code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(style="ticks", font_scale=1.5)
sns.set_context("poster")
import ipywidgets as widgets
from ipywidgets import interactive
# +
# %matplotlib inline
def f(Tmiddle, deltaT, show_deltas, show_tangent):
fig, ax = plt.subplots(figsize=(8,6))
t = np.linspace(0,20,101)
foo = lambda t: (t**3 - 15*t**2 + 300) / 100
x = foo(t)
ax.plot(t, x, color="red", lw=3)
ax.set(xlim=[0,20], ylim=[-5,20], ylabel="y", xticks=np.arange(0,21,5))
ax.set_xlabel("time")
ax.set_ylabel("x(t)", rotation="horizontal",labelpad=15)
t0 = Tmiddle - deltaT/2
t1 = Tmiddle + deltaT/2
x0 = foo(t0)
x1 = foo(t1)
ax.plot([t0,t1], [x0,x1], color="blue", lw=2)
if show_deltas:
ax.plot([t0,t1], [x0,x0], color="gray")
ax.plot([t1,t1], [x0,x1], color="gray")
ax.text(Tmiddle, x0, r"$\Delta t$", fontsize=20,va="top")
ax.text(t1, (x1+x0)/2, r"$\Delta x$", fontsize=20)
if show_tangent:
slope = (3*Tmiddle**2 - 30*Tmiddle) / 100
intercept = foo(Tmiddle) - slope*Tmiddle
Tmin = -100
Tmax = 100
line = lambda t: slope*t + intercept
ax.plot([Tmin, Tmax], [line(Tmin), line(Tmax)],
color="black", ls=":")
interactive_plot = interactive(f, Tmiddle=widgets.FloatSlider(min=1, max=17, step=0.5, value=15),
deltaT=widgets.FloatSlider(min=0, max=8, step=0.2, value=5),
show_deltas=widgets.Checkbox(value=True),
show_tangent=widgets.Checkbox(value=True))
output = interactive_plot.children[-1]
# to avoid flickering: https://ipywidgets.readthedocs.io/en/latest/examples/Using%20Interact.html#Flickering-and-jumping-output
output.layout.height = '450px'
interactive_plot
| archive/physics/lectures/instantaneous_velocity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial: retrieve Gaia EDR3 sources
#
# This notebook illustrates a method to retrieve sources from the Gaia EDR3 catalog.
# The sources within 1 degree around $(\mathrm{ra}, \mathrm{dec}) = (269.27, -18.99)$ are extracted.
# The retrived sources are stored as a pickle dump file.
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pickle as pkl
import warpfield as w
# The center of the search region is specified by a `SkyCoord` instance.
# The radius of the search region is given by an `Angle` instance.
from astropy.coordinates import SkyCoord, Longitude, Latitude, Angle
import astropy.units as u
lon = Longitude(269.267, unit=u.degree)
lat = Latitude(-18.985, unit=u.degree)
radius = Angle(1.0*u.degree)
pointing = SkyCoord(lon, lat, frame='icrs')
# The Gaia EDR3 sources are fetched via `retrieve_gaia_sources`.
# The first and second arguments are the central position and the radius of the search region.
# The sources with `parallax_over_error` larger than 10 are retrieved.
# This possibly takes a few minute to retrieve the sources.
gaia_sources = w.retrieve_gaia_sources(pointing, radius)
# About 18000 sources are retrieved.
print(len(gaia_sources))
# The sources are given as a `SkyCoord` instance. `ra`, `dec`, `pm_ra_cosdec`, and `pm_dec` are imported from the Gaia EDR3 catalog. `distance` is given by a reciprocal of `parallax`.
print(gaia_sources)
# Save the retrieved catalog as `gaia_edr3.pkl`.
with open('gaia_edr3.pkl','wb') as f:
pkl.dump(gaia_sources, f)
| notebook/tutorial_01_retrieve_gaia_edr3_catalog.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
from reconciler import reconcile
import pandas as pd
original_data = "../../analysis/data/panglaodb/"
results_path = "../../analysis/results/true_matches/"
original_cells_organs = pd.read_csv(f"{original_data}cells_organs_germlayers.csv")
original_cells_organs.head()
# -
cell_types = original_cells_organs["cell_type"]
# Reconcile against cell types on Wikidata
reconciled = reconcile(cell_types, type_id="Q189118")
# Now I`ll check the matches labeled as "true" for false positives.
cell_type_matches_considered_true = reconciled[reconciled["match"] == True]
cell_type_matches_considered_true.to_csv("../results/cell_type_matches_considered_true_29_10_2020.csv")
# I have saved the file as a csv and opened on LibreOffice. I am looking at each match for inconsistencies and will note down any inconsistencies found.
#
# - Dopaminergic neuron is matched to "dopaminergic cell group", and the item is a bit weird, it uses info from the Foundational Model of Anatomy (FMA). There is an item for "dopaminergic neuron" on Wikitada (Q66591220) with a different FMA identifier. Arguably is a better match, so I`ll use that instead.
#
#
# Modifications to make:
#
# - Dopaminergic neurons: Q66591220
#
#
# I`ll prepare a verified dict for name to qid reconciliation of cell types.
#
#
# +
panglao_cell_type_to_qid = {}
for i, row in cell_type_matches_considered_true.iterrows():
name = row["input_value"]
qid = row["id"]
panglao_cell_type_to_qid[name] = qid
# Fix manually false positive match
panglao_cell_type_to_qid["Dopaminergic neurons"] = "Q66591220"
# -
# Nice. Now let`s look at the matches that were found, but considered false.
# +
cell_type_matches_considered_false = reconciled[reconciled["match"] == False]
cell_type_matches_considered_false = cell_type_matches_considered_false[cell_type_matches_considered_false["id"].notna()]
cell_type_matches_considered_false.to_csv("../results/cell_type_matches_considered_false_29_10_2020.csv")
# -
# Once again I open it with LibreOffice. Now I will have to correc most of them.
#
# - Perisynaptic Schwann cells (Q17157019) seems to be too specific. Schwann cell (Q465621) is the better match.
# - radial glial cell (Q4387286) is similar, but different of Astrocyte progenitor cells. I have created an item for astrocyte progenitor cells (Q101001053).
# - Adaptive NK cells (Q49000020) is too specific. natural killer cell (Q332181) is the better match.
# - Endoplasmic Reticulum Stress in Beta Cells (Q5376385) is just a topic. beta cell (Q1767180) is the better match.
# - Perineuronal satellite cells (Q70068158) is a wrong match. Satellite cells in PanglaoDB are "precursors to skeletal muscle cells, able to give rise to satellite cells or differentiated skeletal muscle cells." There is a "Satellite cell (Q66592694)" but I am not sure if that is what we want. I`ll create a new term.
# - subependymal glioma (Q18556497) is a wrong match for "Ependymal cells." ependymal cell (Q28646122) is the better match.
# - cardiomyocyte (Q1047227) is too specific for "Myocytes". myocyte (Q428914) is a better match.
# - natural killer T cell (Q224930) is similar, but different from "T cells". T-lymphocytes (Q193529) is a better match.
#
#
#
# As there is a large number of mismatches, I`ll plainly add the ones that need modification to the list below. I`ll note any case that is different from the ones above and any creation of new
#
#
#
#
#
# Modifications to make:
#
# - Schwann cells: Q465621
# - Astrocyte progenitor cells: Q101001053
# - NK cells: Q332181
# - Beta cells: Q1767180
# - Satellite cells: Q101001061
# - Ependymal cells: Q28646122
# - Myocytes: Q428914
# - T cells: Q193529
#
#
#
#
| improvements/src/check_and_add_missing_cell_types_29_10_2020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
soma=0
cont=0
for i in range(1,501,2):
if i%3==0:
cont += 1
soma += i
print('A soma de todos os {} valores solicitados é {}'.format(cont,soma))
| .ipynb_checkpoints/EX048 - Soma Ímpares Múltiplos de Três-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: NV2
# language: python
# name: nv2
# ---
# # Intro
# This notebook serves as a tiny example on how to train a DNN using TensorFlow and export to the NNet format that NeuralVerification.jl can parse.
# # Requirements
# Before we begin make sure your environment satisfies the requirements specified in `requirements.txt`, the easiest way is to create a new virtualenv and run:
#
# `pip install -r requirements.txt`
#
# then you can go ahead and run the following cells, for this make sure that your jupyter notebook is using the kernel corresponding to the virtualenv you just created (Chris: should I expand on this?)
# # imports needed
from tensorflow.examples.tutorials.mnist import input_data
import tensorflow as tf
import math
import numpy as np
# # fetch and load the data
mnist = input_data.read_data_sets('./mnist_data')
model_name = "mnist_network"
# # define auxiliary functions
# +
def weight_variable(shape):
initial = tf.truncated_normal(shape, stddev=0.1)
return tf.Variable(initial)
def bias_variable(shape):
initial = tf.constant(0.1, shape=shape)
return tf.Variable(initial)
# -
# # define the TF variables
# +
INPUT = 784 # mnist images are 28 x 28 = 784 pixels
OUTPUT = 10 # there are 10 possible classes for each image (0-9 digits)
HFC = 50 # hidden layer size
# placeholder used to feed in the input, it is very important to explicitly name
# which will enable the script that generates the nnet file from the protobuffer
# to identify the relevant part of the computational graph
x_in = tf.placeholder(tf.float32, [None, INPUT], name='input_op')
W_fc = weight_variable([INPUT, HFC])
b_fc = bias_variable([HFC])
W_fc1 = weight_variable([HFC, HFC])
b_fc1 = bias_variable([HFC])
#W_fc2 = weight_variable([HFC, HFC])
#b_fc2 = bias_variable([HFC])
#W_fc3 = weight_variable([HFC, HFC])
#b_fc3 = bias_variable([HFC])
#W_fc4 = weight_variable([HFC, HFC])
#b_fc4 = bias_variable([HFC])
#W_fc5 = weight_variable([HFC, HFC])
#b_fc5 = bias_variable([HFC])
W_o = weight_variable([HFC, OUTPUT])
b_o = bias_variable([OUTPUT])
#
x_image = tf.reshape(x_in, [-1, int(math.sqrt(INPUT)), int(math.sqrt(INPUT)), 1])
# -
# # define the network
h_fc = tf.nn.relu(tf.add(tf.matmul(x_in, W_fc), b_fc))
h_fc1 = tf.nn.relu(tf.add(tf.matmul(h_fc, W_fc1), b_fc1))
#h_fc2 = tf.nn.relu(tf.add(tf.matmul(h_fc1, W_fc2), b_fc2))
#h_fc3 = tf.nn.relu(tf.add(tf.matmul(h_fc2, W_fc3), b_fc3))
#h_fc4 = tf.nn.relu(tf.add(tf.matmul(h_fc3, W_fc4), b_fc4))
#h_fc5 = tf.nn.relu(tf.add(tf.matmul(h_fc4, W_fc5), b_fc5))
y = tf.add(tf.matmul(h_fc1, W_o), b_o, name="output_op")
# # defining the loss and optimizer
y_ = tf.placeholder(tf.int64, [None])
cross_entropy = tf.losses.sparse_softmax_cross_entropy(labels=y_, logits=y)
train_step = tf.train.AdamOptimizer(0.0001).minimize(cross_entropy)
# +
epochs = 10 # epochs for training
batch_size = 64 # batch size
N = mnist.train.num_examples
batch_amount = epochs*N//batch_size
# -
# # Train and save protobuffer
# +
epochs = 10 # epochs for training
batch_size = 50 # batch size
N = mnist.train.num_examples
batch_amount = epochs*N//batch_size
with tf.Session() as sess:
epoch_in = 1
tf.global_variables_initializer().run()
for i in range(batch_amount):
batch_xs, batch_ys = mnist.train.next_batch(batch_size)
sess.run(train_step, feed_dict={x_in: batch_xs, y_: batch_ys})
if (i*batch_size) % N == 0:
#print('loss> {}'.format(sess.run(cross_entropy, feed_dict={x_in: batch_xs, y_: batch_ys})))
print("Epoch {} done.".format(epoch_in))
epoch_in += 1
correct_prediction = tf.equal(tf.argmax(y, 1), y_)
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
print("Accuracy: {}".format(sess.run(accuracy, feed_dict={x_in: mnist.test.images, y_: mnist.test.labels})))
output_graph_def = tf.graph_util.convert_variables_to_constants(sess, tf.get_default_graph().as_graph_def(), ['output_op'])
#saver = tf.train.Saver()
#save_path = saver.save(sess, 'models/'+model_name+'.ckpt')
with tf.gfile.GFile('models/' + model_name+'.pb', "wb") as f:
f.write(output_graph_def.SerializeToString())
# -
# # Now let's convert the protobuffer to NNet format
# !python ./NNet/scripts/pb2nnet.py
| examples/cars_workshop/1 training-proto-nnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import pandas_datareader as pdr
nvda = pdr.get_data_yahoo('NVDA', '20181231')
nvda.info()
# drop unwanted column if try to run cell more than once keyerror
nvda.drop(columns=['Adj Close'], inplace=True)
nvda = round(nvda,2)
nvda['Change'] = nvda['Close'] - nvda['Close'].shift(1)
nvda.head()
import numpy as np
nvda['Return'] = np.log(nvda['Close']).diff()
nvda.head()
nvda['Close'].pct_change()
nvda['Volatility'] = nvda['Return'].rolling(21).std().shift(1)
nvda.head(10)
# cleanup missing values
nvda.dropna(inplace=True)
nvda.head()
nvda['Exp Change'] = nvda['Volatility'] * nvda['Close']
nvda['Move Size'] = nvda['Change'] / nvda['Exp Change']
nvda.tail()
# ### Filtering or subsetting
nvda['20181225':]
nvda[nvda['Move Size']> 2]
len(nvda[nvda['Move Size']> 2])
len(nvda)
big_move = nvda[(nvda['Move Size']> 2) | (nvda['Move Size']< -2) ]
len(big_move)
big_move['Move Size'].plot(kind='hist',bins=25)
nvda[['Close', 'Change', 'Exp Change', 'Return']].head()
nvda['Volume'].describe()
nvda[nvda.Volume > nvda.Volume.mean() + 2 * nvda.Volume.std()]
| Section 2/2.6-Filtering_with_pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os, json, random, cv2
import numpy as np, pandas as pd
import matplotlib.pyplot as plt
import tensorflow as tf, re, math
from tqdm import tqdm
from PIL import Image
train = pd.read_csv('../input/landmark-recognition-2021/train.csv')
train
# +
import glob
train_file_list=glob.glob('../input/landmark-recognition-2021/train')
train_file_list
my_train_list=[]
path='../input/landmark-recognition-2021/train'
for file in glob.glob(path):
print(file)
a=cv2.imread(file)
my_train_list.append(a)
my_train_list
# -
def image_grid5x5(image_array, landmarks):
fig = plt.figure(figsize=(15., 15.))
grid = ImageGrid(fig, 111,
nrows_ncols=(5, 5),
axes_pad=1)
for idx, (ax, im) in enumerate(zip(grid, image_array)):
ax.imshow(im)
ax.set_title(landmarks[idx])
ax.set_xlabel(f'{im.shape}')
plt.show()
def make_img_path(img_id):
return "/".join([char for char in img_id[:3]]) + "/" + img_id + ".jpg"
def get_img_numpy(img_id, base="/train"):
img_path = make_img_path(img_id)
img = Image.open(base + "/" + img_path)
return np.asarray(img)
# +
from mpl_toolkits.axes_grid1 import ImageGrid
from PIL import Image
import seaborn as sns
BASE_PATH = "../input/landmark-recognition-2021"
img_array_train = [get_img_numpy(img, BASE_PATH + "/train") for img in train['id'][500:525]]
image_grid5x5(img_array_train, [landmark for landmark in train['landmark_id'][500:525]])
# -
sns.histplot(data=train, x="landmark_id", bins=100)
sns.set()
plt.title('Training set: number of images per class(line plot)')
landmarks_fold = pd.DataFrame(train['landmark_id'].value_counts())
landmarks_fold.reset_index(inplace=True)
landmarks_fold.columns = ['landmark_id','count']
ax = landmarks_fold['count'].plot(logy=True, grid=True)
locs, labels = plt.xticks()
plt.setp(labels, rotation=30)
ax.set(xlabel="Landmarks", ylabel="Number of images")
landmarks_fold.head()
# **here i am tring to get only the landmark image which has landmark_id 20883 **
train = train[train.landmark_id==20883]
train
#
def get_train_file_path(image_id):
return "../input/landmark-recognition-2021/train/{}/{}/{}/{}.jpg".format(
image_id[0], image_id[1], image_id[2], image_id)
train['file_path'] = train['id'].apply(get_train_file_path)
train.head()
# there are two image in train data set which belong to landmark_id , i get the file path and i am going to plot using the file path
fig = plt.figure(figsize=(30,20))
x=1
for i in train.file_path[:20883]:
image = cv2.imread(i)
fig.add_subplot(4, 4, x)
plt.imshow(cv2.cvtColor(image, cv2.COLOR_BGR2RGB))
plt.axis('off')
x+=1
# \I am now going to extract featears from the image
| land-mark-recognition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # BaseTransformer
# This notebook shows the functionality included in the BaseTransformer class. This is the base class for the package and all other transformers within should inherit from it. This means that the functionality below is also present in the other transformers in the package. <br>
# This is more 'behind the scenes' functionality that is useful to be aware of, but not the actual transformations required before building / predicting with models. <br>
# Examples of the actual pre processing transformations can be found in the other notebooks in this folder.
import pandas as pd
import numpy as np
from sklearn.datasets import fetch_california_housing
import tubular
from tubular.base import BaseTransformer
tubular.__version__
# ## Load California housing dataset from sklearn
cali = fetch_california_housing()
cali_df = pd.DataFrame(cali['data'], columns=cali['feature_names'])
cali_df.head()
cali_df.dtypes
# ## Initialising BaseTransformer
# ### Not setting columns
# Columns do not have to be specified when initialising BaseTransformer objects. Both the fit and transform methods call the columns_set_or_check to ensure that columns is set before the transformer has to do any work.
base_1 = BaseTransformer(copy = True, verbose = True)
# ## BaseTransformer fit
# Not all transformers in the package will implement a fit method, if the user directly specifies the values the transformer needs e.g. passes the impute value, there is no need for it.
# ### Setting columns in fit
# If the columns attribute is not set when fit is called, columns_set_or_check will set columns to be all columns in X.
base_1.columns is None
base_1.fit(cali_df)
base_1.columns
# ## BaseTransformer transform
# All transformers will implement a transform method.
# ### Transform with copy
# This ensures that the input dataset is not modified in transform.
cali_df_2 = base_1.transform(cali_df)
pd.testing.assert_frame_equal(cali_df_2, cali_df)
cali_df_2 is cali_df
# ### Transform without copy
# This can be useful if you are working with a large dataset or are concerned about the time to copy.
base_2 = BaseTransformer(copy = False, verbose = True)
cali_df_3 = base_2.fit_transform(cali_df)
cali_df_3 is cali_df
| examples/base/BaseTransformer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # License
# ***
# Copyright (C) 2017 <NAME>, <EMAIL>
#
# Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
# imports
import h2o
import numpy as np
import pandas as pd
from h2o.estimators.deeplearning import H2ODeepLearningEstimator
from h2o.grid.grid_search import H2OGridSearch
# display matplotlib graphics in notebook
# %matplotlib inline
# start and connect to h2o server
h2o.init()
# load clean data
path = '/Users/phall/workspace/GWU_data_mining/03_regression/data/loan_clean.csv'
# define input variable measurement levels
# strings automatically parsed as enums (nominal)
# numbers automatically parsed as numeric
col_types = {'bad_loan': 'enum'}
frame = h2o.import_file(path=path, col_types=col_types) # multi-threaded import
frame.describe()
# split into 40% training, 30% validation, and 30% test
train, valid, test = frame.split_frame([0.4, 0.3])
# assign target and inputs
y = 'bad_loan'
X = [name for name in frame.columns if name not in ['id', '_WARN_', y]]
print(y)
print(X)
# set target to factor - for binary classification
train[y] = train[y].asfactor()
valid[y] = valid[y].asfactor()
test[y] = test[y].asfactor()
# +
# neural network
# initialize nn model
nn_model = H2ODeepLearningEstimator(
epochs=50, # read over the data 50 times, but in mini-batches
hidden=[100], # 100 hidden units in 1 hidden layer
input_dropout_ratio=0.2, # randomly drop 20% of inputs for each iteration, helps w/ generalization
hidden_dropout_ratios=[0.05], # randomly set 5% of hidden weights to 0 each iteration, helps w/ generalization
activation='TanhWithDropout', # bounded activation function that allows for dropout, tanh
l1=0.001, # L1 penalty can help generalization
l2=0.01, # L2 penalty can increase stability in presence of highly correlated inputs
adaptive_rate=True, # adjust magnitude of weight updates automatically (+stability, +accuracy)
stopping_rounds=5, # stop after validation error does not decrease for 5 iterations
score_each_iteration=True, # score validation error on every iteration
model_id='nn_model') # for easy lookup in flow
# train nn model
nn_model.train(
x=X,
y=y,
training_frame=train,
validation_frame=valid)
# print model information
nn_model
# view detailed results at http://localhost:54321/flow/index.html
# -
# measure nn AUC
print(nn_model.auc(train=True))
print(nn_model.auc(valid=True))
print(nn_model.model_performance(test_data=test).auc())
# +
# NN with random hyperparameter search
# train many different NN models with random hyperparameters
# and select best model based on validation error
# define random grid search parameters
hyper_parameters = {'hidden':[[170, 320], [80, 190], [320, 160, 80], [100], [50, 50, 50, 50]],
'l1':[s/1e4 for s in range(0, 1000, 100)],
'l2':[s/1e5 for s in range(0, 1000, 100)],
'input_dropout_ratio':[s/1e2 for s in range(0, 20, 2)]}
# define search strategy
search_criteria = {'strategy':'RandomDiscrete',
'max_models':20,
'max_runtime_secs':600}
# initialize grid search
gsearch = H2OGridSearch(H2ODeepLearningEstimator,
hyper_params=hyper_parameters,
search_criteria=search_criteria)
# execute training w/ grid search
gsearch.train(x=X,
y=y,
training_frame=train,
validation_frame=valid)
# view detailed results at http://localhost:54321/flow/index.html
# +
# show grid search results
gsearch.show()
# select best model
nn_model2 = gsearch.get_grid()[0]
# print model information
nn_model2
# -
# measure nn AUC
print(nn_model2.auc(train=True))
print(nn_model2.auc(valid=True))
print(nn_model2.model_performance(test_data=test).auc())
# partial dependence plots are a powerful machine learning interpretation tool
# to calculate partial dependence across the domain a variable
# hold column of interest at constant value
# find the mean prediction of the model with this column constant
# repeat for multiple values of the variable of interest
# h2o has a built-in function for partial dependence as well
par_dep_dti1 = nn_model2.partial_plot(data=train, cols=['STD_IMP_REP_dti'], server=True, plot=True)
# shutdown h2o
h2o.cluster().shutdown(prompt=False)
| 05_neural_networks/src/py_part_5_neural_networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import os
import cv2
import random
from tqdm import tqdm
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation, Flatten, Conv2D, MaxPooling2D, BatchNormalization, Dropout
from sklearn.model_selection import train_test_split
from tensorflow.keras.preprocessing.image import ImageDataGenerator
# -
DATADIR = "C:/Users/Rawan/Desktop/Classification-Flutter-TFlite-master/Currency/"
CATEGORIE = ["5", "10", "20", "50", "100", "200"]
CLASSES = {
'1': 0,
'5': 1,
'10': 2,
'20': 3,
'50': 4,
'100': 5,
'200': 6
}
# +
dataset = []
for cat in os.listdir(DATADIR):
path = os.path.join(DATADIR, cat)
for img_path in os.listdir(path):
img = cv2.imread(os.path.join(path, img_path))
img = cv2.resize(img, (128, 128))
img = img / 255.0
lab = CLASSES[cat]
dataset.append([img, lab])
random.seed(42)
random.shuffle(dataset)
data = []
labels = []
for d, l in dataset:
data.append(d)
labels.append(l)
data = np.array(data)
labels = np.array(labels)
print("Data shape:", data.shape)
print("Labels shape:", labels.shape)
# +
(trainX, testX, trainY, testY) = train_test_split(data, labels, test_size=0.2, random_state=42, stratify=labels)
print(trainX.shape)
print(trainY.shape)
print(testX.shape)
print(testY.shape)
# +
import seaborn as sns
sns.countplot(x=labels);
# -
sns.countplot(x=trainY);
sns.countplot(x=testY);
# +
def create_model(input_shape):
model= Sequential()
model.add(Conv2D(64,(3,3), padding='same', input_shape = input_shape))
#model.add(BatchNormalization())
model.add(Activation("relu"))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Conv2D(128, (3,3), padding='same'))
#model.add(BatchNormalization())
model.add(Activation("relu"))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Conv2D(256, (3,3), padding='same'))
#model.add(BatchNormalization())
model.add(Activation("relu"))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Flatten())
model.add(Dense(64))
model.add(Dropout(0.5))
model.add(Dense(len(CLASSES)))
model.add(Activation('softmax'))
return model
def create_mobilenet(input_shape):
base_model = tf.keras.applications.MobileNetV2(include_top=False, input_shape=input_shape)
base_model.trainable = False
model = tf.keras.Sequential([
base_model,
tf.keras.layers.GlobalAveragePooling2D(),
tf.keras.layers.Dense(len(CLASSES), activation='softmax')
])
return model
#model = create_model(data.shape[1:])
model = create_mobilenet(data.shape[1:])
model.compile(loss='sparse_categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
aug = ImageDataGenerator(rotation_range=25, width_shift_range=0.1,
height_shift_range=0.1, shear_range=0.2, zoom_range=0.2,
horizontal_flip=True, fill_mode="nearest")
model.summary()
# +
EPOCHS = 80
H = model.fit(aug.flow(trainX, trainY, batch_size=32),
validation_data=(testX, testY),
steps_per_epoch=len(trainX) // 32,
epochs=EPOCHS, verbose=1)
#H = model.fit(trainX, trainY, validation_data=(testX, testY), epochs=20, verbose=1)
# -
plt.style.use("ggplot")
plt.figure()
N = EPOCHS
plt.plot(np.arange(0, N), H.history["loss"], label="train_loss")
plt.plot(np.arange(0, N), H.history["val_loss"], label="val_loss")
plt.plot(np.arange(0, N), H.history["accuracy"], label="train_acc")
plt.plot(np.arange(0, N), H.history["val_accuracy"], label="val_acc")
plt.title("Training Loss and Accuracy")
plt.xlabel("Epoch #")
plt.ylabel("Loss/Accuracy")
plt.legend(loc="upper left");
# +
from sklearn.metrics import classification_report
preds = model.predict(testX)
predIdxs = np.argmax(preds, axis=1)
print(classification_report(testY, predIdxs, target_names=CLASSES.keys()))
# +
# Convert the model.
converter = tf.lite.TFLiteConverter.from_keras_model(model)
tflite_model = converter.convert()
# Save the model.
with open('C:/Users/Rawan/Desktop/Classification-Flutter-TFlite-master/assets/modelMobNet.tflite', 'wb') as f:
f.write(tflite_model)
# -
| models_for_currency/currency_detection (mobilenet).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: zenv
# language: python
# name: zenv
# ---
# Checking to see how much faster things might be if we have the data stored locally, vs doing all the data processing/loading on the each request for a new batch
import torch
import numpy as np
import pandas as pd
import os
import h5py
from exabiome.nn.loader import read_dataset, LazySeqDataset
import argparse
from torch.utils.data import Dataset, DataLoader
import torch.nn as nn
from model import *
from data import *
from fastai.text.all import *
x_df = pd.read_csv('data/toy_x.csv')
x_df.shape
y_df = pd.read_csv('data/toy_y.csv')
y_df.shape
x_df.head()
torch.Tensor(x_df.iloc[0][1:]).shape
torch.Tensor(y_df.iloc[0][1:])
len(x_df)
class local_ds(Dataset):
def __init__(self, xs_df, ys_df):
self.xs = xs_df
self.ys = ys_df
def __len__(self):
return len(self.xs)
def __getitem__(self, idx):
x = torch.Tensor(self.xs.iloc[idx][1:])
y = torch.Tensor(self.ys.iloc[idx][1:])
x = x.to(torch.float)
y = y.to(torch.long)
return (x.unsqueeze(0), y.squeeze())
train_ds = local_ds(x_df[:len(x_df)-2000], y_df[:len(x_df)-2000])
val_ds = local_ds(x_df[-2000:], y_df[-2000:])
len(train_ds), len(val_ds)
train_ds[0]
train_ds[0][1].shape
train_dl = DataLoader(train_ds, batch_size=128, shuffle=True)
valid_dl = DataLoader(val_ds, batch_size=128, shuffle=False)
len(train_dl), len(valid_dl)
dls = DataLoaders(train_dl, valid_dl)
class compress_cb(Callback):
def after_pred(self):
self.learn.pred = self.pred.squeeze(2)
model = EffNet_b0(out_feats=1)
model.to('cuda')
dls.to('cuda')
batch = next(iter(dls.train))
batch[0].shape, batch[1].shape
out = model(batch[0])
out.shape
nn.CrossEntropyLoss()(out.squeeze(2), batch[1])
learn = Learner(dls, model, loss_func=nn.CrossEntropyLoss(),
cbs=[compress_cb],
metrics=[accuracy]).to_fp16()
learn.lr_find()
learn.fine_tune(2)
# This looks 3 times faster when pulling the data from a csv -- probably the way to go for prototyping. Should look into how fast it is with the data in the /tmp folder as per this: https://docs-dev.nersc.gov/cgpu/usage/
| local_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: imbalanced
# language: python
# name: imbalanced
# ---
# # ADASYN
#
# Creates new samples by interpolation of samples of the minority class and its closest neighbours. It creates more samples from samples that are harder to classify.
# +
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.datasets import make_blobs
from imblearn.over_sampling import ADASYN
# -
# ## Create data
#
# https://scikit-learn.org/stable/modules/generated/sklearn.datasets.make_blobs.html
#
# We will create 2 classes, one majority one minority, clearly separated to facilitate the demonstration.
# +
# Configuration options
blobs_random_seed = 42
centers = [(0, 0), (5, 5)]
cluster_std = 1.5
num_features_for_samples = 2
num_samples_total = 1600
# Generate X
X, y = make_blobs(
n_samples=num_samples_total,
centers=centers,
n_features=num_features_for_samples,
cluster_std=cluster_std)
# transform arrays to pandas formats
X = pd.DataFrame(X, columns=['VarA', 'VarB'])
y = pd.Series(y)
# create an imbalancced Xset
# (make blobs creates same number of obs per class
# we need to downsample manually)
X = pd.concat([
X[y == 0],
X[y == 1].sample(200, random_state=42)
], axis=0)
y = y.loc[X.index]
# display size
X.shape, y.shape
# +
sns.scatterplot(
data=X, x="VarA", y="VarB", hue=y, alpha=0.5
)
plt.title('Toy dataset')
plt.show()
# -
# ## ADASYN
#
# https://imbalanced-learn.org/stable/generated/imblearn.over_sampling.ADASYN.html
# +
ada = ADASYN(
sampling_strategy='auto', # samples only the minority class
random_state=0, # for reproducibility
n_neighbors=5,
n_jobs=4
)
X_res, y_res = ada.fit_resample(X, y)
# +
# size of original data
X.shape, y.shape
# +
# size of undersampled data
X_res.shape, y_res.shape
# +
# number of minority class observations
y.value_counts(), y_res.value_counts()
# +
# plot of original data
sns.scatterplot(
data=X, x="VarA", y="VarB", hue=y,alpha=0.5
)
plt.title('Original dataset')
plt.show()
# +
# plot of original data
sns.scatterplot(
data=X_res, x="VarA", y="VarB", hue=y_res, alpha=0.5
)
plt.title('Over-sampled dataset')
plt.show()
# -
# There are now new observations that differ from the original ones. New observations sit at the boundary between the classes.
# **HOMEWORK**
#
# - Test ADASYN using the toy datasets that we created for section 4 and see how the distribution of the newly created data varies with the different separateness of the classes.
| Section-05-Oversampling/05-04-ADASYN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: DESI master
# language: python
# name: desi-master
# ---
# +
#import sys
# #!{sys.executable} -m pip install --user alerce
# -
# # light_transient_matching
# ## Matches DESI observations to ALERCE and DECAM ledger objects
#
# This code predominately takes in data from the ALERCE and DECAM ledger brokers and identifies DESI observations within 2 arcseconds of those objects, suspected to be transients. It then prepares those matches to be fed into our [CNN code](https://github.com/MatthewPortman/timedomain/blob/master/cronjobs/transient_matching/modified_cnn_classify_data_gradCAM.ipynb) which attempts to identify the class of these transients.
#
# The main matching algorithm uses astropy's **match_coordinate_sky** to match 1-to-1 targets with the objects from the two ledgers. Wrapping functions handle data retrieval from both the ledgers as well as from DESI and prepare this data to be fed into **match_coordinate_sky**. Since ALERCE returns a small enough (pandas) dataframe, we do not need to precondition the input much. However, DECAM has many more objects to match so we use a two-stage process: an initial 2 degree match to tile RA's/DEC's and a second closer 1 arcsecond match to individual targets.
#
# As the code is a work in progress, please forgive any redundancies. We are attempting to merge all of the above (neatly) into the same two or three matching/handling functions!
# +
from astropy.io import fits
from astropy.table import Table
from astropy import units as u
from astropy.time import Time
from astropy.coordinates import SkyCoord, match_coordinates_sky, Angle
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from glob import glob
import sys
import sqlite3
import os
from desispec.io import read_spectra, write_spectra
from desispec.spectra import Spectra
# Some handy global variables
global db_filename
db_filename = '/global/cfs/cdirs/desi/science/td/daily-search/transients_search.db'
global exposure_path
exposure_path = os.environ["DESI_SPECTRO_REDUX"]
global color_band
color_band = "r"
global minDist
minDist = {}
global today
today = Time.now()
# -
# ## Necessary functions
# +
# Grabbing the file names
def all_candidate_filenames(transient_dir: str):
# This function grabs the names of all input files in the transient directory and does some python string manipulation
# to grab the names of the input files with full path and the filenames themselves.
try:
filenames_read = glob(transient_dir + "/*.fits") # Hardcoding is hopefully a temporary measure.
except:
print("Could not grab/find any fits in the transient spectra directory:")
print(transient_dir)
filenames_read = [] # Just in case
#filenames_out = [] # Just in case
raise SystemExit("Exiting.")
#else:
#filenames_out = [s.split(".")[0] for s in filenames_read]
#filenames_out = [s.split("/")[-1] for s in filenames_read]
#filenames_out = [s.replace("in", "out") for s in filenames_out]
return filenames_read #, filenames_out
#path_to_transient = "/global/cfs/cdirs/desi/science/td/daily-search/desitrip/out"
#print(all_candidate_filenames(path_to_transient)[1])
# +
# From ALeRCE_ledgermaker https://github.com/alercebroker/alerce_client
# I have had trouble importing this before so I copy, paste it, and modify it here.
# I also leave these imports here because why not?
import requests
from alerce.core import Alerce
from alerce.exceptions import APIError
alerce_client = Alerce()
# Choose cone_radius of diameter of tile so that, whatever coord I choose for ra_in, dec_in, we cover the whole tile
def access_alerts(lastmjd_in=[], ra_in = None, dec_in = None, cone_radius = 3600*4.01, classifier='stamp_classifier', class_names=['SN', 'AGN']):
if type(class_names) is not list:
raise TypeError('Argument `class_names` must be a list.')
dataframes = []
if not lastmjd_in:
date_range = 60
lastmjd_in = [Time.now().mjd - 60, Time.now().mjd]
print('Defaulting to a lastmjd range of', str(date_range), 'days before today.')
#print("lastmjd:", lastmjd_in)
for class_name in class_names:
data = alerce_client.query_objects(classifier=classifier,
class_name=class_name,
lastmjd=lastmjd_in,
ra = ra_in,
dec = dec_in,
radius = cone_radius, # in arcseconds
page_size = 5000,
order_by='oid',
order_mode='DESC',
format='pandas')
#if lastmjd is not None:
# select = data['lastmjd'] >= lastmjd
# data = data[select]
dataframes.append(data)
#print(pd.concat(dataframes).columns)
return pd.concat(dataframes).sort_values(by = 'lastmjd')
# -
# From https://github.com/desihub/timedomain/blob/master/too_ledgers/decam_TAMU_ledgermaker.ipynb
# Function to grab decam data
from bs4 import BeautifulSoup
import json
import requests
def access_decam_data(url, overwrite=False):
"""Download reduced DECam transient data from Texas A&M.
Cache the data to avoid lengthy and expensive downloads.
Parameters
----------
url : str
URL for accessing the data.
overwrite : bool
Download new data and overwrite the cached data.
Returns
-------
decam_transients : pandas.DataFrame
Table of transient data.
"""
folders = url.split('/')
thedate = folders[-1] if len(folders[-1]) > 0 else folders[-2]
outfile = '{}.csv'.format(thedate)
if os.path.exists(outfile) and not overwrite:
# Access cached data.
decam_transients = pd.read_csv(outfile)
else:
# Download the DECam data index.
# A try/except is needed because the datahub SSL certificate isn't playing well with URL requests.
try:
decam_dets = requests.get(url, auth=('decam','tamudecam')).text
except:
requests.packages.urllib3.disable_warnings(requests.packages.urllib3.exceptions.InsecureRequestWarning)
decam_dets = requests.get(url, verify=False, auth=('decam','tamudecam')).text
# Convert transient index page into scrapable data using BeautifulSoup.
soup = BeautifulSoup(decam_dets)
# Loop through transient object summary JSON files indexed in the main transient page.
# Download the JSONs and dump the info into a Pandas table.
decam_transients = None
j = 0
for a in soup.find_all('a', href=True):
if 'object-summary.json' in a:
link = a['href'].replace('./', '')
summary_url = url + link
summary_text = requests.get(summary_url, verify=False, auth=('decam','tamudecam')).text
summary_data = json.loads(summary_text)
j += 1
#print('Accessing {:3d} {}'.format(j, summary_url)) # Modified by Matt
if decam_transients is None:
decam_transients = pd.DataFrame(summary_data, index=[0])
else:
decam_transients = pd.concat([decam_transients, pd.DataFrame(summary_data, index=[0])])
# Cache the data for future access.
print('Saving output to {}'.format(outfile))
decam_transients.to_csv(outfile, index=False)
return decam_transients
# + tags=[]
# Function to read in fits table info, RA, DEC, MJD and targetid if so desired
# Uses control parameter tile to determine if opening tile exposure file or not since headers are different
import logging
def read_fits_info(filepath: str, transient_candidate = True):
'''
if transient_candidate:
hdu_num = 1
else:
hdu_num = 5
'''
# Disabling INFO logging temporarily to suppress INFO level output/print from read_spectra
logging.disable(logging.INFO)
try:
spec_info = read_spectra(filepath).fibermap
except:
filename = filepath.split("/")[-1]
print("Could not open or use:", filename)
#print("In path:", filepath)
#print("Trying the next file...")
return np.array([]), np.array([]), 0, 0
headers = ['TARGETID', 'TARGET_RA', 'TARGET_DEC', 'LAST_MJD']
targ_info = {}
for head in headers:
try:
targ_info[head] = spec_info[head].data
except:
if not head == 'LAST_MJD': print("Failed to read in", head, "data. Continuing...")
targ_info[head] = False
# targ_id = spec_info['TARGETID'].data
# targ_ra = spec_info['TARGET_RA'].data # Now it's a numpy array
# targ_dec = spec_info['TARGET_DEC'].data
# targ_mjd = spec_info['LAST_MJD'] #.data
if np.any(targ_info['LAST_MJD']):
targ_mjd = Time(targ_info['LAST_MJD'][0], format = 'mjd')
elif transient_candidate:
targ_mjd = filepath.split("/")[-1].split("_")[-2] #to grab the date
targ_mjd = Time(targ_mjd, format = 'mjd') #.mjd
else:
print("Unable to determine observation mjd for", filename)
print("This target will not be considered.")
return np.array([]), np.array([]), 0, 0
'''
with fits.open(filepath) as hdu1:
data_table = Table(hdu1[hdu_num].data) #columns
targ_id = data_table['TARGETID']
targ_ra = data_table['TARGET_RA'].data # Now it's a numpy array
targ_dec = data_table['TARGET_DEC'].data
#targ_mjd = data_table['MJD'][0] some have different versions of this so this is a *bad* idea... at least now I know the try except works!
if tile:
targ_mjd = hdu1[hdu_num].header['MJD-OBS']
'''
# if tile and not np.all(targ_mjd):
# print("Unable to grab mjd from spectra, taking it from the filename...")
# targ_mjd = filepath.split("/")[-1].split("_")[-2] #to grab the date
# #targ_mjd = targ_mjd[:4]+"-"+targ_mjd[4:6]+"-"+targ_mjd[6:] # Adding dashes for Time
# targ_mjd = Time(targ_mjd, format = 'mjd') #.mjd
# Re-enabling logging for future calls if necessary
logging.disable(logging.NOTSET)
return targ_info["TARGET_RA"], targ_info["TARGET_DEC"], targ_mjd, targ_info["TARGETID"] #targ_ra, targ_dec, targ_mjd, targ_id
# -
# ## Matching function
#
# More or less the prototype to the later rendition used for DECAM. Will not be around in later versions of this notebook as I will be able to repurpose the DECAM code to do both. Planned obsolescence?
#
# It may not be even worth it at this point... ah well!
# + tags=[]
# Prototype for the later, heftier matching function
# Will be deprecated, please reference commentary in inner_matching later for operation notes
def matching(path_in: str, max_sep: float, tile = False, date_dict = {}):
max_sep *= u.arcsec
#max_sep = Angle(max_sep*u.arcsec)
#if not target_ra_dec_date:
# target_ras, target_decs, obs_mjds = read_fits_ra_dec(path_in, tile)
#else:
# target_ras, target_decs, obs_mjds = target_ra_dec_date
#Look back 60 days from the DESI observations
days_back = 60
if not date_dict:
print("No RA's/DEC's fed in. Quitting.")
return np.array([]), np.array([])
all_trans_matches = []
all_alerts_matches = []
targetid_matches = []
for obs_mjd, ra_dec in date_dict.items():
# Grab RAs and DECs from input.
target_ras = ra_dec[:, 0]
target_decs = ra_dec[:, 1]
target_ids = np.int64(ra_dec[:, 2])
# Check for NaN's and remove which don't play nice with match_coordinates_sky
nan_ra = np.isnan(target_ras)
nan_dec = np.isnan(target_decs)
if np.any(nan_ra) or np.any(nan_dec):
print("NaNs found, removing them from array (not FITS) before match.")
#print("Original length (ra, dec): ", len(target_ras), len(target_decs))
nans = np.logical_not(np.logical_and(nan_ra, nan_dec))
target_ras = target_ras[nans] # Logic masking, probably more efficient
target_decs = target_decs[nans]
#print("Reduced length (ra, dec):", len(target_ras), len(target_decs))
# Some code used to test -- please ignore ******************
# Feed average to access alerts, perhaps that will speed things up/find better results
#avg_ra = np.average(target_ras)
#avg_dec = np.average(target_decs)
# coo_trans_search = SkyCoord(target_ras*u.deg, target_decs*u.deg)
# #print(coo_trans_search)
# idxs, d2d, _ = match_coordinates_sky(coo_trans_search, coo_trans_search, nthneighbor = 2)
# # for conesearch in alerce
# max_sep = np.max(d2d).arcsec + 2.1 # to expand a bit further than the furthest neighbor
# ra_in = coo_trans_search[0].ra
# dec_in = coo_trans_search[0].dec
# Some code used to test -- please ignore ******************
#print([obs_mjd - days_back, obs_mjd])
try:
alerts = access_alerts(lastmjd_in = [obs_mjd - days_back, obs_mjd],
ra_in = target_ras[0],
dec_in = target_decs[0], #cone_radius = max_sep,
class_names = ['SN']
) # Modified Julian Day .mjd
except:
#print("No SN matches ("+str(days_back)+" day range) for", obs_mjd)
#break
continue
# For each fits file, look at one month before the observation from Alerce
# Not sure kdtrees matter
# tree_name = "kdtree_" + str(obs_mjd - days_back)
alerts_ra = alerts['meanra'].to_numpy()
#print("Length of alerts: ", len(alerts_ra))
alerts_dec = alerts['meandec'].to_numpy()
# Converting to SkyCoord type arrays (really quite handy)
coo_trans_search = SkyCoord(target_ras*u.deg, target_decs*u.deg)
coo_alerts = SkyCoord(alerts_ra*u.deg, alerts_dec*u.deg)
# Some code used to test -- please ignore ******************
#ra_range = list(zip(*[(i, j) for i,j in zip(alerts_ra,alerts_dec) if (np.min(target_ras) < i and i < np.max(target_ras) and np.min(target_decs) < j and j < np.max(target_decs))]))
#try:
# ra_range = SkyCoord(ra_range[0]*u.deg, ra_range[1]*u.deg)
#except:
# continue
#print(ra_range)
#print(coo_trans_search)
#idx_alerts, d2d_trans, d3d_trans = match_coordinates_sky(coo_trans_search, ra_range)
#for i in coo_trans_search:
#print(i.separation(ra_range[3]))
#print(idx_alerts)
#print(np.min(d2d_trans))
#break
# Some code used to test -- please ignore ******************
idx_alerts, d2d_trans, d3d_trans = match_coordinates_sky(coo_trans_search, coo_alerts)
# Filtering by maximum separation and closest match
sep_constraint = d2d_trans < max_sep
trans_matches = coo_trans_search[sep_constraint]
alerts_matches = coo_alerts[idx_alerts[sep_constraint]]
targetid_matches = target_ids[sep_constraint]
#print(d2d_trans < max_sep)
minDist[obs_mjd] = np.min(d2d_trans)
# Adding everything to lists and outputting
if trans_matches.size:
all_trans_matches.append(trans_matches)
all_alerts_matches.append(alerts_matches)
sort_dist = np.sort(d2d_trans)
#print("Minimum distance found: ", sort_dist[0])
#print()
#break
#else:
#print("No matches found...\n")
#break
return all_trans_matches, all_alerts_matches, targetid_matches
# -
# ## Matching to ALERCE
# Runs a 5 arcsecond match of DESI to Alerce objects. Since everything is handled in functions, this part is quite clean.
#
# From back when I was going to use *if __name__ == "__main__":*... those were the days
# +
# Transient dir
path_to_transient = "/global/cfs/cdirs/desi/science/td/daily-search/desitrip/out"
# Grab paths
paths_to_fits = all_candidate_filenames(path_to_transient)
#print(len(paths_to_fits))
desi_info_dict = {}
target_ras, target_decs, obs_mjd, targ_ids = read_fits_info(paths_to_fits[0], transient_candidate = True)
desi_info_dict[obs_mjd] = np.column_stack((target_ras, target_decs, targ_ids))
'''
To be used when functions are properly combined.
initial_check(ledger_df = None, ledger_type = '')
closer_check(matches_dict = {}, ledger_df = None, ledger_type = '', exclusion_list = [])
'''
fail_count = 0
# Iterate through every fits file and grab all necessary info and plop it all together
for path in paths_to_fits[1:]:
target_ras, target_decs, obs_mjd, targ_ids = read_fits_info(path, transient_candidate = True)
if not obs_mjd:
fail_count += 1
continue
#try:
if obs_mjd in desi_info_dict.keys():
np.append(desi_info_dict[obs_mjd], np.array([target_ras, target_decs, targ_ids]).T, axis = 0)
else:
desi_info_dict[obs_mjd] = np.column_stack((target_ras, target_decs, targ_ids))
#desi_info_dict[obs_mjd].extend((target_ras, target_decs, targ_ids))
#except:
# continue
#desi_info_dict[obs_mjd] = np.column_stack((target_ras, target_decs, targ_ids))
#desi_info_dict[obs_mjd].append((target_ras, target_decs, targ_ids))
#trans_matches, _ = matching(path, 5.0, (all_desi_ras, all_desi_decs, all_obs_mjd))
# if trans_matches.size:
# all_trans_matches.append(trans_matches)
# all_alerts_matches.append(alerts_matches)
# -
#print([i.mjd for i in sorted(desi_info_dict.keys())])
print(len(paths_to_fits))
print(len(desi_info_dict))
#print(fail_count)
# + active=""
# #print(len(desi_info_dict))
# temp_dict = {a:b for a,b,c in zip(desi_info_dict.keys(), desi_info_dict.values(), range(len(desi_info_dict))) if c > 650}
# +
# I was going to prepare everything by removing duplicate target ids but it's more trouble than it's worth and match_coordinates_sky can handle it
# Takes quite a bit of time... not much more I can do to speed things up though since querying Alerce for every individual date is the hang-up.
#print(len(paths_to_fits) - ledesi_info_dictfo_dict))
#print(fail_count)
#trans_matches, _, target_id_matches = matching("", 2.0, date_dict = temp_dict)
trans_matches, _, target_id_matches = matching("", 2.0, date_dict = desi_info_dict)
print(trans_matches)
print(target_id_matches)
# -
print(sorted(minDist.values())[:5])
#for i in minDist.values():
# print(i)
# ## Matching to DECAM functions
# Overwrite *read_fits_info* with older version to accommodate *read_spectra* error
# Read useful data from fits file, RA, DEC, target ID, and mjd as a leftover from previous use
def read_fits_info(filepath: str, transient_candidate = False):
if transient_candidate:
hdu_num = 1
else:
hdu_num = 5
try:
with fits.open(filepath) as hdu1:
data_table = Table(hdu1[hdu_num].data) #columns
targ_ID = data_table['TARGETID']
targ_ra = data_table['TARGET_RA'].data # Now it's a numpy array
targ_dec = data_table['TARGET_DEC'].data
#targ_mjd = data_table['MJD'][0] some have different versions of this so this is a *bad* idea... at least now I know the try except works!
# if transient_candidate:
# targ_mjd = hdu1[hdu_num].header['MJD-OBS'] # This is a string
# else:
# targ_mjd = data_table['MJD'].data
# targ_mjd = Time(targ_mjd[0], format = 'mjd')
except:
filename = filepath.split("/")[-1]
print("Could not open or use:", filename)
#print("In path:", filepath)
#print("Trying the next file...")
return np.array([]), np.array([]), np.array([])
return targ_ra, targ_dec, targ_ID #targ_mjd, targ_ID
# +
# Grabbing the frame fits files
def glob_frames(exp_d: str):
# This function grabs the names of all input files in the transient directory and does some python string manipulation
# to grab the names of the input files with full path and the filenames themselves.
try:
filenames_read = glob(exp_d + "/cframe-" + color_band + "*.fits") # Only need one of b, r, z
# sframes not flux calibrated
# May want to use tiles... coadd (will need later, but not now)
except:
try:
filenames_read = glob(exp_d + "/frame-" + color_band + "*.fits") # Only need one of b, r, z
except:
print("Could not grab/find any fits in the exposure directory:")
print(exp_d)
filenames_read = [] # Just in case
#filenames_out = [] # Just in case
raise SystemExit("Exitting.")
#else:
#filenames_out = [s.split(".")[0] for s in filenames_read]
#filenames_out = [s.split("/")[-1] for s in filenames_read]
#filenames_out = [s.replace("in", "out") for s in filenames_out]
return filenames_read #, filenames_out
#path_to_transient = "/global/cfs/cdirs/desi/science/td/daily-search/desitrip/out"
#print(all_candidate_filenames(path_to_transient)[1])
# -
# ## Match handling routines
#
# The two functions below perform data handling/calling for the final match step.
#
# The first, **initial_check** grabs all the tile RAs and DECS from the exposures and tiles SQL table, does some filtering, and sends the necessary information to the matching function. Currently designed to handle ALERCE as well but work has to be done to make sure it operates correctly.
def initial_check(ledger_df = None, ledger_type = ''):
query_date_start = "20210301"
#today = Time.now()
smushed_YMD = today.iso.split(" ")[0].replace("-","")
query_date_end = smushed_YMD
# Handy queries for debugging/useful info
query2 = "PRAGMA table_info(exposures)"
query3 = "PRAGMA table_info(tiles)"
# Crossmatch across tiles and exposures to grab obsdate via tileid
query_match = "SELECT distinct tilera, tiledec, obsdate, obsmjd, expid, exposures.tileid from exposures INNER JOIN tiles ON exposures.tileid = tiles.tileid where obsdate BETWEEN " + \
query_date_start + " AND " + query_date_end + ";"
'''
Some handy code for debugging
#cur.execute(query2)
#row2 = cur.fetchall()
#for i in row2:
# print(i[:])
'''
# Querying sql and returning a data type called sqlite3 row, it's kind of like a namedtuple/dictionary
conn = sqlite3.connect(db_filename)
conn.row_factory = sqlite3.Row # https://docs.python.org/3/library/sqlite3.html#sqlite3.Row
cur = conn.cursor()
cur.execute(query_match)
matches_list = cur.fetchall()
cur.close()
# I knew there was a way! THANK YOU!
# https://stackoverflow.com/questions/11276473/append-to-a-dict-of-lists-with-a-dict-comprehension
# Grabbing everything by obsdate from matches_list
date_dict = {k['obsdate'] : list(filter(lambda x:x['obsdate'] == k['obsdate'], matches_list)) for k in matches_list}
alert_matches_dict = {}
all_trans_matches = []
all_alerts_matches = []
# Grabbing DECAM ledger if not already fed in
if ledger_type.upper() == 'DECAM_TAMU':
if ledger_df.empty:
ledger_df = access_decam_data('https://datahub.geos.tamu.edu:8000/decam/LCData_Legacy/')
# Iterating through the dates and checking each tile observed on each date
# It is done in this way to cut down on calls to ALERCE since we go day by day
# It's also a convenient way to organize things
for date, row in date_dict.items():
date_str = str(date)
date_str = date_str[:4]+"-"+date_str[4:6]+"-"+date_str[6:] # Adding dashes for Time
obs_mjd = Time(date_str).mjd
# This method is *technically* safer than doing a double list comprehension with set albeit slower
# The lists are small enough that speed shouldn't matter here
unique_tileid = {i['tileid']: (i['tilera'], i['tiledec']) for i in row}
exposure_ras, exposure_decs = zip(*unique_tileid.values())
# Grabbing alerce ledger if not done already
if ledger_type.upper() == 'ALERCE':
if ledger_df.empty:
ledger_df = access_alerts(lastmjd = obs_mjd - 28) # Modified Julian Day #.mjd
elif ledger_type.upper() == 'DECAM_TAMU':
pass
else:
print("Cannot use alerts broker/ledger provided. Stopping before match.")
return {}
#Reatin tileid
tileid_arr = np.array(list(unique_tileid.keys()))
# Where the magic/matching happens
trans_matches, alert_matches, trans_ids, alerts_ids, _ = \
inner_matching(target_ids_in = tileid_arr, target_ras_in = exposure_ras, target_decs_in = exposure_decs, obs_mjd_in = obs_mjd,
path_in = '', max_sep = 1.8, sep_units = 'deg', ledger_df_in = ledger_df, ledger_type_in = ledger_type)
# Add everything into one giant list for both
if trans_matches.size:
#print(date, "-", len(trans_matches), "matches")
all_trans_matches.append(trans_matches)
all_alerts_matches.append(alert_matches)
else:
#print("No matches on", date)
continue
# Prepping output
# Populating the dictionary by date (a common theme)
# Each element in the dictionary thus contains the entire sqlite3 row (all info from sql tables with said headers)
alert_matches_dict[date] = []
for tup in trans_matches:
ra = tup.ra.deg
dec = tup.dec.deg
match_rows = [i for i in row if (i['tilera'], i['tiledec']) == (ra, dec)] # Just rebuilding for populating, this shouldn't change/exclude anything
alert_matches_dict[date].extend(match_rows)
return alert_matches_dict
# ## closer_check
# **closer_check** is also a handling function but operates differently in that now it is checking individual targets. This *must* be run after **initial_check** because it takes as input the dictionary **initial_check** spits out. It then grabs all the targets from the DESI files and pipes that into the matching function but this time with a much more strict matching radius (in this case 2 arcseconds).
#
# It then preps the data for output and writing.
def closer_check(matches_dict = {}, ledger_df = None, ledger_type = '', exclusion_list = []):
all_exp_matches = {}
if not matches_dict:
print("No far matches fed in for nearby matching. Returning none.")
return {}
# Again just in case the dataframe isn't fed in
if ledger_type.upper() == 'DECAM_TAMU':
id_head = 'ObjectID'
ra_head = 'RA-OBJECT'
dec_head = 'DEC-OBJECT'
if ledger_df.empty:
ledger_df = access_decam_data('https://datahub.geos.tamu.edu:8000/decam/LCData_Legacy/')
count_flag=0
# Iterating through date and all tile information for that date
for date, row in matches_dict.items():
print("\n", date)
if date in exclusion_list:
continue
# Declaring some things
all_exp_matches[date] = []
alert_exp_matches = []
file_indices = {}
all_targ_ras = np.array([])
all_targ_decs = np.array([])
all_targ_ids = np.array([])
all_tileids = np.array([])
all_petals = np.array([])
# Iterating through each initial match tile for every date
for i in row:
# Grabbing the paths and iterating through them to grab the RA's/DEC's
exp_paths = '/'.join((exposure_path, "daily/exposures", str(i['obsdate']), "000"+str(i['expid'])))
#print(exp_paths)
for path in glob_frames(exp_paths):
#print(path)
targ_ras, targ_decs, targ_ids = read_fits_info(path, transient_candidate = False)
h=fits.open(path)
tileid = h[0].header['TILEID']
tileids = np.full(len(targ_ras),tileid).tolist()
petal = path.split("/")[-1].split("-")[1][-1]
petals = np.full(len(targ_ras),petal).tolist()
# This is to retain the row to debug/check the original FITS file
# And to pull the info by row direct if you feel so inclined
all_len = len(all_targ_ras)
new_len = len(targ_ras)
if all_len:
all_len -= 1
file_indices[path] = (all_len, all_len + new_len) # The start and end index, modulo number
else:
file_indices[path] = (0, new_len) # The start and end index, modulo number
if len(targ_ras) != len(targ_decs):
print("Length of all ras vs. all decs do not match.")
print("Something went wrong!")
print("Continuing but not adding those to match...")
continue
# All the ras/decs together!
all_targ_ras = np.append(all_targ_ras, targ_ras)
all_targ_decs = np.append(all_targ_decs, targ_decs)
all_targ_ids = np.append(all_targ_ids, targ_ids)
all_tileids = np.append(all_tileids, tileids)
all_petals = np.append(all_petals, petals)
date_mjd = str(date)[:4]+"-"+str(date)[4:6] + "-" + str(date)[6:] # Adding dashes for Time
date_mjd = Time(date_mjd).mjd
# Grabbing ALERCE just in case
# Slow
if ledger_type.upper() == 'ALERCE':
id_head = 'oid'
ra_head = 'meanra'
dec_head = 'meandec'
if ledger_df.empty:
ledger_df = access_alerts(lastmjd_in = obs_mjd - 45) # Modified Julian Day #.mjd
# Checking for NaNs, again doesn't play nice with match_coordinates_sky
nan_ra = np.isnan(all_targ_ras)
nan_dec = np.isnan(all_targ_decs)
if np.any(nan_ra) or np.any(nan_dec):
print("NaNs found, removing them from array before match.")
#print("Original length (ra, dec): ", len(target_ras), len(target_decs))
nans = np.logical_not(np.logical_and(nan_ra, nan_dec))
all_targ_ras = all_targ_ras[nans] # Logic masking, probably more efficient
all_targ_decs = all_targ_decs[nans]
all_targ_ids = all_targ_ids[nans]
all_tileids = all_tileids[nans]
all_petals = all_petals[nans]
# Where the magic matching happens. This time with separation 2 arcseconds.
# Will be cleaned up (eventually)
alert_exp_matches, alerts_matches, targetid_exp_matches, id_alerts_matches, exp_idx = inner_matching(target_ids_in =all_targ_ids, \
target_ras_in = all_targ_ras, target_decs_in = all_targ_decs, obs_mjd_in = date_mjd,
path_in = '', max_sep = 2, sep_units = 'arcsec', ledger_df_in = ledger_df, ledger_type_in = ledger_type)
date_arr=np.full(alerts_matches.shape[0],date)
#print(date_arr.shape,targetid_exp_matches.shape,alert_exp_matches.shape, id_alerts_matches.shape,alerts_matches.shape )
info_arr_date=np.column_stack((date_arr,all_tileids[exp_idx],all_petals[exp_idx], targetid_exp_matches,alert_exp_matches.ra.deg,alert_exp_matches.dec.deg, \
id_alerts_matches,alerts_matches.ra.deg,alerts_matches.dec.deg ))
all_exp_matches[date].append(info_arr_date)
if count_flag==0:
all_exp_matches_arr=info_arr_date
count_flag=1
else:
#print(all_exp_matches_arr,info_arr_date)
all_exp_matches_arr=np.concatenate((all_exp_matches_arr,info_arr_date))
# Does not easily output to a csv since we have multiple results for each date
# so uh... custom file output for me
return all_exp_matches_arr
# ## inner_matching
# #### aka the bread & butter
# **inner_matching** is what ultimately does the final match and calls **match_coordinates_sky** with everything fed in. So really it doesn't do much other than take in all the goodies and make everyone happy.
#
# It may still be difficult to co-opt for alerce matching but that may be a project for another time.
def inner_matching(target_ids_in = np.array([]), target_ras_in = np.array([]), target_decs_in = np.array([]), obs_mjd_in = '', path_in = '', max_sep = 2, sep_units = 'arcsec', ledger_df_in = None, ledger_type_in = ''): # to be combined with the other matching thing in due time
# Figuring out the units
if sep_units == 'arcsec':
max_sep *= u.arcsec
elif sep_units == 'arcmin':
max_sep *= u.arcmin
elif sep_units == 'deg':
max_sep *= u.deg
else:
print("Separation unit specified is invalid for matching. Defaulting to arcsecond.")
max_sep *= u.arcsec
if not np.array(target_ras_in).size:
return np.array([]), np.array([])
# Checking for NaNs, again doesn't play nice with match_coordinates_sky
nan_ra = np.isnan(target_ras_in)
nan_dec = np.isnan(target_decs_in)
if np.any(nan_ra) or np.any(nan_dec):
print("NaNs found, removing them from array before match.")
#print("Original length (ra, dec): ", len(target_ras), len(target_decs))
nans = np.logical_not(np.logical_and(nan_ra, nan_dec))
target_ras_in = target_ras_in[nans] # Logic masking, probably more efficient
target_decs_in = target_decs_in[nans]
target_ids_in = target_ids_in[nans]
#print("Reduced length (ra, dec):", len(target_ras), len(target_decs))
# For quick matching if said kdtree actually does anything
# Supposed to speed things up on subsequent runs *shrugs*
tree_name = "_".join(("kdtree", ledger_type_in, str(obs_mjd_in)))
# Selecting header string to use with the different alert brokers/ledgers
if ledger_type_in.upper() == 'DECAM_TAMU':
id_head = 'ObjectID'
ra_head = 'RA-OBJECT'
dec_head = 'DEC-OBJECT'
elif ledger_type_in.upper() == 'ALERCE':
id_head = 'oid' #Check this is how id is called!
ra_head = 'meanra'
dec_head = 'meandec'
else:
print("No ledger type specified. Quitting.")
# lofty goals
# Will try to figure it out assuming it's a pandas dataframe.")
#print("Returning empty-handed for now until that is complete - Matthew P.")
return np.array([]), np.array([])
# Convert df RA/DEC to numpy arrays
alerts_id = ledger_df_in[id_head].to_numpy()
alerts_ra = ledger_df_in[ra_head].to_numpy()
alerts_dec = ledger_df_in[dec_head].to_numpy()
# Convert everything to SkyCoord
coo_trans_search = SkyCoord(target_ras_in*u.deg, target_decs_in*u.deg)
coo_alerts = SkyCoord(alerts_ra*u.deg, alerts_dec*u.deg)
# Do the matching!
idx_alerts, d2d_trans, d3d_trans = match_coordinates_sky(coo_trans_search, coo_alerts, storekdtree = tree_name) # store tree to speed up subsequent results
# Filter out the good stuff
sep_constraint = d2d_trans < max_sep
trans_matches = coo_trans_search[sep_constraint]
trans_matches_ids = target_ids_in[sep_constraint]
alerts_matches = coo_alerts[idx_alerts[sep_constraint]]
alerts_matches_ids = alerts_id[idx_alerts[sep_constraint]]
if trans_matches.size:
print(len(trans_matches), "matches with separation -", max_sep)
#sort_dist = np.sort(d2d_trans)
#print("Minimum distance found: ", sort_dist[0])
return trans_matches, alerts_matches, trans_matches_ids, alerts_matches_ids, sep_constraint
# ## Grab DECAM ledger as pandas dataframe
decam_transients = access_decam_data('https://datahub.geos.tamu.edu:8000/decam/LCData_Legacy/', overwrite = True) # If True, grabs a fresh batch
decam_transients_agn = access_decam_data('https://datahub.geos.tamu.edu:8000/decam/LCData_Legacy_AGN/', overwrite = True) # If True, grabs a fresh batch
decam_transients
# ## Run initial check (on tiles) and closer check (on targets)
init_matches_by_date = initial_check(ledger_df = decam_transients, ledger_type = 'DECAM_TAMU')
close_matches = closer_check(init_matches_by_date, ledger_df = decam_transients, ledger_type = 'DECAM_TAMU', exclusion_list = [])
np.save('matches_DECam',close_matches, allow_pickle=True)
init_matches_agn_by_date = initial_check(ledger_df = decam_transients_agn, ledger_type = 'DECAM_TAMU')
close_matches_agn = closer_check(init_matches_agn_by_date, ledger_df = decam_transients_agn, ledger_type = 'DECAM_TAMU', exclusion_list = [])
np.save('matches_DECam_agn',close_matches_agn, allow_pickle=True)
np.save('matches_DECam_agn',close_matches_agn, allow_pickle=True)
# ## A quick plot to see the distribution of target matches
plt.scatter(close_matches[:,4], close_matches[:,5],label='SN')
plt.scatter(close_matches_agn[:,4], close_matches_agn[:,5],label='AGN')
plt.legend()
# ## End notes:
# Double matches are to be expected, could be worthwhile to compare the spectra of both
| cronjobs/transient_matching/light_transient_matching.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (geoenv3)
# language: python
# name: geoenv3
# ---
# # Exploring routing tools for calculating paths for a set of origin-destination pairs.
#
# In this notebook, I explore OSRM service, OSMnx python library, and googlemaps python library (requests made to Google Maps Directions API) for computing routes and corresponding travel times and distances for a set of origin and destination pairs. Based on your use case, you can pick which one works best for you.
# +
#general libraries
from shapely.geometry import shape
from shapely.geometry import LineString
import polyline
from osgeo import ogr, osr
import geopandas as gpd
import pandas as pd
import json
import time
#specific to osrm approach
import requests
#specific to osmnx & networkx approach
import networkx as nx
import osmnx as ox
#specific to googlemaps approach
import googlemaps
#import folium
# -
# # 1. OSRM to find fastest paths
#
# OSRM is particularly useful when you want to calculate fastest routes (traffic independent conditions) and are looking for a free and open-source routing service. For more services by OSRM head to its [documentation](http://project-osrm.org/docs/v5.23.0/api/#). You can also set up your own OSRM server for bulkier requests. If you end up using OSRM in your project, do send them some love <3.
#
# Used the get_route() method discussed by <NAME> [here](https://www.thinkdatascience.com/post/2020-03-03-osrm/osrm/), and collected the 'duration' attribute as well from the response string, in the output dictionary. For more options such as alternative routes, mode of transportation (driving, bike, foot) i.e. the speed profile, do have a look at the OSRM documentation. In this example, I use [driving (car)](https://github.com/Project-OSRM/osrm-backend/blob/master/profiles/car.lua) speeds profile. You can also create your own speed profiles.
# +
#Read the csv with origin-destination pairs arranged row-wise.
#In the table used in this example, o_long and o_lat are coordinates of origin point, similarly for destination
od_table = "E:/OD_pairs1.csv"
df = pd.read_csv(od_table)
df
# -
#Method to send requests to OSRM server and parse the json response to collect distance, duration, and route information
#and return a dictionary
def get_route(origin_lon, origin_lat, dest_lon, dest_lat):
loc = "{},{};{},{}".format(origin_lon, origin_lat, dest_lon, dest_lat)
url = "http://router.project-osrm.org/route/v1/driving/"
r = requests.get(url + loc)
if r.status_code!= 200:
return {}
res = r.json()
routes = polyline.decode(res['routes'][0]['geometry'])
start_point = [res['waypoints'][0]['location'][1], res['waypoints'][0]['location'][0]]
end_point = [res['waypoints'][1]['location'][1], res['waypoints'][1]['location'][0]]
distance = res['routes'][0]['distance']
duration = res['routes'][0]['duration']
out = {'route':routes, #a list of tuples of coordinates along the route
'start_point':start_point, #list of float coords
'end_point':end_point,
'distance':distance, #in metres
'duration': duration #in seconds
}
return out
# +
def reverseTuple(lstOfTuple):
return [tup[::-1] for tup in lstOfTuple]
#extract_route() parses the list of coordinates in the dictionary returned by get_route()
#and returns the route geometry essential for outputting spatial features
def extract_route(routes_dict):
route_coords_list= routes_dict['route']
listnew = reverseTuple(route_coords_list)
return(LineString(listnew))
def extract_duration(routes_dict):
return(routes_dict['duration'])
def extract_distance(routes_dict):
return(routes_dict['distance'])
#You can also wrap above 3 methods into a single one and return a single tuple with the three elements
# +
#applying the above methods to the dataframe to calculate and add attributes to the dataframe
start_time = time.time()
df['routes'] = df.apply(lambda x: get_route(x['o_long'], x['o_lat'], x['d_long'], x['d_lat']), axis=1)
df['geometry'] = df.apply(lambda x: extract_route(x['routes']), axis=1)
df['osrm_dur'] = df.apply(lambda x: extract_duration(x['routes']),axis=1)
df['osrm_dist'] = df.apply(lambda x: extract_distance(x['routes']),axis=1)
print("Time taken: ", (time.time() - start_time), "seconds")
df
# +
#create a geodataframe and pass the geometry column, created using extract_route() method, as the geometry information of the geodataframe
gdf = gpd.GeoDataFrame(df, geometry = df['geometry'])
#Export in whichever spatial formats you need to
gdf.to_file('routes_OSRM.shp')
gdf.to_file('routes_OSRM.geojson', driver= 'GeoJSON')
# -
# # 2. OSMnx and Networkx for shortest paths
#
# OSMnx is a very handy tool for network analysis which also allows you to easily access and import road network graph objects based on OSM data. OSMnx library allows you quite a lot of control over the graph behavior and a neat interface to model the networks. Read more about this project on its [documentation](https://osmnx.readthedocs.io/en/stable/#) page for all its modules and also [here](https://geoffboeing.com/2016/11/osmnx-python-street-networks/).
#
# In this example, I use OSMnx
# 1) to import street graph for the region <br/>
# 2) to find nearest network node for each origin and destination point and
#
# I use Networkx to calculate the shortest path using the graph built by OSMnx.
#My data points lie in Bangalore region.
#OSMnx has a cool feature where you can extract the streets graph just by inputting the region name
G= ox.graph_from_place('Bangalore, India')
fig, ax = ox.plot_graph(G)
# If you prefer importing streets network graph for a specific study region/ neighborhood and have a boundary shapefile of the same,
# you can uncomment and use the lines below and ignore the lines above. For more ways to import graph, check out the osmnx graph module
# [here](https://osmnx.readthedocs.io/en/stable/osmnx.html#module-osmnx.graph).
# +
# boundary = "path_to_file/boundary.shp"
##Extract
# file = ogr.Open(boundary)
# layer = file.GetLayer()
# feature = layer.GetFeature(0)
# geom= feature.GetGeometryRef()
# feature_json = geom.ExportToJson()
# loadasdict = json.loads(feature_json)
# geom2 = shape(loadasdict)
# G = ox.graph_from_polygon(geom2)
# G2 = ox.consolidate_intersections(G, tolerance=10, rebuild_graph=True)
# -
#Read the Origin-Destination csv
od_table = 'E:/OD_pairs1.csv'
df = pd.read_csv(od_table)
# The methods below are built based on the steps outlined by Geoff Boeing (also the author of osmnx) [here](https://stackoverflow.com/a/58311118/7105292).
# +
def nodes_to_linestring(path):
coords_list = [(G.nodes[i]['x'], G.nodes[i]['y']) for i in path ]
#print(coords_list)
line = LineString(coords_list)
return(line)
def shortestpath(o_lat, o_long, d_lat, d_long):
nearestnode_origin, dist_o_to_onode = ox.distance.get_nearest_node(G, (o_lat, o_long), method='haversine', return_dist=True)
nearestnode_dest, dist_d_to_dnode = ox.distance.get_nearest_node(G, (d_lat, d_long), method='haversine', return_dist=True)
#Add up distance to nodes from both o and d ends. This is the distance that's not covered by the network
dist_to_network = dist_o_to_onode + dist_d_to_dnode
shortest_p = nx.shortest_path(G,nearestnode_origin, nearestnode_dest)
route = nodes_to_linestring(shortest_p) #Method defined above
# Calculating length of the route requires projection into UTM system. Using
inSpatialRef = osr.SpatialReference()
inSpatialRef.ImportFromEPSG(4326)
outSpatialRef = osr.SpatialReference()
outSpatialRef.ImportFromEPSG(32643)
coordTransform = osr.CoordinateTransformation(inSpatialRef, outSpatialRef)
#route.wkt returns wkt of the shapely object. This step was necessary as transformation can be applied
#only on an ogr object. Used EPSG 32643 as Bangalore is in 43N UTM grid zone.
geom = ogr.CreateGeometryFromWkt(route.wkt)
geom.Transform(coordTransform)
length = geom.Length()
#Total length to be covered is length along network between the nodes plus the distance from the O,D points to their nearest nodes
total_length = length + dist_to_network
#in metres
return(route, total_length )
# +
start_time = time.time()
df['osmnx_geometry'] = df.apply(lambda x: shortestpath(x['o_lat'], x['o_long'], x['d_lat'], x['d_long'])[0] , axis=1)
df['osmnx_length'] = df.apply(lambda x: shortestpath(x['o_lat'], x['o_long'], x['d_lat'], x['d_long'])[1] , axis=1)
print("Time taken: ", (time.time() - start_time), "seconds")
df
#Note that the lambda function returns a tuple. While applying the function, have add [0] and [1] to return only one of the two outputs.
#There must be a nicer way to add both outputs in one go. This was more of a fluke try which worked. Alternatively, you could define two functions to return both separtely, but might be an overkill
# -
#rename osmnx_geometry column to 'geometry' to pass it as the geometry component to the new geo dataframe
df = df.rename(columns = {'osmnx_geometry': 'geometry'})
gpdf = gpd.GeoDataFrame(df, geometry =df['geometry'])
gpdf.to_file('osmnx_shortestpaths.shp')
# # 3. Googlemaps Directions API for routes with travel times factoring traffic
#
# At the time of this writing, calls to the directions API yields neither the absolute shortest path nor the absolute fastest path. As per the [API documentation](https://developers.google.com/maps/documentation/directions/overview), it returns the "most-efficient" route, taking into account travel-time, distance, number of turns, and so on. However, if you are really looking for shortest route or traffic based fastest route, as pointed out by [this answer](https://stackoverflow.com/a/37844795/7105292), you can request for all alternatives and tweak the code below to select only the shortest or fastest route from all the routes returned. Do go through the documentation for info on how you can pass other parameters meaningful to your use case.
#
# Do note that you might also need to create a Google Maps Platform billing account (separate from the Google Cloud Platform billing account) to be able to access the traffic based travel times. The monthly free credits should allow you to make considerable amount of requests. This [pricing sheet](https://cloud.google.com/maps-platform/pricing/sheet/) might help too.
# +
od_pairs_csv = 'E:/OD_pairs1.csv'
df= pd.read_csv(od_pairs_csv)
df
# -
gmaps = googlemaps.Client(key ='AI..YOUR_API_KEY')
# +
def get_route(pickup_lon, pickup_lat, dropoff_lon, dropoff_lat):
origin = (pickup_lat,pickup_lon)
dest =(dropoff_lat, dropoff_lon)
directions_response = gmaps.directions(origin, dest, mode = "driving", departure_time = 1613397816)
# Always input a future time in epoch seconds. Can use an online converter E.g. https://www.epochconverter.com/
return directions_response
#long, lat has to be passed in x,y format for shapely linestring object, hence
#the tuples in the list returned by directions() function need to be inverted before constructing the linestring geometry
def reverseTuple(lstOfTuple):
return [tup[::-1] for tup in lstOfTuple]
def extract_route_geom(directions_response):
line = directions_response[0]['overview_polyline']['points']
route = polyline.decode(line)
reverse_tup_list = reverseTuple(route)
return(LineString(reverse_tup_list))
def extract_gdis(directions_response):
gdis = directions_response[0]['legs'][0]['distance']['value']
return gdis
def extract_gdur(directions_response):
gdur = directions_response[0]['legs'][0]['duration']['value']
return gdur
def extract_gdur_traffic(directions_response):
gdur_traf = directions_response[0]['legs'][0]['duration_in_traffic']['value']
return gdur_traf
# +
start_time = time.time()
df['responses'] = df.apply(lambda x: get_route(x['o_long'], x['o_lat'], x['d_long'], x['d_lat']), axis = 1)
df['geometry'] = df.apply(lambda x: extract_route_geom(x['responses']), axis = 1)
df['gdis'] = df.apply(lambda x: extract_gdis(x['responses']), axis = 1)
df['gdur'] = df.apply(lambda x: extract_gdur(x['responses']), axis = 1)
df['gdur_traffic'] = df.apply(lambda x: extract_gdur_traffic(x['responses']), axis = 1)
print("Time taken: ", (time.time() - start_time), "seconds")
df
# +
#geodataframe will not accept 'responses' as a valid field. Hence before exporting this into a geodataframe, save the entire dataframe df as a csv
#so that 'responses' can be saved for any future analysis. (avoid re-runs at a later time)
df.to_csv('gmaps_routes_responses.csv')
df = df.drop(columns = ['responses'])
gdf = gpd.GeoDataFrame(df, geometry = df['geometry'])
gdf.to_file("gmaps_routes.shp", ignore_index=True)
# -
# # Visualise the routes extracted
# Go ahead and visualise the shapefiles in QGIS or any spatial software you prefer! You can also use folium to visualise within the notebook. Here I attach an image of how these routes look. As you can see, they do not coincide.
# * <b><font color = 'green'> Green lines </font></b>: Fastest routes (OSRM).
# * <b><font color = 'blue'> Blue lines </font></b>: Shortest routes (OSMnx).
# * <b><font color = 'red'> Red lines </font></b>: Traffic based 'most-efficient' routes (Google Maps)
# * Basemap used is Wikimedia Labelled Layer.
#
# 
| Routing_Libraries_Services.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Analysis
# language: python
# name: ana
# ---
# # Basics of Electoral Bias Measures
#
# *<NAME><br>
# University of Bristol<br>
# <EMAIL><br>
# (Supported by NSF #1657689)*
# So far, we have learned quite a bit about the structural models of elections that model the expected wins a party has in the legislature as a function of how well they do in an election. These *seats-votes* models are fundamental to many different kinds of electoral bias measures. To this point, we've talked about electoral bias a *specific parameter* in a given model of the seats-votes curve (see Chapter 00 in the discussion of the Cube Rule & the Log-Odds election model). This works when we state a specific model for the seats-votes curve that provides a "bias" parameter, but many of the methods (including the simple bootstrap method we've been using so far) do not admit a direct parameter that measures the "bias" of a given seats-votes curve.
#
# So, how can we measure bias? Well, using the simulations from something like the seats-votes curve estimators, we can obtain nearly any measure of bias. Further, armed with a simulator, we can also get somewhat reasonable estimates of uncertainty about the bias (so long as we have confidence that the simulated elections actually represent *realistic* scenarios that might actually be experienced in elections). This notebook will walk through three fundamental ideas in the estimation of "partisan bias," and will talk about four measure of partisan bias, including:
#
# 1. partisan symmetry
# 2. mean-median discrepancy
# 3. attainment gaps
# 4. efficiency gaps
# First, though, like before, we'll need to import some of the Python packages necessary to do the data processing, plotting, and mathematics:
# +
import seatsvotes
import numpy
import pandas
from scipy import stats
numpy.random.seed(111211)
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# This time, we'll use the same example data set provided in the `seatsvotes` package:
df = seatsvotes.data.congress(geo=True)
# And focus only on the state of Illinois:
data = df[df.state_name.str.match('illinois')]
# Then, we'll fit the same Bootstrap-style estimator:
boot = seatsvotes.Bootstrap(data, district_id='lewis_dist')
# And, for visualization purposes, show what the estimated seats-votes curve looks like below:
boot.plot_simulated_seatsvotes(year=2010, n_sims=1000)
plt.xlabel('% Democrat District Average ')
plt.ylabel('% Democrat Seats')
plt.title('Seats Votes Curve, IL 2010')
# Now, many measures of partisan bias focus on this curve. In general, these measures tend to compare the difference in shape between this curve and some *idealized* version of this curve that represents a fair relationship between the seats a party wins in a congressional delegation and its performance in the election. Further, many measures also have a specific reference to a given ethical question, statement, or principle that is considered to be indicative of "fairness" in an electoral system. Thus, we will discuss these concepts for a few common partisan bias measures. Specifically, we'll cover the four bias measures we mention in the introduction, starting with the most commonly-accepted concept, partisan symmetry.
#
#
# # Basic Partisan Symmetry
# The core concept & main idea of partisan symmetry focuses on ensuring parties with similar popular support achieve similar levels of government control. Stated explicitly:
#
# > parties with similar levels of popular support should be given similar levels of representation in Congress
#
# At its core, then, this means that two parties that recieve the same average district vote share should probably also get a similar number of seats in Congress. Since the seats-votes curve proides a prediction of how many seats a party wins as a function of the average district vote share, partisan symmetry measures can assess the asymmetry between the fraction of seats in congress a party wins and their average level of support in districts across the country. In a two-party system like the US, partisan symmetry can be assessed in either of the following scenarios:
#
# 1. Party A wins $s\%$ of the Congress when they win $h\%$ average district vote share. When party B wins $h\%$ average district vote share, does party B also win $s\%$ of the Congress?
# 2. Party A and Party B have equivalent average district vote shares, $h\% = (100-h)\% = 50\%$. Do party A and party B win the same number of seats?
#
# In a strict sense, it's *always* easier to analyze the second scenario than the first. Say Democrats win an average of 44% of the vote in each district. Scenario 1 requires that we construct a prediction about what would happen when Dems win $100-44 = 56\%$ on average. This prediction is $12\%$ away from the reality we *did* observe, where Dems won $44\%$. But, using scenario 2, we only have to create a prediction of what happens when Dems and Republicans both get 50% of the vote. This is a scenario that's only $6\%$ away from the case we did observe. Thus, it's a **much** smaller extrapolation than the one that requires us to predict what happens under a $12\%$ swing. Thus, it's simpler and more effective for analysts to consider what happens when both parties recieve an equivalent percent of the vote.
#
# *(As an aside, in my dissertation, I interviewed officials who worked for the nonpartisan redistricting commissions in both Washington and Arizona. Both groups of participants agreed that scenario 2 is realistic in their home states, but that scenario 1 is unlikely. That is, they said it was indeed possible for parties to win about the same average percentage vote in districts, but they thought it highly improbable that the state would **flip** from one party to the same split for the other party. Thus for both methodological and validity reasons, it's better to analyze scenario 2 than 1.)*
# Focusing on the area of the seats-votes curve near where both parties would win 50% on average:
# +
f,ax,sims,ranks = boot.plot_simulated_seatsvotes(year=2010, n_sims=1000,
band=True, silhouette=True,
return_sims=True)
n_districts = sims[0].shape[0]
boot.plot_empirical_seatsvotes(year=2010,
scatter_kw=dict(marker='x', s=100,
color='blue', label='Observed'),
ax=ax)
win_the_delegation = numpy.hstack([sim[numpy.where(rank==4)]
for sim,rank in zip(sims,ranks)])
ax.hlines(numpy.arange(0,n_districts+1)/n_districts,
0,1, color='lightgrey', zorder=-1)
plt.vlines(.5,0,1, linestyle=':')
ax.set_ylim(.4, .66)
ax.set_xlim(.4,.6)
ax.set_title('The Median Seats')
ax.legend()
plt.show()
# -
# From this, we can see that, when both parties win 50% on average, Democrats tend to win one seat below a majority, or they tend to win one or two seats above a majority. But, we really want to hone in on the replications where Democrats & Republicans win nearly the same average district vote share. Simply looking in aggregate is not enough. To do this, we can see the full distribution of Democrat average voteshares for each simulation, and highlight the band where simulations have Democrats winning between 49.5% & 50.5% district average voteshare on average:
plt.hist(sims.mean(axis=1), density=True, color='k', label='Simulated')
plt.title('Average District Vote Share')
plt.vlines(boot.wide[-1].vote_share.mean(), 0,1, color='r', linewidth=2, label='Observed')
plt.vlines((.495,.505), 0,5.4, color='lightblue', label='Nearly 50-50')
plt.legend(fontsize=14)
# As we'd expect, most of the simulations cluster at or around the observed average vote share for Democrats, about 55%. But, we need to focus on simulations where Democrats and Republicans both win about 50% of the vote in districts, on average. Thus, we will cut out all the simulations where the average district vote share falls in 49.5% & 50.5%:
nearly_even = numpy.abs(sims.mean(axis=1) - .5) < .005
# There are only a few simulations that fall within this range:
nearly_even.sum()
# We can artificially inflate this number using a few numerical tricks, but we'll focus on the raw results without them for now. Assuming that results are functionally the same when observations are between 49.5 & 50.5, we first get the percent of the congress Dems win when they get between 49.5 and 50.5% in districts on average:
dem_wins_in_even_elex = sims[nearly_even] > .5
dem_seatshares_in_even_elex = dem_wins_in_even_elex.mean(axis=1)
# Breaking this down by the number of simulations where dems win a given fraction of seats, we see the following distribution for simulations that fall within 49.5% and 50.5%:
fractions, counts = numpy.unique(dem_seatshares_in_even_elex, return_counts=True)
print('Simulations with Democrat District \nAverage Vote between 49.5% & 50.5%')
pandas.DataFrame({'Dem Wins':fractions, 'N Sims':counts})
# Thus, on average, there's around 11 simulations where the Dems win a majority in congress and 8 simulations where Dems fail to win a majority in Congress when parties achieve about the same percentage of the average district vote share. On average, then, Dems win:
print('{:.2f} percent'.format(dem_seatshares_in_even_elex.mean()*100))
# of the seats in the Illinois congressional delegation when they win the same vote share in districts as Republicans. Thus, you can say the system is biased *very slightly* against Democrats in Illinois congressional elections, but the size of this bias is so small that it's unlikely to be practically significant under any meaningful criteria of interpretation.
# # One Alternative: the Mean-Median Gap
# Related to ideas about partisan symmetry, the mean-median gap is another way that we can characterize the structure of bias. The measure focuses on the fact that the *median* district in each simulation is the district which splits the congressional delegation 50-50. If a party tends to need a lot more votest to get 50% of the congressional delegation, then the system may be biased against that party. Thus, the gap between the mean district and the median district provides an indication of the discrepancy between the votes required to win 50% of the seats in a delegation and the average votes that the party tends to win.
#
# For example, let's focus on the 2010 election:
observed_vote = boot.wide[-1].vote_share.values
# In this election, Democrats had an average district vote share of:
observed_vote.mean()
# but had a median vote share of:
numpy.median(observed_vote)
# So, this suggests that Democrats only needed 48% on average in districts to win a bare majority of the Illinois congressional delegation. But, they actually won 54% of the vote. Thus, there are many heavily Democratic districts which skew the distribution of Democrat district vote shares. With this skew, Dems tend to win more votes in districts than they would really need to in order to win a majority of the congressional delegation. Thus, their votes are wasted, and the system is biased subtly (again) towards Republicans in Illinois.
#
# In general, we can look at this in each simulation, and we'll see that in the simulations, we get a small bias *towards* Democrats, but there's so much variability around this that there's no distinguishable bias either way.
plt.hist(numpy.median(sims, axis=1) - numpy.mean(sims, axis=1),
density=True, color='k')
plt.vlines(numpy.median(observed_vote) - numpy.mean(observed_vote), 0,1,
color='r', linewidth=2, label='Observed')
plt.vlines(numpy.mean(numpy.median(sims, axis=1) - numpy.mean(sims, axis=1)), 0,1,
color='skyblue', linewidth=2, label='Sim. Average')
plt.legend(fontsize=14)
plt.show()
# # Another Alternative: minimum attainment
# In a similar fashion, an attainment gap considers the difference in the voteshares Dems or Republicans need to get a majority of the congressional delgation. That is, we build the average district vote share for when Democrats win the congressional delegation & for when Republicans win the congressional delegation. Then, we look at the average district vote share for times when Republicans win and when Democrats win. Basically, this shows how "difficult" it is for a party to win a majority of the delegation or legislature under study. If Republicans tend to need *way* higher average vote share than Democrats, then it's harder on average for them to win control. Computing this directly, we first grab the simulations where Democrats & Republicans win:
dems_win = (sims > .5).sum(axis=1) > (n_districts * .5)
reps_win = (sims < .5).sum(axis=1) > (n_districts * .5)
# And then get the average vote share *Democrats* recieve in both of these scenarios *(We'll convert the Republican wins to Republican vote shares later, to avoid confusion)*.
average_dem_wins = sims[dems_win].mean(axis=1)
average_rep_wins = sims[reps_win].mean(axis=1)
# First, though, we can see the distributions of average voteshares for when each party wins control:
plt.hist(average_dem_wins, color='b', alpha=.5, density=True, label='D Wins')
plt.hist(average_rep_wins, color='r', alpha=.5, density=True, label='R Wins')
plt.legend(fontsize=14)
plt.show()
# Thus, in general, we see that Democrats tend to win more than 50% of the vote *regardless* of which party wins control of the congressional delegation. This means that the attainment gap in Illinois is rather large, around 7%:
(1-average_rep_wins).mean(), average_dem_wins.mean()
# This can also be assessed using a distributional distance measure, like Cohen's $d$:
n1, = average_dem_wins.shape
n2, = average_rep_wins.shape
div = (average_dem_wins.mean() - average_rep_wins.mean())
disp = ((n1-1)*average_dem_wins.var() + (n2 - 1)*average_rep_wins.var()) / (n1 + n2 - 2)
print("Cohen's d for difference in Democrat & Republican wins: {:.2f}".format(div/(disp**.5)))
# This is pretty large, suggesting that there's a substantial difference between the average vote shares in simulations where Democrats tend to win and those where Republicans tend to win. Further, this suggests that Democrats need way more votes to win a majority of the congressional delegation than Republicans (on average).
# # Another Alternative: Efficiency Gap
# The attainment gap relates conceptually this next estimate of partisan bias, the *efficiency gap*. The efficiency gap is related to the attainment gap, but considers the *turnout* in districts. Thus, it's often stated as the percentage difference in *wasted votes* between seats a party wins versus seats a party loses. Altogether, this is a *seat-level* attainment measure, whereas the attainment measure noted above is a *delegation-level* attainment measure. Together, they both provide a direct measurement of how much easier (or more difficult) it is for a party to win seats; this is unlike the symmetry measures, which only provide an expectation of the wins or losses in a hypothetical scenario.
#
# It's this reason attainment-style measures have been more recently favored in legislation and jurisprudence. Specifically the efficiency gap has seen popular adoption, and is one measure that is gaining significant public attention in addition to academic interest. It is important to note, however: the same thing happened for partisan symmetry measures around the *Bandemer v. Davis (1984)* decision, as well as the *LULAC v. Perry (2006)* decision.
#
# In order for us to examine efficiency gaps, we need the observed turnout:
turnout = boot.wide[-1].turnout.values[None,::]
# Then, the efficiency gap is built from the *raw votes* that are wasted by parties. The efficiency gap considers two kinds of votes as "wasted":
#
# 1. votes cast for a candidate that loses. These can be called *losing votes*, and are all votes cast for the party that loses the district.
# 2. votes cast for a candidate that wins, but aren't needed for that candidate to win. These can be called *excess votes*, and are all votes past the 50%+1th vote that the candidate gets in order to win election.
#
# Thus, the total waste for party *A*, called $w_A$, is the sum of losing & excess votes. The efficiency gap is the difference in waste, divided by the total turnout, $N_v$:
#
# $$ \frac{w_{A} - w_{B}}{N_{v}} $$
#
# If the system tends to favor party $A$, the gap is negative, since party $B$ has a higher waste than party $A$. In general, we can compute this by building the losing & excess votes for each party:
dem_in_excess_of_victory = (turnout * (sims - .5) * (sims > .5)).sum(axis=1)
dem_for_losers = (turnout * sims * (sims < .5)).sum(axis=1)
rep_in_excess_of_victory = (turnout * ( (1-sims) - .5) * ((1-sims) > .5) ).sum(axis=1)
rep_for_losers = (turnout * (1-sims) * ((1-sims) < .5) ).sum(axis=1)
# And then their total waste is the sum of the excess and losing otes:
dem_waste = dem_in_excess_of_victory + dem_for_losers
rep_waste = rep_in_excess_of_victory + rep_for_losers
# Building the empirical waste is also possible using the same steps:
# +
empirical_dem_eovs = ((observed_vote > .5) * (observed_vote - .5) * turnout).sum()
empirical_dem_lost = ((observed_vote < .5) * (observed_vote) * turnout ).sum()
empirical_dem_waste = empirical_dem_eovs + empirical_dem_lost
empirical_rep_eovs = (((1-observed_vote) > .5) * ((1-observed_vote)- .5) * turnout).sum()
empirical_rep_lost = (((1-observed_vote) < .5) * (1-observed_vote) * turnout ).sum()
empirical_rep_waste = empirical_rep_eovs + empirical_rep_lost
# -
# And, finally, we grab the difference in the wastes and divide by the total number of votes cast across all districts. Together this provides an indication of what percentage of the system overall is wasted, and gives a measure of whether Republicans or Democrats tend to waste more. Again, recall that if the statistic is stated:
#
# $$ \frac{w_{D} - w_{R}}{N_{v}} $$
#
# Then a positive efficiency gap will indicate bias *against Democrats*, since their waste tend to be larger than Republicans. Below, we'll see the *observed* and *simulated* gap for Illinois, suggesting again that the Illinois system is slightly biased against Democrats. Here, though, the power of turnout shows through: in general, over 80% of the simulations suggest that the efficiency gap shows bias against Democrats. While this wouldn't rise to a classical *pseudo*-significance level used in typical social science, this is not necessarily the correct comparison for this application. Thus, we'll take this as weak evidence that, even in cases of random re-swinging of votes, there tends to be a slight anti-Democrat bias in Illinois for the 2010 elections (and, thus, the 2000 districting plan).
plt.hist(, density=True, label='Simulated')
plt.vlines((empirical_dem_waste - empirical_rep_waste)/turnout.sum(), 0,1, label='Observed')
plt.legend(fontsize=14)
plt.show()
# What this *does not* suggest is that the districts are *gerrymandered* against Democrats; bias itself is a necessary result of gerrymandering, but not *sufficient to demonstrate gerrymandering*!
# # Conclusion
#
# Gerrymandering is a complicated topic, and many different measures of partisan bias exist. These measures tend to focus on specific, estimated quantities about the electoral system. In general, any method that can construct a *seats-votes* curve, like the ones provided in the Python package `seatsvotes`, can be used to estimate these measures of bias.
#
# That said, many of the measures of partisan bias operationalize what *fairness* is in different ways. Measures of attainment tend to focus on how *easy* it is for a party to win control of a seat or of an entire congressional delegation. Measures of *symmetry* tend to focus instead on the parity in the number of seats parties tend to win when they win similar levels of popular support. In general, measures can disagree with one another, since they all reflect slightly different ways of thinking about and representing partisan bias. However, in my empirical studies in my dissertation, the measures all tend to agree with one another in nearly all cases; situations where measures *disagreed* even within a single simulation were incredibly rare, but do indeed arise
#
# Finally, empirical measurement of bias does *not* account for other structural factors like incumbency or racially-polarized voting, so it's important not to leap directly from
#
# > there is slight bias against party $A$
#
# to
#
# > the system is *gerrymandered* against party $A$
#
#
# The most critical component of using electoral bias measures is to *attempt* to control for these factors and measure the resulting bias *over and above* that accounted for by other factors. Incumbent gerrymandering & racial gerrymandering *are not* partisan gerrymandering, although they may intersect and obscure our ability to estimate one or the other's size, strength, or significance. In practice, models tend to get around these concerns about other kinds of gerrymandering by *attempting* to estimate the size of incumbent advantage & racial voting, and then constructing simulations where these factors *are not present*. The method used so far, bootstrapping, attempts to control for this by randomizing the extent to which *observed* incumbent or racial factors are assigned to specific districts. That said, no method is perfect. In the following chapters, I demonstrate how to use a few of these methods implemented in `seatsvotes` to simulate seats-votes curves under controlled (or random) conditions.
| notebooks/03 - Bias in a single election.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
from gensim.models import Word2Vec, KeyedVectors
import numpy as np
from itertools import product
from tqdm import tqdm
import sys
sys.path.append('../src')
from models import open_pickle
# +
MODEL_NAME = 'glove.840B'
we_model_load = KeyedVectors.load(f'../data/interim/{MODEL_NAME}_norm', mmap='r')
RESULTS_FILEPATH = f'../data/interim/{MODEL_NAME}_association_metric_exps.pickle'
EXPERIMENT_DEFINITION_FILEPATH = f'../data/interim/{MODEL_NAME}_experiment_definitions.pickle'
IMAGE_SAVE_FILEPATH = f'../reports/figures/{MODEL_NAME}_exp_results.png'
NONRELATIVE_IMAGE_SAVE_FILEPATH = f'../reports/figures/{MODEL_NAME}_nonrelative_exp_results.png'
exp_def_dict = open_pickle(EXPERIMENT_DEFINITION_FILEPATH)
results_dict = open_pickle(RESULTS_FILEPATH)
# -
def add_axes_obj_labels(ax, exp_num, target_label, A_label, B_label, n_samples):
TITLE_FONT_SIZE = 12
[target_label, A_label, B_label] = [s.upper() for s in [target_label, A_label, B_label]]
ax.set_title(f'#{exp_num}: {target_label} terms: {B_label} (left) vs. {A_label} (right)',
fontsize=TITLE_FONT_SIZE)
ax.set_xlabel(f'Bias Regions: CI with {n_samples} samples')
ax.set_ylabel(f'Word')
ax.yaxis.set_ticklabels([])
def annotate_points(ax, terms, x_array, y):
POINT_FONT_SIZE = 9
for i, txt in enumerate(terms):
ax.annotate(txt, (x_array[i], y[i]), fontsize=POINT_FONT_SIZE)
def add_scatters_and_lines(ax, arr, threshold,
mean, QR_dict, y):
S = 20 # Marker size
ZERO_LINE_COLOR = 'lime'
POINT_COLOR = 'red'
PERCENTILES_COLOR = 'blue'
SHADE_DARKNESS = 0.2
SHADE_DARKNESS_80CI = 0.1
SHADE_DARKNESS_90CI = 0.15
SHADE_DARKNESS_95CI = 0.25
CI_COLOR = 'black'
XAXIS_LIMIT = 0.2
y = [i for i in range(1,len(arr)+1)]
QR_95 = QR_dict['QR_95']
ax.scatter(arr, y, c=POINT_COLOR, s=S)
ax.xaxis.grid()
ax.axvspan(QR_95[0], QR_95[1], alpha=SHADE_DARKNESS, COLOR=CI_COLOR)
#ax.axvline(threshold_second, color=POINT_COLOR, linestyle='-.', label='threshold')
#ax.axvline(-threshold_second, color=POINT_COLOR, linestyle='-.')
#ax.axvline(mean_second, c=POINT_COLOR, label='second-order mean')
#ax.axvspan(lower_bound, upper_bound, alpha=SHADE_DARKNESS, color=PERCENTILES_COLOR)
#ax.axvspan(ST1_80CI[0], ST1_80CI[1], alpha=SHADE_DARKNESS_80CI, color=CI_COLOR)
#ax.axvspan(ST1_90CI[0], ST1_90CI[1], alpha=SHADE_DARKNESS_90CI, color=CI_COLOR)
#ax.axvspan(ST1_95CI[0], ST1_95CI[1], alpha=SHADE_DARKNESS_95CI, color=CI_COLOR)
#ax.axvspan(pct_5_second, pct_95_second, alpha=SHADE_DARKNESS, color=PERCENTILES_COLOR)
ax.set_xlim(-XAXIS_LIMIT, XAXIS_LIMIT)
results_dict
# +
fig, axs = plt.subplots(10,2, figsize=(15,50))
LEGEND_SIZE = 10
exps = range(1,11)
target_letters = ['X','Y']
for exp_num, target_letter in tqdm(product(exps, target_letters), total=20):
col = 0 if target_letter =='X' else 1
ax = axs[exp_num-1, col]
arr = results_dict[exp_num]['second'][f'{target_letter}_array']
threshold = results_dict[exp_num]['second']['threshold']
mean = results_dict[exp_num]['second'][f'{target_letter}_mean']
n_samples = -1 #len(results_dict[exp_num]['second']['sigtest_dist_1'])
y = [i for i in range(1,len(arr)+1)]
terms = exp_def_dict[exp_num][f'{target_letter}_terms']
target_label = exp_def_dict[exp_num][f'{target_letter}_label']
A_label = exp_def_dict[exp_num]['A_label']
B_label = exp_def_dict[exp_num]['B_label']
QR_dict = results_dict[exp_num]['second']['QR_dict']
add_scatters_and_lines(ax, arr, threshold,
mean, QR_dict, y)
annotate_points(ax, terms, arr, y)
add_axes_obj_labels(ax, exp_num, target_label, A_label, B_label, n_samples)
axs[0,0].legend(loc=2, prop={'size': LEGEND_SIZE})
fig.tight_layout(pad=2)
print('Rendering...')
plt.savefig(IMAGE_SAVE_FILEPATH)
plt.show()
# -
| notebooks/SingleWordViz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # sklearn-svm
#
# > In machine learning, [support vector machines](https://en.wikipedia.org/wiki/Support_vector_machine) (SVMs, also support vector networks) are supervised learning models with associated learning algorithms that analyze data used for classification and regression analysis.
# +
import numpy as np
from sklearn.svm import SVC
import matplotlib.pyplot as plt
# !pip install mlxtend;
from mlxtend.plotting import plot_decision_regions
# -
# First, it's a good idea to define a random seed for reproducibility:
# define a random seed for reproducibility
np.random.seed(6)
# generate some random data
X = np.random.randn(200, 2)
y = X[:, 1] > np.absolute(X[:, 0])
y = np.where(y, 1, -1)
# +
# define our SVM
svm = SVC(kernel='rbf', random_state=0, gamma=0.5, C=10.0)
svm.fit(X, y)
plot_decision_regions(X, y, svm, markers=['o', 'x'], colors='blue,magenta')
plt.show()
| 01_introduction/1.1_sklearn-svm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="QsvsV62klMux" outputId="06635491-dd7e-4646-ddfd-66075915b727"
# ! pip install transformers
# ! pip install sentencepiece
# + colab={"base_uri": "https://localhost:8080/"} id="DEckzhzD1mWl" outputId="87408bfa-7b0d-4d9c-b557-c8209e4b0991"
from google.colab import drive
drive.mount('/content/drive')
# + id="jC8PigZadfmZ"
import json
from collections import Counter
import pandas as pd
# + id="q_lbaaMedXoE"
with open("/content/drive/My Drive/rsicd/dataset_rsicd.json") as f:
ds = json.load(f)
# + id="i9HztzB3dMw8"
# Setting up the device for GPU usage
from torch import cuda
device = 'cuda' if cuda.is_available() else 'cpu'
# + id="jpxQicklr8sz"
# Storing tuples of image id and sentences in a list
all_sentences = []
for image in ds['images']:
for sentence in image['sentences']:
all_sentences.append((image['imgid'],sentence['raw']))
# + id="aHCg8oROsptb"
# Creating a counter to get counts of how many times each sentence was repeated in the same image
sentence_counts = dict(Counter(all_sentences))
# + id="8tj_v25Rs3ex"
# Converting the dictionary to a tuple - (image_id, sentence, count)
image_id_sentences_and_counts = []
for sentence, count in sentence_counts.items():
image_id_sentences_and_counts.append((sentence[0], sentence[1], count))
# + id="2AfT4N67tRNw"
sentences_df = pd.DataFrame(image_id_sentences_and_counts, columns=['image_id', 'sentence', 'count'])
# + id="6nXYXod3ydrC"
translation_languages_priority = ['fr','es','it', 'pt']
# + id="nruQa3uttleK"
# Creating a dictionary with with languages to use for backtranslation
# eg. If the count = 2, one of them has to be backtranslated, and `fr` will be used for that.
languages_to_back_translate_with = {2 : translation_languages_priority[0],
3 : translation_languages_priority[0:2],
4 : translation_languages_priority[0:3],
5 : translation_languages_priority}
# + id="Cz_O-EvduMs0"
# we dont need backtranslation if count = 1
sentences_df.drop(sentences_df[sentences_df['count'] == 1].index, inplace = True)
# + id="4YfcLCGiuTPD"
# Creating a column that has the languages to be used for running the backtranslation
sentences_df['languages_to_back_translate_with'] = sentences_df['count'].map(languages_to_back_translate_with)
# + id="gzyzXuefuljV"
# Creating a new row for each language
sentences_df = sentences_df.explode('languages_to_back_translate_with')
# + id="mspntvShuwVb"
sentences_df = sentences_df.sort_values(by=['languages_to_back_translate_with'], ignore_index=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="asow56CUu2RY" outputId="3579d2d2-6100-4f56-f26f-5061849128e9"
sentences_df.head()
# + [markdown] id="c0cG48Ncn1Bo"
# ## Translation Model.
# + id="3OnL3vN4u30V"
from transformers import MarianMTModel, MarianTokenizer
# + id="OSfXLSTHu-RY"
target_model_name = 'Helsinki-NLP/opus-mt-en-ROMANCE'
target_tokenizer = MarianTokenizer.from_pretrained(target_model_name)
target_model = MarianMTModel.from_pretrained(target_model_name)
# + id="MvSEf1PN0Tov"
en_model_name = 'Helsinki-NLP/opus-mt-ROMANCE-en'
en_tokenizer = MarianTokenizer.from_pretrained(en_model_name)
en_model = MarianMTModel.from_pretrained(en_model_name)
# + id="wam2LUv1vMw5"
def translate(texts, model, tokenizer, language="fr"):
# Prepare the text data into appropriate format for the model
template = lambda text: f"{text}" if language == "en" else f">>{language}<< {text}"
src_texts = [template(text) for text in texts]
# Tokenize the texts
encoded = tokenizer.prepare_seq2seq_batch(src_texts,return_tensors="pt")
model.to(device)
# Generate translation using model
translated = model.generate(**encoded.to(device))
# Convert the generated tokens indices back into text
translated_texts = tokenizer.batch_decode(translated, skip_special_tokens=True)
return translated_texts
# + id="ahhVb37MvZUK"
def back_translate(texts, source_lang="en", target_lang="fr"):
# Translate from source to target language
fr_texts = translate(texts, target_model, target_tokenizer,
language=target_lang)
# Translate from target language back to source language
back_translated_texts = translate(fr_texts, en_model, en_tokenizer,
language=source_lang)
return back_translated_texts
# + id="UydNhD_7vasR"
# Group the dataframe by language to translate with, so that they can be batched together.
grouped_by_language = sentences_df.groupby(['languages_to_back_translate_with'])
# + colab={"base_uri": "https://localhost:8080/"} id="s3AodggLzFXF" outputId="15491af1-507a-4596-bcc1-631d6c321245"
list_of_dataframes_for_each_lang = []
for lang in translation_languages_priority:
df = grouped_by_language.get_group(lang)
sentences = list(df['sentence'])
#Batch the sentences in groups of 64
list_of_sentence_batches = [sentences[i:i + 64] for i in range(0, len(sentences), 64)]
# list to store all translated sentences.
translated_sentences = []
for sentence_batch in list_of_sentence_batches:
batch_of_translated_sentences = back_translate(sentence_batch, target_lang = lang)
translated_sentences += batch_of_translated_sentences
df["back_translated_sentence"] = translated_sentences
list_of_dataframes_for_each_lang.append(df)
# + id="ASJUE0Ujx0jt"
sentences_with_translation_df = pd.concat(list_of_dataframes_for_each_lang, axis=0)
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="In9sYHnpEszd" outputId="b38bb3c7-eed3-40ba-c746-5694c8d5e072"
sentences_with_translation_df.head()
# + id="HEvFK-WmeEkf"
# Creating a copy of the original json, for augmentation.
text_augmented_ds = ds
# + id="MaCUaVjaGFU9"
def find_sentence_in_img(text, sentences):
for sent_index, sentence in enumerate(sentences):
print(f"{text} --AND-- {sentence['raw']}")
print(text == sentence['raw'])
if text == sentence['raw']:
print(sent_index)
return sent_index
# + colab={"base_uri": "https://localhost:8080/"} id="nlOw7FM4m8dg" outputId="19d732eb-e48f-4403-e3ef-68b6bfd89a3d"
for i,row in enumerate(sentences_with_translation_df.itertuples()):
for image_index, image in enumerate(text_augmented_ds['images']):
if image['imgid'] == row.image_id:
sent_index = find_sentence_in_img(row.sentence, image['sentences'])
print(f"image index = {image_index}")
print(f"sent index = {sent_index}")
text_augmented_ds['images'][image_index]['sentences'][sent_index]['raw'] = row.back_translated_sentence
# + colab={"base_uri": "https://localhost:8080/"} id="flpvZOd1is0n" outputId="476d2a04-1138-4df4-9d50-fe52a990cb8b"
text_augmented_ds['images'][0]
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="ENvU69RJ17G5" outputId="65aeff73-d174-4ef5-d3bd-d462434d68c4"
sentences_with_translation_df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 175} id="qa0poi5xhP3f" outputId="c992529c-5c09-4c2b-dc7d-e776363a249f"
sentences_with_translation_df[sentences_with_translation_df.sentence == 'many planes are parked next to a long building in an airport .']
# + id="J8j7IDXOjS2C"
| nbs/rsicd_text_augmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
from sklearn.preprocessing import StandardScaler
from sklearn.cluster import KMeans
from sklearn.mixture import GaussianMixture
from sklearn.cluster import AgglomerativeClustering
import warnings
warnings.filterwarnings("ignore")
import copy
import pdb
def load_data(path):
data = pd.read_csv(path, delimiter=',')
data = data.values
X = data[:, :-1]
y = data[:,-1]
return X,y
def cat2num(y):
for i in range(len(y)):
if y[i] == 'dos':
y[i] = 0
elif y[i] == 'normal':
y[i] = 1
elif y[i] == 'probe':
y[i] = 2
elif y[i] == 'r2l':
y[i] = 3
else:
y[i] = 4
return y
def activation(z, derivative=False, type=None):
# Sigmoid activation function
if type == "sigmoid":
if derivative:
return activation(z, type="sigmoid") * (1 - activation(z, type="sigmoid"))
else:
return 1 / (1 + np.exp(-z))
# ReLu activation function
if type == "relu":
if derivative:
z[z <= 0.0] = 0.
z[z > 0.0] = 1.
return z
else:
return np.maximum(0, z)
# Tanh activation function
if type == "tanh":
if derivative:
return activation(z, type="tanh") * (1 - activation(z, type="tanh"))
else:
return np.tanh(z)
def autoencoder(X,y,hidden_layer,lr,epoch,act_fn):
# np.random.seed(42)
print('\nAutoencoder Running ! \n')
W1 = np.random.randn(X.shape[1],hidden_layer)
W2 = W1.T
losses = []
for e in range(epoch):
#-----------------------Forward--------------------------
z1 = np.dot(X,W1)
a1 = activation(z1,type = act_fn)
z2 = np.dot(a1,W2)
a2 = activation(z2,type = act_fn)
loss = np.sum(np.power((a2-y),2))/len(y)
#--------------------Backpropagate-----------------------
dW2 = np.dot(a1.T,2*(a2-y)*activation(z2,derivative = True,type = act_fn))
delta = np.dot(2*(a2-y)*activation(z2,derivative = True,type = act_fn),W2.T)*activation(z1,derivative = True,type = act_fn)
dW1 = np.dot(X.T,delta)
#--------------------Weight Update------------------------
W2 = W2 - (lr*dW2)
W1 = W1 - (lr*dW1)
losses.append(loss)
print("Epoch : {}/{}".format(e+1, epoch), end='\r')
plt.figure(1)
plt.plot(range(epoch),losses)
plt.xlabel("Epochs")
plt.ylabel("MSE Loss")
plt.title('Plot of Loss vs Epoch to verify correct functioning of NN for {} Epochs'.format(epoch))
print('Data reduced to new dimention {}\n'.format(hidden_layer))
return z1
def dist(a, b):
return np.linalg.norm(a - b, axis=1)
def kmeans_clustering(x_reduced,k,D):
centroid= np.random.randint(0, np.max(x_reduced), size=k)
centroid = centroid.reshape(k,1)
for i in range (1,x_reduced.shape[1]):
centroid_i = np.random.randint(0, np.max(x_reduced), size=k) #Generate random values between 0 and max(x)-20 of k dimention
centroid_i = centroid_i.reshape(k,1)
centroid = np.hstack((centroid,centroid_i))
print("Clustering Started !!!")
print('')
print("Initial Centroids Set to:")
print(centroid)
print('')
centroid_old = np.zeros(centroid.shape)
clusters = np.zeros(len(x_reduced))
error_dist = dist(centroid, centroid_old)
while np.count_nonzero(error_dist) != 0: #Iterate until there is no change in centroid position
for i in range(len(x_reduced)): # Labelling each point w.r.t its nearest cluster
distances = dist(x_reduced[i], centroid)
cluster = np.argmin(distances)
clusters[i] = cluster
centroid_old = copy.deepcopy(centroid)
for i in range(k): #New centroid will be the average of the distances
points = [x_reduced[j] for j in range(len(x_reduced)) if clusters[j] == i] #All the points which are under ith cluster as per current clustering
centroid[i] = np.mean(points, axis=0)
error_dist = dist(centroid, centroid_old)
print('Distance moved by Centroids in next interation')
print(error_dist)
print('')
print('Clustering Completed !!!')
print('')
if D == 0:
plt.figure(2)
color=['cyan','magenta','red','blue','green']
labels=['cluster1','cluster2','cluster3','cluster4','cluster5']
for i in range(k):
points = np.array([x_reduced[j] for j in range(len(x_reduced)) if clusters[j] == i])
print('No. of points in Cluster {} : {} '.format(i,points.shape))
if points.size != 0:
plt.scatter(points[:,0],points[:,1],c=color[i],label=labels[i])
# plt.scatter(centroid[:,0],centroid[:,1],s=300,c='y',label='Centroids')
plt.title('Clusters of Network Attack after reducing data to 2-D using PCA')
plt.xlabel('Attribte 1')
plt.ylabel('Attribute 2')
plt.legend()
plt.show()
return clusters
def cal_purity(labels,y):
cnf_matrix = np.zeros((5,5))
for i in range(len(y)):
cnf_matrix[int(labels[i]),y[i]] +=1
num = 0
for i in range(5):
num += np.max(cnf_matrix[i])
return (num/len(y))
if __name__ == "__main__":
np.random.seed(26)
path = '../Dataset/intrusion_detection/data.csv'
X,y_true = load_data(path)
y_true = cat2num(y_true)
X = StandardScaler().fit_transform(X)
y = copy.deepcopy(X)
#--------------------------------Hyperparameters------------------------------------------------------
hidden_layer = 14
lr = 0.001
epoch = 100
act_fn = 'sigmoid'
print('Hyperparameters set to:\n\n Nuerons in Hidden Layer: {}\n Learning Rate: {}\n Number of Epochs: {}\n Activation Function: {}\n'.format(hidden_layer,lr,epoch,act_fn))
print('===============================================================================================')
#------------------------------------Autoencoder------------------------------------------------------
x_reduced = autoencoder(X,y,hidden_layer,lr,epoch,act_fn)
print('===============================================================================================')
#-----------------------------------------KMeans------------------------------------------------------
print('\nDoing Clustering by selecting reduced number of dimentions in PCA as per threshold of 10%\n')
clusters1 = kmeans_clustering(x_reduced,5,1)
purity1 = cal_purity(clusters1,y_true)
print('===============================================================================================')
print('\nPurity while using own developed KMeans:{}\n '.format(purity1))
kmeans = KMeans(n_clusters=5)
kmeans.fit(x_reduced)
clusters2 = kmeans.labels_
purity2 = cal_purity(clusters2,y_true)
print('Purity while using KMeans from Sci-Kit Learn:{}\n '.format(purity2))
plt.figure(2)
plt.scatter(x_reduced[:,0],x_reduced[:,1], c=kmeans.labels_, cmap='rainbow')
plt.scatter(kmeans.cluster_centers_[:,0] ,kmeans.cluster_centers_[:,1], color='black')
plt.title('Clusters of Network Attack after reducing data using Autoencoder')
plt.xlabel('Attribte 1')
plt.ylabel('Attribute 2')
print('===============================================================================================')
#-----------------------------------------GMM------------------------------------------------------------
print('\nGMM Started !!!\n')
gmm = GaussianMixture(n_components=5).fit(x_reduced)
labels = gmm.predict(x_reduced)
plt.scatter(x_reduced[:, 0], x_reduced[:, 1], c=labels, s=40, cmap='viridis');
print('GMM Completed !!!\n')
purity_gmm = cal_purity(labels,y_true)
print('Purity while using GMM: ', purity_gmm)
print('\n===============================================================================================')
plt.show()
# Hierarchical clustering
print('\nHierarchical clustering Started !!!')
clustering = AgglomerativeClustering(n_clusters=5, affinity='euclidean', linkage='single')
clustering.fit_predict(x_reduced)
labels = clustering.labels_
plt.scatter(x_reduced[:, 0], x_reduced[:, 1], c=labels, s=40, cmap='viridis');
print('Hierarchical clustering Completed !!!')
print('')
purity_hc = cal_purity(labels,y_true)
print('Purity while using HIerarchical Clustering: ', purity_hc)
print('\n===============================================================================================')
| src/q-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Partial Dependence Plots
#
# <NAME> 2019
# <NAME> 2020
#
# .. currentmodule:: skopt
#
# Plot objective now supports optional use of partial dependence as well as
# different methods of defining parameter values for dependency plots.
#
print(__doc__)
import sys
from skopt.plots import plot_objective
from skopt import forest_minimize
import numpy as np
np.random.seed(123)
import matplotlib.pyplot as plt
# ## Objective function
# Plot objective now supports optional use of partial dependence as well as
# different methods of defining parameter values for dependency plots
#
#
# Here we define a function that we evaluate.
def funny_func(x):
s = 0
for i in range(len(x)):
s += (x[i] * i) ** 2
return s
# ## Optimisation using decision trees
# We run forest_minimize on the function
#
#
# +
bounds = [(-1, 1.), ] * 3
n_calls = 150
result = forest_minimize(funny_func, bounds, n_calls=n_calls,
base_estimator="ET",
random_state=4)
# -
# ## Partial dependence plot
# Here we see an example of using partial dependence. Even when setting
# n_points all the way down to 10 from the default of 40, this method is
# still very slow. This is because partial dependence calculates 250 extra
# predictions for each point on the plots.
#
#
_ = plot_objective(result, n_points=10)
# It is possible to change the location of the red dot, which normally shows
# the position of the found minimum. We can set it 'expected_minimum',
# which is the minimum value of the surrogate function, obtained by a
# minimum search method.
#
#
_ = plot_objective(result, n_points=10, minimum='expected_minimum')
# ## Plot without partial dependence
# Here we plot without partial dependence. We see that it is a lot faster.
# Also the values for the other parameters are set to the default "result"
# which is the parameter set of the best observed value so far. In the case
# of funny_func this is close to 0 for all parameters.
#
#
_ = plot_objective(result, sample_source='result', n_points=10)
# ## Modify the shown minimum
# Here we try with setting the `minimum` parameters to something other than
# "result". First we try with "expected_minimum" which is the set of
# parameters that gives the miniumum value of the surrogate function,
# using scipys minimum search method.
#
#
_ = plot_objective(result, n_points=10, sample_source='expected_minimum',
minimum='expected_minimum')
# "expected_minimum_random" is a naive way of finding the minimum of the
# surrogate by only using random sampling:
#
#
_ = plot_objective(result, n_points=10, sample_source='expected_minimum_random',
minimum='expected_minimum_random')
# We can also specify how many initial samples are used for the two different
# "expected_minimum" methods. We set it to a low value in the next examples
# to showcase how it affects the minimum for the two methods.
#
#
_ = plot_objective(result, n_points=10, sample_source='expected_minimum_random',
minimum='expected_minimum_random',
n_minimum_search=10)
_ = plot_objective(result, n_points=10, sample_source="expected_minimum",
minimum='expected_minimum', n_minimum_search=2)
# ## Set a minimum location
# Lastly we can also define these parameters ourself by parsing a list
# as the minimum argument:
#
#
_ = plot_objective(result, n_points=10, sample_source=[1, -0.5, 0.5],
minimum=[1, -0.5, 0.5])
| 0.8/notebooks/auto_examples/plots/partial-dependence-plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from src.models.dcca import DeepCCA
from src.utils.embeddings import retrieve_all_embeds
from src.utils.files import save_as_pickle
from tensorflow import keras
model_path = "data/models/custom"
embed = retrieve_all_embeds([("data/features/use.pkl.train", "data/features/xception.pkl.train","data/features/xception.pkl.train"),
("data/features/use.pkl.dev","data/features/xception.pkl.dev", "data/features/xception.pkl.train"),
("data/features/use.pkl.test", "data/features/xception.pkl.test", "data/features/xception.pkl.train")])
X1_train, X1_dev, X1_test = embed["text only"]
X2_train, X2_dev, X2_test = embed["image only"]
outdim = 100
layers = [1000, 1000, 1000, outdim]
l2 = 1e-5
lr = 1e-3
epochs = 100
batch_size = 800
use_all_singular_values = True
dcca = DeepCCA(X1_train.shape[1], X2_train.shape[1], layers, l2, lr, outdim, use_all_singular_values, model_path)
dcca.train(X1_train, X2_train, X1_dev, X2_dev, batch_size, epochs)
dcca.model.summary()
keras.utils.plot_model(dcca.model, "data/deepcca_100.png", show_shapes=True)
dev_res = dcca.test(X1_dev, X2_dev, batch_size)
cca_embeds_test = dcca.predict(X1_test, X2_test, batch_size)
save_as_pickle(cca_embeds_test, "data/features/dcca_100.pkl.test")
cca_embeds_dev = dcca.predict(X1_dev, X2_dev, batch_size)
save_as_pickle(cca_embeds_dev, "data/features/dcca_100.pkl.dev")
cca_embeds_train = dcca.predict(X1_train, X2_train, batch_size)
save_as_pickle(cca_embeds_train, "data/features/dcca_100.pkl.train")
| deep_cca.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Eurocode 2 - Chapter 3 - Concrete tables
# +
import pandas as pd
import numpy as np
from numpy import vectorize
import matplotlib.pyplot as plt
from streng.codes.eurocodes.ec2.raw.ch3.concrete import strength, stress_strain
# -
fck = np.array([12., 16. ,20., 25., 30., 35., 35., 40., 45., 50., 55., 60., 70., 80., 90.])
# +
fcm = strength.fcm(fck)
vfctm = vectorize(strength.fctm)
vfctk005 = vectorize(strength.fctk005)
vfctk095 = vectorize(strength.fctk095)
fctm = vfctm(fck)
fctk005 = vfctk005(fck)
fctk095 = vfctk095(fck)
# -
dict_strengths = {"fck": fck, "fcm": fcm.round(2), "fctm": fctm.round(2),
"fctk005": fctk005.round(2), "fctk095": fctk095.round(2)}
strengths_table = pd.DataFrame(dict_strengths)
strengths_table
inverted_strengths_table = strengths_table.T
inverted_strengths_table
# +
fig, ax = plt.subplots()
line0, = ax.plot(fck, fck, '--', linewidth=2,
label='fck')
line1, = ax.plot(fck, fcm, '--', linewidth=2,
label='fcm')
line2, = ax.plot(fck, fctm, dashes=[30, 5, 10, 5],
label='fctm')
line3, = ax.plot(fck, fctk005, dashes=[30, 5, 10, 5],
label='fctk005')
line4, = ax.plot(fck, fctk095,
label='fctk095')
ax.legend(loc='right')
ax.grid(True)
plt.show()
# +
vεc1 = vectorize(stress_strain.εc1)
vεc2 = vectorize(stress_strain.εc2)
vεc3 = vectorize(stress_strain.εc3)
vεcu1 = vectorize(stress_strain.εcu1)
vεcu2 = vectorize(stress_strain.εcu2)
vεcu3 = vectorize(stress_strain.εcu3)
vn = vectorize(stress_strain.n)
εc1 = vεc1(fck)
εc2 = vεc2(fck)
εc3 = vεc3(fck)
εcu1 = vεcu1(fck)
εcu2 = vεcu2(fck)
εcu3 = vεcu3(fck)
n = vn(fck)
# -
dict_strains = {"fck": fck, "εc1": εc1.round(2), "εc2": εc2.round(2), "εc3": εc3.round(2),
"εcu1": εcu1.round(2), "εcu2": εcu2.round(2), "εcu3": εcu3.round(2),
"n": n.round(2)}
strains_table = pd.DataFrame(dict_strains)
strains_table
merged_table = pd.merge(left=strains_table, right=strengths_table, left_on='fck', right_on='fck')
merged_table
print(merged_table.to_markdown(index=False))
print(merged_table.to_latex(index=False))
| codes/eurocodes/ec2/raw_ch3_concrete_tables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#https://www.tensorflow.org/tutorials/keras/regression
import tensorflow as tf
from tensorflow import keras
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# +
batch = 1000
half_batch = int(batch/2)
k = 1 #number of covariates
n = 2 #number of hidden nodes
c = 2 #number of classes
example_batch = np.concatenate((np.ones((half_batch,1)),np.zeros((half_batch,1))))
example_classify = np.concatenate((np.ones((half_batch,1)),np.zeros((half_batch,1))))
#print(example_batch)
hidden_layer = tf.keras.layers.Dense(n, kernel_initializer='ones',
bias_initializer='ones', activation='sigmoid', input_shape=(k,))
linear_activation = lambda x: tf.keras.activations.relu(x, alpha=1.0)
log_odds = tf.keras.layers.Dense(c, kernel_initializer='ones',
bias_initializer='ones', activation=linear_activation, input_shape=(n,))
model = tf.keras.Sequential([hidden_layer, log_odds, tf.keras.layers.Softmax()])
scce = tf.keras.losses.SparseCategoricalCrossentropy()
pred = model.apply(example_batch) #note calculated using natural log...
print(scce(example_classify,pred))
#train model
sgd_fast = tf.keras.optimizers.SGD(learning_rate=0.5)
model.compile(loss=scce, optimizer=sgd_fast)
model.fit(example_batch,example_classify, verbose=0, epochs=10)
#predict
pred = model.apply(example_batch) #note calculated using natural log...
print(pred)
print(scce(example_classify,pred))
# +
tf.keras.backend.clear_session()
# -
| misc/iterated_log/tensor_nn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# + [markdown] tags=["meta", "toc_en", "draft_en"]
# # Unconstrained local optimization with Scipy
# -
# https://docs.scipy.org/doc/scipy/reference/optimize.html#local-optimization
#
# * minimize(method=’Nelder-Mead’)
# * minimize(method=’Powell’)
# * minimize(method=’CG’)
# * minimize(method=’BFGS’)
# * minimize(method=’Newton-CG’)
# * minimize(method=’L-BFGS-B’)
# * minimize(method=’TNC’)
# * minimize(method=’COBYLA’)
# * minimize(method=’SLSQP’)
# * minimize(method=’dogleg’)
# * minimize(method=’trust-ncg’)
#
# General-purpose multivariate methods:
# * fmin(func, x0[, args, xtol, ftol, maxiter, ...]) Minimize a function using the downhill simplex algorithm.
# * fmin_powell(func, x0[, args, xtol, ftol, ...]) Minimize a function using modified Powell’s method.
# * fmin_cg(f, x0[, fprime, args, gtol, norm, ...]) Minimize a function using a nonlinear conjugate gradient algorithm.
# * fmin_bfgs(f, x0[, fprime, args, gtol, norm, ...]) Minimize a function using the BFGS algorithm.
# * fmin_ncg(f, x0, fprime[, fhess_p, fhess, ...]) Unconstrained minimization of a function using the Newton-CG method.
#
# **Constrained** multivariate methods: => **A mettre dans un autre notebook "..._constrained_local_optimization_..." !**
# * fmin_l_bfgs_b(func, x0[, fprime, args, ...]) Minimize a function func using the L-BFGS-B algorithm.
# * fmin_tnc(func, x0[, fprime, args, ...]) Minimize a function with variables subject to bounds, using gradient information in a truncated Newton algorithm.
# * fmin_cobyla(func, x0, cons[, args, ...]) Minimize a function using the Constrained Optimization BY Linear Approximation (COBYLA) method.
# * fmin_slsqp(func, x0[, eqcons, f_eqcons, ...]) Minimize a function using Sequential Least SQuares Programming
# * differential_evolution(func, bounds[, args, ...]) Finds the global minimum of a multivariate function.
#
# Univariate (scalar) minimization methods:
# * fminbound(func, x1, x2[, args, xtol, ...]) Bounded minimization for scalar functions.
# * brent(func[, args, brack, tol, full_output, ...]) Given a function of one-variable and a possible bracketing interval, return the minimum of the function isolated to a fractional precision of tol.
# * golden(func[, args, brack, tol, ...]) Return the minimum of a function of one variable.
| nb_dev_python/python_scipy_optimize_local_optimization_en.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
'''
#DEPAI
read a layer of n polygons
Iterate through the points shp
Compute square buffer around the centroid
buffer to raster (resolution 0.5 m)
Save buffer raster, Save polygon, Save polygon as raster --extent is square buffer extent
#background, 0
assumes you have only train and test folders.
'''
import geopandas
import numpy as np
import gdal
import time
import matplotlib.pyplot as plt
import descartes
root_path = 'E:/DEPAI_project'
def gen_buffer(in_file_name, out_file_name, half_window_size):
'''
generate square buffers around centroids
'''
df_shp = geopandas.read_file(in_file_name)
selection = df_shp
for index, row in selection.iterrows():
#index=+1
pt_envelope = selection[index:index+1].centroid.buffer(half_window_size).envelope
# buffer 50 implies (2X50 = 100 x 100)
pt_envelope.to_file(out_file_name+str(index)+"_"+str(selection['class'][index])+'.shp')
#file name xx_yyyy.shp where xx is the patch number and yyyy is the label value
# +
# generate training set buffers
t0 = time.time()
#for the training set tiles
in_shp_train = root_path + '/labelled_slum_data/training/kis_select_points.shp'
out_SqBuffer_train = root_path + '/buffer_train_shp/'
gen_buffer (in_shp_train,out_SqBuffer_train,50)
t1 = time.time()
print("process complete in:",((t1-t0)/60))
# +
#generate testset buffers
t0 = time.time()
# for the testset tiles
in_shp_test = root_path + '/labelled_pop_data/testing/pop_tile_3.shp'
out_SqBuffer_test = root_path + '/buffer_test_shp/'
gen_buffer (in_shp_test,out_SqBuffer_test,50)
t1 = time.time()
print("process complete in:",((t1-t0)/60))
| Gen_Square_Buffer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Shell geometry
# ## Init symbols for *sympy*
# +
from sympy import *
from geom_util import *
from sympy.vector import CoordSys3D
import matplotlib.pyplot as plt
import sys
sys.path.append("../")
# %matplotlib inline
# %reload_ext autoreload
# %autoreload 2
# %aimport geom_util
# +
# Any tweaks that normally go in .matplotlibrc, etc., should explicitly go here
# %config InlineBackend.figure_format='retina'
plt.rcParams['figure.figsize'] = (12, 12)
plt.rc('text', usetex=True)
plt.rc('font', family='serif')
# SMALL_SIZE = 42
# MEDIUM_SIZE = 42
# BIGGER_SIZE = 42
# plt.rc('font', size=SMALL_SIZE) # controls default text sizes
# plt.rc('axes', titlesize=SMALL_SIZE) # fontsize of the axes title
# plt.rc('axes', labelsize=MEDIUM_SIZE) # fontsize of the x and y labels
# plt.rc('xtick', labelsize=SMALL_SIZE) # fontsize of the tick labels
# plt.rc('ytick', labelsize=SMALL_SIZE) # fontsize of the tick labels
# plt.rc('legend', fontsize=SMALL_SIZE) # legend fontsize
# plt.rc('figure', titlesize=BIGGER_SIZE) # fontsize of the figure title
init_printing()
# -
N = CoordSys3D('N')
alpha1, alpha2, alpha3 = symbols("alpha_1 alpha_2 alpha_3", real = True, positive=True)
# ## Cylindrical coordinates
R, L = symbols("R L", real = True, positive=True)
# +
a1 = pi / 2 + (L / 2 - alpha1)/R
x = (R + alpha3)* cos(a1)
y = alpha2
z = (R + alpha3) * sin(a1)
r = x*N.i + y*N.j + z*N.k
# -
R1=r.diff(alpha1)
R2=r.diff(alpha2)
R3=r.diff(alpha3)
R1
R2
R3
# ### Draw
# +
import plot
# %aimport plot
alpha1_x = lambdify([R, L, alpha1, alpha3], x, "numpy")
alpha3_z = lambdify([R, L, alpha1, alpha3], z, "numpy")
R_num = 1/0.8
L_num = 2
h_num = 0.1
x1_start = 0
x1_end = L_num
x3_start = -h_num/2
x3_end = h_num/2
def alpha_to_x(a1, a2, a3):
x=alpha1_x(R_num, L_num, a1, a3)
z=alpha3_z(R_num, L_num, a1, a3)
return x, 0, z
plot.plot_init_geometry_2(x1_start, x1_end, x3_start, x3_end, alpha_to_x)
# +
# %aimport plot
R3_1=R3.dot(N.i)
R3_3=R3.dot(N.k)
R3_1_x = lambdify([R, L, alpha1, alpha3], R3_1, "numpy")
R3_3_z = lambdify([R, L, alpha1, alpha3], R3_3, "numpy")
def R3_to_x(a1, a2, a3):
x=R3_1_x(R_num, L_num, a1, a3)
z=R3_3_z(R_num, L_num, a1, a3)
return x, 0, z
plot.plot_vectors(x1_start, x1_end, 0, alpha_to_x, R3_to_x)
# +
# %aimport plot
R1_1=R1.dot(N.i)
R1_3=R1.dot(N.k)
R1_1_x = lambdify([R, L, alpha1, alpha3], R1_1, "numpy")
R1_3_z = lambdify([R, L, alpha1, alpha3], R1_3, "numpy")
def R1_to_x(a1, a2, a3):
x=R1_1_x(R_num, L_num, a1, a3)
z=R1_3_z(R_num, L_num, a1, a3)
return x, 0, z
plot.plot_vectors(x1_start, x1_end, 0, alpha_to_x, R1_to_x)
# -
# ### Lame params
# +
H1 = 1+alpha3/R
H2=S(1)
H3=S(1)
H=[H1, H2, H3]
DIM=3
dH = zeros(DIM,DIM)
for i in range(DIM):
dH[i,0]=H[i].diff(alpha1)
dH[i,1]=H[i].diff(alpha2)
dH[i,2]=H[i].diff(alpha3)
dH
| py/notebooks/.ipynb_checkpoints/MatricesForPlaneShells-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.4 64-bit
# name: python374jvsc74a57bd07945e9a82d7512fbf96246d9bbc29cd2f106c1a4a9cf54c9563dadf10f2237d4
# ---
# # Python | day 4 | for loop, functions (feat. basics & if/else)
# ### Exercise 0.1 - Python Basics
#
# 1. What do we use to make a line break in a print? And for a tab stop?
#
# have a look ---> http://elclubdelautodidacta.es/wp/2012/04/python-capitulo-31-exprimiendo-la-funcion-print/
print('line_3\nline_4\n')
print('line_1\t.')
# 2. Make a converter from dollars to euros. You'll have to use input.
exchange_rate = 0.84
d_t_c = int(input('Type you $ to convert')) # Dollars to convert
print(f'{d_t_c}$ Exchange rate: {exchange_rate}')
euros = round(d_t_c * exchange_rate, 2)
print(f'{euros}€\n---------\n')
# 3. Declare two strings, one will be your first name and the other your last name. Declare your age in another variable. Print a sentence which include those variables, using ```f{}```. Wtf is that? check this out **-->** https://realpython.com/python-string-formatting/
# +
name = '<NAME>.'
surname = 'Simpson'
age = 38
print(f'{name} {surname}, {age} years old')
# -
# 4. Given the list `[4,7, -3]`, calculate its maximum, minimum and sum.
# +
a_list = [4,7, -3]
print (f'Max: {max(a_list)}')
print (f'Min: {min(a_list)}')
print (f'Sum: {sum(a_list)}')
# -
# 5. Save the number `38.38276252728` to a variable called `number_dec` and round it to 5 decimal places.
#
number_dec = 38.38276252728
rounded = round(number_dec, 5)
print(rounded)
# 6. Declare a variable `phrase` with the value `"Born to be wild"` and make it uppercase, then lowercase, divide it by spaces, and finally, replace `"wild"` with `"Geek."`
# +
phrase = "Born to be wild"
print(phrase.upper())
print(phrase.lower())
print(phrase.split())
print(phrase.replace("wild", "Geek"))
# -
# 7. Create a program where two inputs are collected, and the output of the program is a boolean which tell the user whether those inputs are the same or not.
#
# +
inp_1 = input('Input #1> ')
print(inp_1)
inp_2 = input('Input #2> ')
print(inp_2)
if inp_1 == inp_2:
boolean = True
print("\n- They're equal")
else:
boolean = False
print("\n- They're not equal")
# +
#using a function
def chekcer ():
"""
This function collects two inputs, and the output of the program is a boolean which tell the user whether those inputs are the same or not
"""
inp_1 = input('Input #1> ')
print(inp_1)
inp_2 = input('Input #2> ')
print(inp_2)
if inp_1 == inp_2:
boolean = True
print("\n- They're equal")
else:
boolean = False
print("\n- They're not equal")
return boolean
boolean_returned = chekcer()
print(f'{boolean_returned} <- Value of the returned variable')
print(f'{type(boolean_returned)} <- Type of the returned variable')
# -
# ### Exercise 0.2 - If/else
#
# 1. Create a decission tree using if/else sentences, to determine the price of the movies ticket. If the client's age is between 5 and 15 years, both included, the price will be 5, if she/he is retired and the movie is one of the `peliculas_discount`, the price is 4. In any other case, it will be 7 euros.
#
# You should create the list of `peliculas_discount` with your favourites movies.
# +
age = int(input('Age of client> '))
movie = input('Movie>')
movies_discount = ['The Thing', 'Star Trek VI', 'Lord Of The Rings', 'Aquaman']
if age >= 5 and age <= 15:
price = 5
elif age >= 67 and movie in movies_discount:
price = 4
else:
price = 7
print(f'Age: {age} Movie: {movie}')
print (f'Price of ticket is {price}€')
# -
# ### Exercise 1 - For
# 1. You are witnessing an epic battle between two powerful sorcerers: Gandalf and Saruman. Each sorcerer has 10 spells of variable power in their mind and they are going to throw them one after the other. The winner of the duel will be the one who wins more of those clashes between spells. Spells are represented as a list of 10 integers whose value equals the power of the spell.
#
# ```python
# gandalf = [10, 11, 13, 30, 22, 11, 10, 33, 22, 22]
# saruman = [23, 66, 12, 43, 12, 10, 44, 23, 12, 17]
# ```
#
# For example:
#
# The first clash is won by Saruman: 10 against 23, wins 23
# The second clash wins Saruman: 11 against 66, wins 66
# etc.
# You will create two variables, one for each sorcerer, where the sum of clashes won will be stored. Depending on which variable is greater at the end of the duel, you will show one of the following three results on the screen:
# ```
# Gandalf wins
# Saruman wins
# Tie
# ```
# +
# Assign spell power lists to variables
gandalf = [10, 11, 13, 30, 22, 11, 10, 33, 22, 22]
saruman = [23, 66, 12, 43, 12, 10, 44, 23, 12, 17]
# Assign 0 to each variable that stores the victories
gandalf_victories = 0
saruman_victories = 0
# Execution of spell clashes
for gandalf_spell, saruman_spell in zip(gandalf, saruman):
if gandalf_spell > saruman_spell:
gandalf_victories += 1
print(f'Gandalf value is {gandalf_spell}. Saruman value is {saruman_spell}. \nGandalf wins')
elif gandalf_spell < saruman_spell:
saruman_victories += 1
print(f'Gandalf value is {gandalf_spell}. Saruman value is {saruman_spell}. \nSaruman wins')
else:
print(f"Gandalf value is {gandalf_spell}. Saruman value is {saruman_spell}. \nIt's a tie!")
print(f'Total Gandalf victories are {gandalf_victories}. \nTotal Saruman victories are {saruman_victories}')
# -
# ### Exercise 2 - Functions
# 1. Write a function program to sum up two given different numbers.
def adder (num_1, num_2):
"""
This function sums two different numbers, prints the result and returns it for further use.
"""
num_3 = num_1 + num_2
print(f'Result: {num_3}')
return num_3
# Call the function you've just defined to sum up:
# - 5 and 6
# - -3 and 10
# - 99 and 789
adder(num_1 = 5, num_2 = 6)
adder(num_1 = -3, num_2 = 10)
adder(num_1 = 99, num_2 = 789)
# ### Bonus Track
# 2. Write a Python program to convert a tuple of characters into a string.
#
# - Sample tuple: `("T", "H", "E", " ", "B", "R", "I", "D", "G", "E")`
# - Expected output : `"THE BRIDGE"`
#
# Hint: Turn it into a list.
# +
tup = ("T", "H", "E", " ", "B", "R", "I", "D", "G", "E")
def tup_converter(tupl):
"""
Program to convert a tuple of characters into a string
"""
result = ''.join(tupl)
return result
# -
tup_converter(tup)
# 3. Write a Python program to generate a list where the values are square of numbers between 1 and 30 (both included), and return a list of first and last 5 elements of the list.
# +
def strange_square ():
"""
Write a Python program to generate a list where the values are square of numbers between 1 and 30 (both included), and return a list of first and last 5 elements of the list.
"""
number = 1
number_list = []
for n in range(30):
num_r = number ** 2
number_list.append(num_r)
number += 1
return number_list[:5] + number_list[-5:]
the_list = strange_square()
the_list
# -
# 4. Create a program that calculates the average.
#
# The program will ask the user to specify how many numbers you want to enter, and will show a prompt to include number by number to a list, and then calculate the average of each number in that list.
# +
def average_calculator ():
choice = int(input('How many numbers will you want?: '))
a_list = []
for n in range(choice):
numbers = int(input('Add your number: '))
a_list.append(numbers)
average = int(sum(a_list))/int(len(a_list))
return average
average = average_calculator()
print (average)
# -
# # If you made it till here you did a fantastic work!
# 
| week1_precurse_python_I/day4_python_III/Practice_4_For_functions_RES.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # In-Class Coding Lab: Data Visualization
#
# The goals of this lab are to help you understand:
#
# - The value of visualization: A picture is worth 1,000 words!
# - The various ways to visualize information
# - The basic requirements for any visualization
# - How to plot complex visualizations such as multi-series charts and maps
# - Visualization Tools:
# - Matplolib
# - Plot.ly
# - Folium Maps
#
# +
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
import plotly
import plotly.plotly as py
import plotly.graph_objs as go
import cufflinks as cf
import pandas as pd
import folium
import warnings
#matplotlib.rcParams['figure.figsize'] = (20.0, 10.0) # larger figure size
warnings.filterwarnings('ignore')
# -
# ## Back to the movie goers data set
#
# For this lab, we will once again use the movie goers dataset. As you may recall this data set is a survey demographic survey of people who go to the movies. Let's reload the data and setup our `age_group` feature again.
# +
goers = pd.read_csv('CCL-moviegoers.csv')
goers['age_group'] = ''
goers['age_group'][goers['age'] <=18] = 'Youth'
goers['age_group'][(goers['age'] >=19) & (goers['age'] <=55)] = 'Adult'
goers['age_group'][goers['age'] >=56] = 'Senior'
goers.sample(5)
# -
# ## Visualizing Data
#
# There are many ways your can visualize information. Which one is the most appropriate? It depends on the data, of course.
#
# - **Counting Categorial data** belongs in *charts like pie charts and bar charts*.
# - **Counting Numerical data** is best suited for *histograms*.
# - **Timeseries data and continuous data** belongs in *line charts*.
# - **A comparision of two continuous values** is best suited for a *scatter plot*.
# - **Geographical data** is best displauyed on *maps*.
#
# Let's use this knowledge to plot some data in the `goers` `DataFrame`!
#
# ## Males or Females?
#
# The first think we might want to visualize is a count gender in the dataset. A **pie** chart is well suited for this task as it displays data as a portion of a whole. To create a pie chart we need the data to count and the labels for the counts.
#
# Let's try it.
#
# First we get the value counts as a series `gender`:
gender = goers['gender'].value_counts()
gender
# Then we make it into a dataframe:
gender_df = pd.DataFrame( { 'Gender' : gender.index, "Counts" : gender })
gender_df
# Then we plot! The index has the labels, and the value at the index is what we want to plot:
gender_df.plot.pie( y = 'Counts') # y are the values we are plotting
# ### Now You Try it!
#
# Create a pie chart based on `age_group` first create a series of the `value_counts()` second, create the `DataFrame` with two columns `AgeGroup` and `Counts` then plot with `.plot.pie()`.
#
# Follow the steps we did in the previous three cells, but comvine into one cell!
#todo write code here
age_group = goers['age_group'].value_counts()
age_group_df = pd.DataFrame( { 'AgeGroup' : age_group.index, "Counts" : age_group })
age_group_df.plot.pie( y = 'Counts')
# ## Too many pieces of the pie?
#
# Pie charts are nice, but they are only useful when you have a small number of labels. More that 5-7 labels and the pie becomes messy. For example take a look at this pie chart of `occupation`:
occ = goers['occupation'].value_counts()
occ_df = pd.DataFrame( { 'occupation' : occ.index, "counts" : occ })
occ_df.plot.pie(y = 'counts')
# That's crazy... and difficult to comprehend. Also pie charts visualize data as part of the whole. We have no idea how many students there are. Sometimes we want to know actual counts. This is where the **bar chart** comes in handy!
#
# ## Raising the bar!
#
# Let's reproduce the same plot as a bar:
occ_df.plot.bar()
# Ahh. that's much better. So much easier to understand!
#
# ### Now you try it!
#
# Write a one-liner to plot `groups_df` as a Bar!
# todo write code here
age_group_df.plot.bar()
# ## When bar charts fail...
#
# Bar charts have the same problem as pie charts. Too many categories overcomplicate the chart, or show the data in a meaningless way. For example, let's create a bart chart for ages:
ages = goers['age'].value_counts()
ages_df = pd.DataFrame( { 'age' : ages.index, "counts" : ages })
ages_df.plot.bar(y = 'counts')
# Meaningless. For two key reasons:
#
# 1. too many categories
# 2. age is a continuous variable not a categorical variable. In plain English, this means there's a relationship between one age and the next. 20 < 21 < 22. This is not represented in a bar chart.
#
# ## ...Call in the Histogram!
#
# What we want is a **historgram**, which takes a continuous variable and loads counts into "buckets". Notice how we didn't have to lump data with `value_counts()`. Histograms can do that automatically because the `age` variable is continuous. Let's try it:
goers.hist(column ='age')
# ## Plot.ly
#
# [Plot.ly](https://plot.ly) is data visualization as a service. You give it data, it gives you back a web-based plot. Plot.ly is free and works with a variety of environments and programming languages, including Python.
#
# For Python is has bindings so that you can use it just like `matplotlib`! No need to manually invoke the web service call.
#
# To get started with plot.ly you must sign up for an account and get a set of credentials:
#
# - Visit [https://plot.ly/settings/api](https://plot.ly/settings/api)
# - Create an account or sign-in with Google or GitHub
# - Generate your API key and paste your username and key in the code below:
# todo: setup the credentials replace ??? and ??? with your Plot.ly username and api_key
plotly.tools.set_credentials_file(username='hvpuzzan-su', api_key='<KEY>')
# Using plot.ly is as easy as, or sometimes easier than `matplotlib`. In most cases all you need to do is call `iplot()` on the data frame. For example, here's out first pie chart, plotly style:
gender_df.iplot(kind="pie", labels = 'Gender', values='Counts')
# Notice that plot.ly is a bit more interactive. You can hover over the part of the pie chart and see counts!
#
# ### Now You Try it!
#
# Use plotly's `iplot()` method to create a bar chart on the `occ_df` Data Frame:
#
# todo: write code here
occ_df.iplot(kind="pie", labels = 'occupation', values='counts')
# ## Folium with Leaflet.js
#
# Folium is a Python module wrapper for [Leaflet.js](http://leafletjs.com/), which uses [Open Street Maps](https://www.openstreetmap.us/). These are two, popular open source mapping libraries. Unlike Google maps API, its 100% free!
#
# You can use Folium to render maps in Python and put data on the maps. Here's how easy it is to bring up a map:
#
CENTER_US = (39.8333333,-98.585522)
london = (51.5074, -0.1278)
map = folium.Map(location=CENTER_US, zoom_start=4)
map
# You can zoom right down to the street level and get a amazing detail. There's also different maps you can use, as was covered in this week's reading.
#
# ## Mapping the students.
#
# Let's take the largest category of movie goers and map their whereabouts. We will first need to import a data set to give us a lat/lng for the `zip_code` we have in the dataframe. We could look this up with Google's geolookup API, but that's too slow as we will be making 100's of requests. It's better to have them stored already and merge them with `goers`!
#
# Let's import the zipcode database into a Pandas DataFrame, then merge it with the `goers` DataFrame:
zipcodes = pd.read_csv('https://raw.githubusercontent.com/mafudge/datasets/master/zipcodes/free-zipcode-database-Primary.csv', dtype = {'Zipcode' :object})
data = goers.merge(zipcodes, how ='inner', left_on='zip_code', right_on='Zipcode')
students = data[ data['occupation'] == 'student']
students.sample()
# Let's explain the code, as a Pandas refresher course:
#
# 1. in the first line I added `dtype = {'Zipcode' :object}` to force the `Zipcode` column to be of type `object` without that, it imports as type `int` and cannot match with the `goers` DataFrame.
# 1. the next line merges the two dataframes together where the `zip_code` in `doers` (on_left) matches `Zipcode` in `zipcodes` (on_right)
# 1. the result `data` is a combined DataFrame, which we then filter to only `student` occupations, sorting that in the `students` DataFrame
#
#
# ## Slapping those students on a map!
#
# We're ready to place the students on a map. It's easy:
#
# 1. For each row in the students dataframe:
# 1. get the coordinates (lat /lng )
# 1. make a `marker` with the coordinates
# 1. add the marker to the map with `add_children()`
#
# Here we go!
#
for row in students.to_records():
pos = (row['Lat'],row['Long'])
message = "%d year old %s from %s, %s" % (row['age'],row['gender'], row['City'], row['State'])
marker = folium.Marker(location=pos,
popup=message
)
map.add_children(marker)
map
# ### Now you try it!
#
#
# 1. use the `data` DataFrame to retrieve only the occupation `programmer`
# 1. create another map `map2` plot the programmers on that map!
## todo write code here!
zipcodes = pd.read_csv('https://raw.githubusercontent.com/mafudge/datasets/master/zipcodes/free-zipcode-database-Primary.csv', dtype = {'Zipcode' :object})
data = goers.merge(zipcodes, how ='inner', left_on='zip_code', right_on='Zipcode')
programmer = data[ data['occupation'] == 'programmer']
programmer.sample()
for row in programmer.to_records():
pos = (row['Lat'],row['Long'])
message = "%d year old %s from %s, %s" % (row['age'],row['gender'], row['City'], row['State'])
marker = folium.Marker(location=pos,
popup=message
)
map.add_children(marker)
map
| content/lessons/13/Class-Coding-Lab/CCL-Data-Visualization.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Scala
// language: scala
// name: scala
// ---
// +
import $ivy.`org.proml::proml-core:0.1.0`
import proml._
import $ivy.`com.cibo::evilplot-jupyter-scala:0.6.1-SNAPSHOT`
import $ivy.`com.cibo::evilplot-repl:0.6.1-SNAPSHOT`
import com.cibo.evilplot._
import com.cibo.evilplot.plot._
import com.cibo.evilplot.plot.aesthetics.DefaultTheme._
import com.cibo.evilplot.numeric.Point
import almond.interpreter.api._
import almond.api.helpers.Display.Image
import proml.Model
import proml.distributions.Continuous._
import proml.samplers.MetropolisHastings
import com.cibo.evilplot.geometry.Drawable
val trueA = 2d
val trueB = -2d
val points: Seq[(Double, Double)] = for {
x <- -10d to 10d by 0.1
noise = normal[Double](0, 1).get
} yield (x * 1.0, trueA * x + trueB + noise)
// +
def plot(data: Seq[(Double, Double)]) = {
ScatterPlot(
data.map{p => Point(p._1, p._2)}
).frame()
.xLabel("x")
.yLabel("y")
.frame()
.xbounds(data.map{_._1}.min, data.map{_._1}.max)
.ybounds(data.map{_._2}.min, data.map{_._2}.max)
.xAxis()
.yAxis()
.render()
}
def show(drawable: Drawable) {
import java.io.ByteArrayOutputStream
import javax.imageio.ImageIO
val baos = new ByteArrayOutputStream()
ImageIO.write(drawable.asBufferedImage, "png", baos)
try{Image.fromArray(baos.toByteArray(), Image.PNG)}
catch{
case e: Exception =>
}
}
show(plot(points))
// + active=""
//
// Bayes Theorm:
// =============
//
// posterior Likelihood Prior
// P(θ|x) = P(x|θ) * P(θ)
// ----------------------
// p(x)
// We cant compute this
//
// MCMC Theorm:
// ============
// lets say μ0 is our current position, and μ is our proposal
//
//
// P(x|μ)P(μ) P(x|μ0)P(μ0) P(x|μ) * P(μ)
// ---------- / ---------- = ----------------
// P(x) P(x) P(x|μ0) * P(μ0)
//
//
// https://chi-feng.github.io/mcmc-demo/app.html#RandomWalkMH,banana
// -
case class Params(amu: Double, bmu: Double)
// Blindly assuming, 100 is the slope and -10 is the y-intercept
val priorAssumption = Params(100, -10)
def getLogLikelihood(p: Params): Double = {
normal[Double](priorAssumption.amu, 1).logPdf(p.amu) +
normal[Double](priorAssumption.bmu, 1).logPdf(p.bmu)
}
def getProposal(prior: Params) =
for {
a <- normal[Double](0, 1)
b <- normal[Double](0, 1)
} yield Params(a + prior.amu, b + prior.bmu)
val linearModel = new Model[Params, Double, Double] {
// MCMC proposal to do random walk
override def proposal = (p: Params) => getProposal(p).get
// P(x|θ)
override def logPrior = getLogLikelihood
// P(θ)
override def prior = priorAssumption
// y = ax + b
override def model = (p: Params) => (x: Double) => x * p.amu + p.bmu
}
val (best, samples) = linearModel.fit(
points,
MetropolisHastings(20000, 15000) // MCMC Sampler
)
println(best)
// +
def cplot(data: Seq[(Double, Double)]) = {
ContourPlot(
data.map{p => Point(p._1, p._2)}
).frame()
.xLabel("Slope")
.yLabel("Y-Intercept")
.xbounds(data.map{_._1}.min, data.map{_._1}.max)
.ybounds(data.map{_._2}.min, data.map{_._2}.max)
.xAxis()
.yAxis()
.topPlot(Histogram(data.map{_._1}), 100)
.rightPlot(Histogram(data.map{_._2}), 100)
.render()
}
show(cplot(samples.map(s => (s.amu, s.bmu))))
| notebooks/Linear Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# Fill in any place that says `# YOUR CODE HERE` or YOUR ANSWER HERE, as well as your name and collaborators below.
# Grading for pre-lecture assignments is all or nothing. Partial credit is available for in-class assignments and checkpoints, but **only when code is commented**.
# -
NAME = ""
COLLABORATORS = ""
# ---
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "81de796416747709f44457f5bd0acb9d", "grade": false, "grade_id": "cell-0ace592735c97afd", "locked": true, "schema_version": 3, "solution": false}
# # Learning Objectives
#
# This lecture will show you how to:
# 1. Create sliders and other widgets
# 2. Customize the appearance and behavior of widgets
# 3. Use container widgets to build a Graphical User Interface (GUI)
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "59a1a634c252595291e4c7bd376c4506", "grade": false, "grade_id": "cell-abd1b2cca923116d", "locked": true, "schema_version": 3, "solution": false}
# imports
import numpy as np
import matplotlib.pyplot as plt
import ipywidgets as widgets # widgets that run when the notebook is live
from IPython import display # display.display() is used to draw the widgets
import grading_helper as _test
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "a81ffe38c06f24833123458a5b213cb0", "grade": false, "grade_id": "cell-e2542f1e037a7a13", "locked": true, "schema_version": 3, "solution": false}
# # Simple Widgets
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "83ddb8e877dc9192aae3b0ecf7b010a2", "grade": false, "grade_id": "cell-3e1adb3781236df0", "locked": true, "schema_version": 3, "solution": false}
# %video TFfF3ulSIuU
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "aeae8fd53b1689d4adc1c33c9314b822", "grade": false, "grade_id": "cell-e2542f1e037a7a14", "locked": true, "schema_version": 3, "solution": false}
# Summary:
# - Use `widgets.interactive` to connect one or more widgets (usually sliders) to each argument of a function.
# - Call `display` to show the widget and its output. You can do this in a different cell.
# -
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "ac88a5829d98271d4c4444c64bb25b62", "grade": false, "grade_id": "cell-4d5dd8920080e674", "locked": true, "schema_version": 3, "solution": false}
# ## Your Turn
#
# Use `interactive` to connect a `IntSlider` to the function `powers_of_two` below. Name the interactive object `gui` (like in the video), and show the widget using `display.display`.
#
# > **Important:** For autograding to work (in any assignment), you must display widgets with an explicit call to `display.display`.
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "37102d6345e82e68246ce66f4a2e2d6a", "grade": false, "grade_id": "cell-6f986a9c00e72b61", "locked": true, "schema_version": 3, "solution": false}
def powers_of_two(n):
"""Print the nth power of 2"""
print(f"2^{n} = {2**n}")
# + deletable=false nbgrader={"cell_type": "code", "checksum": "63dd1781e1ce3e67aac74d6face61d5b", "grade": false, "grade_id": "cell-4b9e392447b94fb5", "locked": false, "schema_version": 3, "solution": true}
# %%graded # 2 points
# YOUR CODE HERE
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "ec909df4e572fc18ccfca7a5e3eb1419", "grade": true, "grade_id": "cell-d6eed339bfcc55cb", "locked": true, "points": 2, "schema_version": 3, "solution": false}
# %%tests
_test.code_contains("interactive", "powers_of_two", "display.display", "gui")
_test.widget_snapshot() # print a text description of the widget that will survive when the notebook isn't live
_test.widget_has("IntSlider", "Output") # check that these widgets exist
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "ed79545c5917da2cadb4f0649c2e2f76", "grade": false, "grade_id": "cell-95238e3a02ff852d", "locked": true, "schema_version": 3, "solution": false}
# # Customizing Widgets
# -
# %video judT3RO8MYs
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "6ee8eef1b1feba3ef660e22718ccf447", "grade": false, "grade_id": "cell-5a29017df3fbd834", "locked": true, "schema_version": 3, "solution": false}
# Summary:
# - Sliders can be customized with a `min`, `max`, `step`, and starting `value`
# - You can label widgets by setting the `description` argument.
# - Update lag can avoid that by adding the argument `continuous_update=False`
#
# See https://ipywidgets.readthedocs.io/en/stable/examples/Widget%20List.html for a list of widgets and their available arguments.
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "8b7bbfbead0528c864ba05562fb5da3b", "grade": false, "grade_id": "cell-3cbc586b7dc272ad", "locked": true, "schema_version": 3, "solution": false}
# ## Your Turn
#
# Make a graph of $\sin(nx)$ for $-10\le x\le 10$, where $n$ is a floating point number between 1 and 5 in steps 0.2 set by a `FloatSlider`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "7f839104ac3b3bf2e647532a26773946", "grade": false, "grade_id": "cell-ba4307ca428933fa", "locked": false, "schema_version": 3, "solution": true}
# %%graded # 1 point
x = np.linspace(-10, 10, 200)
def graph(n):
plt.plot(x, np.sin(n*x))
plt.show()
# create your slider here
# YOUR CODE HERE
sin_gui = widgets.interactive(graph, n=s)
display.display(sin_gui)
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "1446d6d6529f5deaf4404930023260d7", "grade": true, "grade_id": "cell-4d643dc04b39e690", "locked": true, "points": 1, "schema_version": 3, "solution": false}
# %%tests
_test.code_contains("display")
_test.widget_snapshot()
_test.widget_has("FloatSlider", "max=5.0", "min=1.0", "step=0.2")
_test.plot_shown()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "f9a5cbe1d295f150a2cd3c8d4ce5e270", "grade": false, "grade_id": "cell-830305f99033ea1c", "locked": true, "schema_version": 3, "solution": false}
# # Container Widgets
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "e18638dbbf459fee6e57b6eebad8d171", "grade": false, "grade_id": "cell-fd2146e4d1348db7", "locked": true, "schema_version": 3, "solution": false}
# %video QhdyzIufMMM
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "f91df3886c102ba9367d121edac2876e", "grade": false, "grade_id": "cell-8e502d4b46ab82d3", "locked": true, "schema_version": 3, "solution": false}
# Summary:
# - You can put widgets inside special layout widgets to control where they are drawn.
# - The basic building blocks are `widgets.HBox` and `widgets.VBox`
# - Output can be captured as a widget using `widgets.interactive_output` in place of `widgets.interactive`
# -
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "e31f27c656494573cc61589bd3f2bf6a", "grade": false, "grade_id": "cell-9b12f67c7faac919", "locked": true, "schema_version": 3, "solution": false}
# ## Your Turn
#
# Use `HBox` and `VBox` and the code below to create the following layout:
#
# <img src=attachment:image.png width=500>
# + deletable=false nbgrader={"cell_type": "code", "checksum": "823a7ef26b795c11d0038d5b714f495b", "grade": false, "grade_id": "cell-0739970a63afc78b", "locked": false, "schema_version": 3, "solution": true}
# %%graded # 2 points
t = np.linspace(0, 2*np.pi, 200)
def parametric_plot(x, y):
plt.plot(np.cos(x*t), np.sin(y*t))
plt.axis("scaled")
plt.show()
x_slider = widgets.IntSlider(min=1, max=10, description="x")
y_slider = widgets.IntSlider(min=1, max=10, description="y")
arg_dict = dict(x=x_slider, y=y_slider)
out = widgets.interactive_output(parametric_plot, arg_dict)
# Use HBox and VBox to place the sliders and plot
# don't forget to display the final result
# YOUR CODE HERE
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "b535034eaee2e9fbc6ed7c12a6c0841f", "grade": true, "grade_id": "cell-36c07d4a6b42a0ec", "locked": true, "points": 2, "schema_version": 3, "solution": false}
# %%tests
_test.code_contains("HBox", "VBox", "display")
_test.widget_snapshot()
_test.widget_has("HBox", "VBox", "IntSlider", "Output")
_test.plot_shown()
# -
# # Additional Resources
#
# - Official documentation for `ipywidgets` https://ipywidgets.readthedocs.io/en/stable/index.html
# - Check out [Voila](https://voila.readthedocs.io/en/latest/index.html). It's a new project that lets you turn notebooks into web applications.
| Assignments/05.1 Lecture - Widgets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# %matplotlib notebook
import matplotlib.pyplot as plt
import numpy as np
import numpy.ma as ma
import pandas as pd
import netCDF4 as nc
import os
import scipy.io as sio
import scipy
from scipy import interpolate, signal
from pyproj import Proj,transform
import sys
sys.path.append('/ocean/ssahu/CANYONS/wcvi/grid/')
from bathy_common import *
from matplotlib import path
from salishsea_tools import viz_tools
from netCDF4 import Dataset
import xarray as xr
from salishsea_tools import nc_tools
import scipy.io as sio
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.cm as cm
import cmocean as cmo
import matplotlib.gridspec as gridspec
# %matplotlib inline
from scipy.io import loadmat
from scipy.interpolate import griddata
from dateutil.parser import parse
from salishsea_tools import geo_tools, viz_tools, tidetools, nc_tools
import gsw
import sklearn.cluster as cluster
from sklearn.mixture import GMM
from scipy import interpolate
import seaborn as sns
sns.set_context('poster')
sns.set_color_codes()
plot_kwds = {'alpha' : 0.25, 's' : 80, 'linewidths':0}
# +
zlevels = nc.Dataset('/data/mdunphy/NEP036-N30-OUT/CDF_COMB_COMPRESSED/NEP036-N30_IN_20140915_00001440_grid_T.nc').variables['deptht'][:32]
y_wcvi_slice = np.arange(180,350)
x_wcvi_slice = np.arange(480,650)
# bathy = nc.Dataset('/data/mdunphy/NEP036-N30-OUT/INV/Bathymetry_EastCoast_NEMO_R036_GEBCO_corr_v14.nc')
# Z = bathy.variables['Bathymetry'][:]
# lon = bathy['nav_lon'][...]
# lat = bathy['nav_lat'][...]
bathy = nc.Dataset('/data/ssahu/WCVI_sliced_bathy_NEP36_original.nc')
Z = bathy.variables['Bathymetry'][:]
# y_wcvi_slice = np.array(np.arange(180,350))
# x_wcvi_slice = np.array(np.arange(480,650))
# z_wcvi = Z[y_wcvi_slice, x_wcvi_slice]
lon_wcvi = bathy['nav_lon'][:]#[180:350, 480:650]
lat_wcvi = bathy['nav_lat'][:]#[180:350, 480:650]
z0 = np.ma.masked_values(Z, 0)
# +
NEP = nc.Dataset('/data/ssahu/NEP36_Extracted_Months/NEP36_2013_T_S_Spice_larger_offshore_rho_correct.nc')
sal = NEP.variables['vosaline']
temp = NEP.variables['votemper']
spic = NEP.variables['spiciness']
rho = NEP.variables['density']
sal_mean_august = np.mean(sal[88:119,...], axis=0)
temp_mean_august = np.mean(sal[88:119,...], axis=0)
spic_mean_august = np.mean(spic[88:119,...], axis=0)
rho_mean_august = np.mean(rho[88:119,...], axis=0)
# +
survey_mat_file_A = '/data/ssahu/Falkor_2013/mvp/surveyA.mat'
survey_mat_file_B = '/data/ssahu/Falkor_2013/mvp/surveyB.mat'
survey_mat_file_D = '/data/ssahu/Falkor_2013/mvp/surveyD.mat'
survey_mat_file_E = '/data/ssahu/Falkor_2013/mvp/surveyE.mat'
survey_mat_file_G = '/data/ssahu/Falkor_2013/mvp/surveyG.mat'
all_surveys = [survey_mat_file_A, survey_mat_file_B, survey_mat_file_D, \
survey_mat_file_E, survey_mat_file_G]
lon_falkor = []
lat_falkor = []
for survey_file in all_surveys:
mat = scipy.io.loadmat(survey_file)
lat_survey = mat['latitude'][:,0]
lon_survey = mat['longitude'][:,0] - 100
lon_falkor = np.append(arr=lon_falkor, values=lon_survey)
lat_falkor = np.append(arr=lat_falkor, values=lat_survey)
# +
import matplotlib as mpl
# y = np.empty_like(lat_location)
# x = np.empty_like(lat_location)
cmap = plt.cm.get_cmap('icefire_r', 20)
# cmap = cmo.cm.turbid
# cmap = plt.cm.jet
# cmaplist = [cmap(i) for i in range(cmap.N)]
# # force the first color entry to be grey
# cmaplist[0] = (.8, .8, .8, 1.0)
# # create the new map
# cmap = mpl.colors.LinearSegmentedColormap.from_list(
# 'Custom cmap', cmaplist, cmap.N)
# cmap.set_bad('burlywood')
cmap.set_bad('#8b7765')
fig, ax = plt.subplots(1, 1, figsize=(12,10)); ax.grid()
ax.set_aspect(viz_tools.set_aspect(ax, coords='map', lats=lat_wcvi, adjustable='box-forced'))
# ax.relim()
# ax.autoscale_view(True,True,True)
# ax.autoscale_view(scalex=True)
# ax.autoscale(enable=True, axis='both', tight=None)
# bathy_tools.plot_colourmesh(bathy, title = 'WCVI Subset', fig_size=(9, 9), axis_limits=None, colour_map='winter_r', bins=15, land_colour='burlywood')
# p = ax.pcolormesh(x_wcvi_slice,y_wcvi_slice,Z[y_wcvi_slice,x_wcvi_slice], cmap=cmap, vmin=0, vmax=500)
p = ax.pcolormesh(lon_wcvi,lat_wcvi,Z, cmap=cmap, vmin=0, vmax=1000)
cbar = fig.colorbar(p, ax=ax)
ax.set_aspect('auto')
CS1 = ax.contour(lon_wcvi, lat_wcvi, Z, np.arange(100,110,10))
CLS1 = plt.clabel(CS1, inline=3,fmt='%0.0f m', fontsize=12)
CS1 = ax.contour(lon_wcvi,lat_wcvi,Z, np.arange(200,210,10))
CLS1 = plt.clabel(CS1, inline=3,fmt='%0.0f m', fontsize=12)
# CS1 = ax.contour(lon_wcvi,lat_wcvi,Z, np.arange(1000,1010,10))
# CLS1 = plt.clabel(CS1, inline=3,fmt='%0.0f m', fontsize=12)
# CS1 = ax.contour(lon_wcvi,lat_wcvi,Z, np.arange(500,510,10))
# CLS1 = plt.clabel(CS1, inline=3,fmt='%0.0f m', fontsize=12)
# aspect = 20
# pad_fraction = 0.001
# divider = make_axes_locatable(ax)
# width = axes_size.AxesY(ax, aspect=1./aspect)
# pad = axes_size.Fraction(pad_fraction, width)
# cax = divider.append_axes("right", size=width, pad=pad)
# cbar = fig.colorbar(p, cax=cax)
# for i in np.arange(lat_location.shape[0]):
# y[i], x[i] = geo_tools.find_closest_model_point(
# lon_location[i],lat_location[i],lon_wcvi,lat_wcvi,tols={
# 'NEMO': {'tol_lon': 0.1, 'tol_lat': 0.1},'GEM2.5': {'tol_lon': 0.1, 'tol_lat': 0.1}})
# n = np.empty_like(lat_location)
# m = np.empty_like(lat_location)
# for i in np.arange(lat_location.shape[0]):
# n[i], m[i] = geo_tools.find_closest_model_point(
# lon_location[i],lat_location[i],lon_wcvi,lat_wcvi,tols={
# 'NEMO': {'tol_lon': 0.1, 'tol_lat': 0.1},'GEM2.5': {'tol_lon': 0.1, 'tol_lat': 0.1}})
# y = y.astype(int)
# x = x.astype(int)
# n = n.astype(int)
# m = m.astype(int)
# ax.scatter(x[:], y[:], c='orangered', s=100, alpha=0.5, label = 'Falkor CTD locations')
ax.set_xlabel('Longitude', fontsize=16)
ax.set_ylabel('Latitude', fontsize=16)
# ax.set_title('WCVI slice of NEP036 Model Domain', fontsize=20)
lon_W02 = -124.89500
lat_W02 = 48.04167
lon_W01 = -124.82500
lat_W01 = 48.31500
lon_swift = -125.00
lat_swift = 48.55
ax.scatter(lon_swift, lat_swift, c='k', marker='*', s=50, alpha=0.8)#, label = 'Swiftsure Bank')
# S = ("SwB")
S = ("Swiftsure\nBank")
ax.text(lon_swift, lat_swift, S, fontsize=11, color = 'k', fontweight = 'heavy',family='serif', style='italic', ha='center', rotation = 10,
va='top', wrap=False)
# ax.text(lon_swift, lat_swift, S, fontsize=11, color = 'k', fontweight = 'heavy',family='serif', style='italic', ha='left', rotation = 10,
# va='bottom', wrap=False)
lat_cape = 48.3831
lon_cape = -124.7144
ax.scatter(lon_cape, lat_cape, c='k', marker='*', s=50, alpha=0.8)#, label = 'Cape Flattery')
CP = ("Cape\nFlattery")
ax.text(lon_cape, lat_cape, CP, fontsize=11, color = 'k', fontweight = 'heavy',family='serif', style='italic', ha='center', rotation = 10,
va='top', wrap=False)
lat_jdfs = 48.5
lon_jdfs = -124.4
JD = ("JdF\nStrait")
ax.text(lon_jdfs, lat_jdfs, JD, fontsize=11, color = 'white', fontweight = 'heavy',family='serif', style='italic', ha='center', rotation = -35,
va='top', wrap=False)
lat_EP = 49.3835
lon_EP = -126.5447
ax.scatter(lon_EP, lat_EP, c='k', marker='*', s=50, alpha=0.8)
EP = ("Estevan\nPoint")
ax.text(lon_EP, lat_EP, EP, fontsize=11, color = 'k', fontweight = 'heavy',family='serif', style='italic', ha='left', rotation = 0,
va='top', wrap=False)
lon_LB08 = -125.4775
lat_LB08 = 48.4217
ax.scatter(lon_falkor[::3], lat_falkor[::3], c='yellow', marker='o', s=10, alpha=0.9, label = 'Pathways MVP')
ax.scatter(lon_glider[::50], lat_glider[::50], c='magenta', marker='o', s=10, alpha=0.3, label = 'CMOP Glider')
ax.scatter(lon_LB08, lat_LB08, c='indigo', marker='*', s=150, linewidths=1.2, alpha=1, label = 'LB08')
lon_A1 = -126.20433
lat_A1 = 48.52958
ax.scatter(lon_A1, lat_A1, c='cyan',marker='*', s=150, alpha=0.8, label = 'A1 Mooring')
# ax.scatter(lon_W01, lat_W01, c='cyan', s=150, alpha=0.8, label = 'W01')
# ax.scatter(lon_W02, lat_W02, c='red', s=150, alpha=0.8, label = 'W02')
# ax.scatter(lon_location[:], lat_location[:], c='purple', s=100, alpha=0.3, label = 'Falkor CTD locations')
legend = ax.legend(loc='lower left', fancybox=True, framealpha=0.85)
# ax.set_xlabel('x_index')
# ax.set_ylabel('y_index')
# ax.set_title('WCVI slice of NEP036 Model Domain', fontsize=20)
# viz_tools.plot_land_mask(ax, bathy)#, yslice=y_wcvi_slice, xslice=x_wcvi_slice, color='burlywood')
# cbar = fig.colorbar(p, cax=ax)
cbar.ax.set_ylabel('Depth [m]', fontsize=16)
ax.tick_params(axis='both',labelsize =16)
cbar.ax.tick_params(labelsize=16)
# t = ("<NAME>")
# ax.text(lon_W01, lat_W01, t, fontsize=11, color = 'darkblue', family='serif', style='italic', ha='left', rotation = 55,
# va='bottom', wrap=True)
t = ("<NAME>")
ax.text(-125.79, 47.58, t, fontsize=11, color = 'k', fontweight = 'heavy',family='serif', style='italic', ha='left', rotation = 55,
va='bottom', wrap=True)
d = ("Clayoquot C")
ax.text( -126.9, 48.64, d, fontsize=11, color = 'k', fontweight = 'heavy',family='serif', style='italic', ha='left', rotation = 55,
va='bottom', wrap=True)
C = ("<NAME>")
ax.text(-125.14, 48.45, C, fontsize=11, color = 'k',fontweight = 'heavy', family='serif', style='italic', ha='right', rotation = 90,
va='bottom', wrap=True)
X = ("<NAME>")
ax.text(-126.0, 48.3, X, fontsize=11, color = 'k', fontweight = 'heavy',family='serif', style='italic', ha='right', rotation = 50,
va='bottom', wrap=True)
Y = ("<NAME>")
ax.text(-125.73, 48.18, Y, fontsize=11, color = 'k', fontweight = 'heavy',family='serif', style='italic', ha='right', rotation = 50,
va='bottom', wrap=True)
A = ("<NAME>")
ax.text(-124.83, 47.4, A, fontsize=11, color = 'k', fontweight = 'heavy',family='serif', style='italic', ha='right', rotation = 20,
va='bottom', wrap=True)
V = ("Vancouver Island")
ax.text(-124.85, 49.1, V, fontsize=18, color = 'k', family='serif', style='italic', ha='center', rotation = -30,
va='bottom', wrap=True)
from matplotlib.patches import Circle,Ellipse
from matplotlib.collections import PatchCollection
patches = []
# circle = Circle((lon_LB08, lat_LB08), 0.35)
# circle = plt.Circle((lon_LB08, lat_LB08), 0.2, color='blue')
circle = Ellipse((lon_LB08+0.15, lat_LB08), 0.65, 0.35,
angle=0, linewidth=2, fill=False, zorder=2, label = 'Eddy Region')
patches.append(circle)
colors = 100*np.random.rand(len(patches))
p = PatchCollection(patches, alpha=0.4)
p.set_array(np.array(colors))
ax.add_collection(p)
ax.grid()
ax.set_ylim(top=49.5, bottom=47.15136337)
ax.set_xlim(left=-128.51552, right=-124.24068451)
path_to_save = '/home/ssahu/saurav/NEP36_Model_Eval_plots/'
# plt.savefig(path_to_save + 'Domain_edited.png')
# +
# glider_data = pd.read_table('/data/ssahu/CMOP_glider_data/phoebe-2013-7-15_0002013-8-7_2359.csv', \
# delim_whitespace=1, parse_dates= True, header = None, skiprows=1)
glider_data = pd.read_table('/data/ssahu/CMOP_glider_data/phoebe-2013-5-8_0002013-6-1_2359.csv', \
delim_whitespace=1, parse_dates= True, header = None, skiprows=1)
# +
df_cut = glider_data
columns = ['Time[PST]', 'Oxygen[ml/l]', 'Fluorescence[ug/l]', 'Salinity[PSU]', \
'Longitude[degree_east]', 'Depth[m]', 'Latitude[degree_north]', 'CDOM[QSDE]', \
'Backscatter[m^-1 sr^-1]', 'Temperature[C]']
# effective = columns[:7]
df_cut.columns = columns
for i in df_cut.select_dtypes([np.object]).columns[:]:
df_cut[i] = df_cut[i].str.replace(',', '')
for i in df_cut.select_dtypes([np.object]).columns[1:]:
df_cut[i] = df_cut[i].str.replace(',', '').astype(np.float)
df = pd.DataFrame()
df['Time[PST]'] = pd.to_datetime(df_cut.iloc[:,0])
df['Depth[m]'] = df_cut.iloc[:,5]
df['Latitude[degree_north]'] = df_cut.iloc[:,6]
df['Longitude[degree_east]'] = df_cut.iloc[:,4]
df['Temperature[C]'] = df_cut.iloc[:,-1]
df['Salinity[PSU]'] = df_cut.iloc[:,3]
# df = df.set_index('Time[PST]').groupby(pd.Grouper(freq='d')).mean().dropna(how='all')
# df = df.groupby(['Depth[m]']).mean().dropna()
# print(df.groupby(['Latitude[degree_north]', 'Longitude[degree_east]']).groups)
# -
df.head(5)
df.shape[0]
# +
lon_glider = df['Longitude[degree_east]']
lat_glider = df['Latitude[degree_north]']
temp_glider = df['Temperature[C]']
sal_glider = df['Salinity[PSU]']
depth_glider = df['Depth[m]']
pressure_loc = np.empty_like(lon_glider)
SA_loc = np.empty_like(pressure_loc)
CT_loc = np.empty_like(pressure_loc)
spic_loc = np.empty_like(pressure_loc)
rho_loc = np.empty_like(pressure_loc)
for i in np.arange(df.shape[0]):
depth_loc = depth_glider[i]
pressure_loc[i] = gsw.p_from_z(-depth_loc, lat_glider[i])
SA_loc[i] = gsw.SA_from_SP(sal_glider[i], pressure_loc[i], lon_glider[i], lat_glider[i])
CT_loc[i] = gsw.CT_from_pt(sal_glider[i], temp_glider[i])
spic_loc[i] = gsw.spiciness0(SA_loc[i], CT_loc[i])
rho_loc[i] = gsw.density.rho(SA_loc[i], CT_loc[i], 0)
# +
df['Potential Density'] = rho_loc
df['Spice'] = spic_loc
# -
dg = df.groupby(['Latitude[degree_north]', 'Longitude[degree_east]', 'Depth[m]']).mean().dropna()
dg.head(20)
# +
#dg.to_csv('/data/ssahu/CMOP_glider_data/glider_data_may.csv')
# -
dg.index.names
np.array(dg.loc[dg.index[400][0],dg.index[400][1]].index)
# +
### Let us make the along-isopycnal spice for three rho 26.3, 26.4 and 26.5 for each of the locations
# dg.loc[46.908,-124.740]
lon_position = np.empty(dg.index.shape[0])
lat_position = np.empty_like(lon_position)
spic_rho_263 = np.empty_like(lat_position)
spic_rho_264 = np.empty_like(lat_position)
spic_rho_265 = np.empty_like(lat_position)
depth_rho_263 = np.empty_like(lat_position)
depth_rho_264 = np.empty_like(lat_position)
depth_rho_265 = np.empty_like(lat_position)
for k in np.arange(dg.index.shape[0]):
lon_position[k] = dg.index[k][1]
lat_position[k] = dg.index[k][0]
depth_position = np.array(dg.loc[dg.index[k][0],dg.index[k][1]].index)
rho_position = dg.loc[dg.index[k][0],dg.index[k][1]].iloc[:,2] -1000
spic_position = dg.loc[dg.index[k][0],dg.index[k][1]].iloc[:,3]
spic_rho_263[k] = np.interp(26.3, rho_position, spic_position)
depth_rho_263[k] = np.interp(26.3, rho_position, depth_position)
spic_rho_264[k] = np.interp(26.4, rho_position, spic_position)
depth_rho_264[k] = np.interp(26.4, rho_position, depth_position)
spic_rho_265[k] = np.interp(26.5, rho_position, spic_position)
depth_rho_265[k] = np.interp(26.5, rho_position, depth_position)
# +
lon_position = np.empty(dg.index.shape[0])
lat_position = np.empty_like(lon_position)
spic_rho_263 = np.empty_like(lat_position)
spic_rho_264 = np.empty_like(lat_position)
spic_rho_265 = np.empty_like(lat_position)
depth_rho_263 = np.empty_like(lat_position)
depth_rho_264 = np.empty_like(lat_position)
depth_rho_265 = np.empty_like(lat_position)
for k in np.arange(dg.index.shape[0]):
lon_position[k] = dg.index[k][1]
lat_position[k] = dg.index[k][0]
depth_position = np.array(dg.loc[dg.index[k][0],dg.index[k][1]].index)
rho_position = dg.loc[dg.index[k][0],dg.index[k][1]].iloc[:,2] -1000
spic_position = dg.loc[dg.index[k][0],dg.index[k][1]].iloc[:,3]
if rho_position.shape[0]>2:
spic_rho_263[k] = interpolate.interp1d(rho_position, spic_position, fill_value='extrapolate', \
assume_sorted=False)(26.3)
depth_rho_263[k] = interpolate.interp1d(rho_position, depth_position, fill_value='extrapolate', \
assume_sorted=False)(26.3)
spic_rho_264[k] = interpolate.interp1d(rho_position, spic_position, fill_value='extrapolate', \
assume_sorted=False)(26.4)
depth_rho_264[k] = interpolate.interp1d(rho_position, depth_position, fill_value='extrapolate', \
assume_sorted=False)(26.4)
spic_rho_265[k] = interpolate.interp1d(rho_position, spic_position, fill_value='extrapolate', \
assume_sorted=False)(26.5)
depth_rho_265[k] = interpolate.interp1d(rho_position, depth_position, fill_value='extrapolate', \
assume_sorted=False)(26.5)
# spic_rho_263[k] = np.interp(26.3, rho_position, spic_position)
# depth_rho_263[k] = np.interp(26.3, rho_position, depth_position)
# spic_rho_264[k] = np.interp(26.4, rho_position, spic_position)
# depth_rho_264[k] = np.interp(26.4, rho_position, depth_position)
# spic_rho_265[k] = np.interp(26.5, rho_position, spic_position)
# depth_rho_265[k] = np.interp(26.5, rho_position, depth_position)
# -
lat_location = lat_position
lon_location = lon_position
np.max(depth_rho_264)
# +
np.save('/data/ssahu/CMOP_glider_data/spice_on_iso_263.npy', arr=spic_rho_263)
np.save('/data/ssahu/CMOP_glider_data/spice_on_iso_264.npy', arr=spic_rho_264)
np.save('/data/ssahu/CMOP_glider_data/spice_on_iso_265.npy', arr=spic_rho_265)
# +
np.save('/data/ssahu/CMOP_glider_data/lat_location.npy', arr=lat_position)
np.save('/data/ssahu/CMOP_glider_data/lon_location.npy', arr=lon_position)
# +
np.save('/data/ssahu/CMOP_glider_data/depth_on_iso_263.npy', arr=depth_rho_263)
np.save('/data/ssahu/CMOP_glider_data/depth_on_iso_264.npy', arr=depth_rho_264)
np.save('/data/ssahu/CMOP_glider_data/depth_on_iso_265.npy', arr=depth_rho_265)
# -
# !ls '/data/ssahu/CMOP_glider_data/'
# +
lon_location = np.load('/data/ssahu/CMOP_glider_data/lon_location.npy')
lat_location = np.load('/data/ssahu/CMOP_glider_data/lat_location.npy')
spic_rho_263 = np.load('/data/ssahu/CMOP_glider_data/spice_on_iso_263.npy')
spic_rho_264 = np.load('/data/ssahu/CMOP_glider_data/spice_on_iso_264.npy')
spic_rho_265 = np.load('/data/ssahu/CMOP_glider_data/spice_on_iso_265.npy')
depth_rho_263 = np.load('/data/ssahu/CMOP_glider_data/depth_on_iso_263.npy')
depth_rho_264 = np.load('/data/ssahu/CMOP_glider_data/depth_on_iso_264.npy')
depth_rho_265 = np.load('/data/ssahu/CMOP_glider_data/depth_on_iso_265.npy')
# +
iso_NEP = nc.Dataset('/data/ssahu/NEP36_Extracted_Months/2013_short_slice_NEP36_along_isopycnal_larger_offshore_rho_correct.nc')
iso_spic = iso_NEP.variables['spiciness'][...];#[:,:,40:100,110:]
isot = iso_NEP.variables['isot']
nc_tools.show_variables(iso_NEP)
# -
iso_spic.shape
# +
iso_spic_glider_data_mean_time = np.mean(iso_spic[8:33,...], axis=0)
model_rho_263 = iso_spic_glider_data_mean_time[3,...]
model_rho_264 = iso_spic_glider_data_mean_time[4,...]
model_rho_265 = iso_spic_glider_data_mean_time[5,...]
y = np.empty_like(lat_location)
x = np.empty_like(lat_location)
for i in np.arange(lon_location.shape[0]):
y[i], x[i] = geo_tools.find_closest_model_point(
lon_location[i],lat_location[i],lon_wcvi,lat_wcvi,tols={
'NEMO': {'tol_lon': 0.1, 'tol_lat': 0.1},'GEM2.5': {'tol_lon': 0.1, 'tol_lat': 0.1}})
# +
y_new = y[~(np.isnan(x) & np.isnan(x))]
x_new = x[~(np.isnan(x) & np.isnan(x))]
model_263 = model_rho_263[y_new.astype(int), x_new.astype(int)]
model_264 = model_rho_264[y_new.astype(int), x_new.astype(int)]
model_265 = model_rho_265[y_new.astype(int), x_new.astype(int)]
spic_rho_263_finite = spic_rho_263[~(np.isnan(y) & np.isnan(x))]
spic_rho_264_finite = spic_rho_264[~(np.isnan(y) & np.isnan(x))]
spic_rho_265_finite = spic_rho_265[~(np.isnan(y) & np.isnan(x))]
depth_rho_263_finite = depth_rho_263[~(np.isnan(y) & np.isnan(x))]
depth_rho_264_finite = depth_rho_264[~(np.isnan(y) & np.isnan(x))]
depth_rho_264_finite = depth_rho_265[~(np.isnan(y) & np.isnan(x))]
# -
spic_rho_264_finite[np.where(depth_rho_264_finite == np.max(depth_rho_264_finite))]
spic_rho_264_finite[np.where(depth_rho_264_finite == np.min(depth_rho_264_finite))]
np.where((spic_rho_264_finite>-1.1) & (spic_rho_264_finite<-0.9))[0].shape[0]
np.mean(depth_rho_264_finite)
np.min(spic_rho_264_finite)
# +
df =pd.DataFrame()
# df['Observed_Spice'] = survey_iso_spic_A
# df['Model_Spice'] = model_spic_A
df['Depth_Observed_264'] = depth_rho_264_finite
df['Spice_Observed_264'] = spic_rho_264_finite
# fig, ax = plt.subplots(1,1, figsize=(10,8))
# q = df.plot.scatter(x="Observed_Spice", y="Model_Spice", ax = ax, alpha = 0.5)
# q = df.plot.hist2d(survey_iso_spic, model_spic, bins=(1, 1), cmap=plt.cm.Reds)
q = sns.jointplot(x="Spice_Observed_264", y="Depth_Observed_264", data=df, legend='brief', kind="kde", space=0, color="r", sizes=(1,10), stat_func=None)
# gamma2,rms_error,ws = get_stat(df['Observed Spice'], df['Model Spice'])
# anchored_text = AnchoredText("Willmott Score \u2248 {w:.2f}, RMSE \u2248 {r:.2f} and Bias \u2248 {b:.2f}". format(w = ws, r = rms_error, b = gamma2),prop=dict(size=14), loc=3, frameon=True,
# bbox_to_anchor=(1., 1.),
# bbox_transform=q.ax_joint.transAxes
# )
# q.ax_joint.add_artist(anchored_text)
# lims = [
# np.min([ax.get_xlim(), ax.get_ylim()]), # min of both axes
# np.max([ax.get_xlim(), ax.get_ylim()]), # max of both axes
# ]
# ax.tick_params(axis='both',labelsize =16)
# q.set_axis_labels('Observed Spice', 'Model Spice')
# sns.plt.ylim(0, 20)
# sns.plt.xlim(-1, 0)
fig = q.fig
fig.set_figwidth(10)
fig.set_figheight(10)
# -
np.where(depth_rho_264_finite == np.min(depth_rho_264_finite))
# +
spic_rho_263_finite_final = spic_rho_263_finite[spic_rho_263_finite>-10]
model_263 = model_263[spic_rho_263_finite>-10]
spic_rho_263_final = spic_rho_263_finite_final[spic_rho_263_finite_final>-2]
model_263 = model_263[spic_rho_263_finite_final>-2]
# -
sns.set()
sns.set_style('darkgrid')
sns.set_context('talk')
# +
df =pd.DataFrame()
# df['Observed_Spice'] = survey_iso_spic_A
# df['Model_Spice'] = model_spic_A
df['Observed Spice'] = spic_rho_263_final
df['Model Spice'] = model_263
# fig, ax = plt.subplots(1,1, figsize=(10,8))
# q = df.plot.scatter(x="Observed_Spice", y="Model_Spice", ax = ax, alpha = 0.5)
# q = df.plot.hist2d(survey_iso_spic, model_spic, bins=(1, 1), cmap=plt.cm.Reds)
q = sns.jointplot(x="Observed Spice", y="Model Spice", data=df, legend='brief', kind="kde", space=0, color="r", sizes=(1,10), stat_func=None)
# gamma2,rms_error,ws = get_stat(df['Observed Spice'], df['Model Spice'])
# anchored_text = AnchoredText("Willmott Score \u2248 {w:.2f}, RMSE \u2248 {r:.2f} and Bias \u2248 {b:.2f}". format(w = ws, r = rms_error, b = gamma2),prop=dict(size=14), loc=3, frameon=True,
# bbox_to_anchor=(1., 1.),
# bbox_transform=q.ax_joint.transAxes
# )
# q.ax_joint.add_artist(anchored_text)
# lims = [
# np.min([ax.get_xlim(), ax.get_ylim()]), # min of both axes
# np.max([ax.get_xlim(), ax.get_ylim()]), # max of both axes
# ]
# ax.tick_params(axis='both',labelsize =16)
q.set_axis_labels('Observed Spice', 'Model Spice')
# sns.plt.ylim(0, 20)
# sns.plt.xlim(-1, 0)
fig = q.fig
fig.set_figwidth(10)
fig.set_figheight(10)
# +
spic_rho_264_finite_final = spic_rho_264_finite[spic_rho_264_finite>-10]
model_264_finitte_final = model_264[spic_rho_264_finite>-10]
spic_rho_264_final = spic_rho_264_finite_final[spic_rho_264_finite_final>-2]
model_264_final = model_264_finitte_final[spic_rho_264_finite_final>-2]
# -
model_264.shape
lat_location.shape
lat_location_actual = lat_location[~(np.isnan(y) & np.isnan(x))]
lon_location_actual = lon_location[~(np.isnan(y) & np.isnan(x))]
lat_location_actual.shape
# +
spic_rho_negative_1 = spic_rho_264_finite[(spic_rho_264_finite>-1.1) & (spic_rho_264_finite<-0.9)]
lat_location_negative_1 = lat_location_actual[(spic_rho_264_finite>-1.1) & (spic_rho_264_finite<-0.9)]
lon_location_negative_1 = lon_location_actual[(spic_rho_264_finite>-1.1) & (spic_rho_264_finite<-0.9)]
depth_rho_264_negative_1 = depth_rho_264_finite[(spic_rho_264_finite>-1.1) & (spic_rho_264_finite<-0.9)]
# -
np.mean(depth_rho_264_negative_1)
np.max(depth_rho_264_negative_1)
np.min(depth_rho_264_negative_1)
lon_location_negative_1.shape
np.max(depth_rho_264_negative_1)
y_new
# +
df =pd.DataFrame()
# df['Observed_Spice'] = survey_iso_spic_A
# df['Model_Spice'] = model_spic_A
df['Observed Spice'] = spic_rho_264_final
df['Model Spice'] = model_264_final
# fig, ax = plt.subplots(1,1, figsize=(10,8))
# q = df.plot.scatter(x="Observed_Spice", y="Model_Spice", ax = ax, alpha = 0.5)
# q = df.plot.hist2d(survey_iso_spic, model_spic, bins=(1, 1), cmap=plt.cm.Reds)
q = sns.jointplot(x="Observed Spice", y="Model Spice", data=df, legend='brief', kind="kde", space=0, color="r", sizes=(1,10), stat_func=None)
# gamma2,rms_error,ws = get_stat(df['Observed Spice'], df['Model Spice'])
# anchored_text = AnchoredText("Willmott Score \u2248 {w:.2f}, RMSE \u2248 {r:.2f} and Bias \u2248 {b:.2f}". format(w = ws, r = rms_error, b = gamma2),prop=dict(size=14), loc=3, frameon=True,
# bbox_to_anchor=(1., 1.),
# bbox_transform=q.ax_joint.transAxes
# )
# q.ax_joint.add_artist(anchored_text)
# lims = [
# np.min([ax.get_xlim(), ax.get_ylim()]), # min of both axes
# np.max([ax.get_xlim(), ax.get_ylim()]), # max of both axes
# ]
# ax.tick_params(axis='both',labelsize =16)
q.set_axis_labels('Observed Spice', 'Model Spice')
# sns.plt.ylim(0, 20)
# sns.plt.xlim(-1, 0)
fig = q.fig
fig.set_figwidth(10)
fig.set_figheight(10)
# -
spic_rho_264_final_zoom = spic_rho_264_final[(spic_rho_264_final<0) &(spic_rho_264_final>-0.2)]
model_264_final_zoom = model_264_final[(spic_rho_264_final<0) &(spic_rho_264_final>-0.2)]
# +
df =pd.DataFrame()
# df['Observed_Spice'] = survey_iso_spic_A
# df['Model_Spice'] = model_spic_A
df['Observed Spice'] = spic_rho_264_final_zoom
df['Model Spice'] = model_264_final_zoom
# fig, ax = plt.subplots(1,1, figsize=(10,8))
# q = df.plot.scatter(x="Observed_Spice", y="Model_Spice", ax = ax, alpha = 0.5)
# q = df.plot.hist2d(survey_iso_spic, model_spic, bins=(1, 1), cmap=plt.cm.Reds)
q = sns.jointplot(x="Observed Spice", y="Model Spice", data=df, legend='brief', kind="kde", space=0, color="r", sizes=(1,10), stat_func=None)
# gamma2,rms_error,ws = get_stat(df['Observed Spice'], df['Model Spice'])
# anchored_text = AnchoredText("Willmott Score \u2248 {w:.2f}, RMSE \u2248 {r:.2f} and Bias \u2248 {b:.2f}". format(w = ws, r = rms_error, b = gamma2),prop=dict(size=14), loc=3, frameon=True,
# bbox_to_anchor=(1., 1.),
# bbox_transform=q.ax_joint.transAxes
# )
# q.ax_joint.add_artist(anchored_text)
# lims = [
# np.min([ax.get_xlim(), ax.get_ylim()]), # min of both axes
# np.max([ax.get_xlim(), ax.get_ylim()]), # max of both axes
# ]
# ax.tick_params(axis='both',labelsize =16)
q.set_axis_labels('Observed Spice', 'Model Spice')
# sns.plt.ylim(0, 20)
# sns.plt.xlim(-1, 0)
fig = q.fig
fig.set_figwidth(10)
fig.set_figheight(10)
# +
# SCIPY
df =pd.DataFrame()
# df['Observed_Spice'] = survey_iso_spic_A
# df['Model_Spice'] = model_spic_A
df['Observed Spice'] = spic_rho_264_final_zoom
df['Model Spice'] = model_264_final_zoom
# fig, ax = plt.subplots(1,1, figsize=(10,8))
# q = df.plot.scatter(x="Observed_Spice", y="Model_Spice", ax = ax, alpha = 0.5)
# q = df.plot.hist2d(survey_iso_spic, model_spic, bins=(1, 1), cmap=plt.cm.Reds)
q = sns.jointplot(x="Observed Spice", y="Model Spice", data=df, legend='brief', kind="kde", space=0, color="r", sizes=(1,10), stat_func=None)
# gamma2,rms_error,ws = get_stat(df['Observed Spice'], df['Model Spice'])
# anchored_text = AnchoredText("Willmott Score \u2248 {w:.2f}, RMSE \u2248 {r:.2f} and Bias \u2248 {b:.2f}". format(w = ws, r = rms_error, b = gamma2),prop=dict(size=14), loc=3, frameon=True,
# bbox_to_anchor=(1., 1.),
# bbox_transform=q.ax_joint.transAxes
# )
# q.ax_joint.add_artist(anchored_text)
# lims = [
# np.min([ax.get_xlim(), ax.get_ylim()]), # min of both axes
# np.max([ax.get_xlim(), ax.get_ylim()]), # max of both axes
# ]
# ax.tick_params(axis='both',labelsize =16)
q.set_axis_labels('Observed Spice', 'Model Spice')
# sns.plt.ylim(0, 20)
# sns.plt.xlim(-1, 0)
fig = q.fig
fig.set_figwidth(10)
fig.set_figheight(10)
# +
spic_rho_265_finite_final = spic_rho_265_finite[spic_rho_265_finite>-10]
model_265 = model_265[spic_rho_265_finite>-10]
spic_rho_265_final = spic_rho_265_finite_final[spic_rho_265_finite_final>-2]
model_265 = model_265[spic_rho_265_finite_final>-2]
# +
spic_rho_265_final_final = spic_rho_265_final[spic_rho_265_final>-0.2]
model_265 = model_265[spic_rho_265_final>-0.2]
# +
df =pd.DataFrame()
# df['Observed_Spice'] = survey_iso_spic_A
# df['Model_Spice'] = model_spic_A
df['Observed Spice'] = spic_rho_265_final
df['Model Spice'] = model_265
# fig, ax = plt.subplots(1,1, figsize=(10,8))
# q = df.plot.scatter(x="Observed_Spice", y="Model_Spice", ax = ax, alpha = 0.5)
# q = df.plot.hist2d(survey_iso_spic, model_spic, bins=(1, 1), cmap=plt.cm.Reds)
q = sns.jointplot(x="Observed Spice", y="Model Spice", data=df, legend='brief', kind="kde", space=0, color="r", sizes=(1,10), stat_func=None)
# gamma2,rms_error,ws = get_stat(df['Observed Spice'], df['Model Spice'])
# anchored_text = AnchoredText("Willmott Score \u2248 {w:.2f}, RMSE \u2248 {r:.2f} and Bias \u2248 {b:.2f}". format(w = ws, r = rms_error, b = gamma2),prop=dict(size=14), loc=3, frameon=True,
# bbox_to_anchor=(1., 1.),
# bbox_transform=q.ax_joint.transAxes
# )
# q.ax_joint.add_artist(anchored_text)
# lims = [
# np.min([ax.get_xlim(), ax.get_ylim()]), # min of both axes
# np.max([ax.get_xlim(), ax.get_ylim()]), # max of both axes
# ]
# ax.tick_params(axis='both',labelsize =16)
q.set_axis_labels('Observed Spice', 'Model Spice')
# sns.plt.ylim(0, 20)
# sns.plt.xlim(-1, 0)
fig = q.fig
fig.set_figwidth(10)
fig.set_figheight(10)
# -
spic_rho_263.shape
model_263.shape
# +
import matplotlib as mpl
# y = np.empty_like(lat_location)
# x = np.empty_like(lat_location)
cmap = plt.cm.get_cmap('icefire_r', 20)
# cmap = cmo.cm.turbid
# cmap = plt.cm.jet
# cmaplist = [cmap(i) for i in range(cmap.N)]
# # force the first color entry to be grey
# cmaplist[0] = (.8, .8, .8, 1.0)
# # create the new map
# cmap = mpl.colors.LinearSegmentedColormap.from_list(
# 'Custom cmap', cmaplist, cmap.N)
# cmap.set_bad('burlywood')
cmap.set_bad('#8b7765')
fig, ax = plt.subplots(1, 1, figsize=(12,10)); ax.grid()
ax.set_aspect(viz_tools.set_aspect(ax, coords='map', lats=lat_wcvi, adjustable='box-forced'))
# ax.relim()
# ax.autoscale_view(True,True,True)
# ax.autoscale_view(scalex=True)
# ax.autoscale(enable=True, axis='both', tight=None)
# bathy_tools.plot_colourmesh(bathy, title = 'WCVI Subset', fig_size=(9, 9), axis_limits=None, colour_map='winter_r', bins=15, land_colour='burlywood')
# p = ax.pcolormesh(x_wcvi_slice,y_wcvi_slice,Z[y_wcvi_slice,x_wcvi_slice], cmap=cmap, vmin=0, vmax=500)
p = ax.pcolormesh(lon_wcvi,lat_wcvi,Z, cmap=cmap, vmin=0, vmax=1000)
cbar = fig.colorbar(p, ax=ax)
ax.set_aspect('auto')
CS1 = ax.contour(lon_wcvi, lat_wcvi, Z, np.arange(100,110,10))
CLS1 = plt.clabel(CS1, inline=3,fmt='%0.0f m', fontsize=12)
CS1 = ax.contour(lon_wcvi,lat_wcvi,Z, np.arange(200,210,10))
CLS1 = plt.clabel(CS1, inline=3,fmt='%0.0f m', fontsize=12)
# CS1 = ax.contour(lon_wcvi,lat_wcvi,Z, np.arange(1000,1010,10))
# CLS1 = plt.clabel(CS1, inline=3,fmt='%0.0f m', fontsize=12)
# CS1 = ax.contour(lon_wcvi,lat_wcvi,Z, np.arange(500,510,10))
# CLS1 = plt.clabel(CS1, inline=3,fmt='%0.0f m', fontsize=12)
# aspect = 20
# pad_fraction = 0.001
# divider = make_axes_locatable(ax)
# width = axes_size.AxesY(ax, aspect=1./aspect)
# pad = axes_size.Fraction(pad_fraction, width)
# cax = divider.append_axes("right", size=width, pad=pad)
# cbar = fig.colorbar(p, cax=cax)
# for i in np.arange(lat_location.shape[0]):
# y[i], x[i] = geo_tools.find_closest_model_point(
# lon_location[i],lat_location[i],lon_wcvi,lat_wcvi,tols={
# 'NEMO': {'tol_lon': 0.1, 'tol_lat': 0.1},'GEM2.5': {'tol_lon': 0.1, 'tol_lat': 0.1}})
# n = np.empty_like(lat_location)
# m = np.empty_like(lat_location)
# for i in np.arange(lat_location.shape[0]):
# n[i], m[i] = geo_tools.find_closest_model_point(
# lon_location[i],lat_location[i],lon_wcvi,lat_wcvi,tols={
# 'NEMO': {'tol_lon': 0.1, 'tol_lat': 0.1},'GEM2.5': {'tol_lon': 0.1, 'tol_lat': 0.1}})
# y = y.astype(int)
# x = x.astype(int)
# n = n.astype(int)
# m = m.astype(int)
# ax.scatter(x[:], y[:], c='orangered', s=100, alpha=0.5, label = 'Falkor CTD locations')
ax.set_xlabel('Longitude', fontsize=16)
ax.set_ylabel('Latitude', fontsize=16)
# ax.set_title('WCVI slice of NEP036 Model Domain', fontsize=20)
lon_W02 = -124.89500
lat_W02 = 48.04167
lon_W01 = -124.82500
lat_W01 = 48.31500
lon_swift = -125.00
lat_swift = 48.55
ax.scatter(lon_swift, lat_swift, c='k', marker='*', s=50, alpha=0.8)#, label = 'Swiftsure Bank')
# S = ("SwB")
S = ("Swiftsure\nBank")
ax.text(lon_swift, lat_swift, S, fontsize=11, color = 'k', fontweight = 'heavy',family='serif', style='italic', ha='center', rotation = 10,
va='top', wrap=False)
# ax.text(lon_swift, lat_swift, S, fontsize=11, color = 'k', fontweight = 'heavy',family='serif', style='italic', ha='left', rotation = 10,
# va='bottom', wrap=False)
lat_cape = 48.3831
lon_cape = -124.7144
ax.scatter(lon_cape, lat_cape, c='k', marker='*', s=50, alpha=0.8)#, label = 'Cape Flattery')
CP = ("Cape\nFlattery")
ax.text(lon_cape, lat_cape, CP, fontsize=11, color = 'k', fontweight = 'heavy',family='serif', style='italic', ha='center', rotation = 10,
va='top', wrap=False)
lat_jdfs = 48.5
lon_jdfs = -124.4
JD = ("JdF\nStrait")
ax.text(lon_jdfs, lat_jdfs, JD, fontsize=11, color = 'white', fontweight = 'heavy',family='serif', style='italic', ha='center', rotation = -35,
va='top', wrap=False)
lat_EP = 49.3835
lon_EP = -126.5447
ax.scatter(lon_EP, lat_EP, c='k', marker='*', s=50, alpha=0.8)
EP = ("Estevan\nPoint")
ax.text(lon_EP, lat_EP, EP, fontsize=11, color = 'k', fontweight = 'heavy',family='serif', style='italic', ha='left', rotation = 0,
va='top', wrap=False)
lon_LB08 = -125.4775
lat_LB08 = 48.4217
ax.scatter(lon_falkor[::3], lat_falkor[::3], c='yellow', marker='o', s=10, alpha=0.9, label = 'Pathways MVP')
ax.scatter(lon_location_negative_1[::50], lat_location_negative_1[::50], c=depth_rho_264_negative_1[::50], cmap= cmap, vmin = 0, vmax = 1000, marker='o',\
s=100,edgecolors = 'k', alpha=0.1, label = 'CMOP Glider -1 Cluster')
ax.scatter(lon_LB08, lat_LB08, c='indigo', marker='*', s=150, linewidths=1.2, alpha=1, label = 'LB08')
lon_A1 = -126.20433
lat_A1 = 48.52958
ax.scatter(lon_A1, lat_A1, c='cyan',marker='*', s=150, alpha=0.8, label = 'A1 Mooring')
# ax.scatter(lon_W01, lat_W01, c='cyan', s=150, alpha=0.8, label = 'W01')
# ax.scatter(lon_W02, lat_W02, c='red', s=150, alpha=0.8, label = 'W02')
# ax.scatter(lon_location[:], lat_location[:], c='purple', s=100, alpha=0.3, label = 'Falkor CTD locations')
legend = ax.legend(loc='lower left', fancybox=True, framealpha=0.85)
# ax.set_xlabel('x_index')
# ax.set_ylabel('y_index')
# ax.set_title('WCVI slice of NEP036 Model Domain', fontsize=20)
# viz_tools.plot_land_mask(ax, bathy)#, yslice=y_wcvi_slice, xslice=x_wcvi_slice, color='burlywood')
# cbar = fig.colorbar(p, cax=ax)
cbar.ax.set_ylabel('Depth [m]', fontsize=16)
ax.tick_params(axis='both',labelsize =16)
cbar.ax.tick_params(labelsize=16)
# t = ("JdF Canyon")
# ax.text(lon_W01, lat_W01, t, fontsize=11, color = 'darkblue', family='serif', style='italic', ha='left', rotation = 55,
# va='bottom', wrap=True)
t = ("<NAME>")
ax.text(-125.79, 47.58, t, fontsize=11, color = 'k', fontweight = 'heavy',family='serif', style='italic', ha='left', rotation = 55,
va='bottom', wrap=True)
d = ("<NAME>")
ax.text( -126.9, 48.64, d, fontsize=11, color = 'k', fontweight = 'heavy',family='serif', style='italic', ha='left', rotation = 55,
va='bottom', wrap=True)
C = ("Sp<NAME>")
ax.text(-125.14, 48.45, C, fontsize=11, color = 'k',fontweight = 'heavy', family='serif', style='italic', ha='right', rotation = 90,
va='bottom', wrap=True)
X = ("<NAME>")
ax.text(-126.0, 48.3, X, fontsize=11, color = 'k', fontweight = 'heavy',family='serif', style='italic', ha='right', rotation = 50,
va='bottom', wrap=True)
Y = ("<NAME>")
ax.text(-125.73, 48.18, Y, fontsize=11, color = 'k', fontweight = 'heavy',family='serif', style='italic', ha='right', rotation = 50,
va='bottom', wrap=True)
A = ("Quinault C")
ax.text(-124.83, 47.4, A, fontsize=11, color = 'k', fontweight = 'heavy',family='serif', style='italic', ha='right', rotation = 20,
va='bottom', wrap=True)
V = ("Vancouver Island")
ax.text(-124.85, 49.1, V, fontsize=18, color = 'k', family='serif', style='italic', ha='center', rotation = -30,
va='bottom', wrap=True)
from matplotlib.patches import Circle,Ellipse
from matplotlib.collections import PatchCollection
patches = []
# circle = Circle((lon_LB08, lat_LB08), 0.35)
# circle = plt.Circle((lon_LB08, lat_LB08), 0.2, color='blue')
circle = Ellipse((lon_LB08+0.15, lat_LB08), 0.65, 0.35,
angle=0, linewidth=2, fill=False, zorder=2, label = 'Eddy Region')
patches.append(circle)
colors = 100*np.random.rand(len(patches))
p = PatchCollection(patches, alpha=0.4)
p.set_array(np.array(colors))
ax.add_collection(p)
ax.grid()
ax.set_ylim(top=49.5, bottom=47.15136337)
ax.set_xlim(left=-128.51552, right=-124.24068451)
path_to_save = '/home/ssahu/saurav/NEP36_Model_Eval_plots/'
# plt.savefig(path_to_save + 'Domain_edited.png')
# -
np.nanmean(model_263)
np.min(spic_rho_263_finite_final)
x_new.shape
y.shape
spic_rho_263_finite.shape
model_rho_264.shape
lat_wcvi.shape
depth_rho_264_negative_1
f[:2]
spic_position.shape
lon_position
trial = np.array(dg.loc[46.908,-124.740].iloc[:,0])
trial
dg.index[:][0][0]
dg.index[405284][0]
dg.index[405284][1]
# +
def add_model_MVP_data(survey_file, iso_level, survey_iso_spic, model_spic):
mask = nc.Dataset('/data/mdunphy/NEP036-N30-OUT/INV/mesh_mask.nc')
mat = scipy.io.loadmat(survey_file)
depths_survey = mat['depths'][:,0]
lat_survey = mat['latitude'][:,0]
lon_survey = mat['longitude'][:,0] - 100
# den_survey = mat['density'][:]
pden_survey = mat['pden'][:]
temp_survey = mat['temp'][:]
sal_survey = mat['salinity'][:]
mtime = mat['mtime'][:,0]
pressure_survey = np.empty_like(temp_survey)
SA_survey = np.empty_like(temp_survey)
CT_survey = np.empty_like(temp_survey)
spic_survey = np.empty_like(temp_survey)
rho_survey = np.empty_like(temp_survey)
bathy = nc.Dataset('/data/mdunphy/NEP036-N30-OUT/INV/Bathymetry_EastCoast_NEMO_R036_GEBCO_corr_v14.nc')
mbathy = mask.variables['mbathy']#[0,220:280, 575:630]
Z = bathy.variables['Bathymetry']
lon = bathy['nav_lon'][180:350,480:650]
lat = bathy['nav_lat'][180:350,480:650]
for j in np.arange(depths_survey.shape[0]):
for i in np.arange(lat_survey.shape[0]):
pressure_survey[j,i] = gsw.p_from_z(-depths_survey[j],lat_survey[i])
SA_survey[j,i] = gsw.SA_from_SP(sal_survey[j,i], pressure_survey[j,i], lon_survey[i], lat_survey[i])
CT_survey[j,i] = gsw.CT_from_pt(sal_survey[j,i], temp_survey[j,i])
spic_survey[j,i] = gsw.spiciness0(SA_survey[j,i], CT_survey[j,i])
rho_survey[j,i] = gsw.density.rho(SA_survey[j,i], CT_survey[j,i], 0)
#Values indicate that pden is the true representative of the potential density calculated by rho_survey here
y = np.empty_like(lat_survey)
x = np.empty_like(y)
for i in np.arange(lat_survey.shape[0]):
y[i], x[i] = geo_tools.find_closest_model_point(
lon_survey[i],lat_survey[i],lon,lat,tols={
'NEMO': {'tol_lon': 0.1, 'tol_lat': 0.1},'GEM2.5': {'tol_lon': 0.1, 'tol_lat': 0.1}})
rho_0 = isot[iso_level]
spic_rho = np.empty((spic_survey.shape[1]))
for i in np.arange(spic_survey.shape[1]):
spic_loc = spic_survey[:,i]
rho_loc = rho_survey[:,i]
spic_rho[i] = np.interp(rho_0, rho_loc[:]-1000, spic_loc[:])
spic_rho_finite = spic_rho[~np.isnan(spic_rho)]
y = y[~np.isnan(spic_rho)]
x = x[~np.isnan(spic_rho)]
if survey_file == survey_mat_file_A:
t =116
if survey_file == survey_mat_file_B:
t =116
if survey_file == survey_mat_file_D:
t =118
if survey_file == survey_mat_file_E:
t =119
if survey_file == survey_mat_file_G:
t =120
model_add = np.empty_like(y)
for i in np.arange(y.shape[0]):
model_add[i] = iso_spic[t,iso_level,np.int(y[i]),np.int(x[i])]
spic_rho_finite = spic_rho_finite[~np.isnan(model_add)]
model_add = model_add[~np.isnan(model_add)]
# model_add[np.isnan(model_add)] = 0
# model_add = np.ma.masked_equal(model_add, value=0)
survey_iso_spic = np.append(arr= survey_iso_spic, values= spic_rho_finite)
model_spic = np.append(arr= model_spic, values=model_add)
return survey_iso_spic, model_spic
import matplotlib
matplotlib.use('TkAgg')
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
import seaborn as sns
import numpy as np
class SeabornFig2Grid():
def __init__(self, seaborngrid, fig, subplot_spec):
self.fig = fig
self.sg = seaborngrid
self.subplot = subplot_spec
if isinstance(self.sg, sns.axisgrid.FacetGrid) or \
isinstance(self.sg, sns.axisgrid.PairGrid):
self._movegrid()
elif isinstance(self.sg, sns.axisgrid.JointGrid):
self._movejointgrid()
self._finalize()
def _movegrid(self):
""" Move PairGrid or Facetgrid """
self._resize()
n = self.sg.axes.shape[0]
m = self.sg.axes.shape[1]
self.subgrid = gridspec.GridSpecFromSubplotSpec(n,m, subplot_spec=self.subplot)
for i in range(n):
for j in range(m):
self._moveaxes(self.sg.axes[i,j], self.subgrid[i,j])
def _movejointgrid(self):
""" Move Jointgrid """
h= self.sg.ax_joint.get_position().height
h2= self.sg.ax_marg_x.get_position().height
r = int(np.round(h/h2))
self._resize()
self.subgrid = gridspec.GridSpecFromSubplotSpec(r+1,r+1, subplot_spec=self.subplot)
self._moveaxes(self.sg.ax_joint, self.subgrid[1:, :-1])
self._moveaxes(self.sg.ax_marg_x, self.subgrid[0, :-1])
self._moveaxes(self.sg.ax_marg_y, self.subgrid[1:, -1])
def _moveaxes(self, ax, gs):
#https://stackoverflow.com/a/46906599/4124317
ax.remove()
ax.figure=self.fig
self.fig.axes.append(ax)
self.fig.add_axes(ax)
ax._subplotspec = gs
ax.set_position(gs.get_position(self.fig))
ax.set_subplotspec(gs)
def _finalize(self):
plt.close(self.sg.fig)
self.fig.canvas.mpl_connect("resize_event", self._resize)
self.fig.canvas.draw()
def _resize(self, evt=None):
self.sg.fig.set_size_inches(self.fig.get_size_inches())
# +
model_spic = []
survey_iso_spic = []
survey_iso_spic_A, model_spic_A = add_model_MVP_data(survey_file=survey_mat_file_A, \
iso_level=4,model_spic=model_spic, \
survey_iso_spic=survey_iso_spic)
survey_iso_spic_B, model_spic_B = add_model_MVP_data(survey_file=survey_mat_file_B, \
iso_level=4,model_spic=model_spic, \
survey_iso_spic=survey_iso_spic)
survey_iso_spic_D, model_spic_D = add_model_MVP_data(survey_file=survey_mat_file_D, \
iso_level=4,model_spic=model_spic, \
survey_iso_spic=survey_iso_spic)
survey_iso_spic_E, model_spic_E = add_model_MVP_data(survey_file=survey_mat_file_E, \
iso_level=4,model_spic=model_spic, \
survey_iso_spic=survey_iso_spic)
survey_iso_spic_G, model_spic_G = add_model_MVP_data(survey_file=survey_mat_file_G, \
iso_level=4,model_spic=model_spic, \
survey_iso_spic=survey_iso_spic)
model_spic = []
survey_iso_spic = []
for survey_file in all_surveys:
survey_iso_spic, model_spic = add_model_MVP_data(survey_file=survey_file, \
iso_level=4,model_spic=model_spic, \
survey_iso_spic=survey_iso_spic)
# -
sns.set()
# +
df =pd.DataFrame()
# df['Observed_Spice'] = survey_iso_spic_A
# df['Model_Spice'] = model_spic_A
df['Observed Spice'] = survey_iso_spic
df['Model Spice'] = model_spic
fig, ax = plt.subplots(1,1, figsize=(10,8))
# q = df.plot.scatter(x="Observed_Spice", y="Model_Spice", ax = ax, alpha = 0.5)
# q = df.plot.hist2d(survey_iso_spic, model_spic, bins=(1, 1), cmap=plt.cm.Reds)
q = sns.jointplot(x="Observed Spice", y="Model Spice", data=df, ax = ax, legend='brief', kind="kde", space=0, color="r", sizes=(1,10),stat_func=None)
# gamma2,rms_error,ws = get_stat(df['Observed Spice'], df['Model Spice'])
# anchored_text = AnchoredText("Willmott Score \u2248 {w:.2f}, RMSE \u2248 {r:.2f} and Bias \u2248 {b:.2f}". format(w = ws, r = rms_error, b = gamma2),prop=dict(size=14), loc=3, frameon=True,
# bbox_to_anchor=(1., 1.),
# bbox_transform=q.ax_joint.transAxes
# )
# q.ax_joint.add_artist(anchored_text)
# lims = [
# np.min([ax.get_xlim(), ax.get_ylim()]), # min of both axes
# np.max([ax.get_xlim(), ax.get_ylim()]), # max of both axes
# ]
# ax.tick_params(axis='both',labelsize =16)
q.set_axis_labels('Observed Spice', 'Model Spice')
# fig = q.fig
# fig.set_figwidth(10)
# fig.set_figheight(10)
# +
### Take the average of the model data for that month and then come up with the along iso-pycnal analysis
| Glider_data_off_Washington.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Preprocess the VAERS data
#
# In this notebook, we prepare the dataset which will be used in the data annotation and training
import os
import pandas as pd
from tqdm import tqdm
# # Load raw data
#
# The raw data files are downloaded from https://vaers.hhs.gov/data.html,
# we could use these raw files for better screening
# +
df_vax = pd.read_csv('2021VAERSVAX.csv', encoding='cp1252')
df_sym = pd.read_csv('2021VAERSSYMPTOMS.csv', encoding='cp1252')
df_rpt = pd.read_csv('2021VAERSData.csv', encoding='cp1252')
print('* df report size:', df_rpt.VAERS_ID.count())
print('* df symptom size:', df_sym.VAERS_ID.count())
print('* df vax size:', df_vax.VAERS_ID.count())
# -
# ## Check data sample
# check the sample data for the df vax
df_vax.head()
df_sym.head(10)
# since there are too many columns, we could revert the matrix to show all columns
df_rpt.head(10).T
# ## Merge the subset
# +
# merge the report text and vaccination info
# we don't need all of the columns in each dataframe
df = df_rpt[['VAERS_ID', 'AGE_YRS', 'SEX', 'VAX_DATE', 'SYMPTOM_TEXT', 'ALLERGIES']].merge(
df_vax[['VAERS_ID', 'VAX_TYPE', 'VAX_MANU']].drop_duplicates(subset='VAERS_ID'),
on='VAERS_ID',
how='left'
)
# merge the first symptom as the main label
# since there are multiple labels for each report
# we could also collect all symptoms for multi-label classification tasks
df = df.merge(
df_sym[['VAERS_ID', 'SYMPTOM1']].drop_duplicates(subset='VAERS_ID'),
on='VAERS_ID',
how='left'
)
# we only use the COVID19 vaccination data
df = df[df['VAX_TYPE']=='COVID19']
# change the default datetime format for eaiser query
df['VAX_DATE'] = pd.to_datetime(df['VAX_DATE'], format='%m/%d/%Y')
# make sure the format is string for this two
df['SYMPTOM_TEXT'] = df['SYMPTOM_TEXT'].astype(str)
df['SYMPTOM1'] = df['SYMPTOM1'].astype(str)
# calcuate the text length for the symptoms
df['TEXT_LEN'] = df['SYMPTOM_TEXT'].apply(lambda r: len(r))
# replace the blank in symptom name with underline
df['SYMPTOM'] = df['SYMPTOM1'].apply(lambda r: r.replace(' ', '_'))
# update some data string
df.loc[df.SYMPTOM=='Chills', 'SYMPTOM'] = 'Chill'
# then we could remove the unused columns
df.drop(columns=['SYMPTOM1'], inplace=True)
# let's see how our dataframe looks like
print('* df size:', df['VAERS_ID'].count())
df.head()
# -
# ## Get a smaller dataset for working
#
# Let's get our working dataset
# +
# first, we only use the data in 2021
dft = df[df['VAX_DATE']>='2021-01-01']
print('* dft size:', dft['VAERS_ID'].count())
# then, we remove those records that are too short
dft = dft[dft['TEXT_LEN']>80]
print('* dft size:', dft['VAERS_ID'].count())
# then drop those not used columns
dft.drop(columns=['TEXT_LEN', 'ALLERGIES', 'VAX_TYPE'], inplace=True)
dft.head()
# -
# let's see how this dataset looks like by the symptom label
# the default groupby result is not well ordered,
# so we sort the results by the number of records
dft.groupby(['SYMPTOM'])[['VAERS_ID']].count().sort_values(by='VAERS_ID').tail(15)
# # Output the selected top 10
# ## Define the top 10
# +
# we could select the top 10 symptoms according to our dataset
aes = dft.groupby(['SYMPTOM'])[['VAERS_ID']].count().sort_values(by='VAERS_ID').tail(10).index.tolist()
print('* top 10 symptoms:', aes)
# or we could specify 10 symptoms directly,
# then we could put our own selection here
aes = [
'Pyrexia', 'Chill', 'Headache', 'Fatigue', 'Pain',
'Nausea', 'Dizziness', 'Pain_in_extremity', 'Injection_site_pain', 'Myalgia'
]
# and let's see how our selected 10 symptoms
dft[dft.SYMPTOM.isin(aes)].groupby(['SYMPTOM'])[['VAERS_ID']].count().sort_values(by='VAERS_ID').tail(15)
# -
# ## output samples
# +
dft_allsamp = []
dft_allae = []
for ae in tqdm(aes):
# get the sample df for this symptom
dft_ae = dft[dft['SYMPTOM']==ae]
# sampling by the default method
dft_sample = dft_ae.sample(n=50)
# gather the dataframe for further use
dft_allsamp.append(dft_sample)
dft_allae.append(dft_ae)
# output the sample symptom txt file
idx = 0
for _, row in dft_sample.iterrows():
# get the text content
txt = row['SYMPTOM_TEXT']
# create a filename
fn = '%s_%02d.txt' % (ae, idx)
# and make it a full path
full_fn = os.path.join(
'sample', fn
)
# write to disk
with open(full_fn, 'w') as f:
f.write(txt)
idx += 1
# put all dataframe into one big dataframe
dft_allsamp = pd.concat(dft_allsamp)
dft_allae = pd.concat(dft_allae)
# get those records are not sampled
dft_notsamp = pd.concat([dft_allsamp, dft_allae])
dft_notsamp = dft_notsamp.drop_duplicates(keep=False)
print('* dft_allae:', dft_allae.shape)
print('* dft_allsamp:', dft_allsamp.shape)
print('* dft_notsamp:', dft_notsamp.shape)
print('* generated samples')
# -
# ## output large and csv
# +
dft_notsamp.to_csv('large.csv', index=False)
dft_allsamp.to_csv('sample.csv', index=False)
print('* generated large.csv and sample.csv')
| data_preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pb111/Python-tutorials-and-projects/blob/master/Python_List_Comprehension.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="VQnCjyt0wlRo"
# # **Python List Comprehension**
#
# - In this article, we will learn about Python list comprehensions, and how to use it.
# + [markdown] id="l9GgpNBMIKnw"
# ## **1. List Comprehension: Elegant way to create new List**
#
# - List comprehension is an elegant and concise way to create a new list from an existing list in Python.
#
# - A list comprehension consists of an expression followed by for statement inside square brackets.
#
# - Here is an example to make a list with each item being increasing power of 2.
# + colab={"base_uri": "https://localhost:8080/"} id="MGwSwTP7IRlb" outputId="a8b0d69e-1622-466c-b2c4-1079e5702a4f"
pow2 = [2 ** x for x in range(10)]
print(pow2)
# + [markdown] id="3Sc6_ZS4JCaq"
# https://www.programiz.com/python-programming/list
# + [markdown] id="LVB4Joe-JFl6"
# This code is equivalent to:
# + colab={"base_uri": "https://localhost:8080/"} id="vczWDfRrJKnd" outputId="c64c9f6d-d39e-4ceb-ae48-f726f96f957f"
pow2 = []
for x in range(10):
pow2.append(2 ** x)
print(pow2)
# + [markdown] id="V6HsGfTHJZ3v"
# - A list comprehension can optionally contain more for or [if statements](https://www.programiz.com/python-programming/if-elif-else).
# - An optional `if` statement can filter out items for the new list.
# - Here are some examples.
# + colab={"base_uri": "https://localhost:8080/"} id="FiokNde5JTmx" outputId="32bf8db4-ac0f-43e8-bd3b-b332ae6b11e0"
pow2 = [2 ** x for x in range(10) if x > 5]
pow2
# + colab={"base_uri": "https://localhost:8080/"} id="4eL9mbkRJ_zj" outputId="da6be7d8-e31a-490b-bea8-47cbb6aa5c17"
odd = [x for x in range(20) if x%2 == 1]
odd
# + colab={"base_uri": "https://localhost:8080/"} id="XBGhwpwBKMVB" outputId="2aae7b99-2627-4713-860f-96717857197b"
even = [x for x in range(20) if x%2 == 0]
even
# + colab={"base_uri": "https://localhost:8080/"} id="9VlhQ4JCKZqo" outputId="e66c2358-ee11-4d0b-dc02-8e2e02d6c55d"
[x+y for x in ['Python ','C '] for y in ['Language','Programming']]
# + [markdown] id="R_V7BM9lLDsP"
# https://www.programiz.com/python-programming/list
# + [markdown] id="iwzPr60vztx8"
# ## **2. List Comprehension vs For Loop in Python**
#
# - Suppose, we want to separate the letters of the word `human` and add the letters as items of a list.
# - The first thing that comes in mind would be using [for loop](https://www.programiz.com/python-programming/for-loop).
# + [markdown] id="U80BLvOj0cWu"
# #### **Example 1: Iterating through a string Using for Loop**
# + colab={"base_uri": "https://localhost:8080/"} id="84W8g6wz1D4U" outputId="6585cde4-8df8-4432-cf68-075d7bf043e2"
h_letters = []
for letter in 'human':
h_letters.append(letter)
print(h_letters)
# + [markdown] id="nUDeVZ6s15Yq"
# - However, Python has an easier way to solve this issue using List Comprehension.
# - List comprehension is an elegant way to define and create lists based on existing lists.
#
# - Let’s see how the above program can be written using list comprehensions.
# + [markdown] id="X9ktct-J2_ap"
# #### **Example 2: Iterating through a string Using List Comprehension**
# + colab={"base_uri": "https://localhost:8080/"} id="8a_evoWG1Mek" outputId="6b38e589-8156-4689-9daa-a55978ee5155"
h_letters = [letter for letter in 'human']
print(h_letters)
# + [markdown] id="pGMCsZm33pLk"
# - In the above example, a new list is assigned to variable `h_letters`, and list contains the items of the iterable string 'human'.
# - We call `print()` function to receive the output.
#
#
# + [markdown] id="3mE3VLkK4F1l"
# #### **Syntax of List Comprehension**
#
# - [expression for item in list]
#
# + [markdown] id="OeAeouB54ZzD"
# 
# + [markdown] id="V7LUZ99h48v_"
# - We can now identify where list comprehensions are used.
#
# - If you noticed, `human` is a string, not a list. This is the power of list comprehension. It can identify when it receives a string or a tuple and work on it like a [list](https://www.programiz.com/python-programming/list).
#
# - You can do that using loops. However, not every loop can be rewritten as list comprehension.
#
# - But as you learn and get comfortable with list comprehensions, you will find yourself replacing more and more loops with this elegant syntax.
#
#
# + [markdown] id="pwFNBSc75bcM"
# ## **3. List Comprehensions vs Lambda functions**
#
# - List comprehensions aren't the only way to work on lists.
# - Various built-in functions and [lambda functions](https://www.programiz.com/python-programming/anonymous-function) can create and modify lists in less lines of code.
# + [markdown] id="1kwrRgiD6JvR"
# #### **Example 3: Using Lambda functions inside List**
# + colab={"base_uri": "https://localhost:8080/"} id="7K20rD_b6kL0" outputId="54040062-c374-4eb0-ae9a-87fa81114cc6"
letters = list(map(lambda x : x, 'human'))
print(letters)
# + [markdown] id="dhadkfcP8mUI"
# - However, list comprehensions are usually more human readable than lambda functions. It is easier to understand what the programmer was trying to accomplish when list comprehensions are used.
# + [markdown] id="91gFHXa_8phV"
# ## **4. Conditionals in List Comprehension**
#
# - List comprehensions can utilize conditional statement to modify existing list (or other tuples).
# - We will create list that uses mathematical operators, integers, and [range()](https://www.programiz.com/python-programming/methods/built-in/range).
# + [markdown] id="LRRgKNf4-27t"
# #### **Example 4: Using if with List Comprehension**
# + colab={"base_uri": "https://localhost:8080/"} id="kJtDsJOS6pwf" outputId="ac4366a7-3811-459f-be65-e7e6c30f291c"
number_list = [x for x in range(20) if x%2 == 0]
number_list
# + [markdown] id="GmkfRf4__Q9u"
# - The list, `number_list`, will be populated by the items in range from 0-19 if the item's value is divisible by 2.
# + [markdown] id="vRv0OJLZ_beE"
# #### **Example 5: Nested IF with List Comprehension**
# + colab={"base_uri": "https://localhost:8080/"} id="vjf-YR2Q_HDj" outputId="16e6af54-de9e-424d-da67-937704e8d54b"
num_list = [y for y in range(100) if y%2 == 0 if y%5 == 0]
num_list
# + [markdown] id="fAX8GsYfAKc9"
# Here, list comprehension checks:
#
# - 1. Is `y` divisible by 2 or not?
# - 2. Is `y` divisible by 5 or not?
#
# If `y` satisfies both conditions, `y` is appended to `num_list`.
#
#
# + [markdown] id="7GAezDzYA0tE"
# #### **Example 6: if...else With List Comprehension**
# + colab={"base_uri": "https://localhost:8080/"} id="nsN8xTC2_2ic" outputId="3c17cf96-b98f-4782-c637-8ce3a95f9dec"
obj = ["Even" if i%2 == 0 else "Odd" for i in range(10)]
obj
# + [markdown] id="goqxpgqSB1Ce"
# - Here, list comprehension will check the 10 numbers from 0 to 9.
# - If `i` is divisible by 2, then `Even` is appended to the `obj` list.
# - If not, `Odd` is appended.
# + [markdown] id="xKVhR42ACjII"
# ## **5. Nested Loops in List Comprehension**
#
# - Suppose, we need to compute the transpose of a matrix that requires nested for loop.
# - Let’s see how it is done using normal for loop first.
# + [markdown] id="cyeblQBwDIZ2"
# #### **Example 7: Transpose of Matrix using Nested Loops**
# + colab={"base_uri": "https://localhost:8080/"} id="4y15dt4eCUOO" outputId="5ec52fd4-a76f-456f-8dc6-18326ac97ffd"
transposed = []
matrix = [[1, 2, 3, 4], [4, 5, 6, 8]]
for i in range(len(matrix[0])):
transposed_row = []
for row in matrix:
transposed_row.append(row[i])
transposed.append(transposed_row)
print(transposed)
# + [markdown] id="boFj0-JZD9FQ"
# - The above code use two for loops to find transpose of the matrix.
#
# - We can also perform nested iteration inside a list comprehension.
#
# - In this section, we will find transpose of a matrix using nested loop inside list comprehension.
# + [markdown] id="-_gRc5n3EHYu"
# #### **Example 8: Transpose of a Matrix using List Comprehension**
# + colab={"base_uri": "https://localhost:8080/"} id="cjbB7YRhDyC8" outputId="c9ed424d-d3a7-4d28-c86b-71ab8639fff4"
matrix = [[1, 2], [3,4], [5,6], [7,8]]
transpose = [[row[i] for row in matrix] for i in range(2)]
print (transpose)
# + [markdown] id="nbx6Dnh2EmR9"
# - In above program, we have a variable `matrix` which have `4` rows and `2` columns.
# - We need to find transpose of the `matrix`.
# - For that, we used list comprehension.
# + [markdown] id="jR80JpWbE6it"
# - **Note**:
#
# - The nested loops in list comprehension don’t work like normal nested loops.
# - In the above program, for i in range(2) is executed before row[i] for row in matrix.
# - Hence at first, a value is assigned to i then item directed by row[i] is appended in the transpose variable.
# + [markdown] id="FO22lARUFP5o"
# ## **6. Key Points to Remember**
#
# - List comprehension is an elegant way to define and create lists based on existing lists.
# - List comprehension is generally more compact and faster than normal functions and loops for creating list.
# - However, we should avoid writing very long list comprehensions in one line to ensure that code is user-friendly.
# - Remember, every list comprehension can be rewritten in for loop, but every for loop can’t be rewritten in the form of list comprehension.
# + [markdown] id="VObMxBUSFfoS"
# https://www.programiz.com/python-programming/list-comprehension
| Python_List_Comprehension.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/piyushjain220/TSAI/blob/main/Edu/Module4/Project4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ewbaAD6rM3OR"
# Project description:
# - Find a dataset to conduct any form of machine learning we have NOT discussed in class
# - Using any form of trainin/testing method we have discussed in class
# - Train the model using appropriate methods and share results.
#
# As I already submittd the original project, I will keep this project and make sure it also meets with the requirements above.
# + [markdown] id="952yWTCgM3Oh"
# Original project Description:
#
# - Download any dataset from Kaggle that contains chat data to conduct natural language processing.
# - Load 2 tables of CSV data into a SQL database of your choice
# - Optimize your database by creating indexes and primary keys on appropriate columns
# - Add a new column of data that has not been included in the csv
# - Conduct any type of NLP (language detection, sentiment analysis, tokenization, etc.) and add to the database
# - Connect to database from Python, select some columns database and produce a visualization/analysis of it.
# + [markdown] id="SEU-MiDTM3Oj"
# ___
# **Important note**: Although I had several questions regarding the project description (email sent to eduonix support - ticket 5623271728) and had some doubt about the use of SQL for this specific ML course, I still tried to finalize the project, resulting in the elaboration below.
# + [markdown] id="EH6N4L16M3Ok"
# ___
# <span style="color:green">**1. Download any dataset from Kaggle that contains chat data to conduct natural language processing.**</span>
#
# For the dataset I choose for the Teld talks set, as can be found on Kaggle:
# https://www.kaggle.com/rounakbanik/ted-talks
#
# Goal is to match the highest rated category with NLP analysis on the transcripts.
# + [markdown] id="Pfnh5hgAM3Ol"
# ___
# <span style="color:green">**2. Load 2 tables of CSV data into a SQL database of your choice**</span>
#
# Both ted_main.csv and transcripts.csv will be loaded into a sqlite database using sqlalchemy
# + id="txsIYppKM3Om"
import matplotlib
import matplotlib.pyplot as plt
from sqlalchemy import Table, MetaData, create_engine
from sqlalchemy.engine import reflection
import pandas as pd
from termcolor import colored
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import MultinomialNB
from sklearn.ensemble import AdaBoostClassifier
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.metrics import accuracy_score
import datetime
# %matplotlib inline
# + id="ZKYU6O49M3On"
engine = create_engine('sqlite:///tedtalk.db', echo=False)
conn = engine.connect()
# + id="k7EW5zbZM3On" outputId="b43fdb41-4f44-47a3-e239-af99e03cc906"
with open('ted_main.csv', 'r', encoding="utf-8") as file:
data_df = pd.read_csv(file, encoding='utf-8')
data_df.to_sql('ted_main', con=engine, index=False, if_exists='replace')
with open('transcripts_unique.csv', 'r', encoding="utf-8") as file:
data_df = pd.read_csv(file, encoding='utf-8')
data_df.to_sql('transcripts', con=engine, index=False, if_exists='replace')
# check if tables are loaded indeed
print(colored('tables:\n', 'blue'), engine.table_names(), '\n')
# check first line of ted_main
sql_text = 'SELECT * FROM ted_main LIMIT 1'
result = conn.execute(sql_text).fetchall()
print(colored('1st line table ted_main:\n', 'blue'), result, '\n')
# check first line of transcripts
sql_text = 'SELECT * FROM transcripts LIMIT 1'
result = conn.execute(sql_text).fetchall()
print(colored('1st line table transcripts:\n', 'blue'), result, '\n')
# + [markdown] id="eOt0DyPkM3Or"
# ___
# <span style="color:green">**3. Optimize your database by creating indexes and primary keys on appropriate columns**</span>
#
# As the ted talk URL is unique and also available in both tables (it defines the join), these will be the primary keys. I don't see any need to index any other columns than the primary keys.
#
# Unfortunately sqlite only allows added primary key on creating table, so I have to copy the contents to a new table.
# + [markdown] id="xboO9-6EM3Ot"
# During this exercise it appeared that 3 shows where duplicated in the transcripts, preventing unique keys. The following duplicates have been removed from the original __*transcripts.csv*__:
# - brene_brown_listening_to_shame
# - jonathan_haidt_humanity_s_stairway_to_self_transcendence
# - rob_reid_the_8_billion_ipod
#
# The result has been saved to the file __*transcripts_unique.csv*__ and can be found [here](https://github.com/pvdwijdeven/ML_eduonix/blob/master/Eduonix%20Edegree/4.Complete%20Guide%20to%20Machine%20Learning%20using%20Python/Project/transcripts_unique.csv)
# + id="WCTiwooPM3Ow" outputId="764564aa-f438-44ef-f904-bc1a17e44f61"
sql_text = 'ALTER TABLE ted_main RENAME TO ted_main_old;'
conn.execute(sql_text)
sql_text = 'CREATE TABLE ted_main ('
sql_text += ' comments BIGINT,'
sql_text += ' description TEXT,'
sql_text += ' duration BIGINT,'
sql_text += ' event TEXT,'
sql_text += ' film_date BIGINT,'
sql_text += ' languages BIGINT,'
sql_text += ' main_speaker TEXT,'
sql_text += ' name TEXT,'
sql_text += ' num_speaker BIGINT,'
sql_text += ' published_date BIGINT,'
sql_text += ' ratings TEXT,'
sql_text += ' related_talks TEXT,'
sql_text += ' speaker_occupation TEXT,'
sql_text += ' tags TEXT,'
sql_text += ' title TEXT,'
sql_text += ' url TEXT PRIMARY KEY,'
sql_text += ' views BIGINT'
sql_text += ');'
conn.execute(sql_text)
sql_text = 'INSERT INTO ted_main SELECT * FROM ted_main_old;'
conn.execute(sql_text)
sql_text = 'DROP TABLE ted_main_old'
conn.execute(sql_text)
sql_text = 'CREATE INDEX url_index1 ON ted_main (url)'
conn.execute(sql_text)
meta = MetaData()
table = Table('ted_main', meta, autoload=True, autoload_with=engine)
result = table.primary_key.columns.values()[0].name
print(colored('Primary key of table ted_main:\n', 'blue'), result, '\n')
insp = reflection.Inspector.from_engine(engine)
print(colored('Index of table ted_main:', 'blue'))
for index in insp.get_indexes('ted_main'):
print(index)
sql_text = 'ALTER TABLE transcripts RENAME TO transcripts_old;'
conn.execute(sql_text)
sql_text = 'CREATE TABLE transcripts ('
sql_text += ' transcript TEXT,'
sql_text += ' url TEXT PRIMARY KEY'
sql_text += ');'
conn.execute(sql_text)
sql_text = 'INSERT INTO transcripts SELECT * FROM transcripts_old WHERE url LIKE "http%";'
conn.execute(sql_text)
sql_text = 'DROP TABLE transcripts_old'
conn.execute(sql_text)
sql_text = 'CREATE INDEX url_index2 ON transcripts (url)'
conn.execute(sql_text)
table = Table('transcripts', meta, autoload=True, autoload_with=engine)
result = table.primary_key.columns.values()[0].name
print(colored('Primary key of table transcripts:\n', 'blue'), result, '\n')
insp = reflection.Inspector.from_engine(engine)
print(colored('Index of table transcripts:', 'blue'))
for index in insp.get_indexes('transcripts'):
print(index)
# + [markdown] id="En8nZ5DVM3Oy"
# ___
# <span style="color:green">**4.Add a new column of data that has not been included in the csv**</span>
#
# Here I have chosen to create a category out of the review column. The category with the highest counts in the reviews column is being assigned.
# + id="subLjcmhM3Oz" outputId="98f133b0-c37e-4dc3-8e49-63081bad85c2"
# Adding the column
sql_text = "ALTER TABLE ted_main ADD category TEXT;"
result = conn.execute(sql_text)
print(colored('Column "category" added', 'blue'))
# + id="GGqMQVMrM3O0" outputId="b38ecd71-f4b7-458d-c954-a4ec5132cf6d"
def max_count(reviews):
reviews = reviews[2:-2]
l_reviews = reviews.split("}, {")
max = 0
category = "DUMMY"
for review in l_reviews:
detail = review.split(",")
count = int(detail[2][detail[2].find(':') + 2:])
if count > max:
category = detail[1][detail[1].find(':') + 2:]
max = count
return category
counter = 0
sql_text = 'SELECT url, ratings FROM ted_main'
result = conn.execute(sql_text).fetchall()
for row in result:
cat = max_count(row[1])
sql_text = "UPDATE ted_main SET category = " \
+ cat + " WHERE url = '" + row[0] + "';"
conn.execute(sql_text)
counter += 1
print(colored('Updated ' + str(counter) + ' rows', 'blue'))
sql_text = 'SELECT category FROM ted_main'
result = conn.execute(sql_text).fetchall()
print(colored('\nList of all most counted categories:', 'blue'))
category_set = set()
for category in result:
category_set.add(category.values()[0])
for category in category_set:
print(category)
# + [markdown] id="SesNly3ZM3O1"
# ___
# <span style="color:green">**5.Conduct any type of NLP (language detection, sentiment analysis, tokenization, etc.) and add to the database**</span>
#
# First the query is read into a dataframe and split into a test/train set. Then a vectorizer is being used (with remal of the stopwords), followed by a TFIDF transformation. An ADABoost with a multinomial Naive Bains as base is being used to train the model.
# + id="Ji56qr03M3O1" outputId="e23218e0-6efd-4d83-af0e-c5e33d2691a2"
sql_text = "SELECT ted_main.category, transcripts.transcript FROM ted_main " \
+ "INNER JOIN transcripts ON ted_main.url = transcripts.url;"
dataset = pd.read_sql_query(sql_text, conn)
train, test = train_test_split(dataset, test_size=0.2, random_state=0)
count_vect = CountVectorizer(stop_words='english')
X_train_counts = count_vect.fit_transform(
train.transcript) # X_train_counts.shape
tf_transformer = TfidfTransformer(use_idf=False).fit(X_train_counts)
X_train_tfidf = tf_transformer.transform(X_train_counts)
model = AdaBoostClassifier(base_estimator=MultinomialNB(alpha=0.0025))
model.fit(X_train_tfidf, train.category)
X_test_counts = count_vect.transform(test.transcript)
X_test_tfidf = tf_transformer.transform(X_test_counts)
predicted = model.predict(X_test_tfidf)
print(colored('Accuracy score:', 'blue'))
print(accuracy_score(test.category, predicted))
# + [markdown] id="K-wdgtSiM3O2"
# The accuracy score is around **59%**, which seems not too bad. This might very well be due to the fact that the categories e.g. "inspirational", "funny", "informative", "confusing" are not only based on single words, but also actual content and of course the way the presentation is being held.
#
# Optimization of the algorithm has been done by changing the alpha factor and removing the stop-words.
#
# The results of the predictions vs actuals can be found below:
# + id="sb1GJcyLM3O2" outputId="b46ba10c-2dfd-4b3b-e44c-12cf9a149941"
for x,y in zip(test.category, predicted):
if x == y:
print (colored('{:<20}'.format(x) + y, 'green'))
else:
print (colored('{:<20}'.format(x) + y, 'red'))
# + [markdown] id="RZ4y25zDM3O3"
# A confusionmatrix gives more insight in the results.
# + id="WhyS3vxWM3O4" outputId="2d3d13ce-1645-4004-fe99-9bf1c9edab6d"
from sklearn.metrics import confusion_matrix
labels = list(category_set)
labels.sort()
cm = confusion_matrix(test.category, predicted, labels=labels)
fig = plt.figure(figsize=(20, 20 ))
ax = fig.add_subplot(111)
cax = ax.matshow(cm)
plt.title('Confusion matrix of the category')
plt.locator_params(nbins=len(labels))
fig.colorbar(cax)
ax.set_xticklabels([''] + labels)
ax.set_yticklabels([''] + labels)
plt.xlabel('Predicted')
plt.ylabel('True')
plt.show()
# + [markdown] id="pqJ75tPdM3O4"
# It appears that especially the categories **informative** and **inspiring** score quite well. These are also the best scored categories in the reviews. So what happens if we only look at these? Let's try!
# + id="QSEtqm1ZM3O6" outputId="4d07a867-f7dd-4ea3-cb41-c2b434c9e9dd"
sql_text = "SELECT ted_main.category, transcripts.transcript FROM ted_main " \
+ "INNER JOIN transcripts ON ted_main.url = transcripts.url WHERE ted_main.category IN ('Inspiring','Informative');"
dataset = pd.read_sql_query(sql_text, conn)
train, test = train_test_split(dataset, test_size=0.2, random_state=0)
count_vect = CountVectorizer(stop_words='english')
X_train_counts = count_vect.fit_transform(
train.transcript) # X_train_counts.shape
tf_transformer = TfidfTransformer(use_idf=False).fit(X_train_counts)
X_train_tfidf = tf_transformer.transform(X_train_counts)
model = AdaBoostClassifier(base_estimator=MultinomialNB(alpha=0.1))
model.fit(X_train_tfidf, train.category)
X_test_counts = count_vect.transform(test.transcript)
X_test_tfidf = tf_transformer.transform(X_test_counts)
predicted = model.predict(X_test_tfidf)
print(colored('Accuracy score:', 'blue'))
print(accuracy_score(test.category, predicted))
labels = ['Inspiring', 'Informative']
cm = confusion_matrix(test.category, predicted, labels=labels)
fig = plt.figure()
ax = fig.add_subplot(111)
cax = ax.matshow(cm)
plt.title('Confusion matrix of the category')
plt.locator_params(nbins=len(labels))
fig.colorbar(cax)
ax.set_xticklabels([''] + labels)
ax.set_yticklabels([''] + labels)
plt.xlabel('Predicted')
plt.ylabel('True')
plt.show()
# + [markdown] id="QACm-aeaM3O8"
# As expected the accuracy is much better now and also the confusion matrix is satisfying. It appears that for these categories the kinds of words used do actually indicate whether a talk is inspiring or informative!
# + [markdown] id="kKQu8W1LM3O9"
# ___
# <span style="color:green">**6.Connect to database from Python, select some columns database and produce a visualization/analysis of it.**</span>
#
# Of course I have been connected to the database all the time as I used sqlalchemy to create and manipulate the sqlite database.
#
# For the visualisation I decided to show the relation between number of views and the number of times the talk was rated as "Funny".
#
# To be sure that the funny count makes sense, the funny count is converted to a percentage of the total counts. To prevent a skewed result, reviews with a total review count of less than 1000 have been omitted.
# + id="5_ve9dTbM3O-" outputId="d12780d4-9a8c-4a0f-b2fc-11c5253f3682"
# Adding the columns
sql_text = "ALTER TABLE ted_main ADD total_reviews BIGINT;"
result = conn.execute(sql_text)
print(colored('Columns "total_review" added', 'blue'))
sql_text = "ALTER TABLE ted_main ADD funny_count BIGINT;"
result = conn.execute(sql_text)
print(colored('Columns "funny_count" added', 'blue'))
sql_text = "ALTER TABLE ted_main ADD insp_count BIGINT;"
result = conn.execute(sql_text)
print(colored('Columns "insp_count" added', 'blue'))
# + id="boUbIUUiM3O-" outputId="82b14fd2-2596-4ac8-b40a-70393a9132a7"
def funny_total_parser(reviews):
reviews = reviews[2:-2]
l_reviews = reviews.split("}, {")
total_count = 0
funny_count = 0
insp_count = 0
for review in l_reviews:
detail = review.split(",")
count = int(detail[2][detail[2].find(':') + 2:])
total_count += count
category = detail[1][detail[1].find(':') + 2:]
if category == "'Funny'":
funny_count = count
if category == "'Inspiring'":
insp_count = count
return str(total_count), str(funny_count), str(insp_count)
counter = 0
sql_text = 'SELECT url, ratings FROM ted_main'
result = conn.execute(sql_text).fetchall()
for row in result:
total_count, funny_count, insp_count = funny_total_parser(row[1])
sql_text = "UPDATE ted_main SET total_reviews = " \
+ total_count + ", funny_count = " \
+ funny_count + ", insp_count = " \
+ insp_count + " WHERE url = '" + row[0] + "';"
conn.execute(sql_text)
counter += 1
print(colored('Updated ' + str(counter) + ' rows', 'blue'))
sql_text = 'SELECT total_reviews, published_date, ROUND(funny_count/(total_reviews*1.0)*100)' \
+ ' AS funny_factor, ROUND(insp_count/(total_reviews*1.0)*100)' \
+ ' AS insp_factor FROM ted_main WHERE total_reviews > 1000;'
query_dataset = pd.read_sql_query(sql_text, conn)
fig, ax = plt.subplots()
ax.plot(query_dataset.total_reviews,
query_dataset.funny_factor, 'm.', alpha=0.3)
ax.set(xlabel='Number of views', ylabel='Funny factor [%]',
title='TED Talks - Funny factor vs #views')
ax.grid()
plt.show()
# + [markdown] id="x5IGauWEM3O_"
# Well this is disappointing. I was hoping that the funnier people think the Ted Talk is, the more views there are, but it is easy to see that this is not true.
#
# As Ted Talks claims to be inspirational, my hope was on the inspirational review count, resulting in the plot below:
# + id="rSLqzvibM3PA" outputId="46969588-be22-4695-ccbe-3ce58ba089ec"
fig, ax = plt.subplots()
ax.plot(query_dataset.total_reviews,
query_dataset.insp_factor, 'b.', alpha=0.3)
ax.set(xlabel='Number of views', ylabel='Inspirational factor [%]',
title='TED Talks - Inspirational factor vs #views')
ax.grid()
plt.show()
# + [markdown] id="LjCU9vbAM3PB"
# Also here you see that the review count "inspirational" is not directly related to the number of views. Of course the number of views is also depending on the published date: the earlier it is published, the more time there has been to view the show...
# + id="cBRQExkyM3PC" outputId="e15dc57c-4b58-483f-b9be-be9310fb08fb"
query_dataset['published_year'] = query_dataset['published_date']. \
apply(lambda x: datetime.datetime.fromtimestamp(int(x)).strftime('%Y'))
fig, ax = plt.subplots()
ax.plot(query_dataset.total_reviews,
query_dataset.published_year, 'go', alpha=0.3)
ax.set(xlabel='Number of views', ylabel='Published year',
title='TED Talks - Published year vs #views')
ax.grid()
plt.show()
# + [markdown] id="dRcyaNPoM3PD"
# ___
# Well that is still disappointing... Let's take a look at the correlation matrix:
# + id="rRl7h3LYM3PD" outputId="671cf0f8-1db0-417a-e1f7-cbaf48180b60"
corr_matrix = query_dataset.corr()
print (corr_matrix)
# + [markdown] id="jrqPoVqGM3PG"
# Here we can see that the correlation coefficients between the data shown in plots above are quite low. One of the bigger reasons might be that once a show has a certain number of views, people are triggered to view the show, simply because of the number of views, which in turn increases even more...
#
# The top 10 most viewed TED talks below show some of the researched parameters. I guess I just have to watch them to know why they are so popular....
# + id="mmzPXRmjM3PG" outputId="951c709e-81c2-4de1-f3b3-1c7e697910b6"
sql_text = 'SELECT title, category, ROUND(funny_count/(total_reviews*1.0)*100)' \
+ ' AS funny_factor, ROUND(insp_count/(total_reviews*1.0)*100)' \
+ ' AS insp_factor FROM ted_main ORDER BY views DESC LIMIT 10'
result = conn.execute(sql_text).fetchall()
query_dataset = pd.read_sql_query(sql_text, conn)
print(query_dataset)
# + [markdown] id="e2wPwbxyM3PH"
# ___
# As the final project description requests to use a ML form that has not been used in the class so far, I tend to add 1 more investigation: predict the classification by means of the number of views.
#
# As far as I can see the following complies to all the project demands:
#
# **- Find a dataset to conduct any form of machine learning we have NOT discussed in class**
#
# **- Using any form of trainin/testing method we have discussed in class**
#
# **- Train the model using appropriate methods and share results.**
# + id="PX-Wb_9RM3PI" outputId="eacc3faa-7f7d-468b-d8b6-2a5786034ec9"
sql_text = "SELECT category, views, comments FROM ted_main;"
dataset = pd.read_sql_query(sql_text, conn)
train, test = train_test_split(dataset, test_size=0.2, random_state=0)
clf = AdaBoostClassifier(base_estimator=MultinomialNB(alpha=1))
clf.fit(train[['views', 'comments']], train['category'])
predicted = clf.predict(test[['views', 'comments']])
print(colored('Accuracy score:', 'blue'))
print(accuracy_score(test.category, predicted))
labels = list(category_set)
labels.sort()
cm = confusion_matrix(test.category, predicted, labels=labels)
fig = plt.figure(figsize=(20, 20))
ax = fig.add_subplot(111)
cax = ax.matshow(cm)
plt.title('Confusion matrix of the category')
plt.locator_params(nbins=len(labels))
fig.colorbar(cax)
ax.set_xticklabels([''] + labels)
ax.set_yticklabels([''] + labels)
plt.xlabel('Predicted')
plt.ylabel('True')
plt.show()
# + [markdown] id="e3SRdaLDM3PJ"
# Here again we see a higher score for informative and inspiring, but you can also see that the predictions are quite random around these 2 categories. Changing parameters did not really help. I tried other models (e.g. scaled vector machine classifiers always ended with category "Inspiring" for every prediction. I guess this is the best I can do for the relation between number of views and the category...
# + [markdown] id="hxxjZiSQM3PL"
# **Conclusion**: The best result so far for predicting the review category was using NLP on the contents of the show.
# + [markdown] id="wveynPTnM3PL"
# ___
# That's it, now close the SQL connection...
# + id="y8gYhN6dM3PM"
# close sql connection
conn.close()
| Edu/Module4/Project4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# #### _Speech Processing Labs 2021: SIGNALS 2: Moving Average as Rectangular Filter (Extension)_
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import cmath
from math import floor
from matplotlib.animation import FuncAnimation
from IPython.display import HTML
plt.style.use('ggplot')
from dspMisc import *
# -
# # The Moving Average Filter as a Rectangular Filter
#
# <div class="alert alert-success">
# <em>This notebook illustrates how we can use the convolution theorem understand why the moving average type filters act like low pass filters, and also the connection with leakage in the DFT magnitude response we saw previously. <b>This is optional extension material</b>.
#
# If you want to see an example of the convolution theorem working in the frequency domain, you can just run the code and have a look at the graph at the end entitled 'Frequency response after applying 5-point weighted average filter'</em>
# </div>
#
# ### Learning Outcomes
# * Understand how the moving average filter relates to a rectangular FIR filter
#
# ### Need to know
# * Topic Videos: Spectral Envelope, Filter, Impulse Train
# * [FIR filters](./signals-2-2-fir-filters.ipynb)
#
# <div class="alert alert-warning">
# <strong>Equation alert</strong>: If you're viewing this on github, please note that the equation rendering is not always perfect. You should view the notebooks through a jupyter notebook server for an accurate view.
# </div>
#
# The convolution theorem tells us that, if we know the frequency response of an FIR filter, we know how it will affect the frequency response of it's input (we just multiply the individual frequency response together).
#
# To understand what filter frequency response will look like, it's helpful to first observe that our unweighted moving average filter is pretty much a rectangular window function. It's easy to see what this means when we plot it. The following function allows us to generate rectangular functions:
def gen_rect_window(start_index, end_index, sample_rate=64, seq_length=64):
nsteps = np.array(range(seq_length))
t_s = 1/sample_rate
time_steps = t_s * nsteps
## Let's make a rectangular window
x_rect = np.zeros(seq_length)
x_rect[start_index:end_index] = 1
return x_rect, time_steps
# ## A noisy sample
#
# As an example, let's make a noisy sample `x_noisy`. This is made up of 4 Hz and 24 Hz cosine functions, which a sampling rate of `f_s=64` Hz. We take `N=64` samples as input.
# +
## Set the number of samples N, sampling rate f_s (hence sampling time t_s)
N=64
f_s = 64
t_s = 1/f_s
print("sampling rate: f_s = %f\nsampling time: t_s: %f" % (f_s, t_s))
## make some sinusoids:
## Since the sample rate and sequence length is the same, the generated time steps will match for
## x1 and x2
x1, time_steps = gen_sinusoid(frequency=4, phase=0, amplitude=1, sample_rate=f_s, seq_length=N, gen_function=np.cos)
x2, time_steps = gen_sinusoid(frequency=24, phase=0, amplitude=1, sample_rate=f_s, seq_length=N, gen_function=np.cos)
# add them up!
x_noisy = x1 + x2
## Plot the compound sinusoid we've just created:
fig, timedom = plt.subplots(figsize=(16, 4))
timedom.plot(time_steps, x_noisy, color='magenta')
timedom.set_xlabel('Time (s)')
timedom.set_ylabel('Amplitude')
# -
# ### Apply a moving average filter to this input
#
# +
## The 5-point moving average coefficients
h_avg = np.array([1/5, 1/5, 1/5, 1/5, 1/5])
## Apply this to x_noisy
y_avg = fir_filter(x_noisy, h_avg)
## Plot the filter outputs
fig, timedom = plt.subplots(figsize=(16, 4))
## The original "noisy" input
timedom.plot(time_steps, x_noisy, color='magenta', label='input x_noisy')
timedom.scatter(time_steps, x_noisy, color='magenta')
## The 5-point moving average
timedom.plot(time_steps, y_avg, color='blue', label='unweighted average: y_avg')
timedom.scatter(time_steps, y_avg, color='blue')
timedom.legend()
timedom.set_xlabel('Time (s)')
timedom.set_ylabel('Amplitude')
# -
# ### Now, we make a rectangular window
# +
## Make rectangular window
N=64
K=16
f_s=64
start_index=24
end_index=start_index+K
x_rect, time_steps = gen_rect_window(start_index=start_index, end_index=end_index, sample_rate=f_s, seq_length=N)
fig, timedom = plt.subplots(figsize=(16, 4))
timedom.scatter(time_steps, x_rect, color='magenta')
timedom.plot(time_steps, x_rect, color='magenta')
timedom.set_xlabel('Time (s)')
timedom.set_ylabel('Amplitude')
timedom.set_title('a rectangular window')
# -
# You should see a sequence with 64 point where the mmiddle 16 points have value 1 and the rest have value 0 (i.e., it looks like a rectangle in the middle).
# ### Now, let's look at the frequency response of the rectangular window
# +
## Now we do the DFT on the rectangular function:
## get the magnitudes and phases
mags_rect, phases_rect = get_dft_mag_phase(x_rect, N)
## the DFT output frequencies
dft_freqs_rect = get_dft_freqs_all(f_s, N)
## let's just look at the magnitudes
fig, fdom = plt.subplots(figsize=(16, 4))
fdom.set(xlim=(-1, N/2))
fdom.scatter(dft_freqs_rect, mags_rect)
fdom.set_xlabel("Frequency (Hz)")
fdom.set_ylabel('Magnitude')
fdom.set_title('Frequency response of rectangular window')
## Looks leaky!
# -
# ### Leaky windows?
#
# The plot of the frequency magnitude response of our rectangular window has the hallmarks of leakiness. That is, the frequency response looks scalloped, with the biggest peak occuring around 0Hz. That is, it looks like a low pass filter!
#
# With a bit of algebra we can derive the frequency for any $m$ (not just the DFT output bins indices) to be the following:
#
# If $x[n]$ is a rectangular function of N samples with $K$ continugous samples of value 1 (starting at index $n_0$), we can figure out what the DFT output will be:
#
# $$X[m] = e^{i(2\pi m/N)(n_0-(K-1)/2)} . \frac{\sin(2\pi mK/2N)}{\sin(2\pi m /2N)}$$
#
# This is called the **Dirichlet kernel**. It has the **sinc** shape we saw when we looked at spectral leakage.
#
# How is this useful? Since we know what the frequency response of a rectangular window is, we know what convolving this with different input sequences will look like in the frequency domain. We just multiply the frequency magnitude responses together.
#
# <div class="alert alert-success">
# On a more general note, this sort of convolution with a (short) window is how we do frequency analysis of speech: we taking windows of speech (aka frames) through time and and apply the DFT to get a frequency response. A rectangular window is the simplest type of window we can take. The equation above tells us that the sinc shaped response is an inherent part of using this sort of window. In fact, we can use other window types (e.g. Hanning) to make the main lobes shaper and the sidelobes flatter, but we never really get away from this sinc shape in real world applications. This is a key component of this soft of <strong>short term analysis</strong>.
#
# </div>
#
# Let's write this up in a function:
def gen_rect_response(n_0, K, N, stepsize=0.01, polar=True, amplitude=1):
ms = np.arange(0.01, N, stepsize)
qs = 2*np.pi*ms/N
## Infact, we can work the frequency response to be the Dirichlet Kernel:
response = (np.exp(-1j*qs*(n_0-(K-1)/2)) * np.sin(qs*K/2))/np.sin(qs/2)
if polar:
response_polar = [cmath.polar(z) for z in response]
mags = np.array([m for m, _ in response_polar]) * amplitude
phases = np.array([ph if round(mag) > 0 else 0 for mag, ph in response_polar])
return (mags, phases, ms)
return response, ms
# Now we can plot the dirichlet kernel with the leaky looking DFT magnitudes we calculated earlier for our rectangular window.
# +
## Overlay the dirichlet kernel onto the DFT magnitudes we calculated earlier
## You should be able to see that the DFT magnitudes appear as discrete samples of the Dirichlet Kernel
mags_rect, phases_rect = get_dft_mag_phase(x_rect, N)
mags_rect_sinc , _ , ms = response = gen_rect_response(start_index, K, N)
fig, ax = plt.subplots(figsize=(16, 4))
ax.scatter(dft_freqs_rect, mags_rect, label='rectangular window')
ax.plot((f_s/N)*ms, mags_rect_sinc, color='C2', label='dirichlet')
ax.set(xlim=(-1,N/2))
ax.set_xlabel('Frequency (Hz)')
ax.set_ylabel('Magnitude')
ax.set_title('Frequency response of a rectangular sequence, %d samples with %d contiguous ones' % (N, K))
# -
# You should be able to see that the DFT magnitudes appear as discrete samples of the sinc shaped Dirichlet Kernel
#
#
#
# ### The unweighted average filter as a rectangular function
#
# We can think of our 5-point unweighted average filter as a 5-point input sequence with all values set to 1/5. We can then deduce that the frequency response of the filter will have the same shape as the frequency response of a rectangular window of all ones, but scaled down by 1/5.
#
# Now let's check:
# +
N_h=5
f_s=64
start_index=0
end_index=N_h - start_index
## A 5 point rectangular window of all ones
h_avg, time_steps = gen_rect_window(start_index=start_index, end_index=end_index, sample_rate=f_s, seq_length=N_h)
h_avg = h_avg/N_h
fig, td = plt.subplots(figsize=(16, 4))
td.scatter(time_steps, h_avg, color='magenta')
td.plot(time_steps, h_avg, color='magenta')
td.set_xlabel('Time (s)')
td.set_ylabel('Amplitude')
td.set_title('5 point unweighted average as a rectangular function')
## Not very exciting looking!
print("h_avg:", h_avg)
# -
# You should just see 5 point in a row, all with value 1/5. Now, we can plot the DFT magnitude response, as well as it's idealized continuous version:
# +
## Get the frequency magnitude response for our rectangular function
mags_h_avg, phases_h_avg = get_dft_mag_phase(h_avg, N_h)
## Get the continuous
rect_mags_h_avg, _ , ms = gen_rect_response(start_index, N_h, N_h, amplitude=np.max(h_avg))
## x-axis as frequencies rather than indices
ms_freqs_h_avg = (f_s/N_h) * ms
dft_freqs_h_avg = (f_s/N_h) * np.arange(N_h)
## Plot the frequency magnitude response
fig, fd = plt.subplots(figsize=(16, 4))
fd.set(xlim=(-1, N/2))
fd.scatter(dft_freqs_h_avg, mags_h_avg)
fd.set_xlabel('Frequency (Hz)')
fd.set_ylabel('Magnitude')
fd.set_title('Frequency response of 5-point unweighter average filter')
#fd.scatter(dft_freqs_rect, mags_rect)
fd.plot(ms_freqs_h_avg, rect_mags_h_avg, color="C2")
# -
# You should see $floor(N/2) = 2$ points, with a main lobe peaking at 0 Hz, and side lobes peaking between each of the DFT output frequencies.
# So, DFT frequencies sit exactly at the zeros of this function when the windown size K is the same as the number of samples.
#
# ### Matching the filter and input size with zero padding
#
# The theorem we saw above told us that we could calculate the frequency response of applying the FIR filter to an input sequence (via convolution), but multiply the DFT outputs of the filter and the input sequence.
#
# Now, the x-axis range matches our that of our noisy input sequence because that is determined by the sampling rate. However, the filter frequency response we have above only has 5 outputs, while our input sample size was 64 because the number of DFT outputs is determined by the number of samples we put into the DFT.
#
# To get things in the right form, we need to do some **zero padding** of the filter. We'll see that this basically gives us more samples of the Dirichlet Kernel corresponding to the filter frequency response.
#
#
# +
N=64
K=5
f_s=64
start_index=0
end_index=K
## Make a rectangular filter: K ones at the start
h_avg_pad, time_steps = gen_rect_window(start_index=start_index, end_index=end_index, sample_rate=f_s, seq_length=N)
## Divide by K to make it an average
h_avg_pad = h_avg_pad/K
## Plot the filter
fig, td = plt.subplots(figsize=(16, 4))
td.scatter(time_steps, h_avg_pad, color='magenta')
td.plot(time_steps, h_avg_pad, color='magenta')
td.set_xlabel('Time (s)')
td.set_title('5 point unweighted average FIR filter padded with zeros')
#print("N=%d, K=%d, start=%d, end=%d" % (N, K, start_index, end_index))
# +
## Get the frequency magnitude response for our rectangular function
mags_havg, phases_havg = get_dft_mag_phase(h_avg_pad, N)
## Plot the frequency magnitude response
## x-axis as actual frequencies rather that DFT indices
dft_freqs_havg = (f_s/N) * np.arange(N)
fig, fd = plt.subplots(figsize=(16, 4))
fd.set(xlim=(-1,N/2))
fd.scatter(dft_freqs_havg, mags_havg)
fd.set_xlabel('Frequency (Hz)')
fd.set_ylabel('Magnitude')
fd.set_title('Magnitude response of 5-point unweighter average filter zero padded to 64 samples')
# -
# You should be able to see more clearly in the frequency response graph that the zero padding doesn't change doesnt change the basic shape of the filter's frequency response, we just get a finer grained representation in terms of samples (red dots).
# ### Calculate the input and filter frequency responses
# +
## Now let's calculate frequency responses of the original input
mags, phases = get_dft_mag_phase(x_noisy, N)
## ... the filter
mags_filter, phases_filter = get_dft_mag_phase(h_avg_pad, N)
## ... and the filtered output that we calculated above
mags_avg, phases_avg = get_dft_mag_phase(y_avg, N)
## Plot with actual frequencies on the x-axis
dft_freqs = get_dft_freqs_all(f_s, N)
# +
## plot frequency responses
fig, fd = plt.subplots(figsize=(16, 4))
fd.set(xlim=(-1,N/2), ylim=(-1, N))
# DFT(input)
fd.scatter(dft_freqs, mags, color='magenta', label='DFT(input)')
# DFT(filter) * DFT(input)
fd.scatter(dft_freqs, mags_filter*mags, color='blue', label='DFT(filter).DFT(input)')
# DFT(filtered input)
fd.scatter(dft_freqs, mags_avg, color='red', label='DFT(filter*input)')
fd.set_xlabel('Frequency (Hz)')
fd.set_ylabel('Magnitude')
fd.set_title('Frequency response after applying 5-point weighted average filter')
fd.legend()
# -
# You should see that the results from multiplying the DFT magnitudes from the input and the filter (blue) is (more or less) the same as the DFT of applying the filter in th time domain via convolution (red)
#
# * Notice that there are some differences between the results from the time domain application of the filter (red) and the frequency domain multiplication (blue). In particular there appears to be some leakage in the time-domain convolution case, possibly due to floating point errors.
# ### Exercise
#
# * Try changing the frequency of the second cosine component of our compound wave in the code below.
# * Does the amount of attenuation of the high frequency component change as suggested by the DFT of the filter?
# * e.g. try 26 Hz vs 19 Hz
# * What does this tell you about how well this low pass filter get's rid of high frequency noise?
#
#
# +
## Change the frequency of x2
x1, time_steps = gen_sinusoid(frequency=4, phase=0, amplitude=1, sample_rate=f_s, seq_length=N, gen_function=np.cos)
x2, time_steps = gen_sinusoid(frequency=19, phase=0, amplitude=1, sample_rate=f_s, seq_length=N, gen_function=np.cos)
# add them up!
x_noisy = x1 + x2
## Now let's calculate frequency responses of the original input
mags, phases = get_dft_mag_phase(x_noisy, N)
## ... the filter
mags_filter, phases_filter = get_dft_mag_phase(h_avg_pad, N)
## Plot with actual frequencies on the x-axis
dft_freqs = get_dft_freqs_all(f_s, N)
## plot frequency responses
fig, fd = plt.subplots(figsize=(16, 4))
fd.set(xlim=(-1,N/2), ylim=(-1, N))
# DFT(input)
fd.scatter(dft_freqs, mags, color='magenta', label='DFT(input)')
# DFT(filter) * DFT(input)
fd.scatter(dft_freqs, mags_filter*mags, color='blue', label='DFT(filter)*DFT(input)')
fd.set_xlabel('Frequency (Hz)')
fd.set_ylabel('Magnitude')
fd.set_title('Frequency response after applying 5-point weighted average filter')
fd.legend()
# -
# ### Notes
| signals/signals-lab-2/signals-2-4-rectangular-filters-extension.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from scipy import stats
# from sklearn.ensemble import ExtraTreesClassifier
malware_data = pd.read_csv('LargeTrain.csv')
print malware_data.columns
# -
malware_data.info()
for column in malware_data.columns:
print column
X, y = malware_data.iloc[:, 0:1803].values, malware_data.iloc[:, 1804].values
# +
entropy_dict = {}
for column in malware_data.columns:
temp = 0
for val in malware_data[column]:
x = malware_data[column].values
x.stats()
# p = malware_data[column].values.pdf(val)
# temp -= p * log(p, 2)
# entropy_dict[column] = temp
# print entropy_dict
# +
# copied from https://github.com/patrick201/information_value/blob/master/src/information_value.py
import numpy as np
import math
from scipy import stats
from sklearn.utils.multiclass import type_of_target
class WOE:
def __init__(self):
self._WOE_MIN = -20
self._WOE_MAX = 20
def woe(self, X, y, event=1):
'''
Calculate woe of each feature category and information value
:param X: 2-D numpy array explanatory features which should be discreted already
:param y: 1-D numpy array target variable which should be binary
:param event: value of binary stands for the event to predict
:return: numpy array of woe dictionaries, each dictionary contains woe values for categories of each feature
numpy array of information value of each feature
'''
self.check_target_binary(y)
X1 = self.feature_discretion(X)
res_woe = []
res_iv = []
for i in range(0, X1.shape[-1]):
x = X1[:, i]
woe_dict, iv1 = self.woe_single_x(x, y, event)
res_woe.append(woe_dict)
res_iv.append(iv1)
return np.array(res_woe), np.array(res_iv)
def woe_single_x(self, x, y, event=1):
'''
calculate woe and information for a single feature
:param x: 1-D numpy starnds for single feature
:param y: 1-D numpy array target variable
:param event: value of binary stands for the event to predict
:return: dictionary contains woe values for categories of this feature
information value of this feature
'''
self.check_target_binary(y)
event_total, non_event_total = self.count_binary(y, event=event)
x_labels = np.unique(x)
woe_dict = {}
iv = 0
for x1 in x_labels:
y1 = y[np.where(x == x1)[0]]
event_count, non_event_count = self.count_binary(y1, event=event)
rate_event = 1.0 * event_count / event_total
rate_non_event = 1.0 * non_event_count / non_event_total
if rate_event == 0:
woe1 = self._WOE_MIN
elif rate_non_event == 0:
woe1 = self._WOE_MAX
else:
woe1 = math.log(rate_event / rate_non_event)
woe_dict[x1] = woe1
iv += (rate_event - rate_non_event) * woe1
return woe_dict, iv
def woe_replace(self, X, woe_arr):
'''
replace the explanatory feature categories with its woe value
:param X: 2-D numpy array explanatory features which should be discreted already
:param woe_arr: numpy array of woe dictionaries, each dictionary contains woe values for categories of each feature
:return: the new numpy array in which woe values filled
'''
if X.shape[-1] != woe_arr.shape[-1]:
raise ValueError('WOE dict array length must be equal with features length')
res = np.copy(X).astype(float)
idx = 0
for woe_dict in woe_arr:
for k in woe_dict.keys():
woe = woe_dict[k]
res[:, idx][np.where(res[:, idx] == k)[0]] = woe * 1.0
idx += 1
return res
def combined_iv(self, X, y, masks, event=1):
'''
calcute the information vlaue of combination features
:param X: 2-D numpy array explanatory features which should be discreted already
:param y: 1-D numpy array target variable
:param masks: 1-D numpy array of masks stands for which features are included in combination,
e.g. np.array([0,0,1,1,1,0,0,0,0,0,1]), the length should be same as features length
:param event: value of binary stands for the event to predict
:return: woe dictionary and information value of combined features
'''
if masks.shape[-1] != X.shape[-1]:
raise ValueError('Masks array length must be equal with features length')
x = X[:, np.where(masks == 1)[0]]
tmp = []
for i in range(x.shape[0]):
tmp.append(self.combine(x[i, :]))
dumy = np.array(tmp)
# dumy_labels = np.unique(dumy)
woe, iv = self.woe_single_x(dumy, y, event)
return woe, iv
def combine(self, list):
res = ''
for item in list:
res += str(item)
return res
def count_binary(self, a, event=1):
event_count = (a == event).sum()
non_event_count = a.shape[-1] - event_count
return event_count, non_event_count
def check_target_binary(self, y):
'''
check if the target variable is binary, raise error if not.
:param y:
:return:
'''
y_type = type_of_target(y)
if y_type not in ['binary']:
raise ValueError('Label type must be binary')
def feature_discretion(self, X):
'''
Discrete the continuous features of input data X, and keep other features unchanged.
:param X : numpy array
:return: the numpy array in which all continuous features are discreted
'''
temp = []
for i in range(0, X.shape[-1]):
x = X[:, i]
x_type = type_of_target(x)
if x_type == 'continuous':
x1 = self.discrete(x)
temp.append(x1)
else:
temp.append(x)
return np.array(temp).T
def discrete(self, x):
'''
Discrete the input 1-D numpy array using 5 equal percentiles
:param x: 1-D numpy array
:return: discreted 1-D numpy array
'''
res = np.array([0] * x.shape[-1], dtype=int)
for i in range(5):
point1 = stats.scoreatpercentile(x, i * 20)
point2 = stats.scoreatpercentile(x, (i + 1) * 20)
x1 = x[np.where((x >= point1) & (x <= point2))]
mask = np.in1d(x, x1)
res[mask] = (i + 1)
return res
@property
def WOE_MIN(self):
return self._WOE_MIN
@WOE_MIN.setter
def WOE_MIN(self, woe_min):
self._WOE_MIN = woe_min
@property
def WOE_MAX(self):
return self._WOE_MAX
@WOE_MAX.setter
def WOE_MAX(self, woe_max):
self._WOE_MAX = woe_max
# -
print WOE.woe(X, y)
| entropy_perspective.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.3 64-bit (''bigdata'': conda)'
# metadata:
# interpreter:
# hash: ddc5808000e670ee531b6d0df5cab3b50e0bc41afef88fb058cf86665e26377c
# name: python3
# ---
# ## Uso dei Data Cube
#
# Utilizzeremo il pacchetto `cubes` che va installato con `pip` nel nostro ambiente. `cubes` comunica con un backend SQL, detto `Store`, che verrà creato con `sqlalchemy` fornisce l'astrazione necessaria per le creazione di diversi modelli di analisi (decsrizioni json del datacube, delle sue dimensioni, categorie e misure di aggregazione) all'interno di un _workspace_.
#
# 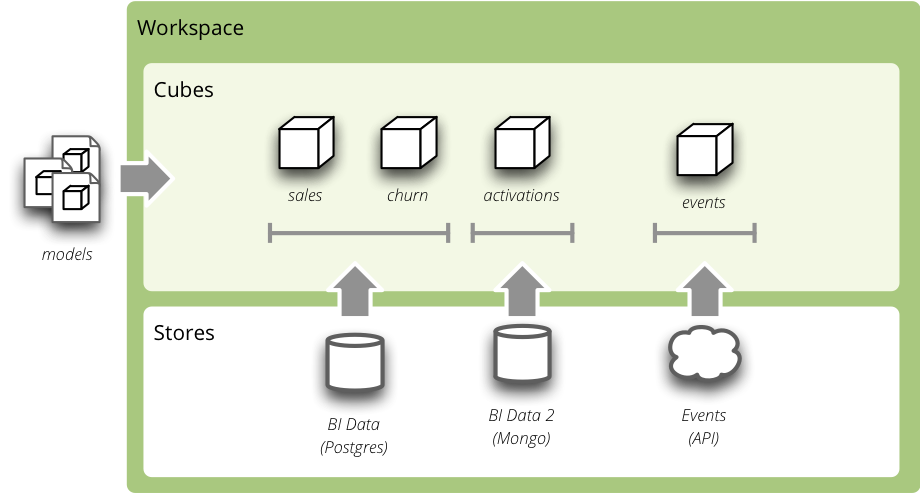
#
#
# La gerarchia delle dimensioni viene sostanziata in dei _mapping_ verso le misure fisiche dei fatti:
#
# 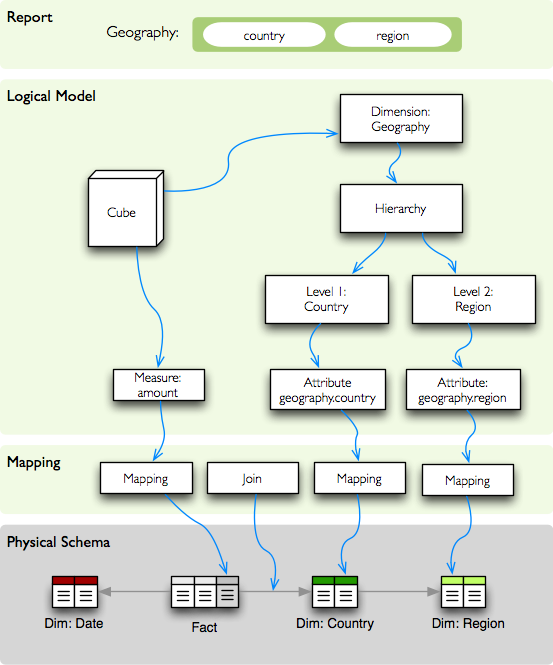
#
# Il modello dei dati è strutturato come un file `json` che ha differenti proprietà mirate a descrivere gli aspetti del modello, dai metadati alla loro corrispondenza fisica con i dati presenti nel database:
#
# 
#
# Segue una breve descrizione
#
# ### Modello logico
#
# - `name`: nome del modello
# - `label`: etichetta opzionale del modello
# - `description`: descrizione estesa opzionale del modello
# - `locale`: etichetta opzionale di localizzazione del modello (ad es. `'en'`, `'it'`, ...)
# - `cubes`: lista dei metadati dei cubi definiti nel modello
# - `dimensions`: lista dei metadati delle dimensioni
# - `public_dimensions`: lista delle dimensioni pubbliche che verrebbero esportate se definite. Tutte le dimensioni sono pubbliche per default
#
# ### Modello fisico
#
# - `store`: nome del data store in cui si trovano i dati: il nome di default è `Default`; si tratta della connessione al database utilizzato
# - `mappings`: dizionario delle corrispondenze tra i nomi logici delle dimensioni e quelli fisici delle colonne nel database che viene ereditata da tutti i cubi del modello
# - `joins`: specifiche di eventuali `join` utilizzati, per esempio con un backend `SQL` nella forma di una lista di `dict` che viene ereditata da tutti i cubi del modello
# - `browser_options`: eventuali opzioni passate al `browser`che è la struttura dati utilizzata per navigare i dati dei cubi.
#
# ### Cubi
#
# Ecco un esempio di cubo, con le proprietà di metadati, di dimensione e fisiche:
#
# ```json
# {
# "name": "sales", // nome del cubo
# "label": "Sales", // etichetta
# "dimensions": [ "date", ... ] // array delle dimensioni come stringhe
#
# "measures": [...], // array delle misure che corrispondono alle colonne del database
# "aggregates": [...], // array delle misure di aggregazione
# "details": [...], // array degli attributi dei fatti che si vogliono eventualmente mostrare
#
# "fact": "fact_table_name", // riferimento esplicito alla tabella dei fatti
# // che dev'essere usata dal backend
# "mappings": { ... }, // mapping di colonne sulla tabella dei fatti
# "joins": [ ... ] // array dei join
# }
# ```
#
# #### Misure di aggregazione
#
# Le funzioni di aggregazione sono gestite dal browser:
#
# 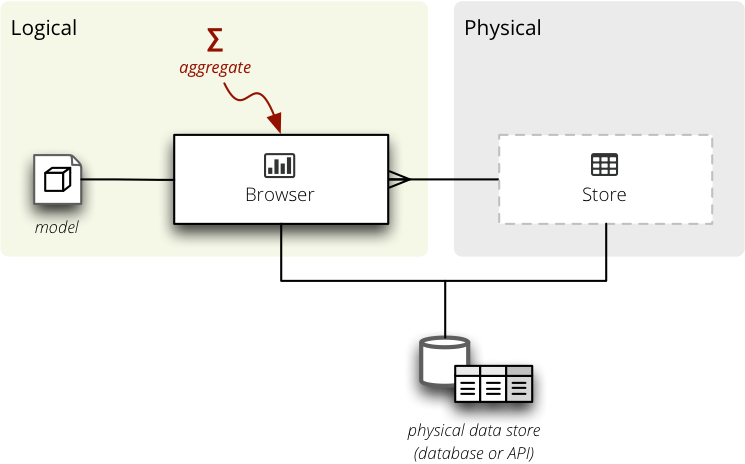
#
#
# Ecco un esempio di definizione nel modello, ma la API consente la definizione di funzioni da parte dell'utente.
#
# ```json
# "aggregates": [
# {
# "name": "amount_sum",
# "label": "Total Sales Amount",
# "measure": "amount",
# "function": "sum"
# },
# {
# "name": "vat_sum",
# "label": "Total VAT",
# "measure": "vat",
# "function": "sum"
# },
# {
# "name": "item_count",
# "label": "Item Count",
# "function": "count"
# }
# ]
# ```
#
# #### Join nel backend SQL
#
# Il beckend `SQL`accetta schemi a stella e snowflake:
#
# 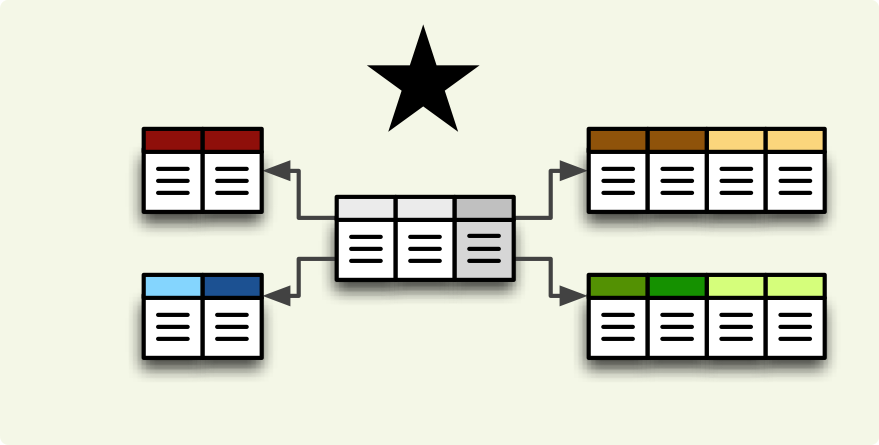 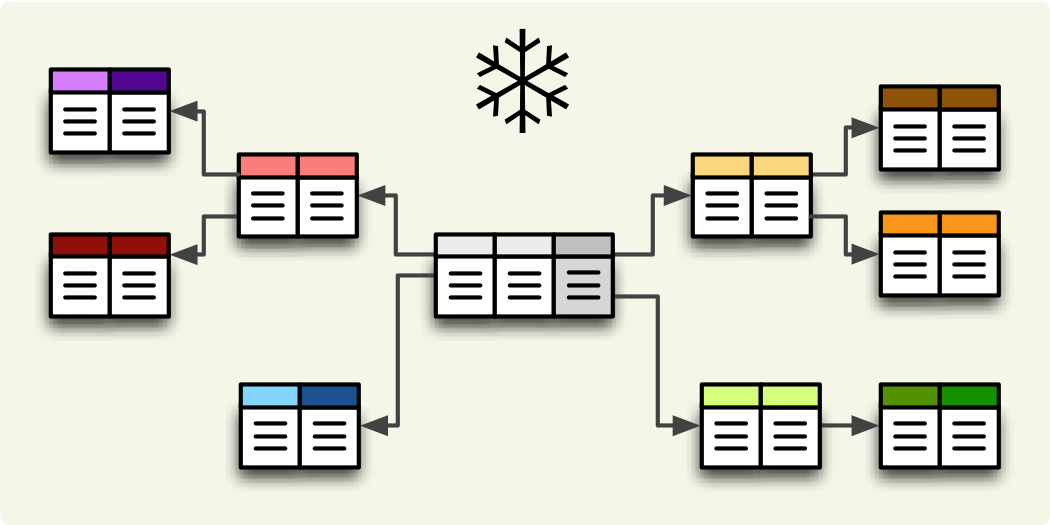
#
# Il join dev'essere specificato esplicitamente per ottenere una rappresentazione tabellare unica di fatti e dimensioni con categorie e sotto-categorie:
#
# 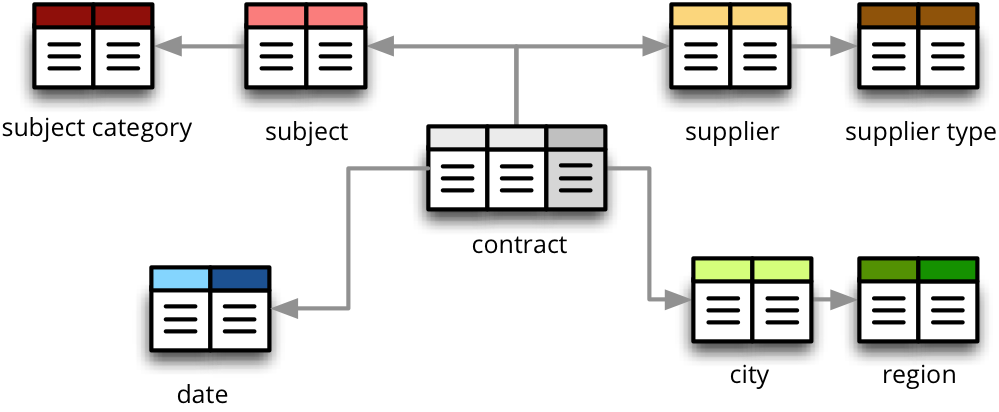
# 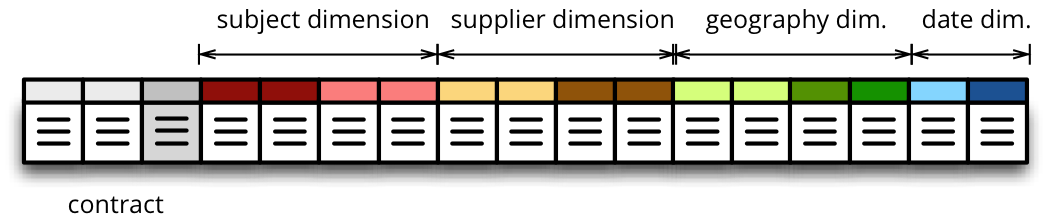
#
# Ecco un esempio di specifica del join:
#
# ```json
# "joins" = [
# {
# "master": "fact_sales.product_id",
# "detail": {
# "schema": "sales",
# "table": "dim_products",
# "column": "id"
# }
# ]
# ```
# +
# importiamo le librerie
import pymysql
from sqlalchemy import create_engine
from cubes.tutorial.sql import create_table_from_csv
from cubes import Workspace
# creiamo l'engine sql e il database fisico
engine = create_engine('mysql+pymysql://mysqluser:mysqlpassword@localhost:3306/data_cube',pool_pre_ping=True)
create_table_from_csv(engine,
"./Data/IBRD_Balance_Sheet__FY2010.csv",
table_name="ibrd_balance",
fields=[
("category", "string"),
("category_label", "string"),
("subcategory", "string"),
("subcategory_label", "string"),
("line_item", "string"),
("year", "integer"),
("amount", "integer")],
create_id=True
)
# -
# Il workspace potrebbe essere creato a partire da un file di configurazione chiamato `slicer.ini` di default e che ha una sintassi simile alla seguente:
#
# ```
# [workspace]
# model = model.json
#
# [store]
# type = sql
# url = postgresql://localhost/database
# ```
# dove viene specificata la coppia store-workspace. In alternativa è possibile inizializzare il workspace direttamente dalla API.
# +
# creiamo il workspace con le API
workspace = Workspace()
workspace.register_default_store("sql", url="mysql+pymysql://mysqluser:mysqlpassword@localhost:3306/data_cube")
# importiamo il modello del data cube che è definito come un file json
# in cui è definito il cubo "ibrd_balance"
workspace.import_model("./Data/tutorial_model.json")
# +
# Creiamo un browser sul data cube per eseguire le operazioni
browser = workspace.browser("ibrd_balance")
# -
# calcoliamo le misure di aggregazione previste dal modello
result = browser.aggregate()
result.summary["record_count"]
result.summary["amount_sum"]
result.aggregates
result = browser.aggregate(drilldown=[("year")])
for record in result:
print(record)
result = browser.aggregate(drilldown=[("item",None,"subcategory")])
for record in result:
print(record)
report=browser.aggregate(drilldown=[('item',None,'category'),'year'])
for record in report:
print(record)
# +
from cubes import Cell, PointCut
cut = [
PointCut('year', [2010]),
PointCut('item', ['l','dl'])
]
cell = Cell(workspace.cube('ibrd_balance'),cut)
report=browser.aggregate(cell,drilldown=[('item',None,'line_item')])
for record in report:
print(record)
# +
cell=Cell(workspace.cube('ibrd_balance'))
cell=cell.drilldown('year',2009)
facts = browser.facts(cell)
for fact in facts:
print(fact)
# +
report = browser.aggregate(cell,drilldown=[('item',None,'category')])
for record in report:
print(record)
report.total_cell_count
| Data Cubes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Affine transforms using scikit-image
# This notebook demonstrates how to apply affine transforms to 3D images.
# + tags=[]
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from skimage.io import imread
# -
# Laod example data
np_array = imread('../../data/Haase_MRT_tfl3d1.tif')
np_array.shape
# To setup an affine transform, you can do this using a 4x4 transform matrix:
transform_matrix = np.asarray([
[1, 0, 0, 50],
[0, 2, 0, 0],
[0, 0, 0.5, 0],
[0, 0, 0, 1]
])
# Scikit-image only supports 2D transforms and thus, we pick a slice to transform it:
# +
# pull image stack from GPU and pick a slice
image = np_array[100]
from skimage.io import imshow
imshow(image)
# -
# We now define an affine transform using scikit-image and apply it to the image.
# +
from skimage import transform as tf
# define transform with #scikit image
transform = tf.AffineTransform(scale=0.5, translation=[10,0])
transformed_image = tf.warp(image, transform.inverse)
imshow(transformed_image)
# -
# ## Interoperability with clesperanto
# Next, we push this single plane image to the GPU and transform it using pyclesperanto
# +
import pyclesperanto_prototype as cle
cle.select_device('RTX')
# +
image_gpu = cle.push(image)
# define transform with #scikit image
from skimage import transform as tf
transform = tf.AffineTransform(scale=0.5, translation=[10,0])
transformed_image = cle.affine_transform(image_gpu, transform=transform)
cle.imshow(transformed_image, color_map="Greys_r")
# -
| docs/19_spatial_transforms/affine_transforms_scikit_image.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import itertools
import json
import operator
import os
from pathlib import Path
from pprint import pprint
import re
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
from matplotlib.lines import Line2D
import numpy as np
import pandas as pd
import seaborn as sns
from scipy import stats
from tqdm.notebook import tqdm
# %matplotlib inline
from IPython.display import set_matplotlib_formats
set_matplotlib_formats('png')
# + [markdown] toc-hr-collapsed=false
# ## Load data and preprocess
# -
# ### Metadata
# +
# Map from test suite tag to high-level circuit.
circuits = {
"Licensing": ["npi", "reflexive"],
"Long-Distance Dependencies": ["fgd", "cleft"],
"Agreement": ["number"],
"Garden-Path Effects": ["npz", "mvrr"],
"Gross Syntactic State": ["subordination"],
"Center Embedding": ["center"],
}
tag_to_circuit = {tag: circuit
for circuit, tags in circuits.items()
for tag in tags}
# +
# Map codenames to readable names for various columns.
def format_pretrained(model_name):
return "%s$^*$" % model_name
PRETTY_COLUMN_MAPS = [
("model_name",
{
"vanilla": "LSTM",
"ordered-neurons": "ON-LSTM",
"rnng": "RNNG",
"ngram": "n-gram",
"random": "Random",
"gpt-2-pretrained": format_pretrained("GPT-2"),
"gpt-2-xl-pretrained": format_pretrained("GPT-2-XL"),
"gpt-2": "GPT-2",
"transformer-xl": format_pretrained("Transformer-XL"),
"grnn": format_pretrained("GRNN"),
"jrnn": format_pretrained("JRNN"),
}),
("corpus", lambda x: x.upper() if x else "N/A"),
]
PRETTY_COLUMNS = ["pretty_%s" % col for col, _ in PRETTY_COLUMN_MAPS]
# -
# Exclusions
exclude_suite_re = re.compile(r"^fgd-embed[34]|^gardenpath|^nn-nv")
exclude_models = ["1gram", "ngram-no-rand"] # "ngram",
# +
ngram_models = ["1gram", "ngram", "ngram-single"]
baseline_models = ["random"]
# Models for which we designed a controlled training regime
controlled_models = ["ngram", "ordered-neurons", "vanilla", "rnng", "gpt-2"]
controlled_nonbpe_models = ["ngram", "ordered-neurons", "vanilla", "rnng"]
# -
# ### Load
ppl_data_path = Path("../data/raw/perplexity.csv")
test_suite_results_path = Path("../data/raw/sg_results")
# +
perplexity_df = pd.read_csv(ppl_data_path, index_col=["model", "corpus", "seed"])
perplexity_df.index.set_names("model_name", level=0, inplace=True)
results_df = pd.concat([pd.read_csv(f) for f in test_suite_results_path.glob("*.csv")])
# Split model_id into constituent parts
model_ids = results_df.model.str.split("_", expand=True).rename(columns={0: "model_name", 1: "corpus", 2: "seed"})
results_df = pd.concat([results_df, model_ids], axis=1).drop(columns=["model"])
results_df["seed"] = results_df.seed.fillna("0").astype(int)
# Add tags
results_df["tag"] = results_df.suite.transform(lambda s: re.split(r"[-_0-9]", s)[0])
results_df["circuit"] = results_df.tag.map(tag_to_circuit)
tags_missing_circuit = set(results_df.tag.unique()) - set(tag_to_circuit.keys())
if tags_missing_circuit:
print("Tags missing circuit: ", ", ".join(tags_missing_circuit))
# +
# Exclude test suites
exclude_filter = results_df.suite.str.contains(exclude_suite_re)
print("Dropping %i results / %i suites due to exclusions:"
% (exclude_filter.sum(), len(results_df[exclude_filter].suite.unique())))
print(" ".join(results_df[exclude_filter].suite.unique()))
results_df = results_df[~exclude_filter]
# Exclude models
exclude_filter = results_df.model_name.isin(exclude_models)
print("Dropping %i results due to dropping models:" % exclude_filter.sum(), list(results_df[exclude_filter].model_name.unique()))
results_df = results_df[~exclude_filter]
# Exclude word-level controlled models with BPE tokenization
exclude_filter = (results_df.model_name.isin(controlled_nonbpe_models)) & (results_df.corpus.str.endswith("bpe"))
results_df = results_df[~exclude_filter]
# Exclude GPT-2 with word-level or SentencePieceBPE tokenization
exclude_filter = ((results_df.model_name=="gpt-2") & ~(results_df.corpus.str.endswith("gptbpe")))
results_df = results_df[~exclude_filter]
# -
# Average across seeds of each ngram model.
# The only difference between "seeds" of these model types are random differences in tie-breaking decisions.
for ngram_model in ngram_models:
# Create a synthetic results_df with one ngram model, where each item is correct if more than half of
# the ngram seeds vote.
ngram_results_df = (results_df[results_df.model_name == ngram_model].copy()
.groupby(["model_name", "corpus", "suite", "item", "tag", "circuit"])
.agg({"correct": "mean"}) > 0.5).reset_index()
ngram_results_df["seed"] = 0
# Drop existing model results.
results_df = pd.concat([results_df[~(results_df.model_name == ngram_model)],
ngram_results_df], sort=True)
# Prettify name columns, which we'll carry through data manipulations
for column, map_fn in PRETTY_COLUMN_MAPS:
pretty_column = "pretty_%s" % column
results_df[pretty_column] = results_df[column].map(map_fn)
if results_df[pretty_column].isna().any():
print("WARNING: In prettifying %s, yielded NaN values:" % column)
print(results_df[results_df[pretty_column].isna()])
# ### Data prep
# +
suites_df = results_df.groupby(["model_name", "corpus", "seed", "suite"] + PRETTY_COLUMNS).correct.mean().reset_index()
suites_df["tag"] = suites_df.suite.transform(lambda s: re.split(r"[-_0-9]", s)[0])
suites_df["circuit"] = suites_df.tag.map(tag_to_circuit)
# For controlled evaluation:
# Compute a model's test suite accuracy relative to the mean accuracy on this test suite.
# Only compute this on controlled models.
def get_controlled_mean(suite_results):
# When computing test suite mean, first collapse test suite accuracies within model--corpus, then combine resulting means.
return suite_results[suite_results.model_name.isin(controlled_models)].groupby(["model_name", "corpus"]).correct.mean().mean()
suite_means = suites_df.groupby("suite").apply(get_controlled_mean)
suites_df["correct_delta"] = suites_df.apply(lambda r: r.correct - suite_means.loc[r.suite] if r.model_name in controlled_models else None, axis=1)
# -
# We'll save this data to a CSV file for access from R, where we do
# linear mixed-effects regression modeling.
suites_df.to_csv("../data/suites_df.csv")
# +
# Join PPL and accuracy data.
joined_data = suites_df.groupby(["model_name", "corpus", "seed"] + PRETTY_COLUMNS)[["correct", "correct_delta"]].agg("mean")
joined_data = pd.DataFrame(joined_data).join(perplexity_df).reset_index()
joined_data.head()
# Track BPE + size separately.
joined_data["corpus_size"] = joined_data.corpus.str.split("-").apply(lambda tokens: tokens[1] if len(tokens) >= 2 else None)
joined_data["corpus_bpe"] = joined_data.corpus.str.split("-").apply(lambda tokens: tokens[2] if len(tokens) > 2 else ("none" if len(tokens) >= 2 else None))
# -
# Join PPL and accuracy data, splitting on circuit.
joined_data_circuits = suites_df.groupby(["model_name", "corpus", "seed", "circuit"] + PRETTY_COLUMNS)[["correct", "correct_delta"]].agg("mean")
joined_data_circuits = pd.DataFrame(joined_data_circuits).reset_index().set_index(["model_name", "corpus", "seed"]).join(perplexity_df).reset_index()
joined_data_circuits.head()
# +
# Analyze stability to modification.
def has_modifier(ts):
if ts.endswith(("_modifier", "_mod")):
return True
else:
return None
suites_df["has_modifier"] = suites_df.suite.transform(has_modifier)
# Mark "non-modifier" test suites
modifier_ts = suites_df[suites_df.has_modifier == True].suite.unique()
no_modifier_ts = [re.sub(r"_mod(ifier)?$", "", ts) for ts in modifier_ts]
suites_df.loc[suites_df.suite.isin(no_modifier_ts), "has_modifier"] = False
# Store subset of test suites which have definite modifier/no-modifier marking
suites_df_mod = suites_df[~(suites_df.has_modifier.isna())].copy()
suites_df_mod["has_modifier"] = suites_df_mod.has_modifier.astype(bool)
# Get base test suite (without modifier/no-modifier marking)
suites_df_mod["test_suite_base"] = suites_df_mod.suite.transform(lambda ts: ts.strip("_no-modifier").strip("_modifier"))
suites_df_mod.head()
# -
# ### Checks
# Each model--corpus--seed should have perplexity data.
ids_from_results = results_df.set_index(["model_name", "corpus", "seed"]).sort_index().index
ids_from_ppl = perplexity_df.sort_index().index
diff = set(ids_from_results) - set(ids_from_ppl)
if diff:
print("Missing perplexity results for:")
pprint(diff)
#raise ValueError("Each model--corpus--seed must have perplexity data.")
# +
# Every model--corpus--seed should have results for all test suite items.
item_list = {model_key: set(results.suite)
for model_key, results in results_df.groupby(["model_name", "corpus", "seed"])}
not_shared = set()
for k1, k2 in itertools.combinations(item_list.keys(), 2):
l1, l2 = item_list[k1], item_list[k2]
if l1 != l2:
print("SyntaxGym test suite results for %s and %s don't match" % (k1, k2))
print("\tIn %s but not in %s:\n\t\t%s" % (k2, k1, l2 - l1))
print("\tIn %s but not in %s:\n\t\t%s" % (k1, k2, l1 - l2))
print()
not_shared |= l2 - l1
not_shared |= l1 - l2
if len(not_shared) > 0:
to_drop = results_df[results_df.suite.isin(not_shared)]
print("Dropping these test suites (%i rows) for now. Yikes:" % len(to_drop))
print(not_shared)
results_df = results_df[~results_df.suite.isin(not_shared)]
else:
print("OK")
# -
# Second sanity check: same number of results per model--corpus--seed
result_counts = results_df.groupby(["model_name", "corpus", "seed"]).item.count()
if len(result_counts.unique()) > 1:
print("WARNING: Some model--corpus--seed combinations have more result rows in results_df than others.")
print(result_counts)
# Second sanity check: same number of suite-level results per model--corpus--seed
suite_result_counts = suites_df.groupby(["model_name", "corpus", "seed"]).suite.count()
if len(suite_result_counts.unique()) > 1:
print("WARNING: Some model--corpus--seed combinations have more result rows in suites_df than others.")
print(suite_result_counts)
# ## Prepare for data rendering
# +
RENDER_FINAL = True
figure_path = Path("../reports/camera_ready_figures")
figure_path.mkdir(exist_ok=True, parents=True)
RENDER_CONTEXT = {
"font_scale": 3.5,
"rc": {"lines.linewidth": 2.5, "hatch.linewidth":3},
"style": "ticks",
"font": "Liberation Sans"
}
sns.set(**RENDER_CONTEXT)
# -
BASELINE_LINESTYLE = {
"color": "gray",
"linestyle": "--",
}
CORPUS_MARKERS = {
"BLLIP-LG": "s",
"BLLIP-MD": "v",
"BLLIP-SM": "P",
"BLLIP-XS": "X",
"BLLIP-LG-BPE": "s",
"BLLIP-MD-BPE": "v",
"BLLIP-LG-GPTBPE": "s",
"BLLIP-MD-GPTBPE": "v",
"BLLIP-SM-GPTBPE": "P",
"BLLIP-XS-GPTBPE": "X"
}
p = sns.color_palette()[:len(joined_data.model_name.unique())]
MODEL_COLORS = {
"GPT-2": p[0],
"LSTM": p[1],
"ON-LSTM": p[2],
"RNNG": p[3],
"n-gram": p[4],
"Random": "darkgrey",
format_pretrained("GPT-2"): "mediumturquoise",
format_pretrained("GPT-2-XL"): p[5],
format_pretrained("Transformer-XL"): "gold",
format_pretrained("GRNN"): p[6],
format_pretrained("JRNN"): "deeppink",
}
def render_final(path):
sns.despine()
plt.tight_layout()
plt.savefig(path)
# Standardize axis labels
SG_ABSOLUTE_LABEL = "SG score"
SG_DELTA_LABEL = "SG score delta"
PERPLEXITY_LABEL = "Test perplexity"
# Establish consistent orderings of model names, corpus names, circuit names
# for figure ordering / coloring. (NB these refer to prettified names)
model_order = sorted(set(results_df.pretty_model_name))
controlled_model_order = ["LSTM", "ON-LSTM", "RNNG", "GPT-2", "n-gram"] #sorted(set(results_df[results_df.model_name.isin(controlled_models)].pretty_model_name))
corpus_order = ["BLLIP-LG", "BLLIP-MD", "BLLIP-SM", "BLLIP-XS",
"BLLIP-LG-BPE", "BLLIP-LG-GPTBPE",
"BLLIP-MD-GPTBPE", "BLLIP-SM-GPTBPE", "BLLIP-XS-GPTBPE"]
corpus_size_order = ["lg", "md", "sm", "xs"]
nobpe_corpus_order = [c for c in corpus_order if "BPE" not in c]
circuit_order = sorted([c for c in results_df.circuit.dropna().unique()])
# ## Reproducing paper figures
# ### Figure 1 (Basic barplot)
# +
f, ax = plt.subplots(figsize=(20, 10))
# Exclude random baseline; will plot as horizontal line
plot_df = suites_df[(suites_df.model_name != "random")]
# Sort by decreasing average accuracy
order = list(plot_df.groupby("pretty_model_name").correct.mean().sort_values(ascending=False).index)
sns.barplot(data=plot_df.reset_index(), x="pretty_model_name", y="correct", order=order, ax=ax, palette=MODEL_COLORS)
# Plot random chance baseline
ax.axhline(suites_df[suites_df.model_name == "random"].correct.mean(), **BASELINE_LINESTYLE)
# Adjust labels and axes
ax.set_xticklabels(ax.get_xticklabels(), rotation=340, horizontalalignment="left")
ax.set_ylim(0,1)
plt.xlabel("Model")
plt.ylabel(SG_ABSOLUTE_LABEL, labelpad=36)
if RENDER_FINAL:
render_final(figure_path / "overall.pdf")
# -
# ### Controlled evaluation of model type + dataset size
controlled_suites_df = suites_df[suites_df.model_name.isin(controlled_models)]
controlled_suites_df_mod = suites_df_mod[suites_df_mod.model_name.isin(controlled_models)]
controlled_joined_data_circuits = joined_data_circuits[joined_data_circuits.model_name.isin(controlled_models)]
# ### Figure 3
# +
_, axes = plt.subplots(nrows=1, ncols=2, sharex=False, sharey=True, figsize=(40,12))
for i, ax in enumerate(axes):
ax.axhline(0, c="gray", linestyle="--")
if i == 0:
kwargs = dict(data=controlled_suites_df.reset_index(), order=controlled_model_order, ax=ax,
x="pretty_model_name", y="correct_delta", palette=MODEL_COLORS)
sns.barplot(**kwargs, units="corpus")
sns.swarmplot(**kwargs, alpha=0.3, size=9)
ax.set_xlabel("Model", labelpad=16)
ax.set_ylabel(SG_DELTA_LABEL)
elif i == 1:
# Estimate error intervals with a structured bootstrap: resampling units = model
kwargs = dict(data=controlled_suites_df.reset_index(), x="pretty_corpus", y="correct_delta", order=nobpe_corpus_order, ax=ax)
sns.barplot(**kwargs, color="Gray", units="pretty_model_name")
sns.swarmplot(**kwargs, hue="pretty_model_name", hue_order=controlled_model_order, palette=MODEL_COLORS, size=9, alpha=0.5)
handles, labels = ax.get_legend_handles_labels()
for h in handles:
h.set_sizes([300.0])
ax.set_xlabel("Corpus", labelpad=16)
ax.set_ylabel("")
ax.legend(handles, labels, loc="upper center", ncol=5, columnspacing=0.3, handletextpad=0.01)
if RENDER_FINAL:
render_final(figure_path / "controlled.pdf")
# -
# ### Figure 4
# +
_, axes = plt.subplots(nrows=1, ncols=2, sharex=True, sharey=True, figsize=(40,15))
legend_params=dict(title="", ncol=5, loc="upper center", columnspacing=1, handlelength=1, handletextpad=0.3)
for i, ax in enumerate(axes):
ax.axhline(0, **BASELINE_LINESTYLE)
if i == 0:
sns.barplot(data=controlled_joined_data_circuits, x="circuit", y="correct_delta",
hue="pretty_model_name", units="corpus", hue_order=controlled_model_order,
ax=ax, palette=MODEL_COLORS)
ax.set_ylabel(SG_DELTA_LABEL)
elif i == 1:
sns.barplot(data=controlled_joined_data_circuits, x="circuit", y="correct_delta",
hue="pretty_corpus", units="model_name", hue_order=nobpe_corpus_order,
ax=ax, palette="Greys_r")
ax.set_ylabel("")
ax.set_xticklabels(ax.get_xticklabels(), rotation=15, ha="right")
ax.set_xlabel("Circuit")
ax.legend(**legend_params)
if RENDER_FINAL:
render_final(figure_path / "controlled_circuit.pdf")
# -
# ### Figure 5
# +
_, ax = plt.subplots(figsize=(40,12))
joined_data_circuits_norandom = joined_data_circuits[joined_data_circuits.pretty_model_name != "Random"]
order = list(plot_df.groupby("pretty_model_name").correct.mean().sort_values(ascending=False).index)
sns.barplot(data=joined_data_circuits_norandom, x="circuit", y="correct",
hue="pretty_model_name", units="corpus", hue_order=order, ax=ax, palette=MODEL_COLORS)
ax.set_xticklabels(ax.get_xticklabels(), rotation=15, ha="right")
ax.set_xlabel("Circuit")
ax.set_ylabel(SG_ABSOLUTE_LABEL)
ax.legend(title="", ncol=int(len(order)/2), loc="upper center", columnspacing=1, handlelength=1, handletextpad=1, bbox_to_anchor=(0.5,1.3))
if RENDER_FINAL:
render_final(figure_path / "allmodels_circuit.pdf")
# -
# ### Figure 6 (Stability to modification)
print("Suites in modification analysis:", controlled_suites_df_mod.suite.unique())
# +
# Sort by decreasing average accuracy.
order = list(plot_df.groupby("pretty_model_name").correct.mean().sort_values(ascending=False).index)
_, ax = plt.subplots(figsize=(20,12))
sns.barplot(data=suites_df_mod, x="pretty_model_name", y="correct", hue="has_modifier", order=order, ax=ax)
# Colors.
sorted_patches = sorted(ax.patches, key=lambda bar: bar.get_x())
colors = [MODEL_COLORS[order[i]] for i in range(len(order))]
for i, bar in enumerate(sorted_patches):
bar.set_facecolor(colors[int(i/2)])
if i % 2 != 0:
bar.set_alpha(0.4)
# Set labels.
ax.set_xlabel("Model", labelpad=16)
ax.set_ylabel(SG_ABSOLUTE_LABEL)
ax.set_xticklabels(ax.get_xticklabels(), rotation=340, horizontalalignment="left")
# Custom legend.
handles, _ = ax.get_legend_handles_labels()
handles[0] = mpatches.Patch(facecolor="k")
handles[1] = mpatches.Patch(facecolor="k", alpha=0.4)
ax.legend(handles, ["No modifier", "With modifier"], loc="upper right", title="")
if RENDER_FINAL:
render_final(figure_path / "stability-all-models.pdf")
# -
# ### Figure 2 (SG score vs perplexity)
# +
# Set limits for broken x-axis to determine proper scaling (ratio of widths).
ax1max = 250
ax2min, ax2max = 520, 540
ax_ratio = ax1max / (ax2max - ax2min)
f, (ax1,ax2) = plt.subplots(1,2,sharey=False,figsize=(19, 20),gridspec_kw={'width_ratios': [ax_ratio, 1]})
sns.despine()
palette = sns.cubehelix_palette(4, reverse=True)
markers = {
"GPT-2": "s",
"RNNG" : "X",
"ON-LSTM" : "v",
"LSTM" : "*",
"n-gram" : "d"
}
for m in joined_data.pretty_model_name.unique():
if m not in markers:
markers[m] = "."
for ax in [ax1,ax2]:
sns.scatterplot(data=joined_data, x="test_ppl", y="correct", hue="corpus_size", hue_order=corpus_size_order,
markers=markers, palette=palette, style_order=model_order,
s=2300, style="pretty_model_name", ax=ax, zorder=2, alpha=0.8)
ax.set_xlabel("")
ax.tick_params(axis='x', which='major', pad=15)
# Add horizontal lines for models without ppl estimates.
no_ppl_data = joined_data[joined_data.test_ppl.isna()]
for model_name, rows in no_ppl_data.groupby("pretty_model_name"):
y = rows.correct.mean()
ax.axhline(y, zorder=1, linewidth=3, **BASELINE_LINESTYLE)
if "GRNN" in model_name: # custom spacing tweaking
y_offset = -0.03
else:
y_offset = 0.006
ax2.text(540, y + y_offset, model_name, fontdict={"size": 38}, ha='right')
plt.subplots_adjust(wspace=0.2)
ax1.get_legend().remove()
ax1.set_ylabel(SG_ABSOLUTE_LABEL)
ax2.set_ylabel("")
plt.xlabel(PERPLEXITY_LABEL, labelpad=10, position=(-6,0))
# Add break in x-axis
ax1.set_xlim(0,ax1max)
ax2.set_xlim(ax2min,ax2max)
# hide the spines between ax1 and ax2
ax1.spines['right'].set_visible(False)
ax2.spines['left'].set_visible(False)
ax2.get_yaxis().set_ticks([])
d = .015 # how big to make the diagonal lines in axes coordinates
kwargs = dict(transform=ax1.transAxes, color='k', clip_on=False)
ax1.plot((1-d,1+d), (-d,+d), **kwargs)
kwargs.update(transform=ax2.transAxes) # switch to the right subplot
ax2.plot((-d*ax_ratio,+d*ax_ratio), (-d,+d), **kwargs)
# Change some legend labels.
handles, labels = ax1.get_legend_handles_labels()
legend_title_map = {"pretty_model_name": "Model",
"pretty_corpus": "Corpus",
"corpus_size": "Corpus size",
"corpus_bpe": "Tokenization"}
# Re-map some labels.
# labels = [legend_title_map.get(l, l) for l in labels]
drop_indices = [i for i,l in enumerate(labels) if l in legend_title_map.keys() or l in no_ppl_data.pretty_model_name.values]
handles = [h for i,h in enumerate(handles) if i not in drop_indices]
labels = [l for i,l in enumerate(labels) if i not in drop_indices]
labels = [l if l not in joined_data.corpus_size.unique() else "BLLIP-%s" % l.upper() for l in labels]
# Add empty handle for legend spacing.
handles.insert(4, mpatches.Patch(facecolor="white"))
labels.insert(4, "")
# Re-order labels.
new_order = ["BLLIP-LG", "LSTM", "BLLIP-MD", "ON-LSTM", "BLLIP-SM", "RNNG", "BLLIP-XS", "GPT-2", "", "n-gram"]
inds = [labels.index(l) for l in new_order]
handles = [handles[i] for i in inds]
labels = [labels[i] for i in inds]
# Set model style markers in legend to outlines only.
for i, (l, h) in enumerate(zip(labels, handles)):
if l != "":
h.set_sizes([500.0])
if l in joined_data.pretty_model_name.unique():
handles[i] = Line2D([0], [0], marker=markers[l], color='k', mew=3, lw=0,
markerfacecolor='w', markersize=27)
plt.legend(handles, labels, bbox_to_anchor=(-16.4,-0.18), ncol=5, loc="center left", columnspacing=0.5, handletextpad=0.05)
if RENDER_FINAL:
# Can't use render_final function because of some spine issues.
plt.savefig(figure_path / "perplexity.pdf", bbox_inches="tight")
# -
| notebooks/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.11 64-bit (''overwatch'': conda)'
# language: python
# name: python3
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# # Custom Raster Datasets
#
# In this tutorial, we demonstrate how to create a custom `RasterDataset` for our own data. We will use the [xView3](https://iuu.xview.us/) tiny dataset as an example.
# ## Setup
# %pip install torchgeo
# ## Imports
# +
from pathlib import Path
from typing import Callable, Dict, Optional
import matplotlib.pyplot as plt
import torch
from rasterio.crs import CRS
from torch import Tensor
from torch.utils.data import DataLoader
from torchgeo.datasets import RasterDataset, stack_samples
from torchgeo.samplers import RandomGeoSampler
# -
# ## Custom RasterDataset
# ### Unzipping the sample xView3 data from the tests folder
# +
from torchgeo.datasets.utils import extract_archive
data_root = Path('../../tests/data/xview3/')
extract_archive(str(data_root / 'sample_data.tar.gz'))
# -
# Now we have the xView3 tiny dataset downloaded and unzipped in our local directory. Note that all the test GeoTIFFs are comprised entirely of zeros. Any plotted image will appear to be entirely uniform.
#
# xview3
# ├── 05bc615a9b0aaaaaa
# │ ├── bathymetry.tif
# │ ├── owiMask.tif
# │ ├── owiWindDirection.tif
# │ ├── owiWindQuality.tif
# │ ├── owiWindSpeed.tif
# │ ├── VH_dB.tif
# │ └── VV_dB.tif
#
# We would like to create a custom Dataset class based off of RasterDataset for this xView3 data. This will let us use `torchgeo` features such as: random sampling, merging other layers, fusing multiple datasets with `UnionDataset` and `IntersectionDataset`, and more. To do this, we can simply subclass `RasterDataset` and define a `filename_glob` property to select which files in a root directory will be included in the dataset. For example:
class XView3Polarizations(RasterDataset):
'''
Load xView3 polarization data that ends in *_dB.tif
'''
filename_glob = "*_dB.tif"
# +
ds = XView3Polarizations(data_root)
sampler = RandomGeoSampler(ds, size=1024, length=5)
dl = DataLoader(ds, sampler=sampler, collate_fn=stack_samples)
for sample in dl:
image = sample['image']
print(image.shape)
image = torch.squeeze(image)
plt.imshow(image, cmap='bone', vmin=-35, vmax=-5)
| docs/tutorials/custom_raster_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import torch
import torchphysics as tp
import math
import numpy as np
import pytorch_lightning as pl
print('Tutorial zu TorchPhysics:')
print('https://torchphysics.readthedocs.io/en/latest/tutorial/tutorial_start.html')
from IPython.display import Image, Math, Latex
from IPython.core.display import HTML
Image(filename='bearing.png',width = 500, height = 250)
# First define all parameters:
h_0 = 16.e-06 #m = 16 um
dh = 14e-06 #m = 14 um
D = 0.01 #m = 10 mm
L = np.pi*D # Länge von Gebiet
u_m = 0.26 #m/s 0.26
beta = 2.2*1e-08 # 2.2e-08 m^2/N
nu_0 = 1.5e-03 # Pa·s = 1.5 mPa·s
# lower and upper bounds of parameters
nu0 = 1.0e-03 # Viskosität
nu1 = 2.5e-03
um0 = 0.2 # Geschwindigkeit
um1 = 0.4
dh0 = 10e-6 # Spaltvariaton
dh1 = 15e-6
p_0 = 1e+5 # 1e+5 N/m^2 = 1 bar
p_rel = 0 # Relativdruck
p_skal = 100000 #Skalierungsdruck für (-1,1) Bereich
# +
# define h:
def h(x, dh): # <- hier jetzt auch dh als input
return h_0 + dh * torch.cos(2*x/D) # x in [0,pi*D]
# and compute h':
def h_x(x, dh): # <- hier jetzt auch dh als input
return -2.0*dh/D * torch.sin(2*x/D) # x in [0,pi*D]
# +
# define the function of the viscosity.
# Here we need torch.tensors, since the function will be evaluated in the pde.
# At the beginng the model will have values close to 0,
# therefore the viscosity will also be close to zero.
# This will make the pde condition unstable, because we divide by nu.
# For now set values smaller then 1e-06 to 1e-06
def nu_func(nu, p):
out = nu * torch.exp(beta * p*p_skal)
return torch.clamp(out, min=1e-06)
def nu_x_func(nu,p):
out = nu* beta*p_skal*torch.exp(beta*p*p_skal)
return out
# -
# Variables:
x = tp.spaces.R1('x')
nu = tp.spaces.R1('nu')
um = tp.spaces.R1('um')
dh = tp.spaces.R1('dh')
# output
p = tp.spaces.R1('p')
A_x = tp.domains.Interval(x, 0, L)
A_nu = tp.domains.Interval(nu, nu0, nu1)
A_um = tp.domains.Interval(um, um0, um1)
A_dh = tp.domains.Interval(dh, dh0, dh1)
#inner_sampler = tp.samplers.AdaptiveRejectionSampler(A_x*A_nu*A_um*A_dh, n_points = 500000)
inner_sampler = tp.samplers.RandomUniformSampler(A_x*A_nu*A_um*A_dh, n_points = 2000000)
# density: 4 Punkte pro Einheitsvolumen
# Boundaries
boundary_v_sampler = tp.samplers.RandomUniformSampler(A_x.boundary*A_nu*A_um*A_dh, n_points = 100000)
# +
#tp.utils.scatter(nu*um*dh, inner_sampler, boundary_v_sampler)
# -
model = tp.models.Sequential(
tp.models.NormalizationLayer(A_x*A_nu*A_um*A_dh),
tp.models.FCN(input_space=x*nu*um*dh, output_space=p, hidden=(20,20,20))
)
display(Math(r'h(x)\frac{d^2 \tilde{p}}{d x^2} +\left( 3 \frac{dh}{dx} - \frac{h}{\nu} \frac{d \nu}{d x} \
\right) \frac{d \tilde{p}}{d x} = \frac{6 u_m \nu}{p_0 h^2} \frac{d h}{d x}\quad \mbox{with} \
\quad \tilde{p}=\frac{p}{p_{skal}} '))
from torchphysics.utils import grad
# Alternativ tp.utils.grad
def pde(nu, p, x, um, dh): # <- brauchen jetzt dh und um auch als input
# evaluate the viscosity and their first derivative
nu = nu_func(nu,p)
nu_x = nu_x_func(nu,p)
# implement the PDE:
# right hand site
rs = 6*um*nu #<- hier jetzt um statt u_m, da deine Variable so heißt
# h und h_x mit Input dh:
h_out = h(x, dh) # nur einmal auswerten
h_x_out = h_x(x, dh) # nur einmal auswerten
#out = h_out * grad(grad(p,x),x)- rs*h_x_out/h_out/h_out/p_skal
out = h_out*grad(grad(p,x),x) + (3*h_x_out -h_out/nu*nu_x)*grad(p,x) - rs*h_x_out/h_out/h_out/p_skal
return out
pde_condition = tp.conditions.PINNCondition(module=model,
sampler=inner_sampler,
residual_fn=pde,
name='pde_condition')
# +
# Hier brauchen wir immer nur den output des modells, da die Bedingung nicht
# von nu, um oder dh abhängt.
def bc_fun(p):
return p-p_rel
boundary_condition = tp.conditions.PINNCondition(module = model,
sampler = boundary_v_sampler,
residual_fn = bc_fun,
name = 'pde_bc')
# -
opt_setting = tp.solver.OptimizerSetting(torch.optim.AdamW, lr=1e-2) #SGD, LBFGS
solver = tp.solver.Solver((pde_condition, boundary_condition),optimizer_setting = opt_setting)
# +
trainer = pl.Trainer(gpus='-1' if torch.cuda.is_available() else None,
num_sanity_val_steps=0,
benchmark=True,
log_every_n_steps=1,
max_steps=1000,
#logger=False, zur Visualisierung im tensorboard
checkpoint_callback=False
)
trainer.fit(solver)
# +
opt_setting = tp.solver.OptimizerSetting(torch.optim.LBFGS, lr=1e-3) #SGD, LBFGS
solver = tp.solver.Solver((pde_condition, boundary_condition),optimizer_setting = opt_setting)
trainer = pl.Trainer(gpus='-1' if torch.cuda.is_available() else None,
num_sanity_val_steps=0,
benchmark=True,
log_every_n_steps=1,
max_steps=600,
#logger=False, zur Visualisierung im tensorboard
checkpoint_callback=False
)
trainer.fit(solver)
# -
import matplotlib.pyplot as plt
solver = solver.to('cpu')
print('nu0= ',nu0,' nu1= ',nu1)
print('dh0= ',dh0, 'dh1= ', dh1, 'm')
print('um0= ', um0, 'um1= ',um1, 'm/s')
# Parameter definieren für Plot
nu_plot = 2.0e-3
um_plot = 0.4
dh_plot = 14.25e-06
print('Minimale Spalthöhe =', h_0-dh_plot)
plot_sampler = tp.samplers.PlotSampler(plot_domain=A_x, n_points=600, device='cpu',
data_for_other_variables={'nu':nu_plot,
'um':um_plot,'dh':dh_plot})
if nu0<=nu_plot and nu_plot<=nu1 and dh0<=dh_plot and dh_plot<=dh1 and um0<=um_plot and um_plot<=um1:
fig = tp.utils.plot(model,lambda p:p,plot_sampler)
else:
print('Ausserhalb des Trainingsbereiches')
print('Skalierungsfaktor = ', p_skal)
plt.savefig(f'p_{um}.png', dpi=300)
# +
import xlsxwriter
#erstellen eines Workbook Objektes mit dem Dateinamen "Gleitlager_***.xlsx"
workbook = xlsxwriter.Workbook('Gleitlager.xlsx')
worksheet = workbook.add_worksheet('Tabelle_1')
worksheet.write('Ergebnistabelle Gleitlager')
worksheet.write('nu', 'dh', 'um')
workbook.close()
# -
import winsound
frequency = 2500 # Set Frequency To 2500 Hertz
duration = 1000 # Set Duration To 1000 ms == 1 second
winsound.Beep(frequency, duration)
| examples/Gleitlager/Gleitlager_V1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="uzvf8uDNAjAy"
from tensorflow import keras
from keras.models import Sequential
from keras.layers import Dense
from tensorflow.keras.utils import plot_model
import tensorflow as tf
from keras import backend as K
# + colab={"base_uri": "https://localhost:8080/", "height": 305} id="vYxRPt4wCzpk" outputId="59f388fe-cbee-43f6-8a29-84cf8d0555cb"
import numpy as np
import matplotlib.pyplot as plt
# !git clone https://github.com/PAULGOYES/pylops_seismic_cs.git
#coil_synt_cspread
Data = np.load('pylops_seismic_cs/syn3D_cross-spread2.npy')[:900,:,:14]
plt.figure()
plt.imshow(Data[:,:,13],aspect='auto',cmap='seismic')
plt.colorbar()
print('the shape is: ', Data.shape)
# + id="L_603TVwKht-" colab={"base_uri": "https://localhost:8080/"} outputId="5f82ef37-65fa-4abc-ae64-da7ed93596d2"
indexDel = [3,5,7,10,12]
IndexSam = [0,1,2,4,6,8,9,11,13]
#idx_Train = indexx[:int(50*50*0.2)]
#idx_Val = indexx[int(50*50*0.2):]
#print(idx)
timesampling = np.linspace(0,1,Data.shape[0])
tracesampling = np.linspace(0,1,Data.shape[1])
shotsampling = np.linspace(0,1,Data.shape[2])
TRACEarray, TIMEarray, SHOTarray = np.meshgrid(tracesampling,timesampling,shotsampling)
COORXYZval = np.stack((TRACEarray.ravel(),TIMEarray.ravel(),SHOTarray.ravel()),axis=-1)
timecoor = np.delete(TIMEarray,indexDel,axis=-1)
tracecoor = np.delete(TRACEarray,indexDel,axis=-1)
shotcoor = np.delete(SHOTarray,indexDel,axis=-1)
COORXYZtrain = np.stack((tracecoor.ravel(),timecoor.ravel(),shotcoor.ravel()),axis=-1)
OUTPUTtrain = np.delete(Data,indexDel,axis=-1).reshape(-1,1)
print(COORXYZtrain.shape)
print(OUTPUTtrain.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="999eJ7Rie1yS" outputId="7c165f98-9c48-4fc8-9d43-f3d5aa8dd86b"
timesamplingval = np.delete(TIMEarray,IndexSam,axis=-1)
tracesamplingval = np.delete(TRACEarray,IndexSam,axis=-1)
shotsamplingval = np.delete(SHOTarray,IndexSam,axis=-1)
COORXYZvaldel = np.stack((tracesamplingval.ravel(),timesamplingval.ravel(),shotsamplingval.ravel()),axis=-1)
OUTPUTval = np.delete( Data,IndexSam,axis=-1).reshape(-1,1)
print(COORXYZvaldel.shape)
print(OUTPUTval.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 593} id="WFFvGjivZ4hm" outputId="007c8821-0abd-4be1-f89b-e6eadabc975a"
fig = plt.figure(figsize = (10, 10))
ax = plt.axes(projection ="3d")
scttssim=ax.scatter3D(COORXYZtrain[:,0],COORXYZtrain[:,1],COORXYZtrain[:,2], c = OUTPUTtrain,s = 0.1, cmap='seismic',alpha=0.1)
scttssim=ax.scatter3D(COORXYZvaldel[:,0],COORXYZvaldel[:,1],COORXYZvaldel[:,2], c = OUTPUTval,s = 0.1,cmap='jet')
plt.title('COORDINATE SURVEY')
ax.set_xlabel('TRACE', fontweight ='bold')
ax.set_ylabel('TIME', fontweight ='bold')
ax.set_zlabel('SHOT', fontweight ='bold')
# + id="S68ki133QHSH" colab={"base_uri": "https://localhost:8080/"} outputId="847ace5a-ba3d-4bd2-8ae4-d2c99e053312"
import numpy as np
def input_mapping(x, L,log_sampling=True):
frequency_bands= None
if log_sampling:
frequency_bands = 2.0**np.linspace(0.0,
L-1,
L)
else:
frequency_bands = np.linspace(
2.0**0.0,
2.0**(L-1),
L)
encoding = []
for idx,freq in enumerate(frequency_bands):
for fun in [np.sin, np.cos]:
encoding.append(fun(x*freq))
return np.concatenate(encoding)
xx = np.array([[0.2,0.25,0.1],[0.1,0.8,0.3]])
a = input_mapping(xx.T,L=1,log_sampling=True)
#ejecute aqui un ejemplo
print(a.shape)
# + [markdown] id="7h3GKNjrlYki"
# - loglinear sampling $\kappa = 2^{i-1} \pi \mathbf{v} \}_{i=1}^{L}$
# - linear sampling $\kappa = \mathbf{v} \pi (i)/2 \}_{i=1}^{L}$
# + colab={"base_uri": "https://localhost:8080/"} id="xhLVkjf0VR3e" outputId="1a9d030c-8f13-44f5-ee7a-f2ec0559a33d"
def auxfun(x,L,sampling):
temp=[]
if sampling == 'loglinear':
kappa = np.pi*2.0**np.arange(L)
elif sampling == 'linear':
kappa = 0.5*np.pi*np.arange(1,L+1)#np.linspace(2.0**0, 2.0**(L-1),L)
for k in kappa:
for fun in [np.sin, np.cos]:
temp.append(fun(x*k))
return temp
def auxfun2(x,L,sampling):
xaxisPE = auxfun(x[0],L[0],sampling)
yaxisPE = auxfun(x[1],L[1],sampling)
zaxisPE = auxfun(x[2],L[2],sampling)
return np.concatenate((xaxisPE,yaxisPE,zaxisPE),axis=0)
def input_mapping_aniso(x,L,sampling='loglinear'):
out = np.zeros((x.shape[0],2*np.sum(L)))
for i in range(x.shape[0]):
out[i,:] = auxfun2(x[i,:],L,sampling)
return out
a = input_mapping_aniso(xx,L=[2,1,1],sampling='loglinear')
print(a)
# + [markdown] id="g8H0EGJwSwN4"
# ## Train networks with different input mappings
#
# We compare the following mappings $\gamma (\mathbf{v})$.
#
# - No mapping: $\gamma(\mathbf{v})= \mathbf{v}$.
#
# - Basic mapping: $\gamma(\mathbf{v})=\left[ \cos(\kappa \mathbf{v}),\sin(\kappa \mathbf{v}) \right]^\mathrm{T}$.
#
# - Positional encoding: $\gamma(\mathbf{v})=\left[ \ldots, \cos(\kappa_j \mathbf{v}),\sin(\kappa_j \mathbf{v}), \ldots \right]^\mathrm{T}$ for $j = 0, \ldots, L-1$
#
# - L-variational positional encoding: $\gamma(\mathbf{x,y,z})=
# \begin{bmatrix}
# \left[ \ldots, \cos(\kappa_i \mathbf{x}),\sin(\kappa_i \mathbf{x}), \ldots \right]^\mathrm{T} for \ i = 0, \ldots, L_\mathbf{x} -1 \\
# \left[ \ldots, \cos(\kappa_j \mathbf{y}),\sin(\kappa_j \mathbf{y}), \ldots \right]^\mathrm{T} for \ j = 0, \ldots, L_\mathbf{y} -1 \\
# \left[ \ldots, \cos(\kappa_k \mathbf{z}),\sin(\kappa_k \mathbf{z}), \ldots \right]^\mathrm{T} for \ k = 0, \ldots, L_\mathbf{z} -1
# \end{bmatrix}$
#
# - Gaussian Fourier feature mapping: $\gamma(\mathbf{v})= \left[ \cos(2 \pi \mathbf B \mathbf{v}), \sin(2 \pi \mathbf B \mathbf{v}) \right]^\mathrm{T}$,
# where each entry in $\mathbf B \in \mathbb R^{m \times d}$ is sampled from $\mathcal N(0,\sigma^2)$, where $m$ is the number of channels and $d$ is the input dimension.
# + id="iiTeUdpzDmjP" colab={"base_uri": "https://localhost:8080/"} outputId="9527afee-3f0d-4c91-a791-88f87abe53cc"
def input_mappingB(x, B):
if B is None:
return x
else:
x_proj = (x) @ B.T
#print(x_proj.shape)
return np.concatenate([np.sin(x_proj), np.cos(x_proj)], axis=-1)
print(input_mappingB(xx,np.eye(3)))
# + id="QK5O7jXvFg9-"
def createMLPFFM(input_dim, channels, Hiddenlayers, output_dim, L=None):
model = keras.Sequential()
if L is None:
print('your input shape in network is: ', input_dim)
for i in range(Hiddenlayers):
model.add(Dense(channels,activation='relu'))
model.add(Dense(output_dim, activation='elu'))
model.build(input_shape=(None, input_dim))
else:
print('your input shape in network is: ', 2*np.sum(L))
for i in range(Hiddenlayers):
model.add(Dense(channels,activation='relu'))
model.add(Dense(output_dim, activation='elu'))
model.build(input_shape=(None, 2*np.sum(L)))
return model
# + colab={"base_uri": "https://localhost:8080/"} id="wM30Q4c6cs7R" outputId="9d27f767-0d21-446a-85b9-47c116f565e1"
#B = np.random.normal(loc=0.0, scale=1e-2, size=(256,2))
#B = np.eye(3)
#datatrainFFM = input_mapping(COORXYZtrain, B)
L = [2,1,1]
sampling = 'loglinear'
datatrainFFM = input_mapping_aniso(COORXYZtrain,L=L,sampling=sampling)
datavalFFM = input_mapping_aniso(COORXYZvaldel,L=L,sampling=sampling)
network = createMLPFFM(input_dim=3, channels=128, Hiddenlayers=15, output_dim=1, L=L)
print(network.summary())
print('shape of train input: ', datatrainFFM.shape)
#print('shape of val input: ', datavalFFM.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="O5wtUAwpdBN-" outputId="7309f536-37c9-4b2f-bb49-b2acff61ad9f"
opt = keras.optimizers.Adam(learning_rate=1e-3)
lossf = keras.losses.Huber(delta=0.5, reduction="auto", name="huber_loss")
msel = tf.keras.losses.MeanSquaredError()
def log10(x):
numerator = K.log(x)
denominator = K.log(K.constant(10, dtype=numerator.dtype))
return numerator / denominator
def PSNR(y_true, y_pred):
max_pixel=1
return 10.0 * log10( (max_pixel) / msel(y_true,y_pred) )
def l5(y_true, y_pred):
return (1+keras.losses.cosine_similarity(y_true, y_pred, axis=0))/2 + keras.losses.huber(y_true, y_pred, delta=0.5)
network.compile(optimizer=opt, loss=l5, metrics=PSNR)
history= network.fit(
x=datatrainFFM,
y=OUTPUTtrain,
batch_size=20000,
epochs=500,
verbose="auto", validation_data=(datavalFFM,OUTPUTval)
)
#validation_data=(datavalFFM,y_output[idx_Val,:])
# + colab={"base_uri": "https://localhost:8080/", "height": 299} id="htcH-OHeg5Uf" outputId="df1235ba-600a-4eb8-895f-4595f6791c86"
print(history.history.keys())
fig, ax1 = plt.subplots()
ax1.tick_params(axis='y', labelcolor='blue')
ax1.plot(history.history['val_loss'][10:],'--b',label='LOSS')
ax1.plot(history.history['loss'][10:],'b',label='LOSS')
ax1.set_ylabel('LOSS', color='blue')
#plt.legend(loc='upper left')
ax2 = ax1.twinx() # instantiate a second axes that shares the same x-axis
ax2.set_ylabel('PSNR', color='red')
ax2.plot(history.history['PSNR'][10:],'red',label='PSNR')
ax2.plot(history.history['val_PSNR'][10:],'--r',label='PSNR')
ax2.tick_params(axis='y', labelcolor='red')
plt.xlabel('epoch')
plt.title('model convergence')
#plt.legend(loc='upper right')
plt.show()
# + id="gJfST_tGseo4"
y_predict = network.predict(input_mapping_aniso(COORXYZval,L=L,sampling=sampling))
# + colab={"base_uri": "https://localhost:8080/", "height": 625} id="PT7R5ZstB0ks" outputId="06643999-df39-44fe-eab4-95052028d7f2"
kk = -4
plt.figure(figsize=(20,10))
plt.subplot(122)
plt.imshow(y_predict.reshape(Data.shape)[:,:,indexDel[kk]],aspect='auto',cmap='seismic'),plt.colorbar()
plt.title('REC-shot '+ str(indexDel[kk]))
plt.subplot(121)
plt.imshow(Data[:,:,indexDel[kk]],aspect='auto',cmap='seismic'), plt.colorbar()
plt.title('GT-shot '+ str(indexDel[kk]))
# + id="Nju3TA8CqGzU" colab={"base_uri": "https://localhost:8080/", "height": 338} outputId="26ac175f-8784-437c-abb8-98ba61475370"
trace = 40
plt.figure(figsize=(10,5))
plt.plot(y_predict.reshape(Data.shape)[:,trace,indexDel[kk]],'r',label='REC')
plt.plot(Data[:,trace,indexDel[kk]],'b',label='GT')
plt.legend()
# + id="mCZXAaTHgEcN"
| PositionalEnconding_SEISMIC_synthetic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basic scVI Tutorial
# cd ../..
# +
import os
import numpy as np
import seaborn as sns
from sklearn.manifold import TSNE
import matplotlib.pyplot as plt
# %matplotlib inline
from scvi.dataset import CortexDataset, RetinaDataset
from scvi.metrics.clustering import entropy_batch_mixing, get_latent
from scvi.metrics.differential_expression import de_stats
from scvi.metrics.imputation import imputation
from scvi.models import VAE, SVAEC
from scvi.inference import VariationalInference
# -
# ## Loading data
#
# Here we load the CORTEX dataset described in
#
# * Zeisel, Amit, et al. "Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq." Science 347.6226 (2015): 1138-1142.
#
# Please see our data loading Jupyter notebook for more examples of data loading -- scVI has many "built-in" datasets, as well as support for loading arbitrary .csv, .loom, and .h5ad (AnnData) files.
gene_dataset = CortexDataset()
# ## Training
# * __n_epochs__: Maximum number of epochs to train the model. If the likelihood change is small than a set threshold training will stop automatically.
# * __lr__: learning rate. Set to 0.001 here.
# * __use_batches__: If the value of true than batch information is used in the training. Here it is set to false because the cortex data only contains one batch.
# * __use_cuda__: Set to true to use CUDA.
#
n_epochs=500
lr=1e-3
use_batches=False
use_cuda=True
# **Train the model and output model likelihood every 5 epochs**
vae = VAE(gene_dataset.nb_genes, n_batch=gene_dataset.n_batches * use_batches)
infer = VariationalInference(vae,
gene_dataset,
train_size=0.9,
use_cuda=use_cuda,
frequency=5)
infer.train(n_epochs=n_epochs, lr=lr)
# ** Plotting the likelihood change across the 500 epochs of training: blue for training error and orange for testing error.**
ll_train = infer.history["ll_train"]
ll_test = infer.history["ll_test"]
x = np.linspace(0,500,(len(ll_train)))
plt.plot(x, ll_train)
plt.plot(x, ll_test)
plt.ylim(1150,1600)
plt.show()
# ## Visualizing the latent space
infer.show_t_sne('sequential', n_samples=False, color_by='labels')
# ## Imputation
#
#
# The ability to impute missing values is useful in practical applications in addition to providing an assay for generalization performance. In the following analysis, we benchmark scVI against BISCUIT, ZINB-WaVE and ZIFA, as well as MAGIC, which provides imputation without explicit statistical modeling. To evaluate these methods on a given dataset, we generated a **corrupted training set**, and then fitted the perturbed dataset with each of the benchmark methods and evaluate them by comparing the imputed values to the original ones (Methods 4.7). Overall, we observe that the imputation accuracy of scVI is higher or comparable (less than one transcript for median error) across all datasets
#
# #### Corrupting the datasets for imputation benchmarking.
#
# Two different approaches to measure the robustness of algorithms to noise in the data:
#
# - **Uniform zero introduction**: select randomly a rate r% of the non-zero entries and multiply the entry n with a Ber(0.9) random variable.
# - **Binomial data corruption**: select a rate r% of the matrix and replace an entry n by a Bin(n, 0.2) random variable.
#
# By default, the rate r is set a 0.1
#
# #### Accuracy of imputing missing data
#
# As imputation tantamount to replace missing data by its mean conditioned on being observed, we use the median L1 distance between the original dataset and the imputed values for corrupted entries only.
# Parameters:
# * The rate of simulated dropout is defined by __rate__, here set ot 0.1
original_list, imputed_list = infer.imputation('train', # the part of the dataset to corrupt
rate=0.1, # the part of the dataset to corrupt
n_epochs=1,
corruption="uniform", # or "binomial"
verbose=True)
# Median of medians for all distances
imputation_errors = np.abs(np.concatenate(original_list) - np.concatenate(imputed_list))
median_imputation_score = np.median(imputation_errors)
print(median_imputation_score)
# We then plot the distribution of absolute errors between the imputed value and the true value at the dropout positions.
#
# Note: The imputed value __px_rate__ is the rate parameter (expected value) of the Zero-Inflated Negative Binomial (ZINB) distribution.
plt.hist(np.log10(imputation_errors))
# ## Differential Expression
# From the trained VAE model we can sample the gene expression rate for each gene in each cell. For the two populations of interest, we can then randomly sample pairs of cells, one from each population to compare their expression rate for a gene. The degree of differential expression is measured by __logit(p/(1-p))__ where __p__ is the probability of a cell from population A having a higher expression than a cell from population B. We can form the null distribution of the DE values by sampling pairs randomly from the combined population.
#
# The following example is implemented for the cortext dataset, vary __cell_types__ and __genes_of_interest__ for other datasets.
# **1. Set population A and population B for comparison**
# +
cell_types = np.array(['astrocytes_ependymal', 'endothelial-mural', 'interneurons', 'microglia',
'oligodendrocytes', 'pyramidal CA1', 'pyramidal SS'], dtype=np.str)
# oligodendrocytes (#4) VS pyramidal CA1 (#5)
couple_celltypes = (4, 5) # the couple types on which to study DE
print("\nDifferential Expression A/B for cell types\nA: %s\nB: %s\n" %
tuple((cell_types[couple_celltypes[i]] for i in [0, 1])))
# -
# **2. Define parameters**
# * __M_sampling__: the number of times to sample __px_scales__ from the vae model for each gene in each cell.
# * __M_permutation__: Number of pairs sampled from the px_scales values for comparison.
M_sampling = 100
M_permutation = 100000
permutation = False
# **3. Sample from the gene expression level from all cells**
# Note: The expectation of the ZINB distribution __px_rate ~ library_size * px_scale__, so __px_scale__ could be understood as the mean gene expression level of each cell after adjusting for the library size factor.
px_scale, all_labels = infer.differential_expression_stats('train')
# **4. Extract the sampled gene expression level for the two populations of interest, and create indexes for the samples**
# +
sample_rate_a = px_scale[all_labels.view(-1) == couple_celltypes[0]].view(-1, px_scale.size(1)).cpu().detach().numpy()
sample_rate_b = px_scale[all_labels.view(-1) == couple_celltypes[1]].view(-1, px_scale.size(1)).cpu().detach().numpy()
list_1 = list(np.arange(sample_rate_a.shape[0]))
list_2 = list(sample_rate_a.shape[0] + np.arange(sample_rate_b.shape[0]))
samples = np.vstack((sample_rate_a, sample_rate_b))
# -
# **5. Compute whether a gene is differentially expressed by computing pairs of cells from population A and population B**
u, v = np.random.choice(list_1, size=M_permutation), np.random.choice(list_2, size=M_permutation)
first_set = samples[u]
second_set = samples[v]
res1 = np.mean(first_set >= second_set, 0)
res1 = np.log(res1 + 1e-8) - np.log(1 - res1 + 1e-8)
# **6. Obtaining the null value by comparing pairs sampled from the combined population**
u, v = (np.random.choice(list_1 + list_2, size=M_permutation),
np.random.choice(list_1 + list_2, size=M_permutation))
first_set = samples[u]
second_set = samples[v]
res2 = np.mean(first_set >= second_set, 0)
res2 = np.log(res2 + 1e-8) - np.log(1 - res2 + 1e-8)
# **7. Print out the differential expression value from both the true comparison and the permuted comparison**
genes_of_interest = ["Thy1", "Mbp"]
gene_names = gene_dataset.gene_names
result = [(gene_name, res1[np.where(gene_names == gene_name.upper())[0]][0],res2[np.where(gene_names == gene_name.upper())[0]][0]) for gene_name in genes_of_interest]
print('\n'.join([gene_name + " : " + str(r1) + " , "+ str(r2) for (gene_name, r1,r2) in result]))
# **8. Plot the null distribution of the DE values**
plt.hist(res2)
# ## Correction for batch effects
#
# First we load the RETINA dataset that is described in
#
# * <NAME>, et al. "Comprehensive classification of retinal bipolar neurons by single-cell transcriptomics." Cell 166.5 (2016): 1308-1323.
gene_dataset = RetinaDataset()
# +
n_epochs=50
lr=1e-3
use_batches=True
use_cuda=True
### Train the model and output model likelihood every 5 epochs
vae = VAE(gene_dataset.nb_genes, n_batch=gene_dataset.n_batches * use_batches)
infer = VariationalInference(vae,
gene_dataset,
train_size=0.9,
use_cuda=use_cuda,
frequency=5)
infer.train(n_epochs=n_epochs, lr=lr)
# +
# Plotting the likelihood change across the 50 epochs of training: blue for training error and orange for testing error.
ll_train = infer.history["ll_train"]
ll_test = infer.history["ll_test"]
x = np.linspace(0,50,(len(ll_train)))
plt.plot(x, ll_train)
plt.plot(x, ll_test)
plt.ylim(min(ll_train)-50, 3500)
plt.show()
# -
# **Computing batch mixing**
print("Entropy batch mixing :", infer.entropy_batch_mixing('sequential'))
# **Coloring by batch and cell type**
# +
# obtaining latent space in the same order as the input data
infer.show_t_sne('sequential',n_samples=1000, color_by='batches and labels')
| docs/notebooks/basic_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
from PIL import Image
import glob
image_list = []
image_name=[]
for filename in glob.glob('tl_images/Green/*.jpg'):
#im=Image.open(filename)
im=cv2.imread(filename)
image_list.append(im)
image_name.append(filename)
n_train = len(image_list)
print("The size of test set is ", n_train)
# +
import numpy as np
import matplotlib.pyplot as plt
import random
# Visualizations will be shown in the notebook.
# %matplotlib inline
num_of_samples=[]
plt.figure(figsize=(12, 50))
for i in range(0, 14):
plt.subplot(11, 4, i+1)
#x_selected = X_train[y_train == i]
img = cv2.imread(image_name[i])
hsv = cv2.cvtColor(img, cv2.COLOR_BGR2HSV)
mask = cv2.inRange(hsv, (36, 25, 25), (70, 255,255))
## slice the green
imask = mask>0
green = np.zeros_like(img, np.uint8)
green[imask] = img[imask]
plt.imshow(green)
title="Class %s \n" % (i)
plt.title(title)
plt.axis('off')
#num_of_samples.append(len(x_selected))
plt.show()
# -
image_list_red = []
image_name_red=[]
for filename in glob.glob('tl_images/Red/*.jpg'):
#im=Image.open(filename)
im=cv2.imread(filename)
image_list_red.append(im)
image_name_red.append(filename)
n_train_red = len(image_list_red)
print("The size of test set is ", n_train_red)
# +
plt.figure(figsize=(12, 50))
for i in range(0, 14):
plt.subplot(11, 4, i+1)
#x_selected = X_train[y_train == i]
img = cv2.imread(image_name_red[i],cv2.IMREAD_COLOR)
b,g,r = cv2.split(img) # get b,g,r
rgb_img = cv2.merge([r,g,b]) # switch it to rgb
#hsv = cv2.cvtColor(rgb_img, cv2.COLOR_RGB2HSV)
hsv = cv2.cvtColor(img, cv2.COLOR_BGR2HSV)
# lower mask (0-10)
#lower_red = np.array([0,50,50])
lower_red = np.array([0,150,150])
upper_red = np.array([10,255,255])
mask0 = cv2.inRange(hsv, lower_red, upper_red)
# upper mask (170-180)
#lower_red = np.array([170,50,50])
lower_red = np.array([160,150,150])
upper_red = np.array([180,200,200])
mask1 = cv2.inRange(hsv, lower_red, upper_red)
# join my masks
mask = mask0+mask1
res = np.count_nonzero(mask)
res0 = np.count_nonzero(mask0)
res1 = np.count_nonzero(mask1)
#print (res)
## slice the green
imask = mask>0
mask_red =cv2.inRange(r,240,255)
red = np.zeros_like(img, np.uint8)
red[imask] = rgb_img[imask]
#plt.imshow(red)
plt.imshow(rgb_img)
title="Class %s %s %s\n" % (res,res0,res1)
plt.title(title)
plt.axis('off')
#num_of_samples.append(len(x_selected))
plt.show()
# -
| .ipynb_checkpoints/test traffic lights-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab_type="code" id="KvPxXdKJguYB" colab={"base_uri": "https://localhost:8080/", "height": 353} outputId="5eca2014-cc63-4710-9dae-229131bc725c"
# Checking out the GPU we have access to
# !nvidia-smi
# + id="rw-juiHfM4MU" colab_type="code" colab={}
# !pip install -q transformers
# + colab_type="code" id="pzM1_ykHaFur" colab={}
# Importing stock libraries needed
import numpy as np
import pandas as pd
import torch
import torch.nn.functional as F
from torch.utils.data import Dataset, DataLoader, RandomSampler, SequentialSampler
from transformers import T5Tokenizer, T5ForConditionalGeneration
# Preparing for TPU usage
#import torch_xla
#import torch_xla.core.xla_model as xm
#device = xm.xla_device()
# + colab_type="code" id="NLxxwd1scQNv" colab={}
# # Setting up the device for GPU usage
from torch import cuda
device = 'cuda' if cuda.is_available() else 'cpu'
# + id="BKhI3WjLM4Mc" colab_type="code" colab={}
df = pd.read_csv('news.csv')
# + colab_type="code" id="FBoPE2bUd_Fx" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="5cbe6e40-07d6-4747-a29d-a90814d2afd8"
df.head()
# + colab_type="code" id="zQ1Jw1Q4hqEi" colab={}
# Sections of config
# Defining some key variables that will be used later on in the training
tokenizer = T5Tokenizer.from_pretrained("t5-base")
MAX_LEN = 512
SUMMARY_LEN = 60
TRAIN_BATCH_SIZE = 4
VALID_BATCH_SIZE = 1
EPOCHS = 2
LEARNING_RATE = 1e-4
# + colab_type="code" id="932p8NhxeNw4" colab={}
# Creating a custom dataset for reading the dataframe and loading it into the dataloader to pass it to the neural network at a later stage for finetuning the model and to prepare it for predictions
class CustomDataset(Dataset):
def __init__(self, dataframe, tokenizer, source_len, summ_len):
self.tokenizer = tokenizer
self.data = dataframe
self.source_len = source_len
self.summ_len = summ_len
self.text = self.data.text
self.ctext = self.data.ctext
def __len__(self):
return len(self.text)
def __getitem__(self, index):
ctext = str(self.ctext[index])
ctext = ' '.join(ctext.split())
text = str(self.text[index])
text = ' '.join(text.split())
source = self.tokenizer.batch_encode_plus([ctext], max_length= self.source_len, pad_to_max_length=True,return_tensors='pt')
target = self.tokenizer.batch_encode_plus([text], max_length= self.summ_len, pad_to_max_length=True,return_tensors='pt')
source_ids = source['input_ids'].squeeze()
source_mask = source['attention_mask'].squeeze()
target_ids = target['input_ids'].squeeze()
target_mask = target['attention_mask'].squeeze()
return {
'source_ids': source_ids.to(dtype=torch.long),
'source_mask': source_mask.to(dtype=torch.long),
'target_ids': target_ids.to(dtype=torch.long),
'target_ids_y': target_ids.to(dtype=torch.long)
}
# + colab_type="code" id="70YVNa-YiSHa" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="910a66ac-5154-44c0-d77d-7ddf1d0336d1"
# Creating the dataset and dataloader for the neural network
train_size = 0.9
train_dataset=df.sample(frac=train_size,random_state=42).reset_index(drop=True)
test_dataset=df.drop(train_dataset.index).reset_index(drop=True)
print("FULL Dataset: {}".format(df.shape))
print("TRAIN Dataset: {}".format(train_dataset.shape))
print("TEST Dataset: {}".format(test_dataset.shape))
training_set = CustomDataset(train_dataset, tokenizer, MAX_LEN, SUMMARY_LEN)
testing_set = CustomDataset(test_dataset, tokenizer, MAX_LEN, SUMMARY_LEN)
# + colab_type="code" id="oZ-Spz29idNS" colab={}
train_params = {'batch_size': TRAIN_BATCH_SIZE,
'shuffle': True,
'num_workers': 0
}
test_params = {'batch_size': VALID_BATCH_SIZE,
'shuffle': False,
'num_workers': 0
}
training_loader = DataLoader(training_set, **train_params)
testing_loader = DataLoader(testing_set, **test_params)
# + colab_type="code" id="51jKmk2eDINe" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="ee2cf4a6-34ff-47d4-873d-8fec1bfb405b"
model = T5ForConditionalGeneration.from_pretrained("t5-base")
model = model.to(device)
# + colab_type="code" id="3wbgnNFCms1O" colab={}
optimizer = torch.optim.Adam(params = model.parameters(), lr=LEARNING_RATE)
# + colab_type="code" id="SaPAR7TWmxoM" colab={}
def train(epoch):
model.train()
for _,data in enumerate(training_loader, 0):
y = data['target_ids'].to(device, dtype = torch.long)
y_ids = y[:, :-1].contiguous()
lm_labels = y[:, 1:].clone().detach()
lm_labels[y[:, 1:] == tokenizer.pad_token_id] = -100
ids = data['source_ids'].to(device, dtype = torch.long)
mask = data['source_mask'].to(device, dtype = torch.long)
outputs = model(input_ids = ids, attention_mask = mask, decoder_input_ids=y_ids, lm_labels=lm_labels)
loss = outputs[0]
if _%1000==0:
print(f'Epoch: {epoch}, Loss: {loss.item()}')
optimizer.zero_grad()
loss.backward()
optimizer.step()
# xm.optimizer_step(optimizer)
# xm.mark_step()
# + colab_type="code" id="6rjDkuihe2iC" colab={}
import logging
logging.basicConfig(level=logging.ERROR)
# + colab_type="code" id="hFAfLq7WoPqi" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="b68bc370-6c28-4e4e-e99e-24b9ed9cfc43"
for epoch in range(EPOCHS):
train(epoch)
# + colab_type="code" id="wK4JcHY64gyk" colab={}
# + colab_type="code" id="1kF98Mao7CXG" colab={}
model.save_pretrained('/content/model/t5')
# + colab_type="code" id="hczENku064dA" colab={}
model = T5ForConditionalGeneration.from_pretrained('/content/model/t5')
# + colab_type="code" id="d0uPhIye8CLj" colab={}
tokenizer = T5Tokenizer.from_pretrained('t5-base')
# + colab_type="code" id="DEWb5ro68aQT" colab={}
#text = 'Hours after Rajasthan chief minister <NAME> reiterated his “forgive and forget” mantra on Thursday with the rebels back in party fold, the Congress revoked the suspension of two MLAs who were suspended on July 17 following allegations that they were involved in a conspiracy to topple the state government. MLAs <NAME> and <NAME> were part of the rebel camp supporting former Rajasthan deputy chief minister <NAME>. The suspension followed the leak of audio tapes with conversations allegedly between <NAME>, <NAME>, an intermediary, and Union minister of <NAME>, which hinted at a conspiracy to bring down the government.'
# + id="SvoKUEZecJ8v" colab_type="code" colab={}
text = df['text'][2]
# + colab_type="code" id="GlBg-0Xx8Fv-" colab={}
input_ids = tokenizer.encode_plus(text, return_tensors="pt")
# + colab_type="code" id="ecB0kP4f8Grj" colab={}
outputs = model.generate(input_ids['input_ids'])
# + colab_type="code" id="fYvIzpjz8NxN" colab={}
pred = [tokenizer.decode(g, skip_special_tokens=True, clean_up_tokenization_spaces=True) for g in outputs]
# + colab_type="code" id="KIDeA3le8P4g" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="033b2e8e-f81b-4358-f931-c56dba500f08"
pred
# + id="ePiJFY5XcBwm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="246434b7-fd31-4474-adce-b7c739360858"
df['ctext'][2]
| transformers_summarization (2).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Importing libraries
# +
import numpy as np
import matplotlib.pyplot as plt
import cv2
import os
import random
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation, Flatten, Conv2D, MaxPooling2D
# -
# # Loading data
# +
train = []
DATA_DIR = '../data/Animals-10'
CATEGORIES = ['butterfly', 'cat', 'chicken', 'cow', 'dog', 'elephant', 'horse', 'sheep', 'spider', 'squirrel']
IMG_SIZE = (32, 32)
for label, category in enumerate(CATEGORIES):
path = os.path.join(DATA_DIR, category)
images = os.listdir(path)
for img in images:
img_array = cv2.imread(os.path.join(path, img))
resized_img = cv2.resize(img_array, IMG_SIZE)
train.append([resized_img, label])
random.shuffle(train)
# +
X = []
y = []
for features, label in train:
X.append(features)
y.append(label)
X = np.array(X).reshape(-1, IMG_SIZE[0], IMG_SIZE[1], 3) / 255.0
y = np.array(y)
# -
# # Building CNN model
# +
model = Sequential()
model.add(Conv2D(32, (3, 3), input_shape=X.shape[1:]))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(64, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(128, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(64))
model.add(Dense(10))
model.add(Activation('softmax'))
model.compile(loss='sparse_categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# -
# # Training & evaluating the model
model.fit(X, y, batch_size=32, epochs=10, validation_split=0.2)
# # Enter new data
# +
img_path = input('Enter image name: ')
img_array = cv2.imread(os.path.join('./', img_path))
rgb_img = cv2.cvtColor(img_array, cv2.COLOR_BGR2RGB)
plt.imshow(rgb_img)
resized_img = cv2.resize(img_array, IMG_SIZE)
input_img = np.array(resized_img).reshape(-1, IMG_SIZE[0], IMG_SIZE[1], 3) / 255.0
prediction = model.predict(input_img)
category = CATEGORIES[np.argmax(prediction)]
description_file = open(f'./descriptions/{category}.txt', 'r')
description = description_file.read()
print(description)
| src/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import numpy as np #linear algebra
import pandas as pd #data preprocessing
import matplotlib.pyplot as plt #data visualization
import h5py
import PIL
import utils.general_utils as util
from utils.general_utils import populate_breeds, get_imgMatrix_from_id, get_breed_value_from_id, get_filename_from_id, get_id_from_filename
#other built utilities
from data_loader import dataFrameBuilder
#using inception_v3 to classify dog breeds
import tensorflow as tf #import tensroflow
import random
# +
#PARAMETERS
image_size = 500 #all images are size image_size x image_size x 3
batch_size = 1 #online learning... batch size = 1
num_classes = 120
train = True #if false, try and load in the model rather than re-construct it
np.set_printoptions(threshold=np.nan) #use entire numpy array
# +
# %matplotlib inline
#prepare csv files
train = pd.read_csv("../data/included/labels.csv")
test = pd.read_csv("../data/included/test_id.csv")
BREED_LIST = "../data/preprocessed_data/breed_list.csv"
# -
#prepare the breed list dataframe
labels = populate_breeds(BREED_LIST) #get the list of all dog breeds
labels_np = np.array(labels).reshape(120,1) #labels list reshaped to numpy array
# +
#x = tf.placeholder(tf.float32, shape=[None, 500,500, 3], name='input_data')
x = tf.keras.layers.Input(shape=(500,500,3), batch_size=batch_size,name='input_data',dtype='float32')
y = tf.placeholder(tf.float32, shape=[None, 120,1], name='correct_labels')
#x=tf.placeholder(tf.float32, shape=[500,500,3],name='input_data')
y_pred = tf.placeholder(tf.float32, shape=[None,120,1], name='predicted_labels')
# +
def train_input_fn(index=0, data_amnt = 1):
input_img_data = dataFrameBuilder(data_amount=data_amnt,
start_index=index)
#df.shuffle().repeat().batch(batch_size)
#print(df.sample(1))
#input_img_data = df.as_matrix(columns=['Image Data'])
#input_img_data.reshape([500,500,3])
input_img_data = np.asarray(input_img_data)
return input_img_data
def train_output_fn(index=0,data_amnt = 1):
output_breed_data = dataFrameBuilder(data_amount=data_amnt,
start_index=index,
ret_input=False,
ret_output=True)
#return df.as_matrix(columns=['Breed'])
output_breed_data = np.asarray(output_breed_data)
return output_breed_data
'''
batch_size
the number of samples returned
features
either 'train' to use training data
or 'test' to return testing data
'''
def generator(batch_size, features = train):
# Create empty arrays to contain batch of features and labels#
batch_features = np.zeros((batch_size, 500, 500, 3))
batch_labels = np.zeros((batch_size,120))
while True:
for i in range(batch_size):
# choose random index in features
index= random.choice([len(features),1])
batch_features[i] = train_input_fn(index=index, data_amnt=1)
batch_labels[i] = train_output_fn(index=index, data_amnt=1)
yield batch_features, batch_labels
# +
inception_v3 = tf.keras.applications.InceptionV3(include_top=False,
input_tensor=x,
classes=120)
#set imagedata to channels_last for best performance
# -
# add a global spatial average pooling layer
output_layer = inception_v3.output
output_layer = tf.keras.layers.GlobalAveragePooling2D()(output_layer)
# let's add a fully-connected layer
output_layer = tf.keras.layers.Dense(1024, activation='relu')(output_layer)
# and a logistic layer -- let's say we have 200 classes
predictions = tf.keras.layers.Dense(120, activation='softmax')(output_layer)
# this is the model we will train
model = tf.keras.Model(inputs=inception_v3.input, outputs=predictions)
for i, layer in enumerate(model.layers):
print(i, layer.name)
config = tf.ConfigProto(allow_soft_placement=True)
config.gpu_options.allocator_type = 'BFC'
config.gpu_options.per_process_gpu_memory_fraction = 0.40
config.gpu_options.allow_growth = True
# +
tf.reset_default_graph()
sess = tf.InteractiveSession(config=tf.ConfigProto(log_device_placement=True))
#with tf.Session(config=tf.ConfigProto(log_device_placement=True)) as ss:
x = tf.keras.layers.Input(shape=(500,500,3), batch_size=batch_size,name='input_data',dtype='float32')
y = tf.placeholder(tf.float32, shape=[None, 120,1], name='correct_labels')
#x=tf.placeholder(tf.float32, shape=[500,500,3],name='input_data')
y_pred = tf.placeholder(tf.float32, shape=[None,120,1], name='predicted_labels')
inception_v3 = tf.keras.applications.InceptionV3(include_top=False,
weights='imagenet',
input_tensor=x,
classes=120)
#steps for adding a new output layer
output_layer = inception_v3.output
output_layer = tf.keras.layers.GlobalAveragePooling2D()(output_layer) #replace the current global avg pool 2d
output_layer = tf.keras.layers.Dense(1024, activation='relu')(output_layer)
predictions = tf.keras.layers.Dense(120, activation='softmax')(output_layer) #120 classes in the new model
model = tf.keras.Model(inputs=inception_v3.input, outputs=predictions)
model.compile(loss=tf.keras.losses.categorical_crossentropy,
optimizer='sgd')
#img_data = train_input_fn(data_amnt=batch_size)
#breed_data = train_output_fn(data_amnt=batch_size)
#model.fit(x=img_data, y=breed_data, batch_size=batch_size)
#inception_v3.fit_generator(generator)
#model.fit_generator(generator(features, labels, batch_size), samples_per_epoch=50, nb_epoch=10)
#print(img_data[0])
model.fit_generator(generator(batch_size), steps_per_epoch=10, epochs=50)
index = 11
img_data = train_input_fn(index=index, data_amnt=batch_size)
breed_data = train_output_fn(index=index, data_amnt=batch_size)
#sample_weight=np.transpose(np.ones(120, dtype='float32'))
#model.evaluate(x=img_data,y=breed_data,batch_size=batch_size, sample_weight=sample_weight)
model.evaluate(x=img_data, y=breed_data,batch_size=batch_size)
# -
def test_input_fn(index=0, data_amnt = 1):
input_img_data = dataFrameBuilder(data_amount=data_amnt,
start_index=index,
dir="../data/preprocessed_data/Test/")
input_img_data = np.asarray(input_img_data)
return input_img_data
# +
'''prepare the labels for the output dataframe'''
df_labels = ['id'] #set up the dataframe column labels
for label in labels_np: #append each breed as a new column
df_labels.append(label[0])
df_labels_np = np.asarray(list(df_labels)).T
# +
data_files = os.listdir("../data/preprocessed_data/Test/") #get a list of all filenames from Test dir
df_data = [] # a list to append all individual id/prediction data to
index = 0 #keep track of which position the file is in the data_files list
for file in data_files:
file_data = [] #the individual file's id and all predictions
file_id = get_id_from_filename(file) #the individual file's id
file_pred_list = model.predict(x=test_input_fn(index=index), batch_size = batch_size) #a list of all predictions for the single file
file_data.append(file_id) #id is first in the list of data
for prediction in file_pred_list[0]: #append all preditions in order
file_data.append(prediction)
index += 1 #next file
if(index >= 10): #for testing/debugging purposes --remove this later
break;
df_data.append(file_data)
# -
df = pd.DataFrame(df_data, columns=df_labels)
df.to_csv('../output.csv')
| src/inceptionClassifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="VXAz3pdWAQgm"
# # Practice with Variables and Types
#
# As you've seen, one of the most basic ways to use Python is as a calculator. Let's step up our calculation game by using Python to solve a math problem.
#
# Let's say you opened up a savings account at your local bank and to start you deposited $100. While your money is in the bank it is *accruing interest* at a rate of 10 percent per year. Now, you want to know how much money you will have at the end of 7 years, assuming you never withdraw or deposit anything from your savings account. The formula for calculating this value is:
#
# <center>total = start_balance * ( 1 + interest_rate )<sup>years</sup></center>
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 410, "status": "ok", "timestamp": 1563150108891, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-WQsOgn6cXFg/AAAAAAAAAAI/AAAAAAAAAVA/ol7X28qyOHc/s64/photo.jpg", "userId": "07259925140867310896"}, "user_tz": 240} id="_qEZ_Vb9AQgq" outputId="7dab0d18-6706-4339-8cd9-85c401eeebb4"
# assign your deposit amount to the variable start_balance
# assign your interest rate to the variable interest_rate (TIP: How do we represent percentages in math?)
# print the types of both start_balance and interest_rate. Make sure they match what you expect
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 649, "status": "ok", "timestamp": 1563150109163, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-WQsOgn6cXFg/AAAAAAAAAAI/AAAAAAAAAVA/ol7X28qyOHc/s64/photo.jpg", "userId": "07259925140867310896"}, "user_tz": 240} id="c-Od3EhdAQg6" outputId="c2b7f31b-5163-4c88-a994-87a25c2f40db"
# assign the number of savings years to the variable years
# calculate the result of your savings problem to the variable total.
# HINT 1: Remember your order of operations!
# HINT 2: Use Google for the Python exponent mathematical operator
# Google is your best friend in coding - don't be afraid to use it!
# print the value of the total variable
# print the type of the total variable. Is this what we expect? Why?
# + [markdown] colab_type="text" id="gNstSD7IAQhE"
# Now that we've had some practice with numerical types, let's get some practice with the other types we learned!
#
# First, let's see if we were able to double our money with our 7 years of interest and patience.
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 632, "status": "ok", "timestamp": 1563150109165, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-WQsOgn6cXFg/AAAAAAAAAAI/AAAAAAAAAVA/ol7X28qyOHc/s64/photo.jpg", "userId": "07259925140867310896"}, "user_tz": 240} id="G6vuBsQpAQhG" outputId="8ad03933-7263-489c-9a50-325deac3d9c8"
# determine if total is greater than or equal to double our start balance
# assign this value to the variable doubled and print it
# What type do we expect this to be? Let's print the type of the variable doubled
# + [markdown] colab_type="text" id="vOGr_zBUAQhR"
# Now, let's play the name game.
# + colab={"base_uri": "https://localhost:8080/", "height": 232} colab_type="code" executionInfo={"elapsed": 633, "status": "error", "timestamp": 1563150109187, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-WQsOgn6cXFg/AAAAAAAAAAI/AAAAAAAAAVA/ol7X28qyOHc/s64/photo.jpg", "userId": "07259925140867310896"}, "user_tz": 240} id="LX0J-tAJAQhT" outputId="88dc461f-a37c-49f7-de7b-61a1a87337e0"
# create a variable called first_name and assign it to your first name as a string
# create a variable called last_name and assign it to your last name as a string
# Now, concatenate (i.e. add) first_name with last_name and assign it a variable called full_name
# print full_name. Does this look like you expect? How can you fix the formatting so it makes sense?
# now try to add full_name to the variable start_balance from above. Does this work?
# read the error message you get carefully!
# learning how to read error messages will give you the keys to fix mistakes (i.e. bugs) in your code
# + [markdown] colab_type="text" id="uzE681JbAQhX"
# Lastly, let's use Python to create a personalized investment report.
#
# We know that we cannot simply add integers, floats, or booleans to strings. But in terms of printing, there are a few ways we can handle the formatting to enable printing of mixed types in the same message.
#
# ### Method 1 - type conversion
# Numeric types and booleans can be converted to strings using the `str()` function.
# + colab={} colab_type="code" id="DLhNTBX9AQhY"
print('ABC' + str(123))
print('LMNOP' + str(8.910))
print('XYZ' + str(True))
# + [markdown] colab_type="text" id="Ki6NJbpXAQhf"
# Some strings can also be converted to other types (if they make sense) using the `int()`, `float()`, and `bool()` functions.
# + colab={} colab_type="code" id="I1hLe-FvAQhg"
print(4 + int('5'))
print(1.2 + float("3.4"))
print(False + bool('True'))
# + [markdown] colab_type="text" id="laceLTnuAQhm"
# ### Method 2 - multiple parameters in `print` function
# The `print` function takes multiple parameters (i.e. inputs), which can be of mixed type and are separated by commas.
# + colab={} colab_type="code" id="aaSBlzpFAQhn"
print("I have", 1 , "dog and", 2, "cats")
print("I ate", 2.5, "ice cream sandwiches")
print("Those statements are both", True)
# + [markdown] colab_type="text" id="HWDI495EAQhr"
# Notice that when using multiple parameters in the `print` functon, Python automatically puts spaces between the different inputs, but when using the type conversion method there are no automatic spaces added.
#
# Now, use either of the above methods to print a report that matches the following format:
#
# Customer Name: <NAME>
# Starting Balance: $100
# Years Saved: 7
# Ending Balance: $194.87
# Doubled Starting: False
#
# Remember: Use the variable names we defined above whenever you can!
# + colab={} colab_type="code" id="qDU2ZGrTAQhs"
# HINT: remember the spacing rules above!
# + colab={} colab_type="code" id="pKQFnZnyAQh1"
| Practices/Practice03_Variables_Types.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="QnXzcZvdc-r6" colab_type="text"
# In this notebook we will demostrate how to perform tokenization,stemming,lemmatization and pos_tagging using libraries like [spacy](https://spacy.io/) and [nltk](https://www.nltk.org/)
# + id="R3xEmJpRc5r8" colab_type="code" colab={}
#This will be our corpus which we will work on
corpus_original = "Need to finalize the demo corpus which will be used for this notebook and it should be done soon !!. It should be done by the ending of this month. But will it? This notebook has been run 4 times !!"
corpus = "Need to finalize the demo corpus which will be used for this notebook & should be done soon !!. It should be done by the ending of this month. But will it? This notebook has been run 4 times !!"
# + id="KHh_33IopPTf" colab_type="code" outputId="37c2db65-d10d-4254-80df-48a3f48713cd" colab={"base_uri": "https://localhost:8080/", "height": 34}
#lower case the corpus
corpus = corpus.lower()
print(corpus)
# + id="3yaGf8RiqgBM" colab_type="code" outputId="5856d53c-63e9-4045-dc53-96bb98fba30e" colab={"base_uri": "https://localhost:8080/", "height": 34}
#removing digits in the corpus
import re
corpus = re.sub(r'\d+','', corpus)
print(corpus)
# + id="v5Q--GItqzfu" colab_type="code" outputId="e0581860-574e-4f1e-f517-29b60b438ec8" colab={"base_uri": "https://localhost:8080/", "height": 34}
#removing punctuations
import string
corpus = corpus.translate(str.maketrans('', '', string.punctuation))
print(corpus)
# + id="zmANqee9rK4N" colab_type="code" outputId="227843ab-d55f-4c9a-adff-cc253eb36450" colab={"base_uri": "https://localhost:8080/", "height": 34}
#removing trailing whitespaces
corpus = corpus.strip()
corpus
# + id="KMuHZTpy9X_u" colab_type="code" outputId="9ec1f466-2e24-4262-b48c-a6c36f3dcb7a" colab={"base_uri": "https://localhost:8080/", "height": 872}
# !pip install spacy
# !python -m spacy download en
# + [markdown] id="nfJx3MnVj_ph" colab_type="text"
# ### Tokenizing the text
# + id="OUz580k2sMqf" colab_type="code" outputId="5195f746-7e75-4cfa-d763-4856359baf05" colab={"base_uri": "https://localhost:8080/", "height": 250}
from pprint import pprint
##NLTK
import nltk
from nltk.corpus import stopwords
nltk.download('stopwords')
nltk.download('punkt')
from nltk.tokenize import word_tokenize
stop_words_nltk = set(stopwords.words('english'))
tokenized_corpus_nltk = word_tokenize(corpus)
print("\nNLTK\nTokenized corpus:",tokenized_corpus)
tokenized_corpus_without_stopwords = [i for i in tokenized_corpus_nltk if not i in stop_words_nltk]
print("Tokenized corpus without stopwords:",tokenized_corpus_without_stopwords)
##SPACY
from spacy.lang.en.stop_words import STOP_WORDS
import spacy
spacy_model = spacy.load('en_core_web_sm')
stopwords_spacy = spacy_model.Defaults.stop_words
print("\nSpacy:")
text_tokens = word_tokenize(corpus)
print("Tokenized Corpus:",text_tokens)
tokens_without_sw= [word for word in tokenized_corpus if not word in stopwords_spacy]
print("Tokenized corpus without stopwords",tokens_without_sw)
# + [markdown] id="eRH_ltkD-HpA" colab_type="text"
# Notice the difference output after stopword removal using nltk and spacy
# + [markdown] id="tGcwD1JlkEao" colab_type="text"
# ### Stemming
# + id="ibEpzcv0sdW8" colab_type="code" outputId="f40aa595-f67f-4042-e9bf-e5b4a160d524" colab={"base_uri": "https://localhost:8080/", "height": 87}
from nltk.stem import PorterStemmer
from nltk.tokenize import word_tokenize
stemmer= PorterStemmer()
print("Before Stemming:")
print(corpus)
print("After Stemming:")
for word in tokenized_corpus_nltk:
print(stemmer.stem(word),end=" ")
# + [markdown] id="9Wy6cwvYkJeR" colab_type="text"
# ### Lemmatization
# + id="27KvL4ZE-fqJ" colab_type="code" outputId="41dc046e-14dd-4d4f-83ca-2b3af4532593" colab={"base_uri": "https://localhost:8080/", "height": 70}
from nltk.stem import WordNetLemmatizer
from nltk.tokenize import word_tokenize
nltk.download('wordnet')
lemmatizer=WordNetLemmatizer()
for word in tokenized_corpus_nltk:
print(lemmatizer.lemmatize(word),end=" ")
# + [markdown] id="h8uCGA8ukMfQ" colab_type="text"
# ### POS Tagging
# + id="kZqBxLDz-6cu" colab_type="code" outputId="9b89af94-c433-4099-b1c3-3adefecae984" colab={"base_uri": "https://localhost:8080/", "height": 1000}
#POS tagging using spacy
doc = spacy_model(corpus_original)
print("POS Tagging using spacy:")
# Token and Tag
for token in doc:
print(token,":", token.pos_)
#pos tagging using nltk
# nltk.download('averaged_perceptron_tagger')
print("POS Tagging using NLTK:")
pprint(nltk.pos_tag(word_tokenize(corpus_original)))
# + [markdown] id="zWdmz6lFkpEI" colab_type="text"
# There are various other libraries you can use to perform these common pre-processing steps
# + id="SRmVqDq5_1C2" colab_type="code" colab={}
| Ch2/04_Tokenization_Stemming_lemmatization_stopword_postagging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Cross-Resonance Gate
#
# *Copyright (c) 2021 Institute for Quantum Computing, Baidu Inc. All Rights Reserved.*
# ## Outline
#
# This tutorial introduces how to generate optimized pulses for Cross-Resonance (CR) gate using Quanlse. Unlike the iSWAP and CZ gate implementation in previous tutorials, CR gate is implemented using an all-microwave drive. The outline of this tutorial is as follows:
# - Introduction
# - Preparation
# - Construct Hamiltonian
# - Generate and optimize pulses via Quanlse Cloud Service
# - Summary
# ## Introduction
#
# **Fundamentals**
#
# Unlike some of the other gates we have seen before, the Cross-Resonance (CR) gate only uses microwaves to implement the two-qubit interaction such that we could avoid noise due to magnetic flux. The physical realization of the CR gate includes two coupled qubits with fixed frequencies. This can be done by driving the control qubit at the frequency of the target qubit. This is shown in the figure below:
#
#
# 
#
#
#
# We will first look at the effective Hamiltonian of the system (for details, please refer to Ref. \[1\] ). In the doubly rotating frame, the effective Hamiltonian for cross-resonance effect in terms of the drive strength $A$, detuning $\Delta$, drive phase $\phi_0$, and coupling strength $g_{01}$ is given \[1\] (for simplicity, we choose $\hbar = 1$) :
#
#
# $$
# \hat{H}_{\rm eff} = \frac{A}{4\Delta}g_{01}(\cos{\phi_0}\hat{\sigma}_0^z\hat{\sigma}_1^x+\sin{\phi_0}\hat{\sigma}_0^z\hat{\sigma}_1^y).
# $$
#
# When $\phi_0=0$, the cross-resonance effect allows for effective coupling of $\hat{\sigma}^z_0\otimes\hat{\sigma}_1^x$. We can thus derive the time evolution matrix from the effective Hamiltonian above:
#
# $$
# U_{\rm CR}(\theta)=e^{-i\frac{\theta}{2}\hat{\sigma}^z_0\otimes\hat{\sigma}^x_1},
# $$
#
# where $\theta = \Omega_0 gt/(2\Delta)$ ($t$ is the gate time). We can see that the cross-resonance effect enables a conditional rotation on qubit 1 (target qubit) depending on the state of qubit 0 (control qubit).
#
#
# Following the derivation above, the matrix form of the CR gate is (refer to \[2\] for more details):
# $$
# U_{\rm CR}(\theta) = \begin{bmatrix}
# \cos{\frac{\theta}{2}} & -i\sin{\frac{\theta}{2}} & 0 & 0 \\
# -i\sin{\frac{\theta}{2}} & \cos{\frac{\theta}{2}} & 0 & 0 \\
# 0 & 0 & \cos{\frac{\theta}{2}} & i\sin{\frac{\theta}{2}} \\
# 0 & 0 & i\sin{\frac{\theta}{2}} & \cos{\frac{\theta}{2}}
# \end{bmatrix}.
# $$
#
#
# In particular, the matrix representation of a CR gate with $\theta = -\frac{\pi}{2}$ is:
#
# $$
# U_{\rm CR}(-\pi/2) = \frac{\sqrt{2}}{2}
# \begin{bmatrix}
# 1 & i & 0 & 0 \\
# i & 1 & 0 & 0 \\
# 0 & 0 & 1 & -i \\
# 0 & 0 & -i & 1
# \end{bmatrix}.
# $$
#
# **Application**
#
# Having analyzed some of the fundamentals of the CR gate, we now switch our focus to the applications of the CR gate in quantum computing - one of which is the implementation of a CNOT gate with a CR gate and two additional single-qubit gates.
#
# 
#
# In this tutorial, we will model the system consisting of two three-level qubits and apply the drive pulse to the control qubit (qubit $q_0$) at the frequency of the target qubit (qubit $q_1$). By performing a rotating wave approximation (RWA), the Hamiltonian can be expressed as (refer to \[1\] for more details):
#
# $$
# \hat{H}_{\rm sys} = (\omega_{\rm q0}-\omega_{\rm d})\hat{a}_{0}^{\dagger }\hat{a}_0 + (\omega_{\rm q1}-\omega_{\rm d})\hat{a}_1^\dagger \hat{a}_1 + \frac{\alpha_0}{2} \hat{a}^{\dagger2}_0\hat{a}^2_0 + \frac{\alpha_1}{2} \hat{a}^{\dagger2}_1\hat{a}^2_1+\frac{g}{2}(\hat{a}_0\hat{a}_1^\dagger + \hat{a}_0^\dagger\hat{a}_1) + \Omega_0^x(t)\frac{\hat{a}^\dagger_0+\hat{a}_0}{2}.
# $$
#
# Please refer to the following chart for symbols' definitions:
#
# | Notation | Definition |
# |:--------:|:----------:|
# |$\omega_{\rm qi}$ | qubit $q_i$'s frequency |
# |$\omega_{\rm d}$|drive frequency|
# |$\hat{a}_i^{\dagger}$ | creation operator |
# |$\hat{a}_i$ | annihilation operator |
# |$\alpha_i$| qubit $q_i$'s anharmonicity |
# | $g$ | coupling strength |
# | $\Omega_0^x$(t) | pulse on the x channel |
# ## Preparation
#
# After you have successfully installed Quanlse, you could run the Quanlse program below following this tutorial. To run this particular tutorial, you would need to import the following packages from Quanlse and other commonly-used Python libraries:
# +
# Import Hamiltonian-related module
from Quanlse.QHamiltonian import QHamiltonian as QHam
from Quanlse.QOperator import driveX, number, duff
# Import optimizer for the cross-resonance gate
from Quanlse.remoteOptimizer import remoteOptimizeCr
# Import tools to analyze the result
from Quanlse.Utils.Functions import project
# Import numpy and math
from numpy import round
from math import pi
# -
# ## Construct Hamiltonian
#
# Now, we need to construct the above Hamiltonian using Quanlse. In Quanlse, all information regarding a Hamiltonian is stored in a dictionary. We start by defining some of the basic parameters needed for constructing a Hamiltonian dictionary: the sampling period, the number of qubits in the system, and the system's energy levels to consider. To initialize our Hamiltonian dictionary, we call the function `QHamiltonian()` from the module `QHamiltonian`.
# +
# Sampling period
dt = 1.0
# Number of qubits
qubits = 2
# System energy level
level = 3
# Initialize the Hamiltonian
ham = QHam(subSysNum=qubits, sysLevel=level, dt=dt)
# -
# Now we can start constructing our Hamiltonian. Before we start, we would need to define a few constants to pass in as the function's arguments:
# Parameters setting
qubitArgs = {
"coupling": 0.0038 * (2 * pi), # Coupling of Q0 and Q1
"qubit_freq0": 5.114 * (2 * pi), # Frequency of Q0
"qubit_freq1": 4.914 * (2 * pi), # Frequency of Q1
"drive_freq0": 4.914 * (2 * pi), # Drive frequency on Q0
"drive_freq1": 4.914 * (2 * pi), # Drive frequency on Q1
"qubit_anharm0": -0.33 * (2 * pi), # Anharmonicity of Q0
"qubit_anharm1": -0.33 * (2 * pi) # Anharmonicity of Q1
}
# Now we need to add the following terms to the Hamiltonian dictionary we initilized earlier:
#
# $$
# \begin{align}
# \hat{H}_{\rm drift} &= (\omega_{\rm q0}-\omega_{\rm d}) \hat{a}_0^\dagger \hat{a}_0 + (\omega_{\rm q1}-\omega_{\rm d}) \hat{a}_1^\dagger \hat{a}_1 + \frac{\alpha_0}{2} \hat{a}_0^{\dagger}\hat{a}_0^{\dagger}\hat{a}_0
# \hat{a}_0 + \frac{\alpha_1}{2} \hat{a}_1^{\dagger}\hat{a}_1^{\dagger}\hat{a}_1 \hat{a}_1 , \\
# \hat{H}_{\rm coup} &= \frac{g_{01}}{2}(\hat{a}_0 \hat{a}_1^\dagger+\hat{a}^\dagger_0 \hat{a}_1). \\
# \end{align}
# $$
#
# In Quanlse's `QOperator` module, we have provided tools that would allow the users to construct the commonly used operators quickly. The detuning term $(\omega_{\rm q}-\omega_{\rm d})\hat{a}^\dagger \hat{a}$ and the anharmonicity term $\frac{\alpha}{2}\hat{a}^\dagger\hat{a}^\dagger \hat{a} \hat{a} $ can be respectively generated using `number(n)` and `duff(n)` from the `QOperator` module: the two functions `number(n)` and `duff(n)` return the $n \times n$ matrices for number operators and duffing operators. Thus we can directly add these terms to the Hamiltonian by using function `addDrift()`. The coupling term, which takes the form, $\frac{g}{2}(\hat{a}_i^\dagger\hat{a}_j+\hat{a}_i\hat{a}_j^\dagger$), can be directly added to the Hamiltonian using function `addCoupling()`.
# +
# Add drift term(s)
for qu in range(2):
# Add detuning term(s)
ham.addDrift(number, qu, (qubitArgs[f"qubit_freq{qu}"] - qubitArgs[f"drive_freq{qu}"]))
# Add anharmonicity term(s)
ham.addDrift(duff, qu, qubitArgs[f"qubit_anharm{qu}"] / 2)
# Add coupling term
ham.addCoupling([0, 1], qubitArgs["coupling"] / 2)
# -
# Noted that the optimizer in Quanlse will automatically add the control term:
#
# $$
# \hat{H}_{\rm ctrl}(t) = \Omega_0^x(t)\frac{\hat{a}^\dagger_0+\hat{a}_0}{2},
# $$
#
# thus we don't need to add this term manually.
# With the system Hamiltonian built, we can now move on to the optimization.
# ## Generate and optimize pulse via Quanlse Cloud Service
# The optimization process usually takes a long time to process on local devices; however, we provide a cloud service that could speed up this process significantly. To use the Quanlse Cloud Service, the users need to acquire a token from http://quantum-hub.baidu.com.
# +
# Import tools to get access to cloud service
from Quanlse import Define
# To use remoteOptimizerCr on cloud, paste your token (a string) here
Define.hubToken = ''
# -
# To find the optimized pulse for CR gate, we use the function `remoteOptimizeCr()`, which takes the Hamiltonian we had previously defined, amplitude's bound, gate time, maximum iterations, and target infidelity. By calling `remoteOptimizeCr()`, the user can submit the optimization task to the Quanlse's server. If the user wants to further mitigate the infidelity, we encourage trying an increased gate time `tg` (the duration of a CR gate is around 200 to 400 nanoseconds). Users can also try increasing the search space by setting larger `aBound` and `maxIter`.
#
# The gate infidelity for performance assessment throughout this tutorial is defined as ${\rm infid} = 1 - \frac{1}{d}\left|{\rm Tr}[U^\dagger_{\rm goal}P(U)]\right|$, where $U_{\rm goal}$ is exactly the target unitary transformation $U_{\rm CR}(-\pi/2)$; $d$ is the dimension of $U_{\rm goal}$; and $U$ is the unitary evolution of the three-level system defined previously. Note that $P(U)$ in particular describes the evolution projected to the computational subspace.
# +
# Set amplitude bound
aBound = (-1.0, 3.0)
# Run the optimization
gateJob, infidelity = remoteOptimizeCr(ham, aBound=aBound, tg=200, maxIter=5, targetInfidelity=0.005)
# -
# We can visualize the generated pulse using `plot()`. (details regarding `plot()` are covered in [single-qubit-gate.ipynb](https://quanlse.baidu.com/#/doc/tutorial-single-qubit))
# Print waves and the infidelity
gateJob.plot()
print(f'infidelity: {infidelity}')
# The users can also print the the projected evolution matrix $P(U)$ using `simulate()` and `project()_`:
# Print the projected evolution
result = ham.simulate(job=gateJob)
process2d = project(result[0]["unitary"], qubits, level, 2)
print("The projected evolution P(U):\n", round(process2d, 2))
# Moreover, for those interested in acquiring the numerical data of the generated pulse for each `dt`, use function `getPulseSequences()`, which takes a Hamiltonian dictionary and channels' names as parameters.
gateJob.generatePulseSequence(driveX(3), 0)
gateJob.plot()
# ## Summary
#
# From constructing the system Hamiltonian to generating an optimized pulse on Quanlse Cloud Service, we have successfully devised a pulse to implement a cross-resonace gate with high fidelity. The users could follow this link [tutorial-cr-gate.ipynb](https://github.com/baidu/Quanlse/blob/main/Tutorial/EN/tutorial-cr.ipynb) to the GitHub page of this Jupyter Notebook document and run this program for themselves. The users are encouraged to try parameter values different from this tutorial to obtain the optimal result.
# + [markdown] pycharm={"name": "#%%\n"}
# ## References
#
# \[1\] [<NAME>, and <NAME>. "Fully microwave-tunable universal gates in superconducting qubits with linear couplings and fixed transition frequencies." *Physical Review B* 81.13 (2010): 134507.](https://journals.aps.org/prb/abstract/10.1103/PhysRevB.81.134507)
#
# \[2\] [<NAME>., and <NAME>. Quantum Computation and Quantum Information: 10th Anniversary Edition. Cambridge University Press, 2010.](https://doi.org/10.1017/CBO9780511976667)
| Tutorial/EN/tutorial-cr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.display import display
import numpy as np
import pandas as pd
import scipy.stats
import seaborn
import xarray as xr
import plot
import util
# ## Load Data ##
sfa = xr.open_dataset("../models/sfa_tcga/sfa.nc").load()
mri_features = xr.open_dataset("../data/processed/mri-features-all.nc").load()
assert all(mri_features['case'] == sfa['case'])
mri_factors = xr.open_dataset("../data/processed/mri-features-all-fa.nc").load()
assert all(mri_factors['case'] == sfa['case'])
sfa = sfa.rename({'factor': 'gexp_factor'})
sfa['gexp_factor'] = ["GF{}".format(i+1) for i in range(len(sfa['gexp_factor']))]
mri_factors = mri_factors.rename({'factor': 'mri_factor'})
mri_factors['mri_factor'] = ["IF{}".format(i+1) for i in range(len(mri_factors['mri_factor']))]
# ## Correlation Factors—MRI features ##
# Compute correlation between all MRI features (except for patient number, Comment and Multifocal) and SFA factors (`factor_feature_cor`). We also compute the nominal p value (`factor_feature_cor_nom_p`) and Bonferroni adjusted p-value (`factor_feature_cor_p`).
numeric_mri_features = list(set(mri_features.keys()) - {'case', 'Comment', 'MultiFocal'})
mri_features_da = mri_features[numeric_mri_features].to_array('cad_feature')
fm_cor = util.cor(mri_features_da, sfa['factors'], 'case')
fm_cor['p'] = np.fmin(1, fm_cor['nominal_p'] * fm_cor['nominal_p'].size)
# Heatmap of correlations. All of them are very low.
plot.heatmap(fm_cor['correlation'], cmap='coolwarm', row_dendrogram=True)
# Heatmap of correlation with nominal p-values < 0.05. This is without multiple testing correction.
plot.heatmap(fm_cor['correlation'], mask=fm_cor['nominal_p'] > 0.05, cmap='coolwarm', row_dendrogram=True)
# None of the correlation are significant after multiple testing correction.
np.min(fm_cor['p'].values)
fm_rcor = util.cor(mri_features_da, sfa['factors'], 'case', method='spearman')
fm_rcor['p'] = np.fmin(1, fm_rcor['nominal_p'] * fm_rcor['nominal_p'].size)
# Heatmap of correlations. All of them are very low.
plot.heatmap(fm_rcor['correlation'], cmap='coolwarm', row_dendrogram=True)
# Heatmap of correlation with nominal p-values < 0.05. This is without multiple testing correction.
plot.heatmap(fm_rcor['correlation'], mask=fm_rcor['nominal_p'] > 0.05, cmap='coolwarm', row_dendrogram=True)
# None of the correlation are significant after multiple testing correction.
np.min(fm_rcor['p'].values)
# ## Factor-Factor Correlation ##
sfa_da = sfa['factors'].reindex_like(mri_factors['factors'])
ff_cor = util.cor(mri_factors['factors'], sfa_da, 'case')
ff_cor['p'] = np.fmin(1, ff_cor['nominal_p'] * ff_cor['nominal_p'].size)
# Heatmap of correlations. All of them are low.
plot.heatmap(ff_cor['correlation'], cmap='coolwarm', row_dendrogram=True, col_dendrogram=True)
# Heatmap of correlation with p-values < 0.05. This is without multiple testing correction.
plot.heatmap(ff_cor['correlation'], mask=ff_cor['p'] > 0.05, cmap='coolwarm', row_dendrogram=True, col_dendrogram=True)
ff_rcor = util.cor(mri_factors['factors'], sfa_da, 'case', method='spearman')
ff_rcor['p'] = np.minimum(ff_rcor['nominal_p'] * len(ff_rcor['nominal_p']), 1.0)
# Heatmap of correlations. All of them are on the low side.
plot.heatmap(ff_rcor['correlation'], cmap='coolwarm', row_dendrogram=True, col_dendrogram=True)
# Heatmap of correlation with p-values < 0.05.
plot.heatmap(ff_rcor['correlation'], mask=ff_rcor['p'] > 0.05, cmap='coolwarm', row_dendrogram=True, col_dendrogram=True)
plot.scatter(sfa_da.sel(gexp_factor='GF8'), mri_factors['factors'].sel(mri_factor='IF1'))
with plot.subplots() as (fig, ax):
seaborn.kdeplot(sfa_da.sel(gexp_factor='GF8'), mri_factors['factors'].sel(mri_factor='IF1'), ax=ax)
plot.scatter(sfa_da.sel(gexp_factor='GF1'), mri_factors['factors'].sel(mri_factor='IF7'))
with plot.subplots() as (fig, ax):
seaborn.kdeplot(sfa_da.sel(gexp_factor='GF1'), mri_factors['factors'].sel(mri_factor='IF7'),
ax=ax, shade=True, gridsize=250)
plot.scatter(sfa_da.sel(gexp_factor='GF1'), mri_factors['factors'].sel(mri_factor='IF1'))
with plot.subplots() as (fig, ax):
seaborn.kdeplot(sfa_da.sel(gexp_factor='GF1'), mri_factors['factors'].sel(mri_factor='IF1'),
ax=ax, shade=True, gridsize=250)
| notebooks/2017-03-01-tb-factor-mri-correlaton.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 class='alert alert-warning'> Aéroports du monde</h1>
#
#
# En prenant pour prétexte un fichier de données recensant un très grands nombre d'aéroports, nous allons travailler les méthodes vues en cours pour :
# - importer un fichier csv
# - sélectionner des données
# - fusionner des données
#
# L'objectif est d'obtenir une carte mondiale des aéroports.
#
# Pour fonctionner, ce notebook doit être dans le même répertoire (ou dossier) que le dossier `./airports/` qui contient deux fichiers CSV : `./airports/airports.csv` et `./airports/countries.csv`
#
# Il est conseillé d'avoir fait le notebook part1 avant de traiter ce notebook.
# # Pour commencer : fichier que nous allons exploiter
#
# C'est la même chose que dans le notebook part 1 on va disposer d'une table des aéroports, c'est à dire une liste de dictionnaires.
# +
import csv
def charger_fichier( nom_fic ):
"""
Permet de lire un fichier CSV en utilisant la ligne d'entêtes
Retourne une liste de dictionnaires.
"""
table = []
fichier_csv = open( nom_fic , "r", newline ="", encoding ="utf-8" )
lecteur_fichier = csv.DictReader( fichier_csv , delimiter =",")
for enreg in lecteur_fichier :
table.append (dict ( enreg )) # enreg est de type OrderedDict on le remet endict
fichier_csv.close()
return table
# -
table_aeroports = charger_fichier('./airports/airports.csv')
table_aeroports[0]
# <div class = "alert alert-danger">
#
#
# ### <NAME>
#
# Rappelez vous que les valeurs de tous les champs sont de type `str` et qu'il faut les convertir en `float` ou en `int` si besoin (et le besoin sera très souvent là).
#
# On appellera **table** une `list` de `dict` python (c'est à dire un tableau de p-uplets nommés en langage "algorithmique").
# # Créer des cartes facilement avec `folium`
# <div class = "alert alert-info">
#
#
# **Question**
#
# Exécuter la cellule de code ci-dessous sans chercher à lire le code.
# +
import folium
import math
import random
def mystere(dictionnaire):
'''
Etant donné un dictionnaire passé en argument retourne une chaîne de caractères
dont la spécification est l'objet d'une question de l'énoncé
'''
code = "<p> <style> table, tr, td {border: 1px solid pink;} </style> <table>"
for cle, valeur in dictionnaire.items():
code = code + ("<tr><td>" + cle + "</td>" + "<td>" + valeur + "</td></tr>")
code = code + "</table></p>"
return code
def generer_popup(dictionnaire):
contenu_de_la_popup = mystere(dictionnaire)
iframe = folium.IFrame(html = contenu_de_la_popup, width = 300, height = 200)
popup = folium.Popup(iframe, max_width = 500)
return popup
def ajouter_marqueur(carte, latitude, longitude, dictionnaire, couleur):
'''
carte : de type folium.Map
latitude et longitude : de type float
dictionnaire : de type dict avec clées et valeurs de type str
couleur : au format '#RRGGBB' où RR, GG, BB sont des entiers entre 0 et 255 en hexadécimal
représentant les composant Rouge, Verte et Bleue de la couleur
'''
radius = 4
folium.CircleMarker(
location = [latitude, longitude],
radius = radius,
popup = generer_popup(dictionnaire),
color = couleur,
fill = True
).add_to(carte)
# -
# Nous allons utiliser le module `folium` qui permet de générer des cartes géographiques interactives (zoomables) avec des marqueurs légendés (cliquables). Lorsqu'on cherche à générer une carte avec ce module le processus de conception est le suivant, qui se déroule en trois étapes :
#
# - Créer une "carte" (de type `folium.Map`)
# - Ajouter des "marqueurs" sur la "carte
# - En spécifiant latitude et longitude
# - En spécifiant les informations que l'on souhaite voir apparaitre lors d'un clic sur le marqueur
# - En spécifiant d'autres paramètres tels que la couleur du marqueur
# - Obtenir la carte en demandant sa représentation
# La fonction `ajouter_marqueur` permet de gérer l'étape 2 (pour un seul marqueur) et prend en paramètres :
# - une latitude (de type `float`)
# - une longitude (de type `float`)
# - une légende sous forme de dictionnaire (clés et valeurs de type `str`)
# - une couleur `"#RRGGBB"` où `RR`,`GG` et `BB` représentent les composantes rouge, verte et bleue en hexadécimal (entre `00` et `FF`)
#
# Elle permet d'ajouter un marqueur ayant la légende donnée, au point de coordonnées données, sur une folium.Map.
# +
#étape 1 : création de la carte
ma_carte = folium.Map(location=(47.5, 1), zoom_start=7)
#étape 2 : ajout de quatre marqueurs
ajouter_marqueur(ma_carte, 47.90, 1.90, {'Ville' : 'ORLEANS', 'Pop.' : '114644'}, "#FF0000")
ajouter_marqueur(ma_carte, 47.39, 0.68, {'Ville' : 'TOURS', 'Pop.' : '136252'}, "#880000")
ajouter_marqueur(ma_carte, 48.73, 1.36, {'Ville' : 'DREUX', 'Pop.' : '30836'}, "#00FFFF")
ajouter_marqueur(ma_carte, 46.81, 1.69, {'Ville' : 'CHATEAUROUX','Pop.' : '43732'}, "#88BB88")
ajouter_marqueur(ma_carte, 47.58, 1.33, {'Ville' : 'BLOIS','Pop.' : '45710'}, "yellow")
#étape 3 : affichage de la carte
ma_carte
# -
# <div class = "alert alert-info">
#
#
# **Question**
#
# Exécuter la cellule de code ci-dessus.
# <div class = "alert alert-info">
#
#
# **Question**
#
# Modifier le code ci-dessus afin d'ajouter un marqueur pour la ville de Blois de couleur jaune (latitude : 47.58, longitude : 1.33, population : 45710).
# <div class = "alert alert-warning">
#
#
# # Avant d'aller plus loin : Synthèse
#
# Nous avons vu comment accéder au fichier des aéroports et comment enrichir une carte avec des marqueurs. Nous allons désormais chercher à créer une carte avec les aéroports de la façon suivante :
#
# - La couleur du marqueur dépendra de l'altitude de l'aéroport (attention dans le fichier elle est exprimée en pieds):
# - entre 0 et 40 mètres : bleu
# - entre 40 et 500 mètres : vert
# - entre 500 et 1000 mètres : jaune
# - entre 1000 et 1500 mètres : rouge
# - au delà de 1500 mètres d'altitude : noir (équivalent : entre 1500 et 99999 mètres d'altitude)
#
#
# - La fenêtre pop-up contiendra les informations suivantes :
# - code IATA de l'aéroport
# - nom de l'aéroport
# - nom de la ville
# - code du pays
#
#
# Pour cela nous allons pour chacune des cinq plages d'altitude :
# - Créer une nouvelle table ne comportant que les aéroports ayant une altitude dans cette plage,
# - Pour chacun des aéroports de cette nouvelle table :
# - Créer un dictionnaire ne comportant que les quatre paires clé-valeur à indiquer dans la pop-up
# - Utiliser la fonction `ajouter_marqueur` pour placer l'aéroport correspondant sur la carte
#
# Bien entendu nous allons travailler avec des fonctions.
# # 1) Quelques fonctions à créer
table_aeroports = charger_fichier('./airports/airports.csv')
# <div class = "alert alert-info">
#
#
# **Question**
#
# Compléter la fonction `extraire_aeroports_dans_plage` qui prend en paramètres :
# - une table d'aéroports `table` comportant un champ `Altitude` exprimée en **pieds**,
# - une altitude minimale `alt_min` exprimée en **mètres**,
# - une altitude maximale `alt_max` exprimée en **mètres**,
#
# et renvoie une table comportant tous les aéroports ayant une altitude comprise entre `alt_min` (compris) et `alt_max` (non compris)
#
#
# **Remarque :** 1 pied = 0,3048 mètre, à utiliser pour convertir en mètres l'altitude des aéroports.
#
def extraire_aeroports_dans_plage(table, alt_min, alt_max):
table_extraite = []
for aero in table_aeroports:
if alt_min<= int(aero["Altitude"]) <= alt_max:
table_extraite.append(aero)
#à compléter pour répondre
return table_extraite
table_out=extraire_aeroports_dans_plage(table_aeroports, 3000, 3500)
for aero in table_out:
print(aero)
print(len(table_out))
# <div class = "alert alert-info">
#
#
# **Question**
#
# Compléter la fonction `creer_dict_popup` qui prend en paramètre :
#
# - un dictionnaire `aeroport` contenant les neuf champs `Airport ID`, `Name`, `City etc.` ,
#
# et renvoie un dictionnaire ne comportant que les quatre champs à afficher dans la pop_up : `Name`, `City`, `Country_Code`, `IATA` .
#
# Votre fonction devra vérifier l'assertion ci-dessous.
def creer_dict_popup(aeroport):
dico_extrait=dict()
for champ in ["Name","City","Country_Code","IATA"]:
dico_extrait[champ]=aeroport[champ]
return dico_extrait
assert(creer_dict_popup({'Airport_ID': '333096',
'Name': 'Qilian Airport',
'Type': 'medium_airport',
'City': 'Qilian',
'Country_Code': 'CN',
'IATA': 'HBQ',
'Latitude': '38.012',
'Longitude': '100.644',
'Altitude': '10377'})
==
{
'Name': 'Qilian Airport',
'City': 'Qilian',
'Country_Code': 'CN',
'IATA': 'HBQ'
}
)
# <div class = "alert alert-info">
#
#
# **Question**
#
# Compléter les fonctions `donner_latitude` et `donner_longitude` qui prennent en paramètre :
#
# - un dictionnaire `aeroport` contenant les neuf champs `Airport ID`, `Name`, `City etc.` ,
#
# et renvoient respectivement la latitude et la longitude de l'aéroport sous forme d'un flottant de type `float`.
#
#
# Votre fonction devra vérifier l'assertion ci-dessous.
# +
def donner_latitude(aeroport):
latitude=float(aeroport["Latitude"])
return latitude
def donner_longitude(aeroport):
longitude=float(aeroport["Longitude"])
return longitude
# -
assert(donner_latitude({'Airport_ID': '333096',
'Name': 'Qilian Airport',
'Type': 'medium_airport',
'City': 'Qilian',
'Country_Code': 'CN',
'IATA': 'HBQ',
'Latitude': '38.012',
'Longitude': '100.644',
'Altitude': '10377'})
==
38.012 )
# # 2) Une première carte
# <div class = "alert alert-info">
#
#
# **Question**
#
# On dispose désormais de cinq fonctions :
# - `ajouter_marqueur`
# - `extraire_aeroports_dans_plage`
# - `creer_dict_popup`
# - `donner_latitude`
# - `donner_longitude`
#
#
# 1) En utilisant ces cinq fonctions, compléter la cellule de code ci-dessous pour obtenir une carte avec les aéroports en bleu ayant une altitude entre 0 et 40 mètres.
#
#
# 2) Compléter le code pour placer sur la carte les 4 autres catégories d'aéroports (40 --> 500 vert, 500-1000 --> jaune, 1000-1500 --> rouge, 1500-99999 --> noir)
#
#
# On pourra :
# - soit créer une fonction qui reprend le code du 1) et qu'on appelera cinq fois,
# - soit faire 4 Copier-Coller (solution moins élégante)
# +
table_aeroports = charger_fichier('./airports/airports.csv')
#étape 1 : création de la carte
ma_carte = folium.Map(location=(45, 0), zoom_start=2)
#étape 2 : ajout des marqueurs
table_extraite = extraire_aeroports_dans_plage(table_aeroports,0,40)
for aero in table_extraite:
ajouter_marqueur(ma_carte, donner_latitude(aero), donner_longitude(aero), {'Ville' : aero["City"], 'Pays' : aero["Country_Code"]}, "#0000FF")
#à compléter (pas mal de ligne de code)
#étape 3 : affichage de la carte
ma_carte
# -
# # 3) Une amélioration qui nécessite une fusion de tables
# On souhaite afficher une variante de cette carte pour laquelle le code du nom du pays a été remplacé par son nom de pays selon la norme ISO-3166
#
# Pour ce dernier point, il convient de savoir que notre table aéroports respectait les noms de pays de la nomre ISO-3166 et que l'on dispose d'une table adéquate permettant d'assurer la correspondance :
# +
import csv
def charger_fichier( nom_fic ):
"""
Permet de lire un fichier CSV en utilisant la ligne d'entêtes
Retourne une liste de dictionnaires.
"""
table = []
fichier_csv = open( nom_fic , "r", newline ="", encoding ="utf-8" )
lecteur_fichier = csv.DictReader( fichier_csv , delimiter =",")
for enreg in lecteur_fichier :
table.append (dict ( enreg )) # enreg est de type OrderedDict on le remet endict
fichier_csv.close()
return table
# -
table_pays = charger_fichier('./airports/countries.csv')
table_aeroports = charger_fichier('./airports/airports.csv')
# Pour l'Andorre, le code alpha-2 est 'AD' (pour votre culture générale, il existe aussi un code alpha-3 sur 3 lettres):
table_pays[0]
# Nous allons fusionner la table des aéroports `table_aeroports` avec la table des pays `table_pays` afin d'obtenir une table fusionnée comportant tous les champs de la table `table_aeroports` plus le champ `Country_Name`
# <div class = "alert alert-info">
#
#
# **Question**
#
#
# Compléter le code de la fonction `fusionner` ci-dessous :
# - prenant en paramètre les tables `table_gauche` et `table_droite`
#
#
# - renvoyant une table `table_fusionnee` telle que :
# - chaque dictionnaire de la table fusionnee comporte tous les champs de la table gauche `table_gauche` et le champ `name` de la `table_droite` renommé en champ `Country_Name`,
#
# - avec le champ `Country_Code` utilisé comme champ de fusion pour `table_gauche` et `code` comme champ de fusion pour `table_droite`
#
#
# (`table_gauche` sera la table des aéroports, `table_droite` sera la table des pays).
#
# Voici un exemple d'enregistrement (de ligne) de la table fusionnée : on remarque le champ `Country_Name` en plus.
#
# ```
# {'Airport_ID': '2596',
# 'Name': 'Sørkjosen Airport',
# 'Type': 'medium_airport',
# 'City': 'Sørkjosen',
# 'Country_Code': 'NO',
# 'IATA': 'SOJ',
# 'Latitude': '69.786796569824',
# 'Longitude': '20.959400177002',
# 'Altitude': '16',
# 'Country_Name': 'Norway'}
# ```
#
# +
import copy
def fusionner(table_gauche, table_droite):
table_fusion = []
for gauche in table_gauche:
for droite in table_droite:
pass
#à compléter
return table_fusion
# -
table_fusionnee = fusionner (table_aeroports, table_pays)
table_fusionne[1500]
# <div class = "alert alert-info">
#
#
# **Question**
#
#
# Modifier le code ci-dessous (à un seul endroit) pour qu'il affiche les aéroports avec le nom des pays au lieu des codes des pays.
# +
def extraire_aeroports_dans_plage(table, alt_min, alt_max):
table_extraite = [aeroport for aeroport in table if
alt_min <= 0.3048 * float(aeroport['Altitude']) and
0.3048 * float(aeroport['Altitude']) < alt_max]
return table_extraite
def creer_dict_popup(aeroport):
dico_extrait = {key:val for key, val in aeroport.items()
if key in ['Name', 'City', 'Country_Code', 'IATA']}
return dico_extrait
def donner_latitude(aeroport):
return float(aeroport['Latitude'])
def donner_longitude(aeroport):
return float(aeroport['Longitude'])
#étape 0 : creer la table des aeroports fusionnee
table_pays = charger_fichier('./airports/countries.csv')
table_aeroports = charger_fichier('./airports/airports.csv')
table_aeroports = fusionner (table_aeroports, table_pays)
#étape 1 : création de la carte
ma_carte = folium.Map(location=(45, 0), zoom_start=2)
#étape 2 : ajout des marqueurs
def ajouter_plage_sur_carte(carte, table_des_aeroports, alt_min, alt_max, couleur):
plage = extraire_aeroports_dans_plage(table_des_aeroports, alt_min, alt_max)
for aeroport in plage :
dict_popup = creer_dict_popup(aeroport)
latitude = donner_latitude(aeroport)
longitude = donner_longitude(aeroport)
ajouter_marqueur(carte, latitude, longitude, dict_popup, couleur)
ajouter_plage_sur_carte(ma_carte, table_aeroports, 0, 40, '#0000FF')
ajouter_plage_sur_carte(ma_carte, table_aeroports, 40, 500, '#00FF00')
ajouter_plage_sur_carte(ma_carte, table_aeroports, 500, 1000, '#FFFF00')
ajouter_plage_sur_carte(ma_carte, table_aeroports, 1000, 1500, '#FF0000')
ajouter_plage_sur_carte(ma_carte, table_aeroports, 1500, 99999, '#000000')
#étape 3 : affichage de la carte
ma_carte
# -
| donnees_en_table/aeroports/airports_part2-correction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
import pandas as pd
# + pycharm={"name": "#%%\n"}
df = pd.read_csv('../../data/DATA.csv', delimiter=',')
# + pycharm={"name": "#%%\n"}
#drop rows without magnitude
df.drop(df[df['Magnitud'] == 'no calculable'].index, inplace = True)
len(df.index)
# + pycharm={"name": "#%%\n"}
df.to_csv("../../data/DATA_0.csv")
# + pycharm={"name": "#%%\n"}
df = pd.read_csv('../../data/DATA_0.csv', delimiter=',')
# + pycharm={"name": "#%%\n"}
# Remove unnecesary rows
df.drop(df.columns[0], axis=1, inplace=True)
df.drop(['Fecha', 'Hora','Referencia de localizacion','Estatus'],axis=1, inplace=True)
# + pycharm={"name": "#%%\n"}
# Remove and complete data accordingly
# Remove rows with Profundidad == 'en revision'
df=df.loc[df.Profundidad != 'en revision',:]
# Fix non float specifications, like Profundidad == 'menos de 1'
df.loc[(df.Profundidad == 'menos de 1'), 'Profundidad'] = 1.0
# + pycharm={"name": "#%%\n"}
# Specify data type of Profundidad
df.astype({'Profundidad': 'float'}).dtypes
# + pycharm={"name": "#%%\n"}
import pandas as pd
# + pycharm={"name": "#%%\n"}
df = pd.read_csv('../../data/DATA_1.csv', delimiter=',')
len(df.index)
# + pycharm={"name": "#%%\n"}
df.columns = ["Magnitude","Latitude","Longitude","Depth","Date","Time"]
# + pycharm={"name": "#%%\n"}
x = df['Date'] + ' ' + df['Time']
df["Datetime"] = pd.to_datetime(x, format='%Y/%m/%d %H:%M:%S')
df.drop(["Date","Time"], axis=1, inplace=True)
df.to_csv("../../data/DATA_2.csv", index=False)
| notebooks/data_prep/1_remove_empty_magnitudes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# name: python3
# ---
# # Betting Framework
#
# In this notebook, I am evaluating the score prediction models versus the betting odds of bookmakers. I identify the best betting opportunity (the one with the highest expected value) and then place a bet using risk management: using the Kelly criterion, I am riding the maximum growth curve.
import pandas as pd
import numpy as np
# Here I use *Bet365* as the benchmark bookmaker.
# +
odds_market = (
pd.read_csv('../data/betting/2021-22.csv')
.loc[:, [
"HomeTeam", "AwayTeam", "FTHG", "FTAG",
"B365H", "B365D", "B365A"]]
.rename(columns={
"HomeTeam": "team1",
"AwayTeam": "team2",
"FTHG": "score1",
"FTAG": "score2",
"B365H": "home_win",
"B365D": "draw",
"B365A": "away_win"})
)
odds_market = odds_market.replace({
'Brighton': 'Brighton and Hove Albion',
'Leicester': 'Leicester City',
'Leeds': 'Leeds United',
'Man City': 'Manchester City',
'Man United': 'Manchester United',
'Norwich': 'Norwich City',
'Tottenham': 'Tottenham Hotspur',
'West Ham': 'West Ham United',
'Wolves': 'Wolverhampton'
})
# +
# Get the model predictions.
outcome = pd.read_csv("../data/fivethirtyeight/spi_matches.csv")
outcome = (
outcome
.loc[
(outcome['league_id'] == 2411) |
(outcome['league_id'] == 2412)]
.dropna()
)
predictions_dixon_coles = (
pd.merge(
pd.read_csv("../data/predictions/fixtures/dixon_coles.csv"),
outcome.loc[:, ['team1', 'team2', 'date', 'score1', 'score2']],
how='left',
left_on=['team1', 'team2', 'date'],
right_on=['team1', 'team2', 'date'])
).dropna()
predictions_spi = pd.read_csv("../data/fivethirtyeight/spi_matches.csv")
predictions_spi = (
predictions_spi
.loc[predictions_spi['league_id'] == 2411]
.rename(columns={
"prob1": "home_win_p",
"probtie": "draw_p",
"prob2": "away_win_p"})
.loc[:, [
'team1', 'team2', 'home_win_p', 'draw_p',
'away_win_p', 'score1', 'score2']]
.loc[predictions_spi['season'] == 2021]
.loc[predictions_spi['score1'].notna()]
)
# +
df_dc = (
pd.merge(
predictions_dixon_coles.loc[:, [
'team1', 'team2', 'home_win_p', 'draw_p', 'away_win_p']],
odds_market,
how='left',
left_on=['team1', 'team2'],
right_on=['team1', 'team2'])
.dropna()
)
df_spi = (
pd.merge(
predictions_spi.loc[:, [
'team1', 'team2', 'home_win_p', 'draw_p', 'away_win_p']],
odds_market,
how='left',
left_on=['team1', 'team2'],
right_on=['team1', 'team2'])
.dropna()
)
# +
def excepted_value(p_win, gain):
return p_win * gain - (1 - p_win) * 0
df_dc['ev_h'] = df_dc.apply(
lambda row: excepted_value(row['home_win_p'], row['home_win']), axis=1)
df_dc['ev_d'] = df_dc.apply(
lambda row: excepted_value(row['draw_p'], row['draw']), axis=1)
df_dc['ev_a'] = df_dc.apply(
lambda row: excepted_value(row['away_win_p'], row['away_win']), axis=1)
df_spi['ev_h'] = df_spi.apply(
lambda row: excepted_value(row['home_win_p'], row['home_win']), axis=1)
df_spi['ev_d'] = df_spi.apply(
lambda row: excepted_value(row['draw_p'], row['draw']), axis=1)
df_spi['ev_a'] = df_spi.apply(
lambda row: excepted_value(row['away_win_p'], row['away_win']), axis=1)
# +
def kelly_criterion(p, q):
return p + (p - 1) / (q - 1)
df_dc['k_h'] = df_dc.apply(
lambda row: kelly_criterion(row['home_win_p'], row['home_win']), axis=1)
df_dc['k_d'] = df_dc.apply(
lambda row: kelly_criterion(row['draw_p'], row['draw']), axis=1)
df_dc['k_a'] = df_dc.apply(
lambda row: kelly_criterion(row['away_win_p'], row['away_win']), axis=1)
df_spi['k_h'] = df_spi.apply(
lambda row: kelly_criterion(row['home_win_p'], row['home_win']), axis=1)
df_spi['k_d'] = df_spi.apply(
lambda row: kelly_criterion(row['draw_p'], row['draw']), axis=1)
df_spi['k_a'] = df_spi.apply(
lambda row: kelly_criterion(row['away_win_p'], row['away_win']), axis=1)
# +
from fixtures.ranked_probability_score import match_outcome
df_dc['winner'] = match_outcome(df_dc)
df_spi['winner'] = match_outcome(df_spi)
# +
def bet(row):
global account_balance
# Get highest potential event
evs = np.array([row['ev_h'], row['ev_d'], row['ev_a']])
edge = np.argmax(evs)
# Ensure its profitable
if evs[edge] > 1:
# Get bet amount
k = np.array([row['k_h'], row['k_d'], row['k_a']])[edge]
# Ensure we bet a positive amount
if k > 0:
bet_value = k * account_balance
account_balance -= bet_value
# Get result of the bet
if edge == row['winner'] == 0:
account_balance += bet_value * row['home_win']
elif edge == row['winner'] == 1:
account_balance += bet_value * row['draw']
elif edge == row['winner'] == 2:
account_balance += bet_value * row['away_win']
return account_balance
account_balance = 100
df_dc['balance'] = df_dc.apply(bet, axis=1)
account_balance = 100
df_spi['balance'] = df_spi.apply(bet, axis=1)
# -
# Well, it turns out that my model is not yet ready to beat the odds. 🤣
# +
import matplotlib.pyplot as plt
import matplotlib as mpl
mpl.rcParams['figure.dpi'] = 300
from highlight_text import fig_text
body_font = "Open Sans"
watermark_font = "DejaVu Sans"
text_color = "w"
background = "#282B2F"
title_font = "DejaVu Sans"
mpl.rcParams['xtick.color'] = text_color
mpl.rcParams['ytick.color'] = text_color
mpl.rcParams['text.color'] = text_color
mpl.rcParams['axes.edgecolor'] = text_color
mpl.rcParams['xtick.labelsize'] = 6
mpl.rcParams['ytick.labelsize'] = 6
# +
fig, ax = plt.subplots(tight_layout=True)
fig.set_facecolor(background)
ax.patch.set_alpha(0)
ax.plot(df_dc['balance'], color='#dab6fc')
ax.plot(df_spi['balance'], color='#815ac0')
ax.set_xlabel('Games', fontsize=8, color=text_color)
ax.set_ylabel('RPS' , fontsize=8, color=text_color)
fig_text(
x=0.1, y=1.075,
s="Balance when Betting using Kelly Criterion Risk Management",
fontsize=12, fontfamily=title_font, color=text_color, alpha=1)
fig_text(
x=0.1, y=1.025,
s="<Dixon-Coles> & <SPI> Model",
highlight_textprops=[
{"color": '#dab6fc'},
{"color": '#815ac0'}
],
fontsize=12, fontfamily=title_font, color=text_color, alpha=1)
fig_text(
x=0.82, y=1.,
s="Updated <15-03-2022>",
highlight_textprops=[{"fontweight": "bold"}],
fontsize=6, fontfamily=title_font, color=text_color, alpha=1)
fig_text(
x=0.8, y=0.01,
s="Created by <<NAME>>",
highlight_textprops=[{"fontstyle": "italic"}],
fontsize=6, fontfamily=watermark_font, color=text_color)
plt.show()
# -
| modeling/fixtures/betting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''bias-vision-language-yfXTBDV4'': pipenv)'
# language: python
# name: python3
# ---
# +
from importlib import reload
import json
import os, sys
from src import PATHS
sys.path.insert(0, os.path.join(PATHS.BASE, "wip", "hugo", "src"))
import parse_config
import numpy as np
import torch
from torch.utils.data import DataLoader, Dataset
import pandas as pd
import matplotlib.pyplot as plt
from itertools import product as iter_product
import src, src.debias, src.models, src.ranking, src.datasets, src.data_utils
from src.models import model_loader
if torch.cuda.device_count() > 1:
use_device_id = int(input(f"Choose cuda index, from [0-{torch.cuda.device_count()-1}]: ").strip())
else: use_device_id = 0
use_device = "cuda:"+str(use_device_id) if torch.cuda.is_available() else "cpu"
if not torch.cuda.is_available():
input("CUDA isn't available, so using cpu. Please press any key to confirm this isn't an error: \n")
print("Using device", use_device)
torch.cuda.set_device(use_device_id)
with open(src.PATHS.TRAINED_MODELS.TEST_PROMPTS, mode="r") as _test_promptsfile:
test_prompts_data = json.load(_test_promptsfile)
# + pycharm={"name": "#%%\n"}
debias_class = "gender"
experiment_results = pd.DataFrame()
test_prompts = test_prompts_data[debias_class]
test_prompts_df = pd.DataFrame({"prompt": test_prompts})
test_prompts_df["group"] = debias_class
test_prompts_df
# + pycharm={"name": "#%%\n"}
eval_dss = {"gender": [("FairFace", "val"), ("UTKFace", "val")], "race": [("FairFace", "val"), ("UTKFace", "val")]}
evaluations = ["maxskew", "ndkl", "clip_audit"]
perf_evaluations = ["cifar10", "flickr1k", "cifar100"] # flickr1k, cifar100, cifar10
all_experiment_results = pd.DataFrame()
clip_audit_results = pd.DataFrame()
batch_sz = 256
try:
with torch.cuda.device(use_device_id):
for model_name in src.models.VALID_MODELS:
print(model_name)
model, preprocess, tokenizer, model_alias = model_loader(model_name, device=use_device, jit=True)
if model_name.startswith("m-bain/"):
model.logit_scale = torch.tensor(-1, dtype=torch.float32, device=use_device)
if "clip_audit" in evaluations:
ca_prompts = test_prompts_data["clip_audit"]
ca_ds = src.datasets.FairFace(iat_type="race", lazy=True, _n_samples=None, transforms=preprocess, mode="val")
ca_dl = DataLoader(ca_ds, batch_size=batch_sz, shuffle=False, num_workers=8) # Shuffling ISN'T(!) reflected in the cache
ca_res = src.ranking.do_clip_audit(ca_dl, ca_prompts, model, model_alias, tokenizer, preprocess, use_device, use_templates=True)
for k, v in {"model_name": model_alias, "dataset": "FairFaceVal",
"evaluation": "clip_audit"}.items():
ca_res[k] = v
clip_audit_results = clip_audit_results.append(ca_res, ignore_index=True)
print("Done with clip audit")
for debias_class in {"gender", "race"}:
experiment_results = pd.DataFrame()
test_prompts = test_prompts_data[debias_class]
test_prompts_df = pd.DataFrame({"prompt": test_prompts})
test_prompts_df["group"] = debias_class
for perf_eval in perf_evaluations:
perf_res = {"model_name": model_alias, "dataset": perf_eval,
"evaluation": perf_eval, "debias_class": debias_class, "mean": src.debias.run_perf_eval(perf_eval, model, tokenizer, preprocess, use_device)}
experiment_results = experiment_results.append(pd.DataFrame([perf_res]), ignore_index=True)
n_imgs = None # First run populates cache, thus run with None first, later runs can reduce number
for dset_name, dset_mode in eval_dss[debias_class]:
ds = getattr(src.datasets, dset_name)(iat_type=debias_class, lazy=True, _n_samples=n_imgs, transforms=preprocess, mode=dset_mode)
dl = DataLoader(ds, batch_size=batch_sz, shuffle=False, num_workers=8) # Shuffling ISN'T(!) reflected in the cache
for evaluation in evaluations:
if evaluation == "clip_audit": continue
model.eval()
_res = src.debias.run_bias_eval(evaluation, test_prompts_df, model, model_alias, tokenizer, dl, use_device, cache_suffix="")
_res = src.debias.mean_of_bias_eval(_res, evaluation, "dem_par")
res = {}
for key, val in _res.items():
for rename in ["mean_", "std_"]:
if key.startswith(rename):
res[rename[:-1]] = val
break
else:
res[key] = val
res["model_name"] = model_alias
res["dataset"] = dset_name+dset_mode.capitalize()
res["evaluation"] = evaluation
experiment_results = experiment_results.append(pd.DataFrame([res]), ignore_index=True)
experiment_results["debias_class"] = debias_class
all_experiment_results = all_experiment_results.append(experiment_results)
#del model, preprocess, tokenizer
finally:
result_name = f"untrained_test_bias_results.csv"
ca_result_name = f"untrained_test_clip_audit_results.csv"
all_experiment_results.to_csv(os.path.join(src.PATHS.PLOTS.BASE, result_name))
clip_audit_results.to_csv(os.path.join(src.PATHS.PLOTS.BASE, ca_result_name))
# + pycharm={"name": "#%%\n"}
display(clip_audit_results)
display(all_experiment_results)
| ranking_experiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Known error: This notebook requires [yt](https://yt-project.org/) to visualize the results. Yt needs to be updated to work properly first. Updates are currently being made to yt's frontends to make this PyNE integration work.
# +
import requests
from os import path, getcwd, remove
from numpy import linspace, bitwise_or
from pyne.mesh import Mesh, NativeMeshTag
from pyne.dagmc import load, discretize_geom
from yt.config import ytcfg; ytcfg["yt","suppressStreamLogging"] = "True"
from yt.frontends.moab.api import PyneMoabHex8Dataset
from yt.visualization.plot_window import SlicePlot
# -
faceted_file = path.join(getcwd(), 'teapot.h5m')
if not path.isfile(faceted_file):
url = "http://data.pyne.io/teapot.h5m"
r = requests.get(url)
with open("teapot.h5m", "wb") as outfile:
outfile.write(r.content)
load(faceted_file)
# +
num_divisions = 50
num_rays = 3
coords0 = linspace(-6, 6, num_divisions)
coords1 = linspace(0, 7, num_divisions)
coords2 = linspace(-4, 4, num_divisions)
# -
m = Mesh(structured=True, structured_coords=[coords0, coords1, coords2], structured_ordering='zyx')
results = discretize_geom(m, num_rays=num_rays, grid=False)
m.vols = NativeMeshTag(1, float)
mask = bitwise_or(results['cell'] == 1, results['cell'] == 2)
m.vols[results['idx'][mask]] = results[mask]['vol_frac']
pf = PyneMoabHex8Dataset(m)
s = SlicePlot(pf, 'z', 'vols')
s.display()
| examples/discretized_teapot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Anatomy of a figure
#
#
# This figure shows the name of several matplotlib elements composing a figure
#
# +
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.ticker import AutoMinorLocator, MultipleLocator, FuncFormatter
np.random.seed(19680801)
X = np.linspace(0.5, 3.5, 100)
Y1 = 3+np.cos(X)
Y2 = 1+np.cos(1+X/0.75)/2
Y3 = np.random.uniform(Y1, Y2, len(X))
fig = plt.figure(figsize=(8, 8))
ax = fig.add_subplot(1, 1, 1, aspect=1)
def minor_tick(x, pos):
if not x % 1.0:
return ""
return "%.2f" % x
ax.xaxis.set_major_locator(MultipleLocator(1.000))
ax.xaxis.set_minor_locator(AutoMinorLocator(4))
ax.yaxis.set_major_locator(MultipleLocator(1.000))
ax.yaxis.set_minor_locator(AutoMinorLocator(4))
ax.xaxis.set_minor_formatter(FuncFormatter(minor_tick))
ax.set_xlim(0, 4)
ax.set_ylim(0, 4)
ax.tick_params(which='major', width=1.0)
ax.tick_params(which='major', length=10)
ax.tick_params(which='minor', width=1.0, labelsize=10)
ax.tick_params(which='minor', length=5, labelsize=10, labelcolor='0.25')
ax.grid(linestyle="--", linewidth=0.5, color='.25', zorder=-10)
ax.plot(X, Y1, c=(0.25, 0.25, 1.00), lw=2, label="Blue signal", zorder=10)
ax.plot(X, Y2, c=(1.00, 0.25, 0.25), lw=2, label="Red signal")
ax.plot(X, Y3, linewidth=0,
marker='o', markerfacecolor='w', markeredgecolor='k')
ax.set_title("Anatomy of a figure", fontsize=20, verticalalignment='bottom')
ax.set_xlabel("X axis label")
ax.set_ylabel("Y axis label")
ax.legend()
def circle(x, y, radius=0.15):
from matplotlib.patches import Circle
from matplotlib.patheffects import withStroke
circle = Circle((x, y), radius, clip_on=False, zorder=10, linewidth=1,
edgecolor='black', facecolor=(0, 0, 0, .0125),
path_effects=[withStroke(linewidth=5, foreground='w')])
ax.add_artist(circle)
def text(x, y, text):
ax.text(x, y, text, backgroundcolor="white",
ha='center', va='top', weight='bold', color='blue')
# Minor tick
circle(0.50, -0.10)
text(0.50, -0.32, "Minor tick label")
# Major tick
circle(-0.03, 4.00)
text(0.03, 3.80, "Major tick")
# Minor tick
circle(0.00, 3.50)
text(0.00, 3.30, "Minor tick")
# Major tick label
circle(-0.15, 3.00)
text(-0.15, 2.80, "Major tick label")
# X Label
circle(1.80, -0.27)
text(1.80, -0.45, "X axis label")
# Y Label
circle(-0.27, 1.80)
text(-0.27, 1.6, "Y axis label")
# Title
circle(1.60, 4.13)
text(1.60, 3.93, "Title")
# Blue plot
circle(1.75, 2.80)
text(1.75, 2.60, "Line\n(line plot)")
# Red plot
circle(1.20, 0.60)
text(1.20, 0.40, "Line\n(line plot)")
# Scatter plot
circle(3.20, 1.75)
text(3.20, 1.55, "Markers\n(scatter plot)")
# Grid
circle(3.00, 3.00)
text(3.00, 2.80, "Grid")
# Legend
circle(3.70, 3.80)
text(3.70, 3.60, "Legend")
# Axes
circle(0.5, 0.5)
text(0.5, 0.3, "Axes")
# Figure
circle(-0.3, 0.65)
text(-0.3, 0.45, "Figure")
color = 'blue'
ax.annotate('Spines', xy=(4.0, 0.35), xytext=(3.3, 0.5),
weight='bold', color=color,
arrowprops=dict(arrowstyle='->',
connectionstyle="arc3",
color=color))
ax.annotate('', xy=(3.15, 0.0), xytext=(3.45, 0.45),
weight='bold', color=color,
arrowprops=dict(arrowstyle='->',
connectionstyle="arc3",
color=color))
ax.text(4.0, -0.4, "Made with http://matplotlib.org",
fontsize=10, ha="right", color='.5')
plt.show()
| matplotlib/gallery_jupyter/showcase/anatomy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sktime.load_data import load_from_tsfile_to_dataframe
cache_path = "C:/temp/sktime_temp_data/"
dataset_name = "JapaneseVowels"
suffix = "_TRAIN.ts"
train_x, train_y = load_from_tsfile_to_dataframe(cache_path + dataset_name + "/", dataset_name + suffix)
type(train_x)
type(train_x.iloc[0])
type(train_x.iloc[0][0])
train_x
| examples/example of loading into pandas DataFrame.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.12 ('base')
# language: python
# name: python3
# ---
# +
from selenium.webdriver import Chrome
from selenium.webdriver.chrome.options import Options
opts = Options()
opts.set_headless()
assert opts.headless # Operating in headless mode
browser = Chrome(options=opts)
browser.get('https://duckduckgo.com')
# -
search_form = browser.find_element_by_id('search_form_input_homepage')
search_form.send_keys('<PASSWORD>')
search_form.submit()
results = browser.find_elements_by_class_name('result')
print(results[0].text)
test = browser.find_element_by_class_name('badge-link__title')
test
browser.close()
quit()
| RealPythonSeleniumExample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Центр непрерывного образования
#
# # Программа «Python для автоматизации и анализа данных»
#
# Неделя 3 - 1
#
# *<NAME>, НИУ ВШЭ*
#
# ## Задачи
# ### Задача 1
#
# Дана строка, состоящая из слов. Сделать из нее аббревиатуру с помощью списковых включений
#
# **Вход:** "Комитет Государственной Безопасности"
#
# **Выход:** "КГБ"
# +
text = "Комитет Государственной Безопасности"
#Решение
res = [i[0] for i in text.split()]
print(*res, sep = '')
# -
# ### Задача 2
#
# На вход принимается число N.
#
# Выведите на печать следующий паттерн:
#
# 1
#
# 1 2
#
# 1 2 3
#
# 1 2 3 4
#
# ...
#
# 1 ... N
#
# **Это не так просто)**
N = 10
tes = [[k for k in range(1, i+1)]for i in range(1, N+1)]
for elem in tes:
print(*elem)
# ### Задача 3
#
# Дана строка. Замените в ней все буквы А на # с помощью спискового включения.
#
input_str = 'АБАЖУР, АБАЗИНСКИЙ, АБАЗИНЫ, АББАТ, АББАТИСА, АББАТСТВО'
res = [i if i != 'А' else '#' for i in input_str]
print(*res, sep='')
# ### Задача 4
#
# В математике существует так называемая последовательность чисел Фибоначчи. Выглядит она так: 1, 1, 2, 3, 5, 8, 13, ...
#
# Каждое последующее число равно сумме двух предыдущих, а первые два числа Фибоначчи - две единицы.
# Запросите с клавиатуры число N и запишите в список первые N чисел Фибоначчи.
# +
N = 15
res = [0, 1]
while len(res)<=N:
newpos = len(res)
res.append(res[newpos-1]+res[newpos-2])
print(res[1:])
# -
# ### Задача 5 Подсчет слов
# Давайте теперь поработаем с настоящим файлом и действительно посчитаем в нем слова. Мы загрузим метаданные почтового сервера университета Мичигана. И попробуем найти, с какого адреса ушло больше всего писем.
# импортируем библиотеку для доступа к файлам в интернете
import requests
# в переменной mbox хранится текст для работы
mbox = requests.get('http://www.py4inf.com/code/mbox.txt').text
# +
import re
counts = dict()
lines = mbox.split('\n')
for line in lines:
x = re.findall('From\s+([a-zA-Z0-9]\S+@\S+[a-zA-Z])', line)
for email in x:
counts[email] = counts.get(email, 0) + 1
highest = None
bigcount = None
for k, v in counts.items():
if highest is None or v > bigcount:
highest = k
bigcount = v
print(highest, counts[highest])
# -
# ### Задача 6 Посчитать среднюю метрику X-DSPAM-Confidence для письма.
# ### (Кажется, это что-то похожее на вероятность того, что письмо - не спам)
#
# Давайте воспользуемся предыдущим кодом и достанем те строчки, где эта метрика записана:
# импортируем библиотеку для доступа к файлам в интернете
import requests
# в переменной mbox хранится текст для работы
mbox = requests.get('http://www.py4inf.com/code/mbox.txt').text
spams = re.findall(f'X-DSPAM-Confidence:\s(\d\.*\d*)', mbox)
spam_num = [float(i) for i in spams]
print(sum(spam_num)/len(spam_num))
# ### Задача 7 Объединение словарей
#
# Напишите программу, которая объединяет значения из двух списков.
#
# Ввод:
#
# shops = [{'товар': 'яблоки', 'количество': 400}, {'товар': 'конфеты', 'количество': 300}, {'товар': 'яблоки', 'количество': 750}]
#
#
# Вывод:
#
# {'яблоки': 1150, 'конфеты': 300}
shops = [{'товар': 'яблоки', 'количество': 400},
{'товар': 'конфеты', 'количество': 300},
{'товар': 'яблоки', 'количество': 750}]
counts = dict()
for elem in shops:
counts[elem['товар']] = counts.get(elem['товар'], 0) + elem['количество']
print(counts)
# ### Задача 8 Найти все числа от 1000 до 3000 включительно, все цифры которых четные.
#
# Программа должна выдавать результат в виде разделенной запятыми строки
print(*[i for i in range(1000, 3000 + 1, 2)], sep=', ')
# ### Задача 9 Найти количество цифр и количество букв в строке
#
# Вход: "абвгд 234"
#
#
# Выход:
#
# Цифры: 3
# Буквы: 5
# %%timeit
str_input = 'абвгд 234'
letters = 'абвгдеёжзийклмнопрстуфхцчшщбъыэюя'
nums = '1234567890'
numcount = 0
lettercount = 0
for letter in str_input:
if letter in letters:
lettercount += 1
elif letter in nums:
numcount += 1
result = f'Цифры: {numcount}\nБуквы: {lettercount}'
# %%timeit
str_input = 'абвгд 234'
import re
nums = re.findall('\d', str_input)
letter = re.findall('[а-яА-яЁёйЙ]', str_input)
result = f'Цифры: {len(nums)}\nБуквы: {len(letter)}'
# значительно быстрее первый вариант
str_input = 'абвгд 234'
letters = 'абвгдеёжзийклмнопрстуфхцчшщбъыэюя'
nums = '1234567890'
numcount = 0
lettercount = 0
for letter in str_input:
if letter in letters:
lettercount += 1
elif letter in nums:
numcount += 1
result = f'Цифры: {numcount}\nБуквы: {lettercount}'
print(result)
# ### Задача 10 Найти частоту слов в строке. Строка вводится с клавиатуры, допустим, она разделена пробелами
#
# Вход: "Я не кидал никого никогда"
#
#
# Выход:
#
# Я:1
# не:1
# кидал:1
# никого:1
# никогда:1
str_input = "Я не кидал никого никогда"
words = str_input.split()
counts = {}
for word in words:
counts[word] = counts.get(word, 0) + 1
for k, v in counts.items():
print(k, v, sep = ':')
# ### Задача 11 Дан массив из чисел. Отсортируйте его, не используя встроенных функций. (циклами for, например). Ожидаю получить сортировку пузырьком)
#
# Вход: На вход дана последовательность из N случайных чисел. Например [48, 59, 9, 90, 15, 58, 19, 49, 13, 7]
#
#
# Выход:
#
# Отсортированная последовательность [7, 9, 13, 15, 19, 48, 49, 58, 59, 90]
# +
from random import randint
N = 10
a = []
for i in range(N):
a.append(randint(1, 99))
print(a)
# -
variants = [(i, j) for i in range(N)
for j in range(N)]
while len(variants) > 0:
d, f = variants.pop()
if a[d] > a[f]:
a[d], a[f] = a[f], a[d]
a
# ### Задача 12 Про <NAME>
#
# На плоскости в точке (0,0) стоит Поросёнок Пётр. Он умеет ходить налево, направо, вверх и вниз. Расстояние его прохода в какую-либо сторону измеряется в шагах. Когда он идет вправо, его первая координата увеличивается, когда влево - уменьшается. Когда он идет вверх, его вторая координата увеличивается, а когда вниз - уменьшается.
#
# С клавиатуры считывается число N - число ходов, которые сделает Пётр. После чего на каждом шаге спрашивается, сколько шагов и в какую сторону за этот ход Пётр сделает. Так происходит, пока Пётр не осуществит все N ходов.
#
# Программа должна вывести, сколько шагов Пётр должен был бы сделать, чтобы кратчайшим путем прибыть из свое начальной точки (0,0) в свою конечную точку. Напоминание: Пётр умеет ходить только вверх-вниз, и влево-вправо, но не по диагонали.
#
# Пример ввода:
#
# Введите N: 3
# Ход 1: Вверх 1
# Ход 2: Вниз 1
# Ход 3: Вверх 1
#
# Пример вывода:
#
# Пётр находится на расстоянии 1 от (0,0)
# +
N = int(input('Введите N: '))
steps = []
i_step = 0
while i_step < N:
step_input = input(f'Ход {i_step+1}: ')
step_input = step_input.split()
direction = step_input[0].lower()
dist = int(step_input[1])
steps.append((direction, dist))
i_step += 1
counts = {}
for elem in steps:
counts[elem[0]] = counts.get(elem[0], 0) + elem[1]
# посчитать сумму + и - движений для каждой оси. забить на другие элементы словаря
y_pos = abs(int(counts.get('вверх') or 0) - int(counts.get('вниз') or 0))
x_pos = abs(int(counts.get('вправо') or 0) - int(counts.get('влево') or 0))
# расчитаны расстояния
pos_new = (y_pos, x_pos)
print(f'Пётр находится на расстоянии {sum(pos_new)} от (0,0)')
| 01 python/lect 4 materials/2020_DPO_4_2_for_exercises_no_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problema Matriz
# Dado uma matriz MxN de números inteiros, busque um número na matriz toda e retorne a posição deste elemento. Retorne também os vizinhos deste elemento.
# +
import numpy as np
# matrix with m rows and n columns
def create_matrix(m, n, min_, max_):
return np.random.random_integers(min_, max_, (m, n))
# -
matrix = create_matrix(3, 4, 1, 23)
matrix
matrix[0][0]
matrix[1][2] # row, column
matrix.shape
def search_element(x, matrix):
rows, columns = matrix.shape
list_index_element = list()
for r in range(rows):
for c in range(columns):
if x == matrix[r][c]:
list_index_element.append((r, c))
return list_index_element
search_element(2, matrix)
search_element(8, matrix)
def return_neighbors(x, m, n, matrix):
list_index_elements = search_element(x, matrix)
list_neighbors = []
for i in list_index_elements:
# Upper
if i[0]-1 >= 0:
list_neighbors.append((i[0]-1, i[1]))
else:
list_neighbors.append(0)
# Down
if i[0]+1 <= m-1:
list_neighbors.append((i[0]+1, i[1]))
else:
list_neighbors.append(0)
# Right
if i[1]-1 >= 0:
list_neighbors.append((i[0], i[1]-1))
else:
list_neighbors.append(0)
# Left
if i[1]+1 <= n-1:
list_neighbors.append((i[0], i[1]+1))
else:
list_neighbors.append(0)
return list_index_elements, list_neighbors # upper, down, left, right
# +
x = 2
m = 3
n = 4
return_neighbors(x, m, n, matrix)
# +
x = 8
m = 3
n = 4
return_neighbors(x, m, n, matrix)
# +
x = 6
m = 3
n = 4
return_neighbors(x, m, n, matrix)
# +
x = 17
return_neighbors(x, m, n, matrix)
# -
list_index, list_neighbors = return_neighbors(12, m, n, matrix)
list_index
list_neighbors
def find_element_and_print(list_index, list_neighbors, matrix):
for i in list_index:
print("Position:", i)
counter = 0
for j in list_neighbors:
if j == 0:
counter += 1
else:
if counter == 0:
print("Upper:", matrix[j[0], j[1]])
if counter == 1:
print("Down:", matrix[j[0], j[1]])
if counter == 2:
print("Right:", matrix[j[0], j[1]])
if counter == 3:
print("Left:", matrix[j[0], j[1]])
counter = 0
find_element_and_print(list_index, list_neighbors, matrix)
# Crie um programa que lê um número X e retorna os vizinhos dele à esquerda, acima, à direita e abaixo de X, quando houver. Exemplo:
# 10 8 15 12\
# 21 11 23 8\
# 14 5 13 9
#
# Busca: 8
#
# Output:
#
# Position 0,1:\
# Left: 10\
# Right: 15\
# Down: 11
#
# Position 1,3:\
# Left: 23\
# Up: 12\
# Down: 19
| 100DOC/006.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_notebooks
# language: python
# name: conda_notebooks
# ---
# +
import sys
import numpy as np
item_count = 50000000
# +
# %%time
nums_list = [ 1 for _ in range(item_count) ]
double_nums_list = [ x * 2 for x in nums_list ]
double_nums_list_sum = sum(double_nums_list)
print(double_nums_list_sum)
print(sys.getsizeof(double_nums_list) / 1000000)
# +
# %%time
nums_arr = np.ones(item_count, dtype="int8")
double_nums_arr = nums_arr * 2
double_nums_arr_sum = np.sum(double_nums_arr)
print(double_nums_arr_sum)
print(double_nums_arr.nbytes / 1000000)
# +
# %%time
nums_arr = np.ones(item_count)
double_nums_arr = nums_arr * 2
double_nums_arr_sum = np.sum(double_nums_arr)
print(double_nums_arr_sum)
print(double_nums_arr.nbytes / 1000000)
# +
# %%time
nums_arr = np.ones(item_count, dtype="int64")
double_nums_arr = nums_arr * 2
double_nums_arr_sum = np.sum(double_nums_arr)
print(double_nums_arr_sum)
print(double_nums_arr.nbytes / 1000000)
# +
# %%time
nums_arr = np.ones(item_count, dtype="int32")
double_nums_arr = nums_arr * 2
double_nums_arr_sum = np.sum(double_nums_arr)
print(double_nums_arr_sum)
print(double_nums_arr.nbytes / 1000000)
# -
| conda_notebooks/01_Numpy_Performance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp data.question_answering
# +
#hide
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
import os
os.environ["TOKENIZERS_PARALLELISM"] = "false"
# -
# # data.question_answering
#
# > This module contains the bits required to use the fastai DataBlock API and/or mid-level data processing pipelines to organize your data for question/answering tasks.
# +
#export
import ast
from functools import reduce
from blurr.utils import *
from blurr.data.core import *
import torch
from transformers import *
from fastai.text.all import *
logging.set_verbosity_error()
# +
#hide
import pdb
from nbdev.showdoc import *
from fastcore.test import *
from fastai import __version__ as fa_version
from torch import __version__ as pt_version
from transformers import __version__ as hft_version
print(f'Using pytorch {pt_version}')
print(f'Using fastai {fa_version}')
print(f'Using transformers {hft_version}')
# -
#cuda
torch.cuda.set_device(1)
print(f'Using GPU #{torch.cuda.current_device()}: {torch.cuda.get_device_name()}')
# ## Question/Answering tokenization, batch transform, and DataBlock methods
#
# Question/Answering tasks are models that require two text inputs (a context that includes the answer and the question). The objective is to predict the start/end tokens of the answer in the context)
path = Path('./')
squad_df = pd.read_csv(path/'squad_sample.csv'); len(squad_df)
# We've provided a simple subset of a pre-processed SQUADv2 dataset below just for demonstration purposes. There is a lot that can be done to make this much better and more fully functional. The idea here is just to show you how things can work for tasks beyond sequence classification.
squad_df.head(2)
# +
model_cls = AutoModelForQuestionAnswering
pretrained_model_name = 'roberta-base' #'xlm-mlm-ende-1024'
hf_arch, hf_config, hf_tokenizer, hf_model = BLURR.get_hf_objects(pretrained_model_name, model_cls=model_cls)
# -
#export
def pre_process_squad(row, hf_arch, hf_tokenizer):
context, qst, ans = row['context'], row['question'], row['answer_text']
tok_kwargs = {}
if(hf_tokenizer.padding_side == 'right'):
tok_input = hf_tokenizer.convert_ids_to_tokens(hf_tokenizer.encode(qst, context, **tok_kwargs))
else:
tok_input = hf_tokenizer.convert_ids_to_tokens(hf_tokenizer.encode(context, qst, **tok_kwargs))
tok_ans = hf_tokenizer.tokenize(str(row['answer_text']), **tok_kwargs)
start_idx, end_idx = 0,0
for idx, tok in enumerate(tok_input):
try:
if (tok == tok_ans[0] and tok_input[idx:idx + len(tok_ans)] == tok_ans):
start_idx, end_idx = idx, idx + len(tok_ans)
break
except: pass
row['tokenized_input'] = tok_input
row['tokenized_input_len'] = len(tok_input)
row['tok_answer_start'] = start_idx
row['tok_answer_end'] = end_idx
return row
# The `pre_process_squad` method is structured around how we've setup the squad DataFrame above.
squad_df = squad_df.apply(partial(pre_process_squad, hf_arch=hf_arch, hf_tokenizer=hf_tokenizer), axis=1)
max_seq_len= 128
squad_df = squad_df[(squad_df.tok_answer_end < max_seq_len) & (squad_df.is_impossible == False)]
#hide
squad_df.head(2)
vocab = dict(enumerate(range(max_seq_len)))
#export
class HF_QuestionAnswerInput(HF_BaseInput): pass
# We'll return a `HF_QuestionAnswerInput` from our custom `HF_BeforeBatchTransform` so that we can customize the show_batch/results methods for this task.
#export
class HF_QABeforeBatchTransform(HF_BeforeBatchTransform):
def __init__(self, hf_arch, hf_config, hf_tokenizer, hf_model,
max_length=None, padding=True, truncation=True, is_split_into_words=False,
tok_kwargs={}, **kwargs):
super().__init__(hf_arch, hf_config, hf_tokenizer, hf_model,
max_length=max_length, padding=padding, truncation=truncation,
is_split_into_words=is_split_into_words, tok_kwargs=tok_kwargs, **kwargs)
def encodes(self, samples):
samples = super().encodes(samples)
for s in samples:
# cls_index: location of CLS token (used by xlnet and xlm); is a list.index(value) for pytorch tensor's
s[0]['cls_index'] = (s[0]['input_ids'] == self.hf_tokenizer.cls_token_id).nonzero()[0]
# p_mask: mask with 1 for token than cannot be in the answer, else 0 (used by xlnet and xlm)
s[0]['p_mask'] = s[0]['special_tokens_mask']
return samples
# By overriding `HF_BeforeBatchTransform` we can add other inputs to each example for this particular task.
# +
before_batch_tfm = HF_QABeforeBatchTransform(hf_arch, hf_config, hf_tokenizer, hf_model,
max_length=max_seq_len, truncation='only_second',
tok_kwargs={ 'return_special_tokens_mask': True })
blocks = (
HF_TextBlock(before_batch_tfm=before_batch_tfm, input_return_type=HF_QuestionAnswerInput),
CategoryBlock(vocab=vocab),
CategoryBlock(vocab=vocab)
)
dblock = DataBlock(blocks=blocks,
get_x=lambda x: (x.question, x.context),
get_y=[ColReader('tok_answer_start'), ColReader('tok_answer_end')],
splitter=RandomSplitter(),
n_inp=1)
# -
dls = dblock.dataloaders(squad_df, bs=4)
b = dls.one_batch(); len(b), len(b[0]), len(b[1]), len(b[2])
b[0]['input_ids'].shape, b[0]['attention_mask'].shape, b[1].shape, b[2].shape
#export
@typedispatch
def show_batch(x:HF_QuestionAnswerInput, y, samples, dataloaders, ctxs=None, max_n=6, trunc_at=None, **kwargs):
# grab our tokenizer
hf_before_batch_tfm = get_blurr_tfm(dataloaders.before_batch)
hf_tokenizer = hf_before_batch_tfm.hf_tokenizer
res = L()
for sample, input_ids, start, end in zip(samples, x, *y):
txt = hf_tokenizer.decode(sample[0], skip_special_tokens=True)[:trunc_at]
ans_toks = hf_tokenizer.convert_ids_to_tokens(input_ids, skip_special_tokens=False)[start:end]
res.append((txt, (start.item(),end.item()), hf_tokenizer.convert_tokens_to_string(ans_toks)))
display_df(pd.DataFrame(res, columns=['text', 'start/end', 'answer'])[:max_n])
return ctxs
# The `show_batch` method above allows us to create a more interpretable view of our question/answer data.
dls.show_batch(dataloaders=dls, max_n=2, trunc_at=500)
# ## Cleanup
#hide
from nbdev.export import notebook2script
notebook2script()
| nbs/04_data-question-answering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import warnings
import itertools
import pandas as pd
import numpy as np
import statsmodels.api as sm
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
# %matplotlib inline
data = sm.datasets.co2.load_pandas()
y = data.data
print(data.data)
# +
# The 'MS' string groups the data in buckets by start of the month
y = y['co2'].resample('MS').mean()
# The term bfill means that we use the value before filling in missing values
y = y.fillna(y.bfill())
print(y)
# -
y.plot(figsize=(15, 6))
plt.show()
# +
# Define the p, d and q parameters to take any value between 0 and 2
p = d = q = range(0, 2)
# Generate all different combinations of p, q and q triplets
pdq = list(itertools.product(p, d, q))
# Generate all different combinations of seasonal p, q and q triplets
seasonal_pdq = [(x[0], x[1], x[2], 12) for x in list(itertools.product(p, d, q))]
print('Examples of parameter combinations for Seasonal ARIMA...')
print('SARIMAX: {} x {}'.format(pdq[1], seasonal_pdq[1]))
print('SARIMAX: {} x {}'.format(pdq[1], seasonal_pdq[2]))
print('SARIMAX: {} x {}'.format(pdq[2], seasonal_pdq[3]))
print('SARIMAX: {} x {}'.format(pdq[2], seasonal_pdq[4]))
# +
warnings.filterwarnings("ignore") # specify to ignore warning messages
for param in pdq:
for param_seasonal in seasonal_pdq:
try:
mod = sm.tsa.statespace.SARIMAX(y,
order=param,
seasonal_order=param_seasonal,
enforce_stationarity=False,
enforce_invertibility=False)
results = mod.fit()
print('ARIMA{}x{}12 - AIC:{}'.format(param, param_seasonal, results.aic))
except:
continue
# +
mod = sm.tsa.statespace.SARIMAX(y,
order=(1, 1, 1),
seasonal_order=(1, 1, 1, 12),
enforce_stationarity=False,
enforce_invertibility=False)
results = mod.fit()
print(results.summary())
# -
results.plot_diagnostics(figsize=(15,12))
plt.show()
pred = results.get_prediction(start=pd.to_datetime('1998-01-01'), dynamic=False)
pred_ci = pred.conf_int()
# +
ax = y['1990':].plot(label='observed')
pred.predicted_mean.plot(ax=ax, label='One-step ahead Forecast', alpha=.7)
ax.fill_between(pred_ci.index,
pred_ci.iloc[:, 0],
pred_ci.iloc[:, 1], color='k', alpha=.2)
ax.set_xlabel('Date')
ax.set_ylabel('CO2 Levels')
plt.legend()
plt.show()
# -
# +
# Extract the predicted and true values of our time series
y_forecasted = pred.predicted_mean
y_truth = y['1998-01-01':]
# Compute the mean square error
mse = ((y_forecasted - y_truth) ** 2).mean()
print('The Mean Squared Error of our forecasts is {}'.format(round(mse, 2)))
# -
pred_dynamic = results.get_prediction(start=pd.to_datetime('1998-01-01'), dynamic=True, full_results=True)
pred_dynamic_ci = pred_dynamic.conf_int()
# +
ax = y['1990':].plot(label='observed', figsize=(20, 15))
pred_dynamic.predicted_mean.plot(label='Dynamic Forecast', ax=ax)
ax.fill_between(pred_dynamic_ci.index,
pred_dynamic_ci.iloc[:, 0],
pred_dynamic_ci.iloc[:, 1], color='k', alpha=.25)
ax.fill_betweenx(ax.get_ylim(), pd.to_datetime('1998-01-01'), y.index[-1],
alpha=.1, zorder=-1)
ax.set_xlabel('Date')
ax.set_ylabel('CO2 Levels')
plt.legend()
plt.show()
# +
# Extract the predicted and true values of our time series
y_forecasted = pred_dynamic.predicted_mean
y_truth = y['1998-01-01':]
# Compute the mean square error
mse = ((y_forecasted - y_truth) ** 2).mean()
print('The Mean Squared Error of our forecasts is {}'.format(round(mse, 2)))
# +
# Get forecast 500 steps ahead in future
pred_uc = results.get_forecast(steps=500)
# Get confidence intervals of forecasts
pred_ci = pred_uc.conf_int()
# +
ax = y.plot(label='observed', figsize=(20, 15))
pred_uc.predicted_mean.plot(ax=ax, label='Forecast')
ax.fill_between(pred_ci.index,
pred_ci.iloc[:, 0],
pred_ci.iloc[:, 1], color='k', alpha=.25)
ax.set_xlabel('Date')
ax.set_ylabel('CO2 Levels')
plt.legend()
plt.show()
# +
| .ipynb_checkpoints/ARIMA-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import statistics as stats
# +
lst = [12, 23, 43, 19, 1, 33, 76 , 100]
lst_range = max(lst) - min(lst)
lst_range
# -
lst = [12, 23, 43, 19, 1, 33, 76, 100, 12, 54, 56, 98, 32, 54, 87]
stats.variance(lst)
stats.stdev(lst)
# # Exercise
# 1.1
import statistics as stats
points = [-4, 17, 25, 19, 6, 11, 7]
points_range = max(points) - min(points)
points_range
# 1.2
data = 2, 4, 7, 1, 6, 8
data2 = 2, 4, 7, 1, 12, 8
stats.stdev(data)
stats.stdev(data2)
Difference = (stats.stdev(data2) - stats.stdev(data))
print("It increases", (Difference))
# 1.3
# yes
#
# any number that is the same or has the same 'mean' as every other element in the data set
data = [0,0,0,0,0,0]
stats.stdev(data)
# 1.4
# What is meant by deviation is the difference between one mumber and the next in each data set. "how far each number is away from one another"
# 1.5
import statistics as stats
# +
data = [23, 12, 34, 65, 34, 81]
for item in data:
print ("Deviation for item: ", item, "is: ", item - stats.mean(data))
# -
# 1.6
# Standard deviation is the square of the unit of the orginial data where as standard deviation is the same as the unit of the orginal data.
| Jupyter/Measures of Dispersion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center>
# <img src="../../img/ods_stickers.jpg">
# ## Open Machine Learning Course
# <center>Author: [<NAME>](https://www.linkedin.com/in/festline/) <br>
# Translated by [<NAME>](https://www.linkedin.com/in/sergeoreshkov/) <br>
# All content is distributed under the [Creative Commons CC BY-NC-SA 4.0](https://creativecommons.org/licenses/by-nc-sa/4.0/) license.
# # <center> Assignment #8 (demo)
#
# ## <center> Implementation of online regressor
# Here we'll implement a regressor trained with stochastic gradient descent (SGD). Fill in the missing code. If you do evething right, you'll pass a simple embedded test.
# ## <center>Linear regression and Stochastic Gradient Descent
import numpy as np
import pandas as pd
from tqdm import tqdm
from sklearn.base import BaseEstimator
from sklearn.metrics import mean_squared_error, log_loss, roc_auc_score
from sklearn.model_selection import train_test_split
# %matplotlib inline
from matplotlib import pyplot as plt
import seaborn as sns
from sklearn.preprocessing import StandardScaler
# Implement class `SGDRegressor`. Specification:
# - class is inherited from `sklearn.base.BaseEstimator`
# - constructor takes parameters `eta` – gradient step ($10^{-3}$ by default) and `n_epochs` – dataset pass count (3 by default)
# - constructor also createы `mse_` and `weights_` lists in order to track mean squared error and weight vector during gradient descent iterations
# - Class has `fit` and `predict` methods
# - The `fit` method takes matrix `X` and vector `y` (`numpy.array` objects) as parameters, appends column of ones to `X` on the left side, initializes weight vector `w` with **zeros** and then makes `n_epochs` iterations of weight updates (you may refer to this [article](https://medium.com/open-machine-learning-course/open-machine-learning-course-topic-8-vowpal-wabbit-fast-learning-with-gigabytes-of-data-60f750086237) for details), and for every iteration logs mean squared error and weight vector `w` in corresponding lists we created in the constructor.
# - Additionally the `fit` method will create `w_` variable to store weights which produce minimal mean squared error
# - The `fit` method returns current instance of the `SGDRegressor` class, i.e. `self`
# - The `predict` method takes `X` matrix, adds column of ones to the left side and returns prediction vector, using weight vector `w_`, created by the `fit` method.
class SGDRegressor(BaseEstimator):
# you code here
def __init__(self):
pass
def fit(self, X, y):
pass
def predict(self, X):
pass
# Let's test out the algorithm on height/weight data. We will predict heights (in inches) based on weights (in lbs).
data_demo = pd.read_csv('../../data/weights_heights.csv')
plt.scatter(data_demo['Weight'], data_demo['Height']);
plt.xlabel('Weight (lbs)')
plt.ylabel('Height (Inch)')
plt.grid();
X, y = data_demo['Weight'].values, data_demo['Height'].values
# Perform train/test split and scale data.
X_train, X_valid, y_train, y_valid = train_test_split(X, y,
test_size=0.3,
random_state=17)
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train.reshape([-1, 1]))
X_valid_scaled = scaler.transform(X_valid.reshape([-1, 1]))
# Train created `SGDRegressor` with `(X_train_scaled, y_train)` data. Leave default parameter values for now.
# +
# you code here
# -
# Draw a chart with training process – dependency of mean squared error from the i-th SGD iteration number.
# +
# you code here
# -
# Print the minimal value of mean squared error and the best weights vector.
# +
# you code here
# -
# Draw chart of model weights ($w_0$ and $w_1$) behavior during training.
# +
# you code here
# -
# Make a prediction for hold-out set `(X_valid_scaled, y_valid)` and check MSE value.
# you code here
sgd_holdout_mse = 10
# Do the same thing for `LinearRegression` class from `sklearn.linear_model`. Evaluate MSE for hold-out set.
# you code here
linreg_holdout_mse = 9
try:
assert (sgd_holdout_mse - linreg_holdout_mse) < 1e-4
print('Correct!')
except AssertionError:
print("Something's not good.\n Linreg's holdout MSE: {}"
"\n SGD's holdout MSE: {}".format(linreg_holdout_mse,
sgd_holdout_mse))
| assignments/demo/assignment08_implement_sgd_regressor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from biom.table import Table
data = np.arange(40).reshape(10, 4)
sample_ids = ['S%d' % i for i in range(4)]
observ_ids = ['O%d' % i for i in range(10)]
sample_metadata = [{'environment': 'A'}, {'environment': 'B'},
{'environment': 'A'}, {'environment': 'B'}]
observ_metadata = [{'taxonomy': ['Bacteria', 'Firmicutes']},
{'taxonomy': ['Bacteria', 'Firmicutes']},
{'taxonomy': ['Bacteria', 'Proteobacteria']},
{'taxonomy': ['Bacteria', 'Proteobacteria']},
{'taxonomy': ['Bacteria', 'Proteobacteria']},
{'taxonomy': ['Bacteria', 'Bacteroidetes']},
{'taxonomy': ['Bacteria', 'Bacteroidetes']},
{'taxonomy': ['Bacteria', 'Firmicutes']},
{'taxonomy': ['Bacteria', 'Firmicutes']},
{'taxonomy': ['Bacteria', 'Firmicutes']}]
table = Table(data, observ_ids, sample_ids, observ_metadata,
sample_metadata, table_id='Example Table')
data
| NOTEBOOKS/songbird.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # CDF and PDF plots of Standard Normal Distribution
# +
import os
try:
import jax
except:
# %pip install jax jaxlib
import jax
import jax.numpy as jnp
from jax.scipy.stats import norm
try:
import matplotlib.pyplot as plt
except:
# %pip install matplotlib
import matplotlib.pyplot as plt
try:
import seaborn as sns
except:
# %pip install seaborn
import seaborn as sns
# +
dev_mode = "DEV_MODE" in os.environ
if dev_mode:
import sys
sys.path.append("scripts")
import pyprobml_utils as pml
from latexify import latexify
latexify(width_scale_factor=2, fig_height=1.5)
# +
# Plot pdf and cdf of standard normal
x = jnp.linspace(-3, 3, 500)
random_var = norm
def make_graph(data, save_name):
fig, ax = plt.subplots()
ax.plot(data["x"], data["y"])
# plt.title("Gaussian pdf")
plt.xlabel("x")
plt.ylabel(data["ylabel"])
sns.despine()
if dev_mode:
pml.save_fig(save_name)
make_graph({"x": x, "y": random_var.pdf(x), "ylabel": "$p(x)$"}, "gaussian1d_latexified.pdf")
make_graph(
{"x": x, "y": random_var.cdf(x), "ylabel": "$Pr(X \leq x)$"},
"gaussianCdf_latexified.pdf",
)
| notebooks/gauss_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="i1Beedm1sgf5"
# # PyTorch Tensor Operations
# + [markdown] colab_type="text" id="HcE5VXEQsgf6"
# This section covers:
# * Indexing and slicing
# * Reshaping tensors (tensor views)
# * Tensor arithmetic and basic operations
# * Dot products
# * Matrix multiplication
# * Additional, more advanced operations
#
# ## Perform standard imports
# + colab={} colab_type="code" id="PGyZhfousgf7"
import torch
import numpy as np
# + [markdown] colab_type="text" id="u5jLdQXxsgf_"
# ## Indexing and slicing
# Extracting specific values from a tensor works just the same as with NumPy arrays<br>
# <img src='https://github.com/shanaka-desoysa/pytorch-deep-learning/blob/master/Images/arrayslicing.png?raw=1' width="500" style="display: inline-block"><br><br>
# Image source: http://www.scipy-lectures.org/_images/numpy_indexing.png
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="RSaAp-XKsgf_" outputId="d5c18bf9-ebf6-4004-ff6f-746bc34068f5"
x = torch.arange(6).reshape(3,2)
print(x)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="QDQyWB5QsggE" outputId="fe17c2a3-3bc0-4316-a5c7-31f3438b0905"
# Grabbing the right hand column values
x[:,1]
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="wS04EViosggH" outputId="0da5c4fb-fe47-4fd3-bb5f-56daf53e7693"
# Grabbing the right hand column as a (3,1) slice
x[:,1:]
# + [markdown] colab_type="text" id="O55YPXiWsggK"
# ## Reshape tensors with <tt>.view()</tt>
# <a href='https://pytorch.org/docs/master/tensors.html#torch.Tensor.view'><strong><tt>view()</tt></strong></a> and <a href='https://pytorch.org/docs/master/torch.html#torch.reshape'><strong><tt>reshape()</tt></strong></a> do essentially the same thing by returning a reshaped tensor without changing the original tensor in place.<br>
# There's a good discussion of the differences <a href='https://stackoverflow.com/questions/49643225/whats-the-difference-between-reshape-and-view-in-pytorch'>here</a>.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="isNkRWBesggK" outputId="3503b087-1965-426c-f6ff-2b4dc19dab72"
x = torch.arange(12)
print(x)
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="SQ9eQePGsggN" outputId="9ea61d78-f284-41a7-bf90-c31542fb3788"
x.view(2,6)
# + colab={"base_uri": "https://localhost:8080/", "height": 119} colab_type="code" id="YGdYsC24sggQ" outputId="c94317b7-1e53-4f7a-b8c6-6282e01058c2"
x.view(6,2)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="AZM0zMaDsggT" outputId="3cfb0553-a725-4a44-8a4d-f4fe84198882"
# x is unchanged
x
# + [markdown] colab_type="text" id="GYOGSgMMsggW"
# ### Views reflect the most current data
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="Hue3Z1d6sggW" outputId="0d1d7555-4762-4e0c-d06f-ed3b043936c7"
z = x.view(2,6)
x[0]=234
print(z)
# + [markdown] colab_type="text" id="CVpG9ShasggZ"
# ### Views can infer the correct size
# By passing in <tt>-1</tt> PyTorch will infer the correct value from the given tensor
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="7NHOw0RCsggZ" outputId="8ae90714-99da-4e6d-f6b2-b863448fad90"
# infer number of columns for given rows
x.view(2,-1)
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="7qxe21nbsggc" outputId="b1cc2113-1ff1-4b2e-f11c-8e9db4a20b70"
# infer number of rows for given columns
x.view(-1,3)
# + [markdown] colab_type="text" id="M7E8Q2Xvsggf"
# ### Adopt another tensor's shape with <tt>.view_as()</tt>
# <a href='https://pytorch.org/docs/master/tensors.html#torch.Tensor.view_as'><strong><tt>view_as(input)</tt></strong></a> only works with tensors that have the same number of elements.
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="mytzLfbmsggf" outputId="d5c3b546-0188-4985-f628-771db2dc0ac4"
x.view_as(z)
# + [markdown] colab_type="text" id="qUxnglnAsggi"
# ## Tensor Arithmetic
# Adding tensors can be performed a few different ways depending on the desired result.<br>
#
# As a simple expression:
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="Nvszuxpbsggj" outputId="6988d9ed-fe80-44c0-e303-939ff553555d"
a = torch.tensor([1,2,3], dtype=torch.float)
b = torch.tensor([4,5,6], dtype=torch.float)
print(a + b)
# + [markdown] colab_type="text" id="hbekEY3Osggl"
# As arguments passed into a torch operation:
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="w5ndFuYRsggm" outputId="e16c093c-dc38-4bc9-fdb6-1ae43d6fcfd6"
print(torch.add(a, b))
# + [markdown] colab_type="text" id="QeJRxC-jsggo"
# With an output tensor passed in as an argument:
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="YORUzt3csggp" outputId="a0a4fafd-dcd5-41d4-f3d9-637960b1e682"
result = torch.empty(3)
torch.add(a, b, out=result) # equivalent to result=torch.add(a,b)
print(result)
# + [markdown] colab_type="text" id="1kxPdCrbsggr"
# **Changing a tensor in-place** with *_*
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="Kr6O6SNbsggs" outputId="c8a58b47-a119-4664-f1b5-9a7e8d64175f"
a.add_(b) # equivalent to a=torch.add(a,b)
print(a)
# + [markdown] colab_type="text" id="YdeCf7XCsggu"
# <div class="alert alert-info"><strong>NOTE:</strong> Any operation that changes a tensor in-place is post-fixed with an underscore _.
# <br>In the above example: <tt>a.add_(b)</tt> changed <tt>a</tt>.</div>
# + [markdown] colab_type="text" id="bNEv8zn-sggu"
# ### Basic Tensor Operations
# <table style="display: inline-block">
# <caption style="text-align: center"><strong>Arithmetic</strong></caption>
# <tr><th>OPERATION</th><th>FUNCTION</th><th>DESCRIPTION</th></tr>
# <tr><td>a + b</td><td>a.add(b)</td><td>element wise addition</td></tr>
# <tr><td>a - b</td><td>a.sub(b)</td><td>subtraction</td></tr>
# <tr><td>a * b</td><td>a.mul(b)</td><td>multiplication</td></tr>
# <tr><td>a / b</td><td>a.div(b)</td><td>division</td></tr>
# <tr><td>a % b</td><td>a.fmod(b)</td><td>modulo (remainder after division)</td></tr>
# <tr><td>a<sup>b</sup></td><td>a.pow(b)</td><td>power</td></tr>
# <tr><td> </td><td></td><td></td></tr>
# </table>
# + [markdown] colab_type="text" id="e-Pxrt6Esggv"
# <table style="display: inline-block">
# <caption style="text-align: center"><strong>Monomial Operations</strong></caption>
# <tr><th>OPERATION</th><th>FUNCTION</th><th>DESCRIPTION</th></tr>
# <tr><td>|a|</td><td>torch.abs(a)</td><td>absolute value</td></tr>
# <tr><td>1/a</td><td>torch.reciprocal(a)</td><td>reciprocal</td></tr>
# <tr><td>$\sqrt{a}$</td><td>torch.sqrt(a)</td><td>square root</td></tr>
# <tr><td>log(a)</td><td>torch.log(a)</td><td>natural log</td></tr>
# <tr><td>e<sup>a</sup></td><td>torch.exp(a)</td><td>exponential</td></tr>
# <tr><td>12.34 ==> 12.</td><td>torch.trunc(a)</td><td>truncated integer</td></tr>
# <tr><td>12.34 ==> 0.34</td><td>torch.frac(a)</td><td>fractional component</td></tr>
# </table>
# + [markdown] colab_type="text" id="DZj8ANYRsggv"
# <table style="display: inline-block">
# <caption style="text-align: center"><strong>Trigonometry</strong></caption>
# <tr><th>OPERATION</th><th>FUNCTION</th><th>DESCRIPTION</th></tr>
# <tr><td>sin(a)</td><td>torch.sin(a)</td><td>sine</td></tr>
# <tr><td>cos(a)</td><td>torch.sin(a)</td><td>cosine</td></tr>
# <tr><td>tan(a)</td><td>torch.sin(a)</td><td>tangent</td></tr>
# <tr><td>arcsin(a)</td><td>torch.asin(a)</td><td>arc sine</td></tr>
# <tr><td>arccos(a)</td><td>torch.acos(a)</td><td>arc cosine</td></tr>
# <tr><td>arctan(a)</td><td>torch.atan(a)</td><td>arc tangent</td></tr>
# <tr><td>sinh(a)</td><td>torch.sinh(a)</td><td>hyperbolic sine</td></tr>
# <tr><td>cosh(a)</td><td>torch.cosh(a)</td><td>hyperbolic cosine</td></tr>
# <tr><td>tanh(a)</td><td>torch.tanh(a)</td><td>hyperbolic tangent</td></tr>
# </table>
# + [markdown] colab_type="text" id="Q__SLr4Lsggv"
# <table style="display: inline-block">
# <caption style="text-align: center"><strong>Summary Statistics</strong></caption>
# <tr><th>OPERATION</th><th>FUNCTION</th><th>DESCRIPTION</th></tr>
# <tr><td>$\sum a$</td><td>torch.sum(a)</td><td>sum</td></tr>
# <tr><td>$\bar a$</td><td>torch.mean(a)</td><td>mean</td></tr>
# <tr><td>a<sub>max</sub></td><td>torch.max(a)</td><td>maximum</td></tr>
# <tr><td>a<sub>min</sub></td><td>torch.min(a)</td><td>minimum</td></tr>
# <tr><td colspan="3">torch.max(a,b) returns a tensor of size a<br>containing the element wise max between a and b</td></tr>
# </table>
# + [markdown] colab_type="text" id="uE5r-GtCsggw"
# <div class="alert alert-info"><strong>NOTE:</strong> Most arithmetic operations require float values. Those that do work with integers return integer tensors.<br>
# For example, <tt>torch.div(a,b)</tt> performs floor division (truncates the decimal) for integer types, and classic division for floats.</div>
# + [markdown] colab_type="text" id="F6kD3S8ssggw"
# #### Use the space below to experiment with different operations
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="HM_Whl8Ksggx" outputId="9e850e02-e0d1-43e2-fdab-621fb30ce907"
a = torch.tensor([1,2,3], dtype=torch.float)
b = torch.tensor([4,5,6], dtype=torch.float)
print(torch.add(a,b).sum())
# + [markdown] colab_type="text" id="oY8zYWgdsgg4"
# ## Dot products
# A <a href='https://en.wikipedia.org/wiki/Dot_product'>dot product</a> is the sum of the products of the corresponding entries of two 1D tensors. If the tensors are both vectors, the dot product is given as:<br>
#
# $\begin{bmatrix} a & b & c \end{bmatrix} \;\cdot\; \begin{bmatrix} d & e & f \end{bmatrix} = ad + be + cf$
#
# If the tensors include a column vector, then the dot product is the sum of the result of the multiplied matrices. For example:<br>
# $\begin{bmatrix} a & b & c \end{bmatrix} \;\cdot\; \begin{bmatrix} d \\ e \\ f \end{bmatrix} = ad + be + cf$<br><br>
# Dot products can be expressed as <a href='https://pytorch.org/docs/stable/torch.html#torch.dot'><strong><tt>torch.dot(a,b)</tt></strong></a> or `a.dot(b)` or `b.dot(a)`
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="G3_VAsSIsgg4" outputId="248eb9b4-35d2-4836-9aa0-22735ce0435e"
a = torch.tensor([1,2,3], dtype=torch.float)
b = torch.tensor([4,5,6], dtype=torch.float)
print(a.mul(b)) # for reference
print()
print(a.dot(b))
# + [markdown] colab_type="text" id="bo1eKBR6sgg6"
# <div class="alert alert-info"><strong>NOTE:</strong> There's a slight difference between <tt>torch.dot()</tt> and <tt>numpy.dot()</tt>. While <tt>torch.dot()</tt> only accepts 1D arguments and returns a dot product, <tt>numpy.dot()</tt> also accepts 2D arguments and performs matrix multiplication. We show matrix multiplication below.</div>
# + [markdown] colab_type="text" id="0GK-l8fcsgg7"
# ## Matrix multiplication
# 2D <a href='https://en.wikipedia.org/wiki/Matrix_multiplication'>Matrix multiplication</a> is possible when the number of columns in tensor <strong><tt>A</tt></strong> matches the number of rows in tensor <strong><tt>B</tt></strong>. In this case, the product of tensor <strong><tt>A</tt></strong> with size $(x,y)$ and tensor <strong><tt>B</tt></strong> with size $(y,z)$ results in a tensor of size $(x,z)$
# <div>
# <div align="left"><img src='https://github.com/shanaka-desoysa/pytorch-deep-learning/blob/master/Images/Matrix_multiplication_diagram.png?raw=1' align="left"><br><br>
#
# $\begin{bmatrix} a & b & c \\
# d & e & f \end{bmatrix} \;\times\; \begin{bmatrix} m & n \\ p & q \\ r & s \end{bmatrix} = \begin{bmatrix} (am+bp+cr) & (an+bq+cs) \\
# (dm+ep+fr) & (dn+eq+fs) \end{bmatrix}$</div></div>
#
# <div style="clear:both">Image source: <a href='https://commons.wikimedia.org/wiki/File:Matrix_multiplication_diagram_2.svg'>https://commons.wikimedia.org/wiki/File:Matrix_multiplication_diagram_2.svg</a></div>
#
# Matrix multiplication can be computed using <a href='https://pytorch.org/docs/stable/torch.html#torch.mm'><strong><tt>torch.mm(a,b)</tt></strong></a> or `a.mm(b)` or `a @ b`
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="mgqreKVMsgg7" outputId="b05c5fd5-5da6-4999-95cb-aecd6d67d519"
a = torch.tensor([[0,2,4],[1,3,5]], dtype=torch.float)
b = torch.tensor([[6,7],[8,9],[10,11]], dtype=torch.float)
print('a: ',a.size())
print('b: ',b.size())
print('a x b: ',torch.mm(a,b).size())
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="PuEDdmqNsgg9" outputId="49ca2c20-45de-464e-806f-3624b3c4db61"
print(torch.mm(a,b))
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="wUGUvlJ_sghA" outputId="2dc0b655-9be2-4868-e214-dfa8f4beed93"
print(a.mm(b))
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="wYnjzhgpsghD" outputId="223fff5c-4009-4388-cb42-4b7042d11d24"
print(a @ b)
# + [markdown] colab_type="text" id="tE0lWX4LsghF"
# ### Matrix multiplication with broadcasting
# Matrix multiplication that involves <a href='https://pytorch.org/docs/stable/notes/broadcasting.html#broadcasting-semantics'>broadcasting</a> can be computed using <a href='https://pytorch.org/docs/stable/torch.html#torch.matmul'><strong><tt>torch.matmul(a,b)</tt></strong></a> or `a.matmul(b)` or `a @ b`
# + colab={} colab_type="code" id="6hZPUNwmsghF"
t1 = torch.randn(2, 3, 4)
t2 = torch.randn(4, 5)
# + colab={"base_uri": "https://localhost:8080/", "height": 136} colab_type="code" id="V1uVYBrrzFUo" outputId="53e4f0f5-8e47-4155-e383-aecb304c0825"
t1
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="SbDboLLNzGbJ" outputId="aa6dce26-f28a-424e-d039-93f54ff8be30"
t2
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="SP4WpsfDzKas" outputId="07db389a-84b2-498a-ad5c-aa45d99a9bd2"
print(torch.matmul(t1, t2).size())
# + [markdown] colab_type="text" id="MAiZRXVmsghH"
# However, the same operation raises a <tt><strong>RuntimeError</strong></tt> with <tt>torch.mm()</tt>:
# + colab={"base_uri": "https://localhost:8080/", "height": 164} colab_type="code" id="B2BFSR2usghI" outputId="19edcada-3494-417b-bc34-67fc4f887882"
print(torch.mm(t1, t2).size())
# + [markdown] colab_type="text" id="BUr9dNHfsghK"
# ___
# # Advanced operations
# + [markdown] colab_type="text" id="4XKUx6m_sghL"
# ## L2 or Euclidian Norm
# See <a href='https://pytorch.org/docs/stable/torch.html#torch.norm'><strong><tt>torch.norm()</tt></strong></a>
#
# The <a href='https://en.wikipedia.org/wiki/Norm_(mathematics)#Euclidean_norm'>Euclidian Norm</a> gives the vector norm of $x$ where $x=(x_1,x_2,...,x_n)$.<br>
# It is calculated as<br>
#
# ${\displaystyle \left\|{\boldsymbol {x}}\right\|_{2}:={\sqrt {x_{1}^{2}+\cdots +x_{n}^{2}}}}$
#
#
# When applied to a matrix, <tt>torch.norm()</tt> returns the <a href='https://en.wikipedia.org/wiki/Matrix_norm#Frobenius_norm'>Frobenius norm</a> by default.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="qxjzWwT0sghL" outputId="ccf2d203-70b0-45ba-b6d8-37237c322a06"
x = torch.tensor([2.,5.,8.,14.])
x.norm()
# + [markdown] colab_type="text" id="IeTxj_IEsghN"
# ## Number of elements
# See <a href='https://pytorch.org/docs/stable/torch.html#torch.numel'><strong><tt>torch.numel()</tt></strong></a>
#
# Returns the number of elements in a tensor.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="jEdsGRCusghO" outputId="825391bf-5ddd-40fb-9578-a8743b240328"
x = torch.ones(3,7)
x.numel()
# + [markdown] colab_type="text" id="rIyW4PVCsghP"
# This can be useful in certain calculations like Mean Squared Error:<br>
# <tt>
# def mse(t1, t2):<br>
# diff = t1 - t2<br>
# return torch.sum(diff * diff) / diff<strong>.numel()</strong></tt>
# -
# <a href="https://colab.research.google.com/github/shanaka-desoysa/notes/blob/master/content/deep_learning/pytorch/tensor_operations.ipynb" target="_blank"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
| docs/PyTorch/tensor_operations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Iris dataset and pandas
#
# 
#
#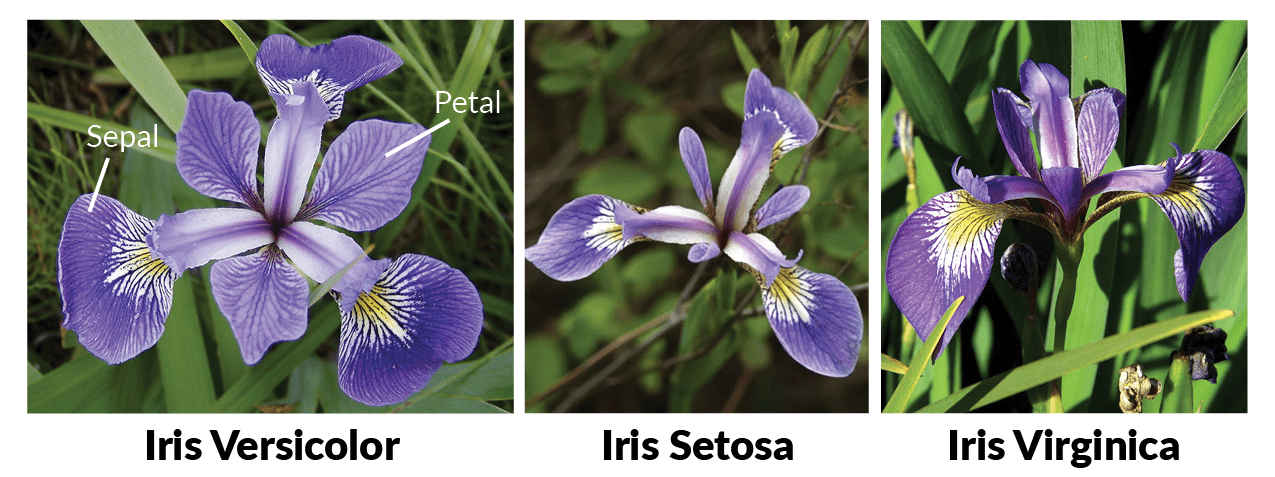 
#
# ***
#
# **[Python Data Analysis Library](https://pandas.pydata.org/)**
#
# *[https://pandas.pydata.org/](https://pandas.pydata.org/)*
#
# The pandas website.
#
# ***
#
# **[<NAME> - 10-minute tour of pandas](https://vimeo.com/59324550)**
#
# *[https://vimeo.com/59324550](https://vimeo.com/59324550)*
#
# A 10 minutes video introduction to pandas.
#
# ***
#
# **[Python for Data Analysis notebooks](https://github.com/wesm/pydata-book)**
#
# *[https://github.com/wesm/pydata-book](https://github.com/wesm/pydata-book)*
#
# Materials and IPython notebooks for "Python for Data Analysis" by <NAME>, published by O'Reilly Media
#
# ***
#
# **[10 Minutes to pandas](http://pandas.pydata.org/pandas-docs/stable/10min.html)**
#
# *[http://pandas.pydata.org/pandas-docs/stable/10min.html](http://pandas.pydata.org/pandas-docs/stable/10min.html)*
#
# Official pandas tutorial.
#
# ***
#
# **[UC Irvine Machine Learning Repository: Iris Data Set](https://archive.ics.uci.edu/ml/datasets/iris)**
#
# *[https://archive.ics.uci.edu/ml/datasets/iris](https://archive.ics.uci.edu/ml/datasets/iris)*
#
# About the Iris data set from UC Irvine's machine learning repository.
# ## Loading Data
# Import Pandas
import pandas as pd
# Load the iris data set from the website
df = pd.read_csv("https://raw.githubusercontent.com/uiuc-cse/data-fa14/gh-pages/data/iris.csv")
df
# ## Selecting Data (the dirty way)
# return a column based on header label
df["species"]
# return 2 columns based on thier labels
df[["petal_width", "species"]]
# return rows from 2 to 6 (exlusive of 6)
df[2:6]
# select a sub set of data based on rows and columns
df[["petal_width", "species"]][2:6]
# This is messy and can cause problems as sometimes is creates a new dataframe for the subset and other times if shoes the same one. better to use iloc[] for returns based on index, and loc[] for returns based on label
df.loc[2:6]
# The 6th row is returned becase for loc[] it returns based on the label
df.iloc[2:6]
# 6th row not returned as iloc[] is based on index
# return a full column
df.loc[:, "species"]
# return specified rows from a column
df.loc[2:6, "species"]
#return an indexed row
df.iloc[2]
# return a indexed range (with an indexed column)
df.iloc[2:4,1]
# at[] is used to return a particular cell
df.at[3,"species"]
# return indexed values from 1 to 10,but just every second value
df.iloc[1:10:2]
# ## Boolean Selects
# use a boolean check to see if the species matches an entered value
df.loc[:,"species"] == 'setosa'
# we can also use this test to return values that suit
df.loc[df.loc[:, "species"] == 'setosa']
# define a new dataframe that is a sub data frame
x = df.loc[df.loc[:, "species"] == 'versicolor']
# loc[] uses the row label
x.loc[51]
# iloc[] uses the row index
x.iloc[1]
# ## Summary Statistics
# show the top 5 items in the list
df.head()
# show the bottom 5 items in the list
df.tail()
# print the summary stats
df.describe()
# get the mean of a column
df.loc[:,"sepal_length"].mean()
# get the mean of a data frame
df.mean()
# ## Plots with Seaborn
# import seaborn
import seaborn as sns
sns.pairplot(df, hue="species")
| Week 05/pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Analytics Project - Backtesting Model Predictions
#
# # HEROMOTOCO STOCK MODELS EVALUATION
#
# ---
# # 1. Importing Required Modules
import pandas as pd
# # 2. Get required datasets
# ## 2.1. Get the orders dataset of HEROMOTOCO
orders_df = pd.read_csv('../Preprocess/Pair1/Pair1_orders.csv')
orders_df.head()
orders_df.tail()
# ## 2.2. Visualize the orders
# +
# Plotting the zscore of the Spread
orders_plt = orders_df.plot(x='Date', y='zscore', figsize=(30,15))
# Plotting the lines at mean, 1 and 2 std. dev.
orders_plt.axhline(0, c='black')
orders_plt.axhline(1, c='red', ls = "--")
orders_plt.axhline(-1, c='red', ls = "--")
# Extracting orders
Orders = orders_df['Orders']
# Plot vertical lines where orders are placed
for order in range(len(Orders)):
if Orders[order] != "FLAT":
# GREEN line for a long position
if Orders[order] == "LONG":
orders_plt.axvline(x=order, c = "green")
# RED line for a short position
elif Orders[order] == "SHORT":
orders_plt.axvline(x=order, c = "red")
# BLACK line for getting out of all positions at that point
else:
orders_plt.axvline(x=order, c = "black")
orders_plt.set_ylabel("zscore")
# -
# __In the figure above:__
# - __Blue line__ - zscore of the Spread
# - __Black horizontal line__ at 0 - Mean
# - __Red dotted horizontal lines__ - at +1 and -1 standard deviations
# - __Green vertical line__ - represents long position taken on that day
# - __Red vertical line__ - represents short position taken on that day
# - __Black vertical line__ - represents getting out of all open positions till that point
#
# ## 2.3. Get the predicitions dataset of HEROMOTOCO
predictions = pd.read_csv("../Models/Pair1/Hero_predicitions.csv")
predictions.head()
predictions = predictions.dropna()
predictions.head()
# ## 2.4 Visualize the predictions by different models
predictions.plot(x='Date', y=['Actual_Close', 'Linear_regression_Close', 'ARIMA_Close', 'LSTM_Close'], figsize=(30,15))
# __In the figure above:__
# - __x-axis__ - Date
# - __y-axis__ - Price in rupees
# - __Blue line__ - Actual data in the time period of correlation
# - __Orange line__ - Linear Regression predictions for the time period
# - __Red line__ - LSTM predictions for the time period
# - __Green line__ - ARIMA predictions for the time period
#
# ---
# # 3. Set parameters of trading
CAPITAL = 1000000
RISK = 20000 # Max risk allowed per trade (2% of capital)
# - CAPITAL - Total amout of money willing to be spent.
# - RISK - Amount to spend per trade (maximum).
# - OPEN POSITION - Buy/Short shares of stock worth 20000.
# - CLOSE POSITION - Buy/Sell shares to consolidate open positions and take profit/loss.
# - TRADE_BOOK - Keeps track of all open positions.
# - For the purpose of this experiment, we assume no brokerage costs, which is a fair assumption as many brokers (such as zerodha, robinhood, sharekhan, etc.) offer equity delivery at no cost.
# ---
# # 4. Evaluate all individual orders
# ## 4.1. Display all orders which are not *FLAT*
not_flat_orders = orders_df[orders_df['Orders'] != 'FLAT']
not_flat_orders
not_flat_orders = not_flat_orders.set_index('Date').reset_index()
not_flat_orders
# ## 4.2 Flip all orders
def flip_orders(orders):
flipped_orders = []
for order in orders:
if order == 'SHORT':
flipped_orders.append('LONG')
elif order == 'LONG':
flipped_orders.append('SHORT')
else:
flipped_orders.append(order)
return flipped_orders
# not_flat_orders['Orders'] = flip_orders(not_flat_orders['Orders'])
not_flat_orders
# ## 4.3. Get predictions of dates where orders are placed
# ### 4.3.1. Filter rows in predictions to only include dates where orders are placed
not_flat_orders = not_flat_orders.set_index('Date')
not_flat_orders
predicitions = predictions.set_index('Date')
predicitions
orders_predictions = predicitions.join(not_flat_orders)
orders_predictions = orders_predictions.dropna()
orders_predictions
# ### 4.3.2 Remove columns in orders and predictions dataframe
orders_predictions = orders_predictions.drop(columns=['BAJAJ-AUTO_Close', 'HEROMOTOCO_Close', 'Spread', 'zscore'])
orders_predictions
# ### 4.3.3 Create function to evaluate orders
def evaluate_orders(orders_df):
actual_profits = []
LR_profits = []
ARIMA_profits = []
LSTM_profits = []
shares = []
num_orders = len(orders_df["Orders"])
for i in range(num_orders):
if i == num_orders - 1:
actual_profits.append(0)
LR_profits.append(0)
ARIMA_profits.append(0)
LSTM_profits.append(0)
shares.append(0)
break
position = orders_df['Orders'][i]
close = orders_df['Actual_Close'][i]
LR_close = orders_df['Linear_regression_Close'][i]
ARIMA_close = orders_df['ARIMA_Close'][i]
LSTM_close = orders_df['LSTM_Close'][i]
print()
print("ORDER: ", i, close, LR_close, ARIMA_close, LSTM_close, position)
if position == 'LONG' or position == 'SHORT':
no_of_shares = 20000//close
for j in range(i+1, num_orders):
if (j == num_orders - 1) or orders_df['Orders'][j] == 'GET_OUT_OF_POSITION':
actual_profit = orders_df['Actual_Close'][j] - orders_df['Actual_Close'][i]
LR_profit = orders_df['Linear_regression_Close'][j] - orders_df['Linear_regression_Close'][i]
ARIMA_profit = orders_df['ARIMA_Close'][j] - orders_df['ARIMA_Close'][i]
LSTM_profit = orders_df['LSTM_Close'][j] - orders_df['LSTM_Close'][i]
actual_profit *= no_of_shares
LR_profit *= no_of_shares
ARIMA_profit *= no_of_shares
LSTM_profit *= no_of_shares
if position == 'SHORT':
actual_profit *= -1
LR_profit *= -1
ARIMA_profit *= -1
LSTM_profit *= -1
print('number of shares: ', no_of_shares)
print('actual profit: ', actual_profit)
print('LR profit: ', LR_profit)
print('ARIMA profit: ', ARIMA_profit)
print('LSTM profit: ', LSTM_profit)
shares.append(no_of_shares)
actual_profits.append(actual_profit)
LR_profits.append(LR_profit)
ARIMA_profits.append(ARIMA_profit)
LSTM_profits.append(LSTM_profit)
break
else:
actual_profits.append(0)
LR_profits.append(0)
ARIMA_profits.append(0)
LSTM_profits.append(0)
shares.append(0)
print()
return actual_profits, LR_profits, ARIMA_profits, LSTM_profits, shares
actual_profits, LR_profits, ARIMA_profits, LSTM_profits, shares = evaluate_orders(orders_predictions)
actual_profits
LR_profits
LSTM_profits
shares
ARIMA_profits
# ### 4.3.4 Adding evaluated orders to dataframe
orders_predictions
orders_predictions['Shares'] = shares
orders_predictions['Actual_profit'] = actual_profits
orders_predictions['Linear_regression_profit'] = LR_profits
orders_predictions['ARIMA_profit'] = ARIMA_profits
orders_predictions['LSTM_profit'] = LSTM_profits
orders_predictions
# ### 4.3.5 Visualizing the profits
orders_predictions = orders_predictions.reset_index()
orders_predictions
orders_predictions.plot(x='Date', y=['Actual_profit', 'Linear_regression_profit', 'ARIMA_profit', 'LSTM_profit'], figsize=(30,15))
# __In the figure above:__
# - __x-axis__ - Date
# - __y-axis__ - Profit
# - __Blue line__ - Actual profit
# - __Orange line__ - Linear Regression predicted profit
# - __Green line__ - ARIMA predicted profit
# - __Red line__ - LSTM predicted profit
# ---
| Backtester/HEROMOTOCO_BACKTESTING.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Axion-neutron coupling limits vs axion mass
#
# Axion mass-PQ scale relation:
# \begin{equation}
# m_{a}=5.70(7) \mu \mathrm{eV}\left(\frac{10^{12} \,\mathrm{GeV}}{f_{a}}\right)
# \end{equation}
#
# Axion-proton coupling
# \begin{equation}
# g_{an} \equiv \frac{C_{ap} m_{p}}{f_{a}} = 1.64\times 10^{-7} C_{ap}\frac{m_a}{\mathrm{eV}}
# \end{equation}
#
# Model dependent constant:
# \begin{equation}
# C_{an} =
# \begin{cases}
# -0.47 & {\rm KSVZ} \\
# [-0.2,-0.6] & {\rm DFSZ\,I} \\
# [-0.2,-0.6] & {\rm DFSZ\,II}
# \end{cases}
# \end{equation}
# Like the axion-electron coupling the range of values for DFSZ I and II come from the perturbativity of the Yukawa couplings with sets the range $0.28<v_{u} / v_{d}<140$ for the Higgs vevs.
# +
% matplotlib inline
from PlotFuncs import BlackHoleSpins, AxionProton, MySaveFig, FigSetup
import matplotlib.pyplot as plt
fig, ax = FigSetup(Shape='Rectangular', ylab='$|g_{ap}|$', mathpazo=True,
g_min=1e-17, g_max=1e-2, m_min=1e-22, m_max=1e-2, FrequencyAxis=True, N_Hz=1)
AxionProton.QCDAxion(ax)
AxionProton.Haloscopes(ax, projection=False)
AxionProton.StellarBounds(ax)
AxionProton.LabExperiments(ax, projection=False)
BlackHoleSpins(ax, 0.93828, [3e-11, 2e-15], text_on=False)
plt.text(1.5e-12, 1e-14, r'{\bf Black}', fontsize=25, ha='center')
plt.text(1.5e-12, 1e-14 / 4, r'{\bf Hole}', fontsize=25, ha='center')
plt.text(1.5e-12, 1e-14 / 16, r'{\bf Spins}', fontsize=25, ha='center');
MySaveFig(fig, 'AxionProton')
# +
fig, ax = FigSetup(Shape='Rectangular', ylab='$|g_{ap}|$', mathpazo=True,
g_min=1e-17, g_max=1e-2, m_min=1e-22, m_max=1e-2, FrequencyAxis=True, N_Hz=1)
AxionProton.QCDAxion(ax)
AxionProton.Haloscopes(ax, projection=True)
AxionProton.StellarBounds(ax)
AxionProton.LabExperiments(ax, projection=True)
BlackHoleSpins(ax, 0.93828, [3e-11, 2e-15], text_on=False)
plt.text(1.5e-12, 0.2e-14, r'{\bf Black}', fontsize=25, ha='center')
plt.text(1.5e-12, 0.2e-14 / 4, r'{\bf Hole}', fontsize=25, ha='center')
plt.text(1.5e-12, 0.2e-14 / 16, r'{\bf Spins}', fontsize=25, ha='center');
MySaveFig(fig, 'AxionProton_with_Projections')
# -
| notebooks/AxionProton.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.1 64-bit (conda)
# name: python3
# ---
# # Exploratory Data Analysis
#
# At this stage data is analysed to capture relationships between the target variable: Brewing Method, and potential predictors: other variables in the table.
# Hence, removing potential outliers.
#
# 1. First data is read from csv file with clean dataset (clean in the previous project) and transformed into a dataframe.
# 2. Then data is plotted for each of the variables against the target:
# - check if there is no direct relationships between predictor and target
# - spot outliers
# - observe the data distribution
#
# *Note:* Some of the variables from the original dataset were omitted:
# - price per kg - price does not influence the brewing method, it can be a result of the latter
# - grind - the state of the beans is irrelevant for the target variable
# - variables previously put into higher positioning were also omitted
import pandas as pd # import necessary library
coffee_df = pd.read_csv('data\coffee_desk_dataset_clean.csv', index_col='idx') # read data to data frame
coffee_df
# ## Data plotting: brewing method
#
# ### Step 1
# I am first plotting the count of tarbet variable to see the distribution of labels.
coffee_df['brewing_method'].value_counts().plot(kind='bar')
coffee_df['brewing_method'].value_counts()
# ### Observation:
# Data is visibly evenly distributed among two principal classes.
# There is the third class of coffees, which can be used for both brewing with traditioanl espresso style or using more altenative techniques.
# Since the principal objective of the client is to be able to buy and offer only the beans which are applicable for the alternative brewing methods (labelled as DRIP), this moxed labels can be changed to DRIP, as in this case they are also good for DRIP but might be cheaper to source, so it would be a missed chance on buiyng cheaper raw materials.
coffee_df['brewing_method_binary'] = coffee_df['brewing_method'].replace({'drip, espresso':'drip'})
coffee_df['brewing_method_binary'].value_counts()
# ## Data plotting: predictor vs. brewing method
#
# ### Step 2
# I am plotting some of the variables I already suspect of outliers. I start with the region of origin (knowing that espresso is a popular style in Italy, Europe), hence using expert knowledge to aid with data analysis.
import plotly.express as px
import seaborn as sns
sns.catplot(x='brewing_method_binary', y='origin_region', data=coffee_df, kind="strip")
# ### Observation:
# It is visible that data from Europe is an outlier, given that from this origin only espresso beans proceed. Hence, it would add unnecessary noise to data and it will be removed.
coffee_df = coffee_df[coffee_df["origin_region"] != "Europe"] # removing data with Europe as origin
coffee_df
# ### Step 3
#
# I also plot process, as I presume some of them to be very specific and hence having direct connection to brewing method.
sns.catplot(x='brewing_method_binary', y='process_general', data=coffee_df, kind="strip")
# ### Observation:
# Another variables to be removed are process types: experimental, cryo and monsooning, as they are clearly pointing to one brewing method, hence no machine learning would be needed to correlate them with the target variable, just as in case of Europe before.
coffee_df = coffee_df[coffee_df["process_general"] != "Experimental"] # removing the rows with unwanted process types
coffee_df = coffee_df[coffee_df["process_general"] != "Monsooning"]
coffee_df = coffee_df[coffee_df["process_general"] != "CRYO"]
coffee_df
# ### Step 4
#
# I now proceed to plotting the only numerical variable.
arabica_fig = px.scatter(coffee_df, x='arabica_percentage', y='brewing_method_binary')
arabica_fig.show()
# ### Observation:
# Another observation to be made is the fact that coffees for drip (alternative) brewing are only pure blends, not mixed between Arabica and Robusta. This relationship can be already drawn from Pue arabica flag variable, hence, this variable will be dropped from dataset.
# ## Plotting categorical variables vs. target variable (brewing method)
#
# ### Step 5
#
# Now I can plot data for two classes only, and the distribution against different categorical variables.
categorical_variables = ['process_general', 'origin_region', 'roast', 'pure_arabica', 'washed', 'natural', 'fermented_traditional', 'fermented_closed_tank']
# +
from plotly.subplots import make_subplots
import plotly.graph_objects as go
rows = (len(categorical_variables) // 2)
cols = 2
subplot_titles = tuple(cat + " vs brewing method" for cat in categorical_variables)
fig = make_subplots(rows=rows, cols=cols, subplot_titles=subplot_titles)
for i, cat in enumerate(categorical_variables):
row = (i // cols) + 1
col = (i % cols) + 1
fig.add_trace(go.Violin(
x=coffee_df[cat], y=coffee_df["brewing_method_binary"], name=cat, box_visible=False
), row=row, col=col)
fig.update_xaxes(patch=dict(type='category', categoryorder='mean ascending'), row=row, col=col)
fig.update_layout(height=2000, width=1500)
fig.show()
# -
# ### Observation:
# Washed and Natural flags add little information given data distribustion, there is no observable patter, hence these variables might be dropped from dataset. There also seems to be direct relation between the brewing method and roast, as well as brewing method and pure arabica flag, which will be analysed further below.
# ## Logistic Regression Data Modelling
#
# ### Step 6
# I will now try to model data using logistic regression from statmodels library with logit function.
# Before I can use it, I need to convert my target variable into binary labels. I' setting drip to 1 as the desired class, and espresso as 0.
coffee_df['brewing_method_binary_num'] = coffee_df['brewing_method_binary'].replace({'drip':1, 'espresso':0})
coffee_df['brewing_method_binary_num'].value_counts()
# ### Step 7
#
# I then prepare a string to be passed to the function, to save time typing.
categorical_variables = ['process_general, Treatment(reference="Honey")', 'origin_region', 'roast', 'pure_arabica', 'washed', 'natural', 'fermented_traditional', 'fermented_closed_tank']
categorical_list = ["C({})".format(var) for var in categorical_variables]
categorical_string = " + ".join(categorical_list)
formula_string = "brewing_method_binary_num ~ {}".format(categorical_string)
formula_string
# +
import statsmodels.formula.api as smf
baseline_model = smf.logit(formula_string, coffee_df).fit()
baseline_model.summary()
# -
# ### Observations:
# From the warning (below the model details) we see that there are variables which can directly point to the target variable, hence making predictions redundan Albert and Anderson (1984) define this as, “there is a vector α that correctly allocates all observations to their group.”
# * Analysing p-values, I come to a conclution that I should drop the following variable values:
# * process_general - Honey
# * process_general - Hybrid
# * process_general - Natural
# * natural - True
# * And possibly drop also the ones with p-value close to 1:
# * process_general - Semi-washed
# * roast - light
# * pure_arabica - True
# * fermented_traditional - True
#
# I will begin by omitting process_general, which seems to have genarated the most issues.
# ### Step 8
#
# I continue with backward stepwise selection.
categorical_variables = ['origin_region', 'roast', 'pure_arabica', 'washed', 'natural', 'fermented_traditional', 'fermented_closed_tank']
categorical_list = ["C({})".format(var) for var in categorical_variables]
categorical_string = " + ".join(categorical_list)
formula_string_2 = "brewing_method_binary_num ~ {}".format(categorical_string)
formula_string_2
# +
import statsmodels.formula.api as smf
baseline_model_2 = smf.logit(formula_string_2, coffee_df).fit()
baseline_model_2.summary()
# -
# I will also remove roast and pure arabica, which seem directly pointing to the brewing method.
categorical_variables = ['origin_region', 'washed', 'natural', 'fermented_traditional', 'fermented_closed_tank']
categorical_list = ["C({})".format(var) for var in categorical_variables]
categorical_string = " + ".join(categorical_list)
formula_string_3 = "brewing_method_binary_num ~ {}".format(categorical_string)
formula_string_3
# +
import statsmodels.formula.api as smf
baseline_model_3 = smf.logit(formula_string_3, coffee_df).fit()
baseline_model_3.summary()
# -
# ### Step 9
#
# I will finally omit the Washed flag, which still has higher p-value.
categorical_variables = ['origin_region', 'natural', 'fermented_traditional', 'fermented_closed_tank']
categorical_list = ["C({})".format(var) for var in categorical_variables]
categorical_string = " + ".join(categorical_list)
formula_string_4 = "brewing_method_binary_num ~ {}".format(categorical_string)
formula_string_4
# +
import statsmodels.formula.api as smf
baseline_model_4 = smf.logit(formula_string_4, coffee_df).fit()
baseline_model_4.summary()
# -
# ### Step 10
#
# After the initial data modelling I have decided to keep the variables: 'origin_region', 'natural', 'fermented_traditional', 'fermented_closed_tank', with lowest p-values.
# Nevertheless, this *backward stepwise selection* is only an attempt at getting the appropriate, statistically significant variables. And I might still need to go back and change the variables I have selected for final model.
# ### Step 11
#
# Save analysed and cleaned data to a new file.
columns = ['origin_region', 'natural', 'fermented_traditional', 'fermented_closed_tank', 'brewing_method_binary_num']
coffee_df.to_csv('data\coffee_desk_dataset_ead_selected.csv', columns=columns)
| tailored_brewing_eda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
from matplotlib import pyplot as plt
import pandas as pd
from scipy.interpolate import splev, splrep
import numpy as np
from matplotlib import rcParams, colors
import seaborn as sns
from matplotlib.colors import ListedColormap
# %matplotlib inline
rcParams['font.sans-serif'] = 'arial'
plot_params = {'markersize': 5.5,
'markeredgewidth': 1.2,
'color': 'white'}
pal = sns.xkcd_palette(['dark sky blue', 'deep red']).as_hex()
h1color=pal[0]
h3color=pal[1]
def get_param(imprinting_file, param, group_name):
df = pd.read_csv(imprinting_file).dropna()
df['CI_upper'] = df.prof_max
df['CI_lower'] = df.prof_min
df.index = df.param
df['group_name'] = group_name
param_values = df.loc[param, :]
return param_values
# Plotting H1N1 results. Profiles are required to make these plots.
all_ages = get_param('../final_results_for_ms/0-100/DAHVage_subtype.profile_liks.csv', 'H1m', '0-100')
exclude_kids = get_param('../final_results_for_ms/15-100/DAHVage_subtype.profile_liks.csv', 'H1m', '15-100')
exclude_adults = get_param('../final_results_for_ms/0-64/DAHVage_subtype.profile_liks.csv', 'H1m', '0-64')
middle_ages = get_param('../final_results_for_ms/15-64/DAHVage_subtype.profile_liks.csv', 'H1m', '15-64')
plotdf = pd.DataFrame([all_ages, exclude_kids ,exclude_adults, middle_ages])
yval = -0.15
for index, row in plotdf.iterrows():
label = row.group_name
plt.plot([row.mle * 100], [-yval], 'o', **plot_params, markeredgecolor=h1color, zorder=1)
plt.hlines(y=-yval, xmin=row.CI_lower*100, xmax=row.CI_upper*100, linewidth=1)
plt.vlines(x=row.CI_lower*100, ymin=-yval-0.2, ymax=-yval+0.2, linewidth=1)
plt.vlines(x=row.CI_upper*100, ymin=-yval-0.2, ymax=-yval+0.2, linewidth=1)
yval += 1
# Plotting H3N2 results. Profiles are required to make these plots.
all_ages = get_param('../final_results_for_ms/0-100/DAHVage_subtype.profile_liks.csv', 'H3m', '0-100')
exclude_kids = get_param('../final_results_for_ms/15-100/DAHVage_subtype.profile_liks.csv', 'H3m', '15-100')
exclude_adults = get_param('../final_results_for_ms/0-64/DAHVage_subtype.profile_liks.csv', 'H3m', '0-64')
middle_ages = get_param('../final_results_for_ms/15-64/DAHVage_subtype.profile_liks.csv', 'H3m', '15-64')
plotdf = pd.DataFrame([all_ages,exclude_kids,exclude_adults,middle_ages])
yval = 0.15
for index, row in plotdf.iterrows():
label = row.group_name
plt.plot([row.mle * 100], [-yval], 'o', **plot_params, markeredgecolor=h3color, zorder=1)
plt.hlines(y=-yval, xmin=row.CI_lower * 100, xmax=row.CI_upper*100, linewidth=1)
plt.vlines(x=row.CI_lower * 100, ymin=-yval-0.2, ymax=-yval+0.2, linewidth=1)
plt.vlines(x=row.CI_upper * 100, ymin=-yval-0.2, ymax=-yval+0.2, linewidth=1)
yval += 1
yval = 0
yticks = []
ylabs = []
for index, row in plotdf.iterrows():
label = row.group_name
yticks.append(yval)
ylabs.append(label)
yval += -1
plt.plot([100], [100], 'o', **plot_params, markeredgecolor=h1color, label='H1N1')
plt.plot([100], [100], 'o', **plot_params, markeredgecolor=h3color, label='H3N2')
plt.legend(ncol=2, loc='center', bbox_to_anchor=(0.5, 1.1))
plt.yticks(yticks,ylabs)
plt.ylabel('Age group (years)', weight='bold')
plt.xlabel('Imprinting protection (%)', weight='bold')
plt.xlim(-5, 105)
plt.ylim(-3.4, 0.4)
plt.gcf().set_size_inches(3, 3)
plt.tight_layout()
# -
# # Ranking by MSE
# +
mse = pd.read_csv('../final_results_for_ms/0-100/loo/loo_mse.csv', index_col=0)
pal = sns.xkcd_palette(['dark sky blue', 'deep red']).as_hex()
h1color=pal[0]
h3color=pal[1]
rcParams['mathtext.default'] = 'regular'
rcParams['font.sans-serif'] = 'arial'
sns.set_context('paper')
# just a dummy dataframe for now, MSE is stored above in the mse dataframe
df = pd.read_csv('../final_results_for_ms/0-100/loo/result_summary.csv', index_col='Unnamed: 0')
pal = sns.color_palette('colorblind').as_hex()
flatui = ['white', pal[3], 'darkgreen', 'lightgreen']
my_cmap = ListedColormap(sns.color_palette(flatui).as_hex())
rcParams['font.sans-serif'] = 'arial'
sns.set_context('paper')
full_model='DAHNV'
final_df = pd.DataFrame(columns=['D', 'E', 'Ap', 'vac_cov', 'Nu', 'A', 'N2', 'H_sub','H_group', 'V_constant', 'V_age', 'V_imprinting', 'V_cohort', 'mse'])
row = 0
df = df.iloc[1:, ]
exclude = ['DAHVage_subtype', 'DAHVcohort_subtype', 'DAHNVseason_subtype', 'DAHNVseason_group', 'DAVseason', 'DAVcohort', 'DAVimprinting', 'DAVage']
for model, r in df.iterrows():
if model not in exclude:
if 'Vage' in model:
V = 'V_age'
final_df.loc[row, V] = 1
elif 'Vseason' in model:
V = 'V_season'
final_df.loc[row, V] = 1
elif 'Vimprinting' in model:
V = 'V_imprinting'
final_df.loc[row, V] = 1
elif 'Vcohort' in model:
V = 'V_cohort'
final_df.loc[row, V] = 1
elif 'Vmean' in model:
V = 'V_constant'
final_df.loc[row, V] = 1
if 'H' in model:
if 'subtype' in model:
final_df.loc[row, 'H_sub'] = 1
elif 'group' in model:
final_df.loc[row, 'H_group'] = 1
if 'N' in model:
if r['N2m'] != 0:
final_df.loc[row, 'N2'] = 0.5
else:
final_df.loc[row, 'N2'] = 0.5
final_df.loc[row, 'A'] = 1
final_df.loc[row, 'D'] = 0.25
final_df.loc[row, 'E'] = 0.25
final_df.loc[row, 'Ap'] = 0.25
final_df.loc[row, 'vac_cov'] = 0.25
final_df.loc[row, 'Nu'] = 0.25
#final_df.loc[row, '']
final_df.loc[row, 'mse'] = mse.loc[model, 'mse']
row += 1
final_df = final_df.sort_values('mse')
final_df = final_df.fillna(0)
#final_df['cAIC'] = [np.exp(-0.5 * (c - min(final_df['cAIC']))) for c in final_df['cAIC']]
#final_df.index = ["%.4f" % (c/sum(final_df.cAIC)) for c in final_df['cAIC']]
final_df.index = ["%.2f" % c for c in final_df['mse']]
final_df = final_df.loc[:, final_df.columns != 'mse']
final_df.columns = ['Demography',
'Enrollment fraction',
'Approachment fraction',
'Healthcare-seeking behavior among vaccinated',
'Nursing home residency',
'Age-specific risk of medically attended influenza A infection',
'N2 imprinting',
'HA imprinting (subtype)',
'HA imprinting (group)',
'Vaccine effectiveness (constant)',
'Vaccine effectiveness (age-specific)',
'Vaccine effectiveness (imprinting-specific)',
'Vaccine effectiveness (cohort-specific)']
sns.heatmap(final_df, cmap=my_cmap, linewidths=1, linecolor='black', cbar=False, yticklabels=1)
ax = plt.gca()
ax.xaxis.tick_top()
plt.yticks(rotation=0, fontsize=10)
plt.xticks(rotation=45, ha='left', weight='bold')
plt.ylabel('MSE', weight='bold')
f = plt.gcf()
f.set_size_inches(5.5, 5)
plt.tight_layout()
# -
# # Prediction of excluded seasons: Unvaccinated cases
# +
inc_data = pd.read_csv('../data/standard_eligible_observed.csv')
seasons = sorted(list(range(2008, 2019)))
h1_seasons = [2009, 2010, 2014, 2016]
p = 1
for season in seasons:
loo = pd.read_csv('../data/out_of_sample_checks/loo_simulation_%s.csv'%season, index_col=0)
if season in h1_seasons:
st = 'h1'
color = 'blue'
else:
st = 'h3'
color = 'red'
if season == 2009.5:
seasonstr = '2009Pan'
title = '2009 pandemic'
else:
seasonstr = str(season)
title = '%s-%s'%(season-1, season)
color='purple'
temp = inc_data[inc_data.season == seasonstr]
loounvac = loo.loc[[str(y) for y in temp.birth_year], ].transpose()
upperunvac = loounvac.quantile(0.975).reset_index()
lowerunvac = loounvac.quantile(0.025).reset_index()
meanunvac = loounvac.mean().reset_index()
loovac = loo.loc[['vac' + str(y) for y in temp.birth_year], ].transpose()
uppervac = loovac.quantile(0.975).reset_index()
lowervac = loovac.quantile(0.025).reset_index()
meanvac = loovac.mean().reset_index()
plt.subplot(4, 3, p)
ax = plt.gca()
temp.plot.scatter(x='birth_year',
y='I_obs_%s'%st,
ax=ax,
alpha=0.7,
edgecolor='k',
s=3,
color=color,
label='Observed')
plt.plot(temp.birth_year,
meanunvac[0],
'-',
color=color,
label='Mean prediction')
ax.fill_between(x=temp.birth_year,
y1=upperunvac[0.975],
y2=lowerunvac[0.025],
alpha=0.3,
color=color,
label='95% prediction interval')
plt.xticks(np.arange(1920, 2016, 15), rotation=45, ha='right')
plt.xlim(1915, 2019)
if p in [1,4,7,10]:
plt.ylabel('Unvaccinated\ncases')
else:
plt.ylabel('')
if p in[9, 10, 11, 12]:
plt.xlabel('Birth year')
else:
plt.xlabel('')
plt.xticks(np.arange(1920, 2016, 15), [])
if p != 12:
ax.legend().set_visible(False)
plt.title(title)
p += 1
plt.gcf().set_size_inches(7,5)
plt.gcf().align_ylabels()
plt.tight_layout()
plt.legend(loc='center', bbox_to_anchor=(0.5,-1.8), ncol=3)
# -
# # Prediction of excluded seasons: Vaccinated cases
# +
inc_data = pd.read_csv('../data/standard_eligible_observed.csv')
seasons = sorted(list(range(2008, 2019)))
h1_seasons = [2009, 2010, 2014, 2016]
p = 1
for season in seasons:
loo = pd.read_csv('../data/out_of_sample_checks/loo_simulation_%s.csv'%season, index_col=0)
if season in h1_seasons:
st = 'h1'
color = 'blue'
else:
st = 'h3'
color = 'red'
if season == 2009.5:
seasonstr = '2009Pan'
title = '2009 pandemic'
else:
seasonstr = str(season)
title = '%s-%s'%(season-1, season)
color='purple'
temp = inc_data[inc_data.season == seasonstr]
loounvac = loo.loc[[str(y) for y in temp.birth_year], ].transpose()
upperunvac = loounvac.quantile(0.975).reset_index()
lowerunvac = loounvac.quantile(0.025).reset_index()
meanunvac = loounvac.mean().reset_index()
loovac = loo.loc[['vac' + str(y) for y in temp.birth_year], ].transpose()
uppervac = loovac.quantile(0.975).reset_index()
lowervac = loovac.quantile(0.025).reset_index()
meanvac = loovac.mean().reset_index()
plt.subplot(4, 3, p)
ax = plt.gca()
temp.plot.scatter(x='birth_year',
y='I_vac_%s'%st,
ax=ax,
alpha=0.7,
edgecolor='k',
s=3,
color=color,
label='Observed')
plt.plot(temp.birth_year,
meanvac[0],
'-',
color=color,
label='Mean prediction')
ax.fill_between(x=temp.birth_year,
y1=uppervac[0.975],
y2=lowervac[0.025],
alpha=0.3,
color=color,
label='95% prediction interval')
plt.xticks(np.arange(1920, 2016, 15), rotation=45, ha='right')
plt.xlim(1915, 2019)
if p in [1,4,7,10]:
plt.ylabel('Vaccinated\ncases')
else:
plt.ylabel('')
if p in[9, 10, 11, 12]:
plt.xlabel('Birth year')
else:
plt.xlabel('')
plt.xticks(np.arange(1920, 2016, 15), [])
if p != 12:
ax.legend().set_visible(False)
plt.title(title)
p += 1
plt.gcf().set_size_inches(7,5)
plt.gcf().align_ylabels()
plt.tight_layout()
plt.legend(loc='center', bbox_to_anchor=(0.5,-1.8), ncol=3)
# -
# # Extrapolating the starts and ends of seasons
# +
import pandas as pd
from dateutil import relativedelta
from datetime import datetime, timedelta
from matplotlib import pyplot as plt
from matplotlib import rcParams
import numpy as np
import pymmwr
from scipy import stats
import seaborn as sns
# %matplotlib inline
rcParams['font.sans-serif'] = 'arial'
cases_per_sampling_day = pd.read_csv('../raw_data/cases_per_sampling_day.csv')
age_classes = [(0, 4),
(5, 9),
(10, 14),
(15, 19),
(20, 29),
(30, 39),
(40, 49),
(50, 64),
(65, 200)]
def dominant_subtype(year):
if year in ['2009', '2009Pan', '2010', '2014', '2016']:
subtypes = ['H1N1', 'H1N1pdm']
else:
subtypes = ['H3N2']
return subtypes
def age_to_age_class(age):
for l, u in age_classes:
if age <= u and age >= l:
age_class = '-'.join([str(l), str(u)])
break
if age_class == '65-200':
age_class = '65+'
return age_class
def season_to_season_float(season_str):
if season_str == '2009Pan':
s = 2009.5
else:
s = float(season_str)
return s
def event_date_to_season_week(date):
event = datetime.strptime(date, '%Y-%m-%d').date()
mmwr_date = pymmwr.date_to_epiweek(event)
epiweek = pymmwr.Epiweek(mmwr_date.year, mmwr_date.week)
return pymmwr.epiweek_to_date(epiweek)
subplot_index = {'2008': 1,
'2009': 2,
'2009Pan': 3,
'2010': 4,
'2011': 5,
'2012': 6,
'2013': 7,
'2014': 8,
'2015': 9,
'2016': 10,
'2017': 11,
'2018': 12}
seasons_to_extrapolate = ['2013', '2014', '2009', '2008', '2015', '2018']
cases_per_sampling_day['Week'] = pd.to_datetime(cases_per_sampling_day['Week'])
cases_to_add = []
pal = sns.color_palette('Set2').as_hex()
fig, axes = plt.subplots(nrows=4, ncols=3)
axes = axes.flatten()
for season, seasondf in cases_per_sampling_day.groupby('Season'):
subplot = subplot_index[season] - 1
if season in seasons_to_extrapolate:
plt.sca(axes[subplot])
plotdf = seasondf.copy()
plotdf.index = plotdf.Week
plotdf = plotdf.fillna(0)
peak_week = plotdf[plotdf['Cases per sampling day'] == max(plotdf['Cases per sampling day'])].index[0]
data_subset = plotdf[plotdf.Week < peak_week].copy()
x = [date.toordinal() for date in data_subset.Week]
y = list(data_subset['Cases per sampling day'])
m, b, r, p, e = stats.linregress(x, y)
x0 = min(x)
y0 = y[0]
new_x = [datetime.fromordinal(X) for X in x]
new_y = [m * X + b for X in x]
case_extrap = []
while y0 > 1:
x0 = x0 - 7
y0 = m * x0 + b
x_test = pymmwr.date_to_epiweek(datetime.fromordinal(x0).date())
new_x.append(datetime.fromordinal(x0))
new_y.append(y0)
case_extrap.append([x_test.year, x_test.week, np.ceil(y0 * 7)])
case_extrap = pd.DataFrame(case_extrap, columns=['Year', 'Week', 'Cases'])
new_x, new_y = zip(*sorted(zip(new_x, new_y)))
plt.plot(new_x, new_y, '--', color=pal[1], label='Extrapolated')
cases_to_add.append([season, 'start', case_extrap[case_extrap.Cases >0].sum().Cases])
# extrapolate ends
seasons_to_extrapolate = ['2008','2009Pan','2016']
for season, seasondf in cases_per_sampling_day.groupby('Season'):
subplot = subplot_index[season] - 1
plt.sca(axes[subplot])
if season in seasons_to_extrapolate:
plotdf = seasondf.copy()
plotdf.index = plotdf.Week
plotdf = plotdf.fillna(0)
peak_week = plotdf[plotdf['Cases per sampling day'] == max(plotdf['Cases per sampling day'])].index[0]
data_subset = plotdf[plotdf.Week > peak_week].copy()
x = [date.toordinal() for date in data_subset.Week]
y = list(data_subset['Cases per sampling day'])
m, b, r, p, e = stats.linregress(x, y)
x0 = max(x)
y0 = y[-1]
new_x = [datetime.fromordinal(X) for X in x]
new_y = [m * X + b for X in x]
case_extrap = []
while y0 > 1:
x0 = x0 + 7
y0 = m * x0 + b
x_test = pymmwr.date_to_epiweek(datetime.fromordinal(x0).date())
new_x.append(datetime.fromordinal(x0))
new_y.append(y0)
case_extrap.append([x_test.year, x_test.week, np.ceil(y0 * 7)])
case_extrap = pd.DataFrame(case_extrap, columns=['Year', 'Week', 'Cases'])
new_x, new_y = zip(*sorted(zip(new_x, new_y)))
plt.plot(new_x, new_y, '--', color=pal[1], label='Extrapolated')
cases_to_add.append([season, 'end', case_extrap[case_extrap.Cases >0].sum().Cases])
plt.ylabel('')
plt.xlabel('')
for season, seasondf in cases_per_sampling_day.groupby('Season'):
subplot = subplot_index[season] - 1
plt.sca(axes[subplot])
plotdf = seasondf.copy()
plotdf.index = plotdf.Week
plotdf = plotdf.fillna(0)
new_x = [datetime.strptime(str(d).split()[0], '%Y-%m-%d') for d in plotdf['Week']]
plt.plot(new_x, plotdf['Cases per sampling day'],
'o', color=pal[0], alpha=0.5, label='Observed')
plt.ylabel('Cases per day')
if season != '2009Pan':
plt.title(str(int(season) - 1) + '-' + season)
else:
plt.title('2009 pandemic')
plt.axhline(1, linestyle='-', linewidth=1, color='black', zorder=1)
#plt.draw()
#ticks, labels = plt.xticks()
#plt.xticks(ticks, labels, rotation=45, ha='right', size=8)
plt.xticks([new_x[int(i)] for i in range(0, len(new_x), 5)],
[datetime.strftime(new_x[int(i)], '%Y-%m-%d') for i in range(0, len(new_x), 5)],
rotation=45,
ha='right')
if subplot in [9, 10, 11]:
plt.xlabel('Start date of week')
else:
plt.xlabel('')
if subplot in [0, 3, 6, 9]:
plt.ylabel('Cases per\nsampling day')
else:
plt.ylabel('')
plt.gcf().align_labels()
plt.gcf().set_size_inches(6, 7)
plt.tight_layout()
plt.legend(loc='center', bbox_to_anchor=(-1.2, -1.7), ncol=2)
# Output extrapolated cases for sensitivity analyses
outdf = pd.DataFrame(cases_to_add, columns=['Season','Period','Cases'])
outdf.to_csv('../data/extrapolated_case_totals.csv')
# -
# # Sensitivity to start and end of season
# +
import glob
import pandas as pd
import seaborn as sns
from matplotlib import pyplot as plt
from matplotlib import rcParams
from matplotlib.colors import ListedColormap
import numpy as np
# %matplotlib inline
df = pd.read_csv('../final_results_for_ms/0-100/sensitivity_summary.csv')
result = pd.read_csv('../final_results_for_ms/0-100/DAHVage_subtype.profile_liks.csv', index_col='param')
pal = sns.xkcd_palette(['dark sky blue', 'deep red']).as_hex()
rcParams['font.sans-serif'] = 'arial'
H1_mle = result.loc['H1m', 'mle']
H3_mle = result.loc['H3m', 'mle']
new_rows = []
for index, row in df.iterrows():
new_rows.append(['H1N1', row.H1m * 100])
new_rows.append(['H3N2', row.H3m * 100])
plotdf = pd.DataFrame(new_rows, columns=['Subtype', 'Imprinting protection (%)'])
sns.set_palette(pal)
sns.violinplot(data=plotdf, x='Subtype', y='Imprinting protection (%)', alpha=0.2, inner=None)
plt.setp(plt.gca().collections, alpha=0.4)
plt.plot([0], [H1_mle * 100], 'o', markeredgecolor='black', label='H1 imprinting strength\nwith no simulated cases',
alpha=0.7)
plt.plot([0],
[np.median(plotdf[plotdf.Subtype=='H1N1']['Imprinting protection (%)'])],
's',
markeredgecolor='black',
label='Median H1 imprinting protection\nfrom simulations',
alpha=0.7,
color=pal[0])
plt.vlines(x=0, ymin=result.loc['H1m', 'prof_min'] * 100, ymax=result.loc['H1m', 'prof_max'] * 100)
plt.plot([1], [H3_mle * 100], 'o', markeredgecolor='black', label='H3 imprinting strength\nwith no simulated cases', alpha=0.7)
plt.plot([1],
[np.median(plotdf[plotdf.Subtype=='H3N2']['Imprinting protection (%)'])],
's',
markeredgecolor='black',
label='Median H3 imprinting protection\nfrom simulations',
alpha=0.7,
color=pal[1])
plt.vlines(x=1, ymin=result.loc['H3m', 'prof_min'] * 100, ymax=result.loc['H3m', 'prof_max'] * 100)
plt.legend()
# -
| figures/Figure_5_imprinting_results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: myenv_new_jupyter
# language: python
# name: myenv_new_jupyter
# ---
# +
# %reset
import sys, os
import numpy as np
import matplotlib
matplotlib.use('Agg')
import matplotlib.pyplot as plt
import scipy.integrate as integrate
import random
import healpy as hp
from astropy.io import fits
from astropy.coordinates import SkyCoord
from numpy.random import rand
import pickle as pk
import matplotlib.cm as cm
import scipy.interpolate as interpolate
import pdb
import time
import multiprocessing as mp
sys.path.insert(0,'/global/project/projectdirs/des/shivamp/cosmosis/y3kp-bias-model/3d_stats/process_measure_data/')
import correlate_gg_gm_3d_class as corr_class
import argparse
import subprocess
# + jupyter={"outputs_hidden": true}
# !python '/global/project/projectdirs/des/shivamp/cosmosis/y3kp-bias-model/3d_stats/process_measure_data/process_cats_funcs_mice_halos.py'
# +
# !python '/global/project/projectdirs/des/shivamp/cosmosis/y3kp-bias-model/3d_stats/process_measure_data/correlate_gg_gm_3d_funcs_mice_halos.py' --bin 5 --lm_min 12.0 --lm_max 12.5 --do_mm 0
# +
# !python '/global/project/projectdirs/des/shivamp/cosmosis/y3kp-bias-model/3d_stats/process_measure_data/correlate_gg_gm_3d_funcs_mice_halos.py' --bin 3 --lm_min 12.5 --lm_max 13.0 --do_mm 0
# +
# !python '/global/project/projectdirs/des/shivamp/cosmosis/y3kp-bias-model/3d_stats/process_measure_data/correlate_gg_gm_3d_funcs_mice_halos.py' --bin 1 --lm_min 12.5 --lm_max 13.0
# +
# !python '/global/project/projectdirs/des/shivamp/cosmosis/y3kp-bias-model/3d_stats/process_measure_data/correlate_gg_gm_3d_funcs_mice_halos.py' --bin 4 --lm_min 12.5 --lm_max 13.0 --do_gg 0 --do_gm 0 --do_mm 0
# +
# !python '/global/project/projectdirs/des/shivamp/cosmosis/y3kp-bias-model/3d_stats/process_measure_data/correlate_gg_gm_3d_funcs_mice_halos.py' --bin 2 --lm_min 13.0 --lm_max 13.5 --do_gg 1 --do_gm 1 --do_mm 1
# +
# !python '/global/project/projectdirs/des/shivamp/cosmosis/y3kp-bias-model/3d_stats/process_measure_data/correlate_gg_gm_3d_funcs_mice_halos.py' --bin 2 --lm_min 12.5 --lm_max 13.0
# +
# !python '/global/project/projectdirs/des/shivamp/cosmosis/y3kp-bias-model/3d_stats/process_measure_data/correlate_gg_gm_3d_funcs_mice_halos.py' --bin 3 --lm_min 12.5 --lm_max 13.0
# +
# !python '/global/project/projectdirs/des/shivamp/cosmosis/y3kp-bias-model/3d_stats/process_measure_data/correlate_gg_gm_3d_funcs_mice_halos.py' --bin 1 --lm_min 13.0 --lm_max 13.5 --do_gg 1 --do_gm 1 --do_mm 1
# +
# !python '/global/project/projectdirs/des/shivamp/cosmosis/y3kp-bias-model/3d_stats/process_measure_data/correlate_gg_gm_3d_funcs_mice_halos.py' --bin 3 --lm_min 13.0 --lm_max 13.5 --do_gg 0 --do_gm 0 --do_mm 1
# +
# !python '/global/project/projectdirs/des/shivamp/cosmosis/y3kp-bias-model/3d_stats/process_measure_data/correlate_gg_gm_3d_funcs_mice_halos.py' --bin 4 --lm_min 13.0 --lm_max 13.5
# +
# !python '/global/project/projectdirs/des/shivamp/cosmosis/y3kp-bias-model/3d_stats/process_measure_data/correlate_gg_gm_3d_funcs_mice_halos.py' --bin 5 --lm_min 13.0 --lm_max 13.5
# -
# !pip install pickle --user
# +
# !python '/global/project/projectdirs/des/shivamp/cosmosis/y3kp-bias-model/3d_stats/process_measure_data/correlate_gg_gm_3d_funcs_mice_halos.py' --bin 3 --lm_min 13.5 --lm_max 14.0 --do_gg 1 --do_gm 1 --do_mm 1
# +
# !python '/global/project/projectdirs/des/shivamp/cosmosis/y3kp-bias-model/3d_stats/process_measure_data/correlate_gg_gm_3d_funcs_mice_halos.py' --bin 1 --lm_min 14.0 --lm_max 14.5
# +
# !python '/global/project/projectdirs/des/shivamp/cosmosis/y3kp-bias-model/3d_stats/process_measure_data/correlate_gg_gm_3d_funcs_mice_halos.py' --bin 4 --lm_min 14.0 --lm_max 14.5
# -
| notebooks/Untitled3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [jupyter notebook 内でのグラフ表示 - Qiita](https://qiita.com/samacoba/items/81093984605abfed70d1)
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
x = np.linspace(0, 10, 100)
y = np.sin(x)
plt.plot(x, y)
# %matplotlib notebook
import matplotlib.pyplot as plt
import numpy as np
x = np.linspace(0, 10, 100)
y = np.sin(x)
plt.plot(x, y)
| markdown/python/anaconda/notebook/Graph/graph01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
from astropy.io import fits
from astropy.io import ascii as asc
import numpy as np
from matplotlib import pyplot as plt
# %matplotlib inline
from spectroscopy import calc_wavelength, apply_redshift, scale_spectra, spectrum1d
# -
spec_file = '../data/spectra/lco/asassn15oz_20151006_redblu_101906.800.fits'
ofile = fits.open(spec_file)
flux = ofile[0].data[0,0,:]
wave = calc_wavelength(ofile[0].header, np.arange(len(flux))+1)
wave = apply_redshift(wave, 0.0069)
spec_obs = spectrum1d(wave, flux)
spec_fit = asc.read('../data/asassn15oz_1006_syn++.txt', names = ['wave', 'flux', 'err'])
spec_fit = spectrum1d(spec_fit['wave'], spec_fit['flux'])
spec_fit = scale_spectra(spec_fit, spec_obs)
plt.plot(spec_obs.wave, spec_obs.flux)
plt.plot(spec_fit.wave, spec_fit.flux)
| notebooks/syn++_comparison.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + hideCode=true
import os, sys
try:
from synapse.lib.jupyter import *
except ImportError as e:
# Insert the root path of the repository to sys.path.
# This assumes the notebook is located three directories away
# From the root synapse directory. It may need to be varied
synroot = os.path.abspath('../../../')
sys.path.insert(0, synroot)
from synapse.lib.jupyter import *
# + hideCode=true
# Get a temp cortex
core = await getTempCoreCmdr()
# + active=""
# .. highlight:: none
#
# .. _storm-ref-data-mod:
#
# Storm Reference - Data Modification
# ===================================
#
# Storm can be used to directly modify the Synapse hypergraph by:
#
# - adding or deleting nodes;
# - setting, modifying, or deleting properties on nodes; and
# - adding or deleting tags from nodes.
#
# Users gain a powerful degree of flexibility and efficiency through the ability to create or modify data on the fly.
#
# (**Note:** For adding or modifying data at scale, we recommend use of the Synapse ``csvtool`` (:ref:`syn-tools-csvtool`), the Synapse ``feed`` utility (:ref:`syn-tools-feed`), or the programmatic ingest of data.)
#
# .. WARNING::
# The ability to add and modify data directly from Storm is powerful and convenient, but also means users can inadvertently modify (or even delete) data inappropriately through mistyped syntax or premature striking of the "enter" key. While some built-in protections exist within Synapse itself it is important to remember that **there is no "are you sure?" prompt before a Storm query executes.**
#
# The following recommended best practices will help prevent inadvertent changes to a Cortex:
#
# - Use extreme caution when constructing complex Storm queries that may modify (or delete) large numbers of nodes. It is **strongly recommended** that you validate the output of a query by first running the query on its own to ensure it returns the expected results (set of nodes) before permanently modifying (or deleting) those nodes.
# - Use the Synapse permissions system to enforce least privilege. Limit users to permissions appropriate for tasks they have been trained for / are responsible for. See :ref:`initial-roles` in the :ref:`quickstart` guide for a basic discussion of users, roles, and permissions.
#
# See :ref:`storm-ref-syntax` for an explanation of the syntax format used below.
#
# See :ref:`storm-ref-type-specific` for details on special syntax or handling for specific data types (:ref:`data-type`).
#
# .. _edit-mode:
#
# Edit Mode
# ---------
#
# To modify data in a Cortex using Storm, you must enter “edit mode”. Edit mode makes use of several conventions to specify what changes should be made and to what data:
#
# - `Edit Brackets`_
# - `Edit Parentheses`_
# - `Edit "Try" Operator (?=)`_
# - `Autoadds and Depadds`_
#
# .. _edit-brackets:
#
# Edit Brackets
# +++++++++++++
#
# The use of square brackets ( ``[ ]`` ) within a Storm query can be thought of as entering edit mode. The data in the brackets specifies the changes to be made and includes changes involving nodes, properties, and tags. The only exception is the deletion of nodes, which is done using the Storm :ref:`storm-delnode` command.
#
# The square brackets used for the Storm data modification syntax indicate "perform the enclosed changes" in a generic way. The brackets are shorthand to request any of the following:
#
# - `Add Nodes`_
# - `Add or Modify Properties`_
# - `Delete Properties`_
# - `Add Light Edges`_
# - `Delete Light Edges`_
# - `Add Tags`_
# - `Modify Tags`_
# - `Remove Tags`_
#
# This means that all of the above directives can be specified within a single set of brackets, in any combination and in any order. The only caveat is that a node must exist before it can be modified, so you must add a node inside the brackets (or lift a node outside of the brackets) before you add a secondary property or a tag.
#
# .. WARNING::
# It is critical to remember that **the brackets are NOT a boundary that segregates nodes;** the brackets simply indicate the start and end of data modification operations. They do **NOT** separate "nodes the modifications should apply to" from "nodes they should not apply to". Storm :ref:`storm-op-chain` with left-to-right processing order still applies. **Any modification request that operates on previous Storm output will operate on EVERYTHING to the left of the modify operation, regardless of whether those nodes are within or outside the brackets.** The only exception is modifications that are placed within :ref:`edit-parens`.
#
# .. NOTE::
# For simplicity, syntax examples below demonstrating how to add nodes, modify properties, etc. only use edit brackets.
#
# See :ref:`data-mod-combo` below for examples showing the use of edit brackets with and without edit parentheses.
#
# .. _edit-parens:
#
# Edit Parentheses
# ++++++++++++++++
#
# Inside of :ref:`edit-brackets`, Storm supports the use of edit parentheses ( ``( )`` ). Edit parentheses ("parens") are used to explicitly limit a set of modifications to a specific node or nodes by enclosing the node(s) and their associated modification(s) within the parentheses. This "overrides" the default behavior for edit brackets, which is that every change specified within the brackets applies to every node generated by the previous Storm output (i.e., every node in the Storm pipeline), whether the node is referenced inside or outside the brackets themselves. Edit parens thus allow you to make limited changes "inline" with a more complex Storm query instead of having to use a smaller, separate query to make those changes.
#
# Similar to edit brackets, a node must exist or be specified before it can be modified, so the data within edit parens must start with the lift or creation of one or more nodes.
#
# In addition, multiple sets of edit parens can be used within a single set of edit brackets; each set of edit parens will delimit a separate set of edits.
#
# See :ref:`data-mod-combo` below for examples showing the use of edit brackets with and without edit parentheses.
#
# .. _edit-try:
#
# Edit "Try" Operator (?=)
# ++++++++++++++++++++++++
#
# Most edit operations will involve explicitly setting a primary or secondary property value using the equivalent ( ``=`` ) comparison operator:
#
# ``[ inet:fqdn = woot.com ]``
#
# ``inet:ipv4 = 1.2.3.4 [ :asn = 444 ]``
#
# Storm also supports the optional "try" operator ( ``?=`` ) within edit brackets or edit parens. The try operator will **attempt** to set a value that may or may not pass :ref:`data-type` enforcement for that property. Similarly, the try operator can also be used when setting tags, e.g. ``[ +?#mytag ]``.
#
# Incorrectly specifying a property value is unlikely to occur for users entering Storm data modification queries at the command line (barring outright user error), as users are directly vetting the data they are entering. However, the try operator may be useful for Storm-based automated ingest of data (such as :ref:`syn-tools-csvtool` or :ref:`syn-tools-feed`) where the data source may contain "bad" data.
#
# Use of the try operator allows Storm to fail silently in the event it encounters a ``BadTypeValu`` error (i.e., skip the bad event but continue processing). Contrast this behavior with using the standard equivalent operator ( ``=`` ), where if Storm encounters an error it will halt processing.
#
# See the :ref:`type-array` section of the :ref:`storm-ref-type-specific` for specialized "edit try" syntax when working with arrays.
#
# Autoadds and Depadds
# ++++++++++++++++++++
#
# Synapse makes use of two optimization features when adding nodes or setting secondary properties: automatic additions (:ref:`gloss-autoadd`) and dependent additions (:ref:`gloss-depadd`).
#
# **Autoadd** is the process where, on node creation, Synapse will automatically set any secondary properties that are derived from a node's primary property. Because these secondary properties are based on the node's primary property (which cannot be changed once set), the secondary properties are read-only.
#
# **Depadd** is the process where, on setting a node's secondary property value, if that property is of a type that is also a form, Synapse will automatically create the form with the corresponding primary property value. (You can view this as the secondary property "depending on" the existing of a node with the corresponding primary property.)
#
# Autoadd and depadd work together (and recursively) to simplify adding data to a Cortex. Properties set via autoadd may result in the creation of nodes via depadd; the new nodes may have secondary properties set via autoadd that result in the creation of additional nodes via depadd, and so on.
#
#
# **Examples:**
#
# .. NOTE::
# The specific syntax and process of node creation, modification, etc. are described in detail below. The examples here are simply meant to illustrate the autoadd and depadd concepts.
#
# *Create a node for the email address <EMAIL>. Note the secondary properties (:fqdn and :user) that are set via autoadd.*
# + hideCode=true
# Make some nodes
q = '[ inet:email = <EMAIL> ]'
# Display the syntax
# Run the query and test
podes = await core.eval(q, num=1, cmdr=True)
# + active=""
# *Create a node to represent the twitter account for The Vertex Project. Note the additional nodes that are created via deppadd (URL, FQDNs, user), as well as the secondary props that are set via autoadd.*
# + hideCode=true
# Remove nodes so we have a clean cortex
q = 'inet:email=<EMAIL> | delnode | inet:fqdn=vertex.link | delnode | inet:fqdn=link | delnode | inet:user=user | delnode'
# Make some nodes
q1 = '[ inet:web:acct=(twitter.com,vertex_project) :webpage=https://vertex.link/]'
print(q1)
q2 = '.created'
# Display the syntax
# Run the query and test
podes = await core.eval(q, num=0, cmdr=False)
podes = await core.eval(q1, num=1, cmdr=False)
podes = await core.eval(q2, num=7, cmdr=True)
# + active=""
# .. _node-add:
#
# Add Nodes
# ---------
#
# Operation to add the specified node(s) to a Cortex.
#
# **Syntax:**
#
# **[** *<form>* **=** | **?=** *<valu>* ... **]**
#
# **Examples:**
#
# *Create a simple node:*
# + hideCode=true
# Make some nodes
q = '[ inet:fqdn = woot.com ]'
# Display the syntax
print(q)
# Run the query and test
podes = await core.eval(q, num=1, cmdr=False)
# + active=""
# *Create a composite (comp) node:*
# + hideCode=true
# Make some nodes
q = '[ inet:dns:a=(woot.com, 172.16.58.3) ]'
# Display the syntax
print(q)
# Run the query and test
podes = await core.eval(q, num=1, cmdr=False)
# + active=""
# *Create a GUID node:*
# + hideCode=true
# Make some nodes
guid = '2f92bc913918f6598bcf310972ebf32e'
q = f'[ ou:org={guid} ]'
# Display the syntax
print(q)
# Run the query and test
podes = await core.eval(q, num=1, cmdr=False)
# + hideCode=true
# Make some nodes
q = '[ ou:org="*" ]'
# Display the syntax
print(q)
# Run the query and test
podes = await core.eval(q, num=1, cmdr=False)
assert podes[0][0] != ('ou:org', guid)
# + active=""
# *Create a digraph (edge) node:*
# + hideCode=true
# Make some nodes
q = '[ edge:refs=((media:news, 00a1f0d928e25729b9e86e2d08c127ce), (inet:fqdn, woot.com)) ]'
# Display the syntax
print(q)
# Run the query and test
podes = await core.eval(q, num=1, cmdr=False)
# + active=""
# *Create multiple nodes:*
# + hideCode=true
# Make some nodes
q = '[ inet:fqdn=woot.com inet:ipv4=12.34.56.78 hash:md5=d41d8cd98f00b204e9800998ecf8427e ]'
# Display the syntax
print(q)
# Run the query and test
podes = await core.eval(q, num=3, cmdr=False)
# + active=""
# **Usage Notes:**
#
# - Storm can create as many nodes as are specified within the brackets. It is not necessary to create only one node at a time.
# - For nodes specified within the brackets that do not already exist, Storm will create and return the node. For nodes that already exist, Storm will simply return that node.
# - When creating a *<form>* whose *<valu>* consists of multiple components, the components must be passed as a comma-separated list enclosed in parentheses.
# - Once a node is created, its primary property (*<form>* = *<valu>*) **cannot be modified.** The only way to "change" a node’s primary property is to create a new node (and optionally delete the old node). "Modifying" nodes therefore consists of adding, modifying, or deleting secondary properties (including universal properties) or adding or removing tags.
# + active=""
# .. _prop-add-mod:
#
# Add or Modify Properties
# ------------------------
#
# Operation to add (set) or change one or more properties on the specified node(s).
#
# The same syntax is used to apply a new property or modify an existing property.
#
# **Syntax:**
#
# *<query>* **[ :** *<prop>* **=** | **?=** *<pval>* ... **]**
#
# .. NOTE::
# Synapse supports secondary properties that are **arrays** (lists of typed forms), such as ``ou:org:names``. See the :ref:`type-array` section of the :ref:`storm-ref-type-specific` guide for slightly modified syntax used to add or modify array properties.
#
# **Examples:**
#
# *Add (or modify) secondary property:*
# + hideCode=true
# Use previous data, define and print test query
q = '<inet:ipv4> '
q1 = 'inet:ipv4=172.16.58.3 '
q2 = '[ :loc=us.oh.wilmington ]'
print(q + q2)
# Execute the query to test it and get the packed nodes (podes).
podes = await core.eval(q1 + q2, num=1, cmdr=False)
assert podes[0][1].get('props').get('loc') == 'us.oh.wilmington'
# + active=""
# *Add (or modify) universal property:*
# + hideCode=true
# Use previous data, define and print test query
q = '<inet:dns:a> '
q1 = 'inet:dns:a=(woot.com,12.34.56.78) '
q2 = '[ .seen=("2017/08/01 01:23", "2017/08/01 04:56") ]'
print(q + q2)
# Execute the query to test it and get the packed nodes (podes).
podes = await core.eval(q1 + q2, num=1, cmdr=False)
# + active=""
# *Add (or modify) a string property to a null value:*
# + hideCode=true
# Use previous data, define and print test query
q = '<media:news> '
q1 = 'media:news=00a1f0d928e25729b9e86e2d08c127ce '
q2 = '[ :summary="" ]'
print(q + q2)
# Execute the query to test it and get the packed nodes (podes).
podes = await core.eval(q1 + q2, num=1, cmdr=False)
assert podes[0][1].get('props').get('summary') == ''
# + active=""
# **Usage Notes:**
#
# - Additions or modifications to properties are performed on the output of a previous Storm query.
# - Storm will set or change the specified properties for all nodes in the current working set (i.e., all nodes resulting from Storm syntax to the left of the *<prop> = <pval>* statement(s)) for which that property is valid, **whether those nodes are within or outside of the brackets** unless :ref:`edit-parens` are used to limit the scope of the modifications.
# - Specifying a property will set the *<prop> = <pval>* if it does not exist, or modify (overwrite) the *<prop> = <pval>* if it already exists. **There is no prompt to confirm overwriting of an existing property.**
# - Storm will return an error if the inbound set of nodes contains any forms for which *<prop>* is not a valid property. For example, attempting to set a ``:loc`` property when the inbound nodes contain both domains and IP addresses will return an error as ``:loc`` is not a valid secondary property for a domain (``inet:fqdn``).
# - Secondary properties **must** be specified by their relative property name. For example, for the form ``foo:bar`` with the property ``baz`` (i.e., ``foo:bar:baz``) the relative property name is specified as ``:baz``.
# - Storm can set or modify any secondary property (including universal properties) except those explicitly defined as read-only (``'ro' : 1``) in the data model. Attempts to modify read only properties will return an error.
# + active=""
# .. _prop-del:
#
# Delete Properties
# -----------------
#
# Operation to delete (fully remove) one or more properties from the specified node(s).
#
# .. WARNING::
# Storm syntax to delete properties has the potential to be destructive if executed following an incorrect, badly formed, or mistyped query. Users are **strongly encouraged** to validate their query by first executing it on its own (without the delete property operation) to confirm it returns the expected nodes before adding the delete syntax. While the property deletion syntax cannot fully remove a node from the hypergraph, it is possible for a bad property deletion operation to irreversibly damage hypergraph pivoting and traversal.
#
# **Syntax:**
#
# *<query>* **[ -:** *<prop>* ... **]**
#
# **Examples:**
#
# *Delete a property:*
# + hideCode=true hideOutput=true
# Make a node
q = '[ inet:ipv4=172.16.58.3 :loc=nl :asn=60781 ]'
# Run the query and test
podes = await core.eval(q, num=1, cmdr=True)
assert podes[0][1].get('props').get('asn') == 60781
assert podes[0][1].get('props').get('loc') == 'nl'
# + hideCode=true
# Use previous data, define and print test query
q = '<inet:ipv4> '
q1 = 'inet:ipv4=172.16.58.3 '
q2 = '[ -:loc ]'
print(q + q2)
# Execute the query to test it and get the packed nodes (podes).
podes = await core.eval(q1 + q2, num=1, cmdr=False)
assert podes[0][1].get('props').get('loc') is None
# + active=""
# *Delete multiple properties:*
# + hideCode=true
# Use previous data, define and print test query
q = '<media:news> '
q1 = 'media:news=00a1f0d928e25729b9e86e2d08c127ce '
q2 = '[ -:author -:summary ]'
print(q + q2)
# Execute the query to test it and get the packed nodes (podes).
podes = await core.eval(q1 + q2, num=1, cmdr=False)
assert podes[0][1].get('props').get('author') is None
assert podes[0][1].get('props').get('summary') is None
# + active=""
# **Usage Notes:**
#
# - Property deletions are performed on the output of a previous Storm query.
# - Storm will delete the specified property / properties for all nodes in the current working set (i.e., all nodes resulting from Storm syntax to the left of the *-:<prop>* statement), **whether those nodes are within or outside of the brackets** unless :ref:`edit-parens` are used to limit the scope of the modifications.
# - Deleting a property fully removes the property from the node; it does not set the property to a null value.
# - Properties which are read-only ( ``'ro' : 1`` ) as specified in the data model cannot be deleted.
# + active=""
# .. _node-del:
#
# Delete Nodes
# ------------
#
# Nodes can be deleted from a Cortex using the Storm :ref:`storm-delnode` command.
# + active=""
# .. _light-edge-add:
#
# Add Light Edges
# ---------------
#
# Operation that links the specified node(s) to another node or set of nodes (as specified by a Storm expression) using a lightweight edge (light edge).
#
# See :ref:`light-edge` for details on light edges.
#
# **Syntax:**
#
# *<query>* **[ +(** *<verb>* **)> {** *<storm>* **} ]**
#
# *<query>* **[ <(** *<verb>* **)+ {** *<storm>* **} ]**
#
# .. NOTE::
# The nodes specified by the Storm expression ( ``{ <storm> }`` ) must either already exist in the Cortex or must be created as part of the Storm expression in order for the light edges to be created.
#
# .. NOTE::
# The query syntax used to create light edges will **yield the nodes that are inbound to the edit brackets** (that is, the nodes represented by *<query>*).
#
# **Examples:**
#
# *Link the specified FQDN and IPv4 to the media:news node referenced by the Storm expression using a "refs" light edge:*
# + hideCode=true
# Make some nodes
q = '[ media:news=(d41d8cd98f00b204e9800998ecf8427e,) inet:fqdn=woot.com inet:ipv4=1.2.3.4]'
# Make some light edges
q1 = "inet:fqdn=woot.com inet:ipv4=1.2.3.4 [ <(refs)+ { media:news=a3759709982377809f28fc0555a38193 } ]"
# Display the syntax
print(q1)
# Run the queries and test
podes = await core.eval(q, num=3, cmdr=False)
podes = await core.eval(q1, num=2, cmdr=False)
# + active=""
# *Link the specified media:news node to the set of indicators tagged APT1 (#aka.feye.thr.apt1) using a "refs" light edge:*
# + hideCode=true
# Make some nodes
q = '[ inet:fqdn=newsonet.net inet:fqdn=staycools.net +#aka.feye.thr.apt1 ]'
# Make some light edges
q1 = "media:news=a3759709982377809f28fc0555a38193 [ +(refs)> { +#aka.feye.thr.apt1 } ]"
# Display the syntax
print(q1)
# Run the queries and test
podes = await core.eval(q, num=2, cmdr=False)
podes = await core.eval(q1, num=1, cmdr=False)
# + active=""
# *Link the specified inet:cidr4 netblock to any IP address within that netblock that already exists in the Cortex (as referenced by the Storm expression) using a "hasip" light edge:*
# + hideCode=true
# Make some nodes
q = '[ inet:cidr4=192.168.127.12/24 inet:ipv4=172.16.58.3 inet:ipv4=172.16.17.32 inet:ipv4=192.168.127.12 ]'
# Make some light edges
q1 = "inet:cidr4=192.168.127.12/24 [ +(hasip)> { inet:ipv4=192.168.127.12/24 } ]"
# Display the syntax
print(q1)
# Run the queries and test
podes = await core.eval(q, num=4, cmdr=False)
podes = await core.eval(q1, num=1, cmdr=False)
# + active=""
# *Link the specified inet:cidr4 netblock to every IP in its range (as referenced by the Storm expression) using a "hasip" light edge, creating the IPs if they don't exist:*
# + hideCode=true
# Make some nodes
q = '[ inet:cidr4=192.168.127.12/24 inet:ipv4=172.16.58.3 inet:ipv4=172.16.17.32 inet:ipv4=192.168.127.12 ]'
# Make some light edges
q1 = "inet:cidr4=192.168.127.12/24 [ +(hasip)> { [ inet:ipv4=192.168.127.12/24 ] } ]"
# Display the syntax
print(q1)
# Run the queries and test
podes = await core.eval(q, num=4, cmdr=False)
podes = await core.eval(q1, num=1, cmdr=False)
# + active=""
# **Usage Notes:**
#
# - No light edge verbs exist in a Cortex by default; they must be created.
# - Light edge verbs are created at the user's discretion "on the fly" (i.e., when they are first used to link nodes); they do not need to be created manually before they can be used.
#
# - We recommend that users agree on a consistent set of light edge verbs and their meanings.
# - The Storm :ref:`storm-model` commands can be used to list and work with any light edge verbs in a Cortex.
#
# - A light edge's verb typically has a logical direction (a report "references" a set of indicators that it contains, but the indicators do not "reference" the report). However, it is up to the user to create the light edges in the correct direction and using forms that are sensical for the light edge verb. That is, there is nothing in the Storm syntax itself to prevent users linking any arbitrary nodes in arbitrary directions using arbitrary light edge verbs.
# - The plus sign ( ``+`` ) used with the light edge expression within the edit brackets is used to create the light edge(s).
# - Light edges can be created in either "direction" (e.g., with the directional arrow pointing either right ( ``+(<verb>)>`` ) or left ( ``<(<verb>)+`` ) - whichever syntax is easier.
# + active=""
# .. _light-edge-del:
#
# Delete Light Edges
# ------------------
#
# Operation that deletes the light edge linking the specified node(s) to the set of nodes specified by a given Storm expression.
#
# See :ref:`light-edge` for details on light edges.
#
# **Syntax:**
#
# *<query>* **[ -(** *<verb>* **)> {** *<storm>* **} ]**
#
# *<query>* **[ <(** *<verb>* **)- {** *<storm>* **} ]**
#
# .. CAUTION::
# The minus sign ( ``-`` ) used to reference a light edge **outside** of edit brackets simply instructs Storm to traverse ("walk") the specified light edge; for example, ``inet:cidr4=192.168.0.0/24 -(hasip)> inet:ipv4`` (see :ref:`walk-light-edge`). The minus sign used to reference a light edge **inside** of edit brackets instructs Storm to **delete** the specified edges (i.e., ``inet:cidr4=192.168.0.0/24 [ -(hasip)> { inet:ipv4=192.168.0.0/24 } ]``).
#
# **Examples:**
#
# *Delete the "refs" light edge linking the MD5 hash of the empty file to the specified media:news node:*
# + hideCode=true
# Make some light edges
q = "hash:md5=d41d8cd98f00b204e9800998ecf8427e [ <(refs)+ { media:news=a3759709982377809f28fc0555a38193 } ]"
# Delete some light edges
q1 = "hash:md5=d41d8cd98f00b204e9800998ecf8427e [ <(refs)- { media:news=a3759709982377809f28fc0555a38193 } ]"
# Display the syntax
print(q1)
# Run the query and test
podes = await core.eval(q, num=1, cmdr=False)
podes = await core.eval(q1, num=1, cmdr=False)
# + active=""
# *Delete the "hasip" light edge linking IP 1.2.3.4 to the specified CIDR block:*
# + hideCode=true
# Make some light edges
q = "inet:cidr4=192.168.127.12/24 [ +(hasip)> { inet:ipv4=1.2.3.4 } ]"
# Delete some light edges
q1 = "inet:cidr4=192.168.127.12/24 [ -(hasip)> { inet:ipv4=1.2.3.4 } ]"
# Display the syntax
print(q1)
# Run the queries and test
podes = await core.eval(q, num=1, cmdr=False)
podes = await core.eval(q1, num=1, cmdr=False)
# + active=""
# **Usage Notes:**
#
# - The minus sign ( ``-`` ) used with the light edge expression within the edit brackets is used to delete the light edge(s).
# - Light edges can be deleted in either "direction" (e.g., with the directional arrow pointiing either right ( ``-(<verb>)>`` ) or left ( ``<(<verb>)-`` ) - whichever syntax is easier.
# + active=""
# .. _tag-add:
#
# Add Tags
# --------
#
# Operation to add one or more tags to the specified node(s).
#
# **Syntax:**
#
# *<query>* **[ +#** *<tag>* ... **]**
#
# **Example:**
#
# *Add multiple tags:*
# + hideCode=true hideOutput=true
# Make a node
q = '[inet:fqdn=blackcake.net]'
# Run the query and test
podes = await core.eval(q, num=1, cmdr=True)
# + hideCode=true
# Use previous data, define and print test query
q = '<inet:fqdn> '
q1 = 'inet:fqdn=blackcake.net '
q2 = '[ +#aka.feye.thr.apt1 +#cno.infra.sink.hole ]'
print(q + q2)
# Execute the query to test it and get the packed nodes (podes).
podes = await core.eval(q1 + q2, num=1, cmdr=False)
assert 'aka.feye.thr.apt1' in podes[0][1].get('tags')
assert 'cno.infra.sink.hole' in podes[0][1].get('tags')
# + active=""
# **Usage Notes:**
#
# - Tag additions are performed on the output of a previous Storm query.
# - Storm will add the specified tag(s) to all nodes in the current working set (i.e., all nodes resulting from Storm syntax to the left of the *+#<tag>* statement) **whether those nodes are within or outside of the brackets** unless :ref:`edit-parens` are used to limit the scope of the modifications.
# + active=""
# .. _tag-prop-add:
#
# Add Tag Timestamps or Tag Properties
# ++++++++++++++++++++++++++++++++++++
#
# Synapse supports the use of :ref:`tag-timestamps` and :ref:`tag-properties` to provide additional context to tags where appopriate.
#
# **Syntax:**
#
# Add tag timestamps:
#
# *<query>* **[ +#** *<tag>* **=** *<time>* | **(** *<min_time>* **,** *<max_time>* **)** ... **]**
#
# Add tag property:
#
# *<query>* **[ +#** *<tag>* **:** *<tagprop>* **=** *<pval>* ... **]**
#
# **Examples:**
# + active=""
# *Add tag with single timestamp:*
# + hideCode=true hideOutput=true
# Make a node
q = '[inet:fqdn=aoldaily.com]'
# Run the query and test
podes = await core.eval(q, num=1, cmdr=True)
# + hideCode=true
# Use previous data, define and print test query
q = '<inet:fqdn> '
q1 = 'inet:fqdn=aoldaily.com '
q2 = '[ +#cno.infra.sink.hole=2018/11/27 ]'
print(q + q2)
# Execute the query to test it and get the packed nodes (podes).
podes = await core.eval(q1 + q2, num=1, cmdr=False)
# + active=""
# *Add tag with a time interval (min / max):*
# + hideCode=true
# Use previous data, define and print test query
q = '<inet:fqdn> '
q1 = 'inet:fqdn=blackcake.net '
q2 = '[ +#cno.infra.sink.hole=(2014/11/06, 2016/11/06) ]'
print(q + q2)
# Execute the query to test it and get the packed nodes (podes).
podes = await core.eval(q1 + q2, num=1, cmdr=False)
assert podes[0][1].get('tags').get('cno.infra.sink.hole') != (None, None)
# + active=""
# *Add tag with custom tag property:*
# + hideCode=true
# Create a custom tag property
await core.core.addTagProp('risk', ('int', {'min': 0, 'max': 100}), {'doc': 'Risk score'})
# + hideCode=true
# Use previous data, define and print test query
q = '<inet:fqdn> '
q1 = 'inet:fqdn=blackcake.net '
q2 = '[ +#rep.symantec:risk = 87 ]'
print(q + q2)
# Execute the query to test it and get the packed nodes (podes).
podes = await core.eval(q1 + q2, num=1, cmdr=False)
# + active=""
# **Usage Notes:**
#
# - :ref:`tag-timestamps` and :ref:`tag-properties` are applied only to the tags to which they are explicitly added. For example, adding a timestamp to the tag ``#foo.bar.baz`` does **not** add the timestamp to tags ``#foo.bar`` and ``#foo``.
# - Tag timestamps are interval (``ival``) types and exhibit behavior specific to that type. See the :ref:`type-ival` section of the :ref:`storm-ref-type-specific` document for additional detail on working with interval types.
# + active=""
# .. _tag-mod:
#
# Modify Tags
# -----------
#
# Tags are "binary" in that they are either applied to a node or they are not. Tag names cannot be changed once set.
#
# To "change" the tag applied to a node, you must add the new tag and delete the old one.
#
# The Storm :ref:`storm-movetag` command can be used to modify tags in bulk - that is, rename an entire set of tags, or move a tag to a different tag tree.
# + active=""
# .. _tag-prop-mod:
#
# Modify Tag Timestamps or Tag Properties
# +++++++++++++++++++++++++++++++++++++++
#
# Tag timestamps or tag properties can be modified using the same syntax used to add the timestamp or property.
#
# Modifications are constrained by the :ref:`data-type` of the timestamp (i.e., :ref:`type-ival`) or property. For example:
#
# - modifying an existing custom property of type integer (``int``) will simply overwrite the old tag property value with the new one.
# - modifying an existing timestamp will only change the timestamp if the new minimum is smaller than the current minimum and / or the new maximum is larger than the current maximum, in accordance with type-specific behavior for intervals (``ival``).
#
# See :ref:`storm-ref-type-specific` for details.
# + active=""
# .. _tag-del:
#
# Remove Tags
# -----------
#
# Operation to delete one or more tags from the specified node(s).
#
# Removing a tag from a node differs from deleting the node representing a tag (a ``syn:tag`` node), which can be done using the Storm :ref:`storm-delnode` command.
#
# .. WARNING::
# Storm syntax to remove tags has the potential to be destructive if executed on an incorrect, badly formed, or mistyped query. Users are **strongly encouraged** to validate their query by first executing it on its own to confirm it returns the expected nodes before adding the tag deletion syntax.
#
# In addition, it is **essential** to understand how removing a tag at a given position in a tag tree affects other tags within that tree. Otherwise, tags may be improperly left in place ("orphaned") or inadvertently removed.
#
# **Syntax:**
#
# *<query>* **[ -#** *<tag>* ... **]**
#
# **Examples:**
#
# *Remove a leaf tag:*
# + hideCode=true hideOutput=true
# Make a node
q = '[inet:ipv4=172.16.58.3 +#cno.infra.anon.tor]'
# Run the query and test
podes = await core.eval(q, num=1, cmdr=True)
# + hideCode=true
# Define and print test query
q = '<inet:ipv4> '
q1 = 'inet:ipv4=172.16.58.3 '
q2 = '[ -#cno.infra.anon.tor ]'
print(q + q2)
# Execute the query to test it and get the packed nodes (podes).
podes = await core.eval(q1 + q2, num=1, cmdr=False)
assert 'cno.infra.anon.tor' not in podes[0][1].get('tags')
# + active=""
# **Usage Notes:**
#
# - Tag deletions are performed on the output of a previous Storm query.
# - Storm will delete the specified tag(s) from all nodes in the current working set (i.e., all nodes resulting from Storm syntax to the left of the -#<tag> statement), **whether those nodes are within or outside of the brackets** unless :ref:`edit-parens` are used to limit the scope of the modifications.
# - Deleting a leaf tag deletes **only** the leaf tag from the node. For example, ``[ -#foo.bar.baz ]`` will delete the tag ``#foo.bar.baz`` but leave the tags ``#foo.bar`` and ``#foo`` on the node.
# - Deleting a non-leaf tag deletes that tag and **all tags below it in the tag hierarchy** from the node. For example, ``[ -#foo ]`` used on a node with tags ``#foo.bar.baz`` and ``#foo.hurr.derp`` will remove **all** of the following tags:
#
# - ``#foo.bar.baz``
# - ``#foo.hurr.derp``
# - ``#foo.bar``
# - ``#foo.hurr``
# - ``#foo``
# + active=""
# .. _tag-prop-del:
#
# Remove Tag Timestamps or Tag Properties
# +++++++++++++++++++++++++++++++++++++++
#
# Currently, it is not possible to remove a tag timestamp or tag property from a tag once it has been applied. Instead, the entire tag must be removed and re-added without the timestamp or property.
# + active=""
# .. _data-mod-combo:
#
# Combining Data Modification Operations
# --------------------------------------
#
# The square brackets representing edit mode are used for a wide range of operations, meaning it is possible to combine operations within a single set of brackets.
#
# Simple Examples
# +++++++++++++++
#
# *Create a node and add secondary properties:*
# + hideCode=true
# Make some nodes
q = '[ inet:ipv4=172.16.58.3 :loc=nl :asn=60781 ]'
# Display the syntax
print(q)
# Run the query and test
podes = await core.eval(q, num=1, cmdr=False)
assert podes[0][1].get('props').get('loc') == 'nl'
assert podes[0][1].get('props').get('asn') == 60781
# + active=""
# *Create a node and add a tag:*
# + hideCode=true
# Make some nodes
q = '[ inet:fqdn=blackcake.net +#aka.feye.thr.apt1 ]'
# Display the syntax
print(q)
# Run the query and test
podes = await core.eval(q, num=1, cmdr=False)
assert 'aka.feye.thr.apt1' in podes[0][1].get('tags')
# + hideCode=true hideOutput=true
# Remove some nodes
q = 'inet:fqdn=blackcake.net | delnode'
# Run the query and test
podes = await core.eval(q, num=0, cmdr=True)
# + active=""
# Edit Brackets and Edit Parentheses Examples
# +++++++++++++++++++++++++++++++++++++++++++
#
# The following examples illustrate the differences in Storm behavior when using :ref:`edit-brackets` alone vs. with :ref:`edit-parens`.
#
# When performing simple edit operations (i.e., Storm queries that add / modify a single node, or apply a tag to the nodes retrieved by a Storm lift operation) users can typically use only edit brackets and not worry about delimiting edit operations within additional edit parens.
#
# That said, edit parens may be necessary when creating and modifying multiple nodes in a single query, or performing edits within a longer or more complex Storm query. In these cases, understanding the difference between edit brackets' "operate on everything inbound" vs. edit parens' "limit modifications to the specified nodes" is critical to avoid unintended data modifications.
#
# **Example 1:**
#
# Consider the following Storm query that uses only edit brackets:
# + hideCode=true
# Define and print test query
q = 'inet:fqdn#aka.feye.thr.apt1 [ inet:fqdn=somedomain.com +#aka.eset.thr.sednit ]'
print(q)
# Execute the query and test
podes = await core.eval(q, cmdr=False)
# + hideCode=true hideOutput=true
# Make some nodes
q = '[inet:fqdn=hugesoft.org inet:fqdn=purpledaily.com +#aka.feye.thr.apt1]'
# Run the query and test
podes = await core.eval(q, num=2, cmdr=True)
# + active=""
# The query will perform the following:
#
# - Lift all domains that FireEye associates with APT1 (i.e., tagged ``#aka.feye.thr.apt1``).
# - Create the new domain ``somedomain.com`` (if it does not already exist) or lift it (if it does).
# - Apply the tag ``#aka.eset.thr.sednit`` to the domain ``somedomain.com`` **and** to all of the domains tagged ``#aka.feye.thr.apt1``.
#
# We can see the effects in the output of our example query:
# + hideCode=true
# Define and print test query
q = 'inet:fqdn#aka.feye.thr.apt1 [ inet:fqdn=somedomain.com +#aka.eset.thr.sednit ]'
# Execute the query and test
podes = await core.eval(q, cmdr=True)
# + hideCode=true hideOutput=true
# Remove some tags for our next example
q = 'inet:fqdn#aka.feye.thr.apt1 [-#aka.eset]'
# Run the query and test
podes = await core.eval(q, num=4, cmdr=True)
# + active=""
# Consider the same query using edit parens inside the brackets:
# + hideCode=true hideOutput=false
# Define and print test query
q = 'inet:fqdn#aka.feye.thr.apt1 [(inet:fqdn=somedomain.com +#aka.eset.thr.sednit)]'
print(q)
# Execute the query and test
podes = await core.eval(q, cmdr=False)
# + active=""
# Because we used the edit parens, the query will now perform the following:
#
# - Lift all domains that FireEye associates with APT1 (i.e., tagged ``#aka.feye.thr.apt1``).
# - Create the new domain ``somedomain.com`` (if it does not already exist) or lift it (if it does).
# - Apply the tag ``aka.eset.thr.sednit`` **only** to the domain ``somedomain.com``.
#
# We can see the difference in the output of the example query:
# + hideCode=true
# Define and print test query
q = 'inet:fqdn#aka.feye.thr.apt1 [(inet:fqdn=somedomain.com +#aka.eset.thr.sednit)]'
# Execute the query and test
podes = await core.eval(q, cmdr=True)
# + active=""
# **Example 2:**
#
# Consider the following Storm query that uses only edit brackets:
# + hideCode=true
# Define and print test query
q = '[inet:ipv4=1.2.3.4 :asn=1111 inet:ipv4=5.6.7.8 :asn=2222]'
print(q)
# Execute the query and test
podes = await core.eval(q, cmdr=False)
# + active=""
# The query will perform the following:
#
# - Create (or lift) the IP address ``1.2.3.4``.
# - Set the IP's ``:asn`` property to ``1111``.
# - Create (or lift) the IP address ``5.6.7.8``.
# - Set the ``:asn`` property for **both** IP addresses to ``2222``.
#
# We can see the effects in the output of our example query:
# + hideCode=true
# Define and print test query
q = '[inet:ipv4=1.2.3.4 :asn=1111 inet:ipv4=5.6.7.8 :asn=2222]'
# Execute the query and test
podes = await core.eval(q, cmdr=True)
# + hideCode=true hideOutput=true
# Delete some nodes for our next example
q = 'inet:ipv4=1.2.3.4 inet:ipv4=5.6.7.8 | delnode'
# Run the query and test
podes = await core.eval(q, num=0, cmdr=True)
# + active=""
# Consider the same query using edit parens inside the brackets:
# + hideCode=true
# Define and print test query
q = '[ (inet:ipv4=1.2.3.4 :asn=1111) (inet:ipv4=5.6.7.8 :asn=2222) ]'
print(q)
# Execute the query and test
podes = await core.eval(q, cmdr=False)
# + active=""
# Because the brackets separate the two sets of modifications, IP ``1.2.3.4`` has its ``:asn`` property set to ``1111`` while IP ``5.6.7.8`` has its ``:asn`` property set to ``2222``:
# + hideCode=true
# Define and print test query
q = '[ (inet:ipv4=1.2.3.4 :asn=1111) (inet:ipv4=5.6.7.8 :asn=2222) ]'
# Execute the query and test
podes = await core.eval(q, cmdr=True)
# + hideCode=true hideOutput=true
# Close cortex because done
await core.fini()
| docs/synapse/userguides/storm_ref_data_mod.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from PIL import Image
import random
import os
# +
rock = Image.open('rock.jpeg')
rock = rock.resize((200,200))
paper = Image.open('paper.jpg')
paper = paper.resize((200,200))
scissor = Image.open('scissors.jpg')
scissor = scissor.resize((200,200))
cards = ['Rock','Paper','Scissor']
rank = {'Rock': rock, 'Paper':paper , 'Scissor':scissor}
# -
class Player:
def __init__(self,name):
self.name = name
self.cards = cards
def shuffle(self):
random.shuffle(self.cards)
def choose(self):
self.card = self.cards[random.randint(0,2)]
self.image = rank[self.card]
return (self.card,self.image)
# +
def win_check(data_1,data_2):
if data_1[0] == data_2[0]:
print('Player_1 {0}'.format(player1.name))
data_1[1].show()
print('Player_2 {0}'.format(player2.name))
data_2[1].show()
print('Tie\n')
return 'Tie'
#--------------------------------------------------------------
elif data_1[0] =='Rock' and data_2[0] =='Scissor':
print('Player_1 {0}'.format(player1.name))
data_1[1].show()
print('Player_2 {0}'.format(player2.name))
data_2[1].show()
print('{} won this Round '.format(player1.name))
print('\n')
return 'Player_1'
elif data_1[0] =='Rock' and data_2[0] =='Paper':
print('Player_1 {0}'.format(player1.name))
data_1[1].show()
print('Player_2 {0}'.format(player2.name))
data_2[1].show()
print('{} won this Round '.format(player2.name))
print('\n')
return 'Player_2'
#--------------------------------------------------------------
elif data_1[0] =='Scissor' and data_2[0] =='Rock':
print('Player_1 {0}'.format(player1.name))
data_1[1].show()
print('Player_2 {0}'.format(player2.name))
data_2[1].show()
print('{} won this Round '.format(player2.name))
print('\n')
return 'Player_2'
elif data_1[0] =='Scissor' and data_2[0] =='Paper':
print('Player_1 {0}'.format(player1.name))
data_1[1].show()
print('Player_2 {0}'.format(player2.name))
data_2[1].show()
print('{} won this Round '.format(player1.name))
print('\n')
return 'Player_1'
#--------------------------------------------------------------
elif data_1[0] =='Paper' and data_2[0] =='Scissor':
print('Player_1 {0}'.format(player1.name))
data_1[1].show()
print('Player_2 {0}'.format(player2.name))
data_2[1].show()
print('{} won this Round '.format(player2.name))
print('\n')
return 'Player_2'
elif data_1[0] =='Paper' and data_2[0] =='Rock':
print('Player_1 {0}'.format(player1.name))
data_1[1].show()
print('Player_2 {0}'.format(player2.name))
data_2[1].show()
print('{} won this Round '.format(player1.name))
print('\n')
return 'Player_1'
# +
# game one
game_on = True
ch = True
count1 = 0
count2 = 0
while ch:
try:
rounds = int(input("Enter number rounds !"))
ch = False
except:
print(" Enter integer only !")
while game_on:
ask1 = input("Enter Player_1's name : ")
player1 = Player(ask1)
ask2 = input("Enter Player_2's name : ")
player2 = Player(ask2)
# shuffle and choose
player1.shuffle()
player2.shuffle()
while rounds !=0:
# choose cards
data1 = player1.choose()
data2 = player1.choose()
ans = win_check(data1,data2)
if ans == 'Player_1':
count1 +=1
elif ans == 'Player_2':
count2 +=1
else:
print(" --------NO-------POINTS-------")
rounds -=1
if count1 > count2:
print("Player_1 {} won the game !".format(player1.name))
elif count2> count1:
print("Player_2 {} won the game !".format(player2.name))
else:
print("No one won Its a Tie")
ask = input("Do you want to play again !")
if ask[0].upper() =='Y':
ch = True
while ch:
try:
rounds = int(input("Enter number rounds !"))
ch = False
except:
print(" Enter integer only !")
continue
else:
game_on = False
# -
# +
# -
| Rock_Paper_Scissors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from __future__ import print_function
from imp import reload
# ## UAT for NbAgg backend.
#
# The first line simply reloads matplotlib, uses the nbagg backend and then reloads the backend, just to ensure we have the latest modification to the backend code. Note: The underlying JavaScript will not be updated by this process, so a refresh of the browser after clearing the output and saving is necessary to clear everything fully.
# +
import matplotlib
reload(matplotlib)
matplotlib.use('nbagg')
import matplotlib.backends.backend_nbagg
reload(matplotlib.backends.backend_nbagg)
# -
# ### UAT 1 - Simple figure creation using pyplot
#
# Should produce a figure window which is interactive with the pan and zoom buttons. (Do not press the close button, but any others may be used).
# +
import matplotlib.backends.backend_webagg_core
reload(matplotlib.backends.backend_webagg_core)
import matplotlib.pyplot as plt
plt.interactive(False)
fig1 = plt.figure()
plt.plot(range(10))
plt.show()
# -
# ### UAT 2 - Creation of another figure, without the need to do plt.figure.
#
# As above, a new figure should be created.
plt.plot([3, 2, 1])
plt.show()
# ### UAT 3 - Connection info
#
# The printout should show that there are two figures which have active CommSockets, and no figures pending show.
print(matplotlib.backends.backend_nbagg.connection_info())
# ### UAT 4 - Closing figures
#
# Closing a specific figure instance should turn the figure into a plain image - the UI should have been removed. In this case, scroll back to the first figure and assert this is the case.
plt.close(fig1)
# ### UAT 5 - No show without plt.show in non-interactive mode
#
# Simply doing a plt.plot should not show a new figure, nor indeed update an existing one (easily verified in UAT 6).
# The output should simply be a list of Line2D instances.
plt.plot(range(10))
# ### UAT 6 - Connection information
#
# We just created a new figure, but didn't show it. Connection info should no longer have "Figure 1" (as we closed it in UAT 4) and should have figure 2 and 3, with Figure 3 without any connections. There should be 1 figure pending.
print(matplotlib.backends.backend_nbagg.connection_info())
# ### UAT 7 - Show of previously created figure
#
# We should be able to show a figure we've previously created. The following should produce two figure windows.
plt.show()
plt.figure()
plt.plot(range(5))
plt.show()
# ### UAT 8 - Interactive mode
#
# In interactive mode, creating a line should result in a figure being shown.
plt.interactive(True)
plt.figure()
plt.plot([3, 2, 1])
# Subsequent lines should be added to the existing figure, rather than creating a new one.
plt.plot(range(3))
# Calling connection_info in interactive mode should not show any pending figures.
print(matplotlib.backends.backend_nbagg.connection_info())
# Disable interactive mode again.
plt.interactive(False)
# ### UAT 9 - Multiple shows
#
# Unlike most of the other matplotlib backends, we may want to see a figure multiple times (with or without synchronisation between the views, though the former is not yet implemented). Assert that plt.gcf().canvas.manager.reshow() results in another figure window which is synchronised upon pan & zoom.
plt.gcf().canvas.manager.reshow()
# ### UAT 10 - Saving notebook
#
# Saving the notebook (with CTRL+S or File->Save) should result in the saved notebook having static versions of the figues embedded within. The image should be the last update from user interaction and interactive plotting. (check by converting with ``ipython nbconvert <notebook>``)
# ### UAT 11 - Creation of a new figure on second show
#
# Create a figure, show it, then create a new axes and show it. The result should be a new figure.
#
# **BUG: Sometimes this doesn't work - not sure why (@pelson).**
# +
fig = plt.figure()
plt.axes()
plt.show()
plt.plot([1, 2, 3])
plt.show()
# -
# ### UAT 12 - OO interface
#
# Should produce a new figure and plot it.
# +
from matplotlib.backends.backend_nbagg import new_figure_manager,show
manager = new_figure_manager(1000)
fig = manager.canvas.figure
ax = fig.add_subplot(1,1,1)
ax.plot([1,2,3])
fig.show()
# -
# ## UAT 13 - Animation
#
# The following should generate an animated line:
# +
import matplotlib.animation as animation
import numpy as np
fig, ax = plt.subplots()
x = np.arange(0, 2*np.pi, 0.01) # x-array
line, = ax.plot(x, np.sin(x))
def animate(i):
line.set_ydata(np.sin(x+i/10.0)) # update the data
return line,
#Init only required for blitting to give a clean slate.
def init():
line.set_ydata(np.ma.array(x, mask=True))
return line,
ani = animation.FuncAnimation(fig, animate, np.arange(1, 200), init_func=init,
interval=32., blit=True)
plt.show()
# -
# ### UAT 14 - Keyboard shortcuts in IPython after close of figure
#
# After closing the previous figure (with the close button above the figure) the IPython keyboard shortcuts should still function.
#
# ### UAT 15 - Figure face colours
#
# The nbagg honours all colours appart from that of the figure.patch. The two plots below should produce a figure with a transparent background and a red background respectively (check the transparency by closing the figure, and dragging the resulting image over other content). There should be no yellow figure.
# +
import matplotlib
matplotlib.rcParams.update({'figure.facecolor': 'red',
'savefig.facecolor': 'yellow'})
plt.figure()
plt.plot([3, 2, 1])
with matplotlib.rc_context({'nbagg.transparent': False}):
plt.figure()
plt.plot([3, 2, 1])
plt.show()
# -
# ### UAT 16 - Events
#
# Pressing any keyboard key or mouse button (or scrolling) should cycle the line line while the figure has focus. The figure should have focus by default when it is created and re-gain it by clicking on the canvas. Clicking anywhere outside of the figure should release focus, but moving the mouse out of the figure should not release focus.
# +
import itertools
fig, ax = plt.subplots()
x = np.linspace(0,10,10000)
y = np.sin(x)
ln, = ax.plot(x,y)
evt = []
colors = iter(itertools.cycle(['r', 'g', 'b', 'k', 'c']))
def on_event(event):
if event.name.startswith('key'):
fig.suptitle('%s: %s' % (event.name, event.key))
elif event.name == 'scroll_event':
fig.suptitle('%s: %s' % (event.name, event.step))
else:
fig.suptitle('%s: %s' % (event.name, event.button))
evt.append(event)
ln.set_color(next(colors))
fig.canvas.draw()
fig.canvas.draw_idle()
fig.canvas.mpl_connect('button_press_event', on_event)
fig.canvas.mpl_connect('button_release_event', on_event)
fig.canvas.mpl_connect('scroll_event', on_event)
fig.canvas.mpl_connect('key_press_event', on_event)
fig.canvas.mpl_connect('key_release_event', on_event)
plt.show()
# -
# ### UAT 17 - Timers
#
# Single-shot timers follow a completely different code path in the nbagg backend than regular timers (such as those used in the animation example above.) The next set of tests ensures that both "regular" and "single-shot" timers work properly.
#
# The following should show a simple clock that updates twice a second:
# +
import time
fig, ax = plt.subplots()
text = ax.text(0.5, 0.5, '', ha='center')
def update(text):
text.set(text=time.ctime())
text.axes.figure.canvas.draw()
timer = fig.canvas.new_timer(500, [(update, [text], {})])
timer.start()
plt.show()
# -
# However, the following should only update once and then stop:
# +
fig, ax = plt.subplots()
text = ax.text(0.5, 0.5, '', ha='center')
timer = fig.canvas.new_timer(500, [(update, [text], {})])
timer.single_shot = True
timer.start()
plt.show()
# -
# And the next two examples should never show any visible text at all:
# +
fig, ax = plt.subplots()
text = ax.text(0.5, 0.5, '', ha='center')
timer = fig.canvas.new_timer(500, [(update, [text], {})])
timer.start()
timer.stop()
plt.show()
# +
fig, ax = plt.subplots()
text = ax.text(0.5, 0.5, '', ha='center')
timer = fig.canvas.new_timer(500, [(update, [text], {})])
timer.single_shot = True
timer.start()
timer.stop()
plt.show()
# -
# ### UAT17 - stoping figure when removed from DOM
#
# When the div that contains from the figure is removed from the DOM the figure should shut down it's comm, and if the python-side figure has no more active comms, it should destroy the figure. Repeatedly running the cell below should always have the same figure number
fig, ax = plt.subplots()
ax.plot(range(5))
plt.show()
# Running the cell below will re-show the figure. After this, re-running the cell above should result in a new figure number.
fig.canvas.manager.reshow()
| matplotlib/backends/web_backend/nbagg_uat.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import torch
import torch.nn as nn
from torch.autograd import Variable
a = np.load('/Users/dkorduban/workspace/sc2/my/replays/npz/a.npz')
batch = a['both']
print(batch.shape)
print(len(batch))
batch = torch.Tensor(batch)
print(batch.shape)
# +
def ortho_weights(shape, scale=1.):
""" PyTorch port of ortho_init from baselines.a2c.utils """
shape = tuple(shape)
if len(shape) == 2:
flat_shape = shape[1], shape[0]
elif len(shape) == 4:
flat_shape = (np.prod(shape[1:]), shape[0])
else:
raise NotImplementedError
a = np.random.normal(0., 1., flat_shape)
u, _, v = np.linalg.svd(a, full_matrices=False)
q = u if u.shape == flat_shape else v
q = q.transpose().copy().reshape(shape)
if len(shape) == 2:
return torch.from_numpy((scale * q).astype(np.float32))
if len(shape) == 4:
return torch.from_numpy((scale * q[:, :shape[1], :shape[2]]).astype(np.float32))
def atari_initializer(module):
""" Parameter initializer for Atari models
Initializes Linear, Conv2d, and LSTM weights.
"""
classname = module.__class__.__name__
if classname == 'Linear':
module.weight.data = ortho_weights(module.weight.data.size(), scale=np.sqrt(2.))
module.bias.data.zero_()
elif classname == 'Conv2d':
module.weight.data = ortho_weights(module.weight.data.size(), scale=np.sqrt(2.))
module.bias.data.zero_()
elif classname == 'LSTM':
for name, param in module.named_parameters():
if 'weight_ih' in name:
param.data = ortho_weights(param.data.size(), scale=1.)
if 'weight_hh' in name:
param.data = ortho_weights(param.data.size(), scale=1.)
if 'bias' in name:
param.data.zero_()
class ConvVAE(nn.Module):
def __init__(self, z_dim):
""" Basic convolutional variational autoencoder
"""
super().__init__()
self.encoder = nn.Sequential(
nn.Conv2d(6, 32, 4, stride=2),
nn.ReLU(inplace=True),
nn.Conv2d(32, 64, 4, stride=2),
nn.ReLU(inplace=True),
nn.Conv2d(64, 128, 4, stride=2),
nn.ReLU(inplace=True)
)
last_dim = 128 * 4
self.mu = nn.Linear(last_dim, z_dim)
self.logvar = nn.Linear(last_dim, z_dim)
self.
def forward(self, conv_in):
""" Module forward pass
Args:
conv_in (Variable): convolutional input, shaped [N x 4 x 84 x 84]
Returns:
pi (Variable): action probability logits, shaped [N x self.num_actions]
v (Variable): value predictions, shaped [N x 1]
"""
N = conv_in.size()[0]
conv_out = self.conv(conv_in).view(N, 64 * 7 * 7)
fc_out = self.fc(conv_out)
pi_out = self.pi(fc_out)
v_out = self.v(fc_out)
return pi_out, v_out
# +
# nn.Conv2d??
# -
import torch.nn.modules.rnn as rnn
# +
# nn.RNNBase??
# -
from torch.utils.data import Dataset,DataLoader
# +
# torch.utils.data.Dataset??
# +
class MySimpleDataset(Dataset):
def __init__(self, path):
a = np.load(path)
self._data = a['both']
def __getitem__(self, index):
return self._data[index]
def __len__(self):
return len(self._data)
dataset = MySimpleDataset('/Users/dkorduban/workspace/sc2/my/replays/npz/a.npz')
loader = DataLoader(dataset=dataset, batch_size=1, shuffle=True)
# -
a = np.array([1.,2])
# +
# a??
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
from lagom.core.networks import make_fc
from lagom.core.networks import make_cnn
from lagom.core.networks import make_transposed_cnn
from lagom.core.networks import ortho_init
from lagom.core.networks import BaseVAE
class ConvVAE(BaseVAE):
def make_encoder(self, config):
out = make_cnn(input_channel=1,
channels=[64, 64, 64],
kernels=[4, 4, 4],
strides=[2, 2, 1],
paddings=[0, 0, 0])
last_dim = 256
return out, last_dim
def make_moment_heads(self, config, last_dim):
out = {}
z_dim = config['network.z_dim']
out['mu_head'] = nn.Linear(in_features=last_dim, out_features=z_dim)
out['logvar_head'] = nn.Linear(in_features=last_dim, out_features=z_dim)
out['z_dim'] = z_dim
return out
def make_decoder(self, config, z_dim):
out = nn.ModuleList()
out.append(nn.Linear(in_features=z_dim, out_features=self.last_dim))
out.extend(make_transposed_cnn(input_channel=64,
channels=[64, 64, 64],
kernels=[4, 4, 4],
strides=[2, 1, 1],
paddings=[0, 0, 0],
output_paddings=[0, 0, 0]))
out.append(nn.Linear(in_features=9216, out_features=28*28*1))
return out
def init_params(self, config):
for layer in self.encoder:
ortho_init(layer, nonlinearity='relu', constant_bias=0.0)
ortho_init(self.mu_head, nonlinearity=None, weight_scale=0.01, constant_bias=0.0)
ortho_init(self.logvar_head, nonlinearity=None, weight_scale=0.01, constant_bias=0.0)
for layer in self.decoder:
ortho_init(layer, nonlinearity='relu', constant_bias=0.0)
def encoder_forward(self, x):
for layer in self.encoder:
x = F.relu(layer(x))
return x
def decoder_forward(self, z):
# Forward of first fully-connected layer
x = F.relu(self.decoder[0](z))
# Reshape as [NxCxHxW]
x = x.view(-1, 64, 2, 2)
# Forward pass through transposed convolutional layer
for layer in self.decoder[1:-1]:
x = F.relu(layer(x))
# Flatten to [N, D]
x = x.flatten(start_dim=1)
# Element-wise binary output
x = torch.sigmoid(self.decoder[-1](x))
return x
vae = ConvVAE(config={'network.z_dim': 32})
x = torch.randn(10, 1, 28, 28)
print(x.shape)
y = vae.encoder_forward(x)
print(y.shape)
y = y.flatten(start_dim=1)
print(y.shape)
# Forward pass through moment heads to obtain mu and logvar for latent variable
mu = vae.mu_head(y)
logvar = vae.logvar_head(y)
print(mu.shape, logvar.shape)
# Sample latent variable by using reparameterization trick
z = vae.reparameterize(mu, logvar)
print(z.shape)
# Forward pass through decoder of sampled latent variable to obtain reconstructed input
re_x = vae.decoder_forward(z)
print(re_x.shape)
# z = vae.reparameterize(y)
# print(z.shape)
# z = vae.decoder_forward(z)
# print(z.shape)
# -
'''
self.encoder = nn.Sequential(
nn.Conv2d(6, 32, 4, stride=2),
nn.ReLU(inplace=True),
nn.Conv2d(32, 64, 4, stride=2),
nn.ReLU(inplace=True),
nn.Conv2d(64, 128, 4, stride=2),
nn.ReLU(inplace=True)
)
last_dim = 128 * 4
self.mu = nn.Linear(last_dim, z_dim)
self.logvar = nn.Linear(last_dim, z_dim)
'''
x = torch.Tensor(batch)
print(x.shape)
x = nn.Conv2d(6, 32, 4, stride=2)(x)
print(x.shape)
x = nn.ReLU(inplace=True)(x)
print(x.shape)
x = nn.Conv2d(32, 64, 4, stride=2)(x)
print(x.shape)
x = nn.ReLU(inplace=True)(x)
print(x.shape)
x = nn.Conv2d(64, 128, 4, stride=2)(x)
print(x.shape)
x = nn.ReLU(inplace=True)(x)
print(x.shape)
x = x.view(-1, 128 * 2 * 2)
print(x.shape)
x = nn.Linear(512, 64)(x)
print(x.shape)
x = nn.Linear(64, 512)(x)
print(x.shape)
x = nn.ReLU(inplace=True)(x)
print(x.shape)
x = x.view(-1, 128, 2, 2)
print(x.shape)
x = nn.ConvTranspose2d(128, 64, 4, stride=2)(x)
print(x.shape)
x = nn.ReLU(inplace=True)(x)
print(x.shape)
x = nn.ConvTranspose2d(64, 32, 4, stride=2)(x)
print(x.shape)
x = nn.ReLU(inplace=True)(x)
print(x.shape)
x = nn.ConvTranspose2d(32, 6, 4, stride=2)(x)
print(x.shape)
x = nn.ReLU(inplace=True)(x)
print(x.shape)
# +
class Sc2ConvVAE(BaseVAE):
def make_encoder(self, config):
out = make_cnn(input_channel=6,
channels=[32, 64, 128],
kernels=[4, 4, 4],
strides=[2, 2, 2],
paddings=[0, 0, 0])
last_dim = 128 * 2 * 2
return out, last_dim
def make_moment_heads(self, config, last_dim):
out = {}
z_dim = config['network.z_dim']
out['mu_head'] = nn.Linear(in_features=last_dim, out_features=z_dim)
out['logvar_head'] = nn.Linear(in_features=last_dim, out_features=z_dim)
out['z_dim'] = z_dim
return out
def make_decoder(self, config, z_dim):
out = nn.ModuleList()
out.append(nn.Linear(in_features=z_dim, out_features=self.last_dim))
out.extend(make_transposed_cnn(input_channel=128,
channels=[64, 32, 6],
kernels=[4, 4, 4],
strides=[2, 2, 2],
paddings=[0, 0, 0],
output_paddings=[0, 0, 0]))
return out
def init_params(self, config):
for layer in self.encoder:
ortho_init(layer, nonlinearity='relu', constant_bias=0.0)
ortho_init(self.mu_head, nonlinearity=None, weight_scale=0.01, constant_bias=0.0)
ortho_init(self.logvar_head, nonlinearity=None, weight_scale=0.01, constant_bias=0.0)
for layer in self.decoder:
ortho_init(layer, nonlinearity='relu', constant_bias=0.0)
def encoder_forward(self, x):
for layer in self.encoder:
x = F.relu(layer(x))
return x
def decoder_forward(self, z):
# Forward of first fully-connected layer
x = F.relu(self.decoder[0](z))
# Reshape as [NxCxHxW]
x = x.view(-1, 128, 2, 2)
# Forward pass through transposed convolutional layer
for layer in self.decoder[1:]:
x = F.relu(layer(x))
# # Flatten to [N, D]
# x = x.flatten(start_dim=1)
# # Element-wise binary output
# x = torch.sigmoid(self.decoder[-1](x))
return x
vae = Sc2ConvVAE(config={'network.z_dim': 64})
x = torch.Tensor(batch)
print(x.shape)
re_x, _, _ = vae.forward(x)
print(re_x.shape)
re_x[0][3]
# +
from lagom import Logger
from lagom.engine import BaseEngine
import torch.optim as optim
from torchvision.utils import save_image
model = Sc2ConvVAE(config={'network.z_dim': 32})
optimizer = optim.Adam(model.parameters(), lr=1e-3)
model.train() # set to training mode
# Create a logger
train_output = Logger()
# Iterate over data batches for one epoch
for epoch in range(10):
print('EPOCH', epoch)
for i, data in enumerate(loader):
# Put data to device
# data = data.to(self.device)
# Zero-out gradient buffer
optimizer.zero_grad()
# Forward pass of data
data = data / 300.
re_x, mu, logvar = model(data)
# Calculate loss
out = model.calculate_loss(re_x=re_x, x=data, mu=mu, logvar=logvar, loss_type='MSE')
loss = out['loss']
# Backward pass to calcualte gradients
loss.backward()
# Take a gradient step
optimizer.step()
# Record train output
# train_output.log('epoch', n)
train_output.log('iteration', i)
train_output.log('train_loss', out['loss'].item()) # item() saves memory
train_output.log('reconstruction_loss', out['re_loss'].item())
train_output.log('KL_loss', out['KL_loss'].item())
# Dump logging
if i == 0 or (i+1) % 10 == 0:
print('-'*50)
train_output.dump(keys=None, index=-1, indent=0)
print('-'*50)
with torch.no_grad(): # fast, disable grad
z = torch.randn(16, 32)
re_x = model.decoder_forward(z)
re_x = re_x.view(16, 6, 30, 30) * 5
me, him = re_x.split(3, dim=1)
save_image(me, f'sample_me_{epoch}.png')
save_image(me, f'sample_him_{epoch}.png')
# train_output.logs
# def log_train(self, train_output, **kwargs):
# logdir = kwargs['logdir']
# epoch = kwargs['epoch']
# mean_loss = np.mean(train_output['train_loss'])
# print(f'====> Average loss: {mean_loss}')
# # Use decoder to sample images from standard Gaussian noise
# with torch.no_grad(): # fast, disable grad
# z = torch.randn(64, self.config['network.z_dim']).to(self.device)
# re_x = self.agent.decoder_forward(z).cpu()
# save_image(re_x.view(64, 1, 28, 28), f'{logdir}/sample_{epoch}.png')
# def eval(self, n=None):
# self.agent.eval() # set to evaluation mode
# # Create a logger
# eval_output = Logger()
# # Iterate over test batches
# for i, (data, label) in enumerate(self.test_loader):
# # Put data to device
# data = data.to(self.device)
# with torch.no_grad(): # fast, disable grad
# # Forward pass of data
# re_x, mu, logvar = self.agent(data)
# # Calculate loss
# out = self.agent.calculate_loss(re_x=re_x, x=data, mu=mu, logvar=logvar, loss_type='BCE')
# # Record eval output
# eval_output.log('eval_loss', out['loss'].item())
# return eval_output.logs
# def log_eval(self, eval_output, **kwargs):
# logdir = kwargs['logdir']
# epoch = kwargs['epoch']
# mean_loss = np.mean(eval_output['eval_loss'])
# print(f'====> Test set loss: {mean_loss}')
# # Reconstruct some test images
# data, label = next(iter(self.test_loader)) # get a random batch
# data = data.to(self.device)
# n = min(data.size(0), 8) # number of images
# D = data[:n]
# with torch.no_grad(): # fast, disable grad
# re_x, _, _ = self.agent(D)
# compare_img = torch.cat([D.cpu(), re_x.cpu().view(-1, 1, 28, 28)])
# save_image(compare_img, f'{logdir}/reconstruction_{epoch}.png', nrow=n)
# +
# optim.Adam??
# -
| experiments/pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # End-to-End FINN Flow for a Simple Convolutional Net
# -----------------------------------------------------------------
#
# In this notebook, we will go through the FINN steps needed to take a binarized convolutional network all the way down to a heterogeneous streaming dataflow accelerator running on the FPGA.
#
# It's recommended to go through the simpler [end-to-end notebook for a fully connected network](tfc_end2end_example.ipynb) first, since many steps here are very similar and we will focus on what is done differently for convolutions.
#
# This notebook is quite lengthy, and some of the cells (involving Vivado synthesis) may take up to an hour to finish running. To let you save and resume your progress, we will save the intermediate ONNX models that are generated in the various steps to disk, so that you can jump back directly to where you left off.
# ## Quick Introduction to the CNV-w1a1 Network
#
# The particular quantized neural network (QNN) we will be targeting in this notebook is referred to as CNV-w1a1 and it classifies 32x32 RGB images into one of ten CIFAR-10 classes. All weights and activations in this network are quantized to bipolar values (either -1 or +1), with the exception of the input (which is RGB with 8 bits per channel) and the final output (which is 32-bit numbers). It first appeared in the original [FINN paper](https://arxiv.org/abs/1612.07119) from ISFPGA'17 with the name CNV, as a variant of the binarized convolutional network from the [BinaryNet paper](https://arxiv.org/abs/1602.02830), in turn inspired by the VGG-11 topology which was the runner-up for the 2014 [ImageNet Large Scale Visual Recognition Challenge](http://www.image-net.org/challenges/LSVRC/).
#
#
# You'll have a chance to interactively examine the layers that make up the network in Netron in a moment, so that's enough about the network for now.
# ## Quick Recap of the End-to-End Flow
#
# The FINN compiler comes with many *transformations* that modify the ONNX representation of the network according to certain patterns. This notebook will demonstrate a *possible* sequence of such transformations to take a particular trained network all the way down to hardware, as shown in the figure below.
# 
# The white fields show the state of the network representation in the respective step. The colored fields represent the transformations that are applied to the network to achieve a certain result. The diagram is divided into 5 sections represented by a different color, each of it includes several flow steps. The flow starts in top left corner with Brevitas export (green section), followed by the preparation of the network (blue section) for the Vivado HLS synthesis and Vivado IPI stitching (orange section), and finally building a PYNQ overlay bitfile and testing it on a PYNQ board (yellow section).
# There is an additional section for functional verification (red section) on the left side of the diagram, which we will not cover in this notebook. For details please take a look in the verification notebook which you can find [here](tfc_end2end_verification.ipynb)
#
#
# We will use the helper function `showInNetron` to show the ONNX model at the current transformation step. The Netron displays are interactive, but they only work when running the notebook actively and not on GitHub (i.e. if you are viewing this on GitHub you'll only see blank squares).
# +
from finn.util.basic import make_build_dir
from finn.util.visualization import showInNetron
build_dir = "/workspace/finn"
# -
# ## 1. Brevitas Export, FINN Import and Tidy-Up
#
# Similar to what we did in the TFC-w1a1 end-to-end notebook, we will start by exporting the [pretrained CNV-w1a1 network](https://github.com/Xilinx/brevitas/tree/master/brevitas_examples/bnn_pynq) to ONNX, importing that into FINN and running the "tidy-up" transformations to have a first look at the topology.
# +
import onnx
from finn.util.test import get_test_model_trained
import brevitas.onnx as bo
from finn.core.modelwrapper import ModelWrapper
from finn.transformation.infer_shapes import InferShapes
from finn.transformation.fold_constants import FoldConstants
from finn.transformation.general import GiveReadableTensorNames, GiveUniqueNodeNames, RemoveStaticGraphInputs
cnv = get_test_model_trained("CNV", 1, 1)
bo.export_finn_onnx(cnv, (1, 3, 32, 32), build_dir + "/end2end_cnv_w1a1_export.onnx")
model = ModelWrapper(build_dir + "/end2end_cnv_w1a1_export.onnx")
model = model.transform(InferShapes())
model = model.transform(FoldConstants())
model = model.transform(GiveUniqueNodeNames())
model = model.transform(GiveReadableTensorNames())
model = model.transform(RemoveStaticGraphInputs())
model.save(build_dir + "/end2end_cnv_w1a1_tidy.onnx")
# -
# Now that the model is exported, let's have a look at its layer structure with Netron. Remember that the visualization below is interactive, you can click on the individual nodes and view the layer attributes, trained weights and so on.
showInNetron(build_dir+"/end2end_cnv_w1a1_tidy.onnx")
# You can see that the network is composed of a repeating convolution-convolution-maxpool layer pattern to extract features using 3x3 convolution kernels (with weights binarized) and `Sign` activations, followed by fully connected layers acting as the classifier. Also notice the initial `MultiThreshold` layer at the beginning of the network, which is quantizing float inputs to 8-bit ones.
# ### Adding Pre- and Postprocessing <a id='prepost'></a>
#
# TODO
# +
from finn.util.pytorch import ToTensor
from finn.transformation.merge_onnx_models import MergeONNXModels
from finn.core.datatype import DataType
model = ModelWrapper(build_dir+"/end2end_cnv_w1a1_tidy.onnx")
global_inp_name = model.graph.input[0].name
ishape = model.get_tensor_shape(global_inp_name)
# preprocessing: torchvision's ToTensor divides uint8 inputs by 255
totensor_pyt = ToTensor()
chkpt_preproc_name = build_dir+"/end2end_cnv_w1a1_preproc.onnx"
bo.export_finn_onnx(totensor_pyt, ishape, chkpt_preproc_name)
# join preprocessing and core model
pre_model = ModelWrapper(chkpt_preproc_name)
model = model.transform(MergeONNXModels(pre_model))
# add input quantization annotation: UINT8 for all BNN-PYNQ models
global_inp_name = model.graph.input[0].name
model.set_tensor_datatype(global_inp_name, DataType.UINT8)
# +
from finn.transformation.insert_topk import InsertTopK
from finn.transformation.infer_datatypes import InferDataTypes
# postprocessing: insert Top-1 node at the end
model = model.transform(InsertTopK(k=1))
chkpt_name = build_dir+"/end2end_cnv_w1a1_pre_post.onnx"
# tidy-up again
model = model.transform(InferShapes())
model = model.transform(FoldConstants())
model = model.transform(GiveUniqueNodeNames())
model = model.transform(GiveReadableTensorNames())
model = model.transform(InferDataTypes())
model = model.transform(RemoveStaticGraphInputs())
model.save(chkpt_name)
showInNetron(build_dir+"/end2end_cnv_w1a1_pre_post.onnx")
# -
# ## 2. How FINN Implements Convolutions: Lowering and Streamlining
#
# In FINN, we implement convolutions with the *lowering* approach: we convert them to matrix-matrix multiply operations, where one of the matrices is generated by sliding a window over the input image. You can read more about the sliding window operator and how convolution lowering works [in this notebook](https://github.com/maltanar/qnn-inference-examples/blob/master/3-convolutional-binarized-gtsrb.ipynb). The streaming dataflow architecture we will end up with is going to look something like this figure from the [FINN-R paper](https://arxiv.org/abs/1809.04570):
#
# 
#
# Note how the convolution layer looks very similar to the fully connected one in terms of the matrix-vector-threshold unit (MVTU), but now the MVTU is preceded by a sliding window unit that produces the matrix from the input image. All of these building blocks, including the `MaxPool` layer you see in this figure, exist as templated Vivado HLS C++ functions in [finn-hlslib](https://github.com/Xilinx/finn-hlslib).
#
#
# To target this kind of hardware architecture with our network we'll apply a convolution lowering transformation, in addition to streamlining. You may recall the *streamlining transformation* that we applied to the TFC-w1a1 network, which is a series of mathematical simplifications that allow us to get rid of floating point scaling operations by implementing few-bit activations as thresholding operations. **The current implementation of streamlining is highly network-specific and may not work for your network if its topology is very different than the example network here. We hope to rectify this in future releases.**
# +
from finn.transformation.streamline import Streamline
from finn.transformation.lower_convs_to_matmul import LowerConvsToMatMul
from finn.transformation.bipolar_to_xnor import ConvertBipolarMatMulToXnorPopcount
import finn.transformation.streamline.absorb as absorb
from finn.transformation.streamline.reorder import MakeMaxPoolNHWC, MoveScalarLinearPastInvariants
from finn.transformation.infer_data_layouts import InferDataLayouts
from finn.transformation.general import RemoveUnusedTensors
model = ModelWrapper(build_dir + "/end2end_cnv_w1a1_pre_post.onnx")
model = model.transform(MoveScalarLinearPastInvariants())
model = model.transform(Streamline())
model = model.transform(LowerConvsToMatMul())
model = model.transform(MakeMaxPoolNHWC())
model = model.transform(absorb.AbsorbTransposeIntoMultiThreshold())
model = model.transform(ConvertBipolarMatMulToXnorPopcount())
model = model.transform(Streamline())
# absorb final add-mul nodes into TopK
model = model.transform(absorb.AbsorbScalarMulAddIntoTopK())
model = model.transform(InferDataLayouts())
model = model.transform(RemoveUnusedTensors())
model.save(build_dir + "/end2end_cnv_w1a1_streamlined.onnx")
# -
# We won't go into too much detail about what happens in each transformation and why they are called in the particular order they are (feel free to visualize the intermediate steps using Netron yourself if you are curious) but here is a brief summmmary:
#
# * `Streamline` moves floating point scaling and addition operations closer to the input of the nearest thresholding activation and absorbs them into thresholds
# * `LowerConvsToMatMul` converts ONNX `Conv` nodes into sequences of `Im2Col, MatMul` nodes as discussed above. `Im2Col` is a custom FINN ONNX high-level node type that implements the sliding window operator.
# * `MakeMaxPoolNHWC` and `AbsorbTransposeIntoMultiThreshold` convert the *data layout* of the network into the NHWC data layout that finn-hlslib primitives use. NCHW means the tensor dimensions are ordered as `(N : batch, H : height, W : width, C : channels)` (assuming 2D images). The ONNX standard ops normally use the NCHW layout, but the ONNX intermediate representation itself does not dictate any data layout.
# * You may recall `ConvertBipolarMatMulToXnorPopcount` from the TFC-w1a1 example, which is needed to implement bipolar-by-bipolar (w1a1) networks correctly using finn-hlslib.
#
# Let's visualize the streamlined and lowered network with Netron. Observe how all the `Conv` nodes have turned into pairs of `Im2Col, MatMul` nodes, and many nodes including `BatchNorm, Mul, Add` nodes have disappeared and replaced with `MultiThreshold` nodes.
showInNetron(build_dir+"/end2end_cnv_w1a1_streamlined.onnx")
# ## 3. Partitioning, Conversion to HLS Layers and Folding
#
# The next steps will be (again) very similar to what we did for the TFC-w1a1 network. We'll first convert the layers that we can put into the FPGA into their HLS equivalents and separate them out into a *dataflow partition*:
#
# +
import finn.transformation.fpgadataflow.convert_to_hls_layers as to_hls
from finn.transformation.fpgadataflow.create_dataflow_partition import (
CreateDataflowPartition,
)
from finn.transformation.move_reshape import RemoveCNVtoFCFlatten
from finn.custom_op.registry import getCustomOp
from finn.transformation.infer_data_layouts import InferDataLayouts
# choose the memory mode for the MVTU units, decoupled or const
mem_mode = "decoupled"
model = ModelWrapper(build_dir + "/end2end_cnv_w1a1_streamlined.onnx")
model = model.transform(to_hls.InferBinaryStreamingFCLayer(mem_mode))
model = model.transform(to_hls.InferQuantizedStreamingFCLayer(mem_mode))
# TopK to LabelSelect
model = model.transform(to_hls.InferLabelSelectLayer())
# input quantization (if any) to standalone thresholding
model = model.transform(to_hls.InferThresholdingLayer())
model = model.transform(to_hls.InferConvInpGen())
model = model.transform(to_hls.InferStreamingMaxPool())
# get rid of Reshape(-1, 1) operation between hlslib nodes
model = model.transform(RemoveCNVtoFCFlatten())
# get rid of Tranpose -> Tranpose identity seq
model = model.transform(absorb.AbsorbConsecutiveTransposes())
# infer tensor data layouts
model = model.transform(InferDataLayouts())
parent_model = model.transform(CreateDataflowPartition())
parent_model.save(build_dir + "/end2end_cnv_w1a1_dataflow_parent.onnx")
sdp_node = parent_model.get_nodes_by_op_type("StreamingDataflowPartition")[0]
sdp_node = getCustomOp(sdp_node)
dataflow_model_filename = sdp_node.get_nodeattr("model")
# save the dataflow partition with a different name for easier access
dataflow_model = ModelWrapper(dataflow_model_filename)
dataflow_model.save(build_dir + "/end2end_cnv_w1a1_dataflow_model.onnx")
# -
# Notice the additional `RemoveCNVtoFCFlatten` transformation that was not used for TFC-w1a1. In the last Netron visualization you may have noticed a `Reshape` operation towards the end of the network where the convolutional part of the network ends and the fully-connected layers started. That `Reshape` is essentialy a tensor flattening operation, which we can remove for the purposes of hardware implementation. We can examine the contents of the dataflow partition with Netron, and observe the `ConvolutionInputGenerator`, `StreamingFCLayer_Batch` and `StreamingMaxPool_Batch` nodes that implement the sliding window, matrix multiply and maxpool operations in hlslib. *Note that the StreamingFCLayer instances following the ConvolutionInputGenerator nodes are really implementing the convolutions, despite the name. The final three StreamingFCLayer instances implement actual FC layers.*
showInNetron(build_dir + "/end2end_cnv_w1a1_dataflow_parent.onnx")
# Note that pretty much everything has gone into the `StreamingDataflowPartition` node; the only operation remaining is to apply a `Transpose` to obtain NHWC input from a NCHW input (the ONNX default).
showInNetron(build_dir + "/end2end_cnv_w1a1_dataflow_model.onnx")
# Now we have to set the *folding factors* for certain layers to adjust the performance of our accelerator, similar to the TFC-w1a1 example. We'll also set the desired FIFO depths around those layers, which are important to achieve full throughput in the accelerator.
# +
model = ModelWrapper(build_dir + "/end2end_cnv_w1a1_dataflow_model.onnx")
fc_layers = model.get_nodes_by_op_type("StreamingFCLayer_Batch")
# each tuple is (PE, SIMD, in_fifo_depth) for a layer
folding = [
(16, 3, 128),
(32, 32, 128),
(16, 32, 128),
(16, 32, 128),
(4, 32, 81),
(1, 32, 2),
(1, 4, 2),
(1, 8, 128),
(5, 1, 3),
]
for fcl, (pe, simd, ififodepth) in zip(fc_layers, folding):
fcl_inst = getCustomOp(fcl)
fcl_inst.set_nodeattr("PE", pe)
fcl_inst.set_nodeattr("SIMD", simd)
fcl_inst.set_nodeattr("inFIFODepth", ififodepth)
# use same SIMD values for the sliding window operators
swg_layers = model.get_nodes_by_op_type("ConvolutionInputGenerator")
for i in range(len(swg_layers)):
swg_inst = getCustomOp(swg_layers[i])
simd = folding[i][1]
swg_inst.set_nodeattr("SIMD", simd)
model = model.transform(GiveUniqueNodeNames())
model.save(build_dir + "/end2end_cnv_w1a1_folded.onnx")
# -
# Below we visualize in Netron to observe the `StreamingDataWidthConverter` and `StreamingFIFO` nodes that have been inserted into graph, as well as the folding factors in the `PE` and `SIMD` attributes of each `StreamingFCLayer_Batch`.
showInNetron(build_dir + "/end2end_cnv_w1a1_folded.onnx")
# Our network is now ready and we can start with the hardware generation.
# ## 4. Hardware Generation
#
# From this point onward, the steps we have to follow do not depend on the particular network and will be exactly the same as the TFC-w1a1 example. **which may take about 30 minutes depending on your host computer**. For more details about what's going on in this step, please consult the [TFC end-to-end notebook](tfc_end2end_example.ipynb) or the appropriate section in the [FINN documentation](https://finn.readthedocs.io/en/latest/hw_build.html).
# +
test_pynq_board = "Pynq-Z1"
target_clk_ns = 10
from finn.transformation.fpgadataflow.make_zynq_proj import ZynqBuild
model = ModelWrapper(build_dir+"/end2end_cnv_w1a1_folded.onnx")
model = model.transform(ZynqBuild(platform = test_pynq_board, period_ns = target_clk_ns))
model.save(build_dir + "/end2end_cnv_w1a1_synth.onnx")
# -
# ## 5. Deployment and Remote Execution
#
# Now that we're done with the hardware generation, we can generate a Python driver for accelerator and copy the necessary files onto our PYNQ board.
#
# **Make sure you've [set up the SSH keys for your PYNQ board](https://finn-dev.readthedocs.io/en/latest/getting_started.html#pynq-board-first-time-setup) before executing this step.**
# +
import os
# set up the following values according to your own environment
# FINN will use ssh to deploy and run the generated accelerator
ip = os.getenv("PYNQ_IP", "192.168.2.99")
username = os.getenv("PYNQ_USERNAME", "xilinx")
password = os.getenv("PYNQ_PASSWORD", "<PASSWORD>")
port = os.getenv("PYNQ_PORT", 22)
target_dir = os.getenv("PYNQ_TARGET_DIR", "/home/xilinx/finn_cnv_end2end_example")
# set up ssh options to only allow publickey authentication
options = "-o PreferredAuthentications=publickey -o PasswordAuthentication=no"
# test access to PYNQ board
# ! ssh {options} {username}@{ip} -p {port} cat /var/run/motd.dynamic
# +
from finn.transformation.fpgadataflow.make_deployment import DeployToPYNQ
model = ModelWrapper(build_dir + "/end2end_cnv_w1a1_synth.onnx")
model = model.transform(DeployToPYNQ(ip, port, username, password, target_dir))
model.save(build_dir + "/end2end_cnv_w1a1_pynq_deploy.onnx")
# -
target_dir_pynq = target_dir + "/" + model.get_metadata_prop("pynq_deployment_dir").split("/")[-1]
target_dir_pynq
# ! ssh {options} {username}@{ip} -p {port} 'ls -l {target_dir_pynq}'
# We only have two more steps to be able to remotely execute the deployed bitfile with some test data from the CIFAR-10 dataset. Let's load up some test data that comes bundled with FINN -- *and before you ask, that's supposed to be a cat (CIFAR-10 class number 3)*.
# +
import pkg_resources as pk
import matplotlib.pyplot as plt
import numpy as np
fn = pk.resource_filename("finn.qnn-data", "cifar10/cifar10-test-data-class3.npz")
x = np.load(fn)["arr_0"]
x = x.reshape(3, 32,32).transpose(1, 2, 0)
plt.imshow(x)
# -
# Recall that we partitioned our original network into a parent graph that contained the non-synthesizable nodes and a child graph that contained the bulk of the network, which we turned into a bitfile. The only operator left outside the FPGA partition was a `Transpose` to convert NCHW images into NHWC ones. Thus, we can skip the execution in the parent as long as we ensure our image has the expected data layout, which we have done above.
# +
import numpy as np
from finn.core.onnx_exec import execute_onnx
model = ModelWrapper(build_dir + "/end2end_cnv_w1a1_pynq_deploy.onnx")
iname = model.graph.input[0].name
oname = model.graph.output[0].name
ishape = model.get_tensor_shape(iname)
input_dict = {iname: x.astype(np.float32).reshape(ishape)}
ret = execute_onnx(model, input_dict, True)
# -
ret[oname]
# We see that the network correctly predicts this as a class 3 ("cat").
# ### Validating the Accuracy on a PYNQ Board <a id='validation'></a>
#
# All the command line prompts here are meant to be executed with `sudo` on the PYNQ board, so we'll use a workaround (`echo password | sudo -S command`) to get that working from this notebook running on the host computer.
#
# **Ensure that your PYNQ board has a working internet connecting for the next steps, since some there is some downloading involved.**
#
# To validate the accuracy, we first need to install the [`dataset-loading`](https://github.com/fbcotter/dataset_loading) Python package to the PYNQ board. This will give us a convenient way of downloading and accessing the MNIST dataset.
#
#
# Command to execute on PYNQ:
#
# ```pip3 install git+https://github.com/fbcotter/dataset_loading.git@0.0.4#egg=dataset_loading```
# ! ssh {options} -t {username}@{ip} -p {port} 'echo {password} | sudo -S pip3 install git+https://github.com/fbcotter/dataset_loading.git@0.0.4#egg=dataset_loading'
# We can now use the `validate.py` script that was generated together with the driver to measure top-1 accuracy on the CIFAR-10 dataset.
#
# Command to execute on PYNQ:
#
# `python3.6 validate.py --dataset cifar10 --batchsize 1000`
# ! ssh {options} -t {username}@{ip} -p {port} 'cd {target_dir_pynq}; echo {password} | sudo -S python3.6 validate.py --dataset cifar10 --batchsize 1000'
# We see that the final top-1 accuracy is 84.19%, which is very close to the 84.22% reported on the [BNN-PYNQ accuracy table in Brevitas](https://github.com/Xilinx/brevitas/tree/master/brevitas_examples/bnn_pynq).
| notebooks/end2end_example/bnn-pynq/cnv_end2end_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <a rel="license" href="http://creativecommons.org/licenses/by-nc-nd/4.0/"><img alt="Creative Commons License" style="border-width:0" src="https://i.creativecommons.org/l/by-nc-nd/4.0/88x31.png" /></a><br />This work is licensed under a <a rel="license" href="http://creativecommons.org/licenses/by-nc-nd/4.0/">Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License</a>.
# 
# # **Click-Through Rate Prediction Lab**
# This lab covers the steps for creating a click-through rate (CTR) prediction pipeline. You will work with the [Criteo Labs](http://labs.criteo.com/) dataset that was used for a recent [Kaggle competition](https://www.kaggle.com/c/criteo-display-ad-challenge).
#
# ** This lab will cover: **
#
# * *Part 1:* Featurize categorical data using one-hot-encoding (OHE)
#
# * *Part 2:* Construct an OHE dictionary
#
# * *Part 3:* Parse CTR data and generate OHE features
# * *Visualization 1:* Feature frequency
#
# * *Part 4:* CTR prediction and logloss evaluation
# * *Visualization 2:* ROC curve
#
# * *Part 5:* Reduce feature dimension via feature hashing
# * *Visualization 3:* Hyperparameter heat map
#
# > Note that, for reference, you can look up the details of:
# > * the relevant Spark methods in [Spark's Python API](https://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD)
# > * the relevant NumPy methods in the [NumPy Reference](http://docs.scipy.org/doc/numpy/reference/index.html)
labVersion = 'cs190.1x-lab4-1.0.4'
print labVersion
# #### ** Part 1: Featurize categorical data using one-hot-encoding **
# ** (1a) One-hot-encoding **
# We would like to develop code to convert categorical features to numerical ones, and to build intuition, we will work with a sample unlabeled dataset with three data points, with each data point representing an animal. The first feature indicates the type of animal (bear, cat, mouse); the second feature describes the animal's color (black, tabby); and the third (optional) feature describes what the animal eats (mouse, salmon).
#
# In a one-hot-encoding (OHE) scheme, we want to represent each tuple of `(featureID, category)` via its own binary feature. We can do this in Python by creating a dictionary that maps each tuple to a distinct integer, where the integer corresponds to a binary feature. To start, manually enter the entries in the OHE dictionary associated with the sample dataset by mapping the tuples to consecutive integers starting from zero, ordering the tuples first by featureID and next by category.
#
# Later in this lab, we'll use OHE dictionaries to transform data points into compact lists of features that can be used in machine learning algorithms.
# Data for manual OHE
# Note: the first data point does not include any value for the optional third feature
sampleOne = [(0, 'mouse'), (1, 'black')]
sampleTwo = [(0, 'cat'), (1, 'tabby'), (2, 'mouse')]
sampleThree = [(0, 'bear'), (1, 'black'), (2, 'salmon')]
sampleDataRDD = sc.parallelize([sampleOne, sampleTwo, sampleThree])
print sampleDataRDD.count()
# print sampleDataRDD.take(5)
# TODO: Replace <FILL IN> with appropriate code
sampleOHEDictManual = {}
sampleOHEDictManual[(0,'bear')] = 0
sampleOHEDictManual[(0,'cat')] = 1
sampleOHEDictManual[(0,'mouse')] = 2
sampleOHEDictManual[(1, 'black')] = 3
sampleOHEDictManual[(1, 'tabby')] = 4
sampleOHEDictManual[(2, 'mouse')] = 5
sampleOHEDictManual[(2, 'salmon')] = 6
print len(sampleOHEDictManual)
# **WARNING:** If *test_helper*, required in the cell below, is not installed, follow the instructions [here](https://databricks-staging-cloudfront.staging.cloud.databricks.com/public/c65da9a2fa40e45a2028cddebe45b54c/8637560089690848/4187311313936645/6977722904629137/05f3c2ecc3.html).
# +
# TEST One-hot-encoding (1a)
from test_helper import Test
Test.assertEqualsHashed(sampleOHEDictManual[(0,'bear')],
'b6589fc6ab0dc82cf12099d1c2d40ab994e8410c',
"incorrect value for sampleOHEDictManual[(0,'bear')]")
Test.assertEqualsHashed(sampleOHEDictManual[(0,'cat')],
'356a192b7913b04c54574d18c28d46e6395428ab',
"incorrect value for sampleOHEDictManual[(0,'cat')]")
Test.assertEqualsHashed(sampleOHEDictManual[(0,'mouse')],
'da4b9237bacccdf19c0760cab7aec4a8359010b0',
"incorrect value for sampleOHEDictManual[(0,'mouse')]")
Test.assertEqualsHashed(sampleOHEDictManual[(1,'black')],
'77de68daecd823babbb58edb1c8e14d7106e83bb',
"incorrect value for sampleOHEDictManual[(1,'black')]")
Test.assertEqualsHashed(sampleOHEDictManual[(1,'tabby')],
'1b6453892473a467d07372d45eb05abc2031647a',
"incorrect value for sampleOHEDictManual[(1,'tabby')]")
Test.assertEqualsHashed(sampleOHEDictManual[(2,'mouse')],
'ac3478d69a3c81fa62e60f5c3696165a4e5e6ac4',
"incorrect value for sampleOHEDictManual[(2,'mouse')]")
Test.assertEqualsHashed(sampleOHEDictManual[(2,'salmon')],
'c1dfd96eea8cc2b62785275bca38ac261256e278',
"incorrect value for sampleOHEDictManual[(2,'salmon')]")
Test.assertEquals(len(sampleOHEDictManual.keys()), 7,
'incorrect number of keys in sampleOHEDictManual')
# -
# ** (1b) Sparse vectors **
#
# Data points can typically be represented with a small number of non-zero OHE features relative to the total number of features that occur in the dataset. By leveraging this sparsity and using sparse vector representations of OHE data, we can reduce storage and computational burdens. Below are a few sample vectors represented as dense numpy arrays. Use [SparseVector](https://spark.apache.org/docs/latest/api/python/pyspark.mllib.html#pyspark.mllib.linalg.SparseVector) to represent them in a sparse fashion, and verify that both the sparse and dense representations yield the same results when computing [dot products](http://en.wikipedia.org/wiki/Dot_product) (we will later use MLlib to train classifiers via gradient descent, and MLlib will need to compute dot products between SparseVectors and dense parameter vectors).
#
# Use `SparseVector(size, *args)` to create a new sparse vector where size is the length of the vector and args is either a dictionary, a list of (index, value) pairs, or two separate arrays of indices and values (sorted by index). You'll need to create a sparse vector representation of each dense vector `aDense` and `bDense`.
import numpy as np
from pyspark.mllib.linalg import SparseVector
# +
# TODO: Replace <FILL IN> with appropriate code
aDense = np.array([0., 3., 0., 4.])
aSparse = SparseVector(len(aDense), range(0,len(aDense)), aDense)
bDense = np.array([0., 0., 0., 1.])
bSparse = SparseVector(len(bDense), range(0,len(bDense)), bDense)
w = np.array([0.4, 3.1, -1.4, -.5])
print aDense.dot(w)
print aSparse.dot(w)
print bDense.dot(w)
print bSparse.dot(w)
print aDense
print bDense
print aSparse
print bSparse
# -
# TEST Sparse Vectors (1b)
Test.assertTrue(isinstance(aSparse, SparseVector), 'aSparse needs to be an instance of SparseVector')
Test.assertTrue(isinstance(bSparse, SparseVector), 'aSparse needs to be an instance of SparseVector')
Test.assertTrue(aDense.dot(w) == aSparse.dot(w),
'dot product of aDense and w should equal dot product of aSparse and w')
Test.assertTrue(bDense.dot(w) == bSparse.dot(w),
'dot product of bDense and w should equal dot product of bSparse and w')
# **(1c) OHE features as sparse vectors **
#
# Now let's see how we can represent the OHE features for points in our sample dataset. Using the mapping defined by the OHE dictionary from Part (1a), manually define OHE features for the three sample data points using SparseVector format. Any feature that occurs in a point should have the value 1.0. For example, the `DenseVector` for a point with features 2 and 4 would be `[0.0, 0.0, 1.0, 0.0, 1.0, 0.0, 0.0]`.
# +
# Reminder of the sample features
# sampleOne = [(0, 'mouse'), (1, 'black')]
# sampleTwo = [(0, 'cat'), (1, 'tabby'), (2, 'mouse')]
# sampleThree = [(0, 'bear'), (1, 'black'), (2, 'salmon')]
# -
# TODO: Replace <FILL IN> with appropriate code
sampleOneOHEFeatManual = SparseVector(7, [2,3], np.array([1.0,1.0]))
sampleTwoOHEFeatManual = SparseVector(7, [1,4,5], np.array([1.0,1.0,1.0]))
sampleThreeOHEFeatManual = SparseVector(7, [0,3,6], np.array([1.0,1.0,1.0]))
print sampleOneOHEFeatManual
print sampleTwoOHEFeatManual
print sampleThreeOHEFeatManual
# TEST OHE Features as sparse vectors (1c)
Test.assertTrue(isinstance(sampleOneOHEFeatManual, SparseVector),
'sampleOneOHEFeatManual needs to be a SparseVector')
Test.assertTrue(isinstance(sampleTwoOHEFeatManual, SparseVector),
'sampleTwoOHEFeatManual needs to be a SparseVector')
Test.assertTrue(isinstance(sampleThreeOHEFeatManual, SparseVector),
'sampleThreeOHEFeatManual needs to be a SparseVector')
Test.assertEqualsHashed(sampleOneOHEFeatManual,
'ecc00223d141b7bd0913d52377cee2cf5783abd6',
'incorrect value for sampleOneOHEFeatManual')
Test.assertEqualsHashed(sampleTwoOHEFeatManual,
'26b023f4109e3b8ab32241938e2e9b9e9d62720a',
'incorrect value for sampleTwoOHEFeatManual')
Test.assertEqualsHashed(sampleThreeOHEFeatManual,
'c04134fd603ae115395b29dcabe9d0c66fbdc8a7',
'incorrect value for sampleThreeOHEFeatManual')
# **(1d) Define a OHE function **
#
# Next we will use the OHE dictionary from Part (1a) to programatically generate OHE features from the original categorical data. First write a function called `oneHotEncoding` that creates OHE feature vectors in `SparseVector` format. Then use this function to create OHE features for the first sample data point and verify that the result matches the result from Part (1c).
# TODO: Replace <FILL IN> with appropriate code
def oneHotEncoding_old(rawFeats, OHEDict, numOHEFeats):
"""Produce a one-hot-encoding from a list of features and an OHE dictionary.
Note:
You should ensure that the indices used to create a SparseVector are sorted.
Args:
rawFeats (list of (int, str)): The features corresponding to a single observation. Each
feature consists of a tuple of featureID and the feature's value. (e.g. sampleOne)
OHEDict (dict): A mapping of (featureID, value) to unique integer.
numOHEFeats (int): The total number of unique OHE features (combinations of featureID and
value).
Returns:
SparseVector: A SparseVector of length numOHEFeats with indices equal to the unique
identifiers for the (featureID, value) combinations that occur in the observation and
with values equal to 1.0.
"""
newFeats = []
idx = []
for k,i in sorted(OHEDict.items(), key=lambda x: x[1]):
if k in rawFeats:
newFeats += [1.0]
idx += [i]
return SparseVector(numOHEFeats, idx, np.array(newFeats))
# +
# TODO: Replace <FILL IN> with appropriate code
def oneHotEncoding(rawFeats, OHEDict, numOHEFeats):
"""Produce a one-hot-encoding from a list of features and an OHE dictionary.
Note:
You should ensure that the indices used to create a SparseVector are sorted.
Args:
rawFeats (list of (int, str)): The features corresponding to a single observation. Each
feature consists of a tuple of featureID and the feature's value. (e.g. sampleOne)
OHEDict (dict): A mapping of (featureID, value) to unique integer.
numOHEFeats (int): The total number of unique OHE features (combinations of featureID and
value).
Returns:
SparseVector: A SparseVector of length numOHEFeats with indices equal to the unique
identifiers for the (featureID, value) combinations that occur in the observation and
with values equal to 1.0.
"""
newFeats = []
idx = []
for f in rawFeats:
if f in OHEDict:
newFeats += [1.0]
idx += [OHEDict[f]]
return SparseVector(numOHEFeats, sorted(idx), np.array(newFeats))
# Calculate the number of features in sampleOHEDictManual
numSampleOHEFeats = len(sampleOHEDictManual)
# Run oneHotEnoding on sampleOne
sampleOneOHEFeat = oneHotEncoding(sampleOne,sampleOHEDictManual,numSampleOHEFeats)
print sampleOneOHEFeat
# -
# TEST Define an OHE Function (1d)
Test.assertTrue(sampleOneOHEFeat == sampleOneOHEFeatManual,
'sampleOneOHEFeat should equal sampleOneOHEFeatManual')
Test.assertEquals(sampleOneOHEFeat, SparseVector(7, [2,3], [1.0,1.0]),
'incorrect value for sampleOneOHEFeat')
Test.assertEquals(oneHotEncoding([(1, 'black'), (0, 'mouse')], sampleOHEDictManual,
numSampleOHEFeats), SparseVector(7, [2,3], [1.0,1.0]),
'incorrect definition for oneHotEncoding')
# **(1e) Apply OHE to a dataset **
#
# Finally, use the function from Part (1d) to create OHE features for all 3 data points in the sample dataset.
# +
# TODO: Replace <FILL IN> with appropriate code
def toOHE(row):
return oneHotEncoding(row,sampleOHEDictManual,numSampleOHEFeats)
sampleOHEData = sampleDataRDD.map(toOHE)
print sampleOHEData.collect()
# -
# TEST Apply OHE to a dataset (1e)
sampleOHEDataValues = sampleOHEData.collect()
Test.assertTrue(len(sampleOHEDataValues) == 3, 'sampleOHEData should have three elements')
Test.assertEquals(sampleOHEDataValues[0], SparseVector(7, {2: 1.0, 3: 1.0}),
'incorrect OHE for first sample')
Test.assertEquals(sampleOHEDataValues[1], SparseVector(7, {1: 1.0, 4: 1.0, 5: 1.0}),
'incorrect OHE for second sample')
Test.assertEquals(sampleOHEDataValues[2], SparseVector(7, {0: 1.0, 3: 1.0, 6: 1.0}),
'incorrect OHE for third sample')
# #### ** Part 2: Construct an OHE dictionary **
# **(2a) Pair RDD of `(featureID, category)` **
#
# To start, create an RDD of distinct `(featureID, category)` tuples. In our sample dataset, the 7 items in the resulting RDD are `(0, 'bear')`, `(0, 'cat')`, `(0, 'mouse')`, `(1, 'black')`, `(1, 'tabby')`, `(2, 'mouse')`, `(2, 'salmon')`. Notably `'black'` appears twice in the dataset but only contributes one item to the RDD: `(1, 'black')`, while `'mouse'` also appears twice and contributes two items: `(0, 'mouse')` and `(2, 'mouse')`. Use [flatMap](https://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD.flatMap) and [distinct](https://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD.distinct).
flat = sampleDataRDD.flatMap(lambda r: r).distinct()
print flat.count()
for i in flat.take(8):
print i
# TODO: Replace <FILL IN> with appropriate code
sampleDistinctFeats = (sampleDataRDD.flatMap(lambda r: r).distinct())
# TEST Pair RDD of (featureID, category) (2a)
Test.assertEquals(sorted(sampleDistinctFeats.collect()),
[(0, 'bear'), (0, 'cat'), (0, 'mouse'), (1, 'black'),
(1, 'tabby'), (2, 'mouse'), (2, 'salmon')],
'incorrect value for sampleDistinctFeats')
# ** (2b) OHE Dictionary from distinct features **
#
# Next, create an `RDD` of key-value tuples, where each `(featureID, category)` tuple in `sampleDistinctFeats` is a key and the values are distinct integers ranging from 0 to (number of keys - 1). Then convert this `RDD` into a dictionary, which can be done using the `collectAsMap` action. Note that there is no unique mapping from keys to values, as all we require is that each `(featureID, category)` key be mapped to a unique integer between 0 and the number of keys. In this exercise, any valid mapping is acceptable. Use [zipWithIndex](https://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD.zipWithIndex) followed by [collectAsMap](https://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD.collectAsMap).
#
# In our sample dataset, one valid list of key-value tuples is: `[((0, 'bear'), 0), ((2, 'salmon'), 1), ((1, 'tabby'), 2), ((2, 'mouse'), 3), ((0, 'mouse'), 4), ((0, 'cat'), 5), ((1, 'black'), 6)]`. The dictionary defined in Part (1a) illustrates another valid mapping between keys and integers.
# TODO: Replace <FILL IN> with appropriate code
sampleOHEDict = sampleDistinctFeats.zipWithIndex().collectAsMap()
print sampleOHEDict
# TEST OHE Dictionary from distinct features (2b)
Test.assertEquals(sorted(sampleOHEDict.keys()),
[(0, 'bear'), (0, 'cat'), (0, 'mouse'), (1, 'black'),
(1, 'tabby'), (2, 'mouse'), (2, 'salmon')],
'sampleOHEDict has unexpected keys')
Test.assertEquals(sorted(sampleOHEDict.values()), range(7), 'sampleOHEDict has unexpected values')
# **(2c) Automated creation of an OHE dictionary **
#
# Now use the code from Parts (2a) and (2b) to write a function that takes an input dataset and outputs an OHE dictionary. Then use this function to create an OHE dictionary for the sample dataset, and verify that it matches the dictionary from Part (2b).
# +
# TODO: Replace <FILL IN> with appropriate code
def createOneHotDict(inputData):
"""Creates a one-hot-encoder dictionary based on the input data.
Args:
inputData (RDD of lists of (int, str)): An RDD of observations where each observation is
made up of a list of (featureID, value) tuples.
Returns:
dict: A dictionary where the keys are (featureID, value) tuples and map to values that are
unique integers.
"""
flat = inputData.flatMap(lambda r: r).distinct()
return flat.zipWithIndex().collectAsMap()
sampleOHEDictAuto = createOneHotDict(sampleDataRDD)
print sampleOHEDictAuto
# -
# TEST Automated creation of an OHE dictionary (2c)
Test.assertEquals(sorted(sampleOHEDictAuto.keys()),
[(0, 'bear'), (0, 'cat'), (0, 'mouse'), (1, 'black'),
(1, 'tabby'), (2, 'mouse'), (2, 'salmon')],
'sampleOHEDictAuto has unexpected keys')
Test.assertEquals(sorted(sampleOHEDictAuto.values()), range(7),
'sampleOHEDictAuto has unexpected values')
# #### **Part 3: Parse CTR data and generate OHE features**
# Before we can proceed, you'll first need to obtain the data from Criteo. Here is the link to Criteo's data sharing agreement:[http://labs.criteo.com/downloads/2014-kaggle-display-advertising-challenge-dataset/](http://labs.criteo.com/downloads/2014-kaggle-display-advertising-challenge-dataset/). After you accept the agreement, you can obtain the download URL by right-clicking on the "Download Sample" button and clicking "Copy link address" or "Copy Link Location", depending on your browser. Paste the URL into the `# TODO` cell below. The script below will download the file and make the sample dataset's contents available in the `rawData` variable.
#
# Note that the download should complete within 30 seconds.
# +
import os.path
baseDir = os.path.join('/Users/bill.walrond/Documents/dsprj/data')
inputPath = os.path.join('CS190_Mod4', 'dac_sample.txt')
fileName = os.path.join(baseDir, inputPath)
if os.path.isfile(fileName):
rawData = (sc
.textFile(fileName, 2)
.map(lambda x: x.replace('\t', ','))) # work with either ',' or '\t' separated data
print rawData.take(1)
print rawData.count()
else:
print 'Couldn\'t find filename: %s' % fileName
# +
# TODO: Replace <FILL IN> with appropriate code
import glob
from io import BytesIO
import os.path
import tarfile
import urllib
import urlparse
# Paste in url, url should end with: dac_sample.tar.gz
url = '<FILL IN>'
url = url.strip()
if 'rawData' in locals():
print 'rawData already loaded. Nothing to do.'
elif not url.endswith('dac_sample.tar.gz'):
print 'Check your download url. Are you downloading the Sample dataset?'
else:
try:
tmp = BytesIO()
urlHandle = urllib.urlopen(url)
tmp.write(urlHandle.read())
tmp.seek(0)
tarFile = tarfile.open(fileobj=tmp)
dacSample = tarFile.extractfile('dac_sample.txt')
dacSample = [unicode(x.replace('\n', '').replace('\t', ',')) for x in dacSample]
rawData = (sc
.parallelize(dacSample, 1) # Create an RDD
.zipWithIndex() # Enumerate lines
.map(lambda (v, i): (i, v)) # Use line index as key
.partitionBy(2, lambda i: not (i < 50026)) # Match sc.textFile partitioning
.map(lambda (i, v): v)) # Remove index
print 'rawData loaded from url'
print rawData.take(1)
except IOError:
print 'Unable to unpack: {0}'.format(url)
# -
# **(3a) Loading and splitting the data **
#
# We are now ready to start working with the actual CTR data, and our first task involves splitting it into training, validation, and test sets. Use the [randomSplit method](https://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD.randomSplit) with the specified weights and seed to create RDDs storing each of these datasets, and then [cache](https://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD.cache) each of these RDDs, as we will be accessing them multiple times in the remainder of this lab. Finally, compute the size of each dataset.
# +
# TODO: Replace <FILL IN> with appropriate code
weights = [.8, .1, .1]
seed = 42
# Use randomSplit with weights and seed
rawTrainData, rawValidationData, rawTestData = rawData.randomSplit(weights, seed)
# Cache the data
rawTrainData.cache()
rawValidationData.cache()
rawTestData.cache()
nTrain = rawTrainData.count()
nVal = rawValidationData.count()
nTest = rawTestData.count()
print nTrain, nVal, nTest, nTrain + nVal + nTest
print rawTrainData.take(1)
# -
# TEST Loading and splitting the data (3a)
Test.assertTrue(all([rawTrainData.is_cached, rawValidationData.is_cached, rawTestData.is_cached]),
'you must cache the split data')
Test.assertEquals(nTrain, 79911, 'incorrect value for nTrain')
Test.assertEquals(nVal, 10075, 'incorrect value for nVal')
Test.assertEquals(nTest, 10014, 'incorrect value for nTest')
# ** (3b) Extract features **
#
# We will now parse the raw training data to create an RDD that we can subsequently use to create an OHE dictionary. Note from the `take()` command in Part (3a) that each raw data point is a string containing several fields separated by some delimiter. For now, we will ignore the first field (which is the 0-1 label), and parse the remaining fields (or raw features). To do this, complete the implemention of the `parsePoint` function.
# +
# TODO: Replace <FILL IN> with appropriate code
def parsePoint(point):
"""Converts a comma separated string into a list of (featureID, value) tuples.
Note:
featureIDs should start at 0 and increase to the number of features - 1.
Args:
point (str): A comma separated string where the first value is the label and the rest
are features.
Returns:
list: A list of (featureID, value) tuples.
"""
# make a list of (featureID, value) tuples, skipping the first element (the label)
return [(k,v) for k,v in enumerate(point[2:].split(','))]
parsedTrainFeat = rawTrainData.map(parsePoint)
print parsedTrainFeat.count()
numCategories = (parsedTrainFeat
.flatMap(lambda x: x)
.distinct()
.map(lambda x: (x[0], 1))
.reduceByKey(lambda x, y: x + y)
.sortByKey()
.collect())
print numCategories[2][1]
# -
# TEST Extract features (3b)
Test.assertEquals(numCategories[2][1], 855, 'incorrect implementation of parsePoint')
Test.assertEquals(numCategories[32][1], 4, 'incorrect implementation of parsePoint')
# **(3c) Create an OHE dictionary from the dataset **
#
# Note that parsePoint returns a data point as a list of `(featureID, category)` tuples, which is the same format as the sample dataset studied in Parts 1 and 2 of this lab. Using this observation, create an OHE dictionary using the function implemented in Part (2c). Note that we will assume for simplicity that all features in our CTR dataset are categorical.
# TODO: Replace <FILL IN> with appropriate code
ctrOHEDict = createOneHotDict(parsedTrainFeat)
print 'Len of ctrOHEDict: {0}'.format(len(ctrOHEDict))
numCtrOHEFeats = len(ctrOHEDict.keys())
print numCtrOHEFeats
print ctrOHEDict.has_key((0, ''))
theItems = ctrOHEDict.items()
for i in range(0,9):
print theItems[i]
# TEST Create an OHE dictionary from the dataset (3c)
Test.assertEquals(numCtrOHEFeats, 233286, 'incorrect number of features in ctrOHEDict')
Test.assertTrue((0, '') in ctrOHEDict, 'incorrect features in ctrOHEDict')
# ** (3d) Apply OHE to the dataset **
#
# Now let's use this OHE dictionary by starting with the raw training data and creating an RDD of [LabeledPoint](http://spark.apache.org/docs/1.3.1/api/python/pyspark.mllib.html#pyspark.mllib.regression.LabeledPoint) objects using OHE features. To do this, complete the implementation of the `parseOHEPoint` function. Hint: `parseOHEPoint` is an extension of the `parsePoint` function from Part (3b) and it uses the `oneHotEncoding` function from Part (1d).
from pyspark.mllib.regression import LabeledPoint
print rawTrainData.count()
r = rawTrainData.first()
l = parsePoint(r)
print 'Length of parsed list: %d' % len(l)
print 'Here\'s the list ...'
print l
sv = oneHotEncoding(l, ctrOHEDict, numCtrOHEFeats)
print 'Here\'s the sparsevector ...'
print sv
lp = LabeledPoint(float(r[:1]), sv)
print 'Here\'s the labeledpoint ...'
print lp
# +
# TODO: Replace <FILL IN> with appropriate code
def parseOHEPoint(point, OHEDict, numOHEFeats):
"""Obtain the label and feature vector for this raw observation.
Note:
You must use the function `oneHotEncoding` in this implementation or later portions
of this lab may not function as expected.
Args:
point (str): A comma separated string where the first value is the label and the rest
are features.
OHEDict (dict of (int, str) to int): Mapping of (featureID, value) to unique integer.
numOHEFeats (int): The number of unique features in the training dataset.
Returns:
LabeledPoint: Contains the label for the observation and the one-hot-encoding of the
raw features based on the provided OHE dictionary.
"""
# first, get the label
label = float(point[:1])
parsed = parsePoint(point)
features = oneHotEncoding(parsed, OHEDict, numOHEFeats)
# return parsed
return LabeledPoint(label,features)
def toOHEPoint(point):
return parseOHEPoint(point, ctrOHEDict, numCtrOHEFeats)
sc.setLogLevel("INFO")
rawTrainData = rawTrainData.repartition(8)
rawTrainData.cache()
OHETrainData = rawTrainData.map(toOHEPoint)
OHETrainData.cache()
print OHETrainData.take(1)
# Check that oneHotEncoding function was used in parseOHEPoint
backupOneHot = oneHotEncoding
oneHotEncoding = None
withOneHot = False
try: parseOHEPoint(rawTrainData.take(1)[0], ctrOHEDict, numCtrOHEFeats)
except TypeError: withOneHot = True
oneHotEncoding = backupOneHot
# -
# TEST Apply OHE to the dataset (3d)
numNZ = sum(parsedTrainFeat.map(lambda x: len(x)).take(5))
numNZAlt = sum(OHETrainData.map(lambda lp: len(lp.features.indices)).take(5))
Test.assertEquals(numNZ, numNZAlt, 'incorrect implementation of parseOHEPoint')
Test.assertTrue(withOneHot, 'oneHotEncoding not present in parseOHEPoint')
# **Visualization 1: Feature frequency **
#
# We will now visualize the number of times each of the 233,286 OHE features appears in the training data. We first compute the number of times each feature appears, then bucket the features by these counts. The buckets are sized by powers of 2, so the first bucket corresponds to features that appear exactly once ( \\( \scriptsize 2^0 \\) ), the second to features that appear twice ( \\( \scriptsize 2^1 \\) ), the third to features that occur between three and four ( \\( \scriptsize 2^2 \\) ) times, the fifth bucket is five to eight ( \\( \scriptsize 2^3 \\) ) times and so on. The scatter plot below shows the logarithm of the bucket thresholds versus the logarithm of the number of features that have counts that fall in the buckets.
x = sc.parallelize([("a", 1), ("b", 1), ("a", 1), ("a", 1),("b", 1), ("b", 1), ("b", 1), ("b", 1)], 3)
y = x.reduceByKey(lambda accum, n: accum + n)
y.collect()
# +
def bucketFeatByCount(featCount):
"""Bucket the counts by powers of two."""
for i in range(11):
size = 2 ** i
if featCount <= size:
return size
return -1
featCounts = (OHETrainData
.flatMap(lambda lp: lp.features.indices)
.map(lambda x: (x, 1))
.reduceByKey(lambda x, y: x + y))
featCountsBuckets = (featCounts
.map(lambda x: (bucketFeatByCount(x[1]), 1))
.filter(lambda (k, v): k != -1)
.reduceByKey(lambda x, y: x + y)
.collect())
print featCountsBuckets
# +
import matplotlib.pyplot as plt
# %matplotlib inline
x, y = zip(*featCountsBuckets)
x, y = np.log(x), np.log(y)
def preparePlot(xticks, yticks, figsize=(10.5, 6), hideLabels=False, gridColor='#999999',
gridWidth=1.0):
"""Template for generating the plot layout."""
plt.close()
fig, ax = plt.subplots(figsize=figsize, facecolor='white', edgecolor='white')
ax.axes.tick_params(labelcolor='#999999', labelsize='10')
for axis, ticks in [(ax.get_xaxis(), xticks), (ax.get_yaxis(), yticks)]:
axis.set_ticks_position('none')
axis.set_ticks(ticks)
axis.label.set_color('#999999')
if hideLabels: axis.set_ticklabels([])
plt.grid(color=gridColor, linewidth=gridWidth, linestyle='-')
map(lambda position: ax.spines[position].set_visible(False), ['bottom', 'top', 'left', 'right'])
return fig, ax
# generate layout and plot data
fig, ax = preparePlot(np.arange(0, 10, 1), np.arange(4, 14, 2))
ax.set_xlabel(r'$\log_e(bucketSize)$'), ax.set_ylabel(r'$\log_e(countInBucket)$')
plt.scatter(x, y, s=14**2, c='#d6ebf2', edgecolors='#8cbfd0', alpha=0.75)
# display(fig)
plt.show()
pass
# -
# **(3e) Handling unseen features **
#
# We naturally would like to repeat the process from Part (3d), e.g., to compute OHE features for the validation and test datasets. However, we must be careful, as some categorical values will likely appear in new data that did not exist in the training data. To deal with this situation, update the `oneHotEncoding()` function from Part (1d) to ignore previously unseen categories, and then compute OHE features for the validation data.
# +
# TODO: Replace <FILL IN> with appropriate code
def oneHotEncoding(rawFeats, OHEDict, numOHEFeats):
"""Produce a one-hot-encoding from a list of features and an OHE dictionary.
Note:
If a (featureID, value) tuple doesn't have a corresponding key in OHEDict it should be
ignored.
Args:
rawFeats (list of (int, str)): The features corresponding to a single observation. Each
feature consists of a tuple of featureID and the feature's value. (e.g. sampleOne)
OHEDict (dict): A mapping of (featureID, value) to unique integer.
numOHEFeats (int): The total number of unique OHE features (combinations of featureID and
value).
Returns:
SparseVector: A SparseVector of length numOHEFeats with indices equal to the unique
identifiers for the (featureID, value) combinations that occur in the observation and
with values equal to 1.0.
"""
newFeats = []
idx = []
for f in rawFeats:
if f in OHEDict:
newFeats += [1.0]
idx += [OHEDict[f]]
return SparseVector(numOHEFeats, sorted(idx), np.array(newFeats))
OHEValidationData = rawValidationData.map(lambda point: parseOHEPoint(point, ctrOHEDict, numCtrOHEFeats))
OHEValidationData.cache()
print OHEValidationData.take(1)
# -
# TEST Handling unseen features (3e)
numNZVal = (OHEValidationData
.map(lambda lp: len(lp.features.indices))
.sum())
Test.assertEquals(numNZVal, 372080, 'incorrect number of features')
# #### ** Part 4: CTR prediction and logloss evaluation **
# ** (4a) Logistic regression **
#
# We are now ready to train our first CTR classifier. A natural classifier to use in this setting is logistic regression, since it models the probability of a click-through event rather than returning a binary response, and when working with rare events, probabilistic predictions are useful.
#
# First use [LogisticRegressionWithSGD](https://spark.apache.org/docs/latest/api/python/pyspark.mllib.html#pyspark.mllib.classification.LogisticRegressionWithSGD) to train a model using `OHETrainData` with the given hyperparameter configuration. `LogisticRegressionWithSGD` returns a [LogisticRegressionModel](https://spark.apache.org/docs/latest/api/python/pyspark.mllib.html#pyspark.mllib.regression.LogisticRegressionModel). Next, use the `LogisticRegressionModel.weights` and `LogisticRegressionModel.intercept` attributes to print out the model's parameters. Note that these are the names of the object's attributes and should be called using a syntax like `model.weights` for a given `model`.
# +
from pyspark.mllib.classification import LogisticRegressionWithSGD
# fixed hyperparameters
numIters = 50
stepSize = 10.
regParam = 1e-6
regType = 'l2'
includeIntercept = True
# -
# TODO: Replace <FILL IN> with appropriate code
model0 = LogisticRegressionWithSGD.train(OHETrainData,
iterations=numIters,
step=stepSize,
regParam=regParam,
regType=regType,
intercept=includeIntercept)
sortedWeights = sorted(model0.weights)
print sortedWeights[:5], model0.intercept
# TEST Logistic regression (4a)
Test.assertTrue(np.allclose(model0.intercept, 0.56455084025), 'incorrect value for model0.intercept')
Test.assertTrue(np.allclose(sortedWeights[0:5],
[-0.45899236853575609, -0.37973707648623956, -0.36996558266753304,
-0.36934962879928263, -0.32697945415010637]), 'incorrect value for model0.weights')
# ** (4b) Log loss **
# Throughout this lab, we will use log loss to evaluate the quality of models. Log loss is defined as: \\[ \scriptsize \ell_{log}(p, y) = \begin{cases} -\log (p) & \text{if } y = 1 \\\ -\log(1-p) & \text{if } y = 0 \end{cases} \\] where \\( \scriptsize p\\) is a probability between 0 and 1 and \\( \scriptsize y\\) is a label of either 0 or 1. Log loss is a standard evaluation criterion when predicting rare-events such as click-through rate prediction (it is also the criterion used in the [Criteo Kaggle competition](https://www.kaggle.com/c/criteo-display-ad-challenge)).
#
# Write a function to compute log loss, and evaluate it on some sample inputs.
# +
# TODO: Replace <FILL IN> with appropriate code
from math import log
def computeLogLoss(p, y):
"""Calculates the value of log loss for a given probabilty and label.
Note:
log(0) is undefined, so when p is 0 we need to add a small value (epsilon) to it
and when p is 1 we need to subtract a small value (epsilon) from it.
Args:
p (float): A probabilty between 0 and 1.
y (int): A label. Takes on the values 0 and 1.
Returns:
float: The log loss value.
"""
epsilon = 10e-12
if p not in [0.0,1.0]:
logeval = p
elif p == 0:
logeval = p+epsilon
else:
logeval = p-epsilon
if y == 1:
return (-log(logeval))
elif y == 0:
return (-log(1-logeval))
print computeLogLoss(.5, 1)
print computeLogLoss(.5, 0)
print computeLogLoss(.99, 1)
print computeLogLoss(.99, 0)
print computeLogLoss(.01, 1)
print computeLogLoss(.01, 0)
print computeLogLoss(0, 1)
print computeLogLoss(1, 1)
print computeLogLoss(1, 0)
# -
# TEST Log loss (4b)
Test.assertTrue(np.allclose([computeLogLoss(.5, 1), computeLogLoss(.01, 0), computeLogLoss(.01, 1)],
[0.69314718056, 0.0100503358535, 4.60517018599]),
'computeLogLoss is not correct')
Test.assertTrue(np.allclose([computeLogLoss(0, 1), computeLogLoss(1, 1), computeLogLoss(1, 0)],
[25.3284360229, 1.00000008275e-11, 25.3284360229]),
'computeLogLoss needs to bound p away from 0 and 1 by epsilon')
# ** (4c) Baseline log loss **
#
# Next we will use the function we wrote in Part (4b) to compute the baseline log loss on the training data. A very simple yet natural baseline model is one where we always make the same prediction independent of the given datapoint, setting the predicted value equal to the fraction of training points that correspond to click-through events (i.e., where the label is one). Compute this value (which is simply the mean of the training labels), and then use it to compute the training log loss for the baseline model. The log loss for multiple observations is the mean of the individual log loss values.
# +
# TODO: Replace <FILL IN> with appropriate code
# Note that our dataset has a very high click-through rate by design
# In practice click-through rate can be one to two orders of magnitude lower
classOneFracTrain = OHETrainData.map(lambda p: p.label).mean()
print classOneFracTrain
logLossTrBase = OHETrainData.map(lambda p: computeLogLoss(classOneFracTrain,p.label) ).mean()
print 'Baseline Train Logloss = {0:.6f}\n'.format(logLossTrBase)
# -
# TEST Baseline log loss (4c)
Test.assertTrue(np.allclose(classOneFracTrain, 0.22717773523), 'incorrect value for classOneFracTrain')
Test.assertTrue(np.allclose(logLossTrBase, 0.535844), 'incorrect value for logLossTrBase')
# ** (4d) Predicted probability **
#
# In order to compute the log loss for the model we trained in Part (4a), we need to write code to generate predictions from this model. Write a function that computes the raw linear prediction from this logistic regression model and then passes it through a [sigmoid function](http://en.wikipedia.org/wiki/Sigmoid_function) \\( \scriptsize \sigma(t) = (1+ e^{-t})^{-1} \\) to return the model's probabilistic prediction. Then compute probabilistic predictions on the training data.
#
# Note that when incorporating an intercept into our predictions, we simply add the intercept to the value of the prediction obtained from the weights and features. Alternatively, if the intercept was included as the first weight, we would need to add a corresponding feature to our data where the feature has the value one. This is not the case here.
# +
# TODO: Replace <FILL IN> with appropriate code
from math import exp # exp(-t) = e^-t
def getP(x, w, intercept):
"""Calculate the probability for an observation given a set of weights and intercept.
Note:
We'll bound our raw prediction between 20 and -20 for numerical purposes.
Args:
x (SparseVector): A vector with values of 1.0 for features that exist in this
observation and 0.0 otherwise.
w (DenseVector): A vector of weights (betas) for the model.
intercept (float): The model's intercept.
Returns:
float: A probability between 0 and 1.
"""
rawPrediction = w.dot(x) + intercept
# Bound the raw prediction value
rawPrediction = min(rawPrediction, 20)
rawPrediction = max(rawPrediction, -20)
return ( 1 / (1 + exp(-1*rawPrediction)) )
trainingPredictions = OHETrainData.map(lambda p: getP(p.features,model0.weights, model0.intercept))
print trainingPredictions.take(5)
# -
# TEST Predicted probability (4d)
Test.assertTrue(np.allclose(trainingPredictions.sum(), 18135.4834348),
'incorrect value for trainingPredictions')
# ** (4e) Evaluate the model **
#
# We are now ready to evaluate the quality of the model we trained in Part (4a). To do this, first write a general function that takes as input a model and data, and outputs the log loss. Then run this function on the OHE training data, and compare the result with the baseline log loss.
a = OHETrainData.map(lambda p: (getP(p.features, model0.weights, model0.intercept), p.label))
print a.count()
print a.take(5)
b = a.map(lambda lp: computeLogLoss(lp[0],lp[1]))
print b.count()
print b.take(5)
# +
# TODO: Replace <FILL IN> with appropriate code
def evaluateResults(model, data):
"""Calculates the log loss for the data given the model.
Args:
model (LogisticRegressionModel): A trained logistic regression model.
data (RDD of LabeledPoint): Labels and features for each observation.
Returns:
float: Log loss for the data.
"""
# Run a map to create an RDD of (prediction, label) tuples
preds_labels = data.map(lambda p: (getP(p.features, model.weights, model.intercept), p.label))
return preds_labels.map(lambda lp: computeLogLoss(lp[0], lp[1])).mean()
logLossTrLR0 = evaluateResults(model0, OHETrainData)
print ('OHE Features Train Logloss:\n\tBaseline = {0:.3f}\n\tLogReg = {1:.6f}'
.format(logLossTrBase, logLossTrLR0))
# -
# TEST Evaluate the model (4e)
Test.assertTrue(np.allclose(logLossTrLR0, 0.456903), 'incorrect value for logLossTrLR0')
# ** (4f) Validation log loss **
#
# Next, following the same logic as in Parts (4c) and 4(e), compute the validation log loss for both the baseline and logistic regression models. Notably, the baseline model for the validation data should still be based on the label fraction from the training dataset.
# +
# TODO: Replace <FILL IN> with appropriate code
logLossValBase = OHEValidationData.map(lambda p: computeLogLoss(classOneFracTrain, p.label)).mean()
logLossValLR0 = evaluateResults(model0, OHEValidationData)
print ('OHE Features Validation Logloss:\n\tBaseline = {0:.3f}\n\tLogReg = {1:.6f}'
.format(logLossValBase, logLossValLR0))
# -
# TEST Validation log loss (4f)
Test.assertTrue(np.allclose(logLossValBase, 0.527603), 'incorrect value for logLossValBase')
Test.assertTrue(np.allclose(logLossValLR0, 0.456957), 'incorrect value for logLossValLR0')
# **Visualization 2: ROC curve **
#
# We will now visualize how well the model predicts our target. To do this we generate a plot of the ROC curve. The ROC curve shows us the trade-off between the false positive rate and true positive rate, as we liberalize the threshold required to predict a positive outcome. A random model is represented by the dashed line.
# +
labelsAndScores = OHEValidationData.map(lambda lp:
(lp.label, getP(lp.features, model0.weights, model0.intercept)))
labelsAndWeights = labelsAndScores.collect()
labelsAndWeights.sort(key=lambda (k, v): v, reverse=True)
labelsByWeight = np.array([k for (k, v) in labelsAndWeights])
length = labelsByWeight.size
truePositives = labelsByWeight.cumsum()
numPositive = truePositives[-1]
falsePositives = np.arange(1.0, length + 1, 1.) - truePositives
truePositiveRate = truePositives / numPositive
falsePositiveRate = falsePositives / (length - numPositive)
# Generate layout and plot data
fig, ax = preparePlot(np.arange(0., 1.1, 0.1), np.arange(0., 1.1, 0.1))
ax.set_xlim(-.05, 1.05), ax.set_ylim(-.05, 1.05)
ax.set_ylabel('True Positive Rate (Sensitivity)')
ax.set_xlabel('False Positive Rate (1 - Specificity)')
plt.plot(falsePositiveRate, truePositiveRate, color='#8cbfd0', linestyle='-', linewidth=3.)
plt.plot((0., 1.), (0., 1.), linestyle='--', color='#d6ebf2', linewidth=2.) # Baseline model
# display(fig)
plt.show()
pass
# -
# #### **Part 5: Reduce feature dimension via feature hashing**
# ** (5a) Hash function **
#
# As we just saw, using a one-hot-encoding featurization can yield a model with good statistical accuracy. However, the number of distinct categories across all features is quite large -- recall that we observed 233K categories in the training data in Part (3c). Moreover, the full Kaggle training dataset includes more than 33M distinct categories, and the Kaggle dataset itself is just a small subset of Criteo's labeled data. Hence, featurizing via a one-hot-encoding representation would lead to a very large feature vector. To reduce the dimensionality of the feature space, we will use feature hashing.
#
# Below is the hash function that we will use for this part of the lab. We will first use this hash function with the three sample data points from Part (1a) to gain some intuition. Specifically, run code to hash the three sample points using two different values for `numBuckets` and observe the resulting hashed feature dictionaries.
# +
from collections import defaultdict
import hashlib
def hashFunction(numBuckets, rawFeats, printMapping=False):
"""Calculate a feature dictionary for an observation's features based on hashing.
Note:
Use printMapping=True for debug purposes and to better understand how the hashing works.
Args:
numBuckets (int): Number of buckets to use as features.
rawFeats (list of (int, str)): A list of features for an observation. Represented as
(featureID, value) tuples.
printMapping (bool, optional): If true, the mappings of featureString to index will be
printed.
Returns:
dict of int to float: The keys will be integers which represent the buckets that the
features have been hashed to. The value for a given key will contain the count of the
(featureID, value) tuples that have hashed to that key.
"""
mapping = {}
for ind, category in rawFeats:
featureString = category + str(ind)
mapping[featureString] = int(int(hashlib.md5(featureString).hexdigest(), 16) % numBuckets)
if(printMapping): print mapping
sparseFeatures = defaultdict(float)
for bucket in mapping.values():
sparseFeatures[bucket] += 1.0
return dict(sparseFeatures)
# Reminder of the sample values:
# sampleOne = [(0, 'mouse'), (1, 'black')]
# sampleTwo = [(0, 'cat'), (1, 'tabby'), (2, 'mouse')]
# sampleThree = [(0, 'bear'), (1, 'black'), (2, 'salmon')]
# +
# TODO: Replace <FILL IN> with appropriate code
# Use four buckets
sampOneFourBuckets = hashFunction(4, sampleOne, True)
sampTwoFourBuckets = hashFunction(4, sampleTwo, True)
sampThreeFourBuckets = hashFunction(4, sampleThree, True)
# Use one hundred buckets
sampOneHundredBuckets = hashFunction(100, sampleOne, True)
sampTwoHundredBuckets = hashFunction(100, sampleTwo, True)
sampThreeHundredBuckets = hashFunction(100, sampleThree, True)
print '\t\t 4 Buckets \t\t\t 100 Buckets'
print 'SampleOne:\t {0}\t\t {1}'.format(sampOneFourBuckets, sampOneHundredBuckets)
print 'SampleTwo:\t {0}\t\t {1}'.format(sampTwoFourBuckets, sampTwoHundredBuckets)
print 'SampleThree:\t {0}\t {1}'.format(sampThreeFourBuckets, sampThreeHundredBuckets)
# -
# TEST Hash function (5a)
Test.assertEquals(sampOneFourBuckets, {2: 1.0, 3: 1.0}, 'incorrect value for sampOneFourBuckets')
Test.assertEquals(sampThreeHundredBuckets, {72: 1.0, 5: 1.0, 14: 1.0},
'incorrect value for sampThreeHundredBuckets')
# ** (5b) Creating hashed features **
#
# Next we will use this hash function to create hashed features for our CTR datasets. First write a function that uses the hash function from Part (5a) with numBuckets = \\( \scriptsize 2^{15} \approx 33K \\) to create a `LabeledPoint` with hashed features stored as a `SparseVector`. Then use this function to create new training, validation and test datasets with hashed features. Hint: `parsedHashPoint` is similar to `parseOHEPoint` from Part (3d).
feats = [(k,v) for k,v in enumerate(rawTrainData.take(1)[0][2:].split(','))]
print feats
hashDict = hashFunction(2 ** 15, feats)
print hashDict
print len(hashDict)
print 2**15
# +
# TODO: Replace <FILL IN> with appropriate code
def parseHashPoint(point, numBuckets):
"""Create a LabeledPoint for this observation using hashing.
Args:
point (str): A comma separated string where the first value is the label and the rest are
features.
numBuckets: The number of buckets to hash to.
Returns:
LabeledPoint: A LabeledPoint with a label (0.0 or 1.0) and a SparseVector of hashed
features.
"""
label = float(point[:1])
rawFeats = [(k,v) for k,v in enumerate(point[2:].split(','))]
hashDict = hashFunction(numBuckets, rawFeats)
return LabeledPoint(label,SparseVector(len(hashDict), sorted(hashDict.keys()), hashDict.values()))
numBucketsCTR = 2 ** 15
hashTrainData = rawTrainData.map(lambda r: parseHashPoint(r,numBucketsCTR))
hashTrainData.cache()
hashValidationData = rawValidationData.map(lambda r: parseHashPoint(r,numBucketsCTR))
hashValidationData.cache()
hashTestData = rawTestData.map(lambda r: parseHashPoint(r,numBucketsCTR))
hashTestData.cache()
a = hashTrainData.take(1)
print a
# +
# TEST Creating hashed features (5b)
hashTrainDataFeatureSum = sum(hashTrainData
.map(lambda lp: len(lp.features.indices))
.take(20))
print hashTrainDataFeatureSum
hashTrainDataLabelSum = sum(hashTrainData
.map(lambda lp: lp.label)
.take(100))
print hashTrainDataLabelSum
hashValidationDataFeatureSum = sum(hashValidationData
.map(lambda lp: len(lp.features.indices))
.take(20))
hashValidationDataLabelSum = sum(hashValidationData
.map(lambda lp: lp.label)
.take(100))
hashTestDataFeatureSum = sum(hashTestData
.map(lambda lp: len(lp.features.indices))
.take(20))
hashTestDataLabelSum = sum(hashTestData
.map(lambda lp: lp.label)
.take(100))
Test.assertEquals(hashTrainDataFeatureSum, 772, 'incorrect number of features in hashTrainData')
Test.assertEquals(hashTrainDataLabelSum, 24.0, 'incorrect labels in hashTrainData')
Test.assertEquals(hashValidationDataFeatureSum, 776,
'incorrect number of features in hashValidationData')
Test.assertEquals(hashValidationDataLabelSum, 16.0, 'incorrect labels in hashValidationData')
Test.assertEquals(hashTestDataFeatureSum, 774, 'incorrect number of features in hashTestData')
Test.assertEquals(hashTestDataLabelSum, 23.0, 'incorrect labels in hashTestData')
# -
# ** (5c) Sparsity **
#
# Since we have 33K hashed features versus 233K OHE features, we should expect OHE features to be sparser. Verify this hypothesis by computing the average sparsity of the OHE and the hashed training datasets.
#
# Note that if you have a `SparseVector` named `sparse`, calling `len(sparse)` returns the total number of features, not the number features with entries. `SparseVector` objects have the attributes `indices` and `values` that contain information about which features are nonzero. Continuing with our example, these can be accessed using `sparse.indices` and `sparse.values`, respectively.
s = sum(hashTrainData.map(lambda lp: len(lp.features.indices) / float(numBucketsCTR) ).collect()) / nTrain
# ratios.count()
s
# +
# TODO: Replace <FILL IN> with appropriate code
def computeSparsity(data, d, n):
"""Calculates the average sparsity for the features in an RDD of LabeledPoints.
Args:
data (RDD of LabeledPoint): The LabeledPoints to use in the sparsity calculation.
d (int): The total number of features.
n (int): The number of observations in the RDD.
Returns:
float: The average of the ratio of features in a point to total features.
"""
return sum(hashTrainData.map(lambda lp: len(lp.features.indices) / float(d) ).collect()) / n
averageSparsityHash = computeSparsity(hashTrainData, numBucketsCTR, nTrain)
averageSparsityOHE = computeSparsity(OHETrainData, numCtrOHEFeats, nTrain)
print 'Average OHE Sparsity: {0:.7e}'.format(averageSparsityOHE)
print 'Average Hash Sparsity: {0:.7e}'.format(averageSparsityHash)
# -
# TEST Sparsity (5c)
Test.assertTrue(np.allclose(averageSparsityOHE, 1.6717677e-04),
'incorrect value for averageSparsityOHE')
Test.assertTrue(np.allclose(averageSparsityHash, 1.1805561e-03),
'incorrect value for averageSparsityHash')
sc.stop()
| jupyter_notebooks/CS190-1x Module 4- Feature Hashing Lab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
from bs4 import BeautifulSoup
import scrapy
from lxml.etree import fromstring
import requests
import time
import re
import json
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
from six.moves.urllib import parse
import selenium.webdriver.chrome.service as service
from pyvirtualdisplay import Display
from selenium.webdriver.support.ui import WebDriverWait
# # scroll page function opens the webpage and keep scrollinig till the end of the page until no more new results are being fetched
def scroll_page(url):
driver = webdriver.Chrome('/Users/mohan/Documents/medius_health_scraper/chromedriver')
driver.get(url)
driver.execute_script("window.scrollTo(0, document.body.scrollHeight);")
time.sleep(3)
html = driver.find_element_by_class_name('css-y1gt6f').get_attribute('innerHTML')
return html
def get_soup(x):
soup = BeautifulSoup(x,'lxml')
return soup
# # extract_name fun returns the name of the disease in the webpage
# # extract_url fun returns the url or link for each disease page
def extract_name(x):
name = x.find('h2',{'class':'h1 css-kjyn3a egb32mv1'})
name = name.text
return name
def extract_url(x):
url = x.find('a',attrs = {'href':re.compile('/health/*')})
url = url.get('href')
complete_url = 'https://www.healthline.com'+url
return complete_url
# # extract_details fun returns the symptoms listed for each disease in the web page
def extract_details(x):
name = extract_name(x)
complete_url = extract_url(x)
response = requests.get(complete_url)
source_code = response.text
sub_soup = get_soup(source_code)
symptoms = sub_soup.find('a',attrs = {'name':re.compile('symptoms*')})
return name,complete_url,symptoms
start_url = 'https://www.healthline.com/symptom/dizziness'
def extract_symptoms(symptoms):
symptoms_list = []
counter = symptoms.findNext('ul').contents
for i in range(len(counter)):
if counter[i] != ' ':
symptoms_list.append(symptoms.findNext('ul').contents[i].string)
symptoms_list = list(filter(None.__ne__, symptoms_list))
return symptoms_list
# # start_scraping fun recursively parse through all the pages of all diseases and get symptoms associated with each disease
# # final_symptoms is a dictionary with key as disease and values being the symptoms associated with each disease
final_symptoms = {}
def start_scraping(soup):
for disease in soup.findAll('li', {'class': 'css-ee6cf6'}):
name,complete_url,symptoms = extract_details(disease)
print(name)
print(complete_url)
if symptoms:
symptoms_list = []
symptoms_list = extract_symptoms(symptoms)
final_symptoms[name] = symptoms_list
#pass
print(symptoms_list)
else:
#sub_html = scroll_page(complete_url)
response = requests.get(complete_url)
source_code = response.text
sub_soup = get_soup(source_code)
start_scraping(sub_soup)
# # i have extracted symptoms for about 30 records by iterating recursively through the web pages for demonstration and saved in the sample_output.json file
homepage_html = scroll_page(start_url)
soup = get_soup(homepage_html)
start_scraping(soup)
len(final_symptoms)
json = json.dumps(final_symptoms)
with open('sample_output.json','w') as output_file:
output_file.write(json)
# # Since the webpage response is slow we extract all the records into our database as json documents and iterate over json documents to get the desired results as we need by parsing through the values/symptoms or combinations as we need.
for key,values in final_symptoms.items():
if 'dizziness' in values:
print(key)
# # A sample interaction which returns a disease having both dizziness and rapid heartbeat as symptoms
for key,values in final_symptoms.items():
if 'dizziness' and 'rapid heartbeat' in values:
print(key)
# # A sample scrap showing list of all 84 possible conditions causing dizziness
for disease in soup.findAll('li', {'class': 'css-ee6cf6'}):
name,complete_url,symptoms = extract_details(disease)
print(name)
print(complete_url)
| scrape_symptoms.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Based on similar work with Twin Cities Pioneer Press [Schools that Work](http://www.twincities.com/2010/07/10/lessons-from-a-school-that-works/)
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
# %matplotlib inline
# ## Setting things up
# Let's load the data and give it a quick look.
df = pd.read_csv('data/apib12tx.csv')
df.describe()
# ## Checking out correlations
# Let's start looking at how variables in our dataset relate to each other so we know what to expect when we start modeling.
df.corr()
# The percentage of students enrolled in free/reduced-price lunch programs is often used as a proxy for poverty.
df.plot(kind="scatter", x="MEALS", y="API12B")
# Conversely, the education level of a student's parents is often a good predictor of how well a student will do in school.
df.plot(kind="scatter", x="AVG_ED", y="API12B")
# ## Running the regression
# Like we did last week, we'll use scikit-learn to run basic single-variable regressions. Let's start by looking at California's Academic Performance index as it relates to the percentage of students, per school, enrolled in free/reduced-price lunch programs.
data = np.asarray(df[['API12B','MEALS']])
x, y = data[:, 1:], data[:, 0]
lr = LinearRegression()
lr.fit(x, y)
# plot the linear regression line on the scatter plot
lr.coef_
lr.score(x, y)
plt.scatter(x, y, color='blue')
plt.plot(x, lr.predict(x), color='red', linewidth=1)
# In our naive universe where we're only paying attention to two variables -- academic performance and free/reduced lunch -- we can clearly see that some percentage of schools is overperforming the performance that would be expected of them, taking poverty out of the equation.
#
# A handful, in particular, seem to be dramatically overperforming. Let's look at them:
df[(df['MEALS'] >= 80) & (df['API12B'] >= 900)]
# Let's look specifically at Solano Avenue Elementary, which has an API of 922 and 80 percent of students being in the free/reduced lunch program. If you were to use the above regression to predict how well Solano would do, it would look like this:
lr.predict(80)
# With an index of 922, clearly the school is overperforming what our simplified model expects.
| 07 Teaching Machines/.ipynb_checkpoints/regression_review-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
from IPython.display import Image
# # CNTK 101: Logistic Regression and ML Primer
#
# This tutorial is targeted to individuals who are new to CNTK and to machine learning. In this tutorial, you will train a simple yet powerful machine learning model that is widely used in industry for a variety of applications. The model trained below scales to massive data sets in the most expeditious manner by harnessing computational scalability leveraging the computational resources you may have (one or more CPU cores, one or more GPUs, a cluster of CPUs or a cluster of GPUs), transparently via the CNTK library.
#
# The following notebook users Python APIs. If you are looking for this example in BrainScript, please look [here](https://github.com/Microsoft/CNTK/tree/v2.0.beta15.0/Tutorials/HelloWorld-LogisticRegression).
#
# ## Introduction
#
# **Problem**:
# A cancer hospital has provided data and wants us to determine if a patient has a fatal [malignant][] cancer vs. a benign growth. This is known as a classification problem. To help classify each patient, we are given their age and the size of the tumor. Intuitively, one can imagine that younger patients and/or patient with small tumor size are less likely to have malignant cancer. The data set simulates this application where the each observation is a patient represented as a dot (in the plot below) where red color indicates malignant and blue indicates benign disease. Note: This is a toy example for learning, in real life there are large number of features from different tests/examination sources and doctors' experience that play into the diagnosis/treatment decision for a patient.
# [malignant]: https://en.wikipedia.org/wiki/Malignancy
# Figure 1
Image(url="https://www.cntk.ai/jup/cancer_data_plot.jpg", width=400, height=400)
# **Goal**:
# Our goal is to learn a classifier that automatically can label any patient into either benign or malignant category given two features (age and tumor size). In this tutorial, we will create a linear classifier that is a fundamental building-block in deep networks.
# Figure 2
Image(url= "https://www.cntk.ai/jup/cancer_classify_plot.jpg", width=400, height=400)
# In the figure above, the green line represents the learnt model from the data and separates the blue dots from the red dots. In this tutorial, we will walk you through the steps to learn the green line. Note: this classifier does make mistakes where couple of blue dots are on the wrong side of the green line. However, there are ways to fix this and we will look into some of the techniques in later tutorials.
#
# **Approach**:
# Any learning algorithm has typically five stages. These are Data reading, Data preprocessing, Creating a model, Learning the model parameters, and Evaluating (a.k.a. testing/prediction) the model.
#
# >1. Data reading: We generate simulated data sets with each sample having two features (plotted below) indicative of the age and tumor size.
# >2. Data preprocessing: Often the individual features such as size or age needs to be scaled. Typically one would scale the data between 0 and 1. To keep things simple, we are not doing any scaling in this tutorial (for details look here: [feature scaling][]).
# >3. Model creation: We introduce a basic linear model in this tutorial.
# >4. Learning the model: This is also known as training. While fitting a linear model can be done in a variety of ways ([linear regression][]), in CNTK we use Stochastic Gradient Descent a.k.a. [SGD][].
# >5. Evaluation: This is also known as testing where one takes data sets with known labels (a.k.a. ground-truth) that was not ever used for training. This allows us to assess how a model would perform in real world (previously unseen) observations.
#
# ## Logistic Regression
# [Logistic regression][] is fundamental machine learning technique that uses a linear weighted combination of features and generates the probability of predicting different classes. In our case the classifier will generate a probability in [0,1] which can then be compared with a threshold (such as 0.5) to produce a binary label (0 or 1). However, the method shown can be extended to multiple classes easily.
# [feature scaling]: https://en.wikipedia.org/wiki/Feature_scaling
# [SGD]: https://en.wikipedia.org/wiki/Stochastic_gradient_descent
# [linear regression]: https://en.wikipedia.org/wiki/Linear_regression
# [logistic regression]: https://en.wikipedia.org/wiki/Logistic_regression
# [softmax]: https://en.wikipedia.org/wiki/Multinomial_logistic_regression
# Figure 3
Image(url= "https://www.cntk.ai/jup/logistic_neuron.jpg", width=300, height=200)
# In the figure above, contributions from different input features are linearly weighted and aggregated. The resulting sum is mapped to a 0-1 range via a [sigmoid][] function. For classifiers with more than two output labels, one can use a [softmax][] function.
# [sigmoid]: https://en.wikipedia.org/wiki/Sigmoid_function
# +
# Import the relevant components
from __future__ import print_function
import numpy as np
import sys
import os
from cntk import *
# Select the right target device when this notebook is being tested:
if 'TEST_DEVICE' in os.environ:
import cntk
if os.environ['TEST_DEVICE'] == 'cpu':
cntk.device.try_set_default_device(cntk.device.cpu())
else:
cntk.device.try_set_default_device(cntk.device.gpu(0))
# -
# ## Data Generation
# Let us generate some synthetic data emulating the cancer example using `numpy` library. We have two features (represented in two-dimensions) each either being to one of the two classes (benign:blue dot or malignant:red dot).
#
# In our example, each observation in the training data has a label (blue or red) corresponding to each observation (set of features - age and size). In this example, we have two classes represented by labels 0 or 1, thus a binary classification task.
# Define the network
input_dim = 2
num_output_classes = 2
# ### Input and Labels
#
# In this tutorial we are generating synthetic data using `numpy` library. In real world problems, one would use a [reader][], that would read feature values (`features`: *age* and *tumor size*) corresponding to each obeservation (patient). The simulated *age* variable is scaled down to have similar range as the other variable. This is a key aspect of data pre-processing that we will learn more in later tutorials. Note, each observation can reside in a higher dimension space (when more features are available) and will be represented as a [tensor][] in CNTK. More advanced tutorials shall introduce the handling of high dimensional data.
#
# [reader]: https://github.com/Microsoft/CNTK/search?p=1&q=reader&type=Wikis&utf8=%E2%9C%93
# [tensor]: https://en.wikipedia.org/wiki/Tensor
# +
# Ensure we always get the same amount of randomness
np.random.seed(0)
# Helper function to generate a random data sample
def generate_random_data_sample(sample_size, feature_dim, num_classes):
# Create synthetic data using NumPy.
Y = np.random.randint(size=(sample_size, 1), low=0, high=num_classes)
# Make sure that the data is separable
X = (np.random.randn(sample_size, feature_dim)+3) * (Y+1)
# Specify the data type to match the input variable used later in the tutorial
# (default type is double)
X = X.astype(np.float32)
# converting class 0 into the vector "1 0 0",
# class 1 into vector "0 1 0", ...
class_ind = [Y==class_number for class_number in range(num_classes)]
Y = np.asarray(np.hstack(class_ind), dtype=np.float32)
return X, Y
# -
# Create the input variables denoting the features and the label data. Note: the input_variable
# does not need additional info on number of observations (Samples) since CNTK creates only
# the network topology first
mysamplesize = 32
features, labels = generate_random_data_sample(mysamplesize, input_dim, num_output_classes)
# Let us visualize the input data.
#
# **Note**: If the import of `matplotlib.pyplot` fails, please run `conda install matplotlib` which will fix the `pyplot` version dependencies. If you are on a python environment different from Anaconda, then use `pip install`.
# +
# Plot the data
import matplotlib.pyplot as plt
# %matplotlib inline
# given this is a 2 class ()
colors = ['r' if l == 0 else 'b' for l in labels[:,0]]
plt.scatter(features[:,0], features[:,1], c=colors)
plt.xlabel("Scaled age (in yrs)")
plt.ylabel("Tumor size (in cm)")
plt.show()
# -
# # Model Creation
#
# A logistic regression (a.k.a. LR) network is the simplest building block but has been powering many ML
# applications in the past decade. LR is a simple linear model that takes as input, a vector of numbers describing the properties of what we are classifying (also known as a feature vector, $\bf{x}$, the blue nodes in the figure) and emits the *evidence* ($z$) (output of the green node, a.k.a. as activation). Each feature in the input layer is connected with a output node by a corresponding weight w (indicated by the black lines of varying thickness).
# Figure 4
Image(url= "https://www.cntk.ai/jup/logistic_neuron2.jpg", width=300, height=200)
# The first step is to compute the evidence for an observation.
#
# $$z = \sum_{i=1}^n w_i \times x_i + b = \textbf{w} \cdot \textbf{x} + b$$
#
# where $\bf{w}$ is the weight vector of length $n$ and $b$ is known as the [bias][] term. Note: we use **bold** notation to denote vectors.
#
# The computed evidence is mapped to a 0-1 scale using a [`sigmoid`][] (when the outcome can take one of two values) or a `softmax` function (when the outcome can take one of more than 2 classes value).
#
# Network input and output:
# - **input** variable (a key CNTK concept):
# >An **input** variable is a user-code facing container where user-provided code fills in different observations (data point or sample, equivalent to a blue/red dot in our example) as inputs to the model function during model learning (a.k.a.training) and model evaluation (a.k.a. testing). Thus, the shape of the `input_variable` must match the shape of the data that will be provided. For example, when data are images each of height 10 pixels and width 5 pixels, the input feature dimension will be 2 (representing image height and width). Similarly, in our example the dimensions are age and tumor size, thus `input_dim` = 2. More on data and their dimensions to appear in separate tutorials.
#
# [bias]: https://www.quora.com/What-does-the-bias-term-represent-in-logistic-regression
#
# [`sigmoid`]: https://en.wikipedia.org/wiki/Sigmoid_function
input = input_variable(input_dim, np.float32)
# ## Network setup
#
# The `linear_layer` function is a straight forward implementation of the equation above. We perform two operations:
# 0. multiply the weights ($\bf{w}$) with the features ($\bf{x}$) using CNTK `times` operator and add individual features' contribution,
# 1. add the bias term $b$.
#
# These CNTK operations are optimized for execution on the available hardware and the implementation hides the complexity away from the user.
# +
# Define a dictionary to store the model parameters
mydict = {"w":None,"b":None}
def linear_layer(input_var, output_dim):
input_dim = input_var.shape[0]
weight_param = parameter(shape=(input_dim, output_dim))
bias_param = parameter(shape=(output_dim))
mydict['w'], mydict['b'] = weight_param, bias_param
return times(input_var, weight_param) + bias_param
# -
# `z` will be used to represent the output of a network.
output_dim = num_output_classes
z = linear_layer(input, output_dim)
# ### Learning model parameters
#
# Now that the network is setup, we would like to learn the parameters $\bf w$ and $b$ for our simple linear layer. To do so we convert, the computed evidence ($z$) into a set of predicted probabilities ($\textbf p$) using a `softmax` function.
#
# $$ \textbf{p} = \mathrm{softmax}(z)$$
#
# The `softmax` is an activation function that maps the accumulated evidences to a probability distribution over the classes (Details of the [softmax function][]). Other choices of activation function can be [found here][].
#
# [softmax function]: https://www.cntk.ai/pythondocs/cntk.ops.html#cntk.ops.softmax
#
# [found here]: https://github.com/Microsoft/CNTK/wiki/Activation-Functions
# ## Training
# The output of the `softmax` is a probability of observations belonging to the respective classes. For training the classifier, we need to determine what behavior the model needs to mimic. In other words, we want the generated probabilities to be as close as possible to the observed labels. This function is called the *cost* or *loss* function and shows what is the difference between the learnt model vs. that generated by the training set.
#
# [`Cross-entropy`][] is a popular function to measure the loss. It is defined as:
#
# $$ H(p) = - \sum_{j=1}^C y_j \log (p_j) $$
#
# where $p$ is our predicted probability from `softmax` function and $y$ represents the label. This label provided with the data for training is also called the ground-truth label. In the two-class example, the `label` variable has dimensions of two (equal to the `num_output_classes` or $C$). Generally speaking, if the task in hand requires classification into $C$ different classes, the label variable will have $C$ elements with 0 everywhere except for the class represented by the data point where it will be 1. Understanding the [details][] of this cross-entropy function is highly recommended.
#
# [`cross-entropy`]: http://cntk.ai/pythondocs/cntk.ops.html#cntk.ops.cross_entropy_with_softmax
# [details]: http://colah.github.io/posts/2015-09-Visual-Information/
label = input_variable((num_output_classes), np.float32)
loss = cross_entropy_with_softmax(z, label)
# #### Evaluation
#
# In order to evaluate the classification, one can compare the output of the network which for each observation emits a vector of evidences (can be converted into probabilities using `softmax` functions) with dimension equal to number of classes.
eval_error = classification_error(z, label)
# ### Configure training
#
# The trainer strives to reduce the `loss` function by different optimization approaches, [Stochastic Gradient Descent][] (`sgd`) being one of the most popular one. Typically, one would start with random initialization of the model parameters. The `sgd` optimizer would calculate the `loss` or error between the predicted label against the corresponding ground-truth label and using [gradient-decent][] generate a new set model parameters in a single iteration.
#
# The aforementioned model parameter update using a single observation at a time is attractive since it does not require the entire data set (all observation) to be loaded in memory and also requires gradient computation over fewer datapoints, thus allowing for training on large data sets. However, the updates generated using a single observation sample at a time can vary wildly between iterations. An intermediate ground is to load a small set of observations and use an average of the `loss` or error from that set to update the model parameters. This subset is called a *minibatch*.
#
# With minibatches we often sample observation from the larger training dataset. We repeat the process of model parameters update using different combination of training samples and over a period of time minimize the `loss` (and the error). When the incremental error rates are no longer changing significantly or after a preset number of maximum minibatches to train, we claim that our model is trained.
#
# One of the key parameter for optimization is called the `learning_rate`. For now, we can think of it as a scaling factor that modulates how much we change the parameters in any iteration. We will be covering more details in later tutorial.
# With this information, we are ready to create our trainer.
#
# [optimization]: https://en.wikipedia.org/wiki/Category:Convex_optimization
# [Stochastic Gradient Descent]: https://en.wikipedia.org/wiki/Stochastic_gradient_descent
# [gradient-decent]: http://www.statisticsviews.com/details/feature/5722691/Getting-to-the-Bottom-of-Regression-with-Gradient-Descent.html
# Instantiate the trainer object to drive the model training
learning_rate = 0.5
lr_schedule = learning_rate_schedule(learning_rate, UnitType.minibatch)
learner = sgd(z.parameters, lr_schedule)
trainer = Trainer(z, (loss, eval_error), [learner])
# First let us create some helper functions that will be needed to visualize different functions associated with training. Note these convinience functions are for understanding what goes under the hood.
# +
from cntk.utils import get_train_eval_criterion, get_train_loss
# Define a utility function to compute the moving average sum.
# A more efficient implementation is possible with np.cumsum() function
def moving_average(a, w=10):
if len(a) < w:
return a[:]
return [val if idx < w else sum(a[(idx-w):idx])/w for idx, val in enumerate(a)]
# Defines a utility that prints the training progress
def print_training_progress(trainer, mb, frequency, verbose=1):
training_loss, eval_error = "NA", "NA"
if mb % frequency == 0:
training_loss = get_train_loss(trainer)
eval_error = get_train_eval_criterion(trainer)
if verbose:
print ("Minibatch: {0}, Loss: {1:.4f}, Error: {2:.2f}".format(mb, training_loss, eval_error))
return mb, training_loss, eval_error
# -
# ### Run the trainer
#
# We are now ready to train our Logistic Regression model. We want to decide what data we need to feed into the training engine.
#
# In this example, each iteration of the optimizer will work on 25 samples (25 dots w.r.t. the plot above) a.k.a. `minibatch_size`. We would like to train on say 20000 observations. If the number of samples in the data is only 10000, the trainer will make 2 passes through the data. This is represented by `num_minibatches_to_train`. Note: In real world case, we would be given a certain amount of labeled data (in the context of this example, observation (age, size) and what they mean (benign / malignant)). We would use a large number of observations for training say 70% and set aside the remainder for evaluation of the trained model.
#
# With these parameters we can proceed with training our simple feedforward network.
# Initialize the parameters for the trainer
minibatch_size = 25
num_samples_to_train = 20000
num_minibatches_to_train = int(num_samples_to_train / minibatch_size)
# +
# Run the trainer and perform model training
training_progress_output_freq = 50
plotdata = {"batchsize":[], "loss":[], "error":[]}
for i in range(0, num_minibatches_to_train):
features, labels = generate_random_data_sample(minibatch_size, input_dim, num_output_classes)
# Specify input variables mapping in the model to actual minibatch data to be trained with
trainer.train_minibatch({input : features, label : labels})
batchsize, loss, error = print_training_progress(trainer, i,
training_progress_output_freq, verbose=1)
if not (loss == "NA" or error =="NA"):
plotdata["batchsize"].append(batchsize)
plotdata["loss"].append(loss)
plotdata["error"].append(error)
# +
# Compute the moving average loss to smooth out the noise in SGD
plotdata["avgloss"] = moving_average(plotdata["loss"])
plotdata["avgerror"] = moving_average(plotdata["error"])
# Plot the training loss and the training error
import matplotlib.pyplot as plt
plt.figure(1)
plt.subplot(211)
plt.plot(plotdata["batchsize"], plotdata["avgloss"], 'b--')
plt.xlabel('Minibatch number')
plt.ylabel('Loss')
plt.title('Minibatch run vs. Training loss')
plt.show()
plt.subplot(212)
plt.plot(plotdata["batchsize"], plotdata["avgerror"], 'r--')
plt.xlabel('Minibatch number')
plt.ylabel('Label Prediction Error')
plt.title('Minibatch run vs. Label Prediction Error')
plt.show()
# -
# ## Evaluation / Testing
#
# Now that we have trained the network. Let us evaluate the trained network on data that hasn't been used for training. This is called **testing**. Let us create some new data and evaluate the average error and loss on this set. This is done using `trainer.test_minibatch`. Note the error on this previously unseen data is comparable to training error. This is a **key** check. Should the error be larger than the training error by a large margin, it indicates that the trained model will not perform well on data that it has not seen during training. This is known as [overfitting][]. There are several ways to address overfitting that is beyond the scope of this tutorial but the Cognitive Toolkit provides the necessary components to address overfitting.
#
# Note: We are testing on a single minibatch for illustrative purposes. In practice one runs several minibatches of test data and reports the average.
#
# **Question** Why is this suggested? Try plotting the test error over several set of generated data sample and plot using plotting functions used for training. Do you see a pattern?
#
# [overfitting]: https://en.wikipedia.org/wiki/Overfitting
#
# +
# Run the trained model on newly generated dataset
test_minibatch_size = 25
features, labels = generate_random_data_sample(test_minibatch_size, input_dim, num_output_classes)
trainer.test_minibatch({input : features, label : labels})
# -
# ### Checking prediction / evaluation
# For evaluation, we map the output of the network between 0-1 and convert them into probabilities for the two classes. This suggests the chances of each observation being malignant and benign. We use a softmax function to get the probabilities of each of the class.
out = softmax(z)
result = out.eval({input : features})
# Let us compare the ground-truth label with the predictions. They should be in agreement.
#
# **Question:**
# - How many predictions were mislabeled? Can you change the code below to identify which observations were misclassified?
print("Label :", [np.argmax(label) for label in labels])
print("Predicted:", [np.argmax(result[i,:,:]) for i in range(result.shape[0])])
# ### Visualization
# It is desirable to visualize the results. In this example, the data is conveniently in two dimensions and can be plotted. For data with higher dimensions, visualization can be challenging. There are advanced dimensionality reduction techniques that allow for such visualizations [t-sne][].
#
# [t-sne]: https://en.wikipedia.org/wiki/T-distributed_stochastic_neighbor_embedding
# +
# Model parameters
print(mydict['b'].value)
bias_vector = mydict['b'].value
weight_matrix = mydict['w'].value
# Plot the data
import matplotlib.pyplot as plt
# given this is a 2 class
colors = ['r' if l == 0 else 'b' for l in labels[:,0]]
plt.scatter(features[:,0], features[:,1], c=colors)
plt.plot([0, bias_vector[0]/weight_matrix[0][1]],
[ bias_vector[1]/weight_matrix[0][0], 0], c = 'g', lw = 3)
plt.xlabel("Scaled age (in yrs)")
plt.ylabel("Tumor size (in cm)")
plt.show()
# -
# **Exploration Suggestions**
# - Try exploring how the classifier behaves with different data distributions - suggest changing the `minibatch_size` parameter from 25 to say 64. Why is the error increasing?
# - Try exploring different activation functions
# - Try exploring different learners
# - You can explore training a [multiclass logistic regression][] classifier.
#
# [multiclass logistic regression]: https://en.wikipedia.org/wiki/Multinomial_logistic_regression
| Tutorials/CNTK_101_LogisticRegression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python Baby Steps - Krakens
# A primeira coisa que devemos fazer antes de começar a programar é...
print('hello world!')
# # Variables
# +
#string
first="Tom"
middle="Cruise"
last="Mapother"
print("Full Name:",first,middle,last)
print("Type:",type(first))
#numbers
base = 15
height = 22
area = 1/2*(base*height)
print("Area:", area)
print("Type:",type(area))
#boolean
learn_python=True
learn_fortran=False
print("Values:", learn_fortran, learn_python)
print("Type:",type(learn_python))
# -
# # Input
n1 = input("Enter first number: ")
n2 = input("Enter second number: ")
sum = float(n1)+float(n2)
# sum = n1 + n2
print("Sum is:",sum)
# # List
colors = ['red','green','blue','yellow', 'black']
colors
type(colors)
colors[0]
colors[-1]
colors[1:3]
colors[:2]
colors[2:]
'red' in colors
'grey' in colors
colors.append('purple')
colors
colors.insert(2,'grey')
colors
# # Dict
# +
members = {
'board': 6,
'hard': 8,
'mark': 7,
'mech': 9
}
def add():
squad = input("Enter squad name to add: ")
squad = squad.lower()
if squad in members:
print("Squad already exist in our dataset. Terminating")
return
p = input(f"Enter population for {squad}")
p = int(p)
members[squad]=p
print_all()
def remove():
squad = input("Enter squad name to remove: ")
squad = squad.lower()
if squad not in members:
print("Squad doesn't exist in our dataset. Terminating")
return
del members[squad]
print_all()
def query():
squad = input("Enter squad name to query: ")
squad = squad.lower()
if squad not in members:
print("Squad doesn't exist in our dataset. Terminating")
return
print(f"Population of {squad} is: {members[squad]}")
def print_all():
for squad, p in members.items():
print(f"{squad} ==> {p}")
def main():
op = input("Enter operation (add, remove, query or print): ")
if op.lower() == 'add':
add()
elif op.lower() == 'remove':
remove()
elif op.lower() == 'query':
query()
elif op.lower() == 'print':
print_all()
# -
main()
# # Tuples (unchangeable)
# +
import math
def circle_calc(radius):
area = math.pi*(radius**2)
circumference = 2*math.pi*radius
diameter = 2*radius
return area, circumference,diameter
r = input("Enter a radius: ")
r = float(r)
area, c, d = circle_calc(r)
print(f"area {area}, circumference {c}, diameter {d}")
# -
# # Strings (and List idx operations)
food='spam and eggs!'
food[0]
food[-1]
food[:4]
food[5:8]
food[9:]
len(food)
food.upper()
# # If
#
n = input("Enter a number: ")
n = int(n)
if n%2 == 0:
print("Number is even")
else:
print("Number is odd")
# Ternário
print("Ternary operator demo")
n = input("Enter a number:")
n = int(n)
message = "Number is even" if n%2==0 else "Number is odd"
print(message)
# +
brazil = ["sao paulo","rio de janeiro","itajuba","florianopolis"]
italy = ["roma","florenca","milao"]
eua = ["new york","los angeles","chicago"]
dish = input("Enter a dish city:")
if dish in brazil:
print(f"{dish} is Brazil")
elif dish in italy:
print(f"{dish} is Italy")
elif dish in eua:
print(f"{dish} is USA")
else:
print(f"Based on my limited knowledge, I don't know which cuisine is {dish}")
# -
# # Loop
#
# +
expenses = [1200,1500,1300,1700]
total = 0
for el in expenses:
total = total + el
print(total)
# -
total = 0
for i in range(len(expenses)):
print(f"Month {i+1}, expense: {expenses[i]}")
total += expenses[i]
print(f"Total expenses is {total}")
num = 0
while num <= 10:
print(num)
num = num+1
# # Functions
# +
def print_pattern(n=5):
for i in range(n):
s = ''
for j in range(i+1):
s = f'{s}*' #using f-string
print(s)
print("Print pattern with input=3")
print_pattern(3)
print("Print pattern with input=4")
print_pattern(4)
# +
def calculate_area(dimension1,dimension2,shape="triangle"):
if shape == "triangle":
area = 1/2*(dimension1*dimension2)
elif shape == "rectangle":
area = dimension1*dimension2
else:
print("***Error: Input shape is neither triangle nor rectangle.")
area = None
return area
base = 10
height = 5
triangle_area = calculate_area(base,height)
print(f"Area of triangle is: {triangle_area}")
length = 20
width = 30
rectangle_area = calculate_area(length,width,"rectangle")
print(f"Area of rectangle is: {rectangle_area}")
# -
# # Read and Write
# read file
f = open("funny.txt","r")
for line in f:
print(line)
f.close()
# write file
f = open("funny.txt","a")
f.write("\nI love python")
f.close()
# # Have Fun
import this
from __future__ import braces
import antigravity
| Lectures/tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
from kaggle_1c_predict_future_sales.cv import sample_no_future, train_no_future, test_dates, \
Pipeline, FeatureExtractor
# +
from sklearn.linear_model import LinearRegression, LogisticRegression
from sklearn.base import BaseEstimator
class LogReg(BaseEstimator):
def __init__(self):
self.logistic_regressor = LogisticRegression()
def fit(self, X, y):
self.logistic_regressor.fit(X, y)
return self
def predict(self, X):
return self.logistic_regressor.predict(X)
pipeline = Pipeline(
LogReg(),
FeatureExtractor(categorical_cols=['shop_id']),
sample_no_future, test_dates)
pipeline.cv()
pipeline.fit()
pipeline.predict()
pipeline.prepare_submit('__deleteme__', force=True)
# -
| 110_logistic_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
a = "hello"
b = a
hex(id(a))
hex(id(b))
a = "hello"
b = "hello"
hex(id(a))
hex(id(b))
b = "Hello World"
hex(id(b))
hex(id(a))
a
b
a = [1, 2, 3]
b = a
hex(id(a))
hex(id(b))
b.append(100)
b
a
hex(id(a))
hex(id(b))
a = 500
b = 500
hex(id(a))
hex(id(b))
| my_classes/variables_memory/shared_references_and_mutability.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install keras tensorflow
from keras import backend
from imutils import paths
image_paths = list(paths.list_images('datasets/animals/'))
# %matplotlib inline
import cv2
import matplotlib.pyplot as plt
import random
import numpy as np
random.shuffle(image_paths)
# +
data = []
labels = []
for image_path in image_paths:
image = cv2.imread(image_path)
image = cv2.resize(image, (32, 32))
label = image_path.split('/')[-2]
data.append(image)
labels.append(label)
# -
data = np.array(data) / 255.0
labels = np.array(labels)
# +
from sklearn.preprocessing import LabelBinarizer
lb = LabelBinarizer()
labels = lb.fit_transform(labels)
# -
data.shape
from keras.models import Sequential
from keras.layers.convolutional import Conv2D
from keras.layers.core import Dense, Flatten
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
data, labels, test_size=0.2, random_state=33
)
model = Sequential()
model.add(
Conv2D(
32,
(3, 3),
input_shape=(32, 32, 3),
activation='relu',
padding='same'
)
)
model.add(Flatten())
model.add(Dense(3, activation='softmax'))
model.summary()
model.compile(
loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy']
)
model.fit(X_train, y_train, epochs=10, validation_split=0.2)
y_pred = model.predict(X_test)
y_pred
y_pred[0]
from sklearn.metrics import classification_report
print(classification_report(
y_test.argmax(axis=1),
y_pred.argmax(axis=1),
target_names=lb.classes_))
from keras.layers.convolutional import MaxPooling2D
from keras.layers.core import Dropout
model = Sequential()
model.add(Conv2D(32, (3, 3), input_shape=(32, 32, 3), activation='relu', padding='same'))
model.add(Conv2D(128, (3, 3), activation='relu', padding='same'))
model.add(Conv2D(128, (3, 3), activation='relu', padding='same'))
model.add(MaxPooling2D(pool_size=(3, 3)))
model.add(Dropout(0.2))
model.add(Flatten())
model.add(Dense(3, activation='softmax'))
model.summary()
model.compile(
loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy']
)
model.fit(X_train, y_train, epochs=10, validation_split=0.2)
y_pred = model.predict(X_test)
print(classification_report(
y_test.argmax(axis=1),
y_pred.argmax(axis=1),
target_names=lb.classes_))
model.save('mymodel.h5')
import pickle
f = open('lb.pkl', 'wb')
pickle.dump(lb, f)
f.close()
https://bit.ly/2OKCTYm
| cv-odds/cnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text"
# # Pretraining a Transformer from scratch with KerasNLP
#
# **Author:** [<NAME>](https://github.com/mattdangerw/)<br>
# **Date created:** 2022/04/18<br>
# **Last modified:** 2022/04/18<br>
# **Description:** Use KerasNLP to train a Transformer model from scratch.
# + [markdown] colab_type="text"
# KerasNLP aims to make it easy to build state-of-the-art text processing models. In this
# guide, we will show how library components simplify pretraining and fine-tuning a
# Transformer model from scratch.
#
# This guide is broken into three parts:
#
# 1. *Setup*, task definition, and establishing a baseline.
# 2. *Pretraining* a Transformer model.
# 3. *Fine-tuning* the Transformer model on our classification task.
# + [markdown] colab_type="text"
# ## Setup
#
# To begin, we can import `keras_nlp`, `keras` and `tensorflow`.
#
# A simple thing we can do right off the bat is to enable
# [mixed precision](https://keras.io/api/mixed_precision/), which will speed up training by
# running most of our computations with 16 bit (instead of 32 bit) floating point numbers.
# Training a Transformer can take a while, so it is important to pull out all the stops for
# faster training!
# + colab_type="code"
# !pip install -q keras-nlp
# + colab_type="code"
import os
import keras_nlp
import tensorflow as tf
from tensorflow import keras
policy = keras.mixed_precision.Policy("mixed_float16")
keras.mixed_precision.set_global_policy(policy)
# + [markdown] colab_type="text"
# Next up, can download two datasets.
#
# - [SST-2](https://paperswithcode.com/sota/sentiment-analysis-on-sst-2-binary) a text
# classification dataset and our "end goal". This dataset is often used to benchmark
# language models.
# - [WikiText-103](https://paperswithcode.com/dataset/wikitext-103): A medium sized
# collection of featured articles from English wikipedia, which we will use for
# pretraining.
#
# Finally, we will download a WordPiece vocabulary, to do sub-word tokenization later on in
# this guide.
# + colab_type="code"
# Download pretraining data.
keras.utils.get_file(
origin="https://s3.amazonaws.com/research.metamind.io/wikitext/wikitext-103-raw-v1.zip",
extract=True,
)
wiki_dir = os.path.expanduser("~/.keras/datasets/wikitext-103-raw/")
# Download finetuning data.
keras.utils.get_file(
origin="https://dl.fbaipublicfiles.com/glue/data/SST-2.zip", extract=True,
)
sst_dir = os.path.expanduser("~/.keras/datasets/SST-2/")
# Download vocabulary data.
vocab_file = keras.utils.get_file(
origin="https://storage.googleapis.com/tensorflow/keras-nlp/examples/bert/bert_vocab_uncased.txt",
)
# + [markdown] colab_type="text"
# Next, we define some hyperparameters we will use during training.
# + colab_type="code"
# Preprocessing params.
PRETRAINING_BATCH_SIZE = 128
FINETUNING_BATCH_SIZE = 32
SEQ_LENGTH = 128
MASK_RATE = 0.25
PREDICTIONS_PER_SEQ = 32
# Model params.
NUM_LAYERS = 3
MODEL_DIM = 256
INTERMEDIATE_DIM = 512
NUM_HEADS = 4
DROPOUT = 0.1
NORM_EPSILON = 1e-5
# Training params.
PRETRAINING_LEARNING_RATE = 5e-4
PRETRAINING_EPOCHS = 8
FINETUNING_LEARNING_RATE = 5e-5
FINETUNING_EPOCHS = 3
# + [markdown] colab_type="text"
# ### Load data
#
# We load our data with [tf.data](https://www.tensorflow.org/guide/data), which will allow
# us to define input pipelines for tokenizing and preprocessing text.
# + colab_type="code"
# Load SST-2.
sst_train_ds = tf.data.experimental.CsvDataset(
sst_dir + "train.tsv", [tf.string, tf.int32], header=True, field_delim="\t"
).batch(FINETUNING_BATCH_SIZE)
sst_val_ds = tf.data.experimental.CsvDataset(
sst_dir + "dev.tsv", [tf.string, tf.int32], header=True, field_delim="\t"
).batch(FINETUNING_BATCH_SIZE)
# Load wikitext-103 and filter out short lines.
wiki_train_ds = (
tf.data.TextLineDataset(wiki_dir + "wiki.train.raw")
.filter(lambda x: tf.strings.length(x) > 100)
.batch(PRETRAINING_BATCH_SIZE)
)
wiki_val_ds = (
tf.data.TextLineDataset(wiki_dir + "wiki.valid.raw")
.filter(lambda x: tf.strings.length(x) > 100)
.batch(PRETRAINING_BATCH_SIZE)
)
# Take a peak at the sst-2 dataset.
print(sst_train_ds.unbatch().batch(4).take(1).get_single_element())
# + [markdown] colab_type="text"
# You can see that our `SST-2` dataset contains relatively short snippets of movie review
# text. Our goal is to predict the sentiment of the snippet. A label of 1 indicates
# positive sentiment, and a label of 0 negative sentiment.
# + [markdown] colab_type="text"
# ### Establish a baseline
#
# As a first step, we will establish a baseline of good performance. We don't actually need
# KerasNLP for this, we can just use core Keras layers.
#
# We will train a simple bag-of-words model, where we learn a positive or negative weight
# for each word in our vocabulary. A sample's score is simply the sum of the weights of all
# words that are present in the sample.
# + colab_type="code"
# This layer will turn our input sentence into a list of 1s and 0s the same size
# our vocabulary, indicating whether a word is present in absent.
multi_hot_layer = keras.layers.TextVectorization(
max_tokens=4000, output_mode="multi_hot"
)
multi_hot_layer.adapt(sst_train_ds.map(lambda x, y: x))
# We then learn a linear regression over that layer, and that's our entire
# baseline model!
regression_layer = keras.layers.Dense(1, activation="sigmoid")
inputs = keras.Input(shape=(), dtype="string")
outputs = regression_layer(multi_hot_layer(inputs))
baseline_model = keras.Model(inputs, outputs)
baseline_model.compile(loss="binary_crossentropy", metrics=["accuracy"])
baseline_model.fit(sst_train_ds, validation_data=sst_val_ds, epochs=5)
# + [markdown] colab_type="text"
# A bag-of-words approach can be a fast and suprisingly powerful, especially when input
# examples contain a large number of words. With shorter sequences, it can hit a
# performance ceiling.
#
# To do better, we would like to build a model that can evaluate words *in context*. Instead
# of evaluating each word in a void, we need to use the information contained in the
# *entire ordered sequence* of our input.
#
# This runs us into a problem. `SST-2` is very small dataset, and there's simply not enough
# example text to attempt to build a larger, more parameterized model that can learn on a
# sequence. We would quickly start to overfit and memorize our training set, without any
# increase in our ability to generalize to unseen examples.
#
# Enter **pretraining**, which will allow us to learn on a larger corpus, and transfer our
# knowledge to the `SST-2` task. And enter **KerasNLP**, which will allow us to pretrain a
# particularly powerful model, the Transformer, with ease.
# + [markdown] colab_type="text"
# ## Pretraining
#
# To beat our baseline, we will leverage the `WikiText103` dataset, an unlabeled
# collection of wikipedia articles that is much bigger than than `SST-2`.
#
# We are going to train a *transformer*, a highly expressive model which will learn
# to embed each word in our input as a low dimentional vector. Our wikipedia dataset has no
# labels, so we will use an unsupervised training objective called the *Masked Language
# Modeling* (MLM) ojective.
#
# Essentially, we will be playing a big game of "guess the missing word". For each input
# sample we will obscure 25% of our input data, and train our model to predict the parts we
# covered up.
# + [markdown] colab_type="text"
# ### Preprocess data for the MLM task
#
# Our text preprocessing for the MLM task will occur in two stages.
#
# 1. Tokenize input text into integer sequences of token ids.
# 2. Mask certain positions in our input to predict on.
#
# To tokenize, we can use a `keras_nlp.tokenizers.Tokenizer` -- the KerasNLP building block
# for transforming text into sequences of integer token ids.
#
# In particular, we will use `keras_nlp.tokenizers.WordPieceTokenizer` which does
# *sub-word* tokenization. Sub-word tokenization is popular when training models on large
# text corpora. Essentially, it allows our model to learn from uncommon words, while not
# requireing a massive vocabulary of every word in our training set.
#
# The second thing we need to do is mask our input for the MLM task. To do this, we can use
# `keras_nlp.layers.MLMMaskGenerator`, which will randomly select a set of tokens in each
# input and mask them out.
#
# The tokenizer and the masking layer can both be used inside a call to
# [tf.data.Dataset.map](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#map).
# We can use `tf.data` to efficiently pre-compute each batch on the CPU, while our GPU or TPU
# works on training with the batch that came before. Because our masking layer will
# choose new words to mask each time, each epoch over our dataset will give us a totally
# new set of labels to train on.
# + colab_type="code"
# Setting sequence_length will trim or pad the token outputs to shape
# (batch_size, SEQ_LENGTH).
tokenizer = keras_nlp.tokenizers.WordPieceTokenizer(
vocabulary=vocab_file, sequence_length=SEQ_LENGTH,
)
# Setting mask_selection_length will trim or pad the mask outputs to shape
# (batch_size, PREDICTIONS_PER_SEQ).
masker = keras_nlp.layers.MLMMaskGenerator(
vocabulary_size=tokenizer.vocabulary_size(),
mask_selection_rate=MASK_RATE,
mask_selection_length=PREDICTIONS_PER_SEQ,
mask_token_id=tokenizer.token_to_id("[MASK]"),
)
def preprocess(inputs):
inputs = tokenizer(inputs)
outputs = masker(inputs)
# Split the masking layer outputs into a (features, labels, and weights)
# tuple that we can use with keras.Model.fit().
features = {
"tokens": outputs["tokens"],
"mask_positions": outputs["mask_positions"],
}
labels = outputs["mask_ids"]
weights = outputs["mask_weights"]
return features, labels, weights
# We use prefetch() to pre-compute preprocessed batches on the fly on the CPU.
pretrain_ds = wiki_train_ds.map(
preprocess, num_parallel_calls=tf.data.AUTOTUNE
).prefetch(tf.data.AUTOTUNE)
pretrain_val_ds = wiki_val_ds.map(
preprocess, num_parallel_calls=tf.data.AUTOTUNE
).prefetch(tf.data.AUTOTUNE)
# Preview a single input example.
# The masks will change each time you run the cell.
print(pretrain_val_ds.take(1).get_single_element())
# + [markdown] colab_type="text"
# The above block sorts our dataset into a `(features, labels, weights)` tuple, which can be
# passed directly to `keras.Model.fit()`.
#
# We have two features:
#
# 1. `"tokens"`, where some tokens have been replaced with our mask token id.
# 2. `"mask_positions"`, which keeps track of which tokens we masked out.
#
# Our labels are simply the ids we masked out.
#
# Because not all sequences will have the same number of masks, we also keep a
# `sample_weight` tensor, which removes padded labels from our loss function by giving them
# zero weight.
# + [markdown] colab_type="text"
# ### Create the Transformer encoder
#
# KerasNLP provides all the building blocks to quickly build a Transformer encoder.
#
# We use `keras_nlp.layers.TokenAndPositionEmbedding` to first embed our input token ids.
# This layer simultaneously learns two embeddings -- one for words in a sentence and another
# for integer positions in a sentence. The output embedding is simply the sum of the two.
#
# Then we can add a series of `keras_nlp.layers.TransformerEncoder` layers. These are the
# bread and butter of the Transformer model, using an attention mechanism to attend to
# different parts of the input sentence, followed by a multi-layer perceptron block.
#
# The output of this model will be a encoded vector per input token id. Unlike the
# bag-of-words model we used as a baseline, this model will embed each token accounting for
# the context in which it appeared.
# + colab_type="code"
inputs = keras.Input(shape=(SEQ_LENGTH,), dtype=tf.int32)
# Embed our tokens with a positional embedding.
embedding_layer = keras_nlp.layers.TokenAndPositionEmbedding(
vocabulary_size=tokenizer.vocabulary_size(),
sequence_length=SEQ_LENGTH,
embedding_dim=MODEL_DIM,
)
outputs = embedding_layer(inputs)
# Apply layer normalization and dropout to the embedding.
outputs = keras.layers.LayerNormalization(epsilon=NORM_EPSILON)(outputs)
outputs = keras.layers.Dropout(rate=DROPOUT)(outputs)
# Add a number of encoder blocks
for i in range(NUM_LAYERS):
outputs = keras_nlp.layers.TransformerEncoder(
intermediate_dim=INTERMEDIATE_DIM,
num_heads=NUM_HEADS,
dropout=DROPOUT,
layer_norm_epsilon=NORM_EPSILON,
)(outputs)
encoder_model = keras.Model(inputs, outputs)
encoder_model.summary()
# + [markdown] colab_type="text"
# ### Pretrain the Transformer
#
# You can think of the `encoder_model` as it's own modular unit, it is the piece of our
# model that we are really interested in for our downstream task. However we still need to
# set up the encoder to train on the MLM task; to do that we attach a
# `keras_nlp.layers.MLMHead`.
#
# This layer will take as one input the token encodings, and as another the positions we
# masked out in the original input. It will gather the token encodings we masked, and
# transform them back in predictions over our entire vocabulary.
#
# With that, we are ready to compile and run pretraining. If you are running this in a
# colab, note that this will take about an hour. Training Transformer is famously compute
# intesive, so even this relatively small Transformer will take some time.
# + colab_type="code"
# Create the pretraining model by attaching a masked language model head.
inputs = {
"tokens": keras.Input(shape=(SEQ_LENGTH,), dtype=tf.int32),
"mask_positions": keras.Input(shape=(PREDICTIONS_PER_SEQ,), dtype=tf.int32),
}
# Encode the tokens.
encoded_tokens = encoder_model(inputs["tokens"])
# Predict an output word for each masked input token.
# We use the input token embedding to project from our encoded vectors to
# vocabulary logits, which has been shown to improve training efficiency.
outputs = keras_nlp.layers.MLMHead(
embedding_weights=embedding_layer.token_embedding.embeddings, activation="softmax",
)(encoded_tokens, mask_positions=inputs["mask_positions"])
# Define and compile our pretraining model.
pretraining_model = keras.Model(inputs, outputs)
pretraining_model.compile(
loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.Adam(learning_rate=PRETRAINING_LEARNING_RATE),
weighted_metrics=["sparse_categorical_accuracy"],
jit_compile=True,
)
# Pretrain the model on our wiki text dataset.
pretraining_model.fit(
pretrain_ds, validation_data=pretrain_val_ds, epochs=PRETRAINING_EPOCHS,
)
# Save this base model for further finetuning.
encoder_model.save("encoder_model")
# + [markdown] colab_type="text"
# ## Fine-tuning
#
# After pretraining, we can now fine-tune our model on the `SST-2` dataset. We can
# leverage the ability of the encoder we build to predict on words in context to boost our
# our performance on the downstream task.
# + [markdown] colab_type="text"
# ### Preprocess data for classification
#
# Preprocessing for fine-tuning is much simpler than for our pretraining MLM task. We just
# tokenize our input sentences and we are ready for training!
# + colab_type="code"
def preprocess(sentences, labels):
return tokenizer(sentences), labels
# We use prefetch() to pre-compute preprocessed batches on the fly on our CPU.
finetune_ds = sst_train_ds.map(
preprocess, num_parallel_calls=tf.data.AUTOTUNE
).prefetch(tf.data.AUTOTUNE)
finetune_val_ds = sst_val_ds.map(
preprocess, num_parallel_calls=tf.data.AUTOTUNE
).prefetch(tf.data.AUTOTUNE)
# Preview a single input example.
print(finetune_val_ds.take(1).get_single_element())
# + [markdown] colab_type="text"
# ### Fine-tune the Transformer
#
# To go from our encoded token output to a classification prediction, we need to attach
# another "head" to our Transformer model. We can afford to be simple here. We pool
# the encoded tokens together, and use a single dense layer to make a prediction.
# + colab_type="code"
# Reload the encoder model from disk so we can restart fine-tuning from scratch.
encoder_model = keras.models.load_model("encoder_model", compile=False)
# Take as input the tokenized input.
inputs = keras.Input(shape=(SEQ_LENGTH,), dtype=tf.int32)
# Encode and pool the tokens.
encoded_tokens = encoder_model(inputs)
pooled_tokens = keras.layers.GlobalAveragePooling1D()(encoded_tokens)
# Predict an output label.
outputs = keras.layers.Dense(1, activation="sigmoid")(pooled_tokens)
# Define and compile our finetuning model.
finetuning_model = keras.Model(inputs, outputs)
finetuning_model.compile(
loss="binary_crossentropy",
optimizer=keras.optimizers.Adam(learning_rate=FINETUNING_LEARNING_RATE),
metrics=["accuracy"],
)
# Finetune the model for the SST-2 task.
finetuning_model.fit(
finetune_ds, validation_data=finetune_val_ds, epochs=FINETUNING_EPOCHS,
)
# + [markdown] colab_type="text"
# Pretraining was enough to boost our performance to 84%, and this is hardly the ceiling
# for Transformer models. You may have noticed during pretraining that our validation
# performance was still steadily increasing. Our model is still significantly undertrained.
# Training for more epochs, training a large Transformer, and training on more unlabeled
# text would all continue to boost performance significantly.
# + [markdown] colab_type="text"
# ### Save a model that accepts raw text
#
# The last thing we can do with our fine-tuned model is saveing including our tokenization
# layer. One of the key advantages of KerasNLP is all preprocessing is done inside the
# [TensorFlow graph](https://www.tensorflow.org/guide/intro_to_graphs), making it possible
# to save and restore a model that can directly run inference on raw text!
# + colab_type="code"
# Add our tokenization into our final model.
inputs = keras.Input(shape=(), dtype=tf.string)
tokens = tokenizer(inputs)
outputs = finetuning_model(tokens)
final_model = keras.Model(inputs, outputs)
final_model.save("final_model")
# This model can predict directly on raw text.
restored_model = keras.models.load_model("final_model", compile=False)
inference_data = tf.constant(["Terrible, no good, trash.", "So great; I loved it!"])
print(restored_model(inference_data))
# + [markdown] colab_type="text"
# One of the key goals of KerasNLP is to provide a modular approach to NLP model building.
# We have shown one approach to building a Transformer here, but KerasNLP supports an ever
# growing array of components for preprocessing text and building models. We hope it makes
# it easier to experiment on solutions to your natural language problems.
| guides/ipynb/keras_nlp/transformer_pretraining.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.029712, "end_time": "2022-02-06T06:59:52.601217", "exception": false, "start_time": "2022-02-06T06:59:52.571505", "status": "completed"} tags=[]
# # Short term trading strategies that work
#
# In this notebook, strategies mentioned in the book 'Short term trading strategies that work' from <NAME> are investigated. This book mainly focuses on short-term mean-reversion strategies in the US equity market.
# + papermill={"duration": 2.639199, "end_time": "2022-02-06T06:59:55.269150", "exception": false, "start_time": "2022-02-06T06:59:52.629951", "status": "completed"} tags=[]
# %matplotlib inline
from datetime import datetime
import logging
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
import matplotlib.pyplot as plt
plt.style.use('bmh')
import pandas as pd
from vivace.backtest import (BacktestEngine, Strategy, Weighting, InstrumentDef,
Performance, signal, processing, rebase, PnLType)
from vivace.backtest.engine import long_only_returns
# + [markdown] papermill={"duration": 0.028022, "end_time": "2022-02-06T06:59:55.325543", "exception": false, "start_time": "2022-02-06T06:59:55.297521", "status": "completed"} tags=[]
# # Strategy 1
# If the following conditions are met, go long the equity market for 5 days.
# - It went down 3 days in a row
# - Price is above 200 day MA. Rationale is to go long when long-term trend is up and capture temporal pullback.
# + papermill={"duration": 0.03358, "end_time": "2022-02-06T06:59:55.387214", "exception": false, "start_time": "2022-02-06T06:59:55.353634", "status": "completed"} tags=[]
instruments = ['SPY', 'QQQ', '^GDAXI', '^N225']
# + papermill={"duration": 0.039236, "end_time": "2022-02-06T06:59:55.455223", "exception": false, "start_time": "2022-02-06T06:59:55.415987", "status": "completed"} tags=[]
def run_strat_1(instrument):
engine = BacktestEngine(
strategy=Strategy.DELTA_ONE.value,
instrument=instrument,
signal=signal.Pipeline([
signal.ParallelSignal([
signal.GreaterThanMA(lookback=200, shift=1),
signal.ConsecutiveSign(n_consecutive=3, lookback_direction=-1, position=1)
], weighting='product', post_process=processing.CarryOver(5))
]),
log_level=logging.WARNING,
)
engine.run()
return engine
def run_backtests(backtest_func, instruments):
result = []
for name in instruments:
instrument = InstrumentDef(name, pnl_type=PnLType.ADJCLOSE_TO_ADJCLOSE.value)
strat_engine = backtest_func(instrument)
strat_equity_curve = strat_engine.calculate_equity_curve(calculate_net=False).rename(f'Strategy ({name})')
long_only = long_only_returns(instrument, strat_equity_curve.index[0], strat_equity_curve.index[-1])
long_only_equity_curve = long_only.add(1).cumprod().rename(f'Long-only ({name})')
result.append(
pd.concat((long_only_equity_curve, strat_equity_curve), axis=1).pipe(rebase)
)
return result
# + papermill={"duration": 1.800287, "end_time": "2022-02-06T06:59:57.284644", "exception": false, "start_time": "2022-02-06T06:59:55.484357", "status": "completed"} tags=[]
strategy_1_result = run_backtests(run_strat_1, instruments)
# + papermill={"duration": 1.784922, "end_time": "2022-02-06T06:59:59.110493", "exception": false, "start_time": "2022-02-06T06:59:57.325571", "status": "completed"} tags=[]
fig, axes = plt.subplots(1, len(instruments), figsize=(14, 5), sharey=True)
for ax, result in zip(axes, strategy_1_result):
result.plot(ax=ax, logy=True)
# + papermill={"duration": 0.342843, "end_time": "2022-02-06T06:59:59.482862", "exception": false, "start_time": "2022-02-06T06:59:59.140019", "status": "completed"} tags=[]
pd.concat([i.pipe(Performance).summary() for i in strategy_1_result], axis=1)
# + papermill={"duration": 1.507879, "end_time": "2022-02-06T07:00:01.056081", "exception": false, "start_time": "2022-02-06T06:59:59.548202", "status": "completed"} tags=[]
fig, axes = plt.subplots(1, len(instruments), figsize=(14, 5), sharey=True)
for ax, result in zip(axes, strategy_1_result):
result.tail(252 * 2).pipe(rebase).plot(ax=ax, logy=True)
# + [markdown] papermill={"duration": 0.032347, "end_time": "2022-02-06T07:00:01.121533", "exception": false, "start_time": "2022-02-06T07:00:01.089186", "status": "completed"} tags=[]
# # Strategy 2
#
# If the following conditions are met, go long the equity market for 5 days.
# - It the market made new 10-day lows
# - Price is above 200 day MA. Rationale is to go long when long-term trend is up and capture temporal pullback.
# + papermill={"duration": 0.048516, "end_time": "2022-02-06T07:00:01.202677", "exception": false, "start_time": "2022-02-06T07:00:01.154161", "status": "completed"} tags=[]
def run_strat_2(instrument):
engine = BacktestEngine(
strategy=Strategy.DELTA_ONE.value,
instrument=instrument,
signal=signal.Pipeline([
signal.ParallelSignal([
signal.GreaterThanMA(lookback=200, shift=1),
signal.LocalMinimum(lookback=10, shift=1)
], weighting='product', post_process=processing.CarryOver(5))
]),
log_level=logging.WARNING,
)
engine.run()
return engine
# + papermill={"duration": 0.973404, "end_time": "2022-02-06T07:00:02.225757", "exception": false, "start_time": "2022-02-06T07:00:01.252353", "status": "completed"} tags=[]
strategy_2_result = run_backtests(run_strat_2, instruments)
# + papermill={"duration": 1.801248, "end_time": "2022-02-06T07:00:04.093881", "exception": false, "start_time": "2022-02-06T07:00:02.292633", "status": "completed"} tags=[]
fig, axes = plt.subplots(1, len(instruments), figsize=(14, 5), sharey=True)
for ax, result in zip(axes, strategy_2_result):
result.plot(ax=ax, logy=True)
# + papermill={"duration": 0.342355, "end_time": "2022-02-06T07:00:04.488091", "exception": false, "start_time": "2022-02-06T07:00:04.145736", "status": "completed"} tags=[]
pd.concat([i.pipe(Performance).summary() for i in strategy_2_result], axis=1)
# + papermill={"duration": 1.658087, "end_time": "2022-02-06T07:00:06.198718", "exception": false, "start_time": "2022-02-06T07:00:04.540631", "status": "completed"} tags=[]
fig, axes = plt.subplots(1, len(instruments), figsize=(14, 5), sharey=True)
for ax, result in zip(axes, strategy_2_result):
result.tail(252 * 2).pipe(rebase).plot(ax=ax, logy=True)
# + [markdown] papermill={"duration": 0.088292, "end_time": "2022-02-06T07:00:06.328076", "exception": false, "start_time": "2022-02-06T07:00:06.239784", "status": "completed"} tags=[]
# # Strategy 3
#
# This strategy is the so-called "Double 7's strategy".
# + papermill={"duration": 0.044792, "end_time": "2022-02-06T07:00:06.414593", "exception": false, "start_time": "2022-02-06T07:00:06.369801", "status": "completed"} tags=[]
def run_strat_3(instrument):
engine = BacktestEngine(
strategy=Strategy.DELTA_ONE.value,
instrument=instrument,
signal=signal.Double7Connors(),
log_level=logging.WARNING,
)
engine.run()
return engine
# + papermill={"duration": 1.454875, "end_time": "2022-02-06T07:00:07.976608", "exception": false, "start_time": "2022-02-06T07:00:06.521733", "status": "completed"} tags=[]
strategy_3_result = run_backtests(run_strat_3, instruments)
# + papermill={"duration": 1.78589, "end_time": "2022-02-06T07:00:09.803374", "exception": false, "start_time": "2022-02-06T07:00:08.017484", "status": "completed"} tags=[]
fig, axes = plt.subplots(1, len(instruments), figsize=(14, 5), sharey=True)
for ax, result in zip(axes, strategy_3_result):
result.plot(ax=ax, logy=True)
# + papermill={"duration": 0.344886, "end_time": "2022-02-06T07:00:10.208103", "exception": false, "start_time": "2022-02-06T07:00:09.863217", "status": "completed"} tags=[]
pd.concat([i.pipe(Performance).summary() for i in strategy_3_result], axis=1)
# + papermill={"duration": 1.680779, "end_time": "2022-02-06T07:00:11.941789", "exception": false, "start_time": "2022-02-06T07:00:10.261010", "status": "completed"} tags=[]
fig, axes = plt.subplots(1, len(instruments), figsize=(14, 5), sharey=True)
for ax, result in zip(axes, strategy_3_result):
result.tail(252 * 2).pipe(rebase).plot(ax=ax, logy=True)
# + [markdown] papermill={"duration": 0.041783, "end_time": "2022-02-06T07:00:12.029677", "exception": false, "start_time": "2022-02-06T07:00:11.987894", "status": "completed"} tags=[]
# # Strategy 4
# Month-end strategy which goes long when the following conditions are met:
# - n days before month-end
# - (optional) only do so when the previous day is down
# - The price is above 200 day MA
# + papermill={"duration": 0.105776, "end_time": "2022-02-06T07:00:12.177520", "exception": false, "start_time": "2022-02-06T07:00:12.071744", "status": "completed"} tags=[]
def run_strat_4(instrument):
engine = BacktestEngine(
strategy=Strategy.DELTA_ONE.value,
instrument=instrument,
signal=signal.Pipeline([
signal.ParallelSignal([
signal.GreaterThanMA(lookback=200, shift=1),
signal.MonthEndLong(n_before=5, n_after=0, is_prev_neg=True),
], weighting='product', post_process=processing.CarryOver(5))
]),
log_level=logging.WARNING,
)
engine.run()
return engine
# + papermill={"duration": 1.00528, "end_time": "2022-02-06T07:00:13.225243", "exception": false, "start_time": "2022-02-06T07:00:12.219963", "status": "completed"} tags=[]
strategy_4_result = run_backtests(run_strat_4, instruments)
# + papermill={"duration": 1.632183, "end_time": "2022-02-06T07:00:14.900813", "exception": false, "start_time": "2022-02-06T07:00:13.268630", "status": "completed"} tags=[]
fig, axes = plt.subplots(1, len(instruments), figsize=(14, 5), sharey=True)
for ax, result in zip(axes, strategy_4_result):
result.plot(ax=ax, logy=True)
# + papermill={"duration": 0.337205, "end_time": "2022-02-06T07:00:15.282537", "exception": false, "start_time": "2022-02-06T07:00:14.945332", "status": "completed"} tags=[]
pd.concat([i.pipe(Performance).summary() for i in strategy_4_result], axis=1)
# + papermill={"duration": 1.693404, "end_time": "2022-02-06T07:00:17.024823", "exception": false, "start_time": "2022-02-06T07:00:15.331419", "status": "completed"} tags=[]
fig, axes = plt.subplots(1, len(instruments), figsize=(14, 5), sharey=True)
for ax, result in zip(axes, strategy_4_result):
result.tail(252 * 2).pipe(rebase).plot(ax=ax, logy=True)
# + [markdown] papermill={"duration": 0.060971, "end_time": "2022-02-06T07:00:17.132602", "exception": false, "start_time": "2022-02-06T07:00:17.071631", "status": "completed"} tags=[]
# # Reference
# - <NAME>. and <NAME>., 2009. Short Term Trading Strategies that Work: A Quantified Guide to Trading Stocks and ETFs. TradingMarkets Publishing Group.
# + papermill={"duration": 0.063203, "end_time": "2022-02-06T07:00:17.242241", "exception": false, "start_time": "2022-02-06T07:00:17.179038", "status": "completed"} tags=[]
print(f'Updated: {datetime.utcnow():%d-%b-%Y %H:%M}')
| equity_short_term_trading_connors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import os
os.getcwd()
path = r"C:\Users\Albert's Laptop\Documents\GitHub\course-project-group_1023\data\raw\weatherAUS.csv"
df=pd.read_csv(path)
df
| analysis/Albert/milestone1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # [Arranging Coins](https://leetcode.com/explore/featured/card/july-leetcoding-challenge/544/week-1-july-1st-july-7th/3377/)
# You have a total of n coins that you want to form in a staircase shape, where every k-th row must have exactly k coins.
#
# Given n, find the total number of full staircase rows that can be formed.
#
# n is a non-negative integer and fits within the range of a 32-bit signed integer.
#
# ***Example 1:***
#
# n = 5
#
# The coins can form the following rows:\
# ¤\
# ¤ ¤\
# ¤ ¤
#
# Because the 3rd row is incomplete, we return 2.
#
# ***Example 2:***
#
# n = 8
#
# The coins can form the following rows:\
# ¤\
# ¤ ¤\
# ¤ ¤ ¤\
# ¤ ¤
#
# Because the 4th row is incomplete, we return 3.
class Solution:
def arrangeCoins(self, n: int) -> int:
c=0
while True:
if (2*n)<=c*(c+1) :
if (2*n)!=c*(c+1):
c-=1
break
else:
c+=1
# print(c*(c+1))
return c
print(Solution().arrangeCoins(int(input())))
# # Ugly Number II
# ***Write a program to find the n-th ugly number.***
#
# Ugly numbers are positive numbers whose prime factors only include 2, 3, 5.
#
# **Example:**
#
# **Input:** n = 10\
# **Output:** 12\
# **Explanation:** 1, 2, 3, 4, 5, 6, 8, 9, 10, 12 is the sequence of the first 10 ugly numbers.\
# **Note:** \
# 1. 1 is typically treated as an ugly number.\
# 2. n does not exceed 1690.
class Solution:
def nthUglyNumber(self, n: int) -> int:
ugly=[0]*1690
ugly[0]=1
i2,i3,i5=[0]*3
next_2=2
next_3=3
next_5=5
for i in range(1,1690):
ugly[i]=min(next_2,next_3,next_5)
if ugly[i]==next_2:
i2+=1
next_2=ugly[i2]*2
if ugly[i]==next_3:
i3+=1
next_3=ugly[i3]*3
if ugly[i]==next_5:
i5+=1
next_5=ugly[i5]*5
return ugly[n-1]
s=Solution()
print(s.nthUglyNumber(int(input())))
# # Hamming Distance
# The Hamming distance between two integers is the number of positions at which the corresponding bits are different.
#
# Given two integers x and y, calculate the Hamming distance.
#
# **Note:**\
# 0 ≤ x, y < 231.
#
# **Example:**
#
# **Input:** x = 1, y = 4
#
# **Output:** 2
#
# **Explanation:**\
# 1 (0 0 0 1)\
# 4 (0 1 0 0)\
# ↑ ↑
#
# The above arrows point to positions where the corresponding bits are different.
bin(int(input())^int(input())).count('1')
float('1_1')
x=float(_)
x
float('1_1_1_1_1')
# # Plus One
# Given a non-empty array of digits representing a non-negative integer, plus one to the integer.
#
# The digits are stored such that the most significant digit is at the head of the list, and each element in the array contain a single digit.
#
# You may assume the integer does not contain any leading zero, except the number 0 itself.
#
# **Example 1:**
#
# **Input:** [1,2,3]\
# **Output:** [1,2,4]\
# **Explanation:** The array represents the integer 123.\
# **Example 2:**
#
# **Input:** [4,3,2,1]\
# **Output:** [4,3,2,2]\
# **Explanation:** The array represents the integer 4321.
a=list(map(int,input().split()))
l=list(str(int(''.join((map(str,a))))+1))
for i in range(len(l)):
l[i]=int(l[i])
print(l)
# # Island Perimeter
# You are given a map in form of a two-dimensional integer grid where 1 represents land and 0 represents water.
#
# Grid cells are connected horizontally/vertically (not diagonally). The grid is completely surrounded by water, and there is exactly one island (i.e., one or more connected land cells).
#
# The island doesn't have "lakes" (water inside that isn't connected to the water around the island). One cell is a square with side length 1. The grid is rectangular, width and height don't exceed 100. Determine the perimeter of the island.
#
#
#
# **Example:**
#
# **Input:**\
# [[0,1,0,0],\
# [1,1,1,0],\
# [0,1,0,0],\
# [1,1,0,0]]
#
# **Output:** 16
#
# **Explanation:** The perimeter is the 16 yellow stripes in the image below:
#
# 
class Solution:
def islandPerimeter(self, mat: List[List[int]]) -> int:
def numofneighbour(mat, i, j):
count = 0;
if (i > 0 and mat[i - 1][j]):
count+= 1;
if (j > 0 and mat[i][j - 1]):
count+= 1;
if (i < R-1 and mat[i + 1][j]):
count+= 1
if (j < C-1 and mat[i][j + 1]):
count+= 1;
return count;
def findperimeter(mat):
perimeter = 0;
for i in range(0, R):
for j in range(0, C):
if (mat[i][j]):
perimeter += (4 - numofneighbour(mat, i, j));
return perimeter;
R = len(mat)
C = len(mat[0])
return(findperimeter(mat));
# # 3Sum
#
# **Solution**\
# Given an array nums of n integers, are there elements a, b, c in nums such that a + b + c = 0? Find all unique triplets in the array which gives the sum of zero.
#
# **Note:**
#
# The solution set must not contain duplicate triplets.
#
# **Example:**
#
# Given array nums = [-1, 0, 1, 2, -1, -4],
#
# A solution set is:\
# [\
# [-1, 0, 1],\
# [-1, -1, 2]\
# ]
class Solution(object):
def threeSum(self, nums):
nums.sort()
result = []
for i in range(len(nums)-2):
if i> 0 and nums[i] == nums[i-1]:
continue
l = i+1
r = len(nums)-1
while(l<r):
sum = nums[i] + nums[l] + nums[r]
if sum<0:
l+=1
elif sum >0:
r-=1
else:
result.append([nums[i],nums[l],nums[r]])
while l<len(nums)-1 and nums[l] == nums[l + 1] : l += 1
while r>0 and nums[r] == nums[r - 1]: r -= 1
l+=1
r-=1
return result
ob1 = Solution()
print(ob1.threeSum([1,-1,-1,0]))
# # [Maximum Width of Binary Tree](https://leetcode.com/explore/featured/card/july-leetcoding-challenge/545/week-2-july-8th-july-14th/3385/)
# Given a binary tree, write a function to get the maximum width of the given tree. The width of a tree is the maximum width among all levels. The binary tree has the same structure as a full binary tree, but some nodes are null.
#
# The width of one level is defined as the length between the end-nodes (the leftmost and right most non-null nodes in the level, where the null nodes between the end-nodes are also counted into the length calculation.
#
# **Example 1:**
#
# **Input:**
#
# 1
# / \
# 3 2
# / \ \
# 5 3 9
#
# **Output:** 4\
# **Explanation:** The maximum width existing in the third level with the length 4 (5,3,null,9).\
# **Example 2:**
#
# **Input:**
#
# 1
# /
# 3
# / \
# 5 3
#
# **Output:** 2\
# **Explanation:** The maximum width existing in the third level with the length 2 (5,3).\
# **Example 3:**
#
# **Input:**
#
# 1
# / \
# 3 2
# /
# 5
#
# **Output:** 2\
# **Explanation:** The maximum width existing in the second level with the length 2 (3,2).\
# **Example 4:**
#
# **Input:**
#
# 1
# / \
# 3 2
# / \
# 5 9
# / \
# 6 7
#
# **Output:** 8\
# **Explanation:**The maximum width existing in the fourth level with the length 8 (6,null,null,null,null,null,null,7).
#
#
# **Note:** Answer will in the range of 32-bit signed integer.
# +
class Node:
def __init__(self,key):
self.right=None
self.left=None
self.val=key
def insert(root,node):
if root is None:
root=node
else:
if root.val<node.val:
if root.right is None:
root.right=node
else:
insert(root.right,node)
else:
if root.left is None:
root.left=node
else:
insert(root.left,node)
h=0
l1=[]
def inorder(root,h):
if root:
inorder(root.left,h+1)
print(root.val,h,sep=' --> ')
l1.append(h)
inorder(root.right,h+1)
n=int(input())
a=list(map(int,input().split()))
for i in range(n):
if i==0:
r=Node(a[i])
else:
insert(r,Node(a[i]))
inorder(r,1)
print(l1.count(max(l1))*2)
# -
# Definition for a binary tree node.
# class TreeNode:
# def __init__(self, val=0, left=None, right=None):
# self.val = val
# self.left = left
# self.right = right
class Solution:
def widthOfBinaryTree(self, root: TreeNode) -> int:
if root is None: return 0
result=1
Queue=[[root,0]]
while len(Queue)>0:
count=len(Queue)
start=Queue[0][1]
end=Queue[-1][0]
result=max(result,end-start+1)
for i in range(count):
p=Queue[0]
idx=p[1]-satrt
Queue.pop()
if p[0].left: Queue.append([p[0].left,2*idx+1])
if p[0].right: Queue.append([p[0].right,2*idx+2])
return result
# # [Subsets](https://leetcode.com/explore/challenge/card/july-leetcoding-challenge/545/week-2-july-8th-july-14th/3387/)
#
# <pre>
# Given a set of distinct integers, nums, return all possible subsets (the power set).
#
# Note: The solution set must not contain duplicate subsets.
#
# <b>Example:</b>
#
# <b>Input:</b> nums = [1,2,3]
# <b>Output:</b>
# [
# [3],
# [1],
# [2],
# [1,2,3],
# [1,3],
# [2,3],
# [1,2],
# []
# ]
# </pre>
l=[]
a=list(map(int,input().split()))
l.append(a.copy())
print(l)
while a:
k=a.pop()
if [k] not in l:
l.append([k])
if a not in l:
l.append(a.copy())
for i in a:
if [i,k] not in l:
l.append([i,k])
l
# +
import itertools as it
def combinations_all(l,i):
return list(map(list, itertools.combinations(l, i)))
a=list(map(int,input().split()))
l=[]
for i in range(len(a)+1):
l.append(combinations_all(a,i))
print(l)
# -
class Solution(object):
def subsets(self, nums):
temp_result = []
self.subsets_util(nums,[0 for i in range(len(nums))],temp_result,0)
main_result = []
for lists in temp_result:
temp = []
for i in range(len(lists)):
if lists[i] == 1:
temp.append(nums[i])
main_result.append(temp)
return main_result
def subsets_util(self,nums,temp,result,index):
if index == len(nums):
result.append([i for i in temp])
#print(temp)
return
temp[index] = 0
self.subsets_util(nums,temp,result,index+1)
temp[index] = 1
self.subsets_util(nums, temp, result,index + 1)
ob1 = Solution()
print(ob1.subsets([1,2,3,4]))
# # [Reverse Bits](https://leetcode.com/explore/challenge/card/july-leetcoding-challenge/545/week-2-july-8th-july-14th/3388/)
#
# <pre>
# Reverse bits of a given 32 bits unsigned integer.
#
#
#
# Example 1:
#
# Input: 00000010100101000001111010011100
# Output: 00111001011110000010100101000000
# Explanation: The input binary string 00000010100101000001111010011100 represents the unsigned integer 43261596, so return 964176192 which its binary representation is 00111001011110000010100101000000.
# Example 2:
#
# Input: 11111111111111111111111111111101
# Output: 10111111111111111111111111111111
# Explanation: The input binary string 11111111111111111111111111111101 represents the unsigned integer 4294967293, so return 3221225471 which its binary representation is 10111111111111111111111111111111.
#
#
# Note:
#
# Note that in some languages such as Java, there is no unsigned integer type. In this case, both input and output will be given as signed integer type and should not affect your implementation, as the internal binary representation of the integer is the same whether it is signed or unsigned.
# In Java, the compiler represents the signed integers using 2's complement notation. Therefore, in Example 2 above the input represents the signed integer -3 and the output represents the signed integer -1073741825.
#
#
# Follow up:
#
# If this function is called many times, how would you optimize it?
# </pre>
x=bin(int(input()))[2:]
int(('0'*(32-len(x))+x)[::-1],2)
class Solution:
def reverseBits(self, n: int) -> int:
x=bin(n)[2:]
return int(('0'*(32-len(x))+x)[::-1],2)
print(Solution().reverseBits(int(input())))
# # WEEK 3
# ### [Reverse Words in a String](https://leetcode.com/explore/challenge/card/july-leetcoding-challenge/546/week-3-july-15th-july-21st/3391/)
s=input()
' '.join(s.strip().split()[::-1])
class Solution:
def reverseWords(self, s: str) -> str:
return ' '.join(s.strip().split()[::-1])
print(Solution().reverseWords(input()))
# ### [ Pow(x, n)](https://leetcode.com/explore/challenge/card/july-leetcoding-challenge/546/week-3-july-15th-july-21st/3392/)
#
# <pre>
# Implement pow(x, n), which calculates x raised to the power n (xn).
#
# Example 1:
#
# Input: 2.00000, 10
# Output: 1024.00000
# Example 2:
#
# Input: 2.10000, 3
# Output: 9.26100
# Example 3:
#
# Input: 2.00000, -2
# Output: 0.25000
# Explanation: 2-2 = 1/22 = 1/4 = 0.25
# Note:
#
# -100.0 < x < 100.0
# n is a 32-bit signed integer, within the range [−231, 231 − 1]
# </pre>
2.00000**10
# ### [Add Binary](https://leetcode.com/explore/challenge/card/july-leetcoding-challenge/546/week-3-july-15th-july-21st/3395/)
#
# <pre>
# Given two binary strings, return their sum (also a binary string).
#
# The input strings are both non-empty and contains only characters 1 or 0.
#
# Example 1:
#
# Input: a = "11", b = "1"
# Output: "100"
# Example 2:
#
# Input: a = "1010", b = "1011"
# Output: "10101"
#
#
# Constraints:
#
# Each string consists only of '0' or '1' characters.
# 1 <= a.length, b.length <= 10^4
# Each string is either "0" or doesn't contain any leading zero.
# </pre>
bin(int('1010',2)+int('1011',2))[2:]
# ### [ Remove Linked List Elements](https://leetcode.com/explore/challenge/card/july-leetcoding-challenge/546/week-3-july-15th-july-21st/3396/)
# +
class node:
def __init__(self,val):
self.data=val
self.next=None
class LinkedList:
def __init__(self):
self.start=None
def add_data(self):
for i in range(int(input())):
if self.start is None:
self.start=node(int(input()))
ptr=self.start
else:
ptr.next=node(int(input()))
ptr=ptr.next
return self.start
def display(self,head):
ptr=head
while ptr:
print(ptr.data,end=' ')
ptr=ptr.next
l=LinkedList()
head=l.add_data()
l.display(head)
# -
def remove_data(root):
data=int(input())
l=LinkedList()
while root:
if root.data!=data:
if l.start is None:
l.start=node(root.data)
ptr=l.start
else:
ptr.next=node(root.data)
ptr=ptr.next
root=root.next
return l.start
head1=remove_data(head)
l.display(head1)
l.display(head)
def getNode(head, p):
l=[]
while head:
l.append(head.data)
head=head.next
return l[-p-1]
getNode(head,2)
def getNode(head, p):
tracked=head
while head:
if p>-1:
p-=1
else:
tracked=tracked.next
head=head.next
return tracked.data
def has_cycle(head):
l=[]
while head:
if head.data in l:
return True
else:
l.append(head.data)
head=head.next
return False
has_cycle(head)
has_cycle(head1)
# ### [Climbing Stairs](https://leetcode.com/explore/challenge/card/july-leetcoding-challenge/548/week-5-july-29th-july-31st/3407/)
n=int(input())
c_2=n//2
c_1=n%2
sum_p=1
while c_2:
sum_p+=get_p(c_2+c_1,c_2,c_1)
c_2-=1
c_1+=2
print(sum_p)
def fact(n):
if n==0:
return 1
return n*fact(n-1)
def get_p(n,n1,n2):
return fact(n)//(fact(n1)*fact(n2))
fact(4)
get_p(3,1,2)
| LeetCode/July LeetCoding Challenge 2020/July_LeetCoding_Challenge_2020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# language: python
# name: python3
# ---
import csv
from enum import Enum
from typing import List
# +
class PlaceType(Enum):
study="STUDY"
classroom="CLASSROOM"
dining="DINING"
class Place(object):
def __init__(self, nodeId: int, place_type: PlaceType):
self.nodeId = nodeId
self.place_type = place_type
def sql_insert(self) -> str:
return f'''INSERT INTO public."Place"(
"NodeID", "PlaceType")
VALUES ({self.nodeId}, '{self.place_type.value}');'''
# +
places: List[Place] = []
with open('classroom-dining-study.csv') as csvfile:
node_classification_reader = csv.reader(csvfile, delimiter=',')
next(node_classification_reader)
for row in node_classification_reader:
nodeId = row[0]
place_type_raw = row[1].strip()
parsedPlace = Place(int(nodeId), PlaceType[place_type_raw])
places.append(parsedPlace)
# -
for place in places:
print(place.sql_insert())
| capybara-import-data/node-classifcation/place-insert-generator copy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] Collapsed="false"
# <img src='./img/LogoWekeo_Copernicus_RGB_0.png' alt='Logo EU Copernicus EUMETSAT' align='right' width='20%'></img>
# + [markdown] Collapsed="false"
# <br>
# + [markdown] Collapsed="false"
# # Functions
# -
# This notebook lists all `functions` that are defined and used throughout the `LTPy course`.
# The following functions are listed:
#
# **[Data loading and re-shaping functions](#load_reshape)**
# * [generate_geographical_subset](#generate_geographical_subset)
# * [select_channels_for_rgb](#rgb_channels)
# * [normalize](#normalize)
#
# **[Data visualization functions](#visualization)**
# * [visualize_pcolormesh](#visualize_pcolormesh)
# * [visualize_s3_pcolormesh](#visualize_s3_pcolormesh)
#
# <hr>
# #### Load required libraries
# + Collapsed="false"
import os
from matplotlib import pyplot as plt
import xarray as xr
from netCDF4 import Dataset
import numpy as np
import glob
from matplotlib import pyplot as plt
import matplotlib.colors
from matplotlib.colors import LogNorm
import cartopy.crs as ccrs
from cartopy.mpl.gridliner import LONGITUDE_FORMATTER, LATITUDE_FORMATTER
import cartopy.feature as cfeature
import warnings
warnings.simplefilter(action = "ignore", category = RuntimeWarning)
warnings.simplefilter(action = "ignore", category = FutureWarning)
# -
# <hr>
# ## <a id="load_reshape"></a>Data loading and re-shaping functions
# + [markdown] Collapsed="false"
# ### <a id='generate_geographical_subset'></a>`generate_geographical_subset`
# + Collapsed="false"
def generate_geographical_subset(xarray, latmin, latmax, lonmin, lonmax):
"""
Generates a geographical subset of a xarray DataArray and shifts the longitude grid from a 0-360 to a -180 to 180 deg grid.
Parameters:
xarray (xarray DataArray): a xarray DataArray with latitude and longitude coordinates
latmin, latmax, lonmin, lonmax (int): boundaries of the geographical subset
Returns:
Geographical subset of a xarray DataArray.
"""
return xarray.where((xarray.latitude < latmax) & (xarray.latitude > latmin) & (xarray.longitude < lonmax) & (xarray.longitude > lonmin),drop=True)
# -
# ### <a id='rgb_channels'></a> `select_channels_for_rgb`
def select_channels_for_rgb(xarray, red_channel, green_channel, blue_channel):
"""
Selects the channels / bands of a multi-dimensional xarray for red, green and blue composites.
Parameters:
xarray(xarray Dataset): xarray Dataset object that stores the different channels / bands.
red_channel(str): Name of red channel to be selected
green_channel(str): Name of green channel to be selected
blue_channel(str): Name of blue channel to be selected
Returns:
Three xarray DataArray objects with selected channels / bands
"""
return xarray[red_channel], xarray[green_channel], xarray[blue_channel]
# ### <a id='normalize'></a> `normalize`
def normalize(array):
"""
Normalizes a numpy array / xarray DataArray object value to values between 0 and 1.
Parameters:
xarray(numpy array or xarray DataArray): xarray DataArray or numpy array object.
Returns:
Normalized array
"""
array_min, array_max = array.min(), array.max()
return ((array - array_min)/(array_max - array_min))
# <hr>
# ## <a id="visualization"></a>Data visualization functions
# + [markdown] Collapsed="false"
# ### <a id='visualize_pcolormesh'></a>`visualize_pcolormesh`
# + Collapsed="false"
def visualize_pcolormesh(data_array, longitude, latitude, projection, color_scale, unit, long_name, vmin, vmax, lonmin, lonmax, latmin, latmax, log=True, set_global=True):
"""
Visualizes a numpy array with matplotlib's 'pcolormesh' function.
Parameters:
data_array: any numpy MaskedArray, e.g. loaded with the NetCDF library and the Dataset function
longitude: numpy Array holding longitude information
latitude: numpy Array holding latitude information
projection: a projection provided by the cartopy library, e.g. ccrs.PlateCarree()
color_scale (str): string taken from matplotlib's color ramp reference
unit (str): the unit of the parameter, taken from the NetCDF file if possible
long_name (str): long name of the parameter, taken from the NetCDF file if possible
vmin (int): minimum number on visualisation legend
vmax (int): maximum number on visualisation legend
lonmin,lonmax,latmin,latmax: geographic extent of the plot
log (logical): set True, if the values shall be represented in a logarithmic scale
set_global (logical): set True, if the plot shall have a global coverage
"""
fig=plt.figure(figsize=(20, 10))
ax = plt.axes(projection=projection)
# define the coordinate system that the grid lons and grid lats are on
if(log):
img = plt.pcolormesh(longitude, latitude, np.squeeze(data_array), norm=LogNorm(),
cmap=plt.get_cmap(color_scale), transform=ccrs.PlateCarree(),
vmin=vmin,
vmax=vmax)
else:
img = plt.pcolormesh(longitude, latitude, data_array,
cmap=plt.get_cmap(color_scale), transform=ccrs.PlateCarree(),
vmin=vmin,
vmax=vmax)
ax.add_feature(cfeature.BORDERS, edgecolor='black', linewidth=1)
ax.add_feature(cfeature.COASTLINE, edgecolor='black', linewidth=1)
if (projection==ccrs.PlateCarree()):
ax.set_extent([lonmin, lonmax, latmin, latmax], projection)
gl = ax.gridlines(draw_labels=True, linestyle='--')
gl.xformatter=LONGITUDE_FORMATTER
gl.yformatter=LATITUDE_FORMATTER
gl.xlabel_style={'size':14}
gl.ylabel_style={'size':14}
if(set_global):
ax.set_global()
ax.gridlines()
cbar = fig.colorbar(img, ax=ax, orientation='horizontal', fraction=0.04, pad=0.1)
cbar.set_label(unit, fontsize=16)
cbar.ax.tick_params(labelsize=14)
ax.set_title(long_name, fontsize=20, pad=40.0)
# plt.show()
return fig, ax
# -
# ### <a id='visualize_s3_pcolormesh'></a>`visualize_s3_pcolormesh`
def visualize_s3_pcolormesh(color_array, array, latitude, longitude, title):
"""
Visualizes a numpy array (Sentinel-3 data) with matplotlib's 'pcolormesh' function as RGB image.
Parameters:
color_array (numpy MaskedArray): any numpy MaskedArray, e.g. loaded with the NetCDF library and the Dataset function
array (xarray.DataArray): in order to get the array dimensions
longitude (numpy Array): array with longitude values
latitude (numpy Array) : array with latitude values
title (str): title of the resulting plot
"""
fig=plt.figure(figsize=(20, 12))
ax=plt.axes(projection=ccrs.Mercator())
ax.coastlines()
gl = ax.gridlines(draw_labels=True, linestyle='--')
gl.xformatter=LONGITUDE_FORMATTER
gl.yformatter=LATITUDE_FORMATTER
gl.xlabel_style={'size':14}
gl.ylabel_style={'size':14}
img1 = plt.pcolormesh(longitude, latitude, array*np.nan, color=color_array,
clip_on = True,
edgecolors=None,
zorder=0,
transform=ccrs.PlateCarree())
ax.set_title(title, fontsize=20, pad=40.0)
plt.show()
# + [markdown] Collapsed="false"
# <hr>
# + [markdown] Collapsed="false"
# <img src='./img/all_partners_wekeo.png' alt='Logo EU Copernicus EUMETSAT' align='right' width='100%'></img>
| atmosphere/functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Example: How to use me-types-mapper
import numpy as np
import pandas as pd
# # Create toy datasets
# +
from sklearn.datasets import make_blobs
N_samples_1, N_types_1 = 100, 6
N_samples_2, N_types_2 = 700, 14
N_features = 200
N_efeatures = 70
# Create datasets using make_blobs from sklearn
dataset_1, labels_1 = make_blobs(n_samples=N_samples_1, centers=N_types_1, n_features=N_features,
cluster_std=3.0, center_box=(- 15.0, 15.0), )
dataset_2, labels_2 = make_blobs(n_samples=N_samples_2, centers=N_types_2, n_features=N_features,
cluster_std=10.0, center_box=(- 10.0, 10.0), )
# Convert into panda DataFrame (needed to use me-types-mapper)
dataset_1 = pd.DataFrame(dataset_1, index = [np.float("100" + str(i)) for i in range(len(labels_1))])
dataset_2 = pd.DataFrame(dataset_2, index = ["dsetB_sample" + str(i) for i in range(len(labels_2))])
# Give cluster labels more explicit names
labels_1 = pd.DataFrame(["dsetA_clstr" + str(x) for x in labels_1],
index = dataset_1.index)
labels_2 = pd.DataFrame(["dsetB_clstr" + str(x) for x in labels_2],
index = dataset_2.index)
labels = pd.concat([labels_1, labels_2], axis=0)[0]
# Create a mask to separate features into two subtypes (e.g. electorphysiological and morphological)
msk_efeat = np.asarray([True] * N_efeatures + [False] * (N_features - N_efeatures))
# -
# # Visualize dataset using PCA
# +
from sklearn.decomposition import PCA
import matplotlib.pyplot as plt
# %matplotlib inline
def plot_pca(X, labels, title):
pca = PCA(n_components=2)
embedding = pca.fit_transform(X)
for lbl in labels.unique():
msk = labels == lbl
plt.scatter(embedding[:,0][msk], embedding[:,1][msk], alpha=0.5)
plt.legend(labels.unique())
plt.xlabel("PC_1")
plt.ylabel("PC_2")
plt.title(title)
plt.show()
return
data = pd.concat([dataset_1, dataset_2], axis=0)
lbls_dset = pd.DataFrame(["dataset A"] * len(labels_1) + ["dataset B"] * len(labels_2))[0]
plot_pca(data, lbls_dset, "PCA")
plot_pca(data, labels, "")
# -
# # Define common clusters using me-types-mapper
# +
from me_types_mapper.mapper.coclustering_functions import cross_predictions_v2
(alpha_opt, map_, c1, c2,
dict_cluster_label, cluster_ref, fig_alpha, fig_d_opt) = cross_predictions_v2(dataset_1, dataset_2,
msk_efeat, ~msk_efeat, labels,
alpha_list_ = np.arange(.1,1.,.1))
# -
# # Probabilistic mapping
# +
# Prepare a dictionnary with sample index from both dataset as keys
# and their attached common cluster (defined above) as values.
cell_id_list = dataset_1.index.tolist() + dataset_2.index.tolist()
cluster_ref_dict = {}
for cell_idx, cell_id in enumerate(cell_id_list):
cluster_ref_dict[cell_id] = cluster_ref[cell_idx]
# Masks to split data from both datasets.
mask_tmp_1 = np.asarray([True] * len(labels_1) + [False] * len(labels_2))
mask_tmp_2 = ~mask_tmp_1
# probabilistic mapping
from me_types_mapper.mapper.coclustering_functions import compute_probabilistic_maps
p_maps = compute_probabilistic_maps(labels_2, 0, mask_tmp_2,
labels_1, 0, mask_tmp_1,
cluster_ref_dict)
# -
# # Plot the probabilistic map
# +
import matplotlib.pyplot as plt
# %matplotlib inline
def plot_matrix(ax, X , axis_labels=True):
X = X.div(np.sum(X, axis=0), axis=1).fillna(0)
ax.imshow(X, vmin=0., vmax=1., cmap='Greys')
if axis_labels:
ax.set_xticks(np.arange(len(X.columns)))
ax.set_xticklabels(X.columns, rotation=90)
ax.set_yticks(np.arange(len(X.index)))
ax.set_yticklabels(X.index)
else:
ax.set_xticks([])
ax.set_yticks([])
# Loop over data dimensions and create text annotations.
for i in range(len(X.index)):
for j in range(len(X.columns)):
if (X.values[i, j] > 0.5):
text = ax.text(j, i, int(X.values[i, j] * 100),
ha="center", va="center", color="w")
elif (0.1 < X.values[i, j] <= 0.5):
text = ax.text(j, i, int(X.values[i, j] * 100),
ha="center", va="center", color="k")
return
fig, ax = plt.subplots(1, figsize=(4,4))
plot_matrix(ax, p_maps[-1], axis_labels=True)
ax.set_title("P(cluster_datasetA|cluster_datasetB)")
fig2, ax2 = plt.subplots(1, figsize=(4,4))
plot_matrix(ax2, p_maps[-2], axis_labels=True)
ax2.set_title("P(cluster_datasetB|cluster_datasetA)")
# -
| Examples/example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import seaborn as sns
import scipy as sp
# %matplotlib inline
plt.rcParams['figure.figsize'] = [15, 5]
# !ls ../input_data/
train = pd.read_csv('../input_data/train.csv')
test = pd.read_csv('../input_data/test.csv')
# # Feature engineering
# +
def preprocess(T):
T = T.set_index('PassengerId')
# Create a marker for missing data
T['UnknownCabin'] = T['Cabin'].isna().astype(int)
T['UnknownAge'] = T['Age'].isna().astype(int)
T['Sp-Pa'] = T['SibSp'] - T['Parch']
# Define fare categories
T['FareCat'] = 1
T.loc[T['Fare'] <= 10.0, 'FareCat'] = 0
T.loc[T['Fare'] >= 50.0, 'FareCat'] = 2
T['AgeCat'] = 0 # Children and babies
T.loc[T['Age'] >= 10.0, 'AgeCat'] = 1 # Adult
# Estimate Age category based on Title
# Mr & Mrs ... are adults
# Miss & Master ... are children
# All the rest should be adults too
T['Title'] = T['Name'].str.split().apply(lambda name: name[1])
T['AgeCatByTitle'] = 1;
T.loc[T['Title'].isin(['Miss.', 'Master.']), 'AgeCatByTitle'] = 0
# For missing entries overwrite AgeCat
T.loc[T['UnknownAge'].astype(bool), 'AgeCat'] = T.loc[T['UnknownAge'].astype(bool), 'AgeCatByTitle']
# Convert to easy to process values
# 0 ... female, 1 ... male
T['ppSex'] = (T['Sex'] == 'male').astype(int)
T['ppEmbarked'] = T['Embarked'].astype('category').cat.codes
if 'Survived' in T.columns:
# Split depended and indepened
Y = pd.DataFrame(T['Survived'])
T.drop('Survived', axis=1, inplace=True)
else:
Y = pd.DataFrame((T['Age']*np.nan).rename('Survived'))
# Only keep some features
keep = ['Pclass', 'AgeCat', 'Sp-Pa', 'SibSp', 'Parch', 'FareCat', 'ppEmbarked', 'UnknownCabin', 'UnknownAge', 'ppSex']
return T[keep], Y
# -
Xtrain, Ytrain = preprocess(train)
Xtest, Ytest = preprocess(test)
print("Number of missing values")
pd.DataFrame(Xtrain.isna().sum(axis=0)).T
pd.DataFrame(Xtest.isna().sum(axis=0)).T
# # Storing results
# write results to a hdf file in ../data
from pandas import HDFStore
with HDFStore('../data/processed.h5', mode='w') as hdf:
hdf.put('Xtrain', Xtrain)
hdf.put('Ytrain', Ytrain)
hdf.put('Xtest', Xtest)
hdf.put('Ytest', Ytest)
| titanic/nb/1 - Data Preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
from __future__ import annotations
import numpy as np
from numpy.linalg import inv, det, slogdet
class UnivariateGaussian:
"""
Class for univariate Gaussian Distribution Estimator
"""
def __init__(self, biased_var: bool = False) -> UnivariateGaussian:
"""
Estimator for univariate Gaussian mean and variance parameters
Parameters
----------
biased_var : bool, default=True
Should fitted estimator of variance be a biased or unbiased estimator
Attributes
----------
fitted_ : bool
Initialized as false indicating current estimator instance has not been fitted.
To be set as True in `UnivariateGaussian.fit` function.
mu_: float
Estimated expectation initialized as None. To be set in `UnivariateGaussian.fit`
function.
var_: float
Estimated variance initialized as None. To be set in `UnivariateGaussian.fit`
function.
"""
self.biased_ = biased_var
self.fitted_, self.mu_, self.var_ = False, None, None
def fit(self, X: np.ndarray) -> UnivariateGaussian:
"""
Estimate Gaussian expectation and variance from given samples
Parameters
----------
X: ndarray of shape (n_samples, )
Training data
Returns
-------
self : returns an instance of self.
Notes
-----
Sets `self.mu_`, `self.var_` attributes according to calculated estimation (where
estimator is either biased or unbiased). Then sets `self.fitted_` attribute to `True`
"""
raise NotImplementedError()
self.fitted_ = True
return self
def pdf(self, X: np.ndarray) -> np.ndarray:
"""
Calculate PDF of observations under Gaussian model with fitted estimators
Parameters
----------
X: ndarray of shape (n_samples, )
Samples to calculate PDF for
Returns
-------
pdfs: ndarray of shape (n_samples, )
Calculated values of given samples for PDF function of N(mu_, var_)
Raises
------
ValueError: In case function was called prior fitting the model
"""
if not self.fitted_:
raise ValueError("Estimator must first be fitted before calling `pdf` function")
raise NotImplementedError()
@staticmethod
def log_likelihood(mu: float, sigma: float, X: np.ndarray) -> float:
"""
Calculate the log-likelihood of the data under a specified Gaussian model
Parameters
----------
mu : float
Expectation of Gaussian
sigma : float
Variance of Gaussian
X : ndarray of shape (n_samples, )
Samples to calculate log-likelihood with
Returns
-------
log_likelihood: float
log-likelihood calculated
"""
raise NotImplementedError()
class MultivariateGaussian:
"""
Class for multivariate Gaussian Distribution Estimator
"""
def __init__(self):
"""
Initialize an instance of multivariate Gaussian estimator
Attributes
----------
fitted_ : bool
Initialized as false indicating current estimator instance has not been fitted.
To be set as True in `MultivariateGaussian.fit` function.
mu_: float
Estimated expectation initialized as None. To be set in `MultivariateGaussian.ft`
function.
cov_: float
Estimated covariance initialized as None. To be set in `MultivariateGaussian.ft`
function.
"""
self.mu_, self.cov_ = None, None
self.fitted_ = False
def fit(self, X: np.ndarray) -> MultivariateGaussian:
"""
Estimate Gaussian expectation and covariance from given samples
Parameters
----------
X: ndarray of shape (n_samples, )
Training data
Returns
-------
self : returns an instance of self.
Notes
-----
Sets `self.mu_`, `self.cov_` attributes according to calculated estimation.
Then sets `self.fitted_` attribute to `True`
"""
raise NotImplementedError()
self.fitted_ = True
return self
def pdf(self, X: np.ndarray):
"""
Calculate PDF of observations under Gaussian model with fitted estimators
Parameters
----------
X: ndarray of shape (n_samples, )
Samples to calculate PDF for
Returns
-------
pdfs: ndarray of shape (n_samples, )
Calculated values of given samples for PDF function of N(mu_, cov_)
Raises
------
ValueError: In case function was called prior fitting the model
"""
if not self.fitted_:
raise ValueError("Estimator must first be fitted before calling `pdf` function")
raise NotImplementedError()
@staticmethod
def log_likelihood(mu: np.ndarray, cov: np.ndarray, X: np.ndarray) -> float:
"""
Calculate the log-likelihood of the data under a specified Gaussian model
Parameters
----------
mu : float
Expectation of Gaussian
cov : float
covariance matrix of Gaussian
X : ndarray of shape (n_samples, )
Samples to calculate log-likelihood with
Returns
-------
log_likelihood: float
log-likelihood calculated
"""
raise NotImplementedError()
# -
| lab/ex 1/.ipynb_checkpoints/pyKernel-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Exercise: price list
# Imports:
import requests
from bs4 import BeautifulSoup
import time # for sleeping between multiple requests
headers = {'user-agent': 'scrapingCourseBot'}
# #### Documentation:
# - [Requests.py](http://docs.python-requests.org)
# - [Beautifulsoup.py](https://www.crummy.com/software/BeautifulSoup/bs4/doc/)
r = requests.get('http://testing-ground.webscraping.pro/price-list-1.html', headers=headers)
print(r.status_code)
soup = BeautifulSoup(r.text, 'lxml')
#print(soup)
# Retrieve the first product name:
result = soup.find("div", class_="name").text
print(result)
# Print all product names contained in the page (both case 1 and 2)
names = soup.find_all("div", class_="name")
for name in names:
print(name.text)
# Print all product names from Case1:
names = soup.find("div", id="case1").find_all("div", class_="name")
for name in names:
print(name.text)
# Option: print all product names + prices, Case1:
names = soup.find("div", id="case1").find_all("div", class_="name")
for name in names:
price = name.parent.next_sibling
print(name.text, price.text)
| 20210913/Answers/Price_List.ipynb |