="RND", alpha=0.6)
ax2.plot([ii, ii+1], rewards_ae[ii:ii+2], ':', \
lw=10, color=my_cmap((max_steps//2 + ii/2) / max_steps), label="AE")
for ii in range(1, len(rewards_ae)-1):
ax[1].plot([ii, ii+1], rewards_rnd[ii:ii+2], '-', \
lw=10, color=my_cmap((max_steps//2 + ii/2) / max_steps), alpha=0.6)
ax2.plot([ii, ii+1], rewards_ae[ii:ii+2], ':', \
lw=10, color=my_cmap((max_steps//2 + ii /2) / max_steps))
ax[1].set_title("Random Network Distillation/Autoencoder Bonus", fontsize=32, fontweight="bold")
ax[1].set_ylabel("random network distillation reward", fontweight="bold")
#ax[1].set_xlabel("step", fontweight="bold")
ax2.set_ylabel("autoecndoer loss reward", fontweight="bold")
ax2.set_xlabel("step", fontweight="bold")
#ax[0].set_xticklabels("")
#ax[0].set_yticklabels("")
ax2.legend(loc=(0.8, .17))
ax[1].legend(loc=(0.775, 0.015))
ax2.axis([-30, 2011, -0.03, 0.31])
ax[1].axis([-30, 2011, -0.0003, 0.0036])
for kk in range(1,5):
ax[0].text(0.385*128, (128*0.25*kk)-3, f"Step {snapshot_steps[kk-1]}",
color=my_cmap((max_steps/2 + snapshot_steps[kk-1]/2)/max_steps))
plt.tight_layout()
# -
| notebooks/reward_figures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploratory Data Analysis in Python
# EDA is an approach to analyzing data sets to summarize their main characteristics, often with visual methods. A statistical model can be used or not, but primarily EDA is for seeing what the data can tell us beyond the formal modeling or hypothesis testing task.
# In this module we will be working on House Price Prediction Dataset.
# we will cover :
# - Basics of EDA
# - Handling Missing values
# - Detecting Outliers
# - Handling Outliers
# - Histogram
# - Correlation Heatmap
# - Scatterplot
# - Boxplot
# - Feature Engineering
# ### Import Libraries
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import os
# %matplotlib inline
# -
# Read CSV file in Pandas dataframe
os.chdir("E:\Project=House Prices Advanced Regression Techniques")
data=pd.read_csv("train.csv")
data.head()
# In this above dataset, SalePrice column is dependent variable and
# other columns are indepandent variables which are correlated to dependent variable.
data.columns
#Check shape of dataframe
data.shape
#Handling Missing values
missing=data.isnull().sum()
missing=missing[missing>0]
missing.sort_values(inplace=True)
plt.figure(figsize=(15,8))
missing.plot.bar()
#plot distplot
sns.set(rc={'figure.figsize':(12,8)})
sns.distplot(data['SalePrice'],kde=True,bins=20);
data['SalePrice'].describe()#data describtion
# ### Correlation
numeric_features=data.select_dtypes(include=[np.number])
numeric_features.columns#numeric_columns
categorical_features=data.select_dtypes(include=[np.object])
categorical_features.columns#categorical_columns
#Correlation between saleprice and numeric_features
#Selecting top correlated features
correlation=numeric_features.corr()
print(correlation['SalePrice'].sort_values(ascending=False),'\n')
# + active=""
# Now,we plot heatmap of correlation of numeric features.
#
# We need a heatmap to visualize two categories, and the values associated with them using colors.
# The use of color shades makes it easy to understand the data in a sight without giving any stress to the brain.
#
# Note that, we can not plot catrogrical heatmap because of string and notation
# -
#plot heatmap of correlation of numeric features with Sale Price
f,ax=plt.subplots(figsize=(14,12))
sns.heatmap(correlation,square=True,vmax=0.8)
plt.title('correlation of numeric features with Sale Price',y=1,size=16)
#plot heatmap of correlation of k features with Sale Price
k=11
cols=correlation.nlargest(k,'SalePrice')['SalePrice'].index
print(cols)
cm=np.corrcoef(data[cols].values.T)
f,ax=plt.subplots(figsize=(14,12))
sns.heatmap(cm,vmax=0.8,linewidths=0.01,square=True,annot=True,cmap='viridis',
linecolor="white",xticklabels=cols.values,annot_kws={'size':12},yticklabels=cols.values)
# ### Scatterplot
# Scatter plots are the graphs that present the relationship between two variables in a data-set. It represents data points on a two-dimensional plane or on a Cartesian system. The variable or attribute which is independent is plotted on the X-axis, while the dependent variable is plotted on the Y-axis. These plots are often called scatter graphs
#plot scattrplot
sns.scatterplot(x='GarageCars',y='SalePrice',data=data)
#plot regplot(to see correlation between two independent variables)
sns.regplot(x='GarageCars',y='GarageArea',data=data,scatter=True,fit_reg=True)
sns.regplot(x='GarageCars',y='GarageArea',data=data,scatter=True,fit_reg=False)
# +
#Scatter plots between the most correlated Variables
fig, ((ax1, ax2),(ax3, ax4),(ax5, ax6))= plt.subplots(nrows=3, ncols=2, figsize=(14,10))
sns.regplot(x='OverallQual',y='SalePrice',data=data,scatter=True,fit_reg=True, ax=ax1)
sns.regplot(x='GrLivArea',y='SalePrice',data=data,scatter=True,fit_reg=True, ax=ax2)
sns.regplot(x='GarageArea',y='SalePrice',data=data,scatter=True,fit_reg=True, ax=ax3)
sns.regplot(x='FullBath',y='SalePrice',data=data,scatter=True,fit_reg=True, ax=ax4)
sns.regplot(x='YearBuilt',y='SalePrice',data=data,scatter=True,fit_reg=True, ax=ax5)
sns.regplot(x='WoodDeckSF',y='SalePrice',data=data,scatter=True,fit_reg=True, ax=ax6)
# -
# ### Boxplot
#plot boxplt to Detecting Outliers
sns.boxplot(x=data["SalePrice"])
f,ax=plt.subplots(figsize=(16,10))
fig=sns.boxplot(x='SaleType', y='SalePrice', data=data)
fig.axis(ymin=0, ymax=800000);
xt = plt.xticks(rotation=45)
f,ax=plt.subplots(figsize=(16,10))
fig=sns.boxplot(x='OverallQual', y='SalePrice', data=data)
fig.axis(ymin=0, ymax=800000);
# ### Remove Outliars
first_quartile=data['SalePrice'].quantile(0.25)
third_quartile=data['SalePrice'].quantile(0.75)
IQR=third_quartile-first_quartile
new_boundary=third_quartile+ 3*IQR
data.drop(data[data["SalePrice"]>new_boundary].index,axis=0,inplace=True)
sns.boxplot(x=data["SalePrice"])
# ### REMOVE BAD FEATURES FROM DATA
# Multicollinear features:
# 1. GarageArea <-> GarageCars
# 2. TotalBsmtSF <-> 1stFlrSF
# 3. TotRmsAbvGrd <-> GrLivArea
# 4. GrLivArea <-> FullBath
#
# Features with missing value more than 20%:
# 5. FireplaceQu 690/1460= 47%
# 6. Fence 1179/1460= 80%
# 7. Alley 1369 > 90%
# 8. MiscFeature 1406 > 90%
# 9. PoolQC 1453 > 90%
#
# Features with poor correlation with the target feature (SalePrice):
# 10. LotFrontage
# 11. WoodDeckSF
# 12. 2ndFlrSF
# 13. OpenPorchSF
# 14. HalfBath
# 15. LotArea
# 16. BedroomAbvGr
# 17. ScreenPorch
# 18. PoolArea
# 19. MoSold
# 20. 3SsnPorch
# 21. BsmtFinSF2
# 22. BsmtHalfBath
# 23. MiscVal
# 24. Id
# 25. LowQualFinSF
# 26. YrSold
# 27. OverallCond
# 28. MSSubClass
# 29. EnclosedPorch
# 30. KitchenAbvGr
#
cols_to_remove = ['BsmtFinSF1','LotFrontage','WoodDeckSF','2ndFlrSF','OpenPorchSF','HalfBath','LotArea',
'BsmtFullBath','BsmtUnfSF','BedroomAbvGr','ScreenPorch','PoolArea','MoSold','3SsnPorch',
'BsmtFinSF2','BsmtHalfBath','MiscVal','Id','LowQualFinSF','YrSold','OverallCond',
'MSSubClass','EnclosedPorch','KitchenAbvGr','FireplaceQu','Fence','Alley','MiscFeature',
'PoolQC','GarageCars','1stFlrSF','GrLivArea','FullBath']
# ### Drop the columns
data.drop(cols_to_remove,axis =1,inplace=True)
data.head()
data.columns
data.shape
| 07Satyam_ML_EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# Overview of the problem to solve with this model
# - Intially train a simple model off few features utilizing a training pipeline to predict patients who 1) are septic or 2) at-risk of becoming septic
# - Develop heuristics to filter out patients who are septic to avoid costs for model inferencing
# - Train more complex models to infer patients at-risk of becoming septic based on more complex data sources and features
# Overview of Sepsis Indicators:
#
# **bold indicates availability from our data**
#
# Sepsis - Systemic inflammatory response syndrome (SIRS) 2 or more are met:
# 1. **Body temperature > 38.5°C or < 35.0°C**
# 1. **Heart rate > 90 beats per minute**
# 1. **Respiratory rate > 20 breaths per minute or arterial CO2 tension < 32 mm Hg or need for mechanical ventilation**
# 1. **White blood cell count > 12,000/mm3 or < 4,000/mm3 or immature forms > 10%**
#
# Severe sepsis - Sepsis and at least one sign of organ hypoperfusion or organ dysfunction:
# 1. Areas of mottled skin
# 1. Capillary refilling time ≥ 3 s
# 1. Urinary output < 0.5 mL/kg for at least 1 h or renal replacement therapy
# 1. **Lactates > 2 mmol/L**
# 1. Abrupt change in mental status or abnormal electroencephalogram
# 1. **Platelet counts < 100,000/mL or disseminated intravascular coagulation**
# 1. Acute lung injury—acute respiratory distress syndrome
# 1. Cardiac dysfunction (echocardiography)
#
# Septic shock - Severe sepsis and one of:
# 1. Systemic mean blood pressure of < 60 mm Hg (< 80 mm Hg if previous hypertension) after 20–30 mL/kg starch or 40–60 mL/kg serum saline, or pulmonary capillary wedge pressure between 12 and 20 mm Hg
# 1. Need for dopamine > 5 μg/kg per min or norepinephrine or epinephrine < 0.25 μg/kg per min to maintain mean blood pressure above 60 mm Hg (> 80 mm Hg if previous hypertension) ### Refractory septic shock
# 1. Need for dopamine > 15 μg/kg per min or norepinephrine or epinephrine > 0.25 μg/kg per min to maintain mean blood pressure above 60 mm Hg (> 80 mm Hg if previous hypertension)
# Overview of this projects current goal for the Data Engineering
# - use only the Patient Vital signs (pat_vitals_labeled-dataSepsis.csv) to indentify predictive signals (columns)
# - generate a data preprocessing pipeline for feeding data to the model
# Overview of the data for this project
# - Data was originally based on a Kaggle project https://www.kaggle.com/maxskoryk/datasepsishttps://www.kaggle.com/maxskoryk/datasepsis
# - Major changes were made due to the data bias for demographics influencing sepsis indicator AND the sepsis indicators were not accurate
# - Patient ID, record date and record time were added
# - HR, Temp and RR were generated to accurately reflect values and patient percentage representation in the believed real world
# - Data was split into 3 separate labeled data files
# - Patient Demographics (pat_demog_labeled_dataSepsis.csv)
# - Patient Laboratory Values (pat_labs_labeled_dataSepsis.csv)
# - Patient Vital Signs (pat_vitals_labeled_dataSepsis.csv)
# Overview of steps in the notebook Overview of steps in the notebook
# - Fetch and write the data for updates using urllib, zipfile, and os for OS agnostic handling
# - Load the data as a Dataframe using Pandas
# - Explore the Dataframe with Pandas
# - Split the data into train and test sets with Scikit-Learn
# - Visualize the train data with Matplotlib and Seaborn
# - Explore correlation among features
# - Feature down selection
# Import Packages
# +
# data ingestion
import urllib.request
import os
import zipfile
# data manipulation
import pandas as pd
import numpy as np
# data visualization
import seaborn as sns
import matplotlib.pyplot as plt
# Add directory above current directory to path
import sys; sys.path.insert(0, '..')
# possible removeable of submodules
#from submodules.fetch_data import fetch_data
#from submodules.load_data import load_data
from pandas.plotting import scatter_matrix
from IPython.display import Image
# data splitting
from sklearn.model_selection import train_test_split
# -
# Fetch the data
# +
# fetch the data using a python function, commented out b/c cannot use with Kaggle source
#fetch_data()
# -
# Load the data
# +
# load the data using a python function
#data = load_data()
# without using a python function
# set for the Signal definitions
attr_path = "../../data/dataSepsis/csv_format/attribute_definitions.csv"
attr = pd.read_csv(attr_path, sep=",")
# set for the Patient Vital Signs
csv_path = "../../data/dataSepsis/csv_format/pat_vitals_labeled-dataSepsis.csv"
data = pd.read_csv(csv_path, sep=",")
# -
# Review of Signal Definitions available from the data source
# list the attributes definition file for the Patient Vital Signs
attr.head(13)
# First glance of raw data. First 10 rows.
data.head(10)
# What types of columns and data do we have?
# 1. are we missing any records?
# 1. are we missing entries?
# 1. do we have any categorical data?
# If yes for any above questions, plan to collect, drop, or convert.
data.info()
data.count(axis=0).sort_values(ascending=True)
# Forward Action
# 1. drop record_date
# 1. drop record_time
# 1. drop EtCO2
# What does calculus expose about the data?
#
# Average (mean), minimum (min), maximum (max) are self-explanatory
#
# Standard deviation (std) how dispersed the values are
# - normal (Gaussian) distribution follows 68-95-99.7 rule
# - % of values are within 1 std
# - % of values are within 2 std
# - % of values are within 3 std
#
# 1st (25%), median (50%), 3rd (75%) quartiles or percentiles, for example:
# - % of the patients had a temp lower than 36.3°C
# - % of the patients had a resp higher than 20.5 breaths per minute
data.describe(include="all").T
# Forward Action
# 1. Scale remaining signals between 0 and 1
# Are Sepsis patient entries representative of the real world? A common split is:
# - 93% of patients are not septic
# - 7% of patients are septic
data["isSepsis"].value_counts(normalize=True).to_frame()
# Create a Test Dataset
#
# - Performing this early minimizes generalization and bias you may inadvertently apply to your system.
# - A test set of data involves: picking ~(10, 15, or 20)% of the instances randomly and setting them aside.
# - you never want your model to see the entire dataset
# - you should plab to fetch new data for training
# - you want to maintain the same percentage of training data against the entire dataset
# - you want a representative training dataset (~7% septic positive)
# +
X_train, X_test, y_train, y_test = \
train_test_split(data.drop("isSepsis", axis=1),
data["isSepsis"], test_size=0.15,
random_state=42, stratify=data["isSepsis"])
print("Training data: ", X_train.shape)
print("Training labels: ", y_train.shape)
print("Testing data: ", X_test.shape)
print("Testing labels: ", y_test.shape)
# -
# Plot the non-septic (0) vs. septic (1) patient records to identify clear distinctions?
vitals = ["HR", # Heart Rate normal adult 60 - 100 beats per minute (bpm)
'O2Sat', # Oxygen saturation normal adult 97% - 100% (%)
'Temp', # Temperature normal 97.8°F/36.5°C - 99°F/37.2°C (°C)
'SBP', # Systolic Blood Pressure normal < 120 mmHg (mm Hg) (indicates how much pressure your blood is exerting against your artery walls when the heart beats.)
'DBP', # beclomethasone dipropionate normal < 80 mmHg (mm Hg) (indicates how much pressure your blood is exerting against your artery walls while the heart is resting between beats.)
'MAP', # Mean Arterial Pressure (mm Hg)
'Resp', # Respiration rate 12<normal<20 breaths / minute (bpm)
"EtCO2" # End-tidal CO2 maximum concentration of CO2 at exhalation normal 35 - 45 mmHg (mm Hg)
]
sns.set_theme(context="notebook", style="ticks", palette="colorblind")
plt.figure(figsize=(18,12))
plt.subplots_adjust(hspace = .5)
for i, column in enumerate(vitals, 1):
plt.subplot(4,2,i)
# stat = Aggregate statistic to compute in each bin density normalizes counts so that the area of the histogram is 1
# common_norm = False = normalize each histogram independently
# kde = If True, compute a kernel density estimate to smooth the distribution and show on the plot as (one or more) line(s)
sns.histplot(data=X_train, x=column, hue=y_train, stat="density", common_norm=False, bins=60, kde=True)
# Observations
# 1. HR, Temp, MAP, Resp all have clear separation between Septic and non-Septic patients
# 1. SBP and DBP are used to calculate MAP, could drop
# 1. no clear pattern in O2Sat and EtCO2
# Is there correlation between signals?
# +
corr_features = [
# vitals
"HR", # Heart Rate normal adult 60 - 100 beats per minute (bpm)
"O2Sat", # Oxygen saturation normal adult 97% - 100% (%)
"Temp", # Temperature normal 97.8°F/36.5°C - 99°F/37.2°C (°C)
"SBP", # Systolic Blood Pressure normal < 120 mmHg (mm Hg)
"DBP", # beclomethasone dipropionate normal < 80 mmHg (mm Hg)
"MAP", # Mean Arterial Pressure (mm Hg)
"Resp", # Respiration rate 12<normal<20 breaths / minute (bpm)
"EtCO2", # End-tidal CO2 maximum concentration of CO2 at exhalation normal 35 - 45 mmHg (mm Hg) = REMOVED missing too much data
]
corr_matrix = data[corr_features].corr()
mask = np.zeros_like(corr_matrix)
mask[np.triu_indices_from(mask)] = True
plt.figure(figsize=(26,22))
sns.heatmap(corr_matrix, mask=mask, square=True, annot=True, fmt=".2f", center=0, linewidths=.5, cmap="RdBu")
# -
Observations
1. DBP and SBP are used to calculate MAP, so they are highly correlated
# Another view is correlation against the label:
corr_matrix = data.corr()
corr_matrix["isSepsis"].sort_values(ascending=False)
# Observation
# 1. Resp, HR, and Temp are highly correlated to Septic patients
# Reference List:
# - https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6304323/
# - https://www.aafp.org/afp/2013/0701/p44.html
# - https://www.kaggle.com/maxskoryk/datasepsis
# - https://www.nursingcenter.com/ncblog/march-2017/elevated-lactate-%E2%80%93-not-just-a-marker-for-sepsis-an
# - https://unitslab.com/node/74
| notebooks/data-exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#-*- coding:utf-8 -*-
from commonTool import *
import glob
# +
#################################################
# EVALUATE-PREPARE.xml 실행되어야 함.
# -
# +
# 기준일 로딩
bf = open(resourceDir + 'BASIS-DAYS.txt', 'r', encoding='utf-8')
# 0.YYYYMM, 1.FIRST_DAY, 2.SECOND_DAY, 3.PREV_DAY, 4.LAST_DAY, 5.NEXT_DAY
lineText = bf.readline() # 제목
basisDays = []
quaterBasisDays = []
while True:
lineText = bf.readline()
if not lineText:
break
lineText = lineText.strip()
if len(lineText) <= 0:
continue
dayData = lineText.split('\t')
if dayData[0][0:4] == '2012' or dayData[0] == '201301': continue # 데이터 부족
basisDays.append([dayData[0], dayData[3], dayData[4]])
if dayData[0] > '202001':
quaterBasisDays.append([dayData[0], dayData[3], dayData[4]])
bf.close()
# -
# +
# 로직 적용한 결과 정리
def makeAnalysisReport(resultFolder, initialCash, period, feeRate = 0.015, withProcess = True):
statResult = [f for f in glob.glob(resultFolder + '*-STAT.txt')]
if len(statResult) == 0:
print('no result file')
return
statResult.sort()
m = 0
y = 0
cash = initialCash
maxV = initialCash
maxMdd = 0
cagr = 0
if withProcess:
print('\t'.join(['Year', '매수일', '매도일', '손익', '투자금', '수익률', 'CAGR', 'MDD']))
for statPath in statResult:
m += period
sf = open(statPath, 'r', encoding='utf-8')
lineText = sf.readline() # 제목: --> 매수일, 매도일, R_SUM
lineText = sf.readline() # 결과
rec = lineText.split('\t')
profitLoss = int(rec[2])
newCash = round((cash + profitLoss) * (1.0 - feeRate))
maxV = max(maxV, newCash)
if newCash < cash:
mdd = (newCash - maxV) / maxV
maxMdd = min(mdd, maxMdd)
y = m / 12
cagr = pow(newCash / initialCash, 1 / y) - 1
cagr = round(cagr * 100, 2)
if withProcess:
print('\t'.join([str(y), rec[0], rec[1],
format(profitLoss, ',d').rjust(12), format(newCash, ',d').rjust(15),
str(round(profitLoss / cash * 100, 2)).rjust(9), str(cagr).rjust(9), str(round(maxMdd * -100, 2))]))
sf.close()
cash = newCash
if withProcess: print('')
print('Year:', y, ' Invest:', format(initialCash, ',d'), ' Cash:', format(cash, ',d'),
' CAGR:', cagr, '%', ' Max MDD:', round(maxMdd * -100, 2), '%\n\n')
# -
# +
testResult = [
'4PXX-25-10T-10-M2',
'5PXR-25-10T-10-M2',
# '4PXX-25-10T-15-M2',
# '4PXX-25-10T-20-M2',
'END' # 콤마 편하게 하려고 넣었음
]
for key in testResult:
if key == 'END': break
print('#', key)
makeAnalysisReport(outputRawPath + 'logicResult' + os.path.sep + key + os.path.sep, 10000000,
(3 if key.endswith('-P3') else 6), 0.015, True)
# -
# +
testResult = [
'4PXX-25-10T-10-M2',
'4PXX-25-10T-10-M3',
'4PXX-25-10T-10-M4',
'4PXX-25-10T-10-M5',
'4PXX-25-10T-10-M6',
'END' # 콤마 편하게 하려고 넣었음
]
for key in testResult:
if key == 'END': break
print('#', key)
makeAnalysisReport(outputRawPath + 'logicResult' + os.path.sep + key + os.path.sep, 10000000, 6, 0.015, True)
# +
testResult = [
'4PXX-25-10T-10-M2',
'4PXX-25-10T-20-M2',
'END' # 콤마 편하게 하려고 넣었음
]
for key in testResult:
if key == 'END': break
print('#', key)
makeAnalysisReport(outputRawPath + 'logicResult' + os.path.sep + key + os.path.sep, 10000000, 6, 0.015, True)
# -
# +
testResult = [
# '4PXX-25-10T-20-M2',
'Q-4PXX-25-10T-10-M2',
'Q-RND-25-10T-10-M2',
'END' # 콤마 편하게 하려고 넣었음
]
for key in testResult:
if key == 'END': break
print('#', key)
makeAnalysisReport(outputRawPath + 'logicResult' + os.path.sep + key + os.path.sep, 10000000, 3, 0.015, False)
# -
makeAnalysisReport(outputRawPath + 'logicResult' + os.path.sep + '4PXX-25-10T-10-M2' + os.path.sep, 10000000, 6, 0.015, True)
# +
analId = '4PXX' # 로직명: 4PXX, MAGIC, %PXR, RND
calcDate = yyyymm() # 로직 실행할 대상 월 '202107'
# sys.argv[0] 은 python 파일
if len(sys.argv) > 1:
analId = sys.argv[1]
if len(sys.argv) > 2:
calcDate = sys.argv[2]
print('running', analId, calcDate)
cash = 10000000 # 초기 투자금
period = 6 # 리밸런싱 기간 (월)
buyCount = 20 # 매수 종목 개수
totalAmount = 25 # 25 # 시가총액 하위 퍼센트 조건
volumeLimit = 10000 # 10000 # 거래량 조건
startMonth = 2
analTitle = analId + '-' + str(totalAmount) + '-' + str(round(volumeLimit / 1000)) + 'T-' + str(buyCount) \
+ '-M' + str(startMonth) + ('-P' + str(period) if period != 6 else '')
# 로직 스크립트
scriptPathName = crawlegoScriptPath + 'ANAL-' + analId + '.xml'
# 결과 저장 위치
resultFolder = outputRawPath + 'logicResult' + os.path.sep + analTitle + os.path.sep
# [os.remove(f) for f in glob.glob(resultFolder + '*.txt')]
mkdir(resultFolder)
done = False
for basis in basisDays:
if basis[0] != calcDate: continue
parameter = {
'BASIS_DAY': basis[1],
'BUY_DAY': basis[2],
'SELL_DAY': basis[2],
'CASH': str(cash),
'TOP_N': str(buyCount),
'PERIOD': 'ANNUAL',
'IN_PATH': outputRawPath + 'index' + os.path.sep + (os.path.sep if runServerType == 1 else ''),
'OUT_PATH': resultFolder + (os.path.sep if runServerType == 1 else ''),
'TOTAL_AMOUNT': str(totalAmount),
'VOL_LIMIT': str(volumeLimit),
'DO_SERVER': '192.168.127.12'
}
print(basis, 'running...')
retCode = runDashScript(scriptPathName, parameter, False)
print(basis, 'returns', retCode)
print('result folder:', resultFolder)
done = True
break
if not done:
print('invalid argument', analId, calcDate)
# -
len(sys.argv)
# +
# 년간 재무 데이터를 이용한 분석
analId = '4PXX' # 로직명: 4PXX, MAGIC, %PXR, RND
inital_cash = 10000000 # 초기 투자금
period = 6 # 리밸런싱 기간 (월)
buyCount = 10 # 매수 종목 개수
totalAmount = 25 # 25 # 시가총액 하위 퍼센트 조건
volumeLimit = 10000 # 10000 # 거래량 조건
startMonth = 2
analTitle = analId + '-' + str(totalAmount) + '-' + str(round(volumeLimit / 1000)) + 'T-' + str(buyCount) \
+ '-M' + str(startMonth) + ('-P' + str(period) if period != 6 else '')
print('running', analTitle)
feeRate = 0.015 # 1.5% 수수료
# 로직 스크립트
scriptPathName = crawlegoScriptPath + 'ANAL-' + analId + '.xml'
# 결과 저장 위치
resultFolder = outputRawPath + 'logicResult' + os.path.sep + analTitle + os.path.sep
# [os.remove(f) for f in glob.glob(resultFolder + '*.txt')]
mkdir(resultFolder)
# 계산 시작
m = 0
cash = inital_cash
for i in range(startMonth - 2, len(basisDays), period):
basis = basisDays[i]
sellIdx = min(i + period, len(basisDays) - 1)
parameter = {
'BASIS_DAY': basis[1],
'BUY_DAY': basis[2],
'SELL_DAY': basisDays[sellIdx][2],
'CASH': str(cash),
'TOP_N': str(buyCount),
'PERIOD': 'ANNUAL',
'IN_PATH': outputRawPath + 'index' + os.path.sep + (os.path.sep if runServerType == 1 else ''),
'OUT_PATH': resultFolder + (os.path.sep if runServerType == 1 else ''),
'TOTAL_AMOUNT': str(totalAmount),
'VOL_LIMIT': str(volumeLimit),
'DO_SERVER': '192.168.127.12'
}
m += period
retCode = runDashScript(scriptPathName, parameter, False)
print(m / 12, basis[0], basis[2], basisDays[sellIdx][2], 'ret code', retCode)
if retCode != 0:
continue
# 결과 정리
statPath = resultFolder + basis[1] + '-STAT.txt'
sf = open(statPath, 'r', encoding='utf-8')
lineText = sf.readline() # 제목: 매수일, 매도일, R_SUM
lineText = sf.readline() # 결과
rec = lineText.split('\t')
profitLoss = int(rec[2])
sf.close()
cash = round((cash + profitLoss) * (1.0 - feeRate))
# end of for
makeAnalysisReport(resultFolder, inital_cash, period, feeRate, True)
# end of script
# -
# +
# 분기 재무 데이터를 이용한 분석
analId = 'RND' # 로직명
inital_cash = 10000000 # 초기 투자금
period = 3 # 리밸런싱 기간 (월)
buyCount = 10 # 매수 종목 개수
totalAmount = 200 # 25 # 시가총액 하위 퍼센트 조건
volumeLimit = 1 # 10000 # 거래량 조건
startMonth = 2
analTitle = 'Q-' + analId + '-' + str(totalAmount) + '-' + str(round(volumeLimit / 1000)) + 'T-' + str(buyCount) + '-M' + str(startMonth)
print('running', analTitle)
feeRate = 0.015 # 1.5% 수수료
# 로직 스크립트
scriptPathName = crawlegoScriptPath + 'ANAL-' + analId + '.xml'
# 결과 저장 위치
resultFolder = outputRawPath + 'logicResult' + os.path.sep + analTitle + os.path.sep
# [os.remove(f) for f in glob.glob(resultFolder + '*.txt')]
mkdir(resultFolder)
# 계산 시작
m = 0
cash = inital_cash
for i in range(startMonth - 2, len(quaterBasisDays), period):
basis = quaterBasisDays[i]
sellIdx = min(i + period, len(quaterBasisDays) - 1)
parameter = {
'BASIS_DAY': basis[1],
'BUY_DAY': basis[2],
'SELL_DAY': quaterBasisDays[sellIdx][2],
'CASH': str(cash),
'TOP_N': str(buyCount),
'PERIOD': 'QUARTER',
'IN_PATH': outputRawPath + 'index' + os.path.sep + (os.path.sep if runServerType == 1 else ''),
'OUT_PATH': resultFolder + (os.path.sep if runServerType == 1 else ''),
'TOTAL_AMOUNT': str(totalAmount),
'VOL_LIMIT': str(volumeLimit),
'DO_SERVER': '192.168.127.12'
}
m += period
retCode = runDashScript(scriptPathName, parameter, False)
print(m / 12, basis[0], 'ret code', retCode)
if retCode != 0:
continue
# 결과 정리
statPath = resultFolder + basis[1] + '-STAT.txt'
sf = open(statPath, 'r', encoding='utf-8')
lineText = sf.readline() # 제목: 매수일, 매도일, R_SUM
lineText = sf.readline() # 결과
rec = lineText.split('\t')
profitLoss = int(rec[2])
sf.close()
cash = round((cash + profitLoss) * (1.0 - feeRate))
# end of for
makeAnalysisReport(resultFolder, inital_cash, period, feeRate, True)
# end of script
# -
# +
resultFolder = outputRawPath + 'logicResult' + os.path.sep + '4PXX-25-10T-10-M2' + os.path.sep
saveFolder = temporaryPath + 'analMM' + os.path.sep
scriptPathName = crawlegoScriptPath + 'CALC-MOMENTUM.xml'
mmCount = 3
analResult = [f for f in glob.glob(resultFolder + '*-RESULT.txt')]
for selFile in analResult:
p = selFile.rfind(os.path.sep)
basisDay = selFile[p+1:p+9]
parameter = {
'BASIS_DAY': basisDay,
'IN_PATH': resultFolder + (os.path.sep if runServerType == 1 else ''),
'OUT_PATH': saveFolder + (os.path.sep if runServerType == 1 else ''),
'MM_COUNT': str(mmCount),
'DO_SERVER': '192.168.127.12'
}
retCode = runDashScript(scriptPathName, parameter, False)
print(basisDay, 'returns', retCode)
# -
# +
resultFolder = outputRawPath + 'logicResult' + os.path.sep + '4PXX-25-10T-10-M2' + os.path.sep
saveFolder = temporaryPath + 'analMM' + os.path.sep
scriptPathName = crawlegoScriptPath + 'CALC-AVGINDEX.xml'
mmCount = 3
analResult = [f for f in glob.glob(resultFolder + '*-RESULT.txt')]
for selFile in analResult:
p = selFile.rfind(os.path.sep)
basisDay = selFile[p+1:p+9]
parameter = {
'BASIS_DAY': basisDay,
'IN_PATH': resultFolder + (os.path.sep if runServerType == 1 else ''),
'OUT_PATH': saveFolder + (os.path.sep if runServerType == 1 else ''),
'DO_SERVER': '192.168.127.12'
}
retCode = runDashScript(scriptPathName, parameter, False)
print(basisDay, 'returns', retCode)
# -
# +
DD = [
['20130227', '20130228', '20130731'],
['20130829', '20130930', '20131129'],
['20140227', '20140228', '20140829'],
['20140828', '20140828', '20140828'],
['20150226', '20150227', '20150630'],
['20150828', '20150828', '20150828'],
['20160226', '20160229', '20160831'],
['20160830', '20160831', '20170228'],
['20170227', '20170331', '20170831'],
['20170830', '20171130', '20180228'],
['20180227', '20180228', '20180731'],
['20180830', '20180830', '20180830'],
['20190227', '20190228', '20190830'],
['20190829', '20191031', '20200228'],
['20200227', '20200228', '20200831'],
['20200828', '20201030', '20210226'],
['20210225', '20210226', '20210621']
]
resultFolder = outputRawPath + 'logicResult' + os.path.sep + '4PXX-25-10T-10-M2' + os.path.sep
saveFolder = 'D:\work\jupyter\krx\intermediate\profit' + os.path.sep
scriptPathName = crawlegoScriptPath + 'CALC-PROFIT.xml'
cash = 10000000
for day in DD:
parameter = {
'BASIS_DAY': day[0],
'BUY_DAY': day[1],
'SELL_DAY': day[2],
'CASH': str(cash),
'IN_PATH': resultFolder + (os.path.sep if runServerType == 1 else ''),
'OUT_PATH': saveFolder + (os.path.sep if runServerType == 1 else ''),
'DO_SERVER': '192.168.127.12'
}
retCode = runDashScript(scriptPathName, parameter, False)
print(day[0], 'returns', retCode)
sf = open(saveFolder + day[0] + '-STAT.txt', 'r', encoding='utf-8')
lineText = sf.readline() # 제목: --> 매수일, 매도일, R_SUM
lineText = sf.readline() # 결과
rec = lineText.split('\t')
profitLoss = int(rec[2])
sf.close()
cash += profitLoss
print(day[0], profitLoss, cash)
# -
# +
#
resultFolder = outputRawPath + 'highlow' + os.path.sep
mkdir(resultFolder)
scriptPathName = crawlegoScriptPath + 'POC-MOMENTUM.xml'
midSizeComp = resourceDir + 'midSizeComp.txt'
mf = open(midSizeComp, 'r', encoding='utf-8')
while True:
lineText = mf.readline()
if not lineText:
break
lineText = lineText.strip()
if len(lineText) <= 0:
continue
# lineText is company code
parameter = {
'COMP_CODE': lineText,
'OUT_PATH': resultFolder,
'DO_SERVER': '13.124.29.70'
}
retCode = runDashScript(scriptPathName, parameter, False)
print(lineText, 'returns', retCode)
break
mf.close()
# -
| applyLogic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from __future__ import print_function
import numpy as np
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers.core import Dense, Activation
from keras.optimizers import SGD
from keras.utils import np_utils
# +
np.random.seed(1671) # for reproducibility
# network and training
NB_EPOCH = 200
BATCH_SIZE = 128
VERBOSE = 1
NB_CLASSES = 10 # number of outputs = number of digits
OPTIMIZER = SGD() # SGD optimizer, explained later in this chapter
N_HIDDEN = 128
VALIDATION_SPLIT=0.2 # how much TRAIN is reserved for VALIDATION
# data: shuffled and split between train and test sets
#
(X_train, y_train), (X_test, y_test) = mnist.load_data()
# +
X_train.shape
# the 3-ed number is 1 accroding to the y_train.
# And we need to turn this 2-dim array to 1 dim.
X_train[3]
X_train.ndim
# -
#X_train is 60000 rows of 28x28 values --> reshaped in 60000 x 784
RESHAPED = 784
#
X_train = X_train.reshape(60000, RESHAPED)
X_test = X_test.reshape(10000, RESHAPED)
X_train = X_train.astype('float32')
X_test = X_test.astype('float32')
X_train.shape
y_train.shape
y_train[:6]
28*28
# +
# X_train[:1]
# -
# normalize
#
X_train /= 255
X_test /= 255
print(X_train.shape[0], 'train samples')
print(X_test.shape[0], 'test samples')
X_train[:1]
# +
# One-Hot Encoder
# convert class vectors to binary class matrices
Y_train = np_utils.to_categorical(y_train, NB_CLASSES)
Y_test = np_utils.to_categorical(y_test, NB_CLASSES)
#Y_train[:6]
Y_train.shape
# +
# 10 outputs
# final stage is softmax
model = Sequential()
model.add(Dense(NB_CLASSES, input_shape=(RESHAPED,)))
model.add(Activation('softmax'))
model.summary()
model.compile(loss='categorical_crossentropy',
optimizer=OPTIMIZER,
metrics=['accuracy'])
history = model.fit(X_train, Y_train,
batch_size=BATCH_SIZE, epochs=NB_EPOCH,
verbose=VERBOSE, validation_split=VALIDATION_SPLIT)
score = model.evaluate(X_test, Y_test, verbose=VERBOSE)
print("\nTest score:", score[0])
print('Test accuracy:', score[1])
| Chapter01/keras_MINST_V1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modeling and Simulation in Python
#
# Chapter 1: Modeling
#
# Copyright 2017 <NAME>
#
# License: [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0)
# ## Jupyter
#
# Welcome to Modeling and Simulation, welcome to Python, and welcome to Jupyter.
#
# This is a Jupyter notebook, which is a development environment where you can write and run Python code. Each notebook is divided into cells. Each cell contains either text (like this cell) or Python code (like the cell below this one).
#
# ### Selecting and running cells
#
# To select a cell, click in the left margin next to the cell. You should see a blue frame surrounding the selected cell.
#
# To edit a code cell, click inside the cell. You should see a green frame around the selected cell, and you should see a cursor inside the cell.
#
# To edit a text cell, double-click inside the cell. Again, you should see a green frame around the selected cell, and you should see a cursor inside the cell.
#
# To run a cell, hold down SHIFT and press ENTER. If you run a text cell, it will typeset the text and display the result.
#
# If you run a code cell, it runs the Python code in the cell and displays the result, if any.
#
# To try it out, edit this cell, change some of the text, and then press SHIFT-ENTER to run it.
# ### Adding and removing cells
#
# You can add and remove cells from a notebook using the buttons in the toolbar and the items in the menu, both of which you should see at the top of this notebook.
#
# You might want to try the following exercises:
#
# 1. From the Insert menu select "Insert cell below" to add a cell below this one. By default, you get a code cell, and you can see in the pulldown menu that says "Code".
#
# 2. In the new cell, add a print statement like `print('Hello')`, and run it.
#
# 3. Add another cell, select the new cell, and then click on the pulldown menu that says "Code" and select "Markdown". This makes the new cell a text cell.
#
# 4. In the new cell, type some text, and then run it.
#
# 5. Use the arrow buttons in the toolbar to move cells up and down.
#
# 6. Use the cut, copy, and paste buttons to delete, add, and move cells.
#
# 7. As you make changes, Jupyter saves your notebook automatically, but if you want to make sure, you can press the save button, which looks like a floppy disk from the 1990s.
#
# 8. Finally, when you are done with a notebook, selection "Close and Halt" from the File menu.
# ### Using the notebooks
#
# The notebooks for each chapter contain the code from the chapter along with addition examples, explanatory text, and exercises. I recommend you read the chapter first to understand the concepts and vocabulary, then run the notebook to review what you learned and see it in action, and then attempt the exercises.
#
# The notebooks contain some explanatory text, but it is probably not enough to make sense if you have not read the book. If you are working through a notebook and you get stuck, you might want to re-read (or read!) the corresponding section of the book.
#
# If you try to work through the notebooks without reading the book, you're gonna have a bad time. If you have previous programming experience, you might get through the first few notebooks, but sooner or later, you will get to the end of your leash, and you won't like it.
# ### Importing modsim
#
# The following cell imports `modsim`, which is a collection of functions we will use throughout the book. Whenever you start the notebook, you will have to run the following cell. It does two things:
#
# 1. It uses a Jupyter "magic command" to specify whether figures should appear in the notebook, or pop up in a new window.
#
# 2. It imports everything defined in `modsim`.
#
# Select the following cell and press SHIFT-ENTER to run it.
# +
# If you want the figures to appear in the notebook,
# and you want to interact with them, use
# # %matplotlib notebook
# If you want the figures to appear in the notebook,
# and you don't want to interact with them, use
# # %matplotlib inline
# If you want the figures to appear in separate windows, use
# # %matplotlib qt5
# To switch from one to another, you have to select Kernel->Restart
# %matplotlib qt5
from modsim import *
print('If this cell runs successfully, it produces no output other than this message.')
# -
# ## The penny myth
#
# The following cells contain code from the beginning of Chapter 1.
#
# `modsim` defines `UNITS`, which contains variables representing pretty much every unit you've ever heard of. The following to lines create new variables named `meter` and `second`.
meter = UNITS.meter
second = UNITS.second
# To find out what units are defined, type `UNITS.` in the next cell and then press TAB. You should see a pop-up menu with a list of units.
UNITS
# Create a variable named `a` and display its value:
a = 9.8 * meter / second**2
a
# Create `t` and display its value:
t = 4 * second
t
# If you create a variable and don't display the value, you don't get any output:
h = a * t**2
# Add a second line to the previous cell to display the value of `h`.
#
# Now let's solve the falling penny problem. The following lines set `h` to the height of the Empire State Building and compute the time it would take a penny to fall, assuming constant acceleration.
h = 381 * meter
t = sqrt(2 * h / a)
t
# Given `t`, we can compute the velocity of the penny when it lands.
v = a * t
v
# We can convert from one set of units to another like this:
mile = UNITS.mile
hour= UNITS.hour
v.to(mile/hour)
# **Exercise:** In reality, air resistance prevents the penny from reaching this velocity. At about 20 meters per second, the force of air resistance equals the force of gravity and the penny stops accelerating.
#
# As a simplification, let's assume that the acceleration of the penny is `a` until the penny reaches 20 meters per second, and then 0 afterwards. What is the total time for the penny to fall 381 meters?
# Solution goes here
from modsim import *
# Make a variable for terminal velocity because that's good practice.
terminal_velocity = 20 * meter / second
# Use v = a * t to find the time spent falling.
time_before_terminal_velocity = terminal_velocity / a
# To find the time spent falling at terminal velocity, first find the height left to fall and then use kinematic equations to find the time.
amount_fallen_before_terminal_velocity = 0.5 * terminal_velocity * time_before_terminal_velocity
# Then, let's find the height left to fall.
height_remaining = h - amount_fallen_before_terminal_velocity
# And then use the distance formula to find how long this takes.
time_after_terminal_velocity = height_remaining / terminal_velocity
# Lastly, we can find and print the sum of both times.
total_time = time_before_terminal_velocity + time_after_terminal_velocity
print(total_time)
# ## Modeling a bikeshare system
# We'll start with a `System` object that represents the number of bikes at each station.
bikeshare = System(olin=10, wellesley=2, babson=0)
# If you display the value of a `System` object, it lists the system variables and their values (not necessarily in the order you defined them):
bikeshare
# We can access the system variables using dot notation.
bikeshare.olin
bikeshare.wellsley
bikeshare.babson
# **Exercise:** What happens if you spell the name of a system variable wrong? Edit the previous cell, change the spelling of `wellesley`, and run the cell again.
#
# The error message uses the word "attribute", which is another name for what we are calling a system variable.
# **Exercise:** Add a third attribute called `babson` with initial value 0, and print the state of `bikeshare` again.
# ## Plotting
#
# `newfig` creates a new figure, which should appear either in the notebook or in a new window, depending on which magic command you ran in the first code cell.
#
# `plot` adds a data point to the figure; in this example, you should see a red square and a blue circle representing the number of bikes at each station.
newfig()
plot(bikeshare.olin, 'rs-')
plot(bikeshare.wellesley, 'bo-')
# We can use the operators `+=` and `-=` to increase and decrease the system variables. The following lines move a bike from Olin to Wellesley.
bikeshare.olin -= 1
bikeshare.wellesley += 1
bikeshare
# And the following lines plot the updated state of the system. You should see two new data points with lines connecting them to the old data points.
plot(bikeshare.olin, 'rs-')
plot(bikeshare.wellesley, 'bo-')
# **Exercise:** In the cell below, write a few lines of code to move a bike from Wellesley to Olin and plot the updated state.
# Solution goes here
# Move a bike to Olin
bikeshare.wellesley -= 1
bikeshare.olin += 1
# Plot the change.
plot(bikeshare.olin, 'rs-')
plot(bikeshare.wellesley, 'bo-')
# ## Functions
#
# Now we can take the code we've written so far and encapsulate it in functions.
def bike_to_wellesley():
bikeshare.olin -= 1
bikeshare.wellesley += 1
# When you define a function, it doesn't run the statements inside the function, yet.
def plot_state():
plot(bikeshare.olin, 'rs-', label='Olin')
plot(bikeshare.wellesley, 'bo-', label='Wellesley')
# Now when we run the functions, it runs the statements inside.
bike_to_wellesley()
plot_state()
bikeshare
# You should see two more data points that represent the current state of the system. If the figure is embedded in the notebook, you might have to scroll up to see the change.
#
# One common error is to omit the parentheses, which has the effect of looking up the function, but not running it.
bike_to_wellesley
# The output indicates that `bike_to_wellesley` is a function defined in a "namespace" called `__main__`, but you don't have to understand what that means.
# **Exercise:** Define a function called `bike_to_olin` that moves a bike from Wellesley to Olin. Run the new function and print or plot the results to confirm that it works.
# Solution goes here
def bike_to_olin():
"""
Moves a bike from Wellesley to Olin
"""
# Move a bike to Olin
bikeshare.wellesley -= 1
bikeshare.olin += 1
# Run the function.
bikeshare
bike_to_olin()
plot_state()
bikeshare
# ## Parameters
# Before we go on, let's start with a new state object and a new plot.
bikeshare = System(olin=10, wellesley=2)
newfig()
plot_state()
# Since we have two similar functions, we can create a new function, `move_bike` that takes a parameter `n`, which indicates how many bikes are moving, and in which direction.
def move_bike(n):
bikeshare.olin -= n
bikeshare.wellesley += n
# Now we can use `move_bike` to write simpler versions of the other functions.
# +
def bike_to_wellesley():
move_bike(1)
def bike_to_olin():
move_bike(-1)
# -
# When we define these functions, we replace the old definitions with the new ones.
#
# Now we can test them and update the figure.
bike_to_wellesley()
plot_state()
bikeshare
# Again, each time you run `plot_state` you should see changes in the figure.
bike_to_olin()
plot_state()
bikeshare
# At this point, `move_bike` is complicated enough that we should add some documentation. The text in triple-quotation marks is in English, not Python. It doesn't do anything when the program runs, but it helps people understand what this function does and how to use it.
def move_bike(n):
"""Move bikes.
n: number of bikes: positive moves from Olin to Wellesley;
negative moves from Wellesley to Olin
"""
bikeshare.olin -= n
bikeshare.wellesley += n
# Whenever you make a figure, you should put labels on the axes to explain what they mean and what units they are measured in. Here's how:
label_axes(title='Olin-Wellesley Bikeshare',
xlabel='Time step (min)',
ylabel='Number of bikes')
# Again, you might have to scroll up to see the effect.
#
# And you can save figures as files; the suffix of the filename indicates the format you want. This example saves the current figure in a PDF file.
savefig('chap01_fig01.pdf')
# **Exercise:** The following function definitions start with print statements so they display messages when they run. Run each of these functions (with appropriate arguments) and confirm that they do what you expect.
#
# Adding print statements like this to functions is a useful debugging technique. Keep it in mind!
# +
def move_bike_debug(n):
print('Running move_bike_debug with argument', n)
bikeshare.olin -= n
bikeshare.wellesley += n
def bike_to_wellesley_debug():
print('Running bike_to_wellesley_debug')
move_bike_debug(1)
def bike_to_olin_debug():
print('Running bike_to_olin_debug')
move_bike_debug(-1)
# -
# Solution goes here
move_bike_debug(2)
# Solution goes here
bike_to_wellesley_debug()
# Solution goes here
bike_to_olin_debug()
# ## Conditionals
# The function `flip` takes a probability and returns either `True` or `False`, which are special values defined by Python.
#
# In the following example, the probability is 0.7 or 70%. If you run this cell several times, you should get `True` about 70% of the time and `False` about 30%.
flip(0.7)
# Modify the argument in the previous cell and see what effect it has.
#
# In the following example, we use `flip` as part of an if statement. If the result from `flip` is `True`, we print `heads`; otherwise we do nothing.
if flip(0.7):
print('heads')
# With an else clause, we can print heads or tails depending on whether `flip` returns `True` or `False`.
if flip(0.7):
print('heads')
else:
print('tails')
# Now let's get back to the bikeshare system. Again let's start with a new `System` object and a new plot.
bikeshare = System(olin=10, wellesley=2)
newfig()
plot_state()
# Suppose that in any given minute, there is a 70% chance that a student picks up a bike at Olin and rides to Wellesley. We can simulate that like this.
# +
if flip(0.7):
bike_to_wellesley()
print('Moving a bike to Wellesley')
plot_state()
bikeshare
# -
# And maybe at the same time, there is also a 60% chance that a student at Wellesley rides to Olin.
# +
if flip(0.6):
bike_to_olin()
print('Moving a bike to Olin')
plot_state()
bikeshare
# -
# We can wrap that code in a function called `step` that simulates one time step. In any given minute, a student might ride from Olin to Wellesley, from Wellesley to Olin, or both, or neither, depending on the results of `flip`.
def step():
if flip(0.7):
bike_to_wellesley()
print('Moving a bike to Wellesley')
if flip(0.6):
bike_to_olin()
print('Moving a bike to Olin')
# If you run `step` a few times, it should update the current figure. In each time step, the number of bikes at each location might go up, down, or stay the same.
step()
plot_state()
bikeshare
# The following function labels the axes and adds a legend to the figure.
def decorate():
legend(loc='random string')
label_axes(title='Olin-Wellesley Bikeshare',
xlabel='Time step (min)',
ylabel='Number of bikes')
# As always, when you define a function, it has no effect until you run it.
decorate()
# **Exercise:** Change the argument of `legend` to `'random string'` and run `decorate` again. You should get an error message that lists the valid location where you can put the legend.
# ## Optional parameters
# Again let's start with a new `System` object and a new plot.
bikeshare = System(olin=10, wellesley=2)
newfig()
plot_state()
# We can make `step` more general by adding parameters. Because these parameters have default values, they are optional.
def step(p1=0.5, p2=0.5):
print('p1 ->', p1)
print('p2 ->', p2)
if flip(p1):
bike_to_wellesley()
if flip(p2):
bike_to_olin()
# I added print statements, so each time we run `step` we can see the arguments.
#
# If you provide no arguments, you get the default values:
step()
plot_state()
# If you provide one argument, it overrides the first parameter.
step(0.4)
plot_state()
# If you provide two arguments, they override both.
step(0.4, 0.2)
plot_state()
# You can specify the names of the parameters you want to override.
step(p1=0.4, p2=0.2)
plot_state()
# Which means you can override the second parameter and use the default for the first.
step(p2=0.2)
plot_state()
# You can combine both forms, but it is not very common:
step(0.4, p2=0.2)
plot_state()
# One reason it's not common is that it's error prone. The following example causes an error.
# +
# If you remove the # at the beginning of the next line and run it, you get
# SyntaxError: positional argument follows keyword argument
step(p1=0.4, 0.2)
# -
# From the error message, you might infer that arguments like `step(0.4, 0.2)` are called "positional" and arguments like `step(p1=0.4, p2=0.2)` are called "keyword arguments".
# **Exercise:** Write a version of `decorate` that takes an optional parameter named `loc` with default value `'best'`. It should pass the value of `loc` along as an argument to `legend.` Test your function with different values of `loc`. [You can see the list of legal values here](https://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.legend).
# Solution goes here
def decorate(loc="best"):
legend(loc=loc)
label_axes(title='Olin-Wellesley Bikeshare',
xlabel='Time step (min)',
ylabel='Number of bikes')
# Solution goes here
decorate("lower right")
plot_state()
decorate("center left")
plot_state()
# ## For loop
# Before we go on, I'll redefine `step` without the print statements.
def step(p1=0.5, p2=0.5):
if flip(p1):
bike_to_wellesley()
if flip(p2):
bike_to_olin()
# And let's start again with a new `System` object and a new figure.
bikeshare = System(olin=10, wellesley=2)
newfig()
plot_state()
decorate()
# We can use a for loop to move 4 bikes from Olin to Wellesley.
for i in range(4):
bike_to_wellesley()
plot_state()
# Or we can simulate 4 random time steps.
for i in range(4):
step()
plot_state()
# If each step corresponds to a minute, we can simulate the rest of the hour like this.
for i in range(52):
step(p1=0.4, p2=0.2)
plot_state()
# **Exercise:** Combine the examples from the previous two sections to write a function named `run_steps` that takes three parameters, named `num_steps`, `p1`, and `p2`. It should use a for loop to run `step` the number of times specified by `num_steps`, passing along the specified values of `p1` and `p2`. After each step, it should plot the updated state.
#
# Test your function by creating a new `System` object, creating a new figure, and running `run_steps`.
# Solution goes here
def run_steps(num_steps, p1, p2):
"""
Simulates a number of steps with specified probabilities for bike movements between Wellesley and Olin and plots changes.
num_steps : The number of steps that the program should run for.
p1 : The probability (should be < 1) that a student bikes from Olin to Wellesley,
p2 : The probability (should be < 1) that a student bikes from Wellesley to Olin.
"""
for i in range(num_steps):
step(p1, p2)
plot_state()
# Solution goes here
bikeshare = System(olin=10, wellesley=2)
newfig()
plot_state()
decorate()
run_steps(50, .4, .3)
| code/chap01mine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.12 64-bit (''codeforecon'': conda)'
# name: python3
# ---
# (regression)=
# # Regression
# ## Introduction
#
# In this chapter, you'll learn how to run linear regressions with code.
#
# If you're running this code (either by copying and pasting it, or by downloading it using the icons at the top of the page), you may need to install the packages it uses first. There's a brief guide to installing packages in the Chapter on {ref}`code-preliminaries`.
#
# Most of this chapter will rely on [statsmodels](https://www.statsmodels.org/stable/index.html) with some use of [**linearmodels**](https://bashtage.github.io/linearmodels/). Some of the material in this chapter follows [Grant McDermott](https://grantmcdermott.com/)'s excellent notes and the [Library of Statistical Translation](https://lost-stats.github.io/).
#
# ### Notation and basic definitions
#
# Greek letters, like $\beta$, are the truth and represent parameters. Modified Greek letters are an estimate of the truth, for example $\hat{\beta}$. Sometimes Greek letters will stand in for vectors of parameters. Most of the time, upper case Latin characters such as $X$ will represent random variables (which could have more than one dimension). Lower case letters from the Latin alphabet denote realised data, for instance $x$ (which again could be multi-dimensional). Modified Latin alphabet letters denote computations performed on data, for instance $\bar{x} = \frac{1}{n} \displaystyle\sum_{i} x_i$ where $n$ is number of samples.
#
# Ordinary least squares (OLS) regression can be used to *estimate* the parameters of certain types of model, most typically models of the form
#
# $$
# y = \beta_0 + \beta_1 \cdot x_1 + \beta_2 \cdot x_2
# $$
#
# This generic model says that the value of an outcome variable $y$ is a linear function of one or more input predictor variables $x_i$, where the $x_i$ could be transforms of original data. But the above equation is a platonic ideal, what we call a data generating process (DGP). OLS allows us to recover *estimates* of the parameters of the model , i.e. to find $\hat{\beta_i}$ and to enable us to write an estimated model:
#
# $$
# y = \hat{\beta_0} + \hat{\beta_1} \cdot x_1 + \hat{\beta_2} \cdot x_2 + \epsilon
# $$
#
# This equation can also be expressed in matrix form as
#
# $$
# y = x'\cdot \hat{\beta} + \epsilon
# $$
#
# where $x' = (1, x_1, \dots, x_{n})'$ and $\hat{\beta} = (\hat{\beta_0}, \hat{\beta_1}, \dots, \hat{\beta_{n}})$.
#
# Given data $y_i$ stacked to make a vector $y$ and $x_{i}$ stacked to make a matrix $X$, this can be solved for the coefficients $\hat{\beta}$ according to
#
# $$
# \hat{\beta} = \left(X'X\right)^{-1} X'y
# $$
#
# To be sure that the estimates of these parameters are the *best linear unbiased estimate*, a few conditions need to hold: the Gauss-Markov conditions:
#
# 1. $y$ is a linear function of the $\beta_i$
# 2. $y$ and the $x_i$ are randomly sampled from the population.
# 3. There is no perfect multi-collinearity of variables.
# 4. $\mathbb{E}(\epsilon | x_1, \dots, x_n) = 0$ (unconfoundedness)
# 5. $\text{Var}(\epsilon | x_1, \dots, x_n) = \sigma^2$ (homoskedasticity)
#
# (1)-(4) also guarantee that OLS estimates are unbiased and $\mathbb{E}(\hat{\beta}_i) = \beta_i$.
#
# The classic linear model requires a 6th assumption; that $\epsilon \thicksim \mathcal{N}(0, \sigma^2)$.
#
# The interpretation of regression coefficients depends on what their units are to begin with, but you can always work it out by differentiating both sides of the model equation with respect to the $x_i$. For example, for the first model equation above
#
# $$
# \frac{\partial y}{\partial x_i} = \beta_i
# $$
#
# so we get the interpretation that $\beta_i$ is the rate of change of y with respect to $x_i$. If $x_i$ and $y$ are in levels, this means that a unit increase in $x_i$ is associated with a $\beta_i$ units increase in $y$. If the right-hand side of the model is $\ln x_i$ then we get
#
# $$
# \frac{\partial y}{\partial x_i} = \beta_i \frac{1}{x_i}
# $$
#
# with some abuse of notation, we can rewrite this as $\partial y = \beta_i \partial x_i/x_i$, which says that a percent change in $x_i$ is associated with a $\beta_i$ unit change in $y$. With a logged $y$ variable, it's a percent change in $x_i$ that is associated with a percent change in $y$, or $\partial y/y = \beta_i \partial x_i/x_i$ (note that both sides of this equation are unitless in this case). Finally, another example that is important in practice is that of log differences, eg $y = \beta_i (\ln x_i - \ln x_i')$. Again, we will abuse notation and say that this case may be represented as $\partial y = \beta_i (\partial x_i/x_i - \partial x_i'/x_i')$, i.e. the difference in two percentages, a *percentage point* change, in $x_i$ is associated with a $\beta_i$ unit change in $y$.
#
# ### Imports
#
# Let's import some of the packages we'll be using:
import numpy as np
import scipy.stats as st
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import statsmodels.api as sm
import statsmodels.formula.api as smf
import os
from pathlib import Path
# Set max rows displayed for readability
pd.set_option("display.max_rows", 6)
# Plot settings
plt.style.use(
"https://github.com/aeturrell/coding-for-economists/raw/main/plot_style.txt"
)
# ## Regression basics
#
# There are two ways to run regressions in [**statsmodels**](https://www.statsmodels.org/stable/index.html); passing the data directly as objects, and using formulae. We'll see both but, just to get things started, let's use the formula API.
#
# We'll use the starwars dataset to run a regression of mass on height for star wars characters. This example borrows very heavily from notes by [<NAME>](https://grantmcdermott.com/). First, let's bring the dataset in:
df = pd.read_csv(
"https://github.com/aeturrell/coding-for-economists/raw/main/data/starwars.csv",
index_col=0,
)
# Look at first few rows
df.head()
# Okay, now let's do a regression using OLS and a formula that says our y-variable is mass and our regressor is height:
results = smf.ols("mass ~ height", data=df).fit()
# Well, where are the results!? They're stored in the object we created. To peek at them we need to call the summary function (and, for easy reading, I'll print it out too using `print`)
print(results.summary())
# What we're seeing here are really several tables glued together. To just grab the coefficients in a tidy format, use
results.summary().tables[1]
# You'll have noticed that we got an intercept, even though we didn't specify one in the formula. **statsmodels** adds in an intercept by default because, most of the time, you will want one. To turn it off, add a `-1` at the end of the formula command, eg in this case you would call `smf.ols('mass ~ height -1', data=df).fit()`.
#
# The fit we got in the case with the intercept was pretty terrible; a low $R^2$ and both of our confidence intervals are large and contain zero. What's going on? If there's one adage in regression that's always worth paying attention to, it's *always plot your data*. Let's see what's going on here:
fig, ax = plt.subplots()
sns.scatterplot(data=df, x="height", y="mass", s=200, ax=ax, legend=False, alpha=0.8)
ax.annotate(
"<NAME>",
df.iloc[df["mass"].idxmax()][["height", "mass"]],
xytext=(0, -50),
textcoords="offset points",
arrowprops=dict(
arrowstyle="fancy",
color="k",
connectionstyle="arc3,rad=0.3",
),
)
ax.set_ylim(0, None)
ax.set_title("Always plot the data", loc="left")
plt.show()
# Oh dear, Jabba's been on the paddy frogs again, and he's a bit of different case. When we're estimating statistical relationships, we have all kinds of choices and should be wary about arbitrary decisions of what to include or exclude in case we fool ourselves about the generality of the relationship we are capturing. Let's say we knew that we weren't interested in Hutts though, but only in other species: in that case, it's fair enough to filter out Jabba and run the regression without this obvious outlier. We'll exclude any entry that contains the string 'Jabba' in the `name` column:
results_outlier_free = smf.ols(
"mass ~ height", data=df[~df["name"].str.contains("Jabba")]
).fit()
print(results_outlier_free.summary())
# This looks a lot more healthy. Not only is the model explaining a *lot* more of the data, but the coefficients are now significant.
# ### Robust regression
#
# Filtering out data is one way to deal with outliers, but it's not the only one; an alternative is to use a regression technique that is robust to such outliers. **statsmodels** has a variety of robust linear models that you can read more about [here](https://www.statsmodels.org/stable/examples/notebooks/generated/robust_models_0.html). To demonstrate the general idea, we will run the regression again but using a robust method.
#
#
results_robust = smf.rlm(
"mass ~ height", data=df, M=sm.robust.norms.TrimmedMean(0.5)
).fit()
print(results_robust.summary())
# There are many different 'M-estimators' available; in this case the TrimmedMean estimator gives a very similar result to the regression with the point excluded. We can visualise this, and, well, the results are not really very different in this case. Note that `abline_plot` just takes an intercept and coefficient from a fitted model and renders the line that they encode.
fig, ax = plt.subplots()
ax.scatter(df["height"], df["mass"])
sm.graphics.abline_plot(model_results=results_robust, ax=ax, alpha=0.5, label="Robust")
sm.graphics.abline_plot(
model_results=results, ax=ax, color="red", label="OLS", alpha=0.5, ls="--"
)
ax.legend()
ax.set_xlabel("Height")
ax.set_ylabel("Mass")
ax.set_ylim(0, None)
plt.show()
# ### Standard errors
#
# You'll have seen that there's a column for the standard error of the estimates in the regression table and a message saying that the covariance type of these is 'nonrobust'. Let's say that, instead, we want to use Eicker-Huber-White robust standard errors, aka "HC2" standard errors. We can specify to use these up front standard errors up front in the fit method:
(smf.ols("mass ~ height", data=df).fit(cov_type="HC2").summary().tables[1])
# Or, alternatively, we can go back to our existing results and recompute the results from those:
print(results.get_robustcov_results("HC2").summary())
# There are several different types of standard errors available in **statsmodels**:
#
# - ‘HC0’, ‘HC1’, ‘HC2’, and ‘HC3’
# - ‘HAC’, for heteroskedasticity and autocorrelation consistent standard errors, for which you may want to also use some keyword arguments
# - 'hac-groupsum’, for Driscoll and Kraay heteroscedasticity and
# autocorrelation robust standard errors in panel data, again for which you may have to specify extra keyword arguments
# - 'hac-panel’, for heteroscedasticity and autocorrelation robust standard
# errors in panel data, again with keyword arguments; and
# - 'cluster' for clustered standard errors.
#
# You can find information on all of these [here](https://www.statsmodels.org/stable/generated/statsmodels.regression.linear_model.OLSResults.get_robustcov_results.html?highlight=get_robustcov_results#statsmodels.regression.linear_model.OLSResults.get_robustcov_results). For more on standard errors in python, [this is a good](http://www.vincentgregoire.com/standard-errors-in-python/) link.
#
# For now, let's look more closely at those last ones: clustered standard errors.
#
#
# #### Clustered standard errors
#
# Often, we know something about the structure of likely errors, namely that they occur in groups. In the below example we use one-way clusters to capture this effect in the errors.
#
# Note that in the below example, we grab a subset of the data for which a set of variables we're interested in are defined, otherwise the below example would execute with an error because of missing cluster-group values.
xf = df.dropna(subset=["homeworld", "mass", "height", "species"])
results_clus = smf.ols("mass ~ height", data=xf).fit(
cov_type="cluster", cov_kwds={"groups": xf["homeworld"]}
)
print(results_clus.summary())
# We can add two-way clustering of standard errors using the following:
xf = df.dropna(subset=["homeworld", "mass", "height", "species"])
two_way_clusters = np.array(xf[["homeworld", "species"]], dtype=str)
results_clus = smf.ols("mass ~ height", data=xf).fit(
cov_type="cluster", cov_kwds={"groups": two_way_clusters}
)
print(results_clus.summary())
# As you would generally expect, the addition of clustering has increased the standard errors.
# ## Fixed effects and categorical variables
#
# Fixed effects are a way of allowing the intercept of a regression model to vary freely across individuals or groups. It is, for example, used to control for any individual-specific attributes that do not vary across time in panel data.
#
# Let's use the 'mtcars' dataset to demonstrate this. We'll read it in and set the datatypes of some of the columns at the same time.
mpg = pd.read_csv(
"https://raw.githubusercontent.com/LOST-STATS/lost-stats.github.io/source/Data/mtcars.csv",
dtype={"model": str, "mpg": float, "hp": float, "disp": float, "cyl": "category"},
)
mpg.head()
# Now we have our data in we want to regress mpg (miles per gallon) on hp (horsepower) with fixed effects for cyl (cylinders). Now we *could* just pop in a formula like this `'mpg ~ hp + cyl'` because we took the trouble to declare that `cyl` was of datatype category when reading it in from the csv file. This means that statsmodels will treat it as a category and use it as a fixed effect by default.
#
# But when I read that formula I get nervous that `cyl` might not have been processed correctly (ie it could have been read in as a float, which is what it looks like) and it might just be treated as a float (aka a continuous variable) in the regression. Which is not what we want at all. So, to be safe, and make our intentions explicit (even when the data is of type 'category'), it's best to use the syntax `C(cyl)` to ask for a fixed effect.
#
# Here's a regression which does that:
results_fe = smf.ols("mpg ~ hp + C(cyl)", data=mpg).fit()
print(results_fe.summary())
# We can see here that two of the three possible values of `cyl`:
mpg["cyl"].unique()
# have been added as fixed effects regressors. The way that `+C(cyl)` has been added makes it so that the coefficients given are relative to the coefficient for the intercept. We can turn the intercept off to get a coefficient per unique `cyl` value:
print(smf.ols("mpg ~ hp + C(cyl) -1", data=mpg).fit().summary().tables[1])
# When there is an intercept, the coefficients of fixed effect variables can be interpreted as being the average of $y$ for that class *compared* to the excluded classes holding all other categories and variables fixed.
# ### High dimensional fixed effects, aka absorbing regression
#
# Sometimes, you just have a LOT of fixed effects (and perhaps you don't particularly care about them individually). A common example is having a large number of firms as part of a panel. Fortunately, there are ways to make regression with high dimensional fixed effects be both fast and concise. (In Stata, this is provided by the `reghdfe` package.) Here, we will use the [**linearmodels**](https://bashtage.github.io/linearmodels/index.html) package, which is built on top of **statsmodels**.
#
# Let's say we have a regression of the form
#
# $$
# y_i = x_i\cdot \beta + z_i\cdot \gamma +\epsilon_i
# $$
#
# where $y_i$ are observations indexed by $i$, $x_i$ are vectors of exogenous variables we care about the coefficients ($\beta$), $z_i$ are vectors of fixed effects we don't care too much about the coefficients (\gamma) for, and the $\epsilon_i$ are errors. Then we can use an *absorbing regression* to solve for the $\beta$ while ignoring the $\gamma$.
#
# Here's an example using simulated data on workers taken from the **linearmodels** docs. Let's simulate some data first, with two fixed effects (state and firm) alongside the two exogenous variables we're interested in.
# +
from numpy.random import default_rng
rng = default_rng() # Random number generator
# Create synthetic input data
nobs = 1_000_000 # No. observations
state_id = rng.integers(50, size=nobs) # State identifier
firm_id = rng.integers(nobs // 5, size=nobs) # Firm identifier (mean of 5 workers/firm)
x = rng.standard_normal((nobs, 2)) # Exogenous variables
sim = pd.DataFrame(
{
"state_id": pd.Categorical(state_id),
"firm_id": pd.Categorical(firm_id),
"exog_0": x[:, 0],
"exog_1": x[:, 1],
}
)
# Create synthetic relationship
beta = [1, 3] # coefficients of interest
state_effects = rng.standard_normal(state_id.max() + 1)
state_effects = state_effects[state_id] # Generate state fixed effects
firm_effects = rng.standard_normal(firm_id.max() + 1)
firm_effects = firm_effects[firm_id] # Generate firm fixed effects
eps = rng.standard_normal(nobs) # Generate errors
# Generate endogeneous outcome variable
sim["y"] = (
sim["exog_0"] * beta[0]
+ sim["exog_1"] * beta[1]
+ firm_effects
+ state_effects
+ eps
)
sim.head()
# -
# Now we pass this to **linearmodels** and with the `state_id` and `firm_id` variables entered via the `absorb` keyword argument:
# +
from linearmodels.iv.absorbing import AbsorbingLS
mod = AbsorbingLS(
sim["y"], sim[["exog_0", "exog_1"]], absorb=sim[["state_id", "firm_id"]]
)
print(mod.fit())
# -
# So, from our 1,000,000 observations, we have roughly 200,000 fixed effects that have been scooped up and packed away, leaving us with just the coefficients, $\beta$, on the exogenous variables of interest.
# ## Transformations of regressors
#
# This chapter is showcasing *linear* regression. What that means is that the model is linear in the regressors: but it doesn't mean that those regressors can't be some kind of (potentially non-linear) transform of the original features $x_i$.
# ### Logs and arcsinh
#
# You have two options for adding in logs: do them before, or do them in the formula. Doing them before just makes use of standard dataframe operations to declare a new column:
#
mpg["lnhp"] = np.log(mpg["hp"])
print(smf.ols("mpg ~ lnhp", data=mpg).fit().summary().tables[1])
# Alternatively, you can specify the log directly in the formula:
results_ln = smf.ols("mpg ~ np.log(hp)", data=mpg).fit()
print(results_ln.summary().tables[1])
# Clearly, the first method will work for `arcsinh(x)` and `log(x+1)`, but you can also pass both of these into the formula directly too. (For more on the pros and cons of arcsinh, see {cite}`bellemare2020elasticities`.) Here it is with arcsinh:
print(smf.ols("mpg ~ np.arcsinh(hp)", data=mpg).fit().summary().tables[1])
# ### Interaction terms and powers
#
# This chapter is showcasing *linear* regression. What that means is that the model is linear in the regressors: but it doesn't mean that those regressors can't be some kind of non-linear transform of the original features $x_i$. Two of the most common transformations that you might want to use are *interaction terms* and *polynomial terms*. An example of an interaction term would be
#
# $$
# y = \beta_0 + \beta_1 x_1 \cdot x_2
# $$
#
# while an example of a polynomial term would be
#
# $$
# y = \beta_0 + \beta_1 x_1^2
# $$
#
# i.e. the last term enters only after it is multiplied by itself.
#
# One note of warning: the interpretation of the effect of a variable is no longer as simple as was set out at the start of this chapter. To work out *what* the new interpretation is, the procedure is the same though: just take the derivative. In the case of the interaction model above, the effect of a unit change in $x_1$ on $y$ is now going to be a function of $x_2$. In the case of the polynomial model above, the effect of a unit change in $x_1$ on $y$ will be $2\beta_1 \cdot x_1$. For more on interaction terms, see {cite}`balli2013interaction`.
#
# Alright, with all of that preamble out of the way, let's see how we actual do some of this! Let's try including a linear and squared term in the regression of `mpg` on `hp` making use of the numpy power function:
res_poly = smf.ols("mpg ~ hp + np.power(hp, 2)", data=mpg).fit()
print(res_poly.summary().tables[1])
# Now let's include the original term in hp, a term in disp, and the interaction between them, which is represented by hp:disp in the table.
res_inter = smf.ols("mpg ~ hp * disp", data=mpg).fit()
print(res_inter.summary().tables[1])
# In the unusual case that you want *only* the interaction term, you write it as it appears in the table above:
print(smf.ols("mpg ~ hp : disp", data=mpg).fit().summary().tables[1])
# ## The formula API explained
#
# As you will have seen `~` separates the left- and right-hand sides of the regression. `+` computes a set union, which will also be familiar from the examples above (ie it inludes two terms as long as they are distinct). `-` computes a set difference; it adds the set of terms to the left of it while removing any that appear on the right of it. As we've seen, `a*b` is a short-hand for `a + b + a:b`, with the last term representing the interaction. `/` is short hand for `a + a:b`, which is useful if, for example `b` is nested within `a`, so it doesn't make sense to control for `b` on its own. Actually, the `:` character can interact multiple terms so that `(a + b):(d + c)` is the same as `a:c + a:d + b:c + b:d`. `C(a)` tells statsmodels to treat `a` as a categorical variable that will be included as a fixed effect. Finally, as we saw above with powers, you can also pass in vectorised functions, such as `np.log` and `np.power`, directly into the formulae.
#
# One gotcha with the formula API is ensuring that you have sensible variable names in your dataframe, i.e. ones that do *not* include whitespace or, to take a really pathological example, have the name 'a + b' for one of the columns that you want to regress on. You can dodge this kind of problem by passing in the variable name as, for example, `Q("a + b")` to be clear that the *column name* is anything within the `Q("...")`.
# ## Multiple regression models
#
# As is so often the case, you're likely to want to run more than one model at once with different specifications. Although there is a base version of this in **statsmodels**, called `summary_col`, which you can find an example of [here](http://aeturrell.com//2018/05/05/running-many-regressions-alongside-pandas/), instead we'll be using the [**stargazer**](https://github.com/mwburke/stargazer) package to assemble the regressions together in a table.
#
# In the above examples, we've collected a few different regression results. Let's put them together:
# +
from stargazer.stargazer import Stargazer
stargazer_tab = Stargazer([results_ln, res_poly, res_inter])
stargazer_tab
# -
# There are lots of customisation options, including ones that add a title, rename variables, add notes, and so on. What is most useful is that as well as the HTML friendly output that you can see above, the package also exports to latex:
print(stargazer_tab.render_latex())
# And of course this can be written to a file using `open('regression.tex', 'w').write(stargazer.render_latex())` where you can get your main latex compilation to scoop it up and use it.
# ## Specifying regressions without formulae, using the array API
#
# As noted, there are two ways to run regressions in [**statsmodels**](https://www.statsmodels.org/stable/index.html); passing the data directly as objects, and using formulae. We've seen the formula API, now let's see how to specify regressions using arrays with the format `sm.OLS(y, X)`.
#
# We will first need to take the data out of the **pandas** dataframe and put it into a couple of arrays. When we're not using the formula API, the default is to treat the array X as the design matrix for the regression-so, if it doesn't have a column of constants in, there will be no intercept in the regression. Therefore, we need to add a constant vector to the matrix `X` if we *do* want an intercept. Use `sm.add_constant(X)` for this.
X = np.array(xf["height"])
y = np.array(xf["mass"])
X = sm.add_constant(X)
results = sm.OLS(y, X).fit()
print(results.summary())
#
# This approach seems a lot less convenient, not to mention less clear, so you may be wondering when it is useful. It's useful when you want to do many regressions in a systematic way or when you don't know what the columns of a dataset will be called ahead of time. It can actually be a little bit simpler to specify for more complex regressions too.
# ### Fixed effects in the array API
#
# If you're using the formula API, it's easy to turn a regressor `x` into a fixed effect by putting `C(x)` into the model formula, as you'll see in the next section.
#
# For the array API, things are not that simple and you need to use dummy variables. Let's say we have some data like this:
# +
from numpy.random import Generator, PCG64
# Set seed for random numbers
seed_for_prng = 78557
prng = Generator(PCG64(seed_for_prng))
no_obs = 200
X = pd.DataFrame(prng.normal(size=no_obs))
X[1] = prng.choice(["a", "b"], size=no_obs)
# Get this a numpy array
X = X.values
# Create the y data, adding in a bit of noise
y = X[:, 0] * 2 + 0.5 + prng.normal(scale=0.1, size=no_obs)
y = [el_y + 1.5 if el_x == "a" else el_y + 3.4 for el_y, el_x in zip(y, X[:, 1])]
X[:5, :]
# -
# The first feature (column) is of numbers and it's clear how we include it. The second, however, is a grouping that we'd like to include as a fixed effect. But if we just throw this matrix into `sm.OLS(y, X)`, we're going to get trouble because **statsmodels** isn't sure what to do with a vector of strings. So, instead, we need to create some dummy variables out of our second column of data
#
# Astonishingly, there are several popular ways to create dummy variables in Python: **scikit-learn**'s `OneHotEncoder` and **pandas**' `get_dummies` being my favourites. Let's use the latter here.
pd.get_dummies(X[:, 1])
# We just need to pop this into our matrix $X$:
X = np.column_stack([X[:, 0], pd.get_dummies(X[:, 1])])
X = np.array(X, dtype=float)
X[:5, :]
# Okay, so now we're ready to do our regression:
print(sm.OLS(y, X).fit().summary())
# Perhaps you can see why I generally prefer the formula API...
# ## Instrumental variables
#
# Rather than use **statsmodels** for IV, we'll use the [**linearmodels**](https://bashtage.github.io/linearmodels/doc/index.html) package, which has very clean documentation (indeed, this sub-section is indebted to that documentation).
#
#
# Recall that a good instrumental variable $z$ has zero covariance with the error from the regression (which is untestable) and non-zero covariance with the variable of interest (which is).
#
#
# Recall that in IV regression, we have a model of the form
#
# $$
# \begin{split}y_i & = x_{1i}\hat{\beta_1} + x_{2i}\hat{\beta_2} + \epsilon_i \\
# x_{2i} & = z_{1i}\hat{\delta} + z_{2i}\hat{\gamma} + \nu_i\end{split}
# $$
#
# where $x_{1i}$ is a set of $k_1$ exogenous regressors and $x_{2i}$ is a set of $k_2$ endogenous regressors such that $\text{Cov}(x_{2i}, \epsilon_i)\neq 0$. This is a problem for the usual OLS assumptions (the right-hand side should be exogenous).
#
#
# To get around this, in 2-stage least squares IV, we first regress $x_{2i}$ on instruments that explain $x_{2i}$ *but not* $y_i$, and then regress $y_i$ only on the predicted/estimated left-hand side from the first regression, ie on $\hat{x_{2i}}$. There are other estimators than IV2SLS, but I think that one has the most intuitive explanation of what's going.
#
# As well as a 2-stage least squares estimator called `IV2SLS`, **linearmodels** has a Limited Information Maximum Likelihood (LIML) estimator `IVLIML`, a Generalized Method of Moments (GMM) estimator `IVGMM`, and a Generalized Method of Moments using the Continuously Updating Estimator (CUE) `IVGMMCUE`.
#
# Just as with OLS via **statsmodels**, there's an option to use an array API for the **linearmodels** IV methods.
#
# It's always easiest to see an example, so let's estimate what might cause (realised) cigarette demand for the 48 continental US states in 1995 with `IV2SLS`. First we need to import the estimator, `IV2SLS`, and the data:
# +
from linearmodels.iv import IV2SLS
df = pd.read_csv(
"https://vincentarelbundock.github.io/Rdatasets/csv/AER/CigarettesSW.csv",
dtype={"state": "category", "year": "category"},
).assign(
rprice=lambda x: x["price"] / x["cpi"],
rincome=lambda x: x["income"] / x["population"] / x["cpi"],
)
df.head()
# -
# Now we'll specify the model. It's going to be in the form `dep ~ exog + [endog ~ instruments]`, where endog will be regressed on instruments and dep will be regressed on both exog and the predicted values of endog.
#
# In this case, the model will be
#
# $$
# \text{Price}_i = \hat{\pi_0} + \hat{\pi_1} \text{SalesTax}_i + v_i
# $$
#
# in the first stage regression and
#
# $$
# \text{Packs}_i = \hat{\beta_0} + \hat{\beta_2}\widehat{\text{Price}_i} + \hat{\beta_1} \text{RealIncome}_i + u_i
# $$
#
# in the second stage.
results_iv2sls = IV2SLS.from_formula(
"np.log(packs) ~ 1 + np.log(rincome) + C(year) + C(state) + [np.log(rprice) ~ taxs]",
df,
).fit(cov_type="clustered", clusters=df["year"])
print(results_iv2sls.summary)
# We sort of skipped a step here and did everything all in one go. If we *did* want to know how our first stage regression went, we can just pass a formula to `IV2SLS` without the part in square brackets, `[...]`, and it will run regular OLS.
#
# But, in this case, there's an easier way: we can print out a set of handy 1st stage statistics from running the full model.
#
print(results_iv2sls.first_stage)
# There are more tests and checks available. For example, Wooldridge’s regression test of exogeneity uses regression residuals from the endogenous variables regressed on the exogenous variables and the instrument to test for endogenity and is available to run on fitted model results. Let's check that:
results_iv2sls.wooldridge_regression
# We can compare the IV results against (naive) OLS. First, run the OLS equivalent:
res_cig_ols = IV2SLS.from_formula(
"np.log(packs) ~ 1 + np.log(rincome) + C(year) + C(state) + np.log(rprice)", df
).fit(cov_type="clustered", clusters=df["year"])
# Now select these two models to compare:
# +
from collections import OrderedDict
from linearmodels.iv.results import compare
res = OrderedDict()
res["OLS"] = res_cig_ols
res["2SLS"] = results_iv2sls
print(compare(res))
# -
# Once we take into account the fact that the real price is endogeneous to (realised) demand, we find that its coefficient is more negative; i.e. an increase in the real price of cigarettes creates a bigger fall in number of packs bought.
# ## Logit, probit, and generalised linear models
#
# ### Logit
#
# A logistical regression, aka a logit, is a statistical method for a best-fit line between a regressors $X$ and an outcome varibale $y$ that takes on values in $(0, 1)$.
#
# The function that we're assuming links the regressors and the outcome has a few different names but the most common is the sigmoid function or the logistic function. The data generating process is assumed to be
#
# $$
# {\displaystyle \mathbb{P}(Y=1\mid X) = \frac{1}{1 + e^{-X'\beta}}}
# $$
#
# we can also write this as $\ln\left(\frac{p}{p-1}\right) = \beta_0 + \sum_i \beta_i x_i$ to get a 'log-odds' relationship. The coefficients from a logit model do not have the same interpration as in an OLS estimation, and you can see this from the fact that $\partial y/\partial x_i \neq \beta_i$ for logit. Of course, you can work out what the partial derivative is for yourself but most packages offer a convenient way to quickly recover the marginal effects.
#
# Logit models are available in **scikit-learn** and **statsmodels** but bear in mind that the **scikit-learn** logit model is, ermm, extremely courageous in that regularisation is applied by default. If you don't know what that means, don't worry, but it's probably best to stick with **statsmodels** as we will do in this example.
#
# We will predict a target `GRADE`, representing whether a grade improved or not, based on some regressors including participation in a programme.
# Load the data from Spector and Mazzeo (1980)
df = sm.datasets.spector.load_pandas().data
# Look at info on data
print(sm.datasets.spector.NOTE)
res_logit = smf.logit("GRADE ~ GPA + TUCE + PSI", data=df).fit()
print(res_logit.summary())
# So, did participation (`PSI`) help increase a grade? Yes. But we need to check the marginal effect to say exactly how much. We'll use `get_margeff` to do this, we'd like the $dy/dx$ effect, and we'll take it at the mean of each regressor.
marg_effect = res_logit.get_margeff(at="mean", method="dydx")
marg_effect.summary()
# So participation gives almost half a grade increase.
# ### Probit
#
# Probit is very similar to logit: it's a statistical method for a best-fit line between regressors $X$ and an outcome varibale $y$ that takes on values in $(0, 1)$. And, just like with logit, the function that we're assuming links the regressors and the outcome has a few different names!
#
# The data generating process is assumed to be
#
# $$
# {\displaystyle \mathbb{P}(Y=1\mid X)=\Phi (X^{T}\beta )}
# $$
#
# where
#
# $$
# {\displaystyle \Phi (x)={\frac {1}{\sqrt {2\pi }}}\int _{-\infty }^{x}e^{-{\frac {y^{2}}{2}}}dy.}
# $$
#
# is the cumulative standard normal (aka Gaussian) distribution. The coefficients from a probit model do not have the same interpration as in an OLS estimation, and you can see this from the fact that $\partial y/\partial x_i \neq \beta_i$ for probit. And, just as with logit, although you can derive the marginal effects, most packages offer a convenient way to quickly recover them.
#
# We can re-use our previous example of predicting a target `GRADE`, representing whether a grade improved or not, based on some regressors including participation (PSI) in a programme.
res_probit = smf.probit("GRADE ~ GPA + TUCE + PSI", data=df).fit()
print(res_probit.summary())
p_marg_effect = res_probit.get_margeff(at="mean", method="dydx")
p_marg_effect.summary()
# It's no coincidence that we find very similar results here because the two functions we're using don't actually look all that different:
# +
import scipy.stats as st
fig, ax = plt.subplots()
support = np.linspace(-6, 6, 1000)
ax.plot(support, st.logistic.cdf(support), "r-", ls="--", label="Logistic")
ax.plot(support, st.norm.cdf(support), label="Probit")
ax.legend()
ax.set_ylim(0, None)
ax.set_ylim(0, None)
plt.show()
# -
# What difference there is, is that logistic regression puts more weight into the tails of the distribution. Arguably, logit is easier to interpret too. With logistic regression, a one unit change in $x_i$ is associated with a $\beta_i$ change in the log odds of a 1 outcome or, alternatively, an $e^{\beta_i}$-fold change in the odds, all else being equal. With a probit, this is a change of $\beta_i z$ for $z$ a normalised variable that you'd have to convert into a predicted probability using the normal CDF.
# ### Generalised linear models
#
# Logit and probit (and OLS for that matter) as special cases of a class of models such that $g$ is a 'link' function connects a function of regressors to the output, and $\mu$ is the mean of a conditional response distribution at a given point in the space of regressors. When $g(\mu) = X'\beta$, we just get regular OLS. When it's logit, we have
#
# $$
# {\displaystyle \mu= \mathbb{E}(Y\mid X=x) =g^{-1}(X'\beta)= \frac{1}{1 + e^{-X'\beta}}.}
# $$
#
# But as well as the ones we've seen, there are many possible link functions one can use via the catch-all `glm` function. These come in different 'families' of distributions, with the default for the binomial family being logit. So, running `smf.glm('GRADE ~ GPA + TUCE + PSI', data=df, family=sm.families.Binomial()).fit()` will produce exactly the same as we got both using the `logit` function. For more on the families of distributions and possible link functions, see the [relevant part](https://www.statsmodels.org/stable/glm.html#) of the **statsmodels** documentation.
#
# If you need a library dedicated to GLMs that has all the bells and whistles you can dream of, you might want to check out [glum](https://glum.readthedocs.io/en/latest/). At the time of writing, it is [faster](https://glum.readthedocs.io/en/latest/benchmarks.html) than either GLMnet or H2O (two other popular GLM libraries).
# ## Linear probability model
#
# When $y$ takes values in $\{0, 1\}$ but the model looks like
#
# $$
# y = x' \cdot \beta
# $$
#
# and is estimated by OLS then you have a linear probability model. In this case, the interpretion of a unit change in $x_i$ is that it induces a $\beta_i$ *change in probability* of $y$. Note that homoskedasticity does not hold for the linear probability model.
# ## Violations of the classical linear model (CLM)
#
# ### Heteroskedasticity
#
# If an estimated model is homoskedastic then its random variables have equal (finite) variance. This is also known as homogeneity of variance. Another way of putting it is that, for all *observations* $i$ in an estimated model $y_i = X_i\hat{\beta} + \epsilon_i$ then
#
# $$
# \mathbb{E}(\epsilon_i \epsilon_i) = \sigma^2
# $$
#
# When this relationship does not hold, an estimated model is said to be heteroskedastic.
#
# To test for heteroskedasticity, you can use **statsmodels**' versions of the [Breusch-Pagan](https://www.statsmodels.org/stable/generated/statsmodels.stats.diagnostic.het_breuschpagan.html#statsmodels.stats.diagnostic.het_breuschpagan) or [White](https://www.statsmodels.org/stable/generated/statsmodels.stats.diagnostic.het_white.html#statsmodels.stats.diagnostic.het_white) tests with the null hypothesis that the estimated model is homoskedastic. If the null hypothesis is rejected, then standard errors, t-statistics, and F-statistics are invalidated. In this case, you will need HAC (heteroskedasticity and auto-correlation consistent) standard errors, t- and F-statistics.
#
# To obtain HAC standard errors from existing regression results in a variable `results`, you can use (for 1 lag):
#
# ```python
# results.get_robustcov_results('HAC', maxlags=1).summary()
# ```
# ## Quantile regression
#
# Quantile regression estimates the conditional quantiles of a response variable. In some cases, it can be more robust to outliers and, in the case of the $q=0.5$ quantile it is equivalent LAD (Least Absolute Deviation) regression. Let's look at an example of quantile regression in action, lifted direct from the **statsmodels** [documentation](https://www.statsmodels.org/dev/examples/notebooks/generated/quantile_regression.html) and based on a Journal of Economic Perspectives paper by <NAME> Hallock.
df = sm.datasets.engel.load_pandas().data
df.head()
# What we have here are two sets of related data. Let's perform several quantile regressions from 0.1 to 0.9 in steps of 0.1
mod = smf.quantreg("foodexp ~ income", df)
quantiles = np.arange(0.1, 1.0, 0.1)
q_results = [mod.fit(q=x) for x in quantiles]
# The $q=0.5$ entry will be at the `4` index; let's take a look at it:
print(q_results[4].summary())
# Let's take a look at the results for all of the regressions *and* let's add in OLS for comparison:
# +
ols_res = smf.ols("foodexp ~ income", df).fit()
get_y = lambda a, b: a + b * x
x = np.arange(df.income.min(), df.income.max(), 50)
# Just to make the plot clearer
x_max = 3000
x = x[x < x_max]
fig, ax = plt.subplots()
df.plot.scatter(
ax=ax, x="income", y="foodexp", alpha=0.7, s=10, zorder=2, edgecolor=None
)
for i, res in enumerate(q_results):
y = get_y(res.params["Intercept"], res.params["income"])
ax.plot(x, y, color="grey", lw=0.5, zorder=0, linestyle=(0, (5, 10)))
ax.annotate(f"$q={quantiles[i]:1.1f}$", xy=(x.max(), y.max()))
y = get_y(ols_res.params["Intercept"], ols_res.params["income"])
ax.plot(x, y, color="red", label="OLS", zorder=0)
ax.legend()
ax.set_xlim(0, x_max)
plt.show()
# -
# This chart shows very clearly how quantile regression differs from OLS. The line fitted by OLS is trying to be all things to all points whereas the line fitted by quantile regression is focused only on its quantile. You can also see how points far from the median (not all shown) may be having a large influence on the OLS line.
# ## Rolling and recursive regressions
#
# Rolling ordinary least squares applies OLS (ordinary least squares) across a fixed window of observations and then rolls (moves or slides) that window across the data set. They key parameter is `window`, which determines the number of observations used in each OLS regression. Recursive regression is equivalent to rolling regression but with a window that expands over time.
#
# Let's first create some synthetic data to perform estimation on:
# +
from statsmodels.regression.rolling import RollingOLS
import statsmodels.api as sm
from sklearn.datasets import make_regression
X, y = make_regression(n_samples=200, n_features=2, random_state=0, noise=4.0, bias=0)
df = pd.DataFrame(X).rename(columns={0: "feature0", 1: "feature1"})
df["target"] = y
df.head()
# -
# Now let's fit the model using a formula and a `window` of 25 steps.
roll_reg = RollingOLS.from_formula(
"target ~ feature0 + feature1 -1", window=25, data=df
)
model = roll_reg.fit()
# Note that -1 in the formala suppresses the intercept. We can see the parameters using `model.params`. Here are the params for time steps between 20 and 30:
model.params[20:30]
# Note that there aren't parameters for entries between 0 and 23 because our window is 25 steps wide. We can easily look at how any of the coefficients are changing over time. Here's an example for 'feature0'.
fig = model.plot_recursive_coefficient(variables=["feature0"])
plt.xlabel("Time step")
plt.ylabel("Coefficient value")
plt.show()
# A rolling regression with an *expanding* rather than *moving* window is effectively a recursive least squares model. We can do this instead using the `RecursiveLS` function from **statsmodels**. Let's fit this to the whole dataset:
reg_rls = sm.RecursiveLS.from_formula("target ~ feature0 + feature1 -1", df)
model_rls = reg_rls.fit()
print(model_rls.summary())
# But now we can look back at how the values of the coefficients changed over time too:
fig = model_rls.plot_recursive_coefficient(
range(reg_rls.k_exog), legend_loc="upper right"
)
ax_list = fig.axes
for ax in ax_list:
ax.set_xlim(0, None)
ax_list[-1].set_xlabel("Time step")
ax_list[0].set_title("Coefficient value");
# ## Regression plots
#
# **statsmodels** has a number of built-in plotting methods to help you understand how well your regression is capturing the relationships you're looking for. Let's see a few examples of these using **statsmodels** built-in Statewide Crime data set:
#
#
crime_data = sm.datasets.statecrime.load_pandas()
print(sm.datasets.statecrime.NOTE)
# First, let's look at a Q-Q plot to get a sense of how the variables are distributed. This uses **scipy**'s stats module. The default distribution is normal but you can use any that **scipy** supports.
st.probplot(crime_data.data["murder"], dist="norm", plot=plt);
# Clearly, this is not quite normal and there are some serious outliers in the tails.
#
# Let's run take a look at the unconditional relationship we're interested in: how murder depends on high school graduation. We'll use [**plotnine**](https://plotnine.readthedocs.io/en/stable/index.html)'s `geom_smooth` to do this but bear in mind it will only run a linear model of `'murder ~ hs_grad'` and ignore the other covariates.
# +
from plotnine import *
(
ggplot(crime_data.data, aes(y="murder", x="hs_grad"))
+ geom_point()
+ geom_smooth(method="lm")
)
# -
# We can take into account those other factors by using a partial regression plot that asks what does $\mathbb{E}(y|X)$ look like as a function of $\mathbb{E}(x_i|X)$? (Use `obs_labels=False` to remove data point labels.)
with plt.rc_context({"font.size": 5}):
sm.graphics.plot_partregress(
endog="murder",
exog_i="hs_grad",
exog_others=["urban", "poverty", "single"],
data=crime_data.data,
obs_labels=True,
)
plt.show()
# At this point, the results of the regression are useful context.
results_crime = smf.ols(
"murder ~ hs_grad + urban + poverty + single", data=crime_data.data
).fit()
print(results_crime.summary())
# Putting the multicollinearity problems to one side, we see that the relationship shown in the partial regression plot is also implied by the coefficient on `hs_grad` in the regression table.
# We can also look at an in-depth summary of one exogenous regressor and its relationship to the outcome variable. Each of these types of regression diagnostic are available individually, or for all regressors at once, too. The first panel is the chart we did with **plotnine** rendered differently (and, one could argue, more informatively). Most of the plots below are self-explanatory except for the third one, the CCPR (Component-Component plus Residual) plot. This provides a way to judge the effect of one regressor on the response variable by taking into account the effects of the other independent variables.
# +
fig = plt.figure(figsize=(8, 6), dpi=150)
sm.graphics.plot_regress_exog(results_crime, "hs_grad", fig=fig)
plt.tight_layout()
plt.show()
# -
#
# **statsmodels** can also produce influence plots of the 'externally studentised' residuals vs. the leverage of each observation as measured by the so-called hat matrix $X(X^{\;\prime}X)^{-1}X^{\;\prime}$ (because it puts the 'hat' on $y$). Externally studentised residuals are residuals that are scaled by their standard deviation. High leverage points could exert an undue influence over the regression line, but only if the predicted $y$ values of a regression that was fit with them excluded was quite different. In the example below, DC is having a big influence.
with plt.rc_context({"font.size": 6}):
sm.graphics.influence_plot(results_crime)
# Finally, it's nice to be able to see plots of our coefficients along with their standard errors. There isn't a built-in **statsmodels** option for this, but happily it's easy to extract the results of regressions in a sensible format. Using the `results` object from earlier, and excluding the intercept, we can get the coefficients from `results.params[1:]` and the associated errors from `results.bse[1:]`.
# Put the results into a dataframe with Name, Coefficient, Error
res_df = (
pd.concat([results_crime.params[1:], results_crime.bse[1:]], axis=1)
.reset_index()
.rename(columns={"index": "Name", 0: "Coefficient", 1: "Error"})
)
# Plot the coefficient values and their errors
(
ggplot(res_df)
+ geom_point(aes("Name", "Coefficient"))
+ geom_errorbar(aes(x="Name", ymin="Coefficient-Error", ymax="Coefficient+Error"))
)
# ## Specification curve analysis
#
# When specifying a model, modellers have many options. These can be informed by field intelligence, priors, and even misguided attempts to find a significant result. Even with the best of intentions, research teams can reach entirely different conclusions using the same, or similar, data because of different choices made in preparing data or in modelling it.
#
# There’s formal evidence that researchers really do make different decisions; one study {cite}`silberzahn2018many` gave the same research question - whether soccer referees are more likely to give red cards to dark-skin-toned players than to light-skin-toned players - to 29 different teams. From the abstract of that paper:
#
# > Analytic approaches varied widely across the teams, and the estimated effect sizes ranged from 0.89 to 2.93 (Mdn = 1.31) in odds-ratio units. Twenty teams (69%) found a statistically significant positive effect, and 9 teams (31%) did not observe a significant relationship. Overall, the 29 different analyses used 21 unique combinations of covariates. Neither analysts’ prior beliefs about the effect of interest nor their level of expertise readily explained the variation in the outcomes of the analyses. Peer ratings of the quality of the analyses also did not account for the variability.
#
# So not only were different decisions made, there seems to be no clearly identifiable reason for them. There is usually scope for reasonable alternative model specifications when estimating coefficients, and those coefficients will vary with those specifications.
#
# Specification curve analysis {cite}`simonsohn2020specification` looks for a more exhaustive way of trying out alternative specifications. The three steps of specification curve analysis are:
#
# 1. identifying the set of theoretically justified, statistically valid, and non-redundant analytic specifications;
#
# 2. displaying alternative results graphically, allowing the identification of decisions producing different results; and
#
# 3. conducting statistical tests to determine whether as a whole results are inconsistent with the null hypothesis.
#
# For a good example of specification curve analysis in action, see this recent Nature Human Behaviour paper {cite}`orben2019association` on the association between adolescent well-being and the use of digital technology.
#
# We'll use the [**specification curve analysis**](https://specification-curve.readthedocs.io/en/latest/readme.html) package to do the first two, which you can install with `pip install specification_curve`. To demonstrate the full functionality, we'll create a second, alternative 'hp' that is a transformed version of the original.
mpg["hp_boxcox"], _ = st.boxcox(mpg["hp"])
# Now let's create a specification curve. We need to specify the data, the different outcome variables we'd like to try, `y_endog`; the different possible versions of the main regressor of interest, `x_exog`; the possible controls, `controls`; any controls that should always be included, `always_include`; and any categorical variables to include class-by-class, `cat_expand`. Some of these accept lists of variables as well as single reggressors. The point estimates that have confidence intervals which include zero are coloured in grey, instead of blue. There is also an `exclu_grps` option to exclude certain combinations of regressors, and you can pass alternative estimators to fit, for example `fit(estimator=sm.Logit)`.
# +
from specification_curve import specification_curve as specy
sc = specy.SpecificationCurve(
mpg,
y_endog="mpg",
x_exog=["lnhp", "hp_boxcox"],
controls=["drat", "qsec", "cyl", "gear"],
always_include=["gear"],
cat_expand="cyl",
)
sc.fit()
sc.plot()
# -
# ## Review
#
# In this very short introduction to regression with code, you should have learned how to:
#
# - ✅ perform linear OLS regressions with code;
# - ✅ add fixed effects/categorical variables to regressions;
# - ✅ use different standard errors;
# - ✅ use models with transformed regressors;
# - ✅ use the formula or array APIs for **statsmodels** and **linearmodels**;
# - ✅ show the results from multiple models;
# - ✅ perform IV regressions;
# - ✅ perform GLM regressions; and
# - ✅ use plots as a way to interrogate regression results.
| econmt-regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
data = {
'city' : ['Toronto', 'Montreal', 'Waterloo', 'Toronto', 'Waterloo', 'Toronto', 'Toronto'],
'points' : [80, 70, 90, 85, 79, 82, 200]
}
data
type(data)
df = pd.DataFrame(data)
df.groupby('city')['points'].apply(lambda x:x.rolling(window=1).mean())
df.groupby('city')['points'].apply(lambda x:x.rolling(window=2).mean())
| notebooks/pandas/apply-function.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py37] *
# language: python
# name: conda-env-py37-py
# ---
# +
import pandas as pd
import sys
from nntools.utils import Config
sys.path.append('../')
from experiment import OCTClassification
# -
resnet = '/home/clement/Documents/Clement/runs/mlruns/1/f225fafe580e48d491e1eebe122d0635/artifacts/private_predictions_log.csv'
vit = '/home/clement/Documents/Clement/runs/mlruns/1/505896c1301b4f6790efd15dca57b8f9/artifacts/private_predictions_log.csv'
df_vit = pd.read_csv(vit)
df_resnet = pd.read_csv(resnet)
# +
config_path = '../config.yaml'
config = Config(config_path)
config['Network']['architecture'] = 'ResNet152'
experiment = OCTClassification(config)
dataset = experiment.test_dataset
# +
labels = {}
for k, v in dataset.map_class.items():
labels[v] = k
df = pd.DataFrame()
import numpy as np
df['image'] = df_vit['file']
df['prediction_vit'] = df_vit.apply(lambda row:labels[row.prediction], axis=1)
df['probas_vit'] = df_vit.apply(lambda row:max([float(_) for _ in row.probas.replace('[', '').replace(']', '').split(' ') if _]), axis=1)
df['prediction_resnet'] = df_resnet.apply(lambda row:labels[row.prediction], axis=1)
df['probas_resnet'] = df_resnet.apply(lambda row:max([float(_) for _ in row.probas.replace('[', '').replace(']', '').split(' ') if _]), axis=1)
df = df.set_index('image')
# -
np.save('prediction', df.to_dict(), allow_pickle=True)
max([float(_) for _ in df_vit.iloc[0].probas.replace('[', '').replace(']', '').split(' ') if _])
df.to_dict()['probas_resnet']
| notebooks/joined_prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
weatherStation = pd.read_csv('weatherStationLocation.csv')
waterSystem = pd.read_csv('waterSystemLocation.csv')
# print(waterSystem.head)
print(waterSystem[:5])
print(weatherStation[:5])
# print(waterSystem['PWSID'])
print(waterSystem.ix[146347])
print(waterSystem.ix[146341])
# +
weatherLatLong = weatherStation[['Lat','Lon']].values
waterLatLong = waterSystem[['LAT','LON']].values
print(waterLatLong.shape)
print(weatherLatLong.shape)
# -
import time
import scipy.spatial.distance
# +
bestWeatherStations = []
start = time.time()
# distances_test = np.sum(np.power(waterLatLong[ii,:] - weatherLatLong,2),axis=1)
# bestWeatherStations.append(weatherStation['Station'][np.argsort(distances_test)[0]])
Y = scipy.spatial.distance.cdist(waterLatLong,weatherLatLong)
print(Y.shape)
end = time.time()
print('Total computation time is ...', end-start, 'seconds.')
# +
start = time.time()
bestOrder = np.argmin(Y,axis=1)
end = time.time()
print('Total computation time is ...', end-start, 'seconds.')
# +
nearestWeatherStations = weatherStation['Station'][bestOrder]
print(nearestWeatherStations)
# -
print(waterSystem[['PWSID']].shape)
print(nearestWeatherStations.shape)
# +
print(isinstance(waterSystem[['PWSID']],pd.DataFrame))
print(isinstance(nearestWeatherStations,pd.DataFrame))
nearestWeatherStations = pd.DataFrame(np.expand_dims(nearestWeatherStations,1))
isinstance(nearestWeatherStations,pd.DataFrame)
# -
nearestWeatherStationResults = waterSystem[['PWSID']].join(nearestWeatherStations)
print(nearestWeatherStationResults)
| projects/exploratory_phase/code/notebooks/match_waterSystems_weatherStations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/wel51x/DS-Unit-4-Sprint-4-Deep-Learning/blob/master/My_LS_DS_441_RNN_and_LSTM_Assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="_IizNKWLomoA"
# # Lambda School Data Science - Recurrent Neural Networks and LSTM
#
# > "Yesterday's just a memory - tomorrow is never what it's supposed to be." -- <NAME>
# + [markdown] id="LAbnJeJ8j0s3" colab_type="text"
# ####have down-version numpy to get "RNN/LSTM Sentiment Classification with Keras" to work in colab
# + id="4yBeveIrj7x4" colab_type="code" outputId="baabc43b-261d-4015-dc55-35a25425a6fb" colab={"base_uri": "https://localhost:8080/", "height": 296}
# !pip install numpy==1.16.2
import numpy as np
# + [markdown] colab_type="text" id="os-szg47dgwf"
# ### Forecasting
#
# Forecasting - at it's simplest, it just means "predict the future":
# + [markdown] colab_type="text" id="0lfZdD_cp1t5"
# # Assignment
#
# 
#
# It is said that [infinite monkeys typing for an infinite amount of time](https://en.wikipedia.org/wiki/Infinite_monkey_theorem) will eventually type, among other things, the complete works of <NAME>. Let's see if we can get there a bit faster, with the power of Recurrent Neural Networks and LSTM.
#
# This text file contains the complete works of Shakespeare: https://www.gutenberg.org/files/100/100-0.txt
#
# Use it as training data for an RNN - you can keep it simple and train character level, and that is suggested as an initial approach.
#
# Then, use that trained RNN to generate Shakespearean-ish text. Your goal - a function that can take, as an argument, the size of text (e.g. number of characters or lines) to generate, and returns generated text of that size.
#
# Note - Shakespeare wrote an awful lot. It's OK, especially initially, to sample/use smaller data and parameters, so you can have a tighter feedback loop when you're trying to get things running. Then, once you've got a proof of concept - start pushing it more!
# + id="3pqhutb1j9sb" colab_type="code" colab={}
# Imports
from random import random
import numpy as np
import requests
# + colab_type="code" id="Ltj1je1fp5rO" colab={}
# TODO - Words, words, mere words, no matter from the heart.
# Grab first ten
r = requests.get('http://www.gutenberg.org/files/100/100-0.txt', verify=True)
x = r.text.find('From')
y = r.text.find('thine or thee.')
article_text = r.text[x : y+14]
# + id="G6X5y_IKixSD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="75ff32c6-195f-4345-cf5b-f54d1687b7d0"
chars = list(set(article_text)) # split and remove duplicate characters. convert to list.
num_chars = len(chars) # the number of unique characters
txt_data_size = len(article_text)
print("unique characters : ", num_chars)
print("txt_data_size : ", txt_data_size)
# + [markdown] id="GRCADQermdIJ" colab_type="text"
# #### one hot encode
# + id="mCkYGITxjK9v" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 176} outputId="033b0b3d-b8e5-4137-8c74-f14b5e52ccb5"
char_to_int = dict((c, i) for i, c in enumerate(chars)) # "enumerate" retruns index and value. Convert it to dictionary
int_to_char = dict((i, c) for i, c in enumerate(chars))
print(char_to_int)
print("----------------------------------------------------")
print(int_to_char)
print("----------------------------------------------------")
# integer encode input data
integer_encoded = [char_to_int[i] for i in article_text] # "integer_encoded" is a list which has a sequence converted from an original data to integers.
print(integer_encoded)
print("----------------------------------------------------")
print("data length : ", len(integer_encoded))
# + [markdown] id="7RysEhSqmYLX" colab_type="text"
# #### hyperparameters
# + id="ubprmX-hjRt2" colab_type="code" colab={}
iteration = 500
sequence_length = 40
batch_size = round((txt_data_size /sequence_length)+0.5) # = math.ceil
hidden_size = 128 # size of hidden layer of neurons.
learning_rate = 1e-1
# model parameters
W_xh = np.random.randn(hidden_size, num_chars)*0.01 # weight input -> hidden.
W_hh = np.random.randn(hidden_size, hidden_size)*0.01 # weight hidden -> hidden
W_hy = np.random.randn(num_chars, hidden_size)*0.01 # weight hidden -> output
b_h = np.zeros((hidden_size, 1)) # hidden bias
b_y = np.zeros((num_chars, 1)) # output bias
h_prev = np.zeros((hidden_size,1)) # h_(t-1)
# + [markdown] id="zjvJhIDqmS1E" colab_type="text"
# #### Forward propagation
# + id="NGDdTnVVkElU" colab_type="code" colab={}
def forwardprop(inputs, targets, h_prev):
# Since the RNN receives the sequence, the weights are not updated during one sequence.
xs, hs, ys, ps = {}, {}, {}, {} # dictionary
hs[-1] = np.copy(h_prev) # Copy previous hidden state vector to -1 key value.
loss = 0 # loss initialization
for t in range(len(inputs)): # t is a "time step" and is used as a key(dic).
xs[t] = np.zeros((num_chars,1))
xs[t][inputs[t]] = 1
hs[t] = np.tanh(np.dot(W_xh, xs[t]) + np.dot(W_hh, hs[t-1]) + b_h) # hidden state.
ys[t] = np.dot(W_hy, hs[t]) + b_y # unnormalized log probabilities for next chars
ps[t] = np.exp(ys[t]) / np.sum(np.exp(ys[t])) # probabilities for next chars.
# Softmax. -> The sum of probabilities is 1 even without the exp() function, but all of the elements are positive through the exp() function.
loss += -np.log(ps[t][targets[t],0]) # softmax (cross-entropy loss). Efficient and simple code
# y_class = np.zeros((num_chars, 1))
# y_class[targets[t]] =1
# loss += np.sum(y_class*(-np.log(ps[t]))) # softmax (cross-entropy loss)
return loss, ps, hs, xs
# + [markdown] id="TgbdPTQHmONx" colab_type="text"
# #### Backward propagation
# + id="s1l4L4Taludi" colab_type="code" colab={}
def backprop(ps, inputs, hs, xs):
dWxh, dWhh, dWhy = np.zeros_like(W_xh), np.zeros_like(W_hh), np.zeros_like(W_hy) # make all zero matrices.
dbh, dby = np.zeros_like(b_h), np.zeros_like(b_y)
dhnext = np.zeros_like(hs[0]) # (hidden_size,1)
# reversed
for t in reversed(range(len(inputs))):
dy = np.copy(ps[t]) # shape (num_chars,1). "dy" means "dloss/dy"
dy[targets[t]] -= 1 # backprop into y. After taking the soft max in the input vector, subtract 1 from the value of the element corresponding to the correct label.
dWhy += np.dot(dy, hs[t].T)
dby += dy
dh = np.dot(W_hy.T, dy) + dhnext # backprop into h.
dhraw = (1 - hs[t] * hs[t]) * dh # backprop through tanh nonlinearity #tanh'(x) = 1-tanh^2(x)
dbh += dhraw
dWxh += np.dot(dhraw, xs[t].T)
dWhh += np.dot(dhraw, hs[t-1].T)
dhnext = np.dot(W_hh.T, dhraw)
for dparam in [dWxh, dWhh, dWhy, dbh, dby]:
np.clip(dparam, -5, 5, out=dparam) # clip to mitigate exploding gradients.
return dWxh, dWhh, dWhy, dbh, dby
# + [markdown] id="u1ae-K8GmJqc" colab_type="text"
# #### Training
# + id="BCykTWesmDfX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 408} outputId="f621b338-f95b-4ea2-e8e1-f4faefdea4b6"
# %%time
data_pointer = 0
# memory variables for Adagrad
mWxh, mWhh, mWhy = np.zeros_like(W_xh), np.zeros_like(W_hh), np.zeros_like(W_hy)
mbh, mby = np.zeros_like(b_h), np.zeros_like(b_y)
for i in range(iteration+1):
h_prev = np.zeros((hidden_size,1)) # reset RNN memory
data_pointer = 0 # go from start of data
for b in range(batch_size):
inputs = [char_to_int[ch] for ch in article_text[data_pointer:data_pointer+sequence_length]]
targets = [char_to_int[ch] for ch in article_text[data_pointer+1:data_pointer+sequence_length+1]] # t+1
if (data_pointer+sequence_length+1 >= len(article_text) and b == batch_size-1): # processing of the last part of the input data.
# targets.append(char_to_int[txt_data[0]]) # When the data doesn't fit, add the first char to the back.
targets.append(char_to_int[" "]) # When the data doesn't fit, add space(" ") to the back.
# forward
loss, ps, hs, xs = forwardprop(inputs, targets, h_prev)
# print(loss)
# backward
dWxh, dWhh, dWhy, dbh, dby = backprop(ps, inputs, hs, xs)
# perform parameter update with Adagrad
for param, dparam, mem in zip([W_xh, W_hh, W_hy, b_h, b_y],
[dWxh, dWhh, dWhy, dbh, dby],
[mWxh, mWhh, mWhy, mbh, mby]):
mem += dparam * dparam # elementwise
param += -learning_rate * dparam / np.sqrt(mem + 1e-8) # adagrad update
data_pointer += sequence_length # move data pointer
if i % 25 == 0:
print ('iter %d, loss: %f' % (i, loss)) # print progress
# + [markdown] id="_vo6x-LMroI2" colab_type="text"
# #### Prediction
# + id="nwAV1xxcrQKu" colab_type="code" colab={}
def predict(test_char, length):
x = np.zeros((num_chars, 1))
x[char_to_int[test_char]] = 1
ixes = []
h = np.zeros((hidden_size,1))
for t in range(length):
h = np.tanh(np.dot(W_xh, x) + np.dot(W_hh, h) + b_h)
y = np.dot(W_hy, h) + b_y
p = np.exp(y) / np.sum(np.exp(y))
ix = np.random.choice(range(num_chars), p=p.ravel()) # ravel -> rank0
# "ix" is a list of indexes selected according to the soft max probability.
x = np.zeros((num_chars, 1)) # init
x[ix] = 1
ixes.append(ix) # list
txt = test_char + ''.join(int_to_char[i] for i in ixes)
print ('----\n %s \n----' % (txt, ))
# + id="cLP6HY4vrquv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="4418e6f1-1dda-430e-c89a-a80ff8198274"
predict('S', 500)
# + id="-LU-bA4-r4vp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 346} outputId="2bc33d73-f6a5-49be-e009-1a9841d880d7"
predict('C', 750)
# + [markdown] colab_type="text" id="zE4a4O7Bp5x1"
# # Resources and Stretch Goals
# + [markdown] colab_type="text" id="uT3UV3gap9H6"
# ## Stretch goals:
# - Refine the training and generation of text to be able to ask for different genres/styles of Shakespearean text (e.g. plays versus sonnets)
# - Train a classification model that takes text and returns which work of Shakespeare it is most likely to be from
# - Make it more performant! Many possible routes here - lean on Keras, optimize the code, and/or use more resources (AWS, etc.)
# - Revisit the news example from class, and improve it - use categories or tags to refine the model/generation, or train a news classifier
# - Run on bigger, better data
#
# ## Resources:
# - [The Unreasonable Effectiveness of Recurrent Neural Networks](https://karpathy.github.io/2015/05/21/rnn-effectiveness/) - a seminal writeup demonstrating a simple but effective character-level NLP RNN
# - [Simple NumPy implementation of RNN](https://github.com/JY-Yoon/RNN-Implementation-using-NumPy/blob/master/RNN%20Implementation%20using%20NumPy.ipynb) - Python 3 version of the code from "Unreasonable Effectiveness"
# - [TensorFlow RNN Tutorial](https://github.com/tensorflow/models/tree/master/tutorials/rnn) - code for training a RNN on the Penn Tree Bank language dataset
# - [4 part tutorial on RNN](http://www.wildml.com/2015/09/recurrent-neural-networks-tutorial-part-1-introduction-to-rnns/) - relates RNN to the vanishing gradient problem, and provides example implementation
# - [RNN training tips and tricks](https://github.com/karpathy/char-rnn#tips-and-tricks) - some rules of thumb for parameterizing and training your RNN
| My_LS_DS_441_RNN_and_LSTM_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Figure 3 main paper
import autodisc as ad
import random
import numpy as np
import collections
import os
import plotly
import plotly.graph_objs as go
plotly.offline.init_notebook_mode(connected=True)
# +
# default print properties
multiplier = 2
pixel_cm_ration = 36.5
width_full = int(13.95 * pixel_cm_ration) * multiplier
width_half = int(13.95/2 * pixel_cm_ration) * multiplier
height_default_1 = int(4.5 * pixel_cm_ration) * multiplier
height_default_2 = int(7 * pixel_cm_ration) * multiplier
# margins in pixel
top_margin = 0 * multiplier
left_margin = 10 * multiplier
right_margin = 0 * multiplier
bottom_margin = 10 * multiplier
font_size = 8 * multiplier
font_family='Times New Roman'
line_width = 2 * multiplier
# +
plotly.offline.init_notebook_mode(connected=True)
org_experiment_definitions = dict()
org_experiment_definitions['main_paper'] = [
dict(id = '1',
directory = '../experiments/IMGEP-VAE',
name = 'IMGEP-VAE',
is_default = True),
dict(id = '2',
directory = '../experiments/IMGEP-HOLMES',
name = 'IMGEP-HOLMES',
is_default = True),
]
repetition_ids = list(range(10))
# define names and load the data
experiment_name_format = '<name>' # <id>, <name>
#global experiment_definitions
experiment_definitions = []
experiment_statistics = []
current_experiment_list = 'main_paper'
experiment_definitions = []
for org_exp_def in org_experiment_definitions[current_experiment_list]:
new_exp_def = dict()
new_exp_def['directory'] = org_exp_def['directory']
if 'is_default' in org_exp_def:
new_exp_def['is_default'] = org_exp_def['is_default']
if 'name' in org_exp_def:
new_exp_def['id'] = ad.gui.jupyter.misc.replace_str_from_dict(experiment_name_format, {'id': org_exp_def['id'], 'name': org_exp_def['name']})
else:
new_exp_def['id'] = ad.gui.jupyter.misc.replace_str_from_dict(experiment_name_format, {'id': org_exp_def['id']})
experiment_definitions.append(new_exp_def)
experiment_statistics = dict()
for experiment_definition in experiment_definitions:
experiment_statistics[experiment_definition['id']] = dict()
for repetition_idx in repetition_ids:
experiment_statistics[experiment_definition['id']][repetition_idx] = ad.gui.jupyter.misc.load_statistics(os.path.join(experiment_definition['directory'], 'repetition_{:06d}'.format(repetition_idx)))
# -
# # Plotting
# ## RSA Matrix
def plot_goalspaces_RSAmatrix(RSA_matrix, config=None, **kwargs):
default_config = dict(
random_seed = 0,
# global style config
global_layout = dict(
xaxis=dict(
showline = True,
linewidth = 1,
zeroline=False,
ticks = "",
tickfont = dict(
family=font_family,
size=12,
),
),
yaxis=dict(
showline = True,
linewidth = 1,
zeroline=False,
ticks = "",
tickfont = dict(
family=font_family,
size=12,
),
),
font = dict(
family=font_family,
size=font_size,
),
width = width_half, # in cm
height = height_default_1 , # in cm
margin = dict(
l=left_margin, #left margin in pixel
r=right_margin, #right margin in pixel
b=bottom_margin, #bottom margin in pixel
t=top_margin, #top margin in pixel
),
title = "",
hovermode='closest',
showlegend = True,
),
colorscale = 'Viridis',
showscale = False
)
config = ad.config.set_default_config(kwargs, config, default_config)
random.seed(config.random_seed)
n_goal_spaces = len(config.space_names)
x = np.array(config.space_names)
y = np.array(config.space_names)
z = np.asarray([[RSA_matrix[m_i, m_j] for m_i in range(n_goal_spaces)] for m_j in range(n_goal_spaces)])
figure = dict(data=[go.Heatmap(x=x, y=y, z=z, colorscale=config.colorscale, showscale=config.showscale)], layout=config.global_layout)
plotly.offline.iplot(figure)
return figure
# # IMGEP-VAE
# +
RSA_VAE = experiment_statistics['IMGEP-VAE'][0]['temporal_RSA']
config = dict()
config["global_layout"] = dict()
config["global_layout"]["margin"] = dict(t=20*multiplier)
config["global_layout"]["xaxis"] = dict(title="training stages")
config["global_layout"]["yaxis"] = dict(title="training stages")
config["global_layout"]["margin"] = dict(l=20*multiplier, b=20*multiplier)
config["global_layout"]["width"] = width_half
config["global_layout"]["height"] = width_half
config["colorscale"] = 'Viridis'
config["space_names"] = [str(i) for i in range(1,50)]
fig = plot_goalspaces_RSAmatrix(RSA_VAE, config=config)
#plotly.io.write_image(fig, 'main_figure_3_VAE.pdf')
# -
# # IMGEP-HOLMES
# +
RSA_HOLMES = experiment_statistics['IMGEP-HOLMES'][0]['holmes_RSA']
order_default = ['0', '00', '000', '0000', '00000', '00001', '0001', '00010', '00011', '001', '01', '010', '011', '0110', '01100', '01101', '0111', '01110', '01111', '011110', '0111100', '0111101', '011111']
order_desired = ['0', '00', '01', '010', '011', '000', '001', '0110', '0111', '0000', '0001', '01110', '01111', '011110', '011111', '01100', '01101', '00010', '00011', '00000', '00001', '0111100' , '0111101']
RSA_HOLMES_copy = RSA_HOLMES
permute_order = []
for i in order_desired:
permute_order.append(order_default.index(i))
RSA_HOLMES = RSA_HOLMES[permute_order, :]
RSA_HOLMES = RSA_HOLMES[:, permute_order]
order_desired = ['BC 0', 'BC 00', 'BC 01', '<b>BC 010</b>', 'BC 011', 'BC 000', '<b>BC 001</b>', 'BC 0110', 'BC 0111', 'BC 0000', 'BC 0001', '<b>BC 01110</b>', 'BC 01111', 'BC 011110', '<b>BC 011111</b>', '<b>BC 01100</b>', '<b>BC 01101</b>', '<b>BC 00010</b>', '<b>BC 00011</b>', '<b>BC 00000</b>', '<b>BC 00001</b>', '<b>BC 0111100</b>', '<b>BC 0111101</b>']
config = dict()
config["global_layout"] = dict()
config["global_layout"]["xaxis"] = dict(title="modules")
config["global_layout"]["yaxis"] = dict(title="modules")
config["global_layout"]["margin"] = dict(l=45*multiplier, b=42*multiplier, r=40*multiplier)
config["global_layout"]["width"] = width_half + 40*multiplier
config["global_layout"]["height"] = width_half
config["colorscale"] = 'Viridis'
config["space_names"] = order_desired
config["showscale"] = True
fig = plot_goalspaces_RSAmatrix(RSA_HOLMES, config=config)
#plotly.io.write_image(fig, 'main_figure_3_HOLMES.pdf')
# -
| reproduce_paper_figures/make_main_figure_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
import requests
# ## Extract all critera
paloma2 = dict()
studytype_url = 'https://clinicaltrials.gov/api/query/field_values?expr=paloma2&field=StudyType&fmt=json'
studytype = requests.get(studytype_url).json()['FieldValuesResponse']['FieldValues'][0]['FieldValue']
paloma2['StudyType'] = studytype
criteria_url = 'https://clinicaltrials.gov/api/query/field_values?expr=paloma2&field=EligibilityCriteria&fmt=json'
criteria = requests.get(criteria_url).json()['FieldValuesResponse']['FieldValues'][0]['FieldValue']
criteria
a = criteria.split('\n')
inclusion = []
exclusion = []
for i in range(0,len(a)):
if a[i] == 'Inclusion Criteria:':
for j in range(i+1,len(a)):
if a[j] == 'Exclusion Criteria:':
break
if a[j] != '':
inclusion.append(a[j])
if a[i] == 'Exclusion Criteria:':
for j in range(i+1,len(a)):
if a[j] != '':
exclusion.append(a[j])
paloma2['InclusionCriteria'] = inclusion
paloma2['ExclusionCriteria'] = exclusion
paloma2
drug_url = 'https://clinicaltrials.gov/api/query/study_fields?expr=paloma-2&fields=ArmGroupInterventionName&min_rnk=2&max_rnk=2&fmt=json'
drug = requests.get(drug_url).json()['StudyFieldsResponse']['StudyFields'][0]['ArmGroupInterventionName']
drug
drugs = [i.strip("Drug:").strip("'").strip() for i in drug]
drugs = list(dict.fromkeys(drugs))
drugs
# API_ENDPOINT = 'https://www.wikidata.org/w/api.php'
# drugs_with_des1 = {}
# for i in drugs:
# query = i
# params = {
# 'action':'wbsearchentities',
# 'format':'json',
# 'language':'en',
# 'search':query,
# }
# drugs_with_des[i] = requests.get(API_ENDPOINT,params=params).json()['search'][0]['description']
API_ENDPOINT = 'https://en.wikipedia.org/w/api.php'
query = 'Placebo'
params = {
'action':'query',
'list':'search',
'srsearch':query,
'format':'json',
}
a = requests.get(API_ENDPOINT,params=params).json()['query']['search'][0]['snippet']
a
API_ENDPOINT = 'https://en.wikipedia.org/w/api.php'
drugs_with_des = {}
for i in drugs:
drugs_with_des[i] = {}
query = i
params = {
'action':'query',
'list':'search',
'srsearch':query,
'format':'json',
}
drugs_with_des[i]['url'] = requests.get(API_ENDPOINT,params=params).url
drugs_with_des[i]['description'] = requests.get(API_ENDPOINT,params=params).json()\
['query']['search'][0]['snippet']
drugs_with_des
API_ENDPOINT = 'https://en.wikipedia.org/w/api.php'
query = 'Letrozole'
params = {
'action':'query',
'list':'search',
'srsearch':query,
'format':'json',
}
res = requests.get(API_ENDPOINT,params=params).url
res
paloma2['Drugs'] = drugs_with_des
monaleesa2 = {}
studytype_url = 'https://clinicaltrials.gov/api/query/field_values?expr=monaleesa2&field=StudyType&fmt=json'
studytype = requests.get(studytype_url).json()['FieldValuesResponse']['FieldValues'][0]['FieldValue']
monaleesa2['StudyType'] = studytype
criteria_url = 'https://clinicaltrials.gov/api/query/field_values?expr=monaleesa2&field=EligibilityCriteria&fmt=json'
criteria = requests.get(criteria_url).json()['FieldValuesResponse']['FieldValues'][0]['FieldValue']
criteria
a = criteria.split('\n')
inclusion = []
exclusion = []
for i in range(0,len(a)):
if a[i] == 'Inclusion Criteria:':
for j in range(i+1,len(a)):
if a[j] == 'Exclusion Criteria:':
break
if a[j] != '':
inclusion.append(a[j])
if a[i] == 'Exclusion Criteria:':
for j in range(i+1,len(a)):
if a[j] != '':
exclusion.append(a[j])
monaleesa2['InclusionCriteria'] = inclusion
monaleesa2['ExclusionCriteria'] = exclusion
monaleesa2
drug_url2 = 'https://clinicaltrials.gov/api/query/study_fields?expr=monaleesa2&fields=ArmGroupInterventionName&min_rnk=1&max_rnk=&fmt=json'
drug2 = requests.get(drug_url2).json()['StudyFieldsResponse']['StudyFields'][0]['ArmGroupInterventionName']
drugs2 = [i.strip("Drug:").strip("'").strip() for i in drug2]
drugs2 = [i.split(" ") for i in drugs2]
drug = set()
for i in drugs2:
for j in range(0,len(i)):
drug.add(i[j])
API_ENDPOINT = 'https://en.wikipedia.org/w/api.php'
drugs_with_des2 = {}
for i in drug:
drugs_with_des2[i] = {}
query = i
params = {
'action':'query',
'list':'search',
'srsearch':query,
'format':'json',
}
drugs_with_des2[i]['url'] = requests.get(API_ENDPOINT,params=params).url
drugs_with_des2[i]['description'] = requests.get(API_ENDPOINT,params=params).json()\
['query']['search'][0]['snippet']
i = 'LEE011'
query = i
params = {
'action':'query',
'list':'search',
'srsearch':query,
'format':'json',
}
drugs_with_des2[i]['url'] = requests.get(API_ENDPOINT,params=params).url
drugs_with_des2[i]['description'] = requests.get(API_ENDPOINT,params=params).json()\
['query']['search'][1]['snippet']
drugs_with_des2
monaleesa2['Drugs'] = drugs_with_des2
monaleesa2
paloma2
import textdistance
# ## compare clinicaltrials.gove materials
import string
table = str.maketrans('', '', string.punctuation)
for i in paloma2['InclusionCriteria']:
score = 0
a1 = i.lower().split(" ")
a2 = [w.translate(table) for w in a1]
pal = ''
mon = ''
for j in monaleesa2['InclusionCriteria']:
b1 = j.lower().split(" ")
b2 = [w.translate(table) for w in b1]
sim = textdistance.jaccard.normalized_similarity(a2,b2)
if sim > score:
score = sim
pal = i
mon = j
if score > 0.1:
print('paloma2',pal)
print('monaleesa2',mon)
print('simmilarity',score)
for i in paloma2['ExclusionCriteria']:
score = 0
a1 = i.lower().split(" ")
a2 = [w.translate(table) for w in a1]
pal = ''
mon = ''
for j in monaleesa2['ExclusionCriteria']:
b1 = j.lower().split(" ")
b2 = [w.translate(table) for w in b1]
sim = textdistance.jaccard.normalized_similarity(a2,b2)
if sim > score:
score = sim
pal = i
mon = j
if score > 0.1:
print('paloma2',pal)
print('monaleesa2',mon)
print('simmilarity',score)
for i in paloma2['Drugs']:
score = 0
a1 = paloma2['Drugs'][i]['description'].lower().split(" ")
a2 = [w.translate(table) for w in a1]
pal = ''
mon = ''
for j in monaleesa2['Drugs']:
b1 = monaleesa2['Drugs'][j]['description'].lower().split(" ")
b2 = [w.translate(table) for w in b1]
sim = textdistance.jaccard.normalized_similarity(a2,b2)
if sim > score:
score = sim
pal = i
mon = j
if score > 0.1:
print('paloma2',pal)
print('monaleesa2',mon)
print('simmilarity',score)
for j in monaleesa2['InclusionCriteria']:
print(j)
from tabula import read_pdf
table = read_pdf('paloma2.pdf',pages = 4,output_format='json')
def get_vals(test_dict, key):
for i, j in test_dict.items():
if i == key:
yield (j)
yield from [] if not isinstance(j, dict) else get_vals(j, key)
data = []
for i in table[0]['data']:
for j in i:
data.append(list(get_vals(j,'text')))
data = list(filter(lambda a: a != [''],data))
data
paloma2['data'] = data
paloma2
from rdflib import Graph, URIRef, Literal, XSD, Namespace, RDF, RDFS
import pandas as pd
import json
CT = Namespace('http://myclinicaltrials.org/CT#')
SCHEMA = Namespace('http://schema.org/')
# +
kg = Graph()
kg.bind('Clinical_Trials', CT)
kg.bind('schema',SCHEMA)
ct_uri = URIRef('https://clinicaltrials.gov/ct2/show/NCT04199078?term=paloma-2&draw=2&rank=1')
kg.add((ct_uri, RDF.type, SCHEMA.MedicalTrial))
# kg.add((ct_uri, RDF.type, CT['Clinical_Trial']))
# -
for i in paloma2['Drugs']:
print(i)
# +
for i in paloma2:
if i == 'InclusionCriteria':
for j in paloma2[i]:
kg.add((ct_uri, CT.hasInclusionCriteria, Literal(j)))
if i == 'ExclusionCriteria':
for j in paloma2[i]:
kg.add((ct_uri, CT.hasExclusionCriteria, Literal(j)))
if i == 'Drugs':
for j in paloma2[i]:
k = URIRef(paloma2[i][j]['url'])
kg.add((ct_uri, CT.hasDrug, k))
kg.add((k, SCHEMA.name, Literal(j)))
kg.add((k, SCHEMA.description, Literal(paloma2[i][j]['description'])))
# +
# kg.add((CT[drug], SCHEMA.Drug, SCHEMA.MedicalTrial))
# -
kg.serialize('paloma-2.ttl', format="turtle")
| CTKG.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sudo-ken/FFmpeg-for-GDrive/blob/master/FFmpeg_for_GDrive.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="pety7rf841Dg" colab_type="text"
# # **<font color='blue'> Unrar, Unzip, Rar, Zip in GDrive - Shared by [SudoKen](https://www.youtube.com/channel/UCZq5YhZIR-250zX_MzkTbxA?sub_confirmation=1) </font>**
# + [markdown] id="EFOqhHG6hOVH" colab_type="text"
# #__1. Install FFmpeg__
# + id="hFeE-qPuhTiK" colab_type="code" cellView="form" colab={}
#@markdown <br><center><img src='https://raw.githubusercontent.com/sudo-ken/FFmpeg-for-GDrive/master/res/ffmpeg.png' height="50" alt="Gdrive-logo"/></center>
#@markdown <center><h3>Upgrade FFmpeg to v4.2.2</h3></center><br>
from IPython.display import clear_output
import os, urllib.request
HOME = os.path.expanduser("~")
pathDoneCMD = f'{HOME}/doneCMD.sh'
if not os.path.exists(f"{HOME}/.ipython/ttmg.py"):
hCode = "https://raw.githubusercontent.com/sudo-ken/FFmpeg-for-GDrive/master/ttmg.py"
urllib.request.urlretrieve(hCode, f"{HOME}/.ipython/ttmg.py")
from ttmg import (
loadingAn,
textAn,
)
loadingAn(name="lds")
textAn("Installing Dependencies...", ty='twg')
os.system('pip install git+git://github.com/AWConant/jikanpy.git')
os.system('add-apt-repository -y ppa:jonathonf/ffmpeg-4')
os.system('apt-get update')
os.system('apt install mediainfo')
os.system('apt-get install ffmpeg')
clear_output()
print('Installation finished.')
# + [markdown] id="CUq1_Dnegrs1" colab_type="text"
# #__2. Mount Google Drive__
#
#
#
#
# + id="ojI73noUg1If" colab_type="code" colab={} cellView="form"
#@markdown <br><center><img src='https://upload.wikimedia.org/wikipedia/commons/thumb/d/da/Google_Drive_logo.png/600px-Google_Drive_logo.png' height="50" alt="Gdrive-logo"/></center>
#@markdown <center><h3>Mount GDrive to /content/drive</h3></center><br>
MODE = "MOUNT" #@param ["MOUNT", "UNMOUNT"]
#Mount your Gdrive!
from google.colab import drive
drive.mount._DEBUG = False
if MODE == "MOUNT":
drive.mount('/content/drive', force_remount=True)
elif MODE == "UNMOUNT":
try:
drive.flush_and_unmount()
except ValueError:
pass
get_ipython().system_raw("rm -rf /root/.config/Google/DriveFS")
# + [markdown] id="KgNPvGccgwd8" colab_type="text"
# #__3. Run FFmpeg Scripts (Convert, Edit, Trim + more)__
# + [markdown] id="RDHuIkoi6l9a" colab_type="text"
# ###__Display Media File Metadata__
# + id="Sv8au_RO6WUs" colab_type="code" cellView="form" colab={}
import os, sys, re
media_file_path = "" #@param {type:"string"}
os.environ['inputFile'] = media_file_path
# !ffmpeg -i "$inputFile" -hide_banner
# + [markdown] id="X4yIG_nqYAoH" colab_type="text"
# > *You can ignore the* "`At least one output file must be specified`" *error after running this.*
#
#
#
# + [markdown] id="NQ0TxfKeghR8" colab_type="text"
# ###__Convert Video File ➔ .mp4 (Lossless)__
# + id="Ls4O5VLwief-" colab_type="code" cellView="form" colab={}
import os, sys, re
video_file_path = "" #@param {type:"string"}
output_file_path = re.search("^[\/].+\/", video_file_path)
output_file_path_raw = output_file_path.group(0)
delsplit = re.search("\/(?:.(?!\/))+$", video_file_path)
filename = re.sub("^[\/]", "", delsplit.group(0))
filename_raw = re.sub(".{4}$", "", filename)
file_extension = re.search(".{3}$", filename)
file_extension_raw = file_extension.group(0)
os.environ['inputFile'] = video_file_path
os.environ['outputPath'] = output_file_path_raw
os.environ['fileName'] = filename_raw
os.environ['fileExtension'] = file_extension_raw
# !ffmpeg -hide_banner -i "$inputFile" -c copy -strict -2 "$outputPath"/"$fileName".mp4
# + [markdown] colab_type="text" id="NObEcBWAJoaz"
# ###__Convert Video File ➔ .mkv (Lossless)__
# + colab_type="code" cellView="form" id="zsx4JFLRJoa0" colab={}
import os, sys, re
video_file_path = "" #@param {type:"string"}
output_file_path = re.search("^[\/].+\/", video_file_path)
output_file_path_raw = output_file_path.group(0)
delsplit = re.search("\/(?:.(?!\/))+$", video_file_path)
filename = re.sub("^[\/]", "", delsplit.group(0))
filename_raw = re.sub(".{4}$", "", filename)
file_extension = re.search(".{3}$", filename)
file_extension_raw = file_extension.group(0)
os.environ['inputFile'] = video_file_path
os.environ['outputPath'] = output_file_path_raw
os.environ['fileName'] = filename_raw
os.environ['fileExtension'] = file_extension_raw
# !ffmpeg -hide_banner -i "$inputFile" -c copy -strict -2 "$outputPath"/"$fileName".mkv
# + [markdown] id="FpJXJiRl6-gK" colab_type="text"
# ###__Trim Video File (Lossless)__
# + id="iFBUeQhn7QTc" colab_type="code" cellView="form" colab={}
import os, sys, re
video_file_path = "" #@param {type:"string"}
start_time = "00:00:00.000" #@param {type:"string"}
end_time = "00:01:00.000" #@param {type:"string"}
output_file_path = re.search("^[\/].+\/", video_file_path)
output_file_path_raw = output_file_path.group(0)
delsplit = re.search("\/(?:.(?!\/))+$", video_file_path)
filename = re.sub("^[\/]", "", delsplit.group(0))
filename_raw = re.sub(".{4}$", "", filename)
file_extension = re.search(".{3}$", filename)
file_extension_raw = file_extension.group(0)
os.environ['inputFile'] = video_file_path
os.environ['outputPath'] = output_file_path_raw
os.environ['startTime'] = start_time
os.environ['endTime'] = end_time
os.environ['fileName'] = filename_raw
os.environ['fileExtension'] = file_extension_raw
# !ffmpeg -hide_banner -i "$inputFile" -ss "$startTime" -to "$endTime" -c copy "$outputPath"/"$fileName"-TRIM."$fileExtension"
# + [markdown] colab_type="text" id="SNDGdMRn3PA-"
# ###__Crop Video__
# + [markdown] id="KFcIThDuBii_" colab_type="text"
# <h3> Crop Variables Explanation:
#
# * `out_width` = The width of your cropped video file.
# * `out_height` = The height of your cropped video file.
# * `starting_position_x` & `starting_position_y` = These values define the x & y coordinates of the top left corner of your original video to start cropping from.
#
# ###### *Example: For cropping the black bars from a video that looked like* [this](https://i.imgur.com/ud8nbvT.png):
# * *For your starting coordinates* (`x` , `y`) *you would use* (`0` , `138`).
# * *For* `out_width` *you would use* `1920`. *And for* `out_height` *you would use `804`.*
#
#
#
#
# + colab_type="code" cellView="form" id="CEHi5EMm9lXG" colab={}
import os, sys, re
video_file_path = "" #@param {type:"string"}
out_width = "1920" #@param {type:"string"}
out_height = "804" #@param {type:"string"}
starting_position_x = "0" #@param {type:"string"}
starting_position_y = "138" #@param {type:"string"}
output_file_path = re.search("^[\/].+\/", video_file_path)
output_file_path_raw = output_file_path.group(0)
delsplit = re.search("\/(?:.(?!\/))+$", video_file_path)
filename = re.sub("^[\/]", "", delsplit.group(0))
filename_raw = re.sub(".{4}$", "", filename)
file_extension = re.search(".{3}$", filename)
file_extension_raw = file_extension.group(0)
os.environ['inputFile'] = video_file_path
os.environ['outputPath'] = output_file_path_raw
os.environ['outWidth'] = out_width
os.environ['outHeight'] = out_height
os.environ['positionX'] = starting_position_x
os.environ['positionY'] = starting_position_y
os.environ['fileName'] = filename_raw
os.environ['fileExtension'] = file_extension_raw
# !ffmpeg -hide_banner -i "$inputFile" -filter:v "crop=$outWidth:$outHeight:$positionX:$positionY" "$outputPath"/"$fileName"-CROP."$fileExtension"
# + [markdown] colab_type="text" id="2f-THZmDoOaY"
# ###__Extract Audio from Video File (Lossless)__
# + id="nSeO98YQoTJe" colab_type="code" cellView="form" colab={}
import os, sys, re
video_file_path = "" #@param {type:"string"}
output_file_extension = 'm4a' #@param ["m4a", "mp3", "opus", "flac", "wav"]
delsplit = re.search("\/(?:.(?!\/))+$", video_file_path)
output_file_path = re.search("^[\/].+\/", video_file_path)
filename = re.sub("^[\/]", "", delsplit.group(0))
filename_raw = re.sub(".{4}$", "", filename)
os.environ['inputFile'] = video_file_path
os.environ['outputPath'] = output_file_path.group(0)
os.environ['fileName'] = filename_raw
os.environ['fileType'] = output_file_extension
# !ffmpeg -hide_banner -i "$inputFile" -vn -c:a copy "$outputPath"/"$fileName"-audio."$fileType"
# + [markdown] id="MSUasbRUDP3B" colab_type="text"
# ###__Re-encode a Video to a Different Resolution__
# + id="nd2LvSRZCxRe" colab_type="code" cellView="form" colab={}
import os, sys, re
video_file_path = '' #@param {type:"string"}
resolution = '1080p' #@param ["2160p", "1440p", "1080p", "720p", "480p", "360p", "240p"]
file_type = 'mp4' #@param ["mkv", "mp4"]
delsplit = re.search("\/(?:.(?!\/))+$", video_file_path)
testsplit = video_file_path.split("/")
filename = re.sub("^[\/]", "", delsplit.group(0))
filename_raw = re.sub(".{4}$", "", filename)
resolution_raw = re.search("[^p]{3,4}", resolution)
output_file_path = re.search("^[\/].+\/", video_file_path)
os.environ['inputFile'] = video_file_path
os.environ['outputPath'] = output_file_path.group(0)
os.environ['fileName'] = filename_raw
os.environ['fileType'] = file_type
os.environ['resolutionHeight'] = resolution_raw.group(0)
# !ffmpeg -hide_banner -i "$inputFile" -vf "scale=-1:"$resolutionHeight"" -c:a copy -strict experimental "$outputPath"/"$fileName"-"$resolutionHeight"p."$fileType"
# + [markdown] colab_type="text" id="9UagRtLPyKoQ"
# ###__Extract Individual Frames from Video__
# + colab_type="code" cellView="form" id="jTnByMhAyKoF" colab={}
#@markdown This will create a folder in the same directory titled "`Extracted Frames`"
#@markdown - [Example](https://i.imgur.com/yPDk1hO.png) of output folder
import os, sys, re
video_file_path = "" #@param {type:"string"}
start_time = "00:01:00.000" #@param {type:"string"}
end_time = "00:01:05.000" #@param {type:"string"}
frame_rate = "23.976" #@param {type:"string"}
output_file_path = re.search("^[\/].+\/", video_file_path)
output_file_path_raw = output_file_path.group(0)
delsplit = re.search("\/(?:.(?!\/))+$", video_file_path)
filename = re.sub("^[\/]", "", delsplit.group(0))
filename_raw = re.sub(".{4}$", "", filename)
file_extension = re.search(".{3}$", filename)
file_extension_raw = file_extension.group(0)
os.environ['inputFile'] = video_file_path
os.environ['outputPath'] = output_file_path_raw
os.environ['startTime'] = start_time
os.environ['endTime'] = end_time
os.environ['frameRate'] = frame_rate
os.environ['fileName'] = filename_raw
os.environ['fileExtension'] = file_extension_raw
# !mkdir "$outputPath"/"Extracted Frames"
# !ffmpeg -hide_banner -i "$inputFile" -ss "$startTime" -to "$endTime" -r "$frameRate"/1 "$outputPath"/"Extracted Frames"/frame%04d.png
# + [markdown] colab_type="text" id="GahMjYf8miNs"
# ###__Generate Thumbnails - Preview from Video (3x2)__
# + colab_type="code" cellView="form" id="J2u-Rha8miNy" colab={}
#@markdown Example of output image: https://i.imgur.com/0ymP144.png <br>
import os, sys, re
video_file_path = "" #@param {type:"string"}
output_file_type = 'png' #@param ["png", "jpg"]
output_file_path = re.search("^[\/].+\/", video_file_path)
output_file_path_raw = output_file_path.group(0)
delsplit = re.search("\/(?:.(?!\/))+$", video_file_path)
filename = re.sub("^[\/]", "", delsplit.group(0))
filename_raw = re.sub(".{4}$", "", filename)
file_extension = re.search(".{3}$", filename)
file_extension_raw = file_extension.group(0)
os.environ['inputFile'] = video_file_path
os.environ['outputPath'] = output_file_path_raw
os.environ['outputExtension'] = output_file_type
os.environ['fileName'] = filename_raw
os.environ['fileExtension'] = file_extension_raw
# !ffmpeg -hide_banner -i "$inputFile" -vframes 1 -q:v 2 -vf "select=not(mod(n\,200)),scale=-1:480,tile=3x2" -an "$outputPath"/"$fileName"_thumbnails."$outputExtension"
# + [markdown] id="7-3O4en4C4IL" colab_type="text"
# ###__Convert Audio Filetype (mp3, m4a, ogg, flac, etc.)__
# + id="aURlOf9BC1P3" colab_type="code" cellView="form" colab={}
import os, sys, re
audio_file_path = "" #@param {type:"string"}
output_file_type = "mp3" #@param ["mp3", "ogg", "m4a", "opus", "flac", "alac", "wav"]
output_file_path = re.search("^[\/].+\/", audio_file_path)
output_file_path_raw = output_file_path.group(0)
delsplit = re.search("\/(?:.(?!\/))+$", audio_file_path)
filename = re.sub("^[\/]", "", delsplit.group(0))
filename_raw = re.sub(".{4}$", "", filename)
file_extension = re.search(".{3}$", filename)
file_extension_raw = file_extension.group(0)
os.environ['inputFile'] = audio_file_path
os.environ['outputPath'] = output_file_path_raw
os.environ['fileExtension'] = output_file_type
os.environ['fileName'] = filename_raw
# !ffmpeg -hide_banner -i "$inputFile" "$outputPath"/"$fileName"converted."$fileExtension"
# + [markdown] colab_type="text" id="VRk2Ye1exWVA"
# ###__Sharable Links of Randomly Extracted Frames from Video__
# + colab_type="code" cellView="form" id="BIGsgarfxWVI" colab={}
import os, re, time, pathlib
import urllib.request
from IPython.display import clear_output
Auto_UP_Gdrive = False
AUTO_MOVE_PATH = "/content"
HOME = os.path.expanduser("~")
pathDoneCMD = f'{HOME}/doneCMD.sh'
if not os.path.exists(f"{HOME}/.ipython/ttmg.py"):
hCode = "https://raw.githubusercontent.com/biplobsd/" \
"Google-Colab-CloudTorrent/master/res/ttmg.py"
urllib.request.urlretrieve(hCode, f"{HOME}/.ipython/ttmg.py")
from ttmg import (
runSh,
findProcess,
loadingAn,
updateCheck,
ngrok
)
video_file_path = "" #@param {type:"string"}
output_file_path = re.search("^[\/].+\/", video_file_path)
output_file_path_raw = output_file_path.group(0)
delsplit = re.search("\/(?:.(?!\/))+$", video_file_path)
filename = re.sub("^[\/]", "", delsplit.group(0))
filename_raw = re.sub(".{4}$", "", filename)
file_extension = re.search(".{3}$", filename)
file_extension_raw = file_extension.group(0)
os.environ['inputFile'] = video_file_path
os.environ['outputPath'] = output_file_path_raw
os.environ['fileName'] = filename_raw
os.environ['fileExtension'] = file_extension_raw
# !mkdir -p "/content/frames"
for i in range(10):
clear_output()
loadingAn()
print("Uploading Frames...")
# %cd "/content/frames"
# !ffmpeg -hide_banner -ss 00:56.0 -i "$inputFile" -vframes 1 -q:v 1 -y "/content/frames/frame1.png"
# !curl --silent -F "reqtype=fileupload" -F "fileToUpload=@frame1.png" https://catbox.moe/user/api.php -o frame1.txt
f1 = open('frame1.txt', 'r')
# %cd "/content"
file_content1 = f1.read()
# %cd "/content/frames"
# !ffmpeg -hide_banner -ss 02:20.0 -i "$inputFile" -vframes 1 -q:v 1 -y "/content/frames/frame2.png"
# !curl --silent -F "reqtype=fileupload" -F "fileToUpload=@frame2.png" https://catbox.moe/user/api.php -o frame2.txt
# %cd "/content/frames"
f2 = open('frame2.txt', 'r')
# %cd "/content"
file_content2 = f2.read()
clear_output()
print ("Screenshot URLs:")
print ("1. " + file_content1)
print ("2. " + file_content2)
# + [markdown] id="tozwpAhhnm69" colab_type="text"
#
# ###__MediaInfo__
# + id="NTULRguzu0b0" colab_type="code" cellView="form" colab={}
path = "" #@param {type:"string"}
save_txt = True #@param {type:"boolean"}
import os, uuid, re, IPython
import ipywidgets as widgets
import time
from glob import glob
from IPython.display import HTML, clear_output
from google.colab import output, drive
def mediainfo():
display(HTML("<br>"))
# print(path.split("/")[::-1][0])
display(HTML("<br>"))
# # media = !mediainfo "$path"
# media = "\n".join(media).replace(os.path.dirname(path)+"/", "")
get_ipython().system_raw("""mediainfo --LogFile="/root/.nfo" "$path" """)
with open('/root/.nfo', 'r') as file:
media = file.read()
media = media.replace(os.path.dirname(path)+"/", "")
print(media)
get_ipython().system_raw("rm -f '/root/.nfo'")
if save_txt:
txt = path.rpartition('.')[0] + ".txt"
if os.path.exists(txt):
get_ipython().system_raw("rm -f '$txt'")
# !curl -s https://pastebin.com/raw/TV8Byydt -o "$txt"
with open(txt, 'a+') as file:
file.write("\n\n")
file.write(media)
while not os.path.exists("/content/drive"):
try:
drive.mount("/content/drive")
clear_output(wait=True)
except:
clear_output()
if not os.path.exists("/usr/bin/mediainfo"):
get_ipython().system_raw("apt-get install mediainfo")
mediainfo()
# + [markdown] id="XDp_IAgx46fP" colab_type="text"
# #◀️ To get more cool stuff like this, subscribe to the Youtube Channel: [SudoKen](https://www.youtube.com/channel/UCZq5YhZIR-250zX_MzkTbxA?sub_confirmation=1)
| FFmpeg_for_GDrive.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assembly statistics
# ---
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
plt.style.use('ggplot')
sns.set_style("whitegrid")
# %matplotlib inline
# %config InlineBackend.figure_format = 'svg'
def read_align(files):
align_stat = pd.DataFrame()
for f in files:
assembly = f.split("/")[-3]
_df = pd.read_csv(f, header=None, names=["sample", "%"], sep="\t")
_df["%"] = [float(x.rstrip("%")) for x in _df["%"].values]
_df = _df.assign(assembly = pd.Series([assembly]*_df.shape[0], index=_df.index))
align_stat = pd.concat([align_stat, _df], sort=True)
return align_stat
# ## Overall assembly statistics
stat_result = pd.read_csv(snakemake.input.stat, sep="\t", header=0)
stat_result_m = pd.melt(stat_result, id_vars=["assembly"])
ax=sns.catplot(kind="bar", x="assembly", y="value", order=sorted(stat_result.assembly),
col="variable", data=stat_result_m, height=2.5,
sharey=False, col_wrap=4)
ax.set_xticklabels(rotation=90)
ax.set_titles("{col_name}")
plt.savefig(snakemake.output[0], dpi=300, bbox_inches="tight")
# ## Distribution of contig lengths
sizedist_result = pd.read_csv(snakemake.input.dist, sep="\t", header=0)
ax = sns.lineplot(data=sizedist_result, hue="assembly", x="min_length", y="%", linewidth=1,
hue_order = sorted(set(sizedist_result.assembly)))
ax.set_ylabel("% of total assembly");
ax.set_xlabel("contig length");
plt.savefig(snakemake.output[1], dpi=300, bbox_inches="tight")
# ## Alignment frequency
align_stat = read_align(snakemake.input.maps)
ax = sns.stripplot(data=align_stat, x="assembly", y="%", hue="assembly",
order=sorted(align_stat.assembly.unique()), hue_order=sorted(align_stat.assembly.unique()))
ax.set_ylabel("% alignment");
ax.set_xlabel("assembly");
plt.savefig(snakemake.output[2], dpi=300, bbox_inches="tight")
| workflow/notebooks/assembly_stats.py.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Random Forest Modeling
# Previously we had used a single decision tree to classify our data, this time around we'll use a random forest.
# +
# import libraries
from warnings import filterwarnings
filterwarnings("ignore")
import pandas as pd
import numpy as np
from scipy import stats
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split, RandomizedSearchCV
from sklearn.metrics import plot_roc_curve
from imblearn.over_sampling import SMOTENC
from src.seed import SEED
from src.helper import confmat, praf1
# %matplotlib inline
sns.set(font_scale=1.2)
# +
# load data, spllt
train = pd.read_csv("../data/processed/train.csv")
X = train.iloc[:, :-1]
y = train.iloc[:, -1]
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=SEED, stratify=y)
# -
# account for class imbalance
sm = SMOTENC(np.arange(19, 69), random_state=SEED, n_jobs=-1)
# X_train, y_train = sm.fit_resample(X_train, y_train)
# ## Baseline Random Forest
# We won't perform any tuning of our model other than specifying a random state
# +
# train and predict a random forest model
rf = RandomForestClassifier(random_state=SEED, n_jobs=-1)
rf.fit(X_train, y_train)
train_pred = rf.predict(X_train)
test_pred = rf.predict(X_test)
# -
# output scores
a = praf1(y_train, train_pred, "Training")
b = praf1(y_test, test_pred, "Testing")
pd.concat([a, b])
# output confusion matrix
confmat([y_train, y_test], [train_pred, test_pred], ["Training", "Testing"])
# +
# output roc/auc curve
fig, ax = plt.subplots(figsize=(12, 8))
plot_roc_curve(rf, X_train, y_train, name="Training", ax=ax)
plot_roc_curve(rf, X_test, y_test, name="Testing", ax=ax)
line = np.linspace(0, 1)
plt.plot(line, line, "--")
plt.title("Baseline Random Forest ROC/AUC Curve")
plt.show()
# -
# This model is definitely interesting, we see in our tests it has an f1 score of .71, with a recall of about .69, and precision of .73, our tests even show an AUC of .86, meaning our model has a good measure of separability. Moving forward we'll aim to increase our auc.
# ## Hyper Parameter Tuning
# Next up is tuning, we'll be looking to increase our performance by optimizing our f1 score. Giving us a balance between precision and recall.
# +
# create param grid, and randomized search
param_grid = {
"max_depth": np.arange(1, 16),
"min_samples_leaf": stats.uniform(),
"max_features": stats.uniform(),
}
rs = RandomizedSearchCV(
RandomForestClassifier(criterion="entropy", random_state=SEED, n_jobs=-1),
param_grid,
n_iter=500,
scoring="f1",
n_jobs=-1,
random_state=SEED,
)
rs.fit(X_train, y_train)
print(rs.best_params_)
# +
# predictions and scoring
train_pred = rs.predict(X_train)
test_pred = rs.predict(X_test)
a = praf1(y_train, train_pred, "Training")
b = praf1(y_test, test_pred, "Testing")
pd.concat([a, b])
# -
# output confusion matrix
confmat([y_train, y_test], [train_pred, test_pred], ["Training", "Testing"])
# +
# output roc/auc curve
fig, ax = plt.subplots(figsize=(12, 8))
plot_roc_curve(rs, X_train, y_train, name="Training", ax=ax)
plot_roc_curve(rs, X_test, y_test, name="Testing", ax=ax)
line = np.linspace(0, 1)
plt.plot(line, line, "--")
plt.title("Baseline Random Forest ROC/AUC Curve")
plt.show()
# -
# After searching through 500 random models, our best one doesn't have a better auc score still at .87, meaning we still differentiate between our classes at the same rate. However, we do see an improvement in accuracy by 2% compared to our baseline model, and our recall has jumped almost 20%. Overall this model is noticeably better, and has less over fitting.
| notebooks/07-Skellet0r-random-forest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''env'': venv)'
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import networkx as nx
import pickle
import json
import math
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(rc={'figure.figsize':(20, 11)}, font_scale=2)
sns.set_style("whitegrid")
from tqdm import tqdm
# Import Custom packages
import sys
sys.path.append('../../')
import utils
import semantic_type_propagation as stp
# -
# # Tripartite vs. Bipartite Graph
# +
df_path_tripartite = '../../output/synthetic_example_tripartite/'
df_path_bipartite = '../../output/synthetic_example_bipartite/'
g_path = '../../../graph_construction/combined_graphs_output/synthetic_benchmark/tripartite/tripartite.graph'
G = pickle.load(open(g_path, "rb"))
df_tr = pd.read_pickle(df_path_tripartite + 'graph_stats_df.pickle')
df_bi = pd.read_pickle(df_path_bipartite + 'graph_stats_with_groundtruth_df.pickle')
# Create the 'graph_stats_with_groundtruth_df' dataframe
df_tr['is_homograph'] = np.nan
is_homograph_map = {}
for node in df_tr[df_tr['node_type']=='cell']['node']:
is_homograph_map[node] = utils.groundtruth.is_cur_node_homograph(G, node)
df_tr['is_homograph'] = df_tr['node'].map(is_homograph_map)
df_tr.to_pickle(df_path_tripartite+'graph_stats_with_groundtruth_df.pickle')
# Remove nodes with degree 1 from the dataframe
df_tr = stp.process_df(df_tr, G)
df_bi = stp.process_df(df_bi, G)
df_tr = df_tr.sort_values(by='betweenness_centrality', ascending=False)
# Compute the number of row nodes each cell node is connected to
df_tr['num_row_nodes'] = [len(utils.graph_helpers.get_neighbors_of_instance(G, node, 'row')) for node in df_tr['node']]
df_tr['num_col_nodes'] = [len(utils.graph_helpers.get_neighbors_of_instance(G, node, 'attr')) for node in df_tr['node']]
df_tr['num_cell_node_neighbors'] = [len(utils.graph_helpers.get_cell_node_neighbors(G, node)) for node in df_tr['node']]
df_tr['num_row_wise_neighbors'] = [len(utils.graph_helpers.get_row_wise_neighbors(G, node)) for node in df_tr['node']]
df_tr.head(50)
# +
ax = sns.scatterplot(data=df_tr, x='betweenness_centrality', y='num_row_nodes', hue='is_homograph', linewidth=0, s=60)
ax.set(xlabel='Betweenness Centrality', ylabel='Number of Rows')
plt.tight_layout()
plt.savefig("../../figures/row_context/bc_vs_num_rows.svg")
# +
ax = sns.scatterplot(data=df_tr, x='betweenness_centrality', y='num_cell_node_neighbors', hue='is_homograph', linewidth=0, s=60)
ax.set(xlabel='Betweenness Centrality', ylabel='Number of Column-wise Neighbors', xscale='log')
plt.tight_layout()
plt.savefig("../../figures/row_context/bc_vs_num_column_neighbors_log.svg")
# -
ax = sns.scatterplot(data=df_tr, x='betweenness_centrality', y='num_row_wise_neighbors', hue='is_homograph', linewidth=0, s=60)
ax.set(xlabel='Betweenness Centrality', ylabel='Number of Row-Wise Neighbors', xscale='log')
plt.tight_layout()
plt.savefig("../../figures/row_context/bc_vs_num_row_neighbors_log.svg")
df_tr.sort_values(by='num_row_nodes', ascending=False)
plt.scatter(df_tr['betweenness_centrality'], df_tr['num_row_nodes'])
plt.xscale('log')
df_bi.head(40)
| network_analysis/notebooks/graph_representation/row_context.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
snacks_df = pd.read_csv('./datasets/snacks.csv', dtype = { 'client_id': str })
snacks_df.head()
snacks_df['country_region'].unique()
snacks_df['country_region'] = snacks_df['country_region'].str.strip()
snacks_df['country_region'].unique()
# +
group = snacks_df.groupby('country_region')['units'].sum().reset_index(name='units').sort_values('units', ascending=False)
plt.figure(figsize=(16, 10))
sns.barplot(x='country_region', y='units', data=group)
# -
clean_df = snacks_df[snacks_df['country_region'] == 'Sur']
clean_df.shape
clean_df['category_name'].isna().sum()
to_rename = {
'pruduction_line': 'production_line',
'office_department_city': 'office_name',
'warehouse_city': 'warehouse_name'
}
clean_df = clean_df.rename(columns=to_rename)
clean_df = clean_df[clean_df['distributor'].notna()]
clean_df['distributor'].isna().any()
clean_df['incentive'] = clean_df['price_type'].apply(lambda x: True if x == 'OFERTADO' else False)
clean_df = clean_df.drop([
'production_line',
'warehouse_id',
'area_name',
'weight',
'currency',
'category_id',
'plant',
'client_id',
'price_type',
'country_region'
], axis=1)
clean_df.shape
float_columns = ['sale_amount', 'sale_discount', 'sale_devolution']
for column in float_columns:
clean_df[column] = clean_df[column].str.replace(',', '').astype(float)
# +
brand_names = {
'M01': 'Cheetos',
'M02': 'Popcorn',
'M03': 'Cheetos',
'M08': 'Criollas',
'M07': 'Ruffles',
'M16': 'Todo en uno',
'Otras': 'Dulces',
'M27': 'Surtidas',
'M25': 'Costillas',
'M24': 'Choclos',
'M11': 'Ruffles econo',
'M20': 'Tortillitas',
'M14': 'Criollas econo',
'M18': 'Caleñas',
'M17': 'Surtidas',
'M23': 'Nachos',
'M28': 'Fritas econo',
'M13': 'Platanos econo',
'M26': 'Costillas econo',
'M09': 'Papas econo'
}
def name_brand(x):
if x in brand_names.keys():
return brand_names[x]
else:
return x
# -
clean_df['brand_id'] = clean_df['brand_id'].apply(name_brand)
clean_df = clean_df.rename(columns={'brand_id': 'brand'})
clean_df['category_name'] = clean_df['category_name'].fillna('Dulces')
# +
def get_warehouse_name(name):
if name is np.nan:
return name
algo = clean_df[clean_df['description'].str.contains(name) | False].groupby('warehouse_name')['date'].count().reset_index().sort_values('date')
if len(algo) == 0:
return np.nan
return algo.iloc[-1]['warehouse_name']
descriptions = clean_df[clean_df['warehouse_name'].isna()]['description'].unique()
desc_warehouse_dict = {}
for each in descriptions:
desc_warehouse_dict[each] = get_warehouse_name(each)
desc_warehouse_dict['VENTA DIRECTA MEDELLIN'] = 'SUR'
# -
clean_df.loc[clean_df['warehouse_name'].isna(), 'warehouse_name'] = clean_df[clean_df['warehouse_name'].isna()]['description'].map(desc_warehouse_dict)
clean_df = clean_df.drop(['description'], axis=1)
to_rename = {
'office_name': 'office',
'warehouse_name': 'warehouse',
'category_name': 'category',
'flavor_name': 'flavor',
'client_name': 'client',
'point_of_sale_name': 'point_of_sale'
}
clean_df = clean_df.rename(columns=to_rename)
# Normalize categorical columns
# office, warehouse, category, brand, flavor, client, point_of_sale, distributor
columns = ['office', 'warehouse', 'category', 'brand', 'flavor', 'client', 'point_of_sale', 'distributor']
for column in columns:
clean_df[column] = clean_df[column].map(lambda name: name.title() if isinstance(name, str) else name)
# +
columns = ['sale_discount', 'sale_devolution']
for column in columns:
if (clean_df[column] <= 0).all():
clean_df[column] *= -1
# -
clean_df.head()
clean_df.to_csv('./datasets/clean_snacks.csv', encoding='utf-8', index=False)
| cleaning_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A hysteresis mechanism
# > The way up is not the same as the way down
#
# - toc: true
# - badges: true
# - comments: false
# - categories: [jupyter]
# > youtube: https://youtu.be/xgRDhOifFow
# ## Introduction
#
# Hysteresis mechanism created by bistability of states.
#
# Energy function:
# $$f = u^4 - 2u^2 + hu$$
#
# ## The code
# comment these lines if you want interactive mode,
# i.e., if you want to see the animation in real time.
import matplotlib
matplotlib.use('Agg')
# +
import matplotlib.pyplot as plt
import numpy as np
import os
import sympy
from scipy.integrate import ode
# learn how to configure: http://matplotlib.sourceforge.net/users/customizing.html
params = {#'backend': 'GTKAgg',
'legend.handlelength': 2.5,
'legend.borderaxespad': 0,
'font.family':'serif',
'font.size': 18,
'font.serif':['Times'], # Times, Palatino, New Century Schoolbook, Bookman, Computer Modern Roman
'ps.usedistiller': 'xpdf',
'text.usetex': True,
}
plt.rcParams.update(params)
fig=plt.figure(1,figsize=(9.6,5.4),dpi=100) # 1920x1080 # figsize accepts only inches. if you rather think in cm, change the code yourself.
fig.clf()
fig.subplots_adjust(left=0.07, right=0.93,top=0.90, bottom=0.12,hspace=0.02,wspace=0.10)
Hlim=2.5 # parameter range from -Hlim to Hlim
ax1=fig.add_subplot(121)
ax1.set_xticks([])
ax1.set_yticks([])
ax1.set_xlabel(r'System response',labelpad=12)
ax1.set_ylabel('Energy',labelpad=12)
ax1.axis([-Hlim,Hlim,-5,5])
ax2=fig.add_subplot(122)
ax2.set_xticks([])
ax2.set_yticks([])
ax2.set_xlabel(r'Parameter',labelpad=12)
ax2.set_ylabel(r'System response',labelpad=12)
ax2.yaxis.set_label_position("right")
ax2.axis([-Hlim*1.2,Hlim*1.2,-2,2])
frame_names = []
frame_index = 0
make_movie=True
plt.ion()
# +
# energy function and its derivative
f = lambda u,h: u**4-2*u**2+h*u
fprime = lambda u,h: sympy.diff(f(u,h),u)
Hinit=Hlim
ulim=2.5 # system response axis, from -ulim to ulim
u = np.linspace(-ulim,ulim,101)
x = sympy.Symbol('x')
def res(h):
"""System response is one of the real roots
of the energy function derivative
"""
# derivative roots, complex
resp = sympy.solvers.solve(fprime(x,h),x)
# numerical evaluation
resp = map(sympy.N,resp)
# let's check which roots are real
isreal = len(resp)*[False]
for i in range(len(resp)):
# negligible imaginary component
if np.abs(sympy.functions.im(resp[i]))<1e-15:
resp[i]=sympy.functions.re(resp[i])
isreal[i]=True
resp = np.array(resp)
# return only real roots
return resp[np.array(isreal)]
# let's plot stuff, and make a nice movie
#### left plot, ax1 ####
line_func, = ax1.plot(u,f(u,Hinit),lw=2,color='black')
# ball color
ball_color = "blue"
# minimum = the smallest root, the leftmost root
mini = np.min(res(Hinit)) # calculated for initial parameter value
boost = 0.22 # so that ball sits on top of the curve
# plot ball
ball_u, = ax1.plot([mini],[f(mini,Hinit)+boost],'o',
markersize=12, markerfacecolor=ball_color)
#### right plot, ax2 ####
# build empty hysteresis array, we will add values
# as simulation progresses
deetype = np.dtype([('h', 'float64'), ('u', 'float64')])
hysteresis = np.array([(Hinit,mini)],dtype=deetype)
line_hyst, = ax2.plot(hysteresis['h'],hysteresis['u'], lw=2,color='black')
ballH, = ax2.plot([hysteresis['h'][-1]],[hysteresis['u'][-1]],'o',
markersize=12, markerfacecolor=ball_color)
plt.show()
# +
# time to simulate
Total_time = 15 # seconds
fps = 24 # frames per second
# divided by 2 because we ramp down then up
param_vec = np.linspace(Hlim,-Hlim,Total_time*fps/2)
# ramp down
for H in param_vec:
line_func.set_data(u,f(u,H)) # update line on the left
mini = np.min(res(H)) # calculate new minimum
ball_u.set_data([mini],[f(mini,H)+boost]) # update ball on the left
new_line = np.array([(H,mini)],dtype=deetype) # create new line
# append new line to hysteresis array
hysteresis = np.concatenate((hysteresis,new_line))
line_hyst.set_data(hysteresis['h'],hysteresis['u']) # update line
ballH.set_data([hysteresis['h'][-1]],[hysteresis['u'][-1]]) # update ball on the right
fig.canvas.draw()
if make_movie:
fname = "_tmp{:05d}.png".format(frame_index)
frame_names.append(fname)
fig.savefig(fname,dpi=200)
frame_index+=1
# ramp up
for H in param_vec[::-1]: # just reverse parameter array
line_func.set_data(u,f(u,H))
maxi = np.max(res(H)) # everything is the same, but now with maximum
ball_u.set_data([maxi],[f(maxi,H)+boost])
new_line = np.array([(H,maxi)],dtype=deetype)
hysteresis = np.concatenate((hysteresis,new_line))
line_hyst.set_data(hysteresis['h'],hysteresis['u'])
ballH.set_data([hysteresis['h'][-1]],[hysteresis['u'][-1]])
fig.canvas.draw()
if make_movie:
fname = "_tmp{:05d}.png".format(frame_index)
frame_names.append(fname)
fig.savefig(fname,dpi=200)
frame_index+=1
if make_movie:
frames = "_tmp%5d.png"
movie_command = "ffmpeg -y -r {:} -i {:} ball.mp4".format(fps,frames)
os.system(movie_command)
for fname in frame_names:
# pass
os.remove(fname)
| _notebooks/2020-01-01-hysteresis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:carnd-term1]
# language: python
# name: conda-env-carnd-term1-py
# ---
# +
import csv
import cv2
import numpy as np
lines = []
with open('/Users/gongjianxin/Desktop/data_sample/driving_log.csv') as csvfile:
reader = csv.reader(csvfile)
for line in reader:
lines.append(line)
images = []
measurements = []
for line in lines:
source_path = line[0]
filename = source_path.split('/')[-1]
current_path = '/Users/gongjianxin/Desktop/data_sample/IMG/' + filename
image = cv2.imread(current_path)
images.append(image)
measurement= float(line[3])
measurements.append(measurement)
augmented_images,augmented_measurements = [],[]
for image,measurement in zip(images,measurements):
augmented_images.append(image)
augmented_measurements.append(measurement)
augmented_images.append(cv2.flip(image,1))
augmented_measurements.append(measurement*-1.0)
X_train = np.array(augmented_images)
y_train = np.array(augmented_measurements)
from keras.models import Sequential
from keras.layers.core import Dense, Activation, Flatten, Dropout, Lambda
from keras.layers import Cropping2D
from keras.layers.convolutional import Convolution2D
from keras.layers.pooling import MaxPooling2D
model = Sequential()
model.add(Lambda(lambda x: x/255.0-0.5, input_shape=(160,320,3)))
model.add(Cropping2D(cropping=((70,25),(0,0))))
model.add(Convolution2D(6, 5, 5, activation="relu"))
#model.add(Activation('relu'))
model.add(MaxPooling2D())
model.add(Convolution2D(6, 5, 5, activation="relu"))
#model.add(Activation('relu'))
model.add(MaxPooling2D())
model.add(Flatten())
model.add(Dense(120))
model.add(Dense(84))
model.add(Dense(1))
model.compile(loss='mse',optimizer='adam')
model.fit(X_train, y_train, validation_split=0.2,shuffle = True,nb_epoch = 2)
model.save('model.h5')
# -
| behavior_cloning_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Input and Output
# + [markdown] slideshow={"slide_type": "fragment"}
# ## Opening files
#
# How to open a file ?
#
# `f = open(filename, mode='r')`
#
# modes can be:
#
# * `r` for read-only mode
# * `w` for write mode
# * `a` for append mode
# * `+` for read+write mode
# * `b` for binary mode (disable encoding handling)
#
#
# Common methods for all file objects:
#
# * `f.close()` → close the file
# * `seek(pos)` → Moves to a given position in the file
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Writing into a file
#
# - `file.write(str)` → write a string to the file
# - `file.writelines([list of strings])` → write multiple lines
# - `file.flush()` → write (actually) the data to the disk
#
# + slideshow={"slide_type": "fragment"}
import time
fn1 = "example_io1"
f = open(fn1, "w")
f.write("It is now: " + time.ctime())
f.flush() #Optional
f.close() # Mandatory
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Reading from a file
#
#
# - `file.read(size)` → read size characters or the whole file in a string
# - `file.readlines()` → read the whole file in a list of lines
# - `file.readline()` → read the next line (by default defined by `\n`)
#
# + slideshow={"slide_type": "fragment"}
print(time.ctime())
print(open(fn1).read())
# + [markdown] slideshow={"slide_type": "-"}
# **Nota:** In the former cell, the file may not be closed immediately as the garbage collection can be delayed. As the number of opened files is limited at ~1000 per process, it is always better to explicitely close files.
#
# **This is best achieved using a `context manager`.**
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Context manager
#
# * Context manager will allocate and release resources 'automatically' when needed.
# * Usually used from the `with` statement.
#
# so to write safelly into a file, instead of having something like:
# + slideshow={"slide_type": "-"}
file = open('my_file', 'w')
try:
file.write('Hi there!')
finally:
file.close()
# + [markdown] slideshow={"slide_type": "-"}
# you will have:
# + slideshow={"slide_type": "-"}
with open('my_file', 'w') as opened_file:
opened_file.write('Hi there!')
# + [markdown] slideshow={"slide_type": "-"}
# The main advantage of using a `with` statement is to make sure your file will be closed (without dealing with the `finally` block in this case)
#
# This will also increase code readability.
#
# A **common use case of context managers is locking and unlocking resources**.
#
# Yasoob Khalid Python tips regarding context manager:
#
# http://book.pythontips.com/en/latest/context_managers.html
# + [markdown] slideshow={"slide_type": "subslide"}
# ## File as iterator
#
# Files can behave as iterators over readlines
# + slideshow={"slide_type": "-"}
for line in open('data/dummy_file.txt'):
print(line)
# + [markdown] slideshow={"slide_type": "fragment"}
# will display:
# ``` text
# first line
# second line
# ...
# ```
#
# * Very concise typing
# * Efficient reading
# * Limited memory footprint (File is not fully loaded in memory → only one line at a time)
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Hands on
#
# write into a file:
#
# - your name
# - the current date
#
# Then read back this file and parse it to retrieve the month of the date.
# Use only the functions given by the file object (the one returned by open(...))
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Solution - writing
# + slideshow={"slide_type": "fragment"}
import time
with open('myoutputfile', mode='w') as opened_file:
opened_file.write('Jerome\n')
opened_file.write(time.ctime())
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Solution - reading
# + slideshow={"slide_type": "fragment"}
# reading
with open('myoutputfile', mode='r') as f:
# read the first line with the name
firstline = f.readline()
# read the second line with the date
secondline = f.readline()
month = secondline.split()[1]
print("month is %s"%month)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Hands on
#
# Read an ascii spreadsheet written by FIT2D:
#
# * The first non commented line looks like:
# * `512 512 Start pixel = ( 1 1 )`
# * Then 512 values per line, 512 lines
# * Read the file as a list of lists and display as an image.
#
# If `data` is a list of lists (of float), this can be done using matlab with:
#
# ``` python
# # # %matplotlib inline
# from matplotlib.pyplot import subplots
#
# fig, ax = subplots()
# ax.imshow(data)
# ```
#
# * Example file in : data/example.spr
#
#
# 
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Solution
# + slideshow={"slide_type": "subslide"}
def readspr(filepath):
"Read a FIT2D ascii spread file"
result = []
xsize = 0
ysize = 0
with open(filepath, 'r') as opened_file:
for idx, line in enumerate(opened_file):
strippedline = line.strip()
# if this is a commented line
if strippedline.startswith('#'):
continue
words = strippedline.split()
if(len(words) == 8) and (words[2:6] == ["Start", "pixel", "=", "("]):
xsize = int(words[0])
ysize = int(words[1])
print("Dimensions of the size are (%s, %s)" %(xsize, ysize))
break
if xsize and ysize:
for line in opened_file:
words = line.split()
if len(words) != xsize:
print("Error !!! Expected entries are %s, got %s"%(xsize, len(words)))
return None
else:
result.append([float(i) for i in words])
return result
data = readspr("data/example.spr")
# + slideshow={"slide_type": "fragment"}
#Display the image
# %matplotlib inline
from matplotlib.pyplot import subplots
fig, ax = subplots()
ax.imshow(data)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Interaction with the console
#
# - Output to the console:
# - `print(str)`
# - Input from the console:
# - `input()` reads the standard input and returns a string (`raw_input` in python2)
# + slideshow={"slide_type": "fragment"}
user_name = input('please enter your name')
print('user name is', user_name)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Hands on
#
#
# Create a function asking for the name and the age of the user and then display it
#
# ```
# >>> What is your name ?
# polo
# >>> How old are you ?
# 22
# >>> Your name is polo and you are 22 years old
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Solution
#
# + slideshow={"slide_type": "fragment"}
def questioner():
print("What is your name ?")
name = input()
print("How old are you ? ")
age = input()
print("Your name is %s and you are %s years old" % (name, age))
questioner()
# + [markdown] slideshow={"slide_type": "subslide"}
# ## sys standard output, input and error
# - `sys.stdout`, `sys.stdin`, `sys.stderr`:
# - File objects used for standard output, input and errors.
# - `sys.stdout` is a file open in write mode
# - `sys.stdin` is a file open in read mode
# - `sys.stdout.write(str+os.linesep)` is equivalent to `print(str)`
# + [markdown] slideshow={"slide_type": "fragment"}
# ``` ipython
# >>> import sys
# >>> import os
# >>> sys.stdout.write('Is winter coming ?' + os.linesep)
# Is winter coming ?
# >>> answer = sys.stdin.readline()
# might
# >>> print(answer)
# might
# ```
# + [markdown] slideshow={"slide_type": "fragment"}
# example of stdin, stdout
# -
import sys
sys.stdout = open('std_out_file', 'w')
print('toto')
sys.stdout.flush()
| sesame/3_1_general_io.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import gpsro_tools
# # Import Occultation Data
# +
# Import occultation data
#import april with data on which year of occultation was taken
march_year_info = np.load('/usb/monthly_diurnal_cycle_data_occultations/march_cosmic_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
april_year_info = np.load('/usb/monthly_diurnal_cycle_data_occultations/april_cosmic_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
may_year_info = np.load('/usb/monthly_diurnal_cycle_data_occultations/may_cosmic_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
march_year_info_metopa = np.load('/usb/monthly_diurnal_cycle_data_occultations/march_metop_A_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
april_year_info_metopa = np.load('/usb/monthly_diurnal_cycle_data_occultations/april_metop_A_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
may_year_info_metopa = np.load('/usb/monthly_diurnal_cycle_data_occultations/may_metop_A_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
march_year_info_metopb = np.load('/usb/monthly_diurnal_cycle_data_occultations/march_metop_B_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
april_year_info_metopb = np.load('/usb/monthly_diurnal_cycle_data_occultations/april_metop_B_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
may_year_info_metopb = np.load('/usb/monthly_diurnal_cycle_data_occultations/may_metop_B_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
march_year_info_grace = np.load('/usb/monthly_diurnal_cycle_data_occultations/march_grace_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
april_year_info_grace = np.load('/usb/monthly_diurnal_cycle_data_occultations/april_grace_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
may_year_info_grace = np.load('/usb/monthly_diurnal_cycle_data_occultations/may_grace_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
march_year_info_tsx = np.load('/usb/monthly_diurnal_cycle_data_occultations/march_tsx_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
april_year_info_tsx = np.load('/usb/monthly_diurnal_cycle_data_occultations/april_tsx_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
may_year_info_tsx = np.load('/usb/monthly_diurnal_cycle_data_occultations/may_tsx_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
march_year_info_kompsat5 = np.load('/usb/monthly_diurnal_cycle_data_occultations/march_kompsat5_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
april_year_info_kompsat5 = np.load('/usb/monthly_diurnal_cycle_data_occultations/april_kompsat5_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
may_year_info_kompsat5 = np.load('/usb/monthly_diurnal_cycle_data_occultations/may_kompsat5_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
march_year_info_paz = np.load('/usb/monthly_diurnal_cycle_data_occultations/march_paz_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
april_year_info_paz = np.load('/usb/monthly_diurnal_cycle_data_occultations/april_paz_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
may_year_info_paz = np.load('/usb/monthly_diurnal_cycle_data_occultations/may_paz_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
march_year_info_cosmic2 = np.load('/usb/monthly_diurnal_cycle_data_occultations/march_cosmic2_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
april_year_info_cosmic2 = np.load('/usb/monthly_diurnal_cycle_data_occultations/april_cosmic2_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
may_year_info_cosmic2 = np.load('/usb/monthly_diurnal_cycle_data_occultations/may_cosmic2_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
march_year_info_sacc = np.load('/usb/monthly_diurnal_cycle_data_occultations/march_sacc_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
april_year_info_sacc = np.load('/usb/monthly_diurnal_cycle_data_occultations/april_sacc_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
may_year_info_sacc = np.load('/usb/monthly_diurnal_cycle_data_occultations/may_sacc_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
march_year_info_tdx = np.load('/usb/monthly_diurnal_cycle_data_occultations/march_tdx_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
april_year_info_tdx = np.load('/usb/monthly_diurnal_cycle_data_occultations/april_tdx_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
may_year_info_tdx = np.load('/usb/monthly_diurnal_cycle_data_occultations/may_tdx_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
march_year_info_metopc = np.load('/usb/monthly_diurnal_cycle_data_occultations/march_metopc_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
april_year_info_metopc = np.load('/usb/monthly_diurnal_cycle_data_occultations/april_metopc_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
may_year_info_metopc = np.load('/usb/monthly_diurnal_cycle_data_occultations/may_metopc_diurnal_cycles_TLS_year_info.npy', allow_pickle=True)
apr_data = np.concatenate((april_year_info.T, april_year_info_metopa.T, april_year_info_metopb.T,
april_year_info_grace.T, april_year_info_tsx.T, april_year_info_kompsat5.T,
april_year_info_cosmic2.T, april_year_info_paz.T, april_year_info_sacc.T,
april_year_info_tdx.T, april_year_info_metopc.T))
may_data = np.concatenate((may_year_info.T, may_year_info_metopa.T, may_year_info_metopb.T,
may_year_info_grace.T, may_year_info_tsx.T, may_year_info_kompsat5.T,
may_year_info_cosmic2.T, may_year_info_paz.T, may_year_info_sacc.T,
may_year_info_tdx.T, may_year_info_metopc.T))
mar_data = np.concatenate((march_year_info.T, march_year_info_metopa.T, march_year_info_metopb.T,
march_year_info_grace.T, march_year_info_tsx.T, march_year_info_kompsat5.T,
march_year_info_cosmic2.T, march_year_info_paz.T, march_year_info_sacc.T,
march_year_info_tdx.T, march_year_info_metopc.T))
#create dataframes for season
apr_year_info_df = pd.DataFrame(apr_data, columns=['Lat', 'Lon', 'Year', 'Day', 'Hour', 'Temp'])
may_year_info_df = pd.DataFrame(may_data, columns=['Lat', 'Lon', 'Year', 'Day', 'Hour', 'Temp'])
mar_year_info_df = pd.DataFrame(mar_data, columns=['Lat', 'Lon', 'Year', 'Day', 'Hour', 'Temp'])
# Round hours to nearest integer
to_int_hour = lambda x: int(x)
mar_year_info_df["Hour"] = mar_year_info_df.Hour.map(to_int_hour)
apr_year_info_df["Hour"] = apr_year_info_df.Hour.map(to_int_hour)
may_year_info_df["Hour"] = may_year_info_df.Hour.map(to_int_hour)
# -
# # Import ERA-5 Data
# +
era_5_apr_06_5x10 = np.load('../../ERA_5_monthly_TLS_maps/april_2006_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_apr_07_5x10 = np.load('../../ERA_5_monthly_TLS_maps/april_2007_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_apr_08_5x10 = np.load('../../ERA_5_monthly_TLS_maps/april_2008_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_apr_09_5x10 = np.load('../../ERA_5_monthly_TLS_maps/april_2009_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_apr_10_5x10 = np.load('../../ERA_5_monthly_TLS_maps/april_2010_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_apr_11_5x10 = np.load('../../ERA_5_monthly_TLS_maps/april_2011_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_apr_12_5x10 = np.load('../../ERA_5_monthly_TLS_maps/april_2012_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_apr_13_5x10 = np.load('../../ERA_5_monthly_TLS_maps/april_2013_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_apr_14_5x10 = np.load('../../ERA_5_monthly_TLS_maps/april_2014_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_apr_15_5x10 = np.load('../../ERA_5_monthly_TLS_maps/april_2015_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_apr_16_5x10 = np.load('../../ERA_5_monthly_TLS_maps/april_2016_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_apr_17_5x10 = np.load('../../ERA_5_monthly_TLS_maps/april_2017_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_apr_18_5x10 = np.load('../../ERA_5_monthly_TLS_maps/april_2018_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_apr_19_5x10 = np.load('../../ERA_5_monthly_TLS_maps/april_2019_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_apr_20_5x10 = np.load('../../ERA_5_monthly_TLS_maps/april_2020_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_may_06_5x10 = np.load('../../ERA_5_monthly_TLS_maps/may_2006_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_may_07_5x10 = np.load('../../ERA_5_monthly_TLS_maps/may_2007_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_may_08_5x10 = np.load('../../ERA_5_monthly_TLS_maps/may_2008_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_may_09_5x10 = np.load('../../ERA_5_monthly_TLS_maps/may_2009_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_may_10_5x10 = np.load('../../ERA_5_monthly_TLS_maps/may_2010_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_may_11_5x10 = np.load('../../ERA_5_monthly_TLS_maps/may_2011_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_may_12_5x10 = np.load('../../ERA_5_monthly_TLS_maps/may_2012_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_may_13_5x10 = np.load('../../ERA_5_monthly_TLS_maps/may_2013_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_may_14_5x10 = np.load('../../ERA_5_monthly_TLS_maps/may_2014_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_may_15_5x10 = np.load('../../ERA_5_monthly_TLS_maps/may_2015_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_may_16_5x10 = np.load('../../ERA_5_monthly_TLS_maps/may_2016_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_may_17_5x10 = np.load('../../ERA_5_monthly_TLS_maps/may_2017_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_may_18_5x10 = np.load('../../ERA_5_monthly_TLS_maps/may_2018_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_may_19_5x10 = np.load('../../ERA_5_monthly_TLS_maps/may_2019_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_may_20_5x10 = np.load('../../ERA_5_monthly_TLS_maps/may_2020_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_mar_06_5x10 = np.load('../../ERA_5_monthly_TLS_maps/march_2006_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_mar_07_5x10 = np.load('../../ERA_5_monthly_TLS_maps/march_2007_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_mar_08_5x10 = np.load('../../ERA_5_monthly_TLS_maps/march_2008_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_mar_09_5x10 = np.load('../../ERA_5_monthly_TLS_maps/march_2009_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_mar_10_5x10 = np.load('../../ERA_5_monthly_TLS_maps/march_2010_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_mar_11_5x10 = np.load('../../ERA_5_monthly_TLS_maps/march_2011_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_mar_12_5x10 = np.load('../../ERA_5_monthly_TLS_maps/march_2012_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_mar_13_5x10 = np.load('../../ERA_5_monthly_TLS_maps/march_2013_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_mar_14_5x10 = np.load('../../ERA_5_monthly_TLS_maps/march_2014_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_mar_15_5x10 = np.load('../../ERA_5_monthly_TLS_maps/march_2015_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_mar_16_5x10 = np.load('../../ERA_5_monthly_TLS_maps/march_2016_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_mar_17_5x10 = np.load('../../ERA_5_monthly_TLS_maps/march_2017_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_mar_18_5x10 = np.load('../../ERA_5_monthly_TLS_maps/march_2018_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_mar_19_5x10 = np.load('../../ERA_5_monthly_TLS_maps/march_2019_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_mar_20_5x10 = np.load('../../ERA_5_monthly_TLS_maps/march_2020_ERA_5_daily_zonal_mean_TLS_map_5_10.npy', allow_pickle=True)
era_5_mar_07_5x10_df = gpsro_tools.era5_df_switcher(era_5_mar_07_5x10)
era_5_mar_08_5x10_df = gpsro_tools.era5_df_switcher(era_5_mar_08_5x10)
era_5_mar_09_5x10_df = gpsro_tools.era5_df_switcher(era_5_mar_09_5x10)
era_5_mar_10_5x10_df = gpsro_tools.era5_df_switcher(era_5_mar_10_5x10)
era_5_mar_11_5x10_df = gpsro_tools.era5_df_switcher(era_5_mar_11_5x10)
era_5_mar_12_5x10_df = gpsro_tools.era5_df_switcher(era_5_mar_12_5x10)
era_5_mar_13_5x10_df = gpsro_tools.era5_df_switcher(era_5_mar_13_5x10)
era_5_mar_14_5x10_df = gpsro_tools.era5_df_switcher(era_5_mar_14_5x10)
era_5_mar_15_5x10_df = gpsro_tools.era5_df_switcher(era_5_mar_15_5x10)
era_5_mar_16_5x10_df = gpsro_tools.era5_df_switcher(era_5_mar_16_5x10)
era_5_mar_17_5x10_df = gpsro_tools.era5_df_switcher(era_5_mar_17_5x10)
era_5_mar_18_5x10_df = gpsro_tools.era5_df_switcher(era_5_mar_18_5x10)
era_5_mar_19_5x10_df = gpsro_tools.era5_df_switcher(era_5_mar_19_5x10)
era_5_mar_20_5x10_df = gpsro_tools.era5_df_switcher(era_5_mar_20_5x10)
era_5_apr_07_5x10_df = gpsro_tools.era5_df_switcher(era_5_apr_07_5x10)
era_5_apr_08_5x10_df = gpsro_tools.era5_df_switcher(era_5_apr_08_5x10)
era_5_apr_09_5x10_df = gpsro_tools.era5_df_switcher(era_5_apr_09_5x10)
era_5_apr_10_5x10_df = gpsro_tools.era5_df_switcher(era_5_apr_10_5x10)
era_5_apr_11_5x10_df = gpsro_tools.era5_df_switcher(era_5_apr_11_5x10)
era_5_apr_12_5x10_df = gpsro_tools.era5_df_switcher(era_5_apr_12_5x10)
era_5_apr_13_5x10_df = gpsro_tools.era5_df_switcher(era_5_apr_13_5x10)
era_5_apr_14_5x10_df = gpsro_tools.era5_df_switcher(era_5_apr_14_5x10)
era_5_apr_15_5x10_df = gpsro_tools.era5_df_switcher(era_5_apr_15_5x10)
era_5_apr_16_5x10_df = gpsro_tools.era5_df_switcher(era_5_apr_16_5x10)
era_5_apr_17_5x10_df = gpsro_tools.era5_df_switcher(era_5_apr_17_5x10)
era_5_apr_18_5x10_df = gpsro_tools.era5_df_switcher(era_5_apr_18_5x10)
era_5_apr_19_5x10_df = gpsro_tools.era5_df_switcher(era_5_apr_19_5x10)
era_5_apr_20_5x10_df = gpsro_tools.era5_df_switcher(era_5_apr_20_5x10)
era_5_may_06_5x10_df = gpsro_tools.era5_df_switcher(era_5_may_06_5x10)
era_5_may_07_5x10_df = gpsro_tools.era5_df_switcher(era_5_may_07_5x10)
era_5_may_08_5x10_df = gpsro_tools.era5_df_switcher(era_5_may_08_5x10)
era_5_may_09_5x10_df = gpsro_tools.era5_df_switcher(era_5_may_09_5x10)
era_5_may_10_5x10_df = gpsro_tools.era5_df_switcher(era_5_may_10_5x10)
era_5_may_11_5x10_df = gpsro_tools.era5_df_switcher(era_5_may_11_5x10)
era_5_may_12_5x10_df = gpsro_tools.era5_df_switcher(era_5_may_12_5x10)
era_5_may_13_5x10_df = gpsro_tools.era5_df_switcher(era_5_may_13_5x10)
era_5_may_14_5x10_df = gpsro_tools.era5_df_switcher(era_5_may_14_5x10)
era_5_may_15_5x10_df = gpsro_tools.era5_df_switcher(era_5_may_15_5x10)
era_5_may_16_5x10_df = gpsro_tools.era5_df_switcher(era_5_may_16_5x10)
era_5_may_17_5x10_df = gpsro_tools.era5_df_switcher(era_5_may_17_5x10)
era_5_may_18_5x10_df = gpsro_tools.era5_df_switcher(era_5_may_18_5x10)
era_5_may_19_5x10_df = gpsro_tools.era5_df_switcher(era_5_may_19_5x10)
era_5_mar_df = pd.concat([era_5_mar_07_5x10_df, era_5_mar_08_5x10_df, era_5_mar_09_5x10_df, era_5_mar_10_5x10_df,
era_5_mar_11_5x10_df, era_5_mar_12_5x10_df, era_5_mar_13_5x10_df, era_5_mar_14_5x10_df,
era_5_mar_15_5x10_df, era_5_mar_16_5x10_df, era_5_mar_17_5x10_df, era_5_mar_18_5x10_df,
era_5_mar_19_5x10_df, era_5_mar_20_5x10_df])
era_5_apr_df = pd.concat([era_5_apr_07_5x10_df, era_5_apr_08_5x10_df, era_5_apr_09_5x10_df, era_5_apr_10_5x10_df,
era_5_apr_11_5x10_df, era_5_apr_12_5x10_df, era_5_apr_13_5x10_df, era_5_apr_14_5x10_df,
era_5_apr_15_5x10_df, era_5_apr_16_5x10_df, era_5_apr_17_5x10_df, era_5_apr_18_5x10_df,
era_5_apr_19_5x10_df, era_5_apr_20_5x10_df])
era_5_may_df = pd.concat([era_5_may_06_5x10_df, era_5_may_07_5x10_df, era_5_may_08_5x10_df, era_5_may_09_5x10_df,
era_5_may_10_5x10_df, era_5_may_11_5x10_df, era_5_may_12_5x10_df, era_5_may_13_5x10_df,
era_5_may_14_5x10_df, era_5_may_15_5x10_df, era_5_may_16_5x10_df, era_5_may_17_5x10_df,
era_5_may_18_5x10_df, era_5_may_19_5x10_df])
# -
# # Begin processing
daily_era5_removed_and_bias_removed_apr = gpsro_tools.background_and_bias_remover(apr_year_info_df, era_5_apr_df)
daily_era5_removed_and_bias_removed_may = gpsro_tools.background_and_bias_remover(may_year_info_df, era_5_may_df)
daily_era5_removed_and_bias_removed_mar = gpsro_tools.background_and_bias_remover(mar_year_info_df, era_5_mar_df)
# #### Regroup data and drop NaN values
data_all_mean_stuff_removed = pd.concat([daily_era5_removed_and_bias_removed_mar,
daily_era5_removed_and_bias_removed_apr,
daily_era5_removed_and_bias_removed_may])
data_all_mean_stuff_removed.dropna(subset=["Temp"], inplace=True)
# <h1> Now create the diurnal cycles
#
cleaned_diurnal_cycle_data = gpsro_tools.box_mean_remover(data_all_mean_stuff_removed)
diurnal_cycles_by_lat, diurnal_cycles_in_boxes = gpsro_tools.diurnal_binner(cleaned_diurnal_cycle_data)
np.save('MAM_GPSRO_5_10_boxes_diurnal_cycles', diurnal_cycles_in_boxes)
| MAMnotebooks/.ipynb_checkpoints/MAMgpsro-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Spyder)
# language: python3
# name: python3
# ---
# Load penetration data based on SBA PPP loans and BLS 2-digit NAICS codes
# +
import pandas as pd
import numpy as np
from scipy.optimize import linprog
from scipy.sparse import identity
from plotnine import * # python lib to use ggplot
# list of 2-digit NAICS sectors to include in the analysis
NAIC2lst = ['23' # Construction
, '44-45' # (retail)
, '54' # Professional and technical services
, '62' # (health & social, includes childcare)
, '71' # Arts, entertainment, and recreation
, '72'] #(accommodation & food)]
fpath = '/Users/aligo/Box/1 RisknDecScience/FEMA recovery/SBA paper/data/'
# read nloans, amount, nestablishments per county and NAIC2 sector
df = pd.read_excel(fpath + 'PPPpenetrationBLS_County_NAICS2US.xlsx', engine='openpyxl' # , dtype={'STATEFP10':'object','COUNTYFP10':'object'}
)
# keep selected NAICS only
pens = df[df['NAICS2'].isin(NAIC2lst)]
# -
# Descriptive: Total loans and amt per state
# Metrics: penetration, loan amount per employee, (tentatively) loan amount / average salary
# Apply metrics on current allocation, average for NAICS2
# - Total loans and amt per 2-dig NAICS
# - Penetration: difference between NAICS 71-72 and other NAICS
# - Look at penetration of subsectors within 71 and 72
#pens['State'].unique()
mbefore = pens.assign( TotLoanAmount = pens['NLoans']*pens['AvgLoanAmount'], TotJobsReported =
pens['NLoans']*pens['AvgJobsReported'] ).groupby('NAICS2').agg(
{'NEstabs':'sum','NLoans':'sum','TotLoanAmount':'sum','TotJobsReported':'sum'} )
mbefore = mbefore.assign( AvgLoanAmount = mbefore['TotLoanAmount'] / mbefore['NLoans']
, penetration = mbefore['NLoans'] / mbefore['NEstabs']
, LoanAmtperEmp = mbefore['TotLoanAmount'] / mbefore['TotJobsReported']
, actual = 1 )
mbefore
# Apply metrics on current allocation, average for NAICS3
# ### Simulate allocation maximizing loan amt per job (will give loans to estabs with least jobs)
# Formulation:
# Decision variables: $NLoans_{n,c}$
# where $n$: NAICS code in [23, 44-45, 54, 62, 71, 72]
# and $c$: county in US
# \begin{equation*}
# max_{NLoans_{n,c}}\ \sum_{n,c}LoanAmtperEmp_{n,c} = \sum_{n,c}\frac{TotLoanAmount_{n,c}}{TotJobsReported_{n,c}} \\
# = \sum_{n,c}NLoans_{n,c}*\frac{L}{TotJobsReported_{n,c}}
# \end{equation*}
# where $L = TotLoanAmount / TotNLoans$ (we assume that loan amount is the same for all loans and is the average of first round)
# Subject to
# $Nloans_{n,c} <= NEstabs_{n,c}$ for any sector n and county c
# $\sum_{n,c}NLoans_{n,c} = TotNLoans$ (preserve same number of loans from first round)
# $Nloans_{n,c} >= 0$
Npairs = pens.shape[0]
TotNLoans = pens['NLoans'].sum()
tmp = pens.assign( TotLoanAmount = pens['AvgLoanAmount'] * pens['NLoans']
, TotJobsReported = pens['AvgJobsReported'] * pens['NLoans'] )
L = tmp['TotLoanAmount'].sum() / TotNLoans
# objective function coefficients
oc = L / tmp['TotJobsReported']
oc[oc.isna()] = 0
oc[np.isinf(oc)] = 0
# inequality constraint matrix and vector A @ x <= b
Au = identity( Npairs, dtype='int8' )
bu = pens['NEstabs']
# inequality constraint matrix
Ae = np.ones([1,Npairs])
# Solver
res = linprog(oc, A_ub=Au, b_ub=bu, A_eq=Ae, b_eq=TotNLoans)
# check solution
print('Total loans from 1st round:', TotNLoans )
print('Total loans from linprog:', sum(res.x) )
nloansim = res.x.round()
print('Total loans, rounded:', sum(nloansim) )
penssim = pens[['State','COUNTYfips','NAICS2','NEstabs','AvgJobsReported']
].assign( NLoans = nloansim )
msim = penssim.assign( TotLoanAmount = penssim['NLoans']*L, TotJobsReported =
penssim['NLoans']*penssim['AvgJobsReported'] ).groupby('NAICS2').agg(
{'NEstabs':'sum','NLoans':'sum','TotLoanAmount':'sum','TotJobsReported':'sum'} )
msim = msim.assign( AvgLoanAmount = msim['TotLoanAmount'] / msim['NLoans']
, penetration = msim['NLoans'] / msim['NEstabs']
, LoanAmtperEmp = msim['TotLoanAmount'] / msim['TotJobsReported']
, actual = 0 )
msim
# +
# join current and simulated allocations
comp = pd.concat([mbefore, msim]).reset_index()
# Total amount per job of current and simulated allocations
compsum = comp.groupby('actual').agg('sum')
compsum = compsum.assign(AvgLoanAmount = compsum['TotLoanAmount'] / compsum['NLoans']
, penetration = compsum['NLoans'] / compsum['NEstabs']
, LoanAmtperEmp = compsum['TotLoanAmount'] / compsum['TotJobsReported']
).reset_index()
ggplot( compsum, aes(x='actual', y='LoanAmtperEmp')
) + geom_bar(stat="identity", position ="identity",alpha=0.5
# ) + scale_alpha_manual(values=[.1, .3]
) + theme_bw(
) + ylab('Average Loan Amount per Job Reported'
) + ggtitle('Actual vs Simulated Allocation: max Loan Amount per Job'
) #+ scale_y_continuous(labels=lambda l: ["%d%%" % (v * 100) for v in l]
#) + coord_flip()
# -
# plot current and simulated allocations per NAICS
ggplot( comp, aes(x='NAICS2', y='LoanAmtperEmp',fill='factor(actual)')
) + geom_bar(stat="identity", position ="identity",alpha=0.5
# ) + scale_alpha_manual(values=[.1, .3]
) + theme_bw(
) + xlab('2-digit NAICS code'
) + ylab('Average Loan Amount per Job Reported'
) + ggtitle('Actual vs Simulated Allocation: max Loan Amount per Job'
) #+ scale_y_continuous(labels=lambda l: ["%d%%" % (v * 100) for v in l]
#) + coord_flip()
# +
tmp = pens['']
# optmize allocation to maximize penetration
#max Sum[all county-naics pairs](Nloansi / NEstabsi)
# s.t. Sum[all county-naics pairs](Nloansi) = Ntotal
# Nloansi <= NEstabsi
# Nloansi >= 0
# Constraints
n = pens.shape[0] # number of decision variables: county-NAICS pairs
NEstabsTot = pens['NEstabs'].sumn()
# Sum[all county-naics pairs](Nloansi) = NEstabsTot
A_eq = np.ones( (1,n) )
b_eq = [NEstabsTot]
# Nloansi <= NEstabsi
A_ub = np.identity( n )
b_ub = pens['NEstabs'].to_numpy()
# vector of coefficients of objective function: 1/NEstabsi
c = 1 / b_ub
c = ( 1 / pens['NEstabs'] ).array
| SBA_paper_Allocation_jobs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Codebook
# **Authors:** <NAME>
# Documenting existing data files of DaanMatch with information about location, owner, "version", source etc.
import boto3
import numpy as np
import pandas as pd
pd.plotting.register_matplotlib_converters()
import matplotlib.pyplot as plt
# %matplotlib inline
from collections import Counter
import statistics
client = boto3.client('s3')
resource = boto3.resource('s3')
my_bucket = resource.Bucket('daanmatchdatafiles')
# # CSR Spent 17-18.xlsx
#
# ## TOC:
# * [About this dataset](#1)
# * [What's in this dataset](#2)
# * [Codebook](#3)
# * [Missing values](#3.1)
# * [Summary statistics](#3.2)
# * [Columns](#4)
# * [Url](#4.1)
# * [Company Name](#4.2)
# * [CSR Spent 17-18](#4.3)
# * [Date of Incorporation](#4.4)
# * [Class](#4.5)
# * [RoC](#4.6)
# * [Category](#4.7)
# * [Sub Category](#4.8)
# * [Listing Status](#4.9)
# * [Registered Address](#4.10)
# * [Zipcode](#4.11)
# * [State](#4.12)
# * [Email ID](#4.13)
# * [Paid-up Capital (in INR Cr.)](#4.14)
# * [Authorized Capital (in INR Cr.)](#4.15)
# **About this dataset** <a class="anchor" id="1"></a>
# Data provided by: Unknown.
# Source: https://daanmatchdatafiles.s3.us-west-1.amazonaws.com/DaanMatch_DataFiles/CSR+Spent+17-18.xlsx
# Type: xlsx
# Last Modified: May 29, 2021, 19:54:24 (UTC-07:00)
# Size: 3.4 MB
path = "s3://daanmatchdatafiles/CSR Spent 17-18.xlsx"
csr_spent_17_18 = pd.ExcelFile(path)
print(csr_spent_17_18.sheet_names)
csr_spent_17_18 = csr_spent_17_18.parse('CSR Spent 17-18')
csr_spent_17_18.head()
# **What's in this dataset?** <a class="anchor" id="2"></a>
print("Shape:", csr_spent_17_18.shape)
print("Rows:", csr_spent_17_18.shape[0])
print("Columns:", csr_spent_17_18.shape[1])
print("Each row is a company.")
# +
csr_spent_17_18_columns = [column for column in csr_spent_17_18.columns]
csr_spent_17_18_description = ["Link to the company's website.",
"Name of Company.",
"Amount of money spent on CSR in the fiscal year 2017-18.",
"Timestamp of date of incorporation: YYYY-MM-DD.",
"Class of Company: Private or Public.",
"Registrar of Companies, an office under the MCA.",
"Category of Company: Limited by Shares, Limited by Guarantee, Unlimited Company.",
"Subcategory of Company: Non-govt, Union Gtvt, State Govt, Subsidiary of Foreign Company, Guarantee and Association Company.",
"Lisitng status: Listed or Unlisited.",
"Address of the registered office.",
"Zipcode of the registered office.",
"State the company is located in.",
"Email address.",
"Actual amount that is paid by shareholders to the company.",
"Maximum value of shares that the company is legally authorized to issue to the shareholders."]
csr_spent_17_18_dtypes = [dtype for dtype in csr_spent_17_18.dtypes]
data = {"Column Name": csr_spent_17_18_columns, "Description": csr_spent_17_18_description, "Type": csr_spent_17_18_dtypes}
csr_spent_17_18_codebook = pd.DataFrame(data)
csr_spent_17_18_codebook.style.set_properties(subset=['Description'], **{'width': '600px'})
# -
# **Missing values** <a class="anchor" id="3.1"></a>
csr_spent_17_18.isnull().sum()
# **Summary statistics** <a class="anchor" id="3.2"></a>
csr_spent_17_18.describe()
# ## Columns
# <a class="anchor" id="4"></a>
# ### Url
# <a class="anchor" id="4.1"></a>
# Link to the company's website.
column = csr_spent_17_18["Url"]
column
# +
print("No. of unique values:", len(column.unique()))
# Check for duplicates
counter = dict(Counter(column))
duplicates = { key:value for key, value in counter.items() if value > 1}
print("Duplicates:", duplicates)
# -
# ### Company Name
# <a class="anchor" id="4.2"></a>
# Name of Company.
column = csr_spent_17_18["Company Name"]
column
# +
print("No. of unique values:", len(column.unique()))
# Check for duplicates
counter = dict(Counter(column))
duplicates = { key:value for key, value in counter.items() if value > 1}
#print("Duplicates:", duplicates)
if len(duplicates) > 0:
print("No. of duplicates:", len(duplicates))
# -
csr_spent_17_18[csr_spent_17_18['Company Name'].isin(duplicates)].sort_values('Company Name')
# Duplicates in ```Companay Name``` does not mean the rows are duplicates. Many of the duplicates appear to be the same company, yet they have different Class, Listing Status, or CSR Spent 17-18 values.
# ### CSR Spent 17-18
# <a class="anchor" id="4.3"></a>
# Amount of money spent on CSR in the fiscal year 2017-18.
column = csr_spent_17_18["CSR Spent 17-18"]
column
# +
#Check how many unique values
print("No. of unique values:", len(column.unique()))
# Check for duplicates
counter = dict(Counter(column))
duplicates = { key:value for key, value in counter.items() if value > 1}
#print("Duplicates:", duplicates)
if len(duplicates) > 0:
print("No. of duplicates:", len(duplicates))
#Check how many values are 0
print("No. of 0 values:", len(column[column == 0]))
print('Max:', max(column))
print("Min:", min(column))
# -
bins= np.linspace(0, 70000000, 15)
plt.figure(figsize = (10, 6))
plt.hist(column[column != 0], bins=bins, edgecolor="k")
plt.title('CSR Spent 17-18')
plt.xlabel('Amount Spent (INR)')
plt.ylabel('Count of Companies')
plt.xticks(bins, rotation = 90);
#Statistics
column.describe()
# ### Date of Incorporation
# <a class="anchor" id="4.4"></a>
# Timestamp of date of incorporation: YYYY-MM-DD.
column = csr_spent_17_18["Date of Incorporation"]
column
# +
#Check how many unique values
print("No. of unique values:", len(column.unique()))
# Check for duplicates
counter = dict(Counter(column))
duplicates = { key:value for key, value in counter.items() if value > 1}
#print("Duplicates:", duplicates)
if len(duplicates) > 0:
print("No. of duplicates:", len(duplicates))
#Check how many values are 0
print("No. of 0 values:", len(column[column == 0]))
print('Max:', max(column))
print("Min:", min(column))
# -
years = column.apply(lambda x: int(x[:4]))
counter = dict(Counter(years))
count = { key:[value] for key, value in counter.items()}
table = pd.DataFrame.from_dict(count)
table = table.melt(var_name="Date", value_name="Count")
print("No. of unique values:", table.shape[0])
table
# Plot of number of each year
plt.figure(figsize = (10, 7))
plt.bar(table["Date"], table["Count"])
plt.title("Count of Year of Incorporation")
plt.ylabel("Count")
plt.xlabel("Year");
# ### Class
# <a class="anchor" id="4.5"></a>
# Class of Company: Private or Public.
column = csr_spent_17_18["Class"]
column
# +
print("Unique values:", column.unique())
print("No. of unique values:", len(column.unique()))
# Number of empty strings
print("Empty strings:", sum(column == " "))
# Table of number of each class
table = column.value_counts().rename_axis('Class').reset_index(name='Count')
table
# -
# Plot number of each class
plt.figure(figsize = (8, 6))
plt.bar(table["Class"], table["Count"])
plt.title("Count of Company Classes")
plt.xlabel("Class")
plt.show()
# ### RoC
# <a class="anchor" id="4.6"></a>
# Registrar of Companies, an office under the MCA.
column = csr_spent_17_18["RoC"]
column
# +
print("Unique values:", column.unique())
print("No. of unique values:", len(column.unique()))
# Table of number of each class
table = column.value_counts().rename_axis('RoC').reset_index(name='Count')
table
# -
# Plot number of each class
plt.figure(figsize = (10, 7))
plt.bar(table["RoC"], table["Count"])
plt.title("Count of RoC")
plt.xlabel("RoC")
plt.ylabel('Count')
plt.xticks(rotation = 90)
plt.show()
# ### Category
# <a class="anchor" id="4.7"></a>
# Category of Company: Limited by Shares, Limited by Guarantee, Unlimited Company.
column = csr_spent_17_18["Category"]
column
# +
print("Unique values:", column.unique())
print("No. of unique values:", len(column.unique()))
# Table of number of each category
table = column.value_counts().rename_axis('Category').reset_index(name='Count')
table
# -
# Plot number of each Category
plt.figure(figsize = (8, 5))
plt.bar(table["Category"], table["Count"])
plt.title("Count of Company Categories")
plt.xlabel("Category")
plt.xticks(rotation = 40)
plt.show()
# ### Sub Category
# <a class="anchor" id="4.8"></a>
# Subcategory of Company: Non-govt, Union Gtvt, State Govt, Subsidiary of Foreign Company, Guarantee and Association Company.
column = csr_spent_17_18["Sub Category"]
column
# +
print("Unique values:", column.unique())
print("No. of unique values:", len(column.unique()))
# Table of number of each sub-category
table = column.value_counts().rename_axis('Sub-category').reset_index(name='Count')
table
# -
# Plot number of each sub-category
plt.figure(figsize = (8, 5))
plt.bar(table["Sub-category"], table["Count"])
plt.title("Count of Company Sub-categories")
plt.xlabel("Sub-category")
plt.xticks(rotation = 30)
plt.show()
# ### Listing Status
# <a class="anchor" id="4.9"></a>
# Lisitng status: Listed or Unlisited.
column = csr_spent_17_18["Listing Status"]
column
# +
print("Unique values:", column.unique())
print("No. of unique values:", len(column.unique()))
# Table of number of each category
table = column.value_counts().rename_axis('Listing Status').reset_index(name='Count')
table
# -
# Plot number of lisiting status'
plt.figure(figsize = (8, 5))
plt.bar(table["Listing Status"], table["Count"])
plt.title("Count of Listing Status")
plt.xlabel("Listing Status")
plt.xticks(rotation = 40)
plt.show()
# ### Registered Address
# <a class="anchor" id="4.10"></a>
# Address of the registered office.
column = csr_spent_17_18["Registered Address"]
column
print("No. of null values:", sum(column.isnull()))
print("No. of unique values:", len(column.unique()) - 1)
# +
# Check for duplicates
counter = dict(Counter(column))
duplicates = { key:[value] for key, value in counter.items() if value > 1}
print("No. of Duplicates:", len(duplicates))
table = pd.DataFrame.from_dict(duplicates)
table = table.melt(var_name="Duplicate Registered Addresses", value_name="Count").sort_values(by=["Count"], ascending=False).reset_index(drop=True)
table
# -
csr_spent_17_18[csr_spent_17_18["Registered Address"].isin(duplicates)].sort_values('Registered Address')
# Duplicates in ```Registered Address``` do not mean duplicates in companies.
# Sometimes a very small NGO cannot afford their own office, and will register their non-profit under another existing one.
# ### Zipcode
# <a class="anchor" id="4.11"></a>
# Zipcode of the registered office.
column = csr_spent_17_18["Zipcode"]
column
# +
print("No. of unique values:", len(column.unique()))
print("No. of zero values:", len(column[column == 0]))
print("No. of null values:", sum(column.isnull()))
# Check for duplicates
counter = dict(Counter(column))
duplicates = { key:value for key, value in counter.items() if value > 1}
#print("Duplicates:", duplicates)
if len(duplicates) > 0:
print("No. of duplicates:", len(duplicates))
# -
# ### State
# <a class="anchor" id="4.12"></a>
# State the company is located in.
column = csr_spent_17_18["State"]
column
# +
print("No. of unique values:", len(column.unique()))
print("No. of zero values:", len(column[column == 0]))
# Check for duplicates
counter = dict(Counter(column))
duplicates = { key:value for key, value in counter.items() if value > 1}
#print("Duplicates:", duplicates)
if len(duplicates) > 0:
print("No. of duplicates:", len(duplicates))
# -
# Table of number of each sub-category
table = column.value_counts().rename_axis('State').reset_index(name='Count')
table.head()
# Plot number of companies in each State
plt.figure(figsize = (10, 6))
plt.bar(table["State"], table["Count"])
plt.title("Count of Companies in each State")
plt.xlabel("State")
plt.xticks(rotation = 90)
plt.show()
# ### Email ID
# <a class="anchor" id="4.13"></a>
# Email address.
column = csr_spent_17_18["Email ID"]
column
# +
print("No. of unique values:", len(column.unique()))
print("No. of null values:", len(column[column == "-NA-"]))
# Check for duplicates
counter = dict(Counter(column))
duplicates = { key:value for key, value in counter.items() if value > 1}
#print("Duplicates:", duplicates)
if len(duplicates) > 0:
print("No. of duplicates:", len(duplicates))
# -
csr_spent_17_18[csr_spent_17_18["Email ID"].isin(duplicates)].sort_values('Email ID', ascending = False)
# Duplicates in ```Email ID``` does not necessarily mean duplicates in rows.
# Some duplicates appear to be the same company, yet the only difference is the "Listing Status" column value.
# ### Paid-up Capital (in INR Cr.)
# <a class="anchor" id="4.14"></a>
# Actual amount that is paid by shareholders to the company.
column = csr_spent_17_18["Paid-up Capital (in INR Cr.)"]
column
#Turning objects into floats
amounts = column.apply(lambda x: str(x[1:]))
amounts = amounts.apply(lambda x: float(x.replace(',','')))
amounts
# +
print("No. of unique values:", len(amounts.unique()))
print("No. of zero values:", len(amounts[amounts == 0]))
# Check for duplicates
counter = dict(Counter(amounts))
duplicates = { key:value for key, value in counter.items() if value > 1}
#print("Duplicates:", duplicates)
if len(duplicates) > 0:
print("No. of duplicates:", len(duplicates))
# -
#Statistics
amounts.describe()
#Histogram of bottom 75% Of Paid-up Capital
bins= np.linspace(0, 18, 25)
plt.figure(figsize = (10, 6))
plt.hist(amounts, bins=bins, edgecolor="k")
plt.title('Paid-up Capital (in INR Cr.)')
plt.ylabel('Count')
plt.xlabel('Amount')
plt.xticks(bins, rotation = 90);
# ### Authorized Capital (in INR Cr.)
# <a class="anchor" id="4.15"></a>
# Maximum value of shares that the company is legally authorized to issue to the shareholders.
column = csr_spent_17_18["Authorized Capital (in INR Cr.)"]
column
#Turning objects into floats
amounts = column.apply(lambda x: str(x[1:]))
amounts = amounts.apply(lambda x: float(x.replace(',','')))
amounts
# +
print("No. of unique values:", len(amounts.unique()))
print("No. of zero values:", len(amounts[amounts == 0]))
# Check for duplicates
counter = dict(Counter(amounts))
duplicates = { key:value for key, value in counter.items() if value > 1}
#print("Duplicates:", duplicates)
if len(duplicates) > 0:
print("No. of duplicates:", len(duplicates))
# -
#Statistics
amounts.describe()
#Histogram of bottom 75% Of Authorized Capital
bins= np.linspace(0, 30, 25)
plt.figure(figsize = (10, 6))
plt.hist(amounts, bins=bins, edgecolor="k")
plt.title('Authorized Capital (in INR Cr.)')
plt.ylabel('Count')
plt.xlabel('Amount')
plt.xticks(bins, rotation = 90);
#For reference, below is CSR Spent 17-18 in INR Cr. for comparison to above histogram
column = csr_spent_17_18["CSR Spent 17-18"] / 1000000
bins= np.linspace(0, 30, 25)
plt.figure(figsize = (10, 6))
plt.hist(column[column != 0], bins=bins, edgecolor="k")
plt.title('CSR Spent 17-18')
plt.xlabel('Amount Spent (INR Cr.)')
plt.ylabel('Count of Companies')
plt.xticks(bins, rotation = 90);
| CSR Spent 17-18/.ipynb_checkpoints/CSR Spent 17-18-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py3.6-env
# language: python
# name: py3.6-env
# ---
# +
# %matplotlib inline
import gym
import matplotlib
import numpy as np
import sys
import plotting
from collections import defaultdict
from envs.blackjack import BlackjackEnv
matplotlib.style.use('ggplot')
# -
env = BlackjackEnv()
def mc_prediction(policy, env, num_episodes, discount_factor=1.0):
"""
蒙特卡罗预测算法。给定策略policy,计算它的价值函数。
参数:
policy: 一个函数,输入是状态(observation),输出是采取不同action的概率。
env: OpenAI gym 环境对象。
num_episodes: 采样的次数。
discount_factor:打折因子。
返回:
一个dictionary state -> value。
state是一个三元组,而value是float。
"""
# 记录每个状态的Return和出现的次数。
returns_sum = defaultdict(float)
returns_count = defaultdict(float)
# 最终的价值函数
V = defaultdict(float)
for i_episode in range(1, num_episodes + 1):
if i_episode % 1000 == 0:
print("\rEpisode {}/{}.".format(i_episode, num_episodes), end="")
sys.stdout.flush()
# 生成一个episode.
# 一个episode是三元组(state, action, reward) 的数组
episode = []
state = env.reset()
for t in range(100):
action = policy(state)
next_state, reward, done, _ = env.step(action)
episode.append((state, action, reward))
if done:
break
state = next_state
# 找到这个episode里出现的所有状态。
states_in_episode = set([tuple(x[0]) for x in episode])
for state in states_in_episode:
# 找到这个状态第一次出现的下标
first_occurence_idx = next(i for i,x in enumerate(episode) if x[0] == state)
# 计算这个状态的Return
G = sum([x[2]*(discount_factor**i) for i,x in enumerate(episode[first_occurence_idx:])])
# 累加
returns_sum[state] += G
returns_count[state] += 1.0
V[state] = returns_sum[state] / returns_count[state]
return V
def sample_policy(observation):
"""
一个简单的策略:如果不到20就继续要牌,否则就停止要牌。
"""
score, dealer_score, usable_ace = observation
return 0 if score >= 20 else 1
# +
V_10k = mc_prediction(sample_policy, env, num_episodes=10000)
plotting.plot_value_function(V_10k, title="10,000 Steps")
V_500k = mc_prediction(sample_policy, env, num_episodes=500000)
plotting.plot_value_function(V_500k, title="500,000 Steps")
| rl/MC Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
Sys.Date()
# +
libraries = c("dplyr","magrittr","tidyr", "purrr", "ggplot2","gridExtra","RColorBrewer","zoo","scales","colorspace","readxl")
for(x in libraries) { library(x,character.only=TRUE,warn.conflicts=FALSE) }
'%&%' = function(x,y) paste0(x,y)
theme_set(theme_classic(base_size=12, base_family="sans"))
fnt_size = 11
CUTOFF_TMAX = as.Date('2020-01-24')
# -
drname = '../../results/Scenario-2/sens_CUTOFF_TIME'
df = data.frame(filename=list.files(drname, pattern="incidence"), stringsAsFactors = FALSE) %>%
mutate(date = stringr::str_split(filename, "_") %>% map_chr(., 1)) %>% filter(date<=CUTOFF_TMAX)
df
for (i in 1:nrow(df)) {
df_ = read.csv(drname%&%"/"%&%df$filename[i])
df_ = df_[df_$var=='r',] %>% select(-time,-var)
if (i==1)
df_r = c(df[i,'date'], df_[1,] %>% as.numeric)
else {
df_r = rbind(df_r, c(df[i,'date'], df_[1,] %>% as.numeric))
}
}
df_r = data.frame(df_r, stringsAsFactors = FALSE, row.names = 1:nrow(df_r))
names(df_r) = c('date','mean','lower','upper')
df_r %<>% mutate(date = as.Date(date)) %>% mutate_each(as.numeric, -date)
df_r
# +
cs = c(4, 4.5)
options(repr.plot.width=cs[1],repr.plot.height=cs[2])
df_r %>%
ggplot(aes(x=date, y=mean, group = 1)) +
geom_line(aes(y=lower), color="black", size=.4) +
geom_line(aes(y=upper), color="black", size=.4) +
geom_ribbon(data=df_r, aes(x=date, ymin = lower, ymax = upper),
alpha = 0.6, fill="grey70", inherit.aes=FALSE) +
geom_line(aes(y=mean), color="black", size=1) +
xlab("Ending timepoint (month day)") + ylab(expression(italic(r))) +
theme(plot.margin = unit(c(.5,.5,1,.25),"lines"),
text = element_text(family="sans",color="black"),
axis.text.x = element_text(angle = 35, hjust = 1),
axis.text =element_text(size=fnt_size, family="sans",color="black"),
panel.grid.major = element_blank(),
panel.grid.minor = element_blank()) -> fig21
fig21
ggsave(plot=fig21, width=cs[1], height=cs[2],filename="../../figures/draft/figS31.pdf",useDingbats=FALSE)
# -
t0 = as.Date('2019-12-08')
for (i in 1:nrow(df)) {
df_ = read.csv(drname%&%"/"%&%df$filename[i])
df_ = df_[df_$var=='Incidence',] %>% select(-var)
if (i==1) {
df_r = df_ %>% as.matrix
df_r = cbind(rep(df$`date`[i], nrow(df_)), df_r)
} else {
df_r = rbind(df_r, cbind(rep(df$`date`[i], nrow(df_)), df_ %>% as.matrix))
}
}
df_r = data.frame(df_r, stringsAsFactors = FALSE, row.names = 1:nrow(df_r))
names(df_r) = c('CUTOFF_TIME','time','mean','lower','upper')
df_r %<>% mutate(CUTOFF_TIME = as.Date(CUTOFF_TIME)) %>% mutate_each(as.numeric, -CUTOFF_TIME) %>% mutate(date = t0+time)
tail(df_r)
date_max = df_r$date %>% max
date_max
for (i in 1:nrow(df)) {
df_ = read.csv(drname%&%"/"%&%df$filename[i])
df_ = df_[df_$var=='CFR',] %>% select(-var)
if (i==1) {
df_CFR = df_ %>% as.matrix
df_CFR = cbind(rep(df$`date`[i], nrow(df_)), df_CFR)
} else {
df_CFR = rbind(df_CFR, cbind(rep(df$`date`[i], nrow(df_)), df_ %>% as.matrix))
}
}
df_CFR = data.frame(df_CFR, stringsAsFactors = FALSE, row.names = 1:nrow(df_CFR))
names(df_CFR) = c('CUTOFF_TIME','time','mean','lower','upper')
t_max = df_CFR$`time` %>% as.numeric %>% max
df_CFR %<>% mutate(CUTOFF_TIME = as.Date(CUTOFF_TIME)) %>% mutate_each(list(~as.numeric(.)*100), -CUTOFF_TIME) %>%
group_by(CUTOFF_TIME) %>%
mutate(time = time/100, date = CUTOFF_TIME-max(time)+time) %>%
ungroup
tail(df_CFR,5)
# +
cs = c(8, 10)
options(repr.plot.width=cs[1],repr.plot.height=cs[2])
xmax = max(df_r$CUTOFF_TIME)
df_r %>% mutate(CUTOFF_TIME = as.factor(CUTOFF_TIME)) %>%
ggplot(aes(x=date, y=mean, group = desc(CUTOFF_TIME), color=CUTOFF_TIME)) +
geom_ribbon(data=df_r, aes(x=date, ymin = lower, ymax = upper, group=CUTOFF_TIME),
alpha = 0.2, fill="grey70", inherit.aes=FALSE) +
geom_line(aes(y=mean), size=1) +
scale_color_brewer(palette = "Spectral", direction=-1) +
xlab("Date of report (month day)") +
ylab("Estimated cumulative incidence\n") +
guides(color=guide_legend(ncol=2,
title = "Ending timepoint (year-month-day)")) +
theme(plot.margin = unit(c(.5,.5,1,.25),"lines"),
text = element_text(family="sans",color="black"),
axis.text.x = element_text(angle = 35, hjust = 1),
axis.text =element_text(size=fnt_size, family="sans",color="black"),
panel.grid.major = element_blank(),
panel.grid.minor = element_blank(),
legend.position = c(0.22, 0.8),
axis.title.y = element_text(vjust = .5),
legend.text = element_text(size = 12, family="sans"),
legend.spacing.y = unit(.25, 'cm')) +
scale_x_date(expand=c(0,0), date_labels="%b %d",date_breaks ="1 day",
limits=c(as.Date('2019-12-31'),xmax+.5)) +
scale_y_continuous(expand=c(0,.01)) +
coord_cartesian(ylim = c(0,20250)) -> fig22
df_CFR %>% mutate(CUTOFF_TIME = as.factor(CUTOFF_TIME)) %>%
ggplot(aes(x=date, y=mean, group = CUTOFF_TIME, color=CUTOFF_TIME)) +
geom_ribbon(data=df_CFR, aes(x=date, ymin = lower, ymax = upper, group=CUTOFF_TIME),
alpha = 0.1, fill="grey70", inherit.aes=FALSE) +
geom_line(aes(y=mean), size=1) +
scale_color_brewer(palette = "Spectral", direction=-1) +
xlab("Date of report (month day)") +
ylab("Case fatality (%)\n") +
guides(color=F) + #guide_legend(ncol=2, title = "Timepoint when incidence equates\n to one (year-month-day)")) +
theme(plot.margin = unit(c(0,.5,1,1.5),"lines"),
text = element_text(family="sans",color="black"),
axis.text.x = element_text(angle = 35, hjust = 1),
axis.text = element_text(size=fnt_size, family="sans",color="black"),
axis.title.y = element_text(vjust = 4.5),
axis.title = element_text(size=fnt_size+1, family="sans",color="black"),
panel.grid.major = element_blank(),
panel.grid.minor = element_blank()) +
scale_x_date(expand=c(0,0), date_labels="%b %d",date_breaks ="1 day",
limits=c(as.Date('2019-12-31'),xmax+.5)) +
scale_y_continuous(expand=c(0,.01)) +
coord_cartesian(ylim = c(0,33)) -> fig23
pFinal = grid.arrange(fig22, fig23, heights=c(1.75,1), nrow=2, ncol=1);
ggsave(plot=pFinal, width=cs[1], height=cs[2],filename="../../figures/draft/figS32.pdf",useDingbats=FALSE)
# -
| scripts/E3b. Figure - sensitivity for CUTOFF_TIME - Scenario 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Create Wayback Machine Snapshots for references
#
# An eLife editor [provided us](https://github.com/greenelab/scihub-manuscript/issues/34) with the following comment:
#
# > Also, for those items that are not references to journal articles, please ensure that the following information is included: Author, year, title, doi or url, date accessed (in YYYY-MM-DD).
#
# However, most references have been accessed on multiple dates, and we have not kept track of this information. Therefore, we thought it'd make sense to access all articles on a given day and take a snapshot of them via the Internet Archive.
#
# See also [Manubot issue](https://github.com/greenelab/manubot-rootstock/issues/85) related to implementing Internet Archive snapshots.
# +
import datetime
import requests
import time
import pandas
# -
print(f'The current time is {datetime.datetime.now()}')
# Read references.json from the Manubot output branch
url = "https://github.com/greenelab/scihub-manuscript/raw/8cd4af0793ccb75c8230763b33869da97d0219d8/references.json"
references = requests.get(url).json()
# Create a dataframe of URLs
reference_df = pandas.DataFrame(references)[['id', 'URL']]
reference_df.head(4)
def create_url_snapshot(url):
"""
Create a snapshot of a URL in Internet Archive's wayback machine.
Returns the snapshot URL.
"""
archive_url = f'http://web.archive.org/save/{url}'
try:
response = requests.get(archive_url)
except Exception:
return None
if response.status_code != 200:
print(f'Failed: {response.url}')
return None
location = response.headers.get('Content-Location')
if not location:
return None
return f'http://web.archive.org{location}'
snapshots = dict()
for url in reference_df.URL:
if url in snapshots:
continue
snapshots[url] = create_url_snapshot(url)
time.sleep(2)
# #### Failures
#
# Certain URLs could not be archived. For example,
#
# + http://web.archive.org/save/http://doi.wiley.com/10.1002/asi.23445 failed due to "Page cannot be displayed due to robots.txt"
reference_df['snapshot'] = reference_df.URL.map(snapshots)
reference_df.head(4)
reference_df.to_csv('reference-snapshots.tsv', sep='\t', index=False)
| custom/archive-references.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
import os
import natsort
import numpy as np
import pickle
out_dir = "/home/amridul/Documents/forward_project/data/Yumi"
in_dir = "/home/amridul/Documents/forward_project/data/Yumi"
subjects = os.listdir(in_dir)
def findClosestTimeStamp(ts, timeStamps):
lo, hi = 0, len(timeStamps)-1
while(lo <= hi):
mid = int((lo+hi)/2)
if(timeStamps[mid] == ts):
return mid
elif (timeStamps[mid] > ts):
hi = mid-1
elif (timeStamps[mid] < ts):
lo = mid+1
if(lo >= len(timeStamps)):
return hi
if(hi < 0):
return lo
if(np.abs(timeStamps[lo] - ts) < np.abs(timeStamps[hi] - ts)):
return lo
return hi
def dissectKinematicSequence(sub, trial):
fileNameLeft = sub+'_'+trial+'_yumi_left.txt'
fileNameRight = sub+'_'+trial+'_yumi_right.txt'
fileNameTSFrame = sub+'_'+trial+'_color_ts.txt'
fileNameAnnotate = sub+'_'+trial+'_color_annot.txt'
print(fileNameLeft, fileNameRight, fileNameTSFrame, fileNameAnnotate)
frameTSArray = []
subDir = os.path.join(in_dir, sub)
with open(os.path.join(subDir, fileNameTSFrame)) as fTS2Frame:
for line in fTS2Frame:
ts = line.split('\n')[0]
frameTSArray.append(float(ts))
frameTSArray = np.array(frameTSArray)
surgemeStartFrame = []
surgemeEndFrame = []
surgemeIndex = []
surgemeSuccess = []
dominantHand = []
surgmemeParams = []
segments = 0
with open(os.path.join(subDir, fileNameAnnotate)) as fAnnotate:
for line in fAnnotate:
start, end, surgeme, success, domHand, param = line.split(' ')
surgemeStartFrame.append(int(start))
surgemeEndFrame.append(int(end))
surgemeIndex.append(int(surgeme[1]))
surgemeSuccess.append(bool(success))
dominantHand.append(int(domHand))
surgmemeParams.append(int(param))
segments += 1
leftTS = []
leftLines = open(os.path.join(subDir, fileNameLeft)).readlines()
for line in leftLines:
parts = line.split(',')[0]
ts = float(parts)
leftTS.append(ts)
rightTS = []
rightLines = open(os.path.join(subDir, fileNameRight)).readlines()
for line in rightLines:
parts = line.split(',')[0]
ts = float(parts)
rightTS.append(ts)
X = []
Y = []
for frame in range(1, surgemeEndFrame[-1]+1):
surgeme = 0
for segment in range(segments):
if(frame in range(surgemeStartFrame[segment], surgemeEndFrame[segment]+1)):
surgeme = surgemeIndex[segment]
break
frameTS = frameTSArray[frame-1]
leftTSIndex = findClosestTimeStamp(frameTS, leftTS)
leftTimeStamp = leftTS[leftTSIndex]
rightTSIndex = findClosestTimeStamp(frameTS, rightTS)
rightTimeStamp = rightTS[rightTSIndex]
leftLine = leftLines[leftTSIndex]
rightLine = rightLines[rightTSIndex]
leftLine = leftLine.replace("\n", '')
leftLine = leftLine.replace("\r", '')
leftLine = leftLine.replace(" ", '')
rightLine = rightLine.replace("\n", '')
rightLine = rightLine.replace("\r", '')
rightLine = rightLine.replace(" ", '')
leftParts = leftLine.split(',')
rightParts = rightLine.split(',')
leftValid = leftParts[-1]
rightValid = rightParts[-1]
kinData = []
for parts in [leftParts, rightParts]:
for part in parts[1:]:
try:
data = float(part)
kinData.append(data)
except:
if(part in ['True', 'False']):
kinData.append(float(bool(part)))
else:
pre, post = part.split('-')
data = float(pre)
kinData.append(data)
data = float(post)
kinData.append(data)
vecSurgeme = np.zeros(8)
vecSurgeme[surgeme] = 1
X.append(kinData)
Y.append(vecSurgeme)
return X, Y
lineNum = 0
arms = ['left', 'right']
for sub in subjects:
trials = ['T1', 'T2', 'T3', 'T4', 'T5', 'T6']
if(sub == 'S8'):
trials = ['T1', 'T2', 'T3', 'T4']
for trial in trials:
X, Y = dissectKinematicSequence(sub, trial)
fileNameX = os.path.join(os.path.join(in_dir, sub), sub+'_'+trial+'_frame_kinematic_data.pkl')
fileNameY = os.path.join(os.path.join(in_dir, sub), sub+'_'+trial+'_frame_surgeme_data.pkl')
pickle_handle = open(fileNameX, 'wb')
pickle.dump(X, pickle_handle)
pickle_handle.close()
pickle_handle = open(fileNameY, 'wb')
pickle.dump(Y, pickle_handle)
pickle_handle.close()
| Generate_Kinematic_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a id='HOME'></a>
# # CHAPTER 11 Concurrency and Networks
# ## 開發與網路
#
# * [11.1 開發](#Concurrency)
# * [11.2 網路](#Networks)
#
# Time is nature’s way of keeping everything from happening at once.
# Space is what preventseverything from happening to me.
# — Quotes about Time
#
# 先前的範例都是在一台電腦一次運行一次,但是可以利用技術做到分散式計算等更強大的功能。
#
# 好處有
# * 提升性能,可以不用因為某部分運行較慢而塞住後面的程序
# * 提升安全性,同時運作多個相同任務防止意外
# * 把程式拆解得更細小更好維護與理解
# * 網路傳送與接收數據
# ---
# <a id='Concurrency'></a>
# ## 11.1 開發
# [回目錄](#HOME)
#
# 一般而言在計算時速度變慢時多半是因為__I/O___的限制以及__CPU限制__
# I/O的速度比起CPU計算上慢上許多,
# CPU算得較慢時可能是遇到科學相關需要大量計算或是圖形相關的計算導致變慢
#
# 程式在開發時一般會有兩種技術,__同步(synchronous)__與__非同步(asynchronous)__
#
# * 同步是指說程式在執行時是一個接著一個,A指令做完才做B指令,最後在C指令
# * 非同步是可以讓A B C三種指令同時在一個程式裡執行
# __佇列(Queue)__中文也翻作隊列,顧名思義是一種像排隊一樣的概念,
# 以生活中的情況為人們一個接一個的從隊伍後面加入排隊,而窗口的服務人員則從最前面的民眾一個接一個處理事務。(http://emn178.pixnet.net/blog/post/93475832-%E4%BD%87%E5%88%97(queue))
#
# 本書作者以洗碗流程作為範例,分成兩段工作過程,清洗與烘乾。
# 假設一個人做事,可以有兩種做法,
# 1. 洗完一個盤子就拿去烘乾
# 2. 先洗完全部的盤子,在統一把盤子烘乾
#
# 若要改善效率最快的方法就是找幫手,一人負責洗碗一人負責烘乾,但是這樣會遇到一個問題,如果洗盤子的速度大於烘乾盤子時,是要等待烘乾的人員閒置後在遞交盤子給他,還是先行放置在桌上,在繼續洗下一個盤子,等他有空時再自行拿取,前者就是同步的概念,後者則為非同步的概念。
#
# 假設水槽中的盤子們是佇列(Queue)中的工作項目,可以進行同步與分同步的工作流程。
# * 同步:把水槽中的髒盤子給第一個閒置的洗碗人員洗,洗完後等烘乾人員閒置後再把盤子給他
# * 非同步:洗盤子的人員洗好就將盤子放置在桌上後繼續清洗下一個,烘乾人員閒置時就去看桌上有無盤子可以清洗。
#
# 
# #### 原始洗碗流程
#
# ```python
# import os
# import time
# from datetime import datetime
#
# def washer(dishes, now_):
# for dish in dishes:
# now = datetime.now()
# print('Washing', dish, ', time:', now - now_, ', pid', os.getpid())
# time.sleep(1)
# dryer(dish, now_)
#
# def dryer(dish, now_):
# now = datetime.now()
# print('Drying ', dish, ', time:', now - now_, ', pid', os.getpid())
# time.sleep(2)
#
# if __name__ == "__main__":
# now_ = datetime.now()
# dishes = ['dish-1', 'dish-2', 'dish-3', 'dish-4']
# washer(dishes, now_)
# ```
# + jupyter={"outputs_hidden": false}
import subprocess
ret = subprocess.getoutput('python Data/dishes.py')
print(ret) #執行需時間,請等候
# -
# ### 行程or處理程序(Processes)
#
# [wiki 行程](https://zh.wikipedia.org/wiki/%E8%A1%8C%E7%A8%8B)
#
# 下列範例可以模擬兩個人員在進行分工合作,洗碗為主行程,烘乾則為另開的行程,所以洗碗者不必等到烘乾完畢就可以洗下一個盤子
#
# ```python
# import multiprocessing as mp
# import os
# import time
# from datetime import datetime
#
# def washer(dishes, output, now_):
# for dish in dishes:
# now = datetime.now()
# print('Washing', dish, ', time:', now - now_, ', pid', os.getpid())
# time.sleep(1)
# #把東西丟給其他行程後繼續執行下一個
# output.put(dish)
#
# def dryer(input, now_):
# while True:
# dish = input.get()
# now = datetime.now()
# print('Drying ', dish, ', time:', now - now_, ', pid', os.getpid())
# time.sleep(2)
# input.task_done()
#
# if __name__ == "__main__":
# now_ = datetime.now()
# #建立佇列
# dish_queue = mp.JoinableQueue()
# #創建行程(烘乾人員)
# dryer_proc = mp.Process(target=dryer, args=(dish_queue, now_,))
# dryer_proc.daemon = True
# #啟動行程(上班囉)
# dryer_proc.start()
# #time.sleep(1)
#
# dishes = ['dish-1', 'dish-2', 'dish-3', 'dish-4']
# washer(dishes, dish_queue, now_)
# dish_queue.join()
# ```
# p.s. 在ipython中非主程式的行程print不出來,請自行在本機端cmd跑,或是把print改成寫成實體檔案方可看見結果,
# Data資料夾中有dishes_process.py檔案可供使用
#
# 結果:
# Washing dish-1 , time: 0:00:00.037144 , pid 10480
# Washing dish-2 , time: 0:00:01.047415 , pid 10480
# Drying dish-1 , time: 0:00:01.047415 , pid 7280
# Washing dish-3 , time: 0:00:02.060229 , pid 10480
# Drying dish-2 , time: 0:00:03.047613 , pid 7280
# Washing dish-4 , time: 0:00:03.063241 , pid 10480
# Drying dish-3 , time: 0:00:05.047959 , pid 7280
# Drying dish-4 , time: 0:00:07.053659 , pid 7280
# ### 執行緒(Threads)
#
# [wiki 執行緒](https://zh.wikipedia.org/wiki/%E7%BA%BF%E7%A8%8B)
#
# 下列範例可以模擬一個人員洗,兩個人烘的分工合作,全部都為同一個主行程,但是另開兩個線程來處理烘乾工作
#
# ```python
#
# import threading, queue
# import os
# import time
# from datetime import datetime
#
# def washer(dishes, dish_queue, now_):
# for dish in dishes:
# now = datetime.now()
# print ("Washing", dish, now - now_, ', pid', os.getpid(), threading.current_thread())
# time.sleep(1)
# dish_queue.put(dish)
#
# def dryer(dish_queue, now_):
# while True:
# dish = dish_queue.get()
# now = datetime.now()
# print ("Drying ", dish, now - now_, ', pid', os.getpid(), threading.current_thread())
# time.sleep(2)
# dish_queue.task_done()
#
# if __name__ == "__main__":
# dish_queue = queue.Queue()
# now_ = datetime.now()
# #控制要開幾條執行緒
# for n in range(2):
# dryer_thread = threading.Thread(target=dryer, args=(dish_queue, now_))
# dryer_thread.daemon = True
# dryer_thread.start()
#
# dishes = ['dishe-1', 'dishe-2', 'dishe-3', 'dishe-4']
# washer(dishes, dish_queue, now_)
# dish_queue.join()
#
# ```
# + jupyter={"outputs_hidden": false}
import subprocess
ret = subprocess.getoutput('python Data/dishes_threads.py')
print(ret) #執行需時間,請等候
# -
# 總而言之,對於 Python,建議如下:
# * 使用執行緒來解決 I/O 限制問題;
# * 使用行程、網絡或者事件(下一節會介紹)來處理 CPU 限制問題。
# 書中還介紹許多函示庫可以辦到各種不同的佇列(Queue)功能
# * gevent
# * twisted
# * asyncio
# * Redis
# ---
# <a id='Networks'></a>
# ## 11.2 網路
# [回目錄](#HOME)
| CHAPTER 11 Concurrency and Networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.metrics import classification_report, confusion_matrix, roc_curve, roc_auc_score, auc, RocCurveDisplay, ConfusionMatrixDisplay, plot_roc_curve, plot_confusion_matrix
from sklearn.model_selection import cross_val_score, KFold, StratifiedKFold, GridSearchCV, RandomizedSearchCV
X_train = pd.read_csv(r'C:\Users\2bogu\Desktop\Sringboard_Materials\capstone2\data\interim\X_train_bad')
X_test = pd.read_csv(r'C:\Users\2bogu\Desktop\Sringboard_Materials\capstone2\data\interim\X_test_bad')
y_train = pd.read_csv(r'C:\Users\2bogu\Desktop\Sringboard_Materials\capstone2\data\interim\y_train_bad')
y_test = pd.read_csv(r'C:\Users\2bogu\Desktop\Sringboard_Materials\capstone2\data\interim\y_test_bad')
# # To Do
#
# Add each metric to a list so it can be graphed
#
# change crossval to use train values
#
# create graphs for metrics to justify model selection
#
# drop deaths
#
#
# # Classification: Good vs Bad
# - Logistic Regression: as a base line. set param class_weight=balenced
# - KNN
# - Random Forest: Boosted, and other variants
#
# Bad is defined as Has been canceled or has given notice that it will br canceled
#
# Minimizing false positives(false bad prediction)
#
# Heavily inblalenced data, accuracy not a helpful measure
#
# Split train and test set to give equal proportion of good and bad? (DONE)
#
# Oversampling/ Undersampling/ hybrid sampling ?
#
# Drop Dead people??
#
# Maybe this https://imbalanced-learn.readthedocs.io/en/stable/generated/imblearn.over_sampling.SMOTE.html
#
# Wouldn't preprocessing also be in this notebook?
#
# #### Entropy vs Gini for each
#
# #### Hyperparam tuning comes last?
#
#
X_train = X_train.values
X_test = X_test.values
y_train = y_train.values.ravel()
y_test= y_test.values.ravel()
# +
from imblearn.over_sampling import SMOTE, ADASYN
X_smo, y_smo = SMOTE(random_state=2).fit_resample(X_train, y_train)
X_ada, y_ada = ADASYN(random_state=2).fit_resample(X_train, y_train)
# +
#some helper functions for my sanity
def classify(model, X_train, y_train, X_test=X_test ):
'''Takes in model, X_train, X_test, y_train
returns tuple of (fit model, y_pred, predict_proba[:,1])'''
m = model.fit(X_train, y_train)
pred = model.predict(X_test)
prob = model.predict_proba(X_test)[:,1]
return (m, pred, prob)
def model_cv(model, X=X_train, y=y_train, cv=5, scoring='roc_auc'):
'''takes the model, X_train, y_train, cv(=5), scoring(='roc_auc')
returns array of scores, prints array, mean, std of scores scores'''
scores = cross_val_score(model, X, y, cv=cv, scoring=scoring)
print(str(model))
print('The scores are', scores)
print('The mean is', scores.mean())
print('The STD is', scores.std())
return (scores)
def model_cv_smo(model, X=X_smo, y=y_smo, cv=5, scoring='roc_auc'):
'''takes the model, X_train, y_train, cv(=5), scoring(='roc_auc')
returns array of scores, prints array, mean, std of scores scores'''
scores = cross_val_score(model, X, y, cv=cv, scoring=scoring)
print(str(model), 'w/ SMOTE')
print('The scores are', scores)
print('The mean is', scores.mean())
print('The STD is', scores.std())
return (scores)
def model_cv_ada(model, X=X_ada, y=y_ada, cv=5, scoring='roc_auc'):
'''takes the model, X_train, y_train, cv(=5), scoring(='roc_auc')
returns array of scores, prints array, mean, std of scores scores'''
scores = cross_val_score(model, X, y, cv=cv, scoring=scoring)
print(str(model), 'w/ ADASYN')
print('The scores are', scores)
print('The mean is', scores.mean())
print('The STD is', scores.std())
return (scores)
def find_threshold(probs, y_test=y_test):
'''takes predict_proba[:,1] for certain model, y_test is set because imonly doing this for the one y_test
uses Youdens J statistic
prints the threshold
returns new predictions given the threshold'''
fpr, tpr, thresholds = roc_curve(y_test, probs)
gmeans = tpr - fpr
thresh = thresholds[np.argmax(gmeans)]
print('The threshold is', thresh)
return (probs >= thresh).astype(int)
# +
model_names = ['Log Reg', 'Log Reg w/ SMOTE', 'Log Reg w/ ADASYN',
'KNN', 'KNN w/ SMOTE','KNN w/ ADASYN',
'Random Forest', 'Random Forest w/ SMOTE', 'Random Forest w/ ADASYN',
'Gradient Boost', 'Gradient Boost w/ SMOTE', 'Gradient Boost w/ ADASYN']
cv_scores_roc = []
cv_scores_f1 = []
# -
# ## Logistic Regression
# +
#from sklearn.linear_model import LogisticRegressionCV
from sklearn.linear_model import LogisticRegression
lr, pred_lr, prob_lr= classify(LogisticRegression(random_state=2, n_jobs=-1), X_train, y_train)
lr_smo, pred_lr_smo, prob_lr_smo= classify(LogisticRegression(random_state=2, n_jobs=-1), X_smo, y_smo)
lr_ada, pred_lr_ada, prob_lr_ada= classify(LogisticRegression(random_state=2, n_jobs=-1), X_ada, y_ada)
# -
# def con_mats(graph_info, y_test=y_test):
# '''accepts list of 3 tuples each with y_pred, title, and axes in that order. and a y_test to test them on
# plots confusion matrixes in a line'''
# fig, (ax1, ax2, ax3) = plt.subplots(1, len(axes), figsize=(20,5))
#
# for pred, title, ax in graph_info:
# cm = confusion_matrix(y_test, pred)
# cm_dis = ConfusionMatrixDisplay(cm).plot(ax=ax, cmap=plt.cm.Blues)
# cm_dis.ax_.set_title(title)
#
# plt.subplots_adjust(wspace=0.08)
#
# plt.show()
#
#
# axes = (ax1, ax2, ax3)
#
# this = [(pred_lr, 'Logistic Regression (LR)', ax1), (pred_lr_smo, 'LR w/ SMOTE', ax2), (pred_lr_ada, 'LR w/ ADASYN', ax3)]
#
# con_mats(this, lr_titles)
# +
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(20,5))
cm1 = confusion_matrix(y_test, pred_lr)
cm2 = confusion_matrix(y_test, pred_lr_smo)
cm3 = confusion_matrix(y_test, pred_lr_ada)
cm1_dis = ConfusionMatrixDisplay(cm1).plot(ax=ax1,cmap=plt.cm.Blues)
cm2_dis = ConfusionMatrixDisplay(cm2).plot(ax=ax2, cmap=plt.cm.Blues)
cm3_dis = ConfusionMatrixDisplay(cm3).plot(ax=ax3, cmap=plt.cm.Blues)
ax1.set_title('Logistic Regression(LR), no resampling')
ax2.set_title("LR w/ SOMTE")
ax3.set_title('LR w/ ADASYN')
plt.subplots_adjust(wspace=0.08)
plt.show()
# +
fig = plt.figure(figsize=(7,7))
lw = 2
plt.plot([0, 1], [0, 1], color='navy', lw=lw, linestyle='--')
fpr, tpr, _ = roc_curve(y_test, prob_lr)
auc = roc_auc_score(y_test, prob_lr)
plt.plot(fpr, tpr, color='darkorange', lw=lw, label='LR no resampling (area = %0.2f)' % auc)
fpr, tpr, _ = roc_curve(y_test, prob_lr_smo)
auc = roc_auc_score(y_test, prob_lr_smo)
plt.plot(fpr, tpr, color='darkred', lw=lw, label='LR w/ SMOTE(area = %0.2f)' % auc)
fpr, tpr, _ = roc_curve(y_test, prob_lr_ada)
auc = roc_auc_score(y_test, prob_lr_ada)
plt.plot(fpr, tpr, color='darkgreen', lw=lw, label='LR w/ ADASYN (area = %0.2f)' % auc)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic (ROC)')
plt.legend(loc="lower right")
plt.show()
# +
#plot_roc_curve(lr, X_test, y_test)
#plot_roc_curve(lr_smo, X_test, y_test)
#plot_roc_curve(lr_ada, X_test, y_test)
# -
print('No resampling')
model_cv(lr)
print('')
print('SMOTE')
model_cv_smo(lr_smo)
print('')
print('ADASYN')
model_cv_ada(lr_ada)
skf = StratifiedKFold(n_splits=5)
print('With Stratified K Fold CV')
print('')
print('No resampling')
model_cv(lr,cv=skf)
print('')
print('SMOTE')
model_cv_smo(lr_smo, cv=skf)
print('')
print('ADASYN')
model_cv_ada(lr_ada, cv=skf)
# +
fpr, tpr, thresholds = roc_curve(y_test, prob_lr)
gmeans = tpr - fpr
thresh_lr = thresholds[np.argmax(gmeans)]
pred_lr_thsh = (prob_lr >= thresh_lr).astype(int)
fpr, tpr, thresholds = roc_curve(y_test, prob_lr_smo)
gmeans = tpr - fpr
thresh_lr_smo = thresholds[np.argmax(gmeans)]
pred_lr_smo_thsh = (prob_lr_smo >= thresh_lr_smo).astype(int)
fpr, tpr, thresholds = roc_curve(y_test, prob_lr_ada)
gmeans = tpr - fpr
thresh_lr_ada = thresholds[np.argmax(gmeans)]
pred_lr_ada_thsh = (prob_lr_ada >= thresh_lr_ada).astype(int)
# -
print('Regular')
print(classification_report(y_test, pred_lr_thsh),'')
print('SMOTE')
print(classification_report(y_test, pred_lr_smo_thsh),'')
print('ADASYN')
print(classification_report(y_test, pred_lr_ada_thsh),'')
# ## KNN
# +
from sklearn.neighbors import KNeighborsClassifier
#param_grid = {'n_neighbors':np.arange(1,10)}
#knn_cv= GridSearchCV(knn,param_grid,cv=5)
knn, pred_knn, prob_knn = classify(KNeighborsClassifier(n_jobs=-1), X_train, y_train)
knn_smo, pred_knn_smo, prob_knn_smo= classify(KNeighborsClassifier(n_jobs=-1), X_smo, y_smo)
knn_ada, pred_knn_ada, prob_knn_ada= classify(KNeighborsClassifier(n_jobs=-1), X_ada, y_ada)
# +
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(20,5))
cm1 = confusion_matrix(y_test, pred_knn)
cm2 = confusion_matrix(y_test, pred_knn_smo)
cm3 = confusion_matrix(y_test, pred_knn_ada)
cm1_dis = ConfusionMatrixDisplay(cm1).plot(ax=ax1,cmap=plt.cm.Blues)
cm2_dis = ConfusionMatrixDisplay(cm2).plot(ax=ax2, cmap=plt.cm.Blues)
cm3_dis = ConfusionMatrixDisplay(cm3).plot(ax=ax3, cmap=plt.cm.Blues)
ax1.set_title('K Nearest Neighbors, no resampling')
ax2.set_title("KNN w/ SOMTE")
ax3.set_title('KNN w/ ADASYN')
plt.subplots_adjust(wspace=0.08)
plt.show()
# +
#print(classification_report(y_test, y_pred_knn))
#print(classification_report(y_test, y_pred_knn_smo))
#print(classification_report(y_test, y_pred_knn_ada))
# +
fig = plt.figure(figsize=(7,7))
lw = 2
plt.plot([0, 1], [0, 1], color='navy', lw=lw, linestyle='--')
fpr, tpr, _ = roc_curve(y_test, prob_knn)
auc = roc_auc_score(y_test, prob_knn)
plt.plot(fpr, tpr, color='darkorange', lw=lw, label='KNN no resampling (area = %0.2f)' % auc)
fpr, tpr, _ = roc_curve(y_test, prob_knn_smo)
auc = roc_auc_score(y_test, prob_knn_smo)
plt.plot(fpr, tpr, color='darkred', lw=lw, label='KNN w/ SMOTE(area = %0.2f)' % auc)
fpr, tpr, _ = roc_curve(y_test, prob_knn_ada)
auc = roc_auc_score(y_test, prob_knn_ada)
plt.plot(fpr, tpr, color='darkgreen', lw=lw, label='KNN w/ ADASYN (area = %0.2f)' % auc)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic (ROC)')
plt.legend(loc="lower right")
plt.show()
# -
print('No resampling')
model_cv(knn)
print('')
print('SMOTE')
model_cv_smo(knn)
print('')
print('ADASYN')
model_cv_ada(knn)
skf = StratifiedKFold(n_splits=5)
print('With Stratified K Fold CV')
print('')
print('No resampling')
model_cv(knn,cv=skf)
print('')
print('SMOTE')
model_cv_smo(knn_smo, cv=skf)
print('')
print('ADASYN')
model_cv_ada(knn_ada, cv=skf)
# ## Random Forest
from sklearn.ensemble import RandomForestClassifier
# +
rf, pred_rf, prob_rf = classify(RandomForestClassifier(random_state=2, n_jobs=-1), X_train, y_train)
rf_smo, pred_rf_smo, prob_rf_smo= classify(RandomForestClassifier(random_state=2, n_jobs=-1), X_smo, y_smo)
rf_ada, pred_rf_ada, prob_rf_ada= classify(RandomForestClassifier(random_state=2, n_jobs=-1), X_ada, y_ada)
# +
#print('Regular')
#print(classification_report(y_test, y_pred_clf),'')
#print('SMOTE')
#print(classification_report(y_test, y_pred_clf_smo),'')
#print('ADASYN')
#print(classification_report(y_test, y_pred_clf_ada),'')
# +
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(20,5))
cm1 = confusion_matrix(y_test, pred_rf)
cm2 = confusion_matrix(y_test, pred_rf_smo)
cm3 = confusion_matrix(y_test, pred_rf_ada)
cm1_dis = ConfusionMatrixDisplay(cm1).plot(ax=ax1,cmap=plt.cm.Blues)
cm2_dis = ConfusionMatrixDisplay(cm2).plot(ax=ax2, cmap=plt.cm.Blues)
cm3_dis = ConfusionMatrixDisplay(cm3).plot(ax=ax3, cmap=plt.cm.Blues)
ax1.set_title('Random Forest, no resampling')
ax2.set_title("Random Forest w/ SOMTE")
ax3.set_title('Random Forest w/ ADASYN')
plt.subplots_adjust(wspace=0.08)
plt.show()
# +
fig = plt.figure(figsize=(7,7))
lw = 2
plt.plot([0, 1], [0, 1], color='navy', lw=lw, linestyle='--')
fpr, tpr, _ = roc_curve(y_test, prob_rf)
auc = roc_auc_score(y_test, prob_rf)
plt.plot(fpr, tpr, color='darkorange', lw=lw, label='RF no resampling (area = %0.2f)' % auc)
fpr, tpr, _ = roc_curve(y_test, prob_rf_smo)
auc = roc_auc_score(y_test, prob_rf_smo)
plt.plot(fpr, tpr, color='darkred', lw=lw, label='RF w/ SMOTE(area = %0.2f)' % auc)
fpr, tpr, _ = roc_curve(y_test, prob_rf_ada)
auc = roc_auc_score(y_test, prob_rf_ada)
plt.plot(fpr, tpr, color='darkgreen', lw=lw, label='RF w/ ADASYN (area = %0.2f)' % auc)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic (ROC)')
plt.legend(loc="lower right")
plt.show()
# -
print('No resampling')
model_cv(rf)
print('')
print('SMOTE')
model_cv_smo(rf_smo)
print('')
print('ADASYN')
model_cv_ada(rf_ada)
skf = StratifiedKFold(n_splits=5)
print('With Stratified K Fold CV')
print('')
print('No resampling')
model_cv(rf,cv=skf)
print('')
print('SMOTE')
model_cv_smo(rf_smo, cv=skf)
print('')
print('ADASYN')
model_cv_ada(rf_ada, cv=skf)
# # Gradient Boosting
# +
from sklearn.ensemble import GradientBoostingClassifier
gb, pred_gb, prob_gb = classify(GradientBoostingClassifier(random_state=2), X_train, y_train)
gb_smo, pred_gb_smo, prob_gb_smo= classify(GradientBoostingClassifier(random_state=2), X_smo, y_smo)
gb_ada, pred_gb_ada, prob_gb_ada= classify(GradientBoostingClassifier(random_state=2), X_ada, y_ada)
# +
#print(classification_report(y_test, y_pred_gb))
#print(classification_report(y_test, y_pred_gb_smo))
#print(classification_report(y_test, y_pred_gb_ada))
# +
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(20,5))
cm1 = confusion_matrix(y_test, pred_gb)
cm2 = confusion_matrix(y_test, pred_gb_smo)
cm3 = confusion_matrix(y_test, pred_gb_ada)
cm1_dis = ConfusionMatrixDisplay(cm1).plot(ax=ax1,cmap=plt.cm.Blues)
cm2_dis = ConfusionMatrixDisplay(cm2).plot(ax=ax2, cmap=plt.cm.Blues)
cm3_dis = ConfusionMatrixDisplay(cm3).plot(ax=ax3, cmap=plt.cm.Blues)
ax1.set_title('Random Forest, no resampling')
ax2.set_title("Random Forest w/ SOMTE")
ax3.set_title('Random Forest w/ ADASYN')
plt.subplots_adjust(wspace=0.08)
plt.show()
# +
fig = plt.figure(figsize=(7,7))
lw = 2
plt.plot([0, 1], [0, 1], color='navy', lw=lw, linestyle='--')
fpr, tpr, _ = roc_curve(y_test, prob_gb)
auc = roc_auc_score(y_test, prob_gb)
plt.plot(fpr, tpr, color='darkorange', lw=lw, label='Gradient Boosting no resampling (area = %0.2f)' % auc)
fpr, tpr, _ = roc_curve(y_test, prob_gb_smo)
auc = roc_auc_score(y_test, prob_gb_smo)
plt.plot(fpr, tpr, color='darkred', lw=lw, label='GB w/ SMOTE(area = %0.2f)' % auc)
fpr, tpr, _ = roc_curve(y_test, prob_gb_ada)
auc = roc_auc_score(y_test, prob_gb_ada)
plt.plot(fpr, tpr, color='darkgreen', lw=lw, label='GB w/ ADASYN (area = %0.2f)' % auc)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic (ROC)')
plt.legend(loc="lower right")
plt.show()
# -
print('No resampling')
model_cv(gb)
print('')
print('SMOTE')
model_cv_smo(gb_smo)
print('')
print('ADASYN')
model_cv_ada(gb_ada)
skf = StratifiedKFold(n_splits=5)
print('With Stratified K Fold CV')
print('')
print('No resampling')
model_cv(gb,cv=skf)
print('')
print('SMOTE')
model_cv_smo(gb_smo, cv=skf)
print('')
print('ADASYN')
model_cv_ada(gb_ada, cv=skf)
# +
#took too long, not better than bg and LR or rf
from sklearn.svm import SVC
#svc, pred_svc, prob_svc = classify(SVC(probability=True, random_state=2, n_jobs=-1), X_train, y_train)
#svc_smo, pred_svc_smo, prob_svc_smo= classify(SVC(probability=True, random_state=2, n_jobs=-1), X_smo, y_smo)
#svc_ada, pred_svc_ada, prob_svc_ada= classify(SVC(probability=True, random_state=2, n_jobs=-1), X_ada, y_ada)
# +
#print('Regular')
#print(classification_report(y_test, y_pred_svc),'')
#print('SMOTE')
#print(classification_report(y_test, y_pred_svc_smo),'')
#print('ADASYN')
#print(classification_report(y_test, y_pred_svc_ada),'')
# -
# # Next Steps
#
# - Remove deaths?
# - Threshold setting: APPLY AFTER GRIDSEARCH
# - Gridsearch: lr, lr_ada, gb, gb_ada
# - DROP SMO IN FAVOR OF ADA
# ## Gridsearch
def gs_tuning(model, params, X=X_train, y=y_train, scoring='roc_auc', n_jobs=-1, cv=3):
m = GridSearchCV(estimator=model,
param_grid=params,
scoring=scoring,
n_jobs=n_jobs,
cv=cv)
m.fit(X,y)
#pred = m.predict_proba(X_test)[:,1]
print(m.best_params_,)
print('Train score:', m.best_score_)
print('Test score:', m.score(X_test, y_test))#roc_auc_score(y_test, pred)
print(m.best_estimator_)
# #### Logistic Regression
# +
lr_ = LogisticRegression(random_state=2, n_jobs=-1)
params = {'penalty':['l1', 'l2', 'elasticnet', 'none'],
'C':[0.01,0.1,1,10],
'class_weight':['balanced', None],
'solver':['newton-cg', 'lbfgs', 'liblinear', 'sag', 'saga']} #'l1_ratio':[.1, .3, .5, .7, .9]
gs_tuning(lr_, params)
# +
lr_best_ = LogisticRegression(C=0.1, n_jobs=-1, penalty='l2', random_state=2)
lr_best, pred_lr_best, prob_lr_best = classify(lr_best_, X_train, y_train)
model_cv(lr_best_, X_test, y_test)
# +
# getting J statistic threshold
pred_lr_best_thsh = find_threshold(prob_lr_best)
# -
# #### Logistic Regression SMOTE
# +
lr_ = LogisticRegression(random_state=2, n_jobs=-1)
params = {'penalty':['l1', 'l2', 'elasticnet', 'none'],
'C':[0.01,0.1,1,10],
'class_weight':['balanced', None],
'solver':['newton-cg', 'lbfgs', 'liblinear', 'sag', 'saga'],
'l1_ratio':[.1, .3, .5, .7, .9]} #
gs_tuning(lr_, params, X=X_smo, y=y_smo)
# +
lr_best_smo_ = LogisticRegression(C=0.1,
class_weight='balanced',
l1_ratio=0.7,
n_jobs=-1,
penalty='elasticnet',
random_state=2,
solver='saga')
lr_best_smo, pred_lr_best_smo, prob_lr_best_smo = classify(lr_best_smo_, X_smo, y_smo)
model_cv(lr_best_smo, X_test, y_test)
# -
pred_lr_best_smo_thsh = find_threshold(prob_lr_best_smo)
# #### Logistic Regression ADASYN ?
# +
lr_ = LogisticRegression(random_state=2, n_jobs=-1)
params = {'penalty':['l1', 'l2', 'elasticnet', 'none'],
'C':[0.01,0.1,1,10],
'class_weight':['balanced', None],
'solver':['newton-cg', 'lbfgs', 'liblinear', 'sag', 'saga']} #'l1_ratio':[.1, .3, .5, .7, .9]
gs_tuning(lr_,params,X=X_ada,y=y_ada)
# +
lr_best_ada_ = LogisticRegression(C=0.1, n_jobs=-1, random_state=2, solver='newton-cg')
lr_best_ada, pred_lr_best_ada, prob_lr_best_ada = classify(lr_best_ada_, X_ada, y_ada)
model_cv(lr_best_ada, X_test, y_test)
# -
pred_lr_best_ada_thsh = find_threshold(prob_lr_best_ada)
# +
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(20,5)) #,
cm1 = confusion_matrix(y_test, pred_lr, normalize='true')
cm2 = confusion_matrix(y_test, pred_lr_smo, normalize='true')
cm3 = confusion_matrix(y_test, pred_lr_ada, normalize='true')
cm1_dis = ConfusionMatrixDisplay(cm1).plot(ax=ax1, cmap=plt.cm.Blues)
cm2_dis = ConfusionMatrixDisplay(cm2).plot(ax=ax2, cmap=plt.cm.Blues)
cm3_dis = ConfusionMatrixDisplay(cm3).plot(ax=ax3, cmap=plt.cm.Blues)
ax1.set_title('Logistic Regression(LR), no resampling')
ax2.set_title('LR w/ SMOTE')
ax3.set_title('LR w/ ADASYN')
fig, (ax4, ax5, ax6)= plt.subplots(1, 3, figsize=(20,5))
cm4 = confusion_matrix(y_test, pred_lr_best, normalize='true')
cm5 = confusion_matrix(y_test, pred_lr_best_smo, normalize='true')
cm6 = confusion_matrix(y_test, pred_lr_best_ada, normalize='true')
cm4_dis = ConfusionMatrixDisplay(cm4).plot(ax=ax4, cmap=plt.cm.Blues)
cm5_dis = ConfusionMatrixDisplay(cm5).plot(ax=ax5, cmap=plt.cm.Blues)
cm6_dis = ConfusionMatrixDisplay(cm6).plot(ax=ax6, cmap=plt.cm.Blues)
ax4.set_title('Optimized Linear Regression (LR), no resampling')
ax5.set_title('Optimized LR w/ SMOTE')
ax6.set_title('Optimized LR w/ ADASYN')
fig, (ax7, ax8, ax9) = plt.subplots(1, 3, figsize=(20,5))
cm7 = confusion_matrix(y_test, pred_lr_best_thsh, normalize='true')
cm8 = confusion_matrix(y_test, pred_lr_best_smo_thsh, normalize='true')
cm9 = confusion_matrix(y_test, pred_lr_best_ada_thsh, normalize='true')
cm7_dis = ConfusionMatrixDisplay(cm7).plot(ax=ax7, cmap=plt.cm.Blues)
cm8_dis = ConfusionMatrixDisplay(cm8).plot(ax=ax8, cmap=plt.cm.Blues)
cm9_dis = ConfusionMatrixDisplay(cm9).plot(ax=ax9, cmap=plt.cm.Blues)
ax7.set_title('Optimized Logistic Regression(LR) w/ J stat threshold')
ax8.set_title('Optimized LR w/ SMOTE, J stat threshold')
ax9.set_title('Optimized LR w/ ADASYN, J stat threshold')
plt.subplots_adjust(wspace=0.08)
# -
print('Regular')
print(classification_report(y_test, pred_lr_best),'')
print('SMOTE')
print(classification_report(y_test, pred_lr_best_smo),'')
print('ADASYN')
print(classification_report(y_test, pred_lr_best_ada),'')
print('WITH THRESHOLD')
print('Regular')
print(classification_report(y_test, pred_lr_thsh),'')
print('SMOTE')
print(classification_report(y_test, pred_lr_smo_thsh),'')
print('ADASYN')
print(classification_report(y_test, pred_lr_ada_thsh),'')
# +
# ADD DOTS FOR THRESHOLDS?
fig = plt.figure(figsize=(7,7))
lw = 2
plt.plot([0, 1], [0, 1], color='black', lw=lw, linestyle='--')
fpr, tpr, _ = roc_curve(y_test, prob_lr_best)
auc = roc_auc_score(y_test, prob_lr_best)
plt.plot(fpr, tpr, color='darkorange', lw=lw, label='Best LR (area = %0.2f)' % auc)
fpr, tpr, _ = roc_curve(y_test, prob_lr_best_smo)
auc = roc_auc_score(y_test, prob_lr_best_smo)
plt.plot(fpr, tpr, color='darkred', lw=lw, label='Best LR /w SMOTE(area = %0.2f)' % auc)
fpr, tpr, _ = roc_curve(y_test, prob_lr_best_ada)
auc = roc_auc_score(y_test, prob_lr_best_ada)
plt.plot(fpr, tpr, color='darkgreen', lw=lw, label='Best LR w/ ADASYN(area = %0.2f)' % auc)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic (ROC)')
plt.legend(loc="lower right")
plt.show()
# -
# #### Gradient Boosted
# +
params1 = {'n_estimators': range(20,81,10)}
gb_gs1_ = GridSearchCV(estimator = GradientBoostingClassifier(random_state=2),
param_grid = params1,
scoring='roc_auc',
n_jobs=-1,
cv=3)
gb_gs1 = gb_gs1_.fit(X_train, y_train)
gb_gs1.best_params_, gb_gs1.best_score_, gb_gs1.best_estimator_,
# +
params2 = {'max_depth': range(2,7) ,
'min_samples_split': np.linspace(.1,1,10)}
gb_gs2 = GridSearchCV(estimator = GradientBoostingClassifier(random_state=2,
n_estimators=60),
param_grid = params2,
scoring='roc_auc',
n_jobs=-1,
cv=3)
gb_gs2.fit(X_train,y_train)
gb_gs2.best_params_, gb_gs2.best_score_, gb_gs2.best_estimator_,
# +
params3 = {'min_samples_leaf': range(1,7) ,
'max_features': ['sqrt', 'log2', None]}
gb_gs3 = GridSearchCV(estimator = GradientBoostingClassifier(random_state=2,
n_estimators=60,
max_depth=3,
min_samples_split=0.7),
param_grid = params3,
scoring='roc_auc',
n_jobs=-1,
cv=3)
gb_gs3.fit(X_train,y_train)
gb_gs3.best_params_, gb_gs3.best_score_, gb_gs3.best_estimator_,
# +
params4 = {'subsample':[0.85, 0.9, 0.95, 1.0, 1.05, 1.1, 1.15]}
gb_gs4 = GridSearchCV(estimator = GradientBoostingClassifier(random_state=2,
n_estimators=60,
max_depth=3,
min_samples_split=0.7,
min_samples_leaf=2),
param_grid = params4,
scoring='roc_auc',
n_jobs=-1,
cv=3)
gb_gs4.fit(X_train,y_train)
gb_gs4.best_params_, gb_gs4.best_score_, gb_gs4.best_estimator_,
# +
# tuning learning rate, half the learning rate, double n_estimators
params5 = {'learning_rate':[.1, .05, .025], 'n_estimators':[60,120,240]}
gb_gs5 = GridSearchCV(estimator = GradientBoostingClassifier(random_state=2,
n_estimators=60,
max_depth=3,
min_samples_split=0.7,
min_samples_leaf=2),
param_grid = params5,
scoring='roc_auc',
n_jobs=-1,
cv=3)
gb_gs5.fit(X_train,y_train)
gb_gs5.best_params_, gb_gs5.best_score_, gb_gs5.best_estimator_,
# +
gb_best, pred_gb_best, prob_gb_best = classify(GradientBoostingClassifier(learning_rate=0.05,
min_samples_leaf=2,
min_samples_split=0.7,
n_estimators=240,
random_state=2), X_train, y_train)
model_cv(gb_best, X_test, y_test)
# -
# +
from sklearn.metrics import make_scorer
def test_score(model, X, y):
model.fit(X,y)
prob = model.predict_proba(X_test)[:,1]
return roc_auc_score(y_test, prob) # tests against probs from on of the folds, not the whole set refit
my_scorer = make_scorer(test_score, needs_proba=True)
# +
params1 = {'n_estimators': range(50,81,10)} # how do I get gridsearch to care about test score to stop overfitting?
gs_tuning(GradientBoostingClassifier(random_state=2),
params1,
X=X_smo,
y=y_smo)
# +
params1 = {'n_estimators': range(50,81,10)} # how do I get gridsearch to care about test score to stop overfitting?
gs_tuning(GradientBoostingClassifier(random_state=2),
params1,
scoring=test_score,
X=X_smo,
y=y_smo)
# -
# #### Gradient Boosted SMOTE
# +
params1 = {'n_estimators': range(20,81,10)}
gs_tuning(GradientBoostingClassifier(random_state=2),
params1,
X=X_smo,
y=y_smo)
# +
params1 = {'n_estimators': [79]} # how do I get gridsearch to care about test score to stop overfitting?
gs_tuning(GradientBoostingClassifier(random_state=2),
params1,
X=X_smo,
y=y_smo)
# +
params2 = {'max_depth': range(2,6) ,
'min_samples_split': [.9]} #np.linspace(.05,.1,1)
gs_tuning(GradientBoostingClassifier(n_estimators=79,
random_state=2),
params2,
X=X_smo,
y=y_smo)
# +
params3 = {'min_samples_leaf': [750] ,
'max_features': ['sqrt']}
gs_tuning(GradientBoostingClassifier(n_estimators=79,
min_samples_split=0.9,
random_state=2),
params3,
X=X_smo,
y=y_smo)
# +
params3 = {'min_samples_leaf': [680] ,
'max_features': ['sqrt'],
'subsample': [1.0]}
gs_tuning(GradientBoostingClassifier(n_estimators=79,
min_samples_split=0.9,
random_state=2),
params3,
X=X_smo,
y=y_smo)
# +
params5 = {'learning_rate':[.5], 'n_estimators':[79]}
gs_tuning(GradientBoostingClassifier(max_features='sqrt',
min_samples_leaf=680,
min_samples_split=0.9,
n_estimators=79,
random_state=2),
params5,
X=X_smo,
y=y_smo)
# +
params5 = {'learning_rate':[.05], 'n_estimators':[158]}
gs_tuning(GradientBoostingClassifier(max_features='sqrt',
min_samples_leaf=680,
min_samples_split=0.9,
n_estimators=79,
random_state=2),
params5,
X=X_smo,
y=y_smo)
# +
params5 = {'learning_rate':[.025], 'n_estimators':[316]}
gs_tuning(GradientBoostingClassifier(max_features='sqrt',
min_samples_leaf=680,
min_samples_split=0.9,
n_estimators=79,
random_state=2),
params5,
X=X_smo,
y=y_smo)
# +
gb_best_smo, pred_gb_best_smo, prob_gb_best_smo = classify(GradientBoostingClassifier(max_features='sqrt',
min_samples_leaf=680,
min_samples_split=0.9,
n_estimators=79,
random_state=2),
X_smo,
y_smo)
model_cv(gb_best_smo, X_test, y_test)
# -
# #### Gradient Boosting ADA
# +
params1 = {'n_estimators': range(20,81,10)}
gs_tuning(GradientBoostingClassifier(random_state=2),
params1,
X=X_ada,
y=y_ada)
# +
params2 = {'max_depth': range(2,6) ,
'min_samples_split': np.linspace(.1,1,20)}
gs_tuning(GradientBoostingClassifier(n_estimators=80, random_state=2),
params2,
X=X_ada,
y=y_ada)
# +
pred_gb_best_thsh = find_threshold(prob_gb_best)
pred_gb_best_smo_thsh = find_threshold(prob_gb_smo)
# +
fig, (ax1, ax2) = plt.subplots(1,2, figsize=(20,5)) #,
cm1 = confusion_matrix(y_test, pred_gb, normalize='true')
cm2 = confusion_matrix(y_test, pred_gb_smo, normalize='true')
cm1_dis = ConfusionMatrixDisplay(cm1).plot(ax=ax1, cmap=plt.cm.Blues)
cm2_dis = ConfusionMatrixDisplay(cm2).plot(ax=ax2, cmap=plt.cm.Blues)
ax1.set_title('GradientBoosting (GB), no resampling')
ax2.set_title('GB w/ SMOTE')
fig, (ax4, ax5) = plt.subplots(1,2, figsize=(20,5))
cm4 = confusion_matrix(y_test, pred_gb_best, normalize='true')
cm5 = confusion_matrix(y_test, pred_gb_best_smo, normalize='true')
cm4_dis = ConfusionMatrixDisplay(cm4).plot(ax=ax4, cmap=plt.cm.Blues)
cm5_dis = ConfusionMatrixDisplay(cm5).plot(ax=ax5, cmap=plt.cm.Blues)
ax4.set_title('Optimized GradientBoosting (GB), no resampling')
ax5.set_title('Optimized GB w/ SMOTE')
fig, (ax7, ax8) = plt.subplots(1,2, figsize=(20,5))
cm7 = confusion_matrix(y_test, pred_gb_best_thsh, normalize='true')
cm8 = confusion_matrix(y_test, pred_gb_best_smo_thsh, normalize='true')
cm7_dis = ConfusionMatrixDisplay(cm7).plot(ax=ax7, cmap=plt.cm.Blues)
cm8_dis = ConfusionMatrixDisplay(cm8).plot(ax=ax8, cmap=plt.cm.Blues)
ax7.set_title('Optimized GradientBoosting w/ J stat threshold')
ax8.set_title('Optimized GB w/ SMOTE, J stat threshold')
plt.subplots_adjust(wspace=0.08)
# -
print(classification_report(y_test, pred_gb_smo),'')
print(classification_report(y_test, pred_gb_best_smo),'')
print('Regular')
print(classification_report(y_test, pred_gb_best),'')
print('SMOTE')
print(classification_report(y_test, pred_gb_best_smo),'')
#print('ADASYN')
#print(classification_report(y_test, pred_gb_best_ada),'')
print('WITH THRESHOLD')
print('Regular')
print(classification_report(y_test, pred_gb_best_thsh),'')
print('SMOTE')
print(classification_report(y_test, pred_gb_best_smo_thsh),'')
#print('ADASYN')
#print(classification_report(y_test, pred_gb_ada_thsh),'')
# +
fig = plt.figure(figsize=(7,7))
lw = 2
plt.plot([0, 1], [0, 1], color='black', lw=lw, linestyle='--')
fpr, tpr, _ = roc_curve(y_test, prob_gb_best)
auc = roc_auc_score(y_test, prob_gb_best)
plt.plot(fpr, tpr, color='darkorange', lw=lw, label='Best GB (area = %0.2f)' % auc)
fpr, tpr, _ = roc_curve(y_test, prob_gb_best_smo)
auc = roc_auc_score(y_test, prob_gb_best_smo)
plt.plot(fpr, tpr, color='darkred', lw=lw, label='Best GB /w SMOTE(area = %0.2f)' % auc)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic (ROC)')
plt.legend(loc="lower right")
plt.show()
# +
models = [clf, clf_smo, gb, gb_smo]
for m in models:
feature_importance = m.feature_importances_
# make importances relative to max importance
feature_importance = 100.0 * (feature_importance / feature_importance.max())[:30]
sorted_idx = np.argsort(feature_importance)[:30]
pos = np.arange(sorted_idx.shape[0]) + .5
print(pos.size)
sorted_idx.size
plt.figure(figsize=(10,10))
plt.barh(pos, feature_importance[sorted_idx], align='center')
plt.yticks(pos, X_train.columns[sorted_idx])
plt.xlabel('Relative Importance')
plt.title(m)
plt.show()
# -
| notebooks/Modeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import MultinomialNB, GaussianNB
from sklearn.linear_model import SGDClassifier
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.pipeline import Pipeline
from sklearn.metrics import classification_report, f1_score
import preprocessing as pp
import pandas as pd
# ### Training naive bayes model on MSR data
# +
msr_data = pd.read_csv('data/msr/raw_msr_dataset.csv', encoding = 'ANSI')
y_msr = msr_data['class']
msr = msr_data.drop(columns=['class'])
X_train, X_test, y_train, y_test = train_test_split(msr_data['token'],
y_msr, train_size=0.8,
random_state=33, shuffle=True)
# +
text_clf1 = Pipeline([
('vectorizer', CountVectorizer()),
('model', MultinomialNB())])
text_clf1.fit(X_train, y_train)
preds = text_clf1.predict(X_test)
print(classification_report(y_test, preds))
# -
# ### Evaluating on new data
new_data = pd.read_csv('data/new/raw_new_dataset.csv')
#new_data['processed_token'] = pp.preprocess_tokens(new_data)
y_new = new_data['class']
new_preds = text_clf1.predict(new_data['token'])
print(classification_report(y_new, new_preds))
print('f1', f1_score(y_new, new_preds))
# +
def show_most_informative_features(vectorizer, clf, n=50):
feature_names = vectorizer.get_feature_names()
coefs_with_fns = sorted(zip(clf.coef_[0], feature_names))
top = zip(coefs_with_fns[:n], coefs_with_fns[:-(n + 1):-1])
for (coef_1, fn_1), (coef_2, fn_2) in top:
#print("\t%.4f\t%-15s\t\t%.4f\t%-15s" % (coef_1, fn_1, coef_2, fn_2))
print(coef_2, fn_2)
show_most_informative_features(text_clf1['vectorizer'], text_clf1['model'])
# -
# The smaller the better.
# ### Testing SGDClassifier
# +
text_clf2 = Pipeline([
('vectorizer', CountVectorizer(max_features=1000)),
('model', SGDClassifier(loss='hinge', penalty='l2',
alpha=1e-3, random_state=42,
max_iter=5, tol=None))])
text_clf2.fit(X_train, y_train)
new_preds = text_clf2.predict(new_data['token'])
print(classification_report(y_new, new_preds))
print('f1', f1_score(y_new, new_preds))
# -
| nb_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from web3 import Web3, HTTPProvider, IPCProvider
from client import Client
# On web3 console:
# web3.personal.importRawKey('00a1e23ea5e858f1ae36c0248ac1286d9a8b69476dff7f8f040eddf3cb504eb1', 'panda')
web3 = Web3(HTTPProvider('http://5192.168.127.12:8545'))
assert web3.isConnected()
MY_ADDRESS = '0xB038C505907F3ba28E65c46d7fEE07077AEc55b8'
DELEGATOR_CONTRACT = '0x1698215A2bea4935bA9E0F5B48347E83450a6774'
# -
clientB = Client(1, web3, clientAddress=MY_ADDRESS, delegatorAddress=DELEGATOR_CONTRACT)
print(clientB.main())
| BEN_UserB_view.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from sklearn.preprocessing import LabelEncoder
import boto3
import json
# # Dataset : downloading - preprocessing - uploading
# First download the [dataset](http://archive.ics.uci.edu/ml/machine-learning-databases/00222/bank-additional.zip) and keep it in the data folder with name 'bankadditionalfull.csv'
raw_data = pd.read_csv('data/bankadditionalfull.csv', sep=';', index_col=0)
raw_data.head(2)
# +
# Finds categorical data from the dataframe
# Needed for creating the Data Schema, we'll see afterwards
def identify_categorical(dataframe):
total = dataframe.columns
numerical = dataframe._get_numeric_data().columns
dictionary = {'CATEGORICAL': list(set(total) - set(numerical)), 'NUMERIC':list(numerical)}
return dictionary
features = identify_categorical(raw_data)
# -
# Before starting off with using amazon services, do [this](https://boto3.amazonaws.com/v1/documentation/api/latest/guide/quickstart.html)
#
# And it is mandatory to save the data in either S3 or RedShift, otherwise you cannot use Amazon ML
# ## If data already exists in S3 Bucket
# +
# Checking if a bucket exists already
s3 = boto3.client('s3')
response = s3.list_buckets()
bucket = [buckets['Name'] for buckets in response['Buckets']]
# If you already have uploaded the data file to S3 Bucket, you would get the list of buckets in bucket variable
print(bucket)
# -
# ## If data is in your local machine and not yet uploaded
# For users who do not have a S3 Bucket created
s3 = boto3.client('s3')
s3.create_bucket(Bucket='thinkdifferentnow') # Specify any name, all the buckets should have a unique name.
s3.upload_file('data/bankadditionalfull.csv', 'thinkdifferentnow', 'bankadditionalfull.csv')
# - To know more about using boto3 to access S3 buckets click [here](https://boto3.amazonaws.com/v1/documentation/api/latest/guide/s3-example-creating-buckets.html)
#
# - Since we have saved our dataset in the S3 Bucket, we can now move forward to creating ML model.
#
# - First we need to create a datasource. A datasource is basically the information of our dataset. Like,
# * Where is it stored
# * Info of the data features (aka categorical/numerical/text/binary)
# 
client = boto3.client('machinelearning')
# #### Creating JSON file for DataSchema
# +
# Copied this from boto3 documentation
# Even you copy it as it is
DataSchema = {
"version": "1.0",
"targetFieldName": "y",
"dataFormat": "CSV",
"dataFileContainsHeader": 'true', # Set it to true because, CSV contains feature names.
}
# Now we will fill the "attributes"
attributes = []
for featureType in list(features.keys()):
for featureName in features[featureType]:
attributes.append({'fieldName':featureName, 'fieldType':featureType})
DataSchema['attributes'] = attributes
# Saving DataSchema in a JSON file
with open('data/dataschema.json', 'w') as outfile:
json.dump(DataSchema, outfile)
# +
# Make sure for Amazon ML you set your region name to 'us-east-1' or 'eu-west-1'
# As AML works only for US East(Virginia) and EU (Ireland) as of now.
# Make sure you wait for 4-5 minutes once you execute this code cell.
_ = client.create_data_source_from_s3(
DataSourceId='ds-sYkrd9KZMme', # Any ID will do
DataSourceName='tryingboto', # Any name will do
DataSpec={
'DataLocationS3': 's3://bankclassification/bankadditionalfull_.csv', # s3://bucket_name/file_name
# DataScehma is the string of the DataSchema dictionary that we created before. You can copy-paste it from dataschema.json that we created.
'DataSchema': '{"version": "1.0", "targetFieldName": "y", "dataFormat": "CSV", "dataFileContainsHeader": "true", "attributes": [{"fieldName": "day_of_week", "fieldType": "CATEGORICAL"}, {"fieldName": "y", "fieldType": "BINARY"}, {"fieldName": "contact", "fieldType": "CATEGORICAL"}, {"fieldName": "education", "fieldType": "CATEGORICAL"}, {"fieldName": "loan", "fieldType": "CATEGORICAL"}, {"fieldName": "poutcome", "fieldType": "CATEGORICAL"}, {"fieldName": "default", "fieldType": "CATEGORICAL"}, {"fieldName": "marital", "fieldType": "CATEGORICAL"}, {"fieldName": "job", "fieldType": "CATEGORICAL"}, {"fieldName": "month", "fieldType": "CATEGORICAL"}, {"fieldName": "housing", "fieldType": "CATEGORICAL"}, {"fieldName": "duration", "fieldType": "NUMERIC"}, {"fieldName": "campaign", "fieldType": "NUMERIC"}, {"fieldName": "pdays", "fieldType": "NUMERIC"}, {"fieldName": "previous", "fieldType": "NUMERIC"}, {"fieldName": "emp.var.rate", "fieldType": "NUMERIC"}, {"fieldName": "cons.price.idx", "fieldType": "NUMERIC"}, {"fieldName": "cons.conf.idx", "fieldType": "NUMERIC"}, {"fieldName": "euribor3m", "fieldType": "NUMERIC"}, {"fieldName": "nr.employed", "fieldType": "NUMERIC"}]}'
},
ComputeStatistics=True
)
# It turns out, surprisingly it took 16 mins of compute time for creating the datasource :(
# -
# Once Data Source is created, you'd get this:
#
# 
__ = client.create_ml_model(
MLModelId='mlmodelid_',
MLModelName='marketingbank',
MLModelType='BINARY', # Amazon ML has 3 types of model types: BINARY | MULTICLASS | REGRESSION
TrainingDataSourceId='ds-sYkrd9KZMme'
)
# Once the model is trained, you would get this in your Dashboard
#
# 
# +
# To create batch predictions on the test data
# We again need to create a datasource for the dataset
# And enter the datasource id below
___ = client.create_batch_prediction(
BatchPredictionId='batchpredictionid_',
BatchPredictionName='predictresults',
MLModelId='mlmodelid_',
BatchPredictionDataSourceId='ds-CmsaR7xPeTU',
OutputUri='s3://bankclassification/'
)
# OutputUri specifies in which S3 bucket directory shall the prediction folder be placed.
# -
# After executing above code cell, if you go and check your Dashboard you would see something like this:
#
# 
#
#
#
# 
#
#
# And you can find your predictions folder with a name of batch-prediction in the give S3 bucket
| bank_try.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,md:myst
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ```{warning}
# This book is a work in progress and should be considered currently to be in a
# **pre**draft state. Work is actively taking place in preparation for October
# 2020.
#
# If you happen to find this and notice any typos and/or have any suggestions
# please open an issue on the github repo: <https://github.com/drvinceknight/pfm>
# ```
#
# # Python for Mathematics
#
# ## Introduction
#
# This book aims to introduce readers to programming for mathematics.
#
# It is assumed that readers are used to solving high school mathematics problems
# of the form:
#
# ---
#
# ```{admonition} Problem
# Given the function $f:\mathbb{R}\to\mathbb{R}$ defined by
# $f(x) = x ^ 2 - 3 x + 1$ obtain the global minima of the function.
# ```
#
# ```{admonition} Solution
# :class: tip
#
# To solve this we need to apply our **mathematical knowledge** which tells us to:
#
# 1. Differentiate $f(x)$ to get $\frac{df}{dx}$;
# 2. Equate $\frac{df}{dx}=0$;
# 3. Use the second derivative test on the solution to the previous equation.
#
# For each of those 3 steps we will usually make use of our **mathematical
# techniques**:
#
# 1. Differentiate $f(x)$:
#
# $$\frac{df}{dx} = 2 x - 3$$
#
# 2. Equate $\frac{df}{dx}=0$:
#
# $$2x-3 =0 \Rightarrow x = 3/2$$
#
# 3. Use the second derivative test on the solution:
#
# $$\frac{d^2f}{dx^2} = 2 > 0\text{ for all values of }x$$
#
# Thus $x=2/3$ is the global minima of the function.
# ```
#
# ```{attention}
# As we progress as mathematicians **mathematical knowledge** is more prominent
# than **mathematical technique**: often knowing what to do is the real problem as
# opposed to having the technical ability to do it.
# ```
#
# This is what this book will cover: **programming** allows us to instruct a
# computer to carry out mathematical techniques.
#
# We will for example learn how to solve the above problem by instructing a
# computer which **mathematical technique** to carry out.
#
# **This book will teach us how to give the correct instructions to a
# computer.**
#
# The following is an example, do not worry too much about the specific code used
# for now:
#
# 1. Differentiate $f(x)$ to get $\frac{df}{dx}$;
# +
import sympy as sym
x = sym.Symbol("x")
sym.diff(x ** 2 - 3 * x + 1, x)
# -
# 2. Equate $\frac{df}{dx}=0$:
sym.solveset(2 * x - 3, x)
# 3. Use the second derivative test on the solution:
sym.diff(x ** 2 - 3 * x + 1, x, 2)
# {ref}`Knowledge versus technique <fig:knowledge_vs_technique>` is a brief summary.
#
# ```{figure} ./img/knowledge_vs_technique/main.png
# ---
# width: 50%
# name: fig:knowledge_vs_technique
# ---
# Knowledge versus technique in this book.
# ```
#
# ## How this book is structured
#
# Most programming texts introduce readers to the building blocks of
# programming and build up to using more sophisticated tools for a specific
# purpose.
#
# This is akin to teaching someone how to forge metal so as to make a nail and
# then slowly work our way to using more sophisticated tools such as power tools
# to build a house.
#
# This book will do thing in a different way: we will start with using and
# understanding tools that are helpful to mathematicians. In the later part of the
# book we will cover the building blocks and you will be able to build your own
# sophisticated tools.
#
# The book is in two parts:
#
# 1. Tools for mathematics;
# 2. Building tools.
#
# The first part of the book will not make use of any novel mathematics.
# Instead we will consider a number of mathematics problem that are often covered
# in secondary school.
#
# - Algebraic manipulation
# - Calculus (differentiation and integration)
# - Permutations and combinations
# - Probability
# - Linear algebra
#
# The questions we will tackle will be familiar in their presentation and
# description. **What will be different** is that no **by hand** calculations will
# be done. We will instead carry them all out using a programming language.
#
# In the second part of the book you will be encouraged to build your own tools
# to be able to tackle a problem type of your choice.
#
# ```{attention}
# Every chapter will have 4 parts:
#
# - A tutorial: you will be walked through solving a problem. You will be
# specifically told what to do and what to expect.
# - A how to section: this will be a shorter more succinct section that will
# detail how to carry out specific things.
# - A reference section: this will be a section with references to further
# resources as well as background information about specific things in the
# chapter.
# - An exercise section: this will be a number of exercises that you can work on.
# ```
| book/.intro.md.bcp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="copyright"
# Copyright 2020 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="title"
# # Vertex client library: AutoML image classification model for export to edge
#
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/GoogleCloudPlatform/ai-platform-samples/blob/master/ai-platform-unified/notebooks/community/gapic/automl/showcase_automl_image_classification_export_edge.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
# </a>
# </td>
# <td>
# <a href="https://github.com/GoogleCloudPlatform/ai-platform-samples/blob/master/ai-platform-unified/notebooks/community/gapic/automl/showcase_automl_image_classification_export_edge.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
# View on GitHub
# </a>
# </td>
# </table>
# <br/><br/><br/>
# + [markdown] id="overview:automl,export_edge"
# ## Overview
#
#
# This tutorial demonstrates how to use the Vertex client library for Python to create image classification models to export as an Edge model using Google Cloud's AutoML.
# + [markdown] id="dataset:flowers,icn"
# ### Dataset
#
# The dataset used for this tutorial is the [Flowers dataset](https://www.tensorflow.org/datasets/catalog/tf_flowers) from [TensorFlow Datasets](https://www.tensorflow.org/datasets/catalog/overview). The version of the dataset you will use in this tutorial is stored in a public Cloud Storage bucket. The trained model predicts the type of flower an image is from a class of five flowers: daisy, dandelion, rose, sunflower, or tulip.
# + [markdown] id="objective:automl,training,export_edge"
# ### Objective
#
# In this tutorial, you create a AutoML image classification model from a Python script using the Vertex client library, and then export the model as an Edge model in TFLite format. You can alternatively create models with AutoML using the `gcloud` command-line tool or online using the Google Cloud Console.
#
# The steps performed include:
#
# - Create a Vertex `Dataset` resource.
# - Train the model.
# - Export the `Edge` model from the `Model` resource to Cloud Storage.
# - Download the model locally.
# - Make a local prediction.
# + [markdown] id="costs"
# ### Costs
#
# This tutorial uses billable components of Google Cloud (GCP):
#
# * Vertex AI
# * Cloud Storage
#
# Learn about [Vertex AI
# pricing](https://cloud.google.com/vertex-ai/pricing) and [Cloud Storage
# pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
# Calculator](https://cloud.google.com/products/calculator/)
# to generate a cost estimate based on your projected usage.
# + [markdown] id="install_aip"
# ## Installation
#
# Install the latest version of Vertex client library.
# + id="install_aip"
import os
import sys
# Google Cloud Notebook
if os.path.exists("/opt/deeplearning/metadata/env_version"):
USER_FLAG = "--user"
else:
USER_FLAG = ""
# ! pip3 install -U google-cloud-aiplatform $USER_FLAG
# + [markdown] id="install_storage"
# Install the latest GA version of *google-cloud-storage* library as well.
# + id="install_storage"
# ! pip3 install -U google-cloud-storage $USER_FLAG
# + [markdown] id="restart"
# ### Restart the kernel
#
# Once you've installed the Vertex client library and Google *cloud-storage*, you need to restart the notebook kernel so it can find the packages.
# + id="restart"
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
# + [markdown] id="before_you_begin"
# ## Before you begin
#
# ### GPU runtime
#
# *Make sure you're running this notebook in a GPU runtime if you have that option. In Colab, select* **Runtime > Change Runtime Type > GPU**
#
# ### Set up your Google Cloud project
#
# **The following steps are required, regardless of your notebook environment.**
#
# 1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
#
# 2. [Make sure that billing is enabled for your project.](https://cloud.google.com/billing/docs/how-to/modify-project)
#
# 3. [Enable the Vertex APIs and Compute Engine APIs.](https://console.cloud.google.com/flows/enableapi?apiid=ml.googleapis.com,compute_component)
#
# 4. [The Google Cloud SDK](https://cloud.google.com/sdk) is already installed in Google Cloud Notebook.
#
# 5. Enter your project ID in the cell below. Then run the cell to make sure the
# Cloud SDK uses the right project for all the commands in this notebook.
#
# **Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
# + id="set_project_id"
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
# + id="autoset_project_id"
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
# shell_output = !gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
# + id="set_gcloud_project_id"
# ! gcloud config set project $PROJECT_ID
# + [markdown] id="region"
# #### Region
#
# You can also change the `REGION` variable, which is used for operations
# throughout the rest of this notebook. Below are regions supported for Vertex. We recommend that you choose the region closest to you.
#
# - Americas: `us-central1`
# - Europe: `europe-west4`
# - Asia Pacific: `asia-east1`
#
# You may not use a multi-regional bucket for training with Vertex. Not all regions provide support for all Vertex services. For the latest support per region, see the [Vertex locations documentation](https://cloud.google.com/vertex-ai/docs/general/locations)
# + id="region"
REGION = "us-central1" # @param {type: "string"}
# + [markdown] id="timestamp"
# #### Timestamp
#
# If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append onto the name of resources which will be created in this tutorial.
# + id="timestamp"
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
# + [markdown] id="gcp_authenticate"
# ### Authenticate your Google Cloud account
#
# **If you are using Google Cloud Notebook**, your environment is already authenticated. Skip this step.
#
# **If you are using Colab**, run the cell below and follow the instructions when prompted to authenticate your account via oAuth.
#
# **Otherwise**, follow these steps:
#
# In the Cloud Console, go to the [Create service account key](https://console.cloud.google.com/apis/credentials/serviceaccountkey) page.
#
# **Click Create service account**.
#
# In the **Service account name** field, enter a name, and click **Create**.
#
# In the **Grant this service account access to project** section, click the Role drop-down list. Type "Vertex" into the filter box, and select **Vertex Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
#
# Click Create. A JSON file that contains your key downloads to your local environment.
#
# Enter the path to your service account key as the GOOGLE_APPLICATION_CREDENTIALS variable in the cell below and run the cell.
# + id="gcp_authenticate"
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# If on Google Cloud Notebook, then don't execute this code
if not os.path.exists("/opt/deeplearning/metadata/env_version"):
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
# %env GOOGLE_APPLICATION_CREDENTIALS ''
# + [markdown] id="bucket:export_edge"
# ### Create a Cloud Storage bucket
#
# **The following steps are required, regardless of your notebook environment.**
#
# This tutorial is designed to use training data that is in a public Cloud Storage bucket and a local Cloud Storage bucket for exporting the trained model. You may alternatively use your own training data that you have stored in a local Cloud Storage bucket.
#
# Set the name of your Cloud Storage bucket below. Bucket names must be globally unique across all Google Cloud projects, including those outside of your organization.
# + id="bucket"
BUCKET_NAME = "gs://[your-bucket-name]" # @param {type:"string"}
# + id="autoset_bucket"
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "gs://[your-bucket-name]":
BUCKET_NAME = "gs://" + PROJECT_ID + "aip-" + TIMESTAMP
# + [markdown] id="create_bucket"
# **Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
# + id="create_bucket"
# ! gsutil mb -l $REGION $BUCKET_NAME
# + [markdown] id="validate_bucket"
# Finally, validate access to your Cloud Storage bucket by examining its contents:
# + id="validate_bucket"
# ! gsutil ls -al $BUCKET_NAME
# + [markdown] id="setup_vars"
# ### Set up variables
#
# Next, set up some variables used throughout the tutorial.
# ### Import libraries and define constants
# + [markdown] id="import_aip:protobuf"
# #### Import Vertex client library
#
# Import the Vertex client library into our Python environment.
# + id="import_aip:protobuf"
import time
from google.cloud.aiplatform import gapic as aip
from google.protobuf import json_format
from google.protobuf.json_format import MessageToJson, ParseDict
from google.protobuf.struct_pb2 import Struct, Value
# + [markdown] id="aip_constants"
# #### Vertex constants
#
# Setup up the following constants for Vertex:
#
# - `API_ENDPOINT`: The Vertex API service endpoint for dataset, model, job, pipeline and endpoint services.
# - `PARENT`: The Vertex location root path for dataset, model, job, pipeline and endpoint resources.
# + id="aip_constants"
# API service endpoint
API_ENDPOINT = "{}-aiplatform.googleapis.com".format(REGION)
# Vertex location root path for your dataset, model and endpoint resources
PARENT = "projects/" + PROJECT_ID + "/locations/" + REGION
# + [markdown] id="automl_constants"
# #### AutoML constants
#
# Set constants unique to AutoML datasets and training:
#
# - Dataset Schemas: Tells the `Dataset` resource service which type of dataset it is.
# - Data Labeling (Annotations) Schemas: Tells the `Dataset` resource service how the data is labeled (annotated).
# - Dataset Training Schemas: Tells the `Pipeline` resource service the task (e.g., classification) to train the model for.
# + id="automl_constants:icn"
# Image Dataset type
DATA_SCHEMA = "gs://google-cloud-aiplatform/schema/dataset/metadata/image_1.0.0.yaml"
# Image Labeling type
LABEL_SCHEMA = "gs://google-cloud-aiplatform/schema/dataset/ioformat/image_classification_single_label_io_format_1.0.0.yaml"
# Image Training task
TRAINING_SCHEMA = "gs://google-cloud-aiplatform/schema/trainingjob/definition/automl_image_classification_1.0.0.yaml"
# + [markdown] id="tutorial_start:automl"
# # Tutorial
#
# Now you are ready to start creating your own AutoML image classification model.
# + [markdown] id="clients:automl,export_edge"
# ## Set up clients
#
# The Vertex client library works as a client/server model. On your side (the Python script) you will create a client that sends requests and receives responses from the Vertex server.
#
# You will use different clients in this tutorial for different steps in the workflow. So set them all up upfront.
#
# - Dataset Service for `Dataset` resources.
# - Model Service for `Model` resources.
# - Pipeline Service for training.
# + id="clients:automl,export_edge"
# client options same for all services
client_options = {"api_endpoint": API_ENDPOINT}
def create_dataset_client():
client = aip.DatasetServiceClient(client_options=client_options)
return client
def create_model_client():
client = aip.ModelServiceClient(client_options=client_options)
return client
def create_pipeline_client():
client = aip.PipelineServiceClient(client_options=client_options)
return client
clients = {}
clients["dataset"] = create_dataset_client()
clients["model"] = create_model_client()
clients["pipeline"] = create_pipeline_client()
for client in clients.items():
print(client)
# + [markdown] id="create_aip_dataset"
# ## Dataset
#
# Now that your clients are ready, your first step in training a model is to create a managed dataset instance, and then upload your labeled data to it.
#
# ### Create `Dataset` resource instance
#
# Use the helper function `create_dataset` to create the instance of a `Dataset` resource. This function does the following:
#
# 1. Uses the dataset client service.
# 2. Creates an Vertex `Dataset` resource (`aip.Dataset`), with the following parameters:
# - `display_name`: The human-readable name you choose to give it.
# - `metadata_schema_uri`: The schema for the dataset type.
# 3. Calls the client dataset service method `create_dataset`, with the following parameters:
# - `parent`: The Vertex location root path for your `Database`, `Model` and `Endpoint` resources.
# - `dataset`: The Vertex dataset object instance you created.
# 4. The method returns an `operation` object.
#
# An `operation` object is how Vertex handles asynchronous calls for long running operations. While this step usually goes fast, when you first use it in your project, there is a longer delay due to provisioning.
#
# You can use the `operation` object to get status on the operation (e.g., create `Dataset` resource) or to cancel the operation, by invoking an operation method:
#
# | Method | Description |
# | ----------- | ----------- |
# | result() | Waits for the operation to complete and returns a result object in JSON format. |
# | running() | Returns True/False on whether the operation is still running. |
# | done() | Returns True/False on whether the operation is completed. |
# | canceled() | Returns True/False on whether the operation was canceled. |
# | cancel() | Cancels the operation (this may take up to 30 seconds). |
# + id="create_aip_dataset"
TIMEOUT = 90
def create_dataset(name, schema, labels=None, timeout=TIMEOUT):
start_time = time.time()
try:
dataset = aip.Dataset(
display_name=name, metadata_schema_uri=schema, labels=labels
)
operation = clients["dataset"].create_dataset(parent=PARENT, dataset=dataset)
print("Long running operation:", operation.operation.name)
result = operation.result(timeout=TIMEOUT)
print("time:", time.time() - start_time)
print("response")
print(" name:", result.name)
print(" display_name:", result.display_name)
print(" metadata_schema_uri:", result.metadata_schema_uri)
print(" metadata:", dict(result.metadata))
print(" create_time:", result.create_time)
print(" update_time:", result.update_time)
print(" etag:", result.etag)
print(" labels:", dict(result.labels))
return result
except Exception as e:
print("exception:", e)
return None
result = create_dataset("flowers-" + TIMESTAMP, DATA_SCHEMA)
# + [markdown] id="dataset_id:result"
# Now save the unique dataset identifier for the `Dataset` resource instance you created.
# + id="dataset_id:result"
# The full unique ID for the dataset
dataset_id = result.name
# The short numeric ID for the dataset
dataset_short_id = dataset_id.split("/")[-1]
print(dataset_id)
# + [markdown] id="data_preparation:image,u_dataset"
# ### Data preparation
#
# The Vertex `Dataset` resource for images has some requirements for your data:
#
# - Images must be stored in a Cloud Storage bucket.
# - Each image file must be in an image format (PNG, JPEG, BMP, ...).
# - There must be an index file stored in your Cloud Storage bucket that contains the path and label for each image.
# - The index file must be either CSV or JSONL.
# + [markdown] id="data_import_format:icn,u_dataset,csv"
# #### CSV
#
# For image classification, the CSV index file has the requirements:
#
# - No heading.
# - First column is the Cloud Storage path to the image.
# - Second column is the label.
# + [markdown] id="import_file:u_dataset,csv"
# #### Location of Cloud Storage training data.
#
# Now set the variable `IMPORT_FILE` to the location of the CSV index file in Cloud Storage.
# + id="import_file:flowers,csv,icn"
IMPORT_FILE = (
"gs://cloud-samples-data/vision/automl_classification/flowers/all_data_v2.csv"
)
# + [markdown] id="quick_peek:csv"
# #### Quick peek at your data
#
# You will use a version of the Flowers dataset that is stored in a public Cloud Storage bucket, using a CSV index file.
#
# Start by doing a quick peek at the data. You count the number of examples by counting the number of rows in the CSV index file (`wc -l`) and then peek at the first few rows.
# + id="quick_peek:csv"
if "IMPORT_FILES" in globals():
FILE = IMPORT_FILES[0]
else:
FILE = IMPORT_FILE
count = ! gsutil cat $FILE | wc -l
print("Number of Examples", int(count[0]))
print("First 10 rows")
# ! gsutil cat $FILE | head
# + [markdown] id="import_data"
# ### Import data
#
# Now, import the data into your Vertex Dataset resource. Use this helper function `import_data` to import the data. The function does the following:
#
# - Uses the `Dataset` client.
# - Calls the client method `import_data`, with the following parameters:
# - `name`: The human readable name you give to the `Dataset` resource (e.g., flowers).
# - `import_configs`: The import configuration.
#
# - `import_configs`: A Python list containing a dictionary, with the key/value entries:
# - `gcs_sources`: A list of URIs to the paths of the one or more index files.
# - `import_schema_uri`: The schema identifying the labeling type.
#
# The `import_data()` method returns a long running `operation` object. This will take a few minutes to complete. If you are in a live tutorial, this would be a good time to ask questions, or take a personal break.
# + id="import_data"
def import_data(dataset, gcs_sources, schema):
config = [{"gcs_source": {"uris": gcs_sources}, "import_schema_uri": schema}]
print("dataset:", dataset_id)
start_time = time.time()
try:
operation = clients["dataset"].import_data(
name=dataset_id, import_configs=config
)
print("Long running operation:", operation.operation.name)
result = operation.result()
print("result:", result)
print("time:", int(time.time() - start_time), "secs")
print("error:", operation.exception())
print("meta :", operation.metadata)
print(
"after: running:",
operation.running(),
"done:",
operation.done(),
"cancelled:",
operation.cancelled(),
)
return operation
except Exception as e:
print("exception:", e)
return None
import_data(dataset_id, [IMPORT_FILE], LABEL_SCHEMA)
# + [markdown] id="train_automl_model"
# ## Train the model
#
# Now train an AutoML image classification model using your Vertex `Dataset` resource. To train the model, do the following steps:
#
# 1. Create an Vertex training pipeline for the `Dataset` resource.
# 2. Execute the pipeline to start the training.
# + [markdown] id="create_pipeline:automl"
# ### Create a training pipeline
#
# You may ask, what do we use a pipeline for? You typically use pipelines when the job (such as training) has multiple steps, generally in sequential order: do step A, do step B, etc. By putting the steps into a pipeline, we gain the benefits of:
#
# 1. Being reusable for subsequent training jobs.
# 2. Can be containerized and ran as a batch job.
# 3. Can be distributed.
# 4. All the steps are associated with the same pipeline job for tracking progress.
#
# Use this helper function `create_pipeline`, which takes the following parameters:
#
# - `pipeline_name`: A human readable name for the pipeline job.
# - `model_name`: A human readable name for the model.
# - `dataset`: The Vertex fully qualified dataset identifier.
# - `schema`: The dataset labeling (annotation) training schema.
# - `task`: A dictionary describing the requirements for the training job.
#
# The helper function calls the `Pipeline` client service'smethod `create_pipeline`, which takes the following parameters:
#
# - `parent`: The Vertex location root path for your `Dataset`, `Model` and `Endpoint` resources.
# - `training_pipeline`: the full specification for the pipeline training job.
#
# Let's look now deeper into the *minimal* requirements for constructing a `training_pipeline` specification:
#
# - `display_name`: A human readable name for the pipeline job.
# - `training_task_definition`: The dataset labeling (annotation) training schema.
# - `training_task_inputs`: A dictionary describing the requirements for the training job.
# - `model_to_upload`: A human readable name for the model.
# - `input_data_config`: The dataset specification.
# - `dataset_id`: The Vertex dataset identifier only (non-fully qualified) -- this is the last part of the fully-qualified identifier.
# - `fraction_split`: If specified, the percentages of the dataset to use for training, test and validation. Otherwise, the percentages are automatically selected by AutoML.
# + id="create_pipeline:automl"
def create_pipeline(pipeline_name, model_name, dataset, schema, task):
dataset_id = dataset.split("/")[-1]
input_config = {
"dataset_id": dataset_id,
"fraction_split": {
"training_fraction": 0.8,
"validation_fraction": 0.1,
"test_fraction": 0.1,
},
}
training_pipeline = {
"display_name": pipeline_name,
"training_task_definition": schema,
"training_task_inputs": task,
"input_data_config": input_config,
"model_to_upload": {"display_name": model_name},
}
try:
pipeline = clients["pipeline"].create_training_pipeline(
parent=PARENT, training_pipeline=training_pipeline
)
print(pipeline)
except Exception as e:
print("exception:", e)
return None
return pipeline
# + [markdown] id="task_requirements:automl,icn,edge"
# ### Construct the task requirements
#
# Next, construct the task requirements. Unlike other parameters which take a Python (JSON-like) dictionary, the `task` field takes a Google protobuf Struct, which is very similar to a Python dictionary. Use the `json_format.ParseDict` method for the conversion.
#
# The minimal fields we need to specify are:
#
# - `multi_label`: Whether True/False this is a multi-label (vs single) classification.
# - `budget_milli_node_hours`: The maximum time to budget (billed) for training the model, where 1000 = 1 hour. For image classification, the budget must be a minimum of 8 hours.
# - `model_type`: The type of deployed model:
# - `CLOUD`: For deploying to Google Cloud.
# - `MOBILE_TF_LOW_LATENCY_1`: For deploying to the edge and optimizing for latency (response time).
# - `MOBILE_TF_HIGH_ACCURACY_1`: For deploying to the edge and optimizing for accuracy.
# - `MOBILE_TF_VERSATILE_1`: For deploying to the edge and optimizing for a trade off between latency and accuracy.
# - `disable_early_stopping`: Whether True/False to let AutoML use its judgement to stop training early or train for the entire budget.
#
# Finally, create the pipeline by calling the helper function `create_pipeline`, which returns an instance of a training pipeline object.
# + id="task_requirements:automl,icn,edge"
PIPE_NAME = "flowers_pipe-" + TIMESTAMP
MODEL_NAME = "flowers_model-" + TIMESTAMP
task = json_format.ParseDict(
{
"multi_label": False,
"budget_milli_node_hours": 8000,
"model_type": "MOBILE_TF_LOW_LATENCY_1",
"disable_early_stopping": False,
},
Value(),
)
response = create_pipeline(PIPE_NAME, MODEL_NAME, dataset_id, TRAINING_SCHEMA, task)
# + [markdown] id="pipeline_id:response"
# Now save the unique identifier of the training pipeline you created.
# + id="pipeline_id:response"
# The full unique ID for the pipeline
pipeline_id = response.name
# The short numeric ID for the pipeline
pipeline_short_id = pipeline_id.split("/")[-1]
print(pipeline_id)
# + [markdown] id="get_training_pipeline"
# ### Get information on a training pipeline
#
# Now get pipeline information for just this training pipeline instance. The helper function gets the job information for just this job by calling the the job client service's `get_training_pipeline` method, with the following parameter:
#
# - `name`: The Vertex fully qualified pipeline identifier.
#
# When the model is done training, the pipeline state will be `PIPELINE_STATE_SUCCEEDED`.
# + id="get_training_pipeline"
def get_training_pipeline(name, silent=False):
response = clients["pipeline"].get_training_pipeline(name=name)
if silent:
return response
print("pipeline")
print(" name:", response.name)
print(" display_name:", response.display_name)
print(" state:", response.state)
print(" training_task_definition:", response.training_task_definition)
print(" training_task_inputs:", dict(response.training_task_inputs))
print(" create_time:", response.create_time)
print(" start_time:", response.start_time)
print(" end_time:", response.end_time)
print(" update_time:", response.update_time)
print(" labels:", dict(response.labels))
return response
response = get_training_pipeline(pipeline_id)
# + [markdown] id="wait_training_complete"
# # Deployment
#
# Training the above model may take upwards of 30 minutes time.
#
# Once your model is done training, you can calculate the actual time it took to train the model by subtracting `end_time` from `start_time`. For your model, you will need to know the fully qualified Vertex Model resource identifier, which the pipeline service assigned to it. You can get this from the returned pipeline instance as the field `model_to_deploy.name`.
# + id="wait_training_complete"
while True:
response = get_training_pipeline(pipeline_id, True)
if response.state != aip.PipelineState.PIPELINE_STATE_SUCCEEDED:
print("Training job has not completed:", response.state)
model_to_deploy_id = None
if response.state == aip.PipelineState.PIPELINE_STATE_FAILED:
raise Exception("Training Job Failed")
else:
model_to_deploy = response.model_to_upload
model_to_deploy_id = model_to_deploy.name
print("Training Time:", response.end_time - response.start_time)
break
time.sleep(60)
print("model to deploy:", model_to_deploy_id)
# + [markdown] id="model_information"
# ## Model information
#
# Now that your model is trained, you can get some information on your model.
# + [markdown] id="evaluate_the_model:automl"
# ## Evaluate the Model resource
#
# Now find out how good the model service believes your model is. As part of training, some portion of the dataset was set aside as the test (holdout) data, which is used by the pipeline service to evaluate the model.
# + [markdown] id="list_model_evaluations:automl,icn"
# ### List evaluations for all slices
#
# Use this helper function `list_model_evaluations`, which takes the following parameter:
#
# - `name`: The Vertex fully qualified model identifier for the `Model` resource.
#
# This helper function uses the model client service's `list_model_evaluations` method, which takes the same parameter. The response object from the call is a list, where each element is an evaluation metric.
#
# For each evaluation (you probably only have one) we then print all the key names for each metric in the evaluation, and for a small set (`logLoss` and `auPrc`) you will print the result.
# + id="list_model_evaluations:automl,icn"
def list_model_evaluations(name):
response = clients["model"].list_model_evaluations(parent=name)
for evaluation in response:
print("model_evaluation")
print(" name:", evaluation.name)
print(" metrics_schema_uri:", evaluation.metrics_schema_uri)
metrics = json_format.MessageToDict(evaluation._pb.metrics)
for metric in metrics.keys():
print(metric)
print("logloss", metrics["logLoss"])
print("auPrc", metrics["auPrc"])
return evaluation.name
last_evaluation = list_model_evaluations(model_to_deploy_id)
# + [markdown] id="export_model:export_edge"
# ## Export as Edge model
#
# You can export an AutoML image classification model as an Edge model which you can then custom deploy to an edge device, such as a mobile phone or IoT device, or download locally. Use this helper function `export_model` to export the model to Google Cloud, which takes the following parameters:
#
# - `name`: The Vertex fully qualified identifier for the `Model` resource.
# - `format`: The format to save the model format as.
# - `gcs_dest`: The Cloud Storage location to store the SavedFormat model artifacts to.
#
# This function calls the `Model` client service's method `export_model`, with the following parameters:
#
# - `name`: The Vertex fully qualified identifier for the `Model` resource.
# - `output_config`: The destination information for the exported model.
# - `artifact_destination.output_uri_prefix`: The Cloud Storage location to store the SavedFormat model artifacts to.
# - `export_format_id`: The format to save the model format as. For AutoML image classification:
# - `tf-saved-model`: TensorFlow SavedFormat for deployment to a container.
# - `tflite`: TensorFlow Lite for deployment to an edge or mobile device.
# - `edgetpu-tflite`: TensorFlow Lite for TPU
# - `tf-js`: TensorFlow for web client
# - `coral-ml`: for Coral devices
#
# The method returns a long running operation `response`. We will wait sychronously for the operation to complete by calling the `response.result()`, which will block until the model is exported.
# + id="export_model:export_edge"
MODEL_DIR = BUCKET_NAME + "/" + "flowers"
def export_model(name, format, gcs_dest):
output_config = {
"artifact_destination": {"output_uri_prefix": gcs_dest},
"export_format_id": format,
}
response = clients["model"].export_model(name=name, output_config=output_config)
print("Long running operation:", response.operation.name)
result = response.result(timeout=1800)
metadata = response.operation.metadata
artifact_uri = str(metadata.value).split("\\")[-1][4:-1]
print("Artifact Uri", artifact_uri)
return artifact_uri
model_package = export_model(model_to_deploy_id, "tflite", MODEL_DIR)
# + [markdown] id="download_model_artifacts:tflite"
# #### Download the TFLite model artifacts
#
# Now that you have an exported TFLite version of your model, you can test the exported model locally, but first downloading it from Cloud Storage.
# + id="download_model_artifacts:tflite"
# ! gsutil ls $model_package
# Download the model artifacts
# ! gsutil cp -r $model_package tflite
tflite_path = "tflite/model.tflite"
# + [markdown] id="instantiate_tflite_interpreter"
# #### Instantiate a TFLite interpreter
#
# The TFLite version of the model is not a TensorFlow SavedModel format. You cannot directly use methods like predict(). Instead, one uses the TFLite interpreter. You must first setup the interpreter for the TFLite model as follows:
#
# - Instantiate an TFLite interpreter for the TFLite model.
# - Instruct the interpreter to allocate input and output tensors for the model.
# - Get detail information about the models input and output tensors that will need to be known for prediction.
# + id="instantiate_tflite_interpreter"
import tensorflow as tf
interpreter = tf.lite.Interpreter(model_path=tflite_path)
interpreter.allocate_tensors()
input_details = interpreter.get_input_details()
output_details = interpreter.get_output_details()
input_shape = input_details[0]["shape"]
print("input tensor shape", input_shape)
# + [markdown] id="get_test_item"
# ### Get test item
#
# You will use an arbitrary example out of the dataset as a test item. Don't be concerned that the example was likely used in training the model -- we just want to demonstrate how to make a prediction.
# + id="get_test_item:image,224x224"
test_items = ! gsutil cat $IMPORT_FILE | head -n1
test_item = test_items[0].split(",")[0]
with tf.io.gfile.GFile(test_item, "rb") as f:
content = f.read()
test_image = tf.io.decode_jpeg(content)
print("test image shape", test_image.shape)
test_image = tf.image.resize(test_image, (224, 224))
print("test image shape", test_image.shape, test_image.dtype)
test_image = tf.cast(test_image, dtype=tf.uint8).numpy()
# + [markdown] id="invoke_tflite_interpreter"
# #### Make a prediction with TFLite model
#
# Finally, you do a prediction using your TFLite model, as follows:
#
# - Convert the test image into a batch of a single image (`np.expand_dims`)
# - Set the input tensor for the interpreter to your batch of a single image (`data`).
# - Invoke the interpreter.
# - Retrieve the softmax probabilities for the prediction (`get_tensor`).
# - Determine which label had the highest probability (`np.argmax`).
# + id="invoke_tflite_interpreter"
import numpy as np
data = np.expand_dims(test_image, axis=0)
interpreter.set_tensor(input_details[0]["index"], data)
interpreter.invoke()
softmax = interpreter.get_tensor(output_details[0]["index"])
label = np.argmax(softmax)
print(label)
# + [markdown] id="cleanup"
# # Cleaning up
#
# To clean up all GCP resources used in this project, you can [delete the GCP
# project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
#
# Otherwise, you can delete the individual resources you created in this tutorial:
#
# - Dataset
# - Pipeline
# - Model
# - Endpoint
# - Batch Job
# - Custom Job
# - Hyperparameter Tuning Job
# - Cloud Storage Bucket
# + id="cleanup"
delete_dataset = True
delete_pipeline = True
delete_model = True
delete_endpoint = True
delete_batchjob = True
delete_customjob = True
delete_hptjob = True
delete_bucket = True
# Delete the dataset using the Vertex fully qualified identifier for the dataset
try:
if delete_dataset and "dataset_id" in globals():
clients["dataset"].delete_dataset(name=dataset_id)
except Exception as e:
print(e)
# Delete the training pipeline using the Vertex fully qualified identifier for the pipeline
try:
if delete_pipeline and "pipeline_id" in globals():
clients["pipeline"].delete_training_pipeline(name=pipeline_id)
except Exception as e:
print(e)
# Delete the model using the Vertex fully qualified identifier for the model
try:
if delete_model and "model_to_deploy_id" in globals():
clients["model"].delete_model(name=model_to_deploy_id)
except Exception as e:
print(e)
# Delete the endpoint using the Vertex fully qualified identifier for the endpoint
try:
if delete_endpoint and "endpoint_id" in globals():
clients["endpoint"].delete_endpoint(name=endpoint_id)
except Exception as e:
print(e)
# Delete the batch job using the Vertex fully qualified identifier for the batch job
try:
if delete_batchjob and "batch_job_id" in globals():
clients["job"].delete_batch_prediction_job(name=batch_job_id)
except Exception as e:
print(e)
# Delete the custom job using the Vertex fully qualified identifier for the custom job
try:
if delete_customjob and "job_id" in globals():
clients["job"].delete_custom_job(name=job_id)
except Exception as e:
print(e)
# Delete the hyperparameter tuning job using the Vertex fully qualified identifier for the hyperparameter tuning job
try:
if delete_hptjob and "hpt_job_id" in globals():
clients["job"].delete_hyperparameter_tuning_job(name=hpt_job_id)
except Exception as e:
print(e)
if delete_bucket and "BUCKET_NAME" in globals():
# ! gsutil rm -r $BUCKET_NAME
| ai-platform-unified/notebooks/unofficial/gapic/automl/showcase_automl_image_classification_export_edge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import os
import json
from shutil import copyfile, rmtree
from pprint import pprint
inputData = []
with open('data/etla_images.json') as json_data:
images = json.load(json_data)
json_data.close()
inputDatum = []
for i in range(len(images)):
if len(inputDatum) == 10:
inputData.append(inputDatum)
inputDatum = []
inputDatum.append(images[i])
if len(inputDatum) == 10:
inputData.append(inputDatum)
inputDatum = []
pprint(len(inputData))
pprint(len(inputData[999]))
if os.path.exists('examples/image_sentence/example_input.txt'):
os.remove('examples/image_sentence/example_input.txt')
with open('examples/image_sentence/example_input.txt', 'a') as outfile:
for inputDatum in inputData:
json.dump(inputDatum, outfile)
outfile.write('\n')
# outfile.close()
# -
| .ipynb_checkpoints/PrepInput-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="5APKUs5EuiUN"
# # Develop Deep Learning Models for Natural Language in Python
# + [markdown] id="XZx-g71Vo2ih"
# ## 4 - How to Develop Deep Learning Models with Keras
#
# + [markdown] id="ofNVY8FJueUO"
# ### 4.1 - Keras Model Life-*Cycle*
# + id="fXqdlqh3uDsJ"
from keras.layers import Dense
# Define the model
model = keras.Sequential()
# Add layers to the model
model.add(Dense(units = 5, input_shape = (2, ), activation = 'relu'))
model.add(Dense(units = 1, activation = 'sigmoid'))
# Compile the model
model.compile(optimizer = 'sgd', loss = 'mean_squered_error', metrics = ['mae'])
# Overall view of the model
model.summary()
# Fit the model
model.fit(X, y, epochs = 100, batch_size = 10, verbose = False)
# Evaluate on validation data
loss, mae = model.evaluate(X, y, verbose = True)
# Predict on test data
predictions = model.predict(X, verbose = False)
# + [markdown] id="pili7To5j57e"
# ### 4.2 - Keras Functional Models
# + id="J7ClXikBkA6U"
from keras.layers import Dense, Input
from keras.models import Model
# Explicate input layer
visible = Input(shape = (2, ))
# Hidden layer & chaining
hidden = Dense(units = 2)(visible)
# Build the model
model = Model(inputs = visible, outputs = hidden)
# + [markdown] id="nUZ2zBtzpcZl"
# ### 4.3 - Standard Network Models
# + [markdown] id="ctWXsMHypisp"
# #### 4.3.1 - Multilayer Perceptron
# + colab={"base_uri": "https://localhost:8080/", "height": 769} id="_hUNd3unpf5A" outputId="21f499fb-a1eb-4d2a-a25a-5b08b82deebf"
from keras.layers import Dense, Input
from keras.models import Model
from keras.utils.vis_utils import plot_model
# Input layer
visible = Input(shape = (10, ))
# Hidden layers
hidden_1 = Dense(units = 10, activation = 'relu')(visible)
hidden_2 = Dense(units = 20, activation = 'relu')(hidden_1)
hidden_3 = Dense(units = 10, activation = 'relu')(hidden_2)
# Output layer
output = Dense(1, activation = 'sigmoid')(hidden_3)
# Build model
model = Model(inputs = visible, outputs = output)
model.summary()
plot_model(model, to_file = 'multilayer perceptron graph.png')
# + [markdown] id="quVCT14Wr1hG"
# #### 4.3.2 - Convolutional Neural Network
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="hS0xuplwqhmO" outputId="7268e5fd-e66c-4960-dda2-fe2078749e05"
from keras.layers import Dense, Input
from keras.models import Model
from keras.utils.vis_utils import plot_model
from keras.layers.convolutional import Conv2D
from keras.layers.pooling import MaxPooling2D
# Input layer
visible = Input(shape = (64, 64, 1))
conv_1 = Conv2D(filters = 32, kernel_size = 4, activation = 'relu')(visible)
pool_1 = MaxPooling2D(pool_size = (2, 2))(conv_1)
conv_2 = Conv2D(filters = 16, kernel_size = 4, activation = 'relu')(pool_1)
pool_2 = MaxPooling2D(pool_size = (2, 2))(conv_2)
hidden_1 = Dense(10, activation = 'relu')(pool_2)
output = Dense(1, activation = 'sigmoid')(hidden_1)
model = Model(inputs = visible, outputs = output)
model.summary()
plot_model(model, to_file = 'convolutional neural network.png')
# + [markdown] id="ZrYZwXrxt2Rc"
# #### 4.3.3 - Recurrent Neural Network
# + colab={"base_uri": "https://localhost:8080/", "height": 638} id="qhJGjF1Zse2N" outputId="6599da4e-91e6-427e-cbef-d041cd939022"
from keras.layers import Dense, Input
from keras.utils.vis_utils import plot_model
from keras.models import Model
from keras.layers.recurrent import LSTM
# Input layer
visible = Input(shape = (100, 1))
# Hidden layers
hidden_1 = LSTM(units = 10)(visible)
hidden_2 = Dense(units = 10, activation = 'relu')(hidden_1)
# Output layer
output = Dense(units = 1, activation = 'sigmoid')(hidden_2)
model = Model(inputs = visible, outputs = output)
model.summary()
plot_model(model, to_file = 'recurrent neural network.png')
| Develop Deep Learning Models for Natural Language in Python/Chapter 4 - How to Develop Deep Learning Models with Keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.12 ('kswe')
# language: python
# name: python3
# ---
# +
import sys
sys.path.append('/Users/aitomatic/src/github/h1st-ai/h1st/h1st/model/kswe')
from kswe_modeler import KSWEModeler
from segmentor import CombinationSegmentor
# +
from sklearn import datasets, metrics
from sklearn.model_selection import train_test_split
def load_data():
df_raw = datasets.load_iris(as_frame=True).frame
df_raw.columns = ['sepal_length','sepal_width','petal_length','petal_width', 'species']
df_raw['sepal_size'] = df_raw['sepal_length'] * df_raw['sepal_width']
df_raw['sepal_aspect_ratio'] = df_raw['sepal_width'] / df_raw['sepal_length']
X_cols = list(df_raw.columns)
# X_cols.remove('species')
X_train, X_test, y_train, y_test = train_test_split(
df_raw[X_cols], df_raw['species'], test_size=0.4, random_state=1)
return {
'dataframe': {
'X_train': X_train,
'y_train': y_train,
'X_test': X_test,
'y_test': y_test,
}
}
# +
from typing import Any, Dict
from h1st.model.ml_model import MLModel
from h1st.model.ml_modeler import MLModeler
from h1st.model.rule_based_modeler import RuleBasedModeler
from sklearn.linear_model import LogisticRegression
from sklearn import metrics as sk_metrics
import pandas as pd
from segmentor import CombinationSegmentor
from ensemble import MajorityVotingEnsemble
class MySubModel(MLModel):
def predict(self, input_data: Dict) -> Dict:
y = self.base_model.predict(input_data['X'])
return {'predictions': y}
class MySubModelModeler(MLModeler):
def __init__(self, model_class=MySubModel):
self.model_class = model_class
self.stats = {}
def evaluate_model(self, data: Dict, model: MLModel) -> Dict:
# super().evaluate_model(data, model)
if 'X_test' not in data:
print('No test data found. evaluating training results')
X, y_true = data['X_train'], data['y_train']
else:
X, y_true = data['X_test'], data['y_test']
y_pred = pd.Series(model.predict({'X': X})['predictions'])
return {'r2_score': sk_metrics.r2_score(y_true, y_pred)}
def train_base_model(self, data: Dict[str, Any]) -> Any:
X, y = data['X_train'], data['y_train']
model = LogisticRegression(random_state=0)
model.fit(X, y)
return model
# +
data = load_data()
segmentation_features = {
'sepal_size': [(None, 18.5), (18.5, None)],
'sepal_aspect_ratio': [(None, 0.65), (0.65, None)],
'species': [[0, 1], [1, 2]]
}
kswe_modeler = KSWEModeler()
kswe = kswe_modeler.build_model(
input_data=data,
segmentation_features=segmentation_features,
min_data_size=30,
segmentor=CombinationSegmentor(),
sub_model_modeler=MySubModelModeler(),
ensemble_modeler=RuleBasedModeler(model_class=MajorityVotingEnsemble)
)
X_features = list(data['dataframe']['X_test'].columns)
for item in segmentation_features.keys(): X_features.remove(item)
pred = kswe.predict({'X': data['dataframe']['X_test'][X_features]})['predictions']
# +
import os
import tempfile
from kswe import KSWE
def test_kswe(kswe, data):
X, y_true = data['X_test'], data['y_test']
y_pred = pd.Series(kswe.predict({'X': X})['predictions'])
return {'r2_score': metrics.r2_score(y_true, y_pred)}
with tempfile.TemporaryDirectory() as path:
os.environ['H1ST_MODEL_REPO_PATH'] = path
print(test_kswe(kswe, data['dataframe']))
kswe.persist('my_v1')
kswe = None
kswe = KSWE(
segmentor=CombinationSegmentor(),
sub_model=MySubModel,
ensemble=MajorityVotingEnsemble()
)
kswe.load_params('my_v1')
print(test_kswe(kswe, data['dataframe']))
# -
test_data = {
'X_test': data['dataframe']['X_test'][X_features],
'y_test': data['dataframe']['y_test']
}
for name, model in kswe.sub_models.items():
metrics = MySubModelModeler().evaluate_model(test_data, model)
print(f'sub model {name} test resluts based on {test_data["X_test"].shape[0]} samples: {metrics}')
kswe.sub_model
assert 1 == 0
data = load_data()
data['dataframe'].keys()
'dataframe' not in data
if 'dataframe' not in data or 'json' not in data:
raise KeyError('key "dataframe" or "json" is not in your input_data')
# +
# df_0['sepal_aspect_ratio'].hist(bins=20
# +
# df_0['sepal_size'].hist(bins=20)
# -
segmentation_features = {
'sepal_size': [(None, 18.5), (18.5, None)],
'sepal_aspect_ratio': [(None, 0.65), (0.65, None)],
# 'species': [[0]]
}
cs = CombinationSegmentor()
# +
features = ['sepal_length', 'sepal_width', 'petal_length', 'petal_width']
results, filter_combinations = cs.process(
data,
by=segmentation_features,
min_data_size=40,
levels=[2]
)
# -
filter_combinations['segment_0_lvl_2']
for k, v in results.items():
print(k, v['X'].shape)
filter_combinations
temp = {'df': 2, 'ddd': 6}
list(temp.values())[0]
X = results['segment_0_lvl_2'].iloc[:, :-1]
X.shape
df_all['species'].loc[X.index]
# +
def save_coco(file, info, licenses, images, annotations, categories):
with open(file, 'wt', encoding='UTF-8') as coco:
json.dump({'info': info, 'licenses': licenses, 'images': images,
'annotations': annotations, 'categories': categories}, coco, indent=2, sort_keys=True)
def filter_annotations(annotations, images):
image_ids = funcy.lmap(lambda i: int(i['id']), images)
return funcy.lfilter(lambda a: int(a['image_id']) in image_ids, annotations)
# +
def main(args):
with open(args.annotations, 'rt', encoding='UTF-8') as annotations:
coco = json.load(annotations)
info = coco['info']
licenses = coco['licenses']
images = coco['images']
annotations = coco['annotations']
categories = coco['categories']
number_of_images = len(images)
images_with_annotations = funcy.lmap(lambda a: int(a['image_id']), annotations)
if args.having_annotations:
images = funcy.lremove(lambda i: i['id'] not in images_with_annotations, images)
x, y = train_test_split(images, train_size=args.split)
save_coco(args.train, info, licenses, x, filter_annotations(annotations, x), categories)
save_coco(args.test, info, licenses, y, filter_annotations(annotations, y), categories)
print("Saved {} entries in {} and {} in {}".format(len(x), args.train, len(y), args.test))
# +
import json
data_path = '/Users/aitomatic/Desktop/dataset/furuno/sample_15mins/annotations.json'
with open(data_path, 'r', encoding='UTF-8') as annotations:
coco = json.load(annotations)
info = coco['info']
licenses = coco['licenses']
images = coco['images']
annotations = coco['annotations']
categories = coco['categories']
# -
logic_example = filter_combinations['segment_0_lvl_2']
logic_example
import funcy
# 1. create new features and save that in annotation
# ex)
# - depth_of_bb
# - aspect_ratio_of_bb
# - size_of_bb
# - datetime
#
# 2. make logics in this format. [('depth_of_bb', (None, 200)), ('aspect_ratio_of_bb', (None, 0.65))]
#
# 3. make a function get_segments_from_json(JSON, segmentation_logics) -> return segmented JSONs
# - make sure images, annotations, and categories are synchronized.
#
# 4. save those JSONs and move around files based on that json
def create_sample_features():
annotation_json_path = '/Users/aitomatic/Desktop/dataset/furuno/sample_15mins/annotations.json'
with open(annotation_json_path, 'r', encoding='UTF-8') as annotations:
coco = json.load(annotations)
info = coco['info']
licenses = coco['licenses']
images = coco['images']
annotations = coco['annotations']
categories = coco['categories']
for idx in range(len(annotations)):
print(idx)
images[0]
annotations[0]
| tests/model/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # Models
#
# Welcome to the "**Models**" tutorial of the "_From Zero to Hero_" series. In this notebook we will talk about the features offered by the `models` _Avalanche_ sub-module.
#
# ### Models Support
#
# Every continual learning experiment needs a model to train incrementally. The `models` sub-module provide ready-to-use **randomly initialized** and **pre-trained** models you can use _off-the-shelf_.
#
# At the moment we support only the following architectures:
# + pycharm={"name": "#%%\n"}
# !pip install git+https://github.com/ContinualAI/avalanche.git
# + pycharm={"name": "#%%\n"}
from avalanche.models import SimpleCNN
from avalanche.models import SimpleMLP
from avalanche.models import SimpleMLP_TinyImageNet
from avalanche.models import MobilenetV1
# -
# However, we plan to support in the near future all the models provided in the [Pytorch](https://pytorch.org/) official ecosystem models as well as the ones provided by [pytorchcv](https://pypi.org/project/pytorchcv/)!
# ## 🤝 Run it on Google Colab
#
# You can run _this chapter_ and play with it on Google Colaboratory: [](https://colab.research.google.com/github/ContinualAI/colab/blob/master/notebooks/avalanche/models.ipynb)
| notebooks/avalanche/models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Effects of borrowers' financial background on their APR
# ## by <NAME>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Investigation Overview
#
# > In this investigation, I wanted to look at the financial background of borrowers that could be used to predict their APR. The main focus was on the five variables: income range, available bank card credit, monthly loan payment, bank card utilization and loan term.
#
#
# ## Dataset Overview
#
# > The data consisted of APR and attributes related to loans during 2007-2014 of approximately 110,000 loans and 81 variables including term, date, APR and information about the borrowers such as income range and available bank card credit when open the current loan. We'll pick up around 10 variables to do further analysis and see what's affecting borrower APR. The data that BankCardUtilization is more than 1 has been removed.
# + slideshow={"slide_type": "skip"}
# import all packages and set plots to be embedded inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sb
# %matplotlib inline
# suppress warnings from final output
import warnings
warnings.simplefilter("ignore")
# + slideshow={"slide_type": "skip"}
# load in the dataset into a pandas dataframe
loan=pd.read_csv('prosperLoanData.csv')
# convert LoanStatus and IncomeRange into ordered categorical types
ordinal_var_dict = {'LoanStatus': ['Completed','FinalPaymentInProgress','Past Due (1-15 days)','Past Due (16-30 days)',
'Past Due (31-60 days)','Past Due (61-90 days)','Past Due (91-120 days)',
'Past Due (>120 days)','Defaulted','Chargedoff','Cancelled'],
'IncomeRange': ['$100,000+', '$75,000-99,999', '$50,000-74,999', '$25,000-49,999',
'$1-24,999', '$0', 'Not employed','Not displayed'],
'Term':[12,36,60]}
for var in ordinal_var_dict:
ordered_var = pd.api.types.CategoricalDtype(ordered = True,
categories = ordinal_var_dict[var])
loan[var] = loan[var].astype(ordered_var)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Distribution of Borrower APR
#
# > The BorrowerAPR distribution looks bimodal,with the first peak 0.1-0.2 and second peak(even more) 0.3-0.4. Most of the loan APRs are in 0.1-0.3, but the second peak is an exception and need more attention.
# + slideshow={"slide_type": "subslide"}
binsize=0.01
bins=np.arange(0,max(loan.BorrowerAPR)+binsize,binsize)
plt.figure(figsize=[6,6])
plt.hist(data=loan,x='BorrowerAPR',bins=bins)
plt.xlabel('BorrowerAPR(%)')
plt.ylabel('Count')
plt.title('Distribution of Borrower APR',fontsize=14,weight='bold')
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ## BorrowerAPR by LoanStatus, IncomeRange and Term
#
# > Clearly, `IncomeRange` seems to be most relevant to the `BorrowerAPR`, with lower income, the APR goes up except when income range is $'0'$. There may be some input error for the $'0'$ as it equals to $'not$ $employed'$. So this exception won't change our conclusion. Term has little effect on the APR. High APR loans are usually easier to be 'past due', which means APR can represent risk level.
# + slideshow={"slide_type": "subslide"}
numeric_vars = ['BorrowerAPR', 'DebtToIncomeRatio', 'MonthlyLoanPayment', 'CurrentCreditLines', 'OpenRevolvingMonthlyPayment',
'RevolvingCreditBalance', 'BankcardUtilization','InquiriesLast6Months','TotalInquiries','DelinquenciesLast7Years',
'AvailableBankcardCredit']
categoric_vars = ['LoanStatus', 'IncomeRange', 'Term']
def boxgrid(x, y, **kwargs):
""" Quick hack for creating box plots with seaborn's PairGrid. """
default_color = sb.color_palette()[0]
sb.boxplot(x, y, color = default_color)
plt.xticks(rotation=60)
g = sb.PairGrid(data = loan, y_vars = ['BorrowerAPR'], x_vars = categoric_vars,
height=8.27/1,aspect = (14.7/3)/(8.27/1));
g.map(boxgrid);
plt.suptitle('Distribution of APR',y=1,fontsize=14,weight='bold')
plt.tight_layout()
plt.show();
# + [markdown] slideshow={"slide_type": "slide"}
# ## Montly Loan Payment by Income Range
#
# There's a declining trend of MonthlyLoanPayment when the income declines, but not very significant.
# + slideshow={"slide_type": "subslide"}
plt.figure(figsize=[10,6]);
default_color = sb.color_palette()[0];
sb.violinplot(data = loan, x = 'IncomeRange', y = 'MonthlyLoanPayment',
color = default_color);
plt.xticks(rotation=60);
plt.title('Montly Loan Payment by Income Range',fontsize=14,weight='bold');
# + [markdown] slideshow={"slide_type": "slide"}
# ## Available Bank Card Credit by Income Range
#
# There's a declining trend of available bank card credit when the income declines, but not very significant.
# + slideshow={"slide_type": "subslide"}
def log_trans(x, inverse = False):
""" quick function for computing log and power operations """
if not inverse:
return np.log10(x+1)
else:
return np.power(10, x+1)
loan['log_AvailableBankcardCredit'] = loan['AvailableBankcardCredit'].apply(log_trans)
plt.figure(figsize=[10,6])
sb.violinplot(data = loan, x = 'IncomeRange',y='log_AvailableBankcardCredit',
color = default_color)
plt.xticks(rotation=60)
plt.title('Available Bank Card Credit by Income Range',fontsize=14,weight='bold')
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Relationship among Income Range, Available Bank Card Credit and Monthly Loan Payment
#
# Looks like high income people usually have high monthly loan payment. It can explain why income range didn't affect the APR when available bank card credit is the same. If a hoursehold has a high income but high monthly loan payment, the bank won't give it a very low APR.
# + slideshow={"slide_type": "subslide"}
bins=list(np.arange(min(loan['MonthlyLoanPayment']),loan['MonthlyLoanPayment'].quantile(0.99)+100,100))+[max(loan['MonthlyLoanPayment'])]
bins=[int(x) for x in bins]
labels=[f"{bins[i]}-{bins[i+1]}" for i in range(len(bins)-1)]
loan['MonthlyLoanPayment_c']=pd.cut(loan['MonthlyLoanPayment'],bins=bins,labels=labels,right=False)
plt.figure(figsize=[15,8]);
sb.stripplot(data=loan,x='IncomeRange',y='log_AvailableBankcardCredit',hue='MonthlyLoanPayment_c',jitter=0.35,dodge=True,
palette='RdYlGn_r');
plt.title('Available Bank Card Credit by Income Range and Monthly Loan Payment',fontsize=14,weight='bold')
plt.legend(loc='center left',bbox_to_anchor=(1,0.5),title='Monthly Loan Payment($)',title_fontsize=12);
# + [markdown] slideshow={"slide_type": "slide"}
# ## APR by IncomeRange and Term
# In terms of high income range, the BorrowerAPR didn't change much across different terms. But for low income range, shorter term tend to have higher APR.
# + slideshow={"slide_type": "subslide"}
fig = plt.figure(figsize = [10,6])
ax = sb.pointplot(data = loan, x = 'IncomeRange', y = 'BorrowerAPR', hue = 'Term',
palette = 'Greens_r', linestyles = '', dodge = 0.4)
plt.title('APR by IncomeRange and Term',fontsize=14,weight='bold')
plt.ylabel('BorrowerAPR(%)')
plt.yticks(np.array([0, 0.1, 0.2, 0.3, 0.4, 0.5,0.6]), [0, 0.1, 0.2, 0.3, 0.4, 0.5,0.6])
ax.set_yticklabels([],minor = True)
plt.xticks(rotation=60)
plt.show();
# + slideshow={"slide_type": "skip"}
# !jupyter nbconvert slide_deck_Loan.ipynb --to slides --post serve --no-input --no-prompt
| slide_deck_Loan-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 위치정보요약 데이터의 좌표 정보로 각 도로명 주소별 평균 위도, 경도 table 만들기
# +
# pyproj로 주소정보 데이터 위도, 경도로 바꿔보기
import pyproj
from pyproj import Proj
from pyproj import transform
# +
#연세대학교 위치. X과표 : 동서, Y좌표 : 남북, GRS80 UTM-K
# # +proj=tmerc, +lat_0=38, +lon_0=127.5, +k=0.9996, +x_0=1000000, +y_0=2000000, +ellps=GRS80, +units=m, +no_defs
lx=950254.198668
ly=1951390.210053
# 구글 지도 epsg 3857 or epsg 900913
# # +proj=merc, +a=6378137, +b=6378137, +lat_ts=0.0, +lon_0=0.0, +x_0=0.0, +y_0=0, +k=1.0, +units=m, +nadgrids=@null, +no_defs ->이건 구글 map 좌표.
# 우리가 필요한 건 경도, 위도
# https://www.osgeo.kr/17
# transfom(p1, p2, x,y, z=None, radians =False) : p1에서 정의된 x1,y1,z1 점을 p2에서 정의된 x2,y2로 바꾸는 것. z, radian은 default가 none,false인 듯
# -
GRS80={'proj':'tmerc', 'lat_0':'38', 'lon_0':'127.5', 'k':0.9996, 'x_0':1000000, 'y_0':2000000, 'ellps':'GRS80', 'units':'m',}
p1=Proj(**GRS80)
#EPSG={'proj':'merc', 'a':6378137, 'b':6378137, 'lat_ts':'0.0', 'lon_0':'0.0', 'x_0':0.0, 'y_0':0, 'k':1.0, 'units':'m'}
p2=Proj(init='epsg:4019')
nlx,nly = transform(p1,p2,lx,ly)
# +
map_pos=list((nly,nlx))
#m=folium.Map(location=map_pos,min_zoom=11, max_zoom=20)
map_pos
# -
import pandas as pd
import numpy as np
import os
import glob #파일 한꺼번에 부르기 위해서 사용
import re
glob.glob(os.path.join("gh-data/loc-data","*.*"))
filenames = glob.glob(os.path.join("gh-data/loc-data","entrc*.xlsx"))
filenames
df0=pd.read_excel('gh-data/loc-data/entrc_busan.xlsx',header = 0,encoding='utf-8')
df0.head()
df=pd.read_excel('gh-data/entrc_seoul_head.xlsx',header = 0,encoding='utf-8')
df_refine=df[['시도','시군구','도로명','x좌표','y좌표']]
df_refine.dtypes
df_refine.keys()
df_refine['x좌표'][0], df_refine['y좌표'][0]
# +
a,b= transform(p1,p2,df_refine['x좌표'][0], df_refine['y좌표'][0])
a,b
# -
len(df_refine)
df_refine.describe()
# +
lx=df_refine['x좌표'][67]
ly=df_refine['y좌표'][67]
lx>0
# +
gx=[]
gy=[]
null=[]
#
#67번 데이터,
for k in list(range(len(df_refine))) :
lx=df_refine['x좌표'][k]
ly=df_refine['y좌표'][k]
if lx>0 and ly>0 :
a,b= transform(p1,p2,lx,ly)
gx.append(a)
gy.append(b)
else :
gx.append(0)
gy.append(0)
null.append(k)
len(null)
# +
df_refine['위도']=gy
df_refine['경도']=gx
df_refine
# -
type(null)
df_refine_final=df_refine.drop(null,0) #가로일때 0, 세로일때 1
df_refine_final.describe()
# +
a1=df_refine_final['시도'][0]
a2=df_refine_final['시군구'][0]
a3=df_refine_final['도로명'][0]
ab=[a1,a2,a3]
new=' '.join(ab) #이렇게 하는게 처리 속도가 더 빠르다고 함
new
# +
df_refine_final.reset_index(inplace=True,drop=True)
address=[]
#list(range(len(df_refine_final)))
for i in list(range(len(df_refine_final))) :
a1=df_refine_final['시도'][i]
a2=df_refine_final['시군구'][i]
a3=df_refine_final['도로명'][i]
ab=[a1,a2,a3]
new=' '.join(ab)
address.append(new)
address
# +
df_refine_final["도로주소"]=address
AveSeoul=pd.pivot_table(df_refine_final, index=["도로주소"])
# -
AveSeoul
AveSeoul.to_excel('gh-data/2018_loc_for_add_in_seoul.xlsx',encoding='utf-8')
# +
AA=pd.read_excel('gh-data/2018_loc_for_add_in_seoul.xlsx',index_col=None,encoding='utf-8')
AA
| .ipynb_checkpoints/nims-academy-190228-yonsei-oh-report02-00-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# metadata:
# interpreter:
# hash: bb6bc14e21ab582ba23f9445a6e8cb0350b4a77bb7b394131a2cec33c0dfc60e
# name: Python 3.8.5 64-bit
# ---
# #### This notebook shows how to read the fastMRI dataset and apply some simple transformations to the data.
# +
# Testing if integration works
# +
# %matplotlib inline
import h5py
import numpy as np
from matplotlib import pyplot as plt
# -
# The fastMRI dataset is distributed as a set of HDF5 files and can be read with the h5py package. Here, we show how to open a file from the multi-coil dataset. Each file corresponds to one MRI scan and contains the k-space data, ground truth and some meta data related to the scan.
file_name = 'multicoil_train/file1000167.h5'
hf = h5py.File(file_name)
# + tags=[]
print('Keys:', list(hf.keys()))
print('Attrs:', dict(hf.attrs))
# -
# In multi-coil MRIs, k-space has the following shape:
# (number of slices, number of coils, height, width)
#
# For single-coil MRIs, k-space has the following shape:
# (number of slices, height, width)
#
# MRIs are acquired as 3D volumes, the first dimension is the number of 2D slices.
# + tags=[]
volume_kspace = hf['kspace'][()]
print(volume_kspace.dtype)
print(volume_kspace.shape)
# -
slice_kspace = volume_kspace[20] # Choosing the 20-th slice of this volume
# Let's see what the absolute value of k-space looks like:
def show_coils(data, slice_nums, cmap=None):
fig = plt.figure()
for i, num in enumerate(slice_nums):
plt.subplot(1, len(slice_nums), i + 1)
plt.imshow(data[num], cmap=cmap)
show_coils(np.log(np.abs(slice_kspace) + 1e-9), [0, 5, 10]) # This shows coils 0, 5 and 10
# The fastMRI repo contains some utlity functions to convert k-space into image space. These functions work on PyTorch Tensors. The to_tensor function can convert Numpy arrays to PyTorch Tensors.
import fastmri
from fastmri.data import transforms as T
slice_kspace2 = T.to_tensor(slice_kspace) # Convert from numpy array to pytorch tensor
slice_image = fastmri.ifft2c(slice_kspace2) # Apply Inverse Fourier Transform to get the complex image
slice_image_abs = fastmri.complex_abs(slice_image) # Compute absolute value to get a real image
show_coils(slice_image_abs, [0, 5, 10], cmap='gray')
# As we can see, each coil in a multi-coil MRI scan focusses on a different region of the image. These coils can be combined into the full image using the Root-Sum-of-Squares (RSS) transform.
slice_image_rss = fastmri.rss(slice_image_abs, dim=0)
plt.imshow(np.abs(slice_image_rss.numpy()), cmap='gray')
# So far, we have been looking at fully-sampled data. We can simulate under-sampled data by creating a mask and applying it to k-space.
from fastmri.data.subsample import RandomMaskFunc
mask_func = RandomMaskFunc(center_fractions=[0.04], accelerations=[8]) # Create the mask function object
masked_kspace, mask = T.apply_mask(slice_kspace2, mask_func) # Apply the mask to k-space
# Let's see what the subsampled image looks like:
sampled_image = fastmri.ifft2c(masked_kspace) # Apply Inverse Fourier Transform to get the complex image
sampled_image_abs = fastmri.complex_abs(sampled_image) # Compute absolute value to get a real image
sampled_image_rss = fastmri.rss(sampled_image_abs, dim=0)
plt.imshow(np.abs(sampled_image_rss.numpy()), cmap='gray')
| fastMRI_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="qDEeD8_NBbSc" colab_type="text"
# # Tuples
#
# In Python tuples are very similar to lists, however, unlike lists they are *immutable* meaning they can not be changed. You would use tuples to present things that shouldn't be changed, such as days of the week, or dates on a calendar.
#
# In this section, we will get a brief overview of the following:
#
# 1.) Constructing Tuples
# 2.) Basic Tuple Methods
# 3.) Immutability
# 4.) When to Use Tuples
#
# You'll have an intuition of how to use tuples based on what you've learned about lists. We can treat them very similarly with the major distinction being that tuples are immutable.
#
# ## Constructing Tuples
#
# The construction of a tuples use () with elements separated by commas. For example:
# + id="PI539KXHBbSe" colab_type="code" colab={}
# Create a tuple
t = (1,2,3)
# + id="dU_pWYhQBbSi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1595194899366, "user_tz": 420, "elapsed": 752, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjNCW3KoaFGo2LYk117G2iPwFMkjFedUPcICW8sxQ=s64", "userId": "02293805457177884744"}} outputId="f61a7518-15a9-49fa-e99b-f2456b22a159"
# Check len just like a list
len(t)
# + id="QtnrN6_PBbSm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1595194899369, "user_tz": 420, "elapsed": 729, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjNCW3KoaFGo2LYk117G2iPwFMkjFedUPcICW8sxQ=s64", "userId": "02293805457177884744"}} outputId="a605d63d-bc64-4a9a-d5d1-8e4876cec8a6"
# Can also mix object types
t = ('one',2)
# Show
t
# + id="xyu4CGwZBbSq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1595194899370, "user_tz": 420, "elapsed": 705, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjNCW3KoaFGo2LYk117G2iPwFMkjFedUPcICW8sxQ=s64", "userId": "02293805457177884744"}} outputId="74b964e4-5f74-4065-8fa5-90c459048608"
# Use indexing just like we did in lists
t[0]
# + id="jVjWrQYUBbSt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1595194899372, "user_tz": 420, "elapsed": 695, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjNCW3KoaFGo2LYk117G2iPwFMkjFedUPcICW8sxQ=s64", "userId": "02293805457177884744"}} outputId="d6e5e968-b9aa-44f5-b8c0-13cb8f020a01"
# Slicing just like a list
t[-1]
# + [markdown] id="DBLmUGHfBbSw" colab_type="text"
# ## Basic Tuple Methods
#
# Tuples have built-in methods, but not as many as lists do. Let's look at two of them:
# + id="9iAxkxEfBbSx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1595194899373, "user_tz": 420, "elapsed": 682, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjNCW3KoaFGo2LYk117G2iPwFMkjFedUPcICW8sxQ=s64", "userId": "02293805457177884744"}} outputId="0706aae9-9e57-44f4-f50d-5ff2b47b8f8c"
# Use .index to enter a value and return the index
t.index('one')
# + id="YxcUO8JhBbS2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1595194899374, "user_tz": 420, "elapsed": 665, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjNCW3KoaFGo2LYk117G2iPwFMkjFedUPcICW8sxQ=s64", "userId": "02293805457177884744"}} outputId="9f509340-d979-45b0-b12a-22fd6682ce31"
# Use .count to count the number of times a value appears
t.count('one')
# + [markdown] id="gchvjroRBbS5" colab_type="text"
# ## Immutability
#
# It can't be stressed enough that tuples are immutable. To drive that point home:
# + id="x5Omi4MfBbS6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 166} executionInfo={"status": "error", "timestamp": 1595194899541, "user_tz": 420, "elapsed": 818, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjNCW3KoaFGo2LYk117G2iPwFMkjFedUPcICW8sxQ=s64", "userId": "02293805457177884744"}} outputId="3a7b0e56-e49b-40f8-d92e-4a105b740a66"
t[0]= 'change'
# + [markdown] id="DSTAAnzkBbS_" colab_type="text"
# Because of this immutability, tuples can't grow. Once a tuple is made we can not add to it.
# + id="jNZlWZrzBbS_" colab_type="code" colab={}
t.append('nope')
# + [markdown] id="rbJsQW4vBbTD" colab_type="text"
# ## When to use Tuples
#
# You may be wondering, "Why bother using tuples when they have fewer available methods?" To be honest, tuples are not used as often as lists in programming, but are used when immutability is necessary. If in your program you are passing around an object and need to make sure it does not get changed, then a tuple becomes your solution. It provides a convenient source of data integrity.
#
# You should now be able to create and use tuples in your programming as well as have an understanding of their immutability.
#
# Up next Sets and Booleans!!
| DigitalHistory/Week-1/Atul_Python_Notes/Tuples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
from tensorflow.keras.datasets import cifar10
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras import Sequential
from tensorflow.keras.layers import Dense, Dropout, Activation, Flatten, BatchNormalization, Conv2D, MaxPooling2D
from tensorflow.keras.regularizers import l2
import numpy as np
import os
import matplotlib.pyplot as plt
# %matplotlib inline
# Defining the parameters
batch_size = 32
num_classes = 10
epochs = 50
# Splitting the data between train and test
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# +
# plotting some random 10 images
class_names = ['airplane','automobile','bird','cat','deer',
'dog','frog','horse','ship','truck']
fig = plt.figure(figsize=(8,3))
for i in range(num_classes):
ax = fig.add_subplot(2, 5, 1 + i, xticks=[], yticks=[])
idx = np.where(y_train[:]==i)[0]
features_idx = x_train[idx,::]
img_num = np.random.randint(features_idx.shape[0])
im = (features_idx[img_num,::])
ax.set_title(class_names[i])
plt.imshow(im)
plt.show()
# -
# Convert class vectors to binary class matrices.
y_train = tf.keras.utils.to_categorical(y_train, num_classes)
y_test = tf.keras.utils.to_categorical(y_test, num_classes)
# Printing sample data
print(y_train[:10])
# +
model = Sequential()
model.add(Conv2D(32, (3, 3), padding='same',
input_shape=x_train.shape[1:]))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Conv2D(32, (3, 3)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Conv2D(64, (3, 3), padding='same'))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Conv2D(64, (3, 3)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Conv2D(128, (3, 3), padding='same'))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Conv2D(128, (3, 3)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(512,kernel_regularizer=l2(0.01)))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes))
model.add(Activation('softmax'))
# -
# summary of the model
print(model.summary())
# +
# compile
model.compile(loss='categorical_crossentropy',
optimizer='sgd',
metrics=['accuracy'])
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
# Normalizing the input image
x_train /= 255
x_test /= 255
# -
# Training the model
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
validation_data=(x_test, y_test),
shuffle=True)
# list all data in history
print(history.history.keys())
# summarize history for accuracy
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# summarize history for loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
| 11. Convolutional Neural Networks/2. Building CNN with Keras and Python/7. Cifar10 morelayer notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# perform sentiment analysis
import numpy as np
import pickle
import collections
import nltk.classify.util, nltk.metrics
from nltk.classify import NaiveBayesClassifier, MaxentClassifier, SklearnClassifier
import csv
from sklearn import cross_validation
from sklearn.svm import LinearSVC, SVC
import random
from nltk.corpus import stopwords
import itertools
from nltk.collocations import BigramCollocationFinder
from nltk.metrics import BigramAssocMeasures
import pandas as pd
# unigrams features
#Finding the unigram representation
from sklearn.feature_extraction.text import CountVectorizer
vectorizer=CountVectorizer()
with open('sentiments/SentimentText_train.txt', 'rb') as f:
SentimentText = pickle.load(f)
X=vectorizer.fit_transform(SentimentText)
train_data = pd.read_csv('data/train.csv', encoding = 'ISO-8859-1')
y = train_data['Sentiment']
y.shape
X.shape
from sklearn.model_selection import train_test_split
X_train, X_test, y_train ,y_test = train_test_split(X,y,train_size = .8 , test_size = .2 , random_state = 0)
from sklearn.naive_bayes import MultinomialNB
clf = MultinomialNB()
clf.fit(X, y)
# training score
clf.score(X_train,y_train)
# testing score
clf.score(X_test, y_test)
# using bigrams features
# #Finding the bigram representation
bigram_vectorizer=CountVectorizer(ngram_range=(1,2))
with open('sentiments/SentimentText_train.txt', 'rb') as f:
SentimentText = pickle.load(f)
X=bigram_vectorizer.fit_transform(SentimentText)
train_data = pd.read_csv('data/train.csv', encoding = 'ISO-8859-1')
y = train_data['Sentiment']
X.shape
y.shape
from sklearn.model_selection import train_test_split
X_train, X_test, y_train ,y_test = train_test_split(X,y,train_size = .8 , test_size = .2 , random_state = 0)
from sklearn.naive_bayes import MultinomialNB
clf = MultinomialNB()
clf.fit(X, y)
# training score
clf.score(X_train, y_train)
# testing score
clf.score(X_test, y_test)
with open('sentiments/SentimentText_test.txt', 'rb') as f:
SentimentText = pickle.load(f)
X_test=bigram_vectorizer.transform(SentimentText)
X_test.shape
predicted = clf.predict(X_test)
predicted.shape
output = pd.read_csv('data/test.csv', encoding = 'ISO-8859-1')
output.drop(['SentimentText'],axis = 1)
output['Sentiment'] = predicted
output.to_csv('output.csv', index = False)
# using both unigrams and bigrams as features set
# +
#Finding the unigram representation
from sklearn.feature_extraction.text import CountVectorizer
vectorizer=CountVectorizer()
# #Finding the bigram representation
bigram_vectorizer=CountVectorizer(ngram_range=(1,2))
# -
with open('sentiments/SentimentText_train.txt', 'rb') as f:
SentimentText = pickle.load(f)
X_unigrams = vectorizer.fit_transform(SentimentText)
X_bigrams = bigram_vectorizer.fit_transform(SentimentText)
X_unigrams.shape
X_bigrams.shape
from scipy.sparse import hstack
X_combined = hstack([X_unigrams , X_bigrams])
X_combined.shape
train_data = pd.read_csv('data/train.csv', encoding = 'ISO-8859-1')
y = train_data['Sentiment']
clf = MultinomialNB()
clf.fit(X_combined, y)
with open('sentiments/SentimentText_test.txt', 'rb') as f:
SentimentText = pickle.load(f)
X_test_unigrams = vectorizer.transform(SentimentText)
X_test_bigrams = bigram_vectorizer.transform(SentimentText)
X_test_unigrams.shape
X_test_bigrams.shape
from scipy.sparse import hstack
X_test_combined = hstack([X_test_unigrams , X_test_bigrams])
X_test_combined.shape
predicted = clf.predict(X_test_combined)
output = pd.read_csv('data/test.csv', encoding = 'ISO-8859-1')
output.drop(['SentimentText'],axis = 1)
output['Sentiment'] = predicted
output.to_csv('output.csv', index = False)
| .ipynb_checkpoints/program-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Solve time dependent Schroedinger equation in 1D, using FFT method
# Shaking harmonic oscillator 2 (quadrupole oscillations)
import numpy as np
from scipy.fftpack import fft, ifft
from scipy.integrate import simps # 'simps' is Simpson's Rule for integrating
import matplotlib.pyplot as plt
# %matplotlib inline
# The general solution follows the method of http://jakevdp.github.com/blog/2012/09/05/quantum-python/
#
# * Use the potential to propagate a half time step in x space
# * FFT
# * Use the kinetic energy operator to propagate one whole time step in k space
# * IFFT
# * Use the potential to propagate a half time step in x space
#
# For time varying potentials, the propagation term is exp(-i integral(V(x),dt)/hbar). The integral can be done analytically or numerically. Here, if numerical integration is used, we use one step of the trapezoidal approximation. This may not work if the time step is too large.
#
# Hard boundaries are assumed. Be sure xmin and xmax are sufficiently far from the region of interest for the initial state you choose.
#constants (change these to fit the problem)
hbar = 1
m = 1 #mass
tmin = 0 # initial time
tmax = 10*2*np.pi # final time
Nt = 4000 # number of time steps
xmin = -20 # minimum x value
xmax = 20 # maximum x value
Nx = 4096 # number of steps in x (and k). Must be even, power of 2 is better
#calculate lists
xlist = np.linspace(xmin,xmax,Nx)
tlist = np.linspace(tmin,tmax,Nt)
dx = xlist[1]-xlist[0] # delta x
dt = tlist[1]-tlist[0] # delta t
dk = 2 * np.pi/np.abs(xmax-xmin) # delta k (from FFT definition)
kmax = 0.5*Nx*dk # (Nyquist limit)
klist = np.roll(np.arange(-Nx//2+1,Nx//2+1),Nx//2+1)*dk #list of k values, indexed according to FFT convention, double // means integer part of quotient
# The potential below is a harmonic oscillator potential with sinusoidally varying spring constant. (Parametric amplification)
#
# Try playing around with the value of f to see how the system behaves. An interesting range is between 2 and 4.
#define potential function (needs to be vectorizable)
a=0.5 #amplitude of spring modulation
f=2.0 # frequency of spring modulation
def V(x,t):
return 0.5*(x*(1+a*np.sin(f*t)))**2
# integral of V dt, evaluated at x
def intV(x,ti,tf):
#indef = lambda x,t: 0.5*x**2*((0.5+0.5*a**2)*t-2*a/f*np.cos(f*t)-0.25*a**2/f*np.sin(2*f*t) )# indefinite integral
#out = indef(x,tf)-indef(x,ti)
out = 0.5*(V(x,ti)+V(x,tf))*(tf-ti) #trapezoidal rule (backup plan)
return out
#initial wavefunction at t=tmin (normalization optional)
def psi0(x):
a=1.0
# this should be the ground state wavefunction, but it needs a bit of a fudge factor to be stationary under the numerical approximations
return np.exp(-np.sqrt(0.505)*(x-a)**2)/np.pi**(0.25)
psilist = np.zeros([Nx,Nt],dtype=np.cfloat) # initialize array to store wavefunction
psilist[:,0]=psi0(xlist) # store initial wavefunction
#main loop
for tindex in np.arange(1,Nt):
psix = psilist[:,tindex-1]*np.exp(-1.j*intV(xlist,tlist[tindex-1],tlist[tindex]+0.5*dt)/hbar)
psix[0:3] = 0; psix[-4:-1] = 0; # enforce boundary conditions
psik = fft(psix)
psik = psik * np.exp(-0.5j*hbar*klist*klist*dt/m)
psix = ifft(psik)
psix = psix*np.exp(-1.j*intV(xlist,tlist[tindex]-0.5*dt,tlist[tindex])/hbar)
psix[0:3] = 0; psix[-4:-1] = 0; # enforce boundary conditions
psilist[:,tindex] = psix
fig, ax1 = plt.subplots()
tdraw = -1 # time index to plot (-1 = final time)
Nf = simps(np.abs(psilist[:,tdraw])**2) # normalization of final state
Ni = simps(np.abs(psilist[:,0])**2) # normalization of initial state
ax1.plot(xlist,np.abs(psilist[:,tdraw])**2/Nf,label='t={0:.1f}'.format(tlist[tdraw]))
ax1.plot(xlist,np.abs(psilist[:,0])**2/Ni,'k',label='t={0:.1f}'.format(tmin))
ax2 = ax1.twinx()
ax2.plot(xlist,V(xlist,0),'r',label='V(x)')
ax1.set_ylabel('$|\psi(x)|^2$')
ax2.set_ylabel('$V(x)$')
ax1.set_xlabel('$x$')
ax1.legend()
ax1.set_title("Initial and final wavefunction")
ax2.legend()
# Calculate expectation values of x, p, x2, p2, E
EVxlist = np.zeros(Nt)
EVplist = np.zeros(Nt)
EVx2list = np.zeros(Nt)
EVp2list = np.zeros(Nt)
Nlistx = np.zeros(Nt)
Nlistp = np.zeros(Nt)
for t in range(Nt):
Nlistx[t] = simps(np.abs(psilist[:,t])**2)
EVxlist[t] = simps(xlist*np.abs(psilist[:,t])**2)/Nlistx[t]
EVx2list[t] = simps(xlist**2*np.abs(psilist[:,t])**2)/Nlistx[t]-EVxlist[t]**2
psik = fft(psilist[:,t])
Nlistp[t] = simps(np.abs(psik)**2)
EVplist[t] = 0.5*hbar/m*simps(klist*np.abs(psik)**2)/Nlistp[t]
EVp2list[t] = (simps((0.5*hbar/m*klist)**2*np.abs(psik)**2)/Nlistp[t]-EVplist[t]**2)
Elist = 0.5*EVp2list + 0.5*EVx2list
plt.plot(tlist,EVxlist,label=r'$\langle x \rangle$')
plt.plot(tlist,np.sqrt(EVx2list),label=r'$\sqrt{\langle x^2 \rangle-\langle x \rangle ^2}$')
plt.legend()
plt.xlabel('Time')
plt.title('Center of mass and width')
plt.plot(tlist,EVplist,label=r'$\langle p \rangle$')
plt.plot(tlist,np.sqrt(EVp2list),label=r'$\sqrt{\langle p^2 \rangle-\langle p \rangle ^2}$')
plt.legend()
plt.xlabel('Time')
plt.title("Average momentum and momentum width")
plt.plot(tlist,Elist,label=r'$\langle E \rangle$')
#plt.plot(tlist,np.sqrt(EVx2list),label=r'$\sqrt{\langle x^2 \rangle-\langle x \rangle ^2}$')
plt.legend()
plt.xlabel('Time')
plt.title('Energy')
| TDSE/TDSE-3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/francomanca93/liberalismo-y-medioambiente/blob/master/Notebooks/1_Extraccion_y_limpieza_de_datos.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="tSX_sZLj-ije" colab_type="text"
# # Extracción de datos. Conociendo a los mismos.
#
# Este documento es sobre como se irán limpiando los datos para el proyecto Liberalismo y Medioambiente para la fundación Republica Liberal
#
# Se irán agregando los enlaces a los datasets de los datos correspondientes.
#
# [EPI](https://epi.yale.edu/downloads)
#
# [Index of Economic Freedom - 2020](https://www.heritage.org/index/explore?u=637340655378618037)
#
# [Index of Economic Freedom - 1995/2020](https://www.heritage.org/index/explore?view=by-region-country-year&u=637340656550516325)
#
#
# + [markdown] id="rMy4Hdp9FRNV" colab_type="text"
# ## Configuramos Google Colab y librerias
# + [markdown] id="iV01F8qgLYSi" colab_type="text"
# El siguiente chunk de codigo sirve para ingresar a mi unidad virtual de Drive para trabajar con los datasets descargados de los enlaces anteriores.
# + id="phKoc0L76Dy8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="6c4f6af4-3924-48af-f091-f4f1e950ab17"
from google.colab import drive
drive.mount('/content/drive/')
# + id="7Ps6nvIfEl24" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0022ddda-2f71-4693-f6e2-9ff3e16cd042"
# ls '/content/drive/My Drive/Colab Notebooks/republica-liberal/data/'
# + id="opT9UwUGFFsu" colab_type="code" colab={}
import pandas as pd
import numpy as np
# + [markdown] id="1SSGkmorFWlp" colab_type="text"
# ## Conociendo Enviroment performance index
#
# Leemos nuestro dataset, lo guardamos en una variable para luego reconocer las labels que tiene el dataframe.
# + id="W3Db53kfRUl8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="d7d934d6-46fc-4f8a-a46c-cd89e6acc34d"
# ls '/content/drive/My Drive/Colab Notebooks/republica-liberal/data/epi'
# + id="rtAGoNf9E3tS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="909cbd55-a7af-43bf-8196-3e0acaadef6a"
# cd '/content/drive/My Drive/Colab Notebooks/republica-liberal/data/epi'
# + [markdown] id="alCEwwl0Jji-" colab_type="text"
# Veamos que tiene el dataset **epi2020results...**
# + id="CtTF2MVDFLew" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 273} outputId="203a7b84-842e-49ca-b44a-fdf36c3328f2"
df_epi = pd.read_csv('epi2020results20200604.csv')
df_epi.head(5)
# + [markdown] id="2g3zpwTdJvOQ" colab_type="text"
# Conozcamos la dimensión del EPI.
# + id="xVZWSX6hJrRB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a69b2054-7916-4988-a766-895baba237c9"
df_epi.shape
# + [markdown] id="1P1_wREYKNI-" colab_type="text"
# Conozcamos el otro dataset, el mismo nos dice que tenemos las regiones, por el titulo, **epi2020resultsregions**
# + id="nUSq_ZnzFutn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 412} outputId="69ec28a6-9ab7-476f-a63d-cd50836a696e"
df_epi_region = pd.read_csv('epi2020resultsregions20200604.csv')
df_epi_region.head(5)
# + id="RIQ_cB_FJaAf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d05cc60a-8459-44f3-ce0a-c529f12dbca1"
df_epi_region.shape
# + [markdown] id="uTgzFmd2Kf16" colab_type="text"
# Tiene 180 filas y 142 columnas.
# + [markdown] id="dcbnxLCBAeKB" colab_type="text"
# Obtengamos una lista de todas las columnas que tiene nuestro dataset.
# + id="x9FRH4520j5J" colab_type="code" colab={}
list(df_epi_region.columns)
# + [markdown] id="21r8RjoSAyWz" colab_type="text"
# Vemos que son muchas columnas. Vamos a buscar que significa cada abreviación.
# + [markdown] id="1rC6E1W8CRKJ" colab_type="text"
# En la siguiente tabla se puede ver mas información acerca de los indicadores anteriores.
#
# La tabla se encuentra en un pdf en [EPI Downloads](https://epi.yale.edu/downloads).
#
# La misma fue obtenida del siguiente enlace:
# [Technical Appendix](https://epi.yale.edu/downloads/epi2020technicalappendix20200803.pdf)
#
# 
#
# El [índice de rendimiento ambiental o índice de desempeño ambiental](https://es.wikipedia.org/wiki/%C3%8Dndice_de_desempe%C3%B1o_ambiental) (Inglés: Environmental Performance Index, siglas EPI) es un método para cuantificar y clasificar numéricamente el desempeño ambiental de las políticas de un país.
#
# Se puede observar 2 grandes indicadores que contienen a un total de 11 y que estos ultimos contienen un total de 32.
#
# Los 2 indicadores que contienen a los demas son:
#
# -----------------------------
#
# 1. **El Environmental Health (HLT) tiene un 40% de peso en el Environment Performance Index (EPI).**
#
# Este indicador nos cuantifica la salud ambiental de un pais. La salud ambiental nos dice los factores en el medio ambiente que pueden potencialmente afectar adversamente la salud de presentes y futuras generaciones.
#
# Los indicadores que afectan a este son:
# - La [calidad de aire](https://es.wikipedia.org/wiki/Calidad_del_aire) (Air Quality). Peso de 50%.
# - El [saneamiento básico](https://es.wikipedia.org/wiki/Saneamiento_ambiental) y el [agua potable](https://es.wikipedia.org/wiki/Agua_potable) (Sanitation and Dringking water). Peso de 40%.
# - Metales pesados (Heavy Metals). Peso de 5%. ([Lead Exposure](https://en.wikipedia.org/wiki/Lead_poisoning))
# - [Gestion de residuos](https://es.wikipedia.org/wiki/Gesti%C3%B3n_de_residuos) (Waste Management). Peso de 5%.
#
# ---------------------
#
# 2. **El Ecosystem Vitality (ECO) tiene un 60% de peso en el Environment Performance Index (EPI).**
#
# La vitalidad del [ecosistema](https://es.wikipedia.org/wiki/Ecosistema) se refiere a la actividad o energía para vivir o desarrollarse del mismo. Si tiene una baja vitalidad, es un indicador que el pais no es amigable con su ecosistema.
#
# Los indicadores que afectan a ese son:
#
# - [Biodiversidad](https://es.wikipedia.org/wiki/Biodiversidad) y [habitat](https://es.wikipedia.org/wiki/H%C3%A1bitat) (Biodiversity & Habitat). Peso de 25%.
# - [Servicios del ecosistema](https://es.wikipedia.org/wiki/Servicios_del_ecosistema) (Ecosystem Services). Peso de 10%.
# - Pesca ([Fisheries](https://en.wikipedia.org/wiki/Fishery)). Peso de 10%.
# - [Cambio climatico](https://es.wikipedia.org/wiki/Cambio_clim%C3%A1tico) (Climate Change). Peso de 40%.
# - Emisiones contaminantes ([Pollution Emissions](https://en.wikipedia.org/wiki/Air_pollution)). Peso de 5%.
# - [Agricultura](https://es.wikipedia.org/wiki/Agricultura) (Agriculture). Peso de 5%.
# - [Recursos hídricos](https://es.wikipedia.org/wiki/Recurso_h%C3%ADdrico). (Water Resources). Peso de 5%.
#
# -----------------
#
# Por lo tanto para calcular el EPI se utiliza una [media ponderada](https://es.wikipedia.org/wiki/Media_ponderada) de los 2 indicadores principales. A su vez se calcula una media ponderada con los indicadores internos de ambos.
# + id="NJdTKw_Y7SR-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 424} outputId="2ed2e18a-862d-4100-833f-c76c684e0439"
df_epi_region[['country', 'region', 'EPI.rgn.rank', 'EPI.rgn.mean', 'EPI.new', 'HLT.new', 'ECO.new']]
# + id="PTP9FF6IKemt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 412} outputId="0e339602-12e5-467b-fa83-61bd9f552163"
df_epi_group = pd.read_csv('epi2020resultsgroups20200604.csv')
df_epi_group.head(5)
# + [markdown] id="HpfZFdULLQrE" colab_type="text"
# Aparentemente se ven columnas adicionales, utilizadas para obtener diferentes indices. No lo tomares a este dataframe.
# + id="4pERhUa6K9NM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 348} outputId="1101be86-c4ce-41f4-f183-2def984a4528"
df_epi_attr = pd.read_csv('epi2020countryattributes20200604.csv')
df_epi_attr.head(5)
# + id="cOSwY2uQLoEt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="cf7b3d39-f1bd-4c1a-9841-e54d254284e9"
df_epi_attr.shape
# + [markdown] id="c735OIR3L53P" colab_type="text"
# Se puede observar que este dataset se encuentra dentro del dataset **epi2020resultsgroups**, asique no lo utilizaremos.
# + id="0iLsLOVOLvz9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 363} outputId="b6eb0377-2697-4cf6-e74f-9e1f8acd4f6e"
df_epi_weights = pd.read_csv('epi2020weights20200604.csv')
df_epi_weights.head(10)
# + id="25Tb_F2wMLOC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="326f4194-b4ed-4113-8274-8d5b501e0d63"
df_epi_weights.shape
# + [markdown] id="iiyv1wmjQvzl" colab_type="text"
# ## Conociendo Human Freedom Index (HFI)
#
# + id="NPpxBRHmRI_R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="cafeac48-6756-4000-967a-e76da5616b9e"
# !ls '/content/drive/My Drive/Colab Notebooks/republica-liberal/data/index_economic_freedom/data-2020'
# + id="nEAcEmbBRc01" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="c6469418-db42-4b53-9ad8-35623840b98d"
# !ls '/content/drive/My Drive/Colab Notebooks/republica-liberal/data/index_economic_freedom/'
# + [markdown] id="8w90O4kxLyx1" colab_type="text"
# Ahora estudiemos un dataset de todos los que hay para saber el formato de los mismos.
# + id="nPaRjrBP318I" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3855b421-6dea-435c-b0df-9177fd1bc8e2"
# cd '/content/drive/My Drive/Colab Notebooks/republica-liberal/data/index_economic_freedom/'
# + id="B31itwxqLyKm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 181} outputId="bd31d051-019a-44c7-dd89-5bb3404f65a5"
df_america = pd.read_csv('data-america.csv')
df_america.head(3)
# + id="hdhcSa-NTD8C" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="718c6b63-65ec-4bbd-bc53-d1e675d3f907"
df_america.shape
# + [markdown] id="SAYTuG4wTLVK" colab_type="text"
# Podemos observar que el dataframe tiene 832 filas y 15 columnas. Veamos las columnas:
# + id="w3IrtR4gTWR4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="9769fb9d-4c4f-4050-e972-352e4a24c592"
df_america.columns
# + [markdown] id="pguQtyMZ4WHC" colab_type="text"
# Estudiaremos los indicadores utilizados en el indice de Libertad Economica. [Indice de Libertad Economica - Español](https://es.wikipedia.org/wiki/%C3%8Dndice_de_Libertad_Econ%C3%B3mica), [Index of Economic Freedom- English](https://en.wikipedia.org/wiki/Index_of_Economic_Freedom#Investment_freedom)
#
# ¿Que significa cada columna?
#
# En el siguiente informe, [2020 Index of Economic Freedom HIGHLIGHTS](https://www.heritage.org/index/pdf/2020/book/2020_IndexofEconomicFreedom_Highlights.pdf), se divide en 4 partes las 12 columnas desde Proporty Rights hasta Financial Freedom. A estas 12 columnas se las llama, **THE 12 ECONOMIC FREEDOMS** o **LAS 12 LIBERTADES ECONOMICAS**. Estudiemos un poco cada una de ellas. No se agregará información al documento, habrán enlaces que nos redirigen a información oficial.
#
# ----------
#
# - [Rule of law](https://www.heritage.org/index/rule-of-law) - [Imperio de la ley](https://es.wikipedia.org/wiki/Imperio_de_la_ley)
#
# 1. [Property Rights](https://www.heritage.org/index/property-rights) - Derechos de propiedad
# 2. Judicial Effectiveness - Efectividad judicial
# 3. [Government Integrity](https://www.heritage.org/index/freedom-from-corruption) - Integridad del gobierno
#
# ----------
#
# - [Government Size](https://www.heritage.org/index/limited-government) - Tamaño del [Gobierno](https://es.wikipedia.org/wiki/Gobierno) ([Big Government](https://en.wikipedia.org/wiki/Big_government) or [Small Government](https://en.wikipedia.org/wiki/Small_government))
#
# 4. [Tax Burden](https://www.heritage.org/index/fiscal-freedom) - Carga fiscal
# 5. [Government Spending](https://www.heritage.org/index/government-spending) - Gastos gubernamentales
# 6. Fiscal Health - <NAME>
#
# ----------
#
# - [Regulatory Efficiency](https://www.heritage.org/index/regulatory-efficiency) - Eficiencia Regulatoria
#
# 7. [Business Freedom](https://www.heritage.org/index/business-freedom) - Libertad comercial
# 8. [Labor Freedom](https://www.heritage.org/index/labor-freedom) - Libertad labotal
# 9. [Monetary Freedom](https://www.heritage.org/index/monetary-freedom) - Libertad monetaria
#
# ----------
#
# - [Open Markets](https://www.heritage.org/index/open-markets) - Apertura de mercados
#
# 10. [Trade Freedom](https://www.heritage.org/index/trade-freedom.aspx) - [Libertad comercial](https://es.wikipedia.org/wiki/Libre_comercio)
# 11. [Investment Freedom](https://www.heritage.org/index/investment-freedom) - Libertad de inversión.
# 12. [Financial Freedom](https://www.heritage.org/index/financial-freedom) - Libertad financiera
#
# ----------
# + [markdown] id="TZo1eZO0TyXh" colab_type="text"
# Veamos algo de información del dataframe
# + id="f2XncyGSTxax" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 399} outputId="cbf2ec12-489d-46fe-e1f0-f95e38585c5a"
df_america.info()
# + [markdown] id="-zHtOUAqcOVS" colab_type="text"
# Agregamos una nueva columna para identificar la region de este dataframe. La misma nos ayudará a futuros analisis por regiones.
# + id="BcoQV8VCa4_R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 531} outputId="9edc053f-b470-41f4-d695-475a79d70902"
df_america['Region'] = 'America'
df_america
# + id="IqQMaD6jJt_1" colab_type="code" colab={}
df_america['Index Year'] = pd.Categorical(df_america['Index Year'].apply(str))
# + [markdown] id="kBIhlTRfg4mT" colab_type="text"
# ### Generalizando y uniendo dataframes.
#
# Sabiendo cual es la estructura de uno de los dataset podemos generalizar a todos los demas, ya que fueron descargados de la misma fuente y de igual forma. Lo que haremos es agregar la region a cada dataframe, unirlos y luego eliminar las columnas que no se utilizarán para este estudio. Las columnas que se eliminarán son los indicadores utilizados para obtener el indice.
#
# > Nota: El indice HFI es una media de estos indicadores.
#
# + [markdown] id="z1ItEDSBiHRK" colab_type="text"
# Crearemos una función para obtener el nombre de los archivos de las carpetas donde se encuentran contenidos los datasets.
# + id="Iws7m-QVgnVY" colab_type="code" colab={}
from os import scandir, getcwd
def ls(ruta): # getcwd()
'''Funcion que retorna una lista con el nombre de los archivos de la ruta que le pasemos
Param:
ruta : str, string de la ruta del directorio. getcwd(): para la ruta actual
'''
return [arch.name for arch in scandir(ruta) if arch.is_file()]
# + id="AYNBtiD8R1Xg" colab_type="code" colab={}
path_data_2020 = '/content/drive/My Drive/Colab Notebooks/republica-liberal/data/index_economic_freedom/data-2020'
path_data = '/content/drive/My Drive/Colab Notebooks/republica-liberal/data/index_economic_freedom/'
# + [markdown] id="SBR9A5bQSt8J" colab_type="text"
# Lista con los nombres de los archivos del año 2020 y del historico de 1995 a 2020.
# + id="boK8xeHQibQE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="db3f2d28-3f34-493a-d119-9a82ff08bc6b"
nombres_datasets_hfi_2020 = ls(path_data_2020)
nombres_datasets_hfi_2020
# + id="XXa2nW_YSIu8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="7a028d0a-f8a7-4be0-e08f-c206a9da777c"
nombres_datasets_hfi = ls(path_data)
nombres_datasets_hfi
# + [markdown] id="rFAGWgdjiS-n" colab_type="text"
# #### Año 2020
#
# Justamos todos los dataframes del año 2020 en uno, tambien le vamos a agregar una columna nueva llamada *Region*. El nombre de la region lo obtenemos del titulo de los csv's.
# + id="MeOlWOzWe7OV" colab_type="code" colab={}
import os
# + colab_type="code" id="LF32_rCXiqOq" colab={}
os.chdir(path_data_2020)
# + colab_type="code" id="nXl1KcWDiqOc" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="9d6ea67e-c64a-46e4-868a-50f6d26b0edf"
os.getcwd()
# + id="_lCoIg2U3iw7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="54f36d69-c6d8-43ef-aeb2-98db1bf89c26"
import re
pattern = 'data-([\w\W]+)-2020.csv'
lista_de_datasets = []
regiones = []
for data_set in nombres_datasets_hfi_2020:
region = re.findall(pattern, data_set)
df = pd.read_csv(data_set)
df['Region'] = region[0]
df['Index Year'] = pd.Categorical(df['Index Year'].apply(str))
lista_de_datasets.append(df)
df_hfi_2020 = pd.concat(lista_de_datasets)
columnas = range(3, len(df_hfi_2020.columns) - 1)
df_hfi_2020 = df_hfi_2020.drop(df_hfi_2020.columns[columnas], axis=1)
df_hfi_2020.head(3)
# + [markdown] id="PmFuonSni7lV" colab_type="text"
# #### Historico 1995-2020
#
# Justamos todos los dataframes desde 1995 al año 2020 en uno, tambien le vamos a agregar una columna nueva llamada *Region*. El nombre de la region lo obtenemos del titulo de los csv's.
# + id="QO1TlmzwfJiS" colab_type="code" colab={}
os.chdir(path_data)
# + id="KNi64YL7fG4H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="b0b9e1d2-83d9-40d8-c9bf-ab006483af7d"
os.getcwd()
# + id="aXe8-c-fVkM6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="d22a42a5-f8fe-4bb5-fe61-92edbcd93cff"
import re
pattern = 'data-([\w\W]+).csv'
lista_de_datasets = []
regiones = []
for data_set in nombres_datasets_hfi:
region = re.findall(pattern, data_set)
df = pd.read_csv(data_set)
df['Region'] = region[0]
df['Index Year'] = pd.Categorical(df['Index Year'].apply(str))
lista_de_datasets.append(df)
df_hfi = pd.concat(lista_de_datasets)
columnas = range(3, len(df_hfi.columns) - 1)
df_hfi = df_hfi.drop(df_hfi.columns[columnas], axis=1)
df_hfi
# + id="CDs06aXxRM4M" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="d9b0a909-a56a-473b-bb34-93f62ba2201d"
df_hfi.info()
# + [markdown] id="AQG_msDbLuaU" colab_type="text"
# # Selección de datasets. Limpiando datasets.
#
# Utilizaremos HFI 2020, HFI 1995-2020, EPI 1950 - 2020.
# + [markdown] id="iW_2C2YVkEH7" colab_type="text"
# ## HFI 1995-2020
# + id="5qlhoieTzNkM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 208} outputId="847b70d0-cd01-41e3-c575-c98715225421"
df_hfi.info()
# + id="xBjDDL5Xm9w_" colab_type="code" colab={}
# + [markdown] id="3HGDqgqrj80L" colab_type="text"
# ## HFI 2020
# + id="WNZvorgzzTmc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 208} outputId="3ab2b9ba-bd59-483d-b1fc-c08ff6e552d1"
df_hfi_2020.info()
# + [markdown] id="BEIQXEHHkIJM" colab_type="text"
# ## EPI por Region
# + [markdown] id="NFp6tXwE07f-" colab_type="text"
# Utilizaremos el dataframe epi, con la columna adicional de las regiones.
# + id="vbnQvaWuMXUi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 104} outputId="5b9e35ff-6104-4a39-82b8-680f724c6e2a"
df_epi_region.info()
# + id="JB9Wwq8piWKM" colab_type="code" colab={}
df_epi_clean = df_epi_region.drop(['code', 'iso'], axis = 1)
# + [markdown] id="lhoDY65wpb81" colab_type="text"
# Borraremos todas las columnas y solo dejaremos las del indice EPI y sus subindices ECO y HLT
# + id="obl5QSegqsZr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="95708f21-01b9-4a86-d941-26526233e007"
list(df_epi_clean.columns)
# + id="A3iyQNU56bUd" colab_type="code" colab={}
columns_selected = ['country','region', 'EPI.new', 'ECO.new', 'HLT.new']
# + id="eEJyn_Xa73WX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="9c6e2e7b-f19c-4bc5-dab2-293285230858"
df_epi_clean = df_epi_clean[columns_selected]
df_epi_clean.head(5)
# + [markdown] id="aQTPLHj3jKF8" colab_type="text"
# # Unificando datasets
# + [markdown] id="FxVGBRYrjR1-" colab_type="text"
# ## Unificando datos 2020
# + [markdown] id="foniEgpfbjk1" colab_type="text"
# Vamos a hacer una unificacion de Regiones de ambos datasets. Veamos las regiones.
# + id="hVks4VHyBDiq" colab_type="code" colab={}
df_epi_clean = df_epi_clean.rename(columns={'EPI.new':'EPI 2020', 'ECO.new': 'ECO 2020', 'HLT.new': 'HLT 2020'})
# + id="b4uRC9fKeXoS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="16ff30fb-0569-4119-92d1-072c51faf9e6"
df_epi_clean.columns
# + id="7CCNKUVIgrfM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="35864865-9e34-4739-d71a-9b0de3eb9176"
df_epi_clean.info()
# + id="vwO0WChJehFC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d180913f-af54-4441-d08a-efb53861d7f7"
df_hfi_2020.columns
# + [markdown] id="rbxtHHVSrpn9" colab_type="text"
# ### Completando datos nulos
# + [markdown] id="sT11yoFt1PpM" colab_type="text"
# A ese diccionario llegamos porque se descubrio cuando se unieron los datasets y habian 36 valores nulos. Se creo una lista con los nombres de los paises nulos, con esa lista creamos un diccionario y modificamos los nombres que tenian espacios inadecuados.
#
#
# El codigo que utilizamos para descubrir y armar el diccionario de abajo es el siguiente:
#
# ```py
# # Merge de datasets
# df_hfi_epi_union = pd.merge(df_hfi_2020, df_epi_clean, how = 'outer', left_on='Name', right_on='country')
#
# # Analisis de datos nulos
# df_hfi_epi_union_null = df_hfi_epi_union[df_hfi_epi_union.country.isnull()]
# list(df_hfi_epi_union_null['Name'].unique())
#
# df_hfi_epi_union_null_2 = df_hfi_epi_union[df_hfi_epi_union.Name.isnull()]
# list(df_hfi_epi_union_null_2['country'].unique())
# ```
#
#
# + id="bQptU72uzB-K" colab_type="code" colab={}
# Diccionario para eliminar espacios al final del valor
dict_para_hfi_Name = {
'<NAME> ' : '<NAME>',
'El Salvador ' : 'El Salvador',
'Guatemala ' : 'Guatemala',
'Honduras ' : 'Honduras',
'Jamaica ' : 'Jamaica',
'Nicaragua ' : 'Nicaragua',
'Panama ' : 'Panama',
'Paraguay ' : 'Paraguay',
'The Bahamas' : 'The Bahamas',
'United States' : 'United States',
'Uruguay ' : 'Uruguay',
'Venezuela ' : 'Venezuela',
'Bangladesh ' : 'Bangladesh',
'Burma' : 'Burma',
'Hong Kong' : 'Hong Kong',
'Kyrgyz Republic ' : 'Kyrgyz Republic',
'Macau' : 'Macau',
'Malaysia ' : 'Malaysia',
'North Korea' : 'North Korea',
'Pakistan ' : 'Pakistan',
'Thailand ' : 'Thailand',
'Vietnam' : 'Vietnam',
'Libya' : 'Libya',
'Syria' : 'Syria',
'Yemen' : 'Yemen',
'Hungary ' : 'Hungary',
'Kosovo' : 'Kosovo',
'Liechtenstein' : 'Liechtenstein',
'Serbia ' : 'Serbia',
"Côte d'Ivoire " : "Côte d'Ivoire",
'Democratic Republic of Congo': 'Democratic Republic of Congo',
'Mozambique ' : 'Mozambique',
'Republic of Congo ' : 'Republic of Congo',
'São Tomé and Príncipe ' : 'São Tomé and Príncipe',
'Somalia' : 'Somalia',
'The Gambia' : 'The Gambia'
}
# + id="TWu60vv_y0UF" colab_type="code" colab={}
# Seteo como indice la columna name, renombro esas columnas y luego reseteo la columna a su estado original
df_hfi_2020 = df_hfi_2020.set_index(['Name']).rename(index = dict_para_hfi_Name)
df_hfi_2020 = df_hfi_2020.reset_index()
# + id="RmauD53Bto-t" colab_type="code" colab={}
df_hfi_epi_union = pd.merge(df_hfi_2020, df_epi_clean, how = 'outer', left_on='Name', right_on='country')
# + id="TAIwq61n1CeQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="381b5093-9956-4b11-c3ba-b89bd646d707"
df_hfi_epi_union.info()
# + [markdown] id="dScCtqtO7Vak" colab_type="text"
# El mismo procedimiento que antes lo hago ahora, pero para los paises del epi (country de epi)
#
#
# El codigo usado fue:
#
# ```py
# # Analisis de datos nulos. Name de hfi
# df_hfi_epi_union_null = df_hfi_epi_union[df_hfi_epi_union.country.isnull()]
# list(df_hfi_epi_union_null['Name'].unique())
#
# # Analisis de datos nulos. country de epi
# df_hfi_epi_union_null_2 = df_hfi_epi_union[df_hfi_epi_union.Name.isnull()]
# list(df_hfi_epi_union_null_2['country'].unique())
# ```
#
#
# + id="6Nb8_OSh400j" colab_type="code" colab={}
dict_para_epi_country = {
'Antigua and Barbuda' : 'Antigua and Barbuda',
'Bahamas' : 'The Bahamas',
"Cote d'Ivoire" : "Côte d'Ivoire",
'Dem. Rep. Congo' : 'Democratic Republic of Congo',
'Gambia' : 'The Gambia',
'Grenada' : 'Grenada',
'Kyrgyzstan' : 'Kyrgyz Republic',
'Marshall Islands' : 'Marshall Islands',
'Myanmar' : 'Burma',
'Sao Tome and Principe' : 'São Tomé and Príncipe',
'United States of America' : 'United States',
'Viet Nam' : 'Vietnam'
}
# + id="dwIXiL4s7tD7" colab_type="code" colab={}
df_epi_clean = df_epi_clean.set_index(['country']).rename(index = dict_para_epi_country)
df_epi_clean = df_epi_clean.reset_index()
# + id="YYaI4G2g76bl" colab_type="code" colab={}
df_hfi_epi_union = pd.merge(df_hfi_2020, df_epi_clean, how = 'outer', left_on='Name', right_on='country')
# + id="g1p6YF0n77_e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="2b399323-392b-470c-ebf9-7cd9e6630e8a"
df_hfi_epi_union.info()
# + [markdown] id="-diCrqcPrWvC" colab_type="text"
# ### Cambiamos los nombres a las columnas.
#
# Cambiamos el nombre de algunas columnas.
# + id="IuX-fjrLE_yu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="aa2ed1cd-49ad-46d0-fdee-ef48eb65e7ef"
list(df_hfi_epi_union.columns)
# + id="KpPod_jDFIMB" colab_type="code" colab={}
dict_columnas = {
'Name' :'Name',
'Index Year' :'year',
'Overall Score':'human_freedom_index',
'Region' :'Region',
'country' :'country',
'region' :'region',
'EPI 2020' :'environmetal_performance_index',
'ECO 2020' :'ecosystem_vitality',
'HLT 2020' :'environmental_health'
}
# + id="9pDBrIFJGcBZ" colab_type="code" colab={}
df_hfi_epi_union = df_hfi_epi_union.rename(columns=dict_columnas)
# + [markdown] id="ZVl1pE2Nra5t" colab_type="text"
# ### Analizamos datos nulos
#
# Analizamos datos nulos que no pudimos completar.
# + id="doq6mRLk8G7J" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="27eae5f3-87a3-42b8-84bc-ec7309495abe"
# Analisis de datos nulos. Name de hfi
df_hfi_epi_union_null = df_hfi_epi_union[df_hfi_epi_union.country.isnull()]
list(df_hfi_epi_union_null['Name'].unique())
# + id="hORJoiEb8JgE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="36e75150-a321-4a05-aa21-52c55b8e03a5"
# Analisis de datos nulos. country de epi
df_hfi_epi_union_null_2 = df_hfi_epi_union[df_hfi_epi_union.Name.isnull()]
list(df_hfi_epi_union_null_2['country'].unique())
# + [markdown] id="zJPVko5HDVH2" colab_type="text"
# Los siguientes paises no tienen datos en las columnas que se observan. Se filtrara un dataset unido sin estos datos.
#
# Luego a modo de IDEA... se podria calcular los valores nulos de estos paises haciendo una regresin lineal con los datos que se saben. El valor que obtendremos será una aproximación con un grado de confiabilidad mayor o menor en funcion del modelo que se obtenda.
# + id="kes5NBaX8bKs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="d092acfa-9ff6-4f8c-e215-99d24326894b"
df_hfi_epi_union_nulls = df_hfi_epi_union[df_hfi_epi_union.Name.isnull()]
df_hfi_epi_union_nulls
# + id="f8Vs2LvS-1K9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 595} outputId="f81dfa51-9de8-42ce-c5f8-9d8c5ac07680"
df_hfi_epi_union_nulls_2 = df_hfi_epi_union[df_hfi_epi_union.country.isnull()]
df_hfi_epi_union_nulls_2
# + [markdown] id="rNTW2gPMoO9h" colab_type="text"
# Creamos un dataframe con datos nulos, el cual nos puede servir para la propuesta que hicimos mas arriba.
#
# Primero concatenamos los datasets
# + id="0jHUJ0OrnhmW" colab_type="code" colab={}
df_hfi_epi_with_nulls = pd.concat([df_hfi_epi_union_nulls, df_hfi_epi_union_nulls_2]).reset_index().drop(columns=['index'])
# + id="vcexk_n-oUtm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 810} outputId="731ffa27-98ab-4122-d54c-41acd334493b"
df_hfi_epi_with_nulls
# + [markdown] id="gqVRsXTNs1YK" colab_type="text"
# Haciendo los cambios correspondiente al dataframe con datos nulos.
# + id="EyviocPIpCqh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 124} outputId="4ce2ec28-cfeb-4e76-f13e-5e3f84b8c345"
df_hfi_epi_with_nulls['Name'].iloc[0:3] = ['Antigua and Barbuda', 'Grenada', 'Marshall Islands']
df_hfi_epi_with_nulls['Region'].iloc[0:3] = ['america', 'america', 'asia-pacific']
df_hfi_epi_with_nulls['year'].iloc[0:3] = '2020'
df_hfi_epi_with_nulls = df_hfi_epi_with_nulls.drop(columns=['country', 'region'])
df_hfi_epi_with_nulls = df_hfi_epi_with_nulls.rename(columns={'Name': 'country', 'Region':'region'})
df_hfi_epi_with_nulls = df_hfi_epi_with_nulls.reindex(columns=['country',
'region',
'year',
'human_freedom_index',
'environmetal_performance_index',
'ecosystem_vitality',
'environmental_health'])
# + [markdown] id="2tpI5NU3tQkY" colab_type="text"
# El siguiente dataframe ya esta adecuado para exportarlo.
# + id="4ABdZEYCsae3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 724} outputId="19a969a9-0bb2-4d4d-bc2e-65c7ed543f87"
df_hfi_epi_with_nulls
# + [markdown] id="qMliiDhttEUa" colab_type="text"
# ### Analizandos datos NO nulos
#
# Reordenamos las columnas y lo adecuamos para exportarlo mas adelante.
# + id="mpNccdKlEDll" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="12c1e289-2301-492b-b04c-6cb8700f4cec"
df_hfi_epi_notnulls = df_hfi_epi_union[df_hfi_epi_union.country.notnull()]
df_hfi_epi_notnulls = df_hfi_epi_notnulls[df_hfi_epi_notnulls.Name.notnull()]
df_hfi_epi_notnulls.info()
# + id="u5fWz7YaisUf" colab_type="code" colab={}
df_hfi_epi_notnulls = df_hfi_epi_notnulls.drop(columns=['Name', 'region'])
# + id="vWSFRQk3jpsk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 139} outputId="956535da-1efe-4b73-9110-39c9492fe3d6"
df_hfi_epi_notnulls.columns.tolist()
# + id="3DWWfiRuj2qC" colab_type="code" colab={}
df_hfi_epi_notnulls = df_hfi_epi_notnulls.reindex(columns=
['country',
'Region',
'year',
'human_freedom_index',
'environmetal_performance_index',
'ecosystem_vitality',
'environmental_health'])
df_hfi_epi_notnulls = df_hfi_epi_notnulls.rename(columns={'Region': 'region'})
# + [markdown] id="MRrZIcHDk9PI" colab_type="text"
# # Exportando datasets
# + id="MQl6nVDVmB7Z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="bb6aeef3-7ac4-4ccb-f4d1-502dfb2f5d3b"
import os
os.getcwd()
# + id="Ch_xcBHlmrPe" colab_type="code" colab={}
os.chdir('/content/drive/My Drive/Colab Notebooks/republica-liberal/data/data_to_work')
# + id="4aqhZNwtmurM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="c205c01e-aaa2-4707-9de8-5a33e93c2353"
os.getcwd()
# + id="lpASUhQAlA76" colab_type="code" colab={}
df_hfi_epi_notnulls.to_csv('dataset_hfi_epi_2020.csv')
df_hfi_epi_with_nulls.to_csv('dataset_hfi_epi_with_nulls_2020.csv')
# + [markdown] id="kynLV6vmtxC_" colab_type="text"
# > NOTA: Los datos historicos se limpiarán en otro notebook.
| Notebooks/1_Extraccion_y_limpieza_de_datos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# ### **Important:**
# As in the last notebook, in the next cell you need to update the `your_username` variable with **Your Username** (between the single quotes). After you have done that, in principle, you should be able to run the notebook all at once instead of cell by cell.
#Please enter your SciServer username between the single quotes below!
your_username = ''
# import matplotlib to show plots inline.
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import glob
import os
# Import the sims_maf modules needed.
# import maf python modules
import lsst.sims.maf.db as db
import lsst.sims.maf.metrics as metrics
import lsst.sims.maf.slicers as slicers
import lsst.sims.maf.stackers as stackers
import lsst.sims.maf.plots as plots
import lsst.sims.maf.metricBundles as metricBundles
# add opsimUtils module path to search
import sys
sys.path.insert(0, '../Scripts_NBs/')
# import convenience functions
from opsimUtils import *
# +
if your_username == '': # do NOT put your username here
raise Exception('Please provide your username! See the top of the notebook.')
# user provided paths
resultDbPath = '/home/idies/workspace/Storage/{}/persistent/MAFOutput/DDF/'.format(your_username)
metricDataPath = '/home/idies/workspace/Storage/{}/persistent/MAFOutput/DDF/MetricData/'.format(your_username)
# +
# get a dictionary of resultDb from given directory
resultDbs = getResultsDbs(resultDbPath)
# the following line will be useful if you did not run MAF on all 75 opsims
runNames = list(resultDbs.keys())
# +
# retrieve metricBundles for each opsim run and store them in a dictionary
bundleDicts = {}
for runName in resultDbs:
bundleDicts[runName] = bundleDictFromDisk(resultDbs[runName], runName, metricDataPath)
# -
# #### **Note:**
# The `metricId` for each metric could vary from opsim to opsim (due to the inconsistency of the proposalId assginment across opsims).
# we see the available metric keys
list(bundleDicts[runNames[0]].keys())
help(getSummary)
getSummary(resultDbs, 'coadd_COSMOS_g', 'Median', pandas=True)
# ### 2. Visualize Coadd Metric
plotSummaryBar(resultDbs, 'coadd_COSMOS_g', 'Median', slicerName = 'HealpixSlicer', axhline=28.7)
plotHist(bundleDicts, (1, 'coadd_COSMOS_g'), axvline=28.7)
# +
# to get exact metricKey for each metric across different opsims
metricName = 'coadd_COSMOS_g'
summary = getSummary(resultDbs, metricName, 'Median', pandas=True)
# loop over all opsims and make plots
for run in runNames:
row = summary[summary.runName == run]
metricKey = (int(row.metricId), metricName)
plotSky_DDF(bundleDicts[run][metricKey], 'COSMOS')
# -
# ### 3. Visualize Nvisit Metric
# The following basically repeats what were done for the coadd metric.
getSummary(resultDbs, 'nvisit_COSMOS_g', 'Median', pandas=True)
plotSummaryBar(resultDbs, 'nvisit_COSMOS_g', 'Median', \
slicerName = 'HealpixSlicer', axhline=900)
plotHist(bundleDicts, (5, 'nvisit_COSMOS_g'), axvline=900)
# +
# to get exact metricKey for each metric across different opsims
metricName = 'nvisit_COSMOS_g'
summary = getSummary(resultDbs, metricName, 'Median', pandas=True)
# loop over all opsims and make plots
for run in runNames:
row = summary[summary.runName == run]
metricKey = (int(row.metricId), metricName)
# specifiy logscale for better visualization
plotSky_DDF(bundleDicts[run][metricKey], 'COSMOS', scale=np.log10)
# -
| contrib/03_plotDDF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="XYYDvoskkE61"
import json
import tensorflow as tf
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
# + colab={} colab_type="code" id="0eJSTTYnkJQd"
vocab_size = 10000
embedding_dim = 16
max_length = 100
trunc_type='post'
padding_type='post'
oov_tok = "<OOV>"
training_size = 20000
# + colab={} colab_type="code" id="BQVuQrZNkPn9"
'''
!wget --no-check-certificate \
https://storage.googleapis.com/laurencemoroney-blog.appspot.com/sarcasm.json \
-O /tmp/sarcasm.json
'''
# -
path = tf.keras.utils.get_file('sarcasm.json',
'https://storage.googleapis.com/laurencemoroney-blog.appspot.com/sarcasm.json')
print (path)
# + colab={} colab_type="code" id="oaLaaqhNkUPd"
with open(path, 'r') as f:
datastore = json.load(f)
sentences = []
labels = []
for item in datastore:
sentences.append(item['headline'])
labels.append(item['is_sarcastic'])
# + colab={} colab_type="code" id="S1sD-7v0kYWk"
training_sentences = sentences[0:training_size]
testing_sentences = sentences[training_size:]
training_labels = labels[0:training_size]
testing_labels = labels[training_size:]
# + colab={} colab_type="code" id="3u8UB0MCkZ5N"
tokenizer = Tokenizer(num_words=vocab_size, oov_token=oov_tok)
tokenizer.fit_on_texts(training_sentences)
word_index = tokenizer.word_index
training_sequences = tokenizer.texts_to_sequences(training_sentences)
training_padded = pad_sequences(training_sequences, maxlen=max_length, padding=padding_type, truncating=trunc_type)
testing_sequences = tokenizer.texts_to_sequences(testing_sentences)
testing_padded = pad_sequences(testing_sequences, maxlen=max_length, padding=padding_type, truncating=trunc_type)
# + colab={} colab_type="code" id="GrAlWBKf99Ya"
# Need this block to get it to work with TensorFlow 2.x
import numpy as np
training_padded = np.array(training_padded)
training_labels = np.array(training_labels)
testing_padded = np.array(testing_padded)
testing_labels = np.array(testing_labels)
# + colab={} colab_type="code" id="FufaT4vlkiDE"
model = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size, embedding_dim, input_length=max_length),
tf.keras.layers.GlobalAveragePooling1D(),
tf.keras.layers.Dense(24, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid')
])
model.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
model.summary()
# + colab={} colab_type="code" id="2DTKQFf1kkyc"
num_epochs = 30
history = model.fit(training_padded,
training_labels,
epochs=num_epochs,
validation_data=(testing_padded, testing_labels), verbose=2)
# + colab={} colab_type="code" id="2HYfBKXjkmU8"
import matplotlib.pyplot as plt
def plot_graphs(history, string):
plt.plot(history.history[string])
plt.plot(history.history['val_'+string])
plt.xlabel("Epochs")
plt.ylabel(string)
plt.legend([string, 'val_'+string])
plt.show()
plot_graphs(history, "accuracy")
plot_graphs(history, "loss")
# + colab={} colab_type="code" id="7SBdAZAenvzL"
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
def decode_sentence(text):
return ' '.join([reverse_word_index.get(i, '?') for i in text])
print(decode_sentence(training_padded[0]))
print(training_sentences[2])
print(labels[2])
# + colab={} colab_type="code" id="c9MqihtEkzQ9"
e = model.layers[0]
weights = e.get_weights()[0]
print(weights.shape) # shape: (vocab_size, embedding_dim)
# + colab={} colab_type="code" id="LoBXVffknldU"
import io
out_v = io.open('vecs.tsv', 'w', encoding='utf-8')
out_m = io.open('meta.tsv', 'w', encoding='utf-8')
for word_num in range(1, vocab_size):
word = reverse_word_index[word_num]
embeddings = weights[word_num]
out_m.write(word + "\n")
out_v.write('\t'.join([str(x) for x in embeddings]) + "\n")
out_v.close()
out_m.close()
# + colab={} colab_type="code" id="U4eZ5HtVnnEE"
try:
from google.colab import files
except ImportError:
pass
else:
files.download('vecs.tsv')
files.download('meta.tsv')
# + colab={} colab_type="code" id="cG8-ArY-qDcz"
sentence = ["granny starting to fear spiders in the garden might be real", "game of thrones season finale showing this sunday night"]
sequences = tokenizer.texts_to_sequences(sentence)
padded = pad_sequences(sequences, maxlen=max_length, padding=padding_type, truncating=trunc_type)
print(model.predict(padded))
| 3. NLP/AZ/Text Classification/01_RNN/07_sarcasm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="8Xdmm79IChoP" colab_type="code" colab={}
import pandas as pd
pd.set_option('display.max_columns', None)
# + id="6o9H0nwZCkJk" colab_type="code" colab={}
df = pd.read_csv('https://raw.githubusercontent.com/niravjdn/Software-Measurement-Project/master/data/pit/lang/mutations.csv', error_bad_lines=False, names = ["Class", "Package", "gc1", "gc2","gc3","Coverage","gc4"])
# + id="zrakV8XyeDVn" colab_type="code" outputId="9c250c09-2a74-4096-e7da-213fad876526" executionInfo={"status": "ok", "timestamp": 1554301520692, "user_tz": 240, "elapsed": 7889, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 218}
df.head()
# + id="OHOvDA44XrhY" colab_type="code" outputId="35de6dfa-b271-43a5-9e45-d848564a23f8" executionInfo={"status": "ok", "timestamp": 1554301520694, "user_tz": 240, "elapsed": 7867, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 198}
df.drop('gc1', axis=1, inplace=True)
df.drop('gc2', axis=1, inplace=True)
df.drop('gc3', axis=1, inplace=True)
df.drop('gc4', axis=1, inplace=True)
df['Package'] = df['Package'].map(lambda x: str(x)[:x.rfind('.')])
df['Package'] = df['Package'].map(lambda x: x if (x.find('$')+1 == 0) else x[:x.find('$')+1] )
df['Class'] = df['Class'].map(lambda x: str(x)[:x.rfind('.java')])
df.rename(columns={'Class':'CLASS',
'Package':'PACKAGE'},
inplace=True)
df.head()
# + id="pg8GRqelYLA7" colab_type="code" outputId="226bfe11-4a52-4532-b1be-b2720767d55f" executionInfo={"status": "ok", "timestamp": 1554301520695, "user_tz": 240, "elapsed": 7846, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 228}
df = df.groupby(['CLASS','PACKAGE','Coverage'],as_index = False).size().unstack(fill_value=0)
df.head()
# + id="qNfjmmXIMyU_" colab_type="code" outputId="cf8221da-9385-4b28-d4df-b4fdc029ad6b" executionInfo={"status": "ok", "timestamp": 1554301520696, "user_tz": 240, "elapsed": 7834, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
#Not MEMORY_ERROR
df['Total_Mutant'] = (df['KILLED'] + df['NO_COVERAGE'] + df['SURVIVED'] + df['TIMED_OUT'])
df['Mutation_Score'] = ((df['KILLED']+df['TIMED_OUT']) / df['Total_Mutant'])*100
print('Total Mutants '+str(df.Total_Mutant.sum()))
# + id="ydFROW0wNWrG" colab_type="code" outputId="c16f577e-d9ca-4d4b-9a58-b28ea43d6f0c" executionInfo={"status": "ok", "timestamp": 1554301520697, "user_tz": 240, "elapsed": 7823, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
print('Total Killed '+str(df.KILLED.sum()+df.TIMED_OUT.sum()))
# + id="Uev4ykUbGf8S" colab_type="code" outputId="afb7e141-e0ad-4c0e-ab9e-0d82bdc8e579" executionInfo={"status": "ok", "timestamp": 1554301520699, "user_tz": 240, "elapsed": 7810, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
print('Total TIMED_OUT '+str(df.TIMED_OUT.sum()))
# + id="vxz5EbA3b6ZR" colab_type="code" outputId="df6be2ce-1aa4-4200-b5d4-8ddfd1c4cecd" executionInfo={"status": "ok", "timestamp": 1554301520700, "user_tz": 240, "elapsed": 7799, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
print('Mutation Score '+str((df.KILLED.sum()/df.Total_Mutant.sum())*100))
#df.reset_index()
#df.columns.tolist()
# + id="jLTPzrXncHH5" colab_type="code" outputId="d8186673-ccac-44eb-fc77-8be83e670378" executionInfo={"status": "ok", "timestamp": 1554301520701, "user_tz": 240, "elapsed": 7789, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 198}
df.to_csv('data.csv')
from google.colab import files
#files.download("data.csv")
df = pd.read_csv('data.csv', error_bad_lines=False)
df.head()
# + id="nVKA6_A-B26C" colab_type="code" outputId="0e08bb4f-d6d8-497a-b1d1-eab07f052d7f" executionInfo={"status": "ok", "timestamp": 1554301520702, "user_tz": 240, "elapsed": 7779, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 199}
df.count()
# + id="-_g7xRKJsH0c" colab_type="code" outputId="bb42424b-5c0f-43b5-e4a6-72138f0b2a67" executionInfo={"status": "ok", "timestamp": 1554301520704, "user_tz": 240, "elapsed": 7770, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 90}
df1 = pd.read_csv('https://raw.githubusercontent.com/niravjdn/Software-Measurement-Project/master/data/jacoco/lang/jacoco/jacoco.csv', error_bad_lines=False)
df1['CLASS'] = df1['CLASS'].map(lambda x: x if (x.find('.')+1 == 0) else x[:x.find('.')] )
#aggregation_functions = {'price': 'sum', 'amount': 'sum', 'name': 'first'}
df1 = df1.groupby(df1['CLASS']).aggregate(sum).reset_index()
#df1.count()
df1.columns
#df1.head()
# + id="G0vrwC_5sP5Y" colab_type="code" colab={}
df1['Statement_Percentage'] = (df1['LINE_COVERED'] / (df1['LINE_COVERED'] + df1['LINE_MISSED'])) * 100
# + id="gdXG_P7KsTXV" colab_type="code" colab={}
df1['Branch_Percentage'] = (df1['BRANCH_COVERED'] / (df1['BRANCH_COVERED'] + df1['BRANCH_MISSED'])) * 100
# + id="C84PQmq0sd4W" colab_type="code" outputId="79876f4c-c48f-4251-e629-d16cc54ae5ec" executionInfo={"status": "ok", "timestamp": 1554301520711, "user_tz": 240, "elapsed": 7758, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
df1['CC'] = df1['COMPLEXITY_COVERED'] + df1['COMPLEXITY_MISSED'];
df1.CLASS.count()
# + id="2h0qVG20CsYt" colab_type="code" outputId="0ef786ee-0afe-4c50-f203-7e7950da6d65" executionInfo={"status": "ok", "timestamp": 1554301520712, "user_tz": 240, "elapsed": 7747, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
merged_inner = pd.merge(left=df,right=df1, left_on='CLASS', right_on='CLASS')
merged_inner.CLASS.count()
# + id="b1FphMdmsfwH" colab_type="code" outputId="a4cd8102-bb02-4d13-e156-09350eef6d76" executionInfo={"status": "ok", "timestamp": 1554301520715, "user_tz": 240, "elapsed": 7736, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 248}
df1[(~df1.CLASS.isin(merged_inner.CLASS))&(~df1.CLASS.isin(merged_inner.CLASS))]
# + [markdown] id="zrZLLr_0EsLC" colab_type="text"
# Difference - Classes not covered in jacoco or PIT
# + id="OzRy-Vbx7NEO" colab_type="code" outputId="b5fe1800-3f02-425d-994b-1563e32031f4" executionInfo={"status": "ok", "timestamp": 1554301520716, "user_tz": 240, "elapsed": 7722, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 218}
df = merged_inner
df.columns
merged_inner.head()
# + id="_zF4OIsgHySd" colab_type="code" outputId="31ff0a11-95c1-4b70-e2a9-9d30c5241749" executionInfo={"status": "ok", "timestamp": 1554301520718, "user_tz": 240, "elapsed": 7708, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 435}
merged_inner.count()
# + id="UZWnbViD7tBc" colab_type="code" outputId="c93990d5-9389-4e6e-c18a-ede6fbeb39a3" executionInfo={"status": "ok", "timestamp": 1554301520719, "user_tz": 240, "elapsed": 7698, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 379}
df.plot(x='Mutation_Score', y='Statement_Percentage', style='o')
# + id="Y8NzDmbc731f" colab_type="code" outputId="4483da81-d2ef-4583-8716-cb8c62a0a36d" executionInfo={"status": "ok", "timestamp": 1554301520721, "user_tz": 240, "elapsed": 7689, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 108}
df[['Mutation_Score','Statement_Percentage']].corr(method ='spearman')
# + id="Zm_9VAV18AR5" colab_type="code" outputId="312fb7e6-bbd8-4707-9085-e5fea63b5fd8" executionInfo={"status": "ok", "timestamp": 1554301520722, "user_tz": 240, "elapsed": 7677, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 379}
df.plot(x='Mutation_Score', y='Branch_Percentage', style='o')
# + id="7jZK7_PH8GLg" colab_type="code" outputId="4cbbc937-b055-44a5-8a43-9702028f77f4" executionInfo={"status": "ok", "timestamp": 1554301520723, "user_tz": 240, "elapsed": 7667, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 108}
df[['Mutation_Score','Branch_Percentage']].corr(method ='spearman')
# + id="JPDvezjA-6Zv" colab_type="code" colab={}
# + id="7ulanV_I8K7S" colab_type="code" colab={}
df.to_csv('lang-mu-st-branch.csv')
from google.colab import files
files.download("lang-mu-st-branch.csv")
| Jupyter Notebook/Jupyter Notebok/Mutation - Statement/Lang.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true
import io, sys
import numpy as np
from sklearn.decomposition import PCA
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# + deletable=true editable=true
def load_vectors(filename):
fin = io.open(filename, 'r', encoding='utf-8', newline='\n')
n, d = map(int, fin.readline().split())
data = {}
for line in fin:
tokens = line.rstrip().split(' ')
data[tokens[0]] = np.asarray([float(x) for x in tokens[1:]])
return data
# + deletable=true editable=true
word_vectors = load_vectors('wiki.en.vec')
# + deletable=true editable=true
vectors = []
for vector in word_vectors.values():
vectors.append(vector)
eigenwords = np.stack(vectors,axis=-1)
eigenwords.shape
# + deletable=true editable=true
# delete few words from the eigen words, latter we show how we able to construct them from the eigen modes
eigenwords = np.delete(eigenwords,np.s_[:3],axis=1)
eigenwords.shape
# + deletable=true editable=true
# SVD decomposition
U, Sigma, Vt = np.linalg.svd(eigenwords, full_matrices=0)
# + deletable=true editable=true
# eigen modes
U.shape
# + deletable=true editable=true
#scaling matrix
Sigma.shape
# + deletable=true editable=true
# visualize 2 pca components from U
plt.figure(figsize=(20,5))
pca = PCA(n_components=2)
wv_transformed_U = pca.fit_transform(U)
sns.jointplot(wv_transformed_U[:,0],wv_transformed_U[:,1])
# +
#visualize 2 pca components frpm eigenwords
wv_transformed_eigenwords = pca.fit_transform(eigenwords)
sns.jointplot(wv_transformed_eigenwords[:,0],wv_transformed_eigenwords[:,1])
# + deletable=true editable=true
plt.figure(figsize=(20,5))
wv_transformed_eigenwords = pca.fit_transform(eigenwords)
plt.scatter(wv_transformed_eigenwords[:,0],wv_transformed_eigenwords[:,1],c='k',label='EigenWords')
wv_transformed_U = pca.fit_transform(U)
plt.scatter(wv_transformed_U[:,0],wv_transformed_U[:,1],c='r',marker='^',label='EigenModes')
plt.legend()
# + deletable=true editable=true
# This function computes the cosine similarity between vectors u and v
def cosine(u, v):
cos = np.dot(u, v)/(np.linalg.norm(u) * np.linalg.norm(v))
return cos
# + deletable=true editable=true
# This function returns the word corresponding to the nearest neighbor vector of x
def nearest_neighbor(x, word_vectors):
best_score = -1.0
best_word = None
for word, v in word_vectors.items():
score = cosine(x, v)
if score > best_score:
best_score = score
best_word = word
return best_word
# + deletable=true editable=true
# return the nearest neighbors of eigen modes(U matrix) in word vectors
best_words = []
for col in range(300):
best_words.append(nearest_neighbor(np.transpose(U[0:300,col]),word_vectors))
print('The best nearest words for Eigen modes in word vectors are:')
print(' ')
print(best_words)
# -
#
#
#
# To make sense from the cell above: I don't really know how construct words from vector(there is no vec2word) in order to see what are the most dominant words in U matrix, so insteed I did my best and found the nearset words in words verctors to the dominant words in U.
#
#
#
#
#
#
#
#
# + deletable=true editable=true
# remmeber the words we previously deleted from our eigen words dataset [',' '.' 'the']
# can we construct them usig compressed informations presented by eigen modes (U matrix)?
# let's construct the word '.' which was left out when we compute the svd decomposition
# R is number of modes we are willing to take from eigen modes (U matrix)
R = [10,30,60,90,120,150,180,210,240,270,300]
for r in R:
#alpha contain compressed information about the word
alpha = np.dot(np.transpose(U[0:300,:r]), np.transpose(word_vectors[',']))
#Project alpha on u, so we can what's word is projected is it same word which has been deleted from eigenwords
x_hat = np.dot(U[0:300,:r],alpha)
print('At rank %i the word is :'%(r))
print(nearest_neighbor(x_hat,word_vectors))
print(' ')
# + deletable=true editable=true
#check the bais in the eigenwords using only the compressed information (alpha)
alpha_man = np.dot(np.transpose(U[0:300,10:12]), np.transpose(word_vectors['man']))
alpha_woman = np.dot(np.transpose(U[0:300,10:12]), np.transpose(word_vectors['woman']))
alpha_executive = np.dot(np.transpose(U[0:300,10:12]), np.transpose(word_vectors['executive']))
plt.plot(alpha_man[0,],alpha_man[1,],'d',color='k',label='man')
plt.plot(alpha_woman[0,],alpha_woman[1,],'^',color='r',label='woman')
plt.plot(alpha_executive[0,],alpha_executive[1,],'*',color='b',label='executive')
plt.legend()
# + deletable=true editable=true
# This function return the words d, such that a:b and c:d verifies the same relation
# compute it, in the compressed domain
def analogy(a, b, c,eigenmodes,word_vectors):
va = np.dot(np.transpose(eigenmodes), np.transpose(word_vectors[a]))
vb = np.dot(np.transpose(eigenmodes), np.transpose(word_vectors[b]))
vc = np.dot(np.transpose(eigenmodes), np.transpose(word_vectors[c]))
va = va/np.linalg.norm(va)
vb = vb/np.linalg.norm(vb)
vc = vc/np.linalg.norm(vc)
analogy_word = np.dot(eigenmodes,((vb - va) + vc))
return nearest_neighbor(analogy_word, word_vectors)
# + deletable=true editable=true
R = [10,30,60,90,120,150,180,210,240,270,300]
for r in R:
print('')
print('france - paris + rome = ' + analogy('paris', 'france', 'rome', U[0:300,0:r],word_vectors))
# -
| EigenWords.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TP2 - Le prix du carburant en France
#
# --
#
# Tous les jeux de données sont issus de la page https://www.prix-carburants.gouv.fr/rubrique/opendata/, ce sont de vraies données datant de 2018 sur les prix de l'essence en France.
#
# Pour le côté pratique, j'ai réduit la base de données à 500 stations d'essence, et j'ai extrait des données .xml un .csv. Il est disponible à l'adresse suivante:
# https://raw.githubusercontent.com/llesoil/modelisation_des_problemes_scientifiques-/master/7.pandas_matplotlib/prix_essence.csv
#
# Pour la signification des variables:
# - IdStation est l'indentifiant de la station, chaque station a un numéro unique
# - IdEssence est l'identifiant du type de carburant
# - Nom est le nom correspondant au type de carburant, par exemple "Gazole" ou "SP95"
# - Date donne la date à laquelle le prix a été observé, au format "AAAA-MM-JJ" + T + "hh:mm:ss"
# - Valeur1000L donne le prix, en euros, de 1000 litres d'essence pour le carburant d'identifiant IdEssence dans la station d'identifiant IdStation à la date Date.
#
# Si vous souhaitez travailler sur toute la France, j'ouvrirai un dépôt le jour du TP. La correction sera faite sur la France, mais le code donné est valable sur le jeu réduit également.
#
#
# Librairies à importer utiles aux scripts
# N'hésitez pas à en ajouter si vous préférez en utiliser d'autres
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# ### 1) À l'aide de la librairie pandas, importez le jeu de données dans ce notebook.
# La fonction read_table de pandas pourra être utile.
# On s'intéresse à la station Total de Bourg-en-Bresse (aux coordonnées 46.20114 5.19791). Son identifiant est le '1000001'.
#
# ### 2) a-] Extraire du jeu importé toutes les lignes concernant cette station.
#
# Pensez à utiliser la fonction where de numpy, et .iloc[] de pandas
# ### 2) b-] Quel est le prix moyen du carburant dans cette station?
# ### 2) c-] Cette station est -elle plus ou moins chère que la moyenne?
# ### 3) Créez une variable valeur1L, correspondant au prix par litre
# On pourra utiliser la fonction assign de pandas
# ### 4) En moyenne, quel est le carburant le plus cher en France en 2018?
# Au contraire, quel est le moins cher?
# Quel est celui qui varie le plus au fil du temps?
#
# On pourra utiliser la fonction groupby de pandas.
# ### 5) Affichez un boxplot des prix par type de carburants.
#
# Indication: fonction boxplot de seaborn.
# Si vous ne vous sentez pas à l'aise, regardez la correction des questions 6) a-] b-] c-] d-] et passez directement à la question 6)e-].
#
# ### 6) Évolution du prix du carburant en 2018
#
# ### 6) a-] Créez une fonction qui récupère le mois dans la variable Date
# ### 6) b-] Créez une variable mois dans le dataframe contenant le mois associé à chaque date
# ### 6) c-] Créez ensuite le tableau des moyennes de prix mensuelles par carburant
#
# En colonnes les mois, en lignes les types de carburants
# ### 6) d-] Puis affichez les courbes d'évolution des prix moyens au cours des mois de l'année 2018
# ### 6) e-] Compte-tenu des courbes, quand pensez vous que le gazole dépassera le E10? Même question pour le SP98.
# ...
# Les informations concernant les stations sont référencées dans le .csv à l'adresse suivante:
# https://raw.githubusercontent.com/llesoil/modelisation_des_problemes_scientifiques-/master/7.pandas_matplotlib/station_essence_total.csv
#
# ### 7) a-] Importez la table
# ### 7) b-] Calculez le prix moyen par station, et stocker le résultat sus la forme d'une table
# Avant, enlevez les valeurs manquantes de la variable Valeur1L à l'aide des fonctions invert et isnan de numpy.
# ### 7) c-] Créez une table station, jointure des tables créées respectivement en 7)a-] et en 7) b-]
#
# La clef de jointure, commune aux deux tables, est l'identifiant des stations. On pourra utiliser join de pandas.
# ### 8) Transformez la variable Valeur1L en variable qualitative à quatre classes en raisonnant de la manière suivante:
# - prix très bas <=> en-dessous du premier quartile
# - prix assez bas <=> entre le premier quartile et la médiane
# - prix assez fort <=> entre la médiane et le troisième quartile
# - prix très fort <=> supérieur au troisième quartile
# ### 9) Vaut-il mieux acheter son carburant sur route ou sur autoroute?
#
# La variable typeRoute vaut "A" pour autoroute et "R" pour route. On pourra faire un tableau de contingence entre typeRoute et la variable créée en 8).
# Les prix sont homogènes sur les routes. Sur les autoroutes, ils sont plutôt plus hauts.
# ### 10) Le département le plus cher de France
#
# ### 10) a-] Créez une variable dep, qui contiendra le numéro de département de la station
# ### 10) b-] Selon la base de données, dans quel département paye-t-on le plus cher?
# ### 11) Affichez sur une carte les dix stations les moins chères de France en bleu, puis les dix stations les plus chères de France en rouge.
#
# Pour les coordonnées, la latitude et la longitude doivent être divisées par $10^5$. On pourra utiliser le module cartopy.
| 7.pandas_matplotlib/TP2 - Le prix du carburant en France.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # データ数が同一になるようにビン分割を行う
# +
import numpy as np
import pandas as pd
# %matplotlib inline
from matplotlib import pyplot as plt
# +
# 年齢データの生成
## 平均40, 標準偏差10, 1000サンプル
age = np.random.normal(40, 10, 1000)
# 最大99.9, 最小0に丸める
age[age < 0] = 0.0
age[age > 99.9] = 99.9
# -
# 年齢データのヒストグラム
plt.hist(age)
# 10分割
age_ctgr = pd.qcut(age, q=10)
# 分割結果を確認
age_ctgr
# +
# カテゴリごとに集計しplot
plot_data = age_ctgr.value_counts().sort_index()
x_labels = [str(ctgr) for ctgr in plot_data.index]
plt.xticks(rotation=90)
plt.bar(x_labels, plot_data)
# +
# カテゴリごとに集計しラベルが昇順になるようにplot
age_ctgr, bin_def = pd.qcut(age, q=10, retbins=True)
age_ctgr_name = ['{:02}_{:.1f}-{:.1f}'.format(i, bin_def[i], bin_def[i+1]) for i in range(len(bin_def)-1)]
age_ctgr = pd.qcut(age, q=10, labels=age_ctgr_name)
plot_data = age_ctgr.value_counts().sort_index()
x_labels = [str(ctgr) for ctgr in plot_data.index]
plt.xticks(rotation=90)
plt.bar(x_labels, plot_data)
| pandas/binning/qcut.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "7a5776cc-1da2-46c2-82fc-a3db8e5a04c4", "showTitle": false, "title": ""}
# # Structured Streaming using the Python DataFrames API
#
# Apache Spark includes a high-level stream processing API, [Structured Streaming](http://spark.apache.org/docs/latest/structured-streaming-programming-guide.html). In this notebook we take a quick look at how to use the DataFrame API to build Structured Streaming applications. We want to compute real-time metrics like running counts and windowed counts on a stream of timestamped actions (e.g. Open, Close, etc).
#
# To run this notebook, import it and attach it to a Spark cluster.
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "39eb54e7-59e7-4217-9619-79e4bf885027", "showTitle": false, "title": ""}
# ## Sample Data
# We have some sample action data as files in `/databricks-datasets/structured-streaming/events/` which we are going to use to build this appication. Let's take a look at the contents of this directory.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "5a9ba90c-2b0d-4385-ab09-1b21179c000d", "showTitle": false, "title": ""}
# %fs ls /databricks-datasets/structured-streaming/events/
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "f4b333bd-50e7-43d5-af62-5a5fa699770a", "showTitle": false, "title": ""}
# There are about 50 JSON files in the directory. Let's see what each JSON file contains.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "d0d0af05-2226-48ba-b6fd-713146ac56fb", "showTitle": false, "title": ""}
# %fs head /databricks-datasets/structured-streaming/events/file-0.json
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "473a6faa-75e9-43d1-8cdc-e16431e44a13", "showTitle": false, "title": ""}
# Each line in the file contains JSON record with two fields - `time` and `action`. Let's try to analyze these files interactively.
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "804bc3ee-1437-461b-8395-e9d18b32a8f3", "showTitle": false, "title": ""}
# ## Batch/Interactive Processing
# The usual first step in attempting to process the data is to interactively query the data. Let's define a static DataFrame on the files, and give it a table name.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "052de597-ddb8-4ff4-9693-57b6706c5156", "showTitle": false, "title": ""}
from pyspark.sql.types import *
inputPath = "/databricks-datasets/structured-streaming/events/"
# Since we know the data format already, let's define the schema to speed up processing (no need for Spark to infer schema)
jsonSchema = StructType([ StructField("time", TimestampType(), True), StructField("action", StringType(), True) ])
# Static DataFrame representing data in the JSON files
staticInputDF = (
spark
.read
.schema(jsonSchema)
.json(inputPath)
)
display(staticInputDF)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "0d5b179e-2575-4d6e-a05f-30949e5c9806", "showTitle": false, "title": ""}
# Now we can compute the number of "open" and "close" actions with one hour windows. To do this, we will group by the `action` column and 1 hour windows over the `time` column.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "d856da4a-d352-42c0-9701-fe91a7eef5e2", "showTitle": false, "title": ""}
from pyspark.sql.functions import * # for window() function
staticCountsDF = (
staticInputDF
.groupBy(
staticInputDF.action,
window(staticInputDF.time, "1 hour"))
.count()
)
staticCountsDF.cache()
# Register the DataFrame as table 'static_counts'
staticCountsDF.createOrReplaceTempView("static_counts")
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "b3627362-e104-40da-b0f9-b764cb0388d1", "showTitle": false, "title": ""}
# Now we can directly use SQL to query the table. For example, here are the total counts across all the hours.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "a9846312-306a-45d8-84ac-de7657ae4e77", "showTitle": false, "title": ""}
# %sql select action, sum(count) as total_count from static_counts group by action
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "19b663d7-aa35-4086-87a9-7fa963df2541", "showTitle": false, "title": ""}
# How about a timeline of windowed counts?
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "31ba8161-4316-4084-96e8-a0857c2ec6fe", "showTitle": false, "title": ""}
# %sql select action, date_format(window.end, "MMM-dd HH:mm") as time, count from static_counts order by time, action
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "617e1996-8950-419f-a829-b1f815af2d83", "showTitle": false, "title": ""}
# Note the two ends of the graph. The close actions are generated such that they are after the corresponding open actions, so there are more "opens" in the beginning and more "closes" in the end.
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "fda2a865-86b6-4963-af5f-385bd2334e25", "showTitle": false, "title": ""}
# ## Stream Processing
# Now that we have analyzed the data interactively, let's convert this to a streaming query that continuously updates as data comes. Since we just have a static set of files, we are going to emulate a stream from them by reading one file at a time, in the chronological order they were created. The query we have to write is pretty much the same as the interactive query above.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "1a940eea-2d4d-431c-ad2e-d9cedcb068d3", "showTitle": false, "title": ""}
from pyspark.sql.functions import *
# Similar to definition of staticInputDF above, just using `readStream` instead of `read`
streamingInputDF = (
spark
.readStream
.schema(jsonSchema) # Set the schema of the JSON data
.option("maxFilesPerTrigger", 1) # Treat a sequence of files as a stream by picking one file at a time
.json(inputPath)
)
# Same query as staticInputDF
streamingCountsDF = (
streamingInputDF
.groupBy(
streamingInputDF.action,
window(streamingInputDF.time, "1 hour"))
.count()
)
# Is this DF actually a streaming DF?
streamingCountsDF.isStreaming
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "1ccbd0af-9c8c-4df2-9387-f1afc5edbcdd", "showTitle": false, "title": ""}
# As you can see, `streamingCountsDF` is a streaming Dataframe (`streamingCountsDF.isStreaming` was `true`). You can start streaming computation, by defining the sink and starting it.
# In our case, we want to interactively query the counts (same queries as above), so we will set the complete set of 1 hour counts to be in a in-memory table.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "fe9e4efa-f07a-4474-a76c-70ef9ba85558", "showTitle": false, "title": ""}
spark.conf.set("spark.sql.shuffle.partitions", "2") # keep the size of shuffles small
query = (
streamingCountsDF
.writeStream
.format("memory") # memory = store in-memory table
.queryName("counts") # counts = name of the in-memory table
.outputMode("complete") # complete = all the counts should be in the table
.start()
)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "36736a03-fed5-4173-9720-6d94659083ab", "showTitle": false, "title": ""}
# `query` is a handle to the streaming query that is running in the background. This query is continuously picking up files and updating the windowed counts.
#
# Note the status of query in the above cell. The progress bar shows that the query is active.
# Furthermore, if you expand the `> counts` above, you will find the number of files they have already processed.
#
# Let's wait a bit for a few files to be processed and then interactively query the in-memory `counts` table.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "fc5b6fc1-f142-4ff8-98de-e719c9405b40", "showTitle": false, "title": ""}
from time import sleep
sleep(5) # wait a bit for computation to start
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "6fee0151-474e-44c5-90ae-24bd154dde16", "showTitle": false, "title": ""}
# %sql select action, date_format(window.end, "MMM-dd HH:mm") as time, count from counts order by time, action
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "4642a107-c2ac-460f-9bea-1d0f6f4b240d", "showTitle": false, "title": ""}
# We see the timeline of windowed counts (similar to the static one earlier) building up. If we keep running this interactive query repeatedly, we will see the latest updated counts which the streaming query is updating in the background.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "49a730c8-66e0-41e3-a0e9-63dff4a52244", "showTitle": false, "title": ""}
sleep(5) # wait a bit more for more data to be computed
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "b72203c7-fffc-4595-8df5-adb442374e23", "showTitle": false, "title": ""}
# %sql select action, date_format(window.end, "MMM-dd HH:mm") as time, count from counts order by time, action
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "f88e440f-6de5-421a-9b37-dbbf455324ac", "showTitle": false, "title": ""}
sleep(5) # wait a bit more for more data to be computed
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "e4a4b8d5-4779-431a-a12c-4f59d683f3f2", "showTitle": false, "title": ""}
# %sql select action, date_format(window.end, "MMM-dd HH:mm") as time, count from counts order by time, action
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "e5caecec-2eb9-4d1f-8694-89760b28abbf", "showTitle": false, "title": ""}
# Also, let's see the total number of "opens" and "closes".
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "4eb9131c-58b5-4d84-90f7-0dd5fc06f079", "showTitle": false, "title": ""}
# %sql select action, sum(count) as total_count from counts group by action order by action
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "68cde8b1-799f-4244-bd32-148ec58260c9", "showTitle": false, "title": ""}
# If you keep running the above query repeatedly, you will always find that the number of "opens" is more than the number of "closes", as expected in a data stream where a "close" always appear after corresponding "open". This shows that Structured Streaming ensures **prefix integrity**. Read the blog posts linked below if you want to know more.
#
# Note that there are only a few files, so consuming all of them there will be no updates to the counts. Rerun the query if you want to interact with the streaming query again.
#
# Finally, you can stop the query running in the background, either by clicking on the 'Cancel' link in the cell of the query, or by executing `query.stop()`. Either way, when the query is stopped, the status of the corresponding cell above will automatically update to `TERMINATED`.
| big-data-school-5/BigDataHomework-main/Spark/Structured Streaming using Python DataFrames.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import nltk
import pandas as pd
import numpy as np
df
df = pd.read_csv(r"C:\Users\lebah\Documents\GitHub\bigco\data\raw\employee_reviews.csv")
df['summary'] = df['summary'].astype(str)
df['cons'] = df['cons'].astype(str)
df['pros'] = df['pros'].astype(str)
# +
df['summary_word_token'] = df['summary'].apply(nltk.word_tokenize)
df['cons_word_token'] = df['cons'].apply(nltk.word_tokenize)
df['pros_word_token'] = df['pros'].apply(nltk.word_tokenize)
df['summary_sent_token'] = df['summary'].apply(nltk.sent_tokenize)
df['cons_sent_token'] = df['cons'].apply(nltk.sent_tokenize)
df['pros_sent_token'] = df['pros'].apply(nltk.sent_tokenize)
# -
df['summary_sent_token'][1]
df
# # Save file
review_interim_csv = df.to_csv(r"C:\Users\lebah\Documents\GitHub\bigco\data\interim\employee_reviews_tokenized.csv")
| notebooks/EDA/Tokenize_EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Find the Newsroom URLs
#
# After collecting information on the Fortune 100 companies from Fortune's website, the next step in the process is to find the links to the newsrooms for each company. Typically, companies will host links to their press releases on a page typically labeled Newsroom, although sometimes labeled Press room. In some instances, a company won't link to its newsroom on the main website at all, and will instead have a link available on the Corporate page.
#
# In order to find the page where each company keeps their press releases, the code below gathers all of the links on the company's website that I collected in the first notebook, and then rates each link based upon their similarity to "news", "press" and "corporate" to help me determine the link's likelihood of being the newsroom link.
# ## Imports
# +
import pandas as pd
from tqdm import tqdm
import time
from selenium import webdriver
from selenium.webdriver.common.by import By
from fuzzywuzzy import fuzz
import warnings
warnings.filterwarnings('ignore')
# -
# read in the data
cos = pd.read_csv('../data/fortune_100_data.csv')
cos.head()
# ## Finding potential newsrooms from corporate websites
# Similar to the situation I found while I was scraping the Fortune website, many companies will use JavaScript on the main pages of their website, which inhibits the use of the `requests` library. To again work around this, I've used `Selenium` in order to gather the links from the company's website.
#
# The below code visits each `co_website` link collected in the previous notebook and scrapes the HTML for links. It then assesses each link using `fuzzywuzzy` to determine its similarity to 'news', 'press' and 'corporate' to account for differences in how each company may refer to their newsroom page. It then pulls the link with the largest value into the `cos` DataFrame for each company.
# + code_folding=[]
# for any companies that simply don't work, catch them in
# this list to review later
error_cos = []
# prepare the options for the chrome driver
options = webdriver.ChromeOptions()
# making headless so as not to bombard my screen
options.add_argument('headless')
# getting around website features that stop bots
options.add_experimental_option("excludeSwitches", ["enable-automation"])
options.add_experimental_option('useAutomationExtension', False)
# start chrome browser
browser = webdriver.Chrome(options=options)
# getting around website features that stop bots
browser.execute_cdp_cmd(
'Network.setUserAgentOverride', {
"userAgent":
'Mozilla/5.0 (Windows NT 10.0; Win64; x64) \
AppleWebKit/537.36 (KHTML, like Gecko) \
Chrome/83.0.4103.53 Safari/537.36'
})
# iterate through each of the companies in the dataframe
# use multiple try/except to be able to continue gathering data
# for other websites even if the website doesnt work on the first pass
# create different print statements in each of the excepts
# to be able to troubleshoot later, if needed
for i in tqdm(range(len(cos))):
try:
# get the site url
site_url = cos.loc[i, 'co_website']
# find the base of the url to use later
base = site_url.split('.')[1]
try:
# open the website & sleep to avoid security features
browser.get(f'{site_url}')
time.sleep(2)
except:
print(f'{cos.loc[i, "company"]} browser')
print(f'{browser.current_url}, {browser.title}')
# get all the links from the main website
links = browser.find_elements(By.TAG_NAME, 'a')
# create list to put all the links from the website into
site_links = []
# iterate through all of the links and add to a dictionary
# that will be added to `site_links`
for l in links:
link_info = {}
url = l.get_attribute('href')
link_info['link'] = url
# use fuzzywuzzy package to assess the similarity of each link to a
# string to find the newsroom, pressroom, and corporate website links
link_info['news_ratio'] = fuzz.partial_ratio('news', url)
link_info['press_ratio'] = fuzz.partial_ratio('press', url)
link_info['corporate_ratio'] = fuzz.partial_ratio('corporate', url)
try:
link_info['url_len'] = len(url)
except:
pass
# append link info to the `site_links` list
site_links.append(link_info)
# putting the site links into a data frame
site_links_df = pd.DataFrame(site_links).drop_duplicates().dropna()
site_links_df = site_links_df[site_links_df['link'].str.contains(base)]
# getting dataframes for each of the series of links and resetting at
# the top of each loop to avoid links from one website being put into
# another company's row
try:
news_link_df = None
news_link_df = site_links_df[
site_links_df['news_ratio'] >
site_links_df['news_ratio'].mean()].sort_values(
'news_ratio', ascending=False).reset_index(drop=True)
except:
print(f'{cos.loc[i, "company"]} news_link_df')
try:
press_link_df = None
press_link_df = site_links_df[
site_links_df['press_ratio'] >
site_links_df['press_ratio'].mean()].sort_values(
'press_ratio', ascending=False).reset_index(drop=True)
except:
print(f'{cos.loc[i, "company"]} press_link_df')
try:
corp_link_df = None
corp_link_df = site_links_df[
site_links_df['corporate_ratio'] >
site_links_df['corporate_ratio'].mean()].sort_values(
'corporate_ratio', ascending=False).reset_index(drop=True)
except:
print(f'{cos.loc[i, "company"]} corp_link_df')
# pulling the top links into cos
try:
cos.loc[i, 'newsroom_link'] = news_link_df.loc[0, 'link']
except:
cos.loc[i, 'newsroom_link'] = 'N/A'
try:
cos.loc[i, 'pressroom_link'] = press_link_df.loc[0, 'link']
except:
cos.loc[i, 'pressroom_link'] = 'N/A'
try:
cos.loc[i, 'corporate_link'] = corp_link_df.loc[0, 'link']
except:
cos.loc[i, 'corporate_link'] = 'N/A'
except:
error_cos.append(cos.loc[i, "company"])
print(f'{cos.loc[i, "company"]}')
print(f'{browser.current_url}, {browser.title}')
# -
cos.head()
# ## Finding the official newsroom link
# After collecting the links for each company that was most similar to 'news', 'press' and 'corporate', I went through and manually investigated whether or not the link was accurate, and saved the information into `final_websites.csv`. While the newsroom links are ultimately not the final links I will use in the function that gathers all of the information, these links were useful in helping me determine what those links would be.
#
# Although doing this manually would not be feasible for larger datasets, in this case it was the best option to make sure I had all of the correct links.
# read in the final_websites csv
final_websites = pd.read_csv('../data/final_websites.csv')
final_websites.head()
cos = cos.merge(final_websites, on='company')
cos.head()
# confirming there are no null values
cos['final'].isna().sum()
# +
# saving dataframe to a new csv
cos.to_csv('../data/fortune_100_data_w_links.csv', index = False)
| code/02-find-newsroom-urls.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
import numpy as np
from random import random, seed
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import StandardScaler
import statistics
from time import time
from scipy.stats import norm
import matplotlib.pyplot as plt
scaler=StandardScaler()
type(scaler)
def FrankeFunction(x,y): #code from task
term1 = 0.75*np.exp(-(0.25*(9*x-2)**2) - 0.25*((9*y-2)**2))
term2 = 0.75*np.exp(-((9*x+1)**2)/49.0 - 0.1*(9*y+1))
term3 = 0.5*np.exp(-(9*x-7)**2/4.0 - 0.25*((9*y-3)**2))
term4 = -0.2*np.exp(-(9*x-4)**2 - (9*y-7)**2)
return term1 + term2 + term3 + term4
#calculates R2 score and MSE
def R2(y_data, y_model): #week 35 exercise
return 1 - np.sum((y_data - y_model) ** 2) / np.sum((y_data - np.mean(y_data)) ** 2)
def MSE(y_data,y_model):
n = np.size(y_model)
return np.sum((y_data-y_model)**2)/n
def SVD(A): #week35 SVD
U, S, VT = np.linalg.svd(A,full_matrices=True)
D = np.zeros((len(U),len(VT)))
print("shape D= ", np.shape(D))
print("Shape S= ",np.shape(S))
print("lenVT =",len(VT))
print("lenU =",len(U))
D = np.eye(len(U),len(VT))*S
"""
for i in range(0,VT.shape[0]): #was len(VT)
D[i,i]=S[i]
print("i=",i)"""
return U @ D @ VT
#Makes a 3d plot of the franke function
def Plot_franke_function(): #code from task
fig = plt.figure()
ax = fig.gca(projection="3d")
# Make data.
x = np.arange(0, 1, 0.05)
y = np.arange(0, 1, 0.05)
x, y = np.meshgrid(x,y)
z = FrankeFunction(x, y)
# Plot the surface.
surf = ax.plot_surface(x, y, z, cmap=cm.coolwarm,
linewidth=0, antialiased=False)
# Customize the z axis.
ax.set_zlim(-0.10, 1.40)
ax.zaxis.set_major_locator(LinearLocator(10))
ax.zaxis.set_major_formatter(FormatStrFormatter('%.02f'))
# Add a color bar which maps values to colors.
fig.colorbar(surf, shrink=0.5, aspect=5)
plt.show()
#Setting up design matrix from week 35-36 lecture slides
def create_X(x, y, n):
if len(x.shape) > 1:
x = np.ravel(x)
y = np.ravel(y)
N = len(x)
l = int((n+1)*(n+2)/2) # Number of elements in beta
X = np.ones((N,l))
for i in range(1,n+1):
q = int((i)*(i+1)/2)
for k in range(i+1):
X[:,q+k] = (x**(i-k))*(y**k)
return X
def OLS_solver(designmatrix, datapoints):
X = designmatrix
z = datapoints
#Splitting training and test data (20%test)
X_train, X_test, z_train, z_test = train_test_split(X, z, test_size=0.2)
#scaling the the input with standardscalar (week35)
scaler = StandardScaler()
scaler.fit(X_train)
X_scaled = scaler.transform(X_train)
#used to scale train and test
z_mean = np.mean(z_train)
z_sigma = np.std(z_train)
z_train = (z_train- z_mean)/z_sigma
#Singular value decomposition (removed as it doesn't work ref group teacher)
#X_train = SVD(X_train)
# Calculating Beta Ordinary Least Square with matrix inversion
ols_beta = np.linalg.pinv(X_train.T @ X_train) @ X_train.T @ z_train #psudoinverse
#Scaling test data
z_test = (z_test- z_mean)/z_sigma
X_test = scaler.transform(X_test)
ztilde = X_train @ ols_beta
#print("Training R2")
#print(R2(z_train,ztilde))
#print("Training MSE")
#print(MSE(z_train,ztilde))
zpredict = X_test @ ols_beta
#print("Test R2")
#print(R2(z_test,zpredict))
#print("Test MSE")
#print(MSE(z_test,zpredict))
print(z_sigma**2 * np.linalg.pinv(X_train.T @ X_train)) #Agree correct? beta_ols_variance =
return ols_beta, MSE(z_train,ztilde), MSE(z_test,zpredict)
# +
#------Task 2------
#setting up data
n = 500 #does it matter?
x = np.linspace(0,1,n)
y = np.linspace(0,1,n)
sigma_N = 0.1; mu_N = 0 #change for value of sigma_N to appropriate values
z = FrankeFunction(x,y) + sigma_N*np.random.randn(n) #adding noise to the dataset
#gives a weird graph which does not bahve as expected
#Because bootsatrap is not implemented?
complexity = []
MSE_train_set = []
MSE_test_set = []
X = create_X(x, y, 40)
ols_beta, MSE_train, MSE_test = OLS_solver(X,z)
#not working as intended
for i in range(2,30): #goes out of range for high i?
X = create_X(x, y, i)
ols_beta, MSE_train, MSE_test = OLS_solver(X,z)
complexity.append(i)
MSE_train_set.append(MSE_train)
MSE_test_set.append(MSE_test)
plt.plot(complexity,MSE_train_set, label ="train")
plt.plot(complexity,MSE_test_set, label ="test")
plt.xlabel("complexity")
plt.ylabel("MSE")
plt.title("Plot of the MSE as a function of complexity of the model")
plt.legend()
plt.grid()
#plt.savefig('Task2plot(n='+str(n)+').pdf')
plt.show()
# -
# +
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
import numpy as np
from random import random, seed
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import StandardScaler
import statistics
from time import time
from scipy.stats import norm
import matplotlib.pyplot as plt
# FrankeFunction: a two-variables function to create the dataset of our vanilla problem
def FrankeFunction(x,y): #code from task
term1 = 0.75*np.exp(-(0.25*(9*x-2)**2) - 0.25*((9*y-2)**2))
term2 = 0.75*np.exp(-((9*x+1)**2)/49.0 - 0.1*(9*y+1))
term3 = 0.5*np.exp(-(9*x-7)**2/4.0 - 0.25*((9*y-3)**2))
term4 = -0.2*np.exp(-(9*x-4)**2 - (9*y-7)**2)
return term1 + term2 + term3 + term4
# Error analysis: MSE and R2 score
def R2(y_data, y_model): #week 35 exercise
return 1 - np.sum((y_data - y_model) ** 2) / np.sum((y_data - np.mean(y_data)) ** 2)
def MSE(y_data,y_model):
n = np.size(y_model)
return np.sum((y_data-y_model)**2)/n
# SVD theorem
def SVD(A): #week35 SVD change to week 36
U, S, VT = np.linalg.svd(A,full_matrices=True)
D = np.zeros((len(U),len(VT)))
print("shape D= ", np.shape(D))
print("Shape S= ",np.shape(S))
print("lenVT =",len(VT))
print("lenU =",len(U))
D = np.eye(len(U),len(VT))*S
"""
for i in range(0,VT.shape[0]): #was len(VT)
D[i,i]=S[i]
print("i=",i)"""
return U @ D @ VT
#Makes a 3d plot of the franke function
def Plot_franke_function(): #code from task
fig = plt.figure()
ax = fig.gca(projection="3d")
# Make data.
x = np.arange(0, 1, 0.05)
y = np.arange(0, 1, 0.05)
x, y = np.meshgrid(x,y)
z = FrankeFunction(x, y)
# Plot the surface.
surf = ax.plot_surface(x, y, z, cmap=cm.coolwarm,
linewidth=0, antialiased=False)
# Customize the z axis.
ax.set_zlim(-0.10, 1.40)
ax.zaxis.set_major_locator(LinearLocator(10))
ax.zaxis.set_major_formatter(FormatStrFormatter('%.02f'))
# Add a color bar which maps values to colors.
fig.colorbar(surf, shrink=0.5, aspect=5)
plt.show()
#Setting up design matrix from week 35-36 lecture slides
def create_X(x, y, n):
if len(x.shape) > 1:
x = np.ravel(x)
y = np.ravel(y)
N = len(x)
l = int((n+1)*(n+2)/2) # Number of elements in beta, number of feutures (order-degree of polynomial)
X = np.ones((N,l))
for i in range(1,n+1):
q = int((i)*(i+1)/2)
for k in range(i+1):
X[:,q+k] = (x**(i-k))*(y**k)
return X
def Split_and_Scale(X,z,test_size=0.2, scale=True):
#Splitting training and test data
X_train, X_test, z_train, z_test = train_test_split(X, z, test_size=test_size)
#scaling the the input with standardscalar (week35)
if scale==True:
scaler=StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
#used to scale train and test --> #why do you do it manually instead of using the Standard scaler?
"""z_mean = np.mean(z_train)
z_sigma = np.std(z_train)
z_train = (z_train- z_mean)/z_sigma"""
#Scaling test data
X_test = scaler.transform(X_test)
#z_test = (z_test- z_mean)/z_sigma
return X_train, X_test, z_train, z_test
def OLS_solver(X_train, X_test, z_train, z_test):
# Calculating Beta Ordinary Least Square with matrix inversion
ols_beta = np.linalg.pinv(X_train.T @ X_train) @ X_train.T @ z_train #psudoinverse
z_tilde = X_train @ ols_beta
z_predict = X_test @ ols_beta
#beta_ols_variance = z_sigma**2 @ np.linalg.pinv(X_train.T @ X_train) #Agree correct?
return ols_beta, z_tilde, z_predict
"""
Task 1 comments:
We still need to find the variance of beta.
What to plot? (use mesh, x,y, z and z_tilda?)
How to find confidence? y-y_tilda = sigma
Sima is the stardard deviation of the error?
print("Beta(ols) variance:") //variance of beta? or = np.mean( np.var(y_pred, axis=1, keepdims=True) )
print(statistics.variance(ols_beta))
plt.plot(X_train,ztilde, label ="u values")
"""
#------Task 1------
# Create vanilla dataset:
n = 1000
x = np.sort(np.random.uniform(0, 1, n))
y = np.sort(np.random.uniform(0, 1, n))
#x, y = np.meshgrid(x,y)
sigma_N = 0.1; mu_N = 0 #change for value of sigma_N to appropriate values
z = FrankeFunction(x,y) + np.random.normal(mu_N,sigma_N,n)#adding noise to the dataset
print(np.max(z),np.min(z))
Plot_franke_function()
degree=5
# OLS
X = create_X(x, y, degree)
X_train, X_test, z_train, z_test = Split_and_Scale(X,z) #StardardScaler, test_size=0.2, scale=true
ols_beta, z_tilde,z_predict = OLS_solver(X_train, X_test, z_train, z_test)
print("Training MSE", MSE(z_train,z_tilde))
print("Test MSE", MSE(z_test,z_predict))
print("-------------------------------------")
print("Training R2", R2(z_train,z_tilde))
print("Test R2", R2(z_test,z_predict))
# Missing confidence interval
# I would plot the data anyway
# -
nx, ny = (3, 2)
print(nx,ny)
x = np.linspace(0, 1, nx)
y = np.linspace(0, 1, ny)
xv, yv = np.meshgrid(x, y)
print(x)
print(y)
print(xv)
print(yv)
xv, yv = np.meshgrid(x, y, sparse=True) # make sparse output arrays
print(xv)
print(yv)
array([[0. , 0.5, 1. ],
[0. , 0.5, 1. ]])
>>> yv
array([[0., 0., 0.],
[1., 1., 1.]])
>>> xv, yv = np.meshgrid(x, y, sparse=True) # make sparse output arrays
>>> xv
array([[0. , 0.5, 1. ]])
>>> yv
array([[0.],
[1.]])
# +
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
import numpy as np
from random import random, seed
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import StandardScaler
import statistics
from time import time
from scipy.stats import norm
import matplotlib.pyplot as plt
# FrankeFunction: a two-variables function to create the dataset of our vanilla problem
def FrankeFunction(x,y): #code from task
term1 = 0.75*np.exp(-(0.25*(9*x-2)**2) - 0.25*((9*y-2)**2))
term2 = 0.75*np.exp(-((9*x+1)**2)/49.0 - 0.1*(9*y+1))
term3 = 0.5*np.exp(-(9*x-7)**2/4.0 - 0.25*((9*y-3)**2))
term4 = -0.2*np.exp(-(9*x-4)**2 - (9*y-7)**2)
return term1 + term2 + term3 + term4
# 3D plot of FrankeFunction
def Plot_franke_function(): #code from task
fig = plt.figure()
ax = fig.gca(projection="3d")
# Make data.
x = np.arange(0, 1, 0.05)
y = np.arange(0, 1, 0.05)
x, y = np.meshgrid(x,y)
z = FrankeFunction(x, y)
# Plot the surface.
surf = ax.plot_surface(x, y, z, cmap=cm.coolwarm,
linewidth=0, antialiased=False)
# Customize the z axis.
ax.set_zlim(-0.10, 1.40)
ax.zaxis.set_major_locator(LinearLocator(10))
ax.zaxis.set_major_formatter(FormatStrFormatter('%.02f'))
# Add a color bar which maps values to colors.
fig.colorbar(surf, shrink=0.5, aspect=5)
plt.show()
# Error analysis: MSE and R2 score
def R2(y_data, y_model): #week 35 exercise
return 1 - np.sum((y_data - y_model) ** 2) / np.sum((y_data - np.mean(y_data)) ** 2)
def MSE(y_data,y_model):
n = np.size(y_model)
return np.sum((y_data-y_model)**2)/n
# SVD theorem
def SVD(A): #week35 SVD change to week 36
U, S, VT = np.linalg.svd(A,full_matrices=True)
D = np.zeros((len(U),len(VT)))
print("shape D= ", np.shape(D))
print("Shape S= ",np.shape(S))
print("lenVT =",len(VT))
print("lenU =",len(U))
D = np.eye(len(U),len(VT))*S
"""
for i in range(0,VT.shape[0]): #was len(VT)
D[i,i]=S[i]
print("i=",i)"""
return U @ D @ VT
# Design matrix
def create_X(x, y, n): # week 35-36 lecture slides
if len(x.shape) > 1:
x = np.ravel(x)
y = np.ravel(y)
N = len(x)
l = int((n+1)*(n+2)/2) # Number of elements in beta, number of feutures (order-degree of polynomial)
X = np.ones((N,l))
for i in range(1,n+1):
q = int((i)*(i+1)/2)
for k in range(i+1):
X[:,q+k] = (x**(i-k))*(y**k)
return X
def Split_and_Scale(X,z,test_size=0.2, scale=True):
#Splitting training and test data
X_train, X_test, z_train, z_test = train_test_split(X, z, test_size=test_size)
#scaling the the input with standardscalar (week35)
if scale==True:
scaler_X = StandardScaler(with_std=False)
scaler_X.fit(X_train)
X_train = scaler_X.transform(X_train)
X_test = scaler_X.transform(X_test)
scaler_z = StandardScaler(with_std=False)
#scaler_z.fit(z_train)
z_train = np.squeeze(scaler_z.fit_transform(z_train.reshape(-1, 1)))
z_test = np.squeeze(scaler_z.transform(z_test.reshape(-1, 1)))
#used to scale train and test --> #why do you do it manually instead of using the Standard scaler?
"""z_mean = np.mean(z_train)
z_sigma = np.std(z_train)
z_train = (z_train- z_mean)/z_sigma"""
#z_test = (z_test- z_mean)/z_sigma
return X_train, X_test, z_train, z_test
def OLS_solver(X_train, X_test, z_train, z_test):
# Calculating Beta Ordinary Least Square with matrix inversion
ols_beta = np.linalg.pinv(X_train.T @ X_train) @ X_train.T @ z_train #psudoinverse
z_tilde = X_train @ ols_beta
z_predict = X_test @ ols_beta
#beta_ols_variance = z_sigma**2 @ np.linalg.pinv(X_train.T @ X_train) #Agree correct?
return ols_beta, z_tilde, z_predict
"""
Task 1 comments:
We still need to find the variance of beta.
What to plot? (use mesh, x,y, z and z_tilda?)
How to find confidence? y-y_tilda = sigma
Sima is the stardard deviation of the error?
print("Beta(ols) variance:") //variance of beta? or = np.mean( np.var(y_pred, axis=1, keepdims=True) )
print(statistics.variance(ols_beta))
plt.plot(X_train,ztilde, label ="u values")
"""
#------Task 1------
# Create vanilla dataset:
n = 1000
x = np.linspace(0,1,n)
y = np.linspace(0,1,n)
sigma_N = 0.1; mu_N = 0 #change for value of sigma_N to appropriate values
z = FrankeFunction(x,y) + np.random.randn(n) #adding noise to the dataset
degree=5
# OLS
X = create_X(x, y, degree)
X_train, X_test, z_train, z_test = Split_and_Scale(X,z) #StardardScaler, test_size=0.2, scale=true
ols_beta, z_tilde,z_predict = OLS_solver(X_train, X_test, z_train, z_test)
print("Training MSE", MSE(z_train,z_tilde))
print("Test MSE", MSE(z_test,z_predict))
print("-------------------------------------")
print("Training R2", R2(z_train,z_tilde))
print("Test R2", R2(z_test,z_predict))
# Missing confidence interval
# I would plot the data anyway
# +
import matplotlib.pyplot as plt
import numpy as np
from sklearn.linear_model import LinearRegression, Ridge, Lasso
from sklearn.preprocessing import PolynomialFeatures
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline
from sklearn.utils import resample
np.random.seed(2018)
n = 40
n_boostraps = 100
maxdegree = 14
# Make data set.
x = np.linspace(-3, 3, n)
print(x.shape)
x=x.reshape(-1, 1)
print(x.shape)
y = np.exp(-x**2) + 1.5 * np.exp(-(x-2)**2)+ np.random.normal(0, 0.1, x.shape)
error = np.zeros(maxdegree)
bias = np.zeros(maxdegree)
variance = np.zeros(maxdegree)
polydegree = np.zeros(maxdegree)
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.2)
for degree in range(maxdegree):
model = make_pipeline(PolynomialFeatures(degree=degree), LinearRegression(fit_intercept=False))
y_pred = np.empty((y_test.shape[0], n_boostraps))
for i in range(n_boostraps):
x_, y_ = resample(x_train, y_train)
y_pred[:, i] = model.fit(x_, y_).predict(x_test).ravel()
polydegree[degree] = degree
print(y_test.shape, y_pred.shape)
error[degree] = np.mean( np.mean((y_test - y_pred)**2, axis=1, keepdims=True) )
bias[degree] = np.mean( (y_test - np.mean(y_pred, axis=1, keepdims=True))**2 )
variance[degree] = np.mean( np.var(y_pred, axis=1, keepdims=True) )
print('Polynomial degree:', degree)
print('Error:', error[degree])
print('Bias^2:', bias[degree])
print('Var:', variance[degree])
print('{} >= {} + {} = {}'.format(error[degree], bias[degree], variance[degree], bias[degree]+variance[degree]))
plt.plot(polydegree, error, label='Error')
plt.plot(polydegree, bias, label='bias')
plt.plot(polydegree, variance, label='Variance')
plt.legend()
plt.show()
# -
| Projects/Project2/Test/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#default_exp data.seq2seq.translation
# +
#all_slow
# +
#hide
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
import os
os.environ["TOKENIZERS_PARALLELISM"] = "false"
# -
# # data.seq2seq.translation
#
# > This module contains the bits required to use the fastai DataBlock API and/or mid-level data processing pipelines to organize your data for translation tasks
# +
#export
import ast
from functools import reduce
import torch
from datasets import list_datasets, load_dataset
from transformers import *
from fastai.text.all import *
from blurr.utils import *
from blurr.data.core import *
from blurr.data.seq2seq.core import *
logging.set_verbosity_error()
# +
#hide
import pdb
from nbdev.showdoc import *
from fastcore.test import *
from fastai import __version__ as fa_version
from torch import __version__ as pt_version
from transformers import __version__ as hft_version
print(f'Using pytorch {pt_version}')
print(f'Using fastai {fa_version}')
print(f'Using transformers {hft_version}')
# -
#cuda
torch.cuda.set_device(1)
print(f'Using GPU #{torch.cuda.current_device()}: {torch.cuda.get_device_name()}')
ds = load_dataset('wmt16', 'de-en', split='train[:1%]')
# ## Translation tokenization, batch transform, and DataBlock methods
#
# Translation tasks attempt to convert text in one language into another
path = Path('./')
wmt_df = pd.DataFrame(ds['translation'], columns=['de', 'en']); len(wmt_df)
wmt_df.head(2)
# +
pretrained_model_name = "facebook/bart-large-cnn"
model_cls = AutoModelForSeq2SeqLM
hf_arch, hf_config, hf_tokenizer, hf_model = BLURR.get_hf_objects(pretrained_model_name, model_cls=model_cls)
hf_arch, type(hf_tokenizer), type(hf_config), type(hf_model)
# -
blocks = (HF_Seq2SeqBlock(hf_arch, hf_config, hf_tokenizer, hf_model), noop)
dblock = DataBlock(blocks=blocks, get_x=ColReader('de'), get_y=ColReader('en'), splitter=RandomSplitter())
# Two lines! Notice we pass in `noop` for our targets (e.g. our summaries) because the batch transform will take care of both out inputs and targets.
# +
# dblock.summary(wmt_df)
# -
dls = dblock.dataloaders(wmt_df, bs=4)
b = dls.one_batch()
len(b), b[0]['input_ids'].shape, b[1].shape
dls.show_batch(dataloaders=dls, max_n=2, input_trunc_at=250, target_trunc_at=250)
# ## Tests
#
# The purpose of the following tests is to ensure as much as possible, that the core DataBlock code above works for the pretrained **translation models** below. These tests are excluded from the CI workflow because of how long they would take to run and the amount of data that would be required to download.
#
# **Note**: Feel free to modify the code below to test whatever pretrained translation models you are working with ... and if any of your pretrained summarization models fail, please submit a github issue *(or a PR if you'd like to fix it yourself)*
[ model_type for model_type in BLURR.get_models(task='ConditionalGeneration')
if (not model_type.__name__.startswith('TF')) ]
pretrained_model_names = [
'facebook/bart-base',
'facebook/wmt19-de-en', # FSMT
'Helsinki-NLP/opus-mt-de-en', # MarianMT
'sshleifer/tiny-mbart',
'google/mt5-small',
't5-small'
]
path = Path('./')
wmt_df = pd.DataFrame(ds['translation'], columns=['de', 'en'])
# +
#slow
#hide_output
model_cls = AutoModelForSeq2SeqLM
bsz = 2
seq_sz = 128
trg_seq_sz = 128
test_results = []
for model_name in pretrained_model_names:
error=None
print(f'=== {model_name} ===\n')
hf_tok_kwargs = {}
if (model_name == 'sshleifer/tiny-mbart'):
hf_tok_kwargs['src_lang'], hf_tok_kwargs['tgt_lang'] = "de_DE", "en_XX"
hf_arch, hf_config, hf_tokenizer, hf_model = BLURR.get_hf_objects(model_name,
model_cls=model_cls,
tokenizer_kwargs=hf_tok_kwargs)
print(f'architecture:\t{hf_arch}\ntokenizer:\t{type(hf_tokenizer).__name__}\n')
# not all architectures include a native pad_token (e.g., gpt2, ctrl, etc...), so we add one here
if (hf_tokenizer.pad_token is None):
hf_tokenizer.add_special_tokens({'pad_token': '<pad>'})
hf_config.pad_token_id = hf_tokenizer.get_vocab()['<pad>']
hf_model.resize_token_embeddings(len(hf_tokenizer))
before_batch_tfm = HF_Seq2SeqBeforeBatchTransform(hf_arch, hf_config, hf_tokenizer, hf_model,
padding='max_length',
max_length=seq_sz,
max_target_length=trg_seq_sz)
def add_t5_prefix(inp): return f'translate German to English: {inp}' if (hf_arch == 't5') else inp
blocks = (HF_Seq2SeqBlock(before_batch_tfm=before_batch_tfm), noop)
dblock = DataBlock(blocks=blocks,
get_x=Pipeline([ColReader('de'), add_t5_prefix]),
get_y=ColReader('en'),
splitter=RandomSplitter())
dls = dblock.dataloaders(wmt_df, bs=bsz)
b = dls.one_batch()
try:
print('*** TESTING DataLoaders ***\n')
test_eq(len(b), 2)
test_eq(len(b[0]['input_ids']), bsz)
test_eq(b[0]['input_ids'].shape, torch.Size([bsz, seq_sz]))
test_eq(len(b[1]), bsz)
test_eq(b[1].shape, torch.Size([bsz, trg_seq_sz]))
if (hasattr(hf_tokenizer, 'add_prefix_space')):
test_eq(hf_tokenizer.add_prefix_space, True)
test_results.append((hf_arch, type(hf_tokenizer).__name__, model_name, 'PASSED', ''))
dls.show_batch(dataloaders=dls, max_n=2, input_trunc_at=1000)
except Exception as err:
test_results.append((hf_arch, type(hf_tokenizer).__name__, model_name, 'FAILED', err))
# -
#slow
#hide_input
test_results_df = pd.DataFrame(test_results, columns=['arch', 'tokenizer', 'model_name', 'result', 'error'])
display_df(test_results_df)
# ## Cleanup
#hide
from nbdev.export import notebook2script
notebook2script()
| nbs/12_data-seq2seq-translation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MODEL PREDICTION
#
# We will predict the results of prediction on the test images provided in the competition using the best models generated from each of the following approaches
# Approaches followed for model creation
# 1. Creating the Self designned CNN from scratch
# 2. Using the Vgg16 architecture as a base and fine tuning the last layer
# +
import os
import json
from keras.models import load_model
import pandas as pd
import pickle
import numpy as np
import shutil
from keras.preprocessing import image
from tqdm.notebook import tqdm
from PIL import ImageFile
# -
BASE_MODEL_PATH = os.path.join(os.getcwd(),"model")
TEST_DIR = os.path.join(os.getcwd(),"csv_files","test.csv")
PREDICT_DIR = os.path.join(os.getcwd(),"pred_dir")
PICKLE_DIR = os.path.join(os.getcwd(),"pickle_files")
JSON_DIR = os.path.join(os.getcwd(),"json_files")
if not os.path.exists(PREDICT_DIR):
os.makedirs(PREDICT_DIR)
else:
shutil.rmtree(PREDICT_DIR)
os.makedirs(PREDICT_DIR)
if not os.path.exists(JSON_DIR):
os.makedirs(JSON_DIR)
# # Prediction using-->Self Trained Model (CNN Scratch)
BEST_MODEL = os.path.join(BASE_MODEL_PATH,"self_trained","distracted-19-0.99.hdf5")
model = load_model(BEST_MODEL)
model.summary()
data_test = pd.read_csv(os.path.join(TEST_DIR))
#testing on the only 10000 images as loading the all test images requires ram>8gb
data_test = data_test[:10000]
data_test.info()
with open(os.path.join(PICKLE_DIR,"labels_list.pkl"),"rb") as handle:
labels_id = pickle.load(handle)
print(labels_id)
# labels_id = {'c1': 0, 'c6': 1, 'c5': 2, 'c8': 3, 'c3': 4, 'c0': 5, 'c7': 6, 'c2': 7, 'c4': 8, 'c9': 9}
# +
def path_to_tensor(img_path):
# loads RGB image as PIL.Image.Image type
img = image.load_img(img_path, target_size=(64, 64))
# convert PIL.Image.Image type to 3D tensor with shape (224, 224, 3)
x = image.img_to_array(img)
# convert 3D tensor to 4D tensor with shape (1, 224, 224, 3) and return 4D tensor
return np.expand_dims(x, axis=0)
def paths_to_tensor(img_paths):
list_of_tensors = [path_to_tensor(img_path) for img_path in tqdm(img_paths)]
return np.vstack(list_of_tensors)
ImageFile.LOAD_TRUNCATED_IMAGES = True
test_tensors = paths_to_tensor(data_test.iloc[:,0]).astype('float32')/255 - 0.5
# +
ypred_test = model.predict(test_tensors,verbose=1)
ypred_class = np.argmax(ypred_test,axis=1)
id_labels = dict()
for class_name,idx in labels_id.items():
id_labels[idx] = class_name
print(id_labels)
for i in range(data_test.shape[0]):
data_test.iloc[i,1] = id_labels[ypred_class[i]]
# +
#to create a human readable and understandable class_name
class_name = dict()
class_name["c0"] = "SAFE_DRIVING"
class_name["c1"] = "TEXTING_RIGHT"
class_name["c2"] = "TALKING_PHONE_RIGHT"
class_name["c3"] = "TEXTING_LEFT"
class_name["c4"] = "TALKING_PHONE_LEFT"
class_name["c5"] = "OPERATING_RADIO"
class_name["c6"] = "DRINKING"
class_name["c7"] = "REACHING_BEHIND"
class_name["c8"] = "HAIR_AND_MAKEUP"
class_name["c9"] = "TALKING_TO_PASSENGER"
with open(os.path.join(JSON_DIR,'class_name_map.json'),'w') as secret_input:
json.dump(class_name,secret_input,indent=4,sort_keys=True)
# +
# creating the prediction results for the image classification and shifting the predicted images to another folder
#with renamed filename having the class name predicted for that image using model
with open(os.path.join(JSON_DIR,'class_name_map.json')) as secret_input:
info = json.load(secret_input)
for i in range(data_test.shape[0]):
new_name = data_test.iloc[i,0].split("/")[-1].split(".")[0]+"_"+info[data_test.iloc[i,1]]+".jpg"
shutil.copy(data_test.iloc[i,0],os.path.join(PREDICT_DIR,new_name))
#saving the model predicted results into a csv file
data_test.to_csv(os.path.join(os.getcwd(),"csv_files","short_test_result.csv"),index=False)
# -
| Model Prediction Results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/devhemza/deeplearningproject/blob/main/text_summarization_feats.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="yfV2WAKJX8Bp"
# %tensorflow_version 1.x
# + colab={"base_uri": "https://localhost:8080/"} id="aY9STWi2f1K9" outputId="66330044-7d7d-4776-ffec-8eccb51f1fe4"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="mxcetbMWf2Ga" outputId="aa54dba1-8b46-4e90-8b9a-f10dda0be3fb"
#Change this with the path to the cloned repo
# %cd /content/drive/MyDrive/M2/DeepLearning/deeplearningproject/
# + id="sU2-hBAFXhjZ"
#imports
import tensorflow as tf
from tensorflow.contrib import rnn
from utils import get_init_embedding_feats
from nltk.tokenize import word_tokenize
from nltk.tag import pos_tag
from nltk import pos_tag, ne_chunk
import re
import collections
import pickle
import numpy as np
from gensim.models.keyedvectors import KeyedVectors
from gensim.test.utils import get_tmpfile
from gensim.scripts.glove2word2vec import glove2word2vec
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
from sklearn.metrics.pairwise import cosine_similarity
# + colab={"base_uri": "https://localhost:8080/"} id="gULhX_mpXpkF" outputId="607cbb9c-4c2d-4c9a-a5f8-9cabb4c0cc4f"
# !pip install wget
# + [markdown] id="zKSTuw63Xhja"
# ### ENV Preparation:
# + [markdown] id="30mF5YvKXhjb"
# - Download the glove if it does not exist.
#
# We used the Wikipedia 2014 + Gigaword 5 Glove (6B tokens, 400K vocab, uncased, 300d vectors). to initialize word embedding.
#
# https://nlp.stanford.edu/projects/glove/
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="cIn3oYBiXhjc" outputId="00ed2018-b74c-426a-9d5e-3a2c487bcdfd"
import os
import wget
from os import path
import nltk
nltk.download('punkt')
nltk.download('averaged_perceptron_tagger')
nltk.download('maxent_ne_chunker')
nltk.download('words')
glove_dir = "glove"
glove_url = "https://nlp.stanford.edu/data/wordvecs/glove.6B.300d.zip"
# Download glove vector if not exit
if not path.exists("glove"):
if not os.path.exists(glove_dir):
os.mkdir(glove_dir)
wget.download(glove_url, out=glove_dir)
# Extract glove file
with zipfile.ZipFile(os.path.join("glove", "glove.6B.300d.zip"), "r") as z:
z.extractall(glove_dir)
# + [markdown] id="FFk7LkooXhjc"
# ### Model
# + [markdown] id="1Yy46NL2nI-K"
# **Architecture** : Encoder-decoder RNN with attention. (Feature-rich encoder).
# * Encoder : Bidirectionnel GRU-RNN.
# * Decoder : Unidirectionnel GRU-RNN with the same hidden-state size as the encoder.
# * Attention : Bahdanau attention mechanism.
# * Softmax layer over the target vocabulary.
# + [markdown] id="JtRdMLfVqjHx"
# **Feature-rich encoder** :
# Capturing additional linguistic features, such as part-of-speech tags (**POS**), named-entity (**NER**) tags, and TF-IDF statics of the words.
#
# All of this features and the word based embeddings are passed as an input to the encoder after a concatenation into one long vector.
# + id="0RhXBQXuXhjd"
import tensorflow as tf
from tensorflow.contrib import rnn
from utils import get_init_embedding
class Model(object):
def __init__(self, reversed_dict, article_max_len, summary_max_len, args, forward_only=False):
self.vocabulary_size = len(reversed_dict)
self.embedding_size = args.embedding_size
self.num_hidden = args.num_hidden
self.num_layers = args.num_layers
self.learning_rate = args.learning_rate
self.beam_width = args.beam_width
if not forward_only:
self.keep_prob = args.keep_prob
else:
self.keep_prob = 1.0
self.cell = tf.nn.rnn_cell.BasicLSTMCell
with tf.variable_scope("decoder/projection"):
self.projection_layer = tf.layers.Dense(self.vocabulary_size, use_bias=False)
self.batch_size = tf.placeholder(tf.int32, (), name="batch_size")
self.X = tf.placeholder(tf.int32, [None, article_max_len])
self.X_len = tf.placeholder(tf.int32, [None])
self.decoder_input = tf.placeholder(tf.int32, [None, summary_max_len])
self.decoder_len = tf.placeholder(tf.int32, [None])
self.decoder_target = tf.placeholder(tf.int32, [None, summary_max_len])
self.global_step = tf.Variable(0, trainable=False)
with tf.name_scope("embedding"):
if not forward_only and args.glove:
init_embeddings = tf.constant(get_init_embedding(reversed_dict, self.embedding_size), dtype=tf.float32)
else:
init_embeddings = tf.random_uniform([self.vocabulary_size, self.embedding_size], -1.0, 1.0)
self.embeddings = tf.get_variable("embeddings", initializer=init_embeddings)
self.encoder_emb_inp = tf.transpose(tf.nn.embedding_lookup(self.embeddings, self.X), perm=[1, 0, 2])
self.decoder_emb_inp = tf.transpose(tf.nn.embedding_lookup(self.embeddings, self.decoder_input), perm=[1, 0, 2])
with tf.name_scope("encoder"):
fw_cells = [self.cell(self.num_hidden) for _ in range(self.num_layers)]
bw_cells = [self.cell(self.num_hidden) for _ in range(self.num_layers)]
fw_cells = [rnn.DropoutWrapper(cell) for cell in fw_cells]
bw_cells = [rnn.DropoutWrapper(cell) for cell in bw_cells]
encoder_outputs, encoder_state_fw, encoder_state_bw = tf.contrib.rnn.stack_bidirectional_dynamic_rnn(
fw_cells, bw_cells, self.encoder_emb_inp,
sequence_length=self.X_len, time_major=True, dtype=tf.float32)
self.encoder_output = tf.concat(encoder_outputs, 2)
encoder_state_c = tf.concat((encoder_state_fw[0].c, encoder_state_bw[0].c), 1)
encoder_state_h = tf.concat((encoder_state_fw[0].h, encoder_state_bw[0].h), 1)
self.encoder_state = rnn.LSTMStateTuple(c=encoder_state_c, h=encoder_state_h)
with tf.name_scope("decoder"), tf.variable_scope("decoder") as decoder_scope:
decoder_cell = self.cell(self.num_hidden * 2)
if not forward_only:
attention_states = tf.transpose(self.encoder_output, [1, 0, 2])
attention_mechanism = tf.contrib.seq2seq.BahdanauAttention(
self.num_hidden * 2, attention_states, memory_sequence_length=self.X_len, normalize=True)
decoder_cell = tf.contrib.seq2seq.AttentionWrapper(decoder_cell, attention_mechanism,
attention_layer_size=self.num_hidden * 2)
initial_state = decoder_cell.zero_state(dtype=tf.float32, batch_size=self.batch_size)
initial_state = initial_state.clone(cell_state=self.encoder_state)
helper = tf.contrib.seq2seq.TrainingHelper(self.decoder_emb_inp, self.decoder_len, time_major=True)
decoder = tf.contrib.seq2seq.BasicDecoder(decoder_cell, helper, initial_state)
outputs, _, _ = tf.contrib.seq2seq.dynamic_decode(decoder, output_time_major=True, scope=decoder_scope)
self.decoder_output = outputs.rnn_output
self.logits = tf.transpose(
self.projection_layer(self.decoder_output), perm=[1, 0, 2])
self.logits_reshape = tf.concat(
[self.logits, tf.zeros([self.batch_size, summary_max_len - tf.shape(self.logits)[1], self.vocabulary_size])], axis=1)
else:
tiled_encoder_output = tf.contrib.seq2seq.tile_batch(
tf.transpose(self.encoder_output, perm=[1, 0, 2]), multiplier=self.beam_width)
tiled_encoder_final_state = tf.contrib.seq2seq.tile_batch(self.encoder_state, multiplier=self.beam_width)
tiled_seq_len = tf.contrib.seq2seq.tile_batch(self.X_len, multiplier=self.beam_width)
attention_mechanism = tf.contrib.seq2seq.BahdanauAttention(
self.num_hidden * 2, tiled_encoder_output, memory_sequence_length=tiled_seq_len, normalize=True)
decoder_cell = tf.contrib.seq2seq.AttentionWrapper(decoder_cell, attention_mechanism,
attention_layer_size=self.num_hidden * 2)
initial_state = decoder_cell.zero_state(dtype=tf.float32, batch_size=self.batch_size * self.beam_width)
initial_state = initial_state.clone(cell_state=tiled_encoder_final_state)
decoder = tf.contrib.seq2seq.BeamSearchDecoder(
cell=decoder_cell,
embedding=self.embeddings,
start_tokens=tf.fill([self.batch_size], tf.constant(2)),
end_token=tf.constant(3),
initial_state=initial_state,
beam_width=self.beam_width,
output_layer=self.projection_layer
)
outputs, _, _ = tf.contrib.seq2seq.dynamic_decode(
decoder, output_time_major=True, maximum_iterations=summary_max_len, scope=decoder_scope)
self.prediction = tf.transpose(outputs.predicted_ids, perm=[1, 2, 0])
with tf.name_scope("loss"):
if not forward_only:
crossent = tf.nn.sparse_softmax_cross_entropy_with_logits(
logits=self.logits_reshape, labels=self.decoder_target)
weights = tf.sequence_mask(self.decoder_len, summary_max_len, dtype=tf.float32)
self.loss = tf.reduce_sum(crossent * weights / tf.to_float(self.batch_size))
params = tf.trainable_variables()
gradients = tf.gradients(self.loss, params)
clipped_gradients, _ = tf.clip_by_global_norm(gradients, 5.0)
optimizer = tf.train.AdamOptimizer(self.learning_rate)
self.update = optimizer.apply_gradients(zip(clipped_gradients, params), global_step=self.global_step)
# + [markdown] id="kUxYn3lkXhjk"
# ### Training
#
# + [markdown] id="MFJbCventeAj"
# Here we tried to reproduce the training phase of the paper.
# * Encoder-decoder hidden-state size : 400
# * Optimizer : Adadelta.
# * Learning rate : 0.001.
# * batch-size : 50.
# * Gradient clipping.
# + [markdown] id="XVM8WAXIyW-z"
# For lack of time and performance reasons, we used a reduced dataset to train and test the model.
#
# **Train** : 10,000 pairs of article, summary.
#
# **Test** : 1000 pairs of article, summary.
# + colab={"base_uri": "https://localhost:8080/"} id="wYdESWI7Xhjo" outputId="b9d19315-dddf-4227-8082-905af5e33912"
import time
start = time.perf_counter()
import tensorflow as tf
import argparse
import pickle
import os
from utils import build_dict, build_dataset, batch_iter
params = {
"batch_size":64,
"beam_width":10,
"embedding_size":300,
"glove":False,
"keep_prob":0.8,
"learning_rate":0.001,
"num_epochs":10,
"num_hidden":150,
"num_layers":2,
"toy":False,
"with_model":False
}
class Arg(dict):
"""dot.notation access to dictionary attributes"""
__getattr__ = dict.get
__setattr__ = dict.__setitem__
__delattr__ = dict.__delitem__
args = Arg(params)
if not os.path.exists("saved_model"):
os.mkdir("saved_model")
else:
if args['with_model']:
old_model_checkpoint_path = open('saved_model/checkpoint', 'r')
old_model_checkpoint_path = "".join(["saved_model/",old_model_checkpoint_path.read().splitlines()[0].split('"')[1] ])
print("Building dictionary...")
word_dict, reversed_dict, article_max_len, summary_max_len = build_dict("train", args.toy)
print("Loading training dataset...")
train_x, train_y = build_dataset("train", word_dict, article_max_len, summary_max_len, args.toy)
with tf.Session() as sess:
model = Model(reversed_dict, article_max_len, summary_max_len, args)
sess.run(tf.global_variables_initializer())
saver = tf.train.Saver(tf.global_variables())
if 'old_model_checkpoint_path' in globals():
print("Continuing from previous trained model:" , old_model_checkpoint_path , "...")
saver.restore(sess, old_model_checkpoint_path )
batches = batch_iter(train_x, train_y, args.batch_size, args.num_epochs)
num_batches_per_epoch = (len(train_x) - 1) // args.batch_size + 1
print("\nIteration starts.")
print("Number of batches per epoch :", num_batches_per_epoch)
for batch_x, batch_y in batches:
batch_x_len = list(map(lambda x: len([y for y in x if y != 0]), batch_x))
batch_decoder_input = list(map(lambda x: [word_dict["<s>"]] + list(x), batch_y))
batch_decoder_len = list(map(lambda x: len([y for y in x if y != 0]), batch_decoder_input))
batch_decoder_output = list(map(lambda x: list(x) + [word_dict["</s>"]], batch_y))
batch_decoder_input = list(
map(lambda d: d + (summary_max_len - len(d)) * [word_dict["<padding>"]], batch_decoder_input))
batch_decoder_output = list(
map(lambda d: d + (summary_max_len - len(d)) * [word_dict["<padding>"]], batch_decoder_output))
train_feed_dict = {
model.batch_size: len(batch_x),
model.X: batch_x,
model.X_len: batch_x_len,
model.decoder_input: batch_decoder_input,
model.decoder_len: batch_decoder_len,
model.decoder_target: batch_decoder_output
}
_, step, loss = sess.run([model.update, model.global_step, model.loss], feed_dict=train_feed_dict)
if step % 1000 == 0:
print("step {0}: loss = {1}".format(step, loss))
if step % num_batches_per_epoch == 0:
hours, rem = divmod(time.perf_counter() - start, 3600)
minutes, seconds = divmod(rem, 60)
saver.save(sess, "./saved_model/model.ckpt", global_step=step)
print(" Epoch {0}: Model is saved.".format(step // num_batches_per_epoch),
"Elapsed: {:0>2}:{:0>2}:{:05.2f}".format(int(hours),int(minutes),seconds) , "\n")
# + [markdown] id="jKcQBM8PXhjq"
# ### Test
#
# + colab={"base_uri": "https://localhost:8080/"} id="7fGxU31MXhjr" outputId="e47f87ea-6ec9-47c0-80e7-a31974d486aa"
import tensorflow as tf
tf.reset_default_graph()
import pickle
from utils import build_dict, build_dataset, batch_iter
with open("args.pickle", "rb") as f:
args = pickle.load(f)
print("Loading dictionary...")
word_dict, reversed_dict, article_max_len, summary_max_len = build_dict("valid", args.toy)
print("Loading validation dataset...")
valid_x = build_dataset("valid", word_dict, article_max_len, summary_max_len, args.toy)
valid_x_len = [len([y for y in x if y != 0]) for x in valid_x]
open("result.txt", 'w').close()
with tf.Session() as sess:
print("Loading saved model...")
model = Model(reversed_dict, article_max_len, summary_max_len, args, forward_only=True)
saver = tf.train.Saver(tf.global_variables())
ckpt = tf.train.get_checkpoint_state("./saved_model/")
saver.restore(sess, ckpt.model_checkpoint_path)
batches = batch_iter(valid_x, [0] * len(valid_x), args.batch_size, 1)
print("Writing summaries to 'result.txt'...")
for batch_x, _ in batches:
batch_x_len = [len([y for y in x if y != 0]) for x in batch_x]
valid_feed_dict = {
model.batch_size: len(batch_x),
model.X: batch_x,
model.X_len: batch_x_len,
}
prediction = sess.run(model.prediction, feed_dict=valid_feed_dict)
prediction_output = [[reversed_dict[y] for y in x] for x in prediction[:, 0, :]]
summaries = []
with open("result.txt", "a") as f:
for line in prediction_output:
summary = list()
for word in line:
if word == "</s>":
break
if word not in summary:
summary.append(word)
s = " ".join(summary)
summaries.append(s)
print(s, file=f)
print('Summaries are saved to "result.txt"...')
# -
# #### Evaluation
#
# For evaluation we used ROUGE-1, ROUGE-2, and ROUGE-L.
#
# Average ROUGE score (R)recall, (P)precision, or F1-score on all summaries is printed.
# + id="RlR0lFM-lXsK"
import numpy as np
# +
summaries_hat = []
with open("result.txt", "r") as f:
for line in f:
summaries_hat.append(line[:-1])
summaries = []
with open("sumdata/train/valid.title.filter.txt","r") as f:
for line in f:
summaries.append(line)
articles = []
with open("sumdata/train/valid.article.filter.txt","r") as f:
for line in f:
articles.append(line)
def printRandomExamples(articles, y, y_hat, n_examples = 3):
for i in range(n_examples):
j = np.random.randint(0, len(articles))
print("-"*40)
print(f'Article {i+1} :')
print(articles[j])
print('Original summary :')
print(y[j])
print('Generated summary :')
print(y_hat[j])
print('\n')
return
# + colab={"base_uri": "https://localhost:8080/"} id="80uf-BSGjYc7" outputId="4e9a4d18-841c-429d-af27-2f9dcecbc665"
printRandomExamples(articles, summaries, summaries_hat)
# + [markdown] id="Ux9xycxu36DF"
# https://pypi.org/project/py-rouge/
# * The folder "rouge" of this package should be placed in the root of the project directory
#
# + id="rhHexz_FvTzq"
import rouge
# + id="_683cKDeywmA"
evaluator = rouge.Rouge(metrics=['rouge-n', 'rouge-l'],
max_n = 2,
limit_length=True,
length_limit=15,
length_limit_type='words',
apply_avg=True,
stemming=False)
# + id="LPii8oa40Mwi"
scores = evaluator.get_scores(summaries_hat, summaries)
# + id="X0xoDoPS2GBM"
def prepare_results(p, r, f):
return '\t{}:\t{}: {:5.2f}\t{}: {:5.2f}\t{}: {:5.2f}'.format(metric, 'P', 100.0 * p, 'R', 100.0 * r, 'F1', 100.0 * f)
# + colab={"base_uri": "https://localhost:8080/"} id="j795OFLG1hoU" outputId="4fd979e1-b8a6-418a-c5e7-240c6b26557d"
for metric, results in sorted(scores.items(), key=lambda x: x[0]):
print(prepare_results(results['p'], results['r'], results['f']))
# -
| text_summarization_feats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # Deep Factorization Machines
#
# Learning effective feature combinations is critical to the success of click-through rate prediction task. Factorization machines model feature interactions in a linear paradigm (e.g., bilinear interactions). This is often insufficient for real-world data where inherent feature crossing structures are usually very complex and nonlinear. What's worse, second-order feature interactions are generally used in factorization machines in practice. Modeling higher degrees of feature combinations with factorization machines is possible theoretically but it is usually not adopted due to numerical instability and high computational complexity.
#
# One effective solution is using deep neural networks. Deep neural networks are powerful in feature representation learning and have the potential to learn sophisticated feature interactions. As such, it is natural to integrate deep neural networks to factorization machines. Adding nonlinear transformation layers to factorization machines gives it the capability to model both low-order feature combinations and high-order feature combinations. Moreover, non-linear inherent structures from inputs can also be captured with deep neural networks. In this section, we will introduce a representative model named deep factorization machines (DeepFM) :cite:`Guo.Tang.Ye.ea.2017` which combine FM and deep neural networks.
#
#
# ## Model Architectures
#
# DeepFM consists of an FM component and a deep component which are integrated in a parallel structure. The FM component is the same as the 2-way factorization machines which is used to model the low-order feature interactions. The deep component is a multi-layered perceptron that is used to capture high-order feature interactions and nonlinearities. These two components share the same inputs/embeddings and their outputs are summed up as the final prediction. It is worth pointing out that the spirit of DeepFM resembles that of the Wide \& Deep architecture which can capture both memorization and generalization. The advantages of DeepFM over the Wide \& Deep model is that it reduces the effort of hand-crafted feature engineering by identifying feature combinations automatically.
#
# We omit the description of the FM component for brevity and denote the output as $\hat{y}^{(FM)}$. Readers are referred to the last section for more details. Let $\mathbf{e}_i \in \mathbb{R}^{k}$ denote the latent feature vector of the $i^\mathrm{th}$ field. The input of the deep component is the concatenation of the dense embeddings of all fields that are looked up with the sparse categorical feature input, denoted as:
#
# $$
# \mathbf{z}^{(0)} = [\mathbf{e}_1, \mathbf{e}_2, ..., \mathbf{e}_f],
# $$
#
# where $f$ is the number of fields. It is then fed into the following neural network:
#
# $$
# \mathbf{z}^{(l)} = \alpha(\mathbf{W}^{(l)}\mathbf{z}^{(l-1)} + \mathbf{b}^{(l)}),
# $$
#
# where $\alpha$ is the activation function. $\mathbf{W}_{l}$ and $\mathbf{b}_{l}$ are the weight and bias at the $l^\mathrm{th}$ layer. Let $y_{DNN}$ denote the output of the prediction. The ultimate prediction of DeepFM is the summation of the outputs from both FM and DNN. So we have:
#
# $$
# \hat{y} = \sigma(\hat{y}^{(FM)} + \hat{y}^{(DNN)}),
# $$
#
# where $\sigma$ is the sigmoid function. The architecture of DeepFM is illustrated below.
# 
#
# It is worth noting that DeepFM is not the only way to combine deep neural networks with FM. We can also add nonlinear layers over the feature interactions :cite:`He.Chua.2017`.
#
# + attributes={"classes": [], "id": "", "n": "2"} origin_pos=1 tab=["mxnet"]
from d2l import mxnet as d2l
from mxnet import init, gluon, np, npx
from mxnet.gluon import nn
import os
npx.set_np()
# + [markdown] origin_pos=2
# ## Implemenation of DeepFM
# The implementation of DeepFM is similar to that of FM. We keep the FM part unchanged and use an MLP block with `relu` as the activation function. Dropout is also used to regularize the model. The number of neurons of the MLP can be adjusted with the `mlp_dims` hyperparameter.
#
# + attributes={"classes": [], "id": "", "n": "2"} origin_pos=3 tab=["mxnet"]
class DeepFM(nn.Block):
def __init__(self, field_dims, num_factors, mlp_dims, drop_rate=0.1):
super(DeepFM, self).__init__()
num_inputs = int(sum(field_dims))
self.embedding = nn.Embedding(num_inputs, num_factors)
self.fc = nn.Embedding(num_inputs, 1)
self.linear_layer = nn.Dense(1, use_bias=True)
input_dim = self.embed_output_dim = len(field_dims) * num_factors
self.mlp = nn.Sequential()
for dim in mlp_dims:
self.mlp.add(nn.Dense(dim, 'relu', True, in_units=input_dim))
self.mlp.add(nn.Dropout(rate=drop_rate))
input_dim = dim
self.mlp.add(nn.Dense(in_units=input_dim, units=1))
def forward(self, x):
embed_x = self.embedding(x)
square_of_sum = np.sum(embed_x, axis=1) ** 2
sum_of_square = np.sum(embed_x ** 2, axis=1)
inputs = np.reshape(embed_x, (-1, self.embed_output_dim))
x = self.linear_layer(self.fc(x).sum(1)) \
+ 0.5 * (square_of_sum - sum_of_square).sum(1, keepdims=True) \
+ self.mlp(inputs)
x = npx.sigmoid(x)
return x
# + [markdown] origin_pos=4
# ## Training and Evaluating the Model
# The data loading process is the same as that of FM. We set the MLP component of DeepFM to a three-layered dense network with the a pyramid structure (30-20-10). All other hyperparameters remain the same as FM.
#
# + attributes={"classes": [], "id": "", "n": "4"} origin_pos=5 tab=["mxnet"]
batch_size = 2048
data_dir = d2l.download_extract('ctr')
train_data = d2l.CTRDataset(os.path.join(data_dir, 'train.csv'))
test_data = d2l.CTRDataset(os.path.join(data_dir, 'test.csv'),
feat_mapper=train_data.feat_mapper,
defaults=train_data.defaults)
field_dims = train_data.field_dims
train_iter = gluon.data.DataLoader(
train_data, shuffle=True, last_batch='rollover', batch_size=batch_size,
num_workers=d2l.get_dataloader_workers())
test_iter = gluon.data.DataLoader(
test_data, shuffle=False, last_batch='rollover', batch_size=batch_size,
num_workers=d2l.get_dataloader_workers())
devices = d2l.try_all_gpus()
net = DeepFM(field_dims, num_factors=10, mlp_dims=[30, 20, 10])
net.initialize(init.Xavier(), ctx=devices)
lr, num_epochs, optimizer = 0.01, 30, 'adam'
trainer = gluon.Trainer(net.collect_params(), optimizer,
{'learning_rate': lr})
loss = gluon.loss.SigmoidBinaryCrossEntropyLoss()
d2l.train_ch13(net, train_iter, test_iter, loss, trainer, num_epochs, devices)
# + [markdown] origin_pos=6
# Compared with FM, DeepFM converges faster and achieves better performance.
#
# ## Summary
#
# * Integrating neural networks to FM enables it to model complex and high-order interactions.
# * DeepFM outperforms the original FM on the advertising dataset.
#
# ## Exercises
#
# * Vary the structure of the MLP to check its impact on model performance.
# * Change the dataset to Criteo and compare it with the original FM model.
#
# + [markdown] origin_pos=7 tab=["mxnet"]
# [Discussions](https://discuss.d2l.ai/t/407)
#
| d2l-en/mxnet/chapter_recommender-systems/deepfm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="99508910-31f0-4712-0bb5-c84db0e84f54"
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the "../input/" directory.
from subprocess import check_output
print(check_output(["ls", "../input"]).decode("utf8"))
# + _cell_guid="6cef15db-1f07-0c33-bb54-b89b3097b38e"
import os
import math
import cv2
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
from mpl_toolkits.axes_grid1 import ImageGrid
from PIL import Image
import seaborn as sns
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten, Lambda
from keras.layers import Conv2D, MaxPooling2D, AveragePooling2D
from keras.layers.normalization import BatchNormalization
from keras.preprocessing.image import ImageDataGenerator
from keras.regularizers import l2
from keras.callbacks import ModelCheckpoint
from sklearn.model_selection import train_test_split
from sklearn.utils import shuffle
# %matplotlib inline
# + _cell_guid="6a8c4ac8-c27d-d785-3e58-ea4fd990eed1"
filenames = os.listdir('../input/train-jpg')
df = pd.read_csv('../input/train_v2.csv')
# + _cell_guid="66f8e984-b612-1acf-b63d-53f933fe9a31"
df.info()
# + _cell_guid="a866e67a-16e1-0624-ee6c-ac98132a7418"
df.describe()
# + _cell_guid="977da6f4-348f-aa8a-c41a-0dae8c8c175a"
df['tag_set'] = df['tags'].map(lambda s: set(s.split(' ')))
tags = set()
for t in df['tags']:
s = set(t.split(' '))
tags = tags | s
tag_list = list(tags)
tag_list.sort()
tag_columns = ['tag_' + t for t in tag_list]
for t in tag_list:
df['tag_' + t] = df['tag_set'].map(lambda x: 1 if t in x else 0)
# + _cell_guid="06dd8dcf-3dcd-44d7-21e0-0a9f6040d755"
df.info()
df.describe()
# + _cell_guid="5d411dc7-63c1-b3cb-3a15-9f0280381894"
df.head()
# + _cell_guid="3aab2a71-8012-5372-8acb-3c0983c0c506"
df[tag_columns].sum()
# + _cell_guid="851ad0fd-9293-aa11-8b9d-c1b478a9e62a"
df[tag_columns].sum().sort_values().plot.bar()
# + _cell_guid="0d97827d-6fbb-84ba-da9e-b9e9ae29c351"
tags_count = df.groupby('tags').count().sort_values(by='image_name', ascending=False)['image_name']
print('There are {} unique tag combinations'.format(len(tags_count)))
print()
print(tags_count)
# + _cell_guid="abfb47f9-2e78-dea8-9c97-572c579731c0"
from textwrap import wrap
def display(images, cols=None, maxcols=10, width=14, titles=None):
if cols is None:
cols = len(images)
n_cols = cols if cols < maxcols else maxcols
plt.rc('axes', grid=False)
fig1 = plt.figure(1, (width, width * math.ceil(len(images)/n_cols)))
grid1 = ImageGrid(
fig1,
111,
nrows_ncols=(math.ceil(len(images)/n_cols), n_cols),
axes_pad=(0.1, 0.6)
)
for index, img in enumerate(images):
grid1[index].grid = False
if titles is not None:
grid1[index].set_title('\n'.join(wrap(titles[index], width=25)))
if len(img.shape) == 2:
grid1[index].imshow(img, cmap='gray')
else:
grid1[index].imshow(img)
# + _cell_guid="d4f9efa7-6627-ea75-1cf4-d2983ffb1f4b"
def load_image(filename, resize=True, folder='train-jpg'):
img = mpimg.imread('../input/{}/{}.jpg'.format(folder, filename))
if resize:
img = cv2.resize(img, (64, 64))
return np.array(img)
def mean_normalize(img):
return (img - img.mean()) / (img.max() - img.min())
def normalize(img):
return img / 127.5 - 1
# + _cell_guid="a6604505-c1cf-669a-5cc3-d53fa60170ce"
samples = df.sample(16)
sample_images = [load_image(fn) for fn in samples['image_name']]
INPUT_SHAPE = sample_images[0].shape
print(INPUT_SHAPE)
display(
sample_images,
cols=4,
titles=[t for t in samples['tags']]
)
# + _cell_guid="fde3db34-bfc5-871c-7c0a-d98effb51c47"
def preprocess(img):
img = normalize(img)
return img
display(
[(127.5 * (preprocess(img) + 1)).astype(np.uint8) for img in sample_images],
cols=4,
titles=[t for t in samples['tags']]
)
# + [markdown] _cell_guid="3476a683-d9b8-abfa-0f41-e4d34376270d"
# # Learn
# + _cell_guid="7bfe0084-81a9-7c4d-6ecd-e07f23b12511"
df_train = df
# + _cell_guid="84bd0c01-4c0f-4921-0cb7-dd7ed704f2ab"
X = df_train['image_name'].values
y = df_train[tag_columns].values
n_features = 1
n_classes = y.shape[1]
X, y = shuffle(X, y)
X_train, X_valid, y_train, y_valid = train_test_split(X, y, test_size=0.1)
print('We\'ve got {} feature rows and {} labels'.format(len(X_train), len(y_train)))
print('Each row has {} features'.format(n_features))
print('and we have {} classes'.format(n_classes))
assert(len(y_train) == len(X_train))
print('We use {} rows for training and {} rows for validation'.format(len(X_train), len(X_valid)))
print('Each image has the shape:', INPUT_SHAPE)
print('So far, so good')
# + _cell_guid="c7746f07-8533-a624-be6c-a7dda16e97ad"
print('Memory usage (train) kB', X_train.nbytes//(1024))
print('Memory usage (valid) kB', X_valid.nbytes//(1024))
# + _cell_guid="f7d93f36-b75f-2564-2ee3-19a2786ec2b3"
def generator(X, y, batch_size=32):
X_copy, y_copy = X, y
while True:
for i in range(0, len(X_copy), batch_size):
X_result, y_result = [], []
for x, y in zip(X_copy[i:i+batch_size], y_copy[i:i+batch_size]):
rx, ry = [load_image(x)], [y]
rx = np.array([preprocess(x) for x in rx])
ry = np.array(ry)
X_result.append(rx)
y_result.append(ry)
X_result, y_result = np.concatenate(X_result), np.concatenate(y_result)
yield shuffle(X_result, y_result)
X_copy, y_copy = shuffle(X_copy, y_copy)
# + _cell_guid="33af6d60-7e6e-a4d5-6651-11daf350cf91"
from keras import backend as K
def fbeta(y_true, y_pred, threshold_shift=0):
beta = 2
# just in case of hipster activation at the final layer
y_pred = K.clip(y_pred, 0, 1)
# shifting the prediction threshold from .5 if needed
y_pred_bin = K.round(y_pred + threshold_shift)
tp = K.sum(K.round(y_true * y_pred_bin)) + K.epsilon()
fp = K.sum(K.round(K.clip(y_pred_bin - y_true, 0, 1)))
fn = K.sum(K.round(K.clip(y_true - y_pred, 0, 1)))
precision = tp / (tp + fp)
recall = tp / (tp + fn)
beta_squared = beta ** 2
return (beta_squared + 1) * (precision * recall) / (beta_squared * precision + recall + K.epsilon())
# define the model
model = Sequential()
model.add(Conv2D(48, (8, 8), strides=(2, 2), input_shape=INPUT_SHAPE, activation='elu'))
model.add(BatchNormalization())
model.add(Conv2D(64, (8, 8), strides=(2, 2), activation='elu'))
model.add(BatchNormalization())
model.add(Conv2D(96, (5, 5), strides=(2, 2), activation='elu'))
model.add(BatchNormalization())
model.add(Conv2D(96, (3, 3), activation='elu'))
model.add(BatchNormalization())
model.add(Flatten())
model.add(Dropout(0.3))
model.add(Dense(256, activation='elu'))
model.add(BatchNormalization())
model.add(Dense(64, activation='elu'))
model.add(BatchNormalization())
model.add(Dense(n_classes, activation='sigmoid'))
model.compile(
optimizer='adam',
loss='binary_crossentropy',
metrics=[fbeta, 'accuracy']
)
model.summary()
# + _cell_guid="0561a61b-ee47-d683-39f1-1e814d59a4e8"
EPOCHS = 3
BATCH = 32
PER_EPOCH = 256
X_train, y_train = shuffle(X_train, y_train)
X_valid, y_valid = shuffle(X_valid, y_valid)
filepath="weights-improvement-{epoch:02d}-{val_fbeta:.3f}.hdf5"
checkpoint = ModelCheckpoint(filepath, monitor='val_fbeta', verbose=1, save_best_only=True, mode='max')
callbacks_list = [checkpoint]
history = model.fit_generator(
generator(X_train, y_train, batch_size=BATCH),
steps_per_epoch=PER_EPOCH,
epochs=EPOCHS,
validation_data=generator(X_valid, y_valid, batch_size=BATCH),
validation_steps=len(y_valid)//(4*BATCH),
callbacks=callbacks_list
)
| Understanding the Amazon from Space/amazon_from_space.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="tG6UIshqzvc3" colab_type="text"
# - check text data again and cleaning
# - USE BERT, SOTA model
#
# + id="Q8kQEHe6t3D6" colab_type="code" colab={}
import pandas as pd
import numpy as np
import os
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
import seaborn as sns
from nltk.corpus import stopwords
from nltk.util import ngrams
from wordcloud import WordCloud
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
from sklearn.model_selection import train_test_split
from sklearn.decomposition import PCA, TruncatedSVD
from sklearn.metrics import classification_report,confusion_matrix
from collections import defaultdict
from collections import Counter
plt.style.use('ggplot')
import re
from nltk.tokenize import word_tokenize
import gensim
import string
from tqdm import tqdm
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.models import Sequential
from keras.layers import Embedding, LSTM,Dense, SpatialDropout1D, Dropout
from keras.initializers import Constant
from keras.optimizers import Adam
# + id="f_Cq4m1QXEzU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 372} outputId="2d8c9e12-c339-4a06-fc34-6982f3e9cc79"
# !nvidia-smi
# + id="4K7ymb39uCmW" colab_type="code" colab={}
tweet= pd.read_csv('train.csv')
test=pd.read_csv('test.csv')
submission = pd.read_csv("sample_submission.csv")
# + id="brAhXGxluPp8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 200} outputId="3731e652-516f-41b8-d11d-49d32122e0c0"
tweet.head()
# + id="l7pwe_c8uXWQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 200} outputId="44f2552c-d5c7-4727-b457-cb943ad5ae24"
test.head()
# + id="zMIujJCBuaPe" colab_type="code" colab={}
# check class distribution
Real_len = tweet[tweet['target']==1].shape[0]
Not_len = tweet[tweet['target']==0].shape[0]
# + id="2StvzfaKu0AH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="aa1aa825-2e9c-4bc9-a6e0-3cd66f3ea3a1"
print(Real_len, Not_len)
# + id="_EG8nF8Vu5-n" colab_type="code" colab={}
# check length of the text
## function to check
def length(text):
return len(text)
# + id="PIdS61s3vGwj" colab_type="code" colab={}
# add length column to the data
tweet['length'] = tweet['text'].apply(length)
# + id="JL1yXJgovKxo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 374} outputId="d512b54f-7500-4fb1-c6e4-5d4ac487e0e0"
# check the length
# disaster red, not disaster blue
plt.rcParams['figure.figsize'] = (18.0, 6.0)
bins = 50
plt.hist(tweet[tweet['target']==0]['length'], alpha=0.6, bins = bins, label='Not')
plt.hist(tweet[tweet['target']==1]['length'], alpha=0.8, bins = bins, label='Real')
plt.grid()
plt.show()
# + id="i2MMwo5pvjOm" colab_type="code" colab={}
# create corpus
def create_corpus(target):
corpus = []
for x in tweet[tweet['target'] == target]['text'].str.split():
for i in x:
corpus.append(i)
return corpus
# + id="ynbduT6rv14y" colab_type="code" colab={}
# create corpus df
def create_corpus_df(tweet, target):
corpus = []
for x in tweet[tweet['target'] == target]['text'].str.split():
for i in x:
corpus.append(i)
return corpus
# + [markdown] id="NmNHcmmhyNti" colab_type="text"
# stop words : tweets with class 0
# + id="GsGGySnjaC-F" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="7e795b68-1704-48a5-9765-8b314d58079d"
import nltk
nltk.download('stopwords')
# + id="JAISQbR_ZgvK" colab_type="code" colab={}
from nltk.corpus import stopwords
stop = set(stopwords.words('english'))
# + id="azAhDtxDyEL6" colab_type="code" colab={}
corpus = create_corpus(0)
# + id="bnw6fOcTYm3v" colab_type="code" colab={}
dic = defaultdict(int)
for word in corpus :
if word in stop:
dic[word] += 1
# + id="kPUZdkGhavlD" colab_type="code" colab={}
top=sorted(dic.items(), key=lambda x:x[1],reverse=True)[:10]
# + id="3LzdohZhYuWX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="8ad0bb20-56b2-442c-907b-5f586d89d111"
np.array(stop) # 해당 텍스트에 있는 불용어 제거
# + [markdown] id="zR3lXTW2azbY" colab_type="text"
# stop words : tweets with class 1
# + id="KsnmOF6Kanhl" colab_type="code" colab={}
corpus = create_corpus(1)
# + id="CdL58KpCa2_w" colab_type="code" colab={}
dic = defaultdict(int)
for word in corpus:
if word in stop:
dic[word] += 1
# + id="23Yaqpzsa8Ds" colab_type="code" colab={}
top = sorted(dic.items(), key = lambda x:x[1], reverse = True)[:10]
# + id="t1tt5n-lbBPC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 392} outputId="f9bc16c7-1267-4a8a-aea0-0c40e09a20b2"
plt.rcParams['figure.figsize'] = (18, 6)
x,y = zip(*top)
plt.bar(x,y)
# + [markdown] id="bCt7bY6LbpRT" colab_type="text"
# N-gram ( bigrams )
# + id="udYLPzhybKs9" colab_type="code" colab={}
def get_top_tweet_bigrams(corpus, n=None) :
vec = CountVectorizer(ngram_range= (2,2)).fit(corpus)
bag_of_words = vec.transform(corpus)
sum_words = bag_of_words.sum(axis = 0)
words_freq = [(word, sum_words[0, idx]) for word, idx in vec.vocabulary_.items()]
words_freq = sorted(words_freq, key = lambda x:x[1], reverse = True)
return words_freq[:n]
# + id="LN2bCiZqchex" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 338} outputId="035f319a-9eeb-49cd-f5c2-345a28fa4363"
plt.figure(figsize = (16,5))
top_tweet_bigrams = get_top_tweet_bigrams(tweet['text'])[:10]
x,y = map(list, zip(*top_tweet_bigrams))
sns.barplot(x=y, y=x)
# + [markdown] id="gNqCzVRGc43d" colab_type="text"
# elimination of stop words
# + id="l8eGTEJhcwIQ" colab_type="code" colab={}
df = pd.concat([tweet, test])
# + id="hlEqJ07rdFnA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 200} outputId="dc121cbc-d5de-48c2-f664-d16776e5a946"
df.head()
# + [markdown] id="RRLlcu6hfftz" colab_type="text"
# 1. remove url
# + id="YNven0zxdGd2" colab_type="code" colab={}
def remove_url(text):
url = re.compile(r'https?://\S+|www\.\S+')
return url.sub(r'', text)
# + id="IJEaYqyuftWl" colab_type="code" colab={}
df['text'] = df['text'].apply(lambda x : remove_url(x))
# + [markdown] id="1vd8BFNaf1g3" colab_type="text"
# 2. remove HTML tags
# + id="urEEebF0fyLQ" colab_type="code" colab={}
def remove_html(text):
html = re.compile(r'<.*?>')
return html.sub(r'', text)
# + id="PTLbyOK5gBgk" colab_type="code" colab={}
df['text'] = df['text'].apply(lambda x : remove_html(x))
# + [markdown] id="v49JM33WiFKM" colab_type="text"
# 3. removing emojis
# + id="my8q3NmqgI8F" colab_type="code" colab={}
# Reference : https://gist.github.com/slowkow/7a7f61f495e3dbb7e3d767f97bd7304b
def remove_emoji(text):
emoji_pattern = re.compile("["
u"\U0001F600-\U0001F64F" # emoticons
u"\U0001F300-\U0001F5FF" # symbols & pictographs
u"\U0001F680-\U0001F6FF" # transport & map symbols
u"\U0001F1E0-\U0001F1FF" # flags (iOS)
u"\U00002702-\U000027B0"
u"\U000024C2-\U0001F251"
"]+", flags=re.UNICODE)
return emoji_pattern.sub(r'', text)
# + id="yi4Ap67niR4a" colab_type="code" colab={}
df['text']=df['text'].apply(lambda x: remove_emoji(x))
# + [markdown] id="kAltARtviV5X" colab_type="text"
# 4. removing punctuation
# + id="WhOIs60MiTnF" colab_type="code" colab={}
def remove_punct(text):
table=str.maketrans('','',string.punctuation)
return text.translate(table)
# + id="k3Cr9DDwiZzO" colab_type="code" colab={}
df['text']=df['text'].apply(lambda x : remove_punct(x))
# + [markdown] id="KiFhh5Xbijwh" colab_type="text"
# Bag of words counts
# + id="wYPYoRNFia_N" colab_type="code" colab={}
def cv(data):
count_vectorizer = CountVectorizer()
emb = count_vectorizer.fit_transform(data)
return emb, count_vectorizer
# + id="a0h-R0T3i9j1" colab_type="code" colab={}
list_corpus = df['text'].tolist()
list_labels = df['target'].tolist()
# + id="gPW_xEZsjDl6" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(list_corpus, list_labels, test_size = 0.2)
# + id="Z1rtfHdJjOid" colab_type="code" colab={}
X_train_counts, count_vectorizer = cv(X_train)
X_test_counts = count_vectorizer.transform(X_test)
# + [markdown] id="Auml1YDDjrA4" colab_type="text"
# using model named BERT
# - Load BERT from the tensorlow hub
# - load tokenizer from the bert layer
# - encode the txt into tokens, masks, and segment flags
# + id="VCMhWLn2je3z" colab_type="code" colab={}
# use the official tokenization script created by google team
# !wget --quiet https://raw.githubusercontent.com/tensorflow/models/master/official/nlp/bert/tokenization.py
# + id="pbgebG4_kQaS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="13e202ec-b96e-4447-ea5a-e413e83c2855"
# !pip install sentencepiece
# + id="keRuAMvbj41j" colab_type="code" colab={}
import tensorflow as tf
from tensorflow.keras.layers import Dense, Input
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.models import Model
from tensorflow.keras.callbacks import ModelCheckpoint
import tensorflow_hub as hub
import tokenization
# + id="mLxbKb3lj8V0" colab_type="code" colab={}
# 논문에서 sentence의 정의가 512 token( max_len으로 지정한다. )
def bert_encode(texts, tokenizer, max_len=512):
all_tokens = []
all_masks = []
all_segments = []
for text in texts:
text = tokenizer.tokenize(text)
text = text[:max_len-2]
input_sequence = ["[CLS]"] + text + ["[SEP]"]
pad_len = max_len - len(input_sequence)
tokens = tokenizer.convert_tokens_to_ids(input_sequence)
tokens += [0] * pad_len
pad_masks = [1] * len(input_sequence) + [0] * pad_len
segment_ids = [0] * max_len
all_tokens.append(tokens)
all_masks.append(pad_masks)
all_segments.append(segment_ids)
return np.array(all_tokens), np.array(all_masks), np.array(all_segments)
# + id="nvUNTW_alVdx" colab_type="code" colab={}
def build_model(bert_layer, max_len=512):
input_word_ids = Input(shape=(max_len,), dtype=tf.int32, name="input_word_ids")
input_mask = Input(shape=(max_len,), dtype=tf.int32, name="input_mask")
segment_ids = Input(shape=(max_len,), dtype=tf.int32, name="segment_ids")
_, sequence_output = bert_layer([input_word_ids, input_mask, segment_ids])
clf_output = sequence_output[:, 0, :]
if Dropout_num == 0:
# Without Dropout
out = Dense(1, activation='sigmoid')(clf_output)
else:
# With Dropout(Dropout_num), Dropout_num > 0
x = Dropout(Dropout_num)(clf_output)
out = Dense(1, activation='sigmoid')(x)
model = Model(inputs=[input_word_ids, input_mask, segment_ids], outputs=out)
model.compile(Adam(lr=learning_rate), loss='binary_crossentropy', metrics=['accuracy'])
return model
# + [markdown] id="N4xpWi6gos7e" colab_type="text"
# - ref) https://www.kaggle.com/rftexas/text-only-bert-keras
# - target correction
# + id="CyuwMd8jnaeH" colab_type="code" colab={}
def clean_tweets(tweet):
"""Removes links and non-ASCII characters"""
tweet = ''.join([x for x in tweet if x in string.printable])
# Removing URLs
tweet = re.sub(r"http\S+", "", tweet)
return tweet
# + id="yDSlexhKoxZy" colab_type="code" colab={}
def remove_emoji(text):
emoji_pattern = re.compile("["
u"\U0001F600-\U0001F64F" # emoticons
u"\U0001F300-\U0001F5FF" # symbols & pictographs
u"\U0001F680-\U0001F6FF" # transport & map symbols
u"\U0001F1E0-\U0001F1FF" # flags (iOS)
u"\U00002702-\U000027B0"
u"\U000024C2-\U0001F251"
"]+", flags=re.UNICODE)
return emoji_pattern.sub(r'', text)
# + id="stImAwoCozj8" colab_type="code" colab={}
def remove_punctuations(text):
punctuations = '@#!?+&*[]-%.:/();$=><|{}^' + "'`"
for p in punctuations:
text = text.replace(p, f' {p} ')
text = text.replace('...', ' ... ')
if '...' not in text:
text = text.replace('..', ' ... ')
return text
# + id="iePM6yLro1eE" colab_type="code" colab={}
abbreviations = {
"$" : " dollar ",
"€" : " euro ",
"4ao" : "for adults only",
"a.m" : "before midday",
"a3" : "anytime anywhere anyplace",
"aamof" : "as a matter of fact",
"acct" : "account",
"adih" : "another day in hell",
"afaic" : "as far as i am concerned",
"afaict" : "as far as i can tell",
"afaik" : "as far as i know",
"afair" : "as far as i remember",
"afk" : "away from keyboard",
"app" : "application",
"approx" : "approximately",
"apps" : "applications",
"asap" : "as soon as possible",
"asl" : "age, sex, location",
"atk" : "at the keyboard",
"ave." : "avenue",
"aymm" : "are you my mother",
"ayor" : "at your own risk",
"b&b" : "bed and breakfast",
"b+b" : "bed and breakfast",
"b.c" : "before christ",
"b2b" : "business to business",
"b2c" : "business to customer",
"b4" : "before",
"b4n" : "bye for now",
"b@u" : "back at you",
"bae" : "before anyone else",
"bak" : "back at keyboard",
"bbbg" : "bye bye be good",
"bbc" : "british broadcasting corporation",
"bbias" : "be back in a second",
"bbl" : "be back later",
"bbs" : "be back soon",
"be4" : "before",
"bfn" : "bye for now",
"blvd" : "boulevard",
"bout" : "about",
"brb" : "be right back",
"bros" : "brothers",
"brt" : "be right there",
"bsaaw" : "big smile and a wink",
"btw" : "by the way",
"bwl" : "bursting with laughter",
"c/o" : "care of",
"cet" : "central european time",
"cf" : "compare",
"cia" : "central intelligence agency",
"csl" : "can not stop laughing",
"cu" : "see you",
"cul8r" : "see you later",
"cv" : "curriculum vitae",
"cwot" : "complete waste of time",
"cya" : "see you",
"cyt" : "see you tomorrow",
"dae" : "does anyone else",
"dbmib" : "do not bother me i am busy",
"diy" : "do it yourself",
"dm" : "direct message",
"dwh" : "during work hours",
"e123" : "easy as one two three",
"eet" : "eastern european time",
"eg" : "example",
"embm" : "early morning business meeting",
"encl" : "enclosed",
"encl." : "enclosed",
"etc" : "and so on",
"faq" : "frequently asked questions",
"fawc" : "for anyone who cares",
"fb" : "facebook",
"fc" : "fingers crossed",
"fig" : "figure",
"fimh" : "forever in my heart",
"ft." : "feet",
"ft" : "featuring",
"ftl" : "for the loss",
"ftw" : "for the win",
"fwiw" : "for what it is worth",
"fyi" : "for your information",
"g9" : "genius",
"gahoy" : "get a hold of yourself",
"gal" : "get a life",
"gcse" : "general certificate of secondary education",
"gfn" : "gone for now",
"gg" : "good game",
"gl" : "good luck",
"glhf" : "good luck have fun",
"gmt" : "greenwich mean time",
"gmta" : "great minds think alike",
"gn" : "good night",
"g.o.a.t" : "greatest of all time",
"goat" : "greatest of all time",
"goi" : "get over it",
"gps" : "global positioning system",
"gr8" : "great",
"gratz" : "congratulations",
"gyal" : "girl",
"h&c" : "hot and cold",
"hp" : "horsepower",
"hr" : "hour",
"hrh" : "his royal highness",
"ht" : "height",
"ibrb" : "i will be right back",
"ic" : "i see",
"icq" : "i seek you",
"icymi" : "in case you missed it",
"idc" : "i do not care",
"idgadf" : "i do not give a damn fuck",
"idgaf" : "i do not give a fuck",
"idk" : "i do not know",
"ie" : "that is",
"i.e" : "that is",
"ifyp" : "i feel your pain",
"IG" : "instagram",
"iirc" : "if i remember correctly",
"ilu" : "i love you",
"ily" : "i love you",
"imho" : "in my humble opinion",
"imo" : "in my opinion",
"imu" : "i miss you",
"iow" : "in other words",
"irl" : "in real life",
"j4f" : "just for fun",
"jic" : "just in case",
"jk" : "just kidding",
"jsyk" : "just so you know",
"l8r" : "later",
"lb" : "pound",
"lbs" : "pounds",
"ldr" : "long distance relationship",
"lmao" : "laugh my ass off",
"lmfao" : "laugh my fucking ass off",
"lol" : "laughing out loud",
"ltd" : "limited",
"ltns" : "long time no see",
"m8" : "mate",
"mf" : "motherfucker",
"mfs" : "motherfuckers",
"mfw" : "my face when",
"mofo" : "motherfucker",
"mph" : "miles per hour",
"mr" : "mister",
"mrw" : "my reaction when",
"ms" : "miss",
"mte" : "my thoughts exactly",
"nagi" : "not a good idea",
"nbc" : "national broadcasting company",
"nbd" : "not big deal",
"nfs" : "not for sale",
"ngl" : "not going to lie",
"nhs" : "national health service",
"nrn" : "no reply necessary",
"nsfl" : "not safe for life",
"nsfw" : "not safe for work",
"nth" : "nice to have",
"nvr" : "never",
"nyc" : "new york city",
"oc" : "original content",
"og" : "original",
"ohp" : "overhead projector",
"oic" : "oh i see",
"omdb" : "over my dead body",
"omg" : "oh my god",
"omw" : "on my way",
"p.a" : "<NAME>",
"p.m" : "after midday",
"pm" : "prime minister",
"poc" : "people of color",
"pov" : "point of view",
"pp" : "pages",
"ppl" : "people",
"prw" : "parents are watching",
"ps" : "postscript",
"pt" : "point",
"ptb" : "please text back",
"pto" : "please turn over",
"qpsa" : "what happens", #"que pasa",
"ratchet" : "rude",
"rbtl" : "read between the lines",
"rlrt" : "real life retweet",
"rofl" : "rolling on the floor laughing",
"roflol" : "rolling on the floor laughing out loud",
"rotflmao" : "rolling on the floor laughing my ass off",
"rt" : "retweet",
"ruok" : "are you ok",
"sfw" : "safe for work",
"sk8" : "skate",
"smh" : "shake my head",
"sq" : "square",
"srsly" : "seriously",
"ssdd" : "same stuff different day",
"tbh" : "to be honest",
"tbs" : "tablespooful",
"tbsp" : "tablespooful",
"tfw" : "that feeling when",
"thks" : "thank you",
"tho" : "though",
"thx" : "thank you",
"tia" : "thanks in advance",
"til" : "today i learned",
"tl;dr" : "too long i did not read",
"tldr" : "too long i did not read",
"tmb" : "tweet me back",
"tntl" : "trying not to laugh",
"ttyl" : "talk to you later",
"u" : "you",
"u2" : "you too",
"u4e" : "yours for ever",
"utc" : "coordinated universal time",
"w/" : "with",
"w/o" : "without",
"w8" : "wait",
"wassup" : "what is up",
"wb" : "welcome back",
"wtf" : "what the fuck",
"wtg" : "way to go",
"wtpa" : "where the party at",
"wuf" : "where are you from",
"wuzup" : "what is up",
"wywh" : "wish you were here",
"yd" : "yard",
"ygtr" : "you got that right",
"ynk" : "you never know",
"zzz" : "sleeping bored and tired"
}
# + id="JsvV65U5o65B" colab_type="code" colab={}
def convert_abbrev_in_text(text):
tokens = word_tokenize(text)
tokens = [convert_abbrev(word) for word in tokens]
text = ' '.join(tokens)
return text
# + id="YuR_GzGfo9hj" colab_type="code" colab={}
module_url = "https://tfhub.dev/tensorflow/bert_en_uncased_L-24_H-1024_A-16/1"
bert_layer = hub.KerasLayer(module_url, trainable=True)
# + id="oF7-pZOQpkWk" colab_type="code" colab={}
train = pd.read_csv("train.csv")
test = pd.read_csv("test.csv")
# + id="DTwz9ktUpo1f" colab_type="code" colab={}
if False:
ids_with_target_error = [328,443,513,2619,3640,3900,4342,5781,6552,6554,6570,6701,6702,6729,6861,7226]
train.loc[train['id'].isin(ids_with_target_error),'target'] = 0
train[train['id'].isin(ids_with_target_error)]
# + id="bhJZIyfQpqnI" colab_type="code" colab={}
if False:
train["text"] = train["text"].apply(lambda x: clean_tweets(x))
test["text"] = test["text"].apply(lambda x: clean_tweets(x))
train["text"] = train["text"].apply(lambda x: remove_emoji(x))
test["text"] = test["text"].apply(lambda x: remove_emoji(x))
train["text"] = train["text"].apply(lambda x: remove_punctuations(x))
test["text"] = test["text"].apply(lambda x: remove_punctuations(x))
train["text"] = train["text"].apply(lambda x: convert_abbrev_in_text(x))
test["text"] = test["text"].apply(lambda x: convert_abbrev_in_text(x))
# + id="4n__r0Kops9x" colab_type="code" colab={}
vocab_file = bert_layer.resolved_object.vocab_file.asset_path.numpy()
do_lower_case = bert_layer.resolved_object.do_lower_case.numpy()
tokenizer = tokenization.FullTokenizer(vocab_file, do_lower_case)
# + id="vnPKDU8TpvqA" colab_type="code" colab={}
# txt를 token, mask, segment flag로 인코딩
train_input = bert_encode(train.text.values, tokenizer, max_len=160)
test_input = bert_encode(test.text.values, tokenizer, max_len=160)
train_labels = train.target.values
# + id="pcXi5Yz2rCuB" colab_type="code" colab={}
Dropout_num = 0
learning_rate = 6e-6
valid = 0.2
epochs_num = 3
batch_size_num = 16
# + id="1Oxn3P58qWE8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 408} outputId="393ef3c0-ab6e-475c-d190-257f7c44c6f1"
model_BERT = build_model(bert_layer, max_len=160)
model_BERT.summary()
# + id="AoR2oYnbqXm8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 124} outputId="5d7a6348-53f7-4c52-f608-13d04bab6ec9"
# train
checkpoint = ModelCheckpoint('model.h5', monitor='val_loss', save_best_only=True)
train_history = model_BERT.fit(
train_input, train_labels,
validation_split=0.2,
epochs=3,
callbacks=[checkpoint],
batch_size=1
)
# + [markdown] id="aaRzfYecrqzd" colab_type="text"
# submit
# + id="oIFgQ2XJrMfB" colab_type="code" colab={}
# Prediction by BERT model with my tuning
model_BERT.load_weights('model.h5')
test_pred_BERT = model_BERT.predict(test_input)
test_pred_BERT_int = test_pred_BERT.round().astype('int')
# + id="YZgy_XE2rtSB" colab_type="code" colab={}
train_pred_BERT = model_BERT.predict(train_input)
train_pred_BERT_int = train_pred_BERT.round().astype('int')
# + id="AemtzfriruhZ" colab_type="code" colab={}
pred = pd.DataFrame(test_pred_BERT, columns=['preds'])
# + id="JKmHA2Kzrw03" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 352} outputId="740bbcc5-8ca6-4666-d0c0-2be076434295"
submission['target'] = test_pred_BERT_int
submission.head(10) # check
# + id="X4lSDuXFryRp" colab_type="code" colab={}
submission.to_csv("submission2.csv", index=False, header=True)
# + id="97oE_CZWB9Hv" colab_type="code" colab={}
| ChungHyunhee/389_NLP with Disasster Tweets/tweeter_disaster_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import xgboost as xgb
from sklearn.cross_validation import train_test_split
import matplotlib.pyplot as plt
import math
# -
# ### Read the training data
train_df = pd.read_csv('data/train.csv', dtype={"place_id": str})
# According to the discussions in the forum, the time is most likely measured in minutes, so let's extract some temporal features that might be useful.
# ### Add temporal features
train_df['hour_of_day'] = (train_df['time'] / 60) % 24
train_df['day_of_week'] = (train_df['time'] / 60 / 24) % 7 # The offset shouldn't matter
# ### Group by place id
gb = train_df.groupby('place_id')
place_dfs = {x:gb.get_group(x) for x in gb.groups}
# ### Filter out places with very low number of check-ins
place_dfs_filtered = {k:place_dfs[k] for k in place_dfs.keys() if place_dfs[k].shape[0] >= 5}
len(place_dfs_filtered)
# ### Compute Gaussian per place id based on (x,y) coords
def get_gauss(df, cols):
X = df[cols].values
mean = np.mean(X, axis = 0)
S = 1/X.shape[0] * np.dot((X - mean).T,(X - mean))
return (mean, S)
gausss = {p:get_gauss(place_dfs_filtered[p], ['x', 'y']) for p in place_dfs_filtered.keys()}
# ### Build the grid and assign places to each cell
def gridify(x): return int(round(min(max(float(x), 0.0), 9.9), 1) * 10)
def grid_str(x, y): return str(gridify(x)) + 'x' + str(gridify(y))
def grid_range(min_x, max_x): return np.arange(gridify(min_x), gridify(max_x) + 1, 1)
places_grid = {grid_str(x,y):[] for x in np.arange(0,10,0.1) for y in np.arange(0,10,0.1)}
deviations = 1.0
for place in gausss.keys():
mean = gausss[place][0]
S = gausss[place][1]
min_x = mean[0] - deviations * S[0][0]
max_x = mean[0] + deviations * S[0][0]
min_y = mean[1] - deviations * S[1][1]
max_y = mean[1] + deviations * S[1][1]
for x in grid_range(min_x, max_x):
for y in grid_range(min_y, max_y):
places_grid[grid_str(x/10.0,y/10.0)].append(place)
index = 0
indices = {}
for p in places_grid.keys():
print("Processing " + p + "(" + str(index) + ")")
current_df = train_df[train_df['place_id'].isin(places_grid[p])]
current_df.to_csv("data/grid_data/train/train_" + str(index) + "_" + p + ".csv", index=False)
indices[p] = index
index = index + 1
test_df = pd.read_csv('data/test.csv')
test_df['hour_of_day'] = (test_df['time'] / 60) % 24
test_df['day_of_week'] = (test_df['time'] / 60 / 24) % 7 # The offset shouldn't matter
test_df['grid_location'] = test_df.apply(lambda r: grid_str(r['x'], r['y']), axis = 1)
for p in places_grid.keys():
print("Processing " + p + "(" + str(indices[p]) + ")")
current_df = test_df[test_df['grid_location'] == p][['row_id','x', 'y', 'accuracy', 'time', 'hour_of_day', 'day_of_week']]
current_df.to_csv("data/grid_data/test/test_" + str(indices[p]) + "_" + p + ".csv", index=False)
| prepare-grid-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Pre-processing Text Data
#
# The purpose of this notebook is to demonstrate how to pre-processing text data for next-step feature engineering and training a machine learning model via Amazon SageMaker. In this notebook we will focus on pre-processing our text data, and we will use the text data we ingested in a [sequel notebook](https://github.com/aws/amazon-sagemaker-examples/blob/master/data_ingestion/012_Ingest_text_data_v2.ipynb) to showcase text data pre-processing methodologies. We are going to discuss many possible methods to clean and enrich your text, but you do not need to run through every single step below. Usually, a rule of thumb is: if you are dealing with very noisy text, like social media text data, or nurse notes, then medium to heavy pre-processing effort might be needed, and if it's domain-specific corpus, text enrichment is helpful as well; if you are dealing with long and well-written documents such as news articles and papers, very light pre-processing is needed; you can add some enrichment to the data to better capture the sentence to sentence relationship and overall meaning.
#
# ## Overview
# ### Input Format
# Labeled text data sometimes are in a structured data format. You might come across this when working on reviews for sentiment analysis, news headlines for topic modeling, or documents for text classification. One column of the dataset could be dedicated for the label, one column for the text, and sometimes other columns as attributes. You can process this dataset format similar to how you would process tabular data and ingest them in the [last section](https://github.com/aws/amazon-sagemaker-examples/blob/master/preprocessing/tabular_data/preprocessing_tabular_data.ipynb). Sometimes text data, especially raw text data, comes as unstructured data and is often in .json or .txt format. To work with this type of formatting, you will need to first extract useful information from the original dataset.
#
# ### Use Cases
# Text data contains rich information and it's everywhere. Applicable use cases include Voice of Customer (VOC), fraud detection, warranty analysis, chatbot and customer service routing, audience analysis, and much more.
#
# ### What's the difference between pre-processing and feature engineering for text data?
# In the pre-processing stage, you want to clean and transfer the text data from human language to standard, machine-analyzable format for further processing. For feature engineering, you extract predictive factors (features) from the text. For example, for a matching equivalent question pairs task, the features you can extract include words overlap, cosine similarity, inter-word relationships, parse tree structure similarity, TF-IDF (frequency-inverse document frequency) scores, etc.; for some language model like topic modeling, words embeddings themselves can also be features.
#
# ### When is my text data ready for feature engineering?
# When the data is ready to be vectorized and fit your specific use case.
# ## Set Up Notebook
# There are several python packages designed specifically for natural language processing (NLP) tasks. In this notebook, you will use the following packages:
#
# * [`nltk`(natrual language toolkit)](https://www.nltk.org/), a leading platform includes multiple text processing libraries, which covers almost all aspects of pre-processing we will discuss in this section: tokenization, stemming, lemmatization, parsing, chunking, POS tagging, stop words, etc.
#
# * [`SpaCy`](https://spacy.io/), offers most functionality provided by `nltk`, and provides pre-trained word vectors and models. It is scalable and designed for production usage.
#
# * [`Gensim` (Generate Similar)](https://radimrehurek.com/gensim/about.html), "designed specifically for topic modeling, document indexing, and similarity retrieval with large corpora".
#
# * [`TextBlob`](), offers POS tagging, noun phrases extraction, sentiment analysis, classification, parsing, n-grams, word inflation, all offered as an API to perform more advanced NLP tasks. It is an easy-to-use wrapper for libraries like `nltk` and `Pattern`. We will use this package for our enrichment tasks.
# %pip install -qU 'sagemaker>=2.15.0' spacy gensim textblob emot autocorrect
import nltk
import spacy
import gensim
from textblob import TextBlob
import re
import string
import glob
import sagemaker
# +
# Get SageMaker session & default S3 bucket
sagemaker_session = sagemaker.Session()
bucket = sagemaker_session.default_bucket() # replace with your own bucket if you have one
s3 = sagemaker_session.boto_session.resource('s3')
prefix = 'text_sentiment140/sentiment140'
filename = 'training.1600000.processed.noemoticon.csv'
# -
# ### Downloading data from Online Sources
# ### Text Data Sets: Twitter -- sentiment140
# **Sentiment140** The sentiment140 dataset contains 1.6M tweets that were extracted using the [Twitter API](https://developer.twitter.com/en/products/twitter-api) . The tweets have been annotated with sentiment (0 = negative, 4 = positive) and topics (hashtags used to retrieve tweets). The dataset contains the following columns:
# * `target`: the polarity of the tweet (0 = negative, 4 = positive)
# * `ids`: The id of the tweet ( 2087)
# * `date`: the date of the tweet (Sat May 16 23:58:44 UTC 2009)
# * `flag`: The query (lyx). If there is no query, then this value is NO_QUERY.
# * `user`: the user that tweeted (robotickilldozr)
# * `text`: the text of the tweet (Lyx is cool)
#
# +
#helper functions to upload data to s3
def write_to_s3(filename, bucket, prefix):
#put one file in a separate folder. This is helpful if you read and prepare data with Athena
filename_key = filename.split('.')[0]
key = "{}/{}/{}".format(prefix,filename_key,filename)
return s3.Bucket(bucket).upload_file(filename,key)
def upload_to_s3(bucket, prefix, filename):
url = 's3://{}/{}/{}'.format(bucket, prefix, filename)
print('Writing to {}'.format(url))
write_to_s3(filename, bucket, prefix)
# +
#run this cell if you are in SageMaker Studio notebook
# #!apt-get install unzip
# -
# !wget http://cs.stanford.edu/people/alecmgo/trainingandtestdata.zip -O sentimen140.zip
# Uncompressing
# !unzip -o sentimen140.zip -d sentiment140
#upload the files to the S3 bucket
csv_files = glob.glob("sentiment140/*.csv")
for filename in csv_files:
upload_to_s3(bucket, 'text_sentiment140', filename)
# ## Read in Data
# We will read the data in as .csv format since the text is embedded in a structured table.
#
# <b>Note:</b> A frequent error when reading in text data is the encoding error. You can try different encoding options with pandas read_csv when "encoding as UTF-8" does not work; see [python encoding documentation](https://docs.python.org/3/library/codecs.html#standard-encodings) for more encodings you may encounter.
import pandas as pd
prefix = 'text_sentiment140/sentiment140'
filename = 'training.1600000.processed.noemoticon.csv'
data_s3_location = "s3://{}/{}/{}".format(bucket, prefix, filename) # S3 URL
# we will showcase with a smaller subset of data for demonstration purpose
text_data = pd.read_csv(data_s3_location, header = None,
encoding = "ISO-8859-1", low_memory=False,
nrows = 10000)
text_data.columns = ['target', 'tw_id', 'date', 'flag', 'user', 'text']
# ## Examine Your Text Data
# Here you will explore common methods and steps for text pre-processing. Text pre-processing is highly specific to each individual corpus and different tasks, so it is important to examine your text data first and decide what steps are necessary.
#
# First, look at your text data. Seems like there are whitespaces to trim, URLs, smiley faces, numbers, abbreviations, spelling, names, etc. Tweets are less than 140 characters so there is less need for document segmentation and sentence dependencies.
pd.set_option('display.max_colwidth', None) #show full content in a column
text_data['text'][:5]
# ## Pre-Processing
#
# ### Step 1: Noise Removal
#
# Start by removing noise from the text data. Removing noise is very task-specific, so you will usually pick and choose from the following to process your text data based on your needs:
# * Remove formatting (HTML, markup, metadata) -- e.g. emails, web-scrapped data
# * Extract text data from full dataset -- e.g. reviews, comments, labeled data from a nested JSON file or from structured data
# * Remove special characters
# * Remove emojis or convert emoji to words -- e.g. reviews, tweets, Instagram and Facebook comments, SMS text with sales
# * Remove URLs -- reviews, web content, emails
# * Convert accented characters to ASCII characters -- e.g. tweets, contents that may contain foreign language
#
# Note that pre-processing is an iterative process, so it is common to revisit any of these steps after you have cleaned and normalized your data.
#
# Here you will look at tweets and decide how you are going to process URL, emojis and emoticons.
#
# Working with text will often means dealing with regular expression. To freshen up on your regex or if you are new, [Pythex](https://pythex.org/) is a good helper page for you to find cheatsheet and test your functions.
# #### Noise Removal - Remove URLs
def remove_urls(text):
url = re.compile(r'https?://\S+|www\.\S+')
return url.sub(r'', text)
# Let's check if our code works with one example:
print (text_data['text'][0])
print ('Removed URL:' + remove_urls(text_data['text'][0]))
# #### Noise Removal - Remove emoticons, or convert emoticons to words
from emot.emo_unicode import UNICODE_EMO, EMOTICONS
def remove_emoticons(text):
"""
This function takes strings containing emoticons and returns strings with emoticons removed.
Input(string): one tweet, contains emoticons
Output(string): one tweet, emoticons removed, everything else unchanged
"""
emoticon = re.compile(u'(' + u'|'.join(k for k in EMOTICONS) + u')')
return emoticon.sub(r'', text)
def convert_emoticons(text):
"""
This function takes strings containing emoticons and convert the emoticons to words that describe the emoticon.
Input(string): one tweet, contains emoticons
Output(string): one tweet, emoticons replaced with words describing the emoticon
"""
for emot in EMOTICONS:
text = re.sub(u'('+emot+')', " ".join(EMOTICONS[emot].replace(",","").split()), text)
return text
# Let's check the results with one example and decide if we should keep the emoticon:
print ('original text: ' +remove_emoticons(text_data['text'][0]))
print ('removed emoticons: ' + convert_emoticons(text_data['text'][0]))
# Assuming our task is sentiment analysis, then converting emoticons to words will be helpful.
# We will apply our remove_URL and convert_emoticons functions to the full dataset:
text_data['cleaned_text'] = text_data['text'].apply(remove_urls).apply(convert_emoticons)
text_data[['text', 'cleaned_text']][:1]
# ## Step 2: Normalization
#
# In the next step, we will further process the text so that all text/words will be put on the same level playing field: all the words should be in the same case, numbers should be also treated as strings, abbreviations and chat words should be recognizable and replaced with the full words, etc. This is important because we do not want two elements in our word list (dictionary) with the same meaning are taken as two non-related different words by machine, and when we eventually convert all words to numbers (vectors), these words will be noises to our model, such as "3" and "three", "Our" and "our", or "urs" and "yours". This process often includes the following steps:
# ### Step 2.1 General Normalization
# * Convert all text to the same case
# * Remove punctuation
# * Convert numbers to word or remove numbers depending on your task
# * Remove white spaces
# * Convert abbreviations/slangs/chat words to word
# * Remove stop words (task specific and general English words); you can also create your own list of stop words
# * Remove rare words
# * Spelling correction
#
# **Note:** some normalization processes are better to perform at sentence and document level, and some processes are word-level and should happen after tokenization and segmentation, which we will cover right after normalization.
#
# Here you will convert the text to lower case, remove punctuation, remove numbers, remove white spaces, and complete other word-level processing steps after tokenizing the sentences.
# #### Normalization - Convert all text to lower case
# Usually, this is a must for all language pre-processing. Since "Word" and "word" will essentially be considered two different elements in word representation, and we want words that have the same meaning to be represented the same in numbers (vectors), we want to convert all text into the same case.
text_data['text_lower'] = text_data['cleaned_text'].str.lower()
text_data[['cleaned_text', 'text_lower']][:1]
# #### Normalization - Remove numbers
# Depending on your use cases, you can either remove numbers or convert numbers into strings. If numbers are not important in your task (e.g. sentiment analysis) you can remove those, and in some cases, numbers are useful (e.g. date), and you can tag these numbers differently. In most pre-trained embeddings, numbers are treated as strings.
#
# In this example, we are using Twitter data (tweets) and typically, numbers are not that important for understanding the meaning or content of a tweet. Therefore, we will remove the numbers.
def remove_numbers(text):
'''
This function takes strings containing numbers and returns strings with numbers removed.
Input(string): one tweet, contains numbers
Output(string): one tweet, numbers removed
'''
return re.sub(r'\d+', '', text)
#let's check the results of our function
remove_numbers(text_data['text_lower'][2])
text_data['normalized_text'] = text_data['text_lower'].apply(remove_numbers)
# #### Twitter data specific: Remove mentions or extract mentions into a different column
# We can remove the mentions in the tweets, but if our task is to monitor VOC, it is helpful to extract the mentions data.
def remove_mentions(text):
'''
This function takes strings containing mentions and returns strings with
mentions (@ and the account name) removed.
Input(string): one tweet, contains mentions
Output(string): one tweet, mentions (@ and the account name mentioned) removed
'''
mentions = re.compile(r'@\w+ ?')
return mentions.sub(r'', text)
print('original text: ' + text_data['text_lower'][0])
print('removed mentions: ' + remove_mentions(text_data['text_lower'][0]))
def extract_mentions(text):
'''
This function takes strings containing mentions and returns strings with
mentions (@ and the account name) extracted into a different element,
and removes the mentions in the original sentence.
Input(string): one sentence, contains mentions
Output:
one tweet (string): mentions (@ and the account name mentioned) removed
mentions (string): (only the account name mentioned) extracted
'''
mentions = [i[1:] for i in text.split() if i.startswith("@")]
sentence = re.compile(r'@\w+ ?').sub(r'', text)
return sentence,mentions
text_data['normalized_text'], text_data['mentions'] = zip(*text_data['normalized_text'].apply(extract_mentions))
text_data[['text','normalized_text', 'mentions']].head(1)
# #### Remove Punctuation
# We will use the `string.punctuation` in python to remove punctuations, which contains the following punctuation symbols`!"#$%&\'()*+,-./:;<=>?@[\\]^_{|}~`
# , you can add or remove more as needed.
punc_list = string.punctuation #you can self define list of punctuation to remove here
def remove_punctuation(text):
"""
This function takes strings containing self defined punctuations and returns
strings with punctuations removed.
Input(string): one tweet, contains punctuations in the self-defined list
Output(string): one tweet, self-defined punctuations removed
"""
translator = str.maketrans('', '', punc_list)
return text.translate(translator)
remove_punctuation(text_data['normalized_text'][2])
text_data['normalized_text'] = text_data['normalized_text'].apply(remove_punctuation)
# #### Remove Whitespaces
# You can also use `trim` functions to trim whitespaces from left and right or in the middle. Here we will just simply utilize the `split` function to extract all words from our text since we already removed all special characters, and combine them with a single whitespace.
def remove_whitespace(text):
'''
This function takes strings containing mentions and returns strings with
whitespaces removed.
Input(string): one tweet, contains whitespaces
Output(string): one tweet, white spaces removed
'''
return " ".join(text.split())
print('original text: ' + text_data['normalized_text'][2])
print('removed whitespaces: ' + remove_whitespace(text_data['normalized_text'][2]))
text_data['normalized_text'] = text_data['normalized_text'].apply(remove_whitespace)
# ## Step 3: Tokenization and Segmentation
#
# After we extracted useful text data from the full dataset, we will split large chunks of text (documents) into sentences, and sentences into words. Most of the times we will use sentence-ending punctuation to split documents into sentences, but it can be ambiguous especially when we are dealing with character conversations ("Are you alright?" said Ron), abbreviations (Dr. Fay would like to see <NAME> now.) and other special use cases. There are Python libraries designed for this task (check [textsplit](https://github.com/chschock/textsplit)), but you can take your own approach depending on your context.
#
# Here for Twitter data, we are only dealing with sentences shorter than 140 characters, so we will just tokenize sentences into words. We do want to normalize the sentence before tokenizing sentences into words, so we will introduce normalization, and tokenize our tweets into words after normalizing sentences.
# ### Tokenizing Sentences into Words
nltk.download('punkt')
from nltk.tokenize import word_tokenize
def tokenize_sent(text):
'''
This function takes strings (a tweet) and returns tokenized words.
Input(string): one tweet
Output(list): list of words tokenized from the tweet
'''
word_tokens = word_tokenize(text)
return word_tokens
text_data['tokenized_text'] = text_data['normalized_text'].apply(tokenize_sent)
text_data[['normalized_text','tokenized_text']][:1]
# ### Continuing Word-level Normalization
# #### Remove Stop Words
# Stop words are common words that does not contribute to the meaning of a sentence, such as 'the', 'a', 'his'. Most of the time we can remove these words without harming further analysis, but if you want to apply Part-of-Speech (POS) tagging later, be careful with what you removed in this step as they can provide valuable information. You can also add stop words to the list based on your use cases.
nltk.download('stopwords')
from nltk.corpus import stopwords
stopwords_list = set(stopwords.words('english'))
# One way to add words to your stopwords list is to check for most frequent words, especially if you are working with a domain-specific corpus and those words sometimes are not covered by general English stop words. You can also remove rare words from your text data.
#
# Let's check for the most common words in our data. All the words we see in the following example are covered in general English stop words, so we will not add any additional stop words.
from collections import Counter
counter = Counter()
for word in [w for sent in text_data["tokenized_text"] for w in sent]:
counter[word] += 1
counter.most_common(10)
# Let's check for the rarest words now. In this example, infrequently used words mostly consist of misspelled words, which we will later correct, but we can add them to our stop words list as well.
#least frequent words
counter.most_common()[:-10:-1]
top_n = 10
bottom_n = 10
stopwords_list |= set([word for (word, count) in counter.most_common(top_n)])
stopwords_list |= set([word for (word, count) in counter.most_common()[:-bottom_n:-1]])
stopwords_list |= {'thats'}
def remove_stopwords(tokenized_text):
'''
This function takes a list of tokenized words from a tweet, removes self-defined stop words from the list,
and returns the list of words with stop words removed
Input(list): a list of tokenized words from a tweet, contains stop words
Output(list): a list of words with stop words removed
'''
filtered_text = [word for word in tokenized_text if word not in stopwords_list]
return filtered_text
print(text_data['tokenized_text'][2])
print(remove_stopwords(text_data['tokenized_text'][2]))
text_data['tokenized_text'] = text_data['tokenized_text'].apply(remove_stopwords)
# #### Convert Abbreviations, slangs and chat words into words
# Sometimes you will need to develop your own mapping for abbreviations/slangs <-> words, for chat data, or for domain-specific data where abbreviations often have different meanings from what is commonly used.
chat_words_map = {
'idk': 'i do not know',
'btw': 'by the way',
'imo': 'in my opinion',
'u': 'you',
'oic': 'oh i see'
}
chat_words_list = set(chat_words_map)
def translator(text):
"""
This function takes a list of tokenized words, finds the chat words in the self-defined chat words list,
and replace the chat words with the mapped full expressions. It returns the list of tokenized words with
chat words replaced.
Input(list): a list of tokenized words from a tweet, contains chat words
Output(list): a list of words with chat words replaced by full expressions
"""
new_text = []
for w in text:
if w in set(chat_words_map):
new_text = new_text + chat_words_map[w].split()
else:
new_text.append(w)
return new_text
print(text_data['tokenized_text'][13])
print(translator(text_data['tokenized_text'][13]))
text_data['tokenized_text'] = text_data['tokenized_text'].apply(translator)
# #### Spelling Correction
# Some common spelling correction packages include `SpellChecker` and `autocorrect`. It might take some time to spell check every sentence of the text, so you can decide if a spell check is absolutely necessary. If you are dealing with documents (news, papers, articles) generally it is not necessary; but if you are dealing with chat data, reviews, notes, it might be a good idea to spell check your text.
from autocorrect import Speller
spell = Speller(lang='en', fast = True)
def spelling_correct(tokenized_text):
"""
This function takes a list of tokenized words from a tweet, spell check every words and returns the
corrected words if applicable. Note that not every wrong spelling words will be identified especially
for tweets.
Input(list): a list of tokenized words from a tweet, contains wrong-spelling words
Output(list): a list of corrected words
"""
corrected = [spell(word) for word in tokenized_text]
return corrected
print(text_data['tokenized_text'][0])
print(spelling_correct(text_data['tokenized_text'][0]))
text_data['tokenized_text'] = text_data['tokenized_text'].apply(spelling_correct)
# ### Step 3.2 [Stemming and Lemmatization](https://en.wikipedia.org/wiki/Lemmatisation)
# **Stemming** is the process of removing affixes from a word to get a word stem, and **lemmatization** can in principle select the appropriate lemma depending on the context. The difference is that a stemmer operates on a single word without knowledge of the context, and therefore cannot discriminate between words that have different meanings depending on part of speech. However, stemmers are typically easier to implement and run faster, and the reduced accuracy may not matter for some applications.
# #### Stemming
# There are several stemming algorithms available, and the most popular ones are Porter, Lancaster, and Snowball. Porter is the most common one, Snowball is an improvement over Porter, and Lancaster is more aggressive. You can check for more algorithms provided by `nltk` [here](https://www.nltk.org/api/nltk.stem.html).
# +
from nltk.stem import SnowballStemmer
from nltk.tokenize import word_tokenize
stemmer = SnowballStemmer("english")
def stem_text(tokenized_text):
"""
This function takes a list of tokenized words from a tweet, and returns the stemmed words by your
defined stemmer.
Input(list): a list of tokenized words from a tweet
Output(list): a list of stemmed words in its root form
"""
stems = [stemmer.stem(word) for word in tokenized_text]
return stems
# -
# #### Lemmatization
nltk.download('wordnet')
from nltk.stem import WordNetLemmatizer
from nltk.tokenize import word_tokenize
lemmatizer = WordNetLemmatizer()
def lemmatize_text(tokenized_text):
'''
This function takes a list of tokenized words from a tweet, and returns the lemmatized words.
you can also provide context for lemmatization, i.e. part-of-speech.
Input(list): a list of tokenized words from a tweet
Output(list): a list of lemmatized words in its base form
'''
lemmas = [lemmatizer.lemmatize(word, pos = 'v') for word in tokenized_text]
return lemmas
# #### Let's compare our stemming and lemmatization results:
# It seems like both processes returned similar results besides some verb being trimmed differently, so it is okay to go with stemming in this case if you are dealing with a lot of data and want a better performance. You can also keep both and experiment with further feature engineering and modeling to see which one produces better results.
print(text_data['tokenized_text'][2])
print(stem_text(text_data['tokenized_text'][2]))
print(lemmatize_text(text_data['tokenized_text'][2]))
# It seems that a stemmer can do the work for our tweets data. You can keep both and decide which one you want to use for feature engineering and modeling.
text_data['stem_text'] = text_data['tokenized_text'].apply(stem_text)
text_data['lemma_text'] = text_data['tokenized_text'].apply(lemmatize_text)
# ## Step 3.5: Re-examine the results
# Take a pause here and examine the results from previous steps to decide if more noise removal/normalization is needed. In this case, you might want to add more words to the stop words list, spell-check more aggressively, or add more mappings to the abbreviation/slang to words list.
text_data.sample(5)[['text', 'stem_text', 'lemma_text']]
# ## Step 4: Enrichment and Augmentation
# After you have cleaned and tokenized your text data into a standard form, you might want to enrich it with more useful information that was not provided directly in the original text or its single-word form. For example:
# * Part-of-speech tagging
# * Extracting phrases
# * Name entity recognition
# * Dependency parsing
# * Word level embeddings
#
# Many Python Packages including `nltk`, `SpaCy`, `CoreNLP`, and here we will use `TextBlob` to illustrate some enrichment methods.
#
# #### Part-of-Speech (POS) Tagging
# Part-of-Speech tagging can assign each word in accordance with its syntactic functions (noun, verb, adjectives, etc.).
nltk.download('averaged_perceptron_tagger')
text_example = text_data.sample()['lemma_text']
text_example
from textblob import TextBlob
result = TextBlob(" ".join(text_example.values[0]))
print(result.tags)
# #### Extracting Phrases
# Sometimes words come in as phrases (noun group phrases, verb group phrases, etc.) and often have discrete grammatical meanings. Extract those words as phrases rather than separate words in this case.
nltk.download('brown')
# orginal text:
text_example = text_data.sample()['lemma_text']
" ".join(text_example.values[0])
#noun phrases that can be extracted from this sentence
result = TextBlob(" ".join(text_example.values[0]))
for nouns in result.noun_phrases:
print(nouns)
# #### Name Entity Recognition (NER)
# You can use pre-trained/pre-defined name entity recognition models to find named entities in text and classify them into pre-defined categories. You can also train your own NER model, especially if you are dealing with domain specific context.
nltk.download('maxent_ne_chunker')
nltk.download('words')
text_example_enr = text_data.sample()['lemma_text'].values[0]
print("original text: " + " ".join(text_example_enr))
from nltk import pos_tag, ne_chunk
print (ne_chunk(pos_tag(text_example_enr)))
# ## Final Dataset ready for feature engineering and modeling
# For this notebook you cleaned and normalized the data, kept mentions as a separate column, and stemmed and lemmatized the tokenized words. You can experiment with these two results to see which one gives you a better model performance.
#
# Twitter data is short and often does not have complex syntax structures, so no enrichment (POS tagging, parsing, etc.) was done at this time; but you can experiment with those when you have more complicated text data.
text_data.head(2)
# ### Save our final dataset to S3 for further process
filename_write_to = 'processed_sentiment_140.csv'
text_data.to_csv(filename_write_to, index = False)
upload_to_s3(bucket, 'text_sentiment140_processed', filename_write_to)
# # Conclusion
#
# Congratulations! You cleaned and prepared your text data and it is now ready to be vectorized or used for feature engineering.
# Now that your data is ready to be converted into machine-readable format (numbers), we will cover extracting features and word embeddings in the next section **text data feature engineering**.
| preprocessing/text_data/04_preprocessing_text_data_v3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Spyder)
# language: python3
# name: python3
# ---
# # Transient Fickian Diffusion with Reaction
# `OpenPNM` supports adding reaction terms to both steady state and transient simulations. `OpenPNM` already includes many different source term models that can be added to simulate a reaction. In this example, we show how to add a `powerlaw` source term model to a transient fickian diffusion simulation.
# Start by importing openpnm
import openpnm as op
%config InlineBackend.figure_formats = ['svg']
import matplotlib.pyplot as plt
import numpy as np
ws = op.Workspace()
proj = ws.new_project()
np.random.seed(10)
# # Define network, geometry, and phase objects
shape = [25, 25, 1]
net = op.network.Cubic(shape=shape, spacing=1e-4)
geo = op.geometry.SpheresAndCylinders(network=net, pores=net.Ps, throats=net.Ts)
phase = op.phases.Water(network=net)
# You can adjust the diffusion coefficient by calling the `pore.diffusivity` dictionary key on the phase object and then setting it's value.
phase['pore.diffusivity'] = 2e-09
# # Define physics object
# Here, we will use `Standard` physics which already includes many standard models. You could also use `Generic` physics and add neccessary models like `throat.diffusive_conductance` after using `phys.add_model()` method.
phys = op.physics.Standard(network=net, phase=phase, geometry=geo)
# # Add reaction model
# Add reaction model to physics object. For this example we use the `powerlaw` reaction model which is of the form:
#
# \begin{equation*}
# \ A_1X^{A_2}
# \end{equation*}
phase['pore.concentration'] = 0
phys['pore.rxnA'] = -1e-10
phys['pore.rxnb'] = 1
phys.add_model(propname='pore.reaction', model=op.models.physics.generic_source_term.power_law,
A1='pore.rxnA', A2='pore.rxnb', X='pore.concentration')
# # Define transient fickian diffusion object
tfd = op.algorithms.TransientFickianDiffusion(network=net, phase=phase)
# # Setup the transient algorithm settings
# To do this you can call the `tfd.setup()` method and use it to set settings such as transient solver scheme, final time, time step, tolerance, etc. The `cranknicolson` scheme used here is the most accurate but slowest. Other time schemes are `implicit` which is faster but less accurate and `steady` which gives the steady state solution.
tfd.settings.update({'t_scheme': 'cranknicolson',
't_final': 1000,
't_output': 100,
't_step': 1,
't_tolerance': 1e-12})
# # Set value boundary conditions
# In this example we set the concentraton of the `front` pores to 1 and the concentration of the `back` pores to 0.
tfd.set_value_BC(pores=net.pores('front'), values=1)
tfd.set_value_BC(pores=net.pores('back'), values=0)
# # Set initial condition
# This command sets all pores to 0 (or whatever value you set) to start the simulation.
tfd.set_IC(values=0)
# # Set source term
# In this example, we apply the source term to pores 212 and 412.
tfd.set_source(propname='pore.reaction', pores=[212, 412])
# # Run the simulation
tfd.run()
# # Visualize results
# Ater simulation runs we can visualize results using a colour plot from `matplotlib.pyplot`. Here we visualize the results at the final time `t_final` by using the `pore.concentration` key on the algorithm object.
c = tfd['pore.concentration'].reshape(shape)
fig, ax = plt.subplots(figsize=(6, 6))
plt.imshow(c[:,:,0])
plt.title('Concentration (mol/m$^3$)')
plt.colorbar();
# If we print the `TransientFickianDiffusion` object we can see a list of the object's properties. Notice how the concentration is recorded here for each output concentration, `t_output`.
print(tfd)
# We can visualize intermediate concentration profiles using a colour plot but use `pore.concentration@100` (or similar) as the dictionary key on the algorithm object.
c = tfd['pore.concentration@100'].reshape(shape)
fig, ax = plt.subplots(figsize=(6, 6))
plt.imshow(c[:,:,0])
plt.title('Concentration (mol/m$^3$)')
plt.colorbar();
| examples/notebooks/algorithms/transient/transient_fickian_diffusion_with_reaction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
print("Four score and seven years ago our fathers brought forth on this continent, a new nation, conceived in Liberty, ")
print("and dedicated to the proposition that all men are created equal.")
print("Now we are engaged in a great civil war, testing whether that nation, or any nation so conceived and so dedicated, ")
print("can long endure. We are met on a great battle-field of that war. We have come to dedicate a portion of that ")
print("field, as a final resting place for those who here gave their lives that that nation might live. It is ")
print("altogether fitting and proper that we should do this.")
print("But, in a larger sense, we can not dedicate -- we can not consecrate -- we can not hallow -- this ground. ")
print("The brave men, living and dead, who struggled here, have consecrated it, far above our poor power to add or detract. ")
print("The world will little note, nor long remember what we say here, but it can never forget what they did here. ")
print("It is for us the living, rather, to be dedicated here to the unfinished work which they who fought here have ")
print("thus far so nobly advanced. It is rather for us to be here dedicated to the great task remaining before us -- ")
print("that from these honored dead we take increased devotion to that cause for which they gave the last full measure ")
print("of devotion -- that we here highly resolve that these dead shall not have died in vain -- that this nation, ")
print("under God, shall have a new birth of freedom -- and that government of the people, by the people, for the people, ")
print("shall not perish from the earth.")
| tests/cells-not-in-order.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Python and APIs
# Angela, Moritz & Thomas
# + [markdown] slideshow={"slide_type": "slide"}
# ### Outline of Session 5
#
# In these slides
# * Introduction
# * JSON
# * Hardcoding access with `requests`
# * Popular pre-written modules for geocoding and mapping
#
# External scripts
# * Trade data with Comtrade API (kudos Federico)
# * The Twitter API (kudos Charles)
# + [markdown] slideshow={"slide_type": "slide"}
# # What is an API?
#
# ## API: Application programming interface
#
# _"a set of clearly defined methods of communication between various software components."_
#
#
# - Pre-internet days: Extension of software beyond its usual capabilities
# - Nowadays: Interface by web service providers for you to connect/retrieve with your own application (i.e. without going through GUI)
# - You send data <-> You get data back
# - Most APIs have a similar structure: the REST architecture (REpresentational State Transfer)
# - Inititatives like the Linux Foundation's [OpenAPI](https://www.openapis.org/) develop these types of standards
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Illustration: General Set-up
#
# <img src="Slide1.png" width="800" height="450">
#
# Source: [MuleSoft Videos](https://www.youtube.com/embed/s7wmiS2mSXY)
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Illustration: Making a request
#
# <img src="Slide2.png" width="800" height="450">
# + [markdown] slideshow={"slide_type": "slide"}
# # Illustration: Returning Content
#
# <img src="Slide3.png" width="800" height="450">
# + [markdown] slideshow={"slide_type": "slide"}
# # Elements of an API
#
# * A protocol (eg: https)
# * A server (eg: httpbin.org)
# * A method name / location (eg: /get)
# * A set of arguments (eg: hello=world and foo=bar)
#
# ➥ https://httpbin.org/get?hello=world&foo=bar
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# # The Request
#
# * You need to read the documentation (e.g. [Eurostat API documentation](https://ec.europa.eu/eurostat/web/json-and-unicode-web-services/getting-started/rest-request))
# * You need to specify the URL (if it is a remote API)
# * You need to import the `requests` module in Python
# + [markdown] slideshow={"slide_type": "slide"}
# # The returning object
#
# * Usually it is encoded in JSON
# * The other popular formats for data structure are XML (mainly old stuff) and CSV (spreadsheet compatible)
# + [markdown] slideshow={"slide_type": "slide"}
# # What is a JSON?
# * JSON = [(JavaScript Object Notation)](https://www.json.org/)
# * It is one of the most popular format for data in the world
# * It looks like a Python dictionary, except for the fact that:
# - JSON is a string (it is inside a text file)
# - JSON must use double quotation mark
# - in JavaScript it is defined as Object
# * In Python there is a built-in module called `json` module
# * It follows this strcture {"key" : "value"}; see example below:
#
#
# + slideshow={"slide_type": "subslide"}
{
"friends": [
{
"name": "Jose",
"degree": "Applied Computing"
},
{
"name": "Rolf",
"degree": "Computer Science"
},
{
"name": "Anna",
"degree": "Physics"
}
]
}
# + [markdown] slideshow={"slide_type": "slide"}
# # What can I do with APIs?
#
# - Retrieve [World Bank Dev Indicators](https://datahelpdesk.worldbank.org/knowledgebase/articles/898581-api-basic-call-structures)
# - Track stock prices with [OpenFIGI](https://www.openfigi.com/api)
# - Geocode an address with [Here Maps](https://developer.here.com/)
# - Convert fiat currency with [opencurrency](https://openexchangerates.org/) or crypto with [alternative.me](https://alternative.me/crypto/api/)
# - Send tweets with [Twitter](https://developer.twitter.com/)
# - Download images from Mars or the Moon from [NASA](https://api.nasa.gov/)
# - ...
# + [markdown] slideshow={"slide_type": "slide"}
#
# <a href="https://imgflip.com/i/3rrkeh"><img src="https://i.imgflip.com/3rrkeh.jpg" title="madeatimgflip.com"/></a>
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# # The Starwars Character API
#
# - Let's get data on Starwars characters!
# - We find [the documentation](https://swapi.co/documentation)
# - We import the `requests` module
# - We set the base URL of the API: `https://swapi.co/api/`
#
#
# + slideshow={"slide_type": "subslide"}
import requests
url = "https://swapi.co/api/"
# Launch request
base_req = requests.get(url)
print(base_req)
# + slideshow={"slide_type": "subslide"}
# Explore json content of object
print(base_req.json())
# + [markdown] slideshow={"slide_type": "slide"}
# # Making sense of the answer
#
# - The call to the basic URL returns various subfields
# `
# {
# 'people': 'https://swapi.co/api/people/',
# 'planets': 'https://swapi.co/api/planets/',
# 'films': 'https://swapi.co/api/films/',
# 'species': 'https://swapi.co/api/species/',
# 'vehicles': 'https://swapi.co/api/vehicles/',
# 'starships': 'https://swapi.co/api/starships/'
# }
# `
# - The documentation adds: `/people/:id/ -- get a specific people resource`
# - Let's request info on the first id of the subfield `people`
# - We can request further info on a sub-subfield (e.g. `starships`)
#
#
#
#
# + slideshow={"slide_type": "subslide"}
# Let's try
url = "https://swapi.co/api/people/1"
# Launch request
req_1 = requests.get(url)
# Extract json
req_1_js = req_1.json()
print(req_1_js)
# + slideshow={"slide_type": "subslide"}
# Let's extract Luke's starships
ships = req_1_js.get("starships")
print(ships)
# And get entry of first starship
ship = requests.get(ships[0]).json()
print(ship)
# + [markdown] slideshow={"slide_type": "slide"}
# # Geocoding
#
# - Most map services like Google Maps, Open Street Maps, Bing, etc. have APIs
# - Let's check out Open Street Maps' (OSM) API called [Nominatim](https://developer.mapquest.com/)
# - We will use the API through the Python module `geopy`
# - To install new modules check out this [tutorial](https://jakevdp.github.io/blog/2017/12/05/installing-python-packages-from-jupyter/)
# - But we still need to sign up with our email to obtain a key for Nominatim
#
#
#
#
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Retriving a location
#
# * We install `geopy`
# * We sign up and obtain the key (in this example we use Moritz's key)
# * We search the location of two popular places among ULB students: the ULB and the bar Tavernier
# * We store and print out their coordinates
# + slideshow={"slide_type": "subslide"}
#import sys
# #!{sys.executable} -m pip install geopy
from geopy.geocoders import Nominatim
# Replace the user agent key with your own key
geolocator = Nominatim(user_agent="jjG4qnPniTAGpG7O0q8XcMhARm0Pxcln")
location_ULB = geolocator.geocode("Universite libre de Bruxelles")
location_tav = geolocator.geocode("Bar Tavernier Ixelles")
# Did it work?
print(location_ULB)
print(location_tav)
# Like a charm...
# Store coordinates in tuple
ulb_coords = [location_ULB.latitude, location_ULB.longitude]
print(ulb_coords)
tav_coords = [location_tav.latitude, location_tav.longitude]
print(tav_coords)
# + [markdown] slideshow={"slide_type": "slide"}
# # Mapping
#
# - Sadly, not everybody can associate a place to its coordinates in their mind :-P
# - With the free [Leaflet API](https://leafletjs.com/reference-1.6.0.html) we can pin point them on a map
# - Unfortunately written in Java Script
# - But no need to learn, use "wrapper" Python Module `Folium`
# - Example: Map of environmental violations of shale gas drilling in Pennsyilvania ([website](http://stateimpact.npr.org/pennsylvania/drilling/violations/))
#
# <img src="map.png" alt="Map" width="250" height="250">
#
#
#
#
#
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Producing a map
#
# * We install `Folium`
# * We create a map for the ULB, specifying the location and a zoom level
# * We display the map
# + slideshow={"slide_type": "subslide"}
# Install Python module Folium, a wrapper of the Leaflet API
#import sys
# #!{sys.executable} -m pip install folium
import folium
from folium.plugins import MarkerCluster
import pandas as pd
#Create the map
my_map = folium.Map(location = ulb_coords, zoom_start = 14)
# + slideshow={"slide_type": "subslide"}
# Display map
my_map
# + [markdown] slideshow={"slide_type": "slide"}
# # Adding markers to the map
#
# * It is still not clear where the ULB and Tavernier are located: we need some markers!
# * We pin point them on the map by adding some representative icons ;-)
# + slideshow={"slide_type": "subslide"}
# Let's add markers
folium.Marker(ulb_coords, popup = 'ULB', icon = folium.Icon(icon='book')).add_to(my_map)
folium.Marker(tav_coords, popup = 'STATA User Meeting', icon = folium.Icon(icon='glass', color='red')).add_to(my_map)
my_map
| Session_5/Session 5 - APIs Intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# name: python3
# ---
# # 语义成分分解
# ## 导入模块
from EduNLP.utils import dict2str4sif
# ## 测试数据
item = {
"stem": r"若复数$z=1+2 i+i^{3}$,则$|z|=$",
"options": ['0', '1', r'$\sqrt{2}$', '2'],
}
item
# ## 区分题目成分
# - 通过添加特殊关键词的方式,区分题目的题干和选项,选项之间按顺序做编号标记。
dict2str4sif(item,key_as_tag=True,
add_list_no_tag=False,
# keys=["options"],
tag_mode="head"
)
dict2str4sif(item, add_list_no_tag=True)
dict2str4sif(item, add_list_no_tag=False)
dict2str4sif(item, tag_mode="head")
dict2str4sif(item, tag_mode="tail")
dict2str4sif(item, key_as_tag=False)
| examples/utils/data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="Tce3stUlHN0L"
# ##### Copyright 2019 The TensorFlow Authors.
#
#
# + colab={} colab_type="code" id="tuOe1ymfHZPu" cellView="form"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="MfBg1C5NB3X0"
# # Multi-worker training with Keras
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/distribute/multi_worker_with_keras"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/distribute/multi_worker_with_keras.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/distribute/multi_worker_with_keras.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/distribute/multi_worker_with_keras.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="xHxb-dlhMIzW"
# ## Overview
#
# This tutorial demonstrates multi-worker distributed training with Keras model using `tf.distribute.Strategy` API. With the help of the strategies specifically designed for multi-worker training, a Keras model that was designed to run on single-worker can seamlessly work on multiple workers with minimal code change.
#
# [Distributed Training in TensorFlow](../../guide/distributed_training.ipynb) guide is available for an overview of the distribution strategies TensorFlow supports for those interested in a deeper understanding of `tf.distribute.Strategy` APIs.
#
#
# + [markdown] colab_type="text" id="MUXex9ctTuDB"
# ## Setup
#
# First, setup TensorFlow and the necessary imports.
# + colab={} colab_type="code" id="IqR2PQG4ZaZ0"
from __future__ import absolute_import, division, print_function, unicode_literals
# + colab={} colab_type="code" id="bnYxvfLD-LW-"
try:
# # %tensorflow_version only exists in Colab.
# !pip install tf-nightly
except Exception:
pass
import tensorflow_datasets as tfds
import tensorflow as tf
tfds.disable_progress_bar()
# + [markdown] colab_type="text" id="hPBuZUNSZmrQ"
# ## Preparing dataset
#
# Now, let's prepare the MNIST dataset from [TensorFlow Datasets](https://www.tensorflow.org/datasets). The [MNIST dataset](http://yann.lecun.com/exdb/mnist/) comprises 60,000
# training examples and 10,000 test examples of the handwritten digits 0–9,
# formatted as 28x28-pixel monochrome images.
# + colab={} colab_type="code" id="dma_wUAxZqo2"
BUFFER_SIZE = 10000
BATCH_SIZE = 64
def make_datasets_unbatched():
# Scaling MNIST data from (0, 255] to (0., 1.]
def scale(image, label):
image = tf.cast(image, tf.float32)
image /= 255
return image, label
datasets, info = tfds.load(name='mnist',
with_info=True,
as_supervised=True)
return datasets['train'].map(scale).cache().shuffle(BUFFER_SIZE)
train_datasets = make_datasets_unbatched().batch(BATCH_SIZE)
# + [markdown] colab_type="text" id="o87kcvu8GR4-"
# ## Build the Keras model
# Here we use `tf.keras.Sequential` API to build and compile a simple convolutional neural networks Keras model to train with our MNIST dataset.
#
# Note: For a more comprehensive walkthrough of building Keras model, please see [TensorFlow Keras Guide](https://www.tensorflow.org/guide/keras#sequential_model).
#
# + colab={} colab_type="code" id="aVPHl0SfG2v1"
def build_and_compile_cnn_model():
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(32, 3, activation='relu', input_shape=(28, 28, 1)),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(10)
])
model.compile(
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
optimizer=tf.keras.optimizers.SGD(learning_rate=0.001),
metrics=['accuracy'])
return model
# + [markdown] colab_type="text" id="2UL3kisMO90X"
# Let's first try training the model for a small number of epochs and observe the results in single worker to make sure everything works correctly. You should expect to see the loss dropping and accuracy approaching 1.0 as epoch advances.
# + colab={} colab_type="code" id="6Qe6iAf5O8iJ"
single_worker_model = build_and_compile_cnn_model()
single_worker_model.fit(x=train_datasets, epochs=3, steps_per_epoch=5)
# + [markdown] colab_type="text" id="8YFpxrcsZ2xG"
# ## Multi-worker Configuration
#
# Now let's enter the world of multi-worker training. In TensorFlow, `TF_CONFIG` environment variable is required for training on multiple machines, each of which possibly has a different role. `TF_CONFIG` is used to specify the cluster configuration on each worker that is part of the cluster.
#
# There are two components of `TF_CONFIG`: `cluster` and `task`. `cluster` provides information about the training cluster, which is a dict consisting of different types of jobs such as `worker`. In multi-worker training, there is usually one `worker` that takes on a little more responsibility like saving checkpoint and writing summary file for TensorBoard in addition to what a regular `worker` does. Such worker is referred to as the 'chief' worker, and it is customary that the `worker` with `index` 0 is appointed as the chief `worker` (in fact this is how `tf.distribute.Strategy` is implemented). `task` on the other hand provides information of the current task.
#
# In this example, we set the task `type` to `"worker"` and the task `index` to `0`. This means the machine that has such setting is the first worker, which will be appointed as the chief worker and do more work than other workers. Note that other machines will need to have `TF_CONFIG` environment variable set as well, and it should have the same `cluster` dict, but different task `type` or task `index` depending on what the roles of those machines are.
#
# For illustration purposes, this tutorial shows how one may set a `TF_CONFIG` with 2 workers on `localhost`. In practice, users would create multiple workers on external IP addresses/ports, and set `TF_CONFIG` on each worker appropriately.
#
# Warning: Do not execute the following code in Colab. TensorFlow's runtime will attempt to create a gRPC server at the specified IP address and port, which will likely fail.
#
# ```
# os.environ['TF_CONFIG'] = json.dumps({
# 'cluster': {
# 'worker': ["localhost:12345", "localhost:23456"]
# },
# 'task': {'type': 'worker', 'index': 0}
# })
# ```
#
#
# + [markdown] colab_type="text" id="P94PrIW_kSCE"
# Note that while the learning rate is fixed in this example, in general it may be necessary to adjust the learning rate based on the global batch size.
# + [markdown] colab_type="text" id="UhNtHfuxCGVy"
# ## Choose the right strategy
#
# In TensorFlow, distributed training consists of synchronous training, where the steps of training are synced across the workers and replicas, and asynchronous training, where the training steps are not strictly synced.
#
# `MultiWorkerMirroredStrategy`, which is the recommended strategy for synchronous multi-worker training, will be demonstrated in this guide.
# To train the model, use an instance of `tf.distribute.experimental.MultiWorkerMirroredStrategy`. `MultiWorkerMirroredStrategy` creates copies of all variables in the model's layers on each device across all workers. It uses `CollectiveOps`, a TensorFlow op for collective communication, to aggregate gradients and keep the variables in sync. The [`tf.distribute.Strategy` guide](../../guide/distributed_training.ipynb) has more details about this strategy.
# + colab={} colab_type="code" id="1uFSHCJXMrQ-"
strategy = tf.distribute.experimental.MultiWorkerMirroredStrategy()
# + [markdown] colab_type="text" id="N0iv7SyyAohc"
# Note: `TF_CONFIG` is parsed and TensorFlow's GRPC servers are started at the time `MultiWorkerMirroredStrategy.__init__()` is called, so `TF_CONFIG` environment variable must be set before a `tf.distribute.Strategy` instance is created.
# + [markdown] colab_type="text" id="FMy2VM4Akzpr"
# `MultiWorkerMirroredStrategy` provides multiple implementations via the [`CollectiveCommunication`](https://github.com/tensorflow/tensorflow/blob/a385a286a930601211d78530734368ccb415bee4/tensorflow/python/distribute/cross_device_ops.py#L928) parameter. `RING` implements ring-based collectives using gRPC as the cross-host communication layer. `NCCL` uses [Nvidia's NCCL](https://developer.nvidia.com/nccl) to implement collectives. `AUTO` defers the choice to the runtime. The best choice of collective implementation depends upon the number and kind of GPUs, and the network interconnect in the cluster.
# + [markdown] colab_type="text" id="H47DDcOgfzm7"
# ## Train the model with MultiWorkerMirroredStrategy
#
# With the integration of `tf.distribute.Strategy` API into `tf.keras`, the only change you will make to distribute the training to multi-worker is enclosing the model building and `model.compile()` call inside `strategy.scope()`. The distribution strategy's scope dictates how and where the variables are created, and in the case of `MultiWorkerMirroredStrategy`, the variables created are `MirroredVariable`s, and they are replicated on each of the workers.
#
# Note: Currently there is a limitation in `MultiWorkerMirroredStrategy` where TensorFlow ops need to be created after the instance of strategy is created. If you see `RuntimeError: Collective ops must be configured at program startup`, try creating the instance of `MultiWorkerMirroredStrategy` at the beginning of the program and put the code that may create ops after the strategy is instantiated.
#
# Note: In this Colab, the following code can run with expected result, but however this is effectively single-worker training since `TF_CONFIG` is not set. Once you set `TF_CONFIG` in your own example, you should expect speed-up with training on multiple machines.
#
# Note: Since `MultiWorkerMirroredStrategy` does not support last partial batch handling, pass the `steps_per_epoch` argument to `model.fit()` when dataset is imbalanced on multiple workers.
# + colab={} colab_type="code" id="BcsuBYrpgnlS"
NUM_WORKERS = 2
# Here the batch size scales up by number of workers since
# `tf.data.Dataset.batch` expects the global batch size. Previously we used 64,
# and now this becomes 128.
GLOBAL_BATCH_SIZE = 64 * NUM_WORKERS
with strategy.scope():
# Creation of dataset, and model building/compiling need to be within
# `strategy.scope()`.
train_datasets = make_datasets_unbatched().batch(GLOBAL_BATCH_SIZE)
multi_worker_model = build_and_compile_cnn_model()
# Keras' `model.fit()` trains the model with specified number of epochs and
# number of steps per epoch. Note that the numbers here are for demonstration
# purposes only and may not sufficiently produce a model with good quality.
multi_worker_model.fit(x=train_datasets, epochs=3, steps_per_epoch=5)
# + [markdown] colab_type="text" id="Rr14Vl9GR4zq"
# ### Dataset sharding and batch size
# In multi-worker training, sharding data into multiple parts is needed to ensure convergence and performance. However, note that in above code snippet, the datasets are directly sent to `model.fit()` without needing to shard; this is because `tf.distribute.Strategy` API takes care of the dataset sharding automatically in multi-worker trainings.
#
# If you prefer manual sharding for your training, automatic sharding can be turned off via `tf.data.experimental.DistributeOptions` api. Concretely,
# + colab={} colab_type="code" id="JxEtdh1vH-TF"
options = tf.data.Options()
options.experimental_distribute.auto_shard_policy = tf.data.experimental.AutoShardPolicy.OFF
train_datasets_no_auto_shard = train_datasets.with_options(options)
# + [markdown] colab_type="text" id="NBCtYvmCH-7g"
# Another thing to notice is the batch size for the `datasets`. In the code snippet above, we use `GLOBAL_BATCH_SIZE = 64 * NUM_WORKERS`, which is `NUM_WORKERS` times as large as the case it was for single worker, because the effective per worker batch size is the global batch size (the parameter passed in `tf.data.Dataset.batch()`) divided by the number of workers, and with this change we are keeping the per worker batch size same as before.
# + [markdown] colab_type="text" id="XVk4ftYx6JAO"
# ## Performance
#
# You now have a Keras model that is all set up to run in multiple workers with `MultiWorkerMirroredStrategy`. You can try the following techniques to tweak performance of multi-worker training.
#
# * `MultiWorkerMirroredStrategy` provides multiple [collective communication implementations](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/distribute/cross_device_ops.py). `RING` implements ring-based collectives using gRPC as the cross-host communication layer. `NCCL` uses [Nvidia's NCCL](https://developer.nvidia.com/nccl) to implement collectives. `AUTO` defers the choice to the runtime. The best choice of collective implementation depends upon the number and kind of GPUs, and the network interconnect in the cluster. To override the automatic choice, specify a valid value to the `communication` parameter of `MultiWorkerMirroredStrategy`'s constructor, e.g. `communication=tf.distribute.experimental.CollectiveCommunication.NCCL`.
# * Cast the variables to `tf.float` if possible. The official ResNet model includes [an example](https://github.com/tensorflow/models/blob/8367cf6dabe11adf7628541706b660821f397dce/official/resnet/resnet_model.py#L466) of how this can be done.
#
#
#
#
# + [markdown] colab_type="text" id="97WhAu8uKw3j"
# ## Fault tolerance
#
# In synchronous training, the cluster would fail if one of the workers fails and no failure-recovery mechanism exists. Using Keras with `tf.distribute.Strategy` comes with the advantage of fault tolerance in cases where workers die or are otherwise unstable. We do this by preserving training state in the distributed file system of your choice, such that upon restart of the instance that previously failed or preempted, the training state is recovered.
#
# Since all the workers are kept in sync in terms of training epochs and steps, other workers would need to wait for the failed or preempted worker to restart to continue.
#
# ### ModelCheckpoint callback
#
# To take advantage of fault tolerance in multi-worker training, provide an instance of `tf.keras.callbacks.ModelCheckpoint` at the `tf.keras.Model.fit()` call. The callback will store the checkpoint and training state in the directory corresponding to the `filepath` argument to `ModelCheckpoint`.
# + colab={} colab_type="code" id="xIY9vKnUU82o"
# Replace the `filepath` argument with a path in the file system
# accessible by all workers.
callbacks = [tf.keras.callbacks.ModelCheckpoint(filepath='/tmp/keras-ckpt')]
with strategy.scope():
multi_worker_model = build_and_compile_cnn_model()
multi_worker_model.fit(x=train_datasets,
epochs=3,
steps_per_epoch=5,
callbacks=callbacks)
# + [markdown] colab_type="text" id="Ii6VmEdOjkZr"
# If a worker gets preempted, the whole cluster pauses until the preempted worker is restarted. Once the worker rejoins the cluster, other workers will also restart. Now, every worker reads the checkpoint file that was previously saved and picks up its former state, thereby allowing the cluster to get back in sync. Then the training continues.
#
# If you inspect the directory containing the `filepath` you specified in `ModelCheckpoint`, you may notice some temporarily generated checkpoint files. Those files are needed for recovering the previously lost instances, and they will be removed by the library at the end of `tf.keras.Model.fit()` upon successful exiting of your multi-worker training.
# + [markdown] colab_type="text" id="ega2hdOQEmy_"
# ## See also
# 1. [Distributed Training in TensorFlow](https://www.tensorflow.org/guide/distributed_training) guide provides an overview of the available distribution strategies.
# 2. [Official models](https://github.com/tensorflow/models/tree/master/official), many of which can be configured to run multiple distribution strategies.
#
| site/en/tutorials/distribute/multi_worker_with_keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py37
# language: python
# name: py37
# ---
import torch; import torch.nn as nn
import sys
sys.path.append('../')
from torchdyn.models import *
# +
import torch
import torch.nn as nn
class HybridNeuralDE(nn.Module):
def __init__(self, jump, flow, readout, s_spans=None, residual=True, reverse=False):
super().__init__()
self.flow, self.jump, self.readout = flow, jump, readout
self.reverse, self.residual = reverse, residual
self.s_spans = s_spans
def forward(self, x):
"s_spans: iterable "
if self.s_spans:
assert len(self.s_spans) == x.shape[0]-1
h = self._init_hidden(x)
Y = torch.zeros_like(x)
if self.reverse: x_t = x_t.flip(0)
for i, x_t in enumerate(x):
h = self.jump(x_t, h) + h if self.residual else self.jump(x_t, h)
if self.s_spans: self.flow.s_span = self.s_spans[i]
h = self.flow(h)
Y[i] = h
return self.readout(Y[-1])
def _init_hidden(self, x):
# determine shape of hidden `h` based on x
if not hasattr(self, 'hidden_dim'):
self.hidden_dim = x.shape[2:] # L, B, [...]
return torch.zeros((x.shape[1], *self.hidden_dim)).to(x.device) # B, [...]
def hidden_trajectory(self, x, s_spans):
h = self._init_hidden(x)
# find global num mesh points across all s_spans
mesh = sum(map(len, s_spans))
Y = torch.zeros((mesh, *x.shape[1:]))
if self.reverse: x_t = x_t.flip(0)
for i, x_t in enumerate(x):
h = self.jump(x_t, h) + h if self.residual else self.jump(x_t, h)
h = self.flow.trajectory(h, s_spans[i])
print(Y.shape, h.shape)
Y[i] = h
return torch.readout(Y)
class NeuralCDE(nn.Module):
def __init__(self):
super().__init__()
# pass
# def forward(self, x):
# pass
# -
s1, s2 = torch.linspace(0, 1, 100), torch.linspace(0, 2, 10)
# +
f = nn.Linear(2, 2)
flow = NeuralODE(f)
jump = nn.RNNCell(2, 2)
readout = nn.Linear(2, 2)
net = HybridNeuralDE(jump, flow, readout)
# -
x0 = torch.randn(10, 2, 2)
s_spans = [torch.linspace(0, 1, i+1) for i in range(10)]
net.hidden_trajectory(x0, s_spans)
| test/benchmark/hybrid_dev/hybrid_ver2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 0. Web APIを用いた目次情報の取得
#
# [文化庁メディア芸術データベース マンガ分野 WebAPI](https://mediaarts-db.bunka.go.jp/webapi_proto_documents.pdf)を用いて,分析に必要なデータを入手します.なお,python3を使ったweb APIの利用については,[Python3でjsonを返却するwebAPIにアクセスして結果を出力するまで](http://qiita.com/sakura1116/items/4a11a9f7db9f535397fa)を参考にさせて頂きました.
#
# ## 環境構築
#
# ```bash
# conda env create -f env.yml
# ```
#
# ## 準備
import json
import urllib.request
from time import sleep
# ## 雑誌巻号検索結果の取得
#
# 以下の関数`search_magazine()`を使って,週刊少年ジャンプの雑誌巻号情報を検索し,結果を変数`magazines`に保存します.ここで取得したユニークIDは,次節の「雑誌巻号情報の取得」に必要になります.
def search_magazine(key='JUMPrgl', n_pages=25):
"""
「ユニークID」「雑誌巻号ID」あるいは「雑誌コード」にkey含む雑誌を,
n_pages分取得する関数です.
"""
url = 'https://mediaarts-db.bunka.go.jp/mg/api/v1/results_magazines?id=' + \
key + '&page='
magazines = []
for i in range(1, n_pages):
response = urllib.request.urlopen(url + str(i))
content = json.loads(response.read().decode('utf8'))
magazines.extend(content['results'])
return magazines
# Web APIでは,パラメータ`id`で「ユニークID」「雑誌巻号ID」あるいは「雑誌コード」を,`page`で検索結果の取得ページ番号(1ページあたり100件,デフォルトは1)を指定することができます.ここで,週刊少年ジャンプは「雑誌巻号ID」に`JUMPrgl`を含むため,`id=JUMPrgl`を指定します.また,週刊少年ジャンプの検索結果は合計24ページ(2320件)あるため,`page`に1から24を順次指定する必要があります.
# 詳細は[WebAPI仕様](https://mediaarts-db.bunka.go.jp/webapi_proto_documents.pdf)をご参照ください.
#
# なお,2017年3月31日より文化庁メディア芸術データベースのURLが変更されたため,[WebAPI仕様](https://mediaarts-db.bunka.go.jp/webapi_proto_documents.pdf)に記載のリクエストURL(`https://mediaarts-db.jp/mg/api/v1/results_magazines`)ではなく,新しいURL(`https://mediaarts-db.bunka.go.jp/mg/api/v1/results_magazines`)を使用する必要があることにご注意ください.
magazines = search_magazine()
len(magazines)
magazines[0]
magazines[-1]
# ## 雑誌巻号情報の取得
#
# 以下の関数`extract_data()`で必要な目次情報を抽出し,`save_data()`で目次情報を保存します.
def extract_data(content):
"""
contentに含まれる目次情報を取得する関数です.
- year: 発行年
- no: 号数
- title: 作品名
- author: 著者
- color: カラーか否か
- pages: 掲載ページ数
- start_page: 作品のスタートページ
- best: 巻頭から数えた掲載順
- worst: 巻末から数えた掲載順
"""
# マンガ作品のみ抽出します.
comics = [comic for comic in content['contents']
if comic['category']=='マンガ作品']
data = []
year = int(content['basics']['date_indication'][:4])
# 号数が記載されていない場合があるので,例外処理が必要です.
try:
no = int(content['basics']['number_indication'])
except ValueError:
no = content['basics']['number_indication']
for comic in comics:
title= comic['work']
if not title:
continue
# ページ数が記載されていない作品があるので,例外処理が必要です.
# 特に理由はないですが,無記載の作品は10ページとして処理を進めます.
try:
pages = int(comic['work_pages'])
except ValueError:
pages = 10
# 「いぬまるだしっ」等,1週に複数話掲載されている作品に対応するため
# data中にすでにtitleが含まれる場合は,新規datumとして登録せずに,
# 既存のdatumのページ数のみ加算します.
if len(data) > 0 and title in [datum['title'] for datum in data]:
data[[datum['title'] for datum in
data].index(title)]['pages'] += pages
else:
data.append({
'year': year,
'no': no,
'title': comic['work'],
'author': comic['author'],
'subtitle': comic['subtitle'],
'color': comic['note'].count('カラー'),
'pages': int(comic['work_pages']),
'start_pages': int(comic['start_page'])
})
# 企画物のミニマンガを除外するため,合計5ページ以下のdatumはリストから除外します.
filterd_data = [datum for datum in data if datum['pages'] > 5]
for n, datum in enumerate(filterd_data):
datum['best'] = n + 1
datum['worst'] = len(filterd_data) - n
return filterd_data
# 泥臭い話ですが,一部のギャグ漫画の扱いに苦労しました.例えば,「[いぬまるだしっ](https://mediaarts-db.bunka.go.jp/mg/magazine_works/545?ids%5B%5D=545)」は,基本的に一週間に2話ずつ掲載していましたが,データベースでは各話が別々の行に記載されています.これらを1つの作品として見なす必要があるので,当該`comic`の`title`が`data`中にある場合は,別`datum`として`data`に追加せず,既存の`datum`の`pages`を加算する処理を行っています.また,例えば「[ピューと吹く!ジャガー](https://mediaarts-db.bunka.go.jp/mg/comic_works/83870)」は,その人気に関係なく(実際めちゃくちゃ面白かったです),連載中は常に雑誌の最後に掲載されていました.これを外れ値として除外するかどうかで悩みましたが,結局残すことにしました.
def save_data(magazines, offset=0, file_name='data/wj-api.json'):
"""
magazinesに含まれる全てのmagazineについて,先頭からoffset以降の巻号の
目次情報を取得し,file_nameに保存する関数です.
"""
url = 'https://mediaarts-db.bunka.go.jp/mg/api/v1/magazine?id='
# ファイル先頭行
if offset == 0:
with open(file_name, 'w') as f:
f.write('[\n')
with open(file_name, 'a') as f:
# magazines中のmagazine毎にWeb APIを叩きます.
for m, magazine in enumerate(magazines[offset:]):
response = urllib.request.urlopen(url + str(magazine['id']),
timeout=30)
content = json.loads(response.read().decode('utf8'))
# 前記の関数extract_data()で,必要な情報を抽出します.
comics = extract_data(content)
print('{0:4d}/{1}: Extracted data from {2}'.\
format(m + offset, len(magazines), url + str(magazine['id'])))
# comics中の各comicについて,file_nameに情報を保存します.
for n, comic in enumerate(comics):
# ファイル先頭以外の,magazineの最初のcomicの場合は,
# まず',\n'を追記.
if m + offset > 0 and n == 0:
f.write(',\n')
json.dump(comic, f, ensure_ascii=False)
# 最後のcomic以外は',\n'を追記.
if not n == len(comics) - 1:
f.write(',\n')
print('{0:9}: Saved data to {1}'.format(' ', file_name))
# サーバへの負荷を抑えるため,必ず一時停止します.
sleep(3)
# ファイル最終行
with open(file_name, 'a') as f:
f.write(']')
# タイムアウトに柔軟に対応するため,目次情報を一括処理せず,無理やり逐次処理にしています.また,サーバに負荷をかけないよう,`sleep()`で一時停止していることにご注意ください.
#
# なお,2017年3月31日より文化庁メディア芸術データベースのURLが変更されたため,[WebAPI仕様](https://mediaarts-db.bunka.go.jp/webapi_proto_documents.pdf)に記載のリクエストURL(`https://mediaarts-db.jp/mg/api/v1/magazine`)ではなく,新しいURL(`https://mediaarts-db.bunka.go.jp/mg/api/v1/magazine`)を使用する必要があることにご注意ください.
#
# 以下を実行し,`wj-api.json`にデータを保存します.
save_data(magazines)
# タイムアウトした場合は,`offset`を利用して再開します.例えば,`447/2320`でタイムアウトした場合は,`save_data(offset=448)`を実行します.
save_data(magazines, offset=448)
save_data(magazines, offset=500)
save_data(magazines, offset=1269)
save_data(magazines, offset=1889)
save_data(magazines, offset=2274)
| .ipynb_checkpoints/0_obtain_comic_data_j-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=false editable=false nbgrader={"checksum": "a569ad9a41c0d43b58bb9425c5bad9df", "grade": false, "grade_id": "cell-2dfc0bc1e6fbbbd3", "locked": true, "schema_version": 1, "solution": false} id="FtKUK-ucMgL9"
# # Part 3: Sequence Classification
# + [markdown] deletable=false editable=false nbgrader={"checksum": "4e155a44e17248e3d102e1b80e24bf6c", "grade": false, "grade_id": "cell-16d5c7a45d3f9b23", "locked": true, "schema_version": 1, "solution": false} id="R9JEgVndMgME"
# __Before starting, we recommend you enable GPU acceleration if you're running on Colab.__
# + deletable=false editable=false nbgrader={"checksum": "b1b77d0af7b67787cd29f90504f29014", "grade": false, "grade_id": "cell-9fa514521b79541d", "locked": true, "schema_version": 1, "solution": false} id="dia60m-hMgMF" colab={"base_uri": "https://localhost:8080/"} outputId="06c9a0e2-6dc0-4ff4-e504-13077fcd8ee5"
# Execute this code block to install dependencies when running on colab
try:
import torch
except:
from os.path import exists
from wheel.pep425tags import get_abbr_impl, get_impl_ver, get_abi_tag
platform = '{}{}-{}'.format(get_abbr_impl(), get_impl_ver(), get_abi_tag())
# cuda_output = !ldconfig -p|grep cudart.so|sed -e 's/.*\.\([0-9]*\)\.\([0-9]*\)$/cu\1\2/'
accelerator = cuda_output[0] if exists('/dev/nvidia0') else 'cpu'
# !pip install -q http://download.pytorch.org/whl/{accelerator}/torch-1.0.0-{platform}-linux_x86_64.whl torchvision
try:
import torchbearer
except:
# !pip install torchbearer
try:
import torchtext
except:
# !pip install torchtext
try:
import spacy
except:
# !pip install spacy
try:
spacy.load('en')
except:
# !python -m spacy download en
# + [markdown] deletable=false editable=false nbgrader={"checksum": "e0fd6d0300fd2e1ce9a1b34ffd2fefe0", "grade": false, "grade_id": "cell-cabb9cac57ae217e", "locked": true, "schema_version": 1, "solution": false} id="Jmg-f_mlMgMG"
# ## Sequence Classification
# The problem that we will use to demonstrate sequence classification in this lab is the IMDB movie review sentiment classification problem. Each movie review is a variable sequence of words and the sentiment of each movie review must be classified.
#
# The Large Movie Review Dataset (often referred to as the IMDB dataset) contains 25,000 highly-polar movie reviews (good or bad) for training and the same amount again for testing. The problem is to determine whether a given movie review has a positive or negative sentiment. The data was collected by Stanford researchers and was used in a 2011 paper where a split of 50-50 of the data was used for training and test. An accuracy of 88.89% was achieved.
#
# We'll be using a **recurrent neural network** (RNN) as they are commonly used in analysing sequences. An RNN takes in sequence of words, $X=\{x_1, ..., x_T\}$, one at a time, and produces a _hidden state_, $h$, for each word. We use the RNN _recurrently_ by feeding in the current word $x_t$ as well as the hidden state from the previous word, $h_{t-1}$, to produce the next hidden state, $h_t$.
#
# $$h_t = \text{RNN}(x_t, h_{t-1})$$
#
# Once we have our final hidden state, $h_T$, (from feeding in the last word in the sequence, $x_T$) we feed it through a linear layer, $f$, (also known as a fully connected layer), to receive our predicted sentiment, $\hat{y} = f(h_T)$.
#
# Below shows an example sentence, with the RNN predicting zero, which indicates a negative sentiment. The RNN is shown in orange and the linear layer shown in silver. Note that we use the same RNN for every word, i.e. it has the same parameters. The initial hidden state, $h_0$, is a tensor initialized to all zeros.
#
# 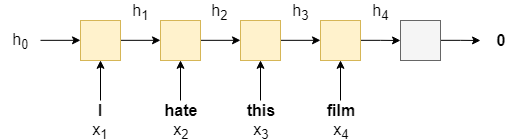
#
# **Note:** some layers and steps have been omitted from the diagram, but these will be explained later.
#
#
# The TorchText library provides easy access to the IMDB dataset. The `IMDB` class allows you to load the dataset in a format that is ready for use in neural network and deep learning models, and TorchText's utility methods allow us to easily create batches of data that are `padded` to the same length (we need to pad shorter sentences in the batch to the length of the longest sentence).
# + [markdown] deletable=false editable=false nbgrader={"checksum": "494c01d089301731550196dabe047067", "grade": false, "grade_id": "cell-23e92e167a2ccd52", "locked": true, "schema_version": 1, "solution": false} id="vDMNVWxDMgMH"
# One of the main concepts of TorchText is the `Field`. These define how your data should be processed. In our sentiment classification task the data consists of both the raw string of the review and the sentiment, either "pos" or "neg".
#
# The parameters of a `Field` specify how the data should be processed.
#
# We use the `TEXT` field to define how the review should be processed, and the `LABEL` field to process the sentiment.
#
# Our `TEXT` field has `tokenize='spacy'` as an argument. This defines that the "tokenization" (the act of splitting the string into discrete "tokens") should be done using the [spaCy](https://spacy.io) tokenizer. If no `tokenize` argument is passed, the default is simply splitting the string on spaces.
#
# `LABEL` is defined by a `LabelField`, a special subset of the `Field` class specifically used for handling labels. We will explain the `dtype` argument later.
#
# For more on `Fields`, go [here](https://github.com/pytorch/text/blob/master/torchtext/data/field.py).
# + deletable=false editable=false nbgrader={"checksum": "ecbcba375e6e1012f085383a6ccee602", "grade": false, "grade_id": "cell-e0561eba5550d048", "locked": true, "schema_version": 1, "solution": false} id="7RfI9lxTMgMH"
import torch
from torchtext.legacy import data
TEXT = data.Field(tokenize='spacy', lower=True, include_lengths=True)
LABEL = data.LabelField(dtype=torch.float)
# + [markdown] deletable=false editable=false nbgrader={"checksum": "c45641b2f9364a3c13d1c00d9c92682e", "grade": false, "grade_id": "cell-0689e30f35617f29", "locked": true, "schema_version": 1, "solution": false} id="W3o9-gUjMgMI"
# The following code automatically downloads the IMDb dataset and splits it into the canonical train/test splits as `torchtext.datasets` objects. It process the data using the `Fields` we have previously defined. Note that the following can take a couple of minutes to run due to the tokenisation:
# + deletable=false editable=false nbgrader={"checksum": "73091f32f831e502e86041779578b7d0", "grade": false, "grade_id": "cell-bfc816072fd54de0", "locked": true, "schema_version": 1, "solution": false} id="kwkvtOzLMgMI" colab={"base_uri": "https://localhost:8080/"} outputId="ed5ee83d-d4ec-4de0-a3f6-2ce3824ff119"
from torchtext.legacy import datasets
train_data, test_data = datasets.IMDB.splits(TEXT, LABEL)
# + [markdown] deletable=false editable=false nbgrader={"checksum": "2ade9f38fd31f83a376798748847ceb4", "grade": false, "grade_id": "cell-a98e9c37bcda1b30", "locked": true, "schema_version": 1, "solution": false} id="viwD9CluMgMJ"
# We can see how many examples are in each split by checking their length.
# + deletable=false editable=false nbgrader={"checksum": "a31d190aee8c06a8f3fa0fe2957bb31e", "grade": false, "grade_id": "cell-a05363648fa59cae", "locked": true, "schema_version": 1, "solution": false} id="MOvqjBNZMgMJ" colab={"base_uri": "https://localhost:8080/"} outputId="1fb4f20f-af4e-4e43-98e0-1c594c494342"
print(f'Number of training examples: {len(train_data)}')
print(f'Number of testing examples: {len(test_data)}')
# + [markdown] deletable=false editable=false nbgrader={"checksum": "f868c66c17e0c39a48728520e07cb468", "grade": false, "grade_id": "cell-83b4651e016c211e", "locked": true, "schema_version": 1, "solution": false} id="c2XwvZwNMgMJ"
# We can also check an example.
# + deletable=false editable=false nbgrader={"checksum": "76a3507d5c228460a24aea95f68c5507", "grade": false, "grade_id": "cell-a3aaf4270ecf8c11", "locked": true, "schema_version": 1, "solution": false} id="7X0ph41-MgMK" colab={"base_uri": "https://localhost:8080/"} outputId="c64f3a41-66d9-4970-95c4-334b8e537d11"
print(vars(train_data.examples[0]))
# + [markdown] deletable=false editable=false nbgrader={"checksum": "5f7210df2512451c6d38c52c76ddf6c1", "grade": false, "grade_id": "cell-1c8ef9d389e1ea7c", "locked": true, "schema_version": 1, "solution": false} id="dwSj5Us-MgMK"
# The IMDb dataset only has train/test splits, so we need to create a validation set. We can do this with the `.split()` method.
#
# By default this splits 70/30, however by passing a `split_ratio` argument, we can change the ratio of the split, i.e. a `split_ratio` of 0.8 would mean 80% of the examples make up the training set and 20% make up the validation set.
#
# We also pass our random seed to the `random_state` argument, ensuring that we get the same train/validation split each time.
# + deletable=false editable=false nbgrader={"checksum": "6282e5f4b4da6ea29f0fa5d349055147", "grade": false, "grade_id": "cell-9a9d0a261cd1d62c", "locked": true, "schema_version": 1, "solution": false} id="fD4OrOVsMgML"
import random
train_data, valid_data = train_data.split()
# + [markdown] deletable=false editable=false nbgrader={"checksum": "8b77b32ded3821f4d03d18d256df19ae", "grade": false, "grade_id": "cell-ebac196da95db0fb", "locked": true, "schema_version": 1, "solution": false} id="lkuV0aZSMgML"
# Again, we'll view how many examples are in each split.
# + deletable=false editable=false nbgrader={"checksum": "8e00a4cf9bddfe86221ed1b820fcea6c", "grade": false, "grade_id": "cell-11de3fcbde1d6f7f", "locked": true, "schema_version": 1, "solution": false} id="QkPL10fPMgML" colab={"base_uri": "https://localhost:8080/"} outputId="aff8b269-67f0-4498-b9e8-edebfa357bce"
print(f'Number of training examples: {len(train_data)}')
print(f'Number of validation examples: {len(valid_data)}')
print(f'Number of testing examples: {len(test_data)}')
# + [markdown] deletable=false editable=false nbgrader={"checksum": "eae6f73416a822fca491ebf20f878aa5", "grade": false, "grade_id": "cell-921d2a5f1737e53b", "locked": true, "schema_version": 1, "solution": false} id="j0hezaJwMgMM"
# Next, we have to build a _vocabulary_. This is effectively a look up table where every unique word in your data set has a corresponding _index_ (an integer).
#
# We do this as our machine learning model cannot operate on strings, only numbers. Each _index_ is used to construct a _one-hot_ vector for each word. A one-hot vector is a vector where all of the elements are 0, except one, which is 1, and dimensionality is the total number of unique words in your vocabulary, commonly denoted by $V$.
#
# 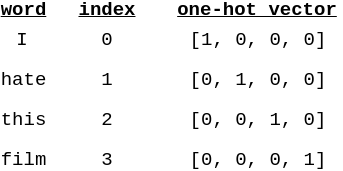
#
# The number of unique words in our training set is over 100,000, which means that our one-hot vectors will have over 100,000 dimensions! This will make training slow and possibly won't fit onto your GPU (if you're using one).
#
# There are two ways to effectively cut-down our vocabulary, we can either only take the top $n$ most common words or ignore words that appear less than $m$ times. We'll do the former, only keeping the top 25,000 words.
#
# What do we do with words that appear in examples but we have cut from the vocabulary? We replace them with a special _unknown_ or `<unk>` token. For example, if the sentence was "This film is great and I love it" but the word "love" was not in the vocabulary, it would become "This film is great and I `<unk>` it".
#
# The following builds the vocabulary, only keeping the most common `max_size` tokens.
# + deletable=false editable=false nbgrader={"checksum": "6746775e3c0746e78afa6b2382d0f249", "grade": false, "grade_id": "cell-1cf0d6f0d09b9333", "locked": true, "schema_version": 1, "solution": false} id="smRlBImEMgMM"
TEXT.build_vocab(train_data, max_size=25000)
LABEL.build_vocab(train_data)
# + [markdown] deletable=false editable=false nbgrader={"checksum": "c406d281eaa4f81948f1e423fd4876ed", "grade": false, "grade_id": "cell-1ca43190cc40ef00", "locked": true, "schema_version": 1, "solution": false} id="YpwwqkDBMgMM"
# Why do we only build the vocabulary on the training set? When testing any machine learning system you do not want to look at the test set in any way. We do not include the validation set as we want it to reflect the test set as much as possible.
# + deletable=false editable=false nbgrader={"checksum": "4e073ef87fd96107616b5016c51b53b7", "grade": false, "grade_id": "cell-79a871ebea509deb", "locked": true, "schema_version": 1, "solution": false} id="hc5h4VZPMgMN" colab={"base_uri": "https://localhost:8080/"} outputId="4cf2265c-10a1-4a94-b1db-74c96302c7a4"
print(f"Unique tokens in TEXT vocabulary: {len(TEXT.vocab)}")
print(f"Unique tokens in LABEL vocabulary: {len(LABEL.vocab)}")
# + [markdown] deletable=false editable=false nbgrader={"checksum": "0d084d49bcadb8addbb1b8cd39bda543", "grade": false, "grade_id": "cell-74d663b76304878f", "locked": true, "schema_version": 1, "solution": false} id="a03IJg-kMgMN"
# Why is the vocab size 25002 and not 25000? One of the addition tokens is the `<unk>` token and the other is a `<pad>` token.
#
# When we feed sentences into our model, we feed a _batch_ of them at a time, i.e. more than one at a time, and all sentences in the batch need to be the same size. Thus, to ensure each sentence in the batch is the same size, any sentences which are shorter than the longest within the batch are padded.
#
# 
#
# We can also view the most common words in the vocabulary and their frequencies.
# + deletable=false editable=false nbgrader={"checksum": "5c353d4eec3c94b10fb4228a9af8f274", "grade": false, "grade_id": "cell-0efc8b86fea8d6e2", "locked": true, "schema_version": 1, "solution": false} id="9NdqHYnEMgMN" colab={"base_uri": "https://localhost:8080/"} outputId="9ec0757a-c948-4987-daf1-eca76bdebde9"
print(TEXT.vocab.freqs.most_common(20))
# + [markdown] deletable=false editable=false nbgrader={"checksum": "6f0044693302832d5b864ea9f20bcaa2", "grade": false, "grade_id": "cell-68985316db3edb24", "locked": true, "schema_version": 1, "solution": false} id="tecCrpOJMgMN"
# We can also see the vocabulary directly using either the `stoi` (**s**tring **to** **i**nt) or `itos` (**i**nt **to** **s**tring) method.
# + deletable=false editable=false nbgrader={"checksum": "862acc66ef827e8a3e33ca822b682728", "grade": false, "grade_id": "cell-3f6931771dfb8b05", "locked": true, "schema_version": 1, "solution": false} id="F_vzIm9nMgMO" colab={"base_uri": "https://localhost:8080/"} outputId="74804b88-a904-44ad-fd1d-9e549d1c4397"
print(TEXT.vocab.itos[:10])
# + [markdown] deletable=false editable=false nbgrader={"checksum": "e5f1fcac73f30d4b4f3fff4d3ba247b3", "grade": false, "grade_id": "cell-126deacfb7443e94", "locked": true, "schema_version": 1, "solution": false} id="fLQxX2TuMgMO"
# We can also check the labels, ensuring 0 is for negative and 1 is for positive.
# + deletable=false editable=false nbgrader={"checksum": "7d95213f5534ac30134149c90fa6c6c7", "grade": false, "grade_id": "cell-c45080555e1e47a8", "locked": true, "schema_version": 1, "solution": false} id="9nKj81XVMgMO" colab={"base_uri": "https://localhost:8080/"} outputId="a08b046c-cc8e-4698-a009-1664b0472093"
print(LABEL.vocab.stoi)
# + [markdown] deletable=false editable=false nbgrader={"checksum": "217fe362a504a59655710774d6bd4f3e", "grade": false, "grade_id": "cell-35a9ad2d4bf17b42", "locked": true, "schema_version": 1, "solution": false} id="ySUc_WTlMgMO"
# The final step of preparing the data is creating the iterators. We iterate over these in the training/evaluation loop, and they return a batch of examples (indexed and converted into tensors) at each iteration.
#
# We'll use a `BucketIterator` which is a special type of iterator that will return a batch of examples where each example is of a similar length, minimizing the amount of padding per example. Torchtext will pad for us automatically (handled by the `Field` object). We'll request the items within each batch produced by the `BucketIterator` are sorted by length.
#
# We also want to place the tensors returned by the iterator on the GPU (if you're using one). PyTorch handles this using `torch.device`, we then pass this device to the iterator.
# + deletable=false editable=false nbgrader={"checksum": "526210054aa53f54d8dad8acf67d1dcf", "grade": false, "grade_id": "cell-722b81c8ccb13d25", "locked": true, "schema_version": 1, "solution": false} id="XtDn1rSjMgMP"
BATCH_SIZE = 64
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
train_iterator, valid_iterator, test_iterator = data.BucketIterator.splits(
(train_data, valid_data, test_data),
batch_size=BATCH_SIZE,
device=device,
sort_key=lambda x: len(x.text),
sort_within_batch=True)
# + [markdown] deletable=false editable=false nbgrader={"checksum": "def10308b65f2ed87e6fe3d3f2d89727", "grade": false, "grade_id": "cell-5d7acf8d6db191d3", "locked": true, "schema_version": 1, "solution": false} id="LHOYgZ5XMgMP"
# ## Build the Model
#
# The next stage is building the model that we'll eventually train and evaluate.
#
# There is a small amount of boilerplate code when creating models in PyTorch, note how our `RNN` class is a sub-class of `nn.Module` and the use of `super`.
#
# Within the `__init__` we define the _layers_ of the module. Our three layers are an _embedding_ layer, our RNN, and a _linear_ layer. All layers have their parameters initialized to random values, unless explicitly specified.
#
# The embedding layer is used to transform our sparse one-hot vector (sparse as most of the elements are 0) into a dense embedding vector (dense as the dimensionality is a lot smaller and all the elements are real numbers). This embedding layer is simply a single fully connected layer. As well as reducing the dimensionality of the input to the RNN, there is the theory that words which have similar impact on the sentiment of the review are mapped close together in this dense vector space. For more information about word embeddings, see [here](https://monkeylearn.com/blog/word-embeddings-transform-text-numbers/).
#
# The RNN layer is our RNN which takes in our dense vector and the previous hidden state $h_{t-1}$, which it uses to calculate the next hidden state, $h_t$.
#
# 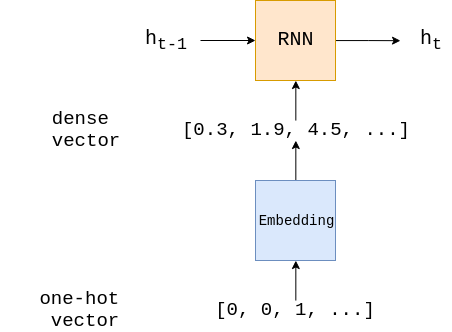
#
# Finally, the linear layer takes the final hidden state and feeds it through a fully connected layer, $f(h_T)$, transforming it to the correct output dimension.
#
# The `forward` method is called when we feed examples into our model.
#
# Each batch, `text_len`, is a tuple containing a tensor of size _**[max_sentence length, batch size]**_ and a tensor of **batch_size** containing the true lengths of each sentence (remember, they won't necessarily be the same; some reviews are much longer than others).
#
# The first tensor in the tuple contains the ordered word indexes for each review in the batch. The act of converting a list of tokens into a list of indexes is commonly called *numericalizing*.
#
# The input batch is then passed through the embedding layer to get `embedded`, which gives us a dense vector representation of our sentences. `embedded` is a tensor of size _**[sentence length, batch size, embedding dim]**_.
#
# `embedded` is then fed into a function called `pack_padded_sequence` before being fed into the RNN. `pack_padded_sequence` is used to create a datastructure that allows the RNN to 'mask' off the padding during the BPTT process (we don't want to learn the padding, as this could drastically influence the results!). In some frameworks you must feed the initial hidden state, $h_0$, into the RNN, however in PyTorch, if no initial hidden state is passed as an argument it defaults to a tensor of all zeros.
#
# The RNN returns 2 tensors, `output` of size _**[sentence length, batch size, hidden dim]**_ and `hidden` of size _**[1, batch size, hidden dim]**_. `output` is the concatenation of the hidden state from every time step, whereas `hidden` is simply the final hidden state.
#
# Finally, we feed the last hidden state, `hidden`, through the linear layer, `fc`, to produce a prediction. Note the `squeeze` method, which is used to remove a dimension of size 1.
# + deletable=false editable=false nbgrader={"checksum": "c785b0842586d20d70510c6f8a2c6b39", "grade": false, "grade_id": "cell-fbb5f94b744dd6db", "locked": true, "schema_version": 1, "solution": false} id="_7IsmK3dMgMQ"
import torch.nn as nn
class RNN(nn.Module):
def __init__(self, input_dim, embedding_dim, hidden_dim, output_dim):
super().__init__()
self.embedding = nn.Embedding(input_dim, embedding_dim)
self.rnn = nn.RNN(embedding_dim, hidden_dim)
self.fc = nn.Linear(hidden_dim, output_dim)
def forward(self, text, lengths):
embedded = self.embedding(text)
embedded = nn.utils.rnn.pack_padded_sequence(embedded, lengths)
packed_output, hidden = self.rnn(embedded)
return self.fc(hidden.squeeze(0))
# + [markdown] deletable=false editable=false nbgrader={"checksum": "3c834acdf19cf4124c69e3b92c9fbd91", "grade": false, "grade_id": "cell-8b4748e05072b330", "locked": true, "schema_version": 1, "solution": false} id="WsyIyUUnMgMQ"
# We now create an instance of our RNN class.
#
# The input dimension is the dimension of the one-hot vectors, which is equal to the vocabulary size.
#
# The embedding dimension is the size of the dense word vectors. This is usually around 50-250 dimensions, but depends on the size of the vocabulary.
#
# The hidden dimension is the size of the hidden states. This is usually around 100-500 dimensions, but also depends on factors such as on the vocabulary size, the size of the dense vectors and the complexity of the task.
#
# The output dimension is usually the number of classes, however in the case of only 2 classes the output value is between 0 and 1 and thus can be 1-dimensional, i.e. a single scalar real number.
# + deletable=false editable=false nbgrader={"checksum": "a7f66791ac9794a7da9ee4e6fc743b24", "grade": false, "grade_id": "cell-751c2df54b71d158", "locked": true, "schema_version": 1, "solution": false} id="8xNhSsw4MgMR"
INPUT_DIM = len(TEXT.vocab)
EMBEDDING_DIM = 50
HIDDEN_DIM = 100
OUTPUT_DIM = 1
model = RNN(INPUT_DIM, EMBEDDING_DIM, HIDDEN_DIM, OUTPUT_DIM)
# + [markdown] deletable=false editable=false nbgrader={"checksum": "591ccfef47e2bcc226185c87e568821a", "grade": false, "grade_id": "cell-daf23924e258a608", "locked": true, "schema_version": 1, "solution": false} id="wjpIQn45MgMR"
# # Train the model
#
# Now we'll set up the training and then train the model.
#
# First, we'll create an optimizer. This is the algorithm we use to update the parameters of the module. Here, we'll use _stochastic gradient descent_ (SGD). The first argument is the parameters that will be updated by the optimizer, the second is the learning rate, i.e. how much we'll change the parameters by when we do a parameter update.
# + deletable=false editable=false nbgrader={"checksum": "4a0e4ee985e7d35fcf80b8b3eaabdbde", "grade": false, "grade_id": "cell-d7566606b6f480ec", "locked": true, "schema_version": 1, "solution": false} id="7HtU_2IhMgMR"
import torch.optim as optim
optimizer = optim.SGD(model.parameters(), lr=0.001)
# + [markdown] deletable=false editable=false nbgrader={"checksum": "072bc1b948afe036e31f60a05469fc27", "grade": false, "grade_id": "cell-8981a2c109df3c53", "locked": true, "schema_version": 1, "solution": false} id="DF_wtJzlMgMS"
# Next, we'll define our loss function. In PyTorch this is commonly called a criterion.
#
# The loss function here is _binary cross entropy with logits_.
#
# Our model currently outputs an unbound real number. As our labels are either 0 or 1, we want to restrict the predictions to a number between 0 and 1. We do this using the _sigmoid_ function.
#
# We then use this this bound scalar to calculate the loss using binary cross entropy.
#
# The `BCEWithLogitsLoss` criterion carries out both the sigmoid and the binary cross entropy steps.
# + deletable=false editable=false nbgrader={"checksum": "dee16f7d05d182806906fc2f3d6c4484", "grade": false, "grade_id": "cell-99068d084d5dfb73", "locked": true, "schema_version": 1, "solution": false} id="OnMi550tMgMS"
criterion = nn.BCEWithLogitsLoss()
# + [markdown] deletable=false editable=false nbgrader={"checksum": "7a7cef0ba9cb2d6b49111244fb8f5841", "grade": false, "grade_id": "cell-71e6ceff6ffba4f7", "locked": true, "schema_version": 1, "solution": false} id="q19_J8ZjMgMS"
# Finally, before creating a Torchbearer trial and copying the model to the GPU (if we have one), we need to adapt the format of the batches returned by TorchText. TorchText's iterators return an object representing a batch of data from which you can access the fields used to store the underlying data (e.g. the labels and words in our case). Torchbearer on the other hand needs to know what the X's and y's are explicitly.
#
# There are a number of ways in which we can adapt the batch data format, but one of the easiest conceptually is to write a simple wrapper iterator that reads the next torchtext batch and returns a tuple of objects (the X and y). Note that because we specified that we want the lengths of the sentences earlier, the X's will be a tuple of the sentences and their lengths. If we had not requested the lengths, then the X's would just be a tensor encoding the padded sentences.
# + deletable=false editable=false nbgrader={"checksum": "fd0f46de194f98826875d7431eee5918", "grade": false, "grade_id": "cell-f011ec7d73d7ef46", "locked": true, "schema_version": 1, "solution": false} id="ENcWrkWQMgMS" colab={"base_uri": "https://localhost:8080/", "height": 671, "referenced_widgets": ["e1b77d3d4c144a99b70a3240b093d5bf", "1084fc42ddbe4ec2a0bb3daf918a32f8", "84cd26f77fd74d62aaedc8b34ab41d46", "afdd5e8c2b494f5596e30cb6c7f9afbd", "061ad94c0ef54dc2ac8ccc7937455998", "096951fea34340269964b0cb4a79cd5e", "ea38ecd3379b4f0aa52956ab5b0fd41a", "b3344aff369b41d198faf25ac6cf2ea2", "11c838b1ea374564a36fdda995c35830", "9456a421b1a44cfd9348550a14f9b01d", "d60509c2a3134a81b3e67e5300705585", "82dc41eee79549a193d961a0f190f341", "4a166af383e84f7eab1dce667105fa11", "88fcc86c46444ce5b521fac49fe5a637", "1c3e58d295a54241b6b825155b1612e8", "42d91beb2cd1466384e58817f672db4e", "4e118ae97f6947ef8672359f9566736b", "<KEY>", "<KEY>", "bb41885e07fa44ad89ef0eadef870a99", "<KEY>", "8ea1e99cada147b4a8a54e7c875ca21a", "<KEY>", "<KEY>", "<KEY>", "665ce851cc36405d86edff086a9c9b5b", "<KEY>", "<KEY>", "3bea6f9dd82e4cc4a3d2911a1047d9cf", "<KEY>", "7a0ca99887d14d41ac35bd4055a0c96f", "949ddf954a8f49d2af19e3300bbec54c", "<KEY>", "1d2021dad0d444b8b01500a2a6e1f075", "676aca2600e142a5858eed6231611469", "<KEY>", "df9a352e74a742adb838ce1514908617", "508f73aadbcc4a1bacde4e3a57a6220c", "<KEY>", "<KEY>", "<KEY>", "0f6f9fea46404e83a963598ae934fcea", "5171f203fa084fc1bae37671f65afca9", "8e03a04b604f472886e5666e58a3f5f5", "a0fe9cabf5e34ab5b7fad83c27ef0f51", "<KEY>", "<KEY>", "b1e7bf4fd5a74c55a56094b205346dd1", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "da1f964f7fcb42af97afcfa180defeef", "f3d7eff72d0843faa4c0687f7ee94fe7", "4aff16461b8640bd9e6a0ec0489ad999", "<KEY>", "07e10922a993434683d7d484845422d3", "81069ba6947c419685801395eb0ead59", "<KEY>", "bd1efce7f45b4dfaa81e3a523be38069", "af1f624446ea443e8e25efa3d003107d", "56aab64aa6d84adbae7eec8709da03b2", "c9ff6865ee17401ea9de9bc2005f2ce6", "<KEY>", "2ed14a796feb490e98985a3e09a3e03e", "<KEY>", "006e1dedafb84caa9320fcb422da96c8", "<KEY>", "<KEY>", "6f6f232362c54729a3facf5ed44ceb11", "9e2d76af191444f1a5ba479ac6d62917", "<KEY>", "<KEY>", "59008e5d88e64607a7262d5830c70f6b", "<KEY>", "6eb9e070ea3f4335a8580ad6698785cb", "<KEY>", "<KEY>", "<KEY>", "d86d904ca88746d1bdb56978218ae39a", "<KEY>", "8c9cd2e02ff2465ca28121d199038015", "<KEY>", "688ded6f6fcd4334a1e8e4310cac702b", "ecc013ae84de419eb4b04490f69e1c01", "<KEY>", "7104a74782ff4956940e3a92b5b58bc5", "<KEY>"]} outputId="6f1b8bc6-195d-4049-c6b5-c2148c5c5110"
from torchbearer import Trial
class MyIter:
def __init__(self, it):
self.it = it
def __iter__(self):
for batch in self.it:
yield (batch.text, batch.label.unsqueeze(1))
def __len__(self):
return len(self.it)
torchbearer_trial = Trial(model, optimizer, criterion, metrics=['acc', 'loss']).to(device)
torchbearer_trial.with_generators(train_generator=MyIter(train_iterator), val_generator=MyIter(valid_iterator), test_generator=MyIter(test_iterator))
torchbearer_trial.run(epochs=5)
torchbearer_trial.predict()
# + [markdown] deletable=false editable=false nbgrader={"checksum": "9853268a5e221536d6e9650e8a77d473", "grade": false, "grade_id": "cell-03b5931999ab62d1", "locked": true, "schema_version": 1, "solution": false} id="pLui9Q6RMgMS"
# __Use the box below to comment on and give insight into the performance of the above model:__
# + [markdown] deletable=false nbgrader={"checksum": "615ab310b18718e361d870399422e5ae", "grade": true, "grade_id": "cell-5bf61cbc741af01b", "locked": false, "points": 5, "schema_version": 1, "solution": true} id="GHBcAZENMgMT"
# YOUR ANSWER HERE
# + [markdown] deletable=false editable=false nbgrader={"checksum": "fbeb865b3ebb7348e9d0e0e2b3f1e6d2", "grade": false, "grade_id": "cell-acff7e648aa99e42", "locked": true, "schema_version": 1, "solution": false} id="kXJoww5QMgMT"
# Now try and build a better model. Rather than using a plain RNN, we'll instead use a (single layer) LSTM, and we'll use Adam with an initial learning rate of 0.01 as the optimiser. __Complete the following code to implement the improved model, and then train it:__
# + deletable=false nbgrader={"checksum": "ef8857370d3a96cdb46acea6f94f99ac", "grade": true, "grade_id": "cell-7c7913d0313ff2e8", "locked": false, "points": 5, "schema_version": 1, "solution": true} id="nj6fAY8EMgMT" colab={"base_uri": "https://localhost:8080/", "height": 671, "referenced_widgets": ["9f417adac6014d8282be9dcb66bf073a", "352a6c58fbd1429797360e8aaaf63e4a", "fcf9f72785824b27a28c28821ef72dc9", "<KEY>", "0722c638811c42e5b93ad1d81a0d5911", "f81abee0764340f7af1ef0cba2686dfa", "<KEY>", "8ec1dc0552744165b03385ed8fee144a", "<KEY>", "f8fcda00a388497789cd1e23d775a16c", "<KEY>", "cba54384a54a4e8daa80ce9d33ec5624", "<KEY>", "<KEY>", "<KEY>", "b56b4a3e82844370802bbfeb5a6ee50f", "c2cd253c280649828d632bc0b715d7d1", "9ad2bfd3855947febd0deca522985136", "<KEY>", "<KEY>", "781448decea84f68a2b7bd1820a027c6", "<KEY>", "d9c4572410674acda9e8c8a17772ff41", "<KEY>", "8cd783ee2c7f456d899af23501c83e5f", "<KEY>", "b40eeec226bd47a9b19ac1c563d9b7b6", "46284bd282764be68c940d4703f5577e", "<KEY>", "<KEY>", "<KEY>", "e6f48f9590044f68aa708787af4da378", "b22d2a3c81484419ad5046314ce7a731", "d37ae625cb144a4f8961be57187817b1", "<KEY>", "<KEY>", "<KEY>", "20378df31da54dbb9eeb1ba06391e063", "b2b186afa365493f911401489a00c7e5", "00a5d0a42349441da91adae11783e6e9", "<KEY>", "bce6719371c94c6c91eeb39e538ef7eb", "<KEY>", "<KEY>", "ddbe89fc895c4c05a0201983980cf795", "e8d494497cba41d392ca718f4119d477", "<KEY>", "e7e2e233f3d94e3e86c60d5540e7bbb2", "d32af090895a4d3e87fcc867a5c381d3", "b60406eb35ca40648243a73eb5be7dab", "b72d148197504436bf0cd76bc17619ca", "<KEY>", "b2422bec65364117a0996a9153cf3dab", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "fb372696b1484e238de6ed9859167567", "b4250f70df884e5c9ebd2c6594a1da11", "<KEY>", "<KEY>", "<KEY>", "7f7ca2b6098a4e9283d9bad7291c73ae", "331830388ce546d49ccb483a85b64c7c", "c2d8fb59ac7a4c93b9dfd30da4da9048", "baf43af1f22e4e8186ed46c814c53756", "<KEY>", "<KEY>", "abfb70870e3d4940bd98e7f679ce7ece", "f533cdb29cea4d8b9df3a6a9d113ff6f", "d7f5695fbe804e6e83cc5fdf40747b17", "e35120e741a146e99dfdd474a245c3a9", "<KEY>", "7c7220a4e5f04d0787f50ad3fca6ff34", "d4f31f1242f34f0d8ce2605860e13e05", "b9f45e7182544b459ecacc5f102ef2ba", "f0fcb536c6ce41c0a52312610b6e1d93", "f9c3c052ed2842cbba33b27815364ae9", "<KEY>", "c94813efd41a431f8abb82b82a36b0f9", "2f9a6f590ced48cc8e150f8a9816ec5a", "<KEY>", "<KEY>", "<KEY>", "a0b87cb0397741b7976d98406854c393", "0097c89238e9480a8aca082fcde0d97a", "c89d0645ec4c47d2b6373bafb6d9d801", "254e959b1605441283b58e9c3899f650"]} outputId="0572b844-1daa-41d4-f2d1-9f76429ccbfb"
class ImprovedRNN(nn.Module):
def __init__(self, input_dim, embedding_dim, hidden_dim, output_dim):
super().__init__()
self.embedding = nn.Embedding(input_dim, embedding_dim)
self.lstm = nn.LSTM(embedding_dim, hidden_dim)
self.fc = nn.Linear(hidden_dim, output_dim)
def forward(self, text, lengths):
embedded = self.embedding(text)
embedded = nn.utils.rnn.pack_padded_sequence(embedded, lengths)
lstm_out, (ht, ct) = self.lstm(embedded)
out = self.fc(ht[-1])
return out
INPUT_DIM = len(TEXT.vocab)
EMBEDDING_DIM = 50
HIDDEN_DIM = 100
OUTPUT_DIM = 1
imodel = ImprovedRNN(INPUT_DIM, EMBEDDING_DIM, HIDDEN_DIM, OUTPUT_DIM)
optimizer = optim.Adam(imodel.parameters(), lr=0.001)
criterion = nn.BCEWithLogitsLoss()
torchbearer_trial = Trial(imodel, optimizer, criterion, metrics=['acc', 'loss']).to(device)
torchbearer_trial.with_generators(train_generator=MyIter(train_iterator), val_generator=MyIter(valid_iterator), test_generator=MyIter(test_iterator))
torchbearer_trial.run(epochs=5)
torchbearer_trial.predict()
# + [markdown] deletable=false editable=false nbgrader={"checksum": "a84da93b0d8631bd2279f0a0ca7cecf7", "grade": false, "grade_id": "cell-9da7d879835eafa4", "locked": true, "schema_version": 1, "solution": false} id="xrpEExtNMgMT"
# __What do you observe about the performance of this model? What would you do next if you wanted to improve it further? Write your answers in the box below:__
# + deletable=false nbgrader={"checksum": "bd9fe3f4f88270f113a2731db2b1b2b7", "grade": true, "grade_id": "cell-856a0834622f664f", "locked": false, "points": 10, "schema_version": 1, "solution": true} id="hUvE_dG6MgMT"
# + [markdown] deletable=false editable=false nbgrader={"checksum": "21752b32fcb29f71d7da22bf21a14a51", "grade": false, "grade_id": "cell-f5e06c722621a0d2", "locked": true, "schema_version": 1, "solution": false} id="cEDImkpBMgMU"
# ## User Input
#
# We can now use our models to predict the sentiment of any sentence we give it. As it has been trained on movie reviews, the sentences provided should also be movie reviews.
#
# Our `predict_sentiment` function does a few things:
# - tokenizes the sentence, i.e. splits it from a raw string into a list of tokens
# - indexes the tokens by converting them into their integer representation from our vocabulary
# - converts the indexes, which are a Python list into a PyTorch tensor
# - add a batch dimension by `unsqueeze`ing
# - squashes the output prediction from a real number between 0 and 1 with the `sigmoid` function
# - converts the tensor holding a single value into an integer with the `item()` method
#
# We are expecting reviews with a negative sentiment to return a value close to 0 and positive reviews to return a value close to 1.
# + deletable=false editable=false nbgrader={"checksum": "03dd780275233d49f92dc46d21217de6", "grade": false, "grade_id": "cell-256f5d0cab3585a2", "locked": true, "schema_version": 1, "solution": false} id="cYeepOtkMgMU"
import spacy
nlp = spacy.load('en')
def predict_sentiment(model, sentence):
tokenized = [tok.text for tok in nlp.tokenizer(sentence)]
indexed = [TEXT.vocab.stoi[t] for t in tokenized]
tensor = torch.LongTensor(indexed).to(device)
tensor = tensor.unsqueeze(1)
prediction = torch.sigmoid(model((tensor, torch.tensor([tensor.shape[0]]))))
return prediction.item()
# + [markdown] deletable=false editable=false nbgrader={"checksum": "2f0ff29f9b4ff26535014b40ac64f159", "grade": false, "grade_id": "cell-44ca76b13f0ef977", "locked": true, "schema_version": 1, "solution": false} id="TksV9Z1SMgMU"
# An example negative review...
# + id="9-cP0QVhzhfp"
predict_sentiment(imodel, "This film is terrible")
# + [markdown] deletable=false editable=false nbgrader={"checksum": "f890392d505e39def4b5dc80d028019f", "grade": false, "grade_id": "cell-78424acf52854f0e", "locked": true, "schema_version": 1, "solution": false} id="r4wDjnqrMgMU"
# and an example positive review...
# + id="W2V92f5fzjxU"
predict_sentiment(imodel, "This film is great")
| Pytorch Practical Tasks/7_3_SequenceClassification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from math import cos, sin, asin, acos, gcd, atan2
import numpy as np
from numpy import array, dot, degrees, cross
from numpy.linalg import inv, det, solve, norm
import pandas as pd
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# import the python scripts:
import gb_code.csl_generator as csl
import gb_code.gb_generator as gbc
# %matplotlib notebook
# -
# ## Produce Lists of CSL boundaries for any given rotation axis (hkl) :
#
#
# +
# for example: [1, 0, 0], [1, 1, 0] or [1, 1, 1]
axis = np.array([1, 1, 1])
# list Sigma boundaries < 50
csl.print_list(axis, 50)
# -
# ## Select a sigma and get the characteristics of the GB:
# +
# pick a sigma for this axis, ex: 19.
sigma = 19
theta, m, n = csl.get_theta_m_n_list(axis, sigma)[0]
R = csl.rot(axis, theta)
# Minimal CSL cells. The plane orientations and the orthogonal cells
# will be produced from these original cells.
M1, M2 = csl.Create_minimal_cell_Method_1(sigma, axis, R)
print('Angle:', degrees(theta), '\n', 'Sigma:', sigma, '\n',
'Minimal cells:', '\n', M1, '\n', M2, '\n')
# -
# ## Produce Lists of GB planes for the chosen boundary :
#
# +
# the higher the limit the higher the indices of GB planes produced.
lim = 3
V1, V2, M, Gb = csl.Create_Possible_GB_Plane_List(axis, m, n, lim)
# the following data frame shows the created list of GB planes and their corresponding types
df = pd.DataFrame(
{'GB1': list(V1),
'GB2': list(V2),
'Type': Gb
})
df.head()
# -
# ## Criteria for finding the GB plane of interest*:
#
# ### 1- Based on the type of GB plane:
# #### _*The following criteria searches the generated data frame. To extend the search you can increase the limit (lim) in the above cell._
# +
# Gb types: Symmetric Tilt, Tilt, Twist, Mixed
df[df['Type'] == 'Tilt'].head()
# -
#
# ### 2 - Based on the minimum number of atoms in the orthogonal cell:
#
#
# #### _This can be of interest for DFT calculations that require smaller cells. The search may take a few minutes if your original limit is large as it must calculate all the orthogonal cells to know the number of atoms._
basis = 'fcc'
Number = np.zeros(len(V1))
Number_atoms = []
for i in range((len(V1))):
Number_atoms.append(csl.Find_Orthogonal_cell(basis,axis,m,n,V1[i])[2])
# +
# show me Gb planes that have orthogonal cells with less than max_num_atoms, here: 500.
df['Number'] = Number_atoms
max_num_atoms = 300
df[df['Number'] < max_num_atoms]
# -
# ### 3- Based on proximity to a particular type boundary, for example the Symmetric tilts:
# #### _This is how I created various steps on grain boundaries with vicinal orientations in the following work: __(https://journals.aps.org/prmaterials/abstract/10.1103/PhysRevMaterials.2.043601)__ See also here:_
# __(https://www.mpie.de/2955247/GrainBoundaryDynamics)__
# +
SymmTiltGbs = []
for i in range(len(V1)):
if str(Gb[i]) == 'Symmetric Tilt':
SymmTiltGbs.append(V1[i])
# Find GBs less than Delta (here 6) degrees from any of the symmetric tilt boundaries in this system
Delta = 6
Min_angles = []
for i in range(len(V1)):
angles = []
for j in range(len(SymmTiltGbs)):
angles.append(csl.angv(V1[i],SymmTiltGbs[j]))
Min_angles.append(min(angles))
# +
df['Angles'] = Min_angles
df[df['Angles'] < Delta]
# -
# ## Select a GB plane and go on:
# ### You only need to pick the GB1 plane, from any of the three criteria in the cells above
GB_plane = [-2, -3, 5]
# lattice parameter
LatP = 4
basis = 'fcc'
# just a piece of info, how much of this mixed boundary is tilt or twist?
csl.Tilt_Twist_comp(GB_plane, axis, m, n)
# +
# instantiate a GB:
my_gb = gbc.GB_character()
# give all the characteristics
my_gb.ParseGB(axis, basis, LatP, m, n, GB_plane)
# Create the bicrystal
my_gb.CSL_Bicrystal_Atom_generator()
# Write a GB :
# by default overlap = 0.0, rigid = False,
# dim1, dim2, dim3 = [1, 1, 1]
# file = 'LAMMPS'
# read the io_file and/or README for more info, briefly: when you give overlap > 0
# you need to decide 'whichG': atoms to be removed from either G1 or G2.
# if rigid = True, then you need two integers a and b to make a mesh on the GB plane
# a and b can be put to 10 and 5 (just a suggestion) respectively for any GB except
# TWIST for the TWISTS the code will handle it internally regardless of your input a and b
# ex:
#my_gb.WriteGB(overlap=0.3, whichG='g1', rigid= True, a=3, b=2, dim2=5 , file='VASP')
my_gb.WriteGB(overlap=0.3, whichG='g1', dim1=3, dim2=3, dim3=2, file='VASP')
# +
# extract atom positions of the two grains:
X = my_gb.atoms1
Y = my_gb.atoms2
# +
# 3d plot of the gb, that can be shown in any cartesian view direction:
def plot_gb(X,Y, view_dir = [0,1,0]):
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.scatter(X[:,0],X[:,1],X[:,2],'o', s = 20, facecolor = 'y',edgecolor='none', alpha=0.2 )
ax.scatter(Y[:,0],Y[:,1],Y[:,2],'o',s = 20,facecolor = 'b',edgecolor='none', alpha=0.2)
# Show [0, 0, 0] as a red point
ax.scatter(0,0,0,'s',s = 200,facecolor = 'r',edgecolor='none')
ax.set_proj_type('ortho')
ax.grid(False)
az = degrees(atan2(view_dir[1],view_dir[0]))
el = degrees(asin(view_dir[2]/norm(view_dir)))
ax.view_init(azim = az, elev = el)
ax.set_xlabel('X axis')
ax.set_ylabel('Y axis')
ax.set_zlabel('Z axis')
return
# -
# %matplotlib notebook
plot_gb(X,Y,[0,1,0])
| Test/Usage_of_GB_code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# This module process the data.
df = pd.read_excel("./data.xlsx", usecols=[0, 1])
df.columns = ["Time", "Flow"]
print(df.head())
print("Length: {}".format(len(df)))
# +
from pandas.plotting import register_matplotlib_converters
register_matplotlib_converters()
plt.plot(df["Time"], df["Flow"])
plt.title("Flow Volumns Distribution Diagram")
plt.xlabel("Time")
plt.ylabel("Flow")
plt.savefig("Flow1.png")
plt.show()
# +
from scipy.fftpack import fft,ifft
from sklearn.metrics import mean_squared_error
result = fft(df["Flow"])
plt.plot(result)
plt.title("FFT Transformation Diagram - 1")
plt.xlabel("Frequency")
plt.ylabel("Weight")
plt.savefig("FFT0.png")
plt.show()
print(result[1])
print(result[2])
print(result[3])
print(result[4])
for i in range(len(result)):
if (abs(result[i]) <= 7000):
result[i] = 0
plt.plot(result)
plt.title("FFT Transformation Diagram - 2")
plt.xlabel("Frequency")
plt.ylabel("Weight")
plt.savefig("FFT.png")
plt.show()
test = ifft(result)
print(test)
plt.plot(df["Time"], test)
plt.title("Flow Volumns Distribution Diagram - 2")
plt.xlabel("Time")
plt.ylabel("Flow")
plt.savefig("Flow2.png")
plt.show()
print(len(df["Flow"]))
print(mean_squared_error(df["Flow"].values, abs(test)))
# -
print(result)
print(fft(df["Flow"]))
| Flow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Kanghee-Lee/Mask-RCNN_TF/blob/master/Mask_RCNN(RPN).ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="3QCILgjXGPLJ" colab_type="code" colab={}
#Imports
import numpy as np
import os
import glob
import cv2
# !pip install xmltodict
import xmltodict
import tensorflow as tf
import math
from tqdm import tqdm
from PIL import Image, ImageDraw
from google.colab.patches import cv2_imshow
from keras import layers as layers
from keras import models as models
# + id="QppSjIWIKhCh" colab_type="code" colab={}
#download VOC data and extract .tar file
# !mkdir train
# !mkdir test
# !wget http://pjreddie.com/media/files/VOCtrainval_06-Nov-2007.tar -P train/
# !wget https://storage.googleapis.com/coco-dataset/external/PASCAL_VOC.zip -P train/
# !wget http://pjreddie.com/media/files/VOCtest_06-Nov-2007.tar -P test/
# !tar -xf test/VOCtest_06-Nov-2007.tar -C test/
# !tar -xf train/VOCtrainval_06-Nov-2007.tar -C train/
# !unzip train/PASCAL_VOC.zip -d train/
# !rm -rf train/PASCAL_VOC.zip train/VOCtrainval_06-Nov-2007.tar
# !rm -rf test/VOCtest_06-Nov-2007.tar
# + id="_JIp9A0LHAwt" colab_type="code" colab={}
#from google.colab import drive
#drive.mount('/content/drive')
# + id="fzYVT9h2Gjo1" colab_type="code" colab={}
data_images_path = os.getcwd()+'/train/VOCdevkit/VOC2007/JPEGImages'
data_annotation_path = os.getcwd()+'/train/VOCdevkit/VOC2007/Annotations'
img_min=800
img_max=1024
image_height = 1024
image_width = 1024
image_depth = 3 # RGB
rpn_kernel_size = 3 # 3x3
subsampled_ratio = [4, 8, 16, 32, 64] # Pooling 3 times
anchor_sizes = [32,64,128, 256, 512] # Using [128, 256, 512] sizes in 1000x600 img size / Used [32, 64, 128] sizes in 224x224
anchor_aspect_ratio = [[1,1],[1/math.sqrt(2),math.sqrt(2)],[math.sqrt(2),1/math.sqrt(2)]]
num_anchors_in_box = len(anchor_aspect_ratio)
neg_threshold = 0.3
pos_threshold = 0.7
nms_threshold = 0.7
anchor_sampling_amount = 128 # 128 for each positive, negative sampling
# + id="zy2n8V-HG2CN" colab_type="code" colab={}
list_images = sorted([x for x in glob.glob(data_images_path + '/**')])
#list_images=list_images[:2000]
total_images = len(list_images)
print(total_images)
list_annotations = sorted([x for x in glob.glob(data_annotation_path + '/**')])
#list_annotations = list_annotations[:2000]
# evaluating data consistency between images and annotations
t1=[]
t2=[]
for i in range(len(list_images)) :
t1.append(list_images[i][-11:-4])
for i in range(len(list_annotations)) :
t2.append(list_annotations[i][-11:-4])
for i in range(len(list_annotations)) :
if t2[i] not in t1 :
print(list_annotations[i])
print(len(list_annotations))
# + id="sXvffpMVG3OC" colab_type="code" colab={}
def get_classes(xml_files=list_annotations):
'''
Input : dataset's annotations
parsing xml data to get objects label from images
Output : class label
'''
classes = []
for file in xml_files:
f = open(file)
doc = xmltodict.parse(f.read()) #parse the xml file to python dict.
# annotation - object - [obj1, obj2 , ...]
try:
for obj in doc['annotation']['object']:
classes.append(obj['name'].lower())
# annotation - object
except TypeError as e:
classes.append(doc['annotation']['object']['name'].lower())
f.close()
classes = list(set(classes))
classes.sort()
return classes
# + id="0W2F4O8XG5ms" colab_type="code" colab={}
classes = get_classes(list_annotations)
print(classes)
num_of_class = len(classes)
print(num_of_class)
# + id="UDwvH-y8G69Y" colab_type="code" colab={}
def get_labels_from_xml(xml_file_path, num_of_class = num_of_class):
'''
Input : 1 xml file
Get class label, gt box coordinates.
Because images are resized to 224x224, coordinates also need to be resized.
Output: Existing class label, ground truth box coordinates
'''
f = open(xml_file_path)
doc = xmltodict.parse(f.read())
ori_img_height = float(doc['annotation']['size']['height'])
ori_img_width = float(doc['annotation']['size']['width'])
class_label = []
bbox_label = []
# multi-objects in image
try:
for each_obj in doc['annotation']['object']:
obj_class = each_obj['name'].lower()
# Get bounding box coordinates
x_min = float(each_obj['bndbox']['xmin']) # top left x-axis coordinate.
x_max = float(each_obj['bndbox']['xmax']) # bottom right x-axis coordinate.
y_min = float(each_obj['bndbox']['ymin']) # top left y-axis coordinate.
y_max = float(each_obj['bndbox']['ymax']) # bottom right y-axis coordinate.
# Images resized to 224x224. So resize the coordinates
x_min = float((image_width/ori_img_width)*x_min)
y_min = float((image_height/ori_img_height)*y_min)
x_max = float((image_width/ori_img_width)*x_max)
y_max = float((image_height/ori_img_height)*y_max)
generated_box_info = [x_min, y_min, x_max, y_max] # [top-left, bottom-right]
index = classes.index(obj_class)
class_label.append(index)
bbox_label.append(np.asarray(generated_box_info, dtype='float32'))
# single-object in image
except TypeError as e :
obj_class = doc['annotation']['object']['name']
x_min = float(doc['annotation']['object']['bndbox']['xmin'])
x_max = float(doc['annotation']['object']['bndbox']['xmax'])
y_min = float(doc['annotation']['object']['bndbox']['ymin'])
y_max = float(doc['annotation']['object']['bndbox']['ymax'])
x_min = float((image_width/ori_img_width)*x_min)
y_min = float((image_height/ori_img_height)*y_min)
x_max = float((image_width/ori_img_width)*x_max)
y_max = float((image_height/ori_img_height)*y_max)
generated_box_info = [x_min, y_min, x_max, y_max]
index = classes.index(obj_class)
class_label.append(index)
bbox_label.append(np.asarray(generated_box_info, dtype='float32'))
return class_label, np.asarray(bbox_label)
xml_file_path=list_annotations[3]
im = cv2.imread(list_images[3])
cv2_imshow(im)
class_label, bbox_label = get_labels_from_xml(xml_file_path, num_of_class = num_of_class)
print(class_label)
print(bbox_label)
# + id="qvoLY11aG8wC" colab_type="code" colab={}
def generate_anchors(one_anchor_size, one_subsampled_ratio,
rpn_kernel_size=rpn_kernel_size, anchor_aspect_ratio=anchor_aspect_ratio):
'''
Input : subsample_ratio (=Pooled ratio)
* 1 anchor size, subsampled ratio are correnpondent to 3 anchor ratio *
generate anchor in feature map. Then project it to original image.
Output : list of anchors (x,y,w,h) and anchor_boolean (ignore anchor if value equals 0)
'''
list_of_anchors = []
anchor_booleans = [] #This is to keep track of an anchor's status. Anchors that are out of boundary are meant to be ignored.
starting_center = divmod(rpn_kernel_size, 2)[0] # rpn kernel's starting center in feature map
anchor_center = [starting_center - 1,starting_center] # -1 on the x-coor because the increment comes first in the while loop
subsampled_height = image_height/one_subsampled_ratio #
subsampled_width = image_width/one_subsampled_ratio #
while (anchor_center != [subsampled_width, subsampled_heigh]): # != [26, 26] -> [N, N]
anchor_center[0] += 1 #Increment x-axis
#If sliding window reached last center, increase y-axis
if anchor_center[0] > subsampled_width - (1 + starting_center):
anchor_center[1] += 1
anchor_center[0] = starting_center
#anchors are referenced to the original image.
#Therefore, multiply downsampling ratio to obtain input image's center
anchor_center_on_image = [anchor_center[0]*one_subsampled_ratio, anchor_center[1]*one_subsampled_ratio]
for size in [one_anchor_size]:
print(size)
for a_ratio in anchor_aspect_ratio:
# [x,y,w,h] --> [y, x, h, w]
anchor_info = [anchor_center_on_image[1], anchor_center_on_image[0], size*a_ratio[1], size*a_ratio[0]]
print(anchor_info)
# check whether anchor crosses the boundary of the image or not
if (anchor_info[0] - anchor_info[2]/2 < 0 or anchor_info[0] + anchor_info[2]/2 > image_width or
anchor_info[1] - anchor_info[3]/2 < 0 or anchor_info[1] + anchor_info[3]/2 > image_height) :
anchor_booleans.append([0.0]) # if anchor crosses boundary, anchor_booleans=0
else:
anchor_booleans.append([1.0])
list_of_anchors.append(anchor_info)
return list_of_anchors, anchor_booleans
def generate_pyramid_anchors(anchor_sizes=anchor_sizes, subsampled_ratio=subsampled_ratio,
rpn_kernel_size=rpn_kernel_size, anchor_aspect_ratio=anchor_aspect_ratio):
list_of_anchors, anchor_booleans=[], []
for i in range(len(anchor_sizes)) :
anchors, bools = generate_anchors(one_subsampled_ratio=subsampled_ratio[i],
one_anchor_size=anchor_sizes[i])
list_of_anchors.append(anchors)
anchor_booleans.append(bools)
return np.concatenate(list_of_anchors, axis=0), np.concatenate(anchor_booleans) # return all of anchors generated from pyramid feature network / shape : N*3
def normalize_box(anchors) :
width=anchors[:,3]-anchors[:,1]
height=anchors[:,2] - anchors[:,0]
scale=np.array([h-1, w-1, h-1, w-1])
shift=np.array([0, 0, 1, 1])
return np.divide((anchors-shift), scale).astype(np.float32)
generate_pyramid_anchors()
# + id="Cy7lR6pLnIFr" colab_type="code" colab={}
import numpy as np
a=np.array([1, 2, 3])
b=np.array([1, 2, 3])
c=[]
c.append(a)
c.append(b)
print(np.concatenate(c))
# + id="joStnOmpHA1c" colab_type="code" colab={}
def generate_label(class_labels, ground_truth_boxes, anchors, anchor_booleans, num_class=num_of_class,
neg_anchor_thresh = neg_threshold, pos_anchor_thresh = pos_threshold):
'''
Input : classes, ground truth box (top-left, bottom-right), all of anchors, anchor booleans.
Compute IoU to get positive, negative samples.
if IoU > 0.7, positive / IoU < 0.3, negative / Otherwise, ignore
Output : anchor booleans (to know which anchor to ignore), objectness label, regression coordinate in one image
'''
number_of_anchors = len(anchors) #Get the total number of anchors.
anchor_boolean_array = np.reshape(np.asarray(anchor_booleans),(number_of_anchors, 1))
# IoU is more than threshold or not.
objectness_label_array = np.zeros((number_of_anchors, 2), dtype=np.float32)
# delta(x, y, w, h)
box_regression_array = np.zeros((number_of_anchors, 4), dtype=np.float32)
# belongs to which object for every anchor
class_array = np.zeros((number_of_anchors, num_class), dtype=np.float32)
for j in range(ground_truth_boxes.shape[0]):
#Get the ground truth box's coordinates.
gt_box_top_left_x = ground_truth_boxes[j][0]
gt_box_top_left_y = ground_truth_boxes[j][1]
gt_box_btm_rght_x = ground_truth_boxes[j][2]
gt_box_btm_rght_y = ground_truth_boxes[j][3]
#Calculate the area of the original bounding box.1 is added since the index starts from 0 not 1.
gt_box_area = (gt_box_btm_rght_x - gt_box_top_left_x + 1)*(gt_box_btm_rght_y - gt_box_top_left_y + 1)
for i in range(number_of_anchors):
######### Compute IoU #########
# Check if the anchor should be ignored or not. If it is to be ignored, it crosses boundary of image.
if int(anchor_boolean_array[i][0]) == 0:
continue
anchor = anchors[i] #Select the i-th anchor [x,y,w,h]
#anchors are in [x,y,w,h] format, convert them to the [top-left-x, top-left-y, btm-right-x, btm-right-y]
anchor_top_left_x = anchor[0] - anchor[2]/2
anchor_top_left_y = anchor[1] - anchor[3]/2
anchor_btm_rght_x = anchor[0] + anchor[2]/2
anchor_btm_rght_y = anchor[1] + anchor[3]/2
# Get the area of the bounding box.
anchor_box_area = (anchor_btm_rght_x - anchor_top_left_x + 1)*(anchor_btm_rght_y - anchor_top_left_y + 1)
# Determine the intersection rectangle.
int_rect_top_left_x = max(gt_box_top_left_x, anchor_top_left_x)
int_rect_top_left_y = max(gt_box_top_left_y, anchor_top_left_y)
int_rect_btm_rght_x = min(gt_box_btm_rght_x, anchor_btm_rght_x)
int_rect_btm_rght_y = min(gt_box_btm_rght_y, anchor_btm_rght_y)
# if the boxes do not intersect, difference = 0
int_rect_area = max(0, int_rect_btm_rght_x - int_rect_top_left_x + 1)*max(0, int_rect_btm_rght_y - int_rect_top_left_y)
# Calculate the IoU
intersect_over_union = float(int_rect_area / (gt_box_area + anchor_box_area - int_rect_area))
# Positive
if intersect_over_union >= pos_anchor_thresh:
objectness_label_array[i][0] = 1.0
objectness_label_array[i][1] = 0.0
#get the class label
class_label = class_labels[j]
class_array[i][int(class_label)] = 1.0 #Denote the label of the class in the array.
#Get the ground-truth box's [x,y,w,h]
gt_box_center_x = ground_truth_boxes[j][0] + ground_truth_boxes[j][2]/2
gt_box_center_y = ground_truth_boxes[j][1] + ground_truth_boxes[j][3]/2
gt_box_width = ground_truth_boxes[j][2] - ground_truth_boxes[j][0]
gt_box_height = ground_truth_boxes[j][3] - ground_truth_boxes[j][1]
#Regression loss / weight
delta_x = (gt_box_center_x - anchor[0])/anchor[2]
delta_y = (gt_box_center_y - anchor[1])/anchor[3]
delta_w = math.log(gt_box_width/anchor[2])
delta_h = math.log(gt_box_height/anchor[3])
box_regression_array[i][0] = delta_x
box_regression_array[i][1] = delta_y
box_regression_array[i][2] = delta_w
box_regression_array[i][3] = delta_h
if intersect_over_union <= neg_anchor_thresh:
if int(objectness_label_array[i][0]) == 0:
objectness_label_array[i][1] = 1.0
if intersect_over_union > neg_anchor_thresh and intersect_over_union < pos_anchor_thresh:
if int(objectness_label_array[i][0]) == 0 and int(objectness_label_array[i][1]) == 0:
anchor_boolean_array[i][0] = 0.0 # ignore this anchor
return anchor_boolean_array, objectness_label_array, box_regression_array, class_array
# + id="m0IThavQHEfv" colab_type="code" colab={}
def anchor_sampling(anchor_booleans, objectness_label, anchor_sampling_amount=anchor_sampling_amount):
'''
Input : anchor booleans and objectness label
fixed amount of negative anchors and positive anchors for training.
If we use all the neg and pos anchors, model will overfit on the negative samples.
Output: Updated anchor booleans.
'''
positive_count = 0
negative_count = 0
for i in range(objectness_label.shape[0]):
if int(objectness_label[i][0]) == 1: #If the anchor is positive
if positive_count > anchor_sampling_amount: #If the positive anchors are more than the threshold amount, set the anchor boolean to 0.
anchor_booleans[i][0] = 0.0
positive_count += 1
if int(objectness_label[i][1]) == 1: #If the anchor is negatively labelled.
if negative_count > anchor_sampling_amount: #If the negative anchors are more than the threshold amount, set the boolean to 0.
anchor_booleans[i][0] = 0.0
negative_count += 1
return anchor_booleans
# + id="nvyt69uEHHU1" colab_type="code" colab={}
def generate_dataset(first_index, last_index, anchors, anchor_booleans):
'''
Input : starting index and final index of the dataset to be generated.
Output: Anchor booleans, Objectness Label and Regression Label in batches.
'''
num_of_anchors = len(anchors)
batch_anchor_booleans = []
batch_objectness_array = []
batch_regression_array = []
batch_class_label_array = []
for i in range(first_index, last_index):
#Get the true labels and the ground truth boxes [x,y,w,h] for every file.
true_labels, ground_truth_boxes = get_labels_from_xml(xml_file_path=list_annotations[i])
# generate_labels for specified batches
anchor_bools, objectness_label_array, box_regression_array, class_array = generate_label(true_labels, ground_truth_boxes,
anchors, anchor_booleans)
#ggenerate_label(class_labels, ground_truth_boxes, anchors, anchor_booleans, num_class=num_of_class,
# neg_anchor_thresh = neg_threshold, pos_anchor_thresh = pos_threshold)
# get the updated anchor bools based on the fixed number of sample
anchor_bools = anchor_sampling(anchor_bools, objectness_label_array)
batch_anchor_booleans.append(anchor_bools)
batch_objectness_array.append(objectness_label_array)
batch_regression_array.append(box_regression_array)
batch_class_label_array.append(class_array)
batch_anchor_booleans = np.reshape(np.asarray(batch_anchor_booleans), (-1,num_of_anchors)) # (1, 6084, 1) -> (1, 6084)
batch_objectness_array = np.asarray(batch_objectness_array)
batch_regression_array = np.asarray(batch_regression_array)
batch_class_label_array = np.asarray(batch_class_label_array)
return (batch_anchor_booleans, batch_objectness_array, batch_regression_array, batch_class_label_array)
# + id="wqQCGHU5HJEe" colab_type="code" colab={}
def read_images(first_index, last_index):
'''
Read the image files, then resize.
Input : first and last index.
Output: numpy array of images.
'''
images_list = []
for i in range(first_index, last_index):
im = cv2.imread(list_images[i])
im = cv2.resize(im, (image_height, image_width))/255
images_list.append(im)
return np.asarray(images_list)
# + id="isM7-kOFHKym" colab_type="code" colab={}
anchors, an_bools = generate_anchors() #We only need to generate the anchors and the anchor booleans once.
num_of_anchors = len(anchors)
# + id="4bo78X7oHM1C" colab_type="code" colab={}
a,b,c,d = generate_dataset(0,1, anchors, an_bools)
a.shape
print(a)
# + id="P9X2aZphHN18" colab_type="code" colab={}
learning_rate = 1e-5
epoch = 100
batch_size = 10
# !mkdir -p '/content/drive/My Drive/VOCdata/trained_weight'
model_checkpoint = './drive/My Drive/VOCdata/trained_weight/model.ckpt'
decay_steps = 10000
decay_rate = 0.99
lambda_value = 10
# + id="acSqLwUD-kLx" colab_type="code" colab={}
def smooth_func(t):
t = tf.abs(t)
comparison_tensor = tf.ones((num_of_anchors, 4))
smoothed = tf.where(tf.less(t, comparison_tensor), 0.5*tf.pow(t,2), t - 0.5)
return smoothed
# + id="AlqpqYl8-ltY" colab_type="code" colab={}
def smooth_L1(pred_box, truth_box):
diff = pred_box - truth_box
smoothed = tf.map_fn(smooth_func, diff)
return smoothed
# + id="h7eN-azgsAog" colab_type="code" colab={}
def identity_block(input, filter_size, filter_num) :
y=layers.Conv2D(filter_num[0], (1, 1), strides=(1, 1), padding='valid', use_bias=True)(input)
y=layers.BatchNormalization(y)
y=layers.Activation('relu')(y)
y=layers.Conv2D(filter_num[1], (3, 3), strides=(1, 1), padding='same', use_bias=True)(y)
y=layers.BatchNormalization(y)
y=layers.Activation('relu')(y)
y=layers.Conv2D(filter_num[2], (1, 1), strides=(1, 1), padding='valid', use_bias=True)(y)
y=layers.BatchNormalization(y)
y=layers.Add()([y, input])
y=layers.Activation('relu')(y)
return y
def conv_block(input, filter_num, strides=(2, 2)) :
y=layers.Conv2D(filter_num[0], (1, 1), strides=strides, padding='valid', use_bias=True)(input)
y=layers.BatchNormalization(y)
y=layers.Activation('relu')(y)
y=layers.Conv2D(filter_num[1], (3, 3), strides=(1, 1), padding='same', use_bias=True)(y)
y=layers.BatchNormalization(y)
y=layers.Activation('relu')(y)
y=layers.Conv2D(filter_num[2], (1, 1), strides=(1, 1), padding='valid', use_bias=True)(y)
y=layers.BatchNormalization(y)
# add shortcut block for conv_block
shortcut=layers.Conv2D(filter_num[2], (1, 1), strides=strides, padding='valid', use_bias=True)(input)
shortcut=layers.BatchNormalization(shortcut)
y=layers.Add()([y, shortcut])
y=layers.Activation('relu')(shortcut)
return y
def resnet101(input) :
# stage 1
y=layers.ZeroPadding2D((3, 3))(input)
y=layers.Conv2D(64, (7, 7), strides=(2, 2), padding='valid', use_bias=True)(y)
y=layers.BatchNormalization(y)
y=layers.Activation('relu')(y)
F1=y=layers.MaxPooling2D((3, 3), strides(2, 2), padding='same')(y)
# stage 2 : In stage 2, use strides 1
y=conv_block(y, [64, 64, 256], strides=(1, 1))
y=identity_block(y, [64, 64, 256])
F2=y=identity_block(y, [64, 64, 256])
# stage 3
y=conv_block(y, [128, 128, 512])
y=identity_block(y, [128, 128, 256])
y=identity_block(y, [128, 128, 256])
F3=y=identity_block(y, [128, 128, 256])
# stage 4 : In stage 4, there are 22 identity blocks
y=conv_block(y, [256, 256, 1024])
block_num=22
for i in range(block_num) :
y=identity_block(y, [256, 256, 1024])
F4=y
# stage 5
y=conv_block(y, [512, 512, 2048])
y=identity_block(y, [512, 512, 2048])
F5=y=identity_block(y, [512, 512, 2048])
return [F1, F2, F3, F4, F5]
# Use 256 filters for FPN
def fpn(input) :
P5=layers.Conv2d(256, (1, 1))(F5)
P5=layers.Conv2D(256, (3, 3), padding='same')(P5)
# Compute P4 by adding feature map4 and Upsampled result of P5
res4=layers.UpSammpling2D(size=(2, 2))(P5)
P4=layers.Add([res4, layers.Conv2D(256, (1, 1)(F4))])
P4=layers.Conv2D(256, (3, 3), padding='same')(P4)
res3=layers.UpSammpling2D(size=(2, 2))(P4)
P3=layers.Add([res4, layers.Conv2D(256, (1, 1)(F3))])
P3=layers.Conv2D(256, (3, 3), padding='same')(P3)
res2=layers.UpSammpling2D(size=(2, 2))(P3)
P2=layers.Add([res4, layers.Conv2D(256, (1, 1)(F2))])
P2=layers.Conv2D(256, (3, 3), padding='same')(P2)
#Issues : Why use (1, 1) for pool size and reason for not computing (3, 3) conv only in P6
P6=layers.MaxPooling2D(pool_size=(1, 1), strides=2)(P5)
return [P2, P3, P4, P5, P6]
# + id="eDEFMKcd-m9-" colab_type="code" colab={}
def rpn(input) :
# All of feature map's depth in FPN is 256
input_featuremap=layers.Input(shape=[None, None, 256])
shared4obj_and_bb=layers.Conv2d(512, (3, 3), padding='same', activation_fn='relu', strides=1)(input)
obj_conv = layers.Conv2d(6, (1, 1), padding='VALID', activation_fn='linear')(shared4obj_and_bb) # None * None * 6
bb_conv = layers.Conv2d(12, (1, 1), padding='VALID', activation_fn='linear')(shared4obj_and_bb) # None * None * 12
class_conv_reshape = tf.reshape(obj_conv, (-1, num_of_anchors, 2)) # 6084x2
anchor_conv_reshape = tf.reshape(bb_conv, (-1, num_of_anchors, 4)) # 6084x4
logits = tf.nn.softmax(class_conv_reshape)
return models.Model([input_featuremap], [class_conv_reshape, logits, anchor_conv_reshape])
# + id="gwyKXQFxeLBn" colab_type="code" colab={}
def apply_box_delta(boxes, delta) :
'''
Input : boxes [N, [y1, x1, y2, x2]]
boxes = anchor boxes (ranked top k of All anchors)
delta = correspondents to anchor boxes' delta (also ranked top k anchors' delta)
apply delta to boxes (* refined boxes' coordinates represent normalized coordinates)
'''
y1, x1, y2, x2 = boxes[:,0], boxes[:,1], boxes[:,2], boxes[:,3]
anchor_height = y2 - y1
anchor_width = x2 - x1
anchor_center_x = x1 + width*0.5
anchor_center_y = y1 + height*0.5
# apply delta to box
refined_center_x = delta[:,1] * width + anchor_center_x
refined_center_y = delta[:,0] * height + anchor_center_y
refined_width = tf.exp(delta[:,3]) * anchor_width
refined_height = tf.exp(delta[:,2]) * anchor_height
# compute refined coordinates : [y1, x1, y2, x2]
refined_x1=refined_center_x - 0.5*width
refined_x2=refined_center_x + 0.5*width
refined_y1=refined_center_y - 0.5*height
refined_y2=refined_center_y + 0.5*height
return tf.stack([refined_y1, refined_x1, refined_y2, refined_x2], axis=1)
# + id="_ql51sMdjPAH" colab_type="code" colab={}
def compute_iou_boxes(one_box, boxes, box_area, boxes_area) :
intersection_y1 = np.maximum(box[0], boxes[:, 0])
intersection_x1 = np.maximum(box[1], boxes[:, 1])
intersection_y2 = np.minimum(box[2], boxes[:, 2])
intersection_x2 = np.minimum(box[3], boxes[:, 3])
inter_area = np.maximum(intersection_x2-intersection_x1, 0) * np.maximum(intersection_y1-intersection_y2, 0)
union_area = box_area + boxes_area[:] - inter_area
iou = inter_area / union_area
return iou
def non_max_suppression(boxes, scores) :
# scores : result from softmax of rpn's output(logits)
y1=boxes[:, 0]
x1=boxes[:, 1]
y2=boxes[:, 2]
x2=boxes[:, 3]
area=(y2-y1)*(x2-x1)
# Use fg score to sort
sorted_idx = scores.argsort()[::-1] # return idx of sorted result
result = []
while len(ixs) > 0:
# Pick top box and add its index to the list
i = sorted_idx[0]
pick.append(i)
# Compute IoU of the picked box with the rest
iou = compute_iou(boxes[i], boxes[ixs[1:]], area[i], area[ixs[1:]])
# Identify boxes with IoU over the 0.7(nms_threshold).
# This returns are correspondents to ixs[1:], so add 1 (ixs are correspondents to ixis[:]).
remove_ixs = np.where(iou > nms_threshold)[0] + 1
# Remove indices of the picked and overlapped boxes.
ixs = np.delete(ixs, remove_ixs)
ixs = np.delete(ixs, 0)
nms_roi = tf.gather(boxes, result)
return nms_roi
# + id="m2JYUXi3-p0a" colab_type="code" colab={}
# Loss Function
loss1 = 1/256*tf.reduce_sum(anch_bool*(tf.nn.softmax_cross_entropy_with_logits(labels=Y_obj, logits=class_conv_reshape))) # positive(128) + negative(128)
# (10, 6084, 2)
loss2 = lambda_value*(1/128)*tf.reduce_sum((tf.reshape(Y_obj[:,:,0], (-1,num_of_anchors,1)))*smooth_L1(anchor_conv_reshape, Y_coor))
total_loss = loss1 + loss2
optimizer = tf.train.AdamOptimizer(decayed_lr).minimize(total_loss, global_step=global_step)
global_step = tf.Variable(0, trainable=False)
decayed_lr = tf.train.exponential_decay(learning_rate,
global_step, decay_steps,
decay_rate, staircase=True)
# + id="M8GmIAcv-wKo" colab_type="code" colab={}
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
saver = tf.train.Saver()
try:
saver.restore(sess, model_checkpoint)
print("Model has been loaded!")
except:
print("Model doens't exist!")
# + id="MI3fc5DQ-ylY" colab_type="code" colab={}
'''
def draw_a_rectangel_in_img(draw_obj, box, color, width):
'''
use draw lines to draw rectangle. since the draw_rectangle func can not modify the width of rectangle
:param draw_obj:
:param box: [x1, y1, x2, y2]
:return:
'''
x1, y1, x2, y2 = box[0], box[1], box[2], box[3]
top_left, top_right = (x1, y1), (x2, y1)
bottom_left, bottom_right = (x1, y2), (x2, y2)
draw_obj.line(xy=[top_left, top_right],
fill=color,
width=width)
draw_obj.line(xy=[top_left, bottom_left],
fill=color,
width=width)
draw_obj.line(xy=[bottom_left, bottom_right],
fill=color,
width=width)
draw_obj.line(xy=[top_right, bottom_right],
fill=color,
width=width)
PIXEL_MEAN = [123.68, 116.779, 103.939]
def draw_boxes_with_label_and_scores(img_array, boxes):
img_array = img_array + np.array(PIXEL_MEAN)
img_array.astype(np.float32)
boxes = boxes.astype(np.int64)
#labels = labels.astype(np.int32)
img_array = np.array(img_array * 255 / np.max(img_array), dtype=np.uint8)
img_obj = Image.fromarray(img_array)
#img_obj=img_array
print('zz')
raw_img_obj = img_obj.copy()
draw_obj = ImageDraw.Draw(img_obj)
num_of_objs = 0
for box in boxes:
draw_a_rectangel_in_img(draw_obj, box, color='Coral', width=3)
out_img_obj = Image.blend(raw_img_obj, img_obj, alpha=0.6)
return np.array(out_img_obj)
'''
#TRAINING
for epoch_idx in range(epoch): #Each epoch.
#Loop through the whole dataset in batches.
for start_idx in tqdm(range(0, total_images, batch_size)):
end_idx = start_idx + batch_size
if end_idx >= total_images : end_idx = total_images - 1 #In case the end index exceeded the dataset.
images = read_images(start_idx, end_idx) #Read images.
#Get the labels needed.
batch_anchor_booleans, batch_objectness_array, batch_regression_array, _ = \
generate_dataset(start_idx,end_idx, anchors, an_bools)
print(batch_objectness_array.shape)
#Optimize the model.
anchor_reshape, _, theloss = sess.run([anchor_conv_reshape, optimizer, total_loss], feed_dict={X: images,
Y_obj:batch_objectness_array,
Y_coor: batch_regression_array,
anch_bool: batch_anchor_booleans})
'''
img_array = cv2.imread(list_images[0])
#img_array = cv2.resize(img_array, (image_height, image_width))/255
img_array = np.array(img_array, np.float32) - np.array(PIXEL_MEAN)
anchor_booleans, objectness_array, regression_array, _ = \
generate_dataset(0,1, anchors, an_bools)
img_array_tensor=read_images(0, 1)
anchor_reshape=sess.run([anchor_conv_reshape], feed_dict={X:img_array_tensor, Y_obj : objectness_array, Y_coor : regression_array, anch_bool : anchor_booleans})
print(anchor_reshape)
print(anchor_reshape[0])
anchor_reshape=anchor_reshape[0]
boxes = np.array(
[[200, 200, 500, 500],
[300, 300, 400, 400],
[200, 200, 400, 400]]
)
anchor_reshape=np.array(anchor_reshape)
im=draw_boxes_with_label_and_scores(img_array, np.array(anchor_reshape))
cv2_imshow(im)
'''
#Save the model periodically.
saver.save(sess, model_checkpoint)
print("Epoch : %d, Loss : %g"%(epoch_idx, theloss))
img_array = cv2.imread(list_images[0])
#img_array = cv2.resize(img_array, (image_height, image_width))/255
img_array = np.array(img_array, np.float32) - np.array(PIXEL_MEAN)
anchor_booleans, objectness_array, regression_array, _ = \
generate_dataset(0,1, anchors, an_bools)
img_array_tensor=read_images(0, 1)
anchor_reshape=sess.run([anchor_conv_reshape], feed_dict={X:img_array_tensor, Y_obj : objectness_array, Y_coor : regression_array, anch_bool : anchor_booleans})
print(anchor_reshape)
print(anchor_reshape[0])
anchor_reshape=anchor_reshape[0][0]
boxes = np.array(
[[200, 200, 500, 500],
[300, 300, 400, 400],
[200, 200, 400, 400]]
)
im=draw_boxes_with_label_and_scores(img_array, np.array(anchor_reshape))
cv2_imshow(im)
# + id="PBbxs5h5bvKz" colab_type="code" colab={}
| Mask_RCNN(RPN).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="wDKRRCv3Dc82"
# # Exercício 01 - PEL201
#
# ## Exercício
# Implemente o algoritmo de Máximo Divisor Comum (MDC) de dois números (a e b) de dois modos: usando a solução iterativa (que percorre as possibilidades) e a solução recursiva.
#
# ### Pequeno texto sobre o que é e como encontrar, computacionalmente, o MDC
# O Máximo Divisor Comum é o maior número que divide dois inteiros em questão. Para encontrá-lo computacionalmente existem duas possbilidades:
#
# 1. Utilizando um algoritmo iterativo que irá percorrer todas as opções até encontrar a melhor.
#
# * Neste cenário é importante compreender que a maioria dos testes resultarão em falha, principalmente se estivermos avaliando números primos. Ou seja, este algoritmo passa a ser pouco eficiente quanto maior forem os números sendo avaliados;
#
#
# 2. A segunda opção é se utilizar de um algoritmo criado por Euclides, aproximadamente em 300 a.C..
#
# * Este algoritmo consiste em subtrair o menor número do maior sucessivamente até chegar em zero. Quando esta convergência ocorrer o MDC será o outro número inteiro. Desta forma podemos calcular o MDC recursivamente utilizando o resto da divisão como entrada para a função até que o resto seja igual a 0.
# + id="qxB6xRTZDW7H"
import time
from tqdm import tqdm
import matplotlib.pyplot as plt
import numpy as np
# + id="BV5Aho-kDiXL"
def read_random_primes():
f = open("./top_primes.txt", "r")
first_half = []
second_half = []
i = 0
for prime in f:
if i < 50000:
first_half.append(int(prime.replace("\n", "")))
else:
second_half.append(int(prime.replace("\n", "")))
i = i + 1
pairs = zip(first_half, second_half)
return pairs
# -
# ### Explique, em palavras, como é a solução iterativa (e depois apresente o código, identificando qual linguagem usou)
# Linguagem: `Python`
#
# O algoritmo iterativo inicia definindo o menor número dentre as duas entradas como primeiro candidato. Após isso, iremos iterar entre todos os números até que o resto da divisão seja zero. Neste caso iremos repetir esta avaliação, no melhor caso `1` vez (isso se o `min(m, n)` for exatamente o MDC) e no pior caso `min(m, n)`.
# Desta forma o número de iterações irá crescer proporcionalmente ao tamanho das entradas.
# + id="BAM7WZbqDlQY"
def mdc_iterativo(m=15485863, n=15485857):
candidato = min(m, n)
while m % candidato != 0 or n % candidato != 0:
candidato = candidato - 1
return candidato
# -
# ### Explique, em palavras, como é a solução recursiva (e depois apresente o código, identificando qual linguagem usou)
# Linguagem: `Python`
#
# Este é o algoritmo de Euclides para encontrar o MDC de dois números inteiros. Como este algoritmo é recursivo, a primeira avaliação é feita para garantir o critério de parada. Isto feito realizamos uma nova chamada à função porém utilizando o resto da divisão como entrada para o segundo parâmetro. Isso será realizado até que a função atinja o critério de parada que é quando o resto da divisão for igual a 0.
#
# Utilizando-se deste algoritmo, a quantidade de execuções nunca será maior do que 5 vezes o tamanho de dígitos do menor número.
# + id="73K_Hob1DuSj"
def mdc_recursivo(m=15485863, n=15485857):
if n == 0:
return m
return mdc_recursivo(n, m % n)
# + id="piwyateMIYk9"
items = read_random_primes()
items = list(items)
# + colab={"base_uri": "https://localhost:8080/"} id="k4tIvvBwDv9P" outputId="3c77aea6-8551-44a7-93a1-bbe57aec9317"
iter_runtime_result = []
for item in tqdm(items):
mdc_iter_start = time.time()
mdc_iter_result = mdc_iterativo(item[0], item[1])
mdc_iter_end = time.time()
iter_runtime_result.append(mdc_iter_end - mdc_iter_start)
# + colab={"base_uri": "https://localhost:8080/"} id="RnzPvPiBIV7T" outputId="5c80bf6a-ba87-406d-b5b0-40e4aeec6e78"
recur_runtime_result = []
for item in tqdm(items):
mdc_recur_start = time.time()
mdc_recur_result = mdc_recursivo(item[0], item[1])
mdc_recur_end = time.time()
recur_runtime_result.append(mdc_recur_end - mdc_recur_start)
# -
# ### Faça uma comparação em gráfico e tabela dos desempenhos de tempo (em segundos) entre elas para várias entradas iguais (faça entradas com números grandes e/ou rode o mesmo algoritmo várias vezes para capturar o tempo)
#
# Utilizei um dataset com os primeiros 100 mil de números primos. Com esta lista montei 50.000 pares e utilizei como dados de teste.
#
# Cada um dos 50.000 pares foram testados com o método iterativo e com o método recursivo.
#
# #### Resultados
#
# Antes de apresentar os resultados dos testes é importante fazer a avaliação conceitual de ambos os algoritmos para compreender a diferença na eficiência.
#
# Vamos comparar os algoritmos utilizando como entrada os seguintes números primos: `m=15485863` e `n=15485857`
#
# * Utilizando o algoritmo iterativo teremos, no pior caso, `15.485.857` iterações;
#
# * Utilizando o algoritmo recursivo teremos, no pior caso, `5 * 8 = 40`. Ou seja, 5 vezes os 8 digitos do menor número, resultando em 40 passos.
#
# Algoritmo Iterativo: `100%|██████████| 50000/50000 [1:09:27<00:00, 12.00it/s]`
#
# Algoritmo Recursivo: `100%|██████████| 50000/50000 [00:00<00:00, 85873.26it/s]`
#
# Ao realizar o teste foi possível perceber que o método iterativo é 99,98% mais lento que o método recursivo. Podemos perceber esta diferença através do gráfico abaixo, onde a curva de tempo de execução do método iterativo se aproxima de uma função linear enquanto a curva de tempo de execução do método recursivo se aproxima de uma curva constante, sendo assim muito mais eficiente do que o anterior.
print("Média - Tempo Algoritmo Iterativo (s): " + str(np.average(iter_runtime_result)))
print("Média - Tempo Algoritmo Recursivo (s): " + str(np.average(recur_runtime_result)))
((np.average(iter_runtime_result) - np.average(recur_runtime_result)) / np.average(iter_runtime_result)) * 100
# + colab={"base_uri": "https://localhost:8080/", "height": 268} id="ttuUgLc0Dze8" outputId="13cd415c-4e91-4ce4-b3b4-d0158523ec2a"
fig, ax = plt.subplots()
line1, = ax.plot(iter_runtime_result, label='Iterativo')
line2, = ax.plot(recur_runtime_result, label='Recursivo')
ax.legend()
plt.show()
# -
# ### Apresente e EXPLIQUE a recorrência T(n) de cada um dos algoritmos implementados
#
# #### Algoritmo Iterativo
#
# ```python
# def mdc_iterativo(m=15485863, n=15485857):
# candidato = min(m, n)
#
# while m / candidato != 0 or n / candidato != 0:
# candidato = candidato - 1
#
# return candidato
# ```
#
# Linha 1: `candidado = min(m, n)` => C * 1
#
# Linha 2: `while m % candidato != 0 or n % candidato != 0:` => C * n + 1
#
# Linha 3: `candidado = candidado - 1` => C * n
#
# Linha 4: `return candidato` => C * 1
#
# Resultado: $$T(n) = 4c + 2n + 1$$
#
# Assumindo `c = 1`, temos: $$T(n) = 2n + 5$$
#
#
# #### Algoritmo Recursivo
#
# ```python
# def mdc_recursivo(m=15485863, n=15485857):
# if n == 0:
# return m
#
# return mdc_recursivo(n, m % n)
# ```
#
# Linha 1: `if n == 0` => C * 1
#
# Linha 2: `return m` => C * 1
#
# Linha 3: `return mdc_recursivo(n, m % n)` => $$T(m / n) + \theta$$
#
# Logo a recorrência deste algoritmo é:
#
# $$ \theta(1) $$
#
# $$ T(m / n) + \theta(1)$$
| PEL201_01_MDC-v1.ipynb |