code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py3.6-env
# language: python
# name: py3.6-env
# ---
# +
# %matplotlib inline
import gym
import matplotlib
import numpy as np
import sys
from collections import defaultdict
from envs.blackjack import BlackjackEnv
import plotting
matplotlib.style.use('ggplot')
# -
env = BlackjackEnv()
def make_epsilon_greedy_policy(Q, epsilon, nA):
"""
给定一个Q函数和epsilon,构建一个ε-贪婪的策略
参数:
Q: 一个dictionary其key-value是state -> action-values.
key是状态s,value是一个长为nA(Action个数)的numpy数组,表示采取行为a的概率。
epsilon: float
nA: action的个数
返回值:
返回一个 函数,这个函数的输入是一个状态/观察(observation),输出是一个长度为nA的numpy数组,表示采取不同Action的概率
"""
def policy_fn(observation):
A = np.ones(nA, dtype=float) * epsilon / nA
best_action = np.argmax(Q[observation])
A[best_action] += (1.0 - epsilon)
return A
return policy_fn
def mc_control_epsilon_greedy(env, num_episodes, discount_factor=1.0, epsilon=0.1):
"""
使用Epsilon-贪婪策略的蒙特卡罗控制,用了找到最优的epsilon-greedy策略
参数:
env: OpenAI gym environment
num_episodes: 采样的episode个数
discount_factor: 打折因子
epsilon: Float
返回:
一个tuple(Q, policy).
Q函数 state -> action values。key是状态,value是长为nA的numpy数组,表示Q(s,a)
policy 最优的策略函数,输入是状态,输出是nA长的numpy数组,表示采取不同action的概率
"""
# 记录每个状态的回报累加值和次数
returns_sum = defaultdict(float)
returns_count = defaultdict(float)
# Q函数state -> (action -> action-value)。key是状态s,value又是一个dict,其key是a,value是Q(s,a)
Q = defaultdict(lambda: np.zeros(env.action_space.n))
# epsilon-贪婪的策略
policy = make_epsilon_greedy_policy(Q, epsilon, env.action_space.n)
for i_episode in range(1, num_episodes + 1):
if i_episode % 1000 == 0:
print("\rEpisode {}/{}.".format(i_episode, num_episodes), end="")
sys.stdout.flush()
# 生成一个episode。
# 一个episode是一个数组,每个元素是一个三元组(state, action, reward)
episode = []
state = env.reset()
for t in range(100):
probs = policy(state)
action = np.random.choice(np.arange(len(probs)), p=probs)
next_state, reward, done, _ = env.step(action)
episode.append((state, action, reward))
if done:
break
state = next_state
# 找到episode里出现的所有(s,a)对epsilon。把它变成tuple以便作为dict的key。
sa_in_episode = set([(tuple(x[0]), x[1]) for x in episode])
for state, action in sa_in_episode:
sa_pair = (state, action)
# 找到(s,a)第一次出现的下标
first_occurence_idx = next(i for i,x in enumerate(episode)
if x[0] == state and x[1] == action)
# 计算(s,a)的回报
G = sum([x[2]*(discount_factor**i) for i,x in enumerate(episode[first_occurence_idx:])])
# 累计计数
returns_sum[sa_pair] += G
returns_count[sa_pair] += 1.0
Q[state][action] = returns_sum[sa_pair] / returns_count[sa_pair]
# 策略已经通过Q“隐式”的提高了!
return Q, policy
Q, policy = mc_control_epsilon_greedy(env, num_episodes=500000, epsilon=0.1)
V = defaultdict(float)
for state, actions in Q.items():
action_value = np.max(actions)
V[state] = action_value
plotting.plot_value_function(V, title="Optimal Value Function")
| rl/On-Policy MC Control.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Gaussian XOR and Gaussian R-XOR Experiment with Task Unaware Settings
# +
# import dependencies
import numpy as np
import random
from proglearn.sims import generate_gaussian_parity
import matplotlib.pyplot as plt
import seaborn as sns
# functions to perform the experiments in this notebook
import functions.xor_rxor_with_unaware_fns as fn
# -
# ## Ksample test
# Using ksample test from hyppo we can determine at which angle rxor is significantly different enough from xor to require a new task/transformer. The following code using functions from xor_rxor_aware_unaware_fns.py to calculate p-values from k sample test dcorr from rxor angles 0 to 90 degrees for sample sizes 100, 500, and 1000.
# number of times to run the experiment, decrease for shorter run times
mc_rep = 10
# set angle range
angle_sweep = range(0, 90, 1)
# calculates and plots angle vs pvalue from ksample test xor vs rxor
# returns numpy array containing all p-vals from all mc_rep experiments at all angles in angle_sweep
pvals_100, pvals_500, pvals_1000 = fn.calc_ksample_pval_vs_angle(mc_rep, angle_sweep)
# sets plotting params
sns.set_context("talk")
# plots the mean p-values of the mc_rep experiments with error bars
# dotted green line at p-value = 0.05
plt.figure(figsize=(8, 8))
fn.plot_pval_vs_angle(pvals_100, pvals_500, pvals_1000, angle_sweep)
# ## Task aware BTE and generalization error (XOR)
# Next, we'll run the progressive learner to see how different angles of rxor affect backward transfer efficiency and multitask generalization error of xor (task1). We start by defining the following hyperparameters.
# number of times to run the experiment, decrease for shorter run times
mc_rep = 100
# samples to use for task1 (xor)
task1_sample = 100
# samples to use for task2 (rxor)
task2_sample = 100
# we will use the same angle_sweep as before
angle_sweep = range(0, 90, 1)
# call the function to run the experiment
# give us arrays with mean_te and mean_error
mean_te, mean_error = fn.bte_ge_v_angle(angle_sweep, task1_sample, task2_sample, mc_rep)
# plot angle vs BTE
fn.plot_bte_v_angle(mean_te)
# plot angle vs generalization error
plt.figure(figsize=(8, 8))
plt.plot(angle_sweep, mean_error[:, 1])
plt.xlabel("Angle of Rotation")
plt.ylabel("Generalization Error (xor)")
plt.show()
# ## Task Unaware: K-sample testing "dcorr"
# Instead of adding a new task for every angle of rxor, we use a k sample test to determine when rxor is different enough to warrant adding a new task. Then we plot the BTE and multitask generalization error of xor (task1). Once again, we start by definining hyperparameters. We will examine BTE and generalization error for 100, 500, and 1000 task samples.
# ### 100 task samples
# number of times to run the experiment, decrease for shorter run times
mc_rep = 100
# samples to use for task1 (xor)
task1_sample = 100
# samples to use for task2 (rxor)
task2_sample = 100
# we will use the same angle_sweep as before
angle_sweep = range(0, 90, 1)
# call our function to run the experiment
un_mean_te, un_mean_error = fn.unaware_bte_v_angle(
angle_sweep, task1_sample, task2_sample, mc_rep
)
# plot angle vs BTE
fn.plot_unaware_bte_v_angle(un_mean_te)
# plot angle vs generalization error
plt.figure(figsize=(8, 8))
plt.plot(angle_sweep, un_mean_error[:, 1])
plt.xlabel("Angle of Rotation")
plt.ylabel("Generalization Error (XOR)")
plt.show()
# ### 500 task samples
# +
# number of times to run the experiment, decrease for shorter run times
mc_rep = 100
# samples to use for task1 (xor)
task1_sample = 500
# samples to use for task2 (rxor)
task2_sample = 500
# we will use the same angle_sweep as before
angle_sweep = range(0, 90, 1)
# call our function to run the experiment
un_mean_te, un_mean_error = fn.unaware_bte_v_angle(
angle_sweep, task1_sample, task2_sample, mc_rep
)
# plot angle vs BTE
fn.plot_unaware_bte_v_angle(un_mean_te)
# -
# plot angle vs generalization error
plt.figure(figsize=(8, 8))
plt.plot(angle_sweep, un_mean_error[:, 1])
plt.xlabel("Angle of Rotation")
plt.ylabel("Generalization Error (XOR)")
plt.show()
# ### 1000 task samples
# +
# number of times to run the experiment, decrease for shorter run times
mc_rep = 100
# samples to use for task1 (xor)
task1_sample = 1000
# samples to use for task2 (rxor)
task2_sample = 1000
# we will use the same angle_sweep as before
angle_sweep = range(0, 90, 1)
# call our function to run the experiment
un_mean_te, un_mean_error = fn.unaware_bte_v_angle(
angle_sweep, task1_sample, task2_sample, mc_rep
)
# plot angle vs BTE
fn.plot_unaware_bte_v_angle(un_mean_te)
# -
# plot angle vs generalization error
plt.figure(figsize=(8, 8))
plt.plot(angle_sweep, un_mean_error[:, 1])
plt.xlabel("Angle of Rotation")
plt.ylabel("Generalization Error (XOR)")
plt.show()
| docs/experiments/xor_rxor_with_unaware.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Download** (right-click, save target as ...) this page as a jupyterlab notebook from: [Lab31](http://192.168.3.11/engr-1330-webroot/8-Labs/Lab31/Lab31.ipynb)
#
# ___
# # <font color=darkred>Laboratory 31: "On The Virtue and Value of Classification" or "Who Ordered a Classy Fire?" </font>
# LAST NAME, FIRST NAME
#
# R00000000
#
# ENGR 1330 Laboratory 14 - In-Lab
#  <br>
#
# ## For the last few sessions we have talked about regression ... <br>
#
#  <br>
#
# ### We discussed ...
# - __The theory and implementation of simple linear regression in Python__<br>
# - __OLS and MLE methods for estimation of slope and intercept coefficients__ <br>
# - __Errors (Noise, Variance, Bias) and their impacts on model's performance__ <br>
# - __Confidence and prediction intervals__
# - __And Multiple Linear Regressions__
#
# <br>  <br>
#
# - __What if we want to predict a discrete variable?__
#
# The general idea behind our efforts was to use a set of observed events (samples) to capture the relationship between one or more predictor (AKA input, indipendent) variables and an output (AKA response, dependent) variable. The nature of the dependent variables differentiates *__regression__* and *__classification__* problems.
# <br> 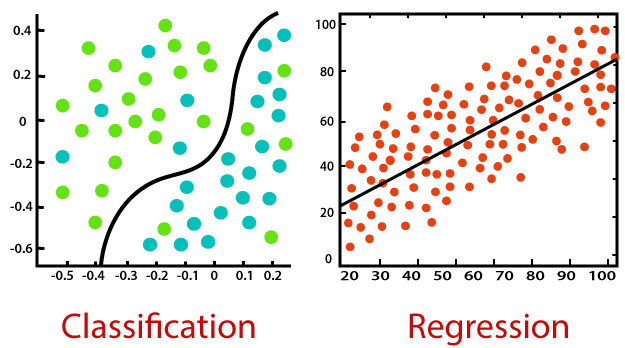 <br>
#
#
# Regression problems have continuous and usually unbounded outputs. An example is when you’re estimating the salary as a function of experience and education level. Or all the examples we have covered so far!
#
# On the other hand, classification problems have discrete and finite outputs called classes or categories. For example, predicting if an employee is going to be promoted or not (true or false) is a classification problem. There are two main types of classification problems:
#
# - Binary or binomial classification:
#
# exactly two classes to choose between (usually 0 and 1, true and false, or positive and negative)
#
# - Multiclass or multinomial classification:
#
# three or more classes of the outputs to choose from
#
#
# - __When Do We Need Classification?__
#
# We can apply classification in many fields of science and technology. For example, text classification algorithms are used to separate legitimate and spam emails, as well as positive and negative comments. Other examples involve medical applications, biological classification, credit scoring, and more.
#
# ## Logistic Regression
#
# - __What is logistic regression?__
# Logistic regression is a fundamental classification technique. It belongs to the group of linear classifiers and is somewhat similar to polynomial and linear regression. Logistic regression is fast and relatively uncomplicated, and it’s convenient for users to interpret the results. Although it’s essentially a method for binary classification, it can also be applied to multiclass problems.
#
# <br>  <br>
#
#
#
# Logistic regression is a statistical method for predicting binary classes. The outcome or target variable is dichotomous in nature. Dichotomous means there are only two possible classes. For example, it can be used for cancer detection problems. It computes the probability of an event occurrence. Logistic regression can be considered a special case of linear regression where the target variable is categorical in nature. It uses a log of odds as the dependent variable. Logistic Regression predicts the probability of occurrence of a binary event utilizing a logit function. HOW?
# Remember the general format of the multiple linear regression model:
# <br> 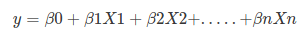 <br>
#
# Where, y is dependent variable and x1, x2 ... and Xn are explanatory variables. This was, as you know by now, a linear function. There is another famous function known as the *__Sigmoid Function__*, also called *__logistic function__*. Here is the equation for the Sigmoid function:
# <br> 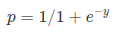 <br>
#
# This image shows the sigmoid function (or S-shaped curve) of some variable 𝑥:
# <br> 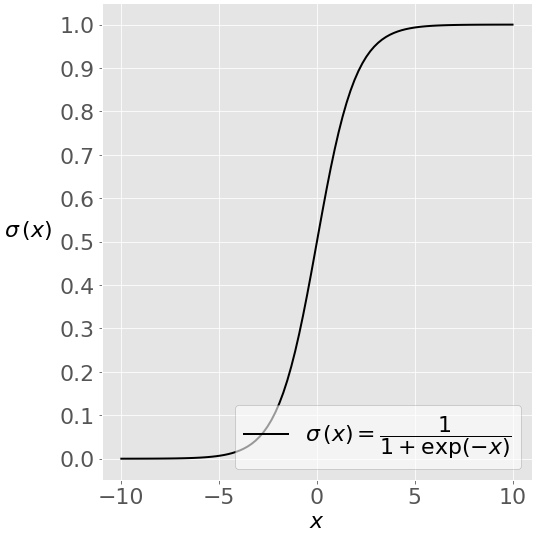 <br>
# As you see, The sigmoid function has values very close to either 0 or 1 across most of its domain. It can take any real-valued number and map it into a value between 0 and 1. If the curve goes to positive infinity, y predicted will become 1, and if the curve goes to negative infinity, y predicted will become 0. This fact makes it suitable for application in classification methods since we are dealing with two discrete classes (labels, categories, ...). If the output of the sigmoid function is more than 0.5, we can classify the outcome as 1 or YES, and if it is less than 0.5, we can classify it as 0 or NO. This cutoff value (threshold) is not always fixed at 0.5. If we apply the Sigmoid function on linear regression:
# <br>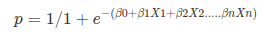 <br>
#
# Notice the difference between linear regression and logistic regression:
# <br>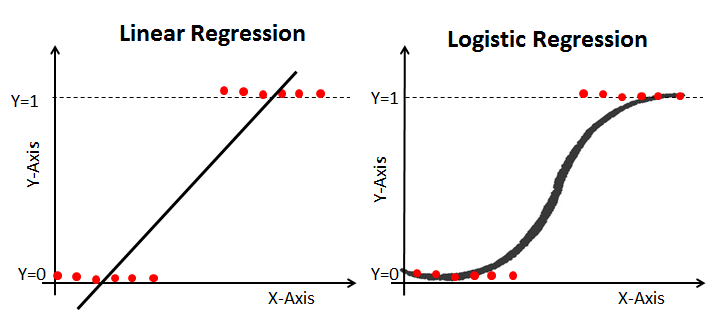 <br>
#
# logistic regression is estimated using Maximum Likelihood Estimation (MLE) approach. Maximizing the likelihood function determines the parameters that are most likely to produce the observed data.
#
# Let's work on an example in Python! <br>
#
#  <br>
# ### Example 1: Diagnosing Diabetes <br>
#
#  <br>
#
#
#
# #### The "diabetes.csv" dataset is originally from the National Institute of Diabetes and Digestive and Kidney Diseases. The objective of the dataset is to diagnostically predict whether or not a patient has diabetes, based on certain diagnostic measurements included in the dataset.
# *Several constraints were placed on the selection of these instances from a larger database. In particular, all patients here are females at least 21 years old of Pima Indian heritage.*
# #### The datasets consists of several medical predictor variables and one target variable, Outcome. Predictor variables includes the number of pregnancies the patient has had, their BMI, insulin level, age, and so on.
#
# |Columns|Info.|
# |---:|---:|
# |Pregnancies |Number of times pregnant|
# |Glucose |Plasma glucose concentration a 2 hours in an oral glucose tolerance test|
# |BloodPressure |Diastolic blood pressure (mm Hg)|
# |SkinThickness |Triceps skin fold thickness (mm)|
# |Insulin |2-Hour serum insulin (mu U/ml)|
# |BMI |Body mass index (weight in kg/(height in m)^2)|
# |Diabetes pedigree |Diabetes pedigree function|
# |Age |Age (years)|
# |Outcome |Class variable (0 or 1) 268 of 768 are 1, the others are 0|
#
#
# #### Let's see if we can build a logistic regression model to accurately predict whether or not the patients in the dataset have diabetes or not?
# *Acknowledgements:
# <NAME>., <NAME>., <NAME>., <NAME>., & <NAME>. (1988). Using the ADAP learning algorithm to forecast the onset of diabetes mellitus. In Proceedings of the Symposium on Computer Applications and Medical Care (pp. 261--265). IEEE Computer Society Press.*
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import sklearn.metrics as metrics
import seaborn as sns
# %matplotlib inline
# Import the dataset:
data = pd.read_csv("diabetes.csv")
data.rename(columns = {'Pregnancies':'pregnant', 'Glucose':'glucose','BloodPressure':'bp','SkinThickness':'skin',
'Insulin ':'Insulin','BMI':'bmi','DiabetesPedigreeFunction':'pedigree','Age':'age',
'Outcome':'label'}, inplace = True)
data.head()
data.describe()
#Check some histograms
sns.distplot(data['pregnant'], kde = True, rug= True, color ='orange')
sns.distplot(data['glucose'], kde = True, rug= True, color ='darkblue')
sns.distplot(data['label'], kde = False, rug= True, color ='purple', bins=2)
sns.jointplot(x ='glucose', y ='label', data = data, kind ='kde')
# #### Selecting Feature: Here, we need to divide the given columns into two types of variables dependent(or target variable) and independent variable(or feature variables or predictors).
#split dataset in features and target variable
feature_cols = ['pregnant', 'glucose', 'bp', 'skin', 'Insulin', 'bmi', 'pedigree', 'age']
X = data[feature_cols] # Features
y = data.label # Target variable
# #### Splitting Data: To understand model performance, dividing the dataset into a training set and a test set is a good strategy. Let's split dataset by using function train_test_split(). You need to pass 3 parameters: features, target, and test_set size. Additionally, you can use random_state to select records randomly. Here, the Dataset is broken into two parts in a ratio of 75:25. It means 75% data will be used for model training and 25% for model testing:
# split X and y into training and testing sets
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.25,random_state=0)
# #### Model Development and Prediction: First, import the Logistic Regression module and create a Logistic Regression classifier object using LogisticRegression() function. Then, fit your model on the train set using fit() and perform prediction on the test set using predict().
# +
# import the class
from sklearn.linear_model import LogisticRegression
# instantiate the model (using the default parameters)
#logreg = LogisticRegression()
logreg = LogisticRegression()
# fit the model with data
logreg.fit(X_train,y_train)
#
y_pred=logreg.predict(X_test)
# -
#  <br>
# - __How to assess the performance of logistic regression?__
#
# Binary classification has four possible types of results:
#
# - True negatives: correctly predicted negatives (zeros)
# - True positives: correctly predicted positives (ones)
# - False negatives: incorrectly predicted negatives (zeros)
# - False positives: incorrectly predicted positives (ones)
#
# We usually evaluate the performance of a classifier by comparing the actual and predicted outputsand counting the correct and incorrect predictions. A confusion matrix is a table that is used to evaluate the performance of a classification model.
#
# <br> 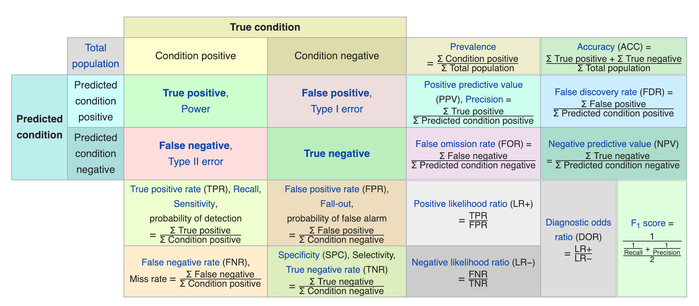 <br>
#
# Some indicators of binary classifiers include the following:
#
# - The most straightforward indicator of classification accuracy is the ratio of the number of correct predictions to the total number of predictions (or observations).
# - The positive predictive value is the ratio of the number of true positives to the sum of the numbers of true and false positives.
# - The negative predictive value is the ratio of the number of true negatives to the sum of the numbers of true and false negatives.
# - The sensitivity (also known as recall or true positive rate) is the ratio of the number of true positives to the number of actual positives.
# - The precision score quantifies the ability of a classifier to not label a negative example as positive. The precision score can be interpreted as the probability that a positive prediction made by the classifier is positive.
# - The specificity (or true negative rate) is the ratio of the number of true negatives to the number of actual negatives.
# <br> 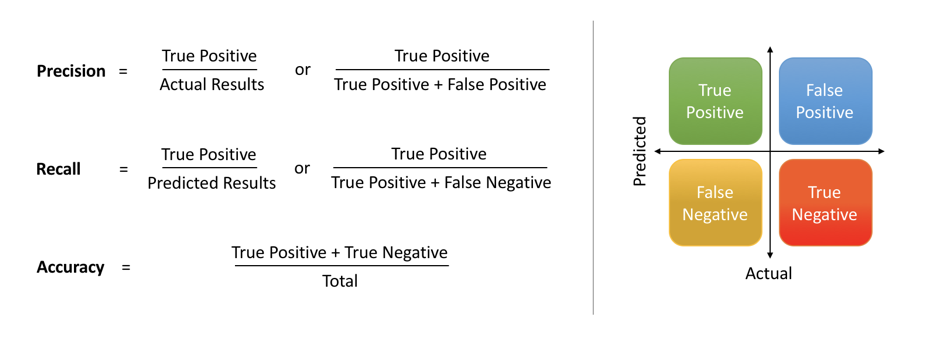 <br>
#
# The extent of importance of recall and precision depends on the problem. Achieving a high recall is more important than getting a high precision in cases like when we would like to detect as many heart patients as possible. For some other models, like classifying whether a bank customer is a loan defaulter or not, it is desirable to have a high precision since the bank wouldn’t want to lose customers who were denied a loan based on the model’s prediction that they would be defaulters.
# There are also a lot of situations where both precision and recall are equally important. Then we would aim for not only a high recall but a high precision as well. In such cases, we use something called F1-score. F1-score is the Harmonic mean of the Precision and Recall:
# <br>  <br>
# This is easier to work with since now, instead of balancing precision and recall, we can just aim for a good F1-score and that would be indicative of a good Precision and a good Recall value as well.
# <br>  <br>
# #### Model Evaluation using Confusion Matrix: A confusion matrix is a table that is used to evaluate the performance of a classification model. You can also visualize the performance of an algorithm. The fundamental of a confusion matrix is the number of correct and incorrect predictions are summed up class-wise.
# import the metrics class
from sklearn import metrics
cnf_matrix = metrics.confusion_matrix(y_pred, y_test)
cnf_matrix
# #### Here, you can see the confusion matrix in the form of the array object. The dimension of this matrix is 2*2 because this model is binary classification. You have two classes 0 and 1. Diagonal values represent accurate predictions, while non-diagonal elements are inaccurate predictions. In the output, 119 and 36 are actual predictions, and 26 and 11 are incorrect predictions.
# #### Visualizing Confusion Matrix using Heatmap: Let's visualize the results of the model in the form of a confusion matrix using matplotlib and seaborn.
class_names=[0,1] # name of classes
fig, ax = plt.subplots()
tick_marks = np.arange(len(class_names))
plt.xticks(tick_marks, class_names)
plt.yticks(tick_marks, class_names)
# create heatmap
sns.heatmap(pd.DataFrame(cnf_matrix), annot=True, cmap="YlGnBu" ,fmt='g')
ax.xaxis.set_label_position("top")
plt.tight_layout()
plt.title('Confusion matrix', y=1.1)
plt.ylabel('Predicted label')
plt.xlabel('Actual label')
# #### Confusion Matrix Evaluation Metrics: Let's evaluate the model using model evaluation metrics such as accuracy, precision, and recall.
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
print("Precision:",metrics.precision_score(y_test, y_pred))
print("Recall:",metrics.recall_score(y_test, y_pred))
print("F1-score:",metrics.f1_score(y_test, y_pred))
# +
from sklearn.metrics import classification_report
print(classification_report(y_test, y_pred))
# -
# 
# ___
# ## Example: Credit Card Fraud Detection <br>
#
#  <br>
#
#
#
# ### For many companies, losses involving transaction fraud amount to more than 10% of their total expenses. The concern with these massive losses leads companies to constantly seek new solutions to prevent, detect and eliminate fraud. Machine Learning is one of the most promising technological weapons to combat financial fraud. The objective of this project is to create a simple Logistic Regression model capable of detecting fraud in credit card operations, thus seeking to minimize the risk and loss of the business.
#
# ### The dataset used contains transactions carried out by European credit card holders that took place over two days in September 2013, and is a shorter version of a dataset that is available on kaggle at https://www.kaggle.com/mlg-ulb/creditcardfraud/version/3.
#
# ### "It contains only numerical input variables which are the result of a PCA transformation. Unfortunately, due to confidentiality issues, we cannot provide the original features and more background information about the data. Features V1, V2, … V28 are the principal components obtained with PCA, the only features which have not been transformed with PCA are 'Time' and 'Amount'. Feature 'Time' contains the seconds elapsed between each transaction and the first transaction in the dataset. The feature 'Amount' is the transaction Amount, this feature can be used for example-dependant cost-senstive learning. Feature 'Class' is the response variable and it takes value 1 in case of fraud and 0 otherwise."
#
#
# |Columns|Info.|
# |---:|---:|
# |Time |Number of seconds elapsed between this transaction and the first transaction in the dataset|
# |V1-V28 |Result of a PCA Dimensionality reduction to protect user identities and sensitive features(v1-v28)|
# |Amount |Transaction amount|
# |Class |1 for fraudulent transactions, 0 otherwise|
#
#
# *NOTE: Principal Component Analysis, or PCA, is a dimensionality-reduction method that is often used to reduce the dimensionality of large data sets, by transforming a large set of variables into a smaller one that still contains most of the information in the large set.*
#
# <hr>
#
# *__Acknowledgements__*
# The dataset has been collected and analysed during a research collaboration of Worldline and the Machine Learning Group (http://mlg.ulb.ac.be) of ULB (Université Libre de Bruxelles) on big data mining and fraud detection.
# More details on current and past projects on related topics are available on https://www.researchgate.net/project/Fraud-detection-5 and the page of the DefeatFraud project
#
# Please cite the following works:
#
# *<NAME>, <NAME>, <NAME> and <NAME>. Calibrating Probability with Undersampling for Unbalanced Classification. In Symposium on Computational Intelligence and Data Mining (CIDM), IEEE, 2015*
#
# *<NAME>, Andrea; <NAME>; <NAME>; <NAME>; <NAME>. Learned lessons in credit card fraud detection from a practitioner perspective, Expert systems with applications,41,10,4915-4928,2014, Pergamon*
#
# *<NAME>, Andrea; <NAME>; <NAME>; <NAME>; <NAME>. Credit card fraud detection: a realistic modeling and a novel learning strategy, IEEE transactions on neural networks and learning systems,29,8,3784-3797,2018,IEEE*
#
# *<NAME>, Andrea Adaptive Machine learning for credit card fraud detection ULB MLG PhD thesis (supervised by G. Bontempi)*
#
# *<NAME>; <NAME>, Andrea; <NAME>, Yann-Aël; <NAME>; <NAME>; <NAME>. Scarff: a scalable framework for streaming credit card fraud detection with Spark, Information fusion,41, 182-194,2018,Elsevier*
#
# *<NAME>; <NAME>, Yann-Aël; <NAME>; <NAME>. Streaming active learning strategies for real-life credit card fraud detection: assessment and visualization, International Journal of Data Science and Analytics, 5,4,285-300,2018,Springer International Publishing*
#
# *<NAME>, <NAME>, <NAME>, <NAME>, <NAME> Deep-Learning Domain Adaptation Techniques for Credit Cards Fraud Detection, INNSBDDL 2019: Recent Advances in Big Data and Deep Learning, pp 78-88, 2019*
#
# *<NAME>, <NAME>, <NAME>, <NAME>, <NAME> Combining Unsupervised and Supervised Learning in Credit Card Fraud Detection Information Sciences, 2019*
# ### As you know by now, the first step is to load some necessary libraries:
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import sklearn.metrics as metrics
import seaborn as sns
# %matplotlib inline
# ### Then, we should read the dataset and explore it using tools such as descriptive statistics:
# Import the dataset:
data = pd.read_csv("creditcard_m.csv")
data.head()
# ### As expected, the dataset has 31 columns and the target variable is located in the last one. Let's check and see whether we have any missing values in the dataset:
data.isnull().sum()
# ### Great! No missing values!
data.describe()
print ('Not Fraud % ',round(data['Class'].value_counts()[0]/len(data)*100,2))
print ()
print (round(data.Amount[data.Class == 0].describe(),2))
print ()
print ()
print ('Fraud % ',round(data['Class'].value_counts()[1]/len(data)*100,2))
print ()
print (round(data.Amount[data.Class == 1].describe(),2))
# ### We have a total of 140000 samples in this dataset. The PCA components (V1-V28) look as if they have similar spreads and rather small mean values in comparison to another predictors such as 'Time'. The majority (75%) of transactions are below 81 euros with some considerably high outliers (the max is 19656.53 euros). Around 0.19% of all the observed transactions were found to be fraudulent which means that we are dealing with an extremely unbalanced dataset. An important characteristic of such problems. Although the share may seem small, each fraud transaction can represent a very significant expense, which together can represent billions of dollars of lost revenue each year.
# ### The next step is to defind our predictors and target:
#split dataset in features and target variable
y = data.Class # Target variable
X = data.loc[:, data.columns != "Class"] # Features
# ### The next step would be to split our dataset and define the training and testing sets. The random seed (np.random.seed) is used to ensure that the same data is used for all runs. Let's do a 70/30 split:
# +
# split X and y into training and testing sets
np.random.seed(123)
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.30,random_state=1)
# -
# ### Now it is time for model development and prediction!
# ### import the Logistic Regression module and create a Logistic Regression classifier object using LogisticRegression() function. Then, fit your model on the train set using fit() and perform prediction on the test set using predict().
# +
# import the class
from sklearn.linear_model import LogisticRegression
# instantiate the model (using the default parameters)
#logreg = LogisticRegression()
logreg = LogisticRegression(solver='lbfgs',max_iter=10000)
# fit the model with data -TRAIN the model
logreg.fit(X_train,y_train)
# -
# TEST the model
y_pred=logreg.predict(X_test)
# ### Once the model and the predictions are ready, we can assess the performance of our classifier. First, we need to get our confusion matrix:
#
# *A confusion matrix is a table that is used to evaluate the performance of a classification model. You can also visualize the performance of an algorithm. The fundamental of a confusion matrix is the number of correct and incorrect predictions are summed up class-wise.*
from sklearn import metrics
cnf_matrix = metrics.confusion_matrix(y_pred, y_test)
print(cnf_matrix)
tpos = cnf_matrix[0][0]
fneg = cnf_matrix[1][1]
fpos = cnf_matrix[0][1]
tneg = cnf_matrix[1][0]
print("True Positive Cases are",tpos) #How many non-fraud cases were identified as non-fraud cases - GOOD
print("True Negative Cases are",tneg) #How many Fraud cases were identified as Fraud cases - GOOD
print("False Positive Cases are",fpos) #How many Fraud cases were identified as non-fraud cases - BAD | (type 1 error)
print("False Negative Cases are",fneg) #How many non-fraud cases were identified as Fraud cases - BAD | (type 2 error)
class_names=[0,1] # name of classes
fig, ax = plt.subplots()
tick_marks = np.arange(len(class_names))
plt.xticks(tick_marks, class_names)
plt.yticks(tick_marks, class_names)
# create heatmap
sns.heatmap(pd.DataFrame(cnf_matrix), annot=True, cmap="YlGnBu" ,fmt='g')
ax.xaxis.set_label_position("top")
plt.tight_layout()
plt.title('Confusion matrix', y=1.1)
plt.ylabel('Predicted label')
plt.xlabel('Actual label')
# ### We should go further and evaluate the model using model evaluation metrics such as accuracy, precision, and recall. These are calculated based on the confustion matrix:
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
# ### That is a fantastic accuracy score, isn't it?
print("Precision:",metrics.precision_score(y_test, y_pred))
print("Recall:",metrics.recall_score(y_test, y_pred))
print("F1-score:",metrics.f1_score(y_test, y_pred))
# +
from sklearn.metrics import classification_report
print(classification_report(y_test, y_pred))
# -
# ### Although the accuracy is excellent, the model struggles with fraud detection and has not captured about 30 out of 71 fraudulent transactions.
# ### Accuracy in a highly unbalanced data set does not represent a correct value for the efficiency of a model. That's where precision, recall and more specifically F1-score as their combinations becomes important:
#
# - *Accuracy is used when the True Positives and True negatives are more important while F1-score is used when the False Negatives and False Positives are crucial*
#
# - *Accuracy can be used when the class distribution is similar while F1-score is a better metric when there are imbalanced classes as in the above case.*
#
# - *In most real-life classification problems, imbalanced class distribution exists and thus F1-score is a better metric to evaluate our model on.*
#  <br>
#
# *This notebook was inspired by several blogposts including:*
#
# - __"Logistic Regression in Python"__ by __<NAME>__ available at* https://realpython.com/logistic-regression-python/ <br>
# - __"Understanding Logistic Regression in Python"__ by __<NAME>__ available at* https://www.datacamp.com/community/tutorials/understanding-logistic-regression-python <br>
# - __"Understanding Logistic Regression with Python: Practical Guide 1"__ by __<NAME>__ available at* https://datascience.foundation/sciencewhitepaper/understanding-logistic-regression-with-python-practical-guide-1 <br>
# - __"Understanding Data Science Classification Metrics in Scikit-Learn in Python"__ by __<NAME>__ available at* https://towardsdatascience.com/understanding-data-science-classification-metrics-in-scikit-learn-in-python-3bc336865019 <br>
#
#
# *Here are some great reads on these topics:*
# - __"Example of Logistic Regression in Python"__ available at* https://datatofish.com/logistic-regression-python/ <br>
# - __"Building A Logistic Regression in Python, Step by Step"__ by __<NAME>__ available at* https://towardsdatascience.com/building-a-logistic-regression-in-python-step-by-step-becd4d56c9c8 <br>
# - __"How To Perform Logistic Regression In Python?"__ by __<NAME>__ available at* https://www.edureka.co/blog/logistic-regression-in-python/ <br>
# - __"Logistic Regression in Python Using Scikit-learn"__ by __<NAME>__ available at* https://heartbeat.fritz.ai/logistic-regression-in-python-using-scikit-learn-d34e882eebb1 <br>
# - __"ML | Logistic Regression using Python"__ available at* https://www.geeksforgeeks.org/ml-logistic-regression-using-python/ <br>
#
# *Here are some great videos on these topics:*
# - __"StatQuest: Logistic Regression"__ by __StatQuest with <NAME>__ available at* https://www.youtube.com/watch?v=yIYKR4sgzI8&list=PLblh5JKOoLUKxzEP5HA2d-Li7IJkHfXSe <br>
# - __"Linear Regression vs Logistic Regression | Data Science Training | Edureka"__ by __edureka!__ available at* https://www.youtube.com/watch?v=OCwZyYH14uw <br>
# - __"Logistic Regression in Python | Logistic Regression Example | Machine Learning Algorithms | Edureka"__ by __edureka!__ available at* https://www.youtube.com/watch?v=VCJdg7YBbAQ <br>
# - __"How to evaluate a classifier in scikit-learn"__ by __Data School__ available at* https://www.youtube.com/watch?v=85dtiMz9tSo <br>
# - __"How to evaluate a classifier in scikit-learn"__ by __Data School__ available at* https://www.youtube.com/watch?v=85dtiMz9tSo <br>
# ___
#  <br>
#
# ## Exercise: Logistic Regression in Engineering <br>
#
# ### Think of a few applications of Logistic Regression in Engineering?
#
# #### _Make sure to cite any resources that you may use._
#  <br>
#
| 8-Labs/Lab31/.ipynb_checkpoints/Lab31-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp gcp_elevation_api
# -
# # Traffic networks with google elevation api
#
# > Integrating traffic networks with google elevation api for an area in Melbourne. Then calculate the shortest path which accounts for grade impedance.
#hide
from nbdev.showdoc import *
import os
import numpy as np
import geopandas as gpd
import osmnx as ox
# Setup google elevation api key
ELEVATION_API = os.environ.get('GCP_ELEVATION_API')
# ### Get a network and add elevation to its nodes
# Check number of nodes and edges of Greater Melbourne
GMB = ox.graph_from_place("Greater Melbourne, Victoria, Australia", network_type='all')
len(GMB.nodes), len(GMB.edges)
# Check number of nodes and edges of City of Monash
GM = ox.graph_from_place("City of Monash, Victoria, Australia", network_type='all')
len(GM.nodes), len(GM.edges)
# ##### Get a network by place
# Get for a small area
place = "Oakleigh"
place_query = "Oakleigh, the city of Monash, Victoria, 3166, Australia"
G = ox.graph_from_place(place_query, network_type='bike')
len(G.nodes), len(G.edges)
fig, ax = ox.plot_graph(G)
# ##### Get elevation data
# Add elevations to nodes, and grades to edges
G = ox.add_node_elevations(G, api_key=ELEVATION_API)
G = ox.add_edge_grades(G)
# ### Calculate several stats
# ##### Average and median grade
edge_grades = [data['grade_abs'] for u, v, k, data in ox.get_undirected(G).edges(keys=True, data=True)]
avg_grade = np.mean(edge_grades)
print('Average street grade in {} is {:.1f}%'.format(place, avg_grade*100))
med_grade = np.median(edge_grades)
print('Median street grade in {} is {:.1f}%'.format(place, med_grade*100))
# ##### Plot nodes by elevation
# get one color for each node by elevation
nc = ox.plot.get_node_colors_by_attr(G, 'elevation', cmap='plasma')
fig, ax = ox.plot_graph(G, node_color=nc, node_size=5, edge_color='#333333', bgcolor='k')
# ##### Plot the edges by grade
# get a color for each edge, by grade, then plot the network
ec = ox.plot.get_edge_colors_by_attr(G, 'grade_abs', cmap='plasma', num_bins=5, equal_size=True)
fig, ax = ox.plot_graph(G, edge_color=ec, edge_linewidth=0.5, node_size=0, bgcolor='k')
# ### Shortest paths account for grade impedance
from shapely.geometry import Polygon, Point
# Select an origin and destination node
orig = (-37.8943, 145.0900)
dest = (-37.9059, 145.1030)
# Check the distance
Point(orig).distance(Point(dest))
# Get nearest nodes and a bounding box
orig = ox.get_nearest_node(G, orig)
dest = ox.get_nearest_node(G, dest)
bbox = ox.utils_geo.bbox_from_point((-37.9001, 145.0965), dist=1500)
# +
# An edge impedance function
def impedance(length, grade):
penalty = grade ** 2
return length * penalty
def impedance_2(length, grade):
penalty = length * (np.abs(grade) ** 3)
return penalty
# -
for u, v, k, data in G.edges(keys=True, data=True):
# data['impedance'] = impedance(data['length'], data['grade_abs'])
data['impedance'] = impedance_2(data['length'], data['grade'])
data['rise'] = data['length'] * data['grade']
# #### First find the shortest path by minimising distance
route_by_length = ox.shortest_path(G, orig, dest, weight='length')
fig, ax = ox.plot_graph_route(G, route_by_length, node_size=0)
# #### Find the shortest path by minimising impedance
route_by_impedance = ox.shortest_path(G, orig, dest, weight='impedance')
fig, ax = ox.plot_graph_route(G, route_by_impedance, node_size=0)
# #### Stats about these two routes
def print_route_stats(route):
route_grades = ox.utils_graph.get_route_edge_attributes(G, route, 'grade_abs')
msg = 'The average grade is {:.1f}% and the max is {:.1f}%'
print(msg.format(np.mean(route_grades)*100, np.max(route_grades)*100))
route_rises = ox.utils_graph.get_route_edge_attributes(G, route, 'rise')
ascent = np.sum([rise for rise in route_rises if rise >= 0])
descent = np.sum([rise for rise in route_rises if rise < 0])
msg = 'Total elevation change is {:.1f} meters: a {:.0f} meter ascent and a {:.0f} meter descent'
print(msg.format(np.sum(route_rises), ascent, abs(descent)))
route_lengths = ox.utils_graph.get_route_edge_attributes(G, route, 'length')
print('Total trip distance: {:,.0f} meters'.format(np.sum(route_lengths)))
# stats of route minimizing length
print_route_stats(route_by_length)
# stats of route minimizing impedance (function of length and grade)
print_route_stats(route_by_impedance)
| 12_gcp_elevation_api.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pre-processing notebook
# In this notebook, we will pre-process the frames. For better visualisation, we will just capture 2 frames and visualise all the steps. The steps are:
# 1. Capture 2 consecutive frames.
# 2. Find difference between the frames to capture the motion.
# 3. Use GaussianBlur, thresholding, dilation and erosion to pre-process the frames.
# 4. Image segmentation using contours. Extract the vehicles during this method.
# 5. Convert contours to hulls.
# +
# Run these if OpenCV doesn't load
import sys
# sys.path.append('/Library/Frameworks/Python.framework/Versions/3.6/lib/python3.6/site-packages/cv2/')
# -
# First, we import the necessary libraries
import cv2
import numpy as np
import math
import matplotlib.pyplot as plt
# %matplotlib inline
# Next, we define variables that will be used through the duration of the code
# Here, we define some colours
SCALAR_BLACK = (0.0,0.0,0.0)
SCALAR_WHITE = (255.0,255.0,255.0)
SCALAR_YELLOW = (0.0,255.0,255.0)
SCALAR_GREEN = (0.0,255.0,0.0)
SCALAR_RED = (0.0,0.0,255.0)
SCALAR_CYAN = (255.0,255.0,0.0)
# Function to draw the image
# function to plot n images using subplots
def plot_image(images, captions=None, cmap=None ):
f, axes = plt.subplots(1, len(images), sharey=True)
f.set_figwidth(15)
for ax,image,caption in zip(axes, images, captions):
ax.imshow(image, cmap)
ax.set_title(caption)
# ### Capturing movement in video
# **Two consecutive frames are required to capture the movement**. If there is movement in vehicle, there will be small change in pixel value in the current frame compared to the previous frame. The change implies movement. Let's capture the first 2 frames now.
# +
SHOW_DEBUG_STEPS = True
# Reading video
cap = cv2.VideoCapture('./AundhBridge.mp4')
# if video is not present, show error
if not(cap.isOpened()):
print("Error reading file")
# Check if you are able to capture the video
ret, fFrame = cap.read()
# Capturing 2 consecutive frames and making a copy of those frame. Perform all operations on the copy frame.
ret, fFrame1 = cap.read()
ret, fFrame2 = cap.read()
img1 = fFrame1.copy()
img2 = fFrame2.copy()
if(SHOW_DEBUG_STEPS):
print ('img1 height' + str(img1.shape[0]))
print ('img1 width' + str(img1.shape[1]))
print ('img2 height' + str(img2.shape[0]))
print ('img2 width' + str(img2.shape[1]))
# Convert the colour images to greyscale in order to enable fast processing
img1 = cv2.cvtColor(img1, cv2.COLOR_BGR2GRAY)
img2 = cv2.cvtColor(img2, cv2.COLOR_BGR2GRAY)
#plotting
plot_image([img1, img2], cmap='gray', captions=["First frame", "Second frame"])
# -
# ### Adding gaussion blur for smoothening
# +
# Add some Gaussian Blur
img1 = cv2.GaussianBlur(img1,(5,5),0)
img2 = cv2.GaussianBlur(img2,(5,5),0)
#plotting
plot_image([img1, img2], cmap='gray', captions=["GaussianBlur first frame", "GaussianBlur second frame"])
# -
# ### Find the movement in video
# If vehicle is moving, there will be **slight change** in pixel value in the next frame compared to previous frame. We then threshold the image. This will be useful further for preprocessing. Pixel value below 30 will be set as 0(black) and above as 255(white)
#
# Thresholding: https://docs.opencv.org/3.0-beta/doc/py_tutorials/py_imgproc/py_thresholding/py_thresholding.html
# +
# This imgDiff variable is the difference between consecutive frames, which is equivalent to detecting movement
imgDiff = cv2.absdiff(img1, img2)
# Thresholding the image that is obtained after taking difference. Pixel value below 30 will be set as 0(black) and above as 255(white)
ret,imgThresh = cv2.threshold(imgDiff,30.0,255.0,cv2.THRESH_BINARY)
ht = np.size(imgThresh,0)
wd = np.size(imgThresh,1)
plot_image([imgDiff, imgThresh], cmap='gray', captions = ["Difference between 2 frames", "Difference between 2 frames after threshold"])
# -
# ### Dilation and erosion in image
# Dilation and erosion in the image with the filter size of structuring elements.
# +
# Now, we define structuring elements
strucEle3x3 = cv2.getStructuringElement(cv2.MORPH_RECT,(3,3))
strucEle5x5 = cv2.getStructuringElement(cv2.MORPH_RECT,(5,5))
strucEle7x7 = cv2.getStructuringElement(cv2.MORPH_RECT,(7,7))
strucEle15x15 = cv2.getStructuringElement(cv2.MORPH_RECT,(15,15))
plot_image([strucEle3x3, strucEle5x5, strucEle7x7, strucEle15x15], cmap='gray', captions = ["strucEle3x3", "strucEle5x5", "strucEle7x7", "strucEle15x15"])
# +
for i in range(2):
imgThresh = cv2.dilate(imgThresh,strucEle5x5,iterations = 2)
imgThresh = cv2.erode(imgThresh,strucEle5x5,iterations = 1)
imgThreshCopy = imgThresh.copy()
if(SHOW_DEBUG_STEPS):
print ('imgThreshCopy height' + str(imgThreshCopy.shape[0]))
print ('imgThreshCopy width' + str(imgThreshCopy.shape[1]))
plt.imshow(imgThresh, cmap = 'gray')
# -
# ## Extracting contours
# Till now, you have a binary image. Next, we will segment the image and find all possible contours(possible vehicles). The shape of the contours will tell us the number of contours that has been identified. Define *drawAndShowContours()* function to plot the contours. You will see that the threshold image above and the countour image will look alike. So, additionally, we also plot a particular '9th' countour for further clarity.
def drawAndShowContours(wd,ht,contours,strImgName):
global SCALAR_WHITE
global SHOW_DEBUG_STEPS
# Defining a blank frame. Since it is initialised with zeros, it will be black. Will add all the coutours in this image.
blank_image = np.zeros((ht,wd,3), np.uint8)
#cv2.drawContours(blank_image,contours,10,SCALAR_WHITE,-1)
# Adding all possible contour to the blank frame. Contour is white
cv2.drawContours(blank_image,contours,-1,SCALAR_WHITE,-1)
# For better clarity, lets just view countour 9
blank_image_contour_9 = np.zeros((ht,wd,3), np.uint8)
# Let's just add contour 9 to the blank image and view it
cv2.drawContours(blank_image_contour_9,contours,8,SCALAR_WHITE,-1)
# Plotting
plot_image([blank_image, blank_image_contour_9], cmap='gray', captions = ["All possible contours", "Only the 9th contour"])
return blank_image
# +
# Now, we move on to the contour mapping portion
im, contours, hierarchy = cv2.findContours(imgThreshCopy,cv2.RETR_EXTERNAL,cv2.CHAIN_APPROX_SIMPLE)
im2 = drawAndShowContours(wd,ht,contours,'imgContours')
# Printing all the coutours in the image.
if(SHOW_DEBUG_STEPS):
print ('contours.shape: ' + str(len(contours)))
# -
# ## Hulls
# Hulls are contours with the "convexHull".
# +
# Next, we define hulls.
# Hulls are contours with the "convexHull" function from cv2
hulls = contours # does it work?
for i in range(len(contours)):
hulls[i] = cv2.convexHull(contours[i])
# Then we draw the contours
im3 = drawAndShowContours(wd,ht,hulls,'imgConvexHulls')
| 06Deep Learning/04Convolutional Neural Networks - Industry Applications/03Industry Demo_ Detecting Vehicles in Videos/07Detecting Object in Video/Pre-processing+notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 3.2
# Import the pySpark libraries
import sys
# !conda install --yes --prefix {sys.prefix} -c conda-forge pyspark
from pyspark.sql import SparkSession
from pyspark.sql.functions import col, split, size
# Connect to a local Spark session
spark = SparkSession.builder.appName("Packt").getOrCreate()
# Read the raw data from a CSV file
data = spark.read.csv('../../Datasets/netflix_titles_nov_2019.csv', header='true')
# Take only the movies
movies = data.filter((col('type') == 'Movie') & (col('release_year') == 2019))
# Add a column with the number of actors
transformed = movies.withColumn('count_cast', size(split(movies['cast'], ',')))
# Select a subset of columns to store
selected = transformed.select('title', 'director', 'count_cast', 'cast', 'rating', 'release_year', 'type')
# Write the contents of the DataFrame to disk
selected.write.csv('transformed.csv', header='true')
| Chapter03/Exercise03.02/spark_etl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import matplotlib .pyplot as plt
# %matplotlib inline
import numpy as np
x = np.linspace(0,20,1000)
y2 = [np.exp(-6*i)*(0.0037*np.cos(8*i)+0.09*np.sin(8*i))+0.017*np.sin(10*i-np.pi*0.5) for i in x]
plt.grid()
plt.plot(x, y2,c = 'r')
| Untitled35.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Hi and welcome to your first code along lesson!
#
# #### Reading, writing and displaying images with OpenCV
# Let's start by importing the OpenCV libary
# +
# Press CTRL + ENTER to run this line
# You should see an * between the [ ] on the left
# OpenCV takes a couple seconds to import the first time
import cv2
# -
# +
# Now let's import numpy
# We use as np, so that everything we call on numpy, we can type np instead
# It's short and looks neater
import numpy as np
# -
# Let's now load our first image
# +
# We don't need to do this again, but it's a good habit
import cv2
# Load an image using 'imread' specifying the path to image
input = cv2.imread('./images/input.jpg')
# Our file 'input.jpg' is now loaded and stored in python
# as a varaible we named 'image'
# To display our image variable, we use 'imshow'
# The first parameter will be title shown on image window
# The second parameter is the image varialbe
cv2.imshow('Hello World', input)
# 'waitKey' allows us to input information when a image window is open
# By leaving it blank it just waits for anykey to be pressed before
# continuing. By placing numbers (except 0), we can specify a delay for
# how long you keep the window open (time is in milliseconds here)
cv2.waitKey()
# This closes all open windows
# Failure to place this will cause your program to hang
cv2.destroyAllWindows()
# +
# Same as above without the extraneous comments
import cv2
input = cv2.imread('./images/input.jpg')
cv2.imshow('<NAME>', input)
cv2.waitKey(0)
cv2.destroyAllWindows()
# -
# ### Let's take a closer look at how images are stored
# Import numpy
import numpy as np
print input.shape
# #### Shape gives the dimensions of the image array
#
# The 2D dimensions are 830 pixels in high bv 1245 pixels wide.
# The '3L' means that there are 3 other components (RGB) that make up this image.
# +
# Let's print each dimension of the image
print 'Height of Image:', int(input.shape[0]), 'pixels'
print 'Width of Image: ', int(input.shape[1]), 'pixels'
# -
# ### How do we save images we edit in OpenCV?
# Simply use 'imwrite' specificing the file name and the image to be saved
cv2.imwrite('output.jpg', input)
cv2.imwrite('output.png', input)
| LECTURES/Lecture 2.4 - Reading, writing and displaying images.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
root_path = os.path.abspath("../../../")
if root_path not in sys.path:
sys.path.append(root_path)
def draw_losses(nn):
el, il = nn.log["epoch_loss"], nn.log["iter_loss"]
ee_base = np.arange(len(el))
ie_base = np.linspace(0, len(el) - 1, len(il))
plt.plot(ie_base, il, label="Iter loss")
plt.plot(ee_base, el, linewidth=3, label="Epoch loss")
plt.legend()
plt.show()
from NN import Basic
from Util.Util import DataUtil
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (18, 8)
# -
x_train, y_train = DataUtil.gen_xor(size=1000, one_hot=False)
x_test, y_test = DataUtil.gen_xor(size=100, one_hot=False)
nn = Basic(model_param_settings={"n_epoch": 200}).fit(x_train, y_train, x_test, y_test, snapshot_ratio=0)
draw_losses(nn)
x_train, y_train = DataUtil.gen_spiral(size=1000, one_hot=False)
x_test, y_test = DataUtil.gen_spiral(size=100, one_hot=False)
nn = Basic(model_param_settings={"n_epoch": 200}).fit(x_train, y_train, x_test, y_test, snapshot_ratio=0)
draw_losses(nn)
x_train, y_train = DataUtil.gen_nine_grid(size=1000, one_hot=False)
x_test, y_test = DataUtil.gen_nine_grid(size=100, one_hot=False)
nn = Basic(model_param_settings={"n_epoch": 200}).fit(x_train, y_train, x_test, y_test, snapshot_ratio=0)
draw_losses(nn)
| _Dist/NeuralNetworks/c_BasicNN/BasicNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Cascade decomposition
#
# This example script shows how to compute and plot the cascade decompositon of
# a single radar precipitation field in pysteps.
#
from matplotlib import cm, pyplot as plt
import numpy as np
import os
from pprint import pprint
from pysteps.cascade.bandpass_filters import filter_gaussian
from pysteps import io, rcparams
from pysteps.cascade.decomposition import decomposition_fft
from pysteps.utils import conversion, transformation
from pysteps.visualization import plot_precip_field
# ## Read precipitation field
#
# First thing, the radar composite is imported and transformed in units
# of dB.
#
#
# +
# Import the example radar composite
root_path = rcparams.data_sources["fmi"]["root_path"]
filename = os.path.join(
root_path, "20160928", "201609281600_fmi.radar.composite.lowest_FIN_SUOMI1.pgm.gz"
)
R, _, metadata = io.import_fmi_pgm(filename, gzipped=True)
# Convert to rain rate
R, metadata = conversion.to_rainrate(R, metadata)
# Nicely print the metadata
pprint(metadata)
# Plot the rainfall field
plot_precip_field(R, geodata=metadata)
plt.show()
# Log-transform the data
R, metadata = transformation.dB_transform(R, metadata, threshold=0.1, zerovalue=-15.0)
# -
# ## 2D Fourier spectrum
#
# Compute and plot the 2D Fourier power spectrum of the precipitaton field.
#
#
# +
# Set Nans as the fill value
R[~np.isfinite(R)] = metadata["zerovalue"]
# Compute the Fourier transform of the input field
F = abs(np.fft.fftshift(np.fft.fft2(R)))
# Plot the power spectrum
M, N = F.shape
fig, ax = plt.subplots()
im = ax.imshow(
np.log(F ** 2), vmin=4, vmax=24, cmap=cm.jet, extent=(-N / 2, N / 2, -M / 2, M / 2)
)
cb = fig.colorbar(im)
ax.set_xlabel("Wavenumber $k_x$")
ax.set_ylabel("Wavenumber $k_y$")
ax.set_title("Log-power spectrum of R")
plt.show()
# -
# ## Cascade decomposition
#
# First, construct a set of Gaussian bandpass filters and plot the corresponding
# 1D filters.
#
#
# +
num_cascade_levels = 7
# Construct the Gaussian bandpass filters
filter = filter_gaussian(R.shape, num_cascade_levels)
# Plot the bandpass filter weights
L = max(N, M)
fig, ax = plt.subplots()
for k in range(num_cascade_levels):
ax.semilogx(
np.linspace(0, L / 2, len(filter["weights_1d"][k, :])),
filter["weights_1d"][k, :],
"k-",
base=pow(0.5 * L / 3, 1.0 / (num_cascade_levels - 2)),
)
ax.set_xlim(1, L / 2)
ax.set_ylim(0, 1)
xt = np.hstack([[1.0], filter["central_wavenumbers"][1:]])
ax.set_xticks(xt)
ax.set_xticklabels(["%.2f" % cf for cf in filter["central_wavenumbers"]])
ax.set_xlabel("Radial wavenumber $|\mathbf{k}|$")
ax.set_ylabel("Normalized weight")
ax.set_title("Bandpass filter weights")
plt.show()
# -
# Finally, apply the 2D Gaussian filters to decompose the radar rainfall field
# into a set of cascade levels of decreasing spatial scale and plot them.
#
#
# +
decomp = decomposition_fft(R, filter, compute_stats=True)
# Plot the normalized cascade levels
for i in range(num_cascade_levels):
mu = decomp["means"][i]
sigma = decomp["stds"][i]
decomp["cascade_levels"][i] = (decomp["cascade_levels"][i] - mu) / sigma
fig, ax = plt.subplots(nrows=2, ncols=4)
ax[0, 0].imshow(R, cmap=cm.RdBu_r, vmin=-5, vmax=5)
ax[0, 1].imshow(decomp["cascade_levels"][0], cmap=cm.RdBu_r, vmin=-3, vmax=3)
ax[0, 2].imshow(decomp["cascade_levels"][1], cmap=cm.RdBu_r, vmin=-3, vmax=3)
ax[0, 3].imshow(decomp["cascade_levels"][2], cmap=cm.RdBu_r, vmin=-3, vmax=3)
ax[1, 0].imshow(decomp["cascade_levels"][3], cmap=cm.RdBu_r, vmin=-3, vmax=3)
ax[1, 1].imshow(decomp["cascade_levels"][4], cmap=cm.RdBu_r, vmin=-3, vmax=3)
ax[1, 2].imshow(decomp["cascade_levels"][5], cmap=cm.RdBu_r, vmin=-3, vmax=3)
ax[1, 3].imshow(decomp["cascade_levels"][6], cmap=cm.RdBu_r, vmin=-3, vmax=3)
ax[0, 0].set_title("Observed")
ax[0, 1].set_title("Level 1")
ax[0, 2].set_title("Level 2")
ax[0, 3].set_title("Level 3")
ax[1, 0].set_title("Level 4")
ax[1, 1].set_title("Level 5")
ax[1, 2].set_title("Level 6")
ax[1, 3].set_title("Level 7")
for i in range(2):
for j in range(4):
ax[i, j].set_xticks([])
ax[i, j].set_yticks([])
plt.tight_layout()
plt.show()
# sphinx_gallery_thumbnail_number = 4
| notebooks/plot_cascade_decomposition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: geopy2019
# language: python
# name: geopy2019
# ---
# # Test that the Python environment is properly installed
#
# - check the "kernel" under "Kernel" -> "change kernel" -> "geopy2019"
# - always close a notebook safely via "File" -> "Save and Checkpoint" and then "Close and Halt"
# - close the Jupyter notebook server via the "Quit" button on the main page
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn
import scipy
import shapely
import gdal
import fiona
import shapely
import geopandas as gpd
import pysal
import bokeh
import cartopy
import mapclassify
import geoviews
import rasterstats
import rasterio
import geoplot
import folium
print("The Pysal package can use additional packages, but we don't need that functionality. The warning is ok.")
| test_install.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.021005, "end_time": "2021-04-23T11:48:51.989118", "exception": false, "start_time": "2021-04-23T11:48:51.968113", "status": "completed"} tags=[]
# # Parcels Experiment:<br><br>Expanding the polyline code to release particles at density based on local velocity normal to section.
#
# _(Based on an experiment originally designed by <NAME>.)_
#
# _(Runs on GEOMAR Jupyter Server at https://schulung3.geomar.de/user/workshop007/lab)_
# + [markdown] papermill={"duration": 0.018031, "end_time": "2021-04-23T11:48:52.025362", "exception": false, "start_time": "2021-04-23T11:48:52.007331", "status": "completed"} tags=[]
# ## To do
#
# - Check/ask how OceanParcels deals with partial cells, if it does.
# - It doesn't. Does it matter?
# + [markdown] papermill={"duration": 0.018189, "end_time": "2021-04-23T11:48:52.063205", "exception": false, "start_time": "2021-04-23T11:48:52.045016", "status": "completed"} tags=[]
# ## Technical preamble
# + papermill={"duration": 3.156467, "end_time": "2021-04-23T11:48:55.237706", "exception": false, "start_time": "2021-04-23T11:48:52.081239", "status": "completed"} tags=[]
# %matplotlib inline
from parcels import (
AdvectionRK4_3D,
ErrorCode,
FieldSet,
JITParticle,
ParticleSet,
Variable
)
# from operator import attrgetter
from datetime import datetime, timedelta
import numpy as np
from pathlib import Path
import matplotlib.pyplot as plt
import cmocean as co
import pandas as pd
import xarray as xr
# import dask as dask
# + [markdown] papermill={"duration": 0.018082, "end_time": "2021-04-23T11:48:55.275850", "exception": false, "start_time": "2021-04-23T11:48:55.257768", "status": "completed"} tags=[]
# ## Experiment settings (user input)
# + [markdown] papermill={"duration": 0.020178, "end_time": "2021-04-23T11:48:55.314305", "exception": false, "start_time": "2021-04-23T11:48:55.294127", "status": "completed"} tags=[]
# ### Parameters
# These can be set in papermill
# + papermill={"duration": 0.02382, "end_time": "2021-04-23T11:48:55.356204", "exception": false, "start_time": "2021-04-23T11:48:55.332384", "status": "completed"} tags=["parameters"]
# OSNAP multiline details
sectionPathname = '../data/external/'
sectionFilename = 'osnap_pos_wp.txt'
sectionname = 'osnap'
# location of input data
path_name = '/data/iAtlantic/ocean-only/VIKING20X.L46-KKG36107B/nemo/output/'
experiment_name = 'VIKING20X.L46-KKG36107B'
data_resolution = '1m'
w_name_extension = '_repaire_depthw_time'
# location of mask data
mask_path_name = '/data/iAtlantic/ocean-only/VIKING20X.L46-KKG36107B/nemo/suppl/'
mesh_mask_filename = '1_mesh_mask.nc_notime_depthw'
# location of output data
outpath_name = '../data/raw/'
year_prefix = 201 # this does from 2000 onwards
# set line segment to use
start_vertex = 4
end_vertex = 12
# experiment duration etc
runtime_in_days = 10
dt_in_minutes = -10
# repeatdt = timedelta(days=3)
# number of particles to track
create_number_particles = 200000 # many will not be ocean points
use_number_particles = 200000
min_release_depth = 0
max_release_depth = 1_000
# max current speed for particle selection
max_current = 1.0
# set base release date and time
t_0_str = '2010-01-16T12:00:00'
t_start_str = '2016-01-16T12:00:00'
# particle positions are stored every x hours
outputdt_in_hours = 120
# select subdomain (to decrease needed resources) comment out to use whole domain
# sd_i1, sd_i2 = 0, 2404 # western/eastern limit (indices not coordinates)
# sd_j1, sd_j2 = 1200, 2499 # southern/northern limit (indices not coordinates)
# sd_z1, sd_z2 = 0, 46
# how to initialize the random number generator
# --> is set in next cell
# RNG_seed = 123
use_dask_chunks = True
# + papermill={"duration": 0.022373, "end_time": "2021-04-23T11:48:55.396851", "exception": false, "start_time": "2021-04-23T11:48:55.374478", "status": "completed"} tags=["injected-parameters"]
# Parameters
path_name = "/gxfs_work1/geomar/smomw355/model_data/ocean-only/VIKING20X.L46-KKG36107B/nemo/output/"
data_resolution = "5d"
w_name_extension = ""
mask_path_name = "/gxfs_work1/geomar/smomw355/model_data/ocean-only/VIKING20X.L46-KKG36107B/nemo/suppl/"
mesh_mask_filename = "1_mesh_mask.nc"
year_prefix = ""
runtime_in_days = 3650
create_number_particles = 4000000
use_number_particles = 4000000
max_release_depth = 1000
max_current = 2.0
t_0_str = "1980-01-03T12:00:00"
t_start_str = "2019-02-22T12:00:00"
use_dask_chunks = False
# + [markdown] papermill={"duration": 0.017956, "end_time": "2021-04-23T11:48:55.432903", "exception": false, "start_time": "2021-04-23T11:48:55.414947", "status": "completed"} tags=[]
# ### Derived variables
# + papermill={"duration": 0.026652, "end_time": "2021-04-23T11:48:55.477730", "exception": false, "start_time": "2021-04-23T11:48:55.451078", "status": "completed"} tags=[]
# times
t_0 = datetime.fromisoformat(t_0_str) # using monthly mean fields. Check dates.
t_start = datetime.fromisoformat(t_start_str)
# RNG seed based on release day (days since 1980-01-03)
RNG_seed = int((t_start - t_0).total_seconds() / (60*60*24))
# names of files to load
fname_U = f'1_{experiment_name}_{data_resolution}_{year_prefix}*_grid_U.nc'
fname_V = f'1_{experiment_name}_{data_resolution}_{year_prefix}*_grid_V.nc'
fname_T = f'1_{experiment_name}_{data_resolution}_{year_prefix}*_grid_T.nc'
fname_W = f'1_{experiment_name}_{data_resolution}_{year_prefix}*_grid_W.nc{w_name_extension}'
sectionPath = Path(sectionPathname)
data_path = Path(path_name)
mask_path = Path(mask_path_name)
outpath = Path(outpath_name)
display(t_0)
display(t_start)
# + papermill={"duration": 0.022805, "end_time": "2021-04-23T11:48:55.519288", "exception": false, "start_time": "2021-04-23T11:48:55.496483", "status": "completed"} tags=[]
if dt_in_minutes > 0:
direction = '_forwards_'
else:
direction = '_backward_'
year_str = str(t_start.year)
month_str = str(t_start.month).zfill(2)
day_str = str(t_start.day).zfill(2)
days = str(runtime_in_days)
seed = str(RNG_seed)
npart= str(use_number_particles)
# + papermill={"duration": 0.022343, "end_time": "2021-04-23T11:48:55.560561", "exception": false, "start_time": "2021-04-23T11:48:55.538218", "status": "completed"} tags=[]
degree2km = 1.852*60.0
# + [markdown] papermill={"duration": 0.020862, "end_time": "2021-04-23T11:48:55.600297", "exception": false, "start_time": "2021-04-23T11:48:55.579435", "status": "completed"} tags=[]
# ## Construct input / output paths etc.
# + papermill={"duration": 0.022398, "end_time": "2021-04-23T11:48:55.641720", "exception": false, "start_time": "2021-04-23T11:48:55.619322", "status": "completed"} tags=[]
mesh_mask = mask_path / mesh_mask_filename
# + [markdown] papermill={"duration": 0.018687, "end_time": "2021-04-23T11:48:55.679163", "exception": false, "start_time": "2021-04-23T11:48:55.660476", "status": "completed"} tags=[]
# ## Load input datasets
# + papermill={"duration": 0.028614, "end_time": "2021-04-23T11:48:55.726606", "exception": false, "start_time": "2021-04-23T11:48:55.697992", "status": "completed"} tags=[]
def fieldset_defintions(
list_of_filenames_U, list_of_filenames_V,
list_of_filenames_W, list_of_filenames_T,
mesh_mask
):
ds_mask = xr.open_dataset(mesh_mask)
filenames = {'U': {'lon': (mesh_mask),
'lat': (mesh_mask),
'depth': list_of_filenames_W[0],
'data': list_of_filenames_U},
'V': {'lon': (mesh_mask),
'lat': (mesh_mask),
'depth': list_of_filenames_W[0],
'data': list_of_filenames_V},
'W': {'lon': (mesh_mask),
'lat': (mesh_mask),
'depth': list_of_filenames_W[0],
'data': list_of_filenames_W},
'T': {'lon': (mesh_mask),
'lat': (mesh_mask),
'depth': list_of_filenames_W[0],
'data': list_of_filenames_T},
'S': {'lon': (mesh_mask),
'lat': (mesh_mask),
'depth': list_of_filenames_W[0],
'data': list_of_filenames_T},
'MXL': {'lon': (mesh_mask),
'lat': (mesh_mask),
'data': list_of_filenames_T}
}
variables = {'U': 'vozocrtx',
'V': 'vomecrty',
'W': 'vovecrtz',
'T': 'votemper',
'S': 'vosaline',
'MXL':'somxl010'
}
dimensions = {'U': {'lon': 'glamf', 'lat': 'gphif', 'depth': 'depthw',
'time': 'time_counter'}, # needs to be on f-nodes
'V': {'lon': 'glamf', 'lat': 'gphif', 'depth': 'depthw',
'time': 'time_counter'}, # needs to be on f-nodes
'W': {'lon': 'glamf', 'lat': 'gphif', 'depth': 'depthw',
'time': 'time_counter'}, # needs to be on f-nodes
'T': {'lon': 'glamf', 'lat': 'gphif', 'depth': 'depthw',
'time': 'time_counter'}, # needs to be on t-nodes
'S': {'lon': 'glamf', 'lat': 'gphif', 'depth': 'depthw',
'time': 'time_counter'}, # needs to be on t-nodes
'MXL': {'lon': 'glamf', 'lat': 'gphif',
'time': 'time_counter'}, # needs to be on t-nodes
}
# exclude the two grid cells at the edges of the nest as they contain 0
# and everything south of 20N
indices = {'lon': range(2, ds_mask.x.size-2), 'lat': range(1132, ds_mask.y.size-2)}
# indices = {
# 'U': {'depth': range(sd_z1, sd_z2), 'lon': range(sd_i1, sd_i2), 'lat': range(sd_j1, sd_j2)},
# 'V': {'depth': range(sd_z1, sd_z2), 'lon': range(sd_i1, sd_i2), 'lat': range(sd_j1, sd_j2)},
# 'W': {'depth': range(sd_z1, sd_z2), 'lon': range(sd_i1, sd_i2), 'lat':range(sd_j1, sd_j2)},
# 'T': {'depth': range(sd_z1, sd_z2), 'lon': range(sd_i1, sd_i2), 'lat':range(sd_j1, sd_j2)},
# 'S': {'depth': range(sd_z1, sd_z2), 'lon': range(sd_i1, sd_i2), 'lat':range(sd_j1, sd_j2)}
# }
if use_dask_chunks:
field_chunksizes = {'U': {'lon':('x', 1024), 'lat':('y',128), 'depth': ('depthw', 64),
'time': ('time_counter',3)}, # needs to be on f-nodes
'V': {'lon':('x', 1024), 'lat':('y',128), 'depth': ('depthw', 64),
'time': ('time_counter',3)}, # needs to be on f-nodes
'W': {'lon':('x', 1024), 'lat':('y',128), 'depth': ('depthw', 64),
'time': ('time_counter',3)}, # needs to be on f-nodes
'T': {'lon':('x', 1024), 'lat':('y',128), 'depth': ('depthw', 64),
'time': ('time_counter',3)}, # needs to be on t-nodes
'S': {'lon':('x', 1024), 'lat':('y',128), 'depth': ('depthw', 64),
'time': ('time_counter',3)}, # needs to be on t-nodes
'MXL': {'lon':('x', 1024), 'lat':('y',128),
'time': ('time_counter',3)}, # needs to be on t-nodes
}
else:
field_chunksizes = None
return FieldSet.from_nemo(
filenames, variables, dimensions,
indices=indices,
chunksize=field_chunksizes, # = None for no chunking
mesh='spherical',
tracer_interp_method='cgrid_tracer'
# ,time_periodic=time_loop_period
# ,allow_time_extrapolation=True
)
# + papermill={"duration": 0.023563, "end_time": "2021-04-23T11:48:55.769023", "exception": false, "start_time": "2021-04-23T11:48:55.745460", "status": "completed"} tags=[]
def create_fieldset(
data_path=data_path, experiment_name=experiment_name,
fname_U=fname_U, fname_V=fname_V, fname_W=fname_W, fname_T=fname_T,
mesh_mask = mesh_mask
):
files_U = list(sorted((data_path).glob(fname_U)))
files_V = list(sorted((data_path).glob(fname_V)))
files_W = list(sorted((data_path).glob(fname_W)))
files_T = list(sorted((data_path).glob(fname_T)))
print(files_U)
fieldset = fieldset_defintions(
files_U, files_V,
files_W, files_T, mesh_mask)
return fieldset
# + papermill={"duration": 280.78306, "end_time": "2021-04-23T11:53:36.570974", "exception": false, "start_time": "2021-04-23T11:48:55.787914", "status": "completed"} tags=[]
fieldset = create_fieldset()
# + [markdown] papermill={"duration": 0.019498, "end_time": "2021-04-23T11:53:36.610897", "exception": false, "start_time": "2021-04-23T11:53:36.591399", "status": "completed"} tags=[]
# ## Create Virtual Particles
# + [markdown] papermill={"duration": 0.01937, "end_time": "2021-04-23T11:53:36.649637", "exception": false, "start_time": "2021-04-23T11:53:36.630267", "status": "completed"} tags=[]
# #### add a couple of simple plotting routines
# + papermill={"duration": 0.02814, "end_time": "2021-04-23T11:53:36.697389", "exception": false, "start_time": "2021-04-23T11:53:36.669249", "status": "completed"} tags=[]
def plot_section_sdist():
plt.figure(figsize=(10,5))
u = np.array([p.uvel for p in pset]) * degree2km * 1000.0 * np.cos(np.radians(pset.lat))
v = np.array([p.vvel for p in pset]) * degree2km * 1000.0
section_index = np.searchsorted(lonlat.lon,pset.lon)-1
u_normal = v * lonlatdiff.costheta[section_index].data - u * lonlatdiff.sintheta[section_index].data
y = (pset.lat - lonlat.lat[section_index]) * degree2km
x = (pset.lon - lonlat.lon[section_index]) * degree2km*np.cos(np.radians(lonlat2mean.lat[section_index+1].data))
dist = np.sqrt(x**2 + y**2) + lonlatdiff.length_west[section_index].data
plt.scatter(
dist,
[p.depth for p in pset],
1,
u_normal,
cmap=co.cm.balance,vmin=-0.3,vmax=0.3
)
plt.ylim(1200,0)
plt.colorbar(label = r'normal velocity [$\mathrm{m\ s}^{-1}$]')
plt.xlabel('distance [km]')
plt.ylabel('depth [m]')
return
# + papermill={"duration": 0.025351, "end_time": "2021-04-23T11:53:36.742399", "exception": false, "start_time": "2021-04-23T11:53:36.717048", "status": "completed"} tags=[]
def plot_section_lon():
plt.figure(figsize=(10,5))
u = np.array([p.uvel for p in pset]) * degree2km * 1000.0 * np.cos(np.radians(pset.lat))
v = np.array([p.vvel for p in pset]) * degree2km * 1000.0
section_index = np.searchsorted(lonlat.lon,pset.lon)-1
u_normal = v * lonlatdiff.costheta[section_index].data - u * lonlatdiff.sintheta[section_index].data
plt.scatter(
[p.lon for p in pset],
[p.depth for p in pset],
1,
u_normal,
cmap=co.cm.balance,vmin=-0.3,vmax=0.3
)
plt.ylim(1200,0)
plt.colorbar(label = r'normal velocity [$\mathrm{m\ s}^{-1}$]');
plt.xlabel('longitude [$\degree$E]')
plt.ylabel('depth [m]')
return
# + papermill={"duration": 0.025549, "end_time": "2021-04-23T11:53:36.787929", "exception": false, "start_time": "2021-04-23T11:53:36.762380", "status": "completed"} tags=[]
class SampleParticle(JITParticle):
"""Add variables to the standard particle class.
Particles will sample temperature and track the age of the particle.
Particles also have a flag `alive` that is 1 if the particle is alive and 0 otherwise.
Furthermore, we have a `speed_param` that scales the velocity with which particles can
swim towards the surface.
Note that we don't initialize temp from the actual data.
This speeds up particle creation, but might render initial data point less useful.
"""
mxl = Variable('mxl', dtype=np.float32, initial=-100)
temp = Variable('temp', dtype=np.float32, initial=-100)
salt = Variable('salt', dtype=np.float32, initial=-100)
uvel = Variable('uvel', dtype=np.float32, initial=0)
vvel = Variable('vvel', dtype=np.float32, initial=0)
# wvel = Variable('wvel', dtype=np.float32, initial=0)
# alive = Variable('alive', dtype=np.int32, initial=1)
# speed_param = Variable('speed_param', dtype=np.float32, initial=1)
# age = Variable('age', dtype=np.int32, initial=0, to_write=True)
# + [markdown] papermill={"duration": 0.019704, "end_time": "2021-04-23T11:53:36.827545", "exception": false, "start_time": "2021-04-23T11:53:36.807841", "status": "completed"} tags=[]
# ## Create a set of particles with random initial positions
#
# We seed the RNG to be reproducible (and to be able to quickly create a second equivalent experiment with differently chosen compatible initial positions), and create arrays of random starting times, lats, lons, depths, and speed parameters (see kernel definitions below for details).
#
# Initially create points on 'rectangle'. Land points are removed later in a OceanParcels 'run' with runtime and timedelta zero.
# + [markdown] papermill={"duration": 0.019686, "end_time": "2021-04-23T11:53:36.867010", "exception": false, "start_time": "2021-04-23T11:53:36.847324", "status": "completed"} tags=[]
# ### First set up the piecewise section
# + papermill={"duration": 0.040124, "end_time": "2021-04-23T11:53:36.926942", "exception": false, "start_time": "2021-04-23T11:53:36.886818", "status": "completed"} tags=[]
lonlat = xr.Dataset(pd.read_csv(sectionPath / sectionFilename,delim_whitespace=True))
# + papermill={"duration": 0.264084, "end_time": "2021-04-23T11:53:37.211718", "exception": false, "start_time": "2021-04-23T11:53:36.947634", "status": "completed"} tags=[]
lonlat.lon.attrs['long_name']='Longitude'
lonlat.lat.attrs['long_name']='Latitude'
lonlat.lon.attrs['standard_name']='longitude'
lonlat.lat.attrs['standard_name']='latitude'
lonlat.lon.attrs['units']='degrees_east'
lonlat.lat.attrs['units']='degrees_north'
lonlatdiff = lonlat.diff('dim_0')
lonlat2mean= lonlat.rolling({'dim_0':2}).mean()
lonlat.plot.scatter(x='lon',y='lat')
lonlat2mean.plot.scatter(x='lon',y='lat')
lonlatdiff = lonlatdiff.assign({'y':lonlatdiff['lat']*degree2km})
lonlatdiff = lonlatdiff.assign({'x':lonlatdiff['lon']*degree2km*np.cos(np.radians(lonlat2mean.lat.data[1:]))})
lonlatdiff=lonlatdiff.assign({'length':np.sqrt(lonlatdiff['x']**2+lonlatdiff['y']**2)})
lonlatdiff=lonlatdiff.assign({'length_west':lonlatdiff.length.sum() - np.cumsum(lonlatdiff.length[::-1])[::-1]})
lonlatdiff=lonlatdiff.assign({'costheta':lonlatdiff['x']/lonlatdiff['length']})
lonlatdiff=lonlatdiff.assign({'sintheta':lonlatdiff['y']/lonlatdiff['length']})
total_length = lonlatdiff.length.sum().data
print(total_length)
# + papermill={"duration": 0.025517, "end_time": "2021-04-23T11:53:37.258536", "exception": false, "start_time": "2021-04-23T11:53:37.233019", "status": "completed"} tags=[]
lonlatdiff.length.shape[0]
# + [markdown] papermill={"duration": 0.020726, "end_time": "2021-04-23T11:53:37.299863", "exception": false, "start_time": "2021-04-23T11:53:37.279137", "status": "completed"} tags=[]
# ### Seed particles uniform random along OSNAP section
# + papermill={"duration": 0.12663, "end_time": "2021-04-23T11:53:37.446757", "exception": false, "start_time": "2021-04-23T11:53:37.320127", "status": "completed"} tags=[]
np.random.seed(RNG_seed)
# define time of release for each particle relative to t0
# can start each particle at a different time if required
# here all start at time t_start.
times = []
lons = []
lats = []
depths = []
# for subsect in range(lonlatdiff.length.shape[0]):
for subsect in range(start_vertex,end_vertex):
number_particles = int(create_number_particles*lonlatdiff.length[subsect]/total_length)
time = np.zeros(number_particles)
time += (t_start - t_0).total_seconds()
# start along a line from west to east
west_lat = lonlat.lat[subsect].data
west_lon = lonlat.lon[subsect].data
east_lat = lonlat.lat[subsect+1].data
east_lon = lonlat.lon[subsect+1].data
lon = np.random.uniform(
low=west_lon, high = east_lon,
size=time.shape
)
lat = west_lat + ((lon - west_lon) * (east_lat - west_lat)/ (east_lon - west_lon))
# at depths from surface to max_release_depth
depth = np.random.uniform(
low=min_release_depth, high=max_release_depth,
size=time.shape
)
times.append(time)
lons.append(lon)
lats.append(lat)
depths.append(depth)
time = np.concatenate(times)
lon = np.concatenate(lons)
lat = np.concatenate(lats)
depth = np.concatenate(depths)
# + [markdown] papermill={"duration": 0.020667, "end_time": "2021-04-23T11:53:37.488685", "exception": false, "start_time": "2021-04-23T11:53:37.468018", "status": "completed"} tags=[]
# ### Build particle set
# + papermill={"duration": 0.92072, "end_time": "2021-04-23T11:53:38.430105", "exception": false, "start_time": "2021-04-23T11:53:37.509385", "status": "completed"} tags=[]
# %%time
pset = ParticleSet(
fieldset=fieldset,
pclass=SampleParticle,
lat=lat,
lon=lon,
# speed_param=speed_param,
depth=depth,
time=time
# repeatdt = repeatdt
)
# + papermill={"duration": 0.025444, "end_time": "2021-04-23T11:53:38.477051", "exception": false, "start_time": "2021-04-23T11:53:38.451607", "status": "completed"} tags=[]
print(f"Created {len(pset)} particles.")
# display(pset[:5])
# display(pset[-5:])
# + [markdown] papermill={"duration": 0.020978, "end_time": "2021-04-23T11:53:38.518851", "exception": false, "start_time": "2021-04-23T11:53:38.497873", "status": "completed"} tags=[]
# ## Compose custom kernel
#
# We'll create three additional kernels:
# - One Kernel adds velocity sampling
# - One Kernel adds temperature sampling
# - One kernel adds salinity sampling
#
# Then, we combine the builtin `AdvectionRK4_3D` kernel with these additional kernels.
# + papermill={"duration": 0.025406, "end_time": "2021-04-23T11:53:38.565202", "exception": false, "start_time": "2021-04-23T11:53:38.539796", "status": "completed"} tags=[]
def velocity_sampling(particle, fieldset, time):
'''Sample velocity.'''
(particle.uvel,particle.vvel) = fieldset.UV[time, particle.depth, particle.lat, particle.lon]
# + papermill={"duration": 0.024873, "end_time": "2021-04-23T11:53:38.611159", "exception": false, "start_time": "2021-04-23T11:53:38.586286", "status": "completed"} tags=[]
def temperature_sampling(particle, fieldset, time):
'''Sample temperature.'''
particle.temp = fieldset.T[time, particle.depth, particle.lat, particle.lon]
# + papermill={"duration": 0.025025, "end_time": "2021-04-23T11:53:38.657255", "exception": false, "start_time": "2021-04-23T11:53:38.632230", "status": "completed"} tags=[]
def salinity_sampling(particle, fieldset, time):
'''Sample salinity.'''
particle.salt = fieldset.S[time, particle.depth, particle.lat, particle.lon]
# + papermill={"duration": 0.024952, "end_time": "2021-04-23T11:53:38.703310", "exception": false, "start_time": "2021-04-23T11:53:38.678358", "status": "completed"} tags=[]
def mxl_sampling(particle, fieldset, time):
'''Sample mixed layer depth.'''
particle.mxl = fieldset.MXL[time, particle.depth, particle.lat, particle.lon]
# + papermill={"duration": 0.257752, "end_time": "2021-04-23T11:53:38.982502", "exception": false, "start_time": "2021-04-23T11:53:38.724750", "status": "completed"} tags=[]
custom_kernel = (
pset.Kernel(AdvectionRK4_3D)
# + pset.Kernel(temperature_sensitivity)
+ pset.Kernel(temperature_sampling)
+ pset.Kernel(salinity_sampling)
+ pset.Kernel(velocity_sampling)
+ pset.Kernel(mxl_sampling)
)
# + [markdown] papermill={"duration": 0.02129, "end_time": "2021-04-23T11:53:39.025721", "exception": false, "start_time": "2021-04-23T11:53:39.004431", "status": "completed"} tags=[]
# ## Be able to handle errors during integration
#
# We have restricted our domain so in principle, particles could reach undefined positions.
# In that case, we want to just delete the particle (without forgetting its history).
# + papermill={"duration": 0.026506, "end_time": "2021-04-23T11:53:39.074065", "exception": false, "start_time": "2021-04-23T11:53:39.047559", "status": "completed"} tags=[]
def DeleteParticle(particle, fieldset, time):
particle.delete()
recovery_cases = {
ErrorCode.ErrorOutOfBounds: DeleteParticle,
ErrorCode.Error: DeleteParticle,
ErrorCode.ErrorInterpolation: DeleteParticle
}
# + [markdown] papermill={"duration": 0.021499, "end_time": "2021-04-23T11:53:39.117079", "exception": false, "start_time": "2021-04-23T11:53:39.095580", "status": "completed"} tags=[]
# ## Run with runtime=0 to initialise fields
# + papermill={"duration": 230.359976, "end_time": "2021-04-23T11:57:29.498491", "exception": false, "start_time": "2021-04-23T11:53:39.138515", "status": "completed"} tags=[]
# %%time
# with dask.config.set(**{'array.slicing.split_large_chunks': False}):
pset.execute(
custom_kernel,
runtime=0,
# dt=timedelta(minutes=0),
# output_file=outputfile,
recovery=recovery_cases
)
# + papermill={"duration": 41.999956, "end_time": "2021-04-23T11:58:11.520812", "exception": false, "start_time": "2021-04-23T11:57:29.520856", "status": "completed"} tags=[]
plot_section_sdist()
# + [markdown] papermill={"duration": 0.025156, "end_time": "2021-04-23T11:58:11.571265", "exception": false, "start_time": "2021-04-23T11:58:11.546109", "status": "completed"} tags=[]
# ## Trim unwanted points from ParticleSet
#
# Use initialised fields to remove land points. We test `temp == 0.0` (the mask value over land).
#
# + papermill={"duration": 5.436886, "end_time": "2021-04-23T11:58:17.033070", "exception": false, "start_time": "2021-04-23T11:58:11.596184", "status": "completed"} tags=[]
t = np.array([p.temp for p in pset])
# u = np.array([p.uvel for p in pset])
# v = np.array([p.vvel for p in pset])
pset.remove_indices(np.argwhere(t == 0).flatten())
# pset.remove(np.argwhere(x * y * z == 0).flatten())
print(len(pset))
# + papermill={"duration": 33.447636, "end_time": "2021-04-23T11:58:50.511102", "exception": false, "start_time": "2021-04-23T11:58:17.063466", "status": "completed"} tags=[]
plot_section_sdist()
# + [markdown] papermill={"duration": 0.028471, "end_time": "2021-04-23T11:58:50.568595", "exception": false, "start_time": "2021-04-23T11:58:50.540124", "status": "completed"} tags=[]
# ### Test velocity normal to section
# + [markdown] papermill={"duration": 0.028324, "end_time": "2021-04-23T11:58:50.625485", "exception": false, "start_time": "2021-04-23T11:58:50.597161", "status": "completed"} tags=[]
# #### Velocity conversions from degrees lat/lon per second to m/s
# + papermill={"duration": 8.256567, "end_time": "2021-04-23T11:58:58.910587", "exception": false, "start_time": "2021-04-23T11:58:50.654020", "status": "completed"} tags=[]
u = np.array([p.uvel for p in pset])
v = np.array([p.vvel for p in pset])
# + papermill={"duration": 0.068444, "end_time": "2021-04-23T11:58:59.011749", "exception": false, "start_time": "2021-04-23T11:58:58.943305", "status": "completed"} tags=[]
u=u * degree2km * 1000.0 * np.cos(np.radians(pset.lat))
v=v * degree2km * 1000.0
# + [markdown] papermill={"duration": 0.029097, "end_time": "2021-04-23T11:58:59.070408", "exception": false, "start_time": "2021-04-23T11:58:59.041311", "status": "completed"} tags=[]
# #### normal velocities
# + papermill={"duration": 0.082545, "end_time": "2021-04-23T11:58:59.182045", "exception": false, "start_time": "2021-04-23T11:58:59.099500", "status": "completed"} tags=[]
section_index = np.searchsorted(lonlat.lon,pset.lon)-1
u_normal = v * lonlatdiff.costheta[section_index].data - u * lonlatdiff.sintheta[section_index].data
# + papermill={"duration": 0.038215, "end_time": "2021-04-23T11:58:59.249653", "exception": false, "start_time": "2021-04-23T11:58:59.211438", "status": "completed"} tags=[]
abs(u_normal).max()
# + [markdown] papermill={"duration": 0.028632, "end_time": "2021-04-23T11:58:59.307064", "exception": false, "start_time": "2021-04-23T11:58:59.278432", "status": "completed"} tags=[]
# #### remove particles randomly with probability proportional to normal speed
# + papermill={"duration": 0.204119, "end_time": "2021-04-23T11:58:59.540203", "exception": false, "start_time": "2021-04-23T11:58:59.336084", "status": "completed"} tags=[]
u_random = np.random.rand(len(u_normal))*max_current
pset.remove_indices(np.argwhere(abs(u_normal) < u_random).flatten())
print(len(pset))
# + papermill={"duration": 1.203511, "end_time": "2021-04-23T11:59:00.773311", "exception": false, "start_time": "2021-04-23T11:58:59.569800", "status": "completed"} tags=[]
plot_section_sdist()
# + [markdown] papermill={"duration": 0.032438, "end_time": "2021-04-23T11:59:00.840906", "exception": false, "start_time": "2021-04-23T11:59:00.808468", "status": "completed"} tags=[]
# ## Prepare output
#
# We define an output file and specify the desired output frequency.
# + papermill={"duration": 0.044628, "end_time": "2021-04-23T11:59:00.918839", "exception": false, "start_time": "2021-04-23T11:59:00.874211", "status": "completed"} tags=[]
# output_filename = 'Parcels_IFFForwards_1m_June2016_2000.nc'
npart = str(len(pset))
output_filename = 'tracks_randomvel_mxl_'+sectionname+direction+year_str+month_str+day_str+'_N'+npart+'_D'+days+'_Rnd'+ seed+'.nc'
outfile = outpath / output_filename
print(outfile)
outputfile = pset.ParticleFile(
name=outfile,
outputdt=timedelta(hours=outputdt_in_hours)
)
# + [markdown] papermill={"duration": 0.033319, "end_time": "2021-04-23T11:59:00.985761", "exception": false, "start_time": "2021-04-23T11:59:00.952442", "status": "completed"} tags=[]
# ## Execute the experiment
#
# We'll evolve particles, log their positions and variables to the output buffer and finally export the output to a the file.
# + [markdown] papermill={"duration": 0.033425, "end_time": "2021-04-23T11:59:01.052590", "exception": false, "start_time": "2021-04-23T11:59:01.019165", "status": "completed"} tags=[]
# ### Run the experiment
# + papermill={"duration": 70465.369189, "end_time": "2021-04-24T07:33:26.455348", "exception": false, "start_time": "2021-04-23T11:59:01.086159", "status": "completed"} tags=[]
# %%time
# with dask.config.set(**{'array.slicing.split_large_chunks': False}):
pset.execute(
custom_kernel,
runtime=timedelta(days=runtime_in_days),
dt=timedelta(minutes=dt_in_minutes),
output_file=outputfile,
recovery=recovery_cases
)
# + papermill={"duration": 0.209089, "end_time": "2021-04-24T07:33:26.867280", "exception": false, "start_time": "2021-04-24T07:33:26.658191", "status": "completed"} tags=[]
# outputfile.export()
# + papermill={"duration": 179.370177, "end_time": "2021-04-24T07:36:26.439918", "exception": false, "start_time": "2021-04-24T07:33:27.069741", "status": "completed"} tags=[]
outputfile.close()
# + papermill={"duration": 2.774444, "end_time": "2021-04-24T07:36:30.188999", "exception": false, "start_time": "2021-04-24T07:36:27.414555", "status": "completed"} tags=[]
conda list
# + papermill={"duration": 1.06214, "end_time": "2021-04-24T07:36:31.454302", "exception": false, "start_time": "2021-04-24T07:36:30.392162", "status": "completed"} tags=[]
pip list
| notebooks/executed/037_afox_RunParcels_TS_MXL_Multiline_Randomvel_Papermill_executed_2019-02-22.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp util
# -
# # Utils for illumination
#
# > API details.
# +
#export
import torch as th
import torch.nn as nn
from smpr3d.util import fftshift_checkerboard, Param
import numpy as np
# def zernike_aberrations(qx, qy, lam, C):
# """
# Calculates the geometrical aberrations up to the order specified in C. C is a complex array specifying the
# aberration coefficients.
#
# :param qx: x coordinate system, can be (i_batch, qxx, qxy)
# :param qy: y coordinate system, can be (i_batch, qyx, qyy)
# :param lam: wavelength in A
# :param C: (n_batch, m_max, n_max) complex array
# :return: chi, the real aberration function
# """
# if isinstance(qx, np.ndarray):
# xp = np
#
# if C.ndim == 3:
# n_batch, m_max, n_max = C.shape
# elif C.ndim == 2:
# m_max, n_max = C.shape
# n_batch = 1
# C = C[None, ...]
# else:
# raise RuntimeError("Aberration coefficient array has wrong dimension")
#
# sh = (n_batch, *qx.shape)
# chi = xp.zeros(sh, dtype=np.float32) + 1j * xp.zeros(sh, dtype=np.float32)
# ω = lam * (qx + 1j * qy)
#
# for i_batch in range(n_batch):
# for m in range(m_max):
# if m <= 3:
# n_max = m + 1
# elif m <= 5:
# n_max = m // 2
# else:
# n_max = 1
# for n in range(n_max):
# if m > 0 or n > 0:
# # print(f"C[{m}, {n}] = {C[i_batch, m, n]}")
# ab = C[i_batch, m, n] * (ω ** m) * (ω.conj() ** n)
# ab /= (m + n)
# chi[i_batch] += ab
# chi = 2 * np.pi / lam * chi.real
# chi = xp.squeeze(chi)
# elif isinstance(qx, th.Tensor):
# if C.ndimension() == 4:
# n_batch, m_max, n_max, _ = C.shape
# elif C.ndimension() == 3:
# m_max, n_max, _ = C.shape
# n_batch = 1
# C = C.unsqueeze(0)
# else:
# raise RuntimeError("Aberration coefficient array has wrong dimension")
#
# sh = (n_batch, *qx.shape)
#
# chi = th.zeros(sh, dtype=th.float32)
#
# # print(chi.shape)
#
# ω = th.zeros_like(qx)
# ω[re] = lam * qx
# ω[im] = lam * qy
# C_mn = make_real(th.zeros(qx.shape))
# for i_batch in range(n_batch):
# for m in range(m_max):
# if m <= 3:
# n_max = m + 1
# elif m <= 5:
# n_max = m // 2
# else:
# n_max = 1
# for n in range(n_max):
# if m > 0 or n > 0:
# # print(f"C[{m}, {n}] = {C[i_batch, m, n]}")
# C_mn[re] = C[i_batch, m, n][re]
# C_mn[im] = C[i_batch, m, n][im]
# ab = complex_mul(C_mn, cpow(ω, n))
# ab = complex_mul(ab, cpow(conj(ω), n))
# ab /= (m + n)
# # print(chi.shape)
# chi[i_batch] += ab
# chi = 2 * np.pi / lam * chi[re]
# chi = chi.squeeze()
# return chi
def cartesian_aberrations_single(qx, qy, lam, C):
"""
Zernike polynomials in the cartesian coordinate system
:param qx:
:param qy:
:param lam: wavelength in Angstrom
:param C: (12 ,)
:return:
"""
u = qx * lam
v = qy * lam
u2 = u ** 2
u3 = u ** 3
u4 = u ** 4
# u5 = u ** 5
v2 = v ** 2
v3 = v ** 3
v4 = v ** 4
# v5 = v ** 5
aberr = Param()
aberr.C1 = C[0]
aberr.C12a = C[1]
aberr.C12b = C[2]
aberr.C21a = C[3]
aberr.C21b = C[4]
aberr.C23a = C[5]
aberr.C23b = C[6]
aberr.C3 = C[7]
aberr.C32a = C[8]
aberr.C32b = C[9]
aberr.C34a = C[10]
aberr.C34b = C[11]
chi = 0
# r-2 = x-2 +y-2.
chi += 1 / 2 * aberr.C1 * (u2 + v2) # r^2
#r-2 cos(2*phi) = x"2 -y-2.
# r-2 sin(2*phi) = 2*x*y.
chi += 1 / 2 * (aberr.C12a * (u2 - v2) + 2 * aberr.C12b * u * v) # r^2 cos(2 phi) + r^2 sin(2 phi)
# r-3 cos(3*phi) = x-3 -3*x*y'2. r"3 sin(3*phi) = 3*y*x-2 -y-3.
chi += 1 / 3 * (aberr.C23a * (u3 - 3 * u * v2) + aberr.C23b * (3 * u2 * v - v3))# r^3 cos(3phi) + r^3 sin(3 phi)
# r-3 cos(phi) = x-3 +x*y-2.
# r-3 sin(phi) = y*x-2 +y-3.
chi += 1 / 3 * (aberr.C21a * (u3 + u * v2) + aberr.C21b * (v3 + u2 * v))# r^3 cos(phi) + r^3 sin(phi)
# r-4 = x-4 +2*x-2*y-2 +y-4.
chi += 1 / 4 * aberr.C3 * (u4 + v4 + 2 * u2 * v2)# r^4
# r-4 cos(4*phi) = x-4 -6*x-2*y-2 +y-4.
chi += 1 / 4 * aberr.C34a * (u4 - 6 * u2 * v2 + v4)# r^4 cos(4 phi)
# r-4 sin(4*phi) = 4*x-3*y -4*x*y-3.
chi += 1 / 4 * aberr.C34b * (4 * u3 * v - 4 * u * v3) # r^4 sin(4 phi)
# r-4 cos(2*phi) = x-4 -y-4.
chi += 1 / 4 * aberr.C32a * (u4 - v4)
# r-4 sin(2*phi) = 2*x-3*y +2*x*y-3.
chi += 1 / 4 * aberr.C32b * (2 * u3 * v + 2 * u * v3)
# r-5 cos(phi) = x-5 +2*x-3*y-2 +x*y-4.
# r-5 sin(phi) = y*x"4 +2*x-2*y-3 +y-5.
# r-5 cos(3*phi) = x-5 -2*x-3*y-2 -3*x*y-4.
# r-5 sin(3*phi) = 3*y*x-4 +2*x-2*y-3 -y-5.
# r-5 cos(5*phi) = x-5 -10*x-3*y-2 +5*x*y-4.
# r-5 sin(5*phi) = 5*y*x-4 -10*x-2*y-3 +y-5.
chi *= 2 * np.pi / lam
return chi
def cartesian_aberrations(qx, qy, lam, C):
"""
Zernike polynomials in the cartesian coordinate system
:param qx:
:param qy:
:param lam: wavelength in Angstrom
:param C: 12 x D
:return:
"""
u = qx * lam
v = qy * lam
u2 = u ** 2
u3 = u ** 3
u4 = u ** 4
# u5 = u ** 5
v2 = v ** 2
v3 = v ** 3
v4 = v ** 4
# v5 = v ** 5
aberr = Param()
aberr.C1 = C[0].unsqueeze(1).unsqueeze(1)
aberr.C12a = C[1].unsqueeze(1).unsqueeze(1)
aberr.C12b = C[2].unsqueeze(1).unsqueeze(1)
aberr.C21a = C[3].unsqueeze(1).unsqueeze(1)
aberr.C21b = C[4].unsqueeze(1).unsqueeze(1)
aberr.C23a = C[5].unsqueeze(1).unsqueeze(1)
aberr.C23b = C[6].unsqueeze(1).unsqueeze(1)
aberr.C3 = C[7].unsqueeze(1).unsqueeze(1)
aberr.C32a = C[8].unsqueeze(1).unsqueeze(1)
aberr.C32b = C[9].unsqueeze(1).unsqueeze(1)
aberr.C34a = C[10].unsqueeze(1).unsqueeze(1)
aberr.C34b = C[11].unsqueeze(1).unsqueeze(1)
chi = 0
# r-2 = x-2 +y-2.
chi += 1 / 2 * aberr.C1 * (u2 + v2) # r^2
#r-2 cos(2*phi) = x"2 -y-2.
# r-2 sin(2*phi) = 2*x*y.
chi += 1 / 2 * (aberr.C12a * (u2 - v2) + 2 * aberr.C12b * u * v) # r^2 cos(2 phi) + r^2 sin(2 phi)
# r-3 cos(3*phi) = x-3 -3*x*y'2. r"3 sin(3*phi) = 3*y*x-2 -y-3.
chi += 1 / 3 * (aberr.C23a * (u3 - 3 * u * v2) + aberr.C23b * (3 * u2 * v - v3))# r^3 cos(3phi) + r^3 sin(3 phi)
# r-3 cos(phi) = x-3 +x*y-2.
# r-3 sin(phi) = y*x-2 +y-3.
chi += 1 / 3 * (aberr.C21a * (u3 + u * v2) + aberr.C21b * (v3 + u2 * v))# r^3 cos(phi) + r^3 sin(phi)
# r-4 = x-4 +2*x-2*y-2 +y-4.
chi += 1 / 4 * aberr.C3 * (u4 + v4 + 2 * u2 * v2)# r^4
# r-4 cos(4*phi) = x-4 -6*x-2*y-2 +y-4.
chi += 1 / 4 * aberr.C34a * (u4 - 6 * u2 * v2 + v4)# r^4 cos(4 phi)
# r-4 sin(4*phi) = 4*x-3*y -4*x*y-3.
chi += 1 / 4 * aberr.C34b * (4 * u3 * v - 4 * u * v3) # r^4 sin(4 phi)
# r-4 cos(2*phi) = x-4 -y-4.
chi += 1 / 4 * aberr.C32a * (u4 - v4)
# r-4 sin(2*phi) = 2*x-3*y +2*x*y-3.
chi += 1 / 4 * aberr.C32b * (2 * u3 * v + 2 * u * v3)
# r-5 cos(phi) = x-5 +2*x-3*y-2 +x*y-4.
# r-5 sin(phi) = y*x"4 +2*x-2*y-3 +y-5.
# r-5 cos(3*phi) = x-5 -2*x-3*y-2 -3*x*y-4.
# r-5 sin(3*phi) = 3*y*x-4 +2*x-2*y-3 -y-5.
# r-5 cos(5*phi) = x-5 -10*x-3*y-2 +5*x*y-4.
# r-5 sin(5*phi) = 5*y*x-4 -10*x-2*y-3 +y-5.
chi *= 2 * np.pi / lam
return chi
def aperture(q: th.Tensor, lam, alpha_max, edge=2):
ktheta = th.asin(q.norm(dim=0) * lam)
qmax = alpha_max / lam
dk = q[0][1][0]
arr = th.zeros_like(q[1])
arr[ktheta < alpha_max] = 1
if edge > 0:
dEdge = edge / (qmax / dk) # fraction of aperture radius that will be smoothed
# some fancy indexing: pull out array elements that are within
# our smoothing edges
ind = (ktheta / alpha_max > (1 - dEdge)) * (ktheta / alpha_max < (1 + dEdge))
arr[ind] = 0.5 * (1 - th.sin(np.pi / (2 * dEdge) * (ktheta[ind] / alpha_max - 1)))
return arr
class ZernikeProbe2(nn.Module):
def __init__(self, q: th.Tensor, lam, fft_shifted=True):
"""
Creates an aberration surface from aberration coefficients. The output is backpropable
:param q: 2 x M1 x M2 tensor of x coefficients of reciprocal space
:param lam: wavelength in Angstrom
:param C: aberration coefficients
:return: (maximum size of all aberration tensors) x MY x MX
"""
super(ZernikeProbe2, self).__init__()
self.q = q
self.lam = lam
self.fft_shifted = fft_shifted
if self.fft_shifted:
cb = fftshift_checkerboard(self.q.shape[1] // 2, self.q.shape[2] // 2)
self.cb = th.from_numpy(cb).float().to(q.device)
def forward(self, C, A):
chi = cartesian_aberrations(self.q[1], self.q[0], self.lam, C)
Psi = th.exp(-1j*chi) * A.expand_as(chi)
if self.fft_shifted:
Psi = Psi * self.cb
return Psi
class ZernikeProbeSingle(nn.Module):
def __init__(self, q: th.Tensor, lam, fft_shifted=True):
"""
Creates an aberration surface from aberration coefficients. The output is backpropable
:param q: 2 x M1 x M2 tensor of x coefficients of reciprocal space
:param lam: wavelength in Angstrom
:param C: aberration coefficients
:return: (maximum size of all aberration tensors) x MY x MX
"""
super(ZernikeProbeSingle, self).__init__()
self.q = q
self.lam = lam
self.fft_shifted = fft_shifted
if self.fft_shifted:
cb = fftshift_checkerboard(self.q.shape[1] // 2, self.q.shape[2] // 2)
self.cb = th.from_numpy(cb).float().to(q.device)
def forward(self, C, A):
chi = cartesian_aberrations_single(self.q[1], self.q[0], self.lam, C)
Psi = th.exp(-1j*chi) * A.expand_as(chi)
if self.fft_shifted:
Psi = Psi * self.cb
return Psi
class ZernikeProbe(nn.Module):
def __init__(self, q: th.Tensor, lam, A_init, A_requires_grad, fft_shifted=True,
C1=th.zeros(1), C12a=th.zeros(1), C12b=th.zeros(1), C21a=th.zeros(1), C21b=th.zeros(1),
C23a=th.zeros(1), C23b=th.zeros(1), C3=th.zeros(1), C32a=th.zeros(1), C32b=th.zeros(1),
C34a=th.zeros(1), C34b=th.zeros(1)):
"""
Creates an aberration surface from aberration coefficients. The output is backpropable
:param qx: M1 x M2 tensor of x coefficients of reciprocal space
:param qy: M1 x M2 tensor of y coefficients of reciprocal space
:param lam: wavelength in Angstrom
:param C: shift in Angstrom
:return: (maximum size of all aberration tensors) x MY x MX
"""
super(ZernikeProbe, self).__init__()
self.qx = q[1]
self.qy = q[0]
self.lam = lam
self.fft_shifted = fft_shifted
self.dtype = q.dtype
d = self.dtype
# defocus
self.C1 = nn.Parameter(data=C1.type(d), requires_grad=th.max(th.abs(C1)).item() > 1e-10)
# two-fold stig
self.C12a = nn.Parameter(data=C12a.type(d), requires_grad=th.max(th.abs(C12a)).item() > 1e-10).to(C1.device)
self.C12b = nn.Parameter(data=C12b.type(d), requires_grad=th.max(th.abs(C12b)).item() > 1e-10).to(C1.device)
# axial coma
self.C21a = nn.Parameter(data=C21a.type(d), requires_grad=th.max(th.abs(C21a)).item() > 1e-10).to(C1.device)
self.C21b = nn.Parameter(data=C21b.type(d), requires_grad=th.max(th.abs(C21b)).item() > 1e-10).to(C1.device)
# 3-fold stig
self.C23a = nn.Parameter(data=C23a.type(d), requires_grad=th.max(th.abs(C23a)).item() > 1e-10).to(C1.device)
self.C23b = nn.Parameter(data=C23b.type(d), requires_grad=th.max(th.abs(C23b)).item() > 1e-10).to(C1.device)
# spherical
self.C3 = nn.Parameter(data=C3.type(d), requires_grad=th.max(th.abs(C3)).item() > 1e-10).to(C1.device)
# star
self.C32a = nn.Parameter(data=C32a.type(d), requires_grad=th.max(th.abs(C32a)).item() > 1e-10).to(C1.device)
self.C32b = nn.Parameter(data=C32b.type(d), requires_grad=th.max(th.abs(C32b)).item() > 1e-10).to(C1.device)
# 4-fold stig
self.C34a = nn.Parameter(data=C34a.type(d), requires_grad=th.max(th.abs(C34a)).item() > 1e-10).to(C1.device)
self.C34b = nn.Parameter(data=C34b.type(d), requires_grad=th.max(th.abs(C34b)).item() > 1e-10).to(C1.device)
self.A = nn.Parameter(data=A_init.type(d), requires_grad=A_requires_grad).to(C1.device)
self.optimized_parameters_ = []
if self.fft_shifted:
self.cb = th.from_numpy(fftshift_checkerboard(self.qx.shape[0] // 2, self.qx.shape[1] // 2)).float().to(
q.device).type(d)
for p in self.parameters():
if p.requires_grad:
self.optimized_parameters_.append(p)
def optimized_parameters(self):
return self.optimized_parameters_
def forward(self):
max_size = np.max(
[a.shape[0] if len(a.shape) > 0 else 1 for a in
[self.C1, self.C12a, self.C12b, self.C21a, self.C21b, self.C23a, self.C23b, self.C3,
self.C32a, self.C32b, self.C34a, self.C34b]])
C = th.stack((self.C1.expand(max_size), self.C12a.expand(max_size), self.C12b.expand(max_size),
self.C21a.expand(max_size), self.C21b.expand(max_size), self.C23a.expand(max_size),
self.C23b.expand(max_size), self.C3.expand(max_size), self.C32a.expand(max_size),
self.C32b.expand(max_size), self.C34a.expand(max_size), self.C34b.expand(max_size)), 0)
chi = cartesian_aberrations(self.qx, self.qy, self.lam, C).type(self.dtype)
Psi = th.exp(-1j*chi) * self.A.expand_as(chi)
if self.fft_shifted:
Psi = Psi * self.cb
return Psi
| nbs/01d_util_illumination.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import init
from SBMLLint.common import constants as cn
from SBMLLint.common.molecule import Molecule, MoleculeStoichiometry
from SBMLLint.common import simple_sbml
from SBMLLint.common.reaction import Reaction
from SBMLLint.tools import sbmllint
from SBMLLint.tools import print_reactions
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# +
from games_setup import *
from SBMLLint.common import constants as cn
from SBMLLint.common.simple_sbml import SimpleSBML
from SBMLLint.common.stoichiometry_matrix import StoichiometryMatrix
from SBMLLint.games.som import SOM
from SBMLLint.games.games_pp import GAMES_PP, SOMStoichiometry, SOMReaction, TOLERANCE
from SBMLLint.games.games_report import GAMESReport, SimplifiedReaction
import matplotlib.pyplot as plt
# %matplotlib inline
import time
from scipy.linalg import lu, inv
# -
data_dir = '/Users/woosubs/Desktop/ModelEngineering/ext_biomodels'
# we can remove EXCEPTIONS from files, as they are not loaded by simpleSBML
files = [f for f in os.listdir(data_dir) if f[-4:] == ".xml"]
len(files)
# data frame structure
# statistics columns
NUM_REACTIONS = "num_reactions(nonbdry)"
LP_ERROR = "lp_error"
GAMES_ERROR = "games_error"
GAMESPP_ERROR = "gamespp_error"
TYPEI_ERROR = "type1_error"
TYPEII_ERROR = "type2_error"
CANCELING_ERROR = "canceling_error"
ECHELON_ERROR = "echelon_error"
TYPEIII_ERROR = "type3_error"
result_columns = [NUM_REACTIONS,
LP_ERROR,
GAMES_ERROR,
GAMESPP_ERROR,
TYPEI_ERROR,
TYPEII_ERROR,
CANCELING_ERROR,
ECHELON_ERROR,
TYPEIII_ERROR]
results = pd.DataFrame(0, index=files, columns=result_columns)
results[:5]
# +
# cannot be initialized by simpleSBML; bad antimony string
EXCEPTIONS = ["BIOMD0000000146_url.xml",
"BIOMD0000000152_url.xml",
"BIOMD0000000608_url.xml",
"BIOMD0000000620_url.xml",
"BIOMD0000000634_url.xml",
]
# simple.initialize(os.path.join(data_dir, EXCEPTIONS[4]))
# s = StoichiometryMatrix(simple)
# num_reactions = s.stoichiometry_matrix.shape[1]
# results.at[file, NUM_REACTIONS] = num_reactions
# if num_reactions:
# consistent = s.isConsistent()
# -
files[0]
files[0][-7:-4]
simple = SimpleSBML()
model_147 = [x for x in files if x[-7:-4] == '147'][0]
simple.initialize(os.path.join(data_dir, model_147))
m = GAMES_PP(simple)
res = m.analyze(simple_games=False, error_details=True, suppress_message=False)
gr = GAMESReport(m)
for reaction in m.reactions:
print(reaction)
for str_obj in gr.reportTypeOneError(m.type_one_errors):
print(str_obj)
s = StoichiometryMatrix(simple)
s.isConsistent()
col_of_interest = s.stoichiometry_matrix.T['IkBa_cytoplasm']
nonzero_reactions = col_of_interest[col_of_interest.to_numpy().nonzero()[0]]
s.stoichiometry_matrix[nonzero_reactions.index[3]]
# +
# LP only
# suppress warnings;
import warnings
warnings.filterwarnings('ignore')
simple = SimpleSBML()
count = 0
lp_start = time.time()
for file in files:
count += 1
if (count%100)==0:
print("we are analyzing Model number:", count)
try:
simple.initialize(os.path.join(data_dir, file))
s = StoichiometryMatrix(simple)
num_reactions = s.stoichiometry_matrix.shape[1]
results.at[file, NUM_REACTIONS] = num_reactions
if num_reactions:
consistent = s.isConsistent()
else:
consistent = -1
results.at[file, LP_ERROR] = 1 - int(consistent)
except:
results.at[file, LP_ERROR] = -1
lp_end = time.time()
lp_time = lp_end - lp_start
print("Analysis finished!")
print("LP time:", lp_time)
# -
lp_results = results[results[LP_ERROR] == 1]
len(lp_results)
print("(Mean) ISS for LP is:", np.mean(lp_results[NUM_REACTIONS]))
print("(STD) ISS for LP is:", np.std(lp_results[NUM_REACTIONS]))
len(results[results[LP_ERROR]==1])
results[results[LP_ERROR]==-1]
# simple bGAMES only
simple = SimpleSBML()
count = 0
games_start = time.time()
for file in files:
count += 1
if (count%100)==0:
print("we are analyzing Model number:", count)
try:
simple.initialize(os.path.join(data_dir, file))
m = GAMES_PP(simple)
if simple.reactions:
res = m.analyze(simple_games=True, error_details=False, suppress_message=True)
results.at[file, GAMES_ERROR] = int(res)
if res:
gr = GAMESReport(m)
summary = m.error_summary
if m.type_one_errors:
results.at[file, TYPEI_ERROR] = len(m.type_one_errors)
report, error_num = gr.reportTypeOneError(m.type_one_errors, explain_details=True)
if m.type_two_errors:
results.at[file, TYPEII_ERROR] = len(m.type_two_errors)
report, error_num = gr.reportTypeTwoError(m.type_two_errors, explain_details=True)
except:
results.at[file, GAMES_ERROR] = -1
games_end = time.time()
games_time = games_end - games_start
print("Analysis finished!")
print("GAMES time:", games_time)
print("number of detected errors: ", len(results[results[GAMES_ERROR]==1]))
print("number of simple GAMES but not in LP", len(results[(results[GAMES_ERROR]==1) & (results[LP_ERROR]!=1)]))
123/158
# GAMES+
# file, GAMES_ERROR coding:
# 0; normal - no error found
# -1; not loaded or error found
# 1; normal - error found
# 2; echelon error found, but it is not explainable
# 3; type III error found, but it is not explainable
simple = SimpleSBML()
count = 0
gamespp_start = time.time()
for file in files:
count += 1
if (count%100)==0:
print("we are analyzing Model number:", count)
try:
simple.initialize(os.path.join(data_dir, file))
m = GAMES_PP(simple)
if simple.reactions:
res = m.analyze(simple_games=False, error_details=False, suppress_message=True)
results.at[file, GAMESPP_ERROR] = int(res)
if res:
# if m.echelon_errors or m.type_three_errors:
# try:
# #k = inv(m.lower)
# k = np.linalg.inv(m.lower)
# except:
# print("model %s has as a singular L matrix:" % file)
# condition_number = np.linalg.cond(m.lower)
# if condition_number > 300:
# print("*****The L matrix of the model %s has a condition number %f*****" % (file, condition_number))
gr = GAMESReport(m)
summary = m.error_summary
if m.type_one_errors:
results.at[file, TYPEI_ERROR] = len(m.type_one_errors)
report, error_num = gr.reportTypeOneError(m.type_one_errors, explain_details=True)
if m.type_two_errors:
results.at[file, TYPEII_ERROR] = len(m.type_two_errors)
report, error_num = gr.reportTypeTwoError(m.type_two_errors, explain_details=True)
if m.canceling_errors:
results.at[file, CANCELING_ERROR] = len(m.canceling_errors)
report, error_num = gr.reportCancelingError(m.canceling_errors, explain_details=True)
if m.echelon_errors:
#print("Model %s has an echelon error:" % file)
results.at[file, ECHELON_ERROR] = len(m.echelon_errors)
report, error_num = gr.reportEchelonError(m.echelon_errors, explain_details=True)
if report is False:
results.at[file, GAMESPP_ERROR] = 2
# print("Model %s has an unexplainable Echelon Error" % file)
# print("As the lower matrix has a condition number %f" % condition_number)
# print("Decide if the matrix is invertible")
if m.type_three_errors:
#print("Model %s has a type III error:" % file)
results.at[file, TYPEIII_ERROR] = len(m.type_three_errors)
report, error_num = gr.reportTypeThreeError(m.type_three_errors, explain_details=True)
if report is False:
results.at[file, GAMESPP_ERROR] = 3
# print("Model %s has an unexplainable Type III Error" % file)
# print("As the lower matrix has a condition number %f" % condition_number)
# print("Decide if the matrix is invertible")
except:
results.at[file, GAMES_ERROR] = -1
gamespp_end = time.time()
gamespp_time = gamespp_end - gamespp_start
print("\nAnalysis finished!")
print("GAMES++ time:", gamespp_time)
print("number of detected errors: ", len(results[results[GAMESPP_ERROR]==1]))
print("number of extended GAMES errors not in LP", len(results[(results[GAMESPP_ERROR]==1) & (results[LP_ERROR]!=1)]))
len(results[results[GAMESPP_ERROR]==-1])
len(results[results[GAMESPP_ERROR]==3])
results[results[GAMES_ERROR]==-1]
150/158
# +
# Finally, model statistics
MODEL_ID = "model_id"
NUM_TOTAL_REACTIONS = "num_total_reactions"
NUM_BDRY_REACTIONS = "num_bdry_reactions"
NUM_UNIUNI_REACTIONS = "num_uniuni_reactions"
NUM_UMMU_REACTIONS = "num_ummu_reactions"
NUM_MULTIMULTI_REACTIONS = "num_multimulti_reactions"
# exceptions for model
EXCEPTIONS = ["BIOMD0000000094.xml",
"BIOMD0000000596.xml",
"BIOMD0000000786.xml",
"BIOMD0000000794.xml",
"BIOMD0000000830.xml"]
# Checking all models to calculate the number of reactions per type
count = 0
simple = SimpleSBML()
res_list = []
false_errors = set()
for file in files:
if file in EXCEPTIONS:
continue
simple.initialize(os.path.join(data_dir, file))
num_uniuni = 0
num_ummu = 0
num_multimulti = 0
num_bdry = 0
count += 1
if (count%100)==0:
print("we are analyzing Model number:", count)
simple.initialize(os.path.join(data_dir, file))
num_dic = {cn.REACTION_1_1: 0,
cn.REACTION_1_n: 0,
cn.REACTION_n_1: 0,
cn.REACTION_n_n: 0,
cn.REACTION_BOUNDARY: 0
}
for reaction in simple.reactions:
for category in num_dic.keys():
if reaction.category == category:
num_dic[category] += 1
res_dic = dict({MODEL_ID: file[:-8],
NUM_TOTAL_REACTIONS: len(simple.reactions),
NUM_BDRY_REACTIONS: num_dic[cn.REACTION_BOUNDARY],
NUM_UNIUNI_REACTIONS: num_dic[cn.REACTION_1_1],
NUM_UMMU_REACTIONS: num_dic[cn.REACTION_1_n] + num_dic[cn.REACTION_n_1],
NUM_MULTIMULTI_REACTIONS: num_dic[cn.REACTION_n_n]})
sum = res_dic[NUM_BDRY_REACTIONS] + res_dic[NUM_UNIUNI_REACTIONS] + res_dic[NUM_UMMU_REACTIONS] + res_dic[NUM_MULTIMULTI_REACTIONS]
if sum != res_dic[NUM_TOTAL_REACTIONS]:
print("Error! The sum doens't match with model %s" % file)
res_list.append(res_dic)
# summarize the results
stats_df = pd.DataFrame(res_list)
reaction_num_cat = [NUM_UNIUNI_REACTIONS,
NUM_UMMU_REACTIONS,
NUM_MULTIMULTI_REACTIONS
]
reaction_prop = [(stats_df[x]/stats_df[NUM_TOTAL_REACTIONS]).mean()*100
for x in
reaction_num_cat]
ext_biomodels_reaction_prop = pd.Series(reaction_prop, index=reaction_num_cat)
ext_biomodels_reaction_prop
# -
EXCEPTIONS
| notebooks/analyze_ext_biomodels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="BJ8psIo-fhhB"
# ### **ABSTRACT**
#
# Stroke is a medical disorder in which the blood arteries in the brain are ruptured, causing damage to the brain. When the supply of blood and other nutrients to the brain is interrupted, symptoms might develop. According to the World Health Organization (WHO), stroke is the greatest cause of death and disability globally. Early recognition of the various warning signs of a stroke can help reduce the severity of the stroke.
#
# Stroke occurs when the blood flow to various areas of the brain is disrupted or diminished, resulting in the cells in those areas of the brain not receiving the nutrients and oxygen they require and dying. A stroke is a medical emergency that requires urgent medical attention. Early detection and appropriate management are required to prevent further damage to the affected area of the brain and other complications in other parts of the body.
#
# The World Health Organization (WHO) estimates that 15 million people worldwide suffer from strokes each year, with one person dying every 4-5 minutes in the affected population. Stroke is the 16 leading cause of mortality in the United States according to the Centers for Disease Control and Prevention (CDC) [1]. Stroke is a noncommunicable disease that kills approximately 11% of the population.
#
# In this project, I am working on stroke dataset. Here AutoMLH20 and Logistic Regression library is used for analysis.
#
# Source- https://pubmed.ncbi.nlm.nih.gov/34868531/
#
#
#
# + [markdown] id="U1Tb2yiGgyxO"
# ### **OBJECTIVE**
#
#
# Determine what makes a person at risk for stroke.
#
# Clean data and create a machine learning model to perform early detection of stroke for patients.
# + colab={"base_uri": "https://localhost:8080/"} id="463IzFiyXvhD" outputId="38751f23-fde3-4c1d-b14c-9d416b01fd50"
# !pip install requests
# !pip install tabulate
# !pip install "colorama>=0.3.8"
# !pip install future
# !pip install seaborn
# !pip install statsmodels
# + id="zJ5u9PGkZdJv"
# Import libraries
# Use pip install or conda install if missing a library
import random, os, sys
import warnings
from datetime import datetime
import pandas as pd
import logging
import csv
import optparse
import time
import json
from distutils.util import strtobool
import psutil
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
warnings.filterwarnings('ignore')
# + id="pK2R-4KQbOH8"
#For Pandas
df_pd=pd.read_csv('https://github.com/trivedi-mi/DataScience/raw/master/healthcare-dataset-stroke-data.csv')
df_pd.head()
df_pd_org=df_pd.copy(deep=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="kelmrIFP3spK" outputId="eaab6a30-686b-4ccd-e937-0671fcd58b5c"
#First five rows
df_pd.head()
# + colab={"base_uri": "https://localhost:8080/"} id="6JtVGCnl4nTa" outputId="231be014-bf6e-44b2-f2cd-8fe37d4029c0"
#Get Stats of all the columns
df_pd.info()
# + [markdown] id="ECmHTQnI5yib"
# There are categorical variables as well as float numerical values in the dataset.
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="mfbO2Xex5O-h" outputId="663b2e45-4d03-4d5f-afd6-5d69fb29de06"
#Description of all the columns
df_pd.describe()
# + [markdown] id="nuoG8xrJfAOS"
# From the table above, we can see there are some missing vales for BMI
# The average age is 43.
# The average bmi is 28
# The minimum age is questionable.
# Average glucose level is 106
# + colab={"base_uri": "https://localhost:8080/"} id="l_gO-SLe963H" outputId="a4a9cbb9-ba2f-437f-c69f-5a9260e6a290"
df_pd.shape
# + [markdown] id="dqgnHlgB99PK"
# We can see here the shape of the dataset is (5110,12)
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="gRQ7xAGp-MDY" outputId="97357d5d-714e-4770-b909-4130d5dd4f3b"
sns.countplot(x='stroke',data=df_pd_org)
# + [markdown] id="N9D5ZSfH-d6y"
# We can see that there is an imbalance for the number of stroke predictions. There are much more 0's than 1's
# + colab={"base_uri": "https://localhost:8080/", "height": 403} id="OsVhs_CwmNzh" outputId="1762e4c8-3ed3-4337-98dc-928e54b8979e"
#Relation between age and stroke
sns.catplot('stroke','age',data=df_pd_org,kind='violin')
plt.title('Relationship between Age and Stroke')
# + [markdown] id="MO3gmEQU590A"
# From the diagram above, we can see age is an important factor in predicting stroke. With the increase in age, the changes of stroke increases
# + colab={"base_uri": "https://localhost:8080/", "height": 319} id="qrxwqVGUmgSM" outputId="4e9e8e45-09ea-46f5-abe4-c94983788bbb"
df_pd_org[['Residence_type','bmi']].groupby('Residence_type').mean('bmi').plot(kind='bar') # Relation residence type and BMI
# + [markdown] id="oFKghzGo6Pnh"
#
# BMI is not varying for Rural and Urban
# + colab={"base_uri": "https://localhost:8080/", "height": 375} id="JbVGA1C7mmHz" outputId="f6b96de7-dec8-4108-d7fe-db5843eca259"
df_pd_org[['work_type','avg_glucose_level']].groupby('work_type').mean('avg_glucose_level').plot(kind='bar')
plt.ylabel('average glucose level')
plt.title('Relationship between glucose level and work type')
# + [markdown] id="df-nCNQoggCB"
# No significant result found
#
# + colab={"base_uri": "https://localhost:8080/", "height": 293} id="zP64dDz5mv6l" outputId="e34cd0cd-c3ed-4ce2-fe95-c53f2d9fa7ef"
# Relation for stroke with bmi and average
df_pd_org[['stroke','avg_glucose_level','bmi']].groupby('stroke').mean(['bmi','avg_glucose_level']).plot(kind='bar')
# + [markdown] id="O8kh7rl86lDx"
# Average glucose level is more with heart patients. No significant variation found for BMI.
# + colab={"base_uri": "https://localhost:8080/", "height": 337} id="5Kp8iJ9Uh59G" outputId="e2dad3a2-ff89-48b1-f1ac-f6a419774e91"
palettes = ['#9B856A', '#475962', '#598392', '#124559', '#540B0E']
ax = sns.barplot(data=df_pd_org, x='smoking_status', y='stroke',
palette=palettes, edgecolor=palettes, ci=None)
ax.set_title('Stroke and Smoking behaviour', y=1.1, weight='bold', fontsize=14)
# + [markdown] id="TT3yOuWkiU4t"
# One who smokes or has formerly smoked may have a higher risk of stroke
# + colab={"base_uri": "https://localhost:8080/", "height": 313} id="URaXK_3UB4UP" outputId="5c44c97c-4c96-4451-f5d0-39c3d7f56d16"
sns.countplot(x='stroke',data=df_pd_org,hue='gender')
plt.title('Gender wise stroke count',fontweight='bold')
# + [markdown] id="6ctBiQ-CDu1j"
# We can see that males are at a higher risk than females
# + colab={"base_uri": "https://localhost:8080/", "height": 269} id="pK8rQRyN8ImZ" outputId="0ce4dddf-6a47-42ec-993d-8f91b5c222ef"
#Find the correlation
corr_matrix = df_pd_org.corr()
corr_matrix
# + [markdown] id="sdqImjqN_0Jf"
# It can be seen that age plays an important factor in stroke with a correlation factor of 0.25. Average glucose level, heart disease and hypertension also play an important role.
# + colab={"base_uri": "https://localhost:8080/"} id="h-iu4MUa81EC" outputId="541a48e0-9523-4778-c52d-7bab407b6cb9"
# Find how much each attribue correlates to Stroke
corr_matrix["stroke"].sort_values(ascending=False)
# + [markdown] id="gmZWCYL29dwF"
# Age, heart_disease. glucose level and hypertension may play a significant role in stroke prediction
# + id="p4LD1cqXdDm_"
df_pd.dropna(inplace=True) # vif can't be calculated with nan values
df_pd = df_pd._get_numeric_data() # drop non-numeric cols
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="xe4QK5iMdqJm" outputId="356f56de-6bf6-4bc2-9046-23b489e3d622"
from statsmodels.stats.outliers_influence import variance_inflation_factor
#Indicate which variables to compute VIF
Multic = df_pd[['age', 'hypertension', 'heart_disease', 'avg_glucose_level', 'bmi']]
#Compute VIF
vif = pd.DataFrame()
vif["variables"] = Multic.columns
vif["VIF"] = [variance_inflation_factor(Multic.values, i) for i in range(Multic.shape[1])]
vif
# + [markdown] id="tie-Fj_D-k4a"
# They may be some collinearity as age, avg glucose level and BMI as their VIF is high
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="Ief2U1gljGL0" outputId="fda48fdd-a668-4455-b4bc-5dd58c063e84"
df_pd_org=df_pd_org.drop('id',axis=1,inplace=False)
df_pd_org.corr(method='pearson')
# + [markdown] id="OZ6vjWF2bu8U"
# Distributions for the numerical variables
# + colab={"base_uri": "https://localhost:8080/", "height": 441} id="SnsUI1S0a7nN" outputId="fce24a50-0208-47cc-a3cb-3d233a796e26"
column_list = ['age', 'avg_glucose_level', 'bmi']
fig, ax = plt.subplots(1, 3, figsize=(12,6))
for i, col in enumerate(column_list):
sns.histplot(data=df_pd_org, x=col, hue='stroke', multiple='stack', ax=ax[i])
plt.tight_layout()
plt.show()
# + [markdown] id="xSSy-Bsdjqjx"
# Average_Glucose_Level and BMI seems to have a right skewed form
# + colab={"base_uri": "https://localhost:8080/", "height": 441} id="xhr-j26w4D9X" outputId="ff63c711-db39-410c-86e1-d3d98d4210db"
column_list = ['age', 'avg_glucose_level', 'bmi']
fig, ax = plt.subplots(1, 3, figsize=(12,6))
for i, col in enumerate(column_list):
sns.boxplot(data=df_pd_org, x=col, ax=ax[i])
plt.tight_layout()
plt.show()
# + [markdown] id="ZeA36zXt_iBa"
# There may be some outliers for avg_glucose_level and bmi
# + colab={"base_uri": "https://localhost:8080/", "height": 367} id="YLMWChATj-Ze" outputId="0ecea405-05d4-47ad-c072-b6c0ff48ff21"
#from IPython.core.pylabtools import figsize
#f,ax=plt.subplots(figsize=(6,6))
#sns.heatmap(df_pd_org.corr(),center=0, linewidths=0.8,cmap='Greens',annot=True)
from matplotlib import colors
cmap = colors.ListedColormap(['#9B856A', '#124559', '#475962', '#598392'])
sns.heatmap(df_pd_org.corr(), annot=True, fmt='.2f', cmap=cmap)
# + [markdown] id="Nil78X9VsHTE"
# ID has no role to predict stroke. Furthermore, correlation between age and stroke is high as well as age and BMI
#
# + colab={"base_uri": "https://localhost:8080/", "height": 287} id="0aUVTom9CtEt" outputId="836bc493-047f-4156-a843-3c99222f166f"
data_onehot=pd.get_dummies(df_pd_org, prefix=None, dummy_na=False, columns=['gender','ever_married','work_type', 'Residence_type','smoking_status'], sparse=False, drop_first=True, dtype=None)
data_onehot.head()
# + [markdown] id="ZF90s1SfC0YW"
# One hot encoding for categorical data.
# + id="3VI3_pOQC64Z"
from sklearn.impute import KNNImputer
imputer = KNNImputer(n_neighbors=5)
data_onehot = pd.DataFrame(imputer.fit_transform(data_onehot),columns = data_onehot.columns)
# + [markdown] id="MAtj7_mEC9Eu"
# KNN imputation to fill values for BMI.
# + colab={"base_uri": "https://localhost:8080/", "height": 381} id="aKgaKDv9EG0v" outputId="7e6ad5e8-0cd3-4f96-9874-918d1da0cc09"
data_onehot.describe()
# + [markdown] id="rLR9O-WwEMDC"
# Missing values filled for BMI.
# + id="I9QyYhqNExpN"
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
y_stroke=data_onehot['stroke']
X_prep = data_onehot.drop(columns=['stroke'], inplace=False)
X_input = pd.DataFrame(data=scaler.fit_transform(X_prep), index=X_prep.index, columns=X_prep.columns)
# + [markdown] id="CGYIgN0ZFA6A"
# Applied Min Max scalar to the inputs to balance the data and reduce effects of the ranges of variables.
# + colab={"base_uri": "https://localhost:8080/", "height": 287} id="BqloQAaUE1gM" outputId="71a5f8aa-e149-416a-e687-89154b5c3d9e"
X_input.head()
# + colab={"base_uri": "https://localhost:8080/"} id="COdSrJwhGpFX" outputId="8967d8fa-69ec-4633-b55d-4bd9c660852d"
df_pd_org['stroke'].value_counts()
# + [markdown] id="uVDc_kkcGusl"
# We can see that the number of positive stroke prediction in the whole data is much less than the negative stroke prediction. We can assume our model shall be struggling with the training as it is not exposed to much equally sampled data i.e the data has much more 0's than 1's
# + id="gA0xcHDIGwKs"
x_infea=data_onehot.drop(columns='stroke')
y_outfea=data_onehot['stroke']
# + id="iHPGkf8CGzqm"
from sklearn.preprocessing import scale
from sklearn.model_selection import train_test_split
x_train_in,x_test_in,y_train_out,y_test_out=train_test_split(x_infea,y_outfea,test_size=0.2,random_state=421)
# + colab={"base_uri": "https://localhost:8080/", "height": 266} id="IzM4dAswG4rE" outputId="0160b2b5-972d-4220-b95f-ae3bb8d544c4"
plt.hist(y_train_out);
# + [markdown] id="Ylg7E-VeG8KS"
# No of ones are very less. Hence there is an imbalance in the output. We can use SMOTE to balance.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="2GLoUeFeHHQX" outputId="99792f23-1c2e-400e-ce17-304789b69e0a"
from imblearn.over_sampling import SMOTE
mod=SMOTE()
x_train1,y_train1=mod.fit_resample(x_train_in,y_train_out)
plt.hist(y_train1);
print(x_train1)
# + [markdown] id="-xTHPgxuHSbV"
# We have balanced number here and we can apply regression again.
# + colab={"base_uri": "https://localhost:8080/"} id="zm2COeWjHQwr" outputId="ddb10f01-9b00-4cc0-bbdb-2320f21b0b65"
#Applying Logistic Regression
#Apply l1 penalty
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
from sklearn.metrics import roc_auc_score
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
#lg=LogisticRegression(max_iter=500,penalty='l1',solver='liblinear',C=3.0)
lg=LogisticRegression()
lg.fit(x_train1,y_train1)
print(confusion_matrix(y_test_out,lg.predict(x_test_in)))
print(classification_report(y_test_out,lg.predict(x_test_in)))
from sklearn.metrics import f1_score
from sklearn.metrics import recall_score
print("F1 Score:")
print(f1_score(y_test_out, lg.predict(x_test_in), average='weighted'))
print("Recall")
(recall_score(y_test_out, lg.predict(x_test_in), average='macro'))
# + id="4_IGTx1Dnclt"
from sklearn.metrics import roc_curve
fpr,tpr,thresh=roc_curve(y_test_out,lg.predict_proba(x_test_in)[:,1])
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="8roOvJF-oOPk" outputId="262bfae5-ccb0-4c68-9336-e3a7e7979e9c"
plt.plot(fpr,tpr,color='blue',label='logistic')
# + [markdown] id="2zi8w_ET4Ff8"
# # The recall is good with l1 penalty and the accuracy is good
# + [markdown] id="LUMVRvB84wpB"
# ### Lets train and test the models by dropping the unimportant variables like work type and residence
# + id="eSyq3xweQHgH" colab={"base_uri": "https://localhost:8080/", "height": 287} outputId="38ddb0a6-aaed-4dbb-8982-8f469a09a52e"
data_onehot.head()
# + id="07DJqwNJTuwW"
data_dropped=data_onehot.copy(deep=True)
# + id="qWovfopAT1NZ"
data_dropped=data_dropped.drop(['gender_Male','bmi', 'gender_Other', 'ever_married_Yes','work_type_Never_worked','work_type_Private','work_type_Self-employed', 'work_type_children','Residence_type_Urban' ], axis=1)
# + id="pwf0GffdUWgK" colab={"base_uri": "https://localhost:8080/", "height": 250} outputId="cd5972a0-078d-4b62-ede9-f58cbbe27274"
data_dropped.head()
# + id="F11iAphMUt1P" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="35057001-ff92-4c32-9cd7-83bbcff4f942"
x_infea3=data_dropped.drop(columns='stroke')
y_outfea3=data_dropped['stroke']
x_train_in3,x_test_in3,y_train_out3,y_test_out3=train_test_split(x_infea3,y_outfea3,test_size=0.2,random_state=431)
mod=SMOTE()
x_train4,y_train4=mod.fit_resample(x_train_in3,y_train_out3)
plt.hist(y_train4);
# + id="_DgD5ubNV51S" colab={"base_uri": "https://localhost:8080/"} outputId="10e83924-4319-4414-fe1a-6ade3193e582"
y_train4.value_counts()
# + [markdown] id="R9qfKPE6ok66"
# ## Now lets test on scaled data
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="I2_5hEE3XDIT" outputId="26a2174a-afee-4ac0-8570-72291a6cc9d1"
x_infea3=data_dropped.drop(columns='stroke')
y_outfea3=data_dropped['stroke']
scaler = MinMaxScaler()
scaler.fit(x_infea3)
X_scaled = scaler.transform(x_infea3)
X = pd.DataFrame(X_scaled, columns=x_infea3.columns)
x_train_in3,x_test_in3,y_train_out3,y_test_out3=train_test_split(X,y_outfea3,test_size=0.2,random_state=431)
mod=SMOTE()
x_train4,y_train4=mod.fit_resample(x_train_in3,y_train_out3)
plt.hist(y_train4);
# + colab={"base_uri": "https://localhost:8080/"} id="aiJqrVmgcAeN" outputId="16b3a05f-f3a9-4d5d-8a3a-bc17f7b610fc"
best_clf2=LogisticRegression()
best_clf2.fit(x_train4, y_train4)
print(best_clf2.coef_, best_clf2.intercept_)
# + [markdown] id="tWrZNCZeadwa"
# Again we can see that age has a high importance here in predicting stroke
# + colab={"base_uri": "https://localhost:8080/"} id="IK5Shr5McL0T" outputId="f62a134f-587d-4427-d236-106215bb8978"
print (f'Accuracy - : {best_clf2.score(x_train4,y_train4):.3f}')
# + colab={"base_uri": "https://localhost:8080/"} id="CXrYXrF1cQB-" outputId="7fc499f1-eaf4-4a8b-cfad-7f6f40d5a37e"
y_prediction3=best_clf2.predict(x_test_in3)
cm2 = confusion_matrix(y_test_out3,y_prediction3)
print(cm2)
cr2=classification_report(y_test_out3,y_prediction3)
print(cr2)
from sklearn.metrics import f1_score
from sklearn.metrics import recall_score
print("F1 Score:")
print(f1_score(y_test_out3, y_prediction3, average='weighted'))
from sklearn.metrics import recall_score
print("Recall")
(recall_score(y_test_out3, y_prediction3, average='weighted'))
# + [markdown] id="p_mUR2zxLMgW"
# Dropping unimportant variables does not make a great difference in F1 score but yes it has significantlty increased
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="_B4pXw56-kL1" outputId="bc65f115-a2e5-4c5d-d892-dbc1b0d1deb0"
from sklearn.metrics import roc_curve
fpr,tpr,thresh=roc_curve(y_test_out3,best_clf2.predict_proba(x_test_in3)[:,1])
plt.plot(fpr,tpr,color='blue',label='logistic')
# + [markdown] id="ybXKRWHx-Ipy"
# Recall on Stroke prediction is good. We have more number of predicting positive stroke and that is great here
#
# + [markdown] id="mnVUegIf5LU1"
# ### Recall and F1 score is good. Seems to be a good model
# + colab={"base_uri": "https://localhost:8080/", "height": 287} id="B14X8m93rlyL" outputId="b48bb745-1d9b-4906-8249-e56e53aa71e1"
data_feature=pd.get_dummies(df_pd_org, prefix=None, dummy_na=False, columns=['gender','ever_married','work_type', 'Residence_type','smoking_status'], sparse=False, drop_first=True, dtype=None)
data_feature.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 381} id="Oeg126ulz95z" outputId="d4a5f2af-114c-4a55-fba6-481ed656ed12"
data_feature.describe()
# + [markdown] id="f5UCDcYb0VRz"
# There are missing values for BMI
# + [markdown] id="jyvl7WLh0eU_"
# To fix the missing values in the BMI columns we use a imputation technics based on the KNN.
# + id="AyFmOqhkrtuY"
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
df_feature = pd.DataFrame(scaler.fit_transform(data_feature), columns = data_feature.columns)
df_feature.head()
df_feature_impute=df_feature.copy(deep=True)
# + id="rSmVQ5gP0lHM"
from sklearn.impute import KNNImputer
imputer = KNNImputer(n_neighbors=5)
df_feature = pd.DataFrame(imputer.fit_transform(df_feature),columns = df_feature.columns)
# + colab={"base_uri": "https://localhost:8080/"} id="X4umhm-121wB" outputId="1b0fe5a5-0d9e-469c-883d-863203779473"
df_feature.isna().sum()
# + [markdown] id="gIvxPqEg-2QS"
# Now BMI has no missing values
# + colab={"base_uri": "https://localhost:8080/"} id="WQ-RQheA7-5B" outputId="e9114bc0-f48e-4a8d-a3d3-3869d4829ab1"
from imblearn.over_sampling import SMOTE
X , y = df_feature.drop(columns='stroke'),df_feature["stroke"]
x_train,x_test,y_train,y_test = train_test_split(X,y,test_size=0.2,random_state=23)
sm = SMOTE()
X_res, y_res = sm.fit_resample(x_train,y_train)
print("Before OverSampling, counts of label '1': {}".format(sum(y==1)))
print("Before OverSampling, counts of label '0': {} \n".format(sum(y==0)))
print('After OverSampling, the shape of train_X: {}'.format(X_res.shape))
print('After OverSampling, the shape of train_y: {} \n'.format(y_res.shape))
print("After OverSampling, counts of label '1': {}".format(sum(y_res==1)))
print("After OverSampling, counts of label '0': {}".format(sum(y_res==0)))
# + [markdown] id="HkLhkpKhrcE4"
# Sampling has balanced our data.
# + colab={"base_uri": "https://localhost:8080/", "height": 505} id="THmtztikA1Nx" outputId="a8a0fb06-7b4d-41ef-8add-a7df2488f3df"
X
# + colab={"base_uri": "https://localhost:8080/"} id="-xiQGFWSA5Pn" outputId="b8bda10c-96a3-4b9e-b03f-64d777fae37d"
y_res.value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="1D4HJKVHA9Bh" outputId="e5aa0a82-1232-42b6-9dd7-d9928eee8d1c"
from sklearn.metrics import classification_report
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
lg=LogisticRegression(max_iter=500,penalty='l1',solver='liblinear',C=3.0)
#lg=LogisticRegression()
lg.fit(X_res,y_res)
print(confusion_matrix(y_test,lg.predict(x_test)))
print(classification_report(y_test,lg.predict(x_test)))
from sklearn.metrics import f1_score
from sklearn.metrics import recall_score
print('Validation Acuuracy: ',accuracy_score(y_test,lg.predict(x_test)))
print('Training Accuracy: ',accuracy_score(y_res,lg.predict(X_res)))
print("F1 Score:")
print(f1_score(y_test, lg.predict(x_test), average='weighted'))
print("Recall")
(recall_score(y_test, lg.predict(x_test), average='macro'))
# + [markdown] id="2sL6bFQczaVj"
# No significant chabge in accuracy after filling BMI.
# + colab={"base_uri": "https://localhost:8080/", "height": 381} id="KZRkswSCFqqX" outputId="b5ec5260-58ec-43f4-de16-a6ff71b422ac"
df_feature.describe()
df_feature_copy=df_feature.copy(deep=True)
df_feature_copy.describe()
# + [markdown] id="RWqwyCderjP2"
# ## Removing *Outliers*
# + id="J-OVqCo2RluT"
cols = ['age','avg_glucose_level','bmi'] # one or more
Q1 = df_feature[cols].quantile(0.25)
Q3 = df_feature[cols].quantile(0.75)
IQR = Q3 - Q1
df_feature = df_feature[~((df_feature[cols] < (Q1 - 1.5 * IQR)) |(df_feature[cols] > (Q3 + 1.5 * IQR))).any(axis=1)]
# + [markdown] id="ONzywjErzhWo"
# Removed outliers
# + colab={"base_uri": "https://localhost:8080/", "height": 381} id="SBW6L4QBTEqV" outputId="fe258454-c809-4800-e7e2-b28129616708"
df_feature.describe()
# + [markdown] id="taZgRoE3M8hA"
# We can see that we have removed the outliers here.
# + id="vHpfcF0UTGe1"
from imblearn.over_sampling import SMOTE
X , y = df_feature.drop(columns='stroke'),df_feature["stroke"]
x_train,x_test,y_train,y_test = train_test_split(X,y,test_size=0.2,random_state=26)
sm = SMOTE()
X_res, y_res = sm.fit_resample(x_train,y_train)
# + [markdown] id="D-fcs1oNNAu9"
# Smote to balance the data.
# + colab={"base_uri": "https://localhost:8080/", "height": 505} id="7J5fHQoOUZAZ" outputId="dd874f7c-0c2f-45f4-837d-51c03af5a48d"
X
# + colab={"base_uri": "https://localhost:8080/"} id="7PxAHVCVUdXC" outputId="dde7d094-7031-43b7-e54b-d20411a36f39"
y_res.value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="MaHcU9QuUB7s" outputId="62020806-461c-48b4-a30f-6a0278b3726b"
from sklearn.metrics import classification_report
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
lg=LogisticRegression(max_iter=500,penalty='l1',solver='liblinear',C=3.0)
#lg=LogisticRegression()
lg.fit(X_res,y_res)
print(confusion_matrix(y_test,lg.predict(x_test)))
print(classification_report(y_test,lg.predict(x_test)))
from sklearn.metrics import f1_score
from sklearn.metrics import recall_score
print('Validation Acuuracy: ',accuracy_score(y_test,lg.predict(x_test)))
print('Training Accuracy: ',accuracy_score(y_res,lg.predict(X_res)))
print("F1 Score:")
print(f1_score(y_test, lg.predict(x_test), average='weighted'))
print("Recall")
(recall_score(y_test, lg.predict(x_test), average='macro'))
# + [markdown] id="52Yug6Nbr3gW"
# F1 score, recall and accuracy improved after removing outliers
# + colab={"base_uri": "https://localhost:8080/", "height": 381} id="8XyPJvvTbMFz" outputId="918bffd7-5b71-4a98-f0b4-98e8c6b7ad63"
df_feature_impute_standard=df_feature_copy.copy(deep=True)
df_feature_impute_standard.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="rD14_bIzn1j5" outputId="a0f2e987-3626-4c50-946e-efdf84c7069b"
df_feature_impute_standard['bmi'].describe()
# + [markdown] id="1YWxC9Y5srl_"
# ## Remove 1%, 5%, and 10% of your data randomly and impute the values back using at least 3 imputation methods
# + colab={"base_uri": "https://localhost:8080/", "height": 423} id="v_xMOKb_pVyK" outputId="2f73f430-3c8c-41c7-8db6-76ea8b45322a"
df_bmi = df_feature_impute_standard[['bmi']]
df_bmi['bmi_copy_1_percent'] = df_bmi[['bmi']]
df_bmi['bmi_copy_5_percent'] = df_bmi[['bmi']]
df_bmi['bmi_copy_10_percent'] = df_bmi[['bmi']]
df_bmi['sample_bmi'] = df_feature_impute_standard[['bmi']]
df_bmi['sample_bmi'] = df_bmi['sample_bmi']/1000
df_bmi
# + id="mn998Spzqg6J"
def get_percent_missing(dataframe):
percent_missing = dataframe.isnull().sum() * 100 / len(dataframe)
missing_value_df = pd.DataFrame({'column_name': dataframe.columns,
'percent_missing': percent_missing})
return missing_value_df
# + id="6itERCtRqyI3" colab={"base_uri": "https://localhost:8080/"} outputId="a2e381e5-d925-4f9a-ee13-3b25c0f4bc5b"
print(get_percent_missing(df_bmi))
# + id="1wscyqQhq5WT"
def create_missing(dataframe, percent, col):
dataframe.loc[dataframe.sample(frac = percent).index, col] = np.nan
# + id="c5k31q0jriXF"
create_missing(df_bmi, 0.01, 'bmi_copy_1_percent')
create_missing(df_bmi, 0.05, 'bmi_copy_5_percent')
create_missing(df_bmi, 0.1, 'bmi_copy_10_percent')
# + colab={"base_uri": "https://localhost:8080/"} id="f86mBPBJrt6Q" outputId="8e1f110f-9f6d-44e5-9c93-3873fb6ce871"
print(get_percent_missing(df_bmi))
# + id="jUu5O9qPr1Vd"
# Store Index of NaN values in each coloumns
bmi_1_idx = list(np.where(df_bmi['bmi_copy_1_percent'].isna())[0])
bmi_5_idx = list(np.where(df_bmi['bmi_copy_5_percent'].isna())[0])
bmi_10_idx = list(np.where(df_bmi['bmi_copy_10_percent'].isna())[0])
# + colab={"base_uri": "https://localhost:8080/"} id="f7c8lfcpwQ90" outputId="679a2dd6-f4bc-4af5-f30b-435447f0b926"
print(f"Length of bmi_1_idx is {len(bmi_1_idx)} and it contains {(len(bmi_1_idx)/len(df_bmi['bmi_copy_1_percent']))*100}% of total data in column | Total rows: {len(df_bmi['bmi_copy_1_percent'])}")
# + colab={"base_uri": "https://localhost:8080/"} id="D83wdkihwpdR" outputId="fe337cf4-91a5-4c99-9abf-0db51df0358c"
print(f"Length of bmi_5_idx is {len(bmi_5_idx)} and it contains {(len(bmi_5_idx)/len(df_bmi['bmi_copy_5_percent']))*100}% of total data in column | Total rows: {len(df_bmi['bmi_copy_5_percent'])}")
# + colab={"base_uri": "https://localhost:8080/"} id="8czqdWOaw4hR" outputId="cd02aa8b-ee03-424b-e945-f0560e889e8a"
print(f"Length of bmi_10_idx is {len(bmi_10_idx)} and it contains {(len(bmi_10_idx)/len(df_bmi['bmi_copy_10_percent']))*100}% of total data in column | Total rows: {len(df_bmi['bmi_copy_10_percent'])}")
# + [markdown] id="<KEY>"
# ## Imputation using KNN
# + id="9an3dbFhw_JQ"
df_bmi1 = df_bmi[['sample_bmi','bmi_copy_10_percent']]
imputer = KNNImputer(n_neighbors=5)
imputed_bmi_df = pd.DataFrame(imputer.fit_transform(df_bmi1), columns = df_bmi1.columns)
# + colab={"base_uri": "https://localhost:8080/", "height": 363} id="K5YZJ4scxaAX" outputId="c7f7ce72-f555-4e67-e6b3-13ffda586d48"
imputed_bmi_df.sample(10)
# + colab={"base_uri": "https://localhost:8080/"} id="w5DMXUzrxfYQ" outputId="21b6cd6f-d457-496f-88be-cdd296b7836c"
print(get_percent_missing(imputed_bmi_df))
# + id="0lMoSpnuyA6-"
imputed_bmi_df['original'] = df_bmi['bmi']
# + [markdown] id="4PACQ6Yv2ucG"
# 10 percent KNN
#
# + id="1cbTijXoyPDj"
imputed_bmi_df['diff'] = imputed_bmi_df['original'] - imputed_bmi_df['bmi_copy_10_percent']
# + colab={"base_uri": "https://localhost:8080/"} id="h5aSWXpvyW0S" outputId="ac6c4b7a-0cb5-4ab5-e093-af18db49e5b8"
for i in bmi_10_idx:
print(str(imputed_bmi_df['bmi_copy_10_percent'][i])+", "+str(imputed_bmi_df['original'][i])+", "+str(imputed_bmi_df['diff'][i]))
# + colab={"base_uri": "https://localhost:8080/"} id="ZeJjla5dPI7a" outputId="2998258e-fbe5-414e-b7ce-5230231774d9"
# create list of difference bwtween imputed and orginal value
bmi_diff_1 = []
bmi_diff_5 = []
bmi_diff_10 = []
count = 0
for i in bmi_10_idx:
diff10 = abs(imputed_bmi_df['bmi_copy_10_percent'][i] - df_bmi['bmi'][i])
bmi_diff_10.append(diff10)
print("Array of differences: ")
print(bmi_diff_10)
print("Length of Array: " + str(len(bmi_diff_10)))
print("Sum of Differences: ")
print(sum(abs(number) for number in bmi_diff_10))
# + [markdown] id="HmirYt4Tz3rY"
# KNN performed well
# + [markdown] id="r29WtuI32zrk"
# 5 percent KNN
# + colab={"base_uri": "https://localhost:8080/"} id="sACdkPpvynyG" outputId="52b0c30f-3ea9-4719-f969-2e61d7ac39e1"
df_bmi1 = df_bmi[['sample_bmi','bmi_copy_5_percent']]
imputer = KNNImputer(n_neighbors=5)
imputed_bmi_df = pd.DataFrame(imputer.fit_transform(df_bmi1), columns = df_bmi1.columns)
imputed_bmi_df.sample(10)
print(get_percent_missing(imputed_bmi_df))
imputed_bmi_df['original'] = df_bmi['bmi']
imputed_bmi_df['diff'] = imputed_bmi_df['original'] - imputed_bmi_df['bmi_copy_5_percent']
for i in bmi_5_idx:
print(str(imputed_bmi_df['bmi_copy_5_percent'][i])+", "+str(imputed_bmi_df['original'][i])+", "+str(imputed_bmi_df['diff'][i]))
# + colab={"base_uri": "https://localhost:8080/"} id="-GJyyk7qPxRF" outputId="a37ce8d8-3018-4db1-e155-636677035ca0"
# create list of difference bwtween imputed and orginal value
bmi_diff_1 = []
bmi_diff_5 = []
bmi_diff_10 = []
count = 0
for i in bmi_5_idx:
diff5 = abs(imputed_bmi_df['bmi_copy_5_percent'][i] - df_bmi['bmi'][i])
bmi_diff_5.append(diff5)
print(len(bmi_diff_5))
print("Sum : ")
print(sum(abs(number) for number in bmi_diff_5))
# + [markdown] id="zUJz2wMA241X"
# 1 percent KNN
# + colab={"base_uri": "https://localhost:8080/"} id="TI0V17JTzbrb" outputId="f1533b18-d590-4472-f2cd-4d6a3ffef903"
df_bmi1 = df_bmi[['sample_bmi','bmi_copy_1_percent']]
imputer = KNNImputer(n_neighbors=5)
imputed_bmi_df = pd.DataFrame(imputer.fit_transform(df_bmi1), columns = df_bmi1.columns)
imputed_bmi_df.sample(10)
print(get_percent_missing(imputed_bmi_df))
imputed_bmi_df['original'] = df_bmi['bmi']
imputed_bmi_df['diff'] = imputed_bmi_df['original'] - imputed_bmi_df['bmi_copy_1_percent']
for i in bmi_1_idx:
print(str(imputed_bmi_df['bmi_copy_1_percent'][i])+", "+str(imputed_bmi_df['original'][i])+", "+str(imputed_bmi_df['diff'][i]))
# + colab={"base_uri": "https://localhost:8080/"} id="oUpa-A_U6xor" outputId="96008baf-db80-4117-ec56-aac58bb643e7"
# create list of difference bwtween imputed and orginal value
bmi_diff_1 = []
bmi_diff_5 = []
bmi_diff_10 = []
count = 0
for i in bmi_1_idx:
count +=1
diff1 = abs(imputed_bmi_df['bmi_copy_1_percent'][i] - df_bmi['bmi'][i])
bmi_diff_1.append(diff1)
print(len(bmi_diff_1))
print("Sum: ")
print(sum(abs(number) for number in bmi_diff_1))
# + [markdown] id="RDhx8hIFXLCv"
# KNN nerarest neighbours performed pretty well in recovering the variables where the differences are minimal between original and imputed values
# + [markdown] id="JEGoQmHX4mPX"
# ## Impute BMI recovery check using mean
#
# + colab={"base_uri": "https://localhost:8080/"} id="MYO66gZ-1tzO" outputId="ab5738c5-f25d-4972-9753-b013deeceb1e"
df_feature_impute_standard['bmi'].describe()
df_bmi = df_feature_impute_standard[['bmi']]
df_bmi['bmi_copy_1_percent'] = df_bmi[['bmi']]
df_bmi['bmi_copy_5_percent'] = df_bmi[['bmi']]
df_bmi['bmi_copy_10_percent'] = df_bmi[['bmi']]
df_bmi['sample_bmi'] = df_feature_impute_standard[['bmi']]
df_bmi['sample_bmi'] = df_bmi['sample_bmi']/1000
df_bmi
def get_percent_missing(dataframe):
percent_missing = dataframe.isnull().sum() * 100 / len(dataframe)
missing_value_df = pd.DataFrame({'column_name': dataframe.columns,
'percent_missing': percent_missing})
return missing_value_df
print(get_percent_missing(df_bmi))
def create_missing(dataframe, percent, col):
dataframe.loc[dataframe.sample(frac = percent).index, col] = np.nan
create_missing(df_bmi, 0.01, 'bmi_copy_1_percent')
create_missing(df_bmi, 0.05, 'bmi_copy_5_percent')
create_missing(df_bmi, 0.1, 'bmi_copy_10_percent')
print(get_percent_missing(df_bmi))
# Store Index of NaN values in each coloumns
bmi_1_idx = list(np.where(df_bmi['bmi_copy_1_percent'].isna())[0])
bmi_5_idx = list(np.where(df_bmi['bmi_copy_5_percent'].isna())[0])
bmi_10_idx = list(np.where(df_bmi['bmi_copy_10_percent'].isna())[0])
print(f"Length of bmi_1_idx is {len(bmi_1_idx)} and it contains {(len(bmi_1_idx)/len(df_bmi['bmi_copy_1_percent']))*100}% of total data in column | Total rows: {len(df_bmi['bmi_copy_1_percent'])}")
print(f"Length of bmi_5_idx is {len(bmi_5_idx)} and it contains {(len(bmi_5_idx)/len(df_bmi['bmi_copy_5_percent']))*100}% of total data in column | Total rows: {len(df_bmi['bmi_copy_5_percent'])}")
print(f"Length of bmi_10_idx is {len(bmi_10_idx)} and it contains {(len(bmi_10_idx)/len(df_bmi['bmi_copy_10_percent']))*100}% of total data in column | Total rows: {len(df_bmi['bmi_copy_10_percent'])}")
# + [markdown] id="7IrvsmCW28VP"
# 5 percent Mean
# + colab={"base_uri": "https://localhost:8080/"} id="XoKomF7A4tq4" outputId="b8dfc495-47c7-4249-ad85-e20455a783ef"
df_bmi1 = df_bmi[['sample_bmi','bmi_copy_5_percent']]
imputed_bmi_df = df_bmi1.fillna(df_bmi1.mean())
imputed_bmi_df.sample(10)
print(get_percent_missing(imputed_bmi_df))
imputed_bmi_df['original'] = df_bmi['bmi']
imputed_bmi_df['diff'] = imputed_bmi_df['original'] - imputed_bmi_df['bmi_copy_5_percent']
for i in bmi_5_idx:
print(str(imputed_bmi_df['bmi_copy_5_percent'][i])+", "+str(imputed_bmi_df['original'][i])+", "+str(imputed_bmi_df['diff'][i]))
# + colab={"base_uri": "https://localhost:8080/"} id="V9_BQhnLQWx0" outputId="c14c6c4f-f397-40bc-ff08-2d1f4de0ced4"
# create list of difference bwtween imputed and orginal value
bmi_diff_1 = []
bmi_diff_5 = []
bmi_diff_10 = []
count = 0
for i in bmi_5_idx:
diff5 = abs(imputed_bmi_df['bmi_copy_5_percent'][i] - df_bmi['bmi'][i])
bmi_diff_5.append(diff5)
print(len(bmi_diff_5))
print("Sum: ")
print(sum(abs(number) for number in bmi_diff_5))
# + [markdown] id="y610lsujt20g"
# Mean difference can be high because of presence of Outliers in the data
#
# + [markdown] id="xOw9Xvzj3BgP"
# 10 percent Mean
#
# + colab={"base_uri": "https://localhost:8080/"} id="4rk0LIrd42-n" outputId="0cb62c92-14f7-4b95-ff1d-452855f09876"
df_bmi1 = df_bmi[['sample_bmi','bmi_copy_10_percent']]
imputed_bmi_df = df_bmi1.fillna(df_bmi1.mean())
imputed_bmi_df.sample(10)
print(get_percent_missing(imputed_bmi_df))
imputed_bmi_df['original'] = df_bmi['bmi']
imputed_bmi_df['diff'] = imputed_bmi_df['original'] - imputed_bmi_df['bmi_copy_10_percent']
for i in bmi_10_idx:
print(str(imputed_bmi_df['bmi_copy_10_percent'][i])+", "+str(imputed_bmi_df['original'][i])+", "+str(imputed_bmi_df['diff'][i]))
# + colab={"base_uri": "https://localhost:8080/"} id="AiHbNU6zQn0N" outputId="b8f50a01-c97b-4fb9-db3d-2ecf948a7b90"
# create list of difference bwtween imputed and orginal value
bmi_diff_1 = []
bmi_diff_5 = []
bmi_diff_10 = []
count = 0
for i in bmi_10_idx:
diff10 = abs(imputed_bmi_df['bmi_copy_10_percent'][i] - df_bmi['bmi'][i])
bmi_diff_10.append(diff10)
print(len(bmi_diff_10))
print("Sum: ")
print(sum(abs(number) for number in bmi_diff_10))
# + [markdown] id="7QFW6r_s3Ga0"
# 1 percent mean
#
# + colab={"base_uri": "https://localhost:8080/"} id="LZKkT4vm5Pzb" outputId="5a4ee8d9-46dc-4057-dc3e-9428b0a8f450"
df_bmi1 = df_bmi[['sample_bmi','bmi_copy_1_percent']]
imputed_bmi_df = df_bmi1.fillna(df_bmi1.mean())
imputed_bmi_df.sample(10)
print(get_percent_missing(imputed_bmi_df))
imputed_bmi_df['original'] = df_bmi['bmi']
imputed_bmi_df['diff'] = imputed_bmi_df['original'] - imputed_bmi_df['bmi_copy_1_percent']
for i in bmi_1_idx:
print(str(imputed_bmi_df['bmi_copy_1_percent'][i])+", "+str(imputed_bmi_df['original'][i])+", "+str(imputed_bmi_df['diff'][i]))
# + colab={"base_uri": "https://localhost:8080/"} id="7l5Dvyo0QwK5" outputId="445ca8b5-910e-47db-807d-20cd95f33a0d"
# create list of difference bwtween imputed and orginal value
bmi_diff_1 = []
bmi_diff_5 = []
bmi_diff_10 = []
count = 0
for i in bmi_1_idx:
count +=1
diff1 = abs(imputed_bmi_df['bmi_copy_1_percent'][i] - df_bmi['bmi'][i])
bmi_diff_1.append(diff1)
print(len(bmi_diff_1))
print("Sum: ")
print(sum(abs(number) for number in bmi_diff_1))
# + [markdown] id="sxM977pZ5gVe"
# ## Impute Check using median
# + colab={"base_uri": "https://localhost:8080/"} id="vH0t9bPC5ayb" outputId="31294f72-6e45-4e3c-f3ec-e79b4ad7d1dc"
df_feature_impute_standard['bmi'].describe()
df_bmi = df_feature_impute_standard[['bmi']]
df_bmi['bmi_copy_1_percent'] = df_bmi[['bmi']]
df_bmi['bmi_copy_5_percent'] = df_bmi[['bmi']]
df_bmi['bmi_copy_10_percent'] = df_bmi[['bmi']]
df_bmi['sample_bmi'] = df_feature_impute_standard[['bmi']]
df_bmi['sample_bmi'] = df_bmi['sample_bmi']/1000
df_bmi
def get_percent_missing(dataframe):
percent_missing = dataframe.isnull().sum() * 100 / len(dataframe)
missing_value_df = pd.DataFrame({'column_name': dataframe.columns,
'percent_missing': percent_missing})
return missing_value_df
print(get_percent_missing(df_bmi))
def create_missing(dataframe, percent, col):
dataframe.loc[dataframe.sample(frac = percent).index, col] = np.nan
create_missing(df_bmi, 0.01, 'bmi_copy_1_percent')
create_missing(df_bmi, 0.05, 'bmi_copy_5_percent')
create_missing(df_bmi, 0.1, 'bmi_copy_10_percent')
print(get_percent_missing(df_bmi))
# Store Index of NaN values in each coloumns
bmi_1_idx = list(np.where(df_bmi['bmi_copy_1_percent'].isna())[0])
bmi_5_idx = list(np.where(df_bmi['bmi_copy_5_percent'].isna())[0])
bmi_10_idx = list(np.where(df_bmi['bmi_copy_10_percent'].isna())[0])
print(f"Length of bmi_1_idx is {len(bmi_1_idx)} and it contains {(len(bmi_1_idx)/len(df_bmi['bmi_copy_1_percent']))*100}% of total data in column | Total rows: {len(df_bmi['bmi_copy_1_percent'])}")
print(f"Length of bmi_5_idx is {len(bmi_5_idx)} and it contains {(len(bmi_5_idx)/len(df_bmi['bmi_copy_5_percent']))*100}% of total data in column | Total rows: {len(df_bmi['bmi_copy_5_percent'])}")
print(f"Length of bmi_10_idx is {len(bmi_10_idx)} and it contains {(len(bmi_10_idx)/len(df_bmi['bmi_copy_10_percent']))*100}% of total data in column | Total rows: {len(df_bmi['bmi_copy_10_percent'])}")
# + [markdown] id="fMZF4UW_3KT2"
# 1 percent Median
# + colab={"base_uri": "https://localhost:8080/"} id="1yhPW7jV5kLS" outputId="253f14c1-e9a1-4553-ece8-3d069a8fe569"
df_bmi1 = df_bmi[['sample_bmi','bmi_copy_1_percent']]
imputed_bmi_df = df_bmi1.fillna(df_bmi1.median())
imputed_bmi_df.sample(10)
print(get_percent_missing(imputed_bmi_df))
imputed_bmi_df['original'] = df_bmi['bmi']
imputed_bmi_df['diff'] = imputed_bmi_df['original'] - imputed_bmi_df['bmi_copy_1_percent']
for i in bmi_1_idx:
print(str(imputed_bmi_df['bmi_copy_1_percent'][i])+", "+str(imputed_bmi_df['original'][i])+", "+str(imputed_bmi_df['diff'][i]))
# + colab={"base_uri": "https://localhost:8080/"} id="JaLPzMwlRBmB" outputId="eea06bc1-5788-4a51-924d-652269a3927d"
# create list of difference bwtween imputed and orginal value
bmi_diff_1 = []
bmi_diff_5 = []
bmi_diff_10 = []
count = 0
for i in bmi_1_idx:
count +=1
diff1 = abs(imputed_bmi_df['bmi_copy_1_percent'][i] - df_bmi['bmi'][i])
bmi_diff_1.append(diff1)
print(len(bmi_diff_1))
print("Sum: ")
print(sum(abs(number) for number in bmi_diff_1))
# + [markdown] id="N14D6ATyuXT8"
# Median performs better than mean when there are outliers
# + [markdown] id="09LevvY13PUe"
# 5 percent Median
#
# + colab={"base_uri": "https://localhost:8080/"} id="1jtXEt-O6CdP" outputId="abb0adcd-e61f-480e-b13b-a159e0ab80b8"
df_bmi1 = df_bmi[['sample_bmi','bmi_copy_5_percent']]
imputed_bmi_df = df_bmi1.fillna(df_bmi1.median())
imputed_bmi_df.sample(10)
print(get_percent_missing(imputed_bmi_df))
imputed_bmi_df['original'] = df_bmi['bmi']
imputed_bmi_df['diff'] = imputed_bmi_df['original'] - imputed_bmi_df['bmi_copy_5_percent']
for i in bmi_5_idx:
print(str(imputed_bmi_df['bmi_copy_5_percent'][i])+", "+str(imputed_bmi_df['original'][i])+", "+str(imputed_bmi_df['diff'][i]))
# + colab={"base_uri": "https://localhost:8080/"} id="PM1QlFUjRbZB" outputId="e52bb524-15c8-4ab5-eb4f-880b219c6b1f"
# create list of difference bwtween imputed and orginal value
bmi_diff_1 = []
bmi_diff_5 = []
bmi_diff_10 = []
count = 0
for i in bmi_5_idx:
diff5 = abs(imputed_bmi_df['bmi_copy_5_percent'][i] - df_bmi['bmi'][i])
bmi_diff_5.append(diff5)
print(len(bmi_diff_5))
print("Sum for 5 percent: ")
print(sum(abs(number) for number in bmi_diff_5))
# + [markdown] id="fbtDEjOq3ZR1"
# 10 percent Median
#
# + colab={"base_uri": "https://localhost:8080/"} id="_1x21wqC6PJX" outputId="208b7d47-cb8f-4b4c-900d-a238c7861bc1"
df_bmi1 = df_bmi[['sample_bmi','bmi_copy_10_percent']]
imputed_bmi_df = df_bmi1.fillna(df_bmi1.median())
imputed_bmi_df.sample(10)
print(get_percent_missing(imputed_bmi_df))
imputed_bmi_df['original'] = df_bmi['bmi']
imputed_bmi_df['diff'] = imputed_bmi_df['original'] - imputed_bmi_df['bmi_copy_10_percent']
for i in bmi_10_idx:
print(str(imputed_bmi_df['bmi_copy_10_percent'][i])+", "+str(imputed_bmi_df['original'][i])+", "+str(imputed_bmi_df['diff'][i]))
# + id="LGpxTeaK6ZS-" colab={"base_uri": "https://localhost:8080/"} outputId="6bdda2b5-4e83-4e82-f36b-5a409c392c94"
# create list of difference bwtween imputed and orginal value
bmi_diff_1 = []
bmi_diff_5 = []
bmi_diff_10 = []
count = 0
for i in bmi_10_idx:
diff10 = abs(imputed_bmi_df['bmi_copy_10_percent'][i] - df_bmi['bmi'][i])
bmi_diff_10.append(diff10)
print("Sum for 10 perecent: ")
print(sum(abs(number) for number in bmi_diff_10))
# + [markdown] id="BN5xgwwGvPua"
# KNN here performs the best here since it has the minimum summation of differences. Then comes median and mean in order.
# + [markdown] id="fPazPl2VwAZG"
# ### IMPORTANT OBSERVATIONS
#
# ## What are the data types? (Only numeric and categorical)
# Datatypes- id,age,avg_glucose_level, bmi are all numerical variables whereas gender, hypertension, heart_disease, work_type, residence_type, smoking_status, stroke are categorical variables
#
# ## Are there missing values?
# There are missing values for BMI in the dataset. Rest of the values counts are balanced in the dataset.
#
# ## What are the likely distributions of the numeric variables?
# The distribution of age is not significantly skewed wheres the avg_glucose_leven and BMI are slightly right skewed
#
# ## Which independent variables are useful to predict a target
# From the correlation, VIF and logistic coefficients, we can infer that age is the most important variable in predicting stroke. Also, hypertension and previous heart jstory disease plays a significant role in predicting stroke. Correlation between age and stroke is high(around 0.34)
#
# ## Which independent variables have missing data? How much?
# BMI has 3.94 percent of missing data in the dataset found by isna function
#
# ## Do the training and test sets have the same data?
# The data is splitted into train and test sets. Train data has been scaled using MinMax Scalar here without touching the test dataset. Moreover, unimportant variables have been removed.
#
# ## In the predictor variables independent of all the other predictor variables?
# From the VIF and correlation, it was found that the glucose level, bmi and age may have a significant correlation.
#
# ## Which predictor variables are the most important?
# The most important predctor variables are age, bmi, hypertension and heart disease with a high correlation with stroke.
#
# ## Do the ranges of the predictor variables make sense?
# Initially it doesn't but after processing, yes the ranges of the standard variables make sense. Though there are some outliers, they have been removed and then the values are handles used min-max scalar tranformation
#
# ## What are the distributions of the predictor variables?
# The numerical predictor variables follows a normal distribution. Though the average glucose level and BMI shows some right skewness.
#
# ## Remove outliers and keep outliers (does if have an effect of the final predictive model)?
# Removing outliers significantly improved the F1 score and recall of logistic regression eg- F1 score went from 0.79 to 0.84
#
# ## Remove 1%, 5%, and 10% of your data randomly and impute the values back using at least 3 imputation methods. How well did the methods recover the missing values?
# All the three methods recovered the missing data well. The methods used were KNN, dropping, mean and median imputation methods. Though median was better than mean when there were outliers present in the data.
#
# Metrics:
# [[647 199]
# [ 9 25]]
# precision recall f1-score support
#
# 0.0 0.99 0.76 0.86 846
# 1.0 0.11 0.74 0.19 34
#
# accuracy 0.76 880
# Validation Acuuracy: 0.7636363636363637
# Training Accuracy: 0.7784174785946265
# F1 Score:
# 0.8357197216021109
# Recall
# 0.7500347656793214
#
#
#
#
# + [markdown] id="ggJZD4vFvbcZ"
# ## Conclusion
#
# Age is the most important variable in the dataset. It has a high correlation factor too. Hypertension and heart disease also play a significant role. Recall has been great with logistic Regression here. Though accuracy is not that great, F1 and recall have improved after balancing the dataset and removing outliers. We can have False negative here but cannot afford false positive in case of heart attacks and thats why recall is important.
# + [markdown] id="AY0e0vEFLG44"
# ### **Citations**
#
#
# https://www.analyticsvidhya.com/blog/2018/03/introduction-k-neighbours-algorithm-clustering/
#
# https://scikit-learn.org/stable/modules/grid_search.html
#
# https://www.knowledgehut.com/tutorials/machine-learning/hyperparameter-tuning-machine-learning
#
# https://www.geeksforgeeks.org/data-cleansing-introduction/
#
# https://stackoverflow.com/questions/52739323/python-data-cleaning
#
# https://towardsdatascience.com/6-different-ways-to-compensate-for-missing-values-data-imputation-with-examples-6022d9ca0779
#
#
#
# + [markdown] id="zbuajVBsMur6"
# # **License**
#
# MIT License
#
# Copyright (c) 2022 trivedi-mi
#
# Permission is hereby granted, free of charge, to any person obtaining a copy
# of this software and associated documentation files (the "Software"), to deal
# in the Software without restriction, including without limitation the rights
# to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
# copies of the Software, and to permit persons to whom the Software is
# furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all
# copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
# AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
# OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
# SOFTWARE.
#
| Data_Clean_Feature.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: uc-python
# language: python
# name: python3
# ---
# # Case Study
# ## Part 1
# *To be completed at the conclusion of Day 1*
# For the following exercises, you should use the data stored at `../data/companies.csv`
# You aren't expected to finish all the exercises; just get through as many as time allows and we will review them together.
#
# 1. Start by becoming familiar with the data. How many rows and how many columns does it have? What are the data types of the columns?
# 2. Set the data's index to be the "Symbol" column.
# 3. Look up the company with the symbol NCLH. What company is this? What sector is it in?
# 4. Filter down to companies that *either* in the "Consumer Discretionary" or the "Consumer Staples" sectors.
# 5. How many companies are left in the data now?
# 6. Create a new column, "Symbol_Length", that is the length of the symbol of each company. *Hint: you may need to reset an index along the way.*
# 7. Find the company named "Kroger Co.". Change its name to "The Kroger Company".
# **Bonus**: *For these two exercises, you won't find examples of the solution in our notebooks.
# You'll need to search for help on the internet.*
#
# *Don't worry if you aren't able to solve them.*
#
# 1. Filter down to companies whose symbol starts with A. How many companies meet this criterion?
# 2. What is the longest company name remaining in the dataset? You could just search the data visually, but try to find a programmatic solution.
# ## Part 2
# *To be completed at the conclusion of Day 2*
# This section again uses the data at `../data/companies.csv`.
#
# 1. Re-create the "Symbol_Length" column (see above).
# 2. What is the average symbol length of companies in the data set?
# 3. What is the average symbol length by sector? That is, after grouping by sector, what is the average symbol length for each group?
# 4. How long is the longest company name? How long is the longest company name by sector?
# Now open the pricing data at `../data/prices.csv`.
# Note that this data is entirely fabricated and does not exhibit the qualities of real stock market data!
#
# 1. Become familiar with this data. What is its shape? What are its data types?
# 2. Get summary metrics (count, min, max, standard deviation, etc) for both the Price and Quarter columns. *Hint: we saw a method of DataFrames that will do this for you in a single line.*
# 3. Perform an inner join between this data set and the companies data, on the Symbol column.
# 4. How many rows does our data have now?
# 5. What do you think this data represents? Form a hypothesis and look through the data more carefully until you are confident you understand what it is and how it is structured.
# 6. Group the data by sector. What is the average first quarter price for a company in the Real Estate sector? What is the minimum fourth quarter price for a company in the Industrials sector?
# 7. Filter the data down to just prices for Apple, Google, Microsoft, and Amazon.
# 8. Save this data as big_4.csv in the `../data` directory.
# 9. Using Seaborn, plot the price of these companies over 4 quarters. Encode the quarter as the x-axis, the price as the y-axis, and the company symbol as the hue.
# **Bonus**:
#
# This data is in a form that is useful for plotting.
# But in this shape, it would be quite difficult to calculate the difference between each company's fourth quarter price and its first quarter price.
#
# Reshape this data so it is of a form like the below:
#
# | Symbol | Name | Sector | Q1 | Q2 | Q3 | Q4 |
# |--------|------|--------|----|----|----|----|
# | AAPL | Apple Inc. | Information Technology | 275.20 | 269.96 | 263.51 | 266.07
#
# From which we could easily calculate Q4 - Q1.
#
# *You will probably want to google something like "python reshaping data". This is a very challenging problem!*
| notebooks/Case-Study.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Preliminares Matemáticos del Algoritmo de Maximización de la Esperanza
# <img style="float: right; margin: 0px 0px 15px 15px;" src="https://upload.wikimedia.org/wikipedia/commons/3/3d/EM_Process.jpg" width="500px" height="300px" />
#
# > En esta serie de cuadernos estaremos estudiando el algoritmo de maximización de la esperanza que introdujimos en la clase pasada en el contexto de mezclas Gaussianas.
#
# > Este interesante algoritmo sirve para entrenar los parámetros de una gran mayoría de los modelos con variables latentes, incluido el modelo de mezclas Gaussianas.
#
# > Sin embargo, para entenderlo es necesario tener claros varios conceptos matemáticos preliminares.
#
# > **Objetivos:**
# > - Explicar la desigualdad de Jensen para funciones cóncavas.
# > - Comprender la divergencia de Kullback-Leibler como una medida de proximidad de distribuciones.
# > - Probar propiedades sencillas relativas a la divergencia de Kullback-Leibler.
#
# > **Referencias:**
# > - Bayesian Methods for Machine Learning course, HSE University, Coursera.
# ## 1. Funciones cóncavas
# > *Definición.* Sea $f:\Omega \subseteq \mathbb{R}^{k} \to \mathbb{R}$ una función real. Decimos que $f$ es cóncava si para todo $x_1, x_2 \in \Omega$ y $\alpha \in (0, 1)$
# >
# > $$
# f(\alpha x_1 + (1 - \alpha) x_2) \geq \alpha f(x_1) + (1 - \alpha) f(x_2).
# $$
#
#
# La definición anterior lo que dice es que una función es cóncava si para cualesquier par de puntos sobre la gráfica de la función $f$, la función supera al segmento de recta que une esos dos puntos.
# Veámoslo a través de un ejemplo:
# Importamos librerías
import numpy as np
from matplotlib import pyplot as plt
# Definimos una función cóncava
def f(x):
return np.log(x)
# Vector de x
x = np.linspace(1, 100)
# Graficamos
alpha = np.linspace(0, 1)
x1 = 5
x2 = 80
plt.figure(figsize=(6, 4))
plt.plot(x, f(x), label="$f(x)$")
plt.plot(alpha * x1 + (1 - alpha) * x2, alpha * f(x1) + (1 - alpha) * f(x2),
label=r"$\alpha f(x_1) + (1 - \alpha) f(x_2)$")
plt.plot(x1, f(x1), 'or', label="$x_1$")
plt.plot(x2, f(x2), 'og', label="$x_2$")
plt.legend()
# **¿Qué ejemplos de funciones cóncavas se les vienen a la mente?**
#
# - $ax^2 + bx + c;$ con $a<0$.
# - $x^a$; con $0 < a < 1$.
# Seguramente recordarán de cálculo un resultado para funciones dos veces diferenciables:
#
# > *Teorema.* Sea $f: \mathbb{R} \to \mathbb{R}$ una función dos veces diferenciable en un intervalo abierto $(a, b)$. Entonces, $f$ es cóncava si y sólo si $f''(x) \leq 0$ para todo $x \in (a, b)$.
# Usar el teorema para comprobar la concavidad de las funciones:
#
# - $f(x) = \log x$.
# - $f(x) = x^a$, con $0 < a < 1$.
# ## 2. Desigualdad de Jensen
#
# Ahora, la definición de concavidad sólo involucra un par de puntos. Sin embargo, es posible extender esto para cualquier cantidad de puntos:
#
# > *Proposición.* Sea $f:\Omega \subseteq \mathbb{R}^{k} \to \mathbb{R}$ una función cóncava. Entonces para cualquier selección de números $\alpha_i \geq 0$, con $i = 1, \dots, m$, tales que $\sum_{i=1}^m \alpha_i = 1$ y cualquier selección de elementos $x_i \in \Omega$, con $i = 1, \dots, m$, se tiene que:
# >
# > $$
# f\left(\sum_{i=1}^{m} \alpha_i x_i\right) \geq \sum_{i=1}^m \alpha_i f(x_i)
# $$
# **¿La propiedad que deben cumplir las $\alpha_i$ se les hace conocida?**
#
# - $\alpha_i \geq 0$, con $i = 1, \dots, m$
# - $\sum_{i=1}^m \alpha_i = 1$
# En términos probabilísticos, esto se podría escribir como:
#
# > *Proposición.* Sea $f:\Omega \subseteq \mathbb{R}^{k} \to \mathbb{R}$ una función cóncava y $X$ una variable aleatoria (multivariable de dimensión $k$). Entonces
# >
# > $$
# f(E[X]) \geq E[f(X)].
# $$
# **Ejercicio.** Sea $X ~ \mathcal{N}(0, 1)$. Definimos la VA $Y$ como una función determinista de $X$, $Y=X^2 + 5$. ¿Cuál de las siguientes afirmaciones es cierta?
#
# - La desigualdad de Jensen no se puede aplicar puesto que $\log (x^2 + 5)$ no es cóncava.
# - $E[\log (x^2 + 5)] \geq \log E[x^2 + 5]$
# - $E[\log (x^2 + 5)] \leq \log E[x^2 + 5]$
# - $E[\log y] \geq \log E[y]$
# ## 3. Divergencia de Kullback-Leibler
# Comúnmente, no solo en el contexto del algoritmo de maximización de la esperanza, necesitaremos medir la diferencia (o similaridad) entre dos distribuciones de probabilidad.
#
# Una manera de medir esto es usando la **divergencia de Kullback-Leibler**.
# **Ejemplo.** Supongamos que tenemos dos Gaussianas:
from matplotlib import pyplot as plt
from scipy.stats import norm
import numpy as np
X = norm(loc=0, scale=1)
Y = norm(loc=1, scale=1)
x = np.linspace(-5, 5, 1001)
plt.plot(x, X.pdf(x), label=r"$\mathcal{N}(0, 1)$")
plt.plot(x, Y.pdf(x), label=r"$\mathcal{N}(1, 1)$")
plt.xlabel("$x$")
plt.legend()
# Una posible manera de medir la diferencia entre estas distribuciones sería medir la "distancia" entre los parámetros, la cual en este caso es 1.
#
# Sin embargo este enfoque tiene dos problemas:
#
# 1. ¿Qué pasa si las distribuciones que quiero comparar son de distintas familias?
#
# 2. Aún cuando las distribuciones son de la misma familia, veamos el siguiente caso:
X = norm(loc=0, scale=10)
Y = norm(loc=1, scale=10)
x = np.linspace(-20, 20, 1001)
plt.plot(x, X.pdf(x), label=r"$\mathcal{N}(0, 100)$")
plt.plot(x, Y.pdf(x), label=r"$\mathcal{N}(1, 100)$")
plt.xlabel("$x$")
plt.legend()
# En este caso, aplicando el mismo principio, la distancia también sería 1. Sin embargo, podemos apreciar que estas distribuciones se parecen muchísimo más que las anteriores. Por esto deberíamos considerar una medida alternativa de similitud / diferencia entre distribuciones.
#
# Esta medida es la **divergencia de Kullback-Leibler**.
#
# > *Definición.* Dadas dos distribuciones de probabilidad, la divergencia de Kullback-Leibler se define como:
# >
# > $$
# \mathcal{KL}(q || p) = \int_{-\infty}^{\infty} q(x) \log \frac{q(x)}{p(x)} d x = E_{q(x)}\left[\log \frac{q(x)}{p(x)}\right]
# $$
# >
# > si las variables son continuas, o
# >
# > $$
# \mathcal{KL}(q || p) = \sum_{x} q(x) \log \frac{q(x)}{p(x)} = E_{q(x)}\left[\log \frac{q(x)}{p(x)}\right]
# $$
# >
# > si las variables son discretas.
# Evaluemos:
from scipy.integrate import quad
# Primer conjunto de distribuciones
def caso1(x):
return norm.pdf(x, 0, 1) * np.log(norm.pdf(x, 0, 1) / norm.pdf(x, 1, 1))
val, err = quad(caso1, -10, 10)
val, err
# Segundo conjunto de distribuciones
def caso2(x):
return norm.pdf(x, 0, 10) * np.log(norm.pdf(x, 0, 10) / norm.pdf(x, 1, 10))
val, err = quad(caso2, -100, 100)
val, err
# Observamos que si evaluamos la diferencia entre las distribuciones en el primer caso (más distintas) obtenemos un valor de divergencia de 0.5, mientras que para el segundo caso (más parecidas) obtenemos un valor de divergencia de 0.005.
# ### Propiedades de la divergencia de Kullback-Leibler
#
# Tenemos las siguientes propiedades:
#
# 1. $\mathcal{KL}(q || p) \neq \mathcal{KL}(p || q)$.
# 2. $\mathcal{KL}(q || q) = 0$
# 3. $\mathcal{KL}(q || p) \geq 0$
#
# *Prueba.* En clase ... (Considerar el negativo de la divergencia y usar la desigualdad de Jensen).
# **Tarea (opcional - válida por una tarea anterior):**
#
# Supongamos que $q(x)=\mathcal{N}(x | \mu_1, \sigma_1^2)$ y $p(x)=\mathcal{N}(x | \mu_2, \sigma_2^2)$.
#
# Calcular:
#
# 1. $\mathcal{KL}(q || p)$.
#
# 2. $\mathcal{KL}(p || q)$.
# <script>
# $(document).ready(function(){
# $('div.prompt').hide();
# $('div.back-to-top').hide();
# $('nav#menubar').hide();
# $('.breadcrumb').hide();
# $('.hidden-print').hide();
# });
# </script>
#
# <footer id="attribution" style="float:right; color:#808080; background:#fff;">
# Created with Jupyter by <NAME>.
# </footer>
| Aplicaciones/AME-GMM/2_preliminares_matematicos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
from controleII import digital
from lcapy.discretetime import z, n
from lcapy import s,exp
from control import tf
# Questão 1
Xz = (10 * z + 5) / ((z-1) * (z-0.2))
teste = digital(Xz)
display(teste.izt())
print(teste.calcn())
# Questão 2
Xz = 1 + 2*z**-1 + 3*z**-2 + 4 *z **-3
teste = digital(Xz)
display(teste.izt())
print(teste.calcn())
# +
# Questão 4
Xz = (z**2 + z + 2) / ((z-1)* (z**2 -z + 1))
Q4 = digital(Xz=Xz)
display(Q4.izt())
Q4.calcn()
# -
# Questão 5
Xz = (2*z**3 + z) / ((z-2)**2 * (z-1))
Q5 = digital(Xz = Xz)
display(Q5.izt())
# Questão 6
Xz1 = 9 / (z * (1-2*z**-1)**2)
Xz2 = z / (z - 2)
Xz3 = 3*z / (z - 1)
Xz = Xz1 + Xz2 + Xz3
teste = digital(Xz)
display(teste.izt())
# +
# Questão 7
Xz = (z+2) / (z**2 * (z-2))
Q7 = digital(Xz = Xz)
display(Q7.izt())
# -
# Questão 13:
gs = 1 / (s * (s + 1))
gsc = tf([1], [1, 1])
teste = digital(Gs = gs, Gsc = gsc, tau=1)
display(teste.s2t())
display(teste.s2z())
# +
# questão 14
gss = 4 / (s * (s+2))
gsc = tf(4, [1, 2])
tau = .1
Q14 = digital(Gs=gss, Gsc=gsc, tau = .1)
display(Q14.s2t())
display(Q14.s2z())
fz, time, mag = Q14.Fz(20)
display(fz, time, mag)
# +
# questão 16
gss = 10 / (s + 10*s**2)
gsc = tf([10], [10, 1])
tau = 2
Q16 = digital(Gs=gss, Gsc=gsc, tau = 2)
display(Q16.s2z())
display(Q16.s2t())
fz, time, mag = Q16.Fz(14, fb = 0.05, plot=True)
display(fz, mag)
# +
g1s = 2 * (s - 6.93) / (s - 1) * (1 / (s * (s+1)))
gsc = tf(1, [1, 1]) #* 2* tf([1, -6.93], [1, -1])
tau = .1
Q17 = digital(Gs=g1s, Gsc=gsc, tau = tau)
display(Q17.s2z())
fz, time, mag = Q17.Fz(30, plot=True, save=True, title='Resposta ao degrau unitário sem o controlador')
display(fz)
# -
Xz = (0.130976*z - 0.26969) / (z**2 - 1.879029*z + 0.7303) * (z / (z-1))
C = N(Xz.partfrac())
C
from sympy import solve, var, N
var('u')
a =(u**2 - 2.01*u + 1) * u/(u-1)
solve(u, a)
# +
from sympy import var
from sympy import solve
k = var('k')
W = var('W')
t = (2 + W) / (2 - W)
Xz = k/(z-1) - ((1-exp(-1))/(z- exp(-1)))*k
#Xz = Xz.subs(z, t)
F = Xz/(Xz + 1)
G = F.simplify().as_numer_denom()[1]
G.subs(k, 25)
| ELT 331/lista3II.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import pymysql
import timeit
conn=pymysql.connect(host='172.16.58.3',port=6823,user='gml_read',passwd='<PASSWORD>',db='gml_base_qa',use_unicode=True, charset="utf8")
# SELECT goods_name,b.`name` FROM common_goods_info a LEFT JOIN category_info b ON a.`category_level2`=b.`id`
sql="select * from common_goods_info"
df_all=pd.read_sql(sql,con=conn)
conn.close()
# -
set(df_all["classify_status"])
df_all["classify_status"].unique()
df_all[["goods_name","goods_element","brand","specification_type"]]
df_all.loc[df_all["classify_status"]==0]
df_all["classify_status"]
df_all.head()
# +
import pandas as pd
import pymysql
import random
conn=pymysql.connect(host='172.16.58.3',port=6823,user='gml_read',passwd='<PASSWORD>',db='gml_base_qa',use_unicode=True, charset="utf8")
sql="""SELECT c.goods_name,d.category_1,d.category_2,c.average_recorded_price,c.brand,c.origin_country FROM common_goods_info c
LEFT JOIN (select a.id,a.name category_2, b.name category_1 from category_info a
LEFT JOIN (select * from category_info where parent_id=0) b on a.parent_id = b.id
where a.parent_id != 0) d ON c.category_level2=d.id"""
df_all=pd.read_sql(sql,con=conn)
conn.close()
# -
df_all[df_all["category_1"].isnull()]
| pytorch_textcnn/data_propre_v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="yf_vvLMZI4MH"
# Berdasarkan isu [#87](https://github.com/taruma/hidrokit/issues/87): **perhitungan debit andalan sungai dengan kurva durasi debit**
#
# Referensi isu:
# - SNI 6738:2015: **Perhitungan debit andalan sungai dengan kurva durasi debit**. Pranala: [sni.litbang.pu.go.id](http://sni.litbang.pu.go.id/index.php?r=/sni/new/sni/detail/id/458) diakses 10 desember 2019.
#
# Deskripsi Permasalahan:
# - Mencari nilai debit andal pada $Q_{80}$, $Q_{90}$, dan $Q_{95}$ pada periode tertentu (dalam satu tahun, atau setiap bulan).
#
# Strategi Penyelesaian Masalah:
# - Mengikuti panduan SNI 6738:2015.
# - Membuat fungsi dengan input `pandas.DataFrame` dengan jumlah observasi sembarang. Sehingga, penggunaan fungsi akan ditentukan.
# - Menyusun berurutan nilai dalam dataframe.
# - Membuat `list/array` yang merupakan nilai (kumulatif) probabilitas.
# - Mengembalikan d
# - Mencari nilai $Q_{80}$, $Q_{90}$, dan $Q_{95}$
#
# Catatan:
# - Dataset yang digunakan serupa dengan lampiran A dalam SNI 6738:2015.
# + [markdown] id="LeEnxFBwPS_r"
# # PERSIAPAN DAN DATASET
# + id="jpwIcCWjPy83"
# Unduh dataset
# !wget -O data.csv "https://taruma.github.io/assets/hidrokit_dataset/data_sni_67382015.csv" -q
FILE = 'data.csv'
# + id="gfn-_rHpIx1M"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + id="EVDi21tlQVOH" outputId="a9f875f5-cecf-4615-da58-e688cdcc02f7" colab={"base_uri": "https://localhost:8080/", "height": 306}
# Import dataset
dataset = pd.read_csv(FILE, index_col=0, header=0, parse_dates=True)
dataset.info()
dataset.head()
# + [markdown] id="AkGyGOpLP3Ut"
# # KODE
# + id="7AyNN8mTP4d0"
import numpy as np
import pandas as pd
def prob_weibull(m, n):
return m / (n + 1) * 100
def _array_weibull(n):
return np.array([prob_weibull(i, n) for i in range(1, n+1)])
def _fdc_xy(df):
n = len(df.index)
x = _array_weibull(n)
y = df.sort_values(ascending=False).values
return x, y
def _interpolate(probability, x, y):
return {p: np.interp(p, x, y) for p in probability}
def debit_andal(df, column, kind='table', prob=[80, 90, 95]):
x, y = _fdc_xy(df.loc[:, column])
if kind.lower() == 'array':
return x, y
if kind.lower() == 'prob':
return _interpolate(prob, x, y)
if kind.lower() == 'table':
data = {
'idx': df.loc[:, column].sort_values(ascending=False).index,
'rank': list(range(1, len(df.index)+1)),
'prob': x,
'data': y,
}
return pd.DataFrame(data)
def debit_andal_bulanan(df, column, **kwargs):
return {
m: debit_andal(df[df.index.month == m], column, **kwargs)
for m in range(1, 13)
}
# + [markdown] id="NZygLBVTnmmF"
# # PENGGUNAAN
# + [markdown] id="xnW38GFqLt0Y"
# ## Fungsi `.debit_andal()`
#
# Pada fungsi ini terdapat parameter yang perlu diperhatikan selain `df` dan `column` yaitu `kind`. Parameter `kind` menentukan hasil keluaran dari fungsi. Berikut nilai yang diterima oleh parameter `kind`:
# - `'array'`: keluaran berupa _tuple_ berisi dua `np.array` yaitu `x` (untuk sumbu x, probabilitas weibull) dan `y` (untuk sumbu y, nilai debit yang telah diurutkan).
# - `'table'` (**default**) : keluaran berupa `pandas.DataFrame` tabelaris yang berisikan kolom `idx` (indeks/tanggal kejadian), `rank` (ranking), `prob` (probabilitas weibull), `data` (nilai yang telah diurutkan).
# - `'prob'`: keluaran berupa _dictionary_ dengan _key_ sebagai nilai probabilitas dan _value_ sebagai nilai data yang dicari. Nilai tersebut diperoleh menggunakan fungsi interpolasi dari `numpy` yaitu `np.interp()`.
# + [markdown] id="os0fxLR3NoOP"
# ### `kind='array'`
# + id="t9b4K-DJTEYY" outputId="6f3b24f1-b31d-4534-fbf4-d2c1f6557421" colab={"base_uri": "https://localhost:8080/", "height": 51}
x, y = debit_andal(dataset, 'debit', kind='array')
print(f'len(x) = {len(x)}\tx[:5] = {x[:5]}')
print(f'len(y) = {len(y)}\ty[:5] = {y[:5]}')
# + [markdown] id="W5AuOxw7OBE4"
# ### `kind='table'` (**default**)
# + id="-Y2YqRHmODvy" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="3931eb74-a86a-4242-e382-1aface33ed10"
debit_andal(dataset, 'debit') # atau debit_andal(dataset, 'debit', kind='table')
# + [markdown] id="WfST4_5lOQQF"
# ### `kind='prob'`
#
# Nilai probabilitas yang digunakan yaitu $Q_{80}$, $Q_{90}$, $Q_{95}$ atau `[80, 90, 95`]
# + id="VS8XMabpOdWL" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1c34c1a9-e43e-4f2a-e08b-1e7979cfa8e6"
debit_andal(dataset, 'debit', kind='prob')
# atau debit_andal(dataset, 'debit', kind='prob', prob=[80, 90, 95])
# + [markdown] id="BvU5_3q_q3Tj"
# Contoh menggunakan nilai probabilitas yang berbeda `[30, 35, 70, 85, 95]`.
# + id="tknFyXIWOnu9" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="294cb9b2-8f31-4785-b356-fd22a21fe379"
debit_andal(dataset, 'debit', kind='prob', prob=[30, 35, 70, 85, 95])
# + [markdown] id="C7UY9njwPX2O"
# ## Fungsi `.debit_andal_bulanan()`
#
# Fungsi ini merupakan pengembangan lebih lanjut dari `.debit_andal()` yang dapat digunakan untuk membuat kurva durasi debit per bulan. Fungsi `.debit_andal_bulanan()` dapat menerima parameter yang sama dengan `.debit_andal()` seperti `kind` dan `prob`.
#
# _Key_ pada hasil keluaran fungsi ini menunjukkan bulan, contoh: `[1]` mengartikan bulan ke-1 (Januari).
# + id="OZfOduuESMEZ" outputId="42ecaca8-26a4-4d81-d248-c8bc6f11111d" colab={"base_uri": "https://localhost:8080/", "height": 51}
bulanan = debit_andal_bulanan(dataset, 'debit')
print(f'keys = {bulanan.keys()}')
print(f'values = {type(bulanan[1])}')
# out: berupa dataframe karena nilai kind='table' (default) fungsi debit_andal()
# + [markdown] id="D936WFTFSMsn"
# Menampilkan tabel untuk bulan Maret (ke-3)
# + id="uThGy3TBSMHX" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="df5fcc76-6416-4551-ee92-32f669b4a735"
bulanan[3].head()
# + [markdown] id="tp-olJpPq_DO"
# Contoh menampilkan nilai $Q_{80}, Q_{85}, Q_{90}, Q_{95}$ untuk setiap bulan
# + id="8Rl1RAKL0e1M" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="771885be-1245-4307-8c2d-b507421af63d"
bulanan_prob = debit_andal_bulanan(
dataset, 'debit', kind='prob', prob=[80, 85, 90, 95]
)
for key, value in bulanan_prob.items():
print('Bulan ke-', key, ':\t', value, sep='')
# + [markdown] id="BAZ739a7STA5"
# # Changelog
#
# ```
# - 20191214 - 1.0.0 - Initial
# ```
#
# #### Copyright © 2019 [<NAME>](https://taruma.github.io)
#
# Source code in this notebook is licensed under a [MIT License](https://choosealicense.com/licenses/mit/). Data in this notebook is licensed under a [Creative Common Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0/).
#
| booklet_hidrokit/0.4.0/ipynb/manual/taruma_0_3_5_hk87_debit_andal_kurva_durasi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Step 4: Create scenarios and edit input data in OpenAgua
#
# ### By <NAME>, Jan 2022
# ## Create new WEAP Scenarios in OpenAgua
# I updated these instructions to make it easy for you. No action is needed here but to view the new scenarios and look into how adding a new scenario works in OpenAgua
# * In OpenAgua, Go to the Bear River Network (2017) in the WEAP model project.
# * Click at the Scenario Icon on the left bar.
#
# #### The previous step has simplified this work and already uploaded the scenarios and their results to OpenAgua.
#
#
# * Click "Add Uncertainty to add a dummy scenario. Define a new scenario to the parent scenario "Bear River WEAP Model 2017 (Figure 1) and you can call it "Test".
#
# You will see these two scenarios are already there :)
# Cons25PercCacheUrbWaterUse
#
# Incr25PercCacheUrbWaterUse
#
# * Check out the WASH model. You'll see these scenarios: ConsDemand and IncrDemand
#
# * Next, you will edit input data for each scenario that changes the demand for conservation and growth cases.(see below)
#
# <img src="https://github.com/WamdamProject/WaMDaM-software-ecosystem/blob/master/mkdocs/Edit_MD_Files/images/Add_Scenarios.PNG?raw=true" style="float:center;width:600px;padding:20px">
# <h3><center>**Figure 1:** Add new scenarios </center></h3>
#
#
#
# # View input data to WEAP model new scenarios within OpenAgua
# * In the WEAP Model (Bear River WEAP Model 2017), we will view input data for three demand sites: Logan Potable, North Cache Potable, and South Cache Potable which are for Cache County urban demand. Its attribute called : Monthly Demand_Se
#
# * Use the drop down menu in OpenAgua to the right top corner to switch between scenarios.
# * For the Cons Demand scenario, we reduced demand by 25%. The easiest way to do it is to copy the Bear River WEAP Model 2017 base scenario demand into excel for all the months. Then create a formula to reduce demand by 25%. One way to do it is to get the 75% of the base demand which is like (=A1*.75). I already did that and you can copy and paste the values for you for both scenarios. So no action is needed here. But feel free to do the same on the "Test" scenario that you defined above.
#
#
# - Logan Potable
#
#
# - North Cache Potable
#
#
# - South Cache Potable
#
#
#
#
# * Do the same for the other scenario (Incr25PercCacheUrbWaterUse). But this time, increase it by 25% (like =A1*1.25)
# * Notice the autogenerated plot to the right and how the new scenarios are shifted up or down from the base scenario.
#
#
# #### Note: This screenshot is from an older OpenAgua version. Now it only shows the data and plot for one scenario at-a-time.
#
# <img src="https://github.com/WamdamProject/WaMDaM-software-ecosystem/blob/master/mkdocs/Edit_MD_Files/images/WEAP_Cons25PercCacheUrbWaterUse.PNG?raw=true" style="float:center;width:800px;padding:20px">
# <h3><center>**Figure 2:** Edit demand data for each new scenario in WEAP</center></h3>
#
#
# # View input data to WASH model new scenarios within OpenAgua
#
# * In the WASH Model, I changed change input data for the demand site called "j3" which is for Cache County urban demand. Its attribute called : dReqBase.
# * Use the drop down menu in OpenAgua to the right top corner to switch between scenarios.
# * For the Cons Demand scenario, we reduce demand by 25%. The easiest way to do it is to copy the OneYear base scenario demand into excel for all the months. Then create a formula to reduce demand by 25%. One way to do it is to get the 75% of the base demand which is like (=A1*.75)
# * No action is needed here as I already copied the new demand values and paste them into the ConsDemand column. (Figure 3).
# * Then I did the same for the other scenario (IncrDemand). But this time, increase it by 25% (like =A1*1.25)
# * Notice the autogenerated plot to the right and how the new scenarios are shifted up or down from the base scenario.
#
# <img src="https://github.com/WamdamProject/WaMDaM-software-ecosystem/blob/master/mkdocs/Edit_MD_Files/images/Edit_WASH.PNG?raw=true" style="float:center;width:900px;padding:20px">
# <h3><center>**Figure 3:** Edit demand data for each new scenario in WASH</center></h3>
#
# # Congratulations!
# * Now you have both WEAP and WASH models with two new scenarios each.
#
# * Next, we will use the WaMDaM Wizard to download them into a WaMDaM Excel Workbook and then into a SQLite database
#
# * We will also download a third model that is publicly available in OpenAgua for Monterrey Mexico
#
| 3_VisualizePublish/04_Step4_Create_New_Scenarios_Edit_OpenAgua.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Pymaceuticals Data Analysis</h1>
#
# ## Observations and Insights
# ###### 1. According to data given the best treatment results were achieved with Ramocane and Capomulin drug regimens. However, these two drug regimens have more data points than other drug regimens. When we check Standard Variantion and SEM we observe more stable results with Capomulin and Ramocane regimens.
# ###### 2. For further analysis we used two the best treatment results and another two from bottom of our list. Mouse gender were equally separated, 50.6% Male and 49.4% Female. While analysing of rpossible data outliers we found only one, which is definitely tells us the good quality of our data.
# ###### 3. Analysis of Capomuline and Ramicane treatment regimens show that tumor volume decrease were achieved. Tables "Results for a411 by Ramicane Regimen" and "Results for b128 by Capomulin Regimen" show the previous statement.
# ###### 4. Also we did analysis of Mouse weight affect to Tumor volume and results show positive regression.
# ###### 5. "Results for a203 by Infubinol Regimen" table shows ineffective treatment regimen where tumor volume increased.
# ###### 6. As an addition I would also consider Metastatic Sites quantity as a data to analyse, we can find that even successful treatment regimens were not able to prevent new metastatic sites appear.
# ## Dependencies and starter code
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import scipy.stats as st
import numpy as np
from scipy.stats import linregress
# Study data files
mouse_metadata = "data/Mouse_metadata.csv"
study_results = "data/Study_results.csv"
# Read the mouse data and the study results
mouse_metadata = pd.read_csv(mouse_metadata)
study_results = pd.read_csv(study_results)
# Combine the data into a single dataset
study_data = pd.merge(mouse_metadata, study_results, on="Mouse ID", how="outer")
study_data.rename(columns={"Tumor Volume (mm3)": "Tumor_Volume_mm3"})
study_data.head()
# -
# ## Summary statistics
# +
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
summary_calc=study_data[["Drug Regimen","Timepoint","Tumor Volume (mm3)","Weight (g)"]]
summary_data_grouped = summary_calc.groupby(['Drug Regimen','Timepoint'])
tumor_mean = summary_data_grouped['Tumor Volume (mm3)'].mean()
df = pd.DataFrame(tumor_mean)
df_pivot = pd.pivot_table(df, values='Tumor Volume (mm3)', index=['Drug Regimen'],
columns=['Timepoint'], aggfunc=np.mean)
df_pivot["Tumor Volume Mean (mm3)"] = df_pivot.iloc[:, 0:10].mean(axis=1)
df_pivot["Tumor Volume Median (mm3)"] = df_pivot.iloc[:, 0:10].median(axis=1)
df_pivot["Tumor Volume Variance (mm3)"] = df_pivot.iloc[:, 0:10].var(axis=1)
df_pivot["Tumor Volume Std Dev (mm3)"] = df_pivot.iloc[:, 0:10].std(axis=1)
df_pivot["Tumor Volume SEM (mm3)"] = df_pivot.iloc[:, 0:10].sem(axis=1)
summary = df_pivot[["Tumor Volume Mean (mm3)","Tumor Volume Median (mm3)","Tumor Volume Variance (mm3)",
"Tumor Volume Std Dev (mm3)", "Tumor Volume SEM (mm3)"]]
summary = summary.sort_values(by="Tumor Volume Mean (mm3)", ascending=True)
summary
# -
# ## Bar plots
# +
# Generate a bar plot showing number of data points for each treatment regimen using pandas
data_points = study_data['Drug Regimen'].value_counts()
dF = pd.DataFrame(data_points)
dF = dF.reset_index(drop=False)
dF = dF.sort_values(by="Drug Regimen", ascending=True)
x_axis= 'index'
y_axis = 'Drug Regimen'
pandas_bar = dF.plot.barh(x_axis, y_axis, title='Treatment Analysis', xlim=(0,240), ylim=(0,10), color="blue",
figsize = (13,5), width = 0.75, stacked=True)
pandas_bar.set_ylabel("Drug Regimen")
pandas_bar.set_xlabel("Data Points")
pandas_bar.grid(True, linestyle='--', which='major',
color='grey', alpha=.25)
# +
# Generate a bar plot showing number of data points for each treatment regimen using pyplot
x_axis= np.arange(len(dF))
y_axis = dF["Drug Regimen"]
plt.figure(figsize=(13,5))
bar_chart = plt.barh(x_axis, y_axis, color = 'b', alpha= 1, align='center', label='Drug Regimen')
plt.xlim(0,240)
plt.ylim(-0.75,9.75)
plt.title('Treatment Analysis')
plt.xlabel("Data Points")
plt.ylabel("Drug Regimen")
plt.legend(loc='best')
tick_locations = [value for value in x_axis]
plt.yticks(tick_locations, dF["index"], rotation="horizontal")
plt.grid(True, linestyle='--', which='major',
color='grey', alpha=.25)
plt.show()
# -
# ## Pie plots
# +
# Generate a pie plot showing the distribution of female versus male mice using pandas
male = (len(study_data.loc[study_data["Sex"] == "Male",:])/len(study_data["Sex"]))*100
female = (len(study_data.loc[study_data["Sex"] == "Female",:])/len(study_data["Sex"]))*100
sex_df = pd.DataFrame({"Sex": [male, female], "Mice":["Male", "Female"]})
sex_df = sex_df.set_index('Mice')
plot = sex_df.plot.pie(y="Sex", figsize =(4, 4), explode=(0.1,0), colors = 'cr', autopct='%1.1f%%',
startangle = 70, title="Male vs Female")
# -
# Generate a pie plot showing the distribution of female versus male mice using pyplot
male = (len(study_data.loc[study_data["Sex"] == "Male",:])/len(study_data["Sex"]))*100
female = (len(study_data.loc[study_data["Sex"] == "Female",:])/len(study_data["Sex"]))*100
sex_df = pd.DataFrame({"Sex": [male, female], "Mice":["Male", "Female"]})
sex_df = sex_df.set_index('Mice')
sizes =[male, female]
colors = ["c", "r"]
labels = ["Male","Female"]
fig, ax = plt.subplots(figsize=(4, 4), subplot_kw=dict(aspect="equal"))
plt.legend(labels, loc='best')
explode = (0.1, 0)
pie_chart = plt.pie(sizes, labels = labels,explode=explode, autopct = '%1.1f%%', colors=colors,
startangle = 70, shadow = True)
plt.legend(labels, loc='best')
plt.title('Male vs Female')
plt.ylabel("Sex")
plt.show()
# ## Quartiles, outliers and boxplots
# +
# Calculate the final tumor volume of each mouse across four of the most promising treatment regimens.
# Calculate the IQR and quantitatively determine if there are any potential outliers.
most_promise=study_data[["Mouse ID","Drug Regimen","Tumor Volume (mm3)"]]\
.groupby(["Mouse ID", "Drug Regimen" ]).last()\
.sort_values(by = "Drug Regimen", ascending = True).reset_index()
most_promise.set_index(["Drug Regimen", "Mouse ID"], inplace =True)
capomulin = most_promise.loc['Capomulin']["Tumor Volume (mm3)"]
quartiles_capomulin = capomulin.quantile([.25,.5,.75])
lower_capomulin = quartiles_capomulin[.25]
upper_capomulin = quartiles_capomulin[.75]
iqr_capomulin = upper_capomulin-lower_capomulin
capomulin_outliers = []
for value in capomulin:
if value > upper_capomulin + 1.5*iqr_capomulin:
capomulin_outliers.append(value)
elif value < lower_capomulin - 1.5*iqr_capomulin:
capomulin_outliers.append(value)
print(f"There is(are) " + str(len(capomulin_outliers)) + " outlier(s) in Campomulin and here is a list: " +
str(capomulin_outliers))
ramicane = most_promise.loc['Ramicane']["Tumor Volume (mm3)"]
quartiles_ramicane = ramicane.quantile([.25,.5,.75])
lower_ramicane = quartiles_ramicane[.25]
upper_ramicane = quartiles_ramicane[.75]
iqr_ramicane = upper_ramicane-lower_ramicane
ramicane_outliers = []
for value in ramicane:
if value > upper_ramicane + 1.5*iqr_ramicane:
ramicane_outliers.append(value)
elif value < lower_ramicane - 1.5*iqr_ramicane:
ramicane_outliers.append(value)
print(f"There is(are) " + str(len(ramicane_outliers)) + " outlier(s) in Ramicane and here is a list: " +
str(ramicane_outliers))
infubinol = most_promise.loc['Infubinol']["Tumor Volume (mm3)"]
quartiles_infubinol = infubinol.quantile([.25,.5,.75])
lower_infubinol = quartiles_infubinol[.25]
upper_infubinol = quartiles_infubinol[.75]
iqr_infubinol = upper_infubinol-lower_infubinol
infubinol_outliers = []
for value in infubinol:
if value > upper_infubinol + 1.5*iqr_infubinol:
infubinol_outliers.append(value)
elif value < lower_infubinol - 1.5*iqr_infubinol:
infubinol_outliers.append(value)
print(f"There is(are) " + str(len(infubinol_outliers)) + " outlier(s) in Infubinol and here is a list: " +
str(infubinol_outliers))
ceftamin = most_promise.loc['Ceftamin']["Tumor Volume (mm3)"]
quartiles_ceftamin = ceftamin.quantile([.25,.5,.75])
lower_ceftamin = quartiles_ceftamin[.25]
upper_ceftamin = quartiles_ceftamin[.75]
iqr_ceftamin = upper_ceftamin-lower_ceftamin
ceftamin_outliers = []
for value in ceftamin:
if value > upper_ceftamin + 1.5*iqr_ceftamin:
ceftamin_outliers.append(value)
elif value < lower_ceftamin - 1.5*iqr_ceftamin:
ceftamin_outliers.append(value)
print(f"There is(are) " + str(len(ceftamin_outliers)) + " outlier(s) in Ceftamin and here is a list: " +
str(ceftamin_outliers))
#Pseudo code: we can use here iteration to iterate through most_promise data frame,
# pick drug regimen and append quartile/iqr results to the list and then print the results.
# +
# Generate a box plot of the final tumor volume of each mouse across four regimens of interest
capomulin = list(capomulin)
ceftamin = list(ceftamin)
infubinol = list(infubinol)
ramicane = list(ramicane)
fig1, ax = plt.subplots()
ax.set_xticklabels(['Capomulin', 'Ceftamin', 'Infubinol', 'Ramicane'])
circle = dict(markerfacecolor='purple', marker='o')
square = dict(markerfacecolor='black', marker='s')
diamond = dict(markerfacecolor='b', marker='d')
pentagon = dict(markerfacecolor='g', marker='p')
data = [capomulin, ceftamin, infubinol, ramicane]
ax.set_title("Treatment Results on Box Plot", fontsize = 18)
ax.yaxis.grid(True, linestyle='-', which='major', color='lightgrey', alpha=0.3)
ax.set_xlabel('Drug Regimen')
ax.set_ylabel('Final Tumor Volume (mm3)')
box1 = ax.boxplot(capomulin, positions = [1], flierprops=circle, showmeans=True, meanline=True, patch_artist=True)
box2 = ax.boxplot(ceftamin, positions = [2], flierprops=square, showmeans=True, meanline=True,patch_artist=True)
box3 = ax.boxplot(infubinol,positions = [3], flierprops=diamond, showmeans=True, meanline=True, patch_artist=True)
box4 = ax.boxplot(ramicane,positions = [4], flierprops=pentagon, showmeans=True, meanline=True, patch_artist=True)
c1=['purple']
c2=['black']
c3=['blue']
c4=['green']
for patch, color in zip(box1['boxes'], c1):
patch.set_fc(color)
for patch, color in zip(box2['boxes'], c2):
patch.set_fc(color)
for patch, color in zip(box3['boxes'], c3):
patch.set_fc(color)
for patch, color in zip(box4['boxes'], c4):
patch.set_fc(color)
plt.show()
# -
# ## Line and scatter plots
# +
# Generate a line plot of time point versus tumor volume for a mouse treated with Capomulin
capomulin_data =study_data[["Drug Regimen","Timepoint", "Mouse ID","Tumor Volume (mm3)"]]
capomulin_data.set_index(["Drug Regimen"], inplace =True)
capomulin_dat = capomulin_data.loc["Capomulin", :]
capomulin_dat = capomulin_dat.sort_values(by="Mouse ID", ascending = True)
capomulin_dat = capomulin_dat.head(10)
capomulin_dat = capomulin_dat.sort_values(by="Timepoint", ascending = True)
line= capomulin_dat.plot.line(x="Timepoint", y="Tumor Volume (mm3)", xlim=(-1,46), ylim=(37,46),color="green",
figsize = (13,5), fontsize = 15, grid=True)
line.set_ylabel("Tumor Volume", fontsize = 15)
line.set_xlabel("Timepoint", fontsize = 15)
line.set_title(' Results for b128 by Capomulin Regimen', fontsize=15)
# Additional plots for comparison
ramicane_data = capomulin_data.loc["Ramicane", :]
ramicane_data = ramicane_data.sort_values(by="Mouse ID", ascending = True)
ramicane_data = ramicane_data.head(10)
ramicane_data = ramicane_data.sort_values(by="Timepoint", ascending = True)
line= ramicane_data.plot.line(x="Timepoint", y="Tumor Volume (mm3)", xlim=(-1,46), ylim=(37,46),color="blue",
figsize = (13,5), fontsize = 15, grid=True)
line.set_ylabel("Tumor Volume", fontsize = 15)
line.set_xlabel("Timepoint", fontsize = 15)
line.set_title(' Results for a411 by Ramicane Regimen', fontsize=15)
infubinol_data = capomulin_data.loc["Infubinol", :]
infubinol_data = infubinol_data.sort_values(by="Mouse ID", ascending = True)
infubinol_data = infubinol_data.head(10)
infubinol_data = infubinol_data.sort_values(by="Timepoint", ascending = True)
line= infubinol_data.plot.line(x="Timepoint", y="Tumor Volume (mm3)", xlim=(-1,46), ylim=(44,70),color="purple",
figsize = (13,5), fontsize = 15, grid=True)
line.set_ylabel("Tumor Volume", fontsize = 15)
line.set_xlabel("Timepoint", fontsize = 15)
line.set_title(' Results for a203 by Infubinol Regimen', fontsize=15)
# +
# Generate a scatter plot of mouse weight versus average tumor volume for the Capomulin regimen
# Additional line added to the plot to make it more readable
weight_df = study_data[["Drug Regimen","Tumor Volume (mm3)","Weight (g)"]]\
.groupby(["Weight (g)"]).mean()
weight_df = weight_df.reset_index(drop=False)
plt.figure(figsize=(15,5))
plt.xlim(14,31)
plt.ylim(35.5,60)
plt.scatter(weight_df['Weight (g)'], weight_df['Tumor Volume (mm3)'], marker="o", facecolors="blue", edgecolors="black")
plt.ylabel("Average Tumor Volume (mm3)", fontsize = 12)
plt.xlabel("Weight (g)", fontsize = 12)
plt.title("Mouse weight vs Average Tumor Volume", fontsize=14)
plt.plot(weight_df['Weight (g)'], weight_df['Tumor Volume (mm3)'], color='blue', label="Weight (g)")
plt.legend(loc="best")
plt.grid(True, linestyle='--', which='major', color='grey', alpha=.25)
plt.show()
# -
# Calculate the correlation coefficient and linear regression model for mouse weight and average tumor volume for the Capomulin regimen
x_values = weight_df["Weight (g)"]
y_values = weight_df["Tumor Volume (mm3)"]
(slope, intercept, rvalue, pvalue, stderr)=linregress(x_values,y_values)
regress_values = x_values*slope + intercept
line_eq ="y= " + str(round(slope,2)) + "x + " +str(round(intercept,2))
coef = x_values.corr(y_values)
plt.figure(figsize=(15,5))
plt.xlim(14,31)
plt.ylim(32,60)
plt.scatter(x_values, y_values, marker="o", facecolors="blue", edgecolors="black" )
plt.plot(x_values, regress_values,"r-", label="Regression")
plt.annotate(line_eq,(18, 40), fontsize=15, color="red", rotation = "15")
plt.ylabel("Average Tumor Volume (mm3)", fontsize = 12)
plt.xlabel("Weight (g)", fontsize = 12)
plt.title("Mouse weight vs Average Tumor Volume", fontsize=14)
plt.plot(weight_df['Weight (g)'], weight_df['Tumor Volume (mm3)'], color='blue', label="Weight (g)")
plt.legend(loc="best")
plt.grid(True, linestyle='--', which='major', color='grey', alpha=.25)
plt.show()
print(f" Correlation coefficient is equal to: " + str(coef))
| Pymaceuticals/pymaceuticals_starter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:Anaconda2]
# language: python
# name: conda-env-Anaconda2-py
# ---
import numpy as np
import random
import matplotlib.pyplot as plt
import pdb
# +
# INPUT PARAMETERS
def input_param():
w = 1000000 # time window [in ms]
g = 0.1 # gamma parameter
d = 10 # base difficulty
MAX_TX = 5000 # number of txs generated
return w, g, d, MAX_TX
# +
####### SINGLE NODE ANALYSIS WITH ADAPTIVE POW ########
#######################################################
tx = []
delay = []
dif = []
mu = 10**-7 # CPU power
[w, g, d, MAX_TX] = input_param()
r = 0.3 + 0.4*np.random.rand(MAX_TX)
x = d
for i in range(MAX_TX):
# PoW time (40 is a scale factor to convert time in ms)
delay.append(mu*3**x*r[i]/40)
if i > 0:
tx.append(tx[i-1] + mu*3**x*r[i]/40)
else:
tx.append(mu*3**x*r[i]/40)
dif.append(x)
# adapt difficulty
a = (sum(1 for j in tx if j <= tx[i] and j > max(tx[i] - w, 0))) # compute how many txs in the prev time window
x = d + np.floor(a*g) # set the new difficulty
if i % 500 == 0:
print(i)
# +
# PoW difficulty plot
fig = plt.figure()
plt.plot(dif)
plt.xlabel("Sequence of txs")
plt.ylabel("PoW difficulty")
plt.grid()
plt.show()
# +
# Time to compute the PoW
fig = plt.figure()
plt.plot(np.array(delay)/1000, linewidth=0.2)
plt.plot(np.mean(delay)*np.ones((MAX_TX))/1000, color="r", linewidth=2)
plt.xlabel("Sequence of txs")
plt.ylabel("PoW time")
plt.grid()
plt.show()
# +
# Throughput [tps]
fig = plt.figure()
plt.semilogy((np.array(range(MAX_TX))+1)*1000/np.array(tx))
plt.grid()
plt.xlabel("Sequence of txs")
plt.ylabel("Throughput")
plt.show()
# +
######## SINGLE NODE ANALYSIS WITH FIXED POW ##########
#######################################################
MAX_TX = 5000
tx = []
delay = []
mu = 10**-7 # CPU power
# set the PoW difficulty x as the base difficulty
x = 14
r = 0.3 + 0.4*np.random.rand(MAX_TX)
for i in range(MAX_TX):
# PoW time (40 is a scale factor to convert time in ms)
delay.append(mu*3**x*r[i]/40)
if i > 0:
tx.append(tx[i-1] + mu*3**x*r[i]/40)
else:
tx.append(mu*3**x*r[i]/40)
if i % 500 == 0:
print(i)
# +
# Time to compute the PoW
fig = plt.figure()
plt.plot(np.array(delay)/1000, linewidth=0.2)
plt.plot(np.mean(delay)*np.ones((MAX_TX))/1000, color="r", linewidth=2)
plt.xlabel("Sequence of txs")
plt.ylabel("PoW time")
plt.grid()
plt.show()
# +
# Throughput [tps]
fig = plt.figure()
plt.plot((np.array(range(MAX_TX))+1)*1000/np.array(tx))
plt.grid()
plt.xlabel("Sequence of txs")
plt.ylabel("Throughput")
plt.show()
# +
######## SENSITIVITY ANALYSIS ON GAMMA PARAMETER #########
##########################################################
MAX_TX = 2000
tps = []
mu_vector = [10**0, 10**-1, 10**-7] # CPU power
g_vector = [0, 0.001, 0.01, 0.1, 0.2, 0.5, 1]
[w, _, d, _] = input_param()
for g in g_vector:
print("Gamma = " + str(g))
for mu in mu_vector:
# set the PoW difficulty x as the base difficulty
x = d
tx = []
r = 0.3 + 0.4*np.random.rand(MAX_TX)
for i in range(MAX_TX):
# PoW time (40 is a scale factor to convert time in ms)
#delay.append(mu*3**x*r[i]/40)
if i > 0:
tx.append(tx[i-1] + mu*3**x*r[i]/40)
else:
tx.append(mu*3**x*r[i]/40)
#dif.append(x)
# adapt difficulty
a = (sum(1 for j in tx if j <= tx[i] and j > max(tx[i] - w, 0))) # compute how many txs in the prev time window
x = d + np.floor(a*g) # set the new difficulty
print((MAX_TX+1)*1000/np.array(tx[MAX_TX - 1]))
tps.append((MAX_TX+1)*1000/np.array(tx[MAX_TX - 1]))
# +
# Plot - sensitivity analysis of gamma parameter
t = np.reshape(tps, (7, 3))
fig = plt.figure(figsize=(10, 6))
plt.loglog(g_vector, t)
plt.grid()
plt.ylabel("throughput [tps]")
plt.xlabel("gamma")
plt.legend(["IoT", "Laptop", "FPGA"])
plt.show()
| adaptive-ratecontrol.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!/usr/bin/env python3
# -*- coding: utf-8 -*-
"""
@author: gkweston
github: https://github.com/gkweston
This script takes merged data files and marks stress moments 60 seconds before and after user
flagged instances. Then data is windowed by 120 second intervals, dropping any window w/ at
least 20 missing heart rate values. As windows are processed, a file is compiled for each
participant with applicably extracted heartrate features.
"""
import glob as gb
import pandas as pd
import numpy as np
# -
masterDir = r"/Users/gkweston/git/ptsd-continuous-monitoring/data/"
inputDir = masterDir + r"prep_output/pre_window/"
outputDir = masterDir + r"windowed/"
featureDir = masterDir + r"/features/"
monitorDir = r"/Users/gkweston/Desktop/workingFiles/acelab/proj2/data_monitor/"
fileList = gb.glob(inputDir + "*.csv")
# +
# Creating empty dataframe to extract features
fCols=['window_num', 'participant', 'mean_hr',
'min_hr', 'max_hr', 'range_hr', 'smoment']
feat_df = pd.DataFrame(columns=fCols)
# Cleaning up dataframe and updating stressmoment 60 s before and 60 after recorded time
for file in fileList:
participant = file[87:89]
open_file = pd.read_csv(file, low_memory=False)
prim_df = pd.DataFrame(open_file)
prim_df.drop('V1', axis=1, inplace=True)
prim_df.drop('Unnamed: 0', axis=1, inplace=True)
prim_df['window'] = np.nan
momentIndex = prim_df[prim_df['smoment']==1].index
for moment in momentIndex:
prim_df['smoment'][moment-60 : moment+60]=1
# Creating temporary dataframe, if leq 20 missing hr in temp append to final dataframe
# and add window num
fin_df = pd.DataFrame()
nanThresh = 20
winNum = 1
for i in range(0, prim_df.shape[0], 120):
temp_df = prim_df[i : i + 120]
hrNanCount = temp_df['hr'].isna().sum()
smCount = temp_df.smoment.count()
if hrNanCount < nanThresh:
# Updating stress moments in each window
if smCount > 0:
temp_df['smoment'] = 1
stress = 1
else:
stress = 0
temp_df['window'] = winNum
# Extracting features by window specific DataFrame
meanHr = temp_df['hr'].mean(skipna=True)
minHr = temp_df['hr'].min(skipna=True)
maxHr = temp_df['hr'].max(skipna=True)
rngeHr = maxHr - minHr
tFeatData = [[winNum, participant, meanHr, minHr, maxHr, rngeHr, stress]]
tFeat_df = pd.DataFrame(data=tFeatData, columns=fCols)
feat_df = feat_df.append(tFeat_df)
fin_df = fin_df.append(temp_df)
winNum += 1
# Output final data sets
fin_df.to_csv(outputDir + file[75:])
fin_df.to_csv(monitorDir + file[75:])
feat_df.to_csv(featureDir + "feature.csv")
print(f"\nAll processes complete succesfuly.")
# -
| windowing_featuring.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Weakly Supervised Recommendation Systems
# Experiments steps:
# 1. **User's Preferences Model**: Leverage the most *explicit* ratings to build a *rate/rank prediction model*. This is a simple *Explicit Matrix Factorization* model.
# 2. **Generate Weak DataSet**: Use the above model to *predict* for all user/item pairs $(u,i)$ in *implicit feedback dataset* to build a new *weak explicit dataset* $(u, i, r^*)$.
# 3. **Evaluate**: Use the intact test split in the most explicit feedback, in order to evaluate the performance of any model.
# ## Explicit Model Experiments
# This section contains all the experiments based on the explicit matrix factorization model.
# ### Explicit Rate Model
# +
import utils
from spotlight.evaluation import rmse_score
dataset_recommend_train, dataset_recommend_test, dataset_recommend_dev, dataset_read, dataset_wish = utils.parse_douban()
print('Explicit dataset (TEST) contains %s interactions of %s users and %s items'%(
format(len(dataset_recommend_test.ratings), ','),
format(dataset_recommend_test.num_users, ','),
format(dataset_recommend_test.num_items, ',')))
print('Explicit dataset (VALID) contains %s interactions of %s users and %s items'%(
format(len(dataset_recommend_dev.ratings), ','),
format(dataset_recommend_dev.num_users, ','),
format(dataset_recommend_dev.num_items, ',')))
print('Explicit dataset (TRAIN) contains %s interactions of %s users and %s items'%(
format(len(dataset_recommend_train.ratings), ','),
format(dataset_recommend_train.num_users, ','),
format(dataset_recommend_train.num_items, ',')))
print('Implicit dataset (READ/READING/TAG/COMMENT) contains %s interactions of %s users and %s items'%(
format(len(dataset_read.ratings), ','),
format(dataset_read.num_users, ','),
format(dataset_read.num_items, ',')))
print('Implicit dataset (WISH) contains %s interactions of %s users and %s items'%(
format(len(dataset_wish.ratings), ','),
format(dataset_wish.num_users, ','),
format(dataset_wish.num_items, ',')))
# train the explicit model based on recommend feedback
model = utils.train_explicit(train_interactions=dataset_recommend_train,
valid_interactions=dataset_recommend_dev,
run_name='model_douban_explicit_rate')
# evaluate the new model
mrr, ndcg, ndcg10, ndcg_5, mmap, success_10, success_5 = utils.evaluate(interactions=dataset_recommend_test,
model=model,
topk=20)
rmse = rmse_score(model=model, test=dataset_recommend_test)
print('-'*20)
print('RMSE: {:.4f}'.format(rmse))
print('MRR: {:.4f}'.format(mrr))
print('nDCG: {:.4f}'.format(ndcg))
print('nDCG@10: {:.4f}'.format(ndcg10))
print('nDCG@5: {:.4f}'.format(ndcg_5))
print('MAP: {:.4f}'.format(mmap))
print('success@10: {:.4f}'.format(success_10))
print('success@5: {:.4f}'.format(success_5))
# -
# ## Remove valid/test ratings
# +
test_interact = set()
for (uid, iid) in zip(dataset_recommend_test.user_ids, dataset_recommend_test.item_ids):
test_interact.add((uid, iid))
for (uid, iid) in zip(dataset_recommend_dev.user_ids, dataset_recommend_dev.item_ids):
test_interact.add((uid, iid))
# clean implicit dataset from test/dev rating
for idx, (uid, iid, r) in enumerate(zip(dataset_read.user_ids, dataset_read.item_ids, dataset_read.ratings)):
if (uid, iid) in test_interact:
dataset_read.ratings[idx] = -1
# -
# ### Explicit Read/Reading/Tag/Comment Model
# Leverage the **explicit rate model** trained at the previous section to annotate **missing values** in the **read/reading/tag/comment** dataset.
# +
# annotate the missing values in the play dataset based on the explicit recommend model
dataset_read = utils.annotate(interactions=dataset_read,
model=model,
run_name='dataset_douban_read_explicit_annotated')
# train the explicit model based on recommend feedback
model = utils.train_explicit(train_interactions=dataset_read,
valid_interactions=dataset_recommend_dev,
run_name='model_douban_explicit_read')
# evaluate the new model
mrr, ndcg, ndcg10, ndcg_5, mmap, success_10, success_5 = utils.evaluate(interactions=dataset_recommend_test,
model=model,
topk=20)
rmse = rmse_score(model=model, test=dataset_recommend_test)
print('-'*20)
print('RMSE: {:.4f}'.format(rmse))
print('MRR: {:.4f}'.format(mrr))
print('nDCG: {:.4f}'.format(ndcg))
print('nDCG@10: {:.4f}'.format(ndcg10))
print('nDCG@5: {:.4f}'.format(ndcg_5))
print('MAP: {:.4f}'.format(mmap))
print('success@10: {:.4f}'.format(success_10))
print('success@5: {:.4f}'.format(success_5))
# -
# ## Implicit Model Experiments
# This section contains all the experiments based on the implicit matrix factorization model.
# ### Implicit Model using Negative Sampling
# +
import utils
from spotlight.evaluation import rmse_score
dataset_recommend_train, dataset_recommend_test, dataset_recommend_dev, dataset_read, dataset_wish = utils.parse_douban()
print('Explicit dataset (TEST) contains %s interactions of %s users and %s items'%(
format(len(dataset_recommend_test.ratings), ','),
format(dataset_recommend_test.num_users, ','),
format(dataset_recommend_test.num_items, ',')))
print('Explicit dataset (VALID) contains %s interactions of %s users and %s items'%(
format(len(dataset_recommend_dev.ratings), ','),
format(dataset_recommend_dev.num_users, ','),
format(dataset_recommend_dev.num_items, ',')))
print('Explicit dataset (TRAIN) contains %s interactions of %s users and %s items'%(
format(len(dataset_recommend_train.ratings), ','),
format(dataset_recommend_train.num_users, ','),
format(dataset_recommend_train.num_items, ',')))
print('Implicit dataset (READ/READING/TAG/COMMENT) contains %s interactions of %s users and %s items'%(
format(len(dataset_read.ratings), ','),
format(dataset_read.num_users, ','),
format(dataset_read.num_items, ',')))
print('Implicit dataset (WISH) contains %s interactions of %s users and %s items'%(
format(len(dataset_wish.ratings), ','),
format(dataset_wish.num_users, ','),
format(dataset_wish.num_items, ',')))
# train the explicit model based on recommend feedback
model = utils.train_implicit_negative_sampling(train_interactions=dataset_read,
valid_interactions=dataset_recommend_dev,
run_name='model_douban_implicit_read2')
# evaluate the new model
mrr, ndcg, ndcg10, ndcg_5, mmap, success_10, success_5 = utils.evaluate(interactions=dataset_recommend_test,
model=model,
topk=20)
rmse = rmse_score(model=model, test=dataset_recommend_test)
print('-'*20)
print('RMSE: {:.4f}'.format(rmse))
print('MRR: {:.4f}'.format(mrr))
print('nDCG: {:.4f}'.format(ndcg))
print('nDCG@10: {:.4f}'.format(ndcg10))
print('nDCG@5: {:.4f}'.format(ndcg_5))
print('MAP: {:.4f}'.format(mmap))
print('success@10: {:.4f}'.format(success_10))
print('success@5: {:.4f}'.format(success_5))
# -
# ## Popularity
# +
import utils
from popularity import PopularityModel
from spotlight.evaluation import rmse_score
dataset_recommend_train, dataset_recommend_test, dataset_recommend_dev, dataset_read, dataset_wish = utils.parse_douban()
print('Explicit dataset (TEST) contains %s interactions of %s users and %s items'%(
format(len(dataset_recommend_test.ratings), ','),
format(dataset_recommend_test.num_users, ','),
format(dataset_recommend_test.num_items, ',')))
print('Explicit dataset (VALID) contains %s interactions of %s users and %s items'%(
format(len(dataset_recommend_dev.ratings), ','),
format(dataset_recommend_dev.num_users, ','),
format(dataset_recommend_dev.num_items, ',')))
print('Explicit dataset (TRAIN) contains %s interactions of %s users and %s items'%(
format(len(dataset_recommend_train.ratings), ','),
format(dataset_recommend_train.num_users, ','),
format(dataset_recommend_train.num_items, ',')))
print('Implicit dataset (READ/READING/TAG/COMMENT) contains %s interactions of %s users and %s items'%(
format(len(dataset_read.ratings), ','),
format(dataset_read.num_users, ','),
format(dataset_read.num_items, ',')))
print('Implicit dataset (WISH) contains %s interactions of %s users and %s items'%(
format(len(dataset_wish.ratings), ','),
format(dataset_wish.num_users, ','),
format(dataset_wish.num_items, ',')))
# train the explicit model based on recommend feedback
model = PopularityModel()
print('fit the model')
model.fit(interactions=dataset_recommend_train)
# evaluate the new model
print('evaluate the model')
mrr, ndcg, ndcg10, ndcg_5, mmap, success_10, success_5 = utils.evaluate(interactions=dataset_recommend_test,
model=model,
topk=20)
# rmse = rmse_score(model=model, test=dataset_recommend_test, batch_size=512)
# print('-'*20)
# print('RMSE: {:.4f}'.format(rmse))
print('MRR: {:.4f}'.format(mrr))
print('nDCG: {:.4f}'.format(ndcg))
print('nDCG@10: {:.4f}'.format(ndcg10))
print('nDCG@5: {:.4f}'.format(ndcg_5))
print('MAP: {:.4f}'.format(mmap))
print('success@10: {:.4f}'.format(success_10))
print('success@5: {:.4f}'.format(success_5))
| weak_recsys_douban.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Regularization and Gradient Descent Exercises
#
# ## Introduction
#
# We will begin with a short tutorial on regression, polynomial features, and regularization based on a very simple, sparse data set that contains a column of `x` data and associated `y` noisy data. The data file is called `X_Y_Sinusoid_Data.csv`.
from __future__ import print_function
import os
data_path = ['data']
# + [markdown] run_control={"marked": true}
# ## Question 1
#
# * Import the data.
#
# * Also generate approximately 100 equally spaced x data points over the range of 0 to 1. Using these points, calculate the y-data which represents the "ground truth" (the real function) from the equation: $y = sin(2\pi x)$
#
# * Plot the sparse data (`x` vs `y`) and the calculated ("real") data.
#
# +
import pandas as pd
import numpy as np
filepath = os.sep.join(data_path + ['X_Y_Sinusoid_Data.csv'])
data = pd.read_csv(filepath)
X_real = np.linspace(0, 1.0, 100)
Y_real = np.sin(2 * np.pi * X_real)
# +
import matplotlib.pyplot as plt
import seaborn as sns
% matplotlib inline
# +
sns.set_style('white')
sns.set_context('talk')
sns.set_palette('dark')
# Plot of the noisy (sparse)
ax = data.set_index('x')['y'].plot(ls='', marker='o', label='data')
ax.plot(X_real, Y_real, ls='--', marker='', label='real function')
ax.legend()
ax.set(xlabel='x data', ylabel='y data');
# -
# ## Question 2
#
#
# * Using the `PolynomialFeatures` class from Scikit-learn's preprocessing library, create 20th order polynomial features.
# * Fit this data using linear regression.
# * Plot the resulting predicted value compared to the calculated data.
#
# Note that `PolynomialFeatures` requires either a dataframe (with one column, not a Series) or a 2D array of dimension (`X`, 1), where `X` is the length.
# +
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import LinearRegression
# Setup the polynomial features
degree = 20
pf = PolynomialFeatures(degree)
lr = LinearRegression()
# Extract the X- and Y- data from the dataframe
X_data = data[['x']]
Y_data = data['y']
# Create the features and fit the model
X_poly = pf.fit_transform(X_data)
lr = lr.fit(X_poly, Y_data)
Y_pred = lr.predict(X_poly)
# Plot the result
plt.plot(X_data, Y_data, marker='o', ls='', label='data', alpha=1)
plt.plot(X_real, Y_real, ls='--', label='real function')
plt.plot(X_data, Y_pred, marker='^', alpha=.5, label='predictions w/ polynomial features')
plt.legend()
ax = plt.gca()
ax.set(xlabel='x data', ylabel='y data');
# -
# ## Question 3
#
# * Perform the regression on using the data with polynomial features using ridge regression ($\alpha$=0.001) and lasso regression ($\alpha$=0.0001).
# * Plot the results, as was done in Question 1.
# * Also plot the magnitude of the coefficients obtained from these regressions, and compare them to those obtained from linear regression in the previous question. The linear regression coefficients will likely need a separate plot (or their own y-axis) due to their large magnitude.
#
# What does the comparatively large magnitude of the data tell you about the role of regularization?
# +
# Mute the sklearn warning about regularization
import warnings
warnings.filterwarnings('ignore', module='sklearn')
from sklearn.linear_model import Ridge, Lasso
# The ridge regression model
rr = Ridge(alpha=0.001)
rr = rr.fit(X_poly, Y_data)
Y_pred_rr = rr.predict(X_poly)
# The lasso regression model
lassor = Lasso(alpha=0.0001)
lassor = lassor.fit(X_poly, Y_data)
Y_pred_lr = lassor.predict(X_poly)
# The plot of the predicted values
plt.plot(X_data, Y_data, marker='o', ls='', label='data')
plt.plot(X_real, Y_real, ls='--', label='real function')
plt.plot(X_data, Y_pred, label='linear regression', marker='^', alpha=.5)
plt.plot(X_data, Y_pred_rr, label='ridge regression', marker='^', alpha=.5)
plt.plot(X_data, Y_pred_lr, label='lasso regression', marker='^', alpha=.5)
plt.legend()
ax = plt.gca()
ax.set(xlabel='x data', ylabel='y data');
# +
# let's look at the absolute value of coefficients for each model
coefficients = pd.DataFrame()
coefficients['linear regression'] = lr.coef_.ravel()
coefficients['ridge regression'] = rr.coef_.ravel()
coefficients['lasso regression'] = lassor.coef_.ravel()
coefficients = coefficients.applymap(abs)
coefficients.describe() # Huge difference in scale between non-regularized vs regularized regression
# +
colors = sns.color_palette()
# Setup the dual y-axes
ax1 = plt.axes()
ax2 = ax1.twinx()
# Plot the linear regression data
ax1.plot(lr.coef_.ravel(),
color=colors[0], marker='o', label='linear regression')
# Plot the regularization data sets
ax2.plot(rr.coef_.ravel(),
color=colors[1], marker='o', label='ridge regression')
ax2.plot(lassor.coef_.ravel(),
color=colors[2], marker='o', label='lasso regression')
# Customize axes scales
ax1.set_ylim(-2e14, 2e14)
ax2.set_ylim(-25, 25)
# Combine the legends
h1, l1 = ax1.get_legend_handles_labels()
h2, l2 = ax2.get_legend_handles_labels()
ax1.legend(h1+h2, l1+l2)
ax1.set(xlabel='coefficients',ylabel='linear regression')
ax2.set(ylabel='ridge and lasso regression')
ax1.set_xticks(range(len(lr.coef_)));
# -
# ## Question 4
#
# For the remaining questions, we will be working with the [data set](https://www.kaggle.com/c/house-prices-advanced-regression-techniques) from last lesson, which is based on housing prices in Ames, Iowa. There are an extensive number of features--see the exercises from week three for a discussion of these features.
#
# To begin:
#
# * Import the data with Pandas, remove any null values, and one hot encode categoricals. Either Scikit-learn's feature encoders or Pandas `get_dummies` method can be used.
# * Split the data into train and test sets.
# * Log transform skewed features.
# * Scaling can be attempted, although it can be interesting to see how well regularization works without scaling features.
filepath = os.sep.join(data_path + ['Ames_Housing_Sales.csv'])
data = pd.read_csv(filepath, sep=',')
# Create a list of categorial data and one-hot encode. Pandas one-hot encoder (`get_dummies`) works well with data that is defined as a categorical.
# +
# Get a Pd.Series consisting of all the string categoricals
one_hot_encode_cols = data.dtypes[data.dtypes == np.object] # filtering by string categoricals
one_hot_encode_cols = one_hot_encode_cols.index.tolist() # list of categorical fields
# Here we see another way of one-hot-encoding:
# Encode these columns as categoricals so one hot encoding works on split data (if desired)
for col in one_hot_encode_cols:
data[col] = pd.Categorical(data[col])
# Do the one hot encoding
data = pd.get_dummies(data, columns=one_hot_encode_cols)
# -
# Next, split the data in train and test data sets.
# +
from sklearn.model_selection import train_test_split
train, test = train_test_split(data, test_size=0.3, random_state=42)
# -
# There are a number of columns that have skewed features--a log transformation can be applied to them. Note that this includes the `SalePrice`, our predictor. However, let's keep that one as is.
# Create a list of float colums to check for skewing
mask = data.dtypes == np.float
float_cols = data.columns[mask]
# +
skew_limit = 0.75
skew_vals = train[float_cols].skew()
skew_cols = (skew_vals
.sort_values(ascending=False)
.to_frame()
.rename(columns={0:'Skew'})
.query('abs(Skew) > {0}'.format(skew_limit)))
skew_cols
# -
# Transform all the columns where the skew is greater than 0.75, excluding "SalePrice".
# +
# OPTIONAL: Let's look at what happens to one of these features, when we apply np.log1p visually.
field = "BsmtFinSF1"
fig, (ax_before, ax_after) = plt.subplots(1, 2, figsize=(10, 5))
train[field].hist(ax=ax_before)
train[field].apply(np.log1p).hist(ax=ax_after)
ax_before.set(title='before np.log1p', ylabel='frequency', xlabel='value')
ax_after.set(title='after np.log1p', ylabel='frequency', xlabel='value')
fig.suptitle('Field "{}"'.format(field));
# a little bit better
# +
# Mute the setting wtih a copy warnings
pd.options.mode.chained_assignment = None
for col in skew_cols.index.tolist():
if col == "SalePrice":
continue
train[col] = np.log1p(train[col])
test[col] = test[col].apply(np.log1p) # same thing
# -
# Separate features from predictor.
# +
feature_cols = [x for x in train.columns if x != 'SalePrice']
X_train = train[feature_cols]
y_train = train['SalePrice']
X_test = test[feature_cols]
y_test = test['SalePrice']
# -
# ## Question 5
#
# * Write a function **`rmse`** that takes in truth and prediction values and returns the root-mean-squared error. Use sklearn's `mean_squared_error`.
#
# +
from sklearn.metrics import mean_squared_error
def rmse(ytrue, ypredicted):
return np.sqrt(mean_squared_error(ytrue, ypredicted))
# -
# * Fit a basic linear regression model
# * print the root-mean-squared error for this model
# * plot the predicted vs actual sale price based on the model.
# +
from sklearn.linear_model import LinearRegression
linearRegression = LinearRegression().fit(X_train, y_train)
linearRegression_rmse = rmse(y_test, linearRegression.predict(X_test))
print(linearRegression_rmse)
# +
f = plt.figure(figsize=(6,6))
ax = plt.axes()
ax.plot(y_test, linearRegression.predict(X_test),
marker='o', ls='', ms=3.0)
lim = (0, y_test.max())
ax.set(xlabel='Actual Price',
ylabel='Predicted Price',
xlim=lim,
ylim=lim,
title='Linear Regression Results');
# -
# ## Question 6
#
# Ridge regression uses L2 normalization to reduce the magnitude of the coefficients. This can be helpful in situations where there is high variance. The regularization functions in Scikit-learn each contain versions that have cross-validation built in.
#
# * Fit a regular (non-cross validated) Ridge model to a range of $\alpha$ values and plot the RMSE using the cross validated error function you created above.
# * Use $$[0.005, 0.05, 0.1, 0.3, 1, 3, 5, 10, 15, 30, 80]$$ as the range of alphas.
# * Then repeat the fitting of the Ridge models using the range of $\alpha$ values from the prior section. Compare the results.
# Now for the `RidgeCV` method. It's not possible to get the alpha values for the models that weren't selected, unfortunately. The resulting error values and $\alpha$ values are very similar to those obtained above.
# +
from sklearn.linear_model import RidgeCV
alphas = [0.005, 0.05, 0.1, 0.3, 1, 3, 5, 10, 15, 30, 80]
ridgeCV = RidgeCV(alphas=alphas,
cv=4).fit(X_train, y_train)
ridgeCV_rmse = rmse(y_test, ridgeCV.predict(X_test))
print(ridgeCV.alpha_, ridgeCV_rmse)
# -
# ## Question 7
#
#
# Much like the `RidgeCV` function, there is also a `LassoCV` function that uses an L1 regularization function and cross-validation. L1 regularization will selectively shrink some coefficients, effectively performing feature elimination.
#
# The `LassoCV` function does not allow the scoring function to be set. However, the custom error function (`rmse`) created above can be used to evaluate the error on the final model.
#
# Similarly, there is also an elastic net function with cross validation, `ElasticNetCV`, which is a combination of L2 and L1 regularization.
#
# * Fit a Lasso model using cross validation and determine the optimum value for $\alpha$ and the RMSE using the function created above. Note that the magnitude of $\alpha$ may be different from the Ridge model.
# * Repeat this with the Elastic net model.
# * Compare the results via table and/or plot.
#
# Use the following alphas:
# `[1e-5, 5e-5, 0.0001, 0.0005]`
# +
from sklearn.linear_model import LassoCV
alphas2 = np.array([1e-5, 5e-5, 0.0001, 0.0005])
lassoCV = LassoCV(alphas=alphas2,
max_iter=5e4,
cv=3).fit(X_train, y_train)
lassoCV_rmse = rmse(y_test, lassoCV.predict(X_test))
print(lassoCV.alpha_, lassoCV_rmse) # Lasso is slower
# -
# We can determine how many of these features remain non-zero.
print('Of {} coefficients, {} are non-zero with Lasso.'.format(len(lassoCV.coef_),
len(lassoCV.coef_.nonzero()[0])))
# + [markdown] run_control={"marked": true}
# Now try the elastic net, with the same alphas as in Lasso, and l1_ratios between 0.1 and 0.9
# +
from sklearn.linear_model import ElasticNetCV
l1_ratios = np.linspace(0.1, 0.9, 9)
elasticNetCV = ElasticNetCV(alphas=alphas2,
l1_ratio=l1_ratios,
max_iter=1e4).fit(X_train, y_train)
elasticNetCV_rmse = rmse(y_test, elasticNetCV.predict(X_test))
print(elasticNetCV.alpha_, elasticNetCV.l1_ratio_, elasticNetCV_rmse)
# -
# Comparing the RMSE calculation from all models is easiest in a table.
# +
rmse_vals = [linearRegression_rmse, ridgeCV_rmse, lassoCV_rmse, elasticNetCV_rmse]
labels = ['Linear', 'Ridge', 'Lasso', 'ElasticNet']
rmse_df = pd.Series(rmse_vals, index=labels).to_frame()
rmse_df.rename(columns={0: 'RMSE'}, inplace=1)
rmse_df
# -
# We can also make a plot of actual vs predicted housing prices as before.
# +
f = plt.figure(figsize=(6,6))
ax = plt.axes()
labels = ['Ridge', 'Lasso', 'ElasticNet']
models = [ridgeCV, lassoCV, elasticNetCV]
for mod, lab in zip(models, labels):
ax.plot(y_test, mod.predict(X_test),
marker='o', ls='', ms=3.0, label=lab)
leg = plt.legend(frameon=True)
leg.get_frame().set_edgecolor('black')
leg.get_frame().set_linewidth(1.0)
ax.set(xlabel='Actual Price',
ylabel='Predicted Price',
title='Linear Regression Results');
# -
# ## Question 8
#
# Let's explore Stochastic gradient descent in this exercise.
# Recall that Linear models in general are sensitive to scaling.
# However, SGD is *very* sensitive to scaling.
# Moreover, a high value of learning rate can cause the algorithm to diverge, whereas a too low value may take too long to converge.
#
# * Fit a stochastic gradient descent model without a regularization penalty (the relevant parameter is `penalty`).
# * Now fit stochastic gradient descent models with each of the three penalties (L2, L1, Elastic Net) using the parameter values determined by cross validation above.
# * Do not scale the data before fitting the model.
# * Compare the results to those obtained without using stochastic gradient descent.
# +
# Import SGDRegressor and prepare the parameters
from sklearn.linear_model import SGDRegressor
model_parameters_dict = {
'Linear': {'penalty': 'none'},
'Lasso': {'penalty': 'l2',
'alpha': lassoCV.alpha_},
'Ridge': {'penalty': 'l1',
'alpha': ridgeCV_rmse},
'ElasticNet': {'penalty': 'elasticnet',
'alpha': elasticNetCV.alpha_,
'l1_ratio': elasticNetCV.l1_ratio_}
}
new_rmses = {}
for modellabel, parameters in model_parameters_dict.items():
# following notation passes the dict items as arguments
SGD = SGDRegressor(**parameters)
SGD.fit(X_train, y_train)
new_rmses[modellabel] = rmse(y_test, SGD.predict(X_test))
rmse_df['RMSE-SGD'] = pd.Series(new_rmses)
rmse_df
# -
# Notice how high the error values are! The algorithm is diverging. This can be due to scaling and/or learning rate being too high. Let's adjust the learning rate and see what happens.
#
# * Pass in `eta0=1e-7` when creating the instance of `SGDClassifier`.
# * Re-compute the errors for all the penalties and compare.
# +
# Import SGDRegressor and prepare the parameters
from sklearn.linear_model import SGDRegressor
model_parameters_dict = {
'Linear': {'penalty': 'none'},
'Lasso': {'penalty': 'l2',
'alpha': lassoCV.alpha_},
'Ridge': {'penalty': 'l1',
'alpha': ridgeCV_rmse},
'ElasticNet': {'penalty': 'elasticnet',
'alpha': elasticNetCV.alpha_,
'l1_ratio': elasticNetCV.l1_ratio_}
}
new_rmses = {}
for modellabel, parameters in model_parameters_dict.items():
# following notation passes the dict items as arguments
SGD = SGDRegressor(eta0=1e-7, **parameters)
SGD.fit(X_train, y_train)
new_rmses[modellabel] = rmse(y_test, SGD.predict(X_test))
rmse_df['RMSE-SGD-learningrate'] = pd.Series(new_rmses)
rmse_df
# -
# Now let's scale our training data and try again.
#
# * Fit a `MinMaxScaler` to `X_train` create a variable `X_train_scaled`.
# * Using the scaler, transform `X_test` and create a variable `X_test_scaled`.
# * Apply the same versions of SGD to them and compare the results. Don't pass in a eta0 this time.
# +
# Please write the code with the scaled train and test data [Hint:use scaler.fit_transform]
| Intel-ML101-Class4/.ipynb_checkpoints/Week4_Regularization_and_Gradient_Descent_HW-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ___
# # Agenda
# - Numerical Optimization Techniques
# - Types of Optimization
# - Programming the Optimization
# - **Whirlwind Lecture Alert**
# - Entire classes cover these concepts in expanded form
# - But we can cover them in one lecture to get a good intuition!
# - And then you can look over this even more for better understanding.
# - If you feel confused after this lecture, that's okay. These are not easy the first time you see them. Keep going, you got this.
#
# <img src="PDF_slides/dont-be-nervous-you-got-this-meme.jpg" width="300">
#
#
#
# ___
#
# # Last Time
#
# |Description| Equations, Derivations, Hessian Calculations, and Miscellaneous|
# |-----------|--------|
# | Sigmoid Definition | $$ p(y^{(i)}=1\text{ | }\mathbf{x}^{(i)},\mathbf{w})=\frac{1}{1+\exp{(-\mathbf{w}^T \mathbf{x}^{(i)}})}$$ |
# | Log Likelihood | $$ l(\mathbf{w}) = \sum_i \left( y^{(i)} \ln [g(\mathbf{w}^T \mathbf{x}^{(i)})] + (1-y^{(i)}) (\ln [1 - g(\mathbf{w}^T \mathbf{x}^{(i)})]) \right) $$ |
# | Vectorized Gradient | $$gradient =\frac{1}{M}\sum_{i=1}^M (y^{(i)}-g(\mathbf{w}^T\mathbf{x}^{(i)}))\mathbf{x}^{(i)}$$ |
# | Regularization | $$ \mathbf{w} \leftarrow \mathbf{w} + \eta \left[\underbrace{\nabla l(\mathbf{w})_{old}}_{\text{old gradient}} - C \cdot 2\mathbf{w} \right]$$|
#
# ___
#
# <img src="PDF_slides/BtJXjJcCAAE7QOB.jpg" width="300">
#
# # More Advanced Optimization for Machine Learning
# From previous notebooks, we know that the logistic regression update equation is given by:
#
# $$ \underbrace{w_j}_{\text{new value}} \leftarrow \underbrace{w_j}_{\text{old value}} + \eta \underbrace{\left[\left(\sum_{i=1}^M (y^{(i)}-g(\mathbf{w}^T\mathbf{x}^{(i)}))x^{(i)}_j\right) - C \cdot 2w_j \right]}_{\nabla l(w)}$$
#
# Which can be made into more generic notation by denoting the objective function as $l(\mathbf{w})$ and the gradient calculation as $\nabla$:
# $$ \mathbf{w} \leftarrow \mathbf{w} + \eta \nabla l(\mathbf{w})$$
#
# One problem is that we still need to set the value of $\eta$, which can drastically change the performance of the optimization algorithm. If $\eta$ is too large, the algorithm might be unstable. If $\eta$ is too small, it might take a long time (i.e., many iterations) to converge.
#
#
# <img src="PDF_slides/batch.gif" width="400">
#
#
# We can solve this issue by performing a line search for the best value of $\eta$ along the direction of the gradient.
#
# $$ \mathbf{w} \leftarrow \mathbf{w} + \underbrace{\eta}_{\text{best step?}} \nabla l(\mathbf{w}) $$
#
# $$ \eta \leftarrow \arg\max_\eta l(\mathbf{w}+\eta\cdot\nabla l(\mathbf{w})) $$
#
# <img src="PDF_slides/line_search copy.gif" width="400">
#
# ## Optimizing Logistic Regression via Line Search
# +
from sklearn.datasets import load_iris
import numpy as np
from sklearn.metrics import accuracy_score
from scipy.special import expit
ds = load_iris()
X = ds.data
y = (ds.target>1).astype(np.int) # make problem binary
# +
# %%time
# from last time, our logistic regression algorithm is given by (including everything we previously had):
class BinaryLogisticRegression:
def __init__(self, eta, iterations=20, C=0.001):
self.eta = eta
self.iters = iterations
self.C = C
# internally we will store the weights as self.w_ to keep with sklearn conventions
def __str__(self):
if(hasattr(self,'w_')):
return 'Binary Logistic Regression Object with coefficients:\n'+ str(self.w_) # is we have trained the object
else:
return 'Untrained Binary Logistic Regression Object'
# convenience, private:
@staticmethod
def _add_bias(X):
return np.hstack((np.ones((X.shape[0],1)),X)) # add bias term
@staticmethod
def _sigmoid(theta):
# increase stability, redefine sigmoid operation
return expit(theta) #1/(1+np.exp(-theta))
# vectorized gradient calculation with regularization using L2 Norm
def _get_gradient(self,X,y):
ydiff = y-self.predict_proba(X,add_bias=False).ravel() # get y difference
gradient = np.mean(X * ydiff[:,np.newaxis], axis=0) # make ydiff a column vector and multiply through
gradient = gradient.reshape(self.w_.shape)
gradient[1:] += -2 * self.w_[1:] * self.C
return gradient
# public:
def predict_proba(self,X,add_bias=True):
# add bias term if requested
Xb = self._add_bias(X) if add_bias else X
return self._sigmoid(Xb @ self.w_) # return the probability y=1
def predict(self,X):
return (self.predict_proba(X)>0.5) #return the actual prediction
def fit(self, X, y):
Xb = self._add_bias(X) # add bias term
num_samples, num_features = Xb.shape
self.w_ = np.zeros((num_features,1)) # init weight vector to zeros
# for as many as the max iterations
for _ in range(self.iters):
gradient = self._get_gradient(Xb,y)
self.w_ += gradient*self.eta # multiply by learning rate
# add bacause maximizing
blr = BinaryLogisticRegression(eta=0.1,iterations=50,C=0.001)
blr.fit(X,y)
print(blr)
yhat = blr.predict(X)
print('Accuracy of: ',accuracy_score(y,yhat))
# +
# %%time
# and we can update this to use a line search along the gradient like this:
from scipy.optimize import minimize_scalar
import copy
from numpy import ma # (masked array) this has most numpy functions that work with NaN data.
class LineSearchLogisticRegression(BinaryLogisticRegression):
# define custom line search for problem
def __init__(self, line_iters=0.0, **kwds):
self.line_iters = line_iters
# but keep other keywords
super().__init__(**kwds) # call parent initializer
# this defines the function with the first input to be optimized
# therefore eta will be optimized, with all inputs constant
@staticmethod
def objective_function(eta,X,y,w,grad,C):
wnew = w - grad*eta
g = expit(X @ wnew)
return -np.sum(ma.log(g[y==1]))-ma.sum(np.log(1-g[y==0])) + C*sum(wnew**2)
def fit(self, X, y):
Xb = self._add_bias(X) # add bias term
num_samples, num_features = Xb.shape
self.w_ = np.zeros((num_features,1)) # init weight vector to zeros
# for as many as the max iterations
for _ in range(self.iters):
gradient = -self._get_gradient(Xb,y)
# minimization inopposite direction
# do line search in gradient direction, using scipy function
opts = {'maxiter':self.line_iters} # unclear exactly what this should be
res = minimize_scalar(self.objective_function, # objective function to optimize
bounds=(0,self.eta*10), #bounds to optimize
args=(Xb,y,self.w_,gradient,self.C), # additional argument for objective function
method='bounded', # bounded optimization for speed
options=opts) # set max iterations
eta = res.x # get optimal learning rate
self.w_ -= gradient*eta # set new function values
# subtract to minimize
lslr = LineSearchLogisticRegression(eta=1,
iterations=5,
line_iters=5,
C=0.001)
lslr.fit(X,y)
yhat = lslr.predict(X)
print(lslr)
print('Accuracy of: ',accuracy_score(y,yhat))
# -
# This performs well, but was not too much faster than previously (this is because $\eta$ was chosen well in the initial example).
# ___
# # Self Test
# How much computation (i.e., how many multiplies) are required for calculating the gradient of:
# $$ \left( \frac{1}{M}\left[\sum_{i=1}^M (y^{(i)}-g(\mathbf{w}^T\mathbf{x}^{(i)}))\mathbf{x}^{(i)}\right] - 2C\cdot \mathbf{w}\right) $$
#
# Where $M$ is the number of instance and $N$ is the number of elements in $\mathbf{w}$.
#
# - A: $ M\cdot N+1$
# - B: $ (M+1)\cdot N$
# - C: $ 2N $
# - D: $ 2N-M$
# _____
#
# # Stochastic Gradient Descent
# Sometimes the gradient calcualtion is too computational:
# $$ \mathbf{w} \leftarrow \mathbf{w} + \eta\left( \frac{1}{M}\left[\sum_{i=1}^M (y^{(i)}-g(\mathbf{w}^T\mathbf{x}^{(i)}))\mathbf{x}^{(i)}\right] - 2C\cdot \mathbf{w}\right) $$
#
# Instead, we can approximate the gradient using one instance, this is called stochastic gradient descent (SGD) because the steps can appear somewhat random.
# $$ \mathbf{w} \leftarrow \mathbf{w} + \eta \underbrace{\left((y^{(i)}-g(\mathbf{w}^T\mathbf{x}^{(i)}))\mathbf{x}^{(i)}-2C\cdot \mathbf{w}\right)}_{\text{approx. gradient}} \text{, where } i\in M$$
#
# <img src="PDF_slides/SGD.gif" width="400">
#
# Let's code up the SGD example.
# +
# %%time
class StochasticLogisticRegression(BinaryLogisticRegression):
# stochastic gradient calculation
def _get_gradient(self,X,y):
idx = int(np.random.rand()*len(y)) # grab random instance
ydiff = y[idx]-self.predict_proba(X[idx],add_bias=False) # get y difference (now scalar)
gradient = X[idx] * ydiff[:,np.newaxis] # make ydiff a column vector and multiply through
gradient = gradient.reshape(self.w_.shape)
gradient[1:] += -2 * self.w_[1:] * self.C
return gradient
slr = StochasticLogisticRegression(eta=0.05, iterations=100, C=0.001) # take a lot more steps!!
slr.fit(X,y)
yhat = slr.predict(X)
print(slr)
print('Accuracy of: ',accuracy_score(y,yhat))
# -
# ___
#
#
#
# <img src="PDF_slides/hessian_second.jpg" width="300">
#
#
#
#
# # Optimizing with Second Order Derivatives
# First, let's look at the one dimensional case when we have a function $l(w)$ where w is a scalar. The optimal value of w is given by:
#
# $$ w \leftarrow w - \underbrace{[\frac{\partial^2}{\partial w}l(w)]^{-1}}_{\text{inverse 2nd deriv}}\underbrace{\frac{\partial}{\partial w}l(w)}_{\text{derivative}} $$
#
# Note that if $l(w)$ is a quadratic function, this solution converges in a single step!
#
#
# **Aside: an example with the second derivative:**
# - Say $l(w)=2w^2+4w+5$, and we want to minimize the function. We have that:
# - $\frac{\partial}{\partial w}l(w)=4w+4$
# - $\frac{\partial^2}{\partial w}l(w)=4$
# - Therefore, if we choose $w_{start}=0$, we have:
# - $\frac{\partial}{\partial w}l(0)=4$
# - $\frac{\partial^2}{\partial w}l(0)=4$
# - So the update becomes
# - $w \leftarrow w_{start} - \frac{1}{4}4 = -1$
# - The solution is found in one step. This works for any initial value of $w_{start}$. Let's verify that the solution worked graphically.
# +
import numpy as np
from matplotlib import pyplot as plt
# %matplotlib inline
plt.style.use('ggplot')
w = np.linspace(-7,5,100)
l = 2*w**2+4*w+5
plt.plot(w,l)
plt.text(-1,2.5,'$\leftarrow$found minimum',fontsize=14)
# -
# ___
#
#
# # Newton's Update Method
# <img src="PDF_slides/newton.png" width="600">
#
# But how do we translate this over to objective funtions with more than one variable? We need a second derivative of a multivariate equation... enter, the hessian. Our new update is defined by Newton's method:
#
# $$ w \leftarrow w - \underbrace{[\frac{\partial^2}{\partial w}l(w)]^{-1}}_{\text{inverse 2nd deriv}}\underbrace{\frac{\partial}{\partial w}l(w)}_{\text{derivative}} $$
#
# such that, in multiple dimensions we can approximate the update as:
#
# $$ \mathbf{w} \leftarrow \mathbf{w} + \eta \cdot \underbrace{\mathbf{H}(\mathbf{w})^{-1}}_{\text{inverse Hessian}}\cdot\underbrace{\nabla l(\mathbf{w})}_{\text{gradient}}$$
#
# where the Hessian is defined as follows for any multivariate equation $l(\mathbf{w})$:
# $$ \nabla^2 l(\mathbf{w}) = \mathbf{H}(\mathbf{w}) $$
#
# $$ \mathbf{H}(\mathbf{w}) = \begin{bmatrix}
# \frac{\partial^2}{\partial w_1}l(\mathbf{w}) & \frac{\partial}{\partial w_1}\frac{\partial}{\partial w_2}l(\mathbf{w}) & \ldots & \frac{\partial}{\partial w_1}\frac{\partial}{\partial w_N}l(\mathbf{w}) \\
# \frac{\partial}{\partial w_2}\frac{\partial}{\partial w_1}l(\mathbf{w}) & \frac{\partial^2}{\partial w_2}l(\mathbf{w}) & \ldots & \frac{\partial}{\partial w_2}\frac{\partial}{\partial w_N}l(\mathbf{w}) \\
# & \vdots &\\
# \frac{\partial}{\partial w_N}\frac{\partial}{\partial w_1}l(\mathbf{w}) & \frac{\partial}{\partial w_N}\frac{\partial}{\partial w_2}l(\mathbf{w}) & \ldots & \frac{\partial^2}{\partial w_N}l(\mathbf{w}) \\
# \end{bmatrix}
# $$
#
# ____
#
# <img src="PDF_slides/spider_hessian.png" width="400">
# ____
# For logistic regression, we can calculate the formula for the $j^{th}$ and $k^{th}$ element of the Hessian as follows:
#
# $$ \mathbf{H}_{j,k}(\mathbf{w}) = \frac{\partial}{\partial w_k} \underbrace{\frac{\partial}{\partial w_j}l(\mathbf{w})}_{\text{first derivative}} $$
#
# But we already know the result of the $j^{th}$ partial derivative from our calculation of $\nabla l(\mathbf{w})$:
#
# $$ \frac{\partial}{\partial w_j}l(\mathbf{w}) = \sum_i \left(y^{(i)}-g(\mathbf{w}^T\cdot\mathbf{x}^{(i)})\right)x_j^{(i)} $$
#
# So we can plug this back into the equation to get:
#
# $$
# \begin{split}
# \mathbf{H}_{j,k}(\mathbf{w}) & = \frac{\partial}{\partial w_k}\sum_i \left(y^{(i)}-g(\mathbf{w}^T\cdot\mathbf{x}^{(i)})\right)x_j^{(i)} \\
# & = \underbrace{\sum_i \frac{\partial}{\partial w_k} y^{(i)}x_j^{(i)}}_{\text{no dependence on }k\text{, zero}} -\sum_i \frac{\partial}{\partial w_k}g(\mathbf{w}^T\cdot\mathbf{x}^{(i)})x_j^{(i)} \\
# & = -\sum_i x_j^{(i)}\underbrace{\frac{\partial}{\partial w_k}g(\mathbf{w}^T\cdot\mathbf{x}^{(i)})}_{\text{already know this as }g(1-g)x_k} \\
# & = -\sum_{i=1}^M \left[g(\mathbf{w}^T\mathbf{x}^{(i)})[1-g(\mathbf{w}^T\mathbf{x}^{(i)})]\right]\cdot{x_k}^{(i)}{x_j}^{(i)} \\
# \end{split}
# $$
#
#
#
#
# Therefore the Hessian for logistic regression becomes (adding in the regularization term also):
# $$ \mathbf{H}_{j,k}(\mathbf{w}) =\left( -\sum_{i=1}^M \underbrace{\left[g(\mathbf{w}^T\mathbf{x}^{(i)})[1-g(\mathbf{w}^T\mathbf{x}^{(i)})]\right]}_{\text{scalar value for each instance}}\cdot\underbrace{{x_k}^{(i)}{x_j}^{(i)}}_{i^{th}\text{ instance elements}} \right) + \underbrace{2\cdot C}_{\text{regularization}} $$
#
#
# This equation can be calcuated in a for loop, for each $j,k$ element in the Hessian and for each instance in the dataset, but this would be **slow** in python. To vectorize this operation, we need to have each operation be linear algebra, so that it can be run efficiently with numpy.
# ____
# ### Caluclating the Hessian for Logistic Regression using Linear Algebra
# First notice that the sum of each terms ${x_k}^{(i)}{x_j}^{(i)}$ that forms a matrix can be calculated as follows:
#
# $$
# \begin{bmatrix}
# \sum_{i=1}^M {x_1}^{(i)}{x_1}^{(i)} & \sum_{i=1}^M {x_1}^{(i)}{x_2}^{(i)} & \ldots & \sum_{i=1}^M {x_1}^{(i)}{x_N}^{(i)} \\
# \sum_{i=1}^M {x_2}^{(i)}{x_1}^{(i)} & \sum_{i=1}^M {x_2}^{(i)}{x_2}^{(i)} & \ldots & \sum_{i=1}^M {x_2}^{(i)}{x_N}^{(i)} \\
# & \vdots & \\
# \sum_{i=1}^M {x_N}^{(i)}{x_1}^{(i)} & \sum_{i=1}^M {x_N}^{(i)}{x_2}^{(i)} & \ldots & \sum_{i=1}^M {x_N}^{(i)}{x_N}^{(i)} \\ \\
# \end{bmatrix}
# %
# = \mathbf{X}^T \cdot\mathbf{I} \cdot\mathbf{X}
# $$
#
# where $\mathbf{I}$ is the identity matrix of size $M\text{x}M$. This can be seen in the following exploded view of the matrix operations:
#
# $$ \mathbf{X}^T \cdot\mathbf{I} \cdot\mathbf{X}=
# \begin{bmatrix}
# \uparrow & \uparrow & & \uparrow \\
# \mathbf{x}^{(1)} & \mathbf{x}^{(2)} & \ldots & \mathbf{x}^{(M)} \\
# \downarrow & \downarrow & & \downarrow \\
# \end{bmatrix}
# %
# \begin{bmatrix}
# 1 & 0 & \ldots & 0 \\
# 0 & 1 & \ldots & 0 \\
# & \vdots & & \\
# 0 & 0 & \ldots & 1 \\
# \end{bmatrix}
# %
# \begin{bmatrix}
# \leftarrow & \mathbf{x}^{(1)} & \rightarrow \\
# \leftarrow & \mathbf{x}^{(2)} & \rightarrow \\
# & \vdots &\\
# \leftarrow & \mathbf{x}^{(M)} & \rightarrow \\
# \end{bmatrix}
# %
# $$
#
# With this equation in mind, we can revisit the calcualtion of the Hessian and use matrix operations to define the needed multiplication in an exploded view of the operations:
#
# $$ \mathbf{H}[l(\mathbf{w})]=
# \begin{bmatrix}
# \uparrow & \uparrow & & \uparrow \\
# \mathbf{x}^{(1)} & \mathbf{x}^{(2)} & \ldots & \mathbf{x}^{(M)} \\
# \downarrow & \downarrow & & \downarrow \\
# \end{bmatrix}
# %
# \begin{bmatrix}
# g(\mathbf{w}^T\mathbf{x}^{(1)})[1-g(\mathbf{w}^T\mathbf{x}^{(1)})] & \ldots & 0 \\
# & \vdots & \\
# 0 & \ldots & g(\mathbf{w}^T\mathbf{x}^{(M)})[1-g(\mathbf{w}^T\mathbf{x}^{(M)})] \\
# \end{bmatrix}
# %
# \begin{bmatrix}
# \leftarrow & \mathbf{x}^{(1)} & \rightarrow \\
# \leftarrow & \mathbf{x}^{(2)} & \rightarrow \\
# & \vdots &\\
# \leftarrow & \mathbf{x}^{(M)} & \rightarrow \\
# \end{bmatrix}
# %
# $$
#
# ___
# Or, more succintly as follows (adding in the regularization term as well):
#
# $$ \mathbf{H}[l(\mathbf{w})] = \mathbf{X}^T \cdot \text{diag}\left[g(\mathbf{X}\cdot\mathbf{w})\odot(1-g(\mathbf{X}\cdot\mathbf{w}))\right]\cdot \mathbf{X} -2C$$
#
# ___
#
# Now we can place the Hessian derivation into the Newton Update Equation, like this:
#
# $$ \mathbf{w} \leftarrow \mathbf{w} + \eta \cdot \underbrace{\mathbf{H}[l(\mathbf{w})]^{-1}}_{\text{inverse Hessian}}\cdot\underbrace{\nabla l(\mathbf{w})}_{\text{gradient}}$$
#
# Adding in the exact equations for the Hessian and gradient, we can finally get:
#
# $$ \mathbf{w} \leftarrow \mathbf{w} + \eta \cdot \underbrace{\left[\mathbf{X}^T \cdot \text{diag}\left[g(\mathbf{X}\cdot\mathbf{w})\odot(1-g(\mathbf{X}\cdot\mathbf{w}))\right] \cdot \mathbf{X} -2C \right]^{-1} }_{\text{inverse Hessian}} \cdot \underbrace{\mathbf{X}\odot y_{diff}}_{\text{gradient}}$$
#
#
# You can see the full derivation of the Hessian in my hand written notes here also:
# - https://raw.githubusercontent.com/eclarson/MachineLearningNotebooks/master/PDF_Slides/HessianCalculation.pdf
#
#
#
# So let's code this up using numpy:
# +
# %%time
from numpy.linalg import pinv
class HessianBinaryLogisticRegression(BinaryLogisticRegression):
# just overwrite gradient function
def _get_gradient(self,X,y):
g = self.predict_proba(X,add_bias=False).ravel() # get sigmoid value for all classes
hessian = X.T @ np.diag(g*(1-g)) @ X - 2 * self.C # calculate the hessian
ydiff = y-g # get y difference
gradient = np.sum(X * ydiff[:,np.newaxis], axis=0) # make ydiff a column vector and multiply through
gradient = gradient.reshape(self.w_.shape)
gradient[1:] += -2 * self.w_[1:] * self.C
return pinv(hessian) @ gradient
hlr = HessianBinaryLogisticRegression(eta=1.0,
iterations=4,
C=0.001) # note that we need only a few iterations here
hlr.fit(X,y)
yhat = hlr.predict(X)
print(hlr)
print('Accuracy of: ',accuracy_score(y,yhat))
# -
# ___
# ### Can we still do better? Problems With the Hessian:
# Quadratic isn’t always a great assumption:
# - highly dependent on starting point
# - jumps can get really random!
# - near saddle points, inverse hessian is unstable
# - hessian not always invertible… or invertible with enough numerical precision
#
# The Hessian can sometimes be ill formed for these problems and can also be highly computational. Thus, we prefer to approximate the Hessian, and approximate its inverse to better control the steps we make and directions we use.
#
# <img src="PDF_slides/gru_hessian.jpg" width="400">
#
# ____
#
# # Quasi-Newton Methods
# In general:
# - Approximate the Hessian with something numerically sound and efficiently invertible
# - Back off to gradient descent when the approximate hessian is not stable
# - Try to create an approximation with as many properties of the Hessian as possible, like being symmetric and positive semi-definite
# - A popular approach: Rank One Hessian Approximation
# - An even more popular appraoch: Rank Two, with Broyden-Fletcher-Goldfarb-Shanno (BFGS)
#
#
# ### Rank One Hessian Approximation
# Let's work our way up to using BFGS by first looking at one quasi-newton method, the rank one Hessian approximation. **Note, I only want you to get an intuition for this process. There is no requirenemtn to understand the derivation completely.**
# Essentially, we want to update the Hessian with an approximation that is easily invertible and based on stable gradient calculations. We can define the approximate Hessian for each iteration, $\mathbf{H}_k$. To start as simple as possible, we will assume the Hessian can be approximated with one vector. Let's start off with a few other assumptions. We wil develop some equations that characterize a family of solutions. Within this family, we will only give one popular example for the rank one family solution and one popular example for the rank two family solution.
#
# ___
# One property of the hessian is called the Secant equation, which relates the change in input to the change in the derivative. The Secant Equation (exact for quadratic functions) is:
# $$ \underbrace{\mathbf{H}_{k+1}}_{\text{approx. Hessian}} \cdot\underbrace{(\mathbf{w}_{k+1} - \mathbf{w}_k)}_{\text{Change in }w} = \underbrace{\nabla l(\mathbf{w}_{k+1}) - \nabla l(\mathbf{w}_k)}_{\text{Change in gradient}}$$
#
# or, using intermediate variables for the differences:
# $$ \mathbf{H}_{k+1} \mathbf{s}_k = \mathbf{v}_k $$
#
# where $ \mathbf{s} = (\mathbf{w}_{k+1} - \mathbf{w}_k) $ and $ \mathbf{v} = (\nabla l(\mathbf{w}_{k+1}) - \nabla l(\mathbf{w}_k)) $, the difference in the gradient. If we enforce this relationship, we can find the hessian, assuming that the current location is approximated well by a quadratic (making the Secant a good assumption also). We also want the Hessian to be symmetric and not too far away from its initial value (for stable optimization).
# ___
#
# For optimizing, we would like to be able to update the Hessian at $\mathbf{w}_{k+1}$ from our previous guess, $\mathbf{H}_k$ at $\mathbf{w}_{k}$, and have the update be easy to calculate. Therefore, we can choose the update of the Hessian to be approximated by the rank one update (one vector). Since the Hessian is the second partial derivative, a starting approximation might be the gradient difference we already defined such that $\mathbf{H} \approx \mathbf{v}\cdot\mathbf{v}^T$ which would form a matrix of the differences of each partial deriviative in the gradient. In practice, we need a vector that is slightly less constrained, such that
# $$ \mathbf{H}_{k+1}=\mathbf{H}_k+\alpha_k\mathbf{u}\cdot\mathbf{u}^T $$
# Where $\mathbf{u}$ and $\alpha_k$ can be anything we want.
#
# Substituting back into the secant formula:
# $$ (\mathbf{H}_k +\alpha_k\mathbf{u}_k\cdot\mathbf{u}_k^T)\mathbf{s}_{k} = \mathbf{v}_{k} $$
#
#
# Many solutions exist for this and they are referred to as a family of rank one Hessian approximations. One solution of this equation (there are many solutions) is to use one that simplifies nicely. For example we can choose the following:
# $$ \mathbf{u}_k=\mathbf{v}_{k}-\mathbf{H}_k \mathbf{s}_{k} \text{ and } \alpha_k=\frac{1}{(\mathbf{v}_{k}-\mathbf{H}_k \mathbf{s}_{k})\mathbf{s}_{k}}=\frac{1}{\mathbf{u}_{k}^T\mathbf{s}_{k}}$$
#
# ___
# and combining this with our initial $\mathbf{H}_{k+1}$ formula:
# $$ \mathbf{H}_{k+1}=\mathbf{H}_k- \frac{\mathbf{u}_k\mathbf{u}_k^T}{\mathbf{u}_k^T\mathbf{s}_{k}} $$
#
# This gives an update for the Hessian, which we can use in our optimization formula. However, we need to define the vectors using the secant equation assumptions, such that $ \mathbf{v}_k $ and $ \mathbf{s}_k $ are the difference in gradients and weights as defined, respectively.
#
# **Now for the power of this method.** We can now assume that the inverse of the Hessian can be optimized and formulate similar equations for its update, based upon the previous inverse. Therefore, we need the inverse of $(\mathbf{H}_k+\mathbf{v}\cdot\mathbf{v}^T)^{-1}$, which luckily has a closed form solution according to the Sherman-Morrison formula:
#
# $$ (\mathbf{A}+\mathbf{v}\cdot\mathbf{v}^T)^{-1} = \mathbf{A}^{-1} - \frac{\mathbf{A}^{-1} \mathbf{v} \mathbf{v}^T\mathbf{A}^{-1}}{1+\mathbf{v}^T \mathbf{A}^{-1} \mathbf{v}} $$
# ___
#
# Now the optimization can be described as a rank one approximation of the Hessian. Placing it all together, we can get the following:
#
# |Description| Equations, Derivations, Hessian Calculations, and Miscellaneous |
# |-----------|--------|
# | **Definitions with Rank 1 Approximation** | |
# |$$ \mathbf{w} \leftarrow \mathbf{w} + \eta \cdot \underbrace{\mathbf{H}[l(\mathbf{w})]^{-1}}_{\text{inverse Hessian}}\cdot\underbrace{\nabla l(\mathbf{w})}_{\text{gradient}}$$ | |
# |1. Initial Approx. Hessian for $k=0$ is identity matrix| $$\mathbf{H}_0=\mathbf{I}$$|
# |2. Find update direction, $\mathbf{p}_k$ | $$ \mathbf{p}_k = -\mathbf{H}_k^{-1} \nabla l(\mathbf{w}_k) $$|
# |3. Update $\mathbf{w}$|$$\mathbf{w}_{k+1}\leftarrow \mathbf{w}_k + \eta \cdot \mathbf{p}_k $$|
# |4. Save scaled direction ($\mathbf{w}_{k+1}-\mathbf{w}_k$)| $$\mathbf{s}_k=\eta \cdot \mathbf{p}_k$$ |
# |5a. Approximate change in derivative | $$\mathbf{v}_k = \nabla l(\mathbf{w}_{k+1}) - \nabla l(\mathbf{w}_k) $$|
# | 5b. Define $\mathbf{u}$ from above: | $$\mathbf{u}_k=\mathbf{v}_k-\mathbf{H}_k\mathbf{s}_k$$|
# |6. Redefine approx Hessian update| $$\mathbf{H}_{k+1}=\mathbf{H}_k+\underbrace{\frac{\mathbf{u}_k \mathbf{u}_k^T}{\mathbf{u}_k^T \mathbf{s}_k}}_{\text{approx. Hessian}} $$ |
# |7. Approx. Inverse $\mathbf{H}_{k+1}^{-1}$ via <NAME>| $$ \mathbf{H}_{k+1}^{-1} = \mathbf{H}_{k}^{-1} - \frac{\mathbf{H}_k^{-1} \mathbf{u}_k \mathbf{u}_k^T\mathbf{H}_k^{-1}}{1+\mathbf{u}_k^T \mathbf{H}_k^{-1} \mathbf{u}_k} $$ |
# | 8. Repeat starting at step 2| $$ k = k+1 $$|
#
#
#
# ___
# ### Rank Two Hessian Approximation: BFGS
#
#
# Although the rank one approximation is a good performer, it can be improved by adding some additional criteria to the Hessian approximation. In this case, we assume that the $\mathbf{H}_k$ needs to also be positive semi-definite, which helps with numerical stability. One of the most popular quasi-Newton methods that does this is known as Broyden-Fletcher-Goldfarb-Shanno (BFGS).
# - https://en.wikipedia.org/wiki/Broyden–Fletcher–Goldfarb–Shanno_algorithm
#
# <img src="PDF_slides/bfgs_meme.png" width="300">
#
# In this formulation we add an additional vector matrix addition to the update equation that ensure the resulting matrix is positive semidefinite:
# $$ \mathbf{H}_{k+1}=\mathbf{H}_k+\alpha_k\mathbf{u}_k\cdot\mathbf{u}_k^T + \beta_k\mathbf{z}_k\cdot\mathbf{z}_k^T$$
#
# The derivation is intuitively similar to the previous rank one approximation. Again, there are many potential solutions, referred to as a family of ranke two solutions. However, it becomes easier to obtain simple solutions for $\mathbf{u}$ and $\mathbf{z}$. THe BFGS solution takes the following form:
# $$ \mathbf{u}_k = \mathbf{v}_k \text{ and } \mathbf{z}_k=\mathbf{H}_k \mathbf{s}_k $$
#
# After solving for the $\alpha_k$ and $\beta_k$ coefficients, we get the update equation as:
# $$ \mathbf{H}_{k+1}=\mathbf{H}_k+\underbrace{\frac{\mathbf{v}_k \mathbf{v}_k^T}{\mathbf{v}_k^T \mathbf{s}_k}}_{\text{previous}} -\underbrace{\frac{\mathbf{H}_k \mathbf{s}_k \mathbf{s}_k^T \mathbf{H}_k}{\mathbf{s}_k^T \mathbf{H}_k \mathbf{s}_k}}_{\text{new}} $$
#
# The complete formulation can replace steps from the previous rank update as follows:
#
# |Description| Equations, Derivations, Hessian Calculations, and Miscellaneous |
# |-----------|--------|
# | **Alternative Definitions with Rank 2 (BFGS)** | |
# | 6. Redefine approx Hessian| $$\mathbf{H}_{k+1}=\mathbf{H}_k+\frac{\mathbf{v}_k \mathbf{v}_k^T}{\mathbf{v}_k^T \mathbf{s}_k} -\frac{\mathbf{H}_k \mathbf{s}_k \mathbf{s}_k^T \mathbf{H}_k}{\mathbf{s}_k^T \mathbf{H}_k \mathbf{s}_k} $$ |
# |7. Approximate Inverse $\mathbf{H}_{k+1}^{-1}$ via <NAME>| $$ \mathbf{H}_{k+1}^{-1} = \mathbf{H}_{k}^{-1} + \frac{(\mathbf{s}_k^T \mathbf{v}_k+\mathbf{H}_{k}^{-1})(\mathbf{s}_k \mathbf{s}_k^T)}{(\mathbf{s}_k^T \mathbf{v}_k)^2}-\frac{\mathbf{H}_{k}^{-1} \mathbf{v}_k \mathbf{s}_k^T+\mathbf{s}_k \mathbf{v}_k^T\mathbf{H}_{k}^{-1}}{\mathbf{s}_k^T \mathbf{v}_k} $$|
#
# ___
# We won't explicitly program the BFGS algorithm--instead we can take advantage of scipy's calculations to do it for us. For using this algorithm, we need to define the objective function and the gradient explicitly for another program to calculate.
# Recall that Logistic regression uses the following objective function:
#
# $$ l(w) = \left(\sum_i y^{(i)} \ln g(\mathbf{x}^{(i)}) + (1-y^{(i)})\ln[1-g(\mathbf{x}^{(i)})]\right) - C \cdot \sum_j w_j^2 $$
# +
# %%time
# for this, we won't perform our own BFGS implementation
# (it takes a fair amount of code and understanding, which we haven't setup yet)
# luckily for us, scipy has its own BFGS implementation:
from scipy.optimize import fmin_bfgs # maybe the most common bfgs algorithm in the world
from numpy import ma
class BFGSBinaryLogisticRegression(BinaryLogisticRegression):
@staticmethod
def objective_function(w,X,y,C):
g = expit(X @ w)
# invert this because scipy minimizes, but we derived all formulas for maximzing
return -np.sum(ma.log(g[y==1]))-np.sum(ma.log(1-g[y==0])) + C*sum(w**2)
#-np.sum(y*np.log(g)+(1-y)*np.log(1-g))
@staticmethod
def objective_gradient(w,X,y,C):
g = expit(X @ w)
ydiff = y-g # get y difference
gradient = np.mean(X * ydiff[:,np.newaxis], axis=0)
gradient = gradient.reshape(w.shape)
gradient[1:] += -2 * w[1:] * C
return -gradient
# just overwrite fit function
def fit(self, X, y):
Xb = self._add_bias(X) # add bias term
num_samples, num_features = Xb.shape
self.w_ = fmin_bfgs(self.objective_function, # what to optimize
np.zeros((num_features,1)), # starting point
fprime=self.objective_gradient, # gradient function
args=(Xb,y,self.C), # extra args for gradient and objective function
gtol=1e-03, # stopping criteria for gradient, |v_k|
maxiter=self.iters, # stopping criteria iterations
disp=False)
self.w_ = self.w_.reshape((num_features,1))
bfgslr = BFGSBinaryLogisticRegression(_,iterations=2,C=0.001) # note that we need only a few iterations here
bfgslr.fit(X,y)
yhat = bfgslr.predict(X)
print(bfgslr)
print('Accuracy of: ',accuracy_score(y,yhat))
# +
#str(bfgslr.eta)
# -
# ### BFGS and Newton's Method for Multiclass Logistic Regression
# Now let's add BFGS and the actual Hessian to non-binary classification. As before, we will use one-versus-all.
# +
# allow for the user to specify the algorithm they want to sovle the binary case
class MultiClassLogisticRegression:
def __init__(self, eta, iterations=20,
C=0.0001,
solver=BFGSBinaryLogisticRegression):
self.eta = eta
self.iters = iterations
self.C = C
self.solver = solver
self.classifiers_ = []
# internally we will store the weights as self.w_ to keep with sklearn conventions
def __str__(self):
if(hasattr(self,'w_')):
return 'MultiClass Logistic Regression Object with coefficients:\n'+ str(self.w_) # is we have trained the object
else:
return 'Untrained MultiClass Logistic Regression Object'
def fit(self,X,y):
num_samples, num_features = X.shape
self.unique_ = np.sort(np.unique(y)) # get each unique class value
num_unique_classes = len(self.unique_)
self.classifiers_ = []
for i,yval in enumerate(self.unique_): # for each unique value
y_binary = np.array(y==yval).astype(int) # create a binary problem
# train the binary classifier for this class
hblr = self.solver(eta=self.eta,iterations=self.iters,C=self.C)
hblr.fit(X,y_binary)
# add the trained classifier to the list
self.classifiers_.append(hblr)
# save all the weights into one matrix, separate column for each class
self.w_ = np.hstack([x.w_ for x in self.classifiers_]).T
def predict_proba(self,X):
probs = []
for hblr in self.classifiers_:
probs.append(hblr.predict_proba(X).reshape((len(X),1))) # get probability for each classifier
return np.hstack(probs) # make into single matrix
def predict(self,X):
return np.argmax(self.predict_proba(X),axis=1) # take argmax along row
# +
from sklearn.preprocessing import StandardScaler
ds = load_iris()
X = ds.data
#X = StandardScaler().fit(X).transform(X)
y_not_binary = ds.target # note problem is NOT binary anymore, there are three classes!
# +
# %%time
lr = MultiClassLogisticRegression(eta=1,
iterations=10,
C=0.01,
solver=BFGSBinaryLogisticRegression
)
lr.fit(X,y_not_binary)
print(lr)
yhat = lr.predict(X)
print('Accuracy of: ',accuracy_score(y_not_binary,yhat))
# +
# %%time
lr = MultiClassLogisticRegression(eta=1,
iterations=10,
C=0.001,
solver=HessianBinaryLogisticRegression
)
lr.fit(X,y_not_binary)
print(lr)
yhat = lr.predict(X)
print('Accuracy of: ',accuracy_score(y_not_binary,yhat))
# +
# %%time
# how do we compare now to sklearn?
from sklearn.linear_model import LogisticRegression
lr_sk = LogisticRegression(solver='lbfgs',n_jobs=1,
multi_class='ovr', C = 1/0.001,
penalty='l2',
max_iter=50) # all params default
# note that sklearn is optimized for using the liblinear library with logistic regression
# ...and its faster than our implementation here
lr_sk.fit(X, y_not_binary) # no need to add bias term, sklearn does it internally!!
print(lr_sk.coef_)
yhat = lr_sk.predict(X)
print('Accuracy of: ',accuracy_score(y_not_binary,yhat))
# +
# %%time
# actually, we aren't quite as good as the lib linear implementation
# how do we compare now to sklearn?
from sklearn.linear_model import LogisticRegression
lr_sk = LogisticRegression(solver='liblinear',n_jobs=1,
multi_class='ovr', C = 1/0.001,
penalty='l2',max_iter=100)
lr_sk.fit(X,y_not_binary) # no need to add bias term, sklearn does it internally!!
print(lr_sk.coef_)
yhat = lr_sk.predict(X)
print('Accuracy of: ',accuracy_score(y_not_binary,yhat))
# -
# Liblinear is a great toolkit for linear modeling (from national Taiwan University) and the paper can be found here:
# - https://www.csie.ntu.edu.tw/~cjlin/papers/liblinear.pdf
#
# Actually, this solves a slightly different problem (known as the 'dual' formulation because its a linear SVM) to make it extremely fast. So this is actually not a fair comparison to ours.
# +
# %%time
# its still faster! Can we fix it with parallelization?
from joblib import Parallel, delayed
class ParallelMultiClassLogisticRegression(MultiClassLogisticRegression):
@staticmethod
def par_logistic(yval,eta,iters,C,X,y,solver):
y_binary = y==yval # create a binary problem
# train the binary classifier for this class
hblr = solver(eta=eta,iterations=iters,C=C)
hblr.fit(X,y_binary)
return hblr
def fit(self,X,y):
num_samples, num_features = X.shape
self.unique_ = np.sort(np.unique(y)) # get each unique class value
num_unique_classes = len(self.unique_)
backend = 'threading' # can also try 'multiprocessing'
self.classifiers_ = Parallel(n_jobs=-1,backend=backend)(
delayed(self.par_logistic)(yval,self.eta,self.iters,self.C,X,y,self.solver) for yval in self.unique_)
# save all the weights into one matrix, separate column for each class
self.w_ = np.hstack([x.w_ for x in self.classifiers_]).T
plr = ParallelMultiClassLogisticRegression(eta=1,iterations=10,C=0.001,solver=HessianBinaryLogisticRegression)
plr.fit(X,y_not_binary)
print(plr)
yhat = plr.predict(X)
print('Accuracy of: ',accuracy_score(y_not_binary,yhat))
# -
# ___
# Please note that the overhead of parallelization is not worth it for this problem!!
#
# **When would it make sense???**
# ___
#
#
# <img src="PDF_slides/mark_scooter.png" width="300">
#
#
# # Extended Logistic Regression Example
#
# In this example we will explore methods of using logistic regression in scikit-learn. A basic understanding of scikit-learn is required to complete this notebook, but we start very basic. Note also that there are more efficient methods of separating testing and training data, but we will leave that for a later lecture.
#
# First let's load a dataset and prepare it for analysis. We will use pandas to load in data, and then prepare it for classification. We will be using the titanic dataset (a very modest sized data set of about 1000 instances). The imputation methods used here are discussed in a previous notebook.
#
# Steps:
# - Load data, impute
# - One hot encode and normalize data
# - Separate into training and testing sets
# - Explore best hyper parameter, C
#
#
# ## Load Titanic Data and Pre-process
#
# +
import pandas as pd
import numpy as np
df = pd.read_csv('data/titanic.csv') # read in the csv file
# 1. Remove attributes that just arent useful for us
del df['PassengerId']
del df['Name']
del df['Cabin']
del df['Ticket']
# 2. Impute some missing values, grouped by their Pclass and SibSp numbers
df_grouped = df.groupby(by=['Pclass','SibSp'])
# # now use this grouping to fill the data set in each group, then transform back
# fill in the numeric values
df_imputed = df_grouped.transform(lambda grp: grp.fillna(grp.median()))
# fill in the categorical values
df_imputed[['Sex','Embarked']] = df_grouped[['Sex','Embarked']].apply(lambda grp: grp.fillna(grp.mode()))
# fillin the grouped variables from original data frame
df_imputed[['Pclass','SibSp']] = df[['Pclass','SibSp']]
# 4. drop rows that still had missing values after grouped imputation
df_imputed.dropna(inplace=True)
# 5. Rearrange the columns
df_imputed = df_imputed[['Survived','Age','Sex','Parch','SibSp','Pclass','Fare','Embarked']]
df_imputed.info()
# +
# perform one-hot encoding of the categorical data "embarked"
tmp_df = pd.get_dummies(df_imputed.Embarked,prefix='Embarked')
df_imputed = pd.concat((df_imputed,tmp_df),axis=1) # add back into the dataframe
# replace the current Sex atribute with something slightly more intuitive and readable
df_imputed['IsMale'] = df_imputed.Sex=='male'
df_imputed.IsMale = df_imputed.IsMale.astype(np.int)
# Now let's clean up the dataset
if 'Sex' in df_imputed:
del df_imputed['Sex'] # if 'Sex' column still exists, delete it (as we created an ismale column)
if 'Embarked' in df_imputed:
del df_imputed['Embarked'] # get reid of the original category as it is now one-hot encoded
# Finally, let's create a new variable based on the number of family members
# traveling with the passenger
# notice that this new column did not exist before this line of code--we use the pandas
# syntax to add it in
df_imputed['FamilySize'] = df_imputed.Parch + df_imputed.SibSp
df_imputed.info()
# -
# ## Training and Testing Split
# For training and testing purposes, let's gather the data we have and grab 80% of the instances for training and the remaining 20% for testing. Moreover, let's repeat this process of separating the testing and training data three times. We will use the hold out cross validation method built into scikit-learn.
# +
from sklearn.model_selection import ShuffleSplit
# we want to predict the X and y data as follows:
if 'Survived' in df_imputed:
y = df_imputed['Survived'].values # get the labels we want
del df_imputed['Survived'] # get rid of the class label
norm_features = ['Age','Fare' ]
df_imputed[norm_features] = (df_imputed[norm_features]-df_imputed[norm_features].mean()) / df_imputed[norm_features].std()
X = df_imputed.to_numpy() # use everything else to predict!
## X and y are now numpy matrices, by calling 'values' on the pandas data frames we
# have converted them into simple matrices to use with scikit learn
# to use the cross validation object in scikit learn, we need to grab an instance
# of the object and set it up. This object will be able to split our data into
# training and testing splits
num_cv_iterations = 3
num_instances = len(y)
cv_object = ShuffleSplit(
n_splits=num_cv_iterations,
test_size = 0.2)
print(cv_object)
# +
# run logistic regression and vary some parameters
from sklearn import metrics as mt
# first we create a reusable logisitic regression object
# here we can setup the object with different learning parameters and constants
lr_clf = HessianBinaryLogisticRegression(eta=0.1,iterations=10) # get object
# now we can use the cv_object that we setup before to iterate through the
# different training and testing sets. Each time we will reuse the logisitic regression
# object, but it gets trained on different data each time we use it.
iter_num=0
# the indices are the rows used for training and testing in each iteration
for train_indices, test_indices in cv_object.split(X,y):
# I will create new variables here so that it is more obvious what
# the code is doing (you can compact this syntax and avoid duplicating memory,
# but it makes this code less readable)
X_train = X[train_indices]
y_train = y[train_indices]
X_test = X[test_indices]
y_test = y[test_indices]
# train the reusable logisitc regression model on the training data
lr_clf.fit(X_train,y_train) # train object
y_hat = lr_clf.predict(X_test) # get test set precitions
# now let's get the accuracy and confusion matrix for this iterations of training/testing
acc = mt.accuracy_score(y_test,y_hat)
conf = mt.confusion_matrix(y_test,y_hat)
print("====Iteration",iter_num," ====")
print("accuracy", acc )
print("confusion matrix\n",conf)
iter_num+=1
# Also note that every time you run the above code
# it randomly creates a new training and testing set,
# so accuracy will be different each time
# +
# this does the exact same thing as the above block of code, but with shorter syntax
for iter_num, (train_indices, test_indices) in enumerate(cv_object.split(X,y)):
lr_clf.fit(X[train_indices],y[train_indices]) # train object
y_hat = lr_clf.predict(X[test_indices]) # get test set precitions
# print the accuracy and confusion matrix
print("====Iteration",iter_num," ====")
print("accuracy", mt.accuracy_score(y[test_indices],y_hat))
print("confusion matrix\n",mt.confusion_matrix(y[test_indices],y_hat))
# -
# ## Interactive Example, adjusting C
# +
from ipywidgets import widgets as wd
num_cv_iterations = 10
num_instances = len(y)
cv_object = ShuffleSplit(n_splits=num_cv_iterations,
test_size = 0.5)
def lr_explor(cost):
print('Running')
lr_clf = HessianBinaryLogisticRegression(eta=0.1,iterations=10,
C=float(cost)) # get object
acc = []
for iter_num, (train_indices, test_indices) in enumerate(cv_object.split(X,y)):
lr_clf.fit(X[train_indices],y[train_indices]) # train object
y_hat = lr_clf.predict(X[test_indices]) # get test set predictions
acc.append(mt.accuracy_score(y[test_indices],y_hat))
acc = np.array(acc)
print(acc.mean(),'+-',2.7*acc.std())
wd.interact(lr_explor,cost=list(np.logspace(-4,1,15)),__manual=True)
# -
# ## Exhaustive Search for C, then Visualize with Boxplots
# +
# %%time
# alternatively, we can also graph out the values using boxplots
num_cv_iterations = 20
num_instances = len(y)
cv_object = ShuffleSplit(n_splits=num_cv_iterations,
test_size = 0.5)
def lr_explor(cost):
lr_clf = BFGSBinaryLogisticRegression(eta=0.1,iterations=10,
C=float(cost)) # get object
acc = []
for iter_num, (train_indices, test_indices) in enumerate(cv_object.split(X,y)):
lr_clf.fit(X[train_indices],y[train_indices]) # train object
y_hat = lr_clf.predict(X[test_indices]) # get test set predictions
acc.append(mt.accuracy_score(y[test_indices],y_hat))
acc = np.array(acc)
return acc
costs = np.logspace(-5,1,20)
accs = []
for c in costs:
accs.append(lr_explor(c))
# +
# now show a boxplot of the data across c
from matplotlib import pyplot as plt
# %matplotlib inline
plt.boxplot(accs)
plt.xticks(range(1,len(costs)+1),['%.4f'%(c) for c in costs],rotation='vertical')
plt.xlabel('C')
plt.ylabel('validation accuracy')
plt.show()
# -
#
# ___
# # Next Time: Neural Networks
# ___
#
# In this notebook you learned:
# - Formulation of Logistic regression with different optimization strategies
# - Line Search
# - Mini-batch
# - Stochastic Gradient
# - Newton's Approach using Hessian
# - Quasi Newton's Method
# - Use Exhaustive Searches for Finding "C"
# - And Training/Testing Splits
| 06. Optimization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Disable the IPython pager
# https://gist.github.com/minrk/7715212
from IPython.core import page
page.page = print
# # 06 - Property Based Testing (Bonus)
#
# SW testing joke
# > A QA engineer walks into a bar. Orders a beer. Orders 0 beers. Orders 99999999999 beers. Orders a lizard. Orders -1 beers. Orders a ueicbksjdhd.
#
# > First real customer walks in and asks where the bathroom is. The bar bursts into flames, killing everyone.
#
# From [hypothesis documentation](https://hypothesis.readthedocs.io/en/latest):
# > [Hypothesis](https://hypothesis.works) is a Python library for creating unit tests which are simpler to write and more powerful when run, finding edge cases in your code you wouldn’t have thought to look for. It is stable, powerful and easy to add to any existing test suite.
import pytest
import hypothesis
# The Hypothesis library provides
# * strategies for generating input data based on given criteria
# * tools for integration those into property based tests
#
# Sligthly modified [definition](https://hypothesis.readthedocs.io/en/latest/):
#
# | Example based tests | Property based tests |
# | :------------------ | :------------------- |
# | 1. Set up *some example* data | 1. for **all data** matching some specification |
# | 2. Perform some operations on the data | 2. Perform some operations on the data |
# | 3. Assert something *specific* about the result | 3. Assert something **generic** about the result |
#
# ## Test data generators
from hypothesis.strategies import floats, integers, text
f = floats()
[f.example() for _ in range(10)]
print(text().example())
# See more examples under [What you can generate and how](https://hypothesis.readthedocs.io/en/latest/data.html)
# ## Example use-case
# Let's assume we need to write a function that returns the 2nd largest value in a list of integers
import math
def second_largest(values):
return values[-2]
# Is this a good implementation?
#
# To find out, we can try a couple of example tests
second_largest([1, 2, 3, 4, 5])
second_largest([1, 2, 3, 4])
# Seems ok? Except it isn't.
#
# These tests are too "weak". Let's use Hypothesis
# %pycat test_stats.py
# !pytest test_stats.py
# ## Pandas and Numpy types
# Read more on [Hypothesis for the Scientific Stack](https://hypothesis.readthedocs.io/en/latest/numpy.html)
# ## See also
# * [Quick Start Guide](https://hypothesis.readthedocs.io/en/latest/quickstart.html)
# * [Introductory articles](https://hypothesis.works/articles/)
| solutions/06_property_based_testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
def f(N):
s = 0
for i in range(N):
for j in range(N):
for k in range(N):
s += i + j + k
return s
# %time f(100)
# -
# %time f(1000)
| all_repository/julia_lab/py3_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def miniMaxSum(arr):
no_max_arr = arr.copy()
no_min_arr = arr.copy()
no_max_arr.remove(max(arr))
no_min_arr.remove(min(arr))
print(str(sum(no_max_arr))+' '+str(sum(no_min_arr)))
a = [1,1,1,1,-3]
miniMaxSum(a)
| HackerRank/Problem Solving/min_max.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/gxflove307/R-CNN_object_detection/blob/main/R_CNN_Implement.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="1FM14BVjC_wl" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fd57b550-9ebb-4626-8872-ead9fbbd4d52"
from google.colab import drive
drive.mount('/content/drive/')
# + id="scmkBKT_f0K1"
import os,cv2,keras,csv
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
import random
#import tensorflow.compat.v1 as tf
#tf.disable_v2_behavior()
# + id="-s0F9K1pyFM_" colab={"base_uri": "https://localhost:8080/", "height": 89} outputId="a7b2f96d-2bfd-44e9-c6de-980bf9bf62c1"
# !pip install pyprind
import pyprind
# + id="uPKzhf9Tgh1U"
PathImg = '/content/drive/My Drive/My MCM Practicum/Insight-MVT_Annotation_Test_part/' #图片目录
PathCsv ='/content/drive/My Drive/XML/CSV/'#注释目录
PathTest ='/content/drive/My Drive/My MCM Practicum/Prediction/'#预测目录
PathSave ='/content/drive/My Drive/My MCM Practicum/Prediction_pict/'#保存目录
# + id="IF9byRZneDTF" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="33f4be64-eb31-4404-f670-7ecb1195d77b"
#python画图保存 以左上角为原点
#a = random.randint(0,60)
for e,i in enumerate(os.listdir(PathTest)):
#if e ==a:
filename = i.split(".")[0]+".jpg"
# print(a)
#print(filename)
img = cv2.imread(os.path.join(PathImg,filename))
#print(img.shape)
df = pd.read_csv(os.path.join(PathTest,i),sep=',')
df.reset_index(inplace=True)
#plt.imshow(img)
gtvalues=[]
#print(df)
for index,row in df.iterrows():
x1 = int(row[0])
y1 = int(row[1])
x2 = int(row[2])
y2 = int(row[3])
gtvalues.append({"x1":x1,"x2":x2,"y1":y1,"y2":y2})
cv2.rectangle(img,(x1,y1),(x2,y2),(255,0,0), 2)
plt.figure()
plt.imshow(img)
plt.savefig(os.path.join(PathSave,filename))
# + id="cimlxEiUoAXm" colab={"base_uri": "https://localhost:8080/", "height": 581} outputId="c91bdb4c-6436-4382-dbba-42a3f7b7878e"
#python画图 以左上角为原点
a = random.randint(0,60)
for e,i in enumerate(os.listdir(PathCsv)):
#if e ==a:
if i=='MVI_20012-img00745.csv':
filename = i.split(".")[0]+".jpg"
print(a)
print(filename)
img = cv2.imread(os.path.join(PathImg,filename))
#print(img.shape)
df = pd.read_csv(os.path.join(PathCsv,i),sep=',')
df.reset_index(inplace=True)
plt.imshow(img)
gtvalues=[]
print(df)
for index,row in df.iterrows():
x1 = int(row[0])
y1 = int(row[1])
x2 = int(row[2])
y2 = int(row[3])
gtvalues.append({"x1":x1,"x2":x2,"y1":y1,"y2":y2})
cv2.rectangle(img,(x1,y1),(x2,y2),(255,0,0), 2)
plt.figure()
plt.imshow(img)
#plt.savefig(os.path.join(PathSave,filename))
break
# + [markdown] id="ngQxZ3sVQNXH"
# 选择性搜索是区域规划算法的方法,被用于目标检测。它具有fast 和 high recall 的特性。算法原理:基于颜色、纹理、大小和形状等,将image分层划分为诸多相似区域。
# + id="2hotZcWzrWzu"
cv2.setUseOptimized(True);#打开优化代码
# create Selective Search Segmentation Object using default parameters 使用默认参数创建选择性搜索分割对象
ss = cv2.ximgproc.segmentation.createSelectiveSearchSegmentation()
# + id="ge4Yqx8QWnr7"
#剔除候选框中面积小于2000px,长宽比大于2的box
def get_usefulbox(bb):
bb=pd.DataFrame(bb,columns=['x','y','w','h'])
bb.drop(bb[(bb.w * bb.h < 2000) | (bb.w/bb.h > 2) | (bb.h/bb.w > 2)].index,inplace=True)
bb.reset_index()
bb = bb.values
return bb
# + id="Ec2WIBi1iJew"
# %%time
im = cv2.imread(os.path.join(PathImg,filename))
ss.setBaseImage(im)#set input image on which we will run segmentation 设置我们将运行分割的输入图像,
ss.switchToSelectiveSearchQuality()#fast but low recall Selective Search method 选择速度快的分隔方法 还可以使用SearchQuality()基于质量不过运行慢
rects = ss.process()#run selective search segmentation on input image 处理图片
rects = get_usefulbox(rects)
imOut = im.copy()# create a copy of original image 创建一个原图片的备份
for i, rect in (enumerate(rects)):# itereate over all the region proposals 迭代显示所有区域
if i < 10: #看20个框框就够了
x, y, w, h = rect
cv2.rectangle(imOut, (x, y), (x+w, y+h), (255, 0, 0), 2, cv2.LINE_AA)
# plt.figure()
plt.imshow(imOut)
# + id="eNhPExHUiWz9"
train_images=[]
train_labels=[]
# + id="M6MnOJFGiXkm"
#iou(Intersection over union)交并比 可用来验证算法画框的准确性(此处用来确定区域选择的生成框是否可取 iou越大说明属性区域选择生成框与备注框重合部分越大 则可取为训练集)
def get_iou(bb1, bb2): #bb1原图片的备注框 bb2选择搜索生成框
assert bb1['x1'] < bb1['x2']
assert bb1['y1'] < bb1['y2']
assert bb2['x1'] < bb2['x2']
assert bb2['y1'] < bb2['y2']
x_left = max(bb1['x1'], bb2['x1'])
y_top = max(bb1['y1'], bb2['y1'])
x_right = min(bb1['x2'], bb2['x2'])
y_bottom = min(bb1['y2'], bb2['y2'])
if x_right < x_left or y_bottom < y_top:
return 0.0
intersection_area = (x_right - x_left) * (y_bottom - y_top)#取两个框的重叠部分的面积
bb1_area = (bb1['x2'] - bb1['x1']) * (bb1['y2'] - bb1['y1'])#原图备注框面积
bb2_area = (bb2['x2'] - bb2['x1']) * (bb2['y2'] - bb2['y1'])#选择搜索生成框
iou = intersection_area / float(bb1_area + bb2_area - intersection_area)#两框重叠部分占两框总面积的比例
assert iou >= 0.0
assert iou <= 1.0
return iou
# + id="4BUFMKbDifET"
ss = cv2.ximgproc.segmentation.createSelectiveSearchSegmentation()# create Selective Search Segmentation Object using default parameters
# + id="Rqsx9E88iicg" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="8ee02271-505f-4a70-a9f4-32b526852a97"
#pbar1 = pyprind.ProgBar(len(os.listdir(PathCsv)),title='Trainset generation')
# %%time
for e,i in enumerate(os.listdir(PathCsv)):
try:
if e == 0:#选择图片数量
filename = i.split(".")[0]+".jpg"
print(e,filename)
image = cv2.imread(os.path.join(PathImg,filename))
df = pd.read_csv(os.path.join(PathCsv,i))
df.reset_index(inplace=True)
gtvalues=[]
for index,row in df.iterrows():
x1 = int(row[0])
y1 = int(row[1])
x2 = int(row[2])
y2 = int(row[3])
gtvalues.append({"x1":x1,"x2":x2,"y1":y1,"y2":y2})
ss.setBaseImage(image)#set input image on which we will run segmentation
ss.switchToSelectiveSearchFast()#fast but low recall Selective Search method
ssresults = ss.process()#run selective search segmentation on input image
ssresults = get_usefulbox(ssresults)
imout = image.copy()# create a copy of original image
#pbar1.update()
counter = 0
falsecounter = 0
flag = 0
fflag = 0
bflag = 0
#pbar2 = pyprind.ProgBar(len(ssresults),title='ssresults images')
for e,result in enumerate(ssresults):
#pbar2.update()
if e < 2000 and flag == 0:
for gtval in gtvalues:
x,y,w,h = result
iou = get_iou(gtval,{"x1":x,"x2":x+w,"y1":y,"y2":y+h})
if counter < 30:
if iou > 0.7:#重合面积大于70%
timage = imout[y:y+h,x:x+w]#合并图片
resized = cv2.resize(timage, (224,224), interpolation = cv2.INTER_AREA)
train_images.append(resized)#将选择划分的图片中与原备注框重合率大于70%的作为训练集并添加标签为1
train_labels.append(1)
counter += 1
#print("counter=%d" %counter)
else :
fflag =1
if falsecounter <30:
if iou < 0.3:
timage = imout[y:y+h,x:x+w]
resized = cv2.resize(timage, (224,224), interpolation = cv2.INTER_AREA)
train_images.append(resized)#将选择划分的图片中与原备注框重合率小于30%的作为训练集并添加标签为0
train_labels.append(0)
falsecounter += 1
#print("falsecounter=%d" %falsecounter)
else :
bflag = 1
if fflag == 1 and bflag == 1:
print("inside")
flag = 1
except Exception as e:
print(e)
print("error in "+filename)
continue
# + id="_WF5qGzbil34"
X_new = np.array(train_images)
y_new = np.array(train_labels)
# + id="g_mG9cZtdZQ7"
np.save("/content/drive/My Drive/XML/550_X_new.npy",np.array(train_images))
# + id="_H1bXiE0iKnX"
np.save("/content/drive/My Drive/XML/550_y_new.npy",np.array(train_labels))
# + id="OQqF-07xegD9"
X_new = np.load("/content/drive/My Drive/My MCM Practicum/400_X_new.npy")
y_new = np.load("/content/drive/My Drive/My MCM Practicum/400_y_new.npy")
# + id="klvA8KV4i08l" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="a44557f9-655d-4a32-ae88-bfecde71c46d"
print(X_new.shape)
print(y_new.shape)
# + id="jZ3Y4iJMi3PQ"
from keras.layers import Dense
from keras import Model
from keras import optimizers
from keras.preprocessing.image import ImageDataGenerator
from keras.applications.vgg16 import VGG16
# + id="fI1V7DJhi9va"
#transfer learning
vggmodel = VGG16(weights='imagenet', include_top=True)
#vggmodel.summary()
# + id="gVvJLVygjAUq" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="02a6b71f-b261-45b4-c4fc-70fd121905ac"
#freezing the first 15 layers of the model 冻结模型的前19层
for layers in (vggmodel.layers)[:15]:
print(layers)
layers.trainable = False
# + id="PbKTtcuzjCJ3"
#取模型的最后2层
X= vggmodel.layers[-2].output
# + id="elZRPbMhjEqF"
#adding a 2 unit softmax dense layer as we have just 2 classes to predict 添加两个softmax层 分类为2
predictions = Dense(2, activation="softmax")(X)
# + id="QIvIcaLpjG7H" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="0f79ce24-65e2-48df-b43f-78b468fffcb2"
model_final = Model(input = vggmodel.input, output = predictions)
# + id="zKsGSWJEjIwN"
from keras.optimizers import Adam
opt = Adam(lr=0.0001)
# + id="WYVsNDz7jKbw"
model_final.compile(loss = tf.keras.losses.categorical_crossentropy, optimizer = opt, metrics=["accuracy"])
# + id="RsdZb_KdjMa8" colab={"base_uri": "https://localhost:8080/", "height": 924} outputId="f63adaaf-2ed3-4b24-9787-c001e4bd2aab"
#模型创建完毕
model_final.summary()
# + id="SAqJcPF_jOMg"
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelBinarizer
# + id="QkhMLuQ5jPtv"
#one-hot encode
class MyLabelBinarizer(LabelBinarizer):
def transform(self, y):
Y = super().transform(y)
if self.y_type_ == 'binary':
return np.hstack((Y, 1-Y))
else:
return Y
def inverse_transform(self, Y, threshold=None):
if self.y_type_ == 'binary':
return super().inverse_transform(Y[:, 0], threshold)
else:
return super().inverse_transform(Y, threshold)
# + id="tiWCiZkhjSmy"
lenc = MyLabelBinarizer()
Y = lenc.fit_transform(y_new)
# + id="dYhjmlK1jXIE"
#split trin and test set
X_train, X_test , y_train, y_test = train_test_split(X_new,Y,test_size=0.10)
# + id="aRB8OsCejash" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f99a24ec-7715-486f-f607-eae9af4751c1"
print(X_train.shape,X_test.shape,y_train.shape,y_test.shape)
# + id="n-4E9Pt_jc7y"
#We will do some augmentation on the dataset like horizontal flip, vertical flip and rotation to increase the dataset. 水平 垂直反转增加数据集
trdata = ImageDataGenerator(horizontal_flip=True, vertical_flip=True, rotation_range=90)
traindata = trdata.flow(x=X_train, y=y_train)
tsdata = ImageDataGenerator(horizontal_flip=True, vertical_flip=True, rotation_range=90)
testdata = tsdata.flow(x=X_test, y=y_test)
# + id="reXiuOftjfAp"
from keras.callbacks import ModelCheckpoint, EarlyStopping
# + id="r8Sg3m34jhLT"
checkpoint = ModelCheckpoint("rcnn_vgg16_1.h5", monitor='val_loss', verbose=1, save_best_only=True, save_weights_only=False, mode='auto', period=1)
early = EarlyStopping(monitor='val_loss', min_delta=0, patience=100, verbose=1, mode='auto')
# + id="cB4Gvu47jkRw" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="0eb414d8-3f05-4091-a4b0-412b6b420b3d"
# %%time
#hist = model_final.fit(x=X_train, y=y_train,batch_size=100, epochs= 1000,validation_data=(X_test,y_test),callbacks=[checkpoint,early])
hist = model_final.fit_generator(generator= traindata, steps_per_epoch= 10, epochs= 1000, validation_data= testdata, validation_steps=2, callbacks=[checkpoint,early])
# + id="krz2NDNdLh3r" colab={"base_uri": "https://localhost:8080/", "height": 304} outputId="e7e11064-8e4e-48fe-97c6-69d8968c0184"
# !/opt/bin/nvidia-smi
# + id="gZ-p_GiJjmgQ" colab={"base_uri": "https://localhost:8080/", "height": 313} outputId="76888152-cc92-47a6-aa32-6709933d90a4"
import matplotlib.pyplot as plt
# plt.plot(hist.history["acc"])
# plt.plot(hist.history['val_acc'])
plt.plot(hist.history['loss'])
plt.plot(hist.history['val_loss'])
plt.title("model loss")
plt.ylabel("Loss")
plt.xlabel("Epoch")
plt.legend(["Loss","Validation Loss"])
plt.show()
plt.savefig('chart loss.png')
# + id="2k-rEVkQjoyx" colab={"base_uri": "https://localhost:8080/", "height": 287} outputId="487c5420-cda7-4a83-b52d-132c7f3e7191"
im = X_test[127]
plt.imshow(im)
img = np.expand_dims(im, axis=0)
out= model_final.predict(img)
if out[0][0] > out[0][1]:
print("vehicle")
else:
print("not vehicle")
# + id="CEgIeIbessFK"
model_final.save('/content/drive/My Drive/My MCM Practicum/400_model.h5')
# + id="2DFaB5G_O2uh"
import os,cv2,keras
from keras.models import load_model
# + id="wjZIsKxuO9hw"
model_final = load_model("/content/drive/My Drive/My MCM Practicum/400_model.h5")
# + id="uaDIc7PEs3RI"
ss = cv2.ximgproc.segmentation.createSelectiveSearchSegmentation() # create Selective Search Segmentation Object using default parameters
# + id="Y5PS1jMqjrd1" colab={"base_uri": "https://localhost:8080/", "height": 307} outputId="845b6445-6b15-4db1-eefc-c5431a74169d"
# %%time
z=0
a = random.randint(0,60)
for e,i in enumerate(os.listdir(PathImg)):
if e == a:
#if i =='MVI_40761_img00166.jpg':
print(i)
#z += 1
img = cv2.imread(os.path.join(PathImg,i))
ss.setBaseImage(img)
ss.switchToSelectiveSearchFast()
ssresults = ss.process()
ssresults = get_usefulbox(ssresults)
imout = img.copy()
for e,result in enumerate(ssresults):
if e < 2000:
x,y,w,h = result
timage = imout[y:y+h,x:x+w]
resized = cv2.resize(timage, (224,224), interpolation = cv2.INTER_AREA)
img = np.expand_dims(resized, axis=0)
out= model_final.predict(img)
if out[0][0] > 0.5:
cv2.rectangle(imout, (x, y), (x+w, y+h), (0, 255, 0),2)
z+=1
#print(out[0][0])
plt.figure()
plt.imshow(imout)
print(z)
# + id="F2tlp7ZQv8rb" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="5d191f77-7ef6-4ce6-e341-10f12481a6ff"
len(os.listdir(PathImg))
#print(PathImg)
# + id="oXePWi7FBQ0N" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="6f2d6a50-0654-4775-9a70-c6e4b3712d03"
# %%time
a = random.randint(0,60)
#pbar = pyprind.ProgBar((2400-len(os.listdir(PathTest))),title='Prediction generation')
Imgfiles = os.listdir(PathImg)
Imgfiles.sort(key=lambda x:int(x[4:8]))
for count,name in enumerate(Imgfiles):
try:
filename = name.split(".")[0]+".csv"
#if not(os.path.exists(os.path.join(PathTest,filename))):
if count == a:
print(name)
#z += 1
#pbar.update(item_id=count)
Annopositions = pd.read_csv(os.path.join(PathCsv,filename),sep=',')
Annopositions.reset_index(inplace=True)
Annovalues=[]
getpositions=[]
getrightpos=[]
delects=[]
for index,row in Annopositions.iterrows():
x1 = int(row[0])
y1 = int(row[1])
x2 = int(row[2])
y2 = int(row[3])
Annovalues.append({"x1":x1,"x2":x2,"y1":y1,"y2":y2})
img = cv2.imread(os.path.join(PathImg,name))
ss.setBaseImage(img)
ss.switchToSelectiveSearchFast()
ssresults = ss.process()
ssresults = get_usefulbox(ssresults)
imout = img.copy()
for area,result in enumerate(ssresults):
if area < 2000:
x,y,w,h = result
timage = imout[y:y+h,x:x+w]
resized = cv2.resize(timage, (224,224), interpolation = cv2.INTER_AREA)
img = np.expand_dims(resized, axis=0)
out= model_final.predict(img)
if out[0][0] > 0.7:
getpositions.append({"x1":x,"x2":x+w,"y1":y,"y2":y+h}) #将所得到的框的坐标放入getpositions中
#cv2.rectangle(imout, (x, y), (x+w, y+h), (0, 255, 0),
#去重,如果所得带坐标框与Annotation的iou大于0.5 就保存
for q in range(len(getpositions)):
for p in range(len(Annovalues)):
iou = get_iou(getpositions[q],Annovalues[p])
if iou > 0.5:
getrightpos.append(getpositions[q])
if iou == 0:
getrightpos.append(getpositions[q])
#去重,如果所得带坐标框之间iou大于0.2 就删除其中一个框
for i in range(len(getrightpos)):
for j in range(i+1,len(getrightpos)):
iou = get_iou(getrightpos[i],getrightpos[j])
if iou >0.2:
delects.append(j)
delects=list(set(delects))
getrightpos = [getrightpos[i] for i in range(len(getrightpos)) if i not in delects]
#在原图的copy中画框
#将框保存在CSV中
frame_num = []
index = []
Resident_data = open(PathTest+name[0:-4]+'.csv', 'w')
csvwriter = csv.writer(Resident_data)
frame_num.append(len(getrightpos))
csvwriter.writerow(frame_num)
frame_num = []
for getpos in getrightpos:
index.append(getpos['x1'])
index.append(getpos['y1'])
index.append(getpos['x2'])
index.append(getpos['y2'])
csvwriter.writerow(index)
index=[]
Resident_data.close()
except Exception as e:
print(e)
print("error in "+filename)
continue
#print(pbar)
| R_CNN_Implement.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # DeadTree Train Notebook
# +
import torch
import pytorch_lightning as pl
from pytorch_lightning import Trainer
from pytorch_lightning.callbacks import ModelCheckpoint, EarlyStopping
from pytorch_lightning.loggers.wandb import WandbLogger
import hydra
from omegaconf import DictConfig
from deadtrees.network.segmodel import SemSegment
from deadtrees.data.deadtreedata import DeadtreesDataModule
from deadtrees.visualization.helper import show
# -
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
print(f"NVIDIA Cuda available: {torch.cuda.is_available()}")
print(f"PyTorch Version: {torch.__version__}")
print(f"PyTorch Lightning Version: {pl.__version__}")
# ## Instantiate DataModule
datamodule = DeadtreesDataModule(
"../data/dataset/train/",
pattern = "train-balanced-000*.tar",
pattern_extra = ["train-negativesamples-000*.tar", "train-randomsamples-000*.tar"],
batch_size_extra = [1, 7],
train_dataloader_conf = {'batch_size': 16, 'num_workers': 2},
val_dataloader_conf = {'batch_size': 16, 'num_workers': 2},
test_dataloader_conf = {'batch_size': 16, 'num_workers': 2},
)
datamodule.setup(in_channels=4, classes=2)
# ## Instantiate the Model
# +
train_config = DictConfig(
dict(learning_rate = 0.0003,
run_test = False,
)
)
network_config = DictConfig(
dict(
# model definitions
architecture = 'unet',
encoder_name = 'resnet34',
encoder_depth = 5,
encoder_weights = "imagenet",
# data specific settings
classes = 2,
in_channels = 4,
)
)
model = SemSegment(train_config, network_config)
model.summarize(max_depth=1);
# -
# ## Instantiate a Trainer
# +
# define some callbacks
model_checkpoint = ModelCheckpoint(
monitor = "val/total_loss",
mode = "min",
save_top_k = 1,
dirpath = "checkpoints/",
filename = "{epoch:02d}"
)
early_stopping = EarlyStopping(
monitor = "val/total_loss",
mode = "min",
patience = 10,
)
# define the Weights&Biases logger
wandb_logger = WandbLogger(
project = "deadtrees",
offline = False,
job_type = "train",
group = "",
save_dir = ".",
)
# -
EPOCHS = 10
trainer = Trainer(
gpus=1,
min_epochs=1,
max_epochs=EPOCHS,
precision=16,
progress_bar_refresh_rate=10,
terminate_on_nan=True,
callbacks=[model_checkpoint, early_stopping],
logger=[wandb_logger],
)
# ## Some experiments
#
# > **NOTE:**
# > Currently not working - skip ?!?
# + active=""
# # Run learning rate finder
# lr_finder = trainer.tuner.lr_find(model, min_lr=1e-07, max_lr=0.01, num_training=4, datamodule=datamodule)
# lr_finder.results
# + active=""
# # Plot with
# fig = lr_finder.plot(suggest=True)
# fig.show()
# + active=""
# # Pick point based on plot, or get suggestion
# new_lr = lr_finder.suggestion()
#
# # update hparams of the model
# model.hparams.learning_rate = new_lr
# -
# ## Train the model
trainer.fit(model=model, datamodule=datamodule)
# ## Test the model
trainer.test()
| notebooks/01-Training.ipynb |
# ##### Copyright 2020 Google LLC.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# # alphametic
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/google/or-tools/blob/master/examples/notebook/contrib/alphametic.ipynb"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/colab_32px.png"/>Run in Google Colab</a>
# </td>
# <td>
# <a href="https://github.com/google/or-tools/blob/master/examples/contrib/alphametic.py"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/github_32px.png"/>View source on GitHub</a>
# </td>
# </table>
# First, you must install [ortools](https://pypi.org/project/ortools/) package in this colab.
# !pip install ortools
# +
# Copyright 2010 <NAME> <EMAIL>
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""
Generic alphametic solver in Google CP Solver.
This is a generic alphametic solver.
Usage:
python alphametic.py
-> solves SEND+MORE=MONEY in base 10
python alphametic.py 'SEND+MOST=MONEY' 11
-> solver SEND+MOST=MONEY in base 11
python alphametic.py TEST <base>
-> solve some test problems in base <base>
(defined in test_problems())
Assumptions:
- we only solves problems of the form
NUMBER<1>+NUMBER<2>...+NUMBER<N-1> = NUMBER<N>
i.e. the last number is the sum
- the only nonletter characters are: +, =, \d (which are splitted upon)
Compare with the following model:
* Zinc: http://www.hakank.org/minizinc/alphametic.zinc
This model was created by <NAME> (<EMAIL>)
Also see my other Google CP Solver models:
http://www.hakank.org/google_or_tools/
"""
import sys
import re
from ortools.constraint_solver import pywrapcp
# Create the solver.
solver = pywrapcp.Solver("Send most money")
# data
print("\nproblem:", problem_str)
# convert to array.
problem = re.split("[\s+=]", problem_str)
p_len = len(problem)
print("base:", base)
# create the lookup table: list of (digit : ix)
a = sorted(set("".join(problem)))
n = len(a)
lookup = dict(list(zip(a, list(range(n)))))
# length of each number
lens = list(map(len, problem))
#
# declare variables
#
# the digits
x = [solver.IntVar(0, base - 1, "x[%i]" % i) for i in range(n)]
# the sums of each number (e.g. the three numbers SEND, MORE, MONEY)
sums = [solver.IntVar(1, 10**(lens[i]) - 1) for i in range(p_len)]
#
# constraints
#
solver.Add(solver.AllDifferent(x))
ix = 0
for prob in problem:
this_len = len(prob)
# sum all the digits with proper exponents to a number
solver.Add(
sums[ix] == solver.Sum([(base**i) * x[lookup[prob[this_len - i - 1]]]
for i in range(this_len)[::-1]]))
# leading digits must be > 0
solver.Add(x[lookup[prob[0]]] > 0)
ix += 1
# the last number is the sum of the previous numbers
solver.Add(solver.Sum([sums[i] for i in range(p_len - 1)]) == sums[-1])
#
# solution and search
#
solution = solver.Assignment()
solution.Add(x)
solution.Add(sums)
db = solver.Phase(x, solver.CHOOSE_FIRST_UNBOUND, solver.ASSIGN_MIN_VALUE)
solver.NewSearch(db)
num_solutions = 0
while solver.NextSolution():
num_solutions += 1
print("\nsolution #%i" % num_solutions)
for i in range(n):
print(a[i], "=", x[i].Value())
print()
for prob in problem:
for p in prob:
print(p, end=" ")
print()
print()
for prob in problem:
for p in prob:
print(x[lookup[p]].Value(), end=" ")
print()
print("sums:", [sums[i].Value() for i in range(p_len)])
print()
print("\nnum_solutions:", num_solutions)
print("failures:", solver.Failures())
print("branches:", solver.Branches())
print("WallTime:", solver.WallTime())
def test_problems(base=10):
problems = [
"SEND+MORE=MONEY", "SEND+MOST=MONEY", "VINGT+CINQ+CINQ=TRENTE",
"EIN+EIN+EIN+EIN=VIER", "DONALD+GERALD=ROBERT",
"SATURN+URANUS+NEPTUNE+PLUTO+PLANETS", "WRONG+WRONG=RIGHT"
]
for p in problems:
main(p, base)
problem = "SEND+MORE=MONEY"
base = 10
| examples/notebook/contrib/alphametic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import json
import subprocess
os.environ["CUDA_VISIBLE_DEVICES"] = '-1' ### run on CPU
import tensorflow as tf
print(tf.__version__)
if tf.__version__[0] == '1':
tf.compat.v1.enable_eager_execution()
import numpy as np
import pandas as pd
import pysam
import matplotlib.pyplot as plt
from cooltools.lib.numutils import set_diag
from basenji import dataset, dna_io, seqnn
# -
# ## Load trained model
# +
### load params, specify model ###
model_dir = './'
params_file = model_dir+'params.json'
model_file = model_dir+'model_best.h5'
with open(params_file) as params_open:
params = json.load(params_open)
params_model = params['model']
params_train = params['train']
seqnn_model = seqnn.SeqNN(params_model)
# -
### restore model ###
# note: run %%bash get_model.sh
# if you have not already downloaded the model
seqnn_model.restore(model_file)
print('successfully loaded')
# +
### names of targets ###
data_dir = './data/'
hic_targets = pd.read_csv(data_dir+'/targets.txt',sep='\t')
hic_file_dict_num = dict(zip(hic_targets['index'].values, hic_targets['file'].values) )
hic_file_dict = dict(zip(hic_targets['identifier'].values, hic_targets['file'].values) )
hic_num_to_name_dict = dict(zip(hic_targets['index'].values, hic_targets['identifier'].values) )
# read data parameters
data_stats_file = '%s/statistics.json' % data_dir
with open(data_stats_file) as data_stats_open:
data_stats = json.load(data_stats_open)
seq_length = data_stats['seq_length']
target_length = data_stats['target_length']
hic_diags = data_stats['diagonal_offset']
target_crop = data_stats['crop_bp'] // data_stats['pool_width']
target_length1 = data_stats['seq_length'] // data_stats['pool_width']
# -
# ## Make predictions for saved tfrecords
# +
### load data ###
# note: run %%bash get_data.sh
# if you have not already downloaded the data
sequences = pd.read_csv(data_dir+'sequences.bed', sep='\t', names=['chr','start','stop','type'])
sequences_test = sequences.iloc[sequences['type'].values=='test']
sequences_test.reset_index(inplace=True, drop=True)
test_data = dataset.SeqDataset(data_dir, 'test', batch_size=8)
# test_targets is a float array with shape
# [#regions, #pixels, target #target datasets]
# representing log(obs/exp)data, where #pixels
# corresponds to the number of entries in the flattened
# upper-triangular representation of the matrix
# test_inputs are 1-hot encoded arrays with shape
# [#regions, 2^20 bp, 4 nucleotides datasets]
test_inputs, test_targets = test_data.numpy(return_inputs=True, return_outputs=True)
# +
### for converting from flattened upper-triangluar vector to symmetric matrix ###
def from_upper_triu(vector_repr, matrix_len, num_diags):
z = np.zeros((matrix_len,matrix_len))
triu_tup = np.triu_indices(matrix_len,num_diags)
z[triu_tup] = vector_repr
for i in range(-num_diags+1,num_diags):
set_diag(z, np.nan, i)
return z + z.T
target_length1_cropped = target_length1 - 2*target_crop
print('flattened representation length:', target_length)
print('symmetrix matrix size:', '('+str(target_length1_cropped)+','+str(target_length1_cropped)+')')
# -
fig2_examples = [ 'chr12:115163136-116211712',
'chr11:75429888-76478464',
'chr15:63281152-64329728' ]
fig2_inds = []
for seq in fig2_examples:
print(seq)
chrm,start,stop = seq.split(':')[0], seq.split(':')[1].split('-')[0], seq.split(':')[1].split('-')[1]
test_ind = np.where( (sequences_test['chr'].values== chrm) *
(sequences_test['start'].values== int(start))*
(sequences_test['stop'].values== int(stop )) )[0][0]
fig2_inds.append(test_ind)
fig2_inds
# +
### make predictions and plot the three examples above ###
target_index = 0 # HFF
for test_index in fig2_inds:
chrm, seq_start, seq_end = sequences_test.iloc[test_index][0:3]
myseq_str = chrm+':'+str(seq_start)+'-'+str(seq_end)
print(' ')
print(myseq_str)
test_target = test_targets[test_index:test_index+1,:,:]
test_pred = seqnn_model.model.predict(test_inputs[test_index:test_index+1,:,:])
plt.figure(figsize=(8,4))
target_index = 0
vmin=-2; vmax=2
# plot pred
plt.subplot(121)
mat = from_upper_triu(test_pred[:,:,target_index], target_length1_cropped, hic_diags)
im = plt.matshow(mat, fignum=False, cmap= 'RdBu_r', vmax=vmax, vmin=vmin)
plt.colorbar(im, fraction=.04, pad = 0.05, ticks=[-2,-1, 0, 1,2]);
plt.title('pred-'+str(hic_num_to_name_dict[target_index]),y=1.15 )
plt.ylabel(myseq_str)
# plot target
plt.subplot(122)
mat = from_upper_triu(test_target[:,:,target_index], target_length1_cropped, hic_diags)
im = plt.matshow(mat, fignum=False, cmap= 'RdBu_r', vmax=vmax, vmin=vmin)
plt.colorbar(im, fraction=.04, pad = 0.05, ticks=[-2,-1, 0, 1,2]);
plt.title( 'target-'+str(hic_num_to_name_dict[target_index]),y=1.15)
plt.tight_layout()
plt.show()
# -
# ## Make a prediction from sequence
# +
### make a prediction from sequence ###
if not os.path.isfile('./data/hg38.ml.fa'):
print('downloading hg38.ml.fa')
subprocess.call('curl -o ./data/hg38.ml.fa.gz https://storage.googleapis.com/basenji_barnyard/hg38.ml.fa.gz', shell=True)
subprocess.call('gunzip ./data/hg38.ml.fa.gz', shell=True)
fasta_open = pysam.Fastafile('./data/hg38.ml.fa')
# +
# this example uses the sequence for the test set region
# with the corresponding test_index, but
# predictions can be made for any DNA sequence of length = seq_length = 2^20
chrm, seq_start, seq_end = sequences_test.iloc[test_index][0:3]
seq = fasta_open.fetch( chrm, seq_start, seq_end ).upper()
if len(seq) != seq_length: raise ValueError('len(seq) != seq_length')
# seq_1hot is a np.array with shape [2^20 bp, 4 nucleotides]
# representing 1-hot encoded DNA sequence
seq_1hot = dna_io.dna_1hot(seq)
# -
# expand input dimensions, as model accepts arrays of size [#regions,2^20bp, 4]
test_pred_from_seq = seqnn_model.model.predict(np.expand_dims(seq_1hot,0))
# +
# plot pred
plt.figure(figsize=(8,4))
target_index = 0
vmin=-2; vmax=2
#transform from flattened representation to symmetric matrix representation
mat = from_upper_triu(test_pred_from_seq[:,:,target_index], target_length1_cropped, hic_diags)
plt.subplot(121)
im = plt.matshow(mat, fignum=False, cmap= 'RdBu_r', vmax=vmax, vmin=vmin)
plt.colorbar(im, fraction=.04, pad = 0.05, ticks=[-2,-1, 0, 1,2]);
plt.title('pred-'+str(hic_num_to_name_dict[target_index]),y=1.15 )
plt.ylabel(myseq_str)
# plot target
plt.subplot(122)
mat = from_upper_triu(test_target[:,:,target_index], target_length1_cropped, hic_diags)
im = plt.matshow(mat, fignum=False, cmap= 'RdBu_r', vmax=vmax, vmin=vmin)
plt.colorbar(im, fraction=.04, pad = 0.05, ticks=[-2,-1, 0, 1,2]);
plt.title( 'target-'+str(hic_num_to_name_dict[target_index]),y=1.15)
plt.tight_layout()
plt.show()
| manuscripts/akita/explore_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# Download from: https://www.kaggle.com/lakshmi25npathi/imdb-dataset-of-50k-movie-reviews
import string
import pandas as pd
from tqdm import tqdm
import transformers
import pandas as pd
import json
import seaborn
df = pd.read_csv("./imdb_data.csv")
df.head()
# +
source_lengths = []
target_lengths = []
tokenizer = transformers.AutoTokenizer.from_pretrained("t5-large", cache_dir="/workspace/cache")
def get_len(text):
return len(tokenizer.encode(text))
label_set = ["positive", "negative"]
def create_pair(s, t, split):
prefix = ""
s = s.replace("<br /><br />", " ")
line = {
"translation": {
"s": prefix+s,
"t": t
}
}
source_lengths.append(get_len(s))
target_lengths.append(get_len(t))
with open("./"+split+"lines.json", 'a+') as outfile:
json.dump(line, outfile)
outfile.write("\n")
# +
x_list = []
y_list = []
count = 0
for index, row in df.iterrows():
s = row["review"].split(" ")[:256]
s = " ".join(s).strip()
t = row["sentiment"].strip()
x_list.append(s)
y_list.append(t)
count += 1
if count > 10000:
break
# -
from sklearn.model_selection import train_test_split
x_train, x_valid, y_train, y_valid = train_test_split(x_list, y_list, test_size=0.2, stratify=y_list)
# +
split = "train"
for s, t in zip(x_train, y_train):
create_pair(s, t, split)
split = "valid"
for s, t in zip(x_valid, y_valid):
create_pair(s, t, split)
# -
seaborn.displot(source_lengths)
seaborn.displot(target_lengths)
| sample_data/make_seq2seq_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (moneyball)
# language: python
# name: moneyball
# ---
# # Building Machine Learning Classifiers: Explore Gradient Boosting model with grid-search
# **Grid-search:** Exhaustively search all parameter combinations in a given grid to determine the best model.
# ### Read in & clean text
# +
import nltk
import pandas as pd
import re
from sklearn.feature_extraction.text import TfidfVectorizer
import string
stopwords = nltk.corpus.stopwords.words('english')
ps = nltk.PorterStemmer()
data = pd.read_csv("SMSSpamCollection.tsv", sep='\t')
data.columns = ['label', 'body_text']
def count_punct(text):
count = sum([1 for char in text if char in string.punctuation])
return round(count/(len(text) - text.count(" ")), 3)*100
data['body_len'] = data['body_text'].apply(lambda x: len(x) - x.count(" "))
data['punct%'] = data['body_text'].apply(lambda x: count_punct(x))
def clean_text(text):
text = "".join([word.lower() for word in text if word not in string.punctuation])
tokens = re.split('\W+', text)
text = [ps.stem(word) for word in tokens if word not in stopwords]
return text
tfidf_vect = TfidfVectorizer(analyzer=clean_text)
X_tfidf = tfidf_vect.fit_transform(data['body_text'])
X_features = pd.concat([data['body_len'], data['punct%'], pd.DataFrame(X_tfidf.toarray())], axis=1)
X_features.head()
# -
# ### Explore GradientBoostingClassifier Attributes & Hyperparameters
from sklearn.ensemble import GradientBoostingClassifier
print(dir(GradientBoostingClassifier))
print(GradientBoostingClassifier())
# ### Build our own Grid-search
from sklearn.metrics import precision_recall_fscore_support as score
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X_features, data['label'],
test_size=0.2)
def train_GB(est, max_depth, lr):
gb = GradientBoostingClassifier(n_estimators=est, max_depth=max_depth,
learning_rate=lr)
gb_model = gb.fit(X_train, y_train)
y_pred = gb_model.predict(X_test)
precision, recall, fscore, train_support = score(y_test, y_pred,
pos_label='spam',
average='binary')
print('Est: {} / Depth: {} / LR: {} ---- Precision: {} / Recall: {} / Accuracy: {}'.format(
est, max_depth, lr, round(precision, 3), round(recall, 3),
round((y_pred==y_test).sum()/len(y_pred), 3)))
for n_est in [50, 100, 150]:
for max_depth in [3, 7, 11, 15]:
for lr in [0.01, 0.1, 1]:
train_GB(n_est, max_depth, lr)
| nlp/Ex_Files_NLP_Python_ML_EssT/Exercise Files/Ch05/05_09/End/05_09.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.2 64-bit (''flair'': conda)'
# name: python392jvsc74a57bd0ec49377734e452f7232cd190f0d9cf7bf2c279fdb44b8c7a9fbd8a2977087685
# ---
# !python -m spacy download en_core_web_sm
import pandas as pd
import spacy_annotator as spa
import spacy
# +
nlp = spacy.load("en_core_web_sm")
# -
annotator = spa.Annotator(labels=["KEYWORD", "NULL"], model=nlp)
df = pd.DataFrame({"text":["This is C++ require"]})
df_labels = annotator.annotate(df = df, col_text = "text")
df_labels
| data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# TensorFlow Learn Tutorial from https://medium.com/@ilblackdragon/tensorflow-tutorial-part-1-c559c63c0cb1#.9p5m28k6d
from tensorflow.contrib import learn
import pandas as pd
from sklearn.cross_validation import train_test_split
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LogisticRegression
df = pd.read_csv('data/titanic_train.csv')
df.shape
df.columns
df.head(1)
y, X = df['Survived'], df[['Age', 'SibSp', 'Fare']].fillna(0)
# With scikit-learn
X_train, X_test, y_train, y_test = train_test_split(X,
y,
test_size=0.2,
random_state=42)
lr = LogisticRegression()
lr.fit(X_train, y_train)
print(accuracy_score(y_test, lr.predict(X_test)))
# With tf.learn
import random
random.seed(42) # to sample data the same way
classifier = learn.TensorFlowLinearClassifier(n_classes=2,
batch_size=128,
steps=500,
learning_rate=0.05)
classifier.fit(X_train, y_train)
print(accuracy_score(classifier.predict(X_test), y_test))
| tensorflow/tflearn-logistic-regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### 1.新环境版本检查
# !pip list --format=columns #不是必要!!
# ### 2. 常用模块版本查询
# +
# 导入所需模块
print('模块版本查询:')
import sys
print("Python version:{}".format(sys.version))
import pandas as pd
print("pandas version:{}".format(pd.__version__))
import numpy as np
print("numpy version:{}".format(np.__version__))
import scipy as sp
print("scipy version%s" %(np.__version__))
import IPython
from IPython import display
print("IPython verison:{}".format(IPython.__version__))
import sklearn
print("sklearn version:{}".format(sklearn.__version__))
import os #has no attribute '__version__'
# misc libraries 杂质
import random
import time
#ignore warnings 无视警告,纯静输出
import warnings
warnings.filterwarnings('ignore')
print('-'*50,end='\n')
print('数据存储情况查看:')
from subprocess import check_output
print(check_output(["ls", "../titanic/data"]).decode("utf8"))
# -
# ### 3. 导入所需模块
# +
# 基础模块
import pandas as pd
import numpy as np
import random as rnd
import os
import re
import itertools
# 可视化
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.pylab as pylab
import seaborn as sns
from pandas.tools.plotting import scatter_matrix
# 可视化参数设定
# %matplotlib inline
# %config InlineBackend.figure_format='retina' #提升mac专有retina屏幕显示效果
mpl.style.use('ggplot')
sns.set_style('white')
pylab.rcParams['figure.figsize'] = 12,8
# plt.rcParams['figure.figsize'] = (8.0, 4.0) # 设置figure_size尺寸
# plt.rcParams['image.interpolation'] = 'nearest' # 设置 interpolation style
# plt.rcParams['image.cmap'] = 'gray' # 设置 颜色 style
# 模型辅助方法
# Pipeline
from sklearn.pipeline import make_pipeline
from sklearn.pipeline import Pipeline
# 特征工程
from sklearn.preprocessing import OneHotEncoder, LabelEncoder
from sklearn import feature_selection
from sklearn import model_selection
from sklearn import metrics
# 模型
from sklearn.linear_model import LogisticRegression #logistic regression
from sklearn.linear_model import Perceptron
from sklearn import svm #support vector Machine
from sklearn.ensemble import RandomForestClassifier #Random Forest
from sklearn.neighbors import KNeighborsClassifier #KNN
from sklearn.naive_bayes import GaussianNB #Naive bayes
from sklearn.tree import DecisionTreeClassifier #Decision Tree
from sklearn.model_selection import train_test_split #training and testing data split
from sklearn import metrics #accuracy measure
from sklearn.metrics import confusion_matrix #for confusion matrix
from sklearn.ensemble import VotingClassifier
from sklearn.ensemble import AdaBoostClassifier
from sklearn.neural_network import MLPClassifier
from xgboost import XGBClassifier
# Grid
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import StratifiedShuffleSplit
from sklearn.feature_selection import RFE
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import RandomizedSearchCV
import scipy.stats as st
# Evalaluation
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn.model_selection import cross_val_score
from sklearn.metrics import confusion_matrix, roc_curve, auc, accuracy_score
# Esemble
#ignore warnings 无视警告,纯静输出
import warnings
warnings.filterwarnings(action='ignore')
warnings.filterwarnings(action='ignore', category=DeprecationWarning)
warnings.filterwarnings(action='ignore', category=FutureWarning)
print('-'*50,end='\n')
print('数据存储情况查看:')
from subprocess import check_output
# print(check_output(["ls", "./data"]).decode("utf8")) 按需改进
#misc
import time
import datetime
import platform
start = time.time()
# -
| Pyframe/import.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#default_exp prodigy_demo
# +
#no_test
#data structure imports
import pandas as pd
import numpy as np
#python imports
import random
#modeling imports
from spacy.util import fix_random_seed
from ssda_nlp.collate import *
from ssda_nlp.split_data import *
from ssda_nlp.modeling import *
from ssda_nlp.preprocessing import *
from ssda_nlp.model_performance_utils import *
# -
#no_test
seed = 2436
random.seed(seed)
fix_random_seed(seed)
# # First Book of Baptisms, Matanzas (15834)
# +
#no_test
collated_df = prodigy_output_to_collated_df("transcriptions//15834_annot.jsonl")
# Split data
train_df, valid_df, test_df = split_data_grp(collated_df, prop_train = 0.7, prop_validation = 0.2, grp_var = 'entry_no', seed=seed)
# Generate Spacy datasets
train_spacy = genSpaCyInput(train_df)
# Look at column names
train_df.head(10)
# +
#no_test
train_spacy[:2]
# +
#no_test
#if you want the same model everytime:
#random.seed(seed)
#fix_random_seed(seed)
#spacy parameters
cping = {'start':16, 'end':32, 'cp_rate':1.05}
solves = {'learn_rate':0.001}
dpout = 0.6
save_mdir = 'models/15834'
#load model from pretrained
nlp_model = load_model()
nlp_model, perf_df = model_meta_training(nlp_model, train_spacy, valid_df, verbose=True, save_dir = save_mdir,
n_iter=10, solver_params=solves, compound_params=cping, dropout=dpout)
# -
# # Second Book of Baptisms, St. Augustine (239746)
# +
#no_test
collated_df = prodigy_output_to_collated_df("transcriptions/239746_annot.jsonl")
# Split data
train_df, valid_df, test_df = split_data_grp(collated_df, prop_train = 0.7, prop_validation = 0.2, grp_var = 'entry_no', seed=seed)
# Generate Spacy datasets
train_spacy = genSpaCyInput(train_df)
# Look at column names
train_df.head(10)
# +
#no_test
train_spacy[:2]
# +
#no_test
#if you want the same model everytime:
#random.seed(seed)
#fix_random_seed(seed)
#spacy parameters
cping = {'start':16, 'end':32, 'cp_rate':1.05}
solves = {'learn_rate':0.001}
dpout = 0.6
save_mdir = 'models/239746'
#load model from pretrained
nlp_model = load_model('es_core_news_md')
nlp_model, perf_df = model_meta_training(nlp_model, train_spacy, valid_df, verbose=True, save_dir = save_mdir,
n_iter=10, solver_params=solves, compound_params=cping, dropout=dpout)
# -
# It's interesting how similar this training performance was to the Matanzas volume above. If this is replicated across more data sets, it probably means something (although I'm not sure what exactly).
# +
#no_test
collated_df = prodigy_output_to_collated_df("transcriptions/166470_annot.jsonl")
# Split data
train_df, valid_df, test_df = split_data_grp(collated_df, prop_train = 0.9, prop_validation = 0.05, grp_var = 'entry_no', seed=seed)
# Generate Spacy datasets
train_spacy = genSpaCyInput(collated_df)
# Look at column names
train_spacy[:5]
# +
#no_test
#if you want the same model everytime:
#random.seed(seed)
#fix_random_seed(seed)
#spacy parameters
cping = {'start':16, 'end':32, 'cp_rate':1.05}
solves = {'learn_rate':0.001}
dpout = 0.6
save_mdir = 'models/166470'
#load model from pretrained
nlp_model = load_model()
nlp_model, perf_df = model_meta_training(nlp_model, train_spacy, valid_df, verbose=True, save_dir = save_mdir,
n_iter=10, solver_params=solves, compound_params=cping, dropout=dpout)
# -
| 61-prodigy-output-training-demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Neural Network Modeling
# ### Import Libraries and Data
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import scipy as sp
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler, PolynomialFeatures, OneHotEncoder
from sklearn.metrics import balanced_accuracy_score, confusion_matrix, mean_squared_error, median_absolute_error, r2_score
from sklearn.pipeline import make_pipeline
from sklearn.metrics import plot_confusion_matrix, balanced_accuracy_score, f1_score, recall_score, precision_score, roc_auc_score, plot_roc_curve, classification_report
from sklearn.model_selection import train_test_split, GridSearchCV, cross_val_score
import tensorflow.keras
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, BatchNormalization, Input
from tensorflow.keras.regularizers import l1, l2, l1_l2
from tensorflow.keras.callbacks import EarlyStopping
from tensorflow.keras.layers import Flatten
from tensorflow.keras.utils import to_categorical
import warnings
warnings.filterwarnings("ignore")
# -
df = pd.read_csv('../../01_data/cleaned_data/school_df_v6.csv')
df = df.iloc[:,:45]
df.dropna(how='all', inplace=True)
# drop 2 schools that have no attendance data
df.drop(df[df['code'] == 201].index, inplace = True)
df.drop(df[df['code'] == 347].index, inplace = True)
df = df.replace(-1,np.nan)
df.dropna(how='any',inplace=True)
df = df.replace(-1,np.nan)
df.dropna(how='any',inplace=True)
df = df[df['star_rating_SY1819'] !=-1]
df = df.replace(-1,np.nan)
df.dropna(how='any',inplace=True)
df = df[df['star_rating_SY1718'] !=-1]
X=df[['enrollment_SY1718', 'enrollment_SY1819', 'capacity_SY1718',
'capacity_SY1819', 'latitude', 'longitude', 'cluster', 'ward','pct_0_SY1819',
'pct_1-5_SY1819', 'pct_6-10_SY1819', 'pct_11-20_SY1819', 'pct_20+_SY1819',
'pct_0_SY1718', 'pct_1-5_SY1718', 'pct_6-10__SY1718', 'pct_11-20_SY1718',
'pct_20+_SY1718', 'budgeted_amount_FY16', 'budgeted_enrollment_FY16',
'budgeted_amount_FY17', 'budgeted_enrollment_FY17', 'pct_meet_exceed_math_SY1718',
'pct_meet_exceed_ela_SY1718', 'pct_meet_exceed_math_SY1819', 'pct_meet_exceed_ela_SY1819']]
y = df['star_rating_SY1718']
y = y-1
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=7)
sscaler = StandardScaler()
X_train_scaled = sscaler.fit_transform(X_train)
X_test_scaled = sscaler.fit_transform(X_test)
y.value_counts()
# ### Build the model
y.value_counts(normalize=True)
y_train_cat = to_categorical(y_train)
y_test_cat = to_categorical(y_test)
model = Sequential()
model.add(Input(shape=(X.shape[1],)))
model.add(BatchNormalization())
model.add(Dense(128, activation='relu', kernel_regularizer=l2(0.1)))
model.add(Dropout(0.2))
model.add(Dense(128, activation='relu', kernel_regularizer=l2(0.1)))
model.add(Dropout(0.2))
model.add(Dense(5, activation='softmax'))
model.compile(loss='sparse_categorical_crossentropy', optimizer='rmsprop', metrics=['accuracy'])
history = model.fit(X_train_scaled, y_train,
validation_data=(X_test_scaled, y_test),
epochs=300,
verbose=0)
model.evaluate(X_test, y_test)
# +
fig = plt.figure(figsize = (10, 5))
ax= fig.add_subplot(1, 2, 1)
plt.plot(history.history['val_loss'], label = 'Validation Loss')
plt.plot(history.history['loss'], label = 'Train Loss')
# plt.title('Loss')
plt.legend()
plt.xlabel('Epoch')
plt.ylabel('Loss')
ax= fig.add_subplot(1, 2, 2)
plt.plot(history.history['val_accuracy'], label = 'Validation Accuracy',)
plt.plot(history.history['accuracy'], label = 'Train Accuracy')
# plt.title('Accuracy')
plt.legend()
plt.xlabel('Epoch')
plt.ylabel('Accuracy');
# plt.savefig('./figures/neural_network.png');
# -
#model evaluation seems to be overfit and accuracy is very low
model.evaluate(X_test, y_test)
| 02_notebooks/03_Modeling/f_neural_network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # motion estimation in spikeinterface
#
# In 2021,the SpikeInterface project has started to implemented `sortingcomponents`, a modular module for spike sorting steps.
#
# Here is an overview or our progress integrating motion (aka drift) estimation and correction.
#
#
# This notebook will be based on the open dataset from <NAME> published in 2021
# "Imposed motion datasets" from Steinmetz et al. Science 2021
# https://figshare.com/articles/dataset/_Imposed_motion_datasets_from_Steinmetz_et_al_Science_2021/14024495
#
#
# The motion estimation is done in several modular steps:
# 1. detect peaks
# 2. localize peaks:
# * **"center of mass"**
# * **"monopolar_triangulation"** by <NAME> and <NAME>
# https://openreview.net/pdf?id=ohfi44BZPC4
# 3. estimation motion:
# * **rigid** or **non rigid**
# * **"decentralized"** by <NAME> and <NAME>
# DOI : 10.1109/ICASSP39728.2021.9414145
# 4. compute motion corrected peak localizations for visualization
#
#
# Here we will show this chain:
# * **detect peaks > localize peaks with "monopolar_triangulation" > estimation motion "decentralized" (both rigid and nonrigid)**
#
#
# %load_ext autoreload
# %autoreload 2
from pathlib import Path
import spikeinterface.full as si
import numpy as np
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (20, 12)
from probeinterface.plotting import plot_probe
# local folder
base_folder = Path('/Users/charlie/data/')
dataset_folder = base_folder / 'dataset1'
preprocess_folder = base_folder / 'dataset1_preprocessed'
peak_folder = base_folder / 'dataset1_peaks'
peak_folder.mkdir(exist_ok=True)
# global kwargs for parallel computing
job_kwargs = dict(
n_jobs=8,
chunk_size=30_000,
progress_bar=True,
)
# read the file
rec = si.read_spikeglx(dataset_folder)
rec
fig, ax = plt.subplots()
si.plot_probe_map(rec, ax=ax)
ax.set_ylim(-150, 200)
# ## preprocess
#
# This takes 4 min for 30min of signals
if not preprocess_folder.exists():
rec_filtered = si.bandpass_filter(rec, freq_min=300., freq_max=6000.)
rec_preprocessed = si.common_reference(rec_filtered, reference='global', operator='median')
rec_preprocessed.save(folder=preprocess_folder, **job_kwargs)
rec_preprocessed = si.load_extractor(preprocess_folder)
# plot and check spikes
si.plot_timeseries(rec_preprocessed, time_range=(100, 110), channel_ids=rec.channel_ids[50:60])
# ## estimate noise
noise_levels = si.get_noise_levels(rec_preprocessed, return_scaled=False)
fig, ax = plt.subplots(figsize=(8,6))
ax.hist(noise_levels, bins=10)
ax.set_title('noise across channel')
# ## detect peaks
#
# This take 1min30s
from spikeinterface.sortingcomponents.peak_detection import detect_peaks
if not (peak_folder / 'peaks.npy').exists():
peaks = detect_peaks(
rec_preprocessed,
method='locally_exclusive',
local_radius_um=100,
peak_sign='neg',
detect_threshold=5,
n_shifts=5,
noise_levels=noise_levels,
**job_kwargs,
)
np.save(peak_folder / 'peaks.npy', peaks)
peaks = np.load(peak_folder / 'peaks.npy')
print(peaks.shape)
print(rec_preprocessed)
# ## localize peaks
#
# Here we chosse **'monopolar_triangulation' with log barrier**
from spikeinterface.sortingcomponents.peak_localization import localize_peaks
if not (peak_folder / 'peak_locations_monopolar_triangulation_log_limit.npy').exists():
peak_locations = localize_peaks(
rec_preprocessed,
peaks,
ms_before=0.3,
ms_after=0.6,
method='monopolar_triangulation',
method_kwargs={
'local_radius_um': 100.,
'max_distance_um': 1000.,
'optimizer': 'minimize_with_log_penality',
},
**job_kwargs,
)
np.save(peak_folder / 'peak_locations_monopolar_triangulation_log_limit.npy', peak_locations)
print(peak_locations.shape)
peak_locations = np.load(peak_folder / 'peak_locations_monopolar_triangulation_log_limit.npy')
print(peak_locations.dtype.fields, peak_locations)
# ## plot on probe
def clip_values_for_cmap(x):
low, high = np.percentile(x, [5, 95])
return np.clip(x, low, high)
fig, axs = plt.subplots(ncols=2, sharey=True, figsize=(15, 10))
ax = axs[0]
si.plot_probe_map(rec_preprocessed, ax=ax)
ax.scatter(peak_locations['x'], peak_locations['y'], c=clip_values_for_cmap(peaks['amplitude']), s=1, alpha=0.002, cmap=plt.cm.plasma)
ax.set_xlabel('x')
ax.set_ylabel('y')
if 'z' in peak_locations.dtype.fields:
ax = axs[1]
ax.scatter(peak_locations['z'], peak_locations['y'], c=clip_values_for_cmap(peaks['amplitude']), s=1, alpha=0.002, cmap=plt.cm.plasma)
ax.set_xlabel('z')
ax.set_xlim(0, 150)
ax.set_ylim(1800, 2500)
# ## plot peak depth vs time
fig, ax = plt.subplots()
x = peaks['sample_ind'] / rec_preprocessed.get_sampling_frequency()
y = peak_locations['y']
ax.scatter(x, y, s=1, c=clip_values_for_cmap(peaks['amplitude']), cmap=plt.cm.plasma, alpha=0.25)
ax.set_ylim(1300, 2500)
# ## motion estimate : rigid with decentralized
from spikeinterface.sortingcomponents.motion_estimation import (
estimate_motion,
make_motion_histogram,
compute_pairwise_displacement,
compute_global_displacement
)
# +
bin_um = 5
bin_duration_s=5.
motion_histogram, temporal_bins, spatial_bins = make_motion_histogram(
rec_preprocessed,
peaks,
peak_locations=peak_locations,
bin_um=bin_um,
bin_duration_s=bin_duration_s,
direction='y',
weight_with_amplitude=False,
)
print(motion_histogram.shape, temporal_bins.size, spatial_bins.size)
# -
fig, ax = plt.subplots()
extent = (temporal_bins[0], temporal_bins[-1], spatial_bins[0], spatial_bins[-1])
motion_histogram_vis = np.zeros_like(motion_histogram)
vals = motion_histogram[motion_histogram > 0]
vals -= vals.min()
vals = np.clip(vals, 0, np.percentile(vals, 95))
vals /= vals.max()
motion_histogram_vis[motion_histogram > 0] = 3 + 20 * vals
im = ax.imshow(
motion_histogram_vis.T,
interpolation='nearest',
origin='lower',
aspect='auto',
extent=extent,
cmap=plt.cm.cubehelix,
)
im.set_clim(0, 30)
ax.set_ylim(1300, 2500)
ax.set_xlabel('time[s]')
ax.set_ylabel('depth[um]')
# ## pariwise displacement from the motion histogram
#
# +
conv_engine = "numpy"
try:
import torch
conv_engine = "torch"
except ImportError:
pass
pairwise_displacement, pairwise_displacement_weight = compute_pairwise_displacement(
motion_histogram, bin_um, method='conv', conv_engine=conv_engine, progress_bar=True, max_displacement_um=600
)
np.save(peak_folder / 'pairwise_displacement_conv2d.npy', pairwise_displacement)
# -
fig, ax = plt.subplots()
extent = (temporal_bins[0], temporal_bins[-1], temporal_bins[0], temporal_bins[-1])
# extent = None
im = ax.imshow(
pairwise_displacement,
interpolation='nearest',
cmap='PiYG',
origin='lower',
aspect='auto',
extent=extent,
)
im.set_clim(-40, 40)
ax.set_aspect('equal')
fig.colorbar(im)
# ## estimate motion (rigid) from the pairwise displacement
pairwise_displacement
# +
# motion = compute_global_displacement(pairwise_displacement)
motion_gd = compute_global_displacement(pairwise_displacement, convergence_method='gradient_descent')
motion_sparse_lsqr = compute_global_displacement(
pairwise_displacement,
# thresholding correlations
sparse_mask=pairwise_displacement_weight > 0.6,
# weighting by correlations
pairwise_displacement_weight=pairwise_displacement_weight,
convergence_method='lsqr_robust',
lsqr_robust_n_iter=20,
robust_regression_sigma=2,
)
# -
fig, ax = plt.subplots()
ax.plot(temporal_bins[:-1], motion_gd, label="convergence_method='gradient_descent'")
ax.plot(temporal_bins[:-1], motion_sparse_lsqr, label="convergence_method='lsqr_robust'")
plt.legend()
# ## motion estimation with one unique funtion
#
# Internally `estimate_motion()` does:
# * make_motion_histogram()
# * compute_pairwise_displacement()
# * compute_global_displacement()
#
from spikeinterface.sortingcomponents.motion_estimation import estimate_motion
from spikeinterface.widgets import plot_pairwise_displacement, plot_displacement
# +
method='decentralized_registration'
method_kwargs = dict(
pairwise_displacement_method='conv',
convergence_method='gradient_descent',
#convergence_method='lsqr_robust',
)
# method='decentralized_registration'
# method_kwargs = dict(
# pairwise_displacement_method='phase_cross_correlation',
# convergence_method='lsqr_robust',
# )
motion, temporal_bins, spatial_bins, extra_check = estimate_motion(
rec_preprocessed,
peaks,
peak_locations=peak_locations,
direction='y',
bin_duration_s=5.,
bin_um=10.,
method=method,
method_kwargs=method_kwargs,
non_rigid_kwargs=None,
output_extra_check=True,
progress_bar=True,
verbose=False,
upsample_to_histogram_bin=False,
)
# -
plot_pairwise_displacement(motion, temporal_bins, spatial_bins, extra_check, ncols=4)
plot_displacement(motion, temporal_bins, spatial_bins, extra_check, with_histogram=True)
fig, ax = plt.subplots()
x = peaks['sample_ind'] / rec_preprocessed.get_sampling_frequency()
y = peak_locations['y']
ax.scatter(x, y, s=1, color='k', alpha=0.05)
plot_displacement(motion, temporal_bins, spatial_bins, extra_check, with_histogram=False, ax=ax)
# ## motion estimation non rigid
#
# +
method='decentralized_registration'
method_kwargs = dict(
pairwise_displacement_method='conv',
convergence_method='gradient_descent',
conv_engine=conv_engine,
batch_size=8,
corr_threshold=0.6,
)
motion, temporal_bins, spatial_bins, extra_check = estimate_motion(
rec_preprocessed,
peaks,
peak_locations=peak_locations,
direction='y',
bin_duration_s=5.,
bin_um=5.,
method=method,
method_kwargs=method_kwargs,
non_rigid_kwargs=dict(bin_step_um=400, sigma=3),
margin_um=-400,
output_extra_check=True,
progress_bar=True,
verbose=False,
upsample_to_histogram_bin=False,
)
# -
fig, ax = plt.subplots()
for win in extra_check['non_rigid_windows']:
ax.plot(win, extra_check['spatial_hist_bins'][:-1])
plot_pairwise_displacement(motion, temporal_bins, spatial_bins, extra_check, ncols=4)
plot_displacement(motion, temporal_bins, spatial_bins, extra_check, with_histogram=True)
fig, ax = plt.subplots()
x = peaks['sample_ind'] / rec_preprocessed.get_sampling_frequency()
y = peak_locations['y']
ax.scatter(x, y, s=1, color='k', alpha=0.05)
plot_displacement(motion, temporal_bins, spatial_bins, extra_check, with_histogram=False, ax=ax)
ax.set_ylim(0, 2000)
fig, ax = plt.subplots()
ax.plot(temporal_bins, motion);
fig, ax = plt.subplots()
im = ax.imshow(motion.T,
interpolation='nearest',
cmap='PiYG',
origin='lower',
aspect='auto',
# extent=extent,
)
im.set_clim(-40, 40)
ax.set_aspect('equal')
fig.colorbar(im);
# ## upsample motion estimate to original domain and apply motion correction to peak localizations
from spikeinterface.sortingcomponents.motion_correction import correct_motion_on_peaks
motion_up, temporal_bins, spatial_bins_up, extra_check_up = estimate_motion(
rec_preprocessed,
peaks,
peak_locations=peak_locations,
direction='y',
bin_duration_s=5.,
bin_um=5.,
method=method,
method_kwargs=method_kwargs,
non_rigid_kwargs=dict(bin_step_um=300, sigma=3),
margin_um=-400,
output_extra_check=True,
progress_bar=True,
verbose=False,
upsample_to_histogram_bin=True,
)
fig, ax = plt.subplots()
im = ax.imshow(motion_up.T,
interpolation='nearest',
cmap='PiYG',
origin='lower',
aspect='auto',
extent=extent,
)
im.set_clim(-40, 40)
ax.set_aspect('equal')
fig.colorbar(im);
times = peaks['sample_ind'] / rec_preprocessed.get_sampling_frequency()
corrected_peak_locations = correct_motion_on_peaks(peaks, peak_locations, times,
motion_up, temporal_bins, spatial_bins_up,
direction='y', progress_bar=False)
fig, ax = plt.subplots()
x = peaks['sample_ind'] / rec_preprocessed.get_sampling_frequency()
y = peak_locations['y']
ax.scatter(x, y, s=1, c=clip_values_for_cmap(peaks['amplitude']), cmap=plt.cm.plasma, alpha=0.25)
ax.set_ylim(1300, 2500)
ax.set_title("unregistered localizations")
ax.set_xlabel("time (s)")
ax.set_ylabel("depth (um)");
fig, ax = plt.subplots()
x = peaks['sample_ind'] / rec_preprocessed.get_sampling_frequency()
y = corrected_peak_locations['y']
ax.scatter(x, y, s=1, c=clip_values_for_cmap(peaks['amplitude']), cmap=plt.cm.plasma, alpha=0.25)
ax.set_ylim(1300, 2500)
ax.set_title("registered localizations")
ax.set_xlabel("time (s)")
ax.set_ylabel("depth (um)");
| projects/motion-correction-in-si/motion_estimation_and_correction_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Week 1: August 1st - August 7th
#
# >All solutions were what I thought by myself.
#
# ## Detect Capital
#
# Given a word, you need to judge whether the usage of capitals in it is right or not.
#
# We define the usage of capitals in a word to be right when one of the following cases holds:
#
# 1. All letters in this word are capitals, like "USA".
# 2. All letters in this word are not capitals, like "leetcode".
# 3. Only the first letter in this word is capital, like "Google".
#
# Otherwise, we define that this word doesn't use capitals in a right way.
#
# **Example 1:**
# ```
# Input: "USA"
# Output: True
# ```
# **Example 2:**
# ```
# Input: "FlaG"
# Output: False
# ```
# +
class Solution:
def detectCapitalUse(self, word):
if word.isalpha() == True:
if word.istitle() == True:
return True
elif word.isupper() == True:
return True
elif word.islower() == True:
return True
else:
return False
Solution = Solution()
# +
inputs = "USA"
result = Solution.detectCapitalUse(inputs)
print(result)
# +
inputs = "FlaG"
result = Solution.detectCapitalUse(inputs)
print(result)
# -
| August_LeetCoding_Challenge/August_LeetCoding_Challenge_Week1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %%
import os
from os import path
from pcdet.utils import common_utils
from pcdet.models import build_network, load_data_to_gpu
from pcdet.datasets import DatasetTemplate
from pcdet.config import cfg, cfg_from_yaml_file
import torch
import numpy as np
import argparse
import glob
from pathlib import Path
from eval_utils import eval_utils
# from tools.visual_utils import visualize_utils as V
ABS_PATH_PREFIX = os.getcwd()
# # %matplotlib inline
# # %matplotlib notebook
# %matplotlib widget
import matplotlib.pyplot as plt
# -
#%%
class Args:
def __init__(self):
self.cfg_file = path.join(
ABS_PATH_PREFIX, 'cfgs/kitti_models/second.yaml')
self.data_path = path.join(
ABS_PATH_PREFIX, '../data/kitti/')
self.ckpt = path.join(ABS_PATH_PREFIX,
'../output/kitti_models/second/def/ckpt/checkpoint_epoch_50.pth')
self.ext = '.bin'
print("cfg _file = ", self.cfg_file)
# + tags=["outputPrepend"]
#%%
from pcdet.datasets.kitti.kitti_dataset import KittiDataset
os.chdir(ABS_PATH_PREFIX)
args = Args()
cfg_from_yaml_file(args.cfg_file, cfg)
# args, cfg = parse_config()
print("args = ", args)
print("cfg = ", cfg)
logger = common_utils.create_logger()
logger.info(
'-----------------Quick Demo of OpenPCDet-------------------------')
demo_dataset = KittiDataset(
dataset_cfg=cfg.DATA_CONFIG, class_names=cfg.CLASS_NAMES, training=False,
root_path=Path(args.data_path), logger=logger
)
logger.info(f'Total number of samples: \t{len(demo_dataset)}')
model = build_network(model_cfg=cfg.MODEL, num_class=len(
cfg.CLASS_NAMES), dataset=demo_dataset)
model.load_params_from_file(filename=args.ckpt, logger=logger, to_cpu=True)
model.cuda()
model.eval()
metric = {
'gt_num' : 0,
}
for cur_thresh in cfg.MODEL.POST_PROCESSING.RECALL_THRESH_LIST:
metric['recall_roi_%s' % str(cur_thresh)] = 0
metric['recall_rcnn_%s' % str(cur_thresh)] = 0
with torch.no_grad():
for idx, data_dict in enumerate(demo_dataset):
if idx == 3:
break
logger.info(f'Visualized sample index: \t{idx + 1}')
data_dict = demo_dataset.collate_batch([data_dict])
load_data_to_gpu(data_dict)
pred_dicts, ret_dict = model.forward(data_dict)
disp_dict = {}
eval_utils.statistics_info(cfg, ret_dict, metric, disp_dict)
annos = demo_dataset.generate_prediction_dicts(
data_dict, pred_dicts, demo_dataset.class_names, None
)
# pred_dicts includes:
# pred_boxes, N x 7
# pred_scores, N
# pred_labels
print(pred_dicts)
print(data_dict.keys())
# V.draw_scenes(
# points=data_dict['points'][:,1:], ref_boxes=pred_dicts[0]['pred_boxes'],
# ref_scores=pred_dicts[0]['pred_scores'], ref_labels=pred_dicts[0]['pred_labels']
# )
# mlab.show(stop=True)
logger.info('Demo done.')
# +
demo_dataset.get_image_shape('000003')
# -
demo_dataset[2]
demo_dataset[13]['gt_boxes']
data_dict['gt_boxes'].shape
pred_dicts[0]
ret_dict
annos
from mpl_toolkits.mplot3d import Axes3D
def visualize_pts(pts, fig=None, bgcolor=(1,1,1), fgcolor=(1.0, 1.0, 1.0),
show_intensity=True, size=(600, 600), draw_origin=True):
if not isinstance(pts, np.ndarray):
pts = pts.cpu().numpy()
if fig is None:
fig = plt.figure(figsize=(20, 10))
ax = fig.add_subplot(111, projection='3d')
ax.set_facecolor(bgcolor)
if show_intensity:
ax.scatter(pts[:, 0], pts[:, 1], pts[:, 2], c=pts[:,3], s=0.01, marker=',')
else:
ax.scatter(pts[:, 0], pts[:, 1], pts[:, 2], s=0.01, marker=',')
if draw_origin:
ax.plot([0,3], [0,0], [0,0], color=(0,0,1))
ax.plot([0,0], [0,3], [0,0], color=(0,1,0))
ax.plot([0,0], [0,0], [0,3], color=(1,0,0))
return ax
# if draw_origin:
# +
fig = plt.figure()
points = data_dict['points']
plt.plot(points[:,2])
fig = plt.figure()
plt.plot(points[:,1])
fig = plt.figure()
plt.plot(points[:,0])
# -
ax = visualize_pts(points)
# +
def draw_grid(x1, y1, x2, y2, ax, color=(1,0.5,0.5)):
ax.plot([x1, x1], [y1, y2], [0, 0], color=color, linewidth=1)
ax.plot([x2, x2], [y1, y2], [0, 0], color=color, linewidth=1)
ax.plot([x1, x2], [y1, y1], [0, 0], color=color, linewidth=1)
ax.plot([x1, x2], [y2, y2], [0, 0], color=color, linewidth=1)
return ax
def draw_multi_grid_range(ax, grid_size=20, bv_range=(-60, -60, 60, 60)):
for x in range(bv_range[0], bv_range[2], grid_size):
for y in range(bv_range[1], bv_range[3], grid_size):
ax = draw_grid(x, y, x + grid_size, y+grid_size, ax)
return ax
# -
draw_multi_grid_range(ax, bv_range=(0, -40, 80, 40))
# +
def check_numpy_to_torch(x):
if isinstance(x, np.ndarray):
return torch.from_numpy(x).float(), True
return x, False
def rotate_points_along_z(points, angle):
"""
Args:
points: (B, N, 3 + C)
angle: (B), angle along z-axis, angle increases x ==> y
Returns:
"""
points, is_numpy = check_numpy_to_torch(points)
angle, _ = check_numpy_to_torch(angle)
cosa = torch.cos(angle)
sina = torch.sin(angle)
zeros = angle.new_zeros(points.shape[0])
ones = angle.new_ones(points.shape[0])
rot_matrix = torch.stack((
cosa, sina, zeros,
-sina, cosa, zeros,
zeros, zeros, ones
), dim=1).view(-1, 3, 3).float()
points_rot = torch.matmul(points[:, :, 0:3], rot_matrix)
points_rot = torch.cat((points_rot, points[:, :, 3:]), dim=-1)
return points_rot.numpy() if is_numpy else points_rot
def boxes_to_corners_3d(boxes3d):
"""
7 -------- 4
/| /|
6 -------- 5 .
| | | |
. 3 -------- 0
|/ |/
2 -------- 1
Args:
boxes3d: (N, 7) [x, y, z, dx, dy, dz, heading], (x, y, z) is the box center
Returns:
"""
boxes3d, is_numpy = check_numpy_to_torch(boxes3d)
template = boxes3d.new_tensor((
[1, 1, -1], [1, -1, -1], [-1, -1, -1], [-1, 1, -1],
[1, 1, 1], [1, -1, 1], [-1, -1, 1], [-1, 1, 1],
)) / 2
corners3d = boxes3d[:, None, 3:6].repeat(1, 8, 1) * template[None, :, :]
corners3d = rotate_points_along_z(corners3d.view(-1, 8, 3), boxes3d[:, 6]).view(-1, 8, 3)
corners3d += boxes3d[:, None, 0:3]
return corners3d.numpy() if is_numpy else corners3d
# +
def draw_corners3d(corners3d, ax, color=(1, 1, 1), line_width=1, cls=None, tag='', max_num=500, tube_radius=None):
"""
:param corners3d: (N, 8, 3)
:param fig:
:param color:
:param line_width:
:param cls:
:param tag:
:param max_num:
:return:
"""
num = min(max_num, len(corners3d))
for n in range(num):
b = corners3d[n] # (8, 3)
if cls is not None:
if isinstance(cls, np.ndarray):
ax.text(b[6, 0], b[6, 1], b[6, 2], '%.2f' % cls[n], fontsize=0.3, color=color)
else:
ax.text(b[6, 0], b[6, 1], b[6, 2], '%s' % cls[n], fontsize=0.3, color=color)
for k in range(0, 4):
i, j = k, (k + 1) % 4
ax.plot([b[i, 0], b[j, 0]], [b[i, 1], b[j, 1]], [b[i, 2], b[j, 2]], color=color,
linewidth=line_width)
i, j = k + 4, (k + 1) % 4 + 4
ax.plot([b[i, 0], b[j, 0]], [b[i, 1], b[j, 1]], [b[i, 2], b[j, 2]], color=color,
linewidth=line_width)
i, j = k, k + 4
ax.plot([b[i, 0], b[j, 0]], [b[i, 1], b[j, 1]], [b[i, 2], b[j, 2]], color=color,
linewidth=line_width)
i, j = 0, 5
ax.plot([b[i, 0], b[j, 0]], [b[i, 1], b[j, 1]], [b[i, 2], b[j, 2]], color=color,
linewidth=line_width)
i, j = 1, 4
ax.plot([b[i, 0], b[j, 0]], [b[i, 1], b[j, 1]], [b[i, 2], b[j, 2]], color=color,
linewidth=line_width)
return ax
# +
ref_boxes=pred_dicts[0]['pred_boxes'].cpu().numpy()
ref_scores=pred_dicts[0]['pred_scores'].cpu().numpy()
ref_labels=pred_dicts[0]['pred_labels'].cpu().numpy()
ref_corners3d = boxes_to_corners_3d(ref_boxes)
ax = draw_corners3d(ref_corners3d, ax, color=(0,1,0), cls=ref_scores, max_num=100)
box_colormap = [
[1, 1, 1],
[0, 1, 0],
[0, 1, 1],
[1, 1, 0],
]
gt_boxes = data_dict['gt_boxes']
if gt_boxes is not None:
corners3d = boxes_to_corners_3d(gt_boxes)
ax = draw_corners3d(corners3d, ax, color=(0, 0, 1), max_num=100)
if ref_boxes is not None:
ref_corners3d = boxes_to_corners_3d(ref_boxes)
if ref_labels is None:
ax = draw_corners3d(ref_corners3d, ax, color=(0, 1, 0), cls=ref_scores, max_num=100)
else:
for k in range(ref_labels.min(), ref_labels.max() + 1):
cur_color = tuple(box_colormap[k % len(box_colormap)])
mask = (ref_labels == k)
ax = draw_corners3d(ref_corners3d[mask], ax, color=cur_color, cls=ref_scores[mask], max_num=100)
# -
data_dict.keys()
| tools/demo2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # BBVDATA Training
# ### Импортирование инструментов и инициализация данных
# +
import math
import numpy
import pandas as pd
import matplotlib.pyplot as plt
from keras.layers import LSTM
from keras.layers import Dense
from keras.models import Sequential
from keras.callbacks import ReduceLROnPlateau
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import mean_squared_error
# -
numpy.random.seed(7)
dataframe = pd.read_csv("preprocessed.csv").drop("Unnamed: 0", axis=1)
learning_rate_reduction = ReduceLROnPlateau(monitor='val_acc', patience=3, verbose=1, factor=0.5, min_lr=0.00001)
def create_dataset(d, look_back=1):
df = pd.DataFrame()
for i, col in enumerate(dataframe.columns):
df[col] = d.T[i]
d = df
# normalize the dataset
x_train, y_train = d.drop(["load", "amount"],axis=1).as_matrix(), d[["load", "amount"]].as_matrix()
scaler_x = MinMaxScaler(feature_range=(0, 1)).fit(x_train)
scaler_y = MinMaxScaler(feature_range=(0, 1)).fit(y_train)
x_train = scaler_x.transform(x_train)
y_train = scaler_y.transform(y_train)
return numpy.array(x_train), numpy.array(y_train), scaler_x, scaler_y
feature_length = dataframe.shape[1]
dataset = dataframe
dataset = dataset.astype('float32').as_matrix()
# split into train and test sets
train_size = int(len(dataset) * 0.67)
test_size = len(dataset) - train_size
train, test = dataset[0:train_size,:], dataset[train_size:len(dataset),:]
# reshape into X=t and Y=t+1
look_back = 1
trainX, trainY, scaler_trainX, scaler_trainY = create_dataset(train, look_back)
testX, testY, scaler_testX, scaler_testY = create_dataset(test, look_back)
# reshape input to be [samples, time steps, features]
trainX = numpy.reshape(trainX, (trainX.shape[0], 1, trainX.shape[1]))
testX = numpy.reshape(testX, (testX.shape[0], 1, testX.shape[1]))
# create and fit the LSTM network
model = Sequential()
model.add(LSTM(4, input_shape=(look_back, feature_length-2)))
model.add(Dense(2))
model.compile(loss='mean_squared_error', optimizer='adam', metrics=['mean_squared_error'])
h = model.fit(trainX, trainY, epochs=100, batch_size=10, verbose=2)
acc = h.history['acc']
epochs = range(1, len(acc) + 1)
plt.plot(epochs, acc, '-', label='Accuracy')
plt.legend()
plt.show()
# make predictions
trainPredict = model.predict(trainX)
testPredict = model.predict(testX)
# invert predictions
trainPredict = scaler_trainY.inverse_transform(trainPredict)
trainY = scaler_trainY.inverse_transform(trainY)
testPredict = scaler_testY.inverse_transform(testPredict)
testY = scaler_testY.inverse_transform(testY)
# calculate root mean squared error
trainScore = math.sqrt(mean_squared_error(trainY, trainPredict))
print('Train Score: %.2f RMSE' % (trainScore))
testScore = math.sqrt(mean_squared_error(testY, testPredict))
print('Test Score: %.2f RMSE' % (testScore))
# shift train predictions for plotting
trainPredictPlot = numpy.empty_like(trainPredict)
trainPredictPlot[:, :] = numpy.nan
trainPredictPlot = trainPredict
# shift test predictions for plotting
testPredictPlot = numpy.empty_like(dataset)
testPredictPlot[:, :] = numpy.nan
testPredictPlot = testPredict
# plot baseline and predictions
plt.plot(scaler_trainY.inverse_transform(trainPredict), alpha=0.5)
plt.plot(scaler_testY.inverse_transform(testPredict), alpha=0.5)
plt.show()
import tensorflow as tf
correct = tf.equal(tf.argmax(testPredict, 1), tf.argmax(testY, 1))
accuracy = tf.reduce_mean(tf.cast(correct, 'float'))
testY[:5]
testPredict[:5]
model.save("model.h5")
| Training.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
# # Running a differential expression (DE) analysis
# ## Introduction
#
# By default, DEAGO will run both a quality control (QC) and a differential expression (DE) analysis. DE analyses try to identify genes whose expression levels differ between experimental conditions. We don’t normally have enough replicates to do traditional tests of significance for RNA-Seq data. So, most methods look for outliers in the relationship between average abundance and fold change and assume most genes are not differentially expressed.
#
# Rather than just using a fold change threshold to determine which genes are differentially expressed, DEAs use a variety of statistical tests for significance. These tests give us a **p-value** which is an estimate of how often your observations would occur by chance.
#
# However, we perform these comparisons for each one of the thousands of genes/transcripts in our dataset. A p-value of 0.01 estimates a probability of 1% for seeing our observation just by chance. In an experiment like ours with 5,000 genes we would expect 5 genes to be significantly differentially expressed by chance (i.e. even if there were no difference between our conditions). Instead of using a p-value we can use an **adjusted p-value**, also known as the **q-value**, which accounts for the multiple testing and adjusts the p-value accordingly.
#
# The objectives of this part of the tutorial are:
#
# * run a DE analysis with DEAGO
# * interpret the output DE report from DEAGO
#
#
# ### Input files
#
# We will need to give DEAGO two bits of information:
#
# * *the name/location of the directory containing our gene count files (counts)*
#
#
# * *the name/location of our sample/condition mapping file (targets.txt)*
#
# We can optionally give DEAGO a formatted annotation file which contains gene names. These are often more recognisable than the unique gene identifiers found in the counts files. To do this, we use the `-a` option.
#
# ### Running a DE analysis with DEAGO
#
# To run a quick, DE analysis with DEAGO the command would be:
#
# ```
# deago -c <counts_directory> -t <targets file>
# ```
#
# As our count files were generated by featureCounts for this tutorial, we need to also tell DEAGO the count format with the `--count_type` option:
#
# ```
# deago -c <counts_directory> -t <targets file> --count_type featurecounts
# ```
#
# As we want to have the gene names in our output tables and plots, we need to provide our formatted annotation file using the `-a` option.
#
#
# Finally, we will be using the `--control` option which tells DEAGO the condition you want to use as your reference or control, in this case **WT_Ctrl**. We use **`--control`** to define our reference condition because, by default, R chooses the reference condition based on alphabetical order. It would assume that from our four conditions (**KO_Ctrl**, **KO_IL22**, **WT_Ctrl** and **WT_IL22**) that KO_Ctrl is our reference condition because it is first alphabetically. The value you use **must** be in the condition column in your targets file and is _case insensitive_.
#
#
# ```
# deago -c <counts directory> -t <targets file> --count_type featurecounts \
# -a <annotation file> --control <control>
# ```
#
# DEAGO also makes an assumption that you want the FDR cutoff (alpha) to be **0.05** (default). If you are expecting to use a different cutoff in your downstream filtering, use the **`-q`** option to define the FDR cutoff (e.g. -q 0.01).
#
#
# ### Output files/directories
#
# Once your DE analysis has finished, you should see several new files and directories:
#
# * **`deago.config`**
# _config file with key/value parameters defining the analysis_
#
#
# * **`deago.rlog`**
# _log of the R output generated when converting the R markdown to HTML_
#
#
# * **`deago_markdown.Rmd`**
# _R markdown used to run the analysis_
#
#
# * **`deago_markdown.html`**
# _HTML report generated from the R markdown_
#
#
# * **`results_<timestammp>`**
# _directory containing unfiltered DE analysis results and normalised counts for all genes, one file per contrast_
#
# #### Results directory
#
# The report tables are limited to genes with an adjusted p-value < 0.01 and a log2 fold change >= 2 or <= -2. However, you are likely to want to explore and filter these results using different thresholds. So, DEAGO also writes the unfiltered results table containing all genes to individual files, one per contrast in your timestamped results directory.
#
# So, for the full results of the contrast between WT and KO cells treated with IL22 you would look at:
#
# ```
# results_<timestamp>/wt_il22_vs_ko_il22_q0.05.txt
# ```
#
#
# ### DE analysis report
#
# The output file we're interested in is **`deago_markdown.html`** which is your DE analysis report. Go ahead and open it in a web browser (e.g. Chrome, Firefox, IE, Safari...). You can do this by going to "File -> Open" in the top navigation or (if you have Firefox installed, use the command:
firefox deago_markdown.html
# In addition to the QC sections we saw before, you should now see a new option in the left-hand sidebar called **`Pairwise contrasts`**. Click on it and it will take you to your DE analysis results.
#
# First there is a **`Contrast summary`** section which contains a summary table showing how many genes are up-regulated or down-regulated in each contrast (comparison between two sample groups). We can see that there were no differentially expressed (DE) genes between the knockout (**KO**) samples induced with IL22 (**ko_il22**) and the control knockout samples. However, there were 860 DE genes between wildtype (**WT**) and knockout (**KO**) samples induced with IL22, 510 up-regulated in the WT samples compared to the KO samples and 350 down-regulated.
# 
# If there are 2-4 contrasts in the analysis, there will also be a Venn diagram showing the overlap/differences in total DE genes between contrasts. We have 6 contrasts in this analysis, so no Venn diagram was generated, but an example would be:
# 
# The DE analysis report then has a series of subsections, one per contrast. Each contrast section has an MA plot and a volcano plot. The top 5 up- and down-regulated gene identifiers are labelled on plots. If an annotation with gene symbols was used then the point labels will be the gene symbols and not the gene identifiers.
# 
# Each contrast section will also have a DE results table which contains genes with an adjusted p-value < 0.01 and a log2 fold change >= 2 or <= -2. This is to reduce the number of genes in the table so that the HTML report is compact and sharable.
# 
# These tables contain the DESeq2 results and are where you will find your adjusted p-values and log2 fold change values. Also, as our gene identifiers are Ensembl identifiers, they have been converted to a link and if you click on one, it will take you to the current Ensembl page for that gene stable ID. In this example, we had include an annotation in the analysis and so the gene symbols are also shown.
#
# All of the tables are interactive and can be searched or filtered. The paper describes the up-regulation of _Fut2_ by IL-22RA1 signalling, so let's take a look. The search box at the top right searches the whole table, so we can use it to search for any _Fut_ genes.
# 
# This gives us two genes: _Fut2_ and _Fut9_.
#
# Now, say we wanted to only see the up-regulated _Fut_ genes. We can limit searches and filters to a single column by using the search/filter boxes at the top of each column. Use the selector at the top of the **`log2FoldChange`** column to only include values greater than 0 (i.e. drag the left selector).
# 
# We are now left with the only up-regulated _Fut_ gene, _Fut2_.
# ## Exercise 4
#
# **First, let's make sure we're in the `data` directory.**
cd data
# **Each DEAGO analysis should be self-contained, so let's create a new directory for our DE analysis.**
mkdir de_analysis
cd de_analysis
# **Now, let's get our DE report.**
deago --build_config -c ../counts -t ../targets.txt \
--count_type featurecounts \
-a ../ensembl_mm10_deago_formatted.tsv \
--control WT_Ctrl
# ## Questions
#
# In [Figure 5C](https://www.ncbi.nlm.nih.gov/pubmed/25263220) (below), the authors have highlighted four genes: ***Fut2***, ***Sec1***, ***Fut8*** and ***B4galt1*** which are associated with glycosylation.
#
# 
#
# **We have already found _Fut2_, answer the following questions for the remaining 3 genes using the contrast results for WT and KO cells treated with IL22.**
#
# **Q1: What is the gene identifier (geneID)?**
#
# **Q2: What is the log2 fold change?**
#
# **Q3: What is the adjusted p-value?**
#
# _Hint: you may want to use `awk` to look at columns 36 and 40 in the unfiltered results files for that contrast if the genes are not found in the report tables_
# ## What's next?
#
# If you want a recap of input file preparation, head back to [running a quality control (QC) analysis](quality-control.ipynb).
#
# Otherwise, let's continue on to [running a GO term enrichment analysis](go-term-enrichment.ipynb).
| practical/Notebooks/differential-expression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf2
# language: python
# name: tf2
# ---
# +
import os, sys
sys.path.append('..')
sys.path.append('../../')
import random
import pandas as pd
import numpy as np
import json
from sklearn.model_selection import train_test_split
import tensorflow as tf
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
from tensorflow.python.keras.preprocessing.sequence import pad_sequences
import matplotlib.pyplot as plt
import glob
import config as conf
# -
DATA_PATH = './tmp/train'
read_file_lst = glob.glob(conf.data_root + 'read/*')
exclude_file_lst = ['read.tar']
read_df_lst = []
for f in read_file_lst:
file_name = os.path.basename(f)
if file_name in exclude_file_lst:
print(file_name)
else:
df_temp = pd.read_csv(f, header=None, names=['raw'])
df_temp['dt'] = file_name[:8]
df_temp['hr'] = file_name[8:10]
df_temp['user_id'] = df_temp['raw'].str.split(' ').str[0]
df_temp['article_id'] = df_temp['raw'].str.split(' ').str[1:].str.join(' ').str.strip()
read_df_lst.append(df_temp)
read = pd.concat(read_df_lst)
read.head()
read.drop('raw', inplace=True, axis=1)
read['cnt'] = read['article_id'].apply(lambda x: len(x.split(' ')))
read_new = read[read.cnt >= 20]
len(read_new['user_id'].unique())
| preprocessing/truncate_train_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# Copyright (c) 2020-2021 Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
#
# # AutoML with FLAML Library
#
#
# ## 1. Introduction
#
# FLAML is a Python library (https://github.com/microsoft/FLAML) designed to automatically produce accurate machine learning models
# with low computational cost. It is fast and cheap. The simple and lightweight design makes it easy
# to use and extend, such as adding new learners. FLAML can
# - serve as an economical AutoML engine,
# - be used as a fast hyperparameter tuning tool, or
# - be embedded in self-tuning software that requires low latency & resource in repetitive
# tuning tasks.
#
# In this notebook, we use one real data example (binary classification) to showcase how to use FLAML library.
#
# FLAML requires `Python>=3.6`. To run this notebook example, please install flaml with the `notebook` option:
# ```bash
# pip install flaml[notebook]
# ```
# -
# !pip install flaml[notebook];
# + [markdown] slideshow={"slide_type": "slide"}
# ## 2. Classification Example
# ### Load data and preprocess
#
# Download [Airlines dataset](https://www.openml.org/d/1169) from OpenML. The task is to predict whether a given flight will be delayed, given the information of the scheduled departure.
# + slideshow={"slide_type": "subslide"} tags=[]
from flaml.data import load_openml_dataset
X_train, X_test, y_train, y_test = load_openml_dataset(dataset_id = 1169, data_dir = './')
# + [markdown] slideshow={"slide_type": "slide"}
# ### Run FLAML
# In the FLAML automl run configuration, users can specify the task type, time budget, error metric, learner list, whether to subsample, resampling strategy type, and so on. All these arguments have default values which will be used if users do not provide them. For example, the default ML learners of FLAML are `['lgbm', 'xgboost', 'catboost', 'rf', 'extra_tree', 'lrl1']`.
# + slideshow={"slide_type": "slide"}
''' import AutoML class from flaml package '''
from flaml import AutoML
automl = AutoML()
# + slideshow={"slide_type": "slide"}
settings = {
"time_budget": 300, # total running time in seconds
"metric": 'accuracy', # primary metrics can be chosen from: ['accuracy','roc_auc','f1','log_loss','mae','mse','r2']
"task": 'classification', # task type
"log_file_name": 'airlines_experiment.log', # flaml log file
}
# + slideshow={"slide_type": "slide"} tags=[]
'''The main flaml automl API'''
automl.fit(X_train = X_train, y_train = y_train, **settings)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Best model and metric
# + slideshow={"slide_type": "slide"} tags=[]
''' retrieve best config and best learner'''
print('Best ML leaner:', automl.best_estimator)
print('Best hyperparmeter config:', automl.best_config)
print('Best accuracy on validation data: {0:.4g}'.format(1-automl.best_loss))
print('Training duration of best run: {0:.4g} s'.format(automl.best_config_train_time))
# + slideshow={"slide_type": "slide"}
automl.model
# + slideshow={"slide_type": "slide"}
''' pickle and save the best model '''
import pickle
with open('best_model.pkl', 'wb') as f:
pickle.dump(automl.model, f, pickle.HIGHEST_PROTOCOL)
# + slideshow={"slide_type": "slide"} tags=[]
''' compute predictions of testing dataset '''
y_pred = automl.predict(X_test)
print('Predicted labels', y_pred)
print('True labels', y_test)
y_pred_proba = automl.predict_proba(X_test)[:,1]
# + slideshow={"slide_type": "slide"} tags=[]
''' compute different metric values on testing dataset'''
from flaml.ml import sklearn_metric_loss_score
print('accuracy', '=', 1 - sklearn_metric_loss_score('accuracy', y_pred, y_test))
print('roc_auc', '=', 1 - sklearn_metric_loss_score('roc_auc', y_pred_proba, y_test))
print('log_loss', '=', sklearn_metric_loss_score('log_loss', y_pred_proba, y_test))
print('f1', '=', 1 - sklearn_metric_loss_score('f1', y_pred, y_test))
# + [markdown] slideshow={"slide_type": "slide"}
# See Section 4 for an accuracy comparison with default LightGBM and XGBoost.
#
# ### Log history
# + slideshow={"slide_type": "subslide"} tags=[]
from flaml.data import get_output_from_log
time_history, best_valid_loss_history, valid_loss_history, config_history, train_loss_history = \
get_output_from_log(filename = settings['log_file_name'], time_budget = 60)
for config in config_history:
print(config)
# + slideshow={"slide_type": "slide"}
import matplotlib.pyplot as plt
import numpy as np
plt.title('Learning Curve')
plt.xlabel('Wall Clock Time (s)')
plt.ylabel('Validation Accuracy')
plt.scatter(time_history, 1-np.array(valid_loss_history))
plt.step(time_history, 1-np.array(best_valid_loss_history), where='post')
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ## 3. Customized Learner
# + [markdown] slideshow={"slide_type": "slide"}
# Some experienced automl users may have a preferred model to tune or may already have a reasonably by-hand-tuned model before launching the automl experiment. They need to select optimal configurations for the customized model mixed with standard built-in learners.
#
# FLAML can easily incorporate customized/new learners (preferably with sklearn API) provided by users in a real-time manner, as demonstrated below.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Example of Regularized Greedy Forest
#
# [Regularized Greedy Forest](https://arxiv.org/abs/1109.0887) (RGF) is a machine learning method currently not included in FLAML. The RGF has many tuning parameters, the most critical of which are: `[max_leaf, n_iter, n_tree_search, opt_interval, min_samples_leaf]`. To run a customized/new learner, the user needs to provide the following information:
# * an implementation of the customized/new learner
# * a list of hyperparameter names and types
# * rough ranges of hyperparameters (i.e., upper/lower bounds)
# * choose initial value corresponding to low cost for cost-related hyperparameters (e.g., initial value for max_leaf and n_iter should be small)
#
# In this example, the above information for RGF is wrapped in a python class called *MyRegularizedGreedyForest* that exposes the hyperparameters.
# + slideshow={"slide_type": "slide"}
''' SKLearnEstimator is the super class for a sklearn learner '''
from flaml.model import SKLearnEstimator
from flaml import tune
from rgf.sklearn import RGFClassifier, RGFRegressor
class MyRegularizedGreedyForest(SKLearnEstimator):
def __init__(self, task = 'binary:logistic', n_jobs = 1, **params):
'''Constructor
Args:
task: A string of the task type, one of
'binary:logistic', 'multi:softmax', 'regression'
n_jobs: An integer of the number of parallel threads
params: A dictionary of the hyperparameter names and values
'''
super().__init__(task, **params)
'''task=regression for RGFRegressor;
binary:logistic and multiclass:softmax for RGFClassifier'''
if 'regression' in task:
self.estimator_class = RGFRegressor
else:
self.estimator_class = RGFClassifier
# convert to int for integer hyperparameters
self.params = {
"n_jobs": n_jobs,
'max_leaf': int(params['max_leaf']),
'n_iter': int(params['n_iter']),
'n_tree_search': int(params['n_tree_search']),
'opt_interval': int(params['opt_interval']),
'learning_rate': params['learning_rate'],
'min_samples_leaf':int(params['min_samples_leaf'])
}
@classmethod
def search_space(cls, data_size, task):
'''[required method] search space
Returns:
A dictionary of the search space.
Each key is the name of a hyperparameter, and value is a dict with
its domain and init_value (optional), cat_hp_cost (optional)
e.g.,
{'domain': tune.randint(lower=1, upper=10), 'init_value': 1}
'''
space = {
'max_leaf': {'domain': tune.qloguniform(lower = 4, upper = data_size, q = 1), 'init_value': 4},
'n_iter': {'domain': tune.qloguniform(lower = 1, upper = data_size, q = 1), 'init_value': 1},
'n_tree_search': {'domain': tune.qloguniform(lower = 1, upper = 32768, q = 1), 'init_value': 1},
'opt_interval': {'domain': tune.qloguniform(lower = 1, upper = 10000, q = 1), 'init_value': 100},
'learning_rate': {'domain': tune.loguniform(lower = 0.01, upper = 20.0)},
'min_samples_leaf': {'domain': tune.qloguniform(lower = 1, upper = 20, q = 1), 'init_value': 20},
}
return space
@classmethod
def size(cls, config):
'''[optional method] memory size of the estimator in bytes
Args:
config - the dict of the hyperparameter config
Returns:
A float of the memory size required by the estimator to train the
given config
'''
max_leaves = int(round(config['max_leaf']))
n_estimators = int(round(config['n_iter']))
return (max_leaves*3 + (max_leaves-1)*4 + 1.0)*n_estimators*8
@classmethod
def cost_relative2lgbm(cls):
'''[optional method] relative cost compared to lightgbm
'''
return 1.0
# + [markdown] slideshow={"slide_type": "slide"}
# ### Add Customized Learner and Run FLAML AutoML
#
# After adding RGF into the list of learners, we run automl by tuning hyperpameters of RGF as well as the default learners.
# + slideshow={"slide_type": "slide"}
automl = AutoML()
automl.add_learner(learner_name = 'RGF', learner_class = MyRegularizedGreedyForest)
# + slideshow={"slide_type": "slide"} tags=[]
settings = {
"time_budget": 60, # total running time in seconds
"metric": 'accuracy',
"estimator_list": ['RGF', 'lgbm', 'rf', 'xgboost'], # list of ML learners
"task": 'classification', # task type
"log_file_name": 'airlines_experiment_custom.log', # flaml log file
"log_training_metric": True, # whether to log training metric
}
'''The main flaml automl API'''
automl.fit(X_train = X_train, y_train = y_train, **settings)
# -
# ## 4. Comparison with alternatives
#
# ### FLAML's accuracy
print('flaml accuracy', '=', 1 - sklearn_metric_loss_score('accuracy', y_pred, y_test))
# ### Default LightGBM
from lightgbm import LGBMClassifier
lgbm = LGBMClassifier()
lgbm.fit(X_train, y_train)
y_pred = lgbm.predict(X_test)
from flaml.ml import sklearn_metric_loss_score
print('default lgbm accuracy', '=', 1 - sklearn_metric_loss_score('accuracy', y_pred, y_test))
# ### Default XGBoost
from xgboost import XGBClassifier
xgb = XGBClassifier()
xgb.fit(X_train, y_train)
y_pred = xgb.predict(X_test)
from flaml.ml import sklearn_metric_loss_score
print('default xgboost accuracy', '=', 1 - sklearn_metric_loss_score('accuracy', y_pred, y_test))
| notebook/flaml_automl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Course Human-Centered Data Science ([HCDS](https://www.mi.fu-berlin.de/en/inf/groups/hcc/teaching/winter_term_2020_21/course_human_centered_data_science.html)) - Winter Term 2020/21 - [HCC](https://www.mi.fu-berlin.de/en/inf/groups/hcc/index.html) | [Freie Universität Berlin](https://www.fu-berlin.de/)
#
# ***
#
# # A2 - Wikipedia, ORES, and Bias in Data
#
# ## Step 1⃣ | Data acquisition
#
# Two data sources are used: (1) Wikipedia articles of politicians and (2) world population data.
#
# **Wikipedia articles -**
# The Wikipedia articles can be found on [Figshare](https://figshare.com/articles/Untitled_Item/5513449). It contains politiciaans by country from the English-language wikipedia. Please read through the documentation for this repository, then download and unzip it to extract the data file, which is called `page_data.csv`.
#
# **Population data -**
# The population data is available in `CSV` format in the `data_raw` folder. The file is named `export_2019.csv`. This dataset is drawn from the [world population datasheet](https://www.prb.org/international/indicator/population/table/) published by the Population Reference Bureau (downloaded 2020-11-13 10:14 AM). I have edited the dataset to make it easier to use in this assignment. The population per country is given in millions!
import pandas as pd
page = pd.read_csv("../data_raw/page_data.csv")
export = pd.read_csv("../data_raw/export_2019.csv", delimiter = ";")
page.head()
export.head()
# ## Step 2⃣ | Data processing and cleaning
# The data in `page_data.csv` contain some rows that you will need to filter out. It contains some page names that start with the string `"Template:"`. These pages are not Wikipedia articles, and should not be included in your analysis. The data in `export_2019.csv` does not need any cleaning.
#
page = page[~page['page'].str.contains("Template")]
page.head()
# ### Getting article quality predictions with ORES
#
# Now you need to get the predicted quality scores for each article in the Wikipedia dataset. We're using a machine learning system called [**ORES**](https://www.mediawiki.org/wiki/ORES) ("Objective Revision Evaluation Service"). ORES estimates the quality of an article (at a particular point in time), and assigns a series of probabilities that the article is in one of the six quality categories. The options are, from best to worst:
#
# | ID | Quality Category | Explanation |
# |----|------------------|----------|
# | 1 | FA | Featured article |
# | 2 | GA | Good article |
# | 3 | B | B-class article |
# | 4 | C | C-class article |
# | 5 | Start | Start-class article |
# | 6 | Stub | Stub-class article |
#
# ### ORES REST API endpoint
#
# The [ORES REST API](https://ores.wikimedia.org/v3/#!/scoring/get_v3_scores_context_revid_model) is configured fairly similarly to the pageviews API we used for the last assignment. It expects the following parameters:
#
# * **project** --> `enwiki`
# * **revid** --> e.g. `235107991` or multiple ids e.g.: `235107991|355319463` (batch)
# * **model** --> `wp10` - The name of a model to use when scoring.
#
# **❗Note on batch processing:** Please read the documentation about [API usage](https://www.mediawiki.org/wiki/ORES#API_usage) if you want to query a large number of revisions (batches).
#
# You will notice that ORES returns a prediction value that contains the name of one category (e.g. `Start`), as well as probability values for each of the six quality categories. For this assignment, you only need to capture and use the value for prediction.
#
# **❗Note:** It's possible that you will be unable to get a score for a particular article. If that happens, make sure to maintain a log of articles for which you were not able to retrieve an ORES score. This log should be saved as a separate file named `ORES_no_scores.csv` and should include the `page`, `country`, and `rev_id` (just as in `page_data.csv`).
#
# It's recommended to batch no more than 50 revisions within a given request. The dataframe is split into equal chunks of size 50.
# +
import requests
import json
import numpy as np
import sys
from math import floor
# Customize these with your own information
headers = {
'User-Agent': 'https://github.com/Francosinus',
'From': '<EMAIL>'
}
def get_ores_data(rev_id,headers):
# Define the endpoint
# https://ores.wikimedia.org/scores/enwiki/?models=wp10&revids=807420979|807422778
endpoint = 'https://ores.wikimedia.org/v3/scores/{project}/?models={model}&revids={revids}'
params = {'project' : 'enwiki',
'model' : 'wp10',
'revids' : rev_id
}
api_call = requests.get(endpoint.format(**params))
response = api_call.json()
data=json.dumps(response)
return data
def createDf(ids,df):
prediction=[]
err=[]
for i in range(0, len(ids) // 50+1):
if i %50 ==0:
print(f"Processing batch: {i} of {len(ids) // 50+1}")
batch = '|'.join(str(x) for x in ids[(i * 50):((i+1) * 50)])
data = get_ores_data(batch,headers)
data = json.loads(data)
for key in data['enwiki']['scores'].keys():
try:
pred = data['enwiki']['scores'][str(key)]['wp10']['score']['prediction']
prediction.append([key, pred])
except KeyError:
err.append(key)
ores = pd.DataFrame(prediction, columns = ["rev_id","prediction"])
ores['rev_id']=ores['rev_id'].astype(int)
full = df.merge(ores, on=['rev_id'], how='left')
return full
# +
ids = [item for item in page['rev_id']]
page_ores = createDf(ids, page)
# -
# **Let's have a look at the data! We can see that the table now contains a column prediction with refers to the quality scores of the articles**
page_ores.head()
# ### Combining the datasets
#
# Both datasets are combined: (1) the wikipedia articles with ORES scores (2) the population data. Both have columns named `country`. Not all entries can be merged, so the non matching rows are filtered out. This can be easily done by removing rows which contain NaN values.
#
# The non matching data can be found in `../data_clean/countries-no_match.csv`. The remaining data can be found in `../data_clean/politicians_by_country.csv`.
#
df = pd.merge(page_ores, export, on='country', how='outer')
df.head()
# **To make further processing easier, the columns are renamed.**
df['population']=df['population']*1e6
df=df.rename(columns={"page": "article_name", "rev_id": "revision_id", "prediction":"article_quality"})
no_match = df[df.isnull().any(axis=1)]
match = df[~df.isnull().any(axis=1)]
no_match.to_csv("../data_clean/countries-no_match.csv")
match.to_csv("../data_clean/politicians_by_country.csv")
# ## Step 3⃣ | Analysis
#
# The analysis consists of calculating the proportion (as a percentage) of articles-per-population and high-quality articles for **each country** and for **each region**. By `"high quality"` arcticle I mean an article that ORES predicted as `FA` (featured article) or `GA` (good article).
#
# **Examples:**
#
# * if a country has a population of `10,000` people, and you found `10` articles about politicians from that country, then the percentage of `articles-per-population` would be `0.1%`.
# * if a country has `10` articles about politicians, and `2` of them are `FA` or `GA` class articles, then the percentage of `high-quality-articles` would be `20%`.
# ### Results
#
# The results from this analysis are six `data tables`.
#
# First we read the cleaned table in again to further process it.
clean = pd.read_csv("../data_clean/politicians_by_country.csv")
clean.head()
# **Table 1:**
#
# **Top 10 countries by coverage**<br>10 highest-ranked countries in terms of number of politician articles as a proportion of country population
top_coverage = clean.groupby(['country'])
top_coverage = top_coverage.agg({'article_name':'count','population':'last'})
top_coverage = top_coverage.reset_index()
top_coverage['coverage'] = (top_coverage['article_name']*100) / (top_coverage['population'])
top_coverage = top_coverage.sort_values('coverage', ascending=False)
top_coverage = top_coverage.reset_index(drop=True)
top_coverage.head(10)
top_coverage[1:11].to_csv("../results/country_coverage_data_top_10.csv")
# **Table 2:**
#
# **Bottom 10 countries by coverage**<br>10 lowest-ranked countries in terms of number of politician articles as a proportion of country population
top_coverage = top_coverage.sort_values('coverage', ascending=True)
top_coverage.head(10)
top_coverage[1:11].to_csv("../results/country_coverage_data_bottom_10.csv")
# **Table 3:**
#
# **Regions by coverage**<br>Ranking of regions (in descending order) in terms of the total count of politician articles from countries in each region as a proportion of total regional population
top_coverag = clean.groupby(['region'])
top_coverag = top_coverag.agg({'article_name':'count','population':'sum'})
top_coverag = top_coverag.reset_index()
top_coverag['coverage'] = (top_coverag['article_name']*100) / (top_coverag['population'])
top_coverag = top_coverag.sort_values('coverage', ascending=False)
top_coverag = top_coverag.reset_index(drop=True)
top_coverag.head(10)
top_coverag[1:11].to_csv("../results/region_coverage_data_top.csv")
# **Table 4: **
# **Top 10 countries by relative quality**<br>10 highest-ranked countries in terms of the relative proportion of politician articles that are of GA and FA-quality
rel = clean.loc[(clean.article_quality =='GA') | (clean.article_quality =='FA')]
relative = rel.groupby(['country'])
relative = relative.agg({'article_name':'count','population':'last'})
relative = relative.reset_index()
relative['rel_proportion'] = (relative['article_name']*100) / (relative['population'])
relative = relative.sort_values('rel_proportion', ascending=False)
relative = relative.reset_index(drop=True)
relative.head(10)
relative[1:11].to_csv("../results/country_proportion_good_quality_data_top10.csv")
# **Table 5:**
#
# **Bottom 10 countries by relative quality**<br>10 lowest-ranked countries in terms of the relative proportion of politician articles that are of GA and FA-quality
relative = relative.sort_values('rel_proportion', ascending=True)
relative[1:11].to_csv("../results/country_proportion_good_quality_data_bottom10.csv")
relative.head(10)
# **Table 6:**
#
# **Regions by coverage**<br>Ranking of regions (in descending order) in terms of the relative proportion of politician articles from countries in each region that are of GA and FA-quality
relative = rel.groupby(['region'])
relative = relative.agg({'article_name':'count','population':'sum'})
relative = relative.reset_index()
relative['rel_proportion'] = (relative['article_name']*100) / (relative['population'])
relative = relative.sort_values('rel_proportion', ascending=False)
relative = relative.reset_index(drop=True)
relative.head(10)
relative[1:11].to_csv("../results/region_proportion_good_quality_data_top.csv")
# ***
#
# #### Credits
#
# This exercise is slighty adapted from the course [Human Centered Data Science (Fall 2019)](https://wiki.communitydata.science/Human_Centered_Data_Science_(Fall_2019)) of [Univeristy of Washington](https://www.washington.edu/datasciencemasters/) by [<NAME>](https://wiki.communitydata.science/User:Jtmorgan).
#
# Same as the original inventors, we release the notebooks under the [Creative Commons Attribution license (CC BY 4.0)](https://creativecommons.org/licenses/by/4.0/).
| src/A3_Wikipedia_ORES_Bias.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="FnKryrWjZqNZ"
import pandas as pd
# + colab={"base_uri": "https://localhost:8080/"} id="ICryGzaRZsXE" outputId="9d4ee463-04ea-4c88-d8f4-5388f8e441c1"
# !ls ./smtph_total.csv
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="IcswWXbIZ74M" outputId="3a1c786d-cc0a-466b-b9fe-c9931738c42d"
df = pd.read_csv('./drive/MyDrive/Test/smtph_total.csv')
df.head(5)
# + colab={"base_uri": "https://localhost:8080/", "height": 71} id="x2LG9rIyaCQU" outputId="7fa19f3f-2433-4175-aae4-d96a9883ce6a"
posts = df['Description']
posts[0]
# + colab={"base_uri": "https://localhost:8080/"} id="Nexh7iTwax3U" outputId="90ba3d12-86d1-4d3e-ceb2-8e28d5fc8a38"
# !python -m pip install konlpy
# + colab={"base_uri": "https://localhost:8080/"} id="m7S94JhFa_uE" outputId="94f9eabb-5b2c-4066-9b91-f6df170858a6"
# !python -m pip install eunjeon
# + id="nQlxYjkibQgf"
# + colab={"base_uri": "https://localhost:8080/", "height": 520} id="-tISoBFlabuL" outputId="d450e4fe-e24e-4e6f-d511-78ff894d77ad"
from konlpy.tag import Mecab
tagger = Mecab()
# + id="icfaLV4GawUr"
| NLTK_korean.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Release the Kraken!
# +
# The next library we're going to look at is called Kraken, which was developed by Université
# PSL in Paris. It's actually based on a slightly older code base, OCRopus. You can see how the
# flexible open-source licenses allow new ideas to grow by building upon older ideas. And, in
# this case, I fully support the idea that the Kraken - a mythical massive sea creature - is the
# natural progression of an octopus!
#
# What we are going to use Kraken for is to detect lines of text as bounding boxes in a given
# image. The biggest limitation of tesseract is the lack of a layout engine inside of it. Tesseract
# expects to be using fairly clean text, and gets confused if we don't crop out other artifacts.
# It's not bad, but Kraken can help us out be segmenting pages. Lets take a look.
# -
# First, we'll take a look at the kraken module itself
import kraken
help(kraken)
# There isn't much of a discussion here, but there are a number of sub-modules that look
# interesting. I spend a bit of time on their website, and I think the pageseg module, which
# handles all of the page segmentation, is the one we want to use. Lets look at it
from kraken import pageseg
help(pageseg)
# So it looks like there are a few different functions we can call, and the segment
# function looks particularly appropriate. I love how expressive this library is on the
# documentation front -- I can see immediately that we are working with PIL.Image files,
# and the author has even indicated that we need to pass in either a binarized (e.g. '1')
# or grayscale (e.g. 'L') image. We can also see that the return value is a dictionary
# object with two keys, "text_direction" which will return to us a string of the
# direction of the text, and "boxes" which appears to be a list of tuples, where each
# tuple is a box in the original image.
#
# Lets try this on the image of text. I have a simple bit of text in a file called
# two_col.png which is from a newspaper on campus here
from PIL import Image
im=Image.open("readonly/two_col.png")
# Lets display the image inline
display(im)
# Lets now convert it to black and white and segment it up into lines with kraken
bounding_boxes=pageseg.segment(im.convert('1'))['boxes']
# And lets print those lines to the screen
print(bounding_boxes)
# +
# Ok, pretty simple two column text and then a list of lists which are the bounding boxes of
# lines of that text. Lets write a little routine to try and see the effects a bit more
# clearly. I'm going to clean up my act a bit and write real documentation too, it's a good
# practice
def show_boxes(img):
'''Modifies the passed image to show a series of bounding boxes on an image as run by kraken
:param img: A PIL.Image object
:return img: The modified PIL.Image object
'''
# Lets bring in our ImageDraw object
from PIL import ImageDraw
# And grab a drawing object to annotate that image
drawing_object=ImageDraw.Draw(img)
# We can create a set of boxes using pageseg.segment
bounding_boxes=pageseg.segment(img.convert('1'))['boxes']
# Now lets go through the list of bounding boxes
for box in bounding_boxes:
# An just draw a nice rectangle
drawing_object.rectangle(box, fill = None, outline ='red')
# And to make it easy, lets return the image object
return img
# To test this, lets use display
display(show_boxes(Image.open("readonly/two_col.png")))
# +
# Not bad at all! It's interesting to see that kraken isn't completely sure what to do with this
# two column format. In some cases, kraken has identified a line in just a single column, while
# in other cases kraken has spanned the line marker all the way across the page. Does this matter?
# Well, it really depends on our goal. In this case, I want to see if we can improve a bit on this.
#
# So we're going to go a bit off script here. While this week of lectures is about libraries, the
# goal of this last course is to give you confidence that you can apply your knowledge to actual
# programming tasks, even if the library you are using doesn't quite do what you want.
#
# I'd like to pause the video for the moment and collect your thoughts. Looking at the image above,
# with the two column example and red boxes, how do you think we might modify this image to improve
# kraken's ability to text lines?
# -
# Thanks for sharing your thoughts, I'm looking forward to seeing the breadth of ideas that everyone
# in the course comes up with. Here's my partial solution -- while looking through the kraken docs on
# the pageseg() function I saw that there are a few parameters we can supply in order to improve
# segmentation. One of these is the black_colseps parameter. If set to True, kraken will assume that
# columns will be separated by black lines. This isn't our case here, but, I think we have all of the
# tools to go through and actually change the source image to have a black separator between columns.
#
# The first step is that I want to update the show_boxes() function. I'm just going to do a quick
# # copy and paste from the above but add in the black_colseps=True parameter
def show_boxes(img):
'''Modifies the passed image to show a series of bounding boxes on an image as run by kraken
:param img: A PIL.Image object
:return img: The modified PIL.Image object
'''
# Lets bring in our ImageDraw object
from PIL import ImageDraw
# And grab a drawing object to annotate that image
drawing_object=ImageDraw.Draw(img)
# We can create a set of boxes using pageseg.segment
#
bounding_boxes=pageseg.segment(img.convert('1'), black_colseps=True)['boxes']
# Now lets go through the list of bounding boxes
for box in bounding_boxes:
# An just draw a nice rectangle
drawing_object.rectangle(box, fill = None, outline ='red')
# And to make it easy, lets return the image object
return img
# +
# The next step is to think of the algorithm we want to apply to detect a white column separator.
# In experimenting a bit I decided that I only wanted to add the separator if the space of was
# at least 25 pixels wide, which is roughly the width of a character, and six lines high. The
# width is easy, lets just make a variable
char_width=25
# The height is harder, since it depends on the height of the text. I'm going to write a routine
# to calculate the average height of a line
def calculate_line_height(img):
'''Calculates the average height of a line from a given image
:param img: A PIL.Image object
:return: The average line height in pixels
'''
# Lets get a list of bounding boxes for this image
bounding_boxes=pageseg.segment(img.convert('1'))['boxes']
# Each box is a tuple of (top, left, bottom, right) so the height is just top - bottom
# So lets just calculate this over the set of all boxes
height_accumulator=0
for box in bounding_boxes:
height_accumulator=height_accumulator+box[3]-box[1]
# this is a bit tricky, remember that we start counting at the upper left corner in PIL!
# now lets just return the average height
# lets change it to the nearest full pixel by making it an integer
return int(height_accumulator/len(bounding_boxes))
# And lets test this with the image with have been using
line_height=calculate_line_height(Image.open("readonly/two_col.png"))
print(line_height)
# -
# Ok, so the average height of a line is 31.
# Now, we want to scan through the image - looking at each pixel in turn - to determine if there
# is a block of whitespace. How bit of a block should we look for? That's a bit more of an art
# than a science. Looking at our sample image, I'm going to say an appropriate block should be
# one char_width wide, and six line_heights tall. But, I honestly just made this up by eyeballing
# the image, so I would encourage you to play with values as you explore.
# Lets create a new box called gap box that represents this area
gap_box=(0,0,char_width,line_height*6)
gap_box
# It seems we will want to have a function which, given a pixel in an image, can check to see
# if that pixel has whitespace to the right and below it. Essentially, we want to test to see
# if the pixel is the upper left corner of something that looks like the gap_box. If so, then
# we should insert a line to "break up" this box before sending to kraken
#
# Lets call this new function gap_check
def gap_check(img, location):
'''Checks the img in a given (x,y) location to see if it fits the description
of a gap_box
:param img: A PIL.Image file
:param location: A tuple (x,y) which is a pixel location in that image
:return: True if that fits the definition of a gap_box, otherwise False
'''
# Recall that we can get a pixel using the img.getpixel() function. It returns this value
# as a tuple of integers, one for each color channel. Our tools all work with binarized
# images (black and white), so we should just get one value. If the value is 0 it's a black
# pixel, if it's white then the value should be 255
#
# We're going to assume that the image is in the correct mode already, e.g. it has been
# binarized. The algorithm to check our bounding box is fairly easy: we have a single location
# which is our start and then we want to check all the pixels to the right of that location
# up to gap_box[2]
for x in range(location[0], location[0]+gap_box[2]):
# the height is similar, so lets iterate a y variable to gap_box[3]
for y in range(location[1], location[1]+gap_box[3]):
# we want to check if the pixel is white, but only if we are still within the image
if x < img.width and y < img.height:
# if the pixel is white we don't do anything, if it's black, we just want to
# finish and return False
if img.getpixel((x,y)) != 255:
return False
# If we have managed to walk all through the gap_box without finding any non-white pixels
# then we can return true -- this is a gap!
return True
# Alright, we have a function to check for a gap, called gap_check. What should we do once
# we find a gap? For this, lets just draw a line in the middle of it. Lets create a new function
def draw_sep(img,location):
'''Draws a line in img in the middle of the gap discovered at location. Note that
this doesn't draw the line in location, but draws it at the middle of a gap_box
starting at location.
:param img: A PIL.Image file
:param location: A tuple(x,y) which is a pixel location in the image
'''
# First lets bring in all of our drawing code
from PIL import ImageDraw
drawing_object=ImageDraw.Draw(img)
# next, lets decide what the middle means in terms of coordinates in the image
x1=location[0]+int(gap_box[2]/2)
# and our x2 is just the same thing, since this is a one pixel vertical line
x2=x1
# our starting y coordinate is just the y coordinate which was passed in, the top of the box
y1=location[1]
# but we want our final y coordinate to be the bottom of the box
y2=y1+gap_box[3]
drawing_object.rectangle((x1,y1,x2,y2), fill = 'black', outline ='black')
# and we don't have anything we need to return from this, because we modified the image
# +
# Now, lets try it all out. This is pretty easy, we can just iterate through each pixel
# in the image, check if there is a gap, then insert a line if there is.
def process_image(img):
'''Takes in an image of text and adds black vertical bars to break up columns
:param img: A PIL.Image file
:return: A modified PIL.Image file
'''
# we'll start with a familiar iteration process
for x in range(img.width):
for y in range(img.height):
# check if there is a gap at this point
if (gap_check(img, (x,y))):
# then update image to one which has a separator drawn on it
draw_sep(img, (x,y))
# and for good measure we'll return the image we modified
return img
# Lets read in our test image and convert it through binarization
i=Image.open("readonly/two_col.png").convert("L")
i=process_image(i)
display(i)
#Note: This will take some time to run! Be patient!
# -
# Not bad at all! The effect at the bottom of the image is a bit unexpected to me, but it makes
# sense. You can imagine that there are several ways we might try and control this. Lets see how
# this new image works when run through the kraken layout engine
display(show_boxes(i))
# +
# Looks like that is pretty accurate, and fixes the problem we faced. Feel free to experiment
# with different settings for the gap heights and width and share in the forums. You'll notice though
# method we created is really quite slow, which is a bit of a problem if we wanted to use
# this on larger text. But I wanted to show you how you can mix your own logic and work with
# libraries you're using. Just because Kraken didn't work perfectly, doesn't mean we can't
# build something more specific to our use case on top of it.
#
# I want to end this lecture with a pause and to ask you to reflect on the code we've written
# here. We started this course with some pretty simple use of libraries, but now we're
# digging in deeper and solving problems ourselves with the help of these libraries. Before we
# go on to our last library, how well prepared do you think you are to take your python
# skills out into the wild?
# -
# ## Comparing Image Data Structures
# +
# OpenCV supports reading of images in most file formats, such as JPEG, PNG, and TIFF. Most image and
# video analysis requires converting images into grayscale first. This simplifies the image and reduces
# noise allowing for improved analysis. Let's write some code that reads an image of as person, Floyd
# Mayweather and converts it into greyscale.
# First we will import the open cv package cv2
import cv2 as cv
# We'll load the floyd.jpg image
img = cv.imread('readonly/floyd.jpg')
# And we'll convert it to grayscale using the cvtColor image
gray = cv.cvtColor(img, cv.COLOR_BGR2GRAY)
# Now, before we get to the result, lets talk about docs. Just like tesseract, opencv is an external
# package written in C++, and the docs for python are really poor. This is unfortunatly quite common
# when python is being used as a wrapper. Thankfully, the web docs for opencv are actually pretty good,
# so hit the website docs.opencv.org when you want to learn more about a particular function. In this
# case cvtColor converts from one color space to another, and we are convering our image to grayscale.
# Of course, we already know at least two different ways of doing this, using binarization and PIL
# color spaces conversions
# Lets instpec this object that has been returned.
import inspect
inspect.getmro(type(gray))
# -
# We see that it is of type ndarray, which is a fundamental list type coming from the numerical
# python project. That's a bit surprising - up until this point we have been used to working with
# PIL.Image objects. OpenCV, however, wants to represent an image as a two dimensional sequence
# of bytes, and the ndarray, which stands for n dimensional array, is the ideal way to do this.
# Lets look at the array contents.
gray
# +
# The array is shown here as a list of lists, where the inner lists are filled with integers.
# The dtype=uint8 definition indicates that each of the items in an array is an 8 bit unsigned
# integer, which is very common for black and white images. So this is a pixel by pixel definition
# of the image.
#
# The display package, however, doesn't know what to do with this image. So lets convert it
# into a PIL object to render it in the browser.
from PIL import Image
# PIL can take an array of data with a given color format and convert this into a PIL object.
# This is perfect for our situation, as the PIL color mode, "L" is just an array of luminance
# values in unsigned integers
image = Image.fromarray(gray, "L")
display(image)
# +
# Lets talk a bit more about images for a moment. Numpy arrays are multidimensional. For
# instance, we can define an array in a single dimension:
import numpy as np
single_dim = np.array([25, 50 , 25, 10, 10])
# In an image, this is analagous to a single row of 5 pixels each in grayscale. But actually,
# all imaging libraries tend to expect at least two dimensions, a width and a height, and to
# show a matrix. So if we put the single_dim inside of another array, this would be a two
# dimensional array with element in the height direction, and five in the width direction
double_dim = np.array([single_dim])
double_dim
# -
# This should look pretty familiar, it's a lot like a list of lists! Lets see what this new
# two dimensional array looks like if we display it
display(Image.fromarray(double_dim, "L"))
# Pretty unexciting - it's just a little line. Five pixels in a row to be exact, of different
# levels of black. The numpy library has a nice attribute called shape that allows us to see how
# many dimensions big an array is. The shape attribute returns a tuple that shows the height of
# the image, by the width of the image
double_dim.shape
# Lets take a look at the shape of our initial image which we loaded into the img variable
img.shape
# This image has three dimensions! That's because it has a width, a height, and what's called
# a color depth. In this case, the color is represented as an array of three values. Lets take a
# look at the color of the first pixel
first_pixel=img[0][0]
first_pixel
# +
# Here we see that the color value is provided in full RGB using an unsigned integer. This
# means that each color can have one of 256 values, and the total number of unique colors
# that can be represented by this data is 256 * 256 *256 which is roughly 16 million colors.
# We call this 24 bit color, which is 8+8+8.
#
# If you find yourself shopping for a television, you might notice that some expensive models
# are advertised as having 10 bit or even 12 bit panels. These are televisions where each of
# the red, green, and blue color channels are represented by 10 or 12 bits instead of 8. For
# ten bit panels this means that there are 1 billion colors capable, and 12 bit panels are
# capable of over 68 billion colors!
# -
# We're not going to talk much more about color in this course, but it's a fun subject. Instead,
# lets go back to this array representation of images, because we can do some interesting things
# with this.
#
# One of the most common things to do with an ndarray is to reshape it -- to change the number
# of rows and columns that are represented so that we can do different kinds of operations.
# Here is our original two dimensional image
print("Original image")
print(gray)
# If we wanted to represent that as a one dimensional image, we just call reshape
print("New image")
# And reshape takes the image as the first parameter, and a new shape as the second
image1d=np.reshape(gray,(1,gray.shape[0]*gray.shape[1]))
print(image1d)
# So, why are we talking about these nested arrays of bytes, we were supposed to be talking
# about OpenCV as a library. Well, I wanted to show you that often libraries working on the
# same kind of principles, in this case images stored as arrays of bytes, are not representing
# data in the same way in their APIs. But, by exploring a bit you can learn how the internal
# representation of data is stored, and build routines to convert between formats.
#
# For instance, remember in the last lecture when we wanted to look for gaps in an image so
# that we could draw lines to feed into kraken? Well, we use PIL to do this, using getpixel()
# to look at individual pixels and see what the luminosity was, then ImageDraw.rectangle to
# actually fill in a black bar separator. This was a nice high level API, and let us write
# routines to do the work we wanted without having to understand too much about how the images
# were being stored. But it was computationally very slow.
#
# Instead, we could write the code to do this using matrix features within numpy. Lets take
# a look.
import cv2 as cv
# We'll load the 2 column image
img = cv.imread('readonly/two_col.png')
# And we'll convert it to grayscale using the cvtColor image
gray = cv.cvtColor(img, cv.COLOR_BGR2GRAY)
# Now, remember how slicing on a list works, if you have a list of number such as
# a=[0,1,2,3,4,5] then a[2:4] will return the sublist of numbers at position 2 through 4
# inclusive - don't forget that lists start indexing at 0!
# If we have a two dimensional array, we can slice out a smaller piece of that using the
# format a[2:4,1:3]. You can think of this as first slicing along the rows dimension, then
# in the columns dimension. So in this example, that would be a matrix of rows 2, and 3,
# and columns 1, and 2. Here's a look at our image.
gray[2:4,1:3]
# So we see that it is all white. We can use this as a "window" and move it around our
# our big image.
#
# Finally, the ndarray library has lots of matrix functions which are generally very fast
# to run. One that we want to consider in this case is count_nonzero(), which just returns
# the number of entries in the matrix which are not zero.
np.count_nonzero(gray[2:4,1:3])
# Ok, the last benefit of going to this low level approach to images is that we can change
# pixels very fast as well. Previously we were drawing rectangles and setting a fill and line
# width. This is nice if you want to do something like change the color of the fill from the
# line, or draw complex shapes. But we really just want a line here. That's really easy to
# do - we just want to change a number of luminosity values from 255 to 0.
#
# As an example, lets create a big white matrix
white_matrix=np.full((12,12),255,dtype=np.uint8)
display(Image.fromarray(white_matrix,"L"))
white_matrix
# looks pretty boring, it's just a giant white square we can't see. But if we want, we can
# easily color a column to be black
white_matrix[:,6]=np.full((1,12),0,dtype=np.uint8)
display(Image.fromarray(white_matrix,"L"))
white_matrix
# +
# And that's exactly what we wanted to do. So, why do it this way, when it seems so much
# more low level? Really, the answer is speed. This paradigm of using matricies to store
# and manipulate bytes of data for images is much closer to how low level API and hardware
# developers think about storing files and bytes in memory.
#
# How much faster is it? Well, that's up to you to discover; there's an optional assignment
# for this week to convert our old code over into this new format, to compare both the
# readability and speed of the two different approaches.
# -
# ## OpenCV
# Ok, we're just about at the project for this course. If you reflect on the specialization
# as a whole you'll realize that you started with probably little or no understanding of python,
# progressed through the basic control structures and libraries included with the language
# with the help of a digital textbook, moved on to more high level representations of data
# and functions with objects, and now started to explore third party libraries that exist for
# python which allow you to manipulate and display images. This is quite an achievement!
#
# You have also no doubt found that as you have progressed the demands on you to engage in self-
# discovery have also increased. Where the first assignments were maybe straight forward, the
# ones in this week require you to struggle a bit more with planning and debugging code as
# you develop.
#
# But, you've persisted, and I'd like to share with you just one more set of features before
# we head over to a project. The OpenCV library contains mechanisms to do face detection on
# images. The technique used is based on Haar cascades, which is a machine learning approach.
# Now, we're not going to go into the machine learning bits, we have another specialization on
# Applied Data Science with Python which you can take after this if you're interested in that topic.
# But here we'll treat OpenCV like a black box.
#
# OpenCV comes with trained models for detecting faces, eyes, and smiles which we'll be using.
# You can train models for detecting other things - like hot dogs or flutes - and if you're
# interested in that I'd recommend you check out the Open CV docs on how to train a cascade
# classifier: https://docs.opencv.org/3.4/dc/d88/tutorial_traincascade.html
# However, in this lecture we just want to use the current classifiers and see if we can detect
# portions of an image which are interesting.
#
# First step is to load opencv and the XML-based classifiers
import cv2 as cv
face_cascade = cv.CascadeClassifier('readonly/haarcascade_frontalface_default.xml')
eye_cascade = cv.CascadeClassifier('readonly/haarcascade_eye.xml')
# Ok, with the classifiers loaded, we now want to try and detect a face. Lets pull in the
# picture we played with last time
img = cv.imread('readonly/floyd.jpg')
# And we'll convert it to grayscale using the cvtColor image
gray = cv.cvtColor(img, cv.COLOR_BGR2GRAY)
# The next step is to use the face_cascade classifier. I'll let you go explore the docs if you
# would like to, but the norm is to use the detectMultiScale() function. This function returns
# a list of objects as rectangles. The first parameter is an ndarray of the image.
faces = face_cascade.detectMultiScale(gray)
# And lets just print those faces out to the screen
faces
faces.tolist()[0]
# +
# The resulting rectangles are in the format of (x,y,w,h) where x and y denote the upper
# left hand point for the image and the width and height represent the bounding box. We know
# how to handle this in PIL
from PIL import Image
# Lets create a PIL image object
pil_img=Image.fromarray(gray,mode="L")
# Now lets bring in our drawing object
from PIL import ImageDraw
# And lets create our drawing context
drawing=ImageDraw.Draw(pil_img)
# Now lets pull the rectangle out of the faces object
rec=faces.tolist()[0]
# Now we just draw a rectangle around the bounds
drawing.rectangle(rec, outline="white")
# And display
display(pil_img)
# -
# So, not quite what we were looking for. What do you think went wrong?
# Well, a quick double check of the docs and it is apparent that OpenCV is return the coordinates
# as (x,y,w,h), while PIL.ImageDraw is looking for (x1,y1,x2,y2). Looks like an easy fix
# Wipe our old image
pil_img=Image.fromarray(gray,mode="L")
# Setup our drawing context
drawing=ImageDraw.Draw(pil_img)
# And draw the new box
drawing.rectangle((rec[0],rec[1],rec[0]+rec[2],rec[1]+rec[3]), outline="white")
# And display
display(pil_img)
# We see the face detection works pretty good on this image! Note that it's apparent that this is
# not head detection, but that the haarcascades file we used is looking for eyes and a mouth.
# Lets try this on something a bit more complex, lets read in our MSI recruitment image
img = cv.imread('readonly/msi_recruitment.gif')
# And lets take a look at that image
display(Image.fromarray(img))
# Whoa, what's that error about? It looks like there is an error on a line deep within the PIL
# Image.py file, and it is trying to call an internal private member called __array_interface__
# on the img object, but this object is None
#
# It turns out that the root of this error is that OpenCV can't work with Gif images. This is
# kind of a pain and unfortunate. But we know how to fix that right? One was is that we could
# just open this in PIL and then save it as a png, then open that in open cv.
#
# Lets use PIL to open our image
pil_img=Image.open('readonly/msi_recruitment.gif')
# now lets convert it to greyscale for opencv, and get the bytestream
open_cv_version=pil_img.convert("L")
# now lets just write that to a file
open_cv_version.save("msi_recruitment.png")
# +
# Ok, now that the conversion of format is done, lets try reading this back into opencv
cv_img=cv.imread('msi_recruitment.png')
# We don't need to color convert this, because we saved it as grayscale
# lets try and detect faces in that image
faces = face_cascade.detectMultiScale(cv_img)
# Now, we still have our PIL color version in a gif
pil_img=Image.open('readonly/msi_recruitment.gif')
# Set our drawing context
drawing=ImageDraw.Draw(pil_img)
# For each item in faces, lets surround it with a red box
for x,y,w,h in faces:
# That might be new syntax for you! Recall that faces is a list of rectangles in (x,y,w,h)
# format, that is, a list of lists. Instead of having to do an iteration and then manually
# pull out each item, we can use tuple unpacking to pull out individual items in the sublist
# directly to variables. A really nice python feature
#
# Now we just need to draw our box
drawing.rectangle((x,y,x+w,y+h), outline="white")
display(pil_img)
# -
# What happened here!? We see that we have detected faces, and that we have drawn boxes
# around those faces on the image, but that the colors have gone all weird! This, it turns
# out, has to do with color limitations for gif images. In short, a gif image has a very
# limited number of colors. This is called a color pallette after the pallette artists
# use to mix paints. For gifs the pallette can only be 256 colors -- but they can be *any*
# 256 colors. When a new color is introduced, is has to take the space of an old color.
# In this case, PIL adds white to the pallette but doesn't know which color to replace and
# thus messes up the image.
#
# Who knew there was so much to learn about image formats? We can see what mode the image
# is in with the .mode attribute
pil_img.mode
# We can see a list of modes in the PILLOW documentation, and they correspond with the
# color spaces we have been using. For the moment though, lets change back to RGB, which
# represents color as a three byte tuple instead of in a pallette.
# Lets read in the image
pil_img=Image.open('readonly/msi_recruitment.gif')
# Lets convert it to RGB mode
pil_img = pil_img.convert("RGB")
# And lets print out the mode
pil_img.mode
# Ok, now lets go back to drawing rectangles. Lets get our drawing object
drawing=ImageDraw.Draw(pil_img)
# And iterate through the faces sequence, tuple unpacking as we go
for x,y,w,h in faces:
# And remember this is width and height so we have to add those appropriately.
drawing.rectangle((x,y,x+w,y+h), outline="white")
display(pil_img)
# +
# Awesome! We managed to detect a bunch of faces in that image. Looks like we have missed
# four faces. In the machine learning world we would call these false negatives - something
# which the machine thought was not a face (so a negative), but that it was incorrect on.
# Consequently, we would call the actual faces that were detected as true positives -
# something that the machine thought was a face and it was correct on. This leaves us with
# false positives - something the machine thought was a face but it wasn't. We see there are
# two of these in the image, picking up shadow patterns or textures in shirts and matching
# them with the haarcascades. Finally, we have true negatives, or the set of all possible
# rectangles the machine learning classifier could consider where it correctly indicated that
# the result was not a face. In this case there are many many true negatives.
# -
# There are a few ways we could try and improve this, and really, it requires a lot of
# experimentation to find good values for a given image. First, lets create a function
# which will plot rectanges for us over the image
def show_rects(faces):
#Lets read in our gif and convert it
pil_img=Image.open('readonly/msi_recruitment.gif').convert("RGB")
# Set our drawing context
drawing=ImageDraw.Draw(pil_img)
# And plot all of the rectangles in faces
for x,y,w,h in faces:
drawing.rectangle((x,y,x+w,y+h), outline="white")
#Finally lets display this
display(pil_img)
# Ok, first up, we could try and binarize this image. It turns out that opencv has a built in
# binarization function called threshold(). You simply pass in the image, the midpoint, and
# the maximum value, as well as a flag which indicates whether the threshold should be
# binary or something else. Lets try this.
cv_img_bin=cv.threshold(img,120,255,cv.THRESH_BINARY)[1] # returns a list, we want the second value
# Now do the actual face detection
faces = face_cascade.detectMultiScale(cv_img_bin)
# Now lets see the results
show_rects(faces)
# +
# That's kind of interesting. Not better, but we do see that there is one false positive
# towards the bottom, where the classifier detected the sunglasses as eyes and the dark shadow
# line below as a mouth.
#
# If you're following in the notebook with this video, why don't you pause things and try a
# few different parameters for the thresholding value?
# -
# The detectMultiScale() function from OpenCV also has a couple of parameters. The first of
# these is the scale factor. The scale factor changes the size of rectangles which are
# considered against the model, that is, the haarcascades XML file. You can think of it as if
# it were changing the size of the rectangles which are on the screen.
#
# Lets experiment with the scale factor. Usually it's a small value, lets try 1.05
faces = face_cascade.detectMultiScale(cv_img,1.05)
# Show those results
show_rects(faces)
# Now lets also try 1.15
faces = face_cascade.detectMultiScale(cv_img,1.15)
# Show those results
show_rects(faces)
# Finally lets also try 1.25
faces = face_cascade.detectMultiScale(cv_img,1.25)
# Show those results
show_rects(faces)
# +
# We can see that as we change the scale factor we change the number of true and
# false positives and negatives. With the scale set to 1.05, we have 7 true positives,
# which are correctly identified faces, and 3 false negatives, which are faces which
# are there but not detected, and 3 false positives, where are non-faces which
# opencv thinks are faces. When we change this to 1.15 we lose the false positives but
# also lose one of the true positives, the person to the right wearing a hat. And
# when we change this to 1.25 we lost more true positives as well.
#
# This is actually a really interesting phenomena in machine learning and artificial
# intelligence. There is a trade off between not only how accurate a model is, but how
# the inaccuracy actually happens. Which of these three models do you think is best?
# -
# Well, the answer to that question is really, "it depends". It depends why you are trying
# to detect faces, and what you are going to do with them. If you think these issues
# are interesting, you might want to check out the Applied Data Science with Python
# specialization Michigan offers on Coursera.
#
# Ok, beyond an opportunity to advertise, did you notice anything else that happened when
# we changed the scale factor? It's subtle, but the speed at which the processing ran
# took longer at smaller scale factors. This is because more subimages are being considered
# for these scales. This could also affect which method we might use.
#
# Jupyter has nice support for timing commands. You might have seen this before, a line
# that starts with a percentage sign in jupyter is called a "magic function". This isn't
# normal python - it's actually a shorthand way of writing a function which Jupyter
# has predefined. It looks a lot like the decorators we talked about in a previous
# lecture, but the magic functions were around long before decorators were part of the
# python language. One of the built-in magic functions in juptyer is called timeit, and this
# repeats a piece of python ten times (by default) and tells you the average speed it
# took to complete.
#
# Lets time the speed of detectmultiscale when using a scale of 1.05
# %timeit face_cascade.detectMultiScale(cv_img,1.05)
# Ok, now lets compare that to the speed at scale = 1.15
# %timeit face_cascade.detectMultiScale(cv_img,1.15)
# +
# You can see that this is a dramatic difference, roughly two and a half times slower
# when using the smaller scale!
#
# This wraps up our discussion of detecting faces in opencv. You'll see that, like OCR, this
# is not a foolproof process. But we can build on the work others have done in machine learning
# and leverage powerful libraries to bring us closer to building a turn key python-based
# solution. Remember that the detection mechanism isn't specific to faces, that's just the
# haarcascades training data we used. On the web you'll be able to find other training data
# to detect other objects, including eyes, animals, and so forth.
# -
# ## More Jupyter Widgets
# One of the nice things about using the Jupyter notebook systems is that there is a
# rich set of contributed plugins that seek to extend this system. In this lecture I
# want to introduce you to one such plugin, call ipy web rtc. Webrtc is a fairly new
# protocol for real time communication on the web. Yup, I'm talking about chatting.
# The widget brings this to the Jupyter notebook system. Lets take a look.
#
# First, lets import from this library two different classes which we'll use in a
# demo, one for the camera and one for images.
from ipywebrtc import CameraStream, ImageRecorder
# Then lets take a look at the camera stream object
help(CameraStream)
# We see from the docs that it's east to get a camera facing the user, and we can have
# the audio on or off. We don't need audio for this demo, so lets create a new camera
# instance
camera = CameraStream.facing_user(audio=False)
# The next object we want to look at is the ImageRecorder
help(ImageRecorder)
# The image recorder lets us actually grab images from the camera stream. There are features
# for downloading and using the image as well. We see that the default format is a png file.
# Lets hook up the ImageRecorder to our stream
image_recorder = ImageRecorder(stream=camera)
# Now, the docs are a little unclear how to use this within Jupyter, but if we call the
# download() function it will actually store the results of the camera which is hooked up
# in image_recorder.image. Lets try it out
# First, lets tell the recorder to start capturing data
image_recorder.recording=True
# Now lets download the image
image_recorder.download()
# Then lets inspect the type of the image
type(image_recorder.image)
# Ok, the object that it stores is an ipywidgets.widgets.widget_media.Image. How do we do
# something useful with this? Well, an inspection of the object shows that there is a handy
# value field which actually holds the bytes behind the image. And we know how to display
# those.
# Lets import PIL Image
import PIL.Image
# And lets import io
import io
# And now lets create a PIL image from the bytes
img = PIL.Image.open(io.BytesIO(image_recorder.image.value))
# And render it to the screen
display(img)
# +
# Great, you see a picture! Hopefully you are following along in one of the notebooks
# and have been able to try this out for yourself!
#
# What can you do with this? This is a great way to get started with a bit of computer vision.
# You already know how to identify a face in the webcam picture, or try and capture text
# from within the picture. With OpenCV there are any number of other things you can do, simply
# with a webcam, the Jupyter notebooks, and python!
| python3/OCR_using_Kraken.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tarea 6. Distribución óptima de capital y selección de portafolios.
#
# <img style="float: right; margin: 0px 0px 15px 15px;" src="https://upload.wikimedia.org/wikipedia/commons/thumb/6/6e/Separation_theorem_of_MPT.svg/2000px-Separation_theorem_of_MPT.svg.png" width="400px" height="400px" />
#
# **Resumen.**
# > En esta tarea, tendrás la oportunidad de aplicar los conceptos y las herramientas que aprendimos en el módulo 3. Específicamente, utilizarás técnicas de optimización media-varianza para construir la frontera de mínima varianza, encontrar el mejor portafolio sobre la frontera mínima varianza, y finalmente, identificar la asignación óptima de capital para un inversionista dado su nivel de averisón al riesgo.
#
# **Criterio de revisión.**
# > Se te calificará de acuerdo a los resultados finales que reportes, basados en tu análisis.
#
# **Antes de comenzar.**
# > Por favor, copiar y pegar este archivo en otra ubicación. Antes de comenzar, nombrarlo *Tarea6_ApellidoNombre*, sin acentos y sin espacios; por ejemplo, en mi caso el archivo se llamaría *Tarea6_JimenezEsteban*. Resolver todos los puntos en dicho archivo y subir en este espacio.
# ## 1. Datos (10 puntos)
#
# Considere los siguientes datos de bonos, índice de acciones, mercados desarrollados, mercados emergentes, fondos privados, activos reales y activos libres de riesgo:
# Importamos pandas y numpy
import pandas as pd
import numpy as np
# +
# Resumen en base anual de rendimientos esperados y volatilidades
annual_ret_summ = pd.DataFrame(columns=['Bonos', 'Acciones', 'Desarrollado', 'Emergente', 'Privados', 'Real', 'Libre_riesgo'], index=['Media', 'Volatilidad'])
annual_ret_summ.loc['Media'] = np.array([0.0400, 0.1060, 0.0830, 0.1190, 0.1280, 0.0620, 0.0300])
annual_ret_summ.loc['Volatilidad'] = np.array([0.0680, 0.2240, 0.2210, 0.3000, 0.2310, 0.0680, None])
annual_ret_summ.round(4)
# -
# Matriz de correlación
corr = pd.DataFrame(data= np.array([[1.0000, 0.4000, 0.2500, 0.2000, 0.1500, 0.2000],
[0.4000, 1.0000, 0.7000, 0.6000, 0.7000, 0.2000],
[0.2500, 0.7000, 1.0000, 0.7500, 0.6000, 0.1000],
[0.2000, 0.6000, 0.7500, 1.0000, 0.2500, 0.1500],
[0.1500, 0.7000, 0.6000, 0.2500, 1.0000, 0.3000],
[0.2000, 0.2000, 0.1000, 0.1500, 0.3000, 1.0000]]),
columns=annual_ret_summ.columns[:-1], index=annual_ret_summ.columns[:-1])
corr.round(4)
# 1. Graficar en el espacio de rendimiento esperado contra volatilidad cada uno de los activos (10 puntos).
# ## 2. Hallando portafolios sobre la frontera de mínima varianza (35 puntos)
#
# Usando los datos del punto anterior:
#
# 1. Halle los pesos del portafolio de mínima varianza considerando todos los activos riesgosos. También reportar claramente el rendimiento esperado, volatilidad y cociente de Sharpe para dicho portafolio (15 puntos).
# 2. Halle los pesos del portafolio EMV considerando todos los activos riesgosos. También reportar claramente el rendimiento esperado, volatilidad y cociente de Sharpe para dicho portafolio (15 puntos).
# 3. Halle la covarianza y la correlación entre los dos portafolios hallados (5 puntos)
# ## 3. Frontera de mínima varianza y LAC (30 puntos)
#
# Con los portafolios que se encontraron en el punto anterior (de mínima varianza y EMV):
#
# 1. Construya la frontera de mínima varianza calculando el rendimiento esperado y volatilidad para varias combinaciones de los anteriores portafolios. Reportar dichas combinaciones en un DataFrame incluyendo pesos, rendimiento, volatilidad y cociente de Sharpe (15 puntos).
# 2. También construya la línea de asignación de capital entre el activo libre de riesgo y el portafolio EMV. Reportar las combinaciones de estos activos en un DataFrame incluyendo pesos, rendimiento, volatilidad y cociente de Sharpe (15 puntos).
# ## 4. Gráficos y conclusiones (25 puntos)
#
# 1. Usando todos los datos obtenidos, grafique:
# - los activos individuales,
# - portafolio de mínima varianza,
# - portafolio eficiente en media-varianza (EMV),
# - frontera de mínima varianza, y
# - línea de asignación de capital,
# en el espacio de rendimiento (eje $y$) vs. volatilidad (eje $x$). Asegúrese de etiquetar todo y poner distintos colores para diferenciar los distintos elementos en su gráfico (15 puntos).
# 2. Suponga que usted está aconsejando a un cliente cuyo coeficiente de aversión al riesgo resultó ser 4. ¿Qué asignación de capital le sugeriría?, ¿qué significa su resultado?(10 puntos)
# <script>
# $(document).ready(function(){
# $('div.prompt').hide();
# $('div.back-to-top').hide();
# $('nav#menubar').hide();
# $('.breadcrumb').hide();
# $('.hidden-print').hide();
# });
# </script>
#
# <footer id="attribution" style="float:right; color:#808080; background:#fff;">
# Created with Jupyter by <NAME>.
# </footer>
| Modulo3/.ipynb_checkpoints/Tarea6_AsignacionOptimaPortafolio-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#R 1 1
def is_multiple(n,m):
if m % n == 0:
i = m/n
return i
else:
return False
is_multiple(3,6)
# -
# Write a short Python function, is even(k), that takes an integer value and
# returns True if k is even, and False otherwise. However, your function
# cannot use the multiplication, modulo, or division operators
#R12
def even(k):
if k % 2 == 0:
return True
else:
return False
# R13
def minmax(a):
minval = 0
maxval = 0
for i in a:
if not minval:
minval = i
elif i > maxval:
maxval = i
elif i < minval:
minval = i
else:
pass
return minval, maxval
a = [10,20, 30, 11, 5, 9, -1, -300, 55]
minmax(a)
# R14
def posint(a):
b = 0
for i in range(0,a):
b += i**2
return b
[i for i in range(8, -9, -2)]
[2**i for i in range(9)]
# Python’s random module includes a function choice(data) that returns a
# random element from a non-empty sequence. The random module includes a more basic function randrange, with parameterization similar to
# the built-in range function, that return a random choice from the given
# range. Using only the randrange function, implement your own version
# of the choice function.
import random
import numpy as np
from scipy.special import factorial, comb
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
# +
def choice(seq):
idx = random.randrange(0, len(seq))
return seq[idx]
x = []
y = []
random.seed(42)
for i in range(500):
x.append(choice([1,2,3,4,5]))
for i in range(500):
y.append(random.choice([1,2,3,4,5]))
plt.figure()
plt.title("Own Function")
sns.countplot(x)
plt.figure()
plt.title("Built-in Function")
sns.countplot(y)
| Data Structure and Algorithms/.ipynb_checkpoints/Section 1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ExternalSource operator
#
# In this example, we will see how to use `ExternalSource` operator which allows us to
# use an external data source as an input to the Pipeline.
# +
import types
import collections
import numpy as np
from random import shuffle
from nvidia.dali.pipeline import Pipeline
import nvidia.dali.ops as ops
import nvidia.dali.types as types
batch_size = 16
# -
# ### Defining the data source
# In this example, we use an infinite iterator as a data source.
class ExternalInputIterator(object):
def __init__(self, batch_size):
self.images_dir = "../../data/images/"
self.batch_size = batch_size
with open(self.images_dir + "file_list.txt", 'r') as f:
self.files = [line.rstrip() for line in f if line is not '']
shuffle(self.files)
def __iter__(self):
self.i = 0
self.n = len(self.files)
return self
def __next__(self):
batch = []
labels = []
for _ in range(self.batch_size):
jpeg_filename, label = self.files[self.i].split(' ')
f = open(self.images_dir + jpeg_filename, 'rb')
batch.append(np.frombuffer(f.read(), dtype = np.uint8))
labels.append(np.array([label], dtype = np.uint8))
self.i = (self.i + 1) % self.n
return (batch, labels)
# ### Defining the pipeline
#
# The next step is to define the Pipeline.
#
# The `ExternalSource` operator accepts an iterable or a callable. If the source provides multiple outputs (e.g. images and labels), that number must also be specified as `num_outputs` argument.
#
# Internally, the pipeline will call `source` (if callable) or run `next(source)` (if iterable) whenever more data is needed to keep the pipeline running.
eii = ExternalInputIterator(batch_size)
class ExternalSourcePipeline(Pipeline):
def __init__(self, batch_size, num_threads, device_id):
super(ExternalSourcePipeline, self).__init__(batch_size,
num_threads,
device_id,
seed=12)
self.source = ops.ExternalSource(source = eii, num_outputs = 2)
self.decode = ops.ImageDecoder(device = "mixed", output_type = types.RGB)
self.enhance = ops.BrightnessContrast(device = "gpu", contrast = 2)
def define_graph(self):
jpegs, labels = self.source()
images = self.decode(jpegs)
output = self.enhance(images)
return (output, labels)
# ### Using the pipeline
pipe = ExternalSourcePipeline(batch_size=batch_size, num_threads=2, device_id = 0)
pipe.build()
pipe_out = pipe.run()
# Notice that labels are still on CPU and no as_cpu call is needed to show them.
batch_cpu = pipe_out[0].as_cpu()
labels_cpu = pipe_out[1]
from __future__ import print_function
import matplotlib.pyplot as plt
img = batch_cpu.at(2)
print(img.shape)
print(labels_cpu.at(2))
plt.axis('off')
plt.imshow(img)
| docs/examples/general/data_loading/external_input.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Automatização
# ## Salvando o Modelo
# +
import pickle
file = 'models/model.sav'
pickle.dump(modelo, open(file, 'wb'))
print('Modelo salvo!')
# -
# ## Carregando um Modelo
# +
file = 'models/model.sav'
modelo_final = pickle.load(open(file, 'rb'))
modelo_prod = modelo_final.score(X_teste, y_teste)
print("Modelo carregado!")
print("Acurácia: %.3f" % (modelo_prod.mean() * 100))
| app/home/vups/lab/.ipynb_checkpoints/08-Automation-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import math
df = pd.read_csv('../data/Residential-Profiles.csv')
df['Time'] = df['Time'].astype('datetime64')
df.plot(x='Time', y='Household 1')
# +
epsilons = [0.1,0.25,0.5,0.75,1.0]
trials = [1,2,3,4,5]
print("********************************")
print("Grid Search on progress")
print("5 trials with epsilon values [0.1, 0.25, 0.5, 0.75, 1]")
print("Take average of 5 iterations")
print("********************************")
print("Prining relative errors...")
for epsilon in epsilons:
for trial in trials:
PEAK_VALUE = 8000
delta = 10e-3
EX = 0
total_relative_error = 0
for house in houses:
for timestamps in range(df.shape[0]):
energy = df.at[timestamps,house]
maxAllowedError = energy * 10 / 100
sgd = maxAllowedError / 2.33
sensitivity = math.sqrt((sgd*sgd) / 2)
exponential = np.random.exponential(scale = sensitivity/epsilon)
noisy_energy = energy + exponential + EX
if noisy_energy > PEAK_VALUE:
EX = noisy_energy - PEAK_VALUE
noisy_energy = PEAK_VALUE
relative_error = abs(energy-noisy_energy)/energy
total_relative_error += relative_error
avg_relative_error = total_relative_error*100/(df.shape[0]*200)
print("epsilon ", epsilon, "iteration", trial, "error", avg_relative_error)
| random_distribution_based/Exponential.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Recruitment Experiment
# In developing omnidirectional learning algorithms, prior work has involved two main approaches: building and reallocating. Building involves adding new resources to support the arrival of new data, whereas reallocation involves compression of representations to make room for new ones. However, biologically, there is a spectrum between these two modes.
#
# In order to examine whether current resources could be better leveraged, we test a range of approaches: **recruitment** of the best-performing existing trees, **building** new trees completely (the default approach used by omnidirectional forests), ignoring all prior trees (essentially an uncertainty forest), and a **hybrid** between building and recruitment.
#
# This experiment examines the performance of these four approaches based on the available training sample size.
# +
import numpy as np
from tensorflow import keras
import matplotlib.pyplot as plt
import functions.recruitment_functions as fn
# -
# **Note:** This notebook tutorial uses functions stored externally within `functions/recruitment_functions.py` to simplify presentation of code. These functions are imported above, along with other libraries.
# ## Recruitment Within Datasets: CIFAR10x10
# ### The CIFAR-100 Dataset
#
# The classification problem that we examine in this tutorial makes use of the [CIFAR-100 dataset](https://www.cs.toronto.edu/~kriz/cifar.html), which is a labeled subset of the 80 million tiny images dataset. There are 100 classes ("fine" labels), containing 600 32x32-pixel images each (500 training and 100 testing). These 100 classes are grouped into 20 superclasses ("coarse" labels). Let's import the data:
# import data
(X_train, y_train), (X_test, y_test) = keras.datasets.cifar100.load_data()
# To give a quick overview of what these data look like, let's plot the first five images in the training dataset.
# plot example data
plt.figure(figsize=(15, 3))
plt.imshow(np.hstack((X_train[0], X_train[1], X_train[2], X_train[3], X_train[4])))
plt.show()
# ### The Experiment
# Now that we have imported the CIFAR-100 dataset, we can prepare to run the experiment. The function for running the experiment, `experiment`, can be found within `functions/recruitment_functions.py`.
# We first declare the hyperparameters to be used for the experiment, which are as follows:
# - `ntrees`: number of trees
# - `reps`: number of repetitions to run the experiment for
# - `estimation_set`: size of set used to train for 10th task, given as proportion (`1-estimation_set` is the size of the set used for validation, the selection of best trees)
# - `num_points_per_task`: total number of points per run
# - `num_points_per_forest`: number of points per forest (each training iteration/task)
# - `task_10_sample`: number of samples to train on for the 10th task
############################
### Main hyperparameters ###
############################
ntrees = 50
reps = 4 # use 30 reps for the paper plot
estimation_set = 0.63
num_points_per_task = 5000
num_points_per_forest = 500
task_10_sample = 10 * np.array([10, 50, 100, 200, 350, 500])
# Because the images are presented as 3D arrays, where the third dimension contains the values of the red, blue, and green channels, respectively, and are divided into training and testing sets of size 500 and 100, as mentioned earlier, we must do a little more preprocessing to get these data into a form suited for the progressive learner.
#
# Here, we recombine the pre-established training and testing sets into one large dataset and reshape the arrays into a 1-dimensional string of numbers. We then call the `sort_data` command to divide the data into training and testing sets for each task.
# +
# reformat data
data_x = np.concatenate([X_train, X_test])
data_x = data_x.reshape(
(data_x.shape[0], data_x.shape[1] * data_x.shape[2] * data_x.shape[3])
)
data_y = np.concatenate([y_train, y_test])
data_y = data_y[:, 0]
# sort data
(
train_x_across_task,
train_y_across_task,
test_x_across_task,
test_y_across_task,
) = fn.sort_data(data_x, data_y, num_points_per_task)
# -
# To replicate the experiment found in [Vogelstein, et al. (2020)](https://arxiv.org/pdf/2004.12908.pdf), we use the hyperparameter values established previously to train a lifelong forest on the first nine CIFAR 10x10 tasks, where we have `50` trees (`ntrees`) and `500` samples (`num_points_per_forest`) for each set.
#
# For the 10th task, we use training sample sizes ranging from `100` to `5000` (`task_10_sample`) and obtain generalization errors for each of the following approaches:
# 1. **Building (default for omnidirectional forests)**, which involves training `ntrees=50` new trees,
# 2. **Uncertainty forest**, which ignores all prior trees,
# 3. **Recruiting**, which selects the `ntrees=50` (out of all 450 existing trees) that perform best on the newly introduced 10th task, and
# 4. **Hybrid**, which both builds `ntrees/2=25` new trees and recruits the `ntrees/2=25` best-performing trees.
#
# Let's call our `experiment` function and give it a run!
# run the experiment
mean_acc_dict, std_acc_dict = fn.experiment(
train_x_across_task,
train_y_across_task,
test_x_across_task,
test_y_across_task,
ntrees,
reps,
estimation_set,
num_points_per_task,
num_points_per_forest,
task_10_sample,
)
# As you can see from the code above, our `experiment` function returns two dictionaries,`mean_acc_dict` and `std_acc_dict`, that respectively contain the means and standard deviations of the generalization errors for each approach (building, UF, recruiting, hybrid) over all repetitions. The dictionary keys correspond to each approach.
# ### Visualizing the Results
# Let's visualize the results of the experiment through the `recruitment_plot` function, which can be found in `functions/recruitment_functions.py`.
# plot results
fn.recruitment_plot(mean_acc_dict, std_acc_dict, task_10_sample)
# From these results, we see that lifelong forests outperform all other approaches except at 5000 task 10 samples, which illustrate how relative performance depends on available resources and sample size.
# ## Recruitment Between Datasets: MNIST/Fashion-MNIST
# Another thing that is of interest is whether we are able to show improvement in performance *between* similar datasets and how the four schemes (recruitment, building, UF, and hybrid) impact this performance. For this, we look to the MNIST and Fashion-MNIST datasets.
# ### MNIST & Fashion-MNIST Datasets
#
# The MNIST dataset contains 60,000 training samples and 10,000 testing samples of handwritten numerical digits (from 0-9), presented as 28x28 grayscale images. Fashion-MNIST, similarly, also contains 60,000 training and 10,000 testing samples in the form of 28x28 grayscale images, and it was intended to serve as an alternative to MNIST for benchmarking machine learning algorithms. These features make the two datasets ideal for studying recruitment between two datasets due to their similarity in structure.
# ### Recruitment on MNIST and Fashion-MNIST Individually
# Before we are able to examine the recruitment performance between datasets, it is important that
# we first look at their individual performance, so that we have a rough benchmark for how these
# datasets will perform. We use the same hyperparameters for the benchmarking recruitment, as follows:
############################
### Main hyperparameters ###
############################
ntrees = 50
n_tasks = 5
reps = 5
estimation_set = 0.63
num_points_per_task = 10000
num_points_per_forest = 1000
test_points_per_task = 1000
task_10_sample = 10 * np.array([10, 50, 100, 200, 350, 500, 1000])
# As can be seen above, we are using 5 tasks for each dataset, meaning that each task contains the data
# for two numbers (in the case of MNIST) or two clothing types (in the case of Fashion-MNIST).
# #### Recruitment on MNIST
#
# The MNIST results are as follows:
# import and sort data
(x_train, y_train), (x_test, y_test) = keras.datasets.mnist.load_data()
(
train_x_across_task,
train_y_across_task,
test_x_across_task,
test_y_across_task,
) = fn.sort_data_mnist(
x_train, y_train, x_test, y_test, num_points_per_task, test_points_per_task, n_tasks
)
# run the experiment
mean_acc_dict, std_acc_dict = fn.experiment_mnist(
train_x_across_task,
train_y_across_task,
test_x_across_task,
test_y_across_task,
ntrees,
n_tasks,
reps,
estimation_set,
num_points_per_task,
num_points_per_forest,
task_10_sample,
)
# plot results
fn.recruitment_plot_mnist(mean_acc_dict, std_acc_dict, task_10_sample)
# #### Recruitment on Fashion-MNIST
# Likewise, the Fashion-MNIST results are as follows:
# import and sort data
(x_train, y_train), (x_test, y_test) = keras.datasets.fashion_mnist.load_data()
(
train_x_across_task,
train_y_across_task,
test_x_across_task,
test_y_across_task,
) = fn.sort_data_mnist(
x_train, y_train, x_test, y_test, num_points_per_task, test_points_per_task, n_tasks
)
# run the experiment
mean_acc_dict, std_acc_dict = fn.experiment_mnist(
train_x_across_task,
train_y_across_task,
test_x_across_task,
test_y_across_task,
ntrees,
n_tasks,
reps,
estimation_set,
num_points_per_task,
num_points_per_forest,
task_10_sample,
)
# plot results
fn.recruitment_plot_mnist(mean_acc_dict, std_acc_dict, task_10_sample)
# Both MNIST and Fashion-MNIST display improvement in generalization error as the number of samples on the last task increases for all schemes except recruiting forests.
# ### Recruitment Between MNIST and Fashion-MNIST
#
# Now that we have the data for the individual datasets, let's look at the performance between the two.
# Essentially, we set the MNIST data as the first task and Fashion-MNIST as the second task, with
# ten labels in each task. For this, we use the following hyperparameters:
############################
### Main hyperparameters ###
############################
ntrees = 50
n_tasks = 2
reps = 5 # 30
estimation_set = 0.63
num_points_per_task = 50000
num_points_per_forest = 25000
test_points_per_task = 10000
task_10_sample = 10 * np.array([10, 50, 100, 500, 1000, 2500, 5000])
# Let's run the experiment, as before.
#
# First, we load the data:
# import data
(
(mnist_x_train, mnist_y_train),
(mnist_x_test, mnist_y_test),
) = keras.datasets.mnist.load_data()
(
(fmnist_x_train, fmnist_y_train),
(fmnist_x_test, fmnist_y_test),
) = keras.datasets.fashion_mnist.load_data()
# Then, we reformat it and sort into train and test samples:
# reformat data
x_train = np.concatenate((mnist_x_train, fmnist_x_train), axis=0)
x_test = np.concatenate((mnist_x_test, fmnist_x_test), axis=0)
y_train = np.concatenate((mnist_y_train, fmnist_y_train + 10), axis=0)
y_test = np.concatenate((mnist_y_test, fmnist_y_test + 10), axis=0)
# sort data
(
train_x_across_task,
train_y_across_task,
test_x_across_task,
test_y_across_task,
) = fn.sort_data_mnist(
x_train, y_train, x_test, y_test, num_points_per_task, test_points_per_task, n_tasks
)
# Next, we run the experiment:
# run the experiment
mean_acc_dict, std_acc_dict = fn.experiment_mnist(
train_x_across_task,
train_y_across_task,
test_x_across_task,
test_y_across_task,
ntrees,
n_tasks,
reps,
estimation_set,
num_points_per_task,
num_points_per_forest,
task_10_sample,
)
# And plot the results:
# plot results
fn.recruitment_plot_mnist_between(mean_acc_dict, std_acc_dict, task_10_sample)
# We can see that when performed between datasets, there is still evidence of recruitment, but the recruiting scheme becomes progressively worse in comparison for this example.
| docs/experiments/recruitment_exp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from quantopian.research import returns, symbols
from quantopian.pipeline.experimental import QTradableStocksUS
from quantopian.pipeline.data import USEquityPricing
from quantopian.pipeline.data.psychsignal import stocktwits
from quantopian.pipeline.factors import SimpleMovingAverage
from quantopian.pipeline.filters import Q500US
from quantopian.research import run_pipeline, prices
from quantopian.pipeline import Pipeline
# Select a time range to inspect
start = '2008-01-01'
end = '2017-12-31'
bench_returns = returns(
assets=symbols('SPY'),
start=start,
end=end,
)
def make_pipeline():
base_universe = QTradableStocksUS()
close_price = USEquityPricing.close.latest
#profitable = mstar.valuation_ratios.ev_to_ebitda > 0 #TODO: incorproate this later
#sentiment_score = SimpleMovingAverage(
# inputs=[stocktwits.bull_minus_bear],
# window_length=3,) #TODO: incorporate this later
return Pipeline(
columns={
'close_price': close_price,
#'sentiment_score': sentiment_score,
},
screen=Q500US()
)
# +
pipeline_output = run_pipeline(
make_pipeline(),
start_date=start,
end_date=end
)
tickers_list = pipeline_output.index.levels[1].unique()
all_t = pipeline_output.index.levels[0].unique()
all_p = prices(tickers_list, start=start, end=end)
# -
class TradingRRL(object):
def __init__(self, T=1000, thisT = 1000, M=300, thisM = 300, N=0, init_t=10000, mu=10000, sigma=0.04, rho=1.0, n_epoch=10):
self.T = T
self.thisT = thisT
self.M = M
self.thisM = thisM
self.N = N
self.TOP = 20
self.threshold = 0.0
self.init_t = init_t
self.mu = mu
self.sigma = sigma
self.rho = rho
self.all_t = None
self.all_p = None
self.t = None
self.p = None
self.bench = None
self.r = None
self.x = np.zeros([T, M + 2])
self.F = np.zeros((T + 1, N))
self.FS = np.zeros((T + 1, N))
self.R = np.zeros((T, N))
self.w = np.ones((M + 2, N))
self.w_opt = np.ones((M + 2, N))
self.epoch_S = pd.DataFrame()
self.n_epoch = n_epoch
self.progress_period = 100
self.q_threshold = 0.5
self.b = np.ones((T+1, N))
self.total = None
self.bench = None
self.tickers_list = None
self.ticker_data = collections.defaultdict(dict)
def quant(self, f):
fc = f.copy()
fc[np.where(np.abs(fc) < self.q_threshold)] = 0
#return np.sign(fc)
return fc
def softmax(self, x):
l2_norm = np.sqrt(x*x).sum()
return x/l2_norm
#e_x = np.exp(x)
#return e_x / e_x.sum()
def set_t_p_r(self, train_phase=True):
if train_phase:
self.t = self.all_t[self.init_t:self.init_t + self.T + self.M + 1]
self.p = self.all_p[self.init_t:self.init_t + self.T + self.M + 1,:] ## TODO: add column dimension for assets > 1
print('p dimension', self.p.shape)
#self.r = -np.diff(self.p, axis=0)
firstr = np.zeros((1, self.p.shape[1]))
self.r = np.diff(self.p, axis=0)/self.p[:-1]
self.r = np.concatenate((firstr, self.r), axis=0)
print('r dimension', self.r.shape)
pd.DataFrame(self.r).to_csv("smallr.csv", header=False, index=False)
else:
self.t = self.all_t[self.init_t:self.init_t + self.thisT + self.thisM + 1]
self.p = self.all_p[self.init_t:self.init_t + self.thisT + self.thisM + 1,:] ## TODO: add column dimension for assets > 1
print('p dimension', self.p.shape)
# self.r = -np.diff(self.p, axis=0)
firstr = np.zeros((1, self.p.shape[1]))
self.r = np.diff(self.p, axis=0) / self.p[:-1]
self.r = np.concatenate((firstr, self.r), axis=0)
def set_x_F(self, train_phase=True):
if train_phase:
for i in range(self.T - 1, -1, -1):
self.x[i] = np.zeros(self.M + 2)
self.x[i][0] = 1.0
self.x[i][self.M + 2 - 1] = self.F[i+1,-1] ## TODO: i used -1 on column
for j in range(1, self.M + 2 - 1, 1):
#self.x[i][j] = self.r[i+ j - 1,0] ## TODO: i used -1 on column:
self.x[i,j] = self.r[i + (j-1), -1] ## TODO: i used -1 on column; and must deal with j
self.F[i] = self.quant(np.tanh(np.dot(self.x[i], self.w)+self.b[i])) ## TODO: test this
else:
thisw = np.ones((self.thisM+2, self.N))
self.x = np.zeros([self.thisT, self.thisM + 2])
self.F = np.zeros((self.thisT + 1, self.N))
for i in range(self.thisT - 1, -1, -1):
self.x[i] = np.zeros(self.thisM + 2)
self.x[i][0] = 1.0
self.x[i][self.thisM + 2 - 1] = self.F[i+1,-1] ## TODO: i used -1 on column
for j in range(1, self.thisM + 2 - 1, 1):
#self.x[i][j] = self.r[i+ j - 1,0] ## TODO: i used -1 on column:
self.x[i,j] = self.r[i + (j-1), -1] ## TODO: i used -1 on column; and must deal with j
self.F[i] = self.quant(np.tanh(np.dot(self.x[i], thisw)+self.b[i])) ## TODO: test this
def calc_R(self):
#self.R = self.mu * (np.dot(self.r[:self.T], self.F[:,1:]) - self.sigma * np.abs(-np.diff(self.F, axis=1)))
#self.R = self.mu * (self.r[:self.T] * self.F[1:]) - self.sigma * np.abs(-np.diff(self.F, axis=0))
#self.R = self.mu * (np.multiply(self.F[1:,], np.reshape(self.r[:self.T], (self.T, -1)))) * (self.sigma) * np.abs(-np.diff(self.F, axis=0))
self.R = ((np.multiply(self.F[1:, ], np.reshape(0+self.r[:self.T], (self.T, -1)))) * (1-self.sigma * np.abs(-np.diff(self.F, axis=0))))
pd.DataFrame(self.R).to_csv('R.csv')
def calc_sumR(self):
self.sumR = np.cumsum(self.R[::-1], axis=0)[::-1] ## TODO: cumsum axis
#self.sumR = np.cumprod(self.R[::-1], axis=0)[::-1] ## TODO: cumsum axis
self.sumR2 = np.cumsum((self.R[::-1] ** 2), axis=0)[::-1] ## TODO: cumsum axis
#self.sumR2 = np.cumprod((self.R[::-1] ** 2), axis=0)[::-1] ## TODO: cumsum axis
#print('cumprod', self.sumR)
def calc_dSdw(self, train_phase=True):
if not train_phase:
self.T = self.thisT
self.M = self.thisM
self.set_x_F(train_phase=train_phase)
self.calc_R()
self.calc_sumR()
self.Sall = np.empty(0) # a list of period-to-date sharpe ratios, for all n investments
self.dSdw = np.zeros((self.M + 2, self.N))
for j in range(self.N):
self.A = self.sumR[0,j] / self.T
self.B = self.sumR2[0,j] / self.T
#self.A = self.sumR / self.T
#self.B = self.sumR2 / self.T
self.S = self.A / np.sqrt(self.B - (self.A ** 2))
#self.S = ((self.B[1:,j]*np.diff(self.A[:,j], axis=0)-0.5*self.A[1:,j]*np.diff(self.B[:,j], axis=0))/ (self.B[1,j] - (self.A[1,j] ** 2))**(3/2))[1]
#self.S = (self.B[1,j] - (self.A[1,j] ** 2))**(3/2)
#print('sharpe checl', np.isnan(self.r).sum())
self.dSdA = self.S * (1 + self.S ** 2) / self.A
self.dSdB = -self.S ** 3 / 2 / self.A ** 2
self.dAdR = 1.0 / self.T
self.dBdR = 2.0 / self.T * self.R[:,j]
self.dRdF = -self.mu * self.sigma * np.sign(-np.diff(self.F, axis=0))
self.dRdFp = self.mu * self.r[:self.T] + self.mu * self.sigma * np.sign(-np.diff(self.F, axis=0)) ## TODO: r needs to be a matrix if assets > 1
self.dFdw = np.zeros(self.M + 2)
self.dFpdw = np.zeros(self.M + 2)
#self.dSdw = np.zeros((self.M + 2, self.N)) ## TODO: should not have put this here. this resets everytime
self.dSdw_j = np.zeros(self.M + 2)
for i in range(self.T - 1, -1, -1):
if i != self.T - 1:
self.dFpdw = self.dFdw.copy()
self.dFdw = (1 - self.F[i,j] ** 2) * (self.x[i] + self.w[self.M + 2 - 1,j] * self.dFpdw)
self.dSdw_j += (self.dSdA * self.dAdR + self.dSdB * self.dBdR[i]) * (
self.dRdF[i,j] * self.dFdw + self.dRdFp[i,j] * self.dFpdw)
self.dSdw[:, j] = self.dSdw_j
self.Sall = np.append(self.Sall, self.S)
def update_w(self):
self.w += self.rho * self.dSdw
def get_investment_weights(self, train_phase=True):
if not train_phase:
self.FS = np.zeros((self.thisT + 1, self.N))
for i in range(self.FS.shape[0]):
self.FS[i] = np.multiply(self.F[i], self.Sall)
tmp = np.apply_along_axis(self.select_n, 1, self.FS) # TODO: conisder taking the abs(): magnitutde
F1 = np.apply_along_axis(self.softmax, 1, tmp)
print('MAKE F1', F1.shape)
print('see F1', F1)
print('see R', self.R)
mask = F1 != 0
_, j = np.where(mask)
for ji in set(j):
self.ticker_data[self.tickers_list[ji]]['inv weight'] = F1[-2, ji]
self.ticker_data[self.tickers_list[ji]]['return'] = self.R[-2, ji]
print(self.ticker_data)
return F1
def select_n(self, array):
threshold = max(heapq.nlargest(self.TOP, array)[-1], self.threshold)
new_array = [x if x >= threshold else 0 for x in array]
return new_array
def fit(self):
pre_epoch_times = len(self.epoch_S)
self.calc_dSdw()
print("Epoch loop start. Initial sharp's ratio is " + str(np.mean(self.Sall)) + ".")
print('s len', len(self.Sall))
self.S_opt = self.Sall
tic = time.clock()
for e_index in range(self.n_epoch):
self.calc_dSdw()
if np.mean(self.Sall) > np.mean(self.S_opt):
self.S_opt = self.Sall
self.w_opt = self.w.copy()
#self.Sall = np.apply_along_axis(self.select_n, 0, self.Sall) # TODO: don't do this here
self.epoch_S[e_index] = np.array(self.S_opt)
self.update_w()
if e_index % self.progress_period == self.progress_period - 1:
toc = time.clock()
print("Epoch: " + str(e_index + pre_epoch_times + 1) + "/" + str(
self.n_epoch + pre_epoch_times) + ". Shape's ratio: " + str(self.Sall[self.Sall.nonzero()].mean()) + ". Elapsed time: " + str(
toc - tic) + " sec.")
toc = time.clock()
print("Epoch: " + str(e_index + pre_epoch_times + 1) + "/" + str(
self.n_epoch + pre_epoch_times) + ". Shape's ratio after iteration: " + str(self.S_opt[self.S_opt.nonzero()].mean()) + ". Elapsed time: " + str(
toc - tic) + " sec.")
self.w = self.w_opt.copy()
self.calc_dSdw()
print("Epoch loop end. Optimized sharp's ratio is " + str(self.S_opt[self.S_opt.nonzero()].mean()) + ".")
print('first check', self.Sall)
print('now check', self.epoch_S)
print('R dimension', self.R.shape)
def save_weight(self, train_phase=True):
if train_phase:
self.F1 = self.get_investment_weights()
pd.DataFrame(self.w).to_csv("w.csv", header=False, index=False)
self.epoch_S.to_csv("epoch_S.csv", header=False, index=False)
pd.DataFrame(self.F).to_csv("f.csv", header=False, index=False)
pd.DataFrame(self.FS).to_csv("fs.csv", header=False, index=False)
pd.DataFrame(self.F1).to_csv("f1.csv", header=False, index=False)
else:
self.F1 = self.get_investment_weights(train_phase=False)
pd.DataFrame().from_dict(self.ticker_data).T.to_csv('ticker_data.csv')
def load_weight(self):
tmp = pd.read_csv("w.csv", header=None)
self.w = tmp.T.values[0]
def get_investment_sum(self, train_phase=True):
firstR = np.zeros((1,self.p.shape[1]))
self.R = np.concatenate((firstR, self.R), axis=0)
tmp = np.multiply(self.R, self.F1)
self.total = self.mu * ((1+tmp.sum(axis=1)).cumprod(axis=0))
print('iam here', self.total.shape, self.total)
if train_phase:
pd.DataFrame(self.total).to_csv('investment_sum.csv')
else:
pd.DataFrame(self.total).to_csv('investment_sum_testphase.csv')
| universal_portfolio/rrl_trading/01_python/stock_selection_Berlin.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Linear Algebra
# > ## Linearity
# > ### superpostion principle (중첩의 원리)
# >> a function $F(x)$ that satisfies the superposition principle is called a linear function
# >> ### additivity
# >>> ### $F(x_1 + x_2) = F(x_1) + F(x_2)$
# >>
# >> ### homogeneity
# >>> ### $F(ax) = aF(x), \text{ for scalar } a.$
# >> $ homogenous solution + particular solution = 0$
# >
# > ## Algebra
# > ### synbols that stand for numbers
# >> ## Geometry of Linear Equations
# # Methord of Solution
# > ## [Row pciture](https://twlab.tistory.com/6?category=668741)
# >> 공간상에서 the line,plane of dot products
# >> $
# \begin{bmatrix}
# 2 & 5 \\ 1 & 3
# \end{bmatrix}
# \begin{bmatrix}
# 1 \\ 2
# \end{bmatrix} =
# \begin{bmatrix}
# 2 & 5 \\ 0 & 0
# \end{bmatrix}
# \begin{bmatrix}
# 1 \\ 2
# \end{bmatrix} +
# \begin{bmatrix}
# 0 & 0 \\ 1 & 3
# \end{bmatrix}
# \begin{bmatrix}
# 1 \\ 2
# \end{bmatrix} =
# \begin{bmatrix}
# 12 \\ 7
# \end{bmatrix}
# $
# > ## [Column pciture](https://twlab.tistory.com/6?category=668741)
# >> 공간상에서 linear combination of vectors
# >> $
# \begin{bmatrix}
# 2 & 5 \\ 1 & 3
# \end{bmatrix}
# \begin{bmatrix}
# 1 \\ 2
# \end{bmatrix} =
# 1\:
# \begin{bmatrix}
# 2 \\ 1
# \end{bmatrix} +
# 2 \:
# \begin{bmatrix}
# 5 \\ 3
# \end{bmatrix} =
# \begin{bmatrix}
# 12 \\ 7
# \end{bmatrix}
# $
import sympy as sm
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib widget
# ### find solution.
# > ### $
# \begin{cases}
# 2x & - & y & & & = 0 \\
# -x & + & 2y & - &z & = -1 \\
# & - &3y & + & 4z & = 4
# \end{cases}$
#
# > #### $
# \begin{bmatrix} 2 & -1 & 0 \\ -1 & 2 & -1 \\ 0 & -3 & 4 \end{bmatrix} \:
# \begin{bmatrix} x \\ y \\ z \end{bmatrix} \: = \:
# \begin{bmatrix} 0 \\ -1 \\ 4 \end{bmatrix}
# $
# +
fig = plt.figure()
ax = fig.add_subplot(projection = '3d')
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_xlim(-10,10)
ax.set_ylim(-10,10)
ax.set_zlim(-10,10)
xi = np.linspace(-5,5,10)
yi = np.linspace(-5,5,10)
xi,yi = np.meshgrid(xi,yi)
ax.plot_surface(xi, 2*xi, yi,alpha=0.5)
ax.plot_surface(xi, yi, -xi+2*yi+1,alpha=0.5)
ax.plot_surface(xi, yi, 3/4*yi+1,alpha=0.5)
# -
x,y,z = sm.symbols('x y z')
sm.solve([2*x-y,-x+2*y-z+1,-3*y+4*z-4],[x,y,z])
ax.scatter(0,0,1,marker='o',color='r',s = 100)
# # Column picture
# > ## $
# x\:
# \begin{bmatrix}
# 2 \\ -1 \\ 0
# \end{bmatrix} +
# y\:
# \begin{bmatrix}
# -1 \\ 2 \\ -3
# \end{bmatrix} +
# z\:
# \begin{bmatrix}
# 0 \\ -1 \\ 4
# \end{bmatrix} \:= \:
# \begin{bmatrix}
# 0 \\ -1 \\ 4
# \end{bmatrix}
# $
# + jupyter={"source_hidden": true} tags=[]
fig1 = plt.figure()
ax = fig1.add_subplot(projection='3d')
ax.set_xlim(-4,4)
ax.set_ylim(-4,4)
ax.set_zlim(-4,4)
ax.quiver(0,0,0,2,-1,0)
ax.quiver(0,0,0,-1,2,-3)
ax.quiver(0,0,0,0,-1,4)
ax.scatter(0,0,0,c='r')
ax.scatter(0,-1,4,c='r')
# -
M = sm.Matrix([[2,-1,0,0],[-1,2,-1,-1],[0,-3,4,4]])
sm.Matrix([(2,-1,0,0),(-1,2,-1,-1),(0,-3,4,4)])
sm.Matrix(((2,-1,0,0),(-1,2,-1,-1),(0,-3,4,4)))
A[:,:-1]
A[:,-1]
sm.linsolve(sm.Matrix([[2,-1,0,0],[-1,2,-1,-1],[0,-3,4,4]]),(x,y,z))
sm.linsolve((M[:,:-1],M[:,-1]),x,y,z)
| python/Vectors/algebra.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="b518b04cbfe0"
# ##### Copyright 2020 The TensorFlow Authors.
# + cellView="form" id="906e07f6e562"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="daf323e33b84"
# # Writing a training loop from scratch
# + [markdown] id="2440f6e0c5ef"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/guide/keras/writing_a_training_loop_from_scratch"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/snapshot-keras/site/en/guide/keras/writing_a_training_loop_from_scratch.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/keras-team/keras-io/blob/master/guides/writing_a_training_loop_from_scratch.py"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/keras/writing_a_training_loop_from_scratch.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="8d4ac441b1fc"
# ## Setup
# + id="ae2407ad926f"
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
import numpy as np
# + [markdown] id="0f5a253901f8"
# ## Introduction
#
# Keras provides default training and evaluation loops, `fit()` and `evaluate()`.
# Their usage is covered in the guide
# [Training & evaluation with the built-in methods](https://www.tensorflow.org/guide/keras/train_and_evaluate/).
#
# If you want to customize the learning algorithm of your model while still leveraging
# the convenience of `fit()`
# (for instance, to train a GAN using `fit()`), you can subclass the `Model` class and
# implement your own `train_step()` method, which
# is called repeatedly during `fit()`. This is covered in the guide
# [Customizing what happens in `fit()`](https://www.tensorflow.org/guide/keras/customizing_what_happens_in_fit/).
#
# Now, if you want very low-level control over training & evaluation, you should write
# your own training & evaluation loops from scratch. This is what this guide is about.
# + [markdown] id="f4f47351a3ec"
# ## Using the `GradientTape`: a first end-to-end example
#
# Calling a model inside a `GradientTape` scope enables you to retrieve the gradients of
# the trainable weights of the layer with respect to a loss value. Using an optimizer
# instance, you can use these gradients to update these variables (which you can
# retrieve using `model.trainable_weights`).
#
# Let's consider a simple MNIST model:
# + id="aaa775ce7dab"
inputs = keras.Input(shape=(784,), name="digits")
x1 = layers.Dense(64, activation="relu")(inputs)
x2 = layers.Dense(64, activation="relu")(x1)
outputs = layers.Dense(10, name="predictions")(x2)
model = keras.Model(inputs=inputs, outputs=outputs)
# + [markdown] id="d8b02a5759cf"
# Let's train it using mini-batch gradient with a custom training loop.
#
# First, we're going to need an optimizer, a loss function, and a dataset:
# + id="f2c6257b8d02"
# Instantiate an optimizer.
optimizer = keras.optimizers.SGD(learning_rate=1e-3)
# Instantiate a loss function.
loss_fn = keras.losses.SparseCategoricalCrossentropy(from_logits=True)
# Prepare the training dataset.
batch_size = 64
(x_train, y_train), (x_test, y_test) = keras.datasets.mnist.load_data()
x_train = np.reshape(x_train, (-1, 784))
x_test = np.reshape(x_test, (-1, 784))
# Reserve 10,000 samples for validation.
x_val = x_train[-10000:]
y_val = y_train[-10000:]
x_train = x_train[:-10000]
y_train = y_train[:-10000]
# Prepare the training dataset.
train_dataset = tf.data.Dataset.from_tensor_slices((x_train, y_train))
train_dataset = train_dataset.shuffle(buffer_size=1024).batch(batch_size)
# Prepare the validation dataset.
val_dataset = tf.data.Dataset.from_tensor_slices((x_val, y_val))
val_dataset = val_dataset.batch(batch_size)
# + [markdown] id="5c30285b1a2e"
# Here's our training loop:
#
# - We open a `for` loop that iterates over epochs
# - For each epoch, we open a `for` loop that iterates over the dataset, in batches
# - For each batch, we open a `GradientTape()` scope
# - Inside this scope, we call the model (forward pass) and compute the loss
# - Outside the scope, we retrieve the gradients of the weights
# of the model with regard to the loss
# - Finally, we use the optimizer to update the weights of the model based on the
# gradients
# + id="5bf4c10ceb50"
epochs = 2
for epoch in range(epochs):
print("\nStart of epoch %d" % (epoch,))
# Iterate over the batches of the dataset.
for step, (x_batch_train, y_batch_train) in enumerate(train_dataset):
# Open a GradientTape to record the operations run
# during the forward pass, which enables auto-differentiation.
with tf.GradientTape() as tape:
# Run the forward pass of the layer.
# The operations that the layer applies
# to its inputs are going to be recorded
# on the GradientTape.
logits = model(x_batch_train, training=True) # Logits for this minibatch
# Compute the loss value for this minibatch.
loss_value = loss_fn(y_batch_train, logits)
# Use the gradient tape to automatically retrieve
# the gradients of the trainable variables with respect to the loss.
grads = tape.gradient(loss_value, model.trainable_weights)
# Run one step of gradient descent by updating
# the value of the variables to minimize the loss.
optimizer.apply_gradients(zip(grads, model.trainable_weights))
# Log every 200 batches.
if step % 200 == 0:
print(
"Training loss (for one batch) at step %d: %.4f"
% (step, float(loss_value))
)
print("Seen so far: %s samples" % ((step + 1) * batch_size))
# + [markdown] id="d600076b7be0"
# ## Low-level handling of metrics
#
# Let's add metrics monitoring to this basic loop.
#
# You can readily reuse the built-in metrics (or custom ones you wrote) in such training
# loops written from scratch. Here's the flow:
#
# - Instantiate the metric at the start of the loop
# - Call `metric.update_state()` after each batch
# - Call `metric.result()` when you need to display the current value of the metric
# - Call `metric.reset_states()` when you need to clear the state of the metric
# (typically at the end of an epoch)
#
# Let's use this knowledge to compute `SparseCategoricalAccuracy` on validation data at
# the end of each epoch:
# + id="2602509b16c7"
# Get model
inputs = keras.Input(shape=(784,), name="digits")
x = layers.Dense(64, activation="relu", name="dense_1")(inputs)
x = layers.Dense(64, activation="relu", name="dense_2")(x)
outputs = layers.Dense(10, name="predictions")(x)
model = keras.Model(inputs=inputs, outputs=outputs)
# Instantiate an optimizer to train the model.
optimizer = keras.optimizers.SGD(learning_rate=1e-3)
# Instantiate a loss function.
loss_fn = keras.losses.SparseCategoricalCrossentropy(from_logits=True)
# Prepare the metrics.
train_acc_metric = keras.metrics.SparseCategoricalAccuracy()
val_acc_metric = keras.metrics.SparseCategoricalAccuracy()
# + [markdown] id="9111a5cc87dc"
# Here's our training & evaluation loop:
# + id="654e2311dbff"
import time
epochs = 2
for epoch in range(epochs):
print("\nStart of epoch %d" % (epoch,))
start_time = time.time()
# Iterate over the batches of the dataset.
for step, (x_batch_train, y_batch_train) in enumerate(train_dataset):
with tf.GradientTape() as tape:
logits = model(x_batch_train, training=True)
loss_value = loss_fn(y_batch_train, logits)
grads = tape.gradient(loss_value, model.trainable_weights)
optimizer.apply_gradients(zip(grads, model.trainable_weights))
# Update training metric.
train_acc_metric.update_state(y_batch_train, logits)
# Log every 200 batches.
if step % 200 == 0:
print(
"Training loss (for one batch) at step %d: %.4f"
% (step, float(loss_value))
)
print("Seen so far: %d samples" % ((step + 1) * batch_size))
# Display metrics at the end of each epoch.
train_acc = train_acc_metric.result()
print("Training acc over epoch: %.4f" % (float(train_acc),))
# Reset training metrics at the end of each epoch
train_acc_metric.reset_states()
# Run a validation loop at the end of each epoch.
for x_batch_val, y_batch_val in val_dataset:
val_logits = model(x_batch_val, training=False)
# Update val metrics
val_acc_metric.update_state(y_batch_val, val_logits)
val_acc = val_acc_metric.result()
val_acc_metric.reset_states()
print("Validation acc: %.4f" % (float(val_acc),))
print("Time taken: %.2fs" % (time.time() - start_time))
# + [markdown] id="940d8d9fae83"
# ## Speeding-up your training step with `tf.function`
#
# The default runtime in TensorFlow 2.0 is
# [eager execution](https://www.tensorflow.org/guide/eager). As such, our training loop
# above executes eagerly.
#
# This is great for debugging, but graph compilation has a definite performance
# advantage. Describing your computation as a static graph enables the framework
# to apply global performance optimizations. This is impossible when
# the framework is constrained to greedly execute one operation after another,
# with no knowledge of what comes next.
#
# You can compile into a static graph any function that takes tensors as input.
# Just add a `@tf.function` decorator on it, like this:
# + id="fdacc2d48ade"
@tf.function
def train_step(x, y):
with tf.GradientTape() as tape:
logits = model(x, training=True)
loss_value = loss_fn(y, logits)
grads = tape.gradient(loss_value, model.trainable_weights)
optimizer.apply_gradients(zip(grads, model.trainable_weights))
train_acc_metric.update_state(y, logits)
return loss_value
# + [markdown] id="ab61b0bf3126"
# Let's do the same with the evaluation step:
# + id="da4828fd8ef7"
@tf.function
def test_step(x, y):
val_logits = model(x, training=False)
val_acc_metric.update_state(y, val_logits)
# + [markdown] id="d552377968f1"
# Now, let's re-run our training loop with this compiled training step:
# + id="d69d73c94e44"
import time
epochs = 2
for epoch in range(epochs):
print("\nStart of epoch %d" % (epoch,))
start_time = time.time()
# Iterate over the batches of the dataset.
for step, (x_batch_train, y_batch_train) in enumerate(train_dataset):
loss_value = train_step(x_batch_train, y_batch_train)
# Log every 200 batches.
if step % 200 == 0:
print(
"Training loss (for one batch) at step %d: %.4f"
% (step, float(loss_value))
)
print("Seen so far: %d samples" % ((step + 1) * batch_size))
# Display metrics at the end of each epoch.
train_acc = train_acc_metric.result()
print("Training acc over epoch: %.4f" % (float(train_acc),))
# Reset training metrics at the end of each epoch
train_acc_metric.reset_states()
# Run a validation loop at the end of each epoch.
for x_batch_val, y_batch_val in val_dataset:
test_step(x_batch_val, y_batch_val)
val_acc = val_acc_metric.result()
val_acc_metric.reset_states()
print("Validation acc: %.4f" % (float(val_acc),))
print("Time taken: %.2fs" % (time.time() - start_time))
# + [markdown] id="8977d77a8095"
# Much faster, isn't it?
# + [markdown] id="b5b5a54d339a"
# ## Low-level handling of losses tracked by the model
#
# Layers & models recursively track any losses created during the forward pass
# by layers that call `self.add_loss(value)`. The resulting list of scalar loss
# values are available via the property `model.losses`
# at the end of the forward pass.
#
# If you want to be using these loss components, you should sum them
# and add them to the main loss in your training step.
#
# Consider this layer, that creates an activity regularization loss:
# + id="4ec7c4b16596"
class ActivityRegularizationLayer(layers.Layer):
def call(self, inputs):
self.add_loss(1e-2 * tf.reduce_sum(inputs))
return inputs
# + [markdown] id="6b12260b8bf2"
# Let's build a really simple model that uses it:
# + id="57afe49e6b93"
inputs = keras.Input(shape=(784,), name="digits")
x = layers.Dense(64, activation="relu")(inputs)
# Insert activity regularization as a layer
x = ActivityRegularizationLayer()(x)
x = layers.Dense(64, activation="relu")(x)
outputs = layers.Dense(10, name="predictions")(x)
model = keras.Model(inputs=inputs, outputs=outputs)
# + [markdown] id="aadb58115c13"
# Here's what our training step should look like now:
# + id="cf674776a0d2"
@tf.function
def train_step(x, y):
with tf.GradientTape() as tape:
logits = model(x, training=True)
loss_value = loss_fn(y, logits)
# Add any extra losses created during the forward pass.
loss_value += sum(model.losses)
grads = tape.gradient(loss_value, model.trainable_weights)
optimizer.apply_gradients(zip(grads, model.trainable_weights))
train_acc_metric.update_state(y, logits)
return loss_value
# + [markdown] id="0af04732fe78"
# ## Summary
#
# Now you know everything there is to know about using built-in training loops and
# writing your own from scratch.
#
# To conclude, here's a simple end-to-end example that ties together everything
# you've learned in this guide: a DCGAN trained on MNIST digits.
# + [markdown] id="9fb325331a1e"
# ## End-to-end example: a GAN training loop from scratch
#
# You may be familiar with Generative Adversarial Networks (GANs). GANs can generate new
# images that look almost real, by learning the latent distribution of a training
# dataset of images (the "latent space" of the images).
#
# A GAN is made of two parts: a "generator" model that maps points in the latent
# space to points in image space, a "discriminator" model, a classifier
# that can tell the difference between real images (from the training dataset)
# and fake images (the output of the generator network).
#
# A GAN training loop looks like this:
#
# 1) Train the discriminator.
# - Sample a batch of random points in the latent space.
# - Turn the points into fake images via the "generator" model.
# - Get a batch of real images and combine them with the generated images.
# - Train the "discriminator" model to classify generated vs. real images.
#
# 2) Train the generator.
# - Sample random points in the latent space.
# - Turn the points into fake images via the "generator" network.
# - Get a batch of real images and combine them with the generated images.
# - Train the "generator" model to "fool" the discriminator and classify the fake images
# as real.
#
# For a much more detailed overview of how GANs works, see
# [Deep Learning with Python](https://www.manning.com/books/deep-learning-with-python).
#
# Let's implement this training loop. First, create the discriminator meant to classify
# fake vs real digits:
# + id="fabf9cef3400"
discriminator = keras.Sequential(
[
keras.Input(shape=(28, 28, 1)),
layers.Conv2D(64, (3, 3), strides=(2, 2), padding="same"),
layers.LeakyReLU(alpha=0.2),
layers.Conv2D(128, (3, 3), strides=(2, 2), padding="same"),
layers.LeakyReLU(alpha=0.2),
layers.GlobalMaxPooling2D(),
layers.Dense(1),
],
name="discriminator",
)
discriminator.summary()
# + [markdown] id="73396eb6daf9"
# Then let's create a generator network,
# that turns latent vectors into outputs of shape `(28, 28, 1)` (representing
# MNIST digits):
# + id="821d203bfb3e"
latent_dim = 128
generator = keras.Sequential(
[
keras.Input(shape=(latent_dim,)),
# We want to generate 128 coefficients to reshape into a 7x7x128 map
layers.Dense(7 * 7 * 128),
layers.LeakyReLU(alpha=0.2),
layers.Reshape((7, 7, 128)),
layers.Conv2DTranspose(128, (4, 4), strides=(2, 2), padding="same"),
layers.LeakyReLU(alpha=0.2),
layers.Conv2DTranspose(128, (4, 4), strides=(2, 2), padding="same"),
layers.LeakyReLU(alpha=0.2),
layers.Conv2D(1, (7, 7), padding="same", activation="sigmoid"),
],
name="generator",
)
# + [markdown] id="f0d6d54a78a0"
# Here's the key bit: the training loop. As you can see it is quite straightforward. The
# training step function only takes 17 lines.
# + id="3a11c875142e"
# Instantiate one optimizer for the discriminator and another for the generator.
d_optimizer = keras.optimizers.Adam(learning_rate=0.0003)
g_optimizer = keras.optimizers.Adam(learning_rate=0.0004)
# Instantiate a loss function.
loss_fn = keras.losses.BinaryCrossentropy(from_logits=True)
@tf.function
def train_step(real_images):
# Sample random points in the latent space
random_latent_vectors = tf.random.normal(shape=(batch_size, latent_dim))
# Decode them to fake images
generated_images = generator(random_latent_vectors)
# Combine them with real images
combined_images = tf.concat([generated_images, real_images], axis=0)
# Assemble labels discriminating real from fake images
labels = tf.concat(
[tf.ones((batch_size, 1)), tf.zeros((real_images.shape[0], 1))], axis=0
)
# Add random noise to the labels - important trick!
labels += 0.05 * tf.random.uniform(labels.shape)
# Train the discriminator
with tf.GradientTape() as tape:
predictions = discriminator(combined_images)
d_loss = loss_fn(labels, predictions)
grads = tape.gradient(d_loss, discriminator.trainable_weights)
d_optimizer.apply_gradients(zip(grads, discriminator.trainable_weights))
# Sample random points in the latent space
random_latent_vectors = tf.random.normal(shape=(batch_size, latent_dim))
# Assemble labels that say "all real images"
misleading_labels = tf.zeros((batch_size, 1))
# Train the generator (note that we should *not* update the weights
# of the discriminator)!
with tf.GradientTape() as tape:
predictions = discriminator(generator(random_latent_vectors))
g_loss = loss_fn(misleading_labels, predictions)
grads = tape.gradient(g_loss, generator.trainable_weights)
g_optimizer.apply_gradients(zip(grads, generator.trainable_weights))
return d_loss, g_loss, generated_images
# + [markdown] id="fa6bd6292488"
# Let's train our GAN, by repeatedly calling `train_step` on batches of images.
#
# Since our discriminator and generator are convnets, you're going to want to
# run this code on a GPU.
# + id="b6a4e3d42262"
import os
# Prepare the dataset. We use both the training & test MNIST digits.
batch_size = 64
(x_train, _), (x_test, _) = keras.datasets.mnist.load_data()
all_digits = np.concatenate([x_train, x_test])
all_digits = all_digits.astype("float32") / 255.0
all_digits = np.reshape(all_digits, (-1, 28, 28, 1))
dataset = tf.data.Dataset.from_tensor_slices(all_digits)
dataset = dataset.shuffle(buffer_size=1024).batch(batch_size)
epochs = 1 # In practice you need at least 20 epochs to generate nice digits.
save_dir = "./"
for epoch in range(epochs):
print("\nStart epoch", epoch)
for step, real_images in enumerate(dataset):
# Train the discriminator & generator on one batch of real images.
d_loss, g_loss, generated_images = train_step(real_images)
# Logging.
if step % 200 == 0:
# Print metrics
print("discriminator loss at step %d: %.2f" % (step, d_loss))
print("adversarial loss at step %d: %.2f" % (step, g_loss))
# Save one generated image
img = tf.keras.preprocessing.image.array_to_img(
generated_images[0] * 255.0, scale=False
)
img.save(os.path.join(save_dir, "generated_img" + str(step) + ".png"))
# To limit execution time we stop after 10 steps.
# Remove the lines below to actually train the model!
if step > 10:
break
# + [markdown] id="a92959ac630b"
# That's it! You'll get nice-looking fake MNIST digits after just ~30s of training on the
# Colab GPU.
| site/en-snapshot/guide/keras/writing_a_training_loop_from_scratch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
url = 'https://www.yahoo.co.jp/'
ua = 'Mozilla/5.0 (Macintosh; Intel Mac OS X 10_13_5) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/67.0.3396.99 Safari/537.36'
headers = {'User-Agent': ua}
r_ua = requests.get(url, headers=headers)
| notebook/requests_request_header.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Generate BQM with random weights
#
# ### The weights are random and normalized in the interval [-1,1].
#
# ### The matrices can be dense: all the (i,j) are not zero
# ### or they can be sparse: the percentage of (i,j) different from zero is passed as connectivity parameter
# +
import dwave_utils.utils as dwu
bqm_dense = dwu.random_dense_bqm(5)
bqm_sparse = dwu.random_sparse_bqm(5, connectivity=0.3)
# -
# ## Print basic info about BQM
#
# This method should be extended to provide more information:
dwu.bqm_info(bqm_dense,verbose=True)
dwu.bqm_info(bqm_sparse,verbose=True)
# ## Display BQMs weight matrices as heatmaps
#
#
dwu.bqm_heatplot(bqm_dense)
dwu.bqm_heatplot(bqm_sparse)
# ## Display the distribution of the weights
dwu.bqm_distplot(bqm_dense)
dwu.bqm_distplot(bqm_sparse)
# ## Save and open the BQM as binary files
dwu.save_bqm(bqm_dense, './dense_bqm.bqm')
# +
bqm_dense_new = dwu.open_bqm('./dense_bqm.bqm')
print(bqm_dense_new)
| examples/BQM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import scipy as sp
import pandas as pd
import seaborn as sns
import copy
import matplotlib
matplotlib.rcParams['pdf.fonttype'] = 42
matplotlib.rcParams['ps.fonttype'] = 42
import matplotlib.pyplot as plt
# # Generate Data
# +
np.random.seed(101)
n = 50
y = sp.stats.expon.rvs(size = n, scale = 1) #survival
c = sp.stats.expon.rvs(size= n,scale = 0.5) #censoring
t = copy.copy(y)
t[y>c] = c[y>c]
delta = np.ones(n,dtype = 'int')
delta[y>c] = 0
dy = 0.1
y_plot = np.arange(dy,10,dy)
#Normalize
scale = np.sum(t)/np.sum(delta)
t = t/scale
# -
# # Parametric Predictive
a_samp_smc = np.load('plot_files/sim_a_samp_smc.npy')
b_samp_smc = np.load('plot_files/sim_b_samp_smc.npy')
log_w_smc = np.load('plot_files/sim_log_w_smc.npy')
particle_ind_smc = np.load('plot_files/sim_particle_ind_smc.npy')
ESS_smc = np.load('plot_files/sim_ESS_smc.npy')
theta_hist_smc = np.load('plot_files/sim_theta_hist_smc.npy')
# +
#SMC
#Compute mean samples
B = np.shape(a_samp_smc)[0]
theta_samp_smc = b_samp_smc/(a_samp_smc-1)
#Normalize IS weights, reweight and resample
log_z_smc = sp.special.logsumexp(log_w_smc)
log_w_smc = log_w_smc - log_z_smc
w_smc = np.exp(log_w_smc)
theta_rw_smc = np.random.choice(theta_samp_smc,size = B, p = w_smc, replace = True) #Sample with replacement from empirical with IS weights
## ##
# +
#Compute analytical posterior
f =plt.figure(figsize=(7,4))
theta_plot= np.arange(0,6,0.01)
a0 = np.load('plot_files/sim_a0.npy')
b0 = 1
a_post = a0 + np.sum(delta)
b_post = b0 + np.sum(t)
pdf_post= sp.stats.invgamma.pdf(theta_plot,a=a_post,scale = b_post)
plt.plot(theta_plot,pdf_post,label = 'Exact Posterior',color = 'k',linestyle = '--', alpha = 0.95)
sns.distplot(theta_rw_smc,label = 'Predictive SMC',color = 'maroon')
plt.legend()
plt.title('Posterior on Mean Parameter')
plt.xlabel(r'$\theta$')
plt.xlim((-0.25, 3.25))
f.savefig("plots/sim_param_smc_truth.pdf",bbox_inches='tight')
# -
# # Supplementary Experiments
# ## Ordering (naive importance sampling)
# +
a_samp_IS = np.load('plot_files/sim_a_samp_IS.npy')
b_samp_IS = np.load('plot_files/sim_b_samp_IS.npy')
log_w_IS = np.load('plot_files/sim_log_w_IS.npy')
a_samp_IS_ord = np.load('plot_files/sim_a_samp_IS_ord.npy')
b_samp_IS_ord = np.load('plot_files/sim_b_samp_IS_ord.npy')
log_w_IS_ord = np.load('plot_files/sim_log_w_IS_ord.npy')
# +
#IS
#Compute mean samples
theta_samp_IS = b_samp_IS/(a_samp_IS-1)
#Normalize IS weights, reweight and resample
log_z_IS = sp.special.logsumexp(log_w_IS)
log_w_IS = log_w_IS - log_z_IS
w_IS = np.exp(log_w_IS)
theta_rw_IS = np.random.choice(theta_samp_IS,size = B, p = w_IS, replace = True) #Sample with replacement from empirical with IS weights
ESS_IS = 1/np.sum(w_IS**2)
## ##
#IS ordered
#Compute mean samples
theta_samp_IS_ord = b_samp_IS_ord/(a_samp_IS_ord-1)
#Normalize IS weights, reweight and resample
log_z_IS_ord = sp.special.logsumexp(log_w_IS_ord)
log_w_IS_ord = log_w_IS_ord - log_z_IS_ord
w_IS_ord = np.exp(log_w_IS_ord)
theta_rw_IS_ord = np.random.choice(theta_samp_IS_ord,size = B, p = w_IS_ord, replace = True) #Sample with replacement from empirical with IS weights
ESS_IS_ord = 1/np.sum(w_IS_ord**2)
## ##
# +
f =plt.figure(figsize=(14,4))
plt.subplot(1,2,1)
#Compute analytical posterior
plt.plot(theta_plot,pdf_post,label = 'Exact Posterior',color = 'k',linestyle = '--', alpha = 0.6)
sns.distplot(theta_rw_IS, label = 'IS',color = 'steelblue')
plt.title('Posterior on Mean Parameter')
plt.xlabel(r'$\theta$' + '\n\n (a)')
plt.legend()
plt.subplot(1,2,2)
plt.title('Proposal on Mean Parameter')
sns.distplot(theta_samp_IS, label = 'Proposal',color = 'steelblue')
plt.xlabel(r'$\theta$'+ '\n\n (b)')
#plt.plot(theta_plot,pdf_post,label = 'Exact Posterior',color = 'k',linestyle = '--', alpha = 0.6)
plt.legend()
print('IS ESS is {}'.format(1/np.sum(w_IS**2)))
f.savefig("plots/sim_param_IS.pdf",bbox_inches='tight')
# +
f =plt.figure(figsize=(14,4))
plt.subplot(1,2,1)
#Compute analytical posterior
plt.plot(theta_plot,pdf_post,label = 'Exact Posterior',color = 'k',linestyle = '--', alpha = 0.6)
sns.distplot(theta_rw_IS_ord, label = 'IS Ordered',color = 'steelblue')
plt.xlabel(r'$\theta$'+ '\n\n (a)')
plt.title('Posterior on Mean Parameter')
plt.legend()
plt.subplot(1,2,2)
plt.title('Proposal on Mean Parameter')
sns.distplot(theta_samp_IS_ord, label = 'Proposal',color = 'steelblue')
plt.xlabel(r'$\theta$'+ '\n\n (b)')
plt.plot(theta_plot,pdf_post,label = 'Exact Posterior',color = 'k',linestyle = '--', alpha = 0.6)
plt.legend()
print('IS ordered ESS is {}'.format(1/np.sum(w_IS_ord**2)))
f.savefig("plots/sim_param_IS_ordered.pdf",bbox_inches='tight')
# -
# ## ESS and Convergence (for SMC)
# +
f =plt.figure(figsize=(14,4))
#SMC weight diagnostics
plt.subplot(1,2,1)
plt.plot(ESS_smc,label = 'ESS',color = 'k',alpha = 0.9)
plt.title('SMC Diagnostics')
n_unique = np.zeros(n+1)
for i in range(n+1):
n_unique[i] = np.shape(np.unique(particle_ind_smc[i]))[0]
plt.plot(n_unique,label = 'Unique Particle Count',color = 'k',alpha = 0.9,linestyle = '--')
plt.xlabel('Datum'+ '\n\n (a)')
plt.ylabel('Sample size')
plt.legend(loc = 3)
plt.ylim(500,2100)
plt.subplot(1,2,2)
plt.title('Predictive Resampling Diagnostics')
T = 2000
for i in range(20):
plt.plot(np.arange(n,n+T), theta_hist_smc[n:,i], color = 'k',alpha = 0.4)
plt.xlabel('Datum'+ '\n\n (b)')
plt.ylabel(r'$\bar{\theta}$')
f.savefig("plots/sim_diagnostics.pdf",bbox_inches='tight')
| run_expts/1_Simulated Plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# +
import tensorflow as tf
from tensorflow import keras
import numpy as np
import src.models.train_model as trainer
import src.models.predict_model as predictor
# -
# # Training
# ## Corn
data_dir = "../data/processed/Corn/"
train_data, train_labels = trainer._load_training_data(data_dir)
eval_data, eval_labels = trainer._load_testing_data(data_dir)
num_classes = train_labels[0].shape[0]
classifier = trainer.model(num_classes)
batch_size = 64
epochs = 20
classifier.fit(train_data, train_labels, batch_size=batch_size, epochs=epochs, validation_data=(eval_data, eval_labels))
classifier.save("../models/Corn.h5")
# ## Pepper
data_dir = "../data/processed/Pepper/"
train_data, train_labels = trainer._load_training_data(data_dir)
eval_data, eval_labels = trainer._load_testing_data(data_dir)
num_classes = train_labels[0].shape[0]
classifier = trainer.model(num_classes)
batch_size = 64
epochs = 20
classifier.fit(train_data, train_labels, batch_size=batch_size, epochs=epochs, validation_data=(eval_data, eval_labels))
classifier.save("../models/Pepper.h5")
# # Test
from tensorflow import keras
# +
import numpy as np
import os
def load_testing_data(base_dir):
"""Load testing data"""
x_test = np.load(os.path.join(base_dir, 'test_data.npy'))
y_test = np.load(os.path.join(base_dir, 'test_labels.npy'))
return x_test, y_test
# -
# ## Corn
trained_model = keras.models.load_model("../models/Corn.h5")
trained_model.summary()
test_images, test_labels = load_testing_data("../data/processed/Corn/")
loss, acc = trained_model.evaluate(test_images, test_labels, verbose=2)
# the old model
trained_model = keras.models.load_model("../models/corn.h5")
trained_model.summary()
test_images, test_labels = load_testing_data("../data/processed/Corn/")
loss, acc = trained_model.evaluate(test_images, test_labels, verbose=2)
predictions = trained_model.predict(test_images)
keras.utils.to_categorical(np.argmax(predictions[500], axis=0), 4)
test_labels[500]
len(test_labels)
# ## Pepper
trained_model = keras.models.load_model("../models/Pepper.h5")
trained_model
trained_model.summary()
test_images, test_labels = load_testing_data("../data/processed/Pepper/")
loss, acc = trained_model.evaluate(test_images, test_labels, verbose=2)
# # Predictor
import src.models.predict_model as predictor
model = predictor.load_model("../models/Corn.h5")
classes = predictor.get_classes("../data/processed/Corn/Corn_(maize)-labels.json")
classes
from src.data.image_processing import preprocess_image
data = preprocess_image("../1621.JPG")
data.shape
test_images.shape
test_images[0].shape
predictions = model.predict(test_images)
# # TensorFlow Lite
# tensorflow lite conversion: https://www.tensorflow.org/lite/convert/python_api
# https://github.com/tensorflow/tensorflow/blob/master/tensorflow/lite/examples/python/label_image.py#L62
converter = tf.lite.TFLiteConverter.from_keras_model(model)
tflite_model = converter.convert()
# +
# Load TFLite model and allocate tensors.
interpreter = tf.lite.Interpreter(model_content=tflite_model)
interpreter.allocate_tensors()
# Get input and output tensors.
input_details = interpreter.get_input_details()
output_details = interpreter.get_output_details()
# -
# Test the TensorFlow Lite model on random input data.
input_details
output_details
input_data = test_images[:1]
expected_output = test_labels[:1]
input_data
expected_output
interpreter.set_tensor(input_details[0]['index'], input_data)
interpreter.invoke()
tflite_results = interpreter.get_tensor(output_details[0]['index'])
tflite_results
tflite_results.shape
prediction = np.argmax(tflite_results[0])
# +
import json
with open("../data/processed/Corn/Corn_(maize)-labels.json") as json_file:
classes = json.load(json_file)
# -
labels = list(classes.values())
labels
with open("../data/processed/Corn/labels.txt", "w") as labels_file:
labels_file.write("\n".join(labels))
with open("../data/processed/Corn/labels.txt", "r") as f:
labels_from_file = f.readlines()
labels_from_file = list(map(str.strip, labels_from_file))
labels_from_file[prediction]
model.name
import os
os.path.basename("../models/Corn.h5")
| notebooks/3.0-agj-training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deep learning from scratch
# # Learning objectives of the notebook
# - Extend the ideas developed in the previous notebook to build a neural network using functions;
# - Implement functions for the initialization of, forward propagation through, and updating of a neural network;
# - Apply the logic of backpropagation in more detail;
# - Hand-code gradient descent using `numpy` for a more realistic size of problem.
# Having spent spent some time working on the ideas of supervised learning and getting familiar with the terminology of neural networks, let's write some code to implement a neural network from scratch. We're going to use a functional programming style to help build intuition. To make matters easier, we'll use a dictionary called `model` to store all data associated with the neural network (the weight matrices, the bias vectors, etc.) and pass that into functions as a single argument. We'll also assume that the activation functions are the same in all the layers (i.e., the *logistic* or *sigmoid* function) to simplify the implementation. Production codes usually use an object-oriented style to build networks and, of course, are optimized for efficiency (unlike what we'll develop here).
# We're going to borrow notation from <NAME>'s [*Neural Networks and Deep Learning*](http://neuralnetworksanddeeplearning.com) to make life easier. In particular, we'll let $W^\ell$ and $b^\ell$ denote the weight matrices & bias vectors respectively associated with the $\ell$th layer of the network. The entry $W^{\ell}_{jk}$ of $W^\ell$ is the weight parameter associated with the link connecting the $k$th neuron in layer $\ell-1$ to the $j$th neuron in layer $\ell$:
#
# [](http://neuralnetworksanddeeplearning.com/chap2.html)
#
# Let's put this altogether now and construct a network from scratch. We start with some typical imports.
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# ## 1. Create an initialization function to set up model
#
# Rather than the fixed constants in `setup` from before, write a function `initialize_model` that accepts a list `dimensions` of positive integer inputs that constructs a `dict` with specific key-value pairs:
# + `model['nlayers']` : number of layers in neural network
# + `model['weights']` : list of NumPy matrices with appropriate dimensions
# + `model['biases']` : list of NumPy (column) vectors of appropriate dimensions
# + The matrices in `model['weights']` and the vectors in `model['biases']` should be initialized as randomly arrays of the appropriate shapes.
#
# If the input list `dimensions` has `L+1` entries, the number of layers is `L` (the first entry of `dimensions` is the input dimension, the next ones are the number of units/neurons in each subsequent layer going up to the output layer).
# Thus, for example:
#
# ```python
# >>> dimensions = [784, 15, 10]
# >>> model = initialize_model(dimensions)
# >>> for k, (W, b) in enumerate(zip(model['weights'], model['biases'])):
# >>> print(f'Layer {k+1}:\tShape of W{k+1}: {W.shape}\tShape of b{k+1}: {b.shape}')
# ```
# ```
# Layer 1: Shape of W1: (15, 784) Shape of b1: (15, 1)
# Layer 2: Shape of W2: (10, 15) Shape of b2: (10, 1)
# ```
def initialize_model(dimensions):
'''Accepts a list of positive integers; returns a dict 'model' with key/values as follows:
model['nlayers'] : number of layers in neural network
model['weights'] : list of NumPy matrices with appropriate dimensions
model['biases'] : list of NumPy (column) vectors of appropriate dimensions
These correspond to the weight matrices & bias vectors associated with each layer of a neural network.'''
weights, biases = [], []
L = len(dimensions) - 1 # number of layers (i.e., excludes input layer)
for l in range(L):
W = np.random.randn(dimensions[l+1], dimensions[l])
b = np.random.randn(dimensions[l+1], 1)
weights.append(W)
biases.append(b)
return dict(weights=weights, biases=biases, nlayers=L)
# Use a test example to illustrate that the network is initialized as needed
dimensions = [784, 15, 10]
model = initialize_model(dimensions)
for k, (W, b) in enumerate(zip(model['weights'], model['biases'])):
print(f'Layer {k+1}:\tShape of W{k+1}: {W.shape}\tShape of b{k+1}: {b.shape}')
# Let's examine the weight matrix & bias vector associated with the second layer.
print(f'W2:\n\n{model["weights"][1]}') # Expect a 10x15 matrix of random numbers
print(f'b2:\n\n{model["biases"][1]}') # Expect a 10x1 vector of random numbers
# ## 2. Implement activation function(s), loss functions, & their derivatives
# For today's purposes, we'll use only the *logistic* or *sigmoid* function as an activation function:
# $$ \sigma(x) = \frac{1}{1+\exp(-x)} = \frac{\exp(x)}{1+\exp(x)}.$$
# A bit of calculus shows that
# $$ \sigma'(x) = \sigma(x)(1-\sigma(x)) .$$
#
# Actually, a more numerically robust formula for $\sigma(x)$ (i.e., one that works for large positive or large negative input equally well) is
# $$
# \sigma(x) = \begin{cases} \frac{1}{1+\exp(-x)} & (x\ge0) \\ 1 - \frac{1}{1+\exp(x)} & \mathrm{otherwise} \end{cases}.
# $$
# For the loss function, we'll use the typical "$L_2$-norm of the error" (alternatively called *mean-square error (MSE)* when averaged over a batch of values:
# $$ \mathcal{E}(\hat{y},y) = \frac{1}{2} \|\hat{y}-y\|^{2} = \frac{1}{2} \sum_{k=1}^{d} \left[ \hat{y}_{k}-y_{k} \right]^{2}.$$
# Again, using multivariable calculus, we can see that
# $$\nabla_{\hat{y}} \mathcal{E}(\hat{y},y) = \hat{y} - y.$$
#
# Implement all four of these functions below.
def sigma(x):
'''The logistic function; accepts arbitrary arrays as input (vectorized)'''
return np.where(x>=0, 1/(1+np.exp(-x)), 1 - 1/(1+np.exp(x))) # piecewise for numerical robustness
def sigma_prime(x):
'''The *derivative* of the logistic function; accepts arbitrary arrays as input (vectorized)'''
return sigma(x)*(1-sigma(x)) # Derivative of logistic function
def loss(yhat, y):
'''The loss as measured by the L2-norm squared of the error'''
return 0.5 * np.square(yhat-y).sum()
def loss_prime(yhat, y):
'''Implementation of the gradient of the loss function'''
return (yhat - y) # gradient w.r.t. yhat
# ## 3. Implement a function for forward propagation
#
# Write a function `forward` that uses the architecture described in a `dict` as created by `initialize_model` to evaluate the output of the neural network for a given input *column* vector `x`.
# + Take $a^{0}=x$ from the input.
# + For $\ell=1,\dotsc,L$, compute & store the intermediate computed vectors $z^{\ell}=W^{\ell}a^{\ell-1}+b^{\ell}$ (the *weighted inputs*) and $a^{\ell}=\sigma\left(z^{\ell}\right)$ (the *activations*) in an updated dictionary `model`. That is, modify the input dictionary `model` so as to accumulate:
# + `model['activations']`: a list with entries $a^{\ell}$ for $\ell=0,\dotsc,L$
# + `model['z_inputs']`: a list with entries $z^{\ell}$ for $\ell=1,\dotsc,L$
# + The function should return the computed output $a^{L}$ and the modified dictionary `model`.
# Notice that input `z` can be a matrix of dimension $n_{0} \times N_{\mathrm{batch}}$ corresponding to a batch of input vectors (here, $n_0$ is the dimension of the expected input vectors).
# Abstract process into function and run tests again.
def forward(x, model):
'''Implementation of forward propagation through a feed-forward neural network.
x : input array oriented column-wise (i.e., features along the rows)
model : dict with same keys as output of initialize_model & appropriate lists in 'weights' & 'biases'
The output dict model is the same as the input with additional keys 'z_inputs' & 'activations';
these are accumulated to be used later for backpropagation. Notice the lists model['z_inputs'] &
model['activations'] both have the same number of entries as model['weights'] & model['biases']
(one for each layer).
'''
a = x
activations = [a]
zs = []
for W, b in zip(model['weights'], model['biases']):
z = W @ a + b
a = sigma(z)
zs.append(z)
activations.append(a)
model['activations'], model['z_inputs'] = activations, zs
return (a, model)
# Use a test example to illustrate that the network is initialized as needed
dimensions = [784, 15, 10]
model = initialize_model(dimensions)
print(f'Before executing *forward*:\nkeys == {model.keys()}')
N_batch = 3 # Let's use, say, 3 random inputs & their corresponding outputs
x_input = np.random.rand(dimensions[0], N_batch)
y = np.random.rand(dimensions[-1], N_batch)
y_hat, model = forward(x_input, model) # the dict model is *updated* by forward propagation
print(f'After executing *forward*:\nkeys == {model.keys()}')
# Observe additional dict keys: 'activations' & 'z_inputs'
# ### Algorithm for backpropagation:
#
# #### (optional reading for the mathematically brave)
#
# The description here is based on the *wonderfully concise* description from <NAME>'s [*Neural Networks and Deep Learning*](http://neuralnetworksanddeeplearning.com/chap2.html). Neilsen has artfully crafted a summary using the bare minimum mathematical prerequisites. The notation elegantly summarises the important ideas in a way to make implementation easy in array-based frameworks like Matlab or NumPy. This is the best description I (Dhavide) know of that does this.
#
# In the following, $\mathcal{E}$ is the loss function and the symbol $\odot$ is the [*Hadamard product*](https://en.wikipedia.org/wiki/Hadamard_product_(matrices)) of two conforming arrays; this is simply a fancy way of writing the usual element-wise product of arrays as computed by NumPy & is sometimes called the *Schur product*. This can be reformulated in usual matrix algebra for analysis.
#
# Given a neural network with $L$ layers (not including the "input layer") described by an appropriate architecture:
#
# 1. Input $x$: Set the corresponding activation $a^{0} \leftarrow x$ for the input layer.
# 2. Feedforward: For each $\ell=1,2,\dotsc,L$, compute *weighted inputs* $z^{\ell}$ & *activations* $a^{\ell}$ using the formulas
# $$
# \begin{aligned}
# z^{\ell} & \leftarrow W^{\ell} a^{\ell-1} + b^{\ell}, \\
# a^{\ell} & \leftarrow \sigma\left( z^{\ell}\right)
# \end{aligned}.
# $$
# 3. Starting from the end, compute the "error" in the output layer $\delta^{L}$ according to the formula
# $$
# \delta^{L} \leftarrow \nabla_{a^{L}} \mathcal{E} \odot \sigma'\left(z^{L}\right)
# $$
#
# 4. *Backpropagate* the "error" for $\ell=L−1\dotsc,1$ using the formula
# $$
# \delta^{\ell} \leftarrow \left[ W^{\ell+1}\right]^{T}\delta^{\ell+1} \odot \sigma'\left(z^{\ell}\right).
# $$
# 5. The required gradients of the loss function $\mathcal{E}$ with respect to the parameters $W^{\ell}_{p,q}$ and $b^{\ell}_{r}$ can be computed directly from the "errors" $\left\{ \delta^{\ell} \right\}$ and the weighted inputs $\left\{ z^{\ell} \right\}$ according to the relations
# $$
# \begin{aligned}
# \frac{\partial \mathcal{E}}{\partial W^{\ell}_{p,q}} &= a^{\ell-1}_{q} \delta^{\ell}_{p} &&(\ell=1,\dotsc,L)\\
# \frac{\partial \mathcal{E}}{\partial b^{\ell}_{r}} &= \delta^{\ell}_{r} &&
# \end{aligned}
# $$
# ## 4. Implement a function for backward propagation
#
# **This one is a freebie!**
#
# Implement a function `backward` that implements the back-propagation algorithm to compute the gradients of the loss function $\mathcal{E}$ with respect to the weight matrices $W^{\ell}$ and the bias vectors $b^{\ell}$.
# + The function should accept a column vector `y` of output labels and an appropriate dictionary `model` as input.
# + The dict `model` is assumed to have been generated *after* a call to `forward`; that is, `model` should have keys `'w_inputs'` and `'activations'` as computed by a call to `forward`.
# + The result will be a modified dictionary `model` with two additional key-value pairs:
# + `model['grad_weights']`: a list with entries $\nabla_{W^{\ell}} \mathcal{E}$ for $\ell=1,\dotsc,L$
# + `model['grad_biases']`: a list with entries $\nabla_{b^{\ell}} \mathcal{E}$ for $\ell=1,\dotsc,L$
# + Notice the dimensions of the matrices $\nabla_{W^{\ell}} \mathcal{E}$ and the vectors $\nabla_{b^{\ell}} \mathcal{E}$ will be identical to those of ${W^{\ell}}$ and ${b^{\ell}}$ respectively.
# + The function's return value is the modified dictionary `model`.
#
#
# We've done this for you (in the interest of time). Notice that input `y` can be a matrix of dimension $n_{L} \times N_{\mathrm{batch}}$ corresponding to a batch of output vectors (here, $n_L$ is the number of units in the output layer).
def backward(y, model):
'''Implementation of backward propagation of data through the network
y : output array oriented column-wise (i.e., features along the rows) as output by forward
model : dict with same keys as output by forward
Note the input needs to have keys 'nlayers', 'weights', 'biases', 'z_inputs', and 'activations'
'''
Nbatch = y.shape[1] # Needed to extend for batches of vectors
# Compute the "error" delta^L for the output layer
yhat = model['activations'][-1]
z, a = model['z_inputs'][-1], model['activations'][-2]
delta = loss_prime(yhat, y) * sigma_prime(z)
# Use delta^L to compute gradients w.r.t b & W in the output layer.
grad_b, grad_W = delta @ np.ones((Nbatch, 1)), np.dot(delta, a.T)
grad_weights, grad_biases = [grad_W], [grad_b]
loop_iterates = zip(model['weights'][-1:0:-1],
model['z_inputs'][-2::-1],
model['activations'][-3::-1])
for W, z, a in loop_iterates:
delta = np.dot(W.T, delta) * sigma_prime(z)
grad_b, grad_W = delta @ np.ones((Nbatch, 1)), np.dot(delta, a.T)
grad_weights.append(grad_W)
grad_biases.append(grad_b)
# We built up lists of gradients backwards, so we reverse the lists
model['grad_weights'], model['grad_biases'] = grad_weights[::-1], grad_biases[::-1]
return model
# Use the test example from above. Assume model, x_input have been initialized & *forward* has been executed already.
print(f'Before executing *backward*:\nkeys == {model.keys()}')
model = backward(y, model) # the dict model is updated *again* by backward propagation
print(f'After executing *backward*:\nkeys == {model.keys()}')
# Observe additional dict keys: 'grad_weights' & 'grad_biases'
# ## 5. Implement a function to update the model parameters using computed gradients.
#
# Given some positive learning rate $\eta>0$, we want to change all the weights and biases using their gradients.
# Write a function `update` to compute a single step of gradient descent assuming that the model gradients have been computed for a given input vector.
# + The functions signature should be `update(eta, model)` where `eta` is a positive scalar value and `model` is a dictionary as output from `backward`.
# + The result will be an updated model with the values updated for `model['weights']` and `model['biases']`.
# + Written using array notations, these updates can be expressed as
# $$
# \begin{aligned}
# W^{\ell} &\leftarrow W^{\ell} - \eta \nabla_{W^{\ell}} \mathcal{E} &&(\ell=1,\dotsc,L)\\
# b^{\ell} &\leftarrow b^{\ell} - \eta \nabla_{b^{\ell}} \mathcal{E} &&
# \end{aligned}.
# $$
# + Written out component-wise, the preceding array expressions would be written as
# $$
# \begin{aligned}
# W^{\ell}_{p,q} &\leftarrow W^{\ell}_{p,q} - \eta \frac{\partial \mathcal{E}}{\partial W^{\ell}_{p,q}}
# &&(\ell=1,\dotsc,L)\\
# b^{\ell}_{r} &\leftarrow b^{\ell}_{r} - \eta \frac{\partial \mathcal{E}}{\partial b^{\ell}_{r}} &&
# \end{aligned}
# $$.
# + For safety, have the update step delete the keys added by calls to `forward` and `backward`, i.e., the keys `'z_inputs'`, `'activations'`, `'grad_weights'`, & `'grad_biases'`.
# + The output should be a dict `model` like before.
def update(eta, model):
'''Use learning rate and gradients to update model parameters
eta : learning rate (positive scalar parameter)
model : dict with same keys as output by backward
Output result is a modified dict model
'''
new_weights, new_biases = [], []
for W, b, dW, db in zip(model['weights'], model['biases'], model['grad_weights'], model['grad_biases']):
new_weights.append(W - (eta * dW))
new_biases.append(b- (eta * db))
model['weights'] = new_weights
model['biases'] = new_biases
# Get rid of extraneous keys/values
for key in ['z_inputs', 'activations', 'grad_weights', 'grad_biases']:
del model[key]
return model
# Use the test example from above. Assume *forward* & *backward* have been executed already.
print(f'Before executing *update*:\nkeys == {model.keys()}')
eta = 0.5 # Choice of learning rate
model = update(eta, model) # the dict model is updated *again* by calling *update*
print(f'After executing *update*:\nkeys == {model.keys()}')
# Observe fewer dict keys: extraneous keys have been freed.
# Observe the required sequence of executions: (forward -> backward -> update -> forward -> backward -> ...)
# If done out of sequence, results in KeyError
backward(y, model) # This should cause an exception (KeyError)
# ## 6. Implement steepest descent in a loop for random training data
#
# Let's now attempt to use our NumPy-based model to implement the steepest descent algorithm. We'll explain these numbers shortly in the context of the MNIST digit classification problem.
#
# + Generate random arrays `X` and `y` of dimensions $28^2 \times N_{\mathrm{batch}}$ and $10\times N_{\mathrm{batch}}$ respectively where $N_{\mathrm{batch}}=10$.
# + Initialize the network architecture using `initialize_model` as above to require an input layer of $28^2$ units, a hidden layer of 15 units, and an output layer of 10 units.
# + Choose a learning rate of, say, $\eta=0.5$ and a number of epochs `n_epoch` of, say, $30$.
# + Construct a for loop with `n_epochs` iterations in which:
# + The output `yhat` is computed from the input`X` using `forward`.
# + The function `backward` is called to compute the gradients of the loss function with respect to the weights and biases.
# + Update the network parameters using the function `update`.
# + Compute and print out the epoch (iteration counter) and the value of the loss function.
# +
N_batch = 10
n_epochs = 30
dimensions = [784, 15, 10]
X = np.random.randn(dimensions[0], N_batch)
y = np.random.randn(dimensions[-1], N_batch)
eta = 0.5
model = initialize_model(dimensions)
for epoch in range(n_epochs):
yhat, model = forward(X, model)
err = loss(yhat, y)
print(f'Epoch: {epoch}\tLoss: {err}')
model = backward(y, model)
model = update(eta, model)
# Expect to see loss values decreasing systematically in each iteration.
# -
# ## 7. Modify the steepest descent loop to make a plot
#
# Let's alter the preceding loop to accumulate selected epoch & loss values in lists for plotting.
#
# + Set `N_batch` and `n_epochs` to be larger, say, $50$ and $30,000$ respectively.
# + Change the preceding `for` loop so that:
# + The `epoch` counter and the loss value are accumulated into lists every, say, `SKIP` iterations where `SKIP==500`.
# + Eliminate the `print` statement(s) to save on output.
# + After the `for` loop terminates, make a `semilogy` plot to verify that the loss function is actually decreasing with sucessive epochs.
# + Use the list `epochs` to accumulate the `epoch` every 500 epochs.
# + Use the list `losses` to accumulate the values of the loss function every 500 epochs.
# +
N_batch = 50
n_epochs = 30000
SKIP = 50
dimensions = [784, 15, 10]
X = np.random.randn(dimensions[0], N_batch)
y = np.random.randn(dimensions[-1], N_batch)
eta = 0.5
model = initialize_model(dimensions)
# accumulate the epoch and loss in these respective lists
epochs, losses = [], []
for epoch in range(n_epochs):
yhat, model = forward(X, model)
model = backward(y, model)
model = update(eta, model)
if (divmod(epoch, SKIP)[1]==0):
err = loss(yhat, y)
epochs.append(epoch)
losses.append(err)
# -
# code for plotting once that the lists epochs and losses are accumulated
fig = plt.figure(); ax = fig.add_subplot(111)
ax.set_xlim([0,n_epochs]); ax.set_ylim([min(losses), max(losses)]);
ax.set_xticks(epochs[::500]); ax.set_xlabel("Epochs"); ax.grid(True);
ax.set_ylabel(r'$\mathcal{E}$');
h1 = ax.semilogy(epochs, losses, 'r-', label=r'$\mathcal{E}$')
plt.title('Loss vs. epochs');
| notebooks/2-Instructor-deep-learning-from-scratch-pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table class="table table-bordered">
# <tr>
# <th style="text-align:center; width:35%"><img src='https://dl.dropbox.com/s/qtzukmzqavebjd2/icon_smu.jpg' style="width: 300px; height: 90px; "></th>
# <th style="text-align:center;"><font size="4"> <br/>IS.215 - Practical 2 Recommenders in Python</font></th>
# </tr>
# </table>
# ### Content-based recommender system for movies
# This recommender is built using content from IMDB top 250 English movies or https://query.data.world/s/uikepcpffyo2nhig52xxeevdialfl7. The metadata used includes movie director, main actors and plot.
#
# Make sure Rake (Rapid Automatic Keyword Extraction algorithm) library is installed. If not, it can be installed via "pip install rake_nltk". Refer to https://pypi.org/project/rake-nltk/ for more information.
#
# This python script is adapted from https://towardsdatascience.com/how-to-build-from-scratch-a-content-based-movie-recommender-with-natural-language-processing-25ad400eb243
import pandas as pd
from rake_nltk import Rake
import numpy as np
from sklearn.metrics.pairwise import cosine_similarity
from sklearn.feature_extraction.text import CountVectorizer
# **Step 1: Read in and analyse input data**
df = pd.read_csv('movies2.csv')
#df = pd.read_csv('https://query.data.world/s/uikepcpffyo2nhig52xxeevdialfl7')
df.head()
df.shape
# Use the following input features to base the recommendations.
# +
df = df[['Title', 'Genre', 'Director']] # we only use these few features
# Question 1: Write the code snippet that state the feature list as Title, Genre, Director.
df.head()
# -
df.shape
# **Step 2a: Data pre-processing** Transforming the full names of actors and directors in single words so they are considered as unique values.
# +
# putting the directors in a list of words
df['Director'] = df['Director'].map(lambda x: x.split(' ')) # split by whitespace, then store as array
df['Genre'] = df['Genre'].map(lambda x: x.split(',')) # split by comma as genre is given as CSV, then store as array
# merging first and last name for each actor and director into one word
# to ensure no mix up between people sharing a first name
for index, row in df.iterrows():
row['Director'] = ''.join(row['Director']).lower()
row['Genre'] = [x.lower().replace(' ','') for x in row['Genre']]
# -
# **Step 2b: Data pre-processing on plot** Extracting the key words from the plot description.
# +
# initializing the new column
df['Key_words'] = ""
for index, row in df.iterrows():
plot = row['Plot']
# instantiating Rake, by default is uses english stopwords from NLTK
# and discard all puntuation characters
r = Rake()
# extracting the words by passing the text
r.extract_keywords_from_text(plot)
# getting the dictionary whith key words and their scores
key_words_dict_scores = r.get_word_degrees()
# assigning the key words to the new column
row['Key_words'] = list(key_words_dict_scores.keys())
# dropping the Plot column
df.drop(columns = ['Plot'], inplace = True)
#if have error - use df.drop('Plot', axis=1, inplace=True)
# -
# check the columns
df.head()
# **Step 3: Create word representation - via bag of words using the values from the columns**
# +
df['bag_of_words'] = ''
#Title should be omitted from bag of words creation
columns = df.columns[1:]
#print(columns)
for index, row in df.iterrows():
words = ''
for col in columns:
if col != 'Director':
words = words + ' '.join(row[col])+ ' '
else:
words = words + row[col]+ ' '
row['bag_of_words'] = words
df = df[['Title','bag_of_words']]
# -
df.head()
# **Step 4: Create the model using count metrics**
# +
# instantiating and generating the count matrix
count = CountVectorizer()
count_matrix = count.fit_transform(df['bag_of_words'])
# creating a Series for the movie titles so they are associated to an ordered numerical
# list that can be used to match the indexes
indices = pd.Series(df['Title'])
indices[:5]
# -
# generating the cosine similarity matrix
cosine_sim = cosine_similarity(count_matrix, count_matrix)
print(cosine_sim)
# **Step 5: Test and run the model (recommender)**
#
# Question 2: Write the code snippet that return top 5 recommended movies.
#
# Question 3: What is the top recommended movie for 'The Green Mile'?
#
#
# function that takes in movie title as input and returns the top 5 recommended movies
def recommendations(title, cosine_sim = cosine_sim):
recommended_movies = []
# getting the index of the movie that matches the title
idx = indices[indices == title].index[0]
# creating a Series with the similarity scores in descending order
score_series = pd.Series(cosine_sim[idx]).sort_values(ascending = False)
# getting the indexes of the 10 most similar movies
top_5_indexes = list(score_series.iloc[1:6].index) # why don't return index 0? because it's the movie itself!
# populating the list with the titles of the best 10 matching movies
for i in top_5_indexes:
recommended_movies.append(list(df['Title'])[i])
return recommended_movies
recommendations('The Green Mile')
| practicaltest_b2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
df = pd.read_csv('CommViolPredUnnormalizedData.txt')
df.head()
df = df.replace('?', np.nan)
# +
age_groups = ['12t21', '12t29', '16t24', '65up']
for group in age_groups:
df['ageCnt' + group] = (df['population'] * df['agePct' + group]).astype(int)
df[['population'] + ['agePct' + group for group in age_groups] + ['ageCnt' + group for group in age_groups]].head()
# -
group_state_df = df.groupby('state')
group_state_df.sum()[['ageCnt' + group for group in age_groups]]
df.describe()
# +
crime_df = df[['burglPerPop','larcPerPop','autoTheftPerPop','arsonsPerPop','nonViolPerPop']]
f, ax = plt.subplots(figsize=(13, 10))
sns.boxplot(data=crime_df)
plt.show()
# +
feature_columns = ['PctPopUnderPov', 'PctLess9thGrade', 'PctUnemployed', 'ViolentCrimesPerPop', 'nonViolPerPop']
filtered_df = df[feature_columns]
f, ax = plt.subplots(figsize=(13, 10))
sns.heatmap(filtered_df.dropna().astype(float).corr(), center=0, annot=True)
bottom, top = ax.get_ylim()
ax.set_ylim(bottom + 0.5, top - 0.5)
plt.show()
# -
| Activity01/Analyzing_the_Communities_and_Crimes_Dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from sklearn.model_selection import train_test_split
import statsmodels.api as sm
# %matplotlib inline
import matplotlib.pyplot as plt
# download weather data from here: https://drive.google.com/file/d/1fiHg5DyvQeRC4SyhsVnje5dhJNyVWpO1/view
data = pd.read_csv('Weather.csv')
data = data.fillna(0)
yVar = data['MaxTemp'].values.reshape(-1,1)
xVar = data['MinTemp'].values.reshape(-1,1)
X_train, X_test, y_train, y_test = train_test_split(xVar, yVar, test_size=0.2, random_state=0)
# +
# Note the difference in argument order
model = sm.OLS(y_train, X_train).fit()
predictions = model.predict(X_test) # make the predictions by the model
# Print out the statistics
model.summary()
# -
predictions
y_test
plt.scatter([X_test], [y_test], color='gray')
plt.plot(X_test, predictions, color='red', linewidth=2)
plt.show()
| Section 3/3.1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: neon
# language: python
# name: neon
# ---
# +
import dicom
from matplotlib import pyplot as plt, cm
import numpy as np
import cv2
# %matplotlib inline
# -
plan0 = dicom.read_file("../../dicom/ProstateX-0000/1.3.6.1.4.1.14519.5.2.1.7311.5101.158323547117540061132729905711/1.3.6.1.4.1.14519.5.2.1.7311.5101.100000082759836574166944843130/000000.dcm")
plan0.BodyPartExamined
plan0.ImagePositionPatient
plan0.Columns
rsImage = cv2.resize(plan0.pixel_array, (64,64))
# +
plt.figure(figsize=(10,10));
plt.subplot(1,2,1);
plt.imshow(plan0.pixel_array, cmap=cm.bone);
plt.title('Original')
plt.subplot(1,2,2);
plt.imshow(rsImage, cmap=cm.bone);
plt.title('Resized');
# -
plan1 = dicom.read_file("../../dicom/ProstateX-0000/1.3.6.1.4.1.14519.5.2.1.7311.5101.158323547117540061132729905711/1.3.6.1.4.1.14519.5.2.1.7311.5101.100000082759836574166944843130/000001.dcm")
plt.imshow(plan1.pixel_array, cmap=cm.bone);
plan1.ImageType
plan1.SliceThickness
plan2 = dicom.read_file("/Volumes/homes/users/anthony.reina/dicom"
"/Lung CT/stage1/00cba091fa4ad62cc3200a657aeb957e/"
"0a291d1b12b86213d813e3796f14b329.dcm")
plt.imshow(plan2.pixel_array, cmap=cm.bone);
plan2.PatientID
plan3 = dicom.read_file("/Volumes/homes/users/anthony.reina/dicom/Lung CT/stage1/00cba091fa4ad62cc3200a657aeb957e/034673134cbef5ea15ff9e0c8090500a.dcm")
plt.imshow(plan3.pixel_array, cmap=cm.bone)
np.shape(plan1.pixel_array)
plan3.Rows
| dicom/DICOM Load.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Predicting energy output of a power plant
#
# ### Import the power.csv
#
# The dataset is obtained from the [UCI Machine Learning Repository](http://archive.ics.uci.edu/ml/datasets/Combined+Cycle+Power+Plant)
#
import numpy
filename = "power.csv"
raw_data = open(filename, 'rt')
data = numpy.loadtxt(raw_data, delimiter=",")
# ### AT,EV,AP,RH,PE
#
# The dataset contains five columns, namely, Ambient Temperature (AT),Exhaust Vacuum (EV), Ambient Pressure (AP), Relative Humidity (RH), and net hourly electrical energy output (PE) of the plant. The first four are the attributes, and are used to predict the output, PE
# We check the size of the data
data.shape
# We check the first 10 rows
data[:10]
#We import the plotting library
import matplotlib.pyplot as plt
import numpy as np
from sklearn import linear_model
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.model_selection import train_test_split
#We separate out the independent variable Ambient Temperature (AT),Exhaust Vacuum (EV), Ambient Pressure (AP), Relative Humidity (RH) into X
#and dependent variable energy output into y
X=data[:,0:4]
y=data[:,4]
#We split the data into train and test using train_test_split
X_trn, X_tst, y_trn, y_tst = train_test_split(X, y, test_size=0.2, random_state=42)
print(X_trn.shape)
print(y_trn.shape)
print(X_tst.shape)
print(y_tst.shape)
# Plot outputs Ambient Temperature vs energy output
plt.scatter(X_trn[:,0], y_trn, color='red')
plt.xlabel('Ambient Temperature')
plt.ylabel('energy output')
plt.show()
# Plot outputs Exhaust Vacuum vs energy output
plt.scatter(X_trn[:,1], y_trn, color='red')
plt.xlabel('Exhaust Vacuum')
plt.ylabel('energy output')
plt.show()
# Plot outputs Ambient Pressure (AP) vs energy output
plt.scatter(X_trn[:,2], y_trn, color='red')
plt.xlabel('Ambient Pressure')
plt.ylabel('energy output')
plt.show()
# Plot outputs Relative Humidity vs energy output
plt.scatter(X_trn[:,3], y_trn, color='red')
plt.xlabel('Relative Humidity')
plt.ylabel('energy output')
plt.show()
# Create linear regression object
regr = linear_model.LinearRegression()
# Train the model using the training sets
regr.fit(X_trn, y_trn)
# Make predictions using the testing set
y_pred = regr.predict(X_tst)
# The coefficients
print('Coefficients: \n', regr.coef_)
# The mean squared error
print("Mean squared error: %.2f"
% mean_squared_error(y_tst, y_pred))
# Explained variance score: 1 is perfect prediction
print('Variance score: %.2f' % r2_score(y_tst, y_pred))
# +
# Plot outputs
plt.scatter(X_tst[:,0], y_tst, color='black')
plt.plot(X_tst[:,0], y_pred, color='blue', linewidth=3)
plt.xticks(())
plt.yticks(())
plt.show()
| Section 3/predicting_energy_output.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This code crawls Instagram accounts and store information on bio, follows, followers, amount of posts, last 100 posts, and last 100 comments for each of posts retrieved. Then the info can be stored and viewed as JSON.
# +
#DEPENDENCIES
from instaparser.agents import AgentAccount
from instaparser.entities import Account, Media, Location, Tag, Comment
import instagram_explore as ie
#signup in your account AgentAccount("username", "password")
agent = AgentAccount("username", "password")
#all information is stored in this dict
data = dict()
#what users to parse. Set account names followed after this https://www.instagram.com/TARGET_ACCOUNT_NAME/
accounts = ["SET_ACCOUNT_NAME", "SET_ACCOUNT_NAME"]
#process
for account_name in accounts:
account = Account(account_name)
agent.update(account)
data[account_name] = []
#general info
data[account_name].append({'biography' : account.biography})
data[account_name].append({'follows' : account.follows_count})
data[account_name].append({'followers' : account.followers_count})
data[account_name].append({'amount_of_posts' : account.media_count})
data[account_name].append({'100_posts' : []})
#post specific info
if account.media_count >= 100:
media, pointer = agent.get_media(account, count=100)
else:
media, pointer = agent.get_media(account, count=int(account.media_count))
c = 0
for url_media in media:
data[account_name][-1]['100_posts'].append({'post#' + str(c) : [] })
data[account_name][-1]['100_posts'][c]['post#' + str(c)].append({'url' : 'https://www.instagram.com/p/' + str(url_media) })
data[account_name][-1]['100_posts'][c]['post#' + str(c)].append({'caption' : url_media.caption })
data[account_name][-1]['100_posts'][c]['post#' + str(c)].append({'likes' : url_media.likes_count })
data[account_name][-1]['100_posts'][c]['post#' + str(c)].append({'comments' : [] })
res = ie.media(str(url_media))
if url_media.comments_count < 100:
for i in res.data['edge_media_to_comment']['edges']:
data[account_name][-1]['100_posts'][c]['post#' + str(c)][-1]['comments'].append(i['node']['text'])
# print(i['node']['owner']['username'])
else:
for i in range(100):
data[account_name][-1]['100_posts'][c]['post#' + str(c)][-1]['comments'].append(res.data['edge_media_to_comment']['edges']
[i]['node']['text'])
c += 1
# +
# write in JSON
import json
with open('data.txt', 'w') as outfile:
json.dump(data, outfile)
| Instagram_parser_working.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Gerekli Kütüphanelerin Yüklenmesi
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.models import Sequential
from keras.layers import Dense,Dropout,BatchNormalization,Activation
# # Veri Setinin Yüklenmesi
# ### NOT:Yapay sinir ağları kısmında makine öğrenmesinde veri ön işleme adımları uygulanan veri seti kullanılmıştır.
veri_seti = pd.read_csv("new_COVID-19_Dataset.csv")
veri_seti.head()
veri_seti.describe()
veri_seti.columns
veri_seti.isnull().sum()
# # Özniteliklerin Bağımlı ve Bağımsız Değişkenler Olarak Ayrılması
X = veri_seti.drop(['COVID-19'],axis=1)
Y = veri_seti['COVID-19']
# # Veri Setinin Eğitim ve Test Olarak Bölünmesi
X_train,X_test,Y_train,Y_test = train_test_split(X,Y,test_size = 0.3,random_state = 42)
# # YSA Mimarisinin Oluşturulması
# +
model=Sequential()
model.add(Dense(128,input_dim=13))
model.add(Activation("relu"))
model.add(BatchNormalization())
model.add(Dense(64))
model.add(Activation("relu"))
model.add(BatchNormalization())
model.add(Dropout(0.4))
model.add(Dense(32))
model.add(Activation("relu"))
model.add(BatchNormalization())
model.add(Dropout(0.3))
model.add(Dense(16))
model.add(Activation("relu"))
model.add(BatchNormalization())
model.add(Dense(128))
model.add(Activation("relu"))
model.add(BatchNormalization())
model.add(Dropout(0.4))
model.add(Dense(64))
model.add(Activation("relu"))
model.add(BatchNormalization())
model.add(Dense(1, activation='sigmoid'))
model.summary()
# -
# # Modelin Derlenmesi
model.compile(optimizer="adam",
loss="binary_crossentropy",
metrics=["acc"])
# # Modelin Eğitilmesi
history=model.fit(X_train,Y_train,
validation_data=(X_test, Y_test),
batch_size=128,
epochs=100,
verbose=1,
shuffle=True)
# # Sonuçların Grafikle Gösterilmesi
plt.figure(figsize=(14,3))
plt.subplot(1, 2, 1)
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.subplot(1, 2, 2)
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# # Model Sınıflandırma Başarımı
model.evaluate(X_test,Y_test)
# # Modele Tahmin Yaptırma
symptom_list1 = np.array([[1,1,1,0,0,0,0,1,1,1,0,1,1]])
x = np.argmax(model.predict(symptom_list1), axis=-1)
if(x == 1):
print("POZİTİF : ",x)
else:
print("NEGATİF : ",x)
symptom_list2 = np.array([[1,0,0,1,1,0,1,0,0,0,0,1,1]])
x = np.argmax(model.predict(symptom_list2), axis=-1)
if(x == 1):
print("POZİTİF : ",x)
else:
print("NEGATİF : ",x)
# # Ağırlık Değerlerini ve Model Konfigürasyonunu .h5 formatında saklayalım.
model.save("model/covid19_model.h5")
| COVID-19 Disease Prediction/COVID-19_Disease_Prediction_DL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
np.set_printoptions(suppress=True)
from sklearn.datasets import make_classification
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('fivethirtyeight')
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
from tensorflow.keras.utils import to_categorical
# -
# generate some data
X, y = make_classification(n_samples=10000, n_features=20, random_state=42)
X
y
# train test split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42, stratify=y)
# standard scale for any model that uses gradient descent
ss = StandardScaler()
X_train_sc = ss.fit_transform(X_train)
X_test_sc = ss.transform(X_test)
# # Topology
# * an input layer
# * one hidden layer
# * relu activation function for all hidden layers
# * sigmoid activation function for output layer to return a value between 0 and 1
# +
model = Sequential()
model.add(Dense(32, # let's choose 32 neurons in the first layer
input_shape=(20,), # 20 features
activation='relu'))
model.add(Dense(1, activation='sigmoid'))
# compile model
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
# fit model
model.fit(X_train_sc, y_train, validation_data=(X_test_sc, y_test), epochs=10, batch_size=512)
# +
# visualization
history = model.fit(X_train_sc, y_train, validation_data=(X_test_sc, y_test), epochs=10, batch_size=512, verbose=0)
plt.plot(history.history['loss'], label='Train loss')
plt.plot(history.history['val_loss'], label='Val Loss')
plt.legend();
# -
plt.plot(history.history['accuracy'], label='Train accuracy')
plt.plot(history.history['val_accuracy'], label='Val accuracy')
plt.legend();
| linear_regression_nn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:capstone] *
# language: python
# name: conda-env-capstone-py
# ---
# # Grouping of USA states
# +
# sub_df = df.loc[df['column_name'].isin(some_values)]
# e.g. us_ne = US_state.loc[US_state["state_code"].isin(us_ne_names)]
# -
# ### Northeast (ne)
# Maine (me), New Hampshire (nh), Vermont (vt), New York (ny), Massachusetts (ma), Rhode Island (ri), Connecticut (ct), Pensylvania (pa), New Jersey (nj), Delaware (de), Maryland (md), Virgina (va), West Virgina (wv), Disrict of Columbia (dc)
#
# ### Pacific West (pw)
# Alaska (ak), Hawaii (hi), Washington (wa), Oregon (or), California (ca), Idaho (id), Nevada (nv), Arizona (az), Utah (ut)
#
# ### Midwest (mw)
# Minnesota (mn), Iowa (ia), Missouri (mo), Wisconsin (wi), Illinois (il), Michigan (mi), Indiana (in), Kentucky (ky), Ohio (oh)
#
# ### Southeast (se)
# Arkansas (ar), Louisiana (la), Mississippi (ms), Tennessee (tn), Alabama (al), North Carolina (nc), South Carolina (sc), Georgia (ga), Florida (fl), Puerto Rico (pr)
#
# ### Central (c)
# Montana (mt), North Dakota (nd), South Dakota (sd), Wyoming (wy), Nebraska (ne), Colorado (co), Kansas (ks), New Mexico (nm), Oklahoma (ok), Texas (tx)
us_ne_names = ["me", "nh", "vt", "ny", "ma", "ri", "ct", "pa", "nj", "de", "md", "va", "wv", "dc"]
us_pw_names = ["ak", "hi", "wa", "or", "ca", "id", "nv", "az", "ut"]
us_mw_names = ["mn", "ia", "mo", "wi", "il", "mi", "in", "ky", "oh"]
us_se_names = ["ar", "la", "ms", "tn", "al", "nc", "sc", "ga", "fl", "pr"]
us_c_names = ["mt", "nd", "sd", "wy", "ne", "co", "ks", "nm", "ok", "tx"]
| groupings/US_state_groupings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# <a href="https://colab.research.google.com/github/ai-fast-track/timeseries/blob/master/nbs/univariate_timeseries_Tutorial.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# # Train univariate time series
# > A tutorial that can be run in Google Colab or on a local machine
# Run this cell to install the latest version of fastcore shared on github
# !pip install git+https://github.com/fastai/fastai2.git
# Run this cell to install the latest version of fastcore shared on github
# !pip install git+https://github.com/fastai/fastcore.git
# Run this cell to install the latest version of timeseries shared on github
# !pip install git+https://github.com/ai-fast-track/timeseries.git
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
from fastai2.basics import *
# hide
# Only for Windows users because symlink to `timeseries` folder is not recognized by Windows
import sys
sys.path.append("..")
from timeseries.all import *
# ## ECG Dataset
# > This dataset was formatted by <NAME> as part of his thesis
# “Generalized feature extraction for structural pattern recognition
# in time-series data,” at Carnegie Mellon University, 2001. Each
# series traces the electrical activity recorded during one
# heartbeat. The two classes are a normal heartbeat and a Myocardial
# Infarction. Cardiac ischemia refers to lack of blood flow and oxygen to the heart muscle. If ischemia is severe or lasts too long, it can cause a heart attack (myocardial infarction) and can lead to heart tissue death.
#
# 
dsname = 'ECG200' # 'ChlorineConcentration', 'Yoga', ECG_200
# url = 'http://www.timeseriesclassification.com/Downloads/Yoga.zip'
path = unzip_data(URLs_TS.UNI_ECG200)
path
fname_train = f'{dsname}_TRAIN.arff'
fname_test = f'{dsname}_TEST.arff'
fnames = [path/fname_train, path/fname_test]
fnames
data = TSData.from_arff(fnames)
print(data)
items = data.get_items()
seed = 42
splits = RandomSplitter(seed=seed)(range_of(items)) #by default 80% for train split and 20% for valid split are chosen
splits
bs = 32
# Normalize at batch time
tfm_norm = Normalize(scale_subtype = 'per_sample_per_channel', scale_range=(0, 1)) # per_sample , per_sample_per_channel
# tfm_norm = Standardize(scale_subtype = 'per_sample')
batch_tfms = [tfm_norm]
default_device()
lbl_dict = dict([
('-1', 'Normal'),
('1', 'Myocardial Infarction')]
)
# dls = TSDataLoaders.from_files(fnames=fnames, batch_tfms=batch_tfms, num_workers=0, device=default_device())
dls = TSDataLoaders.from_files(fnames=fnames, lbl_dict=lbl_dict, num_workers=0, device=default_device())
dls.show_batch(max_n=9)
# ## Training Model
# Number of channels (i.e. dimensions in ARFF and TS files jargon)
c_in = get_n_channels(dls.train) # data.n_channels
# Number of classes
c_out= dls.c
c_in,c_out
model = inception_time(c_in, c_out).to(device=default_device())
model
# opt_func = partial(Adam, lr=3e-3, wd=0.01)
#Or use Ranger
def opt_func(p, lr=slice(3e-3)): return Lookahead(RAdam(p, lr=lr, mom=0.95, wd=0.01))
# +
#Learner
loss_func = LabelSmoothingCrossEntropy()
learn = Learner(dls, model, opt_func=opt_func, loss_func=loss_func, metrics=accuracy)
print(learn.summary())
# -
lr_min, lr_steep = learn.lr_find()
lr_min, lr_steep
learn.fit_one_cycle(25, lr_max=1e-3)
# ## Graphs
learn.recorder.plot_loss()
learn.show_results(max_n=9)
interp = ClassificationInterpretation.from_learner(learn)
interp.plot_confusion_matrix()
interp.most_confused()
# 
# <a href="https://unsplash.com/@clearsky?utm_medium=referral&utm_campaign=photographer-credit&utm_content=creditBadge">Photo Credit: <NAME></a>
| nbs/univariate_timeseries_Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Try Pytorch with Digit Recognition
#
# * Reference: https://www.analyticsvidhya.com/blog/2018/02/pytorch-tutorial/
# * <b>Some functions are wrong or out of dated which will cause so much troubles. Better to try my code below</b>.
# * However, since these deep learning open sources keep updating and deprecating functions, you will never know whether a previous tutorial works for you at the time when you try. This is one of the reasons I hate deep learning.
# * To compare with Keras code in Digit Recognition: https://github.com/hanhanwu/Hanhan_Data_Science_Practice/blob/master/AI_Experiments/digital_recognition_Keras.ipynb
# +
# %pylab inline
import os
import numpy as np
import pandas as pd
import imageio as io
from sklearn.metrics import accuracy_score
import torch
# Get data from here: https://datahack.analyticsvidhya.com/contest/practice-problem-identify-the-digits/
# -
seed = 10
rng = np.random.RandomState(seed)
train = pd.read_csv('Train_digits/train.csv')
train.head()
# +
# randomly display an image
img_name = rng.choice(train.filename)
training_image_path = 'Train_digits/Images/train/' + img_name
training_img = io.imread(training_image_path, as_gray=True)
pylab.imshow(training_img, cmap='gray')
pylab.axis('off')
pylab.show()
# -
# This is just 1 image
print(training_img.shape)
training_img[0] # each image has 28x28 pixel square, 784 pixels in total
# +
# store all images as numpy arrays, to make data manipulation easier
temp = []
for img_name in train.filename:
training_image_path = 'Train_digits/Images/train/' + img_name
training_img = io.imread(training_image_path, as_gray=True) # !!! as_gray param makes a difference here!!
img = training_img.astype('float32')
temp.append(img)
train_x = np.stack(temp)
train_x /= 255.0
train_x = train_x.reshape(-1, 784).astype('float32') # 784 pixels per image
train_y = train.label.values
# -
# With `as_gray` param in `io.imread()`, it will help you get train_x, train_y have the same length. Otherwise there will be so much troubles in creating batches later.
print(train_x.shape)
train_x
print(train_y.shape) # 49000 images in total
train_y
# +
# create validation set
split_size = int(train_x.shape[0]*0.7)
train_x, val_x = train_x[:split_size], train_x[split_size:]
train_y, val_y = train_y[:split_size], train_y[split_size:]
print(train_x.shape, train_y.shape)
print(train_x)
print(train_y)
# +
# Using Pytorch to build the model
from torch.autograd import Variable
## number of neurons in each layer
input_num_units = 28*28 # 784 pixels per image
hidden_num_units = 500
output_num_units = 10 # 0 - 9, 10 digits
## set variables used in NN
epochs = 5
batch_size = 128
learning_rate = 0.001
# +
# define model
model = torch.nn.Sequential(
torch.nn.Linear(input_num_units, hidden_num_units),
torch.nn.ReLU(),
torch.nn.Linear(hidden_num_units, output_num_units),
)
loss_fn = torch.nn.CrossEntropyLoss()
# define optimization algorithm
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
# +
# preprocess a batch of dataset
def preproc(unclean_batch_x):
"""Convert values to range 0-1"""
temp_batch = unclean_batch_x / unclean_batch_x.max()
return temp_batch
# create a batch
def batch_creator(batch_size):
dataset_name = 'train'
dataset_length = eval(dataset_name+'_x').shape[0]
batch_mask = rng.choice(dataset_length, batch_size)
batch_x = eval(dataset_name+'_x')[batch_mask]
batch_x = preproc(batch_x)
batch_y = eval(dataset_name+'_y')[batch_mask] # train_x, train_y has the same length
return batch_x, batch_y
# +
# train network
total_batch = int(train.shape[0]/batch_size)
for epoch in range(epochs):
avg_cost = 0
for i in range(total_batch):
# create batch
batch_x, batch_y = batch_creator(batch_size)
# pass that batch for training
x, y = Variable(torch.from_numpy(batch_x)), Variable(torch.from_numpy(batch_y), requires_grad=False)
pred = model(x)
# get loss
loss = loss_fn(pred, y)
# perform backpropagation
loss.backward()
optimizer.step()
avg_cost += loss.data/total_batch
print(epoch, avg_cost)
# +
# get training accuracy
x, y = Variable(torch.from_numpy(preproc(train_x))), Variable(torch.from_numpy(train_y), requires_grad=False)
pred = model(x)
final_pred = np.argmax(pred.data.numpy(), axis=1)
accuracy_score(train_y, final_pred)
# +
# get validation accuracy
x, y = Variable(torch.from_numpy(preproc(val_x))), Variable(torch.from_numpy(val_y), requires_grad=False)
pred = model(x)
final_pred = np.argmax(pred.data.numpy(), axis=1)
accuracy_score(val_y, final_pred)
# -
# ## Summary
#
# * Again, a simple deep learning problem cost me so much time to debug because of the updates in these open sources!
# * Simple param makes a difference, in this case, `imread()` from scipy got deprecated, you have to change to `imageio.imread()`, which replaced "flatten" with "as_gray", without this param, your train_x, train_y will get different length after converting to numpy array, which will cause so much trouble when creating batches later.
# * This is why I hate deep learing, software updates, and so much dimentions conversion, difficult to find problems and time consuming to debug.
# * For this case, comparing with Keras, I think Pytorch is more complex to use, especially when thinking about it cost me so much time to debug when creating batches, while Keras will just do that for you! Not sure why many people are crazy about Pytorch (maybe it's better than Tensorflow...)
# * When building the model, the neural network structure is easier to understand than keras, since it will show you the order between layers and the activation functions.
| AI_Experiments/digit_recognition_Pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] #student=false tags=["include"]
# # Biogeography Notebook 2- Mapping Responses to Environmental Change
#
# + [markdown] tags=["ignore"]
# Previously in Notebook 1 we explored how to set up functions and pull data from a database. But pulling data usually isn't enough to answer questions or test hypotheses in Biogeography.
#
# The goal of this notebook is to access and integrate diverse data sets to **visualize correlations and discover patterns** to address questions of species’ responses to environmental change. We will use programmatic tools to show how to use Berkeley resources such as the biodiversity data from biocollections and online databases, field stations, climate models, and other environmental data.
#
# This notebook is a continuation of [Biogeography Notebook 1 (GBIF API)](
# http://datahub.berkeley.edu/user-redirect/interact?account=ds-modules&repo=IB-ESPM-105&branch=master&path=fall2019/notebook1.ipynb). While we completed the Geonomic Notebook before this one, Geonomics is its own module. Sorry for any confusion about the numbering!
#
# If you have any questions getting the Jupyter notebook to run, try dropping into [data peer consulting](https://data.berkeley.edu/education/data-peer-consulting).
#
# ## Table of Contents
#
# 3 - [Mapping](#mapping)
#
# 4 - [Comparing California Oak Species](#oak)
#
# 5 - [Cal-Adapt](#caladapt)
# + [markdown] #student=false tags=["ignore"]
# ## Helpful Reminders: ###
#
# ### Text cells
# In a notebook, each rectangle containing text or code is called a *cell*.
#
# Text cells (like this one) can be edited by double-clicking on them. They're written in a simple format called [Markdown](http://daringfireball.net/projects/markdown/syntax) to add formatting and section headings. You don't need to learn Markdown, but you might want to.
#
# After you edit a text cell, click the "run cell" button at the top to confirm any changes. (Try not to delete the instructions of the lab.)
#
# **The only text cells that need to be modified are labeled "YOUR RESPONSE HERE" and are right below yellow question boxes. To edit a response, double click on YOUR RESPONSE HERE and type in your answer. Afterwards, run the cell with Shift-Enter.**
#
# ### Code cells
# Other cells contain code in the Python 3 language. Running a code cell will execute all of the code it contains.
#
# To run the code in a code cell, first click on that cell to activate it. It'll be highlighted with a little green or blue rectangle. Next, either press the Run button or hold down the `shift` key and press `return` or `enter`.
#
# The only code cells that need to be modified are right below a blue exercise box.
#
# ### Comments
# Comments are statements in English that the computer ignores. We use comments to explain what the surrounding code does. Comments appear in green after the `#` symbol like below:
# + #student=false tags=["ignore"]
1 + 2 # After you run this, you should see 3 as the output
# + [markdown] #student=false tags=["ignore"]
# Run this cell to set up the programming environment. It will take a few seconds. **Remember, if you refresh or restart the notebook, you will need to run this code again for any of the functions below to work!**
# + #student=false tags=["ignore"]
# %%capture
# !pip install --no-cache-dir shapely
# !pip install -U folium
# %matplotlib inline
import pandas as pd
import folium
import json
from pandas import json_normalize
from shapely.geometry import Point, mapping
from shapely.geometry.polygon import Polygon
from shapely import geometry as sg, wkt
from scripts.espm_module import *
from IPython.core.display import display, HTML
import matplotlib.pyplot as plt
import otter
from otter.export import export_notebook
grader = otter.Notebook()
plt.style.use('seaborn')
# + [markdown] #student=false tags=["ignore"]
# # Part 3: Mapping Species Distribution <a id='mapping'></a>
# + [markdown] #student=false tags=["ignore"]
# In programming, we often reuse chunks of code. So instead of copy/pasting it and repeating the same code over and over again, we have something called a **function**, which gives a name to a block of code. This allows us to just call the function instead of rewriting code we used before. Our use of functions later in this notebook is complex, and we will use them in order to reduce the amount of code in this notebook. For now, you can just ignore the details and structure of how functions work. Just remember that a **function** is a shortcut to easily re-run old code and that the `def` keyword means we are creating a function.
# + [markdown] #student=false tags=["ignore"]
# For our first exercise, we are going to use some of the functions we explored in Notebook 1 to pull information and records about an organism, then take it one step further and map where each of these records was collected.
# + [markdown] #student=false tags=["ignore"]
# These functions get the species record from the API (like in Part 2 of Notebook 1). The function `get_species_record` gives us the raw/unorganized records, while `get_species_records_df` gives us the data in a **DataFrame** (table of data). It uses the same commands as we used previously in Notebook 1.
# + #student=false tags=["ignore"]
def get_species_records(scientific_name):
req = GBIFRequest() # creating a request to the API
params = {'scientificName': scientific_name} # setting our parameters (the specific species we want)
pages = req.get_pages(params) # using those parameters to complete the request
records = [rec for page in pages for rec in page['results'] if rec.get('decimalLatitude')] # sift out valid records
return records
def get_species_records_df(scientific_name):
records = get_species_records(scientific_name) # Get the records using the function above
records_df = json_normalize(records) # Convert the raw records into a DataFrame
return records_df
# + [markdown] #student=false tags=["ignore"]
# This creates the **DataFrame** (table of data) we used in Notebook 1 using one of the functions we defined in the cell above. It will take a few seconds to get the records from the API.
# + #student=false tags=["ignore"]
argia_agrioides_df = get_species_records_df('Argia agrioides')
argia_agrioides_df.head() # Show the first 5 records
# + [markdown] #student=false tags=["ignore"]
# Now that we have a table of records, let's take it one step further. Our ultimate goal is to map species distribution, and we also want to retain information about the collection records. Just like in Notebook 1, we will use _Argia agrioides_ as our example organism.
#
# Since we are about to map all of the _Argia agrioides_ specimen by their collection, let's assign each collection a color. These colors are chosen randomly each time the cell is run so you can re-run the cell if you don't like them.
# + #student=false tags=["ignore"]
color_dict, html_key = assign_colors(argia_agrioides_df, 'collectionCode')
display(HTML(html_key))
# + [markdown] #student=false tags=["ignore"]
# In order to generate a map, we are going to use an existing tool called Folium. Folium is a useful library for generating map visualizations. Here, we create a function that handles the Folium mapping for us.
# + #student=false tags=["ignore"]
# This function generates a map visualization using data from species_df and child (if a value is given)
# Grouping Criteria tells Folium how to group specimen by color (ex. by collection or by species)
# Child is any secondary data we want to display (ex. UC Reserve boundaries)
def map_species_with_folium(species_df, grouping_criteria, child=None):
map = folium.Map(location=[37.359276, -122.179626], zoom_start=5) # Creates the starting map location & zoom
if child: # If a child is given, add it to the map
map.add_child(child)
for r in species_df.iterrows(): # For ever specimen in the species record, do the following:
lat, long = r[1]['decimalLatitude'], r[1]['decimalLongitude'] # Get the specimen latitude/longitude
# Add the specimen to the map
folium.CircleMarker((lat, long), color=color_dict[r[1][grouping_criteria]]).add_to(map)
return map
# + [markdown] #student=false tags=["ignore"]
# Let's map the _Argia agrioides_ specimen distribution using the function we just created.
# + #student=false tags=["ignore"]
argia_agrioides_map = map_species_with_folium(argia_agrioides_df, 'collectionCode')
argia_agrioides_map
# + [markdown] #student=false tags=["ignore"]
# ### **Concept check:** *What kinds of questions and hypotheses can we explore using this kind of information? Why might mapping species distribution be important?*
# + [markdown] #student=false tags=["ignore"]
# ### **Adding layers of information**
#
# Additional map layers can provide a wealth of information, including land boundaries, topography, average temperature, etc. Now that we have the general species range, let's map the distribution of _Argia agrioides_ with the boundaries of UC Reserves.
#
# To get the boundaries for all the reserves, we will need to send a request to get GeoJSON, which is a format for encoding a variety of geographic data structures. With this code, we can request GeoJSON for all reserves and plot ocurrences of the species.
#
# First, we'll assign the API URL that has the data to a new variable `url`. Then, we make the requests just like we did earlier through the GBIF. You'll see a huge mess of mostly numbers. This is a JSON of all the UC Reserves and the coordinates of their boundaries.
# + #student=false tags=["ignore"]
url = 'https://ecoengine.berkeley.edu/api/layers/reserves/features/'
reserves = requests.get(url, params={'page_size': 30}).json()
reserves
# + [markdown] #student=false tags=["ignore"]
# There are some reserves that the EcoEngine didn't catch. We'll add the information for "Blodgett", "Hopland", and "Sagehen" manually.
# + #student=false tags=["ignore"]
station_urls = {
'Blodgett Reserve': 'https://raw.githubusercontent.com/BNHM/spatial-layers/master/wkt/BlodgettForestResearchStation.wkt',
'Hopland Reserve': 'https://raw.githubusercontent.com/BNHM/spatial-layers/master/wkt/HoplandResearchAndExtensionCenter.wkt',
'Sagehen Reserve': 'https://raw.githubusercontent.com/BNHM/spatial-layers/master/wkt/SagehenCreekFieldStation.wkt'
}
reserves['features'] += [{'type': 'Feature', 'properties': {'name': name}, 'geometry':
mapping(wkt.loads(requests.get(url).text))} for name, url in station_urls.items()]
# + [markdown] #student=false tags=["ignore"]
# This code goes through our list of reserves and outputs their names. Make sure "Blodgett", "Hopland", and "Sagehen" are included!
# + #student=false tags=["ignore"]
[r['properties']['name'] for r in reserves['features']]
# + [markdown] #student=false tags=["ignore"]
# We can send this `geojson` directly to our mapping library `folium`. We already defined a function to do this for us, so the code is much shorter. **You'll have to zoom in, but you should see blue outlined areas.** Those are the reserves!
# + #student=false tags=["ignore"]
reserve_points = folium.features.GeoJson(reserves) # This tells Folium our reserve boundaries
argia_agrioides_and_reserves_map = map_species_with_folium(argia_agrioides_df, 'collectionCode', child=reserve_points)
argia_agrioides_and_reserves_map
# + [markdown] #student=false tags=["ignore"]
# **To answer the question, double click on YOUR RESPONSE HERE. Then run the cell afterwards.**
# + [markdown] #student=false tags=["include"]
# <!-- BEGIN QUESTION -->
#
# <div class="alert alert-block alert-warning">
# <b>QUESTION 1:</b>
# <br />
# The UC Reserves are a tremendous resource for researchers and students. You can zoom in to make the reserve boundaries more visible and see the geographic characteristics of each reserve.
# <br />
# Find one reserve where <i>A. agrioides</i> was collected. Do the characteristics of the reserve fit with what you know about the biology of <i>Agria agrioides</i> (mainly lower elevation, riparian zone). Is there another reserve that also seems like it is a suitable habitat?
# </div>
# + [markdown] #student=true tags=["include"]
#
#
# YOUR RESPONSE HERE
#
# <!-- END QUESTION -->
# + [markdown] #student=false tags=["ignore"]
# **Now that we've mapped the _Agria agrioides_ specimen, let's do that with a different species.** Think about a terrestrial California or North American species that you are interested in!
# + [markdown] #student=false tags=["include"]
# <!-- BEGIN QUESTION -->
#
# <div class="alert alert-block alert-info">
# <b>EXERCISE 1:</b>
# <br />
# Pick a species and replace ... with its scientific name. Make sure to add single quotation marks around the name! Make sure to select a terrestrial species (sorry, no marine species this time!)
#
# _Remember, your organism should be an individual species. The name should consist of a genus (Argia) and a species (agrioides)._
# </div>
#
# **Hint:** Here's what the code looks like if we used _Argia agrioides_ again:
# ```
# my_species_df = get_species_records_df('Argia agrioides')
# my_species_df.head()
# ```
# + #student=false tags=["include"]
# print pdf
my_species_df = get_species_records_df('...')
my_species_df.head() # Show the first 5 records
# + [markdown] #student=false tags=["ignore"]
# <!-- END QUESTION -->
#
# If the output above doesn't contain a table, that means either you didn't enter a name or the scientific name isn't in the database. Make sure you typed it correctly without abbreviating the species name. You might also have to use a different capitalization. If you have the correct species information but still no records, select another species.
# + [markdown] #student=false tags=["include"]
# <!-- BEGIN QUESTION -->
#
#
# <div class="alert alert-block alert-info">
# <b>EXERCISE 2:</b>
# <br />
# Assign colors to each collection by replacing ... with the name of the DataFrame we just created (my_species_df). Make sure you <b>don't</b> add quotation marks this time! Also, be careful to not accidentally delete the comma!
# </div>
#
# **Hint:** Here's what the code looks like with the `argia_agrioides_df` DataFrame:
# ```
# color_dict, html_key = assign_colors(argia_agrioides_df, 'collectionCode')
# display(HTML(html_key))
# ```
# + #student=true tags=["include"]
# print in pdf
color_dict, html_key = assign_colors(..., 'collectionCode')
display(HTML(html_key))
# + [markdown] #student=false tags=["ignore"]
# <!-- END QUESTION -->
#
#
# Let's map your species with Folium!
# + [markdown] #student=false tags=["include"]
# <!-- BEGIN QUESTION -->
#
#
# <div class="alert alert-block alert-info">
# <b>EXERCISE 3:</b>
# <br />
# Now let's map your species and add the UC Reserve boundaries. Replace ... with the name of the DataFrame we just created (my_species_df). Make sure you <b>don't</b> add quotation marks this time! Also, be careful to not accidentally delete the comma!
# </div>
#
# **Hint:** Here's what the code looks like with the `argia_agrioides_df` DataFrame:
# ```
# reserve_points = folium.features.GeoJson(reserves) # Adds reserve boundaries
# my_species_map = map_species_with_folium(argia_agrioides_df, 'collectionCode', child=reserve_points)
# my_species_map
# ```
# + #student=false tags=["include"]
#print out pdf
reserve_points = folium.features.GeoJson(reserves) # Adds reserve boundaries
my_species_map = map_species_with_folium(..., 'collectionCode', child=reserve_points)
my_species_map
# + [markdown] #student=false tags=["include"]
# <!-- END QUESTION -->
#
# <!-- BEGIN QUESTION -->
#
#
#
# <div class="alert alert-block alert-warning">
# <b>QUESTION 2:</b>
# <br />
# Make some inferences about the biology of your mapped organism from the mapped distribution. Consider the end of the species range. What conditions (both biotic and abiotic) might be limiting the range?
# </div>
# + [markdown] #student=true tags=["include"]
#
# YOUR RESPONSE HERE
#
# <!-- END QUESTION -->
# + [markdown] #student=false tags=["ignore"]
# Now let's go back to looking at _Argia argrioides_.
#
# We can also find out which stations have how many _Argia argrioides_. First we'll have to add a column to our DataFrame that makes points out of the latitude and longitude coordinates.
# + #student=false tags=["ignore"]
station_df = argia_agrioides_df
def make_point(row):
return Point(row['decimalLongitude'], row['decimalLatitude'])
station_df['point'] = station_df.apply(lambda row: make_point (row), axis=1)
# + [markdown] #student=false tags=["ignore"]
# Now we can write a little function to check whether that point is in one of the stations, and if it is, we'll add that station in a new column called `station`. Then we'll apply that function the DataFrame.
# + #student=false tags=["ignore"]
def in_station(reserves, row):
reserve_polygons = []
for r in reserves['features']:
name, poly = r['properties']['name'], sg.shape(r['geometry'])
reserve_polygons.append({'id': name, 'geometry': poly})
sid = False
for r in reserve_polygons:
if r['geometry'].contains(row['point']):
sid = r['id']
return sid
station_df['station'] = station_df.apply(lambda row: in_station(reserves, row),axis=1)
in_stations_df = station_df[station_df['station'] != False]
in_stations_df.head()
# + [markdown] #student=false tags=["ignore"]
# Let's see if this corresponds to what we observed on the map:
# + #student=false tags=["ignore"]
in_stations_df.groupby(['species', 'station'])['station'].count().unstack().plot.barh(stacked=True);
# + [markdown] #student=false tags=["ignore"]
# ### **Concept check:** *Why might you want to identify what research stations have specimen records? What kinds of scientific questions could you answer with this information?*
# + [markdown] #student=false tags=["ignore"]
# # Part 4: Comparing California Oak Species <a id='oak'></a>
# + [markdown] #student=false tags=["ignore"]
# Instead of investigating just one species, let’s compare several different species. Mapping the distributions of several different species can illuminate patterns of interactions and ecosystem functions.
#
# California oaks are common woody plants across North America. Almost all oaks are trees, but in drier areas they can be found as shrub oaks in poorer soils. Thus, they can be a great model system to illuminate the processes of speciation, adaptation, and expression. Let’s see their distribution!
# + [markdown] #student=false tags=["ignore"]
# <table style='center'>
# <tr>
# <td style="text-align: center; vertical-align: middle;">
# <img src="https://upload.wikimedia.org/wikipedia/commons/thumb/6/6d/Large_Blue_Oak.jpg/220px-Large_Blue_Oak.jpg" alt="<NAME>" />
# <br />
# <NAME>
# </td>
# <td style="text-align: center; vertical-align: middle;">
# <img src="https://upload.wikimedia.org/wikipedia/commons/thumb/8/86/Valley_Oak_Mount_Diablo.jpg/220px-Valley_Oak_Mount_Diablo.jpg" alt="<NAME>" />
# <br />
# <NAME>
# </td>
# </tr>
# <tr>
# <td style="text-align: center; vertical-align: middle;">
# <img src="https://upload.wikimedia.org/wikipedia/commons/thumb/b/b8/Quercusduratadurata.jpg/220px-Quercusduratadurata.jpg" alt="<NAME>" />
# <br />
# <NAME>
# </td>
# <td style="text-align: center; vertical-align: middle;">
# <img src="https://upload.wikimedia.org/wikipedia/commons/thumb/d/d1/Quercus_agrifolia_foliage.jpg/220px-Quercus_agrifolia_foliage.jpg" alt="Quercus agrifolia" />
# <br />
# Quercus agrifolia
# </td>
# </tr>
# </table>
# + [markdown] #student=false tags=["ignore"]
# Let's get the California oak records. This cell will take a while to run (about 30 seconds).
# + #student=false tags=["ignore"]
species_dfs = []
species = ['Quercus douglassi', 'Quercus lobata', 'Quercus durata', 'Quercus agrifolia']
# Here, we're getting the species record from the API for the four species of oak trees listed above
for s in species:
species_dfs.append(get_species_records_df(s))
# Combine the data we received into one DataFrame
oak_df = pd.concat(species_dfs, axis=0, sort=True)
# + #student=false tags=["ignore"]
oak_df.head() # Show the first 5 rows of our data
# + [markdown] #student=false tags=["ignore"]
# The table above only shows us the first 5 rows. Run the cell below to see how many total records we have.
# + #student=false tags=["ignore"]
len(oak_df)
# + [markdown] #student=false tags=["ignore"]
# Let's see how those records are distributed by species and sub-species.
# + #student=false tags=["ignore"]
# This creates a bar graph showing the distribution of species in our records
oak_df['scientificName'].value_counts().plot.barh();
# + [markdown] #student=false tags=["ignore"]
# We can also map these like we did with the *Argia arioides* above:
# + #student=false tags=["ignore"]
color_dict, html_key = assign_colors(oak_df, 'scientificName')
display(HTML(html_key))
# + #student=false tags=["ignore"]
oak_map = map_species_with_folium(oak_df, 'scientificName', child=folium.features.GeoJson(reserves))
oak_map
# + [markdown] #student=false tags=["include"]
#
# <!-- BEGIN QUESTION -->
#
# <div class="alert alert-block alert-warning">
# <b>QUESTION 3:</b>
# <br />
# Examine the map you generated of <i>Quercus spp</i>. In some places the geographic range of each species overlaps and in other parts of California the range is non-overlapping. Discuss factors that create this patterning in the oak community. Include concepts of niche and competitive exclusion.
# </div>
# + [markdown] #student=true tags=["include"]
#
# YOUR RESPONSE HERE
#
# <!-- END QUESTION -->
#
# + [markdown] #student=false tags=["ignore"]
# # Part 5: Cal-Adapt <a id='caladapt'></a>
# + [markdown] #student=false tags=["ignore"]
# Now that we've demonstrated what can be learned from mapping, let's take this one step futher. Using another resource [Cal-Adapt](http://www.cal-adapt.org/), we will pull past data and future projections to make some inferences about the future distribution of species impacted by climate change.
#
# Let's go back to the data from _Argia agrioides_ with the GBIF API. This will take a few seconds. The output is also really long. Remember you can click the area to the left of the cell (below the red `Out[ ]`) to expand/collapse the output.
# + #student=false tags=["ignore"]
# Get the first five records in raw text form (rather than DataFrame)
argia_agrioides_records = get_species_records('Argia agrioides')
argia_agrioides_records[:5] # Show the first 5 records
# This looks different from the records from our earlier records because these are the raw records
# + [markdown] #student=false tags=["ignore"]
# Now we will use the [Cal-Adapt](http://www.cal-adapt.org/) Web API to work with time series raster data. It will request an entire time series for any geometry and return a DataFrame for each record in all of our _Argia agrioides_ records. This cell also takes a while to run (1-3 minutes).
# + #student=false tags=["ignore"]
req = CalAdaptRequest()
record_geometry = [dict(rec, geometry=sg.Point(rec['decimalLongitude'], rec['decimalLatitude']))
for rec in argia_agrioides_records]
ca_df = req.concat_features(record_geometry, 'gbifID')
ca_df.head() # Show the first five rows
# + [markdown] #student=false tags=["include"]
#
# <!-- BEGIN QUESTION -->
#
# <div class="alert alert-block alert-warning">
# <b>QUESTION 4:</b>
# <br />
# What is Cal-Adapt? What can it be used for?
# </div>
# + [markdown] #student=true tags=["include"]
#
# YOUR RESPONSE HERE
#
# <!-- END QUESTION -->
#
# + [markdown] #student=false tags=["ignore"]
# <!-- END QUESTION -->
# This looks like the time series data we want for each record (the unique ID numbers as the columns). Each record has the projected temperature in Fahrenheit for 170 years (every row!). We can plot predictions for few random records and include a temperature threshold:
# + #student=false tags=["ignore"]
# Make a line plot using the first 9 columns of df.
ca_df.iloc[:,:9].plot(figsize=(18,10));
#set threshold at 90.
plt.axhline(y=90, xmin=0, xmax=1)
# Use matplotlib to title your plot.
plt.title('Argia agrioides - %s' % req.slug)
# Use matplotlib to add labels to the x and y axes of your plot.
plt.xlabel('Year', fontsize=18)
plt.ylabel('Degrees (Fahrenheit)', fontsize=16);
# + [markdown] #student=false tags=["ignore"]
# It looks like temperature is increasing across the board wherever these observations are occuring. We can calculate the average temperature for each year across observations in California:
# + #student=false tags=["ignore"]
tmax_means = ca_df.mean(axis=1)
# This is just some Pandas code to make the data prettier
pd.DataFrame(tmax_means).reset_index().rename(columns={'event':'Year', 0:'Avg Projected Temp'})
# + #student=false tags=["ignore"]
# This plots the data from the previous table into a graph
tmax_means.plot();
# + [markdown] #student=false tags=["include"]
#
# <!-- BEGIN QUESTION -->
#
# <div class="alert alert-block alert-warning">
# <b>QUESTION 5:</b>
# <br />
# What's happening to the average temperature that <i>Argia agrioides</i> is going to experience in the coming years across California? Current research indicates that adult <i>Argia agrioides</i> cannot survive in air temperatures above 100 degrees Fahrenheit, while larvae cannot survive in water temperatures above 75 degrees F. Based on this information, what predictions can you make about the distribution of <i>Argia agrioides</i> in California in 50 years? In 100 years?
# </div>
# + [markdown] #student=true tags=["include"]
#
# YOUR RESPONSE HERE
#
# <!-- END QUESTION -->
# + [markdown] #student=false tags=["include"]
#
# <!-- BEGIN QUESTION -->
#
# <div class="alert alert-block alert-warning">
# <b>QUESTION 6:</b>
# <br />
# What does this tell you about Santa Cruz Island? As time goes on and the temperature increases, might Santa Cruz Island serve as a refuge for <i>Argia agrioides</i>? (<i>Hint: look into the current climate and environmental conditions of Santa Cruz Island</i>)
# </div>
# + [markdown] #student=true tags=["include"]
#
# YOUR RESPONSE HERE
#
# <!-- END QUESTION -->
# + [markdown] #student=false tags=["ignore"]
# **Make sure that you've answered questions 1-5. Also make sure you've done all 3 exercises.**
#
# You are finished with this notebook! Please run the following cell to generate a download link for your submission file to submit on bCourses.
#
# **Note:** If clicking the link above doesn't work for you, don't worry! Simply right click and choose `Save Link As...` to save a copy of your pdf onto your computer.
#
# **Note:** If you made any changes to your responses and/or they are not reflected in your PDF, make sure to **re-run all the cells.** You can do this by going to the top and clicking `Cell >> Run All` and download your new PDF.
#
#
# **Check the PDF before submitting and make sure all of your answers & code changes are shown.**
# + tags=["ignore"]
#This may take a few extra seconds.
from otter.export import export_notebook
from IPython.display import display, HTML
export_notebook("notebook2.ipynb", filtering=True, pagebreaks=False)
display(HTML("Save this notebook, then click <a href='notebook2.pdf' download>here</a> to open the pdf."))
# + [markdown] #student=false tags=["ignore"]
# You are finished with this notebook! Please run the above cell to generate a download link for your submission file.
#
# If the download link does not work, open a new tab and go to https://datahub.berkeley.edu, click the box next to `notebook2_submission.pdf`, then click the "Download" link below the menu bar.
# + [markdown] #student=false tags=["ignore"]
# **Check the PDF before submitting and make sure all of your answers & code changes are shown.**
# + [markdown] #student=false tags=["ignore"]
# ---
#
# Notebook developed by: <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>
#
# [Data Science Modules](http://data.berkeley.edu/education/modules)
| notebook2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: proj_env
# language: python
# name: proj_env
# ---
from fastquant import get_stock_data, backtest
from gaussian_hmm import *
import matplotlib.pyplot as plt
# +
params = {
'n_components': 2,
'algorithm': 'map',
'n_iter': 100,
'd': 5,
'name':'GHMM'
}
ghmm = GHMM(params=params)
# -
# back testing dates on AAPL 2019-01-01 to 2020-05-31
back_test_data = get_stock_data('AAPL', '2019-01-01', '2020-05-31')
# training data AAPL 2017-08-01 to 2019-01-01
training_data = get_stock_data('AAPL', '2017-08-01', '2019-01-01')
ghmm.train(train_data=training_data)
preds,actual = ghmm.predict(test_data=back_test_data)
| frac_change_forecasting/hmms/.ipynb_checkpoints/backtesting-checkpoint.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.1
# language: julia
# name: julia-1.6
# ---
# # User defined equations of state
# In this notebook, we will give an example of how to implement a new equation of state within `Clapeyron.jl`. In this case, it will be the SAFT-VR Mie+AT model developed by Walker _et al._ which is simply a modification of the regular SAFT-VR Mie equation of state.
#
# The first step is to call `Clapeyron.jl` as one normally would only this time importing a few additional functions which we would like to modify:
using Pkg
Pkg.activate("..")
using Clapeyron, PyCall
import PyPlot; const plt = PyPlot
import Clapeyron: SAFTVRMieModel, @f, @newmodel, AssocOptions, has_sites, has_groups, N_A
import Clapeyron: a_res, a_disp, a_hs, d, B, aS_1, KHS, f123456, Cλ, ∑, data
# The next step is to define the parameter object for our equation of state. This is all the parameters we will need within the equation. The parameter type will also need to be defined (`SingleParam` for those parameters which only depend on a single species and `PairParam` for those which depend on two species):
struct SAFTVRMieATParam <: EoSParam
segment::SingleParam{Float64}
sigma::PairParam{Float64}
lambda_a::PairParam{Float64}
lambda_r::PairParam{Float64}
epsilon::PairParam{Float64}
Mw::SingleParam{Float64}
nu::SingleParam{Float64}
end
# The next step is to define the model type. Since SAFT-VR Mie+AT re-uses many of the same functions as SAFT-VR Mie, we define the `SAFTVRMieATModel` as a sub-type of `SAFTVRMieModel`. This effectively means we only need to re-define those functions which are different between the two equations:
abstract type SAFTVRMieATModel <: SAFTVRMieModel end
# We then create the model using the `@newmodel` macro:
@newmodel SAFTVRMieAT SAFTVRMieATModel SAFTVRMieATParam
export SAFTVRMieAT
# The next function that needs to be built is the model constructor (here called `SAFTVRMieAT`). This involves a few different components:
# Although optional, there should be some additional argument if the user would like to modify certain aspects of the model, such as the ideal model used
function SAFTVRMieAT(components;
idealmodel=BasicIdeal,
userlocations=String[],
ideal_userlocations=String[],
verbose=false)
# The next step is to collect all the parameters from the csv files. The getparams function can handle this.
# It is also useful to allow for custom userlocations to be fed into this function
params = Clapeyron.getparams(components, ["properties/molarmass.csv"]; userlocations=append!(userlocations,ideal_userlocations), verbose=verbose)
# The next step is to transform the parameters from the database into the form in which they are useable within the equation of state (e.g. applying combining rules)
params["Mw"].values .*= 1E-3
Mw = params["Mw"]
segment = params["m"]
nu = params["nu"]
params["sigma"].values .*= 1E-10
sigma = Clapeyron.sigma_LorentzBerthelot(params["sigma"])
epsilon = Clapeyron.epsilon_HudsenMcCoubrey(params["epsilon"], sigma)
lambda_a = Clapeyron.lambda_LorentzBerthelot(params["lambda_a"])
lambda_r = Clapeyron.lambda_LorentzBerthelot(params["lambda_r"])
# The struct with all of our parameters is then formed
packagedparams = SAFTVRMieATParam(segment, sigma, lambda_a, lambda_r, epsilon, Mw, nu)
# If the model is based on an existing one, it's generally a good idea to add the reference DOI here
references = [""]
# We then construct and return the model
model = SAFTVRMieAT(packagedparams, idealmodel;ideal_userlocations=ideal_userlocations, references=references, verbose=verbose)
return model
end
# The next step involves writing down the equations used within the equation of state. At the end of the day, the output should be `a_res`, the reduced residual Helmholtz free energy:
# +
function d(model::SAFTVRMieATModel, V, T, z)
ρ = N_A*sum(z)/V
λ = 0.85
ν = model.params.nu.values
ϵ = model.params.epsilon.diagvalues
σ = model.params.sigma.diagvalues
λr = model.params.lambda_r.diagvalues
λa = model.params.lambda_a.diagvalues
return d.(model,V,T,Ref(z),λa,λr,ϵ,σ,ν,ρ,λ)
end
function d(model::SAFTVRMieATModel, V, T, z, λa,λr,ϵ,σ,ν,ρ,λ)
u = Clapeyron.SAFTVRMieconsts.u
w = Clapeyron.SAFTVRMieconsts.w
θ = @f(Clapeyron.Cλ,λa,λr)*ϵ/T*(1-ρ*λ*ν*σ^3)
σ*(1-∑(w[j]*(θ/(θ+u[j]))^(1/λr)*(exp(θ*(1/(θ/(θ+u[j]))^(λa/λr)-1))/(u[j]+θ)/λr) for j ∈ 1:5))
end
function a_disp(model::SAFTVRMieATModel, V, T, z,_data = @f(data))
_d,ρS,ζi,_ζ_X,_ζst,_ = _data
comps = @Clapeyron.comps
l = length(comps)
∑z = ∑(z)
m = model.params.segment.values
_ϵ = model.params.epsilon.values
_λr = model.params.lambda_r.values
_λa = model.params.lambda_a.values
_σ = model.params.sigma.values
_ν = model.params.nu.values
m̄ = Clapeyron.dot(z, m)
m̄inv = 1/m̄
a₁ = zero(V+T+first(z))
a₂ = a₁
a₃ = a₁
_ζst5 = _ζst^5
_ζst8 = _ζst^8
_KHS = @f(KHS,_ζ_X,ρS)
ρ = N_A*sum(z)/V
λ = 0.85
for i ∈ comps
j = i
x_Si = z[i]*m[i]*m̄inv
x_Sj = x_Si
ϵ = _ϵ[i,j]
λa = _λa[i,i]
λr = _λr[i,i]
σ = _σ[i,i]
ν = _ν[i]
AT_corr = (1-ρ*λ*ν*σ^3)
_C = @f(Cλ,λa,λr)
dij = _d[i]
dij3 = dij^3
x_0ij = σ/dij
#calculations for a1 - diagonal
aS_1_a = @f(aS_1,λa,_ζ_X)
aS_1_r = @f(aS_1,λr,_ζ_X)
B_a = @f(B,λa,x_0ij,_ζ_X)
B_r = @f(B,λr,x_0ij,_ζ_X)
a1_ij = (2*π*AT_corr*ϵ*dij3)*_C*ρS*
(x_0ij^λa*(aS_1_a+B_a) - x_0ij^λr*(aS_1_r+B_r))
#calculations for a2 - diagonal
aS_1_2a = @f(aS_1,2*λa,_ζ_X)
aS_1_2r = @f(aS_1,2*λr,_ζ_X)
aS_1_ar = @f(aS_1,λa+λr,_ζ_X)
B_2a = @f(B,2*λa,x_0ij,_ζ_X)
B_2r = @f(B,2*λr,x_0ij,_ζ_X)
B_ar = @f(B,λr+λa,x_0ij,_ζ_X)
α = _C*(1/(λa-3)-1/(λr-3))
f1,f2,f3,f4,f5,f6 = @f(f123456,α)
_χ = f1*_ζst+f2*_ζst5+f3*_ζst8
a2_ij = π*_KHS*(1+_χ)*ρS*AT_corr^2*ϵ^2*dij3*_C^2 *
(x_0ij^(2*λa)*(aS_1_2a+B_2a)
- 2*x_0ij^(λa+λr)*(aS_1_ar+B_ar)
+ x_0ij^(2*λr)*(aS_1_2r+B_2r))
#calculations for a3 - diagonal
a3_ij = -ϵ^3*AT_corr^3*f4*_ζst * exp(f5*_ζst+f6*_ζst^2)
#adding - diagonal
a₁ += a1_ij*x_Si*x_Si
a₂ += a2_ij*x_Si*x_Si
a₃ += a3_ij*x_Si*x_Si
end
a₁ = a₁*m̄/T/∑z
a₂ = a₂*m̄/(T*T)/∑z
a₃ = a₃*m̄/(T*T*T)/∑z
#@show (a₁,a₂,a₃)
adisp = a₁ + a₂ + a₃
return adisp
end
function a_res(model::SAFTVRMieATModel, V, T, z)
_data = @f(data)
return @f(a_hs,_data)+@f(a_disp,_data)
end
# -
# With that, the equation is built and ready to use! As an example:
# +
species = ["He_JHBV","Ne_JHBV","Ar_JHBV","Kr_JHBV","Xe_JHBV"]
model = SAFTVRMieAT.(species;userlocations=["Ab_initio_SAFT.csv"],
ideal_userlocations=["Ab_initio_SAFT_molarmass.csv"]);
# -
# We can then obtain critical and saturation properties, and plot the results:
# +
label=["Helium","Neon","Argon","Krypton","Xenon"]
color=["y","r","b","g","purple"]
crit=crit_pure.(model);
for i ∈ 1:5
T = range(0.7*crit[i][1],crit[i][1],length=100)
sat = saturation_pressure.(model[i],T)
p = [sat[i][1] for i ∈ 1:100]
v_l = [sat[i][2] for i ∈ 1:100]
v_v = [sat[i][3] for i ∈ 1:100]
plt.semilogy(T,p./1e6,color=color[i],label=label[i])
plt.plot(crit[i][1],crit[i][2]./1e6,marker="o",color=color[i])
end
plt.ylim([0.7,1e1])
plt.xlim([0,320])
plt.legend(loc="lower right",frameon=false,fontsize=12)
plt.xlabel("Temperature / K",fontsize=16)
plt.ylabel("Pressure / MPa",fontsize=16)
plt.xticks(fontsize=12)
plt.yticks(fontsize=12)
| examples/user_defined_eos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# LinearSearch
n = int(input())
li = [int(x) for x in input().split()]
li
ele = int(input())
isFound = False
for i in range(len(li)):
if li[i] == ele:
print(i)
isFound = True
break
if isFound is False:
print(-1)
ele = int(input())
isFound = False
for i in range(len(li)):
if li[i] == ele:
print(i)
isFound = True
break
if isFound is False:
print(-1)
| random/LinearSearch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 2 - Transient Conduction
#
# Solve the following problem and explain your results.
# ## Problem 1 - Transient Conduction in a Plane Wall
#
# The problem we consider is that of a plane wall initially at 100$^\circ$C with its outer surfaces exposed to an ambient temperature of 0$^\circ$C. The wall has a thickness 2*L* and may be considered to have an infinite height and a unit depth. The domain is shown schematically below:
#
# 
#
# Initially, the wall only feels the effect of the ambient air very near the surface and thus, the temperature profile inside the solid is quite steep in the vicinity of the surface. An analytical solution for this problem involves several (at least 4) terms of a Fourier series. After some time, however, the influence of the ambient air will have reached the center of the wall and the analytical solution can be approximated by the first term of the Fourier series (see, for example, *Fundamentals of Heat and Mass Transfer* by Incropera et al.). To study the order of accuracy of the fully implicit first and second order time discretization schemes, we will consider the cooling process during a period past the initial transient where the one-term Fourier solution is valid.
#
# The parameters for the problem are:
#
# $$ Bi = \frac{h L}{k}= 1.0 $$
#
# $$ T_i = 100^\circ C $$
#
# $$ T_{\infty}= 0^\circ C $$
#
# The one-term Fourier solution for this problem is:
#
# $$
# \frac{T-T_{\infty}}{T_i-T_{\infty}}=C_1 \exp\left(-\zeta^2\frac{\alpha t}{L^2}\right)\cos\left(\zeta \frac{x}{L}\right)
# $$
#
# where:
#
# $$ T = T(x,t) $$
#
# $$ \alpha = \frac{k}{\rho c_p} $$
#
# $$ C_1 = 1.1191 $$
#
# $$ \zeta = 0.8603 $$
#
# The solution to this problem at the two different dimensionless time levels of interest is:
#
# $$ \text{at } \frac{\alpha t_1}{L^2}= 0.4535,~~~ T(0,t_1)= 80^\circ C $$
#
# $$ \text{at } \frac{\alpha t_2}{L^2}= 3.2632,~~~ T(0,t_2)= 10^\circ C $$
#
# To solve this problem, initialize the temperature field using the analytical solution at $\alpha t_1/L^2 = 0.4535$. This avoids the need for a very small timestep during the initial transient when solution is changing rapidly. Then, use your code to calculate the temporal variation of the temperature field over the time period described above. Solve the problem by employing 2, 4, 8, 16, and 32 time steps using both the first and second order implicit schemes.
#
# At the end of each run, calculate the absolute average error, $\overline{e}$, using the formula:
#
# $$
# \overline{e}= \frac{1}{N_{CV}} \sum_{i=1}^{N_{CV}} |e(i)|
# $$
#
# where
#
# $$ e(i) = T_{exact}(i) - T(i) $$
#
# Then, for each scheme, plot your results of $\overline{e}$ vs. $\Delta t$ (on a log-log scale) and find the value of $p$ in the expression:
#
# $$
# \overline{e}= c (\Delta t)^p
# $$
#
# where $p$ represents the order accuracy of the transient scheme. Also show a separate plot of T(0,$t_2$) verses the number of timesteps used for each scheme employed.
#
# Repeat this problem on at least three different grids to demonstrate grid independence of the solution.
#
# **Bonus**: Solve the same problem using the Crank-Nicolson scheme and compare the results.
#
| cfdcourse/Assignments/Assignment2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# # AutoML 02: Regression with Local Compute
#
# In this example we use the scikit-learn's [diabetes dataset](http://scikit-learn.org/stable/datasets/index.html#diabetes-dataset) to showcase how you can use AutoML for a simple regression problem.
#
# Make sure you have executed the [00.configuration](00.configuration.ipynb) before running this notebook.
#
# In this notebook you will learn how to:
# 1. Create an `Experiment` in an existing `Workspace`.
# 2. Configure AutoML using `AutoMLConfig`.
# 3. Train the model using local compute.
# 4. Explore the results.
# 5. Test the best fitted model.
#
# ## Create an Experiment
#
# As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments.
# +
import logging
import os
import random
from matplotlib import pyplot as plt
from matplotlib.pyplot import imshow
import numpy as np
import pandas as pd
from sklearn import datasets
import azureml.core
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
from azureml.train.automl import AutoMLConfig
from azureml.train.automl.run import AutoMLRun
# +
ws = Workspace.from_config()
# Choose a name for the experiment and specify the project folder.
experiment_name = 'automl-local-regression'
project_folder = './sample_projects/automl-local-regression'
experiment = Experiment(ws, experiment_name)
output = {}
output['SDK version'] = azureml.core.VERSION
output['Subscription ID'] = ws.subscription_id
output['Workspace Name'] = ws.name
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Project Directory'] = project_folder
output['Experiment Name'] = experiment.name
pd.set_option('display.max_colwidth', -1)
pd.DataFrame(data = output, index = ['']).T
# -
# ### Load Training Data
# This uses scikit-learn's [load_diabetes](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_diabetes.html) method.
# +
# Load the diabetes dataset, a well-known built-in small dataset that comes with scikit-learn.
from sklearn.datasets import load_diabetes
from sklearn.linear_model import Ridge
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
X, y = load_diabetes(return_X_y = True)
columns = ['age', 'gender', 'bmi', 'bp', 's1', 's2', 's3', 's4', 's5', 's6']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
# -
# ## Configure AutoML
#
# Instantiate an `AutoMLConfig` object to specify the settings and data used to run the experiment.
#
# |Property|Description|
# |-|-|
# |**task**|classification or regression|
# |**primary_metric**|This is the metric that you want to optimize. Regression supports the following primary metrics: <br><i>spearman_correlation</i><br><i>normalized_root_mean_squared_error</i><br><i>r2_score</i><br><i>normalized_mean_absolute_error</i>|
# |**max_time_sec**|Time limit in seconds for each iteration.|
# |**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|
# |**n_cross_validations**|Number of cross validation splits.|
# |**X**|(sparse) array-like, shape = [n_samples, n_features]|
# |**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]<br>Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers.|
# |**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder.|
automl_config = AutoMLConfig(task = 'regression',
max_time_sec = 600,
iterations = 10,
primary_metric = 'spearman_correlation',
n_cross_validations = 5,
debug_log = 'automl.log',
verbosity = logging.INFO,
X = X_train,
y = y_train,
path = project_folder)
# ## Train the Models
#
# Call the `submit` method on the experiment object and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while.
# In this example, we specify `show_output = True` to print currently running iterations to the console.
local_run = experiment.submit(automl_config, show_output = True)
local_run
# ## Explore the Results
# #### Widget for Monitoring Runs
#
# The widget will first report a "loading" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.
#
# **Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details.
from azureml.train.widgets import RunDetails
RunDetails(local_run).show()
#
# #### Retrieve All Child Runs
# You can also use SDK methods to fetch all the child runs and see individual metrics that we log.
# +
children = list(local_run.get_children())
metricslist = {}
for run in children:
properties = run.get_properties()
metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}
metricslist[int(properties['iteration'])] = metrics
rundata = pd.DataFrame(metricslist).sort_index(1)
rundata
# -
# ### Retrieve the Best Model
#
# Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*.
best_run, fitted_model = local_run.get_output()
print(best_run)
print(fitted_model)
# #### Best Model Based on Any Other Metric
# Show the run and the model that has the smallest `root_mean_squared_error` value (which turned out to be the same as the one with largest `spearman_correlation` value):
lookup_metric = "root_mean_squared_error"
best_run, fitted_model = local_run.get_output(metric = lookup_metric)
print(best_run)
print(fitted_model)
# ### Test the Best Fitted Model
# Predict on training and test set, and calculate residual values.
# +
y_pred_train = fitted_model.predict(X_train)
y_residual_train = y_train - y_pred_train
y_pred_test = fitted_model.predict(X_test)
y_residual_test = y_test - y_pred_test
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from sklearn import datasets
from sklearn.metrics import mean_squared_error, r2_score
# Set up a multi-plot chart.
f, (a0, a1) = plt.subplots(1, 2, gridspec_kw = {'width_ratios':[1, 1], 'wspace':0, 'hspace': 0})
f.suptitle('Regression Residual Values', fontsize = 18)
f.set_figheight(6)
f.set_figwidth(16)
# Plot residual values of training set.
a0.axis([0, 360, -200, 200])
a0.plot(y_residual_train, 'bo', alpha = 0.5)
a0.plot([-10,360],[0,0], 'r-', lw = 3)
a0.text(16,170,'RMSE = {0:.2f}'.format(np.sqrt(mean_squared_error(y_train, y_pred_train))), fontsize = 12)
a0.text(16,140,'R2 score = {0:.2f}'.format(r2_score(y_train, y_pred_train)), fontsize = 12)
a0.set_xlabel('Training samples', fontsize = 12)
a0.set_ylabel('Residual Values', fontsize = 12)
# Plot a histogram.
a0.hist(y_residual_train, orientation = 'horizontal', color = 'b', bins = 10, histtype = 'step');
a0.hist(y_residual_train, orientation = 'horizontal', color = 'b', alpha = 0.2, bins = 10);
# Plot residual values of test set.
a1.axis([0, 90, -200, 200])
a1.plot(y_residual_test, 'bo', alpha = 0.5)
a1.plot([-10,360],[0,0], 'r-', lw = 3)
a1.text(5,170,'RMSE = {0:.2f}'.format(np.sqrt(mean_squared_error(y_test, y_pred_test))), fontsize = 12)
a1.text(5,140,'R2 score = {0:.2f}'.format(r2_score(y_test, y_pred_test)), fontsize = 12)
a1.set_xlabel('Test samples', fontsize = 12)
a1.set_yticklabels([])
# Plot a histogram.
a1.hist(y_residual_test, orientation = 'horizontal', color = 'b', bins = 10, histtype = 'step')
a1.hist(y_residual_test, orientation = 'horizontal', color = 'b', alpha = 0.2, bins = 10)
plt.show()
| Tutorials/MLADS-fall-2018/demo/automl-regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pdx-exomeseq-analysis]
# language: python
# name: conda-env-pdx-exomeseq-analysis-py
# ---
# # Visualizing Read Depth Statistics
#
# The depth and number of mapped reads are visualized in a series of plots. The data come from `mosdepth` and `samtools flagstat` commands.
# +
import os
import csv
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# -
# %matplotlib inline
plt.style.use('seaborn-notebook')
# +
# Load phenotype data to convert sample id
pheno_df = (
pd.read_csv('pdx_phenotype.csv', dtype='str')
.drop_duplicates(subset=['final_id', 'sample_name'])
)
id_updater = dict(zip([x[0] for x in pheno_df.read_id.str.split('_')],
pheno_df.final_id))
id_updater
# -
# ## Mosdepth calculations of Coverage and Depth
# +
depth_dir = os.path.join('results', 'wes_stats')
depth_files = os.listdir(depth_dir)
plt.xlabel('Depth')
plt.ylabel('Proportion of Genome')
full_depth_list = []
all_samples = []
for depth_file in depth_files:
# Load and process file
full_depth_file = os.path.join(depth_dir, depth_file)
# Update sample id from depth file with id_updated dictionary
sample_id = id_updater[depth_file.split('.')[0]]
depth_df = pd.read_table(full_depth_file, names=['chrom', 'depth', 'prop_cov'])
depth_df = depth_df.assign(sample_id = sample_id)
# Each line will be a different type if under a certain depth
half_depth = depth_df.query('chrom == "total"').query('prop_cov >= 0.5').depth.max()
# Ensure that the top half read depth estimations are solid lines
if half_depth > 78 or sample_id == '001-F5':
linestyle = 'solid'
else:
linestyle = 'dotted'
# Store order and half depth of samples (for plot labeling later)
all_samples.append([sample_id, half_depth])
# Set a reasonable cutoff to view bulk of distribution
depth_sub_df = depth_df[depth_df['depth'] < 300]
# There are chromosome specific estimates, here, view total
ax = plt.plot('depth',
'prop_cov',
linestyle=linestyle,
data=depth_sub_df.query('chrom == "total"'),
label=sample_id)
# Save processed files to list for later extraction
full_depth_list.append(depth_df)
# Get the default handles and labels
handles, labels = plt.gca().get_legend_handles_labels()
# Sort the labels by their depth at 50% of the genome
sorted_labels = (
pd.DataFrame(all_samples,
columns=['sample_id', 'half_depth'])
.sort_values(by='half_depth', ascending=False)
.sample_id
.tolist()
)
# Sort the handles (matplotlib legend lines)
sorted_handles = [dict(zip(labels, handles))[x] for x in sorted_labels]
# Show legend in decreasing depth order
lgd = plt.legend(sorted_handles,
sorted_labels,
ncol=2,
bbox_to_anchor=(1.03, 1),
loc=2,
borderaxespad=0.,
fontsize=10)
# Save Figure
depth_fig_file = os.path.join('figures', 'mosdepth_estimation.pdf')
plt.savefig(depth_fig_file, bbox_extra_artists=(lgd,),
bbox_inches='tight')
# +
# Get full depth matrix
full_depth_df = pd.concat(full_depth_list)
# 50% of the exome is covered at what depth?
half_depth_df = full_depth_df[full_depth_df['prop_cov'] == 0.5]
half_depth_df = half_depth_df[half_depth_df['chrom'] == 'total']
# -
half_depth_df.describe()
# 75% of the exome is covered at what depth?
threeq_depth_df = full_depth_df[full_depth_df['prop_cov'] == 0.75]
threeq_depth_df = threeq_depth_df[threeq_depth_df['chrom'] == 'total']
threeq_depth_df.describe()
# ## General flagstat read mapping distributions
read_stat_dir = os.path.join('results', 'read_counts')
read_stat_files = os.listdir(read_stat_dir)
read_stat_file = os.path.join(read_stat_dir, read_stat_files[0])
# +
total_read_counts = []
mapped_read_counts = []
for read_stat_file in read_stat_files:
read_stat_file = os.path.join(read_stat_dir, read_stat_file)
with open(read_stat_file, 'r') as csvfile:
readstat_reader = csv.reader(csvfile, delimiter=' ')
line_idx = 0
for row in readstat_reader:
if line_idx == 0:
total_read_counts.append(row[0])
if line_idx == 2:
mapped_read_counts.append(row[0])
line_idx +=1
# -
# Get flagstat dataframe
read_depth_df = pd.DataFrame([total_read_counts, mapped_read_counts],
index=['total', 'mapped'], dtype='float64').T
read_depth_df.plot(x='total', y='mapped', kind='scatter');
# Output distribution of total and mapped reads
read_depth_df.describe()
| 1.read-depth-stats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#default_exp core.rebuild
# -
# # core.rebuild
# > Functions needed to rebuild vision-based transforms from export
#export
import io, operator, pickle, torch
#export
from pathlib import Path
from numpy import ndarray
from fastcore.utils import _patched
#export
from fastinference_pytorch.utils import to_device, tensor
# ## Unpickling export
#export
def load_data(path:Path=Path('.'), fn='data'):
"Opens `pkl` file containing exported `Transform` information"
if '.pkl' not in fn: fn += '.pkl'
if not isinstance(path, Path): path = Path(path)
with open(path/fn, 'rb') as handle:
tfmd_dict = pickle.load(handle)
return tfmd_dict
#export
def load_model(path:Path='.', fn='model', cpu=True, onnx=False):
if not onnx:
if '.pkl' not in fn: fn += '.pkl'
path = f'{path}/{fn}'
return torch.load(path, map_location='cpu' if cpu else None)
else:
try:
import onnxruntime as ort
except ImportError:
print('to use ONNX you must have `onnxruntime` installed\n\
Install it with `pip install onnxruntime-gpu`')
if '.onnx' not in fn: fn += '.onnx'
path = f'{path}/{fn}'
ort_session = ort.InferenceSession(path)
if not cpu:
try:
ort_session.set_providers(['CUDAExecutionProvider'])
except:
ort_session.set_providers(['CPUExecutionProvider'])
return ort_session
# ## Pipelines
#
# These generate our transform "pipelines" (as we're not using `fastcore`'s `Pipeline`) to pass our data through
#export
def get_tfm(key, tfms):
"Makes a transform from `key`. Class or function must be in global memory (imported)"
args = tfms[key]
return globals()[key](**args)
#export
def generate_pipeline(tfms, order=True) -> dict:
"Generate `pipe` of transforms from dict and (potentially) sort them"
pipe = []
for key in tfms.keys():
tfm = get_tfm(key, tfms)
pipe.append(tfm)
if order: pipe = sorted(pipe, key=operator.attrgetter('order'))
return pipe
#export
def make_pipelines(tfms) -> dict:
"Make `item` and `batch` transform pipelines"
pipe, keys = {}, ['after_item', 'after_batch']
for key in keys:
pipe[key] = generate_pipeline(tfms[key], True)
if not any(isinstance(x, ToTensor) for x in pipe['after_item']):
pipe['after_item'].append(ToTensor(False))
return pipe
| 00d_core.rebuild.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # `layers.backbone.densenet`
# +
# # %load ../../HPA-competition-solutions/bestfitting/src/layers/backbone/densenet.py
# +
#default_exp layers.backbone.densenet
# +
#export
import re
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.utils.model_zoo as model_zoo
from collections import OrderedDict
# +
#export
__all__ = ['DenseNet', 'densenet121', 'densenet169', 'densenet201', 'densenet161']
model_urls = {
'densenet121': 'https://download.pytorch.org/models/densenet121-a639ec97.pth',
'densenet169': 'https://download.pytorch.org/models/densenet169-b2777c0a.pth',
'densenet201': 'https://download.pytorch.org/models/densenet201-c1103571.pth',
'densenet161': 'https://download.pytorch.org/models/densenet161-8d451a50.pth',
}
def densenet121(pretrained=False, **kwargs):
r"""Densenet-121 model from
`"Densely Connected Convolutional Networks" <https://arxiv.org/pdf/1608.06993.pdf>`_
Args:
pretrained (bool): If True, returns a model pre-trained on ImageNet
"""
model = DenseNet(num_init_features=64, growth_rate=32, block_config=(6, 12, 24, 16),
**kwargs)
if pretrained:
# '.'s are no longer allowed in module names, but pervious _DenseLayer
# has keys 'norm.1', 'relu.1', 'conv.1', 'norm.2', 'relu.2', 'conv.2'.
# They are also in the checkpoints in model_urls. This pattern is used
# to find such keys.
pattern = re.compile(
r'^(.*denselayer\d+\.(?:norm|relu|conv))\.((?:[12])\.(?:weight|bias|running_mean|running_var))$')
state_dict = model_zoo.load_url(model_urls['densenet121'])
for key in list(state_dict.keys()):
res = pattern.match(key)
if res:
new_key = res.group(1) + res.group(2)
state_dict[new_key] = state_dict[key]
del state_dict[key]
model.load_state_dict(state_dict)
return model
def densenet169(pretrained=False, **kwargs):
r"""Densenet-169 model from
`"Densely Connected Convolutional Networks" <https://arxiv.org/pdf/1608.06993.pdf>`_
Args:
pretrained (bool): If True, returns a model pre-trained on ImageNet
"""
model = DenseNet(num_init_features=64, growth_rate=32, block_config=(6, 12, 32, 32),
**kwargs)
if pretrained:
# '.'s are no longer allowed in module names, but pervious _DenseLayer
# has keys 'norm.1', 'relu.1', 'conv.1', 'norm.2', 'relu.2', 'conv.2'.
# They are also in the checkpoints in model_urls. This pattern is used
# to find such keys.
pattern = re.compile(
r'^(.*denselayer\d+\.(?:norm|relu|conv))\.((?:[12])\.(?:weight|bias|running_mean|running_var))$')
state_dict = model_zoo.load_url(model_urls['densenet169'])
for key in list(state_dict.keys()):
res = pattern.match(key)
if res:
new_key = res.group(1) + res.group(2)
state_dict[new_key] = state_dict[key]
del state_dict[key]
model.load_state_dict(state_dict)
return model
def densenet201(pretrained=False, **kwargs):
r"""Densenet-201 model from
`"Densely Connected Convolutional Networks" <https://arxiv.org/pdf/1608.06993.pdf>`_
Args:
pretrained (bool): If True, returns a model pre-trained on ImageNet
"""
model = DenseNet(num_init_features=64, growth_rate=32, block_config=(6, 12, 48, 32),
**kwargs)
if pretrained:
# '.'s are no longer allowed in module names, but pervious _DenseLayer
# has keys 'norm.1', 'relu.1', 'conv.1', 'norm.2', 'relu.2', 'conv.2'.
# They are also in the checkpoints in model_urls. This pattern is used
# to find such keys.
pattern = re.compile(
r'^(.*denselayer\d+\.(?:norm|relu|conv))\.((?:[12])\.(?:weight|bias|running_mean|running_var))$')
state_dict = model_zoo.load_url(model_urls['densenet201'])
for key in list(state_dict.keys()):
res = pattern.match(key)
if res:
new_key = res.group(1) + res.group(2)
state_dict[new_key] = state_dict[key]
del state_dict[key]
model.load_state_dict(state_dict)
return model
def densenet161(pretrained=False, **kwargs):
r"""Densenet-161 model from
`"Densely Connected Convolutional Networks" <https://arxiv.org/pdf/1608.06993.pdf>`_
Args:
pretrained (bool): If True, returns a model pre-trained on ImageNet
"""
model = DenseNet(num_init_features=96, growth_rate=48, block_config=(6, 12, 36, 24),
**kwargs)
if pretrained:
# '.'s are no longer allowed in module names, but pervious _DenseLayer
# has keys 'norm.1', 'relu.1', 'conv.1', 'norm.2', 'relu.2', 'conv.2'.
# They are also in the checkpoints in model_urls. This pattern is used
# to find such keys.
pattern = re.compile(
r'^(.*denselayer\d+\.(?:norm|relu|conv))\.((?:[12])\.(?:weight|bias|running_mean|running_var))$')
state_dict = model_zoo.load_url(model_urls['densenet161'])
for key in list(state_dict.keys()):
res = pattern.match(key)
if res:
new_key = res.group(1) + res.group(2)
state_dict[new_key] = state_dict[key]
del state_dict[key]
model.load_state_dict(state_dict)
return model
class _DenseLayer(nn.Sequential):
def __init__(self, num_input_features, growth_rate, bn_size, drop_rate):
super(_DenseLayer, self).__init__()
self.add_module('norm1', nn.BatchNorm2d(num_input_features)),
self.add_module('relu1', nn.ReLU(inplace=True)),
self.add_module('conv1', nn.Conv2d(num_input_features, bn_size *
growth_rate, kernel_size=1, stride=1, bias=False)),
self.add_module('norm2', nn.BatchNorm2d(bn_size * growth_rate)),
self.add_module('relu2', nn.ReLU(inplace=True)),
self.add_module('conv2', nn.Conv2d(bn_size * growth_rate, growth_rate,
kernel_size=3, stride=1, padding=1, bias=False)),
self.drop_rate = drop_rate
def forward(self, x):
new_features = super(_DenseLayer, self).forward(x)
if self.drop_rate > 0:
new_features = F.dropout(new_features, p=self.drop_rate, training=self.training)
return torch.cat([x, new_features], 1)
class _DenseBlock(nn.Sequential):
def __init__(self, num_layers, num_input_features, bn_size, growth_rate, drop_rate):
super(_DenseBlock, self).__init__()
for i in range(num_layers):
layer = _DenseLayer(num_input_features + i * growth_rate, growth_rate, bn_size, drop_rate)
self.add_module('denselayer%d' % (i + 1), layer)
class _Transition(nn.Sequential):
def __init__(self, num_input_features, num_output_features):
super(_Transition, self).__init__()
self.add_module('norm', nn.BatchNorm2d(num_input_features))
self.add_module('relu', nn.ReLU(inplace=True))
self.add_module('conv', nn.Conv2d(num_input_features, num_output_features,
kernel_size=1, stride=1, bias=False))
self.add_module('pool', nn.AvgPool2d(kernel_size=2, stride=2))
class DenseNet(nn.Module):
r"""Densenet-BC model class, based on
`"Densely Connected Convolutional Networks" <https://arxiv.org/pdf/1608.06993.pdf>`_
Args:
growth_rate (int) - how many filters to add each layer (`k` in paper)
block_config (list of 4 ints) - how many layers in each pooling block
num_init_features (int) - the number of filters to learn in the first convolution layer
bn_size (int) - multiplicative factor for number of bottle neck layers
(i.e. bn_size * k features in the bottleneck layer)
drop_rate (float) - dropout rate after each dense layer
num_classes (int) - number of classification classes
"""
def __init__(self, growth_rate=32, block_config=(6, 12, 24, 16),
num_init_features=64, bn_size=4, drop_rate=0, num_classes=1000):
super(DenseNet, self).__init__()
# First convolution
self.features = nn.Sequential(OrderedDict([
('conv0', nn.Conv2d(3, num_init_features, kernel_size=7, stride=2, padding=3, bias=False)),
('norm0', nn.BatchNorm2d(num_init_features)),
('relu0', nn.ReLU(inplace=True)),
('pool0', nn.MaxPool2d(kernel_size=3, stride=2, padding=1)),
]))
# Each denseblock
num_features = num_init_features
for i, num_layers in enumerate(block_config):
block = _DenseBlock(num_layers=num_layers, num_input_features=num_features,
bn_size=bn_size, growth_rate=growth_rate, drop_rate=drop_rate)
self.features.add_module('denseblock%d' % (i + 1), block)
num_features = num_features + num_layers * growth_rate
if i != len(block_config) - 1:
trans = _Transition(num_input_features=num_features, num_output_features=num_features // 2)
self.features.add_module('transition%d' % (i + 1), trans)
num_features = num_features // 2
# Final batch norm
self.features.add_module('norm5', nn.BatchNorm2d(num_features))
# Linear layer
self.classifier = nn.Linear(num_features, num_classes)
# Official init from torch repo.
for m in self.modules():
if isinstance(m, nn.Conv2d):
nn.init.kaiming_normal(m.weight.data)
elif isinstance(m, nn.BatchNorm2d):
m.weight.data.fill_(1)
m.bias.data.zero_()
elif isinstance(m, nn.Linear):
m.bias.data.zero_()
def forward(self, x):
features = self.features(x)
out = F.relu(features, inplace=True)
out = F.adaptive_avg_pool2d(out,1).view(features.size(0), -1)
out = self.classifier(out)
return out
if __name__ == "__main__":
net=densenet121(pretrained=False)
print(net)
input = torch.randn(2, 3, 128, 128)
output = net.forward(input)
print(output.shape)
# -
| nbs/05_layers.backbone.densenet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# librerias necesarias
import numpy as np
import matplotlib.pyplot as plt
import math
import cv2 # OpenCV library for computer vision
from pillow import Image
import time
from keras.models import load_model
# funciones auxiliares
def cargaImagen(ruta):
""" Metodo que carga una imagen de una ruta """
image = cv2.imread(ruta)
image = cv2.cvtColor(imagen, cv2.COLOR_BGR2RGB)
return imagen
def dibujarImagen(imagen, title=''):
""" Funcion que muestra la Imagen """
# Plot our image using subplots to specify a size and title
fig = plt.figure(figsize = (8,8))
ax1 = fig.add_subplot(111)
ax1.set_xticks([])
ax1.set_yticks([])
ax1.set_title(title)
ax1.imshow(imagen)
# -
| Untitled1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="J_MIyKGnM3xE"
# Precisamos instalar primeiro o gym. Executar o seguinte em um notebook Jupyter deve funcionar:
# + colab={"base_uri": "https://localhost:8080/"} id="IjXylIsIMyt9" outputId="220e27fd-cbf1-462e-af57-38656487c51b"
# !pip install gym
# + [markdown] id="LOUggRubLaBr"
# ## Q-Learning com Gym
# Depois de instalado, podemos carregar o ambiente e renderizar sua aparência:
# Abrindo e mostrando um ambiente pronto do gym que simula o problema do Taxi (versão 3):
#
# + id="OOIZcsERLZSP" colab={"base_uri": "https://localhost:8080/"} outputId="ba984837-bdfe-4d4f-90d3-2f137647af18"
import gym #importa o gym
env = gym.make("Taxi-v3").env #carrega o ambiente
env.render() #Renderiza uma estrutura do ambiente (útil na visualização do ambiente)
# + [markdown] id="-tdwiHPjOQtS"
# O quadrado preenchido representa o táxi, que é amarelo sem passageiro e verde com passageiro.
# A barra vertical ("|") representa uma parede que o táxi não pode atravessar.
# R, G, Y, B são os locais de coleta e destino possíveis. A letra azul representa o local atual de embarque do passageiro e a letra roxa é o destino atual.
# + id="IhK_phsuXxox" colab={"base_uri": "https://localhost:8080/"} outputId="e366278e-89bb-4145-ce6d-11f06e4717aa"
print("Total de Ações {}".format(env.action_space))
print("Total de Estados {}".format(env.observation_space))
# + [markdown] id="GNDUC5XNa1AK"
# ### Alterando o estado
# Podemos codificar o estado do agente e fornecê-lo ao ambiente para renderizar no Gym. Lembre-se de que temos o táxi na linha 3, coluna 1, nosso passageiro está no local 2 e nosso destino é o local 0. Usando o método de codificação de estado Taxi-v3, podemos fazer o seguinte:
#
# Use a função env.encode(taxi_linha,taxi_coluna,passageiro_saida,passageiro_chegada)
# + id="hdAlu52DXyDw" colab={"base_uri": "https://localhost:8080/"} outputId="166775ef-ea4e-4a84-c7b1-f8fe68472942"
state = env.encode(3, 1, 2, 0)
print("Número do estado:", state)
env.s = state
env.render()
# + [markdown] id="pO4_XZWhPSeZ"
# Outra forma de definir o estado. Podemos definir o estado do ambiente manualmente env.env.s usando esse número codificado.
#
# 369 correspondente a um estado entre 0 e 499. O ambiente possue 500 estados possíveis
#
# + colab={"base_uri": "https://localhost:8080/"} id="FyTvFT55QznG" outputId="e63f5f22-e287-4d86-cadf-54147dec2309"
env.s = 369
print("Número do estado:", env.s)
env.render()
# + [markdown] id="IpV02LMuc5CT"
# **A Tabela de Recompensas**
# Quando o ambiente Taxi é criado, também é criada uma tabela inicial de Recompensas, chamada `P`. Podemos pensar nisso como uma matriz que tem o número de estados como linhas e o número de ações como colunas, ou seja,
# s t a t e s X a c t i o n s
#
# + id="akeOkzw2Xy1x" colab={"base_uri": "https://localhost:8080/"} outputId="94a3ef00-acc0-4f35-f6db-2b7216e32d32"
env.P[329] #estado atual
# + [markdown] id="LE-PCWT5Q0Ik"
# Este dicionário tem a estrutura {action: [(probability, nextstate, reward, done)]}.
#
# O 0-5 corresponde às ações (sul, norte, leste, oeste, embarque, desembarque) que o táxi pode executar em nosso estado atual na ilustração.
# Neste env, probabilidade é sempre 1.0.
# Este next state é o estado em que estaríamos se tomarmos a ação.
# Todas as ações de movimento têm uma recompensa -1 e as ações pickup (pegar)/dropoff(deixar) têm -10 neste estado particular. Se estivermos em um estado em que o táxi tem um passageiro e está em cima do destino certo, veríamos uma recompensa de 20 na ação.
# O done é usado para nos dizer quando conseguimos deixar um passageiro no local certo.
# + [markdown] id="qSFd4dC_eJvq"
# ### Resolvendo usando Ações Aleatórias, sem Aprendizagem por Reforço
#
# Vamos ver o que aconteceria se tentarmos usar a força bruta para resolver o problema sem aprendizagem por reforço.
#
# Como temos nossa tabela P de recompensas padrão em cada estado, podemos tentar fazer com que nosso táxi navegue usando apenas isso.
#
# Vamos criar um loop infinito que corre até que um passageiro chegue a um destino, ou em outras palavras, quando a recompensa recebida for 20.
#
# O método env.action_space.sample() seleciona automaticamente uma ação aleatória de um conjunto de todas as ações possíveis.
# + id="jSeXUtwveKEa" colab={"base_uri": "https://localhost:8080/"} outputId="25aa45dc-aa53-449a-d4aa-439f7202ec34"
env.s = 329 # começa no estado do exemplo acima
epochs = 0 # total de ações realizadas
penalties = 0 # quantidade de punições recebidas por pegar ou largar no lugar errado
frames = [] # usado para fazer uma animação
done = False
while not done:
action = env.action_space.sample() # escolhe aleatoriamente uma ação
state, reward, done, info = env.step(action) # aplica a ação e pega o resultado
if reward == -10: # conta uma punição
penalties += 1
# Guarda a sequência para poder fazer a animação depois
frames.append({
'frame': env.render(mode='ansi'),
'state': state,
'action': action,
'reward': reward
}
)
epochs += 1
print("Total de ações executadas: {}".format(epochs))
print("Total de penalizações recebidas: {}".format(penalties))
# + [markdown] id="Cn3osqjIeKjL"
# ### Mostrando a animação dos movimentos realizados
# + id="k8uiG5MBgSXv" colab={"base_uri": "https://localhost:8080/"} outputId="058f5349-e593-4632-c84d-65154945d6d3"
from IPython.display import clear_output
from time import sleep
def print_frames(frames):
for i, frame in enumerate(frames):
clear_output(wait=True)
print(frame['frame'])
print(f"Timestep: {i + 1}")
print(f"State: {frame['state']}")
print(f"Action: {frame['action']}")
print(f"Reward: {frame['reward']}")
sleep(.1)
print_frames(frames)
# + [markdown] id="b2bHbRs7S5o5"
# Não é bom. Nosso agente leva milhares de passos de tempo e faz muitos desembarques errados para entregar apenas um passageiro ao destino certo.
#
# Isso ocorre porque não estamos aprendendo com experiências anteriores. Podemos repetir isso repetidamente e nunca será otimizado. O agente não tem memória de qual ação foi melhor para cada estado, que é exatamente o que o Aprendizado por Reforço fará por nós.
# + [markdown] id="r5kNfr9efDvZ"
# ### Aprendendo a resolver usando Q-Learning
#
# Essencialmente, o Q-learning permite que o agente use as recompensas do ambiente para aprender, com o tempo, a melhor ação a ser executada em um determinado estado.
#
# Em nosso ambiente de táxi, temos a tabela de recompensas P, com a qual o agente aprenderá. Ele procura receber uma recompensa por realizar uma ação no estado atual e, em seguida, atualiza um valor Q para lembrar se essa ação foi benéfica.
#
# Os valores armazenados na tabela Q são chamados de valores Q e são mapeados para uma (state, action)combinação.
#
# Um valor Q para uma determinada combinação de estado-ação é representativo da "qualidade" de uma ação realizada a partir desse estado. Melhores valores de Q implicam em melhores chances de obter maiores recompensas.
#
# Por exemplo, se o táxi se depara com um estado que inclui um passageiro em sua localização atual, é altamente provável que o valor Q para pickup seja maior quando comparado a outras ações, como dropoff ou north.
#
# + id="txSfEmTLeKtK" colab={"base_uri": "https://localhost:8080/"} outputId="3ac0be8a-9f7d-48c9-d111-07bea754ecc3"
import numpy as np
# Inicialização com a tabela de valores Q
# Primeiro, precisamos criar nossa tabela Q, que usaremos para rastrear estados,
# ações e recompensas. O número de estados e ações no ambiente Táxi determina o tamanho da nossa mesa.
q_table = np.zeros([env.observation_space.n, env.action_space.n])
import random
from IPython.display import clear_output
# Hiperparâmetros
alpha = 0.1 # taxa de aprendizagem
gamma = 0.6 # fator de desconto
epsilon = 0.1 # chance de escolha aleatória
# Total geral de ações executadas e penalidades recebidas durante a aprendizagem
epochs, penalties = 0,0
# treinamento do agente
for i in range(1, 100001): # Vai rodar 100000 diferentes versões do problema
state = env.reset() # Inicialização aleatoria do ambient
done = False
while not done:
#épsilon entra em ação quando geramos um valor aleatório entre 0 e 1 e o
#comparamos com nosso épsilon (taxa de exploração), se o valor aleatório for menor,
#executamos uma ação aleatória de nosso espaço de ação
#senão olhamos nossa tabela Q atual e tomamos a ação que maximiza a função de valor.
if random.uniform(0, 1) < epsilon:
action = env.action_space.sample() # Escolhe ação aleatoriamente
else:
action = np.argmax(q_table[state]) # Escolhe ação com base no que já aprendeu
#Depois de escolher uma ação, continuamos com ela e medimos a recompensa associada.
#Isso é feito usando o método embutido env.step (ação) que faz um movimento de um passo de tempo
#Ele retorna o próximo estado, a recompensa da ação anterior;
#done indica se nosso agente atingiu a meta (o que significaria redefinir o ambiente);
# info é apenas um diagnóstico de desempenho usado para depuração.
next_state, reward, done, info = env.step(action) # Aplica a ação
old_value = q_table[state, action] # Valor da ação escolhida no estado atual
next_max = np.max(q_table[next_state]) # Melhor valor no próximo estado
# Atualize o valor Q usando a fórmula principal do Q-Learning
# Finalmente, podemos usar as informações coletadas para atualizar a tabela Q usando a equação de Bellman.
# Lembre-se de que alfa representa a taxa de aprendizado aqui.
new_value = (1 - alpha) * old_value + alpha * (reward + gamma * next_max)
q_table[state, action] = new_value
if reward == -10: # Contabiliza as punições por pegar ou deixar no lugar errado
penalties += 1
state = next_state # Muda de estado
epochs += 1
print("Treinamento finalizado.\n")
print("Total de ações executadas: {}".format(epochs))
print("Total de penalizações recebidas: {}".format(penalties))
# + [markdown] id="GWMOzsS0fEZb"
# ### Mostrando a tabela Q para o estado 329
# Agora que a tabela Q foi estabelecida com mais de 100.000 episódios, vamos ver quais são os valores Q no estado de nossa ilustração:
# + id="5cBAaoqGg1y_" colab={"base_uri": "https://localhost:8080/"} outputId="f65d4a6b-f0f7-43f0-98da-682fabd978ab"
env.s = 329
env.render()
q_table[329]
# + [markdown] id="n2L-4fN2g9YO"
# ### Resolvendo o problema com o aprendizado adquirido
# Vamos avaliar o desempenho do nosso agente. Não precisamos explorar mais as ações, então agora a próxima ação é sempre selecionada usando o melhor valor Q:
# + id="mguXdCZ_rwoN" colab={"base_uri": "https://localhost:8080/"} outputId="6042d7c5-347f-4996-f73c-125cc67a30e7"
state = 329
epochs, penalties = 0, 0
done = False
while not done:
action = np.argmax(q_table[state])
state, reward, done, info = env.step(action)
if reward == -10:
penalties += 1
epochs += 1
print("Total de ações executadas: {}".format(epochs))
print("Total de penalizações recebidas: {}".format(penalties))
# + [markdown] id="ZT1GFNlJWR6O"
# Podemos verificar pela avaliação que o desempenho do agente melhorou significativamente e não incorreu em penalidades, o que significa que realizou as ações corretas de embarque / desembarque com 100 passageiros diferentes.
#
| Exemplo_Q_Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1> Text Classification using TensorFlow/Keras on AI Platform </h1>
#
# This notebook illustrates:
# <ol>
# <li> Creating datasets for AI Platform using BigQuery
# <li> Creating a text classification model using the Estimator API with a Keras model
# <li> Training on Cloud AI Platform
# <li> Rerun with pre-trained embedding
# </ol>
# + id="Nny3m465gKkY" colab_type="code" colab={}
# !sudo chown -R jupyter:jupyter /home/jupyter/training-data-analyst
# -
# !pip install --user google-cloud-bigquery==1.25.0
# **Note**: Restart your kernel to use updated packages.
# Kindly ignore the deprecation warnings and incompatibility errors related to google-cloud-storage.
# change these to try this notebook out
BUCKET = 'cloud-training-demos-ml'
PROJECT = 'cloud-training-demos'
REGION = 'us-central1'
# +
import os
os.environ['BUCKET'] = BUCKET
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
os.environ['TFVERSION'] = '2.6'
if 'COLAB_GPU' in os.environ: # this is always set on Colab, the value is 0 or 1 depending on whether a GPU is attached
from google.colab import auth
auth.authenticate_user()
# download "sidecar files" since on Colab, this notebook will be on Drive
# !rm -rf txtclsmodel
# !git clone --depth 1 https://github.com/GoogleCloudPlatform/training-data-analyst
# !mv training-data-analyst/courses/machine_learning/deepdive/09_sequence/txtclsmodel/ .
# !rm -rf training-data-analyst
# downgrade TensorFlow to the version this notebook has been tested with
# #!pip install --upgrade tensorflow==$TFVERSION
# -
import tensorflow as tf
print(tf.__version__)
# We will look at the titles of articles and figure out whether the article came from the New York Times, TechCrunch or GitHub.
#
# We will use [hacker news](https://news.ycombinator.com/) as our data source. It is an aggregator that displays tech related headlines from various sources.
# ### Creating Dataset from BigQuery
#
# Hacker news headlines are available as a BigQuery public dataset. The [dataset](https://bigquery.cloud.google.com/table/bigquery-public-data:hacker_news.stories?tab=details) contains all headlines from the sites inception in October 2006 until October 2015.
#
# Here is a sample of the dataset:
# %load_ext google.cloud.bigquery
# %%bigquery --project $PROJECT
SELECT
url, title, score
FROM
`bigquery-public-data.hacker_news.stories`
WHERE
LENGTH(title) > 10
AND score > 10
AND LENGTH(url) > 0
LIMIT 10
# Let's do some regular expression parsing in BigQuery to get the source of the newspaper article from the URL. For example, if the url is http://mobile.nytimes.com/...., I want to be left with <i>nytimes</i>
# %%bigquery --project $PROJECT
SELECT
ARRAY_REVERSE(SPLIT(REGEXP_EXTRACT(url, '.*://(.[^/]+)/'), '.'))[OFFSET(1)] AS source,
COUNT(title) AS num_articles
FROM
`bigquery-public-data.hacker_news.stories`
WHERE
REGEXP_CONTAINS(REGEXP_EXTRACT(url, '.*://(.[^/]+)/'), '.com$')
AND LENGTH(title) > 10
GROUP BY
source
ORDER BY num_articles DESC
LIMIT 10
# Now that we have good parsing of the URL to get the source, let's put together a dataset of source and titles. This will be our labeled dataset for AI Platform.
# +
from google.cloud import bigquery
bq = bigquery.Client(project=PROJECT)
query="""
SELECT source, LOWER(REGEXP_REPLACE(title, '[^a-zA-Z0-9 $.-]', ' ')) AS title FROM
(SELECT
ARRAY_REVERSE(SPLIT(REGEXP_EXTRACT(url, '.*://(.[^/]+)/'), '.'))[OFFSET(1)] AS source,
title
FROM
`bigquery-public-data.hacker_news.stories`
WHERE
REGEXP_CONTAINS(REGEXP_EXTRACT(url, '.*://(.[^/]+)/'), '.com$')
AND LENGTH(title) > 10
)
WHERE (source = 'github' OR source = 'nytimes' OR source = 'techcrunch')
"""
df = bq.query(query + " LIMIT 5").to_dataframe()
df.head()
# -
# For ML training, we will need to split our dataset into training and evaluation datasets (and perhaps an independent test dataset if we are going to do model or feature selection based on the evaluation dataset).
#
# A simple, repeatable way to do this is to use the hash of a well-distributed column in our data (See https://www.oreilly.com/learning/repeatable-sampling-of-data-sets-in-bigquery-for-machine-learning).
traindf = bq.query(query + " AND ABS(MOD(FARM_FINGERPRINT(title), 4)) > 0").to_dataframe()
evaldf = bq.query(query + " AND ABS(MOD(FARM_FINGERPRINT(title), 4)) = 0").to_dataframe()
# Below we can see that roughly 75% of the data is used for training, and 25% for evaluation.
#
# We can also see that within each dataset, the classes are roughly balanced.
traindf['source'].value_counts()
evaldf['source'].value_counts()
# Finally we will save our data, which is currently in-memory, to disk.
import os, shutil
DATADIR='data/txtcls'
shutil.rmtree(DATADIR, ignore_errors=True)
os.makedirs(DATADIR)
traindf.to_csv( os.path.join(DATADIR,'train.tsv'), header=False, index=False, encoding='utf-8', sep='\t')
evaldf.to_csv( os.path.join(DATADIR,'eval.tsv'), header=False, index=False, encoding='utf-8', sep='\t')
# !head -3 data/txtcls/train.tsv
# !wc -l data/txtcls/*.tsv
# ### TensorFlow/Keras Code
#
# Please explore the code in this <a href="txtclsmodel/trainer">directory</a>: `model.py` contains the TensorFlow model and `task.py` parses command line arguments and launches off the training job.
#
# In particular look for the following:
#
# 1. [tf.keras.preprocessing.text.Tokenizer.fit_on_texts()](https://www.tensorflow.org/api_docs/python/tf/keras/preprocessing/text/Tokenizer#fit_on_texts) to generate a mapping from our word vocabulary to integers
# 2. [tf.keras.preprocessing.text.Tokenizer.texts_to_sequences()](https://www.tensorflow.org/api_docs/python/tf/keras/preprocessing/text/Tokenizer#texts_to_sequences) to encode our sentences into a sequence of their respective word-integers
# 3. [tf.keras.preprocessing.sequence.pad_sequences()](https://www.tensorflow.org/api_docs/python/tf/keras/preprocessing/sequence/pad_sequences) to pad all sequences to be the same length
#
# The embedding layer in the keras model takes care of one-hot encoding these integers and learning a dense emedding represetation from them.
#
# Finally we pass the embedded text representation through a CNN model pictured below
#
# <img src=images/txtcls_model.png width=25%>
# ### Run Locally (optional step)
# Let's make sure the code compiles by running locally for a fraction of an epoch.
# This may not work if you don't have all the packages installed locally for gcloud (such as in Colab).
# This is an optional step; move on to training on the cloud.
# + language="bash"
# pip install google-cloud-storage
# rm -rf txtcls_trained
# gcloud ai-platform local train \
# --module-name=trainer.task \
# --package-path=${PWD}/txtclsmodel/trainer \
# -- \
# --output_dir=${PWD}/txtcls_trained \
# --train_data_path=${PWD}/data/txtcls/train.tsv \
# --eval_data_path=${PWD}/data/txtcls/eval.tsv \
# --num_epochs=0.1
# -
# ### Train on the Cloud
#
# Let's first copy our training data to the cloud:
# + language="bash"
# gsutil cp data/txtcls/*.tsv gs://${BUCKET}/txtcls/
# + language="bash"
# OUTDIR=gs://${BUCKET}/txtcls/trained_fromscratch
# JOBNAME=txtcls_$(date -u +%y%m%d_%H%M%S)
# gsutil -m rm -rf $OUTDIR
# gcloud ai-platform jobs submit training $JOBNAME \
# --region=$REGION \
# --module-name=trainer.task \
# --package-path=${PWD}/txtclsmodel/trainer \
# --job-dir=$OUTDIR \
# --scale-tier=BASIC_GPU \
# --runtime-version 2.3 \
# --python-version 3.7 \
# -- \
# --output_dir=$OUTDIR \
# --train_data_path=gs://${BUCKET}/txtcls/train.tsv \
# --eval_data_path=gs://${BUCKET}/txtcls/eval.tsv \
# --num_epochs=5
# -
# Change the job name appropriately. View the job in the console, and wait until the job is complete.
# !gcloud ai-platform jobs describe txtcls_190209_224828
# ### Results
# What accuracy did you get? You should see around 80%.
# ### Rerun with Pre-trained Embedding
#
# We will use the popular GloVe embedding which is trained on Wikipedia as well as various news sources like the New York Times.
#
# You can read more about Glove at the project homepage: https://nlp.stanford.edu/projects/glove/
#
# You can download the embedding files directly from the stanford.edu site, but we've rehosted it in a GCS bucket for faster download speed.
# !gsutil cp gs://cloud-training-demos/courses/machine_learning/deepdive/09_sequence/text_classification/glove.6B.200d.txt gs://$BUCKET/txtcls/
# Once the embedding is downloaded re-run your cloud training job with the added command line argument:
#
# ` --embedding_path=gs://${BUCKET}/txtcls/glove.6B.200d.txt`
#
# While the final accuracy may not change significantly, you should notice the model is able to converge to it much more quickly because it no longer has to learn an embedding from scratch.
# #### References
# - This implementation is based on code from: https://github.com/google/eng-edu/tree/master/ml/guides/text_classification.
# - See the full text classification tutorial at: https://developers.google.com/machine-learning/guides/text-classification/
#
# ## Next step
# Client-side tokenizing in Python is hugely problematic. See <a href="text_classification_native.ipynb">Text classification with native serving</a> for how to carry out the preprocessing in the serving function itself.
# Copyright 2020 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| courses/machine_learning/deepdive/09_sequence/text_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Fair Dummies: Synthetic Example
#
# This notebook implements a the fair dummies framework for learning predictive models that approximately satisfy the equalized odds notion of fairness.
#
# Paper: "Achieving Equalized Odds by Resampling Sensitive Attributes," <NAME>, <NAME>, and <NAME>, 2020
#
# ### Proposed approach
#
# __Core idea__: fit a regression function, minimizing
#
# $$ \text{loss = prediction error + distance to equalized odds}$$
#
#
# __Input__: $ \{(X_i,A_i,Y_i)\}_{i=1}^n \sim P_{XAY}$ training data
#
#
# __Step 1__: sample dummy protected attributes
# $$
# \tilde{A}_i \sim P_{A|Y}(A \mid Y=y_{i}) \quad \forall \ i=1,2,\dots,n \nonumber
# $$
#
# $A \in \{0,1\}$? generate $\tilde{A}$ using a biased coin-flip, with
# $$
# P\{A=1|Y=y\} = \frac{P\{y \mid A=1\}P\{A=1\}}{P\{y \mid A=1\}P\{A=1\} + P\{y \mid A=0\}P\{A=0\}}
# $$
#
# __Step 2__: fit a regression function on $\{(X_i, A_i, Y_i)\}_{i=1}^n$
#
# $$
# \hat{f}(\cdot) \,= \, \underset{f \in \mathcal{F}}{\mathrm{arg min}} \, \frac{1-\lambda}{n} \sum_{i=1}^n (Y_i - f(X_i))^2 + \lambda \mathcal{D}\left( [\hat{\mathbf{Y}}, \mathbf{A}, \mathbf{Y}] , [\hat{\mathbf{Y}}, \tilde{\mathbf{A}}, \mathbf{Y}] \right) \nonumber
# $$
#
# where
#
# $$
# \hat{\mathbf{Y}} = \left[f(X_{1}), f(X_{2}), \dots, f(X_{n})\right]^T \ ; \ \mathbf{A} = \left[A_{1}, A_{2}, \dots, A_{n}\right]^T \ ; \ \tilde{\mathbf{A}} = \left[\tilde{A}_{1}, \tilde{A}_{2}, \dots, \tilde{A}_{n}\right]^T \ ; \ \mathbf{Y} = \left[Y_{1}, Y_{2}, \dots, Y_{n}\right]^T
# $$
#
# and $\mathcal{D}\left( \mathbf{Z}_1, \mathbf{Z}_2 \right)$ tests whether $P_{Z_1} = \ P_{Z_2}$ given $\mathbf{Z}_1, \mathbf{Z}_2$, here it implemented as a classifier two-sample test
# +
from sklearn.preprocessing import StandardScaler
import matplotlib
matplotlib.use('Agg')
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
np.warnings.filterwarnings('ignore')
import random
import torch
import torch.nn as nn
import pandas as pd
import os
import sys
sys.path.append(os.path.abspath(os.path.join(os.getcwd() + '/others/third_party/cqr')))
base_path = os.getcwd() + '/data/'
from fair_dummies import fair_dummies_learning
from fair_dummies import utility_functions
seed = 123
random.seed(seed)
np.random.seed(seed)
torch.manual_seed(seed)
# maximal number of testpoints to plot
max_show = 100
# save figures?
save_figures = False
# display results
if save_figures:
SMALL_SIZE = 26
else:
SMALL_SIZE = 18
MEDIUM_SIZE = SMALL_SIZE
BIGGER_SIZE = SMALL_SIZE
plt.rc('font', size=SMALL_SIZE) # controls default text sizes
plt.rc('axes', titlesize=SMALL_SIZE) # fontsize of the axes title
plt.rc('axes', labelsize=MEDIUM_SIZE) # fontsize of the x and y labels
plt.rc('xtick', labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc('ytick', labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc('legend', fontsize=SMALL_SIZE) # legend fontsize
plt.rc('figure', titlesize=BIGGER_SIZE) # fontsize of the figure title
plt.rc('text', usetex=True)
# +
def plot_func(x,
y,
y_u=None,
y_l=None,
pred=None,
point_color="b.",
shade_color="",
method_name="",
title="",
filename=None,
save_figures=False,
show_legend=True):
""" Scatter plot of (x,y) points along with the constructed prediction interval
Parameters
----------
x : numpy array, corresponding to the feature of each of the n samples
y : numpy array, target response variable (length n)
pred : numpy array, the estimated prediction. It may be the conditional mean,
or low and high conditional quantiles.
shade_color : string, desired color of the prediciton interval
method_name : string, name of the method
title : string, the title of the figure
filename : sting, name of the file to save the figure
save_figures : boolean, save the figure (True) or not (False)
"""
inds = np.random.permutation(x.shape[0])[:max_show]
x_ = x[inds]
y_ = y[inds]
if y_u is not None:
y_u_ = y_u[inds]
if y_l is not None:
y_l_ = y_l[inds]
if pred is not None:
pred_ = pred[inds]
fig = plt.figure()
inds = np.argsort(np.squeeze(x_))
if (y_u is not None) and (y_l is not None):
plt.fill_between(x_[inds].squeeze(),
y_u_[inds],
y_l_[inds],
alpha=.2,
color=shade_color,
edgecolor="",
label = u'Prediction interval')
if pred is not None:
if pred_.ndim == 2:
plt.plot(x_[inds,:], pred_[inds,0], 'k', lw=2, alpha=0.2,
label=u'Predicted low and high quantiles')
plt.plot(x_[inds,:], pred_[inds,1], 'k', lw=2, alpha=0.2)
else:
plt.plot(x_[inds,:], pred_[inds], point_color, lw=2, alpha=0.8, markersize=15,
fillstyle='none')
min_val = -8
max_val = 8
ident = [min_val, max_val]
plt.xlim(ident[0], ident[1])
plt.ylim(ident[0], ident[1])
plt.plot(ident,ident, ls="--", c=".3")
plt.xlabel(r'$Y$')
plt.ylabel(r'$\hat{Y}$')
ax = plt.gca()
ax.set_aspect('equal', 'box')
plt.title(title)
if save_figures and (filename is not None):
plt.savefig(filename, bbox_inches='tight', dpi=300)
plt.show()
def plot_groups_pointwise(x_axis_0,x_axis_1,A,Y,Yhat,
y_u=None,y_l=None,point_color="b.",shade="",
filename_0=None,filename_1=None,save_figures=False):
inds = A==0
y_u0 = None
y_l0 = None
if (y_u is not None) and (y_l is not None):
y_u0=y_u[inds]
y_l0=y_l[inds]
plot_func(x_axis_0.reshape(x_axis_0.shape[0],1),
Y[inds], y_u=y_u0, y_l=y_l0, pred=Yhat[inds],
point_color=point_color,shade_color=shade, method_name="", title=r'$A=0$',
filename=filename_0, save_figures=save_figures,show_legend=True)
inds = A==1
y_u1 = None
y_l1 = None
if (y_u is not None) and (y_l is not None):
y_u1=y_u[inds]
y_l1=y_l[inds]
plot_func(x_axis_1.reshape(x_axis_1.shape[0],1),
Y[inds], y_u=y_u1, y_l=y_l1, pred=Yhat[inds],
point_color=point_color,shade_color=shade, method_name="", title=r'$A=1$',
filename=filename_1, save_figures=save_figures,show_legend=False)
# -
# ## Generate data
#
# +
def init_seed(seed = 0):
random.seed(seed)
np.random.seed(seed)
torch.manual_seed(seed)
def synthetic_example(include_A=False, n = 6000):
p0 = 0.9
p1 = 1 - p0
A = np.random.binomial(1,p1,n).T
X = np.random.randn(n,2)
beta0 = [2,1]
beta1 = [1,2]
Y = np.random.randn(n)
Y[A==0] = Y[A==0] + np.dot(X[A==0],beta0)
Y[A==1] = Y[A==1] + np.dot(X[A==1],beta1)
x_axis_0 = Y[A==0]
x_axis_1 = Y[A==1]
if include_A:
X = np.concatenate((X,A[:,np.newaxis]),axis=1)
return X, A, Y, x_axis_0, x_axis_1, beta0, beta1
include_A = False
init_seed(seed)
X, A, Y, x_axis_0, x_axis_1, beta0, beta1 = synthetic_example(include_A=include_A, n = 500)
X_cal, A_cal, Y_cal, x_axis_0, x_axis_1, beta0, beta1 = synthetic_example(include_A=include_A, n = 2000)
X_test, A_test, Y_test, x_axis_0_test, x_axis_1_test, beta0, beta1 = synthetic_example(include_A=include_A, n = 2000)
# compute input dimensions
n = X.shape[0]
in_shape = X.shape[1]
print(X.shape)
print(Y.shape)
# -
# Estimate $P_{A|Y}$ using kernel density estimator, then compute $P\{A=1|Y=y\}$
# +
from sklearn.neighbors import KernelDensity
def compute_density(Z,A,show_graphs=False,Z_test=[]):
bandwidth = np.sqrt(np.median(np.abs(Z)))
kde_0 = KernelDensity(kernel='linear', bandwidth=bandwidth).fit(Z[A==0][:, np.newaxis])
kde_1 = KernelDensity(kernel='linear', bandwidth=bandwidth).fit(Z[A==1][:, np.newaxis])
if show_graphs:
plt.clf()
plt.hist(Z[A==0], fc='#AAAAFF', density=True)
plt.show()
X_plot = np.linspace(min(Z[A==0])-1, max(Z[A==0])+1, 1000)[:, np.newaxis]
log_dens = kde_0.score_samples(X_plot)
plt.clf()
plt.fill(X_plot[:, 0], np.exp(log_dens), fc='#AAAAFF')
plt.show()
log_dens_0 = np.exp(np.squeeze(kde_0.score_samples(Z[:, np.newaxis])))
log_dens_1 = np.exp(np.squeeze(kde_1.score_samples(Z[:, np.newaxis])))
p_0 = np.sum(A==0) / A.shape[0]
p_1 = 1 - p_0
# p(A=1|y) = p(y|A=1)p(A=1) / (p(y|A=1)p(A=1) + p(y|A=0)p(A=0))
p_success = (log_dens_1*p_1) / (log_dens_1*p_1 + log_dens_0*p_0)
p_success_test = []
if len(Z_test) > 0:
log_dens_0_test = np.exp(np.squeeze(kde_0.score_samples(Y_test[:, np.newaxis])))
log_dens_1_test = np.exp(np.squeeze(kde_1.score_samples(Y_test[:, np.newaxis])))
p_success_test = (log_dens_1_test*p_1) / (log_dens_1_test*p_1 + log_dens_0_test*p_0)
return p_success, p_success_test
p_success, p_success_test = compute_density(Y,A,True,Y_test)
# -
# Implementation of a fairness-unaware baseline algorithm
# +
# step size
lr = 0.01
# inner epochs to fit adversary
dis_steps = 1
# inner epochs to fit loss
loss_steps = 1
batch_size = 6000
# utility loss
cost_pred = torch.nn.MSELoss()
in_shape = X.shape[1]
out_shape = 1
model_type = "deep_model"
# equalized odds penalty
mu = 0
second_moment_scaling = 0
# total number of epochs
epochs = 500
base_reg = fair_dummies_learning.EquiRegLearner(lr=lr,
pretrain_pred_epochs=0,
pretrain_dis_epochs=0,
epochs=epochs,
loss_steps=loss_steps,
dis_steps=dis_steps,
cost_pred=cost_pred,
in_shape=in_shape,
batch_size=batch_size,
model_type=model_type,
lambda_vec=mu,
second_moment_scaling=second_moment_scaling,
out_shape=out_shape)
init_seed(seed)
input_data_train = np.concatenate((A[:,np.newaxis],X),1)
base_reg.fit(input_data_train, Y)
input_data_cal = np.concatenate((A_cal[:,np.newaxis],X_cal),1)
Yhat_out_cal = base_reg.predict(input_data_cal)
input_data_test = np.concatenate((A_test[:,np.newaxis],X_test),1)
Yhat_out_test = Yhat_test = base_reg.predict(input_data_test)
# +
# bhat, res, rank, s = np.linalg.lstsq(X, Y)
# Yhat_out_cal = np.dot(X_cal,bhat)
# Yhat_out_test = np.dot(X_test,bhat)
print("Baseline (All): Test Error = " + str(np.sqrt(np.mean((Yhat_out_test - Y_test)**2))))
print("Baseline (A=0): Test Error = " + str(np.sqrt(np.mean((Yhat_out_test[A_test==0] - Y_test[A_test==0])**2))))
print("Baseline (A=1): Test Error = " + str(np.sqrt(np.mean((Yhat_out_test[A_test==1] - Y_test[A_test==1])**2))))
# +
plot_groups_pointwise(x_axis_0_test,
x_axis_1_test,
A_test,
Y_test,
Yhat_out_test,
y_u=None,
y_l=None,
point_color='r.',
shade=None,
filename_0="supp_synth_no_use_A_baseline_0.png",
filename_1="supp_synth_no_use_A_baseline_1.png",
save_figures=save_figures)
p_val_base = utility_functions.fair_dummies_test_regression(Yhat_out_cal,
A_cal,
Y_cal,
Yhat_out_test,
A_test,
Y_test,
num_reps = 1,
num_p_val_rep=1000,
reg_func_name="Net",
lr = 0.01,
return_vec=True)
plt.clf()
x, bins, p=plt.hist(x=p_val_base, bins=30, color='salmon', alpha=0.7, range=(0, 1), density=True, ec='red', align='left', label=u'Baseline')
plt.grid(axis='y', alpha=0.75)
plt.xlabel('Value')
plt.ylabel('Frequency')
plt.title('Fair Dummies Test')
for item in p:
item.set_height(item.get_height()/sum(x))
plt.ylim(0, 1)
plt.show()
# +
# step size
lr = 0.01
# inner epochs to fit adversary
dis_steps = 40
# inner epochs to fit loss
loss_steps = 40
batch_size = 6000
# utility loss
cost_pred = torch.nn.MSELoss()
in_shape = X.shape[1]
out_shape = 1
model_type = "deep_model"
# equalized odds penalty
mu = 0.99
second_moment_scaling = 0
# total number of epochs
epochs = 500
# +
fair_reg = fair_dummies_learning.EquiRegLearner(lr=lr,
pretrain_pred_epochs=0,
pretrain_dis_epochs=0,
epochs=epochs,
loss_steps=loss_steps,
dis_steps=dis_steps,
cost_pred=cost_pred,
in_shape=in_shape,
batch_size=batch_size,
model_type=model_type,
lambda_vec=mu,
second_moment_scaling=second_moment_scaling,
out_shape=out_shape)
init_seed(seed)
input_data_train = np.concatenate((A[:,np.newaxis],X),1)
fair_reg.fit(input_data_train, Y)
input_data_cal = np.concatenate((A_cal[:,np.newaxis],X_cal),1)
Yhat_out_cal = fair_reg.predict(input_data_cal)
input_data_test = np.concatenate((A_test[:,np.newaxis],X_test),1)
Yhat_out_test = Yhat_test = fair_reg.predict(input_data_test)
# -
print("Fair Dummies (All): Test Error = " + str(np.sqrt(np.mean((Yhat_out_test - Y_test)**2))))
print("Fair Dummies (A=0): Test Error = " + str(np.sqrt(np.mean((Yhat_out_test[A_test==0] - Y_test[A_test==0])**2))))
print("Fair Dummies (A=1): Test Error = " + str(np.sqrt(np.mean((Yhat_out_test[A_test==1] - Y_test[A_test==1])**2))))
# +
plot_groups_pointwise(x_axis_0_test,
x_axis_1_test,
A_test,
Y_test,
Yhat_out_test,
y_u=None,
y_l=None,
point_color='b.',
shade=None,
filename_0="supp_synth_no_use_A_equi_0.png",
filename_1="supp_synth_no_use_A_equi_1.png",
save_figures=save_figures)
p_val_equi = utility_functions.fair_dummies_test_regression(Yhat_out_cal,
A_cal,
Y_cal,
Yhat_out_test,
A_test,
Y_test,
num_reps = 1,
num_p_val_rep=1000,
reg_func_name="Net",
lr = 0.01,
return_vec=True)
plt.clf()
x_equi, bins_equi, p_equi=plt.hist(x=p_val_equi, bins=30, color='royalblue', alpha=0.7, range=(0, 1), density=True, ec='blue', align='left', label=u'Equitable model')
x_base, bins_base, p_base=plt.hist(x=p_val_base, bins=30, color='salmon', alpha=0.7, range=(0, 1), density=True, ec='red', align='left', label=u'Baseline model')
plt.grid(axis='y', alpha=0.75)
plt.xlabel('p-value')
plt.ylabel('Frequency')
plt.title('Fair Dummies Test')
for item in p_equi:
item.set_height(item.get_height()/sum(x_equi))
for item in p_base:
item.set_height(item.get_height()/sum(x_base))
plt.ylim(0, 1)
plt.legend(bbox_to_anchor=(1.05, 1.0), loc='upper left')
plt.show()
# -
| synthetic_experiment2_without_A_as_feature.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import sys
sys.path.append('../RenderMan/Builds/LinuxMakefile/build/')
sys.path.append('../dexed/Builds/Linux/build/')
import librenderman as rm
# +
# Important settings. These are good general ones.
sampleRate = 44100
bufferSize = 512
fftSize = 512
# This will host a VST. It will render the features and audio we need.
engine = rm.RenderEngine(sampleRate, bufferSize, fftSize)
# Load the VST into the RenderEngine.
path = "../dexed/Builds/Linux/build/Dexed.so"
path = "../amsynth-1.8.0/.libs/amsynth_vst.so"
engine.load_plugin(path)
# Create a patch generator. We can initialise it to generate the correct
# patches for a given synth by passing it a RenderEngine which has
# loaded a instance of the synthesiser.
generator = rm.PatchGenerator(engine)
# We can also get a string of information about the
# available parameters.
print engine.get_plugin_parameters_description()
# +
# Get a random patch and set it.
new_patch = generator.get_random_patch()
engine.set_patch(new_patch)
# We need to override some parameters to prevent hanging notes in
# Dexed.
overriden_parameters = [(26, 1.), (30, 0.), (48, 1.), (52, 0.),
(70, 1.), (74, 0.), (92, 1.), (96, 0.),
(114, 1.), (118, 0.), (136, 1.), (140, 0.)]
# Loop through each tuple, unpack it and override the correct
# parameter with the correct value to prevent hanging notes.
for parameter in overriden_parameters:
index, value = parameter
engine.override_plugin_parameter(index, value)
# Settings to play a note and extract data from the synth.
midiNote = 40
midiVelocity = 127
noteLength = 1.0
renderLength = 1.0
# Render the data.
engine.render_patch(midiNote, midiVelocity, noteLength, renderLength)
# Get the data. Note the audio is automattically made mono, no
# matter what channel size for ease of use.
audio = engine.get_audio_frames()
mfccs = engine.get_mfcc_frames()
# -
from IPython.display import Audio
Audio(audio, rate=sampleRate)
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
plt.contourf(np.array(mfccs).T)
plt.colorbar()
plt.show()
plt.plot(np.mean(np.array(mfccs).T,axis=1))
plt.show()
# + active=""
#
| notebooks/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Search and Geocode Tutorial
# This is the completed solution for the [Search and geocode](https://developers.arcgis.com/labs/develop/python/search-and-geocode/) ArcGIS tutorial.
#
# [ArcGIS tutorials](https://developers.arcgis.com/labs/) are short guides demonstrating the three phases of building geospatial apps: Data, Design, Develop.
from arcgis.gis import GIS
from arcgis.geocoding import geocode, reverse_geocode
from arcgis.geometry import Point
gis = GIS()
# ## Geocode place names
geocode_result = geocode(address="Hollywood sign", as_featureset=True)
geocode_result.features
map = gis.map("Los Angeles, CA", zoomlevel=11)
map
map.draw(geocode_result)
map.clear_graphics()
# ## Reverse geocode a coordinate
# +
location = {
'Y': 34.13419,
'X': -118.29636,
'spatialReference': {
'wkid':4326
}
}
unknown_pt = Point(location)
# -
address = reverse_geocode(unknown_pt)
address
map.draw(address)
| labs/search_and_geocode.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# language: python
# name: python3
# ---
import pandas as pd
import sys
import torch
import pytorch_lightning as pl
import matplotlib.pyplot as plt
import numpy as np
from pytorch_lightning.utilities.cli import LightningCLI
# +
# Add the example model/etc.
import baseline_example
from baseline_example import InvestmentEulerBaseline
# utility to make calling with CLI defaults easier. Does not log or use early stopping callbacks
def solve_model(Model, args, default_seed = 123):
sys.argv = ["dummy.py"] + [f"--{key}={val}" for key, val in args.items()] # hack overwriting argv
cli = LightningCLI(
Model,
run=False,
seed_everything_default=default_seed,
save_config_overwrite=True,
parser_kwargs={"default_config_files": ["baseline_example_defaults.yaml"]},
)
# Solves the model
trainer = cli.instantiate_trainer(
logger=None,
checkpoint_callback=None,
callbacks=[], # not using the early stopping/etc.
)
trainer.fit(cli.model)
# Calculates the "test" values for it
trainer.test(cli.model)
cli.model.eval() # Turn off training mode, where it calculates gradients for every call.
return cli.model, cli
# -
# Load using the cli options, then run to fit the model
model, cli = solve_model(InvestmentEulerBaseline, {"trainer.max_epochs" : 5, "model.verbose": True})
# +
# Plot the results of the previous cell
df = model.test_results[
model.test_results["ensemble"] == 0
] # first ensemble in dataframe from model on test data
fig, ax = plt.subplots()
ax.plot(df["t"], df["u_hat"], label=r"$u(X_t)$, $\phi($ReLU$)$")
if model.hparams.nu == 1.0: # only add reference line if linear
ax.plot(df["t"], df["u_reference"], dashes=[10, 5, 10, 5], label=r"$u(X_t)$, LQ")
ax.legend()
ax.set_title(r"$u(X_t)$ with $\phi($ReLU$)$ : Equilibrium Path")
ax.set_xlabel(r"Time($t$)")
# -
# Example to evaluate model after fitting it, can do at multiple points at same time:
model.eval() # TODO: why is the `no_grad` also needed?
X_points = torch.stack((model.X_0 + 0.001 * torch.randn(model.hparams.N),
model.X_0 + 0.05 * torch.ones(model.hparams.N)))
# evaluate the policy. Doing it with a `no_grad` can speed things up
with torch.no_grad():
u_X = model(X_points)
print(u_X)
| baseline_example.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Q#
# language: qsharp
# name: iqsharp
# ---
# # Grover's Algorithm
#
# The **Grover's Search** quantum kata is a series of exercises designed
# to get you familiar with Grover's search algorithm.
#
# It covers the following topics:
#
# * writing oracles for Grover's search,
# * performing steps of the algorithm, and
# * putting it all together: Grover's search algorithm.
#
# *Reading material:*
#
# * [This Microsoft Learn module](https://docs.microsoft.com/en-us/learn/modules/solve-graph-coloring-problems-grovers-search/) offers a different, visual explanation of Grover's algorithm.
# * The tasks follow the explanation from *Quantum Computation and Quantum Information* by <NAME> Chuang.
# In the 10th anniversary edition, this is section 6.1.2 on pages 248-251.
# * A different explanation of Grover's algorithm can be found in
# [this Wikipedia article](https://en.wikipedia.org/wiki/Grover%27s_algorithm).
# * [An Introduction to Quantum Algorithms](https://people.cs.umass.edu/~strubell/doc/quantum_tutorial.pdf) by <NAME>, pages 20-24.
# * [Lecture 4: Grover's Algorithm](https://www.cs.cmu.edu/~odonnell/quantum15/lecture04.pdf) by <NAME>.
# * Lectures [12](https://cs.uwaterloo.ca/~watrous/QC-notes/QC-notes.12.pdf) and [13](https://cs.uwaterloo.ca/~watrous/QC-notes/QC-notes.13.pdf) by <NAME>.
# * [This page](http://davidbkemp.github.io/animated-qubits/grover.html) has an animated demonstration of Grover's algorithm for a simple case.
#
# Each task is wrapped in one operation preceded by the description of the task.
# Your goal is to fill in the blanks (marked with the `// ...` comments)
# with some Q# code that solves the task. To verify your answer, run the cell with Ctrl+Enter (⌘+Enter on macOS).
#
# Within each section, tasks are given in approximate order of increasing difficulty;
# harder ones are marked with asterisks.
# ## Part I. Oracles for Grover's Search
#
# ### Task 1.1. The $|11...1\rangle$ Oracle
# **Inputs:**
#
# 1. N qubits in an arbitrary state $|x\rangle$ (input/query register)
#
# 2. A qubit in an arbitrary state $|y\rangle$ (target qubit)
#
# **Goal:**
#
# Flip the state of the target qubit (i.e., apply an X gate to it)
# if the query register is in the $|11...1\rangle$ state,
# and leave it unchanged if the query register is in any other state.
# Leave the query register in the same state it started in.
#
# **Examples:**
#
# * If the query register is in state $|00...0\rangle$, leave the target qubit unchanged.
#
# * If the query register is in state $|10...0\rangle$, leave the target qubit unchanged.
#
# * If the query register is in state $|11...1\rangle$, flip the target qubit.
#
# * If the query register is in state $\frac{1}{\sqrt{2}} \big(|00...0\rangle + |11...1\rangle \big)$, and the target is in state $|0\rangle$,
# the joint state of the query register and the target qubit should be $\frac{1}{\sqrt{2}} \big(|00...00\rangle + |11...11\rangle \big)$.
# +
%kata T11_Oracle_AllOnes
operation Oracle_AllOnes (queryRegister : Qubit[], target : Qubit) : Unit is Adj {
Controlled X(queryRegister, target);
}
# -
# ### Task 1.2. The $|1010...\rangle$ Oracle
#
# **Inputs:**
#
# 1. N qubits in an arbitrary state $|x\rangle$ (input/query register)
#
# 2. A qubit in an arbitrary state $|y\rangle$ (target qubit)
#
# **Goal:**
#
# Flip the state of the target qubit if the query register is in the $|1010...\rangle$ state;
# that is, the state with alternating 1 and 0 values, with any number of qubits in the register.
# Leave the state of the target qubit unchanged if the query register is in any other state.
# Leave the query register in the same state it started in.
#
# **Examples:**
#
# * If the register is in state $|0000000\rangle$, leave the target qubit unchanged.
# * If the register is in state $|10101\rangle$, flip the target qubit.
# +
%kata T12_Oracle_AlternatingBits
operation Oracle_AlternatingBits (queryRegister : Qubit[], target : Qubit) : Unit is Adj {
within {
for i in 1 .. 2 .. Length(queryRegister) - 1 {
X(queryRegister[i]);
}
}
apply {
Controlled X(queryRegister, target); }
}
# -
# ### Task 1.3. Arbitrary Bit Pattern Oracle
#
# **Inputs:**
#
# 1. N qubits in an arbitrary state $|x\rangle$ (input/query register)
#
# 2. A qubit in an arbitrary state $|y\rangle$ (target qubit)
#
# 3. A bit pattern of length N represented as `Bool[]`
#
# **Goal:**
#
# Flip the state of the target qubit if the query register is in the state described by the given bit pattern
# (`true` represents qubit state One, and `false` represents Zero).
# Leave the state of the target qubit unchanged if the query register is in any other state.
# Leave the query register in the same state it started in.
#
# **Example:**
#
# If the bit pattern is `[true, false]`, you need to flip the target qubit if and only if the qubits are in the $|10\rangle$ state.
# +
%kata T13_Oracle_ArbitraryPattern
operation Oracle_ArbitraryPattern (queryRegister : Qubit[], target : Qubit, pattern : Bool[]) : Unit is Adj {
// ...
}
# -
# ### Task 1.4. Oracle Converter
#
# **Input:**
#
# A marking oracle: an oracle that takes a register and a target qubit and
# flips the target qubit if the register satisfies a certain condition.
#
# **Output:**
#
# A phase-flipping oracle: an oracle that takes a register and
# flips the phase of the register if it satisfies this condition.
#
# > Grover's algorithm relies on the search condition implemented as a phase-flipping oracle,
# but it is often easier to write a marking oracle for a given condition. This transformation
# allows to convert one type of oracle into the other. The transformation is described in the
# [Wikipedia article on Grover's algorithm](https://en.wikipedia.org/wiki/Grover%27s_algorithm), in the section "Description of ${U_\omega}$".
#
# <br/>
# <details>
# <summary><b>Need a hint? Click here</b></summary>
# Remember that you can define auxiliary operations. To do that, you'll need to create an extra code cell for each new operation and execute it before returning to this cell.
# </details>
# +
%kata T14_OracleConverter
function OracleConverter (markingOracle : ((Qubit[], Qubit) => Unit is Adj)) : (Qubit[] => Unit is Adj) {
// ...
}
# -
# ## Part II. The Grover Iteration
# ### Task 2.1. The Hadamard Transform
#
# **Input:** A register of N qubits in an arbitrary state
#
# **Goal:** Apply the Hadamard transform to each of the qubits in the register.
#
# > If the register started in the $|0...0\rangle$ state, this operation
# will prepare an equal superposition of all $2^{N}$ basis states.
# +
%kata T21_HadamardTransform
operation HadamardTransform (register : Qubit[]) : Unit is Adj {
ApplyToEachA(H, register);
}
# -
# ### Task 2.2. Conditional Phase Flip
#
# **Input:** A register of N qubits in an arbitrary state.
#
# **Goal:** Flip the sign of the state of the register if it is not in the $|0...0\rangle$ state.
#
# **Examples:**
#
# * If the register is in state $|0...0\rangle$, leave it unchanged.
#
# * If the register is in any other basis state, multiply its phase by -1.
#
# > This operation implements operator $2|0...0\rangle\langle0...0| - I$ $ = \left(\begin{matrix}1&0&...&0\\0&-1&...&0\\\vdots&\vdots&\ddots&\vdots\\0&0&...&-1\end{matrix}\right) $
#
# <br/>
# <details>
# <summary><b>Hint #1</b></summary>
# Note that quantum states are defined up to a global phase.
# Thus the state obtained as a result of this operation is equivalent to the state obtained by flipping the sign of only the $|0...0\rangle$ basis state (those states differ by a global phase $-1$).<br>
# $$-\big(2|0...0\rangle\langle0...0| - I\big) = I - 2|0...0\rangle\langle0...0| = \left(\begin{matrix}-1&0&...&0\\0&1&...&0\\\vdots&\vdots&\ddots&\vdots\\0&0&...&1\end{matrix}\right) $$<br>
#
# It doesn't matter for Grover's search algorithm itself, since the global phase is not observable, but can have side effects when used as part of other algorithms.
# > See the extended discussion in this [Quantum Computing SE question](https://quantumcomputing.stackexchange.com/questions/5973/counting-in-q-number-of-solutions/6446#6446)<br>
# </details>
# <br/>
#
# <details>
# <summary><b>Hint #2</b></summary>
# Consider the Controlled Z gate, applied with most of the qubits as control and the last qubit as target:
# $\text{Controlled Z}(|s_0 s_1 \ldots s_{n-2}\rangle, |s_{n-1}\rangle)$ leaves all basis states except $|1...11\rangle$ unchanged, and adds a $-1$ phase to that state: $|1...11\rangle \rightarrow -|1...11\rangle$ (remember that $Z|0\rangle = |0\rangle$ and $Z|1\rangle = -|1\rangle$).
# You need to modify it to add the $-1$ phase to only the $|0...00\rangle$ state instead.
# <br/><br/>
# Alternatively, you can use the same trick as in the oracle converter task.<br>
# </details>
# <br>
#
# <details>
# <summary><b>Hint #3</b></summary>
# You can use the <a href="https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic.r">R gate</a> to correct the global phase.
# </details>
# +
%kata T22_ConditionalPhaseFlip
operation ConditionalPhaseFlip (register : Qubit[]) : Unit is Adj {
let allZerosOracle = Oracle_ArbitraryPattern_Reference(_, _, new Bool[Length(register)]);
// Convert it into a phase-flip oracle and apply it
let flipOracle = OracleConverter_Reference(allZerosOracle);
flipOracle(register);
// To fix the global phase difference, use the following line :
R(PauliI, 2.0 * PI(), register[0]); // Note : We used 2*PI to add a global phase of PI, as R operation rotates qubit by 𝜃/2
// For more details refer to the following Quantum SE question : https://quantumcomputing.stackexchange.com/questions/5973/counting-in-q-number-of-solutions/6446#6446
}
# -
# ### Task 2.3. The Grover Iteration
#
# **Inputs:**
#
# 1. N qubits in an arbitrary state $|x\rangle$ (input/query register)
#
# 2. A phase-flipping oracle that takes an N-qubit register and flips
# the phase of the state if the register is in the desired state.
#
# **Goal:** Perform one Grover iteration.
#
# <br/>
# <details>
# <summary><b>Need a hint? Click here</b></summary>
# A Grover iteration consists of 4 steps:
# <ol>
# <li>Apply the Oracle</li>
# <li>Apply the Hadamard transform</li>
# <li>Perform a conditional phase shift</li>
# <li>Apply the Hadamard transform again</li>
# </ol>
# </details>
# +
%kata T23_GroverIteration
operation GroverIteration (register : Qubit[], oracle : (Qubit[] => Unit is Adj)) : Unit is Adj {
oracle(register);
within {
HadamardTransform_Reference(register);
} apply {
ConditionalPhaseFlip_Reference(register);
}
}
# -
# ## Part III. Putting It All Together: Grover's Search Algorithm
# ### Task 3.1. Grover's Search
#
# **Inputs:**
#
# 1. N qubits in the $|0...0\rangle$ state.
#
# 2. A marking oracle.
#
# 3. The number of Grover iterations to perform.
#
# **Goal:** Use Grover's algorithm to leave the register in the state that is marked by the oracle as the answer (with high probability).
#
# > The number of iterations is passed as a parameter because it is defined by the nature of the problem
# and is easier to configure/calculate outside the search algorithm itself (for example, in the classical driver).
# +
%kata T31_GroversSearch
operation GroversSearch (register : Qubit[], oracle : ((Qubit[], Qubit) => Unit is Adj), iterations : Int) : Unit {
// ...
}
# -
# ### Task 3.2. Using Grover's Search
#
# **Goal:** Use your implementation of Grover's Algorithm from Task 3.1 and the oracles from part 1
# to find the marked elements of the search space. This task is not covered by a test and allows you to experiment with running the algorithm.
#
# > This is an open-ended task, and is not covered by a unit test. To run the code, execute the cell with the definition of the `Run_GroversSearch_Algorithm` operation first; if it compiled successfully without any errors, you can run the operation by executing the next cell (`%simulate Run_GroversSearch_Algorithm`).
#
# > Note that this task relies on your implementations of the previous tasks. If you are getting the "No variable with that name exists." error, you might have to execute previous code cells before retrying this task.
#
# <details closed>
# <summary><b>Hint #1</b></summary>
# To check whether the algorithm found the correct answer (i.e., an answer marked as 1 by the oracle),
# you can apply the oracle once more to the register after you've measured it and an ancilla qubit,
# which will calculate the function of the answer found by the algorithm.
# </details>
# <br/>
# <details closed>
# <summary><b>Hint #2</b></summary>
# Experiment with the number of iterations to see how it affects
# the probability of the algorithm finding the correct answer.
# </details>
# <br/>
# <details closed>
# <summary><b>Hint #3</b></summary>
# You can use the Message function to output the results.
# </details>
operation Run_GroversSearch_Algorithm () : Unit {
// ...
}
%simulate Run_GroversSearch_Algorithm
| GroversAlgorithm/GroversAlgorithm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Machine Learning with Collaborative Filtering
# ## Contents
# 1. **Loading** movie dataset
# 2. Creating your movie **profile**
# 3. **Predicting** ratings
# 4. Finding **recommendations**
# In this challenge exercise, we will implement user-item collaborative filtering.
# We will first make a prediction for the rating you would give to a certain movie. Then we will generate recommendations, based on the movies that people similar to you liked.
#
# You will need two data files, which can be found in the `moviedata` folder. The data is separated with tabs.
# * `ratings.data`, which contains the ratings of 100,000 movies by many users (4 fields : user_id, movie_id, rating, timestamp).
# * `movies.data`, a mapping between movie ID's and titles.
# ## 1. Loading movie dataset
# First, we will load the `ratings.data` and `movies.data`. We will use the Python library [Pandas](http://pandas.pydata.org/) for this and load the data into a [DataFrame](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.html).
#
# Find the function in Pandas that is used to read csv data and use it to load the ratings and titles. Don't forget to specify the separator!
#
# (Hint: if you encounter encoding problems, set the encoding to 'cp1252'.)
# +
import pandas as pd
ratings = # TODO: load moviedata/ratings.data
movies = # TODO: load moviedata/movies.data
# -
# We can look at our new dataframes with the `.head()` function. This gives us the first five rows of the dataframe (unless you put a different number between the brackets).
movies.head()
ratings.head()
# In our ratings dataframe, we don't really need the timestamp column. Remove this column from the dataframe.
ratings = # TODO: remove timestamp column
# Now we will try to combine the ratings with the movie titles. We will write a function that can turn a column with movie id's into a column with movie names. By turning this into a function, we can easily apply it to other dataframes later.
def add_titles(df, movie_titles_df):
df = # merge dataframes
df = # remove movie_id column
return df
ratings = add_titles(ratings, movies)
ratings.head()
# You should not remove the movie id column without thinking though. This id could separate two movies with the same title (and year) that are not the same. However, in this case, this is not necessary so we can safely remove it.
# ## 2. Creating your own movie profile
# Now we have a dataframe that describes how a lot of users rate movies. The idea of collaborative filtering is to find a users that are similar to you and look at the movies they like or don't like. Your own ratings can be predicted based on the ratings your "neighbours" give these movies.
#
# So before we can find your neighbours, you need to rate a couple of movies. Let's create a movie profile for you.
# ### 2.1. Getting the most famous movies
# The first step will be to filter our movies. To make sure you know the movies you're about to rate, we will find the 100 most reviewed movies.
#
# Follow these steps:
# - Group the ratings dataframe by title
# - On these grouped items, apply the count function, this will give you a dataframe
# - Select the rating column. (Hint: `df["colname"]`)
# - Sort the values descending, so the most reviewed titles appear on top.
# - Take the first 100 rows. (Hint: use `head()`)
#
# You can do this all in one long chain and assign it to a new dataframe.
most_famous_movies = # TODO: group, count, select, sort and take first 100
# Have a look at the result with `head()`. The most reviewed movie should have 583 reviews. Can you guess which one it is?
# +
# TODO: check result
# -
# You might also notice that this result is not displayed as a table anymore. That is because the result has only an index and one column, so pandas turned it into a [Series](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.html) instead. Luckily for us, series work almost exactly the same as dataframes.
# ### 2.2. Rating a few of the most famous movies
# Now onto step 2: reviewing these movies. We will write a loop that will present a movie title to you and asks you to rate it. You don't want to rate all 100 movies though, let's just rate five of them.
#
# First, we want a small function that allows you to enter a number. So write a function that:
# - Gets input from the user. (Hint: experiment with `input("Hello")`.)
# - Then check if the entered text is a number.
# - If the text is a number between 1 and 5: return the number
# - Else: return None (python's null value)
def get_int_input(descr):
# TODO: write function
# Now we write a loop that keeps showing you movie titles until you have rated 5 of them.
#
# Here are the steps:
# - Create a new, empty dataframe with columns "title" and "rating".
# - Shuffle the most famous movies. (Hint: check pandas' `sample` function)
# - Ask rating for every movie.
# - Append result to the new dataframe if it is not None. (Hint: it's easy to append a dict to a df if you set `ignore_index` to True)
# - If we have enough ratings: break the loop.
#
# When you are done you can run the cell and rate five movies. You can skip the ones you don't know by pressing enter, or anything other than a number.
# +
user_ratings = # TODO: create new dataframe
ratings_needed = 5
shuffled_movies = # TODO: shuffle most_famous_movies
for title, row in shuffled_movies.iteritems():
rating = get_int_input("What rating (1-5) would you give to the movie " + title + "? ")
# TODO: append rating to user_ratings if it is not None
if len(user_ratings) >= ratings_needed:
break
# -
# Now you can view your profile:
user_ratings
# ## 3. Predicting ratings
# Based on these ratings, we will predict your opinion about other movies. The general approach to find your rating for a certain movie, is to look at all the users that have rated this movie and then take the average of their ratings for this movie. We multiply their rating by a weight though, based on how similar the users's ratings are to your own.
#
# Now let's do that step by step.
# We begin with the most basic function: getting a movie title based on an id. This will make it easier for us to test our code, because we won't need to type the entire movie title every time.
#
# Write a function that uses the `movies` dataframe to return the title of a movie id. You'll need some pandas for this.
def get_movie_title(movie_id):
# TODO: complete function
# Now we can use the function to select a movie:
selected_movie = get_movie_title(1)
selected_movie
# We will have a look at your predicted rating for this movie. If you rated this one, you can change the movie id.
# ### 3.1. Normalising ratings
# The next important part is to normalise your ratings. Basically you divide them all by the average. We do this because, if you always gave movies five stars, that says less about your taste than if you rated some higher and some lower. It also centers your ratings around zero. A negative rating means you enjoyed it less than usual, a positive rating means you liked it more than usual.
#
# From your user_ratings dataframe, calculate the average of your rating column. Store this as a variable, we will need it later.
user_avg_rating = # TODO: calculate your rating average
print(user_avg_rating)
# ### 3.2. Finding voters
# Now we will have a look at other people who rated our movie. We will refer to them as voters.
#
# Filter the ratings dataframe by our selected movie title and then get the id's of those users.
voters = # TODO: find voters id's in rating df
print("Number of voters who rated {}: {}".format(selected_movie, len(voters)))
# If you didn't change the selected movie, there should be 452 voters.
# ### 3.3. Comparing one voter to you
# We will focus on one voter first and compare him to you. After that, we will generalise it for all voters. We will select the first voter for now. We will assign his `user_id` to a variable so we can use it later.
voter = voters.iloc[0] # iloc[0] gets the element at index 0
voter
# Now get all the voter's ratings from the ratings dataframe.
voter_ratings = # TODO: get voter's ratings
voter_ratings.head()
# We would like to know how similar the voter is to you. If you have rated no movies in common, there is no point in considering him. So let's compare your ratings to his.
#
# Combine your ratings with pandas' `merge` function (or alternatives).
mutual_ratings = # TODO: merge voter_ratings and user_ratings
mutual_ratings
# Do you and the voter have the same taste, or in other words: do you have many movies in common? We would like to put a number to that question. So let's calculate the correlation coëfficient of your ratings. This says how similar two arrays are and is a function that can be found in numpy. Google a bit until you find the right function. This function gives you a matrix of multiple values as result, you need the element at `[0,1]`.
#
# Hint: Get your ratings from the mutual_ratings dataframe and convert them to a list.
# +
import numpy as np
corr = # TODO: calculate correlation between your ratings and voter's ratings
corr
# -
# If this number is NaN (not a number), then you have no movies in common with this voter or there is not enough distinction between your or his ratings. Just go back, pick another voter and run all the cells up to this point again. We will deal with these cases later.
# Last but not least, we need the voter's rating for our movie and his average rating. Calculate those two and store them in variables.
# +
voter_movie_rating = # TODO: find voter's rating for selected_movie
print(voter_movie_rating)
voter_avg_rating = # TODO: calculate voter's average rating
print(voter_avg_rating)
# -
# We're almost done with this step. Now comes the magic math: we multiply the voter's normalised rating with his correlation with you. This we divide by the absolute correlation. Then we add your average rating to un-normalise it for you and there is your rating, based on one single voter.
correlated_rating = corr * (voter_movie_rating - voter_avg_rating)
user_movie_rating = user_avg_rating + (correlated_rating / abs(corr))
user_movie_rating
# ### 3.4. Comparing all voters to you
# We don't want to base our rating on one single voter though. We will edit the code a bit to base it on all users that have rated the movie.
#
# We will:
# - Loop over all the voters.
# - Calculate the same things we did before for one voter.
# - Ignore voters with a correlation of NaN.
# - Sum the absolute correlation and the correlated rating for all voters and then divide them in the end.
# - Turn this all into a function that we can use again later
def predicted_rating(user_ratings, ratings, selected_movie):
abs_corr_sum = 0
corr_rating_sum = 0
voters = # TODO: find voters id's in rating df
for voter in voters:
# get voter's movies
voter_ratings = # TODO: get voter's ratings
# check similarity to user not nan
mutual_ratings = # TODO: merge voter_ratings and user_ratings
corr = # TODO: calculate correlation between your ratings and voter's ratings
if not np.isnan(corr):
# get selected movie rating
voter_movie_rating = # TODO: find voter's rating for selected_movie
# get voter's average
voter_avg_rating = # TODO: calculate voter's average rating
# sum similarity
abs_corr_sum += abs(corr)
corr_rating_sum += corr * (voter_movie_rating - voter_avg_rating)
# calculate predicted rating
user_movie_rating = user_avg_rating + (corr_rating_sum / abs_corr_sum)
return user_movie_rating
# Now we can use our function to get the actual prediction for the movie:
print("Predicted rating for {}: {}".format(selected_movie, round(predicted_rating(user_ratings, ratings, selected_movie))))
# If you get a lot of warnings (these come from the correlation function), you can run this cell to turn all warnings off.
import warnings
warnings.filterwarnings('ignore')
# You are now ready to make predictions! Try it for different movies, and see if it accurately reflects your tastes. If it does, you can use it to decide which movie to watch tonight! (Bear in mind though that this rating is only based on five ratings of yours, so it might not always be accurate.)
movie = get_movie_title(2)
print("Predicted rating for {}: {}".format(movie, round(predicted_rating(user_ratings, ratings, movie))))
# ## 4. Finding recommendations
# Now we can predict ratings for a specific movie, but when you're wondering what to watch you won't know which movie to look for. We could calculate the predicted rating for every single movie, but there is another solution as well. We could find the voters that are most similar to you, your neighbours, and see which movies they liked most.
# ### 4.1. Finding your neighbours
# Calculate your correlation with every user in the ratings dataframe.
corr_list = []
for voter in ratings.user_id.unique():
voter_ratings = # TODO: get voter's ratings
mutual_ratings = # TODO: merge voter_ratings and user_ratings
corr = # TODO: calculate correlation between your ratings and voter's ratings
corr_list.append([voter, len(mutual_ratings), corr])
voter_corr = pd.DataFrame(corr_list, columns=["user_id", "movies_in_common", "corr"])
# Now we have a dataframe of our similarity with all users.
voter_corr.head()
# We want to find out which users are most similar to us. In this case, we want the ten users with the most movies in common and the highest correlation.
#
# Create a list of similar voters:
# - Sort the `voter_corr` dataframe, by `movies_in_common` and then by `corr`. Sort the values descending so the highest ones appear on top.
# - Take the first ten rows to get the most similar users.
# - Select the `user_id` column.
# - Cast this result to a list.
similar_voters = # TODO: create list of most similar user_id's
print(similar_voters)
# Those are the id's of your neighbours.
# ### 4.2. Finding your neighbours favourite movies
# Now we have a look at those users and the movies they gave high ratings.
#
# Follow these steps:
# - Start from the ratings dataframe.
# - Filter out user_id's that don't appear in `similar_voters`.
# - Filter out titles that appear in `user_ratings.title`.
# - Group by title.
# - Aggregate by both count and mean.
# - Select rating column.
# - Filter out movies with a mean rating of less than 4.
# - Sort the result, on count and then on mean, descending.
# - Take the top 10 rows.
recommendations = # TODO: filter voters and already seen movies from ratings df
recommendations = # TODO: group by title and aggregate
recommendations = # TODO: filter mean ratings less than 4
recommendations = # TODO: sort and take first 10
recommendations
# There you go, you have just built your own recomendation engine from scratch. Time to start making money!
| PXL_DIGITAL_JAAR_2/Data Advanced/TiX/Data Advanced/DATAMAARTEN/jupyter notebooks/Machine_Learning_3_CollabFiltering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
library(TCGAbiolinks)
library(stringr)
library(tidyverse)
library(readr)
# This notebook uses the [TCGAbiolinks](http://bioconductor.org/packages/release/bioc/html/TCGAbiolinks.html) R package to download TCGA data from GDC. We then combine MAF files with CNV data and add in information on the cellularity of samples so that we can calculate the mutation copy number (MCN) of each variant.
#get all TCGA cancer subtype codes
tcgacodes <- TCGAbiolinks:::getGDCprojects()$project_id
tcgacodes <- unlist(lapply(tcgacodes[str_detect(tcgacodes, "TCGA")],
function(x){strsplit(x, "-")[[1]][2]}))
# +
#download mutect MAF files for all samples and save to csv file
dfhg38 <- data.frame()
for (t in tcgacodes){
maf <- GDCquery_Maf(t, pipelines = "mutect") %>%
mutate(cancertype = t)
dfhg38 <- rbind(dfhg38, maf)
}
dfhg38 <- dfhg38 %>%
mutate(sampleid = str_sub(Tumor_Sample_Barcode, 1, 16)) #annotate so barcode is consistent with TCGA CNV id
write_delim(dfhg38, "data/TCGA-maf-all-hg38.csv", delim = ",")
#dfhg38 <- read_csv("data/TCGA-maf-all-hg38.csv")
# -
dfhg38 <- read_csv("data/TCGA-maf-all-hg38.csv") %>%
dplyr::mutate(VAF = t_alt_count/t_depth) %>%
dplyr::rename(chr = Chromosome, start = Start_Position, end = End_Position) %>%
dplyr::select(sampleid, chr, start, end, Reference_Allele, Tumor_Seq_Allele2, VAF,
t_depth, t_ref_count, t_alt_count, n_depth,
n_ref_count, n_alt_count, cancertype) %>%
dplyr::mutate(nref = str_length(Reference_Allele), nalt = str_length(Tumor_Seq_Allele2)) %>%
dplyr::mutate(mutation_type = ifelse((nref - nalt) == 0, "SNV", "INS/DEL")) %>%
dplyr::select(-nref, -nalt)
dfhg38 %>%
distinct(sampleid, cancertype) %>%
group_by(cancertype) %>%
summarise(n = n())
# ### Download copy number data
# +
### Download copy number data
### For some reason this takes much longer than the SNV files
tcgacodes <- TCGAbiolinks:::getGDCprojects()$project_id
query <- GDCquery(project = tcgacodes[str_detect(tcgacodes, "TCGA")],
data.category = "Copy Number Variation",
data.type = "Copy Number Segment")
cnvsamples <- getResults(query)
GDCdownload(query)
dfhg38cnv <- GDCprepare(query)
write_delim(dfhg38cnv, "data/TCGA-cnv-all-hg38.csv", delim = ",")
#dfhg38cnv <- read.csv("data/TCGA-cnv-all-hg38.csv")
# -
dfhg38cnv <- read_csv("data/TCGA-cnv-all-hg38.csv") %>%
dplyr::mutate(sampleid = str_sub(Sample, 1, 16)) %>%
dplyr::select(-Sample, -X1) %>%
filter(sampleid %in% dfhg38$sampleid)
head(dfhg38)
head(dfhg38cnv)
# ### Read in cellularity
# +
### Import cellularity estimates
cellularity <- read.delim("data/ascat_acf_ploidy.tsv", header = T) %>%
mutate(sampleid = str_sub(gsub("[.]", "-", Sample), 1, 16)) %>%
select(-Sample) %>%
dplyr::rename(ploidy = Ploidy, cellularity = Aberrant_Cell_Fraction.Purity.)
head(cellularity)
# -
# ### Combine all data types
# +
#filter for useful columns and add new ones to snv df
dfsnv <- dfhg38 %>%
dplyr::mutate(VAF = t_alt_count/t_depth) %>%
dplyr::rename(chr = Chromosome, start = Start_Position, end = End_Position) %>%
dplyr::select(sampleid, chr, start, end, Reference_Allele, Tumor_Seq_Allele2, VAF,
t_depth, t_ref_count, t_alt_count, n_depth,
n_ref_count, n_alt_count, cancertype) %>%
dplyr::mutate(nref = str_length(Reference_Allele), nalt = str_length(Tumor_Seq_Allele2)) %>%
dplyr::mutate(mutation_type = ifelse((nref - nalt) == 0, "SNV", "INS/DEL")) %>%
dplyr::select(-nref, -nalt)
# -
#combine snv with cellularity
dfsnv <- dfhg38
df1temp <- left_join(dfsnv, cellularity, by = c("sampleid")) %>%
filter(cellularity > 0.2) #remove cellularity < 20%
#format CNV for easy joining
dfhg38cnvt <- dfhg38cnv %>%
dplyr::rename(chr = Chromosome) %>%#, start = Start, end = End) %>%
dplyr::mutate(chr = paste0("chr", chr)) %>%
dplyr::filter(sampleid %in% unique(df1temp$sampleid))
length(unique(dfhg38cnvt$sampleid))
#join snv, cnv and cellularity
df2temp <- inner_join(df1temp, dfhg38cnvt, by = c("sampleid", "chr")) %>%
filter(start >= Start & end <= End) %>%
select(-Start, -End)
length(unique(df2temp$sampleid))
# +
#calculate copy number and mutation copy number MCN
dfcombinedhg38 <- df2temp %>%
select(-n_ref_count, -n_alt_count, -Num_Probes)
dfout <- dfcombinedhg38 %>%
mutate(cellularity = ifelse(is.na(cellularity), 1, as.numeric(cellularity))) %>%
#calculate CN by correcting for cellularit
mutate(CN = 2^Segment_Mean * 2, CNcorrected = (2^Segment_Mean + cellularity - 1) * (2 / cellularity),
absCN = round(CN), absCNcorrected = round(CNcorrected)) %>%
#don't allow CN == 0
mutate(absCN = ifelse(absCN == 0, 1, absCN), absCNcorrected = ifelse(absCNcorrected == 0, 1, absCNcorrected)) %>%
mutate(MCN = ((CNcorrected - 2) * 1 + 2) * VAF/cellularity)
head(dfout)
# -
#write hg38 file out
write_delim(dfout, "data/TCGA-combined-hg38-2.csv", delim = ",")
# ### Download clinical info
# +
# The below will also download clinical information, although we do not use this
tcgacodes <- TCGAbiolinks:::getGDCprojects()$project_id
tcgacodes <- tcgacodes[str_detect(tcgacodes, "TCGA")]
dfclinical <- data.frame()
for (p in tcgacodes[1:length(tcgacodes)-1]){
print(p)
query <- GDCquery(project = p,
data.category = "Clinical")
GDCdownload(query)
clinical <- GDCprepare_clinic(query, clinical.info = "patient", directory = "GDCdata/")
names(clinical)
if("stage_event_system_version" %in% names(clinical)){
dfclinical <- bind_rows(dfclinical, select(clinical, -stage_event_system_version))
} else if("patient_id" %in% names(clinical)){
dfclinical <- bind_rows(dfclinical, select(clinical, -patient_id))
}
else{
dfclinical <- bind_rows(dfclinical, clinical)
}
}
dfout <- select(dfclinical, bcr_patient_barcode, tumor_tissue_site, histological_type,
vital_status, days_to_birth, days_to_last_known_alive, days_to_death, days_to_last_followup,
stage_event_clinical_stage, stage_event_pathologic_stage, stage_event_tnm_categories,
stage_event_gleason_grading)
write.csv(dfout, "data/TCGA-clinical.csv", row.names = F)
| notebooks/Notebook 4 - Download TCGA Data and Combine MAF with CNV (R).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
# We go over the basic functionality of the code by initializing a detector and creating some time-ordered data.
# We start by importing the `ScanStrategy`, `Instrument` and `Beam` classes from `beamconv`. We we will also need some functions from `healpy`.
# +
import matplotlib
import matplotlib.pyplot as plt
fontsize=14
matplotlib.rcParams.update({'font.size':fontsize})
matplotlib.rcParams.update({'xtick.labelsize':fontsize})
matplotlib.rcParams.update({'ytick.labelsize':fontsize})
matplotlib.rcParams.update({'axes.labelsize':fontsize})
matplotlib.rcParams.update({'lines.markersize': 4})
import numpy as np
import healpy as hp
from beamconv import ScanStrategy
from beamconv import Instrument
from beamconv import Beam
from beamconv import tools
# -
# For simplicity we will only use a single detector in this example. Detectors are represented by instances of the `Beam` class we just imported. A `Beam` instance contains detector properties such as pointing offsets, but also pointers to the harmonic modes of the actual beam.
#
# First lets see what options we are allowed to set set for our `Beam` instance:
# ?Beam
# Let's create a detector without pointing offset (i.e. with beam centre on the telescope's boresight). For now we simply take a Gaussian beam model. We can thus fully specify the beam with the `fwhm` (Full Width at Half Maximum) parameter. Here we pick `fwhm=40`, in units of arcmin. Since the 2D Gaussian is azimuthally symmetric, we may set the `symmetric` option.
beam_opts = dict(az=0,
el=0,
polang=0,
fwhm=40,
btype='Gaussian',
lmax=800,
symmetric=True)
beam = Beam(**beam_opts)
print(beam)
# In order to start simulating data we need to initialize a `ScanStrategy` instance that manages the simulated data-taking. `ScanStrategy` inherents from the `Instrument` class, so by initializing it we also initialize an `Instrument` instance. Let's see what both classes need as input:
# ?Instrument
# ?ScanStrategy
# For simplicity we pick a preset location for the instrument (the south pole). For the scan strategy we mostly leave the defaults but specify that we would like to simulate one hour of data at a 100 Hz sample rate.
# +
instr_opts = dict(location='spole')
scan_opts = dict(duration=3600,
sample_rate=100)
scan_opts.update(instr_opts)
S = ScanStrategy(**scan_opts)
# -
# Lets add our beam to the detectors focal plane. Each `Instrument` instance has a `beams` attribute: a list of detector pairs that represent the focal plane. Because we only have a single detector, our detector will have a empty slot as detector partner.
S.add_to_focal_plane(beam)
print(S.beams)
# We now specify the telescope's pointing (the boresight pointing). For this example we use a preset scan strategy: a constant-elevation scanning pattern. To do so, we use the `constant_el_scan()` method of our `ScanStrategy` instance. Lets see what input we need to provide:
# ?S.constant_el_scan
# We leave the scan centre set to its default value (`ra0`, `dec0` = -10, -57.5, which is always visible from the south pole) and set the angular speed and azimuthal width of the scans.
const_el_opts = dict(az_throw=50.,
scan_speed=10.)
# We also need to specify the input sky. We do this by providing the harmonic coefficients of the sky in a tuple containing unpolarized and E-, B-modes: ($a_{\ell m}$, $a_{E, \ell m}, a_{B, \ell m}$). Here we use the coefficients of a Gaussian sky with covariance given by some best-fit WMAP $\Lambda$CDM power spectrum.
cls = np.loadtxt('../ancillary/wmap7_r0p03_lensed_uK_ext.txt',
unpack=True) # Cl in uK^2
_, cls = cls[0], cls[1:]
np.random.seed(25)
alm = hp.synalm(cls, lmax=800, new=True, verbose=False) # uK
# The actual scanning of the sky can be done by using the `scan_instrument_mpi` method of `ScanStrategy`. Broadly speaking, this function first calls the `init_spinmaps` function that performs the inverse spherical harmonic transforms needed for the beam convolution, then it calls the `constant_el_scan` function we mentioned above to populate the boresight pointing quaternions. Finally, it calls the `scan` function that combines boresight pointing, detector pointing and the beam-convolved maps to calculate the final time-ordered data.
#
# Note that `scan_instrument_mpi` will also work with more than one detector on the focal plane, possibly distributing the workload over available MPI ranks.
#
# For this demonstration, we can separate the inverse spherical harmonic transforms and the scanning. To do so, we first call the `init_detpair` function. This will populate the beam-convolved `spinmaps`. If we then call `scan_instrument_mpi` with `reuse_spinmaps=True` it will use the precomputed `spinmaps`.
beam = S.beams[0][0]
S.init_detpair(alm, beam, beam_b=None, nside_spin=512)
# Then we do the actual scanning of the sky. We set the `save_tod` option to save the data we are about to generate.
S.scan_instrument_mpi(alm, binning=False, reuse_spinmaps=True, interp=False, save_tod=True, **const_el_opts)
# We can extract the data we just generated by calling the `data` method. Data sampling is generally done in periods of time that divide up the full mission in to managable "chunks" of data. In our case we have only used one chunk. The chunks are stokes in the `chunks` attribute.
print('No. of chunks used: {}'.format(len(S.chunks)))
chunk = S.chunks[0]
print(chunk)
tod = S.data(chunk, beam=beam, data_type='tod')
# Lets plot the first 500 data points:
fig, ax = plt.subplots()
ax.plot(tod[:500])
ax.set_xlabel('Time-sample')
ax.set_ylabel(r'Signal [$\mu K_{\mathrm{CMB}}$]')
# We can rerun with the `interp` option set to use bi-linear interpolation while sampling the beam-convolved maps.
S.scan_instrument_mpi(alm, binning=False, reuse_spinmaps=True, interp=True, save_tod=True, **const_el_opts)
tod_interp = S.data(S.chunks[0], beam=beam, data_type='tod')
fig, ax = plt.subplots()
ax.plot(tod[:50], label='raw')
ax.plot(tod_interp[:50], label='interp')
ax.set_xlabel('Time-sample')
ax.set_ylabel(r'Signal [$\mu K_{\mathrm{CMB}}$]')
ax.legend()
# # Temperature-to-Polarization leakage
# With the basic structure of the code set, we take a look at a leading-order optical systematic: temperature-to-polarization leakage due to azimuthally asymmetric beams.
# In this example we use precomputed harmonic modes for a beam from a simple 2-lens refracting telescope. As this beam is not azimuthally symmetric, we have to specify the number of azimuthal modes we consider. As we will see, the beam is only weakly asymmetric so we only use the first 5 modes by setting `mmax = 5`. The equivalent Gaussian beamwidth is roughly 38 arcmin.
beam_dir = '../tests/test_data/example_blms/'
beam_opts = dict(az=0,
el=0,
polang=0,
fwhm=38,
btype='PO', # Physical optics.
lmax=800,
mmax=5,
po_file=beam_dir+'blm_hp_X1T1R1C8A_800_800.npy')
beam2 = Beam(**beam_opts)
# Lets take a look at the beam and compare it to the Gaussian approximation. First, lets plot the two intensity beams:
# +
def blm2bmap(beam, nside=1024):
'''
Compute pixelized map of the intensity beam of
Beam instance.
Arguments
---------
beam : <beamconv.detector.Beam>
Keyword arguments
-----------------
nside : int
Nside of output maps
Returns
-------
beammaps : array-like
HEALPix map of intensity beam.
'''
blm = np.asarray(beam.blm).copy()[0]
# We need to divide out sqrt(4pi / (2 ell + 1)) to get
# correctly normlized spherical harmonic coeffients.
ell = np.arange(hp.Alm.getlmax(blm.size))
q_ell = np.sqrt(4. * np.pi / (2 * ell + 1))
blm = hp.almxfl(blm, 1 / q_ell)
return hp.alm2map(blm, nside, verbose=False)
beam2.btype='PO'
bmap_po = blm2bmap(beam2)
del(beam2.blm)
beam2.btype='Gaussian'
bmap_ga = blm2bmap(beam2)
del(beam2.blm)
cart_opts = dict(rot=[0, 90, 0], lonra=[-3,3], latra=[-3,3], min=0, max=9,
hold=True, unit=r'$\log |b(\theta, \phi)|$)')
fig, axs = plt.subplots(ncols=2, nrows=1, figsize=(10, 14))
plt.axes(axs[0])
hp.cartview(np.log(np.abs(bmap_po)), title="Physical optics", **cart_opts)
plt.axes(axs[1])
hp.cartview(np.log(np.abs(bmap_ga)), title="Gaussian", **cart_opts)
# -
# It is also instructive to compare the harmonic modes. First we take a look at the $m=0$ modes (the azimuthally symmetric components). We see that the physical optics beams are slightly non-Gaussian. Note that we plot spherical harmonic modes that are normalized at the monopole by defining $q_{\ell} = \sqrt{\frac{4 \pi}{(2\ell + 1)}}$.
# +
beam2.btype='PO'
bell_po = np.real(tools.blm2bl(beam2.blm[0]))
del(beam2.blm)
beam2.btype='Gaussian'
bell_ga = np.real(tools.blm2bl(beam2.blm[0]))
del(beam2.blm)
fig, axs = plt.subplots(ncols=1, nrows=2, figsize=(7, 5), sharex=True)
axs[0].plot(bell_po, label='Physical optics')
axs[0].plot(bell_ga, label='Gaussian')
axs[1].plot(bell_po / bell_ga, label='Ratio (PO / Gauss.)')
axs[0].set_ylabel(r'$q_{\ell} b_{\ell 0}$')
axs[1].set_ylabel(r'$b^{\mathrm{PO}}_{\ell 0} / b^{\mathrm{G}}_{\ell 0}$')
axs[1].set_xlabel(r'Multipole [$\ell$]')
axs[0].legend()
axs[1].legend()
fig.tight_layout()
# -
# Now we take a look at the azimuthally asymmetric modes of the physical optics beam. It is clear that most information is stored in the low $m$ modes and that even the largest asymetric components have only few percent of the symmetric component's amplitude. Note that the $m\neq0$ modes are complex numbers so we plot the real and imaginary part.
beam2.btype='PO'
blm = beam2.blm[0]
del(beam2.blm)
fig, axs = plt.subplots(ncols=2, figsize=(12, 5), sharey=True, sharex=True)
for m in range(1,7):
bell = tools.blm2bl(blm, m=m)
axs[0].plot(np.real(bell), label='m={}'.format(m))
axs[1].plot(np.imag(bell), label='m={}'.format(m))
axs[1].legend(ncol=2)
axs[0].set_title(r'$\Re \ b_{\ell m}$')
axs[1].set_title(r'$\Im \ b_{\ell m}$')
axs[0].set_ylabel(r'$q_{\ell} b^{\mathrm{PO}}_{\ell m}$')
axs[0].set_xlabel(r'Multipole [$\ell$]')
axs[1].set_xlabel(r'Multipole [$\ell$]')
fig.tight_layout()
# We initiate a new `ScanStrategy` instance and add our new beam.
S2 = ScanStrategy(**scan_opts)
S2.add_to_focal_plane(beam2)
# This time we let the instrument rotate 36 degrees around the boresight every 100 seconds in addition the to constant-elevation scanning
S2.set_instr_rot(period=100, angles=np.linspace(0, 360, 10))
# We turn off the linearly polarized component of the sky. This means that any sign of a polarized signal in the data will be due to systematic effects.
alm = np.asarray(alm)
alm[1] *= 0
alm[2] *= 0
# Let us start by scanning the sky with the Gaussian beam. This time we also save the indices of the pixels visited by the scan strategy and the position angle `pa` (or $\psi$) at each sample.
S2.set_btypes('Gaussian')
beam2.symmetric = True
S2.reset_instr_rot()
S2.scan_instrument_mpi(alm, binning=False, reuse_spinmaps=False, interp=False,
save_point=True, save_tod=True, nside_spin=512, **const_el_opts)
pix_ga = S2.data(S2.chunks[0], beam=beam2, data_type='pix')
pa_ga = S2.data(S2.chunks[0], beam=beam2, data_type='pa')
tod_ga = S2.data(S2.chunks[0], beam=beam2, data_type='tod')
# Lets now plot the data gathered from a single pixel on the sky. We pick the pixels that is most visited by the scanning. If we now plot the signal gathered from this pixel, we expect no variation with position angle. This is because the Gaussian beam is azimuthally symmetric. So despite the rotation of the boresight every 100 seconds, the signal should remain constant.
common_pix_ga = np.bincount(pix_ga).argmax()
fig, ax = plt.subplots()
ax.scatter(pa_ga[pix_ga==common_pix_ga], tod_ga[pix_ga==common_pix_ga])
ax.set_xlim(-20, 360)
_= ax.set_xticks([0, 90, 180, 270, 360])
_= ax.set_xticklabels([0, 90, 180, 270, 360])
ax.set_xlabel(r"Position angle $\psi$ [$\deg$]")
_= ax.set_ylabel(r"Signal [$\mu K_{\mathrm{CMB}}$]")
# Lets now repeat the scanning, but with the physical optics beam. This time, the (internal) call to `init_detpair` will take longer as the code also has to perform inverse spherical harmonic transforms for all of the $m \neq 0$ modes of the intensity beam (and all the $m \neq 2$ modes of the linearly polarized beam).
S2.set_btypes('PO')
beam2.symmetric = False
S2.reset_instr_rot()
S2.scan_instrument_mpi(alm, binning=False, reuse_spinmaps=False, interp=False,
save_point=True, save_tod=True, nside_spin=512, **const_el_opts)
# Lets plot the signal in the most visited pixel as function of position angle again. This time we see a variation with position angle. Remember that there is no linearly polarized component to the input sky in this simulation. All the variation is thus because of the asymmetry of the beam.
# +
pix_po = S2.data(S2.chunks[0], beam=beam2, data_type='pix')
pa_po = S2.data(S2.chunks[0], beam=beam2, data_type='pa')
tod_po = S2.data(S2.chunks[0], beam=beam2, data_type='tod')
common_pix_po = np.bincount(pix_po).argmax()
fig, ax = plt.subplots()
ax.scatter(pa_po[pix_po==common_pix_po], tod_po[pix_po==common_pix_po])
ax.set_xlim(-20, 360)
_= ax.set_xticks([0, 90, 180, 270, 360])
_= ax.set_xticklabels([0, 90, 180, 270, 360])
ax.set_xlabel(r"Position angle $\psi$ [$\deg$]")
_= ax.set_ylabel(r"Signal [$\mu K_{\mathrm{CMB}}$]")
# -
# The above plot demonstrates the systematic temperature-to-polarization leakage we care about in this example. The point is that a linearly polarized sky signal will produce a sinusoidal signature with period $2 \psi$ in a plot like the above. We see that the signal in this pixel shows this behaviour (plus some other harmonics). A map-making algorithm that uses the time-ordered data and angular information to reconstruct the linearly polarized sky signal will therefore wronly interpret the above as polarized signal.
| notebooks/introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Installing and Configuring Holoviews
# HoloViews can be installed on any platform where [NumPy](http://numpy.org) and Python 2.7 or 3 are available.
#
# That said, HoloViews is designed to work closely with many other libraries, which can make installation and configuration more complicated. This user guide page describes some of these less-common or not-required options that may be helpful for some users.
# ## Other installation options
#
# The main [installation instructions](http://holoviews.org/#installation) should be sufficient for most users, but you may also want the [Matplotlib](http://matplotlib.org) and [Plotly](https://plot.ly/python/) backends, which are required for some of the examples:
#
# conda install matplotlib plotly
#
# HoloViews can also be installed using one of these `pip` commands:
#
# pip install holoviews
# pip install 'holoviews[recommended]'
# pip install 'holoviews[extras]'
# pip install 'holoviews[all]'
#
# The first option installs just the bare library and the [NumPy](http://numpy.org) and [Param](https://github.com/holoviz/param) libraries, which is all you need on your system to generate and work with HoloViews objects without visualizing them. The other options install additional libraries that are often useful, with the `recommended` option being similar to the `conda` install command above.
#
# Between releases, development snapshots are made available as conda packages:
#
# conda install -c pyviz/label/dev holoviews
#
# To get the very latest development version you can clone our git
# repository and put it on the Python path:
#
# git clone https://github.com/holoviz/holoviews.git
# cd holoviews
# pip install -e .
# ## JupyterLab configuration
#
# To work with JupyterLab you will also need the HoloViews JupyterLab
# extension:
#
# ```
# conda install -c conda-forge jupyterlab
# jupyter labextension install @pyviz/jupyterlab_pyviz
# ```
#
# Once you have installed JupyterLab and the extension launch it with:
#
# ```
# jupyter-lab
# ```
# ## ``hv.config`` settings
#
# The default HoloViews installation will use the latest defaults and options available, which is appropriate for new users. If you want to work with code written for older HoloViews versions, you can use the top-level ``hv.config`` object to control various backwards-compatibility options:
#
# * ``future_deprecations``: Enables warnings about future deprecations (introduced in 1.11).
# * ``warn_options_call``: Warn when using the to-be-deprecated ``__call__`` syntax for specifying options, instead of the recommended ``.opts`` method.
#
# It is recommended you set ``warn_options_call`` to ``True`` in your holoviews.rc file (see section below).
#
# It is possible to set the configuration using `hv.config` directly:
import holoviews as hv
hv.config(future_deprecations=True)
# However, because in some cases this configuration needs to be declared before the plotting extensions are imported, the recommended way of setting configuration options is:
hv.extension('bokeh', config=dict(future_deprecations=True))
# In addition to backwards-compatibility options, ``hv.config`` holds some global options:
#
# * ``image_rtol``: The tolerance used to enforce regular sampling for regular, gridded data. Used to validate ``Image`` data.
#
# This option allows you to set the ``rtol`` parameter of [``Image``](../reference/elements/bokeh/Image.ipynb) elements globally.
#
# ## Improved tab-completion
#
# Both ``Layout`` and ``Overlay`` are designed around convenient tab-completion, with the expectation of upper-case names being listed first. In recent versions of Jupyter/IPython there has been a regression whereby the tab-completion is no longer case-sensitive. This can be fixed with:
import holoviews as hv
hv.extension(case_sensitive_completion=True)
# ## The holoviews.rc file
#
# HoloViews searches for the first rc file it finds in the following places (in order):
#
# 1. ``holoviews.rc`` in the parent directory of the top-level ``__init__.py`` file (useful for developers working out of the HoloViews git repo)
# 2. ``~/.holoviews.rc``
# 3. ``~/.config/holoviews/holoviews.rc``
#
# The rc file location can be overridden via the ``HOLOVIEWSRC`` environment variable.
#
# The rc file is a Python script, executed as HoloViews is imported. An example rc file to include various options discussed above might look like this:
#
# ```
# import holoviews as hv
# hv.config(warn_options_call=True)
# hv.extension.case_sensitive_completion=True
# ```
#
| examples/user_guide/Installing_and_Configuring.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="W7rEsKyWcxmu"
# ##### Copyright 2021 The TF-Agents Authors.
#
# + cellView="form" id="nQnmcm0oI1Q-"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="G6aOV15Wc4HP"
# # CheckpointerとPolicySaver
#
# <table class="tfo-notebook-buttons" align="left">
# <td><a target="_blank" href="https://www.tensorflow.org/agents/tutorials/10_checkpointer_policysaver_tutorial"> <img src="https://www.tensorflow.org/images/tf_logo_32px.png"> TensorFlow.org で表示</a></td>
# <td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ja/agents/tutorials/10_checkpointer_policysaver_tutorial.ipynb"> <img src="https://www.tensorflow.org/images/colab_logo_32px.png"> Google Colab で実行</a></td>
# <td><a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ja/agents/tutorials/10_checkpointer_policysaver_tutorial.ipynb"> <img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png"> GitHub でソースを表示</a></td>
# <td><a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ja/agents/tutorials/10_checkpointer_policysaver_tutorial.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png">ノートブックをダウンロード</a></td>
# </table>
# + [markdown] id="M3HE5S3wsMEh"
# ## はじめに
#
# `tf_agents.utils.common.Checkpointer`は、ローカルストレージとの間でトレーニングの状態、ポリシーの状態、およびreplay_bufferの状態を保存/読み込むユーティリティです。
#
# `tf_agents.policies.policy_saver.PolicySaver`は、ポリシーのみを保存/読み込むツールであり、`Checkpointer`よりも軽量です。`PolicySaver`を使用すると、ポリシーを作成したコードに関する知識がなくてもモデルをデプロイできます。
#
# このチュートリアルでは、DQNを使用してモデルをトレーニングし、次に`Checkpointer`と`PolicySaver`を使用して、状態とモデルをインタラクティブな方法で保存および読み込む方法を紹介します。`PolicySaver`では、TF2.0の新しいsaved_modelツールとフォーマットを使用することに注意してください。
#
# + [markdown] id="vbTrDrX4dkP_"
# ## セットアップ
# + [markdown] id="Opk_cVDYdgct"
# 以下の依存関係をインストールしていない場合は、実行します。
# + id="Jv668dKvZmka"
#@test {"skip": true}
# !sudo apt-get update
# !sudo apt-get install -y xvfb ffmpeg python-opengl
# !pip install pyglet
# !pip install 'imageio==2.4.0'
# !pip install 'xvfbwrapper==0.2.9'
# !pip install tf-agents
# + id="bQMULMo1dCEn"
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import base64
import imageio
import io
import matplotlib
import matplotlib.pyplot as plt
import os
import shutil
import tempfile
import tensorflow as tf
import zipfile
import IPython
try:
from google.colab import files
except ImportError:
files = None
from tf_agents.agents.dqn import dqn_agent
from tf_agents.drivers import dynamic_step_driver
from tf_agents.environments import suite_gym
from tf_agents.environments import tf_py_environment
from tf_agents.eval import metric_utils
from tf_agents.metrics import tf_metrics
from tf_agents.networks import q_network
from tf_agents.policies import policy_saver
from tf_agents.policies import py_tf_eager_policy
from tf_agents.policies import random_tf_policy
from tf_agents.replay_buffers import tf_uniform_replay_buffer
from tf_agents.trajectories import trajectory
from tf_agents.utils import common
tempdir = os.getenv("TEST_TMPDIR", tempfile.gettempdir())
# + id="AwIqiLdDCX9Q"
#@test {"skip": true}
# Set up a virtual display for rendering OpenAI gym environments.
import xvfbwrapper
xvfbwrapper.Xvfb(1400, 900, 24).start()
# + [markdown] id="AOv_kofIvWnW"
# ## DQNエージェント
#
# 前のColabと同じように、DQNエージェントを設定します。 このColabでは、詳細は主な部分ではないので、デフォルトでは非表示になっていますが、「コードを表示」をクリックすると詳細を表示できます。
# + [markdown] id="cStmaxredFSW"
# ### ハイパーパラメーター
# + cellView="both" id="yxFs6QU0dGI_"
env_name = "CartPole-v1"
collect_steps_per_iteration = 100
replay_buffer_capacity = 100000
fc_layer_params = (100,)
batch_size = 64
learning_rate = 1e-3
log_interval = 5
num_eval_episodes = 10
eval_interval = 1000
# + [markdown] id="w4GR7RDndIOR"
# ### 環境
# + id="fZwK4d-bdI7Z"
train_py_env = suite_gym.load(env_name)
eval_py_env = suite_gym.load(env_name)
train_env = tf_py_environment.TFPyEnvironment(train_py_env)
eval_env = tf_py_environment.TFPyEnvironment(eval_py_env)
# + [markdown] id="0AvYRwfkeMvo"
# ### エージェント
# + id="cUrFl83ieOvV"
#@title
q_net = q_network.QNetwork(
train_env.observation_spec(),
train_env.action_spec(),
fc_layer_params=fc_layer_params)
optimizer = tf.compat.v1.train.AdamOptimizer(learning_rate=learning_rate)
global_step = tf.compat.v1.train.get_or_create_global_step()
agent = dqn_agent.DqnAgent(
train_env.time_step_spec(),
train_env.action_spec(),
q_network=q_net,
optimizer=optimizer,
td_errors_loss_fn=common.element_wise_squared_loss,
train_step_counter=global_step)
agent.initialize()
# + [markdown] id="p8ganoJhdsbn"
# ### データ収集
# + id="XiT1p78HdtSe"
#@title
replay_buffer = tf_uniform_replay_buffer.TFUniformReplayBuffer(
data_spec=agent.collect_data_spec,
batch_size=train_env.batch_size,
max_length=replay_buffer_capacity)
collect_driver = dynamic_step_driver.DynamicStepDriver(
train_env,
agent.collect_policy,
observers=[replay_buffer.add_batch],
num_steps=collect_steps_per_iteration)
# Initial data collection
collect_driver.run()
# Dataset generates trajectories with shape [BxTx...] where
# T = n_step_update + 1.
dataset = replay_buffer.as_dataset(
num_parallel_calls=3, sample_batch_size=batch_size,
num_steps=2).prefetch(3)
iterator = iter(dataset)
# + [markdown] id="8V8bojrKdupW"
# ### エージェントのトレーニング
# + id="-rDC3leXdvm_"
#@title
# (Optional) Optimize by wrapping some of the code in a graph using TF function.
agent.train = common.function(agent.train)
def train_one_iteration():
# Collect a few steps using collect_policy and save to the replay buffer.
collect_driver.run()
# Sample a batch of data from the buffer and update the agent's network.
experience, unused_info = next(iterator)
train_loss = agent.train(experience)
iteration = agent.train_step_counter.numpy()
print ('iteration: {0} loss: {1}'.format(iteration, train_loss.loss))
# + [markdown] id="vgqVaPnUeDAn"
# ### ビデオ生成
# + id="ZY6w-fcieFDW"
#@title
def embed_gif(gif_buffer):
"""Embeds a gif file in the notebook."""
tag = '<img src="data:image/gif;base64,{0}"/>'.format(base64.b64encode(gif_buffer).decode())
return IPython.display.HTML(tag)
def run_episodes_and_create_video(policy, eval_tf_env, eval_py_env):
num_episodes = 3
frames = []
for _ in range(num_episodes):
time_step = eval_tf_env.reset()
frames.append(eval_py_env.render())
while not time_step.is_last():
action_step = policy.action(time_step)
time_step = eval_tf_env.step(action_step.action)
frames.append(eval_py_env.render())
gif_file = io.BytesIO()
imageio.mimsave(gif_file, frames, format='gif', fps=60)
IPython.display.display(embed_gif(gif_file.getvalue()))
# + [markdown] id="y-oA8VYJdFdj"
# ### ビデオ生成
#
# ビデオを生成して、ポリシーのパフォーマンスを確認します。
# + id="FpmPLXWbdG70"
print ('global_step:')
print (global_step)
run_episodes_and_create_video(agent.policy, eval_env, eval_py_env)
# + [markdown] id="7RPLExsxwnOm"
# ## チェックポインタとPolicySaverのセットアップ
#
# CheckpointerとPolicySaverを使用する準備ができました。
# + [markdown] id="g-iyQJacfQqO"
# ### Checkpointer
#
# + id="2DzCJZ-6YYbX"
checkpoint_dir = os.path.join(tempdir, 'checkpoint')
train_checkpointer = common.Checkpointer(
ckpt_dir=checkpoint_dir,
max_to_keep=1,
agent=agent,
policy=agent.policy,
replay_buffer=replay_buffer,
global_step=global_step
)
# + [markdown] id="MKpWNZM4WE8d"
# ### Policy Saver
# + id="8mDZ_YMUWEY9"
policy_dir = os.path.join(tempdir, 'policy')
tf_policy_saver = policy_saver.PolicySaver(agent.policy)
# + [markdown] id="1OnANb1Idx8-"
# ### 1回のイテレーションのトレーニング
# + id="ql_D1iq8dl0X"
#@test {"skip": true}
print('Training one iteration....')
train_one_iteration()
# + [markdown] id="eSChNSQPlySb"
# ### チェックポイントに保存
# + id="usDm_Wpsl0bu"
train_checkpointer.save(global_step)
# + [markdown] id="gTQUrKgihuic"
# ### チェックポイントに復元
#
# チェックポイントに復元するためには、チェックポイントが作成されたときと同じ方法でオブジェクト全体を再作成する必要があります。
# + id="l6l3EB-Yhwmz"
train_checkpointer.initialize_or_restore()
global_step = tf.compat.v1.train.get_global_step()
# + [markdown] id="Nb8_MSE2XjRp"
# また、ポリシーを保存して指定する場所にエクスポートします。
# + id="3xHz09WCWjwA"
tf_policy_saver.save(policy_dir)
# + [markdown] id="Mz-xScbuh4Vo"
# ポリシーの作成に使用されたエージェントまたはネットワークについての知識がなくても、ポリシーを読み込めるので、ポリシーのデプロイが非常に簡単になります。
#
# 保存されたポリシーを読み込み、それがどのように機能するかを確認します。
# + id="J6T5KLTMh9ZB"
saved_policy = tf.saved_model.load(policy_dir)
run_episodes_and_create_video(saved_policy, eval_env, eval_py_env)
# + [markdown] id="MpE0KKfqjc0c"
# ## エクスポートとインポート
#
# 以下は、後でトレーニングを続行し、再度トレーニングすることなくモデルをデプロイできるように、Checkpointer とポリシーディレクトリをエクスポート/インポートするのに役立ちます。
#
# 「1回のイテレーションのトレーニング」に戻り、後で違いを理解できるように、さらに数回トレーニングします。 結果が少し改善し始めたら、以下に進みます。
# + id="fd5Cj7DVjfH4"
#@title Create zip file and upload zip file (double-click to see the code)
def create_zip_file(dirname, base_filename):
return shutil.make_archive(base_filename, 'zip', dirname)
def upload_and_unzip_file_to(dirname):
if files is None:
return
uploaded = files.upload()
for fn in uploaded.keys():
print('User uploaded file "{name}" with length {length} bytes'.format(
name=fn, length=len(uploaded[fn])))
shutil.rmtree(dirname)
zip_files = zipfile.ZipFile(io.BytesIO(uploaded[fn]), 'r')
zip_files.extractall(dirname)
zip_files.close()
# + [markdown] id="hgyy29doHCmL"
# チェックポイントディレクトリからzipファイルを作成します。
# + id="nhR8NeWzF4fe"
train_checkpointer.save(global_step)
checkpoint_zip_filename = create_zip_file(checkpoint_dir, os.path.join(tempdir, 'exported_cp'))
# + [markdown] id="VGEpntTocd2u"
# zipファイルをダウンロードします。
# + id="upFxb5k8b4MC"
#@test {"skip": true}
if files is not None:
files.download(checkpoint_zip_filename) # try again if this fails: https://github.com/googlecolab/colabtools/issues/469
# + [markdown] id="VRaZMrn5jLmE"
# 10〜15回ほどトレーニングした後、チェックポイントのzipファイルをダウンロードし、[ランタイム]> [再起動してすべて実行]に移動してトレーニングをリセットし、このセルに戻ります。ダウンロードしたzipファイルをアップロードして、トレーニングを続けます。
# + id="kg-bKgMsF-H_"
#@test {"skip": true}
upload_and_unzip_file_to(checkpoint_dir)
train_checkpointer.initialize_or_restore()
global_step = tf.compat.v1.train.get_global_step()
# + [markdown] id="uXrNax5Zk3vF"
# チェックポイントディレクトリをアップロードしたら、「1回のイテレーションのトレーニング」に戻ってトレーニングを続けるか、「ビデオ生成」に戻って読み込まれたポリシーのパフォーマンスを確認します。
# + [markdown] id="OAkvVZ-NeN2j"
# または、ポリシー(モデル)を保存して復元することもできます。Checkpointerとは異なり、トレーニングを続けることはできませんが、モデルをデプロイすることはできます。ダウンロードしたファイルはCheckpointerのファイルよりも大幅に小さいことに注意してください。
# + id="s7qMn6D8eiIA"
tf_policy_saver.save(policy_dir)
policy_zip_filename = create_zip_file(policy_dir, os.path.join(tempdir, 'exported_policy'))
# + id="rrGvCEXwerJj"
#@test {"skip": true}
if files is not None:
files.download(policy_zip_filename) # try again if this fails: https://github.com/googlecolab/colabtools/issues/469
# + [markdown] id="DyC_O_gsgSi5"
# ダウンロードしたポリシーディレクトリ(exported_policy.zip)をアップロードし、保存したポリシーの動作を確認します。
# + id="bgWLimRlXy5z"
#@test {"skip": true}
upload_and_unzip_file_to(policy_dir)
saved_policy = tf.saved_model.load(policy_dir)
run_episodes_and_create_video(saved_policy, eval_env, eval_py_env)
# + [markdown] id="HSehXThTm4af"
# ## SavedModelPyTFEagerPolicy
#
# TFポリシーを使用しない場合は、`py_tf_eager_policy.SavedModelPyTFEagerPolicy`を使用して、Python envでsaved_modelを直接使用することもできます。
#
# これは、eagerモードが有効になっている場合にのみ機能することに注意してください。
# + id="iUC5XuLf1jF7"
eager_py_policy = py_tf_eager_policy.SavedModelPyTFEagerPolicy(
policy_dir, eval_py_env.time_step_spec(), eval_py_env.action_spec())
# Note that we're passing eval_py_env not eval_env.
run_episodes_and_create_video(eager_py_policy, eval_py_env, eval_py_env)
# + [markdown] id="7fvWqfJg00ww"
# ## ポリシーを TFLite に変換する
#
# 詳細については、「[TensorFlow Lite 推論](https://tensorflow.org/lite/guide/inference)」をご覧ください。
# + id="z9zonVBJ0z46"
converter = tf.lite.TFLiteConverter.from_saved_model(policy_dir, signature_keys=["action"])
converter.target_spec.supported_ops = [
tf.lite.OpsSet.TFLITE_BUILTINS, # enable TensorFlow Lite ops.
tf.lite.OpsSet.SELECT_TF_OPS # enable TensorFlow ops.
]
tflite_policy = converter.convert()
with open(os.path.join(tempdir, 'policy.tflite'), 'wb') as f:
f.write(tflite_policy)
# + [markdown] id="rsi3V9QdxJUu"
# ### TFLite モデルで推論を実行する
# + id="4GeUSWyZxMlN"
import numpy as np
interpreter = tf.lite.Interpreter(os.path.join(tempdir, 'policy.tflite'))
policy_runner = interpreter.get_signature_runner()
print(policy_runner._inputs)
# + id="eVVrdTbRxnOC"
policy_runner(**{
'0/discount':tf.constant(0.0),
'0/observation':tf.zeros([1,4]),
'0/reward':tf.constant(0.0),
'0/step_type':tf.constant(0)})
| site/ja/agents/tutorials/10_checkpointer_policysaver_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="PO17wHPShbcw"
# NLTK 토큰화 하기
import nltk
# + colab={"base_uri": "https://localhost:8080/"} id="KK3-MsS6hezy" outputId="1b038d52-e81e-4ce8-8e9b-309a7354b8f1"
# NLTK 다운
nltk.download('punkt')
# + id="7ZZTB3EVhjxl"
sent = "don't worry, be happy. please wake up everbody come on"
# + colab={"base_uri": "https://localhost:8080/"} id="JkfS70Cjhqdo" outputId="7c3f9601-6284-48e3-c89d-7496d8fcb085"
from nltk.tokenize import word_tokenize
print(word_tokenize(sent))
# + colab={"base_uri": "https://localhost:8080/"} id="Sk-v-NWshzEU" outputId="9da6ccb4-c6c5-4d43-b26e-48985a2e387f"
word_tokenize(sent)
# + colab={"base_uri": "https://localhost:8080/"} id="ft6Nbq60iDFJ" outputId="abaad144-8449-4985-e2e4-8d836a13d0ac"
from nltk.tokenize import wordpunct_tokenize
print(wordpunct_tokenize(sent))
# + colab={"base_uri": "https://localhost:8080/"} id="1x2_By5piUBL" outputId="1b260eb9-a557-4de2-a655-7b3fbfdcffa4"
# !pip install konlpy
# + id="H_Qhp9X6iXvv"
from konlpy.tag import *
hannanum = Hannanum()
kkma = Kkma()
komoran = Komoran()
okt = Okt()
# + colab={"base_uri": "https://localhost:8080/"} id="jB4nsj0vimer" outputId="248ccd2a-456f-4c25-90b3-9bfa69f41009"
# 형태소
okt.nouns("아무말, 집에 갑니다, 배고파요, 잠 깨고 있어요, 취업, 커피 마시고 싶어요, 자리 뺏겼어요, 졸려요, 헉")
# + id="yKh04wj9jV8z"
test = "아무말, 집에 갑니다, 배고파요, 잠 깨고 있어요, 취업, 커피 마시고 싶어요, 자리 뺏겼어요, 졸려요, 헉"
# + colab={"base_uri": "https://localhost:8080/"} id="h8gHQRIajg2F" outputId="c187ab31-57dd-48f7-ab36-02b4bf4674dc"
okt.nouns(test)
# + colab={"base_uri": "https://localhost:8080/"} id="EYC1r1oxjiwR" outputId="ee8a4afa-52be-4c48-8698-6516550ad176"
okt.morphs(test) # 형태소
# + colab={"base_uri": "https://localhost:8080/"} id="mqG90jAHjmBV" outputId="76dc2cb3-c9f7-4893-d5b6-4b524d890e3e"
okt.pos(test) # 품사
# + colab={"base_uri": "https://localhost:8080/"} id="H0q1sSJKjsvd" outputId="90b3152f-1c35-45ee-ff73-adf0f9347ee4"
kkma.nouns(test)
# + colab={"base_uri": "https://localhost:8080/"} id="HlePi1g_j9l3" outputId="fc5e2e76-fb7e-4781-9743-379aec19a42d"
kkma.morphs(test)
# + colab={"base_uri": "https://localhost:8080/"} id="NJH37j_ukFAT" outputId="dc164b66-3796-42cb-cee3-3c8b9ddd24f6"
kkma.pos(test)
# + id="FhdWlyrAkJsx"
# sentense 문장 토큰화
text1 = 'Do the boogie like Side step right left to my beat High like the moon rock with me baby Know that I got that heat'
# + colab={"base_uri": "https://localhost:8080/"} id="UsYl8QAWkvAj" outputId="e42396c5-3cd9-4f5f-b3bb-df0f29ba672f"
from nltk.tokenize import sent_tokenize
sent_tokenize(text1)
# + id="a5Hz7ANVk4VD"
text2 = '사랑을 맺었던가 님들의 치맛 소리, 태욱스럽다'
# + colab={"base_uri": "https://localhost:8080/"} id="1cIYSwHvlfIH" outputId="4762d616-8b18-4614-e34a-fe6e4fc8a503"
# !pip install kss
# + colab={"base_uri": "https://localhost:8080/"} id="YSllg4zLlnA3" outputId="bd26ba20-b4dd-49bb-aee7-836318dc9b09"
import kss
kss.split_sentences(text2)
# + id="ZNHKZaW1lx3y"
# 텍스트 정규화
# 1. stem 스테밍, 어감 추출
from nltk.stem import PorterStemmer
ps = PorterStemmer() # 생성자 객체화
# + colab={"base_uri": "https://localhost:8080/"} id="uV8R753ymDa8" outputId="306accce-ff44-49bb-d465-75a6d1d03cb7"
word_tokenize(text1)
# + id="kIhCkLUImNTY"
words = word_tokenize(text1)
# + colab={"base_uri": "https://localhost:8080/"} id="p8g858gPmRlf" outputId="0bd3fffc-fbc1-4fe6-b953-fad01f3ad51a"
[ps.stem(w) for w in words] # list(), [] 이거 많이 씀
# + colab={"base_uri": "https://localhost:8080/"} id="5PR4OyeymgPy" outputId="59c3f7c8-7f47-4ff1-c7fb-54d927a62824"
# lemmatization
nltk.download('wordnet')
# + id="pynIR6mXm0q9"
from nltk.stem import WordNetLemmatizer
wl = WordNetLemmatizer()
# + id="OwBeEbTrm898"
words = ['policy', 'doing', 'organization', 'have']
# + colab={"base_uri": "https://localhost:8080/"} id="jVHFAs7GnKFA" outputId="248ceee1-7c0a-4ec2-a85f-accdabb65500"
[wl.lemmatize(w) for w in words]
# + colab={"base_uri": "https://localhost:8080/"} id="5rScLLKxnTUJ" outputId="36fd7368-e2d8-4aa5-f536-45e35fed696a"
okt.morphs(text2)
# + colab={"base_uri": "https://localhost:8080/"} id="zvYoRFNoneXz" outputId="f7ac69c4-a58d-4188-a4ff-9539aff3bf15"
okt.morphs(text2, stem = True) # 원형 찾기
# + colab={"base_uri": "https://localhost:8080/"} id="gACFqKKinrXC" outputId="bdbf4c6c-a130-40cc-a543-cffcae3e038c"
okt.pos(text2)
# + colab={"base_uri": "https://localhost:8080/"} id="Wj6PnMFonoTJ" outputId="01a417c0-edb9-4b4d-9b29-99486fc8e109"
# stopwords 불용어 사용하지 않는 것은 상대적. 쓸 수도 있고 쓰지 않을 수도 있고.
nltk.download('stopwords')
# + colab={"base_uri": "https://localhost:8080/"} id="IMMP3A2UoMAg" outputId="e2fc7f68-1104-47a2-8ccd-a99f1c7fa80d"
from nltk.corpus import stopwords #말뭉치: corpus
print(stopwords.words('english'))
# + colab={"base_uri": "https://localhost:8080/"} id="gg48l0nSoVnv" outputId="b55b0a31-dad9-4c88-a232-1c8135431d18"
ex = "Se is very import person. It's everytinog"
set(stopwords.words('english')) # set()은 중복은 없음
# + id="NJKrJqbuouc8"
stop_words = set(stopwords.words('english'))
# + colab={"base_uri": "https://localhost:8080/"} id="JrB0tYmWo3Kn" outputId="9848d123-45da-4a96-eb2b-0ddea9410993"
word_tokenize(ex)
# + id="8w4lqGVto5qD"
word_tokens = word_tokenize(ex)
# + colab={"base_uri": "https://localhost:8080/"} id="M9RCeo4Co_gE" outputId="fe543f99-fcbf-4e3c-8a3a-0bf9b0ff5f78"
result = []
for w in word_tokens:
if w not in stop_words:
result.append(w)
print('원문:', word_tokens)
print('불용어 제거문:', result)
# + id="H5qfwoZJpU5t"
example2 = '남자 소개시켜줘, 알지 남자라고 다 남자 아냐, 재밌네요.'
stop_words2 = '아냐 알지'
stop_words2 = stop_words2.split(' ')
word_tokens2 = word_tokenize(example2)
# + colab={"base_uri": "https://localhost:8080/"} id="euext_YQzZ7e" outputId="bd13fe91-bb6a-4e6e-9b25-36cd8878fac3"
[w for w in word_tokens2 if w not in stop_words2] # list comperation
# + colab={"base_uri": "https://localhost:8080/"} id="ABN4C2qe0EZn" outputId="5842e420-7974-4287-d566-1847f32e5507"
result = [w for w in word_tokens2 if w not in stop_words2]
print('원문: ', word_tokens2)
print('불용어 제거 후: ', result)
# + id="vGp0ahsW0ewR"
# 정수 인코딩
text3 = "A barber is a person. a barber is good person. a barber is huge person. \
he Knew A Secret! The Secret He Kept is huge secret. Huge secret. His barber kept his word. a barber kept his word. \
His barber kept his secret. But keeping and keeping such a huge secret to himself was driving the barber crazy. \
the barber went up a huge mountain."
# + colab={"base_uri": "https://localhost:8080/"} id="Qo3NqhEF1AVS" outputId="01c355b9-820e-46e1-da72-68150ddf5d74"
# sentence tokneization
sent_tokenize(text3)
# + colab={"base_uri": "https://localhost:8080/"} id="Bffjkcp41ZFG" outputId="8c6b243f-18ae-44e0-8d73-00ee3fc0350f"
text3 = sent_tokenize(text3)
print(text3)
# + colab={"base_uri": "https://localhost:8080/"} id="H_ITAjUB1gdU" outputId="ba976d0d-4303-403d-c5d4-bcf028242cc0"
# word tokenization
# 이중for문
# 리스트의 리스트 (리스트 내포)
sentences = [ ]
stop_words = set(stopwords.words('english'))
for i in text3:
sentence = word_tokenize(i) # sentence 구분
result = [ ]
for word in sentence:
word = word.lower()
if word not in stop_words:
if len(word) > 2:
result.append(word)
sentences.append(result)
print(sentences)
# + colab={"base_uri": "https://localhost:8080/"} id="g4ixXW1o2TOr" outputId="3123ff17-c213-4c95-9482-68f53490fa3b"
# 단어집합 vocabulary
from collections import Counter
words = sum(sentences, [])
print(words)
# + colab={"base_uri": "https://localhost:8080/"} id="Qm0obp8r36iG" outputId="82025a0a-ac0d-4c93-cb08-29f18121eeb4"
print(sum(sentences, []))
# + [markdown] id="fOkHrf1o-kJi"
# https://wikidocs.net/31766 정수인코딩
# + colab={"base_uri": "https://localhost:8080/"} id="ViZTvamR-Z5A" outputId="746389a1-ef3c-4094-babe-070713012dba"
Counter(words)
# + colab={"base_uri": "https://localhost:8080/"} id="dKb7cdFr_IXX" outputId="55a11cfa-26cf-4687-f9a4-89c2c93f244a"
vocab = Counter(words)
print(vocab)
# + colab={"base_uri": "https://localhost:8080/"} id="6_Gx8LiB_Xyi" outputId="6a424bb0-9fcc-4573-ff57-aacf503814f0"
print(vocab['huge'])
# + colab={"base_uri": "https://localhost:8080/"} id="E6dBpa3m_hOR" outputId="1e8c97e5-cb56-4707-b627-2ff5919745f6"
# 정수 인덱스 (integer encoding)
# 빈도수가 높을수록 낮은 수의 정수 부여
sorted(vocab.items(), key = lambda x: x[1]) # x[0]: key, x[1]: value
# + colab={"base_uri": "https://localhost:8080/"} id="ZLXiWEu_ABh7" outputId="5bd6157e-b32f-436a-c541-21f71d1d87a9"
sorted(vocab.items(), key = lambda x: x[1], reverse = True) # x[0]: key, x[1]: value
# + id="RcFo8bK7AqLD"
vocab_sorted = sorted(vocab.items(), key = lambda x: x[1], reverse = True) # x[0]: key, x[1]: value
# + colab={"base_uri": "https://localhost:8080/"} id="lLPJOOc3A1TY" outputId="fa7a6594-2309-4de3-8b73-6973ba6294f8"
word2idx = { }
i = 0
for (word, frequency) in vocab_sorted:
if frequency > 1:
i = i + 1
word2idx[word] = i
print(word2idx)
# 케라스 라벨 인코더는 알아서 이걸 해줌
# + colab={"base_uri": "https://localhost:8080/"} id="PdwaC0r3BTLD" outputId="1320789f-7272-4b7e-b2c3-0e32c0314462"
vocab_size = 5
word_frequency = [w for w,c in word2idx.items() if c >= vocab_size + 1 ] # 빈도수 높은 단어
for w in word_frequency:
del word2idx[w]
print(word2idx)
# 마스킹 기법
# 정수 인덱스랑 반대니까, 빈도수 많은거 상위 5개만 정하기 위한 코드
# 빈도수 적은 거 제거하는 기법
# + colab={"base_uri": "https://localhost:8080/"} id="MWJRWVSMCh3j" outputId="2c123024-4f32-451c-d9aa-0fb6e10fa601"
len(word2idx) + 1
# + id="tY5nKRgJD8AG"
word2idx['oov'] = len(word2idx) + 1 # out of vocab
# + colab={"base_uri": "https://localhost:8080/"} id="dYAUXU6AEDNq" outputId="9d541056-c84c-4efe-8d75-e4ecbb7aa2b9"
print(word2idx) # 딕셔너리
# + colab={"base_uri": "https://localhost:8080/"} id="nvBqGCkPEG7j" outputId="ef4fde79-b9ce-4509-b6b7-495b3a844b0f"
sentences # 길이가 다 다르네?!
# + colab={"base_uri": "https://localhost:8080/"} id="eGXIncmcEMrX" outputId="180ad254-8b35-4563-d376-c26874f98223"
encoded = [ ]
for s in sentences:
temp = []
for w in s: # 이중for문
try:
temp.append(word2idx[w])
except KeyError:
temp.append(word2idx['oov'])
encoded.append(temp)
encoded
# + id="sUiZ4gQ0Emor"
| practice/2021_06_23_wed.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ewuerfel66/DS-Unit-1-Sprint-3-Statistical-Tests-and-Experiments/blob/master/EricWuerfel_DS_Unit_1_Sprint_Challenge_3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="NooAiTdnafkz" colab_type="text"
# # Data Science Unit 1 Sprint Challenge 4
#
# ## Exploring Data, Testing Hypotheses
#
# In this sprint challenge you will look at a dataset of people being approved or rejected for credit.
#
# https://archive.ics.uci.edu/ml/datasets/Credit+Approval
#
# Data Set Information: This file concerns credit card applications. All attribute names and values have been changed to meaningless symbols to protect confidentiality of the data. This dataset is interesting because there is a good mix of attributes -- continuous, nominal with small numbers of values, and nominal with larger numbers of values. There are also a few missing values.
#
# Attribute Information:
# - A1: b, a.
# - A2: continuous.
# - A3: continuous.
# - A4: u, y, l, t.
# - A5: g, p, gg.
# - A6: c, d, cc, i, j, k, m, r, q, w, x, e, aa, ff.
# - A7: v, h, bb, j, n, z, dd, ff, o.
# - A8: continuous.
# - A9: t, f.
# - A10: t, f.
# - A11: continuous.
# - A12: t, f.
# - A13: g, p, s.
# - A14: continuous.
# - A15: continuous.
# - A16: +,- (class attribute)
#
# Yes, most of that doesn't mean anything. A16 (the class attribute) is the most interesting, as it separates the 307 approved cases from the 383 rejected cases. The remaining variables have been obfuscated for privacy - a challenge you may have to deal with in your data science career.
#
# Sprint challenges are evaluated based on satisfactory completion of each part. It is suggested you work through it in order, getting each aspect reasonably working, before trying to deeply explore, iterate, or refine any given step. Once you get to the end, if you want to go back and improve things, go for it!
# + [markdown] id="Grf4qbAPhDS_" colab_type="text"
# ## Imports
# + id="zpmkaN7OhIw5" colab_type="code" colab={}
import pandas as pd
import numpy as np
import seaborn as sns
import scipy
from statistics import mean, median
from scipy.stats import ttest_ind
# + [markdown] id="5wch6ksCbJtZ" colab_type="text"
# ## Part 1 - Load and validate the data
#
# - Load the data as a `pandas` data frame.
# - Validate that it has the appropriate number of observations (you can check the raw file, and also read the dataset description from UCI).
# - UCI says there should be missing data - check, and if necessary change the data so pandas recognizes it as na
# - Make sure that the loaded features are of the types described above (continuous values should be treated as float), and correct as necessary
#
# This is review, but skills that you'll use at the start of any data exploration. Further, you may have to do some investigation to figure out which file to load from - that is part of the puzzle.
# + [markdown] id="I_77uigly3Qq" colab_type="text"
# ### Load Data & Validate Observations
# + id="Q79xDLckzibS" colab_type="code" colab={}
df = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/credit-screening/crx.data', header = None)
# + id="GeN9_7yIh_TE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="b531c543-44e2-4df4-f1de-d1b384cc6290"
df.shape
# + [markdown] id="FkQe0uogiECI" colab_type="text"
# The dataframe has **the correct number of rows**
#
# According to UCI, there are 690 instances in the dataset and we have 690 rows.
# + [markdown] id="0mY2E33vy_KD" colab_type="text"
# ### Find & Replace Missing Values
# + id="pAyJZeHfhVwi" colab_type="code" colab={}
df = df.replace(to_replace = '+', value = 1)
df = df.replace(to_replace = '-', value = 0)
# + id="IuzsPM7Tj5Dz" colab_type="code" colab={}
df = df.rename(columns={15:'status'})
# + id="NOArNHd-ib-E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="ca225446-95e9-4c2c-8e4e-d830f1a6b555"
df.isnull().sum()
# + id="hHsqeh6phRtv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="a91eaaaa-2ed3-4fc5-d799-0cc2d18b440e"
df['status'].value_counts()
# + [markdown] id="1CdPCUITi7EL" colab_type="text"
# We only have 1s and 0s in 'status'.
# There isn't any missing data in the column we're interested in
# + [markdown] id="_ubi7e5bkzYj" colab_type="text"
# There's our missing value: '?'
# + id="d7DTTP6xkvko" colab_type="code" colab={}
df = df.replace(to_replace = '?', value = np.NaN)
# + [markdown] id="ag1q0jjQkP5-" colab_type="text"
# ### Check and Fix Data Types
#
# col 1,
# col 10,
# col 13,
# col 14 --> float
# + id="F9wG30uLznU1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="fb978197-cada-4d59-cf7b-1d8e43e2704f"
df.dtypes
# + id="pqzhl4GXlwRV" colab_type="code" colab={}
df[1] = df[1].astype(float)
# + id="R_8b1Cu9oBJM" colab_type="code" colab={}
df[10] = df[10].astype(float)
# + id="OIcwO3P6ntpO" colab_type="code" colab={}
df[13] = df[13].astype(float)
# + id="MzMYiHeLoG-R" colab_type="code" colab={}
df[14] = df[14].astype(float)
# + id="dQOnxWkvn7gB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="88480aea-84e2-4f65-89f1-b43796f1abc7"
df.dtypes
# + id="tQaFxKUzjfL3" colab_type="code" outputId="e501fa44-c646-42c8-b0d3-85015c4b2559" colab={"base_uri": "https://localhost:8080/", "height": 0}
df.head()
# + [markdown] id="G7rLytbrO38L" colab_type="text"
# ## Part 2 - Exploring data, Testing hypotheses
#
# The only thing we really know about this data is that A16 is the class label. Besides that, we have 6 continuous (float) features and 9 categorical features.
#
# Explore the data: you can use whatever approach (tables, utility functions, visualizations) to get an impression of the distributions and relationships of the variables. In general, your goal is to understand how the features are different when grouped by the two class labels (`+` and `-`).
#
# For the 6 continuous features, how are they different when split between the two class labels? Choose two features to run t-tests (again split by class label) - specifically, select one feature that is *extremely* different between the classes, and another feature that is notably less different (though perhaps still "statistically significantly" different). You may have to explore more than two features to do this.
#
# For the categorical features, explore by creating "cross tabs" (aka [contingency tables](https://en.wikipedia.org/wiki/Contingency_table)) between them and the class label, and apply the Chi-squared test to them. [pandas.crosstab](http://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.crosstab.html) can create contingency tables, and [scipy.stats.chi2_contingency](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.chi2_contingency.html) can calculate the Chi-squared statistic for them.
#
# There are 9 categorical features - as with the t-test, try to find one where the Chi-squared test returns an extreme result (rejecting the null that the data are independent), and one where it is less extreme.
#
# **NOTE** - "less extreme" just means smaller test statistic/larger p-value. Even the least extreme differences may be strongly statistically significant.
#
# Your *main* goal is the hypothesis tests, so don't spend too much time on the exploration/visualization piece. That is just a means to an end - use simple visualizations, such as boxplots or a scatter matrix (both built in to pandas), to get a feel for the overall distribution of the variables.
#
# This is challenging, so manage your time and aim for a baseline of at least running two t-tests and two Chi-squared tests before polishing. And don't forget to answer the questions in part 3, even if your results in this part aren't what you want them to be.
# + [markdown] id="Dc1UGVozoYIK" colab_type="text"
# ### Split
# + id="8J__9KF_kurC" colab_type="code" colab={}
approved = df[df['status'] == 1.]
denied = df[df['status'] == 0.]
# + [markdown] id="iL2-LENIoxVC" colab_type="text"
# ### Continuous Variables
#
# 1, 2, 7, 10, 13, 14
#
# Blue is Approved, Orange is Denied
# + [markdown] id="dvHET6baz3tR" colab_type="text"
# #### Visualize the Distributions (Histograms)
# + id="jjNfOXlrpqSM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="1682937b-6cec-41e3-9a10-cfa98b0bd8e5"
app_1 = approved[1].dropna().tolist()
den_1 = denied[1].dropna().tolist()
ax = sns.distplot(app_1, kde=False);
ax = sns.distplot(den_1, kde=False);
# + id="V4bh8JPGqBb3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="0fab1128-bf52-4e62-d798-97a598f134c0"
app_2 = approved[2].dropna().tolist();
den_2 = denied[2].dropna().tolist();
ax = sns.distplot(app_2, kde=False);
ax = sns.distplot(den_2, kde=False);
# + id="rMQKlFosqBn2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="0a9cba86-e3ca-47ad-aa37-92932fdc6f6e"
app_7 = approved[7].dropna().tolist();
den_7 = denied[7].dropna().tolist();
ax = sns.distplot(app_7, kde=False);
ax = sns.distplot(den_7, kde=False);
# + id="0zUy6ZB1qBvw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 270} outputId="0fefb13d-a36b-4adc-b624-19d1bb9043db"
app_10 = approved[10].dropna().tolist();
den_10 = denied[10].dropna().tolist();
ax = sns.distplot(app_10, kde=False);
ax = sns.distplot(den_10, kde=False);
# + id="ly8c82tvqB3W" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="2f1757ea-4a92-4c18-b6e3-a46547bf4ff8"
app_13 = approved[13].dropna().tolist();
den_13 = denied[13].dropna().tolist();
ax = sns.distplot(app_13, kde=False);
ax = sns.distplot(den_13, kde=False);
# + id="u0q1_nkgq6CG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="c49ab1a2-691a-4b25-84f6-6864b7c6b7d9"
app_14 = approved[14].dropna().tolist();
den_14 = denied[14].dropna().tolist();
ax = sns.distplot(app_14, kde=False);
ax = sns.distplot(den_14, kde=False);
# + [markdown] id="oiWFQuIxz8bi" colab_type="text"
# #### t-tests
# + [markdown] id="FEXgJ3rXrhs7" colab_type="text"
# Variables **10 and 14** seem to show the biggest differences between the two groups
# + id="r-HBnXGyrppS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="2f535f3e-1b60-4995-8be4-b8cdc6af955c"
ttest_ind(approved[10], denied[10], nan_policy='omit')
# + [markdown] id="fQFjoGSksyCH" colab_type="text"
# The positive t-statistic, and p-value << 0.01 indicate that we can reject the null hypothesis that the mean of column 10 for borrowers who were approved is the same as the mean of column 10 for borrowers who were denied.
# + id="1uBe8xMZtIbd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="02880ea3-6f35-46aa-98fb-86eba1e23893"
ttest_ind(approved[14], denied[14], nan_policy='omit')
# + [markdown] id="yVTdrm1StQlz" colab_type="text"
# The positive t-statistic, and p-value << 0.01 indicate that we can reject the null hypothesis that the mean of column 14 for borrowers who were approved is the same as the mean of column 14 for borrowers who were denied.
# + [markdown] id="gkj2llNatV1J" colab_type="text"
# ### Categorical Variables
# + [markdown] id="gewtyTsF0Mt-" colab_type="text"
# #### Visualize (Crosstabs)
# + id="bZE0dTWgtcKl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="02c62594-8fee-4796-e698-ce76e254487b"
pd.crosstab(df['status'], df[3], margins=True)
# + id="119xl7HtuDIw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="efdef7d4-fca2-42f0-a60f-b58699b0da0b"
pd.crosstab(df['status'], df[4], margins=True)
# + id="pSjUnj_LuDPO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="76f4bade-4c8b-46db-c886-ef226b589851"
pd.crosstab(df['status'], df[5], margins=True)
# + id="Qnzq9660uDVv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="aca782ea-6e12-42e1-853f-ec025a8c3116"
pd.crosstab(df['status'], df[6], margins=True)
# + id="3CIbheoCuDcX" colab_type="code" colab={}
contingency_8 = pd.crosstab(df['status'], df[8], margins=True)
# + id="h0IMebYeuDi-" colab_type="code" colab={}
contingency_9 = pd.crosstab(df['status'], df[9], margins=True)
# + id="85WDbySuuDqO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="31d6ccca-dbe0-4121-ee1f-7bb90e166e7b"
pd.crosstab(df['status'], df[11], margins=True)
# + id="jdQa2YJouDxG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="88436d14-983e-4649-80cc-1c28fefee361"
pd.crosstab(df['status'], df[12], margins=True)
# + [markdown] id="qVgIJyvW0ijL" colab_type="text"
# #### chi2 tests
# + id="4c7qyYohulJw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="93930244-89e6-49a7-eae9-4fe0df445238"
scipy.stats.chi2_contingency(contingency_8)
# + [markdown] id="uRzYdYd4vfru" colab_type="text"
# With a p-value << 0.01 we can reject the null hypothesis that variable 8 and a borrower's approval status are independent.
# + id="s29qbVg_vfAb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="21a7e0a0-b5ed-41cf-8875-ef64362e00ec"
scipy.stats.chi2_contingency(contingency_9)
# + [markdown] id="_WQWujuZwGDA" colab_type="text"
# Th ep-value for this chi2 test isn't nearly as extreme as it was for variable 8, but it's still << 0.01. This allows us to reject the null hypothesis that variable 9 and a borrower's approval status are independent.
# + [markdown] id="ZM8JckA2bgnp" colab_type="text"
# ## Part 3 - Analysis and Interpretation
#
# Now that you've looked at the data, answer the following questions:
#
# - Interpret and explain the two t-tests you ran - what do they tell you about the relationships between the continuous features you selected and the class labels?
# - Interpret and explain the two Chi-squared tests you ran - what do they tell you about the relationships between the categorical features you selected and the class labels?
# - What was the most challenging part of this sprint challenge?
#
# Answer with text, but feel free to intersperse example code/results or refer to it from earlier.
# + [markdown] id="LIozLDNG2Uhu" colab_type="text"
# ### t-test: Variable 10
#
# test-statistic: 11.7
#
# p-value: $7.96 * 10^{-29}$
#
# The borrowing data was first split in two according to the approval status of the borrower. From observing the two resulting distributions of Variable 10 led me to believe that the mean of Variable 10 for those who were approved would be higher than that for those who were denied.
#
# The t-test returned a positive test-statistic, meaning that the calculated mean of Variable 10 for those approved is higher than the calculated mean for those who were denied. But is it **significantly** higher?
#
# The p-value of $7.96 * 10^{-29}$ tells us that there is an extraordinarily small chance that the difference we observed in the means is due to chance.
#
# We can say that the mean of Variable 10 is higher for those approved than it is for those denied, with 99% confidence.
# + [markdown] id="B_i-rwy08Cun" colab_type="text"
# ### t-test: Variable 14
#
# test-statistic: 4.68
#
# p-value: $3.45 * 10^{-6}$
#
# From the distributions, it seems that the mean for Variable 14 is higher for those who were accepted than it is for those who were denied.
#
# The t-test again returned a positive statistic, confirming that the direction of my observation is at least correct.
#
# The p-value of $3.45 * 10^{-6}$ tells us that there is a wildly small probability that the difference in means is due to chance.
#
# We can say that the mean of Variable 14 is higher for those approved than for those denied.
# + [markdown] id="aHDXCbjo83eo" colab_type="text"
# ### chi2-test: Variable 8
#
# chi2: 358
#
# p-value: $3.13 * 10^{-76}$
#
# From the crosstabs, it looked like many more of those who were approved belonged to 't' than belonged to 'f'.
#
# To confirm this suspicion, I ran a chi2 test. The extraordinarily low p-value allows us to reject the null hypothesis that approval status and Variable 8 are independent.
# + id="0ZpfVlqN9pJ7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="617e0c8e-4b27-4b15-96ed-1362251bd305"
contingency_8
# + [markdown] id="XY0ir3LE-XF5" colab_type="text"
# ### chi2-test: Variable 9
#
# chi2: 145
#
# p-value: $2.49 * 10^{-30}$
#
# The crosstab shows that a higher proportion of those approved are in 't' than in 'f'. Vice versa for those who were denied.
#
# The chi2 test returns a p-value of $2.49 * 10^{-30}$ which is again, far less than 0.01. This allows us to reject the null hypothesis that approval status and Variable 9 are independent.
# + id="IEYnX8-C-ZwN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="72109431-c8b2-4d7f-d71a-97477d1d714e"
contingency_9
# + [markdown] id="6fycXcTA_YqV" colab_type="text"
# The most difficult part of this sprint challenge was finding the best way to search for differences in the continuous variables. Luckily, there are a few variables whose differences between approval status is pretty obvious. Once I had the data cleaned up and split into groups based on approval status, performing the tests was pretty simple.
#
# Explaining the results of t-tests and chi2-tests in meaningful ways is a challenge as well.
| EricWuerfel_DS_Unit_1_Sprint_Challenge_3.ipynb |